code stringlengths 2.5k 150k | kind stringclasses 1 value |
|---|---|
```
import os
import sys
import torch
import gpytorch
from tqdm.auto import tqdm
import timeit
if os.path.abspath('..') not in sys.path:
sys.path.insert(0, os.path.abspath('..'))
from gpytorch_lattice_kernel import RBFLattice as BilateralKernel
# device = "cuda" if torch.cuda.is_available() else "cpu"
device = "cpu"
N_vals = torch.linspace(100, 10000000, 10).int().tolist()
D_vals = torch.linspace(1, 100, 10).int().tolist()
```
# Matmul
```
N_vary = []
for N in tqdm(N_vals):
D = 1
x = torch.randn(N, D).to(device)
K = BilateralKernel().to(device)(x)
v = torch.randn(N, 1).to(device)
def matmul():
return K @ v
time = timeit.timeit(matmul , number=10)
N_vary.append([N, D, time])
del x
del K
del v
del matmul
D_vary = []
for D in tqdm(D_vals):
N = 1000
x = torch.randn(N, D).to(device)
K = BilateralKernel().to(device)(x)
v = torch.randn(N, 1).to(device)
def matmul():
return K @ v
time = timeit.timeit(matmul , number=10)
D_vary.append([N, D, time])
del x
del K
del v
del matmul
import pandas as pd
N_vary = pd.DataFrame(N_vary, columns=["N", "D", "Time"])
D_vary = pd.DataFrame(D_vary, columns=["N", "D", "Time"])
import seaborn as sns
ax = sns.lineplot(data=N_vary, x="N", y="Time")
ax.set(title="Matmul (D=1)")
from sklearn.linear_model import LinearRegression
import numpy as np
regr = LinearRegression()
regr.fit(np.log(D_vary["D"].to_numpy()[:, None]), np.log(D_vary["Time"]))
print('Coefficients: \n', regr.coef_)
pred_time = regr.predict(np.log(D_vary["D"].to_numpy()[:, None]))
ax = sns.lineplot(data=D_vary, x="D", y="Time")
ax.set(title="Matmul (N=1000)", xscale="log", yscale="log")
ax.plot(D_vary["D"].to_numpy(), np.exp(pred_time))
```
# Gradient
```
N_vary = []
for N in tqdm(N_vals):
D = 1
x = torch.randn(N, D, requires_grad=True).to(device)
K = BilateralKernel().to(device)(x)
v = torch.randn(N, 1, requires_grad=True).to(device)
sum = (K @ v).sum()
def gradient():
torch.autograd.grad(sum, [x, v], retain_graph=True)
x.grad = None
v.grad = None
return
time = timeit.timeit(gradient, number=10)
N_vary.append([N, D, time])
del x
del K
del v
del gradient
D_vary = []
for D in tqdm(D_vals):
N = 1000
x = torch.randn(N, D, requires_grad=True).to(device)
K = BilateralKernel().to(device)(x)
v = torch.randn(N, 1, requires_grad=True).to(device)
sum = (K @ v).sum()
def gradient():
torch.autograd.grad(sum, [x, v], retain_graph=True)
x.grad = None
v.grad = None
return
time = timeit.timeit(gradient, number=10)
D_vary.append([N, D, time])
del x
del K
del v
del gradient
import pandas as pd
N_vary = pd.DataFrame(N_vary, columns=["N", "D", "Time"])
D_vary = pd.DataFrame(D_vary, columns=["N", "D", "Time"])
import seaborn as sns
ax = sns.lineplot(data=N_vary, x="N", y="Time")
ax.set(title="Gradient computation of (K@v).sum() (D=1)")
from sklearn.linear_model import LinearRegression
import numpy as np
regr = LinearRegression()
regr.fit(np.log(D_vary["D"].to_numpy()[:, None]), np.log(D_vary["Time"]))
print('Coefficients: \n', regr.coef_)
pred_time = regr.predict(np.log(D_vary["D"].to_numpy()[:, None]))
ax = sns.lineplot(data=D_vary, x="D", y="Time")
ax.set(title="Gradient computation of (K@v).sum() (N=100)", xscale="log", yscale="log")
ax.plot(D_vary["D"].to_numpy(), np.exp(pred_time))
```
| github_jupyter |
# Some simple computations
The main purpose of this notebook is to illustrate a bit of relatively simple Python code - the kind of code that I expect folks in my numerical analysis classes to be able to read, understand, and emulate. There is some mathematics behind the code that we'll deal with later.
## The very basics
I suppose we outta be able to add two and two!
```
2+2
```
We could define and evaluate a function.
```
def f(x): return x**2
[f(x) for x in range(5)]
```
Many functions must be loaded from a package.
```
from numpy import sin,pi,arange
[sin(x) for x in arange(-5*pi/2,5*pi/2,pi/2)]
```
Often, we'll just import the whole package into a namespace and then prepend the namespace. Thus, the previous input could be typed as follows:
```
import numpy as np
[np.sin(x) for x in np.arange(-5*np.pi/2,5*np.pi/2,np.pi/2)]
```
While this might seem like a bit of a pain, it's a nice way to avoid name clashes when working with multiple packages. We'll need to do that when we graphing functions, for example.
```
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
x = np.linspace(-2*np.pi,2*np.pi,100)
y = np.sin(x)
plt.plot(x,y)
```
## A bit more
Here's the basic problem: Solve the equation $\cos(x)=x$. Like a lot of interesting problems, this is very easy to state and understand, but quite hard to solve. It's *impossible* to solve symbolically, in fact! As we'll learn in a few weeks, though, there are fabulous algorithms to find *numerical approximations* to solutions. One of these, called Newton's method, searches for a single *root* of a function close to an initial guess. This is an absolutely fundamental technique in numerical analysis and is implemented in Scipy's `optimize` module as `newton`. Here's how to invoke it to find a solution to our equation near the initial guess $x_0=1$.
```
# import the Numpy library, where the cosine is defined.
import numpy as np
# import newton
from scipy.optimize import newton
# define the function whose root is the solution we seek
def f(x): return np.cos(x)-x
# invoke newton
newton(f,1)
```
The comments and spaces are not necessary, so that's really just four lines of code.
Like many algorithms, Newton's algorithm is iterative. As it turns out in fact, we can solve the equation by simply iterating the cosine function! We'll learn a bit later why this works but here's how to implement the idea in Python.
```
x1 = 1
x2 = np.cos(1)
cnt = 0
while np.abs(x1-x2) > 10**(-8) and cnt < 100:
x1 = x2
x2 = np.cos(x2)
cnt = cnt + 1
(x2,cnt)
```
I guess this says that, after 45 iterates, the difference between $x$ and $\cos(x)$ is less than $1/10^8$. So, we're pretty close.
The code so far illustrates that there's a couple of libraries that we'll use a lot - NumPy (which implements a lot of relatively low level functionality) and SciPy (which implements higher level functionality built on top of NumPy).
Very often, it's nice to visualize your results. There are a number of very nice Python libraries for visualization but one of the most popular and widely used is called Matplotlib. In addition, Matplotlib is included in Anaconda, so that's what we'll use for the most part.
My expectation is that you can at least do a little basic plotting with Matplotlib. Here's how to $f(x)=\cos(x)$ together with the line $y=x$, as well as the point where they are equal.
```
# A cell magic to run Matplotlib inside the notebook.
# Not totally necessary, but sometimes nice.
%matplotlib inline
# Import the plotting module
import matplotlib.pyplot as plt
# Set up the x-values we wish to plot.
# 100 of them evenly distributed over [-0.2,1.2]
xs = np.linspace(-0.2,1.2,100)
# Plot the cosine function over those x-values
plt.plot(xs,np.cos(xs))
# Plot the line y=x
plt.plot(xs,xs)
# Plot the soution as a red dot.
plt.plot(x2,x2,'ro')
```
If we keep track of our orbit as we progress, we can go a bit farther and illustrate the convergence of the orbit with a [cobweb plot](https://en.wikipedia.org/wiki/Cobweb_plot).
```
x0 = .1
x1 = x0
orbitx = [x1]
orbity = [x1]
x2 = np.cos(x1)
cnt = 0
while np.abs(x1-x2) > 10**(-8) and cnt < 100:
orbitx.extend([x1,x2])
orbity.extend([x2,x2])
x1 = x2
x2 = np.cos(x2)
cnt = cnt + 1
xs = np.linspace(-0.2,1.2,100)
plt.plot(orbitx,orbity)
plt.plot(xs,np.cos(xs), 'k', linewidth=2)
plt.plot(xs,xs, 'k')
plt.plot(x0,x0,'go')
plt.plot(x2,x2,'ro')
```
In my own images, I will often go a bit overboard with options to specify things exactly the way I want. I emphasize, this level of graphic detail is not your responsibility!
```
import matplotlib as mpl
plt.plot(orbitx,orbity)
plt.plot(xs,np.cos(xs), 'k', linewidth=2)
plt.plot(xs,xs, 'k')
plt.plot(x0,x0,'go')
plt.plot(x2,x2,'ro')
ax = plt.gca()
ax.set_aspect(1)
ax.set_ylim(-0.1,1.1)
ax.set_xlim(-0.1,1.1)
ax.set_xticks([1/2,1])
ax.set_yticks([1/2,1])
ax.spines['left'].set_position('zero')
ax.spines['right'].set_color('none')
ax.spines['bottom'].set_position('zero')
ax.spines['top'].set_color('none')
xaxis,yaxis = ax.findobj(mpl.axis.Axis)
xticks = xaxis.get_major_ticks()
for tick in xticks:
tick.get_children()[1].set_color('w')
yticks = yaxis.get_major_ticks()
for tick in yticks:
tick.get_children()[1].set_color('w')
```
## A disturbing computation
Is addition commutative?
```
0.1 + (0.2 + -0.3)
(0.1+0.2) + -0.3
```
I guess that addition is *not* commutative.
| github_jupyter |
### Linear Classifier Plot
```
import matplotlib.pyplot as plt
import pandas as pd
circles = [15, 10, 15, 16, 15, 18, 20, 20]
crosses = [3, 0.5, 1.2, 3, 2.5, 6.2, 3, 8.3]
def line(m, x, d):
return m*x+d
line1 = []
for i in range(0,10):
line1.append(line(2,i,3.9))
print(line1)
plt.plot( range(0, len(circles) ), circles, "bo")
plt.plot( range(0, len(crosses) ), crosses, "rx")
plt.plot( range(0, len(line1) ), line1, "b-")
plt.show()
##Neural Network - MPLClassifier (Multi-Layer-Perceptron)
from sklearn.datasets import load_digits
from sklearn.cross_validation import train_test_split
from sklearn.neural_network import MLPClassifier
from sklearn.metrics import classification_report, confusion_matrix
import matplotlib.pyplot as plt
#Import Data
digits = load_digits()
print(digits.data)
print(digits.target)
X = digits.data
Y = digits.target
X_train, X_test, y_train, y_test = train_test_split(X, Y, train_size=0.05)
print(len(X_train),"-",len(X_test))
#Create Neural Network
neuralnetwork = MLPClassifier(hidden_layer_sizes = (100, ),
max_iter = 200,
activation = 'logistic',
learning_rate = 'adaptive',
verbose = True)
neuralnetwork.fit(X_train, y_train)
y_prediction = neuralnetwork.predict(X_test)
print(classification_report(y_test, y_prediction))
"""
plt.gray()
plt.matshow(digits.images[12])
plt.show()
"""
"""
from yahoo_finance import Share
tesla = Share('TSLA')
print(dir(tesla))
print(tesla.get_historical('2016-09-22', '2017-08-22'))
"""
def get_y(change):
if change > 0:
return 1
elif change <= 0:
return 0
filename_csv = "../datasets/TSLA.csv"
csv_tesla_data = pd.read_csv(filename_csv)
csv_tesla_data['Change'] = csv_tesla_data['Close']-csv_tesla_data['Open']
csv_tesla_data['Change_p'] = csv_tesla_data['Change']/csv_tesla_data['Open']*100
csv_tesla_data['Target'] = range(0, len(csv_tesla_data))
#print(get_y(csv_tesla_data['Change_p']))
#csv_tesla_data['Target'] = get_y(csv_tesla_data['Change_p'])
for i in range(0, len(csv_tesla_data)):
if i == 0:
continue
csv_tesla_data['Target'][i] = get_y(csv_tesla_data['Change_p'][i-1])
print(csv_tesla_data)
f, axarr = plt.subplots(2, sharex=True, sharey=False)
axarr[0].plot(range(1,len(csv_tesla_data)+1), csv_tesla_data['Close'], 'b-')
axarr[1].plot(range(1,len(csv_tesla_data)+1), csv_tesla_data['Change_p'], 'r-')
plt.show()
classifier_x = csv_tesla_data[['Open', 'High', 'Low', 'Close', 'Volume', 'Change', 'Change_p']]
classifier_y = csv_tesla_data['Target']
#train_test_split
X_train, X_test, y_train, y_test = train_test_split(classifier_x,
classifier_y)
#Fit the model
neuralnetwork = MLPClassifier(hidden_layer_sizes = (100, ),
max_iter = 200,
activation = 'logistic',
learning_rate = 'adaptive',
verbose = True)
neuralnetwork.fit(X_train, y_train)
y_prediction = neuralnetwork.predict(X_test)
print(y_test, " - ", y_prediction)
print(classification_report(y_test, y_prediction))
print(confusion_matrix(y_test, y_prediction))
#plot the predictions
```
| github_jupyter |
# Checking the OA status of an author's papers
```
import json
import orcid
import requests
import sys
import time
from IPython.display import HTML, display
```
ORCID of the author you want to check
```
ORCID = '0000-0001-5318-3910'
```
Sadly, to make requests to ORCID and Base APIs, you will need to register with them, I cannot share my identifiers
```
orcid_api_id = 'APP-XXXXXXXXXXXXXXXX'
orcid_api_secret = 'XXXXXXXX-XXXX-XXXX-XXXX-XXXXXXXXXXXX'
base_api_ua = 'XXXXXXXXXX@XXXXXXXXXXXXXXXXXXXXXX'
orcid_api_email = 'XXXXXXXXX@XXXXXXXXX'
```
Now we retrieve the papers for this author:
```
def findDOI(paper):
for i in paper['external-ids']['external-id']:
if i['external-id-type'] == 'doi':
return i['external-id-value']
return None
api = orcid.PublicAPI(orcid_api_id, orcid_api_secret)
token = api.get_search_token_from_orcid()
record = api.read_record_public(ORCID, 'record', token)
print(f'ORCID: {record["orcid-identifier"]["path"]}')
name = record['person']['name']
print(f'Name: {name["given-names"]["value"]} {name["family-name"]["value"]}')
works = api.read_record_public(ORCID, 'works', token)['group']
print(f'Number of papers in ORCID: {len(works)}\n')
dois = []
for paper in works:
doi = findDOI(paper)
if doi:
dois.append((doi, paper['work-summary'][0]['title']['title']['value']))
else:
title = paper['work-summary'][0]['title']['title']['value']
print(f'No DOI available for paper: {title}')
print(f'\nNumber of papers with DOI: {len(dois)}')
def base_search(doi):
r = requests.get('https://api.base-search.net/cgi-bin/BaseHttpSearchInterface.fcgi',
params={'func': 'PerformSearch', 'query': f'dcdoi:{doi}', 'boost': 'oa', 'format': 'json'},
headers={'user-agent': 'fx.coudert@chimieparistech.psl.eu'})
docs = r.json()['response']['docs']
for doc in docs:
if doc['dcoa'] == 1:
return True
return False
def unpaywall(doi):
r = requests.get(f'https://api.unpaywall.org/v2/{doi}',
params={"email": "fxcoudert@gmail.com"})
r = r.json()
if 'error' in r:
return False
if r['is_oa']:
return True
return False
res = []
print('Be patient, this step takes time (2 to 3 seconds per paper)')
for doi, title in dois:
res.append((doi, title, base_search(doi), unpaywall(doi)))
time.sleep(1)
def YesNoCell(b):
if b:
return '<td style="background-color: #60FF60">Yes</td>'
else:
return '<td style="background-color: #FF6060">No</td>'
s = ''
tot_base = 0
tot_unpaywall = 0
for doi, title, base, unpaywall in res:
s += f'<tr><td><a href="https://doi.org/{doi}" target="_blank">{title}</a></td>{YesNoCell(base)}{YesNoCell(unpaywall)}</td>'
if base:
tot_base += 1
if unpaywall:
tot_unpaywall += 1
s += f'<tr><td>Total</td><td>{100 * tot_base/ len(res):.1f}%</td><td>{100 * tot_unpaywall/ len(res):.1f}%</td></tr>'
header = '<tr><td>Paper title</td><td>OA in BASE</td><td>OA in Unpaywall</td></tr>'
display(HTML('<table>' + header + s + '</table>'))
```
| github_jupyter |
# Savanna Barcode Intelligence - Quickstart.
This notebook shows you how to get started with Zebra's Savanna APIs related to Barcode Intelligence
- UPC Lookup
- Barcode Creation
- FDA recall information for both food and drugs
## Setup
* In order to run the sample code in this guide you will need an API key, to obtain an API key follow the instructions detailed on the Zebra developer portal at **https://developer.zebra.com/getting-started-0**
* Once you have created a login to the developer portal and created an application you must ensure you have selected the **Barcode Intelligence** package and **associated this with your application**
To run a cell:
- Click anywhere in the code block.
- Click the play button in the top-left of the code block
```
# Paste your API key below, this might also be called the 'Consumer Key' on the portal
api_key = ''
print('Your API key is: ' + api_key)
```
## UPC Lookup
The UPC Lookup API provides product descriptions, photos and pricing for a wide range of products
```
# Enter a barcode to look up or uncomment one of the suggested values
barcode = ''
#barcode = '9780141032016' #Book
#barcode = '9780099558453' #Book
#barcode = '9780141039190' #Book
```
The UPC Lookup functionality is delivered as a REST API. Run the below code to return information about your specified barcode
```
import requests
import json
from requests.exceptions import HTTPError
barcode_lookup_url = 'https://api.zebra.com/v2/tools/barcode/lookup?upc=' + barcode
headers = {'apikey': api_key}
try:
response = requests.get(url = barcode_lookup_url, headers = headers, params = {})
response.raise_for_status()
except HTTPError as http_err:
print(f'HTTP error: {http_err}')
print(response.json())
except Exception as err:
print(f'Other error: {err}')
else:
print(json.dumps(response.json(), indent=4))
```
The following code will display the first returned image associated with the previously specified barcode
```
from IPython.display import Image
response_json = response.json()
if ('items' in response_json and 'images' in response_json['items'][0]):
print('Image: ' + response_json['items'][0]['images'][0])
photo_url = response_json['items'][0]['images'][0]
else:
print('Image not found')
photo_url = "https://www.zebra.com/gb/en/header/jcr:content/mainpar/header/navigationComponent/logoImage.adapt.full.png"
Image(photo_url, width=200)
```
## Create Barcode
The [Create Barcode](https://developer.zebra.com/tools-create-barcode/apis) API will generate a barcode image from the specified barcode data.
*Many symbologies are supported but the code snippet below will work with EAN 13 barcodes using the barcode value specified in the previous step*
```
import requests
from requests.exceptions import HTTPError
from IPython.display import Image
barcode_lookup_url = 'https://api.zebra.com/v2/tools/barcode/generate/?symbology=ean13&includeText=true&text=' + barcode
headers = {'apikey': api_key}
try:
response = requests.get(url = barcode_lookup_url, headers = headers, params = {})
response.raise_for_status()
except HTTPError as http_err:
print(f'HTTP error: {http_err}')
print(response.json())
except Exception as err:
print(f'Other error: {err}')
else:
generated_barcode_binary = response.content
Image(generated_barcode_binary)
print('ok - barcode generated')
Image(generated_barcode_binary)
```
## Food Recall
You can search for food recall announcements by UPC
```
# Specify the UPC to search for related FDA food recall announcements
food_recall_upc = ''
#food_recall_upc = '2324617054' # Shrimp Rolls
#food_recall_upc = '691035359586' # Raisens
import requests
from requests.exceptions import HTTPError
food_recall_url = 'https://api.zebra.com/v2/tools/recalls/food/upc?limit=1&val=' + food_recall_upc
headers = {'apikey': api_key}
try:
response = requests.get(url = food_recall_url, headers = headers, params = {})
response.raise_for_status()
except HTTPError as http_err:
print(f'HTTP error: {http_err}')
print(response.json())
except Exception as err:
print(f'Other error: {err}')
else:
print("Food Recall information is listed below:")
print(json.dumps(response.json(), indent=4))
```
## Drug Recall
You can search for medical recall information either by search term (for medical devices or drug recalls) or by specifying a UPC. This sample shows a search for drug recalls related to the specified search term. You can change how many results are returned by modifying the limit parameter in the URL.
```
# Specify a one word description to search for drug recalls about
drug_search_term = ''
#drug_search_term = 'head'
#drug_search_term = 'stomach'
import requests
from requests.exceptions import HTTPError
drug_recall_url = 'https://api.zebra.com/v2/tools/recalls/drug/description?limit=2&val=' + drug_search_term
headers = {'apikey': api_key}
try:
response = requests.get(url = drug_recall_url, headers = headers, params = {})
response.raise_for_status()
except HTTPError as http_err:
print(f'HTTP error: {http_err}')
print(response.json())
except Exception as err:
print(f'Other error: {err}')
else:
print("Drug Recall information is listed below:")
print(json.dumps(response.json(), indent=4))
```
## What's next?
Next, explore the [other APIs](https://developer.zebra.com/apis) available in Savanna or submit a new API suggestion to the [Sandbox](https://developer.zebra.com/sandbox-overview).
If you have questions then please visit the [Developer Portal](https://developer.zebra.com)
| github_jupyter |
# Homework 1: Wine Rating with Pandas and Sklearn
### 1. Read the syllabus in its entirety. Mark “Yes” below.
_____ I have read and understood the syllabus for this class.
### 2. MTC (MegaTelCo) has decided to use supervised learning to address its problem of churn in its wireless phone business. As a consultant to MTC, you realize that a main task in the business understanding/data understanding phases of the data mining process is to define the target variable. In one or two sentences each, please suggest a definition for the target variable. Suggest 3 features and explain why you think they will be give information on the target variable. Be as precise as possible—someone else will be implementing your suggestion. (Remember: your formulation should make sense from a business point of view, and it should be reasonable that MTC would have data available to know the value of the target variable and features for historical customers.)
___
For the remainder of this homework, we explore the chemical characteristics of wine and how those characteristics relate to ratings given to wine by expert wine tasters. The data is described at a high level [here](https://archive.ics.uci.edu/ml/datasets/Wine+Quality). For simplicity, for this assignment, we'll restrict our analyses to white wines.
Tasks for this assignment will extend what we've done in the first lecture, and include data manipulation, analysis, visualation, and modeling.
```
# consider the following url referencing white wines
wine_url = "https://archive.ics.uci.edu/ml/machine-learning-databases/wine-quality/winequality-white.csv"
```
### **3.** Load the data referenced by the above url into a pandas data frame. What are the names of the columns that are present? Hint: columns of this data are delimited by a ;
### 4. What is the distribution of different quality ratings? Print the number of occurrences for each quality rating in the data frame you've created
### 5. Plot the above distribution as a histogram inline in this notebook
### 6. Notice the above distribution is highly imbalanced, some ratings occur much more frequently than others, making it difficult to fully compare ratings to each other. One common way to deal with this kind of imbalance in data visualization is by plotting a histogram of the _log_ of counts. Plot another histogram so that the data is presented on a log scale
### 7. Show descriptive statistics for all the columns "pH", "density", "chlorides" and "fixed acidity".
### 8. Measures of correlation, such as Pearson's correlation coefficient, show whether one numeric variable gives information on another numeric variable. Pandas allows us to compute the Pearson correlation coefficient between all pairs of columns in our dataframe. Display the correlations between all pairs of columns, for those rows with a quality score of at least 5.
### 9. Is this the only sort of correlation that Pandas lets us compute? If so, repeat exercise 6 with the an alternate correlation. If not, name another and explain the difference.
### 10. Heatmaps are a tool for conveniently interpreting correlation data. Plot these correlations as a seaborn heatmap. Which pairs of variables are most closely correlated? Which variable gives the most information on wine quality?
### 11. Following the example in class, build a linear model to predict the quality of wine using the chemical info available. Generate predictions and compare predicted quality to the actual value in a scatter plot
### 12. There are many different types of predictive models, each with their own plusses and minuses. For this task, repeat your modeling performed in step 8, but using a `sklearn.ensemble.RandomForestRegressor`. How does the scatter plot compare with the prior results?
| github_jupyter |
<a href="https://colab.research.google.com/github/skhadem/numerical-analysis-class/blob/master/Homework/Solutions/HW10/hw10.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
import numpy as np
import matplotlib.pyplot as plt
from scipy.interpolate import CubicHermiteSpline
# y'
f = lambda t : np.cos(t)
# True sol
F = lambda t: np.sin(t)
```
# Problem 2d
```
def eq_4(f, F, tspan, y0, h=0.1, dense_output=False):
"""F should be the true solution"""
n = int( (tspan[1] - tspan[0])/h ) + 1
t, h = np.linspace(tspan[0],tspan[1], num=n, retstep=True)
w = np.zeros(n)
dydt = np.zeros(n)
w[0] = y0
w[1] = F(t[1]) # init using the true sol
dydt[0] = f(t[0])
for i in range(2, n):
dydt[i-1] = f(t[i-1])
w[i] = 4*w[i-1] - 3*w[i-2] - 2*h*dydt[i-2]
if dense_output is True:
sol = CubicHermiteSpline(t,w,dydt)
return t, w, sol
else:
return t, w
t_hist, w_hist, sol = eq_4(f, F, [0, 0.1], 0, h=1e-2, dense_output=True)
# Make sure the last value is 0.1097
w_hist[-1]
def eq_5(f, F, tspan, y0, h=0.1, dense_output=False):
"""F should be the true solution"""
n = int( (tspan[1] - tspan[0])/h ) + 1
t, h = np.linspace(tspan[0],tspan[1], num=n, retstep=True)
w = np.zeros(n)
dydt = np.zeros(n)
w[0] = y0
w[1] = F(t[1])
dydt[0] = f(t[0])
dydt[1] = f(t[1])
for i in range(2, n):
dydt[i] = f(t[i])
w[i] = (4*w[i-1] - w[i-2] + 2*h*dydt[i])/3
if dense_output is True:
sol = CubicHermiteSpline(t,w,dydt)
return t, w, sol
else:
return t, w
t_hist, w_hist, sol = eq_5(f, F, [0, 0.1], 0, h=1e-2, dense_output=True)
# Make sure the last value is 0.0998
w_hist[-1]
```
# Problem 2e
## Eq 4
```
hs = np.logspace(-2, -6, 10)
errors = []
true_sol = F(0.1)
for h in hs:
_, w_hist = eq_4(f, F, [0, 0.1], 0, h=h, dense_output=False)
errors.append(np.abs(w_hist[-1] - true_sol))
plt.loglog(hs, errors)
plt.loglog(hs, hs,'--',label='$O(h)$')
plt.loglog(hs, hs**2,'--',label='$O(h^2)$')
plt.grid()
plt.legend()
plt.ylabel('Error $|y(0.1) - w_n|$')
plt.gca().invert_xaxis()
plt.xlabel('$h \propto n^{-1}$')
plt.show()
```
## Eq 5
```
hs = np.logspace(-2, -6, 10)
errors = []
true_sol = F(0.1)
for h in hs:
_, w_hist = eq_5(f, F, [0, 0.1], 0, h=h, dense_output=False)
errors.append(np.abs(w_hist[-1] - true_sol))
plt.loglog(hs, errors, label='BD2')
plt.loglog(hs, hs,'--',label='$O(h)$')
plt.loglog(hs, hs**2,'--',label='$O(h^2)$')
plt.loglog(hs, hs**3,'--',label='$O(h^3)$')
plt.grid()
plt.legend()
plt.ylabel('Error $|y(0.1) - w_n|$')
plt.gca().invert_xaxis()
plt.xlabel('$h \propto n^{-1}$')
plt.show()
errors
hs
```
An error of $10^{-10}$ occurs after h gets smaller than $10^{-4}$
# Problem 1f
As expected, the first method (eq 4) is not stable. This leads to the error blowing up. The second (implicit) method is stable, so it converges at approximately $\mathcal{O}(h^2.5)$
```
```
| github_jupyter |
# Training a model with distributed LightGBM
In this example we will train a model in Ray AIR using distributed LightGBM.
Let's start with installing our dependencies:
```
!pip install -qU "ray[tune]" lightgbm_ray
```
Then we need some imports:
```
import argparse
import math
from typing import Tuple
import pandas as pd
import ray
from ray.train.batch_predictor import BatchPredictor
from ray.train.lightgbm import LightGBMPredictor
from ray.data.preprocessors.chain import Chain
from ray.data.preprocessors.encoder import Categorizer
from ray.train.lightgbm import LightGBMTrainer
from ray.data.dataset import Dataset
from ray.air.result import Result
from ray.data.preprocessors import StandardScaler
from sklearn.datasets import load_breast_cancer
from sklearn.model_selection import train_test_split
```
Next we define a function to load our train, validation, and test datasets.
```
def prepare_data() -> Tuple[Dataset, Dataset, Dataset]:
data_raw = load_breast_cancer()
dataset_df = pd.DataFrame(data_raw["data"], columns=data_raw["feature_names"])
dataset_df["target"] = data_raw["target"]
# add a random categorical column
num_samples = len(dataset_df)
dataset_df["categorical_column"] = pd.Series(
(["A", "B"] * math.ceil(num_samples / 2))[:num_samples]
)
train_df, test_df = train_test_split(dataset_df, test_size=0.3)
train_dataset = ray.data.from_pandas(train_df)
valid_dataset = ray.data.from_pandas(test_df)
test_dataset = ray.data.from_pandas(test_df.drop("target", axis=1))
return train_dataset, valid_dataset, test_dataset
```
The following function will create a LightGBM trainer, train it, and return the result.
```
def train_lightgbm(num_workers: int, use_gpu: bool = False) -> Result:
train_dataset, valid_dataset, _ = prepare_data()
# Scale some random columns, and categorify the categorical_column,
# allowing LightGBM to use its built-in categorical feature support
columns_to_scale = ["mean radius", "mean texture"]
preprocessor = Chain(
Categorizer(["categorical_column"]), StandardScaler(columns=columns_to_scale)
)
# LightGBM specific params
params = {
"objective": "binary",
"metric": ["binary_logloss", "binary_error"],
}
trainer = LightGBMTrainer(
scaling_config={
"num_workers": num_workers,
"use_gpu": use_gpu,
},
label_column="target",
params=params,
datasets={"train": train_dataset, "valid": valid_dataset},
preprocessor=preprocessor,
num_boost_round=100,
)
result = trainer.fit()
print(result.metrics)
return result
```
Once we have the result, we can do batch inference on the obtained model. Let's define a utility function for this.
```
def predict_lightgbm(result: Result):
_, _, test_dataset = prepare_data()
batch_predictor = BatchPredictor.from_checkpoint(
result.checkpoint, LightGBMPredictor
)
predicted_labels = (
batch_predictor.predict(test_dataset)
.map_batches(lambda df: (df > 0.5).astype(int), batch_format="pandas")
.to_pandas(limit=float("inf"))
)
print(f"PREDICTED LABELS\n{predicted_labels}")
shap_values = batch_predictor.predict(test_dataset, pred_contrib=True).to_pandas(
limit=float("inf")
)
print(f"SHAP VALUES\n{shap_values}")
```
Now we can run the training:
```
result = train_lightgbm(num_workers=2, use_gpu=False)
```
And perform inference on the obtained model:
```
predict_lightgbm(result)
```
| github_jupyter |
# Exploratory Data Analysis - Zillow Zestimate Competition
```
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load in
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import matplotlib as plt
import seaborn as sns
%matplotlib inline
# Input data files are available in the "../input/" directory.
# For example, running this (by clicking run or pressing Shift+Enter) will list the files in the input directory
from subprocess import check_output
print(check_output(["ls", "data/"]).decode("utf8"))
# Any results you write to the current directory are saved as output.
```
### Look at the Data Dictionary
```
def print_dictionary_tables(input_path):
try:
excel_file = pd.ExcelFile(input_path)
dfs = []
for sheet in excel_file.sheet_names:
print(sheet)
df = excel_file.parse(sheet)
if df.shape[1] == 2:
for i in range(df.shape[0]):
print(df[df.columns[0]][i], ' - ' + df[df.columns[1]][i])
df['Table'] = sheet
dfs.append(df)
else:
print("Too many dimensions to print.")
print(' ')
print(' ')
return dfs
except:
"Dimensions incorrect."
dict_dfs = print_dictionary_tables('data/zillow_data_dictionary.xlsx')
dict_dfs[4]
```
#### End of data dictionary exploration.
## Explore Training Set File
```
train = pd.read_csv('data/train_2016_v2.csv')
train.head()
train.shape
train.parcelid.value_counts().head()
# Are the 3 rows different?
train[train['parcelid'] == 11842707]
```
So, some are duplicated, but only one happened 3 times.
The parcel with 3 appearances had 3 different transaction dates, and went up with each date. That'll be something to look at in the parcels set.
```
train.logerror.describe()
train.transactiondate.value_counts().head()
```
Lots of duplication in dates. A couple only have one entry.
#### End of training set exploration.
## Explore Properties File
```
properties = pd.read_csv('data/properties_2016.csv', low_memory=False)
properties.head()
properties.shape
```
#### For categoricals, I will fill NaNs with a -1, for 'no category'.
```
ac = dict_dfs[4].copy()
properties = properties.merge(ac, how='left', left_on='airconditioningtypeid', right_on='AirConditioningTypeID')
properties.drop(['AirConditioningTypeID', 'Table'], axis=1, inplace=True)
properties.head()
properties['airconditioningtypeid'].fillna(value=-1, inplace=True)
properties['AirConditioningDesc'].fillna(value='Not Specified', inplace=True)
properties['AirConditioningDesc'].value_counts()
properties.AirConditioningDesc.value_counts()[1:].plot(kind='bar')
arch = dict_dfs[5].copy()
arch.head()
properties = properties.merge(arch, how='left', left_on='architecturalstyletypeid', right_on='ArchitecturalStyleTypeID')
properties.drop(['ArchitecturalStyleTypeID', 'Table'], axis=1, inplace=True)
properties['architecturalstyletypeid'].fillna(value=-1, inplace=True)
properties['ArchitecturalStyleDesc'].fillna(value='Not Specified', inplace=True)
properties.head()
properties['ArchitecturalStyleDesc'].value_counts()
properties['ArchitecturalStyleDesc'].value_counts()[1:].plot(kind='bar')
properties['basementsqft'].hist()
properties['basementsqft'].describe()
# Since there are no zeros, and this won't be a negative value, I'll fill na's with 0
properties['basementsqft'].fillna(value=0, inplace=True)
properties['basementsqft'].describe()
properties['basementsqft'].hist()
properties.shape[0] - 1628
# It looks like a vast majority don't have finished basements!
properties.head()
properties['bathroomcnt'].count()
properties.shape[0]
properties.shape[0] - properties['bathroomcnt'].count()
properties.shape[0] - properties['bedroomcnt'].count()
bc = dict_dfs[7].copy()
properties = properties.merge(bc, how='left', left_on='buildingclasstypeid', right_on='BuildingClassTypeID')
properties.drop(['BuildingClassTypeID', 'Table'], axis=1, inplace=True)
properties.head()
properties['buildingclasstypeid'].fillna(value=-1, inplace=True)
properties['BuildingClassDesc'].fillna(value='Not Specified', inplace=True)
properties['BuildingClassDesc'].value_counts()
properties['BuildingClassDesc'].value_counts()[1:].plot(kind='bar')
properties.head()
properties['buildingqualitytypeid'].value_counts()
properties['buildingqualitytypeid'].hist()
properties['calculatedbathnbr'].hist()
# Percentage which are NaN
(properties.shape[0] - properties['calculatedbathnbr'].count())/properties.shape[0]
# There is only a single value. It's pretty well, has a deck or doesn't have a deck. Going to fill N/A with negative one and treat this as categorical
properties['decktypeid'].value_counts()
properties['decktypeid'].fillna(-1, inplace=True)
properties.head()
properties['finishedfloor1squarefeet'].hist()
properties['finishedfloor1squarefeet'].count()
# Percentage not having a value. Derive what can be derived from similar values, and do a cluster median for the rest.
(properties.shape[0] - properties['finishedfloor1squarefeet'].count())/properties.shape[0]
properties['calculatedfinishedsquarefeet'].hist()
properties['calculatedfinishedsquarefeet'].count()
(properties.shape[0] - properties['calculatedfinishedsquarefeet'].count())/properties.shape[0]
properties['finishedsquarefeet12'].hist()
properties['finishedsquarefeet12'].count()
(properties.shape[0] - properties['finishedsquarefeet12'].count())/properties.shape[0]
properties['finishedsquarefeet13'].hist()
properties['finishedsquarefeet13'].count()
(properties.shape[0] - properties['finishedsquarefeet13'].count())/properties.shape[0]
properties['finishedsquarefeet15'].hist()
properties['finishedsquarefeet15'].count()
(properties.shape[0] - properties['finishedsquarefeet15'].count())/properties.shape[0]
properties['finishedsquarefeet50'].hist()
properties['finishedsquarefeet50'].count()
(properties.shape[0] - properties['finishedsquarefeet50'].count())/properties.shape[0]
properties['finishedsquarefeet6'].hist()
properties['finishedsquarefeet6'].count()
(properties.shape[0] - properties['finishedsquarefeet6'].count())/properties.shape[0]
properties['fips'].value_counts().plot(kind='bar')
(properties.shape[0] - properties['fips'].count())/properties.shape[0]
print(properties.shape[0])
print(properties['fips'].count())
properties.shape[0] - properties['fips'].count()
properties['fireplacecnt'].value_counts().plot(kind='bar')
properties['fireplacecnt'].value_counts()
# No zero, so I CAN do a fillna with zero on this one.
properties['fireplacecnt'].fillna(value=0, inplace=True)
properties['fullbathcnt'].value_counts().plot(kind='bar')
properties['fullbathcnt'].value_counts()
# Again, no zero, so I can fillna with zero.
properties['fullbathcnt'].fillna(0, inplace=True)
properties['garagecarcnt'].value_counts().plot(kind='bar')
properties['garagecarcnt'].value_counts()
properties.shape[0] - properties['garagecarcnt'].count()
properties['garagetotalsqft'].value_counts().plot(kind='bar')
properties['garagetotalsqft'].value_counts()
properties.shape[0] - properties['garagetotalsqft'].count()
properties.shapepe[0]
properties.columns
properties['taxvaluedollarcnt'].describe()
properties.columns
# So, do all records have a target variable?
properties.shape[0] - properties['landtaxvaluedollarcnt'].count()
# TODO: See if any of these have a value at another point in time. Use that to fill n/a if available, or else take a median based on other characteristics.
(properties.shape[0] - properties['landtaxvaluedollarcnt'].count())/properties.shape[0]
# Constructing of list of variables to fill later. Need to be zero, but REAL zeroes are present.
no_zero_na = ['bathroomcnt','bedroomcnt']
fill_with_median_or_mode_from_cluster = ['buildingqualitytypeid','fips', 'garagecarcnt']
fill_w_cluster_median_or_derive = ['finishedfloor1squarefeet', 'calculatedfinishedsquarefeet', 'finishedsquarefeet12',
'finishedsquarefeet13', 'finishedsquarefeet15', 'finishedsquarefeet50',
'finishedsquarefeet6']
```
#### End of properties file exploration.
| github_jupyter |
```
import SimpleITK as sitk
import numpy as np
import os
def normalize_one_volume(volume):
new_volume = np.zeros(volume.shape)
location = np.where(volume != 0)
mean = np.mean(volume[location])
var = np.std(volume[location])
new_volume[location] = (volume[location] - mean) / var
return new_volume
def merge_volumes(*volumes):
return np.stack(volumes, axis=0)
def get_volume(root, patient, desired_depth,
desired_height, desired_width,
normalize_flag,
flip=0):
flair_suffix = "_flair.nii.gz"
t1_suffix = "_t1.nii.gz"
t1ce_suffix = "_t1ce.nii.gz"
t2_suffix = "_t2.nii.gz"
path_flair = os.path.join(root, patient, patient + flair_suffix)
path_t1 = os.path.join(root, patient, patient + t1_suffix)
path_t2 = os.path.join(root, patient, patient + t2_suffix)
path_t1ce = os.path.join(root, patient, patient + t1ce_suffix)
flair = sitk.GetArrayFromImage(sitk.ReadImage(path_flair))
t1 = sitk.GetArrayFromImage(sitk.ReadImage(path_t1))
t2 = sitk.GetArrayFromImage(sitk.ReadImage(path_t2))
t1ce = sitk.GetArrayFromImage(sitk.ReadImage(path_t1ce))
if desired_depth > 155:
flair = np.concatenate([flair, np.zeros((desired_depth - 155, 240, 240))], axis=0)
t1 = np.concatenate([t1, np.zeros((desired_depth - 155, 240, 240))], axis=0)
t2 = np.concatenate([t2, np.zeros((desired_depth - 155, 240, 240))], axis=0)
t1ce = np.concatenate([t1ce, np.zeros((desired_depth - 155, 240, 240))], axis=0)
if normalize_flag == True:
out = merge_volumes(normalize_one_volume(flair), normalize_one_volume(t2), normalize_one_volume(t1ce),
normalize_one_volume(t1))
else:
out = merge_volumes(flair, t2, t1ce, t1)
if flip == 1:
out = out[:, ::-1, :, :]
elif flip == 2:
out = out[:, :, ::-1, :]
elif flip == 3:
out = out[:, :, :, ::-1]
elif flip == 4:
out = out[:, :, ::-1, ::-1]
elif flip == 5:
out = out[:, ::-1, ::-1, ::-1]
return np.expand_dims(out, axis=0)
import sys
import numpy as np
sys.path.remove('/home/sentic/.local/lib/python3.6/site-packages')
import torch
torch.backends.cudnn.benchmark=True
device_id = 0
torch.cuda.set_device(device_id)
root = "/home/sentic/MICCAI/data/train/"
use_gpu = True
n_epochs = 300
batch_size = 1
use_amp = False
import torch.nn as nn
import torch.nn.functional as F
from torch.autograd import Variable
import torch.optim as optim
import torch.optim.lr_scheduler as lr_scheduler
from torch.optim.lr_scheduler import LambdaLR
from torch.utils.data import DataLoader
from model import LargeCascadedModel
from dataset import BraTS
from losses import DiceLoss, DiceLossLoss
from tqdm import tqdm_notebook, tqdm
paths_model = ["./checkpoints/checkpoint_190.pt", "./checkpoints/checkpoint_191.pt", "./checkpoints/checkpoint_192.pt",
"./checkpoints/checkpoint_193.pt", "./checkpoints/checkpoint_194.pt", "./checkpoints/checkpoint_195.pt",
"./checkpoints/checkpoint_196.pt", "./checkpoints/checkpoint_197.pt", "./checkpoints/checkpoint_198.pt",
"./checkpoints/checkpoint_199.pt"]
diceLoss = DiceLossLoss()
intresting_patients = ['BraTS20_Validation_067', 'BraTS20_Validation_068', 'BraTS20_Validation_069', 'BraTS20_Validation_072',
'BraTS20_Validation_083', 'BraTS20_Validation_077', 'BraTS20_Validation_076', 'BraTS20_Validation_074',
'BraTS20_Validation_085', 'BraTS20_Validation_087', 'BraTS20_Validation_088', 'BraTS20_Validation_089',
'BraTS20_Validation_091', 'BraTS20_Validation_092', 'BraTS20_Validation_099', 'BraTS20_Validation_103']
patients_path = "/home/sentic/MICCAI/data/train/"
for patient_name in tqdm(os.listdir(patients_path)):
if patient_name.startswith('BraTS'):
output = np.zeros((3, 155, 240, 240))
for path_model in paths_model:
model = LargeCascadedModel(inplanes_encoder_1=4, channels_encoder_1=16, num_classes_1=3,
inplanes_encoder_2=7, channels_encoder_2=32, num_classes_2=3)
model.load_state_dict(torch.load(path_model, map_location='cuda:0')['state_dict'])
if use_gpu:
model = model.to("cuda")
model.eval()
with torch.no_grad():
for flip in range(0, 6):
volume = get_volume(patients_path, patient_name, 160, 240, 240, True, flip)
volume = torch.FloatTensor(volume.copy())
if use_gpu:
volume = volume.to("cuda")
_, _, decoded_region3, _ = model(volume)
decoded_region3 = decoded_region3.detach().cpu().numpy()
decoded_region3 = decoded_region3.squeeze()
if flip == 1:
decoded_region3 = decoded_region3[:, ::-1, :, :]
elif flip == 2:
decoded_region3 = decoded_region3[:, :, ::-1, :]
elif flip == 3:
decoded_region3 = decoded_region3[:, :, :, ::-1]
elif flip == 4:
decoded_region3 = decoded_region3[:, :, ::-1, ::-1]
elif flip == 5:
decoded_region3 = decoded_region3[:, ::-1, ::-1, ::-1]
output += decoded_region3[:, :155, :, :]
np_array = output
np_array = np_array / (6.0 * 10.0)
np.save("./val_masks_np/" + patient_name + ".np", np_array)
import os
import numpy as np
import SimpleITK as sitk
threshold_wt = 0.7
threshold_tc = 0.6
threshold_et = 0.7
low_threshold_et = 0.6
threshold_num_pixels_et = 150
patients_path = "/home/sentic/MICCAI/data/train/"
intresting_patients = ['BraTS20_Validation_067', 'BraTS20_Validation_068', 'BraTS20_Validation_069', 'BraTS20_Validation_072',
'BraTS20_Validation_083', 'BraTS20_Validation_077', 'BraTS20_Validation_076', 'BraTS20_Validation_074',
'BraTS20_Validation_085', 'BraTS20_Validation_087', 'BraTS20_Validation_088', 'BraTS20_Validation_089',
'BraTS20_Validation_091', 'BraTS20_Validation_092', 'BraTS20_Validation_099', 'BraTS20_Validation_103']
for patient_name in os.listdir(patients_path):
if patient_name.startswith('BraTS'):
path_big_volume = os.path.join(patients_path, patient_name, patient_name + "_flair.nii.gz")
np_array = np.load("./val_masks_np/" + patient_name + ".np.npy")
image = sitk.ReadImage(path_big_volume)
direction = image.GetDirection()
spacing = image.GetSpacing()
origin = image.GetOrigin()
seg_image = np.zeros((155, 240, 240))
label_1 = np_array[2, :, :, :] # where the enhanced tumor is
location_pixels_et = np.where(label_1 >= threshold_et)
num_pixels_et = location_pixels_et[0].shape[0]
label_2 = np_array[1, :, :, :] # locatia 1-urilor si 4-urilor
label_3 = np_array[0, :, :, :] # locatia 1 + 2 + 4
if patient_name in intresting_patients:
print(patient_name, "--->", num_pixels_et)
else:
print(patient_name, "***->", num_pixels_et)
if num_pixels_et > threshold_num_pixels_et: # if there are at least num of pixels
label_1[label_1 >= threshold_et] = 1 # put them in et category
else:
label_1[label_1 >= threshold_et] = 0 # don't put them
label_2[location_pixels_et] = 1 # but put them on tumor core
label_2[(label_1 < threshold_et) & (label_1 >= low_threshold_et)] = 1
label_1[label_1 < threshold_et] = 0
location_1 = np.where(label_1 != 0)
seg_image[location_1] = 4
label_2[label_2 >= threshold_tc] = 1
label_2[label_2 < threshold_tc] = 0
location_2 = np.where((label_2 != 0) & (label_1 == 0))
seg_image[location_2] = 1
label_3[label_3 >= threshold_wt] = 1
label_3[label_3 < threshold_wt] = 0
location_3 = np.where((label_3 != 0) & (label_2 == 0))
seg_image[location_3] = 2
out_image = sitk.GetImageFromArray(seg_image)
out_image.SetDirection(direction)
out_image.SetSpacing(spacing)
out_image.SetOrigin(origin)
sitk.WriteImage(out_image, os.path.join("./final_masks", patient_name + ".nii.gz"))
```
| github_jupyter |
# MadMiner particle physics tutorial
# Part 3a: Training a likelihood ratio estimator
Johann Brehmer, Felix Kling, Irina Espejo, and Kyle Cranmer 2018-2019
In part 3a of this tutorial we will finally train a neural network to estimate likelihood ratios. We assume that you have run part 1 and 2a of this tutorial. If, instead of 2a, you have run part 2b, you just have to load a different filename later.
## Preparations
```
from __future__ import absolute_import, division, print_function, unicode_literals
import logging
import numpy as np
import matplotlib
from matplotlib import pyplot as plt
%matplotlib inline
from madminer.sampling import SampleAugmenter
from madminer import sampling
from madminer.ml import ParameterizedRatioEstimator
# MadMiner output
logging.basicConfig(
format='%(asctime)-5.5s %(name)-20.20s %(levelname)-7.7s %(message)s',
datefmt='%H:%M',
level=logging.INFO
)
# Output of all other modules (e.g. matplotlib)
for key in logging.Logger.manager.loggerDict:
if "madminer" not in key:
logging.getLogger(key).setLevel(logging.WARNING)
```
## 1. Make (unweighted) training and test samples with augmented data
At this point, we have all the information we need from the simulations. But the data is not quite ready to be used for machine learning. The `madminer.sampling` class `SampleAugmenter` will take care of the remaining book-keeping steps before we can train our estimators:
First, it unweights the samples, i.e. for a given parameter vector `theta` (or a distribution `p(theta)`) it picks events `x` such that their distribution follows `p(x|theta)`. The selected samples will all come from the event file we have so far, but their frequency is changed -- some events will appear multiple times, some will disappear.
Second, `SampleAugmenter` calculates all the augmented data ("gold") that is the key to our new inference methods. Depending on the specific technique, these are the joint likelihood ratio and / or the joint score. It saves all these pieces of information for the selected events in a set of numpy files that can easily be used in any machine learning framework.
```
sampler = SampleAugmenter('data/lhe_data_shuffled.h5')
# sampler = SampleAugmenter('data/delphes_data_shuffled.h5')
```
The `SampleAugmenter` class defines five different high-level functions to generate train or test samples:
- `sample_train_plain()`, which only saves observations x, for instance for histograms or ABC;
- `sample_train_local()` for methods like SALLY and SALLINO, which will be demonstrated in the second part of the tutorial;
- `sample_train_density()` for neural density estimation techniques like MAF or SCANDAL;
- `sample_train_ratio()` for techniques like CARL, ROLR, CASCAL, and RASCAL, when only theta0 is parameterized;
- `sample_train_more_ratios()` for the same techniques, but with both theta0 and theta1 parameterized;
- `sample_test()` for the evaluation of any method.
For the arguments `theta`, `theta0`, or `theta1`, you can (and should!) use the helper functions `benchmark()`, `benchmarks()`, `morphing_point()`, `morphing_points()`, and `random_morphing_points()`, all defined in the `madminer.sampling` module.
Here we'll train a likelihood ratio estimator with the ALICES method, so we focus on the `extract_samples_train_ratio()` function. We'll sample the numerator hypothesis in the likelihood ratio with 1000 points drawn from a Gaussian prior, and fix the denominator hypothesis to the SM.
Note the keyword `sample_only_from_closest_benchmark=True`, which makes sure that for each parameter point we only use the events that were originally (in MG) generated from the closest benchmark. This reduces the statistical fluctuations in the outcome quite a bit.
```
x, theta0, theta1, y, r_xz, t_xz, n_effective = sampler.sample_train_ratio(
theta0=sampling.random_morphing_points(10000, [('gaussian', 0., 0.5), ('gaussian', 0., 0.5)]),
theta1=sampling.benchmark('sm'),
n_samples=500000,
folder='./data/samples',
filename='train_ratio',
sample_only_from_closest_benchmark=True,
return_individual_n_effective=True,
)
```
For the evaluation we'll need a test sample:
```
_ = sampler.sample_test(
theta=sampling.benchmark('sm'),
n_samples=1000,
folder='./data/samples',
filename='test'
)
_,_,neff=sampler.sample_train_plain(
theta=sampling.morphing_point([0,0.5]),
n_samples=10000,
)
```
You might notice the information about the "eeffective number of samples" in the output. This is defined as `1 / max_events(weights)`; the smaller it is, the bigger the statistical fluctuations from too large weights. Let's plot this over the parameter space:
```
cmin, cmax = 10., 10000.
cut = (y.flatten()==0)
fig = plt.figure(figsize=(5,4))
sc = plt.scatter(theta0[cut][:,0], theta0[cut][:,1], c=n_effective[cut],
s=30., cmap='viridis',
norm=matplotlib.colors.LogNorm(vmin=cmin, vmax=cmax),
marker='o')
cb = plt.colorbar(sc)
cb.set_label('Effective number of samples')
plt.xlim(-1.0,1.0)
plt.ylim(-1.0,1.0)
plt.tight_layout()
plt.show()
```
## 2. Plot cross section over parameter space
This is not strictly necessary, but we can also plot the cross section as a function of parameter space:
```
thetas_benchmarks, xsecs_benchmarks, xsec_errors_benchmarks = sampler.cross_sections(
theta=sampling.benchmarks(list(sampler.benchmarks.keys()))
)
thetas_morphing, xsecs_morphing, xsec_errors_morphing = sampler.cross_sections(
theta=sampling.random_morphing_points(1000, [('gaussian', 0., 1.), ('gaussian', 0., 1.)])
)
cmin, cmax = 0., 2.5 * np.mean(xsecs_morphing)
fig = plt.figure(figsize=(5,4))
sc = plt.scatter(thetas_morphing[:,0], thetas_morphing[:,1], c=xsecs_morphing,
s=40., cmap='viridis', vmin=cmin, vmax=cmax,
marker='o')
plt.scatter(thetas_benchmarks[:,0], thetas_benchmarks[:,1], c=xsecs_benchmarks,
s=200., cmap='viridis', vmin=cmin, vmax=cmax, lw=2., edgecolor='black',
marker='s')
cb = plt.colorbar(sc)
cb.set_label('xsec [pb]')
plt.xlim(-3.,3.)
plt.ylim(-3.,3.)
plt.tight_layout()
plt.show()
```
What you see here is a morphing algorithm in action. We only asked MadGraph to calculate event weights (differential cross sections, or basically squared matrix elements) at six fixed parameter points (shown here as squares with black edges). But with our knowledge about the structure of the process we can interpolate any observable to any parameter point without loss (except that statistical uncertainties might increase)!
## 3. Train likelihood ratio estimator
It's now time to build the neural network that estimates the likelihood ratio. The central object for this is the `madminer.ml.ParameterizedRatioEstimator` class. It defines functions that train, save, load, and evaluate the estimators.
In the initialization, the keywords `n_hidden` and `activation` define the architecture of the (fully connected) neural network:
```
estimator = ParameterizedRatioEstimator(
n_hidden=(60,60),
activation="tanh"
)
```
To train this model we will minimize the ALICES loss function described in ["Likelihood-free inference with an improved cross-entropy estimator"](https://arxiv.org/abs/1808.00973). Many alternatives, including RASCAL, are described in ["Constraining Effective Field Theories With Machine Learning"](https://arxiv.org/abs/1805.00013) and ["A Guide to Constraining Effective Field Theories With Machine Learning"](https://arxiv.org/abs/1805.00020). There is also SCANDAL introduced in ["Mining gold from implicit models to improve likelihood-free inference"](https://arxiv.org/abs/1805.12244).
```
estimator.train(
method='alices',
theta='data/samples/theta0_train_ratio.npy',
x='data/samples/x_train_ratio.npy',
y='data/samples/y_train_ratio.npy',
r_xz='data/samples/r_xz_train_ratio.npy',
t_xz='data/samples/t_xz_train_ratio.npy',
alpha=10,
n_epochs=10,
scale_parameters=True,
)
estimator.save('models/alices')
```
Let's for fun also train a model that only used `pt_j1` as input observable, which can be specified using the option `features` when defining the `ParameterizedRatioEstimator`
```
estimator_pt = ParameterizedRatioEstimator(
n_hidden=(40,40),
activation="tanh",
features=[0],
)
estimator_pt.train(
method='alices',
theta='data/samples/theta0_train_ratio.npy',
x='data/samples/x_train_ratio.npy',
y='data/samples/y_train_ratio.npy',
r_xz='data/samples/r_xz_train_ratio.npy',
t_xz='data/samples/t_xz_train_ratio.npy',
alpha=8,
n_epochs=10,
scale_parameters=True,
)
estimator_pt.save('models/alices_pt')
```
## 4. Evaluate likelihood ratio estimator
`estimator.evaluate_log_likelihood_ratio(theta,x)` estimated the log likelihood ratio and the score for all combination between the given phase-space points `x` and parameters `theta`. That is, if given 100 events `x` and a grid of 25 `theta` points, it will return 25\*100 estimates for the log likelihood ratio and 25\*100 estimates for the score, both indexed by `[i_theta,i_x]`.
```
theta_each = np.linspace(-1,1,25)
theta0, theta1 = np.meshgrid(theta_each, theta_each)
theta_grid = np.vstack((theta0.flatten(), theta1.flatten())).T
np.save('data/samples/theta_grid.npy', theta_grid)
theta_denom = np.array([[0.,0.]])
np.save('data/samples/theta_ref.npy', theta_denom)
estimator.load('models/alices')
log_r_hat, _ = estimator.evaluate_log_likelihood_ratio(
theta='data/samples/theta_grid.npy',
x='data/samples/x_test.npy',
evaluate_score=False
)
```
Let's look at the result:
```
bin_size = theta_each[1] - theta_each[0]
edges = np.linspace(theta_each[0] - bin_size/2, theta_each[-1] + bin_size/2, len(theta_each)+1)
fig = plt.figure(figsize=(6,5))
ax = plt.gca()
expected_llr = np.mean(log_r_hat,axis=1)
best_fit = theta_grid[np.argmin(-2.*expected_llr)]
cmin, cmax = np.min(-2*expected_llr), np.max(-2*expected_llr)
pcm = ax.pcolormesh(edges, edges, -2. * expected_llr.reshape((25,25)),
norm=matplotlib.colors.Normalize(vmin=cmin, vmax=cmax),
cmap='viridis_r')
cbar = fig.colorbar(pcm, ax=ax, extend='both')
plt.scatter(best_fit[0], best_fit[1], s=80., color='black', marker='*')
plt.xlabel(r'$\theta_0$')
plt.ylabel(r'$\theta_1$')
cbar.set_label(r'$\mathbb{E}_x [ -2\, \log \,\hat{r}(x | \theta, \theta_{SM}) ]$')
plt.tight_layout()
plt.show()
```
Note that in this tutorial our sample size was very small, and the network might not really have a chance to converge to the correct likelihood ratio function. So don't worry if you find a minimum that is not at the right point (the SM, i.e. the origin in this plot). Feel free to dial up the event numbers in the run card as well as the training samples and see what happens then!
| github_jupyter |
<h1> Getting started with TensorFlow </h1>
In this notebook, you play around with the TensorFlow Python API.
```
import tensorflow as tf
import numpy as np
print(tf.__version__)
```
<h2> Adding two tensors </h2>
First, let's try doing this using numpy, the Python numeric package. numpy code is immediately evaluated.
```
a = np.array([5, 3, 8])
b = np.array([3, -1, 2])
c = np.add(a, b)
print(c)
```
The equivalent code in TensorFlow consists of two steps:
<p>
<h3> Step 1: Build the graph </h3>
```
a = tf.constant([5, 3, 8])
b = tf.constant([3, -1, 2])
c = tf.add(a, b)
print(c)
```
c is an Op ("Add") that returns a tensor of shape (3,) and holds int32. The shape is inferred from the computation graph.
Try the following in the cell above:
<ol>
<li> Change the 5 to 5.0, and similarly the other five numbers. What happens when you run this cell? </li>
<li> Add an extra number to a, but leave b at the original (3,) shape. What happens when you run this cell? </li>
<li> Change the code back to a version that works </li>
</ol>
<p/>
<h3> Step 2: Run the graph
```
with tf.Session() as sess:
result = sess.run(c)
print(result)
```
<h2> Using a feed_dict </h2>
Same graph, but without hardcoding inputs at build stage
```
a = tf.placeholder(dtype=tf.int32, shape=(None,)) # batchsize x scalar
b = tf.placeholder(dtype=tf.int32, shape=(None,))
c = tf.add(a, b)
with tf.Session() as sess:
result = sess.run(c, feed_dict={
a: [3, 4, 5],
b: [-1, 2, 3]
})
print(result)
```
<h2> Heron's Formula in TensorFlow </h2>
The area of triangle whose three sides are $(a, b, c)$ is $\sqrt{s(s-a)(s-b)(s-c)}$ where $s=\frac{a+b+c}{2}$
Look up the available operations at https://www.tensorflow.org/api_docs/python/tf
```
def compute_area(sides):
# slice the input to get the sides
a = sides[:,0] # 5.0, 2.3
b = sides[:,1] # 3.0, 4.1
c = sides[:,2] # 7.1, 4.8
# Heron's formula
s = (a + b + c) * 0.5 # (a + b) is a short-cut to tf.add(a, b)
areasq = s * (s - a) * (s - b) * (s - c) # (a * b) is a short-cut to tf.multiply(a, b), not tf.matmul(a, b)
return tf.sqrt(areasq)
with tf.Session() as sess:
# pass in two triangles
area = compute_area(tf.constant([
[5.0, 3.0, 7.1],
[2.3, 4.1, 4.8]
]))
result = sess.run(area)
print(result)
```
<h2> Placeholder and feed_dict </h2>
More common is to define the input to a program as a placeholder and then to feed in the inputs. The difference between the code below and the code above is whether the "area" graph is coded up with the input values or whether the "area" graph is coded up with a placeholder through which inputs will be passed in at run-time.
```
with tf.Session() as sess:
sides = tf.placeholder(tf.float32, shape=(None, 3)) # batchsize number of triangles, 3 sides
area = compute_area(sides)
result = sess.run(area, feed_dict = {
sides: [
[5.0, 3.0, 7.1],
[2.3, 4.1, 4.8]
]
})
print(result)
```
## tf.eager
tf.eager allows you to avoid the build-then-run stages. However, most production code will follow the lazy evaluation paradigm because the lazy evaluation paradigm is what allows for multi-device support and distribution.
<p>
One thing you could do is to develop using tf.eager and then comment out the eager execution and add in the session management code.
<b>You may need to restart the session to try this out.</b>
```
import tensorflow as tf
tf.enable_eager_execution()
def compute_area(sides):
# slice the input to get the sides
a = sides[:,0] # 5.0, 2.3
b = sides[:,1] # 3.0, 4.1
c = sides[:,2] # 7.1, 4.8
# Heron's formula
s = (a + b + c) * 0.5 # (a + b) is a short-cut to tf.add(a, b)
areasq = s * (s - a) * (s - b) * (s - c) # (a * b) is a short-cut to tf.multiply(a, b), not tf.matmul(a, b)
return tf.sqrt(areasq)
area = compute_area(tf.constant([
[5.0, 3.0, 7.1],
[2.3, 4.1, 4.8]
]))
print(area)
```
## Challenge Exercise
Use TensorFlow to find the roots of a fourth-degree polynomial using [Halley's Method](https://en.wikipedia.org/wiki/Halley%27s_method). The five coefficients (i.e. $a_0$ to $a_4$) of
<p>
$f(x) = a_0 + a_1 x + a_2 x^2 + a_3 x^3 + a_4 x^4$
<p>
will be fed into the program, as will the initial guess $x_0$. Your program will start from that initial guess and then iterate one step using the formula:
<img src="https://wikimedia.org/api/rest_v1/media/math/render/svg/142614c0378a1d61cb623c1352bf85b6b7bc4397" />
<p>
If you got the above easily, try iterating indefinitely until the change between $x_n$ and $x_{n+1}$ is less than some specified tolerance. Hint: Use [tf.while_loop](https://www.tensorflow.org/api_docs/python/tf/while_loop)
Copyright 2017 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License
| github_jupyter |
### Linear SCM simulations with variance shift noise interventions in Section 5.2.2
variance shift instead of mean shift
| Sim Num | name | better estimator | baseline |
| :-----------: | :--------------------------------|:----------------:| :-------:|
| (viii) | Single source anti-causal DA without Y interv + variance shift | DIP-std+ | DIP |
| (ix) | Multiple source anti-causal DA with Y interv + variance shift | CIRMweigh-std+ | OLSPool |
```
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
plt.rcParams['axes.facecolor'] = 'lightgray'
np.set_printoptions(precision=3)
sns.set(style="darkgrid")
```
#### Helper functions
```
def boxplot_all_methods(plt_handle, res_all, title='', names=[], colors=[], ylim_option=0):
res_all_df = pd.DataFrame(res_all.T)
res_all_df.columns = names
res_all_df_melt = res_all_df.melt(var_name='methods', value_name='MSE')
res_all_mean = np.mean(res_all, axis=1)
plt_handle.set_title(title, fontsize=20)
plt_handle.axhline(res_all_mean[1], ls='--', color='b')
plt_handle.axhline(res_all_mean[0], ls='--', color='r')
ax = sns.boxplot(x="methods", y="MSE", data=res_all_df_melt,
palette=colors,
ax=plt_handle)
ax.set_xticklabels(ax.get_xticklabels(), rotation=-70, ha='left', fontsize=20)
ax.tick_params(labelsize=20)
# ax.yaxis.set_major_formatter(matplotlib.ticker.FormatStrFormatter('%.2f'))
ax.yaxis.grid(False) # Hide the horizontal gridlines
ax.xaxis.grid(True) # Show the vertical gridlines
# ax.xaxis.set_visible(False)
ax.set_xlabel("")
ax.set_ylabel("MSE", fontsize=20)
if ylim_option == 1:
lower_ylim = res_all_mean[0] - (res_all_mean[1] - res_all_mean[0]) *0.3
# upper_ylim = max(res_all_mean[1] + (res_all_mean[1] - res_all_mean[0]) *0.3, res_all_mean[0]*1.2)
upper_ylim = res_all_mean[1] + (res_all_mean[1] - res_all_mean[0]) *0.3
# get the boxes that are outside of the plot
outside_index = np.where(res_all_mean > upper_ylim)[0]
for oindex in outside_index:
ax.annotate("box\nbeyond\ny limit", xy=(oindex - 0.3, upper_ylim - (upper_ylim-lower_ylim)*0.15 ), fontsize=15)
plt_handle.set_ylim(lower_ylim, upper_ylim)
def scatterplot_two_methods(plt_handle, res_all, index1, index2, names, colors=[], title="", ylimmax = -1):
plt_handle.scatter(res_all[index1], res_all[index2], alpha=1.0, marker='+', c = np.array(colors[index2]).reshape(1, -1), s=100)
plt_handle.set_xlabel(names[index1], fontsize=20)
plt_handle.set_ylabel(names[index2], fontsize=20)
plt_handle.tick_params(labelsize=20)
#
if ylimmax <= 0:
# set ylim automatically
# ylimmax = np.max((np.max(res_all[index1]), np.max(res_all[index2])))
ylimmax = np.percentile(np.concatenate((res_all[index1], res_all[index2])), 90)
print(ylimmax)
plt_handle.plot([0, ylimmax],[0, ylimmax], 'k--', alpha=0.5)
# plt.axis('equal')
plt_handle.set_xlim(0.0, ylimmax)
plt_handle.set_ylim(0.0, ylimmax)
plt_handle.set_title(title, fontsize=20)
```
#### 8. Single source anti-causal DA without Y interv + variance shift - boxplots
boxplots showwing thatDIP-std+ and DIP-MMD works
```
names_short = ["OLSTar", "OLSSrc[1]", "DIP[1]-mean", "DIP[1]-std+", "DIP[1]-MMD"]
COLOR_PALETTE1 = sns.color_palette("Set1", 9, desat=1.)
COLOR_PALETTE2 = sns.color_palette("Set1", 9, desat=.7)
COLOR_PALETTE3 = sns.color_palette("Set1", 9, desat=.5)
COLOR_PALETTE4 = sns.color_palette("Set1", 9, desat=.3)
# this corresponds to the methods in names_short
COLOR_PALETTE = [COLOR_PALETTE2[0], COLOR_PALETTE2[1], COLOR_PALETTE2[3], COLOR_PALETTE2[3], COLOR_PALETTE3[3], COLOR_PALETTE4[3]]
sns.palplot(COLOR_PALETTE)
interv_type = 'sv1'
M = 2
lamL2 = 0.
lamL1 = 0.
epochs = 20000
lr = 1e-4
n = 5000
prefix_template_exp8_box = "simu_results/sim_exp8_box_r0%sd31020_%s_lamMatch%s_n%d_epochs%d_repeats%d"
save_dir = 'paper_figures'
nb_ba = 4
results_src_ba = np.zeros((3, M-1, nb_ba, 2, 10))
results_tar_ba = np.zeros((3, 1, nb_ba, 2, 10))
savefilename_prefix = prefix_template_exp8_box %(interv_type, 'baseline', 1.,
n, epochs, 10)
res_all_ba = np.load("%s.npy" %savefilename_prefix, allow_pickle=True)
for i in range(3):
for j in range(1):
results_src_ba[i, :], results_tar_ba[i, :] = res_all_ba.item()[i, j]
lamMatches = [10.**(k) for k in (np.arange(10)-5)]
print(lamMatches)
nb_damean = 2 # DIP, DIPOracle
results_src_damean = np.zeros((3, len(lamMatches), M-1, nb_damean, 2, 10))
results_tar_damean = np.zeros((3, len(lamMatches), 1, nb_damean, 2, 10))
for k, lam in enumerate(lamMatches):
savefilename_prefix = prefix_template_exp8_box %(interv_type, 'DAmean', lam,
n, epochs, 10)
res_all_damean = np.load("%s.npy" %savefilename_prefix, allow_pickle=True)
for i in range(3):
for j in range(1):
results_src_damean[i, k, :], results_tar_damean[i, k, 0, :] = res_all_damean.item()[i, j]
nb_dastd = 2 # DIP-std, DIP-std+
results_src_dastd = np.zeros((3, len(lamMatches), M-1, nb_dastd, 2, 10))
results_tar_dastd = np.zeros((3, len(lamMatches), 1, nb_dastd, 2, 10))
for k, lam in enumerate(lamMatches):
savefilename_prefix = prefix_template_exp8_box %(interv_type, 'DAstd', lam,
n, epochs, 10)
res_all_dastd = np.load("%s.npy" %savefilename_prefix, allow_pickle=True)
for i in range(3):
for j in range(1):
# run methods on data generated from sem
results_src_dastd[i, k, :], results_tar_dastd[i, k, 0, :] = res_all_dastd.item()[i, j]
nb_dammd = 1 # DIP-MMD
results_src_dammd = np.zeros((3, len(lamMatches), M-1, nb_dammd, 2, 10))
results_tar_dammd = np.zeros((3, len(lamMatches), 1, nb_dammd, 2, 10))
for k, lam in enumerate(lamMatches):
savefilename_prefix = prefix_template_exp8_box %(interv_type, 'DAMMD', lam,
n, 2000, 10)
res_all_dammd = np.load("%s.npy" %savefilename_prefix, allow_pickle=True)
for i in range(3):
for j in range(1):
# run methods on data generated from sem
results_src_dammd[i, k, :], results_tar_dammd[i, k, 0, :] = res_all_dammd.item()[i, j]
# now add the methods
results_tar_plot = {}
lamMatchIndex = 6
print("Fix lambda choice: lmabda = ", lamMatches[lamMatchIndex])
for i in range(3):
results_tar_plot[i] = np.concatenate((results_tar_ba[i, 0, :2, 0, :],
results_tar_damean[i, lamMatchIndex, 0, 0, 0, :].reshape(1, -1),
results_tar_dastd[i, lamMatchIndex, 0, 1, 0, :].reshape(1, -1),
results_tar_dammd[i, lamMatchIndex, 0, 0, 0, :].reshape(1, -1)), axis=0)
ds = [3, 10, 20]
fig, axs = plt.subplots(1, 3, figsize=(20,5))
for i in range(3):
boxplot_all_methods(axs[i], results_tar_plot[i],
title="linear SCM: d=%d" %(ds[i]), names=names_short, colors=COLOR_PALETTE[:len(names_short)])
plt.subplots_adjust(top=0.9, bottom=0.1, left=0.1, right=0.9, hspace=0.6,
wspace=0.2)
plt.savefig("%s/sim_6_2_exp_%s.pdf" %(save_dir, interv_type), bbox_inches="tight")
plt.show()
fig, axs = plt.subplots(1, 1, figsize=(5,5))
boxplot_all_methods(axs, results_tar_plot[1],
title="linear SCM: d=%d" %(10), names=names_short, colors=COLOR_PALETTE[:len(names_short)])
plt.subplots_adjust(top=0.9, bottom=0.1, left=0.1, right=0.9, hspace=0.6,
wspace=0.2)
plt.savefig("%s/sim_6_2_exp_%s_single10.pdf" %(save_dir, interv_type), bbox_inches="tight")
plt.show()
```
#### 8 Single source anti-causal DA without Y interv + variance shift - scatterplots
Scatterplots showing that DIP-std+ and DIP-MMD works
```
interv_type = 'sv1'
M = 2
lamL2 = 0.
lamL1 = 0.
epochs = 20000
lr = 1e-4
n = 5000
prefix_template_exp8_scat = "simu_results/sim_exp8_scat_r0%sd10_%s_lamMatch%s_n%d_epochs%d_seed%d"
nb_ba = 4
repeats = 100
results_scat_src_ba = np.zeros((M-1, nb_ba, 2, repeats))
results_scat_tar_ba = np.zeros((1, nb_ba, 2, repeats))
for myseed in range(100):
savefilename_prefix = prefix_template_exp8_scat %(interv_type, 'baseline', 1.,
n, epochs, myseed)
res_all_ba = np.load("%s.npy" %savefilename_prefix, allow_pickle=True)
results_scat_src_ba[0, :, :, myseed] = res_all_ba.item()['src'][:, :, 0]
results_scat_tar_ba[0, :, :, myseed] = res_all_ba.item()['tar'][:, :, 0]
lamMatches = [10.**(k) for k in (np.arange(10)-5)]
nb_damean = 2 # DIP, DIPOracle
results_scat_src_damean = np.zeros((len(lamMatches), M-1, nb_damean, 2, 100))
results_scat_tar_damean = np.zeros((len(lamMatches), 1, nb_damean, 2, 100))
for myseed in range(100):
for k, lam in enumerate(lamMatches):
savefilename_prefix = prefix_template_exp8_scat %(interv_type, 'DAmean', lam,
n, epochs, myseed)
res_all_damean = np.load("%s.npy" %savefilename_prefix, allow_pickle=True)
results_scat_src_damean[k, 0, :, :, myseed] = res_all_damean.item()['src'][:, :, 0]
results_scat_tar_damean[k, 0, :, :, myseed] = res_all_damean.item()['tar'][:, :, 0]
nb_dastd = 2 # DIP-std, DIP-std+
results_scat_src_dastd = np.zeros((len(lamMatches), M-1, nb_dastd, 2, 100))
results_scat_tar_dastd = np.zeros((len(lamMatches), 1, nb_dastd, 2, 100))
for myseed in range(100):
for k, lam in enumerate(lamMatches):
savefilename_prefix = prefix_template_exp8_scat %(interv_type, 'DAstd', lam,
n, epochs, myseed)
res_all_dastd = np.load("%s.npy" %savefilename_prefix, allow_pickle=True)
results_scat_src_dastd[k, 0, :, :, myseed] = res_all_dastd.item()['src'][:, :, 0]
results_scat_tar_dastd[k, 0, :, :, myseed] = res_all_dastd.item()['tar'][:, :, 0]
nb_dammd = 1 # DIP-MMD
results_scat_src_dammd = np.zeros((len(lamMatches), M-1, nb_dammd, 2, 100))
results_scat_tar_dammd = np.zeros((len(lamMatches), 1, nb_dammd, 2, 100))
for myseed in range(100):
for k, lam in enumerate(lamMatches):
savefilename_prefix = prefix_template_exp8_scat %(interv_type, 'DAMMD', lam,
n, 2000, myseed)
res_all_dammd = np.load("%s.npy" %savefilename_prefix, allow_pickle=True)
results_scat_src_dammd[k, 0, :, :, myseed] = res_all_dammd.item()['src'][:, :, 0]
results_scat_tar_dammd[k, 0, :, :, myseed] = res_all_dammd.item()['tar'][:, :, 0]
# now add the methods
results_tar_plot = {}
lamMatchIndex = 6
print("Fix lambda choice: lmabda = ", lamMatches[lamMatchIndex])
results_scat_tar_plot = np.concatenate((results_scat_tar_ba[0, :2, 0, :],
results_scat_tar_damean[lamMatchIndex, 0, 0, 0, :].reshape(1, -1),
results_scat_tar_dastd[lamMatchIndex, 0, 0, 0, :].reshape(1, -1),
results_scat_tar_dammd[lamMatchIndex, 0, 0, 0, :].reshape(1, -1)), axis=0)
for index1, name1 in enumerate(names_short):
for index2, name2 in enumerate(names_short):
if index2 > index1:
fig, axs = plt.subplots(1, 1, figsize=(5,5))
nb_below_diag = np.sum(results_scat_tar_plot[index1, :] >= results_scat_tar_plot[index2, :])
scatterplot_two_methods(axs, results_scat_tar_plot, index1, index2, names_short, COLOR_PALETTE[:len(names_short)],
title="%d%% of pts below the diagonal" % (nb_below_diag),
ylimmax = -1)
plt.savefig("%s/sim_6_2_exp_%s_single_repeat_%s_vs_%s.pdf" %(save_dir, interv_type, name1, name2), bbox_inches="tight")
plt.show()
```
#### 9 Multiple source anti-causal DA with Y interv + variance shift - scatterplots
Scatterplots showing that CIRM-std+ and CIRM-MMD works
```
interv_type = 'smv1'
M = 15
lamL2 = 0.
lamL1 = 0.
epochs = 20000
lr = 1e-4
n = 5000
prefix_template_exp9 = "simu_results/sim_exp9_scat_r0%sd20x4_%s_lamMatch%s_lamCIP%s_n%d_epochs%d_seed%d"
nb_ba = 4 # OLSTar, SrcPool, OLSTar, SrcPool
results9_src_ba = np.zeros((M-1, nb_ba, 2, 100))
results9_tar_ba = np.zeros((1, nb_ba, 2, 100))
for myseed in range(100):
savefilename_prefix = prefix_template_exp9 %(interv_type, 'baseline',
1., 0.1, n, epochs, myseed)
res_all_ba = np.load("%s.npy" %savefilename_prefix, allow_pickle=True)
results9_src_ba[0, :, :, myseed] = res_all_ba.item()['src'][0, :, 0]
results9_tar_ba[0, :, :, myseed] = res_all_ba.item()['tar'][:, :, 0]
lamMatches = [10.**(k) for k in (np.arange(10)-5)]
nb_damean = 5 # DIP, DIPOracle, DIPweigh, CIP, CIRMweigh
results9_src_damean = np.zeros((len(lamMatches), M-1, nb_damean, 2, 100))-1
results9_tar_damean = np.zeros((len(lamMatches), 1, nb_damean, 2, 100))-1
for myseed in range(100):
for k, lam in enumerate(lamMatches):
savefilename_prefix = prefix_template_exp9 %(interv_type, 'DAmean',
lam, 0.1, 5000, epochs, myseed)
res_all_damean = np.load("%s.npy" %savefilename_prefix, allow_pickle=True)
results9_src_damean[k, :, :, :, myseed] = res_all_damean.item()['src'][:, :, :, 0]
results9_tar_damean[k, 0, :, :, myseed] = res_all_damean.item()['tar'][:, :, 0]
nb_dastd = 4 # DIP-std+, DIPweigh-std+, CIP-std+, CIRMweigh-std+
results9_src_dastd = np.zeros((len(lamMatches), M-1, nb_dastd, 2, 100))-1
results9_tar_dastd = np.zeros((len(lamMatches), 1, nb_dastd, 2, 100))-1
for myseed in range(100):
for k, lam in enumerate(lamMatches):
savefilename_prefix = prefix_template_exp9 %(interv_type, 'DAstd',
lam, 0.1, 5000, epochs, myseed)
res_all_dastd = np.load("%s.npy" %savefilename_prefix, allow_pickle=True)
results9_src_dastd[k, :, :, :, myseed] = res_all_dastd.item()['src'][:, :, :, 0]
results9_tar_dastd[k, 0, :, :, myseed] = res_all_dastd.item()['tar'][:, :, 0]
nb_dammd = 4 # DIP-MMD, DIPweigh-MMD, CIP-MMD, CIRMweigh-MMMD
results9_src_dammd = np.zeros((len(lamMatches), M-1, nb_dammd, 2, 100))-1
results9_tar_dammd = np.zeros((len(lamMatches), 1, nb_dammd, 2, 100))-1
for myseed in range(100):
for k, lam in enumerate(lamMatches):
savefilename_prefix = prefix_template_exp9 %(interv_type, 'DAMMD',
lam, 0.1, 5000, 2000, myseed)
res_all_dammd = np.load("%s.npy" %savefilename_prefix, allow_pickle=True)
results9_src_dammd[k, :, :, :, myseed] = res_all_dammd.item()['src'][:, :, :, 0]
results9_tar_dammd[k, 0, :, :, myseed] = res_all_dammd.item()['tar'][:, :, 0]
# now add the methods
names_short = ["Tar", "SrcPool", "CIRMweigh-mean", "CIRMweigh-std+", "DIPweigh-MMD", "CIRMweigh-MMD"]
COLOR_PALETTE1 = sns.color_palette("Set1", 9, desat=1.)
COLOR_PALETTE = [COLOR_PALETTE1[k] for k in [0, 1, 2, 3, 4, 7, 6]]
COLOR_PALETTE = [COLOR_PALETTE[k] for k in [0, 1, 6, 6, 4, 6]]
sns.palplot(COLOR_PALETTE)
lamMatchIndex = 6
print("Fix lambda choice: lmabda = ", lamMatches[lamMatchIndex])
results9_tar_plot = np.concatenate((results9_tar_ba[0, :2, 0, :],
results9_tar_damean[lamMatchIndex, 0, 4, 0, :].reshape(1, -1),
results9_tar_dastd[lamMatchIndex, 0, 3, 0, :].reshape(1, -1),
results9_tar_dammd[lamMatchIndex, 0, 1, 0, :].reshape(1, -1),
results9_tar_dammd[lamMatchIndex, 0, 3, 0, :].reshape(1, -1)), axis=0)
for index1, name1 in enumerate(names_short):
for index2, name2 in enumerate(names_short):
if index2 > index1:
fig, axs = plt.subplots(1, 1, figsize=(5,5))
nb_below_diag = np.sum(results9_tar_plot[index1, :] >= results9_tar_plot[index2, :])
scatterplot_two_methods(axs, results9_tar_plot, index1, index2, names_short, COLOR_PALETTE[:len(names_short)],
title="%d%% of pts below the diagonal" % (nb_below_diag),
ylimmax = -1)
plt.savefig("%s/sim_6_2_exp_%s_y_shift_single_repeat_%s_vs_%s.pdf" %(save_dir, interv_type, name1, name2), bbox_inches="tight")
plt.show()
```
| github_jupyter |
# General Imports
!! IMPORTANT !!
If you did NOT install opengrid with pip,
make sure the path to the opengrid folder is added to your PYTHONPATH
```
import os
import inspect
import sys
import pandas as pd
import charts
from opengrid_dev.library import houseprint
import matplotlib.pyplot as plt
%matplotlib inline
plt.rcParams['figure.figsize'] = 16,8
```
## Houseprint
```
hp = houseprint.Houseprint()
# for testing:
# hp = houseprint.Houseprint(spreadsheet='unit and integration test houseprint')
hp
hp.sites[:5]
hp.get_devices()[:4]
hp.get_sensors('water')[:3]
```
A Houseprint object can be saved as a pickle. It loses its tmpo session however (connections cannot be pickled)
```
hp.save('new_houseprint.pkl')
hp = houseprint.load_houseprint_from_file('new_houseprint.pkl')
```
### TMPO
The houseprint, sites, devices and sensors all have a get_data method. In order to get these working for the fluksosensors, the houseprint creates a tmpo session.
```
hp.init_tmpo()
hp._tmpos.debug = False
hp.sync_tmpos()
```
## Lookup sites, devices, sensors based on key
These methods return a single object
```
hp.find_site(1)
hp.find_device('FL03001562')
sensor = hp.find_sensor('d5a747b86224834f745f4c9775d70241')
print(sensor.site)
print(sensor.unit)
```
## Lookup sites, devices, sensors based on search criteria
These methods return a list with objects satisfying the criteria
```
hp.search_sites(inhabitants=5)
hp.search_sensors(type='electricity', direction='Import')
```
### Get Data
```
hp.sync_tmpos()
head = pd.Timestamp('20150101')
tail = pd.Timestamp('20160101')
df = hp.get_data(sensortype='water', head=head, tail=tail, diff=True, resample='min', unit='l/min')
#charts.plot(df, stock=True, show='inline')
df.info()
```
## Site
```
site = hp.find_site(1)
site
print(site.size)
print(site.inhabitants)
print(site.postcode)
print(site.construction_year)
print(site.k_level)
print(site.e_level)
print(site.epc_cert)
site.devices
site.get_sensors('electricity')
head = pd.Timestamp('20150617')
tail = pd.Timestamp('20150628')
df=site.get_data(sensortype='electricity', head=head,tail=tail, diff=True, unit='kW')
charts.plot(df, stock=True, show='inline')
```
## Device
```
device = hp.find_device('FL03001552')
device
device.key
device.get_sensors('gas')
head = pd.Timestamp('20151101')
tail = pd.Timestamp('20151104')
df = hp.get_data(sensortype='gas', head=head,tail=tail, diff=True, unit='kW')
charts.plot(df, stock=True, show='inline')
```
## Sensor
```
sensor = hp.find_sensor('53b1eb0479c83dee927fff10b0cb0fe6')
sensor
sensor.key
sensor.type
sensor.description
sensor.system
sensor.unit
head = pd.Timestamp('20150617')
tail = pd.Timestamp('20150618')
df=sensor.get_data(head,tail,diff=True, unit='W')
charts.plot(df, stock=True, show='inline')
```
## Getting data for a selection of sensors
```
sensors = hp.search_sensors(type='electricity', system='solar')
print(sensors)
df = hp.get_data(sensors=sensors, head=head, tail=tail, diff=True, unit='W')
charts.plot(df, stock=True, show='inline')
```
## Dynamically loading data sensor per sensor
A call to `hp.get_data()` is lazy: it creates a big list of Data Series per sensor and concatenates them. This can take a while, specifically when you need many sensors and a large time span.
Often, you don't use the big DataFrame as a whole, you rather re-divide it by using a for loop and looking at each sensor individually.
By using `hp.get_data_dynamic()`, data is fetched from tmpo per sensor at a time, just when you need it.
```
dyn_data = hp.get_data_dynamic(sensortype='electricity', head=head, tail=tail)
ts = next(dyn_data)
df = pd.DataFrame(ts)
charts.plot(df, stock=True, show='inline')
```
You can run the cell above multiple times and eacht time the next sensor will be fetched
| github_jupyter |
<a href="https://colab.research.google.com/github/MoffatKirui/MoffatCoreW2/blob/main/Moringa_Data_Science_Core_W2_Independent_Project_Moffat_Kirui_Python_Notebook.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
## 1. Defining the Question
### a) Specifying the Data Analytic Question
```
#Determine how to predict which individuals are most likely to have or use a bank account.
```
### b) Defining the Metric for Success
coming up with a solution that predict individuals who are likely to have a bank account basing on predictor variables
### c) Understanding the context
Financial Inclusion remains one of the main obstacles to economic and human development in Africa. For example, across Kenya, Rwanda, Tanzania, and Uganda only 9.1 million adults (or 13.9% of the adult population) have access to or use a commercial bank account.
Traditionally, access to bank accounts has been regarded as an indicator of financial inclusion. Despite the proliferation of mobile money in Africa and the growth of innovative fintech solutions, banks still play a pivotal role in facilitating access to financial services. Access to bank accounts enables households to save and facilitate payments while also helping businesses build up their credit-worthiness and improve their access to other financial services. Therefore, access to bank accounts is an essential contributor to long-term economic growth.
### d) Recording the Experimental Design
* loading provided dataset
performing data cleaning
* Finding and dealing with outliers, anomalies, and missing data within the dataset.
* Performing of univariate, bivariate and multivariate analysis recording of observations.
* Implementing the solution .
* Challenge your solution.
### e) Data Relevance
A complete description of the provided dataset is found in the link below.
* Variable Definitions: [link](http://bit.ly/VariableDefinitions )
## 2. Reading the Data
```
# Loading the Data from the source i.e. csv
#import required libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style()
url = "http://bit.ly/FinancialDataset"
findata = pd.read_csv(url)
```
## 3. Checking the Data
```
# Determining the no. of records in our dataset
#
findata.shape
# Previewing the top of our dataset
#
findata.head()
# Previewing the bottom of our dataset
#
findata.tail()
# Checking whether each column has an appropriate datatype
#
findata.dtypes
```
## 4. External Data Source Validation
The main dataset contains demographic information and what financial services are used by individuals across East Africa. This data was extracted from various Finscope surveys ranging from 2016 to 2018, and more information about these surveys can be found here:
* FinAccess Kenya 2018. [[Link]](https://fsdkenya.org/publication/finaccess2019/)
* Finscope Rwanda 2016. [[Link]](http://www.statistics.gov.rw/publication/finscope-rwanda-2016)
* Finscope Tanzania 2017. [[Link]](http://www.fsdt.or.tz/finscope/)
* Finscope Uganda 2018. [[Link]](http://fsduganda.or.ug/finscope-2018-survey-report/)
## 5. Tidying the Dataset
```
# Checking for Outliers
#
sns.boxplot(findata.household_size, showmeans=True)
plt.title('Household size Boxplot')
sns.boxplot(findata['Respondent Age'], showmeans=True)
plt.title('Age Boxplot')
#identifying missing data
#
findata.isnull().sum()
# Dealing with the Missing Data
#dropping records with null values in the 'Has a Bank account' column as it is our main point of focus
findata.dropna(inplace=True)
findata.isnull().sum()
# More data cleaning procedures
findata.drop(['country','uniqueid','year'],axis=1,inplace=True)
#changing column names to upper case and placing underscore to make the column names be more presentable
findata.columns = findata.columns.str.strip().str.upper().str.replace(' ', '_').str.replace('(', '').str.replace(')', '')
```
## 6. Exploratory Analysis
### Univariate Analysis
```
# Ploting the univariate summaries and recording our observations
#
findata.describe(include='all')
#mode for the household size
findata.HOUSEHOLD_SIZE.mode()
#mode for respondent age
findata.RESPONDENT_AGE.mode()
print('Age Variance=',findata.RESPONDENT_AGE.var())
print('Age range=',findata.RESPONDENT_AGE.max()-findata.RESPONDENT_AGE.min())
print('Age IQR=', findata.RESPONDENT_AGE.quantile(0.75)-findata.RESPONDENT_AGE.quantile(0.25))
print('household size variance=',findata.HOUSEHOLD_SIZE.var())
print('household size range=',findata.HOUSEHOLD_SIZE.max()-findata.HOUSEHOLD_SIZE.min())
print('househod size IQR=', findata.HOUSEHOLD_SIZE.quantile(0.75)-findata.HOUSEHOLD_SIZE.quantile(0.25))
sns.distplot(findata.RESPONDENT_AGE)
findata.RESPONDENT_AGE.skew()
findata.RESPONDENT_AGE.kurt()
sns.displot(findata.HOUSEHOLD_SIZE)
findata.HOUSEHOLD_SIZE.skew()
findata.HOUSEHOLD_SIZE.kurt()
```
#### Bar charts
```
#fig, ((ax1, ax2, ax3, ax4), (ax5, ax6, ax7, ax8)) = plt.subplots(4,2, figsize=(20, 10))
#fig.suptitle('Bar charts')
findata.groupby('HAS_A_BANK_ACCOUNT')['HAS_A_BANK_ACCOUNT'].count().plot.barh()
plt.show()
findata.groupby('TYPE_OF_LOCATION')['TYPE_OF_LOCATION'].count().plot.barh()
plt.show()
findata.groupby('CELL_PHONE_ACCESS')['CELL_PHONE_ACCESS'].count().plot.barh()
plt.show()
findata.groupby('GENDER_OF_RESPONDENT')['GENDER_OF_RESPONDENT'].count().plot.barh()
plt.show()
findata.groupby('THE_RELATHIP_WITH_HEAD')['THE_RELATHIP_WITH_HEAD'].count().plot.barh()
plt.show()
findata.groupby('MARITAL_STATUS')['MARITAL_STATUS'].count().plot.barh()
plt.show()
findata.groupby('LEVEL_OF_EDUCUATION')['LEVEL_OF_EDUCUATION'].count().plot.barh()
plt.show()
findata.groupby('TYPE_OF_JOB')['TYPE_OF_JOB'].count().plot.barh()
plt.show()
```
### Bivariate analysis
```
findata.plot(x='RESPONDENT_AGE',y='HOUSEHOLD_SIZE',kind= 'scatter')
pearson_coeff = findata["RESPONDENT_AGE"].corr(findata["HOUSEHOLD_SIZE"], method="pearson")
print(pearson_coeff)
findata.groupby(['TYPE_OF_LOCATION','HAS_A_BANK_ACCOUNT'])['HAS_A_BANK_ACCOUNT'].count().unstack().plot.bar()
findata.groupby(['TYPE_OF_JOB','HAS_A_BANK_ACCOUNT'])['HAS_A_BANK_ACCOUNT'].count().unstack().plot.barh()
findata.groupby(['CELL_PHONE_ACCESS','HAS_A_BANK_ACCOUNT'])['HAS_A_BANK_ACCOUNT'].count().unstack().plot.bar()
findata.groupby(['GENDER_OF_RESPONDENT','HAS_A_BANK_ACCOUNT'])['HAS_A_BANK_ACCOUNT'].count().unstack().plot.bar()
findata.groupby(['THE_RELATHIP_WITH_HEAD','HAS_A_BANK_ACCOUNT'])['HAS_A_BANK_ACCOUNT'].count().unstack().plot.bar()
findata.groupby(['MARITAL_STATUS','HAS_A_BANK_ACCOUNT'])['HAS_A_BANK_ACCOUNT'].count().unstack().plot.bar()
findata.groupby(['LEVEL_OF_EDUCUATION','HAS_A_BANK_ACCOUNT'])['HAS_A_BANK_ACCOUNT'].count().unstack().plot.bar()
findata.groupby(['RESPONDENT_AGE','HAS_A_BANK_ACCOUNT'])['HAS_A_BANK_ACCOUNT'].count().unstack().plot.line()
findata.groupby(['HOUSEHOLD_SIZE','HAS_A_BANK_ACCOUNT'])['HAS_A_BANK_ACCOUNT'].count().unstack().plot.line()
```
### Multivariate Analysis
#### label encoding
```
#label encoding data to binary
from sklearn import preprocessing
label = preprocessing.LabelEncoder()
findata['GENDER_OF_RESPONDENT']= label.fit_transform(findata['GENDER_OF_RESPONDENT'])
findata['HAS_A_BANK_ACCOUNT']= label.fit_transform(findata['HAS_A_BANK_ACCOUNT'])
findata['CELL_PHONE_ACCESS']= label.fit_transform(findata['CELL_PHONE_ACCESS'])
findata['TYPE_OF_LOCATION']= label.fit_transform(findata['TYPE_OF_LOCATION'])
#one hot encoding columns with more than two categories
k=pd.get_dummies(findata.iloc[:,6:],prefix=['THE_RELATHIP_WITH_HEAD','MARITAL_STATUS','LEVEL_OF_EDUCUATION','TYPE_OF_JOB'])
findata.drop(['THE_RELATHIP_WITH_HEAD','MARITAL_STATUS','LEVEL_OF_EDUCUATION','TYPE_OF_JOB'],axis=1,inplace=True)
#concatinating the one hot encoded dataframe to the initial dataframe
t=[findata,k]
j=pd.concat(t,axis=1)
j.head()
```
#### PCA
```
# The first preprocessing step is to divide the dataset into a feature set and corresponding labels.
# The following script performs this task. The script below stores the feature sets into the X variable
# and the series of corresponding labels in to the y variable.
#
X = j.drop('HAS_A_BANK_ACCOUNT', 1)
y = j['HAS_A_BANK_ACCOUNT']
# The next preprocessing step is to divide data into training and test sets.
# We execute the following script to do so:
#
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=0)
# We will perform standard scalar normalization to normalize our feature set.
# To do this, we execute the following code:
#
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
# Performing PCA using Scikit-Learn is a two-step process:
# Initialize the PCA class by passing the number of components to the constructor.
# Call the fit and then transform methods by passing the feature set to these methods.
# The transform method returns the specified number of principal components.
# Let's take a look at the following code. In the code above, we create a PCA object named pca.
# We did not specify the number of components in the constructor.
# Hence, all four of the features in the feature set will be returned for both the training and test sets.
#
from sklearn.decomposition import PCA
pca = PCA()
X_train = pca.fit_transform(X_train)
X_test = pca.transform(X_test)
# The PCA class contains explained_variance_ratio_ which returns the variance caused by each of the principal components.
# We execute the following line of code to find the "explained variance ratio".
#
explained_variance = pca.explained_variance_ratio_
explained_variance
# In this case we'll use random forest classification for making the predictions.
#
from sklearn.ensemble import RandomForestClassifier
classifier = RandomForestClassifier(max_depth=2, random_state=0)
classifier.fit(X_train, y_train)
# Predicting the Test set results
y_pred = classifier.predict(X_test)
# Performance Evaluation
#
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score
cm = confusion_matrix(y_test, y_pred)
print(cm)
print('Accuracy' , accuracy_score(y_test, y_pred))
```
## Challenging the solution
#### LDA
```
# Performing LDA
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis as LDA
lda = LDA(n_components=1)
X_train = lda.fit_transform(X_train, y_train)
X_test = lda.transform(X_test)
# Training and Making Predictions
# We will use the random forest classifier to evaluate the performance of a LDA-reduced algorithms as shown
#
from sklearn.ensemble import RandomForestClassifier
classifier = RandomForestClassifier(max_depth=2, random_state=0)
classifier.fit(X_train, y_train)
y_pred = classifier.predict(X_test)
# Performance Evaluation
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score
cm = confusion_matrix(y_test, y_pred)
print(cm)
print('Accuracy' , accuracy_score(y_test, y_pred))
```
LDA with an accuracy of 88% has a better performance in predicting the category than PCA which had 87%
## Follow-up questions
#### Did we have the right data?
```
#yes we had the right data since it contained information that allow us to make prediction with a relatively high accuracy.
```
#### Do we need other data?
```
#Additional data for example containing details of annual incomes could help improve on the accuracy of our prediction
```
#### Did we have the right question?
```
Yes. because access to bank accounts has been regarded as an indicator of financial inclusion thus implementing a solution to predict the likelihood of an individual having a bank account seems right.
```
| github_jupyter |
```
import pymc3 as pm
import arviz as az
from pymc3.math import switch
import numpy as np
```
## Bayes Factor for Two Different Coin Models
This is taken from chapter 10 of Kruschke's book. We hypothesize two types of coins. One type of coin is hypothesized to be tail-biased and the other type of coin is head-biased. These two possibilities will form our two hypotheses and we will calculate Bayes factors to evaluate their relative credibility.
```
with pm.Model() as eqPrior:
pm1 = pm.Categorical('pm1', [.5, .5])
omega_0 = .25
kappa_0 = 12
theta_0 = pm.Beta('theta_0', mu=.25, sigma=.25)
omega_1 = .75
kappa_1 = 12
theta_1 = pm.Beta('theta_1', mu=.75, sigma=.25)
theta = pm.math.switch(pm.math.eq(pm1, 0), theta_0, theta_1)
y2 = pm.Bernoulli('y2', theta, observed=[1,1,0,0,0,0,0,0])
with eqPrior:
trace2 = pm.sample(10000)
pm1 = trace2['pm1'].mean() # mean value of model indicator variable
```
The posterior is provided by the estimated value of the model indicator variable, `pm1`.
```
print(f'Posterior: p(model 1|data) = {pm1:.2f}')
print(f'Posterior: p(model 2|data) = {(1-pm1):.2f}')
print(f'Posterior odds: p(model 2|data)/p(model 1|data) = {(1-pm1)/pm1:.2f}')
print(f'Bayes factor: p(model 2|data)/p(model 1|data) * p(model 1)/p(model 2) = {(1-pm1)/pm1 * (.5/.5):.2f}')
```
So our posterior odd is identical to our Bayes factor. This is because our prior on the model indicator variable gave equal credibility to each model.
```
with pm.Model() as uneqPrior:
pm1 = pm.Categorical('pm1', [.25, .75])
omega_0 = .25
kappa_0 = 12
theta_0 = pm.Beta('theta_0', mu=.25, sigma=.25)
omega_1 = .75
kappa_1 = 12
theta_1 = pm.Beta('theta_1', mu=.75, sigma=.25)
theta = pm.math.switch(pm.math.eq(pm1, 0), theta_0, theta_1)
y2 = pm.Bernoulli('y2', theta, observed=[1,1,0,0,0,0,0,0])
with uneqPrior:
trace2 = pm.sample(10000)
pm1 = trace2['pm1'].mean() # mean value of model indicator variable
print(f'Posterior: p(model 1|data) = {pm1:.2f}')
print(f'Posterior: p(model 2|data) = {(1-pm1):.2f}')
print(f'Posterior odds: p(model 1|data)/p(model 2|data) = {pm1/(1-pm1):.2f}')
print(f'Bayes factor: p(model 1|data)/p(model 2|data) * p(model 2)/p(model 1) = {pm1/(1-pm1) * (.75/.25):.2f}')
```
Here, the posteroir odds and the Bayes factor are different because we gave more (prior) credibility to model 2. So the posterior probabilies of the two model are nearly identical, but that reflects the our priors (favoring model 2) and the likelihoods (favoring model 1) more or less cancelling each other out.
## Bayes Factor with a "Null" Hypothesis
Let's test a more traditional "null hypothesis". Here, we will posit two types of coins. One type is characterized by a value of theta that is exactly 0.5. We have absolute confidence that such a coin's value of theta is not .4999999999 nor .5000000001, etc. The other type of coin could have any value of theta (0-1) and all values are equally credible a priori. We then observe some data and ask whether such data should convince us that the coin is "fair" (H_0) or not (H_1).
```
n_heads = 2
n_tails = 8
data3 = np.repeat([1, 0], [n_heads, n_tails])
with pm.Model() as model3:
pm1 = pm.Categorical('pm1', [.5, .5])
theta_0 = 0.5
theta_1 = pm.Beta('theta_1', 1, 1)
theta = pm.math.switch(pm.math.eq(pm1, 0), theta_0, theta_1)
y2 = pm.Bernoulli('y2', theta, observed=data3)
with model3:
trace3 = pm.sample(10000, tune=5000)
pm1 = trace3['pm1'].mean() # mean value of model indicator variable
pm1
print(f'Posterior: p(model 1|data) = {pm1:.2f}')
print(f'Posterior: p(model 2|data) = {(1-pm1):.2f}')
print(f'Bayes factor: p(model 1|data)/p(model 2|data) * p(model 2)/p(model 1) = {pm1/(1-pm1) * (.5/.5):.2f}')
```
So we have no good evidence that would allow us to choose between our 2 hypotheses. The data isn't particularly consistent with our "null hypothesis". A priori, the alternative hypothesis entails many credible values of theta that are much more consistent with the observed data (e.g., theta = .2). However, this alternative hypothesis also entails many values of theta that are **highly** inconsistent with the observed data (e.g., theta = .9999). So the "null" suffers because there is poor agreement with the data (i.e., likelihood) whereas the alternative hypothesis suffers because it is too agnostic about the possible values of theta.
Let's compare this to a traditional, frequentist procedure to compare these models. We will first find the most likelihood of theta permitted each model, find the likelihood that each of these values yields, and then take the ratio of these likelihoods.
To get a quick approximation of the maximum likelihood associated with our alternative hypothesis, we can plot the posterior and request a mode from the kernel density estimate.
```
az.plot_posterior(trace3['theta_1'], point_estimate='mode');
```
So the value of theta that gives us the maximum likelihood is 0.2 (which makes sense because we observed 2 heads in our 10 flips). So we can use that. Of course our null hypothesis has theta fixed at 0.5.
```
mle_h0 = .5
mle_h1 = .2 # 20% of flips were heads
```
These are the values of theta that maximize the likelihood of the observed data. Now we need to know what the likelihood of our observed data under each of these values of theta. We know how to calculate the likelihood of a set of flips from earlier.
```
def likelihood(theta, n_flips, n_heads):
return (theta**n_heads) * ( (1-theta)**(n_flips - n_heads) )
likelihood_h0 = likelihood(mle_h0, n_heads+n_tails, n_heads)
print(f'likelihood_h0 = {likelihood_h0:.4f}')
likelihood_h1 = likelihood(mle_h1, n_heads+n_tails, n_heads)
print(f'likelihood_h1 = {likelihood_h1:.4f}')
print(f'Likelihood Ratio = {likelihood_h1 / likelihood_h0:.2f}')
```
In the limit of large data, likelihood ratios (multiplied by 2) are chi-squared distributed, with a degree of freedom equal to the difference in the number of parameters in the two models. Here, our alternative hypothesis has 1 parameter (theta) and our null hypothesis doesn't have any. So the df=1. Let's calculate a p-value.
```
from scipy.stats import chi2
print(f'p = {1 - chi2.cdf(2 * (likelihood_h1 / likelihood_h0), 1):.4f}')
```
So our likelihood-ratio test is suggesting that we should be extremely skeptical of our null hypothesis whereas the Bayes factor was basically ambivalent. What is going on?
The key difference between the Bayes factor and the likelihood-ratio test is that the Bayes factor treats our alternative hypothesis as embodying the full prior (i.e., theta~U(0,1)), whereas the likelihood ratio test, being a frequestist test, doesn't know anything about our priors. As a result, the likelihood-ratio test permits the alternative hypothesis to reflect whatever value of theta is most consistent with the observed data (i.e., the maximium likelihood estimate). But that's an extrodinary degree of flexibility. Our alternative hypothesis gets to adapt itself to the data it is seeking to explain, no matter how credible the final estimate was before we observed the data. This means that we should construct our hypotheses so as to be as open-minded and agnostic as possible, because we are only penalized when we observe data that are inconsistent with any configuration of our hypothesis (e.g., combination of parameter values). We are penalized for being unparsimonious, but only coarsely (i.e., the alternative hypothesis is penalized for having 1 more parameter than our "null").
In the Bayes factor, our agnosticism about the cedible values of theta represents a substantial tradeoff. Being uncertain is good because an uncertain hypothesis will be somewhat consistent with many different patterns of data that **might** be observed. However, an uncertain hypotheiss will also be consistent with many different patterns of data that **were not** observed. The former is good, but the latter is bad. The Bayes factor (and all Bayesian approaches) appropriately balance both of these facets and does so thoroughly (incorporating the prior credibility of each parameter value and how the likleihood of the data in light of each parameter value). This is the sense in which people say that Bayesian approaches naturally ensure parsimony. The more agnostic you are (regardless of how many parameters your model has), the less parsimonious your hypotheses are, and the lower the likelihood of the overall model will be.
To see this in action, let's consider the same 2 hypothesis but evaluate them on a data set that is highly likely under the "null". In this data set, 50% of flips come up heads. In a frequentist context, our two hypotheses are indistinguisable. In a Bayesian context, the parsimony of the "null" should cause it to win out over the more agnostic alterntive hypothesis.
```
n_heads = 10
n_tails = 10
data3 = np.repeat([1, 0], [n_heads, n_tails])
with pm.Model() as model4:
pm1 = pm.Categorical('pm1', [.5, .5])
theta_0 = 0.5
theta_1 = pm.Beta('theta_1', 1, 1)
theta = pm.math.switch(pm.math.eq(pm1, 0), theta_0, theta_1)
y2 = pm.Bernoulli('y2', theta, observed=data3)
with model4:
trace4 = pm.sample(10000, tune=5000)
pm1 = trace4['pm1'].mean() # mean value of model indicator variable
print(f'Posterior: p(model 1|data) = {pm1:.2f}')
print(f'Posterior: p(model 2|data) = {(1-pm1):.2f}')
print(f'Bayes factor: p(model 2|data)/p(model 1|data) * p(model 1)/p(model 2) = {(1-pm1)/pm1 * (.5/.5):.2f}')
```
Using our t-shirt guide to interpreting Bayes factor, we have "substantial evidence" in favor of our "null" hypothesis. Why? Because our alternative hypothesis was agnostic and implied that many theta values were credible. The "null", in contrast, committed to exactly 1. So our null hypothesis is far more parsimonious than the alternative.
What does the likelihood-ratio test have to say about this?
```
az.plot_posterior(trace4['theta_1'], point_estimate='mode');
mle_h0 = .5
mle_h1 = .5 # 50% of flips were heads
likelihood_h0 = likelihood(mle_h0, n_heads+n_tails, n_heads)
print(f'likelihood_h0 = {likelihood_h0:.8f}')
likelihood_h1 = likelihood(mle_h1, n_heads+n_tails, n_heads)
print(f'likelihood_h1 = {likelihood_h1:.8f}')
print(f'Likelihood Ratio = {likelihood_h1 / likelihood_h0:.2f}')
print(f'p = {1 - chi2.cdf(-2 * (likelihood_h1 / likelihood_h0), 1):.4f}')
```
The result of this likelihood-ratio test is pretty trivial, but confirms the expectation described above. By treating each hypothesis as synomymous with the corresponding maximum likelihood estimate of theta, the two hypotheses end up being identical when we observe heads on 50% of our flips.
### Take home message
Bayes factors are fine, but I would almost never recommend them. They are useful for performing NHST-style "tests" in a Bayesian framework, but the idea that you have to **choose** between two or more hypotheses is something that (I think) researchers never should have been doing in the first place.
This is particularly true in the case of these point-estimate, "null"-style models. We either believe that the value of $\theta$ is *exactly* 0.50000000 or whether have no clue whatsoever what the value of $\theta$ is? It seems plausible that we don't actually believe one of these (e.g., as is the case in most NHST settings).
On top of that, Bayes factors do not reflect our prior beliefs about the credibility of the models we are comparing. Bayes factors speak to the "evidential value" of our data. But, as we saw above, the data can strongly imply one model and our priors can strongly favor the other. In such cases, Bayes factors only provide part of the relevant story.
What do I recommend? If you have data and a model and you would like to answer questions about the values of model parameters in light of the data, then **estimate** the credible values of those parameters and use the posterior to answer your questions. If you are not sure whether a coin is fair or not, build a model in which theta can take on many values of theta and ask how credible the values far from 0.5 are (for one or more definitions of "far"). Ask how credible values of theta close to 0.5 are (for one or more definitions of "close"). But dichotomizing the world seems unwise. Also the sampler doesn't like it.
| github_jupyter |
<a href="https://colab.research.google.com/github/rumen-cholakov/SemanticWeb/blob/master/grao_table_parser.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
## Imports
```
import pandas as pd
import requests
import pickle
import regex
import enum
import os
from typing import TypeVar, Callable, Sequence, List, Optional, Tuple
from collections import namedtuple
from bs4 import BeautifulSoup
from functools import reduce
```
## Type Declarations
```
class HeaderEnum(enum.Enum):
Old = 0
New = 1
class TableTypeEnum(enum.Enum):
Qarterly = 0
Yearly = 1
DataTuple = namedtuple('DataTuple', 'data header_type table_type')
MunicipalityIdentifier = namedtuple('MunicipalityIdentifier', 'region municipality')
SettlementInfo = namedtuple('SettlementInfo', 'name permanent_residents current_residents')
FullSettlementInfo = namedtuple('FullSettlementInfo', 'region municipality settlement permanent_residents current_residents')
PopulationInfo = namedtuple('PopulationInfo', 'permanent current')
ParsedLines = namedtuple('ParsedLines', 'municipality_ids settlements_info')
T = TypeVar('T')
```
## Definition of Data Source
```
data_source: List[DataTuple] = [
DataTuple("https://www.grao.bg/tna/t41nm-15-03-2020_2.txt", HeaderEnum.New, TableTypeEnum.Qarterly),
DataTuple("https://www.grao.bg/tna/tadr2019.txt", HeaderEnum.New, TableTypeEnum.Yearly),
DataTuple("https://www.grao.bg/tna/tadr2018.txt", HeaderEnum.New, TableTypeEnum.Yearly),
DataTuple("https://www.grao.bg/tna/tadr2017.txt", HeaderEnum.New, TableTypeEnum.Yearly),
DataTuple("https://www.grao.bg/tna/tadr-2016.txt", HeaderEnum.New, TableTypeEnum.Yearly),
DataTuple("https://www.grao.bg/tna/tadr-2015.txt", HeaderEnum.New, TableTypeEnum.Yearly),
DataTuple("https://www.grao.bg/tna/tadr-2014.txt", HeaderEnum.New, TableTypeEnum.Yearly),
DataTuple("https://www.grao.bg/tna/tadr-2013.txt", HeaderEnum.New, TableTypeEnum.Yearly),
DataTuple("https://www.grao.bg/tna/tadr-2012.txt", HeaderEnum.New, TableTypeEnum.Yearly),
DataTuple("https://www.grao.bg/tna/tadr-2011.txt", HeaderEnum.New, TableTypeEnum.Yearly),
DataTuple("https://www.grao.bg/tna/tadr-2010.txt", HeaderEnum.New, TableTypeEnum.Yearly),
DataTuple("https://www.grao.bg/tna/tadr-2009.txt", HeaderEnum.New, TableTypeEnum.Yearly),
DataTuple("https://www.grao.bg/tna/tadr-2008.txt", HeaderEnum.New, TableTypeEnum.Yearly),
DataTuple("https://www.grao.bg/tna/tadr-2007.txt", HeaderEnum.New, TableTypeEnum.Yearly),
DataTuple("https://www.grao.bg/tna/tadr-2006.txt", HeaderEnum.New, TableTypeEnum.Yearly),
DataTuple("https://www.grao.bg/tna/tadr-2005.txt", HeaderEnum.Old, TableTypeEnum.Yearly),
DataTuple("https://www.grao.bg/tna/tadr-2004.txt", HeaderEnum.Old, TableTypeEnum.Yearly),
DataTuple("https://www.grao.bg/tna/tadr-2003.txt", HeaderEnum.Old, TableTypeEnum.Yearly),
DataTuple("https://www.grao.bg/tna/tadr-2002.txt", HeaderEnum.Old, TableTypeEnum.Yearly),
DataTuple("https://www.grao.bg/tna/tadr-2001.txt", HeaderEnum.Old, TableTypeEnum.Yearly),
DataTuple("https://www.grao.bg/tna/tadr-2000.txt", HeaderEnum.Old, TableTypeEnum.Yearly),
DataTuple("https://www.grao.bg/tna/tadr-1999.txt", HeaderEnum.Old, TableTypeEnum.Yearly),
DataTuple("https://www.grao.bg/tna/tadr-1998.txt", HeaderEnum.Old, TableTypeEnum.Yearly),
]
```
## Regular Expresions Construction
```
# Building regex strings
cap_letter = '\p{Lu}'
low_letter = '\p{Ll}'
separator = '[\||\!]\s*'
number = '\d+'
year_group = '(\d{4})'
name_part = f'\s*{cap_letter}*'
name_part_old = f'\s{cap_letter}*'
type_abbr = f'{cap_letter}+\.'
name = f'{cap_letter}+{name_part * 3}'
name_old = f'{cap_letter}+{name_part_old * 3}'
word = f'{low_letter}+'
number_group = f'{separator}({number})\s*'
old_reg = f'ОБЛАСТ:({name_old})'
print(old_reg)
old_mun = f'ОБЩИНА:({name_old})'
print(old_mun)
region_name_new_re = f'{word} ({name}) {word} ({name})'
print(region_name_new_re)
# Quaterly
settlement_info_quarterly_re = f'({type_abbr}{name})\s*{number_group * 3}'
print(settlement_info_quarterly_re)
# Yearly
settlement_info_yearly_re = f'({type_abbr}{name})\s*{number_group * 6}'
print(settlement_info_yearly_re)
```
## Helper Functions
```
def pipeline(
value: T,
function_pipeline: Sequence[Callable[[T], T]],
) -> T:
'''A generic Unix-like pipeline
:param value: the value you want to pass through a pipeline
:param function_pipeline: an ordered list of functions that
comprise your pipeline
'''
return reduce(lambda v, f: f(v), function_pipeline, value)
def build_pipline(functions: Sequence[Callable[[T], T]]) -> Callable[[T], T]:
return (lambda value: pipeline(value, function_pipeline=functions))
def execute_pipeline(value: T, pipeline: Callable[[T], T]) -> T:
return pipeline(value)
def static_vars_funktion(**kwargs):
def decorate(func):
for k in kwargs:
setattr(func, k, kwargs[k])
return func
return decorate
```
## Parsing Pipeline Definitions
```
def fetch_raw_table(data_tuple: DataTuple) -> DataTuple:
headers = requests.utils.default_headers()
headers.update({
'User-Agent': 'Mozilla/5.0 (X11; Ubuntu; Linux x86_64; rv:52.0) Gecko/20100101 Firefox/52.0'
})
url = data_tuple.data
req = requests.get(url, headers)
req.encoding = 'windows-1251'
return DataTuple(req, data_tuple.header_type, data_tuple.table_type)
def raw_table_to_lines(data_tuple: DataTuple) -> DataTuple:
req = data_tuple.data
soup = BeautifulSoup(req.text, 'lxml').prettify()
split = soup.split('\r\n')
return DataTuple(split, data_tuple.header_type, data_tuple.table_type)
def parse_lines(data_tuple: DataTuple) -> DataTuple:
def parse_data_line(line: str, table_type: TableTypeEnum) -> Optional[SettlementInfo]:
settlement_info_re = ''
permanent_population_position = -1
current_population_position = -1
if table_type == TableTypeEnum.Qarterly:
settlement_info_re = settlement_info_quarterly_re
permanent_population_position = 2
current_population_position = 3
elif table_type == TableTypeEnum.Yearly:
settlement_info_re = settlement_info_yearly_re
permanent_population_position = 2
current_population_position = 6
settlement_info = regex.search(settlement_info_re, line)
if settlement_info:
name, permanent, current = settlement_info.group(1,
permanent_population_position,
current_population_position)
settlement_info = SettlementInfo(name.strip(), permanent, current)
return settlement_info
@static_vars_funktion(region=None)
def parse_header_line(line: str, header_type: HeaderEnum) -> Optional[MunicipalityIdentifier]:
region_name = None
if header_type == HeaderEnum.New:
region_name_re = region_name_new_re
region_gr = regex.search(region_name_re, line)
if region_gr:
region, municipality = region_gr.group(1, 2)
region_name = MunicipalityIdentifier(region.strip(), municipality.strip())
elif header_type == HeaderEnum.Old:
if not parse_header_line.region:
parse_header_line.region = regex.search(old_reg, line)
region_name = None
else:
mun_gr = regex.search(old_mun, line)
if mun_gr:
region, municipality = parse_header_line.region.group(1), mun_gr.group(1)
region_name = MunicipalityIdentifier(region.strip(), municipality.strip())
parse_header_line.region = None
return region_name
municipality_ids = {}
settlements_info = {}
for line_num, line in enumerate(data_tuple.data):
municipality_id = parse_header_line(line, data_tuple.header_type)
if municipality_id:
municipality_ids[line_num] = municipality_id
settlement_info = parse_data_line(line, data_tuple.table_type)
if settlement_info:
settlements_info[line_num] = settlement_info
return DataTuple(ParsedLines(municipality_ids, settlements_info), data_tuple.header_type, data_tuple.table_type)
def parssed_lines_to_full_info_list(data_tuple: DataTuple) -> DataTuple:
regions = data_tuple.data.municipality_ids
settlements_info = data_tuple.data.settlements_info
reg_keys = list(regions.keys())
settlement_keys = list(settlements_info.keys())
reg_keys_pairs = zip(reg_keys[:-1], reg_keys[1:])
sk_index = 0
full_name_settlement_infos = []
for current_mun, next_mun in reg_keys_pairs:
while current_mun < settlement_keys[sk_index] < next_mun:
reg = regions[current_mun]
set_info = settlements_info[settlement_keys[sk_index]]
fnsi = FullSettlementInfo(reg.region,
reg.municipality,
set_info.name,
set_info.permanent_residents,
set_info.current_residents)
full_name_settlement_infos.append(fnsi)
sk_index += 1
return DataTuple(full_name_settlement_infos, data_tuple.header_type, data_tuple.table_type)
def full_info_list_to_data_frame(data_tuple: DataTuple) -> DataTuple:
df = pd.DataFrame(data_tuple.data)
df.set_index(['region', 'municipality', 'settlement'], drop=True, inplace=True)
return DataTuple(df, data_tuple.header_type, data_tuple.table_type)
parsing_pipeline = build_pipline(functions=(
fetch_raw_table,
raw_table_to_lines,
parse_lines,
parssed_lines_to_full_info_list,
full_info_list_to_data_frame
))
```
## Data Processing Pipeline
```
def process_data(data_source: List[DataTuple]) -> List[DataTuple]:
parsed_data = None
data_frame_list = []
for data_tuple in data_source:
year = regex.search(year_group, data_tuple.data).group(1)
data_frame = execute_pipeline(data_tuple, parsing_pipeline).data
data_frame = data_frame.rename(columns={'permanent_residents':f'permanent_{year}',
'current_residents':f'current_{year}'})
data_frame_list.append(data_frame)
if isinstance(parsed_data, pd.DataFrame):
parsed_data = parsed_data.merge(data_frame, sort=False, how='right', left_index=True, right_index=True)
else:
parsed_data = data_frame
return [DataTuple(parsed_data,0,0), DataTuple(data_frame_list,0,0)]
def store_data(processed_data: List[DataTuple]) -> List[DataTuple]:
directory = './grao'
if not os.path.exists(directory):
os.makedirs(directory)
combined_data = processed_data[0].data
combined_data.to_csv(f'{directory}/combined_data.csv')
combined_data.to_pickle(f'{directory}/combined_data.pkl')
data_list = processed_data[1].data
with open(f'{directory}/data_frames_list.pkl', 'wb') as f:
pickle.dump(data_list, f)
return processed_data
processing_pipeline = build_pipline(functions=(
process_data,
store_data
))
```
## Data Processing
```
processed_data = execute_pipeline(data_source, processing_pipeline)
processed_data
!grep "ГОЦЕ ДЕЛЧЕВ" ./grao/combined_data.csv
```
stanislaw trailov
simeon
| github_jupyter |
# Sex differences in Autism Spectrum Disorder, a Comorbidity Pattern Analysis in National Scale Data: (v) PubMed publications for the Phecodes of interest
To check if the PheCodes that we have found have been previous reported in the literature, we will conduct a PubMed (https://www.ncbi.nlm.nih.gov/pubmed/)query for each PheCode. The steps followed are:
1. Map from PheCode to MESH term (https://www.ncbi.nlm.nih.gov/mesh) through UMLS.
3. Generate a PubMed query for each PheCode
### From PheCodes to MESH
From our final results, we previously saved a data.frame that contains the Phecode and the description.
Then, to map the PheCodes to MESH:
- Map from PheCode to ICD9-CM (https://phewascatalog.org/files/phecode_icd9_map_unrolled.csv.zip)
- Query MRCONSO (UMLS) to extract the MESH terms for each ICD9-CM
- Query PubMed to extract the number of publications for each phenotype.
```
#load the Phecodes of interest
phecodesForStudy <- read.delim( "phenosForPubmedCheck.txt" )
#load the mapping file (PheCode to ICD9-CM)
phemapFile <- read.csv( "phecode_icd9_rolled.csv" )
phemapFile <- phemapFile[ , c( "ICD9", "PheCode" ) ]
#we generate the SQL query to extract the MESH terms for each ICD9-CM code
for( i in 1:length( phecodesForStudy$description) ){
icd9Selection <- as.character(phemapFile[ phemapFile$PheCode == phecodesForStudy$phecode[i], "ICD9"])
myquery <- "SELECT unique CUI, CODE, STR
FROM UMLS.MRCONSO mrc
WHERE SAB = 'MSH' AND CUI IN ( SELECT CUI FROM umls.mrconsO WHERE SAB LIKE 'ICD9CM' AND CODE IN ("
for( j in 1:length( icd9Selection ) ){
if(j != length( icd9Selection )){
myquery <- paste0( myquery, "'", icd9Selection[j], "',")
}else{
myquery <- paste0( myquery, "'", icd9Selection[j], "'));")
}
}
print(paste0( phecodesForStudy$description[i], "*******", myquery ) )
}
```
As a result we will generate a SQL query for each one of the PheCodes.
For example, the query for Austism Spectrum Disorder will be:
```
# SELECT unique CUI, CODE, STR
# FROM
# UMLS.MRCONSO mrc
# WHERE
# SAB = 'MSH' AND
# CUI IN
# (
# SELECT CUI FROM umls.mrconsO
# WHERE SAB LIKE 'ICD9CM' AND
# CODE IN ('299.0', '299.00','299.01', '299.8', '299.80', '299.81', '299.9', '299.90', '299.91')
# );
```
### PubMed queries
When executing this and the rest of SQL queries generated, we will create a file for each phenotype of interest, that will contain a list of mapping MESH terms. Using those files as input, we will create and run SQL queries to PubMed using the R package **'rentrez'**.
#### Install and load R libraries
```
install.packages( "rentrez" )
library( "rentrez" )
#select the dbs that we will use, in this case, pubmed
entrez_dbs()
entrez_db_searchable("pubmed")
#define the path were the files with the MESH terms for the PheCodes of interest are located
pth <- "./"
```
#### Extract publications supporting the ASD co-occurrence with each one of the phenotypes.
For all the PubMed queries we will have a common part that will contain:
- The autism MESH terms
- The date range of publication
- The organism, in our case we are interested in Human research
- The type of publications
Then, for each phenotype we will have a variable part of the query, that will contain the list of each MESH terms associated to each phenotype.
```
#load the ASD MESH terms
autismMesh <- read.delim( "autism.dsv", header = TRUE, sep = "\t" )
autismMesh <- as.character( unique( tolower(autismMesh$STR ) ) )
#define the fixed part of the query
commonPart <- "humans[MeSH Terms] AND (\"2009\"[Date - Publication] : \"3000\"[Date - Publication]))
AND ( Classical Article[Publication Type] OR Clinical Study[Publication Type] OR
Comparative Study[Publication Type] OR Randomized Controlled Trial[Publication Type] OR
Observational Study[Publication Type] OR Journal Article[Publication Type]) AND ("
#add the list of ASD MESH terms to the query
for( i in 1:length( autismMesh ) ){
if( i == 1){
queryFirstPart <- paste0( commonPart, autismMesh[i], "[MeSH Terms] " )
}
if( i != 1 & i != length( autismMesh ) ){
queryFirstPart <- paste0( queryFirstPart, "OR ", autismMesh[i], "[MeSH Terms] " )
}
if( i == length( autismMesh ) ){
queryFirstPart <- paste0( queryFirstPart, "OR ", autismMesh[i], "[MeSH Terms] ) " )
}
}
#define the path where the rest of the mapping files are located
myfiles <- list.files( pth )
#generate an empty data frame to fill with the results
mydfPublicationsASD <- as.data.frame( matrix( ncol = 5, nrow = length( myfiles ) ) )
colnames( mydfPublicationsASD ) <- c("phenotype",
"publications",
"MeSH term mapped to the phenotype",
"query",
"timeQuery" )
#for each phenotype, complete the query with the specific MESH terms
for( cont in 1:length( myfiles ) ){
print( cont )
phenoMesh <- read.delim(paste0( pth, myfiles[ cont ]), header = TRUE, sep = "\t" )
if( colnames( phenoMesh )[3] == "STR" ){
phenoMesh <- as.character( unique( tolower(phenoMesh$STR ) ) )
if( length( phenoMesh ) == 0 ){
mydfPublicationsASD[cont,] <- c( myfiles[ cont ], " ", length( phenoMesh ), " ", " ")
next()
}else {
for( i in 1:length( phenoMesh ) ){
if( length( phenoMesh ) != 1 ){
if( i == 1){
querySecondPart <- paste0( " AND (", phenoMesh[i], "[MeSH Terms] " )
}
if( i != 1 & i != length( phenoMesh ) ){
querySecondPart <- paste0( querySecondPart, "OR ", phenoMesh[i], "[MeSH Terms] " )
}
if( i == length( phenoMesh ) ){
querySecondPart <- paste0( querySecondPart, "OR ", phenoMesh[i], "[MeSH Terms] ) " )
}
}
else{
querySecondPart <- paste0( " AND (", phenoMesh[i], "[MeSH Terms] )" )
}
}
}
finalQuery <- paste0( queryFirstPart, querySecondPart )
#sometimes when the query is too long we can get some errors, so we will print the query to run it directly in
#the PubMed web page
#otherwise we run the query from R applying the function "entrez_search"
if( length( phenoMesh ) > 17 ){
mydfPublicationsASD[cont,] <- c( myfiles[ cont ],
"TooLarge",
length( phenoMesh ),
finalQuery,
as.character(Sys.time() ) )
print("###########################")
print( finalQuery )
print("###########################")
}else{
r_search <- entrez_search(db="pubmed", finalQuery, retmax=100, use_history=TRUE )
r_search
mydfPublicationsASD[cont,] <- c( myfiles[ cont ],
r_search$count,
length( phenoMesh ),
finalQuery,
as.character(Sys.time() ) )
}
}
else{
print( myfiles[ cont ] )
}
}
colnames( mydfPublicationsASD ) <- c( "phenotype", "publicationsASD",
"MESHmapped", "queryASD", "timeQueryASD" )
```
#### Extract publications supporting the ASD co-occurrence with each one of the phenotypes and sex differences.
For all the PubMed queries we will have a common part that will contain:
- The autism MESH terms
- The date range of publication
- The organism, in our case we are interested in Human research
- The type of publications
- The MESH terms that define "SEX DIFFERENCES"
Then, for each phenotype we will have a variable part of the query, that will contain the list of each MESH terms associated to each phenotype.
