code stringlengths 2.5k 150k | kind stringclasses 1
value |
|---|---|
# Homework 6
## Due: Tuesday, October 10 at 11:59 PM
# Problem 1: Bank Account Revisited
We are going to rewrite the bank account closure problem we had a few assignments ago, only this time developing a formal class for a Bank User and Bank Account to use in our closure (recall previously we just had a nonlocal var... | github_jupyter |
```
%matplotlib notebook
import numpy as np
import matplotlib.pyplot as plt
```
# Utilizando un modelo pre-entrenado
[`torchvision.models`](https://pytorch.org/vision/stable/models.html) ofrece una serie de modelos famosos de la literatura de *deep learning*
Por defecto el modelo se carga con pesos aleatorios
Si in... | github_jupyter |
<a href="https://colab.research.google.com/github/saketkc/pyFLGLM/blob/master/Chapters/01_Chapter01.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
## Chapter 1 - Introduction to Linear and Generalized Linear Models
```
import warnings
import pand... | github_jupyter |
# Tutorial about loading localization data from file
```
from pathlib import Path
import locan as lc
lc.show_versions(system=False, dependencies=False, verbose=False)
```
Localization data is typically provided as text or binary file with different formats depending on the fitting software. Locan provides functions ... | github_jupyter |
```
# Copyright 2021 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writi... | github_jupyter |
# Pull census data for the neighborhoods in Seattle
Use this link to find tables: https://api.census.gov/data/2018/acs/acs5/variables.html
```
import pandas as pd
import censusdata
import csv
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import scipy
from scipy import stats
sample = censusda... | github_jupyter |
# Playing with Matplotlib
Please note I am making no assumptions nor any conclusions as I have not studied this data, it's actual original source, the source I got it from or even looked at most of the dataset itself. It is just some data to make graphs with and part of a tutorial.
```
import pandas as pd
#Demo use... | github_jupyter |
```
import azureml
from azureml.core import Workspace, Experiment, Datastore, Environment
from azureml.core.runconfig import RunConfiguration
from azureml.data.datapath import DataPath, DataPathComputeBinding
from azureml.data.data_reference import DataReference
from azureml.core.compute import ComputeTarget, AmlComput... | github_jupyter |
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from datetime import datetime, timedelta
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import OneHotEncoder
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from ... | github_jupyter |
# Levy Stable models of Stochastic Volatility
This tutorial demonstrates inference using the Levy [Stable](http://docs.pyro.ai/en/stable/distributions.html#stable) distribution through a motivating example of a non-Gaussian stochastic volatilty model.
Inference with stable distribution is tricky because the density `... | github_jupyter |
### Send email Clint
#### Importing all dependency
```
# ! /usr/bin/python
import smtplib
from email.mime.multipart import MIMEMultipart
from email.mime.text import MIMEText
from email.header import Header
from email.utils import formataddr
import getpass
```
#### User Details Function
```
def user():
# ORG_EMAI... | github_jupyter |
# healthy versus severe
```
import re
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import pydotplus
from IPython.display import Image
from six import StringIO
import matplotlib.image as mpimg
#%pylab inline
from sklearn.tree import DecisionTreeClassifier, DecisionTreeRegressor, plot_tree, ... | github_jupyter |
```
import base64
import imageio
import IPython
import matplotlib
import matplotlib.pyplot as plt
import PIL.Image
import pyvirtualdisplay
import tensorflow as tf
from tf_agents.agents.dqn import dqn_agent
from tf_agents.agents.dqn import q_network
from tf_agents.drivers import dynamic_step_driver
from tf_agents.enviro... | github_jupyter |
```
!pip install --upgrade tables
!pip install eli5
!pip install xgboost
import pandas as pd
import numpy as np
from sklearn.dummy import DummyRegressor
from sklearn.tree import DecisionTreeRegressor
from sklearn.ensemble import RandomForestRegressor
import xgboost as xgb
from sklearn.metrics import mean_absolute_er... | github_jupyter |
<a href="https://colab.research.google.com/github/LeonardoQZ/handson-ml2/blob/master/CaliforniaGeostats.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# California Housing with Geostatistics
```
import pandas as pd
import numpy as np
from sklearn... | github_jupyter |
# Lab Three
Ryan Gonfiantini
---
For this lab we're going to be making and using a bunch of functions.
