text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
# Pandas Data Analysis
## Imports
Import `pandas` and `numpy` into the notebook.
```
import pandas as pd
import numpy as np
```
## Loading data
```
df = pd.read_csv('./data.csv')
df
```
The [`head`](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.head.html) method of our data frame returns... | github_jupyter |
# DefinedAEpTandZ0 media example
```
%load_ext autoreload
%autoreload 2
import skrf as rf
import skrf.mathFunctions as mf
import numpy as np
from numpy import real, log, log10, sum, absolute, pi, sqrt
import matplotlib.pyplot as plt
from matplotlib.ticker import AutoMinorLocator
from scipy.optimize import minimize
rf... | github_jupyter |
# QCodes example with Mercury iPS
## Initial instantiation/connection
```
from qcodes.instrument_drivers.oxford.MercuryiPS_VISA import MercuryiPS
from time import sleep
# Note that the MercuryiPS_VISA is a VISA instrument using
# a socket connection. The VISA resource name therefore
# contains the port number and th... | github_jupyter |
#### Define your project and region below. If you are not authenticated to GCP, do it by oncommenting the line below the definitions.
```
PROJECT_ID = "SOME_PROJECT"
REGION = "YOUR_REGION" #though us-central is cheaper
PIPELINE_ROOT = "gs://SOME_BUCKET/SOME_FOLDER"
#!gcloud auth login
```
#### Imports
Our imports:
... | github_jupyter |
# Knowledge Graph Embeddings
Word embeddings aim at capturing the meaning of words based on very large corpora; however, there are decades of experience and approaches that have tried to capture this meaning by structuring knowledge into semantic nets, ontologies and graphs.
| | Neural | Symbolic ... | github_jupyter |
.. _nb_repair:
## Repair Operator
The repair operator is mostly problem dependent. Most commonly it is used to make sure the algorithm is only searching in the feasible space. It is applied after the offsprings have been reproduced. In the following, we are using the knapsack problem to demonstrate the repair operato... | github_jupyter |
# Creation of the Alternative Classification for Modeling
In this notebook, we create a csv file containing the alternative classification of crimes, in 7 categories.
<br>
We also clean and segment the data according to time, localization and neighborhoods.
# Cleaning of the Data from clean_data.csv
```
data = pd.r... | github_jupyter |
Training and Testing Data
=====================================
To evaluate how well our supervised models generalize, we can split our data into a training and a test set:
<img src="../images/train_test_split.svg" width="80%">
```
from sklearn.datasets import load_iris
from sklearn.neighbors import KNeighborsClass... | github_jupyter |
# SVM classification/SMOTE oversampling for an imbalanced data set
Date created: Oct 14, 2016
Last modified: Nov 16, 2016
Tags: SVM, SMOTE, ROC/AUC, oversampling, imbalanced data set, semiconductor data
About: Rebalance imbalanced semicondutor manufacturing dataset by oversampling the minority class using SMOT... | github_jupyter |
```
import pandas as pd
import numpy as np
import catboost as cat
def reduce_mem_usage(df):
""" iterate through all the columns of a dataframe and modify the data type
to reduce memory usage.
"""
start_mem = df.memory_usage().sum()
for col in df.columns:
col_type = df[col].... | github_jupyter |
# Find when a piece of text appears in an archived web page
This notebook helps you find when a particular piece of text appears in, or disappears from, a web page. Using Memento Timemaps, it gets a list of available captures from the selected web archive. It then searches each capture for the desired text, displaying... | github_jupyter |
Objectives
- Order the rows of a table using a chosen column
- Convert to long format to plot multiple columns at the same time
- Switch between short/long table format
Content to cover
- sort_values
- pivot, pivot_table
- melt
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
%matplotlib i... | github_jupyter |
# Morisita-Horn similarity calculation
```
from __future__ import print_function
from collections import Counter
from datetime import datetime
import itertools
import multiprocessing as mp
import os
import subprocess as sp
import sys
import tempfile
import time
import numpy as np
import pandas as pd
from abutils.ut... | github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
sns.set_style('whitegrid')
%matplotlib inline
default = pd.read_csv('../data/credit_card_default.csv')
default.rename(columns=lambda x: x.lower(), inplace=True)
default.rename(columns={'pay_0':'pay_1','default payment next ... | github_jupyter |
```
import matplotlib.pyplot as plt
import numpy as np
from qiskit import QuantumCircuit, Aer, transpile, assemble
from qiskit.visualization import plot_histogram
from math import gcd
from numpy.random import randint
import pandas as pd
from fractions import Fraction
print("Imports Successful")
def c_amod15(a, power):
... | github_jupyter |
# Import library
```
import os, csv
import pandas as pd
from os import path
import plotly.graph_objs as go
from plotly.offline import plot, init_notebook_mode, iplot
%matplotlib inline
```
# Configure directory
```
userhome = os.path.expanduser('~')
txt_file = open(userhome + r"/DifferentDiffAlgorithms/SZZ/code_doc... | github_jupyter |
```
from PIL import Image
import numpy as np
import matplotlib.pyplot as plt
#load operation
img = Image.open('mouse.png').convert('RGB')
np.shape(img)
# resize opeartion
img = img.resize((14, 14))
plt.imshow(img)
img.size
# opem my write file(should be empty)
f = open('mouse_array.c', 'w')
# fill loop
for y in range(n... | github_jupyter |
## Haensel AMS Homework
#### Paul Teehan
#### June 6, 2016
We are asked to solve the following task:
* Two listings of product-session pairs are provided; one for search results, and one for viewings.