```
#load the ASD MESH terms
autismMesh <- read.delim( "autism.dsv", header = TRUE, sep = "\t" )
autismMesh <- as.character( unique( tolower(autismMesh$STR ) ) )
#define the fixed part of the query
commonPart <- "humans[MeSH Terms] AND (\"2009\"[Date - Publication] : \"3000\"[Date - Publication]))
AND (sex difference[MeSH Terms] OR Sex Factors[MeSH Terms] OR Sex[MeSH Terms] OR
Sex Characteristics[MeSH Terms]) AND ( Classical Article[Publication Type] OR
Clinical Study[Publication Type] OR Comparative Study[Publication Type] OR
Randomized Controlled Trial[Publication Type] OR Observational Study[Publication Type] OR
Journal Article[Publication Type]) AND ("
#add the list of ASD MESH terms to the query
for( i in 1:length( autismMesh ) ){
if( i == 1){
queryFirstPart <- paste0( commonPart, autismMesh[i], "[MeSH Terms] " )
}
if( i != 1 & i != length( autismMesh ) ){
queryFirstPart <- paste0( queryFirstPart, "OR ", autismMesh[i], "[MeSH Terms] " )
}
if( i == length( autismMesh ) ){
queryFirstPart <- paste0( queryFirstPart, "OR ", autismMesh[i], "[MeSH Terms] ) " )
}
}
#define the path where the rest of the mapping files are located
myfiles <- list.files( pth )
#generate an empty data frame to fill with the results
mydfPublicationsSexDiff <- as.data.frame( matrix( ncol = 5, nrow = length( myfiles ) ) )
colnames( mydfPublicationsSexDiff ) <- c("phenotype",
"publications",
"MeSH term mapped to the phenotype",
"query",
"timeQuery" )
#for each phenotype, complete the query with the specific MESH terms
for( cont in 1:length( myfiles ) ){
print( cont )
phenoMesh <- read.delim(paste0( pth, myfiles[ cont ]), header = TRUE, sep = "\t" )
if( colnames( phenoMesh )[3] == "STR" ){
phenoMesh <- as.character( unique( tolower(phenoMesh$STR ) ) )
if( length( phenoMesh ) == 0 ){
mydfPublicationsSexDiff[cont,] <- c( myfiles[ cont ], " ", length( phenoMesh ), " ", " ")
next()
}else {
for( i in 1:length( phenoMesh ) ){
if( length( phenoMesh ) != 1 ){
if( i == 1){
querySecondPart <- paste0( " AND (", phenoMesh[i], "[MeSH Terms] " )
}
if( i != 1 & i != length( phenoMesh ) ){
querySecondPart <- paste0( querySecondPart, "OR ", phenoMesh[i], "[MeSH Terms] " )
}
if( i == length( phenoMesh ) ){
querySecondPart <- paste0( querySecondPart, "OR ", phenoMesh[i], "[MeSH Terms] ) " )
}
}
else{
querySecondPart <- paste0( " AND (", phenoMesh[i], "[MeSH Terms] )" )
}
}
}
finalQuery <- paste0( queryFirstPart, querySecondPart )
#sometimes when the query is too long we can get some errors, so we will print the query to run it directly in
#the PubMed web page
#otherwise we run the query from R applying the function "entrez_search"
if( length( phenoMesh ) > 17 ){
mydfPublicationsSexDiff[cont,] <- c( myfiles[ cont ],
"TooLarge",
length( phenoMesh ),
finalQuery,
as.character(Sys.time() ) )
print("###########################")
print( finalQuery )
print("###########################")
}else{
r_search <- entrez_search(db="pubmed", finalQuery, retmax=100, use_history=TRUE )
r_search
mydfPublicationsSexDiff[cont,] <- c( myfiles[ cont ],
r_search$count,
length( phenoMesh ),
finalQuery,
as.character(Sys.time() ) )
}
}
else{
print( myfiles[ cont ] )
}
}
colnames( mydfPublicationsSexDiff) <- c("phenotype", "publicationsSexDiff",
"MESHmapped", "querySexDiff", "timeQuerySexDiff" )
```
#### Combine both results in one table
As a result we generate a table *(Supplementary table 3)* that contains for each Phenotype the number publications supporting ASD co-occurrence, and the number of publications supporting ASD co-occurrence and sex differences.
Additionally we will also save the date of the publication, the number of MESH terms that mapped to each PheCode and the specific PubMed Query.
And example PubMed query would be:
*humans[MeSH Terms] AND ("2009"[Date - Publication] : "3000"[Date - Publication])) AND ( Classical Article[Publication Type] OR Clinical Study[Publication Type] OR Comparative Study[Publication Type] OR Randomized Controlled Trial[Publication Type] OR Observational Study[Publication Type] OR Journal Article[Publication Type]) AND (infantile autism, early[MeSH Terms] OR autism[MeSH Terms] OR autism, infantile[MeSH Terms] OR autistic disorder[MeSH Terms] OR autism, early infantile[MeSH Terms] OR kanners syndrome[MeSH Terms] OR pervasive development disorders[MeSH Terms] OR early infantile autism[MeSH Terms] OR infantile autism[MeSH Terms] OR disorders, autistic[MeSH Terms] OR disorder, autistic[MeSH Terms] OR kanner's syndrome[MeSH Terms] OR kanner syndrome[MeSH Terms] ) AND (astigmatism[MeSH Terms])*
```
finalPublicationTable <- merge( mydfPublicationsASD, mydfPublicationsSexDiff, by="phenotype" )
write.table( finalPublicationTable,
file = "pubmedCheckOutput.txt",
col.names = TRUE,
row.names = FALSE,
quote = FALSE,
sep = "\t" )
```
| github_jupyter |
# Contribute
Before we can accept contributions, you need to become a CLAed contributor.
E-mail a signed copy of the
[CLAI](https://github.com/openpifpaf/openpifpaf/blob/main/docs/CLAI.txt)
(and if applicable the
[CLAC](https://github.com/openpifpaf/openpifpaf/blob/main/docs/CLAC.txt))
as PDF file to research@svenkreiss.com.
(modify-code)=
## Modify Code
For development of the openpifpaf source code itself, you need to clone this repository and then:
```sh
pip3 install numpy cython
pip3 install --editable '.[dev,train,test]'
```
The last command installs the Python package in the current directory
(signified by the dot) with the optional dependencies needed for training and
testing. If you modify `functional.pyx`, run this last command again which
recompiles the static code.
Develop your features in separate feature branches.
Create a pull request with your suggested changes. Make sure your code passes
`pytest`, `pylint` and `pycodestyle` checks:
```sh
pylint openpifpaf
pycodestyle openpifpaf
pytest
cd guide
python download_data.py
pytest --nbval-lax --current-env *.ipynb
```
## Things to Contribute
This is a research project and changing fast. Contributions can be in many areas:
* Add a new dataset?
* Add a new base network?
* Try a different loss?
* Try a better multi-task strategy?
* Try a different head architecture?
* Add a new task?
* Run on new hardware (mobile phones, embedded devices, ...)?
* Improve training schedule/procedure?
* Use it to build an app?
* Improve documentation (!!)
* ...
## Missing Dependencies
OpenPifPaf has few core requirements so that you can run it efficiently on servers without graphical interface.
Sometimes, you just want to install all possible dependencies. Those are provided as "extra" requirements.
Use the following `pip3` command to install all extras.
```
# NO CODE
import sys
if sys.version_info >= (3, 8):
import importlib.metadata
extras = importlib.metadata.metadata('openpifpaf').get_all('Provides-Extra')
print(f'pip3 install "openpifpaf[{",".join(extras)}]"')
```
## Your Project and OpenPifPaf
Let us know about your open source projects. We would like to feature them in our "related projects" section.
The simplest way to integrate with OpenPifPaf is to write a plugin. If some functionality is not possible through our plugin architecture, open an issue to discuss and if necessary send us a pull request that enables the missing feature you need.
If you do need to make a copy of OpenPifPaf, you must respect our license.
## Build Guide
```sh
cd guide
jb build .
```
If you encounter issues with the kernel spec in a notebook, open the notebook
with a text editor and find `metadata.kernelspec.name` and set it to `python3`.
Alternatively, you can patch your local package yourself. Open
`venv/lib/python3.9/site-packages/jupyter_cache/executors/utils.py`
in your editor and add `kernel_name='python3'` to the arguments of `nbexecute()`
[here](https://github.com/executablebooks/jupyter-cache/blob/1431ed72961fabc2f09a13553b0aa45f4c8a7c23/jupyter_cache/executors/utils.py#L56).
Alternatively, for continuous integration, the `kernel_name` is replace in the json of the
Jupyter Notebook before executing jupyter-book
[here](https://github.com/openpifpaf/openpifpaf/blob/6db797205cd082bf7a80e967372957e56c9835fb/.github/workflows/deploy-guide.yml#L47).
Only use this operation on a discardable copy as `jq` changes all formatting.
## Build Environment
```
%%bash
pip freeze
%%bash
python -m openpifpaf.predict --version
```
| github_jupyter |
# Transfer Learning
In this notebook, you'll learn how to use pre-trained networks to solved challenging problems in computer vision. Specifically, you'll use networks trained on [ImageNet](http://www.image-net.org/) [available from torchvision](http://pytorch.org/docs/0.3.0/torchvision/models.html).
ImageNet is a massive dataset with over 1 million labeled images in 1000 categories. It's used to train deep neural networks using an architecture called convolutional layers. I'm not going to get into the details of convolutional networks here, but if you want to learn more about them, please [watch this](https://www.youtube.com/watch?v=2-Ol7ZB0MmU).
Once trained, these models work astonishingly well as feature detectors for images they weren't trained on. Using a pre-trained network on images not in the training set is called transfer learning. Here we'll use transfer learning to train a network that can classify our cat and dog photos with near perfect accuracy.
With `torchvision.models` you can download these pre-trained networks and use them in your applications. We'll include `models` in our imports now.
```
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
import matplotlib.pyplot as plt
import torch
from torch import nn
from torch import optim
import torch.nn.functional as F
from torchvision import datasets, transforms, models
```
Most of the pretrained models require the input to be 224x224 images. Also, we'll need to match the normalization used when the models were trained. Each color channel was normalized separately, the means are `[0.485, 0.456, 0.406]` and the standard deviations are `[0.229, 0.224, 0.225]`.
```
data_dir = 'Cat_Dog_data'
# TODO: Define transforms for the training data and testing data
train_transforms = transforms.Compose([
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406],
[0.229, 0.224, 0.225])])
test_transforms = transforms.Compose([
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406],
[0.229, 0.224, 0.225])])
# Pass transforms in here, then run the next cell to see how the transforms look
train_data = datasets.ImageFolder(data_dir + '/train', transform=train_transforms)
test_data = datasets.ImageFolder(data_dir + '/test', transform=test_transforms)
trainloader = torch.utils.data.DataLoader(train_data, batch_size=64, shuffle=True)
testloader = torch.utils.data.DataLoader(test_data, batch_size=32)
```
We can load in a model such as [DenseNet](http://pytorch.org/docs/0.3.0/torchvision/models.html#id5). Let's print out the model architecture so we can see what's going on.
```
model = models.densenet121(pretrained=True)
model
```
This model is built out of two main parts, the features and the classifier. The features part is a stack of convolutional layers and overall works as a feature detector that can be fed into a classifier. The classifier part is a single fully-connected layer `(classifier): Linear(in_features=1024, out_features=1000)`. This layer was trained on the ImageNet dataset, so it won't work for our specific problem. That means we need to replace the classifier, but the features will work perfectly on their own. In general, I think about pre-trained networks as amazingly good feature detectors that can be used as the input for simple feed-forward classifiers.
```
# Freeze parameters so we don't backprop through them
for param in model.parameters():
param.requires_grad = False
from collections import OrderedDict
classifier = nn.Sequential(OrderedDict([
('fc1', nn.Linear(1024, 500)),
('relu', nn.ReLU()),
('fc2', nn.Linear(500, 2)),
('output', nn.LogSoftmax(dim=1))
]))
model.classifier = classifier
```
With our model built, we need to train the classifier. However, now we're using a **really deep** neural network. If you try to train this on a CPU like normal, it will take a long, long time. Instead, we're going to use the GPU to do the calculations. The linear algebra computations are done in parallel on the GPU leading to 100x increased training speeds. It's also possible to train on multiple GPUs, further decreasing training time.
PyTorch, along with pretty much every other deep learning framework, uses [CUDA](https://developer.nvidia.com/cuda-zone) to efficiently compute the forward and backwards passes on the GPU. In PyTorch, you move your model parameters and other tensors to the GPU memory using `model.to('cuda')`. You can move them back from the GPU with `model.to('cpu')` which you'll commonly do when you need to operate on the network output outside of PyTorch. As a demonstration of the increased speed, I'll compare how long it takes to perform a forward and backward pass with and without a GPU.
```
import time
#for device in ['cpu', 'cuda']:
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
criterion = nn.NLLLoss()
# Only train the classifier parameters, feature parameters are frozen
optimizer = optim.Adam(model.classifier.parameters(), lr=0.001)
model.to(device)
for ii, (inputs, labels) in enumerate(trainloader):
# Move input and label tensors to the GPU
inputs, labels = inputs.to(device), labels.to(device)
start = time.time()
outputs = model.forward(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
if ii==3:
break
print(f"Device = {device}; Time per batch: {(time.time() - start)/3:.3f} seconds")
# Test out your network!
import helper
model.eval()
dataiter = iter(testloader)
images, labels = dataiter.next()
img = images[0]
print(images[0].size())
helper.imshow(img)
# Convert 2D image to 1D vector
#img = img.resize_(3, 224*224)
img = img.view(1, 3*224*224)
# Calculate the class probabilities (softmax) for img
with torch.no_grad():
output = model.forward(img)
ps = torch.exp(output)
# Plot the image and probabilities
helper.view_classify(img.view(3, 224, 224), ps, version='Fashion')
```
You can write device agnostic code which will automatically use CUDA if it's enabled like so:
```python
# at beginning of the script
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
...
# then whenever you get a new Tensor or Module
# this won't copy if they are already on the desired device
input = data.to(device)
model = MyModule(...).to(device)
```
From here, I'll let you finish training the model. The process is the same as before except now your model is much more powerful. You should get better than 95% accuracy easily.
>**Exercise:** Train a pretrained models to classify the cat and dog images. Continue with the DenseNet model, or try ResNet, it's also a good model to try out first. Make sure you are only training the classifier and the parameters for the features part are frozen.
```
# TODO: Train a model with a pre-trained network
```
| github_jupyter |
## Data Leakage, L1(Lasso) and L2 (Ridge) regularization using Linear Regression
We will use cross validation, lasso and ridge regression in this lab.
Specifically speaking, <br>
Regularization basically adds the penalty as model complexity increases.<br>
Cross validation is used to evaluate how well our model can generalize on the dataset. <br>
We will be using r2 score in this lab. It provides an indication of goodness of fit and therefore a measure of how well unseen samples are likely to be predicted by the model.
In this task, we will explore the following things on linear regression model:
- Cross Validation
- L1 regularization (Lasso regression)
- L2 regularization (Ridge regression)
#### Dataset
The dataset is available at "data/bike.csv" in the respective challenge's repo.<br>
The dataset is __modified version__ of the dataset 'bike.csv' provided by UCI Machine Learning repository.
Original dataset: https://archive.ics.uci.edu/ml/datasets/bike+sharing+dataset
#### Objective
To learn about how cross validation, L1 regularization and L2 regularization work.
#### Tasks
- load the dataset.
- perform pre-processing on the data.
- remove registered feature and keep the casual feature to understand data leakage.
- construct train and test dataset.
- create a linear regression model.
- check the r2 score of the initial linear regression model on train and test dataset
- observe distribution of weights in the initial linear regression model.
- split the dataset into k consecutive folds.
- calculate cross validation score for the k fold and check how well our model can generalize on the training dataset.
- checking the variance threshold of dataset and remove features with low variance.
- apply L1 regularization on the dataset and check the r2_score.
- visualize the distribution of weights on the lasso regression model.
- apply L2 regularization on the dataset and check the r2_score.
- visualize the distribution of weights on the ridge regression model.
#### Further fun
- apply RFE on the dataset to automatically remove uneccessary features which would prevent overfitting.
- don't remove casual and registered features and check the effect of data leakage on the model
- implement lasso and ridge regression without using inbuilt librarires.
- apply elastic net to visualize the effect of both ridge and lasso regression.
#### Helpful links
- Cross validation : https://machinelearningmastery.com/k-fold-cross-validation/#:~:text=Cross%2Dvalidation%20is%20a%20resampling,k%2Dfold%20cross%2Dvalidation.
- Cross validation: https://scikit-learn.org/stable/modules/cross_validation.html
- L1 and L2 regularization : https://towardsdatascience.com/ridge-and-lasso-regression-a-complete-guide-with-python-scikit-learn-e20e34bcbf0b
- L1 and L2 regularization : https://www.youtube.com/watch?v=9lRv01HDU0s&list=PLZoTAELRMXVPBTrWtJkn3wWQxZkmTXGwe&index=30&t=904s
- r2_score: https://scikit-learn.org/stable/modules/generated/sklearn.metrics.r2_score.html#sklearn.metrics.r2_score
- pd.get_dummies() and One Hot Encoding: https://queirozf.com/entries/one-hot-encoding-a-feature-on-a-pandas-dataframe-an-example
- Data Leakage : "https://machinelearningmastery.com/data-leakage-machine-learning/
- sklearn k-fold : https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.KFold.html
- sklearn cross_val_score : https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.cross_val_score.html?highlight=cross_val_score#sklearn.model_selection.cross_val_score
- sklearn lasso regression : https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.Lasso.html?highlight=lasso#sklearn.linear_model.Lasso
- sklearn ridge regression : https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.Ridge.html?highlight=ridge#sklearn.linear_model.Ridge
- RFE : https://machinelearningmastery.com/rfe-feature-selection-in-python/
- RFE sklearn : https://scikit-learn.org/stable/modules/generated/sklearn.feature_selection.RFE.html
- Use slack for doubts:
```
#import the necessary libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# Sklearn processing
from sklearn.preprocessing import MinMaxScaler
from sklearn.model_selection import train_test_split
from sklearn.model_selection import KFold
from sklearn.model_selection import cross_val_score
# Sklearn linear regression model
from sklearn.linear_model import LinearRegression
# Sklearn regression model evaluation functions
from sklearn.metrics import r2_score
# Perform feature selection using a variance threshold
from sklearn.feature_selection import VarianceThreshold
# Feature selection using Recursive Feature Elimimation
from sklearn.feature_selection import RFE
#load the data and inspect the first 5 rows
!wget https://github.com/DeepConnectAI/challenge-week-6/raw/master/data/bike.csv
data = pd.read_csv('./bike.csv')
data.head()
# print the data types of each feature name
data.dtypes
data.columns
# check for null values in each column
data.isnull().sum()
# print out the unique values of the features ['season', 'year', 'weather', 'promotion_type']
print(data['season'].unique())
print(data['year'].unique())
print(data['weather'].unique())
print(data['promotion_type'].unique())
cols = data.columns
for i in cols:
print(i)
print(data[i].unique())
# print out the value counts (frequency of occurence) of the unique values in these features ['season', 'year', 'weather', 'promotion_type']
cols = ['season', 'year', 'weather', 'promotion_type']
for i in cols:
print(i+': ')
print(data[i].value_counts())
print()
# print the shape of data
data.shape
# drop the feature 'id' as it has no information to deliver.
data = data.drop('id', axis = 1)
data = data.drop('year',axis = 1)
# print the shape of data
data.shape
# one hot encode the categorical columns.
categorical_columns = ['season', 'weather']
for i in categorical_columns:
x = pd.get_dummies(data[i], prefix = i)
data = pd.concat([data,x],axis = 1)
data = data.drop(i,axis = 1)
# print the shape of data
# notice the increase in the no. of features
data.shape
```
Notice that our target feature "cnt" is the sum of the features "registered" + "casual"<br>
To avoid data leakage remove the feature "casual" for the training purpose. <br>
To understand more about data leakage refer the article mentioned in the uselful links.
```
# Split the dataset into X and y
# While loading data into X drop the columns "cnt" and "casual".
X_cols = ['holiday', 'weekday', 'workingday', 'temp', 'feel_temp', 'hum',
'windspeed', 'promotion_level', 'promotion_type',
'promotion_level_external', 'promotion_type_external','registered','season_autumn', 'season_spring', 'season_summer',
'season_winter', 'weather_cloud', 'weather_fair', 'weather_rain']
X = data[X_cols]
# notice the target variable is 'cnt'
y = data['cnt'].values
print(X.shape,y.shape)
# store the names of the training features / name of the columns used for training. [Very important step for visualization later.]
train_columns = list(X.columns)
print(train_columns)
# Apply scaling if our data is spread across wide differences of range values.
X.head
# print the type of X
# num = ['holiday', 'weekday', 'workingday', 'temp', 'feel_temp', 'hum', 'windspeed', 'promotion_level', 'promotion_type', 'promotion_level_external', 'promotion_type_external', 'registered', 'season_autumn', 'season_spring', 'season_summer', 'season_winter', 'weather_cloud', 'weather_fair', 'weather_rain']
# for i in num:
# data[i] = (data[i] - data[i].min())/(data[i].max() - data[i].min())
minmax=MinMaxScaler()
X=minmax.fit_transform(X)
#data=pd.DataFrame(data_N,columns=data.columns)
# minmax=MinMaxScaler()
# data_Nm=minmax.fit_transform(data['registered'].values.reshape(-1,1))
# data['registered']=pd.DataFrame(data_Nm,columns=['registered'])
```
Note : <br>
Type of X should be pandas dataframe.
If not then convert X into pandas DataFrame object before proceeding further.
```
# convert X into pandas Dataframe
# in the parameters specify columns = train_columns.
X = pd.DataFrame(X, columns = train_columns)
X.head()
# split the dataset into X_train, X_test, y_train, y_test
# play around with test sizes.
test_size = 0.2
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = test_size, random_state=42)
# print the shapes
print(X_train.shape)
print(X_test.shape)
print(y_train.shape)
print(y_test.shape)
# build the Linear Regression model.
model = LinearRegression()
# fit the model on the training data
model.fit(X_train, y_train)
# print the score on training set
y_pred_train = model.predict(X_train)
print("On Training set : ", r2_score(y_train, y_pred_train))
# print the score on the test set
y_pred_test = model.predict(X_test)
print("On testing set : ", r2_score(y_test,y_pred_test))
```
Do not edit the code given below. Observe the distribution of weights.
Which feature has the maximum coefficient ? <br>
Keep this figure as a base reference for visualizing the effects of l1-norm and l2-norm later in this notebook.
```
# custom summary function to plot the coefficients / weightage of the features.
def custom_summary(model, column_names, title):
'''Show a summary of the trained linear regression model'''
# Plot the coeffients as bars
fig = plt.figure(figsize=(8,len(column_names)/3))
fig.suptitle(title, fontsize=16)
rects = plt.barh(column_names, model.coef_,color="lightblue")
# Annotate the bars with the coefficient values
for rect in rects:
width = round(rect.get_width(),4)
plt.gca().annotate(' {} '.format(width),
xy=(0, rect.get_y()),
xytext=(0,2),
textcoords="offset points",
ha='left' if width<0 else 'right', va='bottom')
plt.show()
# coefficients plot
# let's call the above custom function.
custom_summary(model, train_columns, "Linear Regression coefficients.")
# evaluate the model with k = 10 Fold Cross validation
folds = KFold(n_splits = 10, shuffle = True, random_state = 100)
results = cross_val_score(model, X, y, scoring = 'r2', cv = folds)
print(type(model).__name__)
print("kFoldCV:")
print("Fold R2 scores:", results)
print("Mean R2 score:", results.mean())
print("Std R2 score:", results.std())
print("Generalizability on training set : ", results.mean(), " +/- ", results.std())
```
Feature Selection using Variance Thresholding
```
print("Original shape of X_train : ", X_train.shape)
# check the variance of X.
# Note the type(X) should be a pandas DataFrame as stated earlier.
X.var()
```
Remove low variance features using Variance Threshold.
Note : If the variance is less, it implies the values of that particular feature spans limited range of values.
```
# play around with the threshold values
sel = VarianceThreshold(threshold = (0.01))
sel.fit(X_train)
# do not edit.
selected_features = list(X_train.columns[sel.get_support()])
print("Selected features : ", selected_features)
print("Removed features : ", list(X_train.columns[~sel.get_support()]))
removed_features = list(X_train.columns[~sel.get_support()])
# Delete the removed features from the train_columns list.
for i in removed_features:
train_columns.remove(i)
#train_columns.append(i)
#transform / remove the low variance features
X_train = sel.transform(X_train)
X_test = sel.transform(X_test)
```
## Lasso Regression : L1 - norm
```
from sklearn.linear_model import Lasso
#X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2, random_state = 100)
# hyperparamater alpha : controls the degree of penaliation.
# play around with alpha values.
alpha = 1.0
#create the model
model_lasso = Lasso(alpha = alpha)
#fit the model on training data
model_lasso.fit(X_train, y_train)
#calculate the score on training data
y_pred_train = model_lasso.predict(X_train)
print("On train set : ", r2_score(y_train, y_pred_train))
#evaluate the model on testing data
y_pred_test = model_lasso.predict(X_test)
print("On test set : ", r2_score(y_test, y_pred_test))
# visualize the coefficients.
# compare the results with the plot obtained earlier.
custom_summary(model_lasso, train_columns, "Lasso Regression Coefficients.")
```
We can see that Lasso regression has automatically done a lot of feature selection. Some columns might have zero coefficients. It has been effectively removed. <br>
The model is much more interpretable than the baseline linear regression model.
<br>
Hence, Lasso regression has embedded Feature Selection.
# Ridge Regression : L2 - norm
```
from sklearn.linear_model import Ridge
# hyperparamater alpha : controls the degree of penaliation.
# play around with alpha values.
alpha = 1.0
#create the model
model_ridge = Ridge(alpha = alpha)
#fit the model on training data
model_ridge.fit(X_train, y_train)
#calculate the score on training data
y_pred_train = model_ridge.predict(X_train)
print("On train set : ", r2_score(y_train, y_pred_train))
#evaluate the model on testing data
y_pred_test = model_ridge.predict(X_test)
print("On test set : ", r2_score(y_test, y_pred_test))
# visualize the coefficients.
# compare the results with the plot obtained earlier.
custom_summary(model_ridge, train_columns, "Ridge Regression Coefficients.")
```
Ridge regression doesn't drive smaller coefficients to 0 hence it doesn't possess internal feature selection.
Points to Ponder ! [Optional]
Did you notice the highest dependency on the feature "registered" if you haven't removed it till now ?
Since our target is "cnt" which is the simple combination of "registered" and "casual".
we have removed "casual", but the model was smart enough to predict the target "cnt" simply from one feature "registered" itself.
This is the classic example of Data Leakage. So the aim here is not to make 99 percent accurate predictions, the aim is to take into account the factors for making predictions.
So, to get a detailed report, we should avoid data leakage thereby removing both the features "registered" and "casual".
```
```
| github_jupyter |
```
import os
import sys
import json
import pathlib
cwd = pathlib.Path.cwd()
sys.path.insert(0, str(cwd.parent.joinpath("turk", "analysis")))
from analysis import parse_csv
from collections import defaultdict, OrderedDict
import numpy as np
import seaborn as sns
from matplotlib import pyplot as plt
np.random.seed(12)
def group_by(res_lines, variable_name):
to_ret = defaultdict(list)
for line in res_lines:
interest_key = line[variable_name]
to_ret[interest_key].append(line)
return to_ret
def sort_by_fxn(res_dict, fxn):
new_dict = OrderedDict()
# average = lambda x: sum(x)/len(x)
to_sort = []
for key, values in res_dict.items():
ps = [line['p_true'] for line in values]
score = fxn(ps)
to_sort.append((score, key, values))
to_sort = sorted(to_sort, key = lambda x: x[0])
for __, key, values in to_sort:
new_dict[key] = values
return new_dict
def add_model_data(path, data):
with open(path) as f1:
model_data = json.load(f1)
model_data_dict = {data["questionId"]: data for data in model_data}
for i, item in enumerate(data):
qid = item["question_id"]
model_pred = model_data_dict[qid]['prediction']
model_p_yes = model_pred["yes"]
model_p_no = model_pred["no"]
assert(model_p_yes + model_p_no > 0.98)
model_p_yes *= 100
data[i]["model_p_true"] = model_p_yes
return data
results_path = cwd.parent.parent.joinpath("results")
lxmert_path = results_path.joinpath("lxmert", "gqa")
turk_path = results_path.joinpath("turk", "gqa")
# all_data_paths
all_data = {}
for csv in lxmert_path.glob("*.json"):
filename = csv.stem
unit = filename.split("_")[1]
turk_name = f"clean_and_norm_{unit}.csv"
try:
res_lines = parse_csv(turk_path.joinpath(turk_name))
res_lines = add_model_data(csv, res_lines)
except FileNotFoundError:
print(f"error with {filename}")
continue
by_sent = group_by(res_lines, "question_id")
by_sent_sorted_avg = sort_by_fxn(by_sent, lambda x: np.mean(x))
# by_sent_sorted_std_dev = sort_by_fxn(by_sent, lambda x: np.std(x))
all_data[filename] = by_sent_sorted_avg
figure_path = "/Users/Elias//vagueness-2020/figures/"
# by_sent = group_by(res_lines, "question_id")
# by_sent_sorted_avg = sort_by_fxn(by_sent, lambda x: np.mean(x))
# by_sent_sorted_std_dev = sort_by_fxn(by_sent, lambda x: np.std(x))
from scipy.optimize import curve_fit
np.random.seed(12)
def sigmoid(x, x0, k):
return 1.0 / (1.0 + np.exp(-k * (x-x0)))
def get_rmse(pred_ys, true_ys):
return np.sqrt(np.sum((pred_ys - true_ys)**2) / len(pred_ys))
def fit_sigmoid(data):
xs = np.arange(len(data.keys()))
human_ys = [np.mean([line['p_true'] for line in data[key] ] ) for key in data.keys() ]
human_ys = np.array(human_ys)/100
bounds=([0,len(xs)+2], [0.01, 2])
popt, pcov = curve_fit(sigmoid, xs, human_ys, p0= [20, 0.1], method='dogbox')
pred_ys = np.array([sigmoid(x, popt[0], popt[1]) for x in xs])
# use mean squared error
rmse_pred_to_human = get_rmse(pred_ys, human_ys)
model_ys = [np.mean([line['model_p_true'] for line in data[key] ] ) for key in data.keys()]
model_ys = np.array(model_ys)/100
rmse_pred_to_model = get_rmse(pred_ys, model_ys)
random_ys = np.random.uniform(size=len(human_ys))
rmse_pred_to_random = get_rmse(pred_ys, random_ys)
return rmse_pred_to_human, rmse_pred_to_model, rmse_pred_to_random, popt
import os
from analysis import parse_csv
from collections import defaultdict, OrderedDict
import seaborn as sns
import pandas as pd
import matplotlib
from matplotlib import pyplot as plt
font = {'family' : 'Times New Roman',
'weight' : 'normal',
'size' : 20}
matplotlib.rc('font', **font)
np.random.seed(12)
def plot_data(data, title_text, sigmoid_params = None, of_interest = None, do_save = False):
xs = np.arange(len(data.keys()))
human_ys = [np.mean([line['p_true'] for line in data[key] ] ) for key in data.keys() ]
human_std_dev = np.array([np.std([line['p_true'] for line in data[key] ] ) for key in data.keys() ])
human_std_err = human_std_dev / np.sqrt(10) # np.sqrt(len(human_ys))
# all the same value so mean = any of them
model_ys = [np.mean([line['model_p_true'] for line in data[key]] ) for key in data.keys()]
sents = [line[0]['sent'] for line in data.values()]
qids = [line[0]['question_id'] for line in data.values()]
all_lines = [lines[0] for lines in data.values() ]
true_colors = ["True" if np.mean([line['p_true'] for line in data[key] ]) > 50 else "False" for key in data.keys() ]
palette = {"True": '#4575b4' ,
"False": '#e34a33'}
markers = {"True": "." ,
"False": "."}
plt.figure()
fig, axs = plt.subplots(2, 1, sharex='col', sharey=True, figsize=(6,6))
axs[0].set_title(title_text)
axs[0].fill_between(xs, human_ys - human_std_err, human_ys + human_std_err, color=(0,0,0,0.15))
for x, human_y, model_y, c, s, qid in zip(xs, human_ys, model_ys, true_colors, sents, qids):
color = c
axs[0].scatter([x], [human_y], s = 150, marker = markers[c], color = palette[color])
axs[1].scatter([x], [model_y], s = 150, marker = markers[c], color = palette[color])
# iterate again so rings are on top
for x, human_y, model_y, c, s, qid in zip(xs, human_ys, model_ys, true_colors, sents, qids):
if of_interest is not None and qid in of_interest:
axs[0].scatter([x], [human_y], s = 400, marker = ".", color = (0,0,0))
axs[0].scatter([x], [human_y], s = 100, marker = ".", color = palette[c])
axs[1].scatter([x], [model_y], s = 400, marker = ".", color = (0,0,0))
axs[1].scatter([x], [model_y], s = 100, marker = ".", color = palette[c])
if sigmoid_params is not None:
sigmoid_xs = xs
sigmoid_ys = np.array([sigmoid(x, *sigmoid_params) for x in sigmoid_xs]) * 100
axs[0].plot(sigmoid_xs, sigmoid_ys, "-", color = (0,0,0), linewidth=2)
axs[1].plot(sigmoid_xs, sigmoid_ys, "-", color = (0,0,0), linewidth=2)
for i in range(2):
axs[i].set_yticks([0.0, 100])
axs[i].set_yticklabels([0.0, 100])
axs[i].set_xticks([])
axs[i].set_xticklabels([])
axs[i].spines['right'].set_visible(False)
axs[i].spines['top'].set_visible(False)
axs[i].spines['bottom'].set_visible(False)
# axs.spines['left'].set_visible(False)
axs[i].spines['bottom'].set_visible(True)
axs[0].set_ylabel("Human", rotation = 90)
axs[1].set_ylabel("LXMERT", rotation = 90)
plt.tight_layout()
if do_save:
plt.savefig(os.path.join(figure_path, f"{title_text}_gqa.pdf"))
return plt
from scipy.optimize import curve_fit
np.random.seed(12)
def sigmoid(x, x0, k):
return 1.0 / (1.0 + np.exp(-k * (x-x0)))
def get_rmse(pred_ys, true_ys):
return np.sqrt(np.sum((pred_ys - true_ys)**2) / len(pred_ys))
def fit_sigmoid(data, folds = 10):
xs = np.arange(len(data.keys()))
all_human_ys = [np.mean([line['p_true'] for line in data[key] ] ) for key in data.keys() ]
all_human_ys = np.array(all_human_ys)/100
all_model_ys = [np.mean([line['model_p_true'] for line in data[key] ] ) for key in data.keys()]
all_model_ys = np.array(all_model_ys)/100
bounds=([0,len(xs)+2], [0.01, 2])
all_human_rmse, all_model_rmse, all_random_rmse = [], [], []
all_popts = []
# shuffle data
zipped = list(zip(xs, all_human_ys, all_model_ys))
np.random.seed(12)
np.random.shuffle(zipped)
xs, all_human_ys, all_model_ys = zip(*zipped)
idxs = [i for i in range(len(xs))]
for fold in range(folds):
train_xs, train_human_ys, dev_human_ys, train_model_ys, dev_model_ys = None, None, None, None, None
start = int(fold/folds * len(xs))
end = int((fold+1)/folds * len(xs))
dev_idxs = idxs[start:end]
train_idxs = [idx for idx in idxs if idx not in dev_idxs]
dev_xs = np.array([xs[idx] for idx in dev_idxs])
train_xs = np.array([xs[idx] for idx in train_idxs])
train_human_ys = np.array([all_human_ys[idx] for idx in train_idxs])
dev_human_ys = np.array([all_human_ys[idx] for idx in dev_idxs])
train_model_ys = np.array([all_model_ys[idx] for idx in train_idxs])
dev_model_ys = np.array([all_model_ys[idx] for idx in dev_idxs])
midpoint = len(xs)/2
popt_human, pcov_human = curve_fit(sigmoid,
train_xs,
train_human_ys,
p0 = [midpoint, 0.1],
method='dogbox')
pred_ys_human = np.array([sigmoid(x, popt_human[0], popt_human[1]) for x in dev_xs])
rmse_pred_to_human = get_rmse(pred_ys_human, dev_human_ys)
all_human_rmse.append(rmse_pred_to_human)
all_popts.append(popt_human)
for param in [0.0001, 0.001, 0.01, 0.1, 1, 10]:
try:
popt_model, pcov_model = curve_fit(sigmoid,
train_xs,
train_model_ys,
p0 = [midpoint, param],
method='dogbox')
pred_ys_model = np.array([sigmoid(x, popt_model[0], popt_model[1]) for x in dev_xs])
rmse_pred_to_model = get_rmse(pred_ys_model, dev_model_ys)
all_model_rmse.append(rmse_pred_to_model)
break
except RuntimeError:
continue
train_random_ys = np.random.uniform(size=len(train_human_ys))
dev_random_ys = np.random.uniform(size=len(dev_human_ys))
for param in [0.0001, 0.001, 0.01, 0.1, 1, 10]:
try:
popt_random, pcov_random = curve_fit(sigmoid,
train_xs,
train_random_ys,
p0 = [midpoint, 1],
method='dogbox')
pred_ys_random = np.array([sigmoid(x, popt_random[0], popt_random[1]) for x in dev_xs])
rmse_pred_to_random = get_rmse(pred_ys_random, dev_random_ys)
all_random_rmse.append(rmse_pred_to_random)
break
except RuntimeError:
continue
return (np.mean(all_human_rmse),
np.mean(all_model_rmse),
np.mean(all_random_rmse),
np.mean(np.array(all_popts)))
fits = {}
data_for_latex = {}
for name, data in all_data.items():
human_rmse, model_rmse, random_rmse, popt = fit_sigmoid(data, folds = 10)
fits[name] = (human_rmse, model_rmse, random_rmse, popt)
human_rmse = np.format_float_positional(human_rmse, 3)
model_rmse = np.format_float_positional(model_rmse, 3)
random_rmse = np.format_float_positional(random_rmse, 3)
data_for_latex[name] = [human_rmse, model_rmse,random_rmse]
# print(f"{name} human rmse: {human_rmse}, model rmse: {model_rmse}, random rsme: {random_rmse}")
HEADER = """\\begin{tabular}{llll}
\\hline
Predicate & Human & Model & Random \\\\
\\hline
"""
body = ""
for name in ["output_sunny_yesno_small", "output_cloudy_yesno_small", "output_new_yesno_small", "output_old_yesno_small", "output_young_yesno_small", "output_adult_yesno_small"]:
row_data = data_for_latex[name]
just_name = name.split("_")[1]
row = " & ".join([just_name] + row_data) + "\\\\"
row += "\n"
body+=row
FOOTER = """\\end{tabular}"""
print(HEADER + body + FOOTER)
# filenames = ["output_sunny_yesno_small", "output_cloudy_yesno_small",
# "output_new_yesno_small","output_old_yesno_small",
# "output_young_yesno_small", "output_adult_yesno_small"]
# names = ["sunny","cloudy","new","old", "young","adult"]
# for filename, name in zip(filenames,names):
# plot_data(all_data[filename], name)
filenames = ["output_sunny_yesno_small", "output_cloudy_yesno_small",
"output_new_yesno_small","output_old_yesno_small",
"output_young_yesno_small", "output_adult_yesno_small"]
names = ["sunny","cloudy","new","old", "young","adult"]
of_interest = ['03959852', '18812880', '18590951']
for filename, name in zip(filenames,names):
# plot_data(all_data[filename], name, sigmoid_params = fits[filename][-1], of_interest = of_interest)
plot_data(all_data[filename], name, sigmoid_params = None, of_interest = of_interest, do_save = True)
# example search
filenames = ["output_sunny_yesno_small"] #, "output_cloudy_yesno_small"]
names = ["sunny"] #,"cloudy"]
for filename, name in zip(filenames,names):
data = all_data[filename]
keys = list(data.keys())
# print(filename, len(data))
print("NOT VAGUE NO")
for i in range(0, 3):
print(data[keys[i]][0])
print("==============================")
print("VAGUE")
midpoint = int(len(data)/2)
for i in range(midpoint-5, midpoint+5):
print(data[keys[i]][0])
print("==============================")
print("NOT VAGUE YES")
for i in range(-3, -1):
print(data[keys[i]][0])
print("==============================")
```
| github_jupyter |
# Demonstration of PET OSEM reconstruction with SIRF
This demonstration shows how to use OSEM as implemented in SIRF. It also suggests some exercises for reconstruction with and without attenuation etc.
The notebook is currently set-up to use prepared data with a single slice of an XCAT phantom, with a low resolution scanner, such that all results can be obtained easily on a laptop. Of course, the code will work exactly the same for any sized data.
Authors: Kris Thielemans and Evgueni Ovtchinnikov
First version: June 2021
CCP SyneRBI Synergistic Image Reconstruction Framework (SIRF).
Copyright 2015 - 2018, 2021 University College London.
This is software developed for the Collaborative Computational Project in Synergistic Reconstruction for Biomedical Imaging (http://www.ccpsynerbi.ac.uk/).
Licensed under the Apache License, Version 2.0 (the "License");
you may not use this file except in compliance with the License.
You may obtain a copy of the License at
http://www.apache.org/licenses/LICENSE-2.0
Unless required by applicable law or agreed to in writing, software
distributed under the License is distributed on an "AS IS" BASIS,
WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
See the License for the specific language governing permissions and
limitations under the License.
# What is OSEM?
The following is just a very brief explanation of the concepts behind OSEM.
PET reconstruction is commonly based on the *Maximum Likelihood Estimation (MLE)* principle. The *likelihood* is the probability to observe some measured data given a (known) image. MLE attempts to find the image that maximises this likelihood. This needs to be done iteratively as the system of equations is very non-linear.
A common iterative method uses *Expectation Maximisation*, which we will not explain here. The resulting algorithm is called *MLEM* (or sometimes *EMML*). However, it is rather slow. The most popular method to increase computation speed is to compute every image update based on only a subset of the data. Subsets are nearly always chosen in terms of the "views" (or azimuthal angles). The *Ordered Subsets Expectation Maximisation (OSEM)* cycles through the subsets. More on this in another notebook, but here we just show how to use the SIRF implementation of OSEM.
OSEM is (still) the most common algorithm in use in clinical PET.
# Initial set-up
```
#%% make sure figures appears inline and animations works
%matplotlib notebook
# Setup the working directory for the notebook
import notebook_setup
from sirf_exercises import cd_to_working_dir
cd_to_working_dir('PET', 'OSEM_reconstruction')
#%% Initial imports etc
import numpy
from numpy.linalg import norm
import matplotlib.pyplot as plt
import matplotlib.animation as animation
import os
import sys
import shutil
#import scipy
#from scipy import optimize
import sirf.STIR as pet
from sirf.Utilities import examples_data_path
from sirf_exercises import exercises_data_path
# define the directory with input files for this notebook
data_path = os.path.join(examples_data_path('PET'), 'thorax_single_slice')
#%% our usual handy function definitions
def plot_2d_image(idx,vol,title,clims=None,cmap="viridis"):
"""Customized version of subplot to plot 2D image"""
plt.subplot(*idx)
plt.imshow(vol,cmap=cmap)
if not clims is None:
plt.clim(clims)
plt.colorbar(shrink=.6)
plt.title(title)
plt.axis("off")
```
## We will first create some simulated data from ground-truth images
see previous notebooks for more information.
```
#%% Read in images
image = pet.ImageData(os.path.join(data_path, 'emission.hv'))
attn_image = pet.ImageData(os.path.join(data_path, 'attenuation.hv'))
#%% display
im_slice = image.dimensions()[0]//2
plt.figure(figsize=(9, 4))
plot_2d_image([1,2,1],image.as_array()[im_slice,:,:,], 'emission image')
plot_2d_image([1,2,2],attn_image.as_array()[im_slice,:,:,], 'attenuation image')
plt.tight_layout()
#%% save max for future displays
cmax = image.max()*.6
#%% create acquisition model
acq_model = pet.AcquisitionModelUsingRayTracingMatrix()
template = pet.AcquisitionData(os.path.join(data_path, 'template_sinogram.hs'))
acq_model.set_up(template, image)
#%% simulate data using forward projection
acquired_data=acq_model.forward(image)
#%% Display bitmaps of a middle sinogram
acquired_data.show(im_slice,title='Forward projection')
```
# Reconstruction via a SIRF reconstruction class
While you can write your own reconstruction algorithm by using `AcquisitionModel` etc (see other notebooks), SIRF provides a few reconstruction clases. We show how to use the OSEM implementation here.
## step 1: create the objective function
In PET, the iterative algorithms in SIRF rely on an objective function (i.e. the function to maximise).
In PET, this is normally the Poisson log-likelihood. (We will see later about adding prior information).
```
obj_fun = pet.make_Poisson_loglikelihood(acquired_data)
```
We could set acquisition model but the default (ray-tracing) is in this case ok. See below for more information. You could do this as follows.
```
obj_fun.set_acquisition_model(acq_model)
```
We could also add a prior, but we will not do that here (although the rest of the exercise would still work).
## step 2: create OSMAPOSL reconstructor
The `sirf.STIR.OSMAPOSLReconstructor` class implements the *Ordered Subsets Maximum A-Posteriori One Step Late algorithm*. That's quite a mouthful! We will get round to the "OSL" part, which is used to incorporate prior information. However, without a prior, this algorithm is identical to *Ordered Subsets Expectation Maximisation* (OSEM).
```
recon = pet.OSMAPOSLReconstructor()
recon.set_objective_function(obj_fun)
# use 4 subset and 60 image updates. This is not too far from clinical practice.
recon.set_num_subsets(4)
num_subiters=60
recon.set_num_subiterations(num_subiters)
```
## step 3: use this reconstructor!
We first create an initial image. Passing this image automatically gives the dimensions of the output image.
It is common practice to initialise OSEM with a uniform image. Here we use a value which is roughly of the correct scale, although this value doesn't matter too much (see discussion in the OSEM_DIY notebook).
Then we need to set-up the reconstructor. That will do various checks and initial computations.
And then finally we call the `reconstruct` method.
```
#initialisation
initial_image=image.get_uniform_copy(cmax / 4)
recon.set_current_estimate(initial_image)
# set up the reconstructor
recon.set_up(initial_image)
# do actual recon
recon.process()
reconstructed_image=recon.get_output()
```
display of images
```
plt.figure(figsize=(9, 4))
plot_2d_image([1,2,1],image.as_array()[im_slice,:,:,],'ground truth image',[0,cmax*1.2])
plot_2d_image([1,2,2],reconstructed_image.as_array()[im_slice,:,:,],'reconstructed image',[0,cmax*1.2])
plt.tight_layout();
```
## step 4: write to file
You can ask the `OSMAPOSLReconstructor` to write images to file every few sub-iterations, but this is by default disabled. We can however write the image to file from SIRF.
For each "engine" its default file format is used, which for STIR is Interfile.
```
reconstructed_image.write('OSEM_result.hv')
```
You can also use the `write_par` member to specify a STIR parameter file to write in a different file format, but this is out of scope for this exercise.
# Including a more realistic acquisition model
The above steps were appropriate for an acquisition without attenuation etc. This is of course not appropriate for measured data.
Let us use some things we've learned from the [image_creation_and_simulation notebook](image_creation_and_simulation.ipynb). First thing is to create a new acquisition model, then we need to use it to simulate new data, and finally to use it for the reconstruction.
```
# create attenuation
acq_model_for_attn = pet.AcquisitionModelUsingRayTracingMatrix()
asm_attn = pet.AcquisitionSensitivityModel(attn_image, acq_model_for_attn)
asm_attn.set_up(template)
attn_factors = asm_attn.forward(template.get_uniform_copy(1))
asm_attn = pet.AcquisitionSensitivityModel(attn_factors)
# create acquisition model
acq_model = pet.AcquisitionModelUsingRayTracingMatrix()
# we will increase the number of rays used for every Line-of-Response (LOR) as an example
# (it is not required for the exercise of course)
acq_model.set_num_tangential_LORs(5)
acq_model.set_acquisition_sensitivity(asm_attn)
# set-up
acq_model.set_up(template,image)
# simulate data
acquired_data = acq_model.forward(image)
# let's add a background term of a reasonable scale
background_term = acquired_data.get_uniform_copy(acquired_data.max()/10)
acq_model.set_background_term(background_term)
acquired_data = acq_model.forward(image)
# create reconstructor
obj_fun = pet.make_Poisson_loglikelihood(acquired_data)
obj_fun.set_acquisition_model(acq_model)
recon = pet.OSMAPOSLReconstructor()
recon.set_objective_function(obj_fun)
recon.set_num_subsets(4)
recon.set_num_subiterations(60)
# initialisation and reconstruction
recon.set_current_estimate(initial_image)
recon.set_up(initial_image)
recon.process()
reconstructed_image=recon.get_output()
# display
plt.figure(figsize=(9, 4))
plot_2d_image([1,2,1],image.as_array()[im_slice,:,:,],'ground truth image',[0,cmax*1.2])
plot_2d_image([1,2,2],reconstructed_image.as_array()[im_slice,:,:,],'reconstructed image',[0,cmax*1.2])
plt.tight_layout();
```
# Exercise: write a function to do an OSEM reconstruction
The above lines still are quite verbose. So, your task is now to create a function that includes these steps, such that you can avoid writing all those lines all over again.
For this, you need to know a bit about Python, but mostly you can copy-paste lines from above.
Let's make a function that creates an acquisition model, given some input. Then we can write OSEM function that does the reconstruction.
Below is a skeleton implementation. Look at the code above to fill in the details.
To debug your code, it might be helpful at any messages that STIR writes. By default these are written to the terminal, but this is not helpful when running in a jupyter notebook. The line below will redirect all messages to files which you can open via the `File>Open` menu.
```
msg_red = pet.MessageRedirector('info.txt', 'warnings.txt', 'errors.txt')
```
Note that they will be located in the current directory.
```
%pwd
def create_acq_model(attn_image, background_term):
'''create a PET acquisition model.
Arguments:
attn_image: the mu-map
background_term: bakcground-term as an sirf.STIR.AcquisitionData
'''
# acq_model_for_attn = ...
# asm_model = ...
acq_model = pet.AcquisitionModelUsingRayTracingMatrix();
# acq_model.set_...
return acq_model
def OSEM(acq_data, acq_model, initial_image, num_subiterations, num_subsets=1):
'''run OSEM
Arguments:
acq_data: the (measured) data
acq_model: the acquisition model
initial_image: used for initialisation (and sets voxel-sizes etc)
num_subiterations: number of sub-iterations (or image updates)
num_subsets: number of subsets (defaults to 1, i.e. MLEM)
'''
#obj_fun = ...
#obj_fun.set...
recon = pet.OSMAPOSLReconstructor()
#recon.set_objective_function(...)
#recon.set...
recon.set_current_estimate(initial_image)
recon.set_up(initial_image)
recon.process()
return recon.get_output()
```
Now test it with the above data.
```
acq_model = create_acq_model(attn_image, background_term)
my_reconstructed_image = OSEM(acquired_data, acq_model, image.get_uniform_copy(cmax), 30, 4)
```
# Exercise: reconstruct with and without attenuation
In some cases, it can be useful to reconstruct the emission data without taking attenuation (or even background terms) into account. One common example is to align the attenuation image to the emission image.
It is easy to do such an *no--attenuation-correction (NAC)* recontruction in SIRF. You need to create an `AcquisitionModel` that does not include the attenuation factors, and use that for the reconstruction. (Of course, in a simulation context, you would still use the full model to do the simulation).
Implement that here and reconstruct the data with and without attenuation to see visually what the difference is in the reconstructed images. If you have completed the previous exercise, you can use your own functions to do this.
Hint: the easiest way would be to take the existing attenuation image, and use `get_uniform_copy(0)` to create an image where all $\mu$-values are 0. Another (and more efficient) way would be to avoid creating the `AcquisitionSensitivityModel` at all.
# Final remarks
In these exercises we have used attenuation images of the same size as the emission image. This is easier to code and for display etc, but it is not a requirement of SIRF.
In addition, we have simulated and reconstructed the data with the same `AcquisitionModel` (and preserved image sizes). This is also convenient, but not a requirement (as you've seen in the NAC exercise). In fact, do not write your next paper using this "inverse crime". The problem is too
| github_jupyter |
```
import matplotlib
matplotlib.use('Qt5Agg')
import matplotlib.pyplot as plt
from src.utils import *
from src.InstrumentalVariable import InstrumentalVariable
from tqdm.notebook import tnrange
def run_manifold_tests(X, y, min_p_value=80, max_p_value=95, bootstrap=True, min_l2_reg=0,
max_l2_reg=50, n_tests=100):
n_samples, n_features, _ = X.shape
experiment_coefs = np.zeros((n_tests, n_features))
for i in tnrange(n_tests):
p_value = np.random.uniform(min_p_value, max_p_value)
if max_l2_reg > 0:
l2_reg = np.random.uniform(min_l2_reg, max_l2_reg)
else:
l2_reg = None
iv_model = InstrumentalVariable(p_value, l2_reg)
feature_size = np.random.randint(8, 20)
feature_inds = np.random.choice(n_features, feature_size, replace=False)
if bootstrap:
bootstrap_inds = np.random.choice(len(X), len(X))
X_train, y_train = X[bootstrap_inds], y[bootstrap_inds]
else:
X_train = X
y_train = y
X_train = X_train[:, feature_inds]
iv_model.fit(X_train, y_train)
np.put(experiment_coefs[i], feature_inds, iv_model.coef_)
return experiment_coefs
def filter_metrics(coefs):
positive_coefs = np.apply_along_axis(lambda feature: len(np.where(feature > 0)[0]), 0, coefs)
negative_coefs = np.apply_along_axis(lambda feature: len(np.where(feature < 0)[0]), 0, coefs)
print(positive_coefs)
print(negative_coefs)
filtered_coef_inds = []
for i, feature_coefs in enumerate(coefs.T):
pos = positive_coefs[i]
neg = negative_coefs[i]
if pos + neg == 0:
continue
if pos == 0 or neg == 0 or min(pos/neg, neg/pos) < 0.2:
filtered_coef_inds.append(i)
return np.array(filtered_coef_inds)
def plot_coefficients(coefs, metric_map=None):
n_tests, n_features = coefs.shape
fig, axes = plt.subplots(nrows=n_features, sharex=True)
fig.suptitle("3_metric_stability")
collections = []
for i, metric_coefs in enumerate(coefs.T):
ax = axes[i]
ax.set_title('Weights for short_term_' + str(metric_map[i]), loc='left')
ax.plot([0, 0], [-1, 1], 'r')
metric_coefs = metric_coefs[metric_coefs != 0]
n_tests = len(metric_coefs)
col = ax.scatter(metric_coefs, np.random.rand(n_tests) * 2 - 1,
cmap=plt.get_cmap("RdBu"), picker=5, s=50)
collections.append(col)
plt.show()
short_metrics_p, long_metrics_p = read_data(dataset_name='feed_top_ab_tests_pool_big_dataset.csv' ,shift=True)
short_metrics = short_metrics_p[:, :, 0]
long_metrics = long_metrics_p[:, :, 0]
target_metric_p = long_metrics_p[:, 3, :] # <--- here you can choose target (0, 1, 2, 3)
target_metric = target_metric_p[:, 0]
#main part of the sandbox, as it allows to change the constraints
coefs = run_manifold_tests(short_metrics_p, target_metric,
min_l2_reg=0, max_l2_reg=0.001,
min_p_value=50, max_p_value=95, n_tests=1000)
clear_metrics = filter_metrics(coefs)
filtered_coefs = coefs[:, clear_metrics]
plot_coefficients(coefs, range(np.shape(coefs)[1]))
```
| github_jupyter |
# Time series visualization by `Pandas`
*April 2, 2021 10:53*
## 1. Single time series
```
import tushare as ts
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
# display all rows
pd.set_option('expand_frame_repr', False)
url = "https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_deaths_global.csv"
country = 'Canada'
df = pd.read_csv(url, delimiter=',', )
df_ts = pd.DataFrame(df[df['Country/Region'] == f'{country}'].sum(axis=0, numeric_only=True,), columns=[f'num_death_in_{country}'],).iloc[2:]
df_ts.index = pd.to_datetime(df_ts.index)
df_ts.tail()
ax = df_ts.plot(color = 'tan')
ax.set_xlabel('Pandemic Time')
ax.set_ylabel(f'No.of Death in {country}')
plt.show()
data_filepath = 'C:\\Users\\dengj/ubc_mds_21/ds_block6/574_spat_temp_mod/DSCI_574_lab1_dengjr/data/'
beach_df = pd.read_csv(data_filepath + 'beach_train.csv', delimiter=',', index_col=['Time'], parse_dates=True)
beach_df.head()
beach_df = beach_df.interpolate()
beach_df.info()
ax = beach_df.plot(color = 'tan')
ax.set_xlabel('Time')
ax.set_ylabel(f'Shoreline(m)')
plt.show()
# see the pre-set plot style
styles = (plt.style.available)
probs=np.random.dirichlet(np.ones(len(styles)),size=1).flatten()
style = np.random.choice(styles, p=probs)
plt.style.use(f'{style}')
ax1 = beach_df.plot()
ax1.set_title(f'{style}')
ax1.set_ylabel('Shoreline(m)')
plt.show()
```
## 2. Put more details
> The graph above is a simple line plot, but more details could be added by using the parameters in `plot()`. This will help the graph being more aesthetic and clear by putting up details.
- `figsize(width, height)` for the figure size
- `linewidth` for the size of the lines
- `fontsize` for the font size
- `set_title()` to set the title
```
ax = beach_df.plot(color='tan', figsize=(15, 3), linewidth=2, fontsize=11)
ax.set_title('The record of shoreline(m) over years', fontsize=22)
plt.show()
```
> If you want to see the trend in any sub-time range, you could index such time range and then plotting
```
df_subset = beach_df['1982-05-25':'1996-02-18']
ax = df_subset.plot(color='tan', fontsize=11, figsize=(15, 3))
plt.show()
```
> If you want to emphasize particular observation, you could use `.axvline()` and `.axhline()` to add vertical and horizontal lines.
```
beach_df.index[111:444];
probs=np.random.dirichlet(np.ones(len(styles)),size=1).flatten()
style = np.random.choice(styles, p=probs)
plt.style.use(f'{style}')
ax = beach_df.plot(color='tan', fontsize=9)
ax.set_title(f'{style}')
ax.axvline('1996-02-11', color='red', linestyle='--')
ax.axhline(111.11, color='green', linestyle=':')
plt.show()
```
> `axvspan()` will allow you to add shade for particular areas, where `alpha` is for transparent rate (0 is totally transparent and 1 is fully colored)
```
import matplotlib.colors as mcolors
colors = list(mcolors.CSS4_COLORS.keys())
beach_df.max()
probs=np.random.dirichlet(np.ones(len(styles)),size=1).flatten()
probs2=np.random.dirichlet(np.ones(len(colors)),size=1).flatten()
style = np.random.choice(styles, p=probs)
linecolor = np.random.choice(colors, p=probs2)
shadecolor1 = colors[int(np.random.randint(len(colors), size=1))]
shadecolor2 = colors[int(np.random.randint(len(colors), size=1))]
plt.style.use(f'{style}')
ax = beach_df.plot(color=f'{linecolor}', fontsize=9)
ax.set_title(f'{style} w/ line color = {linecolor}, shade1 = {shadecolor1}, shade2={shadecolor2}')
ax.axvspan('1994-04-19', '2000-11-11', color=f'{shadecolor1}', alpha=.5)
n1=np.random.randint(144, size=1)
n2=np.random.randint(n1, 144, size=1)
ax.axhspan(int(n1), int(n2), color=f'{shadecolor2}', alpha=.88)
plt.show()
```
## 3. `rolling()`
> `rolling()` can help you move the observing window size.
```
w = 33
ma=beach_df.rolling(window=w).mean()
mstd = beach_df.rolling(window=w).std()
ma['upper'] = ma.Shoreline + mstd.Shoreline*2
ma['lower'] = ma.Shoreline - mstd.Shoreline*2
ax = ma.plot(linewidth=.9)
ax.set_xlabel('Time', fontsize=8)
ax.set_ylabel("Shoreline(m)")
ax.set_title(f'{style} w/ line color = {linecolor}, shade1 = {shadecolor1}, shade2={shadecolor2}')
plt.show()
```
## 4. Multiple time-series
```
min_=beach_df.Shoreline.min()
max_=beach_df.Shoreline.max()
beach_df['sync1'] = np.random.gamma(55, 4, 580)
beach_df['sync2'] = np.random.gamma(8, 5, 580)
beach_df
ax = beach_df.plot(linewidth=2, fontsize=12)
ax.set_xlabel('Time')
ax.legend(fontsize=9)
plt.show()
```
### `.plot.area()`
```
ax = beach_df.plot.area(fontsize=12)
ax.set_xlabel('Time')
ax.legend(fontsize=9)
plt.show()
```
### `subplot=True`, `layout=(nrow, ncol)`, `sharex=False/True`, `sharey=False/True`, `colormap='viridis'`
```
beach_df.plot(
figsize=(15, 5),
subplots=True,
layout=(1, 3),
sharex=False,
sharey=False,
colormap='viridis',
fontsize=8,
legend=False,
linewidth=1
)
plt.show()
```
## 5. Conclusion:
- [`matplotlib`](https://matplotlib.org/)
- [`matplotlib.pyplot`](https://matplotlib.org/stable/api/_as_gen/matplotlib.pyplot.html#module-matplotlib.pyplot)
- [datacamp](https://www.datacamp.com/courses/visualizing-time-series-data-in-python)
| github_jupyter |
# Linear algebra
Linear algebra is the branch of mathematics that deals with **vector spaces**.
```
import re, math, random # regexes, math functions, random numbers
import matplotlib.pyplot as plt # pyplot
from collections import defaultdict, Counter
from functools import partial, reduce
```
# Vectors
Vectors are points in some finite-dimensional space.
```
v = [1, 2]
w = [2, 1]
vectors = [v, w]
def vector_add(v, w):
"""adds two vectors componentwise"""
return [v_i + w_i for v_i, w_i in zip(v,w)]
vector_add(v, w)
def vector_subtract(v, w):
"""subtracts two vectors componentwise"""
return [v_i - w_i for v_i, w_i in zip(v,w)]
vector_subtract(v, w)
def vector_sum(vectors):
return reduce(vector_add, vectors)
vector_sum(vectors)
def scalar_multiply(c, v):
# c is a number, v is a vector
return [c * v_i for v_i in v]
scalar_multiply(2.5, v)
def vector_mean(vectors):
"""compute the vector whose i-th element is the mean of the
i-th elements of the input vectors"""
n = len(vectors)
return scalar_multiply(1/n, vector_sum(vectors))
vector_mean(vectors)
def dot(v, w):
"""v_1 * w_1 + ... + v_n * w_n"""
return sum(v_i * w_i for v_i, w_i in zip(v, w))
dot(v, w)
```
The dot product measures how far the vector v extends in the w direction.
- For example, if w = [1, 0] then dot(v, w) is just the first component of v.
The dot product measures the length of the vector you’d get if you projected v onto w.
```
def sum_of_squares(v):
"""v_1 * v_1 + ... + v_n * v_n"""
return dot(v, v)
sum_of_squares(v)
def magnitude(v):
return math.sqrt(sum_of_squares(v))
magnitude(v)
def squared_distance(v, w):
return sum_of_squares(vector_subtract(v, w))
squared_distance(v, w)
def distance(v, w):
return math.sqrt(squared_distance(v, w))
distance(v, w)
```
Using lists as vectors
- is great for exposition
- but terrible for performance.
- to use the NumPy library.
# Matrices
A matrix is a two-dimensional collection of numbers.
- We will represent matrices as lists of lists
- If A is a matrix, then A[i][j] is the element in the ith row and the jth column.
```
A = [[1, 2, 3],
[4, 5, 6]]
B = [[1, 2],
[3, 4],
[5, 6]]
def shape(A):
num_rows = len(A)
num_cols = len(A[0]) if A else 0
return num_rows, num_cols
shape(A)
def get_row(A, i):
return A[i]
get_row(A, 1)
def get_column(A, j):
return [A_i[j] for A_i in A]
get_column(A, 2)
def make_matrix(num_rows, num_cols, entry_fn):
"""returns a num_rows x num_cols matrix
whose (i,j)-th entry is entry_fn(i, j),
entry_fn is a function for generating matrix elements."""
return [[entry_fn(i, j)
for j in range(num_cols)]
for i in range(num_rows)]
def entry_add(i, j):
"""a function for generating matrix elements. """
return i+j
make_matrix(5, 5, entry_add)
def is_diagonal(i, j):
"""1's on the 'diagonal',
0's everywhere else"""
return 1 if i == j else 0
identity_matrix = make_matrix(5, 5, is_diagonal)
identity_matrix
```
### Matrices will be important.
- using a matrix to represent a dataset
- using an n × k matrix to represent a linear function that maps k-dimensional vectors to n-dimensional vectors.
- using matrix to represent binary relationships.
```
friendships = [(0, 1),
(0, 2),
(1, 2),
(1, 3),
(2, 3),
(3, 4),
(4, 5),
(5, 6),
(5, 7),
(6, 8),
(7, 8),
(8, 9)]
friendships = [[0, 1, 1, 0, 0, 0, 0, 0, 0, 0], # user 0
[1, 0, 1, 1, 0, 0, 0, 0, 0, 0], # user 1
[1, 1, 0, 1, 0, 0, 0, 0, 0, 0], # user 2
[0, 1, 1, 0, 1, 0, 0, 0, 0, 0], # user 3
[0, 0, 0, 1, 0, 1, 0, 0, 0, 0], # user 4
[0, 0, 0, 0, 1, 0, 1, 1, 0, 0], # user 5
[0, 0, 0, 0, 0, 1, 0, 0, 1, 0], # user 6
[0, 0, 0, 0, 0, 1, 0, 0, 1, 0], # user 7
[0, 0, 0, 0, 0, 0, 1, 1, 0, 1], # user 8
[0, 0, 0, 0, 0, 0, 0, 0, 1, 0]] # user 9
friendships[0][2] == 1 # True, 0 and 2 are friends
def matrix_add(A, B):
if shape(A) != shape(B):
raise ArithmeticError("cannot add matrices with different shapes")
num_rows, num_cols = shape(A)
def entry_fn(i, j): return A[i][j] + B[i][j]
return make_matrix(num_rows, num_cols, entry_fn)
A = make_matrix(5, 5, is_diagonal)
B = make_matrix(5, 5, entry_add)
matrix_add(A, B)
v = [2, 1]
w = [math.sqrt(.25), math.sqrt(.75)]
c = dot(v, w)
vonw = scalar_multiply(c, w)
o = [0,0]
plt.figure(figsize=(4, 5), dpi = 100)
plt.arrow(0, 0, v[0], v[1],
width=0.002, head_width=.1, length_includes_head=True)
plt.annotate("v", v, xytext=[v[0] + 0.01, v[1]])
plt.arrow(0 ,0, w[0], w[1],
width=0.002, head_width=.1, length_includes_head=True)
plt.annotate("w", w, xytext=[w[0] - 0.1, w[1]])
plt.arrow(0, 0, vonw[0], vonw[1], length_includes_head=True)
plt.annotate(u"(v•w)w", vonw, xytext=[vonw[0] - 0.1, vonw[1] + 0.02])
plt.arrow(v[0], v[1], vonw[0] - v[0], vonw[1] - v[1],
linestyle='dotted', length_includes_head=True)
plt.scatter(*zip(v,w,o),marker='.')
plt.axis('equal')
plt.show()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/vikniksor/DataScience/blob/main/comparing_some_clf_models.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
import itertools
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.ticker import NullFormatter
import pandas as pd
import numpy as np
import matplotlib.ticker as ticker
from sklearn import preprocessing
%matplotlib inline
```
### About dataset
This dataset is about past loans. The __Loan_train.csv__ data set includes details of 346 customers whose loan are already paid off or defaulted. It includes following fields:
| Field | Description |
|----------------|---------------------------------------------------------------------------------------|
| Loan_status | Whether a loan is paid off on in collection |
| Principal | Basic principal loan amount at the |
| Terms | Origination terms which can be weekly (7 days), biweekly, and monthly payoff schedule |
| Effective_date | When the loan got originated and took effects |
| Due_date | Since it’s one-time payoff schedule, each loan has one single due date |
| Age | Age of applicant |
| Education | Education of applicant |
| Gender | The gender of applicant |
Lets download the dataset
```
!wget -O loan_train.csv https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/ML0101ENv3/labs/loan_train.csv
```
### Load Data From CSV File
```
df = pd.read_csv('loan_train.csv')
df.head()
df.shape
```
### Convert to date time object
```
df['due_date'] = pd.to_datetime(df['due_date'])
df['effective_date'] = pd.to_datetime(df['effective_date'])
df.head()
```
# Data visualization and pre-processing
Let’s see how many of each class is in our data set
```
df['loan_status'].value_counts()
```
260 people have paid off the loan on time while 86 have gone into collection
Lets plot some columns to underestand data better:
```
import seaborn as sns
bins = np.linspace(df.Principal.min(), df.Principal.max(), 10)
g = sns.FacetGrid(df, col="Gender", hue="loan_status", palette="Set1", col_wrap=2)
g.map(plt.hist, 'Principal', bins=bins, ec="k")
g.axes[-1].legend()
plt.show()
bins = np.linspace(df.age.min(), df.age.max(), 10)
g = sns.FacetGrid(df, col="Gender", hue="loan_status", palette="Set1", col_wrap=2)
g.map(plt.hist, 'age', bins=bins, ec="k")
g.axes[-1].legend()
plt.show()
```
# Pre-processing: Feature selection/extraction
```
df['dayofweek'] = df['effective_date'].dt.dayofweek
bins = np.linspace(df.dayofweek.min(), df.dayofweek.max(), 10)
g = sns.FacetGrid(df, col="Gender", hue="loan_status", palette="Set1", col_wrap=2)
g.map(plt.hist, 'dayofweek', bins=bins, ec="k")
g.axes[-1].legend()
plt.show()
```
We see that people who get the loan at the end of the week dont pay it off, so lets use Feature binarization to set a threshold values less then day 4
```
df['weekend'] = df['dayofweek'].apply(lambda x: 1 if (x>3) else 0)
df.head()
```
**Convert Categorical features to numerical values**
Lets look at gender:
```
df.groupby(['Gender'])['loan_status'].value_counts(normalize=True)
```
86 % of female pay there loans while only 73 % of males pay there loan
Lets convert male to 0 and female to 1:
```
df['Gender'].replace(to_replace=['male','female'], value=[0,1],inplace=True)
df.head()
```
**One Hot Encoding**
How about education?
```
df.groupby(['education'])['loan_status'].value_counts(normalize=True)
```
Feature befor One Hot Encoding
```
df[['Principal','terms','age','Gender','education']].head()
```
Use one hot encoding technique to conver categorical varables to binary variables and append them to the feature Data Frame
```
Feature = df[['Principal','terms','age','Gender','weekend']]
Feature = pd.concat([Feature,pd.get_dummies(df['education'])], axis=1)
Feature.drop(['Master or Above'], axis = 1,inplace=True)
Feature.head()
```
Feature selection
Lets defind feature sets, X:
```
X = Feature
X[0:5]
```
What are our lables?
```
y = df['loan_status'].values
y[0:5]
```
**Normalize Data**
Data Standardization give data zero mean and unit variance (technically should be done after train test split )
```
X = preprocessing.StandardScaler().fit(X).transform(X)
X[0:5]
```
# Classification
Will use:
- K Nearest Neighbor(KNN)
- Decision Tree
- Support Vector Machine
- Logistic Regression
# K Nearest Neighbor(KNN)
```
from sklearn.model_selection import train_test_split
from sklearn.neighbors import KNeighborsClassifier
X_knn, y_knn = X, y
X_knn_train, X_knn_valid, y_knn_train, y_knn_valid = train_test_split(X_knn, y_knn, test_size = 0.4, random_state = 6)
scores = {}
for k in range(1, 10):
knn_clf = KNeighborsClassifier(k)
knn_clf.fit(X_knn_train, y_knn_train)
knn_clf.predict(X_knn_valid)
scores[k] = knn_clf.score(X_knn_valid, y_knn_valid)
print(scores)
knn_clf = KNeighborsClassifier(n_neighbors = 5)
knn_clf.fit(X, y)
```
# Decision Tree
```
from sklearn.tree import DecisionTreeClassifier
dt_clf = DecisionTreeClassifier()
dt_clf.fit(X, y)
```
# Support Vector Machine
```
from sklearn.svm import SVC
svc_clf = SVC()
svc_clf.fit(X, y)
```
# Logistic Regression
```
from sklearn.linear_model import LogisticRegression
logreg = LogisticRegression(C = 0.1, class_weight = "balanced")
logreg.fit(X, y)
```
# Model Evaluation using Test set
```
from sklearn.metrics import jaccard_score
from sklearn.metrics import f1_score
from sklearn.metrics import log_loss
```
First, download and load the test set:
```
!wget -O loan_test.csv https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/ML0101ENv3/labs/loan_test.csv
```
### Load Test set for evaluation
```
test_df = pd.read_csv('loan_test.csv')
test_df.head()
test_df['due_date'] = pd.to_datetime(test_df['due_date'])
test_df['effective_date'] = pd.to_datetime(test_df['effective_date'])
test_df['dayofweek'] = test_df['effective_date'].dt.dayofweek
# evaulate weekend field
test_df['weekend'] = test_df['dayofweek'].apply(lambda x: 1 if (x>3) else 0)
# work out education level
test_feature = test_df[['Principal','terms','age','Gender','weekend']]
test_feature = pd.concat([test_feature,pd.get_dummies(test_df['education'])], axis=1)
test_feature.drop(['Master or Above'], axis = 1,inplace=True)
test_feature.tail()
# normalize the test data
test_X = preprocessing.StandardScaler().fit(test_feature).transform(test_feature)
y_test = test_df["loan_status"].values
knn_pred = knn_clf.predict(test_X)
#jk1 = jaccard_score(y_test, knn_pred, pos_label = "PAIDOFF")
fs1 = f1_score(y_test, knn_pred, average = "weighted")
dt_pred = dt_clf.predict(test_X)
#jk2 = jaccard_score(y_test, dt_pred, pos_label = "PAIDOFF")
fs2 = f1_score(y_test, dt_pred, average = "weighted")
svm_pred = svc_clf.predict(test_X)
#jk3 = jaccard_score(y_test, svm_pred, pos_label = "PAIDOFF")
fs3 = f1_score(y_test, svm_pred, average = "weighted")
logreg_pred = logreg.predict(test_X)
#jk4 = jaccard_score(y_test, logreg_pred, pos_label = "PAIDOFF")
fs4 = f1_score(y_test, logreg_pred, average = "weighted")
logreg_proba = logreg.predict_proba(test_X)
ll4 = log_loss(y_test, logreg_proba)
list_jk = [JK1, JK2, JK3, JK4]
list_fs = [fs1, fs2, fs3, fs4]
list_ll = ['NA', 'NA', 'NA', ll4]
report_df = pd.DataFrame(list_jk, index=['KNN','Decision Tree','SVM','Logistic Regression'])
report_df.columns = ['Jaccard']
report_df.insert(loc=1, column='F1-score', value=list_fs)
report_df.insert(loc=2, column='LogLoss', value=list_ll)
report_df.columns.name = 'Algorithm'
report_df
```
| github_jupyter |
```
from scipy.cluster.hierarchy import linkage, fcluster
import matplotlib.pyplot as plt
import seaborn as sns, pandas as pd
x_coords = [80.1, 93.1, 86.6, 98.5, 86.4, 9.5, 15.2, 3.4, 10.4, 20.3, 44.2, 56.8, 49.2, 62.5]
y_coords = [87.2, 96.1, 95.6, 92.4, 92.4, 57.7, 49.4, 47.3, 59.1, 55.5, 25.6, 2.1, 10.9, 24.1]
df = pd.DataFrame({"x_coord" : x_coords, "y_coord": y_coords})
df.head()
Z = linkage(df, "ward")
df["cluster_labels"] = fcluster(Z, 3, criterion="maxclust")
df.head(3)
sns.scatterplot(x="x_coord", y="y_coord", hue="cluster_labels", data=df)
plt.show()
```
### K-means clustering in SciPy
#### two steps of k-means clustering:
* Define cluster centers through kmeans() function.
* It has two required arguments: observations and number of clusters.
* Assign cluster labels through the vq() function.
* It has two required arguments: observations and cluster centers.
```
from scipy.cluster.vq import kmeans, vq
import random
# Generate cluster centers
cluster_centers, distortion = kmeans(comic_con[["x_scaled","y_scaled"]], 2)
# Assign cluster labels
comic_con['cluster_labels'], distortion_list = vq(comic_con[["x_scaled","y_scaled"]], cluster_centers)
# Plot clusters
sns.scatterplot(x='x_scaled', y='y_scaled',
hue='cluster_labels', data = comic_con)
plt.show()
random.seed((1000,2000))
centroids, _ = kmeans(df, 3)
df["cluster_labels_kmeans"], _ = vq(df, centroids)
sns.scatterplot(x="x_coord", y="y_coord", hue="cluster_labels_kmeans", data=df)
plt.show()
```
### Normalization of Data
```
# Process of rescaling data to a standard deviation of 1
# x_new = x / std(x)
from scipy.cluster.vq import whiten
data = [5, 1, 3, 3, 2, 3, 3, 8, 1, 2, 2, 3, 5]
scaled_data = whiten(data)
scaled_data
plt.figure(figsize=(12,4))
plt.subplot(131)
plt.plot(data, label="original")
plt.legend()
plt.subplot(132)
plt.plot(scaled_data, label="scaled")
plt.legend()
plt.subplot(133)
plt.plot(data, label="original")
plt.plot(scaled_data, label="scaled")
plt.legend()
plt.show()
```
### Normalization of small numbers
```
# Prepare data
rate_cuts = [0.0025, 0.001, -0.0005, -0.001, -0.0005, 0.0025, -0.001, -0.0015, -0.001, 0.0005]
# Use the whiten() function to standardize the data
scaled_rate_cuts = whiten(rate_cuts)
plt.figure(figsize=(12,4))
plt.subplot(131)
plt.plot(rate_cuts, label="original")
plt.legend()
plt.subplot(132)
plt.plot(scaled_rate_cuts, label="scaled")
plt.legend()
plt.subplot(133)
plt.plot(rate_cuts, label='original')
plt.plot(scaled_rate_cuts, label='scaled')
plt.legend()
plt.show()
```
#### Hierarchical clustering: ward method
```
# Import the fcluster and linkage functions
from scipy.cluster.hierarchy import fcluster, linkage
# Use the linkage() function
distance_matrix = linkage(comic_con[['x_scaled', 'y_scaled']], method = "ward", metric = 'euclidean')
# Assign cluster labels
comic_con['cluster_labels'] = fcluster(distance_matrix, 2, criterion='maxclust')
# Plot clusters
sns.scatterplot(x='x_scaled', y='y_scaled',
hue='cluster_labels', data = comic_con)
plt.show()
```
#### Hierarchical clustering: single method
```
# Use the linkage() function
distance_matrix = linkage(comic_con[["x_scaled", "y_scaled"]], method = "single", metric = "euclidean")
# Assign cluster labels
comic_con['cluster_labels'] = fcluster(distance_matrix, 2, criterion="maxclust")
# Plot clusters
sns.scatterplot(x='x_scaled', y='y_scaled',
hue='cluster_labels', data = comic_con)
plt.show()
```
#### Hierarchical clustering: complete method
```
# Import the fcluster and linkage functions
from scipy.cluster.hierarchy import linkage, fcluster
# Use the linkage() function
distance_matrix = linkage(comic_con[["x_scaled", "y_scaled"]], method = "complete", metric = "euclidean")
# Assign cluster labels
comic_con['cluster_labels'] = fcluster(distance_matrix, 2, criterion="maxclust")
# Plot clusters
sns.scatterplot(x='x_scaled', y='y_scaled',
hue='cluster_labels', data = comic_con)
plt.show()
```
### Visualizing Data
```
# Import the pyplot class
import matplotlib.pyplot as plt
# Define a colors dictionary for clusters
colors = {1:'red', 2:'blue'}
# Plot a scatter plot
comic_con.plot.scatter(x="x_scaled",
y="y_scaled",
c=comic_con['cluster_labels'].apply(lambda x: colors[x]))
plt.show()
# Import the seaborn module
import seaborn as sns
# Plot a scatter plot using seaborn
sns.scatterplot(x="x_scaled",
y="y_scaled",
hue="cluster_labels",
data = comic_con)
plt.show()
```
### Dendogram
```
from scipy.cluster.hierarchy import dendrogram
Z = linkage(df[['x_whiten', 'y_whiten']], method='ward', metric='euclidean')
dn = dendrogram(Z)
plt.show()
### timing using %timeit
%timeit sum([1,3,5])
```
### Finding optimum "k" Elbow Method
```
distortions = []
num_clusters = range(1, 7)
# Create a list of distortions from the kmeans function
for i in num_clusters:
cluster_centers, distortion = kmeans(comic_con[["x_scaled","y_scaled"]], i)
distortions.append(distortion)
# Create a data frame with two lists - num_clusters, distortions
elbow_plot_data = pd.DataFrame({'num_clusters': num_clusters, 'distortions': distortions})
# Creat a line plot of num_clusters and distortions
sns.lineplot(x="num_clusters", y="distortions", data = elbow_plot_data)
plt.xticks(num_clusters)
plt.show()
```
### Elbow method on uniform data
```
# Let us now see how the elbow plot looks on a data set with uniformly distributed points.
distortions = []
num_clusters = range(2, 7)
# Create a list of distortions from the kmeans function
for i in num_clusters:
cluster_centers, distortion = kmeans(uniform_data[["x_scaled","y_scaled"]], i)
distortions.append(distortion)
# Create a data frame with two lists - number of clusters and distortions
elbow_plot = pd.DataFrame({'num_clusters': num_clusters, 'distortions': distortions})
# Creat a line plot of num_clusters and distortions
sns.lineplot(x="num_clusters", y="distortions", data=elbow_plot)
plt.xticks(num_clusters)
plt.show()
```
### Impact of seeds on distinct clusters
Notice that kmeans is unable to capture the three visible clusters clearly, and the two clusters towards the top have taken in some points along the boundary. This happens due to the underlying assumption in kmeans algorithm to minimize distortions which leads to clusters that are similar in terms of area.
### Dominant Colors in Images
#### Extracting RGB values from image
There are broadly three steps to find the dominant colors in an image:
* Extract RGB values into three lists.
* Perform k-means clustering on scaled RGB values.
* Display the colors of cluster centers.
To extract RGB values, we use the imread() function of the image class of matplotlib.
```
# Import image class of matplotlib
import matplotlib.image as img
from matplotlib.pyplot import imshow
# Read batman image and print dimensions
sea_horizon = img.imread("../00_DataSets/img/sea_horizon.jpg")
print(sea_horizon.shape)
imshow(sea_horizon)
# Store RGB values of all pixels in lists r, g and b
r, g, b = [], [], []
for row in sea_horizon:
for temp_r, temp_g, temp_b in row:
r.append(temp_r)
g.append(temp_g)
b.append(temp_b)
sea_horizon_df = pd.DataFrame({'red': r, 'blue': b, 'green': g})
sea_horizon_df.head()
distortions = []
num_clusters = range(1, 7)
# Create a list of distortions from the kmeans function
for i in num_clusters:
cluster_centers, distortion = kmeans(sea_horizon_df[["red", "blue", "green"]], i)
distortions.append(distortion)
# Create a data frame with two lists, num_clusters and distortions
elbow_plot_data = pd.DataFrame({"num_clusters":num_clusters, "distortions":distortions})
# Create a line plot of num_clusters and distortions
sns.lineplot(x="num_clusters", y="distortions", data = elbow_plot_data)
plt.xticks(num_clusters)
plt.show()
# scaling the data
sea_horizon_df["scaled_red"] = whiten(sea_horizon_df["red"])
sea_horizon_df["scaled_blue"] = whiten(sea_horizon_df["blue"])
sea_horizon_df["scaled_green"] = whiten(sea_horizon_df["green"])
distortions = []
num_clusters = range(1, 7)
# Create a list of distortions from the kmeans function
for i in num_clusters:
cluster_centers, distortion = kmeans(sea_horizon_df[["scaled_red", "scaled_blue", "scaled_green"]], i)
distortions.append(distortion)
# Create a data frame with two lists, num_clusters and distortions
elbow_plot_data = pd.DataFrame({"num_clusters":num_clusters, "distortions":distortions})
# Create a line plot of num_clusters and distortions
sns.lineplot(x="num_clusters", y="distortions", data = elbow_plot_data)
plt.xticks(num_clusters)
plt.show()
```
#### Show Dominant colors
To display the dominant colors, convert the colors of the cluster centers to their raw values and then converted them to the range of 0-1, using the following formula: converted_pixel = standardized_pixel * pixel_std / 255
```
# Get standard deviations of each color
r_std, g_std, b_std = sea_horizon_df[['red', 'green', 'blue']].std()
colors = []
for cluster_center in cluster_centers:
scaled_red, scaled_green, scaled_blue = cluster_center
# Convert each standardized value to scaled value
colors.append((
scaled_red * r_std / 255,
scaled_green * g_std / 255,
scaled_blue * b_std / 255
))
# Display colors of cluster centers
plt.imshow([colors])
plt.show()
```
### Document clustering
```
# TF-IDF of movie plots
from sklearn.feature_extraction.text import TfidfVectorizer
# Import TfidfVectorizer class from sklearn
from sklearn.feature_extraction.text import TfidfVectorizer
# Initialize TfidfVectorizer
tfidf_vectorizer = TfidfVectorizer(max_df=0.75, min_df=0.1, max_features=50, tokenizer=remove_noise)
# Use the .fit_transform() method on the list plots
tfidf_matrix = tfidf_vectorizer.fit_transform(plots)
num_clusters = 2
# Generate cluster centers through the kmeans function
cluster_centers, distortion = kmeans(tfidf_matrix.todense(), num_clusters)
# Generate terms from the tfidf_vectorizer object
terms = tfidf_vectorizer.get_feature_names()
for i in range(num_clusters):
# Sort the terms and print top 3 terms
center_terms = dict(zip(terms, list(cluster_centers[i])))
sorted_terms = sorted(center_terms, key=center_terms.get, reverse=True)
print(sorted_terms[:3])
```
| github_jupyter |
# Gradient Descent
:label:`sec_gd`
In this section we are going to introduce the basic concepts underlying gradient descent. This is brief by necessity. See e.g., :cite:`Boyd.Vandenberghe.2004` for an in-depth introduction to convex optimization. Although the latter is rarely used directly in deep learning, an understanding of gradient descent is key to understanding stochastic gradient descent algorithms. For instance, the optimization problem might diverge due to an overly large learning rate. This phenomenon can already be seen in gradient descent. Likewise, preconditioning is a common technique in gradient descent and carries over to more advanced algorithms. Let us start with a simple special case.
## Gradient Descent in One Dimension
Gradient descent in one dimension is an excellent example to explain why the gradient descent algorithm may reduce the value of the objective function. Consider some continuously differentiable real-valued function $f: \mathbb{R} \rightarrow \mathbb{R}$. Using a Taylor expansion (:numref:`sec_single_variable_calculus`) we obtain that
$$f(x + \epsilon) = f(x) + \epsilon f'(x) + \mathcal{O}(\epsilon^2).$$
:eqlabel:`gd-taylor`
That is, in first approximation $f(x+\epsilon)$ is given by the function value $f(x)$ and the first derivative $f'(x)$ at $x$. It is not unreasonable to assume that for small $\epsilon$ moving in the direction of the negative gradient will decrease $f$. To keep things simple we pick a fixed step size $\eta > 0$ and choose $\epsilon = -\eta f'(x)$. Plugging this into the Taylor expansion above we get
$$f(x - \eta f'(x)) = f(x) - \eta f'^2(x) + \mathcal{O}(\eta^2 f'^2(x)).$$
If the derivative $f'(x) \neq 0$ does not vanish we make progress since $\eta f'^2(x)>0$. Moreover, we can always choose $\eta$ small enough for the higher order terms to become irrelevant. Hence we arrive at
$$f(x - \eta f'(x)) \lessapprox f(x).$$
This means that, if we use
$$x \leftarrow x - \eta f'(x)$$
to iterate $x$, the value of function $f(x)$ might decline. Therefore, in gradient descent we first choose an initial value $x$ and a constant $\eta > 0$ and then use them to continuously iterate $x$ until the stop condition is reached, for example, when the magnitude of the gradient $|f'(x)|$ is small enough or the number of iterations has reached a certain value.
For simplicity we choose the objective function $f(x)=x^2$ to illustrate how to implement gradient descent. Although we know that $x=0$ is the solution to minimize $f(x)$, we still use this simple function to observe how $x$ changes. As always, we begin by importing all required modules.
```
%mavenRepo snapshots https://oss.sonatype.org/content/repositories/snapshots/
%maven ai.djl:api:0.7.0-SNAPSHOT
%maven org.slf4j:slf4j-api:1.7.26
%maven org.slf4j:slf4j-simple:1.7.26
%maven ai.djl.mxnet:mxnet-engine:0.7.0-SNAPSHOT
%maven ai.djl.mxnet:mxnet-native-auto:1.7.0-b
%load ../utils/plot-utils
%load ../utils/Functions.java
import ai.djl.ndarray.*;
import tech.tablesaw.plotly.traces.ScatterTrace;
import java.lang.Math;
Function<Float, Float> f = x -> x * x; // Objective Function
Function<Float, Float> gradf = x -> 2 * x; // Its Derivative
NDManager manager = NDManager.newBaseManager();
```
Next, we use $x=10$ as the initial value and assume $\eta=0.2$. Using gradient descent to iterate $x$ for 10 times we can see that, eventually, the value of $x$ approaches the optimal solution.
```
public float[] gd(float eta) {
float x = 10f;
float[] results = new float[11];
results[0] = x;
for (int i = 0; i < 10; i++) {
x -= eta * gradf.apply(x);
results[i + 1] = x;
}
System.out.printf("epoch 10, x: %f\n", x);
return results;
}
float[] res = gd(0.2f);
```
The progress of optimizing over $x$ can be plotted as follows.
```
/* Saved in GradDescUtils.java */
public void plotGD(float[] x, float[] y, float[] segment, Function<Float, Float> func,
int width, int height) {
// Function Line
ScatterTrace trace = ScatterTrace.builder(Functions.floatToDoubleArray(x),
Functions.floatToDoubleArray(y))
.mode(ScatterTrace.Mode.LINE)
.build();
// GD Line
ScatterTrace trace2 = ScatterTrace.builder(Functions.floatToDoubleArray(segment),
Functions.floatToDoubleArray(Functions.callFunc(segment, func)))
.mode(ScatterTrace.Mode.LINE)
.build();
// GD Points
ScatterTrace trace3 = ScatterTrace.builder(Functions.floatToDoubleArray(segment),
Functions.floatToDoubleArray(Functions.callFunc(segment, func)))
.build();
Layout layout = Layout.builder()
.height(height)
.width(width)
.showLegend(false)
.build();
display(new Figure(layout, trace, trace2, trace3));
}
/* Saved in GradDescUtils.java */
public void showTrace(float[] res) {
float n = 0;
for (int i = 0; i < res.length; i++) {
if (Math.abs(res[i]) > n) {
n = Math.abs(res[i]);
}
}
NDArray fLineND = manager.arange(-n, n, 0.01f);
float[] fLine = fLineND.toFloatArray();
plotGD(fLine, Functions.callFunc(fLine, f), res, f, 500, 400);
}
showTrace(res);
```
### Learning Rate
:label:`section_gd-learningrate`
The learning rate $\eta$ can be set by the algorithm designer. If we use a learning rate that is too small, it will cause $x$ to update very slowly, requiring more iterations to get a better solution. To show what happens in such a case, consider the progress in the same optimization problem for $\eta = 0.05$. As we can see, even after 10 steps we are still very far from the optimal solution.
```
showTrace(gd(0.05f));
```
Conversely, if we use an excessively high learning rate, $\left|\eta f'(x)\right|$ might be too large for the first-order Taylor expansion formula. That is, the term $\mathcal{O}(\eta^2 f'^2(x))$ in :eqref:`gd-taylor` might become significant. In this case, we cannot guarantee that the iteration of $x$ will be able to lower the value of $f(x)$. For example, when we set the learning rate to $\eta=1.1$, $x$ overshoots the optimal solution $x=0$ and gradually diverges.
```
showTrace(gd(1.1f));
```
### Local Minima
To illustrate what happens for nonconvex functions consider the case of $f(x) = x \cdot \cos c x$. This function has infinitely many local minima. Depending on our choice of learning rate and depending on how well conditioned the problem is, we may end up with one of many solutions. The example below illustrates how an (unrealistically) high learning rate will lead to a poor local minimum.
```
float c = (float)(0.15f * Math.PI);
Function<Float, Float> f = x -> x * (float)Math.cos(c * x);
Function<Float, Float> gradf = x -> (float)(Math.cos(c * x) - c * x * Math.sin(c * x));
showTrace(gd(2));
```
## Multivariate Gradient Descent
Now that we have a better intuition of the univariate case, let us consider the situation where $\mathbf{x} \in \mathbb{R}^d$. That is, the objective function $f: \mathbb{R}^d \to \mathbb{R}$ maps vectors into scalars. Correspondingly its gradient is multivariate, too. It is a vector consisting of $d$ partial derivatives:
$$\nabla f(\mathbf{x}) = \bigg[\frac{\partial f(\mathbf{x})}{\partial x_1}, \frac{\partial f(\mathbf{x})}{\partial x_2}, \ldots, \frac{\partial f(\mathbf{x})}{\partial x_d}\bigg]^\top.$$
Each partial derivative element $\partial f(\mathbf{x})/\partial x_i$ in the gradient indicates the rate of change of $f$ at $\mathbf{x}$ with respect to the input $x_i$. As before in the univariate case we can use the corresponding Taylor approximation for multivariate functions to get some idea of what we should do. In particular, we have that
$$f(\mathbf{x} + \mathbf{\epsilon}) = f(\mathbf{x}) + \mathbf{\epsilon}^\top \nabla f(\mathbf{x}) + \mathcal{O}(\|\mathbf{\epsilon}\|^2).$$
:eqlabel:`gd-multi-taylor`
In other words, up to second order terms in $\mathbf{\epsilon}$ the direction of steepest descent is given by the negative gradient $-\nabla f(\mathbf{x})$. Choosing a suitable learning rate $\eta > 0$ yields the prototypical gradient descent algorithm:
$\mathbf{x} \leftarrow \mathbf{x} - \eta \nabla f(\mathbf{x}).$
To see how the algorithm behaves in practice let us construct an objective function $f(\mathbf{x})=x_1^2+2x_2^2$ with a two-dimensional vector $\mathbf{x} = [x_1, x_2]^\top$ as input and a scalar as output. The gradient is given by $\nabla f(\mathbf{x}) = [2x_1, 4x_2]^\top$. We will observe the trajectory of $\mathbf{x}$ by gradient descent from the initial position $[-5, -2]$. We need two more helper functions. The first uses an update function and applies it $20$ times to the initial value. The second helper visualizes the trajectory of $\mathbf{x}$.
We also create a `Weights` class to make it easier to store the weight parameters and return them in functions.
```
/* Saved in GradDescUtils.java */
public class Weights {
public float x1, x2;
public Weights(float x1, float x2) {
this.x1 = x1;
this.x2 = x2;
}
}
/* Saved in GradDescUtils.java */
/* Optimize a 2D objective function with a customized trainer. */
public ArrayList<Weights> train2d(Function<Float[], Float[]> trainer, int steps) {
// s1 and s2 are internal state variables and will
// be used later in the chapter
float x1 = -5f, x2 = -2f, s1 = 0f, s2 = 0f;
ArrayList<Weights> results = new ArrayList<>();
results.add(new Weights(x1, x2));
for (int i = 1; i < steps + 1; i++) {
Float[] step = trainer.apply(new Float[]{x1, x2, s1, s2});
x1 = step[0];
x2 = step[1];
s1 = step[2];
s2 = step[3];
results.add(new Weights(x1, x2));
System.out.printf("epoch %d, x1 %f, x2 %f\n", i, x1, x2);
}
return results;
}
import java.util.function.BiFunction;
/* Saved in GradDescUtils.java */
/* Show the trace of 2D variables during optimization. */
public void showTrace2d(BiFunction<Float, Float, Float> f, ArrayList<Weights> results) {
// TODO: add when tablesaw adds support for contour and meshgrids
}
```
Next, we observe the trajectory of the optimization variable $\mathbf{x}$ for learning rate $\eta = 0.1$. We can see that after 20 steps the value of $\mathbf{x}$ approaches its minimum at $[0, 0]$. Progress is fairly well-behaved albeit rather slow.
```
float eta = 0.1f;
BiFunction<Float, Float, Float> f = (x1, x2) -> x1 * x1 + 2 * x2 * x2; // Objective
BiFunction<Float, Float, Float[]> gradf = (x1, x2) -> new Float[]{2 * x1, 4 * x2}; // Gradient
Function<Float[], Float[]> gd = (state) -> {
Float x1 = state[0];
Float x2 = state[1];
Float[] g = gradf.apply(x1, x2); // Compute Gradient
Float g1 = g[0];
Float g2 = g[1];
return new Float[]{x1 - eta * g1, x2 - eta * g2, 0f, 0f}; // Update Variables
};
showTrace2d(f, train2d(gd, 20));
```

## Adaptive Methods
As we could see in :numref:`section_gd-learningrate`, getting the learning rate $\eta$ "just right" is tricky. If we pick it too small, we make no progress. If we pick it too large, the solution oscillates and in the worst case it might even diverge. What if we could determine $\eta$ automatically or get rid of having to select a step size at all? Second order methods that look not only at the value and gradient of the objective but also at its *curvature* can help in this case. While these methods cannot be applied to deep learning directly due to the computational cost, they provide useful intuition into how to design advanced optimization algorithms that mimic many of the desirable properties of the algorithms outlined below.
### Newton's Method
Reviewing the Taylor expansion of $f$ there is no need to stop after the first term. In fact, we can write it as
$$f(\mathbf{x} + \mathbf{\epsilon}) = f(\mathbf{x}) + \mathbf{\epsilon}^\top \nabla f(\mathbf{x}) + \frac{1}{2} \mathbf{\epsilon}^\top \nabla \nabla^\top f(\mathbf{x}) \mathbf{\epsilon} + \mathcal{O}(\|\mathbf{\epsilon}\|^3).$$
:eqlabel:`gd-hot-taylor`
To avoid cumbersome notation we define $H_f := \nabla \nabla^\top f(\mathbf{x})$ to be the *Hessian* of $f$. This is a $d \times d$ matrix. For small $d$ and simple problems $H_f$ is easy to compute. For deep networks, on the other hand, $H_f$ may be prohibitively large, due to the cost of storing $\mathcal{O}(d^2)$ entries. Furthermore it may be too expensive to compute via backprop as we would need to apply backprop to the backpropagation call graph. For now let us ignore such considerations and look at what algorithm we'd get.
After all, the minimum of $f$ satisfies $\nabla f(\mathbf{x}) = 0$. Taking derivatives of :eqref:`gd-hot-taylor` with regard to $\mathbf{\epsilon}$ and ignoring higher order terms we arrive at
$$\nabla f(\mathbf{x}) + H_f \mathbf{\epsilon} = 0 \text{ and hence }
\mathbf{\epsilon} = -H_f^{-1} \nabla f(\mathbf{x}).$$
That is, we need to invert the Hessian $H_f$ as part of the optimization problem.
For $f(x) = \frac{1}{2} x^2$ we have $\nabla f(x) = x$ and $H_f = 1$. Hence for any $x$ we obtain $\epsilon = -x$. In other words, a single step is sufficient to converge perfectly without the need for any adjustment! Alas, we got a bit lucky here since the Taylor expansion was exact. Let us see what happens in other problems.
```
float c = 0.5f;
Function<Float, Float> f = x -> (float)Math.cosh(c * x); // Objective
Function<Float, Float> gradf = x -> c * (float)Math.sinh(c * x); // Derivative
Function<Float, Float> hessf = x -> c * c * (float)Math.cosh(c * x); // Hessian
// Hide learning rate for now
public float[] newton(float eta) {
float x = 10f;
float[] results = new float[11];
results[0] = x;
for (int i = 0; i < 10; i++) {
x -= eta * gradf.apply(x) / hessf.apply(x);
results[i + 1] = x;
}
System.out.printf("epoch 10, x: %f\n", x);
return results;
}
showTrace(newton(1));
```
Now let us see what happens when we have a *nonconvex* function, such as $f(x) = x \cos(c x)$. After all, note that in Newton's method we end up dividing by the Hessian. This means that if the second derivative is *negative* we would walk into the direction of *increasing* $f$. That is a fatal flaw of the algorithm. Let us see what happens in practice.
```
c = 0.15f * (float)Math.PI;
Function<Float, Float> f = x -> x * (float)Math.cos(c * x);
Function<Float, Float> gradf = x -> (float)(Math.cos(c * x) - c * x * Math.sin(c * x));
Function<Float, Float> hessf = x -> (float)(-2 * c * Math.sin(c * x) -
x * c * c * Math.cos(c * x));
showTrace(newton(1));
```
This went spectacularly wrong. How can we fix it? One way would be to "fix" the Hessian by taking its absolute value instead. Another strategy is to bring back the learning rate. This seems to defeat the purpose, but not quite. Having second order information allows us to be cautious whenever the curvature is large and to take longer steps whenever the objective is flat. Let us see how this works with a slightly smaller learning rate, say $\eta = 0.5$. As we can see, we have quite an efficient algorithm.
```
showTrace(newton(0.5f));
```
### Convergence Analysis
We only analyze the convergence rate for convex and three times differentiable $f$, where at its minimum $x^*$ the second derivative is nonzero, i.e., where $f''(x^*) > 0$. The multivariate proof is a straightforward extension of the argument below and omitted since it doesn't help us much in terms of intuition.
Denote by $x_k$ the value of $x$ at the $k$-th iteration and let $e_k := x_k - x^*$ be the distance from optimality. By Taylor series expansion we have that the condition $f'(x^*) = 0$ can be written as
$$0 = f'(x_k - e_k) = f'(x_k) - e_k f''(x_k) + \frac{1}{2} e_k^2 f'''(\xi_k).$$
This holds for some $\xi_k \in [x_k - e_k, x_k]$. Recall that we have the update $x_{k+1} = x_k - f'(x_k) / f''(x_k)$. Dividing the above expansion by $f''(x_k)$ yields
$$e_k - f'(x_k) / f''(x_k) = \frac{1}{2} e_k^2 f'''(\xi_k) / f''(x_k).$$
Plugging in the update equations leads to the following bound $e_{k+1} \leq e_k^2 f'''(\xi_k) / f'(x_k)$. Consequently, whenever we are in a region of bounded $f'''(\xi_k) / f''(x_k) \leq c$, we have a quadratically decreasing error $e_{k+1} \leq c e_k^2$.
As an aside, optimization researchers call this *linear* convergence, whereas a condition such as $e_{k+1} \leq \alpha e_k$ would be called a *constant* rate of convergence.
Note that this analysis comes with a number of caveats: We do not really have much of a guarantee when we will reach the region of rapid convergence. Instead, we only know that once we reach it, convergence will be very quick. Second, this requires that $f$ is well-behaved up to higher order derivatives. It comes down to ensuring that $f$ does not have any "surprising" properties in terms of how it might change its values.
### Preconditioning
Quite unsurprisingly computing and storing the full Hessian is very expensive. It is thus desirable to find alternatives. One way to improve matters is by avoiding to compute the Hessian in its entirety but only compute the *diagonal* entries. While this is not quite as good as the full Newton method, it is still much better than not using it. Moreover, estimates for the main diagonal elements are what drives some of the innovation in stochastic gradient descent optimization algorithms. This leads to update algorithms of the form
$$\mathbf{x} \leftarrow \mathbf{x} - \eta \mathrm{diag}(H_f)^{-1} \nabla \mathbf{x}.$$
To see why this might be a good idea consider a situation where one variable denotes height in millimeters and the other one denotes height in kilometers. Assuming that for both the natural scale is in meters we have a terrible mismatch in parameterizations. Using preconditioning removes this. Effectively preconditioning with gradient descent amounts to selecting a different learning rate for each coordinate.
### Gradient Descent with Line Search
One of the key problems in gradient descent was that we might overshoot the goal or make insufficient progress. A simple fix for the problem is to use line search in conjunction with gradient descent. That is, we use the direction given by $\nabla f(\mathbf{x})$ and then perform binary search as to which step length $\eta$ minimizes $f(\mathbf{x} - \eta \nabla f(\mathbf{x}))$.
This algorithm converges rapidly (for an analysis and proof see e.g., :cite:`Boyd.Vandenberghe.2004`). However, for the purpose of deep learning this is not quite so feasible, since each step of the line search would require us to evaluate the objective function on the entire dataset. This is way too costly to accomplish.
## Summary
* Learning rates matter. Too large and we diverge, too small and we do not make progress.
* Gradient descent can get stuck in local minima.
* In high dimensions adjusting learning the learning rate is complicated.
* Preconditioning can help with scale adjustment.
* Newton's method is a lot faster *once* it has started working properly in convex problems.
* Beware of using Newton's method without any adjustments for nonconvex problems.
## Exercises
1. Experiment with different learning rates and objective functions for gradient descent.
1. Implement line search to minimize a convex function in the interval $[a, b]$.
* Do you need derivatives for binary search, i.e., to decide whether to pick $[a, (a+b)/2]$ or $[(a+b)/2, b]$.
* How rapid is the rate of convergence for the algorithm?
* Implement the algorithm and apply it to minimizing $\log (\exp(x) + \exp(-2*x -3))$.
1. Design an objective function defined on $\mathbb{R}^2$ where gradient descent is exceedingly slow. Hint - scale different coordinates differently.
1. Implement the lightweight version of Newton's method using preconditioning:
* Use diagonal Hessian as preconditioner.
* Use the absolute values of that rather than the actual (possibly signed) values.
* Apply this to the problem above.
1. Apply the algorithm above to a number of objective functions (convex or not). What happens if you rotate coordinates by $45$ degrees?
| github_jupyter |
## Demo 4: HKR multiclass and fooling
[](https://colab.research.google.com/github/deel-ai/deel-lip/blob/master/doc/notebooks/demo4.ipynb)
This notebook will show how to train a lispchitz network in a multiclass setup.
The HKR is extended to multiclass using a one-vs all setup. It will go through
the process of designing and training the network. It will also show how to create robustness certificates from the output of the network. Finally these
certificates will be checked by attacking the network.
### installation
First, we install the required libraries. `Foolbox` will allow to perform adversarial attacks on the trained network.
```
# pip install deel-lip foolbox -qqq
from deel.lip.layers import (
SpectralDense,
SpectralConv2D,
ScaledL2NormPooling2D,
ScaledAveragePooling2D,
FrobeniusDense,
)
from deel.lip.model import Sequential
from deel.lip.activations import GroupSort, FullSort
from deel.lip.losses import MulticlassHKR, MulticlassKR
from deel.lip.callbacks import CondenseCallback
from tensorflow.keras.layers import Input, Flatten
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.datasets import mnist, fashion_mnist, cifar10
from tensorflow.keras.utils import to_categorical
from tensorflow.keras.preprocessing.image import ImageDataGenerator
import numpy as np
```
For this example, the dataset `fashion_mnist` will be used. In order to keep things simple, no data augmentation will be performed.
```
# load data
(x_train, y_train_ord), (x_test, y_test_ord) = fashion_mnist.load_data()
# standardize and reshape the data
x_train = np.expand_dims(x_train, -1) / 255
x_test = np.expand_dims(x_test, -1) / 255
# one hot encode the labels
y_train = to_categorical(y_train_ord)
y_test = to_categorical(y_test_ord)
```
Let's build the network.
### the architecture
The original one vs all setup would require 10 different networks ( 1 per class ), however, in practice we use a network with
a common body and 10 1-lipschitz heads. Experiments have shown that this setup don't affect the network performance. In order to ease the creation of such network, `FrobeniusDense` layer has a parameter for this: whenr `disjoint_neurons=True` it act as the stacking of 10 single neurons head. Note that, altough each head is a 1-lipschitz function the overall network is not 1-lipschitz (Concatenation is not 1-lipschitz). We will see later how this affects the certficate creation.
### the loss
The multiclass loss can be found in `HKR_multiclass_loss`. The loss has two params: `alpha` and `min_margin`. Decreasing `alpha` and increasing `min_margin` improve robustness (at the cost of accuracy). note also in the case of lipschitz networks, more robustness require more parameters. For more information see [our paper](https://arxiv.org/abs/2006.06520).
In this setup choosing `alpha=100`, `min_margin=.25` provide a good robustness without hurting the accuracy too much.
Finally the `KR_multiclass_loss()` indicate the robustness of the network ( proxy of the average certificate )
```
# Sequential (resp Model) from deel.model has the same properties as any lipschitz model.
# It act only as a container, with features specific to lipschitz
# functions (condensation, vanilla_exportation...)
model = Sequential(
[
Input(shape=x_train.shape[1:]),
# Lipschitz layers preserve the API of their superclass ( here Conv2D )
# an optional param is available: k_coef_lip which control the lipschitz
# constant of the layer
SpectralConv2D(
filters=16,
kernel_size=(3, 3),
activation=GroupSort(2),
use_bias=True,
kernel_initializer="orthogonal",
),
# usual pooling layer are implemented (avg, max...), but new layers are also available
ScaledL2NormPooling2D(pool_size=(2, 2), data_format="channels_last"),
SpectralConv2D(
filters=32,
kernel_size=(3, 3),
activation=GroupSort(2),
use_bias=True,
kernel_initializer="orthogonal",
),
ScaledL2NormPooling2D(pool_size=(2, 2), data_format="channels_last"),
# our layers are fully interoperable with existing keras layers
Flatten(),
SpectralDense(
64,
activation=GroupSort(2),
use_bias=True,
kernel_initializer="orthogonal",
),
FrobeniusDense(
y_train.shape[-1], activation=None, use_bias=False, kernel_initializer="orthogonal"
),
],
# similary model has a parameter to set the lipschitz constant
# to set automatically the constant of each layer
k_coef_lip=1.0,
name="hkr_model",
)
# HKR (Hinge-Krantorovich-Rubinstein) optimize robustness along with accuracy
model.compile(
# decreasing alpha and increasing min_margin improve robustness (at the cost of accuracy)
# note also in the case of lipschitz networks, more robustness require more parameters.
loss=MulticlassHKR(alpha=100, min_margin=.25),
optimizer=Adam(1e-4),
metrics=["accuracy", MulticlassKR()],
)
model.summary()
```
### notes about constraint enforcement
There are currently 3 way to enforce a constraint in a network:
1. regularization
2. weight reparametrization
3. weight projection
The first one don't provide the required garanties, this is why `deel-lip` focuses on the later two. Weight reparametrization is done directly in the layers (parameter `niter_bjorck`) this trick allow to perform arbitrary gradient updates without breaking the constraint. However this is done in the graph, increasing ressources consumption. The last method project the weights between each batch, ensuring the constraint at an more affordable computational cost. It can be done in `deel-lip` using the `CondenseCallback`. The main problem with this method is a reduced efficiency of each update.
As a rule of thumb, when reparametrization is used alone, setting `niter_bjorck` to at least 15 is advised. However when combined with weight projection, this setting can be lowered greatly.
```
# fit the model
model.fit(
x_train,
y_train,
batch_size=4096,
epochs=100,
validation_data=(x_test, y_test),
shuffle=True,
verbose=1,
)
```
### model exportation
Once training is finished, the model can be optimized for inference by using the `vanilla_export()` method.
```
# once training is finished you can convert
# SpectralDense layers into Dense layers and SpectralConv2D into Conv2D
# which optimize performance for inference
vanilla_model = model.vanilla_export()
```
### certificates generation and adversarial attacks
```
import foolbox as fb
from tensorflow import convert_to_tensor
import matplotlib.pyplot as plt
import tensorflow as tf
# we will test it on 10 samples one of each class
nb_adv = 10
hkr_fmodel = fb.TensorFlowModel(vanilla_model, bounds=(0., 1.), device="/GPU:0")
```
In order to test the robustness of the model, the first correctly classified element of each class are selected.
```
# strategy: first
# we select a sample from each class.
images_list = []
labels_list = []
# select only a few element from the test set
selected=np.random.choice(len(y_test_ord), 500)
sub_y_test_ord = y_test_ord[:300]
sub_x_test = x_test[:300]
# drop misclassified elements
misclassified_mask = tf.equal(tf.argmax(vanilla_model.predict(sub_x_test), axis=-1), sub_y_test_ord)
sub_x_test = sub_x_test[misclassified_mask]
sub_y_test_ord = sub_y_test_ord[misclassified_mask]
# now we will build a list with input image for each element of the matrix
for i in range(10):
# select the first element of the ith label
label_mask = [sub_y_test_ord==i]
x = sub_x_test[label_mask][0]
y = sub_y_test_ord[label_mask][0]
# convert it to tensor for use with foolbox
images = convert_to_tensor(x.astype("float32"), dtype="float32")
labels = convert_to_tensor(y, dtype="int64")
# repeat the input 10 times, one per misclassification target
images_list.append(images)
labels_list.append(labels)
images = convert_to_tensor(images_list)
labels = convert_to_tensor(labels_list)
```
In order to build a certficate, we take for each sample the top 2 output and apply this formula:
$$ \epsilon \geq \frac{\text{top}_1 - \text{top}_2}{2} $$
Where epsilon is the robustness radius for the considered sample.
```
values, classes = tf.math.top_k(hkr_fmodel(images), k=2)
certificates = (values[:, 0] - values[:, 1]) / 2
certificates
```
now we will attack the model to check if the certificates are respected. In this setup `L2CarliniWagnerAttack` is used but in practice as these kind of networks are gradient norm preserving, other attacks gives very similar results.
```
attack = fb.attacks.L2CarliniWagnerAttack(binary_search_steps=6, steps=8000)
imgs, advs, success = attack(hkr_fmodel, images, labels, epsilons=None)
dist_to_adv = np.sqrt(np.sum(np.square(images - advs), axis=(1,2,3)))
dist_to_adv
```
As we can see the certificate are respected.
```
tf.assert_less(certificates, dist_to_adv)
```
Finally we can take a visual look at the obtained examples.
We first start with utility functions for display.
```
class_mapping = {
0: "T-shirt/top",
1: "Trouser",
2: "Pullover",
3: "Dress",
4: "Coat",
5: "Sandal",
6: "Shirt",
7: "Sneaker",
8: "Bag",
9: "Ankle boot",
}
def adversarial_viz(model, images, advs, class_mapping):
"""
This functions shows for each sample:
- the original image
- the adversarial image
- the difference map
- the certificate and the observed distance to adversarial
"""
scale = 1.5
kwargs={}
nb_imgs = images.shape[0]
# compute certificates
values, classes = tf.math.top_k(model(images), k=2)
certificates = (values[:, 0] - values[:, 1]) / 2
# compute difference distance to adversarial
dist_to_adv = np.sqrt(np.sum(np.square(images - advs), axis=(1,2,3)))
# find classes labels for imgs and advs
orig_classes = [class_mapping[i] for i in tf.argmax(model(images), axis=-1).numpy()]
advs_classes = [class_mapping[i] for i in tf.argmax(model(advs), axis=-1).numpy()]
# compute differences maps
if images.shape[-1] != 3:
diff_pos = np.clip(advs - images, 0, 1.)
diff_neg = np.clip(images - advs, 0, 1.)
diff_map = np.concatenate([diff_neg, diff_pos, np.zeros_like(diff_neg)], axis=-1)
else:
diff_map = np.abs(advs - images)
# expands image to be displayed
if images.shape[-1] != 3:
images = np.repeat(images, 3, -1)
if advs.shape[-1] != 3:
advs = np.repeat(advs, 3, -1)
# create plot
figsize = (3 * scale, nb_imgs * scale)
fig, axes = plt.subplots(
ncols=3,
nrows=nb_imgs,
figsize=figsize,
squeeze=False,
constrained_layout=True,
**kwargs,
)
for i in range(nb_imgs):
ax = axes[i][0]
ax.set_title(orig_classes[i])
ax.set_xticks([])
ax.set_yticks([])
ax.axis("off")
ax.imshow(images[i])
ax = axes[i][1]
ax.set_title(advs_classes[i])
ax.set_xticks([])
ax.set_yticks([])
ax.axis("off")
ax.imshow(advs[i])
ax = axes[i][2]
ax.set_title(f"certif: {certificates[i]:.2f}, obs: {dist_to_adv[i]:.2f}")
ax.set_xticks([])
ax.set_yticks([])
ax.axis("off")
ax.imshow(diff_map[i]/diff_map[i].max())
```
When looking at the adversarial examples we can see that the network has interresting properties:
#### predictability
by looking at the certificates, we can predict if the adversarial example will be close of not
#### disparity among classes
As we can see, the attacks are very efficent on similar classes (eg. T-shirt/top, and Shirt ). This denote that all classes are not made equal regarding robustness.
#### explainability
The network is more explainable: attacks can be used as counterfactuals.
We can tell that removing the inscription on a T-shirt turns it into a shirt makes sense. Non robust examples reveals that the network rely on textures rather on shapes to make it's decision.
```
adversarial_viz(hkr_fmodel, images, advs, class_mapping)
```
| github_jupyter |
```
using Tensorflow;
using static Tensorflow.Binding;
using PlotNET;
using NumSharp;
int training_epochs = 1000;
// Parameters
float learning_rate = 0.01f;
int display_step = 50;
NumPyRandom rng = np.random;
NDArray train_X, train_Y;
int n_samples;
train_X = np.array(3.3f, 4.4f, 5.5f, 6.71f, 6.93f, 4.168f, 9.779f, 6.182f, 7.59f, 2.167f,
7.042f, 10.791f, 5.313f, 7.997f, 5.654f, 9.27f, 3.1f);
train_Y = np.array(1.7f, 2.76f, 2.09f, 3.19f, 1.694f, 1.573f, 3.366f, 2.596f, 2.53f, 1.221f,
2.827f, 3.465f, 1.65f, 2.904f, 2.42f, 2.94f, 1.3f);
n_samples = train_X.shape[0];
// tf Graph Input
var X = tf.placeholder(tf.float32);
var Y = tf.placeholder(tf.float32);
// Set model weights
// We can set a fixed init value in order to debug
// var rnd1 = rng.randn<float>();
// var rnd2 = rng.randn<float>();
var W = tf.Variable(-0.06f, name: "weight");
var b = tf.Variable(-0.73f, name: "bias");
// Construct a linear model
var pred = tf.add(tf.multiply(X, W), b);
// Mean squared error
var cost = tf.reduce_sum(tf.pow(pred - Y, 2.0f)) / (2.0f * n_samples);
// Gradient descent
// Note, minimize() knows to modify W and b because Variable objects are trainable=True by default
var optimizer = tf.train.GradientDescentOptimizer(learning_rate).minimize(cost);
// Initialize the variables (i.e. assign their default value)
var init = tf.global_variables_initializer();
// Start training
using (var sess = tf.Session())
{
// Run the initializer
sess.run(init);
// Fit all training data
for (int epoch = 0; epoch < training_epochs; epoch++)
{
foreach (var (x, y) in zip<float>(train_X, train_Y))
{
sess.run(optimizer,
new FeedItem(X, x),
new FeedItem(Y, y));
}
// Display logs per epoch step
if ((epoch + 1) % display_step == 0)
{
var c = sess.run(cost,
new FeedItem(X, train_X),
new FeedItem(Y, train_Y));
Console.WriteLine($"Epoch: {epoch + 1} cost={c} " + $"W={sess.run(W)} b={sess.run(b)}");
}
}
Console.WriteLine("Optimization Finished!");
var training_cost = sess.run(cost,
new FeedItem(X, train_X),
new FeedItem(Y, train_Y));
var plotter = new Plotter();
plotter.Plot(
train_X,
train_Y,
"Original data", ChartType.Scatter,"markers");
plotter.Plot(
train_X,
sess.run(W) * train_X + sess.run(b),
"Fitted line", ChartType.Scatter, "Fitted line");
plotter.Show();
// Testing example
var test_X = np.array(6.83f, 4.668f, 8.9f, 7.91f, 5.7f, 8.7f, 3.1f, 2.1f);
var test_Y = np.array(1.84f, 2.273f, 3.2f, 2.831f, 2.92f, 3.24f, 1.35f, 1.03f);
Console.WriteLine("Testing... (Mean square loss Comparison)");
var testing_cost = sess.run(tf.reduce_sum(tf.pow(pred - Y, 2.0f)) / (2.0f * test_X.shape[0]),
new FeedItem(X, test_X),
new FeedItem(Y, test_Y));
Console.WriteLine($"Testing cost={testing_cost}");
var diff = Math.Abs((float)training_cost - (float)testing_cost);
Console.WriteLine($"Absolute mean square loss difference: {diff}");
plotter.Plot(
test_X,
test_Y,
"Testing data", ChartType.Scatter, "markers");
plotter.Plot(
train_X,
sess.run(W) * train_X + sess.run(b),
"Fitted line", ChartType.Scatter);
plotter.Show();
return diff < 0.01;
}
```
| github_jupyter |
# Character-Level LSTM in PyTorch
In this notebook, I'll construct a character-level LSTM with PyTorch. The network will train character by character on some text, then generate new text character by character. As an example, I will train on Anna Karenina. **This model will be able to generate new text based on the text from the book!**
This network is based off of Andrej Karpathy's [post on RNNs](http://karpathy.github.io/2015/05/21/rnn-effectiveness/) and [implementation in Torch](https://github.com/karpathy/char-rnn). Below is the general architecture of the character-wise RNN.
<img src="assets/charseq.jpeg" width="500">
First let's load in our required resources for data loading and model creation.
```
import numpy as np
import torch
from torch import nn
import torch.nn.functional as F
```
## Load in Data
Then, we'll load the Anna Karenina text file and convert it into integers for our network to use.
### Tokenization
In the second cell, below, I'm creating a couple **dictionaries** to convert the characters to and from integers. Encoding the characters as integers makes it easier to use as input in the network.
```
# open text file and read in data as `text`
with open('data/anna.txt', 'r') as f:
text = f.read()
```
Now we have the text, encode it as integers.
```
# encode the text and map each character to an integer and vice versa
# we create two dictonaries:
# 1. int2char, which maps integers to characters
# 2. char2int, which maps characters to unique integers
chars = tuple(set(text))
int2char = dict(enumerate(chars))
char2int = {ch: ii for ii, ch in int2char.items()}
encoded = np.array([char2int[ch] for ch in text])
```
Let's check out the first 100 characters, make sure everything is peachy. According to the [American Book Review](http://americanbookreview.org/100bestlines.asp), this is the 6th best first line of a book ever.
```
text[:15]
```
And we can see those same characters encoded as integers.
```
encoded[:15]
```
## Pre-processing the data
As you can see in our char-RNN image above, our LSTM expects an input that is **one-hot encoded** meaning that each character is converted into an integer (via our created dictionary) and *then* converted into a column vector where only it's corresponsing integer index will have the value of 1 and the rest of the vector will be filled with 0's. Since we're one-hot encoding the data, let's make a function to do that!
```
def one_hot_encode(arr, n_labels):
# Initialize the the encoded array
one_hot = np.zeros((np.multiply(*arr.shape), n_labels), dtype=np.float32)
# Fill the appropriate elements with ones
one_hot[np.arange(one_hot.shape[0]), arr.flatten()] = 1.
# Finally reshape it to get back to the original array
one_hot = one_hot.reshape((*arr.shape, n_labels))
return one_hot
```
## Making training mini-batches
To train on this data, we also want to create mini-batches for training. Remember that we want our batches to be multiple sequences of some desired number of sequence steps. Considering a simple example, our batches would look like this:
<img src="assets/sequence_batching@1x.png" width=500px>
<br>
In this example, we'll take the encoded characters (passed in as the `arr` parameter) and split them into multiple sequences, given by `n_seqs` (also refered to as "batch size" in other places). Each of those sequences will be `n_steps` long.
### Creating Batches
**1. The first thing we need to do is discard some of the text so we only have completely full batches. **
Each batch contains $N \times M$ characters, where $N$ is the batch size (the number of sequences) and $M$ is the number of steps. Then, to get the total number of batches, $K$, we can make from the array `arr`, you divide the length of `arr` by the number of characters per batch. Once you know the number of batches, you can get the total number of characters to keep from `arr`, $N * M * K$.
**2. After that, we need to split `arr` into $N$ sequences. **
You can do this using `arr.reshape(size)` where `size` is a tuple containing the dimensions sizes of the reshaped array. We know we want $N$ sequences, so let's make that the size of the first dimension. For the second dimension, you can use `-1` as a placeholder in the size, it'll fill up the array with the appropriate data for you. After this, you should have an array that is $N \times (M * K)$.
**3. Now that we have this array, we can iterate through it to get our batches. **
The idea is each batch is a $N \times M$ window on the $N \times (M * K)$ array. For each subsequent batch, the window moves over by `n_steps`. We also want to create both the input and target arrays. Remember that the targets are the inputs shifted over one character. The way I like to do this window is use `range` to take steps of size `n_steps` from $0$ to `arr.shape[1]`, the total number of steps in each sequence. That way, the integers you get from `range` always point to the start of a batch, and each window is `n_steps` wide.
> **TODO:** Write the code for creating batches in the function below. The exercises in this notebook _will not be easy_. I've provided a notebook with solutions alongside this notebook. If you get stuck, checkout the solutions. The most important thing is that you don't copy and paste the code into here, **type out the solution code yourself.**
```
def get_batches(arr, n_seqs, n_steps):
'''Create a generator that returns batches of size
n_seqs x n_steps from arr.
Arguments
---------
arr: Array you want to make batches from
n_seqs: Batch size, the number of sequences per batch
n_steps: Number of sequence steps per batch
'''
# Get the number of characters per batch
batch_size = n_seqs #* n_steps
## TODO: Get the number of batches we can make
n_batches = len(arr)//(batch_size* n_steps)
## TODO: Keep only enough characters to make full batches
arr = arr[:n_batches*batch_size*n_steps]
## TODO: Reshape into batch_size rows
arr = arr.reshape( (batch_size,n_batches*n_steps) )
print(arr.shape)
## TODO: Make batches
for n in range(0, arr.shape[1], n_steps):
# The features
print(n)
x = arr[:,n*n_steps:(n+1)*n_steps]
# The targets, shifted by one
y = arr[:,n*n_steps+1:(n+1)*n_steps+1]
yield x, y
```
### Test Your Implementation
Now I'll make some data sets and we can check out what's going on as we batch data. Here, as an example, I'm going to use a batch size of 10 and 50 sequence steps.
```
print(len(encoded))
batches = get_batches(encoded, 10, 50)
x, y = next(batches)
print('x\n', x[:10, :10])
print('\ny\n', y[:10, :10])
```
If you implemented `get_batches` correctly, the above output should look something like
```
x
[[55 63 69 22 6 76 45 5 16 35]
[ 5 69 1 5 12 52 6 5 56 52]
[48 29 12 61 35 35 8 64 76 78]
[12 5 24 39 45 29 12 56 5 63]
[ 5 29 6 5 29 78 28 5 78 29]
[ 5 13 6 5 36 69 78 35 52 12]
[63 76 12 5 18 52 1 76 5 58]
[34 5 73 39 6 5 12 52 36 5]
[ 6 5 29 78 12 79 6 61 5 59]
[ 5 78 69 29 24 5 6 52 5 63]]
y
[[63 69 22 6 76 45 5 16 35 35]
[69 1 5 12 52 6 5 56 52 29]
[29 12 61 35 35 8 64 76 78 28]
[ 5 24 39 45 29 12 56 5 63 29]
[29 6 5 29 78 28 5 78 29 45]
[13 6 5 36 69 78 35 52 12 43]
[76 12 5 18 52 1 76 5 58 52]
[ 5 73 39 6 5 12 52 36 5 78]
[ 5 29 78 12 79 6 61 5 59 63]
[78 69 29 24 5 6 52 5 63 76]]
```
although the exact numbers will be different. Check to make sure the data is shifted over one step for `y`.
---
## Defining the network with PyTorch
Below is where you'll define the network. We'll break it up into parts so it's easier to reason about each bit. Then we can connect them up into the whole network.
<img src="assets/charRNN.png" width=500px>
Next, you'll use PyTorch to define the architecture of the network. We start by defining the layers and operations we want. Then, define a method for the forward pass. You've also been given a method for predicting characters.
### Model Structure
In `__init__` the suggested structure is as follows:
* Create and store the necessary dictionaries (this has been done for you)
* Define an LSTM layer that takes as params: an input size (the number of characters), a hidden layer size `n_hidden`, a number of layers `n_layers`, a dropout probability `drop_prob`, and a batch_first boolean (True, since we are batching)
* Define a dropout layer with `dropout_prob`
* Define a fully-connected layer with params: input size `n_hidden` and output size (the number of characters)
* Finally, initialize the weights (again, this has been given)
Note that some parameters have been named and given in the `__init__` function, and we use them and store them by doing something like `self.drop_prob = drop_prob`.
---
### LSTM Inputs/Outputs
You can create a basic LSTM cell as follows
```python
self.lstm = nn.LSTM(input_size, n_hidden, n_layers,
dropout=drop_prob, batch_first=True)
```
where `input_size` is the number of characters this cell expects to see as sequential input, and `n_hidden` is the number of units in the hidden layers in the cell. And we can add dropout by adding a dropout parameter with a specified probability; this will automatically add dropout to the inputs or outputs. Finally, in the `forward` function, we can stack up the LSTM cells into layers using `.view`. With this, you pass in a list of cells and it will send the output of one cell into the next cell.
We also need to create an initial cell state of all zeros. This is done like so
```python
self.init_weights()
```
```
class CharRNN(nn.Module):
def __init__(self, tokens, n_steps=100, n_hidden=256, n_layers=2,
drop_prob=0.5, lr=0.001):
super().__init__()
self.drop_prob = drop_prob
self.n_layers = n_layers
self.n_hidden = n_hidden
self.lr = lr
# creating character dictionaries
self.chars = tokens
self.int2char = dict(enumerate(self.chars))
self.char2int = {ch: ii for ii, ch in self.int2char.items()}
## TODO: define the LSTM, self.lstm
input_size = len(self.chars)
self.lstm = nn.LSTM(input_size, self.n_hidden, self.n_layers,
dropout=self.drop_prob, batch_first=True)
## TODO: define a dropout layer, self.dropout
self.fc1_drop = nn.Dropout(p=self.drop_prob)
## TODO: define the final, fully-connected output layer, self.fc
self.fc = nn.Linear(self.n_hidden, input_size)
# initialize the weights
self.init_weights()
def forward(self, x, hc):
''' Forward pass through the network.
These inputs are x, and the hidden/cell state `hc`. '''
#h, c = hc
## TODO: Get x, and the new hidden state (h, c) from the lstm
x, out = self.lstm(x, hc)
print(x.size())
h,c = out
## TODO: pass x through a droupout layer
x = self.fc1_drop(x)
# Stack up LSTM outputs using view
print('x_size =', x.size())
x = x.view(x.size()[0]*x.size()[1], self.n_hidden)
print('x_size =', x.size())
## TODO: put x through the fully-connected layer
x = self.fc(x)
# return x and the hidden state (h, c)
return x, (h, c)
def predict(self, char, h=None, cuda=False, top_k=None):
''' Given a character, predict the next character.
Returns the predicted character and the hidden state.
'''
if cuda:
self.cuda()
else:
self.cpu()
if h is None:
h = self.init_hidden(1)
x = np.array([[self.char2int[char]]])
x = one_hot_encode(x, len(self.chars))
inputs = torch.from_numpy(x)
if cuda:
inputs = inputs.cuda()
h = tuple([each.data for each in h])
out, h = self.forward(inputs, h)
p = F.softmax(out, dim=1).data
if cuda:
p = p.cpu()
if top_k is None:
top_ch = np.arange(len(self.chars))
else:
p, top_ch = p.topk(top_k)
top_ch = top_ch.numpy().squeeze()
p = p.numpy().squeeze()
char = np.random.choice(top_ch, p=p/p.sum())
return self.int2char[char], h
def init_weights(self):
''' Initialize weights for fully connected layer '''
initrange = 0.1
# Set bias tensor to all zeros
self.fc.bias.data.fill_(0)
# FC weights as random uniform
self.fc.weight.data.uniform_(-1, 1)
def init_hidden(self, n_seqs):
''' Initializes hidden state '''
# Create two new tensors with sizes n_layers x n_seqs x n_hidden,
# initialized to zero, for hidden state and cell state of LSTM
weight = next(self.parameters()).data
return (weight.new(self.n_layers, n_seqs, self.n_hidden).zero_(),
weight.new(self.n_layers, n_seqs, self.n_hidden).zero_())
```
### A note on the `predict` function
The output of our RNN is from a fully-connected layer and it outputs a **distribution of next-character scores**.
To actually get the next character, we apply a softmax function, which gives us a *probability* distribution that we can then sample to predict the next character.