Our Goals are:
- Searching our Documentation
- Using built in functions
- Making our own functions
- Combining functions
- Structuring solutions
```
# For the following built in functions we didn't touch on them i... | github_jupyter |
# 6.7 门控循环单元(GRU)
## 6.7.2 读取数据集
```
import numpy as np
import torch
from torch import nn, optim
import torch.nn.functional as F
import sys
sys.path.append("..")
import d2lzh_pytorch as d2l
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
(corpus_indices, char_to_idx, idx_to_char, vocab_size) =... | github_jupyter |
# The JupyterLab Interface
The JupyterLab interface consists of a main work area containing tabs of documents and activities, a collapsible left sidebar, and a menu bar. The left sidebar contains a file browser, the list of running terminals and kernels, the table of contents, and the extension manager.
, a drop-in replacement for the MNIST dataset. MNIST is actually quite trivial with neural networks where you can easily achieve better than 9... | github_jupyter |
```
# Automatically reload imported modules that are changed outside this notebook
%load_ext autoreload
%autoreload 2
# More pixels in figures
import matplotlib.pyplot as plt
%matplotlib inline
plt.rcParams["figure.dpi"] = 200
# Init PRNG with fixed seed for reproducibility
import numpy as np
np_rng = np.random.defau... | github_jupyter |
### Load Dataset
```
import pandas as pd
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
import seaborn as sns
# 노트북 안에 그래프를 그리기 위해
%matplotlib inline
# 그래프에서 마이너스 폰트 깨지는 문제에 대한 대처
mpl.rcParams['axes.unicode_minus'] = False
import warnings
warnings.filterwarnings('ignore')
train = pd.re... | github_jupyter |
# 数学函数、字符串和对象
## 本章介绍Python函数来执行常见的数学运算
- 函数是完成一个特殊任务的一组语句,可以理解为一个函数相当于一个小功能,但是在开发中,需要注意一个函数的长度最好不要超过一屏
- Python中的内置函数是不需要Import导入的
<img src="../Photo/15.png"></img>
```
a = -10
print(abs(a))
max
b = -10.1
print(abs(b))
c = 0
print(abs(c))
max(1, 2, 3, 4, 5)
min(1, 2, 3, 4, 5)
min(1, 2, 3, -4, 5)
for i in range(10):
... | github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
Licensed under the Apache License, Version 2.0 (the "License");
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.o... | github_jupyter |
<img src="https://cybersecurity-excellence-awards.com/wp-content/uploads/2017/06/366812.png">
<h1><center>Darwin Supervised Classification Model Building </center></h1>
Prior to getting started, there are a few things you want to do:
1. Set the dataset path.
2. Enter your username and password to ensure that you're a... | github_jupyter |
# Hashing
## Lineare Sondierung
Bei einer Kollision versuchen wir die nächste freie Stelle in unserer Hashtabelle zu suchen. Dieses Verhalten wird durch die Formel:<br> $h(k, i) = (h'(k) + i) \bmod m$ mit $h'(k) = k \bmod m$<br>
m ist die Größe der Hashtabelle ausgedrückt.<br>
Im ersten Durchlauf ist i = 0. Tritt eine... | github_jupyter |
# Mandala: self-managing experiments
## What is Mandala?
Mandala enables new, simpler patterns for working with complex and evolving
computational experiments.
It eliminates low-level code and decisions for how to save, load, query,
delete and otherwise organize results. To achieve this, it lets computational
code "m... | github_jupyter |
# notebook for processing fully reduced m3 data "triplets"
This is a notebook for processing L0 / L1B / L2 triplets (i.e.,
the observations that got reduced).
## general notes
We process the reduced data in triplets simply to improve the metadata on the
L0 and L2 products. We convert L1B first to extract several attr... | github_jupyter |
```
import os
from ipywidgets import Output, HBox, Layout
import jupyter_cadquery
icon_path = os.path.join(os.path.dirname(jupyter_cadquery.__file__), "icons")
```
# ipywidgets
```
from ipywidgets import interact, interactive, fixed, interact_manual
import ipywidgets as widgets
def f(x):
return x
interact(f,... | github_jupyter |
# Measure Watson Assistant Performance
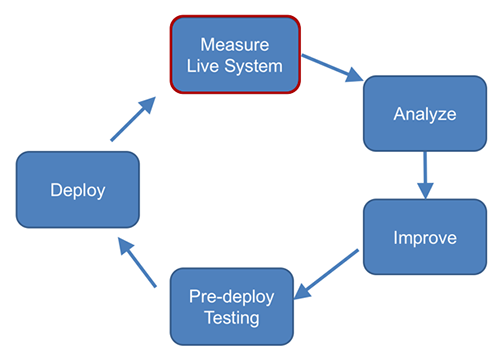
## Introduction
This notebook demonstrates how to setup automated metrics that help you measure, monitor, and und... | github_jupyter |
# Modeling and Simulation in Python
Chapter 4
Copyright 2017 Allen Downey
License: [Creative Commons Attribution 4.0 International](https://creativecommons.org/licenses/by/4.0)
```
# Configure Jupyter so figures appear in the notebook
%matplotlib inline
# Configure Jupyter to display the assigned value after an as... | github_jupyter |
# Training baseline model
This notebook shows the implementation of a baseline model for our movie genre classification problem.
