code stringlengths 2.5k 150k | kind stringclasses 1 value |
|---|---|
# SVM Classification Using Individual Replicas
This notebook analyzes the quality of the classifiers resulting from training on individual replicas of read counts rather than averaged values. Data are adjusted for library size and gene length.
Training data
1. Uses individual replicas (not averaged)
1. Uses all genes
1. Includes time T1 (normoxia is not combined with resuscitation)
```
import init
from common import constants as cn
from common.trinary_data import TrinaryData
from common.data_provider import DataProvider
from common_python.plots import util_plots
from plots import util_plots as xutil_plots
from common_python.classifier import classifier_ensemble
from common_python.classifier import util_classifier
from common_python.classifier import classifier_collection
from common_python.classifier.classifier_ensemble_random_forest import ClassifierEnsembleRandomForest
from common_python.plots import util_plots as common_plots
import collections
import os
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import sklearn
from sklearn.decomposition import PCA
from sklearn import svm
from sklearn.model_selection import cross_val_score
%matplotlib inline
```
## Analyze Replica Data
The following shows the extent to which replicas agree with the tranary values that are assigned.
```
def compareDFValues(df1, df2, title):
RANGE = [-8, 8]
plt.figure()
arr1 = df1.values.flatten()
arr2 = df2.values.flatten()
plt.scatter(arr1, arr2)
# Define region of 0 values
plt.plot([-1, -1], [-1, 1], color="b")
plt.plot([1, 1], [-1, 1], color="b")
plt.plot([-1, 1], [-1, -1], color="b")
plt.plot([-1, 1], [1, 1], color="b")
# Define region of 1 values
plt.plot([1, 1], [1, RANGE[1]], color="b")
plt.plot([1, RANGE[1]], [1, 1], color="b")
# Define region of -1 values
plt.plot([-1, -1], [-1, RANGE[0]], color="b")
plt.plot([-1, RANGE[0]], [-1, -1], color="b")
plt.plot(RANGE, RANGE, color="r")
plt.title(title)
provider = DataProvider()
provider.do()
dfs = []
for idx in range(3):
dfs.append(provider.dfs_adjusted_read_count_wrtT0_log2[idx])
compareDFValues(dfs[0], dfs[1], "0 vs 1")
compareDFValues(dfs[0], dfs[2], "0 vs 2")
compareDFValues(dfs[1], dfs[2], "1 vs 2")
dfs[0].values.flatten()
```
## Accuracy With Replicas
Compares accuracies with replicas (is_averaged=False) vs. with replicas as the data.
```
def clfEval(is_averaged, high_rank=15, ensemble_size=50, is_randomize=False, num_iterations=10):
trinary = TrinaryData(is_averaged=is_averaged, is_dropT1=is_averaged)
df_X = trinary.df_X.copy()
df_X.columns = trinary.features
ser_y = trinary.ser_y.copy()
if is_randomize:
# Randomize the relationship between features and state
df_X = df_X.sample(frac=1)
ser_y = ser_y.sample(frac=1)
#
svm_ensemble = classifier_ensemble.ClassifierEnsemble(
classifier_ensemble.ClassifierDescriptorSVM(), size=ensemble_size,
filter_high_rank=high_rank)
return classifier_ensemble.ClassifierEnsemble.crossValidateByState(
svm_ensemble, df_X, ser_y, num_iterations)
clfEval(True, ensemble_size=50, is_randomize=True)
clfEval(False, ensemble_size=50, is_randomize=True)
clfEval(True)
clfEval(False)
```
## Analysis of Classifier Accuracy by State
```
# Plot values by state
def plotValuesByState(states, values, stds=None, ylabel="percent"):
if stds is None:
plt.bar(states, values)
else:
plt.bar(states, values, yerr=stds, alpha=0.5)
plt.xticks(rotation=45)
plt.xlabel("state")
plt.ylabel(ylabel)
# State statistics
def plotStateDistributions():
PERCENT = "percent"
VALUE = "value"
NAME = "name"
trinary = TrinaryData(is_averaged=False, is_dropT1=False)
df = pd.DataFrame(trinary.ser_y)
df[VALUE] = list(np.repeat(1, len(df)))
df_group = pd.DataFrame(df.groupby(NAME).count())
dct = {v: k for k, v in trinary.state_dict.items()}
df_group.index = [dct[s] for s in df_group.index]
df_group[PERCENT] = 100*df_group[VALUE] / len(df)
plotValuesByState(df_group.index, df_group[PERCENT])
plotStateDistributions()
# Classification accuracy by state
def stateClassificationAccuracy(state, num_iterations=10, is_averaged=False):
NUM_HOLDOUTS = 1
is_dropT1 = is_averaged
trinary = TrinaryData(is_averaged=is_averaged, is_dropT1=is_dropT1)
df_X = trinary.df_X.copy()
df_X.columns = trinary.features
ser_y = trinary.ser_y
results = []
for _ in range(num_iterations):
test_indices = []
ser_sample = ser_y[ser_y == state].sample(n=NUM_HOLDOUTS)
test_indices.extend(list(ser_sample.index))
train_indices = list(set(df_X.index).difference(test_indices))
svm_ensemble = classifier_ensemble.ClassifierEnsemble(
classifier_ensemble.ClassifierDescriptorSVM(), size=30,
filter_high_rank=1500,
classes=list(ser_y.values))
svm_ensemble.fit(df_X.loc[train_indices, :], ser_y.loc[train_indices])
results.append(svm_ensemble.score(df_X.loc[test_indices, :], ser_y[test_indices]))
return results
def plotStateAccuracies(is_averaged=True):
is_dropT1 = is_averaged
trinary = TrinaryData(is_averaged=is_averaged, is_dropT1=is_dropT1)
states = list(trinary.state_dict.values())
avgs = []
stds = []
for state in states:
values = stateClassificationAccuracy(state, is_averaged=is_averaged)
avgs.append(np.mean(values))
stds.append(np.std(values))
plotValuesByState(list(trinary.state_dict.keys()), avgs, stds=stds, ylabel="accuracy")
plotStateAccuracies(is_averaged=True)
plt.figure()
plotStateAccuracies(is_averaged=False)
if False:
is_averaged = False
df = df_confuse.applymap(lambda v: np.nan if v <= 0.2 else 1)
df.columns = [c - 0.5 for c in df.columns]
trinary = TrinaryData(is_averaged=is_averaged, is_dropT1=is_averaged)
states = trinary.ser_y.values
state_colors = ["grey", "orange", "green", "pink", "peru", "greenyellow"]
heatmap = plt.pcolor(df.T, cmap='jet')
#fig = heatmap.get_figure()
#axes = fig.get_axes()[0]
#yaxis = axes.get_yaxis()
#xv = [x + 0.5 for x in range(len(df.T.columns))]
#yv = [y + 0.5 for y in range(len(df.T))]
#plt.xticks(xv)
#plt.yticks(yv)
positions = [p - 0.5 for p in range(-1, len(states))]
labels = [str(int(c-.5)) if c >= 0 else "" for c in positions]
plt.yticks(positions, labels)
for idx, state in enumerate(states):
color = state_colors[state]
plt.scatter(idx, [-1], color=color)
#plt.colorbar(heatmap)
is_averaged = False
trinary = TrinaryData(is_averaged=is_averaged, is_dropT1=is_averaged)
svm_ensemble = classifier_ensemble.ClassifierEnsemble(
classifier_ensemble.ClassifierDescriptorSVM(), size=50,
filter_high_rank=None)
ser_pred = svm_ensemble.makeInstancePredictionDF(trinary.df_X, trinary.ser_y)
util_classifier.plotInstancePredictions(trinary.ser_y, ser_pred, is_plot=True)
```
| github_jupyter |
# Inference in Google Earth Engine + Colab
> Scaling up machine learning with GEE and Google Colab.
- toc: true
- badges: true
- author: Drew Bollinger
- comments: false
- hide: false
- sticky_rank: 11
# Inference in Google Earth Engine + Colab
Here we demonstrate how to take a trained model and apply to to imagery with Google Earth Engine + Colab + Tensorflow. This is adapted from an [Earth Engine <> TensorFlow demonstration notebook](https://developers.google.com/earth-engine/guides/tf_examples). We'll be taking the trained model from the [Deep Learning Crop Type Segmentation Model Example](https://developmentseed.org/sat-ml-training/DeepLearning_CropType_Segmentation).
# Setup software libraries
Authenticate and import as necessary.
```
# Import, authenticate and initialize the Earth Engine library.
import ee
ee.Authenticate()
ee.Initialize()
# Mount our Google Drive
from google.colab import drive
drive.mount('/content/drive')
# Add necessary libraries.
!pip install -q focal-loss
import os
from os import path as op
import tensorflow as tf
import folium
from focal_loss import SparseCategoricalFocalLoss
```
# Variables
Declare the variables that will be in use throughout the notebook.
```
# Specify names locations for outputs in Google Drive.
FOLDER = 'servir-inference-demo'
ROOT_DIR = '/content/drive/My Drive/'
# Specify inputs (Sentinel indexes) to the model.
BANDS = ['NDVI', 'WDRVI', 'SAVI']
# Specify the size and shape of patches expected by the model.
KERNEL_SIZE = 224
KERNEL_SHAPE = [KERNEL_SIZE, KERNEL_SIZE]
```
# Imagery
Gather and setup the imagery to use for inputs. It's important that we match the index inputs from the earlier analysis. This is a three-month Sentinel-2 composite. Display it in the notebook for a sanity check.
```
# Use Sentinel-2 data.
def add_indexes(img):
ndvi = img.expression(
'(nir - red) / (nir + red + a)', {
'a': 1e-5,
'nir': img.select('B8'),
'red': img.select('B4')
}
).rename('NDVI')
wdrvi = img.expression(
'(a * nir - red) / (a * nir + red)', {
'a': 0.2,
'nir': img.select('B8'),
'red': img.select('B4')
}
).rename('WDRVI')
savi = img.expression(
'1.5 * (nir - red) / (nir + red + 0.5)', {
'nir': img.select('B8'),
'red': img.select('B4')
}
).rename('SAVI')
return ee.Image.cat([ndvi, wdrvi, savi])
image = ee.ImageCollection('COPERNICUS/S2') \
.filterDate('2018-01-01', '2018-04-01') \
.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', 20)) \
.map(add_indexes) \
.median()
# Use folium to visualize the imagery.
mapid = image.getMapId({'bands': BANDS, 'min': -1, 'max': 1})
map = folium.Map(location=[
-29.177943749121233,
30.55984497070313,
])
folium.TileLayer(
tiles=mapid['tile_fetcher'].url_format,
attr='Map Data © <a href="https://earthengine.google.com/">Google Earth Engine</a>',
overlay=True,
name='median composite',
).add_to(map)
map.add_child(folium.LayerControl())
map
```
# Load our saved model
```
# Load a trained model.
MODEL_DIR = '/content/drive/Shared drives/servir-sat-ml/data/model_out/10062020/'
model = tf.keras.models.load_model(MODEL_DIR)
model.summary()
```
# Prediction
The prediction pipeline is:
1. Export imagery on which to do predictions from Earth Engine in TFRecord format to Google Drive.
2. Use the trained model to make the predictions.
3. Write the predictions to a TFRecord file in Google Drive.
4. Manually upload the predictions TFRecord file to Earth Engine.
The following functions handle this process. It's useful to separate the export from the predictions so that you can experiment with different models without running the export every time.
```
def doExport(out_image_base, shape, region):
"""Run the image export task. Block until complete.
"""
task = ee.batch.Export.image.toDrive(
image = image.select(BANDS),
description = out_image_base,
fileNamePrefix = out_image_base,
folder = FOLDER,
region = region.getInfo()['coordinates'],
scale = 30,
fileFormat = 'TFRecord',
maxPixels = 1e10,
formatOptions = {
'patchDimensions': shape,
'compressed': True,
'maxFileSize': 104857600
}
)
task.start()
# Block until the task completes.
print('Running image export to Google Drive...')
import time
while task.active():
time.sleep(30)
# Error condition
if task.status()['state'] != 'COMPLETED':
print('Error with image export.')
else:
print('Image export completed.')
def doPrediction(out_image_base, kernel_shape, region):
"""Perform inference on exported imagery.
"""
print('Looking for TFRecord files...')
# Get a list of all the files in the output bucket.
filesList = os.listdir(op.join(ROOT_DIR, FOLDER))
# Get only the files generated by the image export.
exportFilesList = [s for s in filesList if out_image_base in s]
# Get the list of image files and the JSON mixer file.
imageFilesList = []
jsonFile = None
for f in exportFilesList:
if f.endswith('.tfrecord.gz'):
imageFilesList.append(op.join(ROOT_DIR, FOLDER, f))
elif f.endswith('.json'):
jsonFile = f
# Make sure the files are in the right order.
imageFilesList.sort()
from pprint import pprint
pprint(imageFilesList)
print(jsonFile)
import json
# Load the contents of the mixer file to a JSON object.
with open(op.join(ROOT_DIR, FOLDER, jsonFile), 'r') as f:
mixer = json.load(f)
pprint(mixer)
patches = mixer['totalPatches']
# Get set up for prediction.
imageColumns = [
tf.io.FixedLenFeature(shape=kernel_shape, dtype=tf.float32)
for k in BANDS
]
imageFeaturesDict = dict(zip(BANDS, imageColumns))
def parse_image(example_proto):
return tf.io.parse_single_example(example_proto, imageFeaturesDict)
def toTupleImage(inputs):
inputsList = [inputs.get(key) for key in BANDS]
stacked = tf.stack(inputsList, axis=0)
stacked = tf.transpose(stacked, [1, 2, 0])
return stacked
# Create a dataset from the TFRecord file(s) in Cloud Storage.
imageDataset = tf.data.TFRecordDataset(imageFilesList, compression_type='GZIP')
imageDataset = imageDataset.map(parse_image, num_parallel_calls=5)
imageDataset = imageDataset.map(toTupleImage).batch(1)
# Perform inference.
print('Running predictions...')
predictions = model.predict(imageDataset, steps=patches, verbose=1)
# print(predictions[0])
print('Writing predictions...')
out_image_file = op.join(ROOT_DIR, FOLDER, f'{out_image_base}pred.TFRecord')
writer = tf.io.TFRecordWriter(out_image_file)
patches = 0
for predictionPatch in predictions:
print('Writing patch ' + str(patches) + '...')
predictionPatch = tf.argmax(predictionPatch, axis=2)
# Create an example.
example = tf.train.Example(
features=tf.train.Features(
feature={
'class': tf.train.Feature(
float_list=tf.train.FloatList(
value=predictionPatch.numpy().flatten()))
}
)
)
# Write the example.
writer.write(example.SerializeToString())
patches += 1
writer.close()
```
Now there's all the code needed to run the prediction pipeline, all that remains is to specify the output region in which to do the prediction, the names of the output files, where to put them, and the shape of the outputs.
```
# Base file name to use for TFRecord files and assets.
image_base = 'servir_inference_demo_'
# South Africa (near training data)
region = ee.Geometry.Polygon(
[[[
30.55984497070313,
-29.177943749121233
],
[
30.843429565429684,
-29.177943749121233
],
[
30.843429565429684,
-28.994928377910732
],
[
30.55984497070313,
-28.994928377910732
]]], None, False)
# Run the export.
doExport(image_base, KERNEL_SHAPE, region)
# Run the prediction.
doPrediction(image_base, KERNEL_SHAPE, region)
```
# Display the output
One the data has been exported, the model has made predictions and the predictions have been written to a file, we need to [manually import the TFRecord to Earth Engine](https://developers.google.com/earth-engine/guides/tfrecord#uploading-tfrecords-to-earth-engine). Then we can display our crop type predictions as an image asset
```
out_image = ee.Image('users/drew/servir_inference_demo_-mixer')
mapid = out_image.getMapId({'min': 0, 'max': 10, 'palette': ['00A600','63C600','E6E600','E9BD3A','ECB176','EFC2B3','F2F2F2']})
map = folium.Map(location=[
-29.177943749121233,
30.55984497070313,
])
folium.TileLayer(
tiles=mapid['tile_fetcher'].url_format,
attr='Map Data © <a href="https://earthengine.google.com/">Google Earth Engine</a>',
overlay=True,
name='predicted crop type',
).add_to(map)
map.add_child(folium.LayerControl())
map
```
| github_jupyter |
<a href="https://cognitiveclass.ai"><img src = "https://ibm.box.com/shared/static/9gegpsmnsoo25ikkbl4qzlvlyjbgxs5x.png" width = 400> </a>
<h1 align=center><font size = 5>From Understanding to Preparation</font></h1>
## Introduction
In this lab, we will continue learning about the data science methodology, and focus on the **Data Understanding** and the **Data Preparation** stages.
## Table of Contents
<div class="alert alert-block alert-info" style="margin-top: 20px">
1. [Recap](#0)<br>
2. [Data Understanding](#2)<br>
3. [Data Preparation](#4)<br>
</div>
<hr>
# Recap <a id="0"></a>
In Lab **From Requirements to Collection**, we learned that the data we need to answer the question developed in the business understanding stage, namely *can we automate the process of determining the cuisine of a given recipe?*, is readily available. A researcher named Yong-Yeol Ahn scraped tens of thousands of food recipes (cuisines and ingredients) from three different websites, namely:
<img src = "https://ibm.box.com/shared/static/4fruwan7wmjov3gywiz3swlojw0srv54.png" width=500>
www.allrecipes.com
<img src = "https://ibm.box.com/shared/static/cebfdbr22fjxa47lltp0bs533r103g0z.png" width=500>
www.epicurious.com
<img src = "https://ibm.box.com/shared/static/epk727njg7xrz49pbkpkzd05cm5ywqmu.png" width=500>
www.menupan.com
For more information on Yong-Yeol Ahn and his research, you can read his paper on [Flavor Network and the Principles of Food Pairing](http://yongyeol.com/papers/ahn-flavornet-2011.pdf).
We also collected the data and placed it on an IBM server for your convenience.
------------
# Data Understanding <a id="2"></a>
<img src="https://ibm.box.com/shared/static/89geb3m0ge1z73s92hl8o8wdcpcrggtz.png" width=500>
<strong> Important note:</strong> Please note that you are not expected to know how to program in python. The following code is meant to illustrate the stages of data understanding and data preparation, so it is totally fine if you do not understand the individual lines of code. We have a full course on programming in python, <a href="http://cocl.us/PY0101EN_DS0103EN_LAB3_PYTHON">Python for Data Science</a>, so please feel free to complete the course if you are interested in learning how to program in python.
### Using this notebook:
To run any of the following cells of code, you can type **Shift + Enter** to excute the code in a cell.
Get the version of Python installed.
```
# check Python version
!python -V
```
Download the library and dependencies that we will need to run this lab.
```
import pandas as pd # import library to read data into dataframe
pd.set_option('display.max_columns', None)
import numpy as np # import numpy library
import re # import library for regular expression
```
Download the data from the IBM server and read it into a *pandas* dataframe.
```
recipes = pd.read_csv("https://ibm.box.com/shared/static/5wah9atr5o1akuuavl2z9tkjzdinr1lv.csv")
print("Data read into dataframe!") # takes about 30 seconds
```
Show the first few rows.
```
recipes.head()
```
Get the dimensions of the dataframe.
```
recipes.shape
```
So our dataset consists of 57,691 recipes. Each row represents a recipe, and for each recipe, the corresponding cuisine is documented as well as whether 384 ingredients exist in the recipe or not, beginning with almond and ending with zucchini.
We know that a basic sushi recipe includes the ingredients:
* rice
* soy sauce
* wasabi
* some fish/vegetables
Let's check that these ingredients exist in our dataframe:
```
ingredients = list(recipes.columns.values)
print([match.group(0) for ingredient in ingredients for match in [(re.compile(".*(rice).*")).search(ingredient)] if match])
print([match.group(0) for ingredient in ingredients for match in [(re.compile(".*(wasabi).*")).search(ingredient)] if match])
print([match.group(0) for ingredient in ingredients for match in [(re.compile(".*(soy).*")).search(ingredient)] if match])
```
Yes, they do!
* rice exists as rice.
* wasabi exists as wasabi.
* soy exists as soy_sauce.
So maybe if a recipe contains all three ingredients: rice, wasabi, and soy_sauce, then we can confidently say that the recipe is a **Japanese** cuisine! Let's keep this in mind!
----------------
# Data Preparation <a id="4"></a>
<img src="https://ibm.box.com/shared/static/lqc2j3r0ndhokh77mubohwjqybzf8dhk.png" width=500>
In this section, we will prepare data for the next stage in the data science methodology, which is modeling. This stage involves exploring the data further and making sure that it is in the right format for the machine learning algorithm that we selected in the analytic approach stage, which is decision trees.
First, look at the data to see if it needs cleaning.
```
recipes["country"].value_counts() # frequency table
```
By looking at the above table, we can make the following observations:
1. Cuisine column is labeled as Country, which is inaccurate.
2. Cuisine names are not consistent as not all of them start with an uppercase first letter.
3. Some cuisines are duplicated as variation of the country name, such as Vietnam and Vietnamese.
4. Some cuisines have very few recipes.
#### Let's fixes these problems.
Fix the name of the column showing the cuisine.
```
column_names = recipes.columns.values
column_names[0] = "cuisine"
recipes.columns = column_names
recipes
```
Make all the cuisine names lowercase.
```
recipes["cuisine"] = recipes["cuisine"].str.lower()
```
Make the cuisine names consistent.
```
recipes.loc[recipes["cuisine"] == "austria", "cuisine"] = "austrian"
recipes.loc[recipes["cuisine"] == "belgium", "cuisine"] = "belgian"
recipes.loc[recipes["cuisine"] == "china", "cuisine"] = "chinese"
recipes.loc[recipes["cuisine"] == "canada", "cuisine"] = "canadian"
recipes.loc[recipes["cuisine"] == "netherlands", "cuisine"] = "dutch"
recipes.loc[recipes["cuisine"] == "france", "cuisine"] = "french"
recipes.loc[recipes["cuisine"] == "germany", "cuisine"] = "german"
recipes.loc[recipes["cuisine"] == "india", "cuisine"] = "indian"
recipes.loc[recipes["cuisine"] == "indonesia", "cuisine"] = "indonesian"
recipes.loc[recipes["cuisine"] == "iran", "cuisine"] = "iranian"
recipes.loc[recipes["cuisine"] == "italy", "cuisine"] = "italian"
recipes.loc[recipes["cuisine"] == "japan", "cuisine"] = "japanese"
recipes.loc[recipes["cuisine"] == "israel", "cuisine"] = "jewish"
recipes.loc[recipes["cuisine"] == "korea", "cuisine"] = "korean"
recipes.loc[recipes["cuisine"] == "lebanon", "cuisine"] = "lebanese"
recipes.loc[recipes["cuisine"] == "malaysia", "cuisine"] = "malaysian"
recipes.loc[recipes["cuisine"] == "mexico", "cuisine"] = "mexican"
recipes.loc[recipes["cuisine"] == "pakistan", "cuisine"] = "pakistani"
recipes.loc[recipes["cuisine"] == "philippines", "cuisine"] = "philippine"
recipes.loc[recipes["cuisine"] == "scandinavia", "cuisine"] = "scandinavian"
recipes.loc[recipes["cuisine"] == "spain", "cuisine"] = "spanish_portuguese"
recipes.loc[recipes["cuisine"] == "portugal", "cuisine"] = "spanish_portuguese"
recipes.loc[recipes["cuisine"] == "switzerland", "cuisine"] = "swiss"
recipes.loc[recipes["cuisine"] == "thailand", "cuisine"] = "thai"
recipes.loc[recipes["cuisine"] == "turkey", "cuisine"] = "turkish"
recipes.loc[recipes["cuisine"] == "vietnam", "cuisine"] = "vietnamese"
recipes.loc[recipes["cuisine"] == "uk-and-ireland", "cuisine"] = "uk-and-irish"
recipes.loc[recipes["cuisine"] == "irish", "cuisine"] = "uk-and-irish"
recipes
```
Remove cuisines with < 50 recipes.
```
# get list of cuisines to keep
recipes_counts = recipes["cuisine"].value_counts()
cuisines_indices = recipes_counts > 50
cuisines_to_keep = list(np.array(recipes_counts.index.values)[np.array(cuisines_indices)])
rows_before = recipes.shape[0] # number of rows of original dataframe
print("Number of rows of original dataframe is {}.".format(rows_before))
recipes = recipes.loc[recipes['cuisine'].isin(cuisines_to_keep)]
rows_after = recipes.shape[0] # number of rows of processed dataframe
print("Number of rows of processed dataframe is {}.".format(rows_after))
print("{} rows removed!".format(rows_before - rows_after))
```
Convert all Yes's to 1's and the No's to 0's
```
recipes = recipes.replace(to_replace="Yes", value=1)
recipes = recipes.replace(to_replace="No", value=0)
```
#### Let's analyze the data a little more in order to learn the data better and note any interesting preliminary observations.
Run the following cell to get the recipes that contain **rice** *and* **soy** *and* **wasabi** *and* **seaweed**.
```
recipes.head()
check_recipes = recipes.loc[
(recipes["rice"] == 1) &
(recipes["soy_sauce"] == 1) &
(recipes["wasabi"] == 1) &
(recipes["seaweed"] == 1)
]
check_recipes
```
Based on the results of the above code, can we classify all recipes that contain **rice** *and* **soy** *and* **wasabi** *and* **seaweed** as **Japanese** recipes? Why?
Double-click __here__ for the solution.
<!-- The correct answer is:
No, because other recipes such as Asian and East_Asian recipes also contain these ingredients.
-->
Let's count the ingredients across all recipes.
```
# sum each column
ing = recipes.iloc[:, 1:].sum(axis=0)
# define each column as a pandas series
ingredient = pd.Series(ing.index.values, index = np.arange(len(ing)))
count = pd.Series(list(ing), index = np.arange(len(ing)))
# create the dataframe
ing_df = pd.DataFrame(dict(ingredient = ingredient, count = count))
ing_df = ing_df[["ingredient", "count"]]
print(ing_df.to_string())
```
Now we have a dataframe of ingredients and their total counts across all recipes. Let's sort this dataframe in descending order.
```
ing_df.sort_values(["count"], ascending=False, inplace=True)
ing_df.reset_index(inplace=True, drop=True)
print(ing_df)
```
#### What are the 3 most popular ingredients?
Double-click __here__ for the solution.
<!-- The correct answer is:
// 1. Egg with <strong>21,025</strong> occurrences.
// 2. Wheat with <strong>20,781</strong> occurrences.
// 3. Butter with <strong>20,719</strong> occurrences.
-->
However, note that there is a problem with the above table. There are ~40,000 American recipes in our dataset, which means that the data is biased towards American ingredients.
**Therefore**, let's compute a more objective summary of the ingredients by looking at the ingredients per cuisine.
#### Let's create a *profile* for each cuisine.
In other words, let's try to find out what ingredients Chinese people typically use, and what is **Canadian** food for example.
```
cuisines = recipes.groupby("cuisine").mean()
cuisines.head()
```
As shown above, we have just created a dataframe where each row is a cuisine and each column (except for the first column) is an ingredient, and the row values represent the percentage of each ingredient in the corresponding cuisine.
**For example**:
* *almond* is present across 15.65% of all of the **African** recipes.
* *butter* is present across 38.11% of all of the **Canadian** recipes.
Let's print out the profile for each cuisine by displaying the top four ingredients in each cuisine.
```
num_ingredients = 4 # define number of top ingredients to print
# define a function that prints the top ingredients for each cuisine
def print_top_ingredients(row):
print(row.name.upper())
row_sorted = row.sort_values(ascending=False)*100
top_ingredients = list(row_sorted.index.values)[0:num_ingredients]
row_sorted = list(row_sorted)[0:num_ingredients]
for ind, ingredient in enumerate(top_ingredients):
print("%s (%d%%)" % (ingredient, row_sorted[ind]), end=' ')
print("\n")
# apply function to cuisines dataframe
create_cuisines_profiles = cuisines.apply(print_top_ingredients, axis=1)
```
At this point, we feel that we have understood the data well and the data is ready and is in the right format for modeling!
-----------
### Thank you for completing this lab!
This notebook was created by [Alex Aklson](https://www.linkedin.com/in/aklson/). We hope you found this lab session interesting. Feel free to contact us if you have any questions!
This notebook is part of the free course on **Cognitive Class** called *Data Science Methodology*. If you accessed this notebook outside the course, you can take this free self-paced course, online by clicking [here](https://cocl.us/DS0103EN_LAB3_PYTHON).
<hr>
Copyright © 2018 [Cognitive Class](https://cognitiveclass.ai/?utm_source=bducopyrightlink&utm_medium=dswb&utm_campaign=bdu). This notebook and its source code are released under the terms of the [MIT License](https://bigdatauniversity.com/mit-license/).
| github_jupyter |

```
import numpy as np
import keras
from keras.models import Sequential
from keras.layers import Dense
x_train = np.array([[150, 72],
[125, 55],
[115, 50],
[165, 75],
[180, 60],
[135, 56]])
y_train = np.array([1,0,0,1,1,0])
```
## Scale Data using z = (x - m) / s where m = mean, s = standard deviation
```
class scale:
def mean(self, x):
return x.sum(axis=0) / x_train.shape[0]
def std(self, x):
m = self.mean(x)
x_minus_mean = x - m
x_minus_mean2 = x_minus_mean ** 2
summation = x_minus_mean2.sum(axis=0)
divide_by_N = summation / (x_train.shape[0])
standard_deviation = np.sqrt(divide_by_N)
return standard_deviation
def scale_data(self, x):
return (x - self.mean(x)) / self.std(x)
scaler = scale()
print("MEAN = {}".format(scaler.mean(x_train)))
print("STD = {}".format(scaler.std(x_train)))
x_train = scaler.scale_data(x_train)
print(x_train)
#x_train shape = 11x4
# x_train = np.array([[1.2, 4.3, 2.1, 1.9],
# [6.2, 8.3, 5.1, 9.9],
# [2.3, 4.3, 3.1, 0.9],
# [4.1, 4.4, 1.1, 0.3],
# [6.1, 7.1, 8.1, 9.1],
# [1.0, 2.0, 1.0, 1.0],
# [5.1, 5.1, 5.1, 5.1],
# [1.8, 4.0, 3.9, 2.7],
# [4.4, 0.8, 1.9, 2.7],
# [6.9, 8.8, 5.7, 7.1]])
# #y_train shape = 1x11
# y_train = np.array([[1,0],
# [0,1],
# [1,0],
# [1,0],
# [0,1],
# [1,0],
# [0,1],
# [1,0],
# [1,0],
# [0,1]])
class tinyNN:
def __init__(self):
np.random.seed(1)
self.w1 = (2 * np.random.rand(2,50)) - 1
self.w2 = (2 * np.random.rand(50,1)) - 1
self.lr = .001
def tanh(self, x):
return (np.exp(x)-np.exp(-x)) / (np.exp(x)+np.exp(-x))
def derivTanh(self, x):
return (4*np.exp(2*x)) / ((np.exp(2*x) + 1)**2)
def sigmoid(self, x):
return 1 / (1+np.exp(-x))
def derivSigmoid(self, x):
return np.exp(x) / (np.exp(x) + 1)**2
def loss(self, y_pred, y_true):
return (y_pred - y_true)**2
def derivLoss(self, error):
return error
def forward(self, inputs):
self.hidden = np.dot(inputs, self.w1)
self.h1 = self.tanh(self.hidden)
outputs = np.dot(self.h1, self.w2)
return outputs
def calcError(self, y_pred, y):
return y_pred.T - y
def backProp(self, error):
dypred_dw2 = self.h1
dse_dypred = error
delta_w2 = np.dot(dypred_dw2.T, dse_dypred.T)
dse_dh = np.dot(dse_dypred.T, self.w2.T)
dh_dp = self.derivTanh(self.hidden)
dse_dp = dse_dh * dh_dp
delta_w1 = np.dot(x_train.T, dse_dp)
self.w2 -= self.lr * delta_w2
self.w1 -= self.lr * delta_w1
garvis = tinyNN()
for i in range(1000):
outputs = garvis.forward(x_train)
error = garvis.calcError(outputs, y_train)
garvis.backProp(error)
outputs = garvis.forward(x_train)
print(outputs)
outputs = np.where(outputs >= .5, 1, 0)
outputs = np.where(outputs == 1, "Male", "Female")
print(outputs)
```
| github_jupyter |
# <font color='blue'>Data Science Academy - Python Fundamentos - Capítulo 4</font>
## Download: http://github.com/dsacademybr
```
# Versão da Linguagem Python
from platform import python_version
print('Versão da Linguagem Python Usada Neste Jupyter Notebook:', python_version())
```
# Versão da Linguagem Python
from platform import python_version
print('Versão da Linguagem Python Usada Neste Jupyter Notebook:', python_version())
## Exercícios
```
# Exercício 1 - Crie uma lista de 3 elementos e calcule a terceira potência de cada elemento.
list1 = [3,4,5]
quadrado = [item**3 for item in list1]
print(quadrado)
# Exercício 2 - Reescreva o código abaixo, usando a função map(). O resultado final deve ser o mesmo!
palavras = 'A Data Science Academy oferce os melhores cursos de análise de dados do Brasil'.split()
resultado = [[w.upper(), w.lower(), len(w)] for w in palavras]
for i in resultado:
print (i)
resultado = map(lambda w: [w.upper(), w.lower(), len(w)], palavras)
for i in resultado:
print (i)
# Exercício 3 - Calcule a matriz transposta da matriz abaixo.
# Caso não saiba o que é matriz transposta, visite este link: https://pt.wikipedia.org/wiki/Matriz_transposta
# Matriz transposta é um conceito fundamental na construção de redes neurais artificiais, base de sistemas de IA.
matrix = [[1, 2],[3,4],[5,6],[7,8]]
transpose = [[row[i] for row in matrix] for i in range(2)]
print(transpose)
# Exercício 4 - Crie duas funções, uma para elevar um número ao quadrado e outra para elevar ao cubo.
# Aplique as duas funções aos elementos da lista abaixo.
# Obs: as duas funções devem ser aplicadas simultaneamente.
lista = [0, 1, 2, 3, 4]
def square(x):
return (x**2)
def cube(x):
return (x**3)
funcs = [square, cube]
for i in lista:
valor = map(lambda x: x(i), funcs)
print(list((valor)))
# Exercício 5 - Abaixo você encontra duas listas. Faça com que cada elemento da listaA seja elevado
# ao elemento correspondente na listaB.
listaA = [2, 3, 4]
listaB = [10, 11, 12]
list(map(pow, listaA, listaB))
# Exercício 6 - Considerando o range de valores abaixo, use a função filter() para retornar apenas os valores negativos.
range(-5, 5)
list(filter((lambda x: x < 0), range(-5,5)))
# Exercício 7 - Usando a função filter(), encontre os valores que são comuns às duas listas abaixo.
a = [1,2,3,5,7,9]
b = [2,3,5,6,7,8]
print (list(filter(lambda x: x in a, b)))
# Exercício 8 - Considere o código abaixo. Obtenha o mesmo resultado usando o pacote time.
# Não conhece o pacote time? Pesquise!
import datetime
print (datetime.datetime.now().strftime("%d/%m/%Y %H:%M"))
import time
print (time.strftime("%d/%m/%Y %H:%M"))
# Exercício 9 - Considere os dois dicionários abaixo.
# Crie um terceiro dicionário com as chaves do dicionário 1 e os valores do dicionário 2.
dict1 = {'a':1,'b':2}
dict2 = {'c':4,'d':5}
def trocaValores(d1, d2):
dicTemp = {}
for d1key, d2val in zip(d1,d2.values()):
dicTemp[d1key] = d2val
return dicTemp
dict3 = trocaValores(dict1, dict2)
print(dict3)
# Exercício 10 - Considere a lista abaixo e retorne apenas os elementos cujo índice for maior que 5.
lista = ['a', 'b', 'c', 'd', 'e', 'f', 'g', 'h']
for indice, valor in enumerate(lista):
if indice <= 5:
continue
else:
print (valor)
```
# Fim
### Obrigado
### Visite o Blog da Data Science Academy - <a href="http://blog.dsacademy.com.br">Blog DSA</a>
| github_jupyter |
```
import pandas as pd
import bs4
import itertools
from scipy.stats import chi2_contingency
```
## Report Writing Defs (To be moved)
```
from scipy import stats
from scipy.stats import linregress
from scipy.stats import chi2_contingency
import statsmodels.api as sm
from statsmodels.formula.api import ols
# Creating a function that provide an overview of the data
def overview(df, numerical_variable, report):
data_head = df.head()
data_shape = df.shape
data_type = df.dtypes
df = (df.drop(numerical_variable, axis=1).join(df[numerical_variable].apply(pd.to_numeric, errors='coerce'))) # Converts any non-numeric values in a numerical column into NaN
null_values = df.isnull().sum()
zero_prop = ((df[df == 0].count(axis=0)/len(df.index)).round(2)* 100)
data_summary = df.describe()
report.write(f"""<h1>Exploratory data analysis summary of {file_name}</h1><br>
<h3>The first 5 rows of content comprise of:</h3>
<table>{data_head.to_html()}</table><br>
<h3>There are a total of {data_shape[0]} rows and {data_shape[1]} columns.</h3><br>
<table>
<tr>
<th>The data type for each column is:</th>
<th>Number of NaN values for each column:</th>
<th>% of zeros in each column:</th>
</tr><tr>
<td>{data_type.to_frame().to_html()}</td>
<td>{null_values.to_frame().to_html()}</td>
<td>{zero_prop.to_frame().to_html()}</td>
</tr>
</table><br>
<h3>The summary of data:</h3>
<table>{data_summary.to_html()}</table>""")
return df
# Creating report for correlation
def run(num_var_combination, catnum_combination, cat_var_combination, report,data):
## For numeric variables
# Pearson correlation (Numerical)
report.write("<h1>Pearson Correlation Summary (numerical vs. numerical)</h1>")
for i in num_var_combination:
var1 = i[0]
var2 = i[1]
pearson_data = linregress(data[var1], data[var2])
pearson_r2, pearson_pvalue = ((pearson_data[2]**2), pearson_data[3])
report.write(f"<p>The Pearson R_Square and Pearson P-values between {var1} and {var2} are {pearson_r2} and {pearson_pvalue} respectively.</p>")
# Spearsman correlation (Ordinal)
report.write("<h1>Spearsman Correlation Summary (Ordinal numerical vs. numerical)</h1>")
for q in num_var_combination:
var1 = q[0]
var2 = q[1]
spearsman_data = stats.spearmanr(data[var1], data[var2])
spearsman_r2, spearsman_pvalue = ((spearsman_data[0]**2), spearsman_data[1])
report.write(f"<p>The Spearsman R_Square and Spearsman P-values between {var1} and {var2} are {spearsman_r2} and {spearsman_pvalue} respectively.</p>")
## For categorical-categorical variables
# Chi-Sq test
report.write("<h1>Chi Square Test Correlation Summary (categorical - categorical)</h1>")
for k in cat_var_combination:
cat1 = k[0]
cat2 = k[1]
chi_sq = pd.crosstab(data[cat1], data[cat2])
chi_sq_result = chi2_contingency(chi_sq)
report.write(f"<p>The Chi-Square P-value between {cat1} and {cat2} is {chi_sq_result[1]}.")
# ## For numeric-categorical variables
# # ONE WAY ANOVA (Cat-num variables)
# report.write("\n\n\n\n__________Correlation Summary (One Way ANOVA)__________")
# for j in catnum_combination:
# var1 = j[0]
# var2 = j[1]
# lm = ols('{} ~ {}'.format(var1,var2), data = data).fit()
# table = sm.stats.anova_lm(lm)
# one_way_anova_pvalue = table.loc[var2,'PR(>F)']
# report.write("\n\nThe One Way ANOVA P-value between {} and {} is {}."
# .format(var1, var2, one_way_anova_pvalue))
report.write("<h1>Summary Complete</h1>")
report.close()
```
## Plot Creation Defs (to be moved)
```
#TODO: Create HTML docustructure to contain all plots
#TODO: Change to accept path
#TODO: Change plots to be scrollable
import seaborn as sns
import itertools
from sklearn.preprocessing import LabelEncoder
import matplotlib.pyplot as plt
# Create a function for plots
## Look into making count plots for categorical data
## Look into making scatterplots and barplot for numerical data
## Export plots into plot path
def plot_run(data,categorical_variable,numerical_variable):
## Set Unique categorical values that are < 5 as hue
hue_lst = []
for x in categorical_variable:
if len(set(data[x])) <= 5: # if we have less than 5 unique values, we will use it for hue attributes
hue_lst.append(x)
## Creating possible combinations among a list of numerical variables
num_var_combination = list(itertools.combinations(numerical_variable, 2))
## Creating possible combinations among a list of categorical variables
cat_var_combination = list(itertools.combinations(categorical_variable, 2))
## Creating possible combinations among a list of numerical and categorical variuable
catnum_combination = list(itertools.product(numerical_variable, categorical_variable))
## Using scatterplot for numerical-numerical variables
if len(categorical_variable) > 1:
num_var_hue_combination = list(itertools.product(num_var_combination, hue_lst))
for i in num_var_hue_combination:
var1 = i[0][0]
var2 = i[0][1]
hue1 = i[1]
plot1 = sns.scatterplot(data = data, x = var1, y = var2, hue = hue1, figsize = (len(), len()))
plot1.set_xticklabels(plot1.get_xticklabels(), rotation=45, horizontalalignment='right')
fig1 = plot1.get_figure()
fig1.savefig("plots/{} vs {} by {} scatterplot.png".format(var1,var2, hue1))
fig1.clf()
else:
for l in num_var_combination:
var1 = l[0]
var2 = l[1]
plot1 = sns.scatterplot(data = data, x = var1, y = var2)
plot1.set_xticklabels(plot1.get_xticklabels(), rotation=45, horizontalalignment='right')
fig1 = plot1.get_figure()
fig1.savefig("plots/{} vs {} scatterplot.png".format(var1,var2))
fig1.clf()
## Using countplot for categorical data
for j in categorical_variable:
plot2 = sns.countplot(data = data, x = j)
plot2.set_xticklabels(plot2.get_xticklabels(), rotation=45, horizontalalignment='right')
fig2 = plot2.get_figure()
fig2.savefig("plots/{}_countplot.png".format(j))
fig2.clf()
## Using boxplot for numerical + Categorical data
for k in catnum_combination:
num1 = k[0]
cat1 = k[1]
plt.figure(figsize=(len(cat1)/3, 6))
plot3 = sns.boxplot(data = data, x = cat1, y = num1)
plot3.set_xticklabels(plot3.get_xticklabels(), rotation=45, horizontalalignment='right')
fig3 = plot3.get_figure()
fig3.savefig("plots/{}_{}_barplot.png".format(num1,cat1))
fig3.clf()
## Creating heatmap to show correlation
le = LabelEncoder()
for cat in data[categorical_variable]:
data[cat] = le.fit_transform(data[cat])
plt.figure(figsize=(15,10))
corrMatrix = data.corr()
plot4 = sns.heatmap(corrMatrix, annot=True)
plot4.set_xticklabels(plot4.get_xticklabels(), rotation=45, horizontalalignment='right')
fig4 = plot4.get_figure()
fig4.savefig("plots/heatplot.png")
fig4.clf()
```
# Data Intake
```
file_name = 'Video Game Sales'
data = pd.read_csv(r"C:\Users\602387\Desktop\Data sets\vgsales.csv")
#Defining columns as categorical or numerical
#TODO: def geo_variable
def cat_variable(df):
return list(df.select_dtypes(include = ['category', 'object']))
def num_variable(df):
return list(df.select_dtypes(exclude = ['category', 'object']))
categorical_variable = cat_variable(data)
numerical_variable = num_variable(data)
print(categorical_variable)
print(numerical_variable)
#TODO create tab structure for Summary, Correlations, Plots
#creating blank report
report = open("report.html", "w")
#template for report
report.write("""
<!DOCTYPE html>
<html>
<head>
<style>
table {
font-family: "Trebuchet MS", Arial, Helvetica, sans-serif;
border-collapse: collapse;
}
td, th {
border: 1px solid #ddd;
padding: 8px;
}
tr:nth-child(even){background-color: #f2f2f2;}
tr:hover {background-color: #ddd;}
th {
padding-top: 12px;
padding-bottom: 12px;
text-align: left;
background-color: #269bd1;
color: white;
}
</style>
<title>Correlation Reprot</title>
</head>
<body>
</body>
</html>""")
# Execute overview function in model module
data = overview(data, numerical_variable, report)
## Creating possible combinations among a list of numerical variables
num_var_combination = list(itertools.combinations(numerical_variable, 2))
## Creating possible combinations among a list of categorical variables
cat_var_combination = list(itertools.combinations(categorical_variable, 2))
## Creating possible combinations among a list of numerical and categorical variuable
catnum_combination = list(itertools.product(numerical_variable, categorical_variable))
## Running the report now
run(num_var_combination,catnum_combination,cat_var_combination,report,data)
## Running Plots
plot_run(data, categorical_variable,numerical_variable)
```
| github_jupyter |
# Tutorial Part 2: Arrays, Dictionaries and Reference Values
**Table of Content**
* [Isopy Arrays](#Isopy-Arrays)
* [Creating Arrays](#Creating-Arrays)
* [Array Attributes](#Array-Attributes)
* [Array Methods](#Array-Methods)
* [Array Functions](#Array-Functions)
* [Isopy Dictionaries](#Isopy-Dictionaries)
* [Reference Values](#Reference-Values)
```
import isopy
import numpy as np
import pandas as pd
import pyperclip # library for interacting with the clipboard
```
## Isopy Arrays
An isopy array can be seen as a table of data with a certain number of rows and columns. Each column has a key in the form of an isopy key string. These arrays allow you to easily manipulate data while keeping track of what the values represents. Technically, isopy arrays are a custom view of a structured numpy array. This means that they inherit much of the functionality of a numpy array.
### Creating Arrays
You can create arrays directly using the ``array`` and ``asarray`` functions or by directly using the ``IsopyArray`` class. Isopy arrays can be created from a range of different data, described below.
---
When the input is a list/tuple or a numpy array we have to pass along the keys for each column in the input
```
data = [10, 20, 30] # Produces a 0-dimensional array
isopy.array(data, ['ru', 'pd', 'cd'])
data = np.array([[10, 20, 30], [11, 21, 31]]) #Produces a 1-dimensional array
isopy.array(data, ['ru', 'pd', 'cd'])
```
**Note** The data type of each column defaults to ``numpy.float64`` for values that do not have a numpy dtype. If a value cannot be represented as a float the default data type inferred by numpy will be used. In the examples above the first array created will use the default data type while the second array will inherit the data type of the input, ``np.int32`` in this case.
---
Using the ``dtype`` keyword you can specify the data type for columns in the array. Any data type accepted by numpy is valid. You can either pass a single data type or a tuple of data types. The latter will use the first the data type which is valid for all the data in the column.
```
isopy.array([10, 20, 30], ['ru', 'pd', 'cd']).dtype # f8 stands for np.float64
isopy.array([10, 20, 30], ['ru', 'pd', 'cd'], dtype=np.int32).dtype # i4 stands for np.int32
isopy.array(['ten', 20, 30], ['ru', 'pd', 'cd'], dtype=(np.int32, str)).dtype # U stands for unicode string and the next number is the maximum length
```
To specify different data types for different columns pass a list of equal length to the number of columns.
```
isopy.array([10, 20, 30], ['ru', 'pd', 'cd'], dtype=[str, np.int32, np.float64]).dtype
```
---
Using the ``ndim`` keyword you can specify the number of dimensions of the array. To return a 0-dimensional array if possible, otherwise return a 1-dimensional array specify ``ndim`` as ``-1``. The row number of 0-dimensional arrays will appear as ``"None"`` in the ``repr()`` output of an array. You can also check the dimensionality of an array using the ``ndim`` attribute.
```
isopy.array([10, 20, 30], ['ru', 'pd', 'cd'], ndim=0) # Make 0-dimensional
isopy.array([10, 20, 30], ['ru', 'pd', 'cd'], ndim=1) # Make 1-dimensional
isopy.array([10, 20, 30], ['ru', 'pd', 'cd'], ndim=-1) # Make 0-dimesional if possible otherwise 1-dimensional.
isopy.array([[10, 20, 30], [11, 21, 31]], ['ru', 'pd', 'cd'], ndim = -1)
```
---
If the input is a dictionary or a structured numpy array the name of each column will be automatically inferred from the first argument.
```
data = dict(ru = [10, 11], pd= [20, 21], cd = [30, 31])
isopy.array(data)
data = np.array([(10, 20, 30), (11, 21, 31)], dtype = [('ru', float), ('pd', float), ('cd', float)])
isopy.array(data)
```
You can overwrite the inferred column keys by passing keys during creation
```
data = dict(ru = [10, 11], pd= [20, 21], cd = [30, 31])
isopy.array(data, ['101ru', '105pd', '111cd'])
```
You can convert an isopy array back into a numpy array or a dictionary using the ``to_ndarray()`` and ``to_dict()`` methods.
---
There are a number of methods for converting isopy arrays into other python objects
```
a = isopy.array(dict(ru = [10, 11], pd= [20, 21], cd = [30, 31]))
a.to_ndarray() # Converts array into a structured numpy array
a.to_dict() # Converts array into a dictionary
```
---
Similarly you can create isopy arrays from a pandas dataframe
```
df = pd.DataFrame(dict(ru = [10, 11], pd= [20, 21], cd = [30, 31]))
df
isopy.array(df)
```
You can convert a isopy array back into a dataframe using the ``to_dataframe()`` method or by passing it directly to ``DataFrame()``
```
array = isopy.array(df)
array.to_dataframe()
pd.DataFrame(array)
```
---
You can create arrays from existing isopy arrays
```
a = isopy.array([10, 20, 30], ['ru', 'pd', 'cd'])
isopy.array(a)
```
There key difference between ``array`` and ``asarray`` is that if the first argument is an isopy array then ``asarray`` will return a reference that array, rather than a copy, if no other arguments are given while ``array`` will return a copy.
```
a = isopy.array([10, 20, 30], ['ru', 'pd', 'cd'])
isopy.array(a) is a, isopy.asarray(a) is a
```
#### Filled Arrays
You can create an array of uninitiated values, zeros or ones using the ``empty``, ``zeros`` and ``one`` functions.
```
isopy.empty(None, ['ru', 'pd', 'cd']) #None, or -1, creates a 0-dimensional array
isopy.zeros(1, ['ru', 'pd', 'cd'])
isopy.ones(2, ['ru', 'pd', 'cd'])
```
To create an array filled with a specific value use the ``full`` function. The second arguement is the fill value. This can either be a single value used for all rows in the column or a sequence of values of of the same length as the number of rows.
```
isopy.full(2, np.nan, ['ru', 'pd', 'cd'])
isopy.full(2, [1,2], ['ru', 'pd', 'cd'])
```
---
If no keys are given, or can be inferred, a normal numpy array is returned.
```
isopy.ones(5) # same as np.ones(5)
```
---
#### Random Arrays
To create an array of random values use the ``random`` function. The second argument is either a single argument or a tuple of arguments that will be passed to the random generator. By default this function draws values from a normal distribution. The following example draws values from a normal distribution with a center of 1 and standard deviation of 0.1
```
isopy.random(10, (1, 0.1), ['ru', 'pd', 'cd'])
```
You can specify different distributions for different columns by passing a list as the second argument
```
isopy.random(10, [(1, 0.1), (0,1), (10, 1)], ['ru', 'pd', 'cd'])
```
---
If no keys are given, or can be inferred, a normal numpy array is returned.
```
isopy.random(10)
```
### Array Attributes
Since isopy arrays are custom implementation of a numpy arrays they have all the attributes you would find in numpy arrays, e.g. ``size``, ``.ndim``, ``.shape`` and ``.dtype``.
```
a = isopy.array(dict(ru = [10, 11], pd= [20, 21], cd = [30, 31]))
a.size, a.ndim, a.shape, a.dtype
```
In addition to the numpy attributes ``.nrows`` and ``.ncols`` are also available for isopy arrays. There return the number of rows and number of columns in the array respectively.
```
a.nrows, a.ncols
```
**Note** That ``.size`` will return ``1`` for both 0-dimensional arrays and 1-dimensional arrays with 1 row. ``.nrows`` on the other hand will return ``-1`` 0-dimensional arrays.
```
a = isopy.array(dict(ru = 10, pd= 20, cd = 30))
a.size, a.nrows
```
The column keys are available through the ``.keys`` attribute
```
a.keys # a.keys() also works fine
```
### Array Methods
While isopy arrays also contain all the methods found in numpy arrays many of these are not relevant to isopy arrays and may therefore not work as expected, if at all. See the reference documentation for a list of all methods that have been implemented for isopy arrays. Any methods not listed there should be used with **caution** as the behavior is undefined.
---
Isopy arrays have a number of methods that mimic those found in dictionaries. In addition to the ``.keys`` attribute, that can be used as a method, arrays also have ``values()``, ``items()`` and ``get()`` methods.
```
a = isopy.array(dict(ru = [10, 11], pd= [20, 21], cd = [30, 31]))
a.values()
a.items()
```
**Note** both``values()`` and ``items()`` both return a tuple (Unlike dictionaries where they return iterators).
```
a.get('ru')
```
If a column with the specified key is not present in the array a default value is return with the same shape as a column in the array.
```
a.get('ag') # If not specified the default value is np.nan
a.get('ag', 40) # Second argument is the default value
a.get('ag', [40, 41]) # A sequence the same shape as a valid column is also accepted
```
---
The ``filter()`` method return a view containing only the columns that match the supplied key filters. See the ``filter()`` method for the different key lists for available filter keywords.
```
a = isopy.array(dict(ru101 = [10, 11], pd105= [20, 21], cd111 = [30, 31]))
a.filter(mass_number_gt=104) # Return only the columns with that have a mass number greater than 104
```
---
The ``copy()`` method can be used to return a copy of the array
```
a = isopy.array(dict(ru101 = [10, 11], pd105= [20, 21], cd111 = [30, 31]))
b = a.copy()
b is a, a == b
```
You can copy only those columns that meet a certain criteria by passing filter keywords as in the ``filter()`` method described above.
```
a = isopy.array(dict(ru101 = [10, 11], pd105= [20, 21], cd111 = [30, 31]))
a.copy(mass_number_gt = 104) # Return only the columns with that have a mass number greater than 104
```
---
You can create a ratio from data within an array using the ``ratio()`` method
```
c = a.ratio('105Pd'); c
```
Ratio arrays have a ``deratio()`` method for flattening a ratio array. This requires that all column keys in the array have a common denominator
```
c.deratio()
c.deratio([20, 21]) #You can specify the value(s) for the denominator
```
---
The ``normalise`` function allows you to normalise the data in the array to a certian value. Calling the function without any arguments will normalise all the values to that the sum of each row is ``1``
```
a = isopy.array(dict(ru = [10, 11], pd= [20, 21], cd = [30, 31]))
a.normalise()
```
The optional arguments are 1) the value you with to normalise to and 2) the key(s) of the columns that the normalisation should be based on.
```
a.normalise(100, 'pd')
a.normalise([100, 1000], ['ru', 'cd']) # The sum of the specified keys will be equal to the values given
a.normalise([100, 1000], ['ru', 'cd']).normalise([20, 21], 'pd')
```
---
With the ``to_text()`` method of arrays you can convert the contents of an array to a text string. The ``str()`` ``repr()`` functions both call this function to produce two slightly different strings. The *str* function will return the raw data in each column of the array.
```
array = isopy.random(20, keys='ru pd cd'.split())
print(array) # Same as print( str(array) ) and print( array.to_text() )
```
The *repr* function will format *float* values to 5 significant digits, include an additional column with the row number, and it will only show a maximum of 10 rows.
```
array # Same as print( repr(array) ) and print( array.to_text(nrows = 10, include_row=True, f='{:.5g}') )
```
There are a number of optional arguments you can specify to change things like number formats and the delimiter for the string
```
a.to_text(delimiter = '\t', include_row=True) #includes the row number and uses a tab delimiter
```
The method ``to_clipboard()`` takes the same arguments as ``to_text()`` but copies the string to the clipboard.
```
a.to_clipboard() # It also returns the copied string
pyperclip.paste() # Paste whatever is currently is in the clipboard
```
If you are using jupyter you can use the ``display_table()`` method to render a nice table in a cell. Except from *delimiter* it takes the same arguments as ``to_text()``
```
array.display_table(include_row=True)
```
### Array Functions
The isopy package comes with several custom made array functions and isopy arrays support a large number of the numpy array functions. An array function is a function that perform an action on one or more arrays, e.g. adding arrays together or finding the mean of values in an array. See [Introduction Part 3: Working with arrays]() **LINK MISSING** for a comprehensive explanation of array functions with lots examples.
A few quick examples are
```
a1 = isopy.array(dict(ru = 1, pd= 2, cd = 3))
a2 = isopy.array(dict(ru = 10, pd= 20, ag = 25))
a1 + a2 # Columns not present in all arrays are assinged a value of np.nan
a = isopy.random(100, [(1, 0.1), (0, 1), (10, 2)], ['ru', 'pd', 'cd'])
np.mean(a) #Calculate the mean of each column
np.std(a) # Calculate the standard deviation of each column
```
---
One useful feature of the array function implementation for isopy arrays is that they can be used in conjunction with dictionaries. Only keys present in the array are included in the output meanign the dictionary can contain as many keys as you want. Thus dictionaries are useful for storing reference values.
```
a = isopy.array(dict(ru = 10, pd= 20, cd = 30))
d = dict(ru = 1, rh = 2, pd=3, ag=4, cd = 5)
a + d # Only column keys in the array are present in the output
```
**Note** The dictionary keys do not need to formatted to match the proper key string format.
## Isopy Dictionaries
Isopy has two special dictionaries, ``IsopyDict`` and ``ScalarDict``. These function much like normal dictionaries with a few enhancements. First, all values are stored as isopy key strings. Second, they can be readonly and have predefined default values. Third, create a subsection of the dictionary using ``copy()`` by passing filter keywords.
```
d = isopy.IsopyDict(ru101 = 1, rh103 = 2, pd105=3, ag107=4, cd111 = 5, default_value=0); d # The input can also be a another dictionary
d.get('76ge'), d.get('80se', 100) # If not specified the default value of the dictionary is used
d = isopy.IsopyDict(ru101 = 1, rh103 = 2, pd105=3, ag107=4, cd111 = 5, default_value=0)
d.copy(mass_number_gt = 104) # Returns a new dict containing only isotopes with a mass number greater than 104
```
---
``ScalarDict`` works just like an ``IsopyDict`` with three exceptions. First it can only store scalars, that is a single numerical value. Second, the ``get()`` method can calculate the ratio of two values in the dictionary if a ratio key string is not present in the dictionary. Finally, you can create an array directly from the dictionary with the ``to_array()`` method.
```
d = isopy.ScalarDict(ru101 = 1, rh103 = 2, pd105=3, ag107=4, cd111 = 5) # Default value is by default np.nan
d.get('pd105/cd111') # Automatically calculated from the numerator and denominator values
d.to_array()
d.to_array(mass_number_gt = 104) # You can specify key filters too
```
## Reference Values
There are a number of reference values included with isopy under the ``refval`` namespace. You can find the available reference values listed [here](https://isopy.readthedocs.io/en/latest/refpages/reference_values.html) toghether with a short description. There are currently three categories of reference values, ``mass``, ``element`` and ``isotope`` referring to the flavour of the key string of the values in the dictionaries.
```
isopy.refval.element.atomic_number.get('pd') # The atomic number of palladium
isopy.refval.isotope.fraction.get('105pd') # The natural isotope fraction of 105Pd
isopy.refval.element.isotopes.get('pd') # Returns a list of all naturally occuring isotopes of palladium
```
---
Many reference values are isopy dictionaries so the ``copy()`` method accepts filter keywords
```
isopy.refval.isotope.fraction.copy(element_symbol='pd')
```
Similarly many reference values has the ``to_array()`` method.
```
isopy.refval.isotope.fraction.to_array(element_symbol='pd')
```
| github_jupyter |
```
from ipyleaflet import Map, GeoData, basemaps, LayersControl
from ipywidgets import Dropdown, interactive
import geopandas as gpd
import pandas as pd
import json
from pathlib import Path
from IPython.display import display
PROCESSED_DATA = Path("../../data/processed/")
datos = gpd.read_file(PROCESSED_DATA / 'CA_data.shp')
datos.dropna(inplace = True)
datos.crs = {'proj': 'aea',
'lat_1': "29.5",
'lat_2': "45.5",
'lat_0': "37.5",
'lon_0': "-96",
'x_0': "0",
'y_0': "0",
'datum': 'NAD83',
'units': 'm',
'no_defs': "true"}
datos = datos.to_crs(epsg=4326)
def geo_center(df):
_bounds = df.bounds
xmin, xmax, ymin, ymax = _bounds.minx.min(), _bounds.maxx.max(), _bounds.miny.min(), _bounds.maxy.max()
y_center = (ymax + ymin) / 2
x_center = (xmax + xmin) / 2
center = (y_center, x_center)
return center
datos_female = datos[datos['sex'] == 'female'].copy()
datos_female['bins'] = pd.cut(datos_female['count'], bins = 5)
#datos_female['bins'].unique()
datos_female['bins'] = datos_female['bins'].astype(str)
_orden = ["(153686.0, 305094.0]", "(1520.96, 153686.0]", "(607910.0, 759318.0]"]
_colores = "#81d1e6","#dc0d12",["#fbe1e1",]
_escala = dict(zip(_orden, _colores))
def crea_visualizacion(datos):
_centro = geo_center(datos)
m = Map(center = _centro, zoom = 5, basemap = basemaps.OpenStreetMap.BlackAndWhite, scroll_wheel_zoom = True, )
for option in datos["bins"].unique():
option_data = datos[datos["bins"] == option]
geo_data = GeoData(
geo_dataframe = option_data,
style={'color': _escala[option], 'fillColor': _escala[option], 'opacity':0.35, 'weight':1.9, 'dashArray':'2', 'fillOpacity':0.7},
hover_style={'fillColor': _escala[option] , 'fillOpacity': 0.9},
name = f'{option}'
)
# add layers
m.add_layer(geo_data)
m.add_control(LayersControl())
return m
```
<div class='ppic-blog-titlearea'><h1>Where are the nerds?</h1><div class='ppic-author-line'><span class='ppic-blog-authors'><a href='https://www.ppic.org/person/sergio-sanchez/'>Sergio Sánchez</a>, <a href='https://www.ppic.org/person/joseph-hayes/'>Joseph Hayes</a></span> <span class='ppic-date'>June 26, 2019</span></div></div>
Lorem ipsum dolor amet dreamcatcher pabst DIY four loko disrupt. Disrupt literally deep v keytar banh mi live-edge kickstarter locavore af. Trust fund cray vexillologist drinking vinegar blog. Hoodie small batch chartreuse crucifix kickstarter health goth squid raclette 8-bit activated charcoal stumptown dreamcatcher glossier. Gochujang four loko church-key, brunch farm-to-table mlkshk.
Gastropub hashtag jianbing, man bun trust fund street art shaman kombucha retro master cleanse selvage 90's godard. Selfies echo park lumbersexual, migas sartorial four loko master fingerstache shoreditch try-hard. Shaman four loko stumptown viral, cold-pressed fixie pabst. Pickled master cleanse 90's williamsburg kombucha humblebrag wolf.
```
crea_visualizacion(datos_female)
```
| github_jupyter |
# Autoencoder (Semi-supervised)
```
%load_ext autoreload
%autoreload 2
# Seed value
# Apparently you may use different seed values at each stage
seed_value= 0
# 1. Set the `PYTHONHASHSEED` environment variable at a fixed value
import os
os.environ['PYTHONHASHSEED']=str(seed_value)
# 2. Set the `python` built-in pseudo-random generator at a fixed value
import random
random.seed(seed_value)
# 3. Set the `numpy` pseudo-random generator at a fixed value
import numpy as np
np.random.seed(seed_value)
# 4. Set the `tensorflow` pseudo-random generator at a fixed value
import tensorflow as tf
tf.set_random_seed(seed_value)
# 5. Configure a new global `tensorflow` session
from keras import backend as K
session_conf = tf.ConfigProto(intra_op_parallelism_threads=1, inter_op_parallelism_threads=1)
sess = tf.Session(graph=tf.get_default_graph(), config=session_conf)
K.set_session(sess)
import keras
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import sklearn
# plt.style.use('fivethirtyeight')
sns.set_style("whitegrid")
sns.set_context("notebook")
DATA_PATH = '../data/'
VAL_SPLITS = 4
from plot_utils import plot_confusion_matrix
from cv_utils import run_cv_f1
from cv_utils import plot_cv_roc
from cv_utils import plot_cv_roc_prc
from sklearn.model_selection import StratifiedShuffleSplit
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import cross_validate
```
For this part of the project, we will only work with the training set, that we will split again into train and validation to perform the hyperparameter tuning.
We will save the test set for the final part, when we have already tuned our hyperparameters.
```
df = pd.read_csv(os.path.join(DATA_PATH,'df_train.csv'))
df.drop(columns= df.columns[0:2],inplace=True)
df.head()
```
## Preprocessing the data
Although we are always using cross validation with `VAL_SPLITS` folds, (in general, 4), here we are gonna set only one split in order to explore how the Autoencoder works and get intuition.
```
cv = StratifiedShuffleSplit(n_splits=1,test_size=0.15,random_state=0)
# In case we want to select a subset of features
# df_ = df[['Class','V9','V14','V16','V2','V3','V17']]
df_ = df[['Class','V4','V14','V16','V12','V3','V17']]
X = df_.drop(columns='Class').to_numpy()
y = df_['Class'].to_numpy()
for idx_t, idx_v in cv.split(X,y):
X_train = X[idx_t]
y_train = y[idx_t]
X_val = X[idx_v]
y_val = y[idx_v]
# Now we need to erase the FRAUD cases on the TRAINING set
X_train_normal = X_train[y_train==0]
```
## Defining the model
```
# this is the size of our encoded representations
ENCODED_DIM = 2
INPUT_DIM = X.shape[1]
from keras.layers import Input, Dense
from keras.models import Model
from keras.models import Sequential
from keras.layers import Dense, Activation
from keras.layers import LeakyReLU
def create_encoder(input_dim, encoded_dim):
encoder = Sequential([
Dense(32, input_shape=(input_dim,)),
LeakyReLU(),
Dense(16),
LeakyReLU(),
Dense(8),
LeakyReLU(),
Dense(encoded_dim)
], name='encoder')
return encoder
def create_decoder(input_dim, encoded_dim):
decoder = Sequential([
Dense(8, input_shape=(encoded_dim,) ),
LeakyReLU(),
Dense(16),
LeakyReLU(),
Dense(8),
LeakyReLU(),
Dense(input_dim)
],name='decoder')
return decoder
def create_autoencoder(input_dim, encoded_dim, return_encoder = True):
encoder = create_encoder(input_dim,encoded_dim)
decoder = create_decoder(input_dim,encoded_dim)
inp = Input(shape=(INPUT_DIM,),name='Input_Layer')
# a layer instance is callable on a tensor, and returns a tensor
x_enc = encoder(inp)
x_out = decoder(x_enc)
# This creates a model that includes
# the Input layer and three Dense layers
autoencoder = Model(inputs=inp, outputs=x_out)
if return_encoder:
return autoencoder, encoder
else:
return autoencoder
autoencoder, encoder = create_autoencoder(INPUT_DIM,ENCODED_DIM)
print('ENCODER SUMMARY\n')
print(encoder.summary())
print('AUTOENCODER SUMMARY\n')
print(autoencoder.summary())
autoencoder.compile(optimizer='adam',
loss='mean_squared_error')
```
## Training the model
```
autoencoder.fit(x=X_train_normal, y= X_train_normal,
batch_size=512,epochs=40, validation_split=0.1) # starts training
```
## Testing
```
X_enc = encoder.predict(X_val)
X_enc_normal = X_enc[y_val==0]
X_enc_fraud = X_enc[y_val==1]
sns.scatterplot(x = X_enc_normal[:,0], y = X_enc_normal[:,1] ,label='Normal', alpha=0.5)
sns.scatterplot(x = X_enc_fraud[:,0], y = X_enc_fraud[:,1] ,label='Fraud')
X_out = autoencoder.predict(X_val)
print(X_out.shape)
X_val.shape
distances = np.sum((X_out-X_val)**2,axis=1)
bins = np.linspace(0,np.max(distances),40)
sns.distplot(distances[y_val==0],label='Normal',kde=False,
bins=bins, norm_hist=True, axlabel='Distance')
sns.distplot(distances[y_val==1],label='Fraud',kde=False, bins=bins, norm_hist=True)
bins = np.linspace(0,100,40)
sns.distplot(distances[y_val==0],label='Normal',kde=False,
bins=bins, norm_hist=True, axlabel='Distance')
sns.distplot(distances[y_val==1],label='Fraud',kde=False, bins=bins, norm_hist=True)
plt.xlim((0,100))
```
## Validating the model
```
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import cross_validate
from sklearn.metrics import f1_score
def clf_autoencoder(X,autoencoder,threshold):
"""
Classifier based in the autoencoder.
A datapoint is a nomaly if the distance of the original points
and the result of the autoencoder is greater than the threshold.
"""
X_out = autoencoder.predict(X)
distances = np.sum((X_out-X)**2,axis=1).reshape((-1,1))
# y_pred = 1 if it is anomaly
y_pred = 1.*(distances > threshold )
return y_pred
cv = StratifiedShuffleSplit(n_splits=VAL_SPLITS,test_size=0.15,random_state=0)
# Thresholds to validate
thresholds = np.linspace(0,100,100)
# List with the f1 of all the thresholds at each validation fold
f1_all = []
for i,(idx_t, idx_v) in enumerate(cv.split(X,y)):
X_train = X[idx_t]
y_train = y[idx_t]
X_val = X[idx_v]
y_val = y[idx_v]
# Now we need to erase the FRAUD cases on the TRAINING set
X_train_normal = X_train[y_train==0]
# Train the autoencoder
autoencoder, encoder = create_autoencoder(INPUT_DIM,ENCODED_DIM)
autoencoder.compile(optimizer='adam',
loss='mean_squared_error')
autoencoder.fit(x=X_train_normal, y= X_train_normal,
batch_size=512,epochs=30, shuffle=True,
verbose=0) # starts training
# Plot of the validation set in the embedding space
X_enc = encoder.predict(X_val)
X_enc_normal = X_enc[y_val==0]
X_enc_fraud = X_enc[y_val==1]
sns.scatterplot(x = X_enc_normal[:,0], y = X_enc_normal[:,1] ,label='Normal', alpha=0.5)
sns.scatterplot(x = X_enc_fraud[:,0], y = X_enc_fraud[:,1] ,label='Fraud')
plt.show()
# Transformation of the points through the autoencoder
# and calculate the predictions
y_preds=clf_autoencoder(X_val,autoencoder,thresholds)
metrics_f1 = np.array([ f1_score(y_val,y_pred) for y_pred in y_preds.T ])
f1_all.append(metrics_f1)
# Save the models into files for future use
autoencoder.save('models_autoencoder/autoencoder_fold_'+str(i+1)+'.h5')
encoder.save('models_autoencoder/encoder_fold_'+str(i+1)+'.h5')
del(autoencoder,encoder)
f1_mean = np.mean(f1_all,axis=0)
# Plot of F1-Threshold curves
for i,f1_fold in enumerate(f1_all):
sns.lineplot(thresholds,f1_fold, label='Fold '+str(i+1))
sns.scatterplot(thresholds,f1_mean,label='Mean')
plt.show()
f1_opt = f1_mean.max()
threshold_opt = thresholds[np.argmax(f1_mean)]
print('F1 max = {:.3f} at threshold = {:.3f}'.format(f1_opt,threshold_opt))
```
| github_jupyter |
# Facial Keypoint Detection
This project will be all about defining and training a convolutional neural network to perform facial keypoint detection, and using computer vision techniques to transform images of faces. The first step in any challenge like this will be to load and visualize the data you'll be working with.
Let's take a look at some examples of images and corresponding facial keypoints.
<img src='images/key_pts_example.png' width=50% height=50%/>
Facial keypoints (also called facial landmarks) are the small magenta dots shown on each of the faces in the image above. In each training and test image, there is a single face and **68 keypoints, with coordinates (x, y), for that face**. These keypoints mark important areas of the face: the eyes, corners of the mouth, the nose, etc. These keypoints are relevant for a variety of tasks, such as face filters, emotion recognition, pose recognition, and so on. Here they are, numbered, and you can see that specific ranges of points match different portions of the face.
<img src='images/landmarks_numbered.jpg' width=30% height=30%/>
---
## Load and Visualize Data
The first step in working with any dataset is to become familiar with your data; you'll need to load in the images of faces and their keypoints and visualize them! This set of image data has been extracted from the [YouTube Faces Dataset](https://www.cs.tau.ac.il/~wolf/ytfaces/), which includes videos of people in YouTube videos. These videos have been fed through some processing steps and turned into sets of image frames containing one face and the associated keypoints.
#### Training and Testing Data
This facial keypoints dataset consists of 5770 color images. All of these images are separated into either a training or a test set of data.
* 3462 of these images are training images, for you to use as you create a model to predict keypoints.
* 2308 are test images, which will be used to test the accuracy of your model.
The information about the images and keypoints in this dataset are summarized in CSV files, which we can read in using `pandas`. Let's read the training CSV and get the annotations in an (N, 2) array where N is the number of keypoints and 2 is the dimension of the keypoint coordinates (x, y).
---
First, before we do anything, we have to load in our image data. This data is stored in a zip file and in the below cell, we access it by it's URL and unzip the data in a `/data/` directory that is separate from the workspace home directory.
```
# -- DO NOT CHANGE THIS CELL -- #
!mkdir /data
!wget -P /data/ https://s3.amazonaws.com/video.udacity-data.com/topher/2018/May/5aea1b91_train-test-data/train-test-data.zip
!unzip -n /data/train-test-data.zip -d /data
# import the required libraries
import glob
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import cv2
```
Then, let's load in our training data and display some stats about that dat ato make sure it's been loaded in correctly!
```
key_pts_frame = pd.read_csv('/data/training_frames_keypoints.csv')
n = 0
image_name = key_pts_frame.iloc[n, 0]
key_pts = key_pts_frame.iloc[n, 1:].as_matrix()
key_pts = key_pts.astype('float').reshape(-1, 2)
print('Image name: ', image_name)
print('Landmarks shape: ', key_pts.shape)
print('First 4 key pts: {}'.format(key_pts[:4]))
# print out some stats about the data
print('Number of images: ', key_pts_frame.shape[0])
img2 = cv2.imread('/data/training/Nicolas_Sarkozy_02.jpg')
width, height, channels = img2.shape
npixel = img2.size
print("npixel", npixel)
print("width {}, height {}, channels {}".format(width, height, channels))
```
## Look at some images
Below, is a function `show_keypoints` that takes in an image and keypoints and displays them. As you look at this data, **note that these images are not all of the same size**, and neither are the faces! To eventually train a neural network on these images, we'll need to standardize their shape.
```
def show_keypoints(image, key_pts):
"""Show image with keypoints"""
plt.imshow(image)
plt.scatter(key_pts[:, 0], key_pts[:, 1], s=20, marker='.', c='m')
# Display a few different types of images by changing the index n
# select an image by index in our data frame
n = 0
image_name = key_pts_frame.iloc[n, 0]
key_pts = key_pts_frame.iloc[n, 1:].as_matrix()
key_pts = key_pts.astype('float').reshape(-1, 2)
plt.figure(figsize=(5, 5))
show_keypoints(mpimg.imread(os.path.join('/data/training/', image_name)), key_pts)
plt.show()
```
## Dataset class and Transformations
To prepare our data for training, we'll be using PyTorch's Dataset class. Much of this this code is a modified version of what can be found in the [PyTorch data loading tutorial](http://pytorch.org/tutorials/beginner/data_loading_tutorial.html).
#### Dataset class
``torch.utils.data.Dataset`` is an abstract class representing a
dataset. This class will allow us to load batches of image/keypoint data, and uniformly apply transformations to our data, such as rescaling and normalizing images for training a neural network.
Your custom dataset should inherit ``Dataset`` and override the following
methods:
- ``__len__`` so that ``len(dataset)`` returns the size of the dataset.
- ``__getitem__`` to support the indexing such that ``dataset[i]`` can
be used to get the i-th sample of image/keypoint data.
Let's create a dataset class for our face keypoints dataset. We will
read the CSV file in ``__init__`` but leave the reading of images to
``__getitem__``. This is memory efficient because all the images are not
stored in the memory at once but read as required.
A sample of our dataset will be a dictionary
``{'image': image, 'keypoints': key_pts}``. Our dataset will take an
optional argument ``transform`` so that any required processing can be
applied on the sample. We will see the usefulness of ``transform`` in the
next section.
```
from torch.utils.data import Dataset, DataLoader
class FacialKeypointsDataset(Dataset):
"""Face Landmarks dataset."""
def __init__(self, csv_file, root_dir, transform=None):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.key_pts_frame = pd.read_csv(csv_file)
self.root_dir = root_dir
self.transform = transform
def __len__(self):
return len(self.key_pts_frame)
def __getitem__(self, idx):
image_name = os.path.join(self.root_dir,
self.key_pts_frame.iloc[idx, 0])
image = mpimg.imread(image_name)
# if image has an alpha color channel, get rid of it
if(image.shape[2] == 4):
image = image[:,:,0:3]
key_pts = self.key_pts_frame.iloc[idx, 1:].as_matrix()
key_pts = key_pts.astype('float').reshape(-1, 2)
sample = {'image': image, 'keypoints': key_pts}
if self.transform:
sample = self.transform(sample)
return sample
```
Now that we've defined this class, let's instantiate the dataset and display some images.
```
# Construct the dataset
face_dataset = FacialKeypointsDataset(csv_file='/data/training_frames_keypoints.csv',
root_dir='/data/training/')
# print some stats about the dataset
print('Length of dataset: ', len(face_dataset))
# Display a few of the images from the dataset
num_to_display = 3
for i in range(num_to_display):
# define the size of images
fig = plt.figure(figsize=(20,10))
# randomly select a sample
rand_i = np.random.randint(0, len(face_dataset))
sample = face_dataset[rand_i]
# print the shape of the image and keypoints
print(i, sample['image'].shape, sample['keypoints'].shape)
ax = plt.subplot(1, num_to_display, i + 1)
ax.set_title('Sample #{}'.format(i))
# Using the same display function, defined earlier
show_keypoints(sample['image'], sample['keypoints'])
```
## Transforms
Now, the images above are not of the same size, and neural networks often expect images that are standardized; a fixed size, with a normalized range for color ranges and coordinates, and (for PyTorch) converted from numpy lists and arrays to Tensors.
Therefore, we will need to write some pre-processing code.
Let's create four transforms:
- ``Normalize``: to convert a color image to grayscale values with a range of [0,1] and normalize the keypoints to be in a range of about [-1, 1]
- ``Rescale``: to rescale an image to a desired size.
- ``RandomCrop``: to crop an image randomly.
- ``ToTensor``: to convert numpy images to torch images.
We will write them as callable classes instead of simple functions so
that parameters of the transform need not be passed everytime it's
called. For this, we just need to implement ``__call__`` method and
(if we require parameters to be passed in), the ``__init__`` method.
We can then use a transform like this:
tx = Transform(params)
transformed_sample = tx(sample)
Observe below how these transforms are generally applied to both the image and its keypoints.
```
import torch
from torchvision import transforms, utils
# tranforms
class Normalize(object):
"""Convert a color image to grayscale and normalize the color range to [0,1]."""
def __call__(self, sample):
image, key_pts = sample['image'], sample['keypoints']
image_copy = np.copy(image)
key_pts_copy = np.copy(key_pts)
# convert image to grayscale
image_copy = cv2.cvtColor(image, cv2.COLOR_RGB2GRAY)
# scale color range from [0, 255] to [0, 1]
image_copy= image_copy/255.0
# scale keypoints to be centered around 0 with a range of [-1, 1]
# mean = 100, sqrt = 50, so, pts should be (pts - 100)/50
key_pts_copy = (key_pts_copy - 100)/50.0
return {'image': image_copy, 'keypoints': key_pts_copy}
class Rescale(object):
"""Rescale the image in a sample to a given size.
Args:
output_size (tuple or int): Desired output size. If tuple, output is
matched to output_size. If int, smaller of image edges is matched
to output_size keeping aspect ratio the same.
"""
def __init__(self, output_size):
assert isinstance(output_size, (int, tuple))
self.output_size = output_size
def __call__(self, sample):
image, key_pts = sample['image'], sample['keypoints']
h, w = image.shape[:2]
if isinstance(self.output_size, int):
if h > w:
new_h, new_w = self.output_size * h / w, self.output_size
else:
new_h, new_w = self.output_size, self.output_size * w / h
else:
new_h, new_w = self.output_size
new_h, new_w = int(new_h), int(new_w)
img = cv2.resize(image, (new_w, new_h))
# scale the pts, too
key_pts = key_pts * [new_w / w, new_h / h]
return {'image': img, 'keypoints': key_pts}
class RandomCrop(object):
"""Crop randomly the image in a sample.
Args:
output_size (tuple or int): Desired output size. If int, square crop
is made.
"""
def __init__(self, output_size):
assert isinstance(output_size, (int, tuple))
if isinstance(output_size, int):
self.output_size = (output_size, output_size)
else:
assert len(output_size) == 2
self.output_size = output_size
def __call__(self, sample):
image, key_pts = sample['image'], sample['keypoints']
h, w = image.shape[:2]
new_h, new_w = self.output_size
top = np.random.randint(0, h - new_h)
left = np.random.randint(0, w - new_w)
image = image[top: top + new_h,
left: left + new_w]
key_pts = key_pts - [left, top]
return {'image': image, 'keypoints': key_pts}
class ToTensor(object):
"""Convert ndarrays in sample to Tensors."""
def __call__(self, sample):
image, key_pts = sample['image'], sample['keypoints']
# if image has no grayscale color channel, add one
if(len(image.shape) == 2):
# add that third color dim
image = image.reshape(image.shape[0], image.shape[1], 1)
# swap color axis because
# numpy image: H x W x C
# torch image: C X H X W
image = image.transpose((2, 0, 1))
return {'image': torch.from_numpy(image),
'keypoints': torch.from_numpy(key_pts)}
```
## Test out the transforms
Let's test these transforms out to make sure they behave as expected. As you look at each transform, note that, in this case, **order does matter**. For example, you cannot crop a image using a value smaller than the original image (and the orginal images vary in size!), but, if you first rescale the original image, you can then crop it to any size smaller than the rescaled size.
```
# test out some of these transforms
rescale = Rescale(100)
crop = RandomCrop(50)
composed = transforms.Compose([Rescale(250),
RandomCrop(224)])
# apply the transforms to a sample image
test_num = 500
sample = face_dataset[test_num]
fig = plt.figure()
for i, tx in enumerate([rescale, crop, composed]):
transformed_sample = tx(sample)
ax = plt.subplot(1, 3, i + 1)
plt.tight_layout()
ax.set_title(type(tx).__name__)
show_keypoints(transformed_sample['image'], transformed_sample['keypoints'])
plt.show()
```
## Create the transformed dataset
Apply the transforms in order to get grayscale images of the same shape. Verify that your transform works by printing out the shape of the resulting data (printing out a few examples should show you a consistent tensor size).
```
# define the data tranform
# order matters! i.e. rescaling should come before a smaller crop
data_transform = transforms.Compose([Rescale(250),
RandomCrop(224),
Normalize(),
ToTensor()])
# create the transformed dataset
transformed_dataset = FacialKeypointsDataset(csv_file='/data/training_frames_keypoints.csv',
root_dir='/data/training/',
transform=data_transform)
# print some stats about the transformed data
print('Number of images: ', len(transformed_dataset))
# make sure the sample tensors are the expected size
for i in range(5):
sample = transformed_dataset[i]
print(i, sample['image'].size(), sample['keypoints'].size())
```
## Data Iteration and Batching
Right now, we are iterating over this data using a ``for`` loop, but we are missing out on a lot of PyTorch's dataset capabilities, specifically the abilities to:
- Batch the data
- Shuffle the data
- Load the data in parallel using ``multiprocessing`` workers.
``torch.utils.data.DataLoader`` is an iterator which provides all these
features, and we'll see this in use in the *next* notebook, Notebook 2, when we load data in batches to train a neural network!
---
## Ready to Train!
Now that you've seen how to load and transform our data, you're ready to build a neural network to train on this data.
In the next notebook, you'll be tasked with creating a CNN for facial keypoint detection.
| github_jupyter |
```
import os
import pickle
import re
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sn
import numpy as np
import re
import xgboost as xgb
import shap
from sklearn import ensemble
from sklearn import dummy
from sklearn import linear_model
from sklearn import svm
from sklearn import neural_network
from sklearn import metrics
from sklearn import preprocessing
from sklearn.model_selection import train_test_split
from sklearn.utils.fixes import loguniform
import scipy
from misc import save_model, load_model, regression_results, grid_search_cv
# Options of settings with different Xs and Ys
options = ["../data/Train_Compound_Viral_interactions_for_Supervised_Learning_with_LS_LS.csv",
"../data/Train_Compound_Viral_interactions_for_Supervised_Learning_with_MFP_LS.csv",
".."] #(to be continued)
data_type_options = ["LS_Compound_LS_Protein",
"MFP_Compound_LS_Protein",
".."
]
# input option is also used to control the model parameters below
input_option = 0
classification_task = False
classification_th = 85
data_type=data_type_options[input_option]
filename = options[input_option]
with open(filename, "rb") as file:
print("Loading ", filename)
big_df = pd.read_csv(filename, header='infer', delimiter=",")
total_length = len(big_df.columns)
X = big_df.iloc[:,range(5,total_length)]
Y = big_df[['pchembl_value']].to_numpy().flatten()
meta_X = big_df.iloc[:,[0,1,2,3]]
print("Lengths --> X = %d, Y = %d" % (len(X), len(Y)))
print(X.columns)
n_samples = len(X)
indices = np.arange(n_samples)
X_train = X
y_train = Y
print(X_train[:10])
print(X_train.shape,y_train.shape)
print(X_train.columns)
print(big_df.isnull().sum().sum())
def calculate_classification_metrics(labels, predictions):
predictions = predictions.round()
fpr, tpr, thresholds = metrics.roc_curve(labels, predictions)
auc = metrics.auc(fpr, tpr)
aupr = metrics.average_precision_score(labels,predictions)
return metrics.accuracy_score(labels, predictions),\
metrics.f1_score(labels, predictions, average='binary'),\
auc,\
aupr
def calculate_regression_metrics(labels, predictions):
return metrics.mean_absolute_error(labels, predictions),\
metrics.mean_squared_error(labels, predictions),\
metrics.r2_score(labels, predictions),\
scipy.stats.pearsonr(np.array(labels).flatten(),np.array(predictions.flatten()))[0],\
scipy.stats.spearmanr(np.array(labels).flatten(),np.array(predictions.flatten()))[0]
def supervised_learning_steps(method,scoring,data_type,task,model,params,X_train,y_train,n_iter):
gs = grid_search_cv(model, params, X_train, y_train, scoring=scoring, n_iter = n_iter)
y_pred = gs.predict(X_train)
y_pred[y_pred < 0] = 0
if task:
results=calculate_classification_metrics(y_train, y_pred)
print("Acc: %.3f, F1: %.3f, AUC: %.3f, AUPR: %.3f" % (results[0], results[1], results[2], results[3]))
else:
results=calculate_regression_metrics(y_train,y_pred)
print("MAE: %.3f, MSE: %.3f, R2: %.3f, Pearson R: %.3f, Spearman R: %.3f" % (results[0], results[1], results[2], results[3], results[4]))
print('Parameters')
print('----------')
for p,v in gs.best_estimator_.get_params().items():
print(p, ":", v)
print('-' * 80)
if task:
save_model(gs, "%s_models/%s_%s_classifier_gs.pk" % (method,method,data_type))
save_model(gs.best_estimator_, "%s_models/%s_%s_classifier_best_estimator.pk" %(method,method,data_type))
else:
save_model(gs, "%s_models/%s_%s_regressor_gs.pk" % (method,method,data_type))
save_model(gs.best_estimator_, "%s_models/%s_%s_regressor_best_estimator.pk" %(method,method,data_type))
return(gs)
if classification_task:
model = ensemble.RandomForestRegressor(n_estimators=100, criterion='auc',
max_depth=None, min_samples_split=2,
min_samples_leaf=1, min_weight_fraction_leaf=0.0,
max_features='auto', max_leaf_nodes=None,
min_impurity_decrease=0.0, min_impurity_split=None,
bootstrap=True, oob_score=False,
n_jobs=-1, random_state=328, verbose=1,
warm_start=False, ccp_alpha=0.0, max_samples=None)
else:
model = ensemble.RandomForestRegressor(n_estimators=100, criterion='mse',
max_depth=None, min_samples_split=2,
min_samples_leaf=1, min_weight_fraction_leaf=0.0,
max_features='auto', max_leaf_nodes=None,
min_impurity_decrease=0.0, min_impurity_split=None,
bootstrap=True, oob_score=False,
n_jobs=-1, random_state=328, verbose=1,
warm_start=False, ccp_alpha=0.0, max_samples=None)
# Grid parameters
param_rf = {"n_estimators": scipy.stats.randint(20, 500),
"max_depth": scipy.stats.randint(1, 9),
"min_samples_leaf": scipy.stats.randint(1, 10),
"max_features": scipy.stats.uniform.ppf([0.1,0.7])
}
n_iter=200
if classification_task:
rf_gs=supervised_learning_steps("rf","roc_auc",data_type,classification_task,model,param_rf,X_train,y_train,n_iter)
else:
rf_gs=supervised_learning_steps("rf","r2",data_type,classification_task,model,param_rf,X_train,y_train,n_iter)
rf_gs.cv_results_
rf_gs = load_model("rf_models/rf__LS_Drug_LS_Protein_regressor_gs.pk")
np.max(rf_gs.cv_results_["mean_test_score"])
file_list = ["../data/Test_Compound_Viral_interactions_for_Supervised_Learning_with_LS_LS.csv",
"../data/Test_Compound_Viral_interactions_for_Supervised_Learning_with_MFP_LS.csv"]
filename = file_list[input_option]
with open(filename, "rb") as file:
print("Loading ", filename)
big_df = pd.read_csv(filename, header='infer', delimiter=",")
total_length = len(big_df.columns)
X = big_df.iloc[:,range(5,total_length)]
Y = big_df[['pchembl_value']].to_numpy().flatten()
meta_X = big_df.iloc[:,[0,1,2,3]]
print("Lengths --> X = %d, Y = %d" % (len(X), len(Y)))
print(X.columns)
n_samples = len(X)
indices = np.arange(n_samples)
X_test = X
y_test = Y
rf_best = rf_gs.best_estimator_
y_pred_rf=rf_best.predict(X_test)
print(calculate_regression_metrics(y_test,y_pred_rf))
#Write the output in the results folder
meta_X["predictions"]=y_pred_xgb
meta_X["labels"]=y_test
rev_output_df = meta_X.iloc[:,[0,2,4,5]].copy()
rev_output_df.to_csv("../results/RF_"+data_type_options[input_option]+"supervised_test_predictions.csv",index=False)
## load JS visualization code to notebook (Doesn't work for random forest)
#shap.initjs()
## explain the model's predictions using SHAP values
#explainer = shap.TreeExplainer(xgb_gs.best_estimator_)
#shap_values = explainer.shap_values(X_train)
#shap.summary_plot(shap_values, X_train)
##Get results for SARS-COV-2
#big_X_test = pd.read_csv("../data/COVID-19/sars_cov_2_additional_compound_viral_interactions_to_predict_with_LS_v2.csv",header='infer',sep=",")
#total_length = len(big_X_test.columns)
#X_test = big_X_test.iloc[:,range(8,total_length)]
#rf_best = load_model("../models/rf_models/rf__LS_Drug_LS_Protein_regressor_best_estimator.pk")
#y_pred = rf_best.predict(X_test)
#meta_X_test = big_X_test.iloc[:,[0,2]].copy()
#meta_X_test.loc[:,'predictions']=y_pred
#meta_X_test.loc[:,'labels']=0
#meta_X_test.to_csv("../results/RF_supervised_sars_cov2_additional_test_predictions.csv",index=False)
```
| github_jupyter |
# 2.4 Deep Taylor Decomposition Part 2.
## Tensorflow Walkthrough
### 1. Import Dependencies
I made a custom `Taylor` class for Deep Taylor Decomposition. If you are interested in the details, check out `models_3_2.py` in the models directory.
```
import os
import re
from tensorflow.examples.tutorials.mnist import input_data
from tensorflow.python.ops import nn_ops, gen_nn_ops
import matplotlib.pyplot as plt
import tensorflow as tf
import numpy as np
from models.models_2_4 import MNIST_CNN, Taylor
%matplotlib inline
mnist = input_data.read_data_sets("MNIST_data/", one_hot=True)
images = mnist.train.images
labels = mnist.train.labels
logdir = './tf_logs/2_4_DTD/'
ckptdir = logdir + 'model'
if not os.path.exists(logdir):
os.mkdir(logdir)
```
### 2. Building Graph
```
with tf.name_scope('Classifier'):
# Initialize neural network
DNN = MNIST_CNN('CNN')
# Setup training process
X = tf.placeholder(tf.float32, [None, 784], name='X')
Y = tf.placeholder(tf.float32, [None, 10], name='Y')
activations, logits = DNN(X)
tf.add_to_collection('DTD', X)
for activation in activations:
tf.add_to_collection('DTD', activation)
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits=logits, labels=Y))
optimizer = tf.train.AdamOptimizer().minimize(cost, var_list=DNN.vars)
correct_prediction = tf.equal(tf.argmax(logits, 1), tf.argmax(Y, 1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
cost_summary = tf.summary.scalar('Cost', cost)
accuray_summary = tf.summary.scalar('Accuracy', accuracy)
summary = tf.summary.merge_all()
```
### 3. Training Network
This is the step where the DNN is trained to classify the 10 digits of the MNIST images. Summaries are written into the logdir and you can visualize the statistics using tensorboard by typing this command: `tensorboard --lodir=./tf_logs`
```
sess = tf.InteractiveSession()
sess.run(tf.global_variables_initializer())
saver = tf.train.Saver()
file_writer = tf.summary.FileWriter(logdir, tf.get_default_graph())
# Hyper parameters
training_epochs = 15
batch_size = 100
for epoch in range(training_epochs):
total_batch = int(mnist.train.num_examples / batch_size)
avg_cost = 0
avg_acc = 0
for i in range(total_batch):
batch_xs, batch_ys = mnist.train.next_batch(batch_size)
_, c, a, summary_str = sess.run([optimizer, cost, accuracy, summary], feed_dict={X: batch_xs, Y: batch_ys})
avg_cost += c / total_batch
avg_acc += a / total_batch
file_writer.add_summary(summary_str, epoch * total_batch + i)
print('Epoch:', '%04d' % (epoch + 1), 'cost =', '{:.9f}'.format(avg_cost), 'accuracy =', '{:.9f}'.format(avg_acc))
saver.save(sess, ckptdir)
print('Accuracy:', sess.run(accuracy, feed_dict={X: mnist.test.images, Y: mnist.test.labels}))
sess.close()
```
### 4. Restoring Subgraph
Here we first rebuild the DNN graph from metagraph, restore DNN parameters from the checkpoint and then gather the necessary weight and biases for Deep Taylor Decomposition using the `tf.get_collection()` function.
```
tf.reset_default_graph()
sess = tf.InteractiveSession()
new_saver = tf.train.import_meta_graph(ckptdir + '.meta')
new_saver.restore(sess, tf.train.latest_checkpoint(logdir))
weights = tf.get_collection(tf.GraphKeys.TRAINABLE_VARIABLES, scope='CNN')
activations = tf.get_collection('DTD')
X = activations[0]
```
### 5. Attaching Subgraph for Calculating Relevance Scores
```
conv_ksize = [1, 3, 3, 1]
pool_ksize = [1, 2, 2, 1]
conv_strides = [1, 1, 1, 1]
pool_strides = [1, 2, 2, 1]
weights.reverse()
activations.reverse()
taylor = Taylor(activations, weights, conv_ksize, pool_ksize, conv_strides, pool_strides, 'Taylor')
Rs = []
for i in range(10):
Rs.append(taylor(i))
```
### 6. Calculating Relevance Scores $R(x_i)$
```
sample_imgs = []
for i in range(10):
sample_imgs.append(images[np.argmax(labels, axis=1) == i][3])
imgs = []
for i in range(10):
imgs.append(sess.run(Rs[i], feed_dict={X: sample_imgs[i][None,:]}))
```
### 7. Displaying Images
The relevance scores are visualized as heat maps. You can see which features/data points influenced the DNN most its decision making.
```
plt.figure(figsize=(15,15))
for i in range(5):
plt.subplot(5, 2, 2 * i + 1)
plt.imshow(np.reshape(imgs[2 * i], [28, 28]), cmap='hot_r')
plt.title('Digit: {}'.format(2 * i))
plt.colorbar()
plt.subplot(5, 2, 2 * i + 2)
plt.imshow(np.reshape(imgs[2 * i + 1], [28, 28]), cmap='hot_r')
plt.title('Digit: {}'.format(2 * i + 1))
plt.colorbar()
plt.tight_layout()
```
| github_jupyter |
# Initialization
Welcome to the first assignment of "Improving Deep Neural Networks".
Training your neural network requires specifying an initial value of the weights. A well chosen initialization method will help learning.
If you completed the previous course of this specialization, you probably followed our instructions for weight initialization, and it has worked out so far. But how do you choose the initialization for a new neural network? In this notebook, you will see how different initializations lead to different results.
A well chosen initialization can:
- Speed up the convergence of gradient descent
- Increase the odds of gradient descent converging to a lower training (and generalization) error
To get started, run the following cell to load the packages and the planar dataset you will try to classify.
```
import numpy as np
import matplotlib.pyplot as plt
import sklearn
import sklearn.datasets
from init_utils import sigmoid, relu, compute_loss, forward_propagation, backward_propagation
from init_utils import update_parameters, predict, load_dataset, plot_decision_boundary, predict_dec
%matplotlib inline
plt.rcParams['figure.figsize'] = (7.0, 4.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
# load image dataset: blue/red dots in circles
train_X, train_Y, test_X, test_Y = load_dataset()
```
You would like a classifier to separate the blue dots from the red dots.
## 1 - Neural Network model
You will use a 3-layer neural network (already implemented for you). Here are the initialization methods you will experiment with:
- *Zeros initialization* -- setting `initialization = "zeros"` in the input argument.
- *Random initialization* -- setting `initialization = "random"` in the input argument. This initializes the weights to large random values.
- *He initialization* -- setting `initialization = "he"` in the input argument. This initializes the weights to random values scaled according to a paper by He et al., 2015.
**Instructions**: Please quickly read over the code below, and run it. In the next part you will implement the three initialization methods that this `model()` calls.
```
def model(X, Y, learning_rate = 0.01, num_iterations = 15000, print_cost = True, initialization = "he"):
"""
Implements a three-layer neural network: LINEAR->RELU->LINEAR->RELU->LINEAR->SIGMOID.
Arguments:
X -- input data, of shape (2, number of examples)
Y -- true "label" vector (containing 0 for red dots; 1 for blue dots), of shape (1, number of examples)
learning_rate -- learning rate for gradient descent
num_iterations -- number of iterations to run gradient descent
print_cost -- if True, print the cost every 1000 iterations
initialization -- flag to choose which initialization to use ("zeros","random" or "he")
Returns:
parameters -- parameters learnt by the model
"""
grads = {}
costs = [] # to keep track of the loss
m = X.shape[1] # number of examples
layers_dims = [X.shape[0], 10, 5, 1]
# Initialize parameters dictionary.
if initialization == "zeros":
parameters = initialize_parameters_zeros(layers_dims)
elif initialization == "random":
parameters = initialize_parameters_random(layers_dims)
elif initialization == "he":
parameters = initialize_parameters_he(layers_dims)
# Loop (gradient descent)
for i in range(0, num_iterations):
# Forward propagation: LINEAR -> RELU -> LINEAR -> RELU -> LINEAR -> SIGMOID.
a3, cache = forward_propagation(X, parameters)
# Loss
cost = compute_loss(a3, Y)
# Backward propagation.
grads = backward_propagation(X, Y, cache)
# Update parameters.
parameters = update_parameters(parameters, grads, learning_rate)
# Print the loss every 1000 iterations
if print_cost and i % 1000 == 0:
print("Cost after iteration {}: {}".format(i, cost))
costs.append(cost)
# plot the loss
plt.plot(costs)
plt.ylabel('cost')
plt.xlabel('iterations (per hundreds)')
plt.title("Learning rate =" + str(learning_rate))
plt.show()
return parameters
```
## 2 - Zero initialization
There are two types of parameters to initialize in a neural network:
- the weight matrices $(W^{[1]}, W^{[2]}, W^{[3]}, ..., W^{[L-1]}, W^{[L]})$
- the bias vectors $(b^{[1]}, b^{[2]}, b^{[3]}, ..., b^{[L-1]}, b^{[L]})$
**Exercise**: Implement the following function to initialize all parameters to zeros. You'll see later that this does not work well since it fails to "break symmetry", but lets try it anyway and see what happens. Use np.zeros((..,..)) with the correct shapes.
```
# GRADED FUNCTION: initialize_parameters_zeros
def initialize_parameters_zeros(layers_dims):
"""
Arguments:
layer_dims -- python array (list) containing the size of each layer.
Returns:
parameters -- python dictionary containing your parameters "W1", "b1", ..., "WL", "bL":
W1 -- weight matrix of shape (layers_dims[1], layers_dims[0])
b1 -- bias vector of shape (layers_dims[1], 1)
...
WL -- weight matrix of shape (layers_dims[L], layers_dims[L-1])
bL -- bias vector of shape (layers_dims[L], 1)
"""
parameters = {}
L = len(layers_dims) # number of layers in the network
for l in range(1, L):
### START CODE HERE ### (≈ 2 lines of code)
parameters['W' + str(l)] = np.zeros((layers_dims[l], layers_dims[l-1]))
parameters['b' + str(l)] = np.zeros((layers_dims[l],1))
### END CODE HERE ###
return parameters
parameters = initialize_parameters_zeros([3,2,1])
print("W1 = " + str(parameters["W1"]))
print("b1 = " + str(parameters["b1"]))
print("W2 = " + str(parameters["W2"]))
print("b2 = " + str(parameters["b2"]))
```
**Expected Output**:
<table>
<tr>
<td>
**W1**
</td>
<td>
[[ 0. 0. 0.]
[ 0. 0. 0.]]
</td>
</tr>
<tr>
<td>
**b1**
</td>
<td>
[[ 0.]
[ 0.]]
</td>
</tr>
<tr>
<td>
**W2**
</td>
<td>
[[ 0. 0.]]
</td>
</tr>
<tr>
<td>
**b2**
</td>
<td>
[[ 0.]]
</td>
</tr>
</table>
Run the following code to train your model on 15,000 iterations using zeros initialization.
```
parameters = model(train_X, train_Y, initialization = "zeros")
print ("On the train set:")
predictions_train = predict(train_X, train_Y, parameters)
print ("On the test set:")
predictions_test = predict(test_X, test_Y, parameters)
```
The performance is really bad, and the cost does not really decrease, and the algorithm performs no better than random guessing. Why? Lets look at the details of the predictions and the decision boundary:
```
print ("predictions_train = " + str(predictions_train))
print ("predictions_test = " + str(predictions_test))
plt.title("Model with Zeros initialization")
axes = plt.gca()
axes.set_xlim([-1.5,1.5])
axes.set_ylim([-1.5,1.5])
plot_decision_boundary(lambda x: predict_dec(parameters, x.T), train_X, train_Y)
```
The model is predicting 0 for every example.
In general, initializing all the weights to zero results in the network failing to break symmetry. This means that every neuron in each layer will learn the same thing, and you might as well be training a neural network with $n^{[l]}=1$ for every layer, and the network is no more powerful than a linear classifier such as logistic regression.
<font color='blue'>
**What you should remember**:
- The weights $W^{[l]}$ should be initialized randomly to break symmetry.
- It is however okay to initialize the biases $b^{[l]}$ to zeros. Symmetry is still broken so long as $W^{[l]}$ is initialized randomly.
## 3 - Random initialization
To break symmetry, lets intialize the weights randomly. Following random initialization, each neuron can then proceed to learn a different function of its inputs. In this exercise, you will see what happens if the weights are intialized randomly, but to very large values.
**Exercise**: Implement the following function to initialize your weights to large random values (scaled by \*10) and your biases to zeros. Use `np.random.randn(..,..) * 10` for weights and `np.zeros((.., ..))` for biases. We are using a fixed `np.random.seed(..)` to make sure your "random" weights match ours, so don't worry if running several times your code gives you always the same initial values for the parameters.
```
# GRADED FUNCTION: initialize_parameters_random
def initialize_parameters_random(layers_dims):
"""
Arguments:
layer_dims -- python array (list) containing the size of each layer.
Returns:
parameters -- python dictionary containing your parameters "W1", "b1", ..., "WL", "bL":
W1 -- weight matrix of shape (layers_dims[1], layers_dims[0])
b1 -- bias vector of shape (layers_dims[1], 1)
...
WL -- weight matrix of shape (layers_dims[L], layers_dims[L-1])
bL -- bias vector of shape (layers_dims[L], 1)
"""
np.random.seed(3) # This seed makes sure your "random" numbers will be the as ours
parameters = {}
L = len(layers_dims) # integer representing the number of layers
for l in range(1, L):
### START CODE HERE ### (≈ 2 lines of code)
parameters['W' + str(l)] = np.random.randn(layers_dims[l], layers_dims[l-1]) * 10
parameters['b' + str(l)] = np.zeros((layers_dims[l], 1))
### END CODE HERE ###
return parameters
parameters = initialize_parameters_random([3, 2, 1])
print("W1 = " + str(parameters["W1"]))
print("b1 = " + str(parameters["b1"]))
print("W2 = " + str(parameters["W2"]))
print("b2 = " + str(parameters["b2"]))
```
**Expected Output**:
<table>
<tr>
<td>
**W1**
</td>
<td>
[[ 17.88628473 4.36509851 0.96497468]
[-18.63492703 -2.77388203 -3.54758979]]
</td>
</tr>
<tr>
<td>
**b1**
</td>
<td>
[[ 0.]
[ 0.]]
</td>
</tr>
<tr>
<td>
**W2**
</td>
<td>
[[-0.82741481 -6.27000677]]
</td>
</tr>
<tr>
<td>
**b2**
</td>
<td>
[[ 0.]]
</td>
</tr>
</table>
Run the following code to train your model on 15,000 iterations using random initialization.
```
parameters = model(train_X, train_Y, initialization = "random")
print ("On the train set:")
predictions_train = predict(train_X, train_Y, parameters)
print ("On the test set:")
predictions_test = predict(test_X, test_Y, parameters)
```
If you see "inf" as the cost after the iteration 0, this is because of numerical roundoff; a more numerically sophisticated implementation would fix this. But this isn't worth worrying about for our purposes.
Anyway, it looks like you have broken symmetry, and this gives better results. than before. The model is no longer outputting all 0s.
```
print (predictions_train)
print (predictions_test)
plt.title("Model with large random initialization")
axes = plt.gca()
axes.set_xlim([-1.5,1.5])
axes.set_ylim([-1.5,1.5])
plot_decision_boundary(lambda x: predict_dec(parameters, x.T), train_X, train_Y)
```
**Observations**:
- The cost starts very high. This is because with large random-valued weights, the last activation (sigmoid) outputs results that are very close to 0 or 1 for some examples, and when it gets that example wrong it incurs a very high loss for that example. Indeed, when $\log(a^{[3]}) = \log(0)$, the loss goes to infinity.
- Poor initialization can lead to vanishing/exploding gradients, which also slows down the optimization algorithm.
- If you train this network longer you will see better results, but initializing with overly large random numbers slows down the optimization.
<font color='blue'>
**In summary**:
- Initializing weights to very large random values does not work well.
- Hopefully intializing with small random values does better. The important question is: how small should be these random values be? Lets find out in the next part!
## 4 - He initialization
Finally, try "He Initialization"; this is named for the first author of He et al., 2015. (If you have heard of "Xavier initialization", this is similar except Xavier initialization uses a scaling factor for the weights $W^{[l]}$ of `sqrt(1./layers_dims[l-1])` where He initialization would use `sqrt(2./layers_dims[l-1])`.)
**Exercise**: Implement the following function to initialize your parameters with He initialization.
**Hint**: This function is similar to the previous `initialize_parameters_random(...)`. The only difference is that instead of multiplying `np.random.randn(..,..)` by 10, you will multiply it by $\sqrt{\frac{2}{\text{dimension of the previous layer}}}$, which is what He initialization recommends for layers with a ReLU activation.
```
# GRADED FUNCTION: initialize_parameters_he
def initialize_parameters_he(layers_dims):
"""
Arguments:
layer_dims -- python array (list) containing the size of each layer.
Returns:
parameters -- python dictionary containing your parameters "W1", "b1", ..., "WL", "bL":
W1 -- weight matrix of shape (layers_dims[1], layers_dims[0])
b1 -- bias vector of shape (layers_dims[1], 1)
...
WL -- weight matrix of shape (layers_dims[L], layers_dims[L-1])
bL -- bias vector of shape (layers_dims[L], 1)
"""
np.random.seed(3)
parameters = {}
L = len(layers_dims) - 1 # integer representing the number of layers
for l in range(1, L + 1):
### START CODE HERE ### (≈ 2 lines of code)
parameters['W' + str(l)] = np.random.randn(layers_dims[l], layers_dims[l-1]) * np.sqrt(2./layers_dims[l-1])
parameters['b' + str(l)] = np.zeros((layers_dims[l], 1))
### END CODE HERE ###
return parameters
parameters = initialize_parameters_he([2, 4, 1])
print("W1 = " + str(parameters["W1"]))
print("b1 = " + str(parameters["b1"]))
print("W2 = " + str(parameters["W2"]))
print("b2 = " + str(parameters["b2"]))
```
**Expected Output**:
<table>
<tr>
<td>
**W1**
</td>
<td>
[[ 1.78862847 0.43650985]
[ 0.09649747 -1.8634927 ]
[-0.2773882 -0.35475898]
[-0.08274148 -0.62700068]]
</td>
</tr>
<tr>
<td>
**b1**
</td>
<td>
[[ 0.]
[ 0.]
[ 0.]
[ 0.]]
</td>
</tr>
<tr>
<td>
**W2**
</td>
<td>
[[-0.03098412 -0.33744411 -0.92904268 0.62552248]]
</td>
</tr>
<tr>
<td>
**b2**
</td>
<td>
[[ 0.]]
</td>
</tr>
</table>
Run the following code to train your model on 15,000 iterations using He initialization.
```
parameters = model(train_X, train_Y, initialization = "he")
print ("On the train set:")
predictions_train = predict(train_X, train_Y, parameters)
print ("On the test set:")
predictions_test = predict(test_X, test_Y, parameters)
plt.title("Model with He initialization")
axes = plt.gca()
axes.set_xlim([-1.5,1.5])
axes.set_ylim([-1.5,1.5])
plot_decision_boundary(lambda x: predict_dec(parameters, x.T), train_X, train_Y)
```
**Observations**:
- The model with He initialization separates the blue and the red dots very well in a small number of iterations.
## 5 - Conclusions
You have seen three different types of initializations. For the same number of iterations and same hyperparameters the comparison is:
<table>
<tr>
<td>
**Model**
</td>
<td>
**Train accuracy**
</td>
<td>
**Problem/Comment**
</td>
</tr>
<td>
3-layer NN with zeros initialization
</td>
<td>
50%
</td>
<td>
fails to break symmetry
</td>
<tr>
<td>
3-layer NN with large random initialization
</td>
<td>
83%
</td>
<td>
too large weights
</td>
</tr>
<tr>
<td>
3-layer NN with He initialization
</td>
<td>
99%
</td>
<td>
recommended method
</td>
</tr>
</table>
<font color='blue'>
**What you should remember from this notebook**:
- Different initializations lead to different results
- Random initialization is used to break symmetry and make sure different hidden units can learn different things
- Don't intialize to values that are too large
- He initialization works well for networks with ReLU activations.
```
!tar cvfz notebook.tar.gz *
!tar cvfz notebook.tar.gz *
!tar cvfz notebook.tar.gz *
```
| github_jupyter |
# Day 3 - Advent of Code 2020 (https://adventofcode.com/2020/day/3)
## Data Preparation
As with the previous days, I will be starting with getting my input into a good format before even taken a reat pass at the problem...
```
real_run = False
file_name = "day3-input.txt" if real_run else "day3-test.txt"
# create a list from the file, removing any '\n' characters
data = [line.rstrip('\n') for line in open(file_name)]
# print data to check it's what we want it to be
print(data)
```
## Part One
Each line of our data is a layer in a toboggan slope... It repeats inifinitely out to either side and '#' are trees and '.' are open spaces.
We have a route given to us (down 1, right 3) and we want to return the amount of trees we are given. If we start from (0,0), we will go down hitting (1,3) (2,6) (3,9) etc.
```
length = len(data)
for row in range(length):
col = row * 3
item = data[row][col]
print(item)
```
Since the length of each row is less than 3x the length we will get a string index out of range exception. Since we are starting at a 0 index and we know the length of each row, we can use modulus.
```
length = len(data)
row_length = len(data[0])
for row in range(length):
col = row * 3 % row_length
item = data[row][col]
print(item)
```
Now we make sure to keep track of the count as it increases.
```
length = len(data)
row_length = len(data[0])
tree_count = 0
for row in range(length):
col = (row * 3) % row_length
item = data[row][col]
if item == '#':
tree_count += 1
print(tree_count)
```
## Part Two
Now, we do the same for a group of:
* Right 1, down 1.
* Right 3, down 1. (This is the slope we already checked.)
* Right 5, down 1.
* Right 7, down 1.
* Right 1, down 2.
So we can generalise and pass in a parameter for what the column position. We can address 4/5 requirements by just making this one change!
```
def traverse_path(path_data, right):
length = len(path_data)
row_length = len(path_data[0])
tree_count = 0
for row in range(length):
col = (row * right) % row_length
item = data[row][col]
if item == '#':
tree_count += 1
return tree_count
# Test with the one we already know...
r3_d1 = traverse_path(data, 3)
print(r3_d1)
```
And to figure the "down" motion of right 1, down 2 we can make some adjustments to the row number and divide the length so we don't go down beyond the rows.
```
import math
def traverse_path(path_data, right, down=1):
# Use ceil as to make sure we get the final rows! (due to range taking us up to strictly less than the total)
length = math.ceil(len(path_data) / down)
row_length = len(path_data[0])
tree_count = 0
for row in range(length):
col = (row * right) % row_length
row_num = row * down
item = data[row_num][col]
if item == '#':
tree_count += 1
return tree_count
# Test with the one we already know...
r3_d1 = traverse_path(data, 3)
print(r3_d1)
# and do all the others:
r1_d1 = traverse_path(data, 1)
r5_d1 = traverse_path(data, 5)
r7_d1 = traverse_path(data, 7)
r1_d2 = traverse_path(data, 1, 2)
product = r3_d1 * r1_d1 * r5_d1 * r7_d1 * r1_d2
print(product)
```
#### There we have it...
But perhaps we can go one step further, reduce that repeated code and give the instructions as a dict...
```
instructions = [
{'right':1, 'down':1},
{'right':3, 'down':1},
{'right':5, 'down':1},
{'right':7, 'down':1},
{'right':1, 'down':2}
]
def prod_trav_paths(instrs, path_data):
results = []
for instr in instrs:
res = traverse_path(path_data, instr['right'], instr['down'])
results.append(res)
return math.prod(results)
prod_trav_paths(instructions, data)
```
| github_jupyter |
```
# The usual preamble
%matplotlib inline
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
# Make the graphs a bit prettier, and bigger
plt.style.use('ggplot')
plt.rcParams['figure.figsize'] = (15, 5)
plt.rcParams['font.family'] = 'sans-serif'
# This is necessary to show lots of columns in pandas 0.12.
# Not necessary in pandas 0.13.
pd.set_option('display.width', 5000)
pd.set_option('display.max_columns', 60)
```
One of the main problems with messy data is: how do you know if it's messy or not?
We're going to use the NYC 311 service request dataset again here, since it's big and a bit unwieldy.
```
requests = pd.read_csv('data/311-service-requests.csv', low_memory=False)
```
# 7.1 How do we know if it's messy?
We're going to look at a few columns here. I know already that there are some problems with the zip code, so let's look at that first.
To get a sense for whether a column has problems, I usually use `.unique()` to look at all its values. If it's a numeric column, I'll instead plot a histogram to get a sense of the distribution.
When we look at the unique values in "Incident Zip", it quickly becomes clear that this is a mess.
Some of the problems:
* Some have been parsed as strings, and some as floats
* There are `nan`s
* Some of the zip codes are `29616-0759` or `83`
* There are some N/A values that pandas didn't recognize, like 'N/A' and 'NO CLUE'
What we can do:
* Normalize 'N/A' and 'NO CLUE' into regular nan values
* Look at what's up with the 83, and decide what to do
* Make everything strings
```
requests['Incident Zip'].unique()
```
# 7.2 Fixing the nan values and string/float confusion
We can pass a `na_values` option to `pd.read_csv` to clean this up a little bit. We can also specify that the type of Incident Zip is a string, not a float.
```
na_values = ['NO CLUE', 'N/A', '0']
requests = pd.read_csv('data/311-service-requests.csv', na_values=na_values, dtype={'Incident Zip': str})
requests['Incident Zip'].unique()
```
# 7.3 What's up with the dashes?
```
rows_with_dashes = requests['Incident Zip'].str.contains('-').fillna(False)
len(requests[rows_with_dashes])
requests[rows_with_dashes]
```
I thought these were missing data and originally deleted them like this:
`requests['Incident Zip'][rows_with_dashes] = np.nan`
But then my friend Dave pointed out that 9-digit zip codes are normal. Let's look at all the zip codes with more than 5 digits, make sure they're okay, and then truncate them.
```
long_zip_codes = requests['Incident Zip'].str.len() > 5
requests['Incident Zip'][long_zip_codes].unique()
```
Those all look okay to truncate to me.
```
requests['Incident Zip'] = requests['Incident Zip'].str.slice(0, 5)
```
Done.
Earlier I thought 00083 was a broken zip code, but turns out Central Park's zip code 00083! Shows what I know. I'm still concerned about the 00000 zip codes, though: let's look at that.
```
requests[requests['Incident Zip'] == '00000']
```
This looks bad to me. Let's set these to nan.
```
zero_zips = requests['Incident Zip'] == '00000'
requests.loc[zero_zips, 'Incident Zip'] = np.nan
```
Great. Let's see where we are now:
```
unique_zips = requests['Incident Zip'].unique()
unique_zips
```
Amazing! This is much cleaner. There's something a bit weird here, though -- I looked up 77056 on Google maps, and that's in Texas.
Let's take a closer look:
```
zips = requests['Incident Zip']
# Let's say the zips starting with '0' and '1' are okay, for now. (this isn't actually true -- 13221 is in Syracuse, and why?)
is_close = zips.str.startswith('0') | zips.str.startswith('1')
# There are a bunch of NaNs, but we're not interested in them right now, so we'll say they're False
is_far = ~(is_close) & zips.notnull()
zips[is_far]
requests[is_far][['Incident Zip', 'Descriptor', 'City']].sort_values(by='Incident Zip')
```
Okay, there really are requests coming from LA and Houston! Good to know. Filtering by zip code is probably a bad way to handle this -- we should really be looking at the city instead.
```
requests['City'].str.upper().value_counts()
```
It looks like these are legitimate complaints, so we'll just leave them alone.
# 7.4 Putting it together
Here's what we ended up doing to clean up our zip codes, all together:
```
na_values = ['NO CLUE', 'N/A', '0']
requests = pd.read_csv('data/311-service-requests.csv',
na_values=na_values,
dtype={'Incident Zip': str})
def fix_zip_codes(zips):
# Truncate everything to length 5
zips = zips.str.slice(0, 5)
# Set 00000 zip codes to nan
zero_zips = zips == '00000'
zips[zero_zips] = np.nan
return zips
requests['Incident Zip'] = fix_zip_codes(requests['Incident Zip'])
requests['Incident Zip'].unique()
```
<style>
@font-face {
font-family: "Computer Modern";
src: url('http://mirrors.ctan.org/fonts/cm-unicode/fonts/otf/cmunss.otf');
}
div.cell{
width:800px;
margin-left:16% !important;
margin-right:auto;
}
h1 {
font-family: Helvetica, serif;
}
h4{
margin-top:12px;
margin-bottom: 3px;
}
div.text_cell_render{
font-family: Computer Modern, "Helvetica Neue", Arial, Helvetica, Geneva, sans-serif;
line-height: 145%;
font-size: 130%;
width:800px;
margin-left:auto;
margin-right:auto;
}
.CodeMirror{
font-family: "Source Code Pro", source-code-pro,Consolas, monospace;
}
.text_cell_render h5 {
font-weight: 300;
font-size: 22pt;
color: #4057A1;
font-style: italic;
margin-bottom: .5em;
margin-top: 0.5em;
display: block;
}
.warning{
color: rgb( 240, 20, 20 )
}
| github_jupyter |
```
import boto3
import sagemaker
import time
import pandas as pd
import numpy as np
role = sagemaker.get_execution_role()
region = boto3.Session().region_name
sagemaker_session = sagemaker.Session()
bucket_name = sagemaker_session.default_bucket()
prefix = 'endtoendmlsm'
print(region)
print(role)
print(bucket_name)
s3 = boto3.resource('s3')
file_key = 'data/raw/windturbine_raw_data.csv'
copy_source = {
'Bucket': 'gianpo-public',
'Key': 'endtoendml/{0}'.format(file_key)
}
s3.Bucket(bucket_name).copy(copy_source, '{0}/'.format(prefix) + file_key)
#sagemaker_session.upload_data('/home/ec2-user/SageMaker/windturbine_raw_data_2.csv', bucket=bucket_name, key_prefix='endtoendmlsm/data/raw')
sagemaker_session.download_data(path='.', bucket=bucket_name, key_prefix='endtoendmlsm/data/raw/windturbine_raw_data.csv', extra_args=None)
original_ds = pd.read_csv('./windturbine_raw_data.csv', names = ['turbine_id','turbine_type','wind_speed','rpm_blade','oil_temperature','oil_level','temperature','humidity','vibrations_frequency','pressure','wind_direction','breakdown'])
original_ds.head()
original_ds.describe(include='all')
%matplotlib inline
import matplotlib.pyplot as plt
plt.scatter(original_ds.wind_speed, original_ds.rpm_blade)
plt.hist(original_ds.wind_speed, bins=70)
#wind_speed: gaussian, mean=50, std=30
ws = abs(np.random.normal(50, 30, 1000000)).astype(int)
#temperature: gaussian, mean=20, std=18
temp = abs(np.random.normal(20, 18, 1000000)).astype(int)
#humidity: gaussian, mean=50, std=5
hum = abs(np.random.normal(50, 5, 1000000)).astype(int)
#pressure: gaussian, mean=40, std=25
press = abs(np.random.normal(40, 25, 1000000)).astype(int)
#oil_level: uniform, min=5, max=35
oil_lev = np.random.uniform(5,35,1000000).astype(int)
#rpm_blade: alpha*wind_speed + error
alpha = 5
rpm_blade = abs(alpha*ws + np.random.normal(0,30,1000000)).astype(int)
#vibration_freq: beta*rpm_blade + gamma*pressure + error
beta = 3.5
gamma = 2
vibration_freq = abs(beta*rpm_blade + gamma*press + np.random.normal(0,50,1000000)).astype(int)
#oil_temp: delta*temp + error
#delta = 4.5
#oil_temperature = abs(delta*temp + np.random.normal(0,50,1000000)).astype(int)
#breakdown: k1*rpm_blade + k2*vibration_freq + k3*oil_temp + error
new_dataset = pd.DataFrame()
new_dataset['turbine_id'] = original_ds['turbine_id']
new_dataset['turbine_type'] = original_ds['turbine_type']
new_dataset['wind_direction'] = original_ds['wind_direction']
new_dataset['wind_speed'] = ws
new_dataset['temperature'] = temp
new_dataset['humidity'] = hum
new_dataset['pressure'] = press
new_dataset['oil_level'] = oil_lev
new_dataset['rpm_blade'] = rpm_blade
new_dataset['vibrations_frequency'] = vibration_freq
new_dataset['oil_temperature'] = original_ds['oil_temperature']
new_dataset.describe()
plt.scatter(new_dataset['wind_speed'][:10000], new_dataset['rpm_blade'][:10000])
plt.hist(new_dataset['rpm_blade'][:10000])
from scipy.special import expit
k1=0.0003
k2=0.0005
k3=0.0033
breakdown = k1*rpm_blade + k2*vibration_freq + k3*oil_lev + np.random.normal(0,0.1,1000000)
new_dataset['breakdown_num'] = breakdown
new_dataset.loc[new_dataset['breakdown_num'] <= 0.9, 'breakdown'] = 'no'
new_dataset.loc[new_dataset['breakdown_num'] > 0.9, 'breakdown'] = 'yes'
new_dataset.describe(include='all')
plt.scatter(new_dataset['breakdown'][:10000], new_dataset['rpm_blade'][:10000])
plt.scatter(new_dataset['breakdown'][:10000], new_dataset['vibrations_frequency'][:10000])
final_dataset = new_dataset
final_dataset = final_dataset.drop(columns=['breakdown_num'])
final_dataset.to_csv('windturbine_raw_data.csv', index=False, columns = ['turbine_id','turbine_type','wind_speed','rpm_blade','oil_temperature','oil_level','temperature','humidity','vibrations_frequency','pressure','wind_direction','breakdown'])
sagemaker_session.upload_data('windturbine_raw_data.csv', bucket=bucket_name, key_prefix='endtoendmlsm/data/raw')
```
| github_jupyter |
**Important note:** You should always work on a duplicate of the course notebook. On the page you used to open this, tick the box next to the name of the notebook and click duplicate to easily create a new version of this notebook.
You will get errors each time you try to update your course repository if you don't do this, and your changes will end up being erased by the original course version.
# Welcome to Jupyter Notebooks!
If you want to learn how to use this tool you've come to the right place. This article will teach you all you need to know to use Jupyter Notebooks effectively. You only need to go through Section 1 to learn the basics and you can go into Section 2 if you want to further increase your productivity.
You might be reading this tutorial in a web page (maybe Github or the course's webpage). We strongly suggest to read this tutorial in a (yes, you guessed it) Jupyter Notebook. This way you will be able to actually *try* the different commands we will introduce here.
## Section 1: Need to Know
### Introduction
Let's build up from the basics, what is a Jupyter Notebook? Well, you are reading one. It is a document made of cells. You can write like I am writing now (markdown cells) or you can perform calculations in Python (code cells) and run them like this:
```
1+1
```
Cool huh? This combination of prose and code makes Jupyter Notebook ideal for experimentation: we can see the rationale for each experiment, the code and the results in one comprehensive document. In fast.ai, each lesson is documented in a notebook and you can later use that notebook to experiment yourself.
Other renowned institutions in academy and industry use Jupyter Notebook: Google, Microsoft, IBM, Bloomberg, Berkeley and NASA among others. Even Nobel-winning economists [use Jupyter Notebooks](https://paulromer.net/jupyter-mathematica-and-the-future-of-the-research-paper/) for their experiments and some suggest that Jupyter Notebooks will be the [new format for research papers](https://www.theatlantic.com/science/archive/2018/04/the-scientific-paper-is-obsolete/556676/).
### Writing
A type of cell in which you can write like this is called _Markdown_. [_Markdown_](https://en.wikipedia.org/wiki/Markdown) is a very popular markup language. To specify that a cell is _Markdown_ you need to click in the drop-down menu in the toolbar and select _Markdown_.
Click on the the '+' button on the left and select _Markdown_ from the toolbar.
My first markdown cell
Now you can type your first _Markdown_ cell. Write 'My first markdown cell' and press run.

You should see something like this:
My first markdown cell
Now try making your first _Code_ cell: follow the same steps as before but don't change the cell type (when you add a cell its default type is _Code_). Type something like 3/2. You should see '1.5' as output.
```
3/2
3/2
```
### Modes
If you made a mistake in your *Markdown* cell and you have already ran it, you will notice that you cannot edit it just by clicking on it. This is because you are in **Command Mode**. Jupyter Notebooks have two distinct modes:
1. **Edit Mode**: Allows you to edit a cell's content.
2. **Command Mode**: Allows you to edit the notebook as a whole and use keyboard shortcuts but not edit a cell's content.
You can toggle between these two by either pressing <kbd>ESC</kbd> and <kbd>Enter</kbd> or clicking outside a cell or inside it (you need to double click if its a Markdown cell). You can always know which mode you're on since the current cell has a green border if in **Edit Mode** and a blue border in **Command Mode**. Try it!
### Other Important Considerations
1. Your notebook is autosaved every 120 seconds. If you want to manually save it you can just press the save button on the upper left corner or press <kbd>s</kbd> in **Command Mode**.

2. To know if your kernel is computing or not you can check the dot in your upper right corner. If the dot is full, it means that the kernel is working. If not, it is idle. You can place the mouse on it and see the state of the kernel be displayed.

3. There are a couple of shortcuts you must know about which we use **all** the time (always in **Command Mode**). These are:
<kbd>Shift</kbd>+<kbd>Enter</kbd>: Runs the code or markdown on a cell
<kbd>Up Arrow</kbd>+<kbd>Down Arrow</kbd>: Toggle across cells
<kbd>b</kbd>: Create new cell
<kbd>0</kbd>+<kbd>0</kbd>: Reset Kernel
You can find more shortcuts in the Shortcuts section below.
4. You may need to use a terminal in a Jupyter Notebook environment (for example to git pull on a repository). That is very easy to do, just press 'New' in your Home directory and 'Terminal'. Don't know how to use the Terminal? We made a tutorial for that as well. You can find it [here](https://course.fast.ai/terminal_tutorial.html).

That's it. This is all you need to know to use Jupyter Notebooks. That said, we have more tips and tricks below ↓↓↓
## Section 2: Going deeper
### Markdown formatting
#### Italics, Bold, Strikethrough, Inline, Blockquotes and Links
The five most important concepts to format your code appropriately when using markdown are:
1. *Italics*: Surround your text with '\_' or '\*'
2. **Bold**: Surround your text with '\__' or '\**'
3. `inline`: Surround your text with '\`'
4. > blockquote: Place '\>' before your text.
5. [Links](https://course.fast.ai/): Surround the text you want to link with '\[\]' and place the link adjacent to the text, surrounded with '()'
#### Headings
Notice that including a hashtag before the text in a markdown cell makes the text a heading. The number of hashtags you include will determine the priority of the header ('#' is level one, '##' is level two, '###' is level three and '####' is level four). We will add three new cells with the '+' button on the left to see how every level of heading looks.
Double click on some headings and find out what level they are!
#### Lists
There are three types of lists in markdown.
Ordered list:
1. Step 1
2. Step 1B
3. Step 3
Unordered list
* learning rate
* cycle length
* weight decay
Task list
- [x] Learn Jupyter Notebooks
- [x] Writing
- [x] Modes
- [x] Other Considerations
- [ ] Change the world
Double click on each to see how they are built!
### Code Capabilities
**Code** cells are different than **Markdown** cells in that they have an output cell. This means that we can _keep_ the results of our code within the notebook and share them. Let's say we want to show a graph that explains the result of an experiment. We can just run the necessary cells and save the notebook. The output will be there when we open it again! Try it out by running the next four cells.
```
# Import necessary libraries
from fastai.vision import *
import matplotlib.pyplot as plt
from PIL import Image
a = 1
b = a + 1
c = b + a + 1
d = c + b + a + 1
a, b, c ,d
plt.plot([a,b,c,d])
plt.show()
```
We can also print images while experimenting. I am watching you.
```
Image.open('images/notebook_tutorial/cat_example.jpg')
```
### Running the app locally
You may be running Jupyter Notebook from an interactive coding environment like Gradient, Sagemaker or Salamander. You can also run a Jupyter Notebook server from your local computer. What's more, if you have installed Anaconda you don't even need to install Jupyter (if not, just `pip install jupyter`).
You just need to run `jupyter notebook` in your terminal. Remember to run it from a folder that contains all the folders/files you will want to access. You will be able to open, view and edit files located within the directory in which you run this command but not files in parent directories.
If a browser tab does not open automatically once you run the command, you should CTRL+CLICK the link starting with 'https://localhost:' and this will open a new tab in your default browser.
### Creating a notebook
Click on 'New' in the upper right corner and 'Python 3' in the drop-down list (we are going to use a [Python kernel](https://github.com/ipython/ipython) for all our experiments).