```
## ---- keep notebook from crashing during training --- ##
import os
import requests
import time
def train(net, data, epochs=10, n_seqs=10, n_steps=50, lr=0.001, clip=5, val_frac=0.1, cuda=False, print_every=10):
''' Training a network
Arguments
---------
net: CharRNN network
data: text data to train the network
epochs: Number of epochs to train
n_seqs: Number of mini-sequences per mini-batch, aka batch size
n_steps: Number of character steps per mini-batch
lr: learning rate
clip: gradient clipping
val_frac: Fraction of data to hold out for validation
cuda: Train with CUDA on a GPU
print_every: Number of steps for printing training and validation loss
'''
net.train()
opt = torch.optim.Adam(net.parameters(), lr=lr)
criterion = nn.CrossEntropyLoss()
# create training and validation data
val_idx = int(len(data)*(1-val_frac))
data, val_data = data[:val_idx], data[val_idx:]
if cuda:
net.cuda()
counter = 0
n_chars = len(net.chars)
old_time = time.time()
for e in range(epochs):
h = net.init_hidden(n_seqs)
for x, y in get_batches(data, n_seqs, n_steps):
if time.time() - old_time > 60:
old_time = time.time()
requests.request("POST",
"https://nebula.udacity.com/api/v1/remote/keep-alive",
headers={'Authorization': "STAR " + response.text})
counter += 1
# One-hot encode our data and make them Torch tensors
x = one_hot_encode(x, n_chars)
inputs, targets = torch.from_numpy(x), torch.from_numpy(y)
if cuda:
inputs, targets = inputs.cuda(), targets.cuda()
# Creating new variables for the hidden state, otherwise
# we'd backprop through the entire training history
h = tuple([each.data for each in h])
net.zero_grad()
output, h = net.forward(inputs, h)
loss = criterion(output, targets.view(n_seqs*n_steps))
loss.backward()
# `clip_grad_norm` helps prevent the exploding gradient problem in RNNs / LSTMs.
nn.utils.clip_grad_norm_(net.parameters(), clip)
opt.step()
if counter % print_every == 0:
# Get validation loss
val_h = net.init_hidden(n_seqs)
val_losses = []
for x, y in get_batches(val_data, n_seqs, n_steps):
# One-hot encode our data and make them Torch tensors
x = one_hot_encode(x, n_chars)
x, y = torch.from_numpy(x), torch.from_numpy(y)
# Creating new variables for the hidden state, otherwise
# we'd backprop through the entire training history
val_h = tuple([each.data for each in val_h])
inputs, targets = x, y
if cuda:
inputs, targets = inputs.cuda(), targets.cuda()
output, val_h = net.forward(inputs, val_h)
val_loss = criterion(output, targets.view(n_seqs*n_steps))
val_losses.append(val_loss.item())
print("Epoch: {}/{}...".format(e+1, epochs),
"Step: {}...".format(counter),
"Loss: {:.4f}...".format(loss.item()),
"Val Loss: {:.4f}".format(np.mean(val_losses)))
```
## Time to train
Now we can actually train the network. First we'll create the network itself, with some given hyperparameters. Then, define the mini-batches sizes (number of sequences and number of steps), and start the training. With the train function, we can set the number of epochs, the learning rate, and other parameters. Also, we can run the training on a GPU by setting `cuda=True`.
```
if 'net' in locals():
del net
# define and print the net
net = CharRNN(chars, n_hidden=512, n_layers=2)
print(net)
n_seqs, n_steps = 128, 100
# you may change cuda to True if you plan on using a GPU!
# also, if you do, please INCREASE the epochs to 25
# Open the training log file.
log_file = 'training_log.txt'
f = open(log_file, 'w')
response = requests.request("GET",
"http://metadata.google.internal/computeMetadata/v1/instance/attributes/keep_alive_token",
headers={"Metadata-Flavor":"Google"})
# TRAIN
train(net, encoded, epochs=1, n_seqs=n_seqs, n_steps=n_steps, lr=0.001, cuda=False, print_every=10)
# Close the training log file.
f.close()
```
## Getting the best model
To set your hyperparameters to get the best performance, you'll want to watch the training and validation losses. If your training loss is much lower than the validation loss, you're overfitting. Increase regularization (more dropout) or use a smaller network. If the training and validation losses are close, you're underfitting so you can increase the size of the network.
## Hyperparameters
Here are the hyperparameters for the network.
In defining the model:
* `n_hidden` - The number of units in the hidden layers.
* `n_layers` - Number of hidden LSTM layers to use.
We assume that dropout probability and learning rate will be kept at the default, in this example.
And in training:
* `n_seqs` - Number of sequences running through the network in one pass.
* `n_steps` - Number of characters in the sequence the network is trained on. Larger is better typically, the network will learn more long range dependencies. But it takes longer to train. 100 is typically a good number here.
* `lr` - Learning rate for training
Here's some good advice from Andrej Karpathy on training the network. I'm going to copy it in here for your benefit, but also link to [where it originally came from](https://github.com/karpathy/char-rnn#tips-and-tricks).
> ## Tips and Tricks
>### Monitoring Validation Loss vs. Training Loss
>If you're somewhat new to Machine Learning or Neural Networks it can take a bit of expertise to get good models. The most important quantity to keep track of is the difference between your training loss (printed during training) and the validation loss (printed once in a while when the RNN is run on the validation data (by default every 1000 iterations)). In particular:
> - If your training loss is much lower than validation loss then this means the network might be **overfitting**. Solutions to this are to decrease your network size, or to increase dropout. For example you could try dropout of 0.5 and so on.
> - If your training/validation loss are about equal then your model is **underfitting**. Increase the size of your model (either number of layers or the raw number of neurons per layer)
> ### Approximate number of parameters
> The two most important parameters that control the model are `n_hidden` and `n_layers`. I would advise that you always use `n_layers` of either 2/3. The `n_hidden` can be adjusted based on how much data you have. The two important quantities to keep track of here are:
> - The number of parameters in your model. This is printed when you start training.
> - The size of your dataset. 1MB file is approximately 1 million characters.
>These two should be about the same order of magnitude. It's a little tricky to tell. Here are some examples:
> - I have a 100MB dataset and I'm using the default parameter settings (which currently print 150K parameters). My data size is significantly larger (100 mil >> 0.15 mil), so I expect to heavily underfit. I am thinking I can comfortably afford to make `n_hidden` larger.
> - I have a 10MB dataset and running a 10 million parameter model. I'm slightly nervous and I'm carefully monitoring my validation loss. If it's larger than my training loss then I may want to try to increase dropout a bit and see if that helps the validation loss.
> ### Best models strategy
>The winning strategy to obtaining very good models (if you have the compute time) is to always err on making the network larger (as large as you're willing to wait for it to compute) and then try different dropout values (between 0,1). Whatever model has the best validation performance (the loss, written in the checkpoint filename, low is good) is the one you should use in the end.
>It is very common in deep learning to run many different models with many different hyperparameter settings, and in the end take whatever checkpoint gave the best validation performance.
>By the way, the size of your training and validation splits are also parameters. Make sure you have a decent amount of data in your validation set or otherwise the validation performance will be noisy and not very informative.
After training, we'll save the model so we can load it again later if we need too. Here I'm saving the parameters needed to create the same architecture, the hidden layer hyperparameters and the text characters.
```
# change the name, for saving multiple files
model_name = 'rnn_local_epoch.net'
checkpoint = {'n_hidden': net.n_hidden,
'n_layers': net.n_layers,
'state_dict': net.state_dict(),
'tokens': net.chars}
with open(model_name, 'wb') as f:
torch.save(checkpoint, f)
```
## Sampling
Now that the model is trained, we'll want to sample from it. To sample, we pass in a character and have the network predict the next character. Then we take that character, pass it back in, and get another predicted character. Just keep doing this and you'll generate a bunch of text!
### Top K sampling
Our predictions come from a categorcial probability distribution over all the possible characters. We can make the sample text and make it more reasonable to handle (with less variables) by only considering some $K$ most probable characters. This will prevent the network from giving us completely absurd characters while allowing it to introduce some noise and randomness into the sampled text.
Typically you'll want to prime the network so you can build up a hidden state. Otherwise the network will start out generating characters at random. In general the first bunch of characters will be a little rough since it hasn't built up a long history of characters to predict from.
```
def sample(net, size, prime='The', top_k=None, cuda=False):
if cuda:
net.cuda()
else:
net.cpu()
net.eval()
# First off, run through the prime characters
chars = [ch for ch in prime]
h = net.init_hidden(1)
for ch in prime:
char, h = net.predict(ch, h, cuda=cuda, top_k=top_k)
chars.append(char)
# Now pass in the previous character and get a new one
for ii in range(size):
char, h = net.predict(chars[-1], h, cuda=cuda, top_k=top_k)
chars.append(char)
return ''.join(chars)
print(sample(net, 2000, prime='Anna', top_k=5, cuda=False))
```
## Loading a checkpoint
```
# Here we have loaded in a model that trained over 1 epoch `rnn_1_epoch.net`
with open('rnn_1_epoch.net', 'rb') as f:
checkpoint = torch.load(f)
loaded = CharRNN(checkpoint['tokens'], n_hidden=checkpoint['n_hidden'], n_layers=checkpoint['n_layers'])
loaded.load_state_dict(checkpoint['state_dict'])
# Change cuda to True if you are using GPU!
print(sample(loaded, 2000, cuda=False, top_k=5, prime="And Levin said"))
```
| github_jupyter |
# Hiddon Markov Models
**Author: Carleton Smith**
**Reviewer: Jessica Cervi**
**Expected time = 1.5 hours**
**Total points = 45**
## Assignment Overview
In this assignment, we will work on Hidden Markov Models (HMMs). HMMs can be effective in solving problems in reinforcement learning and temporal pattern recognition.
In this assignment, we will:
- Use clustering
- Decide on the type of HMM problem
- Initialize and tune HMM parameters
The implementation of HMM from scratch may involve complex code that might be difficult to comprehend and is not the most robust and efficient.
In this assignment, we:
- Prioritize the demonstration of the algorithm's steps while maintaining reproducibility
- Define and implement certain tasks and non-essential features of the algorithm (example: 'Signature' and 'Parameter' classes) without explanation.
We are not expecting students to understand the entirety of the codebase. Rather, we will test students on their high-level understanding of HMM and some critical functions.
We have designed this assignment to build your familiarity and comfort in coding in Python. It will also help you review the key topics from each module. As you progress through the assignment, answers to the questions will get increasingly complex. You must adopt a data scientist's mindset when completing this assignment. Remember to run your code from each cell before submitting your assignment. Running your code beforehand will notify you of errors and giving you a chance to fix your errors before submitting it. You should view your Vocareum submission as if you are delivering a final project to your manager or client.
***Vocareum Tips***
- Do not add arguments or options to functions unless asked specifically. This will cause an error in Vocareum.
- Do not use a library unless you are explicitly asked in the question.
- You can download the Grading Report after submitting the assignment. It will include the feedback and hints on incorrect questions.
### Learning Objectives
- Describe the three problems that HMMs can solve
- Explain HMM parameters
- Predict using a trained HMM
- Explain the components of discreet HMM
- Implement k-means clustering using sklearn
- Build state transition and emmision matrices for HMM
- Explain forward/backward algorithm to solve HMM
- Explain vitervi algorithm to solve HMM
- Interpret results of forward/backward algorithm to solve HMM
## Index:
#### Hiddon Markov Models
+ [Question 01](#q01)
+ [Question 02](#q02)
+ [Question 03](#q03)
+ [Question 04](#q04)
+ [Question 05](#q05)
+ [Question 06](#q06)
+ [Question 07](#q07)
## Hiddon Markov Models
### Importing the dataset
Throughout this assignment, we will try to predict the weekly price of corn from a dataset taken from Kaggle.
The dataset can be downloaded using [_this link_](https://www.kaggle.com/nickwong64/corn2015-2017) from the Kaggle.
We will begin by importing the libraries that we will use in this assignment.
This dataset file composed of simply two columns. The first column lists the date of the final day of each week and the second column lists corn close price. The timeframe goes from 2015-01-04 until 2017-10-01. The original data is downloaded from Quantopian corn futures price.
Next, we will use the 'pandas' function 'read_csv' to read the dataset. Finally, we will display a sample of the data taken from Amazon.
```
# Import libraries
import numpy as np
import pandas as pd
from collections import namedtuple
from itertools import permutations, chain, product
from sklearn.cluster import KMeans
from inspect import Signature, Parameter
```
Below we read the data using the Pandas' function 'read_csv'. Also, we rename the columns in each dataset for convenience.
```
corn13_17 = pd.read_csv('./data/corn2013-2017.txt', names = ("week","price") )
```
Below, we display the first 5 columns of our dataset.
```
corn13_17.head()
```
Next, we inspect the data for missing values.
```
corn13_17.info()
```
From the output above, the data appear to be complete.
### Preprocess: Discretization
As mentioned in the lectures, there are two types of HMM:
- 1. Discrete HMM
- 2. Continuous HMM
The typical structure of HMM involves a discrete number of latent ("hidden") states that are unobserved. The observations, which in our case are corn prices, are generated from a state dependent "emissions" (or "observations") distribution.
In the discrete HMM, the emissions are discrete values. Conversely, in the continuous HMM, the emissions are continuous. In the latter, the distribution that generates the emissions is usually assumed to be Gaussian.
<a id="q01"></a>
[Return to top](#questions)
### Question 01
*5 points*
Decide wheter the following statement is true or false.
*The number of discrete states is a hyperparameter of HMM.*
Assign a boolean value to the variable ans1.
```
### GRADED
### YOUR SOLUTION HERE
ans1 = None
###
### YOUR CODE HERE
###
###
### AUTOGRADER TEST - DO NOT REMOVE
###
```
In order to simplify the problem, in come cases, it is advisable to use clustering in order to discretize the continuous HMM emissions.
We will see how to do this in the next section.
### Generate Clusters
As noted in the lectures, clustering a sequence of continuous observations is a form of data quantization. This can simplify the learning of HMM.
Instead of calculating A posteriori probability from a continuous emissions sequence, we can use the respective cluster label as the observation. This way, the emission probability matrix can be encoded as a discrete vector of probabilities.
<a id="q02"></a>
[Return to top](#questions)
### Question 02
*10 points*
Define a function called 'generate_cluster_assignments' that accepts the following arguments:
- A 'Pandas' series.
- The desired number of cluster.
Your function should instantiate an 'sklearn' KMeans class using the specified number of clusters and random_state equals to 24. Next, your function should transform the series to a dataframe. Finally, your function should return a 'Pandas' series of cluster labels for each observation in the sequence.
*Hint*: That KMeans object has the '.fit()' and '.predict()' methods to create the cluster labels.
For example, if we set
`data_series = pd.Series([1,2,3,2,1,2,3,2,1,2,3,2,1,2,3,2,1,6,7,8,7,6,7,8,6,7,6,7,8,7,7,8,56,57,58,59,57,58,6,7,8,1,2])`
Then calling the function by using
`labels = generate_cluster_assignments(data_series, clusters = 3)`
should return
`labels = array([1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 2, 2, 2, 2, 2,2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 0, 0, 0, 0, 0, 0, 2, 2, 2, 1, 1])`
Note that the KMeans object can be instantiated via the following command:
'clusterer = KMeans(args)'
Moreover, your particular labels might not match exactly, but the **clusters** should be the same.
```
### GRADED
### YOUR SOLUTION HERE
def generate_cluster_assignments(series, clusters):
return
###
### YOUR CODE HERE
###
###
### AUTOGRADER TEST - DO NOT REMOVE
###
```
<a id="part2"></a>
## Part 2: Components of a Hidden Markov Model
### HMM Parameters
A HMM consists of 5 components:
- N -- The number of hidden states: This is a discrete integer that the practitioner provides. It is the number of assumed hidden states.
- A -- State transition matrix: With N=2, A may look like:
$$ A = \begin{bmatrix}
. & S1 & S2 \\\
S1 & .7 & .3 \\\
S2 & .6 & .4
\end{bmatrix}$$
The second row of A shows the probability of transitioning from state 1 --> (state 1=0.7, state 2=0.3).
The third row of A shows the probability of transitioning from state 2 --> (state 1=0.6, state 2=0.4).
- B -- Emission probability matrix: Setting the number of unique observation M==4, B may look like:
$$ B = \begin{bmatrix}
. &a & b & c & d \\\
S1 & .4 & .3 & .1 & .2 \\\
S2 & .1 & .4 & .1 & .4
\end{bmatrix}$$
The rows correspond to state 1 and state 2, respectively. The columns in B correspond to the probability of that observation, given the respective state.
For example, the probability of observing $d$ in $S1$ = $0.2$.
- $\pi$ -- Starting likelihood (initial state probabilities). $\pi$ may take the following form:
$$\pi = \begin{bmatrix}
.5 \\
.5
\end{bmatrix}$$
In this case, it means that the sequence of states is equally likely to start in $S1$ or $S2$.
- $(x_{i}, ..., x_{T})$ -- Sequence of emissions or observations
The sequence of observations, $(x_{i}, ..., x_{T})$ is always the known component of HMM. We can determine the type of HMM problem by the known components and the motivation of the problem. Next, we will discuss the types of HMM problems.
<a id="q03"></a>
[Return to top](#questions)
### Question 03
*5 points*
If "N" is the number of states, then the shape of "A" (the transition matrix) will always be:
- a) 2 x 2
- b) N x N
- c) N x number of unique observations
- d) None of the above
Assign the character associated with your choice as a string to the variable 'ans3'.
```
### GRADED
### YOUR SOLUTION HERE
ans3 = None
###
### YOUR CODE HERE
###
###
### AUTOGRADER TEST - DO NOT REMOVE
###
```
<a id="q04"></a>
[Return to top](#questions)
### Question 4
*5 points*
If "N" is the number of states, then the shape of "B" (the emission matrix) will always be:
- a) 2 x 4
- b) N x N
- c) N x number of unique observations
- d) None of the above
Assign the character associated with your choice as a string to the variable ans4.
```
### GRADED
### YOUR SOLUTION HERE
ans4 = None
###
### YOUR CODE HERE
###
###
### AUTOGRADER TEST - DO NOT REMOVE
###
```
### Types of Problems using HMM
In Lecture 11-2, we described three HMM estimation problems:
1. State Estimation
2. State Sequence
3. Learn an HMM
Below, we will briefly cover the motivation of each estimation problem.
**1. State Estimation**:
- For the given HMM ($\pi$, _A_, *B*) and the observation sequence $(x_{i}, ..., x_{T})$, estimate the state probability for $x_{i}$.
**2. State Sequence**:
- For the given HMM ($\pi$, _A_, *B*) and the observation sequence $(x_{i}, ..., x_{T})$, estimate the most probable state sequence.
**3. Learn an HMM**:
- For the given observation sequence $(x_{i}, ..., x_{T})$, estimate the HMM parameters ($\pi$, _A_, *B*).
<a id="q05"></a>
[Return to top](#questions)
### Question 5
*5 points*
Which of the following HMM problems uses the Forward-Backward Algorithm to estimate the solution?
- a) State Estimation
- b) State Sequence
- c) Learn an HMM
- d) None of the above
- e) All of the above
List all the answers that apply as 'a', 'b', 'c', 'd', and/or 'e' in the list assigned to the variable `ans5`.
```
### GRADED
### YOUR SOLUTION HERE
ans5 = None
###
### YOUR CODE HERE
###
###
### AUTOGRADER TEST - DO NOT REMOVE
###
```
<a id="q06"></a>
[Return to top](#questions)
### Question 06
*5 points*
Which of the following HMM problems uses the Viterbi Algorithm to estimate the solution?
- a) State Estimation
- b) State Sequence
- c) Learn HMM
- d) None of the above
- e) All of the above
List all the answers that apply as 'a', 'b', 'c', 'd', and/or 'e' in the list assigned to the variable `ans6`.
```
### GRADED
### YOUR SOLUTION HERE
ans6 = None
###
### YOUR CODE HERE
###
###
### AUTOGRADER TEST - DO NOT REMOVE
###
```
<a id="part3"></a>
## Part 3: Estimating a Hidden Markov Model
In this section, you will be guided through an exercise using the third HMM estimation problem: **Learn an HMM**
We will use the the Corn Prices 2013-2017 dataset from Kaggle as our sequence of observations.
In Question 02 we asked you to make a function to discretize a Pandas Series into a specified number of clusters. We will now use this function to discretize our price data into 5 clusters.
```
# Cluster 2013-2017
corn13_17_seq = generate_cluster_assignments(corn13_17[['price']], 5)
corn13_17_seq
```
**NOTE:** There are quite a few functions provided in the following cells. It is not imperative that you completely understand how each of them works. Many of them are helper functions that perform a specific task within another function. You must understand the procedure that is occuring to estimate the HMM parameters. These steps are laid out in the next section.
### Steps for Learning a HMM
The Expectation Maximization (EM) algorithm is used to estimate the parameters of HMM given a sequence of observations (or emissions). Here are the general steps for estimating the parameters in HMM:
1. Initialize a set of parameters for HMM ($\pi$, _A_, *B*)
2. Conduct the EM algorithm:
- The E Step: Use forward-backward algorithm to calculate the probability of observing the emissions with the given HMM parameters ($\pi$, _A_, *B*)
- The M Step: Update the HMM parameters so that the sequence of observations are more likely to have come from this particular HMM
3. Repeat steps 1 and 2 until the HMM parameters have converged.
<br>
The remaining parts of this assignment will perform this procedure.
### Important Constant: Number of Unique States
```
# Almost all the functions require this constant as an argument
STATE_LIST = ['S1', 'S2']
# Initialze the state transition probabilities (2 states)
STATE_TRANS_PROBS = [0.4, 0.6, 0.35, 0.55]
```
### Helper Functions
These functions are used to perform the common tasks.
```
# given a list with unique items, this function will return a new list with all permutations
def make_state_permutations(list_of_unique_states):
l1 = [''.join(tup) for tup in permutations(list_of_unique_states, 2)]
l2 = [state+state for state in list_of_unique_states]
return sorted(l1 + l2)
# helper function in EM function
def _grab_highest_prob_and_state(state_permutations_lst, prob_arr):
return (prob_arr[np.argmax(prob_arr)], state_permutations_lst[np.argmax(prob_arr)])
```
**The following two functions transform a dictionary to a different format.**
```
def dict_to_tuples(list_of_unique_states, d):
"""
list_of_unique_states: List of unique state names, as strings
d: Dictionary of state transition probabilities
EXAMPLE:
s_perms = ['S1S1', 'S1S2', 'S2S1', 'S2S2']
p_list = [0.1, 0.9, 0.4, 0.6]
d = {'S1S1': 0.1, 'S1S2': 0.9, 'S2S1': 0.4, 'S2S2': 0.6}
print(dict_to_tuples(d))
OUTPUT:
{S1: (0.1, 0.9), S2: (0.4, 0.6)}
"""
# Defensive programming to ensure output will be correct
list_of_unique_states = sorted(list_of_unique_states)
assert make_state_permutations(list_of_unique_states) == list(d.keys()), \
"Keys of dictionary must match output of `make_state_permutations(list_of_unique_states)`"
lengths = [len(st) for st in list_of_unique_states]
final_dict = {}
for idx, st in enumerate(list_of_unique_states):
tup = []
for trans_p in d.keys():
if trans_p[:lengths[idx]] == st:
tup.append(d[trans_p])
else:
continue
final_dict[st] = tuple(tup)
return final_dict
def obs_to_tuples(list_of_unique_states, d, sequence):
"""
list_of_unique_states: List of unique state names, as strings
d: Dictionary of obs transition probabilities
sequence: the observation sequence
EXAMPLE:
STATE_LIST = ['S1', 'S2']
d = {'S1_0': 0.1,
'S1_1': 0.3,
'S1_2': 0.4,
'S1_3': 0.15,
'S1_4': 0.05,
'S2_0': 0.15,
'S2_1': 0.2,
'S2_2': 0.3,
'S2_3': 0.05,
'S2_4': 0.3}
corn15_17_seq = generate_cluster_assignments(corn15_17[['price']], 5)
print(obs_to_tuples(STATE_LIST, d))
OUTPUT:
{'S1': (0.1, 0.3, 0.4, 0.15, 0.05), 'S2': (0.15, 0.2, 0.3, 0.05, 0.3)}
"""
# Defensive programming to ensure output will be correct
list_of_unique_states = sorted(list_of_unique_states)
num_unique_obs = len(np.unique(sequence))
lengths = [len(st) for st in list_of_unique_states]
final_dict = {}
for idx, st in enumerate(list_of_unique_states):
tup = []
for e_trans in d.keys():
if e_trans[:lengths[idx]] == st:
tup.append(d[e_trans])
else:
continue
final_dict[st] = tuple(tup)
return final_dict
```
### Define Transition Functions
We will use the following three definitions of functions (all starting with 'generate') to create our initial HMM parameters ($\pi$, $A$, $B$) in the form of a dictionary.
If no values are explicitly given in the '**kwargs' argument**, the functions are flexible enough to create ($\pi$, $A$, $B$) from user-specified values or to provide default uniform probability vectors.
#### An Aside: Why dictionaries and not arrays?
In this exercise, for data structure, we use dictionaries instead of arrays to improve efficiency and accuracy. The functions defined for this procedure frequently involve retrieving values from data structures and altering existing values. We can achieve this using a dictionary rather than an array because dictionaries utilize hashing functions to lookup data instead of indexed positions.
To demonstrate the speed benefits of using dictionaries to retrieve values, consider the time needed to retrieve a piece of data from both an array and a dictionary in the following exercise.
```
import timeit
all_setup = """
import numpy as np
# These hold the same information
arr = np.array([[0.4, 0.6], [0.35, 0.55]])
d = {'S1S1': 0.4, 'S1S2': 0.6, 'S2S1': 0.35, 'S2S2': 0.55}
"""
i = 10_000_000
index_an_array = 'arr[0,0]'
retrieve_value = "d['S1S1']"
print('Seconds to index an array {} times: {}'.format(
i, timeit.timeit(setup=all_setup, stmt=index_an_array, number=i)))
print('\n','#' * 60, '\n')
print('Seconds to retrieve value {} times: {}'.format(i, timeit.timeit(setup=all_setup, stmt=retrieve_value, number=i)))
```
**The following functions generate initial probabilities for ($\pi$, $A$, $B$)**
```
def generate_state_trans_dict(list_of_unique_states, **kwargs):
'''
'list_of_unique_states': list of states as strings
''**kwargs': keyword being the state and value is tuple of state transitions.
<Must be listed in same order as listed in 'list_of_unique_states'>
If **kwargs omitted, transitions are given uniform distribution based on
number of states.
EXAMPLE1:
state_params = generate_state_trans_dict(['S1', 'S2', 'S3'])
OUTPUT1:
{'S1S1': 0.5, 'S2S2': 0.5, 'S1S2': 0.5, 'S2S1': 0.5}
EXAMPLE2:
state_params = generate_state_trans_dict(['S1', 'S2'], S1=(0.1, 0.9), S2=(0.4, 0.6))
OUTPUT2:
{'S1S1': 0.1, 'S1S2': 0.9, 'S2S1': 0.4, 'S2S2': 0.6}
'''
# number of states
N = len(list_of_unique_states)
# this runs if specific transitions are provided
if kwargs:
state_perms = [''.join(tup) for tup in permutations(list(kwargs.keys()), 2)]
all_permutations = [state+state for state in list_of_unique_states] + state_perms
pbs = chain.from_iterable(kwargs.values())
state_trans_dict = {perm:p for perm, p in zip(sorted(all_permutations), pbs)}
return state_trans_dict
state_perms = [''.join(tup) for tup in permutations(list_of_unique_states, 2)]
all_permutations = [state+state for state in list_of_unique_states] + state_perms
state_trans_dict = {perm: (1/N) for perm in all_permutations}
return state_trans_dict
def generate_emission_prob_dist(list_of_unique_states, sequence, **kwargs):
'''
list_of_unique_states: list of states as strings
sequence: array of observations
EXAMPLE1:
corn15_17_seq = generate_cluster_assignments(corn15_17[['price']])
STATE_LIST = ['S1', 'S2']
generate_emission_prob_dist(STATE_LIST, corn15_17_seq, S1=(0.1, 0.3, 0.4, 0.15, 0.05))
OUTPUT1:
{'S1_0': 0.1,
'S1_1': 0.3,
'S1_2': 0.4,
'S1_3': 0.05,
'S1_4': 0.05,
'S2_0': 0.2,
'S2_1': 0.2,
'S2_2': 0.2,
'S2_3': 0.2,
'S2_4': 0.2}
'''
# number of unique obs
B = list(np.unique(sequence).astype(str))
# this runs if specific transitions are provided
if kwargs:
for t in kwargs.values():
assert len(t) == len(B), "Must provide all probabilities for unique emissions in given state."
assert round(np.sum(t)) == 1.0, "Given emission probabilities for a state must add up to 1.0"
for k in kwargs.keys():
assert k in list_of_unique_states, "Keyword arguments must match a value included in `list_of_unique_states`"
diff = list(set(list_of_unique_states).difference(kwargs.keys()))
pbs = chain.from_iterable(kwargs.values())
obs_perms = [state + '_' + str(obs) for state in kwargs.keys() for obs in B]
obs_trans_dict = {perm:p for perm, p in zip(sorted(obs_perms), pbs)}
if diff:
obs_perms_diff = [state + '_' + obs for state in diff for obs in B]
obs_trans_dict.update({perm: (1/len(B)) for perm in obs_perms_diff})
return obs_trans_dict
obs_perms = [state + '_' + obs for state in list_of_unique_states for obs in B]
obs_trans_dict = {perm: (1/len(B)) for perm in obs_perms}
return obs_trans_dict
def generate_init_prob_dist(list_of_unique_states, **kwargs):
"""
Examples:
STATE_LIST = ['S0','S1','S2','S3','S4']
initial_states = {'S1':.2, 'S2':.3, 'S3':.05, 'S4':.25, 'S0':.2}
print(generate_init_prob_dist(STATE_LIST))
# --> {'S0': 0.2, 'S1': 0.2, 'S2': 0.2, 'S3': 0.2, 'S4': 0.2}
print(generate_init_prob_dist(STATE_LIST, **initial_states)) ### NOTE: must unpack dictionary with **
# --> {'S1': 0.2, 'S2': 0.3, 'S3': 0.05, 'S4': 0.25, 'S0': 0.2}
"""
# number of states
N = len(list_of_unique_states)
# this runs if specific transitions are provided
if kwargs:
for t in kwargs.values():
assert isinstance(t, float), "Must provide probabilities as floats."
assert t > 0, "Probabilities must be greater than 0."
assert np.sum(list(kwargs.values())) == 1.0, "Given probabilities must add up to 1.0"
assert len(kwargs) == len(list_of_unique_states), "Please provide initial probabilities for all states, or leave blank"
# build the prob dictionary
init_prob_dict = {item[0]: item[1] for item in kwargs.items()}
return init_prob_dict
init_prob_dist = {state: (1/N) for state in list_of_unique_states}
return init_prob_dist
```
### Create a Priori State Transition
We will use the functions defined above to permutate state transitions and to create a matrix with such entries.
```
# Make permutations of state transition (this len should match len(STATE_TRANS_PROBS))
state_transitions_list = make_state_permutations(STATE_LIST)
# Create the transition matrix in form of dictionary
state_transition_probs = {
trans: prob for trans, prob in zip(state_transitions_list, STATE_TRANS_PROBS)
}
state_transition_probs
```
Above, we formatted the state transition matrix as a dictionary.
We can also represent this dictionary in another convenient format using **'kwargs' argument**. This transformation of the dictionary can be done using the function 'dict_to_tuples' as displayed in the cell below.
```
# Transform dictionary to be in tuple format
#### - this format is required in the `generate_*` functions used later
A_prior = dict_to_tuples(STATE_LIST, state_transition_probs)
A_prior
```
#### NOTE ON FORMATTING
Some functions in this assignment require the transition probabilities to be in a specific format. We will switch between the two formats that hold the same information.
**Format 1**: {'S1S1': 0.4, 'S1S2': 0.6, 'S2S1': 0.35, 'S2S2': 0.55}
The above dictionary contains the state transition matrix. Every key defines the likelihood of transitioning from one state to another. For example, the key-value pair, ''S1S2': 0.6', says the probability of the state moving from 'S1' to 'S2', from observation 'i' to observation 'j', is '0.6'.
**Format 2**: {'S1': (0.4, 0.6), 'S2': (0.35, 0.55)}
The second format contains the same information as the first but encodes the probabilities in tuples. With only two states and assuming 'S1' is the first state, ''S1': (0.4, 0.6)' is interpreted as:
- The probability of staying in 'S1' is 0.4
- The probability of moving from 'S1' to 'S2' is 0.6
### Create Emission Probability Priors
In the above **Format 1**, we will initialize some emission probabilities manually.
```
# In Format 1, manually initialize the emission probabilities
B_format1 = {
'S1_0': 0.1,
'S1_1': 0.3,
'S1_2': 0.4,
'S1_3': 0.15,
'S1_4': 0.05,
'S2_0': 0.15,
'S2_1': 0.2,
'S2_2': 0.3,
'S2_3': 0.05,
'S2_4': 0.3
}
```
The emission probabilities can be stored in dictionary format as well. Using the function `obs_to_tuples()` function in the cell below, we convert the emission probabilities to a dictionary format that is well suited to be provided as an argument in `**kwargs`.
```
# Convert emission matrix to format 2
B_format2 = obs_to_tuples(STATE_LIST, B_format1, corn13_17_seq)
B_format2
```
The emission probabilities can be converted back to the original format by using the previously defined `generate_emission_prob_dist` function. This is demonstrated in the following cell:
```
# Use `generate_emission_prob_dist` to convert B back to format 1
generate_emission_prob_dist(STATE_LIST, corn13_17_seq, **B_format2)
```
We will keep the emission probabilities in the '"key": tuple' format so that it can be used easily as a '**kwargs' argument** later.
```
B_prior = obs_to_tuples(STATE_LIST, B_format1, corn13_17_seq)
```
### Let's recap.
A fair amount of setup has already occurred and we have not yet started the HMM Learning procedure. Let's take a moment to recap the important elements we have established so far.
**We have a state transition matrix, $A$ (prior for $A$)**
```
A_prior
```
**We have the emissions probability matrix, $B$ (prior for $B$)**
```
B_prior
```
**We need the initial state probability matrix, $\pi$. We will use 'generate_init_prob_dist' to do this.**
We can also use the 'generate_init_prob_dist' function without specified parameters to make the uniform initial state probabilities.
_Note: If `pi__init` is not provided, a uniform distribution is produced based on the number of states._
```
# User specified the initial probabilities
pi__init = {'S1': 0.4 , 'S2': 0.6}
# generate the dictionary holding the initial state probabilities
pi = generate_init_prob_dist(STATE_LIST, **pi__init)
pi
# Using default initial parameters - demonstration only, we won't save this dictionary
generate_init_prob_dist(STATE_LIST)
```
**And we have defined a number of functions that will be involved in the EM algorithm in some way**
A summary of the constants and functions defined above is given below:
_Constants_:
STATE_LIST
STATE_TRANS_PROBS
_Functions_:
'generate_cluster_assignments'
'make_state_permutations'
'_grab_highest_prob_and_state'
'dict_to_tuples'
'obs_to_tuples'
'generate_state_trans_dict'
'generate_emission_prob_dist'
'generate_init_prob_dist'
**Finally, we need to create a data structure that will hold all of our probability calculations until we are finished computing the E Step of the EM algorithm**
For this task, we will take the advantage of a powerful data structure from the 'collections' module: 'namedtuple'.
#### NAMED TUPLES
Take a few minutes to review the [documentation](https://docs.python.org/3.6/library/collections.html#collections.namedtuple) on 'namedtuples'. Then answer the following question.
Alternatively, for a short and helpful introduction, review [this tutorial](https://dbader.org/blog/writing-clean-python-with-namedtuples).
<a id="q07"></a>
[Return to top](#questions)
### Question 07
*10 points*
Consider the following array of probabilities:
```
probs = np.array([0.3, 0.7])
```
Create a *namedtuple* factory called 'state' that has two field names: 'prob1' and 'prob2'. After defining the factory, instantiate an instance of the 'state' factory called 'my_state' and store the two probabilities contained in the array 'probs' defined above in the 'prob1' and 'prob2' field names, respectively. Assign the value of the 'prob1' field name to 'ans7' below.
```
### GRADED
### YOUR SOLUTION HERE
my_state = None
ans7 = None
###
### YOUR CODE HERE
###
###
### AUTOGRADER TEST - DO NOT REMOVE
###
```
### Putting it all together
We now have all the pieces to use the EM algorithm. To do this, we require the calculation of forward backward algorithm. So, we will use what we have recently learned about 'namedtuple' from the 'collections' module to do this.
The function below will generate this data structure for us.
```
def generate_obs_data_structure(sequence):
# sequence: 1D numpy array of observations
ObservationData = namedtuple(
'ObservationData',
['prob_lst', 'highest_prob', 'highest_state']
)
return {index+1: ObservationData for index in np.arange(len(sequence)-1)}
```
### STEP 1: ESTIMATE PROBABILITIES:
This step involves using the Forward-Backward algorithm to calculate the probability of observing a sequence, given a set of HMM parameters. We have all the tools to do this now.
```
# Enforce an Argument Signature on the following function to prevent errors with **kwargs
params = [Parameter('list_of_unique_states', Parameter.POSITIONAL_OR_KEYWORD),
Parameter('sequence', Parameter.POSITIONAL_OR_KEYWORD),
Parameter('A', Parameter.KEYWORD_ONLY, default=generate_state_trans_dict),
Parameter('B', Parameter.KEYWORD_ONLY, default=generate_emission_prob_dist),
Parameter('pi', Parameter.KEYWORD_ONLY, default=generate_init_prob_dist)]
sig = Signature(params)
def calculate_probabilities(list_of_unique_states, sequence, **kwargs):
# enforce signature to ensure variable names
bound_values = sig.bind(list_of_unique_states, sequence, **kwargs)
bound_values.apply_defaults()
# grab params that are left to default values
param_defaults = [(name, val) for name, val in bound_values.arguments.items() if callable(val)]
# grab non-default params
set_params = [(name, val) for name, val in bound_values.arguments.items() if isinstance(val, dict)]
# this will run if any default hmm parameters are used
if param_defaults:
for name, val in param_defaults:
if name == 'B':
B = val(list_of_unique_states, sequence)
elif name == 'A':
A = val(list_of_unique_states)
elif name == 'pi':
pi = val(list_of_unique_states)
else:
continue
# this will run if kwargs are provided
if set_params:
for name, val in set_params:
if name == 'B':
B = generate_emission_prob_dist(list_of_unique_states, sequence, **val)
elif name == 'A':
A = generate_state_trans_dict(list_of_unique_states, **val)
elif name == 'pi':
pi = generate_init_prob_dist(list_of_unique_states, **val)
else:
continue
# instantiate the data structure
obs_probs = generate_obs_data_structure(sequence)
# all state transitions
state_perms = make_state_permutations(list_of_unique_states)
# for every transition from one observation to the next, calculate the probability of going from Si to Sj
# loop through observations
for idx, obs in enumerate(sequence):
if idx != 0: # check if this is the first observation
# instantiate the namedtuple for this observation
obs_probs[idx] = obs_probs[idx]([], [], [])
# loop through each possible state transition
for st in state_perms:
# calculate prob of current obs for this state
prev_prob = pi[st[:2]] * B[st[:2]+'_'+str(sequence[idx-1])]
# calculate prob of previous obs for this state
curr_prob = A[st] * B[st[2:]+'_'+str(obs)]
# combine these two probabilities
combined_prob = round(curr_prob * prev_prob, 4)
# append probability to the list in namedtuple
obs_probs[idx].prob_lst.append(combined_prob)
# check for highest prob of observing that sequence
prob_and_state = _grab_highest_prob_and_state(state_perms, obs_probs[idx].prob_lst)
obs_probs[idx].highest_prob.append(prob_and_state[0])
obs_probs[idx].highest_state.append(prob_and_state[1])
else: # this is the first observation, exit loop.
continue
return (obs_probs, A, B, pi)
ob_prob, A, B, pi = calculate_probabilities(STATE_LIST, corn13_17_seq, A=A_prior, B=B_prior, pi=pi)
ob_prob
A
B
pi
```
### STEP 2: UPDATE PARAMETERS
**Update the State Transition Matrix**
```
# This function sums all of the probabilities and
# gives an output of a new (un-normalized) state transition matrix
def new_state_trans(STATE_LIST, probabilities):
state_perms = make_state_permutations(STATE_LIST)
sums_of_st_trans_prob = {p:0 for p in state_perms}
highest_prob_sum = 0
for obs in probabilities:
highest_prob_sum += probabilities[obs].highest_prob[0]
for i, p in enumerate(sums_of_st_trans_prob):
sums_of_st_trans_prob[p] += probabilities[obs].prob_lst[i]
for key in sums_of_st_trans_prob:
sums_of_st_trans_prob[key] = sums_of_st_trans_prob[key] / highest_prob_sum
# finally, normalize so the rows add up to 1
for s in STATE_LIST:
l = []
for k in sums_of_st_trans_prob:
if s == k[:2]:
l.append(sums_of_st_trans_prob[k])
for k in sums_of_st_trans_prob:
if s == k[:2]:
sums_of_st_trans_prob[k] = sums_of_st_trans_prob[k] / sum(l)
return sums_of_st_trans_prob
# Update and normalize the posterior state transition
A_posterior = new_state_trans(STATE_LIST, ob_prob)
A_posterior
# Convert the transition state to "format 2" so it can be
# used as input in the next iteration of "E" step
A_posterior = dict_to_tuples(STATE_LIST, A_posterior)
```
**Update the Emission Probabilities**
Here, we define some functions designed to do specific tasks.
```
##### tally up all observed sequences
def observed_pairs(sequence):
observed_pairs = []
for idx in range(len(sequence)-1):
observed_pairs.append((sequence[idx], sequence[idx+1]))
return observed_pairs
def make_emission_permutations(sequence):
unique_e = np.unique(sequence)
return list(product(unique_e, repeat = 2))
make_emission_permutations([1,1,0, 2])
make_emission_permutations([0,1,0,3,0])
def find_highest_with_state_obs(prob_pairs, state, obs):
for pp in prob_pairs:
if pp[0].count((state,obs))>0:
return pp[1]
def normalize_emissions(b_tuple_format):
new_b_dict = {}
for key, val in b_tuple_format.items():
denominator = sum(val)
new_lst = [v/denominator for v in val]
new_b_dict[key] = tuple(new_lst)
return new_b_dict
```
**Finally, we are ready to update the emission probabilities with the function below.**
```
def emission_matrix_update(sequence, state_list, A, B, pi):
state_pairs = list(product(state_list, repeat = 2))
obs_pairs = observed_pairs(sequence)
new_B = {}
for obs in np.unique(sequence): # For every unique emission
# Find all the sequence-pairs that include that emission
inc_seq = [seq for seq in obs_pairs if seq.count(obs)>0]
# Collector for highest-probabilities
highest_pairs = []
# For each sequence-pair that include that emission
for seq in inc_seq:
prob_pairs = []
# Go through each potential pair of states
for state_pair in state_pairs:
state1, state2 = state_pair
obs1, obs2 = seq
# Match each state with its emission
assoc_tuples = [(state1, obs1),
(state2, obs2)]
# Calculate the probability of the sequence from state
prob = pi[state1] * B[state1+"_"+str(obs1)]
prob *= A[state1+state2]*B[state2+"_"+str(obs2)]
prob = round(prob,5)
# Append the state emission tuples and probability
prob_pairs.append([assoc_tuples, prob])
# Sort probabilities by maximum probability
prob_pairs = sorted(prob_pairs, key = lambda x: x[1], reverse = True)
# Save the highest probability
to_add = {'highest':prob_pairs[0][1]}
# Find the highest probability where each state is associated
# With the current emission
for state in STATE_LIST:
highest_of_state = 0
# Go through sorted list, find first (state,observation) tuple
# save associated probability
for pp in prob_pairs:
if pp[0].count((state,obs))>0:
highest_of_state = pp[1]
break
to_add[state] = highest_of_state
# Save completed dictionary
highest_pairs.append(to_add)
# Total highest_probability
highest_probability =sum([d['highest'] for d in highest_pairs])
# Total highest probabilities for each state; divide by highest prob
# Add to new emission matrix
for state in STATE_LIST:
new_B[state+"_"+str(obs)]= sum([d[state] for d in highest_pairs])/highest_probability
return new_B
```
Run the function:
```
nb = emission_matrix_update(corn13_17_seq,STATE_LIST, A,B,pi)
nb
```
The emission probabilities are updated, but they need to be normalized. To do this, we will convert to dictionary to the 'key: tuple' format and normalize so that the probabilities add up to 1.
```
B_ = obs_to_tuples(STATE_LIST, nb, corn13_17_seq)
B_posterior = normalize_emissions(B_)
# normalized state transition posterior:
A_posterior
# normalized emission posterior probabilities
B_posterior
```
### STEP 3: REPEAT UNTIL PARAMETERS CONVERGE
```
ob_prob2, A2, B2, pi2 = calculate_probabilities(STATE_LIST, corn13_17_seq, A=A_posterior, B=B_posterior, pi=pi)
ob_prob2
A_post2 = new_state_trans(STATE_LIST, ob_prob2) # update and normalize state transition matrix again
A_post2 = dict_to_tuples(STATE_LIST, A2) # convert to `key: tuple` format
A_post2
# update the emissions matrix again
nb2 = emission_matrix_update(corn13_17_seq, STATE_LIST, A2, B2, pi) # update emissions matrix again
B_post2 = obs_to_tuples(STATE_LIST, nb2, corn13_17_seq) # convert emission posterior to `key:tuples` format
B_post2 = normalize_emissions(B_post2) # normalize emissions probabilities
B_post2
```
| github_jupyter |
```
#import libaries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
#save data to dataframe
path = 'https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/DA0101EN/automobileEDA.csv'
df = pd.read_csv(path)
df.head()
#Simple Linear Regression
'''Metode untuk mengetahui hubungan antara dua variabel,
dependen (response) dengan independen (predictor)'''
from sklearn.linear_model import LinearRegressiongression
#objek Linear Regression
lm = LinearRegression()
'''mengetahui hubungan antara variabel highway-mpg dengan price'''
X = df[['highway-mpg']]
y = df [['price']]
'''Fit X dan y dengan model'''
lm.fit(X,y)
'''Prediksi dari model terhadap data baru'''
Yhat = lm.predict(X)
Yhat[0:5]
#nilai dari intercept (a)
lm.intercept_
#nilai dari slope (b)
lm.coef_
#Object LR baru
lm1= LinearRegression()
'''Train model dengan engine-size sebagai predictor'''
X1 = df[['engine-size']]
lm1.fit(X1,y)
print (lm.intercept_)
print("\n", lm.coef_)
#Multi Linear Regression
#Menggunakan lebih dari satu variabel sebagai predictor
#Kasus ini menggunakan horsepower, curb-weight,engine-size
#highway-mpg
Z = df[['horsepower', 'curb-weight', 'engine-size','highway-mpg']]
lm.fit(Z,df['price'])
lm.intercept_
lm.coef_
'''Model Baru Multi Linear Regression'''
lm2 = LinearRegression()
Z2 = df[['normalized-losses', 'highway-mpg']]
lm2.fit(Z2, df['price'])
lm.coef_
#Visualisasi
import seaborn as sns
%matplotlib inline
'''Regression Plot'''
plt.figure (figsize=(12,10))
sns.regplot(x='highway-mpg',y='price', data=df)
plt.ylim(0,)
plt.figure (figsize=(12,10))
sns.regplot(x='peak-rpm',y='price', data=df)
plt.ylim(0,)
df[['peak-rpm','highway-mpg','price']].corr()
'''highway-mpg memiliki nilai korelasi yang paling tinggi dengan
price'''
plt.figure (figsize=(12,10))
sns.residplot(df['highway-mpg'], df['price'])
plt.show()
'''Visualisasi Multiple Linear Regression
menggunakan distribution plot'''
Yhat = lm.predict(Z)
plt.figure(figsize=(12, 10))
ax1 = sns.distplot(df['price'], hist=False, color="r", label="Actual Value")
sns.distplot(Yhat, hist=False, color="b", label="Fitted Values" , ax=ax1)
plt.title('Actual vs Fitted Values for Price')
plt.xlabel('Price (in dollars)')
plt.ylabel('Proportion of Cars')
plt.show()
plt.close()
'''Polynomial Regression'''
#fungsi untuk visualisasi
def PlotPolly(model, independent_variable, dependent_variabble, Name):
x_new = np.linspace(15, 55, 100)
y_new = model(x_new)
plt.plot(independent_variable, dependent_variabble, '.', x_new, y_new, '-')
plt.title('Polynomial Fit with Matplotlib for Price ~ Length')
ax = plt.gca()
ax.set_facecolor((0.898, 0.898, 0.898))
fig = plt.gcf()
plt.xlabel(Name)
plt.ylabel('Price of Cars')
plt.show()
plt.close()
x = df['highway-mpg']
y = df['price']
#menggunakan polynomial orde 3
f = np.polyfit(x,y,3)
p = np.poly1d(f)
p
PlotPolly(p,x,y,'highway-mpg')
'''Membuat dengan orde 11'''
f1 = np.polyfit(x,y,11)
p1 = np.poly1d(f1)
p1
PlotPolly(p1,x,y,'highway-mpg')
from sklearn.preprocessing import PolynomialFeatures
pr = PolynomialFeatures(degree=2)
Z_pr = pr.fit_transform(Z)
Z.shape
Z_pr.shape
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
Input= [('scale', StandardScaler()), ('polynomial', PolynomialFeatures(include_bias=False)), ('model', LinearRegression())]
pipe = Pipeline(Input)
pipe
pipe.fit(Z,y)
ypipe = pipe.predict(Z)
ypipe[0:4]
'''Measures forIn-Sample Evaluation'''
'''R Squared
untuk Linear Regression'''
lm.fit(X, y)
print ('Nilai R-Square : ', lm.score(X,y))
Yhat = lm.predict(X)
print ('Prediksi: ', Yhat[0:4])
from sklearn.metrics import mean_squared_error
mse = mean_squared_error(df['price'], Yhat)
print(mse)
'''Untuk Multi Linear Regression'''
lm.fit (Z, df['price'])
print('R^2 : ', lm.score(Z, df['price']))
y_predict_multi = lm.predict(Z)
print ('MSE: ', mean_squared_error(df['price'], y_predict_multi))
from sklearn.metrics import r2_score
r_squared = r2_score(y,p(x))
print ('Nilai R^2: ', r_squared)
mean_squared_error(df['price'], p(x))
'''New Input'''
new_input=np.arange(1,100,1).reshape(-1,1)
lm.fit(X,y)
lm
yhat = lm.predict(new_input)
yhat[0:5]
plt.plot(new_input, yhat)
plt.show()
'''Model yang bagus adalah yang memiliki nilai R-Square terbesar
dan MSE terkecil'''
```
| github_jupyter |
# Representing Qubit States
You now know something about bits, and about how our familiar digital computers work. All the complex variables, objects and data structures used in modern software are basically all just big piles of bits. Those of us who work on quantum computing call these *classical variables.* The computers that use them, like the one you are using to read this article, we call *classical computers*.
In quantum computers, our basic variable is the _qubit:_ a quantum variant of the bit. These have exactly the same restrictions as normal bits do: they can store only a single binary piece of information, and can only ever give us an output of `0` or `1`. However, they can also be manipulated in ways that can only be described by quantum mechanics. This gives us new gates to play with, allowing us to find new ways to design algorithms.
To fully understand these new gates, we first need to understand how to write down qubit states. For this we will use the mathematics of vectors, matrices and complex numbers. Though we will introduce these concepts as we go, it would be best if you are comfortable with them already. If you need a more in-depth explanation or refresher, you can find a guide [here](../ch-prerequisites/linear_algebra.html).
## Contents
1. [Classical vs Quantum Bits](#cvsq)
1.1 [Statevectors](#statevectors)
1.2 [Qubit Notation](#notation)
1.3 [Exploring Qubits with Qiskit](#exploring-qubits)
2. [The Rules of Measurement](#rules-measurement)
2.1 [A Very Important Rule](#important-rule)
2.2 [The Implications of this Rule](#implications)
3. [The Bloch Sphere](#bloch-sphere)
3.1 [Describing the Restricted Qubit State](#bloch-sphere-1)
3.2 [Visually Representing a Qubit State](#bloch-sphere-2)
## 1. Classical vs Quantum Bits <a id="cvsq"></a>
### 1.1 Statevectors<a id="statevectors"></a>
In quantum physics we use _statevectors_ to describe the state of our system. Say we wanted to describe the position of a car along a track, this is a classical system so we could use a number $x$:

$$ x=4 $$
Alternatively, we could instead use a collection of numbers in a vector called a _statevector._ Each element in the statevector contains the probability of finding the car in a certain place:

$$
|x\rangle = \begin{bmatrix} 0\\ \vdots \\ 0 \\ 1 \\ 0 \\ \vdots \\ 0 \end{bmatrix}
\begin{matrix} \\ \\ \\ \leftarrow \\ \\ \\ \\ \end{matrix}
\begin{matrix} \\ \\ \text{Probability of} \\ \text{car being at} \\ \text{position 4} \\ \\ \\ \end{matrix}
$$
This isn’t limited to position, we could also keep a statevector of all the possible speeds the car could have, and all the possible colours the car could be. With classical systems (like the car example above), this is a silly thing to do as it requires keeping huge vectors when we only really need one number. But as we will see in this chapter, statevectors happen to be a very good way of keeping track of quantum systems, including quantum computers.
### 1.2 Qubit Notation <a id="notation"></a>
Classical bits always have a completely well-defined state: they are either `0` or `1` at every point during a computation. There is no more detail we can add to the state of a bit than this. So to write down the state of a classical bit (`c`), we can just use these two binary values. For example:
c = 0
This restriction is lifted for quantum bits. Whether we get a `0` or a `1` from a qubit only needs to be well-defined when a measurement is made to extract an output. At that point, it must commit to one of these two options. At all other times, its state will be something more complex than can be captured by a simple binary value.
To see how to describe these, we can first focus on the two simplest cases. As we saw in the last section, it is possible to prepare a qubit in a state for which it definitely gives the outcome `0` when measured.
We need a name for this state. Let's be unimaginative and call it $0$ . Similarly, there exists a qubit state that is certain to output a `1`. We'll call this $1$. These two states are completely mutually exclusive. Either the qubit definitely outputs a ```0```, or it definitely outputs a ```1```. There is no overlap. One way to represent this with mathematics is to use two orthogonal vectors.
$$
|0\rangle = \begin{bmatrix} 1 \\ 0 \end{bmatrix} \, \, \, \, |1\rangle =\begin{bmatrix} 0 \\ 1 \end{bmatrix}.
$$
This is a lot of notation to take in all at once. First, let's unpack the weird $|$ and $\rangle$. Their job is essentially just to remind us that we are talking about the vectors that represent qubit states labelled $0$ and $1$. This helps us distinguish them from things like the bit values ```0``` and ```1``` or the numbers 0 and 1. It is part of the bra-ket notation, introduced by Dirac.
If you are not familiar with vectors, you can essentially just think of them as lists of numbers which we manipulate using certain rules. If you are familiar with vectors from your high school physics classes, you'll know that these rules make vectors well-suited for describing quantities with a magnitude and a direction. For example, the velocity of an object is described perfectly with a vector. However, the way we use vectors for quantum states is slightly different from this, so don't hold on too hard to your previous intuition. It's time to do something new!
With vectors we can describe more complex states than just $|0\rangle$ and $|1\rangle$. For example, consider the vector
$$
|q_0\rangle = \begin{bmatrix} \tfrac{1}{\sqrt{2}} \\ \tfrac{i}{\sqrt{2}} \end{bmatrix} .
$$
To understand what this state means, we'll need to use the mathematical rules for manipulating vectors. Specifically, we'll need to understand how to add vectors together and how to multiply them by scalars.
<p>
<details>
<summary>Reminder: Matrix Addition and Multiplication by Scalars (Click here to expand)</summary>
<p>To add two vectors, we add their elements together:
$$|a\rangle = \begin{bmatrix}a_0 \\ a_1 \\ \vdots \\ a_n \end{bmatrix}, \quad
|b\rangle = \begin{bmatrix}b_0 \\ b_1 \\ \vdots \\ b_n \end{bmatrix}$$
$$|a\rangle + |b\rangle = \begin{bmatrix}a_0 + b_0 \\ a_1 + b_1 \\ \vdots \\ a_n + b_n \end{bmatrix} $$
</p>
<p>And to multiply a vector by a scalar, we multiply each element by the scalar:
$$x|a\rangle = \begin{bmatrix}x \times a_0 \\ x \times a_1 \\ \vdots \\ x \times a_n \end{bmatrix}$$
</p>
<p>These two rules are used to rewrite the vector $|q_0\rangle$ (as shown above):
$$
\begin{aligned}
|q_0\rangle & = \tfrac{1}{\sqrt{2}}|0\rangle + \tfrac{i}{\sqrt{2}}|1\rangle \\
& = \tfrac{1}{\sqrt{2}}\begin{bmatrix}1\\0\end{bmatrix} + \tfrac{i}{\sqrt{2}}\begin{bmatrix}0\\1\end{bmatrix}\\
& = \begin{bmatrix}\tfrac{1}{\sqrt{2}}\\0\end{bmatrix} + \begin{bmatrix}0\\\tfrac{i}{\sqrt{2}}\end{bmatrix}\\
& = \begin{bmatrix}\tfrac{1}{\sqrt{2}} \\ \tfrac{i}{\sqrt{2}} \end{bmatrix}\\
\end{aligned}
$$
</details>
</p>
<p>
<details>
<summary>Reminder: Orthonormal Bases (Click here to expand)</summary>
<p>
It was stated before that the two vectors $|0\rangle$ and $|1\rangle$ are orthonormal, this means they are both <i>orthogonal</i> and <i>normalised</i>. Orthogonal means the vectors are at right angles:
</p><p><img src="images/basis.svg"></p>
<p>And normalised means their magnitudes (length of the arrow) is equal to 1. The two vectors $|0\rangle$ and $|1\rangle$ are <i>linearly independent</i>, which means we cannot describe $|0\rangle$ in terms of $|1\rangle$, and vice versa. However, using both the vectors $|0\rangle$ and $|1\rangle$, and our rules of addition and multiplication by scalars, we can describe all possible vectors in 2D space:
</p><p><img src="images/basis2.svg"></p>
<p>Because the vectors $|0\rangle$ and $|1\rangle$ are linearly independent, and can be used to describe any vector in 2D space using vector addition and scalar multiplication, we say the vectors $|0\rangle$ and $|1\rangle$ form a <i>basis</i>. In this case, since they are both orthogonal and normalised, we call it an <i>orthonormal basis</i>.
</details>
</p>
Since the states $|0\rangle$ and $|1\rangle$ form an orthonormal basis, we can represent any 2D vector with a combination of these two states. This allows us to write the state of our qubit in the alternative form:
$$ |q_0\rangle = \tfrac{1}{\sqrt{2}}|0\rangle + \tfrac{i}{\sqrt{2}}|1\rangle $$
This vector, $|q_0\rangle$ is called the qubit's _statevector,_ it tells us everything we could possibly know about this qubit. For now, we are only able to draw a few simple conclusions about this particular example of a statevector: it is not entirely $|0\rangle$ and not entirely $|1\rangle$. Instead, it is described by a linear combination of the two. In quantum mechanics, we typically describe linear combinations such as this using the word 'superposition'.
Though our example state $|q_0\rangle$ can be expressed as a superposition of $|0\rangle$ and $|1\rangle$, it is no less a definite and well-defined qubit state than they are. To see this, we can begin to explore how a qubit can be manipulated.
### 1.3 Exploring Qubits with Qiskit <a id="exploring-qubits"></a>
First, we need to import all the tools we will need:
```
from qiskit import QuantumCircuit, execute, Aer
from qiskit.visualization import plot_histogram, plot_bloch_vector
from math import sqrt, pi
```
In Qiskit, we use the `QuantumCircuit` object to store our circuits, this is essentially a list of the quantum gates in our circuit and the qubits they are applied to.
```
qc = QuantumCircuit(1) # Create a quantum circuit with one qubit
```
In our quantum circuits, our qubits always start out in the state $|0\rangle$. We can use the `initialize()` method to transform this into any state. We give `initialize()` the vector we want in the form of a list, and tell it which qubit(s) we want to initialise in this state:
```
qc = QuantumCircuit(1) # Create a quantum circuit with one qubit
initial_state = [0,1] # Define initial_state as |1>
qc.initialize(initial_state, 0) # Apply initialisation operation to the 0th qubit
qc.draw('text') # Let's view our circuit (text drawing is required for the 'Initialize' gate due to a known bug in qiskit)
```
We can then use one of Qiskit’s simulators to view the resulting state of our qubit. To begin with we will use the statevector simulator, but we will explain the different simulators and their uses later.
```
backend = Aer.get_backend('statevector_simulator') # Tell Qiskit how to simulate our circuit
```
To get the results from our circuit, we use `execute` to run our circuit, giving the circuit and the backend as arguments. We then use `.result()` to get the result of this:
```
qc = QuantumCircuit(1) # Create a quantum circuit with one qubit
initial_state = [0,1] # Define initial_state as |1>
qc.initialize(initial_state, 0) # Apply initialisation operation to the 0th qubit
result = execute(qc,backend).result() # Do the simulation, returning the result
```
from `result`, we can then get the final statevector using `.get_statevector()`:
```
qc = QuantumCircuit(1) # Create a quantum circuit with one qubit
initial_state = [0,1] # Define initial_state as |1>
qc.initialize(initial_state, 0) # Apply initialisation operation to the 0th qubit
result = execute(qc,backend).result() # Do the simulation, returning the result
out_state = result.get_statevector()
print(out_state) # Display the output state vector
```
**Note:** Python uses `j` to represent $i$ in complex numbers. We see a vector with two complex elements: `0.+0.j` = 0, and `1.+0.j` = 1.
Let’s now measure our qubit as we would in a real quantum computer and see the result:
```
qc.measure_all()
qc.draw('text')
```
This time, instead of the statevector we will get the counts for the `0` and `1` results using `.get_counts()`:
```
result = execute(qc,backend).result()
counts = result.get_counts()
plot_histogram(counts)
```
We can see that we (unsurprisingly) have a 100% chance of measuring $|1\rangle$. This time, let’s instead put our qubit into a superposition and see what happens. We will use the state $|q_0\rangle$ from earlier in this section:
$$ |q_0\rangle = \tfrac{1}{\sqrt{2}}|0\rangle + \tfrac{i}{\sqrt{2}}|1\rangle $$
We need to add these amplitudes to a python list. To add a complex amplitude we use `complex`, giving the real and imaginary parts as arguments:
```
initial_state = [1/sqrt(2), 1j/sqrt(2)] # Define state |q>
```
And we then repeat the steps for initialising the qubit as before:
```
qc = QuantumCircuit(1) # Must redefine qc
qc.initialize(initial_state, 0) # Initialise the 0th qubit in the state `initial_state`
state = execute(qc,backend).result().get_statevector() # Execute the circuit
print(state) # Print the result
results = execute(qc,backend).result().get_counts()
plot_histogram(results)
```
We can see we have an equal probability of measuring either $|0\rangle$ or $|1\rangle$. To explain this, we need to talk about measurement.
## 2. The Rules of Measurement <a id="rules-measurement"></a>
### 2.1 A Very Important Rule <a id="important-rule"></a>
There is a simple rule for measurement. To find the probability of measuring a state $|\psi \rangle$ in the state $|x\rangle$ we do:
$$p(|x\rangle) = | \langle x| \psi \rangle|^2$$
The symbols $\langle$ and $|$ tell us $\langle x |$ is a row vector. In quantum mechanics we call the column vectors _kets_ and the row vectors _bras._ Together they make up _bra-ket_ notation. Any ket $|a\rangle$ has a corresponding bra $\langle a|$, and we convert between them using the conjugate transpose.
<details>
<summary>Reminder: The Inner Product (Click here to expand)</summary>
<p>There are different ways to multiply vectors, here we use the <i>inner product</i>. The inner product is a generalisation of the <i>dot product</i> which you may already be familiar with. In this guide, we use the inner product between a bra (row vector) and a ket (column vector), and it follows this rule:
$$\langle a| = \begin{bmatrix}a_0^*, & a_1^*, & \dots & a_n^* \end{bmatrix}, \quad
|b\rangle = \begin{bmatrix}b_0 \\ b_1 \\ \vdots \\ b_n \end{bmatrix}$$
$$\langle a|b\rangle = a_0^* b_0 + a_1^* b_1 \dots a_n^* b_n$$
</p>
<p>We can see that the inner product of two vectors always gives us a scalar. A useful thing to remember is that the inner product of two orthogonal vectors is 0, for example if we have the orthogonal vectors $|0\rangle$ and $|1\rangle$:
$$\langle1|0\rangle = \begin{bmatrix} 0 , & 1\end{bmatrix}\begin{bmatrix}1 \\ 0\end{bmatrix} = 0$$
</p>
<p>Additionally, remember that the vectors $|0\rangle$ and $|1\rangle$ are also normalised (magnitudes are equal to 1):
$$
\begin{aligned}
\langle0|0\rangle & = \begin{bmatrix} 1 , & 0\end{bmatrix}\begin{bmatrix}1 \\ 0\end{bmatrix} = 1 \\
\langle1|1\rangle & = \begin{bmatrix} 0 , & 1\end{bmatrix}\begin{bmatrix}0 \\ 1\end{bmatrix} = 1
\end{aligned}
$$
</p>
</details>
In the equation above, $|x\rangle$ can be any qubit state. To find the probability of measuring $|x\rangle$, we take the inner product of $|x\rangle$ and the state we are measuring (in this case $|\psi\rangle$), then square the magnitude. This may seem a little convoluted, but it will soon become second nature.
If we look at the state $|q_0\rangle$ from before, we can see the probability of measuring $|0\rangle$ is indeed $0.5$:
$$
\begin{aligned}
|q_0\rangle & = \tfrac{1}{\sqrt{2}}|0\rangle + \tfrac{i}{\sqrt{2}}|1\rangle \\
\langle 0| q_0 \rangle & = \tfrac{1}{\sqrt{2}}\langle 0|0\rangle - \tfrac{i}{\sqrt{2}}\langle 0|1\rangle \\
& = \tfrac{1}{\sqrt{2}}\cdot 1 - \tfrac{i}{\sqrt{2}} \cdot 0\\
& = \tfrac{1}{\sqrt{2}}\\
|\langle 0| q_0 \rangle|^2 & = \tfrac{1}{2}
\end{aligned}
$$
You should verify the probability of measuring $|1\rangle$ as an exercise.
This rule governs how we get information out of quantum states. It is therefore very important for everything we do in quantum computation. It also immediately implies several important facts.
### 2.2 The Implications of this Rule <a id="implications"></a>
### #1 Normalisation
The rule shows us that amplitudes are related to probabilities. If we want the probabilities to add up to 1 (which they should!), we need to ensure that the statevector is properly normalized. Specifically, we need the magnitude of the state vector to be 1.
$$ \langle\psi|\psi\rangle = 1 \\ $$
Thus if:
$$ |\psi\rangle = \alpha|0\rangle + \beta|1\rangle $$
Then:
$$ \sqrt{|\alpha|^2 + |\beta|^2} = 1 $$
This explains the factors of $\sqrt{2}$ you have seen throughout this chapter. In fact, if we try to give `initialize()` a vector that isn’t normalised, it will give us an error:
```
vector = [1,1]
qc.initialize(vector, 0)
```
#### Quick Exercise
1. Create a state vector that will give a $1/3$ probability of measuring $|0\rangle$.
2. Create a different state vector that will give the same measurement probabilities.
3. Verify that the probability of measuring $|1\rangle$ for these two states is $2/3$.
You can check your answer in the widget below (you can use 'pi' and 'sqrt' in the vector):
```
# Run the code in this cell to interact with the widget
from qiskit_textbook.widgets import state_vector_exercise
state_vector_exercise(target=1/3)
```
### #2 Alternative measurement
The measurement rule gives us the probability $p(|x\rangle)$ that a state $|\psi\rangle$ is measured as $|x\rangle$. Nowhere does it tell us that $|x\rangle$ can only be either $|0\rangle$ or $|1\rangle$.
The measurements we have considered so far are in fact only one of an infinite number of possible ways to measure a qubit. For any orthogonal pair of states, we can define a measurement that would cause a qubit to choose between the two.
This possibility will be explored more in the next section. For now, just bear in mind that $|x\rangle$ is not limited to being simply $|0\rangle$ or $|1\rangle$.
### #3 Global Phase
We know that measuring the state $|1\rangle$ will give us the output `1` with certainty. But we are also able to write down states such as
$$\begin{bmatrix}0 \\ i\end{bmatrix} = i|1\rangle.$$
To see how this behaves, we apply the measurement rule.
$$ |\langle x| (i|1\rangle) |^2 = | i \langle x|1\rangle|^2 = |\langle x|1\rangle|^2 $$
Here we find that the factor of $i$ disappears once we take the magnitude of the complex number. This effect is completely independent of the measured state $|x\rangle$. It does not matter what measurement we are considering, the probabilities for the state $i|1\rangle$ are identical to those for $|1\rangle$. Since measurements are the only way we can extract any information from a qubit, this implies that these two states are equivalent in all ways that are physically relevant.
More generally, we refer to any overall factor $\gamma$ on a state for which $|\gamma|=1$ as a 'global phase'. States that differ only by a global phase are physically indistinguishable.
$$ |\langle x| ( \gamma |a\rangle) |^2 = | \gamma \langle x|a\rangle|^2 = |\langle x|a\rangle|^2 $$
Note that this is distinct from the phase difference _between_ terms in a superposition, which is known as the 'relative phase'. This becomes relevant once we consider different types of measurements and multiple qubits.
### #4 The Observer Effect
We know that the amplitudes contain information about the probability of us finding the qubit in a specific state, but once we have measured the qubit, we know with certainty what the state of the qubit is. For example, if we measure a qubit in the state:
$$ |q\rangle = \alpha|0\rangle + \beta|1\rangle$$
And find it in the state $|0\rangle$, if we measure again, there is a 100% chance of finding the qubit in the state $|0\rangle$. This means the act of measuring _changes_ the state of our qubits.
$$ |q\rangle = \begin{bmatrix} \alpha \\ \beta \end{bmatrix} \xrightarrow{\text{Measure }|0\rangle} |q\rangle = |0\rangle = \begin{bmatrix} 1 \\ 0 \end{bmatrix}$$
We sometimes refer to this as _collapsing_ the state of the qubit. It is a potent effect, and so one that must be used wisely. For example, were we to constantly measure each of our qubits to keep track of their value at each point in a computation, they would always simply be in a well-defined state of either $|0\rangle$ or $|1\rangle$. As such, they would be no different from classical bits and our computation could be easily replaced by a classical computation. To achieve truly quantum computation we must allow the qubits to explore more complex states. Measurements are therefore only used when we need to extract an output. This means that we often place the all measurements at the end of our quantum circuit.
We can demonstrate this using Qiskit’s statevector simulator. Let's initialise a qubit in superposition:
```
qc = QuantumCircuit(1) # Redefine qc
initial_state = [0.+1.j/sqrt(2),1/sqrt(2)+0.j]
qc.initialize(initial_state, 0)
qc.draw('text')
```
This should initialise our qubit in the state:
$$ |q\rangle = \tfrac{i}{\sqrt{2}}|0\rangle + \tfrac{1}{\sqrt{2}}|1\rangle $$
We can verify this using the simulator:
```
state = execute(qc, backend).result().get_statevector()
print("Qubit State = " + str(state))
```
We can see here the qubit is initialised in the state `[0.+0.70710678j 0.70710678+0.j]`, which is the state we expected.
Let’s now measure this qubit:
```
qc.measure_all()
qc.draw('text')
```
When we simulate this entire circuit, we can see that one of the amplitudes is _always_ 0:
```
state = execute(qc, backend).result().get_statevector()
print("State of Measured Qubit = " + str(state))
```
You can re-run this cell a few times to reinitialise the qubit and measure it again. You will notice that either outcome is equally probable, but that the state of the qubit is never a superposition of $|0\rangle$ and $|1\rangle$. Somewhat interestingly, the global phase on the state $|0\rangle$ survives, but since this is global phase, we can never measure it on a real quantum computer.
### A Note about Quantum Simulators
We can see that writing down a qubit’s state requires keeping track of two complex numbers, but when using a real quantum computer we will only ever receive a yes-or-no (`0` or `1`) answer for each qubit. The output of a 10-qubit quantum computer will look like this:
`0110111110`
Just 10 bits, no superposition or complex amplitudes. When using a real quantum computer, we cannot see the states of our qubits mid-computation, as this would destroy them! This behaviour is not ideal for learning, so Qiskit provides different quantum simulators: The `qasm_simulator` behaves as if you are interacting with a real quantum computer, and will not allow you to use `.get_statevector()`. Alternatively, `statevector_simulator`, (which we have been using in this chapter) does allow peeking at the quantum states before measurement, as we have seen.
## 3. The Bloch Sphere <a id="bloch-sphere"></a>
### 3.1 Describing the Restricted Qubit State <a id="bloch-sphere-1"></a>
We saw earlier in this chapter that the general state of a qubit ($|q\rangle$) is:
$$
|q\rangle = \alpha|0\rangle + \beta|1\rangle
$$
$$
\alpha, \beta \in \mathbb{C}
$$
(The second line tells us $\alpha$ and $\beta$ are complex numbers). The first two implications in section 2 tell us that we cannot differentiate between some of these states. This means we can be more specific in our description of the qubit.
Firstly, since we cannot measure global phase, we can only measure the difference in phase between the states $|0\rangle$ and $|1\rangle$. Instead of having $\alpha$ and $\beta$ be complex, we can confine them to the real numbers and add a term to tell us the relative phase between them:
$$
|q\rangle = \alpha|0\rangle + e^{i\phi}\beta|1\rangle
$$
$$
\alpha, \beta, \phi \in \mathbb{R}
$$
Finally, since the qubit state must be normalised, i.e.
$$
\sqrt{\alpha^2 + \beta^2} = 1
$$
we can use the trigonometric identity:
$$
\sqrt{\sin^2{x} + \cos^2{x}} = 1
$$
to describe the real $\alpha$ and $\beta$ in terms of one variable, $\theta$:
$$
\alpha = \cos{\tfrac{\theta}{2}}, \quad \beta=\sin{\tfrac{\theta}{2}}
$$
From this we can describe the state of any qubit using the two variables $\phi$ and $\theta$:
$$
|q\rangle = \cos{\tfrac{\theta}{2}}|0\rangle + e^{i\phi}\sin{\tfrac{\theta}{2}}|1\rangle
$$
$$
\theta, \phi \in \mathbb{R}
$$
### 3.2 Visually Representing a Qubit State <a id="bloch-sphere-2"></a>
We want to plot our general qubit state:
$$
|q\rangle = \cos{\tfrac{\theta}{2}}|0\rangle + e^{i\phi}\sin{\tfrac{\theta}{2}}|1\rangle
$$
If we interpret $\theta$ and $\phi$ as spherical co-ordinates ($r = 1$, since the magnitude of the qubit state is $1$), we can plot any qubit state on the surface of a sphere, known as the _Bloch sphere._
Below we have plotted a qubit in the state $|{+}\rangle$. In this case, $\theta = \pi/2$ and $\phi = 0$.
(Qiskit has a function to plot a bloch sphere, `plot_bloch_vector()`, but at the time of writing it only takes cartesian coordinates. We have included a function that does the conversion automatically).
```
from qiskit_textbook.widgets import plot_bloch_vector_spherical
coords = [pi/2,0,1] # [Theta, Phi, Radius]
plot_bloch_vector_spherical(coords) # Bloch Vector with spherical coordinates
```
#### Warning!
When first learning about qubit states, it's easy to confuse the qubits _statevector_ with its _Bloch vector_. Remember the statevector is the vector discussed in [1.1](#notation), that holds the amplitudes for the two states our qubit can be in. The Bloch vector is a visualisation tool that maps the 2D, complex statevector onto real, 3D space.
#### Quick Exercise
Use `plot_bloch_vector()` or `plot_bloch_sphere_spherical()` to plot a qubit in the states:
1. $|0\rangle$
2. $|1\rangle$
3. $\tfrac{1}{\sqrt{2}}(|0\rangle + |1\rangle)$
4. $\tfrac{1}{\sqrt{2}}(|0\rangle - i|1\rangle)$
5. $\tfrac{1}{\sqrt{2}}\begin{bmatrix}i\\1\end{bmatrix}$
We have also included below a widget that converts from spherical co-ordinates to cartesian, for use with `plot_bloch_vector()`:
```
from qiskit_textbook.widgets import bloch_calc
bloch_calc()
import qiskit
qiskit.__qiskit_version__
```
| github_jupyter |
### Propósito:
- Realizar uma análise exploratória dos dados, afim de encontrar padrões que nos tragam insights.
### Questões propostas:
● Quais os 3 principais fatores que levam a um diagnóstico positivo de diabetes?
● Qual a chance de uma pessoa do sexo masculino com menos de 40 anos ter o diagnóstico positivo de diabetes?
● Qual a relação entre Polyphagia e o diagnóstico de diabetes?
Autora: Grazielly de Melo Oliveira
## Dependências
```
#Importando bibliotecas
import os
import pandas as pd
import numpy as np
import seaborn as sns
import cloudpickle
import matplotlib.pyplot as plt
from scipy.stats import norm
from pandas.api.types import is_numeric_dtype
from utils.eda import plotar_correlacao
from mlxtend.frequent_patterns import apriori
from mlxtend.frequent_patterns import association_rules
pd.set_option('display.max_rows', 500)
pd.set_option('display.max_columns', 500)
pd.set_option('display.width', 1000)
%matplotlib inline
cmap = sns.diverging_palette(0,100,74,39,19,25, center='light', as_cmap=True) #heatmap
```
# Dados e Constantes
```
# Constantes
DATA_INTER_PATH = '../data/inter'
TRAIN_DATA = 'diabetes_train.parquet'
# Dados
df_train = pd.read_parquet(os.path.join(DATA_INTER_PATH, TRAIN_DATA))
```
# Análise Exploratória
```
# Separando colunas numéricas e categóricas.
todas_as_variaveis = set(df_train.columns.tolist())
variaveis_categoricas = set(df_train.select_dtypes(include=['object']).columns.tolist())
variaveis_numericas = todas_as_variaveis - variaveis_categoricas
# Distribuição do target para os dados de treino
sns.countplot(x=df_train['target'], data=df_train, palette="Set3")
plt.show()
```
## Apriori
O Apriori trabalha com o conceito de padrões frequentes, criando regras de associação entre o que foi pré estabelecido.
Umas das medidas de desempenho consideradas nesta análise é o Lift que indica qual a chance de Y (nosso target) ser dado como positivo, se X (outras features do dataset) forem positivas também, e considerando toda a popularidade de Y. Em outras palavras, ele verifica qual informação nos traz mais conhecimento sobre a possibilidade de esses padrões serem encontrados juntos.
Para realizar a análise, iremos converter nossas classes para inteiros
```
basket_train_features = df_train.drop(['idade'], axis=1)
dict_convert = {'yes':1, 'no':0, 'female':0, 'male':1}
basket_train_features = basket_train_features.replace(dict_convert)
for col in basket_train_features.columns.tolist():
basket_train_features[col] = pd.to_numeric(basket_train_features[col])
# Aplicando a análise de padrões frequentes
# Aqui estamos lidando com padrões que aparecem, no mínimo 10% das vezes
frequent_itemsets = apriori(basket_train_features, min_support=0.1, use_colnames=True)
# Extraindo regras
rules = association_rules(frequent_itemsets, metric="lift", min_threshold=1)
# Extraindo regras, cujo consequente seja diabetes (target = 1)
rules[(rules['consequents']=={('target')}) & (rules['lift']>1)].sort_values(by='lift', ascending=False).set_index('antecedents')
```
Extraímos a 5 regras com maior lift (que mais aumentam a probabilidade a posteriori do target positivo)
Em geral a polyuria é um bom indicador, pois aparece em vários padrões. Para exemplificação, extraímos apenas 5. Uma análise mais profunda é requerida para decidir o conjunto final de regras, para analisar a correlação das regras, por exemplo.
Com base no lift, extraímos:
- Possuir juntos: (polyuria, polydipsia, cicatrizacao_retardada, perda_de_peso_repentina)
- Possuir juntos: (polydipsia, coceira, cicatrizacao_retardada, perda_de_peso_repentina)
- Possuir juntos: (polyphagia, paresia_parcial, polydipsia, perda_de_peso_repentina)
- Possuir Juntos: (polyphagia, polydipsia, cicatrizacao_retardada, perda_de_peso_repentina)
- Possuir juntos: (polyphagia, polydipsia, coceira, perda_de_peso_repentina)
Se o paciente tem polyuria, polydipsia, cicatrização retardada, perda de peso repentina, ele tem 62% a mais de chance de ter diabetes em estágio inicial.
## Questões Propostas
### Observação
* Estou atribuindo que o target POSITIVO representa 1, e NEGATIVO representa 0.
* A partir daqui, estou atribuindo que o sexo feminino será representado por 0, e o sexo masculino por 1.
```
#Plot da distribuição da variável target.
df_train['genero'] = df_train['genero'].map({'female':0, 'male': 1})
df_train['target'].value_counts(normalize=True).plot.bar()
plt.show()
```
Plot do target para verificar se há desbalanceamento, para garantir uma melhor métrica de validação na modelagem
```
df_train['genero'].value_counts(normalize=True).plot.bar()
plt.show()
```
Na base há mais homens (>60%) que mulheres.
```
sns.histplot(df_train["idade"], kde=False, bins=10, palette="RdYlGn_r")
plt.show()
df_train["idade"].mean()
```
A média de idade é de 48 anos. E a grande maioria está acima dos 30.
Vamos ver a taxa de ocorrência de diabetes em obesos.
```
df_train.groupby('obesidade').mean()['target'].plot.bar()
plt.show()
```
Nesta base, cerca de mais de 75% de pessoas que são obesas também estão em estágio inicial de diabetes.
## Qual a relação entre Polyphagia e o diagnóstico de diabetes?
```
sns.countplot(x='polyphagia', hue="target", data=df_train)
plt.show()
df_train.groupby('polyphagia').mean()['target']
df_train.groupby('polyphagia').mean()['target'].plot.bar()
plt.show()
```
Polifagia é um sinal médico que significa fome excessiva e ingestão anormalmente alta de sólidos pela boca, pode-se notar que nesta amostra de dados cerca de 80% de pessoas que possuem`polyphagia` também possuem diabetes. Apenas 40% de quem não apresenta essa condição possui diabetes (o que é menor, inclusive, que a probabilidade apriori)
Dessa forma, isso nos leva a crer que há uma dependência entre esse fator e a ocorrência de diabetes.
## Qual a chance de uma pessoa do sexo masculino com menos de 40 anos ter o diagnóstico positivo de diabetes?
Considerando que a chance é a probabilidade da classe positiva pela probabilidade da classe negativa.
```
genero_masc = df_train[df_train['genero'] == 1]
genero_masc_40 = genero_masc[genero_masc['idade'] < 40]
genero_masc_40.groupby('genero').mean()['target'].plot.bar()
plt.show()
probabilidade_masc_40_positivo = genero_masc_40.groupby('genero').mean()['target'].values[0]
print(f"{probabilidade_masc_40_positivo*100} % dos homens com menos de 40 anos possuem diabetes em estágio inicial")
# A chance será
chance = round((probabilidade_masc_40_positivo/(1-probabilidade_masc_40_positivo)),2)
chance
```
Isto é aproximadamente uma chance de 5 casos de diabetes em estágio inicial para 11 pacientes sem diabetes em estágio inicial, dado que são homens com menos de 40 anos.
# Quais os 3 principais fatores que levam a um diagnóstico positivo de diabetes?
Esta última pergunta pode ser respondida de mais de uma forma. Primeiramente olharemos para medidas de correlação e associação. Posteriormente, tentaremos algo baseado em shapley values.
```
df_train.replace({'yes':1, 'no':0}, inplace=True)
```
Vamos avaliar a correlação point-biserial entre a variável idadee e o target e o cramer phi para as variáveis categóricas e o target. Embora a correlação e o cramer phi não sejam comparáveis diretamente, podemos ter uma noção da força das variáveis e elencar as candidatas as mais fortes.
Em se tratando de variável binária e target binário, o cramer phi é numericamente igual a correlação de pearson, além disso, a correlação point-biserial também é numericamente igual a correlação de pearson, no caso de uma variável numérica e outra binária. Por isso iremos utilizar o método de correlação para calcular ambas
```
plotar_correlacao(todas_as_variaveis,df_train)
```
Logo pode-se notar que as variáveis `polyuria`, `polydipsia`, e `paresia parcial`, possuem uma associação mais alta em relação ao target. Em se tratando de correlação, a idade é correlacionada positivamente, indicando que quanto mais velho o paciente, mais chance dele estar em estágio inicial de diabetes.
Apesar dos número apresentados, iremos submeter cada variável a testes para selecionar as melhroes.Precisamos, por exemplo, checar se a correção de 0.14 da idade tem significancia estatística antes de usá-la no modelo.
Destaco aqui o cramer phi da coceira, que deu 0. Contudo, esta condição compôs vários padrões frequentes. De fato, a análise de cramer e correlação são univariadas e incapazes de capturar interações. Dessa forma, a variável ainda pode ser útil se combinada com outras. A análise de padrões frequentes pode ajudar a criar features combinando coceira a outros fatores e essas novas features serem mais relevantes que as originais.
| github_jupyter |
# Developing an AI application
Going forward, AI algorithms will be incorporated into more and more everyday applications. For example, you might want to include an image classifier in a smart phone app. To do this, you'd use a deep learning model trained on hundreds of thousands of images as part of the overall application architecture. A large part of software development in the future will be using these types of models as common parts of applications.
In this project, you'll train an image classifier to recognize different species of flowers. You can imagine using something like this in a phone app that tells you the name of the flower your camera is looking at. In practice you'd train this classifier, then export it for use in your application. We'll be using [this dataset](http://www.robots.ox.ac.uk/~vgg/data/flowers/102/index.html) of 102 flower categories, you can see a few examples below.
<img src='assets/Flowers.png' width=500px>
The project is broken down into multiple steps:
* Load and preprocess the image dataset
* Train the image classifier on your dataset
* Use the trained classifier to predict image content
We'll lead you through each part which you'll implement in Python.
When you've completed this project, you'll have an application that can be trained on any set of labeled images. Here your network will be learning about flowers and end up as a command line application. But, what you do with your new skills depends on your imagination and effort in building a dataset. For example, imagine an app where you take a picture of a car, it tells you what the make and model is, then looks up information about it. Go build your own dataset and make something new.
First up is importing the packages you'll need. It's good practice to keep all the imports at the beginning of your code. As you work through this notebook and find you need to import a package, make sure to add the import up here.
```
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
import matplotlib.pyplot as plt
import torch
from torch import nn
from torch import optim
import torch.nn.functional as F
from torchvision import datasets, transforms, models
import numpy as np
from PIL import Image
import time
import os
import random
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
```
## Load the data
Here you'll use `torchvision` to load the data ([documentation](http://pytorch.org/docs/0.3.0/torchvision/index.html)). The data should be included alongside this notebook, otherwise you can [download it here](https://s3.amazonaws.com/content.udacity-data.com/nd089/flower_data.tar.gz). The dataset is split into three parts, training, validation, and testing. For the training, you'll want to apply transformations such as random scaling, cropping, and flipping. This will help the network generalize leading to better performance. You'll also need to make sure the input data is resized to 224x224 pixels as required by the pre-trained networks.
The validation and testing sets are used to measure the model's performance on data it hasn't seen yet. For this you don't want any scaling or rotation transformations, but you'll need to resize then crop the images to the appropriate size.
The pre-trained networks you'll use were trained on the ImageNet dataset where each color channel was normalized separately. For all three sets you'll need to normalize the means and standard deviations of the images to what the network expects. For the means, it's `[0.485, 0.456, 0.406]` and for the standard deviations `[0.229, 0.224, 0.225]`, calculated from the ImageNet images. These values will shift each color channel to be centered at 0 and range from -1 to 1.
```
data_dir = 'flowers'
train_dir = data_dir + '/train'
valid_dir = data_dir + '/valid'
test_dir = data_dir + '/test'
#Define transforms for the training, validation, and testing sets
train_transforms = transforms.Compose([transforms.RandomRotation(30),
transforms.RandomResizedCrop(224),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406],
[0.229, 0.224, 0.225])])
test_transforms = transforms.Compose([transforms.Resize(255),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406],
[0.229, 0.224, 0.225])])
# Load the datasets with ImageFolder
train_data = datasets.ImageFolder(train_dir, transform=train_transforms)
validation_data = datasets.ImageFolder(valid_dir, transform=test_transforms)
test_data = datasets.ImageFolder(test_dir, transform=test_transforms)
# Using the image datasets and the trainforms, define the dataloaders
trainloader = torch.utils.data.DataLoader(train_data, batch_size=64, shuffle=True)
validationloader = torch.utils.data.DataLoader(validation_data, batch_size=64)
testloader = torch.utils.data.DataLoader(test_data, batch_size=64)
```
### Label mapping
You'll also need to load in a mapping from category label to category name. You can find this in the file `cat_to_name.json`. It's a JSON object which you can read in with the [`json` module](https://docs.python.org/2/library/json.html). This will give you a dictionary mapping the integer encoded categories to the actual names of the flowers.
```
import json
with open('cat_to_name.json', 'r') as f:
cat_to_name = json.load(f)
```
# Building and training the classifier
Now that the data is ready, it's time to build and train the classifier. As usual, you should use one of the pretrained models from `torchvision.models` to get the image features. Build and train a new feed-forward classifier using those features.
We're going to leave this part up to you. Refer to [the rubric](https://review.udacity.com/#!/rubrics/1663/view) for guidance on successfully completing this section. Things you'll need to do:
* Load a [pre-trained network](http://pytorch.org/docs/master/torchvision/models.html) (If you need a starting point, the VGG networks work great and are straightforward to use)
* Define a new, untrained feed-forward network as a classifier, using ReLU activations and dropout
* Train the classifier layers using backpropagation using the pre-trained network to get the features
* Track the loss and accuracy on the validation set to determine the best hyperparameters
We've left a cell open for you below, but use as many as you need. Our advice is to break the problem up into smaller parts you can run separately. Check that each part is doing what you expect, then move on to the next. You'll likely find that as you work through each part, you'll need to go back and modify your previous code. This is totally normal!
When training make sure you're updating only the weights of the feed-forward network. You should be able to get the validation accuracy above 70% if you build everything right. Make sure to try different hyperparameters (learning rate, units in the classifier, epochs, etc) to find the best model. Save those hyperparameters to use as default values in the next part of the project.
One last important tip if you're using the workspace to run your code: To avoid having your workspace disconnect during the long-running tasks in this notebook, please read in the earlier page in this lesson called Intro to
GPU Workspaces about Keeping Your Session Active. You'll want to include code from the workspace_utils.py module.
**Note for Workspace users:** If your network is over 1 GB when saved as a checkpoint, there might be issues with saving backups in your workspace. Typically this happens with wide dense layers after the convolutional layers. If your saved checkpoint is larger than 1 GB (you can open a terminal and check with `ls -lh`), you should reduce the size of your hidden layers and train again.
```
network = "resnet50"
inputs = 2048
outputs = 102
learning_rate = 0.001
hidden_units = 2
epochs = 15
model = getattr(models, network)(pretrained=True)
for param in model.parameters():
param.requires_grad = False
classifier = nn.Sequential(nn.Linear(inputs, 1024),
nn.ReLU(),
nn.Dropout(0.2),
nn.Linear(1024, 512),
nn.ReLU(),
nn.Dropout(0.2),
nn.Linear(512, outputs),
nn.LogSoftmax(dim=1))
model.fc = classifier
model.to(device)
criterion = nn.NLLLoss()
optimizer = optim.Adam(model.fc.parameters(), lr=learning_rate);
def train_classifier(epochs, trainloader, device, optimizer, model, criterion, validationloader):
running_loss = 0
start = time.time()
print('Training started')
for epoch in range(epochs):
for images, labels in trainloader:
images, labels = images.to(device), labels.to(device)
optimizer.zero_grad()
log_ps = model.forward(images)
loss = criterion(log_ps, labels)
loss.backward()
optimizer.step()
running_loss += loss.item()
else:
validation_loss = 0
accuracy = 0
model.eval()
with torch.no_grad():
for images, labels in validationloader:
images, labels = images.to(device), labels.to(device)
log_ps = model.forward(images)
loss += criterion(log_ps, labels)
validation_loss += loss.item()
ps = torch.exp(log_ps)
top_p, top_class = ps.topk(1, dim=1)
equals = top_class == labels.view(*top_class.shape)
accuracy += torch.mean(equals.type(torch.FloatTensor)).item()
time_elapsed = time.time() - start
print("Epoch: {}/{}.. ".format(epoch+1, epochs),
"Training Loss: {:.3f}.. ".format(running_loss/len(trainloader)),
"Validation Loss: {:.3f}.. ".format(validation_loss/len(validationloader)),
"Validation Accuracy: {:.3f}.. ".format(accuracy/len(validationloader) * 100),
"Elapsed time: {:.0f}m {:.0f}s".format(time_elapsed//60, time_elapsed % 60))
running_loss = 0
model.train()
print("\nTotal time: {:.0f}m {:.0f}s".format(time_elapsed//60, time_elapsed % 60))
train_classifier(epochs, trainloader, device, optimizer, model, criterion, validationloader)
```
## Testing your network
It's good practice to test your trained network on test data, images the network has never seen either in training or validation. This will give you a good estimate for the model's performance on completely new images. Run the test images through the network and measure the accuracy, the same way you did validation. You should be able to reach around 70% accuracy on the test set if the model has been trained well.
```
testing_loss = 0
accuracy = 0
model.eval()
with torch.no_grad():
for images, labels in testloader:
images, labels = images.to(device), labels.to(device)
log_ps = model.forward(images)
ps = torch.exp(log_ps)
top_p, top_class = ps.topk(1, dim=1)
equals = top_class == labels.view(*top_class.shape)
accuracy += torch.mean(equals.type(torch.FloatTensor)).item()
print("Testing Accuracy: {:.3f}".format(accuracy/len(testloader) * 100))
model.train();
```
## Save the checkpoint
Now that your network is trained, save the model so you can load it later for making predictions. You probably want to save other things such as the mapping of classes to indices which you get from one of the image datasets: `image_datasets['train'].class_to_idx`. You can attach this to the model as an attribute which makes inference easier later on.
```model.class_to_idx = image_datasets['train'].class_to_idx```
Remember that you'll want to completely rebuild the model later so you can use it for inference. Make sure to include any information you need in the checkpoint. If you want to load the model and keep training, you'll want to save the number of epochs as well as the optimizer state, `optimizer.state_dict`. You'll likely want to use this trained model in the next part of the project, so best to save it now.
```
model.class_to_idx = train_data.class_to_idx
checkpoint = {'network': network,
'inputs': inputs,
'outputs': outputs,
'learning_rate': learning_rate,
'hidden_units': hidden_units,
'epochs': epochs,
'classifier': model.fc,
'model_state_dict': model.state_dict(),
'class_to_idx': model.class_to_idx,
'optimizer_state_dict': optimizer.state_dict()}
torch.save(checkpoint, 'checkpoint.pth')
```
## Loading the checkpoint
At this point it's good to write a function that can load a checkpoint and rebuild the model. That way you can come back to this project and keep working on it without having to retrain the network.
```
def load_checkpoint(filepath):
checkpoint = torch.load(filepath, map_location=None if torch.cuda.is_available() else "cpu")
network = checkpoint['network']
inputs = checkpoint['inputs']
outputs = checkpoint['outputs']
learning_rate = checkpoint['learning_rate']
hidden_units = checkpoint['hidden_units']
epochs = checkpoint['epochs']
classifier = checkpoint['classifier']
model = getattr(models, network)()
model.fc = classifier
model.load_state_dict(checkpoint['model_state_dict'])
model.class_to_idx = checkpoint['class_to_idx']
model.to(device)
optimizer = optim.Adam(model.fc.parameters(), lr=checkpoint['learning_rate'])
optimizer.load_state_dict(checkpoint['optimizer_state_dict'])
return network, inputs, outputs, learning_rate, hidden_units, epochs, model, optimizer, classifier
network, inputs, outputs, learning_rate, hidden_units, epochs, model, optimizer, classifier = load_checkpoint('checkpoint.pth')
# Invert class index dictionary for easy lookup
idx_to_class = {value: key for key, value in model.class_to_idx.items()}
testing_loss = 0
accuracy = 0
model.eval()
with torch.no_grad():
for images, labels in testloader:
images, labels = images.to(device), labels.to(device)
log_ps = model.forward(images)
ps = torch.exp(log_ps)
top_p, top_class = ps.topk(1, dim=1)
equals = top_class == labels.view(*top_class.shape)
accuracy += torch.mean(equals.type(torch.FloatTensor)).item()
print("Testing Accuracy: {:.3f}".format(accuracy/len(testloader) * 100))
model.train();
```
# Inference for classification
Now you'll write a function to use a trained network for inference. That is, you'll pass an image into the network and predict the class of the flower in the image. Write a function called `predict` that takes an image and a model, then returns the top $K$ most likely classes along with the probabilities. It should look like
```python
probs, classes = predict(image_path, model)
print(probs)
print(classes)
> [ 0.01558163 0.01541934 0.01452626 0.01443549 0.01407339]
> ['70', '3', '45', '62', '55']
```
First you'll need to handle processing the input image such that it can be used in your network.
## Image Preprocessing
You'll want to use `PIL` to load the image ([documentation](https://pillow.readthedocs.io/en/latest/reference/Image.html)). It's best to write a function that preprocesses the image so it can be used as input for the model. This function should process the images in the same manner used for training.
First, resize the images where the shortest side is 256 pixels, keeping the aspect ratio. This can be done with the [`thumbnail`](http://pillow.readthedocs.io/en/3.1.x/reference/Image.html#PIL.Image.Image.thumbnail) or [`resize`](http://pillow.readthedocs.io/en/3.1.x/reference/Image.html#PIL.Image.Image.thumbnail) methods. Then you'll need to crop out the center 224x224 portion of the image.
Color channels of images are typically encoded as integers 0-255, but the model expected floats 0-1. You'll need to convert the values. It's easiest with a Numpy array, which you can get from a PIL image like so `np_image = np.array(pil_image)`.
As before, the network expects the images to be normalized in a specific way. For the means, it's `[0.485, 0.456, 0.406]` and for the standard deviations `[0.229, 0.224, 0.225]`. You'll want to subtract the means from each color channel, then divide by the standard deviation.
And finally, PyTorch expects the color channel to be the first dimension but it's the third dimension in the PIL image and Numpy array. You can reorder dimensions using [`ndarray.transpose`](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.ndarray.transpose.html). The color channel needs to be first and retain the order of the other two dimensions.
```
def process_image(image):
''' Scales, crops, and normalizes a PIL image for a PyTorch model,
returns a Tensor
'''
im = Image.open(image)
return test_transforms(im).to(device)
```
To check your work, the function below converts a PyTorch tensor and displays it in the notebook. If your `process_image` function works, running the output through this function should return the original image (except for the cropped out portions).
```
def imshow(image, ax=None, title=None):
"""Imshow for Tensor."""
if ax is None:
fig, ax = plt.subplots()
# PyTorch tensors assume the color channel is the first dimension
# but matplotlib assumes is the third dimension
image = image.numpy().transpose((1, 2, 0))
# Undo preprocessing
mean = np.array([0.485, 0.456, 0.406])
std = np.array([0.229, 0.224, 0.225])
image = std * image + mean
# Image needs to be clipped between 0 and 1 or it looks like noise when displayed
image = np.clip(image, 0, 1)
ax.imshow(image)
return ax
```
## Class Prediction
Once you can get images in the correct format, it's time to write a function for making predictions with your model. A common practice is to predict the top 5 or so (usually called top-$K$) most probable classes. You'll want to calculate the class probabilities then find the $K$ largest values.
To get the top $K$ largest values in a tensor use [`x.topk(k)`](http://pytorch.org/docs/master/torch.html#torch.topk). This method returns both the highest `k` probabilities and the indices of those probabilities corresponding to the classes. You need to convert from these indices to the actual class labels using `class_to_idx` which hopefully you added to the model or from an `ImageFolder` you used to load the data ([see here](#Save-the-checkpoint)). Make sure to invert the dictionary so you get a mapping from index to class as well.
Again, this method should take a path to an image and a model checkpoint, then return the probabilities and classes.
```python
probs, classes = predict(image_path, model)
print(probs)
print(classes)
> [ 0.01558163 0.01541934 0.01452626 0.01443549 0.01407339]
> ['70', '3', '45', '62', '55']
```
```
def predict(image_path, model, topk=5):
''' Predict the class (or classes) of an image using a trained deep learning model.
'''
image = process_image(image_path)
model.eval()
with torch.no_grad():
image.unsqueeze_(0)
log_ps = model.forward(image)
ps = torch.exp(log_ps)
probs, classes = ps.topk(topk, dim=1)
return probs.cpu().numpy()[0], list(map(lambda c: cat_to_name[idx_to_class.get(c)], classes.cpu().numpy()[0]))
```
## Sanity Checking
Now that you can use a trained model for predictions, check to make sure it makes sense. Even if the testing accuracy is high, it's always good to check that there aren't obvious bugs. Use `matplotlib` to plot the probabilities for the top 5 classes as a bar graph, along with the input image. It should look like this:
<img src='assets/inference_example.png' width=300px>
You can convert from the class integer encoding to actual flower names with the `cat_to_name.json` file (should have been loaded earlier in the notebook). To show a PyTorch tensor as an image, use the `imshow` function defined above.
```
label = random.choice(os.listdir('./flowers/test/'))
image = random.choice(os.listdir('./flowers/test/' + label))
file_path = './flowers/test/'+ label + '/' + image
probs, classes = predict(file_path, model, 5)
fig = plt.figure(figsize=(6,6))
ax1 = plt.subplot2grid((15,9), (0,0), colspan=9, rowspan=9)
ax2 = plt.subplot2grid((15,9), (9,2), colspan=5, rowspan=5)
image = Image.open(file_path)
ax1.axis('off')
ax1.set_title(cat_to_name[label])
ax1.imshow(image)
y_pos = np.arange(len(classes))
ax2.set_yticks(y_pos)
ax2.set_yticklabels(classes)
ax2.set_xlabel('Probability')
ax2.set_ylabel('Classification')
ax2.invert_yaxis()
ax2.barh(y_pos, probs, align='center', color='steelblue')
plt.show()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/baixianger/DLAV-2022/blob/main/homeworks/hw3/CNN_Exercise.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
## Convolutional Networks
We'll check out how to build a **convolutional network** to classify CIFAR10 images. By using weight sharing - multiple units with the same weights - convolutional layers are able to learn repeated patterns in your data. For example, a unit could learn the pattern for an eye, or a face, or lower level features like edges.
```
from google.colab import drive
drive.mount('/content/drive')
import numpy as np
import time
import torch
from torch import nn
from torch import optim
import torch.nn.functional as F
import torch.utils.data as utils
from torchvision import datasets, transforms
from torch.utils.data.sampler import SubsetRandomSampler
import matplotlib.pyplot as plt
%matplotlib inline
label_names = [
'airplane',
'automobile',
'bird',
'cat',
'deer',
'dog',
'frog',
'horse',
'ship',
'truck'
]
def plot_images(images, cls_true, cls_pred=None):
"""
Adapted from https://github.com/Hvass-Labs/TensorFlow-Tutorials/
"""
fig, axes = plt.subplots(3, 3)
for i, ax in enumerate(axes.flat):
# plot img
ax.imshow(images[i, :, :, :], interpolation='spline16')
# show true & predicted classes
cls_true_name = label_names[cls_true[i]]
if cls_pred is None:
xlabel = "{0} ({1})".format(cls_true_name, cls_true[i])
else:
cls_pred_name = label_names[cls_pred[i]]
xlabel = "True: {0}\nPred: {1}".format(
cls_true_name, cls_pred_name
)
ax.set_xlabel(xlabel)
ax.set_xticks([])
ax.set_yticks([])
plt.show()
def get_train_valid_loader(data_dir='data',
batch_size=64,
augment=False,
random_seed = 1,
valid_size=0.02,
shuffle=True,
show_sample=True,
num_workers=4,
pin_memory=False):
"""
Utility function for loading and returning train and valid
multi-process iterators over the CIFAR-10 dataset. A sample
9x9 grid of the images can be optionally displayed.
If using CUDA, num_workers should be set to 1 and pin_memory to True.
Params
------
- data_dir: path directory to the dataset.
- batch_size: how many samples per batch to load.
- augment: whether to apply the data augmentation scheme
mentioned in the paper. Only applied on the train split.
- random_seed: fix seed for reproducibility.
- valid_size: percentage split of the training set used for
the validation set. Should be a float in the range [0, 1].
- shuffle: whether to shuffle the train/validation indices.
- show_sample: plot 9x9 sample grid of the dataset.
- num_workers: number of subprocesses to use when loading the dataset.
- pin_memory: whether to copy tensors into CUDA pinned memory. Set it to
True if using GPU.
Returns
-------
- train_loader: training set iterator.
- valid_loader: validation set iterator.
"""
error_msg = "[!] valid_size should be in the range [0, 1]."
assert ((valid_size >= 0) and (valid_size <= 1)), error_msg
normalize = transforms.Normalize(
mean=[0.4914, 0.4822, 0.4465],
std=[0.2023, 0.1994, 0.2010],
)
# define transforms
valid_transform = transforms.Compose([
transforms.ToTensor(),
normalize,
])
if augment:
train_transform = transforms.Compose([
transforms.RandomCrop(32, padding=4),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
normalize,
])
else:
train_transform = transforms.Compose([
transforms.ToTensor(),
normalize,
])
# load the dataset
train_dataset = datasets.CIFAR10(
root=data_dir, train=True,
download=True, transform=train_transform,
)
valid_dataset = datasets.CIFAR10(
root=data_dir, train=True,
download=True, transform=valid_transform,
)
num_train = len(train_dataset)
indices = list(range(num_train))
split = int(np.floor(valid_size * num_train))
if shuffle:
np.random.seed(random_seed)
np.random.shuffle(indices)
train_idx, valid_idx = indices[split:], indices[:split]
train_sampler = SubsetRandomSampler(train_idx)
valid_sampler = SubsetRandomSampler(valid_idx)
train_loader = torch.utils.data.DataLoader(
train_dataset, batch_size=batch_size, sampler=train_sampler,
num_workers=num_workers, pin_memory=pin_memory,
)
valid_loader = torch.utils.data.DataLoader(
valid_dataset, batch_size=batch_size, sampler=valid_sampler,
num_workers=num_workers, pin_memory=pin_memory,
)
# visualize some images
if show_sample:
sample_loader = torch.utils.data.DataLoader(
train_dataset, batch_size=9, shuffle=shuffle,
num_workers=num_workers, pin_memory=pin_memory,
)
data_iter = iter(sample_loader)
images, labels = data_iter.next()
X = images.numpy().transpose([0, 2, 3, 1])
plot_images(X, labels)
return (train_loader, valid_loader)
# check cuda
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
if torch.cuda.is_available():
print("Found CUDA device, setting copy tensors into CUDA pinned memory")
trainloader, valloader = get_train_valid_loader(num_workers=1, pin_memory=True)
else:
from multiprocessing import cpu_count
print(f"Found {cpu_count()} CPU kernals, load data.....")
trainloader, valloader = get_train_valid_loader(num_workers=cpu_count(), pin_memory=False)
```
Fomular
$$
\text{out}(N_i, C_{\text{out}_j}) = \text{bias}(C_{\text{out}_j}) +
\sum_{k = 0}^{C_{\text{in}} - 1} \text{weight}(C_{\text{out}_j}, k) \star \text{input}(N_i, k)
$$
where :
- $\star$ is the valid 2D `cross-correlation`_ operator,
- `N` is a batch size,
- `C` denotes a number of channels,
- `H` is a height of input planes in pixels,
- `W` iswidth in pixels.
Shape
- Input: $(N, C_{in}, H_{in}, W_{in})$
- Output:$(N, C_{out}, H_{out}, W_{out})$
where:
- $ H_{out} = \left\lfloor\frac{H_{in} + 2 \times \text{padding}[0] - \text{dilation}[0]
\times (\text{kernel\_size}[0] - 1) - 1}{\text{stride}[0]} + 1\right\rfloor
$
- $ W_{out} = \left\lfloor\frac{W_{in} + 2 \times \text{padding}[1] - \text{dilation}[1]
\times (\text{kernel\_size}[1] - 1) - 1}{\text{stride}[1]} + 1\right\rfloor
$
```
class ConvNet(nn.Module):
def __init__(self, n_input_channels=3, n_output=10):
super().__init__()
################################################################################
# TODO: #
# Define 2 or more different layers of the neural network #
################################################################################
# use the LeNet model
# '?' denotes the batch size, which is 64 in our default setting from func above
self.conv1 = nn.Conv2d(3, 6, 5) # (?, 3,32,32) => (?, 6,28,28)
self.pool = nn.MaxPool2d(2, 2) # we use this pooling twice
# 1st time :(?, 6,28,28) => (?, 6,14,14)
# 2nd time :(?,16,10,10) => (?,16, 5, 5)
self.conv2 = nn.Conv2d(6, 16, 5) # (?, 6,14,14) => (?,16,10,10)
self.fc1 = nn.Linear(16 * 5 * 5, 120) # (?,16, 5, 5) ~ (?,400) => (?,120)
self.fc2 = nn.Linear(120, 84) # (?,120) => (?,84)
self.fc3 = nn.Linear(84, 10) # (?, 84) => (?,10)
################################################################################
# END OF YOUR CODE #
################################################################################
def forward(self, x):
################################################################################
# TODO: #
# Set up the forward pass that the input data will go through. #
# A good activation function betweent the layers is a ReLu function. #
# #
# Note that the output of the last convolution layer should be flattened #
# before being inputted to the fully connected layer. We can flatten #
# Tensor `x` with `x.view`. #
################################################################################
x = self.pool(F.relu(self.conv1(x)))
x = self.pool(F.relu(self.conv2(x)))
x = torch.flatten(x, 1) # only keep the batch dim, flatten rest dims
# alternatively
# x = x.view(x.size[0], -1)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.fc3(x)
################################################################################
# END OF YOUR CODE #
################################################################################
return x
def predict(self, x):
logits = self.forward(x)
return F.softmax(logits)
net = ConvNet()
################################################################################
# TODO: #
# Choose an Optimizer that will be used to minimize the loss function. #
# Choose a critera that measures the loss #
################################################################################
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
net.to(device)
criterion = nn.CrossEntropyLoss()
optimizer = optim.SGD(net.parameters(), lr=0.005, momentum=0.9)
epochs = 16
steps = 0
running_loss = 0
print_every = 20
for e in range(epochs):
start = time.time()
for images, labels in iter(trainloader):
steps += 1
################################################################################
# TODO: #
# Run the training process #
# #
# #
################################################################################
images, labels = images.to(device), labels.to(device)
optimizer.zero_grad()
outputs = net(images)
################################################################################
# END OF YOUR CODE #
################################################################################
loss = criterion(outputs, labels)
################################################################################
# TODO: #
# Run the training process #
# #
# HINT: Calculate the gradient and move one step further #
################################################################################
loss.backward()
optimizer.step()
################################################################################
# END OF YOUR CODE #
################################################################################
running_loss += loss.item()
if steps % print_every == 0:
stop = time.time()
# Test accuracy
accuracy = 0
for ii, (images, labels) in enumerate(valloader):
################################################################################
# TODO: #
# Calculate the accuracy #
################################################################################
with torch.no_grad():
images, labels = images.to(device), labels.to(device)
outputs = net(images)
_, predicted = torch.max(outputs.data, 1)
# alternatively
# _, predicted = torch.max(F.softmax(outputs,dim=1).data, 1)
accuracy += (predicted == labels).sum().item() / labels.size(0)
################################################################################
# END OF YOUR CODE #
################################################################################
print("Epoch: {}/{}..".format(e+1, epochs),
"Loss: {:.4f}..".format(running_loss/print_every),
"Test accuracy: {:.4f}..".format(accuracy/(ii+1)),
"{:.4f} s/batch".format((stop - start)/print_every)
)
running_loss = 0
start = time.time()
# 04:lr=0.001 Loss: 1.4490.. Test accuracy: 0.4340.. 0.0277 s/batch
# 08:lr=0.001 Loss: 1.3028.. Test accuracy: 0.5350.. 0.0196 s/batch
# 12:lr=0.001 Loss: 1.0942.. Test accuracy: 0.5613.. 0.0192 s/batch
# 16:lr=0.001 Loss: 0.9728.. Test accuracy: 0.6096.. 0.0204 s/batch
# 20:lr=0.001 Loss: 0.9827.. Test accuracy: 0.6143.. 0.0301 s/batch
# 28:lr=0.0005 Loss: 0.7851.. Test accuracy: 0.6361.. 0.0195 s/batch
# 32:lr=0.001 Loss: 0.7912.. Test accuracy: 0.6262.. 0.0205 s/batch
# 40:lr=0.0001 Loss: 0.6549.. Test accuracy: 0.6561.. 0.0208 s/batch
# 56:lr=0.00005L
```
Save best trained model.
```
## You should be familiar with how to save a pytorch model (Make sure to save the model in your Drive)
torch.save(net.state_dict(), 'drive/MyDrive/Colab Notebooks/Cifar10_Classifier_pytorch_CUDA.ckpt')
checkpoint = torch.load("drive/MyDrive/Colab Notebooks/Cifar10_Classifier_pytorch_CUDA.ckpt")
net.load_state_dict(checkpoint)
```
| github_jupyter |
# Informer
### Uses informer model as prediction of future.
```
import os, sys
from tqdm import tqdm
from subseasonal_toolkit.utils.notebook_util import isnotebook
if isnotebook():
# Autoreload packages that are modified
%load_ext autoreload
%autoreload 2
else:
from argparse import ArgumentParser
import pandas as pd
import numpy as np
from scipy.spatial.distance import cdist, euclidean
from datetime import datetime, timedelta
from ttictoc import tic, toc
from subseasonal_data.utils import get_measurement_variable
from subseasonal_toolkit.utils.general_util import printf
from subseasonal_toolkit.utils.experiments_util import get_id_name, get_th_name, get_first_year, get_start_delta
from subseasonal_toolkit.utils.models_util import (get_submodel_name, start_logger, log_params, get_forecast_filename,
save_forecasts)
from subseasonal_toolkit.utils.eval_util import get_target_dates, mean_rmse_to_score, save_metric
from sklearn.linear_model import *
from subseasonal_data import data_loaders
#
# Specify model parameters
#
if not isnotebook():
# If notebook run as a script, parse command-line arguments
parser = ArgumentParser()
parser.add_argument("pos_vars",nargs="*") # gt_id and horizon
parser.add_argument('--target_dates', '-t', default="std_test")
args, opt = parser.parse_known_args()
# Assign variables
gt_id = get_id_name(args.pos_vars[0]) # "contest_precip" or "contest_tmp2m"
horizon = get_th_name(args.pos_vars[1]) # "12w", "34w", or "56w"
target_dates = args.target_dates
else:
# Otherwise, specify arguments interactively
gt_id = "contest_tmp2m"
horizon = "34w"
target_dates = "std_contest"
#
# Process model parameters
#
# One can subtract this number from a target date to find the last viable training date.
start_delta = timedelta(days=get_start_delta(horizon, gt_id))
# Record model and submodel name
model_name = "informer"
submodel_name = get_submodel_name(model_name)
FIRST_SAVE_YEAR = 2007 # Don't save forecasts from years prior to FIRST_SAVE_YEAR
if not isnotebook():
# Save output to log file
logger = start_logger(model=model_name,submodel=submodel_name,gt_id=gt_id,
horizon=horizon,target_dates=target_dates)
# Store parameter values in log
params_names = ['gt_id', 'horizon', 'target_dates']
params_values = [eval(param) for param in params_names]
log_params(params_names, params_values)
printf('Loading target variable and dropping extraneous columns')
tic()
var = get_measurement_variable(gt_id)
gt = data_loaders.get_ground_truth(gt_id).loc[:,["start_date","lat","lon",var]]
toc()
printf('Pivoting dataframe to have one column per lat-lon pair and one row per start_date')
tic()
gt = gt.set_index(['lat','lon','start_date']).squeeze().unstack(['lat','lon'])
toc()
#
# Make predictions for each target date
#
from fbprophet import Prophet
from pandas.tseries.offsets import DateOffset
def get_first_fourth_month(date):
targets = {(1, 31), (5, 31), (9, 30)}
while (date.month, date.day) not in targets:
date = date - DateOffset(days=1)
return date
def get_predictions(date):
# take the first (12/31, 8/31, 4/30) right before the date.
true_date = get_first_fourth_month(date)
true_date_str = true_date.strftime("%Y-%m-%d")
cmd = f"python -u main_informer.py --model informer --data gt-{gt_id}-14d-{horizon} \
--attn prob --features S --start-date {true_date_str} --freq 'd' \
--train_epochs 20 --gpu 0 &"
os.system(cmd) # comment to not run the actual program.
# open the file where this is outputted.
folder_name = f"results/gt-{gt_id}-14d-{horizon}_{true_date_str}_informer_gt-{gt_id}-14d-{horizon}_ftM_sl192_ll96_pl48_dm512_nh8_el3_dl2_df1024_atprob_ebtimeF_dtTrue_test_0/"
# return the answer.
dates = np.load(folder_name + "dates.npy")
preds = np.load(folder_name + "preds.npy")
idx = -1
for i in range(dates):
if dates[i] == date:
idx = i
return preds[idx]
tic()
target_date_objs = pd.Series(get_target_dates(date_str=target_dates,horizon=horizon))
rmses = pd.Series(index=target_date_objs, dtype=np.float64)
preds = pd.DataFrame(index = target_date_objs, columns = gt.columns,
dtype=np.float64)
preds.index.name = "start_date"
# Sort target_date_objs by day of week
target_date_objs = target_date_objs[target_date_objs.dt.weekday.argsort(kind='stable')]
toc()
for target_date_obj in target_date_objs:
tic()
target_date_str = datetime.strftime(target_date_obj, '%Y%m%d')
# Find the last observable training date for this target
last_train_date = target_date_obj - start_delta
if not last_train_date in gt.index:
printf(f'-Warning: no persistence prediction for {target_date_str}; skipping')
continue
printf(f'Forming persistence prediction for {target_date_obj}')
# key logic here:
preds.loc[target_date_obj,:] = get_predictions(target_date_obj)
# Save prediction to file in standard format
if target_date_obj.year >= FIRST_SAVE_YEAR:
save_forecasts(
preds.loc[[target_date_obj],:].unstack().rename("pred").reset_index(),
model=model_name, submodel=submodel_name,
gt_id=gt_id, horizon=horizon,
target_date_str=target_date_str)
# Evaluate and store error if we have ground truth data
if target_date_obj in gt.index:
rmse = np.sqrt(np.square(preds.loc[target_date_obj,:] - gt.loc[target_date_obj,:]).mean())
rmses.loc[target_date_obj] = rmse
print("-rmse: {}, score: {}".format(rmse, mean_rmse_to_score(rmse)))
mean_rmse = rmses.mean()
print("-mean rmse: {}, running score: {}".format(mean_rmse, mean_rmse_to_score(mean_rmse)))
toc()
printf("Save rmses in standard format")
rmses = rmses.sort_index().reset_index()
rmses.columns = ['start_date','rmse']
save_metric(rmses, model=model_name, submodel=submodel_name, gt_id=gt_id, horizon=horizon, target_dates=target_dates, metric="rmse")
```
| github_jupyter |
# TensorFlow Tutorial
Welcome to this week's programming assignment. Until now, you've always used numpy to build neural networks. Now we will step you through a deep learning framework that will allow you to build neural networks more easily. Machine learning frameworks like TensorFlow, PaddlePaddle, Torch, Caffe, Keras, and many others can speed up your machine learning development significantly. All of these frameworks also have a lot of documentation, which you should feel free to read. In this assignment, you will learn to do the following in TensorFlow:
- Initialize variables
- Start your own session
- Train algorithms
- Implement a Neural Network
Programing frameworks can not only shorten your coding time, but sometimes also perform optimizations that speed up your code.
## <font color='darkblue'>Updates</font>
#### If you were working on the notebook before this update...
* The current notebook is version "v3b".
* You can find your original work saved in the notebook with the previous version name (it may be either TensorFlow Tutorial version 3" or "TensorFlow Tutorial version 3a.)
* To view the file directory, click on the "Coursera" icon in the top left of this notebook.
#### List of updates
* forward_propagation instruction now says 'A1' instead of 'a1' in the formula for Z2;
and are updated to say 'A2' instead of 'Z2' in the formula for Z3.
* create_placeholders instruction refer to the data type "tf.float32" instead of float.
* in the model function, the x axis of the plot now says "iterations (per fives)" instead of iterations(per tens)
* In the linear_function, comments remind students to create the variables in the order suggested by the starter code. The comments are updated to reflect this order.
* The test of the cost function now creates the logits without passing them through a sigmoid function (since the cost function will include the sigmoid in the built-in tensorflow function).
* In the 'model' function, the minibatch_cost is now divided by minibatch_size (instead of num_minibatches).
* Updated print statements and 'expected output that are used to check functions, for easier visual comparison.
## 1 - Exploring the Tensorflow Library
To start, you will import the library:
```
import math
import numpy as np
import h5py
import matplotlib.pyplot as plt
import tensorflow as tf
from tensorflow.python.framework import ops
from tf_utils import load_dataset, random_mini_batches, convert_to_one_hot, predict
%matplotlib inline
np.random.seed(1)
```
Now that you have imported the library, we will walk you through its different applications. You will start with an example, where we compute for you the loss of one training example.
$$loss = \mathcal{L}(\hat{y}, y) = (\hat y^{(i)} - y^{(i)})^2 \tag{1}$$
```
y_hat = tf.constant(36, name='y_hat') # Define y_hat constant. Set to 36.
y = tf.constant(39, name='y') # Define y. Set to 39
loss = tf.Variable((y - y_hat)**2, name='loss') # Create a variable for the loss
init = tf.global_variables_initializer() # When init is run later (session.run(init)),
# the loss variable will be initialized and ready to be computed
with tf.Session() as session: # Create a session and print the output
session.run(init) # Initializes the variables
print(session.run(loss)) # Prints the loss
```
Writing and running programs in TensorFlow has the following steps:
1. Create Tensors (variables) that are not yet executed/evaluated.
2. Write operations between those Tensors.
3. Initialize your Tensors.
4. Create a Session.
5. Run the Session. This will run the operations you'd written above.
Therefore, when we created a variable for the loss, we simply defined the loss as a function of other quantities, but did not evaluate its value. To evaluate it, we had to run `init=tf.global_variables_initializer()`. That initialized the loss variable, and in the last line we were finally able to evaluate the value of `loss` and print its value.
Now let us look at an easy example. Run the cell below:
```
a = tf.constant(2)
b = tf.constant(10)
c = tf.multiply(a,b)
print(c)
```
As expected, you will not see 20! You got a tensor saying that the result is a tensor that does not have the shape attribute, and is of type "int32". All you did was put in the 'computation graph', but you have not run this computation yet. In order to actually multiply the two numbers, you will have to create a session and run it.
```
sess = tf.Session()
print(sess.run(c))
```
Great! To summarize, **remember to initialize your variables, create a session and run the operations inside the session**.
Next, you'll also have to know about placeholders. A placeholder is an object whose value you can specify only later.
To specify values for a placeholder, you can pass in values by using a "feed dictionary" (`feed_dict` variable). Below, we created a placeholder for x. This allows us to pass in a number later when we run the session.
```
# Change the value of x in the feed_dict
x = tf.placeholder(tf.int64, name = 'x')
print(sess.run(2 * x, feed_dict = {x: 3}))
sess.close()
```
When you first defined `x` you did not have to specify a value for it. A placeholder is simply a variable that you will assign data to only later, when running the session. We say that you **feed data** to these placeholders when running the session.
Here's what's happening: When you specify the operations needed for a computation, you are telling TensorFlow how to construct a computation graph. The computation graph can have some placeholders whose values you will specify only later. Finally, when you run the session, you are telling TensorFlow to execute the computation graph.
### 1.1 - Linear function
Lets start this programming exercise by computing the following equation: $Y = WX + b$, where $W$ and $X$ are random matrices and b is a random vector.
**Exercise**: Compute $WX + b$ where $W, X$, and $b$ are drawn from a random normal distribution. W is of shape (4, 3), X is (3,1) and b is (4,1). As an example, here is how you would define a constant X that has shape (3,1):
```python
X = tf.constant(np.random.randn(3,1), name = "X")
```
You might find the following functions helpful:
- tf.matmul(..., ...) to do a matrix multiplication
- tf.add(..., ...) to do an addition
- np.random.randn(...) to initialize randomly
```
# GRADED FUNCTION: linear_function
def linear_function():
"""
Implements a linear function:
Initializes X to be a random tensor of shape (3,1)
Initializes W to be a random tensor of shape (4,3)
Initializes b to be a random tensor of shape (4,1)
Returns:
result -- runs the session for Y = WX + b
"""
np.random.seed(1)
"""
Note, to ensure that the "random" numbers generated match the expected results,
please create the variables in the order given in the starting code below.
(Do not re-arrange the order).
"""
### START CODE HERE ### (4 lines of code)
X = tf.constant(np.random.randn(3,1), name = "X")
W = tf.constant(np.random.randn(4,3), name = "W")
b = tf.constant(np.random.randn(4,1), name = "b")
Y = tf.add(tf.matmul(W,X), b)
### END CODE HERE ###
# Create the session using tf.Session() and run it with sess.run(...) on the variable you want to calculate
### START CODE HERE ###
sess = tf.Session()
result = sess.run(Y)
### END CODE HERE ###
# close the session
sess.close()
return result
print( "result = \n" + str(linear_function()))
```
*** Expected Output ***:
```
result =
[[-2.15657382]
[ 2.95891446]
[-1.08926781]
[-0.84538042]]
```
### 1.2 - Computing the sigmoid
Great! You just implemented a linear function. Tensorflow offers a variety of commonly used neural network functions like `tf.sigmoid` and `tf.softmax`. For this exercise lets compute the sigmoid function of an input.
You will do this exercise using a placeholder variable `x`. When running the session, you should use the feed dictionary to pass in the input `z`. In this exercise, you will have to (i) create a placeholder `x`, (ii) define the operations needed to compute the sigmoid using `tf.sigmoid`, and then (iii) run the session.
** Exercise **: Implement the sigmoid function below. You should use the following:
- `tf.placeholder(tf.float32, name = "...")`
- `tf.sigmoid(...)`
- `sess.run(..., feed_dict = {x: z})`
Note that there are two typical ways to create and use sessions in tensorflow:
**Method 1:**
```python
sess = tf.Session()
# Run the variables initialization (if needed), run the operations
result = sess.run(..., feed_dict = {...})
sess.close() # Close the session
```
**Method 2:**
```python
with tf.Session() as sess:
# run the variables initialization (if needed), run the operations
result = sess.run(..., feed_dict = {...})
# This takes care of closing the session for you :)
```
```
# GRADED FUNCTION: sigmoid
def sigmoid(z):
"""
Computes the sigmoid of z
Arguments:
z -- input value, scalar or vector
Returns:
results -- the sigmoid of z
"""
### START CODE HERE ### ( approx. 4 lines of code)
# Create a placeholder for x. Name it 'x'.
x = tf.placeholder(tf.float32, name="x")
# compute sigmoid(x)
sigmoid = tf.sigmoid(x)
# Create a session, and run it. Please use the method 2 explained above.
# You should use a feed_dict to pass z's value to x.
sess = tf.Session()
# Run the variables initialization (if needed), run the operations
result = sess.run(sigmoid, feed_dict = {x: z})
sess.close() # Close the session
### END CODE HERE ###
return result
print ("sigmoid(0) = " + str(sigmoid(0)))
print ("sigmoid(12) = " + str(sigmoid(12)))
```
*** Expected Output ***:
<table>
<tr>
<td>
**sigmoid(0)**
</td>
<td>
0.5
</td>
</tr>
<tr>
<td>
**sigmoid(12)**
</td>
<td>
0.999994
</td>
</tr>
</table>
<font color='blue'>
**To summarize, you how know how to**:
1. Create placeholders
2. Specify the computation graph corresponding to operations you want to compute
3. Create the session
4. Run the session, using a feed dictionary if necessary to specify placeholder variables' values.
### 1.3 - Computing the Cost
You can also use a built-in function to compute the cost of your neural network. So instead of needing to write code to compute this as a function of $a^{[2](i)}$ and $y^{(i)}$ for i=1...m:
$$ J = - \frac{1}{m} \sum_{i = 1}^m \large ( \small y^{(i)} \log a^{ [2] (i)} + (1-y^{(i)})\log (1-a^{ [2] (i)} )\large )\small\tag{2}$$
you can do it in one line of code in tensorflow!
**Exercise**: Implement the cross entropy loss. The function you will use is:
- `tf.nn.sigmoid_cross_entropy_with_logits(logits = ..., labels = ...)`
Your code should input `z`, compute the sigmoid (to get `a`) and then compute the cross entropy cost $J$. All this can be done using one call to `tf.nn.sigmoid_cross_entropy_with_logits`, which computes
$$- \frac{1}{m} \sum_{i = 1}^m \large ( \small y^{(i)} \log \sigma(z^{[2](i)}) + (1-y^{(i)})\log (1-\sigma(z^{[2](i)})\large )\small\tag{2}$$
```
# GRADED FUNCTION: cost
def cost(logits, labels):
"""
Computes the cost using the sigmoid cross entropy
Arguments:
logits -- vector containing z, output of the last linear unit (before the final sigmoid activation)
labels -- vector of labels y (1 or 0)
Note: What we've been calling "z" and "y" in this class are respectively called "logits" and "labels"
in the TensorFlow documentation. So logits will feed into z, and labels into y.
Returns:
cost -- runs the session of the cost (formula (2))
"""
### START CODE HERE ###
# Create the placeholders for "logits" (z) and "labels" (y) (approx. 2 lines)
z = tf.placeholder(tf.float32, name="z")
y = tf.placeholder(tf.float32, name="y")
# Use the loss function (approx. 1 line)
cost = tf.nn.sigmoid_cross_entropy_with_logits(logits = z, labels = y)
# Create a session (approx. 1 line). See method 1 above.
sess = tf.Session()
# Run the session (approx. 1 line).
cost = sess.run(cost, feed_dict={z: logits, y: labels})
# Close the session (approx. 1 line). See method 1 above.
sess.close()
### END CODE HERE ###
return cost
logits = np.array([0.2,0.4,0.7,0.9])
cost = cost(logits, np.array([0,0,1,1]))
print ("cost = " + str(cost))
```
** Expected Output** :
```
cost = [ 0.79813886 0.91301525 0.40318605 0.34115386]
```
### 1.4 - Using One Hot encodings
Many times in deep learning you will have a y vector with numbers ranging from 0 to C-1, where C is the number of classes. If C is for example 4, then you might have the following y vector which you will need to convert as follows:
<img src="images/onehot.png" style="width:600px;height:150px;">
This is called a "one hot" encoding, because in the converted representation exactly one element of each column is "hot" (meaning set to 1). To do this conversion in numpy, you might have to write a few lines of code. In tensorflow, you can use one line of code:
- tf.one_hot(labels, depth, axis)
**Exercise:** Implement the function below to take one vector of labels and the total number of classes $C$, and return the one hot encoding. Use `tf.one_hot()` to do this.
```
# GRADED FUNCTION: one_hot_matrix
def one_hot_matrix(labels, C):
"""
Creates a matrix where the i-th row corresponds to the ith class number and the jth column
corresponds to the jth training example. So if example j had a label i. Then entry (i,j)
will be 1.