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import re
import json
import nltk
from sklearn.model_selection import train_test_split
from sklearn.preprocessing ... | github_jupyter |
# 张量
[](https://gitee.com/mindspore/docs/blob/master/tutorials/source_zh_cn/tensor.ipynb) [](https://obs.dualstack.cn-north-4.myhuaweicloud.com/mindsp... | github_jupyter |
# Speed benchmarks
This is just for having a quick reference of how the speed of running the program scales
```
from __future__ import print_function
import pprint
import subprocess
import sys
sys.path.append('../')
# sys.path.append('/home/heberto/learning/attractor_sequences/benchmarking/')
import numpy as np
impo... | github_jupyter |
<!--- <div style="text-align: center;">
<font size="5">
<b>Data-driven Design and Analyses of Structures and Materials (3dasm)</b>
</font>
</div>
<br>
</br>
<div style="text-align: center;">
<font size="5">
<b>Lecture 1</b>
</font>
</div>
<center>
<img src=docs/tudelft_logo.jpg width=550px>
</center>
... | github_jupyter |
<a href="https://colab.research.google.com/github/CrucifierBladex/cifar10_convnet/blob/main/convnet_cifar10.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
from keras import layers,models
from tensorflow.python.client import device_lib
device_li... | github_jupyter |
#### Information About the Data
u.data -- The full u data set, 100000 ratings by 943 users on 1682 items.
Each user has rated at least 20 movies. Users and items are
numbered consecutively from 1. The data is randomly
ordered. This is a tab separated list of
user id | item id | rating | timestamp.
The time stamps are ... | github_jupyter |
# Purpose
The purpose of this notebook is to train and export the model configuration selected from previous hyperparameter analysis.
The following are the optimal parameters. Other parameter alignments are also stored in order to be able to compare different iterations of the model
**Parameters Selected**:
* **Embe... | github_jupyter |
```
import pandas as pd
train = pd.read_csv("./datasets/labeledTrainData.tsv", header=0, delimiter='\t', quoting=3)
train.head()
train.shape
train.columns.values
train["review"][0]
from bs4 import BeautifulSoup
example1 = BeautifulSoup(train["review"][0])
example1.get_text()
import re
letters_only = re.sub("[^a-zA-Z]",... | github_jupyter |
## Creating schools.csv
1. Install packages
2. Create cities.csv with full state name/ city column to use in getting school information
3. For persisitance creating a schools csv using selenium to get school information from greatschools.org
4. Clean csv for use in schools endpoint
### 1. Import necessary libraries
... | github_jupyter |
<a href="https://colab.research.google.com/github/stephenbeckr/numerical-analysis-class/blob/master/Demos/Ch4_integration.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Numerical Integration (quadrature)
- See also Prof. Brown's [integration not... | github_jupyter |
```
%matplotlib inline
import pandas as pd
import geopandas
import matplotlib.pyplot as plt
```
# Case study - Conflict mapping: mining sites in eastern DR Congo
In this case study, we will explore a dataset on artisanal mining sites located in eastern DR Congo.
**Note**: this tutorial is meant as a hands-on sessio... | github_jupyter |
```
%load_ext autoreload
%autoreload 2
BASE_PATH="/mnt/Archivos/dataset-xray"
from pathlib import Path
from covidframe.tools.load import load_database
base_dir = Path(BASE_PATH)
DEFAULT_DATABASE_NAME_TRAIN = "database_clean_balanced_train.metadata.csv"
DEFAULT_DATABASE_NAME_TEST = "database_clean_balanced_test.metadata... | github_jupyter |
# Milne
```
#All libraries necesary:
%matplotlib inline
import matplotlib
matplotlib.rcParams['figure.figsize'] = (10, 6)
import matplotlib.pyplot as plt
import numpy as np
from math import pi, sin, cos
from copy import deepcopy
from mutils2 import *
import time
# import seaborn
from milne import *
# PARAMETROS:
nline... | github_jupyter |
```
import math
from tensorflow.python.keras.datasets import imdb
from tensorflow.python.keras.preprocessing import sequence
from tensorflow.python.keras import layers
from tensorflow.python.keras.models import Sequential
import numpy as np
from sklearn.calibration import calibration_curve
from sklearn import metric... | github_jupyter |
```
import gtsam
import numpy as np
from gtsam.gtsam import (Cal3_S2, DoglegOptimizer,
GenericProjectionFactorCal3_S2, NonlinearFactorGraph,
Point3, Pose3, Point2, PriorFactorPoint3, PriorFactorPose3,
Rot3, SimpleCamera, Values)
from utils impo... | github_jupyter |
```
# default_exp resimulation
```
# Match resimulation
> Simulating match outcomes based on the xG of individual shots
```
#hide
from nbdev.showdoc import *
#export
import collections
import itertools
import numpy as np
```
Use Poisson-Binomial distribution calculation from https://github.com/tsakim/poibin
It lo... | github_jupyter |
<div align="center">
<font size="6">Solving the Mystery of Chai Time Data Science</font>
</div>
<br>
<div align="center">
<font size="4">A Data Science podcast series by Sanyam Bhutani</font>
</div>
---
<img src="https://miro.medium.com/max/1400/0*ovcHbNV5470zvsH5.jpeg" alt="drawing"/>
---
<div>
<font si... | github_jupyter |
```
import os
import folium
print(folium.__version__)
```
# How to create Popups
## Simple popups
You can define your popup at the feature creation, but you can also overwrite them afterwards:
```
m = folium.Map([45, 0], zoom_start=4)
folium.Marker([45, -30], popup='inline implicit popup').add_to(m)