Note: You will sometimes hear people talking about the Notebook 'kernel'. The 'kernel' is just the Python engine that performs the computations for you.
### Shortcuts and tricks
#### Command Mode Shortcuts
There are a couple of useful keyboard shortcuts in `Command Mode` that you can leverage to make Jupyter Notebook faster to use. Remember that to switch back and forth between `Command Mode` and `Edit Mode` with <kbd>Esc</kbd> and <kbd>Enter</kbd>.
<kbd>m</kbd>: Convert cell to Markdown
<kbd>y</kbd>: Convert cell to Code
<kbd>D</kbd>+<kbd>D</kbd>: Delete the cell(if it's not the only cell) or delete the content of the cell and reset cell to Code(if only one cell left)
<kbd>o</kbd>: Toggle between hide or show output
<kbd>Shift</kbd>+<kbd>Arrow up/Arrow down</kbd>: Selects multiple cells. Once you have selected them you can operate on them like a batch (run, copy, paste etc).
<kbd>Shift</kbd>+<kbd>M</kbd>: Merge selected cells.
<kbd>Shift</kbd>+<kbd>Tab</kbd>: [press these two buttons at the same time, once] Tells you which parameters to pass on a function
<kbd>Shift</kbd>+<kbd>Tab</kbd>: [press these two buttons at the same time, three times] Gives additional information on the method
#### Cell Tricks
```
from fastai import*
from fastai.vision import *
```
There are also some tricks that you can code into a cell.
`?function-name`: Shows the definition and docstring for that function
```
?ImageDataBunch
```
`??function-name`: Shows the source code for that function
```
??ImageDataBunch
```
`doc(function-name)`: Shows the definition, docstring **and links to the documentation** of the function
(only works with fastai library imported)
```
doc(ImageDataBunch)
```
#### Line Magics
Line magics are functions that you can run on cells and take as an argument the rest of the line from where they are called. You call them by placing a '%' sign before the command. The most useful ones are:
`%matplotlib inline`: This command ensures that all matplotlib plots will be plotted in the output cell within the notebook and will be kept in the notebook when saved.
`%reload_ext autoreload`, `%autoreload 2`: Reload all modules before executing a new line. If a module is edited, it is not necessary to rerun the import commands, the modules will be reloaded automatically.
These three commands are always called together at the beginning of every notebook.
```
%matplotlib inline
%reload_ext autoreload
%autoreload 2
```
`%timeit`: Runs a line a ten thousand times and displays the average time it took to run it.
```
%timeit [i+1 for i in range(1000)]
```
`%debug`: Allows to inspect a function which is showing an error using the [Python debugger](https://docs.python.org/3/library/pdb.html).
```
for i in range(1000):
a = i+1
b = 'string'
c = b+1
%debug
```
| github_jupyter |
---
title: "Create empty feature groups for Online Feature Store"
date: 2021-04-25
type: technical_note
draft: false
---
```
import json
from pyspark.sql.types import StructField, StructType, StringType, DoubleType, TimestampType, LongType, IntegerType
```
# Create empty feature groups
In this demo example we are expecting to recieve data from Kafka topic, read using spark streaming, do streamig aggregations and ingest aggregated data to feature groups. Thus we will create empy feature groups where we will ingest streaming data.
### Define schema for feature groups
```
card_schema = StructType([StructField('tid', StringType(), True),
StructField('datetime', StringType(), True),
StructField('cc_num', LongType(), True),
StructField('amount', DoubleType(), True)])
schema_10m = StructType([StructField('cc_num', LongType(), True),
StructField('num_trans_per_10m', LongType(), True),
StructField('avg_amt_per_10m', DoubleType(), True),
StructField('stdev_amt_per_10m', DoubleType(), True)])
schema_1h = StructType([StructField('cc_num', LongType(), True),
StructField('num_trans_per_1h', LongType(), True),
StructField('avg_amt_per_1h', DoubleType(), True),
StructField('stdev_amt_per_1h', DoubleType(), True)])
schema_12h = StructType([StructField('cc_num', LongType(), True),
StructField('num_trans_per_12h', LongType(), True),
StructField('avg_amt_per_12h', DoubleType(), True),
StructField('stdev_amt_per_12h', DoubleType(), True)])
```
### Create empty spark dataframes
```
empty_card_df = sqlContext.createDataFrame(sc.emptyRDD(), card_schema)
empty_10m_agg_df = sqlContext.createDataFrame(sc.emptyRDD(), schema_10m)
empty_1h_agg_df = sqlContext.createDataFrame(sc.emptyRDD(), schema_1h)
empty_12h_agg_df = sqlContext.createDataFrame(sc.emptyRDD(), schema_12h)
```
### Establish a connection with your Hopsworks feature store.
```
import hsfs
connection = hsfs.connection()
# get a reference to the feature store, you can access also shared feature stores by providing the feature store name
fs = connection.get_feature_store()
```
### Create feature group metadata objects and save empty spark dataframes to materialise them in hopsworks feature store.
Now We will create each feature group and enable online feature store. Since we are plannig to use these feature groups durring online model serving primary key(s) are required to retrieve feature vector from online feature store.
```
card_transactions = fs.create_feature_group("card_transactions",
version = 1,
online_enabled=False,
statistics_config=False,
primary_key=["tid"])
card_transactions.save(empty_card_df)
card_transactions_10m_agg = fs.create_feature_group("card_transactions_10m_agg",
version = 1,
online_enabled=True,
statistics_config=False,
primary_key=["cc_num"])
card_transactions_10m_agg.save(empty_10m_agg_df)
card_transactions_1h_agg = fs.create_feature_group("card_transactions_1h_agg",
version = 1,
online_enabled=True,
statistics_config=False,
primary_key=["cc_num"])
card_transactions_1h_agg.save(empty_1h_agg_df)
card_transactions_12h_agg = fs.create_feature_group("card_transactions_12h_agg",
version = 1,
online_enabled=True,
statistics_config=False,
primary_key=["cc_num"])
card_transactions_12h_agg.save(empty_12h_agg_df)
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
import seaborn as sns
import nltk
train = pd.read_csv("C:\\Users\\Moaz\\Desktop\\moaz\\Jupyter Python NB\\Machine Hack Practice\\Beer Train Data Set.csv")
test = pd.read_csv("C:\\Users\\Moaz\\Desktop\\moaz\\Jupyter Python NB\\Machine Hack Practice\\Beer Test Data Set.csv")
train.head()
train.info()
train.isnull().sum()
train[["Minimum Temperature", "Maximum Temperature"]]=train["Cellar Temperature"].str.split("-", expand=True, n=1).astype(float)
train[["Minimum Serving Temperature", "Maximum Serving Temperature"]]=train["Serving Temperature"].str.split("-", expand=True, n=1).astype(float)
# Filling empty vaues with MEAN value
avg_abv = train["ABV"].astype("float").mean(axis=0)
train["ABV"].replace(np.nan, avg_abv, inplace=True)
avg_min_temp = train["Minimum Temperature"].astype("float").mean(axis=0)
train["Minimum Temperature"].replace(np.nan, avg_min_temp, inplace=True)
avg_min_temp = train["Maximum Temperature"].astype("float").mean(axis=0)
train["Maximum Temperature"].replace(np.nan, avg_min_temp, inplace=True)
avg_minserv_temp = train["Minimum Serving Temperature"].astype("float").mean(axis=0)
train["Minimum Serving Temperature"].replace(np.nan, avg_minserv_temp, inplace=True)
avg_minserv_temp = train["Maximum Serving Temperature"].astype("float").mean(axis=0)
train["Maximum Serving Temperature"].replace(np.nan, avg_minserv_temp, inplace=True)
train.isnull().sum()
freq = nltk.FreqDist(train['Food Paring'])
for key,value in freq.items():
print(str(key)+' : '+str(value))
train['Food Paring'] = train['Food Paring'].replace("(Curried,Thai)Cheese(pepperyMontereyPepperJack,sharpBlue,Cheddar,pungentGorgonzola,Limburger)Meat(Poultry,Fish,Shellfish,Salmon)" , "Thai, Cheese, Meat" )
train['Food Paring'] = train['Food Paring'].replace("(PanAsian)Cheese(earthyCamembert,Fontina,nuttyAsiago,Colby,Parmesan,tangyBrick,Edam,Feta)General(Salad)Meat(Poultry)" , "Pan-Asian, Cheese, General Food, Meat")
train['Food Paring'] = train['Food Paring'].replace("Meat(Pork,Poultry)" , "Meat")
train['Food Paring'] = train['Food Paring'].replace("(Indian,LatinAmerican,PanAsian)General(Aperitif)" , "Indian, Latin-American, Pan-Asian, General Food")
train['Food Paring'] = train['Food Paring'].replace("Meat(Poultry,Fish,Shellfish)" , "Meat")
train['Food Paring'] = train['Food Paring'].replace("(Italian,German)Cheese(nuttyAsiago,Colby,Parmesan)Meat(Fish,Shellfish,Salmon)" , "Italian, German, Cheese, Meat")
train['Food Paring'] = train['Food Paring'].replace("Cheese(earthyCamembert,Fontina,nuttyAsiago,Colby,Parmesan)Meat(Pork,GrilledMeat)" , "Cheese, Meat")
train['Food Paring'] = train['Food Paring'].replace("Cheese(butteryBrie,Gouda,Havarti,Swiss)General(Chocolate,Digestive)Meat(Beef,SmokedMeat,Game,GrilledMeat)" , "Cheese, General Food, Meat")
train['Food Paring'] = train['Food Paring'].replace("(Barbecue,Indian,LatinAmerican,Thai,PanAsian)Cheese(pepperyMontereyPepperJack)Meat(Shellfish)" , "Barbecue, Indian, Latin-American, Thai, Pan-Asian, Cheese, Meat")
train['Food Paring'] = train['Food Paring'].replace("(Barbecue)Cheese(sharpBlue,Cheddar,pungentGorgonzola,Limburger)Meat(Beef,GrilledMeat)" , "Barbecue, Cheese, Meat")
train['Food Paring'] = train['Food Paring'].replace("(Curried,Thai)Cheese(earthyCamembert,Fontina,nuttyAsiago,Colby,Parmesan,pungentGorgonzola,Limburger)General(Salad,Aperitif)Meat(Poultry,Fish,Shellfish)" , "Thai, Cheese, General Food, Meat")
train['Food Paring'] = train['Food Paring'].replace("(Salad)" , "General Food")
train['Food Paring'] = train['Food Paring'].replace("(Barbecue,Curried,Indian,LatinAmerican,Italian,Thai,Chinese,Japanese,PanAsian,Mediterranean,MiddleEastern)" , "Barbecue, Indian, Latin-American, Italian, Thai, Japanese, Pan-Asian, Mediterranean, Middle-East")
train['Food Paring'] = train['Food Paring'].replace("Cheese(tangyBrick,Edam,Feta)" , "Cheese")
train['Food Paring'] = train['Food Paring'].replace("Cheese(sharpBlue,Cheddar)Meat(Beef,Poultry,Fish)" , "Cheese")
train['Food Paring'] = train['Food Paring'].replace("(Barbecue,German)Cheese(pepperyMontereyPepperJack,sharpBlue,Cheddar)Meat(Beef,SmokedMeat,Game,GrilledMeat,Salmon)" , "Barbecue, German, Cheese, Meat")
train['Food Paring'] = train['Food Paring'].replace("Cheese(butteryBrie,Gouda,Havarti,Swiss)Meat(SmokedMeat,Salmon)" , "Cheese, Meat")
train['Food Paring'] = train['Food Paring'].replace("Cheese(pepperyMontereyPepperJack,pungentGorgonzola,Limburger)General(Salad)" , "Cheese, General Food")
train['Food Paring'] = train['Food Paring'].replace("(German)General(Chocolate,Dessert)Meat(GrilledMeat)" , "German, General Food")
train['Food Paring'] = train['Food Paring'].replace("(Curried,Indian)Cheese(nuttyAsiago,Colby,Parmesan,sharpBlue,Cheddar)Meat(Shellfish)" , "Indian, Cheese, Meat")
train['Food Paring'] = train['Food Paring'].replace("(Barbecue,LatinAmerican)Cheese(earthyCamembert,Fontina)General(Chocolate,Dessert)Meat(Beef,Shellfish,SmokedMeat,GrilledMeat)" , "Barbecue, Latin-American, Cheese, General Food, Meat")
train['Food Paring'] = train['Food Paring'].replace("(German)" , "German")
train['Food Paring'] = train['Food Paring'].replace("(Barbecue)Cheese(pepperyMontereyPepperJack,sharpBlue,Cheddar)Meat(Beef,SmokedMeat,Game,GrilledMeat,Salmon)" , "Barbecue, Cheese, Meat")
train['Food Paring'] = train['Food Paring'].replace("Cheese(sharpBlue,Cheddar,pungentGorgonzola,Limburger)General(Digestive)" , "Cheese, General Food")
train['Food Paring'] = train['Food Paring'].replace("(Barbecue,Italian)Cheese(earthyCamembert,Fontina)Meat(Pork,Poultry,Fish,Shellfish)" , "Barbecue, Italian, Cheese, Meat")
train['Food Paring'] = train['Food Paring'].replace("(Curried)Cheese(nuttyAsiago,Colby,Parmesan,pepperyMontereyPepperJack)Meat(Poultry,Fish)" , "Cheese, Meat")
train['Food Paring'] = train['Food Paring'].replace("(German)Cheese(tangyBrick,Edam,Feta)General(Salad)Meat(Poultry,Fish,Shellfish)" , "German, Cheese, General Food, Meat")
train['Food Paring'] = train['Food Paring'].replace("Cheese(butteryBrie,Gouda,Havarti,Swiss,pungentGorgonzola,Limburger)General(Chocolate)Meat(Beef)" , "Cheese, General Food, Meat")
train['Food Paring'] = train['Food Paring'].replace("(Barbecue)Cheese(butteryBrie,Gouda,Havarti,Swiss)General(Chocolate,Dessert)Meat(Beef,SmokedMeat,GrilledMeat)" , "Barbecue, Cheese, General Food, Meat")
train['Food Paring'] = train['Food Paring'].replace("(Thai)Cheese(tangyBrick,Edam,Feta)General(Salad,Aperitif)Meat(Fish)" , "Thai, Cheese, General Food, Meat")
train['Food Paring'] = train['Food Paring'].replace("(LatinAmerican,German)Meat(Pork,Poultry)" , "Latin-American, German, Meat")
train['Food Paring'] = train['Food Paring'].replace("(Barbecue)Cheese(butteryBrie,Gouda,Havarti,Swiss,earthyCamembert,Fontina)General(Chocolate,Dessert)Meat(Beef,Shellfish,SmokedMeat,Game,GrilledMeat)" , "Barbecue, Cheese, General Food, Meat")
train['Food Paring'] = train['Food Paring'].replace("Cheese(butteryBrie,Gouda,Havarti,Swiss,sharpBlue,Cheddar)General(Digestive)Meat(Beef,SmokedMeat,Game)" , "Cheese, General Food")
train['Food Paring'] = train['Food Paring'].replace("(Dessert,Aperitif,Digestive)" , "Dessert")
train['Food Paring'] = train['Food Paring'].replace("(Barbecue)Cheese(pepperyMontereyPepperJack,sharpBlue,Cheddar,pungentGorgonzola,Limburger)Meat(Game,GrilledMeat,Salmon)" , "Barbecue, Cheese, Meat")
train['Food Paring'] = train['Food Paring'].replace("(German)Cheese(butteryBrie,Gouda,Havarti,Swiss,sharpBlue,Cheddar)General(Salad,Aperitif)" , "German, Cheese, General Food")
train['Food Paring'] = train['Food Paring'].replace("(German)Cheese(butteryBrie,Gouda,Havarti,Swiss,earthyCamembert,Fontina)General(Chocolate)Meat(Game)" , "German, Cheese, General Food")
train['Food Paring'] = train['Food Paring'].replace("(Aperitif,Digestive)Meat(Game,Salmon)" , "Dessert, Meat")
train['Food Paring'] = train['Food Paring'].replace("(Barbecue)Cheese(earthyCamembert,Fontina,nuttyAsiago,Colby,Parmesan)General(Chocolate)Meat(Beef)" , "Barbecue, Cheese, General Food")
train['Food Paring'] = train['Food Paring'].replace("(Indian,MiddleEastern)Cheese(nuttyAsiago,Colby,Parmesan,tangyBrick,Edam,Feta)General(Salad,Aperitif)Meat(Fish,Shellfish)" , "Indian, Middle-East, Cheese, General Food, Meat")
train['Food Paring'] = train['Food Paring'].replace("(German)Cheese(earthyCamembert,Fontina)General(Chocolate)Meat(Game)" , "German, Cheese, General Food, Meat")
train['Food Paring'] = train['Food Paring'].replace("Cheese(pepperyMontereyPepperJack,tangyBrick,Edam,Feta)General(Salad)Meat(Poultry,Fish,Shellfish)" , "Cheese, General Food, Meat")
train['Food Paring'] = train['Food Paring'].replace("(Barbecue,LatinAmerican)General(Chocolate)Meat(SmokedMeat,GrilledMeat)" , "Barbecue, Latin-American, General Food, Meat")
train['Food Paring'] = train['Food Paring'].replace("(German)Cheese(butteryBrie,Gouda,Havarti,Swiss)General(Salad)Meat(Pork,Fish,Shellfish)" , "German, Cheese, General Food, Meat")
train['Food Paring'] = train['Food Paring'].replace("(Dessert)Meat(Poultry)" , "Dessert, Meat")
train['Food Paring'] = train['Food Paring'].replace("(German)General(Salad)Meat(Fish)" , "German, General Food, Meat")
train['Food Paring'] = train['Food Paring'].replace("None,yet" , "None yet")
train['Food Paring'] = train['Food Paring'].replace("(Mediterranean)Cheese(sharpBlue,Cheddar,pungentGorgonzola,Limburger)Meat(Pork,Poultry)" , "Mediterranean, Cheese, Meat")
train['Food Paring'] = train['Food Paring'].replace("(Barbecue,Curried,Indian,LatinAmerican,Chinese)Cheese(sharpBlue,Cheddar)General(Aperitif,Digestive)Meat(Shellfish,Game)" , "Barbecue, Indian, Latin-American, Chinese, Cheese, General Food, Meat")
train['Food Paring'] = train['Food Paring'].replace("(German)Cheese(pepperyMontereyPepperJack)General(Salad)Meat(Pork)" , "German, Cheese, General Food, Meat")
train['Food Paring'] = train['Food Paring'].replace("(Italian,MiddleEastern)Cheese(pepperyMontereyPepperJack)General(Salad)Meat(Fish)" , "Italian, Middle-East, Cheese, General Food, Meat")
train['Food Paring'] = train['Food Paring'].replace("(Japanese,German)Cheese(pepperyMontereyPepperJack)General(Aperitif)Meat(Poultry,Fish)" , "Japanese, German, Cheese, General Food, Meat")
train['Food Paring'] = train['Food Paring'].replace("(LatinAmerican,German)Meat(Beef,SmokedMeat,GrilledMeat)" , "Latin-American, German, Meat")
train['Food Paring'] = train['Food Paring'].replace("(Chocolate,Salad,Dessert,Aperitif)" , "Dessert")
train['Food Paring'] = train['Food Paring'].replace("(Barbecue)Cheese(earthyCamembert,Fontina)Meat(Beef,SmokedMeat,Game,GrilledMeat)" , "Barbecue, Cheese, Meat")
train['Food Paring'] = train['Food Paring'].replace("(German)General(Salad)Meat(Pork,Fish,Shellfish)" , "German, General Food, Meat")
train['Food Paring'] = train['Food Paring'].replace("(German)Cheese(butteryBrie,Gouda,Havarti,Swiss)General(Aperitif)Meat(Pork,Poultry,Fish,Shellfish)" , "German, Cheese, General Food, Meat")
train['Food Paring'] = train['Food Paring'].replace("Cheese(earthyCamembert,Fontina,sharpBlue,Cheddar)Meat(GrilledMeat)" , "Cheese, Meat")
train['Food Paring'] = train['Food Paring'].replace("(Curried,Indian,Thai,Chinese,Japanese,PanAsian)Cheese(sharpBlue,Cheddar)" , "Indian, Thai, Chinese, Japanese, Pan-Aisan, Cheese")
train['Food Paring'] = train['Food Paring'].replace("Cheese(sharpBlue,Cheddar,pungentGorgonzola,Limburger)General(Dessert,Digestive)" , "Cheese, General Food")
train['Food Paring'] = train['Food Paring'].replace("(LatinAmerican)Meat(Beef,Poultry)" , "Latin-American, Meat")
train['Food Paring'] = train['Food Paring'].replace("(German)Meat(SmokedMeat,Game,GrilledMeat)" , "German, Meat ")
train['Food Paring'] = train['Food Paring'].replace("Cheese(nuttyAsiago,Colby,Parmesan)General(Digestive)" , "Cheese, General Food")
train['Food Paring'] = train['Food Paring'].replace("(Barbecue)Cheese(pepperyMontereyPepperJack,sharpBlue,Cheddar,tangyBrick,Edam,Feta)General(Salad)Meat(Pork,Poultry,Fish,Shellfish)" , "Barbecue, Cheese, General Food, Meat")
train['Food Paring'] = train['Food Paring'].replace("(Indian,Mediterranean,MiddleEastern)Cheese(pepperyMontereyPepperJack,sharpBlue,Cheddar)Meat(Fish,Shellfish)" , "Indian, Mediterranean, Middle-East, Cheese, Meat")
train['Food Paring'] = train['Food Paring'].replace("Cheese(sharpBlue,Cheddar,pungentGorgonzola,Limburger,tangyBrick,Edam,Feta)General(Salad)" , "Cheese, General")
train['Food Paring'] = train['Food Paring'].replace("(German)Cheese(earthyCamembert,Fontina)Meat(SmokedMeat,Game,GrilledMeat)" , "German, Cheese, Meat")
train['Food Paring'] = train['Food Paring'].replace("Cheese(earthyCamembert,Fontina)General(Chocolate)Meat(GrilledMeat)" , "Cheese, General Food")
train['Food Paring'] = train['Food Paring'].replace("Cheese(sharpBlue,Cheddar,tangyBrick,Edam,Feta)General(Aperitif,Digestive)" , "Cheese, General Food")
train['Food Paring'] = train['Food Paring'].replace("Cheese(pepperyMontereyPepperJack)General(Chocolate)Meat(GrilledMeat)" , "Cheese, General Food, Meat")
train['Food Paring'] = train['Food Paring'].replace("(Curried,German)Cheese(nuttyAsiago,Colby,Parmesan)General(Digestive)Meat(Salmon)" , "German, Cheese, General Food")
train['Food Paring'] = train['Food Paring'].replace("(German)Cheese(butteryBrie,Gouda,Havarti,Swiss,sharpBlue,Cheddar)General(Dessert,Digestive)" , "German, Cheese, General Food")
train['Food Paring'] = train['Food Paring'].replace("Cheese(earthyCamembert,Fontina)General(Aperitif)" , "Cheese, General Food")
train['Food Paring'] = train['Food Paring'].replace("(German)General(Salad)Meat(Poultry,Fish)" , "German, General Food, Meat")
train['Food Paring'] = train['Food Paring'].replace("(Japanese)" , "Japanese")
train['Food Paring'] = train['Food Paring'].replace("(German)Cheese(sharpBlue,Cheddar)General(Salad)Meat(Pork)" , "German, Cheese, General Food, Meat")
train['Food Paring'] = train['Food Paring'].replace("Cheese(pungentGorgonzola,Limburger,tangyBrick,Edam,Feta)General(Digestive)Meat(Shellfish,Game,GrilledMeat)" , "Cheese, General Food, Meat")
train['Food Paring'] = train['Food Paring'].replace("(Barbecue,LatinAmerican)Cheese(pepperyMontereyPepperJack)Meat(Fish,SmokedMeat)" , "Barbecue, Latin-American, Cheese, Meat")
train['Food Paring'] = train['Food Paring'].replace("(Salad)Meat(Poultry,Game)" , "General Food, Meat")
train['Food Paring'] = train['Food Paring'].replace("(Curried,Thai,PanAsian)Cheese(sharpBlue,Cheddar)Meat(Game,GrilledMeat)" , "Thai, Cheese, Meat")
train['Food Paring'] = train['Food Paring'].replace("Cheese(butteryBrie,Gouda,Havarti,Swiss,pungentGorgonzola,Limburger)General(Dessert,Digestive)Meat(Game)" , "Cheese, General Food, Meat")
train['Food Paring'] = train['Food Paring'].replace("Cheese(butteryBrie,Gouda,Havarti,Swiss,sharpBlue,Cheddar,pungentGorgonzola,Limburger)" , "Cheese")
train['Food Paring'] = train['Food Paring'].replace("(Aperitif)Meat(Fish,Shellfish,Salmon)" , "Dessert, Meat")
train['Food Paring'] = train['Food Paring'].replace("(Thai,Chinese,Japanese,PanAsian)Meat(Pork,Poultry,Fish,Shellfish)" , "Thai, Chinese, Japanese, Pan-Asian, Meat")
train['Food Paring'] = train['Food Paring'].replace("(Barbecue,LatinAmerican)Cheese(nuttyAsiago,Colby,Parmesan)General(Chocolate)Meat(Salmon)" , "Barbecue, Latin-American, Geberal Food, Meat")
train['Food Paring'] = train['Food Paring'].replace("Cheese(sharpBlue,Cheddar,pungentGorgonzola,Limburger)General(Aperitif,Digestive)" , "Cheese, General Food")
train['Food Paring'] = train['Food Paring'].replace("(Dessert,Aperitif)" , "Dessert")
train['Food Paring'] = train['Food Paring'].replace("(Chocolate,Salad,Dessert,Apritif)" , "Dessert")
train['Food Paring'].nunique()
freq = nltk.FreqDist(train['Glassware Used'])
for key,value in freq.items():
print(str(key)+' : '+str(value))
train['Glassware Used'] = train['Glassware Used'].replace('PintGlass(orBecker,Nonic,Tumbler),Mug(orSeidel,Stein)','Pint Glass, Mug')
train['Glassware Used'] = train['Glassware Used'].replace('PintGlass(orBecker,Nonic,Tumbler),PilsenerGlass(orPokal),Mug(orSeidel,Stein)','Pint Glass, Pilsener Glass, Mug')
train['Glassware Used'] = train['Glassware Used'].replace('PilsenerGlass(orPokal)','Pilsener Glass')
train['Glassware Used'] = train['Glassware Used'].replace('Flute,PilsenerGlass(orPokal),Mug(orSeidel,Stein)','Flute, Pint Glass, Mug')
train['Glassware Used'] = train['Glassware Used'].replace('PintGlass(orBecker,Nonic,Tumbler),Snifter,OversizedWineGlass','Pint Glass, Snifter, Over-sized Wine Glass')
train['Glassware Used'] = train['Glassware Used'].replace('PintGlass(orBecker,Nonic,Tumbler),PilsenerGlass(orPokal)','Pint Glass, Pilsener Glass')
train['Glassware Used'] = train['Glassware Used'].replace('Snifter,Tulip,Goblet(orChalice),OversizedWineGlass','Snifter, Tulip, Goblet, Over-sized Wine Glass')
train['Glassware Used'] = train['Glassware Used'].replace('PintGlass(orBecker,Nonic,Tumbler),Tulip,OversizedWineGlass','Pint Glass, Tulip, Over-sized Wine Glass')
train['Glassware Used'] = train['Glassware Used'].replace('Mug(orSeidel,Stein),Stange(SlenderCylinder)','Mug, Stange')
train['Glassware Used'] = train['Glassware Used'].replace('PintGlass(orBecker,Nonic,Tumbler),Snifter,Tulip','Pint Glass, Nonic, Tumbler')
train['Glassware Used'] = train['Glassware Used'].replace('Flute,Tulip,OversizedWineGlass','Flute, Tulip, Over-sized Wine Glass')
train['Glassware Used'] = train['Glassware Used'].replace('Flute,WeizenGlass','Flute, Weizen Glass')
train['Glassware Used'] = train['Glassware Used'].replace('Flute,PilsenerGlass(orPokal)','Flute, Pilsener Glass')
train['Glassware Used'] = train['Glassware Used'].replace('PintGlass(orBecker,Nonic,Tumbler)','Pint Glass')
train['Glassware Used'] = train['Glassware Used'].replace('WeizenGlass','Weizen Glass')
train['Glassware Used'] = train['Glassware Used'].replace('Goblet(orChalice)','Goblet')
train['Glassware Used'] = train['Glassware Used'].replace('Snifter,Tulip,OversizedWineGlass','Snifter, Tulip, Over-sized Wine Glass')
train['Glassware Used'] = train['Glassware Used'].replace('Snifter,Tulip,Goblet(orChalice)','Snifter, Tulip, Goblet')
train['Glassware Used'] = train['Glassware Used'].replace('Mug(orSeidel,Stein)','Mug')
train['Glassware Used'] = train['Glassware Used'].replace('PintGlass(orBecker,Nonic,Tumbler),Goblet(orChalice)','Pint Glass, Goblet')
train['Glassware Used'] = train['Glassware Used'].replace('PilsenerGlass(orPokal),Mug(orSeidel,Stein)','Pilsener Glass, Mug')
train['Glassware Used'] = train['Glassware Used'].replace('Tulip,OversizedWineGlass','Tulip, Over-sized Wine Glass')
train['Glassware Used'] = train['Glassware Used'].replace('Flute,PilsenerGlass(orPokal),Mug(orSeidel,Stein),Stange(SlenderCylinder)','Flute, Mug, Stange')
train['Glassware Used'] = train['Glassware Used'].replace('Stange(SlenderCylinder),WeizenGlass','Stange, Weizen Glass')
train['Glassware Used'] = train['Glassware Used'].replace('Snifter,Goblet(orChalice)','Snifter, Goblet')
train['Glassware Used'] = train['Glassware Used'].replace('PintGlass(orBecker,Nonic,Tumbler),Stange(SlenderCylinder)','Pint Glass, Stange')
train['Glassware Used'] = train['Glassware Used'].replace('Flute,Snifter,Tulip,Stange(SlenderCylinder)','Flute, Snifter, Tulip, Stange ')
train['Glassware Used'] = train['Glassware Used'].replace('Stange(SlenderCylinder)','Stange ')
train['Glassware Used'] = train['Glassware Used'].replace('PintGlass(orBecker,Nonic,Tumbler),Mug(orSeidel,Stein),OversizedWineGlass','Pint Glass, Mug, Over-sized Wine Glass')
train['Glassware Used'] = train['Glassware Used'].replace('Flute,Snifter,Tulip','Flute, Snifter, Tulip ')
train['Glassware Used'] = train['Glassware Used'].replace('PintGlass(orBecker,Nonic,Tumbler),Snifter','Pint Glass, Snifter ')
train['Glassware Used'] = train['Glassware Used'].replace('PintGlass(orBecker,Nonic,Tumbler),Snifter,Mug(orSeidel,Stein)','Pint Glass, Snifter, Mug ')
train['Glassware Used'] = train['Glassware Used'].replace('Tulip,Goblet(orChalice),OversizedWineGlass','Tulip, Over-sized Wine Glass')
train['Glassware Used'] = train['Glassware Used'].replace('None,yet','None yet ')
train['Glassware Used'] = train['Glassware Used'].replace('Flute,Snifter,OversizedWineGlass','Flute, Snifter, Over-sized Wine Glass ')
train['Glassware Used'] = train['Glassware Used'].replace('Snifter,OversizedWineGlass','Snifter, Over-sized Wine Glass ')
train['Glassware Used'] = train['Glassware Used'].replace('Flute','Flute ')
train['Glassware Used'] = train['Glassware Used'].replace('PintGlass(orBecker,Nonic,Tumbler),Mug(orSeidel,Stein),WeizenGlass','Pint Glass, Mug, Weizen Glass ')
train['Glassware Used'] = train['Glassware Used'].replace('Flute,Stange(SlenderCylinder)','Flute, Stange ')
train['Glassware Used'].nunique()
train['Style Name'].nunique()
from sklearn.preprocessing import LabelEncoder
label_encoder = LabelEncoder()
train['Food Paring label']= label_encoder.fit_transform(train['Food Paring'])
train['Glassware Used label']= label_encoder.fit_transform(train['Glassware Used'])
train['Style Name label']= label_encoder.fit_transform(train['Style Name'])
train['Ratings'] = pd.to_numeric(train['Ratings'],errors='coerce')
train['Beer Name'] = train['Beer Name'].astype(float)
train['Brewing Company'] = train['Brewing Company'].astype(float)
train.head()
train.dtypes
train1 = train[['ABV', 'Ratings', 'Minimum Temperature','Maximum Temperature','Minimum Serving Temperature','Maximum Serving Temperature', 'Food Paring label', 'Glassware Used label', 'Style Name label', 'Score']]
train1.isnull().sum()
# Replace empty values by mean rating values
avg_rating = train1["Ratings"].astype("float").mean(axis=0)
train1["Ratings"].replace(np.nan, avg_rating, inplace=True)
train1.isnull().sum()
sns.set(style="ticks", color_codes=True)
sns.pairplot(train1)
#A simple correlation plot usong seaborn. The below plot shows how the different variables correlate with each other
corr = train1.corr()
fig, ax = plt.subplots(figsize=(10,10))
ax = sns.heatmap(
corr,
vmin=-1, vmax=1 , center=2,
square=True,
annot=True,
linewidths=.5,
cmap="YlGnBu" )
#Rotating labels on x axis
ax.set_xticklabels(
ax.get_xticklabels(),
rotation=55,
horizontalalignment='right'
)
```
## TEST SET
```
test.head()
test.isnull().sum()
test[["Minimum Temperature", "Maximum Temperature"]] = test["Cellar Temperature"].str.split("-", expand=True, n=1).astype(float)
test[["Minimum Serving Temperature", "Maximum Serving Temperature"]] = test["Serving Temperature"].str.split("-", expand=True, n=1).astype(float)
avg_abv1 = test["ABV"].astype("float").mean(axis=0)
test["ABV"].replace(np.nan, avg_abv1, inplace=True)
avg_min_temp1 = test["Minimum Temperature"].astype("float").mean(axis=0)
test["Minimum Temperature"].replace(np.nan, avg_min_temp1, inplace=True)
avg_max_temp1 = test["Maximum Temperature"].astype("float").mean(axis=0)
test["Maximum Temperature"].replace(np.nan, avg_max_temp1, inplace=True)
avg_minserv_temp1 = test["Minimum Serving Temperature"].astype("float").mean(axis=0)
test["Minimum Serving Temperature"].replace(np.nan, avg_minserv_temp1, inplace=True)
avg_maxserv_temp1 = test["Maximum Serving Temperature"].astype("float").mean(axis=0)
test["Maximum Serving Temperature"].replace(np.nan, avg_maxserv_temp1, inplace=True)
freq = nltk.FreqDist(test['Food Paring'])
for key,value in freq.items():
print(str(key)+' : '+str(value))
test['Food Paring'] = test['Food Paring'].replace("(Curried,Thai)Cheese(pepperyMontereyPepperJack,sharpBlue,Cheddar,pungentGorgonzola,Limburger)Meat(Poultry,Fish,Shellfish,Salmon)" , "Thai, Cheese, Meat")
test['Food Paring'] = test['Food Paring'].replace("(Barbecue)Cheese(butteryBrie,Gouda,Havarti,Swiss)General(Chocolate,Dessert)Meat(Beef,SmokedMeat,GrilledMeat)" , "Barbecue, Cheese, General Food, Meat")
test['Food Paring'] = test['Food Paring'].replace("Cheese(earthyCamembert,Fontina)General(Aperitif)" , "Cheese, General Food")
test['Food Paring'] = test['Food Paring'].replace("(LatinAmerican,German)Meat(Pork,Poultry)" , "Meat")
test['Food Paring'] = test['Food Paring'].replace("(Barbecue)Cheese(pepperyMontereyPepperJack,sharpBlue,Cheddar,pungentGorgonzola,Limburger)Meat(Game,GrilledMeat,Salmon)" , "Barbecue, Cheese, Meat")
test['Food Paring'] = test['Food Paring'].replace("Cheese(butteryBrie,Gouda,Havarti,Swiss)General(Chocolate,Digestive)Meat(Beef,SmokedMeat,Game,GrilledMeat)" , "Cheese, General Food, Meat")
test['Food Paring'] = test['Food Paring'].replace("Meat(Poultry,Fish,Shellfish)" , "Meat")
test['Food Paring'] = test['Food Paring'].replace("Cheese(pepperyMontereyPepperJack,pungentGorgonzola,Limburger)General(Salad)" , "Cheese, General Food")
test['Food Paring'] = test['Food Paring'].replace("(Dessert)Meat(Poultry)" , "Dessert, Meat")
test['Food Paring'] = test['Food Paring'].replace("(Curried,Thai)Cheese(earthyCamembert,Fontina,nuttyAsiago,Colby,Parmesan,pungentGorgonzola,Limburger)General(Salad,Aperitif)Meat(Poultry,Fish,Shellfish)" , "Thai, Cheese, General Food, Meat")
test['Food Paring'] = test['Food Paring'].replace("(Curried,Indian,Thai,Chinese,Japanese,PanAsian)Cheese(sharpBlue,Cheddar)" , "Indian, Thai, Chinese, Japanese, PanAsian, Cheese")
test['Food Paring'] = test['Food Paring'].replace("(PanAsian)Cheese(earthyCamembert,Fontina,nuttyAsiago,Colby,Parmesan,tangyBrick,Edam,Feta)General(Salad)Meat(Poultry)" , "PanAsian, Cheese, General Food, Meat")
test['Food Paring'] = test['Food Paring'].replace("(Indian,Mediterranean,MiddleEastern)Cheese(pepperyMontereyPepperJack,sharpBlue,Cheddar)Meat(Fish,Shellfish)" , "Indian, Mediterranean, MiddleEastern, Cheese, Meat")
test['Food Paring'] = test['Food Paring'].replace("Cheese(sharpBlue,Cheddar,pungentGorgonzola,Limburger)General(Digestive)" , "Cheese, General Food")
test['Food Paring'] = test['Food Paring'].replace("Cheese(sharpBlue,Cheddar,pungentGorgonzola,Limburger)General(Dessert,Digestive)" , "Cheese, General Food")
test['Food Paring'] = test['Food Paring'].replace("Cheese(sharpBlue,Cheddar)Meat(Beef,Poultry,Fish)" , "Cheese, Meat")
test['Food Paring'] = test['Food Paring'].replace("(Salad)" , "Salad")
test['Food Paring'] = test['Food Paring'].replace("(Italian,German)Cheese(nuttyAsiago,Colby,Parmesan)Meat(Fish,Shellfish,Salmon)" , "Italian, German, Cheese, Meat")
test['Food Paring'] = test['Food Paring'].replace("Cheese(pepperyMontereyPepperJack,tangyBrick,Edam,Feta)General(Salad)Meat(Poultry,Fish,Shellfish)" , "Cheese, General Food, Meat")
test['Food Paring'] = test['Food Paring'].replace("(German)" , "German")
test['Food Paring'] = test['Food Paring'].replace("(Thai)Cheese(tangyBrick,Edam,Feta)General(Salad,Aperitif)Meat(Fish)" , "Thai, Cheese, General Food, Meat")
test['Food Paring'] = test['Food Paring'].replace("(Curried,Indian)Cheese(nuttyAsiago,Colby,Parmesan,sharpBlue,Cheddar)Meat(Shellfish)" , "Indian, Cheese, Meat")
test['Food Paring'] = test['Food Paring'].replace("(Barbecue)Cheese(butteryBrie,Gouda,Havarti,Swiss,earthyCamembert,Fontina)General(Chocolate,Dessert)Meat(Beef,Shellfish,SmokedMeat,Game,GrilledMeat)" , "Barbecue, Cheese, General Food, Meat")
test['Food Paring'] = test['Food Paring'].replace("Cheese(pepperyMontereyPepperJack)General(Chocolate)Meat(GrilledMeat)" , "Cheese, General Food, Meat")
test['Food Paring'] = test['Food Paring'].replace("(German)Cheese(butteryBrie,Gouda,Havarti,Swiss,earthyCamembert,Fontina)General(Chocolate)Meat(Game)" , "German, Cheese, General Food, Meat")
test['Food Paring'] = test['Food Paring'].replace("(German)Cheese(earthyCamembert,Fontina)Meat(SmokedMeat,Game,GrilledMeat)" , "German, Cheese, Meat")
test['Food Paring'] = test['Food Paring'].replace("(Dessert,Aperitif,Digestive)" , "Dessert")
test['Food Paring'] = test['Food Paring'].replace("(Indian,LatinAmerican,PanAsian)General(Aperitif)" , "Indian, LatinAmerican, PanAsian, General Food")
test['Food Paring'] = test['Food Paring'].replace("Cheese(butteryBrie,Gouda,Havarti,Swiss,pungentGorgonzola,Limburger)General(Chocolate)Meat(Beef)" , "Cheese, General Food, Meat")
test['Food Paring'] = test['Food Paring'].replace("Cheese(nuttyAsiago,Colby,Parmesan)General(Digestive)" , "Cheese, General Food")
test['Food Paring'] = test['Food Paring'].replace("(German)Cheese(butteryBrie,Gouda,Havarti,Swiss,sharpBlue,Cheddar)General(Salad,Aperitif)" , "German, Cheese, General Food")
test['Food Paring'] = test['Food Paring'].replace("(German)Cheese(tangyBrick,Edam,Feta)General(Salad)Meat(Poultry,Fish,Shellfish)" , "German, Cheese, Meat")
test['Food Paring'] = test['Food Paring'].replace("(Indian,MiddleEastern)Cheese(nuttyAsiago,Colby,Parmesan,tangyBrick,Edam,Feta)General(Salad,Aperitif)Meat(Fish,Shellfish)" , "Indian, MiddleEastern, Cheese, General Food, Meat")
test['Food Paring'] = test['Food Paring'].replace("(German)Cheese(earthyCamembert,Fontina)General(Chocolate)Meat(Game)" , "German, Cheese, General Food, Meat")
test['Food Paring'] = test['Food Paring'].replace("(Italian,MiddleEastern)Cheese(pepperyMontereyPepperJack)General(Salad)Meat(Fish)" , "Italian, MiddleEastern, Cheese, General Food, Meat")
test['Food Paring'] = test['Food Paring'].replace("(German)General(Salad)Meat(Pork,Fish,Shellfish)" , "German, General Food, Meat")
test['Food Paring'] = test['Food Paring'].replace("(Barbecue)Cheese(earthyCamembert,Fontina,nuttyAsiago,Colby,Parmesan)General(Chocolate)Meat(Beef)" , "Barbecue, Cheese, General Food, Meat")
test['Food Paring'] = test['Food Paring'].replace("(Aperitif,Digestive)Meat(Game,Salmon)" , "Meat")
test['Food Paring'] = test['Food Paring'].replace("(Barbecue,Italian)Cheese(earthyCamembert,Fontina)Meat(Pork,Poultry,Fish,Shellfish)" , "Barbecue, Italian, Cheese, Meat")
test['Food Paring'] = test['Food Paring'].replace("(Barbecue,Curried,Indian,LatinAmerican,Italian,Thai,Chinese,Japanese,PanAsian,Mediterranean,MiddleEastern)" , "Barbecue, Curried, Indian, LatinAmerican, Italian, Thai, Chinese, Japanese, PanAsian, Mediterranean, MiddleEastern")
test['Food Paring'] = test['Food Paring'].replace("(Barbecue)Cheese(pepperyMontereyPepperJack,sharpBlue,Cheddar,tangyBrick,Edam,Feta)General(Salad)Meat(Pork,Poultry,Fish,Shellfish)" , "Barbecue, Cheese, General Food, Meat")
test['Food Paring'] = test['Food Paring'].replace("(German)Meat(SmokedMeat,Game,GrilledMeat)" , "German, Meat")
test['Food Paring'] = test['Food Paring'].replace("(Japanese,German)Cheese(pepperyMontereyPepperJack)General(Aperitif)Meat(Poultry,Fish)" , "Japanese, German, Cheese, General Food, Meat")
test['Food Paring'] = test['Food Paring'].replace("(Barbecue)Cheese(earthyCamembert,Fontina)Meat(Beef,SmokedMeat,Game,GrilledMeat)" , "Barbecue, Cheese, Meat")
test['Food Paring'] = test['Food Paring'].replace("Cheese(butteryBrie,Gouda,Havarti,Swiss)Meat(SmokedMeat,Salmon)" , "Cheese, Meat")
test['Food Paring'] = test['Food Paring'].replace("(Mediterranean)Cheese(sharpBlue,Cheddar,pungentGorgonzola,Limburger)Meat(Pork,Poultry)" , "Mediterranean, Cheese, Meat")
test['Food Paring'] = test['Food Paring'].replace("(German)Cheese(pepperyMontereyPepperJack)General(Salad)Meat(Pork)" , "German, Cheese, General Food, Meat")
test['Food Paring'] = test['Food Paring'].replace("(Barbecue,Curried,Indian,LatinAmerican,Chinese)Cheese(sharpBlue,Cheddar)General(Aperitif,Digestive)Meat(Shellfish,Game)" , "Barbecue, Curried, Indian, LatinAmerican, Chinese, Cheese, General Food, Meat")
test['Food Paring'] = test['Food Paring'].replace("(Curried,Thai,PanAsian)Cheese(sharpBlue,Cheddar)Meat(Game,GrilledMeat)" , "Curried, Thai, PanAsian, Cheese, Meat")
test['Food Paring'] = test['Food Paring'].replace("None,yet" , "None")
test['Food Paring'] = test['Food Paring'].replace("Cheese(tangyBrick,Edam,Feta)" , "Cheese")
test['Food Paring'] = test['Food Paring'].replace("(Barbecue,LatinAmerican)Cheese(earthyCamembert,Fontina)General(Chocolate,Dessert)Meat(Beef,Shellfish,SmokedMeat,GrilledMeat)" , "Barbecue, LatinAmerican, Cheese, General Food, Meat")
test['Food Paring'] = test['Food Paring'].replace("Cheese(sharpBlue,Cheddar,pungentGorgonzola,Limburger,tangyBrick,Edam,Feta)General(Salad)" , "Cheese, General Food")
test['Food Paring'] = test['Food Paring'].replace("(German)Cheese(butteryBrie,Gouda,Havarti,Swiss)General(Salad)Meat(Pork,Fish,Shellfish)" , "German, Cheese, General Food, Meat")
test['Food Paring'] = test['Food Paring'].replace("(Barbecue,Indian,LatinAmerican,Thai,PanAsian)Cheese(pepperyMontereyPepperJack)Meat(Shellfish)" , "Barbecue, Indian, LatinAmerican, Thai, PanAsian, Cheese, Meat")
test['Food Paring'] = test['Food Paring'].replace("(German)General(Salad)Meat(Fish)" , "German, General Food, Meat")
test['Food Paring'] = test['Food Paring'].replace("Cheese(earthyCamembert,Fontina,nuttyAsiago,Colby,Parmesan)Meat(Pork,GrilledMeat)" , "Cheese, Meat")
test['Food Paring'] = test['Food Paring'].replace("(LatinAmerican)Meat(Beef,Poultry)" , "LatinAmerican, Meat")
test['Food Paring'] = test['Food Paring'].replace("(Curried,German)Cheese(nuttyAsiago,Colby,Parmesan)General(Digestive)Meat(Salmon)" , "German, Cheese, General Food, Meat")
test['Food Paring'] = test['Food Paring'].replace("Meat(Pork,Poultry)" , "Meat")
test['Food Paring'] = test['Food Paring'].replace("(Barbecue,LatinAmerican)Cheese(nuttyAsiago,Colby,Parmesan)General(Chocolate)Meat(Salmon)" , "Barbecue, LatinAmerican, Cheese, General Food, Meat")
test['Food Paring'] = test['Food Paring'].replace("(German)General(Chocolate,Dessert)Meat(GrilledMeat)" , "German, General Food, Meat")
test['Food Paring'] = test['Food Paring'].replace("Cheese(earthyCamembert,Fontina)General(Chocolate)Meat(GrilledMeat)" , "Cheese, General Food, Meat")
test['Food Paring'] = test['Food Paring'].replace("(Chocolate,Salad,Dessert,Aperitif)" , "Dessert")
test['Food Paring'] = test['Food Paring'].replace("(LatinAmerican,German)Meat(Beef,SmokedMeat,GrilledMeat)" , "LatinAmerican, German, Meat")
test['Food Paring'] = test['Food Paring'].replace("(Barbecue)Cheese(sharpBlue,Cheddar,pungentGorgonzola,Limburger)Meat(Beef,GrilledMeat)" , "Barbecue, Cheese, Meat")
test['Food Paring'] = test['Food Paring'].replace("Cheese(butteryBrie,Gouda,Havarti,Swiss,pungentGorgonzola,Limburger)General(Dessert,Digestive)Meat(Game)" , "Cheese, General Food, Meat")
test['Food Paring'] = test['Food Paring'].replace("Cheese(sharpBlue,Cheddar,tangyBrick,Edam,Feta)General(Aperitif,Digestive)" , "Cheese, General Food")
test['Food Paring'] = test['Food Paring'].replace("Cheese(butteryBrie,Gouda,Havarti,Swiss,sharpBlue,Cheddar)General(Digestive)Meat(Beef,SmokedMeat,Game)" , "Cheese, General Food, Meat")
test['Food Paring'] = test['Food Paring'].replace("(Barbecue,LatinAmerican)General(Chocolate)Meat(SmokedMeat,GrilledMeat)" , "Barbecue, LatinAmerican, General Food, Meat")
test['Food Paring'] = test['Food Paring'].replace("(Salad)Meat(Poultry,Game)" , "Salad, Meat")
test['Food Paring'] = test['Food Paring'].replace("(German)General(Salad)Meat(Poultry,Fish)" , "German, General Food, Meat")
test['Food Paring'] = test['Food Paring'].replace("Cheese(pungentGorgonzola,Limburger,tangyBrick,Edam,Feta)General(Digestive)Meat(Shellfish,Game,GrilledMeat)" , "Cheese, General Food, Meat")
test['Food Paring'] = test['Food Paring'].replace("(Barbecue)Cheese(pepperyMontereyPepperJack,sharpBlue,Cheddar)Meat(Beef,SmokedMeat,Game,GrilledMeat,Salmon)" , "Barbecue, Cheese, Meat")
test['Food Paring'] = test['Food Paring'].replace("(German)Cheese(sharpBlue,Cheddar)General(Salad)Meat(Pork)" , "German, Cheese, General Food, Meat")
test['Food Paring'] = test['Food Paring'].replace("(Thai,Chinese,Japanese,PanAsian)Meat(Pork,Poultry,Fish,Shellfish)" , "Thai, Chinese, Japanese, PanAsian, Meat")
test['Food Paring'] = test['Food Paring'].replace("(Barbecue,LatinAmerican)Cheese(pepperyMontereyPepperJack)Meat(Fish,SmokedMeat)" , "Barbecue, LatinAmerican, Cheese, Meat")
test['Food Paring'] = test['Food Paring'].replace("Cheese(sharpBlue,Cheddar,pungentGorgonzola,Limburger)General(Aperitif,Digestive)" , "Cheese, General Food")
test['Food Paring'] = test['Food Paring'].replace("(Curried)Cheese(nuttyAsiago,Colby,Parmesan,pepperyMontereyPepperJack)Meat(Poultry,Fish)" , "Curried, Cheese, Meat")
test['Food Paring'] = test['Food Paring'].replace("(German)Cheese(butteryBrie,Gouda,Havarti,Swiss,sharpBlue,Cheddar)General(Dessert,Digestive)" , "German, Cheese, General Food")
test['Food Paring'] = test['Food Paring'].replace("Cheese(earthyCamembert,Fontina,sharpBlue,Cheddar)Meat(GrilledMeat)" , "Cheese, Meat")
test['Food Paring'] = test['Food Paring'].replace("Cheese(butteryBrie,Gouda,Havarti,Swiss,sharpBlue,Cheddar,pungentGorgonzola,Limburger)" , "Cheese")
test['Food Paring'] = test['Food Paring'].replace("(Barbecue,German)Cheese(pepperyMontereyPepperJack,sharpBlue,Cheddar)Meat(Beef,SmokedMeat,Game,GrilledMeat,Salmon)" , "Barbecue, German, Cheese, Meat")
test['Food Paring'] = test['Food Paring'].replace("(Dessert,Aperitif)" , "Dessert")
test['Food Paring'] = test['Food Paring'].replace("(German)Cheese(butteryBrie,Gouda,Havarti,Swiss)General(Aperitif)Meat(Pork,Poultry,Fish,Shellfish)" , "German, Cheese, General Food, Meat")
test['Food Paring'] = test['Food Paring'].replace("(Japanese)" , "Japanese")
test['Food Paring'] = test['Food Paring'].replace("(Aperitif)Meat(Fish,Shellfish,Salmon)" , "Meat")
test['Food Paring'].nunique()
freq = nltk.FreqDist(test['Glassware Used'])
for key,value in freq.items():
print(str(key)+' : '+str(value))
test['Glassware Used'] = test['Glassware Used'].replace("PintGlass(orBecker,Nonic,Tumbler),Mug(orSeidel,Stein)" , "PintGlass, Mug")
test['Glassware Used'] = test['Glassware Used'].replace("Snifter,Tulip,OversizedWineGlass" , "Snifter, Tulip, OversizedWineGlass")
test['Glassware Used'] = test['Glassware Used'].replace("Flute,PilsenerGlass(orPokal),Mug(orSeidel,Stein)" , "Flute, PilsenerGlass, Mug")
test['Glassware Used'] = test['Glassware Used'].replace("PintGlass(orBecker,Nonic,Tumbler),Snifter,OversizedWineGlass" , "PintGlass, Snifter, OversizedWineGlass")
test['Glassware Used'] = test['Glassware Used'].replace("PilsenerGlass(orPokal" , "PilsenerGlass")
test['Glassware Used'] = test['Glassware Used'].replace("Flute,Tulip,OversizedWineGlass" , "Flute, Tulip, OversizedWineGlass")
test['Glassware Used'] = test['Glassware Used'].replace("PintGlass(orBecker,Nonic,Tumbler)" , "PintGlass")
test['Glassware Used'] = test['Glassware Used'].replace("PintGlass(orBecker,Nonic,Tumbler),Tulip,OversizedWineGlass" , "PintGlass, Tulip, OversizedWineGlass")
test['Glassware Used'] = test['Glassware Used'].replace("Flute,PilsenerGlass(orPokal),Mug(orSeidel,Stein),Stange(SlenderCylinder)" , "Flute, PilsenerGlass, Mug, Stange")
test['Glassware Used'] = test['Glassware Used'].replace("PintGlass(orBecker,Nonic,Tumbler),Snifter" , "PintGlass, Snifter")
test['Glassware Used'] = test['Glassware Used'].replace("PintGlass(orBecker,Nonic,Tumbler),PilsenerGlass(orPokal),Mug(orSeidel,Stein)" , "PintGlass, PilsenerGlass, Mug")
test['Glassware Used'] = test['Glassware Used'].replace("Mug(orSeidel,Stein)" , "Mug")
test['Glassware Used'] = test['Glassware Used'].replace("PilsenerGlass(orPokal),Mug(orSeidel,Stein)" , "PilsenerGlass, Mug")
test['Glassware Used'] = test['Glassware Used'].replace("Stange(SlenderCylinder)" , "Stange")
test['Glassware Used'] = test['Glassware Used'].replace("Goblet(orChalice)" , "Goblet")
test['Glassware Used'] = test['Glassware Used'].replace("PintGlass(orBecker,Nonic,Tumbler),Snifter,Mug(orSeidel,Stein)","PintGlass, Snifter, Mug")
test['Glassware Used'] = test['Glassware Used'].replace("PintGlass(orBecker,Nonic,Tumbler),Goblet(orChalice)" , "PintGlass, Goblet")
test['Glassware Used'] = test['Glassware Used'].replace("WeizenGlass" , "WeizenGlass")
test['Glassware Used'] = test['Glassware Used'].replace("Tulip,OversizedWineGlass" , "Tulip, OversizedWineGlass")
test['Glassware Used'] = test['Glassware Used'].replace("Flute,PilsenerGlass(orPokal)" , "Flute, PilsenerGlass")
test['Glassware Used'] = test['Glassware Used'].replace("PintGlass(orBecker,Nonic,Tumbler),Snifter,Tulip" , "PintGlass, Snifter, Tulip")
test['Glassware Used'] = test['Glassware Used'].replace("Snifter,Goblet(orChalice)" , "Snifter, Goblet")
test['Glassware Used'] = test['Glassware Used'].replace("PintGlass(orBecker,Nonic,Tumbler),Stange(SlenderCylinder)" , "PintGlass, Stange")
test['Glassware Used'] = test['Glassware Used'].replace("Tulip,Goblet(orChalice),OversizedWineGlass" , "Tulip, Goblet, OversizedWineGlass")
test['Glassware Used'] = test['Glassware Used'].replace("PintGlass(orBecker,Nonic,Tumbler),PilsenerGlass(orPokal)" , "PintGlass, PilsenerGlass")
test['Glassware Used'] = test['Glassware Used'].replace("Stange(SlenderCylinder),WeizenGlass" , "Stange, WeizenGlass")
test['Glassware Used'] = test['Glassware Used'].replace("PintGlass(orBecker,Nonic,Tumbler),Mug(orSeidel,Stein),WeizenGlass" , "PintGlass, Mug, WeizenGlass")
test['Glassware Used'] = test['Glassware Used'].replace("PintGlass(orBecker,Nonic,Tumbler),Mug(orSeidel,Stein),OversizedWineGlass" , "PintGlass, Mug, OversizedWineGlass")
test['Glassware Used'] = test['Glassware Used'].replace("Flute,WeizenGlass" , "Flute, WeizenGlass")
test['Glassware Used'] = test['Glassware Used'].replace("Flute,Snifter,Tulip" , "Flute, Snifter, Tulip")
test['Glassware Used'] = test['Glassware Used'].replace("Flute,Snifter,Tulip,Stange(SlenderCylinder)" , "Flute, Snifter, Tulip, Stange")
test['Glassware Used'] = test['Glassware Used'].replace("Snifter,Tulip,Goblet(orChalice),OversizedWineGlass" , "Snifter, Tulip, Goblet, OversizedWineGlass")
test['Glassware Used'] = test['Glassware Used'].replace("Snifter,OversizedWineGlass" , "Snifter, OversizedWineGlass")
test['Glassware Used'] = test['Glassware Used'].replace("Snifter,Tulip,Goblet(orChalice)" , "Snifter, Tulip, Goblet")
test['Glassware Used'] = test['Glassware Used'].replace("None,yet" , "None")
test['Glassware Used'] = test['Glassware Used'].replace("Flute,Stange(SlenderCylinder)" , "Flute,Stange")
test['Glassware Used'] = test['Glassware Used'].replace("Flute,Snifter,OversizedWineGlass" , "Flute, Snifter, OversizedWineGlass")
test['Glassware Used'] = test['Glassware Used'].replace("Mug(orSeidel,Stein),Stange(SlenderCylinder)" , "Mug, Stange")
test['Glassware Used'] = test['Glassware Used'].replace("Flute" , "Flute")
test['Glassware Used'].nunique()
test.dtypes
test['Food Paring label']= label_encoder.fit_transform(test['Food Paring'])
test['Glassware Used label']= label_encoder.fit_transform(test['Glassware Used'])
test['Style Name label']= label_encoder.fit_transform(test['Style Name'])
test['Ratings'] = pd.to_numeric(test['Ratings'],errors='coerce')
test['Beer Name'] = test['Beer Name'].astype(float)
test['Brewing Company'] = test['Brewing Company'].astype(float)
test.isnull().sum()
test_avg_ratings = test['Ratings'].astype(float).mean()
test['Ratings'].replace(np.nan, test_avg_ratings, inplace=True)
test.isnull().sum()
test.columns
test1 = test[['ABV', 'Ratings', 'Minimum Temperature', 'Maximum Temperature','Minimum Serving Temperature', 'Maximum Serving Temperature', 'Food Paring label', 'Glassware Used label', 'Style Name label']]
test1.isnull().sum()
```
## Data Pre-Processing
```
x = train1.iloc[:,:-1]
y = train1.iloc[:,-1]
print(x.columns)
from sklearn.preprocessing import StandardScaler
x = StandardScaler().fit(x).transform(x)
x[:3]
```
### Regression
```
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(x, y, random_state = 0)
from sklearn.linear_model import LinearRegression
reg = LinearRegression()
reg.fit(x_train, y_train)
y_pred = reg.predict(x_test)
print(y_pred)
# Calculating score from Root Mean Log Squared Error
def rmlse(y_test, y_pred):
error = np.square(np.log10(y_pred +1) - np.log10(y_test +1)).mean() ** 0.5
score = 1 - error
return score
print("\n----------------------------\nRMLSE Score = ", rmlse(y_test, y_pred))
```
### SVR
```
from sklearn.svm import SVR
svr = SVR(kernel='rbf')
# Training the regressor with training data
svr.fit(x_train, y_train)
y_pred2 = svr.predict(x_test)
print(y_pred2)
print("----------------------------\nRMLSE Score = ", rmlse(y_test, y_pred2))
pd.DataFrame({'Score' : y_pred}).to_excel("C:\\Users\\Moaz\\Desktop\\moaz\\Jupyter Python NB\\Machine Hack\\beer_score.xlsx")
```
| github_jupyter |
```
# SINGLE LAYER AND MULTI LAYER NETWORKS FOR MNIST
# BASED ON CODE FROM TENSORFLOW TUTORIAL
import tensorflow as tf
```
# OBTAIN
## (& SCRUB -- this data comes scrubbed)
```
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets("MNIST_data/", one_hot=True)
```
# MODEL
```
# MODEL
# CREATE PLACEHOLDER VARIABLES FOR OPERATION MANIPULATION
# THE 784 MATCHES THE VECTOR SIZE OF THE MNIST IMAGES - 28*28
x = tf.placeholder(tf.float32, [None, 784])
# MODEL
# CREATE WEIGHTS & BIASES VARIABLES
# IN TF, OUR MODEL PARAMETERS ARE OFTEN MANAGED AS VARIABLES
W = tf.Variable(tf.zeros([784, 10]))
b = tf.Variable(tf.zeros([10]))
# MODEL
# CREATE MODEL - USES SOFTMAX AS THE ACTIVATION FUNCTION
# REMEMBER GOAL FOR ACTIVATION FUNCTION IS TO "SHAPE" THE
# OUTPUT INTO A PROBABILITY DISTRO OVER THE 10 CLASSES
y = tf.nn.softmax(tf.matmul(x, W) + b)
# MODEL
# CREATE PREDICTED VARIABLE Y-HAT
# AND USE CROSS ENTROPY TO DETERMINE LOSS
# CROSS ENTROPY - HOW INEFFICIENT ARE OUR PREDICTIONS?
y_ = tf.placeholder(tf.float32, [None, 10])
cross_entropy = tf.reduce_mean(-tf.reduce_sum(y_ * tf.log(y), reduction_indices=[1]))
# MODEL
# TRAIN USING GRADIENT DESCENT
# LEARNING RATE AT MIDPOINT - .5 - MAKE SMALL ADJUSTMENTS TO MINIMIZE COST
train_step = tf.train.GradientDescentOptimizer(0.5).minimize(cross_entropy)
```
## MODEL -- RUN MODEL
```
# MODEL - RUN
sess = tf.InteractiveSession()
tf.global_variables_initializer().run()
for _ in range(10000):
batch_xs, batch_ys = mnist.train.next_batch(100)
sess.run(train_step, feed_dict={x: batch_xs, y_: batch_ys})
```
# EVALUATE
```
# EVALUATE MODEL
correct_prediction = tf.equal(tf.argmax(y,1), tf.argmax(y_,1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
print(sess.run(accuracy, feed_dict={x: mnist.test.images, y_: mnist.test.labels}))
```
# BLOCK TWO
Alternative Approach
```
# WEIGHT INITIALIZATION
def weight_variable(shape):
initial = tf.truncated_normal(shape, stddev=0.1)
return tf.Variable(initial)
def bias_variable(shape):
initial = tf.constant(0.1, shape=shape)
return tf.Variable(initial)
# CREATE CONVOLUTION AND POOLING LAYERS
def conv2d(x, W):
return tf.nn.conv2d(x, W, strides=[1, 1, 1, 1], padding='SAME')
def max_pool_2x2(x):
return tf.nn.max_pool(x, ksize=[1, 2, 2, 1],
strides=[1, 2, 2, 1], padding='SAME')
# FIRST CONVOLUTION LAYER
W_conv1 = weight_variable([5, 5, 1, 32])
b_conv1 = bias_variable([32])
x_image = tf.reshape(x, [-1, 28, 28, 1]) # BASE IMAGE SIZE OF 28 * 28
h_conv1 = tf.nn.relu(conv2d(x_image, W_conv1) + b_conv1)
h_pool1 = max_pool_2x2(h_conv1) # RESULTING IMAGE SIZE IS 14 * 14
# SECOND CONOLUTION LAYER
# MORE THAN ONE LAYER? DEEP LEARNING
W_conv2 = weight_variable([5, 5, 32, 64])
b_conv2 = bias_variable([64])
h_conv2 = tf.nn.relu(conv2d(h_pool1, W_conv2) + b_conv2)
h_pool2 = max_pool_2x2(h_conv2)
# FULLY CONNECTED LAYER - BEFORE OUTPUT
W_fc1 = weight_variable([7 * 7 * 64, 1024])
b_fc1 = bias_variable([1024])
h_pool2_flat = tf.reshape(h_pool2, [-1, 7*7*64])
h_fc1 = tf.nn.relu(tf.matmul(h_pool2_flat, W_fc1) + b_fc1) # ADD THE RECTIFIED LINEAR UNIT
# DROP LAYER - REDUCE OVERFITTING
# USE OF rate IS AN UPDATE BASED ON TF
keep_prob = tf.placeholder(tf.float32)
rate = 1 - keep_prob
h_fc1_drop = tf.nn.dropout(h_fc1, rate)
# LAST LAYER - OUTPUT
W_fc2 = weight_variable([1024, 10])
b_fc2 = bias_variable([10])
y_conv = tf.matmul(h_fc1_drop, W_fc2) + b_fc2
```
## MODEL -- RUN MODEL
```
# RUN THE MODEL
cross_entropy = tf.reduce_mean(
tf.nn.softmax_cross_entropy_with_logits(labels=y_, logits=y_conv))
train_step = tf.train.AdamOptimizer(1e-4).minimize(cross_entropy)
correct_prediction = tf.equal(tf.argmax(y_conv, 1), tf.argmax(y_, 1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
for i in range(10000):
batch = mnist.train.next_batch(50)
if i % 100 == 0:
train_accuracy = accuracy.eval(feed_dict={
x: batch[0], y_: batch[1], rate: 1.0})
print('step %d, training accuracy %g' % (i, train_accuracy))
train_step.run(feed_dict={x: batch[0], y_: batch[1], rate: 0.5})
print('test accuracy %g' % accuracy.eval(feed_dict={
x: mnist.test.images, y_: mnist.test.labels, rate: 1.0}))
```
| github_jupyter |
# Importance of XCov term in loss function
- How does the model behave differently without XCov?
- Does amount of input variation matter? (None,Med,hi)?
```
import os
import json
import numpy as np
import pandas as pd
import scipy
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
from src.data_loader import Shifted_Data_Loader
from src.plot import orig_vs_transformed as plot_ovt
from src.plot import enc_dec_samples
from src.models import GResNet,EResNet,ResBlock,EncResBlock
from src.config import get_config
from src.trainer import Trainer
from src.utils import prepare_dirs_and_logger
from keras.datasets import fashion_mnist,mnist
from keras.layers import Dense
# from tabulate import tabulate
config,_ = get_config()
# Boilerplate
setattr(config, 'proj_root', '/home/elijahc/projects/vae')
setattr(config, 'log_dir', '/home/elijahc/projects/vae/logs')
setattr(config, 'dev_mode',True)
# setattr(config,'model_dir','/home/elijahc/projects/vae/models/2019-01-17/')
# Architecture Params
setattr(config, 'enc_blocks', [32,64,128,256])
setattr(config, 'dec_blocks', [4,2,1])
setattr(config, 'z_dim', 10)
setattr(config, 'y_dim', 10)
# Training Params
setattr(config, 'batch_size', 512)
setattr(config, 'dataset', 'fashion_mnist')
setattr(config, 'epochs', 100)
setattr(config, 'monitor', 'val_G_loss')
setattr(config, 'min_delta', 0.25)
setattr(config, 'optimizer', 'adam')
# Loss Weights
setattr(config, 'xcov', 0)
setattr(config, 'recon', 25)
setattr(config, 'xent', 15)
if not config.dev_mode:
print('setting up...')
prepare_dirs_and_logger(config)
vars(config)
translation_amt = 0.8 # Med
DL = Shifted_Data_Loader(dataset=config.dataset,flatten=False,
rotation=None,
translation=translation_amt,
)
DL.input_shape
G_builder = GResNet(y_dim=config.y_dim,z_dim=config.z_dim,dec_blocks=config.dec_blocks,flatten_out=False)
E_builder = EResNet(blocks=config.enc_blocks,z_dim=config.z_dim,y_dim=config.y_dim)
trainer = Trainer(config,DL,E_builder,G_builder,)
# setattr(trainer.config,'model_dir','/home/elijahc/projects/vae/models/2019-01-22/')
pt,idx = plot_ovt(DL,cmap='gray')
from keras.utils import to_categorical
RF = to_categorical(np.ones(len(DL.sx_train)),num_classes=2)
trainer.compile_model()
trainer.E.summary()
trainer.go(x=DL.sx_train,
y={
'class':DL.y_train_oh,
# 'D_real':RF,
'G':DL.sx_train},
validation_split=0.05,
verbose=0)
hist_df = pd.DataFrame.from_records(trainer.model.history.history)
hist_df.head()
sns.set_context('paper')
metrics = ['loss','G_loss','class_acc']
fig,axs = plt.subplots(nrows=len(metrics),sharex=True,figsize=(5,10))
for metric_name,ax in zip(metrics,axs):
sns.scatterplot(data=hist_df[[metric_name,'val_'+metric_name]],ax=ax)
# if not config.dev_mode:
# trainer.save_model()
from keras.models import Model
from keras.layers import Input
generator = trainer.G
trainer.E.summary()
z_encoder = Model(trainer.input,trainer.z_lat)
classifier = Model(trainer.input,trainer.y_class)
# y_lat_encoder = Model(trainer.E.input,trainer.y_lat)
# decoder_inp = Input(shape=(config.y_dim+config.z_dim,))
# dec_layers = trainer.model.layers[-(1+(5*2)):]
# print(dec_layers)
# _gen_x = dec_layers[0](decoder_inp)
# l = dec_layers[1]
# isinstance(l,keras.layers.core.Reshape)
# F = None
# for l in dec_layers[1:]:
# print(type(l))
# if isinstance(l,keras.layers.merge.Add):
# _gen_x = l([F,_gen_x])
# else:
# _gen_x = l(_gen_x)
# if isinstance(l,keras.layers.convolutional.Conv2DTranspose):
# if l.kernel_size==(1,1):
# F = _gen_x
# # generator = Model(decoder_inp,_gen_x)
classifier.summary()
DL.y_test_oh.shape
classifier.compile(loss='categorical_crossentropy',optimizer='adam',metrics=['acc'])
classifier.evaluate(DL.sx_test,DL.y_test_oh,batch_size=config.batch_size)
z_enc = z_encoder.predict(DL.sx_test,batch_size=config.batch_size)
# y_lat = y_lat_encoder.predict(DL.sx_test,batch_size=config.batch_size)
y_lat = classifier.predict(DL.sx_test,batch_size=config.batch_size)
_lat_vec = np.concatenate([y_lat,z_enc],axis=1)
_lat_vec.shape
z_enc_mu = np.mean(z_enc,axis=0)
z_enc_cov = np.cov(z_enc,rowvar=False)
np.random.multivariate_normal(z_enc_mu,z_enc_cov,size=50).shape
regen = generator.predict(_lat_vec,batch_size=config.batch_size)
rand_im = np.random.randint(0,10000)
plt.imshow(regen[rand_im].reshape(56,56),cmap='gray')
_lat_vec[rand_im]
DL2 = Shifted_Data_Loader(dataset=config.dataset,flatten=False,
rotation=None,
translation=translation_amt,
)
enc_dec_samples(DL.x_test,DL.sx_test,z_enc,y_lat,generator)
z_enc2 = z_encoder.predict(DL2.sx_test,batch_size=config.batch_size)
y_lat2 = classifier.predict(DL2.sx_test,batch_size=config.batch_size)
_lat_vec2 = np.concatenate([y_lat2,z_enc2],axis=1)
regen2 = generator.predict(_lat_vec2,batch_size=config.batch_size)
from src.plot import remove_axes,remove_labels
from src.utils import gen_trajectory
examples = 5
rand_im = np.random.randint(0,10000,size=examples)
fix,axs = plt.subplots(examples,11,figsize=(8,4))
_lat_s = []
regen_s = []
out = gen_trajectory(z_enc[rand_im],z_enc2[rand_im],delta=.25)
out_y = gen_trajectory(y_lat[rand_im],y_lat2[rand_im],delta=.25)
for z,y in zip(out,out_y):
_lat = np.concatenate([y,z],axis=1)
_lat_s.append(_lat)
regen_s.append(generator.predict(_lat,batch_size=config.batch_size))
i=0
for axr,idx in zip(axs,rand_im):
axr[0].imshow(DL.x_test[idx].reshape(28,28),cmap='gray')
axr[1].imshow(DL.sx_test[idx].reshape(56,56),cmap='gray')
axr[2].imshow(regen[idx].reshape(56,56),cmap='gray')
for j,a in enumerate(axr[3:-3]):
a.imshow(regen_s[j][i,:].reshape(56,56),cmap='gray')
# a.imshow(s.reshape(56,56),cmap='gray')
axr[-3].imshow(regen2[idx].reshape(56,56),cmap='gray')
axr[-2].imshow(DL2.sx_test[idx].reshape(56,56),cmap='gray')
axr[-1].imshow(DL2.x_test[idx].reshape(28,28),cmap='gray')
for a in axr:
remove_axes(a)
remove_labels(a)
i+=1
# plt.imshow(regen[rand_im].reshape(56,56),cmap='gray')
fix.savefig('../../updates/2019-02-05/assets/img/translocate_{}.png'.format(translation_amt))
dxs = DL.dx[1]-14
dys = DL.dy[1]-14
from sklearn.preprocessing import MinMaxScaler
feat_range = (0,50)
z_enc_scaled = [MinMaxScaler(feat_range).fit_transform(z_enc[:,i].reshape(-1,1)).tolist() for i in np.arange(config.z_dim+config.y_dim)]
z_enc_scaled = np.squeeze(np.array(z_enc_scaled,dtype=int))
from collections import Counter
import dit
from dit import Distribution
def mutual_information(X,Y):
XY_c = Counter(zip(X,Y))
XY_pmf = {k:v/float(sum(XY_c.values())) for k,v in XY_c.items()}
XY_jdist = Distribution(XY_pmf)
return dit.shannon.mutual_information(XY_jdist,[0],[1])
z_dx_I = [mutual_information(z_enc_scaled[i],dxs.astype(int)+14) for i in np.arange(25)]
z_dy_I = [mutual_information(z_enc_scaled[i],dys.astype(int)+14) for i in np.arange(25)]
z_class_I = [mutual_information(z_enc_scaled[i],DL.y_test) for i in np.arange(25)]
z_I_df = pd.DataFrame.from_records({'class':z_class_I,'dy':z_dy_I,'dx':z_dx_I})
z_I_df['class'] = z_I_df['class'].values.round(decimals=1)
config.translation_amt = translation_amt
config.translation_amt
dir_path = '../data/xcov_importance/dist_{}/'.format(translation_amt)
z_I_df.to_pickle('../data/xcov_importance/dist_{}/z_mutual_info.pk'.format(translation_amt))
np.save('../data/xcov_importance/dist_{}/dxs'.format(translation_amt), DL.dx[1]-14)
np.save('../data/xcov_importance/dist_{}/dys'.format(translation_amt), DL.dy[1]-14)
np.save('../data/xcov_importance/dist_{}/z_enc'.format(translation_amt), z_enc)
hist_df.to_pickle(os.path.join(dir_path,'training_hist.df'))
with open(os.path.join(dir_path,'config.json'), 'w') as fp:
json.dump(vars(config), fp)
sns.set_context('talk')
fig,ax = plt.subplots(1,1,figsize=(6,5))
ax.set_ylim(0,0.9)
ax.set_xlim(0,0.9)
points = plt.scatter(x=z_I_df['dx'],y=z_I_df['dy'],c=z_I_df['class'],cmap='plasma')
plt.colorbar(points)
fig,ax = plt.subplots(1,1,figsize=(5,5))
ax.scatter(z_dx_I,z_dy_I)
# ax.set_ylim(0,0.8)
# ax.set_xlim(0,0.8)
plt.scatter(np.arange(25),sorted(z_class_I,reverse=True))
# plt.scatter(np.arange(25),z_dx_I)
# plt.scatter(np.arange(25),z_dy_I)
from src.metrics import var_expl,norm_var_expl
from collections import Counter
dtheta = DL.dtheta[1]
fve_dx = norm_var_expl(features=z_enc,cond=dxs,bins=21)
fve_dy = norm_var_expl(features=z_enc,cond=dys,bins=21)
fve_class = norm_var_expl(features=z_enc, cond=DL.y_test, bins=21)
# fve_dt = norm_var_expl(features=z_enc,cond=dtheta,bins=21)
# fve_dx_norm = (dxs.var()-fve_dx)/dxs.var()
# fve_dy_norm = (dys.var()-fve_dy)/dys.var()
# fve_dth_norm = (dtheta.var()-fve_dt)/dtheta.var()
fve_dx_norm = fve_dx
fve_dy_norm = fve_dy
import seaborn as sns
sns.set_context('talk')
fve_dx_norm.shape
# np.save(os.path.join(config.model_dir,'fve_dx_norm'),fve_dx_norm)
fig,ax = plt.subplots(1,1,figsize=(5,5))
plt.scatter(fve_dx_norm.mean(axis=0),fve_dy_norm.mean(axis=0))
plt.xlabel('fve_dx')
plt.ylabel('fve_dy')
plt.tight_layout()
# plt.savefig(os.path.join(config.model_dir,'fve_dx.png'))
# plt.ylim(-0.125,0.25)
xdim = np.argmax(fve_dx_norm.mean(axis=0))
fve_dy_norm.mean(axis=0)
# np.save(os.path.join(config.model_dir,'fve_dy_norm'),fve_dy_norm)
plt.scatter(np.arange(config.z_dim),fve_dy_norm.mean(axis=0))
plt.xlabel('Z_n')
plt.ylabel('fve_dy')
plt.tight_layout()
# plt.savefig(os.path.join(config.model_dir,'fve_dy.png'))
# plt.ylim(-0.125,0.25)
ydim = np.argmax(fve_dy_norm.mean(axis=0))
plt.scatter(np.arange(config.z_dim),fve_class.mean(axis=0))
plt.xlabel('Z_n')
plt.ylabel('fve_class')
# plt.ylim(0.0,0.5)
np.argmax(fve_class.mean(axis=0))
from src.plot import Z_color_scatter
Z_color_scatter(z_enc,[xdim,ydim],dxs)
Z_color_scatter(z_enc,[xdim,ydim],dys)
Z_color_scatter(z_enc,[7,18],dtheta)
from plt.
```
| github_jupyter |
# Freesurfer space to native space using `mri_vol2vol`
BMED360-2021: `freesurfer-to-native-space.ipynb`
```
%matplotlib inline
import os
import pathlib
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import nibabel as nib
from os.path import expanduser, join, basename, split
import sys
sys.path.append('.') # path to utils.py
import utils
import time
import shutil # copy files
cwd = os.getcwd()
```
### We will use the `fs711_subjects` Freesurfer tree previously run for the `bids_bg_bmed360` sample
```
fs711_home = '/usr/local/freesurfer'
working_dir = join(cwd, 'data')
bids_dir = '%s/bids_bg_bmed360' % (working_dir)
fs711_subj = '%s/fs711_subjects' % (working_dir)
dmri_res = '%s/dmri_results' % (working_dir)
if not os.path.exists(dmri_res):
os.makedirs(dmri_res)
else:
print('subdirectory dmri_results already exists')
```
The Freesurfer environment:
```
%%bash -s '/usr/local/freesurfer' './data/fs711_subjects'
echo $1
echo $2
FREESURFER_HOME=${1}; export FREESURFER_HOME
PATH=${FREESURFER_HOME}/bin:${PATH}; export PATH
SUBJECTS_DIR=${2}; export SUBJECTS_DIR
FSLDIR=/usr/local/fsl; export FSLDIR
PATH=${FSLDIR}/bin:${PATH}; export PATH
. ${FSLDIR}/etc/fslconf/fsl.sh
source ${FREESURFER_HOME}/SetUpFreeSurfer.sh
```
## How to Convert from FreeSurfer Space Back to Native Anatomical Space
See: https://surfer.nmr.mgh.harvard.edu/fswiki/FsAnat-to-NativeAnat
Question: I have successfully run a subject's data through [FreeSurfer](https://surfer.nmr.mgh.harvard.edu/fswiki/FreeSurfer).
FreeSurfer creates volumes in 1 mm$^3$, 256$^3$ space, but I want the FreeSurfer results in the space of my original anatomical. How do I do this?<br>
The exact command you use depends on what you want to convert, an image (like brain.mgz) or a segmentation (like aseg.mgz).
For an image:
```
# cd $SUBJECTS_DIR/<subjid>/mri
# mri_vol2vol --mov brain.mgz --targ rawavg.mgz --regheader --o brain-in-rawavg.mgz --no-save-reg
```
For a segmentation (aseg.mgz, aparc+aseg.mgz, wmparc.mgz, etc):
```
# cd $SUBJECTS_DIR/<subjid>/mri
# mri_label2vol --seg aseg.mgz --temp rawavg.mgz --o aseg-in-rawavg.mgz --regheader aseg.mgz
```
Map the surface to the native space:
```
# mri_surf2surf --sval-xyz pial --reg register.native.dat rawavg.mgz --tval lh.pial.native --tval-xyz rawavg.mgz --hemi lh --s subjectname
```
The output will be stored in $SUBJECTS_DIR/subjectname/surf/lh.pial.native and can be viewed with freeview rawavg.mgz -f ../surf/lh.pial.native<br>
To verify that this worked, run
```
# freeview -v rawavg.mgz -f lh.pial.native
MRI_VOL2VOL = '%s/bin/mri_vol2vol' % (fs711_home)
print(os.popen(MRI_VOL2VOL).read())
def my_mri_vol2vol(subj_dir, sub, ses, inp_image, targ_image, targ_name, out_dir, interptype):
"""
Ex.
cd $SUBJECTS_DIR/<subjid>/mri
mri_vol2vol --mov brain.mgz --targ rawavg.mgz --regheader --o brain-in-rawavg.mgz --no-save-reg
--interp interptype : interpolation cubic, trilin, nearest (def is trilin)
"""
fs_mri = join('%s' % (subj_dir), 'sub_%d_tp%d/mri' % (sub, ses))
cmd = [
MRI_VOL2VOL,
'--mov', '%s/%s.mgz' % (fs_mri, inp_image),
'--targ', '%s' % (targ_image),
'--regheader',
'--interp', '%s' % (interptype),
'--o', '%s/sub_%d_tp%d_%s_in_%s.nii.gz' % (out_dir, sub, ses, inp_image, targ_name),
'--no-save-reg']
# ' 2>', error_output_log,'>', output_log]
cmd_str = " ".join(cmd)
#print('cmd_str = \n%s\n' % cmd_str)
# EXECUTE
os.system(cmd_str)
```
### Testing the native space conversion on one subject (sub_102_tp1)
**using the `sub-102_ses-1_T1w.nii.gz` in the `bids_bg_bmed360` tree as target image**
```
subj_dir = fs711_subj
out_dir = dmri_res
sub = 102
ses = 1
targ_image = '%s/sub-%d/ses-%d/anat/sub-%d_ses-%d_T1w.nii.gz' % (bids_dir, sub, ses, sub, ses)
targ_name = 'native_space'
```
**Use the `my_mri_vol2vol()`function on different source images and masks using approriate interpolation ('trilinear' and 'nearest neighbour')**
```
%%time
shutil.copy2(targ_image,out_dir) # copy the original anatomy file in bids tree to out_dir
interptype = 'trilin'
inp_image = 'orig'
my_mri_vol2vol(subj_dir, sub, ses, inp_image, targ_image, targ_name, out_dir, interptype)
interptype = 'trilin'
inp_image = 'brain'
my_mri_vol2vol(subj_dir, sub, ses, inp_image, targ_image, targ_name, out_dir, interptype)
interptype = 'nearest'
inp_image = 'ribbon'
my_mri_vol2vol(subj_dir, sub, ses, inp_image, targ_image, targ_name, out_dir, interptype)
interptype = 'nearest'
inp_image = 'aseg'
my_mri_vol2vol(subj_dir, sub, ses, inp_image, targ_image, targ_name, out_dir, interptype)
interptype = 'nearest'
inp_image = 'wmparc'
my_mri_vol2vol(subj_dir, sub, ses, inp_image, targ_image, targ_name, out_dir, interptype)
interptype = 'nearest'
inp_image = 'aparc+aseg'
my_mri_vol2vol(subj_dir, sub, ses, inp_image, targ_image, targ_name, out_dir, interptype)
```
#### Run the native space conversion on all subjects and sessions using the `_T1_biascorr_brain.nii.gz` image obtained from `03-fsl-anat.ipynb` as target image.
```
%%time
subj_dir = fs711_subj
bids_dir = bids_dir
out_dir = dmri_res
targ_name = 'native_space'
for sub in [102, 103, 111, 123]:
for ses in [1, 2]:
print(f'Computing sub:{sub} ses:{ses}')
targ_image = join(bids_dir,'sub-%d/ses-%d/anat/sub-%d_ses-%d_T1w.nii.gz' % (sub, ses, sub, ses))
shutil.copy2(targ_image,out_dir) # copy the original anatomy file in bids tree to out_dir
inp_image = 'orig'
interptype = 'trilin'
my_mri_vol2vol(subj_dir, sub, ses, inp_image, targ_image, targ_name, out_dir, interptype)
inp_image = 'brain'
interptype = 'trilin'
my_mri_vol2vol(subj_dir, sub, ses, inp_image, targ_image, targ_name, out_dir, interptype)
inp_image = 'brainmask'
interptype = 'nearest'
my_mri_vol2vol(subj_dir, sub, ses, inp_image, targ_image, targ_name, out_dir, interptype)
inp_image = 'ribbon'
interptype = 'nearest'
my_mri_vol2vol(subj_dir, sub, ses, inp_image, targ_image, targ_name, out_dir, interptype)
inp_image = 'aseg'
interptype = 'nearest'
my_mri_vol2vol(subj_dir, sub, ses, inp_image, targ_image, targ_name, out_dir, interptype)
inp_image = 'wmparc'
interptype = 'nearest'
my_mri_vol2vol(subj_dir, sub, ses, inp_image, targ_image, targ_name, out_dir, interptype)
inp_image = 'aparc+aseg'
interptype = 'nearest'
my_mri_vol2vol(subj_dir, sub, ses, inp_image, targ_image, targ_name, out_dir, interptype)
```
| github_jupyter |
# Cdc2 Cyclin Model
Drew Willis
Source:</br>
Tyson, J. J. “Modeling the Cell Division Cycle: Cdc2 and Cyclin Interactions.”
Proceedings of the National Academy of Sciences, vol. 88, no. 16, 1991, pp. 7328–7332., doi:10.1073/pnas.88.16.7328. https://www.pnas.org/content/pnas/88/16/7328.full.pdf
This is a recreation of Tyson's Cdc2 and Cyclin Interactions model, and includes extra tools to explore other aspects of the model.
```
import tellurium as te
import numpy as np
import ipywidgets as widgets
import matplotlib.pyplot as plt
from ipywidgets import *
from IPython.display import display, update_display
```
## Background
(still need description here, recommend reading the abstract (bold text) here: https://www.pnas.org/content/pnas/88/16/7328.full.pdf)
The cell cycle is composed of cell growth and mitosis. These processes have been shown to operate differently in relation to each other. We look at the cell cycle as two processes:
* Cell Growth cycle
* Cell Division cycle
The division cycle is determined by an enzyme, the maturation promoting factor (MPF). It is formed by cdc2 and cyclin in the cell and is autocatalytic. The MPF enzyme is necessary for mitotic processes to occur. Once the cell reaches anaphase, MPF is degraded and the cycle repeats.
MPF activation can depend on cell growth, cyclin levels in the cell, or other enzymes, depending on the cell type and environment.

(Tyson)
In the model, this process is represented by 9 steps:
1. cyclin is created
2. cyclin can be unstable and be destroyed
3. cyclin is phosphorylated and forms heterodimer with
4. cdc2-P to form preMPF
5. preMPF is dephosphorylated to form active MPF
6. MPF activation can be opposed
7. Nuclear is division triggered, and active MPF is destroyed, releasing phosphorylated cyclin
8. Phosphorylated cyclin is destroyed
9. cdc2 is phosphorylated
10. cdc2 can be reversed
The original paper uses a series of differential equations, but I have chosen to represent the model in arrow equation format that is functionally the same.
#### Model equations
1. C2 -> CP ; k8*P*C2
2. CP -> C2 ; k9*CP
3. CP -> pM ; k3*CP*Y
4. pM -> M ; pM * (k4p + k4*(M/CT)^2)
5. M -> pM ; k5*P*M
6. M -> C2 ; k6*M
7. -> Y ; k1*aa
8. Y -> ; k2*Y + k3*CP*Y
9. -> YP ; k6*M
10. YP -> ; k7*YP
(will clean up these equations with better formatting for readability)
##### Variable descriptions
* aa : amino acids
* C2 : cdc2
* CP : cdc2-P
* pM : P-cyclin-cdc2-P / preMPF
* M : P-cyclin-cdc2 / active MPF
* Y : cyclin
* CP : cyclin-P
* CT : total cdc2
* k4 > k4p
#### Expected Result
The conclusion of the original paper found three states in which the system would operate:
1. Steady state high MPF activity
2. Autonomous oscillations
3. Excitable steady state
## Cycle Model
```
# ----< DEFINE MODEL >----
model = '''
// Equations
E1: C2 -> CP ; k8*P*C2
E2: CP -> C2 ; k9*CP
E3: CP -> pM ; k3*CP*Y
E4: pM -> M ; pM * (k4p + k4*(M/CT)^2)
E5: M -> pM ; k5*P*M
E6: M -> C2 ; k6*M
E7: -> Y ; k1*aa
E8: Y -> ; k2*Y + k3*CP*Y
E9: -> YP ; k6*M
E10: YP -> ; k7*YP
CT := C2+CP+pM+M
// Inputs
k1 := 0.015*CT/aa
k2 = 0
k3 := 200/CT
k4 = 100
k4p = 0.018
k5 = 0
k6 = 0.5
k7 = 0.6
k8 = 100
k9 = 10
P = 1
aa = 1
C2 = 0.1
CP = 1
pM = 0.1
M = 0.1
Y = 0.1
YP = 0.1
'''
# ----< WIDGETS >----
# model parameters
style = {'description_width': 'initial'}
k4_widget = widgets.FloatSlider(
description='k4 rate constant',
value=100.0,
min=10.0,
max=1000.0,
continuous_update=False,
style=style
)
k6_widget = widgets.FloatSlider(
description='k6 rate constant',
value=0.5,
min=0.1,
max=10.0,
continuous_update=False,
style=style
)
k8_widget = widgets.FloatSlider(
description='k8 rate constant',
value=0.5,
min=10.0,
max=200.0,
continuous_update=False,
style=style
)
k9_widget = widgets.FloatSlider(
description='k9 rate constant',
value=10.0,
min=10.0,
max=200.0,
continuous_update=False,
style=style
)
# simulation settings
sim_length_widget = widgets.IntSlider(
description='simulation length',
value=100,
min=2,
max=1000,
continuous_update=False,
style=style
)
sim_points_widget = widgets.IntSlider(
description='simulated points',
value=1000,
min=2,
max=20000,
continuous_update=False,
style=style
)
# display toggles
C2_widget = widgets.ToggleButton(
description='C2 toggle',
value=True,
)
CP_widget = widgets.ToggleButton(
description='CP toggle',
value=True,
)
pM_widget = widgets.ToggleButton(
description='pM toggle',
value=True,
)
Y_widget = widgets.ToggleButton(
description='Y toggle',
value=True,
)
M_widget = widgets.ToggleButton(
description='M toggle',
value=True,
)
YP_widget = widgets.ToggleButton(
description='YP toggle',
value=True,
)
yscale_widget = widgets.ToggleButton(
description='yscale: linear',
value=False
)
center_align = widgets.Layout(display='justify-content',
flex_flow='column',
align_items='stretch',
width='100%')
right_align = widgets.Layout(display='flex',
flex_flow='column',
align_items='flex-end',
width='100%')
left_vbox = widgets.VBox([k4_widget,k6_widget,k8_widget,k9_widget])
right_vbox = widgets.VBox([sim_length_widget,sim_points_widget])
simulation_hbox = widgets.HBox([left_vbox,right_vbox],layout=center_align)
yscale_hbox = widgets.HBox([yscale_widget], layout=right_align)
display_toggles_hbox = widgets.HBox([C2_widget,CP_widget,pM_widget,Y_widget,M_widget,YP_widget])
# ----< PLOT SETUP >----
fig, axs = plt.subplots(1,figsize=(15,7))
plt.close()
fig2, axs2 = plt.subplots(1, figsize=(7,7))
axs.set(title='cdc2 cyclin model',xlabel='time',ylabel='variable')
axs2.set(title='pM vs YP',xlabel='pM',ylabel='YP')
plt.close()
# ----< INTERACT AND RUN >----
def RunModel(*args):
# reload model in case user interacts with other cells and touches these widgets
m = te.loada(model)
m.k4 = k4_widget.value
m.k6 = k6_widget.value
m.k8 = k8_widget.value
m.k9 = k9_widget.value
s = m.simulate(0,sim_length_widget.value,sim_points_widget.value,['time','C2','CP','pM','Y','M','YP'])
axs.set(xlim=[0,sim_length_widget.value])
linewidth = 2
if C2_widget.value:
axs.plot(s['time'],s['C2'],linewidth=linewidth,label='C2')
if CP_widget.value:
axs.plot(s['time'],s['CP'],linewidth=linewidth,label='CP')
if pM_widget.value:
axs.plot(s['time'],s['pM'],linewidth=linewidth,label='pM')
if Y_widget.value:
axs.plot(s['time'],s['Y'],linewidth=linewidth,label='Y')
if M_widget.value:
axs.plot(s['time'],s['M'],linewidth=linewidth,label='M')
if YP_widget.value:
axs.plot(s['time'],s['YP'],linewidth=linewidth,label='YP')
if axs.lines:
axs.legend(bbox_to_anchor=(0.1, -0.175, 0.8, .102), loc=2, ncol=3, mode="expand",fontsize='large')
axs2.plot(s['pM'],s['YP'])
update_display(display_id="0", obj=fig)
update_display(display_id="1", obj=fig2)
axs.cla()
axs2.cla()
axs.set(title='cdc2 cyclin model',xlabel='time (minutes)',ylabel='variable')
axs2.set(title='pM vs YP',xlabel='pM',ylabel='YP')
for i in range(len(left_vbox.children)):
left_vbox.children[i].observe(RunModel,names='value')
for i in range(len(right_vbox.children)):
right_vbox.children[i].observe(RunModel,names='value')
for i in range(len(display_toggles_hbox.children)):
display_toggles_hbox.children[i].observe(RunModel,names='value')
RunModel()
```
## Model
Use the toggles to show and hide each component. The included rate constants have boundaries defined by the original paper. (Although k6 values should be rather low on the slider.)
```
display(fig, display_id="0")
display(display_toggles_hbox)
display(simulation_hbox)
display(yscale_hbox)
```
## Simulation Energy
Reading this graph can tell you about the energy of the system and whether or not it will continue to oscillate.
I have chosen to graph YP against pM since their oscillations are always out of sync.
* Large loops in this graph indicate continuous oscillations.
* Loops decaying inwards indicate the steady state with high MPF activity.
* Loops that break outwards indicate the excitable switch with low MPF activity.
(still need a better way to describe this section)
```
display(fig2, display_id="1")
display(display_toggles_hbox)
display(simulation_hbox)
```
### Parameter Scan
```
# ----< DEFINE MODEL >----
model = '''
// Equations
E1: C2 -> CP ; k8*P*C2
E2: CP -> C2 ; k9*CP
E3: CP -> pM ; k3*CP*Y
E4: pM -> M ; pM * (k4p + k4*(M/CT)^2)
E5: M -> pM ; k5*P*M
E6: M -> C2 ; k6*M
E7: -> Y ; k1*aa
E8: Y -> ; k2*Y
E9: Y -> ; k3*CP*Y
E10: -> YP ; k6*M
E11: YP -> ; k7*YP
CT := C2+CP+pM+M
// Inputs
k1 := 0.015*CT/aa
k2 = 0
k3 := 200/CT
k4 = 100
k4p = 0.018
k5 = 0
k6 = 0.5
k7 = 0.6
k8 = 100
k9 = 10
P = 1
aa = 1
C2 = 0.1
CP = 1
pM = 0.1
M = 0.1
Y = 0.1
YP = 0.1
'''
# ----< WIDGETS >----
# model parameters
style = {'description_width': 'initial'}
k8_widget = widgets.FloatSlider(
description='k8 rate constant',
value=0.5,
min=100.0,
max=200.0,
continuous_update=False,
style=style
)
k9_widget = widgets.FloatSlider(
description='k9 rate constant',
value=10.0,
min=10.0,
max=200.0,
continuous_update=False,
style=style
)
# simulation settings
sim_length_widget = widgets.IntSlider(
description='simulation length',
value=250,
min=2,
max=1000,
continuous_update=False,
style=style
)
sim_points_widget = widgets.IntSlider(
description='simulated points',
value=500,
min=2,
max=20000,
continuous_update=False,
style=style
)
cycleTimesW = widgets.Textarea(
value='',
placeholder='',
description='min/max cycle times:',
disabled=False
)
left_vbox = widgets.VBox([k8_widget,k9_widget])
right_vbox = widgets.VBox([sim_length_widget,sim_points_widget])
simulation_hbox = widgets.HBox([left_vbox,right_vbox,cycleTimesW])
# ----< PLOT SETUP >----
fig, axs = plt.subplots(1,figsize=(7,7))
axs.set(title='k4 vs k6 oscillation occurences',xlabel='k6',ylabel='k4')
plt.close()
# ----< INTERACT AND RUN >----
def GetState(s):
oscillations=0
flip=False
highYP=0
highpM=0
for i in range(int(len(s)/2),len(s['pM'])):
if flip:
highYP+=1
if s['pM'][i]>s['YP'][i]+0.075:
oscillations += 1
flip=False
else:
highpM+=1
if s['pM'][i]<s['YP'][i]+0.075:
oscillations += 1
flip=True
if oscillations>7:
return "green"
else:
if highYP>highpM:
return "red"
else:
return "orange"
def GetCycleTime(s):
flip=False
times = []
for i in range(int(len(s)/2),len(s['pM'])):
if flip:
if s['pM'][i]>s['YP'][i]+0.075:
times.append(s['time'][i])
flip=False
else:
if s['pM'][i]<s['YP'][i]+0.075:
flip=True
if times:
cycleTime = np.mean(np.diff(np.asarray(times)))
growthRate = 1/cycleTime
if ( growthRate > 1.0):
print("Error: Growth rate too large.")
return cycleTime, np.clip(growthRate, 0.0, 1.0)
else:
return 0.0, 0.0
def RunModel(*args):
# reload model in case user interacts with other cells and touches these widgets
m = te.loada(model)
x=[]
y=[]
color=[]
maxCycleTime=0
maxCT_k4=0
maxCT_k6=0
minCycleTime=99999
minCT_k4=0
minCT_k6=0
for lk4 in np.arange(1.0,3.0,0.1):
for lk6 in np.arange(-1.0,1.0,0.1):
m.resetAll()
m.k4 = 10**lk4
m.k6 = 10**lk6
m.k8 = k8_widget.value
m.k9 = k9_widget.value
s = m.simulate(0,sim_length_widget.value,sim_points_widget.value,['time','C2','CP','pM','Y','M','YP'])
state = GetState(s)
cycleTime, _ = GetCycleTime(s)
if (cycleTime > maxCycleTime):
maxCycleTime = cycleTime
maxCT_k4 = m.k4
maxCT_k6 = m.k6
elif ( (cycleTime < minCycleTime) and (cycleTime > 0.0) ):
minCycleTime = cycleTime
minCT_k4 = m.k4
minCT_k6 = m.k6
x.append(10**lk6)
y.append(10**lk4)
color.append(state)
cycleTimesW.value = "maxCT="+str(round(maxCycleTime,2))+" k4="+str(maxCT_k4)+" k6="+str(maxCT_k6)+"\n"+"minCT="+str(round(minCycleTime,2))+" k4="+str(minCT_k4)+" k6="+str(minCT_k6)
axs.scatter(x=x,y=y,color=color,label=color)
axs.set(yscale='log',xscale='log')
axs.legend(bbox_to_anchor=(0.1, -0.175, 0.8, .102), loc=2, ncol=3, mode="expand",fontsize='large')
#axs.set(xlim=[0,10],ylim=[10,20])
update_display(display_id="2", obj=fig)
#axs.lines=[]
axs.cla()
axs.set(title='k4 vs k6 oscillation occurences',xlabel='k6',ylabel='k4')
for i in range(len(left_vbox.children)):
left_vbox.children[i].observe(RunModel,names='value')
for i in range(len(right_vbox.children)):
right_vbox.children[i].observe(RunModel,names='value')
```
## Parameter Scan
From the model and the paper we see that k4 and k6 are the most important factors that determine the fate of the simulation. We can do a parameter scan of k4 and k6 to see a 2-dimensional plot of the results
Sliders for k8 and k9 are included to observe whether or not they have a significant impact on the state of the simulation.
(The legend is currently broken)
* red : steady state, high MPF activity
* green : continuous oscillations
* yellow : excitable switch, low MPF activity
This parameter scan also scans for the maximum and minimum growth rates within the oscillating simulations. MaxCT is the maximum division time in minutes, followed by the k4 and k6 that caused this result. MinCT is the minimum division time (not including results with 0).
```
display(fig, display_id="2")
display(simulation_hbox)
```
We clearly see how low k6 and high k4 tends to the steady state and the reverse tends to the excitable switch. This result looks very similar to the results of fig. 2 in the original paper.
I believe this graph also definitively shows k8 and k9 have no effect on the simulation.
```
from scipy.signal import argrelextrema
# ----< DEFINE MODEL >----
cycle_model = '''
// Equations
E1: C2 -> CP ; k8*P*C2
E2: CP -> C2 ; k9*CP
E3: CP -> pM ; k3*CP*Y
E4: pM -> M ; pM * (k4p + k4*(M/CT)^2)
E5: M -> pM ; k5*P*M
E6: M -> C2 ; k6*M
E7: -> Y ; k1*aa
E8: Y -> ; k2*Y + k3*CP*Y
E9: -> YP ; k6*M
E10: YP -> ; k7*YP
CT := C2+CP+pM+M
// Inputs
k1 := 0.015*CT/aa
k2 = 0
k3 := 200/CT
k4 = 100
k4p = 0.018
k5 = 0
k6 = 0.5
k7 = 0.6
k8 = 100
k9 = 10
P = 1
aa = 1
C2 = 0.1
CP = 1
pM = 0.1
M = 0.1
Y = 0.1
YP = 0.1
'''
growth_model = '''
// Equations
E1: -> P ; k*P
// Inputs
P = 100.0
k = 0.01
'''
# ----< WIDGETS >----
# model parameters
style = {'description_width': 'initial'}
k4_widget = widgets.FloatSlider(
description='k4 rate constant',
value=100.0,
min=10.0,
max=1000.0,
continuous_update=False,
style=style
)
k6_widget = widgets.FloatSlider(
description='k6 rate constant',
value=0.5,
min=0.1,
max=10.0,
continuous_update=False,
style=style
)
k8_widget = widgets.FloatSlider(
description='k8 rate constant',
value=0.5,
min=10.0,
max=200.0,
continuous_update=False,
style=style
)
k9_widget = widgets.FloatSlider(
description='k9 rate constant',
value=10.0,
min=10.0,
max=200.0,
continuous_update=False,
style=style
)
# simulation settings
sim_length_widget = widgets.IntSlider(
description='simulation length',
value=200,
min=2,
max=1000,
continuous_update=False,
style=style
)
sim_points_widget = widgets.IntSlider(
description='simulated points',
value=2000,
min=2,
max=20000,
continuous_update=False,
style=style
)
# display toggles
C2_widget = widgets.ToggleButton(
description='C2 toggle',
value=True,
)
CP_widget = widgets.ToggleButton(
description='CP toggle',
value=True,
)
pM_widget = widgets.ToggleButton(
description='pM toggle',
value=True,
)
Y_widget = widgets.ToggleButton(
description='Y toggle',
value=True,
)
M_widget = widgets.ToggleButton(
description='M toggle',
value=True,
)
YP_widget = widgets.ToggleButton(
description='YP toggle',
value=True,
)
yscale_widget = widgets.ToggleButton(
description='yscale: linear',
value=False
)
center_align = widgets.Layout(display='justify-content',
flex_flow='column',
align_items='stretch',
width='100%')
right_align = widgets.Layout(display='flex',
flex_flow='column',
align_items='flex-end',
width='100%')
left_vbox = widgets.VBox([k4_widget,k6_widget,k8_widget,k9_widget])
right_vbox = widgets.VBox([sim_length_widget,sim_points_widget])
simulation_hbox = widgets.HBox([left_vbox,right_vbox],layout=center_align)
yscale_hbox = widgets.HBox([yscale_widget], layout=right_align)
display_toggles_hbox = widgets.HBox([C2_widget,CP_widget,pM_widget,Y_widget,M_widget,YP_widget])
# ----< PLOT SETUP >----
fig, axs = plt.subplots(2,figsize=(15,9))
fig.subplots_adjust(left=None, bottom=None, right=None, top=None, wspace=10, hspace=0.4)
axs[0].set(title='cdc2 cyclin model',xlabel='time (minutes)',ylabel='variable')
axs[1].set(title='Growth model',xlabel='time (minutes)',ylabel='Population (cells)')
plt.close()
def GetGrowthRate(s):
flip=False
times = []
for i in range(int(len(s)/2),len(s['pM'])):
if flip:
if s['pM'][i]>s['YP'][i]+0.075:
times.append(s['time'][i])
flip=False
else:
if s['pM'][i]<s['YP'][i]+0.075:
flip=True
if times:
cycleTime = np.mean(np.diff(np.asarray(times)))
growthRate = 1/cycleTime
if ( growthRate > 1.0):
print("Error: Growth rate too large.")
return cycleTime, np.clip(growthRate, 0.0, 1.0)
else:
return 0.0, 0.0
# ----< INTERACT AND RUN >----
def RunModel(*args):
# reload model in case user interacts with other cells and touches these widgets
m = te.loada(cycle_model)
m.k4 = k4_widget.value
m.k6 = k6_widget.value
m.k8 = k8_widget.value
m.k9 = k9_widget.value
s = m.simulate(0,sim_length_widget.value,sim_points_widget.value,['time','C2','CP','pM','Y','M','YP'])
# times of each peak in minutes
#peaks = np.asarray(argrelextrema(s['M'], np.greater)) / (sim_points_widget.value/sim_length_widget.value)
#avgCycle = np.mean(np.diff(peaks)) # in minutes
cycleTime, growthRate = GetGrowthRate(s)
g_m = te.loada(growth_model)
# simulate growth model
g_m.k = growthRate
g_s = g_m.simulate(0,sim_length_widget.value,sim_points_widget.value,['time','P'])
axs[0].set(xlim=[0,sim_length_widget.value])
axs[1].set(xlim=[0,sim_length_widget.value])
axs[1].cla()
axs[1].set(title='Growth model ('+str(round(cycleTime,2))+' minute rate)',xlabel='time (minutes)',ylabel='Population (cells)')
linewidth = 2
if C2_widget.value:
axs[0].plot(s['time'],s['C2'],linewidth=linewidth,label='C2')
if CP_widget.value:
axs[0].plot(s['time'],s['CP'],linewidth=linewidth,label='CP')
if pM_widget.value:
axs[0].plot(s['time'],s['pM'],linewidth=linewidth,label='pM')
if Y_widget.value:
axs[0].plot(s['time'],s['Y'],linewidth=linewidth,label='Y')
if M_widget.value:
axs[0].plot(s['time'],s['M'],linewidth=linewidth,label='M')
if YP_widget.value:
axs[0].plot(s['time'],s['YP'],linewidth=linewidth,label='YP')
if axs[0].lines:
axs[0].legend(bbox_to_anchor=(0.1, -0.175, 0.8, .102), loc=2, ncol=3, mode="expand",fontsize='large')
if yscale_widget.value:
yscale_widget.description = 'yscale: log'
axs[1].set_yscale('log')
else:
yscale_widget.description = 'yscale: linear'
axs[1].set_yscale('linear')
axs[1].plot(g_s['time'],g_s['P'],linewidth=linewidth,label='P')
update_display(display_id="0", obj=fig)
axs[0].cla()
axs[0].set(title='cdc2 cyclin model',xlabel='time (minutes)',ylabel='variable')
for i in range(len(left_vbox.children)):
left_vbox.children[i].observe(RunModel,names='value')
for i in range(len(right_vbox.children)):
right_vbox.children[i].observe(RunModel,names='value')
for i in range(len(yscale_hbox.children)):
yscale_hbox.children[i].observe(RunModel,names='value')
for i in range(len(display_toggles_hbox.children)):
display_toggles_hbox.children[i].observe(RunModel,names='value')
RunModel()
```
# Changes in the model vs cell proliferation
How do the k4 and k6 parameters affect the exponential growth rate?
Using numbers from the parameter scan, we can get the maximum and minimum growth rates within the oscillating simulations. The faster the oscillations, the more cells we should see at the end of the second graph.
```
display(fig, display_id="0")
display(display_toggles_hbox)
display(simulation_hbox)
display(yscale_hbox)
```
# Source
Tyson, J. J. “Modeling the Cell Division Cycle: Cdc2 and Cyclin Interactions.” Proceedings of the National Academy of Sciences, vol. 88, no. 16, 1991, pp. 7328–7332., doi:10.1073/pnas.88.16.7328.
https://www.pnas.org/content/pnas/88/16/7328.full.pdf
| github_jupyter |
# Tokenizers
```
! pipenv install nltk
import nltk
from nltk import tokenize
s1 = """Why wase time say lot word when few word do trick?"""
s2 = """Hickory dickory dock, the mouse ran up the clock."""
from nltk.tokenize import word_tokenize
! df -h /home/christangrant/nltk_data
# nltk.download('punkt') # Download the model
word_tokenize(s1)
word_tokenize(s2)
paragraph = [s1,s2]
paragraph
' '.join(paragraph)
from nltk.tokenize import sent_tokenize
sent_tokenize(' '.join(paragraph))
[ word_tokenize(sent) for sent in sent_tokenize(' '.join(paragraph))]
```
# Vectorization
Given a corpus C, normalize C, then vectorize C.
Vectorizing C gives us a **vocabulary** and it gives us **weights**.
Get vocabulary => vocabulary is a list of terms that appear in the corpus.
Create a vector where each entry represents a voculary item.
If our vocabulary size is 10K, our vector size if 10K. (Good Normalization shrinks this vector size.)
If V is the vocabulary, V + 5 => vocabulary size.
- OOV (Out of Vocabulary) /Unknown terms
- Redacted terms
- grouped terms
-
Corpus =
>Old macdonald had a farm, on his farm he had a cow.
>Old macdonald had a farm, on his farm he had a pig.
>Old macdonald had a farm, on his farm he had a goat.
Normalize(Corpus)
>Old, macdonald, farm, pig, cow, goat
```pre
Old, macdonald, farm, pig, cow, goat
[1, 1, 1, 0, 1 ,0]
[1, 1, 1, 1 ,0, 0]
[1, 1, 1, 0, 0, 1]
```
**One-hot encoding** of text.
How can we get some positional information in this one-hot encoded format?
> Use weights to represent positions?
> Use nggrams to group terms together
### N-gram encoding
Vocabulary size grows with the size of the ngram
```pre
Old, macdonald, farm, pig, cow, goat
<s> Old, Old macdonald, macdonald farm, farm pig, pig cow, cow goat, goat </s>
[1,1,1,0,0,0,0]
[1,1,1,0,0,0,1]
[1,1,1,0,0,0,0]
```
Useful to have a range of n-grams when vectorizing.
## Bag of words model
Unordered bag representation of the vocabulary.
>Old macdonald had a farm, on his farm he had a cow.
>Old macdonald had a farm, on his farm he had a pig.
>Old macdonald had a farm, on his farm he had a goat.
```pre
bow =
Old, macdonald, farm, pig, cow, goat
[1, 1, 2, 0, 1 ,0]
[1, 1, 2, 1 ,0, 0]
[1, 1, 2, 0, 0, 1]
```
Unique words may be important!
## Term frequency
The raw frequency value of a term in a particular document.
$$
tf(word,document) = \sum_{v \in D} 1 (if v == w)
$$
## Document frequency
The number of documents that contain a word w.
## TF*IDF
idf = 1/df = log(N/(df+epslilon)) + epsilon
Term frequency * Inverse document freqency
## Smoothing.
Adding a small term to help handle out of vocabulary errors floating point issues.
```
! pipenv install sklearn
```
Sklearn has an api to documents.
>Transformers take a document and processes it using the function .fit_transform
>Vectorizers take a document and process it using .fit()
>fit create an internal model using the document
```
corpus = [ word_tokenize(sent) for sent in sent_tokenize(' '.join(paragraph))]
corpus
import sklearn
from sklearn.feature_extraction.text import CountVectorizer
cv = CountVectorizer(min_df=0., max_df=1.)
cv_matrix = cv.fit_transform([s1, s2])
print(cv_matrix)
cv_matrix.toarray()
vocab = cv.get_feature_names_out()
vocab
import pandas as pd
df = pd.DataFrame(cv_matrix.toarray(), columns=vocab)
df
bv = CountVectorizer(ngram_range=(2,2))
bv_matrix = bv.fit_transform([s1, s2])
vocab = bv.get_feature_names_out()
df1 = pd.DataFrame(bv_matrix.toarray(), columns=vocab)
df1
```
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
from IPython.display import Markdown, display
def printmd(string):
display(Markdown(string))
def colorize(string,color="red"):
return f"<span style=\"color:{color}\">{string}</span>"
```
# Problem description
### Subtask2: Detecting antecedent and consequence
Indicating causal insight is an inherent characteristic of counterfactual. To further detect the causal knowledge conveyed in counterfactual statements, subtask 2 aims to locate antecedent and consequent in counterfactuals.
According to (Nelson Goodman, 1947. The problem of counterfactual conditionals), a counterfactual statement can be converted to a contrapositive with a true antecedent and consequent. Consider example “Her post-traumatic stress could have been avoided if a combination of paroxetine and exposure therapy had been prescribed two months earlier”; it can be transposed into “because her post-traumatic stress was not avoided, (we know) a combination of paroxetine and exposure therapy was not prescribed”. Such knowledge can be not only used for analyzing the specific statement but also be accumulated across corpora to develop domain causal knowledge (e.g., a combination of paroxetine and exposure may help cure post-traumatic stress).
Please note that __in some cases there is only an antecedent part while without a consequent part in a counterfactual statement__. For example, "Frankly, I wish he had issued this order two years ago instead of this year", in this sentence we could only get the antecedent part. In our subtask2, when locating the antecedent and consequent part, please set '-1' as consequent starting index (character index) and ending index (character index) to refer that there is no consequent part in this sentence. For details, please refer to the 'Evaluation' on this website.
```
!ls
import pandas as pd
!pwd
df = pd.read_csv('../../.data/semeval2020_5/train_task2.csv')
```
We have this amount of data:
```
len(df)
import random
i = random.randint(0,len(df))
print(df.iloc[i])
print("-"*50)
print(df["sentence"].iloc[i])
print("-"*50)
print(df["antecedent"].iloc[i])
print("-"*50)
print(df["consequent"].iloc[i])
import random
i = random.randint(0,len(df))
s = df.loc[df["sentenceID"]==203483]
#print(s)
print("-"*50)
print(s["sentence"].iloc[0])
print("-"*50)
print(s["antecedent"].iloc[0])
print("-"*50)
print(s["consequent"].iloc[0])
df["antecedent"].iloc[0]
df["consequent"].iloc[0]
df["sentence"].iloc[0][df["consequent_startid"].iloc[0]:df["consequent_endid"].iloc[0]]
```
Check whether all indices fit the annotation
_Note: annotation indices are inclusive!_
```
for i in range(len(df)):
assert df["sentence"].iloc[i][df["antecedent_startid"].iloc[i]:df["antecedent_endid"].iloc[i]+1] \
== df["antecedent"].iloc[i]
if df["consequent_startid"].iloc[i]>0:
assert df["sentence"].iloc[i][df["consequent_startid"].iloc[i]:df["consequent_endid"].iloc[i]+1] \
== df["consequent"].iloc[i]
```
__Consequent part might not always exist!__
```
df.loc[df['consequent_startid'] == -1]
```
It does not exist in this number of cases
```
df_without_conseq = df.loc[df['consequent_startid'] == -1]
print(f"{len(df_without_conseq)} / {len(df)}")
```
Lets check what are the lengths of sentences, and how much sentences without consequent correlate with length.
```
all_lens = [len(s.split()) for s in df["sentence"].values.tolist()]
no_conseq_lens = [len(s.split()) for s in df_without_conseq["sentence"].values.tolist()]
all_lens
import matplotlib.pyplot as plt
values1 = all_lens
values2= no_conseq_lens
bins=100
_range=(0,max(all_lens))
fig = plt.figure(figsize=(8,6))
ax = fig.add_subplot(111)
ax.hist(values1, alpha=0.5, bins=bins, range=_range, color= 'b', label='All sentences')
ax.hist(values2, alpha=0.5, bins=bins, range=_range, color= 'r', label='Sentences without consequent')
ax.legend(loc='upper right', prop={'size':14})
plt.show()
```
Distribution is skewed a little bit toward smaller values, but there does not seem to be any big correlation here...
| github_jupyter |
```
import numpy as np
import scipy.io as sio
from sklearn import svm
from sklearn.model_selection import cross_val_score
from sklearn.metrics.pairwise import pairwise_distances
from sklearn import manifold
import matplotlib.pyplot as plt
from sklearn.decomposition import PCA
from sklearn.svm import SVC
from graph_kernels_lib import WeisfeilerLehmanKernel, fit_n_components
ppi = sio.loadmat("PPI.mat")
ppi_graphs = ppi['G'][0]
ppi_labels = ppi['labels'].ravel()
n = ppi_labels.shape[0]
wl_kernel = WeisfeilerLehmanKernel()
K = wl_kernel.eval_similarities(ppi_graphs[:]['am'], 2)
D = pairwise_distances(K, metric='euclidean')
plt.imshow(D, zorder=2, cmap='Blues', interpolation='nearest')
plt.colorbar();
plt.style.use("ggplot")
plt.show()
```
# SVM Linear Classifier
```
from sklearn.model_selection import StratifiedKFold
strat_k_fold = StratifiedKFold(n_splits = 10, shuffle = True) #10
clf = svm.SVC(kernel="linear", C = 1.0)
scores_ln = cross_val_score(clf, D, ppi_labels, cv = strat_k_fold)
print(str(np.min(scores_ln)) +" - "+str(np.mean(scores_ln))+ " - " + str(np.max(scores_ln)) + " - "+ str(np.std(scores_ln)))
PCA_D = PCA(n_components = 2).fit_transform(D)
plt.plot(np.cumsum(PCA().fit(D).explained_variance_ratio_))
plt.show()
np.cumsum(PCA().fit(D).explained_variance_ratio_)[:3]
acidovorax = PCA_D[ppi_labels == 1]
acidobacteria = PCA_D[ppi_labels == 2]
clf = clf.fit(PCA_D, ppi_labels)
w = clf.coef_[0]
a = -w[0] / w[1]
xx = np.linspace(np.min(PCA_D), np.max(PCA_D))
yy = a * xx - (clf.intercept_[0]) / w[1]
plt.figure(figsize=(10,5))
ax_av = plt.scatter(acidovorax[:, 0], acidovorax[:, 1], color = "xkcd:red", marker = "^",label = "Acidovorax", s = 455, alpha = 0.65)
ax_ab = plt.scatter(acidobacteria[:, 0], acidobacteria[:, 1], color = "green", label = "Acidobacteria", s = 250, alpha = 0.75)
svm_line = plt.plot(xx, yy, color = "xkcd:sky blue", linestyle = "--", linewidth = 3.0)
plt.axis('tight');
#plt.grid(True)
plt.legend(prop={'size': 15})
ax_av.set_facecolor('xkcd:salmon')
ax_ab.set_facecolor('xkcd:pale green')
plt.show()
from mpl_toolkits.mplot3d import Axes3D
fig = plt.figure(figsize=(10,8))
ax = fig.add_subplot(111, projection='3d')
PCA_D = PCA(n_components = 3).fit_transform(D)
acidovorax = PCA_D[ppi_labels == 1]
acidobacteria = PCA_D[ppi_labels == 2]
clf = clf.fit(PCA_D, ppi_labels)
w = clf.coef_[0]
a = -w[0] / w[1]
xx = np.linspace(np.min(PCA_D), np.max(PCA_D))
yy = a * xx - (clf.intercept_[0]) / w[1]
#plt.figure(figsize=(10,5))
ax_av = ax.scatter(acidovorax[:, 0], acidovorax[:, 1], acidovorax[:, 2],c = "xkcd:red", marker = "^",label = "Acidovorax", s = 455, alpha = 0.65)
ax_ab = ax.scatter(acidobacteria[:, 0], acidobacteria[:, 1], acidobacteria[:, 2], c = "green", label = "Acidobacteria", s = 250, alpha = 0.75)
#svm_line = plt.plot(xx, yy, color = "xkcd:sky blue", linestyle = "--", linewidth = 3.0)
plt.axis('tight');
#plt.grid(True)
plt.legend(prop={'size': 15})
ax_av.set_facecolor('xkcd:salmon')
ax_ab.set_facecolor('xkcd:pale green')
ax.view_init(azim = 30, elev = 25)
plt.show()
```
# Manifold Learning Isomap
```
n_neighbors = 14#15
n_components = 2
iso_prj_D = manifold.Isomap(n_neighbors, n_components).fit_transform(D)
scores_ln = cross_val_score(clf, iso_prj_D, ppi_labels, cv = strat_k_fold, n_jobs= 8)
np.mean(scores_ln)
```
It seems that manifold learning with Isomap does not improve the performance of our linear svm classifier
### Plots for Isomap
```
acidovorax = iso_prj_D[ppi_labels == 1]
acidobacteria = iso_prj_D[ppi_labels == 2]
clf = clf.fit(iso_prj_D, ppi_labels)
w = clf.coef_[0]
a = -w[0] / w[1]
xx = np.linspace(np.min(iso_prj_D), np.max(iso_prj_D))
yy = a * xx - (clf.intercept_[0]) / w[1]
plt.figure(figsize=(10,5))
ax_av = plt.scatter(acidovorax[:, 0], acidovorax[:, 1], color = "xkcd:red", marker = "^",label = "Acidovorax", s = 455, alpha = 0.65)
ax_ab = plt.scatter(acidobacteria[:, 0], acidobacteria[:, 1], color = "green", label = "Acidobacteria", s = 250, alpha = 0.75)
svm_line = plt.plot(xx, yy, color = "xkcd:sky blue", linestyle = "--", linewidth = 3.0)
plt.axis('tight');
#plt.grid(True)
plt.legend(prop={'size': 15})
ax_av.set_facecolor('xkcd:salmon')
ax_ab.set_facecolor('xkcd:pale green')
plt.show()
```
#### Fit with best n of components
```
opt_n_components = fit_n_components(D, ppi_labels, manifold.Isomap, n_iteration= 10)
opt_iso_prj_D = manifold.Isomap(n_neighbors, opt_n_components).fit_transform(D)
scores_ln = cross_val_score(clf, opt_iso_prj_D, ppi_labels, cv = strat_k_fold, n_jobs= 8)
np.mean(scores_ln)
```
# Manifold Learning LocalLinearEmbedding
```
n_neighbors = 13#15
n_components = 15
lle_prj_D = manifold.LocallyLinearEmbedding(n_neighbors, n_components).fit_transform(D)
scores_ln = cross_val_score(clf, lle_prj_D, ppi_labels, cv = strat_k_fold, n_jobs= 8)
np.mean(scores_ln)
```
It seems that also manifold learning with LocalLinearEmbedding does not improve the performance of our linear svm classifier
### Plots for LLE
```
acidovorax = lle_prj_D[ppi_labels == 1]
acidobacteria = lle_prj_D[ppi_labels == 2]
clf = clf.fit(lle_prj_D, ppi_labels)
w = clf.coef_[0]
a = -w[0] / w[1]
xx = np.linspace(-0.2,0.25)
yy = a * xx - (clf.intercept_[0]) / w[1]
plt.figure(figsize=(10,5))
ax_av = plt.scatter(acidovorax[:, 0], acidovorax[:, 1], color = "xkcd:red", marker = "^",label = "Acidovorax", s = 455, alpha = 0.65)
ax_ab = plt.scatter(acidobacteria[:, 0], acidobacteria[:, 1], color = "green", label = "Acidobacteria", s = 250, alpha = 0.75)
svm_line = plt.plot(xx, yy, color = "xkcd:sky blue", linestyle = "--", linewidth = 3.0)
plt.axis('tight');
#plt.grid(True)
plt.legend(prop={'size': 15})
ax_av.set_facecolor('xkcd:salmon')
ax_ab.set_facecolor('xkcd:pale green')
plt.show()
```
#### Fit with best n of components
```
opt_n_components = fit_n_components(D, ppi_labels, manifold.LocallyLinearEmbedding, n_neighbors=13, n_iteration= 10)
opt_n_components
opt_lle_prj_D = manifold.LocallyLinearEmbedding(13, opt_n_components).fit_transform(D)
scores_ln = cross_val_score(clf, opt_lle_prj_D, ppi_labels, cv = strat_k_fold, n_jobs= 8)
np.mean(scores_ln)
```
# Graphs plots
```
import networkx as nx
G = nx.from_numpy_matrix(ppi_graphs[10]['am'])
#pos=nx.spring_layout(G) # positions for all nodes
pos = nx.spring_layout(G, k = 0.9, iterations = 1000)
nx.draw_networkx_nodes(G, pos, with_labels= False, node_color = "green", node_size = 300, alpha = 0.8)
nx.draw_networkx_edges(G, pos, width = 2, alpha=0.5,edge_color='r')
plt.axis('off')
#plt.savefig("acidovorax_graph_10.png") # save as png
plt.show() # display
G = nx.from_numpy_matrix(ppi_graphs[59]['am'])
#pos=nx.spring_layout(G) # positions for all nodes
pos = nx.spring_layout(G, k = 0.9, iterations = 1000)
nx.draw_networkx_nodes(G, pos, with_labels= False, node_color = "green", node_size = 300, alpha = 0.8)
nx.draw_networkx_edges(G, pos, width = 2, alpha=0.5,edge_color='r')
plt.axis('off')
#plt.savefig("Acidobacteria_graph_59.png") # save as png
plt.show() # display
G = nx.from_numpy_matrix(ppi_graphs[6]['am'])
#pos=nx.spring_layout(G) # positions for all nodes
pos = nx.spring_layout(G, k = 0.9, iterations = 1000)
nx.draw_networkx_nodes(G, pos, with_labels= False, node_color = "green", node_size = 300, alpha = 0.8)
nx.draw_networkx_edges(G, pos, width = 2, alpha=0.5,edge_color='r')
plt.axis('off')
#plt.savefig("acidovorax_graph_2.png") # save as png
plt.show() # display
G = nx.from_numpy_matrix(ppi_graphs[48]['am'])
#pos=nx.spring_layout(G) # positions for all nodes
pos = nx.spring_layout(G, k = 0.9, iterations = 1000)
nx.draw_networkx_nodes(G, pos, with_labels= False, node_color = "green", node_size = 300, alpha = 0.8)
nx.draw_networkx_edges(G, pos, width = 2, alpha=0.5,edge_color='r')
plt.axis('off')
#plt.savefig("Acidobacteria_graph_48.png") # save as png
plt.show() # display
node_labels = wl_kernel.extract_graphs_labels(ppi_graphs[:]['am'])
size = int(np.max(np.concatenate(node_labels)))
degree_component = np.zeros((n, size))
for i in range(len(node_labels)):
for j in node_labels[i]:
degree_component[i,int(j)-1] += 1
degree_component[0]
```
| github_jupyter |
# NumPy Practice Exercise
In this notebook, you have to use the knowledge that you have gathered from the `numpy-intro` notebook and solve all four Questions to pass the second module.
```
import numpy as np
from grader import grader_1
np.random.seed(10) #do not edit
```
#### Question 1
Create two numpy arrays such that `a` should be all integers between 25-35 (inclusive) and `b` should be ten evenly spaced numbers between 1-6 (inclusive). Perform the following operations on these NumPy arrays:
1. Cube (i.e. raise to the power of 3) all the elements in both arrays (element-wise)
2. Add both the cubed arrays (e.g., [1,2] + [3,4] = [4,6])
3. Sum the elements with even indices of the added array.
4. Take the square root of the added array (element-wise square root)
5. Append `b` to `a`, reshape the appended array so that it is a 4x5, 2d array and store the results in a variable `m`
6. Shuffle `m` and extract the third and the fourth column of the m matrix. Store the resulting 4x2 matrix in a new variable `m2`.
```
# Your code goes here
# Answer
```
#### Question 2
Create two numpy arrays such that `A` should be a 4x3, 2d array with integers randomly chosen from 1 to 11 and `b` should be a 4x1, 2d array with integers randomly chosen from 1 to 11. Using numpy functions and routines solve for x in `Ax = b`. Note, you should use numpy's pseudoinverse function while aligning A and b's dimensions.
```
Expected answer: x = [[ 0.64303194]
[ 0.57685743]
[-0.28759588]]
```
```
# Your code goes here
# Answer
```
#### Question 3
Create an 1d numpy array `original_data` with 1000 elements and divide that array into two arrays, `train` with 2/3rd of the elements of `original_data` and `test` with remaining 1/3rd elements of `original_data`.
```
# Your code goes here
# Answer
```
#### Question 4 (Graded)
Let `x` be the number of miles a person drives per day and `y` be the dollars spent on buying car fuel (per day). Created two numpy arrays each of size 100 such that represent `x` (number of miles) ranges from 1 to 10 (hint : use np.linspace()) with a uniform noise of (0,1/2) and `y` (money spent in dollars) will be from 1 to 20 (hint : use np.linspace()) with a uniform noise (0,1). Once these arrays are created, find the:
1. Expected value of x and the expected value of y
2. Variance and co-variance of distributions of x and y
3. Assuming that number of dollars spent in car fuel is only dependant on the miles driven, by a linear relationship. Write code that uses a linear predictor to calculate a predicted value of y for each x i.e `y_estimated = mx + b`. Refer to the image below for the formulae.
4. y_pred for each value in x, put the error into an array called y_error
5. Root mean square error (RMSE)

```
# Your code goes here
# Hint : use np.linspace to generate 100 numbers within the range as specified in the question.
x= #generate x
np.random.seed(0) # do not edit
x= x + # Add uniform noise using np.random.uniform
y= #generate y
np.random.seed(0) # do not edit
y=y + # Add uniform noise using np.random.uniform
# Answer
e_x=
e_y=
v_x=
v_y=
# To calculate covariance of x and y, use the following formula
# cov(x,y)= Expectation of (x*y) - (expectation of (x) * expectation of (y))
cov_xy=
b=
m=
y_estimated=
```
# To calculate RMSE