Arguments:
labels -- vector containing the labels
C -- number of classes, the depth of the one hot dimension
Returns:
one_hot -- one hot matrix
"""
### START CODE HERE ###
# Create a tf.constant equal to C (depth), name it 'C'. (approx. 1 line)
C = tf.constant(C, name="C")
# Use tf.one_hot, be careful with the axis (approx. 1 line)
one_hot_matrix = tf.one_hot(indices=labels, depth=C, axis=0)
# Create the session (approx. 1 line)
sess = tf.Session()
# Run the session (approx. 1 line)
one_hot = sess.run(one_hot_matrix)
# Close the session (approx. 1 line). See method 1 above.
sess.close()
### END CODE HERE ###
return one_hot
labels = np.array([1,2,3,0,2,1])
one_hot = one_hot_matrix(labels, C = 4)
print ("one_hot = \n" + str(one_hot))
```
**Expected Output**:
```
one_hot =
[[ 0. 0. 0. 1. 0. 0.]
[ 1. 0. 0. 0. 0. 1.]
[ 0. 1. 0. 0. 1. 0.]
[ 0. 0. 1. 0. 0. 0.]]
```
### 1.5 - Initialize with zeros and ones
Now you will learn how to initialize a vector of zeros and ones. The function you will be calling is `tf.ones()`. To initialize with zeros you could use tf.zeros() instead. These functions take in a shape and return an array of dimension shape full of zeros and ones respectively.
**Exercise:** Implement the function below to take in a shape and to return an array (of the shape's dimension of ones).
- tf.ones(shape)
```
# GRADED FUNCTION: ones
def ones(shape):
"""
Creates an array of ones of dimension shape
Arguments:
shape -- shape of the array you want to create
Returns:
ones -- array containing only ones
"""
### START CODE HERE ###
# Create "ones" tensor using tf.ones(...). (approx. 1 line)
ones = tf.ones(1,3)
# Create the session (approx. 1 line)
sess = tf.Session()
# Run the session to compute 'ones' (approx. 1 line)
ones = sess.run(ones)
# Close the session (approx. 1 line). See method 1 above.
sess.close()
### END CODE HERE ###
return ones
print ("ones = " + str(ones([3])))
```
**Expected Output:**
<table>
<tr>
<td>
**ones**
</td>
<td>
[ 1. 1. 1.]
</td>
</tr>
</table>
# 2 - Building your first neural network in tensorflow
In this part of the assignment you will build a neural network using tensorflow. Remember that there are two parts to implement a tensorflow model:
- Create the computation graph
- Run the graph
Let's delve into the problem you'd like to solve!
### 2.0 - Problem statement: SIGNS Dataset
One afternoon, with some friends we decided to teach our computers to decipher sign language. We spent a few hours taking pictures in front of a white wall and came up with the following dataset. It's now your job to build an algorithm that would facilitate communications from a speech-impaired person to someone who doesn't understand sign language.
- **Training set**: 1080 pictures (64 by 64 pixels) of signs representing numbers from 0 to 5 (180 pictures per number).
- **Test set**: 120 pictures (64 by 64 pixels) of signs representing numbers from 0 to 5 (20 pictures per number).
Note that this is a subset of the SIGNS dataset. The complete dataset contains many more signs.
Here are examples for each number, and how an explanation of how we represent the labels. These are the original pictures, before we lowered the image resolutoion to 64 by 64 pixels.
<img src="images/hands.png" style="width:800px;height:350px;"><caption><center> <u><font color='purple'> **Figure 1**</u><font color='purple'>: SIGNS dataset <br> <font color='black'> </center>
Run the following code to load the dataset.
```
# Loading the dataset
X_train_orig, Y_train_orig, X_test_orig, Y_test_orig, classes = load_dataset()
```
Change the index below and run the cell to visualize some examples in the dataset.
```
# Example of a picture
index = 0
plt.imshow(X_train_orig[index])
print ("y = " + str(np.squeeze(Y_train_orig[:, index])))
```
As usual you flatten the image dataset, then normalize it by dividing by 255. On top of that, you will convert each label to a one-hot vector as shown in Figure 1. Run the cell below to do so.
```
# Flatten the training and test images
X_train_flatten = X_train_orig.reshape(X_train_orig.shape[0], -1).T
X_test_flatten = X_test_orig.reshape(X_test_orig.shape[0], -1).T
# Normalize image vectors
X_train = X_train_flatten/255.
X_test = X_test_flatten/255.
# Convert training and test labels to one hot matrices
Y_train = convert_to_one_hot(Y_train_orig, 6)
Y_test = convert_to_one_hot(Y_test_orig, 6)
print ("number of training examples = " + str(X_train.shape[1]))
print ("number of test examples = " + str(X_test.shape[1]))
print ("X_train shape: " + str(X_train.shape))
print ("Y_train shape: " + str(Y_train.shape))
print ("X_test shape: " + str(X_test.shape))
print ("Y_test shape: " + str(Y_test.shape))
```
**Note** that 12288 comes from $64 \times 64 \times 3$. Each image is square, 64 by 64 pixels, and 3 is for the RGB colors. Please make sure all these shapes make sense to you before continuing.
**Your goal** is to build an algorithm capable of recognizing a sign with high accuracy. To do so, you are going to build a tensorflow model that is almost the same as one you have previously built in numpy for cat recognition (but now using a softmax output). It is a great occasion to compare your numpy implementation to the tensorflow one.
**The model** is *LINEAR -> RELU -> LINEAR -> RELU -> LINEAR -> SOFTMAX*. The SIGMOID output layer has been converted to a SOFTMAX. A SOFTMAX layer generalizes SIGMOID to when there are more than two classes.
### 2.1 - Create placeholders
Your first task is to create placeholders for `X` and `Y`. This will allow you to later pass your training data in when you run your session.
**Exercise:** Implement the function below to create the placeholders in tensorflow.
```
# GRADED FUNCTION: create_placeholders
def create_placeholders(n_x, n_y):
"""
Creates the placeholders for the tensorflow session.
Arguments:
n_x -- scalar, size of an image vector (num_px * num_px = 64 * 64 * 3 = 12288)
n_y -- scalar, number of classes (from 0 to 5, so -> 6)
Returns:
X -- placeholder for the data input, of shape [n_x, None] and dtype "tf.float32"
Y -- placeholder for the input labels, of shape [n_y, None] and dtype "tf.float32"
Tips:
- You will use None because it let's us be flexible on the number of examples you will for the placeholders.
In fact, the number of examples during test/train is different.
"""
### START CODE HERE ### (approx. 2 lines)
X = tf.placeholder(tf.float32, name="n_x")
Y = tf.placeholder(tf.float32, name="n_y")
### END CODE HERE ###
return X, Y
X, Y = create_placeholders(12288, 6)
print ("X = " + str(X))
print ("Y = " + str(Y))
```
**Expected Output**:
<table>
<tr>
<td>
**X**
</td>
<td>
Tensor("Placeholder_1:0", shape=(12288, ?), dtype=float32) (not necessarily Placeholder_1)
</td>
</tr>
<tr>
<td>
**Y**
</td>
<td>
Tensor("Placeholder_2:0", shape=(6, ?), dtype=float32) (not necessarily Placeholder_2)
</td>
</tr>
</table>
### 2.2 - Initializing the parameters
Your second task is to initialize the parameters in tensorflow.
**Exercise:** Implement the function below to initialize the parameters in tensorflow. You are going use Xavier Initialization for weights and Zero Initialization for biases. The shapes are given below. As an example, to help you, for W1 and b1 you could use:
```python
W1 = tf.get_variable("W1", [25,12288], initializer = tf.contrib.layers.xavier_initializer(seed = 1))
b1 = tf.get_variable("b1", [25,1], initializer = tf.zeros_initializer())
```
Please use `seed = 1` to make sure your results match ours.
```
# GRADED FUNCTION: initialize_parameters
def initialize_parameters():
"""
Initializes parameters to build a neural network with tensorflow. The shapes are:
W1 : [25, 12288]
b1 : [25, 1]
W2 : [12, 25]
b2 : [12, 1]
W3 : [6, 12]
b3 : [6, 1]
Returns:
parameters -- a dictionary of tensors containing W1, b1, W2, b2, W3, b3
"""
tf.set_random_seed(1) # so that your "random" numbers match ours
### START CODE HERE ### (approx. 6 lines of code)
W1 = tf.get_variable("W1", [25,12288], initializer = tf.contrib.layers.xavier_initializer(seed = 1))
b1 = tf.get_variable("b1", [25,1], initializer = tf.zeros_initializer())
W2 = tf.get_variable("W2", [12, 25], initializer = tf.contrib.layers.xavier_initializer(seed = 1))
b2 = tf.get_variable("b2", [12,1], initializer = tf.zeros_initializer())
W3 = tf.get_variable("W3", [6, 12], initializer = tf.contrib.layers.xavier_initializer(seed = 1))
b3 = tf.get_variable("b3", [6, 1], initializer = tf.zeros_initializer())
### END CODE HERE ###
parameters = {"W1": W1,
"b1": b1,
"W2": W2,
"b2": b2,
"W3": W3,
"b3": b3}
return parameters
tf.reset_default_graph()
with tf.Session() as sess:
parameters = initialize_parameters()
print("W1 = " + str(parameters["W1"]))
print("b1 = " + str(parameters["b1"]))
print("W2 = " + str(parameters["W2"]))
print("b2 = " + str(parameters["b2"]))
```
**Expected Output**:
<table>
<tr>
<td>
**W1**
</td>
<td>
< tf.Variable 'W1:0' shape=(25, 12288) dtype=float32_ref >
</td>
</tr>
<tr>
<td>
**b1**
</td>
<td>
< tf.Variable 'b1:0' shape=(25, 1) dtype=float32_ref >
</td>
</tr>
<tr>
<td>
**W2**
</td>
<td>
< tf.Variable 'W2:0' shape=(12, 25) dtype=float32_ref >
</td>
</tr>
<tr>
<td>
**b2**
</td>
<td>
< tf.Variable 'b2:0' shape=(12, 1) dtype=float32_ref >
</td>
</tr>
</table>
As expected, the parameters haven't been evaluated yet.
### 2.3 - Forward propagation in tensorflow
You will now implement the forward propagation module in tensorflow. The function will take in a dictionary of parameters and it will complete the forward pass. The functions you will be using are:
- `tf.add(...,...)` to do an addition
- `tf.matmul(...,...)` to do a matrix multiplication
- `tf.nn.relu(...)` to apply the ReLU activation
**Question:** Implement the forward pass of the neural network. We commented for you the numpy equivalents so that you can compare the tensorflow implementation to numpy. It is important to note that the forward propagation stops at `z3`. The reason is that in tensorflow the last linear layer output is given as input to the function computing the loss. Therefore, you don't need `a3`!
```
# GRADED FUNCTION: forward_propagation
def forward_propagation(X, parameters):
"""
Implements the forward propagation for the model: LINEAR -> RELU -> LINEAR -> RELU -> LINEAR -> SOFTMAX
Arguments:
X -- input dataset placeholder, of shape (input size, number of examples)
parameters -- python dictionary containing your parameters "W1", "b1", "W2", "b2", "W3", "b3"
the shapes are given in initialize_parameters
Returns:
Z3 -- the output of the last LINEAR unit
"""
# Retrieve the parameters from the dictionary "parameters"
W1 = parameters['W1']
b1 = parameters['b1']
W2 = parameters['W2']
b2 = parameters['b2']
W3 = parameters['W3']
b3 = parameters['b3']
### START CODE HERE ### (approx. 5 lines) # Numpy Equivalents:
Z1 = tf.add(tf.matmul(W1, X), b1) # Z1 = np.dot(W1, X) + b1
A1 = tf.nn.relu(Z1) # A1 = relu(Z1)
Z2 = tf.add(tf.matmul(W2, A1), b2) # Z2 = np.dot(W2, A1) + b2
A2 = tf.nn.relu(Z2) # A2 = relu(Z2)
Z3 = tf.add(tf.matmul(W3, A2), b3) # Z3 = np.dot(W3, A2) + b3
### END CODE HERE ###
return Z3
tf.reset_default_graph()
with tf.Session() as sess:
X, Y = create_placeholders(12288, 6)
parameters = initialize_parameters()
Z3 = forward_propagation(X, parameters)
print("Z3 = " + str(Z3))
```
**Expected Output**:
<table>
<tr>
<td>
**Z3**
</td>
<td>
Tensor("Add_2:0", shape=(6, ?), dtype=float32)
</td>
</tr>
</table>
You may have noticed that the forward propagation doesn't output any cache. You will understand why below, when we get to brackpropagation.
### 2.4 Compute cost
As seen before, it is very easy to compute the cost using:
```python
tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits = ..., labels = ...))
```
**Question**: Implement the cost function below.
- It is important to know that the "`logits`" and "`labels`" inputs of `tf.nn.softmax_cross_entropy_with_logits` are expected to be of shape (number of examples, num_classes). We have thus transposed Z3 and Y for you.
- Besides, `tf.reduce_mean` basically does the summation over the examples.
```
# GRADED FUNCTION: compute_cost
def compute_cost(Z3, Y):
"""
Computes the cost
Arguments:
Z3 -- output of forward propagation (output of the last LINEAR unit), of shape (6, number of examples)
Y -- "true" labels vector placeholder, same shape as Z3
Returns:
cost - Tensor of the cost function
"""
# to fit the tensorflow requirement for tf.nn.softmax_cross_entropy_with_logits(...,...)
logits = tf.transpose(Z3)
labels = tf.transpose(Y)
### START CODE HERE ### (1 line of code)
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits = logits, labels = labels))
### END CODE HERE ###
return cost
tf.reset_default_graph()
with tf.Session() as sess:
X, Y = create_placeholders(12288, 6)
parameters = initialize_parameters()
Z3 = forward_propagation(X, parameters)
cost = compute_cost(Z3, Y)
print("cost = " + str(cost))
```
**Expected Output**:
<table>
<tr>
<td>
**cost**
</td>
<td>
Tensor("Mean:0", shape=(), dtype=float32)
</td>
</tr>
</table>
### 2.5 - Backward propagation & parameter updates
This is where you become grateful to programming frameworks. All the backpropagation and the parameters update is taken care of in 1 line of code. It is very easy to incorporate this line in the model.
After you compute the cost function. You will create an "`optimizer`" object. You have to call this object along with the cost when running the tf.session. When called, it will perform an optimization on the given cost with the chosen method and learning rate.
For instance, for gradient descent the optimizer would be:
```python
optimizer = tf.train.GradientDescentOptimizer(learning_rate = learning_rate).minimize(cost)
```
To make the optimization you would do:
```python
_ , c = sess.run([optimizer, cost], feed_dict={X: minibatch_X, Y: minibatch_Y})
```
This computes the backpropagation by passing through the tensorflow graph in the reverse order. From cost to inputs.
**Note** When coding, we often use `_` as a "throwaway" variable to store values that we won't need to use later. Here, `_` takes on the evaluated value of `optimizer`, which we don't need (and `c` takes the value of the `cost` variable).
### 2.6 - Building the model
Now, you will bring it all together!
**Exercise:** Implement the model. You will be calling the functions you had previously implemented.
```
def model(X_train, Y_train, X_test, Y_test, learning_rate = 0.0001,
num_epochs = 1500, minibatch_size = 32, print_cost = True):
"""
Implements a three-layer tensorflow neural network: LINEAR->RELU->LINEAR->RELU->LINEAR->SOFTMAX.
Arguments:
X_train -- training set, of shape (input size = 12288, number of training examples = 1080)
Y_train -- test set, of shape (output size = 6, number of training examples = 1080)
X_test -- training set, of shape (input size = 12288, number of training examples = 120)
Y_test -- test set, of shape (output size = 6, number of test examples = 120)
learning_rate -- learning rate of the optimization
num_epochs -- number of epochs of the optimization loop
minibatch_size -- size of a minibatch
print_cost -- True to print the cost every 100 epochs
Returns:
parameters -- parameters learnt by the model. They can then be used to predict.
"""
ops.reset_default_graph() # to be able to rerun the model without overwriting tf variables
tf.set_random_seed(1) # to keep consistent results
seed = 3 # to keep consistent results
(n_x, m) = X_train.shape # (n_x: input size, m : number of examples in the train set)
n_y = Y_train.shape[0] # n_y : output size
costs = [] # To keep track of the cost
# Create Placeholders of shape (n_x, n_y)
### START CODE HERE ### (1 line)
X = tf.placeholder(tf.float32, name="n_x")
Y = tf.placeholder(tf.float32, name="n_y")
### END CODE HERE ###
# Initialize parameters
### START CODE HERE ### (1 line)
parameters = None
### END CODE HERE ###
# Forward propagation: Build the forward propagation in the tensorflow graph
### START CODE HERE ### (1 line)
Z3 = None
### END CODE HERE ###
# Cost function: Add cost function to tensorflow graph
### START CODE HERE ### (1 line)
cost = None
### END CODE HERE ###
# Backpropagation: Define the tensorflow optimizer. Use an AdamOptimizer.
### START CODE HERE ### (1 line)
optimizer = None
### END CODE HERE ###
# Initialize all the variables
init = tf.global_variables_initializer()
# Start the session to compute the tensorflow graph
with tf.Session() as sess:
# Run the initialization
sess.run(init)
# Do the training loop
for epoch in range(num_epochs):
epoch_cost = 0. # Defines a cost related to an epoch
num_minibatches = int(m / minibatch_size) # number of minibatches of size minibatch_size in the train set
seed = seed + 1
minibatches = random_mini_batches(X_train, Y_train, minibatch_size, seed)
for minibatch in minibatches:
# Select a minibatch
(minibatch_X, minibatch_Y) = minibatch
# IMPORTANT: The line that runs the graph on a minibatch.
# Run the session to execute the "optimizer" and the "cost", the feedict should contain a minibatch for (X,Y).
### START CODE HERE ### (1 line)
_ , minibatch_cost = None
### END CODE HERE ###
epoch_cost += minibatch_cost / minibatch_size
# Print the cost every epoch
if print_cost == True and epoch % 100 == 0:
print ("Cost after epoch %i: %f" % (epoch, epoch_cost))
if print_cost == True and epoch % 5 == 0:
costs.append(epoch_cost)
# plot the cost
plt.plot(np.squeeze(costs))
plt.ylabel('cost')
plt.xlabel('iterations (per fives)')
plt.title("Learning rate =" + str(learning_rate))
plt.show()
# lets save the parameters in a variable
parameters = sess.run(parameters)
print ("Parameters have been trained!")
# Calculate the correct predictions
correct_prediction = tf.equal(tf.argmax(Z3), tf.argmax(Y))
# Calculate accuracy on the test set
accuracy = tf.reduce_mean(tf.cast(correct_prediction, "float"))
print ("Train Accuracy:", accuracy.eval({X: X_train, Y: Y_train}))
print ("Test Accuracy:", accuracy.eval({X: X_test, Y: Y_test}))
return parameters
```
Run the following cell to train your model! On our machine it takes about 5 minutes. Your "Cost after epoch 100" should be 1.048222. If it's not, don't waste time; interrupt the training by clicking on the square (⬛) in the upper bar of the notebook, and try to correct your code. If it is the correct cost, take a break and come back in 5 minutes!
```
parameters = model(X_train, Y_train, X_test, Y_test)
```
**Expected Output**:
<table>
<tr>
<td>
**Train Accuracy**
</td>
<td>
0.999074
</td>
</tr>
<tr>
<td>
**Test Accuracy**
</td>
<td>
0.716667
</td>
</tr>
</table>
Amazing, your algorithm can recognize a sign representing a figure between 0 and 5 with 71.7% accuracy.
**Insights**:
- Your model seems big enough to fit the training set well. However, given the difference between train and test accuracy, you could try to add L2 or dropout regularization to reduce overfitting.
- Think about the session as a block of code to train the model. Each time you run the session on a minibatch, it trains the parameters. In total you have run the session a large number of times (1500 epochs) until you obtained well trained parameters.
### 2.7 - Test with your own image (optional / ungraded exercise)
Congratulations on finishing this assignment. You can now take a picture of your hand and see the output of your model. To do that:
1. Click on "File" in the upper bar of this notebook, then click "Open" to go on your Coursera Hub.
2. Add your image to this Jupyter Notebook's directory, in the "images" folder
3. Write your image's name in the following code
4. Run the code and check if the algorithm is right!
```
import scipy
from PIL import Image
from scipy import ndimage
## START CODE HERE ## (PUT YOUR IMAGE NAME)
my_image = "thumbs_up.jpg"
## END CODE HERE ##
# We preprocess your image to fit your algorithm.
fname = "images/" + my_image
image = np.array(ndimage.imread(fname, flatten=False))
image = image/255.
my_image = scipy.misc.imresize(image, size=(64,64)).reshape((1, 64*64*3)).T
my_image_prediction = predict(my_image, parameters)
plt.imshow(image)
print("Your algorithm predicts: y = " + str(np.squeeze(my_image_prediction)))
```
You indeed deserved a "thumbs-up" although as you can see the algorithm seems to classify it incorrectly. The reason is that the training set doesn't contain any "thumbs-up", so the model doesn't know how to deal with it! We call that a "mismatched data distribution" and it is one of the various of the next course on "Structuring Machine Learning Projects".
<font color='blue'>
**What you should remember**:
- Tensorflow is a programming framework used in deep learning
- The two main object classes in tensorflow are Tensors and Operators.
- When you code in tensorflow you have to take the following steps:
- Create a graph containing Tensors (Variables, Placeholders ...) and Operations (tf.matmul, tf.add, ...)
- Create a session
- Initialize the session
- Run the session to execute the graph
- You can execute the graph multiple times as you've seen in model()
- The backpropagation and optimization is automatically done when running the session on the "optimizer" object.
| github_jupyter |
```
import sys, os
import argparse
sys.path.insert(0, os.path.abspath(os.path.join(os.getcwd(), "./../../../../")))
sys.path.insert(0, os.path.abspath(os.path.join(os.getcwd(), "./../../../")))
sys.path.insert(0, os.path.abspath(os.path.join(os.getcwd(), "")))
from fedml_api.data_preprocessing.cifar10.data_loader import load_partition_data_cifar10
from fedml_api.standalone.fedavg.my_model_trainer_classification import MyModelTrainer as MyModelTrainerCLS
# from fedml_api.model.cv.resnet import resnet56
from fedml_api.model.contrastive_cv.resnet_with_embedding import Resnet56
import torch
from torch import nn
from collections import OrderedDict
import torch.nn.functional as F
import matplotlib.pyplot as plt
from sklearn.manifold import TSNE
import numpy as np
import random
import pickle
dataset = 'cifar10'
data_dir = "./../../../data/cifar10"
partition_method = 'hetero'
# partition_method = 'homo'
partition_alpha = 100
client_num_in_total = 10
batch_size = 100
total_epochs = 500
save_model_path = 'model/client_{0}_triplet_epochs_{1}.pt'
device = 'cuda:1'
train_data_num, test_data_num, train_data_global, test_data_global, \
train_data_local_num_dict, train_data_local_dict, test_data_local_dict, \
class_num,traindata_cls_counts = load_partition_data_cifar10(dataset, data_dir, partition_method,
partition_alpha, client_num_in_total, batch_size)
dataset = [train_data_num, test_data_num, train_data_global, test_data_global, \
train_data_local_num_dict, train_data_local_dict, test_data_local_dict, \
class_num,traindata_cls_counts]
with open(f'dataset_{partition_method}_{partition_alpha}_{client_num_in_total}_with_cls_counts.pickle', 'wb') as f:
pickle.dump(dataset, f)
# pickle.save()
# print(train_data_num)
# print(train_data_local_dict)
with open(f'dataset_{partition_method}_{client_num_in_total}.pickle', 'rb') as f:
dataset = pickle.load(f)
class Client(object):
def __init__(self, client_index, train_data_local_dict, train_data_local_num_dict, test_data_local_dict, device, model):
self.id = client_index
self.train_data = train_data_local_dict[self.id]
self.local_sample_number = train_data_local_num_dict[self.id]
self.test_local = test_data_local_dict[self.id]
self.device = device
self.model = model
model = Resnet56(class_num=dataset[-1], neck='bnneck')
model.load_state_dict(torch.load(str.format('model/cs_{0}_{1}_client_{2}_triplet_epochs_{3}.pt', client_num_in_total, partition_method, 0, 300)))
# [train_data_num, test_data_num, train_data_global, test_data_global, \
# # train_data_local_num_dict, train_data_local_dict, test_data_local_dict, \
# # class_num]
# model.load_state_dict(torch.load(str.format('model/client_{0}_triplet_epochs_{1}.pt', 0, 399)))
client_1 = Client(0, dataset[5], dataset[4], dataset[6], device, model)
# client_1.train_data.shuffle = False
def extract_features(model, data_loader,device):
model.to(device)
model.eval()
features = []
labels = []
with torch.no_grad():
for batch_idx, (x, l) in enumerate(data_loader):
x, l = x.to(device), l.to(device)
score, feats = model(x)
# print(feats.shape)
for feat, label in zip(feats, l):
features.append(feat.cpu())
labels.append(label.cpu())
return features, labels
features, labels = extract_features(client_1.model, client_1.test_local, device)
def extract_cnn_feature(model, input, modules):
# model.eval()
outputs = OrderedDict()
handles = []
# outputs = None
for m in modules:
# print(id(m))
outputs[id(m)] = None
def func(m, i, o): outputs[id(m)] = o.data.cpu()
handles.append(module.register_forward_hook(func))
model(input)
for h in handles:
h.remove()
# print(outputs.values())
#return: [1, 64, 256, 8, 8]
return list(outputs.values())[0]
def extract_features(model, data_loader,device, module):
model.to(device)
model.eval()
features = []
labels = []
with torch.no_grad():
for batch_idx, (x, l) in enumerate(data_loader):
x, l = x.to(device), l.to(device)
#[batch_size, channel, w, h]
outputs = extract_cnn_feature(model, x, module)
# features.append(outputs)
# labels.append(labels)
# print(len(outputs))
for output, label in zip(outputs, l):
# features[batch_idx] = output
# labels[batch_idx] = label
# print(output.shape)
# output: [64, 256, 8, 8]
features.append(output.view(-1, 256*8*8))
labels.append(label)
# print(len(outputs))
# print(outputs[0].shape)
#[79, 64, 256, 8, 8]
return features, labels
module = None
for name, m in client_1.model.named_modules():
if name == 'layer3.5.relu':
module = m
print(module)
# features, labels = extract_features(client_1.model, test_data_global, device, [module])
features, labels = extract_features(client_1.model, client_1.train_data, device, [module])
from collections import Counter
ls = np.array([int(l) for l in labels])
counter_result = Counter(ls)
print(Counter(ls))
# class imbalance sample
# ks = counter_result.keys()
def get_index(lst, item):
return [i for i in range(len(lst)) if lst[i]==item]
sampled_idx = []
# ids = np.random.randint(0, 9, size=5)
# print(idx)
for k, v in counter_result.items():
# print(f'{k}-{v}')
idx = get_index(ls, k)
# print(len(idx))
# print(k)
sampled_idx.extend(random.sample(idx, 100 if v>=100 else 0))
# if k in ids:
# sampled_idx.extend(random.sample(idx, 100 if v>=100 else 0))
# print(len(sampled_idx))
# print(shape(features))
# print(np.array(features[0][0]))
# print(features[0])
# print(np.array(features[0][0]).shape)
# print(torch.stack(labels).shape)
plot_features = torch.squeeze(torch.stack(features),dim=1)
plot_features = plot_features.numpy()[sampled_idx]
# for idx in sampled_idx:
# plt_f.append(plot_features[idx])
print(len(plot_features))
source_features = F.normalize(torch.from_numpy(plot_features), dim=1)
tsne = TSNE(n_components=2, init='pca', perplexity=30)
Y = tsne.fit_transform(source_features)
plt.scatter(Y[:, 0], Y[:, 1], c=ls[sampled_idx])
plt.show()
model = Resnet56(class_num=dataset[-1], neck='bnneck')
model.load_state_dict(torch.load(str.format('model/cs_{0}_{1}_client_{2}_oral_epochs_{3}.pt', client_num_in_total, partition_method, 0, 200)))
# [train_data_num, test_data_num, train_data_global, test_data_global, \
# # train_data_local_num_dict, train_data_local_dict, test_data_local_dict, \
# # class_num]
# model.load_state_dict(torch.load(str.format('model/client_{0}_triplet_epochs_{1}.pt', 0, 399)))
client_2 = Client(0, dataset[5], dataset[4], dataset[6], device, model)
features2, labels2 = extract_features(client_2.model, client_2.test_local, device)
ls = np.array([int(l) for l in labels2])
counter_result = Counter(ls)
print(Counter(ls))
# # class imbalance sample
# # ks = counter_result.keys()
# def get_index(lst, item):
# return [i for i in range(len(lst)) if lst[i]==item]
# sampled_idx = []
# # ids = np.random.randint(0, 9, size=5)
# # print(idx)
# for k, v in counter_result.items():
# # print(f'{k}-{v}')
# idx = get_index(ls, k)
# # print(len(idx))
# # print(k)
# sampled_idx.extend(random.sample(idx, 100 if v>=100 else 0))
# # if k in ids:
# # sampled_idx.extend(random.sample(idx, 100 if v>=100 else 0))
# # print(len(sampled_idx))
# print(shape(features))
# print(np.array(features[0][0]))
# print(features[0])
# print(np.array(features[0][0]).shape)
# print(torch.stack(labels).shape)
plot_features = torch.squeeze(torch.stack(features2),dim=1)
plot_features = plot_features.numpy()[sampled_idx]
# for idx in sampled_idx:
# plt_f.append(plot_features[idx])
print(len(plot_features))
source_features = F.normalize(torch.from_numpy(plot_features), dim=1)
tsne = TSNE(n_components=2, init='pca', perplexity=30)
Y = tsne.fit_transform(source_features)
plt.scatter(Y[:, 0], Y[:, 1], c=ls[sampled_idx])
plt.show()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/yahyanh21/Machine-Learning-Homework/blob/main/Week8.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
#https://www.kaggle.com/azzion/svm-for-beginners-tutorial/notebook
#https://www.kaggle.com/gulsahdemiryurek/mobile-price-classification-with-svm
#Mobile Price Classification
#The below topics are covered in this Kernal.
#1. Data prepocessing
#2. Target value Analysis
#3. SVM
#4. Linear SVM
#5. SV Regressor
#6. Non Linear SVM with kernal - RBF ( note: you can also try poly )
#7. Non Linear SVR
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import seaborn as sns
import matplotlib.pyplot as plt
from matplotlib.colors import ListedColormap
#connect to google drive
from google.colab import drive
drive.mount('/content/drive')
# A.DATA PREPROCESSING
# save filepath to variable for easier access
train_file_path = '../content/drive/My Drive/dataset/train1.csv'
test_file_path = '../content/drive/My Drive/dataset/test1.csv'
df = pd.read_csv(train_file_path)
test = pd.read_csv(test_file_path)
df.head()
df.info()
#battery_power: Total energy a battery can store in one time measured in mAh
#blue: Has bluetooth or not
#clock_speed: speed at which microprocessor executes instructions
#dual_sim: Has dual sim support or not
#fc: Front Camera mega pixels
#four_g: Has 4G or not
#int_memory: Internal Memory in Gigabytes
#m_dep: Mobile Depth in cm
#mobile_wt: Weight of mobile phone
#n_cores: Number of cores of processor
#pc: Primary Camera mega pixels
#px_height: Pixel Resolution Height
#px_width: Pixel Resolution Width
#ram: Random Access Memory in Mega Bytes
#sc_h: Screen Height of mobile in cm
#sc_w: Screen Width of mobile in cm
#talk_time: longest time that a single battery charge will last when you are
#three_g: Has 3G or not
#touch_screen: Has touch screen or not
#wifi: Has wifi or not
#price_range: This is the target variable with value of 0(low cost), 1(medium cost), 2(high cost) and 3(very high cost).
#cek missing values
import missingno as msno
import matplotlib.pyplot as plt
msno.bar(df)
plt.show()
#B. TARGET VALUE ANALYSIS
#understanding the predicted value - which is hot encoded, in real life price won't be hot encoded.
df['price_range'].describe(), df['price_range'].unique()
# there are 4 classes in the predicted value
#correlation matrix with heatmap (mencari korelasi antar features, salah satu teknik features selection)
corrmat = df.corr()
f,ax = plt.subplots(figsize=(12,10))
sns.heatmap(corrmat,vmax=0.8,square=True,annot=True,annot_kws={'size':8})
#price range correlation
corrmat.sort_values(by=["price_range"],ascending=False).iloc[0].sort_values(ascending=False)
f, ax = plt.subplots(figsize=(10,4))
plt.scatter(y=df['price_range'],x=df['battery_power'],color='red')
plt.scatter(y=df['price_range'],x=df['ram'],color='Green')
plt.scatter(y=df['price_range'],x=df['n_cores'],color='blue')
plt.scatter(y=df['price_range'],x=df['mobile_wt'],color='orange')
# clearly we can see that each of the category has different set of value ranges
# SUPPORT VECTOR MACHINES AND METHODS :
from sklearn.svm import SVC
from sklearn.model_selection import train_test_split
y_t = np.array(df['price_range'])
X_t = df
X_t = df.drop(['price_range'],axis=1)
X_t = np.array(X_t)
print("shape of Y :"+str(y_t.shape))
print("shape of X :"+str(X_t.shape))
from sklearn.preprocessing import MinMaxScaler #(scaling rentang 0-1)
scaler = MinMaxScaler()
X_t = scaler.fit_transform(X_t)
X_train,X_test,Y_train,Y_test = train_test_split(X_t,y_t,test_size=.20,random_state=42)
print("shape of X Train :"+str(X_train.shape))
print("shape of X Test :"+str(X_test.shape))
print("shape of Y Train :"+str(Y_train.shape))
print("shape of Y Test :"+str(Y_test.shape))
for this_C in [1,3,5,10,40,60,80,100]: #parameter C di SVM Linier
clf = SVC(kernel='linear',C=this_C).fit(X_train,Y_train) #clf = cross validate metrics for evaluating classification
scoretrain = clf.score(X_train,Y_train)
scoretest = clf.score(X_test,Y_test)
print("Linear SVM value of C:{}, training score :{:2f} , Test Score: {:2f} \n".format(this_C,scoretrain,scoretest))
from sklearn.model_selection import cross_val_score,StratifiedKFold,LeaveOneOut
clf1 = SVC(kernel='linear',C=20).fit(X_train,Y_train)
scores = cross_val_score(clf1,X_train,Y_train,cv=5)
strat_scores = cross_val_score(clf1,X_train,Y_train,cv=StratifiedKFold(5,random_state=10,shuffle=True))
#Loo = LeaveOneOut()
#Loo_scores = cross_val_score(clf1,X_train,Y_train,cv=Loo)
print("The Cross Validation Score :"+str(scores))
print("The Average Cross Validation Score :"+str(scores.mean()))
print("The Stratified Cross Validation Score :"+str(strat_scores))
print("The Average Stratified Cross Validation Score :"+str(strat_scores.mean()))
#print("The LeaveOneOut Cross Validation Score :"+str(Loo_scores))
#print("The Average LeaveOneOut Cross Validation Score :"+str(Loo_scores.mean()))
from sklearn.dummy import DummyClassifier
for strat in ['stratified', 'most_frequent', 'prior', 'uniform']:
dummy_maj = DummyClassifier(strategy=strat).fit(X_train,Y_train)
print("Train Stratergy :{} \n Score :{:.2f}".format(strat,dummy_maj.score(X_train,Y_train)))
print("Test Stratergy :{} \n Score :{:.2f}".format(strat,dummy_maj.score(X_test,Y_test)))
# plotting the decision boundries for the data
#converting the data to array for plotting.
X = np.array(df.iloc[:,[0,13]])
y = np.array(df['price_range'])
print("Shape of X:"+str(X.shape))
print("Shape of y:"+str(y.shape))
X = scaler.fit_transform(X)
# custome color maps
cm_dark = ListedColormap(['#ff6060', '#8282ff','#ffaa00','#fff244','#4df9b9','#76e8fc','#3ad628'])
cm_bright = ListedColormap(['#ffafaf', '#c6c6ff','#ffaa00','#ffe2a8','#bfffe7','#c9f7ff','#9eff93'])
plt.scatter(X[:,0],X[:,1],c=y,cmap=cm_dark,s=10,label=y)
plt.show()
h = .02 # step size in the mesh
C_param = 1 # No of neighbours
for weights in ['uniform', 'distance']:
# we create an instance of Neighbours Classifier and fit the data.
clf1 = SVC(kernel='linear',C=C_param)
clf1.fit(X, y)
# Plot the decision boundary. For that, we will assign a color to each
# point in the mesh [x_min, x_max]x[y_min, y_max].
x_min, x_max = X[:, 0].min()-.20, X[:, 0].max()+.20
y_min, y_max = X[:, 1].min()-.20, X[:, 1].max()+.20
xx, yy = np.meshgrid(np.arange(x_min, x_max, h),
np.arange(y_min, y_max, h))
Z = clf1.predict(np.c_[xx.ravel(), yy.ravel()]) # ravel to flatten the into 1D and c_ to concatenate
# Put the result into a color plot
Z = Z.reshape(xx.shape)
plt.figure()
plt.pcolormesh(xx, yy, Z, cmap=cm_bright)
# Plot also the training points
plt.scatter(X[:, 0], X[:, 1], c=y, cmap=cm_dark,
edgecolor='k', s=20)
plt.xlim(xx.min(), xx.max())
plt.ylim(yy.min(), yy.max())
plt.title("SVM Linear Classification (kernal = linear, Gamma = '%s')"% (C_param))
plt.show()
print("The score of the above :"+str(clf1.score(X,y)))
# Linear Support vector machine with only C Parameter
from sklearn.svm import LinearSVC
for this_C in [1,3,5,10,40,60,80,100]:
clf2 = LinearSVC(C=this_C).fit(X_train,Y_train)
scoretrain = clf2.score(X_train,Y_train)
scoretest = clf2.score(X_test,Y_test)
print("Linear SVM value of C:{}, training score :{:2f} , Test Score: {:2f} \n".format(this_C,scoretrain,scoretest))
from sklearn.svm import SVR
svr = SVR(kernel='linear',C=1,epsilon=.01).fit(X_train,Y_train)
print("{:.2f} is the accuracy of the SV Regressor".format(svr.score(X_train,Y_train)))
#NON LINIER SVM
# SMV with RBF KERNAL AND ONLY C PARAMETER
for this_C in [1,5,10,25,50,100]:
clf3 = SVC(kernel='rbf',C=this_C).fit(X_train,Y_train)
clf3train = clf3.score(X_train,Y_train)
clf3test = clf3.score(X_test,Y_test)
print("SVM for Non Linear \n C:{} Training Score : {:2f} Test Score : {:2f}\n".format(this_C,clf3train,clf3test))
# SVM WITH RBF KERNAL, C AND GAMMA HYPERPARAMTER
for this_gamma in [.1,.5,.10,.25,.50,1]:
for this_C in [1,5,7,10,15,25,50]:
clf3 = SVC(kernel='rbf',C=this_C,gamma=this_gamma).fit(X_train,Y_train)
clf3train = clf3.score(X_train,Y_train)
clf3test = clf3.score(X_test,Y_test)
print("SVM for Non Linear \n Gamma: {} C:{} Training Score : {:2f} Test Score : {:2f}\n".format(this_gamma,this_C,clf3train,clf3test))
# grid search method
from sklearn.model_selection import GridSearchCV
param_grid = {'C': [1,5,7,10,15,25,50],
'gamma': [.1,.5,.10,.25,.50,1]}
GS = GridSearchCV(SVC(kernel='rbf'),param_grid,cv=5)
GS.fit(X_train,Y_train)
print("the parameters {} are the best.".format(GS.best_params_))
print("the best score is {:.2f}.".format(GS.best_score_))
# Kernalized SVM machine
svr2 = SVR(degree=2,C=100,epsilon=.01).fit(X_train,Y_train)
print("{:.2f} is the accuracy of the SV Regressor".format(svr2.score(X_train,Y_train)))
test = test.drop(['id'],axis=1)
test.head()
test_mat = np.array(test)
test = scaler.fit_transform(test_mat)
clf4 = SVC(kernel='rbf',C=25,gamma=.1).fit(X_train,Y_train)
prediction = clf4.predict(test_mat)
pred = pd.DataFrame(prediction)
pred.head()
pred.info()
prediction = svr2.predict(test_mat)
pred = pd.DataFrame(prediction)
pred.head()
```
| github_jupyter |
```
import os
import glob
import json
import pandas as pd
def load_gpu_util(dlprof_summary_file):
with open(dlprof_summary_file) as json_file:
summary = json.load(json_file)
gpu_util_raw = summary["Summary Report"]
gpu_util = {
"sm_util": float(100 - gpu_util_raw["GPU Idle %"][0]),
"tc_util": float(gpu_util_raw["Tensor Core Kernel Utilization %"][0])
}
return gpu_util
def parse_pl_timings(pl_profile_file):
lines = [line.rstrip("\n") for line in open(pl_profile_file)]
mean_timings = {}
for l in lines[7:]:
if "|" in l:
l = l.split("|")
l = [i.strip() for i in l]
mean_timings[l[0]] = float(l[1])
return mean_timings
gpu_names = [
"v100-16gb-300w",
"a100-40gb-400w"
]
compute_types = [
"amp"
]
model_names = [
"distilroberta-base",
"roberta-base",
"roberta-large"
]
columns = ["gpu", "compute", "model", "seq_len", "batch_size",
"cpu_time", "forward", "backward", "train_loss",
"vram_usage", "vram_io",
"sm_util", "tc_util", ]
rows = []
cpu_time_sections = ["get_train_batch", "on_batch_start", "on_train_batch_start",
"training_step_end", "on_after_backward",
"on_batch_end", "on_train_batch_end"]
for gn in gpu_names:
for ct in compute_types:
for mn in model_names:
path = "/".join(["./results", gn, ct, mn])+"/*"
configs = glob.glob(path)
configs.sort(reverse=True)
for c in configs:
print(c)
try:
seq_len, batch_size = c.split("/")[-1].split("-")
row_1 = [gn, ct, mn, int(seq_len), int(batch_size)]
pl_timings = parse_pl_timings(c+"/pl_profile.txt")
cpu_time = sum([pl_timings[k] for k in cpu_time_sections])
metrics_0 = pd.read_csv(c+"/version_0/metrics.csv")
metrics_1 = pd.read_csv(c+"/version_1/metrics.csv")
sm_util = metrics_0["gpu_id: 0/utilization.gpu (%)"].mean()
vram_usage = metrics_0["gpu_id: 0/memory.used (MB)"].mean()
vram_io = metrics_0["gpu_id: 0/utilization.memory (%)"].mean()
test_loss = (metrics_0["train_loss"].mean() + metrics_1["train_loss"].mean())/2
row_2 = [cpu_time, pl_timings["model_forward"], pl_timings["model_backward"], test_loss]
util_data = load_gpu_util(c+"/dlprof_summary.json")
sm_util = (sm_util + util_data["sm_util"])/2
row_3 = [vram_usage, vram_io, sm_util, util_data["tc_util"]]
row = row_1 + row_2 + row_3
print(row)
rows.append(row)
except Exception as e:
print(e)
df = pd.DataFrame(rows, columns=columns)
df.head(20)
df.to_csv("./results.csv")
```
| github_jupyter |
## Define the Convolutional Neural Network
After you've looked at the data you're working with and, in this case, know the shapes of the images and of the keypoints, you are ready to define a convolutional neural network that can *learn* from this data.
In this notebook and in `models.py`, you will:
1. Define a CNN with images as input and keypoints as output
2. Construct the transformed FaceKeypointsDataset, just as before
3. Train the CNN on the training data, tracking loss
4. See how the trained model performs on test data
5. If necessary, modify the CNN structure and model hyperparameters, so that it performs *well* **\***
**\*** What does *well* mean?
"Well" means that the model's loss decreases during training **and**, when applied to test image data, the model produces keypoints that closely match the true keypoints of each face. And you'll see examples of this later in the notebook.
---
## CNN Architecture
Recall that CNN's are defined by a few types of layers:
* Convolutional layers
* Maxpooling layers
* Fully-connected layers
You are required to use the above layers and encouraged to add multiple convolutional layers and things like dropout layers that may prevent overfitting. You are also encouraged to look at literature on keypoint detection, such as [this paper](https://arxiv.org/pdf/1710.00977.pdf), to help you determine the structure of your network.
### TODO: Define your model in the provided file `models.py` file
This file is mostly empty but contains the expected name and some TODO's for creating your model.
---
## PyTorch Neural Nets
To define a neural network in PyTorch, you define the layers of a model in the function `__init__` and define the feedforward behavior of a network that employs those initialized layers in the function `forward`, which takes in an input image tensor, `x`. The structure of this Net class is shown below and left for you to fill in.
Note: During training, PyTorch will be able to perform backpropagation by keeping track of the network's feedforward behavior and using autograd to calculate the update to the weights in the network.
#### Define the Layers in ` __init__`
As a reminder, a conv/pool layer may be defined like this (in `__init__`):
```
# 1 input image channel (for grayscale images), 32 output channels/feature maps, 3x3 square convolution kernel
self.conv1 = nn.Conv2d(1, 32, 3)
# maxpool that uses a square window of kernel_size=2, stride=2
self.pool = nn.MaxPool2d(2, 2)
```
#### Refer to Layers in `forward`
Then referred to in the `forward` function like this, in which the conv1 layer has a ReLu activation applied to it before maxpooling is applied:
```
x = self.pool(F.relu(self.conv1(x)))
```
Best practice is to place any layers whose weights will change during the training process in `__init__` and refer to them in the `forward` function; any layers or functions that always behave in the same way, such as a pre-defined activation function, should appear *only* in the `forward` function.
#### Why models.py
You are tasked with defining the network in the `models.py` file so that any models you define can be saved and loaded by name in different notebooks in this project directory. For example, by defining a CNN class called `Net` in `models.py`, you can then create that same architecture in this and other notebooks by simply importing the class and instantiating a model:
```
from models import Net
net = Net()
```
```
# import the usual resources
import matplotlib.pyplot as plt
import numpy as np
# watch for any changes in model.py, if it changes, re-load it automatically
%load_ext autoreload
%autoreload 2
## TODO: Define the Net in models.py
import torch
import torch.nn as nn
import torch.nn.functional as F
## TODO: Once you've define the network, you can instantiate it
# one example conv layer has been provided for you
from models import Net
net = Net()
print(net)
```
## Transform the dataset
To prepare for training, create a transformed dataset of images and keypoints.
### TODO: Define a data transform
In PyTorch, a convolutional neural network expects a torch image of a consistent size as input. For efficient training, and so your model's loss does not blow up during training, it is also suggested that you normalize the input images and keypoints. The necessary transforms have been defined in `data_load.py` and you **do not** need to modify these; take a look at this file (you'll see the same transforms that were defined and applied in Notebook 1).
To define the data transform below, use a [composition](http://pytorch.org/tutorials/beginner/data_loading_tutorial.html#compose-transforms) of:
1. Rescaling and/or cropping the data, such that you are left with a square image (the suggested size is 224x224px)
2. Normalizing the images and keypoints; turning each RGB image into a grayscale image with a color range of [0, 1] and transforming the given keypoints into a range of [-1, 1]
3. Turning these images and keypoints into Tensors
These transformations have been defined in `data_load.py`, but it's up to you to call them and create a `data_transform` below. **This transform will be applied to the training data and, later, the test data**. It will change how you go about displaying these images and keypoints, but these steps are essential for efficient training.
As a note, should you want to perform data augmentation (which is optional in this project), and randomly rotate or shift these images, a square image size will be useful; rotating a 224x224 image by 90 degrees will result in the same shape of output.
```
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
# the dataset we created in Notebook 1 is copied in the helper file `data_load.py`
from data_load import FacialKeypointsDataset
# the transforms we defined in Notebook 1 are in the helper file `data_load.py`
from data_load import Rescale, RandomCrop, Normalize, ToTensor
## TODO: define the data_transform using transforms.Compose([all tx's, . , .])
# order matters! i.e. rescaling should come before a smaller crop
data_transform = transforms.Compose([
Rescale(256), RandomCrop(224), Normalize(), ToTensor()
])
# testing that you've defined a transform
assert(data_transform is not None), 'Define a data_transform'
# create the transformed dataset
transformed_dataset = FacialKeypointsDataset(csv_file='data/training_frames_keypoints.csv',
root_dir='data/training/',
transform=data_transform)
print('Number of images: ', len(transformed_dataset))
# iterate through the transformed dataset and print some stats about the first few samples
for i in range(4):
sample = transformed_dataset[i]
print(i, sample['image'].size(), sample['keypoints'].size())
```
## Batching and loading data
Next, having defined the transformed dataset, we can use PyTorch's DataLoader class to load the training data in batches of whatever size as well as to shuffle the data for training the model. You can read more about the parameters of the DataLoader, in [this documentation](http://pytorch.org/docs/master/data.html).
#### Batch size
Decide on a good batch size for training your model. Try both small and large batch sizes and note how the loss decreases as the model trains.
**Note for Windows users**: Please change the `num_workers` to 0 or you may face some issues with your DataLoader failing.
```
# load training data in batches
batch_size = 10
train_loader = DataLoader(transformed_dataset,
batch_size=batch_size,
shuffle=True,
num_workers=4)
```
## Before training
Take a look at how this model performs before it trains. You should see that the keypoints it predicts start off in one spot and don't match the keypoints on a face at all! It's interesting to visualize this behavior so that you can compare it to the model after training and see how the model has improved.
#### Load in the test dataset
The test dataset is one that this model has *not* seen before, meaning it has not trained with these images. We'll load in this test data and before and after training, see how your model performs on this set!
To visualize this test data, we have to go through some un-transformation steps to turn our images into python images from tensors and to turn our keypoints back into a recognizable range.
```
# load in the test data, using the dataset class
# AND apply the data_transform you defined above
# create the test dataset
test_dataset = FacialKeypointsDataset(csv_file='data/test_frames_keypoints.csv',
root_dir='data/test/',
transform=data_transform)
# load test data in batches
batch_size = 16
test_loader = DataLoader(test_dataset,
batch_size=batch_size,
shuffle=True,
num_workers=4)
```
## Apply the model on a test sample
To test the model on a test sample of data, you have to follow these steps:
1. Extract the image and ground truth keypoints from a sample
2. Make sure the image is a FloatTensor, which the model expects.
3. Forward pass the image through the net to get the predicted, output keypoints.
This function test how the network performs on the first batch of test data. It returns the images, the transformed images, the predicted keypoints (produced by the model), and the ground truth keypoints.
```
# test the model on a batch of test images
def net_sample_output():
# iterate through the test dataset
for i, sample in enumerate(test_loader):
# get sample data: images and ground truth keypoints
images = sample['image']
key_pts = sample['keypoints']
# convert images to FloatTensors
images = images.type(torch.FloatTensor)
# forward pass to get net output
output_pts = net(images)
# reshape to batch_size x 68 x 2 pts
output_pts = output_pts.view(output_pts.size()[0], 68, -1)
# break after first image is tested
if i == 0:
return images, output_pts, key_pts
```
#### Debugging tips
If you get a size or dimension error here, make sure that your network outputs the expected number of keypoints! Or if you get a Tensor type error, look into changing the above code that casts the data into float types: `images = images.type(torch.FloatTensor)`.
```
# call the above function
# returns: test images, test predicted keypoints, test ground truth keypoints
test_images, test_outputs, gt_pts = net_sample_output()
# print out the dimensions of the data to see if they make sense
print(test_images.data.size())
print(test_outputs.data.size())
print(gt_pts.size())
```
## Visualize the predicted keypoints
Once we've had the model produce some predicted output keypoints, we can visualize these points in a way that's similar to how we've displayed this data before, only this time, we have to "un-transform" the image/keypoint data to display it.
Note that I've defined a *new* function, `show_all_keypoints` that displays a grayscale image, its predicted keypoints and its ground truth keypoints (if provided).
```
def show_all_keypoints(image, predicted_key_pts, gt_pts=None):
"""Show image with predicted keypoints"""
# image is grayscale
plt.imshow(image, cmap='gray')
plt.scatter(predicted_key_pts[:, 0], predicted_key_pts[:, 1], s=20, marker='.', c='m')
# plot ground truth points as green pts
if gt_pts is not None:
plt.scatter(gt_pts[:, 0], gt_pts[:, 1], s=20, marker='.', c='g')
```
#### Un-transformation
Next, you'll see a helper function. `visualize_output` that takes in a batch of images, predicted keypoints, and ground truth keypoints and displays a set of those images and their true/predicted keypoints.
This function's main role is to take batches of image and keypoint data (the input and output of your CNN), and transform them into numpy images and un-normalized keypoints (x, y) for normal display. The un-transformation process turns keypoints and images into numpy arrays from Tensors *and* it undoes the keypoint normalization done in the Normalize() transform; it's assumed that you applied these transformations when you loaded your test data.
```
# visualize the output
# by default this shows a batch of 10 images
def visualize_output(test_images, test_outputs, gt_pts=None, batch_size=10):
for i in range(batch_size):
plt.figure(figsize=(20,10))
ax = plt.subplot(1, batch_size, i+1)
# un-transform the image data
image = test_images[i].data # get the image from it's wrapper
image = image.numpy() # convert to numpy array from a Tensor
image = np.transpose(image, (1, 2, 0)) # transpose to go from torch to numpy image
# un-transform the predicted key_pts data
predicted_key_pts = test_outputs[i].data
predicted_key_pts = predicted_key_pts.numpy()
# undo normalization of keypoints
predicted_key_pts = predicted_key_pts*50.0+100
# plot ground truth points for comparison, if they exist
ground_truth_pts = None
if gt_pts is not None:
ground_truth_pts = gt_pts[i]
ground_truth_pts = ground_truth_pts*50.0+100
# call show_all_keypoints
show_all_keypoints(np.squeeze(image), predicted_key_pts, ground_truth_pts)
plt.axis('off')
plt.show()
# call it
visualize_output(test_images, test_outputs, gt_pts)
```
## Training
#### Loss function
Training a network to predict keypoints is different than training a network to predict a class; instead of outputting a distribution of classes and using cross entropy loss, you may want to choose a loss function that is suited for regression, which directly compares a predicted value and target value. Read about the various kinds of loss functions (like MSE or L1/SmoothL1 loss) in [this documentation](http://pytorch.org/docs/master/_modules/torch/nn/modules/loss.html).
### TODO: Define the loss and optimization
Next, you'll define how the model will train by deciding on the loss function and optimizer.
---
```
## TODO: Define the loss and optimization
import torch.optim as optim
criterion = F.smooth_l1_loss
optimizer = optim.AdamW(net.parameters(), lr=0.0001)
# to plot the training loss
training_loss = []
real_epoch = 0
```
## Training and Initial Observation
Now, you'll train on your batched training data from `train_loader` for a number of epochs.
To quickly observe how your model is training and decide on whether or not you should modify it's structure or hyperparameters, you're encouraged to start off with just one or two epochs at first. As you train, note how your the model's loss behaves over time: does it decrease quickly at first and then slow down? Does it take a while to decrease in the first place? What happens if you change the batch size of your training data or modify your loss function? etc.
Use these initial observations to make changes to your model and decide on the best architecture before you train for many epochs and create a final model.
```
def train_net(n_epochs):
# prepare the net for training
net.train()
# train with gpu
device = torch.device('cpu')
if torch.cuda.is_available():
device = torch.device('cuda:0')
net.to(device)
for epoch in range(n_epochs): # loop over the dataset multiple times
running_loss = 0.0
# train on batches of data, assumes you already have train_loader
for batch_i, data in enumerate(train_loader):
# get the input images and their corresponding labels
images = data['image']
key_pts = data['keypoints']
# flatten pts
key_pts = key_pts.view(key_pts.size(0), -1)
# convert variables to floats for regression loss
key_pts = key_pts.type(torch.FloatTensor)
images = images.type(torch.FloatTensor)
# forward pass to get outputs
output_pts = net(images.to(device))
# calculate the loss between predicted and target keypoints
loss = criterion(output_pts, key_pts.to(device))
# zero the parameter (weight) gradients
optimizer.zero_grad()
# backward pass to calculate the weight gradients
loss.backward()
# update the weights
optimizer.step()
# print loss statistics
# to convert loss into a scalar and add it to the running_loss, use .item()
running_loss += loss.item()
if batch_i % batch_size == (batch_size-1): # print every 10 batches
print('Epoch: {}, Batch: {}, Avg. Loss: {}'.format(epoch + 1, batch_i+1, running_loss/(batch_size*100)))
training_loss.append(running_loss/1000)
running_loss = 0.0
print('Finished Training')
# train your network
n_epochs = 6 # start small, and increase when you've decided on your model structure and hyperparams
train_net(n_epochs)
plt.plot(training_loss, color="m")
```
## Test data
See how your model performs on previously unseen, test data. We've already loaded and transformed this data, similar to the training data. Next, run your trained model on these images to see what kind of keypoints are produced. You should be able to see if your model is fitting each new face it sees, if the points are distributed randomly, or if the points have actually overfitted the training data and do not generalize.
```
# get a sample of test data again
net.to(torch.device('cpu'))
test_images, test_outputs, gt_pts = net_sample_output()
print(test_images.data.size())
print(test_outputs.data.size())
print(gt_pts.size())
## TODO: visualize your test output
# you can use the same function as before, by un-commenting the line below:
visualize_output(test_images, test_outputs, gt_pts)
```
Once you've found a good model (or two), save your model so you can load it and use it later!
```
## TODO: change the name to something uniqe for each new model
model_dir = 'saved_models/'
model_name = 'keypoints_model_1.pt'
# after training, save your model parameters in the dir 'saved_models'
torch.save(net.state_dict(), model_dir+model_name)
```
After you've trained a well-performing model, answer the following questions so that we have some insight into your training and architecture selection process. Answering all questions is required to pass this project.
### Question 1: What optimization and loss functions did you choose and why?
**Answer**:
Started with SGD and the MSE Loss function. Loss decreased slowly and taken around 32 iterations to bring it to 0.002***. Changed to AdamW , which after couple of epochs jumped to a minimal value. Smooth_L1_Loss had more accuracy compared to the MSE_loss , hence choosed Smooth_L1_Loss.
### Question 2: What kind of network architecture did you start with and how did it change as you tried different architectures? Did you decide to add more convolutional layers or any layers to avoid overfitting the data?
Started with a minimal convolution network with 02 convolution layers and 2 fully connected layers. Network performed well on the training data , but failed on the test data. Changed to network with 3 convolution layers and 3 FC layers. added Dropout between the FC layers. Tuned the dropout probability to improve the train-test loss.
### Question 3: How did you decide on the number of epochs and batch_size to train your model?
Started with minimal Epoch - 2 and default batch_size 10. After couple of iterations changed the batch_size to 16. Plotted the number of Epoch vs Loss to see whether the loss increases or explodes after certain epochs. but after 6 epochs, the loss was stagnant, so stopped with 6 epochs.
## Feature Visualization
Sometimes, neural networks are thought of as a black box, given some input, they learn to produce some output. CNN's are actually learning to recognize a variety of spatial patterns and you can visualize what each convolutional layer has been trained to recognize by looking at the weights that make up each convolutional kernel and applying those one at a time to a sample image. This technique is called feature visualization and it's useful for understanding the inner workings of a CNN.
In the cell below, you can see how to extract a single filter (by index) from your first convolutional layer. The filter should appear as a grayscale grid.
```
# Get the weights in the first conv layer, "conv1"
# if necessary, change this to reflect the name of your first conv layer
weights1 = net.conv1.weight.data
w = weights1.numpy()
filter_index = 0
print(w[filter_index][0])
print(w[filter_index][0].shape)
# display the filter weights
plt.imshow(w[filter_index][0], cmap='gray')
```
## Feature maps
Each CNN has at least one convolutional layer that is composed of stacked filters (also known as convolutional kernels). As a CNN trains, it learns what weights to include in it's convolutional kernels and when these kernels are applied to some input image, they produce a set of **feature maps**. So, feature maps are just sets of filtered images; they are the images produced by applying a convolutional kernel to an input image. These maps show us the features that the different layers of the neural network learn to extract. For example, you might imagine a convolutional kernel that detects the vertical edges of a face or another one that detects the corners of eyes. You can see what kind of features each of these kernels detects by applying them to an image. One such example is shown below; from the way it brings out the lines in an the image, you might characterize this as an edge detection filter.
<img src='images/feature_map_ex.png' width=50% height=50%/>
Next, choose a test image and filter it with one of the convolutional kernels in your trained CNN; look at the filtered output to get an idea what that particular kernel detects.
### TODO: Filter an image to see the effect of a convolutional kernel
---
```
##TODO: load in and display any image from the transformed test dataset
import cv2
image = cv2.imread('./data/sample_image.jpg')
image = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
weights1 = net.conv1.weight.data
filter_index = 0
w = weights1.numpy()
filtered_image = cv2.filter2D(image, -1, w[filter_index][0])
## TODO: Using cv's filter2D function,
## apply a specific set of filter weights (like the one displayed above) to the test image
plt.subplots(4,4, figsize=(10,5))
for filter in range(16):
plt.subplot(4,4,filter+1)
plt.imshow(cv2.filter2D(image, -1, w[filter][0]), cmap="gray")
```
### Question 4: Choose one filter from your trained CNN and apply it to a test image; what purpose do you think it plays? What kind of feature do you think it detects?
Most filters on the first layer seem to blur the image. Filter 4, 7 , 10 detected the edges
---
## Moving on!
Now that you've defined and trained your model (and saved the best model), you are ready to move on to the last notebook, which combines a face detector with your saved model to create a facial keypoint detection system that can predict the keypoints on *any* face in an image!
| github_jupyter |
## Advanced Lane Finding Project
The goals / steps of this project are the following:
* Compute the camera calibration matrix and distortion coefficients given a set of chessboard images.
* Apply a distortion correction to raw images.
* Use color transforms, gradients, etc., to create a thresholded binary image.
* Apply a perspective transform to rectify binary image ("birds-eye view").
* Detect lane pixels and fit to find the lane boundary.
* Determine the curvature of the lane and vehicle position with respect to center.
* Warp the detected lane boundaries back onto the original image.
* Output visual display of the lane boundaries and numerical estimation of lane curvature and vehicle position.
---
## First, I'll compute the camera calibration using chessboard images
```
import numpy as np
import cv2
import glob
import matplotlib.pyplot as plt
%matplotlib inline
# prepare object points, like (0,0,0), (1,0,0), (2,0,0) ....,(6,5,0)
objp = np.zeros((6*9,3), np.float32)
objp[:,:2] = np.mgrid[0:9,0:6].T.reshape(-1,2)
# Arrays to store object points and image points from all the images.
objpoints = [] # 3d points in real world space
imgpoints = [] # 2d points in image plane.
# Make a list of calibration images
images = glob.glob('../camera_cal/calibration*.jpg')
# Step through the list and search for chessboard corners
for fname in images:
img = cv2.imread(fname)
gray = cv2.cvtColor(img,cv2.COLOR_BGR2GRAY)
# Find the chessboard corners
ret, corners = cv2.findChessboardCorners(gray, (9,6),None)
# If found, add object points, image points
if ret == True:
objpoints.append(objp)
imgpoints.append(corners)
# Draw and display the corners
img = cv2.drawChessboardCorners(img, (9,6), corners, ret)
cv2.imshow('img',img)
cv2.waitKey(500)
cv2.destroyAllWindows()
ret, mtx, dist, tvecs, rvecs = cv2.calibrateCamera(objpoints, imgpoints, img.shape[::-1][1:3],None,None)
images = glob.glob('../test_images/*.jpg')
img = cv2.imread(images[3])
undist = cv2.undistort(img, mtx, dist, None, mtx)
```
## Use color transforms and gradients to create a thresholded binary image.
```
hls = cv2.cvtColor(img, cv2.COLOR_BGR2HLS)
l_channel = hls[:,:,1]
s_channel = hls[:,:,2]
sobelx = cv2.Sobel(l_channel, cv2.CV_64F, 1, 0)
abs_sobelx = np.absolute(sobelx)
scaled_sobel = np.uint8(255*abs_sobelx/np.max(abs_sobelx))
sxbinary = np.zeros_like(scaled_sobel)
sx_thresh=(20, 100)
sxbinary[(scaled_sobel >= sx_thresh[0]) & (scaled_sobel <= sx_thresh[1])] = 1
s_thresh=(170, 255)
s_binary = np.zeros_like(s_channel)
s_binary[(s_channel >= s_thresh[0]) & (s_channel <= s_thresh[1])] = 1
combined_binary = np.zeros_like(sxbinary)
combined_binary[(s_binary == 1) | (sxbinary == 1)] =1
plt.imshow(combined_binary*255, cmap = 'gray')
```
## Apply a perspective transform to rectify binary image ("birds-eye view").
```
def warp(img, src, dst):
M = cv2.getPerspectiveTransform(src, dst)
warped = cv2.warpPerspective(img, M, (img.shape[1],img.shape[0]))
return warped
src = np.float32([(300,650),(500,470), (710,470), (1160,650)])
dst = np.float32([(200,img.shape[0]),(200,0), (img.shape[1]-200,0), (img.shape[1]-200,img.shape[0])])
reverse_M = cv2.getPerspectiveTransform(dst, src)
warped = warp(combined_binary, src, dst)
plt.imshow(warped*255, cmap = 'gray')
```
## Detect lane pixels and fit to find the lane boundary.
```
def laneFinder(img):
histogram = np.sum(img[img.shape[0]//2:,:], axis=0)
out_img = np.dstack((img, img, img))
# Find the peak of the left and right halves of the histogram
# These will be the starting point for the left and right lines
midpoint = np.int(histogram.shape[0]//2)
leftx_base = np.argmax(histogram[:midpoint])
rightx_base = np.argmax(histogram[midpoint:]) + midpoint
#Then choose window parameters and build windows
nwindows = 9
margin = 100
minpix = 50
window_height = np.int(img.shape[0]//nwindows)
# Identify the x and y positions of all nonzero pixels in the image
nonzero = img.nonzero()
nonzeroy = np.array(nonzero[0])
nonzerox = np.array(nonzero[1])
# Current positions to be updated later for each window in nwindows
leftx_current = leftx_base
rightx_current = rightx_base
# Create empty lists to receive left and right lane pixel indices
left_lane_inds = []
right_lane_inds = []
# Step through the windows one by one
for window in range(nwindows):
# Identify window boundaries in x and y (and right and left)
win_y_low = img.shape[0] - (window+1)*window_height
win_y_high = img.shape[0] - window*window_height
win_xleft_low = leftx_current - margin
win_xleft_high = leftx_current + margin
win_xright_low = rightx_current - margin
win_xright_high = rightx_current + margin
# Draw the windows on the visualization image
cv2.rectangle(out_img,(win_xleft_low,win_y_low),
(win_xleft_high,win_y_high),(0,255,0), 2)
cv2.rectangle(out_img,(win_xright_low,win_y_low),
(win_xright_high,win_y_high),(0,255,0), 2)
# Identify the nonzero pixels in x and y within the window #
good_left_inds = ((nonzeroy >= win_y_low) & (nonzeroy < win_y_high) &
(nonzerox >= win_xleft_low) & (nonzerox < win_xleft_high)).nonzero()[0]
good_right_inds = ((nonzeroy >= win_y_low) & (nonzeroy < win_y_high) &
(nonzerox >= win_xright_low) & (nonzerox < win_xright_high)).nonzero()[0]
# Append these indices to the lists
left_lane_inds.append(good_left_inds)
right_lane_inds.append(good_right_inds)
# If you found > minpix pixels, recenter next window on their mean position
if len(good_left_inds) > minpix:
leftx_current = np.int(np.mean(nonzerox[good_left_inds]))
if len(good_right_inds) > minpix:
rightx_current = np.int(np.mean(nonzerox[good_right_inds]))
# Concatenate the arrays of indices (previously was a list of lists of pixels)
try:
left_lane_inds = np.concatenate(left_lane_inds)
right_lane_inds = np.concatenate(right_lane_inds)
except ValueError:
# Avoids an error if the above is not implemented fully
pass
# Extract left and right line pixel positions
leftx = nonzerox[left_lane_inds]
lefty = nonzeroy[left_lane_inds]
rightx = nonzerox[right_lane_inds]
righty = nonzeroy[right_lane_inds]
return leftx, lefty, rightx, righty, out_img
leftx, lefty, rightx, righty, out_img = laneFinder(warped)
```
## Determine the curvature of the lane and vehicle position with respect to center.
```
def fit_polynomial(img):
# Find our lane pixels first
leftx, lefty, rightx, righty, out_img = laneFinder(img)
# Fit a second order polynomial to each using `np.polyfit`
left_fit = np.polyfit(lefty, leftx, 2)
right_fit = np.polyfit(righty, rightx, 2)
# Generate x and y values for plotting
ploty = np.linspace(0, img.shape[0]-1, img.shape[0] )
try:
left_fitx = left_fit[0]*ploty**2 + left_fit[1]*ploty + left_fit[2]
right_fitx = right_fit[0]*ploty**2 + right_fit[1]*ploty + right_fit[2]
except TypeError:
# Avoids an error if `left` and `right_fit` are still none or incorrect
print('The function failed to fit a line!')
left_fitx = 1*ploty**2 + 1*ploty
right_fitx = 1*ploty**2 + 1*ploty
## Visualization ##
# Colors in the left and right lane regions
out_img[lefty, leftx] = [255, 0, 0]
out_img[righty, rightx] = [0, 0, 255]
# Plots the left and right polynomials on the lane lines
plt.plot(left_fitx, ploty, color='yellow')
plt.plot(right_fitx, ploty, color='yellow')
return out_img
out_img = fit_polynomial(warped)
plt.imshow(out_img)
```
## Transfer back to original image and plot detected lanes
```
def drawOnOriginal(org_img, warped, left_fit, right_fit, reverse_M):
base_new = np.zeros_like(org_img)
color_base = np.dstack((base_new, base_new, base_new))
```
| github_jupyter |
# COMP305 -> 2-median problem on Optimal Placement of 2 Hospitals
## Imports
```
import time
import heapq
import numpy as np
from collections import defaultdict
from collections import Counter
from random import choice
from random import randint
```
## Data Read
```
#with open("tests/test1_new.txt") as f:
# test2 = f.read().splitlines()
#with open("tests/test2_new.txt") as f:
# test2 = f.read().splitlines()
#with open("tests/test3_new.txt") as f:
#test3 = f.read().splitlines()
#with open("tests/test1_aycan.txt") as f:
# test01 = f.read().splitlines()
#with open("tests/test_aycan.txt") as f:
# test001 = f.read().splitlines()
```
# txt -> Graph
```
with open("tests/test2_new.txt") as f:
test2 = f.read().splitlines()
lines=test2
number_of_vertices = int(lines[0])
number_of_edges = int(lines[1])
vertices = lines[2:2+number_of_vertices]
edges = lines[2+number_of_vertices:]
ids_and_populations = [tuple(map(int, vertices[i].split(" "))) for i in range(len(vertices))]
populations = dict(sorted(dict(ids_and_populations).items())) #redundant sort
mydict = lambda: defaultdict(lambda: defaultdict())
G = mydict()
for i in range(len(edges)):
source, target, weight = map(int, edges[i].split(" "))
G[source][target] = weight
G[target][source] = weight
```
## Random spawned k-th neighbor Subgraph expansion
```
def dijkstra_path(G, population_dict, source):
costs = dict()
for key in G:
costs[key] = np.inf
costs[source] = 0
#display(source,costs)
pq = []
for node in G:
heapq.heappush(pq, (node, costs[node]))
while len(pq) != 0:
current_node, current_node_distance = heapq.heappop(pq)
for neighbor_node in G[current_node]:
#print(current_node,costs[source])
weight = G[current_node][neighbor_node]
distance = current_node_distance + weight
if distance < costs[neighbor_node]:
#if source==neighbor_node:
#print('here')
costs[neighbor_node] = distance
heapq.heappush(pq, (neighbor_node, distance))
sorted_costs_lst=list(dict(sorted(costs.items())).values())
sorted_populations_lst = list(dict(sorted(population_dict.items())).values())
#print(np.array(sorted_costs_lst) ,np.array(sorted_populations_lst))
return np.array(sorted_costs_lst) * np.array(sorted_populations_lst)
#return list(dict(sorted(costs.items())).values())
# V4 because runs in V^4
def V4(G):
APSP = np.zeros((number_of_vertices,number_of_vertices))
population_dict = dict(sorted([(k, populations[k]) for k in G.keys()]))
for vertex in vertices:
vertex= int(vertex.split()[0])
APSP[vertex] = [e for e in dijkstra_path(G, population_dict,vertex)]
global glob
res = {}
n = len(APSP)
temp_arr = APSP.copy()
count=0
count2=0
for first in range(n):
for second in range(first+1,n):
if first==second:
continue
count+=1
#print(count)
#print(first,second)
temp_arr = APSP.copy()
for row in temp_arr:
if row[first]<row[second]:
row[second]=0
else:
row[first]=0
#print(temp_arr,count)
to_be_summed = temp_arr[:,[first,second]]
summed = sum(sum(to_be_summed))
res[(first,second)]=summed
ret=min(res, key=res.get)
#display(len(res))
#display(res)
#print('pick {}th and {}th vertices to place hospitals!'.format(ret[0],ret[1]))
return ret, res[ret], res
```
# ML
```
class Vertex:
def __init__(self, id, weight):
self.key = 0
self.id = id
self.visited = False
self.weight = weight
self.neighbour_count = 0
self.neighbour_weight = 0
self.distance = 0
self.sum = 0
self.neighbour_neighbour_count = 0
self.neighbour_neighbour_weight = 0
self.neighbour_distance = 0
self.neighbour_sum = 0
self.neighbour_list = []
def add_neighbour(self, Vertex, distance):
self.neighbour_list += [Vertex]
self.neighbour_count = self.neighbour_count + 1
self.distance = self.distance + distance
self.neighbour_weight = self.neighbour_weight + Vertex.weight
self.sum = self.sum + distance * Vertex.weight
self.visited = True
def add_neighbour_neighbour(self, Vertex):
self.neighbour_neighbour_count += Vertex.neighbour_count
self.neighbour_neighbour_weight += Vertex.neighbour_weight
self.neighbour_distance += Vertex.distance
self.neighbour_sum += Vertex.sum
self.visited = True
def calculate(self, a1, a2, a3):
self.key = self.weight * a1 + self.sum * a2 + self.neighbour_sum * a3
return self.key
def get_key(self):
return self.key
def __str__(self):
return "Key: " + self.key.__str__() + " Id: " + self.id.__str__() + \
" Visited: " + self.visited.__str__() + " Weight: " + self.weight.__str__() + \
" Neighbour_count: " + self.neighbour_count.__str__() + " Neighbour_weight: " + self.neighbour_weight.__str__() + \
" Distance: " + self.distance.__str__() + " Sum: " + self.sum.__str__() + \
" N_N_count: " + self.neighbour_neighbour_count.__str__() + " N_N_weight: " + self.neighbour_neighbour_weight.__str__() + \
" N_Distance: " + self.neighbour_distance.__str__() + " N_Sum: " + self.neighbour_sum.__str__() + "\n"
answer = V4(G)
answer=answer[2]
vertex_list = []
for i in range(len(populations)):
v = Vertex(i, populations[i])
vertex_list += [v]
for i in range(len(edges)):
index1, index2, distance = map(int, edges[i].split(" "))
v0 = vertex_list[index1]
v1 = vertex_list[index2]
v0.add_neighbour(v1, distance)
v1.add_neighbour(v0, distance)
for i in range(len(vertex_list)):
Vlist = vertex_list[i].neighbour_list
v0 = vertex_list[i]
for j in range(len(Vlist)):
v0.add_neighbour_neighbour(vertex_list[j])
ret=answer[min(answer.keys())]
ret_val=min(answer.keys())
dict2 = {}
arr = []
for (key, value) in answer.items():
dict2[value] = key
arr += [value]
with open("tests/test2_new.txt") as f:
lines = f.read().splitlines()
number_of_vertices = int(lines[0])
number_of_edges = int(lines[1])
vertices = lines[2:2+number_of_vertices]
edges = lines[2+number_of_vertices:]
ids_and_populations = [tuple(map(int, vertices[i].split(" "))) for i in range(len(vertices))]
populations = dict(ids_and_populations)
mydict = lambda: defaultdict(lambda: defaultdict())
G = mydict()
for i in range(len(edges)):
source, target, weight = map(int, edges[i].split(" "))
G[source][target] = weight
G[target][source] = weight
vertex_list = []
for i in range(len(populations)):
v = Vertex(i, populations[i])
vertex_list += [v]
for i in range(len(edges)):
index1, index2, distance = map(int, edges[i].split(" "))
v0 = vertex_list[index1]
v1 = vertex_list[index2]
v0.add_neighbour(v1, distance)
v1.add_neighbour(v0, distance)
for i in range(len(vertex_list)):
Vlist = vertex_list[i].neighbour_list
v0 = vertex_list[i]
for j in range(len(Vlist)):
v0.add_neighbour_neighbour(vertex_list[j])
ret=answer[min(answer.keys())]
ret_val=min(answer.keys())
dict2 = {}
arr = []
for (key, value) in answer.items():
dict2[value] = key
arr += [value]
learning_rate = 1.0e-8
epochs = 10
bias = 1
sum = 0
def Preceptator(Vertex1, Vertex2, a1, a2, a3, output):
outputP = Vertex1.calculate(a1, a2, a3) + Vertex2.calculate(a1, a2, a3)
error1 = learning_rate * (output - outputP) * a1
error2 = learning_rate * (output - outputP) * a2
error3 = learning_rate * (output - outputP) * a3
a1 += error1
a2 += error2
a3 += error3
return abs(output - outputP)
def getClosestValue(k):
b = int(k)
lst = list(range(b - 1, b + 2, 1)) # lst = [b-1,b,b+1]
return lst[min(range(len(lst)), key=lambda i: abs(lst[i] - k))]
def Predict(Vertex1, Vertex2, a1, a2, a3):
value = Vertex1.calculate(a1, a2, a3) + Vertex2.calculate(a1, a2, a3)
if (value < 0):
print("value : ", int(value) - 1)
elif value == 0:
print("value : ", value)
else:
print("value : ", int(value))
print("y = ", getClosestValue(w[0]), " + ", getClosestValue(w[1]), " * x1 + ", getClosestValue(w[2]), " * x2")
def calculate(Vertex, a1, a2, a3, a4, a5, a6, a7, a8, a9):
return Vertex.weight * a1 + Vertex.neighbour_count * a2 + \
Vertex.neighbour_weight * a3 + Vertex.distance * a4 + \
Vertex.sum * a5 + Vertex.neighbour_neighbour_count * a6 + \
Vertex.neighbour_neighbour_weight * a7 + Vertex.neighbour_distance * a8 + \
Vertex.neighbour_sum * a9
def sigmoid(X, w, w0):
return (1 / (1 + np.exp(-(np.matmul(X, w) + w0))))
def gradient_W(X, y_truth, y_predicted):
return (np.asarray(
[-np.sum(np.repeat((y_truth[:, c] - y_predicted[:, c])[:, None], X.shape[1], axis=1) * X, axis=0) for c in
range(K)]).transpose()) / 3
def gradient_w0(Y_truth, Y_predicted):
return (-np.sum(Y_truth - Y_predicted, axis=0))
# training sets
#for i in range(epochs):
a1 = np.random.rand(1) * 100
a2 = np.random.rand(1) * 10
a3 = np.random.rand(1) * 1
count = 0
for i in range(len(vertex_list)):
for j in range(int(len(vertex_list))):
if i>j:
v0 = vertex_list[i]
v1 = vertex_list[j]
value = arr[count]
count += 1
for j in range(epochs):
val = Preceptator(v0, v1, a1, a2, a3, value)
if j == epochs-1:
sum += val
print(sum/(count)/max(arr))
#Normalizing the variables
coef_sum = a1 + a2 + a3
a1 = a1 / coef_sum
a2 = a2 / coef_sum
a3 = a3 / coef_sum
#with open("tests/test3_new.txt") as f:
#lines = f.read().splitlines()
with open("tests/test3_new.txt") as f:
lines = f.read().splitlines()
number_of_vertices = int(lines[0])
number_of_edges = int(lines[1])
vertices = lines[2:2+number_of_vertices]
edges = lines[2+number_of_vertices:]
ids_and_populations = [tuple(map(int, vertices[i].split(" "))) for i in range(len(vertices))]
populations = dict(ids_and_populations)
mydict = lambda: defaultdict(lambda: defaultdict())
G = mydict()
for i in range(len(edges)):
source, target, weight = map(int, edges[i].split(" "))
G[source][target] = weight
G[target][source] = weight
start = time.time()
vertex_list = []
for i in range(len(populations)):
v = Vertex(i, populations[i])
vertex_list += [v]
for i in range(len(edges)):
index1, index2, distance = map(int, edges[i].split(" "))
v0 = vertex_list[index1]
v1 = vertex_list[index2]
v0.add_neighbour(v1, distance)
v1.add_neighbour(v0, distance)
for i in range(len(vertex_list)):
Vlist = vertex_list[i].neighbour_list
v0 = vertex_list[i]
for j in range(len(Vlist)):
v0.add_neighbour_neighbour(vertex_list[j])
a1 = 92.72
a2 = 9.87
a3 = 0.89
for i in range(len(vertex_list)):
v0 = vertex_list[i]
v0.calculate(a1, a2, a3)
K_arr = []
for i in range(len(vertex_list)):
K_arr += [vertex_list[i].get_key()]
min_val = min(K_arr)
K_arr.sort()
index1 = 0
index2 = 0
for i in range(len(vertex_list)):
v = vertex_list[i]
if K_arr[0] == v.get_key():
index1 = i
if K_arr[1] == v.get_key():
index2 = i
diff = time.time()-start
print('time took: '+str(diff))
print('first node:{}, second node:{} ; associated cost:{}'.format(index1,index2,min_val))
def select_neighbors(G, sub_graph, current_node, k):
if k == 0:
return sub_graph
for j in G[current_node].items():
sub_graph[current_node][j[0]] = j[1]
sub_graph[j[0]][current_node] = j[1]
sub_graph = select_neighbors(G, sub_graph, j[0], k - 1)
return sub_graph
def merge_graph(dict1, dict2):
for key, value in dict2.items():
for subkey, subvalue in value.items():
dict1[key][subkey] = subvalue
def dijkstra_q_impl(G, populations, source):
costs = dict()
for key in G:
costs[key] = np.inf
costs[source] = 0
pq = []
for node in G:
pq.append((node, costs[node]))
while len(pq) != 0:
current_node, current_node_distance = pq.pop(0)
for neighbor_node in G[current_node]:
weight = G[current_node][neighbor_node]
distance = current_node_distance + weight
if distance < costs[neighbor_node]:
costs[neighbor_node] = distance
pq.append((neighbor_node, distance))
#return (costs.values(),population_dict[])
sorted_costs_lst=list(dict(sorted(costs.items())).values())
populations_values_lst = list(dict(sorted(populations.items())).values())
return np.sum(np.array(sorted_costs_lst) * np.array(populations_values_lst))
def random_start(G):
res = [choice(list(G.keys())), choice(list(G.keys()))]
if res[0] == res [1]:
return random_start(G)
print(f"Random start: {res}")
return res
#return [929940, 301820]
#//2 * O((V+E)*logV) = O(E*logV) //
def allocation_cost(G, population_dict, i,j):
return [dijkstra_q_impl(G,population_dict, i),dijkstra_q_impl(G,population_dict, j)]
# V times Dijkstra
def sub_graph_apsp(G, dijkstra_func):
population_dict = dict(sorted([(k, populations[k]) for k in G.keys()]))
selected_vertex = choice(list(G.keys()))
selected_cost = dijkstra_func(G,population_dict, selected_vertex)
for node in G.keys():
if node is not selected_vertex:
this_cost = dijkstra_func(G, population_dict, node)
if this_cost < selected_cost:
selected_cost = this_cost
selected_vertex = node
return selected_vertex, selected_cost
def algorithm_sub_graph_apsp(G, starting_node, k, hop_list, dijkstra_func):
sub_graph = lambda: defaultdict(lambda: defaultdict())
sub_graph = sub_graph()
sub_graph = select_neighbors(G, sub_graph, current_node=starting_node, k=k)
next_node, cost = sub_graph_apsp(sub_graph, dijkstra_func)
#print(next_node)
if len(hop_list) > 0 and next_node == hop_list[-1][0]:
return next_node, cost
hop_list.append((next_node, cost))
return algorithm_sub_graph_apsp(G, next_node, k, hop_list, dijkstra_func)
# 2*O(V)*O(E*logV) = O(E*V*logV) #
def Greedy_Heuristic_Add_Drop(G, dijkstra_func):
population_dict = dict(sorted([(k, populations[k]) for k in G.keys()]))
#population_dict = [populations[i] for i in G.keys()]
selected_vertices = random_start(G)
selected_costs = allocation_cost(G,population_dict, selected_vertices[0],selected_vertices[1])
for not_selected in G.keys():
if not_selected not in selected_vertices:
bigger = max(selected_costs)
this_cost = dijkstra_func(G,population_dict, not_selected)
if this_cost < bigger:
bigger_index = selected_costs.index(bigger)
selected_costs[bigger_index] = this_cost
selected_vertices[bigger_index] = not_selected
return(selected_vertices,selected_costs)
def Greedy_Heuristic_Subgraph_Expansion(G, k, dijkstra_func, bootstrap_cnt=10):
nodes = []
costs = []
for i in range(bootstrap_cnt):
#print("iter")
node, cost = algorithm_sub_graph_apsp(G, choice(list(G.keys())), k, [], dijkstra_func=dijkstra_func)
nodes.append(node)
costs.append(cost)
counter = Counter(nodes)
most_commons = counter.most_common(2)
target_nodes = (most_commons[0][0], most_commons[1][0])
sub_graph1 = lambda: defaultdict(lambda: defaultdict())
sub_graph1 = sub_graph1()
sub_graph1 = select_neighbors(G, sub_graph1, target_nodes[0], k=k)
sub_graph2 = lambda: defaultdict(lambda: defaultdict())
sub_graph2 = sub_graph2()
sub_graph2 = select_neighbors(G, sub_graph2, target_nodes[1], k=k)
merge_graph(sub_graph1, sub_graph2)
points, costs = Greedy_Heuristic_Add_Drop(sub_graph1, dijkstra_func)
if np.inf in costs:
print("INF")
sub_graph1 = lambda: defaultdict(lambda: defaultdict())
sub_graph1 = sub_graph1()
sub_graph1 = select_neighbors(G, sub_graph1, current_node=points[0], k=k+1)
sub_graph2 = lambda: defaultdict(lambda: defaultdict())
sub_graph2 = sub_graph2()
sub_graph2 = select_neighbors(G, sub_graph2, current_node=points[1], k=k+1)
merge_graph(sub_graph1, sub_graph2)
points, costs = Greedy_Heuristic_Add_Drop(sub_graph1, dijkstra_func)
if np.inf not in costs:
return points, costs
else:
print("Graphs are disconnected. Total cost is inf")
return points, costs
return points, costs
start = time.time()
res = Greedy_Heuristic_Subgraph_Expansion(G, 5, bootstrap_cnt=10, dijkstra_func=dijkstra_q_impl) #q for direct Queue based PQ impl (py's pop(0))
diff = time.time()-start
print('\npick cities #'+ str(res[0]) +' with costs '+ str(res[1]))
print('\ntotal time using our Queue-based PQ: '+ str(diff)+ ' sec')
```
| github_jupyter |
# Loading Medicare and Medicaid Claims data into i2b2
[CMS RIF][] docs
This notebook is on demographics.
[CMS RIF]: https://www.resdac.org/cms-data/file-availability#research-identifiable-files
## Python Data Science Tools
especially [pandas](http://pandas.pydata.org/pandas-docs/)
```
import pandas as pd
import numpy as np
import sqlalchemy as sqla
dict(pandas=pd.__version__, numpy=np.__version__, sqlalchemy=sqla.__version__)
```
## DB Access: Luigi Config, Logging
[luigi docs](https://luigi.readthedocs.io/en/stable/)
```
# Passwords are expected to be in the environment.
# Prompt if it's not already there.
def _fix_password():
from os import environ
import getpass
keyname = getpass.getuser().upper() + '_SGROUSE'
if keyname not in environ:
environ[keyname] = getpass.getpass()
_fix_password()
import luigi
def _reset_config(path):
'''Reach into luigi guts and reset the config.
Don't ask.'''
cls = luigi.configuration.LuigiConfigParser
cls._instance = None # KLUDGE
cls._config_paths = [path]
return cls.instance()
_reset_config('luigi-sgrouse.cfg')
luigi.configuration.LuigiConfigParser.instance()._config_paths
import cx_ora_fix
help(cx_ora_fix)
cx_ora_fix.patch_version()
import cx_Oracle as cx
dict(cx_Oracle=cx.__version__, version_for_sqlalchemy=cx.version)
import logging
concise = logging.Formatter(fmt='%(asctime)s %(levelname)s %(message)s',
datefmt='%02H:%02M:%02S')
def log_to_notebook(log,
formatter=concise):
log.setLevel(logging.DEBUG)
to_notebook = logging.StreamHandler()
to_notebook.setFormatter(formatter)
log.addHandler(to_notebook)
return log
from cms_etl import CMSExtract
try:
log.info('Already logging to notebook.')
except NameError:
cms_rif_task = CMSExtract()
log = log_to_notebook(logging.getLogger())
log.info('We try to log non-trivial DB access.')
with cms_rif_task.connection() as lc:
lc.log.info('first bene_id')
first_bene_id = pd.read_sql('select min(bene_id) bene_id_first from %s.%s' % (
cms_rif_task.cms_rif, cms_rif_task.table_eg), lc._conn)
first_bene_id
```
## Demographics: MBSF_AB_SUMMARY, MAXDATA_PS
### Breaking work into groups by beneficiary
```
from cms_etl import BeneIdSurvey
from cms_pd import MBSFUpload
survey_d = BeneIdSurvey(source_table=MBSFUpload.table_name)
chunk_m0 = survey_d.results()[0]
chunk_m0 = pd.Series(chunk_m0, index=chunk_m0.keys())
chunk_m0
dem = MBSFUpload(bene_id_first=chunk_m0.bene_id_first,
bene_id_last=chunk_m0.bene_id_last,
chunk_rows=chunk_m0.chunk_rows)
dem
```
## Column Info: Value Type, Level of Measurement
```
with dem.connection() as lc:
col_data_d = dem.column_data(lc)
col_data_d.head(3)
colprops_d = dem.column_properties(col_data_d)
colprops_d.sort_values(['valtype_cd', 'column_name'])
with dem.connection() as lc:
for x, pct_in in dem.obs_data(lc, upload_id=100):
break
pct_in
x.sort_values(['instance_num', 'valtype_cd']).head(50)
```
### MAXDATA_PS: skip custom for now
```
from cms_pd import MAXPSUpload
survey_d = BeneIdSurvey(source_table=MAXPSUpload.table_name)
chunk_ps0 = survey_d.results()[0]
chunk_ps0 = pd.Series(chunk_ps0, index=chunk_ps0.keys())
chunk_ps0
dem2 = MAXPSUpload(bene_id_first=chunk_ps0.bene_id_first,
bene_id_last=chunk_ps0.bene_id_last,
chunk_rows=chunk_ps0.chunk_rows)
dem2
with dem2.connection() as lc:
col_data_d2 = dem2.column_data(lc)
col_data_d2.head(3)
```
`maxdata_ps` has many groups of columns with names ending in `_1`, `_2`, `_3`, and so on:
```
col_groups = col_data_d2[col_data_d2.column_name.str.match('.*_\d+$')]
col_groups.tail()
pd.DataFrame([dict(all_cols=len(col_data_d2),
cols_in_groups=len(col_groups),
plain_cols=len(col_data_d2) - len(col_groups))])
from cms_pd import col_valtype
def _cprop(cls, valtype_override, info: pd.DataFrame) -> pd.DataFrame:
info['valtype_cd'] = [col_valtype(c).value for c in info.column.values]
for cd, pat in valtype_override:
info.valtype_cd = info.valtype_cd.where(~ info.column_name.str.match(pat), cd)
info.loc[info.column_name.isin(cls.i2b2_map.values()), 'valtype_cd'] = np.nan
return info.drop('column', 1)
_vo = [
('@', r'.*race_code_\d$'),
('@custom_postpone', r'.*_\d+$')
]
#dem2.column_properties(col_data_d2)
colprops_d2 = _cprop(dem2.__class__, _vo, col_data_d2)
colprops_d2.query('valtype_cd != "@custom_postpone"').sort_values(['valtype_cd', 'column_name'])
colprops_d2.dtypes
```
## Patient, Encounter Mapping
```
obs_facts = obs_dx.append(obs_cd).append(obs_num).append(obs_txt).append(obs_dt)
with cc.connection('patient map') as lc:
pmap = cc.patient_mapping(lc, (obs_facts.bene_id.min(), obs_facts.bene_id.max()))
from etl_tasks import I2B2ProjectCreate
obs_patnum = obs_facts.merge(pmap, on='bene_id')
obs_patnum.sort_values('start_date').head()[[
col.name for col in I2B2ProjectCreate.observation_fact_columns
if col.name in obs_patnum.columns.values]]
with cc.connection() as lc:
emap = cc.encounter_mapping(lc, (obs_dx.bene_id.min(), obs_dx.bene_id.max()))
emap.head()
'medpar_id' in obs_patnum.columns.values
obs_pmap_emap = cc.pat_day_rollup(obs_patnum, emap)
x = obs_pmap_emap
(x[(x.encounter_num > 0) | (x.encounter_num % 8 == 0) ][::5]
.reset_index().set_index(['patient_num', 'start_date', 'encounter_num']).sort_index()
.head(15)[['medpar_id', 'start_day', 'admsn_dt', 'dschrg_dt', 'concept_cd']])
```
### Provider etc. done?
```
obs_mapped = cc.with_mapping(obs_dx, pmap, emap)
obs_mapped.columns
[col.name for col in I2B2ProjectCreate.observation_fact_columns
if not col.nullable and col.name not in obs_mapped.columns.values]
test_run = False
if test_run:
cc.run()
```
## Drugs: PDE
```
from cms_pd import DrugEventUpload
du = DrugEventUpload(bene_id_first=bene_chunks.iloc[0].bene_id_first,
bene_id_last=bene_chunks.iloc[0].bene_id_last,
chunk_rows=bene_chunks.iloc[0].chunk_rows,
chunk_size=1000)
with du.connection() as lc:
du_cols = du.column_data(lc)
du.column_properties(du_cols).sort_values('valtype_cd')
with du.connection() as lc:
for x, pct_in in du.obs_data(lc, upload_id=100):
break
x.sort_values(['instance_num', 'valtype_cd']).head(50)
```
## Performance Results
```
bulk_migrate = '''
insert /*+ parallel(24) append */ into dconnolly.observation_fact
select * from dconnolly.observation_fact_2440
'''
with cc.connection() as lc:
lc.execute('truncate table my_plan_table')
print(lc._conn.engine.url.query)
print(pd.read_sql('select count(*) from my_plan_table', lc._conn))
lc._conn.execute('explain plan into my_plan_table for ' + bulk_migrate)
plan = pd.read_sql('select * from my_plan_table', lc._conn)
plan
with cc.connection() as lc:
lc.execute('truncate table my_plan_table')
print(pd.read_sql('select * from my_plan_table', lc._conn))
db = lc._conn.engine
cx = db.dialect.dbapi
dsn = cx.makedsn(db.url.host, db.url.port, db.url.database)
conn = cx.connect(db.url.username, db.url.password, dsn,
threaded=True, twophase=True)
cur = conn.cursor()
cur.execute('explain plan into my_plan_table for ' + bulk_migrate)
cur.close()
conn.commit()
conn.close()
plan = pd.read_sql('select * from my_plan_table', lc._conn)
plan
select /*+ parallel(24) */ max(bene_enrollmt_ref_yr)
from cms_deid.mbsf_ab_summary;
select * from upload_status
where upload_id >= 2799 -- and message is not null -- 2733
order by upload_id desc;
-- order by end_date desc;
select load_status, count(*), min(upload_id), max(upload_id), min(load_date), max(end_date)
, to_char(sum(loaded_record), '999,999,999') loaded_record
, round(sum(loaded_record) / 1000 / ((max(end_date) - min(load_date)) * 24 * 60)) krows_min
from (
select upload_id, loaded_record, load_status, load_date, end_date, end_date - load_date elapsed
from upload_status
where upload_label like 'MBSFUp%'
)
group by load_status
;
```
## Reimport code into running notebook
```
import importlib
import cms_pd
import cms_etl
import etl_tasks
import eventlog
import script_lib
importlib.reload(script_lib)
importlib.reload(eventlog)
importlib.reload(cms_pd)
importlib.reload(cms_etl)
importlib.reload(etl_tasks);
```
| github_jupyter |
# Project - Feature Engineering on the Titanic
The titanic dataset has a few columns from which you can use regex to extract information from. Feature engineering involves using existing columns of data to create new columns of data. You will work on doing just that in these exercises. Read it in and then answer the following questions.
```
import pandas as pd
titanic = pd.read_csv('../data/titanic.csv')
titanic.head()
```
## Exercises
### Exercise 1
<span style="color:green; font-size:16px">Extract the first character of the `Ticket` column and save it as a new column `ticket_first`. Find the total number of survivors, the total number of passengers, and the percentage of those who survived **by this column**. Next find the total survival rate for the entire dataset. Does this new column help predict who survived?</span>
### Exercise 2
<span style="color:green; font-size:16px">If you did Exercise 2 correctly, you should see that only 7% of the people with tickets that began with 'A' survived. Find the survival rate for all those 'A' tickets by `Sex`.</span>
### Exercise 3
<span style="color:green; font-size:16px">Find the survival rate by the last letter of the ticket. Is there any predictive power here?</span>
### Exercise 4
<span style="color:green; font-size:16px">Find the length of each passengers name and assign to the `name_len` column. What is the minimum and maximum name length?</span>
### Exercise 5
<span style="color:green; font-size:16px">Pass the `name_len` column to the `pd.cut` function. Also, pass a list of equal-sized cut points to the `bins` parameter. Assign the resulting Series to the `name_len_cat` column. Find the frequency count of each bin in this column.</span>
### Exercise 6
<span style="color:green; font-size:16px">Is name length a good predictor of survival?<span>
### Exercise 7
<span style="color:green; font-size:16px">Why do you think people with longer names had a better chance at survival?</span>
### Exercise 8
<span style="color:green; font-size:16px">Using the titanic dataset, do your best to extract the title from a person's name. Examples of title are 'Mr.', 'Dr.', 'Miss', etc... Save this to a column called `title`. Find the frequency count of the titles.</span>
### Exercise 9
<span style="color:green; font-size:16px">Does the title have good predictive value of survival?</span>
### Exercise 10
<span style="color:green; font-size:16px">Create a pivot table of survival by title and sex. Use two aggregation functions, mean and size</span>
### Exercise 11
<span style="color:green; font-size:16px">Attempt to extract the first name of each passenger into the column `first_name`. Are there are males and females with the same first name?</span>
### Exercise 12
<span style="color:green; font-size:16px">The exercises have been an exercise in feature engineering. Several new features (columns) have been created from existing columns. Come up with your own feature and test it out on survival.</span>
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/Join/inverted_joins.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/Join/inverted_joins.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/Join/inverted_joins.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API and geemap
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geemap](https://github.com/giswqs/geemap). The **geemap** Python package is built upon the [ipyleaflet](https://github.com/jupyter-widgets/ipyleaflet) and [folium](https://github.com/python-visualization/folium) packages and implements several methods for interacting with Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, and `Map.centerObject()`.
The following script checks if the geemap package has been installed. If not, it will install geemap, which automatically installs its [dependencies](https://github.com/giswqs/geemap#dependencies), including earthengine-api, folium, and ipyleaflet.
**Important note**: A key difference between folium and ipyleaflet is that ipyleaflet is built upon ipywidgets and allows bidirectional communication between the front-end and the backend enabling the use of the map to capture user input, while folium is meant for displaying static data only ([source](https://blog.jupyter.org/interactive-gis-in-jupyter-with-ipyleaflet-52f9657fa7a)). Note that [Google Colab](https://colab.research.google.com/) currently does not support ipyleaflet ([source](https://github.com/googlecolab/colabtools/issues/60#issuecomment-596225619)). Therefore, if you are using geemap with Google Colab, you should use [`import geemap.eefolium`](https://github.com/giswqs/geemap/blob/master/geemap/eefolium.py). If you are using geemap with [binder](https://mybinder.org/) or a local Jupyter notebook server, you can use [`import geemap`](https://github.com/giswqs/geemap/blob/master/geemap/geemap.py), which provides more functionalities for capturing user input (e.g., mouse-clicking and moving).
```
# Installs geemap package
import subprocess
try:
import geemap
except ImportError:
print('geemap package not installed. Installing ...')
subprocess.check_call(["python", '-m', 'pip', 'install', 'geemap'])
# Checks whether this notebook is running on Google Colab
try:
import google.colab
import geemap.eefolium as geemap
except:
import geemap
# Authenticates and initializes Earth Engine
import ee
try:
ee.Initialize()
except Exception as e:
ee.Authenticate()
ee.Initialize()
```
## Create an interactive map
The default basemap is `Google Maps`. [Additional basemaps](https://github.com/giswqs/geemap/blob/master/geemap/basemaps.py) can be added using the `Map.add_basemap()` function.
```
Map = geemap.Map(center=[40,-100], zoom=4)
Map
```
## Add Earth Engine Python script
```
# Add Earth Engine dataset
# Load a Landsat 8 image collection at a point of interest.
collection = ee.ImageCollection('LANDSAT/LC08/C01/T1_TOA') \
.filterBounds(ee.Geometry.Point(-122.09, 37.42))
# Define start and end dates with which to filter the collections.
april = '2014-04-01'
may = '2014-05-01'
june = '2014-06-01'
july = '2014-07-01'
# The primary collection is Landsat images from April to June.
primary = collection.filterDate(april, june)
# The secondary collection is Landsat images from May to July.
secondary = collection.filterDate(may, july)
# Use an equals filter to define how the collections match.
filter = ee.Filter.equals(**{
'leftField': 'system:index',
'rightField': 'system:index'
})
# Define the join.
invertedJoin = ee.Join.inverted()
# Apply the join.
invertedJoined = invertedJoin.apply(primary, secondary, filter)
# Display the result.
print('Inverted join: ', invertedJoined.getInfo())
```
## Display Earth Engine data layers
```
Map.addLayerControl() # This line is not needed for ipyleaflet-based Map.
Map
```
| github_jupyter |
# Függvények
A legtöbb programozási nyelv engedi, hogy a programozó által írt programokat külön, újrafelhasználható részekbe, függvényekbe szervezze. Ezek segítségével a kód olvashatóbb és fenntarthatóbb lesz.
A függvényekbe a következőket lehet írni:
* A paraméterek listája, ami lehet üres is, ekkor nem vár a függvény paramétert.
* Az első sor tartalmazhat egy string-et, ami a függvény dokumentációja lesz (docstring). Ezt használják bizonyos programok dokumentáció generálására.
* A függvény törzsébe a lefuttatni kívánt utasítások kerülnek.
* A függvénynek lehet explicit visszatérési értéke, amit a `return` kulcsszóval adunk meg. Ha ez nem szerepel, akkor van egy implicit visszatérési érték: `None`
Függvény definiálása a `def` kulcsszóval lehetséges, amit a függvény neve és a paraméterek listája követ:
```
def addtwo(a):
r = a + 2
print(r)
```
Figyeljünk az indentálásra! Ha már definiálva van a függvény, *meghívhatjuk* tetszőleges paraméterrel:
```
addtwo(2)
```
Ha rossz *típusú* dolgot adunk át, hibát kaphatunk:
```
addtwo("Hello")
```
A paraméter nélküli függvények meghívásakor is ki kell írni a zárójeleket:
```
def hello():
print("Hello")
hello()
```
A ciklusoknál írtunk egy ciklust, ami megadja az első `n` darab Fibonacci-számot. Írjuk át ezt egy függvénybe, ami azt várja paraméterül, hogy hány számot írjon ki a függvény!
```
def fibonacci(n):
a, b = 0, 1
while n > 0:
print(a)
a, b = b, a + b
n -= 1
fibonacci(10)
```
## Visszatérési értékek megadása
Egy függvény rendelkezhet visszatérési értékkel annak érdekében, hogy a függvény eredményét később fel tudjuk használni. A visszatérési értéket a `return` kulcsszóval adhatjuk meg. Ahogy korábban volt szó róla, ha nem adunk meg egy visszatérési értéket, akkor is rendelkezni fog egyel a függvény. Ez lesz a `None`, amit más nyelvekben null, nil néven neveznek. A `None` egy speciális érték, aminek az intuitív jelentése, hogy "nincs itt semmi", és a típusa `NoneType`.
A következő függvénynek tehát `None` lesz a visszatérési értéke, mert nincs benne `return` érték:
```
def timesthree(n):
r = n * 3
```
A `print()` függvényt használva kiírható a visszatérési érték ilyenkor is:
```
print(timesthree(3))
```
Az előző függvény visszatérési értéke legyen a művelet eredménye:
```
def timesthree(n):
r = n * 3
return r
print(timesthree(3))
```
A függvények visszatérési értékét természetesen átadhatjuk egy változónak:
```
x = timesthree(3)
print(x)
```
## Alapértelmezett paraméterek
Ha egy argumentumnak alapértelmezett értéket akarunk adni, akkor a függvény definiálásakor megtehetjük kulcs-érték párok segítségével:
```
def hello(who, num=10):
return ("Hello, " + who + "\n") * num
```
Alapértelmezett argumentummal rendelkező függvényt többféleképpen meg lehet hívni:
```
print(hello("world"))
print(hello(who="Zénó"))
print(hello(who="Zebulon", num=1))
```
De ha a kötelezően megadandó argumentumot nem adjuk meg, hibát kapunk:
```
print(hello(num=1))
```
Az alapértelmezett argumentumokra a következő szabályok vonatkoznak:
* a kötelezően megadandó argumentum a lista elejére kerül
* az kulcs-érték párokkal megadott alapértelmezett argumentumok a kötelezően megadandó argumentumok után kerülnek
* egy argumentumot legfeljebb egyszer lehet átadni
* legfeljebb annyi argumentumot lehet megadni a híváskor, amennyit definiáláskor megadtunk (ez kiterjeszthető)
**Fontos**: az alapértelmezett argumentum pontosan egyszer, a függvény definiálásakor értékelődik ki:
```
i = 5
def f(arg=i):
print(arg)
i = 6
f()
```
A Python ezen tulajdonsága miatt mutable objektumot (pl: listát) nem szabad megadni alapértelmezett argumentumként (habár a Python nem szól miatta):
```
def f(a, L=[]):
L.append(a)
return L
print(f(1))
print(f(2))
print(f(3))
```
Lista esetén a fenti függvény összes meghívásakor a legfrisebb lista lesz az alapértelmezett argumentum.
## Dokumentációs string (*docstring*)
A dokumentációs string szerepe, hogy elmagyarázza a felhasználónak, mit csinál a függvény. A szabályok a következők:
- az első sor egy rövid, tömör leírása a függvény működésének
- további bekezdések hozzáadhatók, ebben részletezni lehet a működést
- megadhatók továbbá a paraméterek jelentései és a visszatérési érték
A dokumentációs string nem csinál semmit, csak dokumentálja a függvény működését. Ebből szoktak az automatikus dokumentum-generáló programok dokumentumot generálni.
```
def f():
"""Rövid leírása a függvénynek.
Innentől meg jöhetnek a hosszabb dolgok,
mint paraméterek és visszatérési érték leírása.
Külön bekezdésekbe illik szedni a nagyobb
egységeket.
Ez a docstring nem csinál semmit, csak dokumentál.
"""
print("Hello")
```
Jupyter-ben két módja van a docstring lekérésének. Az egyik a Python-ba beépített `help()` függvény, aminek a paraméterébe megadjuk a függvény *nevét* zárójelek nélkül:
```
help(f)
```
A másik módja: a program írása közben a függvény zárójelei között Shift+Tab billentyűkömbinációval lekérhetjük a dokumentációt. Más fejlesztői környezetekben ez máshogy megy.
```
f()
```
## Lambda kifejezések
Egyszerű függvények megadhatók névtelenül a `lambda` kulcsszóval. A következő függvény visszaadja a paraméterül kapott két szám összegét:
```
f = lambda x, y: x + y
```
Ami ezzel a függvénnyel ekvivalens
```
def f(x, y):
return x + y
```
A `lambda` függvény használatára egy példa:
```
f(3, 4)
```
Akkor hasznosak, ha egy függvényt szeretnénk átadni egy másik függvénynek és nem akarjuk fölöslegesen deiniálni azt (mert mondjuk csak egyszer használnánk). A listáknál a `sort()` metódus ilyen, ott megadhatjuk, hogy milyen elv szerint rendezze a lista elemeit:
```
pairs = [(1, 'one'), (2, 'two'), (3, 'three'), (4, 'four')]
pairs.sort(key=lambda pair: pair[1])
print(pairs)
```
A fenti kód ekvivalens ezzel:
```
pairs = [(1, 'one'), (2, 'two'), (3, 'three'), (4, 'four')]
def f(pair):
return pair[1]
pairs.sort(key=f)
print(pairs)
```
## Feladatok
### **1. feladat**: Írjuk át a Fibonacci-sorozatot megadó függvényt úgy, hogy a sorozat első `n` elemének a listájával térjen vissza! Írjunk hozzá docstring-et is! A sorozat elemeit ne írja ki a függvény.
```
# megoldás:
def fib(n):
"""Megadja a Fibonacci-sorozat első n elemét."""
a, b = 0, 1
res = []
while n > 0:
res.append(a)
a, b = b, a + b
n -= 1
return res
print(fib(10))
```
### **2. feladat**: A szinusz függvény közelíthető az alábbi sorozattal:
$$\sin x \simeq (-1)^n \frac{x^{2n+1}}{(2n+1)!}$$
Ez a sorozat az első `n` tag összegével közelíti a szinuszt, de figyeljünk, hogy az `x` radiánban, és nem fokban van megadva! Ha például csak az első 3 tagot vesszük figyelembe:
$$\sin x \simeq x - \frac{x^3}{3!} + \frac{x^5}{5!}$$
Írjunk egy függvényt, ami a fenti végtelen sorozat alapján megadja a szinusz közelítő értékét!
Tipp: használjuk az itt definiált `factorial()` függvényt, ami egy természetes szám faktoriálisát számolja ki!
```
def factorial(n):
if n == 0 or n == 1:
return 1
result = 1
for i in range(1, n+1):
result *= i
return result
# megoldás:
def sin_approx(x, n):
res = 0
x /= 57.29577951 # a szöget radiánba számoljuk át
for i in range(n):
res += (-1) ** i * x ** (2*i + 1) / factorial(2*i + 1)
return res
print(sin_approx(45, 10)) # 45 fok szinusza
```
### **3. feladat**: Írjunk egy függvényt, ami két `float` típusú számról megmondja, hogy *nagyjából* egyenlők-e!
A *nagyjából* egyenlő azt jelenti, hogy a két szám valamilyen pontossággal megegyezik. Ehhez vegyünk fel egy argumentumot, aminek a változtatásával tetszőlegesen pontosan össze tudunk számokat hasonlítani. Legyen az alapértelmezett pontosság `0.00001`! Lásd a példákat!
Tipp: használjuk a beépített `abs()` függvényt, ami megadja egy szám abszolút értékét!
A függvény futására példák:
```python
print(isclose(0.33333, 1/3))
True
print(isclose(0.3333, 1/3))
False
print(isclose(0.3333, 1/3, 1e-4))
True
```
```
# megoldás:
def isclose(a, b, eps=1e-5):
return abs(a - b) <= eps
print(isclose(0.33333, 1/3))
print(isclose(0.3333, 1/3))
print(isclose(0.3333, 1/3, 1e-4))
```
| github_jupyter |
```
pwd
import pandas as pd
df=pd.read_csv('/home/sf/fresh_start/3_class_combination_10-5.csv')
df
test_subject=[int(i) for i in list(df[df['acc']==df['acc'].max()]['subjects_in_test'])[0][1:-1].split(',')]
train_subject=[int(i) for i in list(df[df['acc']==df['acc'].max()]['subjects_in_train'])[0][1:-1].split(',')]
test_subject
import numpy as np
from sklearn.ensemble import ExtraTreesClassifier
from sklearn.metrics import classification_report
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier
from sklearn.tree import DecisionTreeClassifier
from imblearn.over_sampling import SMOTE
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import Normalizer
from itertools import combinations
from sklearn import model_selection
import copy
from statistics import mean,mode
from itertools import combinations
from sklearn.ensemble import GradientBoostingClassifier
df=pd.read_csv('/home/sf/fresh_start/60s_window_wrist_chest.csv',index_col=0)
df=df[df['label']<3]
features=df.columns.tolist()
features
removed = ['label']
for rem in removed:
features.remove(rem)
features_with_sub=[]
features_with_sub[:]=features
removed = ['subject']
for rem in removed:
features.remove(rem)
feature=features
print(len(feature))
len(features_with_sub)
sm = SMOTE(random_state=2)
X, y= sm.fit_sample(df[features_with_sub], df['label'])
df_new=pd.concat([pd.DataFrame(X,columns=features_with_sub),pd.DataFrame(y,columns=['label'])],axis=1)
df_new
for i in range (len(list(df_new['subject']))):
df_new['subject'][i] = min([2,3,4,5,6,7,8,9,10,11,13,14,15,16,17], key=lambda x:abs(x-df_new['subject'][i]))
df_new['subject']=df_new['subject'].astype(int)
p_d=pd.read_csv('/home/sf/fresh_start/personal_detail.csv',index_col=0)
df_new_1=df_new.merge(p_d,on='subject')
df_new_1
sel_fea=['EDA_tonic_mean',
'EDA_tonic_max',
'EDA_tonic_min',
'EDA_phasic_mean',
'EDA_smna_mean',
'EDA_phasic_min',
'EMG_std',
'c_ACC_y_min',
'sport_today_YES',
'ECG_std',
'c_ACC_x_std',
'c_ACC_y_std']
# subjects_in_train = []
# subjects_in_test = []
# best_acc = []
# mean_acc = []
# min_acc = []
# acc = []
# for cp in range (1,len(train_subject)):
# print ('*'*20)
# print ("10C"+str(cp))
# print ('*'*20)
# com = cp # combination number, If any doubt plz call me
# combi = combinations(train_subject, com)
# tot = str(len(list(copy.deepcopy(combi))))
# list_combi = list(combi)
# for lc in list_combi:
# print (list(lc))
# train= df_new_1.loc[df_new_1.subject.isin(list(lc))]
# test= df_new_1.loc[df_new_1.subject.isin(test_subject)]
# print ("TRAIN",lc)
# print ("TEST",test_subject )
# scaler = Normalizer()
# scaled_data_train = scaler.fit_transform(train[sel_fea])
# scaled_data_test = scaler.transform(test[sel_fea])
# clf = ExtraTreesClassifier(n_estimators=100,n_jobs=10)
# clf.fit(scaled_data_train,train['label'])
# y_pred=clf.predict(scaled_data_test)
# #print (classification_report(test['label'],y_pred))
# rpt = classification_report(test['label'],y_pred,output_dict=True)['accuracy']
# acc.append(rpt)
# subjects_in_train.append(str(list(lc)))
# subjects_in_test.append(str(test_subject))
# combi_dict = {'subjects_in_train':subjects_in_train,'subjects_in_test':subjects_in_test, 'acc':acc}
# df_plot_combi = pd.DataFrame(combi_dict)
for cp in range (1,len(train_subject)):
print ('*'*20)
print ("10C"+str(cp))
print ('*'*20)
com = cp # combination number, If any doubt plz call me
combi = combinations(train_subject, com)
tot = str(len(list(copy.deepcopy(combi))))
list_combi = list(combi)
subjects_in_train = []
subjects_in_test = []
best_acc = []
mean_acc = []
min_acc = []
acc = []
for lc in list_combi:
print (list(lc))
train= df_new_1.loc[df_new_1.subject.isin(list(lc))]
test= df_new_1.loc[df_new_1.subject.isin(test_subject)]
scaler = Normalizer()
scaled_data_train = scaler.fit_transform(train[sel_fea])
scaled_data_test = scaler.transform(test[sel_fea])
clf = DecisionTreeClassifier()
clf.fit(scaled_data_train,train['label'])
y_pred=clf.predict(scaled_data_test)
rpt = classification_report(test['label'],y_pred,output_dict=True)['accuracy']
acc.append(rpt)
subjects_in_train.append(str(list(lc)))
subjects_in_test.append(str(test_subject))
combi_dict = {'subjects_in_train':subjects_in_train,'subjects_in_test':subjects_in_test, 'acc':acc}
df_plot_combi = pd.DataFrame(combi_dict)
file_name = '3_class_combination_'+str(cp)+'-'+str(5)+'.csv'
print (file_name)
df_plot_combi.to_csv(file_name)
df_plot_combi
df_plot_combi.to_csv("as_you_asked_ram.csv")
# %%time
# for cp in range (1,len(user_list)):
# print ('*'*20)
# print ("15C"+str(cp))
# print ('*'*20)
# com = cp # combination number, If any doubt plz call me
# combi = combinations(user_list, com)
# tot = str(len(list(copy.deepcopy(combi))))
# # getting the best random state
# best_random_state_train = user_list[0:com]
# best_random_state_test = user_list[com:]
# # print (best_random_state_train)
# # print (best_random_state_test)
# train= df_new_1.loc[df_new_1.subject.isin(best_random_state_train)]
# test= df_new_1.loc[df_new_1.subject.isin(best_random_state_test)]
# scaler = Normalizer()
# scaled_data_train = scaler.fit_transform(train[sel_fea])
# scaled_data_test = scaler.transform(test[sel_fea])
# rnd_loc_acc = []
# for i in range (101):
# clf = ExtraTreesClassifier(n_estimators=100,n_jobs=10,random_state=i)
# clf.fit(scaled_data_train,train['label'])
# y_pred=clf.predict(scaled_data_test)
# #print (classification_report(test['label'],y_pred))
# rpt = classification_report(test['label'],y_pred,output_dict=True)['accuracy']
# rnd_loc_acc.append(rpt)
# rnd_index = rnd_loc_acc.index(max(rnd_loc_acc))
# index = 1
# subjects_in_train = []
# subjects_in_test = []
# best_acc = []
# mean_acc = []
# min_acc = []
# acc = []
# for c in list(combi):
# local_acc = []
# # print (str(index)+" of "+ tot)
# train_sub = list(c)
# test_sub = list(set(user_list)-set(train_sub))
# print (train_sub,test_sub)
# train= df_new_1.loc[df_new_1.subject.isin(train_sub)]
# test= df_new_1.loc[df_new_1.subject.isin(test_sub)]
# scaler = Normalizer()
# scaled_data_train = scaler.fit_transform(train[sel_fea])
# scaled_data_test = scaler.transform(test[sel_fea])
# clf = ExtraTreesClassifier(n_estimators=100,n_jobs=10,random_state=rnd_index)
# clf.fit(scaled_data_train,train['label'])
# y_pred=clf.predict(scaled_data_test)
# #print (classification_report(test['label'],y_pred))
# rpt = classification_report(test['label'],y_pred,output_dict=True)['accuracy']
# acc.append(rpt)
# subjects_in_train.append(str(train_sub))
# subjects_in_test.append(str(test_sub))
# # for i in range (51):
# # print (i)
# # clf = ExtraTreesClassifier(n_estimators=100,n_jobs=10,random_state=i)
# # clf.fit(scaled_data_train,train['label'])
# # y_pred=clf.predict(scaled_data_test)
# # # print (classification_report(test['label'],y_pred))
# # rpt = classification_report(test['label'],y_pred,output_dict=True)['accuracy']
# # local_acc.append(rpt)
# # best_acc.append(max(local_acc))
# # mean_acc.append(mean(local_acc))
# # min_acc.append(min(local_acc))
# # subjects_in_train.append(str(train_sub))
# # subjects_in_test.append(str(test_sub))
# # print ("*"*10)
# # print (acc)
# # print ("*"*10)
# index += 1
# combi_dict = {'subjects_in_train':subjects_in_train,'subjects_in_test':subjects_in_test, 'acc':acc}
# df_plot_combi = pd.DataFrame(combi_dict)
# temp = df_plot_combi[df_plot_combi['acc']>=max(df_plot_combi['acc'])]
# subjects_in_train = eval(temp['subjects_in_train'].values[0])
# subjects_in_test = eval(temp['subjects_in_test'].values[0])
# train= df_new_1.loc[df_new_1.subject.isin(subjects_in_train)]
# test= df_new_1.loc[df_new_1.subject.isin(subjects_in_test)]
# scaler = Normalizer()
# scaled_data_train = scaler.fit_transform(train[sel_fea])
# scaled_data_test = scaler.transform(test[sel_fea])
# print("****** Testing on Model ********")
# #extra tree
# print ("Extra tree")
# loc_acc = []
# for i in range (101):
# clf = ExtraTreesClassifier(n_estimators=100,n_jobs=10,random_state=i)
# clf.fit(scaled_data_train,train['label'])
# y_pred=clf.predict(scaled_data_test)
# #print (classification_report(test['label'],y_pred))
# rpt = classification_report(test['label'],y_pred,output_dict=True)['accuracy']
# loc_acc.append(rpt)
# index = loc_acc.index(max(loc_acc))
# clf = ExtraTreesClassifier(n_estimators=100,n_jobs=10,random_state=index)
# clf.fit(scaled_data_train,train['label'])
# y_pred=clf.predict(scaled_data_test)
# rpt = classification_report(test['label'],y_pred,output_dict=True)['accuracy']
# print (classification_report(test['label'],y_pred))
# df_plot_combi.at[df_plot_combi[df_plot_combi['acc'] == max(df_plot_combi['acc'])].index[0],'acc'] = rpt
# #random forest
# print ("Random Forest")
# loc_acc = []
# for i in range (101):
# clf=RandomForestClassifier(n_estimators=50,random_state=i)
# clf.fit(scaled_data_train,train['label'])
# y_pred=clf.predict(scaled_data_test)
# #print (classification_report(test['label'],y_pred))
# rpt = classification_report(test['label'],y_pred,output_dict=True)['accuracy']
# loc_acc.append(rpt)
# index = loc_acc.index(max(loc_acc))
# clf=RandomForestClassifier(n_estimators=50,random_state=index)
# clf.fit(scaled_data_train,train['label'])
# y_pred=clf.predict(scaled_data_test)
# print (classification_report(test['label'],y_pred))
# #Decision-Tree
# print ("Decision Tree")
# loc_acc = []
# for i in range (101):
# clf= DecisionTreeClassifier(random_state=i)
# clf.fit(scaled_data_train,train['label'])
# y_pred=clf.predict(scaled_data_test)
# #print (classification_report(test['label'],y_pred))
# rpt = classification_report(test['label'],y_pred,output_dict=True)['accuracy']
# loc_acc.append(rpt)
# index = loc_acc.index(max(loc_acc))
# clf= DecisionTreeClassifier(random_state=index)
# clf.fit(scaled_data_train,train['label'])
# y_pred=clf.predict(scaled_data_test)
# print (classification_report(test['label'],y_pred))
# #GradientBoosting
# print ("Gradient Boosting")
# loc_acc = []
# for i in range (101):
# clf= GradientBoostingClassifier(random_state=i)
# clf.fit(scaled_data_train,train['label'])
# y_pred=clf.predict(scaled_data_test)
# #print (classification_report(test['label'],y_pred))
# rpt = classification_report(test['label'],y_pred,output_dict=True)['accuracy']
# loc_acc.append(rpt)
# index = loc_acc.index(max(loc_acc))
# clf= GradientBoostingClassifier(random_state=index)
# clf.fit(scaled_data_train,train['label'])
# y_pred=clf.predict(scaled_data_test)
# print (classification_report(test['label'],y_pred))
# print("****** Writing to File ********")
# # Plz cross check with the file name before saving to df to csv file
# file_name = '3_class_combination_'+str(com)+'-'+str(15-com)+'.csv'
# print (file_name)
# df_plot_combi.to_csv(file_name)
# temp = df_plot_combi[df_plot_combi['acc']>=max(df_plot_combi['acc'])]
# print("Max:",max(df_plot_combi['acc']))
# print("Min:",min(df_plot_combi['acc']))
# print("Mean:",mean(df_plot_combi['acc']))
list_combi
```
| github_jupyter |
```
# Copyright 2021 NVIDIA Corporation. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# =======
```
# ETL with NVTabular
In this notebook we are going to generate synthetic data and then create sequential features with [NVTabular](https://github.com/NVIDIA-Merlin/NVTabular). Such data will be used in the next notebook to train a session-based recommendation model.
NVTabular is a feature engineering and preprocessing library for tabular data designed to quickly and easily manipulate terabyte scale datasets used to train deep learning based recommender systems. It provides a high level abstraction to simplify code and accelerates computation on the GPU using the RAPIDS cuDF library.
### Import required libraries
```
import os
import glob
import numpy as np
import pandas as pd
import cudf
import cupy as cp
import nvtabular as nvt
```
### Define Input/Output Path
```
INPUT_DATA_DIR = os.environ.get("INPUT_DATA_DIR", "/workspace/data/")
```
## Create a Synthetic Input Data
```
NUM_ROWS = 100000
long_tailed_item_distribution = np.clip(np.random.lognormal(3., 1., NUM_ROWS).astype(np.int32), 1, 50000)
# generate random item interaction features
df = pd.DataFrame(np.random.randint(70000, 80000, NUM_ROWS), columns=['session_id'])
df['item_id'] = long_tailed_item_distribution
# generate category mapping for each item-id
df['category'] = pd.cut(df['item_id'], bins=334, labels=np.arange(1, 335)).astype(np.int32)
df['timestamp/age_days'] = np.random.uniform(0, 1, NUM_ROWS)
df['timestamp/weekday/sin']= np.random.uniform(0, 1, NUM_ROWS)
# generate day mapping for each session
map_day = dict(zip(df.session_id.unique(), np.random.randint(1, 10, size=(df.session_id.nunique()))))
df['day'] = df.session_id.map(map_day)
```
- Visualize couple of rows of the synthetic dataset
```
df.head()
```
## Feature Engineering with NVTabular
Deep Learning models require dense input features. Categorical features are sparse, and need to be represented by dense embeddings in the model. To allow for that, categorical features need first to be encoded as contiguous integers `(0, ..., |C|)`, where `|C|` is the feature cardinality (number of unique values), so that their embeddings can be efficiently stored in embedding layers. We will use NVTabular to preprocess the categorical features, so that all categorical columns are encoded as contiguous integers. Note that in the `Categorify` op we set `start_index=1`, the reason for that we want the encoded null values to start from `1` instead of `0` because we reserve `0` for padding the sequence features.
Here our goal is to create sequential features. In this cell, we are creating temporal features and grouping them together at the session level, sorting the interactions by time. Note that we also trim each feature sequence in a session to a certain length. Here, we use the NVTabular library so that we can easily preprocess and create features on GPU with a few lines.
```
# Categorify categorical features
categ_feats = ['session_id', 'item_id', 'category'] >> nvt.ops.Categorify(start_index=1)
# Define Groupby Workflow
groupby_feats = categ_feats + ['day', 'timestamp/age_days', 'timestamp/weekday/sin']
# Groups interaction features by session and sorted by timestamp
groupby_features = groupby_feats >> nvt.ops.Groupby(
groupby_cols=["session_id"],
aggs={
"item_id": ["list", "count"],
"category": ["list"],
"day": ["first"],
"timestamp/age_days": ["list"],
'timestamp/weekday/sin': ["list"],
},
name_sep="-")
# Select and truncate the sequential features
sequence_features_truncated = (groupby_features['category-list', 'item_id-list', 'timestamp/age_days-list', 'timestamp/weekday/sin-list']) >>nvt.ops.ListSlice(0,20) >> nvt.ops.Rename(postfix = '_trim')
# Filter out sessions with length 1 (not valid for next-item prediction training and evaluation)
MINIMUM_SESSION_LENGTH = 2
selected_features = groupby_features['item_id-count', 'day-first', 'session_id'] + sequence_features_truncated
filtered_sessions = selected_features >> nvt.ops.Filter(f=lambda df: df["item_id-count"] >= MINIMUM_SESSION_LENGTH)
workflow = nvt.Workflow(filtered_sessions)
dataset = nvt.Dataset(df, cpu=False)
# Generating statistics for the features
workflow.fit(dataset)
# Applying the preprocessing and returning an NVTabular dataset
sessions_ds = workflow.transform(dataset)
# Converting the NVTabular dataset to a Dask cuDF dataframe (`to_ddf()`) and then to cuDF dataframe (`.compute()`)
sessions_gdf = sessions_ds.to_ddf().compute()
sessions_gdf.head(3)
```
It is possible to save the preprocessing workflow. That is useful to apply the same preprocessing to other data (with the same schema) and also to deploy the session-based recommendation pipeline to Triton Inference Server.
```
workflow.save('workflow_etl')
```
## Export pre-processed data by day
In this example we are going to split the preprocessed parquet files by days, to allow for temporal training and evaluation. There will be a folder for each day and three parquet files within each day folder: train.parquet, validation.parquet and test.parquet
```
OUTPUT_FOLDER = os.environ.get("OUTPUT_FOLDER",os.path.join(INPUT_DATA_DIR, "sessions_by_day"))
!mkdir -p $OUTPUT_FOLDER
from transformers4rec.data.preprocessing import save_time_based_splits
save_time_based_splits(data=nvt.Dataset(sessions_gdf),
output_dir= OUTPUT_FOLDER,
partition_col='day-first',
timestamp_col='session_id',
)
```
## Checking the preprocessed outputs
```
TRAIN_PATHS = sorted(glob.glob(os.path.join(OUTPUT_FOLDER, "1", "train.parquet")))
gdf = cudf.read_parquet(TRAIN_PATHS[0])
gdf.head()
```
You have just created session-level features to train a session-based recommendation model using NVTabular. Now you can move to the the next notebook,`02-session-based-XLNet-with-PyT.ipynb` to train a session-based recommendation model using [XLNet](https://arxiv.org/abs/1906.08237), one of the state-of-the-art NLP model.
| github_jupyter |
# Pixelwise Segmentation
Use the `elf.segmentation` module for feature based instance segmentation from pixels.
Note that this example is educational and there are easier and better performing method for the image used here. These segmentation methods are very suitable for pixel embeddings learned with neural networks, e.g. with methods like [Semantic Instance Segmentation with a Discriminateive Loss Function](https://arxiv.org/abs/1708.02551).
## Image and Features
Load the relevant libraries. Then load an image from the skimage examples and compute per pixel features.
```
%gui qt5
import time
import numpy as np
# import napari for data visualisation
import napari
# import vigra to compute per pixel features
import vigra
# elf segmentation functionality we need for the problem setup
import elf.segmentation.features as feats
from elf.segmentation.utils import normalize_input
# we use the coins example image
from skimage.data import coins
image = coins()
# We use blurring and texture filters from vigra.filters computed for different scales to obain pixel features.
# Note that it's certainly possible to compute better features for the segmentation problem at hand.
# But for our purposes, these features are good enough.
im_normalized = normalize_input(image)
scales = [4., 8., 12.]
image_features = [im_normalized[None]] # use the normal image as
for scale in scales:
image_features.append(normalize_input(vigra.filters.gaussianSmoothing(im_normalized, scale))[None])
feats1 = vigra.filters.hessianOfGaussianEigenvalues(im_normalized, scale)
image_features.append(normalize_input(feats1[..., 0])[None])
image_features.append(normalize_input(feats1[..., 1])[None])
feats2 = vigra.filters.structureTensorEigenvalues(im_normalized, scale, 1.5 * scale)
image_features.append(normalize_input(feats2[..., 0])[None])
image_features.append(normalize_input(feats2[..., 1])[None])
image_features = np.concatenate(image_features, axis=0)
print("Feature shape:")
print(image_features.shape)
# visualize the image and the features with napari
viewer = napari.Viewer()
viewer.add_image(im_normalized)
viewer.add_image(image_features)
```
## Segmentation Problem
Set up a graph segmentation problem based on the image and features with elf functionality.