folium.Circl... | github_jupyter |
# Лабораторная работа 9. ООП.
```
import numpy as np
import matplotlib.pyplot as plt
```
# 1. Создание классов и объектов
В языке программирования Python классы создаются с помощью инструкции `class`, за которой следует произвольное имя класса, после которого ставится двоеточие; далее с новой строки и с отступом реал... | github_jupyter |
# Qcodes example with InstrumentGroup driver
This notebooks explains how to use the `InstrumentGroup` driver.
## About
The goal of the `InstrumentGroup` driver is to combine several instruments as submodules into one instrument. Typically, this is meant to be used with the `DelegateInstrument` driver. An example usag... | github_jupyter |
```
from __future__ import print_function
import sisl
import numpy as np
import matplotlib.pyplot as plt
from functools import partial
%matplotlib inline
```
TBtrans is capable of calculating transport in $N\ge 1$ electrode systems. In this example we will explore a 4-terminal graphene GNR cross-bar (one zGNR, the oth... | github_jupyter |
```
#pip install xlwt openpyxl xlsxwriter xlrd
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import matplotlib as mpl
import seaborn as sns
```
# Loading in Calibration datasets
```
#CO2 only
df_Eguchi_CO2= pd.read_excel('Solubility_Datasets_V1.xlsx', sheet_name='Eguchi_CO2', index_col=0)
d... | github_jupyter |
# Обратные связи в контуре управления
Для рассмотренных в предыдущих лекциях регуляторов требуется оценивать состояние объекта управления. Для построения таких оценок необходимо реализовать обратные связи в контуре управления. На практике для этого используются специальные устройства: датчики.
# Случайные величины
... | github_jupyter |
```
import spotipy
from spotipy.oauth2 import SpotifyOAuth
import pandas as pd
import time
scope = 'user-top-read user-library-read'
sp = spotipy.Spotify(client_credentials_manager=SpotifyOAuth(scope=scope))
sp.user_playlists(sp.current_user()['id'])
results = sp.current_user_top_artists(time_range='short_term', limit=... | github_jupyter |
# Graphics
```
import matplotlib.pyplot as plt
import numpy as np
from PIL import Image
```
## Heat Kernel
```
alpha = 1 / 2
d = 1
K = lambda t, x, y: (4 * np.pi * alpha * t) ** (-d / 2) * np.exp(-(np.abs(x - y) ** 2) / (4 * alpha * t))
t = 1
x = 0
y = np.linspace(-5, 5, 100)
plt.figure(figsize=(10, 2.5))
#plt.title... | github_jupyter |
# Latitude, Longitude for any pixel in a GeoTiff File
How to generate the latitude and longitude for a pixel at any given position in a GeoTiff file.
```
from osgeo import ogr, osr, gdal
# opening the geotiff file
ds = gdal.Open('G:\BTP\Satellite\Data\Test2\LE07_L1GT_147040_20050506_20170116_01_T2\LE07_L1GT_147040_200... | github_jupyter |
```
import os, sys
module_path = os.path.abspath(os.path.join('..'))