```
# write code
rmse =
print(rmse)
```
### Submit assignment
Run the cell below to grade and submit your assignment
```
grader_1(rmse)
```
| github_jupyter |
# Phase 2 Review
```
import pandas as pd
import numpy as np
from sklearn.preprocessing import StandardScaler
from statsmodels.formula.api import ols
pd.set_option('display.max_columns', 100)
```
### Check Your Data … Quickly
The first thing you want to do when you get a new dataset, is to quickly to verify the contents with the .head() method.
```
df = pd.read_csv('movie_metadata.csv')
print(df.shape)
df.head()
```
## Question 1
A Hollywood executive wants to know how much an R-rated movie released after 2000 will earn. The data above is a sample of some of the movies with that rating during that timeframe, as well as other movies. How would you go about answering her question? Talk through it theoretically and then do it in code.
What is the 95% confidence interval for a post-2000 R-rated movie's box office gross?
```
df.isna().sum()
# talk through your answer here
'''
Drop null values.
Filter dataframe for movies after 2000, content rating is 'R'.
Calculate mean, standard deviation, sample size, and plug those into the confidence interval formula to
find the lower and upper bounds of what the executive can expect such a movie to make.
'''
# do it in code here
df.dropna(subset=['gross'], inplace=True)
df_2000R = df[(df['title_year'] > 2000) & (df['content_rating'] == 'R')]
mean = df_2000R.gross.mean()
sd = df_2000R.gross.std()
n = df_2000R.gross.count()
mean, sd, n
se = sd/n**.5
# 95% confidence interval
mean - 1.96 * (sd / n**.5), mean + 1.96 * (sd / n**.5)
```
## Question 2a
Your ability to answer the first question has the executive excited and now she has many other questions about the types of movies being made and the differences in those movies budgets and gross amounts.
Read through the questions below and **determine what type of statistical test you should use** for each question and **write down the null and alternative hypothesis for those tests**.
- Is there a relationship between the number of Facebook likes for a cast and the box office gross of the movie?
- Do foreign films perform differently at the box office than non-foreign films?
- Of all movies created are 40% rated R?
- Is there a relationship between the language of a film and the content rating (G, PG, PG-13, R) of that film?
- Is there a relationship between the content rating of a film and its budget?
# your answer here
### Facebook Likes (cast) and Box Office Gross
'''
Correlation/simple linear regression
'''
Ho: Beta =0
Ha: Beta != 0
### Domestic vs. Foreign and Box Office Gross
'''
Two-sample T-Test
'''
Ho: mu_domestic = mu_foreign
Ha: mu_domestic != mu_foreign
### Rated R
'''
One-sample Z-Test of proportion
'''
Ho: P = 0.4
Ha: P != 0.4
### Language and Content rating
'''
Chi-square
'''
Ho: distributions are equal
Ha: distributions are not equal
### Content rating and budget
'''
ANOVA
'''
Ho: mu_r = mu_PG13 = mu_PG = mu_G
Ha: They are not all equal
## Question 2b
Calculate the answer for the second question:
- Do foreign films perform differently at the box office than non-foreign films?
```
df.head()
import scipy
import numpy as np
USA_array = np.array(df[df.country == "USA"].gross)
Foreign_array = np.array(df[df.country != "USA"].gross)
scipy.stats.ttest_ind(USA_array,Foreign_array, nan_policy = 'omit')
# your answer here
df_foreign = df[df.country != 'USA'].dropna(subset=['country'])
df_domestic = df[df.country == 'USA']
df_foreign.shape, df_domestic.shape
from scipy.stats import ttest_ind
ttest_ind(df_foreign.gross, df_domestic.gross)
'''
Yes! There is a statistically significant difference between the box office gross of foreign and domestic films.
'''
```
## Question 3
Now that you have answered all of those questions, the executive wants you to create a model that predicts the money a movie will make if it is released next year in the US. She wants to use this to evaluate different scripts and then decide which one has the largest revenue potential.
Below is a list of potential features you could use in the model. Create a new frame containing only those variables.
Would you use all of these features in the model?
Identify which features you might drop and why.
*Remember you want to be able to use this model to predict the box office gross of a film **before** anyone has seen it.*
- **budget**: The amount of money spent to make the movie
- **title_year**: The year the movie first came out in the box office
- **years_old**: How long has it been since the movie was released
- **genres**: Each movie is assigned one genre category like action, horror, comedy
- **imdb_score**: This rating is taken from Rotten tomatoes, and is the average rating given to the movie by the audience
- **actor_1_facebook_likes**: The number of likes that the most popular actor in the movie has
- **cast_total_facebook_likes**: The sum of likes for the three most popular actors in the movie
- **language**: the original spoken language of the film
```
df.loc[0, 'genres'].split('|')
df['genres'] = df.genres.apply(lambda x: x.split('|')[0])
df.genre.head()
df.columns
# your answer here
model_data = df[[
'gross', 'budget', 'actor_1_facebook_likes', 'cast_total_facebook_likes',
'title_year', 'content_rating', 'genres'
]]
model_data.corr()
# '''
# drop either `cast_total_facebook_likes` or `actor_1_facebook_likes` due to multicollinearity
# '''
'''
`num_critic_for_reviews` and `imdb_score` can't be known before the movie is released.
we'll drop them from the model.
drop either `cast_total_facebook_likes` or `actor_1_facebook_likes` due to multicollinearity.
'''
```
## Question 4a
Create the following variables:
- `years_old`: The number of years since the film was released.
- Dummy categories for each of the following ratings:
- `G`
- `PG`
- `R`
Once you have those variables, create a summary output for the following OLS model:
`gross~cast_total_facebook_likes+budget+years_old+G+PG+R`
```
import pandas as pd
model_data['years_old'] = 2020 - model_data.title_year
model_data = pd.get_dummies(model_data, columns=['content_rating']).drop(columns='content_rating_PG-13')
model_data.columns
from statsmodels.formula.api import ols writing out the formula
from statsmodels.api import OLS using x,y
# your answer here
lr_model = ols(formula='gross~cast_total_facebook_likes+budget+years_old+G+PG+R', data=model_data).fit()
lr_model.summary()
```
## Question 4b
Below is the summary output you should have gotten above. Identify any key takeaways from it.
- How ‘good’ is this model?
- Which features help to explain the variance in the target variable?
- Which do not?
<img src="ols_summary.png" style="withd:300px;">
```
'''
The model is not very good in that it only explains about 7.9% (13.9% in mine) of the variation
in the data around the mean. (based on R-squared value)
In the photo, Total Facebook likes, budget, age, PG rating, and R rating help to explain the variance,
whereas G rating does not. (based on p-values)
In mine, everything other than years old helps to explain the variance.
'''
```
## Question 5
**Bayes Theorem**
An advertising executive is studying television viewing habits of married men and women during prime time hours. Based on the past viewing records he has determined that during prime time wives are watching television 60% of the time. It has also been determined that when the wife is watching television, 40% of the time the husband is also watching. When the wife is not watching the television, 30% of the time the husband is watching the television. Find the probability that if the husband is watching the television, the wife is also watching the television.
```
# your answer here
'''
P(A) = Probability wife is watching tv
P(B) = Probability husband is watching tv
P(A|B) = Probability wife is watching tv given husband is
P(B|A) = Probability husband is watching tv given wife is
'''
p_A = 0.6
p_notA = 1 - p_A
p_B_given_A = 0.4
p_B_given_notA = 0.3
p_A_given_B = (p_B_given_A * p_A) / (p_B_given_A * p_A + p_B_given_notA * p_notA)
p_A_given_B
```
## Question 6
Explain what a Type I error is and how it relates to the significance level when doing a statistical test.
```
# your answer here
'''
A Type I error occurs when you reject the null hypothesis even though the null hypothesis is True.
The likelihood of a Type I error is directly related to changes in the significance level. If you
increase the significance level, the likelihood of a Type I error also increases and vice versa.
If our significane lecel is 95%, that means we have a 5% chance of making a type one error.
'''
```
## Question 7
How is the confidence interval for a sample related to a one sample t-test?
The range of a confidence interval sets the limits of the values for which you would reject a null hypothesis. For example, if a confidence interval for a population mean was 100 to 105, we would reject any null hypothesis where the proposed population mean is outside of that range.
| github_jupyter |
# Stirlingの公式(対数近似)
* $\log n! \sim n\log n - n$
* $n!$はおおよそ$\left(\frac{n}{e}\right)^n$になる
* 参考: [スターリングの公式(対数近似)の導出](https://starpentagon.net/analytics/stirling_log_formula/)
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
```
## $\log n!$の上からの評価
```
MIN_X = 0.5
MAX_X = 10
x = np.linspace(MIN_X, MAX_X, 100)
y = np.log(x)
p = plt.plot(x, y, label='$\log x$')
p = plt.hlines([0], MIN_X, MAX_X)
p = plt.xlim(MIN_X, MAX_X-0.5)
p = plt.xticks(range(1, MAX_X+1))
p = plt.ylim([-0.2, 2.3])
# 面積log kの矩形を描画
for k in range(2, MAX_X):
p = plt.vlines(k, 0, np.log(k), linestyles='dashed')
p = plt.hlines(np.log(k), k, k+1, linestyles='dashed')
p = plt.legend()
plt.show(p)
```
## $\log n!$の下からの評価
```
MIN_X = 0.5
MAX_X = 10
x = np.linspace(MIN_X, MAX_X, 100)
y = np.log(x)
p = plt.plot(x, y, label='$\log x$')
p = plt.hlines([0], MIN_X, MAX_X)
p = plt.xlim(MIN_X, MAX_X-0.5)
p = plt.xticks(range(1, MAX_X+1))
p = plt.ylim([-0.2, 2.3])
# 面積log kの矩形を描画
for k in range(2, MAX_X):
p = plt.vlines(k-1, 0, np.log(k), linestyles='dashed')
p = plt.hlines(np.log(k), k-1, k, linestyles='dashed')
p = plt.vlines(MAX_X-1, 0, np.log(MAX_X), linestyles='dashed')
p = plt.legend()
plt.show(p)
```
## $n \log n - n$の近似精度
```
def log_factorial(n):
'''log n!を返す'''
val = 0.0
for i in range(1, n+1):
val += np.log(i)
return val
# test of log_factorial
eps = 10**-5
assert abs(log_factorial(1) - 0.0) < eps
assert abs(log_factorial(2) - np.log(2)) < eps
assert abs(log_factorial(5) - np.log(120)) < eps
def log_factorial_approx(n):
'''log n!の近似: n log n - nを返す'''
return n * np.log(n) - n
# test of log_factorial_approx
assert abs(log_factorial_approx(1) - (-1)) < eps
assert abs(log_factorial_approx(2) - (2 * np.log(2) - 2)) < eps
# log_factorial, log_factorial_approxをplot
n_list = range(1, 50+1)
y_fact = [log_factorial(n) for n in n_list]
y_approx = [log_factorial_approx(n) for n in n_list]
p = plt.plot(n_list, y_fact, label='$\log n!$')
p = plt.plot(n_list, y_approx, label='$n \log n - n$')
p = plt.legend()
plt.show(p)
# 近似精度を評価
n_list = [5, 10, 20, 50, 100, 1000]
approx_df = pd.DataFrame()
approx_df['n'] = n_list
approx_df['log n!'] = [log_factorial(n) for n in n_list]
approx_df['n log(n)-n'] = [log_factorial_approx(n) for n in n_list]
approx_df['error(%)'] = 100 * (approx_df['log n!'] - approx_df['n log(n)-n']) / approx_df['log n!']
pd.options.display.float_format = '{:.1f}'.format
approx_df
```
## $n!$と$\left(\frac{n}{e}\right)^n$の比較
```
n_list = [5, 10, 20, 50, 100]
approx_df = pd.DataFrame()
approx_df['n'] = n_list
approx_df['n!'] = [np.exp(log_factorial(n)) for n in n_list]
approx_df['(n/e)^n'] = [np.exp(log_factorial_approx(n)) for n in n_list]
approx_df['error(%)'] = 100 * (approx_df['n!'] - approx_df['(n/e)^n']) / approx_df['n!']
pd.options.display.float_format = None
pd.options.display.precision = 2
approx_df
```
| github_jupyter |
# Final Project: Earth Analytics Python Course, Spring 2020
Steph Shepherd & Lauren Herwehe
A Big Dam Problem:
Global Dam Watch (http://globaldamwatch.org/) maintains a database of existing (GRandD)and future (FHReD) dams across the globe. In this project we explore the future dams database by continent and country, identifying any proposed dams that will potentially impact Ramsar sites - wetlands designated of critical importance under the Ramsar Convention (1971).
## Import packages, define functions, and acquire data
```
# Imports
import warnings
import os
import sys
import numpy as np
import numpy.ma as ma
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
import matplotlib.lines as mlines
import geopandas as gpd
from geopandas import GeoDataFrame as gdf
from geopandas import GeoSeries as gs
from shapely.geometry import Point, Polygon
import contextily as ctx
import earthpy as et
import earthpy.plot as ep
# Check path and set working directory.
wd_path = os.path.join(et.io.HOME, 'earth-analytics', 'data')
if os.path.exists(wd_path):
os.chdir(wd_path)
else:
print("Path does not exist")
# Download Data stored on figshare
# Ramsar Sites
et.data.get_data(url="https://ndownloader.figshare.com/files/22507082")
# Future dams
et.data.get_data(url="https://ndownloader.figshare.com/files/22486157")
# Country boundaries
et.data.get_data(url="https://ndownloader.figshare.com/files/22507058")
# Open the ramsar shapefile with geopandas
ramsar_all = gpd.read_file(os.path.join(
"earthpy-downloads", "ramsar-site-data", "ramsar-boundaries",
"features_publishedPolygon.shp"))
# Check the crs of the ramsar sites
print(ramsar_all.crs)
# Open the dams csv files with pandas
fname = os.path.join("earthpy-downloads", "future_dams_2015.csv")
df = pd.read_csv(fname)
# Covert the pandas dataframe to a shapefile for plotting
# Set output path for shp
dams_path = os.path.join('earthpy-downloads', 'fhred-proposed-dams')
if not os.path.exists(dams_path):
os.mkdir(dams_path)
# Define the geometry for the points
geometry = [Point(xy) for xy in zip(df.Lon_Cleaned, df.LAT_cleaned)]
crs = {'init': 'epsg:4326'}
geo_df = gdf(df, crs=crs, geometry=geometry)
geo_df.to_file(driver='ESRI Shapefile', filename=os.path.join(
dams_path, 'proposed_dams.shp'))
# Open the proposed dams shapefile with geopandas
dams_all = gpd.read_file(os.path.join(dams_path, "proposed_dams.shp"))
# Pull only the columns that we need from each gdf to save processing time
proposed_dams = dams_all[['Country',
'Continent', 'Major Basi', 'Stage', 'geometry']]
ramsar_areas = ramsar_all[['country_en', 'geometry']]
# Open country borders shapefile for adding boundary of study area
country_borders_path = os.path.join("earthpy-downloads", "country-borders",
"99bfd9e7-bb42-4728-87b5-07f8c8ac631c2020328-1-1vef4ev.lu5nk.shp")
country_borders = gpd.read_file(country_borders_path)
```
# Figures 2-3: Plots of Future Dams by Continent and Selected Countries
```
# Getting Number of Dams by Continent and Development State
# Extact the columns needed for analysis
dams_continent = dams_all[['Continent', 'Country', 'Stage']]
# Group and count data by stage.
dams_stage = dams_continent.groupby(['Continent', 'Country'])[
['Stage']].count().reset_index()
dams_stage
# Group and count data by stage.
dams_stage_alt = dams_continent.groupby(['Continent', 'Country', 'Stage'])[
['Stage']].count().reset_index()
dams_stage_alt
# PLOT - NUMBER OF DAMS PROPOSED VS UNDER CONSTRUCTION BY CONTINENT
# Create a bar plot of the dams by continent and stage of process.
labels = ['Africa', 'Asia', 'Europe', 'N. America', 'Oceania', 'S. America']
proposed = [179, 937, 611, 143, 7, 1188]
under_const = [21, 424, 41, 34, 1, 114]
x = np.arange(len(labels)) # the label locations
width = 0.35 # the width of the bars
fig, ax = plt.subplots(figsize=(10, 10))
rects1 = ax.bar(x - width/2, proposed, width, label='Proposed')
rects2 = ax.bar(x + width/2, under_const, width, label='Under Construction')
# Add some text for labels, title and custom x-axis tick labels, etc.
ax.set_ylabel('Count', size=15)
ax.set_title('Figure 2: Future Dams by Continent, 2015', size=20)
ax.set_xticks(x)
ax.set_xticklabels(labels, size=15, rotation=45)
ax.legend()
autolabel(rects1)
autolabel(rects2)
ax.text(0.5, -0.2, "Data Source: Global Dam Watch Future Hydropower "
"Reservoirs and Dams Database (http://globaldamwatch.org/fhred/)",
size=12, ha="center", transform=ax.transAxes)
fig.tight_layout()
plt.show()
# Extract data by continent.
africa = dams_continent[dams_continent["Continent"] == "Africa"]
# Group and count country data by stage.
africa_stage = africa.groupby(['Country', 'Stage'])[['Stage']].count()
africa_stage
#Extract data by continent.
asia = dams_continent[dams_continent["Continent"] == "Asia"]
#Group and count country data by stage.
asia_stage = asia.groupby(['Country','Stage'])[['Stage']].count()
asia_stage
#Extract data by continent.
europe = dams_continent[dams_continent["Continent"] == "Europe"]
#Group and count country data by stage.
europe_stage = europe.groupby(['Country','Stage'])[['Stage']].count()
europe_stage
#Extract data by continent.
n_america = dams_continent[dams_continent["Continent"] == "North America"]
#Group and count country data by stage.
n_america_stage = n_america.groupby(['Country','Stage'])[['Stage']].count()
n_america_stage
#Extract data by continent.
oceania = dams_continent[dams_continent["Continent"] == "Oceania"]
#Group and count country data by stage.
oceania_stage = oceania.groupby(['Country','Stage'])[['Stage']].count()
oceania_stage
#Extract data by continent.
s_america = dams_continent[dams_continent["Continent"] == "South America"]
#Group and count country data by stage.
s_america_stage = s_america.groupby(['Country','Stage'])[['Stage']].count()
s_america_stage
# Create a bar plot of the dams by countries in Africa comparing stage of process.
africa_labels = ['Benin', 'Burkina Faso', 'Gabon', 'Guinea', 'Malawi', 'Mali', 'Morocco',
'Mozambique', 'Namibia', 'Niger', 'Nigeria', 'Rwanda', 'South Africa', 'Zimbabwe']
africa_proposed = [6, 2, 1, 23, 2, 12, 0, 3, 3, 1, 1, 0, 3, 2]
africa_under_const = [0, 0, 1, 0, 0, 1, 2, 1, 0, 1, 2, 1, 0, 1]
x = np.arange(len(africa_labels)) # the label locations
width = 0.35 # the width of the bars
fig, ax = plt.subplots(figsize=(10, 10))
rects1 = ax.bar(x - width/2, africa_proposed, width, label='Proposed')
rects2 = ax.bar(x + width/2, africa_under_const,
width, label='Under Construction')
# Add some text for labels, title and custom x-axis tick labels, etc.
ax.set_ylabel('Count', size=15)
ax.set_title(
'Figure 3A: Future Dam Construction by Selected Countries in Africa', size=20)
ax.set_xticks(x)
ax.set_xticklabels(africa_labels, size=15, rotation=45)
ax.legend()
def autolabel(rects):
"""Attach a text label above each bar in *rects*, displaying its value."""
for rect in rects:
height = rect.get_height()
ax.annotate('{}'.format(height),
xy=(rect.get_x() + rect.get_width() / 2, height),
xytext=(0, 3), # 3 points vertical offset
textcoords="offset points",
ha='center', va='bottom')
autolabel(rects1)
autolabel(rects2)
ax.text(0.5, -0.2, "Data Source: Global Dam Watch Future Hydropower Reservoirs "
"and Dams Database (http://globaldamwatch.org/fhred/)",
size=12, ha="center", transform=ax.transAxes)
fig.tight_layout()
plt.show()
# Create a bar plot of the dams by country in N. America comparing stage of process.
na_labels = ['Belize', 'Canada', 'Costa Rica', 'El Salvador', 'Guatemala', 'Haiti', 'Honduras', 'Mexico', 'Nicaragua', 'Panama', 'United States']
na_proposed = [1, 26, 40, 1, 0, 1, 0, 4, 17, 43, 10]
na_under_const = [0, 8, 3, 0, 2, 0, 2, 3, 1, 15, 0]
x = np.arange(len(na_labels)) # the label locations
width = 0.35 # the width of the bars
fig, ax = plt.subplots(figsize=(10, 10))
rects1 = ax.bar(x - width/2, na_proposed, width, label='Proposed')
rects2 = ax.bar(x + width/2, na_under_const, width, label='Under Construction')
# Add some text for labels, title and custom x-axis tick labels, etc.
ax.set_ylabel('Count', size=15)
ax.set_title('Figure 3B: Future Dam Construction by Countries in N. America', size=20)
ax.set_xticks(x)
ax.set_xticklabels(na_labels, size=15, rotation=45)
ax.legend()
autolabel(rects1)
autolabel(rects2)
fig.tight_layout()
plt.show()
```
# Overlay future dams and Ramsar sites datasets for Africa.
```
# Change the datas CRS to projected for Africa (WGS 84 World Mercator)
# To make this data more accurate, for the next course we can create
#list of EPSG for each country in Africa to include in function
proposed_dams = proposed_dams.to_crs('epsg:3765')
ramsar_areas = ramsar_areas.to_crs('epsg:3765')
# Get dam impact by African country
# List of African country names
# Data cleaning issues: Removed 'Côte d'Ivoire' bc in Ramsar dataset it's called Cite D'ivore and don't know how to deal with additional ' in a string; also removed Congo bc in Ramsar it's called Congo & Democratic Republic of Congo and in FhRED it's called Congo, Rep.
africa_cntry = ['Algeria', 'Angola', 'Benin', 'Botswana', 'Burkina Faso',
'Burundi', 'Cabo Verde', 'Cameroon', 'Central African Republic',
'Chad', 'Comoros', 'Djibouti', 'Equatorial Guinea', 'Eritrea',
'Ethiopia', 'Gabon', 'Gambia', 'Ghana', 'Guinea', 'Guinea-Bissau',
'Kenya', 'Lesotho', 'Liberia', 'Libya', 'Madagascar', 'Malawi',
'Mali', 'Mauritania', 'Mauritius', 'Morocco', 'Mozambique', 'Namibia',
'Niger', 'Nigeria', 'Rwanda', 'Sao Tome and Principe', 'Senegal',
'Seychelles', 'Sierra Leone', 'Somalia', 'South Africa', 'South Sudan',
'Sudan', 'Tanzania', 'Togo', 'Tunisia', 'Uganda', 'Zambia', 'Zimbabwe']
# Empty Africa dict
africa_dams = {}
# Append dam_impact function data to africa_dams
for i in africa_cntry:
try:
africa_dams[i] = {"5km Buffer Area": dam_impact(
5, i), "10km Buffer Area": dam_impact(10, i)}
except:
dams = None
# Turn it into a pandas dataframe for plotting
africa_df = pd.DataFrame.from_dict(africa_dams)
# Some values in the dataframe are zero and some are NaN, make it the same
africa_df = africa_df.fillna('None')
africa_df.replace({0: 'None'})
#Plot data to illustrate which countires have potential imapcts from proposed dams.
# Create legend so only countries with potential impacts are listed.
blue_patch = mpatches.Patch(color='dodgerblue', label='Beinin')
green_patch = mpatches.Patch(color='green', label='Gabon')
red_patch = mpatches.Patch(color='red', label='Guinea')
teal_patch = mpatches.Patch(color='c', label='Niger')
# Create the figure
fig, ax = plt.subplots(figsize=(8, 8))
africa_df.plot(ax=ax, kind='barh', stacked=True, legend=True)
# Add some text for labels, title and custom x-axis tick labels, etc.
ax.set_xlabel('Affected Area (km)', size=15)
ax.set_title('Figure 4: Dam Impact on Ramsar Areas in Africa', size=20)
ax.text(0.5, -0.2, "Data Sources: Global Dam Watch Future Hydropower Reservoirs "
"and Dams Database (http://globaldamwatch.org/fhred/),\nRamsar Sites "
"Information Service (https://rsis.ramsar.org/)",
size=12, ha="center", transform=ax.transAxes)
ax.legend(handles=[blue_patch, green_patch, red_patch, teal_patch],
fontsize=15,
frameon=True,
loc=('lower right'),
title="Country")
```
# Map buffer results for Guinea
```
# Analyze Guinea
# Pull only the data for Guinea
proposed_dams_guin = proposed_dams[proposed_dams['Country'] == "Guinea"]
ramsar_areas_guin = ramsar_areas[ramsar_areas['country_en'] == "Guinea"]
guinea_border = country_borders[country_borders['CNTRY_NAME'] == "Guinea"]
# Get the CRS right for plotting
proposed_dams_guin = proposed_dams_guin.to_crs('epsg:3462')
ramsar_areas_guin = ramsar_areas_guin.to_crs('epsg:3462')
guinea_border = guinea_border.to_crs('epsg:3462')
# Buffer the dams to 5km & 10km for plotting
proposed_dams_guin_5k_buff = proposed_dams_guin.buffer(5000)
proposed_dams_guin_10k_buff = proposed_dams_guin.buffer(10000)
# Create a map of the dams and the ramsar sites for Guinea
black_line = mlines.Line2D([], [], color='black', label='Country Border')
yellow_patch = mpatches.Patch(color='yellow', label='Ramsar Area')
green_circle = mlines.Line2D([], [], color='white', marker='o',
markerfacecolor='forestgreen', markersize=18,
label='10km Buffer')
lime_circle = mlines.Line2D([], [], color='white', marker='o',
markerfacecolor='lime', markersize=12,
label='5km Buffer')
red_dot = mlines.Line2D([], [], color='white', marker='o',
markerfacecolor='red', label='Proposed Dam Site')
fig, ax = plt.subplots(figsize=(15, 15))
ramsar_areas_guin.plot(ax=ax, facecolor='yellow')
proposed_dams_guin_10k_buff.plot(facecolor='forestgreen',
ax=ax)
proposed_dams_guin_5k_buff.plot(facecolor='lime',
ax=ax)
proposed_dams_guin.plot(ax=ax,
markersize=5,
color='red')
guinea_border.plot(ax=ax, color="none", edgecolor="black", linewidth=2)
ax.legend(handles=[black_line, yellow_patch, green_circle, lime_circle, red_dot],
fontsize=15,
frameon=True,
loc=('upper right'),
title="LEGEND")
ctx.add_basemap(ax, url=ctx.providers.Stamen.Terrain, zoom=0)
ax.set_axis_off()
ax.set_title(
'Figure 5: Guinea Ramsar Areas, Proposed Dams, and Dam Buffer Areas', size=20)
ax.text(0.5, -0.1, "Data Sources: Global Dam Watch Future Hydropower Reservoirs "
"and Dams Database (http://globaldamwatch.org/fhred/), \n Ramsar Sites "
"Information Service (https://rsis.ramsar.org/)",
size=12, ha="center", transform=ax.transAxes)
```
| github_jupyter |
```
%%HTML
<link rel="stylesheet" type="text/css" href="style/template.css">
```
# 今さらOpen MVを触ってみた ーその2ー
## K.Miura
# 自己紹介
- 三浦 耕生(こうき)
- 大学院生、ロボット工学専攻
- TechAcademyのジュニアメンター
- Twitter: @k_miura_io
- Facebook: koki.miura05
# スライドのリンク
※このスライドはjupyter-notebookを使用しています
```
print("Hello World")
```
# さて、本題です
# 前回までのあらすじ
- kickstarterでOpen MVを買ったので遊んでみた
- 小さいのに顔認識がサクサク動いて面白かった
<div align="center">
<img src="./imgs/IMG_7078.jpeg" width=30%>
</div>
# でその翌日
# 東京でイベントにてブース出展
- 朝一の新幹線で東京で1日IBMのブースでお手伝い(詳しくは懇親会で)
- その帰りに久々に秋葉原を散策してたらM5Stickを発見
- 前から気になってたしつい買ってしまった
<div align="center">
<img src="./imgs/img001.jpeg" width=15%>
</div>
# というわけで
# ~今さらOpen MVを触ってみた ーその2ー~
# 今さらM5Stick-Vを触ってみた
## K.Miura
# Maker界隈でアツイM5シリーズ
- 様々な機種があり、Twitterで遊んでみた投稿が多く存在する
- M5Stick-Vでも画像認識を使った作品が豊富にある
<div align="center">
<img src="https://media.giphy.com/media/Tfp1lxLUrStcYv2Cud/giphy.gif" width="15%">
</div>
# どうやらM5Stick-V用のモデルを作成するサイトがあるらしい
- その名も**V-Training**
- トレーニング用のファームウェアで画像を撮影して、サイトにアップロードすると学習モデルをダウンロードして自作の画像分類機を簡単にできるらしい
<div align="center">
<img src="./imgs/V-traning.png" width="50%">
</div>
# 使ってみた
# その前に今回のネタ
- 最近カルロス・ゴーンが何かと話題になっている
- カルロス・ゴーンってMr.Beanと顔がよく似ている
- この2人の顔を識別してみよう
<div align="center">
<img src="./imgs/0035.jpg">
</div>
# データ作成
- スクレイピングでgoogleの画像検索で出てきた画像を取得
- V-Traningを使う場合は1クラスあたり最低35枚の画像が必要
- 最大10クラスを学習させることが可能
```
import argparse
import json
import os
import urllib
from bs4 import BeautifulSoup
import requests
from termcolor import cprint
__version__ = "1.0.0"
class GoogleImageSerch(object):
def __init__(self):
self.GOOGLE_IMAGE_SEARCH_URL = "https://www.google.co.jp/search"
self.session = requests.session()
self.session.headers.update(
{
"User-Agent": "Mozilla/5.0 (X11; Linux x86_64; rv:10.0) \
Gecko/20100101 Firefox/10.0"
}
)
def search(self, keyword, maximum):
print(f"Searching {keyword}.")
query = self.generate_query(keyword)
return self.serch_images(query, maximum)
def generate_query(self, keyword):
# search query generator
page = 0
while True:
params = urllib.parse.urlencode(
{"q": keyword, "tbm": "isch", "ijn": str(page)}
)
yield self.GOOGLE_IMAGE_SEARCH_URL + "?" + params
page += 1
def serch_images(self, generate_query, maximum):
results = []
total = 0
while True:
# search
html = self.session.get(next(generate_query)).text
soup = BeautifulSoup(html, "lxml")
elements = soup.select(".rg_meta.notranslate")
jsons = [json.loads(e.get_text()) for e in elements]
image_url_list = [js["ou"] for js in jsons]
# add search results
if not image_url_list:
cprint("No more images.", "yellow")
break
elif len(image_url_list) > maximum - total:
results += image_url_list[: maximum - total]
break
else:
results += image_url_list
total += len(image_url_list)
cprint(f"Found {len(results)} images.", "green")
return results
def main(args):
os.makedirs(args.download_dir, exist_ok=True)
os.makedirs(os.path.join(args.download_dir, args.target_name), exist_ok=True)
google_image_serch = GoogleImageSerch()
# search images
results = google_image_serch.search(args.target_name, maximum=args.num_images)
# download
download_errors = []
for i, url in enumerate(results):
download_name = f"{(i + 1):>0{max(4, len(str(args.num_images)))}}.jpg"
download_path = os.path.join(args.download_dir, args.target_name, download_name)
if os.path.exists(download_path) and not args.is_overwrite:
print(f"{download_path} is already exists.")
download_errors.append(i + 1)
continue
print(f"Downloading image {download_name}.", end=" ")
try:
urllib.request.urlretrieve(url, download_path)
cprint("Successful.", "green")
except urllib.error.HTTPError:
cprint("Failed. (HTTP Error)", "yellow")
download_errors.append(i + 1)
continue
except urllib.error.URLError:
cprint("Failed. (SSL Error)", "yellow")
download_errors.append(i + 1)
continue
except UnicodeEncodeError:
cprint("Failed. (Encoding Error)", "yellow")
download_errors.append(i + 1)
continue
cprint("Download complete.", "blue")
cprint(f"Successful: {len(results) - len(download_errors)} images.", "blue")
if download_errors:
cprint(f"Failed: {len(download_errors)} images.", "yellow")
if __name__ == "__main__":
cprint("-" * 50, "magenta")
cprint((f"Image Collector v{__version__}").center(50), "magenta")
cprint("-" * 50, "magenta")
parser = argparse.ArgumentParser(description=f"Image Collector v{__version__}")
parser.add_argument(
"-t",
"--target",
dest="target_name",
help="Target name",
type=str,
required=True,
)
parser.add_argument(
"-n",
"--number",
dest="num_images",
help="Number of images",
type=int,
required=True,
)
parser.add_argument(
"-d",
"--directory",
dest="download_dir",
help="Download location",
type=str,
default="./data",
)
parser.add_argument(
"-f",
"--force",
dest="is_overwrite",
action="store_true",
help="Whether to overwrite existing files",
)
args = parser.parse_args()
main(args)
```
# Let's Take Photo
- そのまま学習させてもいいけど、M5目線の画像を学習したいので、M5Stick-Vを撮影して学習
- データ作成をするためのプログラムはV-TraningのサイトからダウンロードしてSDカードに保存して動かす
- とりあえず最低量の35枚の画像を収集
<div align="center">
<img src="./imgs/IMG_7358.JPG" width="20%">
</div>
# 画像データをアップロード
- 専用のサイトへSDに保存されたデータセットを圧縮してアップロード
- アップロードするとステータスIDを付与されて裏側で学習される
- 学習が完了するとメールアドレスに学習したモデルと実行用のコードをセットで送られる
<div align="center">
<img src="./imgs/0002.png" width="50%">
</div>
# 1回目の学習結果
- データの質が悪い&量が少ないせいで精度が悪かった
- 「もう1回送ったほうがいいよ」と一言アドバイス ~(余計なお世話)~
- 35枚の画像では精度がでるような仕様ではないようだ
<div align="center">
<img src="./imgs/first_results.png" width="50%">
</div>
# もう一度データを用意
- 今度は画像を100枚用意して撮影
- もちろんキーワードと関係ない画像も含まれているので実質学習に使えたのは60枚程度
- あとは単純に疲れた(笑)
# 2回目の学習結果
<div align="center">
<img src="./imgs/new_results.png" width="50%">
</div>
```
from PIL import Image
im1 = Image.open("./imgs/0006.png")
im2 = Image.open("./imgs/0005.png")
def get_concat_h(im1, im2):
dst = Image.new('RGB', (im1.width + im2.width, im1.height))
dst.paste(im1, (0, 0))
dst.paste(im2, (im1.width, 0))
return dst
get_concat_h(im1, im2).save('./imgs/new_results.png')
```
# END
| github_jupyter |
```
!date
```
# Figure 1b and 1d
```
import glob
import pandas as pd
import numpy as np
import pandas as pd
import scipy as scp
import sklearn
import itertools
from scipy.optimize import fsolve
from upsetplot import generate_data, plot, from_memberships
from collections import Counter
from matplotlib.ticker import FormatStrFormatter
from matplotlib.ticker import StrMethodFormatter
from mpl_toolkits.axes_grid1.inset_locator import inset_axes
import matplotlib.patches as mpatches
import matplotlib.ticker as ticker
import matplotlib.pyplot as plt
import matplotlib
matplotlib.rcParams.update({'font.size': 22})
%config InlineBackend.figure_format = 'retina'
v2_names = np.array(['SRR8599150_v2',
'heart1k_v2', 'SRR8611943_v2',
'SRR8257100_v2', 'EMTAB7320_v2',
'SRR7299563_v2', 'SRR8513910_v2',
'SRR8639063_v2', 'SRR8524760_v2',
'SRR6956073_v2', 'hgmm1k_v2',
'SRR8206317_v2', 'SRR8327928_v2',
'SRR6998058_v2'], dtype=object)
v3_names = np.array(['pbmc_1k_v3', 'hgmm10k_v3',
'neuron_10k_v3', 'pbmc_10k_v3',
'heart1k_v3', 'hgmm1k_v3'], dtype=object)
v2_data = {}
v2_data["EMTAB7320_v2"] = {"n_reads": 335147976}
v2_data["heart1k_v2"] = {"n_reads": 88872840}
v2_data["hgmm1k_v2"] = {"n_reads": 75225120}
v2_data["SRR6956073_v2"] = {"n_reads": 161274652}
v2_data["SRR6998058_v2"] = {"n_reads": 37227612}
v2_data["SRR7299563_v2"] = {"n_reads": 112176350}
v2_data["SRR8206317_v2"] = {"n_reads": 85992089}
v2_data["SRR8257100_v2"] = {"n_reads": 189337914}
v2_data["SRR8327928_v2"] = {"n_reads": 190094560}
v2_data["SRR8513910_v2"] = {"n_reads": 146617182}
v2_data["SRR8524760_v2"] = {"n_reads": 97106426}
v2_data["SRR8599150_v2"] = {"n_reads": 8860361}
v2_data["SRR8611943_v2"] = {"n_reads": 21574502}
v2_data["SRR8639063_v2"] = {"n_reads": 416437344}
v2_data["EMTAB7320_v2"]["n_cells"] = 4510
v2_data["heart1k_v2"]["n_cells"] = 712
v2_data["hgmm1k_v2"]["n_cells"] = 1079
v2_data["SRR6956073_v2"]["n_cells"] = 4168
v2_data["SRR6998058_v2"]["n_cells"] = 575
v2_data["SRR7299563_v2"]["n_cells"] = 1660
v2_data["SRR8206317_v2"]["n_cells"] = 4418
v2_data["SRR8257100_v2"]["n_cells"] = 11685
v2_data["SRR8327928_v2"]["n_cells"] = 10396
v2_data["SRR8513910_v2"]["n_cells"] = 726
v2_data["SRR8524760_v2"]["n_cells"] = 3064
v2_data["SRR8599150_v2"]["n_cells"] = 3949
v2_data["SRR8611943_v2"]["n_cells"] = 5194
v2_data["SRR8639063_v2"]["n_cells"] = 6614
v3_data = {}
v3_data["hgmm1k_v3"] = {"n_reads": 63105786}
v3_data["neuron_10k_v3"] = {"n_reads": 357111595}
v3_data["pbmc_10k_v3"] = {"n_reads": 638901019}
v3_data["pbmc_1k_v3"] = {"n_reads": 66601887}
v3_data["heart1k_v3"] = {"n_reads": 84512390}
v3_data["hgmm10k_v3"] = {"n_reads": 721180737}
v3_data["hgmm1k_v3"]["n_cells"] = 1011
v3_data["neuron_10k_v3"]["n_cells"] = 11477
v3_data["pbmc_10k_v3"]["n_cells"] = 1045
v3_data["pbmc_1k_v3"]["n_cells"] = 11790
v3_data["heart1k_v3"]["n_cells"] = 11692
v3_data["hgmm10k_v3"]["n_cells"] = 1227
w = 67365891
c = 345420
u = 2013414
v2_data["heart1k_v2"]["barcode_error_correction"] = (w, c, u)
w = 57345535
c = 176786
u = 1849405
v3_data["heart1k_v3"]["barcode_error_correction"] = (w, c, u)
w = 58523823
c = 358110
u = 2035210
v2_data["hgmm1k_v2"]["barcode_error_correction"] = (w, c, u)
w = 46243317
c = 132278
u = 1394347
v3_data["hgmm1k_v3"]["barcode_error_correction"] = (w, c, u)
w = 499346666
c = 2613284
u = 20298095
v3_data["hgmm10k_v3"]["barcode_error_correction"] = (w, c, u)
w = 227709973
c = 659929
u = 7299697
v3_data["neuron_10k_v3"]["barcode_error_correction"] = (w, c, u)
w = 353379492
c = 1912254
u = 14819352
v3_data["pbmc_10k_v3"]["barcode_error_correction"] = (w, c, u)
w = 39178903
c = 190366
u = 1538993
v3_data["pbmc_1k_v3"]["barcode_error_correction"] = (w, c, u)
w = 28344188
c = 231718
u = 625557
v2_data["SRR6998058_v2"]["barcode_error_correction"] = (w, c, u)
w = 66294966
c = 782287
u = 1728840
v2_data["SRR8206317_v2"]["barcode_error_correction"] = (w, c, u)
w = 111254198
c = 1567548
u = 4904318
v2_data["SRR8327928_v2"]["barcode_error_correction"] = (w, c, u)
w = 348557155
c = 1857224
u = 1836077
v2_data["SRR8639063_v2"]["barcode_error_correction"] = (w, c, u)
w = 258864227
c = 4111830
u = 9256167
v2_data["EMTAB7320_v2"]["barcode_error_correction"] = (w, c, u)
w = 107572180
c = 1082195
u = 2639035
v2_data["SRR6956073_v2"]["barcode_error_correction"] = (w, c, u)
w = 64690144
c = 477618
u = 1520183
v2_data["SRR7299563_v2"]["barcode_error_correction"] = (w, c, u)
w = 173540630
c = 1094514
u = 4191648
v2_data["SRR8257100_v2"]["barcode_error_correction"] = (w, c, u)
w = 131004911
c = 910116
u = 3772762
v2_data["SRR8513910_v2"]["barcode_error_correction"] = (w, c, u)
w = 3420063
c = 38493
u = 117197
v2_data["SRR8599150_v2"]["barcode_error_correction"] = (w, c, u)
w = 16021922
c = 206410
u = 518515
v2_data["SRR8611943_v2"]["barcode_error_correction"] = (w, c, u)
w = 68514365
c = 615351
u = 1748491
v2_data["SRR8524760_v2"]["barcode_error_correction"] = (w, c, u)
%%time
for f in glob.glob("/home/sina/projects/bus/validate/all_bus/k*"):
name = "_".join(f.split("/")[7].split("_")[1:])
print(name)
if "v2" in name:
# loss counts
v2_data[name]["loss_ratio"] = pd.read_csv(f + "/lossratio.txt", header=None).values.flatten()
elif "v3" in name:
v3_data[name]["loss_ratio"] = pd.read_csv(f + "/lossratio.txt", header=None).values.flatten()
```
# Figure 1b
```
# (inwhitelist, correct, uncorrected)
w = [v2_data[i]["barcode_error_correction"][0]/(v2_data[i]["barcode_error_correction"][0] + v2_data[i]["barcode_error_correction"][1] + v2_data[i]["barcode_error_correction"][2]) for i in v2_names]
[w.append(v3_data[i]["barcode_error_correction"][0]/(v3_data[i]["barcode_error_correction"][0] + v3_data[i]["barcode_error_correction"][1] + v3_data[i]["barcode_error_correction"][2])) for i in v3_names]
c = [v2_data[i]["barcode_error_correction"][1]/(v2_data[i]["barcode_error_correction"][0] + v2_data[i]["barcode_error_correction"][1] + v2_data[i]["barcode_error_correction"][2]) for i in v2_names]
[c.append(v3_data[i]["barcode_error_correction"][1]/(v3_data[i]["barcode_error_correction"][0] + v3_data[i]["barcode_error_correction"][1] + v3_data[i]["barcode_error_correction"][2])) for i in v3_names]
u = [v2_data[i]["barcode_error_correction"][2]/(v2_data[i]["barcode_error_correction"][0] + v2_data[i]["barcode_error_correction"][1] + v2_data[i]["barcode_error_correction"][2]) for i in v2_names]
[u.append(v3_data[i]["barcode_error_correction"][2]/(v3_data[i]["barcode_error_correction"][0] + v3_data[i]["barcode_error_correction"][1] + v3_data[i]["barcode_error_correction"][2])) for i in v3_names]
nreads = [v2_data[i]["n_reads"] for i in v2_names]
[nreads.append(v3_data[i]["n_reads"]) for i in v3_names]
idx_sorted = np.argsort(nreads)
names = v2_names
n3 = v3_names
names = np.append(names, n3)
names = names[idx_sorted]
sorted_nreads = np.sort(nreads)
w = np.array(w)[idx_sorted]
c = np.array(c)[idx_sorted]
u = np.array(u)[idx_sorted]
data = [w, c, u]
p = data[1]/(16*data[0] + data[1])
fig, ax=plt.subplots(figsize=(15, 7))
width=0.8
p1 = ax.bar(names, data[0], width, color='white', label="Retained", edgecolor="black")
p2 = ax.bar(names, data[1], width, bottom=data[0], color='black', label="Corrected", edgecolor="black")
p3 = ax.bar(names, data[2], width,
bottom=np.array(data[0])+np.array(data[1]), color='#cccccc', label="Uncorrected", edgecolor="black")
plt.xticks(rotation=45, ha="right")
xlocs, xlabs = plt.xticks()
for i, v in enumerate(["{:.3f}".format(i) for i in 16*p*(1-p)**15]):
plt.text(xlocs[i]-0.4, 1 + 0.01, str(v), size=12)
ax.set_ylim(bottom=0)
plt.ylabel("Fraction of total barcodes")
plt.legend(loc="lower right")
plt.gca().yaxis.set_major_formatter(StrMethodFormatter('{x:,.1f}'))
plt.subplots_adjust(bottom=0.15, wspace=0.05)
plt.tight_layout()
#plt.savefig("barcodecorrection.pdf")
plt.show()
```
### Data from Figure 1
```
for i in range(len(names)):
print(p[i], names[i])
p.mean()
(16*p*(1-p)**15).mean()
(120*p**2*(1-p)**14).mean()*100
w = [v2_data[i]["barcode_error_correction"][0] for i in v2_names]
[w.append(i) for i in [v3_data[i]["barcode_error_correction"][0] for i in v3_names]]
w = np.array(w)
b = [v2_data[i]["barcode_error_correction"][1] for i in v2_names]
[b.append(i) for i in [v3_data[i]["barcode_error_correction"][1] for i in v3_names]]
b = np.array(b)
g = [v2_data[i]["barcode_error_correction"][2] for i in v2_names]
[g.append(i) for i in [v3_data[i]["barcode_error_correction"][2] for i in v3_names]]
g = np.array(g)
t = w+b+g
w/(1-(p)**16)
t
b/w/(1-(p)**16)
16*p*(1-p)**15
10*p*(1-p)**9*100
(c/(w+c+u)*100).mean()
```
# Figure 1d
```
data = [v2_data[i]["loss_ratio"] for i in v2_names]
[data.append(v3_data[i]["loss_ratio"]) for i in v3_names]
names = v2_names
n3 = v3_names
names = np.append(names, n3)
nreads = [v2_data[i]["n_reads"] for i in v2_names]
[nreads.append(v3_data[i]["n_reads"]) for i in v3_names]
idx_sorted = np.argsort(nreads)
sorted_nreads = np.sort(nreads)
data = np.array(data)[idx_sorted]
names = names[idx_sorted]
kallisto_color_v2 = '#cccccc'
kallisto_color_v3 = 'black'
def adjacent_values(vals, q1, q3):
upper_adjacent_value = q3 + (q3 - q1) * 1.5
upper_adjacent_value = np.clip(upper_adjacent_value, q3, vals[-1])
lower_adjacent_value = q1 - (q3 - q1) * 1.5
lower_adjacent_value = np.clip(lower_adjacent_value, vals[0], q1)
return lower_adjacent_value, upper_adjacent_value
def set_axis_style(ax, labels):
ax.get_xaxis().set_tick_params(direction='out')
ax.xaxis.set_ticks_position('bottom')
ax.set_xticks(np.arange(1, len(labels) + 1))
ax.set_xticklabels(labels)
ax.set_xlim(0.25, len(labels) + 0.75)
ax.set_ylim(0, 1)
#ax.set_xlabel('Sample name')
fig, ax = plt.subplots(figsize=(20, 10), nrows=1, ncols=1)
################
parts = ax.violinplot(data, showmeans=False, showmedians=False,showextrema=False)
for pc_i in range(len(parts['bodies'])):
if "v2" in names[pc_i]:
parts['bodies'][pc_i].set_facecolor(kallisto_color_v2)
elif "v3" in names[pc_i]:
parts['bodies'][pc_i].set_facecolor(kallisto_color_v2)
parts['bodies'][pc_i].set_edgecolor('black')
parts['bodies'][pc_i].set_alpha(1)
# set style for the axes
set_axis_style(ax, names)
inset = inset_axes(ax, width="50%", height="50%", loc=1)
parts = inset.violinplot(data, showmeans=False, showmedians=False,showextrema=False)
for pc_i in range(len(parts['bodies'])):
if "v2" in names[pc_i]:
parts['bodies'][pc_i].set_facecolor(kallisto_color_v2)
elif "v3" in names[pc_i]:
parts['bodies'][pc_i].set_facecolor(kallisto_color_v3)
parts['bodies'][pc_i].set_edgecolor('black')
parts['bodies'][pc_i].set_alpha(1)
set_axis_style(inset, names)
inset.set_xticklabels([])
inset.set_ylim(0, 0.001)
ticks = ax.get_yticks().tolist()
ticks = ["{:.1f}".format(i) for i in ticks]
ticks[0] = "No counts lost 0.0"
ticks[-1] = "All counts lost 1.0"
#for tick in ax.get_xticklabels():
# tick.set_rotation(45, ha="right")
ax.set_xticklabels(ax.xaxis.get_majorticklabels(), rotation=45, ha="right")
grey_patch = mpatches.Patch(color=kallisto_color_v2, label='v2 Chemistry')
black_patch = mpatches.Patch(color=kallisto_color_v3, label='v3 Chemistry')
ax.figure.legend(handles=[grey_patch, black_patch], loc=1, bbox_to_anchor=(0.34,0.94))
ax.yaxis.set_ticklabels(ticks)
plt.subplots_adjust(bottom=0.15, wspace=0.05)
plt.tight_layout()
plt.savefig("lossratio.pdf")
plt.show()
```
### Data from Figure 1
```
loss2 = [v2_data[i]["loss_ratio"] for i in v2_names]
loss3 = [v3_data[i]["loss_ratio"] for i in v3_names]
for i in range(len(v2_names)):
print("{:,.4f}% \t {}".format(loss2[i].mean()*100, v2_names[i]))
for i in range(len(v3_names)):
print("{:,.4f}% \t {}".format(loss3[i].mean(), v3_names[i]))
(np.array([loss2[i].mean() for i in range(len(loss2))])).mean()*100
(np.array([loss3[i].mean() for i in range(len(loss3))])).mean()*100
print("Number of Reads per Cell")
print("------------------------")
for i in v2_names:
print("{:,.0f} \t {}".format(v2_data[i]["n_reads"]/v2_data[i]["n_cells"], i))
print("Number of Reads per Cell")
print("------------------------")
for i in v3_names:
print("{:,.0f} \t {}".format(v3_data[i]["n_reads"]/v3_data[i]["n_cells"], i))
```
| github_jupyter |
```
try:
# %tensorflow_version only exists in Colab.
%tensorflow_version 2.x
except Exception:
pass
```
We need to tell Google Colab that we want the TF 2.0 version so the code can work properly.
```
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
import random
tf.executing_eagerly()
```
We also need to import our required libraries so we can use them in the next parts.
```
a1 = tf.random.uniform(shape=[], minval=-5, maxval=5)
b1 = tf.random.uniform(shape=[], minval=-5, maxval=5)
c1 = tf.random.uniform(shape=[], minval=-5, maxval=5)
```
At this part we get some random values for a, b and c variables since our polynomial equation looks something like this: a*x^2 + b*x + c
```
a2 = tf.random.uniform(shape=[], minval=-5, maxval=5)
b2 = tf.random.uniform(shape=[], minval=-5, maxval=5)
c2 = tf.random.uniform(shape=[], minval=-5, maxval=5)
xs = tf.constant(range(0, 20), dtype=tf.float32)
ys = tf.constant(tf.add(tf.add(tf.multiply(tf.pow(xs, 2), a2), tf.multiply(xs, b2)), c2), dtype=tf.float32)
print(f"Start values: \nModel: ({a1})*x^2 + ({b1})*x + ({c1})\nRandom Values: ({a2})*x^2 + ({b2})*x + ({c2})")
# xs = tf.constant([1.0, 2.0, 3.0, 4.0, 5.0, 6.0, 7.0, 8.0, 9.0, 10.0], dtype=tf.float32)
# ys = tf.constant([45000.0, 50000.0, 60000.0, 80000.0, 110000.0, 150000.0, 200000.0, 300000.0, 500000.0, 1000000.0], dtype=tf.float32)
```
You can either use the upper one or the bottom one while trying this out. The top one generates another random polynomial equation which we then try to find using our model. The bottom one on the other hand, is kind of well known dataset for trying out the polynomial regression. You may come accross that dataset whenever you are looking for polynomial regression tutorials.
```
plt.plot(xs, ys, 'bo')
```
Plotting the X and Y values before training our model so we can get an idea of how our data looks like.
```
def mean_squared_error(predictions, labels):
return tf.reduce_mean(tf.square(predictions - labels))
def stochastic_gradient_descent_optimizer(indexes, labels , predictions):
result = tf.reduce_mean(2 * indexes * (predictions - labels)).numpy()
print(f"SGD --> Indexes: {indexes.numpy()} | Labels: {labels.numpy()} | Predictions: {predictions.numpy()} | Result: {result}")
return result
def predict(indexes):
prediction = tf.add(tf.add(tf.multiply(tf.pow(indexes, 2), a1), tf.multiply(indexes, b1)), c1)
print(f"Incoming: {indexes.numpy()} | Prediction: {prediction.numpy()}")
return prediction
```
Here, we declare our 3 main functions we need for our "thing" to become a bit of "Machine Learning Model". First one is Mean Squared Error. This function will tell a number based on how off our predictions are from the actual values. Next one is Stochastic Gradient Descent. This function will be acting as our optimizer in our model so we will be changing our a1, b1 and c1 values based on this value. And the final and the most important one(yes, all of them are very important but they won't make any sense without this one :D), our connection to model! With the prediction function, we can comminucate with our model and ask for predictions from it.
```
EPOCHS = 25
SAMPLES = xs.shape[0]
BATCH_SIZE = 1
LEARNING_RATE = 0.0001
dataset = tf.data.Dataset.from_tensor_slices((xs , ys))
dataset = dataset.repeat(EPOCHS).batch(BATCH_SIZE)
iterator = dataset.__iter__()
```
At this step, we are preparing our dataset to become iterable so we can train our model with the batches of data we just make here.
```
num_features = len(xs)
epochs_plot = list()
loss_plot = list()
for i in range(EPOCHS):
epoch_loss = list()
for Q in range(int(SAMPLES/BATCH_SIZE)):
x_batch, y_batch = iterator.get_next()
output = predict(x_batch)
loss_val = epoch_loss.append(mean_squared_error(y_batch , output).numpy())
deriv_val = stochastic_gradient_descent_optimizer(x_batch, y_batch , output)
# print(f"deriv_val: {deriv_val}")
a1 -= (LEARNING_RATE * deriv_val)
b1 -= (LEARNING_RATE * deriv_val)
c1 -= (LEARNING_RATE * deriv_val)
loss_val = np.array(epoch_loss).mean()
epochs_plot.append(i + 1)
loss_plot.append(loss_val)
print('Loss is {}'.format(loss_val))
```
And yet another very important step, training! At this step, we train our model using the functions we have defined a few steps ago.
```
plt.plot(epochs_plot, loss_plot)
plt.show()
```
Here, we can see how our loss value lowered as we trained our model at each epoch.
```
polynomial_points = list()
for i in range(len(xs)):
polynomial_points.append(predict(xs[i]).numpy())
plt.plot(xs, ys, 'bo', xs, polynomial_points, 'r')
```
And the final step! First, we just predict our values on the same X values as our dataset so we can match each other in the plot. After making the predictions, we just plot them both together and voila! We have just created and trained our own model for polynomial regression! You can get more information about this project (such as the math behind MSE and SGD) at my blog post which you can go with [this](https://blog.tekno.icu/2020/01/22/polynomial-regression-in-python-using-tensorflow-2-0/) link.
We can also check how close our model became to the randomly generated quadratic equation if you have chosen to generate a random quadratic equation at the first step.
```
print(f"End values: \nModel: ({a1})*x^2 + ({b1})*x + ({c1})\nRandom Values: ({a2})*x^2 + ({b2})*x + ({c2})")
```
| github_jupyter |
<font size="+5">#09. Cluster Analysis con k-Means</font>
- Book + Private Lessons [Here ↗](https://sotastica.com/reservar)
- Subscribe to my [Blog ↗](https://blog.pythonassembly.com/)
- Let's keep in touch on [LinkedIn ↗](www.linkedin.com/in/jsulopz) 😄
# Load the Data
> - Simply execute the following lines of code to load the data
> - This dataset contains **statistics** (columns)
> - About **Car Models** (rows)
```
import seaborn as sns
df = sns.load_dataset(name='mpg', index_col='name')
df.sample(10)
dfclean=df.dropna()
dfclean
X=dfclean[['mpg','horsepower']]
```
# Data `preprocessing`
> - Do you need to *transform* the data
> - To get a **truthful insight** of the model?
```
from sklearn.preprocessing import MinMaxScaler
scaler=MinMaxScaler()
import pandas as pd
vnorm = scaler.fit_transform(X)
X.columns
X.index
dfnorm = pd.DataFrame(vnorm, columns=X.columns, index=X.index)
dfnorm.head()
```
# `KMeans()` Model in Python
## Build the Model
> 1. **Necesity**: Build Model
> 2. **Google**: How do you search for the solution?
> 3. **Solution**: Find the `function()` that makes it happen
```
from sklearn.cluster import KMeans
error=[]
error
model=KMeans(n_clusters=1)
model.fit(X=dfnorm)
inercia=model.score(X=dfnorm)
inercia
error.append(inercia)
error
model=KMeans(n_clusters=2)
model.fit(X=dfnorm)
inercia = model.score(X=dfnorm)
inercia
error.append(inercia)
error
error = []
for pepa in [1, 2, 3, 4, 5, 6, 7, 8, 9, 10]:
model=KMeans(n_clusters=pepa)
model.fit(X=dfnorm)
inercia = model.score(X=dfnorm)
error.append(inercia)
error
sns.scatterplot(y=error, x=list(range(1, 11)))
sns.lineplot(y=error, x=list(range(1, 11)));
import matplotlib.pyplot as plt
dfgroup = dfsel.groupby('cluster')
dfgroup.get_group(0)
dfgroup.get_group(1)
plt.scatter(x='mpg', y='horsepower', data=dfgroup.get_group(0))
plt.scatter(x='mpg', y='horsepower', data=dfgroup.get_group(1))
plt.scatter(x='mpg', y='horsepower', data=dfgroup.get_group(2))
for pepa in [0, 1, 2]:
plt.scatter(x='mpg', y='horsepower', data=dfgroup.get_group(pepa))
error
df.mpg
df.mpg.hist();
df.mpg.sort_values()
df.mpg.quantile(q=[0.1, 0.9])
model=KMeans(n_clusters=3)
```
## Code Thinking
> Which function computes the Model?
> - `fit()`
>
> How could can you **import the function in Python**?
```
X
model.fit(X)
```
### Separate Variables for the Model
> Regarding their role:
> 1. **Target Variable `y`**
>
> - [ ] What would you like **to predict**?
>
> Total number of accients? Or Alcohol?
>
> 2. **Explanatory Variable `X`**
>
> - [ ] Which variable will you use **to explain** the target?
### Data Visualization to Analyize Patterns
> - Visualize the 2 variables with a `scatterplot()`
> - And decide *how many `clusters`* you'd like to calculate
```
sns.scatterplot(x='mpg', y='horsepower', data=dfnorm)
```
### Finally `fit()` the Model
## `predict()` the *cluster* for every row
> - `model.` + `↹`
```
pred=model.predict(X=dfnorm)
dfnorm
pred
```
> - Create a `dfsel` DataFrame
> - That contains the **columns you used for the model**
```
dfsel=dfnorm.copy()
```
> - Add a **new column**
> - That **contains the `cluster` prediction** for every USA State
```
dfsel['cluster'] = pred
pred
dfsel.sample(20)
```
## Model Visualization
> - You may `hue=` the points with the `cluster` column
```
dfsel.cluster
sns.scatterplot(x='mpg', y='horsepower', data=dfsel,hue=dfsel.cluster, palette='Set1');
```
## Model Interpretation
```
%%HTML
<iframe width="560" height="315" src="https://www.youtube.com/embed/4b5d3muPQmA" title="YouTube video player" frameborder="0" allow="accelerometer; autoplay; clipboard-write; encrypted-media; gyroscope; picture-in-picture" allowfullscreen></iframe>
```
> - Can you put a **`nickname` to each group**?
> - Observe the `centroids` within `model.` + `↹`
```
model.__dict__
dfcentroides = pd.DataFrame(model.cluster_centers_, columns=['mpg', 'horsepower'])
dfcentroides
```
## Model Visualization with Centroids
> - I want to see the `centroid`
> - with a **big `markers="X"`** in the plot
```
sns.scatterplot(x='mpg', y='horsepower', data=dfsel,hue=dfsel.cluster, palette='Set1');
sns.scatterplot(x='mpg', y='horsepower', data=dfcentroides,hue=dfcentroides.index, palette='Set1', s=500);
a = dfnorm.sample()
a
model.predict(a)
model.__dict__
```
# Achieved Goals
_Double click on **this cell** and place an `X` inside the square brackets (i.e., [X]) if you think you understand the goal:_
- [X] Understand how the **machine optimizes a model**
- No more than to find the best numbers for a mathematical equation
- [X] **Residual Sum of Squares (RSS)** as a fundamental measure for the **error**. We see it on ↓
- Neural Networks
- Linear Regression
- Variance
- [X] Understand the necessity to **Scale** the Data
- For all algorithms that involves **distance calculation**.
- [X] Understand that programming is not an end itself, but a tool to achieve the end
- We need to understand the problem and design the solution before coding
- But we won't need how to design the solution if we don't know how to code first
- Solution? Apply the discipline
- [X] There is **not a unique way to group data**. The same way it is not a unique way ↓
- To predict a number **Regression Mathematical Equations**
- To predict a category **Classification Mathematical Equations**
## Machine Learning
- Modelos Supervisados
- Objetivo: tipo numerica → Regresion
- Objetivo: tipo categorica → clasificacion
- Modelos No Supervisados
- No hay objetivo: queremos agrupar
| github_jupyter |
# Part 1: Extracting a Journal's Publications+Researchers Datasets
In this notebook we are going to
* extract all publications data for a given journal
* have a quick look at the publications' authors and affiliations
* review how many authors have been disambiguated with a Dimensions Researcher ID
* produce a dataset of non-disambiguated authors that can be used for manual disambiguation
## Prerequisites: Installing the Dimensions Library and Logging in
```
# @markdown # Get the API library and login
# @markdown Click the 'play' button on the left (or shift+enter) after entering your API credentials
username = "" #@param {type: "string"}
password = "" #@param {type: "string"}
endpoint = "https://app.dimensions.ai" #@param {type: "string"}
!pip install dimcli plotly tqdm -U --quiet
import dimcli
from dimcli.shortcuts import *
dimcli.login(username, password, endpoint)
dsl = dimcli.Dsl()
#
# load common libraries
import time
import sys
import json
import os
import pandas as pd
from pandas.io.json import json_normalize
from tqdm.notebook import tqdm as progress
#
# charts libs
# import plotly_express as px
import plotly.express as px
if not 'google.colab' in sys.modules:
# make js dependecies local / needed by html exports
from plotly.offline import init_notebook_mode
init_notebook_mode(connected=True)
#
# create output data folder
if not(os.path.exists("data")):
os.mkdir("data")
```
## Selecting a Journal and Extracting All Publications Metadata
```
#@title Select a journal from the dropdown
#@markdown If the journal isn't there, you can try type in the exact name instead.
journal_title = "Nature Genetics" #@param ['Nature', 'The Science of Nature', 'Nature Communications', 'Nature Biotechnology', 'Nature Medicine', 'Nature Genetics', 'Nature Neuroscience', 'Nature Structural & Molecular Biology', 'Nature Methods', 'Nature Cell Biology', 'Nature Immunology', 'Nature Reviews Drug Discovery', 'Nature Materials', 'Nature Physics', 'Nature Reviews Neuroscience', 'Nature Nanotechnology', 'Nature Reviews Genetics', 'Nature Reviews Urology', 'Nature Reviews Molecular Cell Biology', 'Nature Precedings', 'Nature Reviews Cancer', 'Nature Photonics', 'Nature Reviews Immunology', 'Nature Reviews Cardiology', 'Nature Reviews Gastroenterology & Hepatology', 'Nature Reviews Clinical Oncology', 'Nature Reviews Endocrinology', 'Nature Reviews Neurology', 'Nature Chemical Biology', 'Nature Reviews Microbiology', 'Nature Geoscience', 'Nature Reviews Rheumatology', 'Nature Climate Change', 'Nature Reviews Nephrology', 'Nature Chemistry', 'Nature Digest', 'Nature Protocols', 'Nature Middle East', 'Nature India', 'Nature China', 'Nature Plants', 'Nature Microbiology', 'Nature Ecology & Evolution', 'Nature Astronomy', 'Nature Energy', 'Nature Human Behaviour', 'AfCS-Nature Molecule Pages', 'Human Nature', 'Nature Reviews Disease Primers', 'Nature Biomedical Engineering', 'Nature Reports Stem Cells', 'Nature Reviews Materials', 'Nature Sustainability', 'Nature Catalysis', 'Nature Electronics', 'Nature Reviews Chemistry', 'Nature Metabolism', 'Nature Reviews Physics', 'Nature Machine Intelligence', 'NCI Nature Pathway Interaction Database', 'Nature Reports: Climate Change'] {allow-input: true}
start_year = 2015 #@param {type: "number"}
#@markdown ---
# PS
# To get titles from the API one can do this:
# > %dsldf search publications where journal.title~"Nature" and publisher="Springer Nature" return journal limit 100
# > ", ".join([f"'{x}'" for x in list(dsl_last_results.title)])
#
q_template = """search publications where
journal.title="{}" and
year>={}
return publications[basics+altmetric+times_cited]"""
q = q_template.format(journal_title, start_year)
print("DSL Query:\n----\n", q, "\n----")
pubs = dsl.query_iterative(q.format(journal_title, start_year), limit=500)
```
Save the data as a CSV file in case we want to reuse it later
```
dfpubs = pubs.as_dataframe()
dfpubs.to_csv("data/1.pubs_metadata_with_metrics.csv")
# preview the publications
dfpubs.head(10)
```
Extract the authors data
```
# preview the authors data
authors = pubs.as_dataframe_authors()
authors.to_csv("data/1.publications_authors.csv", index=False)
authors.head(10)
```
Extract the affiliations data
```
affiliations = pubs.as_dataframe_authors_affiliations()
affiliations.to_csv("data/1.publications_authors_affiliations.csv", index=False)
affiliations.head(10)
```
## Some stats about authors
* count how many authors in total
* count how many authors have a researcher ID
* count how many unique researchers IDs we have in total
```
researchers = authors.query("researcher_id!=''")
#
df = pd.DataFrame({
'measure' : ['Authors in total (non unique)', 'Authors with a researcher ID', 'Authors with a researcher ID (unique)'],
'count' : [len(authors), len(researchers), researchers['researcher_id'].nunique()],
})
px.bar(df, x="measure", y="count", title=f"Author stats for {journal_title} (from {start_year})")
# save the researchers data to a file
researchers.to_csv("data/1.authors_with_researchers_id.csv")
```
## Apprendix: A quick look at authors *without a Researcher ID*
We're not going to try to disambiguate them here, but still it's good to have a quick look at them...
Looks like the most common surname is `Wang`, while the most common first name is an empty value
```
authors_without_id = authors.query("researcher_id==''")
authors_without_id[['first_name', 'last_name']].describe()
```
Top Ten surnames seem all Chinese..
```
authors_without_id['last_name'].value_counts()[:10]
```
### Any common patterns?
If we try to group the data by name+surname we can see some interesting patterns
* some entries are things which are not persons (presumably the results of bad source data in Dimensions, eg from the publisher)
* there are some apparently meaningful name+surname combinations with a lot of hits
* not many Chinese names in the top ones
```
test = authors_without_id.groupby(["first_name", "last_name"]).size()
test.sort_values(ascending=False, inplace=True)
test.head(50)
```
## Conclusion and next steps
For the next tasks, we will focus on the disambiguated authors as the Researcher ID links will let us carry out useful analyses.
Still, we can **save the authors with missing IDs** results and try to do some manual disambiguation later. To this end, adding a simple google-search URL can help in making sense of these data quickly.
```
from urllib.parse import quote
out = []
for index, value in test.items():
# compose a simple URL of the form 'https://www.google.com/search?q=tonu+esko'
if index[0] or index[1]:
n, s = quote(index[0]), quote(index[1])
url = f"https://www.google.com/search?q={n}+{s}"
else:
url = ""
d = {'name': index[0] , 'surname' : index[1] , 'frequency' : value , 'search_url' : url }
out.append(d)
dftest = pd.DataFrame.from_dict(out)
# set order of columns
dftest = dftest[['name', 'surname', 'frequency', 'search_url']]
dftest.head(20)
# save the data
#
dftest.to_csv("data/1.authors_not_disambiguated_frequency.csv", header=True)
if COLAB_ENV:
files.download("data/1.authors_not_disambiguated_frequency.csv")
files.download("data/1.authors_with_researchers_id.csv")
files.download("data/1.publications_authors.csv")
files.download("data/1.publications_authors_affiliations.csv")
files.download("data/1.pubs_metadata_with_metrics.csv")
```
That's it!
Now let's go and open this in [Google Sheets](https://docs.google.com/spreadsheets/)...
| github_jupyter |
```
%pylab inline
import os
import sys
src_dir = os.path.join(os.getcwd(), os.pardir, 'src')
sys.path.append(src_dir)
import pickle
import numpy as np
import pandas as pd
# Set the seed for Keras
np.random.seed(4)
from model.model import get_model, split_train_test
from model.predictions import fit_model, predict, assemble_submission, convert_predictions, AMAPE
from utils.visualisations import PenguinVisualisation
# Use the same features without noise added computed for Submission 2
fname = '../data/interim/Submission_02/features.p'
df_train, df_test, df_features, _, features, scaler = pickle.load(open(fname, 'rb'))
print(df_train.shape, df_test.shape)
df_train.head()
df_train.columns
# Get the model
ts_steps = 4
aux_input_size = 6
model = get_model(ts_steps, aux_input_size=aux_input_size)
# Run the fitting
batch_size = 256
model, history = fit_model(df_train, model, df_test, batch_size=batch_size, epochs=300, verbose=0)
# Show the history of the validation loss and select the "best" number of epochs to run
val_loss = history.history['val_loss']
best_epoch_auto = np.argmin(val_loss) + 1
best_epoch = best_epoch_auto
print("Epoch with lowest validation loss is epoch %i with a loss of %.2f" %(best_epoch_auto, val_loss[best_epoch_auto]))
print("Manually chosen epoch is epoch %i with a loss of %.2f" %(best_epoch, val_loss[best_epoch]))
fig = plt.figure(figsize=(13,5))
ax = fig.add_subplot(111)
ax.plot(val_loss)
ax.axvline(best_epoch-1, color='green', linestyle='dashed');
ax.axvline(best_epoch_auto-1, color='red');
# Get the model
np.random.seed(4)
ts_steps = 4
aux_input_size = 6
model = get_model(ts_steps, aux_input_size=aux_input_size)
model, history = fit_model(df_train, model, df_test, batch_size=batch_size, epochs=best_epoch, verbose=0)
val_loss_re = history.history['val_loss']
best_epoch_re = np.argmin(val_loss_re) + 1
print("Epoch with lowest validation loss is epoch %i with a loss of %.2f" %(best_epoch_re, val_loss_re[best_epoch_re-1]))
fig = plt.figure(figsize=(13,5))
ax = fig.add_subplot(111)
ax.plot(val_loss_re)
ax.axvline(best_epoch_re-1, color='red');
# Predict the steps ahead
df_predictions = predict(df_features, 4, model, features)
df_predictions.head()
# Rescale the predictions
df_predictions = convert_predictions(df_predictions, scaler)
df_predictions = df_predictions.round()
df_predictions.head()
# Show some estimate of what the final score for the test set would be.
# First, show the score for all data (i.e. including the interpolated counts)
amape = AMAPE(interpolated=True)
amapeScore = amape.amape(df_predictions)
print("AMAPE 2011: %.2f" %amapeScore.loc['2011'])
print("AMAPE 2012: %.2f" %amapeScore.loc['2012'])
print("AMAPE 2013: %.2f" %amapeScore.loc['2013'])
print("Projected AMAPE score: %.2f" %(amapeScore.loc['2011']+amapeScore.loc['2012']+amapeScore.loc['2013']))
plt.figure(figsize=(10,5))
plt.plot(amapeScore);
# Show the score only for the truly observed predictions. This will be harder to predict since the interpolated
# data is smooth and thus much easier to predict.
amape = AMAPE(interpolated=False)
amapeScore = amape.amape(df_predictions)
print("AMAPE 2011: %.2f" %amapeScore.loc['2011'])
print("AMAPE 2012: %.2f" %amapeScore.loc['2012'])
print("AMAPE 2013: %.2f" %amapeScore.loc['2013'])
print("Projected AMAPE score: %.2f" %(amapeScore.loc['2011']+amapeScore.loc['2012']+amapeScore.loc['2013']))
plt.figure(figsize=(10,5))
plt.plot(amapeScore);
# Have look at some random sites and species and check if the model seems to make sense
vis = PenguinVisualisation(df_predictions)
vis.plot_random()
# Finally, retrain the model with the full data and make the predictions for the submission
np.random.seed(4)
model = get_model(ts_steps, aux_input_size=aux_input_size)
model, _ = fit_model(df_features, model, None, batch_size=batch_size, epochs=best_epoch, verbose=0)
# Make the predictions
df_predictions = predict(df_features, 4, model, features)
df_predictions = convert_predictions(df_predictions, scaler)
df_predictions = df_predictions.round()
# Prepare the submission
df_submission = assemble_submission(df_predictions)
df_submission.head()
# Check if the score changes much compared to the run without the test data.
amape = AMAPE(interpolated=False)
amapeScore = amape.amape(df_predictions)
print("AMAPE 2011: %.2f" %amapeScore.loc['2011'])
print("AMAPE 2012: %.2f" %amapeScore.loc['2012'])
print("AMAPE 2013: %.2f" %amapeScore.loc['2013'])
print("Projected AMAPE score: %.2f" %(amapeScore.loc['2011']+amapeScore.loc['2012']+amapeScore.loc['2013']))
plt.figure(figsize=(10,5))
plt.plot(amapeScore);
fname_submission = '../data/submission/submission_03.csv'
df_submission.to_csv(fname_submission, index=True)
```
| github_jupyter |
# Hill Climbing
---
In this notebook, we will train hill climbing with adaptive noise scaling with OpenAI Gym's Cartpole environment.
### 1. Import the Necessary Packages
```
import gym
import numpy as np
from collections import deque
import matplotlib.pyplot as plt
%matplotlib inline
!python -m pip install pyvirtualdisplay
from pyvirtualdisplay import Display
display = Display(visible=0, size=(1400, 900))
display.start()
is_ipython = 'inline' in plt.get_backend()
if is_ipython:
from IPython import display
plt.ion()
```
### 2. Define the Policy
```
env = gym.make('CartPole-v0')
print('observation space:', env.observation_space)
print('action space:', env.action_space)
class Policy():
def __init__(self, s_size=4, a_size=2):
self.w = 1e-4*np.random.rand(s_size, a_size) # weights for simple linear policy: state_space x action_space
def forward(self, state):
x = np.dot(state, self.w)
return np.exp(x)/sum(np.exp(x))
def act(self, state):
probs = self.forward(state)
#action = np.random.choice(2, p=probs) # option 1: stochastic policy
action = np.argmax(probs) # option 2: deterministic policy
return action
```
### 3. Train the Agent with Stochastic Policy Search
```
env = gym.make('CartPole-v0')
env.seed(0)
np.random.seed(0)
policy = Policy()
def hill_climbing(n_episodes=1000, max_t=1000, gamma=1.0, print_every=100, noise_scale=1e-2):
"""Implementation of hill climbing with adaptive noise scaling.
Params
======
n_episodes (int): maximum number of training episodes
max_t (int): maximum number of timesteps per episode
gamma (float): discount rate
print_every (int): how often to print average score (over last 100 episodes)
noise_scale (float): standard deviation of additive noise
"""
scores_deque = deque(maxlen=100)
scores = []
best_R = -np.Inf
best_w = policy.w
for i_episode in range(1, n_episodes+1):
rewards = []
state = env.reset()
for t in range(max_t):
action = policy.act(state)
state, reward, done, _ = env.step(action)
rewards.append(reward)
if done:
break
scores_deque.append(sum(rewards))
scores.append(sum(rewards))
discounts = [gamma**i for i in range(len(rewards)+1)]
R = sum([a*b for a,b in zip(discounts, rewards)])
if R >= best_R: # found better weights
best_R = R
best_w = policy.w
noise_scale = max(1e-3, noise_scale / 2)
policy.w += noise_scale * np.random.rand(*policy.w.shape)
else: # did not find better weights
noise_scale = min(2, noise_scale * 2)
policy.w = best_w + noise_scale * np.random.rand(*policy.w.shape)
if i_episode % print_every == 0:
print('Episode {}\tAverage Score: {:.2f}'.format(i_episode, np.mean(scores_deque)))
if np.mean(scores_deque)>=195.0:
print('Environment solved in {:d} episodes!\tAverage Score: {:.2f}'.format(i_episode-100, np.mean(scores_deque)))
policy.w = best_w
break
return scores
scores = hill_climbing()
```
### 4. Plot the Scores
```
fig = plt.figure()
ax = fig.add_subplot(111)
plt.plot(np.arange(1, len(scores)+1), scores)
plt.ylabel('Score')
plt.xlabel('Episode #')
plt.show()
```
### 5. Watch a Smart Agent!
```
env = gym.make('CartPole-v0')
state = env.reset()
img = plt.imshow(env.render(mode='rgb_array'))
for t in range(200):
action = policy.act(state)
img.set_data(env.render(mode='rgb_array'))
plt.axis('off')
display.display(plt.gcf())
display.clear_output(wait=True)
state, reward, done, _ = env.step(action)
if done:
break
env.close()
```
| github_jupyter |
# Challenge
Another approach to identifying fraudulent transactions is to look for outliers in the data. Standard deviation or quartiles are often used to detect outliers. Using this starter notebook, code two Python functions:
* One that uses standard deviation to identify anomalies for any cardholder.
* Another that uses interquartile range to identify anomalies for any cardholder.
## Identifying Outliers using Standard Deviation
```
# Initial imports
import pandas as pd
import numpy as np
import random
from sqlalchemy import create_engine
# Create a connection to the database
engine = create_engine("postgresql://postgres:postgres@localhost:5432/fraud_detection")
# Write function that locates outliers using standard deviation
def find_outliers_sd(card_holder=1):
query = (
"SELECT t.date, t.amount, t.card "
+ "FROM transaction AS t "
+ "JOIN credit_card AS cc ON cc.card = t.card "
+ "JOIN card_holder AS ch ON ch.id = cc.id_card_holder "
+ "WHERE ch.id = "
+ str(card_holder)
+ " ORDER BY date"
)
data = pd.read_sql(query, engine)
elements = data["amount"]
mean = np.mean(elements, axis=0)
sd = np.std(elements, axis=0)
# 2 standard deviations are taken for analysis purposes
low_transactions = [x for x in elements if (x < mean - 2 * sd)]
high_transaction = [x for x in elements if (x > mean + 2 * sd)]
final_list = low_transactions + high_transaction
if len(final_list) > 0:
query = (
"SELECT t.date, t.amount, t.card "
+ "FROM transaction AS t "
+ "JOIN credit_card AS cc ON cc.card = t.card "
+ "JOIN card_holder AS ch ON ch.id = cc.id_card_holder "
+ "WHERE ch.id = "
+ str(card_holder)
+ " AND t.amount IN ("
+ str(final_list)[1:-1]
+ ") "
+ "ORDER BY date"
)
data = pd.read_sql(query, engine)
return data
else:
return "There are no fraudulent transactions identified for this card holder"
# Find anomalous transactions for 3 random card holders
for i in range(1, 4):
card_holder = random.randint(1, 25)
print("*" * 60)
print(f"Looking for fraudulent transactions for card holder id {card_holder}")
print(find_outliers_sd(card_holder))
```
## Identifying Outliers Using Interquartile Range
```
# Write a function that locates outliers using interquartile range
def find_outliers_iqr(card_holder=1):
query = (
"SELECT t.date, t.amount, t.card "
+ "FROM transaction AS t "
+ "JOIN credit_card AS cc ON cc.card = t.card "
+ "JOIN card_holder AS ch ON ch.id = cc.id_card_holder "
+ "WHERE ch.id = "
+ str(card_holder)
+ " ORDER BY date"
)
data = pd.read_sql(query, engine)
# calculate interquartile range
q25, q75 = np.percentile(data["amount"], 25), np.percentile(data["amount"], 75)
iqr = q75 - q25
# calculate the outlier cutoff
cut_off = iqr * 1.5
lower, upper = q25 - cut_off, q75 + cut_off
# identify outliers
outliers = [x for x in data["amount"] if x < lower or x > upper]
if len(outliers) > 0:
query = (
"SELECT t.date, t.amount, t.card "
+ "FROM transaction AS t "
+ "JOIN credit_card AS cc ON cc.card = t.card "
+ "JOIN card_holder AS ch ON ch.id = cc.id_card_holder "
+ "WHERE ch.id = "
+ str(card_holder)
+ " AND t.amount IN ("
+ str(outliers)[1:-1]
+ ") "
+ "ORDER BY date"
)
data = pd.read_sql(query, engine)
return data
else:
return "There are no fraudulent transactions identified for this card holder"
# Find anomalous transactions for 3 random card holders
for i in range(1, 4):
card_holder = random.randint(1, 25)
print("*" * 60)
print(f"Looking for fraudulent transactions for card holder id {card_holder}")
print(find_outliers_iqr(card_holder))
```
| github_jupyter |
# Using the PyTorch JIT Compiler with Pyro
This tutorial shows how to use the PyTorch [jit compiler](https://pytorch.org/docs/master/jit.html) in Pyro models.
#### Summary:
- You can use compiled functions in Pyro models.
- You cannot use pyro primitives inside compiled functions.
- If your model has static structure, you can use a `Jit*` version of an `ELBO` algorithm, e.g.
```diff
- Trace_ELBO()
+ JitTrace_ELBO()
```
- The [HMC](http://docs.pyro.ai/en/dev/mcmc.html#pyro.infer.mcmc.HMC) and [NUTS](http://docs.pyro.ai/en/dev/mcmc.html#pyro.infer.mcmc.NUTS) classes accept `jit_compile=True` kwarg.
- Models should input all tensors as `*args` and all non-tensors as `**kwargs`.
- Each different value of `**kwargs` triggers a separate compilation.
- Use `**kwargs` to specify all variation in structure (e.g. time series length).
- To ignore jit warnings in safe code blocks, use `with pyro.util.ignore_jit_warnings():`.
- To ignore all jit warnings in `HMC` or `NUTS`, pass `ignore_jit_warnings=True`.
#### Table of contents
- [Introduction](#Introduction)
- [A simple model](#A-simple-model)
- [Varying structure](#Varying-structure)
```
import os
import torch
import pyro
import pyro.distributions as dist
from torch.distributions import constraints
from pyro import poutine
from pyro.distributions.util import broadcast_shape
from pyro.infer import Trace_ELBO, JitTrace_ELBO, TraceEnum_ELBO, JitTraceEnum_ELBO, SVI
from pyro.infer.mcmc import MCMC, NUTS
from pyro.infer.autoguide import AutoDiagonalNormal
from pyro.optim import Adam
smoke_test = ('CI' in os.environ)
assert pyro.__version__.startswith('1.3.1')
pyro.enable_validation(True) # <---- This is always a good idea!
```
## Introduction
PyTorch 1.0 includes a [jit compiler](https://pytorch.org/docs/master/jit.html) to speed up models. You can think of compilation as a "static mode", whereas PyTorch usually operates in "eager mode".
Pyro supports the jit compiler in two ways. First you can use compiled functions inside Pyro models (but those functions cannot contain Pyro primitives). Second, you can use Pyro's jit inference algorithms to compile entire inference steps; in static models this can reduce the Python overhead of Pyro models and speed up inference.
The rest of this tutorial focuses on Pyro's jitted inference algorithms: [JitTrace_ELBO](http://docs.pyro.ai/en/dev/inference_algos.html#pyro.infer.trace_elbo.JitTrace_ELBO), [JitTraceGraph_ELBO](http://docs.pyro.ai/en/dev/inference_algos.html#pyro.infer.tracegraph_elbo.JitTraceGraph_ELBO), [JitTraceEnum_ELBO](http://docs.pyro.ai/en/dev/inference_algos.html#pyro.infer.traceenum_elbo.JitTraceEnum_ELBO), [JitMeanField_ELBO](http://docs.pyro.ai/en/dev/inference_algos.html#pyro.infer.trace_mean_field_elbo.JitTraceMeanField_ELBO), [HMC(jit_compile=True)](http://docs.pyro.ai/en/dev/mcmc.html#pyro.infer.mcmc.HMC), and [NUTS(jit_compile=True)](http://docs.pyro.ai/en/dev/mcmc.html#pyro.infer.mcmc.NUTS). For further reading, see the [examples/](https://github.com/pyro-ppl/pyro/tree/dev/examples) directory, where most examples include a `--jit` option to run in compiled mode.
## A simple model
Let's start with a simple Gaussian model and an [autoguide](http://docs.pyro.ai/en/dev/infer.autoguide.html).
```
def model(data):
loc = pyro.sample("loc", dist.Normal(0., 10.))
scale = pyro.sample("scale", dist.LogNormal(0., 3.))
with pyro.plate("data", data.size(0)):
pyro.sample("obs", dist.Normal(loc, scale), obs=data)
guide = AutoDiagonalNormal(model)
data = dist.Normal(0.5, 2.).sample((100,))
```
First let's run as usual with an SVI object and `Trace_ELBO`.
```
%%time
pyro.clear_param_store()
elbo = Trace_ELBO()
svi = SVI(model, guide, Adam({'lr': 0.01}), elbo)
for i in range(2 if smoke_test else 1000):
svi.step(data)
```
Next to run with a jit compiled inference, we simply replace
```diff
- elbo = Trace_ELBO()
+ elbo = JitTrace_ELBO()
```
Also note that the `AutoDiagonalNormal` guide behaves a little differently on its first invocation (it runs the model to produce a prototype trace), and we don't want to record this warmup behavior when compiling. Thus we call the `guide(data)` once to initialize, then run the compiled SVI,
```
%%time
pyro.clear_param_store()
guide(data) # Do any lazy initialization before compiling.
elbo = JitTrace_ELBO()
svi = SVI(model, guide, Adam({'lr': 0.01}), elbo)
for i in range(2 if smoke_test else 1000):
svi.step(data)
```
Notice that we have a more than 2x speedup for this small model.
Let us now use the same model, but we will instead use MCMC to generate samples from the model's posterior. We will use the No-U-Turn(NUTS) sampler.
```
%%time
nuts_kernel = NUTS(model)
pyro.set_rng_seed(1)
mcmc_run = MCMC(nuts_kernel, num_samples=100).run(data)
```
We can compile the potential energy computation in NUTS using the `jit_compile=True` argument to the NUTS kernel. We also silence JIT warnings due to the presence of tensor constants in the model by using `ignore_jit_warnings=True`.
```
%%time
nuts_kernel = NUTS(model, jit_compile=True, ignore_jit_warnings=True)
pyro.set_rng_seed(1)
mcmc_run = MCMC(nuts_kernel, num_samples=100).run(data)
```
We notice a significant increase in sampling throughput when JIT compilation is enabled.
## Varying structure
Time series models often run on datasets of multiple time series with different lengths. To accomodate varying structure like this, Pyro requires models to separate all model inputs into tensors and non-tensors.$^\dagger$
- Non-tensor inputs should be passed as `**kwargs` to the model and guide. These can determine model structure, so that a model is compiled for each value of the passed `**kwargs`.
- Tensor inputs should be passed as `*args`. These must not determine model structure. However `len(args)` may determine model structure (as is used e.g. in semisupervised models).
To illustrate this with a time series model, we will pass in a sequence of observations as a tensor `arg` and the sequence length as a non-tensor `kwarg`:
```
def model(sequence, num_sequences, length, state_dim=16):
# This is a Gaussian HMM model.
with pyro.plate("states", state_dim):
trans = pyro.sample("trans", dist.Dirichlet(0.5 * torch.ones(state_dim)))
emit_loc = pyro.sample("emit_loc", dist.Normal(0., 10.))
emit_scale = pyro.sample("emit_scale", dist.LogNormal(0., 3.))
# We're doing manual data subsampling, so we need to scale to actual data size.
with poutine.scale(scale=num_sequences):
# We'll use enumeration inference over the hidden x.
x = 0
for t in pyro.markov(range(length)):
x = pyro.sample("x_{}".format(t), dist.Categorical(trans[x]),
infer={"enumerate": "parallel"})
pyro.sample("y_{}".format(t), dist.Normal(emit_loc[x], emit_scale),
obs=sequence[t])
guide = AutoDiagonalNormal(poutine.block(model, expose=["trans", "emit_scale", "emit_loc"]))
# This is fake data of different lengths.
lengths = [24] * 50 + [48] * 20 + [72] * 5
sequences = [torch.randn(length) for length in lengths]
```
Now lets' run SVI as usual.
```
%%time
pyro.clear_param_store()
elbo = TraceEnum_ELBO(max_plate_nesting=1)
svi = SVI(model, guide, Adam({'lr': 0.01}), elbo)
for i in range(1 if smoke_test else 10):
for sequence in sequences:
svi.step(sequence, # tensor args
num_sequences=len(sequences), length=len(sequence)) # non-tensor args
```
Again we'll simply swap in a `Jit*` implementation
```diff
- elbo = TraceEnum_ELBO(max_plate_nesting=1)
+ elbo = JitTraceEnum_ELBO(max_plate_nesting=1)
```
Note that we are manually specifying the `max_plate_nesting` arg. Usually Pyro can figure this out automatically by running the model once on the first invocation; however to avoid this extra work when we run the compiler on the first step, we pass this in manually.
```
%%time
pyro.clear_param_store()
# Do any lazy initialization before compiling.
guide(sequences[0], num_sequences=len(sequences), length=len(sequences[0]))
elbo = JitTraceEnum_ELBO(max_plate_nesting=1)
svi = SVI(model, guide, Adam({'lr': 0.01}), elbo)
for i in range(1 if smoke_test else 10):
for sequence in sequences:
svi.step(sequence, # tensor args
num_sequences=len(sequences), length=len(sequence)) # non-tensor args
```
Again we see more than 2x speedup. Note that since there were three different sequence lengths, compilation was triggered three times.
$^\dagger$ Note this section is only valid for SVI, and HMC/NUTS assume fixed model arguments.
| github_jupyter |
Your name here.
Your section number here.
# Workshop 1: Python basics, and a little plotting
**Submit this notebook to bCourses to receive a grade for this Workshop.**
Please complete workshop activities in code cells in this iPython notebook. The activities titled **Practice** are purely for you to explore Python, and no particular output is expected. Some of them have some code written, and you should try to modify it in different ways to understand how it works. Although no particular output is expected at submission time, it is _highly_ recommended that you read and work through the practice activities before or alongside the exercises. However, the activities titled **Exercise** have specific tasks and specific outputs expected. Include comments in your code when necessary. Enter your name in the cell at the top of the notebook. The workshop should be submitted on bCourses under the Assignments tab.
To submit the assignment, click File->Download As->Notebook (.ipynb). Then upload the completed (.ipynb) file to the corresponding bCourses assignment.
## Practice: Writing Python code
### The iPython Interpreter
Time to write your first python code! In Jupyter, the code is written in "Cells". Click on the "+" button above to create a new cell and type in "2+2" (without the quotes ... or with them!) and see what happens! To execute, click "Run" button or press "Shift-Enter". Also try switching the type of the cell from "Code" to "Markdown" and see what happens
```
2+2
```
## Practice: Peforming arithmetic in Python
If you get bored of using WolframAlpha to help with physics homework, Python can also be used as a "glorified calculator". Python code must follow certain syntax in order to run properly--it tends to be a bit more picky than Wolfram. However, once you get used to the Python language, you have the freedom to calculate pretty much anything you can think of.
To start, let's see how to perform the basic arithmetic operations. The syntax is
<h3><center><i>number</i> operator <i>number</i></center></h3>
Run the cells below and take a look at several of the different operators that you can use in Python (text after "#" are non-executable comments).
```
3+2 #addition
3-2 #subtraction
3*2 #multiplication
3/2 #division
3%2 #modulus (remainder after division) see https://en.wikipedia.org/wiki/Modulo_operation
3**2 #exponentiation, note: 3^2 means something different in Python
```
Python cares __*a lot*__ about the spaces, tabs, and enters you type (this is known as whitespace in programming). Many of your errors this semester will involve improper indentation. However, in this case, you are free to put a lot of space between numbers and operators as long as you keep everything in one line.
```
5 * 3 #This is valid code
```
You are not limited to just 2 numbers and a single operator; you can put a whole bunch of operations on one line.
```
5 * 4 + 3 / 2
```
Python follows the standard order of operations (PEMDAS) : Parentheses -> Exponentiation -> Multiplication/Division -> Addition/Subtraction. If you use parentheses, make sure every ```(``` has a corresponding ```)```
```
5 * (4 + 3) / 2
```
## Practice: Strings vs numbers
If you're familiar with programming in other languages, you are probably aware that different [_types_](https://realpython.com/python-data-types/) of things exist--you can do more than work with numbers (and not all numbers are the same type). If you'd like to work with letters, words, or sentences in Python, then you'll be using something called a string. To input a string, simply put single `' '` or double `" "` quotes around your desired phrase.
```
"Hello world"
```
Some (but not all) of the arithmetic operations also work with strings; you can add two of them together.
```
"Phys" + "ics"
```
You can multiply a string by a number.
```
"ha"*3
```
This one doesn't work; try reading the error message and see if you understand what it's saying (this is a useful skill to develop).
```
"error"/3
```
## Practice: Printing
Up until this point, we've just been typing a single line of code in each Jupyter cell and running it. Most Python interpreters will display the result of the final thing you typed, but occassionally you want to display the results of many things in a single Python script.
```
"These are some numbers:"
3*2
3*3
3*4
```
In the cell above, there are several multiplications happening but only the final result is displayed. To display everything, we simply use a "print statement" on each line.
```
print("These are some numbers:")
print(3*2)
print(3*3)
print(3*4)
```
If you'd like to print multiple things on one line, you can separate them by commas within the print statement.
```
print("These are some numbers:", 3*2, 3*3, 3*4)
```
## Exercise 1: Four Fours
[Inspired by Harvey Mudd College's CS5 course] Here's an arithmetic game to try your hand at. Your task is to compute each of the numbers, from 1 through 11, using exactly four 4's and simple math operations. You're allowed to use `+` (addition), `-` (subtraction), `*` (multiplication), `/` (division), `sqrt()` (square root), `factorial()` (factorial), and `%` (modulus). You're also allowed to use `.4` (that's one 4) or `44` (that's two 4's) if you'd like. Just remember, you must use exactly four 4 digits total!
As a reminder, four factorial (denoted by $!$ in mathematics) is $4! = 4 \cdot 3 \cdot 2 \cdot 1$, and the modulus operator (usually denoted by $\text{mod}$ in mathematics) is the remainder after division. For instance, $\ 5\ \text{mod}\ 2 = 1$, $\ 13\ \text{mod}\ 7 = 6$, and $\ 14\ \text{mod}\ 7 = 0$.
We've given you `zero` for free, as `4 - 4 + 4 - 4`. Of course, we could have also done `44 * (.4 - .4)` or `factorial(4) - 4 * (4 + sqrt(4))`, since both of those also yield `0` (or rather, `0.0`. Why is that?) and use exactly four 4's.
```
### Exercise 1
from math import factorial, sqrt
print('Zero:', 4 - 4 + 4 - 4)
print('One:')
print('Two:')
print('Three:')
print('Four:')
print('Five:')
print('Six:')
print('Seven:')
print('Eight:')
print('Nine:')
print('Ten:')
print('Eleven:')
```
Your final source code will be full of four fours formulas, but your final output should look like this:
Zero: 0
One: 1
Two: 2
Three: 3
Four: 4
Five: 5
Six: 6
Seven: 7
Eight: 8
Nine: 9
Ten: 10
Eleven: 11
It's ok if some of these have a trailing `.0` (`0.0`, for instance), but make sure you understand why they do!
## Practice: Variables, functions, namespaces
### Variables
Suppose you calculate something in Python and would like to use the result later in your program (instead of just printing it and immediately throwing it away). One big difference between a calculator and a computer language is an ability to store the values in memory, give that memory block a name, and use the value in later calculations. Such named memory block is called a _variable_. To create a variable, use an _assignment_ opperator = . Once you have created the variable, you can use it in the calculations.
```
x = "Phys"
y = "ics!"
z = x + y # Put 'em together
z # See what we got!
y + x # Backwards!
len(z) # 8 characters in total ...
len(z)**2 # Computing the area?
z[0] # Grab the first character
z[1:3] # Grab the next two characters
z[:4]
z[:4] == x # Test a match!
z[4:] == y
z[:] # The whole string
z[::-1] # The whole string, right to left
z[1::3] # Start at the second character and take every third character from there
z*3 + 5*z[-1] # Woo!
```
### Namespaces
This notebook and interpreter are a great place to test things out and mess around. Some interpreters (like Canopy) comes preloaded with a couple libraries (like numpy and matplotlib) that we will use a lot in this course. In Jupyter, you have to pre-load each package before using it. This is a good python practice anyway ! Here is an example.
```
log(e)
```
Both the function `log` and the number `e` are from the `numpy` library, which needs to be loaded into Jupyter. "pylab" adds `matplotlib` (the standard plotting tool) to `numpy`, so we will use that.
```
from pylab import *
log(e)
```
Or type `pie([1,2,3])`, since `pie` is defined by matplotlib!
```
pie([1,2,3])
matplotlib.pyplot.show() #This line is needed so matplotlib actually displays the plot
```
Note that we imported all library definitions from `pylab` into the default <i>namespace</i>, and can use the functions directly instead of having to add the name or alias of the package:
```
import numpy as np
np.log(np.e)
```
Loading into the default namespace can be convenient, but also confusing since many names and variables are already used in ways you might not expect. When writing scripts you'll have to manually import any library you want to use. This little inconvenience is greatly worth the confusion it can save.
### Functions (looking a bit ahead)
You'll often find yourself performing the same operations on several different variables. For example, we might want to convert heights from feet to meters.
```
burj_khalifa = 2717 #height in feet
shanghai_tower = 2073 #height in feet
print(burj_khalifa / 3.281) #height in meters
print(shanghai_tower / 3.281) #height in meters
```
You could just type the same thing over and over (or copy and paste), but this becomes tedious as your operations become more complex. To simplify things, you can define a function in Python (above, you were able to use the `log()` function from the `numpy` library).
```
'''A function definition starts with the 'def' keyword,
followed by the function name. The input variables are then
placed in parentheses after the function name. The first line
ends with a colon'''
def feet_to_meters(height):
#The operations to be performed by the function are now written out at the first indentation level
#You can indent with tabs or a constant number of spaces; just be consistent
converted_height = height / 3.281
print("Your height is being converted to meters.")
return converted_height #To return a value from a function, use the 'return' keyword
```
To use a function, simply type its name with the appropriate input variables in parentheses.
```
feet_to_meters(burj_khalifa)
```
If you'd like a function with multiple input variables, simply separate them with commas in the function declaration.
```
def difference_in_meters(height1, height2):
converted_height1 = height1 / 3.281
converted_height2 = height2 / 3.281
return converted_height1 - converted_height2
difference_in_meters(burj_khalifa, shanghai_tower)
```
## Practice: Formatted output
Usually the data you manipulate has finate precision. You do not know it absolutely precisely, and therefore you should not report it with an arbitrary number of digits. One of the cardinal rules of a good science paper: round off all your numbers to the precision you know them (or care about) -- and no more !
#### Examples:
```
x = 20.0 # I only know 3 digits
print(x) # OK, let Python handle it
```
That's actually pretty good -- Python remembered stored precision !
What happens if you now use x in a calculation ?
```
print(sqrt(x))
```
Do we really know the output to 10 significant digits ? No ! So let's truncate it
```
print('sqrt(x) = {0:5.3f}'.format(sqrt(x)))
```
There are several formatting options available to you, but the basic idea is this:
place `{:.#f}` wherever you'd like to insert a variable into your string (where `#` is
the number of digits you'd like after the decimal point). Then type `.format()` after
the string and place the variable names within the parentheses.
```
from math import e
print("Euler's number with 5 decimal places is {:.5f} and with 3 decimal places is {:.3f}".format(e,e))
```
For more formatting options, see https://pyformat.info/
### Practice
Using what you just learned, try writing program to print only 4 decimal places of $\pi$ and $\log\pi$. The result should look like:
Hello world! Have some pie! 3.1416
And some pie from a log! 1.1447
```
from math import pi
#Your print statement here
```
## Exercise 2: Coulomb force
Write a function that calculates the magnitude of the force between two charged particles. The function should take the charge of each particle ($q_1$ and $q_2$) and the distance between them, $r$, as input (three input variables total). The electrostatic force between two particles is given by:
$ F = k\frac{q_1 q_2}{r^2}$
```
k = 8.99e9 #Coulomb constant, units: N * m**2 / C**2
def calculate_force(q1, q2, r):
#calculate (and return) the force between the two particles
```
Now call the function with random input values (of your choosing) and print the result with 3 decimal places. What happens if you call the function with the value $r=0$ ?
## Practice: Simple plotting
In order to do some plotting, we'll need the tools from two commonly used Python libraries: `matplotlib` (similar to Matlab plotting) and `numpy` (NUMerical PYthon). You've seen importing at work before with `from math import sqrt`; we can also import an entire library (or a large part of it) with the following syntax:
```
import numpy as np
import matplotlib.pyplot as plt
```
You could have also typed `import numpy`, but programmers are lazy when it comes to typing. By including `as np`, you now only have to type the two-letter word `np` when you'd like to use functions from the library. The `np` and `plt` part of the import statements can be whatever you like--these are just the standard names.
Numpy has a lot of the same functions as the `math` library; for example we have `sqrt`, `log`, and `exp`:
```
np.sqrt(4)
np.log(4)
np.exp(3)
np.log(np.exp(5))
```
We could have just gotten these functions from the `math` library, so why bother with `numpy`? There's another variable type in Python known as a *__list__*, which is exactly like it sounds--just a list of some things (numbers, strings, more lists, etc.). We'll talk about these more at some point, but the important thing is that `numpy` has a way better alternative: the `numpy` array. Usually anything you'd want to do with a list can also be done with a `numpy` array, but faster.
Let's just demonstrate by example. Suppose we want to plot the function `x**2`. To do this, we'll plot a collection of (x,y) points and connect them with lines. If the points are spaced closely enough, the plot will look nice and smooth on the screen.
```
x_values = np.linspace(-5, 5, 11)
print(x_values)
```
The `linspace` function from `numpy` gave us an array of 11 numbers, evenly spaced between -5 and 5. We'll want our points a bit closer, so let's change 11 to something larger.
```
x_values = np.linspace(-5, 5 , 1000)
y_values = x_values**2
```
To get the corresponding y values, we can just perform operations on the entire array of x values. Now, we can plot these using the `matplotlib` library.
```
plt.plot(x_values, y_values)
```
There's a ton of stuff you can do with `matplotlib.pyplot` or the `matplotlib` library as a whole, but here are a few basics to get you started.
```
plt.plot(x_values, x_values**3) #As before, this plots the (x,y) points and connects them with lines
plt.show() #This forces matplotlib to display the current figure
plt.figure() #This creates a new, empty figure
plt.plot(x_values, np.exp(x_values), 'g--') #There are lots of optional arguments that do cool things
plt.title(r'$e^x$') #Creates a title; you can use LaTeX formatting in matplotlib as shown here
plt.xlabel('y values') #Label for x-axis
plt.ylabel('exp(x)') #Label for y-axis
plt.show()
```
## Exercise 3: Plotting Radioactivity Data
[Adapted from Ayars, Problem 0-2]
The file Ba137.txt contains two columns. The first is counts from a Geiger counter, the second is time in seconds. If you opened this Workshop notebook using the Interact Link (from the bCourses page), then you should already have Ba137.txt in your datahub directory.
If not, it's available [here](https://raw.githubusercontent.com/celegante/code_chapter_0-_github/master/Ba137.txt). Open the link, right-click and save as a .txt file. Then upload to datahub.berkeley.edu or move it to whichever folder you're keeping this notebook.
1. Make a useful graph of this data, with axes labels and a title.
2. If this data follows an exponential curve, then plotting the natural log of the data (or plotting the raw data on a logarithmic scale) will result in a straight line. Determine whether this is the case, and explain your conclusion with---you guessed it---an appropriate graph.
Be sure to add comments throughout your code so it's clear what each section of the code is doing! It may help to refer to the lecture notes or Ayars Chapter 0.
Try using `'x'` or `'^'` as the marker type in your `plt.plot()` functions (instead of `'g-'`, for instance), to get a single x or triangle for each data point instead of a connected line. Google if you'd like to learn more options!
Once you're through, your code should produce two graphs, one with the data, another with the natural log of the data, both labelled appropriately. It should also print out a clear answer to the question in part 2 (e.g., `Yes, the data follows an exponential curve`, or `No, the data does not follow an exponential curve`).
```
### Exercise 3
import numpy as np
import matplotlib.pyplot as plt
### Load the data here
counts, times = np.loadtxt('Ba137.txt', unpack = True)
plt.figure() # Start a clean figure for your first plot
### Your code for the first plot here!
plt.figure() # Start a clean figure for your second plot
### Your code for the second plot here!
plt.show() # This tells python to display the plots you've made
```
#### Hints
Put the file in the same directory as your python file, and use numpy's `loadtxt` or `genfromtxt` function to load each column into an array for use in your plots.
If your file isn't loading correctly, it might be because your IPython working directory isn't the same as the directory your script and Ba137.txt are in.
If you'd like to learn more about what `loadtxt` does (or why the `unpack = True` option is important), type `loadtxt?` or `help(loadtxt)` into the python interpreter for documentation. Press `q` to get out of the documentation.
## Practice: Debugging
[Adapted from Langtangen, Exercise 1.16] Working with a partner, type these statements into your python interpreter. Figure out why some statements fail and correct the errors.
*Hint: Try testing the left- and right-hand sides seperately before you put them together in statements. It's ok if you don't understand yet why some of the expressions evaluate to the results they do (like the last one).*
1a = 2
a1 = b
x = 2
y = X + 4 # is it 6?
5 = 5 # is it True?
4/5 == 4.0/5.0 # is it True? (this depends on which version of Python you're using)
type(10/2) == type(10/2.) # is it True? (again, this depends on the Python version)
from Math import factorial
print factorial(pi)
discount = 12%
## You're done!
Congratulations, you've finished this week's workshop! You're welcome to leave early or get started on this week's homework.
| github_jupyter |
# 4.3.2 有価証券報告書内のMD&Aが記載されているhtmファイルを抽出
Zipファイルで送られてきた大量の有価証券書のデータをPythonにより自動的に解凍した。その後、有価証券報告書内のMD&A情報のみが記載されたhtmファイルのみを抽出をした。
```
import pandas as pd
import glob
import os
import zipfile
from datetime import datetime as dt
import bs4
import re
def make_industry_zip_list_hash(data_frame, data_frame_name):
industry_zip_list_hash = {}
industry_li = make_type_of_industry_list(data_frame)
zip_files = call_zip_files(data_frame_name)
for industry in industry_li :
industry_zip_list_hash[industry]= list(filter(lambda x: industry in x , zip_files))
return industry_zip_list_hash
def make_type_of_industry_list(data_frame : pd.DataFrame, industry_col="[業種(東証)]"):
return list(set(data_frame[industry_col]))
def call_zip_files(data_frame_name):
zip_files = glob.glob(f"../**/SampleData/{data_frame_name}/**/**.zip", recursive=True)
return zip_files
def call_unziped_htm_files_dir(data_frame_name: str):
unziped_htm_files_dir = os.getcwd()+ "/UnzipedHtmFiles"
if not os.path.exists(unziped_htm_files_dir) :
os.mkdir(unziped_htm_files_dir)
unziped_htm_files_dir_with_df_name = unziped_htm_files_dir + f"/{data_frame_name}"
if not os.path.exists(unziped_htm_files_dir_with_df_name) :
os.mkdir(unziped_htm_files_dir_with_df_name)
return unziped_htm_files_dir_with_df_name
#--------------------------------------------------------------------
def unzip_html_to_unziped_htm_files_dir(industry_zip_list_hash, filepath_unziped):
sum_files_len = sum(map(len, industry_zip_list_hash.values()))
zip_files_len = 1
for industy_name, zip_files in industry_zip_list_hash.items() :
zip_files_len += len(zip_files)
industry_dir = call_industry_dir(filepath_unziped, industy_name)
for idx ,zip_file in enumerate(zip_files):
try:
with zipfile.ZipFile(zip_file) as existing_zip:
candidate_files = existing_zip.namelist()
for c_f in candidate_files:
basename = os.path.basename(c_f)
#第2部のもの(MD&Aが記載されている見出しの部分)を判定
if re.match(r'01020.*htm', basename) != None :
print(c_f)
date_str = c_f[-20:-10]
date_dt = dt.strptime(date_str, '%Y-%m-%d')
existing_zip.extract(c_f, industry_dir)
print(f"{idx + 1} / {len(zip_files)} || {zip_files_len - 1}/{sum_files_len}")
except Exception :
print(zip_file)
with open("unzipping_html_error.txt", "a") as f:
f.write(zip_file + "\n")
def call_industry_dir(filepath_unziped, industry_name: str):
industry_dir = filepath_unziped+ f"/{industry_name}"
if not os.path.exists(industry_dir) :
os.mkdir(industry_dir)
return industry_dir
f = call_zip_files("renketsu") + call_zip_files("hirenketsu")
len_f = len(f)
print(len_f, len_f/3)
#renkestsuとhirenketsuを設定
data_frame_name ="renketsu"
data_frame = pd.read_csv("/home/jovyan/1CalliingEdinetApi"+f"/EdinetIdxFiles/edinet_{data_frame_name}.csv", skiprows=4)
industry_zip_list_hash = make_industry_zip_list_hash(data_frame, data_frame_name)
filepath_unziped = call_unziped_htm_files_dir(data_frame_name)
unzip_html_to_unziped_htm_files_dir(industry_zip_list_hash, filepath_unziped)
```
| github_jupyter |
# Computer Vision Nanodegree
## Project: Image Captioning
---
In this notebook, you will train your CNN-RNN model.
You are welcome and encouraged to try out many different architectures and hyperparameters when searching for a good model.
This does have the potential to make the project quite messy! Before submitting your project, make sure that you clean up:
- the code you write in this notebook. The notebook should describe how to train a single CNN-RNN architecture, corresponding to your final choice of hyperparameters. You should structure the notebook so that the reviewer can replicate your results by running the code in this notebook.
- the output of the code cell in **Step 2**. The output should show the output obtained when training the model from scratch.
This notebook **will be graded**.
Feel free to use the links below to navigate the notebook:
- [Step 1](#step1): Training Setup
- [Step 2](#step2): Train your Model
- [Step 3](#step3): (Optional) Validate your Model
<a id='step1'></a>
## Step 1: Training Setup
In this step of the notebook, you will customize the training of your CNN-RNN model by specifying hyperparameters and setting other options that are important to the training procedure. The values you set now will be used when training your model in **Step 2** below.
You should only amend blocks of code that are preceded by a `TODO` statement. **Any code blocks that are not preceded by a `TODO` statement should not be modified**.
### Task #1
Begin by setting the following variables:
- `batch_size` - the batch size of each training batch. It is the number of image-caption pairs used to amend the model weights in each training step.
- `vocab_threshold` - the minimum word count threshold. Note that a larger threshold will result in a smaller vocabulary, whereas a smaller threshold will include rarer words and result in a larger vocabulary.
- `vocab_from_file` - a Boolean that decides whether to load the vocabulary from file.
- `embed_size` - the dimensionality of the image and word embeddings.
- `hidden_size` - the number of features in the hidden state of the RNN decoder.
- `num_epochs` - the number of epochs to train the model. We recommend that you set `num_epochs=3`, but feel free to increase or decrease this number as you wish. [This paper](https://arxiv.org/pdf/1502.03044.pdf) trained a captioning model on a single state-of-the-art GPU for 3 days, but you'll soon see that you can get reasonable results in a matter of a few hours! (_But of course, if you want your model to compete with current research, you will have to train for much longer._)
- `save_every` - determines how often to save the model weights. We recommend that you set `save_every=1`, to save the model weights after each epoch. This way, after the `i`th epoch, the encoder and decoder weights will be saved in the `models/` folder as `encoder-i.pkl` and `decoder-i.pkl`, respectively.
- `print_every` - determines how often to print the batch loss to the Jupyter notebook while training. Note that you **will not** observe a monotonic decrease in the loss function while training - this is perfectly fine and completely expected! You are encouraged to keep this at its default value of `100` to avoid clogging the notebook, but feel free to change it.
- `log_file` - the name of the text file containing - for every step - how the loss and perplexity evolved during training.
If you're not sure where to begin to set some of the values above, you can peruse [this paper](https://arxiv.org/pdf/1502.03044.pdf) and [this paper](https://arxiv.org/pdf/1411.4555.pdf) for useful guidance! **To avoid spending too long on this notebook**, you are encouraged to consult these suggested research papers to obtain a strong initial guess for which hyperparameters are likely to work best. Then, train a single model, and proceed to the next notebook (**3_Inference.ipynb**). If you are unhappy with your performance, you can return to this notebook to tweak the hyperparameters (and/or the architecture in **model.py**) and re-train your model.
### Question 1
**Question:** Describe your CNN-RNN architecture in detail. With this architecture in mind, how did you select the values of the variables in Task 1? If you consulted a research paper detailing a successful implementation of an image captioning model, please provide the reference.
**Answer:** I referenced the two papers suggested above to come up with an initial design of my CNN-RNN architecture. The CNN architecture was provided in the initial project code and is a pre-trained ResNet-50 model. My RNN architecture is based on the second paper, "Show and Tell: A Neural Image Caption Generator". Thus, I chose `vocab_threshold` of 5, `embed_size` of 512, and `hidden_size` of 512. I think 512 is a good choice because a large word embedding increases the chance of learning useful information. Additionally, I selected a `batch_size` of 128, since it is a power of 2 (taking advantage of vector optimizations) and batch sizes of 128 and 256 are commonly used.
### (Optional) Task #2
Note that we have provided a recommended image transform `transform_train` for pre-processing the training images, but you are welcome (and encouraged!) to modify it as you wish. When modifying this transform, keep in mind that:
- the images in the dataset have varying heights and widths, and
- if using a pre-trained model, you must perform the corresponding appropriate normalization.
### Question 2
**Question:** How did you select the transform in `transform_train`? If you left the transform at its provided value, why do you think that it is a good choice for your CNN architecture?
**Answer:** I left `transform_train` at its provided value. Since I used the CNN architecture as provided, I kept the transform function unchanged. By applying random cropping, the image transform extends the amount of data for training and makes the neural net more robust. Additionally, horizontal flipping makes sense because images are more likely to be mirrored across the vertical axis. A dog facing left and a dog facing right should be interpreted as dogs in a similar position. Normalization is also an important step. The data augmentation introduced by the image transformation function makes it a good choice for the CNN architecture.
### Task #3
Next, you will specify a Python list containing the learnable parameters of the model. For instance, if you decide to make all weights in the decoder trainable, but only want to train the weights in the embedding layer of the encoder, then you should set `params` to something like:
```
params = list(decoder.parameters()) + list(encoder.embed.parameters())
```
### Question 3
**Question:** How did you select the trainable parameters of your architecture? Why do you think this is a good choice?
**Answer:** I selected the trainable parameters of my architecture based on the recommended values. All the weights in the decoder and only the weights in the embedding layer of the encoder are trained, while the other parameters of the encoder won't be trained since we're using a pre-trained model.
### Task #4
Finally, you will select an [optimizer](http://pytorch.org/docs/master/optim.html#torch.optim.Optimizer).
### Question 4
**Question:** How did you select the optimizer used to train your model?
**Answer:** I initially used I used SGD since the paper recommends it. After experimentation, I decided to go with the Adam optimizer to train my final model. SGD was very slow and Adam was faster and produced significantly better perplexity scores (with perplexity <30). Models that are better at predicting a sample have low perplexity.
```
import nltk
nltk.download('punkt')
% load_ext autoreload
import torch
import torch.nn as nn
from torchvision import transforms
import sys
sys.path.append('/opt/cocoapi/PythonAPI')
from pycocotools.coco import COCO
from data_loader import get_loader
from model import EncoderCNN, DecoderRNN
import math
## TODO #1: Select appropriate values for the Python variables below.
batch_size = 128 # batch size
vocab_threshold = 5 # minimum word count threshold
vocab_from_file = True # if True, load existing vocab file
embed_size = 512 # dimensionality of image and word embeddings
hidden_size = 512 # number of features in hidden state of the RNN decoder
num_epochs = 3 # number of training epochs
save_every = 1 # determines frequency of saving model weights
print_every = 100 # determines window for printing average loss
log_file = 'training_log.txt' # name of file with saved training loss and perplexity
# (Optional) TODO #2: Amend the image transform below.
transform_train = transforms.Compose([
transforms.Resize(256), # smaller edge of image resized to 256
transforms.RandomCrop(224), # get 224x224 crop from random location
transforms.RandomHorizontalFlip(), # horizontally flip image with probability=0.5
transforms.ToTensor(), # convert the PIL Image to a tensor
transforms.Normalize((0.485, 0.456, 0.406), # normalize image for pre-trained model
(0.229, 0.224, 0.225))])
# Build data loader.
data_loader = get_loader(transform=transform_train,
mode='train',
batch_size=batch_size,
vocab_threshold=vocab_threshold,
vocab_from_file=vocab_from_file)
# The size of the vocabulary.
vocab_size = len(data_loader.dataset.vocab)
# Initialize the encoder and decoder.
encoder = EncoderCNN(embed_size)
decoder = DecoderRNN(embed_size, hidden_size, vocab_size)
# Move models to GPU if CUDA is available.
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
encoder.to(device)
decoder.to(device)
# Define the loss function.
criterion = nn.CrossEntropyLoss().cuda() if torch.cuda.is_available() else nn.CrossEntropyLoss()
# TODO #3: Specify the learnable parameters of the model.
params = list(decoder.parameters()) + list(encoder.embed.parameters())
# TODO #4: Define the optimizer.
optimizer = torch.optim.Adam(params)
# Set the total number of training steps per epoch.
total_step = math.ceil(len(data_loader.dataset.caption_lengths) / data_loader.batch_sampler.batch_size)
```
<a id='step2'></a>
## Step 2: Train your Model
Once you have executed the code cell in **Step 1**, the training procedure below should run without issue.
It is completely fine to leave the code cell below as-is without modifications to train your model. However, if you would like to modify the code used to train the model below, you must ensure that your changes are easily parsed by your reviewer. In other words, make sure to provide appropriate comments to describe how your code works!
You may find it useful to load saved weights to resume training. In that case, note the names of the files containing the encoder and decoder weights that you'd like to load (`encoder_file` and `decoder_file`). Then you can load the weights by using the lines below:
```python
# Load pre-trained weights before resuming training.
encoder.load_state_dict(torch.load(os.path.join('./models', encoder_file)))
decoder.load_state_dict(torch.load(os.path.join('./models', decoder_file)))
```
While trying out parameters, make sure to take extensive notes and record the settings that you used in your various training runs. In particular, you don't want to encounter a situation where you've trained a model for several hours but can't remember what settings you used :).
### A Note on Tuning Hyperparameters
To figure out how well your model is doing, you can look at how the training loss and perplexity evolve during training - and for the purposes of this project, you are encouraged to amend the hyperparameters based on this information.
However, this will not tell you if your model is overfitting to the training data, and, unfortunately, overfitting is a problem that is commonly encountered when training image captioning models.
For this project, you need not worry about overfitting. **This project does not have strict requirements regarding the performance of your model**, and you just need to demonstrate that your model has learned **_something_** when you generate captions on the test data. For now, we strongly encourage you to train your model for the suggested 3 epochs without worrying about performance; then, you should immediately transition to the next notebook in the sequence (**3_Inference.ipynb**) to see how your model performs on the test data. If your model needs to be changed, you can come back to this notebook, amend hyperparameters (if necessary), and re-train the model.
That said, if you would like to go above and beyond in this project, you can read about some approaches to minimizing overfitting in section 4.3.1 of [this paper](http://ieeexplore.ieee.org/stamp/stamp.jsp?arnumber=7505636). In the next (optional) step of this notebook, we provide some guidance for assessing the performance on the validation dataset.
```
import torch.utils.data as data
import numpy as np
import os
import requests
import time
# Open the training log file.
f = open(log_file, 'w')
old_time = time.time()
response = requests.request("GET",
"http://metadata.google.internal/computeMetadata/v1/instance/attributes/keep_alive_token",
headers={"Metadata-Flavor":"Google"})
for epoch in range(1, num_epochs+1):
for i_step in range(1, total_step+1):
if time.time() - old_time > 60:
old_time = time.time()
requests.request("POST",
"https://nebula.udacity.com/api/v1/remote/keep-alive",
headers={'Authorization': "STAR " + response.text})
# Randomly sample a caption length, and sample indices with that length.
indices = data_loader.dataset.get_train_indices()
# Create and assign a batch sampler to retrieve a batch with the sampled indices.
new_sampler = data.sampler.SubsetRandomSampler(indices=indices)
data_loader.batch_sampler.sampler = new_sampler
# Obtain the batch.
images, captions = next(iter(data_loader))
# Move batch of images and captions to GPU if CUDA is available.
images = images.to(device)
captions = captions.to(device)
# Zero the gradients.
decoder.zero_grad()
encoder.zero_grad()
# Pass the inputs through the CNN-RNN model.
features = encoder(images)
outputs = decoder(features, captions)
# Calculate the batch loss.
loss = criterion(outputs.view(-1, vocab_size), captions.view(-1))
# Backward pass.
loss.backward()
# Update the parameters in the optimizer.
optimizer.step()
# Get training statistics.
stats = 'Epoch [%d/%d], Step [%d/%d], Loss: %.4f, Perplexity: %5.4f' % (epoch, num_epochs, i_step, total_step, loss.item(), np.exp(loss.item()))
# Print training statistics (on same line).
print('\r' + stats, end="")
sys.stdout.flush()
# Print training statistics to file.
f.write(stats + '\n')
f.flush()
# Print training statistics (on different line).
if i_step % print_every == 0:
print('\r' + stats)
# Save the weights.
if epoch % save_every == 0:
torch.save(decoder.state_dict(), os.path.join('./models', 'decoder-%d.pkl' % epoch))
torch.save(encoder.state_dict(), os.path.join('./models', 'encoder-%d.pkl' % epoch))
# Close the training log file.
f.close()
```
<a id='step3'></a>
## Step 3: (Optional) Validate your Model
To assess potential overfitting, one approach is to assess performance on a validation set. If you decide to do this **optional** task, you are required to first complete all of the steps in the next notebook in the sequence (**3_Inference.ipynb**); as part of that notebook, you will write and test code (specifically, the `sample` method in the `DecoderRNN` class) that uses your RNN decoder to generate captions. That code will prove incredibly useful here.
If you decide to validate your model, please do not edit the data loader in **data_loader.py**. Instead, create a new file named **data_loader_val.py** containing the code for obtaining the data loader for the validation data. You can access:
- the validation images at filepath `'/opt/cocoapi/images/train2014/'`, and
- the validation image caption annotation file at filepath `'/opt/cocoapi/annotations/captions_val2014.json'`.
The suggested approach to validating your model involves creating a json file such as [this one](https://github.com/cocodataset/cocoapi/blob/master/results/captions_val2014_fakecap_results.json) containing your model's predicted captions for the validation images. Then, you can write your own script or use one that you [find online](https://github.com/tylin/coco-caption) to calculate the BLEU score of your model. You can read more about the BLEU score, along with other evaluation metrics (such as TEOR and Cider) in section 4.1 of [this paper](https://arxiv.org/pdf/1411.4555.pdf). For more information about how to use the annotation file, check out the [website](http://cocodataset.org/#download) for the COCO dataset.
```
# (Optional) TODO: Validate your model.
```
| github_jupyter |
# Detect data bias with Amazon SageMaker Clarify
### Introduction
Bias can be present in your data before any model training occurs. Inspecting the dataset for bias can help detect collection gaps, inform your feature engineering, and understand societal biases the dataset may reflect. In this lab you will analyze bias on the dataset, generate and analyze bias report, and prepare the dataset for the model training.
### Table of Contents
- [1. Analyze the dataset](#c1w2-1.)
- [1.1. Create a pandas data frame from the CSV file](#c1w2-1.1.)
- [1.2. Upload the dataset to S3 bucket](#c1w2-1.2.)
- [2. Analyze class imbalance on the dataset with Amazon SageMaker Clarify](#c1w2-2.)
- [2.1. Configure a `DataConfig`](#c1w2-2.1.)
- [Exercise 1](#c1w2-ex-1)
- [2.2. Configure `BiasConfig`](#c1w2-2.2.)
- [2.3. Configure Amazon SageMaker Clarify as a processing job](#c1w2-2.3.)
- [2.4. Run the Amazon SageMaker Clarify processing job](#c1w2-2.4.)
- [Exercise 2](#c1w2-ex-2)
- [2.5. Run and review the Amazon SageMaker Clarify processing job on the unbalanced dataset](#c1w2-2.5.)
- [2.6. Analyze unbalanced bias report](#c1w2-2.6.)
- [3. Balance the dataset by `product_category` and `sentiment`](#c1w2-3.)
- [4. Analyze bias on balanced dataset with Amazon SageMaker Clarify](#c1w2-4.)
- [4.1. Configure a `DataConfig`](#c1w2-4.1.)
- [Exercise 3](#c1w2-ex-3)
- [4.2. Configure `BiasConfig`](#c1w2-4.2.)
- [4.3. Configure SageMaker Clarify as a processing job](#c1w2-4.3.)
- [4.4. Run the Amazon SageMaker Clarify processing job](#c1w2-4.4.)
- [Exercise 4](#c1w2-ex-4)
- [4.5. Run and review the Clarify processing job on the balanced dataset](#c1w2-4.5.)
- [4.6. Analyze balanced bias report](#c1w2-4.6.)
First, let's install and import required modules.
```
# please ignore warning messages during the installation
!pip install --disable-pip-version-check -q sagemaker==2.35.0
import boto3
import sagemaker
import pandas as pd
import numpy as np
sess = sagemaker.Session()
bucket = sess.default_bucket()
role = sagemaker.get_execution_role()
region = boto3.Session().region_name
import matplotlib.pyplot as plt
%matplotlib inline
%config InlineBackend.figure_format='retina'
```
<a name='c1w2-1.'></a>
# 1. Analyze the dataset
<a name='c1w2-1.1.'></a>
### 1.1. Create a pandas data frame from the CSV file
Create a pandas dataframe from each of the product categories and concatenate them into one.
```
!aws s3 cp 's3://dlai-practical-data-science/data/transformed/womens_clothing_ecommerce_reviews_transformed.csv' ./
path = './womens_clothing_ecommerce_reviews_transformed.csv'
df = pd.read_csv(path)
df.head()
```
As you saw in the previous lab, there are way more positive reviews than negative or neutral. Such a dataset is called unbalanced.
In this case, using a relatively small data subset you could visualize the occurring unbalances. At scale, you would need to perform bias analysis. Let's use this dataset as an example.
```
import seaborn as sns
sns.countplot(data=df, x='sentiment', hue='product_category')
plt.legend(loc='upper right',bbox_to_anchor=(1.3, 1.1))
```
<a name='c1w2-1.2.'></a>
### 1.2. Upload the dataset to S3 bucket
Upload the dataset to a private S3 bucket in a folder called `bias/unbalanced`.
```
data_s3_uri_unbalanced = sess.upload_data(bucket=bucket,
key_prefix='bias/unbalanced',
path='./womens_clothing_ecommerce_reviews_transformed.csv')
data_s3_uri_unbalanced
```
You can review the uploaded CSV file in the S3 bucket.
**Instructions**:
- open the link
- click on the S3 bucket name `sagemaker-us-east-1-ACCOUNT`
- go to the folder `bias/unbalanced`
- check the existence of the file `womens_clothing_ecommerce_reviews_transformed.csv`
```
from IPython.core.display import display, HTML
display(HTML('<b>Review <a target="top" href="https://s3.console.aws.amazon.com/s3/home?region={}#">Amazon S3 bucket</a></b>'.format(region)))
```
<a name='c1w2-2.'></a>
# 2. Analyze class imbalance on the dataset with Amazon SageMaker Clarify
Let's analyze bias in `sentiment` with respect to the `product_category` facet on the dataset.
<a name='c1w2-2.1.'></a>
### 2.1. Configure a `DataConfig`
Information about the input data needs to be provided to the processor. This can be done with the `DataConfig` of the Clarify container. It stores information about the dataset to be analyzed, for example the dataset file, its format, headers and labels.
<a name='c1w2-ex-1'></a>
### Exercise 1
Configure a `DataConfig` for Clarify.
**Instructions**: Use `DataConfig` to configure the target column (`'sentiment'` label), data input (`data_s3_uri_unbalanced`) and output paths (`bias_report_unbalanced_output_path`) with their formats (header names and the dataset type):
```python
data_config_unbalanced = clarify.DataConfig(
s3_data_input_path=..., # S3 object path containing the unbalanced dataset
s3_output_path=..., # path to store the output
label='...', # target column
headers=df_unbalanced.columns.to_list(),
dataset_type='text/csv'
)
```
```
from sagemaker import clarify
bias_report_unbalanced_output_path = 's3://{}/bias/generated_bias_report/unbalanced'.format(bucket)
data_config_unbalanced = clarify.DataConfig(
### BEGIN SOLUTION - DO NOT delete this comment for grading purposes
s3_data_input_path=data_s3_uri_unbalanced, # Replace None
s3_output_path=bias_report_unbalanced_output_path, # Replace None
label='sentiment', # Replace None
### END SOLUTION - DO NOT delete this comment for grading purposes
headers=df.columns.to_list(),
dataset_type='text/csv'
)
```
<a name='c1w2-2.2.'></a>
### 2.2. Configure `BiasConfig`
Bias is measured by calculating a metric and comparing it across groups. To compute it, you will specify the required information in the `BiasConfig` API. SageMaker Clarify needs the sensitive columns (`facet_name`) and the desirable outcomes (`label_values_or_threshold`). Here `product_category` is the sensitive facet and the desired outcome is with the `sentiment==1`.
SageMaker Clarify can handle both categorical and continuous data for `label_values_or_threshold`. In this case you are using categorical data.
```
bias_config_unbalanced = clarify.BiasConfig(
label_values_or_threshold=[1], # desired sentiment
facet_name='product_category' # sensitive column (facet)
)
```
<a name='c1w2-2.3.'></a>
### 2.3. Configure Amazon SageMaker Clarify as a processing job
Now you need to construct an object called `SageMakerClarifyProcessor`. This allows you to scale the process of data bias detection using two parameters, `instance_count` and `instance_type`. `Instance_count` represents how many nodes you want in the distributor cluster during the data detection. `Instance_type` specifies the processing capability (compute capacity, memory capacity) available for each one of those nodes.
```
clarify_processor_unbalanced = clarify.SageMakerClarifyProcessor(role=role,
instance_count=1,
instance_type='ml.m5.large',
sagemaker_session=sess)
```
<a name='c1w2-2.4.'></a>
### 2.4. Run the Amazon SageMaker Clarify processing job
<a name='c1w2-ex-2'></a>
### Exercise 2
Run the configured processing job to compute the requested bias `methods` of the input data
**Instructions**: Apply the `run_pre_training_bias` method to the configured Clarify processor, passing the configured input/output data (`data_config_unbalanced`), configuration of sensitive groups (`bias_config_unbalanced`) with the other job setup parameters:
```python
clarify_processor_unbalanced.run_pre_training_bias(
data_config=..., # configured input/output data
data_bias_config=..., # configured sensitive groups
methods=["CI", "DPL", "KL", "JS", "LP", "TVD", "KS"], # selector of a subset of potential metrics
wait=False, # whether the call should wait until the job completes (default: True)
logs=False # whether to show the logs produced by the job. Only meaningful when wait is True (default: True)
)
```
```
clarify_processor_unbalanced.run_pre_training_bias(
### BEGIN SOLUTION - DO NOT delete this comment for grading purposes
data_config=data_config_unbalanced, # Replace None
data_bias_config=bias_config_unbalanced, # Replace None
### END SOLUTION - DO NOT delete this comment for grading purposes
methods=["CI", "DPL", "KL", "JS", "LP", "TVD", "KS"],
wait=False,
logs=False
)
run_unbalanced_bias_processing_job_name = clarify_processor_unbalanced.latest_job.job_name
print(run_unbalanced_bias_processing_job_name)
```
<a name='c1w2-2.5.'></a>
### 2.5. Run and review the Amazon SageMaker Clarify processing job on the unbalanced dataset
Review the created Amazon SageMaker Clarify processing job and the Cloud Watch logs.
**Instructions**:
- open the link
- note that you are in the section Amazon SageMaker -> Processing jobs
- check the processing job name
- note which other properties of the processing job you can see in the console
```
from IPython.core.display import display, HTML
display(HTML('<b>Review <a target="blank" href="https://console.aws.amazon.com/sagemaker/home?region={}#/processing-jobs/{}">processing job</a></b>'.format(region, run_unbalanced_bias_processing_job_name)))
```
**Instructions**:
- open the link
- open the log stream with the name, which starts from the processing job name
- have a quick look at the log messages
```
from IPython.core.display import display, HTML
display(HTML('<b>Review <a target="blank" href="https://console.aws.amazon.com/cloudwatch/home?region={}#logStream:group=/aws/sagemaker/ProcessingJobs;prefix={};streamFilter=typeLogStreamPrefix">CloudWatch logs</a> after about 5 minutes</b>'.format(region, run_unbalanced_bias_processing_job_name)))
running_processor = sagemaker.processing.ProcessingJob.from_processing_name(processing_job_name=run_unbalanced_bias_processing_job_name,
sagemaker_session=sess)
```
### _This cell will take approximately 5-10 minutes to run._
```
%%time
running_processor.wait(logs=False)
```
<a name='c1w2-2.6.'></a>
### 2.6. Analyze unbalanced bias report
In this run, you analyzed bias for `sentiment` relative to the `product_category` for the unbalanced data. Let's have a look at the bias report.
List the files in the output path `bias_report_unbalanced_output_path`:
```
!aws s3 ls $bias_report_unbalanced_output_path/
```
Download generated bias report from S3 bucket:
```
!aws s3 cp --recursive $bias_report_unbalanced_output_path ./generated_bias_report/unbalanced/
```
Review the downloaded bias report (in HTML format):
```
from IPython.core.display import display, HTML
display(HTML('<b>Review <a target="blank" href="./generated_bias_report/unbalanced/report.html">unbalanced bias report</a></b>'))
```
The bias report shows a number of metrics, but here you can focus on just two of them:
- Class Imbalance (CI). Measures the imbalance in the number of members between different facet values. Answers the question, does a `product_category` have disproportionately more reviews than others? Values of CI will become equal for even distribution between facets. Here, different CI values show the existence of imbalance.
- Difference in Positive Proportions in Labels (DPL). Measures the imbalance of positive outcomes between different facet values. Answers the question, does a `product_category` have disproportionately higher ratings than others? With the range over the interval from -1 to 1, if there is no bias, you want to see this value as close as possible to zero. Here, non-zero values indicate the imbalances.
<a name='c1w2-3.'></a>
# 3. Balance the dataset by `product_category` and `sentiment`
Let's balance the dataset by `product_category` and `sentiment`. Then you can configure and run SageMaker Clarify processing job to analyze the bias of it. Which metrics values do you expect to see in the bias report?
```
df_grouped_by = df.groupby(['product_category', 'sentiment'])
df_balanced = df_grouped_by.apply(lambda x: x.sample(df_grouped_by.size().min()).reset_index(drop=True))
df_balanced
```
Visualize the distribution of review sentiment in the balanced dataset.
```
import seaborn as sns
sns.countplot(data=df_balanced, x='sentiment', hue='product_category')
plt.legend(loc='upper right',bbox_to_anchor=(1.3, 1.1))
```
<a name='c1w2-4.'></a>
# 4. Analyze bias on balanced dataset with Amazon SageMaker Clarify
Let's analyze bias in `sentiment` with respect to the `product_category` facet on your balanced dataset.
Save and upload balanced data to S3 bucket.
```
path_balanced = './womens_clothing_ecommerce_reviews_balanced.csv'
df_balanced.to_csv(path_balanced, index=False, header=True)
data_s3_uri_balanced = sess.upload_data(bucket=bucket, key_prefix='bias/balanced', path=path_balanced)
data_s3_uri_balanced
```
You can review the uploaded CSV file in the S3 bucket and prefix `bias/balanced`.
```
from IPython.core.display import display, HTML
display(HTML('<b>Review <a target="top" href="https://s3.console.aws.amazon.com/s3/home?region={}#">Amazon S3 bucket</a></b>'.format(region)))
```
<a name='c1w2-4.1.'></a>
### 4.1. Configure a `DataConfig`
<a name='c1w2-ex-3'></a>
### Exercise 3
Configure a `DataConfig` for Clarify to analyze bias on the balanced dataset.
**Instructions**: Pass the S3 object path containing the balanced dataset, the path to store the output (`bias_report_balanced_output_path`) and the target column. You can use exercise 1 as an example.
```
from sagemaker import clarify
bias_report_balanced_output_path = 's3://{}/bias/generated_bias_report/balanced'.format(bucket)
data_config_balanced = clarify.DataConfig(
### BEGIN SOLUTION - DO NOT delete this comment for grading purposes
s3_data_input_path=data_s3_uri_balanced, # Replace None
s3_output_path=bias_report_balanced_output_path, # Replace None
label='sentiment', # Replace None
### END SOLUTION - DO NOT delete this comment for grading purposes
headers=df_balanced.columns.to_list(),
dataset_type='text/csv'
)
```
<a name='c1w2-4.2.'></a>
### 4.2. Configure `BiasConfig`
`BiasConfig` for the balanced dataset will have the same settings as before.
```
bias_config_balanced = clarify.BiasConfig(
label_values_or_threshold=[1], # desired sentiment
facet_name='product_category' # sensitive column (facet)
)
```
<a name='c1w2-4.3.'></a>
### 4.3. Configure SageMaker Clarify as a processing job
`SageMakerClarifyProcessor` object will also have the same parameters.
```
clarify_processor_balanced = clarify.SageMakerClarifyProcessor(role=role,
instance_count=1,
instance_type='ml.m5.large',
sagemaker_session=sess)
```
<a name='c1w2-4.4.'></a>
### 4.4. Run the Amazon SageMaker Clarify processing job
<a name='c1w2-ex-4'></a>
### Exercise 4
Run the configured processing job for the balanced dataset.
**Instructions**: Apply the `run_pre_training_bias` method to the configured Clarify processor, passing the input/output data, configuration of sensitive groups with the other job setup parameters. You can use exercise 2 as an example.
```
clarify_processor_balanced.run_pre_training_bias(
### BEGIN SOLUTION - DO NOT delete this comment for grading purposes
data_config=data_config_balanced, # Replace None
data_bias_config=bias_config_balanced, # Replace None
### END SOLUTION - DO NOT delete this comment for grading purposes
methods=["CI", "DPL", "KL", "JS", "LP", "TVD", "KS"],
wait=False,
logs=False
)
run_balanced_bias_processing_job_name = clarify_processor_balanced.latest_job.job_name
print(run_balanced_bias_processing_job_name)
```
<a name='c1w2-4.5.'></a>
### 4.5. Run and review the Clarify processing job on the balanced dataset
Review the results of the run following the links:
```
from IPython.core.display import display, HTML
display(HTML('<b>Review <a target="blank" href="https://console.aws.amazon.com/sagemaker/home?region={}#/processing-jobs/{}">processing job</a></b>'.format(region, run_balanced_bias_processing_job_name)))
from IPython.core.display import display, HTML
display(HTML('<b>Review <a target="blank" href="https://console.aws.amazon.com/cloudwatch/home?region={}#logStream:group=/aws/sagemaker/ProcessingJobs;prefix={};streamFilter=typeLogStreamPrefix">CloudWatch logs</a> after about 5 minutes</b>'.format(region, run_balanced_bias_processing_job_name)))
running_processor = sagemaker.processing.ProcessingJob.from_processing_name(processing_job_name=run_balanced_bias_processing_job_name,
sagemaker_session=sess)
```
### _This cell will take approximately 5-10 minutes to run._
```
%%time
running_processor.wait(logs=False)
```
<a name='c1w2-4.6.'></a>
### 4.6. Analyze balanced bias report
List the files in the output path `bias_report_balanced_output_path`:
```
!aws s3 ls $bias_report_balanced_output_path/
```
Download generated bias report from S3 bucket:
```
!aws s3 cp --recursive $bias_report_balanced_output_path ./generated_bias_report/balanced/
```
Review the downloaded bias report (in HTML format):
```
from IPython.core.display import display, HTML
display(HTML('<b>Review <a target="blank" href="./generated_bias_report/balanced/report.html">balanced bias report</a></b>'))
```
In this run, you analyzed bias for `sentiment` relative to the `product_category` for the balanced data. Note that the Class Imbalance (CI) metric is equal across all product categories for the target label, `sentiment`. And Difference in Positive Proportions in Labels (DPL) metric values are zero.
Upload the notebook into S3 bucket for grading purposes.
**Note**: you may need to click on "Save" button before the upload.
```
!aws s3 cp ./C1_W2_Assignment.ipynb s3://$bucket/C1_W2_Assignment_Learner.ipynb
```
Please go to the main lab window and click on `Submit` button (see the `Finish the lab` section of the instructions).
| github_jupyter |
## Recursive Functions
A recursive function is a function that makes calls to itself. It works like the loops we described before, but sometimes it the situation is better to use recursion than loops.
Every recursive function has two components: a base case and a recursive step. The base case is usually the smallest input and has an easily verifiable solution. This is also the mechanism that stops the function from calling itself forever. The recursive step is the set of all cases where a recursive call, or a function call to itself, is made.
Consider the example of computing the factorial of a number. For example, the factorial of a number $n$ is given by $f(n) = 1 \ \times \ 2 \ \times \ 3 \ \times \ \dots \ \times \ (n-1) \ \times \ n$.
The recursive from of a factorial is
$$
f(n) = \left\{ \begin{array}{ll} 1 & if \ n=1 \\
n \ \times \ f(n-1) & otherwise\end{array} \right.
$$
which can be expressed in code as
```
def factorial_n(n):
assert type(n) == int, 'Input must be an integer'
if n == 1: #this is the base case
return 1
else: #this is the recursive step
return n * factorial_n(n-1)
factorial_n(1)
factorial_n(2)
factorial_n(5)
1*2*3*4*5
#We can use debbuging tools to understand the code
from pdb import set_trace
def factorial_n(n):
assert type(n) == int, 'Input must be an integer'
set_trace()
if n == 1: #this is the base case
return 1
else: #this is the recursive step
return n * factorial_n(n-1)
factorial_n(1)
factorial_n(3)
```
## mini challenge 1
Fibonacci numbers were originally developed to model the idealized population growth of rabbits. Since then, they have been found to be significant in any naturally occurring phenomena.
Use recursivity to compute the Fibonacci numbers.
The recursive form of the Fibonacci numbers.
$$
f(n) = \left\{ \begin{array}{ll} 1 & if \ n=1 \\
1 & if \ n=2 \\
f(n-1) + f(n-2) & otherwise\end{array} \right.
$$
```
#examples
fibonacci(1) = 1
fibonacci(2) = 1
fibonacci(3) = 2
fibonacci(4) = 3
fibonacci(5) = 5
fibonacci(35) = 9227465
def fibonacci(n) :
assert type(n) == int, 'Input must be an integer'
if n == 1:
return 1
if n == 2:
return 1
else:
return fibonacci(n-1) + fibonacci(n-2)
```
## mini challenge 2
An integer number $n$ is said to be **prime** if is divisible only by itself and one. If $n$ is divisible by any other number between $1$ and $n$, the the number is not prime.
Write a recursive function to verify if a number n is prime.
```
def prime(N, div = 2):
if N == 1:
return True
else:
if N == 2:
return True
elif (N%div) == 0 :
return False
else:
prime(N,div+1)
return True
prime(7)
```
| github_jupyter |
```
!pip install pytest ipytest pytest-csv pytest-benchmark
import numpy as np
import pytest
%matplotlib inline
from coffea import hist
import coffea.processor as processor
import awkward as ak
from dask.distributed import Client, LocalCluster
import time
import os
import ipytest
ipytest.config(rewrite_asserts=True, magics=True)
fileset = {'SingleMu' : ["root://eospublic.cern.ch//eos/root-eos/benchmark/Run2012B_SingleMu.root"]}
from dask.distributed import Client, Worker, WorkerPlugin
from typing import List
import os
class DependencyInstaller(WorkerPlugin):
def __init__(self, dependencies: List[str]):
self._depencendies = " ".join(f"'{dep}'" for dep in dependencies)
def setup(self, worker: Worker):
os.system(f"pip install {self._depencendies}")
dependency_installer = DependencyInstaller([
"pytest-benchmark",
])
client = Client("tls://localhost:8786")
#Uncomment only if we would like to compare the same number of workers
#cluster = CoffeaCasaCluster()
#cluster.scale(10)
#client = Client(cluster)
client.register_worker_plugin(dependency_installer)
# This program plots an event-level variable (in this case, MET, but switching it is as easy as a dict-key change). It also demonstrates an easy use of the book-keeping cutflow tool, to keep track of the number of events processed.
# The processor class bundles our data analysis together while giving us some helpful tools. It also leaves looping and chunks to the framework instead of us.
class Processor(processor.ProcessorABC):
def __init__(self):
# Bins and categories for the histogram are defined here. For format, see https://coffeateam.github.io/coffea/stubs/coffea.hist.hist_tools.Hist.html && https://coffeateam.github.io/coffea/stubs/coffea.hist.hist_tools.Bin.html
dataset_axis = hist.Cat("dataset", "")
MET_axis = hist.Bin("MET", "MET [GeV]", 50, 0, 100)
# The accumulator keeps our data chunks together for histogramming. It also gives us cutflow, which can be used to keep track of data.
self._accumulator = processor.dict_accumulator({
'MET': hist.Hist("Counts", dataset_axis, MET_axis),
'cutflow': processor.defaultdict_accumulator(int)
})
@property
def accumulator(self):
return self._accumulator
def process(self, events):
output = self.accumulator.identity()
# This is where we do our actual analysis. The dataset has columns similar to the TTree's; events.columns can tell you them, or events.[object].columns for deeper depth.
dataset = events.metadata["dataset"]
MET = events.MET.pt
# We can define a new key for cutflow (in this case 'all events'). Then we can put values into it. We need += because it's per-chunk (demonstrated below)
output['cutflow']['all events'] += ak.size(MET)
output['cutflow']['number of chunks'] += 1
# This fills our histogram once our data is collected. The hist key ('MET=') will be defined in the bin in __init__.
output['MET'].fill(dataset=dataset, MET=MET)
return output
def postprocess(self, accumulator):
return accumulator
# Function which we are interested to benchmark where chunk_size is changed dependending on iteration of benchmark run.
def coffea_processor_adlexample1(chunk_size):
output = processor.run_uproot_job(fileset,
treename = 'Events',
processor_instance = Processor(),
executor = processor.dask_executor,
chunksize = chunk_size,
executor_args = {'schema': processor.NanoAODSchema,
'client': client,
'savemetrics': True}
)
return output
@pytest.mark.parametrize("chunk_size", range(100000, 200000, 100000))
def test_coffea_processor_adlexample1(benchmark, chunk_size):
output = benchmark(coffea_processor_adlexample1, chunk_size)
# Custom metrics available with `savemetrics` option
benchmark.extra_info['events_s_thread'] = output[1]['entries'] / output[1]['processtime']
ipytest.run("-qq")
benchmark1.json
{
"min": 55.1104,
"max": 69.7225,
"mean": 61.5893
}
```
| github_jupyter |
```
from astropy.constants import G
import astropy.coordinates as coord
import astropy.table as at
import astropy.units as u
from astropy.time import Time
import matplotlib as mpl
import matplotlib.pyplot as plt
%matplotlib inline
import numpy as np
from gala.units import galactic, UnitSystem
from twobody import TwoBodyKeplerElements, KeplerOrbit
from twobody.anomaly import mean_anomaly_from_eccentric_anomaly
usys = UnitSystem(1e12*u.Msun, u.kpc, u.Gyr, u.radian)
true_m31_sky_c = coord.SkyCoord(
10.64628564*u.deg,
41.23456631*u.deg
)
```
## Simulate some Keplerian data
```
M1 = 1.4e12 * u.Msun
M2 = 2.4e12 * u.Msun
M = M1 + M2
a = 511 * u.kpc
eta = 4.3 * u.rad
e = 0.981
mean_anomaly = mean_anomaly_from_eccentric_anomaly(eta, e)
elem = TwoBodyKeplerElements(
a=a, m1=M1, m2=M2, e=e,
omega=0*u.rad, i=90*u.deg,
M0=0.*u.rad, t0=0. * u.Gyr,
units=galactic
)
orb1 = KeplerOrbit(elem.primary)
orb2 = KeplerOrbit(elem.secondary)
Romega = coord.matrix_utilities.rotation_matrix(elem.secondary.omega, 'z')
xyz1 = orb1.orbital_plane(0. * u.Gyr)
xyz2 = orb2.orbital_plane(0. * u.Gyr).transform(Romega)
xyz1, xyz2
time = (elem.P * (mean_anomaly / (2*np.pi*u.rad))).to(u.Gyr)
xyz1 = orb1.orbital_plane(time)
xyz2 = orb2.orbital_plane(time).transform(Romega)
(xyz1.without_differentials()
- xyz2.without_differentials()).norm().to(u.kpc)
a * (1 - e * np.cos(eta))
times = np.linspace(0, 1, 1024) * elem.P
xyzs1 = orb1.orbital_plane(times)
xyzs2 = orb2.orbital_plane(times).transform(Romega)
rs = (xyzs1.without_differentials()
- xyzs2.without_differentials()).norm().to(u.kpc)
fig, ax = plt.subplots(1, 1, figsize=(8, 8))
ax.plot(xyzs1.x, xyzs1.y, marker='')
ax.plot(xyzs2.x, xyzs2.y, marker='')
ax.plot(xyz1.x, xyz1.y, zorder=100, ms=10, color='tab:orange')
ax.plot(xyz2.x, xyz2.y, zorder=100, ms=10, color='tab:red')
ax.set_xlim(-2*a.value, 2*a.value)
ax.set_ylim(-2*a.value, 2*a.value)
plt.plot(times.value, rs.value)
dxs = xyzs1.without_differentials() - xyzs2.without_differentials()
dvs = xyzs1.differentials['s'] - xyzs2.differentials['s']
dx_cyl = dxs.represent_as(coord.CylindricalRepresentation)
dv_cyl = dvs.represent_as(coord.CylindricalDifferential, dxs)
vrads = dv_cyl.d_rho
vtans = (dx_cyl.rho * dv_cyl.d_phi).to(u.km/u.s, u.dimensionless_angles())
etas = np.linspace(0, 2*np.pi, 1024) * u.rad
mean_anoms = mean_anomaly_from_eccentric_anomaly(etas, e)
eq_times = elem.P * (mean_anoms / (2*np.pi*u.rad))
eq_vrad = np.sqrt(G * M / a) * (e * np.sin(etas)) / (1 - e * np.cos(etas))
eq_vtan = np.sqrt(G * M / a) * np.sqrt(1 - e**2) / (1 - e * np.cos(etas))
plt.plot(times.value, vrads.to_value(u.km/u.s))
plt.plot(times.value, vtans.to_value(u.km/u.s))
plt.plot(eq_times.value, eq_vrad.to_value(u.km/u.s))
plt.plot(eq_times.value, eq_vtan.to_value(u.km/u.s))
plt.ylim(-500, 500)
```
### Transform to ICRS
```
dx = xyz1.without_differentials() - xyz2.without_differentials()
dv = xyz1.differentials['s'] - xyz2.differentials['s']
dx_cyl = dx.represent_as(coord.CylindricalRepresentation)
dv_cyl = dv.represent_as(coord.CylindricalDifferential, dx)
vrad = dv_cyl.d_rho.to(u.km/u.s)
vtan = (dx_cyl.rho * dv_cyl.d_phi).to(u.km/u.s, u.dimensionless_angles())
r = dx.norm()
sun_galcen_dist = coord.Galactocentric().galcen_distance
gamma = coord.Galactocentric().galcen_coord.separation(true_m31_sky_c)
sun_m31_dist = (sun_galcen_dist * np.cos(gamma)) + np.sqrt(
r**2 - sun_galcen_dist**2 * np.sin(gamma)**2
)
r, sun_m31_dist
vscale = np.sqrt(G * M / a)
print(vscale.decompose(usys).value,
vrad.decompose(usys).value,
vtan.decompose(usys).value)
alpha = 32.4 * u.deg
galcen_pos = coord.SkyCoord(true_m31_sky_c.ra,
true_m31_sky_c.dec,
distance=sun_m31_dist)
galcen_pos = galcen_pos.transform_to(coord.Galactocentric())
# galcen_pos = coord.CartesianRepresentation(
# -375 * u.kpc, 605 * u.kpc, -279 * u.kpc)
# galcen_pos = coord.Galactocentric(galcen_pos / galcen_pos.norm() * r)
galcen_sph = galcen_pos.represent_as('spherical')
gc_Rz = coord.matrix_utilities.rotation_matrix(-galcen_sph.lon, 'z')
gc_Ry = coord.matrix_utilities.rotation_matrix(galcen_sph.lat, 'y')
gc_Rx = coord.matrix_utilities.rotation_matrix(alpha, 'x')
R = gc_Rz @ gc_Ry @ gc_Rx
fake_X = R @ [r.value, 0, 0] * r.unit
fake_V = R @ [vrad.to_value(u.km/u.s), vtan.to_value(u.km/u.s), 0.] * u.km/u.s
fake_galcen = coord.Galactocentric(*fake_X, *fake_V)
fake_icrs = fake_galcen.transform_to(coord.ICRS())
fake_icrs
```
## Check roundtripping
```
def tt_sph_to_xyz(r, lon, lat):
return [
r * np.cos(lon) * np.cos(lat),
r * np.sin(lon) * np.cos(lat),
r * np.sin(lat)
]
def tt_cross(a, b):
return np.array([
a[1]*b[2] - a[2]*b[1],
a[2]*b[0] - a[0]*b[2],
a[0]*b[1] - a[1]*b[0]
])
def tt_rotation_matrix(angle_rad, axis):
s = np.sin(angle_rad)
c = np.cos(angle_rad)
if axis == 'x':
R = np.array([
1., 0, 0,
0, c, s,
0, -s, c
])
elif axis == 'y':
R = np.array([
c, 0, -s,
0, 1., 0,
s, 0, c
])
elif axis == 'z':
R = np.array([
c, s, 0,
-s, c, 0,
0, 0, 1.
])
else:
raise ValueError('bork')
return np.reshape(R, (3, 3))
def ugh(m31_ra_rad, m31_dec_rad, m31_distance_kpc, r, vrad, vtan):
galcen_frame = coord.Galactocentric()
# tangent bases: ra, dec, r
M = np.array([
[-np.sin(m31_ra_rad), np.cos(m31_ra_rad), 0.],
[-np.sin(m31_dec_rad)*np.cos(m31_ra_rad), -np.sin(m31_dec_rad)*np.sin(m31_ra_rad), np.cos(m31_dec_rad)],
[np.cos(m31_dec_rad)*np.cos(m31_ra_rad), np.cos(m31_dec_rad)*np.sin(m31_ra_rad), np.sin(m31_dec_rad)]
])
# Matrix to go from ICRS to Galactocentric
R_I2G, offset_I2G = coord.builtin_frames.galactocentric.get_matrix_vectors(
galcen_frame, inverse=False)
dxyz_I2G = offset_I2G.xyz.to_value(usys['length'])
dvxyz_I2G = offset_I2G.differentials['s'].d_xyz.to_value(usys['velocity'])
# Matrix to go from Galactocentric to ICRS
R_G2I, offset_G2I = coord.builtin_frames.galactocentric.get_matrix_vectors(
galcen_frame, inverse=True)
dxyz_G2I = offset_G2I.xyz.to_value(usys['length'])
dvxyz_G2I = offset_G2I.differentials['s'].d_xyz.to_value(usys['velocity'])
m31_icrs_xyz = tt_sph_to_xyz(m31_distance_kpc,
m31_ra_rad, m31_dec_rad)
m31_galcen_xyz = np.dot(R_I2G, m31_icrs_xyz) + dxyz_I2G
m31_galcen_lon = np.arctan2(m31_galcen_xyz[1], m31_galcen_xyz[0])
m31_galcen_lat = np.arcsin(m31_galcen_xyz[2] / r)
xhat = m31_galcen_xyz / r
Rz = tt_rotation_matrix(-m31_galcen_lon, 'z')
print(gc_Ry)
Ry = tt_rotation_matrix(m31_galcen_lat, 'y')
print(Ry)
Rx = tt_rotation_matrix(alpha, 'x')
yhat = np.dot(np.dot(Rz, np.dot(Ry, Rx)), [0, 1, 0.])
zhat = tt_cross(xhat, yhat)
R_LGtoG = np.stack((xhat, yhat, zhat), axis=1)
print(R_LGtoG - R)
x_LG = np.array([r, 0., 0.])
v_LG = np.array([vrad, vtan, 0.])
x_I = np.dot(R_G2I, np.dot(R_LGtoG, x_LG)) + dxyz_G2I
v_I = np.dot(R_G2I, np.dot(R_LGtoG, v_LG)) + dvxyz_G2I
v_I_tangent_plane = np.dot(M, v_I) # alpha, delta, radial
shit1 = coord.CartesianRepresentation(*((R @ x_LG) * usys['length']))
shit2 = coord.CartesianDifferential(*((R @ v_LG) * usys['velocity']))
shit = coord.SkyCoord(shit1.with_differentials(shit2), frame=coord.Galactocentric())
return x_I, v_I, shit.transform_to(coord.ICRS()).velocity
ugh(fake_icrs.ra.radian, fake_icrs.dec.radian, fake_icrs.distance.to_value(u.kpc),
r.decompose(usys).value, vrad.decompose(usys).value, vtan.decompose(usys).value)
fake_icrs.velocity
def ugh2():
galcen_frame = coord.Galactocentric()
# Matrix to go from ICRS to Galactocentric
R_I2G, offset_I2G = coord.builtin_frames.galactocentric.get_matrix_vectors(
galcen_frame, inverse=False)
dxyz_I2G = offset_I2G.xyz.to_value(usys['length'])
dvxyz_I2G = offset_I2G.differentials['s'].d_xyz.to_value(usys['velocity'])
# Matrix to go from Galactocentric to ICRS
R_G2I, offset_G2I = coord.builtin_frames.galactocentric.get_matrix_vectors(
galcen_frame, inverse=True)
dxyz_G2I = offset_G2I.xyz.to_value(usys['length'])
dvxyz_G2I = offset_G2I.differentials['s'].d_xyz.to_value(usys['velocity'])
m31_icrs_xyz = tt_sph_to_xyz(m31_distance_kpc,
m31_ra_rad, m31_dec_rad)
m31_galcen_xyz = np.dot(R_I2G, m31_icrs_xyz) + dxyz_I2G
m31_galcen_lon = np.arctan2(m31_galcen_xyz[1], m31_galcen_xyz[0])
m31_galcen_lat = np.arcsin(m31_galcen_xyz[2] / r)
xhat = m31_galcen_xyz / r
Rz = tt_rotation_matrix(-m31_galcen_lon, 'z')
Ry = tt_rotation_matrix(m31_galcen_lat, 'y')
Rx = tt_rotation_matrix(alpha, 'x')
yhat = np.dot(np.dot(Rz, np.dot(Ry, Rx)), [0, 1, 0.])
zhat = tt_cross(xhat, yhat)
R_LGtoG = np.stack((xhat, yhat, zhat), axis=1)
x_LG = np.array([r, 0., 0.])
v_LG = np.array([vrad, vtan, 0.])
x_I = np.dot(R_G2I, np.dot(R_LGtoG, x_LG)) + dxyz_G2I
v_I = np.dot(R_G2I, np.dot(R_LGtoG, v_LG)) + dvxyz_G2I
v_I_tangent_plane = np.dot(M, v_I) # alpha, delta, radial
shit1 = coord.CartesianRepresentation(*((R @ x_LG) * usys['length']))
shit2 = coord.CartesianDifferential(*((R @ v_LG) * usys['velocity']))
shit = coord.SkyCoord(shit1.with_differentials(shit2), frame=coord.Galactocentric())
return x_I, v_I, shit.transform_to(coord.ICRS()).velocity
```
## Write data to files:
```
rng = np.random.default_rng(seed=42)
dist_err = 11. * u.kpc
pmra_err = 3 * u.microarcsecond / u.yr
pmdec_err = 4 * u.microarcsecond / u.yr
rv_err = 2. * u.km/u.s
t_err = 0.11 * u.Gyr
tbl = {}
tbl['ra'] = u.Quantity(fake_icrs.ra)
tbl['dec'] = u.Quantity(fake_icrs.dec)
tbl['distance'] = rng.normal(fake_icrs.distance.to_value(u.kpc),
dist_err.to_value(u.kpc)) * u.kpc
tbl['distance_err'] = dist_err
tbl['pm_ra_cosdec'] = rng.normal(
fake_icrs.pm_ra_cosdec.to_value(pmra_err.unit),
pmra_err.value) * pmra_err.unit
tbl['pm_ra_cosdec_err'] = pmra_err
tbl['pm_dec'] = rng.normal(
fake_icrs.pm_dec.to_value(pmdec_err.unit),
pmdec_err.value) * pmdec_err.unit
tbl['pm_dec_err'] = pmdec_err
tbl['radial_velocity'] = rng.normal(
fake_icrs.radial_velocity.to_value(rv_err.unit),
rv_err.value) * rv_err.unit
tbl['radial_velocity_err'] = rv_err
tbl['tperi'] = rng.normal(
time.to_value(t_err.unit),
t_err.value) * t_err.unit
tbl['tperi_err'] = t_err
t = at.QTable({k: [] * tbl[k].unit for k in tbl})
t.add_row(tbl)
t.meta['title'] = 'Simulated Two-body'
t.write('../datasets/apw-simulated.ecsv', overwrite=True)
rng = np.random.default_rng(seed=42)
dist_err = 1. * u.kpc
pmra_err = 0.1 * u.microarcsecond / u.yr
pmdec_err = 0.1 * u.microarcsecond / u.yr
rv_err = 0.1 * u.km/u.s
t_err = 0.02 * u.Gyr
tbl = {}
tbl['ra'] = u.Quantity(fake_icrs.ra)
tbl['dec'] = u.Quantity(fake_icrs.dec)
tbl['distance'] = rng.normal(fake_icrs.distance.to_value(u.kpc),
dist_err.to_value(u.kpc)) * u.kpc
tbl['distance_err'] = dist_err
tbl['pm_ra_cosdec'] = rng.normal(
fake_icrs.pm_ra_cosdec.to_value(pmra_err.unit),
pmra_err.value) * pmra_err.unit
tbl['pm_ra_cosdec_err'] = pmra_err
tbl['pm_dec'] = rng.normal(
fake_icrs.pm_dec.to_value(pmdec_err.unit),
pmdec_err.value) * pmdec_err.unit
tbl['pm_dec_err'] = pmdec_err
tbl['radial_velocity'] = rng.normal(
fake_icrs.radial_velocity.to_value(rv_err.unit),
rv_err.value) * rv_err.unit
tbl['radial_velocity_err'] = rv_err
tbl['tperi'] = rng.normal(
time.to_value(t_err.unit),
t_err.value) * t_err.unit
tbl['tperi_err'] = t_err
t = at.QTable({k: [] * tbl[k].unit for k in tbl})
t.add_row(tbl)
t.meta['title'] = 'Simulated Two-body - precise'
t.write('../datasets/apw-simulated-precise.ecsv', overwrite=True)
rng = np.random.default_rng(42)
tbl = {}
vrad_err = 1 * u.km/u.s
vtan_err = 1 * u.km/u.s
t_err = 0.1 * u.Gyr
r_err = 1 * u.kpc
tbl['vrad'] = rng.normal(
vrad.to_value(vrad_err.unit),
vrad_err.value) * vrad_err.unit
tbl['vrad_err'] = vrad_err
tbl['vtan'] = rng.normal(
vtan.to_value(vtan_err.unit),
vtan_err.value) * vtan_err.unit
tbl['vtan_err'] = vtan_err
tbl['r'] = rng.normal(
r.to_value(r_err.unit),
r_err.value) * r_err.unit
tbl['r_err'] = r_err
tbl['tperi'] = rng.normal(
time.to_value(t_err.unit),
t_err.value) * t_err.unit
tbl['tperi_err'] = t_err
t = at.QTable({k: [] * tbl[k].unit for k in tbl})
t.add_row(tbl)
t.meta['title'] = 'Simulated Two-body - simple vrad, vtan'
t.write('../datasets/apw-simulated-simple.ecsv', overwrite=True)
```
| github_jupyter |
```
from torch import nn
from torchvision import models
# load model
model_resnet18 = models.resnet18(pretrained=False)
num_ftrs = model_resnet18.fc.in_features
# change last layer
num_classes=10
model_resnet18.fc = nn.Linear(num_ftrs, num_classes)
import torch
# load state_dict into model
path2weights="./models/resnet18_pretrained.pt"
model_resnet18.load_state_dict(torch.load(path2weights))
# set model in evaluation mode
model_resnet18.eval()
# move model to cuda/gpu device
if torch.cuda.is_available():
device = torch.device("cuda")
model_resnet18=model_resnet18.to(device)
def deploy_model(model,dataset,device, num_classes=10,sanity_check=False):
len_data=len(dataset)
# initialize output tensor on CPU: due to GPU memory limits
y_out=torch.zeros(len_data,num_classes)
# initialize ground truth on CPU: due to GPU memory limits
y_gt=np.zeros((len_data),dtype="uint8")
# move model to device
model=model.to(device)
elapsed_times=[]
with torch.no_grad():
for i in range(len_data):
x,y=dataset[i]
y_gt[i]=y
start=time.time()
yy=model(x.unsqueeze(0).to(device))
y_out[i]=torch.softmax(yy,dim=1)
elapsed=time.time()-start
elapsed_times.append(elapsed)
if sanity_check is True:
break
inference_time=np.mean(elapsed_times)*1000
print("average inference time per image on %s: %.2f ms " %(device,inference_time))
return y_out.numpy(),y_gt
```
## Loading Test Dataset
```
from torchvision import datasets
import torchvision.transforms as transforms
# define transformation
data_transformer = transforms.Compose([transforms.ToTensor()])
path2data="./data"
# loading data
test0_ds=datasets.STL10(path2data, split='test', download=True,transform=data_transformer)
print(test0_ds.data.shape)
from sklearn.model_selection import StratifiedShuffleSplit
sss = StratifiedShuffleSplit(n_splits=1, test_size=0.2, random_state=0)
indices=list(range(len(test0_ds)))
y_test0=[y for _,y in test0_ds]
for test_index, val_index in sss.split(indices, y_test0):
print("test:", test_index, "val:", val_index)
print(len(val_index),len(test_index))
from torch.utils.data import Subset
val_ds=Subset(test0_ds,val_index)
test_ds=Subset(test0_ds,test_index)
mean=[0.4467106, 0.43980986, 0.40664646]
std=[0.22414584,0.22148906,0.22389975]
test0_transformer = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize(mean, std),
])
test0_ds.transform=test0_transformer
import time
import numpy as np
# deploy model
y_out,y_gt=deploy_model(model_resnet18,val_ds,device=device,sanity_check=False)
print(y_out.shape,y_gt.shape)
from sklearn.metrics import accuracy_score
# get predictions
y_pred = np.argmax(y_out,axis=1)
print(y_pred.shape,y_gt.shape)
# compute accuracy
acc=accuracy_score(y_pred,y_gt)
print("accuracy: %.2f" %acc)
y_out,y_gt=deploy_model(model_resnet18,test_ds,device=device)
y_pred = np.argmax(y_out,axis=1)
acc=accuracy_score(y_pred,y_gt)
print(acc)
from torchvision import utils
import matplotlib.pyplot as plt
import numpy as np
%matplotlib inline
np.random.seed(1)
def imshow(inp, title=None):
mean=[0.4467106, 0.43980986, 0.40664646]
std=[0.22414584,0.22148906,0.22389975]
inp = inp.numpy().transpose((1, 2, 0))
mean = np.array(mean)
std = np.array(std)
inp = std * inp + mean
inp = np.clip(inp, 0, 1)
plt.imshow(inp)
if title is not None:
plt.title(title)
plt.pause(0.001) # pause a bit so that plots are updated
grid_size=4
rnd_inds=np.random.randint(0,len(test_ds),grid_size)
print("image indices:",rnd_inds)
x_grid_test=[test_ds[i][0] for i in rnd_inds]
y_grid_test=[(y_pred[i],y_gt[i]) for i in rnd_inds]
x_grid_test=utils.make_grid(x_grid_test, nrow=4, padding=2)
print(x_grid_test.shape)
plt.rcParams['figure.figsize'] = (10, 5)
imshow(x_grid_test,y_grid_test)
device_cpu = torch.device("cpu")
y_out,y_gt=deploy_model(model_resnet18,val_ds,device=device_cpu,sanity_check=False)
print(y_out.shape,y_gt.shape)
```
| github_jupyter |
# Fusing graphblas.matrix_multiply with graphblas.matrix_apply
This example will go over how to use the `--graphblas-optimize` pass from `graphblas-opt` to fuse `graphblas.matrix_multiply` ops with `graphblas.matrix_apply` ops into `graphblas.matrix_multiply` ops with a region attached.
Let's first import some necessary libraries.
```
import tempfile
from mlir_graphblas.cli import GRAPHBLAS_OPT_EXE
```
Since [sparse tensor encodings](https://mlir.llvm.org/docs/Dialects/SparseTensorOps/#sparsetensorencodingattr) can be very verbose in MLIR, let's write some helpers to make the MLIR code more readable.
```
def tersify_mlir(input_string: str) -> str:
terse_string = input_string
terse_string = terse_string.replace(
'''#sparse_tensor.encoding<{ '''
'''dimLevelType = [ "dense", "compressed" ], '''
'''dimOrdering = affine_map<(d0, d1) -> (d0, d1)>, '''
'''pointerBitWidth = 64, '''
'''indexBitWidth = 64 '''
'''}>''',
"#CSR64")
terse_string = terse_string.replace(
'''#sparse_tensor.encoding<{ '''
'''dimLevelType = [ "dense", "compressed" ], '''
'''dimOrdering = affine_map<(d0, d1) -> (d1, d0)>, '''
'''pointerBitWidth = 64, '''
'''indexBitWidth = 64 '''
'''}>''',
"#CSC64")
return terse_string
```
## Fusion Details
Recall that `graphblas.matrix_multiply` can take an optional region, e.g. this code squares each element of the matrix multiply product:
```
%answer = graphblas.matrix_multiply %argA, %argB { semiring = "plus_times" } : (tensor<?x?xf64, #CSR64>, tensor<?x?xf64, #CSC64>) to tensor<?x?xf64, #CSR64> {
^bb0(%value: f64):
%result = std.mulf %value, %value: f64
graphblas.yield %result : f64
}
```
Since `graphblas.matrix_apply` ops only change tensors in an element-wise fashion, we can perform these element-wise changes in the region of a `graphblas.matrix_multiply` op if the `graphblas.matrix_apply` op is run on the result of a `graphblas.matrix_multiply` op.
## Simple Fusion
Here, we'll show the simplest example of how we can fuse a `graphblas.matrix_multiply` op with a `graphblas.matrix_apply` op.
```
mlir_text = """
#CSR64 = #sparse_tensor.encoding<{
dimLevelType = [ "dense", "compressed" ],
dimOrdering = affine_map<(i,j) -> (i,j)>,
pointerBitWidth = 64,
indexBitWidth = 64
}>
#CSC64 = #sparse_tensor.encoding<{
dimLevelType = [ "dense", "compressed" ],
dimOrdering = affine_map<(i,j) -> (j,i)>,
pointerBitWidth = 64,
indexBitWidth = 64
}>
func @fuse_adjacent(%A: tensor<?x?xf64, #CSR64>, %B: tensor<?x?xf64, #CSC64>, %thunk: f64) -> tensor<?x?xf64, #CSR64> {
%C = graphblas.matrix_multiply %A, %B { semiring = "plus_plus" } : (tensor<?x?xf64, #CSR64>, tensor<?x?xf64, #CSC64>) to tensor<?x?xf64, #CSR64>
%apply_result = graphblas.matrix_apply %C, %thunk { apply_operator = "min" } : (tensor<?x?xf64, #CSR64>, f64) to tensor<?x?xf64, #CSR64>
return %apply_result : tensor<?x?xf64, #CSR64>
}
"""
with tempfile.NamedTemporaryFile() as temp:
temp_file_name = temp.name
with open(temp_file_name, 'w') as f:
f.write(mlir_text)
temp.flush()
output_mlir = ! cat $temp_file_name | $GRAPHBLAS_OPT_EXE --graphblas-optimize
output_mlir = "\n".join(output_mlir)
output_mlir = tersify_mlir(output_mlir)
print(output_mlir)
```
The code in the region attached to the `graphblas.matrix_multiply` in the lowered MLIR here may seem confusing at first, but it's simply calculating the minimum of each element (i.e. `%arg3`) and the thunk (i.e. `%thunk` or `%arg2`).
It's noteworthy that this fusion also works if the `graphblas.matrix_multiply` use takes a mask. Rather than explicitly demonstrating this, we'll leave it as an exercise for the reader as it's a fairly straightforward.
Similar to our previous `graphblas.matrix_multiply_reduce_to_scalar` examples, if the intermediate result from the `graphblas.matrix_multiply` op is used in other places outside of the `graphblas.matrix_apply` op, this fusion cannot apply.
| github_jupyter |
Peakcalling Bam Stats and Filtering Report - Insert Sizes
================================================================
This notebook is for the analysis of outputs from the peakcalling pipeline
There are severals stats that you want collected and graphed (topics covered in this notebook in bold).
These are:
- how many reads input
- how many reads removed at each step (numbers and percentages)
- how many reads left after filtering
- inset size distribution pre filtering for PE reads
- how many reads mapping to each chromosome before filtering?
- how many reads mapping to each chromosome after filtering?
- X:Y reads ratio
- **inset size distribution after filtering for PE reads**
- samtools flags - check how many reads are in categories they shouldn't be
- picard stats - check how many reads are in categories they shouldn't be
This notebook takes the sqlite3 database created by cgat peakcalling_pipeline.py and uses it for plotting the above statistics
It assumes a file directory of:
location of database = project_folder/csvdb
location of this notebook = project_folder/notebooks.dir/
Firstly lets load all the things that might be needed
Insert size distribution
------------------------
This section get the size distribution of the fragements that have been sequeced in paired-end sequencing. The pipeline calculates the size distribution by caluculating the distance between the most 5' possition of both reads, for those mapping to the + stand this is the left-post possition, for those mapping to the - strand is the rightmost coordinate.
This plot is especially useful for ATAC-Seq experiments as good samples should show peaks with a period approximately equivelent to the length of a nucleosome (~ 146bp) a lack of this phasing might indicate poor quality samples and either over (if lots of small fragments) or under intergration (if an excess of large fragments) of the topoisomerase.
```
import sqlite3
import pandas as pd
import numpy as np
%matplotlib inline
import matplotlib
import numpy as np
import matplotlib.pyplot as plt
#import cgatcore.pipeline as P
import os
import statistics
#import collections
#load R and the R packages required
#%load_ext rpy2.ipython
#%R require(ggplot2)
# use these functions to display tables nicely as html
from IPython.display import display, HTML
plt.style.use('ggplot')
#plt.style.available
```
This is where we are and when the notebook was run
```
!pwd
!date
```
First lets set the output path for where we want our plots to be saved and the database path and see what tables it contains
```
database_path = '../csvdb'
output_path = '.'
#database_path= "/ifs/projects/charlotteg/pipeline_peakcalling/csvdb"
```
This code adds a button to see/hide code in html
```
HTML('''<script>
code_show=true;
function code_toggle() {
if (code_show){
$('div.input').hide();
} else {
$('div.input').show();
}
code_show = !code_show
}
$( document ).ready(code_toggle);
</script>
<form action="javascript:code_toggle()"><input type="submit" value="Click here to toggle on/off the raw code."></form>''')
```
The code below provides functions for accessing the project database and extract a table names so you can see what tables have been loaded into the database and are available for plotting. It also has a function for geting table from the database and indexing the table with the track name
```
def getTableNamesFromDB(database_path):
# Create a SQL connection to our SQLite database
con = sqlite3.connect(database_path)
cur = con.cursor()
# the result of a "cursor.execute" can be iterated over by row
cur.execute("SELECT name FROM sqlite_master WHERE type='table' ORDER BY name;")
available_tables = (cur.fetchall())
#Be sure to close the connection.
con.close()
return available_tables
db_tables = getTableNamesFromDB(database_path)
print('Tables contained by the database:')
for x in db_tables:
print('\t\t%s' % x[0])
#This function retrieves a table from sql database and indexes it with track name
def getTableFromDB(statement,database_path):
'''gets table from sql database depending on statement
and set track as index if contains track in column names'''
conn = sqlite3.connect(database_path)
df = pd.read_sql_query(statement,conn)
if 'track' in df.columns:
df.index = df['track']
return df
```
Insert Size Summary
====================
1) lets getthe insert_sizes table from database
Firsly lets look at the summary statistics that us the mean fragment size, sequencing type and mean read length. This table is produced using macs2 for PE data, or bamtools for SE data
If IDR has been run the insert_size table will contain entries for the pooled and pseudo replicates too - we don't really want this as it will duplicate the data from the origional samples so we subset this out
```
insert_df = getTableFromDB('select * from insert_sizes;',database_path)
insert_df = insert_df[insert_df["filename"].str.contains('pseudo')==False].copy()
insert_df = insert_df[insert_df["filename"].str.contains('pooled')==False].copy()
def add_expt_to_insertdf(dataframe):
''' splits track name for example HsTh1-RATotal-R1.star into expt
featues, expt, sample_treatment and replicate and adds these as
collumns to the dataframe'''
expt = []
treatment = []
replicate = []
for value in dataframe.filename:
x = value.split('/')[-1]
x = x.split('_insert')[0]
# split into design features
y = x.split('-')
expt.append(y[-3])
treatment.append(y[-2])
replicate.append(y[-1])
if len(expt) == len(treatment) and len(expt)== len(replicate):
print ('all values in list correctly')
else:
print ('error in loading values into lists')
#add collums to dataframe
dataframe['expt_name'] = expt
dataframe['sample_treatment'] = treatment
dataframe['replicate'] = replicate
return dataframe
insert_df = add_expt_to_insertdf(insert_df)
insert_df
```
lets graph the fragment length mean and tag size grouped by sample so we can see if they are much different
```
ax = insert_df.boxplot(column='fragmentsize_mean', by='sample_treatment')
ax.set_title('for mean fragment size',size=10)
ax.set_ylabel('mean fragment length')
ax.set_xlabel('sample treatment')
ax = insert_df.boxplot(column='tagsize', by='sample_treatment')
ax.set_title('for tag size',size=10)
ax.set_ylabel('tag size')
ax.set_xlabel('sample treatment')
ax.set_ylim(((insert_df.tagsize.min()-2),(insert_df.tagsize.max()+2)))
```
Ok now get get the fragment length distributiions for each sample and plot them
```
def getFraglengthTables(database_path):
'''Takes path to sqlite3 database and retrieves fraglengths tables for individual samples
, returns a dictionary where keys = sample table names, values = fraglengths dataframe'''
frag_tabs = []
db_tables = getTableNamesFromDB(database_path)
for table_name in db_tables:
if 'fraglengths' in str(table_name[0]):
tab_name = str(table_name[0])
statement ='select * from %s;' % tab_name
df = getTableFromDB(statement,database_path)
frag_tabs.append((tab_name,df))
print('detected fragment length distribution tables for %s files: \n' % len(frag_tabs))
for val in frag_tabs:
print(val[0])
return frag_tabs
def getDFofFragLengths(database_path):
''' this takes a path to database and gets a dataframe where length of fragments is the index,
each column is a sample and values are the number of reads that have that fragment length in that
sample
'''
fraglength_dfs_list = getFraglengthTables(database_path)
dfs=[]
for item in fraglength_dfs_list:
track = item[0].split('_filtered_fraglengths')[0]
df = item[1]
#rename collumns so that they are correct - correct this in the pipeline then delete this
#df.rename(columns={'frequency':'frag_length', 'frag_length':'frequency'}, inplace=True)
df.index = df.frag_length
df.drop('frag_length',axis=1,inplace=True)
df.rename(columns={'frequency':track},inplace=True)
dfs.append(df)
frag_length_df = pd.concat(dfs,axis=1)
frag_length_df.fillna(0, inplace=True)
return frag_length_df
#Note the frequency and fragment lengths are around the wrong way!
#frequency is actually fragment length, and fragement length is the frequency
#This gets the tables from db and makes master df of all fragment length frequencies
frag_length_df = getDFofFragLengths(database_path)
#plot fragment length frequencies
ax = frag_length_df.divide(1000).plot()
ax.set_ylabel('Number of fragments\n(thousands)')
ax.legend(loc=2,bbox_to_anchor=(1.05, 1),borderaxespad=0. )
ax.set_title('fragment length distribution')
ax.set_xlabel('fragment length (bp)')
ax.set_xlim()
```
Now lets zoom in on the interesting region of the plot (the default in the code looks at fragment lengths from 0 to 800bp - you can change this below by setting the tuple in the ax.set_xlim() function
```
ax = frag_length_df.divide(1000).plot(figsize=(9,9))
ax.set_ylabel('Number of fragments\n(thousands)')
ax.legend(loc=2,bbox_to_anchor=(1.05, 1),borderaxespad=0. )
ax.set_title('fragment length distribution')
ax.set_xlabel('fragment length (bp)')
ax.set_xlim((0,800))
```
it is a bit trickly to see differences between samples of different library sizes so lets look and see if the reads for each fragment length is similar
```
percent_frag_length_df = pd.DataFrame(index=frag_length_df.index)
for column in frag_length_df:
total_frags = frag_length_df[column].sum()
percent_frag_length_df[column] = frag_length_df[column].divide(total_frags)*100
ax = percent_frag_length_df.plot(figsize=(9,9))
ax.set_ylabel('Percentage of fragments')
ax.legend(loc=2,bbox_to_anchor=(1.05, 1),borderaxespad=0. )
ax.set_title('percentage fragment length distribution')
ax.set_xlabel('fragment length (bp)')
ax.set_xlim((0,800))
```
SUMMARISE HERE
==============
From these plots you should be able to tell wether there are any distinctive patterns in the size of the fragment lengths,this is especially important for ATAC-Seq data as in successful experiments you should be able to detect nucleosome phasing - it can also indicate over fragmentation or biases in cutting.
Lets looks at the picard insert size metrics also
```
insert_df = getTableFromDB('select * from picard_stats_insert_size_metrics;',database_path)
for c in insert_df.columns:
print (c)
insert_df
```
These metrics are actually quite different to the ones we calculate themselves - for some reason it seems to split the files into 2 and dives a distribution for smaller fragments and for larger fragments- not sure why at the moment
| github_jupyter |
```
import keras
from keras.models import Sequential, Model, load_model
from keras.layers import Dense, Dropout, Flatten, Input, Lambda, Concatenate
from keras.layers import Conv1D, MaxPooling1D
from keras.callbacks import ModelCheckpoint, EarlyStopping
from keras import backend as K
import keras.losses
import tensorflow as tf
import pandas as pd
import os
import numpy as np
import scipy.sparse as sp
import scipy.io as spio
import matplotlib.pyplot as plt
import matplotlib.cm as cm
import isolearn.io as isoio
import isolearn.keras as iso
from scipy.stats import pearsonr
```
<h2>Load 5' Alternative Splicing Data</h2>
- Load a Pandas DataFrame + Matlab Matrix of measured Splicing Sequences<br/>
- isolearn.io loads all .csv and .mat files of a directory into memory as a dictionary<br/>
- The DataFrame has one column - padded_sequence - containing the splice donor sequence<br/>
- The Matrix contains RNA-Seq counts of measured splicing at each position across the sequence<br/>
```
#Load Splicing Data
splicing_dict = isoio.load('data/processed_data/splicing_5ss_data/splicing_5ss_data')
```
<h2>Create a Training and Test Set</h2>
- We create an index containing row numbers corresponding to training and test sequences<br/>
- Notice that we do not alter the underlying DataFrame, we only make lists of pointers to rows<br/>
```
#Generate training, validation and test set indexes
valid_set_size = 0.10
test_set_size = 0.10
data_index = np.arange(len(splicing_dict['df']), dtype=np.int)
train_index = data_index[:-int(len(data_index) * (valid_set_size + test_set_size))]
valid_index = data_index[train_index.shape[0]:-int(len(data_index) * test_set_size)]
test_index = data_index[train_index.shape[0] + valid_index.shape[0]:]
print('Training set size = ' + str(train_index.shape[0]))
print('Validation set size = ' + str(valid_index.shape[0]))
print('Test set size = ' + str(test_index.shape[0]))
```
<h2>Create Data Generators</h2>
- In Isolearn, we always build data generators that will encode and feed us the data on the fly<br/>
- Here, for example, we create a training and test generator separately (using list comprehension)<br/>
- First argument: The list of row indices (of data points) for this generator<br/>
- Second argument: Dictionary or data sources<br/>
- Third argument: Batch size for the data generator
- Fourth argument: List of inputs, where each input is specified as a dictionary of attributes<br/>
- Fifth argument: List of outputs<br/>
- Sixth argument: List of any randomizers (see description below)<br/>
- Seventh argument: Shuffle the dataset or not<br/>
- Eight argument: True if some data source matrices are in sparse format<br/>
- Ninth argument: In Keras, we typically want to specfiy the Outputs as Inputs when training. <br/>This argument achieves this by moving the outputs over to the input list and replaces the output with a dummy encoder.<br/>
In this example, we specify a One-Hot encoder as the input encoder for the entire splice donor sequence (centered on the splice donor).<br/>
We also specify the target output as the normalized RNA-Seq count at position 120 in the count matrix for each cell line (4 outputs).<br/>
Besides the canonical splice donor at position 120 in the sequence, there are many other splice donors inserted randomly at neighboring positions. If we wanted to learn a general model of splicing, it would be a lot better if we could stochastically "align" sequences on any of the possible splice donors, perturbing both the input sequence and the RNA-Seq count matrix that we estimate splice donor usage from.<br/>
This is achieved using the built-in CutAlignSampler class, which allows us to randomly sample a position in the sequence with supporting splice junction counts, and shift both the sequence and splice count vector to be centered around that position. In this example, we specfiy the sampling rate of splice donors to be 0.5 (p_pos) and the rate of sampling some other, non-splice-site, position at a rate of 0.5 (p_neg).<br/>
```
#Create a One-Hot data generator, to be used for a convolutional net to regress SD1 Usage
total_cuts = splicing_dict['hek_count'] + splicing_dict['hela_count'] + splicing_dict['mcf7_count'] + splicing_dict['cho_count']
shifter = iso.CutAlignSampler(total_cuts, 240, 120, [], 0.0, p_pos=0.5, p_neg=0.5, sparse_source=True)
splicing_gens = {
gen_id : iso.DataGenerator(
idx,
{
'df' : splicing_dict['df'],
'hek_count' : splicing_dict['hek_count'],
'hela_count' : splicing_dict['hela_count'],
'mcf7_count' : splicing_dict['mcf7_count'],
'cho_count' : splicing_dict['cho_count'],
},
batch_size=32,
inputs = [
{
'id' : 'seq',
'source_type' : 'dataframe',
'source' : 'df',
'extractor' : iso.SequenceExtractor('padded_sequence', start_pos=0, end_pos=240, shifter=shifter if gen_id == 'train' else None),
'encoder' : iso.OneHotEncoder(seq_length=240),
'dim' : (240, 4),
'sparsify' : False
}
],
outputs = [
{
'id' : cell_type + '_sd1_usage',
'source_type' : 'matrix',
'source' : cell_type + '_count',
'extractor' : iso.CountExtractor(start_pos=0, end_pos=240, static_poses=[-1], shifter=shifter if gen_id == 'train' else None, sparse_source=False),
'transformer' : lambda t: t[120] / np.sum(t)
} for cell_type in ['hek', 'hela', 'mcf7', 'cho']
],
randomizers = [shifter] if gen_id in ['train'] else [],
shuffle = True if gen_id in ['train'] else False,
densify_batch_matrices=True,
move_outputs_to_inputs=True if gen_id in ['train', 'valid'] else False
) for gen_id, idx in [('train', train_index), ('valid', valid_index), ('test', test_index)]
}
```
<h2>Keras Loss Functions</h2>
Here we specfiy a few loss function (Cross-Entropy and KL-divergence) to be used when optimizing our Splicing CNN.<br/>
```
#Keras loss functions
def sigmoid_entropy(inputs) :
y_true, y_pred = inputs
y_pred = K.clip(y_pred, K.epsilon(), 1. - K.epsilon())
return -K.sum(y_true * K.log(y_pred) + (1.0 - y_true) * K.log(1.0 - y_pred), axis=-1)
def mean_sigmoid_entropy(inputs) :
y_true, y_pred = inputs
y_pred = K.clip(y_pred, K.epsilon(), 1. - K.epsilon())
return -K.mean(y_true * K.log(y_pred) + (1.0 - y_true) * K.log(1.0 - y_pred), axis=-1)
def sigmoid_kl_divergence(inputs) :
y_true, y_pred = inputs
y_pred = K.clip(y_pred, K.epsilon(), 1. - K.epsilon())
y_true = K.clip(y_true, K.epsilon(), 1. - K.epsilon())
return K.sum(y_true * K.log(y_true / y_pred) + (1.0 - y_true) * K.log((1.0 - y_true) / (1.0 - y_pred)), axis=-1)
def mean_sigmoid_kl_divergence(inputs) :
y_true, y_pred = inputs
y_pred = K.clip(y_pred, K.epsilon(), 1. - K.epsilon())
y_true = K.clip(y_true, K.epsilon(), 1. - K.epsilon())
return K.mean(y_true * K.log(y_true / y_pred) + (1.0 - y_true) * K.log((1.0 - y_true) / (1.0 - y_pred)), axis=-1)
```
<h2>Splicing Model Definition</h2>
Here we specfiy the Keras Inputs that we expect to receive from the data generators.<br/>
We also define the model architecture (2 convolutional-layer CNN with MaxPooling).<br/>
```
#Splicing Model Definition (CNN)
#Inputs
seq_input = Input(shape=(240, 4))
#Outputs
true_usage_hek = Input(shape=(1,))
true_usage_hela = Input(shape=(1,))
true_usage_mcf7 = Input(shape=(1,))
true_usage_cho = Input(shape=(1,))
#Shared Model Definition (Applied to each randomized sequence region)
layer_1 = Conv1D(64, 8, padding='valid', activation='relu')
layer_1_pool = MaxPooling1D(pool_size=2)
layer_2 = Conv1D(128, 6, padding='valid', activation='relu')
def shared_model(seq_input) :
return Flatten()(
layer_2(
layer_1_pool(
layer_1(
seq_input
)
)
)
)
shared_out = shared_model(seq_input)
#Layers applied to the concatenated hidden representation
layer_dense = Dense(256, activation='relu')
layer_drop = Dropout(0.2)
dropped_dense_out = layer_drop(layer_dense(shared_out))
#Final cell-line specific regression layers
layer_usage_hek = Dense(1, activation='sigmoid', kernel_initializer='zeros')
layer_usage_hela = Dense(1, activation='sigmoid', kernel_initializer='zeros')
layer_usage_mcf7 = Dense(1, activation='sigmoid', kernel_initializer='zeros')
layer_usage_cho = Dense(1, activation='sigmoid', kernel_initializer='zeros')
pred_usage_hek = layer_usage_hek(dropped_dense_out)
pred_usage_hela = layer_usage_hela(dropped_dense_out)
pred_usage_mcf7 = layer_usage_mcf7(dropped_dense_out)
pred_usage_cho = layer_usage_cho(dropped_dense_out)
#Compile Splicing Model
splicing_model = Model(
inputs=[
seq_input
],
outputs=[
pred_usage_hek,
pred_usage_hela,
pred_usage_mcf7,
pred_usage_cho
]
)
```
<h2>Loss Model Definition</h2>
Here we specfiy our loss function, and we build it as a separate Keras Model.<br/>
In our case, our loss model averages the KL-divergence of predicted vs. true Splice Donor Usage across the 4 different cell types.<br/>
```
#Loss Model Definition
loss_hek = Lambda(sigmoid_kl_divergence, output_shape = (1,))([true_usage_hek, pred_usage_hek])
loss_hela = Lambda(sigmoid_kl_divergence, output_shape = (1,))([true_usage_hela, pred_usage_hela])
loss_mcf7 = Lambda(sigmoid_kl_divergence, output_shape = (1,))([true_usage_mcf7, pred_usage_mcf7])
loss_cho = Lambda(sigmoid_kl_divergence, output_shape = (1,))([true_usage_cho, pred_usage_cho])
total_loss = Lambda(
lambda l: (l[0] + l[1] + l[2] + l[3]) / 4.,
output_shape = (1,)
)(
[
loss_hek,
loss_hela,
loss_mcf7,
loss_cho
]
)
loss_model = Model([
#Inputs
seq_input,
#Target SD Usages
true_usage_hek,
true_usage_hela,
true_usage_mcf7,
true_usage_cho
], total_loss)
```
<h2>Optimize the Loss Model</h2>
Here we use SGD to optimize the Loss Model (defined in the previous notebook cell).<br/>
Since our Loss Model indirectly depends on predicted outputs from our CNN Splicing Model, SGD will optimize the weights of our CNN<br/>
<br/>
Note that we very easily pass the data generators, and run them in parallel, by simply calling Keras fit_generator.<br/>
```
#Optimize CNN with Keras using the Data Generators to stream genomic data features
opt = keras.optimizers.SGD(lr=0.1, decay=1e-6, momentum=0.9, nesterov=True)
loss_model.compile(loss=lambda true, pred: pred, optimizer=opt)
callbacks =[
EarlyStopping(monitor='val_loss', min_delta=0.001, patience=2, verbose=0, mode='auto')
]
loss_model.fit_generator(
generator=splicing_gens['train'],
validation_data=splicing_gens['valid'],
epochs=10,
use_multiprocessing=True,
workers=4,
callbacks=callbacks
)
#Save model
save_dir = os.path.join(os.getcwd(), 'saved_models')
if not os.path.isdir(save_dir):
os.makedirs(save_dir)
model_name = 'splicing_cnn_perturbed_multicell.h5'
model_path = os.path.join(save_dir, model_name)
splicing_model.save(model_path)
print('Saved trained model at %s ' % model_path)
#Load model
save_dir = os.path.join(os.getcwd(), 'saved_models')
model_name = 'splicing_cnn_perturbed_multicell.h5'
model_path = os.path.join(save_dir, model_name)
splicing_model = load_model(model_path)
```
<h2>Evaluate the Splicing CNN</h2>
Here we run our Splicing CNN on the Test set data generator (using Keras predict_generator).<br/>
We then compare our predictions of splice donor usage against the true RNA-Seq measurements.<br/>
```
#Evaluate predictions on test set
predictions = splicing_model.predict_generator(splicing_gens['test'], workers=4, use_multiprocessing=True)
pred_usage_hek, pred_usage_hela, pred_usage_mcf7, pred_usage_cho = [np.ravel(prediction) for prediction in predictions]
targets = zip(*[splicing_gens['test'][i][1] for i in range(len(splicing_gens['test']))])
true_usage_hek, true_usage_hela, true_usage_mcf7, true_usage_cho = [np.concatenate(list(target)) for target in targets]
cell_lines = [
('hek', (pred_usage_hek, true_usage_hek)),
('hela', (pred_usage_hela, true_usage_hela)),
('mcf7', (pred_usage_mcf7, true_usage_mcf7)),
('cho', (pred_usage_cho, true_usage_cho))
]
for cell_name, [y_true, y_pred] in cell_lines :
r_val, p_val = pearsonr(y_pred, y_true)
print("Test set R^2 = " + str(round(r_val * r_val, 2)) + ", p = " + str(p_val))
#Plot test set scatter
f = plt.figure(figsize=(4, 4))
plt.scatter(y_pred, y_true, color='black', s=5, alpha=0.05)
plt.xticks([0.0, 0.25, 0.5, 0.75, 1.0], fontsize=14)
plt.yticks([0.0, 0.25, 0.5, 0.75, 1.0], fontsize=14)
plt.xlabel('Predicted SD1 Usage', fontsize=14)
plt.ylabel('True SD1 Usage', fontsize=14)
plt.title(str(cell_name), fontsize=16)
plt.xlim(0, 1)
plt.ylim(0, 1)
plt.tight_layout()
plt.show()
```
| github_jupyter |
# Computer Vision Nanodegree
## Project: Image Captioning
---
In this notebook, you will learn how to load and pre-process data from the [COCO dataset](http://cocodataset.org/#home). You will also design a CNN-RNN model for automatically generating image captions.
Note that **any amendments that you make to this notebook will not be graded**. However, you will use the instructions provided in **Step 3** and **Step 4** to implement your own CNN encoder and RNN decoder by making amendments to the **models.py** file provided as part of this project. Your **models.py** file **will be graded**.
Feel free to use the links below to navigate the notebook:
- [Step 1](#step1): Explore the Data Loader
- [Step 2](#step2): Use the Data Loader to Obtain Batches
- [Step 3](#step3): Experiment with the CNN Encoder
- [Step 4](#step4): Implement the RNN Decoder
<a id='step1'></a>
## Step 1: Explore the Data Loader
We have already written a [data loader](http://pytorch.org/docs/master/data.html#torch.utils.data.DataLoader) that you can use to load the COCO dataset in batches.
In the code cell below, you will initialize the data loader by using the `get_loader` function in **data_loader.py**.
> For this project, you are not permitted to change the **data_loader.py** file, which must be used as-is.
The `get_loader` function takes as input a number of arguments that can be explored in **data_loader.py**. Take the time to explore these arguments now by opening **data_loader.py** in a new window. Most of the arguments must be left at their default values, and you are only allowed to amend the values of the arguments below:
1. **`transform`** - an [image transform](http://pytorch.org/docs/master/torchvision/transforms.html) specifying how to pre-process the images and convert them to PyTorch tensors before using them as input to the CNN encoder. For now, you are encouraged to keep the transform as provided in `transform_train`. You will have the opportunity later to choose your own image transform to pre-process the COCO images.
2. **`mode`** - one of `'train'` (loads the training data in batches) or `'test'` (for the test data). We will say that the data loader is in training or test mode, respectively. While following the instructions in this notebook, please keep the data loader in training mode by setting `mode='train'`.
3. **`batch_size`** - determines the batch size. When training the model, this is number of image-caption pairs used to amend the model weights in each training step.
4. **`vocab_threshold`** - the total number of times that a word must appear in the in the training captions before it is used as part of the vocabulary. Words that have fewer than `vocab_threshold` occurrences in the training captions are considered unknown words.
5. **`vocab_from_file`** - a Boolean that decides whether to load the vocabulary from file.
We will describe the `vocab_threshold` and `vocab_from_file` arguments in more detail soon. For now, run the code cell below. Be patient - it may take a couple of minutes to run!
```
import sys
sys.path.append('/opt/cocoapi/PythonAPI')
from pycocotools.coco import COCO
!pip install nltk
import nltk
nltk.download('punkt')
from data_loader import get_loader
from torchvision import transforms
# Define a transform to pre-process the training images.
transform_train = transforms.Compose([
transforms.Resize(256), # smaller edge of image resized to 256
transforms.RandomCrop(224), # get 224x224 crop from random location
transforms.RandomHorizontalFlip(), # horizontally flip image with probability=0.5
transforms.ToTensor(), # convert the PIL Image to a tensor
transforms.Normalize((0.485, 0.456, 0.406), # normalize image for pre-trained model
(0.229, 0.224, 0.225))])
# Set the minimum word count threshold.
vocab_threshold = 5
# Specify the batch size.
batch_size = 10
# Obtain the data loader.
data_loader = get_loader(transform=transform_train,
mode='train',
batch_size=batch_size,
vocab_threshold=vocab_threshold,
vocab_from_file=False)
```
When you ran the code cell above, the data loader was stored in the variable `data_loader`.
You can access the corresponding dataset as `data_loader.dataset`. This dataset is an instance of the `CoCoDataset` class in **data_loader.py**. If you are unfamiliar with data loaders and datasets, you are encouraged to review [this PyTorch tutorial](http://pytorch.org/tutorials/beginner/data_loading_tutorial.html).
### Exploring the `__getitem__` Method
The `__getitem__` method in the `CoCoDataset` class determines how an image-caption pair is pre-processed before being incorporated into a batch. This is true for all `Dataset` classes in PyTorch; if this is unfamiliar to you, please review [the tutorial linked above](http://pytorch.org/tutorials/beginner/data_loading_tutorial.html).
When the data loader is in training mode, this method begins by first obtaining the filename (`path`) of a training image and its corresponding caption (`caption`).
#### Image Pre-Processing
Image pre-processing is relatively straightforward (from the `__getitem__` method in the `CoCoDataset` class):
```python
# Convert image to tensor and pre-process using transform
image = Image.open(os.path.join(self.img_folder, path)).convert('RGB')
image = self.transform(image)
```
After loading the image in the training folder with name `path`, the image is pre-processed using the same transform (`transform_train`) that was supplied when instantiating the data loader.
#### Caption Pre-Processing
The captions also need to be pre-processed and prepped for training. In this example, for generating captions, we are aiming to create a model that predicts the next token of a sentence from previous tokens, so we turn the caption associated with any image into a list of tokenized words, before casting it to a PyTorch tensor that we can use to train the network.
To understand in more detail how COCO captions are pre-processed, we'll first need to take a look at the `vocab` instance variable of the `CoCoDataset` class. The code snippet below is pulled from the `__init__` method of the `CoCoDataset` class:
```python
def __init__(self, transform, mode, batch_size, vocab_threshold, vocab_file, start_word,
end_word, unk_word, annotations_file, vocab_from_file, img_folder):
...
self.vocab = Vocabulary(vocab_threshold, vocab_file, start_word,
end_word, unk_word, annotations_file, vocab_from_file)
...
```
From the code snippet above, you can see that `data_loader.dataset.vocab` is an instance of the `Vocabulary` class from **vocabulary.py**. Take the time now to verify this for yourself by looking at the full code in **data_loader.py**.
We use this instance to pre-process the COCO captions (from the `__getitem__` method in the `CoCoDataset` class):
```python
# Convert caption to tensor of word ids.
tokens = nltk.tokenize.word_tokenize(str(caption).lower()) # line 1
caption = [] # line 2
caption.append(self.vocab(self.vocab.start_word)) # line 3
caption.extend([self.vocab(token) for token in tokens]) # line 4
caption.append(self.vocab(self.vocab.end_word)) # line 5
caption = torch.Tensor(caption).long() # line 6
```
As you will see soon, this code converts any string-valued caption to a list of integers, before casting it to a PyTorch tensor. To see how this code works, we'll apply it to the sample caption in the next code cell.
```
sample_caption = 'A person doing a trick on a rail while riding a skateboard.'
```
In **`line 1`** of the code snippet, every letter in the caption is converted to lowercase, and the [`nltk.tokenize.word_tokenize`](http://www.nltk.org/) function is used to obtain a list of string-valued tokens. Run the next code cell to visualize the effect on `sample_caption`.
```
import nltk
sample_tokens = nltk.tokenize.word_tokenize(str(sample_caption).lower())
print(sample_tokens)
```
In **`line 2`** and **`line 3`** we initialize an empty list and append an integer to mark the start of a caption. The [paper](https://arxiv.org/pdf/1411.4555.pdf) that you are encouraged to implement uses a special start word (and a special end word, which we'll examine below) to mark the beginning (and end) of a caption.
This special start word (`"<start>"`) is decided when instantiating the data loader and is passed as a parameter (`start_word`). You are **required** to keep this parameter at its default value (`start_word="<start>"`).
As you will see below, the integer `0` is always used to mark the start of a caption.
```
sample_caption = []
start_word = data_loader.dataset.vocab.start_word
print('Special start word:', start_word)
sample_caption.append(data_loader.dataset.vocab(start_word))
print(sample_caption)
```
In **`line 4`**, we continue the list by adding integers that correspond to each of the tokens in the caption.
```
sample_caption.extend([data_loader.dataset.vocab(token) for token in sample_tokens])
print(sample_caption)
```
In **`line 5`**, we append a final integer to mark the end of the caption.
Identical to the case of the special start word (above), the special end word (`"<end>"`) is decided when instantiating the data loader and is passed as a parameter (`end_word`). You are **required** to keep this parameter at its default value (`end_word="<end>"`).
As you will see below, the integer `1` is always used to mark the end of a caption.
```
end_word = data_loader.dataset.vocab.end_word
print('Special end word:', end_word)
sample_caption.append(data_loader.dataset.vocab(end_word))
print(sample_caption)
```
Finally, in **`line 6`**, we convert the list of integers to a PyTorch tensor and cast it to [long type](http://pytorch.org/docs/master/tensors.html#torch.Tensor.long). You can read more about the different types of PyTorch tensors on the [website](http://pytorch.org/docs/master/tensors.html).
```
import torch
sample_caption = torch.Tensor(sample_caption).long()
print(sample_caption)
```
And that's it! In summary, any caption is converted to a list of tokens, with _special_ start and end tokens marking the beginning and end of the sentence:
```
[<start>, 'a', 'person', 'doing', 'a', 'trick', 'while', 'riding', 'a', 'skateboard', '.', <end>]
```
This list of tokens is then turned into a list of integers, where every distinct word in the vocabulary has an associated integer value:
```
[0, 3, 98, 754, 3, 396, 207, 139, 3, 753, 18, 1]
```
Finally, this list is converted to a PyTorch tensor. All of the captions in the COCO dataset are pre-processed using this same procedure from **`lines 1-6`** described above.
As you saw, in order to convert a token to its corresponding integer, we call `data_loader.dataset.vocab` as a function. The details of how this call works can be explored in the `__call__` method in the `Vocabulary` class in **vocabulary.py**.
```python
def __call__(self, word):
if not word in self.word2idx:
return self.word2idx[self.unk_word]
return self.word2idx[word]
```
The `word2idx` instance variable is a Python [dictionary](https://docs.python.org/3/tutorial/datastructures.html#dictionaries) that is indexed by string-valued keys (mostly tokens obtained from training captions). For each key, the corresponding value is the integer that the token is mapped to in the pre-processing step.
Use the code cell below to view a subset of this dictionary.
```
# Preview the word2idx dictionary.
dict(list(data_loader.dataset.vocab.word2idx.items())[:10])
```
We also print the total number of keys.
```
# Print the total number of keys in the word2idx dictionary.
print('Total number of tokens in vocabulary:', len(data_loader.dataset.vocab))
```
As you will see if you examine the code in **vocabulary.py**, the `word2idx` dictionary is created by looping over the captions in the training dataset. If a token appears no less than `vocab_threshold` times in the training set, then it is added as a key to the dictionary and assigned a corresponding unique integer. You will have the option later to amend the `vocab_threshold` argument when instantiating your data loader. Note that in general, **smaller** values for `vocab_threshold` yield a **larger** number of tokens in the vocabulary. You are encouraged to check this for yourself in the next code cell by decreasing the value of `vocab_threshold` before creating a new data loader.
```
# Modify the minimum word count threshold.
vocab_threshold = 4
# Obtain the data loader.
data_loader = get_loader(transform=transform_train,
mode='train',
batch_size=batch_size,
vocab_threshold=vocab_threshold,
vocab_from_file=False)
# Print the total number of keys in the word2idx dictionary.
print('Total number of tokens in vocabulary:', len(data_loader.dataset.vocab))
```
There are also a few special keys in the `word2idx` dictionary. You are already familiar with the special start word (`"<start>"`) and special end word (`"<end>"`). There is one more special token, corresponding to unknown words (`"<unk>"`). All tokens that don't appear anywhere in the `word2idx` dictionary are considered unknown words. In the pre-processing step, any unknown tokens are mapped to the integer `2`.
```
unk_word = data_loader.dataset.vocab.unk_word
print('Special unknown word:', unk_word)
print('All unknown words are mapped to this integer:', data_loader.dataset.vocab(unk_word))
```
Check this for yourself below, by pre-processing the provided nonsense words that never appear in the training captions.
```
print(data_loader.dataset.vocab('jfkafejw'))
print(data_loader.dataset.vocab('ieowoqjf'))
```
The final thing to mention is the `vocab_from_file` argument that is supplied when creating a data loader. To understand this argument, note that when you create a new data loader, the vocabulary (`data_loader.dataset.vocab`) is saved as a [pickle](https://docs.python.org/3/library/pickle.html) file in the project folder, with filename `vocab.pkl`.
If you are still tweaking the value of the `vocab_threshold` argument, you **must** set `vocab_from_file=False` to have your changes take effect.
But once you are happy with the value that you have chosen for the `vocab_threshold` argument, you need only run the data loader *one more time* with your chosen `vocab_threshold` to save the new vocabulary to file. Then, you can henceforth set `vocab_from_file=True` to load the vocabulary from file and speed the instantiation of the data loader. Note that building the vocabulary from scratch is the most time-consuming part of instantiating the data loader, and so you are strongly encouraged to set `vocab_from_file=True` as soon as you are able.
Note that if `vocab_from_file=True`, then any supplied argument for `vocab_threshold` when instantiating the data loader is completely ignored.
```
# Obtain the data loader (from file). Note that it runs much faster than before!
data_loader = get_loader(transform=transform_train,
mode='train',
batch_size=batch_size,
vocab_from_file=True)
```
In the next section, you will learn how to use the data loader to obtain batches of training data.
<a id='step2'></a>
## Step 2: Use the Data Loader to Obtain Batches
The captions in the dataset vary greatly in length. You can see this by examining `data_loader.dataset.caption_lengths`, a Python list with one entry for each training caption (where the value stores the length of the corresponding caption).
In the code cell below, we use this list to print the total number of captions in the training data with each length. As you will see below, the majority of captions have length 10. Likewise, very short and very long captions are quite rare.
```
from collections import Counter
# Tally the total number of training captions with each length.
counter = Counter(data_loader.dataset.caption_lengths)
lengths = sorted(counter.items(), key=lambda pair: pair[1], reverse=True)
for value, count in lengths:
print('value: %2d --- count: %5d' % (value, count))
```
To generate batches of training data, we begin by first sampling a caption length (where the probability that any length is drawn is proportional to the number of captions with that length in the dataset). Then, we retrieve a batch of size `batch_size` of image-caption pairs, where all captions have the sampled length. This approach for assembling batches matches the procedure in [this paper](https://arxiv.org/pdf/1502.03044.pdf) and has been shown to be computationally efficient without degrading performance.
Run the code cell below to generate a batch. The `get_train_indices` method in the `CoCoDataset` class first samples a caption length, and then samples `batch_size` indices corresponding to training data points with captions of that length. These indices are stored below in `indices`.
These indices are supplied to the data loader, which then is used to retrieve the corresponding data points. The pre-processed images and captions in the batch are stored in `images` and `captions`.
```
import numpy as np
import torch.utils.data as data
# Randomly sample a caption length, and sample indices with that length.
indices = data_loader.dataset.get_train_indices()
print('sampled indices:', indices)
# Create and assign a batch sampler to retrieve a batch with the sampled indices.
new_sampler = data.sampler.SubsetRandomSampler(indices=indices)
data_loader.batch_sampler.sampler = new_sampler
# Obtain the batch.
images, captions = next(iter(data_loader))
print('images.shape:', images.shape)
print('captions.shape:', captions.shape)
# (Optional) Uncomment the lines of code below to print the pre-processed images and captions.
print('images:', images)
print('captions:', captions)
```
Each time you run the code cell above, a different caption length is sampled, and a different batch of training data is returned. Run the code cell multiple times to check this out!
You will train your model in the next notebook in this sequence (**2_Training.ipynb**). This code for generating training batches will be provided to you.
> Before moving to the next notebook in the sequence (**2_Training.ipynb**), you are strongly encouraged to take the time to become very familiar with the code in **data_loader.py** and **vocabulary.py**. **Step 1** and **Step 2** of this notebook are designed to help facilitate a basic introduction and guide your understanding. However, our description is not exhaustive, and it is up to you (as part of the project) to learn how to best utilize these files to complete the project. __You should NOT amend any of the code in either *data_loader.py* or *vocabulary.py*.__
In the next steps, we focus on learning how to specify a CNN-RNN architecture in PyTorch, towards the goal of image captioning.
<a id='step3'></a>
## Step 3: Experiment with the CNN Encoder
Run the code cell below to import `EncoderCNN` and `DecoderRNN` from **model.py**.
```
# Watch for any changes in model.py, and re-load it automatically.
% load_ext autoreload
% autoreload 2
# Import EncoderCNN and DecoderRNN.
from model import EncoderCNN, DecoderRNN
```
In the next code cell we define a `device` that you will use move PyTorch tensors to GPU (if CUDA is available). Run this code cell before continuing.
```
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
```
Run the code cell below to instantiate the CNN encoder in `encoder`.
The pre-processed images from the batch in **Step 2** of this notebook are then passed through the encoder, and the output is stored in `features`.
```
# Specify the dimensionality of the image embedding.
embed_size = 256
#-#-#-# Do NOT modify the code below this line. #-#-#-#
# Initialize the encoder. (Optional: Add additional arguments if necessary.)
encoder = EncoderCNN(embed_size)
# Move the encoder to GPU if CUDA is available.
encoder.to(device)
# Move last batch of images (from Step 2) to GPU if CUDA is available.
images = images.to(device)
# Pass the images through the encoder.
features = encoder(images)
print('type(features):', type(features))
print('features.shape:', features.shape)
# Check that your encoder satisfies some requirements of the project! :D
assert type(features)==torch.Tensor, "Encoder output needs to be a PyTorch Tensor."
assert (features.shape[0]==batch_size) & (features.shape[1]==embed_size), "The shape of the encoder output is incorrect."
```
The encoder that we provide to you uses the pre-trained ResNet-50 architecture (with the final fully-connected layer removed) to extract features from a batch of pre-processed images. The output is then flattened to a vector, before being passed through a `Linear` layer to transform the feature vector to have the same size as the word embedding.