To this end, we construct a grid graph and compute edge features for the inter pixel edges in this graph.
```
# compute a grid graph for the image
shape = image.shape
grid_graph = feats.compute_grid_graph(shape)
# compute the edge features
# elf supports three different distance metrics to compute edge features
# from the image features:
# - 'l1': the l1 distance
# - 'l2': the l2 distance
# - 'cosine': the cosine distance (= 1. - cosine similarity)
# here, we use the l2 distance
distance_type = 'l2'
# 'compute_grid-graph-image_features' returns both the edges (=list of node ids connected by the edge)
# and the edge weights. Here, the edges are the same as grid_graph.uvIds()
edges, edge_weights = feats.compute_grid_graph_image_features(grid_graph, image_features, distance_type)
# we normalize the edge weigths to the range [0, 1]
edge_weights = normalize_input(edge_weights)
# simple post-processing to ensure the background label is '0',
# filter small segments with a size of below 100 pixels
# and ensure that the segmentation ids are consecutive
def postprocess_segmentation(seg, shape, min_size=100):
if seg.ndim == 1:
seg = seg.reshape(shape)
ids, sizes = np.unique(seg, return_counts=True)
bg_label = ids[np.argmax(sizes)]
if bg_label != 0:
if 0 in seg:
seg[seg == 0] = seg.max() + 1
seg[seg == bg_label] = 0
filter_ids = ids[sizes < min_size]
seg[np.isin(seg, filter_ids)] = 0
vigra.analysis.relabelConsecutive(seg, out=seg, start_label=1, keep_zeros=True)
return seg
```
## Multicut
As the first segmentation method, we use Multicut segmentation, based on the grid graph and the edge weights we have just computed.
```
# the elf multicut funtionality
import elf.segmentation.multicut as mc
# In order to apply multicut segmentation, we need to map the edge weights from their initial value range [0, 1]
# to [-inf, inf]; where positive values represent attractive edges and negative values represent repulsive edges.
# When computing these "costs" for the multicut, we can set the threshold for when an edge is counted
# as repulsive with the so called boundary bias, or beta, parameter.
# For values smaller than 0.5 the multicut segmentation will under-segment more,
# for values larger than 0.4 it will over-segment more.
beta = .75
costs = mc.compute_edge_costs(edge_weights, beta=beta)
print("Mapped edge weights in range", edge_weights.min(), edge_weights.max(), "to multicut costs in range", costs.min(), costs.max())
# compute the multicut segmentation
t = time.time()
mc_seg = mc.multicut_kernighan_lin(grid_graph, costs)
print("Computing the segmentation with multicut took", time.time() - t, "s")
mc_seg = postprocess_segmentation(mc_seg, shape)
# visualize the multicut segmentation
viewer = napari.Viewer()
viewer.add_image(image)
viewer.add_labels(mc_seg)
```
## Long-range Segmentation Problem
For now, we have only taken "local" information into account for the segmentation problem.
More specifically, we have only solved the Multicut with edges derived from nearest neighbor pixel transitions.
Next, we will use two algorithms, Mutex Watershed and Lifted Multicut, that can take long range edges into account. This has the advantage that feature differences are often more pronounced along larger distances, thus yielding much better information with respect to label transition.
Here, we extract this information by defining a "pixel offset pattern" and comparing the pixel features for these offsets. For details about this segmentation approach check out [The Mutex Watershed: Efficient, Parameter-Free Image Partitioning](https://openaccess.thecvf.com/content_ECCV_2018/html/Steffen_Wolf_The_Mutex_Watershed_ECCV_2018_paper.html).
```
# here, we define the following offset pattern:
# straight and diagonal transitions at a radius of 3, 9 and 27 pixels
# note that the offsets [-1, 0] and [0, -1] would correspond to the edges of the grid graph
offsets = [
[-3, 0], [0, -3], [-3, 3], [3, 3],
[-9, 0], [0, -9], [-9, 9], [9, 9],
[-27, 0], [0, -27], [-27, 27], [27, 27]
]
# we have significantly more long range than normal edges.
# hence, we subsample the offsets, for which actual long range edges will be computed by setting a stride factor
strides = [2, 2]
distance_type = 'l2' # we again use l2 distance
lr_edges, lr_edge_weights = feats.compute_grid_graph_image_features(grid_graph, image_features, distance_type,
offsets=offsets, strides=strides,
randomize_strides=False)
lr_edge_weights = normalize_input(lr_edge_weights)
print("Have computed", len(lr_edges), "long range edges, compared to", len(edges), "normal edges")
```
## Mutex Watershed
We use the Mutex Watershed to segment the image. This algorithm functions similar to (Lifted) Multicut, but is greedy and hence much faster. Despite its greedy nature, for many problems the solutions are of similar quality than Multicut segmentation.
```
# elf mutex watershed functionality
import elf.segmentation.mutex_watershed as mws
t = time.time()
mws_seg = mws.mutex_watershed_clustering(edges, lr_edges, edge_weights, lr_edge_weights)
print("Computing the segmentation with mutex watershed took", time.time() - t, "s")
mws_seg = postprocess_segmentation(mws_seg, shape)
viewer = napari.Viewer()
viewer.add_image(image)
viewer.add_labels(mws_seg)
```
## Lifted Multicut
Finally, we use Lifted Multicut segmentation. The Lifted Multicut is an extension to the Multicut, which can incorporate long range edges.
```
# elf lifted multicut functionality
import elf.segmentation.lifted_multicut as lmc
# For the lifted multicut, we again need to transform the edge weights in [0, 1] to costs in [-inf, inf]
beta = .75 # we again use a boundary bias of 0.75
lifted_costs = mc.compute_edge_costs(lr_edge_weights, beta=beta)
t = time.time()
lmc_seg = lmc.lifted_multicut_kernighan_lin(grid_graph, costs, lr_edges, lifted_costs)
print("Computing the segmentation with lifted multicut took", time.time() - t, "s")
lmc_seg = postprocess_segmentation(lmc_seg, shape)
viewer = napari.Viewer()
viewer.add_image(image)
viewer.add_labels(lmc_seg)
```
## Comparing the segmentations
We can now compare the three different segmentation. Note that the comparison is not quite fair here, because we have used the beta parameter to bias the segmentation to more over-segmentation for Multicut and Lifted Multicut while applying the Mutex Watershed to unbiased edge weights.
```
viewer = napari.Viewer()
viewer.add_image(image)
viewer.add_labels(mc_seg)
viewer.add_labels(mws_seg)
viewer.add_labels(lmc_seg)
```
| github_jupyter |
```
!jupyter-nbconvert --to python --template python_clean epd2in7.ipynb
%load_ext autoreload
%autoreload 2
# import epdconfig
# epdconfig = epdconfig.RaspberryPi()
import logging
try:
from . import epdconfig
except ImportError:
import epdconfig
# Constants
EPD_WIDTH = 176
EPD_HEIGHT = 264
# colors
GRAY1 = 0xff #white
GRAY2 = 0xC0
GRAY3 = 0x80 #gray
GRAY4 = 0x00 #Blackest
# Display resolution
class EPD:
def __init__(self):
self.reset_pin = epdconfig.RST_PIN
self.dc_pin = epdconfig.DC_PIN
self.busy_pin = epdconfig.BUSY_PIN
self.cs_pin = epdconfig.CS_PIN
self.width = EPD_WIDTH
self.height = EPD_HEIGHT
self.GRAY1 = GRAY1 #white
self.GRAY2 = GRAY2
self.GRAY3 = GRAY3 #gray
self.GRAY4 = GRAY4 #Blackest
# lookup tables
lut_vcom_dc = [0x00, 0x00,
0x00, 0x08, 0x00, 0x00, 0x00, 0x02,
0x60, 0x28, 0x28, 0x00, 0x00, 0x01,
0x00, 0x14, 0x00, 0x00, 0x00, 0x01,
0x00, 0x12, 0x12, 0x00, 0x00, 0x01,
0x00, 0x00, 0x00, 0x00, 0x00, 0x00,
0x00, 0x00, 0x00, 0x00, 0x00, 0x00,
0x00, 0x00, 0x00, 0x00, 0x00, 0x00
]
# white to white transition
lut_ww = [
0x40, 0x08, 0x00, 0x00, 0x00, 0x02,
0x90, 0x28, 0x28, 0x00, 0x00, 0x01,
0x40, 0x14, 0x00, 0x00, 0x00, 0x01,
0xA0, 0x12, 0x12, 0x00, 0x00, 0x01,
0x00, 0x00, 0x00, 0x00, 0x00, 0x00,
0x00, 0x00, 0x00, 0x00, 0x00, 0x00,
0x00, 0x00, 0x00, 0x00, 0x00, 0x00,
]
# black to white transition
lut_bw = [
0x40, 0x08, 0x00, 0x00, 0x00, 0x02,
0x90, 0x28, 0x28, 0x00, 0x00, 0x01,
0x40, 0x14, 0x00, 0x00, 0x00, 0x01,
0xA0, 0x12, 0x12, 0x00, 0x00, 0x01,
0x00, 0x00, 0x00, 0x00, 0x00, 0x00,
0x00, 0x00, 0x00, 0x00, 0x00, 0x00,
0x00, 0x00, 0x00, 0x00, 0x00, 0x00,
]
# black to black transition
lut_bb = [
0x80, 0x08, 0x00, 0x00, 0x00, 0x02,
0x90, 0x28, 0x28, 0x00, 0x00, 0x01,
0x80, 0x14, 0x00, 0x00, 0x00, 0x01,
0x50, 0x12, 0x12, 0x00, 0x00, 0x01,
0x00, 0x00, 0x00, 0x00, 0x00, 0x00,
0x00, 0x00, 0x00, 0x00, 0x00, 0x00,
0x00, 0x00, 0x00, 0x00, 0x00, 0x00,
]
# white to black transition
lut_wb = [
0x80, 0x08, 0x00, 0x00, 0x00, 0x02,
0x90, 0x28, 0x28, 0x00, 0x00, 0x01,
0x80, 0x14, 0x00, 0x00, 0x00, 0x01,
0x50, 0x12, 0x12, 0x00, 0x00, 0x01,
0x00, 0x00, 0x00, 0x00, 0x00, 0x00,
0x00, 0x00, 0x00, 0x00, 0x00, 0x00,
0x00, 0x00, 0x00, 0x00, 0x00, 0x00,
]
###################full screen update LUT######################
#0~3 gray
gray_lut_vcom = [
0x00, 0x00,
0x00, 0x0A, 0x00, 0x00, 0x00, 0x01,
0x60, 0x14, 0x14, 0x00, 0x00, 0x01,
0x00, 0x14, 0x00, 0x00, 0x00, 0x01,
0x00, 0x13, 0x0A, 0x01, 0x00, 0x01,
0x00, 0x00, 0x00, 0x00, 0x00, 0x00,
0x00, 0x00, 0x00, 0x00, 0x00, 0x00,
0x00, 0x00, 0x00, 0x00, 0x00, 0x00,
]
#R21
gray_lut_ww =[
0x40, 0x0A, 0x00, 0x00, 0x00, 0x01,
0x90, 0x14, 0x14, 0x00, 0x00, 0x01,
0x10, 0x14, 0x0A, 0x00, 0x00, 0x01,
0xA0, 0x13, 0x01, 0x00, 0x00, 0x01,
0x00, 0x00, 0x00, 0x00, 0x00, 0x00,
0x00, 0x00, 0x00, 0x00, 0x00, 0x00,
0x00, 0x00, 0x00, 0x00, 0x00, 0x00,
]
#R22H r
gray_lut_bw =[
0x40, 0x0A, 0x00, 0x00, 0x00, 0x01,
0x90, 0x14, 0x14, 0x00, 0x00, 0x01,
0x00, 0x14, 0x0A, 0x00, 0x00, 0x01,
0x99, 0x0C, 0x01, 0x03, 0x04, 0x01,
0x00, 0x00, 0x00, 0x00, 0x00, 0x00,
0x00, 0x00, 0x00, 0x00, 0x00, 0x00,
0x00, 0x00, 0x00, 0x00, 0x00, 0x00,
]
#R23H w
gray_lut_wb =[
0x40, 0x0A, 0x00, 0x00, 0x00, 0x01,
0x90, 0x14, 0x14, 0x00, 0x00, 0x01,
0x00, 0x14, 0x0A, 0x00, 0x00, 0x01,
0x99, 0x0B, 0x04, 0x04, 0x01, 0x01,
0x00, 0x00, 0x00, 0x00, 0x00, 0x00,
0x00, 0x00, 0x00, 0x00, 0x00, 0x00,
0x00, 0x00, 0x00, 0x00, 0x00, 0x00,
]
#R24H b
gray_lut_bb =[
0x80, 0x0A, 0x00, 0x00, 0x00, 0x01,
0x90, 0x14, 0x14, 0x00, 0x00, 0x01,
0x20, 0x14, 0x0A, 0x00, 0x00, 0x01,
0x50, 0x13, 0x01, 0x00, 0x00, 0x01,
0x00, 0x00, 0x00, 0x00, 0x00, 0x00,
0x00, 0x00, 0x00, 0x00, 0x00, 0x00,
0x00, 0x00, 0x00, 0x00, 0x00, 0x00,
]
# # Hardware reset
def reset(self):
epdconfig.digital_write(self.reset_pin, 1)
epdconfig.delay_ms(200)
epdconfig.digital_write(self.reset_pin, 0)
epdconfig.delay_ms(5)
epdconfig.digital_write(self.reset_pin, 1)
epdconfig.delay_ms(200)
def send_command(self, command):
epdconfig.digital_write(self.dc_pin, 0)
epdconfig.digital_write(self.cs_pin, 0)
epdconfig.spi_writebyte([command])
epdconfig.digital_write(self.cs_pin, 1)
def send_data(self, data):
epdconfig.digital_write(self.dc_pin, 1)
epdconfig.digital_write(self.cs_pin, 0)
epdconfig.spi_writebyte([data])
epdconfig.digital_write(self.cs_pin, 1)
#### REVIEW ####
# consider making a break/raise after N loops of attempting to release busy status
def ReadBusy(self):
logging.debug("e-Paper busy")
while(epdconfig.digital_read(self.busy_pin) == 0): # 0: idle, 1: busy
epdconfig.delay_ms(200)
logging.debug("e-Paper busy release")
def set_lut(self):
self.send_command(0x20) # vcom
for count in range(0, 44):
self.send_data(self.lut_vcom_dc[count])
self.send_command(0x21) # ww --
for count in range(0, 42):
self.send_data(self.lut_ww[count])
self.send_command(0x22) # bw r
for count in range(0, 42):
self.send_data(self.lut_bw[count])
self.send_command(0x23) # wb w
for count in range(0, 42):
self.send_data(self.lut_bb[count])
self.send_command(0x24) # bb b
for count in range(0, 42):
self.send_data(self.lut_wb[count])
# def set_lut_gray(self):
def gray_SetLut(self):
self.send_command(0x20)
for count in range(0, 44): #vcom
self.send_data(self.gray_lut_vcom[count])
self.send_command(0x21) #red not use
for count in range(0, 42):
self.send_data(self.gray_lut_ww[count])
self.send_command(0x22) #bw r
for count in range(0, 42):
self.send_data(self.gray_lut_bw[count])
self.send_command(0x23) #wb w
for count in range(0, 42):
self.send_data(self.gray_lut_wb[count])
self.send_command(0x24) #bb b
for count in range(0, 42):
self.send_data(self.gray_lut_bb[count])
self.send_command(0x25) #vcom
for count in range(0, 42):
self.send_data(self.gray_lut_ww[count])
def init(self):
logging.info('init EPD screen (Black & White)')
if (epdconfig.module_init() != 0):
return -1
self.reset()
# see pp35: https://www.waveshare.com/w/upload/2/2d/2.7inch-e-paper-Specification.pdf
logging.debug('booster soft start')
self.send_command(0x06) # BOOSTER_SOFT_START see PP13-14
self.send_data(0x07) # PHA
self.send_data(0x07) # PHB
self.send_data(0x17) # PHC
# poorly documented -- unclear what this does
# see https://github.com/waveshare/e-Paper/issues/155
logging.debug('power optimization stages 1-4')
# Power optimization stage 1
self.send_command(0xF8)
self.send_data(0x60)
self.send_data(0xA5)
# Power optimization stage 2
self.send_command(0xF8)
self.send_data(0x89)
self.send_data(0xA5)
# Power optimization stage 3
self.send_command(0xF8)
self.send_data(0x90)
self.send_data(0x00)
# Power optimization stage 4
self.send_command(0xF8)
self.send_data(0x93)
self.send_data(0x2A)
logging.debug('reset DFV_EN')
self.send_command(0x16) # PARTIAL_DISPLAY_REFRESH
self.send_data(0x00)
logging.debug('power setting')
self.send_command(0x01) # POWER_SETTING
self.send_data(0x03) # VDS_EN, VDG_EN
self.send_data(0x00) # VCOM_HV, VGHL_LV[1], VGHL_LV[0]
self.send_data(0x2b) # VDH
self.send_data(0x2b) # VDL
# self.send_data(0x09) # VDHR -- set voltage for red pixel (uneeded in B&W)
logging.debug('power on')
self.send_command(0x04) # POWER_ON
self.ReadBusy()
logging.debug('pannel setting')
self.send_command(0x00) # PANEL_SETTING (see p11)
self.send_data(0xAF) # res 296x160, use LUT, USE B/W/R pixels, scan up, shift rt, booster on, booster off
logging.debug('PLL control frequency setting (100Hz)')
self.send_command(0x30) # PLL_CONTROL
self.send_data(0x3A) # 3A 100HZ 29 150Hz 39 200HZ 31 171HZ
logging.debug('resolution setting (176x264)')
self.send_command(0x61)
self.send_data(0x00) #176
self.send_data(0xb0)
self.send_data(0x01) #264
self.send_data(0x08)
# order of VCOM and VCM_DC is swapped in spec; must be swapped as shown below
logging.debug('Vcom and data interval setting')
self.send_command(0X50) #VCOM AND DATA INTERVAL SETTING
self.send_data(0x57) # 0x57 in original code
logging.debug('VCM_DC setting ')
self.send_command(0x82) # VCM_DC_SETTING_REGISTER
self.send_data(0x12)
logging.debug('set b&w lookup tables')
self.set_lut()
return 0
def Init_4Gray(self):
logging.info('init EPD screen (4Gray & White)')
if (epdconfig.module_init() != 0):
return -1
self.reset()
logging.debug('booster soft start')
self.send_command(0x06) # BOOSTER_SOFT_START see PP13-14
self.send_data(0x07) # PHA
self.send_data(0x07) # PHB
self.send_data(0x17) # PHC
# poorly documented -- unclear what this does
# see https://github.com/waveshare/e-Paper/issues/155
logging.debug('power optimization stages 1-4')
# Power optimization stage 1
self.send_command(0xF8)
self.send_data(0x60)
self.send_data(0xA5)
# Power optimization stage 2
self.send_command(0xF8)
self.send_data(0x89)
self.send_data(0xA5)
# Power optimization stage 3
self.send_command(0xF8)
self.send_data(0x90)
self.send_data(0x00)
# Power optimization stage 4
self.send_command(0xF8)
self.send_data(0x93)
self.send_data(0x2A)
logging.debug('reset DFV_EN')
self.send_command(0x16) # PARTIAL_DISPLAY_REFRESH
self.send_data(0x00)
logging.debug('power setting')
self.send_command(0x01) # POWER_SETTING
self.send_data(0x03) # VDS_EN, VDG_EN
self.send_data(0x00) # VCOM_HV, VGHL_LV[1], VGHL_LV[0]
self.send_data(0x2b) # VDH
self.send_data(0x2b) # VDL
logging.debug('power on')
self.send_command(0x04) # POWER_ON
self.ReadBusy()
logging.debug('pannel setting')
self.send_command(0x00) # PANEL_SETTING (see p13)
self.send_data(0xbf) # res 296z160, use LUT, use BW pixles LUT1, scan up, shift rt, booster on, booster off
logging.debug('PLL control frequency setting')
self.send_command(0x30) # PLL_CONTROL
self.send_data(0x90) # original from code 100Hz?
logging.debug('resolution setting (176x264)')
self.send_command(0x61)
self.send_data(0x00) #176
self.send_data(0xb0)
self.send_data(0x01) #264
self.send_data(0x08)
# order of VCOM and VCM_DC is swapped in spec; must be swapped as shown below
logging.debug('Vcom and data interval setting')
self.send_command(0X50) #VCOM AND DATA INTERVAL SETTING
self.send_data(0x57) # 0x57 in original code
logging.debug('VCM_DC setting ')
self.send_command(0x82) # VCM_DC_SETTING_REGISTER
self.send_data(0x12)
return 0
def getbuffer(self, image):
# logging.debug("bufsiz = ",int(self.width/8) * self.height)
buf = [0xFF] * (int(self.width/8) * self.height)
image_monocolor = image.convert('1')
imwidth, imheight = image_monocolor.size
pixels = image_monocolor.load()
# logging.debug("imwidth = %d, imheight = %d",imwidth,imheight)
if(imwidth == self.width and imheight == self.height):
logging.debug("Vertical")
for y in range(imheight):
for x in range(imwidth):
# Set the bits for the column of pixels at the current position.
if pixels[x, y] == 0:
buf[int((x + y * self.width) / 8)] &= ~(0x80 >> (x % 8))
elif(imwidth == self.height and imheight == self.width):
logging.debug("Horizontal")
for y in range(imheight):
for x in range(imwidth):
newx = y
newy = self.height - x - 1
if pixels[x, y] == 0:
buf[int((newx + newy*self.width) / 8)] &= ~(0x80 >> (y % 8))
return buf
def getbuffer_4Gray(self, image):
# logging.debug("bufsiz = ",int(self.width/8) * self.height)
buf = [0xFF] * (int(self.width / 4) * self.height)
image_monocolor = image.convert('L')
imwidth, imheight = image_monocolor.size
pixels = image_monocolor.load()
i=0
# logging.debug("imwidth = %d, imheight = %d",imwidth,imheight)
if(imwidth == self.width and imheight == self.height):
logging.debug("Vertical")
for y in range(imheight):
for x in range(imwidth):
# Set the bits for the column of pixels at the current position.
if(pixels[x, y] == 0xC0):
pixels[x, y] = 0x80
elif (pixels[x, y] == 0x80):
pixels[x, y] = 0x40
i= i+1
if(i%4 == 0):
buf[int((x + (y * self.width))/4)] = ((pixels[x-3, y]&0xc0) | (pixels[x-2, y]&0xc0)>>2 | (pixels[x-1, y]&0xc0)>>4 | (pixels[x, y]&0xc0)>>6)
elif(imwidth == self.height and imheight == self.width):
logging.debug("Horizontal")
for x in range(imwidth):
for y in range(imheight):
newx = y
newy = x
if(pixels[x, y] == 0xC0):
pixels[x, y] = 0x80
elif (pixels[x, y] == 0x80):
pixels[x, y] = 0x40
i= i+1
if(i%4 == 0):
buf[int((newx + (newy * self.width))/4)] = ((pixels[x, y-3]&0xc0) | (pixels[x, y-2]&0xc0)>>2 | (pixels[x, y-1]&0xc0)>>4 | (pixels[x, y]&0xc0)>>6)
return buf
def display(self, image):
# load image into buffer
self.send_command(0x10) # start transmission 1
for i in range(0, int(self.width * self.height / 8)):
self.send_data(0xFF)
self.send_command(0x13) # start transmisison 2
for i in range(0, int(self.width * self.height / 8)):
self.send_data(image[i])
# write buffer to display
self.send_command(0x12) # display refresh
self.ReadBusy()
def display_4Gray(self, image):
self.send_command(0x10)
for i in range(0, 5808): #5808*4 46464
temp3=0
for j in range(0, 2):
temp1 = image[i*2+j]
for k in range(0, 2):
temp2 = temp1&0xC0
if(temp2 == 0xC0):
temp3 |= 0x01#white
elif(temp2 == 0x00):
temp3 |= 0x00 #black
elif(temp2 == 0x80):
temp3 |= 0x01 #gray1
else: #0x40
temp3 |= 0x00 #gray2
temp3 <<= 1
temp1 <<= 2
temp2 = temp1&0xC0
if(temp2 == 0xC0): #white
temp3 |= 0x01
elif(temp2 == 0x00): #black
temp3 |= 0x00
elif(temp2 == 0x80):
temp3 |= 0x01 #gray1
else : #0x40
temp3 |= 0x00 #gray2
if(j!=1 or k!=1):
temp3 <<= 1
temp1 <<= 2
self.send_data(temp3)
self.send_command(0x13)
for i in range(0, 5808): #5808*4 46464
temp3=0
for j in range(0, 2):
temp1 = image[i*2+j]
for k in range(0, 2):
temp2 = temp1&0xC0
if(temp2 == 0xC0):
temp3 |= 0x01#white
elif(temp2 == 0x00):
temp3 |= 0x00 #black
elif(temp2 == 0x80):
temp3 |= 0x00 #gray1
else: #0x40
temp3 |= 0x01 #gray2
temp3 <<= 1
temp1 <<= 2
temp2 = temp1&0xC0
if(temp2 == 0xC0): #white
temp3 |= 0x01
elif(temp2 == 0x00): #black
temp3 |= 0x00
elif(temp2 == 0x80):
temp3 |= 0x00 #gray1
else: #0x40
temp3 |= 0x01 #gray2
if(j!=1 or k!=1):
temp3 <<= 1
temp1 <<= 2
self.send_data(temp3)
self.gray_SetLut()
self.send_command(0x12)
epdconfig.delay_ms(200)
self.ReadBusy()
# pass
#### REVIEW ####
# color arg is not needed -- keep for consistency between color and non-color models?
def Clear(self, color):
# send blank (0xFF) into buffer
self.send_command(0x10) # start data transmission 1
for i in range(0, int(self.width * self.height / 8)):
self.send_data(0xFF)
# send blank (0xFF) into buffer
self.send_command(0x13) # start data transmission 2
for i in range(0, int(self.width * self.height / 8)):
self.send_data(0xFF)
# write buffer to display
self.send_command(0x12) # display refresh
self.ReadBusy()
def sleep(self):
logging.info('enter sleep')
self.send_command(0X50) # Vcom and data interval setting
self.send_data(0xf7)
logging.debug('power off')
self.send_command(0X02) # power off
logging.debug('enter deep sleep')
self.send_command(0X07) # deep sleep
self.send_data(0xA5) #deep sleep
epdconfig.delay_ms(2000)
epdconfig.module_exit()
### END OF FILE ###
from pathlib import Path
import time
from PIL import Image,ImageDraw,ImageFont
import logging
logging.root.setLevel('INFO')
font_file = str(Path('../../pic/Font.ttc').resolve())
pic = Path('../../pic/2in7.bmp')
picH = Path('../../pic/2in7_Scale.bmp')
epd = EPD()
epd.init()
# epd.Clear(0xFF)
epd.init()
font24 = ImageFont.truetype(font_file, 24)
font18 = ImageFont.truetype(font_file, 18)
font35 = ImageFont.truetype(font_file, 35)
# Drawing on the Horizontal image
logging.info("1.Drawing on the Horizontal image...")
Himage = Image.new('1', (epd.height, epd.width), 255) # 255: clear the frame
draw = ImageDraw.Draw(Himage)
draw.text((10, 0), 'hello world', font = font24, fill = 0)
draw.text((150, 0), u'微雪电子', font = font24, fill = 0)
draw.line((20, 50, 70, 100), fill = 0)
draw.line((70, 50, 20, 100), fill = 0)
draw.rectangle((20, 50, 70, 100), outline = 0)
draw.line((165, 50, 165, 100), fill = 0)
draw.line((140, 75, 190, 75), fill = 0)
draw.arc((140, 50, 190, 100), 0, 360, fill = 0)
draw.rectangle((80, 50, 130, 100), fill = 0)
draw.chord((200, 50, 250, 100), 0, 360, fill = 0)
epd.display(epd.getbuffer(Himage))
time.sleep(2)
Himage = Image.open(pic)
epd.display(epd.getbuffer(Himage))
'''4Gray display'''
logging.info("4Gray display--------------------------------")
epd.Init_4Gray()
Limage = Image.new('L', (epd.width, epd.height), 0) # 255: clear the frame
draw = ImageDraw.Draw(Limage)
draw.text((20, 0), u'微雪电子', font = font35, fill = epd.GRAY1)
draw.text((20, 35), u'微雪电子', font = font35, fill = epd.GRAY2)
draw.text((20, 70), u'微雪电子', font = font35, fill = epd.GRAY3)
draw.text((40, 110), 'hello world', font = font18, fill = epd.GRAY1)
draw.line((10, 140, 60, 190), fill = epd.GRAY1)
draw.line((60, 140, 10, 190), fill = epd.GRAY1)
draw.rectangle((10, 140, 60, 190), outline = epd.GRAY1)
draw.line((95, 140, 95, 190), fill = epd.GRAY1)
draw.line((70, 165, 120, 165), fill = epd.GRAY1)
draw.arc((70, 140, 120, 190), 0, 360, fill = epd.GRAY1)
draw.rectangle((10, 200, 60, 250), fill = epd.GRAY1)
draw.chord((70, 200, 120, 250), 0, 360, fill = epd.GRAY1)
epd.display_4Gray(epd.getbuffer_4Gray(Limage))
time.sleep(2)
#display 4Gra bmp
Himage = Image.open(picH)
epd.display_4Gray(epd.getbuffer_4Gray(Himage))
time.sleep(2)
logging.info("Clear...")
epd.Clear(0xFF)
logging.info("Goto Sleep...")
epd.sleep()
```
| github_jupyter |
# Chapter 8: ニューラルネット
第6章で取り組んだニュース記事のカテゴリ分類を題材として,ニューラルネットワークでカテゴリ分類モデルを実装する.なお,この章ではPyTorch, TensorFlow, Chainerなどの機械学習プラットフォームを活用せよ.
## 70. 単語ベクトルの和による特徴量
問題50で構築した学習データ,検証データ,評価データを行列・ベクトルに変換したい.例えば,学習データについて,すべての事例$x_i$の特徴ベクトル$\boldsymbol{x}_i$を並べた行列$X$と,正解ラベルを並べた行列(ベクトル)$Y$を作成したい.
$$
X = \begin{pmatrix}
\boldsymbol{x}_1 \\
\boldsymbol{x}_2 \\
\dots \\
\boldsymbol{x}_n \\
\end{pmatrix} \in \mathbb{R}^{n \times d},
Y = \begin{pmatrix}
y_1 \\
y_2 \\
\dots \\
y_n \\
\end{pmatrix} \in \mathbb{N}^{n}
$$
ここで,$n$は学習データの事例数であり,$\boldsymbol{x}_i \in \mathbb{R}^d$と$y_i \in \mathbb{N}$はそれぞれ,$i \in \{1, \dots, n\}$番目の事例の特徴量ベクトルと正解ラベルを表す.
なお,今回は「ビジネス」「科学技術」「エンターテイメント」「健康」の4カテゴリ分類である.$\mathbb{N}_{<4}$で$4$未満の自然数($0$を含む)を表すことにすれば,任意の事例の正解ラベル$y_i$は$y_i \in \mathbb{N}_{<4}$で表現できる.
以降では,ラベルの種類数を$L$で表す(今回の分類タスクでは$L=4$である).
$i$番目の事例の特徴ベクトル$\boldsymbol{x}_i$は,次式で求める.
$$
\boldsymbol{x}_i = \frac{1}{T_i} \sum_{t=1}^{T_i} \mathrm{emb}(w_{i,t})
$$
ここで,$i$番目の事例は$T_i$個の(記事見出しの)単語列$(w_{i,1}, w_{i,2}, \dots, w_{i,T_i})$から構成され,$\mathrm{emb}(w) \in \mathbb{R}^d$は単語$w$に対応する単語ベクトル(次元数は$d$)である.すなわち,$i$番目の事例の記事見出しを,その見出しに含まれる単語のベクトルの平均で表現したものが$\boldsymbol{x}_i$である.今回は単語ベクトルとして,問題60でダウンロードしたものを用いればよい.$300$次元の単語ベクトルを用いたので,$d=300$である.
$i$番目の事例のラベル$y_i$は,次のように定義する.
$$
y_i = \begin{cases}
0 & (\mbox{記事}x_i\mbox{が「ビジネス」カテゴリの場合}) \\
1 & (\mbox{記事}x_i\mbox{が「科学技術」カテゴリの場合}) \\
2 & (\mbox{記事}x_i\mbox{が「エンターテイメント」カテゴリの場合}) \\
3 & (\mbox{記事}x_i\mbox{が「健康」カテゴリの場合}) \\
\end{cases}
$$
なお,カテゴリ名とラベルの番号が一対一で対応付いていれば,上式の通りの対応付けでなくてもよい.
以上の仕様に基づき,以下の行列・ベクトルを作成し,ファイルに保存せよ.
+ 学習データの特徴量行列: $X_{\rm train} \in \mathbb{R}^{N_t \times d}$
+ 学習データのラベルベクトル: $Y_{\rm train} \in \mathbb{N}^{N_t}$
+ 検証データの特徴量行列: $X_{\rm valid} \in \mathbb{R}^{N_v \times d}$
+ 検証データのラベルベクトル: $Y_{\rm valid} \in \mathbb{N}^{N_v}$
+ 評価データの特徴量行列: $X_{\rm test} \in \mathbb{R}^{N_e \times d}$
+ 評価データのラベルベクトル: $Y_{\rm test} \in \mathbb{N}^{N_e}$
なお,$N_t, N_v, N_e$はそれぞれ,学習データの事例数,検証データの事例数,評価データの事例数である.
```
import pandas as pd
import numpy as np
train_df = pd.read_table('../data/news_aggregator/train.txt')
valid_df = pd.read_table('../data/news_aggregator/valid.txt')
test_df = pd.read_table('../data/news_aggregator/test.txt')
categories = ['b', 't', 'e', 'm']
from gensim.models import KeyedVectors
model = KeyedVectors.load_word2vec_format('../data/google_news.bin.gz', binary=True)
# tokenize, make lowercase and make stem
import re
import string
def tokenize(x):
table = str.maketrans(string.punctuation, ' ' * len(string.punctuation))
x = x.translate(table).split()
x = [w.lower() for w in x]
return x
train_df['tokens'] = train_df['title'].map(tokenize)
valid_df['tokens'] = valid_df['title'].map(tokenize)
test_df['tokens'] = test_df['title'].map(tokenize)
def labelize(x):
return categories.index(x)
train_df['label'] = train_df['category'].map(labelize)
valid_df['label'] = valid_df['category'].map(labelize)
test_df['label'] = test_df['category'].map(labelize)
import torch
def vectorize(tokens):
vec = np.array([model[w] for w in tokens if w in model])
return torch.tensor(np.mean(vec, axis=0))
X_train = torch.stack([vectorize(tokens) for tokens in train_df['tokens']])
X_valid = torch.stack([vectorize(tokens) for tokens in valid_df['tokens']])
X_test = torch.stack([vectorize(tokens) for tokens in test_df['tokens']])
y_train = torch.tensor(train_df['label'])
y_valid = torch.tensor(valid_df['label'])
y_test = torch.tensor(test_df['label'])
torch.save(X_train, '../data/X_train.pt')
torch.save(X_valid, '../data/X_valid.pt')
torch.save(X_test, '../data/X_test.pt')
torch.save(y_train, '../data/y_train.pt')
torch.save(y_valid, '../data/y_valid.pt')
torch.save(y_test, '../data/y_test.pt')
```
## 71. 単層ニューラルネットワークによる予測
問題70で保存した行列を読み込み,学習データについて以下の計算を実行せよ.
$$
\hat{\boldsymbol{y}}_1 = {\rm softmax}(\boldsymbol{x}_1 W), \\
\hat{Y} = {\rm softmax}(X_{[1:4]} W)
$$
ただし,${\rm softmax}$はソフトマックス関数,$X_{[1:4]} \in \mathbb{R}^{4 \times d}$は特徴ベクトル$\boldsymbol{x}_1, \boldsymbol{x}_2, \boldsymbol{x}_3, \boldsymbol{x}_4$を縦に並べた行列である.
$$
X_{[1:4]} = \begin{pmatrix}
\boldsymbol{x}_1 \\
\boldsymbol{x}_2 \\
\boldsymbol{x}_3 \\
\boldsymbol{x}_4 \\
\end{pmatrix}
$$
行列$W \in \mathbb{R}^{d \times L}$は単層ニューラルネットワークの重み行列で,ここではランダムな値で初期化すればよい(問題73以降で学習して求める).なお,$\hat{\boldsymbol{y}}_1 \in \mathbb{R}^L$は未学習の行列$W$で事例$x_1$を分類したときに,各カテゴリに属する確率を表すベクトルである.
同様に,$\hat{Y} \in \mathbb{R}^{n \times L}$は,学習データの事例$x_1, x_2, x_3, x_4$について,各カテゴリに属する確率を行列として表現している.
```
X_train = torch.load('../data/X_train.pt')
X_valid = torch.load('../data/X_valid.pt')
X_test = torch.load('../data/X_test.pt')
y_train = torch.load('../data/y_train.pt')
y_valid = torch.load('../data/y_valid.pt')
y_test = torch.load('../data/y_test.pt')
from torch import nn
class SLPNet(nn.Module):
def __init__(self, input_size, output_size):
super().__init__()
self.fc = nn.Linear(input_size, output_size, bias=False)
nn.init.normal_(self.fc.weight, 0.0, 1.0)
def forward(self, x):
x = self.fc(x)
return x
model = SLPNet(300, 4)
y_hat_1 = torch.softmax(model(X_train[:1]), dim=-1)
y_hat_1
Y_hat = torch.softmax(model(X_train[:4]), dim=-1)
Y_hat
```
## 72. 損失と勾配の計算
学習データの事例$x_1$と事例集合$x_1, x_2, x_3, x_4$に対して,クロスエントロピー損失と,行列$W$に対する勾配を計算せよ.なお,ある事例$x_i$に対して損失は次式で計算される.
$$
l_i = - \log [\mbox{事例}x_i\mbox{が}y_i\mbox{に分類される確率}]
$$
ただし,事例集合に対するクロスエントロピー損失は,その集合に含まれる各事例の損失の平均とする.
```
criterion = nn.CrossEntropyLoss()
loss_1 = criterion(model(X_train[:1]), y_train[:1])
model.zero_grad()
loss_1.backward()
print(f'損失: {loss_1}')
print(f'勾配: {model.fc.weight.grad}')
loss = criterion(model(X_train[:4]), y_train[:4])
model.zero_grad()
loss.backward()
print(f'損失: {loss}')
print(f'勾配: {model.fc.weight.grad}')
```
## 73. 確率的勾配降下法による学習
確率的勾配降下法(SGD: Stochastic Gradient Descent)を用いて,行列Wを学習せよ.なお,学習は適当な基準で終了させればよい(例えば「100エポックで終了」など).
```
from torch.utils.data import Dataset, DataLoader
class NewsDataset(Dataset):
def __init__(self, X, y):
self.X = X
self.y = y
def __len__(self):
return len(self.y)
def __getitem__(self, idx):
return [self.X[idx], self.y[idx]]
dataset_train = NewsDataset(X_train, y_train)
dataset_valid = NewsDataset(X_valid, y_valid)
dataset_test = NewsDataset(X_test, y_test)
dataloader_train = DataLoader(dataset_train, batch_size=1, shuffle=True)
dataloader_valid = DataLoader(dataset_valid, batch_size=len(dataset_valid), shuffle=False)
dataloader_test = DataLoader(dataset_test, batch_size=len(dataset_test), shuffle=False)
model = SLPNet(300, 4)
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.SGD(model.parameters(), lr=1e-1)
num_epochs = 100
for epoch in range(num_epochs):
model.train()
loss_train = 0.0
for i, (inputs, labels) in enumerate(dataloader_train):
optimizer.zero_grad()
outputs = model(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
loss_train += loss.item()
loss_train = loss_train / i
model.eval()
with torch.no_grad():
inputs, labels = next(iter(dataloader_valid))
outputs = model(inputs)
loss_valid = criterion(outputs, labels)
print(f'epoch: {epoch + 1}, loss_train: {loss_train:.4f}, loss_valid: {loss_valid:.4f}')
```
## 74. 正解率の計測
問題73で求めた行列を用いて学習データおよび評価データの事例を分類したとき,その正解率をそれぞれ求めよ.
```
def calculate_accuracy(model, loader):
model.eval()
total = 0
correct = 0
with torch.no_grad():
for inputs, labels in loader:
inputs, labels = inputs.to(device), labels.to(device)
outputs = model(inputs)
pred = torch.argmax(outputs, dim=-1)
total += len(inputs)
correct += (pred == labels).sum().item()
return correct / total
acc_train = calculate_accuracy(model, dataloader_train)
acc_test = calculate_accuracy(model, dataloader_test)
print(f'train accuracy: {acc_train:.3f}')
print(f'test accuracy: {acc_test:.3f}')
```
## 75. 損失と正解率のプロット
問題73のコードを改変し,各エポックのパラメータ更新が完了するたびに,訓練データでの損失,正解率,検証データでの損失,正解率をグラフにプロットし,学習の進捗状況を確認できるようにせよ.
```
import matplotlib.pyplot as plt
from IPython.display import clear_output
%matplotlib inline
accuracy_res = {
'train': [],
'valid': []
}
loss_res = {
'train': [],
'valid': []
}
fig = plt.figure(figsize=(16, 6))
ax1 = fig.add_subplot(1, 2, 1)
ax2 = fig.add_subplot(1, 2, 2, sharey=ax1)
ax1.set_xlabel('Train')
ax2.set_xlabel('Valid')
num_epochs = 100
for epoch in tqdm(range(num_epochs)):
model.train()
loss_train = 0.0
for i, (inputs, labels) in enumerate(dataloader_train):
optimizer.zero_grad()
outputs = model(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
loss_train += loss.item()
loss_train = loss_train / i
model.eval()
with torch.no_grad():
inputs, labels = next(iter(dataloader_valid))
outputs = model(inputs)
loss_valid = criterion(outputs, labels)
train_accuracy = calculate_accuracy(model, dataloader_train)
valid_accuracy = calculate_accuracy(model, dataloader_valid)
accuracy_res['train'].append(train_accuracy)
accuracy_res['valid'].append(valid_accuracy)
loss_res['train'].append(loss_train)
loss_res['valid'].append(loss_valid)
clear_output(wait=True)
line1, = ax1.plot(range(1, epoch + 2), loss_res['train'], label='loss', color='royalblue')
line2, = ax1.plot(range(1, epoch + 2), accuracy_res['train'], label='train', color='darkorange')
line3, = ax2.plot(range(1, epoch + 2), loss_res['valid'], label='loss', color='royalblue')
line4, = ax2.plot(range(1, epoch + 2), accuracy_res['valid'], label='accuracy', color='darkorange')
display(fig)
line1.remove()
line2.remove()
line3.remove()
line4.remove()
plt.close(fig)
```
## 76. チェックポイント
問題75のコードを改変し,各エポックのパラメータ更新が完了するたびに,チェックポイント(学習途中のパラメータ(重み行列など)の値や最適化アルゴリズムの内部状態)をファイルに書き出せ.
```
accuracy_res = {
'train': [],
'valid': []
}
loss_res = {
'train': [],
'valid': []
}
fig = plt.figure(figsize=(16, 6))
ax1 = fig.add_subplot(1, 2, 1)
ax2 = fig.add_subplot(1, 2, 2, sharey=ax1)
ax1.set_xlabel('Train')
ax2.set_xlabel('Valid')
num_epochs = 5
for epoch in tqdm(range(num_epochs)):
model.train()
loss_train = 0.0
for i, (inputs, labels) in enumerate(dataloader_train):
optimizer.zero_grad()
outputs = model(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
loss_train += loss.item()
loss_train = loss_train / i
model.eval()
with torch.no_grad():
inputs, labels = next(iter(dataloader_valid))
outputs = model(inputs)
loss_valid = criterion(outputs, labels)
train_accuracy = calculate_accuracy(model, dataloader_train)
valid_accuracy = calculate_accuracy(model, dataloader_valid)
accuracy_res['train'].append(train_accuracy)
accuracy_res['valid'].append(valid_accuracy)
loss_res['train'].append(loss_train)
loss_res['valid'].append(loss_valid)
clear_output(wait=True)
line1, = ax1.plot(range(1, epoch + 2), loss_res['train'], label='loss', color='royalblue')
line2, = ax1.plot(range(1, epoch + 2), accuracy_res['train'], label='train', color='darkorange')
line3, = ax2.plot(range(1, epoch + 2), loss_res['valid'], label='loss', color='royalblue')
line4, = ax2.plot(range(1, epoch + 2), accuracy_res['valid'], label='accuracy', color='darkorange')
display(fig)
line1.remove()
line2.remove()
line3.remove()
line4.remove()
torch.save({
'epoch': 'epoch',
'model_state_dict': model.state_dict(),
'optimizer_state_dict': optimizer.state_dict()
}, f'../data/checkpoints/checkpoint_{epoch}.pt')
plt.close(fig)
```
## 77. ミニバッチ化
問題76のコードを改変し,$B$事例ごとに損失・勾配を計算し,行列$W$の値を更新せよ(ミニバッチ化).$B$の値を$1, 2, 4, 8, \dots$と変化させながら,1エポックの学習に要する時間を比較せよ.
```
import datetime
batch_sizes = [2 ** x for x in range(8)]
times = []
fig = plt.figure(figsize=(8, 6))
ax = fig.add_subplot(1, 1, 1)
ax.set_xlabel('Epoch')
ax.set_ylabel('Seconds/Epoch')
num_epochs = 5
for idx, batch_size in enumerate(batch_sizes):
dataloader_train = DataLoader(dataset_train, batch_size=batch_size, shuffle=True)
model = SLPNet(300, 4)
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.SGD(model.parameters(), lr=1e-1)
start_time = datetime.datetime.now()
for epoch in tqdm(range(num_epochs)):
model.train()
loss_train = 0.0
for i, (inputs, labels) in enumerate(dataloader_train):
optimizer.zero_grad()
outputs = model(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
loss_train += loss.item()
loss_train = loss_train / i
model.eval()
with torch.no_grad():
inputs, labels = next(iter(dataloader_valid))
outputs = model(inputs)
loss_valid = criterion(outputs, labels)
end_time = datetime.datetime.now()
average_time = (end_time - start_time) / num_epochs
times.append(average_time.seconds)
clear_output(wait=True)
try:
line.remove()
except:
pass
line, = ax.plot(batch_sizes[:idx+1], times, color='royalblue')
display(fig)
plt.close(fig)
```
## 78. GPU上での学習
問題77のコードを改変し,GPU上で学習を実行せよ.
```
device = torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')
dataloader_train = DataLoader(dataset_train, batch_size=5, shuffle=True)
dataloader_valid = DataLoader(dataset_valid, batch_size=len(dataset_valid), shuffle=False)
dataloader_test = DataLoader(dataset_test, batch_size=len(dataset_test), shuffle=False)
model = SLPNet(300, 4).to(device)
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.SGD(model.parameters(), lr=1e-1)
accuracy_res = {
'train': [],
'valid': []
}
loss_res = {
'train': [],
'valid': []
}
fig = plt.figure(figsize=(16, 6))
ax1 = fig.add_subplot(1, 2, 1)
ax2 = fig.add_subplot(1, 2, 2, sharey=ax1)
ax1.set_xlabel('Train')
ax2.set_xlabel('Valid')
num_epochs = 100
for epoch in tqdm(range(num_epochs)):
model.train()
loss_train = 0.0
for i, (inputs, labels) in enumerate(dataloader_train):
optimizer.zero_grad()
inputs, labels = inputs.to(device), labels.to(device)
outputs = model(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
loss_train += loss.item()
loss_train = loss_train / i
model.eval()
with torch.no_grad():
inputs, labels = next(iter(dataloader_valid))
inputs, labels = inputs.to(device), labels.to(device)
outputs = model(inputs)
loss_valid = criterion(outputs, labels)
train_accuracy = calculate_accuracy(model, dataloader_train)
valid_accuracy = calculate_accuracy(model, dataloader_valid)
accuracy_res['train'].append(train_accuracy)
accuracy_res['valid'].append(valid_accuracy)
loss_res['train'].append(loss_train)
loss_res['valid'].append(loss_valid)
clear_output(wait=True)
line1, = ax1.plot(range(1, epoch + 2), loss_res['train'], label='loss', color='royalblue')
line2, = ax1.plot(range(1, epoch + 2), accuracy_res['train'], label='train', color='darkorange')
line3, = ax2.plot(range(1, epoch + 2), loss_res['valid'], label='loss', color='royalblue')
line4, = ax2.plot(range(1, epoch + 2), accuracy_res['valid'], label='accuracy', color='darkorange')
display(fig)
line1.remove()
line2.remove()
line3.remove()
line4.remove()
plt.close(fig)
```
## 79. 多層ニューラルネットワーク
問題78のコードを改変し,バイアス項の導入や多層化など,ニューラルネットワークの形状を変更しながら,高性能なカテゴリ分類器を構築せよ.
```
from torch import nn
class MLPNet(nn.Module):
def __init__(self, input_size, output_size):
super().__init__()
self.fc1 = nn.Linear(input_size, 512, bias=True)
self.fc2 = nn.Linear(512, 128, bias=True)
self.fc3 = nn.Linear(128, output_size, bias=True)
self.relu = nn.ReLU()
self.dropout = nn.Dropout(0.2)
nn.init.kaiming_normal_(self.fc1.weight)
nn.init.kaiming_normal_(self.fc2.weight)
nn.init.kaiming_normal_(self.fc3.weight)
def forward(self, x):
x = self.fc1(x)
x = self.relu(x)
x = self.fc2(x)
x = self.relu(x)
x = self.dropout(x)
x = self.fc3(x)
return x
dataloader_train = DataLoader(dataset_train, batch_size=20, shuffle=True)
dataloader_valid = DataLoader(dataset_valid, batch_size=len(dataset_valid), shuffle=False)
dataloader_test = DataLoader(dataset_test, batch_size=len(dataset_test), shuffle=False)
model = MLPNet(300, 4).to(device)
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.SGD(model.parameters(), lr=1e-1)
accuracy_res = {
'train': [],
'valid': []
}
loss_res = {
'train': [],
'valid': []
}
fig = plt.figure(figsize=(16, 6))
ax1 = fig.add_subplot(1, 2, 1)
ax2 = fig.add_subplot(1, 2, 2, sharey=ax1)
ax1.set_xlabel('Train')
ax2.set_xlabel('Valid')
num_epochs = 50
for epoch in tqdm(range(num_epochs)):
model.train()
loss_train = 0.0
for i, (inputs, labels) in enumerate(dataloader_train):
optimizer.zero_grad()
inputs, labels = inputs.to(device), labels.to(device)
outputs = model(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
loss_train += loss.item()
loss_train = loss_train / i
model.eval()
with torch.no_grad():
inputs, labels = next(iter(dataloader_valid))
inputs, labels = inputs.to(device), labels.to(device)
outputs = model(inputs)
loss_valid = criterion(outputs, labels)
train_accuracy = calculate_accuracy(model, dataloader_train)
valid_accuracy = calculate_accuracy(model, dataloader_valid)
accuracy_res['train'].append(train_accuracy)
accuracy_res['valid'].append(valid_accuracy)
loss_res['train'].append(loss_train)
loss_res['valid'].append(loss_valid)
clear_output(wait=True)
line1, = ax1.plot(range(1, epoch + 2), loss_res['train'], label='loss', color='royalblue')
line2, = ax1.plot(range(1, epoch + 2), accuracy_res['train'], label='train', color='darkorange')
line3, = ax2.plot(range(1, epoch + 2), loss_res['valid'], label='loss', color='royalblue')
line4, = ax2.plot(range(1, epoch + 2), accuracy_res['valid'], label='accuracy', color='darkorange')
display(fig)
line1.remove()
line2.remove()
line3.remove()
line4.remove()
plt.close(fig)
acc_train = calculate_accuracy(model, dataloader_train)
acc_test = calculate_accuracy(model, dataloader_test)
print(f'train accuracy: {acc_train:.3f}')
print(f'test accuracy: {acc_test:.3f}')
```
| github_jupyter |
# Vowpal Wabbit and LightGBM for a Regression Problem
This notebook shows how to build simple regression models by using
[Vowpal Wabbit (VW)](https://github.com/VowpalWabbit/vowpal_wabbit) and
[LightGBM](https://github.com/microsoft/LightGBM) with SynapseML.
We also compare the results with
[Spark MLlib Linear Regression](https://spark.apache.org/docs/latest/ml-classification-regression.html#linear-regression).
```
import os
if os.environ.get("AZURE_SERVICE", None) == "Microsoft.ProjectArcadia":
from pyspark.sql import SparkSession
spark = SparkSession.builder.getOrCreate()
import math
from synapse.ml.train import ComputeModelStatistics
from synapse.ml.vw import VowpalWabbitRegressor, VowpalWabbitFeaturizer
from synapse.ml.lightgbm import LightGBMRegressor
import numpy as np
import pandas as pd
from pyspark.ml.feature import VectorAssembler
from pyspark.ml.regression import LinearRegression
from sklearn.datasets import load_boston
```
## Prepare Dataset
We use [*Boston house price* dataset](https://scikit-learn.org/stable/modules/generated/sklearn.datasets.load_boston.html)
.
The data was collected in 1978 from Boston area and consists of 506 entries with 14 features including the value of homes.
We use `sklearn.datasets` module to download it easily, then split the set into training and testing by 75/25.
```
boston = load_boston()
feature_cols = ['f' + str(i) for i in range(boston.data.shape[1])]
header = ['target'] + feature_cols
df = spark.createDataFrame(pd.DataFrame(data=np.column_stack((boston.target, boston.data)), columns=header)).repartition(1)
print("Dataframe has {} rows".format(df.count()))
display(df.limit(10).toPandas())
train_data, test_data = df.randomSplit([0.75, 0.25], seed=42)
```
Following is the summary of the training set.
```
display(train_data.summary().toPandas())
```
Plot feature distributions over different target values (house prices in our case).
```
features = train_data.columns[1:]
values = train_data.drop('target').toPandas()
ncols = 5
nrows = math.ceil(len(features) / ncols)
```
## Baseline - Spark MLlib Linear Regressor
First, we set a baseline performance by using Linear Regressor in Spark MLlib.
```
featurizer = VectorAssembler(inputCols=feature_cols, outputCol='features')
lr_train_data = featurizer.transform(train_data)['target', 'features']
lr_test_data = featurizer.transform(test_data)['target', 'features']
display(lr_train_data.limit(10).toPandas())
# By default, `maxIter` is 100. Other params you may want to change include: `regParam`, `elasticNetParam`, etc.
lr = LinearRegression(labelCol='target')
lr_model = lr.fit(lr_train_data)
lr_predictions = lr_model.transform(lr_test_data)
display(lr_predictions.limit(10).toPandas())
```
We evaluate the prediction result by using `synapse.ml.train.ComputeModelStatistics` which returns four metrics:
* [MSE (Mean Squared Error)](https://en.wikipedia.org/wiki/Mean_squared_error)
* [RMSE (Root Mean Squared Error)](https://en.wikipedia.org/wiki/Root-mean-square_deviation) = sqrt(MSE)
* [R quared](https://en.wikipedia.org/wiki/Coefficient_of_determination)
* [MAE (Mean Absolute Error)](https://en.wikipedia.org/wiki/Mean_absolute_error)
```
metrics = ComputeModelStatistics(
evaluationMetric='regression',
labelCol='target',
scoresCol='prediction').transform(lr_predictions)
results = metrics.toPandas()
results.insert(0, 'model', ['Spark MLlib - Linear Regression'])
display(results)
```
## Vowpal Wabbit
Perform VW-style feature hashing. Many types (numbers, string, bool, map of string to (number, string)) are supported.
```
vw_featurizer = VowpalWabbitFeaturizer(
inputCols=feature_cols,
outputCol='features')
vw_train_data = vw_featurizer.transform(train_data)['target', 'features']
vw_test_data = vw_featurizer.transform(test_data)['target', 'features']
display(vw_train_data.limit(10).toPandas())
```
See [VW wiki](https://github.com/vowpalWabbit/vowpal_wabbit/wiki/Command-Line-Arguments) for command line arguments.
```
# Use the same number of iterations as Spark MLlib's Linear Regression (=100)
args = "--holdout_off --loss_function quantile -l 7 -q :: --power_t 0.3"
vwr = VowpalWabbitRegressor(
labelCol='target',
passThroughArgs=args,
numPasses=100)
# To reduce number of partitions (which will effect performance), use `vw_train_data.repartition(1)`
vw_train_data_2 = vw_train_data.repartition(1).cache()
print(vw_train_data_2.count())
vw_model = vwr.fit(vw_train_data_2.repartition(1))
vw_predictions = vw_model.transform(vw_test_data)
display(vw_predictions.limit(10).toPandas())
metrics = ComputeModelStatistics(
evaluationMetric='regression',
labelCol='target',
scoresCol='prediction').transform(vw_predictions)
vw_result = metrics.toPandas()
vw_result.insert(0, 'model', ['Vowpal Wabbit'])
results = results.append(
vw_result,
ignore_index=True)
display(results)
```
## LightGBM
```
lgr = LightGBMRegressor(
objective='quantile',
alpha=0.2,
learningRate=0.3,
numLeaves=31,
labelCol='target',
numIterations=100)
# Using one partition since the training dataset is very small
repartitioned_data = lr_train_data.repartition(1).cache()
print(repartitioned_data.count())
lg_model = lgr.fit(repartitioned_data)
lg_predictions = lg_model.transform(lr_test_data)
display(lg_predictions.limit(10).toPandas())
metrics = ComputeModelStatistics(
evaluationMetric='regression',
labelCol='target',
scoresCol='prediction').transform(lg_predictions)
lg_result = metrics.toPandas()
lg_result.insert(0, 'model', ['LightGBM'])
results = results.append(
lg_result,
ignore_index=True)
display(results)
```
Following figure shows the actual-vs.-prediction graphs of the results:
<img width="1102" alt="lr-vw-lg" src="https://user-images.githubusercontent.com/42475935/64071975-4c3e9600-cc54-11e9-8b1f-9a1ee300f445.png" />
```
if os.environ.get("AZURE_SERVICE", None) != "Microsoft.ProjectArcadia":
from matplotlib.colors import ListedColormap, Normalize
from matplotlib.cm import get_cmap
import matplotlib.pyplot as plt
f, axes = plt.subplots(nrows, ncols, sharey=True, figsize=(30,10))
f.tight_layout()
yy = [r['target'] for r in train_data.select('target').collect()]
for irow in range(nrows):
axes[irow][0].set_ylabel('target')
for icol in range(ncols):
try:
feat = features[irow*ncols + icol]
xx = values[feat]
axes[irow][icol].scatter(xx, yy, s=10, alpha=0.25)
axes[irow][icol].set_xlabel(feat)
axes[irow][icol].get_yaxis().set_ticks([])
except IndexError:
f.delaxes(axes[irow][icol])
cmap = get_cmap('YlOrRd')
target = np.array(test_data.select('target').collect()).flatten()
model_preds = [
("Spark MLlib Linear Regression", lr_predictions),
("Vowpal Wabbit", vw_predictions),
("LightGBM", lg_predictions)]
f, axes = plt.subplots(1, len(model_preds), sharey=True, figsize=(18, 6))
f.tight_layout()
for i, (model_name, preds) in enumerate(model_preds):
preds = np.array(preds.select('prediction').collect()).flatten()
err = np.absolute(preds - target)
norm = Normalize()
clrs = cmap(np.asarray(norm(err)))[:, :-1]
axes[i].scatter(preds, target, s=60, c=clrs, edgecolors='#888888', alpha=0.75)
axes[i].plot((0, 60), (0, 60), linestyle='--', color='#888888')
axes[i].set_xlabel('Predicted values')
if i ==0:
axes[i].set_ylabel('Actual values')
axes[i].set_title(model_name)
```
| github_jupyter |
```
# https://www.lucypark.kr/courses/2015-dm/text-mining.html
import nltk
import konlp
import gensim
import twython
nltk.download('gutenberg')
nltk.download('maxent_treebank_pos_tagger')
#사용예시: "Samsung (삼성)" 관련 트윗 받기
from twython import Twython
import settings as s # Create a file named settings.py, and put oauth KEY values inside
twitter = Twython(s.APP_KEY, s.APP_SECRET, s.OAUTH_TOKEN, s.OAUTH_TOKEN_SECRET)
tweets = twitter.search(q='삼성', count=100)
data = [(t['user']['screen_name'], t['text'], t['created_at']) for t in tweets['statuses']]
```
1. Read document
이 튜토리얼에서는 NLTK, KoNLPy에서 제공하는 문서들을 사용한다.
영어: Jane Austen의 소설 Emma
한국어: 대한민국 국회 제 1809890호 의안
할 수 있는 사람은, 위의 문서 대신 다른 텍스트 데이터를 로딩하여 사용해보자.
```
from nltk.corpus import gutenberg # Docs from project gutenberg.org
files_en = gutenberg.fileids() # Get file ids
doc_en = gutenberg.open('austen-emma.txt').read()
from konlpy.corpus import kobill # Docs from pokr.kr/bill
files_ko = kobill.fileids() # Get file ids
doc_ko = kobill.open('1809890.txt').read()
```
2. Tokenize
문서를 토큰으로 나누는 방법은 다양하다. 여기서는 영어에는 nltk.regexp_tokenize, 한국어에는 konlpy.tag.Twitter.morph를 사용해보자.
English
```
from nltk import regexp_tokenize
pattern = r'''(?x) ([A-Z]\.)+ | \w+(-\w+)* | \$?\d+(\.\d+)?%? | \.\.\. | [][.,;"'?():-_`]'''
tokens_en = regexp_tokenize(doc_en, pattern)
from konlpy.tag import Twitter; t = Twitter()
tokens_ko = t.morphs(doc_ko)
```
3. Load tokens with nltk.Text()
nltk.Text()는 문서 하나를 편리하게 탐색할 수 있는 다양한 기능을 제공한다.
```
import nltk
en = nltk.Text(tokens_en)
# Korean (For Python 2, name has to be input as u'유니코드'. If you are using Python 2, use u'유니코드' for input of all following Korean text.)
import nltk
ko = nltk.Text(tokens_ko, name='대한민국 국회 의안 제 1809890호') # For Python 2, input `name` as u'유니코드'
```
지금부터 nltk.Text()가 제공하는 다양한 기능을 하나씩 살펴보자. (참고링크: class nltk.text.Text API 문서)
```
# 1. Tokens
# English
print(len(en.tokens)) # returns number of tokens (document length)
print(len(set(en.tokens))) # returns number of unique tokens
en.vocab()
```
... 건너뛰고
Tagging and chunking
Until now, we used delimited text, namely tokens, to explore our sample document. Now let's classify words into given classes, namely part-of-speech tags, and chunk text into larger pieces.
1. POS tagging
There are numerous ways of tagging a text. Among them, the most frequently used, and developed way of tagging is arguably POS tagging.
Since one document is too long to observe a parsed structure, lets use one short sentence for each language.
```
# English
tokens = "The little yellow dog barked at the Persian cat".split()
tags_en = nltk.pos_tag(tokens)
# Korean
from konlpy.tag import Twitter; t = Twitter()
tags_ko = t.pos("작고 노란 강아지가 페르시안 고양이에게 짖었다")
# 2. Noun phrase chunking
nltk.RegexpParser() is a great way to start chunking.
# English
parser_en = nltk.RegexpParser("NP: {<DT>?<JJ>?<NN.*>*}")
chunks_en = parser_en.parse(tags_en)
chunks_en.draw()
# Korean
parser_ko = nltk.RegexpParser("NP: {<Adjective>*<Noun>*}")
chunks_ko = parser_ko.parse(tags_ko)
chunks_ko.draw()
```
| github_jupyter |
# CC3501 - Aux 7: Método de Diferencias Finitas
#### **Profesor: Daniel Calderón**
#### **Auxiliares: Diego Donoso y Pablo Pizarro**
#### **Ayudantes: Francisco Muñoz, Matías Rojas y Sebastián Contreras**
##### Fecha: 31/05/2019
---
#### Objetivos:
* Ejercitar el método de diferencias finitas en una aplicación práctica
* Aprender a escribir ecuaciones en un sistema lineal y a resolverlo con numpy
* Visualizar la solución utilizando matplotlib/mayavi
[Markdowns for Jupyter Notebooks](https://medium.com/ibm-data-science-experience/markdown-for-jupyter-notebooks-cheatsheet-386c05aeebed)
_Cuando te enteras de que puedes poner gifs en Notebooks_
```
from IPython.display import HTML
HTML('<center><img src="https://media.giphy.com/media/2xRWvsvjyrO2k/giphy.gif"></center>')
```
#### Problemas
1. Estudie el ejemplo ex_finite_differences.py. La figura ex_finite_differences.png es complementaria a dicha solución.
_Nota: Se usa comando %load filename para cargar archivo en notebook_
```
# %load ex_finite_differences_laplace_neumann.py
%matplotlib inline
"""
Daniel Calderon, CC3501, 2019-1
Finite Differences for Partial Differential Equations
Solving the Laplace equation in 2D with Dirichlet and
Neumann border conditions over a square domain.
"""
import numpy as np
import matplotlib.pyplot as mpl
# Problem setup
H = 4
W = 3
F = 2
h = 0.1
# Boundary Dirichlet Conditions:
TOP = 20
BOTTOM = 0
LEFT = 5
RIGHT = 15
# Number of unknowns
# left, bottom and top sides are known (Dirichlet condition)
# right side is unknown (Neumann condition)
nh = int(W / h)
nv = int(H / h) - 1
print(nh, nv)
# In this case, the domain is just a rectangle
N = nh * nv
# We define a function to convert the indices from i,j to k and viceversa
# i,j indexes the discrete domain in 2D.
# k parametrize those i,j, this way we can tidy the unknowns
# in a column vector and use the standard algebra
def getK(i,j):
return j * nh + i
def getIJ(k):
i = k % nh
j = k // nh
return (i, j)
"""
# This code is useful to debug the indexation functions above
print("="*10)
print(getK(0,0), getIJ(0))
print(getK(1,0), getIJ(1))
print(getK(0,1), getIJ(2))
print(getK(1,1), getIJ(3))
print("="*10)
import sys
sys.exit(0)
"""
# In this matrix we will write all the coefficients of the unknowns
A = np.zeros((N,N))
# In this vector we will write all the right side of the equations
b = np.zeros((N,))
# Note: To write an equation is equivalent to write a row in the matrix system
# We iterate over each point inside the domain
# Each point has an equation associated
# The equation is different depending on the point location inside the domain
for i in range(0, nh):
for j in range(0, nv):
# We will write the equation associated with row k
k = getK(i,j)
# We obtain indices of the other coefficients
k_up = getK(i, j+1)
k_down = getK(i, j-1)
k_left = getK(i-1, j)
k_right = getK(i+1, j)
# Depending on the location of the point, the equation is different
# Interior
if 1 <= i and i <= nh - 2 and 1 <= j and j <= nv - 2:
A[k, k_up] = 1
A[k, k_down] = 1
A[k, k_left] = 1
A[k, k_right] = 1
A[k, k] = -4
b[k] = 0
# left side
elif i == 0 and 1 <= j and j <= nv - 2:
A[k, k_up] = 1
A[k, k_down] = 1
A[k, k_right] = 1
A[k, k] = -4
b[k] = -LEFT
# right side
elif i == nh - 1 and 1 <= j and j <= nv - 2:
A[k, k_up] = 1
A[k, k_down] = 1
A[k, k_left] = 2
A[k, k] = -4
b[k] = -2 * h * F
# bottom side
elif 1 <= i and i <= nh - 2 and j == 0:
A[k, k_up] = 1
A[k, k_left] = 1
A[k, k_right] = 1
A[k, k] = -4
b[k] = -BOTTOM
# top side
elif 1 <= i and i <= nh - 2 and j == nv - 1:
A[k, k_down] = 1
A[k, k_left] = 1
A[k, k_right] = 1
A[k, k] = -4
b[k] = -TOP
# corner lower left
elif (i, j) == (0, 0):
A[k, k] = 1
b[k] = (BOTTOM + LEFT) / 2
# corner lower right
elif (i, j) == (nh - 1, 0):
A[k, k] = 1
b[k] = BOTTOM
# corner upper left
elif (i, j) == (0, nv - 1):
A[k, k] = 1
b[k] = (TOP + LEFT) / 2
# corner upper right
elif (i, j) == (nh - 1, nv - 1):
A[k, k] = 1
b[k] = TOP
else:
print("Point (" + str(i) + ", " + str(j) + ") missed!")
print("Associated point index is " + str(k))
raise Exception()
# A quick view of a sparse matrix
#mpl.spy(A)
# Solving our system
x = np.linalg.solve(A, b)
# Now we return our solution to the 2d discrete domain
# In this matrix we will store the solution in the 2d domain
u = np.zeros((nh,nv))
for k in range(0, N):
i,j = getIJ(k)
u[i,j] = x[k]
# Adding the borders, as they have known values
ub = np.zeros((nh + 1, nv + 2))
ub[1:nh + 1, 1:nv + 1] = u[:,:]
# Dirichlet boundary condition
# top
ub[0:nh + 2, nv + 1] = TOP
# bottom
ub[0:nh + 2, 0] = BOTTOM
# left
ub[0, 1:nv + 1] = LEFT
# this visualization locates the (0,0) at the lower left corner
# given all the references used in this example.
fig, ax = mpl.subplots(1,1)
pcm = ax.pcolormesh(ub.T, cmap='RdBu_r')
fig.colorbar(pcm)
ax.set_xlabel('x')
ax.set_ylabel('y')
ax.set_title('Laplace equation solution.\n Neumann Condition at the right side.')
ax.set_aspect('equal', 'datalim')
# Note:
# imshow is also valid but it uses another coordinate system,
# a data transformation is required
#ax.imshow(ub.T)
mpl.show()
%run ex_finite_differences_laplace_neumann.py
from ex_finite_differences_laplace_neumann import *
print(getK(1, 2))
```
2. Añada gráficos para visualizar la solución como:
1. Superficie
2. Curvas de nivel
> Utilice distintas paletas de colores y rotule correctamente cada eje.
_Hint: Guía de Laplace-Dirichlet o revisar los links_
_[Link](https://matplotlib.org/3.1.0/gallery/mplot3d/surface3d.html)_
[Link](https://matplotlib.org/3.1.0/api/_as_gen/matplotlib.pyplot.contour.html)
```
from mpl_toolkits.mplot3d import Axes3D
import matplotlib.pyplot as plt
from matplotlib import cm
from matplotlib.ticker import LinearLocator, FormatStrFormatter
import numpy as np
fig = plt.figure()
ax = fig.gca(projection='3d')
# Make data.
X = np.arange(0, nh + 1, 1)
Y = np.arange(0, nv + 2, 1)
X, Y = np.meshgrid(X, Y)
Z = ub.T
# Plot the surface.
surf = ax.plot_surface(X, Y, Z, cmap=cm.coolwarm,
linewidth=0, antialiased=False)
plt.contour(X, Y, Z)
plt.xlabel('x')
plt.ylabel('y')
```
3. Modifique el programa para que resuelva el sistema utilizando h = 0.5, 0.1, 0.05. Adjunte gráficos de todas sus soluciones.
_Nota: Para espaciados más pequeños se hace necesario utilizar otra forma para guardar las matrices, de ahí sale sparse_
_Nota 2: Con h= 0.01 se cae (probablemente)_
```
def problem(h_p):
# Problem setup
H = 4
W = 3
F = 2
# Boundary Dirichlet Conditions:
TOP = 20
BOTTOM = 0
LEFT = 5
RIGHT = 15
# Number of unknowns
# left, bottom and top sides are known (Dirichlet condition)
# right side is unknown (Neumann condition)
nh_p = int(W / h_p) - 1
nv_p = int(H / h_p) - 1
print(nh_p, nv_p)
# In this case, the domain is just a rectangle
N_p = nh_p * nv_p
# We define a function to convert the indices from i,j to k and viceversa
# i,j indexes the discrete domain in 2D.
# k parametrize those i,j, this way we can tidy the unknowns
# in a column vector and use the standard algebra
def newgetK(i, j):
return j * nh_p + i
def newgetIJ(k):
i = k % nh_p
j = k // nh_p
return (i, j)
"""
# This code is useful to debug the indexation functions above
print("="*10)
print(getK(0,0), getIJ(0))
print(getK(1,0), getIJ(1))
print(getK(0,1), getIJ(2))
print(getK(1,1), getIJ(3))
print("="*10)
import sys
sys.exit(0)
"""
# In this matrix we will write all the coefficients of the unknowns
A_p = np.zeros((N_p, N_p))
# In this vector we will write all the right side of the equations
b_p = np.zeros((N_p,))
# Note: To write an equation is equivalent to write a row in the matrix system
# We iterate over each point inside the domain
# Each point has an equation associated
# The equation is different depending on the point location inside the domain
for i in range(0, nh_p):
for j in range(0, nv_p):
# We will write the equation associated with row k
k = newgetK(i, j)
# We obtain indices of the other coefficients
k_up = newgetK(i, j + 1)
k_down = newgetK(i, j - 1)
k_left = newgetK(i - 1, j)
k_right = newgetK(i + 1, j)
# Depending on the location of the point, the equation is different
# Interior
if 1 <= i and i <= nh_p - 2 and 1 <= j and j <= nv_p - 2:
A_p[k, k_up] = 1
A_p[k, k_down] = 1
A_p[k, k_left] = 1
A_p[k, k_right] = 1
A_p[k, k] = -4
b_p[k] = 0
# left side
elif i == 0 and 1 <= j and j <= nv_p - 2:
A_p[k, k_up] = 1
A_p[k, k_down] = 1
A_p[k, k_right] = 1
A_p[k, k] = -4
b_p[k] = -LEFT
# right side
elif i == nh_p - 1 and 1 <= j and j <= nv_p - 2:
A_p[k, k_up] = 1
A_p[k, k_down] = 1
A_p[k, k_left] = 2
A_p[k, k] = -4
b_p[k] = -2 * h_p * F
# bottom side
elif 1 <= i and i <= nh_p - 2 and j == 0:
A_p[k, k_up] = 1
A_p[k, k_left] = 1
A_p[k, k_right] = 1
A_p[k, k] = -4
b_p[k] = -BOTTOM
# top side
elif 1 <= i and i <= nh_p - 2 and j == nv_p - 1:
A_p[k, k_down] = 1
A_p[k, k_left] = 1
A_p[k, k_right] = 1
A_p[k, k] = -4
b_p[k] = -TOP
# corner lower left
elif (i, j) == (0, 0):
A_p[k, k] = 1
b_p[k] = (BOTTOM + LEFT) / 2
# corner lower right
elif (i, j) == (nh_p - 1, 0):
A_p[k, k] = 1
b_p[k] = BOTTOM
# corner upper left
elif (i, j) == (0, nv_p - 1):
A_p[k, k] = 1
b_p[k] = (TOP + LEFT) / 2
# corner upper right
elif (i, j) == (nh_p - 1, nv_p - 1):
A_p[k, k] = 1
b_p[k] = TOP
else:
print("Point (" + str(i) + ", " + str(j) + ") missed!")
print("Associated point index is " + str(k))
raise Exception()
# A quick view of a sparse matrix
# mpl.spy(A)
# Solving our system
x_p = np.linalg.solve(A_p, b_p)
# Now we return our solution to the 2d discrete domain
# In this matrix we will store the solution in the 2d domain
u_p = np.zeros((nh_p, nv_p))
for k in range(0, N_p):
i, j = newgetIJ(k)
u_p[i, j] = x_p[k]
# Adding the borders, as they have known values
ub_p = np.zeros((nh_p + 2, nv_p + 2))
ub_p[1:nh_p + 1, 1:nv_p + 1] = u_p[:, :]
# Dirichlet boundary condition
# top
ub_p[0:nh_p + 2, nv_p + 1] = TOP
# bottom
ub_p[0:nh_p + 2, 0] = BOTTOM
# left
ub_p[0, 1:nv_p + 1] = LEFT
# right
ub_p[nh_p + 1, 1:nv_p + 1] = RIGHT
# this visualization locates the (0,0) at the lower left corner
# given all the references used in this example.
return ub_p
res = []
hs = [0.5, 0.1, 0.05]
for hi in hs:
res.append(problem(hi))
```
4. Utilizando el módulo time de Python, registre el tiempo que tarda resolver el problema para los espaciados h indicados en el problema anterior. Genere un gráfico que relacione h con el tiempo que se tarda.
```
import time
times = []
for hi in hs:
start = time.time()
problem(hi)
end = time.time()
times.append(end - start)
print("Tiempos :", times)
plt.plot(hs, times)
```
5. Modifique el programa para que el problema modelado posea sólo condiciones Dirichlet:
1. Borde superior: 10
2. Borde inferior: 5
3. Borde derecho: 0
4. Borde izquierdo: $f(y)=\sin(\pi\cdot y/H)$
5. Y se mide desde la esquina inferior izquierda hacia arriba.
> Presente su solución utilizando h=0.1
_Para esta pregunta se deben reemplazar todas las condiciones de Dirichlet con las mencionadas en el enunciado, para E) en vez de usar la constante, se debe llamar a una función evaluada en la altura del punto_
6. Modifique el programa de ejemplo para que modele la ecuación de Poisson utilizando $f(x,y) = \cos(x)\cdot\sin(y)$
_Ahora las ecuaciones deben ser de la forma: (en el interior, para los bordes es similar)_
$$U_{i-1, j} + U_{i+1, j} + U_{i, j-1} + U_{i, j+1} - 4\cdot U_{i, j} = h^2 f_{i, j}$$
7. Volviendo al programa de ejemplo, modifíquelo para que los espaciados horizontal y vertical sean diferentes. Es decir, su programa debe utilizar un hx y un hy.
8. ¿Como cambian sus ecuaciones si el borde izquierdo también posee una condición de Neumann?. Implemente este caso, considerando que el borde izquierdo cumple con la siguiente condición de borde:
$$F(y) = \sin\left( 2 \pi \cdot \frac{x}{H}\right)$$
9. Modifique el programa de ejemplo para que el dominio representado sea una L con condiciones de borde exclusivamente Dirichlet. Para esto:
1. Será necesario calcular correctamente la cantidad de incógnitas.
2. Reservar memoria para la matriz de coeficientes A y el vector del lado derecho b
3. Generar una indexación del dominio. Esto es, asociar a cada i,j del interior del dominio un índice k. Esto se logra modificando convenientemente las funciones getIJ y getK.
4. Una forma simple de abordar este problema es ir construyendo una tabla a medida que se recorre el dominio. Esta tabla debe almacenar el valor de i y j en cada fila k. De esta forma, dado un k, encontramos los i y j asociados. Y por otro lado, una búsqueda simple le permitirá encontrar la k dados un i y j.
5. Para generar los gráficos 2D, necesitara asignar un valor a puntos que se encuentran fuera del dominio. Para evitar que matplotlib los grafique, puede utilizar el valor NaN (Not a Number).
##### Si usted realizó exitosamente todas las actividades anteriores: ¡Felicitaciones!, ¡es un experto en diferencias finitas!.
```
HTML('<center><img src="https://media.giphy.com/media/3o8doT9BL7dgtolp7O/giphy.gif"></center>')
```
Hints:
- matplotlib.pyplot.spy permite visualizar rápidamente el contenido de una matriz esparsa
| github_jupyter |
```
# Discretization example
# We will use the titanic dataset
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.tree import DecisionTreeClassifier
# Load the dataset. We will only load 4 features.
# The class is "Survived" (0 or 1)
data = pd.read_csv('./data/titanic.csv',usecols =['Name','Sex','Age','Fare','Survived'])
# Ignore rows with null values
data = data.dropna()
# Keep data where Age >= 1
data = data.loc[data['Age'] >= 1]
data.head()
# We want to discretize "Age"
# So let's build a decision tree using the Age to predict Survive
# because the decision tree takes into account the class label
# Create a model with max depth = 2
tree_model = DecisionTreeClassifier(max_depth=2)
# Fit only using Age data
tree_model.fit(data.Age.to_frame(), data.Survived)
# And use the Age variable to predict the class
data['Age_DT']=tree_model.predict_proba(data.Age.to_frame())[:,1]
data.head(10)
# The "Age_DT" column contains the probability of the data point belonging to the corresponding class
# Check the unique values of the Age_DT attribute
data.Age_DT.unique()
# We have 4 unique values of probabilities.
# A tree of depth 2, makes 2 splits, therefore generating 4 buckets.
# That is why we see 4 different probabilities in the output above.
# Check the number of samples per probabilistic bucket
data.groupby(['Age_DT'])['Survived'].count()
# Check the age limits for each bucket
# i.e. let's see the boundaries of each bucket
pd.concat( [data.groupby(['Age_DT'])['Age'].min(),
data.groupby(['Age_DT'])['Age'].max()], axis=1)
# Visualize the tree
from six import StringIO
from IPython.display import Image
from sklearn.tree import export_graphviz
import pydotplus
dot_data = StringIO()
export_graphviz(tree_model, out_file=dot_data,
filled=True, rounded=True,
special_characters=True)
graph = pydotplus.graph_from_dot_data(dot_data.getvalue())
Image(graph.create_png())
# We obtained 4 bins (the leaf nodes).
##########################################
##########################################
##########################################
##########################################
##########################################
##########################################
##########################################
# Let's see a classification example now
##########################################
##########################################
##########################################
##########################################
##########################################
##########################################
##########################################
# We will use the iris dataset.
# This is perhaps the best known database to be found in the pattern recognition literature.
# Classify the Iris plant based on the 4 features:
# sepal length in cm
# sepal width in cm
# petal length in cm
# petal width in cm
# To one of these classes:
# -- Iris Setosa
# -- Iris Versicolour
# -- Iris Virginica
from sklearn import datasets
import numpy as np
import matplotlib.pyplot as plt
import numpy as np
from sklearn.model_selection import train_test_split
from helper_funcs import plot_decision_regions
# Load dataset (this comes directly in sklearn.datasets)
# We will only load 2 features: petal length and petal width
iris = datasets.load_iris()
X = iris.data[:, [2, 3]]
y = iris.target
print(f'X shape: {X.shape}')
print('Class labels:', np.unique(y))
# Split the data to training and test sets
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=0)
# Fit decision tree
tree = DecisionTreeClassifier(criterion='entropy', max_depth=3, random_state=0)
tree.fit(X_train, y_train)
X_combined = np.vstack((X_train, X_test))
y_combined = np.hstack((y_train, y_test))
plot_decision_regions(X_combined, y_combined,
classifier=tree, test_idx=range(105, 150))
plt.xlabel('petal length [cm]')
plt.ylabel('petal width [cm]')
plt.legend(loc='upper left')
plt.tight_layout()
plt.show()
export_graphviz(tree,
out_file='tree.dot',
feature_names=['petal length', 'petal width'])
dot_data = export_graphviz(
tree,
out_file=None,
feature_names=['petal length', 'petal width'],
class_names=['setosa', 'versicolor', 'virginica'],
filled=True,
rounded=True)
graph = pydotplus.graph_from_dot_data(dot_data)
Image(graph.create_png())
# RULES EXTRACTION
from sklearn.tree import export_text
tree_rules = export_text(tree, feature_names=['petal length', 'petal width'])
print(tree_rules)
# 5 Rules:
# Just follow the paths:
# R1: IF petal width <= 0.75 THEN class=0 (setosa)
# R2: IF petal width > 0.75 AND petal length <= 4.95 AND petal width <= 1.65 THEN class = 1 (versicolor)
# R3: IF petal width > 0.75 AND petal length <= 4.95 AND petal width > 1.65 THEN class = 2 (virginica)
# R4: IF petal width > 0.75 AND petal length > 4.95 AND petal length <= 5.05 THEN class = 2 (virginica)
# R5: IF petal width > 0.75 AND petal length > 4.95 AND petal length > 5.05 THEN class = 2 (virginica)
```
| github_jupyter |
## test.ipynb: Test the training result and Evaluate model
```
# Import the necessary libraries
from sklearn.decomposition import PCA
import os
import scipy.io as sio
import numpy as np
from keras.models import load_model
from keras.utils import np_utils
from sklearn.metrics import classification_report, confusion_matrix
import itertools
import spectral
# Define the neccesary functions for later use
# load the Indian pines dataset which is the .mat format
def loadIndianPinesData():
data_path = os.path.join(os.getcwd(),'data')
data = sio.loadmat(os.path.join(data_path, 'Indian_pines.mat'))['indian_pines']
labels = sio.loadmat(os.path.join(data_path, 'Indian_pines_gt.mat'))['indian_pines_gt']
return data, labels
# load the Indian pines dataset which is HSI format
# refered from http://www.spectralpython.net/fileio.html
def loadHSIData():
data_path = os.path.join(os.getcwd(), 'HSI_data')
data = spectral.open_image(os.path.join(data_path, '92AV3C.lan')).load()
data = np.array(data).astype(np.int32)
labels = spectral.open_image(os.path.join(data_path, '92AV3GT.GIS')).load()
labels = np.array(labels).astype(np.uint8)
labels.shape = (145, 145)
return data, labels
# Get the model evaluation report,
# include classification report, confusion matrix, Test_Loss, Test_accuracy
target_names = ['Alfalfa', 'Corn-notill', 'Corn-mintill', 'Corn'
,'Grass-pasture', 'Grass-trees', 'Grass-pasture-mowed',
'Hay-windrowed', 'Oats', 'Soybean-notill', 'Soybean-mintill',
'Soybean-clean', 'Wheat', 'Woods', 'Buildings-Grass-Trees-Drives',
'Stone-Steel-Towers']
def reports(X_test,y_test):
Y_pred = model.predict(X_test)
y_pred = np.argmax(Y_pred, axis=1)
classification = classification_report(np.argmax(y_test, axis=1), y_pred, target_names=target_names)
confusion = confusion_matrix(np.argmax(y_test, axis=1), y_pred)
score = model.evaluate(X_test, y_test, batch_size=32)
Test_Loss = score[0]*100
Test_accuracy = score[1]*100
return classification, confusion, Test_Loss, Test_accuracy
# apply PCA preprocessing for data sets
def applyPCA(X, numComponents=75):
newX = np.reshape(X, (-1, X.shape[2]))
pca = PCA(n_components=numComponents, whiten=True)
newX = pca.fit_transform(newX)
newX = np.reshape(newX, (X.shape[0],X.shape[1], numComponents))
return newX, pca
def Patch(data,height_index,width_index):
#transpose_array = data.transpose((2,0,1))
#print transpose_array.shape
height_slice = slice(height_index, height_index+PATCH_SIZE)
width_slice = slice(width_index, width_index+PATCH_SIZE)
patch = data[height_slice, width_slice, :]
return patch
# Global Variables
windowSize = 5
numPCAcomponents = 30
testRatio = 0.50
# show current path
PATH = os.getcwd()
print (PATH)
# Read PreprocessedData from file
X_test = np.load("./predata/XtestWindowSize"
+ str(windowSize) + "PCA" + str(numPCAcomponents) + "testRatio" + str(testRatio) + ".npy")
y_test = np.load("./predata/ytestWindowSize"
+ str(windowSize) + "PCA" + str(numPCAcomponents) + "testRatio" + str(testRatio) + ".npy")
# X_test = np.load("./predata/XAllWindowSize"
# + str(windowSize) + "PCA" + str(numPCAcomponents) + "testRatio" + str(testRatio) + ".npy")
# y_test = np.load("./predata/yAllWindowSize"
# + str(windowSize) + "PCA" + str(numPCAcomponents) + "testRatio" + str(testRatio) + ".npy")
X_test = np.reshape(X_test, (X_test.shape[0], X_test.shape[3], X_test.shape[1], X_test.shape[2]))
y_test = np_utils.to_categorical(y_test)
# load the model architecture and weights
model = load_model('./model/HSI_model_epochs100.h5')
# calculate result, loss, accuray and confusion matrix
classification, confusion, Test_loss, Test_accuracy = reports(X_test,y_test)
classification = str(classification)
confusion_str = str(confusion)
# show result and save to file
print('Test loss {} (%)'.format(Test_loss))
print('Test accuracy {} (%)'.format(Test_accuracy))
print("classification result: ")
print('{}'.format(classification))
print("confusion matrix: ")
print('{}'.format(confusion_str))
file_name = './result/report' + "WindowSize" + str(windowSize) + "PCA" + str(numPCAcomponents) + "testRatio" + str(testRatio) +".txt"
with open(file_name, 'w') as x_file:
x_file.write('Test loss {} (%)'.format(Test_loss))
x_file.write('\n')
x_file.write('Test accuracy {} (%)'.format(Test_accuracy))
x_file.write('\n')
x_file.write('\n')
x_file.write(" classification result: \n")
x_file.write('{}'.format(classification))
x_file.write('\n')
x_file.write(" confusion matrix: \n")
x_file.write('{}'.format(confusion_str))
import matplotlib.pyplot as plt
%matplotlib inline
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.get_cmap("Blues")):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
Normalized = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
if normalize:
cm = Normalized
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
print(cm)
plt.imshow(Normalized, interpolation='nearest', cmap=cmap)
plt.colorbar()
plt.title(title)
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=90)
plt.yticks(tick_marks, classes)
fmt = '.4f' if normalize else 'd'
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
thresh = cm[i].max() / 2.
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
plt.figure(figsize=(10,10))
plot_confusion_matrix(confusion, classes=target_names, normalize=False,
title='Confusion matrix, without normalization')
plt.savefig("./result/confusion_matrix_without_normalization.svg")
plt.show()
plt.figure(figsize=(15,15))
plot_confusion_matrix(confusion, classes=target_names, normalize=True,
title='Normalized confusion matrix')
plt.savefig("./result/confusion_matrix_with_normalization.svg")
plt.show()
# load the original image
# X, y = loadIndianPinesData()
X, y = loadHSIData()
X, pca = applyPCA(X, numComponents=numPCAcomponents)
height = y.shape[0]
width = y.shape[1]
PATCH_SIZE = 5
numComponents = 30
# calculate the predicted image
outputs = np.zeros((height,width))
for i in range(height-PATCH_SIZE+1):
for j in range(width-PATCH_SIZE+1):
p = int(PATCH_SIZE/2)
# print(y[i+p][j+p])
# target = int(y[i+PATCH_SIZE/2, j+PATCH_SIZE/2])
target = y[i+p][j+p]
if target == 0 :
continue
else :
image_patch=Patch(X,i,j)
# print (image_patch.shape)
X_test_image = image_patch.reshape(1,image_patch.shape[2],image_patch.shape[0],image_patch.shape[1]).astype('float32')
prediction = (model.predict_classes(X_test_image))
outputs[i+p][j+p] = prediction+1
ground_truth = spectral.imshow(classes=y, figsize=(10, 10))
predict_image = spectral.imshow(classes=outputs.astype(int), figsize=(10, 10))
```
| github_jupyter |
# Pendulum Environment, OpenAI Gym
* Left force: -50N, Right force: 50N, Nothing: 0N, with some amount of noise added to the action
* Generate trajectories by starting upright, and then applying random forces.
* Failure if the pendulum exceeds +/- pi/2
* Setting this problem up: how to encode Newtons? I'm starting things upright- how do we determine success?
```
import configargparse
import torch
import torch.optim as optim
import sys
sys.path.append('../')
from environments import PendulumEnv
from models.agents import NFQAgent
from models.networks import NFQNetwork, ContrastiveNFQNetwork
from util import get_logger, close_logger, load_models, make_reproducible, save_models
import matplotlib.pyplot as plt
import numpy as np
import itertools
import seaborn as sns
import tqdm
env = PendulumEnv()
rollouts = []
for i in range(10):
rollout, episode_cost = env.generate_rollout()
rollouts.extend(rollout)
rewards = [r[2] for r in rollouts]
sns.distplot(rewards)
def generate_data(init_experience=100, bg_only=False, agent=None):
env_bg = PendulumEnv(group=0)
env_fg = PendulumEnv(group=1)
bg_rollouts = []
fg_rollouts = []
if init_experience > 0:
for _ in range(init_experience):
rollout_bg, episode_cost = env_bg.generate_rollout(
agent, render=False, group=0
)
bg_rollouts.extend(rollout_bg)
if not bg_only:
rollout_fg, episode_cost = env_fg.generate_rollout(
agent, render=False, group=1
)
fg_rollouts.extend(rollout_fg)
bg_rollouts.extend(fg_rollouts)
all_rollouts = bg_rollouts.copy()
return all_rollouts, env_bg, env_fg
train_rollouts, train_env_bg, train_env_fg = generate_data(init_experience=200, bg_only=True)
test_rollouts, eval_env_bg, eval_env_fg = generate_data(init_experience=200, bg_only=True)
is_contrastive=True
epoch = 1000
hint_to_goal = False
if hint_to_goal:
goal_state_action_b_bg, goal_target_q_values_bg, group_bg = train_env_bg.get_goal_pattern_set(group=0)
goal_state_action_b_fg, goal_target_q_values_fg, group_fg = train_env_fg.get_goal_pattern_set(group=1)
goal_state_action_b_bg = torch.FloatTensor(goal_state_action_b_bg)
goal_target_q_values_bg = torch.FloatTensor(goal_target_q_values_bg)
goal_state_action_b_fg = torch.FloatTensor(goal_state_action_b_fg)
goal_target_q_values_fg = torch.FloatTensor(goal_target_q_values_fg)
nfq_net = ContrastiveNFQNetwork(state_dim=train_env_bg.state_dim, is_contrastive=is_contrastive, deep=False)
optimizer = optim.Adam(nfq_net.parameters(), lr=1e-1)
nfq_agent = NFQAgent(nfq_net, optimizer)
bg_success_queue = [0] * 3
fg_success_queue = [0] * 3
eval_fg = 0
evaluations = 5
for k, ep in enumerate(tqdm.tqdm(range(epoch + 1))):
state_action_b, target_q_values, groups = nfq_agent.generate_pattern_set(train_rollouts)
if hint_to_goal:
goal_state_action_b = torch.cat([goal_state_action_b_bg, goal_state_action_b_fg], dim=0)
goal_target_q_values = torch.cat([goal_target_q_values_bg, goal_target_q_values_fg], dim=0)
state_action_b = torch.cat([state_action_b, goal_state_action_b], dim=0)
target_q_values = torch.cat([target_q_values, goal_target_q_values], dim=0)
goal_groups = torch.cat([group_bg, group_fg], dim=0)
groups = torch.cat([groups, goal_groups], dim=0)
if not nfq_net.freeze_shared:
loss = nfq_agent.train((state_action_b, target_q_values, groups))
eval_episode_length_fg, eval_success_fg, eval_episode_cost_fg = 0, 0, 0
if nfq_net.freeze_shared:
eval_fg += 1
if eval_fg > 50:
loss = nfq_agent.train((state_action_b, target_q_values, groups))
(eval_episode_length_bg, eval_success_bg, eval_episode_cost_bg) = nfq_agent.evaluate_pendulum(eval_env_bg, render=False)
bg_success_queue = bg_success_queue[1:]
bg_success_queue.append(1 if eval_success_bg else 0)
(eval_episode_length_fg, eval_success_fg, eval_episode_cost_fg) = nfq_agent.evaluate_pendulum(eval_env_fg, render=False)
fg_success_queue = fg_success_queue[1:]
fg_success_queue.append(1 if eval_success_fg else 0)
if sum(bg_success_queue) == 3 and not nfq_net.freeze_shared == True:
nfq_net.freeze_shared = True
print("FREEZING SHARED")
if is_contrastive:
for param in nfq_net.layers_shared.parameters():
param.requires_grad = False
for param in nfq_net.layers_last_shared.parameters():
param.requires_grad = False
for param in nfq_net.layers_fg.parameters():
param.requires_grad = True
for param in nfq_net.layers_last_fg.parameters():
param.requires_grad = True
else:
for param in nfq_net.layers_fg.parameters():
param.requires_grad = False
for param in nfq_net.layers_last_fg.parameters():
param.requires_grad = False
optimizer = optim.Adam(
itertools.chain(
nfq_net.layers_fg.parameters(),
nfq_net.layers_last_fg.parameters(),
),
lr=1e-1,
)
nfq_agent._optimizer = optimizer
if sum(fg_success_queue) == 3:
print("Done Training")
break
if ep % 300 == 0:
perf_bg = []
perf_fg = []
for it in range(evaluations):
(eval_episode_length_bg,eval_success_bg,eval_episode_cost_bg) = nfq_agent.evaluate_pendulum(eval_env_bg, render=False)
(eval_episode_length_fg,eval_success_fg,eval_episode_cost_fg) = nfq_agent.evaluate_pendulum(eval_env_fg, render=False)
perf_bg.append(eval_episode_cost_bg)
perf_fg.append(eval_episode_cost_fg)
train_env_bg.close()
train_env_fg.close()
eval_env_bg.close()
eval_env_fg.close()
print("Evaluation bg: " + str(perf_bg) + " Evaluation fg: " + str(perf_fg))
perf_bg = []
perf_fg = []
for it in range(evaluations*10):
(eval_episode_length_bg,eval_success_bg,eval_episode_cost_bg) = nfq_agent.evaluate_car(eval_env_bg, render=False)
(eval_episode_length_fg,eval_success_fg,eval_episode_cost_fg) = nfq_agent.evaluate_car(eval_env_fg, render=False)
perf_bg.append(eval_episode_cost_bg)
perf_fg.append(eval_episode_cost_fg)
eval_env_bg.close()
eval_env_fg.close()
print("Evaluation bg: " + str(sum(perf_bg)/len(perf_bg)) + " Evaluation fg: " + str(sum(perf_fg)/len(perf_fg)))
```
| github_jupyter |
# NumPy
numpy is python's package for doing math that is more advanced than +-*/
This includes special functions like cosine, exponential, sqrt, ...
On top of this we can use numpy to generate samples from many types of random variables
numpy also has a powerful data type to define vectors, matrices, and tensors
With these data types numpy also allows us to do linear algebra - matrix multiplication and matrix-vector solutions
```
# the first step of using numpy is to tell python to use it
import numpy as np
print(np.cos(np.pi))
print(np.sqrt(1.21))
print(np.log(np.exp(5.2)))
# we can create numpy arrays by converting lists
# this is a vector
vec = np.array([1,2,3])
print(vec)
# we can create matrices by converting lists of lists
mat = np.array([[1,2,1],[4,5,9],[1,8,9]])
print('')
print(mat)
print('')
print(mat.T)
# there are lots of other ways to create numpy arrays
vec2 = np.arange(0,15)
print(vec2)
print('')
vec3 = np.arange(3,21,6)
print(vec3)
vec4 = np.linspace(0,2.7,10)
print(vec4)
print('')
print(vec4.reshape(5,2))
vec4_reshaped = vec4.reshape(5,2)
print(vec4_reshaped)
print(vec4)
mat2 = np.zeros([5,3])
print(mat2)
mat3 = np.ones([3,5])
print('')
print(mat3)
mat4 = np.eye(5)
print('')
print(mat4)
mat5 = np.identity(3)
print('')
print(mat5)
# we can +-*/ arrays together if they're the right size
vec5 = np.arange(1,6)
vec6 = np.arange(3,8)
print(vec5)
print(vec6)
print(vec5+vec6)
print(vec5*vec6)
print(1/vec5)
print(np.sqrt(vec6))
# we can do matrix multiplication
print(mat)
print('')
print(vec)
print()
product = np.matmul(mat,vec)
print(product)
np.dot(mat,vec)
print(np.linalg.solve(mat,product))
print('')
print(np.linalg.inv(mat))
# we can find the unique values in an array
vec7 = np.array(['blue','red','orange','purple','purple','orange','Red',6])
print(vec7)
print(np.unique(vec7))
# we can also use numpy to generate samples of a random variable
rand_mat = np.random.rand(5,5) # uniform random variable
print(rand_mat)
rand_mat2 = np.random.randn(10,5) # standard normal random variable
print('')
print(rand_mat2)
# we can also use numpy for statistical tools on arrays
print(np.mean(rand_mat))
print(np.std(rand_mat2))
print(np.min(rand_mat))
print(np.max(rand_mat2))
# break here for next video!
# how do we access entries in a numpy vector
rand_vec = np.random.randn(19)
print(rand_vec)
print(rand_vec[6])
# we can access multiple entries at once using :
print(rand_vec[4:9])
# we can also access multiple non-consecutive entries using np.arange
print(np.arange(0,15,3))
print(rand_vec[np.arange(0,15,3)])
# what about matrices
print(rand_mat)
print(rand_mat[1][2])
print(rand_mat[1,2])
print(rand_mat[0:2,1:3])
# let's change some values in an array!
print(rand_vec)
rand_vec[3:5] = 4
print('')
print(rand_vec)
rand_vec[3:5] = [1,2]
print('')
print(rand_vec)
print(rand_mat)
rand_mat[1:3,3:5] = 0
print('')
print(rand_mat)
sub_mat = rand_mat[0:2,0:3]
print(sub_mat)
sub_mat[:] = 3
print(sub_mat)
print(rand_mat)
sub_mat2 = rand_mat[0:2,0:3].copy()
sub_mat2[:] = 99
print(sub_mat2)
print(rand_mat)
# break here for next video
# we can also access entries with logicals
rand_vec = np.random.randn(15)
print(rand_vec)
print(rand_vec>0)
print(rand_vec[rand_vec>0])
print(rand_mat2)
print(rand_mat2[rand_mat2>0])
print(rand_vec)
print('')
rand_vec[rand_vec>0.5] = -5
print(rand_vec)
# let's save some arrays on the disk for use later!
np.save('saved_file_name',rand_mat2)
np.savez('zipped_file_name',rand_mat=rand_mat,rand_mat2=rand_mat2)
# now let's load it
loaded_vec = np.load('saved_file_name.npy')
loaded_zip = np.load('zipped_file_name.npz')
print(loaded_vec)
print('')
print(loaded_zip)
print(loaded_zip['rand_mat'])
print('')
print(loaded_zip['rand_mat2'])
new_array = loaded_zip['rand_mat']
print(new_array)
# we can also save/load as text files...but only single variables
np.savetxt('text_file_name.txt',rand_mat,delimiter=',')
rand_mat_txt = np.loadtxt('text_file_name.txt',delimiter=',')
print(rand_mat)
print('')
print(rand_mat_txt)
```
| github_jupyter |
```
class Scanner():
def __init__():
pass
def scan():
print('scan')
class Printer():
def __init__():
pass
def print():
print('printer print')
class Fax():
def print():
print('fax print')
def send():
print('send')
class MDFSPF(Scanner,Printer,Fax):
pass
class MDFSFP(Scanner,Fax,Printer):
pass
MDFSPF.print()
MDFSFP.print()
from datetime import datetime
def add_time(func):
def wrapper(*args):
print('this is the datetime',datetime.now())
for x in args:
print('one more time')
print(func(x))
return wrapper
def add_one(x):
return x + 1
def minus_one(x):
return x - 1
@add_time
def print_out(x):
return f'print {x}'
minus_one(1)
print_out(100000000000000000000000000000000000,123)
def decorator(func):
def inner(var):
func(var)
func(var)
return inner
@decorator
def xprint(var):
print('x',var)
xprint('poop')
class Yes():
__internal = 0
def __init__(self):
Yes.__internal += 1
@classmethod
def get_internal(cls):
return cls.__internal
Yes.get_internal()
yes = Yes()
yes.get_internal()
class Dog():
def __init__(self,name):
print('og __init__')
self.name = name
self.breed = ''
@classmethod
def add_breed(cls,name,breed):
print('new __init__ by the class method')
_dog = cls(name)
_dog.breed = breed
return _dog
dog = Dog('tim')
dog.name
dog.breed
dog2 = Dog.add_breed('bob','retriever')
print(dog2.name)
print(dog2.breed)
# class Error(BaseException):
class Watch():
created = 0
def __init__(self):
Watch.created += 1
self.text = ''
@classmethod
def engraved_watch(cls,text):
_watch = cls()
_watch.text = text
if Watch.validate(_watch):
return _watch
else:
raise Exception('oops')
@staticmethod
def validate(instance):
print(instance.text)
if len(instance.text) <= 40:
for char in instance.text:
if not char.isalnum():
return False
return True
else:
return False
mywatch = Watch()
mywatch.text = ' '
Watch.validate(mywatch)
Watch.created
newwatch = Watch.engraved_watch('9392')
newwatch.text
'a'.isalnum()
import abc
from random import randint
class Scanner(abc.ABC):
def scan_doc(self):
pass
def get_scan_stat(self):
pass
class Printer(abc.ABC):
def print_doc(self):
pass
def get_print_stat(self):
pass
class MFD1(Scanner,Printer):
def scan_doc(self):
return randint(1,100000),'has been scanned'
def get_scan_stat(self):
return 'succ'
def print_doc(self):
return randint(1,100000),'has been printed'
def get_print_stat(self):
return 'succ'
class MFD2(Scanner,Printer):
def scan_doc(self):
return randint(1,100000),'has been scanned'
def get_scan_stat(self):
return 'eh'
def print_doc(self):
return randint(1,100000),'has been printed'
def get_print_stat(self):
return 'eh'
def dance_for_me(self):
return 'dancing'
class MFD3(Scanner,Printer):
# def scan_doc(self):
# return randint(1,100000),'has been scanned'
def get_scan_stat(self):
return 'decent'
def print_doc(self):
return randint(1,100000),'has been printed'
def get_print_stat(self):
return 'decent'
def dance_for_me(self):
return 'wheeeeeeee'
def amazing(self):
return 'SUGEIIIIIIIIIIIIIIII-------------'
mfd1 = MFD1()
mfd2 = MFD2()
mfd3 = MFD3()
print(mfd2.print_doc())
print(mfd3.scan_doc())
class BankAccount():
def __init__(self,balance):
self.__balance = balance
@property
def balance(self):
return self.__balance
@balance.setter
def balance(self,value):
self.__balance = value
@balance.deleter
def balance(self):
self.__balance = None
mybankaccount = BankAccount(100)
print(mybankaccount.balance)
mybankaccount.balance += 10
print(mybankaccount.balance)
del mybankaccount.balance
print(mybankaccount.balance)
from random import randint
class Tires():
def __init__(self,max_pressure,size):
self.size = size
self.pressure = max_pressure
self.max_pressure = max_pressure
self.pressure -= randint(0,self.max_pressure)
def pump(self):
self.pressure = self.max_pressure
def get_pressure(self):
return self.pressure
class Engine():
def _init__(self,fuel_type):
self.fuel_type = fuel_type
def stop(self):
print('the engine is stopping')
def start(self):
print('the engine is starting')
def get_state(self):
print(self.fuel_type)
class CityCar(Tires,Engine):
def __init__(self):
super().__init__(30,15)
self.fuel_type = 'petrol'
citycar = CityCar()
citycar.pump()
citycar.get_pressure()
citycar.fuel_type
CityCar.mro()
from datetime import datetime
class MonitoredDict(dict):
def __init__(self):
self.log = list()
super().__init__()
def addlog(self,action,item):
now = datetime.now()
self.log.append(f'{item} was {action} on {now}')
def __setitem__(self,k,v):
super().__setitem__(k,v)
self.addlog('set_item',(k,v))
def __getitem__(self,k):
super().__getitem__(k)
self.addlog('get_item',(k))
mydict = MonitoredDict()
mydict[0] = 'a'
mydict[1] = 'asdfasd'
mydict.log
mydict[0]
try:
int('asd')
except ValueError as e:
print(e)
print(e.args)
print(e.name)
print(e.path)
print(e.encoding)
print(e.reason)
dir(Exception)
try:
b'\x80'.decode("utf-8")
except UnicodeError as e: # only for unicode
print(e)
print(e.encoding)
print(e.reason)
print(e.object)
print(e.start)
print(e.end)
a_list = ['First error', 'Second error']
class myError(Exception):
pass
def myfuncwentwrong():
raise myError('i wroteith is')
try:
print(a_list[36])
except Exception as f:
print(f,f.__context__)
myfuncwentwrong() # explicit
a_list = ['First error', 'Second error']
class myError(Exception):
pass
try:
print(a_list[3])
except Exception as e: # implicit
0/0
import traceback
class myException(Exception):
pass
def myfunc():
try:
print(X)
except Exception as e:
raise myException('i wrotethi stoo') from e
try:
myfunc()
except Exception as e:
print(e,e.__cause__,'and',e.__context__)
details = traceback.print_tb(e.__traceback__)
print(details)
astring = 'omg!'
bstring = astring
print(astring == bstring)
print(astring is bstring)
# both point to the same place in the memory
print('\n\n')
astring = 'omg!'
bstring = 'omg!'
print(astring == bstring)
print(astring is bstring)
# same content but points to different places in memory
import copy
astring = 'asdfasdfsdfadf'
bstring = copy.copy(astring)
print(id(astring) == id(bstring))
print(astring == bstring)
print(astring is bstring)
# shallow same mem spot
astring = 'pppp'
print(astring)
print(bstring)
astring = 'asdfasdfsdfadf'
bstring = copy.deepcopy(astring)
print(id(astring) == id(bstring))
bstring = 'puopopuiiop'
print(astring)
print(bstring)
import copy
# initializing list 1
li1 = [1, 2, [3,5], 4]
# using deepcopy to deep copy
li2 = copy.deepcopy(li1)
# original elements of list
print ("The original elements before deep copying")
for i in range(0,len(li1)):
print (li1[i],end=" ")
print("\r")
# adding and element to new list
li2[2][0] = 7
# Change is reflected in l2
print ("The new list of elements after deep copying ")
for i in range(0,len( li1)):
print (li2[i],end=" ")
print("\r")
# Change is NOT reflected in original list
# as it is a deep copy
print ("The original elements after deep copying")
for i in range(0,len( li1)):
print (li1[i],end=" ")
import copy
# initializing list 1
li1 = [1, 2, [3,5], 4]
string1 = 'abc'
# using copy to shallow copy
li2 = copy.copy(li1)
string2 = copy.copy(string1)
# original elements of list
print ("The original elements before shallow copying")
print(li1)
print("\r")
# adding and element to new list
li2[2][0] = 7
string2 += 'de'
# checking if change is reflected
print ("The original elements after shallow copying")
print(li1)
print(string1)
print('.')
print('\r')
print('.')
```
# COPY ONLY WORKS ON CONTAINERS
```
warehouse = list()
warehouse.append({'name': 'Lolly Pop', 'price': 0.4, 'weight': 133})
warehouse.append({'name': 'Licorice', 'price': 0.1, 'weight': 251})
warehouse.append({'name': 'Chocolate', 'price': 1, 'weight': 601})
warehouse.append({'name': 'Sours', 'price': 0.01, 'weight': 513})
warehouse.append({'name': 'Hard candies', 'price': 0.3, 'weight': 433})
print('Source list of candies')
warehouse2 = list()
for item in warehouse:
warehouse2.append(copy.deepcopy(item))
for item in warehouse2:
if item['weight'] > 300:
item['price'] *= .8
warehouse2
import pickle
a_dict = dict()
a_dict['EUR'] = {'code':'Euro', 'symbol': '€'}
a_dict['GBP'] = {'code':'Pounds sterling', 'symbol': '£'}
a_dict['USD'] = {'code':'US dollar', 'symbol': '$'}
a_dict['JPY'] = {'code':'Japanese yen', 'symbol': '¥'}
a_list = ['a', 123, [10, 100, 1000]]
with open('multidata.pckl', 'wb') as file_out:
pickle.dump(a_dict, file_out)
pickle.dump(a_list, file_out)
with open('multidata.pckl','rb') as file_in:
data1 = pickle.load(file_in)
data2 = pickle.load(file_in)
print(type(data1))
print(data1)
print(type(data2))
print(data2)
# pickles -> bytes = serilization
# dict of pickled stuff = shelves
for t in (type,list,str):
print(type(t))
type.__class__
dir(type)
type.__subclasses__(type)
class Class():
pass
Class.__class__
Class.__bases__
Class.__dict__
Class.__name__
bool.__class__
object.__class__
type.__class__
class Dog():
pass
dog = Dog()
Dog.__name__ = 'do'
Dog.__name__
Dog.__name__
do
Dog = type('Dog',(),{'age':2})
Dog.__bases__
dog = Dog()
dog.age
def greeting(self):
return 'hello how are you?'
class MetaClass(type):
def __new__(mcs,name,bases,dictionary):
if 'greeting' not in dictionary:
dictionary['greeting'] = greeting
obj = super().__new__(mcs,name,bases,dictionary)
return obj
class MyClass(metaclass=MetaClass):
pass
class OtherClass(metaclass=MetaClass):
def greeting(self):
return 'new thing'
thing = MyClass()
thing.greeting()
otherthing = OtherClass()
otherthing.greeting()
from datetime import datetime
def get_time():
return datetime.now()
def get_instantiation(self):
return self.instantiation_time
class Metaclass(type):
def __new__(mcs,name,bases,dictionary):
dictionary['instantiation_time'] = get_time()
dictionary['get_instantiation_time'] = get_instantiation
obj = super().__new__(mcs,name,bases,dictionary)
return obj
class myClass(metaclass=Metaclass):
pass
myobject = myClass()
myobject.get_instantiation_time()
myClass.__help__()
myobject.help()
class Thing():
Thing.y = 'z'
def __init__(self):
self.x = 'y'
thing = Thing()
getattr(Thing,'y')
setattr(thing,'123',123)
hasattr(Thing,'y')
Thing.__dict__
isinstance(thing,Thing)
class MyMeta(type):
def __new__(mcs,name,bases,dictionary):
obj = super().__new__(mcs,name,bases,dictionary)
return obj
class Mything(metaclass=MyMeta):
def __init__(self):
pass
class Mynewthing(Mything):
def __init__(self):
super().__init__()
mything = Mything()
issubclass(Mynewthing,Mything)
int.__dict__
3.sub()
class MyInt(int):
def __init__(self):
super().__init__()
def __eq__(self,num):
return not super().__eq__(num)
x = MyInt()
x == x
dir(10)
int.__dict__
y = 0xee1
int(y)
oct(12)
y = 0x1000000000
for x in range(10):
y = int(str(y)[2:])
y = hex(y)
print(y)
class MyInt(int):
def __init__(self):
super().__init__()
def __add__(self,num):
return super().__add__(num) + 348917256034
x = MyInt()
x + 10
x = x + 100123123
x
x + 1
a = 1
a.__add__(123)
import shelve
with shelve.open('spam') as db:
db['eggs'] = 'eggs'
l = [0,1,2,3,4,5]
print(l[-2:-1])
1%2
None + None
5 + 7 * 8
v = []
l = v[:]
l.append(1)
v
for x in range(-2,1):
print(x)
v = 1
while v < 10:
print(v)
v = v << 1
v = 1
if v == 1:
continue
print('something')
0 or 0
```
| github_jupyter |
# __Conceptos de estadística e introducción al análisis estadístico de datos usando Python__
```
#Importa las paqueterías necesarias
import numpy as np
import matplotlib.pyplot as pit
import pandas as pd
import seaborn as sns
import pandas_profiling as pp
from joblib import load, dump
import statsmodels.api as sm
```
Para este ejemplo ocuparemos bases de datos abiertas de crimen, registrados en Estados Unidos, específicamente una submuestra de la base de datos de crimen en Nueva York.
```
#Usar pandas para leer los datos y guardarlos como data frame
df_NY = load( "./datos_NY_crimen_limpios.pkl")
#Revisar si los datos fueron leidos correctamente
df_NY.head()
```
Diccionario de variables de la base de datos de crimenes en NY.
1. Ciudad: lugar en el que ocurrio el incidente
2. Fecha: año, mes y día en el que ocurrio el incidente
3. Hora: hora en la que ocurrio el incidente
4. Estatus: indicador de si el incidente fue completado o no
5. Gravedad: nivel del incidente; violación, delito mayor, delito menor
6. Lugar: lugar de ocurrencia del incidente; dentro, detras de, enfrente a y opuesto a...
7. Lugar especifico: lugar específico dónde ocurrio el incidente; tienda, casa habitación...
8. Crimen_tipo: descripción del tipo de delito
9. Edad_sospechoso: grupo de edad del sospechoso
10. Raza_sospechoso: raza del sospechoso
11. Sexo_sospechoso: sexo del sospechoso; M hombre, F mujer, U desconocido
12. Edad_victima: grupo de edad de la victima
13. Raza_victima: raza a la que pertenece la víctima
14. Sexo_victima: sexo de la victima; M hombre, F mujer, U desconocido
## 1.0 __Estadística descriptiva__
## 1.1 Conceptos de estadística descriptiva:
**Población**: conjunto de todos los elementos de interés (N).
**Parámetros**: métricas que obtenemos al trabajar con una población.
**Muestra**: subgrupo de la población (n).
**Estadísticos**: métricas que obtenemos al trabajar con poblaciones.

## 1.2 Una muestra debe ser:
**Representativa**: una muestra representativa es un subgrupo de la poblaciòn que refleja exactamente a los miembros de toda la población.
**Tomada al azar**: una muestra azarosa es recolectada cuando cada miembro de la muestra es elegida de la población estrictamente por casualidad
*¿Cómo sabemos que una muestra es representativa?¿Cómo calculamos el tamaño de muestra?*
Depende de los siguientes factores:
1. **Nivel de confianza**: ¿qué necesitamos para estar seguros de que nuestros resultados no ocurrieron solo por azar? Tipicamente se utiliza un nivel de confianza del _95% al 99%_
2. **Porcentaje de diferencia que deseemos detectar**: entre más pequeña sea la diferencia que quieres detectar, más grande debe ser la muestra
3. **Valor absoluto de las probabilidades en las que desea detectar diferencias**: depende de la prueba con la que estamos trabajando. Por ejemplo, detectar una diferencia entre 50% y 51% requiere un tamaño de muestra diferente que detectar una diferencia entre 80% y 81%. Es decir que, el tamaño de muestra requerido es una función de N1.
4. **La distribución de los datos (principalmente del resultado)**
## 1.3 ¿Qué es una variable?
**Variable**: es una característica, número o cantidad que puede ser descrita, medida o cuantíficada.
__Tipos de variables__:
1. Cualitativas o catégoricas: ordinales y nominales
2. Cuantitativas o numericas: discretas y continuas
```
#ORDINALES
#
#NOMINALES
#
#DISCRETAS
#
#CONTINUA
#
```
Variables de nuestra base de datos
```
df_NY.columns
```
## 1.4 ¿Cómo representar correctamente los diferentes tipos de variables?
__Datos categóricos:__ gráfica de barras, pastel, diagrama de pareto (tienen ambas barras y porcentajes)
__Datos numéricos:__ histograma y scatterplot
## 1.5 Atributos de las variables: medidas de tendencia central
Medidas de tendencia central: __media, mediana y moda__
1. **Media**: es la más común y la podemos obtener sumando todos los elementos de una variable y dividiéndola por el número de ellos. Es afectada por valores extremos
2. **Mediana**: número de la posición central de las observaciones (en orden ascendente). No es afectada por valores extremos.
3. **Moda**: el dato más común (puede existir más de una moda).

## 1.6 Atributos de las variables: medidas de asimetría (sesgo) o dispersión
__Sesgo__: indica si los datos se concentran en un lado de la curva
Por ejemplo:
1) cuando la medias es > que la mediana los datos se concentran del lado izquierdo de la curva, es decir que los outlier se encuentra del lado derecho de la distribución.
2) cuando la mediana < que la media, la mayor parte de los datos se concentran del lado derecho de la distribución y los outliers se encuentran en el lado izquierdo de la distribución.
En ambos casos la moda es la medida con mayor representación.
__Sin sesgo__: cuando la mediana, la moda y la media son iguales, la distribución es simétrica.
__El sesgo nos habla de donde se encuentran nuestros datos!__
## 1.7 Varianza
La __varianza__ es una medida de dispersión de un grupo de datos alrededor de la media.
Una forma más fácil de “visualizar” la varianza es por medio de la __desviación estandar__, en la mayoría de los casos esta es más significativa.
El __coeficiente de variación__ es igual a la desviación estándar dividida por el promedio
La desviación estandar es la medida más común de variabilidad para una base de datos única. Una de las principales ventajas de usar desviación estandar es que las unidades no estan elevadas al cuadrado y son más facil de interpretar
## 1.8 Relación entre variables
__Covarianza y Coeficiente de correlación lineal__
La covarianza puede ser >0, =0 o <0:
1. >0 las dos variables se mueven juntas
2. <0 las dos variables se mueven en direcciones opuestas
3. =0 las dos variables son independientes
El coeficiente de correlación va de -1 a 1
__Para explorar los atributos de cada una de las variables dentro de nuestra base de datos podemos hacer un profile report (podemos resolver toda la estadística descrptiva con un solo comando!!). Este reporte es el resultado de un análisis de cada una de las variables que integran la base de datos. Por medio de este, podemos verificar a que tipo de dato pertenece cada variable y obtener las medidas de tendencia central y asímetria. Con el fin de tener una idea general del comportamiento de nuestras variables.
Además, el profile report arroja un análisis de correlación entre variables (ver más adelante), que nos indica que tan relacionadas están entre si dos pares de variables__.
```
#pp.ProfileReport(df_NY[['Ciudad', 'Fecha', 'Hora', 'Estatus', 'Gravedad', 'Lugar','Crimen_tipo', 'Lugar_especifico', 'Edad_sospechoso', 'Raza_sospechoso','Sexo_sospechoso', 'Edad_victima', 'Raza_victima', 'Sexo_victima']])
```
## __2.0 Estadística inferencial__
## 2.1 Distribuciónes de probabilidad
Una __distribución__ es una función que muestra los valores posibles de una variable y que tan frecuentemente ocurren.
Es decir la __frecuencia__ en la que los posibles valores de una variable ocurren en un intervalo.
Las distribución más famosa en estadística(no precisamente la más común)es la __distribución normal__, donde la media moda y mediana son =. Es decir no hay sesgo
Frecuentemente, cuando los valores de una variable no tienen una distribución normal se recurre a transformaciones o estandarizaciones.
## 2.2 Regresión lineal
Una __regresión lineal__ es un modelo matemático para aproximar la relación de dependencia entre dos variables, una variable independiente y otra dependiente.
*Los valores de las variables dependientes dependen de los valores de las variables independientes*
## 2.3 Análisis de varianza
__Analisis de Varianza (ANOVA)__ se utiliza para comparar los promedios de dos o más grupos. Una prueba de ANOVA puede indicarte si hay diferencia en el promedio entre los grupos. Sin embargo,no nos da información sobre dónde se encuentra la diferencia (entre cuál y cuál grupo). Para resolver esto, podemos realizar una prueba post-hoc.
## __Análisis de base de datos abierta de delitos en NY__
### 1.0 Evaluar frecuencia de delitos
Podemos empezar por análizar los tipos de crimenes registrados, así como frecuencia de cada tipo de crimen.
```
#Usar value_counts en Pandas para cuantificar y organizar el tipo de crimenes
df_NY.Crimen_tipo.value_counts().iloc[:10]
df_NY.Crimen_tipo.value_counts().iloc[:10]
```
Ahora vamos a crear una grafica de los resultados para tener una mejor visualización de los datos.
```
df_NY.Crimen_tipo.value_counts().iloc[:10].plot(kind= "barh")
```
Podemos observar que los crimenes con mayor ocurrencia son "Petit larceny" y "Harraament 2"
### 1.1 Evaluar frecuencia de un delito específico: por ejemplo "Harrassment"
```
df_NY.dropna(inplace=True)
acoso = df_NY[df_NY["Crimen_tipo"].str.contains("HARRASSMENT 2")]
acoso.head(5)
```
## 2.0 Relaciones entre dos variables dependiente e independiente (de manera visual).
### 2.1 Análisis de la ocurrencia del __delito__ por __sitio__
¿Existen diferencias en la frecuencia de acoso en las diferentes localidades en NY? Es decir, qué lugares son más peligrosos.
En este ejemplo, la variable dependiente sería la ocurrecia del delito y la indenpendiente el sitio.
Para ello, usaremos la función __"groupby"__ de Pandas para agrupar por el tipo de localidades, y la función __size__ para revisar el número registrado en cada localidad.
```
acoso.columns
acoso.head()
acoso.groupby("Ciudad").size().sort_values(ascending=False)
acoso.Ciudad.value_counts().iloc[:10].plot(kind= "barh")
```
Al observar los resultados podemos distinguir en cuál de las localidades de NY hay mayores reportes de acoso. Brooklyn presenta más reportes de acoso.
```
acoso.Lugar_especifico.value_counts().iloc[:10].plot(kind= "barh")
```
El acoso ocurrió con mayor frecuencia dentro de casas y lugares de residencia.
### 2.2. Análisis de la ocurrencia del delito en el tiempo
Si queremos saber la frecuencia de ocurrencia del delito en diferentes años (2004-2018) y meses del año.
Aquí la variable dependiente es nuevamente la ocurrencia del delito y la independiente el tiempo.
```
acoso.groupby("anio").size().plot(kind="bar")
```
Podemos observar la mayoria de los resportes de acoso ocurrieron del 2016 al 2018. El 2011 fue el año con menor número de reportes de la ocurrencia de acoso
### 2.3. Analisis de ocurrencia del delito por sexo de la víctima y del agresor
En este ejemplo, la variable dependiente es el sexo de la víctima y la independiente el sexo del agresor
#### VICTIMAS
```
acoso.groupby("Sexo_victima").size().sort_values(ascending=False)
acoso.Sexo_victima.value_counts().iloc[:10].plot(kind= "pie")
acoso.groupby("Edad_victima").size().sort_values(ascending=False)
acoso.Edad_victima.value_counts().iloc[:10].plot(kind= "pie")
```
#### SOSPECHOSOS
```
acoso.groupby("Sexo_sospechoso").size().sort_values(ascending=False)
acoso.Sexo_sospechoso.value_counts().iloc[:10].plot(kind= "pie")
acoso.groupby("Edad_sospechoso").size().sort_values(ascending=False)
acoso.Edad_sospechoso.value_counts().iloc[:10].plot(kind= "pie")
```
### 2.4. Analisis de ocurrencia del delito por raza de la víctima y del agresor
En este ultimo ejemplo de relación entre variables, la variable dependiente es la raza de la víctima y la independiente es la raza del agresor.
#### VICTIMAS
```
acoso.groupby("Raza_victima").size().sort_values(ascending=False)
acoso.Raza_victima.value_counts().iloc[:10].plot(kind= "pie")
```
#### SOSPECHOSOS
```
acoso.groupby("Raza_sospechoso").size().sort_values(ascending=False)
acoso.Raza_sospechoso.value_counts().iloc[:10].plot(kind= "pie")
```
## 3.0 Regresión lineal
Pongamos a prueba la relación entre un par de variables. Por ejemplo, pero de la victima y peso del agresor. La relación puede ser negativa o positiva.
```
import pandas as pd
import statsmodels.api as sm
from sklearn import datasets, linear_model
df_w = pd.read_csv('Weight.csv')
df_w.head()
model = sm.OLS(y,X).fit()
predictions = model.predict(X)
print_model = model.summary()
print(print_model)
from scipy.stats import shapiro
stat, p = shapiro (y)
print('statistics=%.3f, p=%.3f' % (stat, p))
alpha = 0.05
if p > alpha:
print('its Gaussian')
else:
print('not Gaussian')
import statsmodels.api as sm
import pylab
sm.qqplot(y, loc = 4, scale = 3, line = 's')
pylab.show()
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
sns.set(color_codes = True)
sns.regplot(x = "AGRE_Weight ", y = "VIC_Weight", data = tamano);
```
## 4.0 ANOVA
Para realizar un análisis de variaza utilizando nuestros datos inicialmente debemos plantearnos una hipótesis. Por ejemplo: Existe diferencias en la edad de las víctimas entre los sitios donde ocurre ocoso.
Podemos probar nuetra hipótesis de manera estadística.
En este caso generaremos una columna extra de datos numericos continuos aproximados de "Edad_calculada_victima" y "Edad_calculada_agresor" para hacer el análisis
```
import pandas as pd
import scipy.stats as stats
import statsmodels. api as sm
from statsmodels.formula.api import ols
acoso["Edad_sospechoso"].unique()
from random import randint
def rango_a_random(s):
if type(s)==str:
s = s.split('-')
s = [int(i) for i in s]
s = randint(s[0],s[1]+1)
return s
acoso["Edad_calculada_victima"] = acoso["Edad_victima"]
acoso["Edad_calculada_victima"] = acoso["Edad_calculada_victima"].replace("65+","65-90").replace("<18","15-18").replace("UNKNOWN",np.nan)
acoso["Edad_calculada_victima"] = acoso["Edad_calculada_victima"].apply(rango_a_random)
acoso["Edad_calculada_sospechoso"] = acoso["Edad_sospechoso"]
acoso["Edad_calculada_sospechoso"] = acoso["Edad_calculada_sospechoso"].replace("65+","65-90").replace("<18","15-18").replace("UNKNOWN",np.nan)
acoso["Edad_calculada_sospechoso"] = acoso["Edad_calculada_sospechoso"].apply(rango_a_random)
acoso.head(5)
acoso.dropna ()
results = ols('Edad_calculada_victima ~ C(Ciudad)', data = acoso).fit()
results.summary()
```
En un análisis de varianza los dos "datos" de mayor importancia son el valor de F (F-statistic) y el valor de P (Prof F-statistic). Debemos obtener un avalor de P <0.05 para poder aceptar nuestra hipótesis.