sys.path.append(module_path)
import random
from src.loader import *
from src.metrics import Metrics, avg_dicts
from tqdm import tqdm
class Random:
""" Random baseline: probability of 1/(Avg seg length)
that a sentence ends a seg
"""
... | github_jupyter |
# TV Script Generation
In this project, you'll generate your own [Simpsons](https://en.wikipedia.org/wiki/The_Simpsons) TV scripts using RNNs. You'll be using part of the [Simpsons dataset](https://www.kaggle.com/wcukierski/the-simpsons-by-the-data) of scripts from 27 seasons. The Neural Network you'll build will gen... | github_jupyter |
<a href="https://colab.research.google.com/github/tvml/ml2021/blob/main/codici/ae.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
IS_COLAB = ('google.colab' in str(get_ipython()))
if IS_COLAB:
%tensorflow_version 2.x
import tensorflow as tf
fr... | github_jupyter |
```
import tensorflow as tf
from matplotlib import pyplot as plt
%matplotlib inline
plt.style.use('ggplot')
```
## Data
```
n_observations = 10000
xs = np.linspace(-3,3,n_observations)
ys = np.sin(xs) + np.random.uniform(-0.5,0.5,n_observations)
plt.plot(xs,ys, marker='+',alpha=0.4)
```
## Cost
```
sess = tf.Sessio... | github_jupyter |
```
from google.colab import drive
drive.mount('/content/drive', force_remount=True)
cd /content/drive/MyDrive/sop-covid/voice/model_rnn/breath
!unzip ../../data_rnn/data_breath.zip
import numpy as np
import tensorflow as tf
import tensorflow.keras as keras
import matplotlib.pyplot as plt
import pickle
import os
impor... | github_jupyter |
ERROR: type should be string, got "https://www.testdome.com/questions/python/two-sum/14289?questionIds=14288,14289&generatorId=92&type=fromtest&testDifficulty=Easy\n\nWrite a function that, given a list and a target sum, returns zero-based indices of any two distinct elements whose sum is equal to the target sum. If there are no such elements, the function should return (-1, -1).\n\nFor example, `find_two_sum([1, 3, 5, 7, 9], 12)` should return a tuple containing any of the following pairs of indices:\n```\n1 and 4 (3 + 9 = 12)\n2 and 3 (5 + 7 = 12)\n3 and 2 (7 + 5 = 12)\n4 and 1 (9 + 3 = 12)\n```\n\n```\n# Это единственный комментарий который имеет смысл\n# I s\ndef find_index(m,a):\n try:\n return a.index(m)\n except :\n return -1\n \n \ndef find_two_sum(a, s):\n '''\n >>> (3, 5) == find_two_sum([1, 3, 5, 7, 9], 12)\n True\n '''\n if len(a)<2: \n return (-1,-1)\n\n idx = dict( (v,i) for i,v in enumerate(a) )\n\n for i in a:\n m = s - i\n k = idx.get(m,-1)\n if k != -1 :\n return (i,k)\n\n return (-1, -1)\n\n\nprint(find_two_sum([1, 3, 5, 7, 9], 12))\n\n\nif __name__ == '__main__':\n import doctest; doctest.testmod()\n```\n\nhttps://stackoverflow.com/questions/28309430/edit-ipython-cell-in-an-external-editor\n\n\nEdit IPython cell in an external editor\n---\n\nThis is what I came up with. I added 2 shortcuts:\n\n- 'g' to launch gvim with the content of the current cell (you can replace gvim with whatever text editor you like).\n- 'u' to update the content of the current cell with what was saved by gvim.\nSo, when you want to edit the cell with your preferred editor, hit 'g', make the changes you want to the cell, save the file in your editor (and quit), then hit 'u'.\n\nJust execute this cell to enable these features:\n\n```\n%%javascript\n\nIPython.keyboard_manager.command_shortcuts.add_shortcut('g', {\n handler : function (event) {\n \n var input = IPython.notebook.get_selected_cell().get_text();\n \n var cmd = \"f = open('.toto.py', 'w');f.close()\";\n if (input != \"\") {\n cmd = '%%writefile .toto.py\\n' + input;\n }\n IPython.notebook.kernel.execute(cmd);\n //cmd = \"import os;os.system('open -a /Applications/MacVim.app .toto.py')\";\n //cmd = \"!open -a /Applications/MacVim.app .toto.py\";\n cmd = \"!code .toto.py\";\n\n IPython.notebook.kernel.execute(cmd);\n return false;\n }}\n);\n\nIPython.keyboard_manager.command_shortcuts.add_shortcut('u', {\n handler : function (event) {\n function handle_output(msg) {\n