You are welcome (and encouraged) to amend the encoder in **model.py**, to experiment with other architectures. In particular, consider using a [different pre-trained model architecture](http://pytorch.org/docs/master/torchvision/models.html). You may also like to [add batch normalization](http://pytorch.org/docs/master/nn.html#normalization-layers).
> You are **not** required to change anything about the encoder.
For this project, you **must** incorporate a pre-trained CNN into your encoder. Your `EncoderCNN` class must take `embed_size` as an input argument, which will also correspond to the dimensionality of the input to the RNN decoder that you will implement in Step 4. When you train your model in the next notebook in this sequence (**2_Training.ipynb**), you are welcome to tweak the value of `embed_size`.
If you decide to modify the `EncoderCNN` class, save **model.py** and re-execute the code cell above. If the code cell returns an assertion error, then please follow the instructions to modify your code before proceeding. The assert statements ensure that `features` is a PyTorch tensor with shape `[batch_size, embed_size]`.
<a id='step4'></a>
## Step 4: Implement the RNN Decoder
Before executing the next code cell, you must write `__init__` and `forward` methods in the `DecoderRNN` class in **model.py**. (Do **not** write the `sample` method yet - you will work with this method when you reach **3_Inference.ipynb**.)
> The `__init__` and `forward` methods in the `DecoderRNN` class are the only things that you **need** to modify as part of this notebook. You will write more implementations in the notebooks that appear later in the sequence.
Your decoder will be an instance of the `DecoderRNN` class and must accept as input:
- the PyTorch tensor `features` containing the embedded image features (outputted in Step 3, when the last batch of images from Step 2 was passed through `encoder`), along with
- a PyTorch tensor corresponding to the last batch of captions (`captions`) from Step 2.
Note that the way we have written the data loader should simplify your code a bit. In particular, every training batch will contain pre-processed captions where all have the same length (`captions.shape[1]`), so **you do not need to worry about padding**.
> While you are encouraged to implement the decoder described in [this paper](https://arxiv.org/pdf/1411.4555.pdf), you are welcome to implement any architecture of your choosing, as long as it uses at least one RNN layer, with hidden dimension `hidden_size`.
Although you will test the decoder using the last batch that is currently stored in the notebook, your decoder should be written to accept an arbitrary batch (of embedded image features and pre-processed captions [where all captions have the same length]) as input.

In the code cell below, `outputs` should be a PyTorch tensor with size `[batch_size, captions.shape[1], vocab_size]`. Your output should be designed such that `outputs[i,j,k]` contains the model's predicted score, indicating how likely the `j`-th token in the `i`-th caption in the batch is the `k`-th token in the vocabulary. In the next notebook of the sequence (**2_Training.ipynb**), we provide code to supply these scores to the [`torch.nn.CrossEntropyLoss`](http://pytorch.org/docs/master/nn.html#torch.nn.CrossEntropyLoss) optimizer in PyTorch.
```
# Specify the number of features in the hidden state of the RNN decoder.
hidden_size = 512
#-#-#-# Do NOT modify the code below this line. #-#-#-#
# Store the size of the vocabulary.
vocab_size = len(data_loader.dataset.vocab)
# Initialize the decoder.
decoder = DecoderRNN(embed_size, hidden_size, vocab_size)
# Move the decoder to GPU if CUDA is available.
decoder.to(device)
# Move last batch of captions (from Step 1) to GPU if CUDA is available
captions = captions.to(device)
# Pass the encoder output and captions through the decoder.
outputs = decoder(features, captions)
print('type(outputs):', type(outputs))
print('outputs.shape:', outputs.shape)
# Check that your decoder satisfies some requirements of the project! :D
assert type(outputs)==torch.Tensor, "Decoder output needs to be a PyTorch Tensor."
assert (outputs.shape[0]==batch_size) & (outputs.shape[1]==captions.shape[1]) & (outputs.shape[2]==vocab_size), "The shape of the decoder output is incorrect."
```
When you train your model in the next notebook in this sequence (**2_Training.ipynb**), you are welcome to tweak the value of `hidden_size`.
| github_jupyter |
Im Phishing-Register Phishtank.com sind auch Schweizer Top-Level-Domains, die von der Melani gemeldet wurden, zu finden. Welche davon waren von Phishing wirklich betroffen? Und was sagen Sie dazu, dass Sie auf Phishtank.com zu finden sind?
```
import pandas as pd
import numpy as np
import progressbar
import matplotlib.pyplot as plt
import matplotlib
plt.style.use('fivethirtyeight')
%matplotlib inline
ph = pd.read_csv('d/goverCERTphishes.csv')
ph.index = pd.to_datetime(ph['Date'], format='%b %d %Y %I:%M %p')
ph.info()
```
## Finding Swiss sites
```
def com(elem):
if ".com" in elem:
return True
else:
return False
ph['com check'] = ph['URL'].apply(com)
ch = ph[ph['com check']==True]
ch.info()
```
Finding Top Level Domain
```
def toplevel(elem):
elem = elem.replace("https://","").replace("http://","").split("/")[0]
return elem
ch['toplevel'] = ch['URL'].apply(toplevel)
ch
def endinginch(elem):
if elem[-4:] == '.com':
return True
else:
return False
ch['ch toplevel'] = ch['toplevel'].apply(endinginch)
```
# Examining Swiss toplevel domains
```
chtl = ch[ch['ch toplevel']==True]
chtl.info()
chtl[chtl['toplevel']=='www.restauranteabiss.com']
chtl['toplevel'].value_counts()
```
## Grundlage.ch
```
chtl[chtl['toplevel']=='www.grundlage.ch']
#Links:
#https://www.phishtank.com/phish_detail.php?phish_id=4676051
#https://www.phishtank.com/phish_detail.php?phish_id=4648313
```
## Singleactive.ch
```
chtl[chtl['toplevel']=='dating.singleactive.ch']
#Links
#https://www.phishtank.com/phish_detail.php?phish_id=4576196
#https://www.phishtank.com/phish_detail.php?phish_id=4556999
```
## Der Bund
```
chtl[chtl['toplevel']=='der-bund.ch']
```
Domain name:
der-bund.ch
Holder of domain name:
Schlagwort AG
Massat Remo Reimut
Abteilung Technik
Schauenberg 99
CH-7421 Summaprada - Cazis
Switzerland
Technical contact:
Schlagwort AG
Massat Remo Reimut
Abteilung Technik
Schauenberg 99
CH-7421 Summaprada - Cazis
Switzerland
Registrar:
EuroDNS S.A.
First registration date:
2008-11-07
## Nach Datum sortiert
```
pd.options.display.max_rows = 999
chtl.sort_index(ascending=False)
#kruesi-ag.ch
#https://www.phishtank.com/phish_detail.php?phish_id=5580598
#http://kruesi-ag.ch/fileadmin/
#https://www.furtbaechler.ch/
#https://www.phishtank.com/phish_detail.php?phish_id=5550973
#http://www.furtbaechler.ch/europe/
#Lorente.ch
#https://whois.domaintools.com/lorente.ch
#https://www.phishtank.com/phish_detail.php?phish_id=5546192
#http://www.lorente.ch/wildturkeyrocks/guestbook/go.php?url=https://redirect.billing.info-live.cgi-bin.webapps-mpp.home.verified-suiss.ch/iService/
#doctorbook24
#https://doctorbook24.ch/assets/data/Login/
#https://www.phishtank.com/phish_detail.php?phish_id=5261716
#https://doctorbook24.ch/assets/data/Login/
#https://www.stu.ch/
#https://www.phishtank.com/phish_detail.php?phish_id=5216948
#https://www.stu.ch/dd.php
#https://docs.google.com/document/d/1NAh5sciO_q2VoTLKJk6Y5O_pmCmGwXkufsDBAvzp2nU/edit
#http://buergisserweb.ch/
#https://www.phishtank.com/phish_detail.php?phish_id=5089593
#http://buergisserweb.ch/de/index.php
#https://docs.google.com/document/d/1NAh5sciO_q2VoTLKJk6Y5O_pmCmGwXkufsDBAvzp2nU/edit
#lovelysilk.ch
#https://www.phishtank.com/phish_detail.php?phish_id=4869162
#http://lovelysilk.ch/add/dpbx/index.php, Joomla.
#https://docs.google.com/document/d/1NAh5sciO_q2VoTLKJk6Y5O_pmCmGwXkufsDBAvzp2nU/edit
#Metanet AG in Zürich, Hardstrasse 235 Zürich. Heidi Ullmann, Tel.: +41 79 357 88 82. Heutiger Inhaber
#Herr Zahnd weiss nichts davon. Auch Heidi Ullmann. Wie steht es um die Metanet AG. Metanet AG sagt
#allerdings, dass sie die Site nur registriert hat. Zuständig ist eventNET.ch: Guido Blanke. +41 71 560 5445.
#Was sagt eventNET. Frau Tilia Schnarwiler, tis@metanet.ch
# DNS: "0848 830 740"
#https://www.clubmove.ch/
#https://www.phishtank.com/phish_detail.php?phish_id=4844761
#http://clubmove.ch/8NPjX4N/index.php
#https://docs.google.com/document/d/1NAh5sciO_q2VoTLKJk6Y5O_pmCmGwXkufsDBAvzp2nU/edit
#Gamelab.ch
#https://www.phishtank.com/phish_detail.php?phish_id=4756603
#http://gamelab.ch/wp-content/uploads/2pQEzvfeZT8/index.php
##https://docs.google.com/document/d/1NAh5sciO_q2VoTLKJk6Y5O_pmCmGwXkufsDBAvzp2nU/edit
#teatime.ch
#https://www.phishtank.com/phish_detail.php?phish_id=4717935
#https://www.teatime.ch/modules/blockadvertising/2016/client_data/
#https://docs.google.com/document/d/1NAh5sciO_q2VoTLKJk6Y5O_pmCmGwXkufsDBAvzp2nU/edit
#hin.ch
#https://www.phishtank.com/phish_detail.php?phish_id=4662453
#http://my-hin.ch/?uid=WVEJINJE
#https://docs.google.com/document/d/1RAqrNdV-2hY01ZDEZDpKhWiL-WrfkJgHeIr7K9_aBlM/edit
#world-of-grappa.ch
#http://www.world-of-grappa.ch/Googledocwwq/Googledoc1/index.html
#https://www.phishtank.com/phish_detail.php?phish_id=2644206
#https://www.phishtank.com/phish_detail.php?phish_id=2644206
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import glob
from pathlib import Path
tmpl_files = glob.glob('/Users/adamamiller/Downloads/templates-2.0/*lnw')
for tf in tmpl_files:
if '87A' in tf:
print(tf)
with open(tf) as f:
ll = f.readlines()
```
### Loop over all files to find matches
```
sn_name = np.empty(len(tmpl_files)).astype(str)
for sn_num, tf in enumerate(tmpl_files):
name = tf.split('/')[-1].split('.lnw')[0]
if len(name.split('sn')) > 1:
this_sn = name.split('sn')[1].split('_b')[0]
if this_sn[0:2] == '19' or this_sn[0:2] == '20':
sn_name[sn_num] = this_sn[2:]
else:
sn_name[sn_num] = this_sn
else:
sn_name[sn_num] = name
match_list = []
for snn in sn_name:
match = np.where(sn_name == snn)
if len(match[0]) > 1 and snn not in match_list:
match_list.append(snn)
```
### Given 2 random files, check for matching dates and remove them
```
match = match_list[-4]
print(match)
for match in match_list:
matches = np.where(sn_name == match)
nspec = np.empty(len(matches[0])).astype(int)
for m_num, m in enumerate(matches[0]):
line1 = pd.read_csv(tmpl_files[m], nrows=1, header=None, delim_whitespace=True)
nspec[m_num] = int(line1[0])
order = np.argsort(nspec)[::-1]
for m_num, m in enumerate(matches[0][order]):
line1 = pd.read_csv(tmpl_files[m], nrows=1, header=None, delim_whitespace=True)
nskip = int(line1[4])
line_ages = pd.read_csv(tmpl_files[m], nrows=1,
skiprows = nskip + 2,
header=None, delim_whitespace=True)
if m_num == 0:
ages = line_ages.iloc[0].values[1:]
continue
else:
dup_idx = []
for spec_num, age in enumerate(line_ages.iloc[0].values[1:]):
if age in list(ages):
dup_idx.append(spec_num)
if len(dup_idx) > 0:
print('warning ', sn_name[m])
# loop to create new file
nspec_this_sn = int(line1[0])
with open(tmpl_files[m]) as tf:
ll = tf.readlines()
with open(tmpl_files[m].replace('.lnw','_new.lnw'), 'w') as tfw:
new_line1 = '{0:>5}'.format(nspec_this_sn - len(dup_idx)) + ll[0][5:]
print(new_line1[:-1], file=tfw)
for l in ll[1:nskip+2]:
new_line = l[0:7]
for dup_num, di in enumerate(dup_idx):
if dup_num == 0:
new_line += l[7:7 + di*16]
else:
new_line += l[23 + dup_idx[dup_num-1]*16:7 + di*16]
if dup_num == len(dup_idx) - 1:
new_line += l[23 + di*16:]
print(new_line[:-1], file=tfw)
for l in ll[nskip+2:]:
new_line = l[0:8]
for dup_num, di in enumerate(dup_idx):
if dup_num == 0:
new_line += l[8:8 + di*9]
else:
new_line += l[17 + dup_idx[dup_num-1]*9:8 + di*9]
if dup_num == len(dup_idx) - 1:
new_line += l[17 + di*9:]
print(new_line[:-1], file=tfw)
Path(tmpl_files[m]).rename(tmpl_files[m].replace('templates-2.0','templates-2.0/old_with_duplicates'))
if m_num + 1 < len(matches[0]):
ages = np.append(ages, line_ages.iloc[0].values[1:])
```
| github_jupyter |
# INFO 7390
# Advances in Data Science and Architecture
# Bantaba Project on Convolutional Neural Network Using TensorFlow

# 1. Abstract
The focus of the project is to understand Convolutional Neural Network(CNN) as a beginner. A very easy way to explain and interpret a CNN is using classification model. In this project, we will be learning how CNN classifies Kannada digits, from 1 through 9
This is a 5 layers Sequential Convolutional Neural Network for digits recognition trained on Kannada digits dataset. I have chosen to build it with keras API (Tensorflow backend) which is very intuitive. To ensure the model did not overfit, we used Keras callbacks
# 2. Table of Contents
1. Abstract
2. Table of Contents
3. Acknowledgement
4. Introduction
5. Running the Notebook
6. Importing packages and collecting data
7. Data Description and preparation
7.1 Checking the Target Distribution
8. Data pre-processing
8.1 Image Normalization
8.2 Reshaping the images
8.3 Splitting training and Validation Set
8.4 Label Encoding target variables
8.5 Data Augmentation
9. Building the Neural Network
9.1 Defining and Outlining the model
9.2 Complie and Traine the model
9.3 Visualizing the Accuracy and Loss
9.4 Prediction
9.4.1 Evaluating Some Wrong Predictions
9.4.2 Evaluating Right Predictions
10. Conclusion
11. Citation
12. Licensing
# 3. Acknowledgment
I acknowledge that the kannada_MNIST dataset was taken from https://www.kaggle.com/c/Kannada-MNIST/overview/description and was created and provided by Vinay Uday Prabhu
https://arxiv.org/abs/1908.01242
https://vinayprabhu.github.io/
# 4. Introduction

There are hundreds of thousand languages world wide. Some have scripts, some don't. **Kannada** is one of the oldest languages that originated in **South India** and is still spoken by majority of the people espicially in **Karnataka**. I am a kannadiga and I can speak, read and write kannada.
The main focus of the project is to write a chapter on CNN that could be read and understood easily; especially for a beginner.
The sole purpose for having chosen this data set is because it is easier to understand the operation of CNNs through MNIST datasets.
The dataset has 3 parts namely, train,test and Dig-MNIST. Dig-MNIST is a real world handwritten dataset (with 10k images), that can serve as an out-of-domain test dataset.
# 5. Running the Notebook
This file was ran on **Google CoLab** with runtime set to **GPU**. For all who don't what a google colab is, here is a breif inroduction- Google Colab is a free cloud service and now it supports free GPU! You can; improve your Python programming language coding skills. develop deep learning applications using popular libraries such as Keras, TensorFlow, PyTorch, and OpenCV.
1. Open **google colab** Jupyter Notebook
2. **Rename** the file
3. Set the runtime to **GPU** [Runtime>change runtime type>hardware accelerator>GPU]
4. To the right side of the page, there is an option to upload files from the local system. Click on upload button and upload the zip file
5. Run the file
# 6. Importing packages and collecting data
### We will mainly use 4 libraries.
pandas and numpy : It's used to handle our csv files.
matplotlib & seaborn : Used for charting and plotting.
sklearn : Popular ML library.We will use it for splitting our data.
Keras : Popular Deep learning library,we will use it to build our CNN Network.
```
'''Importing Data Manipulattion Moduls'''
import numpy as np
import pandas as pd
'''Seaborn and Matplotlib Visualization'''
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
'''Importing preprocessing libraries'''
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix
from zipfile import ZipFile
file_name = "Kannada-MNIST.zip"
import os
import cv2
'''Display markdown formatted output like bold, italic bold etc.'''
from IPython.display import Markdown
def bold(string):
display(Markdown(string))
import tensorflow as tf
print(tf.__version__)
from tensorflow.keras import layers, models
from keras.models import Sequential
from keras.preprocessing.image import ImageDataGenerator
from keras.utils import to_categorical
from keras.layers import Dense,Conv2D,Flatten,MaxPooling2D,Dropout,BatchNormalization
from keras.optimizers import Adam
from keras.callbacks import ReduceLROnPlateau
from keras.callbacks import EarlyStopping, ModelCheckpoint
with ZipFile(file_name,'r') as zip:
zip.extractall()
print('Done')
train_kannadaMnist = pd.read_csv("train.csv")
test_kannadaMnist = pd.read_csv("test.csv")
dig_kannadaMnist = pd.read_csv("Dig-MNIST.csv")
```
# 7. Data Description and Preparation
```
'''Train and test data at a glance.'''
bold('**Preview of Train Data:**')
display(train_kannadaMnist.head(3))
bold('**Preview of Test Data:**')
display(test_kannadaMnist.head(3))
'''Ckecking for null and missing values'''
bold('**Train Data**')
display(train_kannadaMnist.isnull().any(). describe())
bold('**Test Data**')
display(test_kannadaMnist.isnull().any(). describe())
'''Seting X and Y'''
y_train = train_kannadaMnist['label']
# Drop 'label' column
X_train = train_kannadaMnist.drop('label', axis = 1)
X_test = test_kannadaMnist.drop('id', axis = 1)
dig_img = dig_kannadaMnist.drop('label', axis = 1)
"""Let's have a final look at our data"""
bold('**Data Dimension for Model Building:**')
print('Input matrix dimension:', X_train.shape)
print('Output vector dimension:',y_train.shape)
print('Test data dimension:', X_test.shape)
```
## 7.1 Checking the Target Distribution
```
'''Visualizating the taget distribution'''
plt.figure(figsize = (8,8))
sns.countplot(y_train, palette='cubehelix')
plt.show()
print("Cool")
images = train_kannadaMnist.iloc[:,1:].values
images = images.astype(np.float)
# convert from [0:255] => [0.0:1.0]
images = np.multiply(images, 1.0 / 255.0)
image_size = images.shape[1]
print('image_size => {0}'.format(image_size))
# in this case all images are square
image_width = image_height = np.ceil(np.sqrt(image_size)).astype(np.uint8)
print('image_width => {0}\nimage_height => {1}'.format(image_width, image_height))
'''Displaying image'''
# display image
def display(img):
# (784) => (28,28)
one_image = img.reshape(image_width,image_height)
plt.axis('off')
plt.imshow(one_image, cmap='binary')
# output image
display(images[4])
'''Converting X_train to numpy array'''
X_train_array = X_train.to_numpy()
'''Displaying images'''
n=10
fig = plt.figure(figsize=(10,10))
for i in range(n):
ax = fig.add_subplot(2, n, i+1, xticks=[], yticks=[])
ax.imshow(X_train_array[i].reshape(image_width,image_height), cmap='viridis')
ax.axis("off")
plt.tight_layout()
plt.show()
```
# 8. Data Preprocessing
## 8.1 Image Normalization
Normalize Pixel Values
For most image data, the pixel values are integers with values between 0 and 255.
Neural networks process inputs using small weight values, and inputs with large integer values can disrupt or slow down the learning process. As such it is good practice to normalize the pixel values so that each pixel value has a value between 0 and 1.
It is valid for images to have pixel values in the range 0-1 and images can be viewed normally.
This can be achieved by dividing all pixels values by the largest pixel value; that is 255. This is performed across all channels, regardless of the actual range of pixel values that are present in the image.
```
'''Normalizing the data'''
X_train = X_train / 255.0
X_test = X_test / 255.0
dig_img = dig_img/255.0
```
## 8.2 Reshaping the images
Train and test images (28 x 28) has been stock into pandas.Dataframe as 1D vectors of 784 values. We reshape all data to 28x28x1 3D matrices.
Keras requires an extra dimension in the end which correspond to channels. MNIST images are gray scaled so it use only one channel. For RGB images, there is 3 channels, we would have reshaped 784px vectors to 28x28x3 3D matrices.
```
'''Reshape image in 3 dimensions (height = 28px, width = 28px , canal = 1)'''
X_train = X_train.values.reshape(-1,28,28,1)
X_test = X_test.values.reshape(-1,28,28,1)
dig_img = dig_img.values.reshape(-1,28,28,1)
print(X_train.shape, X_test.shape, dig_img.shape)
```
Data reshape into 60000 examples of height 28 and width 28 and 1 channel.
## 8.3 Splitting Training and Validation set
Now we will split out training data into train and validation data 10 percent of the training data will be used for validation purpose.
```
'''Set the random seed'''
seed = 44
'''Split the train and the validation set for the fitting'''
X_train, X_val, y_train, y_val = train_test_split(X_train, y_train, test_size = 0.1, random_state=seed)
```
## 8.4 Label encoding of Target Variable
```
'''convert class labels from scalars to one-hot vectors'''
# 0 => [1 0 0 0 0 0 0 0 0 0]
# 1 => [0 1 0 0 0 0 0 0 0 0]
# ...
# 9 => [0 0 0 0 0 0 0 0 0 1]
y_train = tf.keras.utils.to_categorical(y_train, num_classes = 10, dtype='uint8')
y_val = tf.keras.utils.to_categorical(y_val, num_classes = 10, dtype='uint8')
```
## 8.5 Data Augmentation
To get more training data, to avoid overfitting, data augmentation is used. Data augmentation is the creation of altered copies of each training instance (image) within a training dataset.
1. Ramdomly rotate the images by 10 degrees
2. Randomly zoom the images by 25%
3. Randomly shift its height and width by 25%
In order to avoid overfitting problem, we need to expand artificially our handwritten digit dataset. We can make your existing dataset even larger. The idea is to alter the training data with small transformations to reproduce the variations occuring when someone is writing a digit.
For example, the number is not centered The scale is not the same (some who write with big/small numbers) The image is rotated...
Approaches that alter the training data in ways that change the array representation while keeping the label the same are known as data augmentation techniques. Some popular augmentations people use are grayscales, horizontal flips, vertical flips, random crops, color jitters, translations, rotations, and much more.
By applying just a couple of these transformations to our training data, we can easily double or triple the number of training examples and create a very robust model.
```
# Artificially increase training set
train_datagen = ImageDataGenerator(rescale=1./255.,
rotation_range=10,
width_shift_range=0.25,
height_shift_range=0.25,
shear_range=0.1,
zoom_range=0.25,
horizontal_flip=False)
train_datagen.fit(X_train)
```
# 9. CNN Model
## 9.1 Defining and Outlining the model
I used the Keras Sequential API, where you have just to add one layer at a time, starting from the input.

The first is the ***convolutional (Conv2D) layer***. It is like a set of ***learnable filters***. I choosed to set 32 filters for the two firsts conv2D layers and 64 filters for the two last ones. Each filter transforms a part of the image (defined by the kernel size) using the kernel filter. The kernel filter matrix is applied on the whole image. Filters can be seen as a transformation of the image.

The CNN can extract features that are useful everywhere from these transformed images (feature maps).
***Padding***
we can pad the image with an additional border, i.e., we add one pixel all around the edges.
***Valid:*** It means no padding. If we are using valid padding, the output will be (n-f+1) X (n-f+1)
***Same:*** Here, we apply padding so that the output size is the same as the input size, i.e.,
n+2p-f+1 = n
So, p = (f-1)/2

The second important layer in CNN is the ***pooling (MaxPool2D) layer***. This layer simply acts as a ***downsampling filter***. It looks at the 2 neighboring pixels and picks the maximal value. These are used to reduce computational cost, and to some extent also reduce overfitting. We have to choose the pooling size (i.e the area size pooled each time) more the pooling dimension is high, more the downsampling is important.

Combining convolutional and pooling layers, CNN are able to combine local features and learn more global features of the image.
***Dropout is a regularization method***, where a proportion of nodes in the layer are randomly ignored (setting their wieghts to zero) for each training sample. This drops randomly a propotion of the network and forces the network to learn features in a distributed way. This technique also improves generalization and reduces the overfitting.

***'relu'*** is the rectifier (activation function max(0,x). The rectifier activation function is used to add non linearity to the network.
The ***Flatten layer*** is use to convert the final feature maps into a one single 1D vector. This flattening step is needed so that you can make use of fully connected layers after some convolutional/maxpool layers. It combines all the found local features of the previous convolutional layers.

In the end I have used the features in two ***fully-connected (Dense) layers*** which is just artificial an neural networks (ANN) classifier. In the last layer(Dense(10,activation="softmax")) the net outputs distribution of probability of each class.

Fully Connected Layer and Output Layer Fully connected layers or dense layers are the same hidden layers consisting of defined number of neurons connected with elements of another layer that we discussed in simple ANN. However the output layer is also the same but the number of neurons depend on our task.
In summary, the architecture of CNN , we can simply understand that it consist of an input layer followed by a Conv layer. The dimensions of conv layer depends on the data and problem, hence changing the dimensions accordingly. After the Conv Layer there is a activation layer , usually ReLU since it gives better results. After some conv and relu combination , pooling layer is used to reduce the size. Then after some combination of previously defined architecture, flattening layer is used to flatten the input for fully connected layer. Next to these layer, the last layer is the output layer.
```
'''Set the CNN model'''
# CNN architechture is In -> [[Conv2D->relu]*2 -> MaxPool2D -> Dropout]*2 -> Flatten -> Dense -> Dropout -> Out
#model = tensorflow.keras.Sequential()
model = Sequential()
model.add(Conv2D(32, (5, 5), activation='relu', padding='same', input_shape=(28,28,1)))
model.add(Conv2D(32, (5, 5), activation='relu', padding='same'))
model.add(BatchNormalization(momentum=0.15))
model.add(MaxPooling2D((2, 2)))
model.add(Dropout(0.15))
model.add(Conv2D(64, (5, 5), activation='relu', padding='same'))
model.add(Conv2D(64, (5, 5), activation='relu', padding='same'))
model.add(BatchNormalization(momentum=0.15))
model.add(MaxPooling2D((2, 2)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(128, activation = "relu"))
model.add(Dropout(0.40))
model.add(Dense(64, activation = "relu"))
model.add(Dropout(0.40))
model.add(Dense(10, activation = "softmax"))
model.summary()
```
# 9.2 Complie and Train
In simpler terms, ***optimizers*** shape and mold your model into its most accurate possible form by futzing with the weights. The loss function is the guide to the terrain, telling the optimizer when it’s moving in the right or wrong direction
```
model.compile(optimizer="adam", loss=['categorical_crossentropy'], metrics=['accuracy'])
```
A ***callback*** is a set of functions to be applied at given stages of the training procedure. You can use callbacks to get a view on internal states and statistics of the model during training.
Keras callbacks can help you fix bugs more quickly, and can help you build better models. They can help you visualize how your model’s training is going, and can even help prevent overfitting by implementing early stopping or customizing the learning rate on each iteration.
Here we use 2 callback functions:
***Early Stopping*** - One technique to reduce overfitting in neural networks is to use early stopping. Early stopping prevents overtraining of the model by terminating the training process if it’s not really learning anything.
***Learning Rate Reduction*** - The learning rate determines the size of the steps taken during the gradient descent process.
With the ***ReduceLROnPlateau*** function from Keras.callbacks, i choose to reduce the LR by half if the accuracy is not improved after 3 epochs.
```
# Set a learning rate annealer. Learning rate will be half after 3 epochs if accuracy is not increased
learning_rate_reduction = ReduceLROnPlateau(monitor='val_accuracy',
patience=3,
verbose=1,
factor=0.5,
min_lr=0.00001)
```
### What is the Difference Between a Batch and an Epoch in a Neural Network?
The training process will run for a fixed number of iterations through the dataset called epochs, that we must specify using the epochs argument. We must also set the number of dataset rows that are considered before the model weights are updated within each epoch, called the batch size and set using the batch_size argument.
```
batch_size=75
epochs = 50
#num_classes = 10
#learning_rate = 0.001
filepath = "model.h5"
earlystopper = EarlyStopping(patience=10, verbose=1)
checkpoint = ModelCheckpoint(filepath, monitor='val_loss', verbose=1,
save_best_only=True, mode='min')
callbacks_list = [earlystopper, checkpoint]
history = model.fit(X_train, y_train, batch_size = batch_size , epochs = epochs,
validation_data=(X_val, y_val),
verbose=1,
callbacks=callbacks_list)
```
## 9.3 Visulizing Accuracy and Loss
```
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title('Model accuracy')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['Train', 'Validation'], loc='upper left')
plt.show()
# Plot training & validation loss values
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('Model loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train', 'Validation'], loc='upper left')
plt.show()
# Predict the values from the validation dataset
Y_pred = model.predict(X_val)
# Convert predictions classes to one hot vectors
Y_pred_classes = np.argmax(Y_pred,axis = 1)
# Convert validation observations to one hot vectors
Y_true = np.argmax(y_val,axis = 1)
'''confusion matrix'''
# compute the confusion matrix
confusion_mtx = confusion_matrix(Y_true, Y_pred_classes)
# plot the confusion matrix
f,ax = plt.subplots(figsize=(8, 8))
sns.heatmap(confusion_mtx, annot=True, linewidths=0.01,cmap="Greens",linecolor="gray", fmt= '.1f',ax=ax)
plt.xlabel("Predicted Label")
plt.ylabel("True Label")
plt.title("Confusion Matrix")
plt.show()
```
## 9.4 Making Predictions
```
'''predict results'''
pred_dig = model.predict(dig_img)
dig_kannadaMnist['pred'] = np.argmax(pred_dig, axis=1)
#'''select the indix with the maximum probability'''
#results = np.argmax(results,axis = 1)
# look at those that were classified wrongly in X_dig
dig_kannadaMnist['correct'] = dig_kannadaMnist['label'] - dig_kannadaMnist['pred']
errors = dig_kannadaMnist[dig_kannadaMnist['correct'] != 0]
error_list = errors.index
print('Number of errors is ', len(errors))
print('The indices are ', error_list)
```
## 9.4.1 Some of the wrong predictions for dig dataset:
### Further Investigation..
We will go on to see some of the misclassified images.We will simply inspect them to understand if it was a tough one to predict or not.Let's see...
```
# plot images of some of the wrong predictions for X_dig
plt.figure(figsize=(15,10))
for i in range(40):
plt.subplot(6, 10, i+1)
plt.imshow(dig_img[error_list[i]].reshape((28,28)),cmap=plt.cm.binary)
plt.title("true={}\npredict={}".format(dig_kannadaMnist['label'][error_list[i]],
dig_kannadaMnist['pred'][error_list[i]]), y=0.9)
plt.axis('off')
plt.subplots_adjust(wspace=0.3, hspace=-0.1)
plt.show()
```
Looking at those that were predicted wrongly, there are quite several difficult and ambiguous ones.
## 9.4.2 Correct Predictions
**Some examples of predictions made**
```
# predict on test set
predictions = model.predict(X_test)
print(predictions.shape)
# set the predicted labels to be the one with the highest probability
predicted_labels = np.argmax(predictions, axis=1)
# look at some of the predictions for test_X
plt.figure(figsize=(15,6))
for i in range(40):
plt.subplot(4, 10, i+1)
plt.imshow(X_test[i].reshape((28,28)),cmap=plt.cm.binary)
plt.title("predict=%d" % predicted_labels[i],y=0.9)
plt.axis('off')
plt.subplots_adjust(wspace=0.3, hspace=-0.1)
plt.show()
```
**My eyeball reading of the prediction is:**
3 0 2 6 7 7 1 9 3 4
8 8 1 7 8 1 5 1 5 9
3 7 6 0 2 0 8 7 0 0
8 9 2 3 2 4 6 0 7 8
**Looks reasonable.**
***I am a kannadiga(Native kannada speaker) and I am fluent in reading, writing and speaking in this language***
# 10.Conclusion
The above model works pretty well on the traing, validation and test dataset. The Notebook is designed in such a way that it could be easily understood by a Newbie(beginner). I believe that I have covered and explained all the concepts of Convolutional Neural Network.
Further, the model can be applied to a whole new image which is not previously seen/learnt by the model
# 11. Author
This notebook was created by ***Indupriya Kompi Sadasivappa***, currently enrolled student at ***Northeastern University, Boston*** as a contribution to ***Bantaba Projects*** in the month of ***April,2020*** under the guidance and supervision of ***Nicholas Brown***, Assistant Teaching Professor, Multidisciplinary Graduate Engineering Programs at ***Northeastern University, Boston***
kompisadasivappa.i@husky.neu.edu
Bearing **NUID 001051831**