En el ejemplo nuestro valor de F=4.129 y el de P=0.002. Es decir que podemos aceptar nuestra hipótesis.
| github_jupyter |
```
import json
import numpy as np
import tensorflow as tf
import collections
from sklearn.cross_validation import train_test_split
with open('ctexts.json','r') as fopen:
ctexts = json.loads(fopen.read())[:200]
with open('headlines.json','r') as fopen:
headlines = json.loads(fopen.read())[:200]
def build_dataset(words, n_words):
count = [['GO', 0], ['PAD', 1], ['EOS', 2], ['UNK', 3]]
count.extend(collections.Counter(words).most_common(n_words))
dictionary = dict()
for word, _ in count:
dictionary[word] = len(dictionary)
data = list()
unk_count = 0
for word in words:
index = dictionary.get(word, 0)
if index == 0:
unk_count += 1
data.append(index)
count[0][1] = unk_count
reversed_dictionary = dict(zip(dictionary.values(), dictionary.keys()))
return data, count, dictionary, reversed_dictionary
concat_from = ' '.join(ctexts).split()
vocabulary_size_from = len(list(set(concat_from)))
data_from, count_from, dictionary_from, rev_dictionary_from = build_dataset(concat_from, vocabulary_size_from)
print('vocab from size: %d'%(vocabulary_size_from))
print('Most common words', count_from[4:10])
print('Sample data', data_from[:10], [rev_dictionary_from[i] for i in data_from[:10]])
concat_to = ' '.join(headlines).split()
vocabulary_size_to = len(list(set(concat_to)))
data_to, count_to, dictionary_to, rev_dictionary_to = build_dataset(concat_to, vocabulary_size_to)
print('vocab to size: %d'%(vocabulary_size_to))
print('Most common words', count_to[4:10])
print('Sample data', data_to[:10], [rev_dictionary_to[i] for i in data_to[:10]])
for i in range(len(headlines)):
headlines[i] = headlines[i] + ' EOS'
headlines[0]
GO = dictionary_from['GO']
PAD = dictionary_from['PAD']
EOS = dictionary_from['EOS']
UNK = dictionary_from['UNK']
def str_idx(corpus, dic):
X = []
for i in corpus:
ints = []
for k in i.split():
try:
ints.append(dic[k])
except Exception as e:
print(e)
ints.append(UNK)
X.append(ints)
return X
X = str_idx(ctexts, dictionary_from)
Y = str_idx(headlines, dictionary_to)
train_X, test_X, train_Y, test_Y = train_test_split(X, Y, test_size = 0.2)
class Summarization:
def __init__(self, size_layer, num_layers, embedded_size,
from_dict_size, to_dict_size, batch_size):
def lstm_cell(reuse=False):
return tf.nn.rnn_cell.LSTMCell(size_layer, initializer=tf.orthogonal_initializer(),
reuse=reuse)
def attention(encoder_out, seq_len, reuse=False):
attention_mechanism = tf.contrib.seq2seq.LuongAttention(num_units = size_layer,
memory = encoder_out,
memory_sequence_length = seq_len)
return tf.contrib.seq2seq.AttentionWrapper(
cell = tf.nn.rnn_cell.MultiRNNCell([lstm_cell(reuse) for _ in range(num_layers)]),
attention_mechanism = attention_mechanism,
attention_layer_size = size_layer)
self.X = tf.placeholder(tf.int32, [None, None])
self.Y = tf.placeholder(tf.int32, [None, None])
self.X_seq_len = tf.placeholder(tf.int32, [None])
self.Y_seq_len = tf.placeholder(tf.int32, [None])
# encoder
encoder_embeddings = tf.Variable(tf.random_uniform([from_dict_size, embedded_size], -1, 1))
encoder_embedded = tf.nn.embedding_lookup(encoder_embeddings, self.X)
encoder_cells = tf.nn.rnn_cell.MultiRNNCell([lstm_cell() for _ in range(num_layers)])
self.encoder_out, self.encoder_state = tf.nn.dynamic_rnn(cell = encoder_cells,
inputs = encoder_embedded,
sequence_length = self.X_seq_len,
dtype = tf.float32)
self.encoder_state = tuple(self.encoder_state[-1] for _ in range(num_layers))
main = tf.strided_slice(self.Y, [0, 0], [batch_size, -1], [1, 1])
decoder_input = tf.concat([tf.fill([batch_size, 1], GO), main], 1)
# decoder
decoder_embeddings = tf.Variable(tf.random_uniform([to_dict_size, embedded_size], -1, 1))
decoder_cell = attention(self.encoder_out, self.X_seq_len)
dense_layer = tf.layers.Dense(to_dict_size)
training_helper = tf.contrib.seq2seq.TrainingHelper(
inputs = tf.nn.embedding_lookup(decoder_embeddings, decoder_input),
sequence_length = self.Y_seq_len,
time_major = False)
training_decoder = tf.contrib.seq2seq.BasicDecoder(
cell = decoder_cell,
helper = training_helper,
initial_state = decoder_cell.zero_state(batch_size, tf.float32).clone(cell_state=self.encoder_state),
output_layer = dense_layer)
training_decoder_output, _, _ = tf.contrib.seq2seq.dynamic_decode(
decoder = training_decoder,
impute_finished = True,
maximum_iterations = tf.reduce_max(self.Y_seq_len))
predicting_helper = tf.contrib.seq2seq.GreedyEmbeddingHelper(
embedding = encoder_embeddings,
start_tokens = tf.tile(tf.constant([GO], dtype=tf.int32), [batch_size]),
end_token = EOS)
predicting_decoder = tf.contrib.seq2seq.BasicDecoder(
cell = decoder_cell,
helper = predicting_helper,
initial_state = decoder_cell.zero_state(batch_size, tf.float32).clone(cell_state=self.encoder_state),
output_layer = dense_layer)
predicting_decoder_output, _, _ = tf.contrib.seq2seq.dynamic_decode(
decoder = predicting_decoder,
impute_finished = True,
maximum_iterations = 2 * tf.reduce_max(self.X_seq_len))
self.training_logits = training_decoder_output.rnn_output
self.predicting_ids = predicting_decoder_output.sample_id
masks = tf.sequence_mask(self.Y_seq_len, tf.reduce_max(self.Y_seq_len), dtype=tf.float32)
self.cost = tf.contrib.seq2seq.sequence_loss(logits = self.training_logits,
targets = self.Y,
weights = masks)
self.optimizer = tf.train.AdamOptimizer().minimize(self.cost)
size_layer = 128
num_layers = 2
embedded_size = 32
batch_size = 32
epoch = 5
tf.reset_default_graph()
sess = tf.InteractiveSession()
model = Summarization(size_layer, num_layers, embedded_size, len(dictionary_from),
len(dictionary_to), batch_size)
sess.run(tf.global_variables_initializer())
def pad_sentence_batch(sentence_batch, pad_int, maxlen=500):
padded_seqs = []
seq_lens = []
max_sentence_len = min(max([len(sentence) for sentence in sentence_batch]),maxlen)
for sentence in sentence_batch:
sentence = sentence[:maxlen]
padded_seqs.append(sentence + [pad_int] * (max_sentence_len - len(sentence)))
seq_lens.append(len(sentence))
return padded_seqs, seq_lens
def check_accuracy(logits, Y):
acc = 0
for i in range(logits.shape[0]):
internal_acc = 0
for k in range(len(Y[i])):
try:
if logits[i][k] == -1 and Y[i][k] == 1:
internal_acc += 1
elif Y[i][k] == logits[i][k]:
internal_acc += 1
except:
continue
acc += (internal_acc / len(Y[i]))
return acc / logits.shape[0]
for i in range(epoch):
total_loss, total_accuracy = 0, 0
for k in range(0, (len(train_X) // batch_size) * batch_size, batch_size):
batch_x, seq_x = pad_sentence_batch(train_X[k: k+batch_size], PAD)
batch_y, seq_y = pad_sentence_batch(train_Y[k: k+batch_size], PAD)
step, predicted, loss, _ = sess.run([model.global_step,
model.predicting_ids, model.cost, model.optimizer],
feed_dict={model.X:batch_x,
model.Y:batch_y,
model.X_seq_len:seq_x,
model.Y_seq_len:seq_y})
total_loss += loss
total_accuracy += check_accuracy(predicted,batch_y)
if step % 5 == 0:
rand = np.random.randint(0, len(test_Y)-batch_size)
batch_x, seq_x = pad_sentence_batch(test_X[rand:rand+batch_size], PAD)
batch_y, seq_y = pad_sentence_batch(test_Y[rand:rand+batch_size], PAD)
predicted, test_loss = sess.run([model.predicting_ids,model.cost], feed_dict={model.X:batch_x,
model.Y:batch_y,
model.X_seq_len:seq_x,
model.Y_seq_len:seq_y})
print('epoch %d, step %d, train loss %f, valid loss %f'%(i+1,step,loss,test_loss))
print('expected output:',' '.join([rev_dictionary_to[n] for n in batch_y[0] if n not in [-1,0,1,2,3]]))
print('predicted output:',' '.join([rev_dictionary_to[n] for n in predicted[0] n not in [-1,0,1,2,3]]),'\n')
total_loss /= (len(train_X) // batch_size)
total_accuracy /= (len(train_X) // batch_size)
print('epoch: %d, avg loss: %f, avg accuracy: %f'%(i+1, total_loss, total_accuracy))
```
| github_jupyter |
<a href="https://colab.research.google.com/github/2walkingfish/snow-activity-exploration/blob/main/Snow_instability_data_explaration.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
pd.set_option('display.max_columns', None)
df_features = pd.read_csv('/content/StevensPass-GraceLakes_4790_feet_2019.csv')
print(df_features.head())
```
**Statistical exploration**
```
print(df_features.info())
print(df_features.describe(include='all'))
```
**Ascending data sort by data**
```
df_features['Date/Time (PST)'] = pd.to_datetime(df_features['Date/Time (PST)'])
df_features.sort_values(by=['Date/Time (PST)'], ascending=True,inplace = True)
df_features.reset_index(drop=True, inplace = True)
print(df_features['Date/Time (PST)'])
```
**Visual feature exploration**
```
def plot_time_series(cl_time, cl_feature,time_from, time_to):
fig, ax = plt.subplots(figsize=(16, 8))
# Add x-axis and y-axis
df_month = df_features.loc[(time_from < df_features[cl_time]) & (df_features[cl_time] < time_to)]
ax.bar(df_month[cl_time],df_month[cl_feature], width = 0.1)
#ax.bar(df_features[cl_time], df_features[cl_feature])
# Set title and labels for axes
ax.set(xlabel="Date",
ylabel=cl_feature,
title= cl_feature + "\n From " + time_from + " to "+ time_to + " for Grace Lakes")
plt.show()
for i in df_features.columns[1:]:
plot_time_series('Date/Time (PST)',i,'2019-01-01','2019-03-01')
df_correlation = df_features.corr()
mask = np.zeros_like(df_correlation, dtype=np.bool)
mask[np.triu_indices_from(mask)] = True
# correlation matrix
f, ax = plt.subplots(figsize=(23, 20))
sns.heatmap(df_correlation, vmax=1, vmin = -1, center = 0, cmap='RdBu', mask=mask, annot = True, fmt="0.2f",robust=True, cbar_kws={'label':
'Correlation key: Exactly –1 a perfect downhill (negative) linear relationship, 0 no linear relationship,'
' +0.50 a moderate uphill (positive) relationship, +0.70 a strong uphill (positive) linear relationship,'
' Exactly +1 a perfect uphill (positive) linear relationship', 'orientation': 'horizontal'})
# ax.set(xlabel="Envirnment features", ylabel="Envirnment features")
ax.set_title(label='Envirnment features correlation heatmap',fontsize=20)
plt.setp( ax.get_xticklabels(), rotation=45, ha="right")
plt.tight_layout()
plt.show()
```
**Data engeneering**
```
df_features['Total Snow Depth Dayly Difference'] = df_features['Total Snow Depth (")'].diff()
print(df_features ['Total Snow Depth Dayly Difference'].describe())
fig, ax = plt.subplots(figsize=(16, 8))
df_features['Total Snow Depth Dayly Difference'].plot.box(fontsize=15)
from sklearn.cluster import DBSCAN
X = pd.DataFrame(df_features['Total Snow Depth Dayly Difference'])
X.fillna(value = 0,inplace=True)
#arbitrary substitution to detect only drop of snow level
X.loc[X['Total Snow Depth Dayly Difference'] > 5, 'Total Snow Depth Dayly Difference'] = 0
clustering = DBSCAN(eps=3, min_samples=15).fit(X)
df_features['Snow instability labels'] = clustering.labels_
```
**Visual exploration if significant drop of snow level was classified correctly**
```
for i in df_features.columns[3:]:
plot_time_series('Date/Time (PST)',i,'2019-01-01','2019-03-01')
df_features.groupby('Snow instability labels').size()
```
**Save enhanced data set for further avalanche predictions**
```
df_features.to_csv('/content/enhanced_StevensPass-GraceLakes_4790_feet_2019.csv')
```
| github_jupyter |
```
import random
from collections import deque
from copy import deepcopy
import gym
import matplotlib.pyplot as plt
import numpy as np
import torch
import torch.nn as nn
import torch.optim as optim
from torch.utils.data import DataLoader
from torch.distributions import Categorical
from IPython.display import clear_output
SEED = 1
BATCH_SIZE = 256
LR = 0.0003
UP_COEF = 0.25
GAMMA = 0.99
EPS = 1e-6
GRAD_NORM = False
# set device
use_cuda = torch.cuda.is_available()
device = torch.device('cuda' if use_cuda else 'cpu')
# random seed
random.seed(SEED)
np.random.seed(SEED)
torch.manual_seed(SEED)
if use_cuda:
torch.cuda.manual_seed_all(SEED)
class DQN(nn.Module):
def __init__(self, obs_space, action_space):
super().__init__()
H = 32
self.head = nn.Sequential(
nn.Linear(obs_space, H),
nn.Tanh()
)
self.fc = nn.Sequential(
nn.Linear(H, H),
nn.Tanh(),
nn.Linear(H, action_space)
)
def forward(self, x):
out = self.head(x)
q = self.fc(out).reshape(out.shape[0], -1)
return q
losses = []
def learn(net, tgt_net, optimizer, rep_memory):
global action_space
net.train()
tgt_net.train()
train_data = random.sample(rep_memory, BATCH_SIZE)
dataloader = DataLoader(
train_data, batch_size=BATCH_SIZE, pin_memory=use_cuda)
for i, (s, a, r, _s, d) in enumerate(dataloader):
s_batch = s.to(device).float()
a_batch = a.to(device).long()
_s_batch = _s.to(device).float()
r_batch = r.to(device).float()
done_mask = 1 - d.to(device).float()
with torch.no_grad():
_q_batch_tgt = tgt_net(_s_batch)
_q_max = torch.max(_q_batch_tgt, dim=1)[0]
q_batch = net(s_batch)
q_acting = q_batch[range(BATCH_SIZE), a_batch]
# loss
loss = (r_batch + GAMMA * done_mask * _q_max - q_acting).pow(2).mean()
losses.append(loss)
optimizer.zero_grad()
loss.backward()
if GRAD_NORM:
nn.utils.clip_grad_norm_(net.parameters(), max_norm=0.5)
optimizer.step()
def select_action(obs, tgt_net):
tgt_net.eval()
with torch.no_grad():
state = torch.tensor([obs]).to(device).float()
q = tgt_net(state)
action = torch.argmax(q)
return action.item()
def plot():
clear_output(True)
plt.figure(figsize=(16, 5))
plt.subplot(121)
plt.plot(rewards)
plt.title('Reward')
plt.subplot(122)
plt.plot(losses)
plt.title('Loss')
plt.show()
```
## Main
```
# make an environment
# env = gym.make('CartPole-v0')
env = gym.make('CartPole-v1')
# env = gym.make('MountainCar-v0')
# env = gym.make('LunarLander-v2')
env.seed(SEED)
obs_space = env.observation_space.shape[0]
action_space = env.action_space.n
# hyperparameter
n_episodes = 1000
learn_start = 1500
memory_size = 50000
update_frq = 1
use_eps_decay = False
epsilon = 0.001
eps_min = 0.001
decay_rate = 0.0001
n_eval = 10
# global values
total_steps = 0
learn_steps = 0
rewards = []
reward_eval = deque(maxlen=n_eval)
is_learned = False
is_solved = False
# make two nerual networks
net = DQN(obs_space, action_space).to(device)
target_net = deepcopy(net)
# make optimizer
optimizer = optim.AdamW(net.parameters(), lr=LR, eps=EPS)
# make a memory
rep_memory = deque(maxlen=memory_size)
use_cuda
env.spec.max_episode_steps
env.spec.reward_threshold
# play
for i in range(1, n_episodes + 1):
obs = env.reset()
done = False
ep_reward = 0
while not done:
# env.render()
if np.random.rand() < epsilon:
action = env.action_space.sample()
else:
action = select_action(obs, target_net)
_obs, reward, done, _ = env.step(action)
rep_memory.append((obs, action, reward, _obs, done))
obs = _obs
total_steps += 1
ep_reward += reward
if use_eps_decay:
epsilon -= epsilon * decay_rate
epsilon = max(eps_min, epsilon)
if len(rep_memory) >= learn_start:
if len(rep_memory) == learn_start:
print('\n============ Start Learning ============\n')
learn(net, target_net, optimizer, rep_memory)
learn_steps += 1
if learn_steps == update_frq:
# target smoothing update
with torch.no_grad():
for t, n in zip(target_net.parameters(), net.parameters()):
t.data = UP_COEF * n.data + (1 - UP_COEF) * t.data
learn_steps = 0
if done:
rewards.append(ep_reward)
reward_eval.append(ep_reward)
plot()
# print('{:3} Episode in {:5} steps, reward {:.2f}'.format(
# i, total_steps, ep_reward))
if len(reward_eval) >= n_eval:
if np.mean(reward_eval) >= env.spec.reward_threshold:
print('\n{} is sloved! {:3} Episode in {:3} steps'.format(
env.spec.id, i, total_steps))
torch.save(target_net.state_dict(),
f'./test/saved_models/{env.spec.id}_ep{i}_clear_model_dqn.pt')
break
env.close()
[
('CartPole-v0', 207, 0.25),
('CartPole-v1', 346, 0.25),
('MountainCar-v0', 304, 0.25),
('LunarLander-v2', 423, 0.25)
]
```
| github_jupyter |
# BFO Experiments
In these experiments we compared the ***Bacterial Foraging Optimization***
(BFO) algorithm, with 3 other popular nature-inspired algorithms:
***Differential Evolution*** (DE),
***Particle Swarm Optimization*** (PSO) and the ***Bat Algorithm*** (BA).
Their performance was evaluated on 10 popular test functions of different dimensions.
## Test Functions
| Name | Function | Search range | Global optimum |
| :- | :-: | :-: | :-: |
| Sphere | $$f_1(\vec{x}) = \sum_{i=1}^{D} x_i^2 $$ | $$[-5.12, 5.12]^D$$ | $$f_1(\vec{0}) = 0$$ |
| Rosenbrock | $$f_2(\vec{x}) = \sum_{i=1}^{D-1} (100 (x_{i+1} - x_i^2)^2 + (x_i - 1)^2) $$ | $$[-2.048, 2.048]^D$$ | $$f_2(\vec{1}) = 0$$ |
| Rastrigin | $$f_3(\vec{x}) = 10D + \sum_{i=1}^D \left(x_i^2 -10\cos(2\pi x_i)\right) $$ | $$[-5.12, 5.12]^D$$ | $$f_3(\vec{0}) = 0$$ |
| Griewank | $$f_4(\vec{x}) = \sum_{i=1}^D \frac{x_i^2}{4000} - \prod_{i=1}^D \cos(\frac{x_i}{\sqrt{i}}) + 1 $$ | $$[-600, 600]^D$$ | $$f_4(\vec{0}) = 0$$ |
| Ackley | $$f_5(\vec{x}) = -a\;\exp\left(-b \sqrt{\frac{1}{D} \sum_{i=1}^D x_i^2}\right) - \exp\left(\frac{1}{D} \sum_{i=1}^D cos(c\;x_i)\right) + a + \exp(1) $$ | $$[-32.768, 32.768]^D$$ | $$f_5(\vec{0}) = 0$$ |
| Schwefel | $$f_6(\vec{x}) = 418.9829d - \sum_{i=1}^{D} x_i \sin(\sqrt{\lvert x_i \rvert}) $$ | $$[-500, 500]^D$$ | $$f_6(\vec{420.9687}) = 0$$ |
| Alpine | $$f_7(\vec{x}) = \sum_{i=1}^{D} \lvert x_i \sin(x_i)+0.1x_i \rvert $$ | $$[-10, 10]^D$$ | $$f_7(\vec{0}) = 0$$ |
| Whitley | $$f_8(\vec{x}) = \sum_{i=1}^D \sum_{j=1}^D \left(\frac{(100(x_i^2-x_j)^2 + (1-x_j)^2)^2}{4000} - \cos(100(x_i^2-x_j)^2 + (1-x_j)^2)+1\right) $$ | $$[-10, 10]^D$$ | $$f_8(\vec{1}) = 0$$ |
| Csendes | $$f_9(\vec{x}) = \sum_{i=1}^D x_i^6\left( 2 + \sin \frac{1}{x_i}\right) $$ | $$[-1, 1]^D$$ | $$f_9(\vec{0}) = 0$$ |
| Dixon Price | $$f_{10}(\vec{x}) = (x_1 - 1)^2 + \sum_{i = 2}^D i (2x_i^2 - x_{i - 1})^2 $$ | $$[-10, 10]^D$$ | $$f_{10}(x^*) = 0, \quad x_i = 2^{-\frac{2^i - 2}{2^i}}$$|
## Parameter Settings
We selected the population size of 100 for all algorithms. The algorithm specific
parameters are described below.
### BFO
We used the classic version of BFO as presented by K. M. Passino,
with the following parameters:
$N_c = \frac{N_{gen}}{N_{re} \times N_{ed}}$, where $N_{gen}$ is the number
of iterations. This ensures that $N_c \times N_{re} \times N_{ed} = N_{gen}$,
if $N_{gen}$ is divisible by $N_{re} \times N_{ed}$, of course<br>
$N_s = 4$<br>
$N_{re} = 5$<br>
$N_{ed} = 2$<br>
$P_{ed} = 0.25$<br>
$C(i) = 0.1$<br>
$d_{attract} = 0.1$<br>
$w_{attract} = 0.2$<br>
$h_{repellent} = d_{attract}$<br>
$w_{repellent} = 10.0$<br>
### DE
We used the DE/rand/1/bin variant with ${CR}=0.9$ and $F=0.8$.
### PSO
We used the global best PSO with velocity clamping and a constant inertia
weight. Parameters: $w=0.9$, $c1=0.5$, $c2=0.3$, $v_{min} = -1.0$, $v_{max}=1.0$.
### BA
The standard Bat algorithm was used with: $A=1.0$, $r_0=1.0$, $\alpha = 0.97$, $\gamma=0.1$,
$Q_{min} = 0.0$, $Q_{max}=2.0$.
## The Experiments
We performed 25 independent runs for each algorithm on 10, 20 and 30 dimensional problems,
logging the best, worst and mean fitness values, along with the standard deviation.
The stopping condition was set as reaching $1000 \times D$, where $D$ is the dimension of the problem to be optimized.
```
import pandas as pd
df_10d = pd.read_pickle('results/10d.pkl')
df_20d = pd.read_pickle('results/20d.pkl')
df_30d = pd.read_pickle('results/30d.pkl')
```
#### Results on 10D functions
```
df_10d
```
#### Results on 20D functions
```
df_20d
```
#### Results on 30D functions
```
df_30d
```
#### Statistical significance
Here we will perform a Wilcoxon signed-rank test to see if our results carry any statistical significance.
```
from scipy.stats import wilcoxon
def perform_wilcoxon(df):
algorithms = df.columns
bfo_vals = df['BFO'].values
results = {}
for algorithm in algorithms[1:]:
s, p = wilcoxon(bfo_vals, df[algorithm].values)
results[algorithm] = p
return results
records = [perform_wilcoxon(df_10d), perform_wilcoxon(df_20d), perform_wilcoxon(df_30d)]
df_wilcoxon = pd.DataFrame.from_records(records)
df_wilcoxon['Dimension'] = [10, 20, 30]
df_wilcoxon = df_wilcoxon[['Dimension', 'DE', 'WVCPSO', 'BA']]
df_wilcoxon
```
| github_jupyter |
<!-- dom:TITLE: From Variational Monte Carlo to Boltzmann Machines and Machine Learning. Notebook 1: Variational Monte Carlo -->
# From Variational Monte Carlo to Boltzmann Machines and Machine Learning. Notebook 1: Variational Monte Carlo
<!-- dom:AUTHOR: Morten Hjorth-Jensen Email hjensen@msu.edu Department of Physics and Astronomy and National Superconducting Cyclotron Laboratory, Michigan State University, East Lansing, 48824 MI, USA -->
<!-- Author: -->
**Morten Hjorth-Jensen Email hjensen@msu.edu Department of Physics and Astronomy and National Superconducting Cyclotron Laboratory, Michigan State University, East Lansing, 48824 MI, USA**
Date: **Mar 21, 2019**
Copyright 1999-2019, Morten Hjorth-Jensen Email hjensen@msu.edu Department of Physics and Astronomy and National Superconducting Cyclotron Laboratory, Michigan State University, East Lansing, 48824 MI, USA. Released under CC Attribution-NonCommercial 4.0 license
## Introduction
### Structure and Aims
These notebooks serve the aim of linking traditional variational Monte
Carlo VMC calculations methods with recent progress on solving
many-particle problems using Machine Learning algorithms.
Furthermore, when linking with Machine Learning algorithms, in particular
so-called Boltzmann Machines, there are interesting connections between
these algorithms and so-called [Shadow Wave functions (SWFs)](https://journals.aps.org/pre/abstract/10.1103/PhysRevE.90.053304) (and references therein). The implications of the latter have been explored in various Monte Carlo calculations.
In total there are three notebooks:
1. the one you are reading now on Variational Monte Carlo methods,
2. notebook 2 on Machine Learning and quantum mechanical problems and in particular on Boltzmann Machines,
3. and finally notebook 3 on the link between Boltzmann machines and SWFs.
### This notebook
In this notebook the aim is to give you an introduction as well as an
understanding of the basic elements that are needed in order to
develop a professional variational Monte Carlo code. We will focus on
a simple system of two particles in an oscillator trap (or
alternatively two fermions moving in a Coulombic potential). The particles can
interact via a repulsive or an attrative force.
The advantage of these systems is that for two particles (boson or
fermions) we have analytical solutions for the eigenpairs of the
non-interacting case. Furthermore, for a two- or three-dimensional
system of two electrons moving in a harmonic oscillator trap, we have
[analytical solutions for the interacting case as well](https://iopscience.iop.org/article/10.1088/0305-4470/27/3/040/meta).
Having analytical eigenpairs is an invaluable feature that allows us
to assess the physical relevance of the trial wave functions, be
these either from a standard VMC procedure, from Boltzmann Machines or
from Shadow Wave functions.
In this notebook we start with the basics of a VMC calculation and
introduce concepts like Markov Chain Monte Carlo methods and the
Metropolis algorithm, importance sampling and Metropolis-Hastings
algorithm, resampling methods to obtain better estimates of the
statistical errors and minimization of the expectation values of the
energy and the variance. The latter is done in order to obtain the
best possible variational parameters. Furthermore it will define the
so-called **cost** function, a commonly encountered quantity in Machine
Learning algorithms. Minimizing the latter is the one which leads to
the determination of the optimal parameters in basically all Machine Learning algorithms.
For our purposes, it will serve as the first link between VMC methods and Machine Learning methods.
Topics like Markov Chain Monte Carlo and various resampling techniques
are also central to Machine Learning methods. Presenting them in the
context of VMC approaches leads hopefully to an easier starting point
for the understanding of these methods.
Finally, the reader may ask what do we actually want to achieve with
complicating life with Machine Learning methods when we can easily
study interacting systems with standard Monte Carlo approaches. Our
hope is that by adding additional degrees of freedom via Machine
Learning algorithms, we can let the algorithms we employ learn the
parameters of the model via a given optimization algorithm. In
standard Monte Carlo calculations the practitioners end up with fine tuning
the trial wave function using all possible insights about the system
understudy. This may not always lead to the best possible ansatz and
can in the long run be rather time-consuming. In fields like nuclear
many-body physics with complicated interaction terms, guessing an
analytical form for the trial wave fuction can be difficult. Letting
the machine learn the form of the trial function or find the optimal
parameters may lead to insights about the problem which cannot be
obtained by selecting various trial wave functions.
The emerging and rapidly expanding fields of Machine Learning and Quantum Computing hold also great promise in tackling the
dimensionality problems (the so-called dimensionality curse in many-body problems) we encounter when studying
complicated many-body problems.
The approach to Machine Learning we will focus on
is inspired by the idea of representing the wave function with
a restricted Boltzmann machine (RBM), presented recently by [G. Carleo and M. Troyer, Science **355**, Issue 6325, pp. 602-606 (2017)](http://science.sciencemag.org/content/355/6325/602). They
named such a wave function/network a *neural network quantum state* (NQS). In their article they apply it to the quantum mechanical
spin lattice systems of the Ising model and Heisenberg model, with
encouraging results.
Machine learning (ML) is an extremely rich field, in spite of its young age. The
increases we have seen during the last three decades in computational
capabilities have been followed by developments of methods and
techniques for analyzing and handling large data sets, relying heavily
on statistics, computer science and mathematics. The field is rather
new and developing rapidly.
Machine learning is the science of giving computers the ability to
learn without being explicitly programmed. The idea is that there
exist generic algorithms which can be used to find patterns in a broad
class of data sets without having to write code specifically for each
problem. The algorithm will build its own logic based on the data.
Machine learning is a subfield of computer science, and is closely
related to computational statistics. It evolved from the study of
pattern recognition in artificial intelligence (AI) research, and has
made contributions to AI tasks like computer vision, natural language
processing and speech recognition. It has also, especially in later
years, found applications in a wide variety of other areas, including
bioinformatics, economy, physics, finance and marketing.
An excellent reference we will come to back to is [Mehta *et al.*, arXiv:1803.08823](https://arxiv.org/abs/1803.08823).
Our focus will first be on the basics of VMC calculations.
## Basic Quantum Monte Carlo
We start with the variational principle.
Given a hamiltonian $H$ and a trial wave function $\Psi_T(\boldsymbol{R};\boldsymbol{\alpha})$, the variational principle states that the expectation value of $\cal{E}[H]$, defined through
$$
\cal {E}[H] =
\frac{\int d\boldsymbol{R}\Psi^{\ast}_T(\boldsymbol{R};\boldsymbol{\alpha})H(\boldsymbol{R})\Psi_T(\boldsymbol{R};\boldsymbol{\alpha})}
{\int d\boldsymbol{R}\Psi^{\ast}_T(\boldsymbol{R};\boldsymbol{\alpha})\Psi_T(\boldsymbol{R};\boldsymbol{\alpha})},
$$
is an upper bound to the ground state energy $E_0$ of the hamiltonian $H$, that is
$$
E_0 \le {\cal E}[H].
$$
In general, the integrals involved in the calculation of various
expectation values are multi-dimensional ones. Traditional integration
methods such as Gauss-Legendre quadrature will not be adequate for say the
computation of the energy of a many-body system.
Here we have defined the vector $\boldsymbol{R} = [\boldsymbol{r}_1,\boldsymbol{r}_2,\dots,\boldsymbol{r}_n]$ as an array that contains the positions of all particles $n$ while the vector $\boldsymbol{\alpha} = [\alpha_1,\alpha_2,\dots,\alpha_m]$ contains the variational parameters of the model, $m$ in total.
The trial wave function can be expanded in the eigenstates $\Psi_i(\boldsymbol{R})$
of the hamiltonian since they form a complete set, viz.,
$$
\Psi_T(\boldsymbol{R};\boldsymbol{\alpha})=\sum_i a_i\Psi_i(\boldsymbol{R}),
$$
and assuming that the set of eigenfunctions are normalized, one obtains
$$
\frac{\sum_{nm}a^*_ma_n \int d\boldsymbol{R}\Psi^{\ast}_m(\boldsymbol{R})H(\boldsymbol{R})\Psi_n(\boldsymbol{R})}
{\sum_{nm}a^*_ma_n \int d\boldsymbol{R}\Psi^{\ast}_m(\boldsymbol{R})\Psi_n(\boldsymbol{R})} =\frac{\sum_{n}a^2_n E_n}
{\sum_{n}a^2_n} \ge E_0,
$$
where we used that $H(\boldsymbol{R})\Psi_n(\boldsymbol{R})=E_n\Psi_n(\boldsymbol{R})$.
In general, the integrals involved in the calculation of various expectation
values are multi-dimensional ones.
The variational principle yields the lowest energy of states with a given symmetry.
In most cases, a wave function has only small values in large parts of
configuration space, and a straightforward procedure which uses
homogenously distributed random points in configuration space
will most likely lead to poor results. This may suggest that some kind
of importance sampling combined with e.g., the Metropolis algorithm
may be a more efficient way of obtaining the ground state energy.
The hope is then that those regions of configurations space where
the wave function assumes appreciable values are sampled more
efficiently.
The tedious part in a VMC calculation is the search for the variational
minimum. A good knowledge of the system is required in order to carry out
reasonable VMC calculations. This is not always the case,
and often VMC calculations
serve rather as the starting
point for so-called diffusion Monte Carlo calculations (DMC). Diffusion Monte Carlo is a way of
solving exactly the many-body Schroedinger equation by means of
a stochastic procedure. A good guess on the binding energy
and its wave function is however necessary.
A carefully performed VMC calculation can aid in this context.
The basic procedure of a Variational Monte Carlo calculations consists thus of
1. Construct first a trial wave function $\psi_T(\boldsymbol{R};\boldsymbol{\alpha})$, for a many-body system consisting of $n$ particles located at positions $\boldsymbol{R}=(\boldsymbol{R}_1,\dots ,\boldsymbol{R}_n)$. The trial wave function depends on $\alpha$ variational parameters $\boldsymbol{\alpha}=(\alpha_1,\dots ,\alpha_M)$.
2. Then we evaluate the expectation value of the hamiltonian $H$
$$
\overline{E}[\boldsymbol{\alpha}]=\frac{\int d\boldsymbol{R}\Psi^{\ast}_{T}(\boldsymbol{R},\boldsymbol{\alpha})H(\boldsymbol{R})\Psi_{T}(\boldsymbol{R},\boldsymbol{\alpha})}
{\int d\boldsymbol{R}\Psi^{\ast}_{T}(\boldsymbol{R},\boldsymbol{\alpha})\Psi_{T}(\boldsymbol{R},\boldsymbol{\alpha})}.
$$
1. Thereafter we vary $\boldsymbol{\alpha}$ according to some minimization algorithm and return eventually to the first step if we are not satisfied with the results.
Here we have used the notation $\overline{E}$ to label the expectation value of the energy.
### Linking with standard statistical expressions for expectation values
In order to bring in the Monte Carlo machinery, we define first a likelihood distribution, or probability density distribution (PDF). Using our ansatz for the trial wave function $\psi_T(\boldsymbol{R};\boldsymbol{\alpha})$ we define a PDF
$$
P(\boldsymbol{R})= \frac{\left|\psi_T(\boldsymbol{R};\boldsymbol{\alpha})\right|^2}{\int \left|\psi_T(\boldsymbol{R};\boldsymbol{\alpha})\right|^2d\boldsymbol{R}}.
$$
This is our model for probability distribution function.
The approximation to the expectation value of the Hamiltonian is now
$$
\overline{E}[\boldsymbol{\alpha}] =
\frac{\int d\boldsymbol{R}\Psi^{\ast}_T(\boldsymbol{R};\boldsymbol{\alpha})H(\boldsymbol{R})\Psi_T(\boldsymbol{R};\boldsymbol{\alpha})}
{\int d\boldsymbol{R}\Psi^{\ast}_T(\boldsymbol{R};\boldsymbol{\alpha})\Psi_T(\boldsymbol{R};\boldsymbol{\alpha})}.
$$
We define a new quantity
<!-- Equation labels as ordinary links -->
<div id="eq:locale1"></div>
$$
E_L(\boldsymbol{R};\boldsymbol{\alpha})=\frac{1}{\psi_T(\boldsymbol{R};\boldsymbol{\alpha})}H\psi_T(\boldsymbol{R};\boldsymbol{\alpha}),
\label{eq:locale1} \tag{1}
$$
called the local energy, which, together with our trial PDF yields a new expression (and which look simlar to the the expressions for moments in statistics)
<!-- Equation labels as ordinary links -->
<div id="eq:vmc1"></div>
$$
\overline{E}[\boldsymbol{\alpha}]=\int P(\boldsymbol{R})E_L(\boldsymbol{R};\boldsymbol{\alpha}) d\boldsymbol{R}\approx \frac{1}{N}\sum_{i=1}^NE_L(\boldsymbol{R_i};\boldsymbol{\alpha})
\label{eq:vmc1} \tag{2}
$$
with $N$ being the number of Monte Carlo samples. The expression on the right hand side follows from Bernoulli's law of large numbers, which states that the sample mean, in the limit $N\rightarrow \infty$ approaches the true mean
The Algorithm for performing a variational Monte Carlo calculations runs as this
* Initialisation: Fix the number of Monte Carlo steps. Choose an initial $\boldsymbol{R}$ and variational parameters $\alpha$ and calculate $\left|\psi_T^{\alpha}(\boldsymbol{R})\right|^2$.
* Initialise the energy and the variance and start the Monte Carlo calculation.
* Calculate a trial position $\boldsymbol{R}_p=\boldsymbol{R}+r*step$ where $r$ is a random variable $r \in [0,1]$.
* Metropolis algorithm to accept or reject this move $w = P(\boldsymbol{R}_p)/P(\boldsymbol{R})$.
* If the step is accepted, then we set $\boldsymbol{R}=\boldsymbol{R}_p$.
* Update averages
* Finish and compute final averages.
Observe that the jumping in space is governed by the variable *step*. This is called brute-force sampling and is normally replaced by what is called **importance sampling**, discussed in more detail below here.
### Simple example, the hydrogen atom
The radial Schroedinger equation for the hydrogen atom can be
written as (when we have gotten rid of the first derivative term in the kinetic energy and used $rR(r)=u(r)$)
$$
-\frac{\hbar^2}{2m}\frac{d^2 u(r)}{d r^2}-
\left(\frac{ke^2}{r}-\frac{\hbar^2l(l+1)}{2mr^2}\right)u(r)=Eu(r).
$$
We will specialize to the case with $l=0$ and end up with
$$
-\frac{\hbar^2}{2m}\frac{d^2 u(r)}{d r^2}-
\left(\frac{ke^2}{r}\right)u(r)=Eu(r).
$$
Then we introduce a dimensionless variable $\rho=r/a$ where $a$ is a constant with dimension length.
Multiplying with $ma^2/\hbar^2$ we can rewrite our equations as
$$
-\frac{1}{2}\frac{d^2 u(\rho)}{d \rho^2}-
\frac{ke^2ma}{\hbar^2}\frac{u(\rho)}{\rho}-\lambda u(\rho)=0.
$$
Since $a$ is just a parameter we choose to set
$$
\frac{ke^2ma}{\hbar^2}=1,
$$
which leads to $a=\hbar^2/mke^2$, better known as the Bohr radius with value $0.053$ nm. Scaling the equations this way does not only render our numerical treatment simpler since we avoid carrying with us all physical parameters, but we obtain also a **natural** length scale. We will see this again and again. In our discussions below with a harmonic oscillator trap, the **natural** lentgh scale with be determined by the oscillator frequency, the mass of the particle and $\hbar$. We have also defined a dimensionless 'energy' $\lambda = Ema^2/\hbar^2$.
With the rescaled quantities, the ground state energy of the hydrogen atom is $1/2$.
The equation we want to solve is now defined by the Hamiltonian
$$
H=-\frac{1}{2}\frac{d^2 }{d \rho^2}-\frac{1}{\rho}.
$$
As trial wave function we peep now into the analytical solution for
the hydrogen atom and use (with $\alpha$ as a variational parameter)
$$
u_T^{\alpha}(\rho)=\alpha\rho \exp{-(\alpha\rho)}.
$$
Inserting this wave function into the expression for the
local energy $E_L$ gives
$$
E_L(\rho)=-\frac{1}{\rho}-
\frac{\alpha}{2}\left(\alpha-\frac{2}{\rho}\right).
$$
To have analytical local energies saves us from computing numerically
the second derivative, a feature which often increases our numerical
expenditure with a factor of three or more. Integratng up the local energy (recall to bring back the PDF in the integration) gives $\overline{E}[\boldsymbol{\alpha}]=\alpha(\alpha/2-1)$.
### Second example, the harmonic oscillator in one dimension
We present here another well-known example, the harmonic oscillator in
one dimension for one particle. This will also serve the aim of
introducing our next model, namely that of interacting electrons in a
harmonic oscillator trap.
Here as well, we do have analytical solutions and the energy of the
ground state, with $\hbar=1$, is $1/2\omega$, with $\omega$ being the
oscillator frequency. We use the following trial wave function
$$
\psi_T(x;\alpha) = \exp{-(\frac{1}{2}\alpha^2x^2)},
$$
which results in a local energy
$$
\frac{1}{2}\left(\alpha^2+x^2(1-\alpha^4)\right).
$$
We can compare our numerically calculated energies with the exact energy as function of $\alpha$
$$
\overline{E}[\alpha] = \frac{1}{4}\left(\alpha^2+\frac{1}{\alpha^2}\right).
$$
Similarly, with the above ansatz, we can also compute the exact variance which reads
$$
\sigma^2[\alpha]=\frac{1}{4}\left(1+(1-\alpha^4)^2\frac{3}{4\alpha^4}\right)-\overline{E}.
$$
Our code for computing the energy of the ground state of the harmonic oscillator follows here. We start by defining directories where we store various outputs.
```
# Common imports
import os
# Where to save the figures and data files
PROJECT_ROOT_DIR = "Results"
FIGURE_ID = "Results/FigureFiles"
DATA_ID = "Results/VMCHarmonic"
if not os.path.exists(PROJECT_ROOT_DIR):
os.mkdir(PROJECT_ROOT_DIR)
if not os.path.exists(FIGURE_ID):
os.makedirs(FIGURE_ID)
if not os.path.exists(DATA_ID):
os.makedirs(DATA_ID)
def image_path(fig_id):
return os.path.join(FIGURE_ID, fig_id)
def data_path(dat_id):
return os.path.join(DATA_ID, dat_id)
def save_fig(fig_id):
plt.savefig(image_path(fig_id) + ".png", format='png')
outfile = open(data_path("VMCHarmonic.dat"),'w')
```
We proceed with the implementation of the Monte Carlo algorithm but list first the ansatz for the wave function and the expression for the local energy
```
%matplotlib inline
# VMC for the one-dimensional harmonic oscillator
# Brute force Metropolis, no importance sampling and no energy minimization
from math import exp, sqrt
from random import random, seed
import numpy as np
import matplotlib.pyplot as plt
from numba import jit
from decimal import *
# Trial wave function for the Harmonic oscillator in one dimension
def WaveFunction(r,alpha):
return exp(-0.5*alpha*alpha*r*r)
# Local energy for the Harmonic oscillator in one dimension
def LocalEnergy(r,alpha):
return 0.5*r*r*(1-alpha**4) + 0.5*alpha*alpha
```
Note that in the Metropolis algorithm there is no need to compute the
trial wave function, mainly since we are just taking the ratio of two
exponentials. It is then from a computational point view, more
convenient to compute the argument from the ratio and then calculate
the exponential. Here we have refrained from this purely of
pedagogical reasons.
```
# The Monte Carlo sampling with the Metropolis algo
# The jit decorator tells Numba to compile this function.
# The argument types will be inferred by Numba when the function is called.
@jit
def MonteCarloSampling():
NumberMCcycles= 100000
StepSize = 1.0
# positions
PositionOld = 0.0
PositionNew = 0.0
# seed for rng generator
seed()
# start variational parameter
alpha = 0.4
for ia in range(MaxVariations):
alpha += .05
AlphaValues[ia] = alpha
energy = energy2 = 0.0
#Initial position
PositionOld = StepSize * (random() - .5)
wfold = WaveFunction(PositionOld,alpha)
#Loop over MC MCcycles
for MCcycle in range(NumberMCcycles):
#Trial position
PositionNew = PositionOld + StepSize*(random() - .5)
wfnew = WaveFunction(PositionNew,alpha)
#Metropolis test to see whether we accept the move
if random() <= wfnew**2 / wfold**2:
PositionOld = PositionNew
wfold = wfnew
DeltaE = LocalEnergy(PositionOld,alpha)
energy += DeltaE
energy2 += DeltaE**2
#We calculate mean, variance and error
energy /= NumberMCcycles
energy2 /= NumberMCcycles
variance = energy2 - energy**2
error = sqrt(variance/NumberMCcycles)
Energies[ia] = energy
Variances[ia] = variance
outfile.write('%f %f %f %f \n' %(alpha,energy,variance,error))
return Energies, AlphaValues, Variances
```
Finally, the results are presented here with the exact energies and variances as well.
```
#Here starts the main program with variable declarations
MaxVariations = 20
Energies = np.zeros((MaxVariations))
ExactEnergies = np.zeros((MaxVariations))
ExactVariance = np.zeros((MaxVariations))
Variances = np.zeros((MaxVariations))
AlphaValues = np.zeros(MaxVariations)
(Energies, AlphaValues, Variances) = MonteCarloSampling()
outfile.close()
ExactEnergies = 0.25*(AlphaValues*AlphaValues+1.0/(AlphaValues*AlphaValues))
ExactVariance = 0.25*(1.0+((1.0-AlphaValues**4)**2)*3.0/(4*(AlphaValues**4)))-ExactEnergies*ExactEnergies
#simple subplot
plt.subplot(2, 1, 1)
plt.plot(AlphaValues, Energies, 'o-',AlphaValues, ExactEnergies,'r-')
plt.title('Energy and variance')
plt.ylabel('Dimensionless energy')
plt.subplot(2, 1, 2)
plt.plot(AlphaValues, Variances, '.-',AlphaValues, ExactVariance,'r-')
plt.xlabel(r'$\alpha$', fontsize=15)
plt.ylabel('Variance')
save_fig("VMCHarmonic")
plt.show()
#nice printout with Pandas
import pandas as pd
from pandas import DataFrame
data ={'Alpha':AlphaValues, 'Energy':Energies,'Exact Energy':ExactEnergies,'Variance':Variances,'Exact Variance':ExactVariance,}
frame = pd.DataFrame(data)
print(frame)
```
For $\alpha=1$ we have the exact eigenpairs, as can be deduced from the
table here. With $\omega=1$, the exact energy is $1/2$ a.u. with zero
variance, as it should. We see also that our computed variance follows rather well the exact variance.
Increasing the number of Monte Carlo cycles will improve our statistics (try to increase the number of Monte Carlo cycles).
The fact that the variance is exactly equal to zero when $\alpha=1$ is that
we then have the exact wave function, and the action of the hamiltionan
on the wave function
$$
H\psi = \mathrm{constant}\times \psi,
$$
yields just a constant. The integral which defines various
expectation values involving moments of the hamiltonian becomes then
$$
\langle H^n \rangle =
\frac{\int d\boldsymbol{R}\Psi^{\ast}_T(\boldsymbol{R})H^n(\boldsymbol{R})\Psi_T(\boldsymbol{R})}
{\int d\boldsymbol{R}\Psi^{\ast}_T(\boldsymbol{R})\Psi_T(\boldsymbol{R})}=
\mathrm{constant}\times\frac{\int d\boldsymbol{R}\Psi^{\ast}_T(\boldsymbol{R})\Psi_T(\boldsymbol{R})}
{\int d\boldsymbol{R}\Psi^{\ast}_T(\boldsymbol{R})\Psi_T(\boldsymbol{R})}=\mathrm{constant}.
$$
**This gives an important information: the exact wave function leads to zero variance!**
As we will see below, many practitioners perform a minimization on both the energy and the variance.
## The Metropolis algorithm
Till now we have not yet discussed the derivation of the Metropolis algorithm. We assume the reader has some familiarity with the mathematics of Markov chains.
The Metropolis algorithm , see [the original article](http://scitation.aip.org/content/aip/journal/jcp/21/6/10.1063/1.1699114), was invented by Metropolis et. al
and is often simply called the Metropolis algorithm.
It is a method to sample a normalized probability
distribution by a stochastic process. We define $\mathbf{P}_i^{(n)}$ to
be the probability for finding the system in the state $i$ at step $n$.
The algorithm is then
* Sample a possible new state $j$ with some probability $T_{i\rightarrow j}$.
* Accept the new state $j$ with probability $A_{i \rightarrow j}$ and use it as the next sample. With probability $1-A_{i\rightarrow j}$ the move is rejected and the original state $i$ is used again as a sample.
We wish to derive the required properties of $T$ and $A$ such that
$\mathbf{P}_i^{(n\rightarrow \infty)} \rightarrow p_i$ so that starting
from any distribution, the method converges to the correct distribution.
Note that the description here is for a discrete probability distribution.
Replacing probabilities $p_i$ with expressions like $p(x_i)dx_i$ will
take all of these over to the corresponding continuum expressions.
The dynamical equation for $\mathbf{P}_i^{(n)}$ can be written directly from
the description above. The probability of being in the state $i$ at step $n$
is given by the probability of being in any state $j$ at the previous step,
and making an accepted transition to $i$ added to the probability of
being in the state $i$, making a transition to any state $j$ and
rejecting the move:
$$
\mathbf{P}^{(n)}_i = \sum_j \left [
\mathbf{P}^{(n-1)}_jT_{j\rightarrow i} A_{j\rightarrow i}
+\mathbf{P}^{(n-1)}_iT_{i\rightarrow j}\left ( 1- A_{i\rightarrow j} \right)
\right ] \,.
$$
Since the probability of making some transition must be 1,
$\sum_j T_{i\rightarrow j} = 1$, and the above equation becomes
$$
\mathbf{P}^{(n)}_i = \mathbf{P}^{(n-1)}_i +
\sum_j \left [
\mathbf{P}^{(n-1)}_jT_{j\rightarrow i} A_{j\rightarrow i}
-\mathbf{P}^{(n-1)}_iT_{i\rightarrow j}A_{i\rightarrow j}
\right ] \,.
$$
For large $n$ we require that $\mathbf{P}^{(n\rightarrow \infty)}_i = p_i$,
the desired probability distribution. Taking this limit, gives the
balance requirement
$$
\sum_j \left [
p_jT_{j\rightarrow i} A_{j\rightarrow i}
-p_iT_{i\rightarrow j}A_{i\rightarrow j}
\right ] = 0 \,.
$$
The balance requirement is very weak. Typically the much stronger detailed
balance requirement is enforced, that is rather than the sum being
set to zero, we set each term separately to zero and use this
to determine the acceptance probabilities. Rearranging, the result is
$$
\frac{ A_{j\rightarrow i}}{A_{i\rightarrow j}}
= \frac{p_iT_{i\rightarrow j}}{ p_jT_{j\rightarrow i}} \,.
$$
The Metropolis choice is to maximize the $A$ values, that is
$$
A_{j \rightarrow i} = \min \left ( 1,
\frac{p_iT_{i\rightarrow j}}{ p_jT_{j\rightarrow i}}\right ).
$$
Other choices are possible, but they all correspond to multilplying
$A_{i\rightarrow j}$ and $A_{j\rightarrow i}$ by the same constant
smaller than unity.\footnote{The penalty function method uses just such
a factor to compensate for $p_i$ that are evaluated stochastically
and are therefore noisy.}
Having chosen the acceptance probabilities, we have guaranteed that
if the $\mathbf{P}_i^{(n)}$ has equilibrated, that is if it is equal to $p_i$,
it will remain equilibrated. Next we need to find the circumstances for
convergence to equilibrium.
The dynamical equation can be written as
$$
\mathbf{P}^{(n)}_i = \sum_j M_{ij}\mathbf{P}^{(n-1)}_j
$$
with the matrix $M$ given by
$$
M_{ij} = \delta_{ij}\left [ 1 -\sum_k T_{i\rightarrow k} A_{i \rightarrow k}
\right ] + T_{j\rightarrow i} A_{j\rightarrow i} \,.
$$
Summing over $i$ shows that $\sum_i M_{ij} = 1$, and since
$\sum_k T_{i\rightarrow k} = 1$, and $A_{i \rightarrow k} \leq 1$, the
elements of the matrix satisfy $M_{ij} \geq 0$. The matrix $M$ is therefore
a stochastic matrix.
The Metropolis method is simply the power method for computing the
right eigenvector of $M$ with the largest magnitude eigenvalue.
By construction, the correct probability distribution is a right eigenvector
with eigenvalue 1. Therefore, for the Metropolis method to converge
to this result, we must show that $M$ has only one eigenvalue with this
magnitude, and all other eigenvalues are smaller.
## The system: two electrons in a harmonic oscillator trap in two dimensions
The Hamiltonian of the quantum dot is given by
$$
\hat{H} = \hat{H}_0 + \hat{V},
$$
where $\hat{H}_0$ is the many-body HO Hamiltonian, and $\hat{V}$ is the
inter-electron Coulomb interactions. In dimensionless units,
$$
\hat{V}= \sum_{i < j}^N \frac{1}{r_{ij}},
$$
with $r_{ij}=\sqrt{\mathbf{r}_i^2 - \mathbf{r}_j^2}$.
This leads to the separable Hamiltonian, with the relative motion part given by ($r_{ij}=r$)
$$
\hat{H}_r=-\nabla^2_r + \frac{1}{4}\omega^2r^2+ \frac{1}{r},
$$
plus a standard Harmonic Oscillator problem for the center-of-mass motion.
This system has analytical solutions in two and three dimensions ([M. Taut 1993 and 1994](https://journals.aps.org/pra/abstract/10.1103/PhysRevA.48.3561)).
We want to perform a Variational Monte Carlo calculation of the ground state of two electrons in a quantum dot well with different oscillator energies, assuming total spin $S=0$.
Our trial wave function has the following form
<!-- Equation labels as ordinary links -->
<div id="eq:trial"></div>
$$
\begin{equation}
\psi_{T}(\boldsymbol{r}_1,\boldsymbol{r}_2) =
C\exp{\left(-\alpha_1\omega(r_1^2+r_2^2)/2\right)}
\exp{\left(\frac{r_{12}}{(1+\alpha_2 r_{12})}\right)},
\label{eq:trial} \tag{3}
\end{equation}
$$
where the $\alpha$s represent our variational parameters, two in this case.
Why does the trial function look like this? How did we get there?
**This will be one of our main motivations** for switching to Machine Learning later.
To find an ansatz for the correlated part of the wave function, it is
useful to rewrite the two-particle local energy in terms of the
relative and center-of-mass motion.
Let us denote the distance
between the two electrons as $r_{12}$. We omit the center-of-mass
motion since we are only interested in the case when $r_{12}
\rightarrow 0$. The contribution from the center-of-mass (CoM)
variable $\boldsymbol{R}_{\mathrm{CoM}}$ gives only a finite contribution. We
focus only on the terms that are relevant for $r_{12}$ and for three
dimensions.
The relevant local energy becomes then
$$
\lim_{r_{12} \rightarrow 0}E_L(R)= \frac{1}{{\calR}_T(r_{12})}\left(2\frac{d^2}{dr_{ij}^2}+\frac{4}{r_{ij}}\frac{d}{dr_{ij}}+\frac{2}{r_{ij}}-\frac{l(l+1)}{r_{ij}^2}+2E \right){\cal R}_T(r_{12})
= 0.
$$
Set $l=0$ and we have the so-called **cusp** condition
$$
\frac{d {\cal R}_T(r_{12})}{dr_{12}} = -\frac{1}{2(l+1)} {\cal R}_T(r_{12})\qquad r_{12}\to 0
$$
The above results in
$$
{\cal R}_T \propto \exp{(r_{ij}/2)},
$$
for anti-parallel spins and
$$
{\cal R}_T \propto \exp{(r_{ij}/4)},
$$
for anti-parallel spins.
This is the so-called cusp condition for the relative motion, resulting in a minimal requirement
for the correlation part of the wave fuction.
For general systems containing more than say two electrons, we have this
condition for each electron pair $ij$.
### First code attempt for the two-electron case
First, as with the hydrogen case, we declare where to store files.
```
# Common imports
import os
# Where to save the figures and data files
PROJECT_ROOT_DIR = "Results"
FIGURE_ID = "Results/FigureFiles"
DATA_ID = "Results/VMCQdotMetropolis"
if not os.path.exists(PROJECT_ROOT_DIR):
os.mkdir(PROJECT_ROOT_DIR)
if not os.path.exists(FIGURE_ID):
os.makedirs(FIGURE_ID)
if not os.path.exists(DATA_ID):
os.makedirs(DATA_ID)
def image_path(fig_id):
return os.path.join(FIGURE_ID, fig_id)
def data_path(dat_id):
return os.path.join(DATA_ID, dat_id)
def save_fig(fig_id):
plt.savefig(image_path(fig_id) + ".png", format='png')
outfile = open(data_path("VMCQdotMetropolis.dat"),'w')
```
Thereafter we set up the analytical expressions for the wave functions and the local energy
```
# 2-electron VMC for quantum dot system in two dimensions
# Brute force Metropolis, no importance sampling and no energy minimization
from math import exp, sqrt
from random import random, seed
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from matplotlib import cm
from matplotlib.ticker import LinearLocator, FormatStrFormatter
import sys
from numba import jit
# Trial wave function for the 2-electron quantum dot in two dims
def WaveFunction(r,alpha,beta):
r1 = r[0,0]**2 + r[0,1]**2
r2 = r[1,0]**2 + r[1,1]**2
r12 = sqrt((r[0,0]-r[1,0])**2 + (r[0,1]-r[1,1])**2)
deno = r12/(1+beta*r12)
return exp(-0.5*alpha*(r1+r2)+deno)
# Local energy for the 2-electron quantum dot in two dims, using analytical local energy
def LocalEnergy(r,alpha,beta):
r1 = (r[0,0]**2 + r[0,1]**2)
r2 = (r[1,0]**2 + r[1,1]**2)
r12 = sqrt((r[0,0]-r[1,0])**2 + (r[0,1]-r[1,1])**2)
deno = 1.0/(1+beta*r12)
deno2 = deno*deno
return 0.5*(1-alpha*alpha)*(r1 + r2) +2.0*alpha + 1.0/r12+deno2*(alpha*r12-deno2+2*beta*deno-1.0/r12)
```
The Monte Carlo sampling without importance sampling is set up here.
```
# The Monte Carlo sampling with the Metropolis algo
# The jit decorator tells Numba to compile this function.
# The argument types will be inferred by Numba when the function is called.
@jit
def MonteCarloSampling():
NumberMCcycles= 10000
StepSize = 1.0
# positions
PositionOld = np.zeros((NumberParticles,Dimension), np.double)
PositionNew = np.zeros((NumberParticles,Dimension), np.double)
# seed for rng generator
seed()
# start variational parameter
alpha = 0.9
for ia in range(MaxVariations):
alpha += .025
AlphaValues[ia] = alpha
beta = 0.2
for jb in range(MaxVariations):
beta += .01
BetaValues[jb] = beta
energy = energy2 = 0.0
DeltaE = 0.0
#Initial position
for i in range(NumberParticles):
for j in range(Dimension):
PositionOld[i,j] = StepSize * (random() - .5)
wfold = WaveFunction(PositionOld,alpha,beta)
#Loop over MC MCcycles
for MCcycle in range(NumberMCcycles):
#Trial position moving one particle at the time
for i in range(NumberParticles):
for j in range(Dimension):
PositionNew[i,j] = PositionOld[i,j] + StepSize * (random() - .5)
wfnew = WaveFunction(PositionNew,alpha,beta)
#Metropolis test to see whether we accept the move
if random() < wfnew**2 / wfold**2:
for j in range(Dimension):
PositionOld[i,j] = PositionNew[i,j]
wfold = wfnew
DeltaE = LocalEnergy(PositionOld,alpha,beta)
energy += DeltaE
energy2 += DeltaE**2
#We calculate mean, variance and error ...
energy /= NumberMCcycles
energy2 /= NumberMCcycles
variance = energy2 - energy**2
error = sqrt(variance/NumberMCcycles)
Energies[ia,jb] = energy
Variances[ia,jb] = variance
outfile.write('%f %f %f %f %f\n' %(alpha,beta,energy,variance,error))
return Energies, Variances, AlphaValues, BetaValues
```
And finally comes the main part with the plots as well.
```
#Here starts the main program with variable declarations
NumberParticles = 2
Dimension = 2
MaxVariations = 10
Energies = np.zeros((MaxVariations,MaxVariations))
Variances = np.zeros((MaxVariations,MaxVariations))
AlphaValues = np.zeros(MaxVariations)
BetaValues = np.zeros(MaxVariations)
(Energies, Variances, AlphaValues, BetaValues) = MonteCarloSampling()
outfile.close()
# Prepare for plots
fig = plt.figure()
ax = fig.gca(projection='3d')
# Plot the surface.
X, Y = np.meshgrid(AlphaValues, BetaValues)
surf = ax.plot_surface(X, Y, Energies,cmap=cm.coolwarm,linewidth=0, antialiased=False)
# Customize the z axis.
zmin = np.matrix(Energies).min()
zmax = np.matrix(Energies).max()
ax.set_zlim(zmin, zmax)
ax.set_xlabel(r'$\alpha$')
ax.set_ylabel(r'$\beta$')
ax.set_zlabel(r'$\langle E \rangle$')
ax.zaxis.set_major_locator(LinearLocator(10))
ax.zaxis.set_major_formatter(FormatStrFormatter('%.02f'))
# Add a color bar which maps values to colors.
fig.colorbar(surf, shrink=0.5, aspect=5)
save_fig("QdotMetropolis")
plt.show()
```
## Importance sampling
The above way of performing a Monte Carlo calculation is not the most efficient one.
We need to replace the brute force Metropolis algorithm with a walk in
coordinate space biased by the trial wave function. This approach is
based on the Fokker-Planck equation and the Langevin equation for
generating a trajectory in coordinate space. The link between the
Fokker-Planck equation and the Langevin equations are explained, only
partly, in the slides below. An excellent reference on topics like
Brownian motion, Markov chains, the Fokker-Planck equation and the
Langevin equation is the text by [Van Kampen](http://www.elsevier.com/books/stochastic-processes-in-physics-and-chemistry/van-kampen/978-0-444-52965-7)
Here we will focus first on the implementation part first.
For a diffusion process characterized by a time-dependent probability density $P(x,t)$ in one dimension the Fokker-Planck
equation reads (for one particle /walker)
$$
\frac{\partial P}{\partial t} = D\frac{\partial }{\partial x}\left(\frac{\partial }{\partial x} -F\right)P(x,t),
$$
where $F$ is a drift term and $D$ is the diffusion coefficient.
The new positions in coordinate space are given as the solutions of the Langevin equation using Euler's method, namely,
we go from the Langevin equation
$$
\frac{\partial x(t)}{\partial t} = DF(x(t)) +\eta,
$$
with $\eta$ a random variable,
yielding a new position
$$
y = x+DF(x)\Delta t +\xi\sqrt{\Delta t},
$$
where $\xi$ is gaussian random variable and $\Delta t$ is a chosen time step.
The quantity $D$ is, in atomic units, equal to $1/2$ and comes from the factor $1/2$ in the kinetic energy operator. Note that $\Delta t$ is to be viewed as a parameter. Values of $\Delta t \in [0.001,0.01]$ yield in general rather stable values of the ground state energy.
The process of isotropic diffusion characterized by a time-dependent probability density $P(\mathbf{x},t)$ obeys (as an approximation) the so-called Fokker-Planck equation
$$
\frac{\partial P}{\partial t} = \sum_i D\frac{\partial }{\partial \mathbf{x_i}}\left(\frac{\partial }{\partial \mathbf{x_i}} -\mathbf{F_i}\right)P(\mathbf{x},t),
$$
where $\mathbf{F_i}$ is the $i^{th}$ component of the drift term (drift velocity) caused by an external potential, and $D$ is the diffusion coefficient. The convergence to a stationary probability density can be obtained by setting the left hand side to zero. The resulting equation will be satisfied if and only if all the terms of the sum are equal zero,
$$
\frac{\partial^2 P}{\partial {\mathbf{x_i}^2}} = P\frac{\partial}{\partial {\mathbf{x_i}}}\mathbf{F_i} + \mathbf{F_i}\frac{\partial}{\partial {\mathbf{x_i}}}P.
$$
The drift vector should be of the form $\mathbf{F} = g(\mathbf{x}) \frac{\partial P}{\partial \mathbf{x}}$. Then,
$$
\frac{\partial^2 P}{\partial {\mathbf{x_i}^2}} = P\frac{\partial g}{\partial P}\left( \frac{\partial P}{\partial {\mathbf{x}_i}} \right)^2 + P g \frac{\partial ^2 P}{\partial {\mathbf{x}_i^2}} + g \left( \frac{\partial P}{\partial {\mathbf{x}_i}} \right)^2.
$$
The condition of stationary density means that the left hand side equals zero. In other words, the terms containing first and second derivatives have to cancel each other. It is possible only if $g = \frac{1}{P}$, which yields
$$
\mathbf{F} = 2\frac{1}{\Psi_T}\nabla\Psi_T,
$$
which is known as the so-called *quantum force*. This term is responsible for pushing the walker towards regions of configuration space where the trial wave function is large, increasing the efficiency of the simulation in contrast to the Metropolis algorithm where the walker has the same probability of moving in every direction.
The Fokker-Planck equation yields a (the solution to the equation) transition probability given by the Green's function
$$
G(y,x,\Delta t) = \frac{1}{(4\pi D\Delta t)^{3N/2}} \exp{\left(-(y-x-D\Delta t F(x))^2/4D\Delta t\right)}
$$
which in turn means that our brute force Metropolis algorithm
$$
A(y,x) = \mathrm{min}(1,q(y,x))),
$$
with $q(y,x) = |\Psi_T(y)|^2/|\Psi_T(x)|^2$ is now replaced by the [Metropolis-Hastings algorithm](http://scitation.aip.org/content/aip/journal/jcp/21/6/10.1063/1.1699114). See also [Hasting's original article](http://biomet.oxfordjournals.org/content/57/1/97.abstract),
$$
q(y,x) = \frac{G(x,y,\Delta t)|\Psi_T(y)|^2}{G(y,x,\Delta t)|\Psi_T(x)|^2}
$$
### Code example for the interacting case with importance sampling
We are now ready to implement importance sampling. This is done here for the two-electron case with the Coulomb interaction, as in the previous example. We have two variational parameters $\alpha$ and $\beta$. After the set up of files
```
# Common imports
import os
# Where to save the figures and data files
PROJECT_ROOT_DIR = "Results"
FIGURE_ID = "Results/FigureFiles"
DATA_ID = "Results/VMCQdotImportance"
if not os.path.exists(PROJECT_ROOT_DIR):
os.mkdir(PROJECT_ROOT_DIR)
if not os.path.exists(FIGURE_ID):
os.makedirs(FIGURE_ID)
if not os.path.exists(DATA_ID):
os.makedirs(DATA_ID)
def image_path(fig_id):
return os.path.join(FIGURE_ID, fig_id)
def data_path(dat_id):
return os.path.join(DATA_ID, dat_id)
def save_fig(fig_id):
plt.savefig(image_path(fig_id) + ".png", format='png')
outfile = open(data_path("VMCQdotImportance.dat"),'w')
```
we move on to the set up of the trial wave function, the analytical expression for the local energy and the analytical expression for the quantum force.
```
# 2-electron VMC code for 2dim quantum dot with importance sampling
# Using gaussian rng for new positions and Metropolis- Hastings
# No energy minimization
from math import exp, sqrt
from random import random, seed, normalvariate
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from matplotlib import cm
from matplotlib.ticker import LinearLocator, FormatStrFormatter
import sys
from numba import jit,njit
# Trial wave function for the 2-electron quantum dot in two dims
def WaveFunction(r,alpha,beta):
r1 = r[0,0]**2 + r[0,1]**2
r2 = r[1,0]**2 + r[1,1]**2
r12 = sqrt((r[0,0]-r[1,0])**2 + (r[0,1]-r[1,1])**2)
deno = r12/(1+beta*r12)
return exp(-0.5*alpha*(r1+r2)+deno)
# Local energy for the 2-electron quantum dot in two dims, using analytical local energy
def LocalEnergy(r,alpha,beta):
r1 = (r[0,0]**2 + r[0,1]**2)
r2 = (r[1,0]**2 + r[1,1]**2)
r12 = sqrt((r[0,0]-r[1,0])**2 + (r[0,1]-r[1,1])**2)
deno = 1.0/(1+beta*r12)
deno2 = deno*deno
return 0.5*(1-alpha*alpha)*(r1 + r2) +2.0*alpha + 1.0/r12+deno2*(alpha*r12-deno2+2*beta*deno-1.0/r12)
# Setting up the quantum force for the two-electron quantum dot, recall that it is a vector
def QuantumForce(r,alpha,beta):
qforce = np.zeros((NumberParticles,Dimension), np.double)
r12 = sqrt((r[0,0]-r[1,0])**2 + (r[0,1]-r[1,1])**2)
deno = 1.0/(1+beta*r12)
qforce[0,:] = -2*r[0,:]*alpha*(r[0,:]-r[1,:])*deno*deno/r12
qforce[1,:] = -2*r[1,:]*alpha*(r[1,:]-r[0,:])*deno*deno/r12
return qforce
```
The Monte Carlo sampling includes now the Metropolis-Hastings algorithm, with the additional complication of having to evaluate the **quantum force** and the Green's function which is the solution of the Fokker-Planck equation.
```
# The Monte Carlo sampling with the Metropolis algo
# jit decorator tells Numba to compile this function.
# The argument types will be inferred by Numba when function is called.
@jit()
def MonteCarloSampling():
NumberMCcycles= 100000
# Parameters in the Fokker-Planck simulation of the quantum force
D = 0.5
TimeStep = 0.05
# positions
PositionOld = np.zeros((NumberParticles,Dimension), np.double)
PositionNew = np.zeros((NumberParticles,Dimension), np.double)
# Quantum force
QuantumForceOld = np.zeros((NumberParticles,Dimension), np.double)
QuantumForceNew = np.zeros((NumberParticles,Dimension), np.double)
# seed for rng generator
seed()
# start variational parameter loops, two parameters here
alpha = 0.9
for ia in range(MaxVariations):
alpha += .025
AlphaValues[ia] = alpha
beta = 0.2
for jb in range(MaxVariations):
beta += .01
BetaValues[jb] = beta
energy = energy2 = 0.0
DeltaE = 0.0
#Initial position
for i in range(NumberParticles):
for j in range(Dimension):
PositionOld[i,j] = normalvariate(0.0,1.0)*sqrt(TimeStep)
wfold = WaveFunction(PositionOld,alpha,beta)
QuantumForceOld = QuantumForce(PositionOld,alpha, beta)
#Loop over MC MCcycles
for MCcycle in range(NumberMCcycles):
#Trial position moving one particle at the time
for i in range(NumberParticles):
for j in range(Dimension):
PositionNew[i,j] = PositionOld[i,j]+normalvariate(0.0,1.0)*sqrt(TimeStep)+\
QuantumForceOld[i,j]*TimeStep*D
wfnew = WaveFunction(PositionNew,alpha,beta)
QuantumForceNew = QuantumForce(PositionNew,alpha, beta)
GreensFunction = 0.0
for j in range(Dimension):
GreensFunction += 0.5*(QuantumForceOld[i,j]+QuantumForceNew[i,j])*\
(D*TimeStep*0.5*(QuantumForceOld[i,j]-QuantumForceNew[i,j])-\
PositionNew[i,j]+PositionOld[i,j])
GreensFunction = exp(GreensFunction)
ProbabilityRatio = GreensFunction*wfnew**2/wfold**2
#Metropolis-Hastings test to see whether we accept the move
if random() <= ProbabilityRatio:
for j in range(Dimension):
PositionOld[i,j] = PositionNew[i,j]
QuantumForceOld[i,j] = QuantumForceNew[i,j]
wfold = wfnew
DeltaE = LocalEnergy(PositionOld,alpha,beta)
energy += DeltaE
energy2 += DeltaE**2
# We calculate mean, variance and error (no blocking applied)
energy /= NumberMCcycles
energy2 /= NumberMCcycles
variance = energy2 - energy**2
error = sqrt(variance/NumberMCcycles)
Energies[ia,jb] = energy
outfile.write('%f %f %f %f %f\n' %(alpha,beta,energy,variance,error))
return Energies, AlphaValues, BetaValues
```
The main part here contains the setup of the variational parameters, the energies and the variance.
```
#Here starts the main program with variable declarations
NumberParticles = 2
Dimension = 2
MaxVariations = 10
Energies = np.zeros((MaxVariations,MaxVariations))
AlphaValues = np.zeros(MaxVariations)
BetaValues = np.zeros(MaxVariations)
(Energies, AlphaValues, BetaValues) = MonteCarloSampling()
outfile.close()
# Prepare for plots
fig = plt.figure()
ax = fig.gca(projection='3d')
# Plot the surface.
X, Y = np.meshgrid(AlphaValues, BetaValues)
surf = ax.plot_surface(X, Y, Energies,cmap=cm.coolwarm,linewidth=0, antialiased=False)
# Customize the z axis.
zmin = np.matrix(Energies).min()
zmax = np.matrix(Energies).max()
ax.set_zlim(zmin, zmax)
ax.set_xlabel(r'$\alpha$')
ax.set_ylabel(r'$\beta$')
ax.set_zlabel(r'$\langle E \rangle$')
ax.zaxis.set_major_locator(LinearLocator(10))
ax.zaxis.set_major_formatter(FormatStrFormatter('%.02f'))
# Add a color bar which maps values to colors.
fig.colorbar(surf, shrink=0.5, aspect=5)
save_fig("QdotImportance")
plt.show()
```
## Technical aspects, improvements and how to define the cost function
The above procedure is also not the smartest one. Looping over all
variational parameters becomes expensive and we see from the previous
plot that the surface is not very smooth, indicating that we need many more
Monte Carlo cycles in order to reliably define an energy minimum.
What we can do however is to perform some preliminary calculations
with selected variational parameters (normally with less Monte Carlo
cycles than those used in a full production calculation). For every
step we evaluate the derivatives of the energy as functions of the
variational parameters. When the derivatives disappear we have
hopefully reached the global minimum.
At this point we have the optimal variational parameters and can start
our large-scale production run. To find the optimal parameters
entails the computation of the gradients of the energy and
optimization algorithms like various **gradient descent** methods.
This is an art by itself and is discussed for example in [our lectures on optimization methods](http://compphysics.github.io/ComputationalPhysics2/doc/pub/cg/html/cg.html). We refer the reader to these notes for more details.
This part allows us also to link with the true working horse of every
Machine Learning algorithm, namely the optimization part. This
normally involves one of the stochastic gradient descent algorithms
discussed in the above lecture notes. We will come back to these topics in the second notebook.
In order to apply these optmization algortihms we anticipate partly what is to come in notebook 2 on
Boltzmann machines. Our cost (or loss) function is here given by the
expectation value of the energy as function of the variational
parameters.
To find the derivatives of the local energy expectation value as
function of the variational parameters, we can use the chain rule and
the hermiticity of the Hamiltonian.
Let us define
$$
\bar{E}_{\alpha_i}=\frac{d\langle E_L\rangle}{d\alpha_i}.
$$
as the derivative of the energy with respect to the variational parameter $\alpha_i$
We define also the derivative of the trial function (skipping the subindex $T$) as
$$
\bar{\Psi}_{i}=\frac{d\Psi}{d\alpha_i}.
$$
The elements of the gradient of the local energy are then (using the
chain rule and the hermiticity of the Hamiltonian)
$$
\bar{E}_{i}=
2\left( \langle \frac{\bar{\Psi}_{i}}{\Psi}E_L\rangle -\langle
\frac{\bar{\Psi}_{i}}{\Psi}\rangle\langle E_L \rangle\right).
$$
From a computational point of view it means that we need to compute
the expectation values of
$$
\langle
\frac{\bar{\Psi}_{i}}{\Psi}E_L\rangle,
$$
and
$$
\langle
\frac{\bar{\Psi}_{i}}{\Psi}\rangle\langle E_L\rangle
$$
These integrals are evaluted using MC intergration (with all its possible
error sources). We can then use methods like stochastic gradient or
other minimization methods to find the optimal variational parameters
As an alternative to the energy as cost function, we could use the variance as the cost function.
As discussed earlier, if we have the exact wave function, the variance is exactly equal to zero.
Suppose the trial function (our model) is the exact wave function.
The variance is defined as
$$
\sigma_E = \langle E^2\rangle - \langle E\rangle^2.
$$
Some practitioners perform Monte Carlo calculations by minimizing both the energy and the variance.
In order to minimize the variance we need the derivatives of
$$
\sigma_E = \langle E^2\rangle - \langle E\rangle^2,
$$
with respect to the variational parameters. The derivatives of the variance can then be used to defined the
so-called Hessian matrix, which in turn allows us to use minimization methods like Newton's method or
standard gradient methods.
This leads to however a more complicated expression, with obvious errors when evaluating many more integrals by Monte Carlo integration. It is normally less used, see however [Filippi and Umrigar](https://journals.aps.org/prl/abstract/10.1103/PhysRevLett.94.150201). The expression becomes complicated
$$
\bar{E}_{ij} = 2\left[ \langle (\frac{\bar{\Psi}_{ij}}{\Psi}+\frac{\bar{\Psi}_{j}}{\Psi}\frac{\bar{\Psi}_{i}}{\Psi})(E_L-\langle E\rangle)\rangle -\langle \frac{\bar{\Psi}_{i}}{\Psi}\rangle\bar{E}_j-\langle \frac{\bar{\Psi}_{j}}{\Psi}\rangle\bar{E}_i\right] +\langle \frac{\bar{\Psi}_{i}}{\Psi}E_L{_j}\rangle +\langle \frac{\bar{\Psi}_{j}}{\Psi}E_L{_i}\rangle -\langle \frac{\bar{\Psi}_{i}}{\Psi}\rangle\langle E_L{_j}\rangle \langle \frac{\bar{\Psi}_{j}}{\Psi}\rangle\langle E_L{_i}\rangle.
$$
Evaluating the cost function means having to evaluate the above second derivative of the energy.
Before we proceed with code examples, let us look at some simple examples, here the one-particle harmonic oscillator in one dimension. This serves as a very useful check when developing a code. The first code discussed is the two-dimensional non-interacting harmonic oscillator.
### Simple example
Let us illustrate what is needed in our calculations using a simple
example, the harmonic oscillator in one dimension. For the harmonic
oscillator in one-dimension we have a trial wave function and
probability
$$
\psi_T(x) = e^{-\alpha^2 x^2} \qquad P_T(x)dx = \frac{e^{-2\alpha^2 x^2}dx}{\int dx e^{-2\alpha^2 x^2}}
$$
with $\alpha$ being the variational parameter.
We obtain then the following local energy
$$
E_L[\alpha] = \alpha^2+x^2\left(\frac{1}{2}-2\alpha^2\right),
$$
which results in the expectation value for the local energy
$$
\langle E_L[\alpha]\rangle = \frac{1}{2}\alpha^2+\frac{1}{8\alpha^2}
$$
The derivative of the energy with respect to $\alpha$ gives
$$
\frac{d\langle E_L[\alpha]\rangle}{d\alpha} = \alpha-\frac{1}{4\alpha^3}
$$
and a second derivative which is always positive (meaning that we find a minimum)
$$
\frac{d^2\langle E_L[\alpha]\rangle}{d\alpha^2} = 1+\frac{3}{4\alpha^4}
$$
The condition
$$
\frac{d\langle E_L[\alpha]\rangle}{d\alpha} = 0,
$$
gives the optimal $\alpha=1/\sqrt{2}$, as expected.
We can also minimize the variance. In our simple model the variance is
$$
\sigma^2[\alpha] = \frac{1}{2}\alpha^4-\frac{1}{4}+\frac{1}{32\alpha^4},
$$
with first derivative
$$
\frac{d \sigma^2[\alpha]}{d\alpha} = 2\alpha^3-\frac{1}{8\alpha^5}
$$
and a second derivative which is always positive (as expected for a convex function)
$$
\frac{d^2\sigma^2[\alpha]}{d\alpha^2} = 6\alpha^2+\frac{5}{8\alpha^6}
$$
In general we end up computing the expectation value of the energy in
terms of some parameters $\alpha_0,\alpha_1,\dots,\alpha_n$ and we
search for a minimum in this multi-variable parameter space. This
leads to an energy minimization problem *where we need the derivative
of the energy as a function of the variational parameters*.
In the above example this was easy and we were able to find the
expression for the derivative by simple derivations. However, in our
actual calculations the energy is represented by a multi-dimensional
integral with several variational parameters.
### Finding the minima
Perhaps the most celebrated of all one-dimensional root-finding
routines is Newton's method, also called the Newton-Raphson
method. This method requires the evaluation of both the
function $f$ and its derivative $f'$ at arbitrary points.
If you can only calculate the derivative
numerically and/or your function is not of the smooth type, we
normally discourage the use of this method.
The Newton-Raphson formula consists geometrically of extending the
tangent line at a current point until it crosses zero, then setting
the next guess to the abscissa of that zero-crossing. The mathematics
behind this method is rather simple. Employing a Taylor expansion for
$x$ sufficiently close to the solution $s$, we have
<!-- Equation labels as ordinary links -->
<div id="eq:taylornr"></div>
$$
f(s)=0=f(x)+(s-x)f'(x)+\frac{(s-x)^2}{2}f''(x) +\dots.
\label{eq:taylornr} \tag{4}
$$
For small enough values of the function and for well-behaved
functions, the terms beyond linear are unimportant, hence we obtain
$$
f(x)+(s-x)f'(x)\approx 0,
$$
yielding
$$
s\approx x-\frac{f(x)}{f'(x)}.
$$
Having in mind an iterative procedure, it is natural to start iterating with
$$
x_{n+1}=x_n-\frac{f(x_n)}{f'(x_n)}.
$$
The above is Newton-Raphson's method. It has a simple geometric
interpretation, namely $x_{n+1}$ is the point where the tangent from
$(x_n,f(x_n))$ crosses the $x$-axis. Close to the solution,
Newton-Raphson converges fast to the desired result. However, if we
are far from a root, where the higher-order terms in the series are
important, the Newton-Raphson formula can give grossly inaccurate
results. For instance, the initial guess for the root might be so far
from the true root as to let the search interval include a local
maximum or minimum of the function. If an iteration places a trial
guess near such a local extremum, so that the first derivative nearly
vanishes, then Newton-Raphson may fail totally
Newton's method can be generalized to systems of several non-linear equations
and variables. Consider the case with two equations
$$
\begin{array}{cc} f_1(x_1,x_2) &=0\\
f_2(x_1,x_2) &=0,\end{array}
$$
which we Taylor expand to obtain
$$
\begin{array}{cc} 0=f_1(x_1+h_1,x_2+h_2)=&f_1(x_1,x_2)+h_1
\partial f_1/\partial x_1+h_2
\partial f_1/\partial x_2+\dots\\
0=f_2(x_1+h_1,x_2+h_2)=&f_2(x_1,x_2)+h_1
\partial f_2/\partial x_1+h_2
\partial f_2/\partial x_2+\dots
\end{array}.
$$
Defining the Jacobian matrix $\hat{J}$ we have
$$
\hat{J}=\left( \begin{array}{cc}
\partial f_1/\partial x_1 & \partial f_1/\partial x_2 \\
\partial f_2/\partial x_1 &\partial f_2/\partial x_2
\end{array} \right),
$$
we can rephrase Newton's method as
$$
\left(\begin{array}{c} x_1^{n+1} \\ x_2^{n+1} \end{array} \right)=
\left(\begin{array}{c} x_1^{n} \\ x_2^{n} \end{array} \right)+
\left(\begin{array}{c} h_1^{n} \\ h_2^{n} \end{array} \right),
$$
where we have defined
$$
\left(\begin{array}{c} h_1^{n} \\ h_2^{n} \end{array} \right)=
-{\bf \hat{J}}^{-1}
\left(\begin{array}{c} f_1(x_1^{n},x_2^{n}) \\ f_2(x_1^{n},x_2^{n}) \end{array} \right).
$$
We need thus to compute the inverse of the Jacobian matrix and it
is to understand that difficulties may
arise in case $\hat{J}$ is nearly singular.
It is rather straightforward to extend the above scheme to systems of
more than two non-linear equations. In our case, the Jacobian matrix is given by the Hessian that represents the second derivative of the cost function.
If we are able to evaluate the second derivative of the energy with
respect to the variational parameters, we can also set up the Hessian
matrix. However, as we saw earlier, the second derivative of the
energy with respect to these parameters involves the evaluation of
more complicated integrals, leading in turn to more statistical
errors.
This means that we normally try to avoid evaluating the second derivative and use rather simpler methods like
the gradient descent family of methods.
### Steepest descent
The basic idea of gradient descent is
that a function $F(\mathbf{x})$,
$\mathbf{x} \equiv (x_1,\cdots,x_n)$, decreases fastest if one goes from $\bf {x}$ in the
direction of the negative gradient $-\nabla F(\mathbf{x})$.
It can be shown that if
$$
\mathbf{x}_{k+1} = \mathbf{x}_k - \gamma_k \nabla F(\mathbf{x}_k),
$$
with $\gamma_k > 0$.
For $\gamma_k$ small enough, then $F(\mathbf{x}_{k+1}) \leq
F(\mathbf{x}_k)$. This means that for a sufficiently small $\gamma_k$
we are always moving towards smaller function values, i.e a minimum.
The previous observation is the basis of the method of steepest
descent, which is also referred to as just gradient descent (GD). One
starts with an initial guess $\mathbf{x}_0$ for a minimum of $F$ and
computes new approximations according to
$$
\mathbf{x}_{k+1} = \mathbf{x}_k - \gamma_k \nabla F(\mathbf{x}_k), \ \ k \geq 0.
$$
The parameter $\gamma_k$ is often referred to as the step length or
the learning rate within the context of Machine Learning.
Ideally the sequence $\{\mathbf{x}_k \}_{k=0}$ converges to a global
minimum of the function $F$. In general we do not know if we are in a
global or local minimum. In the special case when $F$ is a convex
function, all local minima are also global minima, so in this case
gradient descent can converge to the global solution. The advantage of
this scheme is that it is conceptually simple and straightforward to
implement. However the method in this form has some severe
limitations:
In machine learing we are often faced with non-convex high dimensional
cost functions with many local minima. Since GD is deterministic we
will get stuck in a local minimum, if the method converges, unless we
have a very good intial guess. This also implies that the scheme is
sensitive to the chosen initial condition.
Note that the gradient is a function of $\mathbf{x} =
(x_1,\cdots,x_n)$ which makes it expensive to compute numerically.
The gradient descent method
is sensitive to the choice of learning rate $\gamma_k$. This is due
to the fact that we are only guaranteed that $F(\mathbf{x}_{k+1}) \leq
F(\mathbf{x}_k)$ for sufficiently small $\gamma_k$. The problem is to
determine an optimal learning rate. If the learning rate is chosen too
small the method will take a long time to converge and if it is too
large we can experience erratic behavior.
Many of these shortcomings can be alleviated by introducing
randomness. One such method is that of Stochastic Gradient Descent
(SGD). This is not discussed in this notebook.
### The code for two electrons in two dims with no Coulomb interaction
We present here the code (including importance sampling) for finding the optimal parameter $\alpha$ using gradient descent with a given learning rate $\eta$. In principle we should run calculations for various learning rates.
Again, we start first with set up of various files.
```
# 2-electron VMC code for 2dim quantum dot with importance sampling
# No Coulomb interaction
# Using gaussian rng for new positions and Metropolis- Hastings
# Energy minimization using standard gradient descent
# Common imports
import os
# Where to save the figures and data files
PROJECT_ROOT_DIR = "Results"
FIGURE_ID = "Results/FigureFiles"
if not os.path.exists(PROJECT_ROOT_DIR):
os.mkdir(PROJECT_ROOT_DIR)
if not os.path.exists(FIGURE_ID):
os.makedirs(FIGURE_ID)
def image_path(fig_id):
return os.path.join(FIGURE_ID, fig_id)
def save_fig(fig_id):
plt.savefig(image_path(fig_id) + ".png", format='png')
```
Thereafter we define the wave function, the local energy and the quantum force.
We include also the derivative of the wave function as function of the variational parameter $\alpha$.
```
from math import exp, sqrt
from random import random, seed, normalvariate
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from matplotlib import cm
from matplotlib.ticker import LinearLocator, FormatStrFormatter
import sys
from numba import jit
from scipy.optimize import minimize
# Trial wave function for the 2-electron quantum dot in two dims
def WaveFunction(r,alpha):
r1 = r[0,0]**2 + r[0,1]**2
r2 = r[1,0]**2 + r[1,1]**2
return exp(-0.5*alpha*(r1+r2))
# Local energy for the 2-electron quantum dot in two dims, using analytical local energy
def LocalEnergy(r,alpha):
r1 = (r[0,0]**2 + r[0,1]**2)
r2 = (r[1,0]**2 + r[1,1]**2)
return 0.5*(1-alpha*alpha)*(r1 + r2) +2.0*alpha
# Derivate of wave function ansatz as function of variational parameters
def DerivativeWFansatz(r,alpha):
r1 = (r[0,0]**2 + r[0,1]**2)
r2 = (r[1,0]**2 + r[1,1]**2)
WfDer = -0.5*(r1+r2)
return WfDer
# Setting up the quantum force for the two-electron quantum dot, recall that it is a vector
def QuantumForce(r,alpha):
qforce = np.zeros((NumberParticles,Dimension), np.double)
qforce[0,:] = -2*r[0,:]*alpha
qforce[1,:] = -2*r[1,:]*alpha
return qforce
```
Then comes our Monte Carlo sampling.
```
# Computing the derivative of the energy and the energy
# jit decorator tells Numba to compile this function.
# The argument types will be inferred by Numba when function is called.
@jit
def EnergyMinimization(alpha):
NumberMCcycles= 1000
# Parameters in the Fokker-Planck simulation of the quantum force
D = 0.5
TimeStep = 0.05
# positions
PositionOld = np.zeros((NumberParticles,Dimension), np.double)
PositionNew = np.zeros((NumberParticles,Dimension), np.double)
# Quantum force
QuantumForceOld = np.zeros((NumberParticles,Dimension), np.double)
QuantumForceNew = np.zeros((NumberParticles,Dimension), np.double)
# seed for rng generator
seed()
energy = 0.0
DeltaE = 0.0
EnergyDer = 0.0
DeltaPsi = 0.0
DerivativePsiE = 0.0
#Initial position
for i in range(NumberParticles):
for j in range(Dimension):
PositionOld[i,j] = normalvariate(0.0,1.0)*sqrt(TimeStep)
wfold = WaveFunction(PositionOld,alpha)
QuantumForceOld = QuantumForce(PositionOld,alpha)
#Loop over MC MCcycles
for MCcycle in range(NumberMCcycles):
#Trial position moving one particle at the time
for i in range(NumberParticles):
for j in range(Dimension):
PositionNew[i,j] = PositionOld[i,j]+normalvariate(0.0,1.0)*sqrt(TimeStep)+\
QuantumForceOld[i,j]*TimeStep*D
wfnew = WaveFunction(PositionNew,alpha)
QuantumForceNew = QuantumForce(PositionNew,alpha)
GreensFunction = 0.0
for j in range(Dimension):
GreensFunction += 0.5*(QuantumForceOld[i,j]+QuantumForceNew[i,j])*\
(D*TimeStep*0.5*(QuantumForceOld[i,j]-QuantumForceNew[i,j])-\
PositionNew[i,j]+PositionOld[i,j])
GreensFunction = 1.0#exp(GreensFunction)
ProbabilityRatio = GreensFunction*wfnew**2/wfold**2
#Metropolis-Hastings test to see whether we accept the move
if random() <= ProbabilityRatio:
for j in range(Dimension):
PositionOld[i,j] = PositionNew[i,j]
QuantumForceOld[i,j] = QuantumForceNew[i,j]
wfold = wfnew
DeltaE = LocalEnergy(PositionOld,alpha)
DerPsi = DerivativeWFansatz(PositionOld,alpha)
DeltaPsi +=DerPsi
energy += DeltaE
DerivativePsiE += DerPsi*DeltaE
# We calculate mean values
energy /= NumberMCcycles
DerivativePsiE /= NumberMCcycles
DeltaPsi /= NumberMCcycles
EnergyDer = 2*(DerivativePsiE-DeltaPsi*energy)
return energy, EnergyDer
```
Finally, here we use the gradient descent method with a fixed learning rate and a fixed number of iterations.
This code is meant for illustrative purposes only. We could for example add a test which stops the number of
terations when the derivative has reached a certain by us fixed minimal value.
```
#Here starts the main program with variable declarations
NumberParticles = 2
Dimension = 2
# guess for variational parameters
x0 = 0.5
# Set up iteration using stochastic gradient method
Energy =0 ; EnergyDer = 0
Energy, EnergyDer = EnergyMinimization(x0)
# No adaptive search for a minimum
eta = 2.0
Niterations = 50
Energies = np.zeros(Niterations)
EnergyDerivatives = np.zeros(Niterations)
AlphaValues = np.zeros(Niterations)
Totiterations = np.zeros(Niterations)
for iter in range(Niterations):
gradients = EnergyDer
x0 -= eta*gradients
Energy, EnergyDer = EnergyMinimization(x0)
Energies[iter] = Energy
EnergyDerivatives[iter] = EnergyDer
AlphaValues[iter] = x0
Totiterations[iter] = iter
plt.subplot(2, 1, 1)
plt.plot(Totiterations, Energies, 'o-')
plt.title('Energy and energy derivatives')
plt.ylabel('Dimensionless energy')
plt.subplot(2, 1, 2)
plt.plot(Totiterations, EnergyDerivatives, '.-')
plt.xlabel(r'$\mathrm{Iterations}$', fontsize=15)
plt.ylabel('Energy derivative')
save_fig("QdotNonint")
plt.show()
#nice printout with Pandas
import pandas as pd
from pandas import DataFrame
data ={'Alpha':AlphaValues, 'Energy':Energies,'Derivative':EnergyDerivatives}
frame = pd.DataFrame(data)
print(frame)
```
We see that the first derivative becomes smaller and smaller and after
some forty iterations, it is for all practical purposes almost
vanishing. The exact energy is $2.0$ and the optimal variational
parameter is $1.0$, as it should.
Next, we extend the above code to include the Coulomb interaction and the Jastrow factor as well. This is done here.
```
# 2-electron VMC code for 2dim quantum dot with importance sampling
# Using gaussian rng for new positions and Metropolis- Hastings
# Added energy minimization
# Common imports
from math import exp, sqrt
from random import random, seed, normalvariate
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from matplotlib import cm
from matplotlib.ticker import LinearLocator, FormatStrFormatter
import sys
# Trial wave function for the 2-electron quantum dot in two dims
def WaveFunction(r,alpha,beta):
r1 = r[0,0]**2 + r[0,1]**2
r2 = r[1,0]**2 + r[1,1]**2
r12 = sqrt((r[0,0]-r[1,0])**2 + (r[0,1]-r[1,1])**2)
deno = r12/(1+beta*r12)
return exp(-0.5*alpha*(r1+r2)+deno)
# Local energy for the 2-electron quantum dot in two dims, using analytical local energy
def LocalEnergy(r,alpha,beta):
r1 = (r[0,0]**2 + r[0,1]**2)
r2 = (r[1,0]**2 + r[1,1]**2)
r12 = sqrt((r[0,0]-r[1,0])**2 + (r[0,1]-r[1,1])**2)
deno = 1.0/(1+beta*r12)
deno2 = deno*deno
return 0.5*(1-alpha*alpha)*(r1 + r2) +2.0*alpha + 1.0/r12+deno2*(alpha*r12-deno2+2*beta*deno-1.0/r12)
# Derivate of wave function ansatz as function of variational parameters
def DerivativeWFansatz(r,alpha,beta):
WfDer = np.zeros((2), np.double)
r1 = (r[0,0]**2 + r[0,1]**2)
r2 = (r[1,0]**2 + r[1,1]**2)
r12 = sqrt((r[0,0]-r[1,0])**2 + (r[0,1]-r[1,1])**2)
deno = 1.0/(1+beta*r12)
deno2 = deno*deno
WfDer[0] = -0.5*(r1+r2)
WfDer[1] = -r12*r12*deno2
return WfDer
# Setting up the quantum force for the two-electron quantum dot, recall that it is a vector
def QuantumForce(r,alpha,beta):
qforce = np.zeros((NumberParticles,Dimension), np.double)
r12 = sqrt((r[0,0]-r[1,0])**2 + (r[0,1]-r[1,1])**2)
deno = 1.0/(1+beta*r12)
qforce[0,:] = -2*r[0,:]*alpha*(r[0,:]-r[1,:])*deno*deno/r12
qforce[1,:] = -2*r[1,:]*alpha*(r[1,:]-r[0,:])*deno*deno/r12
return qforce
# Computing the derivative of the energy and the energy
def EnergyMinimization(alpha, beta):
NumberMCcycles= 10000
# Parameters in the Fokker-Planck simulation of the quantum force
D = 0.5
TimeStep = 0.05
# positions
PositionOld = np.zeros((NumberParticles,Dimension), np.double)
PositionNew = np.zeros((NumberParticles,Dimension), np.double)
# Quantum force
QuantumForceOld = np.zeros((NumberParticles,Dimension), np.double)
QuantumForceNew = np.zeros((NumberParticles,Dimension), np.double)
# seed for rng generator
seed()
energy = 0.0
DeltaE = 0.0
EnergyDer = np.zeros((2), np.double)
DeltaPsi = np.zeros((2), np.double)
DerivativePsiE = np.zeros((2), np.double)
#Initial position
for i in range(NumberParticles):
for j in range(Dimension):
PositionOld[i,j] = normalvariate(0.0,1.0)*sqrt(TimeStep)
wfold = WaveFunction(PositionOld,alpha,beta)
QuantumForceOld = QuantumForce(PositionOld,alpha, beta)
#Loop over MC MCcycles
for MCcycle in range(NumberMCcycles):
#Trial position moving one particle at the time
for i in range(NumberParticles):
for j in range(Dimension):
PositionNew[i,j] = PositionOld[i,j]+normalvariate(0.0,1.0)*sqrt(TimeStep)+\
QuantumForceOld[i,j]*TimeStep*D
wfnew = WaveFunction(PositionNew,alpha,beta)
QuantumForceNew = QuantumForce(PositionNew,alpha, beta)
GreensFunction = 0.0
for j in range(Dimension):
GreensFunction += 0.5*(QuantumForceOld[i,j]+QuantumForceNew[i,j])*\
(D*TimeStep*0.5*(QuantumForceOld[i,j]-QuantumForceNew[i,j])-\
PositionNew[i,j]+PositionOld[i,j])
GreensFunction = exp(GreensFunction)
ProbabilityRatio = GreensFunction*wfnew**2/wfold**2
#Metropolis-Hastings test to see whether we accept the move
if random() <= ProbabilityRatio:
for j in range(Dimension):
PositionOld[i,j] = PositionNew[i,j]
QuantumForceOld[i,j] = QuantumForceNew[i,j]
wfold = wfnew
DeltaE = LocalEnergy(PositionOld,alpha,beta)
DerPsi = DerivativeWFansatz(PositionOld,alpha,beta)
DeltaPsi += DerPsi
energy += DeltaE
DerivativePsiE += DerPsi*DeltaE
# We calculate mean values
energy /= NumberMCcycles
DerivativePsiE /= NumberMCcycles
DeltaPsi /= NumberMCcycles
EnergyDer = 2*(DerivativePsiE-DeltaPsi*energy)
return energy, EnergyDer
#Here starts the main program with variable declarations
NumberParticles = 2
Dimension = 2
# guess for variational parameters
alpha = 0.95
beta = 0.3
# Set up iteration using stochastic gradient method
Energy = 0
EDerivative = np.zeros((2), np.double)
# Learning rate eta, max iterations, need to change to adaptive learning rate
eta = 0.01
MaxIterations = 50
iter = 0
Energies = np.zeros(MaxIterations)
EnergyDerivatives1 = np.zeros(MaxIterations)
EnergyDerivatives2 = np.zeros(MaxIterations)
AlphaValues = np.zeros(MaxIterations)
BetaValues = np.zeros(MaxIterations)
while iter < MaxIterations:
Energy, EDerivative = EnergyMinimization(alpha,beta)
alphagradient = EDerivative[0]
betagradient = EDerivative[1]
alpha -= eta*alphagradient
beta -= eta*betagradient
Energies[iter] = Energy
EnergyDerivatives1[iter] = EDerivative[0]
EnergyDerivatives2[iter] = EDerivative[1]
AlphaValues[iter] = alpha
BetaValues[iter] = beta
iter += 1
#nice printout with Pandas
import pandas as pd
from pandas import DataFrame
pd.set_option('max_columns', 6)
data ={'Alpha':AlphaValues,'Beta':BetaValues,'Energy':Energies,'Alpha Derivative':EnergyDerivatives1,'Beta Derivative':EnergyDerivatives2}
frame = pd.DataFrame(data)
print(frame)
```
The exact energy is $3.0$ for an oscillator frequency $\omega =1$
(with $\hbar =1$). We note however that with this learning rate and
number of iterations, the energies and the derivatives are not yet
converged.
We can improve upon this by using the algorithms provided by the **optimize** package in Python.
One of these algorithms is Broyden–Fletcher–Goldfarb–Shanno (BFGS) algorithm.
The optimization problem is to minimize $f(\mathbf {x} )$ where
$\mathbf {x}$ is a vector in $R^{n}$, and $f$ is a differentiable
scalar function. There are no constraints on the values that $\mathbf{x}$ can take.
The algorithm begins at an initial estimate for the optimal value
$\mathbf {x}_{0}$ and proceeds iteratively to get a better estimate at
each stage.
The search direction $p_k$ at stage $k$ is given by the solution of the analogue of the Newton equation
$$
B_{k}\mathbf {p} _{k}=-\nabla f(\mathbf {x}_{k}),
$$
where $B_{k}$ is an approximation to the Hessian matrix, which is
updated iteratively at each stage, and $\nabla f(\mathbf {x} _{k})$
is the gradient of the function
evaluated at $x_k$.
A line search in the direction $p_k$ is then used to
find the next point $x_{k+1}$ by minimising
$$
f(\mathbf {x}_{k}+\alpha \mathbf {p}_{k}),
$$
over the scalar $\alpha > 0$.
The modified code here uses the BFGS algorithm but performs now a
production run and writes to file all average values of the
energy.
```
# 2-electron VMC code for 2dim quantum dot with importance sampling
# Using gaussian rng for new positions and Metropolis- Hastings
# Added energy minimization
from math import exp, sqrt
from random import random, seed, normalvariate
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from matplotlib import cm
from matplotlib.ticker import LinearLocator, FormatStrFormatter
from scipy.optimize import minimize
import sys
import os
# Where to save data files
PROJECT_ROOT_DIR = "Results"
DATA_ID = "Results/EnergyMin"
if not os.path.exists(PROJECT_ROOT_DIR):
os.mkdir(PROJECT_ROOT_DIR)
if not os.path.exists(DATA_ID):
os.makedirs(DATA_ID)
def data_path(dat_id):
return os.path.join(DATA_ID, dat_id)
outfile = open(data_path("Energies.dat"),'w')
# Trial wave function for the 2-electron quantum dot in two dims
def WaveFunction(r,alpha,beta):
r1 = r[0,0]**2 + r[0,1]**2
r2 = r[1,0]**2 + r[1,1]**2
r12 = sqrt((r[0,0]-r[1,0])**2 + (r[0,1]-r[1,1])**2)
deno = r12/(1+beta*r12)
return exp(-0.5*alpha*(r1+r2)+deno)
# Local energy for the 2-electron quantum dot in two dims, using analytical local energy
def LocalEnergy(r,alpha,beta):
r1 = (r[0,0]**2 + r[0,1]**2)
r2 = (r[1,0]**2 + r[1,1]**2)
r12 = sqrt((r[0,0]-r[1,0])**2 + (r[0,1]-r[1,1])**2)
deno = 1.0/(1+beta*r12)
deno2 = deno*deno
return 0.5*(1-alpha*alpha)*(r1 + r2) +2.0*alpha + 1.0/r12+deno2*(alpha*r12-deno2+2*beta*deno-1.0/r12)
# Derivate of wave function ansatz as function of variational parameters
def DerivativeWFansatz(r,alpha,beta):
WfDer = np.zeros((2), np.double)
r1 = (r[0,0]**2 + r[0,1]**2)
r2 = (r[1,0]**2 + r[1,1]**2)
r12 = sqrt((r[0,0]-r[1,0])**2 + (r[0,1]-r[1,1])**2)
deno = 1.0/(1+beta*r12)
deno2 = deno*deno
WfDer[0] = -0.5*(r1+r2)
WfDer[1] = -r12*r12*deno2
return WfDer
# Setting up the quantum force for the two-electron quantum dot, recall that it is a vector
def QuantumForce(r,alpha,beta):
qforce = np.zeros((NumberParticles,Dimension), np.double)
r12 = sqrt((r[0,0]-r[1,0])**2 + (r[0,1]-r[1,1])**2)
deno = 1.0/(1+beta*r12)
qforce[0,:] = -2*r[0,:]*alpha*(r[0,:]-r[1,:])*deno*deno/r12
qforce[1,:] = -2*r[1,:]*alpha*(r[1,:]-r[0,:])*deno*deno/r12
return qforce
# Computing the derivative of the energy and the energy
def EnergyDerivative(x0):
# Parameters in the Fokker-Planck simulation of the quantum force
D = 0.5
TimeStep = 0.05
# positions
PositionOld = np.zeros((NumberParticles,Dimension), np.double)
PositionNew = np.zeros((NumberParticles,Dimension), np.double)
# Quantum force
QuantumForceOld = np.zeros((NumberParticles,Dimension), np.double)
QuantumForceNew = np.zeros((NumberParticles,Dimension), np.double)
energy = 0.0
DeltaE = 0.0
alpha = x0[0]
beta = x0[1]
EnergyDer = 0.0
DeltaPsi = 0.0
DerivativePsiE = 0.0
#Initial position
for i in range(NumberParticles):
for j in range(Dimension):
PositionOld[i,j] = normalvariate(0.0,1.0)*sqrt(TimeStep)
wfold = WaveFunction(PositionOld,alpha,beta)
QuantumForceOld = QuantumForce(PositionOld,alpha, beta)
#Loop over MC MCcycles
for MCcycle in range(NumberMCcycles):
#Trial position moving one particle at the time
for i in range(NumberParticles):
for j in range(Dimension):
PositionNew[i,j] = PositionOld[i,j]+normalvariate(0.0,1.0)*sqrt(TimeStep)+\
QuantumForceOld[i,j]*TimeStep*D
wfnew = WaveFunction(PositionNew,alpha,beta)
QuantumForceNew = QuantumForce(PositionNew,alpha, beta)
GreensFunction = 0.0
for j in range(Dimension):
GreensFunction += 0.5*(QuantumForceOld[i,j]+QuantumForceNew[i,j])*\
(D*TimeStep*0.5*(QuantumForceOld[i,j]-QuantumForceNew[i,j])-\
PositionNew[i,j]+PositionOld[i,j])
GreensFunction = exp(GreensFunction)
ProbabilityRatio = GreensFunction*wfnew**2/wfold**2
#Metropolis-Hastings test to see whether we accept the move
if random() <= ProbabilityRatio:
for j in range(Dimension):
PositionOld[i,j] = PositionNew[i,j]
QuantumForceOld[i,j] = QuantumForceNew[i,j]
wfold = wfnew
DeltaE = LocalEnergy(PositionOld,alpha,beta)
DerPsi = DerivativeWFansatz(PositionOld,alpha,beta)
DeltaPsi += DerPsi
energy += DeltaE
DerivativePsiE += DerPsi*DeltaE
# We calculate mean values
energy /= NumberMCcycles
DerivativePsiE /= NumberMCcycles
DeltaPsi /= NumberMCcycles
EnergyDer = 2*(DerivativePsiE-DeltaPsi*energy)
return EnergyDer
# Computing the expectation value of the local energy
def Energy(x0):
# Parameters in the Fokker-Planck simulation of the quantum force
D = 0.5
TimeStep = 0.05
# positions
PositionOld = np.zeros((NumberParticles,Dimension), np.double)
PositionNew = np.zeros((NumberParticles,Dimension), np.double)
# Quantum force
QuantumForceOld = np.zeros((NumberParticles,Dimension), np.double)
QuantumForceNew = np.zeros((NumberParticles,Dimension), np.double)
energy = 0.0
DeltaE = 0.0
alpha = x0[0]
beta = x0[1]
#Initial position
for i in range(NumberParticles):
for j in range(Dimension):
PositionOld[i,j] = normalvariate(0.0,1.0)*sqrt(TimeStep)
wfold = WaveFunction(PositionOld,alpha,beta)
QuantumForceOld = QuantumForce(PositionOld,alpha, beta)
#Loop over MC MCcycles
for MCcycle in range(NumberMCcycles):
#Trial position moving one particle at the time
for i in range(NumberParticles):
for j in range(Dimension):
PositionNew[i,j] = PositionOld[i,j]+normalvariate(0.0,1.0)*sqrt(TimeStep)+\
QuantumForceOld[i,j]*TimeStep*D
wfnew = WaveFunction(PositionNew,alpha,beta)
QuantumForceNew = QuantumForce(PositionNew,alpha, beta)
GreensFunction = 0.0
for j in range(Dimension):
GreensFunction += 0.5*(QuantumForceOld[i,j]+QuantumForceNew[i,j])*\
(D*TimeStep*0.5*(QuantumForceOld[i,j]-QuantumForceNew[i,j])-\
PositionNew[i,j]+PositionOld[i,j])
GreensFunction = exp(GreensFunction)
ProbabilityRatio = GreensFunction*wfnew**2/wfold**2
#Metropolis-Hastings test to see whether we accept the move
if random() <= ProbabilityRatio:
for j in range(Dimension):
PositionOld[i,j] = PositionNew[i,j]
QuantumForceOld[i,j] = QuantumForceNew[i,j]
wfold = wfnew
DeltaE = LocalEnergy(PositionOld,alpha,beta)
energy += DeltaE
if Printout:
outfile.write('%f\n' %(energy/(MCcycle+1.0)))
# We calculate mean values
energy /= NumberMCcycles
return energy
#Here starts the main program with variable declarations
NumberParticles = 2
Dimension = 2
# seed for rng generator
seed()
# Monte Carlo cycles for parameter optimization
Printout = False
NumberMCcycles= 10000
# guess for variational parameters
x0 = np.array([0.9,0.2])
# Using Broydens method to find optimal parameters
res = minimize(Energy, x0, method='BFGS', jac=EnergyDerivative, options={'gtol': 1e-4,'disp': True})
x0 = res.x
# Compute the energy again with the optimal parameters and increased number of Monte Cycles
NumberMCcycles= 2**19
Printout = True
FinalEnergy = Energy(x0)
EResult = np.array([FinalEnergy,FinalEnergy])
outfile.close()
#nice printout with Pandas
import pandas as pd
from pandas import DataFrame
data ={'Optimal Parameters':x0, 'Final Energy':EResult}
frame = pd.DataFrame(data)
print(frame)
```
Note that the **minimize** function returns the final values for the
variable $\alpha=x0[0]$ and $\beta=x0[1]$ in the array $x$.
When we have found the minimum, we use these optimal parameters to perform a production run of energies.
The output is in turn written to file and is used, together with resampling methods like the **blocking method**,
to obtain the best possible estimate for the standard deviation. The optimal minimum is, even with our guess, rather close to the exact value of $3.0$ a.u.
The [sampling
functions](https://github.com/CompPhysics/ComputationalPhysics2/tree/gh-pages/doc/Programs/Resampling)
can be used to perform both a blocking analysis, or a standard
bootstrap and jackknife analysis.
### How do we proceed?
There are several paths which can be chosen. One is to extend the
brute force gradient descent method with an adapative stochastic
gradient. There are several examples of this. A recent approach based
on [the Langevin equations](https://arxiv.org/pdf/1805.09416.pdf)
seems like a promising approach for general and possibly non-convex
optimization problems.
Here we would like to point out that our next step is now to use the
optimal values for our variational parameters and use these as inputs
to a production run. Here we would output values of the energy and
perform for example a blocking analysis of the results in order to get
a best possible estimate of the standard deviation.
## Resampling analysis
The next step is then to use the above data sets and perform a
resampling analysis, either using say the Bootstrap method or the
Blocking method. Since the data will be correlated, we would recommend
to use the non-iid Bootstrap code here. The theoretical background for these resampling methods is found in the [statistical analysis lecture notes](http://compphysics.github.io/ComputationalPhysics2/doc/pub/statanalysis/html/statanalysis.html)
Here we have tailored the codes to the output file from the previous example. We present first the bootstrap resampling with non-iid stochastic event.
```
# Common imports
import os
# Where to save the figures and data files
DATA_ID = "Results/EnergyMin"
def data_path(dat_id):
return os.path.join(DATA_ID, dat_id)
infile = open(data_path("Energies.dat"),'r')
from numpy import std, mean, concatenate, arange, loadtxt, zeros, ceil
from numpy.random import randint
from time import time
def tsboot(data,statistic,R,l):
t = zeros(R); n = len(data); k = int(ceil(float(n)/l));
inds = arange(n); t0 = time()
# time series bootstrap
for i in range(R):
# construct bootstrap sample from
# k chunks of data. The chunksize is l
_data = concatenate([data[j:j+l] for j in randint(0,n-l,k)])[0:n];
t[i] = statistic(_data)
# analysis
print ("Runtime: %g sec" % (time()-t0)); print ("Bootstrap Statistics :")
print ("original bias std. error")
print ("%8g %14g %15g" % (statistic(data), \
mean(t) - statistic(data), \
std(t) ))
return t
# Read in data
X = loadtxt(infile)
# statistic to be estimated. Takes two args.
# arg1: the data
def stat(data):
return mean(data)
t = tsboot(X, stat, 2**12, 2**10)
```
The blocking code, based on the article of [Marius Jonsson](https://journals.aps.org/pre/abstract/10.1103/PhysRevE.98.043304) is given here
```
# Common imports
import os
# Where to save the figures and data files
DATA_ID = "Results/EnergyMin"
def data_path(dat_id):
return os.path.join(DATA_ID, dat_id)
infile = open(data_path("Energies.dat"),'r')
from numpy import log2, zeros, mean, var, sum, loadtxt, arange, array, cumsum, dot, transpose, diagonal, sqrt
from numpy.linalg import inv
def block(x):
# preliminaries
n = len(x)
d = int(log2(n))
s, gamma = zeros(d), zeros(d)
mu = mean(x)
# estimate the auto-covariance and variances
# for each blocking transformation
for i in arange(0,d):
n = len(x)
# estimate autocovariance of x
gamma[i] = (n)**(-1)*sum( (x[0:(n-1)]-mu)*(x[1:n]-mu) )
# estimate variance of x
s[i] = var(x)
# perform blocking transformation
x = 0.5*(x[0::2] + x[1::2])
# generate the test observator M_k from the theorem
M = (cumsum( ((gamma/s)**2*2**arange(1,d+1)[::-1])[::-1] ) )[::-1]
# we need a list of magic numbers
q =array([6.634897,9.210340, 11.344867, 13.276704, 15.086272, 16.811894, 18.475307, 20.090235, 21.665994, 23.209251, 24.724970, 26.216967, 27.688250, 29.141238, 30.577914, 31.999927, 33.408664, 34.805306, 36.190869, 37.566235, 38.932173, 40.289360, 41.638398, 42.979820, 44.314105, 45.641683, 46.962942, 48.278236, 49.587884, 50.892181])
# use magic to determine when we should have stopped blocking
for k in arange(0,d):
if(M[k] < q[k]):
break
if (k >= d-1):
print("Warning: Use more data")
return mu, s[k]/2**(d-k)
x = loadtxt(infile)
(mean, var) = block(x)
std = sqrt(var)
import pandas as pd
from pandas import DataFrame
data ={'Mean':[mean], 'STDev':[std]}
frame = pd.DataFrame(data,index=['Values'])
print(frame)
```
## Additional notes
What we have not done here is to parallelize the codes. This material will be added later.
The material we have developed serves thus the aim to bridge the gap between traditional Monte Carlo calculations and Machine Learning methods. The most important ingredients here are
1. The definition of the cost function (in our case the energy as function of the variational parameters)
2. The optimization methods like gradient descent and stochastic gradient descent
3. Metropolis sampling (and later also Gibbs sampling) and Markov chain Monte Carlo approaches
We will meet these concepts again in our notes on Boltzmann machines (notebook 2) and on shadow wave functions (notebook 3).
| github_jupyter |
```
#Show ALL outputs in cell, not only last result
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
#Set relative path mapping for module imports
import sys
sys.path.append("../../")
#for path in sys.path:
# print(path)
# External Dependencies
import numpy as np
import pandas as pd
#Read in pickled data
X_y_data = pd.read_pickle("../data/interim/X_y_data.pkl")
X = pd.read_pickle("../data/interim/X.pkl")
y = pd.read_pickle("../data/interim/y.pkl")
#Recap data structure
X_y_data.head()
X_y_data.shape
```
## h) Dabl
#### ii) Dabl baseline model
```
# Initial Model Building with dabl
# The SimpleClassifier implements the familiar scikit-learn API of fit and predict
dabl_X_train = dabl_X_y_train_clean.drop([target], axis=1)
dabl_y_train = dabl_X_y_train_clean[target]
dabl_X_test = dabl_X_y_test_clean.drop([target], axis=1)
dabl_y_test = dabl_X_y_test_clean[target]
sc = dabl.SimpleClassifier(random_state=0) \
.fit(dabl_X_train, dabl_y_train)
print("Mean Accuracy on the given TEST data and labels:", sc.score(dabl_X_test, dabl_y_test))
dabl.explain(sc, dabl_X_test, dabl_y_test)
# Initial Model Building with dabl
# The SimpleClassifier implements the familiar scikit-learn API of fit and predict
sc = dabl.SimpleClassifier(random_state=0) \
.fit(X_train, y_train)
print("Mean Accuracy on the given TEST data and labels:", sc.score(X_test, y_test))
dabl.explain(sc, X_test, y_test)
#ac = dabl.AnyClassifier().fit(X_train, y_train)
#dabl.explain(ac)
#ACTION: What is a baseline model?
```
## Model Selection
```
from IPython.display import Image
from IPython.core.display import HTML
Image(url= "https://scikit-learn.org/stable/_static/ml_map.png")
#Which algorithms/estimators are options?
#The estimator you choose for your project will depend on the data set you have and the problem that you are trying to solve.
```
# Fit Model & Predict
```
https://pandas-ml.readthedocs.io/en/latest/index.html
https://www.scikit-yb.org/en/latest/
#Decide on appropriate algorithm/estimator(s)
from sklearn.linear_model import LogisticRegression
lr = LogisticRegression()
#Fit model to training data
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0)
model = lr.fit(X_train, y_train)
#Predict on previously unseen test data
predictions = model.predict(X_test)
#Easy to repeat analysis for another algorithm/estimator
from sklearn.ensemble import RandomForestClassifier
rf = RandomForestClassifier()
rf_model = rf.fit(X_train, y_train)
rf_predictions = rf_model.predict(X_test)
```
| github_jupyter |
```
from utils.prometheus import *
from utils.data import *
import pickle
hf_data = get_hf_data("../data/hf-data.pkl.gzip")
hf_static = get_hf_static_data(hf_data)
durations = hf_data["end"] - hf_data["start"]
```
# Montage2
| | |
|:-:|:-:|
|duration |>= 5 min |
|split |0.75 - 0 - 0.25|
|steps |[5, 10, 15] |
|past_sec |[60, 120, 180] |
|future_sec|[60, 120, 180] |
```
dataset = "montage2_gte_5min"
montage2_gte_5min = hf_data[(hf_data["workflowName"] == "montage2") & (durations >= timedelta(minutes=5)) & (durations <= timedelta(hours=10))].copy()
montage2_gte_5min.loc[montage2_gte_5min["size"] == 619.0, "size"] = 0.25
montage2_gte_5min["size"] = pd.to_numeric(montage2_gte_5min["size"], downcast="float")
steps = [5, 10, 15]
n_pasts = [1 * 60, 2 * 60, 3 * 60]
n_futures = [1 * 60, 2 * 60, 3 * 60]
create_dataset(hf_data, dataset, montage2_gte_5min, steps, n_pasts, n_futures)
```
# Montage2
| | |
|:-:|:-:|
|duration |>= 10 min |
|split |0.75 - 0 - 0.25 |
|steps |[5, 10, 15, 30] |
|past_sec |[60, 120, 180, 240, 300] |
|future_sec|[60, 120, 180, 240, 300] |
```
dataset = "montage2_gte_10min"
montage2_gte_10min = hf_data[(hf_data["workflowName"] == "montage2") & (durations >= timedelta(minutes=10)) & (durations <= timedelta(hours=10))].copy()
montage2_gte_10min["size"] = pd.to_numeric(montage2_gte_10min["size"], downcast="float")
steps = [5, 10, 15, 30]
n_pasts = [1 * 60, 2 * 60, 3 * 60, 4 * 60, 5 * 60]
n_futures = [1 * 60, 2 * 60, 3 * 60, 4 * 60, 5 * 60]
create_dataset(hf_data, dataset, montage2_gte_5min, steps, n_pasts, n_futures)
```
# Montage
| | |
|:-:|:-:|
|duration |>= 5 min |
|split |0.75 - 0 - 0.25 |
|steps |[5, 10, 15, 30] |
|past_sec |[60, 120, 180] |
|future_sec|[60, 120, 180] |
```
dataset = "montage_gte_5min"
montage_gte_5min = hf_data[(hf_data["workflowName"] == "montage") & (durations >= timedelta(minutes=5)) & (durations < timedelta(minutes=60))].copy()
montage_gte_5min["size"] = pd.to_numeric(montage_gte_5min["size"], downcast="float")
steps = [5, 10, 15]
n_pasts = [1 * 60, 2 * 60, 3 * 60]
n_futures = [1 * 60, 2 * 60, 3 * 60]
create_dataset(hf_data, dataset, montage2_gte_5min, steps, n_pasts, n_futures)
```
# SoyKB
| | |
|:-:|:-:|
|split |0.75 - 0 - 0.25 |
|steps |[10, 15, 30, 60] |
|past_sec |[120, 180, 240, 300, 600] |
|future_sec|[120, 180, 240, 300, 600] |
```
dataset = "soykb"
soykb = hf_data[(hf_data["workflowName"] == "soykb") & (durations < timedelta(hours=13))].copy()
soykb["size"] = pd.to_numeric(soykb["size"], downcast="float")
steps = [10, 15, 30, 60]
n_pasts = [2 * 60, 3 * 60, 4 * 60, 5 * 60, 10 * 60]
n_futures = [2 * 60, 3 * 60, 4 * 60, 5 * 60, 10 * 60]
create_dataset(hf_data, dataset, montage2_gte_5min, steps, n_pasts, n_futures)
```
| github_jupyter |
# Basic chemical, electrical, and thermodynamic principles
To develop a quantitative understanding of how these processes work, we start with a set of definitions of the some quantities and concepts with which we are concerned. Specifically, this section reviews basic biochemical, thermodynamic, and related concepts that are particularly relevant to the quantitative analysis of mitochondrial ATP synthesis.
```{figure} Figure1.png
------
name: mitofig
------
Diagram of a mitochondrion with the cytosol, intermembrane space (IMS), and matrix indicated. *Inset from left to right:* Protein channels and complexes associated with oxidative phosphorylation in the cristae of the mitochondrion. Complex I (C1) catalyzes the oxidation of NADH$^{2-}$ to NAD$^{-}$ and reduction of ubiquinone (Q) to QH$_2$. Complex II (C2) catalyzes the oxidation of FADH$_2$ to FAD coupled to the reduction of Q. Complex III (C3) catalyzes the oxidation of QH$_2$ coupled to the reduction of cytochrome c (Cyt c). Complex IV (C4) catalyzes the oxidation of Cyt c coupled to the reduction of oxygen to water. These redox transfers drive pumping of H$^+$ ions out of the matrix, establishing the proton motive force across the inner mitochondrial membrane (IMM) that drives ATP synthesis at complex V, or the F$_0$F$_1$-ATPase (F$_0$F$_1$). The adenine nucleotide translocase (ANT) exchanges matrix ATP for IMS ADP. The inorganic phosphate cotransporter (PiC) brings protons and Pi from the IMS to the matrix. Lastly, there is a passive H$^{+}$ leak across the IMM. (Figure created with Biorender.com.)
```
## Mitochondrial anatomy
The mitochondrion is a membrane-bound, rod-shaped organelle that is responsible for generating most of the chemical energy needed to power the cell's biochemical reactions by respiration {cite}`Nicholls2013`. Mitochondria are comprised of an outer and inner membrane that are separated by the intermembrane space (IMS) ({numref}`mitofig`). The outer mitochondrial membrane is freely permeable to small molecules and ions. The IMM folds inward to make cristae that extend into the matrix. Transmembrane channels called porins and the respiratory complexes involved in oxidative phosphorylation and ATP synthesis allow for more selective IMM permeability. The IMM encloses the mitochondrial matrix, which contains mitochondrial deoxyribonucleic acid (DNA), the majority of mitochondrial proteins, soluble metabolic intermediates including ATP, ADP, and Pi, and the enzymes catalyzing the tricarboxylic acid (TCA) cycle and $\beta$-oxidation.
## IMM capacitance
The IMM acts as an electrical capacitor to store energy in an electrostatic potential difference between the milieu on each side. Electrical capacitance of a membrane ($C_m$) is the proportionality between the rate of charge transport across the membrane, i.e. current ($I$), to the rate of membrane potential ($\Delta \Psi$) change, that is,
```{math}
C_m \dfrac{ {\rm d} {\Delta\Psi}}{{\rm d} t} = I.
```
In the model and associated calculations presented below, we express fluxes in units of moles per unit time per unit volume of mitochondria. Thus, it is convenient to obtain an estimate of $C_m$ in units of mole per volt per volume of mitochondria. Mitochondria take on a roughly ellipsoid shape in vivo, and a more spherical morphometry in suspension of purified mitochondria {cite}`Picard2011`. To estimate the mitochondrial surface area-to-volume ratio, we take a representative mitochondrion as a sphere with radius $r = 1 \ \mu\text{m}$ and obtain a surface area-to-volume ratio of $3 \ \mu\text{m}^{-1}$. Furthermore, we estimate that the IMM has ten-fold greater surface area than the outer membrane, yielding a surface area to volume ratio of $30 \ \mu\text{m}^{-1}$ for the IMM. Since the capacitance density of biological membranes ranges from $0.5\text{-}1.0 \mu\text{F cm}^{-2}$, or $0.5 \text{-} 1.0 \times \ 10^{-8} \ \mu\text{F} \ \mu\text{m}^{-2}$ {cite}`Nicholls2013`, $C_m$ is approximately $30 \times 10^{-8} \ \mu\text{F} \ \mu\text{m}^{-3} = 300 \ \text{F (L mito)}^{-1}$. To convert to the units used in the calculations below, we have
```{math}
C_m = 300 \ \frac{\rm F}{\rm L \ mito} = 300 \ \frac{\rm C}{\rm V \cdot L \, mito}\cdot
\frac{1}{F}\, \frac{\rm mol}{\rm C} =
3.1 \times 10^{-3} \,
\frac{\rm mol}{\rm V \cdot L \, mito}, \,
```
where $F = 96,485 \ \text{C mol}^{-1}$ is Faraday's constant.
## Gibbs free energy
A *free energy* is a thermodynamic quantity that relates a change in the thermodynamic state of a system to an associated change in total entropy of the system plus its environment. Chemical reaction processes necessarily proceed in the direction associated with a reduction in free energy {cite}`Nicholls2013`. When free energy of a system is reduced, total entropy (of the universe) is increased. The form of free energy that is operative in constant-temperature and constant-pressure systems (most relevant for biochemistry) is the Gibbs free energy, or simply the *Gibbs energy*.
For a chemical reaction of reactants $A_i$ and products $B_j$,
```{math}
\sum_{i = 1}^M m_i A_i \rightleftharpoons \sum_{j = 1}^N n_j B_j
```
where $M$ and $N$ are the total number of reactants and products, respectively, and $m_i$ and $n_j$ are the coefficients of reactant $i$ and product $j$, respectively, the Gibbs energy can be expressed as
```{math}
:label: Delta_rG
\Delta_r G = \Delta_r G^\circ + R{\rm T} \ln \left( \dfrac{ \prod_{i = 1}^{N} [\text{B}_j]^{n_i}}{ \prod_{i = 1}^{M} [\text{A}_i]^{m_i}} \right),
```
where $\Delta_r G^\circ$ is the reference Gibbs energy for the reaction (a constant at given constant chemical conditions of temperature, pressure, ionic conditions, etc.), $R = 8.314 \ \text{J mol}^{-1} \ \text{K}^{-1}$ is the gas constant, and $\text{T} = 310.15 \ \text{K}$ is the temperature. The second term on the right hand side of Equation {eq}`Delta_rG` governs how changes in concentrations of species affects $\Delta_r G$. Applications of Equation {eq}`Delta_rG` to reactions in aqueous solution usually adopt the convention that all solute concentrations are measured relative to 1 Molar, ensuring that the argument of the logarithm is unitless regardless of the stoichiometry of the reaction.
A system is in chemical equilibrium when there is no thermodynamic driving force, that is, $\Delta_r G = 0$. Thus, for this chemical reaction the reference Gibbs energy is related to the equilibrium constant as
```{math}
K_{eq} = \left( \frac{\prod_{i = 1}^{N} [\text{B}_j]^{n_i}}{\prod_{i = 1}^{M} [\text{A}_i]^{m_i}} \right)_{eq}
= \exp\left\{ -\frac{\Delta_r G^\circ}{R{\rm T}} \right\} .
```
## Membrane potential and proton motive force
Free energy associated with the oxidation of primary fuels is transduced to generate the chemical potential across the IMM known as the {\em proton motive force}, which is used to synthesize ATP in the matrix and transport ATP out of the matrix to the cytosol {cite}`Nicholls2013`. The thermodynamic driving force for translocation of hydrogen ions ($\text{H}^{+}$) across the IMM has two components: the difference in electrostatic potential across the membrane, $\Delta\Psi$ (V), and the difference in $\text{H}^{+}$ concentration (or activity) between the media on either side of the membrane, $\Delta\text{pH}$, that is
```{math}
:label: DG_H
\Delta G_{\rm H} &=& -F\Delta\Psi + R{\rm T}\ln\left( [{\rm H}^+]_x/[{\rm H}^+]_c \right) \nonumber \\
&=& -F\Delta\Psi - 2.3 R{\rm T} \, \Delta{\rm pH},
```
where the subscripts $x$ and $c$ indicate matrix and external (cytosol) spaces. $\Delta\Psi$ is defined as the cytosolic potential minus matrix potential, yielding a negative change in free energy for a positive potential. Membrane potential in respiring mitochondria is approximately $150 \text{-} 200 \ \text{mV}$, yielding a contribution to $\Delta G_{\rm H}$ on the order of $15 \text{-} 20 \ \text{kJ mol}^{-1}$ {cite}`Bazil2016`. Under in vitro conditions, $\Delta\text{pH}$ between the matrix and external buffer is on the order of $0.1 \ \text{pH}$ units {cite}`Bazil2016`. Thus, the contribution to proton motive force from a pH difference is less than $1 \ \text{kJ mol}^{-1}$ and substantially smaller than that from $\Delta\Psi$.
## Thermodynamics of ATP synthesis/hydrolysis
Under physiological conditions the ATP hydrolysis reaction
```{math}
:label: ATP1
\text{ATP}^{4-} + \text{H}_2\text{O} \rightleftharpoons
\text{ADP}^{3-} + \text{HPO}_4^{2-} + \text{H}^{+}
```
is thermodynamically favored to proceed from the left-to-right direction. The Gibbs energy associated with turnover of this reaction is
```{math}
:label: DrG_ATP
\Delta_r G_{\rm ATP} = \Delta_r G^o_\text{ATP} + R{\rm T} \ln
\left( \frac{ [\text{ADP}^{3-}] [\text{HPO}_4^{2-}] [{\rm H}^{+}] }
{ [\text{ATP}^{4-}] }\right),
```
where the Gibbs energy for ATP hydrolysis under physiological conditions is approximately $\Delta_r G^o_\text{ATP} = 4.99 \ \text{kJ mol}^{-1}$ {cite}`Li2011`. Using the convention that all concentrations are formally defined as measured relative to 1 Molar, the argument of the logarithm in Equation {eq}`DrG_ATP` is unitless.
### Calculation of the ATP hydrolysis potential
Equation {eq}`DrG_ATP` expresses the Gibbs energy of chemical Equation {eq}`ATP1` in terms of its *chemical species*. In practice, biochemistry typically deals with biochemical *reactants*, which are comprised of sums of rapidly interconverting chemical species. We calculate the total ATP concentration, $[\Sigma \text{ATP}]$, in terms of its bound and unbound species, that is,
```{math}
:label: sumATP
[\Sigma \text{ATP}] &= [\text{ATP}^{4-}] + [\text{MgATP}^{2-}] + [\text{HATP}^{3-}] + [\text{KATP}^{3-}] \nonumber\\
&= [\text{ATP}^{4-}] + \frac{[\text{Mg}^{2+}] [\text{ATP}^{4-}]}{K_{\text{MgATP}}} + \frac{ [\text{H}^{+}] [\text{ATP}^{4-}]}{K_{\text{HATP}}} + \frac{ [\text{K}^{+}] [\text{ATP}^{4-}]}{K_{\text{KATP}}} \nonumber \\
&= [\text{ATP}^{4-}] \left( 1 + \frac{[\text{Mg}^{2+}]}{K_{\text{MgATP}}} + \frac{ [\text{H}^{+}]}{K_{\text{HATP}}} + \frac{ [\text{K}^{+}]}{K_{\text{KATP}}} \right) \nonumber \\
&= [\text{ATP}^{4-}] P_{\text{ATP}},
```
where $P_{\text{ATP}}$ is a *binding polynomial*. Here, we we account for only the single cation-bound species. (Free $\text{H}^+$ in solution associates with water to form $\text{H}_3\text{O}^+$. Here we use [$\text{H}^+$] to indicate hydrogen ion activity, which is equal to $10^{-\text{pH}}$.) {numref}`table-dissociationconstants` lists the dissociation constants used in this study from {cite}`Li2011`. Similarly, total ADP, [$\Sigma \text{ADP}$], and inorganic phosphate, [$\Sigma \text{Pi}$], concentrations are
```{math}
:label: sumADP
[\Sigma {\rm ADP} ] &= [{\rm ADP}^{3-}]\left( 1 + \frac{[{\rm Mg}^{2+}]}{K_{\rm MgADP}} + \frac{ [{\rm H}^{+}]}{K_{\rm HADP}} + \frac{ [{\rm K}^{+}]}{K_{\rm KADP}} \right) \nonumber \\
&= [{\rm ADP}^{3-}]P_{\rm ADP}
```
and
```{math}
:label: sumPi
[\Sigma {\rm Pi} ] &= [{\rm HPO}_4^{2-}] \left( 1 + \frac{[{\rm Mg}^{2+}]}{K_{\rm MgPi}} + \frac{ [{\rm H}^{+}]}{K_{\rm HPi}} + \frac{ [{\rm K}^{+}]}{K_{\rm KPi}} \right) \nonumber \\
&= [{\rm HPO}_4^{2-}] P_{\rm Pi},
```
for binding polynomials $P_{\text{ADP}}$ and $P_{\text{Pi}}$.