var ret = msg.content.text;\n IPython.notebook.get_selected_cell().set_text(ret);\n }\n var callback = {'output': handle_output};\n var cmd = \"f = open('.toto.py', 'r');print(f.read())\";\n IPython.notebook.kernel.execute(cmd, {iopub: callback}, {silent: false});\n return false;\n }}\n);\n# v=getattr(a, 'pop')(1)\ns='print 4 7 '\ncommands={\n 'print':print,\n 'len':len\n }\n\n\ndef exec_string(s):\n global commands\n chunks=s.split()\n func_name=chunks[0] if len(chunks) else 'blbl'\n func=commands.get(func_name,None)\n \n params=[int(x) for x in chunks[1:]]\n if func:\n func(*params)\n\nexec_string(s)\n```\n\n# Symmetric Difference\n\nhttps://www.hackerrank.com/challenges/symmetric-difference/problem\n\n#### Task \nGiven sets of integers, and , print their symmetric difference in ascending order. The term symmetric difference indicates those values that exist in either or but do not exist in both.\n\n#### Input Format\n\nThe first line of input contains an integer, . \nThe second line contains space-separated integers. \nThe third line contains an integer, . \nThe fourth line contains space-separated integers.\n\n##### Output Format\n\nOutput the symmetric difference integers in ascending order, one per line.\n\n#### Sample Input\n````\n4\n2 4 5 9\n4\n2 4 11 12\n````\n##### Sample Output\n````\n5\n9\n11\n12\n````\n\n```\nM = int(input())\nm =set((map(int,input().split())))\nN = int(input())\nn =set((map(int,input().split())))\nm ^ n\nS='add 5 6'\nmethod, *args = S.split()\nprint(method)\nprint(*map(int,args))\nmethod,(*map(int,args))\n\n# methods\n# (*map(int,args))\n\n# command='add'.split()\n# method, args = command[0], list(map(int,command[1:]))\n# method, args\nfor _ in range(2):\n met, *args = input().split()\n print(met, args)\n try:\n pass\n\n# methods[met](*list(map(int,args)))\n except:\n pass\nclass Stack:\n def __init__(self):\n self.data = []\n\n def is_empty(self):\n return self.data == []\n\n def size(self):\n return len(self.data)\n\n def push(self, val):\n self.data.append(val)\n\n def clear(self):\n self.data.clear()\n \n def pop(self):\n return self.data.pop()\n\n def __repr__(self):\n return \"Stack(\"+str(self.data)+\")\"\ndef sum_list(ls):\n if len(ls)==0:\n return 0\n elif len(ls)==1:\n return ls[0]\n else:\n return ls[0] + sum_list(ls[1:])\n\ndef max_list(ls):\n print(ls)\n if len(ls)==0:\n return None\n elif len(ls)==1:\n return ls[0]\n else:\n\n m = max_list(ls[1:])\n return ls[0] if ls[0]>m else m\n \ndef reverse_list(ls):\n if len(ls)<2:\n return ls\n \n return reverse_list(ls[1:])+ls[0:1]\n\n\ndef is_ana(s=''):\n if len(s)<2:\n return True\n return s[0]==s[-1] and is_ana(s[1:len(s)-1])\n \n \n \nprint(is_ana(\"abc\"))\nimport turtle\n\nmyTurtle = turtle.Turtle()\nmyWin = turtle.Screen()\n\ndef drawSpiral(myTurtle, lineLen):\n if lineLen > 0:\n myTurtle.forward(lineLen)\n myTurtle.right(90)\n drawSpiral(myTurtle,lineLen-5)\n\ndrawSpiral(myTurtle,100)\n# myWin.exitonclick()\nt.forward(100)\nfrom itertools import combinations_with_replacement\nlist(combinations_with_replacement([1,1,3,3,3],2))\nhash((1,2))\n# 4 \n# a a c d\n# 2\n\n\nfrom itertools import combinations\n\n# N=int(input())\n# s=input().split()\n# k=int(input())\n\ns='a a c d'.split()\nk=2\n\n\ncombs=list(combinations(s,k))\n\n\nprint('{:.4f}'.format(len([x for x in combs if 'a' in x])/len(combs)))\n\n# ------------------------------------------\n\nimport random\n\nnum_trials=10000\nnum_found=0\n\nfor i in range(num_trials):\n if 'a' in random.sample(s,k):\n num_found+=1\n \n\n\nprint('{:.4f}'.format(num_found/num_trials))\ndir(5)\n```\n\n" | github_jupyter |
```
!pip install unidecode googletrans
!pip install squarify
import re
import time
import tweepy
import folium
import squarify
import warnings
import collections
import numpy as np
import pandas as pd
from PIL import Image
from folium import plugins
from datetime import datetime
from textblob import TextBlob
import ma... | github_jupyter |
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import math