* For each product that was viewed, find which three products are most often viewed or displayed in the same session.
... | github_jupyter |
```
import cirq
import numpy as np
import tensorflow as tf
import tensorflow_quantum as tfq
import pandas as pd
from qite import QITE
from qbm import QBM
from circuit import build_ansatz, initialize_ansatz_symbols
from problem import build_ising_model_hamiltonian
from hamiltonian import Hamiltonian
from utils import e... | github_jupyter |
# Exercise: putting everything together
In this you will write code for a model that learns to classify mnist digits. You will use tensorflow, tracking training progress with matplotlib.
For each sub-exercise, you have seen an example solution for it in one of the colabs leading up to this one.
```
from __future__ i... | github_jupyter |
```
import cv2
import numpy as np
import math
import cv2
import random
import time
import sys
import operator
import os
from numpy import zeros, newaxis
import re
import sys
import matplotlib.pyplot as plt
import glob
import skimage
import skimage.io
import scipy.io as scp
from sklearn.utils import shuffle
from __futu... | github_jupyter |
```
%matplotlib inline
import sys
import numpy as np
import numpy.random as rnd
import time
import GPflow
import tensorflow as tf
import matplotlib
import matplotlib.pyplot as plt
plt.style.use('ggplot')
M = 50
```
# Create a dataset and initialise model
```
def func(x):
return np.sin(x * 3*3.14) + 0.3*np.cos(x *... | github_jupyter |
# Train and deploy on Kubeflow from Notebooks
This notebook introduces you to using Kubeflow Fairing to train and deploy a model to Kubeflow on Google Kubernetes Engine (GKE), and Google Cloud ML Engine. This notebook demonstrate how to:
* Train an XGBoost model in a local notebook,
* Use Kubeflow Fairing to train a... | github_jupyter |
# Run fitsverify
```
import os
import sys
import re
import shutil
import subprocess as sp
from configparser import ConfigParser
from random import choice
specprod = 'everest'
specprod_path = os.path.join(os.environ['DESI_SPECTRO_REDUX'], specprod)
```
## Create input file
```
fits_files = os.path.join(os.environ['CS... | github_jupyter |
```
# Putting the initialisation at the top now!
import veneer
%matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
v = veneer.Veneer(port=9876)
```
# Session 6 - Model Setup and Reconfiguration
This session covers functionality in Veneer and veneer-py for making larger changes t... | github_jupyter |
```
import requests
import json
#res = requests.get("https://api.airtable.com/v0/appNcYtL8fFZa1STA/iris?api_key=keyshdNC8CZdj1xgo")
Base_ID = 'appNcYtL8fFZa1STA'
Table_name = 'iris'
# url格式: API URL/v版本/Base_ID/Table_Name
url = 'https://api.airtable.com/v0/{0}/{1}'.format(Base_ID, Table_name);
API_KEY = {'api_key': ... | github_jupyter |
# Mask R-CNN - Compare ouptuts from Heatmap layer and FCN layer
```
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:95% !important; }</style>"))
%matplotlib inline
%load_ext autoreload
%autoreload 2
import sys
sys.path.append('../')
import tensorflow as tf
import keras.backend as... | github_jupyter |
# [Hashformers](https://github.com/ruanchaves/hashformers)
Hashformers is a framework for hashtag segmentation with transformers. For more information, please check the [GitHub repository](https://github.com/ruanchaves/hashformers).