Expressing the Gibbs energy of ATP hydrolysis in Equation {eq}`ATP1` in terms of biochemical reactant concentrations, we obtain
```{math}
:label: ATP2
\Delta_r G_{\rm ATP} &= \Delta_r G^o_\text{ATP} + R{\rm T} \ln \left(
\frac{[\Sigma{\rm ADP}][\Sigma{\rm Pi}]}
{[\Sigma{\rm ATP}]}\cdot\frac{[{\rm H}^+]P_{\rm ATP}}{P_{\rm ADP}P_{\rm Pi}}
\right) \nonumber \\
&= \Delta_r G^o_\text{ATP}
+ R{\rm T} \ln \left(\frac{[{\rm H}^+]P_{\rm ATP}}{P_{\rm ADP}P_{\rm Pi}} \right)
+ R{\rm T} \ln \left(\frac{[\Sigma{\rm ADP}][\Sigma{\rm Pi}]}
{[\Sigma{\rm ATP}]}\right) \nonumber \\
&= \Delta_r G'^o_\text{ATP}
+ R{\rm T} \ln \left(\frac{[\Sigma{\rm ADP}][\Sigma{\rm Pi}]}
{[\Sigma{\rm ATP}]}\right)
```
where $\Delta_r G'^o_\text{ATP}$ is a transformed, or *apparent*, reference Gibbs energy for the reaction.
```{list-table} Dissociation constants given as 10$^{-\text{p}K_a}$.
:header-rows: 2
:name: table-dissociationconstants
* -
-
- Ligand ($L$)
-
* -
- Mg$^{2+}$
- H$^{+}$
- K$^{+}$
* - $K_{L-\text{ATP}}$
- $10^{-3.88}$
- $10^{-6.33}$
- $10^{-1.02}$
* - $K_{L-\text{ADP}}$
- $10^{-3.00}$
- $10^{-6.26}$
- $10^{-0.89}$
* - $K_{L-\text{Pi}}$
- $10^{-1.66}$
- $10^{-6.62}$
- $10^{-0.42}$
```
The following code computes the apparent Gibbs energy with $\text{pH} = 7$, $[\text{K}^{+}] = 150 \ \text{mM}$, and $[\text{Mg}^{2+}] = 1 \ \text{mM}$. Biochemical reactant concentrations are set such that the total adenine nucleotide (TAN) pool inside the mitochondrion is $10 \ \text{mM}$, $[\Sigma \text{ATP}] = 0.5 \ \text{mM}$, $[\Sigma \text{ADP}] = 9.5 \ \text{mM}$, and $[\Sigma \text{Pi}] = 1 \ \text{mM}$. Here, we obtain a value of approximately $\text{-}45 \ \text{kJ mol}^{-1}$.
```
# Import numpy package for calculations
import numpy as np
# Dissociation constants
K_MgATP = 10**(-3.88)
K_MgADP = 10**(-3.00)
K_MgPi = 10**(-1.66)
K_HATP = 10**(-6.33)
K_HADP = 10**(-6.26)
K_HPi = 10**(-6.62)
K_KATP = 10**(-1.02)
K_KADP = 10**(-0.89)
K_KPi = 10**(-0.42)
# Gibbs energy under physiological conditions(J mol^(-1))
DrGo_ATP = 4990
# Thermochemical constants
R = 8.314 # J (mol * K)**(-1)
T = 310.15 # K
F = 96485 # C mol**(-1)
# Environment concentrations
pH = 7
H = 10**(-pH) # Molar
K = 150e-3 # Molar
Mg = 1e-3 # Molar
# Binding polynomials
P_ATP = 1 + H/K_HATP + K/K_KATP + Mg/K_MgATP # equation 6
P_ADP = 1 + H/K_HADP + K/K_KADP + Mg/K_MgADP # equation 7
P_Pi = 1 + H/K_HPi + K/K_KPi + Mg/K_MgPi # equation 8
# Total concentrations
sumATP = 0.5e-3 # Molar
sumADP = 9.5e-3 # Molar
sumPi = 1.0e-3 # Molar
# Reaction:
# ATP4− + H2O ⇌ ADP3− + HPO2−4 + H+
# Use equation 8 to calcuate apparent reference Gibbs energy
DrG_ATP_apparent = DrGo_ATP + R * T * np.log(H * P_ATP / (P_ADP * P_Pi))
# Use equation 8 to calculate reaction Gibbs energy
DrG_ATP = DrG_ATP_apparent + R * T * np.log((sumADP * sumPi / sumATP))
print('Gibbs energy of ATP hydrolysis (kJ mol^(-1))')
print(DrG_ATP / 1000)
```
The reactant concentrations used in the above calculation represent reasonable values for concentrations in the mitochondrial matrix. In the cytosol, the ATP/ADP ratio is on the order of 100:1, yielding a $\Delta_r G_\text{ATP}$ of approximately $\text{-}64 \ \text{kJ mol}^{-1}$.
Note the large difference in magnitude of the estimated Gibbs energy of ATP hydrolysis inside (-$45 \ \text{kJ mol}^{-1}$) versus outside (-$64 \ \text{kJ mol}^{-1}$) of the mitochondrial matrix. Light will be shed on the mechanisms underlying this difference via the calculations and analyses presented below.
### ATP synthesis in the mitochondrial matrix
The F$_0$F$_1$ ATP synthase catalyzes the synthesis of ATP from ADP and Pi by coupling to the translocation of $n_{\text{F}} = 8/3$ protons from the cytosol to the matrix via the combined reaction
```{math}
:label: ATP3
({\rm ADP}^{3-})_x + ({\rm HPO}_4^{2-})_x + ({\rm H}^+)_x + n_{\text{F}} (\text{H}^{+})_c
\rightleftharpoons
({\rm ATP})^{4-}_x + {\rm H_2O} + n_{\text{F}} (\text{H}^{+})_x \, .
```
Using the Gibbs energy of the reaction of Equation {eq}`ATP2` and the proton motive force in Equation {eq}`DG_H`, the overall Gibbs energy for the coupled process of ATP synthesis and proton transport via the F$_0$F$_1$ ATP synthase is
```{math}
:label: DG_F
\Delta G_{\text{F}} &=& -\Delta_r G_{\rm ATP} + n_\text{F} \Delta G_{\rm H} \nonumber \\
&=& -\Delta_r G'^o_\text{ATP} - R{\rm T} \ln \left(\frac{[\Sigma{\rm ADP}]_x[\Sigma{\rm Pi}]_x}
{[\Sigma{\rm ATP}]_x}\right) - n_\text{F} F \Delta \Psi + R{\rm T} \ln \left(
\frac{ [{\rm H}^{+}]_x }{ [{\rm H}^{+}]_c } \right)^{n_{\rm F}} .
```
Note that the negative before $\Delta_r G_\text{ATP}$ indicates that the reaction of Equation {eq}`ATP1` is reversed in Equation {eq}`ATP3`. The equilibrium concentration ratio occurs when $\Delta G_{\text{F}} = 0$. Solving for the second term in Equation {eq}`DG_F`, we calculate the apparent equilibrium constant for ATP synthesis as
```{math}
:label: Kapp_F
K_{eq,\text{F}}^\prime =
\left( \frac{[\Sigma{\rm ATP}]_x}{[\Sigma{\rm ADP}]_x[\Sigma{\rm Pi}]_x} \right)_{eq} = \exp\left\{\frac{ \Delta_rG'^o_{\rm ATP} + n_{\rm F} F \Delta\Psi}{R{\rm T}}\right\}
\left( \frac{[{\rm H^+}]_c}{[{\rm H^+}]_x} \right)^{n_{\rm F}}.
```
(modelATPsynthesis)=
### Mathematical modeling ATP synthesis
A simple model of ATP synthesis kinetics can be constructed using the apparent equilibrium constant and mass-action kinetics in the form
```{math}
:label: J_F
J_{\text{F}} = X_{\text{F}} (K_{eq,\text{F}}^\prime [\Sigma \text{ADP}]_x [\Sigma \text{Pi}]_x - [\Sigma \text{ATP}]_x),
```
where $X_{\text{F}} = 1000 \ \text{mol s}^{-1} \ \text{(L mito)}^{-1}$ is a rate constant set to an arbitrarily high value that maintains the reaction in equilibrium in model simulations. To simulate ATP synthesis at a given membrane potential, matrix pH, cytosolic pH, and cation concentrations, we have
```{math}
:label: system-ATPase
\left\{
\renewcommand{\arraystretch}{2}
\begin{array}{rl}
\dfrac{ {\rm d} [\Sigma \text{ATP}]_x }{{\rm d} t} &= J_\text{F} / W_x \\
\dfrac{ {\rm d} [\Sigma \text{ADP}]_x }{{\rm d} t} &= -J_\text{F} / W_x \\
\dfrac{ {\rm d} [\Sigma \text{Pi}]_x }{{\rm d} t} &= -J_\text{F} / W_x,
\end{array}
\renewcommand{\arraystretch}{1}
\right.
```
where $W_x \ \text{((L matrix water) (L mito)}^{-1}$) is the fraction of water volume in the mitochondrial matrix to total volume of the mitochondrion. Dissociation constants are listed in {numref}`table-dissociationconstants` and all other parameters are listed in {numref}`table-biophysicalconstants`.
```{list-table} Parameters for ATP synthesis in vitro.
:header-rows: 1
:name: table-biophysicalconstants
* - Symbol
- Units
- Description
- Value
- Source
* - F$_0$F$_1$ ATP synthase constants
-
-
-
-
* - $n_{\text{F}}$
-
- Protons translocated
- $8/3 $
- {cite}`Nicholls2013`
* - $X_\text{F}$
- mol s$^{-1}$ (L mito)$^{-1}$
- Rate constant
- $1000 $
-
* - $\Delta_r G_\text{ATP}^\circ$
- kJ mol$^{-1}$
- Reference Gibbs energy
- $4.99 $
- {cite}`Li2011`
* - Biophysical constants
-
-
-
-
* - $R$
- J mol$^{-1}$ K$^{-1}$
- Gas constant
- $8.314 $
-
* - $T$
- K
- Temperature
- $310.15 $
-
* - $F$
- C mol$^{-1}$
- Faraday's constant
- $96485$
-
* - $C_m$
- mol V$^{-1}$ (L mito)$^{-1}$
- IMM capacitance
- $3.1\text{e-}3$
- {cite}`Beard2005`
* - Volume ratios
-
-
-
-
* - $V_c$
- (L cyto) (L cell)$^{-1}$
- Cyto to cell ratio
- $0.6601$
- {cite}`Bazil2016`
* - $V_m$
- (L mito) (L cell)$^{-1}$
- Mito to cell ratio
- $0.2882$
- {cite}`Bazil2016`
* - $V_{m2c}$
- (L mito) (L cyto)$^{-1}$
- Mito to cyto ratio
- $V_m / V_c$
-
* - $W_c$
- (L cyto water) (L cyto)$^{-1}$
- Cyto water space ratio
- $0.8425$
- {cite}`Bazil2016`
* - $W_m$
- (L mito water) (L mito)$^{-1}$
- Mito water space ratio
- $0.7238 $
- {cite}`Bazil2016`
* - $W_x$
- (L matrix water) (L mito)$^{-1}$
- Mito matrix water space ratio
- $0.9$ $W_m$
- {cite}`Bazil2016`
* - $W_i$
- (L IM water) (L mito)$^{-1}$
- IMS water space ratio
- $0.1$ $W_m$
- {cite}`Bazil2016`
```
The following code simulates steady state ATP, ADP, and Pi concentrations for $\Delta \Psi = 175 \ \text{mV}$. Here, a pH gradient is fixed across the IMM such that the pH in the matrix is slightly more basic than the cytosol, $\text{pH}_x = 7.4$ and $\text{pH}_c = 7.2$. All other conditions remain unchanged.
```
import matplotlib.pyplot as plt
import numpy as np
!pip install scipy
from scipy.integrate import solve_ivp
# Define system of ordinary differential equations from equation (13)
def dXdt(t, X, DPsi, pH_c):
# Unpack X state variable
sumATP, sumADP, sumPi = X
# Biophysical constants
R = 8.314 # J (mol * K)**(-1)
T = 310.15 # K
F = 96485 # C mol**(-1)
# F0F1 constants
n_F = 8/3
X_F = 1000 # mol (s * L mito)**(-1)
DrGo_F = 4990 # (J mol**(-1))
# Dissociation constants
K_MgATP = 10**(-3.88)
K_MgADP = 10**(-3.00)
K_MgPi = 10**(-1.66)
K_HATP = 10**(-6.33)
K_HADP = 10**(-6.26)
K_HPi = 10**(-6.62)
K_KATP = 10**(-1.02)
K_KADP = 10**(-0.89)
K_KPi = 10**(-0.42)
# Environment concentrations
pH_x = 7.4 # pH in matrix
H_x = 10**(-pH_x) # M
H_c = 10**(-pH_c) # M
K_x = 150e-3 # M
Mg_x = 1e-3 # M
# Volume ratios
W_m = 0.7238 # (L mito water) (L mito)**(-1)
W_x = 0.9 * W_m # (L matrix water) (L mito)**(-1)
# Binding polynomials
P_ATP = 1 + H_x/K_HATP + K_x/K_KATP + Mg_x/K_MgATP # equation 5
P_ADP = 1 + H_x/K_HADP + K_x/K_KADP + Mg_x/K_MgADP # equation 6
P_Pi = 1 + H_x/K_HPi + K_x/K_KPi + Mg_x/K_MgPi # equation 7
# Gibbs energy (equation 9)
DrGapp_F = DrGo_F + R * T * np.log(H_x * P_ATP / (P_ADP * P_Pi))
# Apparent equilibrium constant
Kapp_F = np.exp((DrGapp_F + n_F * F * DPsi)/ (R * T)) * (H_c / H_x) ** n_F
# Flux (mol (s * L mito)**(-1))
J_F = X_F * (Kapp_F * sumADP * sumPi - sumATP)
###### Differential equations (equation 13) ######
dATP = J_F / W_x
dADP = -J_F / W_x
dPi = -J_F / W_x
dX = (dATP, dADP, dPi)
return dX
# Simple steady state simulation at 175 mV membrane potential
# Initial conditions (M)
sumATP_0 = 0.5e-3
sumADP_0 = 9.5e-3
sumPi_0 = 1e-3
X_0 = np.array([sumATP_0, sumADP_0, sumPi_0])
# Inputs
DPsi = 175e-3 # Constant membrane potential (V)
pH_c = 7.2 # IMS/buffer pH
solutions = solve_ivp(dXdt, [0, 1], X_0, method = 'Radau', args = (DPsi,pH_c))
t = solutions.t
results = solutions.y
results = results * 1000
# Plot figure
plt.figure()
plt.plot(t, results[0,:], label = '[$\Sigma$ATP]$_x$')
plt.plot(t, results[1,:], label = '[$\Sigma$ADP]$_x$')
plt.plot(t, results[2,:], label = '[$\Sigma$Pi]$_x$')
plt.legend()
plt.xlabel('Time (s)')
plt.ylabel('Concentration (mM)')
plt.ylim(0, 10)
plt.show()
```
**Figure 2:** Steady state solution from Equation {eq}`system-ATPase` for $\Delta \Psi = 175$ mV, $\text{pH}_x = 7.4$, and $\text{pH}_c = 7.2$.
The above simulation shows that under the clamped pH and $\Delta\Psi$ conditions simulated here, the model quickly approaches an equilibrium steady state. (Even though all reaction fluxes go to zero in the final steady state, the ATP hydrolysis potential attains a finite nonzero value because of the energy supplied by the clamped proton motive force.) Most of the adenine nucleotide remains in the form of ADP and the final ATP/ADP ratio in the matrix is approximately $1$:$20$, with the inorganic phosphate concentration of approximately $1 \ \text{mM}$.
To explore how the equilibrium changes with membrane potential, the following code computes the predicted equilibrium steady-state over a ranges of $\Delta\Psi$ from $100$ to $250 \ \text{mV}$.
```
### Simulate over a range of membrane potential from 100 mV to 250 mV ###
# Define array to iterate over
membrane_potential = np.linspace(100,250) # mV
# Constant external pH
pH_c = 7.2 # IMS/buffer pH
# Define arrays to store steady state results
ATP_steady_DPsi = np.zeros(len(membrane_potential))
ADP_steady_DPsi = np.zeros(len(membrane_potential))
Pi_steady_DPsi = np.zeros(len(membrane_potential))
# Iterate through range of membrane potentials
for i in range(len(membrane_potential)):
DPsi = membrane_potential[i] / 1000 # convert to V
temp_results = solve_ivp(dXdt, [0, 5], X_0, method = 'Radau', args = (DPsi, pH_c,)).y*1000 # Concentration in mM
ATP_steady_DPsi[i] = temp_results[0,-1]
ADP_steady_DPsi[i] = temp_results[1,-1]
Pi_steady_DPsi[i] = temp_results[2,-1]
# Concentration vs DPsi
plt.figure()
plt.plot(membrane_potential, ATP_steady_DPsi, label = '[$\Sigma$ATP]$_x$')
plt.plot(membrane_potential, ADP_steady_DPsi, label = '[$\Sigma$ADP]$_x$')
plt.plot(membrane_potential, Pi_steady_DPsi, label = '[$\Sigma$Pi]$_x$')
plt.legend()
plt.xlabel('Membrane potential (mV)')
plt.ylabel('Concentration (mM)')
plt.xlim([100, 250])
plt.show()
```
**Figure 3:** Simulation of concentration versus $\Delta \Psi$ for Equation {eq}`system-ATPase` for $\Delta \Psi$ from $100$ to $250$ mV.
The above simulations show that under physiological levels of $\Delta$pH, matrix ATP concentrations become essentially zero for values of the membrane potential less than approximately $150 \ \text{mV}$. At higher levels of $\Delta\Psi$, all of the available phosphate is used to phosphorylate ADP to ATP. Since the initial $[\text{Pi}]$ and $[\text{ATP}]$ are $1 \ \text{mM}$ and $0.5 \ \text{mM}$, respectively, the maximum ATP obtained at the maximal $\Delta\Psi$ is $1.5 \ \text{mM}$.
| github_jupyter |
# Default of credit card clients Data Set
### Data Set Information:
This research aimed at the case of customers’ default payments in Taiwan and compares the predictive accuracy of probability of default among six data mining methods. From the perspective of risk management, the result of predictive accuracy of the estimated probability of default will be more valuable than the binary result of classification - credible or not credible clients. Because the real probability of default is unknown, this study presented the novel “Sorting Smoothing Method†to estimate the real probability of default. With the real probability of default as the response variable (Y), and the predictive probability of default as the independent variable (X), the simple linear regression result (Y = A + BX) shows that the forecasting model produced by artificial neural network has the highest coefficient of determination; its regression intercept (A) is close to zero, and regression coefficient (B) to one. Therefore, among the six data mining techniques, artificial neural network is the only one that can accurately estimate the real probability of default.
### Attribute Information:
This research employed a binary variable, default payment (Yes = 1, No = 0), as the response variable. This study reviewed the literature and used the following 23 variables as explanatory variables:
X1: Amount of the given credit (NT dollar): it includes both the individual consumer credit and his/her family (supplementary) credit.
X2: Gender (1 = male; 2 = female).
X3: Education (1 = graduate school; 2 = university; 3 = high school; 4 = others).
X4: Marital status (1 = married; 2 = single; 3 = others).
X5: Age (year).
X6 - X11: History of past payment. We tracked the past monthly payment records (from April to September, 2005) as follows:
X6 = the repayment status in September, 2005;
X7 = the repayment status in August, 2005;
. . .;
X11 = the repayment status in April, 2005. The measurement scale for the repayment status is: -1 = pay duly; 1 = payment delay for one month; 2 = payment delay for two months; . . .; 8 = payment delay for eight months; 9 = payment delay for nine months and above.
X12-X17: Amount of bill statement (NT dollar).
X12 = amount of bill statement in September, 2005;
X13 = amount of bill statement in August, 2005;
. . .;
X17 = amount of bill statement in April, 2005.
X18-X23: Amount of previous payment (NT dollar).
X18 = amount paid in September, 2005;
X19 = amount paid in August, 2005;
. . .;
X23 = amount paid in April, 2005.
```
%matplotlib inline
import os
import json
import time
import pickle
import requests
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
URL = "http://archive.ics.uci.edu/ml/machine-learning-databases/00350/default%20of%20credit%20card%20clients.xls"
def fetch_data(fname='default_of_credit_card_clients.xls'):
"""
Helper method to retreive the ML Repository dataset.
"""
response = requests.get(URL)
outpath = os.path.abspath(fname)
with open(outpath, 'w') as f:
f.write(response.content)
return outpath
# Fetch the data if required
# DATA = fetch_data()
# IMPORTANT - Issue saving xls file needed to be fix in fetch_data. Using a valid manually downloaded files instead for this example.
DATA = "./default_of_credit_card_clients2.xls"
FEATURES = [
"ID",
"LIMIT_BAL",
"SEX",
"EDUCATION",
"MARRIAGE",
"AGE",
"PAY_0",
"PAY_2",
"PAY_3",
"PAY_4",
"PAY_5",
"PAY_6",
"BILL_AMT1",
"BILL_AMT2",
"BILL_AMT3",
"BILL_AMT4",
"BILL_AMT5",
"BILL_AMT6",
"PAY_AMT1",
"PAY_AMT2",
"PAY_AMT3",
"PAY_AMT4",
"PAY_AMT5",
"PAY_AMT6",
"label"
]
LABEL_MAP = {
1: "Yes",
0: "No",
}
# Read the data into a DataFrame
df = pd.read_excel(DATA,header=None, skiprows=2, names=FEATURES)
# Convert class labels into text
for k,v in LABEL_MAP.items():
df.ix[df.label == k, 'label'] = v
# Describe the dataset
print df.describe()
df.head(5)
# Determine the shape of the data
print "{} instances with {} features\n".format(*df.shape)
# Determine the frequency of each class
print df.groupby('label')['label'].count()
from pandas.tools.plotting import scatter_matrix
scatter_matrix(df, alpha=0.2, figsize=(12, 12), diagonal='kde')
plt.show()
from pandas.tools.plotting import parallel_coordinates
plt.figure(figsize=(12,12))
parallel_coordinates(df, 'label')
plt.show()
from pandas.tools.plotting import radviz
plt.figure(figsize=(12,12))
radviz(df, 'label')
plt.show()
```
## Data Extraction
One way that we can structure our data for easy management is to save files on disk. The Scikit-Learn datasets are already structured this way, and when loaded into a `Bunch` (a class imported from the `datasets` module of Scikit-Learn) we can expose a data API that is very familiar to how we've trained on our toy datasets in the past. A `Bunch` object exposes some important properties:
- **data**: array of shape `n_samples` * `n_features`
- **target**: array of length `n_samples`
- **feature_names**: names of the features
- **target_names**: names of the targets
- **filenames**: names of the files that were loaded
- **DESCR**: contents of the readme
**Note**: This does not preclude database storage of the data, in fact - a database can be easily extended to load the same `Bunch` API. Simply store the README and features in a dataset description table and load it from there. The filenames property will be redundant, but you could store a SQL statement that shows the data load.
In order to manage our data set _on disk_, we'll structure our data as follows:
```
with open('./../data/cc_default/meta.json', 'w') as f:
meta = {'feature_names': FEATURES, 'target_names': LABEL_MAP}
json.dump(meta, f, indent=4)
from sklearn.datasets.base import Bunch
DATA_DIR = os.path.abspath(os.path.join(".", "..", "data", "cc_default"))
# Show the contents of the data directory
for name in os.listdir(DATA_DIR):
if name.startswith("."): continue
print "- {}".format(name)
def load_data(root=DATA_DIR):
# Construct the `Bunch` for the wheat dataset
filenames = {
'meta': os.path.join(root, 'meta.json'),
'rdme': os.path.join(root, 'README.md'),
'data_xls': os.path.join(root, 'default_of_credit_card_clients.xls'),
'data': os.path.join(root, 'default_of_credit_card_clients.csv'),
}
# Load the meta data from the meta json
with open(filenames['meta'], 'r') as f:
meta = json.load(f)
target_names = meta['target_names']
feature_names = meta['feature_names']
# Load the description from the README.
with open(filenames['rdme'], 'r') as f:
DESCR = f.read()
# Load the dataset from the EXCEL file.
df = pd.read_excel(filenames['data_xls'],header=None, skiprows=2, names=FEATURES)
df.to_csv(filenames['data'],header=False)
dataset = np.loadtxt(filenames['data'],delimiter=",")
# Extract the target from the data
data = dataset[:, 0:-1]
target = dataset[:, -1]
# Create the bunch object
return Bunch(
data=data,
target=target,
filenames=filenames,
target_names=target_names,
feature_names=feature_names,
DESCR=DESCR
)
# Save the dataset as a variable we can use.
dataset = load_data()
print dataset.data.shape
print dataset.target.shape
```
## Classification
Now that we have a dataset `Bunch` loaded and ready, we can begin the classification process. Let's attempt to build a classifier with kNN, SVM, and Random Forest classifiers.
```
from sklearn import metrics
from sklearn import cross_validation
from sklearn.cross_validation import KFold
from sklearn.svm import SVC
from sklearn.neighbors import KNeighborsClassifier
from sklearn.ensemble import RandomForestClassifier
def fit_and_evaluate(dataset, model, label, **kwargs):
"""
Because of the Scikit-Learn API, we can create a function to
do all of the fit and evaluate work on our behalf!
"""
start = time.time() # Start the clock!
scores = {'precision':[], 'recall':[], 'accuracy':[], 'f1':[]}
for train, test in KFold(dataset.data.shape[0], n_folds=12, shuffle=True):
X_train, X_test = dataset.data[train], dataset.data[test]
y_train, y_test = dataset.target[train], dataset.target[test]
estimator = model(**kwargs)
estimator.fit(X_train, y_train)
expected = y_test
predicted = estimator.predict(X_test)
# Append our scores to the tracker
scores['precision'].append(metrics.precision_score(expected, predicted, average="weighted"))
scores['recall'].append(metrics.recall_score(expected, predicted, average="weighted"))
scores['accuracy'].append(metrics.accuracy_score(expected, predicted))
scores['f1'].append(metrics.f1_score(expected, predicted, average="weighted"))
# Report
print "Build and Validation of {} took {:0.3f} seconds".format(label, time.time()-start)
print "Validation scores are as follows:\n"
print pd.DataFrame(scores).mean()
# Write official estimator to disk
estimator = model(**kwargs)
estimator.fit(dataset.data, dataset.target)
outpath = label.lower().replace(" ", "-") + ".pickle"
with open(outpath, 'w') as f:
pickle.dump(estimator, f)
print "\nFitted model written to:\n{}".format(os.path.abspath(outpath))
# Perform SVC Classification
#fit_and_evaluate(dataset, SVC, "CC Defaut - SVM Classifier")
# Perform kNN Classification
fit_and_evaluate(dataset, KNeighborsClassifier, "CC Defaut - kNN Classifier", n_neighbors=12)
# Perform Random Forest Classification
fit_and_evaluate(dataset, RandomForestClassifier, "CC Defaut - Random Forest Classifier")
```
| github_jupyter |
```
from google.colab import drive
drive.mount('/content/gdrive')
!ls gdrive/My\ Drive/real-estate
!cp -r gdrive/My\ Drive/real-estate .
!ls real-estate
import os
import glob
import re
from pathlib import Path
import numpy as np
import pandas as pd
glob.glob('real-estate/*')
filenames = glob.glob(str(Path() / 'real-estate' / '*'))
filenames
sorted(filenames)
sorted_filenames = sorted(
filenames,
key=lambda s: int(s[:-5].split('_')[-1]),
)
sorted_filenames
price1 = pd.read_excel(sorted_filenames[0], header=16)
price1.head()
price1.info()
price1 = pd.read_excel(
sorted_filenames[0],
header=16,
usecols=[
'시군구',
'주택유형',
'도로조건',
'연면적(㎡)',
'대지면적(㎡)',
'계약년월',
'거래금액(만원)',
'건축년도',
],
)
price1.columns = [
'si_gun_gu',
'housing_type',
'distance_to_road',
'total_floor_area',
'plottage',
'date_of_contract',
'price',
'construction_year'
]
price1.head()
price1 = pd.read_excel(
sorted_filenames[0],
header=16,
usecols=[
'시군구',
'주택유형',
'도로조건',
'연면적(㎡)',
'대지면적(㎡)',
'계약년월',
'거래금액(만원)',
'건축년도',
],
thousands=',',
)
price1.columns = [
'si_gun_gu',
'housing_type',
'distance_to_road',
'total_floor_area',
'plottage',
'date_of_contract',
'price',
'construction_year'
]
price1.head()
price1['price'].dtype
price1['construction_year'].describe()
price1['construction_year'].nsmallest(5)
price1.loc[price1['construction_year'] >= 1900, 'construction_year'].plot.hist()
(
price1[price1['construction_year'] > 1900]
.dropna()
.astype({'construction_year': int})
).head()
price1['distance_to_road'].unique()
price1['distance_to_road'].replace(
['8m미만', '12m미만', '25m미만', '-', '25m이상'],
[4.0, 10.0, 18.5, np.nan, 50.0],
).unique()
price1['date_of_contract'].head()
pd.to_datetime(price1['date_of_contract'].astype(str), format='%Y%m').head()
price1['housing_type'].unique()
price1['housing_type'].astype('category').head()
print(price1['housing_type'].memory_usage())
print(price1['housing_type'].astype('category').memory_usage())
prices = []
for filename in filenames:
prices.append(pd.read_excel(
filename,
header=16,
usecols=[
'시군구',
'주택유형',
'도로조건',
'연면적(㎡)',
'대지면적(㎡)',
'계약년월',
'거래금액(만원)',
'건축년도',
],
thousands=',',
dtype={
'주택유형': 'category',
}
).dropna())
price = pd.concat(prices).reset_index(drop=True)
price.columns = [
'si_gun_gu',
'housing_type',
'distance_to_road',
'total_floor_area',
'plottage',
'date_of_contract',
'price',
'construction_year'
]
price = price[price['construction_year'] > 1900].astype(
{'construction_year': int}
)
price['distance_to_road'] = price['distance_to_road'].replace(
['8m미만', '12m미만', '25m미만', '-', '25m이상'],
[4.0, 10.0, 18.5, np.nan, 50.0],
)
price = price.dropna().reset_index(drop=True)
price['date_of_contract'] = pd.to_datetime(
price['date_of_contract'].astype(str),
format='%Y%m'
)
price.head()
price.describe()
si_gun_gu = price['si_gun_gu'].str.split(expand=True)
si_gun_gu.head()
si_gun_gu.notna().sum()
si_gun_gu = price['si_gun_gu'].str.split(n=2, expand=True)
si_gun_gu.head()
price.info()
si_gun_gu.index[si_gun_gu[2].isna()]
si_gun_gu.isna().sum()
price.loc[60005]
si_gun_gu = si_gun_gu.fillna('')
si_gun_gu.columns = ['sido', 'sigungu', 'dongli']
si_gun_gu.head()
price = pd.concat([price, si_gun_gu], axis=1).drop(columns='si_gun_gu')
price.head()
```
# Visualization
```
!apt-get update -qq
!apt-get install fonts-nanum* -qq
import matplotlib as mpl
import matplotlib.font_manager as fm
import matplotlib.pyplot as plt
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
pd.plotting.register_matplotlib_converters()
fm._rebuild()
mpl.rc('font', family='NanumGothic')
price['housing_type'].value_counts()
fig, axes = plt.subplots(1, 2)
price['housing_type'].value_counts().plot.pie(
autopct='%.2f %%',
ax=axes[0]
)
price['housing_type'].value_counts().plot.bar(ax=axes[1])
fig.tight_layout()
fig, ax = plt.subplots()
price['price'].plot.hist(ax=ax)
# price.plot.hist(y='price', ax=ax)
ax.set(
xlabel='거래 금액(만 원)',
ylabel='거래 건수',
)
price['price'].nlargest(10)
price[price['price'] > 100_0000]
fig, ax = plt.subplots()
price[price['price'] < 25_0000].plot.hist(y='price', ax=ax)
ax.set(
xlabel='거래 금액(만 원)',
ylabel='거래 건수',
)
fig, ax = plt.subplots()
price[price['price'] > 250_000].plot.hist(y='price', ax=ax)
ax.set(
xlabel='거래 금액(만 원)',
ylabel='거래 건수',
)
price_lower = price[price['price'] < 25_0000]
fig, ax = plt.subplots()
price_lower['date_of_contract'].dt.month.value_counts()
(
price_lower['date_of_contract']
.dt.month
.value_counts()
.sort_index()
.plot.bar()
)
pd.plotting.scatter_matrix(
price_lower[['price', 'construction_year', 'total_floor_area', 'plottage']],
figsize=(10, 10),
)
```
| github_jupyter |
**NODAL POINT ANALYSIS:**
---
**Analysis With the Bottom Hole Node**
- Nodal analysis is performed on the principle
of pressure continuity.
- When the bottom-hole is used as a solution node in Nodal
analysis, the inflow performance is the well inflow performance relationship (IPR) and the outflow performance
is the tubing performance relationship (TPR)
**Well Data:**
```
# Given Data of Oil well
Pres = 3500 # psia (above Bubble Point)
D = 1.66 # (tubing ID)
Pwellhead = 500 # psia
J = 1 # (above bubble Point)
GLR = 1000 # scf/stb
WC = 0.25 # (watercut)
oil_API = 30 # degree
Yw = 1.05
Yg = 0.65
Bo = 1.2 # rb/stb
Twellhead = 100 # degree F
Tubing_shoe_depth = 5000 # ft
Tbottomhole = 150 # degree F
```
**Importing Required Libraries**
```
# Importing libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
```
**IPR MODEL USED:**
The most simplest model as the reservoir pressure is above Pb
Qo = J*(Pres - Pwf)
**TPR MODEL USED:**
Poettmann–Carpenter’s model which considers multiphase flow in the tubing. It uses friction factor but doesnot take liquid viscosity into account.
Pwf = Pwh + (Rho_mix + k/Rho_mix)H/144
where k = (f Qo^2 M^2) / (7.4137 10^10 D^5))
**Calculating Parameters that will not change during Iteration**
```
# Oil specific Gravity
Yo = 141.5/(131.5+oil_API)
# Mass associated with 1 stb of oil
GOR = GLR /(1-WC)
WOR = WC/(1-WC)
M = (350.17 * (Yo + 0.33 * Yw) )+ (GOR*Yg*0.0765)
# WELL HEAD parameters
Rs_head = Yg*pow(( (Pwellhead/18)* (pow(10, 0.0125*oil_API)/pow(10, 0.00091*Twellhead))), 1.2048)
Bo_head = 0.9759 + 0.00012*pow((Rs_head*pow(Yg/Yo, 0.5) + 1.25*Twellhead), 1.2)
Vm_head = 5.615*(Bo_head + WC * Bo) +(GOR - Rs_head)*(14.7/Pwellhead)*((Twellhead+460)/520)*(0.9318)
Rho_head = M/Vm_head
```
**ITERATION**
```
Ql = 500 # Assumed Initial Value
Aof = J*Pres # Max value of Q
# Conatainers for ipr, tpr pressures and flow rates
ipr = []
tpr = []
Q = []
while Ql<Aof:
Qo = Ql-Ql*WC
# PWf from IPR model
Pwf_ipr = Pres - Qo/J
ipr.append(Pwf_ipr)
# BOTTOM HOLE PARAMETERS
Rs_bottom = Yg*pow(( (Pwf_ipr/18)* (pow(10, 0.0125*oil_API)/pow(10, 0.00091*Tbottomhole))), 1.2048)
Bo_bottom = 0.9759 + 0.00012*pow((Rs_bottom*pow(Yg/Yo, 0.5) + 1.25*Tbottomhole), 1.2)
Vm_bottom = 5.615*(Bo_bottom + WC * Bo) +(GOR - Rs_bottom)*(14.7/Pwf_ipr)*((Tbottomhole+460)/520)*(0.9318)
Rho_bottom = M/Vm_bottom
Rho_mixture = (Rho_head + Rho_bottom)/2
# Inertial Force
rdv = 1.4737*pow(10,-5)*M*(Qo)*12/D
# Friction Factor
f = 4*(pow(10, 1.444 - 2.5*np.log10(rdv)))
# Friction term
K = f*Qo*Qo*M*M/(7.4137*pow(10,10)*pow(D/12,5))
# Pwf from TPR model
Pwf_tpr = Pwellhead + (Rho_mixture + K/Rho_mixture )*(Tubing_shoe_depth/144)
tpr.append(Pwf_tpr)
# For Q when Error is very less than that is our operational Point
error = abs(Pwf_ipr - Pwf_tpr)
if(error<4):print("Operational Bottom Hole Pressure: ",(Pwf_ipr)); print("Operational Flow Rate: ",(Qo))
Q.append(Qo)
Ql = Ql + 10
```
**TPR and IPR Plot**
```
# Plotting Curves
plt.style.use('default')
plt.plot(Q, ipr, label='IPR')
plt.plot(Q, tpr, label='TPR')
plt.title('Operational Point Visualization')
plt.xlabel('Qo, stb/day')
plt.ylabel("Pwf, Psia")
plt.legend()
plt.grid()
```
| github_jupyter |
```
#hide
#skip
! [[ -e /content ]] && pip install -Uqq fastai # upgrade fastai on colab
#default_exp vision.widgets
#export
from fastai.torch_basics import *
from fastai.data.all import *
from fastai.vision.core import *
from ipywidgets import HBox,VBox,widgets,Button,Checkbox,Dropdown,Layout,Box,Output,Label,FileUpload
#hide
from nbdev.showdoc import *
#export
_all_ = ['HBox','VBox','widgets','Button','Checkbox','Dropdown','Layout','Box','Output','Label','FileUpload']
```
# Vision widgets
> ipywidgets for images
```
#export
@patch
def __getitem__(self:Box, i): return self.children[i]
#export
def widget(im, *args, **layout):
"Convert anything that can be `display`ed by IPython into a widget"
o = Output(layout=merge(*args, layout))
with o: display(im)
return o
im = Image.open('images/puppy.jpg').to_thumb(256,512)
VBox([widgets.HTML('Puppy'),
widget(im, max_width="192px")])
#export
def _update_children(change):
for o in change['owner'].children:
if not o.layout.flex: o.layout.flex = '0 0 auto'
#export
def carousel(children=(), **layout):
"A horizontally scrolling carousel"
def_layout = dict(overflow='scroll hidden', flex_flow='row', display='flex')
res = Box([], layout=merge(def_layout, layout))
res.observe(_update_children, names='children')
res.children = children
return res
ts = [VBox([widget(im, max_width='192px'), Button(description='click')])
for o in range(3)]
carousel(ts, width='450px')
#export
def _open_thumb(fn, h, w): return Image.open(fn).to_thumb(h, w).convert('RGBA')
#export
class ImagesCleaner:
"A widget that displays all images in `fns` along with a `Dropdown`"
def __init__(self, opts=(), height=128, width=256, max_n=30):
opts = ('<Keep>', '<Delete>')+tuple(opts)
store_attr('opts,height,width,max_n')
self.widget = carousel(width='100%')
def set_fns(self, fns):
self.fns = L(fns)[:self.max_n]
ims = parallel(_open_thumb, self.fns, h=self.height, w=self.width, progress=False,
n_workers=min(len(self.fns)//10,defaults.cpus))
self.widget.children = [VBox([widget(im, height=f'{self.height}px'), Dropdown(
options=self.opts, layout={'width': 'max-content'})]) for im in ims]
def _ipython_display_(self): display(self.widget)
def values(self): return L(self.widget.children).itemgot(1).attrgot('value')
def delete(self): return self.values().argwhere(eq('<Delete>'))
def change(self):
idxs = self.values().argwhere(negate_func(in_(['<Delete>','<Keep>'])))
return idxs.zipwith(self.values()[idxs])
fns = get_image_files('images')
w = ImagesCleaner(('A','B'))
w.set_fns(fns)
w
w.delete(),w.change()
#export
def _get_iw_info(learn, ds_idx=0):
dl = learn.dls[ds_idx].new(shuffle=False, drop_last=False)
inp,probs,targs,preds,losses = learn.get_preds(dl=dl, with_input=True, with_loss=True, with_decoded=True)
inp,targs = L(zip(*dl.decode_batch((inp,targs), max_n=9999)))
return L([dl.dataset.items,targs,losses]).zip()
#export
@delegates(ImagesCleaner)
class ImageClassifierCleaner(GetAttr):
"A widget that provides an `ImagesCleaner` with a CNN `Learner`"
def __init__(self, learn, **kwargs):
vocab = learn.dls.vocab
self.default = self.iw = ImagesCleaner(vocab, **kwargs)
self.dd_cats = Dropdown(options=vocab)
self.dd_ds = Dropdown(options=('Train','Valid'))
self.iwis = _get_iw_info(learn,0),_get_iw_info(learn,1)
self.dd_ds.observe(self.on_change_ds, 'value')
self.dd_cats.observe(self.on_change_ds, 'value')
self.on_change_ds()
self.widget = VBox([self.dd_cats, self.dd_ds, self.iw.widget])
def _ipython_display_(self): display(self.widget)
def on_change_ds(self, change=None):
info = L(o for o in self.iwis[self.dd_ds.index] if o[1]==self.dd_cats.value)
self.iw.set_fns(info.sorted(2, reverse=True).itemgot(0))
```
# Export -
```
#hide
from nbdev.export import notebook2script
notebook2script()
```
| github_jupyter |
```
#IMPORT SEMUA LIBRARY DISINI
#IMPORT LIBRARY PANDAS
import pandas as pd
#IMPORT LIBRARY POSTGRESQL
import psycopg2
from psycopg2.extensions import ISOLATION_LEVEL_AUTOCOMMIT
#IMPORT LIBRARY CHART
from matplotlib import pyplot as plt
from matplotlib import style
#IMPORT LIBRARY PDF
from fpdf import FPDF
#IMPORT LIBRARY BASEPATH
import io
#IMPORT LIBRARY BASE64 IMG
import base64
#IMPORT LIBRARY NUMPY
import numpy as np
#IMPORT LIBRARY EXCEL
import xlsxwriter
#IMPORT LIBRARY SIMILARITAS
import n0similarities as n0
#FUNGSI UNTUK MENGUPLOAD DATA DARI CSV KE POSTGRESQL
def uploadToPSQL(host, username, password, database, port, table, judul, filePath, name, subjudul, dataheader, databody):
#TEST KONEKSI KE DATABASE
try:
for t in range(0, len(table)):
#DATA DIJADIKAN LIST
rawstr = [tuple(x) for x in zip(dataheader, databody[t])]
#KONEKSI KE DATABASE
connection = psycopg2.connect(user=username,password=password,host=host,port=port,database=database)
cursor = connection.cursor()
connection.set_isolation_level(ISOLATION_LEVEL_AUTOCOMMIT);
#CEK TABLE
cursor.execute("SELECT * FROM information_schema.tables where table_name=%s", (table[t],))
exist = bool(cursor.rowcount)
#KALAU ADA DIHAPUS DULU, TERUS DICREATE ULANG
if exist == True:
cursor.execute("DROP TABLE "+ table[t] + " CASCADE")
cursor.execute("CREATE TABLE "+table[t]+" (index SERIAL, tanggal date, total varchar);")
#KALAU GA ADA CREATE DATABASE
else:
cursor.execute("CREATE TABLE "+table[t]+" (index SERIAL, tanggal date, total varchar);")
#MASUKAN DATA KE DATABASE YANG TELAH DIBUAT
cursor.execute('INSERT INTO '+table[t]+'(tanggal, total) values ' +str(rawstr)[1:-1])
#JIKA BERHASIL SEMUA AKAN MENGHASILKAN KELUARAN BENAR (TRUE)
return True
#JIKA KONEKSI GAGAL
except (Exception, psycopg2.Error) as error :
return error
#TUTUP KONEKSI
finally:
if(connection):
cursor.close()
connection.close()
#FUNGSI UNTUK MEMBUAT CHART, DATA YANG DIAMBIL DARI DATABASE DENGAN MENGGUNAKAN ORDER DARI TANGGAL DAN JUGA LIMIT
#DISINI JUGA MEMANGGIL FUNGSI MAKEEXCEL DAN MAKEPDF
def makeChart(host, username, password, db, port, table, judul, filePath, name, subjudul, A2, B2, C2, D2, E2, F2, G2, H2,I2, J2, K2, limitdata, wilayah, tabledata, basePath):
try:
datarowsend = []
for t in range(0, len(table)):
#TEST KONEKSI KE DATABASE
connection = psycopg2.connect(user=username,password=password,host=host,port=port,database=db)
cursor = connection.cursor()
#MENGAMBIL DATA DARI DATABASE DENGAN LIMIT YANG SUDAH DIKIRIMKAN DARI VARIABLE DIBAWAH
postgreSQL_select_Query = "SELECT * FROM "+table[t]+" ORDER BY tanggal DESC LIMIT " + str(limitdata)
cursor.execute(postgreSQL_select_Query)
mobile_records = cursor.fetchall()
uid = []
lengthx = []
lengthy = []
#MENYIMPAN DATA DARI DATABASE KE DALAM VARIABLE
for row in mobile_records:
uid.append(row[0])
lengthx.append(row[1])
lengthy.append(row[2])
datarowsend.append(mobile_records)
#JUDUL CHART
judulgraf = A2 + " " + wilayah[t]
#bar
style.use('ggplot')
fig, ax = plt.subplots()
#DATA CHART DIMASUKAN DISINI
ax.bar(uid, lengthy, align='center')
#JUDUL CHART
ax.set_title(judulgraf)
ax.set_ylabel('Total')
ax.set_xlabel('Tanggal')
ax.set_xticks(uid)
ax.set_xticklabels((lengthx))
b = io.BytesIO()
#BUAT CHART MENJADI FORMAT PNG
plt.savefig(b, format='png', bbox_inches="tight")
#CHART DIJADIKAN BASE64
barChart = base64.b64encode(b.getvalue()).decode("utf-8").replace("\n", "")
plt.show()
#line
#DATA CHART DIMASUKAN DISINI
plt.plot(lengthx, lengthy)
plt.xlabel('Tanggal')
plt.ylabel('Total')
#JUDUL CHART
plt.title(judulgraf)
plt.grid(True)
l = io.BytesIO()
#CHART DIJADIKAN GAMBAR
plt.savefig(l, format='png', bbox_inches="tight")
#GAMBAR DIJADIKAN BAS64
lineChart = base64.b64encode(l.getvalue()).decode("utf-8").replace("\n", "")
plt.show()
#pie
#JUDUL CHART
plt.title(judulgraf)
#DATA CHART DIMASUKAN DISINI
plt.pie(lengthy, labels=lengthx, autopct='%1.1f%%',
shadow=True, startangle=180)
plt.plot(legend=None)
plt.axis('equal')
p = io.BytesIO()
#CHART DIJADIKAN GAMBAR
plt.savefig(p, format='png', bbox_inches="tight")
#CHART DICONVERT KE BASE64
pieChart = base64.b64encode(p.getvalue()).decode("utf-8").replace("\n", "")
plt.show()
#CHART DISIMPAN KE DIREKTORI DIJADIKAN FORMAT PNG
#BARCHART
bardata = base64.b64decode(barChart)
barname = basePath+'jupyter/CEIC/23. Sektor Petambangan dan Manufaktur/img/'+name+''+table[t]+'-bar.png'
with open(barname, 'wb') as f:
f.write(bardata)
#LINECHART
linedata = base64.b64decode(lineChart)
linename = basePath+'jupyter/CEIC/23. Sektor Petambangan dan Manufaktur/img/'+name+''+table[t]+'-line.png'
with open(linename, 'wb') as f:
f.write(linedata)
#PIECHART
piedata = base64.b64decode(pieChart)
piename = basePath+'jupyter/CEIC/23. Sektor Petambangan dan Manufaktur/img/'+name+''+table[t]+'-pie.png'
with open(piename, 'wb') as f:
f.write(piedata)
#MEMANGGIL FUNGSI EXCEL
makeExcel(datarowsend, A2, B2, C2, D2, E2, F2, G2, H2,I2, J2, K2, name, limitdata, table, wilayah, basePath)
#MEMANGGIL FUNGSI PDF
makePDF(datarowsend, judul, barChart, lineChart, pieChart, name, subjudul, A2, B2, C2, D2, E2, F2, G2, H2,I2, J2, K2, limitdata, table, wilayah, basePath)
#JIKA KONEKSI GAGAL
except (Exception, psycopg2.Error) as error :
print (error)
#TUTUP KONEKSI
finally:
if(connection):
cursor.close()
connection.close()
#FUNGSI UNTUK MEMBUAT PDF YANG DATANYA BERASAL DARI DATABASE DIJADIKAN FORMAT EXCEL TABLE F2
#PLUGIN YANG DIGUNAKAN ADALAH FPDF
def makePDF(datarow, judul, bar, line, pie, name, subjudul, A2, B2, C2, D2, E2, F2, G2, H2,I2, J2, K2, lengthPDF, table, wilayah, basePath):
#PDF DIATUR DENGAN SIZE A4 DAN POSISI LANDSCAPE
pdf = FPDF('L', 'mm', [210,297])
#TAMBAH HALAMAN PDF
pdf.add_page()
#SET FONT DAN JUGA PADDING
pdf.set_font('helvetica', 'B', 20.0)
pdf.set_xy(145.0, 15.0)
#TAMPILKAN JUDUL PDF
pdf.cell(ln=0, h=2.0, align='C', w=10.0, txt=judul, border=0)
#SET FONT DAN JUGA PADDING
pdf.set_font('arial', '', 14.0)
pdf.set_xy(145.0, 25.0)
#TAMPILKAN SUB JUDUL PDF
pdf.cell(ln=0, h=2.0, align='C', w=10.0, txt=subjudul, border=0)
#BUAT GARIS DIBAWAH SUB JUDUL
pdf.line(10.0, 30.0, 287.0, 30.0)
pdf.set_font('times', '', 10.0)
pdf.set_xy(17.0, 37.0)
pdf.set_font('Times','B',11.0)
pdf.ln(0.5)
th1 = pdf.font_size
#BUAT TABLE DATA DATA DI DPF
pdf.cell(100, 2*th1, "Kategori", border=1, align='C')
pdf.cell(177, 2*th1, A2, border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Region", border=1, align='C')
pdf.cell(177, 2*th1, B2, border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Frekuensi", border=1, align='C')
pdf.cell(177, 2*th1, C2, border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Unit", border=1, align='C')
pdf.cell(177, 2*th1, D2, border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Sumber", border=1, align='C')
pdf.cell(177, 2*th1, E2, border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Status", border=1, align='C')
pdf.cell(177, 2*th1, F2, border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "ID Seri", border=1, align='C')
pdf.cell(177, 2*th1, G2, border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Kode SR", border=1, align='C')
pdf.cell(177, 2*th1, H2, border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Tanggal Obs. Pertama", border=1, align='C')
pdf.cell(177, 2*th1, str(I2.date()), border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Tanggal Obs. Terakhir ", border=1, align='C')
pdf.cell(177, 2*th1, str(J2.date()), border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Waktu pembaruan terakhir", border=1, align='C')
pdf.cell(177, 2*th1, str(K2.date()), border=1, align='C')
pdf.ln(2*th1)
pdf.set_xy(17.0, 125.0)
pdf.set_font('Times','B',11.0)
epw = pdf.w - 2*pdf.l_margin
col_width = epw/(lengthPDF+1)
pdf.ln(0.5)
th = pdf.font_size
#HEADER TABLE DATA F2
pdf.cell(col_width, 2*th, str("Wilayah"), border=1, align='C')
#TANGAL HEADER DI LOOPING
for row in datarow[0]:
pdf.cell(col_width, 2*th, str(row[1]), border=1, align='C')
pdf.ln(2*th)
#ISI TABLE F2
for w in range(0, len(table)):
data=list(datarow[w])
pdf.set_font('Times','B',10.0)
pdf.set_font('Arial','',9)
pdf.cell(col_width, 2*th, wilayah[w], border=1, align='C')
#DATA BERDASARKAN TANGGAL
for row in data:
pdf.cell(col_width, 2*th, str(row[2]), border=1, align='C')
pdf.ln(2*th)
#PEMANGGILAN GAMBAR
for s in range(0, len(table)):
col = pdf.w - 2*pdf.l_margin
pdf.ln(2*th)
widthcol = col/3
#TAMBAH HALAMAN
pdf.add_page()
#DATA GAMBAR BERDASARKAN DIREKTORI DIATAS
pdf.image(basePath+'jupyter/CEIC/23. Sektor Petambangan dan Manufaktur/img/'+name+''+table[s]+'-bar.png', link='', type='',x=8, y=80, w=widthcol)
pdf.set_xy(17.0, 144.0)
col = pdf.w - 2*pdf.l_margin
pdf.image(basePath+'jupyter/CEIC/23. Sektor Petambangan dan Manufaktur/img/'+name+''+table[s]+'-line.png', link='', type='',x=103, y=80, w=widthcol)
pdf.set_xy(17.0, 144.0)
col = pdf.w - 2*pdf.l_margin
pdf.image(basePath+'jupyter/CEIC/23. Sektor Petambangan dan Manufaktur/img/'+name+''+table[s]+'-pie.png', link='', type='',x=195, y=80, w=widthcol)
pdf.ln(4*th)
#PDF DIBUAT
pdf.output(basePath+'jupyter/CEIC/23. Sektor Petambangan dan Manufaktur/pdf/'+A2+'.pdf', 'F')
#FUNGSI MAKEEXCEL GUNANYA UNTUK MEMBUAT DATA YANG BERASAL DARI DATABASE DIJADIKAN FORMAT EXCEL TABLE F2
#PLUGIN YANG DIGUNAKAN ADALAH XLSXWRITER
def makeExcel(datarow, A2, B2, C2, D2, E2, F2, G2, H2, I2, J2, K2, name, limit, table, wilayah, basePath):
#BUAT FILE EXCEL
workbook = xlsxwriter.Workbook(basePath+'jupyter/CEIC/23. Sektor Petambangan dan Manufaktur/excel/'+A2+'.xlsx')
#BUAT WORKSHEET EXCEL
worksheet = workbook.add_worksheet('sheet1')
#SETTINGAN UNTUK BORDER DAN FONT BOLD
row1 = workbook.add_format({'border': 2, 'bold': 1})
row2 = workbook.add_format({'border': 2})
#HEADER UNTUK TABLE EXCEL F2
header = ["Wilayah", "Kategori","Region","Frekuensi","Unit","Sumber","Status","ID Seri","Kode SR","Tanggal Obs. Pertama","Tanggal Obs. Terakhir ","Waktu pembaruan terakhir"]
#DATA DATA DITAMPUNG PADA VARIABLE
for rowhead2 in datarow[0]:
header.append(str(rowhead2[1]))
#DATA HEADER DARI VARIABLE DIMASUKAN KE SINI UNTUK DITAMPILKAN BERDASARKAN ROW DAN COLUMN
for col_num, data in enumerate(header):
worksheet.write(0, col_num, data, row1)
#DATA ISI TABLE F2 DITAMPILKAN DISINI
for w in range(0, len(table)):
data=list(datarow[w])
body = [wilayah[w], A2, B2, C2, D2, E2, F2, G2, H2, str(I2.date()), str(J2.date()), str(K2.date())]
for rowbody2 in data:
body.append(str(rowbody2[2]))
for col_num, data in enumerate(body):
worksheet.write(w+1, col_num, data, row2)
#FILE EXCEL DITUTUP
workbook.close()
#DISINI TEMPAT AWAL UNTUK MENDEFINISIKAN VARIABEL VARIABEL SEBELUM NANTINYA DIKIRIM KE FUNGSI
#PERTAMA MANGGIL FUNGSI UPLOADTOPSQL DULU, KALAU SUKSES BARU MANGGIL FUNGSI MAKECHART
#DAN DI MAKECHART MANGGIL FUNGSI MAKEEXCEL DAN MAKEPDF
#BASE PATH UNTUK NANTINYA MENGCREATE FILE ATAU MEMANGGIL FILE
basePath = 'C:/Users/ASUS/Documents/bappenas/'
#FILE SIMILARITY WILAYAH
filePathwilayah = basePath+'data mentah/CEIC/allwilayah.xlsx';
#BACA FILE EXCEL DENGAN PANDAS
readexcelwilayah = pd.read_excel(filePathwilayah)
dfwilayah = list(readexcelwilayah.values)
readexcelwilayah.fillna(0)
allwilayah = []
#PEMILIHAN JENIS DATA, APA DATA ITU PROVINSI, KABUPATEN, KECAMATAN ATAU KELURAHAN
tipewilayah = 'prov'
if tipewilayah == 'prov':
for x in range(0, len(dfwilayah)):
allwilayah.append(dfwilayah[x][1])
elif tipewilayah=='kabkot':
for x in range(0, len(dfwilayah)):
allwilayah.append(dfwilayah[x][3])
elif tipewilayah == 'kec':
for x in range(0, len(dfwilayah)):
allwilayah.append(dfwilayah[x][5])
elif tipewilayah == 'kel':
for x in range(0, len(dfwilayah)):
allwilayah.append(dfwilayah[x][7])
semuawilayah = list(set(allwilayah))
#SETTING VARIABLE UNTUK DATABASE DAN DATA YANG INGIN DIKIRIMKAN KE FUNGSI DISINI
name = "01. Produksi Industri (BAA001-BAA008)"
host = "localhost"
username = "postgres"
password = "1234567890"
port = "5432"
database = "ceic"
judul = "Produk Domestik Bruto (AA001-AA007)"
subjudul = "Badan Perencanaan Pembangunan Nasional"
filePath = basePath+'data mentah/CEIC/23. Sektor Petambangan dan Manufaktur/'+name+'.xlsx';
limitdata = int(8)
readexcel = pd.read_excel(filePath)
tabledata = []
wilayah = []
databody = []
#DATA EXCEL DIBACA DISINI DENGAN MENGGUNAKAN PANDAS
df = list(readexcel.values)
head = list(readexcel)
body = list(df[0])
readexcel.fillna(0)
#PILIH ROW DATA YANG INGIN DITAMPILKAN
rangeawal = 106
rangeakhir = 107
rowrange = range(rangeawal, rangeakhir)
#INI UNTUK MEMFILTER APAKAH DATA YANG DIPILIH MEMILIKI SIMILARITAS ATAU TIDAK
#ISIKAN 'WILAYAH' UNTUK SIMILARITAS
#ISIKAN BUKAN WILAYAH JIKA BUKAN WILAYAH
jenisdata = "Indonesia"
#ROW DATA DI LOOPING UNTUK MENDAPATKAN SIMILARITAS WILAYAH
#JIKA VARIABLE JENISDATA WILAYAH AKAN MASUK KESINI
if jenisdata == 'Wilayah':
for x in rowrange:
rethasil = 0
big_w = 0
for w in range(0, len(semuawilayah)):
namawilayah = semuawilayah[w].lower().strip()
nama_wilayah_len = len(namawilayah)
hasil = n0.get_levenshtein_similarity(df[x][0].lower().strip()[nama_wilayah_len*-1:], namawilayah)
if hasil > rethasil:
rethasil = hasil
big_w = w
wilayah.append(semuawilayah[big_w].capitalize())
tabledata.append('produkdomestikbruto_'+semuawilayah[big_w].lower().replace(" ", "") + "" + str(x))
testbody = []
for listbody in df[x][11:]:
if ~np.isnan(listbody) == False:
testbody.append(str('0'))
else:
testbody.append(str(listbody))
databody.append(testbody)
#JIKA BUKAN WILAYAH MASUK KESINI
else:
for x in rowrange:
wilayah.append(jenisdata.capitalize())
tabledata.append('produkdomestikbruto_'+jenisdata.lower().replace(" ", "") + "" + str(x))
testbody = []
for listbody in df[x][11:]:
if ~np.isnan(listbody) == False:
testbody.append(str('0'))
else:
testbody.append(str(listbody))
databody.append(testbody)
#HEADER UNTUK PDF DAN EXCEL
A2 = "Data Migas"
B2 = df[rangeawal][1]
C2 = df[rangeawal][2]
D2 = df[rangeawal][3]
E2 = df[rangeawal][4]
F2 = df[rangeawal][5]
G2 = df[rangeawal][6]
H2 = df[rangeawal][7]
I2 = df[rangeawal][8]
J2 = df[rangeawal][9]
K2 = df[rangeawal][10]
#DATA ISI TABLE F2
dataheader = []
for listhead in head[11:]:
dataheader.append(str(listhead))
#FUNGSI UNTUK UPLOAD DATA KE SQL, JIKA BERHASIL AKAN MAMANGGIL FUNGSI UPLOAD CHART
sql = uploadToPSQL(host, username, password, database, port, tabledata, judul, filePath, name, subjudul, dataheader, databody)
if sql == True:
makeChart(host, username, password, database, port, tabledata, judul, filePath, name, subjudul, A2, B2, C2, D2, E2, F2, G2, H2,I2, J2, K2, limitdata, wilayah, tabledata, basePath)
else:
print(sql)
```
| github_jupyter |
# Interact Exercise 4
## Imports
```
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
from IPython.html.widgets import interact, interactive, fixed
from IPython.display import display
```
## Line with Gaussian noise
Write a function named `random_line` that creates `x` and `y` data for a line with y direction random noise that has a normal distribution $N(0,\sigma^2)$:
$$
y = m x + b + N(0,\sigma^2)
$$
Be careful about the `sigma=0.0` case.
```
def random_line(m, b, sigma, size=10):
"""Create a line y = m*x + b + N(0,sigma**2) between x=[-1.0,1.0]
Parameters
----------
m : float
The slope of the line.
b : float
The y-intercept of the line.
sigma : float
The standard deviation of the y direction normal distribution noise.
size : int
The number of points to create for the line.
Returns
-------
x : array of floats
The array of x values for the line with `size` points.
y : array of floats
The array of y values for the lines with `size` points.
"""
# YOUR CODE HERE
#raise NotImplementedError()
x = np.linspace(-1.0,1.0,size)
if sigma==0:
y=m*x+b
else:
#np.random.normal() creates normal distribution array
y = (m*x)+b+np.random.normal(0.0, sigma**2, size)
return x,y
m = 0.0; b = 1.0; sigma=0.0; size=3
x, y = random_line(m, b, sigma, size)
assert len(x)==len(y)==size
assert list(x)==[-1.0,0.0,1.0]
assert list(y)==[1.0,1.0,1.0]
sigma = 1.0
m = 0.0; b = 0.0
size = 500
x, y = random_line(m, b, sigma, size)
assert np.allclose(np.mean(y-m*x-b), 0.0, rtol=0.1, atol=0.1)
assert np.allclose(np.std(y-m*x-b), sigma, rtol=0.1, atol=0.1)
```
Write a function named `plot_random_line` that takes the same arguments as `random_line` and creates a random line using `random_line` and then plots the `x` and `y` points using Matplotlib's `scatter` function:
* Make the marker color settable through a `color` keyword argument with a default of `red`.
* Display the range $x=[-1.1,1.1]$ and $y=[-10.0,10.0]$.
* Customize your plot to make it effective and beautiful.
```
def ticks_out(ax):
"""Move the ticks to the outside of the box."""
ax.get_xaxis().set_tick_params(direction='out', width=1, which='both')
ax.get_yaxis().set_tick_params(direction='out', width=1, which='both')
def plot_random_line(m, b, sigma, size=10, color='red'):
"""Plot a random line with slope m, intercept b and size points."""
# YOUR CODE HERE
#raise NotImplementedError()
x,y=random_line(m, b, sigma, size)
plt.scatter(x,y,color=color)
plt.xlim(-1.1,1.1)
plt.ylim(-10.0,10.0)
plt.box(False)
plt.xlabel('x')
plt.ylabel('y(x)')
plt.title('Random Line')
plt.tick_params(axis='y', right='off', direction='out')
plt.tick_params(axis='x', top='off', direction='out')
plt.grid(True)
plot_random_line(5.0, -1.0, 2.0, 50)
assert True # use this cell to grade the plot_random_line function
```
Use `interact` to explore the `plot_random_line` function using:
* `m`: a float valued slider from `-10.0` to `10.0` with steps of `0.1`.
* `b`: a float valued slider from `-5.0` to `5.0` with steps of `0.1`.
* `sigma`: a float valued slider from `0.0` to `5.0` with steps of `0.01`.
* `size`: an int valued slider from `10` to `100` with steps of `10`.
* `color`: a dropdown with options for `red`, `green` and `blue`.
```
# YOUR CODE HERE
#raise NotImplementedError()
interact(plot_random_line, m=(-10.0,10.0), b=(-5.0,5.0),sigma=(0.0,5.0,0.01),size=(10,100,10), color={'red':'r','blue':'b','green':'g'})
#### assert True # use this cell to grade the plot_random_line interact
```
| github_jupyter |
# Udemy Amazon Reviews extraction
```
import requests # Importing requests to extract content from a url
from bs4 import BeautifulSoup as bs
import re
import nltk
from nltk.corpus import stopwords
import matplotlib.pyplot as plt
from wordcloud import WordCloud
# creating empty reviews list
iphone_reviews=[]
#forest = ["the","king","of","jungle"]
# Data extraction - Amazon reviews extraction
for i in range(1,20):
ip=[]
url = "https://www.amazon.in/Test-Exclusive-669/product-reviews/B07HGH8ML7/ref=cm_cr_getr_d_paging_btm_prev"
if i==1:
url = url+"_"+str(i)+"?ie=UTF8&reviewerType=all_reviews&pageNumber="+str(i)
else:
url = url+"_"+str(i)+"?ie=UTF8&reviewerType=all_reviews&pageNumber="+str(i)
#print(url)
response = requests.get(url)
soup = bs(response.content, "html.parser")# creating soup object to iterate over the extracted content
reviews = soup.findAll("span", attrs={"class","a-size-base review-text review-text-content"})# Extracting the content under specific
for i in range(len(reviews)):
iphone_reviews.append(reviews[i].text)
print(iphone_reviews)# adding the reviews of one page to empty list which in future contai
iphone_reviews
# Data cleansing and Wordcloud
# writing reviews in a text file
with open("samsungm21.txt","w",encoding='utf8') as output:
output.write(str(iphone_reviews))
# Joining all the reviews into single paragraph
ip_rev_string = " ".join(iphone_reviews)
ip_rev_string
# Removing unwanted symbols incase if exists
ip_rev_string = re.sub("[^A-Za-z" "]+"," ",ip_rev_string).lower()
ip_rev_string = re.sub("[0-9" "]+"," ",ip_rev_string)
ip_rev_string
# words that contained in iphone 8 plus reviews
ip_reviews_words = ip_rev_string.split(" ")
ip_reviews_words
# stop_words = stopwords.words('english') can use this command if nltk loaded
with open("C:\\Users\\V Sudheer Kumar\\stopwords_en.txt") as sw:
stopwords = sw.read()
stopwords = stopwords.split("\n")
temp = ["this","is","awsome","Data","Science"]
[i for i in temp if i not in "is"]
stopwords
ip_reviews_words = [w for w in ip_reviews_words if not w in stopwords]
# Joining all the reviews into single paragraph
ip_rev_string = " ".join(ip_reviews_words)
ip_rev_string
# WordCloud can be performed on the string inputs. That is the reason we have combined
# entire reviews into single paragraph
# Simple word cloud
wordcloud_ip = WordCloud(
background_color='black',
width=1800,
height=1400
).generate(ip_rev_string)
plt.imshow(wordcloud_ip)
#$
# positive words # Choose the path for +ve words stored in system
with open("C:\\Users\\V Sudheer Kumar\\Documents\\poswords.txt","r") as pos:
poswords = pos.read().split("\n")
poswords = poswords[36:]
# negative words # Choose the path for -ve words stored in system
with open("C:\\Users\\V Sudheer Kumar\\Documents\\negwords.txt","r") as neg:
negwords = neg.read().split("\n")
negwords = negwords[37:]
# negative word cloud
# Choosing the only words which are present in negwords
ip_neg_in_neg = " ".join([w for w in ip_reviews_words if w in negwords])
wordcloud_neg_in_neg = WordCloud(
background_color='black',
width=1800,
height=1400
).generate(ip_neg_in_neg)
plt.figure(1)
plt.imshow(wordcloud_neg_in_neg)
# positive word cloud
# Choosing the only words which are present in poswords
ip_pos_in_pos = " ".join([w for w in ip_reviews_words if w in poswords])
wordcloud_pos_in_pos = WordCloud(
background_color='black',
width=1800,
height=1400
).generate(ip_pos_in_pos)
plt.figure(2)
plt.imshow(wordcloud_pos_in_pos)
# With white background
# negative word cloud
# Choosing the only words which are present in negwords
ip_neg_in_neg = " ".join([w for w in ip_reviews_words if w in negwords])
wordcloud_neg_in_neg = WordCloud(
background_color='white',
width=1800,
height=1400
).generate(ip_neg_in_neg)
plt.figure(1)
plt.imshow(wordcloud_neg_in_neg)
# positive word cloud
# Choosing the only words which are present in poswords
ip_pos_in_pos = " ".join([w for w in ip_reviews_words if w in poswords])
wordcloud_pos_in_pos = WordCloud(
background_color='white',
width=1800,
height=1400
).generate(ip_pos_in_pos)
plt.figure(2)
plt.imshow(wordcloud_pos_in_pos)
```
| github_jupyter |
# Translating Story Map from one language to another using Deep Learning
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc">
<ul class="toc-item">
<li><span><a href="#Introduction" data-toc-modified-id="Introduction-1">Introduction</a></span></li>
<li><span><a href="#Prerequisites" data-toc-modified-id="Prerequisites-2">Prerequisites</a></span></li>
<li><span><a href="#Imports" data-toc-modified-id="Imports-3">Imports</a></span></li>
<li><span><a href="#Translate-Story-Map-from-English-to-Spanish" data-toc-modified-id="Translate-Story-Map-from-English-to-Spanish-4">Translate Story Map from English to Spanish</a></span></li>
<ul class="toc-item">
<li><span><a href="#Connect-to-GIS-and-clone-Story-Map" data-toc-modified-id="Connect-to-GIS-and-clone-Story-Map-4.1">Connect to GIS and clone Story Map</a></span></li>
<li><span><a href="#Instantiate-text-translator" data-toc-modified-id="Instantiate-text-translator-4.2">Instantiate text translator</a></span></li>
<li><span><a href="#Translate-Story-Map-content" data-toc-modified-id="Translate-Story-Map-content-4.3">Translate Story Map content</a></span></li>
<li><span><a href="#Update-cloned-Story-Map-item" data-toc-modified-id="Update-cloned-Story-Map-item-4.4">Update cloned Story Map item</a></span></li>
</ul>
<li><span><a href="#Conclusion" data-toc-modified-id="Conclusion-5">Conclusion</a></span></li>
<li><span><a href="#References" data-toc-modified-id="References-6">References</a></span></li>
</ul>
</div>
# Introduction
A [story map](https://www.esri.com/en-us/arcgis/products/arcgis-storymaps/overview) is a web map that is created for a given context with supporting information so that it becomes a stand-alone resource. It integrates maps, legends, text, photos, and video. Story maps can be built using Esri's story map templates and are a great way to quickly build useful information products tailored to one's organization needs. Using the templates, one can publish a story map without writing any code. One can simply create a web map, supply the text and images for the story, and configure the template files to create a story map.
Sometimes, there is a need to convert the text of a story map from one language to another so that it can be understood by nonnative language speaker as well. This can be done either by employing a human translator or by using a machine translation system to automatically convert the text from one language to another.
Machine translation is a sub-field of computational linguistics that deals with the problem of translating an input text or speech from one language to another. With the recent advancements in **Natural Language Processing (NLP)** and **Deep Learning**, it is now possible for a machine translation system to reach human like performance in translating a text from one language to another.
In this notebook, we will pick a story map written in English language, and create another story map with the text translated to Spanish language using the `arcgis.learn.text`'s **TextTranslator** class. The **TextTranslator** class is part of inference-only classes offerered by the `arcgis.learn.text` submodule. These inference-only classes offer a simple API dedicated to several **Natural Language Processing (NLP)** tasks including **Masked Language Modeling**, **Text Generation**, **Sentiment Analysis**, **Summarization**, **Machine Translation** and **Question Answering**.
# Prerequisites
- Inferencing workflows for Inference-only Text models of `arcgis.learn.text` submodule is based on [Hugging Face Transformers](https://huggingface.co/transformers/v3.0.2/index.html) library.
- Refer to the section [Install deep learning dependencies of arcgis.learn module](https://developers.arcgis.com/python/guide/install-and-set-up/#Install-deep-learning-dependencies) for detailed explanation about deep learning dependencies.
- [Beautiful Soup](https://anaconda.org/anaconda/beautifulsoup4) python library to pull text out from the HTML content of the story map.
- **Choosing a pretrained model**: Depending on the task and the language of the input text, user might need to choose an appropriate transformer backbone to generate desired inference. This [link](https://huggingface.co/models?search=helsinki) lists out all the pretrained models offered by [Hugging Face Transformers](https://huggingface.co/transformers/v3.0.2/index.html) library that allows translation from a source language to one or more target languages.
# Imports
```
from arcgis import GIS
from bs4 import BeautifulSoup
from arcgis.learn.text import TextTranslator
```
# Translate Story Map from English to Spanish
In this notebook we have picked up a story map written in **English** language, which talks about the near-term interim improvements (for the new bicycle and pedestrian bridge design over Lady Bird Lake) to South Pleasant Valley Road. Our goal will be to create a clone of this story map with the text translated to **Spanish** language
## Connect to GIS and clone Story Map
To achieve our goal, we will first connect to an ArcGIS online account, get the desired content by passing the appropriate item-id and cloning the item into our GIS account. We will then apply the `TextTranslator` model of `arcgis.learn.text` submodule to this cloned item to convert the content of the story map to **Spanish** language.
```
agol = GIS()
gis = GIS('home')
storymapitem = agol.content.get('c8eef2a96c88489c92010a63d0944881')
storymapitem
cloned_items = gis.content.clone_items([storymapitem], search_existing_items=False)
cloned_items
cloned_item = cloned_items[0]
cloned_item
cloned_item.id
```
## Instantiate text translator
Next, we will instantiate the class object for the `TextTranslator` model. We wish to translate text from **English** language into **Spanish** language. So will invoke the object by passing the corresponding ISO language codes [[1]](#References) in the model constructor.
```
translator = TextTranslator(source_language='en', target_language='es')
```
We will also write some helper functions to help translate the content of story map into the desired language. The `replace` function is a recursive [[2]](#References) function that accepts a json (story map item dictionary) object `obj` and applies the function `func` on the values `v` of the `keys` list passed in the `replace` function argument.
```
def replace(obj, keys, func):
return {k: replace(func(v) if k in keys else v, keys, func)
for k,v in obj.items()} if isinstance(obj, dict) else obj
```
The `translate` function will translate the English `text` (passed in the function argument) into Spanish language. The story map text sometimes contain text wrapped inside HTML [[3]](#References) tags. We will use `BeautifulSoup` library to get the non-HTML part of the input text content and use the `translator`'s `translate` method to translate the non-HTML part of the input text into desired language (Spanish in this case).
```
def translate(text):
if text == '':
return text
soup = BeautifulSoup(text, "html.parser")
for txt in soup.find_all(text=True):
translation = translator.translate(txt)[0]['translated_text'] if txt.strip() != '' else txt
txt.string.replace_with(" " + translation + " ")
return str(soup)
```
## Translate Story Map content
We will call the story map item's `get_data()` method to retrieves the data associated with the item.
```
smdata = storymapitem.get_data()
```
The call to the above method will return a python dictionary containing the contents of the story map which we wish to translate. We wish to translate not only the `text` content of the story map but also things like `title`, `summary`, `captions`, etc. To do so we will call the `replace` function defined above with the desired arguments.
```
result = replace(smdata, ['text', 'alt', 'title', 'summary', 'byline', 'caption', 'storyLogoAlt'], translate)
```
## Update cloned Story Map item
This cloned story map item doesn't contain the translated version of the story map until this point. But this can be achieved by calling the `update()` method of the cloned item and passing a dictionary of the item attributes we wish to translate.
```
cloned_item.update({'url': cloned_item.url.replace(storymapitem.id, cloned_item.id),
'text': result,
'title': translator.translate(storymapitem['title'])[0]['translated_text'],
'description': translator.translate(storymapitem['description'])[0]['translated_text'],
'snippet': translator.translate(storymapitem['snippet'])[0]['translated_text']})
```
The cloned story map text is now translated into Spanish language and is ready to be shared.
```
cloned_item
cloned_item.share(True)
```
# Conclusion
This sample demonstrates how inference only `TextTranslator` class of `arcgis.learn.text` submodule can be used to perform machine translation task to translate text from one language to another. We showed, how easy it is to translate a story map which is written in English language into Spanish language. Similar workflow can be followed to automate the task of translating story maps or other ArcGIS items into various languages.
# References
[1] [ISO Language Codes](https://en.wikipedia.org/wiki/List_of_ISO_639-1_codes)
[2] [Recursion](https://en.wikipedia.org/wiki/Recursion_(computer_science))
[3] [HTML](https://en.wikipedia.org/wiki/HTML)
| github_jupyter |
<a href="https://colab.research.google.com/github/maximematerno/DS-Unit-1-Sprint-3-Statistical-Tests-and-Experiments/blob/master/module3-introduction-to-bayesian-inference/LS_DS_133_Introduction_to_Bayesian_Inference_Assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Lambda School Data Science Module 133
## Introduction to Bayesian Inference
## Assignment - Code it up!
Most of the above was pure math - now write Python code to reproduce the results! This is purposefully open ended - you'll have to think about how you should represent probabilities and events. You can and should look things up, and as a stretch goal - refactor your code into helpful reusable functions!
Specific goals/targets:
1. Write a function `def prob_drunk_given_positive(prob_drunk_prior, prob_positive, prob_positive_drunk)` that reproduces the example from lecture, and use it to calculate and visualize a range of situations
2. Explore `scipy.stats.bayes_mvs` - read its documentation, and experiment with it on data you've tested in other ways earlier this week
3. Create a visualization comparing the results of a Bayesian approach to a traditional/frequentist approach
4. In your own words, summarize the difference between Bayesian and Frequentist statistics
If you're unsure where to start, check out [this blog post of Bayes theorem with Python](https://dataconomy.com/2015/02/introduction-to-bayes-theorem-with-python/) - you could and should create something similar!
Stretch goals:
- Apply a Bayesian technique to a problem you previously worked (in an assignment or project work) on from a frequentist (standard) perspective
- Check out [PyMC3](https://docs.pymc.io/) (note this goes beyond hypothesis tests into modeling) - read the guides and work through some examples
- Take PyMC3 further - see if you can build something with it!
##Bayes_theorem
```
def bayes_theorem(prior, false_positive, false_negative, num_iterations):
probabilities = []
for _ in range(num_iterations):
posterior_probability = (false_positive * prior) / ((false_positive * prior) + ((1-false_negative) * (1-prior)))
probabilities.append(posterior_probability)
prior = posterior_probability
return probabilities
bayes_theorem(0.005, .99, .99, 5)
```
##experiment with it on data you've tested in other ways earlier this week
```
import scipy
from scipy import stats
import pandas as pd
import numpy as np
from scipy.stats import ttest_1samp
from scipy.stats import ttest_ind, ttest_ind_from_stats, ttest_rel
from scipy import stats
df = pd.read_csv('https://archive.ics.uci.edu/ml/machine-learning-databases/voting-records/house-votes-84.data',
header=None,
names=['party','handicapped-infants','water-project',
'budget','physician-fee-freeze', 'el-salvador-aid',
'religious-groups','anti-satellite-ban',
'aid-to-contras','mx-missile','immigration',
'synfuels', 'education', 'right-to-sue','crime','duty-free',
'south-africa'])
print(df.shape)
df.head()
df= df.replace({'?':np.NaN, 'n':0, 'y':1})
df.head()
df.isnull().sum()
rep = df[df.party == 'republican']
print(rep.shape)
rep.head()
dem = df[df.party=='democrat']
print(dem.shape)
dem.head()
df.party.value_counts()
df=df.fillna(0)
print(df['water-project'].shape)
sample_Water_project= df['water-project'].sample(100)
print(sample_Water_project.shape)
sample_Water_project.head()
```
#confidence_interval
```
def confidence_interval(data, confidence_level=0.95):
data = np.array(data)
sample_mean = np.mean(data)
sample_size = len(data)
sample_std_dev = np.std(data, ddof=1)
standard_error = sample_std_dev / (sample_size**.5)
margin_of_error = standard_error * stats.t.ppf((1 + confidence_level) / 2.0, sample_size - 1)
return (sample_mean, sample_mean - margin_of_error, sample_mean + margin_of_error)
confidence_interval(sample_Water_project)
```
#Calculating Bayesian
```
bayesian_confidence_interval = stats.bayes_mvs(sample_Water_project, alpha=0.95)
print(bayesian_confidence_interval[0])
```
#visualization comparing the results
```
import seaborn as sns
import matplotlib.pyplot as plt
sns.kdeplot(sample_Water_project)
CI= confidence_interval(sample_Water_project)
plt.axvline(x=CI[1], color='red')
plt.axvline(x=CI[2], color='red')
plt.axvline(x=CI[0], color='k');
sns.distplot(sample_Water_project, kde=False, rug=True)
CI= confidence_interval(sample_Water_project)
plt.axvline(x=CI[1], color='red')
plt.axvline(x=CI[2], color='red')
plt.axvline(x=CI[0], color='k');
mean, variance, std = stats.bayes_mvs(sample_Water_project)
print(mean)
print(variance)
print(std)
res_mean, res_var, res_std = stats.bayes_mvs(sample_Water_project, alpha=0.95)
print(res_mean)
mean_confidence_interval, _, _ = stats.bayes_mvs(sample_Water_project, alpha=.95)
print(mean_confidence_interval)
print(mean_confidence_interval)
import matplotlib.pyplot as plt
fig = plt.figure()
ax = fig.add_subplot(111)
ax.hist(sample_Water_project, bins=100, density=True, label='Histogram of data')
ax.vlines(res_mean.statistic, 0, 0.5, colors='r', label='Estimated mean')
ax.axvspan(res_mean.minmax[0],res_mean.minmax[1], facecolor='r', alpha=0.2, label=r'Estimated mean (95% limits)')
ax.vlines(res_std.statistic, 0, 0.5, colors='g', label='Estimated scale')
ax.axvspan(res_std.minmax[0],res_std.minmax[1], facecolor='b', alpha=0.2, label=r'Estimated scale (95% limits)')
ax.legend(fontsize=10);
```
### We can see that Bayesian and Frequentist statistics have the same mean and intervall
- [Worked example of Bayes rule calculation](https://en.wikipedia.org/wiki/Bayes'_theorem#Examples) (helpful as it fully breaks out the denominator)
- [Source code for mvsdist in scipy](https://github.com/scipy/scipy/blob/90534919e139d2a81c24bf08341734ff41a3db12/scipy/stats/morestats.py#L139)
| github_jupyter |
```
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
from sklearn.metrics import confusion_matrix
from sklearn.neighbors import KNeighborsClassifier
from sklearn.model_selection import train_test_split
import seaborn as sns
sns.set()
df = pd.read_csv('Real_fake_news.csv')
df.head()
df.tail()
df.shape
df.info()
df.describe()
# Dropping the unnecessary columns
df.drop(['Unnamed: 0','Unnamed: 0.1'], axis=1, inplace=True)
df.head()
# Dropping the rows with null values
df.dropna()
df.shape
#FInding rows that have duplicate values
df[df.duplicated(keep = 'last')]
#Removing the duplicates
df = df.drop_duplicates(subset = None, keep ='first')
df.head()
df.BinaryNumTarget.value_counts()
# Defining numerical and categorical variables
num_atr=[]
cat_atr=['author' , 'statement' , 'source']
# Correlation Matrix
df.corr()
df.dtypes
df.columns
df.head()
# KNN Classification Algorithm
from sklearn.model_selection import train_test_split
X= df.drop(['BinaryNumTarget', 'BinaryTarget', 'target'], axis=1)
y= df['BinaryNumTarget']
X_train,X_test,y_train,y_test=train_test_split(X,y,test_size=0.3,random_state=1)
from sklearn.preprocessing import StandardScaler
from sklearn.metrics import mean_squared_error,r2_score
from sklearn.preprocessing import OneHotEncoder as onehot
from sklearn.preprocessing import LabelEncoder
le = LabelEncoder()
X_train[cat_atr] = X_train[cat_atr].apply(le.fit_transform)
X_train[cat_atr].head()
Xtr = X_train[cat_atr]
Xtr.shape
knn = KNeighborsClassifier(n_neighbors=5, metric='euclidean')
knn.fit(Xtr, y_train)
le = LabelEncoder()
X_test[cat_atr] = X_test[cat_atr].apply(le.fit_transform)
Xtr1 = X_test[cat_atr]
Xtr1.shape
y_pred = knn.predict(Xtr1)
confusion_matrix(y_test, y_pred)
from sklearn.metrics import accuracy_score
accuracy_score(y_test, y_pred)
from sklearn.metrics import precision_recall_fscore_support
precision_recall_fscore_support(y_test, y_pred)
from sklearn.metrics import precision_score
precision_score(y_test, y_pred)
from sklearn.metrics import recall_score
recall_score(y_test, y_pred)
from sklearn.metrics import f1_score
f1_score(y_test, y_pred)
error_rate = []
for i in range(1,40):
knn = KNeighborsClassifier(n_neighbors=i)
knn.fit(Xtr,y_train)
pred_i = knn.predict(Xtr1)
error_rate.append(1-accuracy_score(y_test, pred_i))
plt.figure(figsize=(10,6))
plt.plot(range(1,40),error_rate,color='blue', linestyle='dashed',
marker='o',markerfacecolor='red', markersize=10)
plt.title('Error Rate vs. K Value')
plt.xlabel('K')
plt.ylabel('Error Rate')
plt.show()
print("Minimum error:-",min(error_rate),"at K =",error_rate.index(min(error_rate))+1)
knn = KNeighborsClassifier(n_neighbors=36, metric='euclidean')
knn.fit(Xtr, y_train)
y_pred = knn.predict(Xtr1)
accuracy_score(y_test, y_pred)
```
| github_jupyter |
MNIST classification (drawn from sklearn example)
=====================================================
MWEM is not particularly well suited for image data (where there are tons of features with relatively large ranges) but it is still able to capture some important information about the underlying distributions if tuned correctly.
We use a feature included with MWEM that allows a column to be specified for a custom bin count, if we are capping every other bin count at a small value. In this case, we specify that the numerical column (784) has 10 possible values. We do this with the dict {'784': 10}.
Here we borrow from a scikit-learn example, and insert MWEM synthetic data into their training example/visualization, to understand the tradeoffs.
https://scikit-learn.org/stable/auto_examples/linear_model/plot_sparse_logistic_regression_mnist.html#sphx-glr-download-auto-examples-linear-model-plot-sparse-logistic-regression-mnist-py
```
import time
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from sklearn.datasets import fetch_openml
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.utils import check_random_state
# pip install scikit-image
from skimage import data, color
from skimage.transform import rescale
# Author: Arthur Mensch <arthur.mensch@m4x.org>
# License: BSD 3 clause
# Turn down for faster convergence
t0 = time.time()
train_samples = 5000
# Load data from https://www.openml.org/d/554
data = fetch_openml('mnist_784', version=1, return_X_y=False)
data_np = np.hstack((data.data,np.reshape(data.target.astype(int), (-1, 1))))
from opendp.smartnoise.synthesizers.mwem import MWEMSynthesizer
# Here we set max bin count to be 10, so that we retain the numeric labels
synth = MWEMSynthesizer(10.0, 40, 15, 10, split_factor=1, max_bin_count = 128, custom_bin_count={'784':10})
synth.fit(data_np)
sample_size = 2000
synthetic = synth.sample(sample_size)
from sklearn.linear_model import RidgeClassifier
import utils
real = pd.DataFrame(data_np[:sample_size])
model_real, model_fake = utils.test_real_vs_synthetic_data(real, synthetic, RidgeClassifier, tsne=True)
# Classification
coef = model_real.coef_.copy()
plt.figure(figsize=(10, 5))
scale = np.abs(coef).max()
for i in range(10):
l1_plot = plt.subplot(2, 5, i + 1)
l1_plot.imshow(coef[i].reshape(28, 28), interpolation='nearest',
cmap=plt.cm.RdBu, vmin=-scale, vmax=scale)
l1_plot.set_xticks(())
l1_plot.set_yticks(())
l1_plot.set_xlabel('Class %i' % i)
plt.suptitle('Classification vector for...')
run_time = time.time() - t0
print('Example run in %.3f s' % run_time)
plt.show()
coef = model_fake.coef_.copy()
plt.figure(figsize=(10, 5))
scale = np.abs(coef).max()
for i in range(10):
l1_plot = plt.subplot(2, 5, i + 1)
l1_plot.imshow(coef[i].reshape(28, 28), interpolation='nearest',
cmap=plt.cm.RdBu, vmin=-scale, vmax=scale)
l1_plot.set_xticks(())
l1_plot.set_yticks(())
l1_plot.set_xlabel('Class %i' % i)
plt.suptitle('Classification vector for...')
run_time = time.time() - t0
print('Example run in %.3f s' % run_time)
plt.show()
```
| github_jupyter |
# Dataset Descriptions
This notebook contains most of the datasets used in Pandas Cookbook along with the names, types, descriptions and some summary statistics of each column. This is not an exhaustive list as several datasets used in the book are quite small and are explained with enough detail in the book itself. The datasets presented here are the prominent ones that appear most frequently throughout the book.
## Datasets in order of appearance
* [Movie](#Movie-Dataset)
* [College](#College-Dataset)
* [Employee](#Employee-Dataset)
* [Flights](#Flights-Dataset)
* [Chinook Database](#Chinook-Database)
* [Crime](#Crime-Dataset)
* [Meetup Groups](#Meetup-Groups-Dataset)
* [Diamonds](#Diamonds-Dataset)
```
import pandas as pd
pd.options.display.max_columns = 80
```
# Movie Dataset
### Brief Overview
28 columns from 4,916 movies scraped from the popular website IMDB. Each row contains information on a single movie dating back to 1916 to 2015. Actor and director facebook likes should be constant for all instances across all movies. For instance, Johnny Depp should have the same number of facebook likes regardless of which movie he is in. Since each movie was not scraped at the same exact time, there are some inconsistencies in these counts. The dataset **movie_altered.csv** is a much cleaner version of this dataset.
```
movie = pd.read_csv('data/movie.csv')
movie.head()
movie.shape
pd.read_csv('data/descriptions/movie_decsription.csv', index_col='Column Name')
```
# College Dataset
### Brief Overview
US department of education data on 7,535 colleges. Only a sample of the total number of columns available were used in this dataset. Visit [the website](https://collegescorecard.ed.gov/data/) for more info. Data was pulled in January, 2017.
```
college = pd.read_csv('data/college.csv')
college.head()
college.shape
pd.read_csv('data/descriptions/college_decsription.csv')
```
# Employee Dataset
### Brief Overview
The city of Houston provides information on all its employees to the public. This is a random sample of 2,000 employees with a few of the more interesting columns. For more on [open Houston data visit their website](http://data.houstontx.gov/). Data was pulled in December, 2016.
```
employee = pd.read_csv('data/employee.csv')
employee.head()
employee.shape
pd.read_csv('data/descriptions/employee_description.csv')
```
# Flights Dataset
### Brief Overview
A random sample of three percent of the US domestic flights originating from the ten busiest airports. Data is from the U.S. Department of Transportation's (DOT) Bureau of Transportation Statistics. [See here for more info](https://www.kaggle.com/usdot/flight-delays).
```
flights = pd.read_csv('data/flights.csv')
flights.head()
flights.shape
pd.read_csv('data/descriptions/flights_description.csv')
```
### Airline Codes
```
pd.read_csv('data/descriptions/airlines.csv')
```
### Airport codes
```
pd.read_csv('data/descriptions/airports.csv').head()
```
# Chinook Database
### Brief Overview
This is a sample database of a music store provided by SQLite with 11 tables. The table description image is an excellent way to get familiar with the database. [Visit the sqlite website](http://www.sqlitetutorial.net/sqlite-sample-database/) for more detail.

# Crime Dataset
### Brief Overview
All crime and traffic accidents for the city of Denver from January to September of 2017. This dataset is stored in special binary form called *hdf5*. Pandas uses the PyTables library to help read the data into a DataFrame. [Read the documentation](http://pandas.pydata.org/pandas-docs/stable/io.html#io-hdf5) for more info on hdf5 formatted data.
```
crime = pd.read_hdf('data/crime.h5')
crime.head()
crime.shape
pd.read_csv('data/descriptions/crime_description.csv')
```
# Meetup Groups Dataset
### Brief Overview
Data was collected through the [meetup.com API](https://www.meetup.com/meetup_api/) on five Houston-area data science meetup groups. Each row represents a member joining a particular group.
```
meetup = pd.read_csv('data/meetup_groups.csv')
meetup.head()
meetup.shape
pd.read_csv('data/descriptions/meetup_description.csv')
```
# Diamonds Dataset
### Brief Overview
Quality, size and price of nearly 54,000 diamonds scraped from the [Diamond Search Engine](http://www.diamondse.info/) by Hadley Wickham. [Visit blue nile](https://www.bluenile.com/ca/education/diamonds?track=SideNav) for a beginners guide to diamonds.
```
diamonds = pd.read_csv('data/diamonds.csv')
diamonds.head()
diamonds.shape
pd.read_csv('data/descriptions/diamonds_description.csv')
```
| github_jupyter |
___
<a href='https://www.udemy.com/user/joseportilla/'><img src='../Pierian_Data_Logo.png'/></a>
___
<center><em>Content Copyright by Pierian Data</em></center>
# Working with CSV Files
Welcome back! Let's discuss how to work with CSV files in Python. A file with the CSV file extension is a Comma Separated Values file. All CSV files are plain text, contain alphanumeric characters, and structure the data contained within them in a tabular form. Don't confuse Excel Files with csv files, while csv files are formatted very similarly to excel files, they don't have data types for their values, they are all strings with no font or color. They also don't have worksheets the way an excel file does. Python does have several libraries for working with Excel files, you can check them out [here](http://www.python-excel.org/) and [here](https://www.xlwings.org/).
Files in the CSV format are generally used to exchange data, usually when there's a large amount, between different applications. Database programs, analytical software, and other applications that store massive amounts of information (like contacts and customer data), will usually support the CSV format.
Let's explore how we can open a csv file with Python's built-in csv library.
____
## Notebook Location.
Run **pwd** inside a notebook cell to find out where your notebook is located
```
pwd
```
____
## Reading CSV Files
```
import csv
```
When passing in the file path, make sure to include the extension if it has one, you should be able to Tab Autocomplete the file name. If you can't Tab autocomplete, that is a good indicator your file is not in the same location as your notebook. You can always type in the entire file path (it will look similar in formatting to the output of **pwd**.
```
data = open('example.csv')
data
```
### Encoding
Often csv files may contain characters that you can't interpret with standard python, this could be something like an **@** symbol, or even foreign characters. Let's view an example of this sort of error ([its pretty common, so its important to go over](https://stackoverflow.com/questions/9233027/unicodedecodeerror-charmap-codec-cant-decode-byte-x-in-position-y-character)).
```
csv_data = csv.reader(data)
```
Cast to a list will give an error, note the **can't decode** line in the error, this is a giveaway that we have an encoding problem!
```
data_lines = list(csv_data)
```
Let's not try reading it with a "utf-8" encoding.
```
data = open('example.csv',encoding="utf-8")
csv_data = csv.reader(data)
data_lines = list(csv_data)
# Looks like it worked!
data_lines[:3]
```
Note the first item in the list is the header line, this contains the information about what each column represents. Let's format our printing just a bit:
```
for line in data_lines[:5]:
print(line)
```
Let's imagine we wanted a list of all the emails. For demonstration, since there are 1000 items plus the header, we will only do a few rows.
```
len(data_lines)
all_emails = []
for line in data_lines[1:15]:
all_emails.append(line[3])
print(all_emails)
```
What if we wanted a list of full names?
```
full_names = []
for line in data_lines[1:15]:
full_names.append(line[1]+' '+line[2])
full_names
```
## Writing to CSV Files
We can also write csv files, either new ones or add on to existing ones.
### New File
**This will also overwrite any exisiting file with the same name, so be careful with this!**
```
# newline controls how universal newlines works (it only applies to text
# mode). It can be None, '', '\n', '\r', and '\r\n'.
file_to_output = open('to_save_file.csv','w',newline='')
csv_writer = csv.writer(file_to_output,delimiter=',')
csv_writer.writerow(['a','b','c'])
csv_writer.writerows([['1','2','3'],['4','5','6']])
file_to_output.close()
```
____
### Existing File
```
f = open('to_save_file.csv','a',newline='')
csv_writer = csv.writer(f)
csv_writer.writerow(['new','new','new'])
f.close()
```
That is all for the basics! If you believe you will be working with CSV files often, you may want to check out the powerful [pandas library](https://pandas.pydata.org/).
| github_jupyter |
```
import gym
from gym import spaces
from gym.utils import seeding
import pandas as pd
import numpy as np
import typing
from datetime import datetime
import ray
# Start up Ray. This must be done before we instantiate any RL agents.
ray.init(num_cpus=10, ignore_reinit_error=True, log_to_driver=False,_temp_dir="/rds/general/user/asm119/ephemeral")
tom_path = "../Thomas/"
def load_data(
price_source: str,
tickers: typing.List[str],
start: datetime,
end: datetime,
features: typing.List[str],
):
"""Returned price data to use in gym environment"""
# Load data
# Each dataframe will have columns date and a collection of fields
# TODO: DataLoader from mongoDB
# Raw price from DB, forward impute on the trading days for missing date
# calculate the features (log return, volatility)
if price_source in ["csvdata"]:
feature_df = []
for t in tickers:
df1 = pd.read_csv(tom_path + "csvdata/{}.csv".format(t))
df1['datetime'] = pd.to_datetime(df1['datetime'])
df1 = df1[(df1['datetime']>=start) & (df1['datetime']<=end)]
df1.set_index("datetime",inplace=True)
selected_features = ['return','tcost'] + features
feature_df.append(df1[selected_features])
ref_df_columns = df1[selected_features].columns
# assume all the price_df are aligned and cleaned in the DataLoader
merged_df = pd.concat(feature_df, axis=1, join="outer")
# Imputer missing values with zeros
price_tensor = merged_df['return'].fillna(0.0).values
tcost = merged_df['tcost'].fillna(0.0).values
return {
"dates": merged_df.index,
"fields": ref_df_columns,
"data": merged_df.fillna(0.0).values,
"pricedata": price_tensor,
"tcost": tcost,
}
load_data('csvdata',['SPY','QQQ',], datetime(2010, 5, 4), datetime(2020, 12, 31), ["volatility_20", "skewness_20", "kurtosis_20"] ) ['data'][:10,:]
from empyrical import max_drawdown, alpha_beta, sharpe_ratio, annual_return
from sklearn.preprocessing import StandardScaler
class Equitydaily(gym.Env):
def __init__(self,env_config):
self.tickers = env_config['tickers']
self.lookback = env_config['lookback']
# Load price data, to be replaced by DataLoader class
raw_data = load_data(env_config['pricing_source'],env_config['tickers'],env_config['start'],env_config['end'],env_config['features'])
# Set the trading dates, features and price data
self.dates = raw_data['dates']
self.fields = raw_data['fields']
self.pricedata = raw_data['pricedata']
self.featuredata = raw_data['data']
self.tcostdata = raw_data['tcost']
# Set up historical actions and rewards
self.n_assets = len(self.tickers) + 1
self.n_metrics = 2
self.n_assets_fields = len(self.fields)
self.n_features = self.n_assets_fields * len(self.tickers) + self.n_assets + self.n_metrics # reward function
#self.n_features = self.n_assets_fields * len(self.tickers)
# Set up action and observation space
# The last asset is cash
self.action_space = spaces.Box(low=-1, high=1, shape=(len(self.tickers)+1,), dtype=np.float32)
self.observation_space = spaces.Box(low=-np.inf, high=np.inf,
shape=(self.lookback,self.n_features,1), dtype=np.float32)
self.reset()
def step(self, action):
# Trade every 10 days
# Normalise action space
if self.index % 10 == 0:
normalised_action = action / np.sum(np.abs(action))
self.actions = normalised_action
done = False
# Rebalance portfolio at close using return of the next date
next_day_log_return = self.pricedata[self.index,:]
# transaction cost
transaction_cost = self.transaction_cost(self.actions,self.position_series[-1])
# Rebalancing
self.position_series = np.append(self.position_series, [self.actions], axis=0)
# Portfolio return
today_portfolio_return = np.sum(self.actions[:-1] * next_day_log_return) + np.sum(transaction_cost)
self.log_return_series = np.append(self.log_return_series, [today_portfolio_return], axis=0)
# Calculate reward
# Need to cast log_return in pd series to use the functions in empyrical
recent_series = pd.Series(self.log_return_series)[-100:]
rolling_volatility = np.std(recent_series)
self.metric = today_portfolio_return / rolling_volatility
reward = self.metric
self.metric_series = np.append(self.metric_series, [self.metric], axis=0)
# Check if the end of backtest
if self.index >= self.pricedata.shape[0]-2:
done = True
# Prepare observation for next day
self.index += 1
self.observation = self.get_observation()
return self.observation, reward, done, {}
def reset(self):
self.log_return_series = np.zeros(shape=self.lookback)
self.metric_series = np.zeros(shape=self.lookback)
self.position_series = np.zeros(shape=(self.lookback,self.n_assets))
self.metric = 0
self.index = self.lookback
self.actions = np.zeros(shape=self.n_assets)
self.observation = self.get_observation()
return self.observation
def get_observation(self):
# Can use simple moving average data here
price_lookback = self.featuredata[self.index-self.lookback:self.index,:]
metrics = np.vstack((self.log_return_series[self.index-self.lookback:self.index],
self.metric_series[self.index-self.lookback:self.index])).transpose()
positions = self.position_series[self.index-self.lookback:self.index]
scaler = StandardScaler()
price_lookback = scaler.fit_transform(price_lookback)
observation = np.concatenate((price_lookback, metrics, positions), axis=1)
return observation.reshape((observation.shape[0], observation.shape[1], 1))
# 0.05% and spread to model t-cost for institutional portfolios
def transaction_cost(self, new_action, old_action,):
turnover = np.abs(new_action - old_action)
fees = 0.9995 - self.tcostdata[self.index,:]
fees = np.array(list(fees) + [0.9995])
tcost = turnover * np.log(fees)
return tcost
config = {'pricing_source':'csvdata', 'tickers':['BRK','TLT','QQQ','GLD',],
'lookback':10, 'start':'2008-01-02', 'end':'2018-12-31', 'features':["volatility_20", "skewness_20", "kurtosis_20"]}
EQ_env = Equitydaily(config)
EQ_env.observation.shape, EQ_env.n_features
EQ_env.observation
state = EQ_env.reset()
done = False
reward_list = []
cum_reward = 0
actions = list()
while not done:
#action = agent.compute_action(state)
action = np.array([1,0,0,0,0])
state, reward, done, future_price = EQ_env.step(action)
cum_reward += reward
actions.append(action)
reward_list.append(reward)
pd.Series(reward_list).cumsum().plot()
```
PPO policy
```
from ray.rllib.agents.ppo import PPOTrainer, DEFAULT_CONFIG
from ray.tune.logger import pretty_print
config = DEFAULT_CONFIG.copy()
config['num_workers'] = 1
config["num_envs_per_worker"] = 1
config["rollout_fragment_length"] = 20
config["train_batch_size"] = 5000
config["batch_mode"] = "complete_episodes"
config['num_sgd_iter'] = 20
config['sgd_minibatch_size'] = 200
config['model']['dim'] = 200
config['model']['conv_filters'] = [[32, [5, 1], 5], [32, [5, 1], 5], [4, [5, 1], 5]]
config['num_cpus_per_worker'] = 2 # This avoids running out of resources in the notebook environment when this cell is re-executed
config['env_config'] = {'pricing_source':'csvdata', "tickers": ["BRK_A", "GE_","GOLD_", "AAPL_","GS_","T_",],
'lookback':200, 'start':'1995-01-02', 'end':'2018-12-31', 'features':["return_volatility_20", "return_skewness_20", "return_kurtosis_20"]}
config
```
Check to see if agents can be trained
```
agent = PPOTrainer(config, Equitydaily)
best_reward = -np.inf
for i in range(2):
result = agent.train()
if result['episode_reward_mean'] > best_reward + 10:
path = agent.save('sampleagent')
print(path)
best_reward = result['episode_reward_mean']
print(best_reward)
result
agent.restore('sampleagent/checkpoint_1/checkpoint-1')
for i in range(5):
result = agent.train()
if result['episode_reward_mean'] > best_reward + 1:
path = agent.save('sampleagent')
print(path)
best_reward = result['episode_reward_mean']
print(best_reward)
result
```
SAC
```
from ray.rllib.agents.sac import SACTrainer, DEFAULT_CONFIG
from ray.tune.logger import pretty_print
config = DEFAULT_CONFIG.copy()
config['num_workers'] = 1
config["num_envs_per_worker"] = 1
config["rollout_fragment_length"] = 10
config["train_batch_size"] = 50
config["timesteps_per_iteration"] = 10
config["buffer_size"] = 10000
config["Q_model"]["fcnet_hiddens"] = [10, 10]
config["policy_model"]["fcnet_hiddens"] = [10, 10]
config["num_cpus_per_worker"] = 2
config["env_config"] = {
"pricing_source": "csvdata",
"tickers": ["QQQ", "EEM", "TLT", "SHY", "GLD", "SLV"],
"lookback": 1,
"start": "2007-01-02",
"end": "2015-12-31",
}
# Train agent
agent = SACTrainer(config, Equitydaily)
best_reward = -np.inf
for i in range(20):
result = agent.train()
if result['episode_reward_mean'] > best_reward + 0.01:
path = agent.save('sampleagent')
print(path)
best_reward = result['episode_reward_mean']
print(result['episode_reward_mean'])
result
```
Run environment
```
config
agent = PPOTrainer(config, Equitydaily)
env = Equitydaily({'pricing_source':'Alpaca_Equity_daily', 'tickers':['SPY','QQQ','SHY','GLD','TLT','EEM'], 'lookback':50, 'start':'2011-01-02', 'end':'2020-12-31'})
agent.restore('checkpoint_1087/checkpoint-1087')
state = env.reset()
done = False
reward_list = []
cum_reward = 0
actions = list()
while not done:
#action = agent.compute_action(state)
action = np.array([0,0,0,0,0,0,1])
state, reward, done, _ = env.step(action)
cum_reward += reward
actions.append(action)
reward_list.append(reward)
pd.Series(env.log_return_series).cumsum().plot()
pd.Series(reward_list).plot()
pd.DataFrame(actions)
```
Run environment for RNN environment
```
env = Equitydaily({'pricing_source':'Alpaca_Equity_daily', 'tickers':['SPY','QQQ'], 'lookback':50, 'start':'2018-01-02', 'end':'2020-12-31'})
state = env.reset()
done = False
cum_reward = 0
actions = list()
rnn_state = agent.get_policy().get_initial_state()
while not done:
action, rnn_state, _ = agent.compute_action(state,rnn_state)
#action = np.array([1,-1])
state, reward, done, _ = env.step(action)
cum_reward += reward
actions.append(actions)
pd.Series(env.log_return_series).cumsum().plot()
max_drawdown(pd.Series(env.log_return_series))
annual_return(pd.Series(env.log_return_series))
class Equitydaily_v1(gym.Env):
def __init__(self,env_config):
self.tickers = env_config['tickers']
self.lookback = env_config['lookback']
# Load price data, to be replaced by DataLoader class
raw_data = load_data(env_config['pricing_source'],env_config['tickers'],env_config['start'],env_config['end'])
# Set the trading dates, features and price data
self.dates = raw_data['dates']
self.fields = raw_data['fields']
self.pricedata = raw_data['pricedata']
self.featuredata = raw_data['data']
self.tcostdata = raw_data['tcost']
# Set up historical actions and rewards
self.n_assets = len(self.tickers) + 1
self.n_metrics = 2
self.n_assets_fields = len(self.fields)
self.n_features = self.n_assets_fields * len(self.tickers) + self.n_assets + self.n_metrics # reward function
# Set up action and observation space
# The last asset is cash
self.action_space = spaces.Box(low=-1, high=1, shape=(len(self.tickers)+1,), dtype=np.float32)
self.observation_space = spaces.Box(low=-np.inf, high=np.inf,
shape=(self.lookback,self.n_features), dtype=np.float32)
self.reset()
def step(self, action):
## Normalise action space
normalised_action = action / np.sum(np.abs(action))
done = False
# Rebalance portfolio at close using return of the next date
next_day_log_return = self.pricedata[self.index,:]
# transaction cost
transaction_cost = self.transaction_cost(normalised_action,self.position_series[-1])
# Rebalancing
self.position_series = np.append(self.position_series, [normalised_action], axis=0)
# Portfolio return
today_portfolio_return = np.sum(normalised_action[:-1] * next_day_log_return) + np.sum(transaction_cost)
self.log_return_series = np.append(self.log_return_series, [today_portfolio_return], axis=0)
# Calculate reward
# Need to cast log_return in pd series to use the functions in empyrical
live_days = self.index - self.lookback
burnin = 250
recent_series = pd.Series(self.log_return_series)[-100:]
whole_series = pd.Series(self.log_return_series)
if live_days > burnin:
self.metric = annual_return(whole_series) + 0.5* max_drawdown(whole_series)
else:
self.metric = annual_return(whole_series) + 0.5* max_drawdown(whole_series) *live_days / burnin
reward = self.metric - self.metric_series[-1]
#reward = self.metric
self.metric_series = np.append(self.metric_series, [self.metric], axis=0)
# Check if the end of backtest
if self.index >= self.pricedata.shape[0]-2:
done = True
# Prepare observation for next day
self.index += 1
self.observation = self.get_observation()
return self.observation, reward, done, {'current_price':next_day_log_return}
def reset(self):
self.log_return_series = np.zeros(shape=self.lookback)
self.metric_series = np.zeros(shape=self.lookback)
self.position_series = np.zeros(shape=(self.lookback,self.n_assets))
self.metric = 0
self.index = self.lookback
self.observation = self.get_observation()
return self.observation
def get_observation(self):
# Can use simple moving average data here
price_lookback = self.featuredata[self.index-self.lookback:self.index,:]
metrics = np.vstack((self.log_return_series[self.index-self.lookback:self.index],
self.metric_series[self.index-self.lookback:self.index])).transpose()
positions = self.position_series[self.index-self.lookback:self.index]
observation = np.concatenate((price_lookback, metrics, positions), axis=1)
return observation
# 0.05% and spread to model t-cost for institutional portfolios
def transaction_cost(self,new_action,old_action,):
turnover = np.abs(new_action - old_action)
fees = 0.9995 - self.tcostdata[self.index,:]
tcost = turnover * np.log(fees)
return tcost
```
| github_jupyter |
# T-ABSA Logistic Regression Model using word2vec
#### Preprocessing the reviews
Importing the libraries for preprocessing the reviews
```
import os
import pandas as pd
import nltk
from gensim.models import Word2Vec, word2vec
import matplotlib.pyplot as plt
import numpy as np
from nltk.corpus import stopwords
import os
import re
```
Loading the training dataset into python
```
data_dir = 'D:/jupyter/Year2_Research/Generate_Data/data/5_aspects/'
df_train = pd.read_csv(os.path.join(data_dir, "train_NLI.tsv"),sep="\t")
df_dev = pd.read_csv(os.path.join(data_dir, "train_NLI.tsv"),sep="\t")
df_test = pd.read_csv(os.path.join(data_dir, "test_NLI.tsv"),sep="\t")
df_train.tail(2)
frames = [df_train, df_dev, df_test]
combined_dataframe = pd.concat(frames)
combined_dataframe.iloc[3000]
combined_dataframe.tail()
combined_dataframe['concatinated'] = combined_dataframe['sentence1'] + ' ' + combined_dataframe['sentence2']
combined_dataframe['concatinated'][4102]
word2vec_training_dataset = combined_dataframe['concatinated'].values
word2vec_training_dataset[4000]
```
### Preprocessing the data
Convert each review in the training set to a list of sentences where each sentence is in turn a list of words.
Besides splitting reviews into sentences, non-letters and stop words are removed and all words
coverted to lower case.
```
def review_to_wordlist(review, remove_stopwords=True):
"""
Convert a review to a list of words.
"""
# remove non-letters
review_text = re.sub("[^a-zA-Z]"," ", review)
# convert to lower case and split at whitespace
words = review_text.lower().split()
# remove stop words (false by default)
if remove_stopwords:
stops = set(stopwords.words("english"))
words = [w for w in words if not w in stops]
return words
# Load the punkt tokenizer used for splitting reviews into sentences
tokenizer = nltk.data.load('tokenizers/punkt/english.pickle')
def review_to_sentences(review, tokenizer, remove_stopwords=True):
"""
Split review into list of sentences where each sentence is a list of words.
"""
# use the NLTK tokenizer to split the paragraph into sentences
raw_sentences = tokenizer.tokenize(review.strip())
# each sentence is furthermore split into words
sentences = []
for raw_sentence in raw_sentences:
# If a sentence is empty, skip it
if len(raw_sentence) > 0:
sentences.append(review_to_wordlist(raw_sentence, remove_stopwords))
return sentences
train_sentences = [] # Initialize an empty list of sentences
for review in word2vec_training_dataset:
train_sentences += review_to_sentences(review, tokenizer)
train_sentences[4000]
```
### Training a word2vec model
```
model_name = 'train_model'
# Set values for various word2vec parameters
num_features = 300 # Word vector dimensionality
min_word_count = 40 # Minimum word count
num_workers = 3 # Number of threads to run in parallel
context = 10 # Context window size
downsampling = 1e-3 # Downsample setting for frequent words
if not os.path.exists(model_name):
# Initialize and train the model (this will take some time)
model = word2vec.Word2Vec(train_sentences, workers=num_workers, \
size=num_features, min_count = min_word_count, \
window = context, sample = downsampling)
# If you don't plan to train the model any further, calling
# init_sims will make the model much more memory-efficient.
model.init_sims(replace=True)
# It can be helpful to create a meaningful model name and
# save the model for later use. You can load it later using Word2Vec.load()
model.save(model_name)
else:
model = Word2Vec.load(model_name)
```
```
model.most_similar("internet")
```
### Building a Classifier
```
# shape of the data
df_train.shape
```
Encoding the labels of the dataset
```
y_train = df_train['label'].replace(['None','Positive','Negative'],[1,2,0])
x_cols = [x for x in df_train.columns if x != 'label']
# Split the data into two dataframes (one for the labels and the other for the independent variables)
X_data = df_train[x_cols]
X_data.tail()
X_data['concatinated'] = X_data['sentence1'] + ' ' + X_data['sentence2']
X_data['concatinated'][9]
X_train = X_data['concatinated'].values
X_train[100]
y_train[100]
```
## 3. Build classifier using word embedding
Each review is mapped to a feature vector by averaging the word embeddings of all words in the review. These features are then fed into a random forest classifier.
```
def make_feature_vec(words, model, num_features):
"""
Average the word vectors for a set of words
"""
feature_vec = np.zeros((num_features,),dtype="float32") # pre-initialize (for speed)
nwords = 0
#index2word_set = set(model.index2word) # words known to the model
index2word_set = set(model.wv.index2word) # words known to the model
for word in words:
if word in index2word_set:
nwords = nwords + 1
feature_vec = np.add(feature_vec,model[word])
feature_vec = np.divide(feature_vec, nwords)
return feature_vec
def get_avg_feature_vecs(reviews, model, num_features):
"""
Calculate average feature vectors for all reviews
"""
counter = 0
review_feature_vecs = np.zeros((len(reviews),num_features), dtype='float32') # pre-initialize (for speed)
for review in reviews:
review_feature_vecs[counter] = make_feature_vec(review, model, num_features)
counter = counter + 1
return review_feature_vecs
# calculate average feature vectors for training and test sets
clean_train_reviews = []
for review in X_train:
clean_train_reviews.append(review_to_wordlist(review, remove_stopwords=True))
trainDataVecs = get_avg_feature_vecs(clean_train_reviews, model, num_features)
```
#### Fit a random forest classifier to the training data
```
from sklearn.linear_model import LogisticRegression
print("Fitting a weighted logistic regression to the labeled training data...")
model_lr = LogisticRegression(class_weight='balanced')
model_lr = model_lr.fit(trainDataVecs, y_train)
print("Fitting Completed")
```
## 4. Prediction
### Test set data preparation
```
# Split the data into two dataframes (one for the labels and the other for the independent variables)
x_cols = [x for x in df_test.columns if x != 'label']
X_data_test = df_test[x_cols]
# Combining the review with the generated auxilliary sentence
X_data_test['concatinated'] = X_data_test['sentence1'] + ' ' + X_data_test['sentence2']
# X test data
X_test = X_data_test['concatinated'].values
print(X_test[100:108])
# y test data
y_test = df_test['label'].replace(['None','Positive','Negative'],[1,2,0])
y_test[100:108]
clean_test_reviews = []
for review in X_test:
clean_test_reviews.append(review_to_wordlist(review, remove_stopwords=True))
testDataVecs = get_avg_feature_vecs(clean_test_reviews, model, num_features)
# remove instances in test set that could not be represented as feature vectors
nan_indices = list({x for x,y in np.argwhere(np.isnan(testDataVecs))})
if len(nan_indices) > 0:
print('Removing {:d} instances from test set.'.format(len(nan_indices)))
testDataVecs = np.delete(testDataVecs, nan_indices, axis=0)
test_reviews.drop(test_reviews.iloc[nan_indices, :].index, axis=0, inplace=True)
assert testDataVecs.shape[0] == len(test_reviews)
print("Predicting labels for test data..")
Y_predicted = model_lr.predict(testDataVecs)
```
Evaluating the performance of the model
```
from sklearn.metrics import accuracy_score
print(accuracy_score(y_test, Y_predicted))
Y_forest_score = model_lr.predict_proba(testDataVecs)
Y_forest_score
import csv
# Open/Create a file to append data
csvFile_pred = open('prediction_score.csv', 'w')
#Use csv Writer
csvWriter_pred = csv.writer(csvFile_pred)
csvWriter_pred.writerow(['predicted','score_none','score_pos','score_neg'])
for f in range(len(Y_predicted)):
csvWriter_pred.writerow([Y_predicted[f],Y_forest_score[f][1], Y_forest_score[f][0], Y_forest_score[f][2]])
csvFile_pred.close()
dataframe = pd.read_csv('prediction_score.csv')
dataframe.tail()
```
### Evaluating the model
```
# -*- coding: utf-8 -*-
"""
Created on Sun Oct 20 11:40:28 2019
@author: David
"""
import collections
import numpy as np
import pandas as pd
from sklearn import metrics
def get_y_true():
# """
# Read file to obtain y_true.
#
# """
true_data_file = "D:/jupyter/Year2_Research/Generate_Data/data/5_aspects/test_NLI.tsv"
df = pd.read_csv(true_data_file,sep='\t')
y_true = []
for i in range(len(df)):
label = df['label'][i]
assert label in ['None', 'Positive', 'Negative'], "error!"
if label == 'None':
n = 1
elif label == 'Positive':
n = 2
else:
n = 0
y_true.append(n)
print(len(y_true))
return y_true
def get_y_pred():
# """
# Read file to obtain y_pred and scores.
# """
dataframe = pd.read_csv('prediction_score.csv')
pred=[]
score=[]
for f in range(len(dataframe)):
pred.append(dataframe.predicted[f])
score.append([float(dataframe.score_pos[f]),float(dataframe.score_none[f]),float(dataframe.score_neg[f])])
return pred, score
def sentitel_strict_acc(y_true, y_pred):
"""
Calculate "strict Acc" of aspect detection task of sentitel.
"""
total_cases=int(len(y_true)/5)
true_cases=0
for i in range(total_cases):
if y_true[i*5]!=y_pred[i*5]:continue
if y_true[i*5+1]!=y_pred[i*5+1]:continue
if y_true[i*5+2]!=y_pred[i*5+2]:continue
if y_true[i*5+3]!=y_pred[i*5+3]:continue
if y_true[i*5+4]!=y_pred[i*5+4]:continue
true_cases+=1
aspect_strict_Acc = true_cases/total_cases
return aspect_strict_Acc
def sentitel_macro_F1(y_true, y_pred):
"""
Calculate "Macro-F1" of aspect detection task of sentitel.
"""
p_all=0
r_all=0
count=0
for i in range(len(y_pred)//5):
a=set()
b=set()
for j in range(5):
if y_pred[i*5+j]!=1:
a.add(j)
if y_true[i*5+j]!=1:
b.add(j)
if len(b)==0:continue
a_b=a.intersection(b)
if len(a_b)>0:
p=len(a_b)/len(a)
r=len(a_b)/len(b)
else:
p=0
r=0
count+=1
p_all+=p
r_all+=r
Ma_p=p_all/count
Ma_r=r_all/count
aspect_Macro_F1 = 2*Ma_p*Ma_r/(Ma_p+Ma_r)
return aspect_Macro_F1
def sentitel_AUC_Acc(y_true, score):
"""
Calculate "Macro-AUC" of both aspect detection and sentiment classification tasks of sentitel.
Calculate "Acc" of sentiment classification task of sentitel.
"""
# aspect-Macro-AUC
aspect_y_true=[]
aspect_y_score=[]
aspect_y_trues=[[],[],[],[],[]]
aspect_y_scores=[[],[],[],[],[]]
for i in range(len(y_true)):
if y_true[i]>0:
aspect_y_true.append(0)
else:
aspect_y_true.append(1) # "None": 1
tmp_score=score[i][0] # probability of "None"
aspect_y_score.append(tmp_score)
aspect_y_trues[i%5].append(aspect_y_true[-1])
aspect_y_scores[i%5].append(aspect_y_score[-1])
aspect_auc=[]
for i in range(5):
aspect_auc.append(metrics.roc_auc_score(aspect_y_trues[i], aspect_y_scores[i]))
aspect_Macro_AUC = np.mean(aspect_auc)
# sentiment-Macro-AUC
sentiment_y_true=[]
sentiment_y_pred=[]
sentiment_y_score=[]
sentiment_y_trues=[[],[],[],[],[]]
sentiment_y_scores=[[],[],[],[],[]]
for i in range(len(y_true)):
if y_true[i]>0:
sentiment_y_true.append(y_true[i]-1) # "Postive":0, "Negative":1
tmp_score=score[i][2]/(score[i][1]+score[i][2]) # probability of "Negative"
sentiment_y_score.append(tmp_score)
if tmp_score>0.5:
sentiment_y_pred.append(1) # "Negative": 1
else:
sentiment_y_pred.append(0)
sentiment_y_trues[i%5].append(sentiment_y_true[-1])
sentiment_y_scores[i%5].append(sentiment_y_score[-1])
sentiment_auc=[]
for i in range(5):
sentiment_auc.append(metrics.roc_auc_score(sentiment_y_trues[i], sentiment_y_scores[i]))
sentiment_Macro_AUC = np.mean(sentiment_auc)
# sentiment Acc
sentiment_y_true = np.array(sentiment_y_true)
sentiment_y_pred = np.array(sentiment_y_pred)
sentiment_Acc = metrics.accuracy_score(sentiment_y_true,sentiment_y_pred)
return aspect_Macro_AUC, sentiment_Acc, sentiment_Macro_AUC
#####################################################################
y_true = (get_y_true())
y_pred, score = get_y_pred()
result = collections.OrderedDict()
aspect_strict_Acc = sentitel_strict_acc(y_true, y_pred)
aspect_Macro_F1 = sentitel_macro_F1(y_true, y_pred)
aspect_Macro_AUC, sentiment_Acc, sentiment_Macro_AUC = sentitel_AUC_Acc(y_true, score)
result = {'aspect_strict_Acc': aspect_strict_Acc,
'aspect_Macro_F1': aspect_Macro_F1,
'aspect_Macro_AUC': aspect_Macro_AUC,
'sentiment_Acc': sentiment_Acc,
'sentiment_Macro_AUC': sentiment_Macro_AUC}
print(result)
nameHandle = open('LR_word2vec_evaluation_results.txt', 'w')
nameHandle.write('aspect_strict_Acc:\t'+ str(aspect_strict_Acc))
nameHandle.write('\naspect_Macro_F1:\t' + str(aspect_Macro_F1))
nameHandle.write('\naspect_Macro_AUC:\t' + str(aspect_Macro_AUC))
nameHandle.write('\n\nsentiment_Acc:\t' + str(sentiment_Acc))
nameHandle.write('\nsentiment_Macro_AUC:\t' + str(sentiment_Macro_AUC))
nameHandle.close()
```
| github_jupyter |
```
import pandas as pd
import numpy as np
from scipy.spatial.distance import cdist
from scipy.stats import multivariate_normal
from matplotlib import pyplot as plt
n_dots = 5000
data_20x = pd.read_csv("/mnt/data/Imaging/202105-Deconwolf/data_210726/20x_dw_calc/001_allDots.csv").sort_values("Value", ascending=False).iloc[:n_dots, :].reset_index(drop=True)
data_60x = pd.read_csv("/mnt/data/Imaging/202105-Deconwolf/data_210726/60x_dw_calc/001_allDots.csv").sort_values("Value", ascending=False).iloc[:n_dots, :].reset_index(drop=True)
voxel_side_20x = 325
voxel_side_60x = 108.3
rad = 16
def get_nn_ids(A: np.ndarray, B: np.ndarray, rad: float, verbose: bool = True) -> pd.Series:
paired_dist = cdist(A, B)
if verbose:
print(f"Smallest distance = {paired_dist.min():.3f}")
dist_threshold = rad * 2
paired_dist[paired_dist >= dist_threshold] = np.nan
not_singletons_idx = np.where(np.isnan(paired_dist).sum(1) != paired_dist.shape[0])[0]
if verbose:
print(f"Dots with neighbor within {dist_threshold} => {not_singletons_idx.shape[0]} / {paired_dist.shape[0]}")
return(pd.Series(data=np.nanargmin(paired_dist[not_singletons_idx, :], axis=1), index=not_singletons_idx, name="neighbour_id"))
id_match = get_nn_ids(data_20x.loc[:, ("x", "y")].values*3, data_60x.loc[:, ("x", "y")].values, rad)
id_match
plt.figure(figsize=(10, 10))
plt.plot(data_20x.loc[id_match.index, "x"], data_20x.loc[id_match.index, "y"], "r.")
plt.plot(data_60x.loc[id_match.values, "x"]/3, data_60x.loc[id_match.values, "y"]/3, "b.")
plt.title("Red: 20x. Blue: 60x.")
plt.show()
def get_deltas(A: np.ndarray, B: np.ndarray, rad: float) -> np.ndarray:
id_match = get_id_match(A, B, rad, False)
return(A[id_match.index] - B[id_match.values])
deltas0 = get_deltas(data_20x.loc[:, ("x", "y")].values*3, data_60x.loc[:, ("x", "y")].values, rad)
def get_mean_shift(deltas: np.ndarray, rad: float) -> np.ndarray:
v = np.array([0, 0])
for r in np.linspace(rad*2, .1, 200):
w = multivariate_normal.pdf(deltas, v, np.array([[2*rad, 0], [0, 2*rad]]))
v = w.dot(deltas)/w.sum()
return(v)
shift = get_mean_shift(deltas0, rad)
plt.figure(figsize=(10, 10))
plt.plot(data_20x.loc[id_match.index, "x"], data_20x.loc[id_match.index, "y"], "r.")
plt.plot(data_60x.loc[id_match.values, "x"]/3-delta[0], data_60x.loc[id_match.values, "y"]/3-delta[1], "b.")
plt.title("Red: 20x. Blue: 60x.")
plt.show()
deltas = get_deltas(data_20x.loc[:, ("x", "y")].values*3, data_60x.loc[:, ("x", "y")].values-shift, rad)
fig, ax = plt.subplots(figsize=(20, 10), nrows=1, ncols=2)
ax[0].plot(deltas0[:, 0], deltas0[:, 1], "k.")
ax[1].plot(deltas[:, 0], deltas[:, 1], "k.")
plt.show()
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from ipywidgets import interact, fixed
from scipy import stats
# Cofiguraciones
%matplotlib inline
plt.rcParams['figure.figsize'] = (10,10)
```
# Lectura de los datos
```
vehiculos = pd.read_pickle("./Data/vehiculos_variables_agrupadas.pkl")
vehiculos.head()
vehiculos.dtypes
```
# Distribución de variables numéricas
Podemos usar histogramas/diagrama de barras para ver la distribución de una variable
```
vehiculos["co2"].plot.hist()
```
También podemos usar un diagrama KDE para ver la función estimada de la distribución de una variable
```
vehiculos["co2"].plot.kde()
```
Para poder analizar la distribución de las variables numéricas de una forma más sencilla pordemos usar la librería ipywidgets, para ello tenemos que definir una función que dibuje el kde/histograma de una forma genérica y pasarle las series de la columnas que queremos analizar.
```
def distribucion_variable_numerica(df, col):
df[col].plot.kde()
plt.xlabel("Distribución de la variable {}".format(col))
plt.show()
columnas_numericas = vehiculos.select_dtypes(["int", "float"]).columns
interact(distribucion_variable_numerica, col=columnas_numericas, df=fixed(vehiculos))
```
A excepción de la variable cilindros, el resto parece seguir una distribucón normal. Para ver si sigue una distribución normal podemos realizar un gráfico de probabilidades.
```
def normalidad_variable_numerica(col):
stats.probplot(vehiculos[col], plot=plt)
plt.xlabel('Diagrama de Probabilidad(normal) de la variable {}'.format(col))
plt.show()
interact(normalidad_variable_numerica, col=columnas_numericas, df=fixed(vehiculos));
```
Las variables desplazamiento y cilindros no siguen una distribución normal, ya que de lo contrario mostraría la línea de 45º como sucenden en las otras variables.
Parece que siguen una distribución normal (co2, consumo_litros_milla), no obstante, conviene asegurarse haciendo un test de normalidad. En un test de normalidad, lo que queremos es rechazar la hipótesis nula de que la variable a analizar se ha obtenido de una población que sigue una distribución normal. Para un nivel de confianza de 95%, rechazamos la hipótesis nula si el p-value es inferior a 0.05. Esto es, si se obtiene un valor P (p-value) menor de 0.05, significa que las probabilidades de que la hipótesis nula sean ciertas es tan baja (menos de un 5%) que la rechazamos.
```
for num_col in columnas_numericas:
_, pval = stats.normaltest(vehiculos[num_col])
if(pval < 0.05):
print("Columna {} no sigue una distribución normal".format(num_col))
```
Como podemos observar ninguna de la variables númericas sigue una distribución normal de forma correcta.
# Distribución variables categóricas
La mejor forma de analizar la distribución de una variable categórica es usando value_count.