# Define functions
# Scale values
def scale(arr):
return (np.array(arr) - np.mean(arr)) / np.std(arr)
# Find the slope of the best fitting line
def fit_slope(x, y):
return (np.mean(x) * np.mean(y) - np.mean(x * y)) / (np.me... | github_jupyter |
# Document Classification & Clustering - Lecture
What could we do with the document-term-matrices (dtm[s]) created in the previous notebook? We could visualize them or train an algorithm to do some specific task. We have covered both classification and clustering before, so we won't focus on the particulars of algorit... | github_jupyter |
```
a = 'ok'
b = 'test'
print(a+b)
print(a*2)
name = 'Bob'
print(f'Hello, {name}')
greeting = 'Hello, {}'
with_name = greeting.format(name)
print(with_name)
size = input('Enter the size of your house: ')
integer = int(size)
floating = float(size)
print(integer, floating)
square_meters = integer / 10.8
print(f'{integer}... | github_jupyter |
# Kestrel+Model
### A [Bangkit 2021](https://grow.google/intl/id_id/bangkit/) Capstone Project
Kestrel is a TensorFlow powered American Sign Language translator Android app that will make it easier for anyone to seamlessly communicate with people who have vision or hearing impairments. The Kestrel model builds on the ... | github_jupyter |
## Libraries
```
import numpy as np
import pandas as pd
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
from sklearn.cluster import KMeans
import pickle
from sklearn.linear_model import LogisticRegression
import matplotlib.pyplot as plt
import matplotlib.axes as axs
import s... | github_jupyter |
# Lambda School Data Science Module 141
## Statistics, Probability, and Inference
## Prepare - examine what's available in SciPy
As we delve into statistics, we'll be using more libraries - in particular the [stats package from SciPy](https://docs.scipy.org/doc/scipy/reference/tutorial/stats.html).
```
from scipy im... | github_jupyter |
# Musicians- Medium
```
# Prerequesites
from pyhive import hive
%load_ext sql
%sql hive://cloudera@quickstart.cloudera:10000/sqlzoo
%config SqlMagic.displaylimit = 20
```
## 6.
**List the names, dates of birth and the instrument played of living musicians who play a instrument which Theo also plays.**
```
%%sql
WITH... | github_jupyter |
```
project = 'saga-trafikkdata-prod-pz8l'
use_colab_auth = True
# Legg inn ditt eget prosjekt her, f.eks. 'saga-olanor-playground-ab12'
bq_job_project = ''
if (use_colab_auth):
from google.colab import auth
auth.authenticate_user()
print('Authenticated')
import warnings
from google.cloud import bigquery
warnin... | github_jupyter |
```
import sys
print(f'Interpreter dir: {sys.executable}')
import os
import warnings
warnings.filterwarnings("ignore")
if os.path.basename(os.getcwd()) == 'notebooks':
os.chdir('../')
print(f'Working dir: {os.getcwd()}')
%load_ext autoreload
%autoreload 2
import xgboost as xgb
import lightgbm as lgb
import pan... | github_jupyter |
# PyTorch: Tabular Classify Binary

```
import torch
import torch.nn as nn
from torch import optim
import torchmetrics
from sklearn.preprocessing import LabelBinarizer, StandardScaler
import aiqc
from aiqc import datum
```
---
## Example Data
Reference [Example Datasets](example_data... | github_jupyter |
## Import
```
import numpy as np
import functions as fc
from timeit import default_timer as time
from fatiando.gravmag import polyprism
from fatiando import mesher, gridder
from fatiando.gravmag import prism
from fatiando.constants import G, SI2MGAL
from scipy.sparse import diags
from matplotlib import pyplot as plt
... | github_jupyter |
<div class="contentcontainer med left" style="margin-left: -50px;">
<dl class="dl-horizontal">
<dt>Title</dt> <dd> Scatter Element</dd>
<dt>Dependencies</dt> <dd>Matplotlib</dd>
<dt>Backends</dt>
<dd><a href='./Scatter.ipynb'>Matplotlib</a></dd>
<dd><a href='../bokeh/Scatter.ipynb'>Bokeh</a></dd>
<dd>... | github_jupyter |
```
import numpy as np
import scipy
import matplotlib.pyplot as plt
%matplotlib inline
plt.style.use('jf')