# 11.Citation
1. https://www.kaggle.com/c/Kannada-MNIST
2. https://www.kaggle.com/kaushal2896/kannada-mnist-using-cnn/notebook
3. https://www.kaggle.com/shahules/indian-way-to-learn-cnn
4. https://towardsdatascience.com/
5. Picture are taken from https://www.google.com/imghp?hl=en, a few of the images/giphys are snipped from https://towardsdatascience.com/
# 12.Licensing
Copyright 2020 Indupriya Kompi Sadasivappa
Permission is hereby granted, free of charge, to any person obtaining a copy of this software and associated documentation files (the "Software"), to deal in the Software without restriction, including without limitation the rights to use, copy, modify, merge, publish, distribute, sublicense, and/or sell copies of the Software, and to permit persons to whom the Software is furnished to do so, subject to the following conditions:
The above copyright notice and this permission notice shall be included in all copies or substantial portions of the Software.
THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE SOFTWARE.
| github_jupyter |
```
import multiprocessing as mp
import psutil
import pickle
import time
import sys
import pandas as pd
import numpy as np
from tqdm.notebook import tqdm
# from google.colab import drive
# drive.mount('/content/drive')
# !xz -d -v Moscow_mkad.osm.xz
# парсинг XML
import xml.etree.ElementTree as ET
tree = ET.parse('./Moscow_mkad.osm')
root=tree.getroot()
element = root[0]
"""
Создание матрицы высота
Минимум: 75
Максимум: 317
Генерация np матрицы высот
"""
cell_size = 0.00083333333333333
lat_start = 55.0
lon_start = 35.0
matrix_of_heights = []
with open('./srtm_44_01.asc') as f:
data_str = f.readlines()
for row in data_str[6:]:
row_list = list(map(int, row.split()))
matrix_of_heights.append(row_list)
del row_list
del data_str
matrix_of_heights = np.array(matrix_of_heights, np.ushort)
def haversine_np(lon1, lat1, lon2, lat2):
"""
Calculate the great circle distance between two points
on the earth (specified in decimal degrees)
All args must be of equal length.
"""
lon1, lat1, lon2, lat2 = map(np.radians, [lon1, lat1, lon2, lat2])
dlon = lon2 - lon1
dlat = lat2 - lat1
a = np.sin(dlat/2.0)**2 + np.cos(lat1) * np.cos(lat2) * np.sin(dlon/2.0)**2
c = 2 * np.arcsin(np.sqrt(a))
m = 6367 * c * 1000
return m
def get_height(lat: float, lon: float) -> float:
"""
Получение высоты над уровнем моря по координатам
:param lat: Долгота точки
:param lon: Широта точки
:return: Высота над уровнем моря
"""
semi_x = (lat - lat_start) / cell_size
semi_y = (lon - lon_start) / cell_size
if semi_x > 6000 or semi_y > 6000:
raise Exception('Out of bounds, not in Moscow')
points = [matrix_of_heights[int(np.floor(semi_x)), int(np.floor(semi_y))],
matrix_of_heights[int(np.floor(semi_x)), int(np.ceil(semi_y))],
matrix_of_heights[int(np.ceil(semi_x)), int(np.floor(semi_y))],
matrix_of_heights[int(np.ceil(semi_x)), int(np.ceil(semi_y))]]
floor_lat = np.floor(semi_x) * cell_size + lat_start
floor_lon = np.floor(semi_y) * cell_size + lon_start
ceil_lat = np.ceil(semi_x) * cell_size + lat_start
ceil_lon = np.ceil(semi_y) * cell_size + lon_start
coordinates = [[floor_lat, floor_lon], [floor_lat, ceil_lon], [ceil_lat, floor_lon],
[ceil_lat, ceil_lon]]
idx_min, min_ = 0, 200
for idx, point in enumerate(coordinates):
dist_ = haversine_np(lon, lat, point[1], point[0])
if dist_ < min_:
min_ = dist_
idx_min = idx
triangle = [points[idx_min-1], points[idx_min], points[(idx_min+1)%4]]
return sum(triangle)/3
get_height(55.7558, 37.6173)
"""
Генерация листа эджей формата
[{"nodes": [], "highway":"", "surface":"asphalt", "lanes":1, "width":2}]
А также листа нод формата [{id, lat, lon}]
"""
nodes = []
edges = []
for i in tqdm(range(1, len(root))):
obj = root[i]
if obj.tag == 'node':
nodes.append({'id': obj.attrib['id'],
'lat': obj.attrib['lat'], 'lon': obj.attrib['lon']})
if obj.tag == 'way':
tmp = {"nodes": [], "highway":"", "surface":"asphalt", "lanes":1,
"width":2}
ok = False
for child in obj:
if child.tag == 'nd':
tmp['nodes'].append(child.attrib["ref"])
if child.tag == 'tag':
if child.attrib['k'] == 'highway':
ok = True
if child.attrib['k'] in tmp:
tmp[child.attrib['k']] = child.attrib['v']
if ok and tmp['highway'] in ['pedestrian', 'bridleway', 'cycleway',
'footway', 'living_street', 'path',
'steps', 'residential', 'service']:
edges.append(tmp)
print(f'Edges: {len(edges):,}, Nodes: {len(nodes):,}')
# Удаления парсера XML
del tree
del root
# словарь айди_ноды: её координаты
id2cor = {}
for i in nodes:
id2cor[int(i['id'])] = {'lat': float(i['lat']), 'lon': float(i['lon'])}
# Генерация листа словарей edge
# Медленная!
count = 0
edge_list = []
for edge in tqdm(edges):
for i in range(len(edge['nodes'])-1):
node_1 = id2cor[edge['nodes'][i]]
node_2 = id2cor[edge['nodes'][i+1]]
lon1 = float(node_1['lon'])
lat1 = float(node_1['lat'])
lon2 = float(node_2['lon'])
lat2 = float(node_2['lat'])
dist = haversine_np(lon1, lat1, lon2, lat2)
hight1 = get_height(lat1, lon1)
hight2 = get_height(lat2, lon2)
if isinstance(edge['width'], str):
try:
width_f = float(edge['width'])
except ValueError:
width_f = 1
else:
width_f = float(edge['width'])
edge_list.append({"edge_id":count, "id1":int(edge['nodes'][i]),
"id2":int(edge['nodes'][i+1]), "dist": dist,
"highway": edge['highway'], "surface":edge['surface'],
"lanes": int(edge['lanes']), "width":width_f,
'hight1': hight1, 'hight2': hight2})
count += 1
edge_list[22:25]
import pandas as pd
graph = pd.DataFrame(edge_list, columns=edge_list[0].keys())
graph.to_csv('edge_list.csv', index=False)
graph.head()
# Генерация локации еджа
edge_cors = []
for i in tqdm(range(len(graph))):
cor1 = np.array([id2cor[graph.id1.iloc[i]]['lat'], id2cor[graph.id1.iloc[i]]['lon']])
cor2 = np.array([id2cor[graph.id2.iloc[i]]['lat'], id2cor[graph.id2.iloc[i]]['lon']])
cor = (np.sum([cor1, cor2], axis=0)) / 2
edge_cors.append(cor)
edge_cors[:5]
# Добавление локации еджа в граф
graph["lat"] = np.array(edge_cors)[:, 0]
graph["lon"] = np.array(edge_cors)[:, 1]
graph.head()
graph.to_csv("edge_with_location.csv")
tmp_graph = graph.drop(columns=['highway', 'surface', 'width', 'hight1', 'hight2', 'dist', 'lanes'])
graph_np = tmp_graph.values
del tmp_graph
graph_np[:2], len(graph_np)
# uint
# max id: 8912249766
# min id: 27717690
fast_graph_np = np.genfromtxt('fast_graph.csv', dtype=np.uint, delimiter=',')[1:]
# fast_graph_np = []
# for edge in tqdm(graph_np):
# fast_graph_np.append([edge[0], int(edge[1]), int(edge[2])])
# fast_graph_np = np.array(fast_graph_np, dtype=np.uint)
fast_graph_np[:5]
fast_graph_df = pd.DataFrame(data=fast_graph_np)
fast_graph_df.to_csv('fast_graph.csv', index=False)
fast_graph_np[:,1].min(), fast_graph_np[:,1].max(), fast_graph_np[:,2].min(), fast_graph_np[:,2].max()
from tqdm import tqdm
size = len(fast_graph_np)+1
workers_count = mp.cpu_count() - 10
part_size = len(fast_graph_np[:size])//workers_count
workers_count
print(part_size, mp.cpu_count(), workers_count, len(fast_graph_np[:size]))
def spawn():
dataframe_list = []
procs = list()
manager = mp.Manager()
return_list = manager.list()
for cpu in range(workers_count):
up_border = part_size*(cpu+1)
p = mp.Process(target=run_child, args=(up_border, return_list))
p.start()
procs.append(p)
for idx, p in enumerate(procs):
p.join()
print('Done: {}'.format(idx), end=' ')
return return_list
return_list = spawn()
def run_child(up_border, return_list):
# print('Run child: ', up_border, end='\t')
adj = []
for i in tqdm(range(up_border - part_size, up_border)):
a = fast_graph_np[fast_graph_np[:,2] == fast_graph_np[i,1]][:, 0]
b = fast_graph_np[fast_graph_np[:,1] == fast_graph_np[i,1]][:, 0].tolist()
b.remove(i)
df = b + a.tolist()
adj.append(df)
return_list.extend(adj)
adj = list(return_list)
len(adj)
# import pickle
# with open('adj.pkl', 'wb') as fp:
# pickle.dump(adj, fp, protocol=pickle.HIGHEST_PROTOCOL)
adj[:10]
# adj_str = cast_to_string(return_list)
adj_df = pd.DataFrame(data=adj)
adj_df.to_csv('duo_edge.csv', index=False)
def cast_to_string(routes_property):
return [str(int).replace("[", "").replace("]", "") for int in routes_property]
adj_str = cast_to_string(adj)
adj_df = pd.DataFrame(adj_str, columns = ["adjacent"])
import pickle
with open('nodes.pkl', 'wb') as fp:
pickle.dump(nodes, fp, protocol=pickle.HIGHEST_PROTOCOL)
import pickle
with open('edges.pkl', 'wb') as fp:
pickle.dump(edges, fp, protocol=pickle.HIGHEST_PROTOCOL)
count_pos = 0
count_neg = 0
edge_list = []
for edge in edges:
tmp = []
for node in edge:
if node in id2cor:
count_pos += 1
tmp.append(node)
else:
count_neg += 1
edge_list.append(tmp)
count_pos
a = []
for edge in edges[102]:
a.append({"id": edge, "lat": id2cor[edge]['lat'], "lon": id2cor[edge]['lon']})
df = pd.DataFrame(a, columns=["id", "lat", 'lon'])
df.to_csv('list2.csv', index=False)
dist_edge = []
for edge in tqdm(edges):
for i in range(len(edge)-1):
dist = haversine_np(float(id2cor[edge[i]]['lon']), float(id2cor[edge[i]]['lat']), float(id2cor[edge[i + 1]]['lon']), float(id2cor[edge[i + 1]]['lat']))
dist_edge.append({"id1": edge[i], "id2": edge[i + 1], 'd': dist * 1000})
import pickle
with open('edge_dist.pkl', 'wb') as fp:
pickle.dump(dist_edge, fp, protocol=pickle.HIGHEST_PROTOCOL)
edge_list_for_kepler = []
for edge in tqdm(edges):
for i in range(len(edge['nodes'])-1):
node_1 = id2cor[edge['nodes'][i]]
node_2 = id2cor[edge['nodes'][i+1]]
lon1 = float(node_1['lon'])
lat1 = float(node_1['lat'])
lon2 = float(node_2['lon'])
lat2 = float(node_2['lat'])
dist = haversine_np(lon1, lat1, lon2, lat2)
hight1 = get_height(lat1, lon1)
hight2 = get_height(lat2, lon2)
if isinstance(edge['width'], str):
try:
width_f = float(edge['width'])
except ValueError:
width_f = 1
else:
width_f = float(edge['width'])
edge_list_for_kepler.append({"dist": dist, "highway": edge['highway'], "surface":edge['surface'],
"lanes": int(edge['lanes']), "width":edge['width'],
'hight1': hight1, 'hight2': hight2, 'lat1': lat1, 'lon1': lon1,
'lat2': lat2, 'lon2': lon2})
df = pd.DataFrame(edge_list_for_kepler, columns=edge_list_for_kepler[0].keys())
df.to_csv('edge_list_for_kepler.csv', index=False)
import pickle
with open('nodes.p', 'rb') as fp:
nodes = pickle.load(fp)
edges_pickles = pickle.load(open('nodes.p', 'rb'))
pd.DataFrame(hist_nodes).to_csv("hist_nodes.csv")
import json
with open('dtp_moskva.geojson') as fp:
data = json.load(fp)
data['features'][1]['geometry']
len(data['features'])
dtp = []
for i in tqdm(range(len(data['features']))):
dtp.append({"lat": data['features'][i]['geometry']['coordinates'][1], "lon":data['features'][i]['geometry']['coordinates'][0]})
```
| github_jupyter |
# Stage 0: SETUP
The below libraries are used for this project. For a full list of requirements and versions, please see the requirements.txt file included in the repository.
```
import json
import requests
import os
import pandas as pd
import numpy as np
import matplotlib.dates as mdates
import matplotlib.pyplot as plt
```
# Stage 1: DATA ACQUISITION
## Overview
Data is acquired through the Wikimedia REST API and saved as json files. These files are included in the repository in the *data* folder; you made skip to Stage 2 and use the included files if desired, or skip to Stage 3 to skip processing entirely.
We will request data from both the [Legacy](https://wikitech.wikimedia.org/wiki/Analytics/AQS/Legacy_Pagecounts) and [Pageviews](https://wikitech.wikimedia.org/wiki/Analytics/AQS/Pageviews) API.
We define base templates for the parameters. English wikipedia with monthyl granularity will always be requested, and on the pageviews api we always request agent=user to filter out crawler and bot traffic. We also request consistent dateranges for each api
```
endpoint_legacy = 'https://wikimedia.org/api/rest_v1/metrics/legacy/pagecounts/aggregate/{project}/{site}/{granularity}/{start}/{end}'
endpoint_pageviews = 'https://wikimedia.org/api/rest_v1/metrics/pageviews/aggregate/{project}/{site}/{agent}/{granularity}/{start}/{end}'
params_legacy = {"project" : "en.wikipedia.org",
"granularity" : "monthly",
"start" : "2008010100",
"end" : "2016080100"
}
params_pageviews = {"project" : "en.wikipedia.org",
"agent" : "user",
"granularity" : "monthly",
"start" : "2015070100",
"end" : "2021090100"
}
```
We request each endpoint for each access type, except for aggregates. All data is saved in the *data* folder.
```
def api_call(endpoint,parameters):
headers = {
'User-Agent': 'https://github.com/Cain93',
'From': 'ccase20@uw.edu'
}
call = requests.get(endpoint.format(**parameters), headers=headers)
response = call.json()
return response
legacy_sites = ["desktop-site", "mobile-site"]
pageview_sites = ["desktop", "mobile-app", "mobile-web"]
file_template = "data/{apiname}_{site}_{daterange}.json"
for site in legacy_sites:
data = api_call(endpoint_legacy, {**params_legacy, "site":site})
fileName = file_template.format(apiname="pagecount", site=site, daterange = "200801-201607")
with open(fileName, 'w') as outfile:
json.dump(data, outfile)
for site in pageview_sites:
data = api_call(endpoint_pageviews, {**params_pageviews, "site":site})
fileName = file_template.format(apiname="pageview", site=site, daterange = "201507-202108")
with open(fileName, 'w') as outfile:
json.dump(data, outfile)
```
# Stage 2: DATA PROCESSING
Data is consoldiated and formatted into a single file. This file is included in the repository as *en-wikipedia_traffic_200712-202108.csv*; you may skip to Stage 3 and use the included file if desired.
First we open each file and combine into a dataframe. While doing, we rename columns to make them consistent between legacy and pageview data.
```
combined_data = pd.DataFrame()
col_names = {
"access-site": "access",
"count": "views"
}
for filename in os.listdir("data"):
file = open("data/" + filename, "r")
file_data = json.loads(file.read())
file_df = pd.DataFrame.from_records(file_data["items"]).rename(columns = col_names)
combined_data = combined_data.append(file_df)
combined_data.head()
```
Then we parse the timestamp into year and month, and remove unused columns.
```
combined_data["year"] = combined_data["timestamp"].apply(lambda x: x[0:4])
combined_data["month"] = combined_data["timestamp"].apply(lambda x: x[4:6])
cleaned_data = combined_data.drop(columns=["timestamp", "granularity", "project", "agent"])
cleaned_data.head()
```
Now data is pivoted to create a new column for each type of view. After pivoting:
1. Mobile-web and mobile-app columns are combined into mobile
1. Columns are rename into more descriptive names
1. Aggregate columns for all pageview and pagecount views are created
1. Unused columns are dropped
```
# Pivot
pivot_data = cleaned_data.pivot(index = ["year", "month"], columns=["access"])
pivot_data.columns = pivot_data.columns.droplevel()
# Replace NaN with 0
pivot_data = pivot_data.fillna(0)
print(pivot_data.head())
# Combine mobil views
pivot_data["mobile"] = pivot_data["mobile-web"] + pivot_data["mobile-app"]
pivot_data = pivot_data.drop(columns = ["mobile-web", "mobile-app"])
# Rename and aggregate
pivot_data = pivot_data.rename(columns = {"desktop-site":"pagecount_desktop_views",
"mobile-site": "pagecount_mobile_views",
"desktop":"pageview_desktop_views",
"mobile":"pageview_mobile_views",
})
pivot_data["pagecount_all_views"] = pivot_data["pagecount_desktop_views"] + pivot_data["pagecount_mobile_views"]
pivot_data["pageview_all_views"] = pivot_data["pageview_desktop_views"] + pivot_data["pageview_mobile_views"]
pivot_data.head()
```
The data is converted to csv and saved.
```
pivot_data.to_csv('en-wikipedia_traffic_200712-202108.csv')
```
# Stage 3: ANALYSIS
Some minor processing is necessary to dispaly. The year and month are combined into a datetime index, and 0's are replaced with Nan to avoid plotting them. Values re-scaled to be in billions for easier interpretation.
```
display_data = pd.read_csv('en-wikipedia_traffic_200712-202108.csv')
# Set the index as dates
display_data.index = pd.to_datetime(display_data[['year', 'month']].assign(DAY=1))
display_data.drop(columns=["year", "month"], inplace=True)
# Repplce 0 with Nan
display_data = display_data.replace(0, np.nan)
# Rescale to billions
display_data = display_data / 1000000000
```
With some minor formatting, we see the trend in views. In 2016 we can see the difference in the new pageview API totals compared to the legacy, attributable to the amount of web crawlers accessing pages.
Also visible is the growth of mobile access from when it was first recorded in late 2014.
```
labels = ["Desktop", "Desktop (Legacy)", "Mobile (Legacy)", "Mobile", "Total (Legacy)", "Total"]
styles = ["g--","b--", "b:", "g:", "b", "g"]
widths = [3,3,3,3,6,6]
ax = display_data.plot(figsize=(15, 6), style=styles, title="English Wikipedia Traffic")
# Reorder legend
handles = plt.gca().get_legend_handles_labels()[0]
order = [5,0,3,4,1,2]
plt.legend([handles[idx] for idx in order],[labels[idx] for idx in order])
# Axis titles
ax.set_xlabel("Date")
ax.set_ylabel("Views in Billions")
plt.show()
# Save plot
ax.get_figure().savefig("english_wikipedia_traffic.png")
```
| github_jupyter |
# Working with MODFLOW-NWT v 1.1 option blocks
In MODFLOW-NWT an option block is present for the WEL file, UZF file, and SFR file. This block takes keyword arguments that are supplied in an option line in other versions of MODFLOW.
The `OptionBlock` class was created to provide combatibility with the MODFLOW-NWT option block and allow the user to easily edit values within the option block
```
import os
import sys
import platform
try:
import flopy
except:
fpth = os.path.abspath(os.path.join("..", ".."))
sys.path.append(fpth)
import flopy
from flopy.utils import OptionBlock
print(sys.version)
print("flopy version: {}".format(flopy.__version__))
load_ws = os.path.join("..", "data", "options", "sagehen")
model_ws = os.path.join("temp", "nwt_options", "output")
```
## Loading a MODFLOW-NWT model that has option block options
It is critical to set the `version` flag in `flopy.modflow.Modflow.load()` to `version='mfnwt'`
We are going to load a modified version of the Sagehen test problem from GSFLOW to illustrate compatibility
```
mfexe = "mfnwt"
if platform.system() == "Windows":
mfexe += ".exe"
ml = flopy.modflow.Modflow.load(
"sagehen.nam", model_ws=load_ws, exe_name=mfexe, version="mfnwt"
)
ml.change_model_ws(new_pth=model_ws)
ml.write_input()
success, buff = ml.run_model(silent=True)
if not success:
print("Something bad happened.")
```
## Let's look at the options attribute of the UZF object
The `uzf.options` attribute is an `OptionBlock` object. The representation of this object is the option block that will be written to output, which allows the user to easily check to make sure the block has the options they want.
```
uzf = ml.get_package("UZF")
uzf.options
```
The `OptionBlock` object also has attributes which correspond to the option names listed in the online guide to modflow
The user can call and edit the options within the option block
```
print(uzf.options.nosurfleak)
print(uzf.options.savefinf)
uzf.options.etsquare = False
uzf.options
uzf.options.etsquare = True
uzf.options
```
### The user can also see the single line representation of the options
```
uzf.options.single_line_options
```
### And the user can easily change to single line options writing
```
uzf.options.block = False
# write out only the uzf file
uzf_name = "uzf_opt.uzf"
uzf.write_file(os.path.join(model_ws, uzf_name))
```
Now let's examine the first few lines of the new UZF file
```
f = open(os.path.join(model_ws, uzf_name))
for ix, line in enumerate(f):
if ix == 3:
break
else:
print(line)
```
And let's load the new UZF file
```
uzf2 = flopy.modflow.ModflowUzf1.load(
os.path.join(model_ws, uzf_name), ml, check=False
)
```
### Now we can look at the options object, and check if it's block or line format
`block=False` indicates that options will be written as line format
```
print(uzf2.options)
print(uzf2.options.block)
```
### Finally we can convert back to block format
```
uzf2.options.block = True
uzf2.write_file(os.path.join(model_ws, uzf_name))
ml.remove_package("UZF")
uzf3 = flopy.modflow.ModflowUzf1.load(
os.path.join(model_ws, uzf_name), ml, check=False
)
print("\n")
print(uzf3.options)
print(uzf3.options.block)
```
## We can also look at the WEL object
```
wel = ml.get_package("WEL")
wel.options
```
Let's write this out as a single line option block and examine the first few lines
```
wel_name = "wel_opt.wel"
wel.options.block = False
wel.write_file(os.path.join(model_ws, wel_name))
f = open(os.path.join(model_ws, wel_name))
for ix, line in enumerate(f):
if ix == 4:
break
else:
print(line)
```
And we can load the new single line options WEL file and confirm that it is being read as an option line
```
ml.remove_package("WEL")
wel2 = flopy.modflow.ModflowWel.load(
os.path.join(model_ws, wel_name), ml, nper=ml.nper, check=False
)
wel2.options
wel2.options.block
```
# Building an OptionBlock from scratch
The user can also build an `OptionBlock` object from scratch to add to a `ModflowSfr2`, `ModflowUzf1`, or `ModflowWel` file.
The `OptionBlock` class has two required parameters and one optional parameter
`option_line`: a one line, string based representation of the options
`package`: a modflow package object
`block`: boolean flag for line based or block based options
```
opt_line = "specify 0.1 20"
options = OptionBlock(opt_line, flopy.modflow.ModflowWel, block=True)
options
```
from here we can set the noprint flag by using `options.noprint`
```
options.noprint = True
```
and the user can also add auxillary variables by using `options.auxillary`
```
options.auxillary = ["aux", "iface"]
```
### Now we can create a new wel file using this `OptionBlock`
and write it to output
```
wel3 = flopy.modflow.ModflowWel(
ml,
stress_period_data=wel.stress_period_data,
options=options,
unitnumber=99,
)
wel3.write_file(os.path.join(model_ws, wel_name))
```
And now let's examine the first few lines of the file
```
f = open(os.path.join(model_ws, wel_name))
for ix, line in enumerate(f):
if ix == 8:
break
else:
print(line)
```
We can see that everything that the OptionBlock class writes out options in the correct location.
### The user can also switch the options over to option line style and write out the output too!
```
wel3.options.block = False
wel3.write_file(os.path.join(model_ws, wel_name))
f = open(os.path.join(model_ws, wel_name))
for ix, line in enumerate(f):
if ix == 6:
break
else:
print(line)
```
| github_jupyter |
```
from __future__ import print_function
import collections
import numpy as np
import pandas as pd
import pickle
import sklearn
from keras.datasets import imdb
from keras.models import Sequential
from keras.layers import Dense, Dropout
from keras.layers import LSTM
from keras.layers.embeddings import Embedding
from keras.preprocessing import sequence
from keras.regularizers import L1L2
from sklearn.model_selection import train_test_split
# fix random seed for reproducibility
np.random.seed(7)
from pprint import pprint
# Global config variables
model_name = "streeteye_lstm"
#data_file = "lstm_dump_test.txt"
data_file = "dump_2017_words.txt"
checkpoint_dir = "/home/ubuntu/mount/Notebooks/checkpoints"
tensorboard_dir ="/home/ubuntu/mount/Notebooks/tensorboard"
############################################################
# 1. load data
############################################################
# load dataset
print("Loading data...")
data=[]
y=[]
# count words
c = collections.Counter()
with open(data_file, "r") as infile:
for line in infile:
l = line.rstrip('\n').split(",")
label = l.pop(0)
# skip empty headlines
if len(l[0]) == 0:
continue
if '' in l:
l = [w for w in l if w]
data.append(l)
y.append(label)
c.update(l)
print("Loaded data.")
# create a list of top words
vocabulary_size = 10000 # set this to have ~20 for least popular
count = [['UNK', -1]]
count.extend(c.most_common(vocabulary_size - 1))
print(count[:10])
print(count[-10:])
dictionary = dict()
# map words into a dict of ints
for word, _ in count:
dictionary[word] = len(dictionary)
data_embeddings=[]
unk_count = 0
for obs in data:
embedlist = []
for word in obs:
if word in dictionary:
index = dictionary[word]
else:
index = 0 # dictionary['UNK']
unk_count = unk_count + 1
embedlist.append(index)
data_embeddings.append(embedlist)
count[0][1] = unk_count
reverse_dictionary = dict(zip(dictionary.values(), dictionary.keys()))
print(dictionary['trump'])
print(reverse_dictionary[3])
%matplotlib inline
ls = (map(len, data_embeddings))
pd.DataFrame(ls).hist()
MAX_LENGTH = 120
X = sequence.pad_sequences(data_embeddings, maxlen=MAX_LENGTH)
X[0]
X.shape
y=np.array(np.float32(y))
y=y.reshape((y.shape[0],1))
print(y.shape)
num_labels=1
num_obs, num_features = X.shape
print("Observations: %d\nFeatures: %d" % (num_obs, num_features))
# split into training, xval, test, 60/20/20
print("Split into training, temp")
X_train, X_temp, y_train, y_temp = train_test_split(X, y, test_size=0.4)
print("Split into xval, test")
X_xval, X_test, y_xval, y_test = train_test_split(X_temp, y_temp, test_size=0.5)
print("Training set")
print(X_train.shape)
print("Xval set")
print(X_xval.shape)
print("Test set")
print(X_test.shape)
num_training_samples = X_train.shape[0]
num_xval_samples = X_xval.shape[0]
num_test_samples = X_test.shape[0]
print ("\nTraining observations: %d \nXval observations: %d \nTest observations: %d\n" % (num_training_samples, num_xval_samples, num_test_samples))
# initialize embeddings to Google vals
pkl_file = open('embeddings.pkl', 'rb')
embeddings_dict, embeddings_reverse_dict, embeddings_data = pickle.load(pkl_file)
EMBEDDING_DIM=300
embedding_matrix = np.zeros((len(dictionary) + 1, EMBEDDING_DIM))
count = 0
for word, i in dictionary.items():
#print(word)
embed_i = embeddings_dict.get(word)
if embed_i is not None:
embedding_vector = embeddings_data[i]
count +=1
embedding_matrix[i] = embedding_vector
print("initialized %d embeddings" % count)
# function to generate model
def create_model(lstm_size=30, lstm_reg_penalty=0.0, sigmoid_dropout=(1.0/3.0), sigmoid_reg_penalty=0.0001):
# create model
model = Sequential()
model.add(Embedding(len(dictionary) + 1,
embedding_vector_length,
weights=[embedding_matrix],
input_length=MAX_LENGTH,
trainable=False))
# LSTM with lstm_size units
model.add(LSTM(lstm_size,
kernel_regularizer=L1L2(l1=lstm_reg_penalty, l2=lstm_reg_penalty)))
model.add(Dropout(sigmoid_dropout))
model.add(Dense(1,
activation='sigmoid',
kernel_initializer='TruncatedNormal',
kernel_regularizer=L1L2(l1=sigmoid_reg_penalty, l2=sigmoid_reg_penalty)))
model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
print(model.summary())
return model
def selectThreshold (logits, labels, beta=(2.0/3)):
# return threshold, f-score that yields best F-score
# predict using true if >= threshold
precision, recall, thresholds = sklearn.metrics.precision_recall_curve(labels, logits)
bb = beta**2
f1_scores = (1 + bb) * precision * recall / (bb * precision + recall)
best_index = np.argmax(f1_scores)
best_threshold = thresholds[best_index]
best_score = f1_scores[best_index]
return (best_threshold, best_score)
# create the model
embedding_vector_length = EMBEDDING_DIM
for sig_reg_penalty in [0.00003]:
for dropout in [0.5]:
for lstm_units in [16,]:
#, 32, 64]:
for lstm_reg_penalty in [0.00000,]:
#0.000001, 0.000003, 0.00001, 0.00003]:
model = create_model(lstm_size=lstm_units,
lstm_reg_penalty=lstm_reg_penalty,
sigmoid_dropout=dropout,
sigmoid_reg_penalty=sig_reg_penalty)
print ("LSTM units %d" % lstm_units)
print ("LSTM reg_penalty %.8f" % lstm_reg_penalty)
print ("Sigmoid dropout %.4f" % dropout)
print ("Sigmoid reg_penalty %.8f" % sig_reg_penalty)
model.fit(X_train, y_train, validation_data=(X_xval, y_xval), epochs=100, batch_size=128)
y_train_prob = model.predict(X_train)
beta=(2.0/3.0) # penalize false positives more than false negatives
thresh, score = selectThreshold(y_train_prob, y_train, beta=beta)
y_train_pred = y_train_prob >= thresh
print("Train Accuracy %.3f, Train F1 %.3f, f_score %.3f (beta %.3f)" %
(sklearn.metrics.accuracy_score(y_train_pred, y_train),
sklearn.metrics.f1_score(y_train_pred, y_train),
score, beta))
print(sklearn.metrics.confusion_matrix(y_train_pred, y_train))
y_xval_prob = model.predict(X_xval)
thresh, score = selectThreshold(y_xval_prob, y_xval, beta=beta)
y_xval_pred = y_xval_prob >= thresh
print ("LSTM units %d" % lstm_units)
print ("LSTM reg_penalty %.8f" % lstm_reg_penalty)
print ("Sigmoid dropout %.4f" % dropout)
print ("Sigmoid reg_penalty %.8f" % sig_reg_penalty)
print("Xval Accuracy %.3f, Xval F1 %.3f, f_score %.3f (beta %.3f)" %
(sklearn.metrics.accuracy_score(y_xval_pred, y_xval),
sklearn.metrics.f1_score(y_xval_pred, y_xval),
score, beta))
confusion_matrix = sklearn.metrics.confusion_matrix(y_xval_pred, y_xval)
print(confusion_matrix)
false_positive = confusion_matrix[1][0]
false_negative = confusion_matrix[0][1]
raw_score = (2.0*false_positive + false_negative) / np.sum(confusion_matrix)
print ("Raw score %f" % raw_score)
y_train_prob = model.predict(X_train)
beta=(2.0/3.0) # penalize false positives more than false negatives
thresh, score = selectThreshold(y_train_prob, y_train, beta=beta)
y_train_pred = y_train_prob >= thresh
print("Train Accuracy %.3f, Train F1 %.3f, f_score %.3f (beta %.3f)" %
(sklearn.metrics.accuracy_score(y_train_pred, y_train),
sklearn.metrics.f1_score(y_train_pred, y_train),
score, beta))
print(sklearn.metrics.confusion_matrix(y_train_pred, y_train))
y_xval_prob = model.predict(X_xval)
thresh, score = selectThreshold(y_xval_prob, y_xval, beta=beta)
y_xval_pred = y_xval_prob >= thresh
print("Xval Accuracy %.3f, Xval F1 %.3f, f_score %.3f (beta %.3f)" %
(sklearn.metrics.accuracy_score(y_xval_pred, y_xval),
sklearn.metrics.f1_score(y_xval_pred, y_xval),
score, beta))
print(sklearn.metrics.confusion_matrix(y_xval_pred, y_xval))
y_test_prob = model.predict(X_test)
beta=(2.0/3.0) # penalize false positives more than false negatives
y_test_pred = y_test_prob >= thresh
print("Test Accuracy %.3f, Test F1 %.3f, f_score %.3f (beta %.3f)" %
(sklearn.metrics.accuracy_score(y_test_pred, y_test),
sklearn.metrics.f1_score(y_test_pred, y_test),
score, beta))
print(sklearn.metrics.confusion_matrix(y_test_pred, y_test))
```
| github_jupyter |
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc"><ul class="toc-item"><li><span><a href="#Matplotlib" data-toc-modified-id="Matplotlib-1"><span class="toc-item-num">1 </span>Matplotlib</a></span><ul class="toc-item"><li><span><a href="#Customization" data-toc-modified-id="Customization-1.1"><span class="toc-item-num">1.1 </span>Customization</a></span></li></ul></li><li><span><a href="#subplot" data-toc-modified-id="subplot-2"><span class="toc-item-num">2 </span>subplot</a></span></li></ul></div>
# Intermediate Python for Data Science
## Matplotlib
- source: https://www.datacamp.com/courses/intermediate-python-for-data-science
- color code: https://matplotlib.org/examples/color/named_colors.html
```
# Quick cheat sheet
# to change plot size
plt.figure(figsize=(20,8))
'''Line Plot'''
# Print the last item from years and populations
print(year[-1])
print(pop[-1])
# Import matplotlib.pyplot as plt
import matplotlib.pyplot as plt
# Make a line plot: year on the x-axis, pop on the y-axis
plt.plot(year, pop)
# Display the plot with plt.show()
plt.show()
# Print the last item of gdp_cap and life_exp
print(gdp_cap[-1])
print(life_exp[-1])
# Make a line plot, gdp_cap on the x-axis, life_exp on the y-axis
plt.plot(gdp_cap, life_exp)
# Display the plot
plt.show()
'''Scatter Plot'''
# Change the line plot below to a scatter plot
plt.scatter(gdp_cap, life_exp)
# Put the x-axis on a logarithmic scale
plt.xscale('log')
# Show plot
plt.show()
'''Scatter Plot'''
# Import package
import matplotlib.pyplot as plt
# Build Scatter plot
plt.scatter(pop, life_exp)
# Show plot
plt.show()
'''Histogram'''
# Create histogram of life_exp data
plt.hist(life_exp)
# Display histogram
plt.show()
'''Histogram bins'''
# Build histogram with 5 bins
plt.hist(life_exp, bins = 5)
# Show and clear plot
plt.show()
plt.clf() # cleans it up again so you can start afresh.
# Build histogram with 20 bins
plt.hist(life_exp, bins = 20)
# Show and clear plot again
plt.show()
plt.clf()
'''Histogram compare'''
# Histogram of life_exp, 15 bins
plt.hist(life_exp, bins = 15)
# Show and clear plot
plt.show()
plt.clf()
# Histogram of life_exp1950, 15 bins
plt.hist(life_exp1950, bins = 15)
# Show and clear plot again
plt.show()
plt.clf()
```
### Customization
```
'''Label'''
# Basic scatter plot, log scale
plt.scatter(gdp_cap, life_exp)
plt.xscale('log')
# Strings
xlab = 'GDP per Capita [in USD]'
ylab = 'Life Expectancy [in years]'
title = 'World Development in 2007'
# Add axis labels
plt.xlabel(xlab)
plt.ylabel(ylab)
# Add title
plt.title(title)
# After customizing, display the plot
plt.show()
'''Ticks'''
# Scatter plot
plt.scatter(gdp_cap, life_exp)
# Previous customizations
plt.xscale('log')
plt.xlabel('GDP per Capita [in USD]')
plt.ylabel('Life Expectancy [in years]')
plt.title('World Development in 2007')
# Definition of tick_val and tick_lab
tick_val = [1000,10000,100000]
tick_lab = ['1k','10k','100k']
# Adapt the ticks on the x-axis
plt.xticks(tick_val, tick_lab)
# After customizing, display the plot
plt.show()
'''Sizes
Wouldn't it be nice if the size of the dots corresponds to the population?
'''
# Import numpy as np
import numpy as np
# Store pop as a numpy array: np_pop
np_pop = np.array(pop)
# Double np_pop
np_pop = np_pop * 2
# Update: set s argument to np_pop # s is size
plt.scatter(gdp_cap, life_exp, s = np_pop)
# Previous customizations
plt.xscale('log')
plt.xlabel('GDP per Capita [in USD]')
plt.ylabel('Life Expectancy [in years]')
plt.title('World Development in 2007')
plt.xticks([1000, 10000, 100000],['1k', '10k', '100k'])
# Display the plot
plt.show()
'''Colors
The next step is making the plot more colorful! To do this, a list col has been created for you. It's a list with a color for each corresponding country, depending on the continent the country is part of.
How did we make the list col you ask? The Gapminder data contains a list continent with the continent each country belongs to. A dictionary is constructed that maps continents onto colors:
'''
dict = {
'Asia':'red',
'Europe':'green',
'Africa':'blue',
'Americas':'yellow',
'Oceania':'black'
}
# c = color, alpha = opacity
# Specify c and alpha inside plt.scatter()
plt.scatter(x = gdp_cap, y = life_exp, s = np.array(pop) * 2, c = col, alpha = 0.8)
# Previous customizations
plt.xscale('log')
plt.xlabel('GDP per Capita [in USD]')
plt.ylabel('Life Expectancy [in years]')
plt.title('World Development in 2007')
plt.xticks([1000,10000,100000], ['1k','10k','100k'])
# Show the plot
plt.show()
'''Additional Customizations'''
# Scatter plot
plt.scatter(x = gdp_cap, y = life_exp, s = np.array(pop) * 2, c = col, alpha = 0.8)
# Previous customizations
plt.xscale('log')
plt.xlabel('GDP per Capita [in USD]')
plt.ylabel('Life Expectancy [in years]')
plt.title('World Development in 2007')
plt.xticks([1000,10000,100000], ['1k','10k','100k'])
# Additional customizations
plt.text(1550, 71, 'India')
plt.text(5700, 80, 'China')
# Add grid() call
plt.grid(True)
# Show the plot
plt.show()
from sklearn.datasets import load_iris
data = load_iris()
data.target[[10, 25, 50]]
list(data.target_names)
```
## subplot
source: https://matplotlib.org/examples/pylab_examples/subplot_demo.html
```
# subplot(nrows, ncols, plot_number)
import numpy as np
import matplotlib.pyplot as plt
x1 = np.linspace(0.0, 5.0)
x2 = np.linspace(0.0, 2.0)
y1 = np.cos(2 * np.pi * x1) * np.exp(-x1)
y2 = np.cos(2 * np.pi * x2)
plt.subplot(2, 1, 1)
plt.plot(x1, y1, 'ko-')
plt.title('A tale of 2 subplots')
plt.ylabel('Damped oscillation')
plt.subplot(2, 1, 2)
plt.plot(x2, y2, 'r.-')
plt.xlabel('time (s)')
plt.ylabel('Undamped')
plt.show()
plt.subplot(2, 1, 1)
plt.plot(x1, y1, 'ko-')
plt.title('A tale of 2 subplots')
plt.ylabel('Damped oscillation')
plt.subplot(2, 1, 2)
plt.plot(x2, y2, 'r.-')
plt.xlabel('time (s)')
plt.ylabel('Undamped')
plt.show()
plt.subplots(2, 2, sharex=True, sharey=True)
plt.show()
x = np.linspace(0, 2*np.pi, 400)
y = np.sin(x**2)
fig, axes = plt.subplots(1,2, sharey=True)
axes[0].plot(x, y)
axes[1].scatter(x, y)
plt.show()
# Two subplots, unpack the axes array immediately
f, (ax1, ax2) = plt.subplots(1, 2, sharey=True)
ax1.plot(x, y)
ax1.set_title('Sharing Y axis')
ax2.scatter(x, y)
plt.show()
fig, axes = plt.subplots(1,3, sharey=True, sharex=True)
for i in range(3):
axes[i].scatter(center[i],xn)
axes[i].set_title('Cluster ' + str(i+1))
axes[i].grid(True)
plt.yticks(xn,var)
plt.subplots_adjust(wspace=0, hspace=0)
#plt.grid(True)
plt.show()
```
| github_jupyter |
```
import seaborn as sns
import pandas as pd
import numpy as np
import altair as alt
from markdown import markdown
from IPython.display import Markdown
from ipywidgets.widgets import HTML, Tab
from ipywidgets import widgets
from datetime import timedelta
from matplotlib import pyplot as plt
import os.path as op
from mod import load_data, alt_theme
def author_url(author):
return f"https://github.com/{author}"
# Parameters
fmt_date = "{:%Y-%m-%d}"
n_days = 90
start_date = fmt_date.format(pd.datetime.today() - timedelta(days=n_days))
end_date = fmt_date.format(pd.datetime.today())
renderer = "jupyterlab"
github_orgs = ["jupyterhub", "jupyter", "jupyterlab", "jupyter-widgets", "ipython", "binder-examples", "nteract"]
# Parameters
renderer = "kaggle"
start_date = "2019-02-01"
end_date = "2019-03-01"
comments, issues, prs = load_data('../data/')
bot_names = pd.read_csv('bot_names.csv')['names'].tolist()
comments = comments.query('author not in @bot_names').drop_duplicates()
issues = issues.query('author not in @bot_names').drop_duplicates()
prs = prs.query('author not in @bot_names').drop_duplicates()
# Only keep the dates we want
comments = comments.query('updatedAt > @start_date and updatedAt < @end_date')
issues = issues.query('updatedAt > @start_date and updatedAt < @end_date')
prs = prs.query('updatedAt > @start_date and updatedAt < @end_date')
alt.renderers.enable(renderer);
alt.themes.register('my_theme', alt_theme)
alt.themes.enable("my_theme")
# Information about out time window
time_delta = pd.to_datetime(end_date) - pd.to_datetime(start_date)
n_days = time_delta.days
# Information about the data we loaded
github_orgs = comments['org'].unique()
```
# GitHub activity
Jupyter also has lots of activity across GitHub repositories. The following sections contain
overviews of recent activity across the following GitHub organizations:
```
# Define colors we'll use for GitHub membership
author_types = ['MEMBER', 'CONTRIBUTOR', 'COLLABORATOR', "NONE"]
author_palette = sns.palettes.blend_palette(["lightgrey", "lightgreen", "darkgreen"], 4)
author_colors = ["rgb({}, {}, {}, {})".format(*(ii*256)) for ii in author_palette]
author_color_dict = {key: val for key, val in zip(author_types, author_palette)}
orgs_md = []
for org in github_orgs:
orgs_md.append(f'* [github.com/{org}](https://github.com/{org})')
Markdown('\n'.join(orgs_md))
Markdown(f"Showing GitHub activity from **{start_date}** to **{end_date}**")
```
## List of all contributors per organization
First, we'll list each contributor that has contributed to each organization in the last several days.
Contributions to open source projects are diverse, and involve much more than just contributing code and
code review. Thanks to everybody in the Jupyter communities for all that they do.
```
n_plot = 5
tabs = widgets.Tab(children=[])
for ii, org in enumerate(github_orgs):
authors_comments = comments.query('org == @org')['author']
authors_prs = prs.query('org == @org')['author']
unique_participants = np.unique(np.hstack([authors_comments.values, authors_prs.values]).astype(str)).tolist()
unique_participants.sort(key=lambda a: a.lower())
all_participants = [f"[{participant}](https://github.com/{participant})" for participant in unique_participants]
participants_md = " | ".join(all_participants)
md_html = HTML("<center>{}</center>".format(markdown(participants_md)))
children = list(tabs.children)
children.append(md_html)
tabs.children = tuple(children)
tabs.set_title(ii, org)
display(Markdown(f"All participants across issues and pull requests in each org in the last {n_days} days"))
display(tabs)
```
## Merged Pull requests
Here's an analysis of **merged pull requests** across each of the repositories in the Jupyter
ecosystem.
```
merged = prs.query('state == "MERGED" and closedAt > @start_date and closedAt < @end_date')
prs_by_repo = merged.groupby(['org', 'repo']).count()['author'].reset_index().sort_values(['org', 'author'], ascending=False)
alt.Chart(data=prs_by_repo, title=f"Merged PRs in the last {n_days} days").mark_bar().encode(
x=alt.X('repo', sort=prs_by_repo['repo'].values.tolist()),
y='author',
color='org'
)
```
### A list of merged PRs by project
Below is a tabbed readout of recently-merged PRs. Check out the title to get an idea for what they
implemented, and be sure to thank the PR author for their hard work!
```
tabs = widgets.Tab(children=[])
merged_by = {}
pr_by = {}
for ii, (org, idata) in enumerate(merged.groupby('org')):
issue_md = []
issue_md.append(f"#### Closed PRs for org: `{org}`")
issue_md.append("")
for (org, repo), prs in idata.groupby(['org', 'repo']):
issue_md.append(f"##### [{org}/{repo}](https://github.com/{org}/{repo})")
for _, pr in prs.iterrows():
user_name = pr['author']
user_url = author_url(user_name)
pr_number = pr['number']
pr_html = pr['url']
pr_title = pr['title']
pr_closedby = pr['mergedBy']
pr_closedby_url = f"https://github.com/{pr_closedby}"
if user_name not in pr_by:
pr_by[user_name] = 1
else:
pr_by[user_name] += 1
if pr_closedby not in merged_by:
merged_by[pr_closedby] = 1
else:
merged_by[pr_closedby] += 1
text = f"* [(#{pr_number})]({pr_html}): _{pr_title}_ by **[@{user_name}]({user_url})** merged by **[@{pr_closedby}]({pr_closedby_url})**"
issue_md.append(text)
issue_md.append('')
markdown_html = markdown('\n'.join(issue_md))
children = list(tabs.children)
children.append(HTML(markdown_html))
tabs.children = tuple(children)
tabs.set_title(ii, org)
tabs
```
### Authoring and merging stats by repository
Let's see who has been doing most of the PR authoring and merging. The PR author is generally the
person that implemented a change in the repository (code, documentation, etc). The PR merger is
the person that "pressed the green button" and got the change into the main codebase.
```
# Prep our merging DF
merged_by_repo = merged.groupby(['org', 'repo', 'author'], as_index=False).agg({'id': 'count', 'authorAssociation': 'first'}).rename(columns={'id': "authored", 'author': 'username'})
closed_by_repo = merged.groupby(['org', 'repo', 'mergedBy']).count()['id'].reset_index().rename(columns={'id': "closed", "mergedBy": "username"})
n_plot = 50
charts = []
for ii, (iorg, idata) in enumerate(merged_by_repo.replace(np.nan, 0).groupby(['org'])):
title = f"PR authors for {iorg} in the last {n_days} days"
idata = idata.groupby('username', as_index=False).agg({'authored': 'sum', 'authorAssociation': 'first'})
idata = idata.sort_values('authored', ascending=False).head(n_plot)
ch = alt.Chart(data=idata, width=1000, title=title).mark_bar().encode(
x='username',
y='authored',
color=alt.Color('authorAssociation', scale=alt.Scale(domain=author_types, range=author_colors))
)
charts.append(ch)
alt.hconcat(*charts)
charts = []
for ii, (iorg, idata) in enumerate(closed_by_repo.replace(np.nan, 0).groupby(['org'])):
title = f"Merges for {iorg} in the last {n_days} days"
ch = alt.Chart(data=idata, width=1000, title=title).mark_bar().encode(
x='username',
y='closed',
)
charts.append(ch)
alt.hconcat(*charts)
```
## Issues
Issues are **conversations** that happen on our GitHub repositories. Here's an
analysis of issues across the Jupyter organizations.
```
created = issues.query('state == "OPEN" and createdAt > @start_date and createdAt < @end_date')
closed = issues.query('state == "CLOSED" and closedAt > @start_date and closedAt < @end_date')
created_counts = created.groupby(['org', 'repo']).count()['number'].reset_index()
created_counts['org/repo'] = created_counts.apply(lambda a: a['org'] + '/' + a['repo'], axis=1)
sorted_vals = created_counts.sort_values(['org', 'number'], ascending=False)['repo'].values
alt.Chart(data=created_counts, title=f"Issues created in the last {n_days} days").mark_bar().encode(
x=alt.X('repo', sort=alt.Sort(sorted_vals.tolist())),
y='number',
color='org',
)
closed_counts = closed.groupby(['org', 'repo']).count()['number'].reset_index()
closed_counts['org/repo'] = closed_counts.apply(lambda a: a['org'] + '/' + a['repo'], axis=1)
sorted_vals = closed_counts.sort_values(['org', 'number'], ascending=False)['repo'].values
alt.Chart(data=closed_counts, title=f"Issues closed in the last {n_days} days").mark_bar().encode(
x=alt.X('repo', sort=alt.Sort(sorted_vals.tolist())),
y='number',
color='org',
)
created_closed = pd.merge(created_counts.rename(columns={'number': 'created'}).drop(columns='org/repo'),
closed_counts.rename(columns={'number': 'closed'}).drop(columns='org/repo'),
on=['org', 'repo'], how='outer')
created_closed = pd.melt(created_closed, id_vars=['org', 'repo'], var_name="kind", value_name="count").replace(np.nan, 0)
charts = []
for org in github_orgs:
# Pick the top 10 repositories
this_issues = created_closed.query('org == @org')
top_repos = this_issues.groupby(['repo']).sum().sort_values(by='count', ascending=False).head(10).index
ch = alt.Chart(this_issues.query('repo in @top_repos'), width=120).mark_bar().encode(
x=alt.X("kind", axis=alt.Axis(labelFontSize=15, title="")),
y=alt.Y('count', axis=alt.Axis(titleFontSize=15, labelFontSize=12)),
color='kind',
column=alt.Column("repo", header=alt.Header(title=f"Issue activity, last {n_days} days for {org}", titleFontSize=15, labelFontSize=12))
)
charts.append(ch)
alt.hconcat(*charts)
# Set to datetime
for kind in ['createdAt', 'closedAt']:
closed.loc[:, kind] = pd.to_datetime(closed[kind])
closed.loc[:, 'time_open'] = closed['closedAt'] - closed['createdAt']
closed.loc[:, 'time_open'] = closed['time_open'].dt.total_seconds()
time_open = closed.groupby(['org', 'repo']).agg({'time_open': 'median'}).reset_index()
time_open['time_open'] = time_open['time_open'] / (60 * 60 * 24)
time_open['org/repo'] = time_open.apply(lambda a: a['org'] + '/' + a['repo'], axis=1)
sorted_vals = time_open.sort_values(['org', 'time_open'], ascending=False)['repo'].values
alt.Chart(data=time_open, title=f"Time to close for issues closed in the last {n_days} days").mark_bar().encode(
x=alt.X('repo', sort=alt.Sort(sorted_vals.tolist())),
y=alt.Y('time_open', title="Median Days Open"),
color='org',
)
```
### A list of recent issues
Below is a list of issues with recent activity in each repository. If they seem of interest
to you, click on their links and jump in to participate!
```
# Add comment count data to issues and PRs
comment_counts = (
comments
.query("createdAt > @start_date and createdAt < @end_date")
.groupby(['org', 'repo', 'issue_id'])
.count().iloc[:, 0].to_frame()
)
comment_counts.columns = ['n_comments']
comment_counts = comment_counts.reset_index()
n_plot = 5
tabs = widgets.Tab(children=[])
for ii, (org, idata) in enumerate(comment_counts.groupby('org')):
issue_md = []
issue_md.append(f"#### {org}")
issue_md.append("")
for repo, i_issues in idata.groupby('repo'):
issue_md.append(f"##### [{org}/{repo}](https://github.com/{org}/{repo})")
top_issues = i_issues.sort_values('n_comments', ascending=False).head(n_plot)
top_issue_list = pd.merge(issues, top_issues, left_on=['org', 'repo', 'number'], right_on=['org', 'repo', 'issue_id'])
for _, issue in top_issue_list.sort_values('n_comments', ascending=False).head(n_plot).iterrows():
user_name = issue['author']
user_url = author_url(user_name)
issue_number = issue['number']
issue_html = issue['url']
issue_title = issue['title']
text = f"* [(#{issue_number})]({issue_html}): _{issue_title}_ by **[@{user_name}]({user_url})**"
issue_md.append(text)
issue_md.append('')
md_html = HTML(markdown('\n'.join(issue_md)))
children = list(tabs.children)
children.append(HTML(markdown('\n'.join(issue_md))))
tabs.children = tuple(children)
tabs.set_title(ii, org)
display(Markdown(f"Here are the top {n_plot} active issues in each repository in the last {n_days} days"))
display(tabs)
```
## Commenters across repositories
These are commenters across all issues and pull requests in the last several days.
These are colored by the commenter's association with the organization. For information
about what these associations mean, [see this StackOverflow post](https://stackoverflow.com/a/28866914/1927102).
```
commentors = (
comments
.query("createdAt > @start_date and createdAt < @end_date")
.groupby(['org', 'repo', 'author', 'authorAssociation'])
.count().rename(columns={'id': 'count'})['count']
.reset_index()
.sort_values(['org', 'count'], ascending=False)
)
n_plot = 50
charts = []
for ii, (iorg, idata) in enumerate(commentors.groupby(['org'])):
title = f"Top {n_plot} commentors for {iorg} in the last {n_days} days"
idata = idata.groupby('author', as_index=False).agg({'count': 'sum', 'authorAssociation': 'first'})
idata = idata.sort_values('count', ascending=False).head(n_plot)
ch = alt.Chart(data=idata.head(n_plot), width=1000, title=title).mark_bar().encode(
x='author',
y='count',
color=alt.Color('authorAssociation', scale=alt.Scale(domain=author_types, range=author_colors))
)
charts.append(ch)
alt.hconcat(*charts)
```
## First responders
First responders are the first people to respond to a new issue in one of the repositories.
The following plots show first responders for recently-created issues.
```
first_comments = []
for (org, repo, issue_id), i_comments in comments.groupby(['org', 'repo', 'issue_id']):
ix_min = pd.to_datetime(i_comments['createdAt']).idxmin()
first_comment = i_comments.loc[ix_min]
if isinstance(first_comment, pd.DataFrame):
first_comment = first_comment.iloc[0]
first_comments.append(first_comment)
first_comments = pd.concat(first_comments, axis=1).T
first_responder_counts = first_comments.groupby(['org', 'author', 'authorAssociation'], as_index=False).\
count().rename(columns={'id': 'n_first_responses'}).sort_values(['org', 'n_first_responses'], ascending=False)
n_plot = 50
charts = []
for ii, (iorg, idata) in enumerate(first_responder_counts.groupby(['org'])):
title = f"Top {n_plot} first responders for {iorg} in the last {n_days} days"
idata = idata.groupby('author', as_index=False).agg({'n_first_responses': 'sum', 'authorAssociation': 'first'})
idata = idata.sort_values('n_first_responses', ascending=False).head(n_plot)
ch = alt.Chart(data=idata.head(n_plot), width=1000, title=title).mark_bar().encode(
x='author',
y='n_first_responses',
color=alt.Color('authorAssociation', scale=alt.Scale(domain=author_types, range=author_colors))
)
charts.append(ch)
alt.hconcat(*charts)
%%html
<script src="https://cdn.rawgit.com/parente/4c3e6936d0d7a46fd071/raw/65b816fb9bdd3c28b4ddf3af602bfd6015486383/code_toggle.js"></script>
```
| github_jupyter |
# Speed from Position Data
In this Notebook you'll work with data just like the data you'll be using in the final project for this course. That data comes from CSVs that looks like this:
| timestamp | displacement | yaw_rate | acceleration |
| :-------: | :----------: | :------: | :----------: |
| 0.0 | 0 | 0.0 | 0.0 |
| 0.25 | 0.0 | 0.0 | 19.6 |
| 0.5 | 1.225 | 0.0 | 19.6 |
| 0.75 | 3.675 | 0.0 | 19.6 |
| 1.0 | 7.35 | 0.0 | 19.6 |
| 1.25 | 12.25 | 0.0 | 0.0 |
| 1.5 | 17.15 | -2.82901631903 | 0.0 |
| 1.75 | 22.05 | -2.82901631903 | 0.0 |
| 2.0 | 26.95 | -2.82901631903 | 0.0 |
| 2.25 | 31.85 | -2.82901631903 | 0.0 |
```
from helpers import process_data
from matplotlib import pyplot as plt
PARALLEL_PARK_DATA = process_data("parallel_park.pickle")
# This is what the first few entries in the parallel
# park data look like.
PARALLEL_PARK_DATA[:5]
# In this exercise we'll be differentiating (taking the
# derivative of) displacement data. This will require
# using only the first two columns of this data.
timestamps = [row[0] for row in PARALLEL_PARK_DATA]
displacements = [row[1] for row in PARALLEL_PARK_DATA]
# You'll use these data in the next lesson on integration
# You can ignore them for now.
yaw_rates = [row[2] for row in PARALLEL_PARK_DATA]
accelerations = [row[3] for row in PARALLEL_PARK_DATA]
plt.title("Displacement vs Time while Parallel Parking")
plt.xlabel("Time (seconds)")
plt.ylabel("Displacement (meters)")
plt.scatter(timestamps, displacements)
plt.show()
```
In the graph above, you can see displacement vs time data for a car as it parallel parks. Note that backwards motion winds back the odometer and reduces displacement (this isn't actually how odometers work on modern cars. Sorry Ferris Bueller)
Note how for approximately 4 seconds the motion is backwards and then for the last two the car goes forwards.
Let's look at some data somewhere in the middle of this trajectory
```
print(timestamps[20:22])
print(displacements[20:22])
```
So you can see that at $t=1.25$ the car has displacement $x=-1.40875$ and at $t=1.3125$ the car has displacement $x=-1.53125$
This means we could calculate the speed / slope as follows:
$$\text{slope} = \frac{\text{vertical change}}{\text{horizontal change}} = \frac{\Delta x}{\Delta t}$$
and for the numbers I just mentioned this would mean:
$$\frac{\Delta x}{\Delta t} = \frac{-1.53125 - -1.40875}{1.3125 - 1.25} = \frac{-0.1225 \text{ meters}}{0.0625\text{ seconds}} = -1.96 \frac{m}{s}$$
So I can say the following:
> Between $t=1.25$ and $t=1.3125$ the vehicle had an **average speed** of **-1.96 meters per second**
I could make this same calculation in code as follows
```
delta_x = displacements[21] - displacements[20]
delta_t = timestamps[21] - timestamps[20]
slope = delta_x / delta_t
print(slope)
```
Earlier in this lesson you worked with truly continuous functions. In that situation you could make $\Delta t$ as small as you wanted!
But now we have real data, which means the size of $\Delta t$ is dictated by how frequently we made measurements of displacement. In this case it looks like subsequent measurements are separated by
$$\Delta t = 0.0625 \text{ seconds}$$
In the `get_derivative_from_data` function below, I demonstrate how to "take a derivative" of real data. Read through this code and understand how it works: in the next notebook you'll be asked to reproduce this code yourself.
```
def get_derivative_from_data(position_data, time_data):
"""
Calculates a list of speeds from position_data and
time_data.
Arguments:
position_data - a list of values corresponding to
vehicle position
time_data - a list of values (equal in length to
position_data) which give timestamps for each
position measurement
Returns:
speeds - a list of values (which is shorter
by ONE than the input lists) of speeds.
"""
# 1. Check to make sure the input lists have same length
if len(position_data) != len(time_data):
raise(ValueError, "Data sets must have same length")
# 2. Prepare empty list of speeds
speeds = []
# 3. Get first values for position and time
previous_position = position_data[0]
previous_time = time_data[0]
# 4. Begin loop through all data EXCEPT first entry
for i in range(1, len(position_data)):
# 5. get position and time data for this timestamp
position = position_data[i]
time = time_data[i]
# 6. Calculate delta_x and delta_t
delta_x = position - previous_position
delta_t = time - previous_time
# 7. Speed is slope. Calculate it and append to list
speed = delta_x / delta_t
speeds.append(speed)
# 8. Update values for next iteration of the loop.
previous_position = position
previous_time = time
return speeds
# 9. Call this function with appropriate arguments
speeds = get_derivative_from_data(displacements, timestamps)
# 10. Prepare labels for a plot
plt.title("Speed vs Time while Parallel Parking")
plt.xlabel("Time (seconds)")
plt.ylabel("Speed (m / s)")
# 11. Make the plot! Note the slicing of timestamps!
plt.scatter(timestamps[1:], speeds)
plt.show()
```
Now that you've read through the code and seen how it's used (and what the resulting plot looks like), I want to discuss the numbered sections of the code.
1. The time and position data need to have equal lengths, since each position measurement is meant to correspond to one of those timestamps.
2. The `speeds` list will eventually be returned at the end of the function.
3. The use of the word "previous" in these variable names will be clearer in step 8. But basically we need to have TWO positions if we're ever going to calculate a delta X. This is where we grab the first position in the position_data list.
4. Note that we loop from `range(1, len(position_data))`, which means that the first value for `i` will be `1` and **not** `0`. That's because we already grabbed element 0 in step 3.
5. Get the data for this `i`.
6. Calculate the change in position and time.
7. Find the slope (which is the speed) and append it to the `speeds` list.
8. This sets the values of `previous_position` and `previous_time` so that they are correct for the *next* iteration of this loop.
9. Here we call the function with the `displacements` and `timestamps` data that we used before.
10. Self-explanatory
11. This part is interesting. Note that we only plot `timestamps[1:]`. This means "every element in `timestamps` except the first one". Remember how in step 4 we looped through every element except the first one? That means that our `speeds` array ends up being 1 element shorter than our original data.
## What to Remember
You don't need to memorize any of this. The important thing to remember is this:
When you're working with real time-series data, you calculate the "derivative" by finding the slope between adjacent data points.
You'll be implementing this on your own in the next notebook. Feel free to come back here if you need help, but try your best to get it on your own.
| github_jupyter |
# MovieLens Data Processing
```
# Import packages
import os
import pandas as pd
# Define file directories
MOVIELENS_DIR = 'dat'
USER_DATA_FILE = 'users.dat'
MOVIE_DATA_FILE = 'movies.dat'
RATING_DATA_FILE = 'ratings.dat'
# Specify User's Age and Occupation Column
AGES = { 1: "Under 18", 18: "18-24", 25: "25-34", 35: "35-44", 45: "45-49", 50: "50-55", 56: "56+" }
OCCUPATIONS = { 0: "other or not specified", 1: "academic/educator", 2: "artist", 3: "clerical/admin",
4: "college/grad student", 5: "customer service", 6: "doctor/health care",
7: "executive/managerial", 8: "farmer", 9: "homemaker", 10: "K-12 student", 11: "lawyer",
12: "programmer", 13: "retired", 14: "sales/marketing", 15: "scientist", 16: "self-employed",
17: "technician/engineer", 18: "tradesman/craftsman", 19: "unemployed", 20: "writer" }
# Define csv files to be saved into
USERS_CSV_FILE = 'users.csv'
MOVIES_CSV_FILE = 'movies.csv'
RATINGS_CSV_FILE = 'ratings.csv'
# Read the Ratings File
ratings = pd.read_csv(os.path.join(MOVIELENS_DIR, RATING_DATA_FILE),
sep='::',
engine='python',
encoding='latin-1',
names=['user_id', 'movie_id', 'rating', 'timestamp'])
# Set max_userid to the maximum user_id in the ratings
max_userid = ratings['user_id'].drop_duplicates().max()
# Set max_movieid to the maximum movie_id in the ratings
max_movieid = ratings['movie_id'].drop_duplicates().max()
# Process ratings dataframe for Keras Deep Learning model
# Add user_emb_id column whose values == user_id - 1
ratings['user_emb_id'] = ratings['user_id'] - 1
# Add movie_emb_id column whose values == movie_id - 1
ratings['movie_emb_id'] = ratings['movie_id'] - 1
print(len(ratings), 'ratings loaded')
# Save into ratings.csv
ratings.to_csv(RATINGS_CSV_FILE,
sep='\t',
header=True,
encoding='latin-1',
columns=['user_id', 'movie_id', 'rating', 'timestamp', 'user_emb_id', 'movie_emb_id'])
print('Saved to', RATINGS_CSV_FILE)
# Read the Users File
users = pd.read_csv(os.path.join(MOVIELENS_DIR, USER_DATA_FILE),
sep='::',
engine='python',
encoding='latin-1',
names=['user_id', 'gender', 'age', 'occupation', 'zipcode'])
users['age_desc'] = users['age'].apply(lambda x: AGES[x])
users['occ_desc'] = users['occupation'].apply(lambda x: OCCUPATIONS[x])
print(len(users), 'descriptions of', max_userid, 'users loaded.')
# Save into users.csv
users.to_csv(USERS_CSV_FILE,
sep='\t',
header=True,
encoding='latin-1',
columns=['user_id', 'gender', 'age', 'occupation', 'zipcode', 'age_desc', 'occ_desc'])
print('Saved to', USERS_CSV_FILE)
# Read the Movies File
movies = pd.read_csv(os.path.join(MOVIELENS_DIR, MOVIE_DATA_FILE),
sep='::',
engine='python',
encoding='latin-1',
names=['movie_id', 'title', 'genres'])
print(len(movies), 'descriptions of', max_movieid, 'movies loaded.')
# Save into movies.csv
movies.to_csv(MOVIES_CSV_FILE,
sep='\t',
header=True,
columns=['movie_id', 'title', 'genres'])
print('Saved to', MOVIES_CSV_FILE)
```
| github_jupyter |
## Herencia
Una clase puede heredar todo los métodos y atributos de otra con esta sintaxis. Ejemplo de una clase heredada:
```
class ClaseA(object):
a = 1
b = 2
def c(self):
return 3
def d(self):
return 4
class ClaseB(ClaseA):
b = 5
def d(self):
return 6
clase_a = ClaseA()
clase_b = ClaseB()
print('1a:', clase_a.a)
print('2a:', clase_a.b)
print('3a:', clase_a.c())
print('4a:', clase_a.d())
print('1b:', clase_b.a)
print('2b:', clase_b.b)
print('3b:', clase_b.c())
print('4b:', clase_b.d())
```
# Appium
___
## Conectar un dispositivo
___
### Pasos comunes
Para conectar un dispositivo de Android hay que seguir los siguientes pasos:
1. Descargar e instalar Java jdk 1.8: https://www.oracle.com/technetwork/java/javase/downloads/jdk8-downloads-2133151.html
2. Añadir la variable de entorno JAVA_HOME = "C:\Program Files\Java\jdk {version} "
3. Descargar e instalar Android Studio: https://developer.android.com/studio
4. Añadir la variable de entorno ANDROID_HOME = "C:\Users\\ {user} \AppData\Local\Android\Sdk\"
5. Añadir el directorio "C:\Users\\ {user} \AppData\Local\Android\Sdk\platform-tools\" al Path de Windows
#### Emulador
Para crear un emulador hay que seguir los siguientes pasos:
1. Lanzar Android Studio, si pide crear un proyecto se crea un vacío (que no usaremos para nada)
2. Dejar que se actualice con las actualizaciones por defecto (puede variar dependiendo de la versión)
3. Ir a "Tools" > "AVD Manager"
4. CLick en "Create Virtual Device".
5. Seleccionar "Phone" > "Nexus 5X", "Next"
6. Seleccionar "Oreo" (API Level 27, Android 8.1), si no está disponible click en descargar, "Next"
7. Nombrar y "Fisinsh"
#### Real
Para conectar un dispositivo real hay que seguir los siguientes pasos (No todos los dispositivos son compatibles):
1. En el dispositivo: Ir a "Settings" > "About phone" > "Software information" y pulsar "Build number" 7 veces
2. En el dispositivo: Ir a "Settings" > "Developer options" y activar "Stay awake" y "USB debugging"
3. Conectar por USB y aceptar permisos
### Comprobar la conexión
Par comprobar que todo funciona correctamente ejecutar:
```
! adb devices
```
___
## Levantar un servidor de Appium en local
___
1. Descargar e instalar Appium-Desktop: https://github.com/appium/appium-desktop/releases/
2. Iniciar Appium (tarda)
3. Poner Host: 0.0.0.0 y Puerto: 4723, pulsar "Start Server"
___
## Crear un script con el cliente de Appium para Python
___
Se instalan los sdk's de Appium para Python:
```
! pip install Appium-Python-Client --trusted-host pypi.python.org
```
___
Importamos la librería:
```
from appium import webdriver
import os
desired_caps = {}
desired_caps['platformName'] = 'Android'
desired_caps['deviceName'] = 'Android Emulator'
desired_caps['app'] = os.path.join(os.getcwd(), 'example.apk') # ruta a una apk de ejemplo
driver = webdriver.Remote('http://localhost:4723/wd/hub', desired_caps)
from appium.webdriver.common.mobileby import MobileBy
class CommonPage(object):
def __init__(self, driver):
self.driver = driver
class InitPage(CommonPage):
def btn__add_contact(self):
return driver.find_element(MobileBy.ACCESSIBILITY_ID, "Add Contact")
class AddContactPage(CommonPage):
def tbx__contact_name(self):
return driver.find_element(MobileBy.ID, "com.example.android.contactmanager:id/contactNameEditText")
def tbx__contact_phone(self):
return driver.find_element(MobileBy.ID, "com.example.android.contactmanager:id/contactPhoneEditText")
InitPage(driver).btn__add_contact().click()
import time
time.sleep(1)
page__add_contact = AddContactPage(driver)
page__add_contact.tbx__contact_name().send_keys('Alejandro')
page__add_contact.tbx__contact_phone().send_keys('987654321')
driver.quit()
```
___
## Obtener los localizadores de objectos manualmente
___
1. Desde Appium, ir a "File" > "New Session Window..."
2. Rellenar la tabla con los valores:
Name | Type | Value
-----|------|------
platformName | text | Android
deviceName | text | Android Emulator
app | text | C:\Users\mtp1923\test-lab\example.apk
3. Pulsar en "Start Session"
Se abrirá una ventana que es similar a pulsar F12 en Chrome
| github_jupyter |
# Introduction to Logistic Regression
## Learning Objectives
1. Create Seaborn plots for Exploratory Data Analysis
2. Train a Logistic Regression Model using Scikit-Learn
## Introduction
This lab is an introduction to logistic regression using Python and Scikit-Learn. This lab serves as a foundation for more complex algorithms and machine learning models that you will encounter in the course. In this lab, we will use a synthetic advertising data set, indicating whether or not a particular internet user clicked on an Advertisement on a company website. We will try to create a model that will predict whether or not they will click on an ad based off the features of that user.
Each learning objective will correspond to a __#TODO__ in this student lab notebook -- try to complete this notebook first and then review the [solution notebook](../solutions/intro_logistic_regression.ipynb).
### Import Libraries
```
import os
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
```
### Load the Dataset
We will use a synthetic [advertising](https://www.kaggle.com/fayomi/advertising) dataset. This data set contains the following features:
* 'Daily Time Spent on Site': consumer time on site in minutes
* 'Age': customer age in years
* 'Area Income': Avg. Income of geographical area of consumer
* 'Daily Internet Usage': Avg. minutes a day consumer is on the internet
* 'Ad Topic Line': Headline of the advertisement
* 'City': City of consumer
* 'Male': Whether or not consumer was male
* 'Country': Country of consumer
* 'Timestamp': Time at which consumer clicked on Ad or closed window
* 'Clicked on Ad': 0 or 1 indicated clicking on Ad
```
# TODO 1: Read in the advertising.csv file and set it to a data frame called ad_data.
# TODO: Your code goes here
```
**Check the head of ad_data**
```
ad_data.head()
```
**Use info and describe() on ad_data**
```
ad_data.info()
ad_data.describe()
```
Let's check for any null values.
```
ad_data.isnull().sum()
```
## Exploratory Data Analysis (EDA)
Let's use seaborn to explore the data! Try recreating the plots shown below!
TODO 1: **Create a histogram of the Age**
```
# TODO: Your code goes here
```
TODO 1: **Create a jointplot showing Area Income versus Age.**
```
# TODO: Your code goes here
```
TODO 2: **Create a jointplot showing the kde distributions of Daily Time spent on site vs. Age.**
```
# TODO: Your code goes here
```
TODO 1: **Create a jointplot of 'Daily Time Spent on Site' vs. 'Daily Internet Usage'**
```
# TODO: Your code goes here
```
# Logistic Regression
Logistic regression is a supervised machine learning process. It is similar to linear regression, but rather than predict a continuous value, we try to estimate probabilities by using a logistic function. Note that even though it has regression in the name, it is for classification.
While linear regression is acceptable for estimating values, logistic regression is best for predicting the class of an observation
Now it's time to do a train test split, and train our model! You'll have the freedom here to choose columns that you want to train on!
```
from sklearn.model_selection import train_test_split
```
Next, let's define the features and label. Briefly, feature is input; label is output. This applies to both classification and regression problems.
```
X = ad_data[['Daily Time Spent on Site', 'Age', 'Area Income','Daily Internet Usage', 'Male']]
y = ad_data['Clicked on Ad']
```
TODO 2: **Split the data into training set and testing set using train_test_split**
```
# TODO: Your code goes here
```
**Train and fit a logistic regression model on the training set.**
```
from sklearn.linear_model import LogisticRegression
logmodel = LogisticRegression()
logmodel.fit(X_train,y_train)
```
## Predictions and Evaluations
**Now predict values for the testing data.**
```
predictions = logmodel.predict(X_test)
```
**Create a classification report for the model.**
```
from sklearn.metrics import classification_report
print(classification_report(y_test,predictions))
```
Copyright 2021 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at
http://www.apache.org/licenses/LICENSE-2.0
Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
| github_jupyter |
# Generative Adversarial Network
In this notebook, we'll be building a generative adversarial network (GAN) trained on the MNIST dataset. From this, we'll be able to generate new handwritten digits!
GANs were [first reported on](https://arxiv.org/abs/1406.2661) in 2014 from Ian Goodfellow and others in Yoshua Bengio's lab. Since then, GANs have exploded in popularity. Here are a few examples to check out:
* [Pix2Pix](https://affinelayer.com/pixsrv/)
* [CycleGAN & Pix2Pix in PyTorch, Jun-Yan Zhu](https://github.com/junyanz/pytorch-CycleGAN-and-pix2pix)
* [A list of generative models](https://github.com/wiseodd/generative-models)
The idea behind GANs is that you have two networks, a generator $G$ and a discriminator $D$, competing against each other. The generator makes "fake" data to pass to the discriminator. The discriminator also sees real training data and predicts if the data it's received is real or fake.
> * The generator is trained to fool the discriminator, it wants to output data that looks _as close as possible_ to real, training data.
* The discriminator is a classifier that is trained to figure out which data is real and which is fake.
What ends up happening is that the generator learns to make data that is indistinguishable from real data to the discriminator.
<img src='assets/gan_pipeline.png' width=70% />
The general structure of a GAN is shown in the diagram above, using MNIST images as data. The latent sample is a random vector that the generator uses to construct its fake images. This is often called a **latent vector** and that vector space is called **latent space**. As the generator trains, it figures out how to map latent vectors to recognizable images that can fool the discriminator.
If you're interested in generating only new images, you can throw out the discriminator after training. In this notebook, I'll show you how to define and train these adversarial networks in PyTorch and generate new images!
```
%matplotlib inline
import numpy as np
import torch
import matplotlib.pyplot as plt
from torchvision import datasets
import torchvision.transforms as transforms
# number of subprocesses to use for data loading
num_workers = 0
# how many samples per batch to load
batch_size = 64
# convert data to torch.FloatTensor
transform = transforms.ToTensor()
# get the training datasets
train_data = datasets.MNIST(root='data', train=True,
download=True, transform=transform)
# prepare data loader
train_loader = torch.utils.data.DataLoader(train_data, batch_size=batch_size,
num_workers=num_workers)
```
### Visualize the data
```
# obtain one batch of training images
dataiter = iter(train_loader)
images, labels = dataiter.next()
images = images.numpy()
# get one image from the batch
img = np.squeeze(images[0])
fig = plt.figure(figsize = (3,3))
ax = fig.add_subplot(111)
ax.imshow(img, cmap='gray')
```
---
# Define the Model
A GAN is comprised of two adversarial networks, a discriminator and a generator.
## Discriminator
The discriminator network is going to be a pretty typical linear classifier. To make this network a universal function approximator, we'll need at least one hidden layer, and these hidden layers should have one key attribute:
> All hidden layers will have a [Leaky ReLu](https://pytorch.org/docs/stable/nn.html#torch.nn.LeakyReLU) activation function applied to their outputs.
<img src='assets/gan_network.png' width=70% />
#### Leaky ReLu
We should use a leaky ReLU to allow gradients to flow backwards through the layer unimpeded. A leaky ReLU is like a normal ReLU, except that there is a small non-zero output for negative input values.
<img src='assets/leaky_relu.png' width=40% />
#### Sigmoid Output
We'll also take the approach of using a more numerically stable loss function on the outputs. Recall that we want the discriminator to output a value 0-1 indicating whether an image is _real or fake_.
> We will ultimately use [BCEWithLogitsLoss](https://pytorch.org/docs/stable/nn.html#bcewithlogitsloss), which combines a `sigmoid` activation function **and** and binary cross entropy loss in one function.
So, our final output layer should not have any activation function applied to it.
```
import torch.nn as nn
import torch.nn.functional as F
class Discriminator(nn.Module):
def __init__(self, input_size, hidden_dim, output_size):
super(Discriminator, self).__init__()
# define all layers
self.fc1 = nn.Linear(input_size, hidden_dim * 4)
self.fc2 = nn.Linear(hidden_dim * 4, hidden_dim * 2)
self.fc3 = nn.Linear(hidden_dim * 2, hidden_dim)
self.fc4 = nn.Linear(hidden_dim, output_size)
self.dropout = nn.Dropout(0.3)
def forward(self, x):
# flatten image
x = x.view(-1, 28 * 28)
# pass x through all layers
# apply leaky relu activation to all hidden layers
x = F.leaky_relu(self.fc1(x), 0.2)
x = self.dropout(x)
x = F.leaky_relu(self.fc2(x), 0.2)
x = self.dropout(x)
x = F.leaky_relu(self.fc3(x), 0.2)
x = self.dropout(x)
out = self.fc4(x)
return out
```
## Generator
The generator network will be almost exactly the same as the discriminator network, except that we're applying a [tanh activation function](https://pytorch.org/docs/stable/nn.html#tanh) to our output layer.
#### tanh Output
The generator has been found to perform the best with $tanh$ for the generator output, which scales the output to be between -1 and 1, instead of 0 and 1.
<img src='assets/tanh_fn.png' width=40% />
Recall that we also want these outputs to be comparable to the *real* input pixel values, which are read in as normalized values between 0 and 1.
> So, we'll also have to **scale our real input images to have pixel values between -1 and 1** when we train the discriminator.
I'll do this in the training loop, later on.
```
class Generator(nn.Module):
def __init__(self, input_size, hidden_dim, output_size):
super(Generator, self).__init__()
# define all layers
self.fc1 = nn.Linear(input_size, hidden_dim)
self.fc2 = nn.Linear(hidden_dim, hidden_dim * 2)
self.fc3 = nn.Linear(hidden_dim * 2, hidden_dim * 4)
self.fc4 = nn.Linear(hidden_dim * 4, output_size)
self.dropout = nn.Dropout(0.2)
def forward(self, x):
# pass x through all layers
x = F.leaky_relu(self.fc1(x), 0.2)
x = self.dropout(x)
x = F.leaky_relu(self.fc2(x), 0.2)
x = self.dropout(x)
x = F.leaky_relu(self.fc3(x), 0.2)
x = self.dropout(x)
# final layer should have tanh applied
out = F.tanh(self.fc4(x))
return out
```
## Model hyperparameters
```
# Discriminator hyperparams
# Size of input image to discriminator (28*28)
input_size = 784
# Size of discriminator output (real or fake)
d_output_size = 1
# Size of *last* hidden layer in the discriminator
d_hidden_size = 32
# Generator hyperparams
# Size of latent vector to give to generator
z_size = 100
# Size of discriminator output (generated image)
g_output_size = 784
# Size of *first* hidden layer in the generator
g_hidden_size = 32
```
## Build complete network
Now we're instantiating the discriminator and generator from the classes defined above. Make sure you've passed in the correct input arguments.
```
# instantiate discriminator and generator
D = Discriminator(input_size, d_hidden_size, d_output_size)
G = Generator(z_size, g_hidden_size, g_output_size)
# check that they are as you expect
print(D)
print()
print(G)
```
---
## Discriminator and Generator Losses
Now we need to calculate the losses.
### Discriminator Losses
> * For the discriminator, the total loss is the sum of the losses for real and fake images, `d_loss = d_real_loss + d_fake_loss`.
* Remember that we want the discriminator to output 1 for real images and 0 for fake images, so we need to set up the losses to reflect that.
<img src='assets/gan_pipeline.png' width=70% />
The losses will by binary cross entropy loss with logits, which we can get with [BCEWithLogitsLoss](https://pytorch.org/docs/stable/nn.html#bcewithlogitsloss). This combines a `sigmoid` activation function **and** and binary cross entropy loss in one function.
For the real images, we want `D(real_images) = 1`. That is, we want the discriminator to classify the the real images with a label = 1, indicating that these are real. To help the discriminator generalize better, the labels are **reduced a bit from 1.0 to 0.9**. For this, we'll use the parameter `smooth`; if True, then we should smooth our labels. In PyTorch, this looks like `labels = torch.ones(size) * 0.9`
The discriminator loss for the fake data is similar. We want `D(fake_images) = 0`, where the fake images are the _generator output_, `fake_images = G(z)`.
### Generator Loss
The generator loss will look similar only with flipped labels. The generator's goal is to get `D(fake_images) = 1`. In this case, the labels are **flipped** to represent that the generator is trying to fool the discriminator into thinking that the images it generates (fakes) are real!
```
# Calculate losses
def real_loss(D_out, smooth=False):
# compare logits to real labels
# smooth labels if smooth=True
batch_size = D_out.size(0)
if smooth:
labels = torch.ones(batch_size) * 0.9
else:
labels = torch.ones(batch_size)
criterion = nn.BCEWithLogitsLoss()
loss = criterion(D_out.squeeze(), labels)
return loss
def fake_loss(D_out):
# compare logits to fake labels
batch_size = D_out.size(0)
labels = torch.zeros(batch_size)
criterion = nn.BCEWithLogitsLoss()
loss = criterion(D_out.squeeze(), labels)
return loss
```
## Optimizers
We want to update the generator and discriminator variables separately. So, we'll define two separate Adam optimizers.
```
import torch.optim as optim
# learning rate for optimizers
lr = 0.002
# Create optimizers for the discriminator and generator
d_optimizer = optim.Adam(D.parameters(), lr)
g_optimizer = optim.Adam(G.parameters(), lr)
```
---
## Training
Training will involve alternating between training the discriminator and the generator. We'll use our functions `real_loss` and `fake_loss` to help us calculate the discriminator losses in all of the following cases.
### Discriminator training
1. Compute the discriminator loss on real, training images
2. Generate fake images
3. Compute the discriminator loss on fake, generated images
4. Add up real and fake loss
5. Perform backpropagation + an optimization step to update the discriminator's weights
### Generator training
1. Generate fake images
2. Compute the discriminator loss on fake images, using **flipped** labels!
3. Perform backpropagation + an optimization step to update the generator's weights
#### Saving Samples
As we train, we'll also print out some loss statistics and save some generated "fake" samples.
```
import pickle as pkl
# training hyperparams
num_epochs = 40
# keep track of loss and generated, "fake" samples
samples = []
losses = []
print_every = 400
# Get some fixed data for sampling. These are images that are held
# constant throughout training, and allow us to inspect the model's performance
sample_size=16
fixed_z = np.random.uniform(-1, 1, size=(sample_size, z_size))
fixed_z = torch.from_numpy(fixed_z).float()
# train the network
D.train()
G.train()
for epoch in range(num_epochs):
for batch_i, (real_images, _) in enumerate(train_loader):
batch_size = real_images.size(0)
## Important rescaling step ##
real_images = real_images*2 - 1 # rescale input images from [0,1) to [-1, 1)
# ============================================
# TRAIN THE DISCRIMINATOR
# ============================================
d_optimizer.zero_grad()
# 1. Train with real images
# Compute the discriminator losses on real images
# use smoothed labels
D_real = D(real_images)
d_real_loss = real_loss(D_real, smooth=True)
# 2. Train with fake images
# Generate fake images
z = np.random.uniform(-1, 1, size=(batch_size, z_size))
z = torch.from_numpy(z).float()
fake_images = G(z)
# Compute the discriminator losses on fake images
D_fake = D(fake_images)
d_fake_loss = fake_loss(D_fake)
# add up real and fake losses and perform backprop
d_loss = d_real_loss + d_fake_loss
d_loss.backward()
d_optimizer.step()
# =========================================
# TRAIN THE GENERATOR
# =========================================
g_optimizer.zero_grad()
# 1. Train with fake images and flipped labels
# Generate fake images
z = np.random.uniform(-1, 1, size=(batch_size, z_size))
z = torch.from_numpy(z).float()
fake_images = G(z)
# Compute the discriminator losses on fake images
# using flipped labels!
D_fake = D(fake_images)
g_loss = real_loss(D_fake)
# perform backprop
g_loss.backward()
g_optimizer.step()
# Print some loss stats
if batch_i % print_every == 0:
# print discriminator and generator loss
print('Epoch [{:5d}/{:5d}] | d_loss: {:6.4f} | g_loss: {:6.4f}'.format(
epoch+1, num_epochs, d_loss.item(), g_loss.item()))
## AFTER EACH EPOCH##
# append discriminator loss and generator loss
losses.append((d_loss.item(), g_loss.item()))
# generate and save sample, fake images
G.eval() # eval mode for generating samples
samples_z = G(fixed_z)
samples.append(samples_z)
G.train() # back to train mode
# Save training generator samples
with open('train_samples.pkl', 'wb') as f:
pkl.dump(samples, f)
```
## Training loss
Here we'll plot the training losses for the generator and discriminator, recorded after each epoch.
```
fig, ax = plt.subplots()
losses = np.array(losses)
plt.plot(losses.T[0], label='Discriminator')
plt.plot(losses.T[1], label='Generator')
plt.title("Training Losses")
plt.legend()
```
## Generator samples from training
Here we can view samples of images from the generator. First we'll look at the images we saved during training.
```
# helper function for viewing a list of passed in sample images
def view_samples(epoch, samples):
fig, axes = plt.subplots(figsize=(7,7), nrows=4, ncols=4, sharey=True, sharex=True)
for ax, img in zip(axes.flatten(), samples[epoch]):
img = img.detach()
ax.xaxis.set_visible(False)
ax.yaxis.set_visible(False)
im = ax.imshow(img.reshape((28,28)), cmap='Greys_r')
# Load samples from generator, taken while training
with open('train_samples.pkl', 'rb') as f:
samples = pkl.load(f)
```
These are samples from the final training epoch. You can see the generator is able to reproduce numbers like 1, 7, 3, 2. Since this is just a sample, it isn't representative of the full range of images this generator can make.
```
# -1 indicates final epoch's samples (the last in the list)
view_samples(-1, samples)
```
Below I'm showing the generated images as the network was training, every 10 epochs.
```
rows = 10 # split epochs into 10, so 100/10 = every 10 epochs
cols = 6
fig, axes = plt.subplots(figsize=(7,12), nrows=rows, ncols=cols, sharex=True, sharey=True)
for sample, ax_row in zip(samples[::int(len(samples)/rows)], axes):
for img, ax in zip(sample[::int(len(sample)/cols)], ax_row):
img = img.detach()
ax.imshow(img.reshape((28,28)), cmap='Greys_r')
ax.xaxis.set_visible(False)
ax.yaxis.set_visible(False)
```
It starts out as all noise. Then it learns to make only the center white and the rest black. You can start to see some number like structures appear out of the noise like 1s and 9s.
## Sampling from the generator
We can also get completely new images from the generator by using the checkpoint we saved after training. **We just need to pass in a new latent vector $z$ and we'll get new samples**!
```
# randomly generated, new latent vectors
sample_size=16
rand_z = np.random.uniform(-1, 1, size=(sample_size, z_size))
rand_z = torch.from_numpy(rand_z).float()
G.eval() # eval mode
# generated samples
rand_images = G(rand_z)
# 0 indicates the first set of samples in the passed in list
# and we only have one batch of samples, here
view_samples(0, [rand_images])
```
| github_jupyter |
```
"""
We use following lines because we are running on Google Colab
If you are running notebook on a local computer, you don't need this cell
"""
from google.colab import drive
drive.mount('/content/gdrive')
import os
os.chdir('/content/gdrive/My Drive/finch/tensorflow1/free_chat/chinese/main')
%tensorflow_version 1.x
!pip install texar
import tensorflow as tf
import texar.tf as tx
import numpy as np
import copy
from texar.tf.modules import TransformerEncoder, TransformerDecoder
print("TensorFlow Version", tf.__version__)
print('GPU Enabled:', tf.test.is_gpu_available())
def forward(features, labels, mode):
if isinstance(features, dict):
words = features['words']
else:
words = features
words_len = tf.count_nonzero(words, 1, dtype=tf.int32)
is_training = (mode == tf.estimator.ModeKeys.TRAIN)
batch_sz = tf.shape(words)[0]
with tf.variable_scope('Embedding'):
embedding = tf.Variable(np.load('../vocab/char.npy'),
dtype=tf.float32,
name='fasttext_vectors')
embedding = tf.concat([tf.zeros(shape=[1, params['embed_dim']]), embedding[1:, :]], axis=0)
x = tf.nn.embedding_lookup(embedding, words)
pos_embedder = tx.modules.SinusoidsPositionEmbedder(
position_size = 2*params['max_len'],
hparams = config_model.position_embedder_hparams)
x = (x * config_model.hidden_dim ** 0.5) + pos_embedder(sequence_length=words_len)
with tf.variable_scope('Encoder'):
encoder = TransformerEncoder(hparams=config_model.encoder)
enc_out = encoder(inputs=x, sequence_length=words_len)
with tf.variable_scope('Decoder'):
decoder = TransformerDecoder(vocab_size=len(params['char2idx'])+1,
output_layer=tf.transpose(embedding, (1, 0)),
hparams=config_model.decoder)
start_tokens = tf.fill([batch_sz], 1)
def _embedding_fn(x, y):
x_w_embed = tf.nn.embedding_lookup(embedding, x)
y_p_embed = pos_embedder(y)
return x_w_embed * config_model.hidden_dim ** 0.5 + y_p_embed
predictions = decoder(
memory=enc_out,
memory_sequence_length=words_len,
beam_width=params['beam_width'],
length_penalty=params['length_penalty'],
start_tokens=start_tokens,
end_token=2,
embedding=_embedding_fn,
max_decoding_length=params['max_len'],
mode=tf.estimator.ModeKeys.PREDICT)
return predictions['sample_id'][:, :, :params['top_k']]
def model_fn(features, labels, mode, params):
logits_or_ids = forward(features, labels, mode)
if mode == tf.estimator.ModeKeys.PREDICT:
return tf.estimator.EstimatorSpec(mode, predictions=logits_or_ids)
class config_model:
hidden_dim = 300
num_heads = 8
dropout_rate = .2
num_blocks = 6
position_embedder_hparams = {
'dim': hidden_dim
}
encoder = {
'dim': hidden_dim,
'embedding_dropout': dropout_rate,
'residual_dropout': dropout_rate,
'num_blocks': num_blocks,
'initializer': {
'type': 'variance_scaling_initializer',
'kwargs': {
'scale': 1.0,
'mode': 'fan_avg',
'distribution': 'uniform',
},
},
'multihead_attention': {
'dropout_rate': dropout_rate,
'num_heads': num_heads,
'output_dim': hidden_dim,
'use_bias': True,
},
'poswise_feedforward': {
'name': 'fnn',
'layers': [
{
'type': 'Dense',
'kwargs': {
'name': 'conv1',
'units': hidden_dim * 2,
'activation': 'gelu',
'use_bias': True,
},
},
{
'type': 'Dropout',
'kwargs': {
'rate': dropout_rate,
}
},
{
'type': 'Dense',
'kwargs': {
'name': 'conv2',
'units': hidden_dim,
'use_bias': True,
}
}
],
},
}
decoder = copy.deepcopy(encoder)
decoder['output_layer_bias'] = True
params = {
'model_dir': '../model/transformer',
'export_dir': '../model/transformer_export',
'vocab_path': '../vocab/char.txt',
'max_len': 15,
'embed_dim': config_model.hidden_dim,
'beam_width': 5,
'top_k': 3,
'length_penalty': .6,
}
def serving_input_receiver_fn():
words = tf.placeholder(tf.int32, [None, None], 'words')
features = {'words': words}
receiver_tensors = features
return tf.estimator.export.ServingInputReceiver(features, receiver_tensors)
def get_vocab(f_path):
word2idx = {}
with open(f_path) as f:
for i, line in enumerate(f):
line = line.rstrip('\n')
word2idx[line] = i
return word2idx
params['char2idx'] = get_vocab(params['vocab_path'])
params['idx2char'] = {idx: char for char, idx in params['char2idx'].items()}
estimator = tf.estimator.Estimator(model_fn, params['model_dir'])
estimator.export_saved_model(params['export_dir'], serving_input_receiver_fn)
```
| github_jupyter |
```
from scipy.misc import derivative
import scipy.integrate
from math import *
import sympy as sp
import matplotlib.pyplot as plt
import pandas as pd
import random
from sklearn.datasets.samples_generator import make_regression
from scipy import stats
import seaborn as sns
def f1(x):
return sin(x) * cos(x) + exp(2*x) + 2*x**4 - 10
f1_result = derivative(f1, 2, dx = 1e-10)
f1_result
def f2(x):
return (5*(x**2))/sin(x)
f2_result = derivative(f2, 2, dx = 1e-10)
f2_result
```
## Partial Derivative
```
x = sp.Symbol('x')
y = sp.Symbol('y')
z = sp.Symbol('z')
w = (x**2)*y - 10*(y**2)*(z**3) + 43*x - 7*sp.tan(4*y)
w_p = sp.diff(w, x)
w_p
w_p_v = w_p.subs({x:3})
w_p_v
```
## Derivarive Rule
```
def f(x):
return 4*(x**3) + 3*(x**2)
def g(x):
return 5*(x**6) + 4*x
x = 2
fg_prime_of_x = derivative(f, x, dx=1e-10)*g(x) + f(x)*derivative(g, x, dx=1e-10)
fg_prime_of_x
f_prime_g_prime_of_x = derivative(f, x, dx=1e-10) * derivative(g, x, dx=1e-10)
f_prime_g_prime_of_x
```
## Definite Integral
```
f = lambda x,y: 1 + 8*x*y
g = lambda x:0
h = lambda x:3
i, e = scipy.integrate.dblquad(f, 1,2, g, h)
i
```
## Gradient Descent
```
h = [74, 74, 72, 72, 73, 69, 69, 71, 76, 71, 73, 73, 74, 74, 69, 70, 73, 75, 78, 79, 76, 74, 76, 72, 71, 75, 77, 74, 73, 74, 78, 73, 75, 73, 75, 75, 74, 69, 71, 74, 73, 73, 76, 74, 74, 70, 72, 77, 74, 70, 73, 75, 76, 76, 78, 74, 74, 76, 77, 81, 78, 75, 77, 75, 76, 74, 72, 72, 75, 73, 73, 73, 70, 70, 70, 76, 68, 71, 72, 75, 75, 75, 75, 68, 74, 78, 71, 73, 76, 74, 74, 79, 75, 73, 76, 74, 74, 73, 72, 74, 73, 74, 72, 73, 69, 72, 73, 75, 75, 73, 72, 72, 76, 74, 72, 77, 74, 77, 75, 76, 80, 74, 74, 75, 78, 73, 73, 74, 75, 76, 71, 73, 74, 76, 76, 74, 73, 74, 70, 72, 73, 73, 73, 73, 71, 74, 74, 72, 74, 71, 74, 73, 75, 75, 79, 73, 75, 76, 74, 76, 78, 74, 76, 72, 74, 76, 74, 75, 78, 75, 72, 74, 72, 74, 70, 71, 70, 75, 71, 71, 73, 72, 71, 73, 72, 75, 74, 74, 75, 73, 77, 73, 76, 75, 74, 76, 75, 73, 71, 76, 75, 72, 71, 77, 73, 74, 71, 72, 74, 75, 73, 72, 75, 75, 74, 72, 74, 71, 70, 74, 77, 77, 75, 75, 78, 75, 76, 73, 75, 75, 79, 77, 76, 71, 75, 74, 69, 71, 76, 72, 72, 70, 72, 73, 71, 72, 71, 73, 72, 73, 74, 74, 72, 75, 74, 74, 77, 75, 73, 72, 71, 74, 77, 75, 75, 75, 78, 78, 74, 76, 78, 76, 70, 72, 80, 74, 74, 71, 70, 72, 71, 74, 71, 72, 71, 74, 69, 76, 75, 75, 76, 73, 76, 73, 77, 73, 72, 72, 77, 77, 71, 74, 74, 73, 78, 75, 73, 70, 74, 72, 73, 73, 75, 75, 74, 76, 73, 74, 75, 75, 72, 73, 73, 72, 74, 78, 76, 73, 74, 75, 70, 75, 71, 72, 78, 75, 73, 73, 71, 75, 77, 72, 69, 73, 74, 72, 70, 75, 70, 72, 72, 74, 73, 74, 76, 75, 80, 72, 75, 73, 74, 74, 73, 75, 75, 71, 73, 75, 74, 74, 72, 74, 74, 74, 73, 76, 75, 72, 73, 73, 73, 72, 72, 72, 72, 71, 75, 75, 74, 73, 75, 79, 74, 76, 73, 74, 74, 72, 74, 74, 75, 78, 74, 74, 74, 77, 70, 73, 74, 73, 71, 75, 71, 72, 77, 74, 70, 77, 73, 72, 76, 71, 76, 78, 75, 73, 78, 74, 79, 75, 76, 72, 75, 75, 70, 72, 70, 74, 71, 76, 73, 76, 71, 69, 72, 72, 69, 73, 69, 73, 74, 74, 72, 71, 72, 72, 76, 76, 76, 74, 76, 75, 71, 72, 71, 73, 75, 76, 75, 71, 75, 74, 72, 73, 73, 73, 73, 76, 72, 76, 73, 73, 73, 75, 75, 77, 73, 72, 75, 70, 74, 72, 80, 71, 71, 74, 74, 73, 75, 76, 73, 77, 72, 73, 77, 76, 71, 75, 73, 74, 77, 71, 72, 73, 69, 73, 70, 74, 76, 73, 73, 75, 73, 79, 74, 73, 74, 77, 75, 74, 73, 77, 73, 77, 74, 74, 73, 77, 74, 77, 75, 77, 75, 71, 74, 70, 79, 72, 72, 70, 74, 74, 72, 73, 72, 74, 74, 76, 82, 74, 74, 70, 73, 73, 74, 77, 72, 76, 73, 73, 72, 74, 74, 71, 72, 75, 74, 74, 77, 70, 71, 73, 76, 71, 75, 74, 72, 76, 79, 76, 73, 76, 78, 75, 76, 72, 72, 73, 73, 75, 71, 76, 70, 75, 74, 75, 73, 71, 71, 72, 73, 73, 72, 69, 73, 78, 71, 73, 75, 76, 70, 74, 77, 75, 79, 72, 77, 73, 75, 75, 75, 73, 73, 76, 77, 75, 70, 71, 71, 75, 74, 69, 70, 75, 72, 75, 73, 72, 72, 72, 76, 75, 74, 69, 73, 72, 72, 75, 77, 76, 80, 77, 76, 79, 71, 75, 73, 76, 77, 73, 76, 70, 75, 73, 75, 70, 69, 71, 72, 72, 73, 70, 70, 73, 76, 75, 72, 73, 79, 71, 72, 74, 74, 74, 72, 76, 76, 72, 72, 71, 72, 72, 70, 77, 74, 72, 76, 71, 76, 71, 73, 70, 73, 73, 72, 71, 71, 71, 72, 72, 74, 74, 74, 71, 72, 75, 72, 71, 72, 72, 72, 72, 74, 74, 77, 75, 73, 75, 73, 76, 72, 77, 75, 72, 71, 71, 75, 72, 73, 73, 71, 70, 75, 71, 76, 73, 68, 71, 72, 74, 77, 72, 76, 78, 81, 72, 73, 76, 72, 72, 74, 76, 73, 76, 75, 70, 71, 74, 72, 73, 76, 76, 73, 71, 68, 71, 71, 74, 77, 69, 72, 76, 75, 76, 75, 76, 72, 74, 76, 74, 72, 75, 78, 77, 70, 72, 79, 74, 71, 68, 77, 75, 71, 72, 70, 72, 72, 73, 72, 74, 72, 72, 75, 72, 73, 74, 72, 78, 75, 72, 74, 75, 75, 76, 74, 74, 73, 74, 71, 74, 75, 76, 74, 76, 76, 73, 75, 75, 74, 68, 72, 75, 71, 70, 72, 73, 72, 75, 74, 70, 76, 71, 82, 72, 73, 74, 71, 75, 77, 72, 74, 72, 73, 78, 77, 73, 73, 73, 73, 73, 76, 75, 70, 73, 72, 73, 75, 74, 73, 73, 76, 73, 75, 70, 77, 72, 77, 74, 75, 75, 75, 75, 72, 74, 71, 76, 71, 75, 76, 83, 75, 74, 76, 72, 72, 75, 75, 72, 77, 73, 72, 70, 74, 72, 74, 72, 71, 70, 71, 76, 74, 76, 74, 74, 74, 75, 75, 71, 71, 74, 77, 71, 74, 75, 77, 76, 74, 76, 72, 71, 72, 75, 73, 68, 72, 69, 73, 73, 75, 70, 70, 74, 75, 74, 74, 73, 74, 75, 77, 73, 74, 76, 74, 75, 73, 76, 78, 75, 73, 77, 74, 72, 74, 72, 71, 73, 75, 73, 67, 67, 76, 74, 73, 70, 75, 70, 72, 77, 79, 78, 74, 75, 75, 78, 76, 75, 69, 75, 72, 75, 73, 74, 75, 75, 73]
w = [180, 215, 210, 210, 188, 176, 209, 200, 231, 180, 188, 180, 185, 160, 180, 185, 189, 185, 219, 230, 205, 230, 195, 180, 192, 225, 203, 195, 182, 188, 200, 180, 200, 200, 245, 240, 215, 185, 175, 199, 200, 215, 200, 205, 206, 186, 188, 220, 210, 195, 200, 200, 212, 224, 210, 205, 220, 195, 200, 260, 228, 270, 200, 210, 190, 220, 180, 205, 210, 220, 211, 200, 180, 190, 170, 230, 155, 185, 185, 200, 225, 225, 220, 160, 205, 235, 250, 210, 190, 160, 200, 205, 222, 195, 205, 220, 220, 170, 185, 195, 220, 230, 180, 220, 180, 180, 170, 210, 215, 200, 213, 180, 192, 235, 185, 235, 210, 222, 210, 230, 220, 180, 190, 200, 210, 194, 180, 190, 240, 200, 198, 200, 195, 210, 220, 190, 210, 225, 180, 185, 170, 185, 185, 180, 178, 175, 200, 204, 211, 190, 210, 190, 190, 185, 290, 175, 185, 200, 220, 170, 220, 190, 220, 205, 200, 250, 225, 215, 210, 215, 195, 200, 194, 220, 180, 180, 170, 195, 180, 170, 206, 205, 200, 225, 201, 225, 233, 180, 225, 180, 220, 180, 237, 215, 190, 235, 190, 180, 165, 195, 200, 190, 190, 185, 185, 205, 190, 205, 206, 220, 208, 170, 195, 210, 190, 211, 230, 170, 185, 185, 241, 225, 210, 175, 230, 200, 215, 198, 226, 278, 215, 230, 240, 184, 219, 170, 218, 190, 225, 220, 176, 190, 197, 204, 167, 180, 195, 220, 215, 185, 190, 205, 205, 200, 210, 215, 200, 205, 211, 190, 208, 200, 210, 232, 230, 210, 220, 210, 202, 212, 225, 170, 190, 200, 237, 220, 170, 193, 190, 150, 220, 200, 190, 185, 185, 200, 172, 220, 225, 190, 195, 219, 190, 197, 200, 195, 210, 177, 220, 235, 180, 195, 195, 190, 230, 190, 200, 190, 190, 200, 200, 184, 200, 180, 219, 187, 200, 220, 205, 190, 170, 160, 215, 175, 205, 200, 214, 200, 190, 180, 205, 220, 190, 215, 235, 191, 200, 181, 200, 210, 240, 185, 165, 190, 185, 175, 155, 210, 170, 175, 220, 210, 205, 200, 205, 195, 240, 150, 200, 215, 202, 200, 190, 205, 190, 160, 215, 185, 200, 190, 210, 185, 220, 190, 202, 205, 220, 175, 160, 190, 200, 229, 206, 220, 180, 195, 175, 188, 230, 190, 200, 190, 219, 235, 180, 180, 180, 200, 234, 185, 220, 223, 200, 210, 200, 210, 190, 177, 227, 180, 195, 199, 175, 185, 240, 210, 180, 194, 225, 180, 205, 193, 230, 230, 220, 200, 249, 190, 208, 245, 250, 160, 192, 220, 170, 197, 155, 190, 200, 220, 210, 228, 190, 160, 184, 180, 180, 200, 176, 160, 222, 211, 195, 200, 175, 206, 240, 185, 260, 185, 221, 205, 200, 170, 201, 205, 185, 205, 245, 220, 210, 220, 185, 175, 170, 180, 200, 210, 175, 220, 206, 180, 210, 195, 200, 200, 164, 180, 220, 195, 205, 170, 240, 210, 195, 200, 205, 192, 190, 170, 240, 200, 205, 175, 250, 220, 224, 210, 195, 180, 245, 175, 180, 215, 175, 180, 195, 230, 230, 205, 215, 195, 180, 205, 180, 190, 180, 190, 190, 220, 210, 255, 190, 230, 200, 205, 210, 225, 215, 220, 205, 200, 220, 197, 225, 187, 245, 185, 185, 175, 200, 180, 188, 225, 200, 210, 245, 213, 231, 165, 228, 210, 250, 191, 190, 200, 215, 254, 232, 180, 215, 220, 180, 200, 170, 195, 210, 200, 220, 165, 180, 200, 200, 170, 224, 220, 180, 198, 240, 239, 185, 210, 220, 200, 195, 220, 230, 170, 220, 230, 165, 205, 192, 210, 205, 200, 210, 185, 195, 202, 205, 195, 180, 200, 185, 240, 185, 220, 205, 205, 180, 201, 190, 208, 240, 180, 230, 195, 215, 190, 195, 215, 215, 220, 220, 230, 195, 190, 195, 209, 204, 170, 185, 205, 175, 210, 190, 180, 180, 160, 235, 200, 210, 180, 190, 197, 203, 205, 170, 200, 250, 200, 220, 200, 190, 170, 190, 220, 215, 206, 215, 185, 235, 188, 230, 195, 168, 190, 160, 200, 200, 189, 180, 190, 200, 220, 187, 240, 190, 180, 185, 210, 220, 219, 190, 193, 175, 180, 215, 210, 200, 190, 185, 220, 170, 195, 205, 195, 210, 190, 190, 180, 220, 190, 186, 185, 190, 180, 190, 170, 210, 240, 220, 180, 210, 210, 195, 160, 180, 205, 200, 185, 245, 190, 210, 200, 200, 222, 215, 240, 170, 220, 156, 190, 202, 221, 200, 190, 210, 190, 200, 165, 190, 185, 230, 208, 209, 175, 180, 200, 205, 200, 250, 210, 230, 244, 202, 240, 200, 215, 177, 210, 170, 215, 217, 198, 200, 220, 170, 200, 230, 231, 183, 192, 167, 190, 180, 180, 215, 160, 205, 223, 175, 170, 190, 240, 175, 230, 223, 196, 167, 195, 190, 250, 190, 190, 190, 170, 160, 150, 225, 220, 209, 210, 176, 260, 195, 190, 184, 180, 195, 195, 219, 225, 212, 202, 185, 200, 209, 200, 195, 228, 210, 190, 212, 190, 218, 220, 190, 235, 210, 200, 188, 210, 235, 188, 215, 216, 220, 180, 185, 200, 210, 220, 185, 231, 210, 195, 200, 205, 200, 190, 250, 185, 180, 170, 180, 208, 235, 215, 244, 220, 185, 230, 190, 200, 180, 190, 196, 180, 230, 224, 160, 178, 205, 185, 210, 180, 190, 200, 257, 190, 220, 165, 205, 200, 208, 185, 215, 170, 235, 210, 170, 180, 170, 190, 150, 230, 203, 260, 246, 186, 210, 198, 210, 215, 180, 200, 245, 200, 192, 192, 200, 192, 205, 190, 186, 170, 197, 219, 200, 220, 207, 225, 207, 212, 225, 170, 190, 210, 230, 210, 200, 238, 234, 222, 200, 190, 170, 220, 223, 210, 215, 196, 175, 175, 189, 205, 210, 180, 180, 197, 220, 228, 190, 204, 165, 216, 220, 208, 210, 215, 195, 200, 215, 229, 240, 207, 205, 208, 185, 190, 170, 208, 225, 190, 225, 185, 180, 165, 240, 220, 212, 163, 215, 175, 205, 210, 205, 208, 215, 180, 200, 230, 211, 230, 190, 220, 180, 205, 190, 180, 205, 190, 195]
heights = np.array(h)
weights = np.array(w)
heights = heights*0.0254
weights = weights * 0.453592
plt.figure(figsize=(12,8))
sns.jointplot(x=heights, y = weights)
plt.show()
# weights = m * heights + b
print(heights.shape)
print(weights.shape)
weights = weights.reshape(weights.size, 1)
heights = heights.reshape(weights.size, 1)
import sys
sys.path.insert(
0,
r"C:\Users\DELL\Desktop\Sentifi\machine_learning\linear_regression_from_scratch\gradient_descent",
)
import GD_linear_regression as glr
builder = glr.GDLinearRegression(iterations=200)
reg = builder.fit(heights, weights)
predictions = reg.predict(heights)
costs = reg.costs
thetas = reg.theta
print(len(costs))
print(reg.theta[-1])
print(costs[-1])
plt.plot(costs)
plt.show()
```
| github_jupyter |
# PyIndMach012: an example of user-model using DSS Python
This example runs a modified example from the OpenDSS distribution for the induction machine model with a sample PyIndMach012 implementation, written in Python, and the original, built-in IndMach012.
Check the `PyIndMach012.py` file for more comments. Comparing it to [the Pascal code for IndMach012](https://github.com/dss-extensions/dss_capi/blob/master/Version7/Source/PCElements/IndMach012.pas) can be useful to understand some of the inner workings of OpenDSS.
The user-model code in DSS Python is not stable yet but can be used to develop new ideas before commiting the final model in a traditional DLL user-model. Particularly, I (@PMeira) found some issues with callbacks with newer Version 8 COM DLLs, so changes related to that are expected.
```
%matplotlib inline
import os
import numpy as np
from matplotlib import pyplot as plt
from dss.UserModels import GenUserModel # used to get the DLL path
import PyIndMach012 # we need to import the model so it gets registered
```
## The model class
```
??PyIndMach012
```
## OpenDSS setup
For this example, we can use either COM or DSS Python (DSS C-API). The IndMach012 model in DSS C-API seems to have a bug somewhere though -- this is being tracked in [dss_capi#62](https://github.com/dss-extensions/dss_capi/issues/62).
```
original_dir = os.getcwd() # same the original working directory since the COM module messes with it
USE_COM = True # toggle this value to run with DSS C-API
if USE_COM:
from dss import patch_dss_com
import win32com.client
DSS = patch_dss_com(win32com.client.gencache.EnsureDispatch('OpenDSSengine.DSS'))
DSS.DataPath = original_dir
os.chdir(original_dir)
else:
from dss import DSS
DSS.Version
Text = DSS.Text
Monitors = DSS.ActiveCircuit.Monitors
```
## Using the model
To use a Python model for generators:
- the model class needs to be registered in advance
- create a generator with `model=6`
- pass a `usermodel="{dll_path}"` as in the following DSS command in the `run` function
- pass a `"pymodel=MODELNAME"` parameter in the userdata property, where MODELNAME is the name of the model class in Python
```
def run(pymodel):
Text.Command = 'redirect "master.dss"'
if pymodel:
# This uses our custom user-model in Python
Text.Command = 'New "Generator.Motor1" bus1=Bg2 kW=1200 conn=delta kVA=1500.000 H=6 model=6 kv=0.48 usermodel="{dll_path}" userdata=(pymodel=PyIndMach012 purs=0.048 puxs=0.075 purr=0.018 puxr=0.12 puxm=3.8 slip=0.02 SlipOption=variableslip)'.format(
dll_path=GenUserModel.dll_path,
)
Text.Command = 'New "Monitor.mfr2" element=Generator.Motor1 terminal=1 mode=3'
else:
# This uses the built-in model for comparison
Text.Command = 'New "IndMach012.Motor1" bus1=Bg2 kW=1200 conn=delta kVA=1500.000 H=6 purs=0.048 puxs=0.075 purr=0.018 puxr=0.12 puxm=3.8 slip=0.02 SlipOption=variableslip kv=0.48'
Text.Command = 'New "Monitor.mfr2" element=IndMach012.Motor1 terminal=1 mode=3'
# This will run a power-flow solution
Text.Command = 'Solve'
# This will toggle to the dynamics mode
Text.Command = 'Set mode=dynamics number=1 h=0.000166667'
# And finally run 5000 steps for the dynamic simulation
Text.Command = f'Solve number=5000'
# There are the channels from the Pascal/built-in IndMach012
channels_pas = (' Frequency', 'Theta (deg)', 'E1', 'dSpeed (deg/sec)', 'dTheta (deg)', 'Slip', 'Is1', 'Is2', 'Ir1', 'Ir2', 'Stator Losses', 'Rotor Losses', 'Shaft Power (hp)', 'Power Factor', 'Efficiency (%)')
# There are the channels from the Python module -- we define part of these and part come from the generator model itself
channels_py = (' Frequency', 'Theta (Deg)', 'E1_pu', 'dSpeed (Deg/sec)', 'dTheta (Deg)', 'Slip', 'Is1', 'Is2', 'Ir1', 'Ir2', 'StatorLosses', 'RotorLosses', 'ShaftPower_hp', 'PowerFactor', 'Efficiency_pct')
```
## Running and saving the outputs
Let's run the Pascal/built-in version of IndMach012 and our custom Python version for comparison:
```
run(False)
Monitors.Name = 'mfr2'
outputs_pas = {channel: Monitors.Channel(Monitors.Header.index(channel) + 1) for channel in channels_pas}
run(True)
Monitors.Name = 'mfr2'
outputs_py = {channel: Monitors.Channel(Monitors.Header.index(channel) + 1) for channel in channels_py}
time = np.arange(1, 5000 + 1) * 0.000166667
offset = int(0.1 / 0.000166667)
```
## Plotting the various output channels
The example circuit applies a fault at 0.3 s, isolating the machine at 0.4s (check `master.dss` for more details).
As we can see from the figures below, the outputs match very closely. After the induction machine is isolated, the efficiency and power factor values can misbehave as the power goes to zero, seem especially in the Pascal version.
```
for ch_pas, ch_py in zip(channels_pas, channels_py):
plt.figure(figsize=(8,4))
plt.plot(time, outputs_pas[ch_pas], label='IndMach012', lw=3)
plt.plot(time, outputs_py[ch_py], label='PyIndMach012', ls='--', lw=2)
plt.axvline(0.3, linestyle=':', color='k', alpha=0.5, label='Fault occurs')
plt.axvline(0.4, linestyle='--', color='r', alpha=0.5, label='Relays operate')
plt.legend()
plt.xlabel('Time (s)')
plt.ylabel(ch_pas)
if ch_pas == 'Efficiency (%)':
# Limit efficiency to 0-100
plt.ylim(0, 100)
plt.xlim(0, time[-1])
plt.tight_layout()
```
| github_jupyter |
# A Two-Level, Six-Factor Full Factorial Design
<br />
<br />
<br />
### Table of Contents
* [Introduction](#intro)
* Factorial Experimental Design:
* [Two-Level Six-Factor Full Factorial Design](#fullfactorial)
* [Variables and Variable Labels](#varlabels)
* [Computing Main and Interaction Effects](#computing_effects)
* Analysis of results:
* [Analyzing Effects](#analyzing_effects)
* [Quantile-Quantile Effects Plot](#quantile_effects)
* [Utilizing Degrees of Freedom](#dof)
* [Ordinary Least Squares Regression Model](#ols)
* [Goodness of Fit](#goodness_of_fit)
* [Distribution of Error](#distribution_of_error)
* [Aggregating Results](#aggregating)
* [Distribution of Variance](#dist_variance)
* [Residual vs. Response Plots](#residual)
<br />
<br />
<br />
<a name="intro"></a>
## Introduction
This notebook roughly follows content from Box and Draper's _Empirical Model-Building and Response Surfaces_ (Wiley, 1984). This content is covered by Chapter 4 of Box and Draper.
In this notebook, we'll carry out an anaylsis of a full factorial design, and show how we can obtain inforomation about a system and its responses, and a quantifiable range of certainty about those values. This is the fundamental idea behind empirical model-building and allows us to construct cheap and simple models to represent complex, nonlinear systems.
```
%matplotlib inline
import pandas as pd
import numpy as np
from numpy.random import rand, seed
import seaborn as sns
import scipy.stats as stats
from matplotlib.pyplot import *
seed(10)
```
<a name="fullfactorial"></a>
## Two-Level Six-Factor Full Factorial Design
Let's start with our six-factor factorial design example. Six factors means there are six input variables; this is still a two-level experiment, so this is now a $2^6$-factorial experiment.
Additionally, there are now three response variables, $(y_1, y_2, y_3)$.
To generate a table of the 64 experiments to be run at each factor level, we will use the ```itertools.product``` function below. This is all put into a DataFrame.
This example generates some random response data, by multiplying a vector of random numbers by the vector of input variable values. (Nothing too complicated.)
```
import itertools
# Create the inputs:
encoded_inputs = list( itertools.product([-1,1],[-1,1],[-1,1],[-1,1],[-1,1],[-1,1]) )
# Create the experiment design table:
doe = pd.DataFrame(encoded_inputs,columns=['x%d'%(i+1) for i in range(6)])
# "Manufacture" observed data y
doe['y1'] = doe.apply( lambda z : sum([ rand()*z["x%d"%(i)]+0.01*(0.5-rand()) for i in range(1,7) ]), axis=1)
doe['y2'] = doe.apply( lambda z : sum([ 5*rand()*z["x%d"%(i)]+0.01*(0.5-rand()) for i in range(1,7) ]), axis=1)
doe['y3'] = doe.apply( lambda z : sum([ 100*rand()*z["x%d"%(i)]+0.01*(0.5-rand()) for i in range(1,7) ]), axis=1)
print(doe[['y1','y2','y3']])
```
<a name="varlablels"></a>
## Defining Variables and Variable Labels
Next we'll define some containers for input variable labels, output variable labels, and any interaction terms that we'll be computing:
```
labels = {}
labels[1] = ['x1','x2','x3','x4','x5','x6']
for i in [2,3,4,5,6]:
labels[i] = list(itertools.combinations(labels[1], i))
obs_list = ['y1','y2','y3']
for k in labels.keys():
print(str(k) + " : " + str(labels[k]))
```
Now that we have variable labels for each main effect and interaction effect, we can actually compute those effects.
<a name="computing_effects"></a>
## Computing Main and Interaction Effects
We'll start by finding the constant effect, which is the mean of each response:
```
effects = {}
# Start with the constant effect: this is $\overline{y}$
effects[0] = {'x0' : [doe['y1'].mean(),doe['y2'].mean(),doe['y3'].mean()]}
print(effects[0])
```
Next, compute the main effect of each variable, which quantifies the amount the response changes by when the input variable is changed from the -1 to +1 level. That is, it computes the average effect of an input variable $x_i$ on each of the three response variables $y_1, y_2, y_3$.
```
effects[1] = {}
for key in labels[1]:
effects_result = []
for obs in obs_list:
effects_df = doe.groupby(key)[obs].mean()
result = sum([ zz*effects_df.ix[zz] for zz in effects_df.index ])
effects_result.append(result)
effects[1][key] = effects_result
effects[1]
```
Our next step is to crank through each variable interaction level: two-variable, three-variable, and on up to six-variable interaction effects. We compute interaction effects for each two-variable combination, three-variable combination, etc.
```
for c in [2,3,4,5,6]:
effects[c] = {}
for key in labels[c]:
effects_result = []
for obs in obs_list:
effects_df = doe.groupby(key)[obs].mean()
result = sum([ np.prod(zz)*effects_df.ix[zz]/(2**(len(zz)-1)) for zz in effects_df.index ])
effects_result.append(result)
effects[c][key] = effects_result
def printd(d):
for k in d.keys():
print("%25s : %s"%(k,d[k]))
for i in range(1,7):
printd(effects[i])
```
We've computed the main and interaction effects for every variable combination (whew!), but now we're at a point where we want to start doing things with these quantities.
<a name="analyzing_effects"></a>
## Analyzing Effects
The first and most important question is, what variable, or combination of variables, has the strongest effect on the three responses $y_1$? $y_2$? $y_3$?
To figure this out, we'll need to use the data we computed above. Python makes it easy to slice and dice data. In this case, we've constructed a nested dictionary, with the outer keys mapping to the number of variables and inner keys mapping to particular combinations of input variables. Its pretty easy to convert this to a flat data structure that we can use to sort by variable effects. We've got six "levels" of variable combinations, so we'll flatten ```effects``` by looping through all six dictionaries of variable combinations (from main effects to six-variable interaction effects), and adding each entry to a master dictionary.
The master dictionary will be a flat dictionary, and once we've populated it, we can use it to make a DataFrame for easier sorting, printing, manipulating, aggregating, and so on.
```
print(len(effects))
master_dict = {}
for nvars in effects.keys():
effect = effects[nvars]
for k in effect.keys():
v = effect[k]
master_dict[k] = v
master_df = pd.DataFrame(master_dict).T
master_df.columns = obs_list
y1 = master_df['y1'].copy()
y1.sort_values(inplace=True,ascending=False)
print("Top 10 effects for observable y1:")
print(y1[:10])
y2 = master_df['y2'].copy()
y2.sort_values(inplace=True,ascending=False)
print("Top 10 effects for observable y2:")
print(y2[:10])
y3 = master_df['y3'].copy()
y3.sort_values(inplace=True,ascending=False)
print("Top 10 effects for observable y3:")
print(y3[:10])
```
If we were only to look at the list of rankings of each variable, we would see that each response is affected by different input variables, listed below in order of descending importance:
* $y_1$: 136254
* $y_2$: 561234
* $y_3$: 453216
This is a somewhat mixed message that's hard to interpret - can we get rid of variable 2? We can't eliminate 1, 4, or 5, and probably not 3 or 6 either.
However, looking at the quantile-quantile plot of the effects answers the question in a more visual way.
<a name="quantile_effects"></a>
## Quantile-Quantile Effects Plot
We can examine the distribution of the various input variable effects using a quantile-quantile plot of the effects. Quantile-quantile plots arrange the effects in order from least to greatest, and can be applied in several contexts (as we'll see below, when assessing model fits). If the quantities plotted on a quantile-qantile plot are normally distributed, they will fall on a straight line; data that do not fall on the straight line indicate significant deviations from normal behavior.
In the case of a quantile-quantile plot of effects, non-normal behavior means the effect is paticularly strong. By identifying the outlier points on thse quantile-quantile plots (they're ranked in order, so they correspond to the lists printed above), we can identify the input variables most likely to have a strong impact on the responses.
We need to look both at the top (the variables that have the largest overall positive effect) and the bottom (the variables that have the largest overall negative effect) for significant outliers. When we find outliers, we can add them to a list of variabls that we have decided are important and will keep in our analysis.
```
# Quantify which effects are not normally distributed,
# to assist in identifying important variables
fig = figure(figsize=(14,4))
ax1 = fig.add_subplot(131)
ax2 = fig.add_subplot(132)
ax3 = fig.add_subplot(133)
stats.probplot(y1, dist="norm", plot=ax1)
ax1.set_title('y1')
stats.probplot(y2, dist="norm", plot=ax2)
ax2.set_title('y2')
stats.probplot(y3, dist="norm", plot=ax3)
ax3.set_title('y3')
```
Normally, we would use the main effects that were computed, and their rankings, to eliminate any variables that don't have a strong effect on any of our variables. However, this analysis shows that sometimes we can't eliminate any variables.
All six input variables are depicted as the effects that fall far from the red line - indicating all have a statistically meaningful (i.e., not normally distributed) effect on all three response variables. This means we should keep all six factors in our analysis.
There is also a point on the $y_3$ graph that appears significant on the bottom. Examining the output of the lists above, this point represents the effect for the six-way interaction of all input variables. High-order interactions are highly unlikely (and in this case it is a numerical artifact of the way the responses were generated), so we'll keep things simple and stick to a linear model.
Let's continue our analysis without eliminating any of the six factors, since they are important to all of our responses.
<a name="dof"></a>
## Utilizing Degrees of Freedom
Our very expensive, 64-experiment full factorial design (the data for which maps $(x_1,x_2,\dots,x_6)$ to $(y_1,y_2,y_3)$) gives us 64 data points, and 64 degrees of freedom. What we do with those 64 degrees of freedom is up to us.
We _could_ fit an empirical model, or response surface, that has 64 independent parameters, and account for many of the high-order interaction terms - all the way up to six-variable interaction effects. However, high-order effects are rarely important, and are a waste of our degrees of freedom.
Alternatively, we can fit an empirical model with fewer coefficients, using up fewer degrees of freedom, and use the remaining degrees of freedom to characterize the error introduced by our approximate model.
To describe a model with the 6 variables listed above and no other variable interaction effects would use only 6 degrees of freedom, plus 1 degree of freedom for the constant term, leaving 57 degrees of freedom available to quantify error, attribute variance, etc.
Our goal is to use least squares to compute model equations for $(y_1,y_2,y_3)$ as functions of $(x_1,x_2,x_3,x_4,x_5,x_6)$.
```
xlabs = ['x1','x2','x3','x4','x5','x6']
ylabs = ['y1','y2','y3']
ls_data = doe[xlabs+ylabs]
import statsmodels.api as sm
import numpy as np
x = ls_data[xlabs]
x = sm.add_constant(x)
```
The first ordinary least squares linear model is created to predict values of the first variable, $y_1$, as a function of each of our input variables, the list of which are contained in the ```xlabs``` variable. When we perform the linear regression fitting, we see much of the same information that we found in the prior two-level three-factor full factorial design, but here, everything is done automatically.
The model is linear, meaning it's fitting the coefficients of the function:
$$
\hat{y} = a_0 + a_1 x_1 + a_2 x_2 + a_3 + x_3 + a_4 x_4 + a_5 x_5 + a_6 x_6
$$
(here, the variables $y$ and $x$ are vectors, with one component for each response; in our case, they are three-dimensional vectors.)
Because there are 64 observations and 7 coefficients, the 57 extra observations give us extra degrees of freedom with which to assess how good the model is. That analysis can be done with an ordinary least squares (OLS) model, available through the statsmodel library in Python.
<a name="ols"></a>
## Ordinary Least Squares Regression Model
This built-in OLS model will fit an input vector $(x_1,x_2,x_3,x_4,x_5,x_6)$ to an output vector $(y_1,y_2,y_3)$ using a linear model; the OLS model is designed to fit the model with more observations than coefficients, and utilize the remaining data to quantify the fit of the model.
Let's run through one of these, and analyze the results:
```
y1 = ls_data['y1']
est1 = sm.OLS(y1,x).fit()
print(est1.summary())
```
The StatsModel OLS object prints out quite a bit of useful information, in a nicely-formatted table. Starting at the top, we see a couple of important pieces of information: specifically, the name of the dependent variable (the response) that we're looking at, the number of observations, and the number of degrees of freedom.
We can see an $R^2$ statistic, which indicates how well this data is fit with our linear model, and an adjusted $R^2$ statistic, which accounts for the large nubmer of degrees of freedom. While an adjusted $R^2$ of 0.73 is not great, we have to remember that this linear model is trying to capture a wealth of complexity in six coefficients. Furthermore, the adjusted $R^2$ value is too broad to sum up how good our model actually is.
The table in the middle is where the most useful information is located. The `coef` column shows the coefficients $a_0, a_1, a_2, \dots$ for the model equation:
$$
\hat{y} = a_0 + a_1 x_1 + a_2 x_2 + a_3 + x_3 + a_4 x_4 + a_5 x_5 + a_6 x_6
$$
Using the extra degrees of freedom, an estime $s^2$ of the variance in the regression coefficients is also computed, and reported in the the `std err` column. Each linear term is attributed the same amount of variance, $\pm 0.082$.
```
y2 = ls_data['y2']
est2 = sm.OLS(y2,x).fit()
print(est2.summary())
y3 = ls_data['y3']
est3 = sm.OLS(y3,x).fit()
print(est3.summary())
```
<a name="goodness_of_fit"></a>
## Quantifying Model Goodness-of-Fit
We can now use these linear models to evaluate each set of inputs and compare the model response $\hat{y}$ to the actual observed response $y$. What we would expect to see, if our model does an adequate job of representing the underlying behavior of the model, is that in each of the 64 experiments, the difference between the model prediction $M$ and the measured data $d$, defined as the residual $r$,
$$
r = \left| d - M \right|
$$
should be comparable across all experiments. If the residuals appear to have functional dependence on the input variables, it is an indication that our model is missing important effects and needs more or different terms. The way we determine this, mathematically, is by looking at a quantile-quantile plot of our errors (that is, a ranked plot of our error magnitudes).
If the residuals are normally distributed, they will follow a straight line; if the plot shows the data have significant wiggle and do not follow a line, it is an indication that the errors are not normally distributed, and are therefore skewed (indicating terms missing from our OLS model).
```
%matplotlib inline
import seaborn as sns
import scipy.stats as stats
from matplotlib.pyplot import *
# Quantify goodness of fit
fig = figure(figsize=(14,4))
ax1 = fig.add_subplot(131)
ax2 = fig.add_subplot(132)
ax3 = fig.add_subplot(133)
r1 = y1 - est1.predict(x)
r2 = y2 - est2.predict(x)
r3 = y3 - est3.predict(x)
stats.probplot(r1, dist="norm", plot=ax1)
ax1.set_title('Residuals, y1')
stats.probplot(r2, dist="norm", plot=ax2)
ax2.set_title('Residuals, y2')
stats.probplot(r3, dist="norm", plot=ax3)
ax3.set_title('Residuals, y3')
```
Determining whether significant trends are being missed by the model depends on how many points deviate from the red line, and how significantly. If there is a single point that deviates, it does not necessarily indicate a problem; but if there is significant wiggle and most points deviate significantly from the red line, it means that there is something about the relationship between the inputs and the outputs that our model is missing.
There are only a few points deviating from the red line. We saw from the effect quantile for $y_3$ that there was an interaction variable that was important to modeling the response $y_3$, and it is likely this interaction that is leading to noise at the tail end of these residuals. This indicates residual errors (deviations of the model from data) that do not follow a natural, normal distribution, which indicates there is a _pattern_ in the deviations - namely, the interaction effect.
The conclusion about the error from the quantile plots above is that there are only a few points deviation from the line, and no particularly significant outliers. Our model can use some improvement, but it's a pretty good first-pass model.
<a name="distribution_of_error"></a>
## Distribution of Error
Another thing we can look at is the normalized error: what are the residual errors (differences between our model prediction and our data)? How are their values distributed?
A kernel density estimate (KDE) plot, which is a smoothed histogram, shows the probability distribution of the normalized residual errors. As expected, they're bunched pretty close to zero. There are some bumps far from zero, corresponding to the outliers on the quantile-quantile plot of the errors above. However, they're pretty close to randomly distributed, and therefore it doesn't look like there is any systemic bias there.
```
fig = figure(figsize=(10,12))
ax1 = fig.add_subplot(311)
ax2 = fig.add_subplot(312)
ax3 = fig.add_subplot(313)
axes = [ax1,ax2,ax3]
colors = sns.xkcd_palette(["windows blue", "amber", "faded green", "dusty purple","aqua blue"])
#resids = [r1, r2, r3]
normed_resids = [r1/y1, r2/y2, r3/y3]
for (dataa, axx, colorr) in zip(normed_resids,axes,colors):
sns.kdeplot(dataa, bw=1.0, ax=axx, color=colorr, shade=True, alpha=0.5);
ax1.set_title('Probability Distribution: Normalized Residual Error, y1')
ax2.set_title('Normalized Residual Error, y2')
ax3.set_title('Normalized Residual Error, y3')
```
Note that in these figures, the bumps at extreme value are caused by the fact that the interval containing the responses includes 0 and values close to 0, so the normalization factor is very tiny, leading to large values.
<a name="aggregating"></a>
## Aggregating Results
Let's next aggregate experimental results, by taking the mean over various variables to compute the mean effect for regressed varables. For example, we may want to look at the effects of variables 2, 3, and 4, and take the mean over the other three variables.
This is simple to do with Pandas, by grouping the data by each variable, and applying the mean function on all of the results. The code looks like this:
```
# Our original regression variables
xlabs = ['x2','x3','x4']
doe.groupby(xlabs)[ylabs].mean()
# If we decided to go for a different variable set
xlabs = ['x2','x3','x4','x6']
doe.groupby(xlabs)[ylabs].mean()
```
This functionality can also be used to determine the variance in all of the experimental observations being aggregated. For example, here we aggregate over $x_3 \dots x_6$ and show the variance broken down by $x_1, x_2$ vs $y_1, y_2, y_3$.
```
xlabs = ['x1','x2']
doe.groupby(xlabs)[ylabs].var()
```
Or even the number of experimental observations being aggregated!
```
doe.groupby(xlabs)[ylabs].count()
```
<a name="dist_variance"></a>
## Distributions of Variance
We can convert these dataframes of averages, variances, and counts into data for plotting. For example, if we want to make a histogram of every value in the groupby dataframe, we can use the ```.values``` method, so that this:
doe.gorupby(xlabs)[ylabs].mean()
becomes this:
doe.groupby(xlabs)[ylabs].mean().values
This $M \times N$ array can then be flattened into a vector using the ```ravel()``` method from numpy:
np.ravel( doe.groupby(xlabs)[ylabs].mean().values )
The resulting data can be used to generate histograms, as shown below:
```
# Histogram of means of response values, grouped by xlabs
xlabs = ['x1','x2','x3','x4']
print("Grouping responses by %s"%( "-".join(xlabs) ))
dat = np.ravel(doe.groupby(xlabs)[ylabs].mean().values) / np.ravel(doe.groupby(xlabs)[ylabs].var().values)
hist(dat, 10, normed=False, color=colors[3]);
xlabel(r'Relative Variance ($\mu$/$\sigma^2$)')
show()
# Histogram of variances of response values, grouped by xlabs
print("Grouping responses by %s"%( "-".join(xlabs) ))
dat = np.ravel(doe.groupby(xlabs)['y1'].var().values)
hist(dat, normed=True, color=colors[4])
xlabel(r'Variance in $y_{1}$ Response')
ylabel(r'Frequency')
show()
```
The distribution of variance looks _mostly_ normal, with some outliers. These are the same outliers that showed up in our quantile-quantile plot, and they'll show up in the plots below as well.
<a name="residual"></a>
## Residual vs. Response Plots
Another thing we can do, to look for uncaptured effects, is to look at our residuals vs. $\hat{y}$. This is a further effort to look for underlying functional relationships between $\hat{y}$ and the residuals, which would indicate that our system exhibits behavior not captured by our linear model.
```
# normal plot of residuals
fig = figure(figsize=(14,4))
ax1 = fig.add_subplot(131)
ax2 = fig.add_subplot(132)
ax3 = fig.add_subplot(133)
ax1.plot(y1,r1,'o',color=colors[0])
ax1.set_xlabel('Response value $y_1$')
ax1.set_ylabel('Residual $r_1$')
ax2.plot(y2,r2,'o',color=colors[1])
ax2.set_xlabel('Response value $y_2$')
ax2.set_ylabel('Residual $r_2$')
ax2.set_title('Response vs. Residual Plots')
ax3.plot(y1,r1,'o',color=colors[2])
ax3.set_xlabel('Response value $y_3$')
ax3.set_ylabel('Residual $r_3$')
show()
```
Notice that each plot is trending up and to the right - indicative of an underlying trend that our model $\hat{y}$ is not capturing. The trend is relatively weak, however, indicating that our linear model does a good job of capturing _most_ of the relevant effects of this system.
# Discussion
The analysis shows that there are some higher-order or nonlinear effects in the system that a purely linear model does not account for. Next steps would involve adding higher order points for a quadratic or higher order polynomial model to gather additional data to fit the higher-degree models.
| github_jupyter |
<a href="https://colab.research.google.com/github/NikolaZubic/AppliedGameTheoryHomeworkSolutions/blob/main/domaci3.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# TREĆI DOMAĆI ZADATAK iz predmeta "Primenjena teorija igara" (Applied Game Theory)
Razvoj bota za igranje igre Ajnc (BlackJack) koristeći "Q-learning" pristup.
# Potrebni import-i
```
import gym
import numpy as np
import matplotlib.pyplot as plt
from gym import spaces
import seaborn as sns
```
# Definisanje Ajnc okruženja koristeći "Open AI Gym" toolkit
```
class BlackJackEnvironment(gym.Env):
# Because of human-friendly output
metadata = {'render.modes':['human']}
def __init__(self):
"""
We will define possible number of states with observation_space.
Player's sum can go from 4 to 32: Now when the sum is 22, and the player chooses to hit, he may get a card with value 10, resulting in a sum of 32, and thus loosing the game.
Dealer's card can be from 1 to 10 and we have 2 actions.
Total number of states: 29 * 10 * 2 = 580
Total number of actions = 2 = len( {"HIT", "STAND"} )
"""
self.observation_space = spaces.Discrete(580)
self.action_space = spaces.Discrete(2)
self.step_count = 0 # at the beginning of the game we have 0 actions taken
def check_usable_ace(self,hand):
"""
If someone has an usable ace, we will replace that ace (1) with 11.
:param hand: player's or dealer's card
:return: True if we have usable ace, False otherwise
"""
temp_hand = hand.copy()
# Check if there is ace in hand
if np.any(temp_hand == 1):
# If we have any ace then replace it with 11, but if we have more than one ace replace the first one with 11
temp_hand[np.where(temp_hand == 1)[0][0]] = 11
# If the sum is less or equal than 21 then we can use it
if temp_hand.sum() <= 21:
return True
return False
def use_ace(self,hand):
"""
If there is usable ace in function above, then replace 1 with 11.
:param hand: player's or dealer's hand
:return: new hand where 1 is replaced with 11
"""
temp_hand = hand.copy()
temp_hand[np.where(temp_hand == 1)[0][0]] = 11
return temp_hand
def reset(self):
# Resets the environment after one game.
# Initialize player's hand
self.current_hand = np.random.choice(range(1,11),2)
# Initialize usable Ace to False, since we don't have it at the very beginning
self.usable_ace = False
self.dealer_stand, self.player_stand = False, False
# Replace usable ace in the player's hand
if self.check_usable_ace(self.current_hand):
self.usable_ace = True
self.current_hand = self.use_ace(self.current_hand)
# Player's current sum
self.current_sum = self.current_hand.sum()
# Dealer's hand
self.dealer_hand = np.random.choice(range(1,11),2)
# Dealer's sum
self.dealer_sum = self.dealer_hand.sum()
# First element of self.dealer_hand is the current showing card of dealer
self.dealer_showing_card = self.dealer_hand[0]
# Replace usable ace in the dealer's hand
if self.check_usable_ace(self.dealer_hand):
temp_dealer_hand = self.use_ace(self.dealer_hand)
self.dealer_sum = temp_dealer_hand.sum()
def take_turn(self, current_player):
"""
Play one turn for the player. This function will be called from step() function, directly depending on the game state.
We will take new random card, add it to the current_player hand.
:param player: {"player", "dealer"}
:return: None
"""
if current_player == 'dealer':
# Take new random card
new_card = np.random.choice(range(1,11))
# Add new card to the current_player hand
new_dealer_hand = np.array(self.dealer_hand.tolist() + [new_card])
# Check for usable ace and replace if found
if self.check_usable_ace(new_dealer_hand):
new_dealer_hand = self.use_ace(new_dealer_hand)
self.dealer_hand = new_dealer_hand
# Update his sum
self.dealer_sum = self.dealer_hand.sum()
if current_player == 'player':
new_card = np.random.choice(range(1,11))
new_player_hand = np.array(self.current_hand.tolist()+ [new_card])
if self.check_usable_ace(new_player_hand):
self.usable_ace = True
new_player_hand = self.use_ace(new_player_hand)
self.current_hand = new_player_hand
self.current_sum = self.current_hand.sum()
def check_game_status(self, mode = 'normal'):
"""
Check the current status of the game.
During the 'normal' we check after each turn whether we got in the terminal state.
In the 'compare' mode we compare the totals of both players (player vs dealer) in order to pronounce the winner.
:param mode: {'normal', 'compare'}
:return: dictionary with the winner, whether the game is finished and the reward of the game
"""
result = {'winner':'',
'is_done': False,
'reward':0}
if mode == 'normal':
if self.current_sum > 21:
result['winner'] = 'dealer'
result['is_done'] = True
result['reward'] = -1
elif self.dealer_sum > 21:
result['winner'] = 'player'
result['is_done'] = True
result['reward'] = 1
elif self.current_sum == 21:
result['winner'] = 'player'
result['is_done'] = True
result['reward'] = 1
elif self.dealer_sum == 21:
result['winner'] = 'dealer'
result['is_done'] = True
result['reward'] = -1
elif mode == 'compare':
result['is_done'] = True
diff_21_player = 21 - self.current_sum
diff_21_dealer = 21 - self.dealer_sum
if diff_21_player > diff_21_dealer:
result['reward'] = -1
result['winner'] = 'dealer'
elif diff_21_player < diff_21_dealer:
result['reward'] = 1
result['winner'] = 'player'
else:
result['reward'] = 0
result['winner'] = 'draw'
return result
return result
def step(self,action):
"""
Performs one action.
:param action:
:return: dictionary with the winner, whether the game is finished and the reward of the game
"""
# Increase number of actions that are taken during the game.
self.step_count += 1
result = {'winner':'',
'is_done': False,
'reward':0}
"""
Before taking the first step of the game, we need to ensure that there is no winning condition.
Check if the initial two cards of the players are 21. If anyone has 21, then that player wins.
If both players have 21, then the game is DRAW. Otherwise, we will continue with the game.
"""
if self.step_count == 1:
if self.check_usable_ace(self.current_hand):
self.current_hand = self.use_ace(self.current_hand)
if self.check_usable_ace(self.dealer_hand):
self.current_hand = self.use_ace(self.dealer_hand)
if self.current_sum == 21 and self.dealer_sum == 21:
result['is_done'] = True
result['reward'] = 0
result['winner'] = 'draw'
return result
elif self.current_sum == 21 and self.dealer_sum < 21:
result['is_done'] = True
result['reward'] = 1
result['winner'] = 'player'
return result
elif self.dealer_sum == 21 and self.current_sum < 21:
result['is_done'] = True
result['reward'] = -1
result['winner'] = 'dealer'
return result
if self.dealer_sum >= 17:
self.dealer_stand = True
# action = 0 means "HIT"
if action == 0:
self.take_turn('player')
result = self.check_game_status()
if result['is_done'] == True:
return result
# action = 1 means "STAND"
if action == 1:
if self.dealer_stand == True:
return self.check_game_status(mode = 'compare')
"""
If the dealer hasn't stand, he will hit unless his sum is greater than or equal to 17.
After that, he will stand.
"""
while self.dealer_sum < 17:
self.take_turn('dealer')
result = self.check_game_status()
# After dealer stands, check the game status.
if result['is_done'] == True:
return result
# If the game hasn't finished yet, we set dealer_stand to True, so the player will either HIT or STAND
self.dealer_stand = True
return result
def get_current_state(self):
"""
Get current state which is comprised of current player's sum, dealer's showing card and usable ace presence.
:return: return current state variables
"""
current_state = {}
current_state['current_sum'] = self.current_sum
current_state['dealer_showing_card'] = self.dealer_showing_card
current_state['usable_ace'] = self.usable_ace
return current_state
def render(self):
print("OBSERVABLE STATES")
print("Current player's sum: {}".format(self.current_sum))
print("Dealer's showing card: {}".format(self.dealer_showing_card))
print("Player has usable Ace: {}".format(self.usable_ace))
print("INFORMATION ABOUT CARDS AND DEALER'S SUM")
print("Player's hand: {}".format(self.current_hand))
print("Dealer's hand: {}".format(self.dealer_hand))
print("Dealer's sum: {}".format(self.dealer_sum))
```
# Pomoćne funkcije za Q-learning
```
# dictionaries used for converting the state values to indexes in the Q table
current_sum_to_index = dict(zip(np.arange(4,33),np.arange(29)))
dealer_showing_card_to_index = dict(zip(np.arange(1,11),np.arange(10)))
usable_ace_index = dict(zip([False,True],[0,1]))
action_index = dict(zip(['HIT','STAND'],[0,1]))
def get_state_q_indices(current_state):
"""
Get indexes of Q table for any given state.
:param current_state: comprised of current player's sum, dealer's showing card and usable ace presence.
:return: get table indexes for a state
"""
current_sum_idx = current_sum_to_index[current_state['current_sum']]
dealer_showing_card_idx = dealer_showing_card_to_index[current_state['dealer_showing_card']]
usable_ace_idx = usable_ace_index[current_state['usable_ace']]
return [current_sum_idx,dealer_showing_card_idx,usable_ace_idx]
def get_max_action(Q_sa, current_state):
"""
Get the action with the max Q-value for the given current state and the Q table.
:param Q_sa: given Q table
:param current_state: current state
:return: best action for given state and Q table
"""
state_q_idxs = get_state_q_indices(current_state)
action = Q_sa[state_q_idxs[0],state_q_idxs[1],state_q_idxs[2],:].argmax()
return action
def get_q_value(Q_sa, state, action):
"""
Get Q(s,a) value for state and action in certain Q table.
:param Q_sa: given Q table
:param state: given state
:param action: given action
:return: Q(s, a)
"""
state_q_idxs = get_state_q_indices(state)
q_value = Q_sa[state_q_idxs[0],state_q_idxs[1],state_q_idxs[2],action]
return q_value
```
# Q-learning
Inicijalizacija Q tabele.
```
"""
Player's current sum is ranging from 4 to 32 => 32 - 4 + 1 = 29
Dealer's showing card can be one from the following set {1, 2, 3, 4, 5, 6, 7, 8, 9, 10} => 10 values
Ace can be usable or not => 2
Actions are from the following set {"HIT", "STAND"} => 2
"""
Q = np.zeros((29,10,2,2))
```
Proces treniranja.
```
episode_count = 0
total_episodes = 2000000
# Discounting factor
gamma = 0.9
# Used for filtering q-values, learning rate
LAMBDA = 0.1
# Defined Black Jack Environment
environment = BlackJackEnvironment()
while episode_count < total_episodes:
environment.reset()
current_state = environment.get_current_state()
current_action = get_max_action(Q, current_state)
# Take action
step_result = environment.step(current_action)
# Get into next state and get the reward
next_state = environment.get_current_state()
next_max_action = get_max_action(Q, next_state)
immediate_reward = step_result['reward']
next_state_q_idxs = get_state_q_indices(next_state)
# Get the q-value for the next state and max action in the next state
q_max_s_a = get_q_value(Q, next_state, next_max_action)
td_target = immediate_reward + gamma * q_max_s_a
# Get the q-value for the current state and action
q_current_s_a = get_q_value(Q, current_state, current_action)
td_error = td_target - q_current_s_a
state_q_idxs = get_state_q_indices(current_state)
# Update the current Q(s, a)
Q[state_q_idxs[0],state_q_idxs[1],state_q_idxs[2],current_action] = q_current_s_a + LAMBDA * td_error
# get into the next state
current_state = next_state
if step_result['is_done']:
episode_count += 1
if episode_count % 100000 == 0:
print("Episode number: {}".format(episode_count))
```
# Diskusija rezultata
```
fig, ax = plt.subplots(ncols= 2,figsize=(16,8))
sns.heatmap(Q[:,:,0,0],cmap = sns.light_palette((210, 90, 60), input="husl"), ax = ax[0],
xticklabels=np.arange(1,11),yticklabels=np.arange(4,33))
ax[0].set_title("Usable Ace = False, Action = HIT")
ax[0].set_xlabel("Dealer's Showing Card")
ax[0].set_ylabel("Current Player's Sum")
sns.heatmap(Q[:,:,0,1],cmap = sns.light_palette((210, 90, 60), input="husl"), ax = ax[1],
xticklabels=np.arange(1,11),yticklabels=np.arange(4,33))
ax[1].set_title("Usable Ace = False, Action = STAND")
ax[1].set_xlabel("Dealer's Showing Card")
ax[1].set_ylabel("Current Player's Sum")
```
Na osnovu gornjih heatmapa možemo uočiti koje je to akcije dobro izvršiti u kojem stanju.
**Zaključak sa lijeve heatmape**: kada je ukupna suma igrača manja od 12, 13 onda je najbolje da se izvršava akcija "HIT".
**Zaključak sa desne heatmape**: Za veće vrijednosti otkrivene karte djelitelja i veće vrijednosti ukupne sume igrača bolje je izvršiti akciju "STAND".
```
fig, ax = plt.subplots(ncols = 2, figsize=(16,8))
sns.heatmap(Q[:,:,1,0],cmap = sns.light_palette((210, 90, 60), input="husl"), ax = ax[0],
xticklabels=np.arange(1,11),yticklabels=np.arange(4,33))
ax[0].set_title("Usable Ace = True, Action = HIT")
ax[0].set_xlabel("Dealer's Showing Card")
ax[0].set_ylabel("Current Player's Sum")
sns.heatmap(Q[:,:,1,1],cmap = sns.light_palette((210, 90, 60), input="husl"), ax = ax[1],
xticklabels=np.arange(1,11),yticklabels=np.arange(4,33))
ax[1].set_title("Usable Ace = True, Action = STAND")
ax[1].set_xlabel("Dealer's Showing Card")
ax[1].set_ylabel("Current Player's Sum")
```
U slučaju kad imamo iskoristiv kec, broj semplova je znatno manji, tako da paterni Q-vrijednosti nisu baš potpuno jasni, ali može se zaključiti da je najbolje izvršiti akciju **"HIT" u slučajevima kad je suma igrača oko 12**, dok se akcija **"STAND" izvršava u slučaju kada je igra pri kraju po pitanju sume igrača**.
Sada ćemo pogledati naučene politike (za slučaj pohlepne politike, jer želimo da naš igrač uvijek bira onako da najbolje igra).
**Sa crnim blokovima označeno je kada treba izvršiti akciju "HIT"**, a imamo 2 heatmape za slučaj kad nemamo i imamo iskoristiv kec.
```
fig, ax = plt.subplots(ncols= 1,figsize=(8,6))
sns.heatmap(np.argmax(Q[:17,:,0,:],axis=2),cmap = sns.cubehelix_palette(8, start=2, rot=0, dark=0, light=.95, reverse=True)\
,linewidths=1,xticklabels=np.arange(1,11),yticklabels=np.arange(4,33))
ax.set_title("Usable Ace = False")
ax.set_xlabel("Dealer's Showing Card")
ax.set_ylabel("Current Player's Sum")
fig, ax = plt.subplots(ncols= 1,figsize=(8,6))
sns.heatmap(np.argmax(Q[:17,:,1,:],axis=2),cmap = sns.cubehelix_palette(8, start=2, rot=0, dark=0, light=.95, reverse=True)\
,linewidths=1,xticklabels=np.arange(1,11),yticklabels=np.arange(4,33))
ax.set_title("Usable Ace = True")
ax.set_xlabel("Dealer's Showing Card")
ax.set_ylabel("Current Player's Sum")
```
# Na kraju, nakon 2 miliona iteracija treniranja, testiraćemo algoritam na 10 000 partija.
```
player_wins = 0
dealer_wins = 0
NUMBER_OF_GAMES = 10000
for i in range(NUMBER_OF_GAMES):
environment.reset()
while True:
current_state = environment.get_current_state()
current_action = get_max_action(Q, current_state)
# Take action
step_result = environment.step(current_action)
#environment.render()
next_state = environment.get_current_state()
current_state = next_state
if step_result['is_done']:
break
if step_result['winner'] == 'player':
player_wins += 1
elif step_result['winner'] == 'dealer':
dealer_wins += 1
print("Player wins: " + str(player_wins))
print("Dealer wins: " + str(dealer_wins))
print("Player wins percentage = " + str(round(100 * (player_wins / (player_wins + dealer_wins)), 2)) + "%")
```
| github_jupyter |
```
# Copyright 2021 NVIDIA Corporation. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Accelerate BERT encoder with TF-TRT
## Introduction
The NVIDIA TensorRT is a C++ library that facilitates high performance inference on NVIDIA graphics processing units (GPUs). TensorFlow™ integration with TensorRT™ (TF-TRT) optimizes TensorRT compatible parts of your computation graph, allowing TensorFlow to execute the remaining graph. While you can use TensorFlow's wide and flexible feature set, TensorRT will produce a highly optimized runtime engine for the TensorRT compatible subgraphs of your network.
In this notebook, we demonstrate accelerating BERT inference using TF-TRT. We focus on the encoder.
## Requirements
This notebook requires at least TF 2.5 and TRT 7.1.3.
## 1. Download the model
We will download a bert base model from [TF-Hub](https://tfhub.dev/tensorflow/bert_en_uncased_L-12_H-768_A-12/3).
```
!pip install -q tf-models-official
import tensorflow as tf
import tensorflow_hub as hub
tfhub_handle_encoder = 'https://tfhub.dev/tensorflow/bert_en_uncased_L-12_H-768_A-12/3'
bert_saved_model_path = 'bert_base'
bert_model = hub.load(tfhub_handle_encoder)
tf.saved_model.save(bert_model, bert_saved_model_path)
```
## 2. Inference
In this section we will convert the model using TF-TRT and run inference.
```
import matplotlib.pyplot as plt
import numpy as np
from tensorflow.python.saved_model import signature_constants
from tensorflow.python.saved_model import tag_constants
from tensorflow.python.compiler.tensorrt import trt_convert as trt
from timeit import default_timer as timer
tf.get_logger().setLevel('ERROR')
```
### 2.1 Helper functions
```
def get_func_from_saved_model(saved_model_dir):
saved_model_loaded = tf.saved_model.load(
saved_model_dir, tags=[tag_constants.SERVING])
graph_func = saved_model_loaded.signatures[
signature_constants.DEFAULT_SERVING_SIGNATURE_DEF_KEY]
return graph_func, saved_model_loaded
def predict_and_benchmark_throughput(input_dict, model, N_warmup_run=50, N_run=500,
result_key='predictions', batch_size=None):
elapsed_time = []
for val in input_dict.values():
input_batch_size = val.shape[0]
break
if batch_size is None or batch_size > input_batch_size:
batch_size = input_batch_size
print('Benchmarking with batch size', batch_size)
elapsed_time = np.zeros(N_run)
for i in range(N_warmup_run):
preds = model(**input_dict)
# Force device synchronization with .numpy()
tmp = preds[result_key][0].numpy()
for i in range(N_run):
start_time = timer()
preds = model(**input_dict)
# Synchronize
tmp += preds[result_key][0].numpy()
end_time = timer()
elapsed_time[i] = end_time - start_time
if i>=50 and i % 50 == 0:
print('Steps {}-{} average: {:4.1f}ms'.format(i-50, i, (elapsed_time[i-50:i].mean()) * 1000))
latency = elapsed_time.mean() * 1000
print('Latency: {:5.2f}+/-{:4.2f}ms'.format(latency, elapsed_time.std() * 1000))
print('Throughput: {:.0f} samples/s'.format(N_run * batch_size / elapsed_time.sum()))
return latency
def trt_convert(input_path, output_path, input_shapes, explicit_batch=False,
dtype=np.float32, precision='FP32', prof_strategy='Optimal'):
conv_params=trt.TrtConversionParams(
precision_mode=precision, minimum_segment_size=50,
max_workspace_size_bytes=12*1<<30, maximum_cached_engines=1)
converter = trt.TrtGraphConverterV2(
input_saved_model_dir=input_path, conversion_params=conv_params,
use_dynamic_shape=explicit_batch,
dynamic_shape_profile_strategy=prof_strategy)
converter.convert()
def input_fn():
for shapes in input_shapes:
# return a list of input tensors
yield [np.ones(shape=x).astype(dtype) for x in shapes]
converter.build(input_fn)
converter.save(output_path)
def random_input(batch_size, seq_length):
# Generate random input data
mask = tf.convert_to_tensor(np.ones((batch_size, seq_length), dtype=np.int32))
type_id = tf.convert_to_tensor(np.zeros((batch_size, seq_length), dtype=np.int32))
word_id = tf.convert_to_tensor(np.random.randint(0, 1000, size=[batch_size, seq_length], dtype=np.int32))
return {'input_mask':mask, 'input_type_ids': type_id, 'input_word_ids':word_id}
```
### 2.2 Convert the model with TF-TRT
```
bert_trt_path = bert_saved_model_path + '_trt'
input_shapes = [[(1, 128), (1, 128), (1, 128)]]
trt_convert(bert_saved_model_path, bert_trt_path, input_shapes, True, np.int32, precision='FP16')
```
### 2.3 Run inference with converted model
```
trt_func, _ = get_func_from_saved_model(bert_trt_path)
input_dict = random_input(1, 128)
result_key = 'bert_encoder_1' # 'classifier'
res = predict_and_benchmark_throughput(input_dict, trt_func, result_key=result_key)
```
### Compare to the original function
```
func, model = get_func_from_saved_model(bert_saved_model_path)
res = predict_and_benchmark_throughput(input_dict, func, result_key=result_key)
```
## 3. Dynamic sequence length
The sequence length for the encoder is dynamic, we can use different input sequence lengths. Here we call the original model for two sequences.
```
seq1 = random_input(1, 128)
res1 = func(**seq1)
seq2 = random_input(1, 180)
res2 = func(**seq2)
```
The converted model is optimized for a sequnce length of 128 (and batch size 8). If we infer the converted model using a different sequence length, then two things can happen:
1. If `TrtConversionParams.allow_build_at_runtime` == False: native TF model is inferred
2. if `TrtConversionParams.allow_build_at_runtime` == True a new TRT engine is created which is optimized for the new sequence length.
The first option do not provide TRT accelaration while the second one creates a large overhead while the new engine is constructed. In the next section we convert the model to handle multiple sequence lengths.
### 3.1 TRT Conversion with dynamic sequence length
```
bert_trt_path = bert_saved_model_path + '_trt2'
input_shapes = [[(1, 128), (1, 128), (1, 128)], [(1, 180), (1, 180), (1, 180)]]
trt_convert(bert_saved_model_path, bert_trt_path, input_shapes, True, np.int32, precision='FP16',
prof_strategy='Range')
trt_func_dynamic, _ = get_func_from_saved_model(bert_trt_path)
trt_res = trt_func_dynamic(**seq1)
result_key = 'bert_encoder_1' # 'classifier'
res = predict_and_benchmark_throughput(seq1, trt_func_dynamic, result_key=result_key)
res = predict_and_benchmark_throughput(seq2, trt_func_dynamic, result_key=result_key)
```
| github_jupyter |
```
import sys
sys.path.append('..') # for import src
import os
import cloudpickle
import lzma
import pandas as pd
import numpy as np
from sklearn.linear_model import Ridge
from sklearn.model_selection import cross_val_predict
from scipy.stats import pearsonr
import matplotlib.pyplot as plt
import lightgbm as lgb
import talib
import src
from src.ml_utils import (
fetch_ohlcv,
visualize_result,
normalize_position,
calc_position_cv,
get_feature_columns,
get_symbols,
unbiased_rank,
ewm_finite,
)
cloudpickle.register_pickle_by_value(src) # for model portability
# symbols = 'BTC,ETH'.split(',')
symbols = os.getenv('ALPHASEA_SYMBOLS').split(',') # 売買代金が多く、古いもの
df = fetch_ohlcv(symbols=symbols, with_target=True)
df.to_pickle('/tmp/df.pkl')
display(df)
class ExampleModelRank:
def __init__(self):
self._model = Ridge(fit_intercept=False, alpha=1e5)
self.max_data_sec = 7 * 24 * 60 * 60 # for predict script
def fit(self, df):
df = self._calc_features(df)
features = get_feature_columns(df)
df['ret_rank'] = unbiased_rank(df.groupby('timestamp')['ret']) - 0.5
df = df.dropna()
self.symbols = get_symbols(df) # for predict script
return self._model.fit(df[features], df['ret_rank'])
def predict(self, df):
df = self._calc_features(df)
features = get_feature_columns(df)
y_pred = self._model.predict(df[features])
df['position'] = np.sign(y_pred)
normalize_position(df)
return df['position']
def _calc_features(self, df):
df = df.copy()
for i in [2, 4, 8, 24, 48, 72]:
df['feature_momentum_{}'.format(i)] = (df['cl'] / df.groupby('symbol')['cl'].shift(i) - 1).fillna(0)
for i in [2, 4, 8, 24, 48, 72]:
df['feature_rsi_{}'.format(i)] = df.groupby('symbol')['cl'].transform(lambda x: talib.RSI(x, timeperiod=i).fillna(50))
for col in get_feature_columns(df):
df[col] = unbiased_rank(df.groupby('timestamp')[col]) - 0.5
return df
df = pd.read_pickle('/tmp/df.pkl')
model = ExampleModelRank()
# cv
calc_position_cv(model, df)
visualize_result(df.dropna())
# fit and save model as portable format
model.fit(df)
data = cloudpickle.dumps(model)
data = lzma.compress(data)
with open('/home/jovyan/data/example_model_rank.xz', 'wb') as f:
f.write(data)
# model validation (Just run this cell in the new kernel to make sure you saved it in a portable format.)
import os
import joblib
import pandas as pd
model = joblib.load('/home/jovyan/data/example_model_rank.xz')
df = pd.read_pickle('/tmp/df.pkl')
df = df[['op', 'hi', 'lo', 'cl']]
max_timestamp = df.index.get_level_values('timestamp').max()
df = df.loc[max_timestamp - pd.to_timedelta(model.max_data_sec, unit='S') <= df.index.get_level_values('timestamp')]
print(model.predict(df))
print(model.symbols)
```
| github_jupyter |
# Seasonal Changes in Diet and Size Selectivity
## Introduction
The relationship between numbers of different types of prey items eaten by fish and the availability of those items in the environment is not clear. In some cases a particular type of prey may show a high frequency in stomach contents and yet not be the most abundant item in the resource. _Pomatoschistus microps_ was shown to consume _Corophium_ more than any other prey even when it was not the most abundant prey species, although in laboratory experiements this fish always took the closest moving prey regardless of species and size (Magnhagen and Wiederholm, 1982). Observation of the diet of pinfish, _Lagadon rhomboides_, indicates that the predation of amphipods in seagrass is directly affected by the relative susceptibility of different species to predation than their abundance (Nelson, 1979). Other studies have also indicated similar findings that predation is affected more by other factors than abundance of prey (Ware, 1973; Moore and Moore, 1976).
The problem of availability of prey items in assessing selectivity of prey is difficult to overcome. If relative availability between prey species is assumed to be constant, this problem may be avoided by investigating the effects of abundance over time in a comparative sense.
Changes in the abundance of prey items in both stomach contents of _F. melanobranchus_ and benthic samples are examined for each sampling session to determine if any trends exist between prey abundance and number eaten.
## Methods
Bimonthly samples of fish and benthos were collected on 26th February, 23-26th April, 25-26th June, and 25-26th August 1983, during afternoon low tides.
To obtain a larger sample size of fish for February, specimens collected during January were included for analysis. Only fish of 20mm and above have been used in the analysis. In the benthic samples analysis has been restricted to amphipods retained by a 0.5mm sieve. Methods for sampling and data collection are as previously described.
## Results
The diet of gobies in terms of frequency of prey items shows a marked variation between bimonthly samples (Fig. 11). Overall there is a reduction in the total numbers of prey items eaten (Fig. 12a), and an increase in the number of prey categories. Harpacticoids and calanoids are the major cause of this drop in frequency of prey items, with a very marked change in the numbers of calanoids. Amphipod A shows a decrease through to June and remaining prey categories increase over this period. In August the trends are reversed, amphipod A increasing in frequency and other categories decreasing.
Calculation of Shannon-Wiener diversity index summarises these changes, and shows increasing diversity through to June and then falling in August (Fig. 12b). To determine if these changes reflect significant differences in the diet a t-test as proposed by Hutchenson (in Zar, 1974) was employed to test for significant differences in dietary diversity between adjacent months. The results of this test indicate that the diet of fish in April and June is not significantly different, but the months of January/February and August show significant differences to this period (Table 4).
The abundance of amphipods throughout the total sampling period drops considerable during June (Fig. 13a). This decrease is due to changes in the abundance of amphipod A, whereas the combined abundance of all other amphipods gradually increases throughout the season. Observation of the average size of amphipods (Fig. 13b) indicates little change in the size frequency distribution. This is confirmed by the observation of size frequency distributions of amphipods collected from benthic samples (Figs. 14,15). Shown alongside the size frequency distribution of amphipods in the environment is the size frequency of amphipods eaten by fish. A very marked shift to smaller amphipods in the diet of fish occurs in April and June (Fig. 14). This appears to be related to the abundance of amphipods available, although the observed total abundance of amphipods in benthic samples is not greatly different in February and April. For amphipod A alone (Fig. 15) the shift to smaller amphipods occurs in June and is more clearly related to the drop in abundance of these amphipods.
These changes are summarised in Figure 16. The number of fish eating amphipods and the number of amphipods eaten per fish show an increase in April and then a steady decrease to August (Fig. 16a,b). Amphipod A follows the above trend with the exception of June which shows a low frequency of occurrence and average number eaten. This corresponds with the low abundance of amphipods in June. The average size of amphipods eaten shows a decrease through to June and then returns to prior levels in August (Fig. 16c). These changes appear to be closely related to the changes in abundance of amphipod A (Fig. 13a), whilst all other amphipods show a steady increase throughout the sampling period. The fish appear to be switching from larger sizes of amphipod A to smaller sizes of other amphipods.
The relationship between prey abundance and numbers of each prey type eaten as shown in Figure 16 is not completely clear. When the abundance of amphipod A is compared with the proportion of amphipod A eaten a much clearer trend is apparent (Fig. 17).
## 5.4 Discussion
If the assumption of constant relative availability between different prey items is valid, the results obtained indicate that the abundance of prey items has a considerable affect on the diet of F. melanobranchus. The relationship is not a simple one involving the abundance of all prey items. Although the abundance of amphipods other than amphipod A show an increase throughout the sampling period, predation by gobies appears to be more keyed to the abundance of amphipod A. This may indicate that this amphipod is a preferred prey item.
In terms of optimal foraging theory, diversity of diet is predicted to increase as the abundance of preferred food items decreases. This prediction appears to have support from the data presented here. As the abundance of amphipod A declines so does the dietary diversity increase. The size of prey item shows a simular trend. As abundance of amphipod A decreases so does the average size of amphipod eaten. This is a result of the fish eating more number of amphipods other than amphipod A. As the other amphipods are generally smaller in size, the diet reflects this switch. Similar results have been obtained with planktivorous freshwater fish, where an active preference is shown for larger items. When the abundance of these falls, smaller items will be pursued (Bartell, 1982; Eggers, 1982).
As the size frequency of amphipods does not markedly change whilst abundance changes, the behaviour exhibited by F. melanobranchus indicates a density dependent aspect to predation for larger prey items. Evidence for a density dependent relationship is presented in Chapter 6, and will be further discussed in that section.
The habits of the various prey items may also be of relevance. Only one amphipod (AMA), has been identified as epifaunal. All other amphipods appear to be epibenthic or domicolous (see Appendix A). As amphipod A is epifaunal its availability to fish may be greater than the other amphipods (Nelson, 1979). This may be one possible causal factor in the preference of fish.
Although the results appear to be consistent with theoretical predictions with respect to amphipods, the trends observed for harpacticoid and calanoid copepods do not appear to fit. These are relatively small prey items which if the above discussion is applicable should vary in dietary abundance in a similar manner to the small amphipods. Without knowledge of the changes in benthic abundance of these items it is difficult to propose any explanation. Bartell (1982), also notes that one prey item did not follow the trends as evidenced by other prey items.
| github_jupyter |
# Packages
```
#!/usr/bin/env python
# coding: utf-8
import requests
import numpy as np
import json
import os
import time as tm
import pandas as pd
import http.client
import io
import boto3
import zipfile
from threading import Thread
import logging
from datetime import datetime
import time
from operator import itemgetter
import xlsxwriter
```
# Define API Call Variables
In the following codeblock you'll need to input data for the following variables:
***app_group_api_key*** - This is the string that allows you to access the API calls for an App Group. The App Group API Key can be found in the Braze Dashboard under Settings -> Developer Console -> Rest API Keys. An App Group API key that does not have sufficient access grants may result in an error.
***API_URL*** - This is the URL used to make the Rest API Call.The current value of 'https://rest.iad-01.braze.com' may need to be updated to match the Cluster of your Braze instance. For example, your the cluster of your Braze instance may be 02, and you would update the url to 'https://rest.iad-02.braze.com'. You can find the integer value for the API URL by checking the same value next to "dashboard-0" in the URL you use to access the Braze Dashboard.
***EXPORT_DATE*** - This field is optional, only if you have run the segment export on a prior date to the same S3 bucket. It can be left blank and will export the most recent user profile data for the selected SEGMENT_ID. If not, enter a date when the export was run previously in the following format: 'YYYY-MM-DD'. All other date formats will fail to return results
***SEGMENT_ID*** - This is the Segment API Identifier used to return user data from the segment for the API call. This script can only return results for one segment at a time, and it is recommmended that the segment have no more than 200k users due to hardware limitations that were verified during testing. The Segment API Identifier can be found in the Braze Dashboard under Settings -> Developer Console -> Additional API Identifiers. Under the dropdown menu select 'Segments' and then click the 'Search for a value' dropdown to see a list of segments. Select the segement name that you wish to return results for and copy the value listed under "API Identifier".
***The App Group API Key and Segment ID should never be shared outside of your organization, or be saved in a publically accessible workspace.***
```
app_group_api_key =
now = datetime.now().strftime("%Y-%m-%d")
API_URL = "https://rest.iad-01.braze.com"
EXPORT_DATE = []
SEGMENT_ID =
REQUEST_HEADERS = {
'Content-Type': 'application/json',
'Authorization': 'Bearer ' + app_group_api_key
}
FIELDS_TO_EXPORT = ["braze_id", "custom_attributes", "country", "total_revenue", "push_subscribe",
"email_subscribe", "custom_events", "purchases", "devices", "created_at", "apps",
"campaigns_received", "canvases_received", "cards_clicked", "push_tokens"]
```
# Define S3 Client Variables & Initializing the S3 Client
The codeblock below will initialize the client for Amazon S3 once the following values for the following variables have been added:
***access_key*** - Listed under "AWS Access ID"
***secret_key*** - Listed under "AWS Secret Access Key"
***region_name*** - The region that your S3 bucket is listed under
***user_export_bucket_name*** - The name of the S3 storage bucket that you would like to store the User Profile Export in.
All of these values, with the exception of the user_export_bucket_name can be found in the Braze Dashboard under "Integrations" -> "Technology Partners" -> "AWS Cloud Storage" -> "Data Export Using AWS Credentials".
If there are no values currently listed in this section of the Braze Dashboard, you will need to work with your System Admin to either create them for the first time, or access them. In the event that you are using MFA for AWS S3, you will need to create an account that does not require the use of MFA, as otherwise the export will fail.
***This script will not function without the proper integration between Braze and Amazon S3. While it is possible to modify the script so that the files are returned to your local machine, that functionality requires additional development.***
*You can test your credentials by entering the proper values under 'AWS Access Key', 'AWS Secret Access Key' and 'AWS S3 Bucket Name' and then press 'Test Credentials'. If you see a success message, press save. If you do not see the success message, you'll need to work with your System Admin. to create an account and S3 bucket with the correct access controls.*
**Necessary to point out**: Keep in mind costs related to a high amount of `GET` requests for the user profiles. While these costs are minimal, S3 storage is not free, so keep that in mind before making a high volume of API requests.
Once the S3 credentials have been tested and verified via the Braze Dashboard, you should be all set to store files from the `POST` request for the [User Profiles by Semgent endpoint](https://www.braze.com/docs/api/endpoints/export/user_data/post_users_segment/).
After the variables have been entered, the S3 client will be initialized, and functions will be created so that the ZIP files returned from the API request to the S3 bucket can be processed and transformed into a pandas dataframe.
```
access_key =
secret_key =
region_name =
user_export_bucket_name =
s3 = boto3.resource(
service_name='s3',
aws_access_key_id=access_key,
aws_secret_access_key=secret_key,
region_name=region_name
)
user_export_bucket = s3.Bucket(user_export_bucket_name)
```
# Segment List Endpoint
Here we'll call the [Segment List API Endpoint](https://www.braze.com/docs/api/endpoints/export/segments/get_segment/) in order to return some data needed to build the dataframe and later to return user data from that segment.
```
page, finished = 0, False
braze_segments = []
while True:
endpoint = f"{API_URL}/segments/list?page={page}&sort_direction=desc"
results = requests.get(endpoint, headers=REQUEST_HEADERS).json()['segments']
if not results:
break
braze_segments.extend(results)
page += 1
braze_segments_df = pd.DataFrame(braze_segments)
braze_segments_df.columns = ['segment_id', 'segment_name',
'segment_analytics_tracking_enabled', 'segment_tags']
braze_segments_df = braze_segments_df[braze_segments_df['segment_id'] == SEGMENT_ID]
```
# Defining Functions to Process User Profiles Stored in S3
```
def process_s3_profiles_to_dataframe(objects):
"""Build a DataFrame chunk by chunk and return it.
Temporary function for testing efficiency of building a DataFrame as we go.
There are number of great hosting solutions for us but most come with memory limit. Wanted to leave this function
for troubleshooting potential memory issues there.
Parameters
----------
objects: s3.ObjectSummary
S3 object iterator returned from `bucket.objects.filter` method
Returns
-------
pd.DataFrame
New dataframe with exported user profile from the selected objects
"""
frame_chunks = []
for obj in objects:
segment_id = obj.key.split('/')[1]
with io.BytesIO(obj.get()["Body"].read()) as zip_bytes:
user_chunk = process_s3_zip_object(zip_bytes, segment_id)
frame_chunk = pd.DataFrame(user_chunk)
frame_chunks.append(frame_chunk)
return pd.concat(frame_chunks, ignore_index=True)
def process_s3_profiles(objects, user_data):
"""Extract and process zip user profiles obtained from user segment export.
Parameters
----------
objects : s3.ObjectSummary
S3 object iterator returned from `bucket.objects.filter` method
user_data : list
Store for the extracted profile objects
"""
for obj in objects:
segment_id = obj.key.split('/')[1]
with io.BytesIO(obj.get()["Body"].read()) as zip_bytes:
user_chunk = process_s3_zip_object(zip_bytes, segment_id)
user_data.extend(user_chunk)
def process_s3_zip_object(zip_bytes, segment_id):
"""Extract the zip file contents and process each text file within that zip file.
Text files extracted contain user data JSONs, separated by new line.
Parameters
----------
zip_bytes : io.BytesIO
segment_id : string
Returns
-------
list
Extracted user profile dictionaries from the zip file
"""
profiles = []
with zipfile.ZipFile(zip_bytes) as open_zip:
for user_file in open_zip.namelist():
with open_zip.open(user_file) as users:
for line in users:
user_profile = json.loads(line.decode('utf-8'))
user_profile['segment_id'] = segment_id
profiles.append(user_profile)
return profiles
```
# Define Functions for Processing Campaign Data
The below codeblock:defines functions to enable `GET` requests from the [Campaign Details Endpoint](https://www.braze.com/docs/api/endpoints/export/campaigns/get_campaign_details/) for one or many campaign_ids. It also creates functions to enable the creation of the Channel Combo and Custom Events used in Campaigns.
The URL may need to be updated in the same manner as above following 'iad-0' depending on the cluster of your Braze Instance. For example, you may need to update the string "https://rest.iad-01.braze.com/campaigns/" to "https://rest.iad-02.braze.com/campaigns/" if the Cluster for your Braze instance is 02.
The MAX_RETRIES variable is the number of times that the script will attempt to make a request to the the API Endpoint. If the number is increased the script will take longer to return results from the Campaign Details Endpoint.
```
MAX_RETRIES=3
def process_campaign_id(campaign_id, endpoint):
requests_made = 0
while requests_made < MAX_RETRIES:
try:
response = requests.get("https://rest.iad-01.braze.com/campaigns/"+endpoint+"?campaign_id="+campaign_id, headers=REQUEST_HEADERS
)
return response.json()
except requests.exceptions.HTTPError:
requests_made += 1
tm.sleep(0.5)
if requests_made >= MAX_RETRIES:
raise
### processes a range of ids
def process_campaign_id_range(campaign_id_range, endpoint, store=None):
"""process a number of ids, storing the results in a dict"""
if store is None:
store = {}
for campaign_id in campaign_id_range:
store[campaign_id] = process_campaign_id(campaign_id, endpoint)
return store
def threaded_process_campaigns(nthreads, campaign_id_range, endpoint):
"""process the id range in a specified number of threads"""
try:
store = {}
threads = []
for i in range(nthreads):
campaign_ids = campaign_id_range[i::nthreads]
t = Thread(target=process_campaign_id_range,
args=(campaign_ids, endpoint, store))
threads.append(t)
for t in threads:
t.start()
for t in threads:
t.join()
return store
except Exception as e:
logging.error("Threading exception: "+str(e))
tm.sleep(30)
def get_campaign_id(df_column):
try:
return df_column.get('api_campaign_id')
except AttributeError:
return float('NaN')
def get_message_variation_id(df_column):
try:
return df_column.get('variation_api_id')
except AttributeError:
return float('NaN')
def parse_channel(row):
if row.num_channels > 0:
return row.channel
elif type(row.campaigns_received) != dict:
return "No Messages Received"
else:
return "Unable to Retrieve Campaign Details"
def parse_channel_combo(row):
if type(row.channel_combo) != float:
return row.channel_combo
elif row.channel == "No Messages Received":
return "No Messages Received"
else:
return "Unable to Retrieve Campaign Details"
def get_campaign_custom_event(df_column):
try:
return df_column.get('custom_event_name')
except AttributeError:
return float('NaN')
```
# Define Field Getters to Enable Segment Analytics
The functions defined in the codeblocks below will get the corresponding fields from nested dictionaries stored in dataframes columns that are returned from the User Profiles Endpoint.
```
def get_email_open_engagement(df_column):
try:
return df_column.get('opened_email')
except AttributeError:
return False
def get_email_click_engagement(df_column):
try:
return df_column.get('clicked_email')
except AttributeError:
return False
def get_push_engagement(df_column):
try:
return df_column.get('opened_push')
except AttributeError:
return False
def get_iam_engagement(df_column):
try:
return df_column.get('clicked_in_app_message')
except AttributeError:
return False
def get_conversions(df_column):
try:
return df_column.get('converted')
except AttributeError:
return False
### Create get engagement
def calc_engagement(series):
return series.sum()/series.count()
def get_cards_clicked(row):
if row.channel == 'No Messages Received':
return 0
else:
return len(row.cards_clicked)
def days_between(d1, d2):
d1 = datetime.datetime.strptime(str(d1), '%Y-%m-%dT%H:%M:%S.%f%z')
d2 = datetime.datetime.strptime(str(d2), '%Y-%m-%dT%H:%M:%S.%f%z')
return (d2 - d1).days
def get_custom_event_name(df_column):
try:
return df_column.get('name')
except AttributeError:
return float('NaN')
def get_custom_event_count(df_column):
try:
return df_column.get('count')
except AttributeError:
return float('NaN')
def get_custom_event_first_date(df_column):
try:
return df_column.get('first')
except AttributeError:
return float('NaN')
def get_custom_event_last_date(df_column):
try:
return df_column.get('last')
except AttributeError:
return float('NaN')
def get_notifications_enabled(df_column):
try:
return df_column.get('notifications_enabled')
except AttributeError:
return False
def get_token(df_column):
try:
return df_column.get('token')
except AttributeError:
return 'None'
def get_platform(df_column):
try:
return df_column.get('platform')
except AttributeError:
return 'No platform token'
```
# Export Data from S3 for either Today or a Prior Date
The below codeblock will do the following depending on the value entered above for the EXPORT_DATE variable:
***If the EXPORT_DATE is left blank***:
- Make a request to the [Users by Segment Endpoint](https://www.braze.com/docs/api/endpoints/export/user_data/post_users_segment/)
- Process the user profile data returned to S3 following the successful request for the selected SEGMENT_ID
- Displays the number of user profiles that have been returned.
If the value returned is 0 it is likely that some of the above variables were not configured properly. You'll need to double check and try again.
If the number of user profiles exported is low, it could be because of latency between the Braze API and S3. Try running the code block again and see if the number of users returned increases
***If the EXPORT_DATE is a properly formatted data from a prior export***
- Process the user profile data returned to S3 following the successful request for the selected SEGMENT_ID
- Displays the number of user profiles that have been returned.
If the EXPORT_DATE is not formatted 'YYYY-MM-DD' the below codeblock will fail and you will be asked to try again.
If completed successfully, the segment_df data should return a dataframe for all user profiles from the segment, along with data from the fields listed out in the *FIELDS_TO_EXPORT* variable. Each row in the dataframe corresponds to one user profile within the selected segment.
```
if len(EXPORT_DATE) == 0:
object_prefix_by_segment_id = []
payload = {
"segment_id": SEGMENT_ID,
"fields_to_export": FIELDS_TO_EXPORT
}
res = requests.post(f"{API_URL}/users/export/segment",
headers=REQUEST_HEADERS, json=payload)
res_data = res.json()
print(res_data)
EXPORT_DATE = datetime.today().strftime('%Y-%m-%d')
objects = user_export_bucket.objects.filter(
Prefix=f"segment-export/{SEGMENT_ID}/{EXPORT_DATE}")
tm.sleep(300)
print("Waiting for data to be returned from the Users by Segment Endpoint.")
start = time.time()
user_data = []
print("Reading exported user data from S3")
process_s3_profiles(objects, user_data)
print(f"Took {(time.time() - start):.2f}s")
print(len(user_data))
elif len(EXPORT_DATE) == 10 and EXPORT_DATE.count('-') == 2 and len(EXPORT_DATE) > 0:
year, month, day = EXPORT_DATE.split('-')
isValidDate = True
try:
datetime(int(year), int(month), int(day))
except ValueError:
print("Input date is not the valid YYYY-MM-DD format. Please return to the Define Variables cell and try again enter a properly formatted Date.")
isValidDate = False
if(isValidDate):
objects = user_export_bucket.objects.filter(
Prefix=f"segment-export/{SEGMENT_ID}/{EXPORT_DATE}")
start = time.time()
user_data = []
print("Reading exported user data from S3")
process_s3_profiles(objects, user_data)
print(f"Took {(time.time() - start):.2f}s")
print(len(user_data))
else:
print("This is the text that will display if export date is neither blank nor properly formatted.")
segment_df_raw = pd.DataFrame(user_data)\
.dropna(subset=['braze_id'])
segment_df = pd.merge(segment_df_raw, braze_segments_df,
how='left',
left_on=['segment_id'],
right_on=['segment_id'],
suffixes=['_from_user_segment_endpoint', '_from_segment_list'])
```
# Creating Separate Dataframes for Each KPI
The below codeblock will split the segment_df into the appropriate dataframes so that the following analytical outputs can be viewed from the selected Segment:
1. Rolling Retention
2. Purchasing Rates &
3. Purchase Retention
4. Session Engagement Metrics
5. Custom Event Metrics
5. Message Engagement Rates
6. Custom Events used in Campaigns
7. Opt-In Rates for Push and Email
1-5 will also be crossed by the following dimensions from Message Engagement so that the impact of different messaging strategies can be viewed at the segement level:
- Channel
- Channel Combo
- Campaign Tag
In the event that a segment fails one of the checks below, you can skip those sections in the script. For example, say you are tracking session data, but not purchasing data. Skip the purchasing codeblocks and comment out the final outputs associated with those metrics.
```
rolling_retention_columns = ['braze_id', 'segment_id',
'apps', 'segment_name', 'segment_tags']
purchasing_stats_columns = ['braze_id',
'segment_id', 'apps', 'segment_name', 'segment_tags', 'purchases', 'total_revenue']
sessions_stats_columns = ['braze_id', 'segment_id',
'apps', 'segment_name', 'segment_tags']
custom_events_stats_columns = ['braze_id', 'segment_id', 'apps',
'segment_name', 'segment_tags', 'custom_events']
engagement_stats_columns_all = ['braze_id', 'segment_id', 'country', 'apps', 'segment_name',
'segment_tags', 'campaigns_received', 'canvases_received', 'cards_clicked']
engagement_stats_columns_canvas = ['braze_id', 'segment_id', 'country', 'apps',
'segment_name', 'segment_tags', 'canvases_received', 'cards_clicked']
engagement_stats_columns_campaigns = ['braze_id', 'segment_id', 'country', 'apps',
'segment_name', 'segment_tags', 'campaigns_received', 'cards_clicked']
opt_ins_columns_all = ['braze_id', 'segment_id', 'segment_name', 'apps', 'push_tokens', 'email_subscribe', 'email_opted_in_at', 'push_subscribe',
'push_opted_in_at', 'email_unsubscribed_at', 'push_unsubscribed_at']
opt_ins_columns_email = ['braze_id', 'segment_id', 'segment_name', 'apps', 'push_tokens',
'email_subscribe', 'email_opted_in_at', 'email_unsubscribed_at']
opt_ins_columns_push = ['braze_id', 'segment_id', 'segment_name', 'apps', 'push_tokens',
'push_subscribe', 'push_opted_in_at', 'push_unsubscribed_at']
users_have_sessions = "apps" in segment_df
users_have_purchases = "purchases" in segment_df
users_have_custom_events = "custom_events" in segment_df
users_received_campaigns = "campaigns_received" in segment_df
users_received_canvas = "canvases_received" in segment_df
users_subscribed_email = "email_subscribe" in segment_df
users_subscribed_push = "push_subscribe" in segment_df
if users_have_sessions == True:
segment_rolling_retention_pre_apps = segment_df[rolling_retention_columns]
segment_rolling_retention_pre_apps = segment_rolling_retention_pre_apps.reset_index()
else:
print("Users in these Segments do not have Retention Data")
if users_have_purchases == True:
segment_purchasing_stats_pre_apps = segment_df[purchasing_stats_columns]
segment_purchasing_stats_pre_apps = segment_purchasing_stats_pre_apps.reset_index()
else:
print("Users in these Segments do not have Purchasing Data")
if users_have_sessions == True:
segment_sessions_stats_pre_apps = segment_df[sessions_stats_columns]
segment_sessions_stats_pre_apps = segment_sessions_stats_pre_apps.reset_index()
else:
print("Users in these Segments do not have Session Data")
if users_have_custom_events == True:
segment_custom_event_stats_pre_custom_event = segment_df[custom_events_stats_columns]
segment_custom_event_stats_pre_custom_event = segment_custom_event_stats_pre_custom_event.reset_index()
else:
print("Users in these Segments do not have Custom Event Data")
if (users_received_campaigns == True and users_received_canvas == True):
segment_engagement_stats_pre_apps = segment_df[engagement_stats_columns_all]
elif (users_received_campaigns == False and users_received_canvas == True):
segment_engagement_stats_pre_apps = segment_df[engagement_stats_columns_canvas]
elif (users_received_campaigns == True and users_received_canvas == False):
segment_engagement_stats_pre_apps = segment_df[engagement_stats_columns_campaigns]
elif (users_received_campaigns == False and users_received_canvas == False):
print("Users in these Segments do not have Engagement Data")
if (users_subscribed_email == True and users_subscribed_push == True):
segment_opt_in_stats_pre_apps = segment_df[opt_ins_columns_all]
elif (users_subscribed_email == False and users_subscribed_push == True):
segment_opt_in_stats_pre_apps = segment_df[users_subscribed_push]
elif (users_subscribed_email == True and users_subscribed_push == False):
segment_opt_in_stats_pre_apps = segment_df[users_subscribed_email]
elif (users_subscribed_email == False and users_subscribed_push == False):
print("Users in these Segments do not have Opt-In Data")
```
# Campaign & Engagement Data
The below codeblocks will complete the following tasks:
- Return all of the campaign ids received by the exported Segment
- Send `GET` results from the [Campaign Details API](https://www.braze.com/docs/api/endpoints/export/campaigns/get_campaign_details/#campaign-details-endpoint-api-response) and process the data that is returned.
- Users that received messages from campaign_ids that do not have details returned will be assigned the 'Unable to Retrieve Campaign Details' value for both Channel and Channel Combo.
- Create the Channel_Combo dimension. Please note that the Channel Combo is being created at the Campaign Level and not the User Level.
- Removing Users in the control_group for multivariate campaigns
- Cleaning the Channel names and Channel Combo names
- Creating the dataframe used to caclulate Message Engagement Metrics
- Creating dataframes used to cross other metrics with Channel, Channel Combo, and Campaign Tag
```
segment_engagement_temp = segment_engagement_stats_pre_apps.explode(
'campaigns_received')
segment_engagement_temp['campaign_id'] = list(
map(get_campaign_id, segment_engagement_temp['campaigns_received']))
braze_campaigns = segment_engagement_temp[segment_engagement_temp['campaign_id'].isnull(
) == False]['campaign_id']
braze_campaigns = list(set(braze_campaigns))
campaign_dict = threaded_process_campaigns(
10, braze_campaigns, 'details')
campaign_details_df = pd.DataFrame.from_dict(campaign_dict, orient='index')
campaign_details_df = campaign_details_df.reset_index()
campaign_details_df.rename(columns={"index": "campaign_id"},
inplace=True)
campaign_details_df = campaign_details_df[campaign_details_df['message'] == 'success']
campaign_details_df['num_channels'] = campaign_details_df.channels.apply(len)
campaign_details_df = campaign_details_df[campaign_details_df['num_channels'] > 0]
joined_campaign = pd.merge(segment_engagement_temp, campaign_details_df,
how='left',
left_on=['campaign_id'],
right_on=['campaign_id'],
suffixes=['_from_segments', '_from_campaigns'])
segment_data_engagement_stats_temp = joined_campaign
segment_data_engagement_stats_temp.rename(columns={"channels": "channel"},
inplace=True)
segment_data_engagement_stats_temp['in_control']=segment_data_engagement_stats_temp.campaigns_received.apply(
lambda x: x.get('in_control') if type(x) != float else x)
segment_data_engagement_stats_temp=segment_data_engagement_stats_temp[segment_data_engagement_stats_temp['in_control']!=True]
segment_data_engagement_stats_temp.loc[:, 'channel'] = segment_data_engagement_stats_temp.apply(
parse_channel, axis=1)
segment_data_engagement_stats_temp = segment_data_engagement_stats_temp.explode(
'channel')
segment_data_engagement_stats_temp['channel'] = segment_data_engagement_stats_temp.channel.apply(
lambda x: 'mobile_push' if x == 'android_push' or x == 'ios_push' else x)
segment_data_engagement_stats_temp['channel'] = segment_data_engagement_stats_temp.channel.apply(
lambda x: 'in_app_message' if x == 'legacy_in_app_message' or x == 'trigger_in_app_message ' or x == 'trigger_in_app_message' else x)
segment_data_engagement_stats_temp['channel'] = segment_data_engagement_stats_temp.channel.apply(
lambda x: x.replace("_", " "))
segment_data_engagement_stats_temp['channel'] = segment_data_engagement_stats_temp.channel.apply(
lambda x: x.title())
segment_data_channel_combo = segment_data_engagement_stats_temp[(segment_data_engagement_stats_temp['channel'] != 'No Messages Received')]
segment_data_channel_combo = segment_data_engagement_stats_temp[segment_data_engagement_stats_temp['channel'] != 'Unable To Retrieve Campaign Details']
segment_data_channel_combo = segment_data_channel_combo[[
'braze_id', 'channel']].drop_duplicates()
segment_data_channel_combo = segment_data_channel_combo.dropna(subset=[
'channel'])
segment_data_channel_combo = segment_data_channel_combo.groupby('braze_id')
segment_data_channel_combo = segment_data_channel_combo.apply(
lambda x: x['channel'].unique()).reset_index()
segment_data_channel_combo.columns = ['braze_id', 'channel_combo']
segment_data_channel_combo['channel_combo'] = segment_data_channel_combo.channel_combo.apply(
lambda x: np.ndarray.tolist(x))
segment_data_channel_combo['channel_combo'] = segment_data_channel_combo['channel_combo'].apply(
lambda x: list(set(x)))
segment_data_channel_combo['channel_combo'] = segment_data_channel_combo.channel_combo.apply(
sorted)
segment_data_channel_combo['channel_combo'] = [
', '.join(map(str, l)) for l in segment_data_channel_combo['channel_combo']]
segment_data_channel_combo = segment_data_channel_combo.drop_duplicates()
segment_data_engagement_stats = pd.merge(segment_data_engagement_stats_temp, segment_data_channel_combo,
how='left',
left_on=['braze_id'],
right_on=['braze_id'],
suffixes=['_from_engagement', '_from_channel_combo'])
segment_data_engagement_stats.loc[:, 'channel_combo'] = segment_data_engagement_stats.apply(
parse_channel_combo, axis=1)
users_per_channel_df = segment_data_engagement_stats.groupby(
['segment_name', 'segment_id','channel']).agg(num_users=('braze_id', 'nunique'))
users_per_channel_df = users_per_channel_df.reset_index(level=[0, 1, 2])
users_per_channel_combo_df = segment_data_engagement_stats.groupby(
['segment_name', 'segment_id','channel_combo']).agg(num_users=('braze_id', 'nunique'))
users_per_channel_combo_df = users_per_channel_combo_df.reset_index(level=[
0, 1, 2])
users_per_campaign_tags_df = segment_data_engagement_stats.explode('tags')
users_per_campaign_tags_df['tags'] = users_per_campaign_tags_df.tags.fillna(
'No Messages')
users_per_campaign_tags_df = users_per_campaign_tags_df.groupby(
['segment_name', 'segment_id','tags']).agg(num_users=('braze_id', 'nunique'))
users_per_campaign_tags_df = users_per_campaign_tags_df.reset_index(level=[
0, 1, 2])
```
# Calculate Engagement
The below codeblocks will return Messge Engagement rates for all channels. If the segment did not receive a channel it will simply return a value of zero under the engagement metric.
The following Message Engagement Rates will be returned:
- Number of Users
- Email Open Rate
- Email Click Rate
- Push Open Rate
- In-App Message Click Rate
- Message Conversion Rates (of all Conversion Criteria)
- Content Card Click Rate
Message Engagement Rates will be returned by:
- Segment
- Channel
- Channel Combo
- Campaign Tag
```
segment_data_engagement_stats['campaign_engaged'] = segment_data_engagement_stats.campaigns_received.apply(
lambda x: x.get('engaged') if type(x) != float else x)
segment_data_engagement_stats['opened_email'] = list(
map(get_email_open_engagement, segment_data_engagement_stats['campaign_engaged']))
segment_data_engagement_stats['clicked_email'] = list(map(
get_email_click_engagement, segment_data_engagement_stats['campaign_engaged']))
segment_data_engagement_stats['opened_push'] = list(
map(get_push_engagement, segment_data_engagement_stats['campaign_engaged']))
segment_data_engagement_stats['clicked_iam'] = list(
map(get_iam_engagement, segment_data_engagement_stats['campaign_engaged']))
segment_data_engagement_stats['converted'] = list(
map(get_conversions, segment_data_engagement_stats['campaigns_received']))
segment_data_engagement_stats['converted'] = segment_data_engagement_stats.converted.fillna(
value=False)
segment_data_engagement_stats['cards_clicked'] = segment_data_engagement_stats.cards_clicked.fillna(
value='')
segment_data_engagement_stats.loc[:, 'cards_clicked'] = segment_data_engagement_stats.apply(
get_cards_clicked, axis=1)
engagement_by_segment_preagg = segment_data_engagement_stats.groupby(
['segment_name', 'segment_id'])
engagement_by_segment = engagement_by_segment_preagg.agg(
num_users=('braze_id', 'nunique'), email_open_rate=('opened_email', calc_engagement), email_click_rate=('clicked_email', calc_engagement),
push_open_rate=('opened_push', calc_engagement), iam_click_rate=('clicked_iam', calc_engagement), conversion_rate=('converted', calc_engagement), content_card_click_rate=('cards_clicked', calc_engagement))
engagement_by_segment_and_channel_preagg = segment_data_engagement_stats.groupby(
['segment_name', 'segment_id', 'channel'])
engagement_by_segment_and_channel = engagement_by_segment_and_channel_preagg.agg(
num_users=('braze_id', 'nunique'), email_open_rate=('opened_email', calc_engagement), email_click_rate=('clicked_email', calc_engagement),
push_open_rate=('opened_push', calc_engagement), iam_click_rate=('clicked_iam', calc_engagement), conversion_rate=('converted', calc_engagement), content_card_click_rate=('cards_clicked', calc_engagement))
engagement_by_segment_and_channel_combo_preagg = segment_data_engagement_stats.groupby(
['segment_name', 'segment_id', 'channel_combo'])
engagement_by_segment_and_channel_combo = engagement_by_segment_and_channel_combo_preagg.agg(
num_users=('braze_id', 'nunique'), email_open_rate=('opened_email', calc_engagement), email_click_rate=('clicked_email', calc_engagement),
push_open_rate=('opened_push', calc_engagement), iam_click_rate=('clicked_iam', calc_engagement), conversion_rate=('converted', calc_engagement), content_card_click_rate=('cards_clicked', calc_engagement))
engagement_by_segment_and_channel_combo = engagement_by_segment_and_channel_combo
segment_data_engagement_stats_by_campaign_tags = segment_data_engagement_stats.explode(
'tags')
engagement_by_segment_and_campaign_tag_preagg = segment_data_engagement_stats_by_campaign_tags.groupby([
'segment_name', 'segment_id', 'tags'])
engagement_by_segment_and_campaign_tag = engagement_by_segment_and_campaign_tag_preagg.agg(
num_users=('braze_id', 'nunique'), email_open_rate=('opened_email', calc_engagement), email_click_rate=('clicked_email', calc_engagement),
push_open_rate=('opened_push', calc_engagement), iam_click_rate=('clicked_iam', calc_engagement), conversion_rate=('converted', calc_engagement), content_card_click_rate=('cards_clicked', calc_engagement))
```
# Rolling Retention
The below codeblocks will return Rolling Retetion Rates. You can view the Rolling Retention Methodology [here](https://www.braze.com/resources/articles/calculate-retention-rate).
Rolling Retention Rates will be returned by:
- Segment
- Channel
- Channel Combo
- Campaign Tag
```
segment_rolling_retention_temp = segment_rolling_retention_pre_apps.explode(
'apps')
segment_rolling_retention_temp = segment_rolling_retention_temp.dropna(subset=[
'apps'])
segment_rolling_retention_temp['first_used'] = segment_rolling_retention_temp['apps'].apply(
lambda x: x.get('first_used'))
segment_rolling_retention_temp['last_used'] = segment_rolling_retention_temp['apps'].apply(
lambda x: x.get('last_used'))
segment_rolling_retention_temp['platform'] = segment_rolling_retention_temp['apps'].apply(
lambda x: x.get('platform'))
segment_rolling_retention_temp[['first_used', 'last_used']] = segment_rolling_retention_temp[[
'first_used', 'last_used']].apply(pd.to_datetime)
segment_rolling_retention_temp['day_num'] = (
segment_rolling_retention_temp['last_used'] - segment_rolling_retention_temp['first_used']).dt.days
segment_rolling_retention_temp['day_num'] = segment_rolling_retention_temp['day_num'].astype(
'int')
segment_rolling_retention_raw = pd.pivot_table(segment_rolling_retention_temp,
values=("braze_id"),
index=("segment_name", 'segment_id',
"platform"),
columns="day_num",
aggfunc='nunique')\
.fillna(0)
segment_rolling_retention_raw = segment_rolling_retention_raw[segment_rolling_retention_raw
.columns[::-1]].cumsum(axis=1)
segment_rolling_retention_raw = segment_rolling_retention_raw[
segment_rolling_retention_raw.columns[::-1]]
segment_rolling_retention_raw["num_users"] = segment_rolling_retention_raw[0]
segment_rolling_retention_raw = segment_rolling_retention_raw.groupby(
['segment_name', 'segment_id', 'platform']).sum()
segment_rolling_retention = pd.concat([segment_rolling_retention_raw["num_users"],
segment_rolling_retention_raw
.drop(["num_users"], axis=1)
.div(segment_rolling_retention_raw["num_users"], axis=0)],
axis=1).fillna(0)
segment_rolling_retention=segment_rolling_retention.drop(0,axis=1)
segment_engagement_user_data = segment_data_engagement_stats[[
'braze_id', 'segment_id', 'segment_name', 'apps', 'channel', 'tags', 'channel_combo']]
segment_engagement_data_for_retention = segment_engagement_user_data.explode(
'apps')
segment_engagement_data_for_retention = segment_engagement_data_for_retention.dropna(subset=[
'apps'])
segment_engagement_data_for_retention['platform'] = segment_engagement_data_for_retention['apps'].apply(
lambda x: x.get('platform'))
segment_rolling_retention_by_engagement_temp = pd.merge(segment_rolling_retention_temp.reset_index(), segment_engagement_data_for_retention.reset_index(),
how='left',
left_on=[
'braze_id', 'platform', 'segment_id', 'segment_name'],
right_on=[
'braze_id', 'platform', 'segment_id', 'segment_name'],
suffixes=['_from_retention', '_from_engagement'])
segment_rolling_retention_by_engagement_temp['day_num'] = segment_rolling_retention_by_engagement_temp['day_num'].astype(
'int')
segment_rolling_retention_by_engagement_raw = pd.pivot_table(segment_rolling_retention_by_engagement_temp,
values=(
"braze_id"),
index=(
"segment_name", "segment_id", "platform", "channel"),
columns="day_num",
aggfunc='nunique')\
.fillna(0)
segment_rolling_retention_by_engagement_raw = segment_rolling_retention_by_engagement_raw[segment_rolling_retention_by_engagement_raw
.columns[::-1]].cumsum(axis=1)
segment_rolling_retention_by_engagement_raw = segment_rolling_retention_by_engagement_raw[
segment_rolling_retention_by_engagement_raw.columns[::-1]]
segment_rolling_retention_by_engagement_raw["num_users"] = segment_rolling_retention_by_engagement_raw[0]
segment_rolling_retention_by_engagement_raw = segment_rolling_retention_by_engagement_raw.groupby(
['segment_name', 'segment_id', 'platform', "channel"]).sum()
segment_rolling_retention_by_engagement = pd.concat([segment_rolling_retention_by_engagement_raw["num_users"],
segment_rolling_retention_by_engagement_raw
.drop(["num_users"], axis=1)
.div(segment_rolling_retention_by_engagement_raw["num_users"], axis=0)],
axis=1).fillna(0)
segment_rolling_retention_by_engagement=segment_rolling_retention_by_engagement.drop(0,axis=1)
segment_rolling_retention_by_engagement_temp['day_num'] = segment_rolling_retention_by_engagement_temp['day_num'].astype(
'int')
segment_campaign_tag_data_for_retention_temp = segment_rolling_retention_by_engagement_temp.explode(
'tags')
segment_campaign_tag_data_for_retention_temp = segment_campaign_tag_data_for_retention_temp.dropna(subset=[
'tags'])
segment_rolling_retention_by_campaign_tag_raw = pd.pivot_table(segment_campaign_tag_data_for_retention_temp,
values=(
"braze_id"),
index=(
"segment_name", "segment_id", "platform", "tags"),
columns="day_num",
aggfunc='nunique')\
.fillna(0)
segment_rolling_retention_by_campaign_tag_raw = segment_rolling_retention_by_campaign_tag_raw[segment_rolling_retention_by_campaign_tag_raw
.columns[::-1]].cumsum(axis=1)
segment_rolling_retention_by_campaign_tag_raw = segment_rolling_retention_by_campaign_tag_raw[
segment_rolling_retention_by_campaign_tag_raw.columns[::-1]]
segment_rolling_retention_by_campaign_tag_raw["num_users"] = segment_rolling_retention_by_campaign_tag_raw[0]
segment_rolling_retention_by_campaign_tag_raw = segment_rolling_retention_by_campaign_tag_raw.groupby(
['segment_name', 'segment_id', 'platform', "tags"]).sum()
segment_rolling_retention_by_campaign_tag = pd.concat([segment_rolling_retention_by_campaign_tag_raw["num_users"],
segment_rolling_retention_by_campaign_tag_raw
.drop(["num_users"], axis=1)
.div(segment_rolling_retention_by_campaign_tag_raw["num_users"], axis=0)],
axis=1).fillna(0)
segment_rolling_retention_by_campaign_tag =segment_rolling_retention_by_campaign_tag.drop(0,axis=1)
segment_rolling_retention_by_engagement_temp['day_num'] = segment_rolling_retention_by_engagement_temp['day_num'].astype(
'int')
segment_rolling_retention_by_channel_combo_raw = pd.pivot_table(segment_rolling_retention_by_engagement_temp,
values=(
"braze_id"),
index=(
"segment_name", "segment_id", "platform", "channel_combo"),
columns="day_num",
aggfunc='nunique')\
.fillna(0)
segment_rolling_retention_by_channel_combo_raw = segment_rolling_retention_by_channel_combo_raw[segment_rolling_retention_by_channel_combo_raw
.columns[::-1]].cumsum(axis=1)
segment_rolling_retention_by_channel_combo_raw = segment_rolling_retention_by_channel_combo_raw[
segment_rolling_retention_by_channel_combo_raw.columns[::-1]]
segment_rolling_retention_by_channel_combo_raw["num_users"] = segment_rolling_retention_by_channel_combo_raw[0]
segment_rolling_retention_by_channel_combo_raw = segment_rolling_retention_by_channel_combo_raw.groupby(
['segment_name', 'segment_id', 'platform', "channel_combo"]).sum()
segment_rolling_retention_by_channel_combo = pd.concat([segment_rolling_retention_by_channel_combo_raw["num_users"],
segment_rolling_retention_by_channel_combo_raw
.drop(["num_users"], axis=1)
.div(segment_rolling_retention_by_channel_combo_raw["num_users"], axis=0)],
axis=1).fillna(0)
segment_rolling_retention_by_channel_combo=segment_rolling_retention_by_channel_combo.drop(0,axis=1)
```
# Purchasing Stats
The following purchasing metrics will be returned in the first purchasing stats dataframe:
- Number of Buyers
- Number of Repeat Buyers
- % Buyers
- % Repeat Buyers
- Number of Purchases
- Total Revenue
- Average Revenue per Buyer
- Average time to Purchase
- Purchases per Buyer
The second purchasing stats dataframe will return purchase retention rates.
Both purchasing stats dataframes will returned by:
- Segment
- Channel
- Channel Combo
- Campaign Tag
```
num_users = segment_df.braze_id.nunique()
segment_purchasing_stats_temp = segment_purchasing_stats_pre_apps.dropna(
subset=['apps', 'purchases'])
segment_purchasing_dates = segment_purchasing_stats_pre_apps.dropna(
subset=['apps', 'purchases'])
segment_purchasing_dates = segment_purchasing_dates.explode(
'purchases')
segment_purchasing_dates = segment_purchasing_dates.explode(
'apps')
segment_purchasing_stats_temp['num_purchases'] = segment_purchasing_stats_temp['purchases'].apply(
lambda x: sum(map(itemgetter('count'), x)))
segment_purchasing_dates['first_purchase'] = segment_purchasing_dates['purchases'].apply(
lambda x: x.get('first'))
segment_purchasing_dates['last_purchase'] = segment_purchasing_dates['purchases'].apply(
lambda x: x.get('last'))
segment_purchasing_dates['first_session'] = segment_purchasing_dates['apps'].apply(
lambda x: x.get('first_used'))
segment_purchasing_dates['first_purchase'] = pd.to_datetime(
segment_purchasing_dates['first_purchase'])
segment_purchasing_dates['last_purchase'] = pd.to_datetime(
segment_purchasing_dates['last_purchase'])
segment_purchasing_dates['first_session'] = pd.to_datetime(
segment_purchasing_dates['first_session'])
segment_purchasing_dates_temp = segment_purchasing_dates.groupby(['segment_name', 'segment_id', 'braze_id']).agg(first_purchase_date=(
'first_purchase', 'min'), last_purchase_date=('last_purchase', 'max'), first_session_date=('first_session', 'min'))
segment_purchasing_dates_temp = segment_purchasing_dates_temp.reset_index(level=[
0, 1, 2])
segment_purchasing_stats_temp = pd.merge(segment_purchasing_stats_temp, segment_purchasing_dates_temp,
how='left',
left_on=[
'braze_id', 'segment_id', 'segment_name'],
right_on=[
'braze_id', 'segment_id', 'segment_name'])
segment_purchasing_stats_temp['repeat_buyer'] = segment_purchasing_stats_temp[
'first_purchase_date'] != segment_purchasing_stats_temp['last_purchase_date']
segment_purchasing_stats_temp['repeat_buyer_id'] = segment_purchasing_stats_temp.apply(
lambda row: row.braze_id if row.repeat_buyer == True else 'NaN', axis=1)
segment_purchasing_stats_temp['days_to_purchase'] = (
segment_purchasing_stats_temp['first_purchase_date'] - segment_purchasing_stats_temp['first_session_date']).dt.seconds
segment_purchasing_stats_temp['days_to_purchase'] = segment_purchasing_stats_temp['days_to_purchase']/86400
segment_purchase_retention_temp = segment_purchasing_stats_temp
segment_purchase_data = segment_purchasing_stats_temp
segment_purchasing_stats_temp = segment_purchasing_stats_temp.groupby(['segment_name', 'segment_id']).agg(buyers=('braze_id', 'nunique'), repeat_buyers=('repeat_buyer_id', 'nunique'), num_purchases=(
'num_purchases', 'sum'), total_revenue=('total_revenue', 'sum'), avg_revenue_per_buyer=('total_revenue', 'mean'), avg_time_to_purchase=('days_to_purchase', 'mean'))
segment_purchasing_stats_temp['pct_repeat_buyers'] = round(
segment_purchasing_stats_temp.repeat_buyers/segment_purchasing_stats_temp.buyers, 2)
segment_purchasing_stats_temp['purchases_per_buyer'] = round(
segment_purchasing_stats_temp.num_purchases/segment_purchasing_stats_temp.buyers, 2)
segment_purchasing_stats_temp['revenue_per_item_purchased'] = round(
segment_purchasing_stats_temp.total_revenue/segment_purchasing_stats_temp.num_purchases, 2)
segment_purchasing_stats_temp['purchases_per_user'] = round(
segment_purchasing_stats_temp.num_purchases/num_users, 2)
segment_purchasing_stats_temp['pct_buyer'] = round(
segment_purchasing_stats_temp.buyers/num_users, 2)
segment_purchasing_stats = segment_purchasing_stats_temp
segment_purchase_retention_temp['day_num'] = (
segment_purchase_retention_temp['last_purchase_date'] - segment_purchase_retention_temp['first_purchase_date']).dt.days
segment_purchase_retention_temp['day_num'] = segment_purchase_retention_temp['day_num'].astype(
'int')
segment_purchase_retention_raw = pd.pivot_table(segment_purchase_retention_temp,
values=("braze_id"),
index=("segment_name",
"segment_id"),
columns="day_num",
aggfunc='nunique')\
.fillna(0)
segment_purchase_retention_raw = segment_purchase_retention_raw[segment_purchase_retention_raw
.columns[::-1]].cumsum(axis=1)
segment_purchase_retention_raw = segment_purchase_retention_raw[
segment_purchase_retention_raw.columns[::-1]]
segment_purchase_retention_raw["num_users"] = segment_purchase_retention_raw[0]
segment_purchase_retention_raw = segment_purchase_retention_raw.groupby(
['segment_name', 'segment_id']).sum()
segment_purchase_retention = pd.concat([segment_purchase_retention_raw["num_users"],
segment_purchase_retention_raw
.drop(["num_users"], axis=1)
.div(segment_purchase_retention_raw["num_users"], axis=0)],
axis=1).fillna(0)
segment_purchase_retention=segment_purchase_retention.drop(0,axis=1)
segment_purchase_stats_by_engagement_temp = pd.merge(segment_purchase_data, segment_engagement_user_data,
how='left',
left_on=[
'braze_id', 'segment_id', 'segment_name'],
right_on=[
'braze_id', 'segment_id', 'segment_name'],
suffixes=['_from_retention', '_from_engagement'])
segment_purchase_stats_by_engagement_temp['day_num'] = (
segment_purchase_stats_by_engagement_temp['last_purchase_date'] - segment_purchase_stats_by_engagement_temp['first_purchase_date']).dt.days
segment_purchase_stats_by_engagement_temp['channel'] = segment_purchase_stats_by_engagement_temp.channel.fillna(
'No Messages')
segment_purchase_stats_by_engagement_temp['channel_combo'] = segment_purchase_stats_by_engagement_temp.channel_combo.fillna(
'No Messages')
segment_purchase_stats_by_channel_temp = segment_purchase_stats_by_engagement_temp.groupby(['segment_name', 'segment_id', 'channel']).agg(buyers=('braze_id', 'nunique'), repeat_buyers=('repeat_buyer_id', 'nunique'), num_purchases=(
'num_purchases', 'sum'), total_revenue=('total_revenue', 'sum'), avg_revenue_per_buyer=('total_revenue', 'mean'), avg_time_to_purchase=('days_to_purchase', 'mean'), total_buyers=('braze_id', 'count'), total_repeat_buyers=('repeat_buyer_id', 'count'))
segment_purchase_stats_by_channel_temp['pct_repeat_buyers'] = round(
segment_purchase_stats_by_channel_temp.repeat_buyers/segment_purchase_stats_by_channel_temp.buyers, 2)
segment_purchase_stats_by_channel_temp['purchases_per_buyer'] = round(
segment_purchase_stats_by_channel_temp.num_purchases/segment_purchase_stats_by_channel_temp.total_buyers, 2)
segment_purchase_stats_by_channel_temp['revenue_per_item_purchased'] = round(
segment_purchase_stats_by_channel_temp.total_revenue/segment_purchase_stats_by_channel_temp.num_purchases, 2)
segment_purchase_stats_by_channel = pd.merge(segment_purchase_stats_by_channel_temp, users_per_channel_df,
how='left',
left_on=[
'segment_name', 'segment_id','channel'],
right_on=['segment_name', 'segment_id','channel'])
segment_purchase_stats_by_channel['pct_buyers'] = round(
segment_purchase_stats_by_channel.buyers/segment_purchase_stats_by_channel.num_users, 2)
segment_purchase_stats_by_channel = segment_purchase_stats_by_channel[['segment_name', 'segment_id', 'channel', 'buyers', 'repeat_buyers', 'num_users', 'pct_buyers',
'pct_repeat_buyers', 'purchases_per_buyer', 'avg_revenue_per_buyer', 'avg_time_to_purchase', 'revenue_per_item_purchased']].set_index(['segment_name', 'segment_id', 'channel'])
segment_purchase_stats_by_channel_combo_temp = segment_purchase_stats_by_engagement_temp.groupby(['segment_name', 'segment_id', 'channel_combo']).agg(buyers=('braze_id', 'nunique'), repeat_buyers=('repeat_buyer_id', 'nunique'), num_purchases=(
'num_purchases', 'sum'), total_revenue=('total_revenue', 'sum'), avg_revenue_per_buyer=('total_revenue', 'mean'), avg_time_to_purchase=('days_to_purchase', 'mean'), total_buyers=('braze_id', 'count'), total_repeat_buyers=('repeat_buyer_id', 'count'))
segment_purchase_stats_by_channel_combo_temp['pct_repeat_buyers'] = round(
segment_purchase_stats_by_channel_combo_temp.repeat_buyers/segment_purchase_stats_by_channel_combo_temp.buyers, 2)
segment_purchase_stats_by_channel_combo_temp['purchases_per_buyer'] = round(
segment_purchase_stats_by_channel_combo_temp.num_purchases/segment_purchase_stats_by_channel_combo_temp.total_buyers, 2)
segment_purchase_stats_by_channel_combo_temp['revenue_per_item_purchased'] = round(
segment_purchase_stats_by_channel_combo_temp.total_revenue/segment_purchase_stats_by_channel_combo_temp.num_purchases, 2)
segment_purchase_stats_by_channel_combo = pd.merge(segment_purchase_stats_by_channel_combo_temp, users_per_channel_combo_df,
how='left',
left_on=[
'segment_name', 'segment_id','channel_combo'],
right_on=['segment_name', 'segment_id','channel_combo'])
segment_purchase_stats_by_channel_combo['pct_buyers'] = round(
segment_purchase_stats_by_channel_combo.buyers/segment_purchase_stats_by_channel_combo.num_users, 2)
segment_purchase_stats_by_channel_combo = segment_purchase_stats_by_channel_combo[['segment_name', 'segment_id', 'channel_combo', 'buyers', 'repeat_buyers', 'num_users', 'pct_buyers',
'pct_repeat_buyers', 'purchases_per_buyer', 'avg_revenue_per_buyer', 'avg_time_to_purchase', 'revenue_per_item_purchased']].set_index(['segment_name', 'segment_id', 'channel_combo'])
segment_purchase_stats_by_campaign_tag_temp = segment_purchase_stats_by_engagement_temp.explode(
'tags')
segment_purchase_stats_by_campaign_tag_temp = segment_purchase_stats_by_campaign_tag_temp.groupby(['segment_name', 'segment_id', 'tags']).agg(buyers=('braze_id', 'nunique'), repeat_buyers=('repeat_buyer_id', 'nunique'), num_purchases=(
'num_purchases', 'sum'), total_revenue=('total_revenue', 'sum'), avg_revenue_per_buyer=('total_revenue', 'mean'), avg_time_to_purchase=('days_to_purchase', 'mean'), total_buyers=('braze_id', 'count'), total_repeat_buyers=('repeat_buyer_id', 'count'))
segment_purchase_stats_by_campaign_tag_temp['pct_repeat_buyers'] = round(
segment_purchase_stats_by_campaign_tag_temp.repeat_buyers/segment_purchase_stats_by_campaign_tag_temp.repeat_buyers, 2)
segment_purchase_stats_by_campaign_tag_temp['purchases_per_buyer'] = round(
segment_purchase_stats_by_campaign_tag_temp.num_purchases/segment_purchase_stats_by_campaign_tag_temp.total_buyers, 2)
segment_purchase_stats_by_campaign_tag_temp['revenue_per_item_purchased'] = round(
segment_purchase_stats_by_campaign_tag_temp.total_revenue/segment_purchase_stats_by_campaign_tag_temp.num_purchases, 2)
segment_purchase_stats_by_campaign_tag = pd.merge(segment_purchase_stats_by_campaign_tag_temp, users_per_campaign_tags_df,
how='left',
left_on=[
'segment_name', 'tags'],
right_on=['segment_name', 'tags'])
segment_purchase_stats_by_campaign_tag['pct_buyers'] = round(
segment_purchase_stats_by_campaign_tag.buyers/segment_purchase_stats_by_campaign_tag.num_users, 2)
segment_purchase_stats_by_campaign_tag = segment_purchase_stats_by_campaign_tag[['segment_name', 'segment_id', 'tags', 'buyers', 'repeat_buyers', 'num_users', 'pct_buyers',
'pct_repeat_buyers', 'purchases_per_buyer', 'avg_revenue_per_buyer', 'avg_time_to_purchase', 'revenue_per_item_purchased']].set_index(['segment_name', 'segment_id', 'tags'])
segment_purchase_stats_by_engagement_temp['day_num'] = segment_purchase_stats_by_engagement_temp['day_num'].astype(
'int')
segment_purchase_retention_by_channel_raw = pd.pivot_table(segment_purchase_stats_by_engagement_temp,
values=("braze_id"),
index=(
"segment_name", "segment_id", "channel"),
columns="day_num",
aggfunc='nunique')\
.fillna(0)
segment_purchase_retention_by_channel_raw = segment_purchase_retention_by_channel_raw[segment_purchase_retention_by_channel_raw
.columns[::-1]].cumsum(axis=1)
segment_purchase_retention_by_channel_raw = segment_purchase_retention_by_channel_raw[
segment_purchase_retention_by_channel_raw.columns[::-1]]
segment_purchase_retention_by_channel_raw["num_users"] = segment_purchase_retention_by_channel_raw[0]
segment_purchase_retention_by_channel_raw = segment_purchase_retention_by_channel_raw.groupby(
['segment_name', 'segment_id', "channel"]).sum()
segment_purchase_retention_by_channel = pd.concat([segment_purchase_retention_by_channel_raw["num_users"],
segment_purchase_retention_by_channel_raw
.drop(["num_users"], axis=1)
.div(segment_purchase_retention_by_channel_raw["num_users"], axis=0)],
axis=1).fillna(0)
segment_purchase_retention_by_channel=segment_purchase_retention_by_channel.drop(0, axis=1)
segment_purchase_stats_by_engagement_temp['day_num'] = segment_purchase_stats_by_engagement_temp['day_num'].astype(
'int')
segment_purchase_retention_by_channel_combo_raw = pd.pivot_table(segment_purchase_stats_by_engagement_temp,
values=(
"braze_id"),
index=(
"segment_name", "segment_id", "channel_combo"),
columns="day_num",
aggfunc='nunique')\
.fillna(0)
segment_purchase_retention_by_channel_combo_raw = segment_purchase_retention_by_channel_combo_raw[segment_purchase_retention_by_channel_combo_raw
.columns[::-1]].cumsum(axis=1)
segment_purchase_retention_by_channel_combo_raw = segment_purchase_retention_by_channel_combo_raw[
segment_purchase_retention_by_channel_combo_raw.columns[::-1]]
segment_purchase_retention_by_channel_combo_raw[
"num_users"] = segment_purchase_retention_by_channel_combo_raw[0]
segment_purchase_retention_by_channel_combo_raw = segment_purchase_retention_by_channel_combo_raw.groupby(
['segment_name', 'segment_id', "channel_combo"]).sum()
segment_purchase_retention_by_channel_combo = pd.concat([segment_purchase_retention_by_channel_combo_raw["num_users"],
segment_purchase_retention_by_channel_combo_raw
.drop(["num_users"], axis=1)
.div(segment_purchase_retention_by_channel_combo_raw["num_users"], axis=0)],
axis=1).fillna(0)
segment_purchase_retention_by_channel_combo=segment_purchase_retention_by_channel_combo.drop(0,axis=1)
segment_purchase_stats_by_engagement_temp['day_num'] = segment_purchase_stats_by_engagement_temp['day_num'].astype(
'int')
segment_purchase_stats_by_campaign_tag_temp = segment_purchase_stats_by_engagement_temp.explode(
'tags')
segment_purchase_retention_by_campaign_tags_raw = pd.pivot_table(segment_purchase_stats_by_campaign_tag_temp,
values=(
"braze_id"),
index=(
"segment_name", "segment_id", "tags"),
columns="day_num",
aggfunc='nunique')\
.fillna(0)
### Get the cumulative sum of users based on "last day"
segment_purchase_retention_by_campaign_tags_raw = segment_purchase_retention_by_campaign_tags_raw[segment_purchase_retention_by_campaign_tags_raw
.columns[::-1]].cumsum(axis=1)
segment_purchase_retention_by_campaign_tags_raw = segment_purchase_retention_by_campaign_tags_raw[
segment_purchase_retention_by_campaign_tags_raw.columns[::-1]]
segment_purchase_retention_by_campaign_tags_raw[
"num_users"] = segment_purchase_retention_by_campaign_tags_raw[0]
segment_purchase_retention_by_campaign_tags_raw = segment_purchase_retention_by_campaign_tags_raw.groupby(
['segment_name', 'segment_id', "tags"]).sum()
segment_purchase_retention_by_campaign_tags = pd.concat([segment_purchase_retention_by_campaign_tags_raw["num_users"],
segment_purchase_retention_by_campaign_tags_raw
.drop(["num_users"], axis=1)
.div(segment_purchase_retention_by_campaign_tags_raw["num_users"], axis=0)],
axis=1).fillna(0)
segment_purchase_retention_by_campaign_tags=segment_purchase_retention_by_campaign_tags.drop(0,axis=1)
```
# Session Stats
The following Session Engagement Metrics will be returned by the codeblocks below:
- Number of Users
- Sessions per User
Session Engagement Metrics will returned by:
- Segment
- Channel
- Channel Combo
- Campaign Tag
```
segment_sessions_stats_temp = segment_sessions_stats_pre_apps.explode('apps')
segment_sessions_stats_temp = segment_sessions_stats_temp.dropna(subset=[
'apps'])
segment_sessions_stats_temp['sessions'] = segment_sessions_stats_temp['apps'].apply(
lambda x: x.get('sessions'))
segment_sessions_stats_temp['platform'] = segment_sessions_stats_temp['apps'].apply(
lambda x: x.get('platform'))
segment_sessions_stats_temp = segment_sessions_stats_temp.groupby(['segment_name', 'segment_id']).agg(
num_users=("braze_id", 'nunique'), total_sessions=('sessions', 'sum'))
segment_sessions_stats_temp['sessions_per_user'] = segment_sessions_stats_temp.total_sessions / \
segment_sessions_stats_temp.num_users
segment_sessions_stats = segment_sessions_stats_temp
segment_sessions_stats_by_engagement_temp = segment_engagement_user_data.explode(
'apps')
segment_sessions_stats_by_engagement_temp = segment_sessions_stats_by_engagement_temp.dropna(subset=[
'apps'])
segment_sessions_stats_by_engagement_temp['sessions'] = segment_sessions_stats_by_engagement_temp['apps'].apply(
lambda x: x.get('sessions'))
segment_sessions_stats_by_engagement_temp['platform'] = segment_sessions_stats_by_engagement_temp['apps'].apply(
lambda x: x.get('platform'))
segment_sessions_stats_by_channel_temp = segment_sessions_stats_by_engagement_temp.groupby(['segment_name', 'segment_id', 'channel']).agg(
total_users=("braze_id", 'count'), total_sessions=('sessions', 'sum'), num_users=("braze_id", 'nunique'))
segment_sessions_stats_by_channel_temp = segment_sessions_stats_by_channel_temp.reset_index()
segment_sessions_stats_by_channel_temp['sessions_per_user'] = segment_sessions_stats_by_channel_temp.total_sessions / \
segment_sessions_stats_by_channel_temp.total_users
segment_sessions_stats_by_channel = segment_sessions_stats_by_channel_temp[[
'segment_name', 'segment_id', 'channel', 'num_users', 'sessions_per_user']].set_index(['segment_name', 'segment_id', 'channel'])
segment_sessions_stats_by_channel_combo_temp = segment_sessions_stats_by_engagement_temp.groupby(['segment_name', 'segment_id', 'channel_combo']).agg(
total_users=("braze_id", 'count'), total_sessions=('sessions', 'sum'), num_users=("braze_id", 'nunique'))
segment_sessions_stats_by_channel_combo_temp = segment_sessions_stats_by_channel_combo_temp.reset_index()
segment_sessions_stats_by_channel_combo_temp['sessions_per_user'] = segment_sessions_stats_by_channel_combo_temp.total_sessions / \
segment_sessions_stats_by_channel_combo_temp.total_users
segment_sessions_stats_by_channel_combo = segment_sessions_stats_by_channel_combo_temp[[
'segment_name', 'segment_id', 'channel_combo', 'num_users', 'sessions_per_user']].set_index(['segment_name', 'segment_id', 'channel_combo'])
segment_sessions_stats_by_campaign_tag_temp = segment_sessions_stats_by_engagement_temp.explode(
'tags')
segment_sessions_stats_by_campaign_tag_temp = segment_sessions_stats_by_campaign_tag_temp.groupby(['segment_name', 'segment_id', 'tags']).agg(
total_users=("braze_id", 'count'), total_sessions=('sessions', 'sum'), num_users=("braze_id", 'nunique'))
segment_sessions_stats_by_campaign_tag_temp = segment_sessions_stats_by_campaign_tag_temp.reset_index()
segment_sessions_stats_by_campaign_tag_temp['sessions_per_user'] = segment_sessions_stats_by_campaign_tag_temp.total_sessions / \
segment_sessions_stats_by_campaign_tag_temp.total_users
segment_sessions_stats_by_campaign_tag = segment_sessions_stats_by_campaign_tag_temp[[
'segment_name', 'segment_id', 'tags', 'num_users', 'sessions_per_user']].set_index(['segment_name', 'segment_id', 'tags'])
```
# Custom Event Stats
The following Custom Events Stats will be calculated:
- Number of Users Completing the Custom Event
- Number of Users
- Total Count of Custom Event
- % of Users Completing Custom Events
- Custom Events per User
- Avg. Days between each occurence of a Custom Event
- Avg. Custom Event Completion per Day
Custom Event stats dataframes will returned by:
- Segment
- Channel
- Channel Combo
- Campaign Tag
```
segment_custom_event_stats_temp = segment_custom_event_stats_pre_custom_event.explode(
'custom_events')
segment_custom_event_stats_temp['custom_event_name'] = list(
map(get_custom_event_name, segment_custom_event_stats_temp['custom_events']))
segment_custom_event_stats_temp['custom_event_count'] = list(
map(get_custom_event_count, segment_custom_event_stats_temp['custom_events']))
segment_custom_event_stats_temp['custom_event_first_date'] = list(map(
get_custom_event_first_date, segment_custom_event_stats_temp['custom_events']))
segment_custom_event_stats_temp['custom_event_last_date'] = list(
map(get_custom_event_last_date, segment_custom_event_stats_temp['custom_events']))
segment_custom_event_stats_temp[['custom_event_first_date', 'custom_event_last_date']] = segment_custom_event_stats_temp[[
'custom_event_first_date', 'custom_event_last_date']].apply(pd.to_datetime)
segment_custom_event_stats_temp['days_between_events'] = (
segment_custom_event_stats_temp['custom_event_last_date'] - segment_custom_event_stats_temp['custom_event_first_date']).dt.days
segment_custom_event_stats_temp['custom_event_per_day'] = np.round(np.where(segment_custom_event_stats_temp['days_between_events'] > 0,
segment_custom_event_stats_temp.custom_event_count/segment_custom_event_stats_temp.days_between_events, segment_custom_event_stats_temp.custom_event_count), 1)
total_segment_users_custom_event = segment_custom_event_stats_temp.braze_id.nunique()
segment_custom_event_stats_by_segment = segment_custom_event_stats_temp.groupby(
['segment_name', 'segment_id', 'custom_event_name']).agg(num_users_completing_custom_event=(
'braze_id', 'nunique'), total_custom_events=('custom_event_count', 'sum'), avg_days_between_events=('days_between_events', 'mean'), avg_custom_event_per_day=('custom_event_per_day', 'mean'))
segment_custom_event_stats_by_segment['custom_event_per_user'] = segment_custom_event_stats_by_segment.total_custom_events / \
total_segment_users_custom_event
segment_custom_event_stats_by_segment['pct_users_completing_custom_event'] = segment_custom_event_stats_by_segment.num_users_completing_custom_event / \
total_segment_users_custom_event
segment_custom_event_stats_by_segment['num_users'] = total_segment_users_custom_event
segment_custom_event_stats_by_segment = segment_custom_event_stats_by_segment[[
'num_users_completing_custom_event', 'num_users', 'total_custom_events', 'pct_users_completing_custom_event', 'custom_event_per_user', 'avg_days_between_events', 'avg_custom_event_per_day']]
segment_custom_event_stats_by_engagement_temp = pd.merge(segment_custom_event_stats_temp, segment_engagement_user_data,
how='left',
left_on=[
'braze_id', 'segment_id', 'segment_name'],
right_on=[
'braze_id', 'segment_id', 'segment_name'],
suffixes=['_from_custom_events', '_from_engagement'])
segment_custom_event_stats_by_engagement_temp['channel'] = segment_custom_event_stats_by_engagement_temp.channel.fillna(
'No Messages')
segment_custom_event_stats_by_engagement_temp['channel_combo'] = segment_custom_event_stats_by_engagement_temp.channel_combo.fillna(
'No Messages')
segment_custom_event_stats_by_engagement_temp['tags'] = segment_custom_event_stats_by_engagement_temp.tags.fillna(
'No Messages')
segment_custom_event_stats_by_segment_and_channel_temp = segment_custom_event_stats_by_engagement_temp.groupby(['segment_name', 'segment_id', 'channel', 'custom_event_name']).agg(num_users_completing_custom_event=(
'braze_id', 'nunique'), total_custom_events=('custom_event_count', 'sum'), avg_days_between_events=('days_between_events', 'mean'), avg_custom_event_per_day=('custom_event_per_day', 'mean'))
segment_custom_event_stats_by_segment_and_channel_temp = segment_custom_event_stats_by_segment_and_channel_temp.reset_index()
segment_custom_event_stats_by_segment_and_channel = pd.merge(segment_custom_event_stats_by_segment_and_channel_temp, users_per_channel_df,
how='left',
left_on=[
'segment_name', 'segment_id','channel'],
right_on=['segment_name', 'segment_id','channel'])
segment_custom_event_stats_by_segment_and_channel['custom_event_per_user'] = segment_custom_event_stats_by_segment_and_channel.total_custom_events / \
segment_custom_event_stats_by_segment_and_channel.num_users
segment_custom_event_stats_by_segment_and_channel['pct_users_completing_custom_event'] = segment_custom_event_stats_by_segment_and_channel.num_users_completing_custom_event / \