# Installation
The steps below will install the hashformers framework on Google Cola... | github_jupyter |
In the previous tutorial, we introduced you to the basics of binary finite fields, but didn't really dive into the math or the implementation. In this tutorial, we're going to go deeper and actually walk through the mathematics of how binary fields actually work.
# What is “binary finite fields”?
Finite fields of ord... | github_jupyter |
A quick look at GAMA bulge and disk colours in multi-band GALAPAGOS fits versus single-band GALAPAGOS and SIGMA fits.
Pretty plots at the bottom.
```
%matplotlib inline
from matplotlib import pyplot as plt
# better-looking plots
plt.rcParams['font.family'] = 'serif'
plt.rcParams['figure.figsize'] = (10.0*1.3, 8*1.3)
... | github_jupyter |
```
%matplotlib inline
```
PyTorch是什么?
================
基于Python的科学计算包,服务于以下两种场景:
- 作为NumPy的替代品,可以使用GPU的强大计算能力
- 提供最大的灵活性和高速的深度学习研究平台
开始
---------------
Tensors(张量)
^^^^^^^
Tensors与Numpy中的 ndarrays类似,但是在PyTorch中
Tensors 可以使用GPU进行计算.
```
from __future__ import print_function
import torch
```
创建一个 5x3 矩阵... | github_jupyter |
```
import pandas as pd
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
import seaborn as sns
from scipy import stats
# 노트북 안에 그래프를 그리기 위해
%matplotlib inline
# 그래프에서 격자로 숫자 범위가 눈에 잘 띄도록 ggplot 스타일을 사용
plt.style.use('ggplot')
# 그래프에서 마이너스 폰트 깨지는 문제에 대한 대처
mpl.rcParams['axes.unicode_minus']... | github_jupyter |
<i>Copyright (c) Microsoft Corporation. All rights reserved.</i>
<i>Licensed under the MIT License.</i>
# Spark Collaborative Filtering (ALS) Deep Dive
Spark MLlib provides a collaborative filtering algorithm that can be used for training a matrix factorization model, which predicts explicit or implicit ratings of u... | github_jupyter |
```
%run "Retropy_framework.ipynb"
conf_cache_disk = True
conf_cache_memory = True
t = get("FDN").s
https://docs.scipy.org/doc/scipy/reference/tutorial/interpolate.html
https://docs.scipy.org/doc/scipy/reference/interpolate.html
https://docs.scipy.org/doc/scipy-0.18.1/reference/generated/scipy.interpolate.UnivariateSpl... | github_jupyter |
# Sparse Sinkhorn Transformer (PyTorch/GPU) (Ver 1.0)
***
### Credit for the PyTorch Reformer implementation goes out to @lucidrains of GitHub:
https://github.com/lucidrains/sinkhorn-transformer
***
This is a work in progress so please check back for updates.
***
Project Los Angeles
Tegridy Code 2021
# Setup E... | github_jupyter |
```
import torch
from torch import nn, optim
from neurodiffeq import diff
from neurodiffeq.networks import FCNN
from neurodiffeq.temporal import generator_2dspatial_rectangle, generator_2dspatial_segment, generator_temporal
from neurodiffeq.temporal import FirstOrderInitialCondition, BoundaryCondition