```
def distribucion_variable_categorica(col, df):
df[col].value_counts(ascending=True, normalize=True).tail(20).plot.barh()
plt.show()
columnas_categoricas = vehiculos.select_dtypes(["object", "category"]).columns
interact(distribucion_variable_categorica, col=columnas_categoricas, df=fixed(vehiculos))
```
# Conclusiones
- Ninguna variable numérica sigue una distribución normal.
- La variable numérica cilindros tiene una distribución de valores discretos no balanceada (cilindrada de 2 y 4 y 8 suman el 95% de los vehiculos). Podria agruparse como variable categórica (2, 4 , 8 y otro).
- El fabricante con la mayor cantidad de modelos es Chevrolet (10% del total).
- 65% de los vehiculos usan gasolina normal.
- La distribución de tamaños de motor y de consumo y co2 está equilibrada en todo el rango.
- 70% de los vehiculos usan traccion a las dos ruedas.
- Dos tercios de los coches tienen transmision automática.
- La clase mayoritaria de vehiculos es la de coches pequeños (35% del total).
- Los mayores fabricantes en cuanto a vehiculos analizados son los estadounidenses. Esto tiene sentido ya que la EPA es la agencia americana y probablemente es la que tiene más interes en estudiar coches de USA.
| github_jupyter |
### Naver
```
## Naver
"""Before execute code below, check out your kernel or jupyter notebook kernel environment
If you have problem, just copy this code and paste to yout jupyter notebook (recommended)
Also, before execute this page, execute this first >> "Get Chrome driver & dir setting.ipynb"
browser must be pop up on the screen : if the browser is in a state of minimization, results may go bad
(It does not matter covering the page with other page like jupyternotebook >> you can do other works)
If you have trouble with lxml, selenium, bs4, try to isntall module in anaconda prompt
>>> execute anconda prompt, try to [conda install lxml], [conda install selenium], [conda install bs4]
warning : If you try this code with high frequency, Search engine may ban your ip temporarily (for 5~10 minutes)
Refer to : Scroll_cnt=5 >>> about 100~200 imgs (depending on the searching word)"""
## Install module required
#!pip install lxml
#!pip install selenium
#!pip install bs4
## Import modules
import urllib.request
from bs4 import BeautifulSoup
from selenium import webdriver
from selenium.webdriver.common.keys import Keys
import time
##### Path ######################################################################
Chromedriver_PATH = 'c:\\chrome_webdriver\\chromedriver.exe' # Chromedriver PATH
save_path = 'D:\\images\\naver\\' #save_path
#################################################################################
## get userdata & parameters
Search_Tag = input("Input Search_Tag : ") # Search_Tag
scroll_cnt = int(input("Input scroll_cnt : ")) #scroll count
scrolltime = float(input("Input scroll_sleep_second >>> range(5~10) : ")) #Sleep time
## Get driver & open
driver = webdriver.Chrome(Chromedriver_PATH) # Chromedriver PATH
driver.get("https://search.naver.com/search.naver?where=image&sm=stb_nmr&")
driver.maximize_window()
time.sleep(1)
## input Search_Tag & Submit
elem = driver.find_element_by_id("nx_query")
elem.send_keys(Search_Tag)
time.sleep(1.5) #Do not remove >> if you remove this line, can't go next step
elem.submit()
time.sleep(3.0) #Do not remove
############## Functions ################################################################################
def fetch_list_url(): #parsing src url
imgList = soup.find_all("img", class_="_image")
for im in imgList:
params.append(im["src"])
return params
def fetch_detail_url(): #save src to local #changing save_path : Go to the top of this page (Path)
for idx,p in enumerate(params,1): #enumerate idx option 1 : get start index from 1 (default=0)
urllib.request.urlretrieve(p, save_path + Search_Tag + '_' + str(idx) + "_naver" + ".jpg")
###########################################################################################################
## Scrolling & Parsing
params=[]
for i in range(scroll_cnt):
html = driver.page_source #get source
soup = BeautifulSoup(html, "lxml")
params = fetch_list_url() #save the img_url to params
driver.execute_script("window.scrollTo(0, document.body.scrollHeight);") #scroll
time.sleep(scrolltime)
## Save imgs
print('')
print("Overlaped srcs : ", len(params))
params=list(dict.fromkeys(params)) #delete overlap #index URL >> https://m31phy.tistory.com/130
fetch_detail_url() #save img
print("Non_Overlap srcs : ", len(params))
driver.close() #close browser
```
| github_jupyter |
# k-NN movie reccomendation
| User\Film | Movie A | Movie B | Movie C | ... | Movie # |
|---------------------------------------------------------|
| **User A**| 3 | 4 | 0 | ... | 5 |
| **User B**| 0 | 3 | 2 | ... | 0 |
| **User C**| 4 | 1 | 3 | ... | 4 |
| **User D**| 5 | 3 | 2 | ... | 3 |
| ... | ... | ... | ... | ... | ... |
| **User #**| 2 | 1 | 1 | ... | 4 |
Task: For a new user find k similar users based on movie rating and recommend few new, previously unseen, movies to the new user. Use mean rating of k users to find which one to recommend. Use cosine similarity as distance function. User didnt't see a movie if he didn't rate the movie.
```
# Import necessary libraries
import tensorflow as tf
import numpy as np
# Define paramaters
set_size = 1000 # Number of users in dataset
n_features = 300 # Number of movies in dataset
K = 3 # Number of similary users
n_movies = 6 # Number of movies to reccomend
# Generate dummy data
data = np.array(np.random.randint(0, 6, size=(set_size, n_features)), dtype=np.float32)
new_user = np.array(np.random.randint(0, 6, size=(1, n_features)), dtype=np.float32)
# Find the number of movies that user did not rate
not_rated = np.count_nonzero(new_user == 0)
# Case in which the new user rated all movies in our dataset
if not_rated == 0:
print('Regenerate new user')
# Case in which we try to recommend more movies than user didn't see
if not_rated < n_movies:
print('Regenerate new user')
# Print few examples
# print(data[:3])
# print(new_user)
# Input train vector
X1 = tf.placeholder(dtype=tf.float32, shape=[None, n_features], name="X1")
# Input test vector
X2 = tf.placeholder(dtype=tf.float32, shape=[1, n_features], name="X2")
# Cosine similarity
norm_X1 = tf.nn.l2_normalize(X1, axis=1)
norm_X2 = tf.nn.l2_normalize(X2, axis=1)
cos_similarity = tf.reduce_sum(tf.matmul(norm_X1, tf.transpose(norm_X2)), axis=1)
with tf.Session() as sess:
# Find all distances
distances = sess.run(cos_similarity, feed_dict={X1: data, X2: new_user})
# print(distances)
# Find indices of k user with highest similarity
_, user_indices = sess.run(tf.nn.top_k(distances, K))
# print(user_indices)
# Get users rating
# print(data[user_indices])
# New user ratings
# print(new_user[0])
# NOTICE:
# There is a possibility that we can incorporate
# user for e.g. movie A which he didn't see.
movie_ratings = sess.run(tf.reduce_mean(data[user_indices], axis=0))
# print(movie_ratings)
# Positions where the new user doesn't have rating
# NOTICE:
# In random generating there is a possibility that
# the new user rated all movies in data set, if that
# happens regenerate the new user.
movie_indices = sess.run(tf.where(tf.equal(new_user[0], 0)))
# print(movie_indices)
# Pick only the avarege rating of movies that have been rated by
# other users and haven't been rated by the new user and among
# those movies pick n_movies for recommend to the new user
_, top_rated_indices = sess.run(tf.nn.top_k(movie_ratings[movie_indices].reshape(-1), n_movies))
# print(top_rated_indices)
# Indices of the movies with the highest mean rating, which new user did not
# see, from the k most similary users based on movie ratings
print('Movie indices to reccomend: ', movie_indices[top_rated_indices].T)
```
# Locally weighted regression (LOWESS)
```
# Import necessary libraries
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
tf.reset_default_graph()
# Load data as numpy array
x, y = np.loadtxt('../../data/02_LinearRegression/polynomial.csv', delimiter=',', unpack=True)
m = x.shape[0]
x = (x - np.mean(x, axis=0)) / np.std(x, axis=0)
y = (y - np.mean(y)) / np.std(y)
# Graphical preview
%matplotlib inline
fig, ax = plt.subplots()
ax.set_xlabel('X Labe')
ax.set_ylabel('Y Label')
ax.scatter(x, y, edgecolors='k', label='Data')
ax.grid(True, color='gray', linestyle='dashed')
X = tf.placeholder(tf.float32, name='X')
Y = tf.placeholder(tf.float32, name='Y')
w = tf.Variable(0.0, name='weights')
b = tf.Variable(0.0, name='bias')
point_x = -0.5
tau = 0.15 # 0.22
t_w = tf.exp(tf.div(-tf.pow(tf.subtract(X, point_x), 2), tf.multiply(tf.pow(tau, 2), 2)))
Y_predicted = tf.add(tf.multiply(X, w), b)
cost = tf.reduce_mean(tf.multiply(tf.square(Y - Y_predicted), t_w), name='cost')
optimizer = tf.train.GradientDescentOptimizer(learning_rate=0.001).minimize(cost)
with tf.Session() as sess:
# Initialize the necessary variables, in this case, w and b
sess.run(tf.global_variables_initializer())
# Train the model in 50 epochs
for i in range(500):
total_cost = 0
# Session runs train_op and fetch values of loss
for sample in range(m):
# Session looks at all trainable variables that loss depends on and update them
_, l = sess.run([optimizer, cost], feed_dict={X: x[sample], Y:y[sample]})
total_cost += l
# Print epoch and loss
if i % 50 == 0:
print('Epoch {0}: {1}'.format(i, total_cost / m))
# Output the values of w and b
w1, b1 = sess.run([w, b])
print(sess.run(t_w, feed_dict={X: 1.4}))
print('W: %f, b: %f' % (w1, b1))
print('Cost: %f' % sess.run(cost, feed_dict={X: x, Y: y}))
# Append hypothesis that we found on the plot
x1 = np.linspace(-1.0, 0.0, 50)
ax.plot(x1, x1 * w1 + b1, color='r', label='Predicted')
ax.plot(x1, np.exp(-(x1 - point_x) ** 2 / (2 * 0.15 ** 2)), color='g', label='Weight function')
ax.legend()
fig
```
| github_jupyter |
```
import pandas as pd
from pathlib import Path
import numpy as np
import seaborn as sns
from sklearn.pipeline import make_pipeline
import statsmodels.api as sm
from yellowbrick.model_selection import LearningCurve
from yellowbrick.regressor import ResidualsPlot
from yellowbrick.regressor import PredictionError
from sklearn.model_selection import train_test_split,RepeatedKFold
from sklearn.feature_selection import VarianceThreshold
from sklearn.preprocessing import StandardScaler, RobustScaler
import xgboost as xgb
from sklearn.externals import joblib
from sklearn.model_selection import GridSearchCV
from xgboost import XGBRegressor
from sklearn.metrics import mean_squared_error,mean_absolute_error,r2_score
import matplotlib.pyplot as plt
#import xgboost as xgb
%matplotlib inline
#dataframe final
df_final = pd.read_csv("../data/DF_train15_skempiAB_modeller_final.csv",index_col=0)
pdb_names = df_final.index
features_names = df_final.drop('ddG_exp',axis=1).columns
X = df_final.drop('ddG_exp',axis=1).astype(float)
y = df_final['ddG_exp']
# binned split
bins = np.linspace(0, len(X), 200)
y_binned = np.digitize(y, bins)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, stratify=y_binned,random_state=1)
sns.distplot( y_test , color="red", label="ddG_exp_test")
sns.distplot( y_train , color="skyblue", label="ddG_exp_train")
import numpy as np
from yellowbrick.model_selection import ValidationCurve
#1)
selector = VarianceThreshold()
#2)
lr_model = XGBRegressor(random_state=1212,n_jobs=-1,n_estimators=100)
#3) Crear pipeline
pipeline1 = make_pipeline(selector,lr_model)
viz = ValidationCurve(
pipeline1, njobs=-1,param_name="xgbregressor__reg_alpha",
param_range=[0.1,0.5,0.7,0.9,1.2], cv=10, scoring="r2"
)
#plt.ylim(0,0.6)
# Fit and poof the visualizer
viz.fit(X_train, y_train)
viz.poof()
XGBRegressor?
#1)
selector = VarianceThreshold()
#2)
xg_model = XGBRegressor(random_state=1212)
#3) Crear pipeline
pipeline1 = make_pipeline(selector,xg_model)
# grid params
param_grid = {'xgbregressor__colsample_bytree': [0.3],
'xgbregressor__subsample': [0.5],
'xgbregressor__n_estimators':[100],
'xgbregressor__max_depth': [6],
'xgbregressor__gamma': [3],
'xgbregressor__learning_rate': [0.07],
'xgbregressor__min_child_weight':[20],
'xgbregressor__reg_lambda': [2.7],
'xgbregressor__reg_alpha': [1.9],
'variancethreshold__threshold':[0.0]
}
cv = RepeatedKFold(n_splits=5,n_repeats=10,random_state=13)
# Instantiate the grid search model
grid1 = GridSearchCV(pipeline1, param_grid, verbose=5, n_jobs=-1,cv=cv,scoring=['neg_mean_squared_error','r2'],
refit='r2',return_train_score=True)
grid1.fit(X_train,y_train)
# index of best scores
rmse_bestCV_test_index = grid1.cv_results_['mean_test_neg_mean_squared_error'].argmax()
rmse_bestCV_train_index = grid1.cv_results_['mean_train_neg_mean_squared_error'].argmax()
r2_bestCV_test_index = grid1.cv_results_['mean_test_r2'].argmax()
r2_bestCV_train_index = grid1.cv_results_['mean_train_r2'].argmax()
# scores
rmse_bestCV_test_score = grid1.cv_results_['mean_test_neg_mean_squared_error'][rmse_bestCV_test_index]
rmse_bestCV_test_std = grid1.cv_results_['std_test_neg_mean_squared_error'][rmse_bestCV_test_index]
rmse_bestCV_train_score = grid1.cv_results_['mean_train_neg_mean_squared_error'][rmse_bestCV_train_index]
rmse_bestCV_train_std = grid1.cv_results_['std_train_neg_mean_squared_error'][rmse_bestCV_train_index]
r2_bestCV_test_score = grid1.cv_results_['mean_test_r2'][r2_bestCV_test_index]
r2_bestCV_test_std = grid1.cv_results_['std_test_r2'][r2_bestCV_test_index]
r2_bestCV_train_score = grid1.cv_results_['mean_train_r2'][r2_bestCV_train_index]
r2_bestCV_train_std = grid1.cv_results_['std_train_r2'][r2_bestCV_train_index]
print('CV test RMSE {:f} +/- {:f}'.format(np.sqrt(-rmse_bestCV_test_score),np.sqrt(rmse_bestCV_test_std)))
print('CV train RMSE {:f} +/- {:f}'.format(np.sqrt(-rmse_bestCV_train_score),np.sqrt(rmse_bestCV_train_std)))
print('CV test r2 {:f} +/- {:f}'.format(r2_bestCV_test_score,r2_bestCV_test_std))
print('CV train r2 {:f} +/- {:f}'.format(r2_bestCV_train_score,r2_bestCV_train_std))
print(r2_bestCV_train_score-r2_bestCV_test_score)
print("",grid1.best_params_)
y_test_pred = grid1.best_estimator_.predict(X_test)
y_train_pred = grid1.best_estimator_.predict(X_train)
print("\nRMSE for test dataset: {}".format(np.round(np.sqrt(mean_squared_error(y_test, y_test_pred)), 2)))
print("RMSE for train dataset: {}".format(np.round(np.sqrt(mean_squared_error(y_train, y_train_pred)), 2)))
print("pearson corr {:f}".format(np.corrcoef(y_test_pred,y_test)[0][1]))
print('R2 test',grid1.score(X_test,y_test))
print('R2 train',grid1.score(X_train,y_train))
visualizer = ResidualsPlot(grid1.best_estimator_,hist=False)
visualizer.fit(X_train, y_train) # Fit the training data to the model
visualizer.score(X_test, y_test) # Evaluate the model on the test data
visualizer.poof() # Draw/show/poof the data
perror = PredictionError(grid1.best_estimator_)
perror.fit(X_train, y_train) # Fit the training data to the visualizer
perror.score(X_test, y_test) # Evaluate the model on the test data
g = perror.poof()
viz = LearningCurve(grid1.best_estimator_, cv=cv, n_jobs=-1,scoring='r2',train_sizes=np.linspace(0.3, 1.0, 10))
viz.fit(X, y)
plt.ylim(0,0.6)
viz.poof()
final_xgb = grid1.best_estimator_.fit(X,y)
# save final model
joblib.dump(final_xgb, 'XGBmodel_train15skempiAB_FINAL.pkl')
from sklearn.base import BaseEstimator
from sklearn.model_selection import train_test_split
from xgboost import XGBRegressor, XGBClassifier
class XGBoostWithEarlyStop(BaseEstimator):
def __init__(self, early_stopping_rounds=5, test_size=0.1,
eval_metric='rmse', **estimator_params):
self.early_stopping_rounds = early_stopping_rounds
self.test_size = test_size
self.eval_metric=eval_metric='rmse'
if self.estimator is not None:
self.set_params(**estimator_params)
def set_params(self, **params):
return self.estimator.set_params(**params)
def get_params(self, **params):
return self.estimator.get_params()
def fit(self, X, y):
x_train, x_val, y_train, y_val = train_test_split(X, y, test_size=self.test_size)
self.estimator.fit(x_train, y_train,
early_stopping_rounds=self.early_stopping_rounds,
eval_metric=self.eval_metric, eval_set=[(x_val, y_val)])
return self
def predict(self, X):
return self.estimator.predict(X)
class XGBoostRegressorWithEarlyStop(XGBoostWithEarlyStop):
def __init__(self, *args, **kwargs):
self.estimator = XGBRegressor()
super(XGBoostRegressorWithEarlyStop, self).__init__(*args, **kwargs)
class XGBoostClassifierWithEarlyStop(XGBoostWithEarlyStop):
def __init__(self, *args, **kwargs):
self.estimator = XGBClassifier()
super(XGBoostClassifierWithEarlyStop, self).__init__(*args, **kwargs)
https://www.kaggle.com/c/santander-customer-satisfaction/discussion/20662
https://jakevdp.github.io/PythonDataScienceHandbook/05.03-hyperparameters-and-model-validation.html
print('CV test RMSE',np.sqrt(-grid.best_score_))
print('CV train RMSE',np.sqrt(-grid.cv_results_['mean_train_score'].max()))
#print('Training score (r2): {}'.format(r2_score(X_train, y_train)))
#print('Test score (r2): {}'.format(r2_score(X_test, y_test)))
print(grid.best_params_)
y_test_pred = grid.best_estimator_.predict(X_test)
y_train_pred = grid.best_estimator_.predict(X_train)
print("\nRoot mean square error for test dataset: {}".format(np.round(np.sqrt(mean_squared_error(y_test, y_test_pred)), 2)))
print("Root mean square error for train dataset: {}".format(np.round(np.sqrt(mean_squared_error(y_train, y_train_pred)), 2)))
print("pearson corr: ",np.corrcoef(y_test_pred,y_test)[0][1])
viz = LearningCurve(grid.best_estimator_, cv=5, nthreads=10,scoring='neg_mean_squared_error',train_sizes=np.linspace(.1, 1.0, 10))
viz.fit(X, y)
viz.poof()
```
# Metodologia completa Gridsearch , early stop
```
XGBRegressor?
import numpy as np
from yellowbrick.model_selection import ValidationCurve
#1)
selector = VarianceThreshold()
#2)
lr_model = XGBoostRegressorWithEarlyStop(random_state=1212,n_jobs=-1,n_estimators=30)
#3) Crear pipeline
pipeline1 = make_pipeline(selector,lr_model)
viz = ValidationCurve(
pipeline1, njobs=-1,param_name="xgboostregressorwithearlystop__subsample",
param_range=[0.5,0.6,0.7,0.8,0.9,1], cv=10, scoring="r2"
)
#plt.ylim(0,0.6)
# Fit and poof the visualizer
viz.fit(X_train, y_train)
viz.poof()
import os
if __name__ == "__main__":
# NOTE: on posix systems, this *has* to be here and in the
# `__name__ == "__main__"` clause to run XGBoost in parallel processes
# using fork, if XGBoost was built with OpenMP support. Otherwise, if you
# build XGBoost without OpenMP support, you can use fork, which is the
# default backend for joblib, and omit this.
try:
from multiprocessing import set_start_method
except ImportError:
raise ImportError("Unable to import multiprocessing.set_start_method."
" This example only runs on Python 3.4")
#set_start_method("forkserver")
import numpy as np
import pandas as pd
from pathlib import Path
from sklearn.metrics import mean_squared_error,mean_absolute_error,r2_score
from sklearn.model_selection import GridSearchCV
from sklearn.model_selection import train_test_split
from sklearn.datasets import load_boston
import xgboost as xgb
from sklearn.base import BaseEstimator
from sklearn.model_selection import train_test_split
from xgboost import XGBRegressor, XGBClassifier
class XGBoostWithEarlyStop(BaseEstimator):
def __init__(self, early_stopping_rounds=10, test_size=0.15,
eval_metric='rmse', random_state=1212,**estimator_params):
self.early_stopping_rounds = early_stopping_rounds
self.test_size = test_size
self.eval_metric=eval_metric='rmse'
if self.estimator is not None:
self.set_params(**estimator_params)
def set_params(self, **params):
return self.estimator.set_params(**params)
def get_params(self, **params):
return self.estimator.get_params()
def fit(self, X, y):
x_train, x_val, y_train, y_val = train_test_split(X, y, test_size=self.test_size)
self.estimator.fit(x_train, y_train,
early_stopping_rounds=self.early_stopping_rounds,
eval_metric=self.eval_metric, eval_set=[(x_val, y_val)])
return self
def predict(self, X):
return self.estimator.predict(X)
class XGBoostRegressorWithEarlyStop(XGBoostWithEarlyStop):
def __init__(self, *args, **kwargs):
self.estimator = XGBRegressor()
super(XGBoostRegressorWithEarlyStop, self).__init__(*args, **kwargs)
class XGBoostClassifierWithEarlyStop(XGBoostWithEarlyStop):
def __init__(self, *args, **kwargs):
self.estimator = XGBClassifier()
super(XGBoostClassifierWithEarlyStop, self).__init__(*args, **kwargs)
# Load data
ABPRED_DIR = Path().cwd().parent
DATA = ABPRED_DIR / "data"
#dataframe final
df_final = pd.read_csv(DATA/"../data/DF_contact400_energy_sasa.FcorrZero.csv",index_col=0)
pdb_names = df_final.index
features_names = df_final.drop('ddG_exp',axis=1).columns
# Data final
X = df_final.drop('ddG_exp',axis=1).astype(float)
y = df_final['ddG_exp']
#Split data
# split for final test
X_train, X_test, y_train, y_test = train_test_split(X, y,train_size=0.75,random_state=1212)
njob = 4
os.environ["OMP_NUM_THREADS"] = str(njob) # or to whatever you want
xgb_model = XGBoostRegressorWithEarlyStop()
param_grid = {'colsample_bytree': [0.7],
'subsample': [1],
'n_estimators':[1000],
'max_depth': [10],
'gamma': [1],
'learning_rate': [0.05],
'min_child_weight':[1],
'reg_lambda': [0],
'reg_alpha': [10],
'colsample_bylevel':[0.9],
'random_state':[1212]}
grid = GridSearchCV(xgb_model, param_grid, verbose=5, n_jobs=njob,cv=10,scoring=['neg_mean_squared_error','r2'],
refit='r2',return_train_score=True)
grid.fit(X_train, y_train)
# index of best scores
rmse_bestCV_test_index = grid.cv_results_['mean_test_neg_mean_squared_error'].argmax()
rmse_bestCV_train_index = grid.cv_results_['mean_train_neg_mean_squared_error'].argmax()
r2_bestCV_test_index = grid.cv_results_['mean_test_r2'].argmax()
r2_bestCV_train_index = grid.cv_results_['mean_train_r2'].argmax()
# scores
rmse_bestCV_test_score = grid.cv_results_['mean_test_neg_mean_squared_error'][rmse_bestCV_test_index]
rmse_bestCV_test_std = grid.cv_results_['std_test_neg_mean_squared_error'][rmse_bestCV_test_index]
rmse_bestCV_train_score = grid.cv_results_['mean_train_neg_mean_squared_error'][rmse_bestCV_train_index]
rmse_bestCV_train_std = grid.cv_results_['std_train_neg_mean_squared_error'][rmse_bestCV_train_index]
r2_bestCV_test_score = grid.cv_results_['mean_test_r2'][r2_bestCV_test_index]
r2_bestCV_test_std = grid.cv_results_['std_test_r2'][r2_bestCV_test_index]
r2_bestCV_train_score = grid.cv_results_['mean_train_r2'][r2_bestCV_train_index]
r2_bestCV_train_std = grid.cv_results_['std_train_r2'][r2_bestCV_train_index]
print('CV test RMSE {:f} +/- {:f}'.format(np.sqrt(-rmse_bestCV_test_score),np.sqrt(rmse_bestCV_test_std)))
print('CV train RMSE {:f} +/- {:f}'.format(np.sqrt(-rmse_bestCV_train_score),np.sqrt(rmse_bestCV_train_std)))
print('CV test r2 {:f} +/- {:f}'.format(r2_bestCV_test_score,r2_bestCV_test_std))
print('CV train r2 {:f} +/- {:f}'.format(r2_bestCV_train_score,r2_bestCV_train_std))
print(r2_bestCV_train_score-r2_bestCV_test_score)
print("",grid.best_params_)
y_test_pred = grid.best_estimator_.predict(X_test)
y_train_pred = grid.best_estimator_.predict(X_train)
print("\nRMSE for test dataset: {}".format(np.round(np.sqrt(mean_squared_error(y_test, y_test_pred)), 2)))
print("RMSE for train dataset: {}".format(np.round(np.sqrt(mean_squared_error(y_train, y_train_pred)), 2)))
print("pearson corr {:f}".format(np.corrcoef(y_test_pred,y_test)[0][1]))
viz = LearningCurve(grid.best_estimator_, n_jobs=-1,cv=10, scoring='neg_mean_squared_error')
viz.fit(X, y)
viz.poof()
perror = PredictionError(grid.best_estimator_)
perror.fit(X_train, y_train) # Fit the training data to the visualizer
perror.score(X_test, y_test) # Evaluate the model on the test data
g = perror.poof()
visualizer = ResidualsPlot(grid.best_estimator_)
visualizer.fit(X_train, y_train) # Fit the training data to the model
visualizer.score(X_test, y_test) # Evaluate the model on the test data
visualizer.poof() # Draw/show/poof the data
perror = PredictionError(model)
perror.fit(X_train_normal, y_train_normal) # Fit the training data to the visualizer
perror.score(X_test_normal.values, y_test_normal.values) # Evaluate the model on the test data
g = perror.poof()
```
| github_jupyter |
# Django2.2
**Python Web Framework**:<https://wiki.python.org/moin/WebFrameworks>
先说句大实话,Web端我一直都是`Net技术站`的`MVC and WebAPI`,Python我一直都是用些数据相关的知识(爬虫、简单的数据分析等)Web这块只是会Flask,其他框架也都没怎么接触过,都说`Python`的`Django`是`建站神器`,有`自动生成后端管理页面`的功能,于是乎就接触了下`Django2.2`(目前最新版本)
> 逆天点评:Net的MVC最擅长的就是(通过Model+View)`快速生成前端页面和对应的验证`,而Python的`Django`最擅长的就是(通过注册Model)`快速生成后台管理页面`。**这两个语言都是快速建站的常用编程语言**(项目 V1~V2 阶段)
网上基本上都是Django1.x的教程,很多东西在2下都有点不适用,所以简单记录下我的学习笔记以及一些心得:
> PS:ASP.Net MVC相关文章可以参考我16年写的文章:<https://www.cnblogs.com/dunitian/tag/MVC/>
官方文档:<https://docs.djangoproject.com/zh-hans/2.2/releases/2.2/>
## 1.环境
### 1.虚拟环境
这个之前的确没太大概念,我一直都是用Conda来管理不同版本的包,现在借助Python生态圈里的工具`virtualenv`和`virtualenvwapper`
---
### 2.Django命令
1.**创建一个空项目:`django-admin startproject 项目名称`**
> PS:项目名不要以数字开头哦~
```shell
# 创建一个base_demo的项目
django-admin startproject base_demo
# 目录结构
|-base_demo (文件夹)
|---__init__.py(说明这个文件夹是一个Python包)
|---settings.py(项目配置文件:创建应用|模块后进行配置)
|---urls.py(URL路由配置)
|---wsgi.py(遵循wsgi协议:web服务器和Django的交互入口)
|-manage.py(项目管理文件,用来生成应用|模块)
```
2.**创建一个应用:`python manage.py startapp 应用名称`**
> 项目中一个模块就是一个应用,eg:商品模块、订单模块等
```shell
# 创建一个用户模块
python manage.py startapp users
├─base_demo
│ __init__.py
│ settings.py
│ urls.py
│ wsgi.py
├─manage.py(项目管理文件,用来生成应用|模块)
│
└─users(新建的模块|应用)
│ │ __init__.py
│ │ admin.py(后台管理相关)
│ │ models.py(数据库相关模型)
│ │ views.py(相当于MVC中的C,用来定义处理|视图函数)
│ │ tests.py(写测试代码)
│ │ apps.py:配置应用的元数据(可选)
│ │
│ └─migrations:数据迁移模块(根据Model内容生成的)
│ __init__.py
```
**PS:记得在项目(`base_demo`)的settings.py注册一下应用模块哦~**
```py
INSTALLED_APPS = [
......
'users', # 注册自己创建的模块|应用
]
```
3.**运行项目:`python manage.py runserver`**
> PS:指定端口:`python manage.py runserver 8080`
## 2.MVT入门
**大家都知道MVC(模型-视图-控制器),而Django的MVC叫做MVT(模型-视图-模版)**
> PS:Django出来很早,名字是自己定义的,用法和理念是一样的
### 2.1.M(模型)
#### 2.1.1.类的定义
- 1.**生成迁移文件:`python manage.py makemigrations`**
- PS:根据编写好的Model文件生成(模型里面可以不用定义ID属性)
- 2.**执行迁移生成表:`python mange.py migrate`**
- PS:执行生成的迁移文件
PS:类似于EF的`CodeFirst`,Django默认使用的是`sqlite`,更改数据库后面会说的
先看个演示案例:
**1.定义类文件**(会根据Code来生成DB)
```py
# users > models.py
from django.db import models
# 用户信息表
class UserInfo(models.Model):
# 字符串类型,最大长度为20
name = models.CharField(max_length=20)
# 创建时间:日期类型
create_time = models.DateTimeField()
# 更新时间
update_time = models.DateTimeField()
```
**2. 生成数据库**
```shell
# 生成迁移文件
> python manage.py makemigrations
Migrations for 'userinfo':
userinfo\migrations\0001_initial.py
- Create model UserInfo
# 执行迁移生成表
> python manage.py migrate
Operations to perform:
Apply all migrations: admin, auth, contenttypes, sessions, userinfo
Running migrations:
Applying contenttypes.0001_initial... OK
Applying auth.0001_initial... OK
Applying admin.0001_initial... OK
Applying admin.0002_logentry_remove_auto_add... OK
Applying admin.0003_logentry_add_action_flag_choices... OK
Applying contenttypes.0002_remove_content_type_name... OK
Applying auth.0002_alter_permission_name_max_length... OK
Applying auth.0003_alter_user_email_max_length... OK
Applying auth.0004_alter_user_username_opts... OK
Applying auth.0005_alter_user_last_login_null... OK
Applying auth.0006_require_contenttypes_0002... OK
Applying auth.0007_alter_validators_add_error_messages... OK
Applying auth.0008_alter_user_username_max_length... OK
Applying auth.0009_alter_user_last_name_max_length... OK
Applying sessions.0001_initial... OK
Applying userinfo.0001_initial... OK
```
然后就自动生成对应的表了 ==> **`users_userinfo`**(应用名_模块中的类名)
知识拓展:默认时间相关文章:<https://www.cnblogs.com/huchong/p/7895263.html>
#### 2.1.2.生成后台
##### 1.配置本地化(设置后台管理页面是中文)
主要就是修改`settings.py`文件的`语言`和`时区`(后台管理的语言和时间)
```py
# 使用中文(zh-hans可以这么记==>zh-汉'字')
LANGUAGE_CODE = 'zh-hans'
# 设置中国时间
TIME_ZONE = 'Asia/Shanghai'
```
##### 2.创建管理员
**创建系统管理员:`python manage.py createsuperuser`**
```shell
python manage.py createsuperuser
用户名 (leave blank to use 'win10'): dnt # 如果不填,默认是计算机用户名
电子邮件地址: # 可以不设置
Password:
Password (again):
Superuser created successfully.
```
**经验:如果忘记密码可以创建一个新管理员账号,然后把旧的删掉就行了**
> PS:根据新password字段,修改下旧账号的password也可以
课后拓展:<a href="https://blog.csdn.net/dsjakezhou/article/details/84319228">修改django后台管理员密码</a>
##### 3.后台管理页面
主要就是**在admin中注册模型类**
比如给之前创建的UserInfo类创建对应的管理页面:
```py
# base_demo > users > admin.py
from users.models import UserInfo
# from .models import UserInfo
# 注册模型类(自动生成后台管理页面)
admin.site.register(UserInfo) # .site别忘记
```
然后运行Django(`python manage.py runserver`),访问"127.0.0.1:8080/admin",登录后就就可以管理了
> PS:如果不想交admin,而是想在root下。那么可以修改项目的`urls.py`(后面会说)
##### 4.制定化显示
注册模型类就ok了,但是显示稍微有点不人性化,eg:
列表页显示出来的标题是UserInfo对象,而我们平时一般显示用户名等信息
so ==> 可以自己改写下
回顾下之前讲的:(程序是显示的`str(对象)`,那么我们重写魔方方法`__str__`即可改写显示了)
```py
# base_demo > users > models.py
# 用户信息表
class UserInfo(models.Model):
# 字符串类型,最大长度为20
name = models.CharField(max_length=20)
# 创建时间:日期类型
create_time = models.DateTimeField()
# 更新时间
update_time = models.DateTimeField()
def __str__(self):
"""为了后台管理页面的美化"""
return self.name
```
这时候再访问就美化了:(**不用重启Django**)
Django就没有提供对应的方法?NoNoNo,我们继续看:
```py
# base_demo > users > admin.py
from .models import UserInfo
# 自定义模型管理页面
class UserInfoAdmin(admin.ModelAdmin):
# 自定义管理页面的列表显示字段(和类属性相对应)
list_display = ["id", "name", "create_time", "update_time", "datastatus"]
# 注册模型类和模型管理类(自动生成后台管理页面)
admin.site.register(UserInfo, UserInfoAdmin)
```
其他什么都不用修改,后端管理列表的布局就更新了:
> PS:设置Model的`verbose_name`就可以在后台显示中文,eg:`name = models.CharField(max_length=25, verbose_name="姓名")`
还有更多个性化的内容后面会继续说的~
### 2.3.V(视图)
这个类比于MVC的C(控制器)
> PS:这块比Net的MVC和Python的Flask要麻烦点,url地址要简单配置下映射关系(小意思,不花太多时间)
这块刚接触稍微有点绕,所以我们借助图来看:
**比如我们想访问users应用下的首页(`/users/index`)**
#### 2.3.1.设置视图函数
这个和定义控制器里面的方法没区别:
> PS:函数必须含`request`(类比下类方法必须含的self)
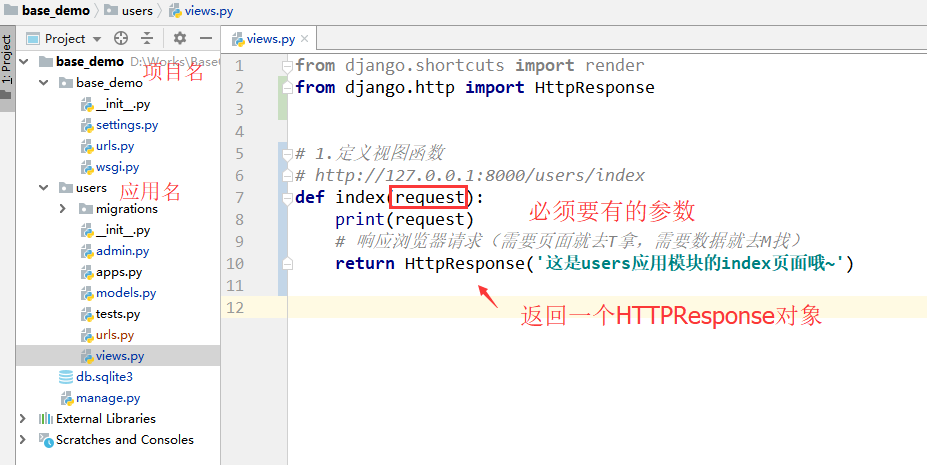
```py
from django.http import HttpResponse
# 1.定义视图函数
# http://127.0.0.1:8000/users/index
def index(request):
print(request)
# 响应浏览器请求(需要页面就去T拿,需要数据就去M找)
return HttpResponse('这是users应用模块的index页面哦~')
```
#### 2.3.2.配置路由
因为我想要的地址是:`/users/index`,那么我在项目urls中也需要配置下访问`/users`的路由规则:
> PS:我是防止以后模块多了管理麻烦,所以分开写,要是你只想在一个urls中配置也无妨
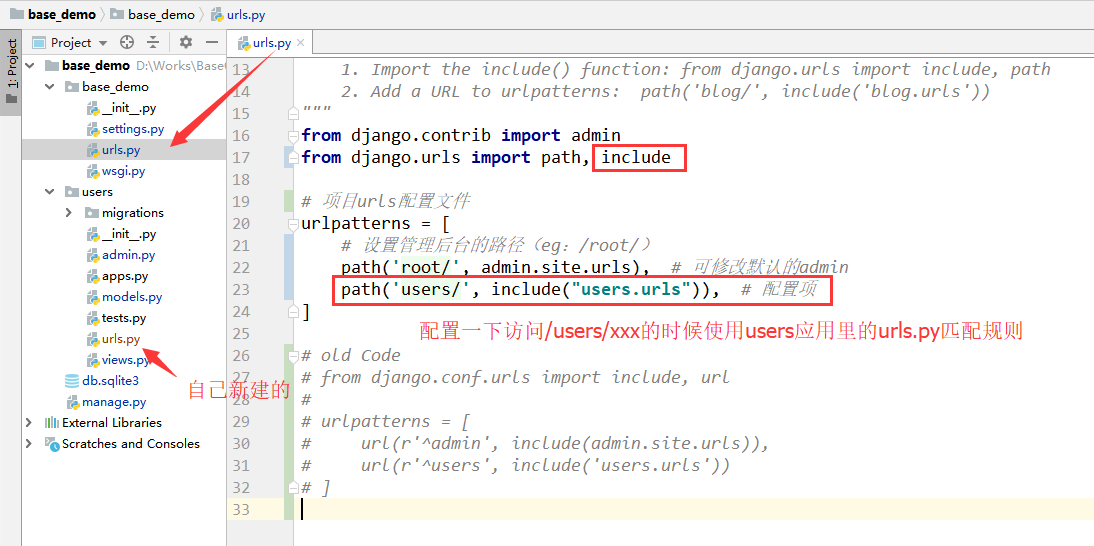
```py
# base_demo > urls.py
from django.contrib import admin
from django.urls import path, include
# 项目urls配置文件
urlpatterns = [
path('users/', include("users.urls")), # 配置项
]
```
最后再贴一下users应用模块的匹配:
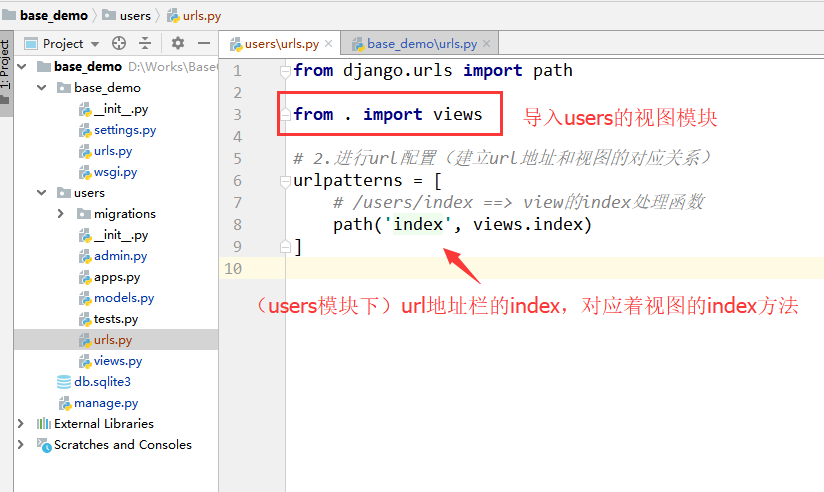
```py
# users > urls.py
from django.urls import path
from . import views
# 2.进行url配置(建立url地址和视图的对应关系)
urlpatterns = [
# /users/index ==> view的index处理函数
path('index', views.index),
]
```
#### 2.3.3.url访问
这时候你访问`127.0.0.1:8000/users/index`就可以了:
简单说下这个过程:
1. 先去项目的urls.py中进行匹配
- `path('users/', include("users.urls")), # 配置项`
2. 发现只要是以`/users/`开头的都使用了`users`模块自己的`urls.py`来匹配
- `path('index', views.index),`
3. 发现访问`/users/index`最后进入的视图函数是`index`
4. 然后执行`def index(request):pass`里面的内容并返回
### 2.4.T(模版)
这个类比于MVC的V,我们来看个简单案例:
#### 2.4.1.创建模版
Django1.x版本需要配置下模版路径之类的,现在只要在对应模块下创建`templates`文件夹就可以直接访问了
我们来定义一个list的模版:
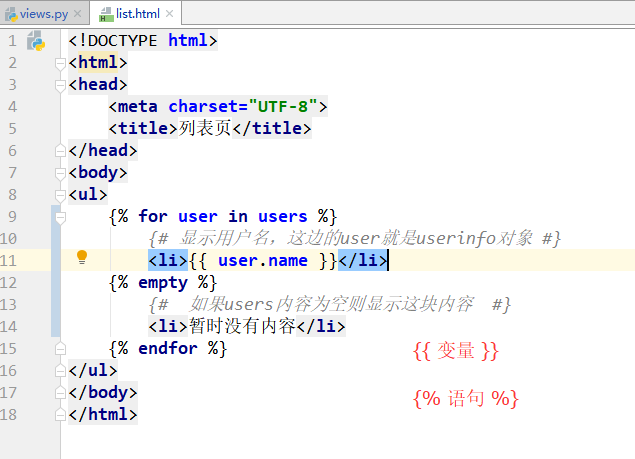
定义视图函数(类比定义控制器方法)
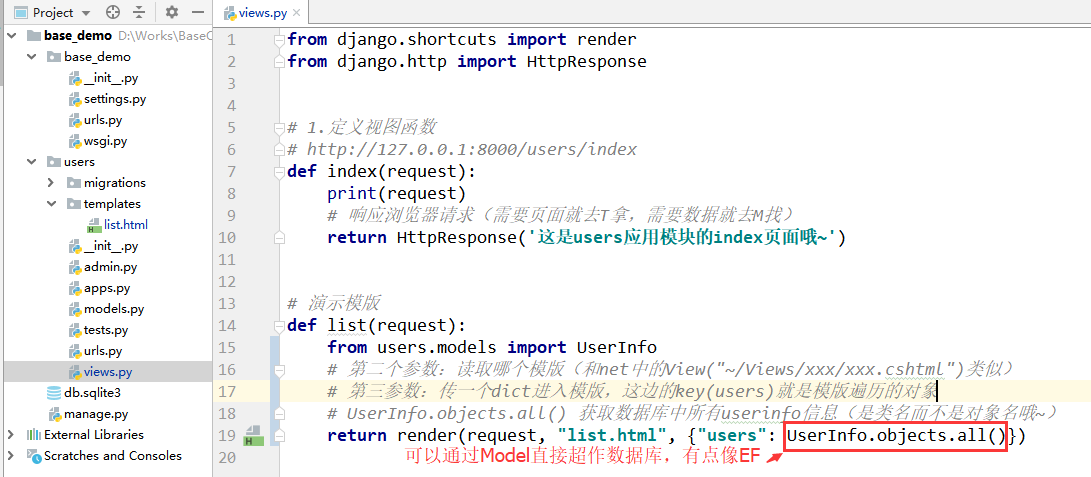
配置对应的路由:
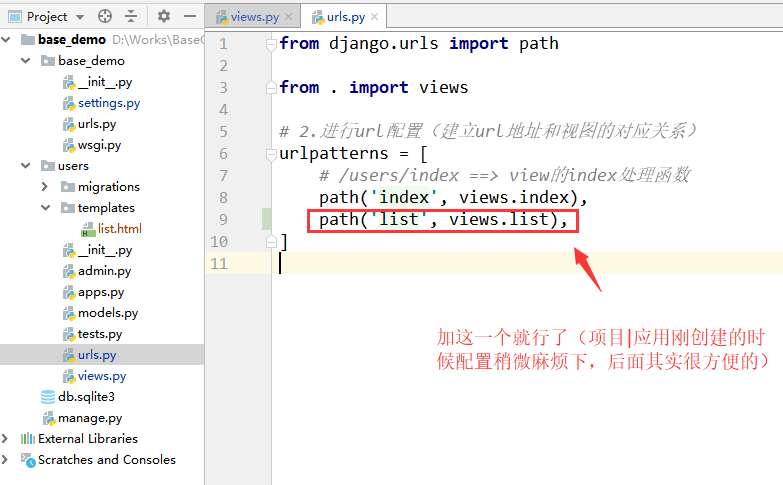
然后就出效果了:
如果把之前添加的数据删除掉,也会显示默认效果:
#### 2.4.2.指定模版
也可以指定模版位置:(看个人习惯)
打开项目`settings.py`文件,设置`TEMPLATES`的`DIRS`值,来指定默认模版路径:
```py
# Base_dir:当前项目的绝对路径
# BASE_DIR = os.path.dirname(os.path.dirname(os.path.abspath(__file__)))
# https://docs.djangoproject.com/zh-hans/2.2/ref/settings/#templates
TEMPLATES = [
{
...
'DIRS': [os.path.join(BASE_DIR, 'templates')], # 模版文件的绝对路径
...
},
]
```
### 扩展:使用MySQL数据库
这篇详细流程可以查看之前写的文章:<a href="" title="https://www.cnblogs.com/dotnetcrazy/p/10782441.html" target="_blank">稍微记录下Django2.2使用MariaDB和MySQL遇到的坑</a>
这边简单过下即可:
#### 1.创建数据库
Django不会帮你创建数据库,需要自己创建,eg:`create database django charset=utf8;`
#### 2.配置数据库
我把对应的文档url也贴了:
```py
# https://docs.djangoproject.com/en/2.2/ref/settings/#databases
DATABASES = {
'default': {
'ENGINE': 'django.db.backends.mysql',
'NAME': 'django', # 使用哪个数据库
'USER': 'root', # mysql的用户名
'PASSWORD': 'dntdnt', # 用户名对应的密码
'HOST': '127.0.0.1', # 数据库服务的ip地址
'PORT': 3306, # 对应的端口
# https://docs.djangoproject.com/en/2.2/ref/settings/#std:setting-OPTIONS
'OPTIONS': {
# https://docs.djangoproject.com/zh-hans/2.2/ref/databases/#setting-sql-mode
# SQLMode可以看我之前写的文章:https://www.cnblogs.com/dotnetcrazy/p/10374091.html
'init_command': "SET sql_mode='STRICT_TRANS_TABLES'", # 设置SQL_Model
},
}
}
```
最小配置:

项目init.py文件中配置:
```py
import pymysql
# Django使用的MySQLdb对Python3支持力度不够,我们用PyMySQL来代替
pymysql.install_as_MySQLdb()
```
图示:
#### 3.解决干扰
如果你的Django是最新的2.2,PyMySQL也是最新的0.93的话,你会发现Django会报错:
> django.core.exceptions.ImproperlyConfigured: mysqlclient 1.3.13 or newer is required; you have 0.9.3.
这个是Django对MySQLdb版本的限制,我们使用的是PyMySQL,所以不用管它
再继续运行发现又冒了个错误:`AttributeError: 'str' object has no attribute 'decode'`
这个就不能乱改了,所以先调试输出下:
发现是对字符串进行了decode解码操作:(一般对字符串进行编码,二进制进行解码)
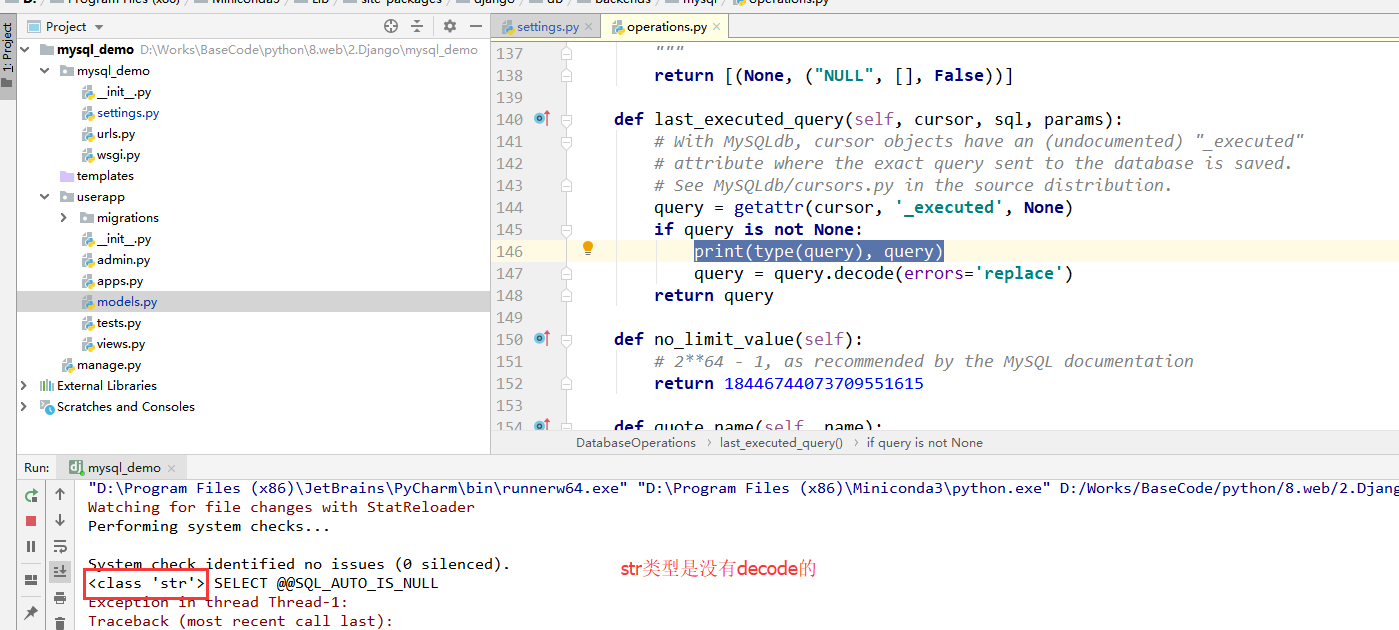
解决也很简单,改成encode即可
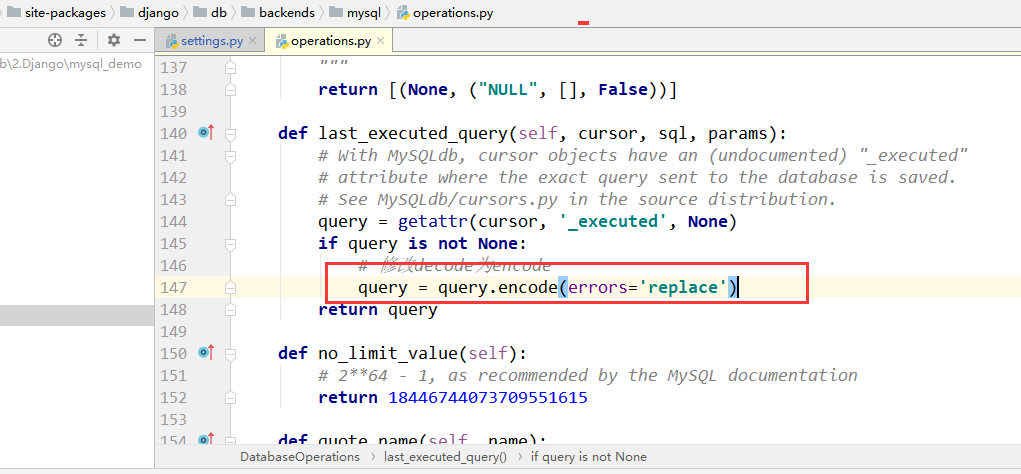
然后就没问题了,之后创建新项目也不会有问题了
### 扩展:避免命令忘记
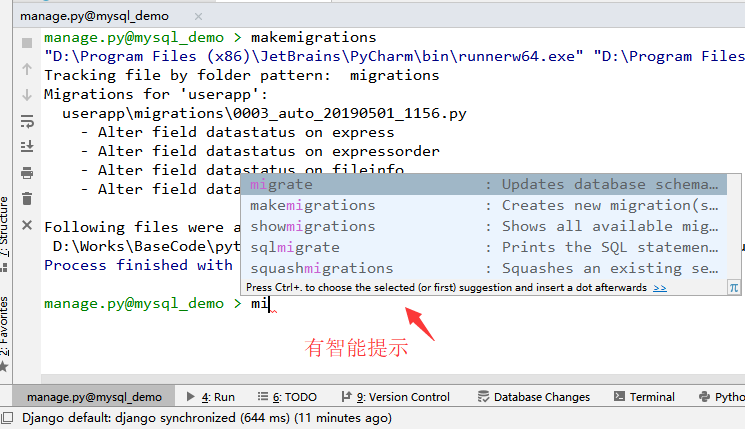
## 3.MVT基础
### 3.1.M基础
#### 3.1.字段类型
模型类的命名规则基本上和变量命名一致,然后添加一条:**不能含`__`**(双下划线)
> PS:这个后面讲查询的时候你就秒懂了(`__`来间隔关键词)
这边简单罗列下**常用字段类型**:**模型类数据库字段的定义:`属性名=models.字段类型(选项)`**
| 字段类型 | 备注 |
| ----------------------------------------------------------------------- | ---------------------------------------------------------------------------- |
| `AutoField` | **自增长的int类型**(Django默认会自动创建属性名为id的自动增长属性) |
| `BigAutoField` | **自增长的bigint类型**(Django默认会自动创建属性名为id的自动增长属性) |
| **`BooleanField`** | **布尔类型**,值为True或False |
| `NullBooleanField` | **可空布尔类型**,支持Null、True、False三种值 |
| **`CharField(max_length=最大长度)`** | `varchar`**字符串**。参数max_length表示最大字符个数 |
| **`TextField`** | **大文本类型**,一般超过4000个字符时使用。 |
| **`IntegerField`** | **整型** |
| `BigIntegerField` | **长整型** |
| **`DecimalField(max_digits=None, decimal_places=None)`** | **十进制浮点数**,`max_digits`:总位数。`decimal_places`:小数占几位 |
| `FloatField(max_digits=None, decimal_places=None)` | **浮点数**,`max_digits`:总位数。`decimal_places`:小数占几位 |
| `DateField([auto_now=True] | [auto_now_add=True])` | **日期类型**,`auto_now_add`:自动设置创建时间,`auto_now`:自动设置修改时间 |
| `TimeField([auto_now=True] | [auto_now_add=True])` | **时间类型**,参数同`DateField` |
| **`DateTimeField([auto_now=True] | [auto_now_add=True])`** | **日期时间类型**,参数同`DateField` |
| **`UUIDField([primary_key=True,] default=uuid.uuid4, editable=False)`** | **UUID字段** |
后端常用字段:
| 字段类型 | 备注 |
| ---------------- | ----------------------------------------------------------- |
| `EmailField` | `CharField`子类,专门用于**Email**的字段 |
| **`FileField`** | **文件字段** |
| **`ImageField`** | **图片字段**,FileField子类,对内容进行校验以保证是有效图片 |
#### 3.2.字段选项
通过选项可以约束数据库字段,简单罗列几个常用的:
| 字段选项 | 描述 |
| -------------- | -------------------------------------------------------- |
| **`default`** | `default=函数名|默认值`设置字段的**默认值** |
| `primary_key` | `primary_key=True`设置**主键**(一般在`AutoField`中使用) |
| `unique` | `unique=True`设置**唯一键** |
| **`db_index`** | `db_index=True`设置**索引** |
| `db_column` | `db_column='xx'`设置数据库的字段名(默认就是属性名) |
| `null` | 字段是否可以为`null`(现在基本上都是不为`null`) |
和Django自动生成的后台管理相关的选项:(后台管理页面表单验证)
| 管理页的表单选项 | 描述 |
| ------------------ | ---------------------------------------------------------------------- |
| **`blank`** | `blank=True`设置表单验证中字段**是否可以为空** |
| **`verbose_name`** | `verbose_name='xxx'`设置字段对应的**中文显示**(下划线会转换为空格) |
| **`help_text`** | `help_text='xxx'`设置字段对应的**文档提示**(可以包含HTML) |
| `editable` | `editable=False`设置字段**是否可编辑**(不可编辑就不显示在后台管理页) |
| `validators` | `validators=xxx`设置字段**验证**(<http://mrw.so/4LzsEq>) |
补充说明:
1. 除非要覆盖默认的主键行为,否则不需要设置任何字段的`primary_key=True`(Django默认会创建`AutoField`来保存主键)
2. ***`auto_now_add`和`auto_now`是互斥的,一个字段中只能设置一个,不能同时设置**
3. 修改模型类时:如果添加的选项不影响表的结构,就不需要重新迁移
- 字段选项中`default`和`管理表单选项`(`blank`、`verbose_name`、`help_text`等)不影响表结构
### ORM基础
官方文档:<https://docs.djangoproject.com/zh-hans/2.2/ref/models/querysets/>
`O`(objects):类和对象,`R`(Relation):关系型数据库,`M`(Mapping):映射
> PS:表 --> 类、每行数据 --> 对象、字段 --> 对象的属性
进入命令模式:python manager.py shell
增(有连接关系的情况)
删(逻辑删除、删)
改(内连接关联修改)
查(总数、条件查询据、分页查询)
表与表之间的关系(relation),主要有这三种:
1. 一对一(one-to-one):一种对象与另一种对象是一一对应关系
- eg:一个学生只能在一个班级。
2. 一对多(one-to-many): 一种对象可以属于另一种对象的多个实例
- eg:一张唱片包含多首歌。
3. 多对多(many-to-many):两种对象彼此都是"一对多"关系
- eg:比如一张唱片包含多首歌,同时一首歌可以属于多张唱片。
#### 执行SQL语句
官方文档:<https://docs.djangoproject.com/zh-hans/2.2/topics/db/sql/>
#### 扩展:查看生成SQL
课后拓展:<https://www.jianshu.com/p/b69a7321a115>
#### 3.3.模型管理器类
1. 改变查询的结果集
- eg:程序里面都是假删除,而默认的`all()`把那些假删除的数据也查询出来了
2. 添加额外的方法
- eg:
---
### 3.2.V基础
返回Json格式(配合Ajax)
JsonResponse
#### 3.2.1.
#### URL路由
#### 模版配置
#### 重定向
跳转到视图函数
知识拓展:<https://www.cnblogs.com/attila/p/10420702.html>
#### 动态生成URL
模版中动态生成URL地址,类似于Net里面的`@Url.Action("Edit","Home",new {id=13})`
> <https://docs.djangoproject.com/zh-hans/2.2/intro/tutorial03/#removing-hardcoded-urls-in-templates>
#### 404和500页面
调试信息关闭
### 扩展:HTTPRequest
**HTTPRequest的常用属性**:
1. **`path`**:请求页面的完整路径(字符串)
- 不含域名和参数
2. **`method`**:请求使用的HTTP方式(字符串)
- eg:`GET`、`POST`
3. `encoding`:提交数据的编码方式(字符串)
- 如果为`None`:使用浏览器默认设置(PS:一般都是UTF-8)
4. **`GET`**:包含get请求方式的所有参数(QueryDict类型,类似于Dict)
5. **`POST`**:包含post请求方式的所有参数(类型同上)
6. **`FILES`**:包含所有上传的文件(MultiValueDict类型,类似于Dict)
7. **`COOKIES`**:以key-value形式包含所有cookie(Dict类型)
8. **`session`**:状态保持使用(类似于Dict)
#### 获取URL参数
复选框:勾选on,不勾选None
reques.POST.get("xxx")
#### Cookies
基于域名来存储的,如果不指定过期时间则关闭浏览器就过期
```py
# set cookie
response = Htpresponse对象(eg:JsonResponse,Response)
# max_age:多少秒后过期
# response.set_cookie(key,value,max_age=7*24*3600) # 1周过期
# expires:过期时间,timedelta:时间间隔
response.set_cookie(key,value,expires=datatime.datatime.now()+datatime.timedelta(days=7))
return response;
# get cookie
value = request.COOKIES.get(key)
```
**PS:Cookie不管保存什么类型,取出来都是str字符串**
知识拓展:<https://blog.csdn.net/cuishizun/article/details/81537316>
#### Session
Django的Session信息存储在`django_session`里面,可以根据sessionid(`session_key`)获取对应的`session_data`值
**PS:Session之所以依赖于Cookie,是因为Sessionid(唯一标识)存储在客户端,没有sessionid你怎么获取?**
```py
# 设置
request.session["key"] = value
# 获取
request.session.get("key",默认值)
# 删除指定session
del request.session["key"] # get(key) ?
# 删除所有session的value
# sessiondata里面只剩下了sessionid,而对于的value变成了{}
request.session.clear()
# 删除所有session(数据库内容全删了)
request.session.flush()
# 设置过期时间(默认过期时间是2周)
request.session.set_expiry(不活动多少秒后失效)
# PS:都是request里面的方法
```
**PS:Session保存什么类型,取出来就是什么类型**(Cookie取出来都是str)
---
#### 文件上传
---
### 3.3.T基础
自定义404页面
## 4.Admin后台
上面演示了一些简单的制定化知识点:<a href="#2.1.2.生成后台">上节回顾</a>,现在简单归纳下`Django2.2`admin相关设置:
### 4.1.修改后台管理页面的标题
大致效果如下:
在`admin.py`中设置`admin.site.site_header`和`admin.site.site_title`:
### 4.2.修改app在Admin后台显示的名称
大致效果如下:

先设置应用模块的中文名:**`verbose_name = 'xxx'`**
让配置生效:**`default_app_config = '应用名.apps.应用名Config'`**
### 4.3.汉化显示应用子项
大致效果如下:
在每个模型类中设置**`Meta`**类,并设置`verbose_name`和`verbose_name_plural`
### 4.4.汉化表单字段和提示
大致效果如下:
汉化表单的字段:**`verbose_name`**,显示字段提示:**`help_text`**
### 4.5.
列表显示
状态显示+字体颜色
文件上传
文本验证
Tag过滤
apt install sqliteman
| github_jupyter |
```
ls -l| tail -10
#G4
from google.colab import drive
drive.mount('/content/gdrive')
cp gdrive/My\ Drive/fingerspelling5.tar.bz2 fingerspelling5.tar.bz2
# rm fingerspelling5.tar.bz2
# cd /media/datastorage/Phong/
!tar xjf fingerspelling5.tar.bz2
cd dataset5
mkdir surrey
mkdir surrey/E
mv dataset5/* surrey/E/
cd ..
#remove depth files
import glob
import os
import shutil
# get parts of image's path
def get_image_parts(image_path):
"""Given a full path to an image, return its parts."""
parts = image_path.split(os.path.sep)
#print(parts)
filename = parts[2]
filename_no_ext = filename.split('.')[0]
classname = parts[1]
train_or_test = parts[0]
return train_or_test, classname, filename_no_ext, filename
#del_folders = ['A','B','C','D','E']
move_folders_1 = ['A','B','C','D']
move_folders_2 = ['E']
# look for all images in sub-folders
for folder in move_folders_1:
class_folders = glob.glob(os.path.join(folder, '*'))
for iid_class in class_folders:
#move depth files
class_files = glob.glob(os.path.join(iid_class, 'depth*.png'))
print('copying %d files' %(len(class_files)))
for idx in range(len(class_files)):
src = class_files[idx]
if "0001" not in src:
train_or_test, classname, _, filename = get_image_parts(src)
dst = os.path.join('train_depth', classname, train_or_test+'_'+ filename)
# image directory
img_directory = os.path.join('train_depth', classname)
# create folder if not existed
if not os.path.exists(img_directory):
os.makedirs(img_directory)
#copying
shutil.copy(src, dst)
else:
print('ignor: %s' %src)
#move color files
for iid_class in class_folders:
#move depth files
class_files = glob.glob(os.path.join(iid_class, 'color*.png'))
print('copying %d files' %(len(class_files)))
for idx in range(len(class_files)):
src = class_files[idx]
train_or_test, classname, _, filename = get_image_parts(src)
dst = os.path.join('train_color', classname, train_or_test+'_'+ filename)
# image directory
img_directory = os.path.join('train_color', classname)
# create folder if not existed
if not os.path.exists(img_directory):
os.makedirs(img_directory)
#copying
shutil.copy(src, dst)
# look for all images in sub-folders
for folder in move_folders_2:
class_folders = glob.glob(os.path.join(folder, '*'))
for iid_class in class_folders:
#move depth files
class_files = glob.glob(os.path.join(iid_class, 'depth*.png'))
print('copying %d files' %(len(class_files)))
for idx in range(len(class_files)):
src = class_files[idx]
if "0001" not in src:
train_or_test, classname, _, filename = get_image_parts(src)
dst = os.path.join('test_depth', classname, train_or_test+'_'+ filename)
# image directory
img_directory = os.path.join('test_depth', classname)
# create folder if not existed
if not os.path.exists(img_directory):
os.makedirs(img_directory)
#copying
shutil.copy(src, dst)
else:
print('ignor: %s' %src)
#move color files
for iid_class in class_folders:
#move depth files
class_files = glob.glob(os.path.join(iid_class, 'color*.png'))
print('copying %d files' %(len(class_files)))
for idx in range(len(class_files)):
src = class_files[idx]
train_or_test, classname, _, filename = get_image_parts(src)
dst = os.path.join('test_color', classname, train_or_test+'_'+ filename)
# image directory
img_directory = os.path.join('test_color', classname)
# create folder if not existed
if not os.path.exists(img_directory):
os.makedirs(img_directory)
#copying
shutil.copy(src, dst)
# #/content
%cd ..
ls -l
mkdir surrey/E/checkpoints
cd surrey/
#MUL 1 - Inception - ST
from keras.applications import MobileNet
# from keras.applications import InceptionV3
# from keras.applications import Xception
# from keras.applications.inception_resnet_v2 import InceptionResNetV2
from tensorflow.keras.applications import EfficientNetB0
from keras.models import Model
from keras.layers import concatenate
from keras.layers import Dense, GlobalAveragePooling2D, Input, Embedding, SimpleRNN, LSTM, Flatten, GRU, Reshape
# from keras.applications.inception_v3 import preprocess_input
# from tensorflow.keras.applications.efficientnet import preprocess_input
from keras.applications.mobilenet import preprocess_input
from keras.layers import GaussianNoise
def get_adv_model():
# f1_base = EfficientNetB0(include_top=False, weights='imagenet',
# input_shape=(299, 299, 3),
# pooling='avg')
# f1_x = f1_base.output
f1_base = MobileNet(weights='imagenet', include_top=False, input_shape=(224,224,3))
f1_x = f1_base.output
f1_x = GlobalAveragePooling2D()(f1_x)
# f1_x = f1_base.layers[-151].output #layer 5
# f1_x = GlobalAveragePooling2D()(f1_x)
# f1_x = Flatten()(f1_x)
# f1_x = Reshape([1,1280])(f1_x)
# f1_x = SimpleRNN(2048,
# return_sequences=False,
# # dropout=0.8
# input_shape=[1,1280])(f1_x)
#Regularization with noise
f1_x = GaussianNoise(0.1)(f1_x)
f1_x = Dense(1024, activation='relu')(f1_x)
f1_x = Dense(24, activation='softmax')(f1_x)
model_1 = Model(inputs=[f1_base.input],outputs=[f1_x])
model_1.summary()
return model_1
from keras.callbacks import Callback
import pickle
import sys
#Stop training on val_acc
class EarlyStoppingByAccVal(Callback):
def __init__(self, monitor='val_acc', value=0.00001, verbose=0):
super(Callback, self).__init__()
self.monitor = monitor
self.value = value
self.verbose = verbose
def on_epoch_end(self, epoch, logs={}):
current = logs.get(self.monitor)
if current is None:
warnings.warn("Early stopping requires %s available!" % self.monitor, RuntimeWarning)
if current >= self.value:
if self.verbose > 0:
print("Epoch %05d: early stopping" % epoch)
self.model.stop_training = True
#Save large model using pickle formate instead of h5
class SaveCheckPoint(Callback):
def __init__(self, model, dest_folder):
super(Callback, self).__init__()
self.model = model
self.dest_folder = dest_folder
#initiate
self.best_val_acc = 0
self.best_val_loss = sys.maxsize #get max value
def on_epoch_end(self, epoch, logs={}):
val_acc = logs['val_acc']
val_loss = logs['val_loss']
if val_acc > self.best_val_acc:
self.best_val_acc = val_acc
# Save weights in pickle format instead of h5
print('\nSaving val_acc %f at %s' %(self.best_val_acc, self.dest_folder))
weigh= self.model.get_weights()
#now, use pickle to save your model weights, instead of .h5
#for heavy model architectures, .h5 file is unsupported.
fpkl= open(self.dest_folder, 'wb') #Python 3
pickle.dump(weigh, fpkl, protocol= pickle.HIGHEST_PROTOCOL)
fpkl.close()
# model.save('tmp.h5')
elif val_acc == self.best_val_acc:
if val_loss < self.best_val_loss:
self.best_val_loss=val_loss
# Save weights in pickle format instead of h5
print('\nSaving val_acc %f at %s' %(self.best_val_acc, self.dest_folder))
weigh= self.model.get_weights()
#now, use pickle to save your model weights, instead of .h5
#for heavy model architectures, .h5 file is unsupported.
fpkl= open(self.dest_folder, 'wb') #Python 3
pickle.dump(weigh, fpkl, protocol= pickle.HIGHEST_PROTOCOL)
fpkl.close()
# Training
import tensorflow as tf
from keras.preprocessing.image import ImageDataGenerator
from keras.callbacks import TensorBoard, ModelCheckpoint, EarlyStopping, CSVLogger, ReduceLROnPlateau
from keras.optimizers import Adam
import time, os
from math import ceil
train_datagen = ImageDataGenerator(
# rescale = 1./255,
rotation_range=30,
width_shift_range=0.3,
height_shift_range=0.3,
shear_range=0.3,
zoom_range=0.3,
# horizontal_flip=True,
# vertical_flip=True,##
# brightness_range=[0.5, 1.5],##
channel_shift_range=10,##
fill_mode='nearest',
# preprocessing_function=get_cutout_v2(),
preprocessing_function=preprocess_input,
)
test_datagen = ImageDataGenerator(
# rescale = 1./255
preprocessing_function=preprocess_input
)
NUM_GPU = 1
batch_size = 64
train_set = train_datagen.flow_from_directory('surrey/E/train_color/',
target_size = (224, 224),
batch_size = batch_size,
class_mode = 'categorical',
shuffle=True,
seed=7,
# subset="training"
)
valid_set = test_datagen.flow_from_directory('surrey/E/test_color/',
target_size = (224, 224),
batch_size = batch_size,
class_mode = 'categorical',
shuffle=False,
seed=7,
# subset="validation"
)
model_txt = 'st'
# Helper: Save the model.
savedfilename = os.path.join('surrey', 'E', 'checkpoints', 'Surrey_MobileNet_E_tmp.hdf5')
checkpointer = ModelCheckpoint(savedfilename,
monitor='val_accuracy', verbose=1,
save_best_only=True, mode='max',save_weights_only=True)########
# Helper: TensorBoard
tb = TensorBoard(log_dir=os.path.join('svhn_output', 'logs', model_txt))
# Helper: Save results.
timestamp = time.time()
csv_logger = CSVLogger(os.path.join('svhn_output', 'logs', model_txt + '-' + 'training-' + \
str(timestamp) + '.log'))
earlystopping = EarlyStoppingByAccVal(monitor='val_accuracy', value=0.9900, verbose=1)
epochs = 40##!!!
lr = 1e-3
decay = lr/epochs
optimizer = Adam(lr=lr, decay=decay)
# train on multiple-gpus
# Create a MirroredStrategy.
strategy = tf.distribute.MirroredStrategy()
print("Number of GPUs: {}".format(strategy.num_replicas_in_sync))
# Open a strategy scope.
with strategy.scope():
# Everything that creates variables should be under the strategy scope.
# In general this is only model construction & `compile()`.
model_mul = get_adv_model()
model_mul.compile(optimizer=optimizer,loss='categorical_crossentropy',metrics=['accuracy'])
step_size_train=ceil(train_set.n/train_set.batch_size)
step_size_valid=ceil(valid_set.n/valid_set.batch_size)
# step_size_test=ceil(testing_set.n//testing_set.batch_size)
# result = model_mul.fit_generator(
# generator = train_set,
# steps_per_epoch = step_size_train,
# validation_data = valid_set,
# validation_steps = step_size_valid,
# shuffle=True,
# epochs=epochs,
# callbacks=[checkpointer],
# # callbacks=[csv_logger, checkpointer, earlystopping],
# # callbacks=[tb, csv_logger, checkpointer, earlystopping],
# verbose=1)
# Training
import tensorflow as tf
from keras.preprocessing.image import ImageDataGenerator
from keras.callbacks import TensorBoard, ModelCheckpoint, EarlyStopping, CSVLogger, ReduceLROnPlateau
from keras.optimizers import Adam
import time, os
from math import ceil
train_datagen = ImageDataGenerator(
# rescale = 1./255,
rotation_range=30,
width_shift_range=0.3,
height_shift_range=0.3,
shear_range=0.3,
zoom_range=0.3,
# horizontal_flip=True,
# vertical_flip=True,##
# brightness_range=[0.5, 1.5],##
channel_shift_range=10,##
fill_mode='nearest',
# preprocessing_function=get_cutout_v2(),
preprocessing_function=preprocess_input,
)
test_datagen = ImageDataGenerator(
# rescale = 1./255
preprocessing_function=preprocess_input
)
NUM_GPU = 1
batch_size = 64
train_set = train_datagen.flow_from_directory('surrey/E/train_color/',
target_size = (224, 224),
batch_size = batch_size,
class_mode = 'categorical',
shuffle=True,
seed=7,
# subset="training"
)
valid_set = test_datagen.flow_from_directory('surrey/E/test_color/',
target_size = (224, 224),
batch_size = batch_size,
class_mode = 'categorical',
shuffle=False,
seed=7,
# subset="validation"
)
model_txt = 'st'
# Helper: Save the model.
savedfilename = os.path.join('gdrive', 'My Drive', 'Surrey_ASL', '4_Surrey_MobileNet_E.hdf5')
checkpointer = ModelCheckpoint(savedfilename,
monitor='val_accuracy', verbose=1,
save_best_only=True, mode='max',save_weights_only=True)########
# Helper: TensorBoard
tb = TensorBoard(log_dir=os.path.join('svhn_output', 'logs', model_txt))
# Helper: Save results.
timestamp = time.time()
csv_logger = CSVLogger(os.path.join('svhn_output', 'logs', model_txt + '-' + 'training-' + \
str(timestamp) + '.log'))
earlystopping = EarlyStoppingByAccVal(monitor='val_accuracy', value=0.9900, verbose=1)
epochs = 40##!!!
lr = 1e-3
decay = lr/epochs
optimizer = Adam(lr=lr, decay=decay)
# train on multiple-gpus
# Create a MirroredStrategy.
strategy = tf.distribute.MirroredStrategy()
print("Number of GPUs: {}".format(strategy.num_replicas_in_sync))
# Open a strategy scope.
with strategy.scope():
# Everything that creates variables should be under the strategy scope.
# In general this is only model construction & `compile()`.
model_mul = get_adv_model()
model_mul.compile(optimizer=optimizer,loss='categorical_crossentropy',metrics=['accuracy'])
step_size_train=ceil(train_set.n/train_set.batch_size)
step_size_valid=ceil(valid_set.n/valid_set.batch_size)
# step_size_test=ceil(testing_set.n//testing_set.batch_size)
result = model_mul.fit_generator(
generator = train_set,
steps_per_epoch = step_size_train,
validation_data = valid_set,
validation_steps = step_size_valid,
shuffle=True,
epochs=epochs,
callbacks=[checkpointer],
# callbacks=[csv_logger, checkpointer, earlystopping],
# callbacks=[tb, csv_logger, checkpointer, earlystopping],
verbose=1)
print(savedfilename)
ls -l
# Open a strategy scope.
with strategy.scope():
model_mul.load_weights(os.path.join('gdrive', 'My Drive', 'Surrey_ASL', '4_Surrey_MobileNet_E.hdf5'))
# Helper: Save the model.
savedfilename = os.path.join('gdrive', 'My Drive', 'Surrey_ASL', '4_Surrey_MobileNet_E_L2.hdf5')
checkpointer = ModelCheckpoint(savedfilename,
monitor='val_accuracy', verbose=1,
save_best_only=True, mode='max',save_weights_only=True)########
epochs = 15##!!!
lr = 1e-4
decay = lr/epochs
optimizer = Adam(lr=lr, decay=decay)
# Open a strategy scope.
with strategy.scope():
model_mul.compile(optimizer=optimizer,loss='categorical_crossentropy',metrics=['accuracy'])
result = model_mul.fit_generator(
generator = train_set,
steps_per_epoch = step_size_train,
validation_data = valid_set,
validation_steps = step_size_valid,
shuffle=True,
epochs=epochs,
callbacks=[checkpointer],
# callbacks=[csv_logger, checkpointer, earlystopping],
# callbacks=[tb, csv_logger, checkpointer, earlystopping],
verbose=1)
# Open a strategy scope.
with strategy.scope():
model_mul.load_weights(os.path.join('gdrive', 'My Drive', 'Surrey_ASL', '4_Surrey_MobileNet_E_L2.hdf5'))
# Helper: Save the model.
savedfilename = os.path.join('gdrive', 'My Drive', 'Surrey_ASL', '4_Surrey_MobileNet_E_L3.hdf5')
checkpointer = ModelCheckpoint(savedfilename,
monitor='val_accuracy', verbose=1,
save_best_only=True, mode='max',save_weights_only=True)########
epochs = 15##!!!
lr = 1e-5
decay = lr/epochs
optimizer = Adam(lr=lr, decay=decay)
# Open a strategy scope.
with strategy.scope():
model_mul.compile(optimizer=optimizer,loss='categorical_crossentropy',metrics=['accuracy'])
result = model_mul.fit_generator(
generator = train_set,
steps_per_epoch = step_size_train,
validation_data = valid_set,
validation_steps = step_size_valid,
shuffle=True,
epochs=epochs,
callbacks=[checkpointer],
# callbacks=[csv_logger, checkpointer, earlystopping],
# callbacks=[tb, csv_logger, checkpointer, earlystopping],
verbose=1)
from sklearn.metrics import classification_report, confusion_matrix
import numpy as np
test_datagen = ImageDataGenerator(
rescale = 1./255)
testing_set = test_datagen.flow_from_directory('dataset5/test_color/',
target_size = (224, 224),
batch_size = 32,
class_mode = 'categorical',
seed=7,
shuffle=False
# subset="validation"
)
y_pred = model.predict_generator(testing_set,steps = testing_set.n//testing_set.batch_size)
y_pred = np.argmax(y_pred, axis=1)
y_true = testing_set.classes
print(confusion_matrix(y_true, y_pred))
# print(model.evaluate_generator(testing_set,
# steps = testing_set.n//testing_set.batch_size))
```
| github_jupyter |
# Am I feeding my network crap
Given that my research on the image content of optical flow images shows such huge variety is my image generation doing anything useful to it??? Perhaps experiment with a very small network for say only 10 classes??
First lets look at the output for something relatively easy like cricket
```
import os
import sys
up1 = os.path.abspath('../../utils/')
up2 = os.path.abspath('../../models/')
sys.path.insert(0, up1)
sys.path.insert(0, up2)
from optical_flow_data_gen import DataGenerator
from ucf101_data_utils import get_test_data_opt_flow, get_train_data_opt_flow
from motion_network import getKerasCifarMotionModel2, getKerasCifarMotionModelOnly
from keras.optimizers import SGD
from matplotlib import pyplot as plt
from keras.optimizers import SGD
import cv2
import numpy as np
```
# Is it the data or my classifier
I am starting to wonder what it is about my optical flow data that might be causing so much easier. Regardless about the unconverged flow images I feel the author of the data still managed with it. So there's essentially two things I can either get a large amount of improvement simply on how I train my classifier (slower?), or my data set is not quite right. I've already seen that I wasn't even doing any random transforms on my opt flow images courtesy my badly written opt flow data generator.
Any how what I am aiming to do is use a stinkingly cheap data model to explore what might be wrong.
```
from keras.models import Sequential
from keras.layers import Conv2D, MaxPooling2D
from keras.layers import Activation, Dropout, Flatten, Dense
from keras.initializers import Ones
from keras import optimizers
def getModel(lr=1e-2):
model = Sequential()
model.add(Conv2D(32, (3, 3), input_shape=(224, 224, 2)))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(32, (3, 3)))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(64, (3, 3)))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Flatten()) # this converts our 3D feature maps to 1D feature vectors
model.add(Dense(64))
model.add(Activation('relu'))
model.add(Dropout(0.5))
model.add(Dense(10))
model.add(Activation('sigmoid'))
optimizers.SGD(lr=lr)
model.compile(loss='categorical_crossentropy',
optimizer='sgd',
metrics=['accuracy'])
return model
from optical_flow_data_gen import DataGenerator
from ucf101_data_utils import get_test_data_opt_flow, get_train_data_opt_flow
training_options = { 'rescale' : 1./255,
'shear_range' : 0.2,
'zoom_range' : 0.2,
'horizontal_flip' : True,
'rotation_range':20,
'width_shift_range':0.2,
'height_shift_range':0.2}
validation_options = { 'rescale' : 1./255 }
params_train = { 'data_dir' : "/data/tvl1_flow",
'dim': (224,224),
'batch_size': 16,
'n_frames': 1,
'n_frequency': 1,
'shuffle': True,
'n_classes' : 10,
'validation' : False,
'enable_augmentation' : True}
params_valid = { 'data_dir' : "/data/tvl1_flow",
'dim': (224,224),
'batch_size': 16,
'n_frames': 1,
'n_frequency': 1,
'shuffle': True,
'n_classes' : 10,
'validation' : True,
'enable_augmentation' : False}
id_labels_train = get_train_data_opt_flow('../../data/ucf101_splits/trainlist01_small.txt')
labels = id_labels_train[1]
id_test = get_test_data_opt_flow('../../data/ucf101_splits/testlist01_small.txt', \
'../../data/ucf101_splits/classInd_small.txt')
training_generator = DataGenerator(*id_labels_train, **params_train)
validation_generator = DataGenerator(id_test[0], id_test[1], **params_valid)
model_with_data_aug = getModel(lr=1e-2)
mod1 = model_with_data_aug.fit_generator(generator=training_generator,
validation_data=validation_generator,
use_multiprocessing=True,
workers=2, epochs=5,
verbose=1)
mod1 = model_fast_lr.fit_generator(generator=training_generator,
validation_data=validation_generator,
use_multiprocessing=True,
workers=2, epochs=10,
verbose=1)
plt.plot(mod1.history['acc'])
model_slow_lr = getModel(lr=1e-4)
mod1 = model_slow_lr.fit_generator(generator=training_generator,
validation_data=validation_generator,
use_multiprocessing=True,
workers=2, epochs=15,
verbose=1)
```
| github_jupyter |
# Mean Reversion on Futures
by Rob Reider and Maxwell Margenot
Part of the Quantopian Lecture Series:
* [www.quantopian.com/lectures](https://www.quantopian.com/lectures)
* [github.com/quantopian/research_public](https://github.com/quantopian/research_public)
Notebook released under the Creative Commons Attribution 4.0 License.
Introducing futures as an asset opens up trading opportunities that were previously unavailable. In this lecture we will look at strategies involving one future against another. We will also look at strategies involving trades of futures and stocks at the same time. Other strategies, like trading calendar spreads (futures on the same commodity but with different delivery months) will be topics for future lectures.
```
import numpy as np
import pandas as pd
import statsmodels.api as sm
from statsmodels.tsa.stattools import coint, adfuller
import matplotlib.pyplot as plt
from quantopian.research.experimental import continuous_future, history
```
## Pairs of Futures (or Spreads)
Before we look at trading pairs of futures contracts, let's quickly review pairs trading in general and cointegration. For full lectures on these topics individually, see the [lecture on stationarity and cointegration](https://www.quantopian.com/lectures/integration-cointegration-and-stationarity) and the [lecture on pairs trading](https://www.quantopian.com/lectures/introduction-to-pairs-trading).
When markets are efficient, the prices of assets are often modeled as [random walks](https://en.wikipedia.org/wiki/Random_walk_hypothesis):
$$P*t=\mu+P*{t-1}+\epsilon_t$$
We can then take the difference of the prices to get white noise, stationary time series:
$$r*t=P_t-P*{t-1}= \mu +\epsilon_t$$
Of course, if prices follow a random walk, they are completely unforecastable, so the goal is to find returns that are correlated with something in the past. For example, if we can find an asset whose price is a little mean-reverting (and therefore its returns are negatively autocorrelated), we can use that to forecast future returns.
## Cointegration and Mean Reversion
The idea behind cointegration is that even if the prices of two different assets both follow random walks, it is still possible that a linear combination of them is not a random walk. A common analogy is of a dog owner walking his dog with a retractable leash. If you look at the position of the dog owner, it may follow a random walk, and if you look at the dog separately, it also may follow a random walk, but the distance between them, the difference of their positions, may very well be mean reverting. If the dog is behind the owner, he may run to catch up and if the dog is ahead, the length of the leash may prevent him from getting too far away.
The dog and its owner are linked together and their distance is a mean reverting process. In cointegration, we look for assets that are economically linked, so that if $P_t$ and $Q_t$ are both random walks, the linear combination, $P_t - b Q_t$, may not itself be a random walk and may be forecastable.
## Finding a Feasible Spread
For stocks, a natural starting point for identifying cointegrated pairs is looking at stocks in the same industry. However, competitors are not necessarily economic substitutes. Think of Apple and Blackberry. It's not always the case that when one of those company's stock price jumps up, the other catches up. The economic link is fairly tenuous. Here it is more like the dog broke the leash and ran away from the owner.
However, with pairs of futures, there may be economic forces that link the two prices. Consider heating oil and natural gas. Some power plants have the ability to use either one, depending on which has become cheaper. So when heating oil has dipped below natural gas, increased demand for heating oil will push it back up. Platinum and Palladium are substitutes for some types of catalytic converters used for emission control. Corn and wheat are substitutes for animal feed. Corn and sugar are substitutes as sweeteners. There are many potential links to examine and test.
Let's go through a specific example of futures prices that might be cointegrated.
## Soybean Crush
The difference in price between soybeans and their refined products is referred to as the "crush spread". It represents the processing margin from "crushing" a soybean into its refined products. Note that we scale up the futures price of soybean oil so that it is the same magnitude as the price for soybean meal. It certainly seems from the plots that the prices of the refined products move together.
```
soy_meal_mult = symbols('SMF17').multiplier
soy_oil_mult = symbols('BOF17').multiplier
soybean_mult = symbols('SYF17').multiplier
sm_future = continuous_future('SM', offset=0, roll='calendar', adjustment='mul')
sm_price = history(sm_future, fields='price', start_date='2014-01-01', end_date='2017-01-01')
bo_future = continuous_future('BO', offset=0, roll='calendar', adjustment='mul')
bo_price = history(bo_future, fields='price', start_date='2014-01-01', end_date='2017-01-01')
sm_price.plot()
bo_price.multiply(soy_oil_mult//soy_meal_mult).plot()
plt.ylabel('Price')
plt.legend(['Soybean Meal', 'Soybean Oil']);
```
However, from looking at the p-value for our test, we conclude that soybean meal and soybean meal and soybean oil are not cointegrated.
```
print 'p-value: ', coint(sm_price, bo_price)[1]
```
We still have this compelling economic link, though. Both soybean oil and soybean meal have a root product in soybeans themselves. Let's see if we can suss out any signal by creating a spread between soybean prices and the refined products together, by implementing the [crush spread](https://en.wikipedia.org/wiki/Crush_spread).
```
sm_future = continuous_future('SM', offset=1, roll='calendar', adjustment='mul')
sm_price = history(sm_future, fields='price', start_date='2014-01-01', end_date='2017-01-01')
bo_future = continuous_future('BO', offset=1, roll='calendar', adjustment='mul')
bo_price = history(bo_future, fields='price', start_date='2014-01-01', end_date='2017-01-01')
sy_future = continuous_future('SY', offset=0, roll='calendar', adjustment='mul')
sy_price = history(sy_future, fields='price', start_date='2014-01-01', end_date='2017-01-01')
crush = sy_price - (sm_price + bo_price)
crush.plot()
plt.ylabel('Crush Spread');
```
In the above plot, we offset the refined products by one month to roughly match the time it takes to crush the soybeans and we set `roll='calendar'` so that all three contracts are rolled at the same time.
To test whether this spread is stationary, we will use the augmented Dickey-Fuller test.
```
print 'p-value for stationarity: ', adfuller(crush)[1]
```
The test confirms that the spread is stationary. And it makes sense, economically, that the crush spread may exhibit some mean reversion due to simple supply and demand.
Note that there is usually a little more finesse required to obtain a mean reverting spread. We usually find a linear combination that would make the spread between the assets stationary after discovering cointegration. For more details on this, see the [lecture on cointegration](https://www.quantopian.com/lectures/integration-cointegration-and-stationarity). We skipped these steps to test this known spread off the bat.
Here are a few other examples of economically-linked futures:
* **3:2:1 Crack Spread**: Buy three crude oil, sell two gasoline, Sell one heating oil (this represents the profitability of oil refining)
* **8:4:3 Cattle Crush** Buy 8 October live-cattle, Sell 4 May feeder cattle, Sell 3 July corn (this represents the profitability of fattening feeder cattle, where the 3 corn contracts are enough to feed the young feeder cattle)
For widely-followed spreads like the crush spread or the crack spread, it would be surprising if the depth of mean reversion became so large that you could easily profit from it. If we consider futures that are linked to stocks, however, the number of potential pairs grows.
## Futures and Stocks
There are many examples of potential relationships between futures and stocks. We already discussed one of them - the relationship between the crush spread and the price of soybean processors. Here are several more, though this is not meant to be a complete list:
* Crude oil futures and oil stocks
* Gold futures and gold mining stocks
* Crude oil futures and airline stocks
* Currency futures and exporters
* Interest rate futures and utilities
* Interest rate futures and Real Estate Investment Trusts (REITs)
* Corn futures and agricultural processing companies (e.g., ADM)
Consider the relationship between ten-year interest rate futures and the price of EQR, a large REIT. Interest rates heavily influence the value of real estate, so there is a strong economic connection between the value of interest rate futures and the value of REITs.
```
ty_future = continuous_future('TY', offset=0, roll='calendar', adjustment='mul')
ty_prices = history(ty_future, fields='price', start_date='2009-01-01', end_date='2017-01-01')
ty_prices.name = ty_future.root_symbol
equities = symbols(['EQR', 'SPY'])
equity_prices = get_pricing(equities, fields='price', start_date='2009-01-01', end_date='2017-01-01')
equity_prices.columns = map(lambda x: x.symbol, equity_prices.columns)
data = pd.concat([ty_prices, equity_prices], axis=1)
data = data.dropna()
data.plot()
plt.legend();
```
If we apply a hypothesis test to the two price series we find that they are indeed cointegrated, corroborating our economic hypothesis.
```
print 'Cointegration test p-value: ', coint(data['TY'], data['EQR'])[1]
```
The next step would be to test if this signal is viable once we include market impact by trading EQR against the futures contract as a pair in a backtest.
Trading strategies based on cointegrated pairs form buy and sell signals based on the *relative prices* of the pair. We can also form trading signals based on *changes in prices*, or returns. Of course we would expect that changes in futures prices to be contemporaneously correlated with stock prices, which is not forecastable. If crude oil prices rise today, oil company stocks are likely to rise today also. But perhaps there are lead/lag effects also between changes in futures and stocks returns. We know there is evidence that the market can systematically underreact or overreact to other news releases, leading to trending and mean reversion. Let's look at a few examples with futures.
The first example looks at crude oil futures and oil company stocks.
```
cl_future = continuous_future('CL', offset=0, roll='calendar', adjustment='mul')
cl_prices = history(cl_future, fields='price', start_date='2007-01-01', end_date='2017-04-06')
cl_prices.name = cl_future.root_symbol
equities = symbols(['XOM', 'SPY'])
equity_prices = get_pricing(equities, fields='price', start_date='2007-01-01', end_date='2017-04-06')
equity_prices.columns = map(lambda x: x.symbol, equity_prices.columns)
data = pd.concat([cl_prices, equity_prices],axis=1)
data = data.dropna()
#Take log of prices
data['stock_ret'] = np.log(data['XOM']).diff()
data['spy_ret'] = np.log(data['SPY']).diff()
data['futures_ret'] = np.log(data['CL']).diff()
# Compute excess returns in excess of SPY
data['stock_excess'] = data['stock_ret'] - data['spy_ret']
#Compute lagged futures returns
data['futures_lag_diff'] = data['futures_ret'].shift(1)
data = data[2:].dropna()
data.tail(5)
```
We have a high positive contemporaneous correlation, but a slightly negative lagged correlation.
```
#Compute contemporaneous correlation
contemp_corr = data['stock_excess'].shift(1).corr(data['futures_lag_diff'])
#Compute correlation of excess stock returns with lagged futures returns
lagged_corr = data['stock_excess'].corr(data['futures_lag_diff'])
print 'Contemporaneous correlation: ', contemp_corr
print 'Lagged correlation : ', lagged_corr
```
And when we form a linear regression of the excess returns of XOM on the lagged futures returns, the coefficient is significant and negative. This and the above correlations indicate that there might be a slight overreaction to the shift in oil prices.
```
result = sm.OLS(data['stock_excess'], sm.add_constant(data['futures_lag_diff'])).fit()
result.summary2()
```
A coefficient of around $-0.02$ on the lagged futures return implies that if the oil price increased by 1% yesterday, the pure-play refiner is expected to go down by $2$ bp today. This would require more testing to formulate a functioning model, but it indicates that there might be some signal in drawing out the underreaction or overreaction of equity prices to changes in futures prices.
This conjecture could be total data mining, but perhaps when the connection between the futures and stock is exceedingly obvious, like oil stocks and oil exploration companies or gold stocks and gold miners, the market overreacts to fundamental information, but when the relationship is more subtle, the market underreacts.
Also, there may be other lead/lag effects over longer time scales than one-day, but as always, this could also lead to more data mining.
```
data['futures_lag_diff'].plot(alpha=0.50, legend=True)
data['stock_excess'].plot(alpha=0.50, legend=True);
```
*This presentation is for informational purposes only and does not constitute an offer to sell, a solicitation to buy, or a recommendation for any security; nor does it constitute an offer to provide investment advisory or other services by Quantopian, Inc. ("Quantopian"). Nothing contained herein constitutes investment advice or offers any opinion with respect to the suitability of any security, and any views expressed herein should not be taken as advice to buy, sell, or hold any security or as an endorsement of any security or company. In preparing the information contained herein, Quantopian, Inc. has not taken into account the investment needs, objectives, and financial circumstances of any particular investor. Any views expressed and data illustrated herein were prepared based upon information, believed to be reliable, available to Quantopian, Inc. at the time of publication. Quantopian makes no guarantees as to their accuracy or completeness. All information is subject to change and may quickly become unreliable for various reasons, including changes in market conditions or economic circumstances.*
| github_jupyter |
# Dijkstra's Algorithm
In this exercise, you'll implement Dijkstra's algorithm. First, let's build the graph.
## Graph Representation
In order to run Dijkstra's Algorithm, we'll need to add distance to each edge. We'll use the `GraphEdge` class below to represent each edge between a node.
```
class GraphEdge(object):
def __init__(self, node, distance):
self.node = node
self.distance = distance
```
The new graph representation should look like this:
```
class GraphNode(object):
def __init__(self, val):
self.value = val
self.edges = []
def add_child(self, node, distance):
self.edges.append(GraphEdge(node, distance))
def remove_child(self, del_node):
if del_node in self.edges:
self.edges.remove(del_node)
class Graph(object):
def __init__(self, node_list):
self.nodes = node_list
def add_edge(self, node1, node2, distance):
if node1 in self.nodes and node2 in self.nodes:
node1.add_child(node2, distance)
node2.add_child(node1, distance)
def remove_edge(self, node1, node2):
if node1 in self.nodes and node2 in self.nodes:
node1.remove_child(node2)
node2.remove_child(node1)
```
Now let's create the graph.
```
node_u = GraphNode('U')
node_d = GraphNode('D')
node_a = GraphNode('A')
node_c = GraphNode('C')
node_i = GraphNode('I')
node_t = GraphNode('T')
node_y = GraphNode('Y')
graph = Graph([node_u, node_d, node_a, node_c, node_i, node_t, node_y])
graph.add_edge(node_u, node_a, 4)
graph.add_edge(node_u, node_c, 6)
graph.add_edge(node_u, node_d, 3)
graph.add_edge(node_d, node_u, 3)
graph.add_edge(node_d, node_c, 4)
graph.add_edge(node_a, node_u, 4)
graph.add_edge(node_a, node_i, 7)
graph.add_edge(node_c, node_d, 4)
graph.add_edge(node_c, node_u, 6)
graph.add_edge(node_c, node_i, 4)
graph.add_edge(node_c, node_t, 5)
graph.add_edge(node_i, node_a, 7)
graph.add_edge(node_i, node_c, 4)
graph.add_edge(node_i, node_y, 4)
graph.add_edge(node_t, node_c, 5)
graph.add_edge(node_t, node_y, 5)
graph.add_edge(node_y, node_i, 4)
graph.add_edge(node_y, node_t, 5)
```
## Implementation
Using what you've learned, implement Dijkstra's Algorithm to find the shortest distance from the "U" node to the "Y" node.
```
import math
def dijkstra(start_node, end_node):
pass
print('Shortest Distance from {} to {} is {}'.format(node_u.value, node_y.value, dijkstra(node_u, node_y)))
```
<span class="graffiti-highlight graffiti-id_6vmf0hp-id_cjtybve"><i></i><button>Show Solution</button></span>
| github_jupyter |
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
cases = pd.read_csv('./ecdc/new_cases_per_million.csv')
tests = pd.read_csv('./testing/covid-testing-all-observations.csv')
inds = np.array(tests['Entity'].str.find('-'))
countstr = tests['Entity'].tolist()
countries = [countstr[i][:inds[i]-1] for i in range(len(inds))]
tests['Country']=countries
countries_unique = tests['Country'].unique()
erase = ['Iceland','Turkey','Tunisia','Australia','Austria','Bahrain','Ireland','Indonesia','Estonia','South Africa','United States','Uruguay','Vietnam']
list2 = []
for i in range(len(countries_unique)):
if countries_unique[i] not in erase:
list2.append(countries_unique[i])
countries_unique = np.array(list2)
dates = cases['date'].unique()
values = np.nan*np.zeros((len(dates),2, len(countries_unique)))
days = np.nan*np.zeros((len(dates),2, len(countries_unique)))
for d in range(len(dates)):
date = dates[d]
days[d,:,:] = d
cases_day = cases[cases['date']==date][countries_unique]
tests_day = tests[tests['Date']==date]['Daily change in cumulative total per thousand'].tolist()
tests_day_country = tests[tests['Date']==date]['Country'].tolist()
#print(tests_day_country)
for i in range(len(tests_day_country)):
count = np.where(countries_unique==tests_day_country[i])[0]
#print(count)
if(len(count)>0):
values[d,0,count[0]] = tests_day[i]
for i in range(len(countries_unique)):
values[d,1,i] = cases_day[countries_unique[i]]/1000
fig,ax = plt.subplots(4,7,figsize=(20,15))
ax=ax.flatten()
ini=60
maxy = np.nanmax(np.log(values[ini:,0,:]))
ll=np.log(values[ini:,1,:])[:]
ll=ll[ll>-np.inf]
miny = np.nanmin(ll)
for i in range(len(countries_unique)):
ax[i].plot(np.log(values[ini:,0,i]))
ax[i].plot(np.log(values[ini:,1,i]),'red')
ax[i].set_title(countries_unique[i],fontsize=25)
ax[i].set_ylim([miny,maxy])
if i>=21:
ax[i].set_xlabel('day',fontsize=15)
if i%7 ==0:
ax[i].set_ylabel('test per day/thousand',fontsize=15)
d = 90
fig,ax = plt.subplots(figsize=(10,10))
ax.scatter(values[d,0,:], values[d,1,:],100)
ax.set_title(dates[d],fontsize=20)
ax.set_xlabel('Tests today (per thousand)',fontsize=20)
ax.set_ylabel('Positives today (per thousand',fontsize=20)
for i, txt in enumerate(countries_unique):
ax.annotate(txt[:], (values[d,0,i], values[d,1,i]),size=15)
fig,ax = plt.subplots(1,2,figsize=(15,10))
#ax.set_yscale('log')
#ax[0].set_xscale('log')
ax[0].scatter(values[d,1,:], np.log(values[d,1,:]/values[d,0,:]),100)
ax[0].set_title(dates[d],fontsize=20)
ax[0].set_xlabel('Positives today (per thousand)',fontsize=20)
ax[0].set_ylabel('Positives/ Tests today ',fontsize=20)
for i, txt in enumerate(countries_unique):
ax[0].annotate(txt[:6], (values[d,1,i], np.log(values[d,1,i]/values[d,0,i])),size=15)
#ax[1].set_xscale('log')
ax[1].scatter(values[d,0,:], np.log(values[d,1,:]/values[d,0,:]),100)
ax[1].set_title(dates[d],fontsize=20)
ax[1].set_xlabel('Tests today (per thousand)',fontsize=20)
ax[1].set_ylabel('Positives/ Tests today ',fontsize=20)
for i, txt in enumerate(countries_unique):
ax[1].annotate(txt[:6], (values[d,0,i], np.log(values[d,1,i]/values[d,0,i])),size=15)
a=values[:,1,:]/values[:,0,:]
b=values[:,1,:]
c=values[:,0,:]
da = days[:,0,:]
ind=~np.isnan(a)
a=a[ind]
b=b[ind]
c=c[ind]
da=da[ind]
print(len(b))
thrs = np.linspace(0,100,10)
fig,ax=plt.subplots(3,len(thrs)-2,figsize=(20,5))
for i in range(0,len(thrs)-2):
#thr=thrs[i]
#print(a<np.percentile(a,thrs[i-1]))
#print([thrs[i],thrs[i+1]])
b1=b[(a>=np.percentile(a,thrs[i]))*(a<=np.percentile(a,thrs[i+1]))]
c1=c[(a>=np.percentile(a,thrs[i]))*(a<=np.percentile(a,thrs[i+1]))]
# print('b')
#print([np.min(b1),np.max(b1),np.mean(b1)])
#print('c')
#print([np.min(c1),np.max(c1),np.mean(c1)])
#print(len(b1))
#print(len(a))
#b2=b[a>thr]
#a=a[a<10000000000]
ax[0,i].hist(b1)
ax[1,i].hist(c1)
ax[2,i].scatter(b1,c1)
ax[2,i].set_xlim([0,0.1])
ax[2,i].set_ylim([0,1])
ax[2,i].set_xlabel('Positives')
ax[2,i].set_ylabel('Tests')
#ax[i].hist(b2)
#print(np.mean(b1))
#print(np.mean(b2))
N=50
prctile = 2
axb = np.linspace(np.min(b), np.max(b), N)
fit = np.zeros(N)
for i in range(len(axb)-1):
#print(i)
#print(np.percentile(b,100*(i+1)/N))
#print((b<=np.percentile(b,np.floor(100*(i+1)/N))))
inds = (b<=np.percentile(b,np.floor(100*(i+1)/N)))*(b>=np.percentile(b,np.floor(100*(i)/N)))
#print(inds)
logs=np.log(a[inds])
#logs = logs[logs>-np.inf]
#if(len(logs)==0):
# logs=-0
ll = np.percentile(logs,prctile)
fit[i] = ll
axb[i] = np.median(b[inds])
fig,ax=plt.subplots(1,figsize=(10,10))
im=ax.scatter(b,np.log(a),30,da)
fig.colorbar(im,ax=ax)
ax.plot(axb,fit,linewidth=3)
ax.set_xlabel('positives',fontsize=20)
ax.set_ylabel('log positives/tested',fontsize=20)
def f(x,axb,fit):
if(x<=axb[0]):
y = axb[0]
return y
if(x>=axb[-1]):
y = axb[-1]
return y
ind1 = np.where(axb<=x)
ind2 = np.where(axb>x)
return fit[ind1[0][-1]]/2+fit[ind2[0][0]]/2
N2 = 10000
xx = np.linspace(np.min(b), np.max(b), N2)
yy = np.array([f(xx[i],axb,fit) for i in range(len(xx))])
indgood = np.where(b>np.percentile(b,75))
inds=50
indsample = indgood[0][inds]
ax.scatter(b[indsample],np.log(a[indsample]),100,color='black')
#ax[0].plot(xx,yy,color='red',linewidth=2)
sample_b = b[indsample]
factors = np.linspace(0.01,10,100)
bnews = factors*sample_b
anews = bnews/c[indsample]
ff = np.array([f(axb[i],axb,fit) for i in range(len(axb))])
#ax[1].plot(axb,ff,linewidth=2)
#ax[1].plot(axb,fit,linewidth=2)
ax.plot(axb,np.log(a[indsample])+np.log(axb/sample_b),linewidth=2)#ax[1].plot(factors,np.log(factors)+np.log(sample_b/c[indsample]),linewidth=2)
ax.axvline(sample_b)
#ax[1].set_xscale('log')
#plt.plot(axb,fit)
plt.savefig('countries.png')
aaa=np.log(a[a<np.inf])
plt.hist(np.log(a[a<np.inf]))
```
| github_jupyter |
# Pipelining In Machine Learning
```
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split
# save filepath to variable for easier access
melbourne_file_path = '../hitchhikersGuideToMachineLearning/home-data-for-ml-course/train.csv'
# read the data and store data in DataFrame titled melbourne_data
train_data = pd.read_csv(melbourne_file_path)
# print a summary of the data in Melbourne data
train_data.head()
```
Instead of directly attacking the dataset I will cover the capabilities of Piplelining library from scikit-learn package!
- Pipelines and composite estimators
- Pipeline: chaining estimators
- Transforming target in regression
- FeatureUnion: composite feature spaces
- ColumnTransformer for heterogeneous data
Especially FetureUnion and ColumnTransformers are important.
Also I will demontrate how to incorporate custom techniques and tricks into piplines.
We will use few of this tricks on our dataset!
Data cleaning and preprocessing are a crucial step in the machine learning project.
Whenever new data points are added to the existing data, we need to perform the same preprocessing steps again before we can use the machine learning model to make predictions. This becomes a tedious and time-consuming process!
An alternate to this is creating a machine learning pipeline that remembers the complete set of preprocessing steps in the exact same order. So that whenever any new data point is introduced, the machine learning pipeline performs the steps as defined and uses the machine learning model to predict the target variable.
Setting up a machine learning algorithm involves more than the algorithm itself. You need to preprocess the data in order for it to fit the algorithm. It's this preprocessing pipeline that often requires a lot of work. Building a flexible pipeline is key. Here's how you can build it in python.
#### What is a pipeline?
A pipeline in sklearn is a set of chained algorithms to extract features, preprocess them and then train or use a machine learning algorith
```
from sklearn.datasets import load_iris
from sklearn.preprocessing import MinMaxScaler
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
from sklearn.pipeline import Pipeline
# Load and split the data
iris = load_iris()
X_train, X_test, y_train, y_test = train_test_split(iris.data, iris.target, test_size= 0.2,random_state=42 )
X_train.shape
```
#### Construction
The Pipeline is built using a list of (key, value) pairs, where the key is a string containing the name you want to give this step and value is an estimator object:
```
estimators = [('minmax', MinMaxScaler()),('lr', LogisticRegression(C=1))]
pipe = Pipeline(estimators)
pipe.fit(X_train, y_train)
score = pipe.score(X_test, y_test)
print('Logistic Regression pipeline test accuracy: %.3f' % score)
```
The utility function make_pipeline is a shorthand for constructing pipelines; it takes a variable number of estimators and returns a pipeline, filling in the names automatically:
```
from sklearn.pipeline import make_pipeline
pipe2= make_pipeline(MinMaxScaler(), LogisticRegression(C=10))
pipe2
```
###### Accessing steps
The estimators of a pipeline are stored as a list in the steps attribute, but can be accessed by index or name by indexing (with [idx]) the Pipeline:
```
pipe.steps[0]
pipe['minmax']
```
Pipeline’s named_steps attribute allows accessing steps by name with tab completion in interactive environments:
```
pipe.named_steps.minmax
```
A sub-pipeline can also be extracted using the slicing notation commonly used for Python Sequences such as lists or strings (although only a step of 1 is permitted). This is convenient for performing only some of the transformations (or their inverse):
```
pipe2[:1]
```
###### Nested parameters
Parameters of the estimators in the pipeline can be accessed using the <estimator>__<parameter> syntax:
```
pipe.set_params(lr__C=2)
```
This is particularly important for doing grid searches!
Individual steps may also be replaced as parameters, and non-final steps may be ignored by setting them to 'passthrough'
```
from sklearn.model_selection import GridSearchCV
import warnings
warnings.filterwarnings='ignore'
param_grid = dict(minmax=['passthrough'],lr__C=[1, 2, 3])
grid_search = GridSearchCV(pipe, param_grid=param_grid)
gd=grid_search.fit(X_train, y_train)
gd.best_estimator_
score = pipe.score(X_test, y_test)
score
```
Let's also explore a regression dataset so that i can drill a few more useful points in your skull!
```
from sklearn.datasets import load_boston
from sklearn.compose import TransformedTargetRegressor
from sklearn.preprocessing import QuantileTransformer
from sklearn.linear_model import LinearRegression
X, y = load_boston(return_X_y=True)
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0)
estimators = [('minmax', MinMaxScaler()),('lr', LinearRegression())]
raw_Y_regr = Pipeline(estimators)
raw_Y_regr.fit(X_train, y_train)
print('R2 score: {0:.2f}'.format(raw_Y_regr.score(X_test, y_test)))
```
Okay I can transform the input features and do all kinds of preprocessing , but what if i want to transform
Y. In many regression tasks you may have to transform Y while feeding the input to the model.For example taing log of Y.
But when we are predicting we ill have to scale it back to original unit basically we will have apply inverse of the transformation on predicted Y.
All this can be done in PIpeline very easily!
```
transformer = QuantileTransformer(output_distribution='normal')
regressor = LinearRegression()
regr = TransformedTargetRegressor(regressor=regressor,transformer=transformer)
regr.fit(X_train, y_train)
print('R2 score: {0:.2f}'.format(regr.score(X_test, y_test)))
```
For simple transformations, instead of a Transformer object, a pair of functions can be passed, defining the transformation and its inverse mapping. It means you can make your custom transformers!
```
def func(x):
return np.log(x)
def inverse_func(x):
return np.exp(x)
regr = TransformedTargetRegressor(regressor=regressor,func=func,inverse_func=inverse_func)
regr.fit(X_train, y_train)
print('R2 score: {0:.2f}'.format(regr.score(X_test, y_test)))
```
###### FeaturUnions
This estimator applies a list of transformer objects in parallel to the input data, then concatenates the results. This is useful to combine several feature extraction mechanisms into a single transformer.
```
from sklearn.pipeline import FeatureUnion
from sklearn.decomposition import PCA
from sklearn.decomposition import KernelPCA
from sklearn.feature_selection import SelectKBest
estimators = [('linear_pca', PCA()), ('select_k_best', SelectKBest(k=10))]
combined_features = FeatureUnion(estimators)
```
From PCA I am selecting 8 and from select K best I am slecting 7
```
X.shape[1]
# Use combined features to transform dataset:
X_features = combined_features.fit(X, y).transform(X)
print("Combined space has", X_features.shape[1], "features")
pipeline = Pipeline([("features", combined_features), ("lr", LinearRegression())])
param_grid = dict(features__linear_pca__n_components=[4, 6],
features__select_k_best__k=[5, 8])
grid_search = GridSearchCV(pipeline, param_grid=param_grid, verbose=10)
grid_search.fit(X, y)
print(grid_search.best_estimator_)
```
###### ColumnTransformer for heterogeneous data
Many datasets contain features of different types, say text, floats, and dates, where each type of feature requires separate preprocessing or feature extraction steps.
```
from sklearn.compose import ColumnTransformer
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.preprocessing import OneHotEncoder
from sklearn.preprocessing import LabelEncoder
from sklearn.impute import SimpleImputer
```
Lets use subset of our data to illustrate few more points.
```
X=train_data.iloc[:,0:10].drop(['Alley'],axis=1)
X
X.columns
```
For this data, we might want to encode the 'street' column as a categorical variable using preprocessing.
As we might use multiple feature extraction methods on the same column, we give each transformer a unique name, say 'street_category'.
By default, the remaining rating columns are ignored (remainder='drop').
We can keep the remaining rating columns by setting remainder='passthrough' also the remainder parameter can be set to an estimator to transform the remaining rating columns.
```
column_trans = ColumnTransformer(
[('street_category', OneHotEncoder(dtype='int'),['Street'])],
remainder='drop')
column_trans.fit(X)
column_trans.get_feature_names()
```
The make_column_selector is used to select columns based on data type or column name. Lets use OneHotEncoder on categorical data
```
from sklearn.preprocessing import StandardScaler
from sklearn.compose import make_column_selector
ct = ColumnTransformer([
('scale', StandardScaler(),
make_column_selector(dtype_include=np.number)),
('ohe',OneHotEncoder(),
make_column_selector(pattern='Street', dtype_include=object))])
ct.fit_transform(X)
```
If you will use LabelEncoder() in place of OneHotEncoder() you will run into this error
fit_transform() takes 2 positional arguments but 3 were given
Read here for workaround
https://stackoverflow.com/questions/46162855/fit-transform-takes-2-positional-arguments-but-3-were-given-with-labelbinarize
Didn't I told you that that we can use custom functionalities! Let's Change the labelencoder is implemented so that we can use it in our pipeline!
```
from sklearn.base import BaseEstimator, TransformerMixin
```
If you want to add some custom functionallity to your pipeline it will be basically two things either you will want to do some transformation which is not present in sklearn(or present but not in suitable format) or some estimator!
You should have obsereved by now that we have to make a object of every transformer or estimator before calling functions over it.For example
>lr=LinearRegression()
>lr.fit(data)
Pipelines also work in the same way so we will need to implememnt our functionallities as Classes!But you dont have to do everything from scratch Scikit-Learn got you covered.
Writing custom functionallity in SKlearn depends upon the inheritence of two classes:
- class sklearn.base.TransformerMixi:
Mixin class for all transformers in scikit-learn.This is the base class for writting all kinds of transformation you want ! That is all other classes will derive it as a parent class.
- class sklearn.base.BaseEstimator:
Base class for all estimators in scikit-learn and it for writting estimators!
So this is your custom functionality
```
class MultiColumnLabelEncoder(BaseEstimator,TransformerMixin):
def __init__(self, columns = None):
self.columns = columns # list of column to encode
def fit(self, X, y=None):
return self
def transform(self, X):
'''
Transforms columns of X specified in self.columns using
LabelEncoder(). If no columns specified, transforms all
columns in X.
'''
output = X.copy()
if self.columns is not None:
for col in self.columns:
output[col] = LabelEncoder().fit_transform(output[col])
else:
for colname, col in output.iteritems():
output[colname] = LabelEncoder().fit_transform(col)
return output
def fit_transform(self, X, y=None):
return self.fit(X, y).transform(X)
sk_pipe = Pipeline([ ("mssing", SimpleImputer(Strategy='most_frequent'))
,("MLCLE", MultiColumnLabelEncoder()),
("lr", LinearRegression())])
sk_pipe.fit(X,y)
```
One can also exploit featureUnion
scikit created a FunctionTransformer as part of the preprocessing class. It can be used in a similar manner as above but with less flexibility. If the input/output of the function is configured properly, the transformer can implement the fit/transform/fit_transform methods for the function and thus allow it to be used in the scikit pipeline.
For example, if the input to a pipeline is a series, the transformer would be as follows:
```
def trans_func(input_series):
return output_series
from sklearn.preprocessing import FunctionTransformer
name_transformer = FunctionTransformer(trans_func)
sk_pipe = Pipeline([("trans", name_transformer), ("lr", LinearRegression())])
sk_pipe
```
Lets put this all concept together and create simple pipleine!
before jumping to pipelines a few thigs still needed to be taken careof:
```
train_data = pd.read_csv('../hitchhikersGuideToMachineLearning/home-data-for-ml-course/train.csv' , index_col ='Id')
X_test_full = pd.read_csv('../hitchhikersGuideToMachineLearning/home-data-for-ml-course/test.csv', index_col='Id')
X_test_full['Neighborhood'].unique()
train_data['Neighborhood']
train_data.dropna(axis=0, subset=['SalePrice'], inplace=True)
y = train_data.SalePrice
train_data.drop(['SalePrice'], axis=1, inplace=True)
# Break off validation set from training data
X_train_full, X_valid_full, y_train, y_valid = train_test_split(train_data, y, train_size=0.8, test_size=0.2,
random_state=0)
# "Cardinality" means the number of unique values in a column
# Select categorical columns with relatively low cardinality (convenient but arbitrary) for one hot encoding
categorical_cols_type1 = [cname for cname in X_train_full.columns if
X_train_full[cname].nunique() <= 10 and
X_train_full[cname].dtype == "object"]
# Select categorical columns with high cardinality (convenient but arbitrary) for one hot encoding
categorical_cols_type2 = [cname for cname in X_train_full.columns if
X_train_full[cname].nunique() > 10 and
X_train_full[cname].dtype == "object"]
# Select numerical columns
numerical_cols = [cname for cname in X_train_full.columns if
X_train_full[cname].dtype in ['int64', 'float64']]
# Keep selected columns only
my_cols = categorical_cols_type1+categorical_cols_type2 + numerical_cols
X_train = X_train_full[my_cols].copy()
X_valid = X_valid_full[my_cols].copy()
X_test = X_test_full[my_cols].copy()
class LabelOneHotEncoder():
def __init__(self):
self.ohe = OneHotEncoder()
self.le = LabelEncoder()
def fit_transform(self, x):
features = self.le.fit_transform( x)
return self.ohe.fit_transform( features.reshape(-1,1))
def transform( self, x):
return self.ohe.transform( self.la.transform( x.reshape(-1,1)))
def inverse_tranform( self, x):
return self.le.inverse_transform( self.ohe.inverse_tranform( x))
def inverse_labels( self, x):
return self.le.inverse_transform( x)
class ModifiedLabelEncoder(LabelEncoder):
def fit_transform(self, y, *args, **kwargs):
return super().fit_transform(y).reshape(-1, 1)
def transform(self, y, *args, **kwargs):
return super().transform(y).reshape(-1, 1)
pipe = Pipeline([("le", ModifiedLabelEncoder()), ("ohe", OneHotEncoder())])
pipe.fit_transform(['dog', 'cat', 'dog'])
pipe.fit_transform(X_train["Street"])
from sklearn.compose import ColumnTransformer
from sklearn.pipeline import Pipeline
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import OneHotEncoder
from sklearn.ensemble import RandomForestRegressor
from sklearn.metrics import mean_absolute_error
# Preprocessing for numerical data
numerical_transformer = SimpleImputer(strategy='constant')
# Preprocessing for categorical data
categorical_transformer = Pipeline(steps=[
('imputer', SimpleImputer(strategy='most_frequent')),
('onehot', OneHotEncoder(handle_unknown='ignore'))
])
# Bundle preprocessing for numerical and categorical data
preprocessor = ColumnTransformer(
transformers=[
('num', numerical_transformer, numerical_cols),
('cat', categorical_transformer, categorical_cols)
])
# Define model
model = RandomForestRegressor(n_estimators=100, random_state=0)
# Bundle preprocessing and modeling code in a pipeline
clf = Pipeline(steps=[('preprocessor', preprocessor),
('model', model)
])
# Preprocessing of training data, fit model
clf.fit(X_train, y_train)
# Preprocessing of validation data, get predictions
preds = clf.predict(X_valid)
print('MAE:', mean_absolute_error(y_valid, preds))
# Preprocessing for numerical data
numerical_transformer1 = SimpleImputer(strategy='constant') # Your code here
# Preprocessing for categorical data
categorical_transformer1 = Pipeline(steps=[
('imputer', SimpleImputer(strategy='most_frequent')),
('onehot', OneHotEncoder(handle_unknown='ignore'))
]) # Your code here
# Bundle preprocessing for numerical and categorical data
preprocessor1 = ColumnTransformer(
transformers=[
('num', numerical_transformer1, numerical_cols),
('cat', categorical_transformer1, categorical_cols)
])
# Define model
model1 = RandomForestRegressor(n_estimators=150, random_state=0)
# Your code here
# Check your answer
step_1.a.check()
# Bundle preprocessing and modeling code in a pipeline
my_pipeline = Pipeline(steps=[('preprocessor', preprocessor1),
('model', model1)
])
# Preprocessing of training data, fit model
my_pipeline.fit(X_train, y_train)
# Preprocessing of validation data, get predictions
preds = my_pipeline.predict(X_valid)
# Evaluate the model
score = mean_absolute_error(y_valid, preds)
print('MAE:', score)
# Check your answer
step_1.b.check()
```
https://www.datanami.com/2018/09/05/how-to-build-a-better-machine-learning-pipeline/
https://www.analyticsvidhya.com/blog/2020/01/build-your-first-machine-learning-pipeline-using-scikit-learn/
https://cloud.google.com/ai-platform/prediction/docs/custom-pipeline
https://stackoverflow.com/questions/31259891/put-customized-functions-in-sklearn-pipeline
http://zacstewart.com/2014/08/05/pipelines-of-featureunions-of-pipelines.html
https://g-stat.com/using-custom-transformers-in-your-machine-learning-pipelines-with-scikit-learn/
| github_jupyter |
```
%matplotlib inline
!pip install annoy
!pip install nmslib
```
# Approximate nearest neighbors in TSNE
This example presents how to chain KNeighborsTransformer and TSNE in a
pipeline. It also shows how to wrap the packages `annoy` and `nmslib` to
replace KNeighborsTransformer and perform approximate nearest neighbors.
These packages can be installed with `pip install annoy nmslib`.
Note: In KNeighborsTransformer we use the definition which includes each
training point as its own neighbor in the count of `n_neighbors`, and for
compatibility reasons, one extra neighbor is computed when
`mode == 'distance'`. Please note that we do the same in the proposed wrappers.
Sample output::
Benchmarking on MNIST_2000:
---------------------------
AnnoyTransformer: 0.583 sec
NMSlibTransformer: 0.321 sec
KNeighborsTransformer: 1.225 sec
TSNE with AnnoyTransformer: 4.903 sec
TSNE with NMSlibTransformer: 5.009 sec
TSNE with KNeighborsTransformer: 6.210 sec
TSNE with internal NearestNeighbors: 6.365 sec
Benchmarking on MNIST_10000:
----------------------------
AnnoyTransformer: 4.457 sec
NMSlibTransformer: 2.080 sec
KNeighborsTransformer: 30.680 sec
TSNE with AnnoyTransformer: 30.225 sec
TSNE with NMSlibTransformer: 43.295 sec
TSNE with KNeighborsTransformer: 64.845 sec
TSNE with internal NearestNeighbors: 64.984 sec
```
# Author: Tom Dupre la Tour
#
# License: BSD 3 clause
import time
import sys
try:
import annoy
except ImportError:
print("The package 'annoy' is required to run this example.")
sys.exit()
try:
import nmslib
except ImportError:
print("The package 'nmslib' is required to run this example.")
sys.exit()
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.ticker import NullFormatter
from scipy.sparse import csr_matrix
from sklearn.base import BaseEstimator, TransformerMixin
from sklearn.neighbors import KNeighborsTransformer
from sklearn.utils._testing import assert_array_almost_equal
from sklearn.datasets import fetch_openml
from sklearn.pipeline import make_pipeline
from sklearn.manifold import TSNE
from sklearn.utils import shuffle
print(__doc__)
class NMSlibTransformer(TransformerMixin, BaseEstimator):
"""Wrapper for using nmslib as sklearn's KNeighborsTransformer"""
def __init__(self, n_neighbors=5, metric='euclidean', method='sw-graph',
n_jobs=1):
self.n_neighbors = n_neighbors
self.method = method
self.metric = metric
self.n_jobs = n_jobs
def fit(self, X):
self.n_samples_fit_ = X.shape[0]
# see more metric in the manual
# https://github.com/nmslib/nmslib/tree/master/manual
space = {
'euclidean': 'l2',
'cosine': 'cosinesimil',
'l1': 'l1',
'l2': 'l2',
}[self.metric]
self.nmslib_ = nmslib.init(method=self.method, space=space)
self.nmslib_.addDataPointBatch(X)
self.nmslib_.createIndex()
return self
def transform(self, X):
n_samples_transform = X.shape[0]
# For compatibility reasons, as each sample is considered as its own
# neighbor, one extra neighbor will be computed.
n_neighbors = self.n_neighbors + 1
results = self.nmslib_.knnQueryBatch(X, k=n_neighbors,
num_threads=self.n_jobs)
indices, distances = zip(*results)
indices, distances = np.vstack(indices), np.vstack(distances)
indptr = np.arange(0, n_samples_transform * n_neighbors + 1,
n_neighbors)
kneighbors_graph = csr_matrix((distances.ravel(), indices.ravel(),
indptr), shape=(n_samples_transform,
self.n_samples_fit_))
return kneighbors_graph
class AnnoyTransformer(TransformerMixin, BaseEstimator):
"""Wrapper for using annoy.AnnoyIndex as sklearn's KNeighborsTransformer"""
def __init__(self, n_neighbors=5, metric='euclidean', n_trees=10,
search_k=-1):
self.n_neighbors = n_neighbors
self.n_trees = n_trees
self.search_k = search_k
self.metric = metric
def fit(self, X):
self.n_samples_fit_ = X.shape[0]
self.annoy_ = annoy.AnnoyIndex(X.shape[1], metric=self.metric)
for i, x in enumerate(X):
self.annoy_.add_item(i, x.tolist())
self.annoy_.build(self.n_trees)
return self
def transform(self, X):
return self._transform(X)
def fit_transform(self, X, y=None):
return self.fit(X)._transform(X=None)
def _transform(self, X):
"""As `transform`, but handles X is None for faster `fit_transform`."""
n_samples_transform = self.n_samples_fit_ if X is None else X.shape[0]
# For compatibility reasons, as each sample is considered as its own
# neighbor, one extra neighbor will be computed.
n_neighbors = self.n_neighbors + 1
indices = np.empty((n_samples_transform, n_neighbors),
dtype=int)
distances = np.empty((n_samples_transform, n_neighbors))
if X is None:
for i in range(self.annoy_.get_n_items()):
ind, dist = self.annoy_.get_nns_by_item(
i, n_neighbors, self.search_k, include_distances=True)
indices[i], distances[i] = ind, dist
else:
for i, x in enumerate(X):
indices[i], distances[i] = self.annoy_.get_nns_by_vector(
x.tolist(), n_neighbors, self.search_k,
include_distances=True)
indptr = np.arange(0, n_samples_transform * n_neighbors + 1,
n_neighbors)
kneighbors_graph = csr_matrix((distances.ravel(), indices.ravel(),
indptr), shape=(n_samples_transform,
self.n_samples_fit_))
return kneighbors_graph
def test_transformers():
"""Test that AnnoyTransformer and KNeighborsTransformer give same results
"""
X = np.random.RandomState(42).randn(10, 2)
knn = KNeighborsTransformer()
Xt0 = knn.fit_transform(X)
ann = AnnoyTransformer()
Xt1 = ann.fit_transform(X)
nms = NMSlibTransformer()
Xt2 = nms.fit_transform(X)
assert_array_almost_equal(Xt0.toarray(), Xt1.toarray(), decimal=5)
assert_array_almost_equal(Xt0.toarray(), Xt2.toarray(), decimal=5)
def load_mnist(n_samples):
"""Load MNIST, shuffle the data, and return only n_samples."""
mnist = fetch_openml("mnist_784", as_frame=False)
X, y = shuffle(mnist.data, mnist.target, random_state=2)
return X[:n_samples] / 255, y[:n_samples]
def run_benchmark():
datasets = [
('MNIST_2000', load_mnist(n_samples=2000)),
('MNIST_10000', load_mnist(n_samples=10000)),
]
n_iter = 500
perplexity = 30
metric = "euclidean"
# TSNE requires a certain number of neighbors which depends on the
# perplexity parameter.
# Add one since we include each sample as its own neighbor.
n_neighbors = int(3. * perplexity + 1) + 1
tsne_params = dict(perplexity=perplexity, method="barnes_hut",
random_state=42, n_iter=n_iter,
square_distances=True)
transformers = [
('AnnoyTransformer',
AnnoyTransformer(n_neighbors=n_neighbors, metric=metric)),
('NMSlibTransformer',
NMSlibTransformer(n_neighbors=n_neighbors, metric=metric)),
('KNeighborsTransformer',
KNeighborsTransformer(n_neighbors=n_neighbors, mode='distance',
metric=metric)),
('TSNE with AnnoyTransformer',
make_pipeline(
AnnoyTransformer(n_neighbors=n_neighbors, metric=metric),
TSNE(metric='precomputed', **tsne_params))),
('TSNE with NMSlibTransformer',
make_pipeline(
NMSlibTransformer(n_neighbors=n_neighbors, metric=metric),
TSNE(metric='precomputed', **tsne_params))),
('TSNE with KNeighborsTransformer',
make_pipeline(
KNeighborsTransformer(n_neighbors=n_neighbors, mode='distance',
metric=metric),
TSNE(metric='precomputed', **tsne_params))),
('TSNE with internal NearestNeighbors',
TSNE(metric=metric, **tsne_params)),
]
# init the plot
nrows = len(datasets)
ncols = np.sum([1 for name, model in transformers if 'TSNE' in name])
fig, axes = plt.subplots(nrows=nrows, ncols=ncols, squeeze=False,
figsize=(5 * ncols, 4 * nrows))
axes = axes.ravel()
i_ax = 0
for dataset_name, (X, y) in datasets:
msg = 'Benchmarking on %s:' % dataset_name
print('\n%s\n%s' % (msg, '-' * len(msg)))
for transformer_name, transformer in transformers:
start = time.time()
Xt = transformer.fit_transform(X)
duration = time.time() - start
# print the duration report
longest = np.max([len(name) for name, model in transformers])
whitespaces = ' ' * (longest - len(transformer_name))
print('%s: %s%.3f sec' % (transformer_name, whitespaces, duration))
# plot TSNE embedding which should be very similar across methods
if 'TSNE' in transformer_name:
axes[i_ax].set_title(transformer_name + '\non ' + dataset_name)
axes[i_ax].scatter(Xt[:, 0], Xt[:, 1], c=y.astype(np.int32),
alpha=0.2, cmap=plt.cm.viridis)
axes[i_ax].xaxis.set_major_formatter(NullFormatter())
axes[i_ax].yaxis.set_major_formatter(NullFormatter())
axes[i_ax].axis('tight')
i_ax += 1
fig.tight_layout()
plt.show()
if __name__ == '__main__':
test_transformers()
run_benchmark()
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.