segment_custom_event_stats_by_segment_and_channel.num_users
segment_custom_event_stats_by_segment_and_channel = segment_custom_event_stats_by_segment_and_channel[['segment_name', 'segment_id', 'channel','custom_event_name', 'num_users_completing_custom_event', 'num_users',
'total_custom_events', 'pct_users_completing_custom_event', 'custom_event_per_user', 'avg_days_between_events', 'avg_custom_event_per_day']].set_index(['segment_name', 'segment_id', 'channel'])
segment_custom_event_stats_by_segment_and_channel_combo_temp = segment_custom_event_stats_by_engagement_temp.groupby(['segment_name', 'segment_id', 'channel_combo', 'custom_event_name']).agg(num_users_completing_custom_event=(
'braze_id', 'nunique'), total_custom_events=('custom_event_count', 'sum'), avg_days_between_events=('days_between_events', 'mean'), avg_custom_event_per_day=('custom_event_per_day', 'mean'))
segment_custom_event_stats_by_segment_and_channel_combo_temp = segment_custom_event_stats_by_segment_and_channel_combo_temp.reset_index()
segment_custom_event_stats_by_segment_and_channel_combo = pd.merge(segment_custom_event_stats_by_segment_and_channel_combo_temp, users_per_channel_combo_df,
how='left',
left_on=[
'segment_name', 'segment_id','channel_combo'],
right_on=['segment_name', 'segment_id','channel_combo'])
segment_custom_event_stats_by_segment_and_channel_combo['custom_event_per_user'] = segment_custom_event_stats_by_segment_and_channel_combo.total_custom_events / \
segment_custom_event_stats_by_segment_and_channel_combo.num_users
segment_custom_event_stats_by_segment_and_channel_combo['pct_users_completing_custom_event'] = segment_custom_event_stats_by_segment_and_channel_combo.num_users_completing_custom_event / \
segment_custom_event_stats_by_segment_and_channel_combo.num_users
segment_custom_event_stats_by_segment_and_channel_combo = segment_custom_event_stats_by_segment_and_channel_combo[['segment_name', 'segment_id', 'channel_combo', 'custom_event_name','num_users_completing_custom_event', 'num_users',
'total_custom_events', 'pct_users_completing_custom_event', 'custom_event_per_user', 'avg_days_between_events', 'avg_custom_event_per_day']].set_index(['segment_name', 'segment_id', 'channel_combo'])
segment_custom_event_stats_by_segment_and_campaign_tags_df = segment_custom_event_stats_by_engagement_temp.explode(
'tags')
segment_custom_event_stats_by_segment_and_campaign_tags_temp = segment_custom_event_stats_by_segment_and_campaign_tags_df.groupby(['segment_name', 'segment_id', 'tags', 'custom_event_name']).agg(num_users_completing_custom_event=(
'braze_id', 'nunique'), total_custom_events=('custom_event_count', 'sum'), avg_days_between_events=('days_between_events', 'mean'), avg_custom_event_per_day=('custom_event_per_day', 'mean'))
segment_custom_event_stats_by_segment_and_campaign_tags_temp = segment_custom_event_stats_by_segment_and_campaign_tags_temp.reset_index()
segment_custom_event_stats_by_segment_and_campaign_tags = pd.merge(segment_custom_event_stats_by_segment_and_campaign_tags_temp, users_per_campaign_tags_df,
how='left',
left_on=[
'segment_name', 'segment_id','tags'],
right_on=['segment_name', 'segment_id','tags'])
segment_custom_event_stats_by_segment_and_campaign_tags['custom_event_per_user'] = segment_custom_event_stats_by_segment_and_campaign_tags.total_custom_events / \
segment_custom_event_stats_by_segment_and_campaign_tags.num_users
segment_custom_event_stats_by_segment_and_campaign_tags['pct_users_completing_custom_event'] = segment_custom_event_stats_by_segment_and_campaign_tags.num_users_completing_custom_event / \
segment_custom_event_stats_by_segment_and_campaign_tags.num_users
segment_custom_event_stats_by_segment_and_campaign_tags = segment_custom_event_stats_by_segment_and_campaign_tags[[
'segment_name', 'segment_id', 'tags', 'custom_event_name','num_users_completing_custom_event', 'num_users', 'total_custom_events', 'pct_users_completing_custom_event', 'custom_event_per_user', 'avg_days_between_events', 'avg_custom_event_per_day']].set_index(['segment_name', 'segment_id', 'tags'])
```
## Custom Events Used in Campaigns
The codeblock below will return all custom events that used in campaigns received by the selected segment.
```
campaign_details_custom_event_temp = campaign_details_df[[
'campaign_id', 'conversion_behaviors']]
campaign_details_custom_event_temp = campaign_details_custom_event_temp.dropna(
subset=['conversion_behaviors'])
campaign_details_custom_event_temp = campaign_details_custom_event_temp.explode(
'conversion_behaviors')
campaign_details_custom_event_temp['custom_event_conversion_behavior'] = list(map(
get_campaign_custom_event, campaign_details_custom_event_temp['conversion_behaviors']))
campaign_details_custom_event_temp = campaign_details_custom_event_temp.dropna(
subset=['custom_event_conversion_behavior'])
campaign_details_custom_event = campaign_details_custom_event_temp[[
'campaign_id', 'custom_event_conversion_behavior']].drop_duplicates()
campaign_details_custom_event = campaign_details_custom_event.set_index(
'campaign_id')
```
# Segment Opt-In Rates
The codeblock below will return the opt-in rates for Push and Email for all users across the following platforms:
- iOS
- Android
- Web
```
segment_opt_ins_temp = segment_opt_in_stats_pre_apps.explode('apps')
segment_opt_ins_temp = segment_opt_ins_temp.dropna(subset=['apps'])
segment_opt_ins_temp = segment_opt_ins_temp.explode('push_tokens')
segment_opt_ins_temp['notifications_enabled'] = list(
map(get_notifications_enabled, segment_opt_ins_temp['push_tokens']))
segment_opt_ins_temp['token'] = list(
map(get_token, segment_opt_ins_temp['push_tokens']))
segment_opt_ins_temp['push_token_platform'] = list(
map(get_platform, segment_opt_ins_temp['push_tokens']))
segment_opt_ins_temp['app_platform'] = segment_opt_ins_temp['apps'].apply(
lambda x: x.get('platform'))
segment_opt_ins_temp_android = segment_opt_ins_temp[segment_opt_ins_temp['app_platform'] == 'Android'].copy()
segment_opt_ins_temp_android['push_opted_in'] = segment_opt_ins_temp.apply(lambda x: True
if x['notifications_enabled'] == True and x['token'] != "None" else False, axis=1)
segment_opt_ins_temp_android['email_opted_in'] = segment_opt_ins_temp.apply(lambda x: True
if x['email_subscribe'] == 'opted_in' else False, axis=1)
segment_opt_ins_temp_ios = segment_opt_ins_temp[segment_opt_ins_temp['app_platform'] == 'iOS'].copy()
segment_opt_ins_temp_ios['push_opted_in'] = segment_opt_ins_temp.apply(lambda x: True
if x['notifications_enabled'] == True and x['token'] != "None" else False, axis=1)
segment_opt_ins_temp_ios['email_opted_in'] = segment_opt_ins_temp.apply(lambda x: True
if x['email_subscribe'] == 'opted_in' else False, axis=1)
segment_opt_ins_temp_web = segment_opt_ins_temp[segment_opt_ins_temp['app_platform'] == 'Web'].copy()
segment_opt_ins_temp_web['push_opted_in'] = segment_opt_ins_temp.apply(lambda x: True
if x['notifications_enabled'] == True and x['token'] != "None" else False, axis=1)
segment_opt_ins_temp_web['email_opted_in'] = segment_opt_ins_temp.apply(lambda x: True
if x['email_subscribe'] == 'opted_in' else False, axis=1)
segment_opt_ins_android_pre_agg = segment_opt_ins_temp_android.groupby(
['segment_id', 'segment_name', 'app_platform'])
opt_ins_aggregator = {'push_opted_in': calc_engagement,
'email_opted_in': calc_engagement}
segment_opt_ins_android = segment_opt_ins_android_pre_agg.agg(
opt_ins_aggregator)
segment_opt_ins_ios_pre_agg = segment_opt_ins_temp_ios.groupby(
['segment_id', 'segment_name', 'app_platform'])
segment_opt_ins_ios = segment_opt_ins_ios_pre_agg.agg(opt_ins_aggregator)
segment_opt_ins_web_pre_agg = segment_opt_ins_temp_web.groupby(
['segment_id', 'segment_name', 'app_platform'])
segment_opt_ins_web = segment_opt_ins_web_pre_agg.agg(opt_ins_aggregator)
segment_opt_ins = pd.concat(
[segment_opt_ins_android, segment_opt_ins_ios, segment_opt_ins_web])
```
## Exporting Outputs to Excel
Please note that attempting to export dataframes that were not created will result in an error.
```
file_name = "Segment Analytics {date}.xlsx".format(date = datetime.now().date())
writer = pd.ExcelWriter(file_name, engine='xlsxwriter')
engagement_by_segment.to_excel(writer, sheet_name='Eng. by Segment')
engagement_by_segment_and_channel.to_excel(
writer, sheet_name='Eng. by Channel')
engagement_by_segment_and_channel_combo.to_excel(
writer, sheet_name='Eng. by Channel Combo')
engagement_by_segment_and_campaign_tag.to_excel(
writer, sheet_name='Eng. by Campaign Tag')
segment_rolling_retention.to_excel(writer, sheet_name='Ret. by Segment')
segment_rolling_retention_by_engagement.to_excel(
writer, sheet_name='Ret. by Channel')
segment_rolling_retention_by_channel_combo.to_excel(
writer, sheet_name='Ret. by Channel Combo')
segment_rolling_retention_by_campaign_tag.to_excel(
writer, sheet_name='Ret. by Campaign Tag')
segment_purchasing_stats.to_excel(writer, sheet_name='Purch. Stats by Segment')
segment_purchase_stats_by_channel.to_excel(
writer, sheet_name='Purch. Stats by Channel')
segment_purchase_stats_by_channel_combo.to_excel(
writer, sheet_name='Purch. Stats by Combo')
segment_purchase_stats_by_campaign_tag.to_excel(
writer, sheet_name='Purch. Stats by Campaign Tag')
segment_purchase_retention.to_excel(writer, sheet_name='Purch. Ret by Segment')
segment_purchase_retention_by_channel.to_excel(
writer, sheet_name='Purch. Ret by Channel')
segment_purchase_retention_by_channel_combo.to_excel(
writer, sheet_name='Purch. Ret by Combo')
segment_purchase_retention_by_campaign_tags.to_excel(
writer, sheet_name='Purch. Ret by Campaign Tag')
segment_sessions_stats.to_excel(writer, sheet_name='Sess. Stats by Segment')
segment_sessions_stats_by_channel.to_excel(
writer, sheet_name='Sess. Stats by Channel')
segment_sessions_stats_by_channel_combo.to_excel(
writer, sheet_name='Sess. Stats by Combo')
segment_sessions_stats_by_campaign_tag.to_excel(
writer, sheet_name='Sess. Stats by Campaign Tag')
segment_custom_event_stats_by_segment.to_excel(
writer, sheet_name='CE Stats by Segment')
segment_custom_event_stats_by_segment_and_channel.to_excel(
writer, sheet_name='CE Stats by Channel')
segment_custom_event_stats_by_segment_and_channel_combo.to_excel(
writer, sheet_name='CE Stats by Combo')
segment_custom_event_stats_by_segment_and_campaign_tags.to_excel(
writer, sheet_name='CE Stats by Campaign Tag')
campaign_details_custom_event.to_excel(
writer, sheet_name='CE Used in Campaigns')
segment_opt_ins.to_excel(writer, sheet_name='Opt-Ins by Segment')
writer.save()
```
| github_jupyter |
### Importing the libraries
```
import torch
import torch.nn as nn
from torch import unsqueeze
class SelfAttention(nn.Module):
def __init__(self, embed_size, heads):
"""
We are going to split the embed size between these heads. If we have 256 size embedding and 8 heads then we will have 32 embed size for each embedding
"""
super(SelfAttention,self).__init__()
self.embed_size = embed_size
self.heads = heads
assert self.embed_size%self.heads == 0 , "To make sure that embed size is properly divisible by heads"
self.head_dim = embed_size//heads
"""
Now we are defining the Query value and key vectors as Linear layers.
We are setting bias = False, because we dont need that
"""
self.values = nn.Linear(self.head_dim, self.head_dim, bias = False)
self.keys = nn.Linear(self.head_dim, self.head_dim, bias = False)
self.queries = nn.Linear(self.head_dim, self.head_dim, bias = False)
self.fc_out = nn.Linear(heads*self.head_dim, embed_size)
def forward(self, values, keys, queries, mask):
N = queries.shape[0]
value_len, key_len, query_len = values.shape[1], keys.shape[1], queries.shape[1]
"""
split embedding into self.heads pieces
"""
values = values.reshape(N, value_len, self.heads, self.head_dim)
keys = keys.reshape(N, key_len, self.heads, self.head_dim)
queries = queries.reshape(N, query_len, self.heads, self.head_dim)
values = self.values(values)
keys = self.keys(keys)
queries = self.queries(queries)
#step 1: multiply query and key
# queries shape : (N, query_len, heads, heads_dim)
# keys shape: (N, key_len, heads, heads_dim)
# energy shape: (N, heads,query_len = target source sentence, key_len = source sentence)
"""
As we have a batch matrix multiplier einsum is quite handy for it
"""
energy = torch.einsum("nqhd,nkhd->nhqk", [queries,keys]) #it is used for matrix multiplication where we have several other dimensions
if mask is not None:
energy = energy.masked_fill(mask==0, float("-1e20"))
attention = torch.softmax(energy/(self.embed_size**(1/2)),dim=3)
#attention shape: (N,heads, query_len, key_len)
#value shape: (N, value_len, heads, heads_dim)
#out shape: (N, Query_len, heads, heads_dim)
out = torch.einsum("nhql,nlhd->nqhd", [attention,values])
#concatanation part
out = out.reshape(N,query_len, self.heads*self.head_dim)
out = self.fc_out(out)
return out
```
### Transformer Block
```
class TransformerBlock(nn.Module):
"""
embedding -> multiheaded_attention -> add&norm -> feed forward -> add&norm
"""
def __init__(self, embed_size, heads, dropout, forward_expansion):
super(TransformerBlock,self).__init__()
self.attention = SelfAttention(embed_size=embed_size, heads = heads)
self.norm1 = nn.LayerNorm(embed_size) #layernorm and batchnorm are almost similar...but layer norm has more computation
self.norm2 = nn.LayerNorm(embed_size)
self.feed_forward = nn.Sequential(
nn.Linear(embed_size, forward_expansion*embed_size),
nn.ReLU(),
nn.Linear(forward_expansion*embed_size,embed_size)
)
self.dropout = nn.Dropout(dropout)
def forward(self, value, key, query, mask):
attention = self.attention(value, key, query, mask)
"""
we needed a skip connection. query is for the skip connection
"""
x = self.dropout(self.norm1(attention + query))
forward = self.feed_forward(x)
out = self.dropout(self.norm2(forward+x))
print(out)
return out
class Encoder(nn.Module):
def __init__(self, src_vocab_size, embed_size, num_layers, heads, device, forward_expansion, dropout, max_length):
"""
Encoder block takes a lot of parameters due to hyperparameter. The parameters are explained below:
---------------------------------------------------------------------------------------------------
src_vocab_size = size of source vocabulary
embed_size = dimension of embedding
num_layers = number of transformer layer in encoder
heads = number of heads in multiheads
device = the device on which we want to train
forward_expansion = the ratio by which we want to expand the size
dropout = dropout probability
max_length = max sentence length.
maximum length of string to ensure positional embedding which is requeired for ensuring we have attention.
What transformer does is we wnat to ensure that some sort of sequence is maintained even is the layer does not have any recurrent unit. It helps the transformer for ensuring parallelization
"""
super(Encoder,self).__init__()
self.embed_size = embed_size
self.device = device
self.word_embedding = nn.Embedding(src_vocab_size, embed_size)
self.positional_embedding = nn.Embedding(max_length, embed_size)
self.layers = nn.ModuleList(
[
TransformerBlock(embed_size=embed_size, heads=heads, dropout=dropout, forward_expansion=forward_expansion) for _ in range(num_layers)
]
)
self.dropout = nn.Dropout(dropout)
def forward(self, x, mask):
N, seq_length = x.shape
positions = torch.arange(0,seq_length).expand(N,seq_length).to(self.device)
out = self.dropout(self.word_embedding(x)+self.positional_embedding(positions))
for layer in self.layers:
out = layer(out,out,out,mask)
return out
class DecoderBlock(nn.Module):
def __init__(self, embed_size, heads, forward_expansion, dropout, device):
super(DecoderBlock,self).__init__()
self.attention = SelfAttention(embed_size, heads)
self.norm = nn.LayerNorm(embed_size)
self.transformer_block = TransformerBlock(embed_size, heads, dropout, forward_expansion)
self.dropout = nn.Dropout(dropout)
def forward(self, x, value, key, src_mask, trg_mask):
"""
Decoder block takes a lot of parameters. The parameters are explained below:
----------------------------------------------------------------------------
x : input
value, key : for self_attention
src_mask: source mask. Although it is optional still we need it. For example, let we have more than one example in the input. In those cases src_mask is needed to make all the sentences equal also we dont need to to extra computations for the masks that are padded
trg_mask: trg_mask is required to make sure that everything works fine
"""
attention = self.attention(x,x,x,trg_mask)
query = self.dropout(self.norm(attention+x))
out = self.transformer_block(value, key, query, src_mask)
return out
class Decoder(nn.Module):
def __init__(self, trg_vocab_size, embed_size, num_layers, heads, forward_expansion, dropout, device, max_length):
super(Decoder,self).__init__()
self.device = device
self.word_embedding = nn.Embedding(trg_vocab_size, embed_size)
self.positional_embedding = nn.Embedding(max_length,embed_size)
self.layers = nn.ModuleList(
[DecoderBlock(embed_size, heads, forward_expansion, dropout, device) for _ in range(num_layers)]
)
self.fc_out = nn.Linear(embed_size, trg_vocab_size)
self.dropout = nn.Dropout(dropout)
def forward(self, x, enc_out, src_mask, trg_mask):
N, seq_length = x.shape
positions = torch.arange(0,seq_length).expand(N,seq_length).to(self.device)
x = self.dropout(self.word_embedding(x)+self.positional_embedding(positions))
for layer in self.layers:
x = layer(x,enc_out, enc_out, src_mask, trg_mask)
out = self.fc_out(x)
return out
class Transformer(nn.Module):
def __init__(self, src_vocab_size, trg_vocab_size, src_pad_idx, trg_pad_idx, embed_size = 256, num_layers = 6, forward_expansion = 4, heads = 8, dropout = 0, device = "cuda", max_length = 100):
super(Transformer,self).__init__()
self.encoder = Encoder(src_vocab_size,embed_size, num_layers, heads, device, forward_expansion, dropout, max_length)
self.decoder = Decoder(trg_vocab_size,embed_size, num_layers, heads, forward_expansion, dropout, device, max_length)
self.src_pad_idx = src_pad_idx
self.trg_pad_idx = trg_pad_idx
self.device = device
def make_src_mask(self,src):
src_mask = (src != self.src_pad_idx).unsqueeze(1).unsqueeze(2)
return src_mask.to(self.device)
def make_trg_mask(self,trg):
N, trg_len = trg.shape
trg_mask = torch.tril(torch.ones((trg_len,trg_len))).expand(N,1,trg_len, trg_len)
return trg_mask.to(self.device)
def forward(self, src, trg):
src_mask = self.make_src_mask(src)
trg_mask = self.make_trg_mask(trg)
enc_src = self.encoder(src, src_mask)
out = self.decoder(trg, enc_src, src_mask, trg_mask)
return out
if __name__ == '__main__':
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
x = torch.tensor([[1,5,6,4,3,9,5,2,0],[1,8,7,3,4,5,6,7,2]]).to(device)
trg = torch.tensor([[1,7,4,3,5,9,2,0],[1,8,7,3,4,5,6,2]]).to(device)
src_pad_idx = 0
trg_pad_idx = 0
src_vocab_size = 10
trg_vocab_Size = 10
model = Transformer(src_vocab_size, trg_vocab_Size, src_pad_idx, trg_pad_idx).to(device)
out = model(x,trg[:,:-1])
print(out.shape)
```
| github_jupyter |
# DJL BERT Inference Demo
## Introduction
In this tutorial, you walk through running inference using DJL on a [BERT](https://towardsdatascience.com/bert-explained-state-of-the-art-language-model-for-nlp-f8b21a9b6270) QA model trained with MXNet.
You can provide a question and a paragraph containing the answer to the model. The model is then able to find the best answer from the answer paragraph.
Example:
```text
Q: When did BBC Japan start broadcasting?
```
Answer paragraph:
```text
BBC Japan was a general entertainment channel, which operated between December 2004 and April 2006.
It ceased operations after its Japanese distributor folded.
```
And it picked the right answer:
```text
A: December 2004
```
## Preparation
This tutorial requires the installation of Java Kernel. To install the Java Kernel, see the [README](https://github.com/awslabs/djl/blob/master/jupyter/README.md).
```
%maven ai.djl:api:0.2.0
%maven ai.djl.mxnet:mxnet-engine:0.2.0
%maven ai.djl:repository:0.2.0
%maven ai.djl.mxnet:mxnet-model-zoo:0.2.0
%maven org.slf4j:slf4j-api:1.7.26
%maven org.slf4j:slf4j-simple:1.7.26
%maven net.java.dev.jna:jna:5.3.0
```
### Include MXNet engine dependency
This tutorial uses MXNet engine as its backend. MXNet has different [build flavor](https://mxnet.apache.org/get_started?version=v1.5.1&platform=linux&language=python&environ=pip&processor=cpu) and it is platform specific.
Please read [here](https://github.com/awslabs/djl/blob/master/examples/README.md#engine-selection) for how to select MXNet engine flavor.
```
String classifier = System.getProperty("os.name").startsWith("Mac") ? "osx-x86_64" : "linux-x86_64";
%maven ai.djl.mxnet:mxnet-native-mkl:jar:${classifier}:1.6.0-a
```
### Import java packages by running the following:
```
import java.io.*;
import java.nio.charset.*;
import java.nio.file.*;
import java.util.*;
import com.google.gson.*;
import com.google.gson.annotations.*;
import ai.djl.*;
import ai.djl.inference.*;
import ai.djl.metric.*;
import ai.djl.mxnet.zoo.*;
import ai.djl.mxnet.zoo.nlp.qa.*;
import ai.djl.repository.zoo.*;
import ai.djl.ndarray.*;
import ai.djl.ndarray.types.*;
import ai.djl.training.util.*;
import ai.djl.translate.*;
import ai.djl.util.*;
```
Now that all of the prerequisites are complete, start writing code to run inference with this example.
## Load the model and input
The model requires three inputs:
- word indices: The index of each word in a sentence
- word types: The type index of the word. All Questions will be labelled with 0 and all Answers will be labelled with 1.
- sequence length: You need to limit the length of the input. In this case, the length is 384
- valid length: The actual length of the question and answer tokens
**First, load the input**
```
var question = "When did BBC Japan start broadcasting?";
var resourceDocument = "BBC Japan was a general entertainment Channel.\n" +
"Which operated between December 2004 and April 2006.\n" +
"It ceased operations after its Japanese distributor folded.";
QAInput input = new QAInput(question, resourceDocument, 384);
```
Then load the model and vocabulary. Create a variable `model` by using the `ModelZoo` as shown in the following code.
```
Map<String, String> criteria = new ConcurrentHashMap<>();
criteria.put("backbone", "bert");
criteria.put("dataset", "book_corpus_wiki_en_uncased");
ZooModel<QAInput, String> model = MxModelZoo.BERT_QA.loadModel(criteria, new ProgressBar());
```
## Run inference
Once the model is loaded, you can call `Predictor` and run inference as follows
```
Predictor<QAInput, String> predictor = model.newPredictor();
String answer = predictor.predict(input);
answer
```
Running inference on DJL is that easy. In the example, you use a model from the `ModelZoo`. However, you can also load the model on your own and use custom classes as the input and output. The process for that is illustrated in greater detail later in this tutorial.
## Dive deep into Translator
Inference in deep learning is the process of predicting the output for a given input based on a pre-defined model.
DJL abstracts away the whole process for ease of use. It can load the model, perform inference on the input, and provide
output. DJL also allows you to provide user-defined inputs. The workflow looks like the following:
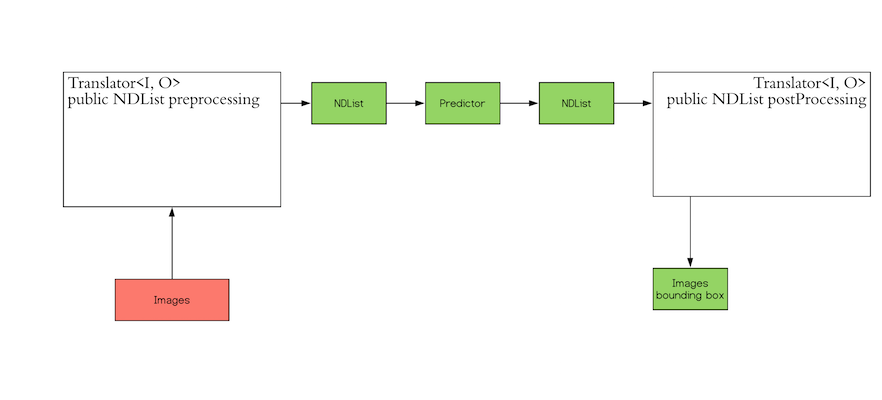
The red block ("Images") in the workflow is the input that DJL expects from you. The green block ("Images
bounding box") is the output that you expect. Because DJL does not know which input to expect and which output format that you prefer, DJL provides the `Translator` interface so you can define your own
input and output.
The `Translator` interface encompasses the two white blocks: Pre-processing and Post-processing. The pre-processing
component converts the user-defined input objects into an NDList, so that the `Predictor` in DJL can understand the
input and make its prediction. Similarly, the post-processing block receives an NDList as the output from the
`Predictor`. The post-processing block allows you to convert the output from the `Predictor` to the desired output
format.
### Pre-processing
Now, you need to convert the sentences into tokens. You can use `BertDataParser.tokenizer` to convert questions and answers into tokens. Then, use `BertDataParser.formTokens` to create Bert-Formatted tokens. Once you have properly formatted tokens, use `parser.token2idx` to create the indices.
The following code block converts the question and answer defined earlier into bert-formatted tokens and creates word types for the tokens.
```
// Create token lists for question and answer
List<String> tokenQ = BertDataParser.tokenizer(question.toLowerCase());
List<String> tokenA = BertDataParser.tokenizer(resourceDocument.toLowerCase());
int validLength = tokenQ.size() + tokenA.size();
System.out.println("Question Token: " + tokenQ);
System.out.println("Answer Token: " + tokenA);
System.out.println("Valid length: " + validLength);
```
Normally, words/sentences are represented as indices instead of Strings for training. They typically work like a vector in a n-dimensional space. In this case, you need to map them into indices. The form tokens also pad the sentence to the required length.
```
// Create Bert-formatted tokens
List<String> tokens = BertDataParser.formTokens(tokenQ, tokenA, 384);
// Convert tokens into indices in the vocabulary
BertDataParser parser = model.getArtifact("vocab.json", BertDataParser::parse);
List<Integer> indices = parser.token2idx(tokens);
```
Finally, the model needs to understand which part is the Question and which part is the Answer. Mask the tokens as follows:
```
[Question tokens...AnswerTokens...padding tokens] => [000000...11111....0000]
```
```
// Get token types
List<Float> tokenTypes = BertDataParser.getTokenTypes(tokenQ, tokenA, 384);
```
To properly convert them into `float[]` for `NDArray` creation, here is the helper function:
```
/**
* Convert a List of Number to float array.
*
* @param list the list to be converted
* @return float array
*/
public static float[] toFloatArray(List<? extends Number> list) {
float[] ret = new float[list.size()];
int idx = 0;
for (Number n : list) {
ret[idx++] = n.floatValue();
}
return ret;
}
float[] indicesFloat = toFloatArray(indices);
float[] types = toFloatArray(tokenTypes);
```
Now that you have everything you need, you can create an NDList and populate all of the inputs you formatted earlier. You're done with pre-processing!
#### Construct `Translator`
You need to do this processing within an implementation of the `Translator` interface. `Translator` is designed to do pre-processing and post-processing. You must define the input and output objects. It contains the following two override classes:
- `public NDList processInput(TranslatorContext ctx, I)`
- `public String processOutput(TranslatorContext ctx, O)`
Every translator takes in input and returns output in the form of generic objects. In this case, the translator takes input in the form of `QAInput` (I) and returns output as a `String` (O). `QAInput` is just an object that holds questions and answer; We have prepared the Input class for you.
Armed with the needed knowledge, you can write an implementation of the `Translator` interface. `BertTranslator` uses the code snippets explained previously to implement the `processInput`method. For more information, see [`NDManager`](https://javadoc.djl.ai/api/0.2.0/ai/djl/ndarray/NDManager.html).
```
manager.create(Number[] data, Shape)
manager.create(Number[] data)
```
The `Shape` for `data0` and `data1` is (num_of_batches, sequence_length). For `data2` is just 1.
```
public class BertTranslator implements Translator<QAInput, String> {
private BertDataParser parser;
private List<String> tokens;
private int seqLength;
BertTranslator(BertDataParser parser) {
this.parser = parser;
this.seqLength = 384;
}
@Override
public Batchifier getBatchifier() {
return null;
}
@Override
public NDList processInput(TranslatorContext ctx, QAInput input) throws IOException {
BertDataParser parser = ctx.getModel().getArtifact("vocab.json", BertDataParser::parse);
// Pre-processing - tokenize sentence
// Create token lists for question and answer
List<String> tokenQ = BertDataParser.tokenizer(question.toLowerCase());
List<String> tokenA = BertDataParser.tokenizer(resourceDocument.toLowerCase());
// Calculate valid length (length(Question tokens) + length(resourceDocument tokens))
var validLength = tokenQ.size() + tokenA.size();
// Create Bert-formatted tokens
tokens = BertDataParser.formTokens(tokenQ, tokenA, 384);
if (tokens == null) {
throw new IllegalStateException("tokens is not defined");
}
// Convert tokens into indices in the vocabulary
List<Integer> indices = parser.token2idx(tokens);
// Get token types
List<Float> tokenTypes = BertDataParser.getTokenTypes(tokenQ, tokenA, 384);
NDManager manager = ctx.getNDManager();
// Using the manager created, create NDArrays for the indices, types, and valid length.
// in that order. The type of the NDArray should all be float
NDArray indicesNd = manager.create(toFloatArray(indices), new Shape(1, 384));
indicesNd.setName("data0");
NDArray typesNd = manager.create(toFloatArray(tokenTypes), new Shape(1, 384));
typesNd.setName("data1");
NDArray validLengthNd = manager.create(new float[]{validLength});
validLengthNd.setName("data2");
NDList list = new NDList(3);
list.add(indicesNd);
list.add(typesNd);
list.add(validLengthNd);
return list;
}
@Override
public String processOutput(TranslatorContext ctx, NDList list) {
NDArray array = list.singletonOrThrow();
NDList output = array.split(2, 2);
// Get the formatted logits result
NDArray startLogits = output.get(0).reshape(new Shape(1, -1));
NDArray endLogits = output.get(1).reshape(new Shape(1, -1));
// Get Probability distribution
NDArray startProb = startLogits.softmax(-1);
NDArray endProb = endLogits.softmax(-1);
int startIdx = (int) startProb.argMax(1).getFloat();
int endIdx = (int) endProb.argMax(1).getFloat();
return tokens.subList(startIdx, endIdx + 1).toString();
}
}
```
Congrats! You have created your first Translator! We have pre-filled the `processOutput()` that will process the `NDList` and return it in a desired format. `processInput()` and `processOutput()` offer the flexibility to get the predictions from the model in any format you desire.
With the Translator implemented, you need to bring up the predictor that uses your `Translator` to start making predictions. You can find the usage for `Predictor` in the [Predictor Javadoc](https://javadoc.djl.ai/api/0.2.0/ai/djl/inference/Predictor.html). Create a translator and use the `question` and `resourceDocument` provided previously.
```
String predictResult = null;
QAInput input = new QAInput(question, resourceDocument, 384);
BertTranslator translator = new BertTranslator(parser);
// Create a Predictor and use it to predict the output
try (Predictor<QAInput, String> predictor = model.newPredictor(translator)) {
predictResult = predictor.predict(input);
}
System.out.println(question);
System.out.println(predictResult);
```
Based on the input, the following result will be shown:
```
[december, 2004]
```
That's it!
You can try with more questions and answers. Here are the samples:
**Answer Material**
The Normans (Norman: Nourmands; French: Normands; Latin: Normanni) were the people who in the 10th and 11th centuries gave their name to Normandy, a region in France. They were descended from Norse ("Norman" comes from "Norseman") raiders and pirates from Denmark, Iceland and Norway who, under their leader Rollo, agreed to swear fealty to King Charles III of West Francia. Through generations of assimilation and mixing with the native Frankish and Roman-Gaulish populations, their descendants would gradually merge with the Carolingian-based cultures of West Francia. The distinct cultural and ethnic identity of the Normans emerged initially in the first half of the 10th century, and it continued to evolve over the succeeding centuries.
**Question**
Q: When were the Normans in Normandy?
A: 10th and 11th centuries
Q: In what country is Normandy located?
A: france
| github_jupyter |
# Run Train of Bubble-Agent (DQN)
- Team: TToBoT
- Member: { Sejun, Steve, Victor } @kaist
## Objective
- run training simultaneously w/ notebook
- to compare the performance of traing
## For Competition
1. prepare the final trained IQN Model (checkpoint w/ 100 iteration)
2. need to customize of env.step()
- it should work only 1 live. (later, we can use 3 lives)
- need to enumerate all stage (1~99) level w/ at least 10 (250,000 x 10) (8hr x 100 = 800/33d) iteration. (model should be same)
- using origin step(). do train w/ random level, do loop (iteration) forever! (final training)
3. in final competion, it will load the latest checkpoint for initial Model paramters.
4. win the competition!!
```
import os, sys, gin
# use parent folder as shared lib path..
if "../" not in sys.path:
sys.path.append("../")
# major libraries
import gin.tf
import seaborn as sns
import matplotlib.pyplot as plt
from absl import flags
import numpy as np
import tensorflow as tf
# show tf version.
print('! tf.ver = {}'.format(tf.__version__))
# Globals
# BASE_PATH = './!experimental_results_bubble/run3'
# let Dopamine .py files to be imported as modules in Jupiter notebook
module_path = os.path.abspath(os.path.join('../dopamine'))
if module_path not in sys.path:
sys.path.append(module_path)
print(module_path)
# try to load `Dopamine` libraries
import bubble
from dopamine.colab import utils as colab_utils
```
## Train Bubble w/ DQN
```
# @title Load the configuration for DQN.
# DQN_PATH = os.path.join(BASE_PATH, 'rainbow')
# Modified from dopamine/agents/dqn/config/dqn_cartpole.gin
# CONFIG FOR DQN (see @bubble/dqn_nature.gin)
gin_config = '''
# run_experiment
# python -um dopamine.discrete_domains.train --base_dir=/tmp/bubble --gin_files='bubble/dqn_nature.gin'
# python -um dopamine.discrete_domains.train --base_dir=/tmp/bubble --gin_files='bubble/dqn_nature.gin' --gin_bindings='DQNAgent.tf_device="/cpu:*"'
# Hyperparameters used in Mnih et al. (2015).
import dopamine.discrete_domains.atari_lib
import dopamine.discrete_domains.run_experiment
import dopamine.agents.dqn.dqn_agent
import dopamine.replay_memory.circular_replay_buffer
import gin.tf.external_configurables
import bubble.retro_lib
import bubble.bubble_agent
retro_lib.create_retro_environment.game_name = 'BubbleBobble'
retro_lib.create_retro_environment.level = 1
Runner.create_environment_fn = @retro_lib.create_retro_environment
create_agent.agent_name = 'dqn'
RetroPreprocessing.wall_offset = 0
DQNAgent.gamma = 0.99
DQNAgent.update_horizon = 1
DQNAgent.min_replay_history = 50000 # agent steps
DQNAgent.update_period = 4
DQNAgent.target_update_period = 10000 # agent steps
DQNAgent.epsilon_train = 0.1
DQNAgent.epsilon_eval = 0.05
DQNAgent.epsilon_decay_period = 1000000 # agent steps
DQNAgent.tf_device = '/gpu:1' # use '/cpu:*' for non-GPU version
DQNAgent.optimizer = @tf.train.RMSPropOptimizer()
tf.train.RMSPropOptimizer.learning_rate = 0.00025
tf.train.RMSPropOptimizer.decay = 0.95
tf.train.RMSPropOptimizer.momentum = 0.0
tf.train.RMSPropOptimizer.epsilon = 0.00001
tf.train.RMSPropOptimizer.centered = True
# atari_lib.create_atari_environment.game_name = 'Pong'
# Deterministic ALE version used in the DQN Nature paper (Mnih et al., 2015).
# atari_lib.create_atari_environment.sticky_actions = False
# create_agent.agent_name = 'dqn'
Runner.num_iterations = 200
Runner.training_steps = 250000 # agent steps
Runner.evaluation_steps = 125000 # agent steps
Runner.max_steps_per_episode = 27000 # agent steps
AtariPreprocessing.terminal_on_life_loss = True
WrappedReplayBuffer.replay_capacity = 1000000
WrappedReplayBuffer.batch_size = 32
'''
# parse this config
gin.parse_config(gin_config, skip_unknown=False)
# Train DQN on Cartpole
#dqn_runner = create_runner(DQN_PATH, schedule='continuous_train')
#print('\n\n\nStart Training...\n\n\n')
#dqn_runner.run_experiment()
#print('\n\n\nDone training\n\n\n')
#dqn4 (5/28) - reward := -0.01 + 1*K - 3*D + log(S,100) + 5*L
#dqn5 (6/02) - same reward, but wall_offset = 0
#dqn7 (6/04) - final reward
DQN_PATH = '/tmp/bubble_dqn7'
# import main run()
from dopamine.discrete_domains import run_experiment
# config main file
gin_files = []
# bindings.....
gin_bindings = ['Runner.evaluation_steps=0']
# # code from train.main()
# tf.logging.set_verbosity(tf.logging.INFO)
# run_experiment.load_gin_configs(gin_files, gin_bindings)
# runner = run_experiment.create_runner(DQN_PATH)
# # start run
# runner.run_experiment()
```
## Thread for updating status
```
# Thread for update canvas
import threading, time
def get_ioloop():
import IPython, zmq
ipython = IPython.get_ipython()
if ipython and hasattr(ipython, 'kernel'):
return zmq.eventloop.ioloop.IOLoop.instance()
# The IOloop is shared
ioloop = get_ioloop()
# Main Thread
class MyThread(threading.Thread):
'''Thread for drawing into canvas in live'''
def __init__(self, sleep = 0.5, name = 'my'):
super().__init__()
self._quit = threading.Event()
self.sleep = 0.5
self.name = name
self.start()
def run(self):
while not self._quit.isSet():
def update_progress():
if self._quit.isSet():
return
self.display()
time.sleep(self.sleep)
ioloop.add_callback(update_progress)
print("! T[{}].Quit()".format(self.name))
def quit(self):
self._quit.set()
def display(self):
pass
# display basic
from ipycanvas import Canvas
canvas = Canvas(width=640, height=480)
if canvas:
canvas.stroke_text('hello canvas! -------------', 0, 10)
# show canvas in here.
canvas
# Helper for Canvas
#canvas.fill_style = 'green'
#canvas.fill_rect(25, 25, 100, 100)
#canvas.clear_rect(45, 45, 60, 60)
def drawPlot2Canvas(fig = None, x=0, y=0):
'''draw current plt to canvas at (x,y)'''
fig = plt.gcf() if fig is None else fig
plt.close() # not to update on screen.
fig.canvas.draw() # draw fig to canvas
arr = np.array(fig.canvas.renderer._renderer)
print('! arr = {}'.format(np.shape(arr)))
h, w, d = np.shape(arr)
print('! w,h,d = {}'.format(w))
cv = Canvas(width=w, height=h)
cv.put_image_data(arr, 0, 0)
cv.stroke_rect(x, y, x+w-1, y+h-1)
canvas.clear_rect(x,y,x+w,y+h)
canvas.draw_image(cv, x, y)
def drawText2Canvas(txt='msg!', x=10, y=10):
w,h,o = 200,10,10
#canvas.fill_style = 'green'
#canvas.fill_rect(x, y-o, x+w, y+h-o)
canvas.clear_rect(x, y-o, x+w, y+h-o)
canvas.stroke_text(txt, x, y)
# draw plot....
fig = plt.figure(1)
plt.plot([[1,3],[3,3],[7,1]])
# draw plot-to-canvas
drawPlot2Canvas(fig, x=0)
drawText2Canvas('hello world')
#drawText2Canvas('......................')
```
### support Multi-Processing
```
from multiprocessing import Process, Queue
# process list
proc_list = []
proc_queue = None
# train function
def processTrain(name = 'train', Q = None):
global gin_files, gin_bindings, DQN_PATH
from dopamine.discrete_domains import run_experiment
Q.put('init!') if Q else None
tf.logging.set_verbosity(tf.logging.INFO)
run_experiment.load_gin_configs(gin_files, gin_bindings)
runner = run_experiment.create_runner(DQN_PATH)
# access to env
env = runner._environment
o = env.reset()
Q.put('! o({}) = {}'.format(type(o), o[0:10,0,]))
Q.put('start!') if Q else None
runner.run_experiment()
Q.put('! P[{}].stop()'.format(name))
# train thread
def startProcessTrain(target = None):
global proc_queue, proc_list
target = target if target is not None else processTrain
proc_queue = Queue() if proc_queue is None else proc_queue
proc = Process(target = target, args = ('T0', proc_queue))
proc_list.append(proc)
proc.start()
return proc
# stop(or kill) processes
def stopProcess():
global proc_list
for proc in proc_list:
proc.terminate()
proc.join()
# trainer = startProcessTrain()
# stop
# stopProcess()
# show process
# !ps -ax | grep python
# proc_queue
```
### MyTrainer and MyThread
```
from dopamine.discrete_domains import run_experiment
# MyRunner for Train
# - report every episode status.
class MyRunner(run_experiment.Runner):
def __init__(self, base_dir, create_agent_fn):
'''initialize runner'''
super(MyRunner, self).__init__(base_dir, create_agent_fn)
self._load_logger()
def _run_one_episode(self):
'''override to post episode status'''
global proc_queue
episode_length, episode_return = super(MyRunner, self)._run_one_episode()
data = {'episode':{'length': episode_length, 'return': episode_return }}
#proc_queue.put('! epsode[len,ret] = {},{}'.format(episode_length, episode_return))
proc_queue.put(data)
return episode_length, episode_return
def _load_logger(self):
'''load logger to save into file'''
import logging, os
# get TF logger
log = logging.getLogger('tensorflow')
log.setLevel(logging.DEBUG)
# create file handler which logs even debug messages
formatter = logging.Formatter('%(asctime)s - %(name)s - %(levelname)s - %(message)s')
fh = logging.FileHandler(os.path.join(DQN_PATH, 'tensorflow.log'))
fh.setLevel(logging.INFO)
fh.setFormatter(formatter)
log.addHandler(fh)
#! start runner
def startMyRunner(name = 'train', Q = None):
global gin_files, gin_bindings, DQN_PATH
from dopamine.discrete_domains import run_experiment
Q.put('! start: my-runner') if Q else None
tf.logging.set_verbosity(tf.logging.INFO)
run_experiment.load_gin_configs(gin_files, gin_bindings)
runner = MyRunner(DQN_PATH, run_experiment.create_agent)
runner.run_experiment()
Q.put('! P[{}].stop()'.format(name)) if Q else None
#! start process of runner
startProcessTrain(target = startMyRunner)
```
#### Train Results (01/Jun/2020)
```pre
INFO:tensorflow:Starting iteration 87
INFO:tensorflow:Average undiscounted return per training episode: 19.29
INFO:tensorflow:Average training steps per second: 98.36
INFO:tensorflow:Starting iteration 88
INFO:tensorflow:Average undiscounted return per training episode: 17.34
INFO:tensorflow:Starting iteration 89
INFO:tensorflow:Average undiscounted return per training episode: 18.19
INFO:tensorflow:Starting iteration 90
INFO:tensorflow:Average undiscounted return per training episode: 16.46
INFO:tensorflow:Starting iteration 91
INFO:tensorflow:Average undiscounted return per training episode: 18.53
INFO:tensorflow:Starting iteration 92
INFO:tensorflow:Average undiscounted return per training episode: 18.22
INFO:tensorflow:Starting iteration 99
INFO:tensorflow:Average undiscounted return per training episode: 17.893
INFO:tensorflow:Starting iteration 100
INFO:tensorflow:Average undiscounted return per training episode: 18.24
INFO:tensorflow:Starting iteration 101
INFO:tensorflow:Average undiscounted return per training episode: 19.01
INFO:tensorflow:Starting iteration 102
INFO:tensorflow:Average undiscounted return per training episode: 19.94
INFO:tensorflow:Starting iteration 103
INFO:tensorflow:Average undiscounted return per training episode: 17.44
INFO:tensorflow:Starting iteration 104
INFO:tensorflow:Average undiscounted return per training episode: 17.876
INFO:tensorflow:Starting iteration 105
INFO:tensorflow:Average undiscounted return per training episode: 17.42
INFO:tensorflow:Starting iteration 106
INFO:tensorflow:Average undiscounted return per training episode: 17.595
INFO:tensorflow:Starting iteration 107
INFO:tensorflow:Average undiscounted return per training episode: 17.779
```
```
# MyThread for status display
class MyTrainStatus(MyThread):
def __init__(self):
super().__init__(name='status')
self.episodes = np.array([[0,0]])
print('! MyTrainStatus({})'.format(self.name))
def display(self):
global canvas, proc_queue, plt
episodes = []
# pop all queue...
while not proc_queue.empty():
msg = proc_queue.get()
if msg and 'episode' in msg:
E = msg['episode']
episodes.append([E['length'], E['return']])
# print('>> episodes = {}'.format(episodes))
# draw plot if len > 0
if len(episodes) > 0:
arr = np.array(episodes)
print('>> arr = {}'.format(arr))
# draw plot...
if 1>0:
self.episodes = np.vstack((self.episodes, arr))
#print('>> self.episodes = {}'.format(self.episodes))
#fig = plt.figure(1)
#plt.plot(self.episodes)
fig, ax1 = plt.subplots()
ax2 = ax1.twinx()
ax1.plot(self.episodes[:,0], 'g-')
ax2.plot(self.episodes[:,1], 'b-')
ax1.set_xlabel('episode count')
ax1.set_ylabel('length', color='g')
ax2.set_ylabel('return', color='b')
drawPlot2Canvas(fig)
#! start thread for status
tstatus = MyTrainStatus()
episode_length, episode_return = 1,3
msg = {'episode':{'length': episode_length, 'return': episode_return }}
proc_queue.put(msg)
print('> msg.org = {}'.format(msg))
# stop - thread of status
tstatus.quit() if tstatus else None
# stop - process of train
stopProcess()
```
| github_jupyter |
```
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
from keras.models import Sequential
from keras.layers import Dense, Flatten, LSTM, Conv1D, MaxPooling1D, Dropout, Activation
from keras.layers.embeddings import Embedding
import nltk
import time
import string
import numpy as np
import pandas as pd
import html,re
from nltk.corpus import stopwords
from nltk import word_tokenize
from nltk.stem.wordnet import WordNetLemmatizer
from nltk.stem.porter import *
from sklearn.model_selection import train_test_split
import plotly.offline as py
import plotly.graph_objs as go
py.init_notebook_mode(connected=True)
from google.colab import drive
drive.mount('/content/drive/')
nltk.download('stopwords')
stopwords = nltk.corpus.stopwords.words("english")
stopwords.extend('rt')
#Sorting out the comments
TwComments = pd.read_csv ('/content/drive/My Drive/Extras/tweetsbitcoin.csv',delimiter=",", index_col=None)
TwComments = TwComments.dropna()
TwComments=TwComments.drop_duplicates()
TwComments=TwComments.sort_values(['date','time'], ascending=[True,True])
i = TwComments[(TwComments.date == '2017-09-30')].index
TwComments = TwComments.drop(i)
TwComments["DateTime"]=pd.to_datetime(TwComments['date'] + ' ' + TwComments['time'])
TwComments["hour"] = TwComments.DateTime.dt.hour
TwComments["day"] = TwComments.DateTime.dt.weekday_name
TwComments["DateTime"] = TwComments.DateTime.values.astype(np.int64) // 10 ** 9
TwComments["TimeDiff"]= TwComments["DateTime"] - (TwComments["DateTime"] % 86400)
TwComments = TwComments[TwComments.TimeDiff > 0]
startTime = int(round(time.time()*60))
tcomm=pd.DataFrame()
grouped_terms = TwComments.groupby(["TimeDiff"])
#This is done to combine the tweets in that hour into a list
i = 0
tweets = []
for name, group in grouped_terms:
t = []
for row, data in group.iterrows():
t.append(data['tweet'])
tweets.append(''.join(t))
Price = pd.read_csv ('/content/drive/My Drive/Extras/1Daybinanceprices.csv',delimiter="\t", index_col=None)
Price["Date"] = pd.to_datetime(Price['ClTime'], unit = 'ms')
Price = Price.sort_values(['OpTime'], ascending=True)
def preprocess(tweets):
t = html.unescape(tweets)
z = lambda x: re.compile('\#').sub('', re.compile('RT @').sub('@', x).strip())
t = z(t)
tweet = ' '.join(re.sub("(@[_A-Za-z0-9]+)|([^0-9A-Za-z \t])|(\w+:\/\/\S+)"," ",t).split())
return tweet
#Tokenizing the tweets
def tokenizer(tweets):
tweet = " ".join(re.split("[^a-zA-Z.,!?]*", tweets.lower())).strip()
stemmer = PorterStemmer()
tweets = [stemmer.stem(tweet.lower()) for tweet in tweets.split()]
return tweets
def basic_tokenizer(tweet):
#Same as tokenize but without the stemmer
tweet = " ".join(re.split("[^a-zA-Z.,!?]*", tweet.lower())).strip()
return tweet.split()
for i,t in enumerate(tweets):
tweets[i] = tokenizer(preprocess(t))
vocabulary_size = 1000000
tokenizer = Tokenizer(num_words= vocabulary_size)
tokenizer.fit_on_texts(tweets)
sequences = tokenizer.texts_to_sequences(tweets)
data = pad_sequences(sequences, maxlen=15000)
Price['PriceDiff'] = Price['Close']-Price['Open']
price_diff = []
for p in Price['PriceDiff']:
if p >= 0:
price_diff.append(1)
else:
price_diff.append(0)
X_train, X_test, y_train, y_test = train_test_split(data, price_diff, test_size=0.1, shuffle=False)
model = Sequential()
model.add(Embedding(1000000, 100, input_length=15000))
model.add(LSTM(100, dropout=0.2, recurrent_dropout=0.2))
model.add(Dense(1, activation='sigmoid'))
model.compile(loss='mean_squared_error', optimizer='adam', metrics=['accuracy'])
model.fit(X_train, y_train, epochs=10)
y = model.predict(X_test)
t = y
for i,z in enumerate(y):
if z>0.4:
t[i]=1
else:
t[i]=0
t = list(map(int, t))
print(t)
su = 0
print(23/37)
print(y_test)
```
| github_jupyter |
```
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
```
`mushi` demo
==
This notebook demonstrates how to use the Python module [`mushi`](https://github.com/harrispopgen/mushi/blob/master/mushi.py) for...
## Inferring mutation spectrum history (and demography)
>*the thing I came for:*\
*the wreck and not the story of the wreck*\
*the thing itself and not the myth*
>
> – Adrienne Rich, Diving Into the Wreck
### Canon ball metaphor stolen from Erick Matsen

In this metaphor, the pile of canonballs represents the mutation spectra we compute from SNPs, and the canon represents the mutational process.
Just as the history of the canon's firing rate and direction explains where we find the piles, the history of the mutation process explains the SNP mutation spectra we find in modern genomes.
We will use `mushi` to infer history of the mutation process, which we can think of as the mutation rate function over time for each triplet mutation type.
In `mushi`, we use coalescent theory and optimization techniques to learn about this history from the $k$-SFS.
### $3$-SFS from the 1000 Genomes Finnish population (previously computed with [`mutyper ksfs`](https://github.com/harrispopgen/mutyper))
Load the $k$-SFS
```
from mushi.ksfs import kSFS
ksfs = kSFS(file='../example_data/3-SFS.EUR.FIN.tsv')
```
Plot the population variant spectrum (summing the $k$-SFS over sample frequency)
```
ksfs.as_df().sum(0).plot.bar(figsize=(17, 3))
plt.xticks(family='monospace')
plt.ylabel('number of variants')
plt.show()
```
Plot the total SFS (summing the $k$-SFS over mutation types)
```
ksfs.plot_total()
plt.yscale('log')
```
plot k-SFS composition as a scatter (a color for each mutation type)
```
ksfs.plot(clr=True)
plt.show()
```
...and as a heatmap (a column for each mutation type)
```
g = ksfs.clustermap(figsize=(17, 7), col_cluster=False, xticklabels=True, cmap='RdBu_r', rasterized=True, robust=True)
g.ax_heatmap.set_xticklabels(g.ax_heatmap.get_xmajorticklabels(), fontsize = 9, family='monospace')
plt.show()
```
We will also need the masked genome size for each mutation type, which we've also previously computed with `mutyper targets`. This defines mutational target sizes.
```
masked_genome_size = pd.read_csv(f'../example_data/masked_size.tsv', sep='\t', header=None, index_col=0)
masked_genome_size.index.name='mutation type'
masked_genome_size.plot.bar(figsize=(6, 3), legend=False)
plt.xticks(family='monospace')
plt.ylabel('mutational target size (sites)')
plt.show()
```
With this we can compute the number of SNPs per target in each mutation type. Notice the enrichment of C>T transitions at CpG sites.
```
normalized_hit_rates = ksfs.as_df().sum(0).to_frame(name='variant count')
normalized_hit_rates['target size'] = [int(masked_genome_size.loc[context])
for context, _ in normalized_hit_rates['variant count'].index.str.split('>')]
(normalized_hit_rates['variant count'] /
normalized_hit_rates['target size']).plot.bar(figsize=(17, 3), legend=False)
plt.xticks(family='monospace')
plt.ylabel('variants per target')
plt.show()
```
To compute the total mutation rate in units of mutations per masked genome per generation, we multiply an estimate of the site-wise rate by the target size
```
μ0 = 1.25e-8 * masked_genome_size[1].sum()
μ0
```
To render time in years rather than generations, we use an estimate of the generation time
```
t_gen = 29
```
### Joint coalescent inference of demography and mutation spectrum history
To access time-calibrated mutation spectrum histories, we first need to estimate the demographic history, since this defines the diffusion timescale of the coalescent process.
We first define a grid of times will represent history on, measured retrospectively from the present in units of Wright-Fisher generations.
```
t = np.logspace(np.log10(1), np.log10(200000), 200)
```
We now run the optimization, setting a few parameters to control how complicated we let the histories look.
```
ksfs.infer_history(t, μ0, alpha_tv=1e2, alpha_spline=3e3, alpha_ridge=1e-10,
beta_rank=1e1, beta_tv=7e1, beta_spline=1e1, beta_ridge=1e-10,
tol=1e-11)
```
Hopefully you agree that was fast 🏎
We'll now check that the demography has a few features we expect in the Finnish population: the out-of-Africa bottleneck shared by all Eurasians, a later bottleneck associated with northward migration, and exponential population growth toward the present.
- The plot on the left will show fit to the SFS
- The plot on the right will show the inferred haploid effective population size history.
```
plt.figure(figsize=(10, 5))
plt.subplot(121)
ksfs.plot_total()
plt.yscale('log')
plt.subplot(122)
ksfs.eta.plot(t_gen=t_gen)
plt.xlim([1e3, 1e6])
plt.show()
```
Now let's take a look at the inferred mutation spectrum history (MuSH).
- The plot on the left will show the measured $k$-SFS composition (points) and the fit from `mushi` (lines)
- The plot on the right will show the inferred MuSH
```
plt.figure(figsize=(16, 5))
plt.subplot(121)
ksfs.plot(clr=True)
plt.subplot(122)
ksfs.μ.plot(t_gen=t_gen, clr=True, alpha=0.75)
ksfs.μ.plot(('TCC>TTC',), t_gen=t_gen, clr=True, lw=5)
plt.xscale('log')
plt.xlim([1e3, 1e6])
plt.show()
```
We can also plot the MuSH as a heatmap with the y axis representing time.
```
g = ksfs.μ.clustermap(t_gen=t_gen, figsize=(17, 7), col_cluster=True, xticklabels=True, robust=False, cmap='RdBu_r')
g.ax_heatmap.set_xticklabels(g.ax_heatmap.get_xmajorticklabels(), fontsize = 9, family='monospace')
g.ax_heatmap.set_ylim([172, 58])
plt.show()
```
Now that you have a MuSH, you can start answering questions about mutation spectrum history!🤸
| github_jupyter |
```
import sys, os; sys.path.append('..')
import pyzx as zx
import random
import math
from fractions import Fraction
%config InlineBackend.figure_format = 'svg'
c = zx.qasm("""
qreg q[3];
cx q[0], q[1];
""")
zx.d3.draw(c)
c = zx.qasm("""
qreg q[2];
rx(0.5*pi) q[1];
t q[0];
cx q[0], q[1];
cx q[1], q[0];
cx q[0], q[1];
tdg q[1];
rx(-0.5*pi) q[0];
""")
zx.d3.draw(c)
c.gates
g = c.to_graph()
g
zx.d3.draw(g)
zx.simplify.spider_simp(g)
zx.d3.draw(g)
zx.full_reduce(g)
zx.d3.draw(g)
g = zx.sqasm("""
qreg S[1];
qreg q[3];
t S[0];
cx q[0], S[0];
cx q[1], S[0];
cx q[2], S[0];
""")
zx.d3.draw(g)
g = zx.sqasm("""
qreg S[2];
qreg q[3];
t S[0];
cx q[0], S[0];
cx q[1], S[0];
cx q[2], S[0];
tdg S[1];
cx q[0], S[1];
cx q[1], S[1];
cx q[2], S[1];
""")
zx.d3.draw(g)
zx.clifford_simp(g)
zx.d3.draw(g)
zx.full_reduce(g)
zx.d3.draw(g)
g = zx.Circuit.load("test.qsim").to_graph()
zx.d3.draw(g)
g1 = g.copy()
g1.map_qubits([
(0,0), (1, 0), (2, 0), (3, 0),
(0,1), (1, 1), (2, 1), (3, 1),
(0,2), (1, 2), (2, 2), (3, 2),
(0,3), (1, 3), (2, 3), (3, 3)
])
zx.d3.draw(g1)
zx.full_reduce(g1)
zx.d3.draw(g1)
def t_optimiser(c):
g = c.to_graph()
g = zx.simplify.teleport_reduce(g)
c_opt = zx.Circuit.from_graph(g).split_phase_gates().to_basic_gates()
return zx.optimize.basic_optimization(c_opt).to_basic_gates()
c = zx.Circuit.load('../circuits/Fast/grover_5.qc')
zx.d3.draw(c.to_graph())
print(zx.tcount(c))
c1 = t_optimiser(c)
zx.d3.draw(c1.to_graph())
print(zx.tcount(c1))
c.verify_equality(c1)
c2 = c1.copy()
c2.add_circuit(c.adjoint())
g = c2.to_graph()
zx.simplify.full_reduce(g)
zx.d3.draw(g)
c1.gates[10] = zx.gates.T(6, adjoint=True)
c.verify_equality(c1)
c2 = c1.copy()
c2.add_circuit(c.adjoint())
g = c2.to_graph()
zx.simplify.full_reduce(g)
zx.d3.draw(g)
g = zx.Circuit.load('../circuits/Fast/hwb6.qc').to_graph()
zx.d3.draw(g)
print(zx.tcount(g))
zx.simplify.full_reduce(g)
zx.d3.draw(g)
print(zx.tcount(g))
g.apply_state("++---+-")
g.apply_effect("+011-1-")
zx.simplify.full_reduce(g)
print(zx.tcount(g))
zx.drawing.arrange_scalar_diagram(g)
zx.d3.draw(g)
def compute_decomp(g):
if zx.tcount(g) >= 6:
gsum = zx.simulate.replace_magic_states(g)
gsum.reduce_scalar()
terms = 0
vals = 0
for g1 in gsum.graphs:
t,v = compute_decomp(g1)
terms += t
vals += v
return (terms, vals)
else:
return (2 ** math.ceil(zx.tcount(g)/2), g.to_matrix())
math.ceil(2**(0.468 * zx.tcount(g)))
compute_decomp(g)
zx.simulate.calculate_path_sum(g)
```
| github_jupyter |
# Aqua 0.7 Operator Redesign
_17-Jan-19, donny@_
| **Status** | **Accepted** |
|:------------------|:----------------------------------------------|
| **RFC #** | 0003 |
| **Authors** | Donny Greenberg (donny@ibm.com) |
| **Deprecates** | NA |
| **Submitted** | 2020-01-17 |
| **Updated** | 2020-01-23 |
## Purpose
To improve the transparency, ease of understanding, and programming power of Aqua’s operator logic and usage. Specifically, to reconcile with the Terra operator hierarchy and make the Aqua algorithmic flow more visible, explicit, and extensible.
Throughout this doc, we rely on definitions of Operators roughly derived from the first chapter of John Watrous's "The Theory of Quantum Information," with a focus on Square Operators over binary alphabets.
## Background: Motivation and Opportunities
The representation of matrices sparsely as linear combinations of Pauli operators is critical in many quantum algorithms. As such, the Operator classes are the workhorses of Aqua today (0.6.2), containing both the expectation value and evolution logic used by most of its algorithms.
However, there are several opportunities for improvement:
* **Basic Construction & Rapid Protoyping:** Aqua's Operators were initially built as procedural infrastructure rather than first-class programming primitives. Improvements to syntax and interfaces can enable the succinctness and power typical of mathematical Operator language
* **Separation of Operator Math and Operator Algorithms**
* Ease of understanding: The "Operator algorithm" logic - the ExpectationValue, Evolution, grouping, and symmetry analysis - is mostly spread across the 3000-line operator hierarchy, and is very branchy for different modes of execution
* Ease of extension: Modification to the expectation value, evolution, grouping, and symmetry logic is a core use case (e.g. the [CVaR expectation](https://arxiv.org/abs/1907.04769), [linear combination evolution](https://arxiv.org/abs/1202.5822), or the many recent papers on [Pauli grouping](https://www.nature.com/articles/nature23879)), but not explicitly supported today
* **Smooth Borders with Broader Qiskit**
* Terra's `quantum_info` module also supports operator math, but is mostly matrix-based
* **Remote Operator Algorithms:** Aer's fast ExpectationValue is not transparently or cleanly interchangeable with Aqua's local ExpectationValue today. The concept of an Algorithm not provided by Aqua is not yet defined to support this type of interchangeability cleanly
### Present State of Operators in Qiskit
Both Aqua and Terra include suites of modules to support Operator math, but do so very differently.
* Aqua
* Operators are focused primarily on the procedural requirements of algorithmic execution
* Modules are very large and include hundreds of lines of procedural algorithm code
* Interfaces were not initial built for end-user usage as a programming primitive, and are therefore wordy and difficult for users to understand
* Syntax is not built for rapid prototyping and lacks syntactic power of mathematical Operator language
* Primarily focused on Pauli-basis Operators
* WeightedPauli - $2^n\times 2^n$ Operators sparsely represented as complex combination of Paulis
* MatrixOperator in the standard basis with $2^n\times 2^n$ elements was initially built for performance improvements which are no longer relevant
* Only dependency on Terra is through Pauli module, but this is largely symbolic (not an inexorable component)
* Terra
* Operator math is mostly built around QCVV (Quantum Characterization Verification & Validation) and open Quantum systems modelling use cases
* Support for Channel, Choi, Superoperator, Kraus, etc.
* Operators are largely matrix-based and therefore do not support the Pauli-basis operations necessary to non-exponentially execute quantum algorithms
* Used by:
* Aqua, 29 dependencies - Only Pauli module
* Aer, 10 dependencies
* Ignis, 2 dependencies
* Ignis includes a `clifford.py` module somewhat specific to characterization needs.
### Aqua Present Usage (0.6.2)
Within Aqua, the primary uses of Operators are:
* Qubit Observable (Hamiltonian, Cost Function, etc.) Construction
* Used as sparse representations of large observables when constructing problems in Chemistry, Physics, Optimization, and Finance today
* Also often a translation step between domain-specific problems and Quantum hardware-addressable equivalents
* ExpectationValues
* Primarily used in VQE (and derivatives QAOA, UCCSD, etc.) as a device-executable cost function of the ansatz state
* Expectation values can only be taken of Operators in the Pauli basis on Quantum hardware
* Also present in the "Evolution of Hamiltonian" algorithm, which is simply state evolution by one operator followed by an expectation value by another operator
* State Evolution
* Used in QPE (and derivatives HHL, iQPE, etc.) as a Quantum circuit-representable matrix exponentiation
* Used in UCCSD and QAOA ansatze and EOH algorithm as representation of system dynamics to simulate time evolution of a system on quantum hardware
* Evolution can only be taken by Operators in the Pauli basis on Quantum hardware
#### Other Important Aqua Operator Features
* __Grouping__ - Grouping is a technique to reduce the number of circuit evaluations required to compute an ExpectationValue based on mutually commuting Paulis in the Operator decomposition.
* __Tapering__ - Tapering is a technique to remove qubits from a Hamiltonian of interest by identifying Z2 symmetries in the Hamiltonian.
* __Gradients__ - Many variational algorithms are improved dramatically when exact gradients of gate parameters with respect to the cost function observable are computed analytically rather than numerically. Aqua can compute these gradients and provide them to the optimizer directly.
### Aqua Present (0.6.2) Operator Object Model and Hierarchy
Aqua's Operators are organized as follows:
* `qiskit.aqua.operators`
* base_operator.py: `BaseOperator(ABC)`
* matrix_operator.py: `MatrixOperator(BaseOperator)`
* weighted_pauli_operator.py: `WeightedPauliOperator(BaseOperator)`, __and__ `Z2Symmetries`
* tpb_grouped_weighted_pauli_operator.py: `TPBGroupedWeightedPauliOperator(WeightedPauliOperator)`, essentially a wrapper around `WeightedPauliOperator` for backward compatibility.
* pauli_graph: `PauliGraph`
* op_converter.py: `to_weighted_pauli_operator(operator)`, `to_matrix_operator(operator)`, `to_tpb_grouped_weighted_pauli_operator(operator, grouping_func, **kwargs)`
* common.py: Utility functions, inc. `evolution_instruction`, `pauli_measurement(circuit, pauli, qr, cr, barrier=False)`, `measure_pauli_z(data, pauli)`, `covariance(data, pauli_1, pauli_2, avg_1, avg_2)`, etc.
* `qiskit.chemistry` __- OUT OF SCOPE OF THIS DOC__
* fermionic_operator.py: `FermionicOperator`, contains `jordan_wigner`, `parity`, `bravyi_kitaev` Fermion-to-qubit operator mappings.
* bksf.py: Another mapping
* `.core`
* chemistry_operator.py: `ChemistryOperator(ABC)`
* hamiltonian.py: `Hamiltonian(ChemistryOperator)`
### Terra Present (0.11.0) Operator Object Model and Hierarchy
Terra's Operators are organized as follows:
* `qiskit.quantum_info`
* `.operators`
* base_operator.py, pauli.py, operator.py (matrix operator), measures.py (`process_fidelity`), predicates.py (`is_unitary_matrix`, `is_hermitian_matrix`, `matrix_equal`, etc.), quaternion.py
* `.channel`
* quantum_channel.py (base), chi.py, choi.py, kraus.py, ptm.py, stinespring.py, superop.py, transformations.py
* `.states`
* quantum_state.py (base), densitymatrix.py, statevector.py, measures.py (`state_fidelity`), states.py (`basis_state`, `projector`, `purity`)
* `.analysis`
* average.py - ExpectationValue of diagonal operator
* make_observable.py - Convert an observable in matrix form to dictionary form
#### WeightedPauliOperator Not Available in Terra
Terra does not contain any of the logic for working in the Pauli-basis implemented in Aqua today, and is not interoptable with Aqua's operator algorithms. As such, these utilities are only accessible to Aqua users.
### Operator Construction and Manipulation Present State
The center of Qiskit's algorithmic Operator logic is the WeightedPauli, being the only non-exponential scaling operator basis available today (the only other being the standard basis).
Qiskit supports several methods of WeightedPauli operator construction, none of which are self explanatory to a new user:
```
# from qiskit.quantum_info.operators import WeightedPauliOperator
from qiskit.aqua.operators import WeightedPauliOperator, MatrixOperator, op_converter
from qiskit.quantum_info.operators import Pauli
pauli_op = WeightedPauliOperator([
[.5, Pauli.from_label('IX')],
[.2, Pauli.from_label('ZY')],
[.1j, Pauli.from_label('ZZ')],
])
pauli_op = WeightedPauliOperator.from_list(
paulis=[Pauli.from_label('IX'),
Pauli.from_label('ZY'),
Pauli.from_label('ZZ')],
weights=[.5, .2, .1j])
mat = [[0. +0.1j, 0.5-0.2j, 0. +0.j , 0. +0.j ],
[0.5+0.2j, 0. -0.1j, 0. +0.j , 0. +0.j ],
[0. +0.j , 0. +0.j , 0. -0.1j, 0.5+0.2j],
[0. +0.j , 0. +0.j , 0.5-0.2j, 0. +0.1j]]
mat_op = MatrixOperator(mat)
pauli_op_from_mat = op_converter.to_weighted_pauli_operator(mat_op)
pauli_op == pauli_op_from_mat
```
Classical matrices can be exported for classical usage, again if the user already knows the Operator hierarchy somewhat well:
```
op_converter.to_matrix_operator(pauli_op).matrix.toarray()
```
Composition uses the `*` operator, while Terra's operators and Python use `@`.
```
3*pauli_op + .2j*pauli_op == (3+.2j)*pauli_op
print((pauli_op * pauli_op).print_details())
```
### Aqua's ExpectationValue is Procedural and Inextensible
Aqua's ExpectationValue is not contained within a single function or module, but rather split into several functions without a clear interface or flow for user usage. This is due to structural constraints in Aqua which are no longer present, where the algorithm requiring the expectation value held the backend object and could run circuits, but the operator could not. We encourage the reader to scan lines [361-395 of Aqua 6.1 VQE’s](https://github.com/Qiskit/qiskit-aqua/blob/stable/qiskit/aqua/algorithms/adaptive/vqe/vqe.py#L361) ExpectationValue calculation to try to understand where and how the expectation is computed. We’ve been asked by numerous Aqua users to explain how this code works, and most do not attempt to use it on their own.
The following is the shortest possible way to write an expectation value in Aqua. Note that it fundamentally requires the user to understand a certain execution flow, the correct functions to use to do this, and how those functions work with their execution mode. This takes a few hours to understand at least, often days. Further, there are no hints that a change from the Z basis for each Pauli is being performed here, or matrix multiplication if the system chooses to do that instead.
```
from qiskit.aqua.operators import WeightedPauliOperator
from qiskit.aqua.components.variational_forms import RY
from qiskit.quantum_info import Pauli
from qiskit import BasicAer, execute, QuantumCircuit
from qiskit.circuit import Parameter
qasm_sim = BasicAer.get_backend('qasm_simulator')
op = WeightedPauliOperator([
[.5, Pauli.from_label('IX')],
[.2j, Pauli.from_label('ZY')],
])
circuit = QuantumCircuit(2)
circuit.h([0,1])
evaluation_circuits = op.construct_evaluation_circuit(wave_function=circuit, statevector_mode=False)
result = execute(evaluation_circuits, qasm_sim).result()
expect, std = op.evaluate_with_result(result=result, statevector_mode=False)
expect
```
#### Alternative Expectation Values and the Aer Expectation Value
Because the ExpectationValue logic is embedded directly in the Operator, modifications to the ExpectationValue (e.g. CVaR) are impossible without editing the Operator directly with heavy branching or duplicating the entire Operator. This branching is already in effect within Aqua, automatically choosing between several execution modes mostly opaquely to the user. This is also the case for grouping, evolution, and symmetry logic.
The most dramatic example of this is the Aer-provided fast ExpectationValue simulation, which is so buried into the Operator it is effectively a super-superuser feature today. It was introduced quickly to achieve critical performance gains, but must be formalized to become a true first-class feature.
* In Aqua, there is no simple way to specify which ExpectationValue algorithm the user wants, Aer or otherwise, and most users do not know that the Aer Expectation Exists
* Aer's ExpectationValue is woven throughout the core operator code in a way that is branchy, inexorable, and difficult for users to understand and control
* A new ExpectationValue, such as one provided by BasicAer or IBMQProvider, would simply introduce additional branches following the existing style
### Aqua's State Evolution is Inextensible and Difficult to Navigate
Evolution is somewhat more succinct, but more difficult to navigate in code. The logic for evolution is distributed over several branchy static modules, and the evolution is pre-compiled as a CNOT-chain circuit, which is often not the ideal evaluation format (e.g. matrix multiplication if simulating, or Swap Networks).
```
from qiskit.circuit import Parameter
op = WeightedPauliOperator([
[.5, Pauli.from_label('IX')],
[.2, Pauli.from_label('ZY')],
])
circuit = QuantumCircuit(2)
θ = Parameter('θ')
instr = op.evolve_instruction(evo_time=θ)
circuit.append(instr, [0,1])
print(circuit.draw(fold=4000))
print('Decomposed:')
circuit.decompose().draw(fold=4000)
```
## Requirements and Design
1. Location and Ownership
1. Operators
1. Provider-specific Algorithms
1. Object Model
1. Operator Definition - Primitives and Composites
1. Algorithms Definition - Primitives and Composite Operations
1. Parameterization and Eagerness
1. Changes to Terra
1. Changes to Aqua
1. Algorithms as composite Operations
1. Circuit Execution Algorithms
1. Expectation Algorithms
1. Evolution Algorithms
1. Other Primitive Algorithms
### Location and Ownership in Qiskit
Given the presence of Operator logic in both Aqua and Terra, there are several options for their placement within Qiskit. The primary considerations here relate to which master branch tests them, who owns what in the case of breakage, and who owns what in the case of design.
In addition, some remote Operator algorithms are being discussed, with one already in production - the Aer Expectation Value. The location of these algorithms is also an important question.
#### Operator Location Considerations
* The Operator's centrality to Aqua means relying on an external library is a big overhead
* Reliance on Terra has created frequent firedrills because behavior and interfaces change without integration testing
* Firedrills are very difficult to troubleshoot because presently there is no integration testing between Terra and Aqua or design review to check whether a change will have downstream implications
* Operator is so central to Aqua that it will require strong ownership by the Aqua team, constant maintenance and changes
* Centralized Operator primitives can simplify interfaces across Qiskit
* By accepting a common Operator format derived from Terra, methods in different areas of Qiskit can communicate in a consistent format without dependencies
* For example, Aer's expectation value can take a circuit and an Operator, rather than depend on Aqua to define its interface, or rely on an informal interface (e.g. lists) which must be validated
* Terra and Aqua's respective Operators can be delineated somewhat cleanly
* Aqua and Terra's operators are seemingly used by completely different users for very different tasks (QA&A vs. QCVV or circuit analysis)
* Terra's Operators are primarily matrix-based, while Aqua's are primarily composites of sparse representations (e.g. sums of Paulis or Circuits)
* Though some are definitely shared, such as Pauli
* Operators and Gates may need to be reconciled at some point
* The X, Y, and Z Paulis are not different from the X, Y, and Z Gates
* Both the gate and operator models include functionality for converting unitary matrices to circuit operations
#### Operator Location Options
**A.** Move Aqua Operators into Terra, with:
1. Joint ownership by Aqua team
2. Aqua integration tests run on Terra's master branch (e.g. pulling in Aqua's master branch to execute tests). _Unit tests alone are not sufficient, as they are usually modified along with breaking changes to pass._
3. Aligned release cycles so Aqua does not need to scramble to release when Terra does
**Big-A.** Combine Aqua and Terra into a single repo and jointly own Operators
**B.** Move all operators and states into Aqua, jointly owned by Terra team
**C.** Leave Operators split between Aqua and Terra, with dependency on Terra for some primitives (QuantumCircuit, Pauli), with joint ownership and Aqua integration testing
##### **Decision:** Following a discussion in Aqua Design Review, option **A** will be pursued for the remainder of this doc.
#### Provider-Specific Algorithm Location Options (Decision)
**A.** Remote algorithms live in provider repo, and are tested and released at provider’s discretion
**B.** Remote algorithms live in Aqua, with Aqua integration testing of functionality in provider repo
**C.** Remote algorithms live in Aqua, with agreed upon interface to enforce consistency, and data interchange (e.g. an Operator format defined in Terra) tested in provider repo
### Object Model and Hierarchy
What is an Operator _to a QA&A (Quantum Algorithms & Applications) programmer?_
Ignoring the Physical definition of an Operator for a moment, as a _Quantum programming primitive,_ the Operator is:
* __Recursively defined__ - Operators can be one of several _primitives_ - e.g. Matrix, Pauli, Clifford, QuantumCircuit, or an arbitrary combination of these primitives, e.g. Addition, Tensor, Composition.
* It makes complete mathematical sense to add two primitives together, e.g. `(my_matrix+my_circuit)@my_pauli`. In classical programming, this would be like `5.7 + "pickle"`.
* __Both code and data__ - The Operator encodes both data (e.g. a matrix for eigensolution or a wavefunction being prepared) and computation (measure my wavefunction in this basis). There is little distinction between the two in Quantum programming.
* __Linear__ - The Operator is a recursively powerful construct, allowing algorithmic rearrangement not typically allowed in classical computation.
* `op1(op2(A,B)) == op1(op2(A)), op2(B))` in many cases, e.g. Expectation(A+B).
* The idea that `program(a*circuita + b*circuitb)` gives a mathematically valid result is highly surprising.
* __Algorithmically ubiquitous__ - Every quantum algorithm uses Operators. Algorithms are nearly always defined in literature by Operator operations. This language is rigorous, accepted, and compact.
* __Eagerly Computable__ - In most cases, Operator computation can be partially compiled as parameters become available, allowing improved performance, functional modularity (e.g. passing a ready-to-run algorithm), and inspection transparency. For example:
* A circuit can be compiled to a Qobj with parameters missing, to be filled in later
* The full list of circuits necessary to execute an algorithm can be prepared pending some operator coefficients
* A full algorithm can be prepared and passed to a user pending the insertion of some subcomponent (a choice of ExpectationValue algorithm) or parameters
#### Operator Definition: Primitives and Combinations
Operators can be _primitives_ or _combinations._ Primitives are base-level Operator representations which are not defined in terms of other primitives, but can be converted into one another with some computational work. Combinations are Operators which are constructed from functions of multiple primitives, such as sums and tensors. Combinations store the primitives from which they are constructed. Note that many Gates are present in other classes of primitives, and this must be reconciled as a follow-on to this redesign. The following should all be modules in the Operator hierarchy:
* Primitives
* Matrix
* Pauli - X, Y, Z, I
* QuantumCircuit, Gate
* Clifford
* Projector - Ze, O, P, M
* Stabilizer
* Graph State - Stored as a graph
* QuantumCircuit - Implicitly starts from |0⟩⟨0|
* Others (follow-on): ZX, MPS, Dirac Matrix, Gell-Mann matrix
* Combinations
* OpSum - Generalization of WeightedPauli. Stores a list of Operators of equal dimension and complex weights
* OpComposition - Stores a list of Operators which are all of equal dimension
* OpKron - Stores a list of Operators of any size
* OpVec - Stores a list of Operators of any size
* OpExp - Stores a single Operator, acting as a placeholder for some Evolution algorithm to replace later
* OpCombo - custom, user-defined recombination function
```
from qiskit.aqua.operators.pauli import X, Y, Z, I
op_new = .5*(I^X) + .2*(Z^Y) + .1j*(Z^Z)
op_new == pauli_op
```
Note that to support the above, the existing Pauli in Terra would need to support Tensor, sum, and scalar multiplication which can return an OpSum and OpKron.
The following overload operations are also desirable:
* Operator composition using `@` overload
* __Decision:__ deprecate the `*` overload for composition?
* Power (`**3`), kronpower (`^3`)
```
(pauli_op^2)**2 == (pauli_op^pauli_op)@(pauli_op^pauli_op)
from qiskit.aqua.ansatz import Ry
from qiskit.aqua.operators.projectors import Ze, O, P
ansatz = Ry(qubits=2, depth=3) @ (P^(-.1*O + 3*Ze))
# This is an OpSum of two circuits!
```
#### Algorithms Definition: Primitives and Composites
Operations on Operators also can be described as primitives or combinations of such. Primitives are computations which can be performed directly on some available computation engine, such as Numpy or Quantum Hardware, while composites are constructed from piping primitives together. Algorithms accept only _specific primitives,_ so an algorithm taking a Pauli vs. one taking a matrix are fundamentally different, but are also defined over certain combinations of their input primitives. For example, a Change-of-Basis Expectation Value is defined to accept a Pauli and a Projector (or QuantumCircuit acting as one from Zero implicitly), but can also accept sums, tensors, and vectorizations of Paulis and Projectors. If an unsupported primitive, such as Matrix or OpComposition were passed in, an exception would be thrown.
* Primitives
* Classical sum, product, tensor, trace, etc.
* Z-Basis QuantumCircuit measurement / Trace (traditional QASM backend)
* Primitive Conversion - Pauli to matrix, matrix to Pauli, etc.
* Evolution Conversion - Trotter, Suzuki, etc.
* Pauli Sum, Composition, Tensor
* Change of Basis - Pauli, Fourier
* Optimizers
* External functions, such as Drivers or imports
* Composites
* ExpectationValue
* Existing Aqua Algorithms: VQE, QPE, HHL, etc.
* Gradients
Over time, we have found that it is easiest to describe the behavior of Algorithms in terms of the flow of Operators through various components and subroutines. This description is naturally recursive, and considerably easier to understand than the present presentation of algorithmic flow in Aqua.
To demonstrate this, consider the following VQE coded from scratch in this model:
```
ansatz = Ry(qubits=2, depth=3) @ (P^P)
# Ansatz state = Ry(θ)|++⟩
hamiltonian = 3*(I^Z) + .4j*(X^Z)
expectation = PauliExpectation(ansatz, hamiltonian, backend)
print(expectation.run({ansatz.params: np.zeroes(len(ansatz.params))}))
# Print starting expectation
gradient = ParamShiftGradient(expectation)
optimizer = AQGD(initial_point=np.zeroes(len(ansatz.params)))
my_vqe = AQGD(cost_fn=expectation.run, grad_fn=gradient.run)
min_eig = my_vqe.run()
```
#### Parameterization and Eagerness
Operators and algorithms can be _parameterized,_ or missing some key information in order to execute. For Operators these may be sum coefficients, evolution times, QuantumCircuit parameters, and more. For Algorithms these may be input operators, execution parameters, or instances of algorithms used in computation which cannot be inferred by default (e.g. backend on which to execute, optimizer, etc.).
##### Eager Parameterization+Execution Interface Options:
An algorithm should execute as soon as it has filled the parameters necessary to do so. This is called **Eager Execution.** In a similar vein, OpSum can be seen as eagerly waiting for the contained operators to be summable, e.g. replaced with scalars by an expectation value. (**Decision**) Some interface options for eagerness:
**Option A**: Algorithms should be **callable** with a parameter dictionary, triggering a breadth-first search to parameterize any sub-objects with the parameter dictionary. This may be too much hocus pocus and difficult for implementers of algorithms to understand. A user may want to parameterize without executing, so an `execute` parameter should be available in the parameterization function.
```
my_op = Parameter('t1')*(Z^Z) + .6*(X^I)
my_vqe = VQE(backend=Parameter('backend'),
operator=my_op,
ansatz=Ry(qubits=2, reps=3),
optimizer=SLSQP(initial_point=Parameter('initial_rotations')))
my_vqe({'t1': .2j, 'backend': Aer.get_backend('qasm_simulator')})
# Didn't return anything yet
rots = np.zeros(len(my_vqe.ansatz.params))
min_eig = my_vqe({'initial_rotations': rots})
# Now a value is returned, and other execution information can be found inside the object
# Alternatively
my_vqe({'initial_rotations': rots}, execute=False)
min_eig = my_vqe()
```
**Option B:** Algorithms should have a `.run(param_dict)` method which accepts parameters and performs the breadth-first parameterization. The form factor of this would be similar to the above, but with `run()` instead of direct function calls. This has the benefit of some backward compatibility.
**Option C:** Algorithms should support separate parameterization and execution functions. This is the most explicit, but is clunky in an eager execution regime, where execution is automatic if the algorithm is sufficiently parameterized.
All of an Algorithm or Operator's pending Parameters should be recursively returned by a `.params` function. _(Tentative)_ A `deepcopy` option should be available to return a deep copy of the algorithm with the desired parameterization, rather than parameterize the algorithm in-place (this is evaluated with `execute=False` by default).
##### Eager Partial Computation
Aqua should be **eager** in partial computation while some parameters necessary for execution are not yet available, to allow for inspection transparency and performance. For example, once backend information is available, circuits should be transpiled for the backend or otherwise prepared for execution. This can avoid many transpilations or preparations later if the circuits are duplicated for Operator composition, as in Change-of-Basis expectation values or gradients.
The choice of which partial computation to perform is left to the algorithm, so only worthwhile partial computations are performed. If parameters change, re-preparing the partial computation can be expensive, so a `lazy` parameter should be available in the callable function.
### Changes to Terra
The `quantum_info` directory should be organized as follows:
* channel
* ...
* matrix.py **- Decision: Rename operator.py to matrix.py or matrix_op.py?**
* pauli.py
* clifford.py **- Decision: Use the Ignis's Clifford?**
* projector.py
* stabilizer.py
* Composites
* op_sum.py, op_composite.py, op_kron.py, op_vec.py, op_exp.py
In addition to the functionality detailed in [Object Model and Hierarchy](#Object-Model-and-Hierarchy) above, Terra should support the following for all of the above Non-matrix-based operators:
* `to_matrix()` - Method to allow quick access to unscalable classical tools, e.g. numpy eigensolution
* `to_quantum_circuits()` - returns a single or list of quantum circuits and coefficients representing the full Operator, including any distributive composition, tensors, etc.
* Trace, Partial Trace, Determinant, Norms, Adjoints - Where possible, linear algebra should be easily accessible
##### Follow-on: Terra Reconciliation Between Operators and Gates
Terra's Operators and Gates are currently fully distinct from one another. The X, Y, Z, Clifford Gates, Evolution by a matrix-specified Unitary (UnitaryGate), and more are direct overlaps between the two, but not interoperable. At some point, Terra should address this difference to allow Operators to be inserted onto a circuit, maintain only a single set of primitive unitaries, allow Gates to be composed with Operators, etc.
### Changes to Aqua
The changes to Aqua are basically just to
* deprecate the Operators after moving their logic into Terra,
* change the Aqua algorithms to rely on the new Terra operators,
* break up the Expectation, Evolution, circuit execution, and gradient code to be first-class algorithms users can extend and understand,
* and change the exsting Aqua algorithms to rely on these new algorithms.
##### Change Algorithms to rely on Terra operators and new Operator algorithms
In particular, algorithms should be accessible with only Terra-defined inputs (meaning constructed using Terra alone) to provide a seamless experience between Terra and Aqua usage, and extensible interfaces. For example, a VQE should be runnable by passing only a parameterized QuantumCircuit and Terra-defined Operator, allowing a provider or collaborator to share a custom VQE without an unnecessary dependency on Aqua. In particular, this allows the Aer Expectation Value to be defined with the same interface as Aqua's Pauli Expectation, without a dependency on Aqua.
##### Circuit Execution Algorithms - **Decision: Name - CircuitExecution? QCExecute? QuantumMeasureZ? RunCircuit?**
Circuit execution is a utility in Aqua today, mediated by the QuantumInstance, which most users do not understand, and growing increasingly branchy to accommodate more and more execution variants. Measurement error mitigation, noisy simulation setup, hardware API fault handling, and more all fall into the same execution flow in various branches.
Circuit execution is an algorithm for sampling a circuit's expectation in exponentially many ${Z, I}^{\otimes n}$ bases, but is not reflected an an algorithm today. It should be promoted to be a first-class algorithm to be more transparent and compartmental, wherein for example, code for simulation and code for execution on hardware can be kept distinct. A CircuitExecution Algorithm accepts a backend and interacts with it in some well-defined way - in way breaking up and organizing of the functionality of the QuantumInstance. Some examples of CircuitExecution algorithms are:
1. QuantumHardware - An Execution algorithm tailored for execution on remote hardware, including fault handling, slicing to limit job sizes, etc. Can stack up a queue of circuits for batch execution, or accept a list of jobids to use as the first n results objects, allowing the user to reuse results from a terminated execution.
1. IdealSimulator - Algorithm tailored for execution in ideal simulation.
1. NoisySimulator - Utility for querying a Hardware backend's properties and providing a noisy simulator using Aer's "noise config from device" functionality.
1. ErrorMitigatedExecutor - OUT OF SCOPE, BEING COVERED IN ANOTHER DOC.
If none is explicitly specified, Aqua should aggressively guess the preferred execution algorithm for the user given the backend and other execution parameters.
##### Expectation Algorithms
Aqua should support the following ExpectationValue algorithms. An `ExpectationBase` class should allow automatic selection of an evolution algorithm by default if none is specified - e.g. if the user has Aer installed, VQE will use the AerExpectation by default instead of QASM execution. Other possible expectation values include:
1. PauliExpectation (Change-of-Basis)
1. CVaRExpectation
1. AerExpectation - relies on Aer's fast expectation feature
1. MatrixExpectation
1. (Tentative) BasicAerExpectation
1. RichardsonExpectation - OUT OF SCOPE, BEING COVERED IN ANOTHER DOC.
##### Grouping
Grouping is an important feature within the PauliExpectation in Aqua today, but is not used by default, and has an interface which is not obvious. Grouping should be moved into the PauliExpectation, with a simple interface for the user to specify whether to group the Paulis, or how aggresively to do so. By default, the PauliExpectation should group Paulis as aggressively as is performant on the given execution backend.
##### Circuit Evolution Algorithms
And similarly for Evolution, a variety of algorithms should be available for converting a OpExp composite operator into a sum, composition, etc. More specifically, circuit evolution algorithms take an OpExp placeholder and return operators which approximate the value of the exponentiation. For example, the PauliEvolution accepts a Pauli and returns a QuantumCircuit representing the unitary evolution of that Pauli. An `EvolutionBase` class should allow automatic selection of an evolution algorithm by default if none is specified.
1. PauliEvolution (Change-of-Basis)
1. SumEvolution
1. Trotter
1. Suzuki
1. MatrixEvolution
1. (Tentative) [LinCombEvolution](https://arxiv.org/abs/1202.5822)
1. (Tentative) AerEvolution
1. (Tentative) BasicAerEvolution
##### Other algorithms to build out into first-class Algorithm groups
1. Converters - convert lazily between Operator types
1. Gradient
1. Optimization
## Timeline and Gameplan
Stage 1: Implement new Operators in Terra with thorough unit and integration tests.
Stage 2: Implement Operator algorithms in Aqua, relying on Terra Operators
Stage 3: Migrate Aqua algorithms to rely on new Operator algorithms and new Terra Operators
Stage 4: Deprecate Present Aqua Operators (0.7 release)
Stage 5: Delete Present Aqua Operators (0.8 release)
## ⚰️⚰️⚰️⚰️⚰️⚰️ Graveyard ⚰️⚰️⚰️⚰️⚰️⚰️
### Other Benefits of OperatorFlow
* Obedient Eager Evaluation - Best of Eager and Lazy evaluation:
* Partially evaluate whatever you can with the parameters you have
* Allows transparency, inspection, rapid prototyping (e.g. Users couldn't find circuits or operator when working through JSON dictionaries)
* Performance - partially compiled algorithms save massive amounts of compilation and deepcopy time
* But not too early, not compiling preemptively for a possible parameter value
* Objects can be returned without being totally incorrectly constructed for the next step or engine (e.g. building massive CNOT chains for UCCSD simulations)
* Intractable but possible computations (e.g. convert to matrix and solve) are avoided
* Natural, Powerful, and Self-defining Programming Interfaces
* __An algorithm's behavior is simply defined by the operator primitives it accepts and returns__
* Nesting of algorithms is identical to user algorithm execution
* Ubiquitous parameters, and obvious interface for Optimization
* OpCombo coefficients, primitive parameters, and algorithm parameters can all be parameterized
* Algorithms of any level of completeness can be returned
* Optimization is universal - simply pass a nearly-complete algorithm to an optimizer and the callable interface executes when the optimizer provides the parameters
#### Grouping
Aqua's grouping functionality is only relevant to ExpectationValues today.
```
qaoa_cost_op = WeightedPauliOperator([
[.5, Pauli.from_label('ZIZ')],
[.2, Pauli.from_label('ZZI')],
[.1j, Pauli.from_label('IZZ')],
])
grouped_cost_op = TPBGroupedWeightedPauliOperator.sorted_grouping(qaoa_cost_op)
grouped_cost_op._basis
class VQE(QuantumAlgorithm):
def __init__(self, operator, var_form, optimizer,
initial_point=None, backend=backend, callback=None, ...):
...
self._expectation_value = ExpectationValue(self._operator, self._backend)
def _energy_evaluation(self, params):
circuits = self._var_form.construct_circuit(params)
energy, stdev = self._expectation_value.run(circuits)
return energy
```
| github_jupyter |
# PDOS data analysis and plotting
---
### Import Modules
```
import os
print(os.getcwd())
import sys
import plotly.graph_objs as go
import matplotlib.pyplot as plt
from scipy import stats
# #########################################################
from methods import get_df_features_targets
from proj_data import scatter_marker_props, layout_shared
```
### Read Data
```
df_features_targets = get_df_features_targets()
```
```
# df_features_targets.columns.tolist()
pband_indices = df_features_targets[[
(
'features',
# 'oh',
'o',
'p_band_center',
)
]].dropna().index.tolist()
df_i = df_features_targets.loc[
pband_indices
][[
("targets", "g_oh", ""),
("targets", "g_o", ""),
("targets", "g_o_m_oh", ""),
("targets", "e_oh", ""),
("targets", "e_o", ""),
("targets", "e_o_m_oh", ""),
("features", "o", "p_band_center"),
]]
# pband_indices =
df_features_targets[[
(
'features',
# 'oh',
'o',
'p_band_center',
)
]]
# ]].dropna().index.tolist()
# assert False
# df_features_targets.shape
# (288, 7)
# (311, 7)
# (312, 7)
# (316, 7)
# df_i.shape
# assert False
# df_i[""]
df = df_i
df = df[
(df["features", "o", "p_band_center"] > -3.5) &
(df["features", "o", "p_band_center"] < -2.) &
# (df[""] == "") &
# (df[""] == "") &
[True for i in range(len(df))]
]
df_i = df
x = df_i["features", "o", "p_band_center"]
# y = df_i["targets", "g_oh", ""]
# y = df_i["targets", "g_o", ""]
y = df_i["targets", "g_o_m_oh", ""]
# y = df_i["targets", "e_o_m_oh", ""]
res = stats.linregress(x, y)
y_new_fit = res.intercept + res.slope * x
def colin_fit(p_i):
g_o_m_oh_i = 0.94 * p_i + 3.58
return(g_o_m_oh_i)
trace_colin_fit = go.Scatter(
x=[-6, 0],
y=[colin_fit(-6), colin_fit(0)],
mode="lines",
name="Colin fit (G_OmOH = 0.94 * p_i + 3.58)",
)
trace_my_fit = go.Scatter(
x=x,
y=y_new_fit,
mode="lines",
name="Colin fit (G_OmOH = 0.94 * p_i + 3.58)",
)
y_new_fit
trace = go.Scatter(
x=x, y=y,
mode="markers",
name="My DFT data",
)
x_i = x.to_numpy()
X = x_i.reshape(-1, 1)
import numpy as np
from sklearn.linear_model import LinearRegression
# X = np.array([[1, 1], [1, 2], [2, 2], [2, 3]])
# y = 1 * x_0 + 2 * x_1 + 3
# y = np.dot(X, np.array([1, 2])) + 3
reg = LinearRegression().fit(X, y)
reg.score(X, y)
print(
reg.coef_,
reg.intercept_,
)
# reg.predict(np.array([[3, 5]]))
y_pred_mine = reg.predict(
[[-6], [2]],
)
trace_my_fit = go.Scatter(
x=[-6, 2],
y=y_pred_mine,
mode="lines",
name="My fit (G_OmOH = 0.75 * p_i + 3.55)",
)
data = [trace, trace_colin_fit, trace_my_fit]
# data = [trace, trace_colin_fit, trace_my_fit]
layout_mine = go.Layout(
showlegend=False,
xaxis=go.layout.XAxis(
title=go.layout.xaxis.Title(
text="ε<sub>2p</sub>",
),
range=[-6, 0, ]
),
yaxis=go.layout.YAxis(
title=go.layout.yaxis.Title(
text="ΔE<sub>O-OH</sub>",
),
range=[-3, 4, ]
),
)
# #########################################################
layout_shared_i = layout_shared.update(layout_mine)
fig = go.Figure(data=data, layout=layout_shared_i)
fig.show()
```
```
# df_i
# df_features_targets
# (0.94 * 0 + 3.58) - (0.94 * 3 + 3.58)
# 0.94 * 0.3
# res.intercept
# res.slope
# layout = go.Layout(
# xaxis=go.layout.XAxis(
# title=go.layout.xaxis.Title(
# text="ε<sub>2p</sub>",
# ),
# ),
# yaxis=go.layout.YAxis(
# title=go.layout.yaxis.Title(
# text="ΔE<sub>O-OH</sub>",
# ),
# ),
# )
```
| github_jupyter |
### Basic Functions for Interactively Exploring the CORTX Metrics Stored in Pickles
```
%cd /home/johnbent/cortx/metrics
import cortx_community
import cortx_graphing
import os
from github import Github
gh = Github(os.environ.get('GH_OATH'))
stx = gh.get_organization('Seagate')
repos = cortx_community.get_repos()
ps = cortx_community.PersistentStats()
# a function which can test the progress of a running scrape_metrics.py process
def check_scan_progress(date,ps):
done=0
for repo in ps.get_repos():
(a,b)=ps.get_latest(repo)
if b == date:
done+=1
#print("Last report for %s is %s" % (repo,b))
print("%d out of %d repos have been scanned" % (done,len(ps.get_repos())))
# a function for comparing a field in a repo over time
# for example, if you want to see the change in innersource_committers over time, you can use this function
def compare_fields(ps,repo,field,verbose=False):
last = None
first = None
for date in sorted(ps.stats[repo].keys()):
try:
last = ps.stats[repo][date][field]
except KeyError:
pass # many fields didn't always exist so might not appear in old stats
if first is None:
first = last
if verbose:
print("%s -> %s" % (date, last))
print("Difference between first and last is: %s" % (first-last))
print("Difference between last and first is: %s" % (last-first))
compare_fields(ps,'GLOBAL','external_committers',verbose=True)
#print(ps.get_values('GLOBAL','external_committers'))
# manually add some values so they can be in executive report
repo='GLOBAL'
slack_members_key='slack_total_members'
slack_wau_key='slack_weekly_ave_active'
newsletter_key='newsletter_subscribers'
webinar_key='webinar_attendees'
integrations_key='integrations'
dates=ps.get_dates(repo)
ec_key='external_committers'
ep_key='external_participants'
hc_key='hackathon_committers'
hp_key='hackathon_participants'
eup_key='eu_r&d_participants'
# the helper function to load them
def add_stats(List,Key):
for item in List:
date=item[0]
value=item[1]
print("Repo",repo,"Date",date,"Stat",Key,"Value",value)
ps.add_stat(date=date,repo=repo,stat=Key,value=value)
# the date strings to use
jun_date='2020-07-06'
sep_date='2020-10-07'
oct_date='2020-11-03'
nov_date='2020-12-21'
dec_date='2020-12-30'
last_date='2021-01-06' # must add metrics for the latest date or else they don't show up in plots . . .
def update_slack():
# adding the slack metrics for executive report
slack_members =[(jun_date,97), (sep_date,212), (nov_date,329), (dec_date,355),(last_date,355)]
slack_weekly_average_active=[(jun_date,11.3),(sep_date,53.0),(nov_date,69.9),(dec_date,59.9),(last_date,59.9)]
add_stats(slack_members,slack_members_key)
add_stats(slack_weekly_average_active,slack_wau_key)
print(ps.get_values(repo,slack_members_key))
print(ps.get_values(repo,slack_wau_key))
def update_newsletter():
# adding the newsletter metrics for executive report
dec_date='2020-12-30'
feb_date='2021-02-13'
newsletter_feb=set(["zengyuan.liu@seagate.com","zayne@uchicago.edu","yuriy.umanets@seagate.com","yuji_yazawa@mail.toyota.co.jp","yoongchin.lim@seagate.com","yihua.jiang@seagate.com","yichun.fan@seagate.com","yeshpal.jain@seagate.com","yazid.muhammad@seagate.com","yatin.mahajan@seagate.com","yashodhan.pise@seagate.com","yash.bhamare@seagate.com","yanqing.f.fu@seagate.com","xiaoyong.d.dai@seagate.com","xiaohui.d.dong@seagate.com","woosik.kim@seagate.com","wendell.wenjen@seagate.com","wei.x.xie@seagate.com","wdchien@kth.se","wangm12@uchicago.edu","vivek.m@seagate.com","vishwas.bhat@seagate.com","vishal.maurya@seagate.com","vishal.dhobale@seagate.com","vinoth.v@seagate.com","vimlesh.kumar@seagate.com","vikram.jadhav@seagate.com","vikas.kumar@seagate.com","vijaykumar.thakkar@seagate.com","vicente.armendariz@seagate.com","vibhuti.singh@seagate.com","venkateswarlu.payidimarry@seagate.com","venkatesh.k@seagate.com","venkatesh.balagani@seagate.com","venkataraman.padmanabhan@seagate.com","varunreddy.boddu@seagate.com","vaibhav.paratwar@seagate.com","utz-uwe.haus@hpe.com","upendra.patwardhan@seagate.com","ujjwal.lanjewar@seagate.com","udayan.yaragattikar@seagate.com","Tushar.gohad@intel.com","trupti.patil@seagate.com","trent.geerdes@seagate.com","toru.takano@seagate.com","tony.tian@seagate.com","tom.z.zhao@seagate.com","tom.r.prohofsky@seagate.com","tom.petaja@seagate.com","tom.dimauro@seagate.com","tobby.yl.tong@seagate.com","tim.t.walker@seagate.com","thavanathan.thangaraj@seagate.com","thanit.karnthak@seagate.com","tejal.sheth@seagate.com","tdavis@deepspacestorage.com","tasneem.vijapure@seagate.com","taro.iwata@seagate.com","tadeu.bastos@seagate.com","t.kai@fujitsu.com","swati.magar@seagate.com","swapnil.khandare@seagate.com","swapnil.chaudhary@seagate.com","suvrat.joshi@seagate.com","suraj.kadam@seagate.com","suprit.shinde@seagate.com","suppakrit.kirdponpattara@seagate.com","sunil.sonale@seagate.com","sunil.savanur@seagate.com","sumit.sharma@seagate.com","sumit.gupta@seagate.com","sumedh.a.kulkarni@seagate.com","subhash.arya@seagate.com","steven.sanchez@seagate.com","stephen.muhs@seagate.com","sridhar.dubbaka@seagate.com","sreenivasulu.bachu@seagate.com","sradhanand.pati@seagate.com","soniya.moholkar@seagate.com","songwee.teo@seagate.com","sonal.kalbende@seagate.com","sining.wu@seagate.com","sienhuay.chong@seagate.com","shubham.patnaik@seagate.com","shubham.bhosale@seagate.com","shriya.deshmukh@seagate.com","shrihari.waskar@seagate.com","shri.metta@seagate.com","shreyas.vidvans@seagate.com","shreya.karmakar@seagate.com","shreekant.upadhyay@seagate.com","shraddha.chaudhari@seagate.com","shipra.gupta@seagate.com","shazia.ahmad@seagate.com","shaun.de-witt@ukaea.uk","shaun.bruce@seagate.com","shashank.parulekar@seagate.com","shankar.more@seagate.com","shalaka.dharap@seagate.com","shailesh@weka.io","shailesh.vaidya@seagate.com","sergey.shilov@seagate.com","sebastien.valat@atos.net","scott.tolson@seagate.com","scott.fast@wwt.com","savitharani.ravichandran@seagate.com","saurabh.khanvilkar@seagate.com","saumya.sunder@seagate.com","saumitra.kulkarni@seagate.com","satish.darade@seagate.com","sanmitra.sinha@seagate.com","sanjog.naik@seagate.com","sandeep.mathew@seagate.com","sandeep.anjara@seagate.com","samuel.duncanson@seagate.com","sampada.petkar@seagate.com","samantha.clarke@seagate.com","sakchai.suntinuraks@seagate.com","sai.narasimhamurthy@seagate.com","sachitanand.shelake@seagate.com","sachin.punadikar@seagate.com","sachin.jagtap@seagate.com","ryan.goss@seagate.com","rupasree.roy@seagate.com","rross@mcs.anl.gov","rplaster@deepspacestorage.com","rizvi.ahmed@seagate.com","richard.buchan@seagate.com","ricardo.alvarez@seagate.com","remi.seghier@seagate.com","raymond.chang@seagate.com","ravindra.choudhari@seagate.com","randy.neill@seagate.com","ramesh.vegesna@seagate.com","ramesh.potu@seagate.com","ramakrishna.chintalapati@seagate.com","rakesh.surve@seagate.com","rakesh.sahuu@seagate.com","rajkumar.patel@SeagateTechnology.onmicrosoft.com","rajesh.nambiar@seagate.com","rajesh.deshmukh@seagate.com","rajesh.chouhan@seagate.com","raja.mohanty@seagate.com","raj.das@seagate.com","rahul.telawade@seagate.com","rahul.ranjan@seagate.com","rahul.modi@seagate.com","rahul.kumar@seagate.com","rahul.jyoti@seagate.com","radha.gulhane@seagate.com","rachel.novak@seagate.com","raaf@uni-mainz.de","puja.mudaliar@seagate.com","priyanka.rathi@seagate.com","priyanka.borawake@seagate.com","priyank.p.dalal@seagate.com","pritam.bhavsar@seagate.com","pratyush.k.khan@seagate.com","prathamesh.rodi@seagate.com","pranay.kumar@seagate.com","pranav.risbud@seagate.com","pranav.pawar@seagate.com","pranali.ugale@seagate.com","pranali.tirkhunde@seagate.com","praful.sambe@seagate.com","pradeep.kumbhre@seagate.com","prabhsimran.singh@seagate.com","pooja.pandey@seagate.com","pierre.lebars@seagate.com","philippe.couvee@atos.net","phil.ruff@seagate.com","peyton.mcnully@dcblox.com","peter.williams@seagate.com","peter.maddocks@seagate.com","pcpeng@uchicago.edu","pawan.kumarsrivastava@seagate.com","paul.woods@seagate.com","paul.heath@seagate.com","paul.croft@seagate.com","patricia.simon@seagate.com","parikshit.dharmale@seagate.com","parag.1.joshi@seagate.com","papan.kumarsingh@seagate.com","padmaja.kannan@seagate.com","nyunt.n.ho@seagate.com","nitesh.mahajan@seagate.com","nino.wicaksono@seagate.com","nilesh.govande@seagate.com","nikos.nikoleris@arm.com","nikita.danilov@seagate.com","nikhil.sawake@seagate.com","nikhil.birgade@seagate.com","nihar.nayak@seagate.com","neha.singh@seagate.com","neerav.choudhari@seagate.com","naval.patel@seagate.com","nate.nally@seagate.com","natalie.mujicaschwahn@seagate.com","narayanan.krishnamurthy@seagate.com","namrata.khake@seagate.com","nalinikanta.jena@seagate.com","nahoosh.mandlik@seagate.com","n.krauter@uni-mainz.de","mukund.kanekar@seagate.com","mukul.malhotra@seagate.com","muhammadfairuz.anwar@seagate.com","muhammad.ahmad@seagate.com","mohit.pathak@seagate.com","mohamadarafat.abdulraheem@seagate.com","mohamad.chaarawi@intel.com","mnizamshah.azzudinshah@seagate.com","mlcurry@sandia.gov","mingjin.w.wang@seagate.com","ming.yik@seagate.com","milind.naik@seagate.com","michelle.e.langan@seagate.com","michele.fagan@seagate.com","mehul.joshi@seagate.com","mazhar.inamdar@seagate.com","mayur.dharmik@seagate.com","maxim.malezhin@seagate.com","max.medved@seagate.com","matthew.halcomb@wwt.com","matt.james@seagate.com","mason.swarr@seagate.com","markidis@kth.se","mark.wiggins@seagate.com","mark.sprouse@dcblox.com","mark.a.penas@seagate.com","mariyappan.ponnusamy@seagate.com","manueljrturao.cabusas@seagate.com","manoj.patil@seagate.com","mandar.sawant@seagate.com","mandar.sabhapatikar@seagate.com","mandar.joshi@seagate.com","mandar.d.joshi@seagate.com","mahima.gupta@seagate.com","mahesh.agarkar@seagate.com","mahendra.shinde@seagate.com","madhura.mande@seagate.com","madhav.vemuri@seagate.com","lynette.sy.neo@seagate.com","luhchyuan.lau@seagate.com","liping.ding@seagate.com","linqiang.luo@seagate.com","lianhoo.tan@seagate.com","liang.gan@seagate.com","lenin.jegatheesan@seagate.com","ladislav.hudec@seagate.com","konstantin.nekrasov@seagate.com","kishan.nayak@seagate.com","kishan.gelli@seagate.com","kiran.mangalore@seagate.com","kimchui.lee@seagate.com","kianwee.tan@seagate.com","kevin.t.james@seagate.com","kevin.a.price@seagate.com","ketan.arlulkar@seagate.com","ken.haugen@seagate.com","kavya.motwani@seagate.com","kaustubh.deorukhkar@seagate.com","kathy.chun@seagate.com","karun.sharma@seagate.com","kapil.jinna@seagate.com","kanchan.chaudhari@seagate.com","kalyan.dandu@seagate.com","kalpesh.chhajed@seagate.com","ka.naidu@seagate.com","jyoti.baral@seagate.com","justin.woo@seagate.com","junichi.hyodo@seagate.com","jugal.patil@seagate.com","josiah.gelsinger@seagate.com","josh.z.li@seagate.com","joseph.rettinghouse@seagate.com","joseph.maniaci@seagate.com","johnbent@gmail.com","john.forgan@seagate.com","john.callender@seagate.com","john.bent@seagate.com","johann.lombardi@intel.com","jin.m.li@seagate.com","jiahui.sun@seagate.com","jerry.l.zhen@seagate.com","jeetandra.kella@seagate.com","jayshree.more@seagate.com","jaydeep.mohite@seagate.com","jayashree.thite@seagate.com","jay.jaiantilal@seagate.com","james.mayfield@seagate.com","james.cleverley@seagate.com","jacques-charles.lafoucriere@cea.fr","ivy.guo@seagate.com","ivan.tishchenko@seagate.com","ivan.poddubnyy@seagate.com","ivan.alekhin@seagate.com","isaacyizhe.teoh@seagate.com","inghong.kong@seagate.com","indrajit.zagade@seagate.com","iman.anvari@seagate.com","igor.pivovarov@seagate.com","ian.davies@seagate.com","i.panourgias@epcc.ed.ac.uk","hua.huang@seagate.com","hemant.raut@seagate.com","harshitkumar.jain@seagate.com","harrison.seow@seagate.com","haibing.ma@seagate.com","gregory.touretsky@seagate.com","graham.d.ferris@seagate.com","gowthaman.chinnathambi@seagate.com","girard.goder@seagate.com","ginann.c.cowen@seagate.com","gheewooi.ong@seagate.com","gaurav.gaur@seagate.com","gaurav.chaudhari@seagate.com","gary.phillips@seagate.com","ganesan.umanesan@seagate.com","gabriel.ruiz@seagate.com","fred.zellinger@seagate.com","evgeniy.brazhnikov@seagate.com","evelyn.godwyn@seagate.com","erik.d.salo@seagate.com","eduard.aleksandrov@seagate.com","earl.dodd@wwt.com","don.r.bloyer@seagate.com","dmitriy.chumak@seagate.com","dmitrii.surnin@seagate.com","divyam.singla@seagate.com","divya.kachhwaha@seagate.com","dipendra.m.bagchee@seagate.com","dinesh.mohanakrishnan@seagate.com","dimitry.didenko@seagate.com","dhananjay.dandapat@seagate.com","deepak.choudhary@seagate.com","debasmita.samaddar@ukaea.uk","david.white@seagate.com","dattaprasad.govekar@seagate.com","daniel.lerner@seagate.com","dahlan.saragih@seagate.com","d.pleiter@fz-juelich.de","cornel.crisan@atos.net","clinton.l.rowe@seagate.com","christopher.haine@hpe.com","christopher.bishop@seagate.com","christina.ku@seagate.com","chris.maio@seagate.com","chong.li@seagate.com","chinfong.hee@seagate.com","chiawhueoy.chin@seagate.com","chetan.srivastava@seagate.com","chetan.deshmukh@seagate.com","cheryl.roberts@seagate.com","cheewai.lum@seagate.com","cheesiong.toh@seagate.com","charles.kunkel@seagate.com","cary.dickens@seagate.com","carrie.harris@seagate.com","bryan.j.baker@seagate.com","bruno.j.masson@seagate.com","brinkman@uni-mainz.de","brian.lavash@seagate.com","brian.ch.chen@seagate.com","brian.a.resch@seagate.com","bikrant.singh@seagate.com","bhavin.katira@seagate.com","bhargava.sreepathi@seagate.com","bhairavi.alurkar@seagate.com","beilei.w.wang@seagate.com","basavaraj.kirunge@seagate.com","bansidhar.soni@seagate.com","balaji.ramachandran@seagate.com","azahar.khan@seagate.com","atul.deshmukh@seagate.com","atita.shirwaikar@seagate.com","ashwini.borse@seagate.com","ashwath.gundepally@seagate.com","ashish.dhavalikar@seagate.com","archana.patil@seagate.com","archana.limaye@seagate.com","arati.kulkarni@seagate.com","apurwa.mohite@seagate.com","anudeep.kankipati@seagate.com","AnneMarie.DeBoard@seagate.com","ankit.yadav@seagate.com","anjali.somwanshi@seagate.com","anirudh.n.joshi@seagate.com","anilkumar.sindhi@seagate.com","andriy.tkachuk@seagate.com","andrey.kononykhin@seagate.com","andrei.zheregelia@seagate.com","anatoliy.bilenko@seagate.com","anagha.g.latne@seagate.com","anagha.deshmukh@seagate.com","amol.shinde@seagate.com","amol.j.kongre@seagate.com","amit.kumar@seagate.com","amit.kapil@seagate.com","alphonsus.kh.kwok@seagate.com","alexander.voronov@seagate.com","alexander.sukhachev@seagate.com","alan.gilda@seagate.com","ajinkya.dhumal@seagate.com","ajay.srivastava@seagate.com","ajay.shingare@seagate.com","ajay.paratmandali@seagate.com","agnieszka.zielinska@seagate.com","abhishek.saha@seagate.com","abhilekh.mustapure@seagate.com","abdul.habeeb@seagate.com"," charles.lafoucriere@cea.fr"," daniar@uchicago.edu"," dbonnie@lanl.gov"," dev@tiger-technology.com"," don.molaro@wwt.com"," ftessier@cscs.ch"," furuta.tomonori@fujitsu.com"," gunawi@gmail.com"," jeff.fonke@wwt.com"," lposton@nvidia.com"," lucy.gonzalez@arm.com"," m.weiland@epcc.ed.ac.uk"," miyamoto.kouki@fujitsu.com"," philippe.deniel@cea.fr"," rich.harper@wwt.com"," shail.manjrekar@gmail.com"," shaun.de-witt@ukaea.com"," sunny.raskar@msystechnologies.com"," theo@stillwater-sc.com"," tiago.quintino@ecmwf.int","sujata.taware@seagate.com","snehal.virkud@seagate.com","terry.harper@intel.com","drew.stern@nb.com","sati.viensiri@seagate.com","pedro.fernandez@seagate.com","rose.hiu@seagate.com","yoram.novick@gmail.com","pnicolas@coldago.com","navneet479@yahoo.co.in","patrick.bickley@seagate.com","gilbertsosi@gmail.com","mondpl@gmail.com","thomas.doran@seagate.com","pupmaison@neuf.fr","dh0253@yahoo.com","eng.amr214@yahoo.com","boyelroyjetson@yahoo.com","src223@gmail.com","drived@yandex.ru","kimchui.lee@gmail.com","peter.r.janik@seagate.com","renee.wilson@seagate.com","chuan.zhong@seagate.com","eddietolleson21@gmail.com","discoverypark1026@gmail.com","daniar@uchicago.edu","dbmuse@gmail.com","chris.cramer@seagate.com","jazztan1986@gmail.com","rrai@msystechnologies.com","ffrankenstein920@gmail.com","Jak12445@mail.ru","krn@krn.dk","kianchye.tan@seagate.com","lamarahenderson@gmail.com","SECRETARIA3134@GMAIL.COM","georgewu@qnap.com","theminimidistudio@gmail.com","ganliang0122@gmail.com","robbie.cazzaniga@gmail.com","j8862j@naver.com","474trizmo@gmail.com","torreschristina0541@gmail.com","hakimwong71@gmail.com","chintumoturu106@gmail.com","tovijayn@yahoo.com","Jeovalima54@gmail.com","lemperezgeoffre@gmail.com","gwaschke@videotron.ca","adeeba.rashid@marriott.com","mmichaelhooper208@gmail.com","abdulla.fayez@believerit.com","Lior.reinitz@gmail.com","bittorrentsoftware91@gmail.com","yasin@sahinyapi.com.tr","stevencheok@gmail.com","vitaliivladimirivich@gmail.com","julialima.jbl@gmail.com","matthew@tldus.com","cody01100011@gmail.com","patrickhession97@gmail.com","richardreed01@outlook.com","cathy101296@hotmail.com","karamello5150@gmail.com","jan.duymelinck@gmail.com","xrdeem@gmail.com","tom1910az@yahoo.com","salgadofranco303@gmail.com","leenaasiltanen@gmail.com","kevinma.vn@gmail.com","jennelswick5366@gmail.com","gemastudios@gmail.com","vladimirvazquezambriz@gmail.com","erin.markert@seagate.com","cethhamner67@gmail.com","smnspnl9@gmail.com","adalidportillo766@gmail.com","robert@rcl-associates.com","chris.wood@cmesecurity.ca","kimberlee.imig@gmail.com","thombrepk@outlook.com","datawormxproject@gmail.com","garyp04@rogers.com","emanuele.dec@gmail.com","elliemeistopforth@gmail.com","jmt@amco999.com","mohammeddawod2007@hotmail.com","gemdavis7@gmail.com","winter@example.com","r_gaisbauer@charter.net","artemfaber4@gmail.com","icatchmedia8@gmail.com","nitaaa2019@gmail.com","libin.l.cai@seagate.com","vpsinghindia@gmail.com","giomay110315@gmail.com"])
newsletter_members_dec=set(["zengyuan.liu@seagate.com","zayne@uchicago.edu","yuriy.umanets@seagate.com","yuji_yazawa@mail.toyota.co.jp","yoongchin.lim@seagate.com","yihua.jiang@seagate.com","yichun.fan@seagate.com","yeshpal.jain@seagate.com","yazid.muhammad@seagate.com","yatin.mahajan@seagate.com","yashodhan.pise@seagate.com","yash.bhamare@seagate.com","yanqing.f.fu@seagate.com","xiaoyong.d.dai@seagate.com","xiaohui.d.dong@seagate.com","woosik.kim@seagate.com","wendell.wenjen@seagate.com","wei.x.xie@seagate.com","wdchien@kth.se","wangm12@uchicago.edu","vivek.m@seagate.com","vishwas.bhat@seagate.com","vishal.maurya@seagate.com","vishal.dhobale@seagate.com","vinoth.v@seagate.com","vimlesh.kumar@seagate.com","vikram.jadhav@seagate.com","vikas.kumar@seagate.com","vijaykumar.thakkar@seagate.com","vicente.armendariz@seagate.com","vibhuti.singh@seagate.com","venkateswarlu.payidimarry@seagate.com","venkatesh.k@seagate.com","venkatesh.balagani@seagate.com","venkataraman.padmanabhan@seagate.com","varunreddy.boddu@seagate.com","vaibhav.paratwar@seagate.com","utz-uwe.haus@hpe.com","upendra.patwardhan@seagate.com","ujjwal.lanjewar@seagate.com","udayan.yaragattikar@seagate.com","Tushar.gohad@intel.com","trupti.patil@seagate.com","trent.geerdes@seagate.com","toru.takano@seagate.com","tony.tian@seagate.com","tom.z.zhao@seagate.com","tom.r.prohofsky@seagate.com","tom.petaja@seagate.com","tom.dimauro@seagate.com","tobby.yl.tong@seagate.com","tim.t.walker@seagate.com","thavanathan.thangaraj@seagate.com","thanit.karnthak@seagate.com","tejal.sheth@seagate.com","tdavis@deepspacestorage.com","tasneem.vijapure@seagate.com","taro.iwata@seagate.com","tadeu.bastos@seagate.com","t.kai@fujitsu.com","swati.magar@seagate.com","swapnil.khandare@seagate.com","swapnil.chaudhary@seagate.com","suvrat.joshi@seagate.com","suraj.kadam@seagate.com","suprit.shinde@seagate.com","suppakrit.kirdponpattara@seagate.com","sunil.sonale@seagate.com","sunil.savanur@seagate.com","sumit.sharma@seagate.com","sumit.gupta@seagate.com","sumedh.a.kulkarni@seagate.com","subhash.arya@seagate.com","steven.sanchez@seagate.com","stephen.muhs@seagate.com","sridhar.dubbaka@seagate.com","sreenivasulu.bachu@seagate.com","sradhanand.pati@seagate.com","soniya.moholkar@seagate.com","songwee.teo@seagate.com","sonal.kalbende@seagate.com","sining.wu@seagate.com","sienhuay.chong@seagate.com","shubham.patnaik@seagate.com","shubham.bhosale@seagate.com","shriya.deshmukh@seagate.com","shrihari.waskar@seagate.com","shri.metta@seagate.com","shreyas.vidvans@seagate.com","shreya.karmakar@seagate.com","shreekant.upadhyay@seagate.com","shraddha.chaudhari@seagate.com","shipra.gupta@seagate.com","shazia.ahmad@seagate.com","shaun.de-witt@ukaea.uk","shaun.bruce@seagate.com","shashank.parulekar@seagate.com","shankar.more@seagate.com","shalaka.dharap@seagate.com","shailesh@weka.io","shailesh.vaidya@seagate.com","sergey.shilov@seagate.com","sebastien.valat@atos.net","scott.tolson@seagate.com","scott.fast@wwt.com","savitharani.ravichandran@seagate.com","saurabh.khanvilkar@seagate.com","saumya.sunder@seagate.com","saumitra.kulkarni@seagate.com","satish.darade@seagate.com","sanmitra.sinha@seagate.com","sanjog.naik@seagate.com","sandeep.mathew@seagate.com","sandeep.anjara@seagate.com","samuel.duncanson@seagate.com","sampada.petkar@seagate.com","samantha.clarke@seagate.com","sakchai.suntinuraks@seagate.com","sai.narasimhamurthy@seagate.com","sachitanand.shelake@seagate.com","sachin.punadikar@seagate.com","sachin.jagtap@seagate.com","ryan.goss@seagate.com","rupasree.roy@seagate.com","rross@mcs.anl.gov","rplaster@deepspacestorage.com","rizvi.ahmed@seagate.com","richard.buchan@seagate.com","ricardo.alvarez@seagate.com","remi.seghier@seagate.com","raymond.chang@seagate.com","ravindra.choudhari@seagate.com","randy.neill@seagate.com","ramesh.vegesna@seagate.com","ramesh.potu@seagate.com","ramakrishna.chintalapati@seagate.com","rakesh.surve@seagate.com","rakesh.sahuu@seagate.com","rajkumar.patel@SeagateTechnology.onmicrosoft.com","rajesh.nambiar@seagate.com","rajesh.deshmukh@seagate.com","rajesh.chouhan@seagate.com","raja.mohanty@seagate.com","raj.das@seagate.com","rahul.telawade@seagate.com","rahul.ranjan@seagate.com","rahul.modi@seagate.com","rahul.kumar@seagate.com","rahul.jyoti@seagate.com","radha.gulhane@seagate.com","rachel.novak@seagate.com","raaf@uni-mainz.de","puja.mudaliar@seagate.com","priyanka.rathi@seagate.com","priyanka.borawake@seagate.com","priyank.p.dalal@seagate.com","pritam.bhavsar@seagate.com","pratyush.k.khan@seagate.com","prathamesh.rodi@seagate.com","pranay.kumar@seagate.com","pranav.risbud@seagate.com","pranav.pawar@seagate.com","pranali.ugale@seagate.com","pranali.tirkhunde@seagate.com","praful.sambe@seagate.com","pradeep.kumbhre@seagate.com","prabhsimran.singh@seagate.com","pooja.pandey@seagate.com","pierre.lebars@seagate.com","philippe.couvee@atos.net","phil.ruff@seagate.com","peyton.mcnully@dcblox.com","peter.williams@seagate.com","peter.maddocks@seagate.com","pcpeng@uchicago.edu","pawan.kumarsrivastava@seagate.com","paul.woods@seagate.com","paul.heath@seagate.com","paul.croft@seagate.com","patricia.simon@seagate.com","parikshit.dharmale@seagate.com","parag.1.joshi@seagate.com","papan.kumarsingh@seagate.com","padmaja.kannan@seagate.com","nyunt.n.ho@seagate.com","nitesh.mahajan@seagate.com","nino.wicaksono@seagate.com","nilesh.govande@seagate.com","nikos.nikoleris@arm.com","nikita.danilov@seagate.com","nikhil.sawake@seagate.com","nikhil.birgade@seagate.com","nihar.nayak@seagate.com","neha.singh@seagate.com","neerav.choudhari@seagate.com","naval.patel@seagate.com","nate.nally@seagate.com","natalie.mujicaschwahn@seagate.com","narayanan.krishnamurthy@seagate.com","namrata.khake@seagate.com","nalinikanta.jena@seagate.com","nahoosh.mandlik@seagate.com","n.krauter@uni-mainz.de","mukund.kanekar@seagate.com","mukul.malhotra@seagate.com","muhammadfairuz.anwar@seagate.com","muhammad.ahmad@seagate.com","mohit.pathak@seagate.com","mohamadarafat.abdulraheem@seagate.com","mohamad.chaarawi@intel.com","mnizamshah.azzudinshah@seagate.com","mlcurry@sandia.gov","mingjin.w.wang@seagate.com","ming.yik@seagate.com","milind.naik@seagate.com","michelle.e.langan@seagate.com","michele.fagan@seagate.com","mehul.joshi@seagate.com","mazhar.inamdar@seagate.com","mayur.dharmik@seagate.com","maxim.malezhin@seagate.com","max.medved@seagate.com","matthew.halcomb@wwt.com","matt.james@seagate.com","mason.swarr@seagate.com","markidis@kth.se","mark.wiggins@seagate.com","mark.sprouse@dcblox.com","mark.a.penas@seagate.com","mariyappan.ponnusamy@seagate.com","manueljrturao.cabusas@seagate.com","manoj.patil@seagate.com","mandar.sawant@seagate.com","mandar.sabhapatikar@seagate.com","mandar.joshi@seagate.com","mandar.d.joshi@seagate.com","mahima.gupta@seagate.com","mahesh.agarkar@seagate.com","mahendra.shinde@seagate.com","madhura.mande@seagate.com","madhav.vemuri@seagate.com","lynette.sy.neo@seagate.com","luhchyuan.lau@seagate.com","liping.ding@seagate.com","linqiang.luo@seagate.com","lianhoo.tan@seagate.com","liang.gan@seagate.com","lenin.jegatheesan@seagate.com","ladislav.hudec@seagate.com","konstantin.nekrasov@seagate.com","kishan.nayak@seagate.com","kishan.gelli@seagate.com","kiran.mangalore@seagate.com","kimchui.lee@seagate.com","kianwee.tan@seagate.com","kevin.t.james@seagate.com","kevin.a.price@seagate.com","ketan.arlulkar@seagate.com","ken.haugen@seagate.com","kavya.motwani@seagate.com","kaustubh.deorukhkar@seagate.com","kathy.chun@seagate.com","karun.sharma@seagate.com","kapil.jinna@seagate.com","kanchan.chaudhari@seagate.com","kalyan.dandu@seagate.com","kalpesh.chhajed@seagate.com","ka.naidu@seagate.com","jyoti.baral@seagate.com","justin.woo@seagate.com","junichi.hyodo@seagate.com","jugal.patil@seagate.com","josiah.gelsinger@seagate.com","josh.z.li@seagate.com","joseph.rettinghouse@seagate.com","joseph.maniaci@seagate.com","johnbent@gmail.com","john.forgan@seagate.com","john.callender@seagate.com","john.bent@seagate.com","johann.lombardi@intel.com","jin.m.li@seagate.com","jiahui.sun@seagate.com","jerry.l.zhen@seagate.com","jeetandra.kella@seagate.com","jayshree.more@seagate.com","jaydeep.mohite@seagate.com","jayashree.thite@seagate.com","jay.jaiantilal@seagate.com","james.mayfield@seagate.com","james.cleverley@seagate.com","jacques-charles.lafoucriere@cea.fr","ivy.guo@seagate.com","ivan.tishchenko@seagate.com","ivan.poddubnyy@seagate.com","ivan.alekhin@seagate.com","isaacyizhe.teoh@seagate.com","inghong.kong@seagate.com","indrajit.zagade@seagate.com","iman.anvari@seagate.com","igor.pivovarov@seagate.com","ian.davies@seagate.com","i.panourgias@epcc.ed.ac.uk","hua.huang@seagate.com","hemant.raut@seagate.com","harshitkumar.jain@seagate.com","harrison.seow@seagate.com","haibing.ma@seagate.com","gregory.touretsky@seagate.com","graham.d.ferris@seagate.com","gowthaman.chinnathambi@seagate.com","girard.goder@seagate.com","ginann.c.cowen@seagate.com","gheewooi.ong@seagate.com","gaurav.gaur@seagate.com","gaurav.chaudhari@seagate.com","gary.phillips@seagate.com","ganesan.umanesan@seagate.com","gabriel.ruiz@seagate.com","fred.zellinger@seagate.com","evgeniy.brazhnikov@seagate.com","evelyn.godwyn@seagate.com","erik.d.salo@seagate.com","eduard.aleksandrov@seagate.com","earl.dodd@wwt.com","don.r.bloyer@seagate.com","dmitriy.chumak@seagate.com","dmitrii.surnin@seagate.com","divyam.singla@seagate.com","divya.kachhwaha@seagate.com","dipendra.m.bagchee@seagate.com","dinesh.mohanakrishnan@seagate.com","dimitry.didenko@seagate.com","dhananjay.dandapat@seagate.com","deepak.choudhary@seagate.com","debasmita.samaddar@ukaea.uk","david.white@seagate.com","dattaprasad.govekar@seagate.com","daniel.lerner@seagate.com","dahlan.saragih@seagate.com","d.pleiter@fz-juelich.de","cornel.crisan@atos.net","clinton.l.rowe@seagate.com","christopher.haine@hpe.com","christopher.bishop@seagate.com","christina.ku@seagate.com","chris.maio@seagate.com","chong.li@seagate.com","chinfong.hee@seagate.com","chiawhueoy.chin@seagate.com","chetan.srivastava@seagate.com","chetan.deshmukh@seagate.com","cheryl.roberts@seagate.com","cheewai.lum@seagate.com","cheesiong.toh@seagate.com","charles.kunkel@seagate.com","cary.dickens@seagate.com","carrie.harris@seagate.com","bryan.j.baker@seagate.com","bruno.j.masson@seagate.com","brinkman@uni-mainz.de","brian.lavash@seagate.com","brian.ch.chen@seagate.com","brian.a.resch@seagate.com","bikrant.singh@seagate.com","bhavin.katira@seagate.com","bhargava.sreepathi@seagate.com","bhairavi.alurkar@seagate.com","beilei.w.wang@seagate.com","basavaraj.kirunge@seagate.com","bansidhar.soni@seagate.com","balaji.ramachandran@seagate.com","azahar.khan@seagate.com","atul.deshmukh@seagate.com","atita.shirwaikar@seagate.com","ashwini.borse@seagate.com","ashwath.gundepally@seagate.com","ashish.dhavalikar@seagate.com","archana.patil@seagate.com","archana.limaye@seagate.com","arati.kulkarni@seagate.com","apurwa.mohite@seagate.com","anudeep.kankipati@seagate.com","AnneMarie.DeBoard@seagate.com","ankit.yadav@seagate.com","anjali.somwanshi@seagate.com","anirudh.n.joshi@seagate.com","anilkumar.sindhi@seagate.com","andriy.tkachuk@seagate.com","andrey.kononykhin@seagate.com","andrei.zheregelia@seagate.com","anatoliy.bilenko@seagate.com","anagha.g.latne@seagate.com","anagha.deshmukh@seagate.com","amol.shinde@seagate.com","amol.j.kongre@seagate.com","amit.kumar@seagate.com","amit.kapil@seagate.com","alphonsus.kh.kwok@seagate.com","alexander.voronov@seagate.com","alexander.sukhachev@seagate.com","alan.gilda@seagate.com","ajinkya.dhumal@seagate.com","ajay.srivastava@seagate.com","ajay.shingare@seagate.com","ajay.paratmandali@seagate.com","agnieszka.zielinska@seagate.com","abhishek.saha@seagate.com","abhilekh.mustapure@seagate.com","abdul.habeeb@seagate.com","charles.lafoucriere@cea.fr","daniar@uchicago.edu","dbonnie@lanl.gov","dev@tiger-technology.com","don.molaro@wwt.com","ftessier@cscs.ch","furuta.tomonori@fujitsu.com","gunawi@gmail.com","jeff.fonke@wwt.com","lposton@nvidia.com","lucy.gonzalez@arm.com","m.weiland@epcc.ed.ac.uk","miyamoto.kouki@fujitsu.com","philippe.deniel@cea.fr","rich.harper@wwt.com","shail.manjrekar@gmail.com","shaun.de-witt@ukaea.com","sunny.raskar@msystechnologies.com","theo@stillwater-sc.com","tiago.quintino@ecmwf.int","sujata.taware@seagate.com","snehal.virkud@seagate.com","terry.harper@intel.com","drew.stern@nb.com","sati.viensiri@seagate.com","pedro.fernandez@seagate.com","rose.hiu@seagate.com","yoram.novick@gmail.com","pnicolas@coldago.com","navneet479@yahoo.co.in","patrick.bickley@seagate.com","gilbertsosi@gmail.com","mondpl@gmail.com","thomas.doran@seagate.com","pupmaison@neuf.fr","dh0253@yahoo.com","eng.amr214@yahoo.com","boyelroyjetson@yahoo.com","src223@gmail.com","drived@yandex.ru","kimchui.lee@gmail.com","peter.r.janik@seagate.com","renee.wilson@seagate.com","chuan.zhong@seagate.com","eddietolleson21@gmail.com","discoverypark1026@gmail.com","daniar@uchicago.edu","dbmuse@gmail.com","chris.cramer@seagate.com","jazztan1986@gmail.com","rrai@msystechnologies.com","ffrankenstein920@gmail.com","Jak12445@mail.ru","krn@krn.dk","kianchye.tan@seagate.com","lamarahenderson@gmail.com","SECRETARIA3134@GMAIL.COM","georgewu@qnap.com","theminimidistudio@gmail.com","ganliang0122@gmail.com","robbie.cazzaniga@gmail.com","j8862j@naver.com","474trizmo@gmail.com","torreschristina0541@gmail.com","hakimwong71@gmail.com","chintumoturu106@gmail.com","tovijayn@yahoo.com","Jeovalima54@gmail.com","lemperezgeoffre@gmail.com","gwaschke@videotron.ca","adeeba.rashid@marriott.com","mmichaelhooper208@gmail.com","abdulla.fayez@believerit.com","Lior.reinitz@gmail.com","bittorrentsoftware91@gmail.com","yasin@sahinyapi.com.tr","stevencheok@gmail.com","vitaliivladimirivich@gmail.com","julialima.jbl@gmail.com","matthew@tldus.com"])
print(len(newsletter_feb))
newsletter = [(sep_date,429),(oct_date,459),(nov_date,477),(dec_date,newsletter_members_dec),(last_date,newsletter_members_dec)]
newsletter = [(feb_date,newsletter_feb)]
add_stats(newsletter,newsletter_key)
print(ps.get_values_as_numbers(repo,newsletter_key))
def update_webinar():
# adding the webinar metrics for executive report
webinar_nov=set(["Andrei Zheregelia","Andrew List","Andriy Tkachuk","Anitoliy Bilenko","Anthony Toccco","Ben Wason","Charles Kunkel","Chetan Deshmukh","Chetan Kumar","Clay Curry","Daniar Kurniawan","Dima.c","Dmitri Sandler","Dmytro Podgornyi","Eduard Aleksandrov","Gary Phillips","Guy Carbonneau","Hanesan Umanesan","Igor Pivovarov","Iman Anvari","Ivan Alekhin","Jason Sliger-Sparks","Jjohn Carrier","Julia Rubtsov","Kalpesh Chhajed","Kaustubh Suresh Deorukhkar","Ken Haugen","Ketan Anil Arlulkar","Konstatin Nekrasov","Lance Blumberg","Madhavrao Vemuri","Mark Jedraszek","Maxim Malezhin","Max Medved","Mehul Joshi","Nicholas Krauter","Nigel Hart","Nikita Danilov","Patrick Raaf","Paul Woods","Philippe Nicolas","Phil Ruff","Raydon Gordon","Ricardo Alvarez-Miranda","Sachin Punadikar","Sailesh Manjrekar","Sai Narasimhamurthy","Sarang Sawant","Serkay Olmez","Shankar More","Shiji Zhang","Swapnil Khandare","Taro Iwata","Ujjwal Lanjewar"])
webinar_dec=set(["Andriy Tkachuk","Anthony Tocco","Charles Kunkel","Daniar Kurniawan","Dan Olster","Ganesan Umanesan","Gary Phillips","Guy Carbonneau","Ivan Poddubnyy","Justin Rackowski","Nicolas Krauter","Nigel Hart","Paul Benn","Praveen Viraraghavan","Rajesh Nambiar","Ricardo Alvarez-miranda","Sachin Punadikar","Sarang Sawant","Shankar More","Shiji Zhang","Swapnil Khandare","Trend Geerdes","Ujjwal Lanjewar","Walter Lopatka",])
webinar_jan=set(["Unknown1","Anatoliy Bilenko","Andrea Chamorro","Andriy Tkachuk","Anthony Tocco","Unknown2","Charles Kunkel","Chetan Kumar","Chirs Cramer","Erin Foley","Gabe Wham","Gary Grider","Gregory Touretsky","Iman Anvari","Joseph Rebovich","Justin Rackowski","Keith Pine","Ken Haugen","Ketan Anil Arlulkar","Madhavrao Vemuri","Amandar Sawant","Mark Jedraszek","Mark Sprouse","Matthew Halcomb","Matthew L Curry Sandia","Max","Mehul Joshi","Meng Wang","Mike Sevilla","Muhul Malhotra","Nedko Amaudov","Oded Kellner","Paul Kusbel","Pedro Fernandez","Pritesh Pawar","Priyanka Borawake","Quinn D Mitchell","Rajesh Bhalerao","Ricardo Alvarez-Miranda","Robert Pechman","Rohan Puri","Sachin Punadikar","Sai Narasimhamurthy","Sarang Sawant","Shailesh","Shankar More","Sharad Mehrotra","Shlomi Avihou","Shreya Karmakar","Shrihari Waskar","Unknown","Sridbar Dubhaka","Stephane Thiell","Swapril Khandare","Tong Shi","Ujjwal Lanjewar","Venky P","Vijay Nanjunda Swamy","Vikram","Vojtech Juranek","Walkter Lopatka","Ziv","Theodore Omtzigt","Rajkumar Patel","Anjinkya Deshpande","Anatoliy Bilenko","Chetan Deshmukh","Henry Newman","Paul Benn","Paul Woods","Kyle Lamb"])
webinar_feb=set(["Ashwin Agrawal","Jean Luca Bez","Rex Tanakit","Samuel Spencer","Shailesh Vaidya","Tripti Srivastava","Abraham Checkoway","Abhijeet Dhumal","Anatoliy Bilenko","Anthony Tocco","Antoine Le Bideau","Basavaraj Kirunge","BK Singh","Branislav Radovanovic","Charles Kunkel","Chetan Deshmukh","Carlos Thomaz","Dan Olster","Debashish Pal","Geert Wenes","Gary Grider","Gary Lowell","Jason Sliger-Sparks","Jean-Thomas","Justin Rackowski","Justin Woo","Kalpesh Chhajed","Keith Pine","Ken Haugen","Ketan Anil Arlulkar","Kiran Mangalore","Liang Gan","Madhavrao Vemuri","Mandar Sawant","Mark Jedraszek","Mehul Joshi","Mukul Malhotra","Nicolau Manubens","Nigel Hart","Nilesh Navale","Parag Joshi","Parks Fields","Paul Benn","Paul Woods","Peyton McNully","Prudence Huang","Philippe Nicolas","Pranali Ramdas Tirkhunde","Ryan Cassidy","Rob Wilson","Robert Read","Rohan Puri","Ryan Tyler","Sarang Sawant","Serkay Olmez","Shankar More","Seth Kindley","Swarajya Pendharkar","Sumedh Kulkarni","Sven Breuner","Sven Breuner","Theodore Omtzigt","Tim Coullter","Ravi Tripathi","Tushar Tarkas","Ujjwal Lanjewar","Venky P","Walter Lopatka","Earl Dodd","Wendell Wenjen","Weikuan Yu","George Zhi Qiao",])
jan_date='2021-01-06'
feb_date='2021-02-06'
dec_date='2020-12-21'
nov_date='2020-11-03'
#print(len(webinar_nov),len(webinar_dec),len(webinar_jan))
#webinar = [(nov_date,webinar_nov),(dec_date,webinar_dec),(jan_date,webinar_jan)]
webinar = [(feb_date,webinar_feb),(dec_date,webinar_dec),(jan_date,webinar_jan),(nov_date,webinar_nov)]
add_stats(webinar,webinar_key)
print(ps.get_values_as_numbers(repo,webinar_key))
def update_integrations():
# add the integrations metric for executive report
integrations = [(sep_date,0),(oct_date,1),(nov_date,1),(dec_date,6),(last_date,6)]
add_stats(integrations,integrations_key)
print(ps.get_values_as_numbers(repo,integrations_key))
def update_external(repo):
def update_external_single(repo,date,original,removals,key):
if original is None:
return
for r in removals:
try:
original -= r
except TypeError:
pass # can't subtract None from a set
ps.add_stat(repo=repo,date=date,stat=key,value=original)
# so we used to double count hackathon folks as external folks
# we fixed that but now the external counts suddenly unexpectedly dropped
# let's fix that in the historical record
# actually this should be easy
# just iterate through every date and subtract hackathon folks from external folks and resave the difference as external
#external participants
for date in dates:
ec = ps.get_values(repo, ec_key,[date])[0]
hc = ps.get_values(repo, hc_key,[date])[0]
ep = ps.get_values(repo, ep_key,[date])[0]
hp = ps.get_values(repo, hp_key,[date])[0]
eup = {'u-u-h', 'jayeshbadwaik'}
update_external_single(repo=repo,date=date,original=ec,removals=[hc], key=ec_key)
update_external_single(repo=repo,date=date,original=ep,removals=[hp,eup], key=ep_key)
def manually_add_historical_committers():
# for external committers also go through and manually add them in the early months
sep_committers=set(['cotigao','daniarherikurniawan','jan--f'])
oct_committers=sep_committers | set(['raikrahul'])
committers = [(jun_date,set()),(sep_date,sep_committers),(oct_date,oct_committers),('2020-12-20',oct_committers),('2020-12-21',oct_committers),('2020-12-23',oct_committers)]
add_stats(committers,ec_key)
#print(ps.get_values_as_numbers(repo, 'external_participants'))
def clean_external():
print(dates)
for r in ps.get_repos():
print("Will clean external committers and participants for %s" % r)
update_external(r)
print(ps.get_values_as_numbers(r, ec_key))
print(ps.get_values_as_numbers(r, ep_key))
print(ps.get_values(repo, ec_key))
update_webinar()
#print(ps.get_dates(repo))
update_newsletter()
ps.persist()
#manually_add_historical_committers()
#update_slack()
#update_webinar()
#update_integrations()
#for i in [3,4,5,10,11,12,20]:
# print(ps.get_dates('GLOBAL')[i], '->', ps.get_values_as_numbers('GLOBAL',webinar_key)[i])
def clean_external_participants():
ep1=ps.stats[repo][dates[4]][ep_key]
ep2=ps.stats[repo][dates[5]][ep_key]
print(len(ep1),len(ep2))
print(dates[4],ep1)
print(dates[5],ep2)
print(ep1-ep2)
ps.stats[repo][dates[5]][ep_key]=ep2|{'jan--f'}
ps.persist()
print(ps.get_values_as_numbers(repo, ep_key))
ep3=ps.stats[repo][dates[-1]][ep_key]
ep4=ps.stats[repo][dates[-2]][ep_key]
print(ep4-ep3)
#clean_external_participants()
def get_logins(Type):
folks=set()
people=cortx_community.CortxCommunity()
for p in people.values():
if p.type == Type:
folks.add(p.login)
return folks
# we once mistakenly characterized a few team folks as innersource folks
def clean_innersource(repo,Type,folks):
key='innersource_%s'%Type
for d in ps.get_dates(repo):
try:
values=ps.stats[repo][d][key]
except KeyError:
continue # some keys didn't always exist
if isinstance(values,set):
bad = [v for v in values if v not in folks]
if len(bad)>0:
print("Need to remove",bad,"from %s:%s on %s" % (repo,key,d))
new_values = values - set(bad)
print("Will reduce from %d to %d" % (len(values),len(new_values)))
ps.stats[repo][d][key]=new_values
def print_values(repo,key,dates=None):
for v in ps.get_values(repo,key,dates):
try:
print(len(v),sorted(v))
except:
print(v)
def bulk_clean():
for r in ps.get_repos():
for t in ('participants','committers'):
clean_innersource(repo=r,Type=t,folks=folks)
print(r,t,ps.get_values_as_numbers(r,'innersource_%s'%t))
folks=get_logins('Innersource')
print(folks)
#bulk_clean()
#ps.persist()
#print("all cleaned?!?!")
def clean_ip():
repo='GLOBAL'
key='innersource_participants'
dates=ps.get_dates(repo)
good_values=ps.get_values(repo=repo,key=key,dates=[dates[5]])[0]
for d in [dates[2],dates[3],dates[4]]:
ps.add_stat(date=d,repo=repo,stat=key,value=good_values)
print_values(repo,key,dates)
print(len(values),values)
def clean_ic():
repo='GLOBAL'
key='innersource_committers'
clean_innersource(repo,'committers',folks)
#clean_ip()
def print_innersource(Key):
for r in ps.get_repos():
print(r, ps.get_values_as_numbers(repo=r,key=Key))
clean_ic()
print_innersource('innersource_committers')
ps.persist()
compare_fields(ps=ps,repo='cortx',field='innersource_committers',verbose=False)
compare_fields(ps=ps,repo='cortx',field='external_email_addresses',verbose=False)
targets=['issues_closed_ave_age_in_s','issues_closed']
for target in targets:
for r in ['GLOBAL','cortx-ha','cortx-hare']:
print("%s %s -> %d " % (r, target, ps.stats[r]['2020-12-29'][target]))
check_scan_progress('2021-01-06',ps)
(a,b)=ps.get_latest('cortx-hare')
a['issues_open']
b
ps.stats['GLOBAL']['2021-01-02']['stars']
# this block is a one-time thing to add historical data from before we automated the scraping
d1={'innersource_participants' : 5, 'pull_requests_external' : 0,
'external_participants' : 0,
'watchers' : 34, 'stars' : 19, 'forks' : 13, 'views_unique_14_days' : 106,
'clones_count_14_days' : 38, 'clones_unique_14_days' : 4,
'seagate_blog_referrer_uniques' : 0, 'seagate_referrer_uniques' : 0,
'downloads_vms' : 0}
d1_date='2020-05-19'
d2={'innersource_participants' : 8, 'pull_requests_external' : 0,
'external_participants' : 0,
'watchers' : 69, 'stars' : 52, 'forks' : 42,
'views_unique_14_days' : 86,
'clones_count_14_days' : 15, 'clones_unique_14_days' : 6,
'seagate_blog_referrer_uniques' : 0, 'seagate_referrer_uniques' : 0,
'downloads_vms' : 0}
d2_date='2020-07-06'
d3={'innersource_participants' : 18, 'pull_requests_external' : 1,
'external_participants' : 0,
'watchers' : 62, 'stars' : 116, 'forks' : 31,
'views_unique_14_days' : 1817,
'clones_count_14_days' : 468, 'clones_unique_14_days' : 224,
'seagate_blog_referrer_uniques' : 0, 'seagate_referrer_uniques' : 0,
'downloads_vms' : 130}
d3_date='2020-10-07'
d4={'innersource_participants' : 18, 'pull_requests_external' : 4,
'external_participants' : 0,
'watchers' : 65, 'stars' : 159, 'forks' : 45,
'views_unique_14_days' : 817,
'clones_count_14_days' : 1851, 'clones_unique_14_days' : 259,
'seagate_blog_referrer_uniques' : 0, 'seagate_referrer_uniques' : 0,
'downloads_vms' : 363}
d4_date='2020-11-03'
print(d1)
#ps.add_stats(date=d1_date,repo='GLOBAL',stats=d1)
#ps.add_stats(date=d2_date,repo='GLOBAL',stats=d2)
#ps.add_stats(date=d3_date,repo='GLOBAL',stats=d3)
#ps.add_stats(date=d4_date,repo='GLOBAL',stats=d4)
ps.get_dates('GLOBAL')
```
| github_jupyter |
```
from pydrive.auth import GoogleAuth
from pydrive.drive import GoogleDrive
from google.colab import auth
from oauth2client.client import GoogleCredentials
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
import torch
import torchaudio
```
# Part 1 : contrastive predictive coding
Contrastive Predictive Coding (CPC) is a method of unsupervised training for speech models. The idea behind it is pretty simple:
1. The raw audio wave is passed through a convolutional network: the ```encoder```
2. Then, the encoder's output is given to a recurrent network the ```context```
3. A third party network, the ```prediction_network``` will try to predict the future embeddings of the encoder using the output of the context network.
In order to avoid a collapse to trivial solutions, the prediction_network doesn't try to reconstruct the future features. Instead, using the context output $c_t$ at time $t$ it is trained to discriminate the real encoder representatioin $g_{t+k}$ at time $t+k$ from several other features $(g_n)_n$ taken elsewhere in the batch. Thus the loss becomes:
\\[ \mathcal{L}_c = - \frac{1}{K} \sum_{k=1}^K \text{Cross_entropy}(\phi_k(c_t), g_{t+k}) \\]
Or:
\\[ \mathcal{L}_c = - \frac{1}{K} \sum_{k=1}^K \log \frac{ \exp\left(\phi_k(c_t)^\top g_{t+k}\right) }{ \sum_{\mathbf{n}\in\mathcal{N}_t} \exp\left(\phi_k(c_t)^\top g_n\right)} \\]
Where:
* $\phi_k$ is the prediction network for the kth timestep
* $\mathcal{N}_t$ is the set of all negative examples sampled for timestep $t$
## Exercice 1 : Building the model
In this exercise, we will build and train a small CPC model using the repository CPC_audio.
The code below loads a context and an encoder newtorks.
```
%cd /content/CPC_audio
from cpc.model import CPCEncoder, CPCAR
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
DIM_ENCODER=256
DIM_CONTEXT=256
KEEP_HIDDEN_VECTOR=False
N_LEVELS_CONTEXT=1
CONTEXT_RNN="LSTM"
N_PREDICTIONS=12
LEARNING_RATE=2e-4
N_NEGATIVE_SAMPLE =128
encoder = CPCEncoder(DIM_ENCODER).to(device)
context = CPCAR(DIM_ENCODER, DIM_CONTEXT, KEEP_HIDDEN_VECTOR, 1, mode=CONTEXT_RNN).to(device)
# Several functions that will be necessary to load the data later
from cpc.dataset import findAllSeqs, AudioBatchData, parseSeqLabels
SIZE_WINDOW = 20480
BATCH_SIZE=8
def load_dataset(path_dataset, file_extension='.flac', phone_label_dict=None):
data_list, speakers = findAllSeqs(path_dataset, extension=file_extension)
dataset = AudioBatchData(path_dataset, SIZE_WINDOW, data_list, phone_label_dict, len(speakers))
return dataset
```
Now build a new class, ```CPCModel``` which will
```
class CPCModel(torch.nn.Module):
def __init__(self,
encoder,
AR):
super(CPCModel, self).__init__()
self.gEncoder = encoder
self.gAR = AR
def forward(self, batch_data):
encoder_output = self.gEncoder(batch_data)
#print(encoder_output.shape)
# The output of the encoder data does not have the good format
# indeed it is Batch_size x Hidden_size x temp size
# while the context requires Batch_size x temp size x Hidden_size
# thus you need to permute
context_input = encoder_output.permute(0, 2, 1)
context_output = self.gAR(context_input)
#print(context_output.shape)
return context_output, encoder_output
```
Let's test your code !
```
audio = torchaudio.load("/content/train_data/831/130739/831-130739-0048.flac")[0]
audio = audio.view(1, 1, -1)
cpc_model = CPCModel(encoder, context).to(device)
context_output, encoder_output = cpc_model(audio.to(device))
```
## Exercise 2 : CPC loss
We will define a class ```CPCCriterion``` which will hold the prediction networks $\phi_k$ defined above and perform the classification loss $\mathcal{L}_c$.
a) In this exercise, the $\phi_k$ will be a linear transform, ie:
\\[ \phi_k(c_t) = \mathbf{A}_k c_t\\]
Using the class [torch.nn.Linear](https://pytorch.org/docs/stable/nn.html#torch.nn.Linear), define the transformations $\phi_k$ in the code below and complete the function ```get_prediction_k``` which computes $\phi_k(c_t)$ for a given batch of vectors $c_t$.
b) Using both ```get_prediction_k``` and ```sample_negatives``` defined below, write the forward function which will take as input two batches of features $c_t$ and $g_t$ and outputs the classification loss $\mathcal{L}_c$ and the average acuracy for all predictions.
```
# Exercice 2: write the CPC loss
# a) Write the negative sampling (with some help)
# ERRATUM: it's really hard, the sampling will be provided
class CPCCriterion(torch.nn.Module):
def __init__(self,
K,
dim_context,
dim_encoder,
n_negative):
super(CPCCriterion, self).__init__()
self.K_ = K
self.dim_context = dim_context
self.dim_encoder = dim_encoder
self.n_negative = n_negative
self.predictors = torch.nn.ModuleList()
for k in range(self.K_):
# TO COMPLETE !
# A affine transformation in pytorch is equivalent to a nn.Linear layer
# To get a linear transformation you must set bias=False
# input dimension of the layer = dimension of the encoder
# output dimension of the layer = dimension of the context
self.predictors.append(torch.nn.Linear(dim_context, dim_encoder, bias=False))
def get_prediction_k(self, context_data):
#TO COMPLETE !
output = []
# For each time step k
for k in range(self.K_):
# We need to compute phi_k = A_k * c_t
phi_k = self.predictors[k](context_data)
output.append(phi_k)
return output
def sample_negatives(self, encoded_data):
r"""
Sample some negative examples in the given encoded data.
Input:
- encoded_data size: B x T x H
Returns
- outputs of size B x (n_negative + 1) x (T - K_) x H
outputs[:, 0, :, :] contains the positive example
outputs[:, 1:, :, :] contains negative example sampled in the batch
- labels, long tensor of size B x (T - K_)
Since the positive example is always at coordinates 0 for all sequences
in the batch and all timestep in the sequence, labels is just a tensor
full of zeros !
"""
batch_size, time_size, dim_encoded = encoded_data.size()
window_size = time_size - self.K_
outputs = []
neg_ext = encoded_data.contiguous().view(-1, dim_encoded)
n_elem_sampled = self.n_negative * window_size * batch_size
# Draw nNegativeExt * batchSize negative samples anywhere in the batch
batch_idx = torch.randint(low=0, high=batch_size,
size=(n_elem_sampled, ),
device=encoded_data.device)
seq_idx = torch.randint(low=1, high=time_size,
size=(n_elem_sampled, ),
device=encoded_data.device)
base_idx = torch.arange(0, window_size, device=encoded_data.device)
base_idx = base_idx.view(1, 1, window_size)
base_idx = base_idx.expand(1, self.n_negative, window_size)
base_idx = base_idx.expand(batch_size, self.n_negative, window_size)
seq_idx += base_idx.contiguous().view(-1)
seq_idx = torch.remainder(seq_idx, time_size)
ext_idx = seq_idx + batch_idx * time_size
neg_ext = neg_ext[ext_idx].view(batch_size, self.n_negative,
window_size, dim_encoded)
label_loss = torch.zeros((batch_size, window_size),
dtype=torch.long,
device=encoded_data.device)
for k in range(1, self.K_ + 1):
# Positive samples
if k < self.K_:
pos_seq = encoded_data[:, k:-(self.K_-k)]
else:
pos_seq = encoded_data[:, k:]
pos_seq = pos_seq.view(batch_size, 1, pos_seq.size(1), dim_encoded)
full_seq = torch.cat((pos_seq, neg_ext), dim=1)
outputs.append(full_seq)
return outputs, label_loss
def forward(self, encoded_data, context_data):
# TO COMPLETE:
# Perform the full cpc criterion
# Returns 2 values:
# - the average classification loss avg_loss
# - the average classification acuracy avg_acc
# Reminder : The permuation !
encoded_data = encoded_data.permute(0, 2, 1)
# First we need to sample the negative examples
negative_samples, labels = self.sample_negatives(encoded_data)
# Then we must compute phi_k
phi_k = self.get_prediction_k(context_data)
# Finally we must get the dot product between phi_k and negative_samples
# for each k
#The total loss is the average of all losses
avg_loss = 0
# Average acuracy
avg_acc = 0
for k in range(self.K_):
B, N_sampled, S_small, H = negative_samples[k].size()
B, S, H = phi_k[k].size()
# As told before S = S_small + K. For segments too far in the sequence
# there are no positive exmples anyway, so we must shorten phi_k
phi = phi_k[k][:, :S_small]
# Now the dot product
# You have several ways to do that, let's do the simple but non optimal
# one
# pytorch has a matrix product function https://pytorch.org/docs/stable/torch.html#torch.bmm
# But it takes only 3D tensors of the same batch size !
# To begin negative_samples is a 4D tensor !
# We want to compute the dot product for each features, of each sequence
# of the batch. Thus we are trying to compute a dot product for all
# B* N_sampled * S_small 1D vector of negative_samples[k]
# Or, a 1D tensor of size H is also a matrix of size 1 x H
# Then, we must view it as a 3D tensor of size (B* N_sampled * S_small, 1, H)
negative_sample_k = negative_samples[k].view(B* N_sampled* S_small, 1, H)
# But now phi and negative_sample_k no longer have the same batch size !
# No worries, we can expand phi so that each sequence of the batch
# is repeated N_sampled times
phi = phi.view(B, 1,S_small, H).expand(B, N_sampled, S_small, H)
# And now we can view it as a 3D tensor
phi = phi.contiguous().view(B * N_sampled * S_small, H, 1)
# We can finally get the dot product !
scores = torch.bmm(negative_sample_k, phi)
# Dot_product has a size (B * N_sampled * S_small , 1, 1)
# Let's reorder it a bit
scores = scores.reshape(B, N_sampled, S_small)
# For each elements of the sequence, and each elements sampled, it gives
# a floating score stating the likelihood of this element being the
# true one.
# Now the classification loss, we need to use the Cross Entropy loss
# https://pytorch.org/docs/master/generated/torch.nn.CrossEntropyLoss.html
# For each time-step of each sequence of the batch
# we have N_sampled possible predictions.
# Looking at the documentation of torch.nn.CrossEntropyLoss
# we can see that this loss expect a tensor of size M x C where
# - M is the number of elements with a classification score
# - C is the number of possible classes
# There are N_sampled candidates for each predictions so
# C = N_sampled
# Each timestep of each sequence of the batch has a prediction so
# M = B * S_small
# Thus we need an input vector of size B * S_small, N_sampled
# To begin, we need to permute the axis
scores = scores.permute(0, 2, 1) # Now it has size B , S_small, N_sampled
# Then we can cast it into a 2D tensor
scores = scores.reshape(B * S_small, N_sampled)
# Same thing for the labels
labels = labels.reshape(B * S_small)
# Finally we can get the classification loss
loss_criterion = torch.nn.CrossEntropyLoss()
loss_k = loss_criterion(scores, labels)
avg_loss+= loss_k
# And for the acuracy
# The prediction for each elements is the sample with the highest score
# Thus the tensors of all predictions is the tensors of the index of the
# maximal score for each time-step of each sequence of the batch
predictions = torch.argmax(scores, 1)
acc_k = (labels == predictions).sum() / (B * S_small)
avg_acc += acc_k
# Normalization
avg_loss = avg_loss / self.K_
avg_acc = avg_acc / self.K_
return avg_loss , avg_acc
```
Don't forget to test !
```
audio = torchaudio.load("/content/train_data/831/130739/831-130739-0048.flac")[0]
audio = audio.view(1, 1, -1)
cpc_criterion = CPCCriterion(N_PREDICTIONS, DIM_CONTEXT,
DIM_ENCODER, N_NEGATIVE_SAMPLE).to(device)
context_output, encoder_output = cpc_model(audio.to(device))
loss, avg = cpc_criterion(encoder_output,context_output)
```
## Exercise 3: Full training loop !
You have the model, you have the criterion. All you need now are a data loader and an optimizer to run your training loop.
We will use an Adam optimizer:
```
parameters = list(cpc_criterion.parameters()) + list(cpc_model.parameters())
optimizer = torch.optim.Adam(parameters, lr=LEARNING_RATE)
```
And as far as the data loader is concerned, we will rely on the data loader provided by the CPC_audio library.
```
dataset_train = load_dataset('/content/train_data')
dataset_val = load_dataset('/content/val_data')
data_loader_train = dataset_train.getDataLoader(BATCH_SIZE, "speaker", True)
data_loader_val = dataset_train.getDataLoader(BATCH_SIZE, "sequence", False)
```
Now that everything is ready, complete and test the ```train_step``` function below which trains the model for one epoch.
```
def train_step(data_loader,
cpc_model,
cpc_criterion,
optimizer):
avg_loss = 0
avg_acc = 0
n_items = 0
for step, data in enumerate(data_loader):
x,y = data
bs = len(x)
optimizer.zero_grad()
context_output, encoder_output = cpc_model(x.to(device))
loss , acc = cpc_criterion(encoder_output, context_output)
loss.backward()
n_items+=bs
avg_loss+=loss.item()*bs
avg_acc +=acc.item()*bs
avg_loss/=n_items
avg_acc/=n_items
return avg_loss, avg_acc
```
## Exercise 4 : Validation loop
Now complete the validation loop.
```
def validation_step(data_loader,
cpc_model,
cpc_criterion):
avg_loss = 0
avg_acc = 0
n_items = 0
for step, data in enumerate(data_loader):
x,y = data
bs = len(x)
context_output, encoder_output = cpc_model(x.to(device))
loss , acc = cpc_criterion(encoder_output, context_output)
n_items+=bs
avg_loss+=loss.item()*bs
avg_acc+=acc.item()*bs
avg_loss/=n_items
avg_acc/=n_items
return avg_loss, avg_acc
```
## Exercise 5: Run everything
```
def run(train_loader,
val_loader,
cpc_model,
cpc_criterion,
optimizer,
n_epochs):
for epoch in range(n_epochs):
print(f"Running epoch {epoch+1} / {n_epochs}")
avg_loss_train, avg_acc_train = train_step(train_loader, cpc_model, cpc_criterion, optimizer)
print("----------------------")
print(f"Training dataset")
print(f"- average loss : {avg_loss_train}")
print(f"- average acuracy : {avg_acc_train}")
print("----------------------")
with torch.no_grad():
cpc_model.eval()
cpc_criterion.eval()
avg_loss_val, avg_acc_val = validation_step(val_loader, cpc_model, cpc_criterion)
print(f"Validation dataset")
print(f"- average loss : {avg_loss_val}")
print(f"- average acuracy : {avg_acc_val}")
print("----------------------")
print()
cpc_model.train()
cpc_criterion.train()
run(data_loader_train, data_loader_val, cpc_model,cpc_criterion,optimizer,1)
```
Once everything is donw, clear the memory.
```
del dataset_train
del dataset_val
del cpc_model
del context
del encoder
```
# Part 2 : Fine tuning
## Exercice 1 : Phone separability with aligned phonemes.
One option to evaluate the quality of the features trained with CPC can be to check if they can be used to recognize phonemes.
To do so, we can fine-tune a pre-trained model using a limited amount of labelled speech data.
We are going to start with a simple evaluation setting where we have the phone labels for each timestep corresponding to a CPC feature.
We will work with a model already pre-trained on English data. As far as the fine-tuning dataset is concerned, we will use a 1h subset of [librispeech-100](http://www.openslr.org/12/).
```
!mkdir checkpoint_data
!wget https://dl.fbaipublicfiles.com/librilight/CPC_checkpoints/not_hub/2levels_6k_top_ctc/checkpoint_30.pt -P checkpoint_data
!wget https://dl.fbaipublicfiles.com/librilight/CPC_checkpoints/not_hub/2levels_6k_top_ctc/checkpoint_logs.json -P checkpoint_data
!wget https://dl.fbaipublicfiles.com/librilight/CPC_checkpoints/not_hub/2levels_6k_top_ctc/checkpoint_args.json -P checkpoint_data
!ls checkpoint_data
%cd /content/CPC_audio
from cpc.dataset import parseSeqLabels
from cpc.feature_loader import loadModel
checkpoint_path = 'checkpoint_data/checkpoint_30.pt'
cpc_model, HIDDEN_CONTEXT_MODEL, HIDDEN_ENCODER_MODEL = loadModel([checkpoint_path])
cpc_model = cpc_model.cuda()
label_dict, N_PHONES = parseSeqLabels('/content/converted_aligned_phones.txt')
dataset_train = load_dataset('/content/train_data', file_extension='.flac', phone_label_dict=label_dict)
dataset_val = load_dataset('/content/val_data', file_extension='.flac', phone_label_dict=label_dict)
data_loader_train = dataset_train.getDataLoader(BATCH_SIZE, "speaker", True)
data_loader_val = dataset_val.getDataLoader(BATCH_SIZE, "sequence", False)
??cpc_model
```
Then we will use a simple linear classifier to recognize the phonemes from the features produced by ```cpc_model```.
### a) Build the phone classifier
Design a class of linear classifiers, ```PhoneClassifier``` that will take as input a batch of sequences of CPC features and output a score vector for each phoneme
```
class PhoneClassifier(torch.nn.Module):
def __init__(self,
input_dim : int,
n_phones : int):
super(PhoneClassifier, self).__init__()
self.linear = torch.nn.Linear(input_dim, n_phones)
def forward(self, x):
return self.linear(x)
```
Our phone classifier will then be:
```
phone_classifier = PhoneClassifier(HIDDEN_CONTEXT_MODEL, N_PHONES).to(device)
```
### b - What would be the correct loss criterion for this task ?
```
loss_criterion = torch.nn.CrossEntropyLoss()
```
To perform the fine-tuning, we will also need an optimization function.
We will use an [Adam optimizer ](https://pytorch.org/docs/stable/optim.html#torch.optim.Adam).
```
parameters = list(phone_classifier.parameters()) + list(cpc_model.parameters())
LEARNING_RATE = 2e-4
optimizer = torch.optim.Adam(parameters, lr=LEARNING_RATE)
```
You might also want to perform this training while freezing the weights of the ```cpc_model```. Indeed, if the pre-training was good enough, then ```cpc_model``` phonemes representation should be linearly separable. In this case the optimizer should be defined like this:
```
optimizer_frozen = torch.optim.Adam(list(phone_classifier.parameters()), lr=LEARNING_RATE)
```
### c- Now let's build a training loop.
Complete the function ```train_one_epoch``` below.
```
def train_one_epoch(cpc_model,
phone_classifier,
loss_criterion,
data_loader,
optimizer):
cpc_model.train()
loss_criterion.train()
avg_loss = 0
avg_accuracy = 0
n_items = 0
for step, full_data in enumerate(data_loader):
# Each batch is represented by a Tuple of vectors:
# sequence of size : N x 1 x T
# label of size : N x T
#
# With :
# - N number of sequence in the batch
# - T size of each sequence
sequence, label = full_data
bs = len(sequence)
seq_len = label.size(1)
optimizer.zero_grad()
context_out, enc_out, _ = cpc_model(sequence.to(device),label.to(device))
scores = phone_classifier(context_out)
scores = scores.permute(0,2,1)
loss = loss_criterion(scores,label.to(device))
loss.backward()
optimizer.step()
avg_loss+=loss.item()*bs
n_items+=bs
correct_labels = scores.argmax(1)
avg_accuracy += ((label==correct_labels.cpu()).float()).mean(1).sum().item()
avg_loss/=n_items
avg_accuracy/=n_items
return avg_loss, avg_accuracy
```
Don't forget to test it !
```
avg_loss, avg_accuracy = train_one_epoch(cpc_model, phone_classifier, loss_criterion, data_loader_train, optimizer_frozen)
avg_loss, avg_accuracy
```
### d- Build the validation loop
```
def validation_step(cpc_model,
phone_classifier,
loss_criterion,
data_loader):
cpc_model.eval()
phone_classifier.eval()
avg_loss = 0
avg_accuracy = 0
n_items = 0
with torch.no_grad():
for step, full_data in enumerate(data_loader):
# Each batch is represented by a Tuple of vectors:
# sequence of size : N x 1 x T
# label of size : N x T
#
# With :
# - N number of sequence in the batch
# - T size of each sequence
sequence, label = full_data
bs = len(sequence)
seq_len = label.size(1)
context_out, enc_out, _ = cpc_model(sequence.to(device),label.to(device))
scores = phone_classifier(context_out)
scores = scores.permute(0,2,1)
loss = loss_criterion(scores,label.to(device))
avg_loss+=loss.item()*bs
n_items+=bs
correct_labels = scores.argmax(1)
avg_accuracy += ((label==correct_labels.cpu()).float()).mean(1).sum().item()
avg_loss/=n_items
avg_accuracy/=n_items
return avg_loss, avg_accuracy
```
### e- Run everything
Test this functiion with both ```optimizer``` and ```optimizer_frozen```.
```
def run(cpc_model,
phone_classifier,
loss_criterion,
data_loader_train,
data_loader_val,
optimizer,
n_epoch):
for epoch in range(n_epoch):
print(f"Running epoch {epoch + 1} / {n_epoch}")
loss_train, acc_train = train_one_epoch(cpc_model, phone_classifier, loss_criterion, data_loader_train, optimizer)
print("-------------------")
print(f"Training dataset :")
print(f"Average loss : {loss_train}. Average accuracy {acc_train}")
print("-------------------")
print("Validation dataset")
loss_val, acc_val = validation_step(cpc_model, phone_classifier, loss_criterion, data_loader_val)
print(f"Average loss : {loss_val}. Average accuracy {acc_val}")
print("-------------------")
print()
run(cpc_model,phone_classifier,loss_criterion,data_loader_train,data_loader_val,optimizer_frozen,n_epoch=10)
```
## Exercise 2 : Phone separability without alignment (PER)
Aligned data are very practical, but un real life they are rarely available. That's why in this excercise we will consider a fine-tuning with non-aligned phonemes.
The model, the optimizer and the phone classifier will stay the same. However, we will replace our phone criterion with a [CTC loss](https://pytorch.org/docs/master/generated/torch.nn.CTCLoss.html).
```
loss_ctc = torch.nn.CTCLoss()
```
Besides, we will use a siglthy different dataset class.
```
%cd /content/CPC_audio
from cpc.eval.common_voices_eval import SingleSequenceDataset, parseSeqLabels, findAllSeqs
path_train_data_per = '/content/per_data/pack_master/1h'
path_val_data_per = '/content/per_data/pack_master/10min'
path_phone_data_per = '/content/per_data/pack_master/10h_phones.txt'
BATCH_SIZE=8
phone_labels, N_PHONES = parseSeqLabels(path_phone_data_per)
data_train_per, _ = findAllSeqs(path_train_data_per, extension='.flac')
dataset_train_non_aligned = SingleSequenceDataset(path_train_data_per, data_train_per, phone_labels)
data_loader_train = torch.utils.data.DataLoader(dataset_train_non_aligned, batch_size=BATCH_SIZE,
shuffle=True)
data_val_per, _ = findAllSeqs(path_val_data_per, extension='.flac')
dataset_val_non_aligned = SingleSequenceDataset(path_val_data_per, data_val_per, phone_labels)
data_loader_val = torch.utils.data.DataLoader(dataset_val_non_aligned, batch_size=BATCH_SIZE,
shuffle=True)
```
### a- Training
Since the phonemes are not aligned, there is no simple direct way to get the classification acuracy of a model. Write and test the three functions ```train_one_epoch_ctc```, ```validation_step_ctc``` and ```run_ctc``` as before but without considering the average acuracy of the model.
```
from cpc.feature_loader import loadModel
checkpoint_path = 'checkpoint_data/checkpoint_30.pt'
cpc_model, HIDDEN_CONTEXT_MODEL, HIDDEN_ENCODER_MODEL = loadModel([checkpoint_path])
cpc_model = cpc_model.cuda()
phone_classifier = PhoneClassifier(HIDDEN_CONTEXT_MODEL, N_PHONES).to(device)
parameters = list(phone_classifier.parameters()) + list(cpc_model.parameters())
LEARNING_RATE = 2e-4
optimizer = torch.optim.Adam(parameters, lr=LEARNING_RATE)
optimizer_frozen = torch.optim.Adam(list(phone_classifier.parameters()), lr=LEARNING_RATE)
import torch.nn.functional as F
def train_one_epoch_ctc(cpc_model,
phone_classifier,
loss_criterion,
data_loader,
optimizer):
cpc_model.train()
loss_criterion.train()
avg_loss = 0
avg_accuracy = 0
n_items = 0
for step, full_data in enumerate(data_loader):
x, x_len, y, y_len = full_data
x_batch_len = x.shape[-1]
x, y = x.to(device), y.to(device)
bs=x.size(0)
optimizer.zero_grad()
context_out, enc_out, _ = cpc_model(x.to(device),y.to(device))
scores = phone_classifier(context_out)
scores = scores.permute(1,0,2)
scores = F.log_softmax(scores,2)
yhat_len = torch.tensor([int(scores.shape[0]*x_len[i]/x_batch_len) for i in range(scores.shape[1])]) # this is an approximation, should be good enough
loss = loss_criterion(scores,y.to(device),yhat_len,y_len)
loss.backward()
optimizer.step()
avg_loss+=loss.item()*bs
n_items+=bs
avg_loss/=n_items
return avg_loss
def validation_step(cpc_model,
phone_classifier,
loss_criterion,
data_loader):
cpc_model.eval()
phone_classifier.eval()
avg_loss = 0
avg_accuracy = 0
n_items = 0
with torch.no_grad():
for step, full_data in enumerate(data_loader):
x, x_len, y, y_len = full_data
x_batch_len = x.shape[-1]
x, y = x.to(device), y.to(device)
bs=x.size(0)
context_out, enc_out, _ = cpc_model(x.to(device),y.to(device))
scores = phone_classifier(context_out)
scores = scores.permute(1,0,2)
scores = F.log_softmax(scores,2)
yhat_len = torch.tensor([int(scores.shape[0]*x_len[i]/x_batch_len) for i in range(scores.shape[1])]) # this is an approximation, should be good enough
loss = loss_criterion(scores,y.to(device),yhat_len,y_len)
avg_loss+=loss.item()*bs
n_items+=bs
avg_loss/=n_items
return avg_loss
def run_ctc(cpc_model,
phone_classifier,
loss_criterion,
data_loader_train,
data_loader_val,
optimizer,
n_epoch):
for epoch in range(n_epoch):
print(f"Running epoch {epoch + 1} / {n_epoch}")
loss_train = train_one_epoch_ctc(cpc_model, phone_classifier, loss_criterion, data_loader_train, optimizer)
print("-------------------")
print(f"Training dataset :")
print(f"Average loss : {loss_train}.")
print("-------------------")
print("Validation dataset")
loss_val = validation_step(cpc_model, phone_classifier, loss_criterion, data_loader_val)
print(f"Average loss : {loss_val}")
print("-------------------")
print()
run_ctc(cpc_model,phone_classifier,loss_ctc,data_loader_train,data_loader_val,optimizer_frozen,n_epoch=10)
```
### b- Evaluation: the Phone Error Rate (PER)
In order to compute the similarity between two sequences, we can use the [Levenshtein distance](https://en.wikipedia.org/wiki/Levenshtein_distance). This distance estimates the minimum number of insertion, deletion and addition to move from one sequence to another. If we normalize this distance by the number of characters in the reference sequence we get the Phone Error Rate (PER).
This value can be interpreted as :
\\[ PER = \frac{S + D + I}{N} \\]
Where:
* N is the number of characters in the reference
* S is the number of substitutiion
* I in the number of insertion
* D in the number of deletion
For the best possible alignment of the two sequences.
```
import numpy as np
def get_PER_sequence(ref_seq, target_seq):
# re = g.split()
# h = h.split()
n = len(ref_seq)
m = len(target_seq)
D = np.zeros((n+1,m+1))
for i in range(1,n+1):
D[i,0] = D[i-1,0]+1
for j in range(1,m+1):
D[0,j] = D[0,j-1]+1
### TODO compute the alignment
for i in range(1,n+1):
for j in range(1,m+1):
D[i,j] = min(
D[i-1,j]+1,
D[i-1,j-1]+1,
D[i,j-1]+1,
D[i-1,j-1]+ 0 if ref_seq[i-1]==target_seq[j-1] else float("inf")
)
return D[n,m]/len(ref_seq)
#return PER
```
You can test your function below:
```
ref_seq = [0, 1, 1, 2, 0, 2, 2]
pred_seq = [1, 1, 2, 2, 0, 0]
expected_PER = 4. / 7.
print(get_PER_sequence(ref_seq, pred_seq) == expected_PER)
```
## c- Evaluating the PER of your model on the test dataset
Evaluate the PER on the validation dataset. Please notice that you should usually use a separate dataset, called the dev dataset, to perform this operation. However for the sake of simplicity we will work with validation data in this exercise.
```
import progressbar
from multiprocessing import Pool
def cut_data(seq, sizeSeq):
maxSeq = sizeSeq.max()
return seq[:, :maxSeq]
def prepare_data(data):
seq, sizeSeq, phone, sizePhone = data
seq = seq.cuda()
phone = phone.cuda()
sizeSeq = sizeSeq.cuda().view(-1)
sizePhone = sizePhone.cuda().view(-1)
seq = cut_data(seq.permute(0, 2, 1), sizeSeq).permute(0, 2, 1)
return seq, sizeSeq, phone, sizePhone
def get_per(test_dataloader,
cpc_model,
phone_classifier):
downsampling_factor = 160
cpc_model.eval()
phone_classifier.eval()
avgPER = 0
nItems = 0
print("Starting the PER computation through beam search")
bar = progressbar.ProgressBar(maxval=len(test_dataloader))
bar.start()
for index, data in enumerate(test_dataloader):
bar.update(index)
with torch.no_grad():
seq, sizeSeq, phone, sizePhone = prepare_data(data)
c_feature, _, _ = cpc_model(seq.to(device),phone.to(device))
sizeSeq = sizeSeq / downsampling_factor
predictions = torch.nn.functional.softmax(
phone_classifier(c_feature), dim=2).cpu()
phone = phone.cpu()
sizeSeq = sizeSeq.cpu()
sizePhone = sizePhone.cpu()
bs = c_feature.size(0)
data_per = [(predictions[b].argmax(1), phone[b]) for b in range(bs)]
# data_per = [(predictions[b], sizeSeq[b], phone[b], sizePhone[b],
# "criterion.module.BLANK_LABEL") for b in range(bs)]
with Pool(bs) as p:
poolData = p.starmap(get_PER_sequence, data_per)
avgPER += sum([x for x in poolData])
nItems += len(poolData)
bar.finish()
avgPER /= nItems
print(f"Average PER {avgPER}")
return avgPER
get_per(data_loader_val,cpc_model,phone_classifier)
```
## Exercice 3 : Character error rate (CER)
The Character Error Rate (CER) is an evaluation metric similar to the PER but with characters insterad of phonemes. Using the following data, run the functions you defined previously to estimate the CER of your model after fine-tuning.
```
# Load a dataset labelled with the letters of each sequence.
%cd /content/CPC_audio
from cpc.eval.common_voices_eval import SingleSequenceDataset, parseSeqLabels, findAllSeqs
path_train_data_cer = '/content/per_data/pack_master/1h'
path_val_data_cer = '/content/per_data/pack_master/10min'
path_letter_data_cer = '/content/per_data/pack_master/chars.txt'
BATCH_SIZE=8
letters_labels, N_LETTERS = parseSeqLabels(path_letter_data_cer)
data_train_cer, _ = findAllSeqs(path_train_data_cer, extension='.flac')
dataset_train_non_aligned = SingleSequenceDataset(path_train_data_cer, data_train_cer, letters_labels)
data_val_cer, _ = findAllSeqs(path_val_data_cer, extension='.flac')
dataset_val_non_aligned = SingleSequenceDataset(path_val_data_cer, data_val_cer, letters_labels)
# The data loader will generate a tuple of tensors data, labels for each batch
# data : size N x T1 x 1 : the audio sequence
# label : size N x T2 the sequence of letters corresponding to the audio data
# IMPORTANT NOTE: just like the PER the CER is computed with non-aligned phone data.
data_loader_train_letters = torch.utils.data.DataLoader(dataset_train_non_aligned, batch_size=BATCH_SIZE,
shuffle=True)
data_loader_val_letters = torch.utils.data.DataLoader(dataset_val_non_aligned, batch_size=BATCH_SIZE,
shuffle=True)
from cpc.feature_loader import loadModel
checkpoint_path = 'checkpoint_data/checkpoint_30.pt'
cpc_model, HIDDEN_CONTEXT_MODEL, HIDDEN_ENCODER_MODEL = loadModel([checkpoint_path])
cpc_model = cpc_model.cuda()
character_classifier = PhoneClassifier(HIDDEN_CONTEXT_MODEL, N_LETTERS).to(device)
parameters = list(character_classifier.parameters()) + list(cpc_model.parameters())
LEARNING_RATE = 2e-4
optimizer = torch.optim.Adam(parameters, lr=LEARNING_RATE)
optimizer_frozen = torch.optim.Adam(list(character_classifier.parameters()), lr=LEARNING_RATE)
loss_ctc = torch.nn.CTCLoss()
run_ctc(cpc_model,character_classifier,loss_ctc,data_loader_train_letters,data_loader_val_letters,optimizer_frozen,n_epoch=10)
get_per(data_loader_val_letters,cpc_model,character_classifier)
```
| github_jupyter |
```
# Copyright 2021 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Vertex Pipelines: AutoML text classification pipelines using google-cloud-pipeline-components
<table align="left">
<td>
<a href="https://colab.research.google.com/github/GoogleCloudPlatform/ai-platform-unified/notebooks/official/google_cloud_pipeline_components_automl_text.ipynb">
<img src="https://cloud.google.com/ml-engine/images/colab-logo-32px.png" alt="Colab logo"> Run in Colab
</a>
</td>
<td>
<a href="https://github.com/GoogleCloudPlatform/ai-platform-unified/notebooks/official/google_cloud_pipeline_components_automl_text.ipynb">
<img src="https://cloud.google.com/ml-engine/images/github-logo-32px.png" alt="GitHub logo">
View on GitHub
</a>
</td>
<td>
<a href="https://console.cloud.google.com/ai/platform/notebooks/deploy-notebook?download_url=https://github.com/GoogleCloudPlatform/ai-platform-unified/notebooks/official/google_cloud_pipeline_components_automl_text.ipynb">
Open in Google Cloud Notebooks
</a>
</td>
</table>
<br/><br/><br/>
## Overview
This notebook shows how to use the components defined in [`google_cloud_pipeline_components`](https://github.com/kubeflow/pipelines/tree/master/components/google-cloud) to build an AutoML text classification workflow on [Vertex Pipelines](https://cloud.google.com/vertex-ai/docs/pipelines).
### Dataset
The dataset used for this tutorial is the [Happy Moments dataset](https://www.kaggle.com/ritresearch/happydb) from [Kaggle Datasets](https://www.kaggle.com/ritresearch/happydb). The version of the dataset you will use in this tutorial is stored in a public Cloud Storage bucket.
### Objective
In this tutorial, you create an AutoML text classification using a pipeline with components from `google_cloud_pipeline_components`.
The steps performed include:
- Create a `Dataset` resource.
- Train an AutoML `Model` resource.
- Creates an `Endpoint` resource.
- Deploys the `Model` resource to the `Endpoint` resource.
The components are [documented here](https://google-cloud-pipeline-components.readthedocs.io/en/latest/google_cloud_pipeline_components.aiplatform.html#module-google_cloud_pipeline_components.aiplatform).
### Costs
This tutorial uses billable components of Google Cloud:
* Vertex AI
* Cloud Storage
Learn about [Vertex AI
pricing](https://cloud.google.com/vertex-ai/pricing) and [Cloud Storage
pricing](https://cloud.google.com/storage/pricing), and use the [Pricing
Calculator](https://cloud.google.com/products/calculator/)
to generate a cost estimate based on your projected usage.
### Set up your local development environment
If you are using Colab or Google Cloud Notebook, your environment already meets all the requirements to run this notebook. You can skip this step.
Otherwise, make sure your environment meets this notebook's requirements. You need the following:
- The Cloud Storage SDK
- Git
- Python 3
- virtualenv
- Jupyter notebook running in a virtual environment with Python 3
The Cloud Storage guide to [Setting up a Python development environment](https://cloud.google.com/python/setup) and the [Jupyter installation guide](https://jupyter.org/install) provide detailed instructions for meeting these requirements. The following steps provide a condensed set of instructions:
1. [Install and initialize the SDK](https://cloud.google.com/sdk/docs/).
2. [Install Python 3](https://cloud.google.com/python/setup#installing_python).
3. [Install virtualenv](Ihttps://cloud.google.com/python/setup#installing_and_using_virtualenv) and create a virtual environment that uses Python 3.
4. Activate that environment and run `pip3 install Jupyter` in a terminal shell to install Jupyter.
5. Run `jupyter notebook` on the command line in a terminal shell to launch Jupyter.
6. Open this notebook in the Jupyter Notebook Dashboard.
## Installation
Install the latest version of Vertex SDK for Python.
```
import os
# Google Cloud Notebook
if os.path.exists("/opt/deeplearning/metadata/env_version"):
USER_FLAG = "--user"
else:
USER_FLAG = ""
! pip3 install --upgrade google-cloud-aiplatform $USER_FLAG
```
Install the latest GA version of *google-cloud-storage* library as well.
```
! pip3 install -U google-cloud-storage $USER_FLAG
```
Install the latest GA version of *google-cloud-pipeline-components* library as well.
```
! pip3 install $USER kfp google-cloud-pipeline-components --upgrade
```
### Restart the kernel
Once you've installed the additional packages, you need to restart the notebook kernel so it can find the packages.
```
import os
if not os.getenv("IS_TESTING"):
# Automatically restart kernel after installs
import IPython
app = IPython.Application.instance()
app.kernel.do_shutdown(True)
```
Check the versions of the packages you installed. The KFP SDK version should be >=1.6.
```
! python3 -c "import kfp; print('KFP SDK version: {}'.format(kfp.__version__))"
! python3 -c "import google_cloud_pipeline_components; print('google_cloud_pipeline_components version: {}'.format(google_cloud_pipeline_components.__version__))"
```
## Before you begin
### GPU runtime
This tutorial does not require a GPU runtime.
### Set up your Google Cloud project
**The following steps are required, regardless of your notebook environment.**
1. [Select or create a Google Cloud project](https://console.cloud.google.com/cloud-resource-manager). When you first create an account, you get a $300 free credit towards your compute/storage costs.
2. [Make sure that billing is enabled for your project.](https://cloud.google.com/billing/docs/how-to/modify-project)
3. [Enable the Vertex AI APIs, Compute Engine APIs, and Cloud Storage.](https://console.cloud.google.com/flows/enableapi?apiid=ml.googleapis.com,compute_component,storage-component.googleapis.com)
4. [The Google Cloud SDK](https://cloud.google.com/sdk) is already installed in Google Cloud Notebook.
5. Enter your project ID in the cell below. Then run the cell to make sure the
Cloud SDK uses the right project for all the commands in this notebook.
**Note**: Jupyter runs lines prefixed with `!` as shell commands, and it interpolates Python variables prefixed with `$`.
```
PROJECT_ID = "[your-project-id]" # @param {type:"string"}
if PROJECT_ID == "" or PROJECT_ID is None or PROJECT_ID == "[your-project-id]":
# Get your GCP project id from gcloud
shell_output = ! gcloud config list --format 'value(core.project)' 2>/dev/null
PROJECT_ID = shell_output[0]
print("Project ID:", PROJECT_ID)
! gcloud config set project $PROJECT_ID
```
#### Region
You can also change the `REGION` variable, which is used for operations
throughout the rest of this notebook. Below are regions supported for Vertex AI. We recommend that you choose the region closest to you.
- Americas: `us-central1`
- Europe: `europe-west4`
- Asia Pacific: `asia-east1`
You may not use a multi-regional bucket for training with Vertex AI. Not all regions provide support for all Vertex AI services.
Learn more about [Vertex AI regions](https://cloud.google.com/vertex-ai/docs/general/locations)
```
REGION = "us-central1" # @param {type: "string"}
```
#### Timestamp
If you are in a live tutorial session, you might be using a shared test account or project. To avoid name collisions between users on resources created, you create a timestamp for each instance session, and append the timestamp onto the name of resources you create in this tutorial.
```
from datetime import datetime
TIMESTAMP = datetime.now().strftime("%Y%m%d%H%M%S")
```
### Authenticate your Google Cloud account
**If you are using Google Cloud Notebook**, your environment is already authenticated. Skip this step.
**If you are using Colab**, run the cell below and follow the instructions when prompted to authenticate your account via oAuth.
**Otherwise**, follow these steps:
In the Cloud Console, go to the [Create service account key](https://console.cloud.google.com/apis/credentials/serviceaccountkey) page.
**Click Create service account**.
In the **Service account name** field, enter a name, and click **Create**.
In the **Grant this service account access to project** section, click the Role drop-down list. Type "Vertex" into the filter box, and select **Vertex Administrator**. Type "Storage Object Admin" into the filter box, and select **Storage Object Admin**.
Click Create. A JSON file that contains your key downloads to your local environment.
Enter the path to your service account key as the GOOGLE_APPLICATION_CREDENTIALS variable in the cell below and run the cell.
```
# If you are running this notebook in Colab, run this cell and follow the
# instructions to authenticate your GCP account. This provides access to your
# Cloud Storage bucket and lets you submit training jobs and prediction
# requests.
import os
import sys
# If on Google Cloud Notebook, then don't execute this code
if not os.path.exists("/opt/deeplearning/metadata/env_version"):
if "google.colab" in sys.modules:
from google.colab import auth as google_auth
google_auth.authenticate_user()
# If you are running this notebook locally, replace the string below with the
# path to your service account key and run this cell to authenticate your GCP
# account.
elif not os.getenv("IS_TESTING"):
%env GOOGLE_APPLICATION_CREDENTIALS ''
```
### Create a Cloud Storage bucket
**The following steps are required, regardless of your notebook environment.**
When you initialize the Vertex SDK for Python, you specify a Cloud Storage staging bucket. The staging bucket is where all the data associated with your dataset and model resources are retained across sessions.
Set the name of your Cloud Storage bucket below. Bucket names must be globally unique across all Google Cloud projects, including those outside of your organization.
```
BUCKET_NAME = "gs://[your-bucket-name]" # @param {type:"string"}
if BUCKET_NAME == "" or BUCKET_NAME is None or BUCKET_NAME == "gs://[your-bucket-name]":
BUCKET_NAME = "gs://" + PROJECT_ID + "aip-" + TIMESTAMP
```
**Only if your bucket doesn't already exist**: Run the following cell to create your Cloud Storage bucket.
```
! gsutil mb -l $REGION $BUCKET_NAME
```
Finally, validate access to your Cloud Storage bucket by examining its contents:
```
! gsutil ls -al $BUCKET_NAME
```
#### Service Account
**If you don't know your service account**, try to get your service account using `gcloud` command by executing the second cell below.
```
SERVICE_ACCOUNT = "[your-service-account]" # @param {type:"string"}
if (
SERVICE_ACCOUNT == ""
or SERVICE_ACCOUNT is None
or SERVICE_ACCOUNT == "[your-service-account]"
):
# Get your GCP project id from gcloud
shell_output = !gcloud auth list 2>/dev/null
SERVICE_ACCOUNT = shell_output[2].strip()
print("Service Account:", SERVICE_ACCOUNT)
```
#### Set service account access for Vertex Pipelines
Run the following commands to grant your service account access to read and write pipeline artifacts in the bucket that you created in the previous step -- you only need to run these once per service account.
```
! gsutil iam ch serviceAccount:{SERVICE_ACCOUNT}:roles/storage.objectCreator $BUCKET_NAME
! gsutil iam ch serviceAccount:{SERVICE_ACCOUNT}:roles/storage.objectViewer $BUCKET_NAME
```
### Set up variables
Next, set up some variables used throughout the tutorial.
### Import libraries and define constants
```
import google.cloud.aiplatform as aip
```
#### Vertex AI constants
Setup up the following constants for Vertex AI:
- `API_ENDPOINT`: The Vertex AI API service endpoint for `Dataset`, `Model`, `Job`, `Pipeline` and `Endpoint` services.
```
# API service endpoint
API_ENDPOINT = "{}-aiplatform.googleapis.com".format(REGION)
```
#### Vertex Pipelines constants
Setup up the following constants for Vertex Pipelines:
```
PIPELINE_ROOT = "{}/pipeline_root/happydb".format(BUCKET_NAME)
```
Additional imports.
```
import kfp
from google_cloud_pipeline_components import aiplatform as gcc_aip
```
## Initialize Vertex SDK for Python
Initialize the Vertex SDK for Python for your project and corresponding bucket.
```
aip.init(project=PROJECT_ID, staging_bucket=BUCKET_NAME)
```
## Define AutoML text classification model pipeline that uses components from `google_cloud_pipeline_components`
Next, you define the pipeline.
Create and deploy an AutoML text classification `Model` resource using a `Dataset` resource.
```
IMPORT_FILE = "gs://cloud-ml-data/NL-classification/happiness.csv"
@kfp.dsl.pipeline(name="automl-text-classification" + TIMESTAMP)
def pipeline(project: str = PROJECT_ID, import_file: str = IMPORT_FILE):
dataset_create_task = gcc_aip.TextDatasetCreateOp(
display_name="train-automl-happydb",
gcs_source=import_file,
import_schema_uri=aip.schema.dataset.ioformat.text.multi_label_classification,
project=project,
)
training_run_task = gcc_aip.AutoMLTextTrainingJobRunOp(
dataset=dataset_create_task.outputs["dataset"],
display_name="train-automl-happydb",
prediction_type="classification",
multi_label=True,
training_fraction_split=0.6,
validation_fraction_split=0.2,
test_fraction_split=0.2,
model_display_name="train-automl-happydb",
project=project,
)
model_deploy_op = gcc_aip.ModelDeployOp( # noqa: F841
model=training_run_task.outputs["model"], project=project
)
```
## Compile the pipeline
Next, compile the pipeline.
```
from kfp.v2 import compiler # noqa: F811
compiler.Compiler().compile(
pipeline_func=pipeline,
package_path="text classification_pipeline.json".replace(" ", "_"),
)
```
## Run the pipeline
Next, run the pipeline.
```
DISPLAY_NAME = "happydb_" + TIMESTAMP
job = aip.PipelineJob(
display_name=DISPLAY_NAME,
template_path="text classification_pipeline.json".replace(" ", "_"),
pipeline_root=PIPELINE_ROOT,
)
job.run()
```
Click on the generated link to see your run in the Cloud Console.
<!-- It should look something like this as it is running:
<a href="https://storage.googleapis.com/amy-jo/images/mp/automl_tabular_classif.png" target="_blank"><img src="https://storage.googleapis.com/amy-jo/images/mp/automl_tabular_classif.png" width="40%"/></a> -->
In the UI, many of the pipeline DAG nodes will expand or collapse when you click on them. Here is a partially-expanded view of the DAG (click image to see larger version).
<a href="https://storage.googleapis.com/amy-jo/images/mp/automl_text_classif.png" target="_blank"><img src="https://storage.googleapis.com/amy-jo/images/mp/automl_text_classif.png" width="40%"/></a>
# Cleaning up
To clean up all Google Cloud resources used in this project, you can [delete the Google Cloud
project](https://cloud.google.com/resource-manager/docs/creating-managing-projects#shutting_down_projects) you used for the tutorial.
Otherwise, you can delete the individual resources you created in this tutorial -- *Note:* this is auto-generated and not all resources may be applicable for this tutorial:
- Dataset
- Pipeline
- Model
- Endpoint
- Batch Job
- Custom Job
- Hyperparameter Tuning Job
- Cloud Storage Bucket
```
delete_dataset = True
delete_pipeline = True
delete_model = True
delete_endpoint = True
delete_batchjob = True
delete_customjob = True
delete_hptjob = True
delete_bucket = True
try:
if delete_model and "DISPLAY_NAME" in globals():
models = aip.Model.list(
filter=f"display_name={DISPLAY_NAME}", order_by="create_time"
)
model = models[0]
aip.Model.delete(model)
print("Deleted model:", model)
except Exception as e:
print(e)
try:
if delete_endpoint and "DISPLAY_NAME" in globals():
endpoints = aip.Endpoint.list(
filter=f"display_name={DISPLAY_NAME}_endpoint", order_by="create_time"
)
endpoint = endpoints[0]
endpoint.undeploy_all()
aip.Endpoint.delete(endpoint.resource_name)
print("Deleted endpoint:", endpoint)
except Exception as e:
print(e)
if delete_dataset and "DISPLAY_NAME" in globals():
if "text" == "tabular":
try:
datasets = aip.TabularDataset.list(
filter=f"display_name={DISPLAY_NAME}", order_by="create_time"
)
dataset = datasets[0]
aip.TabularDataset.delete(dataset.resource_name)
print("Deleted dataset:", dataset)
except Exception as e:
print(e)
if "text" == "image":
try:
datasets = aip.ImageDataset.list(
filter=f"display_name={DISPLAY_NAME}", order_by="create_time"
)
dataset = datasets[0]
aip.ImageDataset.delete(dataset.resource_name)
print("Deleted dataset:", dataset)
except Exception as e:
print(e)
if "text" == "text":
try:
datasets = aip.TextDataset.list(
filter=f"display_name={DISPLAY_NAME}", order_by="create_time"
)
dataset = datasets[0]
aip.TextDataset.delete(dataset.resource_name)
print("Deleted dataset:", dataset)
except Exception as e:
print(e)
if "text" == "video":
try:
datasets = aip.VideoDataset.list(
filter=f"display_name={DISPLAY_NAME}", order_by="create_time"
)
dataset = datasets[0]
aip.VideoDataset.delete(dataset.resource_name)
print("Deleted dataset:", dataset)
except Exception as e:
print(e)
try:
if delete_pipeline and "DISPLAY_NAME" in globals():
pipelines = aip.PipelineJob.list(
filter=f"display_name={DISPLAY_NAME}", order_by="create_time"
)
pipeline = pipelines[0]
aip.PipelineJob.delete(pipeline.resource_name)
print("Deleted pipeline:", pipeline)
except Exception as e:
print(e)
if delete_bucket and "BUCKET_NAME" in globals():
! gsutil rm -r $BUCKET_NAME
```
| github_jupyter |
<a href="https://colab.research.google.com/github/souvikg123/Python-NLTK-Sentiment-Analysis./blob/master/LSTM_model_for__RNN.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
#LSTM
from pandas import DataFrame
from pandas import Series
from pandas import concat
from pandas import read_csv
from pandas import datetime
from sklearn.metrics import mean_squared_error
from sklearn.preprocessing import MinMaxScaler
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import LSTM
from math import sqrt
from matplotlib import pyplot
import numpy
def parser(x):
return datetime.strptime('190'+x, '%Y-%m')
# frame a sequence as a supervised learning problem
def timeseries_to_supervised(data, lag=1):
df = DataFrame(data)
columns = [df.shift(i) for i in range(1, lag+1)]
columns.append(df)
df = concat(columns, axis=1)
df.fillna(0, inplace=True)
return df
# create a differenced series
def difference(dataset, interval=1):
diff = list()
for i in range(interval, len(dataset)):
value = dataset[i] - dataset[i - interval]
diff.append(value)
return Series(diff)
# invert differenced value
def inverse_difference(history, yhat, interval=1):
return yhat + history[-interval]
# scale train and test data to [-1, 1]
def scale(train, test):
# fit scaler
scaler = MinMaxScaler(feature_range=(-1, 1))
scaler = scaler.fit(train)
# transform train
train = train.reshape(train.shape[0], train.shape[1])
train_scaled = scaler.transform(train)
# transform test
test = test.reshape(test.shape[0], test.shape[1])
test_scaled = scaler.transform(test)
return scaler, train_scaled, test_scaled
def invert_scale(scaler, X, value):
new_row = [x for x in X] + [value]
array = numpy.array(new_row)
array = array.reshape(1, len(array))
inverted = scaler.inverse_transform(array)
return inverted[0, -1]
# fit an LSTM network to training data
def fit_lstm(train, batch_size, nb_epoch, neurons):
X, y = train[:, 0:-1], train[:, -1]
X = X.reshape(X.shape[0], 1, X.shape[1])
model = Sequential()
model.add(LSTM(neurons, batch_input_shape=(batch_size, X.shape[1], X.shape[2]), stateful=True))
model.add(Dense(1))
model.compile(loss='mean_squared_error', optimizer='adam')
for i in range(nb_epoch):
model.fit(X, y, epochs=1, batch_size=batch_size, verbose=0, shuffle=False)
model.reset_states()
return model
# make a one-step forecast
def forecast_lstm(model, batch_size, X):
X = X.reshape(1, 1, len(X))
yhat = model.predict(X, batch_size=batch_size)
return yhat[0,0]
# make a one-step forecast
def forecast_lstm(model, batch_size, X):
X = X.reshape(1, 1, len(X))
yhat = model.predict(X, batch_size=batch_size)
return yhat[0,0]
# load dataset
series = read_csv('shampoo.csv', header=0, parse_dates=[0], index_col=0, squeeze=True, date_parser=parser)
# transform data to be stationary
raw_values = series.values
diff_values = difference(raw_values, 1)
# transform data to be supervised learning
supervised = timeseries_to_supervised(diff_values, 1)
supervised_values = supervised.values
# split data into train and test-sets
train, test = supervised_values[0:-12], supervised_values[-12:]
# transform the scale of the data
scaler, train_scaled, test_scaled = scale(train, test)
# fit the model
lstm_model = fit_lstm(train_scaled, 1, 3000, 4)
# forecast the entire training dataset to build up state for forecasting
train_reshaped = train_scaled[:, 0].reshape(len(train_scaled), 1, 1)
lstm_model.predict(train_reshaped, batch_size=1)
predictions = list()
for i in range(len(test_scaled)):
# make one-step forecast
X, y = test_scaled[i, 0:-1], test_scaled[i, -1]
yhat = forecast_lstm(lstm_model, 1, X)
# invert scaling
yhat = invert_scale(scaler, X, yhat)
# invert differencing
yhat = inverse_difference(raw_values, yhat, len(test_scaled)+1-i)
# store forecast
predictions.append(yhat)
expected = raw_values[len(train) + i + 1]
print('Month=%d, Predicted=%f, Expected=%f' % (i+1, yhat, expected))
# report performance
rmse = sqrt(mean_squared_error(raw_values[-12:], predictions))
print('Test RMSE: %.3f' % rmse)
# line plot of observed vs predicted
pyplot.plot(raw_values[-12:])
pyplot.plot(predictions)
pyplot.show()
```
| github_jupyter |
# Ray Serve - Model Serving
© 2019-2022, Anyscale. All Rights Reserved

Now we'll explore a short example for Ray Serve. This example is from the Ray Serve [scikit-learn example.](https://docs.ray.io/en/latest/serve/tutorials/sklearn.html)
See also the Serve documentation's [mini-tutorials](https://docs.ray.io/en/latest/serve/tutorials/index.html) for using Serve with various frameworks.
<img src="../images/ray_serve_deployment_workflow.png" width="90%" height="50%">
```
import ray
from ray import serve
import os
import requests # for making web requests
import tempfile
serve.start()
```
## Get a Model to Serve
We'll begin by training a classifier with the Iris data we used before, this time using [scikit-learn](https://scikit-learn.org/stable/). The details aren't too important for our purposes, except for the fact we'll save the trained model to disk for subsequent serving.
```
import pickle
import json
import numpy as np
import sklearn
from sklearn.datasets import load_iris
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.metrics import mean_squared_error
# Load data
iris_dataset = load_iris()
data, target, target_names = iris_dataset["data"], iris_dataset[
"target"], iris_dataset["target_names"]
# Instantiate model
model = GradientBoostingClassifier()
# Training and validation split
data, target = sklearn.utils.shuffle(data, target)
train_x, train_y = data[:100], target[:100]
val_x, val_y = data[100:], target[100:]
# Train and evaluate models
model.fit(train_x, train_y)
print("MSE:", mean_squared_error(model.predict(val_x), val_y))
# Save the model and label to file
MODEL_PATH = os.path.join(tempfile.gettempdir(),
"iris_model_logistic_regression.pkl")
LABEL_PATH = os.path.join(tempfile.gettempdir(), "iris_labels.json")
# Save the model and label to file. (This could also be S3 or other "global" place)
with open(MODEL_PATH, "wb") as f:
pickle.dump(model, f)
with open(LABEL_PATH, "w") as f:
json.dump(target_names.tolist(), f)
```
## Create a Model and Serve It
Next, we define a servable model by instantiating a class and defining the `__call__` method that Ray Serve will use.
```
@serve.deployment(route_prefix="/regressor")
class BoostingModel:
def __init__(self):
with open(MODEL_PATH, "rb") as f:
self.model = pickle.load(f)
with open(LABEL_PATH) as f:
self.label_list = json.load(f)
# async allows us to have this call concurrently
async def __call__(self, starlette_request):
payload = await starlette_request.json()
print("Worker: received starlette request with data", payload)
input_vector = [
payload["sepal length"],
payload["sepal width"],
payload["petal length"],
payload["petal width"],
]
prediction = self.model.predict([input_vector])[0]
human_name = self.label_list[prediction]
return {"result": human_name}
```
## Deploy the model
```
BoostingModel.deploy()
```
## Score the model
Internally, Serve stores the model as a Ray actor and routes traffic to it as the endpoint is queried, in this case over HTTP.
Now let’s query the endpoint to see results.
```
sample_request_input = {
"sepal length": 1.2,
"sepal width": 1.0,
"petal length": 1.1,
"petal width": 0.9,
}
```
We can now send HTTP requests to our route `route_prefix=/regressor` at the default port 8000
```
response = requests.get(
"http://localhost:8000/regressor", json=sample_request_input)
print(response.text)
for i in range(10):
response = requests.get("http://localhost:8000/regressor", json=sample_request_input).json()
print(response)
```
## Cleanup
```
deployments = serve.list_deployments()
print(f'deployments: {deployments}')
serve.shutdown()
```
## Exercise - Try Adding more examples
Here are some things you can try:
1. Send more input requests.
2. Add a small model of your own
| github_jupyter |
# GAN Flavours
This jupyter notebook contains a training script for the https://github.com/beresandras/gan-flavours-keras repository, and is intended to be used in a Google Colab environment.
```
# uncomment on first run
# !pip install tensorflow_addons
# !git clone https://github.com/beresandras/gan-flavours-keras
import sys
import tensorflow as tf
from tensorflow import keras
sys.path.insert(0,'/content/WGAN')
from dataset import prepare_dataset
from architecture import get_generator, get_discriminator
from augmentation import AdaptiveAugmenter
from losses import (
MiniMaxGAN,
NonSaturatingGAN,
LeastSquaresGAN,
HingeGAN,
WassersteinGAN,
RelativisticGAN,
RelativisticAverageGAN,
)
from utils import generate_images_with, plot_history
# hyperparameters
# data
# some datasets might be unavailable for download at times
dataset_name = "playable" # "oxford_flowers102", "celeb_a", "cifar10"
image_size = 64 # 64, 64, 32
num_epochs = 400 # 500, 25, 100
kid_image_size = 75 # resolution of KID measurement, default 299
plot_interval = 10 # 10, 1, 2
# optimization
batch_size = 128
one_sided_label_smoothing = 0.0 # can be 0.1
ema = 0.99
generator_lr = 2e-4
discriminator_lr = 2e-4
beta_1 = 0.5
beta_2 = 0.999
# architecture
noise_size = 64
depth = 4 # number of up- and downsampling layers, change with resolution
width = 128
initializer = "glorot_uniform"
residual = False
transposed = True # transposed convs vs upsampling + convs in generator
leaky_relu_slope = 0.2
dropout_rate = 0.4
spectral_norm = False
# adaptive discriminator augmentation
target_accuracy = None # 0.85, set to None to disable
integration_steps = 1000
max_probability = 0.8 # maximal augmentation probability
id = 0
# load dataset
train_dataset = prepare_dataset(dataset_name, "train", image_size, batch_size)
val_dataset = prepare_dataset(dataset_name, "validation", image_size, batch_size)
# create model
model = NonSaturatingGAN(
id=id,
generator=get_generator(
noise_size, depth, width, initializer, residual, transposed
),
discriminator=get_discriminator(
image_size,
depth,
width,
initializer,
residual,
leaky_relu_slope,
dropout_rate,
spectral_norm,
),
augmenter=AdaptiveAugmenter(
target_accuracy=target_accuracy,
integration_steps=integration_steps,
max_probability=max_probability,
input_shape=(image_size, image_size, 3),
),
one_sided_label_smoothing=one_sided_label_smoothing,
ema=ema,
kid_image_size=kid_image_size,
plot_interval=plot_interval,
is_jupyter=True,
)
model.compile(
generator_optimizer=keras.optimizers.Adam(
learning_rate=generator_lr, beta_1=beta_1, beta_2=beta_2
),
discriminator_optimizer=keras.optimizers.Adam(
learning_rate=discriminator_lr, beta_1=beta_1, beta_2=beta_2
),
)
# checkpointing
checkpoint_path = "checkpoints/model_{}".format(id)
checkpoint_callback = tf.keras.callbacks.ModelCheckpoint(
filepath=checkpoint_path,
save_weights_only=True,
monitor="val_kid",
mode="min",
save_best_only=True,
)
# run training
history = model.fit(
train_dataset,
epochs=num_epochs,
validation_data=val_dataset,
callbacks=[
keras.callbacks.LambdaCallback(on_epoch_end=model.plot_images),
checkpoint_callback,
],
)
# load best model
model.load_weights(checkpoint_path)
generate_images_with(model, history, id, is_jupyter=True)
# plot history
plot_history(history, id, is_jupyter=True)
```
| github_jupyter |
# Furniture Rearrangement - How to setup a new interaction task in Habitat-Lab
This tutorial demonstrates how to setup a new task in Habitat that utilizes interaction capabilities in Habitat Simulator.

## Task Definition:
The working example in this demo will be the task of **Furniture Rearrangement** - The agent will be randomly spawned in an environment in which the furniture are initially displaced from their desired position. The agent is tasked with navigating the environment, picking furniture and putting them in the desired position. To keep the tutorial simple and easy to follow, we will rearrange just a single object.
To setup this task, we will build on top of existing API in Habitat-Simulator and Habitat-Lab. Here is a summary of all the steps involved in setting up this task:
1. **Setup the Simulator**: Using existing functionalities of the Habitat-Sim, we can add or remove objects from the scene. We will use these methods to spawn the agent and the objects at some pre-defined initial configuration.
2. **Create a New Dataset**: We will define a new dataset class to save / load a list of episodes for the agent to train and evaluate on.
3. **Grab / Release Action**: We will add the "grab/release" action to the agent's action space to allow the agent to pickup / drop an object under a crosshair.
4. **Extend the Simulator Class**: We will extend the Simulator Class to add support for new actions implemented in previous step and add other additional utility functions
5. **Create a New Task**: Create a new task definition, implement new *sensors* and *metrics*.
6. **Train an RL agent**: We will define rewards for this task and utilize it to train an RL agent using the PPO algorithm.
Let's get started!
```
# @title Installation { display-mode: "form" }
# @markdown (double click to show code).
!curl -L https://raw.githubusercontent.com/facebookresearch/habitat-sim/master/examples/colab_utils/colab_install.sh | NIGHTLY=true bash -s
%cd /content
!gdown --id 1Pc-J6pZzXEd8RSeLM94t3iwO8q_RQ853
!unzip -o /content/coda.zip -d /content/habitat-sim/data/scene_datasets
# reload the cffi version
import sys
if "google.colab" in sys.modules:
import importlib
import cffi
importlib.reload(cffi)
# @title Path Setup and Imports { display-mode: "form" }
# @markdown (double click to show code).
%cd /content/habitat-lab
## [setup]
import gzip
import json
import os
import sys
from typing import Any, Dict, List, Optional, Type
import attr
import cv2
import git
import magnum as mn
import numpy as np
%matplotlib inline
from matplotlib import pyplot as plt
from PIL import Image
import habitat
import habitat_sim
from habitat.config import Config
from habitat.core.registry import registry
from habitat_sim.utils import viz_utils as vut
if "google.colab" in sys.modules:
os.environ["IMAGEIO_FFMPEG_EXE"] = "/usr/bin/ffmpeg"
repo = git.Repo(".", search_parent_directories=True)
dir_path = repo.working_tree_dir
%cd $dir_path
data_path = os.path.join(dir_path, "data")
output_directory = "data/tutorials/output/" # @param {type:"string"}
output_path = os.path.join(dir_path, output_directory)
if __name__ == "__main__":
import argparse
parser = argparse.ArgumentParser()
parser.add_argument("--no-display", dest="display", action="store_false")
parser.add_argument(
"--no-make-video", dest="make_video", action="store_false"
)
parser.set_defaults(show_video=True, make_video=True)
args, _ = parser.parse_known_args()
show_video = args.display
display = args.display
make_video = args.make_video
else:
show_video = False
make_video = False
display = False
if make_video and not os.path.exists(output_path):
os.makedirs(output_path)
# @title Util functions to visualize observations
# @markdown - `make_video_cv2`: Renders a video from a list of observations
# @markdown - `simulate`: Runs simulation for a given amount of time at 60Hz
# @markdown - `simulate_and_make_vid` Runs simulation and creates video
def make_video_cv2(
observations, cross_hair=None, prefix="", open_vid=True, fps=60
):
sensor_keys = list(observations[0])
videodims = observations[0][sensor_keys[0]].shape
videodims = (videodims[1], videodims[0]) # flip to w,h order
print(videodims)
video_file = output_path + prefix + ".mp4"
print("Encoding the video: %s " % video_file)
writer = vut.get_fast_video_writer(video_file, fps=fps)
for ob in observations:
# If in RGB/RGBA format, remove the alpha channel
rgb_im_1st_person = cv2.cvtColor(ob["rgb"], cv2.COLOR_RGBA2RGB)
if cross_hair is not None:
rgb_im_1st_person[
cross_hair[0] - 2 : cross_hair[0] + 2,
cross_hair[1] - 2 : cross_hair[1] + 2,
] = [255, 0, 0]
if rgb_im_1st_person.shape[:2] != videodims:
rgb_im_1st_person = cv2.resize(
rgb_im_1st_person, videodims, interpolation=cv2.INTER_AREA
)
# write the 1st person observation to video
writer.append_data(rgb_im_1st_person)
writer.close()
if open_vid:
print("Displaying video")
vut.display_video(video_file)
def simulate(sim, dt=1.0, get_frames=True):
# simulate dt seconds at 60Hz to the nearest fixed timestep
print("Simulating " + str(dt) + " world seconds.")
observations = []
start_time = sim.get_world_time()
while sim.get_world_time() < start_time + dt:
sim.step_physics(1.0 / 60.0)
if get_frames:
observations.append(sim.get_sensor_observations())
return observations
# convenience wrapper for simulate and make_video_cv2
def simulate_and_make_vid(sim, crosshair, prefix, dt=1.0, open_vid=True):
observations = simulate(sim, dt)
make_video_cv2(observations, crosshair, prefix=prefix, open_vid=open_vid)
def display_sample(
rgb_obs,
semantic_obs=np.array([]),
depth_obs=np.array([]),
key_points=None, # noqa: B006
):
from habitat_sim.utils.common import d3_40_colors_rgb
rgb_img = Image.fromarray(rgb_obs, mode="RGB")
arr = [rgb_img]
titles = ["rgb"]
if semantic_obs.size != 0:
semantic_img = Image.new(
"P", (semantic_obs.shape[1], semantic_obs.shape[0])
)
semantic_img.putpalette(d3_40_colors_rgb.flatten())
semantic_img.putdata((semantic_obs.flatten() % 40).astype(np.uint8))
semantic_img = semantic_img.convert("RGBA")
arr.append(semantic_img)
titles.append("semantic")
if depth_obs.size != 0:
depth_img = Image.fromarray(
(depth_obs / 10 * 255).astype(np.uint8), mode="L"
)
arr.append(depth_img)
titles.append("depth")
plt.figure(figsize=(12, 8))
for i, data in enumerate(arr):
ax = plt.subplot(1, 3, i + 1)
ax.axis("off")
ax.set_title(titles[i])
# plot points on images
if key_points is not None:
for point in key_points:
plt.plot(
point[0], point[1], marker="o", markersize=10, alpha=0.8
)
plt.imshow(data)
plt.show(block=False)
```
## 1. Setup the Simulator
---
```
# @title Setup simulator configuration
# @markdown We'll start with setting up simulator with the following configurations
# @markdown - The simulator will render both RGB, Depth observations of 256x256 resolution.
# @markdown - The actions available will be `move_forward`, `turn_left`, `turn_right`.
def make_cfg(settings):
sim_cfg = habitat_sim.SimulatorConfiguration()
sim_cfg.gpu_device_id = 0
sim_cfg.default_agent_id = settings["default_agent_id"]
sim_cfg.scene.id = settings["scene"]
sim_cfg.enable_physics = settings["enable_physics"]
sim_cfg.physics_config_file = settings["physics_config_file"]
# Note: all sensors must have the same resolution
sensors = {
"rgb": {
"sensor_type": habitat_sim.SensorType.COLOR,
"resolution": [settings["height"], settings["width"]],
"position": [0.0, settings["sensor_height"], 0.0],
},
"depth": {
"sensor_type": habitat_sim.SensorType.DEPTH,
"resolution": [settings["height"], settings["width"]],
"position": [0.0, settings["sensor_height"], 0.0],
},
}
sensor_specs = []
for sensor_uuid, sensor_params in sensors.items():
if settings[sensor_uuid]:
sensor_spec = habitat_sim.SensorSpec()
sensor_spec.uuid = sensor_uuid
sensor_spec.sensor_type = sensor_params["sensor_type"]
sensor_spec.resolution = sensor_params["resolution"]
sensor_spec.position = sensor_params["position"]
sensor_specs.append(sensor_spec)
# Here you can specify the amount of displacement in a forward action and the turn angle
agent_cfg = habitat_sim.agent.AgentConfiguration()
agent_cfg.sensor_specifications = sensor_specs
agent_cfg.action_space = {
"move_forward": habitat_sim.agent.ActionSpec(
"move_forward", habitat_sim.agent.ActuationSpec(amount=0.1)
),
"turn_left": habitat_sim.agent.ActionSpec(
"turn_left", habitat_sim.agent.ActuationSpec(amount=10.0)
),
"turn_right": habitat_sim.agent.ActionSpec(
"turn_right", habitat_sim.agent.ActuationSpec(amount=10.0)
),
}
return habitat_sim.Configuration(sim_cfg, [agent_cfg])
settings = {
"max_frames": 10,
"width": 256,
"height": 256,
"scene": "data/scene_datasets/coda/coda.glb",
"default_agent_id": 0,
"sensor_height": 1.5, # Height of sensors in meters
"rgb": True, # RGB sensor
"depth": True, # Depth sensor
"seed": 1,
"enable_physics": True,
"physics_config_file": "data/default.phys_scene_config.json",
"silent": False,
"compute_shortest_path": False,
"compute_action_shortest_path": False,
"save_png": True,
}
cfg = make_cfg(settings)
# @title Spawn the agent at a pre-defined location
def init_agent(sim):
agent_pos = np.array([-0.15776923, 0.18244143, 0.2988735])
# Place the agent
sim.agents[0].scene_node.translation = agent_pos
agent_orientation_y = -40
sim.agents[0].scene_node.rotation = mn.Quaternion.rotation(
mn.Deg(agent_orientation_y), mn.Vector3(0, 1.0, 0)
)
cfg.sim_cfg.default_agent_id = 0
with habitat_sim.Simulator(cfg) as sim:
init_agent(sim)
if make_video:
# Visualize the agent's initial position
simulate_and_make_vid(
sim, None, "sim-init", dt=1.0, open_vid=show_video
)
# @title Set the object's initial and final position
# @markdown Defines two utility functions:
# @markdown - `remove_all_objects`: This will remove all objects from the scene
# @markdown - `set_object_in_front_of_agent`: This will add an object in the scene in front of the agent at the specified distance.
# @markdown Here we add a chair *3.0m* away from the agent and the task is to place the agent at the desired final position which is *7.0m* in front of the agent.
def remove_all_objects(sim):
for id in sim.get_existing_object_ids():
sim.remove_object(id)
def set_object_in_front_of_agent(sim, obj_id, z_offset=-1.5):
r"""
Adds an object in front of the agent at some distance.
"""
agent_transform = sim.agents[0].scene_node.transformation_matrix()
obj_translation = agent_transform.transform_point(
np.array([0, 0, z_offset])
)
sim.set_translation(obj_translation, obj_id)
obj_node = sim.get_object_scene_node(obj_id)
xform_bb = habitat_sim.geo.get_transformed_bb(
obj_node.cumulative_bb, obj_node.transformation
)
# also account for collision margin of the scene
scene_collision_margin = 0.04
y_translation = mn.Vector3(
0, xform_bb.size_y() / 2.0 + scene_collision_margin, 0
)
sim.set_translation(y_translation + sim.get_translation(obj_id), obj_id)
def init_objects(sim):
# Manager of Object Attributes Templates
obj_attr_mgr = sim.get_object_template_manager()
# Add a chair into the scene.
obj_path = "test_assets/objects/chair"
chair_template_id = obj_attr_mgr.load_object_configs(
str(os.path.join(data_path, obj_path))
)[0]
chair_attr = obj_attr_mgr.get_template_by_ID(chair_template_id)
obj_attr_mgr.register_template(chair_attr)
# Object's initial position 3m away from the agent.
object_id = sim.add_object_by_handle(chair_attr.handle)
set_object_in_front_of_agent(sim, object_id, -3.0)
sim.set_object_motion_type(
habitat_sim.physics.MotionType.STATIC, object_id
)
# Object's final position 7m away from the agent
goal_id = sim.add_object_by_handle(chair_attr.handle)
set_object_in_front_of_agent(sim, goal_id, -7.0)
sim.set_object_motion_type(habitat_sim.physics.MotionType.STATIC, goal_id)
return object_id, goal_id
with habitat_sim.Simulator(cfg) as sim:
init_agent(sim)
init_objects(sim)
# Visualize the scene after the chair is added into the scene.
if make_video:
simulate_and_make_vid(
sim, None, "object-init", dt=1.0, open_vid=show_video
)
```
## Rearrangement Dataset

In the previous section, we created a single episode of the rearrangement task. Let's define a format to store all the necessary information about a single episode. It should store the *scene* the episode belongs to, *initial spawn position and orientation* of the agent, *object type*, object's *initial position and orientation* as well as *final position and orientation*.
The format will be as follows:
```
{
'episode_id': 0,
'scene_id': 'data/scene_datasets/coda/coda.glb',
'goals': {
'position': [4.34, 0.67, -5.06],
'rotation': [0.0, 0.0, 0.0, 1.0]
},
'objects': {
'object_id': 0,
'object_template': 'data/test_assets/objects/chair',
'position': [1.77, 0.67, -1.99],
'rotation': [0.0, 0.0, 0.0, 1.0]
},
'start_position': [-0.15, 0.18, 0.29],
'start_rotation': [-0.0, -0.34, -0.0, 0.93]}
}
```
Once an episode is defined, a dataset will just be a collection of such episodes. For simplicity, in this notebook, the dataset will only contain one episode defined above.
```
# @title Create a new dataset
# @markdown Utility functions to define and save the dataset for the rearrangement task
def get_rotation(sim, object_id):
quat = sim.get_rotation(object_id)
return np.array(quat.vector).tolist() + [quat.scalar]
def init_episode_dict(episode_id, scene_id, agent_pos, agent_rot):
episode_dict = {
"episode_id": episode_id,
"scene_id": "data/scene_datasets/coda/coda.glb",
"start_position": agent_pos,
"start_rotation": agent_rot,
"info": {},
}
return episode_dict
def add_object_details(sim, episode_dict, id, object_template, object_id):
object_template = {
"object_id": id,
"object_template": object_template,
"position": np.array(sim.get_translation(object_id)).tolist(),
"rotation": get_rotation(sim, object_id),
}
episode_dict["objects"] = object_template
return episode_dict
def add_goal_details(sim, episode_dict, object_id):
goal_template = {
"position": np.array(sim.get_translation(object_id)).tolist(),
"rotation": get_rotation(sim, object_id),
}
episode_dict["goals"] = goal_template
return episode_dict
# set the number of objects to 1 always for now.
def build_episode(sim, episode_num, object_id, goal_id):
episodes = {"episodes": []}
for episode in range(episode_num):
agent_state = sim.get_agent(0).get_state()
agent_pos = np.array(agent_state.position).tolist()
agent_quat = agent_state.rotation
agent_rot = np.array(agent_quat.vec).tolist() + [agent_quat.real]
episode_dict = init_episode_dict(
episode, settings["scene"], agent_pos, agent_rot
)
object_attr = sim.get_object_initialization_template(object_id)
object_path = os.path.relpath(
os.path.splitext(object_attr.render_asset_handle)[0]
)
episode_dict = add_object_details(
sim, episode_dict, 0, object_path, object_id
)
episode_dict = add_goal_details(sim, episode_dict, goal_id)
episodes["episodes"].append(episode_dict)
return episodes
with habitat_sim.Simulator(cfg) as sim:
init_agent(sim)
object_id, goal_id = init_objects(sim)
episodes = build_episode(sim, 1, object_id, goal_id)
dataset_content_path = "data/datasets/rearrangement/coda/v1/train/"
if not os.path.exists(dataset_content_path):
os.makedirs(dataset_content_path)
with gzip.open(
os.path.join(dataset_content_path, "train.json.gz"), "wt"
) as f:
json.dump(episodes, f)
print(
"Dataset written to {}".format(
os.path.join(dataset_content_path, "train.json.gz")
)
)
# @title Dataset class to read the saved dataset in Habitat-Lab.
# @markdown To read the saved episodes in Habitat-Lab, we will extend the `Dataset` class and the `Episode` base class. It will help provide all the relevant details about the episode through a consistent API to all downstream tasks.
# @markdown - We will first create a `RearrangementEpisode` by extending the `NavigationEpisode` to include additional information about object's initial configuration and desired final configuration.
# @markdown - We will then define a `RearrangementDatasetV0` class that builds on top of `PointNavDatasetV1` class to read the JSON file stored earlier and initialize a list of `RearrangementEpisode`.
from habitat.core.utils import DatasetFloatJSONEncoder, not_none_validator
from habitat.datasets.pointnav.pointnav_dataset import (
CONTENT_SCENES_PATH_FIELD,
DEFAULT_SCENE_PATH_PREFIX,
PointNavDatasetV1,
)
from habitat.tasks.nav.nav import NavigationEpisode
@attr.s(auto_attribs=True, kw_only=True)
class RearrangementSpec:
r"""Specifications that capture a particular position of final position
or initial position of the object.
"""
position: List[float] = attr.ib(default=None, validator=not_none_validator)
rotation: List[float] = attr.ib(default=None, validator=not_none_validator)
info: Optional[Dict[str, str]] = attr.ib(default=None)
@attr.s(auto_attribs=True, kw_only=True)
class RearrangementObjectSpec(RearrangementSpec):
r"""Object specifications that capture position of each object in the scene,
the associated object template.
"""
object_id: str = attr.ib(default=None, validator=not_none_validator)
object_template: Optional[str] = attr.ib(
default="data/test_assets/objects/chair"
)
@attr.s(auto_attribs=True, kw_only=True)
class RearrangementEpisode(NavigationEpisode):
r"""Specification of episode that includes initial position and rotation
of agent, all goal specifications, all object specifications
Args:
episode_id: id of episode in the dataset
scene_id: id of scene inside the simulator.
start_position: numpy ndarray containing 3 entries for (x, y, z).
start_rotation: numpy ndarray with 4 entries for (x, y, z, w)
elements of unit quaternion (versor) representing agent 3D
orientation.
goal: object's goal position and rotation
object: object's start specification defined with object type,
position, and rotation.
"""
objects: RearrangementObjectSpec = attr.ib(
default=None, validator=not_none_validator
)
goals: RearrangementSpec = attr.ib(
default=None, validator=not_none_validator
)
@registry.register_dataset(name="RearrangementDataset-v0")
class RearrangementDatasetV0(PointNavDatasetV1):
r"""Class inherited from PointNavDataset that loads Rearrangement dataset."""
episodes: List[RearrangementEpisode]
content_scenes_path: str = "{data_path}/content/{scene}.json.gz"
def to_json(self) -> str:
result = DatasetFloatJSONEncoder().encode(self)
return result
def __init__(self, config: Optional[Config] = None) -> None:
super().__init__(config)
def from_json(
self, json_str: str, scenes_dir: Optional[str] = None
) -> None:
deserialized = json.loads(json_str)
if CONTENT_SCENES_PATH_FIELD in deserialized:
self.content_scenes_path = deserialized[CONTENT_SCENES_PATH_FIELD]
for i, episode in enumerate(deserialized["episodes"]):
rearrangement_episode = RearrangementEpisode(**episode)
rearrangement_episode.episode_id = str(i)
if scenes_dir is not None:
if rearrangement_episode.scene_id.startswith(
DEFAULT_SCENE_PATH_PREFIX
):
rearrangement_episode.scene_id = (
rearrangement_episode.scene_id[
len(DEFAULT_SCENE_PATH_PREFIX) :
]
)
rearrangement_episode.scene_id = os.path.join(
scenes_dir, rearrangement_episode.scene_id
)
rearrangement_episode.objects = RearrangementObjectSpec(
**rearrangement_episode.objects
)
rearrangement_episode.goals = RearrangementSpec(
**rearrangement_episode.goals
)
self.episodes.append(rearrangement_episode)
# @title Load the saved dataset using the Dataset class
config = habitat.get_config("configs/datasets/pointnav/habitat_test.yaml")
config.defrost()
config.DATASET.DATA_PATH = (
"data/datasets/rearrangement/coda/v1/{split}/{split}.json.gz"
)
config.DATASET.TYPE = "RearrangementDataset-v0"
config.freeze()
dataset = RearrangementDatasetV0(config.DATASET)
# check if the dataset got correctly deserialized
assert len(dataset.episodes) == 1
assert dataset.episodes[0].objects.position == [
1.770593523979187,
0.6726829409599304,
-1.9992598295211792,
]
assert dataset.episodes[0].objects.rotation == [0.0, 0.0, 0.0, 1.0]
assert (
dataset.episodes[0].objects.object_template
== "data/test_assets/objects/chair"
)
assert dataset.episodes[0].goals.position == [
4.3417439460754395,
0.6726829409599304,
-5.0634379386901855,
]
assert dataset.episodes[0].goals.rotation == [0.0, 0.0, 0.0, 1.0]
```
## Implement Grab/Release Action
```
# @title RayCast utility to implement Grab/Release Under Cross-Hair Action
# @markdown Cast a ray in the direction of crosshair from the camera and check if it collides with another object within a certain distance threshold
def raycast(sim, sensor_name, crosshair_pos=(128, 128), max_distance=2.0):
r"""Cast a ray in the direction of crosshair and check if it collides
with another object within a certain distance threshold
:param sim: Simulator object
:param sensor_name: name of the visual sensor to be used for raycasting
:param crosshair_pos: 2D coordiante in the viewport towards which the
ray will be cast
:param max_distance: distance threshold beyond which objects won't
be considered
"""
visual_sensor = sim._sensors[sensor_name]
scene_graph = sim.get_active_scene_graph()
scene_graph.set_default_render_camera_parameters(
visual_sensor._sensor_object
)
render_camera = scene_graph.get_default_render_camera()
center_ray = render_camera.unproject(mn.Vector2i(crosshair_pos))
raycast_results = sim.cast_ray(center_ray, max_distance=max_distance)
closest_object = -1
closest_dist = 1000.0
if raycast_results.has_hits():
for hit in raycast_results.hits:
if hit.ray_distance < closest_dist:
closest_dist = hit.ray_distance
closest_object = hit.object_id
return closest_object
# Test the raycast utility.
with habitat_sim.Simulator(cfg) as sim:
init_agent(sim)
obj_attr_mgr = sim.get_object_template_manager()
obj_path = "test_assets/objects/chair"
chair_template_id = obj_attr_mgr.load_object_configs(
str(os.path.join(data_path, obj_path))
)[0]
chair_attr = obj_attr_mgr.get_template_by_ID(chair_template_id)
obj_attr_mgr.register_template(chair_attr)
object_id = sim.add_object_by_handle(chair_attr.handle)
print(f"Chair's object id is {object_id}")
set_object_in_front_of_agent(sim, object_id, -1.5)
sim.set_object_motion_type(
habitat_sim.physics.MotionType.STATIC, object_id
)
if make_video:
# Visualize the agent's initial position
simulate_and_make_vid(
sim, [190, 128], "sim-before-grab", dt=1.0, open_vid=show_video
)
# Distance threshold=2 is greater than agent-to-chair distance.
# Should return chair's object id
closest_object = raycast(
sim, "rgb", crosshair_pos=[128, 190], max_distance=2.0
)
print(f"Closest Object ID: {closest_object} using 2.0 threshold")
assert (
closest_object == object_id
), f"Could not pick chair with ID: {object_id}"
# Distance threshold=1 is smaller than agent-to-chair distance .
# Should return -1
closest_object = raycast(
sim, "rgb", crosshair_pos=[128, 190], max_distance=1.0
)
print(f"Closest Object ID: {closest_object} using 1.0 threshold")
assert closest_object == -1, "Agent shoud not be able to pick any object"
# @title Define a Grab/Release action and create a new action space.
# @markdown Each new action is defined by a `ActionSpec` and an `ActuationSpec`. `ActionSpec` is mapping between the action name and its corresponding `ActuationSpec`. `ActuationSpec` contains all the necessary specifications required to define the action.
from habitat.config.default import _C, CN
from habitat.core.embodied_task import SimulatorTaskAction
from habitat.sims.habitat_simulator.actions import (
HabitatSimActions,
HabitatSimV1ActionSpaceConfiguration,
)
from habitat_sim.agent.controls.controls import ActuationSpec
from habitat_sim.physics import MotionType
# @markdown For instance, `GrabReleaseActuationSpec` contains the following:
# @markdown - `visual_sensor_name` defines which viewport (rgb, depth, etc) to to use to cast the ray.
# @markdown - `crosshair_pos` stores the position in the viewport through which the ray passes. Any object which intersects with this ray can be grabbed by the agent.
# @markdown - `amount` defines a distance threshold. Objects which are farther than the treshold cannot be picked up by the agent.
@attr.s(auto_attribs=True, slots=True)
class GrabReleaseActuationSpec(ActuationSpec):
visual_sensor_name: str = "rgb"
crosshair_pos: List[int] = [128, 128]
amount: float = 2.0
# @markdown Then, we extend the `HabitatSimV1ActionSpaceConfiguration` to add the above action into the agent's action space. `ActionSpaceConfiguration` is a mapping between action name and the corresponding `ActionSpec`
@registry.register_action_space_configuration(name="RearrangementActions-v0")
class RearrangementSimV0ActionSpaceConfiguration(
HabitatSimV1ActionSpaceConfiguration
):
def __init__(self, config):
super().__init__(config)
if not HabitatSimActions.has_action("GRAB_RELEASE"):
HabitatSimActions.extend_action_space("GRAB_RELEASE")
def get(self):
config = super().get()
new_config = {
HabitatSimActions.GRAB_RELEASE: habitat_sim.ActionSpec(
"grab_or_release_object_under_crosshair",
GrabReleaseActuationSpec(
visual_sensor_name=self.config.VISUAL_SENSOR,
crosshair_pos=self.config.CROSSHAIR_POS,
amount=self.config.GRAB_DISTANCE,
),
)
}
config.update(new_config)
return config
# @markdown Finally, we extend `SimualtorTaskAction` which tells the simulator which action to call when a named action ('GRAB_RELEASE' in this case) is predicte by the agent's policy.
@registry.register_task_action
class GrabOrReleaseAction(SimulatorTaskAction):
def step(self, *args: Any, **kwargs: Any):
r"""This method is called from ``Env`` on each ``step``."""
return self._sim.step(HabitatSimActions.GRAB_RELEASE)
_C.TASK.ACTIONS.GRAB_RELEASE = CN()
_C.TASK.ACTIONS.GRAB_RELEASE.TYPE = "GrabOrReleaseAction"
_C.SIMULATOR.CROSSHAIR_POS = [128, 160]
_C.SIMULATOR.GRAB_DISTANCE = 2.0
_C.SIMULATOR.VISUAL_SENSOR = "rgb"
```
##Setup Simulator Class for Rearrangement Task