from neurodiffeq.... | github_jupyter |
# Creating your own dataset from Google Images
*by: Francisco Ingham and Jeremy Howard. Inspired by [Adrian Rosebrock](https://www.pyimagesearch.com/2017/12/04/how-to-create-a-deep-learning-dataset-using-google-images/)*
In this tutorial we will see how to easily create an image dataset through Google Images. **Note*... | github_jupyter |
# Matplotlib Bars
## Creating Bars
With Pyplot, you can use the `bar()` function to draw bar graphs:
```
# Draw 4 bars:
import matplotlib.pyplot as plt
import numpy as np
x = np.array(["A", "B", "C", "D"])
y = np.array([4, 9, 1, 11])
plt.bar(x,y)
plt.show()
```
The `bar()` function takes arguments that describes ... | github_jupyter |
Query NASA/Ads from python
https://github.com/adsabs/adsabs-dev-api/blob/master/README.md
```
from astroquery.ned import Ned
from astroquery.nasa_ads import ADS
ADS.TOKEN = open('ADS_DEV_KEY','r').read()
token = open('ADS_DEV_KEY','r').read()
import requests
import urllib
import json
from pnlf.constants import tab10
... | github_jupyter |
# NLP Feature Engineering
## Feature Creation
```
# Read in the text data
import pandas as pd
data = pd.read_csv("./data/SMSSpamCollection.tsv", sep='\t')
data.columns = ['label', 'body_text']
```
### Create feature for text message length
```
data['body_len'] = data['body_text'].apply(lambda x: len(x) - x.count("... | github_jupyter |
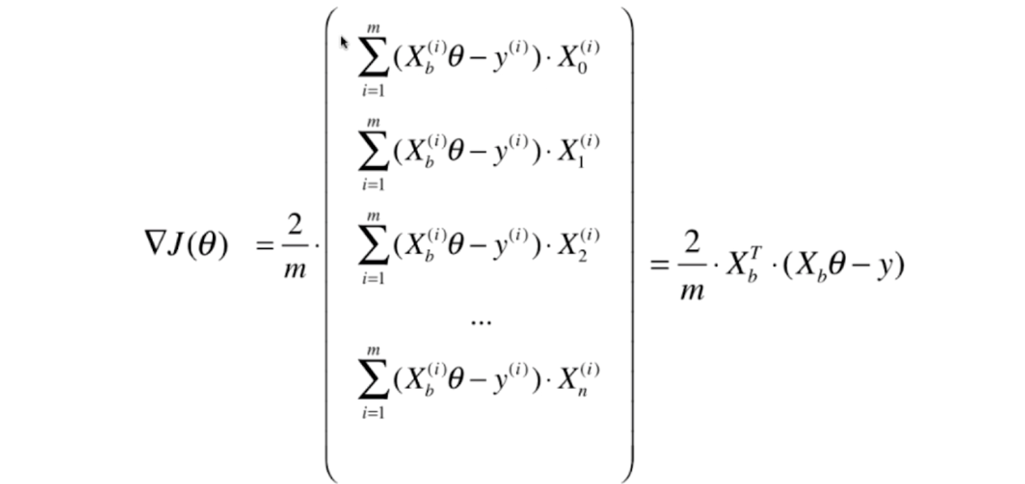
# Stochastic Gradient Descent
- 上节梯度下降法如图所示
[](https://imgchr.com/i/8mATJK)
- 我们每次都把所有的梯度算出来,称为**批量梯度下降法**
- 但是这样在样本容量很大时,也是比较耗时的,解决方法是**随机梯度下降法**
[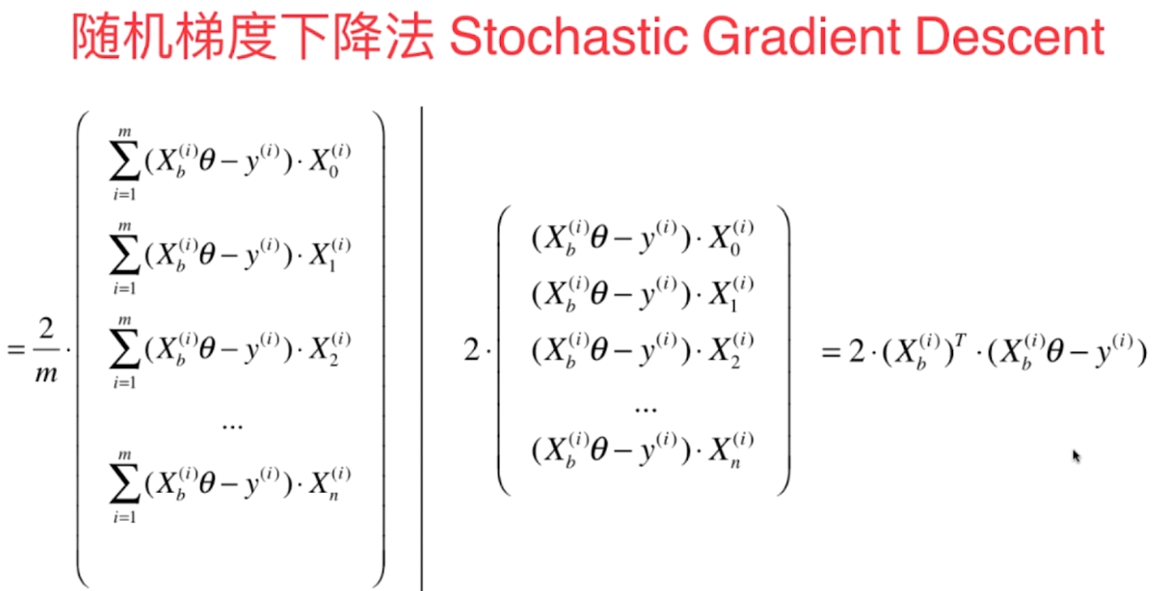](https://imgchr.com/i/8mALsH)
- 我们随机的取一个 $i$ ,然后用这个 $i... | github_jupyter |
```
pip install pandas
pip install gym
pip install matplotlib
pip install tensorflow
import numpy as np
import pandas as pd
import inspect
import random
import gym
import sys
import tensorflow as tf
import tensorflow.keras.layers as kl
import tensorflow.keras.losses as kls
import tensorflow.keras.optimizers as ko
impor... | github_jupyter |
```
import os
import json
import pathlib
import random
import numpy as np
import matplotlib.pyplot as plt
import imageio
from skimage import transform
from IPython import display
try:
os.mkdir('data')
except FileExistsError:
pass
import tensorflow as tf
# Makes it so any changes in pymedphys is automatically... | github_jupyter |
# Siamese Convolutional Neural Network
```
from model import siamese_CNN
import os
os.environ['TF_CPP_MIN_LOG_LEVEL'] = '2'
import pickle
import numpy as np
from pandas import DataFrame
import tensorflow as tf
import keras.backend as K
# model imports
from keras.models import Sequential, Model, Input
from keras.lay... | github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