from jftools import fedvr
# 5 points (element boundaries) gives 4 elements
# very low order to have only a few basis functions for plot
# g = fedvr_grid(4,np.linspace(0,8,5))
g = fedvr.fedvr_grid(4,np.array([0,2,3... | github_jupyter |
# Steps to Tackle a Time Series Problem (with Codes in Python)
Note: These are just the codes from article
## Loading and Handling TS in Pandas
```
import pandas as pd
import numpy as np
import matplotlib.pylab as plt
%matplotlib inline
from matplotlib.pylab import rcParams
rcParams['figure.figsize'] = 15, 6
#Note: a... | github_jupyter |
# Gameplan:
1. Set up data
2. Create subset for Excel
3. Make a prediction w Dot Product
4. Analyze results
5. Try a neural net.
```
from theano.sandbox import cuda
%matplotlib inline
import utils; reload(utils)
from utils import *
from __future__ import division, print_function
path = "data/ml-small/ml-latest-small/... | github_jupyter |
```
### Pytorch geometry (グラフニューラルネットワークライブラリ) のインストール(only first time)
!pip install -q torch-scatter -f https://pytorch-geometric.com/whl/torch-1.9.0+cu102.html
!pip install -q torch-sparse -f https://pytorch-geometric.com/whl/torch-1.9.0+cu102.html
!pip install -q git+https://github.com/rusty1s/pytorch_geometric.git
... | github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import sys
sys.path.append('../../pyutils')
import metrics
import utils
```
# Introduction
In unsupervised learing, one has a set of $N$ observations $x_i \in \mathbb{R}^p$, having joint density $P(X)$.
The goal is to infer properties of th... | github_jupyter |
# Wie Sie dieses Notebook nutzen:
- Führen Sie diesen Code Zelle für Zelle aus.
- Um die Variableninhalte zu beobachten, nutzen Sie in Jupyter-Classic den "Variable Inspektor". Falls Sie dieses Notebook in Jupyter-Lab verwenden, nutzen Sie hierfür den eingebauten Debugger.
- Wenn Sie "Code Tutor" zur Visualisierung des... | github_jupyter |
# The Assemble Module
The `assemble` module of the `repytah` package finds and forms essential structure components. These components are the smallest building blocks that form the basis for every repeat in the song. The functions in this module ensure that each time step of a song is contained in at most one of the s... | github_jupyter |
# Linear algebra in Python with NumPy
In this lab, you will have the opportunity to remember some basic concepts about linear algebra and how to use them in Python.
Numpy is one of the most used libraries in Python for arrays manipulation. It adds to Python a set of functions that allows us to operate on large multid... | github_jupyter |
# Hinge Loss
In this project you will be implementing linear classifiers beginning with the Perceptron algorithm. You will begin by writing your loss function, a hinge-loss function. For this function you are given the parameters of your model θ and θ0
Additionally, you are given a feature matrix in which the rows ar... | github_jupyter |
(Feedforward)=
# Chapter 8 -- Feedforward
Let's take a look at how feedforward is processed in a three layers neural net.
<img src="images/feedForward.PNG" width="500">
Figure 8.1
From the figure 8.1 above, we know that the two input values for the first and the second neuron in the hidden layer are
$$
h_1^{(1)} = ... | github_jupyter |
In this lab, we will optimize the weather simulation application written in Fortran (if you prefer to use C++, click [this link](../../C/jupyter_notebook/profiling-c.ipynb)).
Let's execute the cell below to display information about the GPUs running on the server by running the pgaccelinfo command, which ships with t... | github_jupyter |
```
import matplotlib
import matplotlib.pyplot as plt
import os
import random
import io
import imageio
import glob
import scipy.misc
import numpy as np
from six import BytesIO
from PIL import Image, ImageDraw, ImageFont
from IPython.display import display, Javascript
from IPython.display import Image as IPyImage
impo... | github_jupyter |
This script generates a zone plate pattern (based on partial filling) given the material, energy, grid size and number of zones as input
```
import numpy as np
import matplotlib.pyplot as plt
from numba import njit
from joblib import Parallel, delayed
from tqdm import tqdm, trange
import urllib,os,pickle
from os.path... | github_jupyter |
```
%matplotlib nbagg
import os
os.environ["PYOPENCL_COMPILER_OUTPUT"]="1"
import numpy
import fabio
import pyopencl
from pyopencl import array as cla
from matplotlib.pyplot import subplots
ctx = pyopencl.create_some_context(interactive=True)
queue = pyopencl.CommandQueue(ctx, properties=pyopencl.command_queue_properti... | github_jupyter |
```
# default_exp downloaders
#export
import requests
import pathspec
import time
from pathlib import Path, PurePosixPath
from tightai.lookup import Lookup
from tightai.conf import CLI_ENDPOINT
#hide
test = False
if test:
CLI_ENDPOINT = "http://cli.desalsa.io:8000"
#export
class DownloadVersion(Lookup):
path =... | github_jupyter |
# Import Required Modules
```
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import re
from IPython.display import HTML
%matplotlib inline
HTML('''<script>
code_show=true;
function code_toggle() {
if (code_show){
$('div.input').hide();
} else {
$('div.input'... | github_jupyter |

[](https://colab.research.google.com/github/JohnSnowLabs/spark-nlp-workshop/blob/master/tutorials/Certification_Trainings/Healthcare/Spark%20v2.7.6%20Notebooks/21.Gender_Classi... | github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.