```
# @title RearrangementSim Class
# @markdown Here we will extend the `HabitatSim` class for the rearrangement task. We will make the following changes:
# @markdown - define a new `_initialize_objects` function which will load the object in its initial configuration as defined by the episode.
# @markdown - define a `gripped_object_id` property that stores whether the agent is holding any object or not.
# @markdown - modify the `step` function of the simulator to use the `grab/release` action we define earlier.
# @markdown #### Writing the `step` function:
# @markdown Since we added a new action for this task, we have to modify the `step` function to define what happens when `grab/release` action is called. If a simple navigation action (`move_forward`, `turn_left`, `turn_right`) is called, we pass it forward to `act` function of the agent which already defines the behavior of these actions.
# @markdown For the `grab/release` action, if the agent is not already holding an object, we first call the `raycast` function using the values from the `ActuationSpec` to see if any object is grippable. If it returns a valid object id, we put the object in a "invisible" inventory and remove it from the scene.
# @markdown If the agent was already holding an object, `grab/release` action will try release the object at the same relative position as it was grabbed. If the object can be placed without any collision, then the `release` action is successful.
from habitat.sims.habitat_simulator.habitat_simulator import HabitatSim
from habitat_sim.nav import NavMeshSettings
from habitat_sim.utils.common import quat_from_coeffs, quat_to_magnum
@registry.register_simulator(name="RearrangementSim-v0")
class RearrangementSim(HabitatSim):
r"""Simulator wrapper over habitat-sim with
object rearrangement functionalities.
"""
def __init__(self, config: Config) -> None:
self.did_reset = False
super().__init__(config=config)
self.grip_offset = np.eye(4)
agent_id = self.habitat_config.DEFAULT_AGENT_ID
agent_config = self._get_agent_config(agent_id)
self.navmesh_settings = NavMeshSettings()
self.navmesh_settings.set_defaults()
self.navmesh_settings.agent_radius = agent_config.RADIUS
self.navmesh_settings.agent_height = agent_config.HEIGHT
def reconfigure(self, config: Config) -> None:
super().reconfigure(config)
self._initialize_objects()
def reset(self):
sim_obs = super().reset()
if self._update_agents_state():
sim_obs = self.get_sensor_observations()
self._prev_sim_obs = sim_obs
self.did_reset = True
self.grip_offset = np.eye(4)
return self._sensor_suite.get_observations(sim_obs)
def _initialize_objects(self):
objects = self.habitat_config.objects[0]
obj_attr_mgr = self.get_object_template_manager()
# first remove all existing objects
existing_object_ids = self.get_existing_object_ids()
if len(existing_object_ids) > 0:
for obj_id in existing_object_ids:
self.remove_object(obj_id)
self.sim_object_to_objid_mapping = {}
self.objid_to_sim_object_mapping = {}
if objects is not None:
object_template = objects["object_template"]
object_pos = objects["position"]
object_rot = objects["rotation"]
object_template_id = obj_attr_mgr.load_object_configs(
object_template
)[0]
object_attr = obj_attr_mgr.get_template_by_ID(object_template_id)
obj_attr_mgr.register_template(object_attr)
object_id = self.add_object_by_handle(object_attr.handle)
self.sim_object_to_objid_mapping[object_id] = objects["object_id"]
self.objid_to_sim_object_mapping[objects["object_id"]] = object_id
self.set_translation(object_pos, object_id)
if isinstance(object_rot, list):
object_rot = quat_from_coeffs(object_rot)
object_rot = quat_to_magnum(object_rot)
self.set_rotation(object_rot, object_id)
self.set_object_motion_type(MotionType.STATIC, object_id)
# Recompute the navmesh after placing all the objects.
self.recompute_navmesh(self.pathfinder, self.navmesh_settings, True)
def _sync_gripped_object(self, gripped_object_id):
r"""
Sync the gripped object with the object associated with the agent.
"""
if gripped_object_id != -1:
agent_body_transformation = (
self._default_agent.scene_node.transformation
)
self.set_transformation(
agent_body_transformation, gripped_object_id
)
translation = agent_body_transformation.transform_point(
np.array([0, 2.0, 0])
)
self.set_translation(translation, gripped_object_id)
@property
def gripped_object_id(self):
return self._prev_sim_obs.get("gripped_object_id", -1)
def step(self, action: int):
dt = 1 / 60.0
self._num_total_frames += 1
collided = False
gripped_object_id = self.gripped_object_id
agent_config = self._default_agent.agent_config
action_spec = agent_config.action_space[action]
if action_spec.name == "grab_or_release_object_under_crosshair":
# If already holding an agent
if gripped_object_id != -1:
agent_body_transformation = (
self._default_agent.scene_node.transformation
)
T = np.dot(agent_body_transformation, self.grip_offset)
self.set_transformation(T, gripped_object_id)
position = self.get_translation(gripped_object_id)
if self.pathfinder.is_navigable(position):
self.set_object_motion_type(
MotionType.STATIC, gripped_object_id
)
gripped_object_id = -1
self.recompute_navmesh(
self.pathfinder, self.navmesh_settings, True
)
# if not holding an object, then try to grab
else:
gripped_object_id = raycast(
self,
action_spec.actuation.visual_sensor_name,
crosshair_pos=action_spec.actuation.crosshair_pos,
max_distance=action_spec.actuation.amount,
)
# found a grabbable object.
if gripped_object_id != -1:
agent_body_transformation = (
self._default_agent.scene_node.transformation
)
self.grip_offset = np.dot(
np.array(agent_body_transformation.inverted()),
np.array(self.get_transformation(gripped_object_id)),
)
self.set_object_motion_type(
MotionType.KINEMATIC, gripped_object_id
)
self.recompute_navmesh(
self.pathfinder, self.navmesh_settings, True
)
else:
collided = self._default_agent.act(action)
self._last_state = self._default_agent.get_state()
# step physics by dt
super().step_world(dt)
# Sync the gripped object after the agent moves.
self._sync_gripped_object(gripped_object_id)
# obtain observations
self._prev_sim_obs = self.get_sensor_observations()
self._prev_sim_obs["collided"] = collided
self._prev_sim_obs["gripped_object_id"] = gripped_object_id
observations = self._sensor_suite.get_observations(self._prev_sim_obs)
return observations
```
## Create the Rearrangement Task

```
# @title Implement new sensors and measurements
# @markdown After defining the dataset, action space and simulator functions for the rearrangement task, we are one step closer to training agents to solve this task.
# @markdown Here we define inputs to the policy and other measurements required to design reward functions.
# @markdown **Sensors**: These define various part of the simulator state that's visible to the agent. For simplicity, we'll assume that agent knows the object's current position, object's final goal position relative to the agent's current position.
# @markdown - Object's current position will be made given by the `ObjectPosition` sensor
# @markdown - Object's goal position will be available through the `ObjectGoal` sensor.
# @markdown - Finally, we will also use `GrippedObject` sensor to tell the agent if it's holding any object or not.
# @markdown **Measures**: These define various metrics about the task which can be used to measure task progress and define rewards. Note that measurements are *privileged* information not accessible to the agent as part of the observation space. We will need the following measurements:
# @markdown - `AgentToObjectDistance` which measure the euclidean distance between the agent and the object.
# @markdown - `ObjectToGoalDistance` which measures the euclidean distance between the object and the goal.
from gym import spaces
import habitat_sim
from habitat.config.default import CN, Config
from habitat.core.dataset import Episode
from habitat.core.embodied_task import Measure
from habitat.core.simulator import Observations, Sensor, SensorTypes, Simulator
from habitat.tasks.nav.nav import PointGoalSensor
@registry.register_sensor
class GrippedObjectSensor(Sensor):
cls_uuid = "gripped_object_id"
def __init__(
self, *args: Any, sim: RearrangementSim, config: Config, **kwargs: Any
):
self._sim = sim
super().__init__(config=config)
def _get_uuid(self, *args: Any, **kwargs: Any) -> str:
return self.cls_uuid
def _get_observation_space(self, *args: Any, **kwargs: Any):
return spaces.Discrete(self._sim.get_existing_object_ids())
def _get_sensor_type(self, *args: Any, **kwargs: Any):
return SensorTypes.MEASUREMENT
def get_observation(
self,
observations: Dict[str, Observations],
episode: Episode,
*args: Any,
**kwargs: Any,
):
obj_id = self._sim.sim_object_to_objid_mapping.get(
self._sim.gripped_object_id, -1
)
return obj_id
@registry.register_sensor
class ObjectPosition(PointGoalSensor):
cls_uuid: str = "object_position"
def _get_observation_space(self, *args: Any, **kwargs: Any):
sensor_shape = (self._dimensionality,)
return spaces.Box(
low=np.finfo(np.float32).min,
high=np.finfo(np.float32).max,
shape=sensor_shape,
dtype=np.float32,
)
def get_observation(
self, *args: Any, observations, episode, **kwargs: Any
):
agent_state = self._sim.get_agent_state()
agent_position = agent_state.position
rotation_world_agent = agent_state.rotation
object_id = self._sim.get_existing_object_ids()[0]
object_position = self._sim.get_translation(object_id)
pointgoal = self._compute_pointgoal(
agent_position, rotation_world_agent, object_position
)
return pointgoal
@registry.register_sensor
class ObjectGoal(PointGoalSensor):
cls_uuid: str = "object_goal"
def _get_observation_space(self, *args: Any, **kwargs: Any):
sensor_shape = (self._dimensionality,)
return spaces.Box(
low=np.finfo(np.float32).min,
high=np.finfo(np.float32).max,
shape=sensor_shape,
dtype=np.float32,
)
def get_observation(
self, *args: Any, observations, episode, **kwargs: Any
):
agent_state = self._sim.get_agent_state()
agent_position = agent_state.position
rotation_world_agent = agent_state.rotation
goal_position = np.array(episode.goals.position, dtype=np.float32)
point_goal = self._compute_pointgoal(
agent_position, rotation_world_agent, goal_position
)
return point_goal
@registry.register_measure
class ObjectToGoalDistance(Measure):
"""The measure calculates distance of object towards the goal."""
cls_uuid: str = "object_to_goal_distance"
def __init__(
self, sim: Simulator, config: Config, *args: Any, **kwargs: Any
):
self._sim = sim
self._config = config
super().__init__(**kwargs)
@staticmethod
def _get_uuid(*args: Any, **kwargs: Any):
return ObjectToGoalDistance.cls_uuid
def reset_metric(self, episode, *args: Any, **kwargs: Any):
self.update_metric(*args, episode=episode, **kwargs)
def _geo_dist(self, src_pos, goal_pos: np.array) -> float:
return self._sim.geodesic_distance(src_pos, [goal_pos])
def _euclidean_distance(self, position_a, position_b):
return np.linalg.norm(
np.array(position_b) - np.array(position_a), ord=2
)
def update_metric(self, episode, *args: Any, **kwargs: Any):
sim_obj_id = self._sim.get_existing_object_ids()[0]
previous_position = np.array(
self._sim.get_translation(sim_obj_id)
).tolist()
goal_position = episode.goals.position
self._metric = self._euclidean_distance(
previous_position, goal_position
)
@registry.register_measure
class AgentToObjectDistance(Measure):
"""The measure calculates the distance of objects from the agent"""
cls_uuid: str = "agent_to_object_distance"
def __init__(
self, sim: Simulator, config: Config, *args: Any, **kwargs: Any
):
self._sim = sim
self._config = config
super().__init__(**kwargs)
@staticmethod
def _get_uuid(*args: Any, **kwargs: Any):
return AgentToObjectDistance.cls_uuid
def reset_metric(self, episode, *args: Any, **kwargs: Any):
self.update_metric(*args, episode=episode, **kwargs)
def _euclidean_distance(self, position_a, position_b):
return np.linalg.norm(
np.array(position_b) - np.array(position_a), ord=2
)
def update_metric(self, episode, *args: Any, **kwargs: Any):
sim_obj_id = self._sim.get_existing_object_ids()[0]
previous_position = np.array(
self._sim.get_translation(sim_obj_id)
).tolist()
agent_state = self._sim.get_agent_state()
agent_position = agent_state.position
self._metric = self._euclidean_distance(
previous_position, agent_position
)
# -----------------------------------------------------------------------------
# # REARRANGEMENT TASK GRIPPED OBJECT SENSOR
# -----------------------------------------------------------------------------
_C.TASK.GRIPPED_OBJECT_SENSOR = CN()
_C.TASK.GRIPPED_OBJECT_SENSOR.TYPE = "GrippedObjectSensor"
# -----------------------------------------------------------------------------
# # REARRANGEMENT TASK ALL OBJECT POSITIONS SENSOR
# -----------------------------------------------------------------------------
_C.TASK.OBJECT_POSITION = CN()
_C.TASK.OBJECT_POSITION.TYPE = "ObjectPosition"
_C.TASK.OBJECT_POSITION.GOAL_FORMAT = "POLAR"
_C.TASK.OBJECT_POSITION.DIMENSIONALITY = 2
# -----------------------------------------------------------------------------
# # REARRANGEMENT TASK ALL OBJECT GOALS SENSOR
# -----------------------------------------------------------------------------
_C.TASK.OBJECT_GOAL = CN()
_C.TASK.OBJECT_GOAL.TYPE = "ObjectGoal"
_C.TASK.OBJECT_GOAL.GOAL_FORMAT = "POLAR"
_C.TASK.OBJECT_GOAL.DIMENSIONALITY = 2
# -----------------------------------------------------------------------------
# # OBJECT_DISTANCE_TO_GOAL MEASUREMENT
# -----------------------------------------------------------------------------
_C.TASK.OBJECT_TO_GOAL_DISTANCE = CN()
_C.TASK.OBJECT_TO_GOAL_DISTANCE.TYPE = "ObjectToGoalDistance"
# -----------------------------------------------------------------------------
# # OBJECT_DISTANCE_FROM_AGENT MEASUREMENT
# -----------------------------------------------------------------------------
_C.TASK.AGENT_TO_OBJECT_DISTANCE = CN()
_C.TASK.AGENT_TO_OBJECT_DISTANCE.TYPE = "AgentToObjectDistance"
from habitat.config.default import CN, Config
# @title Define `RearrangementTask` by extending `NavigationTask`
from habitat.tasks.nav.nav import NavigationTask, merge_sim_episode_config
def merge_sim_episode_with_object_config(
sim_config: Config, episode: Type[Episode]
) -> Any:
sim_config = merge_sim_episode_config(sim_config, episode)
sim_config.defrost()
sim_config.objects = [episode.objects.__dict__]
sim_config.freeze()
return sim_config
@registry.register_task(name="RearrangementTask-v0")
class RearrangementTask(NavigationTask):
r"""Embodied Rearrangement Task
Goal: An agent must place objects at their corresponding goal position.
"""
def __init__(self, **kwargs) -> None:
super().__init__(**kwargs)
def overwrite_sim_config(self, sim_config, episode):
return merge_sim_episode_with_object_config(sim_config, episode)
```
## Implement a hard-coded and an RL agent
```
# @title Load the `RearrangementTask` in Habitat-Lab and run a hard-coded agent
import habitat
config = habitat.get_config("configs/tasks/pointnav.yaml")
config.defrost()
config.ENVIRONMENT.MAX_EPISODE_STEPS = 50
config.SIMULATOR.TYPE = "RearrangementSim-v0"
config.SIMULATOR.ACTION_SPACE_CONFIG = "RearrangementActions-v0"
config.SIMULATOR.GRAB_DISTANCE = 2.0
config.SIMULATOR.HABITAT_SIM_V0.ENABLE_PHYSICS = True
config.TASK.TYPE = "RearrangementTask-v0"
config.TASK.SUCCESS_DISTANCE = 1.0
config.TASK.SENSORS = [
"GRIPPED_OBJECT_SENSOR",
"OBJECT_POSITION",
"OBJECT_GOAL",
]
config.TASK.GOAL_SENSOR_UUID = "object_goal"
config.TASK.MEASUREMENTS = [
"OBJECT_TO_GOAL_DISTANCE",
"AGENT_TO_OBJECT_DISTANCE",
]
config.TASK.POSSIBLE_ACTIONS = ["STOP", "MOVE_FORWARD", "GRAB_RELEASE"]
config.DATASET.TYPE = "RearrangementDataset-v0"
config.DATASET.SPLIT = "train"
config.DATASET.DATA_PATH = (
"data/datasets/rearrangement/coda/v1/{split}/{split}.json.gz"
)
config.freeze()
def print_info(obs, metrics):
print(
"Gripped Object: {}, Distance To Object: {}, Distance To Goal: {}".format(
obs["gripped_object_id"],
metrics["agent_to_object_distance"],
metrics["object_to_goal_distance"],
)
)
try: # Got to make initialization idiot proof
sim.close()
except NameError:
pass
with habitat.Env(config) as env:
obs = env.reset()
obs_list = []
# Get closer to the object
while True:
obs = env.step(1)
obs_list.append(obs)
metrics = env.get_metrics()
print_info(obs, metrics)
if metrics["agent_to_object_distance"] < 2.0:
break
# Grab the object
obs = env.step(2)
obs_list.append(obs)
metrics = env.get_metrics()
print_info(obs, metrics)
assert obs["gripped_object_id"] != -1
# Get closer to the goal
while True:
obs = env.step(1)
obs_list.append(obs)
metrics = env.get_metrics()
print_info(obs, metrics)
if metrics["object_to_goal_distance"] < 2.0:
break
# Release the object
obs = env.step(2)
obs_list.append(obs)
metrics = env.get_metrics()
print_info(obs, metrics)
assert obs["gripped_object_id"] == -1
if make_video:
make_video_cv2(
obs_list,
[190, 128],
"hard-coded-agent",
fps=5.0,
open_vid=show_video,
)
# @title Create a task specific RL Environment with a new reward definition.
# @markdown We create a `RearragenmentRLEnv` class and modify the `get_reward()` function.
# @markdown The reward sturcture is as follows:
# @markdown - The agent gets a positive reward if the agent gets closer to the object otherwise a negative reward.
# @markdown - The agent gets a positive reward if it moves the object closer to goal otherwise a negative reward.
# @markdown - The agent gets a positive reward when the agent "picks" up an object for the first time. For all other "grab/release" action, it gets a negative reward.
# @markdown - The agent gets a slack penalty of -0.01 for every action it takes in the environment.
# @markdown - Finally the agent gets a large success reward when the episode is completed successfully.
from typing import Optional, Type
import numpy as np
import habitat
from habitat import Config, Dataset
from habitat_baselines.common.baseline_registry import baseline_registry
from habitat_baselines.common.environments import NavRLEnv
@baseline_registry.register_env(name="RearrangementRLEnv")
class RearrangementRLEnv(NavRLEnv):
def __init__(self, config: Config, dataset: Optional[Dataset] = None):
self._prev_measure = {
"agent_to_object_distance": 0.0,
"object_to_goal_distance": 0.0,
"gripped_object_id": -1,
"gripped_object_count": 0,
}
super().__init__(config, dataset)
self._success_distance = self._core_env_config.TASK.SUCCESS_DISTANCE
def reset(self):
self._previous_action = None
observations = super().reset()
self._prev_measure.update(self.habitat_env.get_metrics())
self._prev_measure["gripped_object_id"] = -1
self._prev_measure["gripped_object_count"] = 0
return observations
def step(self, *args, **kwargs):
self._previous_action = kwargs["action"]
return super().step(*args, **kwargs)
def get_reward_range(self):
return (
self._rl_config.SLACK_REWARD - 1.0,
self._rl_config.SUCCESS_REWARD + 1.0,
)
def get_reward(self, observations):
reward = self._rl_config.SLACK_REWARD
gripped_success_reward = 0.0
episode_success_reward = 0.0
agent_to_object_dist_reward = 0.0
object_to_goal_dist_reward = 0.0
action_name = self._env.task.get_action_name(
self._previous_action["action"]
)
# If object grabbed, add a success reward
# The reward gets awarded only once for an object.
if (
action_name == "GRAB_RELEASE"
and observations["gripped_object_id"] >= 0
):
obj_id = observations["gripped_object_id"]
self._prev_measure["gripped_object_count"] += 1
gripped_success_reward = (
self._rl_config.GRIPPED_SUCCESS_REWARD
if self._prev_measure["gripped_object_count"] == 1
else 0.0
)
# add a penalty everytime grab/action is called and doesn't do anything
elif action_name == "GRAB_RELEASE":
gripped_success_reward += -0.1
self._prev_measure["gripped_object_id"] = observations[
"gripped_object_id"
]
# If the action is not a grab/release action, and the agent
# has not picked up an object, then give reward based on agent to
# object distance.
if (
action_name != "GRAB_RELEASE"
and self._prev_measure["gripped_object_id"] == -1
):
agent_to_object_dist_reward = self.get_agent_to_object_dist_reward(
observations
)
# If the action is not a grab/release action, and the agent
# has picked up an object, then give reward based on object to
# to goal distance.
if (
action_name != "GRAB_RELEASE"
and self._prev_measure["gripped_object_id"] != -1
):
object_to_goal_dist_reward = self.get_object_to_goal_dist_reward()
if (
self._episode_success(observations)
and self._prev_measure["gripped_object_id"] == -1
and action_name == "STOP"
):
episode_success_reward = self._rl_config.SUCCESS_REWARD
reward += (
agent_to_object_dist_reward
+ object_to_goal_dist_reward
+ gripped_success_reward
+ episode_success_reward
)
return reward
def get_agent_to_object_dist_reward(self, observations):
"""
Encourage the agent to move towards the closest object which is not already in place.
"""
curr_metric = self._env.get_metrics()["agent_to_object_distance"]
prev_metric = self._prev_measure["agent_to_object_distance"]
dist_reward = prev_metric - curr_metric
self._prev_measure["agent_to_object_distance"] = curr_metric
return dist_reward
def get_object_to_goal_dist_reward(self):
curr_metric = self._env.get_metrics()["object_to_goal_distance"]
prev_metric = self._prev_measure["object_to_goal_distance"]
dist_reward = prev_metric - curr_metric
self._prev_measure["object_to_goal_distance"] = curr_metric
return dist_reward
def _episode_success(self, observations):
r"""Returns True if object is within distance threshold of the goal."""
dist = self._env.get_metrics()["object_to_goal_distance"]
if (
abs(dist) > self._success_distance
or observations["gripped_object_id"] != -1
):
return False
return True
def _gripped_success(self, observations):
if (
observations["gripped_object_id"] >= 0
and observations["gripped_object_id"]
!= self._prev_measure["gripped_object_id"]
):
return True
return False
def get_done(self, observations):
done = False
action_name = self._env.task.get_action_name(
self._previous_action["action"]
)
if self._env.episode_over or (
self._episode_success(observations)
and self._prev_measure["gripped_object_id"] == -1
and action_name == "STOP"
):
done = True
return done
def get_info(self, observations):
info = self.habitat_env.get_metrics()
info["episode_success"] = self._episode_success(observations)
return info
import os
import time
from collections import defaultdict, deque
from typing import Any, Dict, List, Optional
import numpy as np
from torch.optim.lr_scheduler import LambdaLR
from habitat import Config, logger
from habitat.utils.visualizations.utils import observations_to_image
from habitat_baselines.common.baseline_registry import baseline_registry
from habitat_baselines.common.environments import get_env_class
from habitat_baselines.common.rollout_storage import RolloutStorage
from habitat_baselines.common.tensorboard_utils import TensorboardWriter
from habitat_baselines.rl.models.rnn_state_encoder import RNNStateEncoder
from habitat_baselines.rl.ppo import PPO
from habitat_baselines.rl.ppo.policy import Net, Policy
from habitat_baselines.rl.ppo.ppo_trainer import PPOTrainer
from habitat_baselines.utils.common import (
batch_obs,
generate_video,
linear_decay,
)
from habitat_baselines.utils.env_utils import make_env_fn
def construct_envs(
config,
env_class,
workers_ignore_signals=False,
):
r"""Create VectorEnv object with specified config and env class type.
To allow better performance, dataset are split into small ones for
each individual env, grouped by scenes.
:param config: configs that contain num_processes as well as information
:param necessary to create individual environments.
:param env_class: class type of the envs to be created.
:param workers_ignore_signals: Passed to :ref:`habitat.VectorEnv`'s constructor
:return: VectorEnv object created according to specification.
"""
num_processes = config.NUM_PROCESSES
configs = []
env_classes = [env_class for _ in range(num_processes)]
dataset = habitat.datasets.make_dataset(config.TASK_CONFIG.DATASET.TYPE)
scenes = config.TASK_CONFIG.DATASET.CONTENT_SCENES
if "*" in config.TASK_CONFIG.DATASET.CONTENT_SCENES:
scenes = dataset.get_scenes_to_load(config.TASK_CONFIG.DATASET)
if num_processes > 1:
if len(scenes) == 0:
raise RuntimeError(
"No scenes to load, multiple process logic relies on being able to split scenes uniquely between processes"
)
if len(scenes) < num_processes:
scenes = scenes * num_processes
random.shuffle(scenes)
scene_splits = [[] for _ in range(num_processes)]
for idx, scene in enumerate(scenes):
scene_splits[idx % len(scene_splits)].append(scene)
assert sum(map(len, scene_splits)) == len(scenes)
for i in range(num_processes):
proc_config = config.clone()
proc_config.defrost()
task_config = proc_config.TASK_CONFIG
task_config.SEED = task_config.SEED + i
if len(scenes) > 0:
task_config.DATASET.CONTENT_SCENES = scene_splits[i]
task_config.SIMULATOR.HABITAT_SIM_V0.GPU_DEVICE_ID = (
config.SIMULATOR_GPU_ID
)
task_config.SIMULATOR.AGENT_0.SENSORS = config.SENSORS
proc_config.freeze()
configs.append(proc_config)
envs = habitat.ThreadedVectorEnv(
make_env_fn=make_env_fn,
env_fn_args=tuple(zip(configs, env_classes)),
workers_ignore_signals=workers_ignore_signals,
)
return envs
class RearrangementBaselinePolicy(Policy):
def __init__(self, observation_space, action_space, hidden_size=512):
super().__init__(
RearrangementBaselineNet(
observation_space=observation_space, hidden_size=hidden_size
),
action_space.n,
)
def from_config(cls, config, envs):
pass
class RearrangementBaselineNet(Net):
r"""Network which passes the input image through CNN and concatenates
goal vector with CNN's output and passes that through RNN.
"""
def __init__(self, observation_space, hidden_size):
super().__init__()
self._n_input_goal = observation_space.spaces[
ObjectGoal.cls_uuid
].shape[0]
self._hidden_size = hidden_size
self.state_encoder = RNNStateEncoder(
2 * self._n_input_goal,
self._hidden_size,
)
self.train()
@property
def output_size(self):
return self._hidden_size
@property
def is_blind(self):
return False
@property
def num_recurrent_layers(self):
return self.state_encoder.num_recurrent_layers
def forward(self, observations, rnn_hidden_states, prev_actions, masks):
object_goal_encoding = observations[ObjectGoal.cls_uuid]
object_pos_encoding = observations[ObjectPosition.cls_uuid]
x = [object_goal_encoding, object_pos_encoding]
x = torch.cat(x, dim=1)
x, rnn_hidden_states = self.state_encoder(x, rnn_hidden_states, masks)
return x, rnn_hidden_states
@baseline_registry.register_trainer(name="ppo-rearrangement")
class RearrangementTrainer(PPOTrainer):
supported_tasks = ["RearrangementTask-v0"]
def _setup_actor_critic_agent(self, ppo_cfg: Config) -> None:
r"""Sets up actor critic and agent for PPO.
Args:
ppo_cfg: config node with relevant params
Returns:
None
"""
logger.add_filehandler(self.config.LOG_FILE)
self.actor_critic = RearrangementBaselinePolicy(
observation_space=self.envs.observation_spaces[0],
action_space=self.envs.action_spaces[0],
hidden_size=ppo_cfg.hidden_size,
)
self.actor_critic.to(self.device)
self.agent = PPO(
actor_critic=self.actor_critic,
clip_param=ppo_cfg.clip_param,
ppo_epoch=ppo_cfg.ppo_epoch,
num_mini_batch=ppo_cfg.num_mini_batch,
value_loss_coef=ppo_cfg.value_loss_coef,
entropy_coef=ppo_cfg.entropy_coef,
lr=ppo_cfg.lr,
eps=ppo_cfg.eps,
max_grad_norm=ppo_cfg.max_grad_norm,
use_normalized_advantage=ppo_cfg.use_normalized_advantage,
)
def train(self) -> None:
r"""Main method for training PPO.
Returns:
None
"""
self.envs = construct_envs(
self.config, get_env_class(self.config.ENV_NAME)
)
ppo_cfg = self.config.RL.PPO
self.device = (
torch.device("cuda", self.config.TORCH_GPU_ID)
if torch.cuda.is_available()
else torch.device("cpu")
)
if not os.path.isdir(self.config.CHECKPOINT_FOLDER):
os.makedirs(self.config.CHECKPOINT_FOLDER)
self._setup_actor_critic_agent(ppo_cfg)
logger.info(
"agent number of parameters: {}".format(
sum(param.numel() for param in self.agent.parameters())
)
)
rollouts = RolloutStorage(
ppo_cfg.num_steps,
self.envs.num_envs,
self.envs.observation_spaces[0],
self.envs.action_spaces[0],
ppo_cfg.hidden_size,
)
rollouts.to(self.device)
observations = self.envs.reset()
batch = batch_obs(observations, device=self.device)
for sensor in rollouts.observations:
rollouts.observations[sensor][0].copy_(batch[sensor])
# batch and observations may contain shared PyTorch CUDA
# tensors. We must explicitly clear them here otherwise
# they will be kept in memory for the entire duration of training!
batch = None
observations = None
current_episode_reward = torch.zeros(self.envs.num_envs, 1)
running_episode_stats = dict(
count=torch.zeros(self.envs.num_envs, 1),
reward=torch.zeros(self.envs.num_envs, 1),
)
window_episode_stats = defaultdict(
lambda: deque(maxlen=ppo_cfg.reward_window_size)
)
t_start = time.time()
env_time = 0
pth_time = 0
count_steps = 0
count_checkpoints = 0
lr_scheduler = LambdaLR(
optimizer=self.agent.optimizer,
lr_lambda=lambda x: linear_decay(x, self.config.NUM_UPDATES),
)
with TensorboardWriter(
self.config.TENSORBOARD_DIR, flush_secs=self.flush_secs
) as writer:
for update in range(self.config.NUM_UPDATES):
if ppo_cfg.use_linear_lr_decay:
lr_scheduler.step()
if ppo_cfg.use_linear_clip_decay:
self.agent.clip_param = ppo_cfg.clip_param * linear_decay(
update, self.config.NUM_UPDATES
)
for _step in range(ppo_cfg.num_steps):
(
delta_pth_time,
delta_env_time,
delta_steps,
) = self._collect_rollout_step(
rollouts, current_episode_reward, running_episode_stats
)
pth_time += delta_pth_time
env_time += delta_env_time
count_steps += delta_steps
(
delta_pth_time,
value_loss,
action_loss,
dist_entropy,
) = self._update_agent(ppo_cfg, rollouts)
pth_time += delta_pth_time
for k, v in running_episode_stats.items():
window_episode_stats[k].append(v.clone())
deltas = {
k: (
(v[-1] - v[0]).sum().item()
if len(v) > 1
else v[0].sum().item()
)
for k, v in window_episode_stats.items()
}
deltas["count"] = max(deltas["count"], 1.0)
writer.add_scalar(
"reward", deltas["reward"] / deltas["count"], count_steps
)
# Check to see if there are any metrics
# that haven't been logged yet
for k, v in deltas.items():
if k not in {"reward", "count"}:
writer.add_scalar(
"metric/" + k, v / deltas["count"], count_steps
)
losses = [value_loss, action_loss]
for l, k in zip(losses, ["value, policy"]):
writer.add_scalar("losses/" + k, l, count_steps)
# log stats
if update > 0 and update % self.config.LOG_INTERVAL == 0:
logger.info(
"update: {}\tfps: {:.3f}\t".format(
update, count_steps / (time.time() - t_start)
)
)
logger.info(
"update: {}\tenv-time: {:.3f}s\tpth-time: {:.3f}s\t"
"frames: {}".format(
update, env_time, pth_time, count_steps
)
)
logger.info(
"Average window size: {} {}".format(
len(window_episode_stats["count"]),
" ".join(
"{}: {:.3f}".format(k, v / deltas["count"])
for k, v in deltas.items()
if k != "count"
),
)
)
# checkpoint model
if update % self.config.CHECKPOINT_INTERVAL == 0:
self.save_checkpoint(
f"ckpt.{count_checkpoints}.pth", dict(step=count_steps)
)
count_checkpoints += 1
self.envs.close()
def eval(self) -> None:
r"""Evaluates the current model
Returns:
None
"""
config = self.config.clone()
if len(self.config.VIDEO_OPTION) > 0:
config.defrost()
config.NUM_PROCESSES = 1
config.freeze()
logger.info(f"env config: {config}")
with construct_envs(config, get_env_class(config.ENV_NAME)) as envs:
observations = envs.reset()
batch = batch_obs(observations, device=self.device)
current_episode_reward = torch.zeros(
envs.num_envs, 1, device=self.device
)
ppo_cfg = self.config.RL.PPO
test_recurrent_hidden_states = torch.zeros(
self.actor_critic.net.num_recurrent_layers,
config.NUM_PROCESSES,
ppo_cfg.hidden_size,
device=self.device,
)
prev_actions = torch.zeros(
config.NUM_PROCESSES, 1, device=self.device, dtype=torch.long
)
not_done_masks = torch.zeros(
config.NUM_PROCESSES, 1, device=self.device
)
rgb_frames = [
[] for _ in range(self.config.NUM_PROCESSES)
] # type: List[List[np.ndarray]]
if len(config.VIDEO_OPTION) > 0:
os.makedirs(config.VIDEO_DIR, exist_ok=True)
self.actor_critic.eval()
for _i in range(config.TASK_CONFIG.ENVIRONMENT.MAX_EPISODE_STEPS):
current_episodes = envs.current_episodes()
with torch.no_grad():
(
_,
actions,
_,
test_recurrent_hidden_states,
) = self.actor_critic.act(
batch,
test_recurrent_hidden_states,
prev_actions,
not_done_masks,
deterministic=False,
)
prev_actions.copy_(actions)
outputs = envs.step([a[0].item() for a in actions])
observations, rewards, dones, infos = [
list(x) for x in zip(*outputs)
]
batch = batch_obs(observations, device=self.device)
not_done_masks = torch.tensor(
[[0.0] if done else [1.0] for done in dones],
dtype=torch.float,
device=self.device,
)
rewards = torch.tensor(
rewards, dtype=torch.float, device=self.device
).unsqueeze(1)
current_episode_reward += rewards
# episode ended
if not_done_masks[0].item() == 0:
generate_video(
video_option=self.config.VIDEO_OPTION,
video_dir=self.config.VIDEO_DIR,
images=rgb_frames[0],
episode_id=current_episodes[0].episode_id,
checkpoint_idx=0,
metrics=self._extract_scalars_from_info(infos[0]),
tb_writer=None,
)
print("Evaluation Finished.")
print("Success: {}".format(infos[0]["episode_success"]))
print(
"Reward: {}".format(current_episode_reward[0].item())
)
print(
"Distance To Goal: {}".format(
infos[0]["object_to_goal_distance"]
)
)
return
# episode continues
elif len(self.config.VIDEO_OPTION) > 0:
frame = observations_to_image(observations[0], infos[0])
rgb_frames[0].append(frame)
%load_ext tensorboard
%tensorboard --logdir data/tb
# @title Train an RL agent on a single episode
!if [ -d "data/tb" ]; then rm -r data/tb; fi
import random
import numpy as np
import torch
import habitat
from habitat import Config
from habitat_baselines.config.default import get_config as get_baseline_config
baseline_config = get_baseline_config(
"habitat_baselines/config/pointnav/ppo_pointnav.yaml"
)
baseline_config.defrost()
baseline_config.TASK_CONFIG = config
baseline_config.TRAINER_NAME = "ddppo"
baseline_config.ENV_NAME = "RearrangementRLEnv"
baseline_config.SIMULATOR_GPU_ID = 0
baseline_config.TORCH_GPU_ID = 0
baseline_config.VIDEO_OPTION = ["disk"]
baseline_config.TENSORBOARD_DIR = "data/tb"
baseline_config.VIDEO_DIR = "data/videos"
baseline_config.NUM_PROCESSES = 2
baseline_config.SENSORS = ["RGB_SENSOR", "DEPTH_SENSOR"]
baseline_config.CHECKPOINT_FOLDER = "data/checkpoints"
if vut.is_notebook():
baseline_config.NUM_UPDATES = 400 # @param {type:"number"}
else:
baseline_config.NUM_UPDATES = 1
baseline_config.LOG_INTERVAL = 10
baseline_config.CHECKPOINT_INTERVAL = 50
baseline_config.LOG_FILE = "data/checkpoints/train.log"
baseline_config.EVAL.SPLIT = "train"
baseline_config.RL.SUCCESS_REWARD = 2.5 # @param {type:"number"}
baseline_config.RL.SUCCESS_MEASURE = "object_to_goal_distance"
baseline_config.RL.REWARD_MEASURE = "object_to_goal_distance"
baseline_config.RL.GRIPPED_SUCCESS_REWARD = 2.5 # @param {type:"number"}
baseline_config.freeze()
random.seed(baseline_config.TASK_CONFIG.SEED)
np.random.seed(baseline_config.TASK_CONFIG.SEED)
torch.manual_seed(baseline_config.TASK_CONFIG.SEED)
if __name__ == "__main__":
trainer = RearrangementTrainer(baseline_config)
trainer.train()
trainer.eval()
if make_video:
video_file = os.listdir("data/videos")[0]
vut.display_video(os.path.join("data/videos", video_file))
```
| github_jupyter |
# Language Translation
In this project, you’re going to take a peek into the realm of neural network machine translation. You’ll be training a sequence to sequence model on a dataset of English and French sentences that can translate new sentences from English to French.
## Get the Data
Since translating the whole language of English to French will take lots of time to train, we have provided you with a small portion of the English corpus.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
import helper
import problem_unittests as tests
source_path = 'data/small_vocab_en'
target_path = 'data/small_vocab_fr'
source_text = helper.load_data(source_path)
target_text = helper.load_data(target_path)
```
## Explore the Data
Play around with view_sentence_range to view different parts of the data.
```
view_sentence_range = (0, 10)
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
import numpy as np
print('Dataset Stats')
print('Roughly the number of unique words: {}'.format(len({word: None for word in source_text.split()})))
sentences = source_text.split('\n')
word_counts = [len(sentence.split()) for sentence in sentences]
print('Number of sentences: {}'.format(len(sentences)))
print('Average number of words in a sentence: {}'.format(np.average(word_counts)))
print()
print('English sentences {} to {}:'.format(*view_sentence_range))
print('\n'.join(source_text.split('\n')[view_sentence_range[0]:view_sentence_range[1]]))
print()
print('French sentences {} to {}:'.format(*view_sentence_range))
print('\n'.join(target_text.split('\n')[view_sentence_range[0]:view_sentence_range[1]]))
```
## Implement Preprocessing Function
### Text to Word Ids
As you did with other RNNs, you must turn the text into a number so the computer can understand it. In the function `text_to_ids()`, you'll turn `source_text` and `target_text` from words to ids. However, you need to add the `<EOS>` word id at the end of `target_text`. This will help the neural network predict when the sentence should end.
You can get the `<EOS>` word id by doing:
```python
target_vocab_to_int['<EOS>']
```
You can get other word ids using `source_vocab_to_int` and `target_vocab_to_int`.
```
def text_to_ids(source_text, target_text, source_vocab_to_int, target_vocab_to_int):
"""
Convert source and target text to proper word ids
:param source_text: String that contains all the source text.
:param target_text: String that contains all the target text.
:param source_vocab_to_int: Dictionary to go from the source words to an id
:param target_vocab_to_int: Dictionary to go from the target words to an id
:return: A tuple of lists (source_id_text, target_id_text)
"""
# Split text into sentences
source_text_lst = source_text.split('\n')
target_text_lst = target_text.split('\n')
# Append <EOS> at the end of each sententence
target_text_lst = [sentence + ' <EOS>' for sentence in target_text_lst]
# Make lists using vocab to int mapping
source_id_text = [[source_vocab_to_int[word] for word in sentence.split()] for sentence in source_text_lst]
target_id_text = [[target_vocab_to_int[word] for word in sentence.split()] for sentence in target_text_lst]
return source_id_text, target_id_text
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_text_to_ids(text_to_ids)
```
### Preprocess all the data and save it
Running the code cell below will preprocess all the data and save it to file.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
helper.preprocess_and_save_data(source_path, target_path, text_to_ids)
```
# Check Point
This is your first checkpoint. If you ever decide to come back to this notebook or have to restart the notebook, you can start from here. The preprocessed data has been saved to disk.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
import numpy as np
import helper
(source_int_text, target_int_text), (source_vocab_to_int, target_vocab_to_int), _ = helper.load_preprocess()
```
### Check the Version of TensorFlow and Access to GPU
This will check to make sure you have the correct version of TensorFlow and access to a GPU
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
from distutils.version import LooseVersion
import warnings
import tensorflow as tf
from tensorflow.python.layers.core import Dense
# Check TensorFlow Version
assert LooseVersion(tf.__version__) >= LooseVersion('1.1'), 'Please use TensorFlow version 1.1 or newer'
print('TensorFlow Version: {}'.format(tf.__version__))
# Check for a GPU
if not tf.test.gpu_device_name():
warnings.warn('No GPU found. Please use a GPU to train your neural network.')
else:
print('Default GPU Device: {}'.format(tf.test.gpu_device_name()))
```
## Build the Neural Network
You'll build the components necessary to build a Sequence-to-Sequence model by implementing the following functions below:
- `model_inputs`
- `process_decoder_input`
- `encoding_layer`
- `decoding_layer_train`
- `decoding_layer_infer`
- `decoding_layer`
- `seq2seq_model`
### Input
Implement the `model_inputs()` function to create TF Placeholders for the Neural Network. It should create the following placeholders:
- Input text placeholder named "input" using the TF Placeholder name parameter with rank 2.
- Targets placeholder with rank 2.
- Learning rate placeholder with rank 0.
- Keep probability placeholder named "keep_prob" using the TF Placeholder name parameter with rank 0.
- Target sequence length placeholder named "target_sequence_length" with rank 1
- Max target sequence length tensor named "max_target_len" getting its value from applying tf.reduce_max on the target_sequence_length placeholder. Rank 0.
- Source sequence length placeholder named "source_sequence_length" with rank 1
Return the placeholders in the following the tuple (input, targets, learning rate, keep probability, target sequence length, max target sequence length, source sequence length)
```
def model_inputs():
"""
Create TF Placeholders for input, targets, learning rate, and lengths of source and target sequences.
:return: Tuple (input, targets, learning rate, keep probability, target sequence length,
max target sequence length, source sequence length)
"""
inputs = tf.placeholder(tf.int32, shape=(None, None), name='input')
targets = tf.placeholder(tf.int32, shape=(None, None))
learn_rate = tf.placeholder(tf.float32, shape=None)
keep_prob = tf.placeholder(tf.float32, shape=None, name='keep_prob')
target_sequence_length = tf.placeholder(tf.int32, (None,), name='target_sequence_length')
max_target_sequence_length = tf.reduce_max(target_sequence_length, name='max_target_len')
source_sequence_length = tf.placeholder(tf.int32, (None,), name='source_sequence_length')
return inputs, targets, learn_rate, keep_prob, target_sequence_length, max_target_sequence_length, source_sequence_length
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_model_inputs(model_inputs)
```
### Process Decoder Input
Implement `process_decoder_input` by removing the last word id from each batch in `target_data` and concat the GO ID to the begining of each batch.
```
def process_decoder_input(target_data, target_vocab_to_int, batch_size):
"""
Preprocess target data for encoding
:param target_data: Target Placehoder
:param target_vocab_to_int: Dictionary to go from the target words to an id
:param batch_size: Batch Size
:return: Preprocessed target data
"""
ending = tf.strided_slice(target_data, [0, 0], [batch_size, -1], [1, 1])
dec_input = tf.concat([tf.fill([batch_size, 1], target_vocab_to_int['<GO>']), ending], 1)
return dec_input
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_process_encoding_input(process_decoder_input)
```
### Encoding
Implement `encoding_layer()` to create a Encoder RNN layer:
* Embed the encoder input using [`tf.contrib.layers.embed_sequence`](https://www.tensorflow.org/api_docs/python/tf/contrib/layers/embed_sequence)
* Construct a [stacked](https://github.com/tensorflow/tensorflow/blob/6947f65a374ebf29e74bb71e36fd82760056d82c/tensorflow/docs_src/tutorials/recurrent.md#stacking-multiple-lstms) [`tf.contrib.rnn.LSTMCell`](https://www.tensorflow.org/api_docs/python/tf/contrib/rnn/LSTMCell) wrapped in a [`tf.contrib.rnn.DropoutWrapper`](https://www.tensorflow.org/api_docs/python/tf/contrib/rnn/DropoutWrapper)
* Pass cell and embedded input to [`tf.nn.dynamic_rnn()`](https://www.tensorflow.org/api_docs/python/tf/nn/dynamic_rnn)
```
from imp import reload
reload(tests)
def encoding_layer(rnn_inputs, rnn_size, num_layers, keep_prob,
source_sequence_length, source_vocab_size,
encoding_embedding_size):
"""
Create encoding layer
:param rnn_inputs: Inputs for the RNN
:param rnn_size: RNN Size
:param num_layers: Number of layers
:param keep_prob: Dropout keep probability
:param source_sequence_length: a list of the lengths of each sequence in the batch
:param source_vocab_size: vocabulary size of source data
:param encoding_embedding_size: embedding size of source data
:return: tuple (RNN output, RNN state)
"""
# Encoder embedding
enc_embed_input = tf.contrib.layers.embed_sequence(rnn_inputs, source_vocab_size, encoding_embedding_size)
# RNN cell
def make_cell(rnn_size):
enc_cell = tf.contrib.rnn.LSTMCell(rnn_size,
initializer=tf.random_uniform_initializer(-0.1, 0.1, seed=2))
return enc_cell
enc_cell = tf.contrib.rnn.MultiRNNCell([make_cell(rnn_size) for _ in range(num_layers)])
drop = tf.contrib.rnn.DropoutWrapper(enc_cell, output_keep_prob=keep_prob)
enc_output, enc_state = tf.nn.dynamic_rnn(drop, enc_embed_input, sequence_length=source_sequence_length, dtype=tf.float32)
return enc_output, enc_state
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_encoding_layer(encoding_layer)
```
### Decoding - Training
Create a training decoding layer:
* Create a [`tf.contrib.seq2seq.TrainingHelper`](https://www.tensorflow.org/api_docs/python/tf/contrib/seq2seq/TrainingHelper)
* Create a [`tf.contrib.seq2seq.BasicDecoder`](https://www.tensorflow.org/api_docs/python/tf/contrib/seq2seq/BasicDecoder)
* Obtain the decoder outputs from [`tf.contrib.seq2seq.dynamic_decode`](https://www.tensorflow.org/api_docs/python/tf/contrib/seq2seq/dynamic_decode)
```
def decoding_layer_train(encoder_state, dec_cell, dec_embed_input,
target_sequence_length, max_summary_length,
output_layer, keep_prob):
"""
Create a decoding layer for training
:param encoder_state: Encoder State
:param dec_cell: Decoder RNN Cell
:param dec_embed_input: Decoder embedded input
:param target_sequence_length: The lengths of each sequence in the target batch
:param max_summary_length: The length of the longest sequence in the batch
:param output_layer: Function to apply the output layer
:param keep_prob: Dropout keep probability
:return: BasicDecoderOutput containing training logits and sample_id
"""
# Helper for the training process. Used by BasicDecoder to read inputs.
training_helper = tf.contrib.seq2seq.TrainingHelper(inputs=dec_embed_input,
sequence_length=target_sequence_length,
time_major=False)
# Basic decoder
training_decoder = tf.contrib.seq2seq.BasicDecoder(dec_cell,
training_helper,
encoder_state,
output_layer)
# Perform dynamic decoding using the decoder
training_decoder_output, _ = tf.contrib.seq2seq.dynamic_decode(training_decoder,
impute_finished=True,
maximum_iterations=max_summary_length)
return training_decoder_output
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_decoding_layer_train(decoding_layer_train)
```
### Decoding - Inference
Create inference decoder:
* Create a [`tf.contrib.seq2seq.GreedyEmbeddingHelper`](https://www.tensorflow.org/api_docs/python/tf/contrib/seq2seq/GreedyEmbeddingHelper)
* Create a [`tf.contrib.seq2seq.BasicDecoder`](https://www.tensorflow.org/api_docs/python/tf/contrib/seq2seq/BasicDecoder)
* Obtain the decoder outputs from [`tf.contrib.seq2seq.dynamic_decode`](https://www.tensorflow.org/api_docs/python/tf/contrib/seq2seq/dynamic_decode)
```
def decoding_layer_infer(encoder_state, dec_cell, dec_embeddings, start_of_sequence_id,
end_of_sequence_id, max_target_sequence_length,
vocab_size, output_layer, batch_size, keep_prob):
"""
Create a decoding layer for inference
:param encoder_state: Encoder state
:param dec_cell: Decoder RNN Cell
:param dec_embeddings: Decoder embeddings
:param start_of_sequence_id: GO ID
:param end_of_sequence_id: EOS Id
:param max_target_sequence_length: Maximum length of target sequences
:param vocab_size: Size of decoder/target vocabulary
:param decoding_scope: TenorFlow Variable Scope for decoding
:param output_layer: Function to apply the output layer
:param batch_size: Batch size
:param keep_prob: Dropout keep probability
:return: BasicDecoderOutput containing inference logits and sample_id
"""
# Reuses the same parameters trained by the training process
start_tokens = tf.tile(tf.constant([start_of_sequence_id], dtype=tf.int32), [batch_size], name='start_tokens')
# Helper for the inference process.
inference_helper = tf.contrib.seq2seq.GreedyEmbeddingHelper(dec_embeddings,
start_tokens,
end_of_sequence_id)
# Basic decoder
inference_decoder = tf.contrib.seq2seq.BasicDecoder(dec_cell,
inference_helper,
encoder_state,
output_layer)
# Perform dynamic decoding using the decoder
inference_decoder_output, _ = tf.contrib.seq2seq.dynamic_decode(inference_decoder,
impute_finished=True,
maximum_iterations=max_target_sequence_length)
return inference_decoder_output
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_decoding_layer_infer(decoding_layer_infer)
```
### Build the Decoding Layer
Implement `decoding_layer()` to create a Decoder RNN layer.
* Embed the target sequences
* Construct the decoder LSTM cell (just like you constructed the encoder cell above)
* Create an output layer to map the outputs of the decoder to the elements of our vocabulary
* Use the your `decoding_layer_train(encoder_state, dec_cell, dec_embed_input, target_sequence_length, max_target_sequence_length, output_layer, keep_prob)` function to get the training logits.
* Use your `decoding_layer_infer(encoder_state, dec_cell, dec_embeddings, start_of_sequence_id, end_of_sequence_id, max_target_sequence_length, vocab_size, output_layer, batch_size, keep_prob)` function to get the inference logits.
Note: You'll need to use [tf.variable_scope](https://www.tensorflow.org/api_docs/python/tf/variable_scope) to share variables between training and inference.
```
def decoding_layer(dec_input, encoder_state,
target_sequence_length, max_target_sequence_length,
rnn_size,
num_layers, target_vocab_to_int, target_vocab_size,
batch_size, keep_prob, decoding_embedding_size):
"""
Create decoding layer
:param dec_input: Decoder input
:param encoder_state: Encoder state
:param target_sequence_length: The lengths of each sequence in the target batch
:param max_target_sequence_length: Maximum length of target sequences
:param rnn_size: RNN Size
:param num_layers: Number of layers
:param target_vocab_to_int: Dictionary to go from the target words to an id
:param target_vocab_size: Size of target vocabulary
:param batch_size: The size of the batch
:param keep_prob: Dropout keep probability
:param decoding_embedding_size: Decoding embedding size
:return: Tuple of (Training BasicDecoderOutput, Inference BasicDecoderOutput)
"""
# Embed the target sequences
dec_embeddings = tf.Variable(tf.random_uniform([target_vocab_size, decoding_embedding_size]))
dec_embed_inputs = tf.nn.embedding_lookup(dec_embeddings, dec_input)
# Construct the decoder LSTM cell (just like you constructed the encoder cell above)
def make_cell(rnn_size):
dec_cell = tf.contrib.rnn.LSTMCell(rnn_size, initializer=tf.random_uniform_initializer(-0.1, 0.1, seed=2))
return dec_cell
dec_cell = tf.contrib.rnn.MultiRNNCell([make_cell(rnn_size) for _ in range(num_layers)])
# Create an output layer to map the outputs of the decoder to the elements of our vocabulary
output_layer = Dense(target_vocab_size, kernel_initializer = tf.truncated_normal_initializer(mean = 0.0, stddev=0.1))
with tf.variable_scope("decode"):
# Helper for the training process. Used by BasicDecoder to read inputs.
training_helper = tf.contrib.seq2seq.TrainingHelper(inputs=dec_embed_inputs,
sequence_length=target_sequence_length,
time_major=False)
# Basic decoder
training_decoder = tf.contrib.seq2seq.BasicDecoder(dec_cell,
training_helper,
encoder_state,
output_layer)
# Perform dynamic decoding using the decoder
training_decoder_output, _ = tf.contrib.seq2seq.dynamic_decode(training_decoder,
impute_finished=True,
maximum_iterations=max_target_sequence_length)
# 5. Inference Decoder
# Reuses the same parameters trained by the training process
with tf.variable_scope("decode", reuse=True):
start_tokens = tf.tile(tf.constant([target_vocab_to_int['<GO>']], dtype=tf.int32), [batch_size], name='start_tokens')
# Helper for the inference process.
inference_helper = tf.contrib.seq2seq.GreedyEmbeddingHelper(dec_embeddings,
start_tokens,
target_vocab_to_int['<EOS>'])
# Basic decoder
inference_decoder = tf.contrib.seq2seq.BasicDecoder(dec_cell,
inference_helper,
encoder_state,
output_layer)
# Perform dynamic decoding using the decoder
inference_decoder_output, _ = tf.contrib.seq2seq.dynamic_decode(inference_decoder,
impute_finished=True,
maximum_iterations=max_target_sequence_length)
return training_decoder_output, inference_decoder_output
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_decoding_layer(decoding_layer)
```
### Build the Neural Network
Apply the functions you implemented above to:
- Encode the input using your `encoding_layer(rnn_inputs, rnn_size, num_layers, keep_prob, source_sequence_length, source_vocab_size, encoding_embedding_size)`.
- Process target data using your `process_decoder_input(target_data, target_vocab_to_int, batch_size)` function.
- Decode the encoded input using your `decoding_layer(dec_input, enc_state, target_sequence_length, max_target_sentence_length, rnn_size, num_layers, target_vocab_to_int, target_vocab_size, batch_size, keep_prob, dec_embedding_size)` function.
```
def seq2seq_model(input_data, target_data, keep_prob, batch_size,
source_sequence_length, target_sequence_length,
max_target_sentence_length,
source_vocab_size, target_vocab_size,
enc_embedding_size, dec_embedding_size,
rnn_size, num_layers, target_vocab_to_int):
"""
Build the Sequence-to-Sequence part of the neural network
:param input_data: Input placeholder
:param target_data: Target placeholder
:param keep_prob: Dropout keep probability placeholder
:param batch_size: Batch Size
:param source_sequence_length: Sequence Lengths of source sequences in the batch
:param target_sequence_length: Sequence Lengths of target sequences in the batch
:param source_vocab_size: Source vocabulary size
:param target_vocab_size: Target vocabulary size
:param enc_embedding_size: Decoder embedding size
:param dec_embedding_size: Encoder embedding size
:param rnn_size: RNN Size
:param num_layers: Number of layers
:param target_vocab_to_int: Dictionary to go from the target words to an id
:return: Tuple of (Training BasicDecoderOutput, Inference BasicDecoderOutput)
"""
# Pass the input data through the encoder. We'll ignore the encoder output, but use the state
_, enc_state = encoding_layer(input_data,
rnn_size,
num_layers,
keep_prob,
source_sequence_length,
source_vocab_size,
enc_embedding_size)
# Prepare the target sequences we'll feed to the decoder in training mode
dec_input = process_decoder_input(target_data, target_vocab_to_int, batch_size)
# Pass encoder state and decoder inputs to the decoders
training_decoder_output, inference_decoder_output = decoding_layer(dec_input,
enc_state,
target_sequence_length,
max_target_sentence_length,
rnn_size,
num_layers,
target_vocab_to_int,
target_vocab_size,
batch_size,
keep_prob,
dec_embedding_size)
return training_decoder_output, inference_decoder_output
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_seq2seq_model(seq2seq_model)
```
## Neural Network Training
### Hyperparameters
Tune the following parameters:
- Set `epochs` to the number of epochs.
- Set `batch_size` to the batch size.
- Set `rnn_size` to the size of the RNNs.
- Set `num_layers` to the number of layers.
- Set `encoding_embedding_size` to the size of the embedding for the encoder.
- Set `decoding_embedding_size` to the size of the embedding for the decoder.
- Set `learning_rate` to the learning rate.
- Set `keep_probability` to the Dropout keep probability
- Set `display_step` to state how many steps between each debug output statement
```
# Number of Epochs
epochs = 30
# Batch Size
batch_size = 128
# RNN Size
rnn_size = 50
# Number of Layers
num_layers = 2
# Embedding Size
encoding_embedding_size = 13
decoding_embedding_size = 13
# Learning Rate
learning_rate = 0.001
# Dropout Keep Probability
keep_probability = 0.8
display_step = 10
```
### Build the Graph
Build the graph using the neural network you implemented.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
save_path = 'checkpoints/dev'
(source_int_text, target_int_text), (source_vocab_to_int, target_vocab_to_int), _ = helper.load_preprocess()
max_target_sentence_length = max([len(sentence) for sentence in source_int_text])
train_graph = tf.Graph()
with train_graph.as_default():
input_data, targets, lr, keep_prob, target_sequence_length, max_target_sequence_length, source_sequence_length = model_inputs()
#sequence_length = tf.placeholder_with_default(max_target_sentence_length, None, name='sequence_length')
input_shape = tf.shape(input_data)
train_logits, inference_logits = seq2seq_model(tf.reverse(input_data, [-1]),
targets,
keep_prob,
batch_size,
source_sequence_length,
target_sequence_length,
max_target_sequence_length,
len(source_vocab_to_int),
len(target_vocab_to_int),
encoding_embedding_size,
decoding_embedding_size,
rnn_size,
num_layers,
target_vocab_to_int)
training_logits = tf.identity(train_logits.rnn_output, name='logits')
inference_logits = tf.identity(inference_logits.sample_id, name='predictions')
masks = tf.sequence_mask(target_sequence_length, max_target_sequence_length, dtype=tf.float32, name='masks')
with tf.name_scope("optimization"):
# Loss function
cost = tf.contrib.seq2seq.sequence_loss(
training_logits,
targets,
masks)
# Optimizer
optimizer = tf.train.AdamOptimizer(lr)
# Gradient Clipping
gradients = optimizer.compute_gradients(cost)
capped_gradients = [(tf.clip_by_value(grad, -1., 1.), var) for grad, var in gradients if grad is not None]
train_op = optimizer.apply_gradients(capped_gradients)
```
Batch and pad the source and target sequences
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
def pad_sentence_batch(sentence_batch, pad_int):
"""Pad sentences with <PAD> so that each sentence of a batch has the same length"""
max_sentence = max([len(sentence) for sentence in sentence_batch])
return [sentence + [pad_int] * (max_sentence - len(sentence)) for sentence in sentence_batch]
def get_batches(sources, targets, batch_size, source_pad_int, target_pad_int):
"""Batch targets, sources, and the lengths of their sentences together"""
for batch_i in range(0, len(sources)//batch_size):
start_i = batch_i * batch_size
# Slice the right amount for the batch
sources_batch = sources[start_i:start_i + batch_size]
targets_batch = targets[start_i:start_i + batch_size]
# Pad
pad_sources_batch = np.array(pad_sentence_batch(sources_batch, source_pad_int))
pad_targets_batch = np.array(pad_sentence_batch(targets_batch, target_pad_int))
# Need the lengths for the _lengths parameters
pad_targets_lengths = []
for target in pad_targets_batch:
pad_targets_lengths.append(len(target))
pad_source_lengths = []
for source in pad_sources_batch:
pad_source_lengths.append(len(source))
yield pad_sources_batch, pad_targets_batch, pad_source_lengths, pad_targets_lengths
```
### Train
Train the neural network on the preprocessed data. If you have a hard time getting a good loss, check the forms to see if anyone is having the same problem.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
def get_accuracy(target, logits):
"""
Calculate accuracy
"""
max_seq = max(target.shape[1], logits.shape[1])
if max_seq - target.shape[1]:
target = np.pad(
target,
[(0,0),(0,max_seq - target.shape[1])],
'constant')
if max_seq - logits.shape[1]:
logits = np.pad(
logits,
[(0,0),(0,max_seq - logits.shape[1])],
'constant')
return np.mean(np.equal(target, logits))
# Split data to training and validation sets
train_source = source_int_text[batch_size:]
train_target = target_int_text[batch_size:]
valid_source = source_int_text[:batch_size]
valid_target = target_int_text[:batch_size]
(valid_sources_batch, valid_targets_batch, valid_sources_lengths, valid_targets_lengths ) = next(get_batches(valid_source,
valid_target,
batch_size,
source_vocab_to_int['<PAD>'],
target_vocab_to_int['<PAD>']))
with tf.Session(graph=train_graph) as sess:
sess.run(tf.global_variables_initializer())
for epoch_i in range(epochs):
for batch_i, (source_batch, target_batch, sources_lengths, targets_lengths) in enumerate(
get_batches(train_source, train_target, batch_size,
source_vocab_to_int['<PAD>'],
target_vocab_to_int['<PAD>'])):
_, loss = sess.run(
[train_op, cost],
{input_data: source_batch,
targets: target_batch,
lr: learning_rate,
target_sequence_length: targets_lengths,
source_sequence_length: sources_lengths,
keep_prob: keep_probability})
if batch_i % display_step == 0 and batch_i > 0:
batch_train_logits = sess.run(
inference_logits,
{input_data: source_batch,
source_sequence_length: sources_lengths,
target_sequence_length: targets_lengths,
keep_prob: 1.0})
batch_valid_logits = sess.run(
inference_logits,
{input_data: valid_sources_batch,
source_sequence_length: valid_sources_lengths,
target_sequence_length: valid_targets_lengths,
keep_prob: 1.0})
train_acc = get_accuracy(target_batch, batch_train_logits)
valid_acc = get_accuracy(valid_targets_batch, batch_valid_logits)
print('Epoch {:>3} Batch {:>4}/{} - Train Accuracy: {:>6.4f}, Validation Accuracy: {:>6.4f}, Loss: {:>6.4f}'
.format(epoch_i, batch_i, len(source_int_text) // batch_size, train_acc, valid_acc, loss))
# Save Model
saver = tf.train.Saver()
saver.save(sess, save_path)
print('Model Trained and Saved')
```
### Save Parameters
Save the `batch_size` and `save_path` parameters for inference.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
# Save parameters for checkpoint
helper.save_params(save_path)
```
# Checkpoint
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
import tensorflow as tf
import numpy as np
import helper
import problem_unittests as tests
_, (source_vocab_to_int, target_vocab_to_int), (source_int_to_vocab, target_int_to_vocab) = helper.load_preprocess()
load_path = helper.load_params()
```
## Sentence to Sequence
To feed a sentence into the model for translation, you first need to preprocess it. Implement the function `sentence_to_seq()` to preprocess new sentences.
- Convert the sentence to lowercase
- Convert words into ids using `vocab_to_int`
- Convert words not in the vocabulary, to the `<UNK>` word id.
```
def sentence_to_seq(sentence, vocab_to_int):
"""
Convert a sentence to a sequence of ids
:param sentence: String
:param vocab_to_int: Dictionary to go from the words to an id
:return: List of word ids
"""
# TODO: Implement Function
return None
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_sentence_to_seq(sentence_to_seq)
```
## Translate
This will translate `translate_sentence` from English to French.
```
translate_sentence = 'he saw a old yellow truck .'
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
translate_sentence = sentence_to_seq(translate_sentence, source_vocab_to_int)
loaded_graph = tf.Graph()
with tf.Session(graph=loaded_graph) as sess:
# Load saved model
loader = tf.train.import_meta_graph(load_path + '.meta')
loader.restore(sess, load_path)
input_data = loaded_graph.get_tensor_by_name('input:0')
logits = loaded_graph.get_tensor_by_name('predictions:0')
target_sequence_length = loaded_graph.get_tensor_by_name('target_sequence_length:0')
source_sequence_length = loaded_graph.get_tensor_by_name('source_sequence_length:0')
keep_prob = loaded_graph.get_tensor_by_name('keep_prob:0')
translate_logits = sess.run(logits, {input_data: [translate_sentence]*batch_size,
target_sequence_length: [len(translate_sentence)*2]*batch_size,
source_sequence_length: [len(translate_sentence)]*batch_size,
keep_prob: 1.0})[0]
print('Input')
print(' Word Ids: {}'.format([i for i in translate_sentence]))
print(' English Words: {}'.format([source_int_to_vocab[i] for i in translate_sentence]))
print('\nPrediction')
print(' Word Ids: {}'.format([i for i in translate_logits]))
print(' French Words: {}'.format(" ".join([target_int_to_vocab[i] for i in translate_logits])))
```
## Imperfect Translation
You might notice that some sentences translate better than others. Since the dataset you're using only has a vocabulary of 227 English words of the thousands that you use, you're only going to see good results using these words. For this project, you don't need a perfect translation. However, if you want to create a better translation model, you'll need better data.
You can train on the [WMT10 French-English corpus](http://www.statmt.org/wmt10/training-giga-fren.tar). This dataset has more vocabulary and richer in topics discussed. However, this will take you days to train, so make sure you've a GPU and the neural network is performing well on dataset we provided. Just make sure you play with the WMT10 corpus after you've submitted this project.
## Submitting This Project
When submitting this project, make sure to run all the cells before saving the notebook. Save the notebook file as "dlnd_language_translation.ipynb" and save it as a HTML file under "File" -> "Download as". Include the "helper.py" and "problem_unittests.py" files in your submission.
| github_jupyter |
# Linear Support Vector Classification with StandardScaler
This Code template is for the Classification task using a simple Linear Support Vector Classifier(LinearSVC) based on the Support Vector Machine algorithm and feature rescaling technique StandardScaler in a pipeline.
### Required Packages
```
!pip install imblearn
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as se
import warnings
from sklearn.model_selection import train_test_split
from imblearn.over_sampling import RandomOverSampler
from sklearn.svm import LinearSVC
from sklearn.preprocessing import LabelEncoder,StandardScaler
from sklearn.pipeline import make_pipeline
from sklearn.metrics import classification_report,plot_confusion_matrix
warnings.filterwarnings('ignore')
```
### Initialization
Filepath of CSV file
```
#filepath
file_path= ""
```
List of features which are required for model training .
```
#x_values
features=[]
```
Target variable for prediction.
```
#y_value
target=""
```
### Data Fetching
Pandas is an open-source, BSD-licensed library providing high-performance, easy-to-use data manipulation and data analysis tools.
We will use panda's library to read the CSV file using its storage path.And we use the head function to display the initial row or entry.
```
df=pd.read_csv(file_path)
df.head()
```
### Feature Selections
It is the process of reducing the number of input variables when developing a predictive model. Used to reduce the number of input variables to both reduce the computational cost of modelling and, in some cases, to improve the performance of the model.
We will assign all the required input features to X and target/outcome to Y.
```
X=df[features]
Y=df[target]
```
### Data Preprocessing
Since the majority of the machine learning models in the Sklearn library doesn't handle string category data and Null value, we have to explicitly remove or replace null values. The below snippet have functions, which removes the null value if any exists. And convert the string classes data in the datasets by encoding them to integer classes.
```
def NullClearner(df):
if(isinstance(df, pd.Series) and (df.dtype in ["float64","int64"])):
df.fillna(df.mean(),inplace=True)
return df
elif(isinstance(df, pd.Series)):
df.fillna(df.mode()[0],inplace=True)
return df
else:return df
def EncodeX(df):
return pd.get_dummies(df)
def EncodeY(df):
if len(df.unique())<=2:
return df
else:
un_EncodedT=np.sort(pd.unique(df), axis=-1, kind='mergesort')
df=LabelEncoder().fit_transform(df)
EncodedT=[xi for xi in range(len(un_EncodedT))]
print("Encoded Target: {} to {}".format(un_EncodedT,EncodedT))
return df
```
Calling preprocessing functions on the feature and target set.
```
x=X.columns.to_list()
for i in x:
X[i]=NullClearner(X[i])
X=EncodeX(X)
Y=EncodeY(NullClearner(Y))
X.head()
```
#### Correlation Map
In order to check the correlation between the features, we will plot a correlation matrix. It is effective in summarizing a large amount of data where the goal is to see patterns.
```
f,ax = plt.subplots(figsize=(18, 18))
matrix = np.triu(X.corr())
se.heatmap(X.corr(), annot=True, linewidths=.5, fmt= '.1f',ax=ax, mask=matrix)
plt.show()
```
#### Distribution Of Target Variable
```
plt.figure(figsize = (10,6))
se.countplot(Y)
```
### Data Splitting
The train-test split is a procedure for evaluating the performance of an algorithm. The procedure involves taking a dataset and dividing it into two subsets. The first subset is utilized to fit/train the model. The second subset is used for prediction. The main motive is to estimate the performance of the model on new data.
```
x_train,x_test,y_train,y_test=train_test_split(X,Y,test_size=0.2,random_state=123)
```
#### Handling Target Imbalance
The challenge of working with imbalanced datasets is that most machine learning techniques will ignore, and in turn have poor performance on, the minority class, although typically it is performance on the minority class that is most important.
One approach to addressing imbalanced datasets is to oversample the minority class. The simplest approach involves duplicating examples in the minority class.We will perform overspampling using imblearn library.
```
x_train,y_train = RandomOverSampler(random_state=123).fit_resample(x_train, y_train)
```
### Model
Support vector machines (SVMs) are a set of supervised learning methods used for classification, regression and outliers detection.
A Support Vector Machine is a discriminative classifier formally defined by a separating hyperplane. In other terms, for a given known/labelled data points, the SVM outputs an appropriate hyperplane that classifies the inputted new cases based on the hyperplane. In 2-Dimensional space, this hyperplane is a line separating a plane into two segments where each class or group occupied on either side.
LinearSVC is similar to SVC with kernel=’linear’. It has more flexibility in the choice of tuning parameters and is suited for large samples.
* #### Model Tuning Parameters
> * penalty -> Specifies the norm used in the penalization. The ‘l2’ penalty is the standard used in SVC. The ‘l1’ leads to coef_ vectors that are sparse.
> * Loss -> Specifies the loss function. ‘hinge’ is the standard SVM loss (used e.g. by the SVC class) while ‘squared_hinge’ is the square of the hinge loss. The combination of penalty='l1' and loss='hinge' is not supported.
> * C -> Regularization parameter. The strength of the regularization is inversely proportional to C. Must be strictly positive.
> * tolerance -> Tolerance for stopping criteria.
> * dual -> Select the algorithm to either solve the dual or primal optimization problem. Prefer dual=False when n_samples > n_features.
```
model=make_pipeline(StandardScaler(),LinearSVC(random_state=123))
model.fit(x_train,y_train)
```
#### Model Accuracy
score() method return the mean accuracy on the given test data and labels.
In multi-label classification, this is the subset accuracy which is a harsh metric since you require for each sample that each label set be correctly predicted.
```
print("Accuracy score {:.2f} %\n".format(model.score(x_test,y_test)*100))
```
#### Confusion Matrix
A confusion matrix is utilized to understand the performance of the classification model or algorithm in machine learning for a given test set where results are known.
```
plot_confusion_matrix(model,x_test,y_test,cmap=plt.cm.Blues)
```
#### Classification Report
A Classification report is used to measure the quality of predictions from a classification algorithm. How many predictions are True, how many are False.
* where:
- Precision:- Accuracy of positive predictions.
- Recall:- Fraction of positives that were correctly identified.
- f1-score:- percent of positive predictions were correct
- support:- Support is the number of actual occurrences of the class in the specified dataset.
```
print(classification_report(y_test,model.predict(x_test)))
```
#### Creator: Thilakraj Devadiga , Github: [Profile](https://github.com/Thilakraj1998)
| github_jupyter |
<a href="https://colab.research.google.com/github/yohanesnuwara/machine-learning/blob/master/06_simple_linear_regression/simple_linear_reg_algorithm.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# **Simple Linear Regression**
```
import numpy as np
import matplotlib.pyplot as plt
```
## Method 1 ("Traditional")
Calculate bias (or intercept $B_0$) and slope ($B_1$) using:
$$B_1 = \frac{\sum_{i=1}^{n}(x_i-mean(x))(y_i-mean(y))}{\sum_{i=1}^{n}(x_i-mean(x))^2}$$
$$B_0 = mean(y) - B_1 \cdot mean(x)$$
to construct simple linear regression model: $$y = B_0 + B_1 \cdot x$$
```
x = [1, 2, 4, 3, 5]
y = [1, 3, 3, 2, 5]
# visualize our data
plt.plot(x, y, 'o')
```
Calculate mean of data
```
mean_x = np.mean(x)
mean_y = np.mean(y)
print(mean_x, mean_y)
```
Calculate error
```
err_x = x - mean_x
err_y = y - mean_y
print(err_x)
print(err_y)
```
Multiply error of x and error of y
```
err_mult = err_x * err_y
print(err_mult)
```
Calculate numerator by summing up the errors
```
numerator = np.sum(err_mult)
numerator
```
Calculate denominator by squaring the x error and summing them up
```
err_x_squared = err_x**2
denominator = np.sum(err_x_squared)
print(denominator)
```
Calculate the **slope (B1)** !
```
B1 = numerator / denominator
print(B1)
```
And we can calculate the **intercept (c)** !
```
B0 = mean_y - B1 * mean_x
print(B0)
```
We now have the coefficents for our simple linear regression equation.
$$y = B_0 + B_1 x = 0.4 + 0.8 x$$
### Test the model to our training data
```
x_test = np.array([1, 2, 3, 4, 5])
y_predicted = B0 + B1 * x_test
p1 = plt.plot(x, y, 'o')
p2 = plt.plot(x_test, y_predicted, 'o-', color='r')
plt.legend((p1[0], p2[0]), (['y data', 'predicted y']))
```
### Estimating Error (Root Mean Squared Error)
$$RMSE = \sqrt{\frac{\sum_{i=1}^{n} (p_i - y_i)^2}{n}}$$
```
numerator = np.sum((y_predicted - y)**2)
denominator = len(y)
rmse = np.sqrt(numerator / denominator)
rmse
```
### Wrap all up
```
def simple_linear_regression_traditional(x, y, x_test):
import numpy as np
x = np.array(x); y = np.array(y); x_test = np.array(x_test)
mean_x = np.mean(x)
mean_y = np.mean(y)
err_x = x - mean_x
err_y = y - mean_y
err_mult = err_x * err_y
numerator = np.sum(err_mult)
err_x_squared = err_x**2
denominator = np.sum(err_x_squared)
B1 = numerator / denominator
B0 = mean_y - B1 * mean_x
y_predicted = B0 + B1 * x_test
return(B0, B1, y_predicted)
def linreg_error(y, y_predicted):
import numpy as np
y = np.array(y); y_predicted = np.array(y_predicted)
numerator = np.sum((y_predicted - y)**2)
denominator = len(y)
rmse = np.sqrt(numerator / denominator)
return(rmse)
```
## Method 2 ("Advanced")
Calculate bias (or intercept $B_0$) and slope ($B_1$) using:
$$B_1 = corr(x, y) \cdot \frac{stdev(y)}{stdev(x)}$$
Then, similar to **Method 1**.
$$B_0 = mean(y) - B_1 \cdot mean(x)$$
to construct simple linear regression model: $$y = B_0 + B_1 \cdot x$$
Calculate the **pearson's correlation coefficient $corr(x,y)$**. First, calculate mean and standard deviation.
```
import statistics as stat
mean_x = np.mean(x)
mean_y = np.mean(y)
stdev_x = stat.stdev(x)
stdev_y = stat.stdev(y)
print(stdev_x, stdev_y)
```
Calculate **covariance**. Covariance is the relationship that can be summarized between two variables. The sign of the covariance can be interpreted as whether the two variables change in the same direction (positive) or change in different directions (negative). A covariance value of zero indicates that both variables are completely independent.
```
cov_x_y = (np.sum((x - mean_x) * (y - mean_y))) * (1 / (len(x) - 1))
cov_x_y
```
Calculate **Pearson's Correlation Coefficient**. It summarizes the strength of the linear relationship between two data samples. It is the normalization of the covariance between the two variables. The coefficient returns a value between -1 and 1 that represents the limits of correlation from a full negative correlation to a full positive correlation. A value of 0 means no correlation. The value must be interpreted, where often a value below -0.5 or above 0.5 indicates a notable correlation, and values below those values suggests a less notable correlation.
```
corr_x_y = cov_x_y / (stdev_x * stdev_y)
corr_x_y
```
Calculate slope $B_1$
```
B1 = corr_x_y * (stdev_y / stdev_x)
B1
```
Next, is similar to **Method 1**.
```
B0 = mean_y - B1 * mean_x
x_test = np.array([1, 2, 3, 4, 5])
y_predicted = B0 + B1 * x_test
p1 = plt.plot(x, y, 'o')
p2 = plt.plot(x_test, y_predicted, 'o-', color='r')
plt.legend((p1[0], p2[0]), (['y data', 'predicted y']))
```
Calculate RMSE
```
rmse = linreg_error(y, y_predicted)
rmse
```
### Wrap all up
```
def simple_linear_regression_advanced(x, y, x_test):
import numpy as np
import statistics as stat
x = np.array(x); y = np.array(y); x_test = np.array(x_test)
mean_x = np.mean(x)
mean_y = np.mean(y)
stdev_x = stat.stdev(x)
stdev_y = stat.stdev(y)
cov_x_y = (np.sum((x - mean_x) * (y - mean_y))) * (1 / (len(x) - 1))
corr_x_y = cov_x_y / (stdev_x * stdev_y)
B1 = corr_x_y * (stdev_y / stdev_x)
B0 = mean_y - B1 * mean_x
y_predicted = B0 + B1 * x_test
return(B0, B1, y_predicted)
```
## Implement to Real Dataset
Simple linear regression to WTI and Brent Daily Oil Price (1980-2020)
```
!git clone https://www.github.com/yohanesnuwara/machine-learning
import pandas as pd
brent = pd.read_csv('/content/machine-learning/datasets/brent-daily_csv.csv')
wti = pd.read_csv('/content/machine-learning/datasets/wti-daily_csv.csv')
# Converting to Panda datetime
brent['Date'] = pd.to_datetime(brent['Date'], format='%Y-%m-%d') # depends on the data, format check web: https://strftime.org/
wti['Date'] = pd.to_datetime(wti['Date'], format='%Y-%m-%d') # depends on the data, format check web: https://strftime.org/
brent.head(10)
```
Visualize data
```
from pandas.plotting import register_matplotlib_converters
register_matplotlib_converters()
plt.figure(figsize=(15, 6))
plt.plot(brent.Date, brent.Price, '.', color='blue')
plt.plot(wti.Date, wti.Price, '.', color='red')
plt.title('Daily Oil Price')
plt.xlabel('Year'); plt.ylabel('Price ($/bbl)')
# convert datetime to ordinal
import datetime as dt
brent_date = np.array(brent['Date'].map(dt.datetime.toordinal))
brent_price = brent.Price
brent_test = brent_date
B0_brent, B1_brent, brent_price_predicted = simple_linear_regression_advanced(brent_date, brent_price, brent_test)
wti_date = np.array(wti['Date'].map(dt.datetime.toordinal))
wti_price = wti.Price
wti_test = wti_date
B0_wti, B1_wti, wti_price_predicted = simple_linear_regression_advanced(wti_date, wti_price, wti_test)
plt.figure(figsize=(15, 6))
p1 = plt.plot(brent.Date, brent.Price, '.', color='blue')
p2 = plt.plot(wti.Date, wti.Price, '.', color='red')
p3 = plt.plot(brent_test, brent_price_predicted, color='blue')
p4 = plt.plot(wti_test, wti_price_predicted, color='red')
plt.legend((p1[0], p2[0], p3[0], p4[0]), (['Brent data', 'WTI data', 'Brent predicted', 'WTI predicted']))
plt.title('Daily Oil Price')
plt.xlabel('Year'); plt.ylabel('Price ($/bbl)')
```
| github_jupyter |
```
import matplotlib.pyplot as plt
%matplotlib inline
import pickle
import numpy as np
from scipy.spatial.distance import pdist, squareform
with open('exp_features.p', 'rb') as f:
data = pickle.load(f)
```
## visualize
```
def get_continuous_quantile(x, y, n_interval=100, q=1):
"""
Take continuous x and y, bin the data according to the intervals of x
and then calculate the quantiles of y within this bin
Args:
x (list): array of x values
y (list): array of y values
n_interval (int): number of intervals on x
q (float): quantile value [0, 1]
"""
ind = np.argsort(x)
x = x[ind]
y = y[ind]
boundaries = np.linspace(x[0], x[-1], n_interval+1)
dx = boundaries[1] - boundaries[0]
x_center = np.linspace(x[0]+dx/2, x[-1]-dx/2, n_interval)
y_q = []
for x_min, x_max in zip(boundaries[:-1], boundaries[1:]):
ind = (x>=x_min) & (x<x_max)
ys = y[ind]
if len(ys) > 0:
y_q.append(np.quantile(ys, q))
else:
y_q.append(y_q[-1])
y_q = np.array(y_q)
return x_center, y_q
def visualize(key, n_interval=100, interval=5, alpha=0.5, data_file="100_0.xlsx"):
"""
Visualize the data specified by key.
Args:
key (str): key in data
n_interval (int): number of intervals for drawing the quantile bounds
interval (int): subsamping of the data. Sometimes the input data is too large for visualization
we just subsample the data
"""
keys = list(data['band_gap'].keys())
f = np.concatenate([data[key][i] for i in keys], axis=0)
values = np.array([data['band_gap'][i] for i in keys])
sort_index = np.argsort(values)
fscale = (f-np.min(f, axis=0)) / (np.max(f, axis=0) - np.min(f, axis=0))
d = pdist(fscale)
v_dist = pdist(values.reshape((-1, 1)))
ind = (d>0) & (d<1)
d_ = d[ind]
v_ = v_dist[ind]
#print(d_.shape, v_.shape)
x_center, y_q = get_continuous_quantile(d_, v_, n_interval=n_interval, q=1)
plt.rcParams['font.size'] = 22
plt.rcParams['font.family'] = 'Arial'
plt.figure(figsize=(5.7, 5.0 ))
d_ = d_[::interval]
v_ = v_[::interval]
print(v_.shape)
plt.plot(d_, v_, 'o', alpha=alpha, c='#21c277')
plt.plot(x_center, y_q, '--', c='#21c277', lw=2, alpha=0.5)
import pandas as pd
x = np.round(np.concatenate([d_, x_center]), 3)
y = np.round(np.concatenate([v_, y_q]), 3)
df = pd.DataFrame({"dF": x, "dEg": y})
with pd.ExcelWriter(data_file) as writer:
df.to_excel(writer)
plt.xlim([0, 1])
plt.ylim([0, 13])
plt.xticks(np.linspace(0, 1, 5))
plt.yticks(np.linspace(0, 12.5, 6))
plt.xlabel('$d_{F}$ (a.u.)')
plt.ylabel("$\Delta E_{g}$ (eV)")
plt.tight_layout()
visualize('100_0', n_interval=100, interval=15, alpha=0.08, data_file='100_0.xlsx')
plt.savefig("100_0.pdf")
visualize('100_41000', n_interval=100, interval=15, alpha=0.08, data_file='100_41000.xlsx')
plt.savefig("100_41000.pdf")
```
| github_jupyter |
```
import sklearn
from sklearn.model_selection import train_test_split
from matplotlib import pyplot as plt
%matplotlib inline
import pandas
import numpy as np
import mglearn
from collections import Counter
from sklearn.metrics import cohen_kappa_score
from sklearn import preprocessing
df = pandas.read_excel('house_price_label.xlsx')
# combine multipl columns into a 2D array
# also convert the integer data to float data
X = np.column_stack((df.built_in.astype(float),df.price.astype(float)))
X = preprocessing.scale(X) # scale the data before training the model
y = df.house_type
X_train, X_test, y_train, y_test = train_test_split(X, y,test_size =0.3,stratify = y, random_state=0)
# for classification, make sure a stratify splitting method is selected
mglearn.discrete_scatter(X[:,0],X[:,1],y) # use mglearn to visualize data
plt.legend(y,loc='best')
plt.xlabel('build_in')
plt.ylabel('house price')
plt.show()
from sklearn.neural_network import MLPClassifier
mlp = MLPClassifier(solver='lbfgs',hidden_layer_sizes=(10,), random_state=0).fit(X_train, y_train)
mglearn.discrete_scatter(X_train[:, 0], X_train[:, 1],mlp.predict(X_train))
plt.legend(y,loc='best')
plt.xlabel('build_in')
plt.ylabel('house price')
plt.show()
print("Training set accuracy: {:.2f}".format(mlp.score(X_train, y_train)))
print ("Training Kappa: {:.3f}".format(cohen_kappa_score(y_train,mlp.predict(X_train))))
print("Test set accuracy: {:.2f}".format(mlp.score(X_test, y_test)))
print ("Test Kappa: {:.3f}".format(cohen_kappa_score(y_test,mlp.predict(X_test))))
from sklearn.neural_network import MLPClassifier
mlp = MLPClassifier(solver='lbfgs', hidden_layer_sizes=(20,20,20), random_state=0).fit(X_train, y_train)
mglearn.discrete_scatter(X_train[:, 0], X_train[:, 1],mlp.predict(X_train))
plt.legend(y,loc='best')
plt.xlabel('build_in')
plt.ylabel('house price')
plt.show()
print("Training set accuracy: {:.2f}".format(mlp.score(X_train, y_train)))
print ("Training Kappa: {:.3f}".format(cohen_kappa_score(y_train,mlp.predict(X_train))))
print("Test set accuracy: {:.2f}".format(mlp.score(X_test, y_test)))
print ("Test Kappa: {:.3f}".format(cohen_kappa_score(y_test,mlp.predict(X_test))))
fig, axes = plt.subplots(2, 4, figsize=(20, 8))
for axx, n_hidden_nodes in zip(axes, [10, 20]):
for ax, alpha in zip(axx, [0.0001, 0.01, 0.1, 1]):
mlp = MLPClassifier(solver='lbfgs', random_state=0,
hidden_layer_sizes=[n_hidden_nodes, n_hidden_nodes],
alpha=alpha)
mlp.fit(X_train, y_train)
mglearn.discrete_scatter(X_train[:, 0], X_train[:, 1], mlp.predict(X_train), ax=ax)
ax.set_title("n_hidden=[{}, {}]\nalpha={:.4f}\nkapa={:.4f}".format(
n_hidden_nodes, n_hidden_nodes, alpha,cohen_kappa_score(y_train,mlp.predict(X_train))))
plt.subplots_adjust(hspace=0.5)
mlp = MLPClassifier(solver='lbfgs', hidden_layer_sizes=(20,20), random_state=0).fit(X_train, y_train)
fig, axes = plt.subplots(1, 3, figsize=(20, 8))
for i , ax in zip(range(3),axes):
img = ax.imshow(mlp.coefs_[i], interpolation='none', cmap='viridis')
ax.set_title(" No.{} layer".format(i))
ax.set_xlabel("Columns in weight matrix")
ax.set_ylabel("Input feature")
fig.colorbar(img, ax = ax)
```
| github_jupyter |
# **Lab III : Nombres et signaux complexes**
-----------------
+ **Cours "Physique du Numérique"** - Portail René Descartes - AMU
Préparé par :
- Jean-Marc Themlin (v. 2021-09), Aix-Marseille Université © Contenus à diffusion restreinte, dans le cadre de ce cours.
------------------
#### La cellule ci-dessous, à exécuter (`Shift-Enter`) en tout premier lieu, contient les appels aux librairies nécessaires pour exécuter la suite du TP, ainsi que la fonction `cmplxdraw` qui vous sera utile pour visualiser des nombres complexes dans le diagramme d'Argand.
```
import math as m
import cmath as c
import numpy as np
import matplotlib.pyplot as plt
def cmplxdraw(z1):
""" Draws a list of complex numbers as vectors in the complex plane
:z1: array. Array of complex numbers [za,zb,zc,...]
:lim: float/int. Limits of the (square) graph
"""
tt=np.arange(0,1,1/2**6)
z = np.exp(1j*2*np.pi*tt)
fig = plt.figure() # initialise la figure
ax=fig.add_subplot(1,1,1)
plt.plot(np.real(z),np.imag(z))
plt.plot(np.real(z1),np.imag(z1),'or')
plt.axis('square')
plt.plot(0,0,'xb')
m1 = max(abs(np.real(z1)))
m2 = max(abs(np.imag(z1)))
m = max(m1,m2)
zoom=1.35
plt.xlim(-zoom*m,zoom*m)
plt.ylim(-zoom*m,zoom*m)
plt.grid('on')
plt.quiver([0, 0], [0, 0], [0, 1], [1, 0], angles='xy', scale_units='xy', scale=1)
plt.quiver([0, 0], [0, 0], [0, m], [m, 0], angles='xy', scale_units='xy', scale=1)
i=-1
for ind in z1:
i=i+1
ax.annotate('z'+str(i+1), (np.real(z1[i])+0.3,np.imag(z1[i])),fontsize=10)
ax.quiver(0, 0, np.real(z1[i]), np.imag(z1[i]), angles='xy', scale_units='xy', scale=1)
plt.title('Plan complexe')
plt.xlabel('Axe des réels x')
plt.ylabel('Axe des imaginaires y')
ax.set_facecolor("pink")
```
## **III.1 Nombres complexes avec Python**
### III.1.A Définition d'une variable contenant un nombre complexe
Il y a deux manières possibles pour définir le nombre complexe sous sa forme cartésienne. On peut également utiliser la forme polaire $z=|z| \ e^{Arg(z)}$, le résultat est affiché sous forme cartésienne.
```
y=complex(1,1)
print(y)
print(y.real,y.imag) # real et imag sont des attributs d'une variable complexe
print(np.real(y),np.imag(y),abs(y))
f"Le module de y vaut {abs(y):.4f}" # This is a formatted string literal
x=1+1j
print(x)
print(x.real,x.imag,abs(y))
## ou bien
print(np.real(x),np.imag(x),abs(y))
print("Conversion de x sous forme polaire :")
print(" module et argument : ",c.polar(x))
z=2*np.exp(1j*np.pi/4)
print(z)
print("Conversion de z sous forme cartésienne :")
print(" Partie réelle : ",z.real," Partie imaginaire",z.imag)
print("Autre manière possible, à partir des coordonnées polaires :")
print(" Partie réelle : ",c.rect(2,np.pi/4))
```
### III.1.B Tracé d'un nombre dans le plan complexe
La fonction `cmplxdraw` définie ci-dessus affiche dans le plan complexe les nombres complexes contenus dans un *array* de *numpy* de type [z1,z2,...].
```
z=[1+1j]
cmplxdraw(z)
z
z = [1+1j,1j,2*np.exp(1j*3*np.pi/4),np.exp(1j*3*np.pi/2)]
cmplxdraw(z)
y=complex(1,1)
y
```
### III.1.C Opérations sur les nombres complexes
```
x=np.sqrt(2)*(1+1j)/2
y=np.sqrt(2)*(1-1j)/2
za=x+y
zb=x-y
zc=y-x
cmplxdraw([x,y,za,zb,zc])
x=3*np.exp(2j*np.pi/3)
y=4*np.exp(-1j*np.pi/6)
z=x-y
cmplxdraw([x,y,z])
print(-1.5-2*np.sqrt(3))
np.sqrt(3)/2+2
argz=m.atan(-2/np.sqrt(2))
print(argz)
print(3*argz)
print(180*argz/np.pi)
8*np.sqrt(6)
4*np.sqrt(2)
z=np.sqrt(2)-2j
# z=2*np.exp(1j*np.pi)
z8=z**3
z9=1/z
z10=np.sqrt(z)
z11=(1-1j)**2
z12=-4+3j
z13=1/z12
print(abs(z),abs(z8),abs(z9),abs(z10))
cmplxdraw([z,z9,z10,z11,z12,z13])
6**(3/2)
6**0.25
np.sin(np.pi/6)
m.atan(-3/4)
np.exp(4j)
5*m.atan(-4/3)
5**5
13**2
```
## **Exponentielles complexes avec Python**
| github_jupyter |
```
print("Hello world!")
a=10
a
b=5
b
#addition demo
sum=a+b
print("the sum of a and b is:",sum)
x=2**3
x
y=5/2
y
y=5//2
y
input("Enter some variable")
a=int(input("enter the first number"))
b=int(input("enter the second number"))
int("The sum of first number and second number is:",a+b)
int("The difference of the first and second number is:",a-b)
int("The product of the first and second numberis:",a*b)
int("The quotient of the first and second number is:",a//b)
new=("String demo")
new
#Slice operator
#slice operator works like this
# string variable[x::y::z]
#[x::y::z]
#[start_index=:end index 1:skip index:2]
new[0:5:2]
new[0:]
new[5:]
new*2
new*5
#repetition operator
new*3
#concatenation operator
new +" Test"
new
#short way a=10
new += " Extra"
listt= [ "Kar",'']
listt
#length method to find the value of n(the number of elements in list)
#len()
len(listt)
students=["Akhil","Akila","John","sonum","khushi"]
students
#append function- add a new element to the existing list
#append9()
students.append("jyoti")
students
#Insert function- add a new eleent at the end of index position given
#insert (index,value)
students.insert(3,"papa")
students
students
#if else in python
age=int(input("enter your age:"))
if age>=18:
print("You are eligible")
else:
print("You are not eligible")
#No switch case in python
#if-elif-lse block
a=200
b=33
if b>a:
print("B is greater than A")
elif a==b :
print("A is equal to B")
else:
print("A is greater than B")
```
### Nested if example
age=18
if age>=18:
print("Allow inside club")
if age>=21:
print("Drinking allowed")
else:
print("Drinking not allowed")
else :
print("Not allowed inside club")
```
# For loop
#range function = for var in range(int,int,int):
#range(value)= start from 0 upto value
#range(value 1,value2)= start from value 1 and go upto value2
#range(v1,v2,v3)= start from v1 and go upto v2 and skip every v3
#for x in range(2,9,2):
#x=[2,4,6,8]
#print all value of variable one by one
for x in range(2,9,2):
print("The value of x is:",x)
# i=[1,2,3,4,5]
# i=5
for i in range(1,6):
print(i*"*")
# while loop
# while condition:
# statement
# incrementer
x=0
while x<4:
print(x)
x+=1
for x in range(1,10):
if x==6:
print("Existing loop")
break
else:
print("The value of x is :",x)
def add(a,b):
print("a=",a)
print("b=",b)
return a+b
c= add(5,2)
print(c)
```
###### for i=[1,2,3,4,5,6]
#fori=5
for i in range(1,6):
print("i=1,i<=6,i++")
```
i=1
for i in range(1,6):
j=1
for j in range(i,i+1):
print(j,end=" ")
print()
class student:
def_init_(self,sname,sage,spercent):
listt=['kar','abcd',706,2.33,'johny',70.2,36.3,755]
listt
type(listt[0])
type(listt[3])
int i2;
double d2;
char s2[100]; // this is not scalable for input of unknown size
// Read inputs from stdin
scan("%d", &i2);
scan("%lf", &d2);
scan("%*[\n] %[^\n]", s2);
// Print outputs to stdout
print("%d\n", i + i2);
print("%.01lf\n", d + d2);
print("%s%s", s, s2);
i=int(input(4))
d=int(input(4.0))
s=int(input(Hackerank))
i2 = int(input(12)) # read int
d2 = float(input()) # read double
s2 = input() # read string
# print summed and concatenated values
print(i + i2)
print(d + d2)
print(s + s2)
>>>str="hello"
>>>str[:2]
>>>
import maths
def add(a,b):
return a+b
def sub(a,b):
return a-b
def mul(a,b):
return a*b
def div(a,b):
return a/b
def sqrt(a)
return (math,sqrt.(a))
def powa
return (a**a)
while true:
enter = input("enter function name:")
if function== 'addition'
try:
a =int(input("enter first number"))
b=int(input("enter second number"))
print(add(a,b))
except Value error:
print("please provide valid numbers")
if function == 'subtraction'
try:
a= int(input("enter first number"))
b= int(input("enter second number"))
print(sub(a,b))
except Value error:
print("please provide valid numbers")
if function == 'multiplication'
try:
a=int(input("enter first number"))
b= int(input("enter second number"))
print(mul(a,b))
except Value error:
print("please provide valid numbers")
if function ==;'division'
try :
a=int(input("enter first number"))
b= int(input("enter second number"))
print(div(a,b))
except Value error:
print("please")
import numpy as np
a= np.arange(30).reshape(2,15)
a
```
| github_jupyter |
### Note
* Instructions have been included for each segment. You do not have to follow them exactly, but they are included to help you think through the steps.
```
# Dependencies and Setup
import pandas as pd
# File to Load (Remember to Change These)
file_to_load = "Resources/purchase_data.csv"
# Read Purchasing File and store into Pandas data frame
purchase_data = pd.read_csv(file_to_load)
#Display the purchase DataFrame
purchase_data.head()
```
## Player Count
* Display the total number of players
```
#Create a dataframe using the lenght of the unique values in the "SN" column
players = pd.DataFrame([{"Total Players" : len(purchase_data["SN"].value_counts())}])
players
```
## Purchasing Analysis (Total)
* Run basic calculations to obtain number of unique items, average price, etc.
* Create a summary data frame to hold the results
* Optional: give the displayed data cleaner formatting
* Display the summary data frame
```
#Counting the number of unique items
unique_items = len(purchase_data["Item Name"].value_counts())
#Calculate the average price
avg_price = purchase_data["Price"].mean()
#Total the number of purchases
purchases = purchase_data["Purchase ID"].count()
#Calculate the total revenue
total_revenue = purchase_data["Price"].sum()
#Create a DataFrame using the created values
summary_table = pd.DataFrame([{"Number of Unique Items": unique_items,
"Average Price": avg_price,
"Number of Purchases" : purchases,
"Total Revenue": total_revenue}])
#Changing the format so Average price and total revenue are shown as a currency
summary_table["Average Price"] = summary_table["Average Price"].map('${:,.2f}'.format)
summary_table["Total Revenue"] = summary_table["Total Revenue"].map('${:,.2f}'.format)
#Display the summary table
summary_table
```
## Gender Demographics
* Percentage and Count of Male Players
* Percentage and Count of Female Players
* Percentage and Count of Other / Non-Disclosed
```
#First remove the duplicates from the 'SN' column so the counts are accurate
unique_df = purchase_data.drop_duplicates(subset = ['SN'])
#Get the total count of players
total_players = unique_df['SN'].count()
#Get the value counts for the genders
gender_counts = unique_df['Gender'].value_counts()
#Calculate the percentages of each gender
male_percent = (gender_counts[0] / total_players)
female_percent = (gender_counts[1] / total_players)
other_percent = (gender_counts[2] / total_players)
#Create a percentage list for a pd dataframe
percentages = [male_percent, female_percent, other_percent]
#Create a pandas DataFrame and add the percentage column
gender_df = pd.DataFrame(gender_counts)
gender_df['Percentage of Players'] = percentages
#Format the percentage column and rename the gender column
gender_df['Percentage of Players'] = gender_df['Percentage of Players'].map('{:.2%}'.format)
gender_df = gender_df.rename(columns = {"Gender" : "Total Count"})
#Display the gender dataframe
gender_df
```
## Purchasing Analysis (Gender)
* Run basic calculations to obtain purchase count, avg. purchase price, avg. purchase total per person etc. by gender
* Create a summary data frame to hold the results
* Optional: give the displayed data cleaner formatting
* Display the summary data frame
```
#Create a df for the gender counts then sort so the index is alphabetized
gender_df = pd.DataFrame(purchase_data['Gender'].value_counts())
gender_df = gender_df.sort_index()
#Getting a df for each gender
female_df = purchase_data.loc[purchase_data['Gender'] == "Female", :]
male_df = purchase_data.loc[purchase_data['Gender'] == "Male", :]
other_df = purchase_data.loc[purchase_data['Gender'] == "Other / Non-Disclosed", :]
#Calculating the total purchase for each gender
female_total = female_df['Price'].sum()
male_total = male_df['Price'].sum()
other_total = other_df['Price'].sum()
#Calculate the average price for each gender
female_mean = female_df['Price'].mean()
male_mean = male_df['Price'].mean()
other_mean = other_df['Price'].mean()
#Getting the unique gender counts from the df created earlier
un_gender =unique_df['Gender'].value_counts()
#Calculating the average total per Person
female_avg = female_total / un_gender[1]
male_avg = male_total / un_gender[0]
other_avg = other_total / un_gender[2]
#Creating the lists to append to the summary df
avg_price = [female_mean, male_mean, other_mean]
total_value = [female_total, male_total, other_total]
avg_per_person = [female_avg, male_avg, other_avg]
#Appending the new columns to the dataframe
gender_df['Average Purchase Price'] = avg_price
gender_df['Total Purchase Value'] = total_value
gender_df['Avg Total Purchase Per Person'] = avg_per_person
#Renaming the gender column and formatting the values
gender_df['Average Purchase Price'] = gender_df['Average Purchase Price'].map('${:,.2f}'.format)
gender_df['Total Purchase Value'] = gender_df['Total Purchase Value'].map('${:,.2f}'.format)
gender_df['Avg Total Purchase Per Person'] = gender_df['Avg Total Purchase Per Person'].map('${:,.2f}'.format)
gender_df = gender_df.rename(columns = {"Gender" : "Purchase Count"})
gender_df
```
## Age Demographics
* Establish bins for ages
* Categorize the existing players using the age bins. Hint: use pd.cut()
* Calculate the numbers and percentages by age group
* Create a summary data frame to hold the results
* Optional: round the percentage column to two decimal points
* Display Age Demographics Table
```
#First we will create the bins for the ages
bins = [0,9.9,14.9,19.9,24.9,29.9,34.9,39.9,1000]
#Create a name for each of the bins
group_names = ["<10", "10 to 14", "15 to 19",
"20 to 24", "25 to 29", "30 to 34",
"35 to 39", "40+"]
#Us pd.cut to create a column for the demographics using the unique df from earlier
demographics = pd.DataFrame(pd.cut(unique_df["Age"], bins, labels=group_names, include_lowest=True))
#Use value_counts to total the brackets and create a dataframe
summary_demo = pd.DataFrame(demographics["Age"].value_counts())
#Sorting and renaming the column
summary_demo = summary_demo.sort_index()
summary_demo = summary_demo.rename(columns = {"Age" : "Total Count"})
#Calculating percentage by age group
#First get the total count for all groups
total_count = summary_demo["Total Count"].sum()
#Calculate the average for each age band
averages = [(value / total_count) for value in summary_demo["Total Count"]]
#Add the averages to the DataFrame and format the column to percent
summary_demo["Percentage of Players"] = averages
summary_demo["Percentage of Players"] = summary_demo["Percentage of Players"].map('{:.2%}'.format)
#Display the data
summary_demo
```
## Purchasing Analysis (Age)
* Bin the purchase_data data frame by age
* Run basic calculations to obtain purchase count, avg. purchase price, avg. purchase total per person etc. in the table below
* Create a summary data frame to hold the results
* Optional: give the displayed data cleaner formatting
* Display the summary data frame
```
#Note: at this point, I realized I had not been using groupby and I did not feel like going back and re-doing
#the first parts again, groupby made this a lot easier
#Using the bins from before, we will append the age bands to the entire df and the unique df
purchase_data['Age Band'] = pd.cut(purchase_data["Age"], bins, labels=group_names, include_lowest=True)
#Next we will make a groupby for the age bands
grouped = purchase_data.groupby("Age Band")
#We will count the totals for the age bands
band_counts = grouped['Purchase ID'].count()
#Using groupby we will get the total and the average per band
total_bands = grouped['Price'].sum()
avg_bands = grouped['Price'].mean()
#Get the individual counts from the unique df earlier
u_grouped = unique_df.groupby("Age Band")
Uband_counts = u_grouped['Purchase ID'].count()
#Calcuate the average per person using the total purchase and the unique counts
per_person = total_bands / Uband_counts
#Add all of the calculations to a summary df
analysis = pd.DataFrame(band_counts)
analysis['Average Purchase Price'] = avg_bands
analysis['Total Purchase Value'] = total_bands
analysis['Avg Total Purchase Per Person'] = per_person
#Formatting the columns
analysis['Average Purchase Price'] = analysis['Average Purchase Price'].map('${:,.2f}'.format)
analysis['Total Purchase Value'] = analysis['Total Purchase Value'].map('${:,.2f}'.format)
analysis['Avg Total Purchase Per Person'] = analysis['Avg Total Purchase Per Person'].map('${:,.2f}'.format)
analysis = analysis.rename(columns = {"Purchase ID" : "Purchase Count"})
#Display the results
analysis
```
## Top Spenders
* Run basic calculations to obtain the results in the table below
* Create a summary data frame to hold the results
* Sort the total purchase value column in descending order
* Optional: give the displayed data cleaner formatting
* Display a preview of the summary data frame
```
#Using groupby we will group by the sn
sn_group = purchase_data.groupby('SN')
#First we will total the SN
sn_total = sn_group['Item ID'].count()
#Next we get the purchase sum for each SN
sn_price = sn_group['Price'].sum()
#Calculate the average for each SN
sn_mean = sn_group['Price'].mean()
#Create a summary dataframe
sn_data = pd.DataFrame(sn_total)
sn_data['Average Purchase Price'] = sn_mean
sn_data['Total Purchase Value'] = sn_price
#Sort and format the columns
sn_data['Average Purchase Price'] = sn_data['Average Purchase Price'].map('${:,.2f}'.format)
sn_data['Total Purchase Value'] = sn_data['Total Purchase Value'].map('${:,.2f}'.format)
sn_data = sn_data.sort_values(by='Item ID', ascending = False)
sn_data = sn_data.rename(columns = {"Item ID" : "Purchase Count"})
#Display the data
sn_data.head()
```
## Most Popular Items
* Retrieve the Item ID, Item Name, and Item Price columns
* Group by Item ID and Item Name. Perform calculations to obtain purchase count, average item price, and total purchase value
* Create a summary data frame to hold the results
* Sort the purchase count column in descending order
* Optional: give the displayed data cleaner formatting
* Display a preview of the summary data frame
```
#I used the same code as before but I changed the grouping from SN to Items and Item ID
#Using groupby we will group by the Item ID and Item Name
item_group = purchase_data.groupby(['Item ID', 'Item Name'])
#First we will total the SN
item_total = item_group['SN'].count()
#Next we get the purchase sum for each Item
item_price = item_group['Price'].sum()
#Calculate the average for each Item
item_mean = item_group['Price'].mean()
#Create a summary dataframe
item_data = pd.DataFrame(item_total)
item_data['Item Price'] = item_mean
item_data['Total Purchase Value'] = item_price
# #Sort and format the columns
item_data['Item Price'] = item_data['Item Price'].map('${:,.2f}'.format)
item_data['Total Purchase Value'] = item_data['Total Purchase Value'].map('${:,.2f}'.format)
item_data = item_data.sort_values(by='SN', ascending = False)
item_data = item_data.rename(columns = {"SN" : "Purchase Count"})
# #Display the data
item_data.head()
```
## Most Profitable Items
* Sort the above table by total purchase value in descending order
* Optional: give the displayed data cleaner formatting
* Display a preview of the data frame
| github_jupyter |
```
# export
import pandas as pd
import numpy as np
from PIL import Image
from sklearn.model_selection import train_test_split
import torch
from torch.utils.data import Dataset, DataLoader
from fastcore.all import *
# default_exp data_anime_heads
data_dir = Path('../data/tiny_data/anime_heads')
```
# Data AnimeHeads
>
## Items
```
# exporti
def get_items(data_dir, pct=1, valid_pct=0.2):
df = pd.read_csv(data_dir/'tags.csv', header=None, names=['id', 'cap'])
df = df[:int(len(df)*pct)]
if valid_pct==0:
return df, pd.DataFrame(data=None, columns=['id', 'cap'])
train_items, valid_items = train_test_split(df, test_size=valid_pct, random_state=42, shuffle=True, stratify=df.cap)
return train_items, valid_items
train_items, valid_items = get_items(data_dir)
test_eq(len(train_items), 240)
test_eq(len(valid_items), 60)
train_items[:5]
```
## Datasets
```
# export
class Tokenizer():
def __init__(self):
self.vocab = [
'<pad>', 'orange hair', 'white hair', 'aqua hair', 'gray hair','green hair', 'red hair',
'purple hair', 'pink hair','blue hair', 'black hair', 'brown hair', 'blonde hair', 'black eyes', 'orange eyes',
'purple eyes', 'pink eyes', 'yellow eyes', 'aqua eyes', 'green eyes', 'brown eyes', 'red eyes', 'blue eyes',
]
self.o2i = {v:k for k,v in enumerate(self.vocab)}
self.max_seq_len = 2
self.vocab_sz = len(self.vocab)
self.pad_id = 0
def encode(self, cap):
"cap: 'aqua hair aqua eyes', returns: tag: [2, 17], tag_len: 2"
cap = cap.split()
tags = [' '.join(cap[:2]), ' '.join(cap[2:])]
return [self.o2i[tags[0]], self.o2i[tags[1]]], self.max_seq_len
def decode(self, o):
"o: [2, 17], returns: 'aqua hair aqua eyes'"
tags = [self.vocab[idx] for idx in o]
# tags = [self.vocab[o[0]], self.vocab[o[1]]]
return ' '.join(tags)
tokenizer = Tokenizer()
ori_cap = 'aqua hair aqua eyes'
tags, tag_len = tokenizer.encode(ori_cap)
test_eq(tags, [3, 18])
test_eq(tag_len, 2)
out_cap = tokenizer.decode(tags)
test_eq(out_cap, ori_cap)
# exporti
class AnimeHeadsDataset(Dataset):
def __init__(self, items, data_dir):
"items: df of id and cap"
self.data_dir = data_dir
self.items = list(items.itertuples(index=False, name=None))
self.tokenizer = Tokenizer()
def __len__(self):
return len(self.items)
def __getitem__(self, idx):
return self.tfm(self.items[idx])
def tfm(self, item):
''' item: (0, aqua hair aqua eyes),
returns: tag: (2,), tag_len: (), img64: (64, 64, 3) '''
img_id, cap = item
tag, tag_len = self.tokenizer.encode(cap)
img_path = self.data_dir/f'imgs/{img_id}.jpg'
img64 = np.array(Image.open(img_path))
if len(img64.shape)==2:
img64 = np.repeat(img64[...,None], 3, axis=2)
return torch.tensor(tag), torch.tensor(tag_len), torch.tensor(img64)
ds = AnimeHeadsDataset(train_items, data_dir)
tag, tag_len, img64 = ds[0]
test_eq(tag.shape, (2,))
test_eq(tag_len.shape, ())
test_eq(img64.shape, (64, 64, 3))
print(tag, tag_len)
plt.imshow(img64)
# export
class Datasets():
def __init__(self, data_dir, pct=1, valid_pct=0.2):
train_items, valid_items = get_items(data_dir, pct=pct, valid_pct=valid_pct)
self.train = AnimeHeadsDataset(train_items, data_dir)
self.valid = AnimeHeadsDataset(valid_items, data_dir)
dsets = Datasets(data_dir)
test_eq(len(dsets.train), 240)
test_eq(len(dsets.valid), 60)
```
## DataLoaders
```
# export
class DataLoaders():
def __init__(self, dsets, bs=64):
self.dsets = dsets
self.train = DataLoader(dsets.train, batch_size=bs, shuffle=True, num_workers=2, drop_last=True)
self.valid = DataLoader(dsets.valid, batch_size=bs, shuffle=False, num_workers=2)
dls = DataLoaders(dsets, bs=16)
for tag, tag_len, img in dls.train:
test_eq(tag.shape, (16, 2))
test_eq(tag_len.shape, (16,))
test_eq(img.shape, (16, 64, 64, 3))
break
```
## Export -
```
# hide
from nbdev.export import notebook2script
notebook2script()
```
| github_jupyter |
```
# check out list words and the apostrophe situation in the step for loop
# check out allowing more than maxseqlen in generate text
# look into batch size
#text generation apostrpphe breaking
import pandas as pd
import numpy as np
from keras.preprocessing.sequence import pad_sequences
from keras.optimizers import RMSprop
from keras import optimizers
import sys
from keras.callbacks import LambdaCallback
import random
import matplotlib.pyplot as plt
def sample(preds, temperature=1.0):
# helper function to sample an index from a probability array
preds = np.asarray(preds).astype('float64')
preds = np.log(preds) / temperature
exp_preds = np.exp(preds)
preds = exp_preds / np.sum(exp_preds)
probas = np.random.multinomial(1, preds, 1)
return np.argmax(probas)
def generate_text(seed_text, numb_next_words):
output=seed_text
for i in range (numb_next_words):
words_gen = set(seed_text.split())
words_gen=list(words_gen) #create list of unique words in seed text
# for i in range (len(words_gen)): #replace all ' in seed text
# words_gen[i]=words_gen[i].replace("‘", '').replace("’", '').replace("'", '')
#create a dictionary with index and word
word_indices_gen = dict((c, i) for i, c in enumerate(words_gen, 1))
#turn sentence into a sequence of numbers
sequence=[]
for word in seed_text.split():
sequence.append(word_indices_gen[word])
sequence_padded = pad_sequences([sequence], maxlen=10, padding='pre')
# sequence_padded=sequence
#create an embedding matrix with same indices as word_index
EMBEDDING_DIM=25
total_words=len(word_indices_gen)+1
embedding_matrix = np.zeros((total_words, EMBEDDING_DIM))
for word, i in word_indices_gen.items():
embedding_vector = embeddings_index.get(word)
if embedding_vector is not None:
# words not found in embedding index will be all-zeros.
embedding_matrix[i] = embedding_vector
#create X input with embedding matrix for specific words (by their index)
gener=[]
for number in sequence_padded:
gener.append(embedding_matrix[number])
predicted=model.predict([gener], verbose=0)
predicted=sample(predicted[0])
output_word=""
for word, index in word_indices.items():
if index == predicted:
output_word = word
break
output+=" " + output_word
seed_text+=" " + output_word
seed_text=seed_text.split(' ', 1)[1]
return output
def on_epoch_end(epoch, _):
# Function invoked at end of each epoch. Prints generated text.
print()
print('----- Generating text after Epoch: %d' % epoch)
start_index = random.randint(0, len(listofwords) - maxlen - 1)
for diversity in [0.5, 1.0]:
print('----- diversity:', diversity)
generated = ''
sentence = listofwords[start_index: start_index + maxlen].str.cat(sep=' ')
generated += sentence
print('----- Generating with seed: "' + sentence + '"')
sys.stdout.write(generate_text(generated, 20))
tweet_data=pd.read_csv('../Load_Tweets/data/tweet_data.csv')
tweet_text = tweet_data['TEXT']
tweet_text_all = tweet_data['TEXT'].str.cat(sep=' ')
listofwords=pd.Series(tweet_text_all.split())
tweet_text.head()
# top_words=listofwords.value_counts()
# top_words_percent= top_words/len(listofwords)
# top_words.head(50).plot.bar()
# # top_words.head(50)
total_chars=len(tweet_text_all)
total_chars
total_wordz=len((tweet_text_all.split()))
total_wordz
chars = set(tweet_text_all)
words = set(tweet_text_all.split())
print ("total number of unique words", len(words))
print ("total number of unique chars", len(chars))
words=list(words)
#replace apostrophes in dictionary keys
for i in range (len(words)):
words[i]=words[i].replace("‘", '').replace("’", '').replace("'", '')
words=set(words)
len(words)
#create forward and reverse word index
word_indices = dict((c, i) for i, c in enumerate(words, 1))
indices_word = dict((i, c) for i, c in enumerate(words,1 ))
len(word_indices)
max(word_indices.values())
#choose step
maxlen = 10
step = 2
sentences = []
next_words = []
next_words = []
list_words = []
sentences2 = []
for i in range (len(tweet_text)):
list_words = tweet_text.iloc[i].split()
for i in range(len( list_words)):
list_words[i]=list_words[i].replace("‘", '').replace("’", '').replace("'", '')
for i in range(0, len(list_words) - maxlen, step):
sentences2 = ' '.join(list_words[i: i + maxlen])
sentences.append(sentences2)
next_words.append((list_words[i + maxlen]))
print ('length of sentence list:', len(sentences))
print ("length of next_word list", len(next_words))
sequences=[]
y=[]
for i, sentence in enumerate(sentences):
sequence=[]
for j, word in enumerate(sentence.split()):
sequence.append(word_indices[word])
sequences.append(sequence)
y.append(word_indices[next_words[i]])
sequences=np.asarray(sequences)
sequences.shape
sequences
total_words= len(word_indices)+1
total_words
embeddings_index = {}
f = open('../word_embeding/glove.twitter.27B.25d.txt')
for line in f:
values = line.split()
word = values[0]
coefs = np.asarray(values[1:], dtype='float32')
embeddings_index[word] = coefs
f.close()
print('Found %s word vectors.' % len(embeddings_index))
max(word_indices.values())
len(word_indices)
EMBEDDING_DIM=25
embedding_matrix = np.zeros((total_words, EMBEDDING_DIM))
for word, i in word_indices.items():
embedding_vector = embeddings_index.get(word)
if embedding_vector is not None:
# words not found in embedding index will be all-zeros.
embedding_matrix[i] = embedding_vector
embedding_matrix.shape
X=sequences[0:100]
y=y[0:100]
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import Flatten
from keras.layers import Embedding
from keras.layers import Dropout
from keras.layers import LSTM
from keras.layers import Input
from keras.regularizers import L1L2
from keras import regularizers
from keras import metrics
# embedding_layer= Embedding(total_words, EMBEDDING_DIM, weights=[embedding_matrix],input_length=max_seq,trainable=False)
# sequence_input = Input(shape=(max_seq,), dtype='int32')
# embedded_sequences= embedding_layer(sequence_input)
model=Sequential()
# e=Embedding(total_words, EMBEDDING_DIM, weights=[embedding_matrix],input_length=maxlen,trainable=False)
# model.add(e)
from keras.layers import Embedding
model.add( Embedding(total_words,
EMBEDDING_DIM,
weights=[embedding_matrix],
input_length=maxlen,
input_shape= (maxlen,),
trainable=False))
model.add(LSTM(128, bias_regularizer=regularizers.l1(0.01)))
model.add(Dropout(0.2))
# model.add(LSTM(512, return_sequences=False))
# model.add(Dropout(0.1))
# model.add(Flatten())
model.add(Dense(total_words, activation="softmax"))
optimizer = RMSprop(lr=0.01)
# sgd = optimizers.SGD(lr=0.01, clipvalue=0.5)
model.compile(loss='sparse_categorical_crossentropy', metrics=['accuracy'], optimizer=optimizer)
model.summary()
# model.add(LSTM(128, input_shape=(maxlen, len(chars))))
# model.add(Dense(len(chars), activation='softmax'))
# model.compile(loss='categorical_crossentropy', optimizer=optimizer)
# from keras.models import load_model
# model= load_model("../Saved_models/failed_on_99th_epoch_word_embedding")
# print_callback = LambdaCallback(on_epoch_end=on_epoch_end)
from keras.callbacks import EarlyStopping
early_stopping = EarlyStopping(monitor='val_acc', patience=3)
model.fit(X, y, validation_split=0.2, epochs=10,callbacks=[early_stopping])
loss, accuracy = model.evaluate(X_test_sample, y_test_sample, verbose=0)
print('Accuracy: %f' % (accuracy*100))
print('loss: %f' % (loss))
perplexity = np.exp2(loss)
print ('perplexity: {}'.format(perplexity))
print (generate_text("i will", 20))
# model.save('../failed_on_99th_epoch_word_embedding')
predictions_test=model.predict(X_test)
len(predictions_test)
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from IPython.core.display import display, HTML
plt.style.use('fivethirtyeight')
plt.rc('figure', figsize=(5.0, 2.0))
pokemon=pd.read_csv("../dataset/pokemon.csv")
# Which pokémon is the most difficult to catch?
pokemon['capture_rate']=pd.to_numeric(pokemon['capture_rate'])
pokemon['name'][pokemon['capture_rate']==min(pokemon['capture_rate'])]
#Which no-legendary pokémon is the most diffucult to catch? </b>
no_legendary=pokemon[pokemon['is_legendary']==False]
no_legendary['name'][no_legendary['capture_rate']==min(no_legendary['capture_rate'])]
display(HTML("<img src='../img/beldum.png' width='200px' height='200px'>"))
rate=pokemon[pokemon['name']=='Beldum']['capture_rate'].values
beldum_rate=num = "{0:.2f}".format((rate[0]*100)/255)
print("Name: Beldum\n"+"Percentage of catch: " + beldum_rate + " %")
```
<div class="alert alert-info">
The min value for attack/special-attack and defense/special-defense statistics was calculated from the subsets of the pokemon which have the highest physical/special statistic compared to the special/physical equivalent. In this way the results acquire greater relevance.
</div>
```
#Speed
pokemon['name'][pokemon['speed']==max(pokemon['speed'])]
pokemon['name'][pokemon['speed']==min(pokemon['speed'])]
```
<b> Atk </b>
```
pokemon['name'][pokemon['attack']==max(pokemon['attack'])]
physical_atk=pokemon[pokemon['attack']>=pokemon['sp_attack']]
physical_atk['name'][physical_atk['attack']==min(physical_atk['attack'])]
```
<b>Def</b>
```
pokemon['name'][pokemon['defense']==max(pokemon['defense'])]
physical_def=pokemon[pokemon['defense']>=pokemon['sp_defense']]
physical_def['name'][physical_def['defense']==min(physical_def['defense'])]
```
<b> Sp.Atk</b>
```
pokemon['name'][pokemon['sp_attack']==max(pokemon['sp_attack'])]
special_atk=pokemon[pokemon['sp_attack']>=pokemon['attack']]
special_atk['name'][special_atk['sp_attack']==min(special_atk['sp_attack'])]
```
<b>Sp.Def</b>
```
pokemon['name'][pokemon['sp_defense']==max(pokemon['sp_defense'])]
special_def=pokemon[pokemon['sp_defense']>=pokemon['defense']]
special_def['name'][special_def['sp_defense']==min(special_def['sp_defense'])]
```
<b>Hp</b>
```
pokemon['name'][pokemon['hp']==max(pokemon['hp'])]
pokemon['name'][pokemon['hp']==min(pokemon['hp'])]
```
Combining all the information we can see how <code>Shuckle</code> is a pokémon with <b>very particular statistics</b>. Look at them:
```
display(HTML("<img src='../img/shuckle.png' width='200px' height='200px'>"))
pokemon.iloc[212][['name','hp','attack','sp_attack','defense','sp_defense','speed']]
```
# Which type is the most common?
To answer this question, I think it's more interesting seeing the <b>absolute frequencies</b> for each type of pokémon in a <b>bar chart</b>.
```
types_abs_freq=(pokemon['type1'].value_counts()+pokemon['type2'].value_counts()).sort_values(ascending=False)
x=types_abs_freq.index
y=types_abs_freq.values
types_abs_freq.plot.bar()
plt.show()
```
<div class="alert alert-info">
Absolute frequencies were calculated from a set constructed as the union between the set of types 1 and 2 of each pokémon.
</div>
The result obtained shows us a subdivision of the pokémon by type rather conform to reality: the pokémon closest to have an animal correspondent in the real world are the most widespread.<br>
<b>The most common type is water</b> but the most interesting data is that the psychic type is the fifth most common type, even exceeding the bug type.
# Which ability is the most common?
We answer this question by printing the top 10 most common abilities.
```
ser_abilities=pokemon['abilities']
abilities=[]
for i in range(0,801):
arr_ab=ser_abilities[i].split(',')
for j in range(0,len(arr_ab)):
ability=arr_ab[j].replace("[","").replace("'","").replace("]","")
abilities.append(ability)
abilities_freq=pd.Series(abilities).value_counts().sort_values(ascending=False)
abilities_freq.head(10)
```
<b> Be very careful to do earthquake! </b>
# Correlation
```
import seaborn as sns
plt.figure(figsize=(20,20))
sns.heatmap(pokemon.corr(), linewidths=.5)
plt.show()
```
There is a strong positive correlation between:
- generation and pokédex number (good information for building generation clusters),
- against_ghost and against_dark (thanks to ghost type),
- base_egg_steps and is_legendary (good information for building legendary classifier).
There is a good positive correlation between:
- single stats and base_total,
- height and weight.
There is a strong negative correlation between:
- capture_rate and base_total,
- single stats and capture_rate,
- against_fight and against_ghost (thanks to normal type),
- against_psychic and against_bug (thanks to dark type),
- against_ground and against_ice (Why?),
- base_happiness and base_egg_steps.
There is a good negative correlation between:
- base_happiness and weight,
- base_happiness and is_legendary.
And so on.
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.