from skhep.dataset.numpydataset import *
import uproot
from skhep.dataset.selection import Selection
import ROOT
from Utilities.utilities import destruct_objects
from Utilities.RooFit import RooDataset, RemoveEmptyBins
from PyLHCb.Root.RooFitUtils import ResidualPl... | github_jupyter |
# Example 10 A: Inverted Pendulum with Wall
```
import numpy as np
import scipy.linalg as spa
import pypolycontain as pp
import pydrake.solvers.mathematicalprogram as MP
import pydrake.solvers.gurobi as Gurobi_drake
# use Gurobi solver
global gurobi_solver, license
gurobi_solver=Gurobi_drake.GurobiSolver()
license = g... | github_jupyter |
# ETHZ: 227-0966-00L
# Quantitative Big Imaging
# May 2, 2018
## Statistics and Reproducibility
```
%load_ext autoreload
%autoreload 2
import seaborn as sns
import matplotlib.pyplot as plt
plt.rcParams["figure.figsize"] = (8, 8)
plt.rcParams["figure.dpi"] = 150
plt.rcParams["font.size"] = 14
plt.rcParams['font.family... | github_jupyter |
## TODO: Convert to Python
## Setup Connection to Kafka
```
import org.apache.spark.sql.functions.get_json_object
import org.apache.spark.sql.functions.json_tuple
streamingInputDF = \
spark.readStream \
.format("kafka") \
.option("kafka.bootstrap.servers", "<server:ip>") \
.option("subscribe", "topic... | github_jupyter |
## WORD2VEC
```
import collections
import math
import os
import random
import zipfile
import numpy as np
from six.moves import urllib
from six.moves import xrange # pylint: disable=redefined-builtin
from sklearn.manifold import TSNE
import matplotlib.pyplot as plt
import tensorflow as tf
%matplotlib inline
print ("... | github_jupyter |
# End-to-end demo of the ``stadv`` package
We use a small CNN pre-trained on MNIST and try and fool the network using *Spatially Transformed Adversarial Examples* (stAdv).
### Import the relevant libraries
```
%matplotlib inline
from __future__ import absolute_import
from __future__ import division
from __future__ i... | github_jupyter |
# Setup
```
from math import floor, ceil
from multiprocessing import Pool, cpu_count
from pathlib import Path
from python_speech_features import logfbank
from python_speech_features import mfcc
from scipy.io import wavfile
from time import time
import glob
import hashlib
import numpy as np
import os
import pickle
impo... | github_jupyter |
```
import numpy as np
import itertools
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.tree import DecisionTreeClassifier, export_graphviz
from sklearn import preprocessing
from sklearn.model_selection import train_test_split
from six import StringIO
import pydotplus#
import matplotlib.image as mpimg
... | github_jupyter |
```
# default_exp utils
```
# utils
> Provides different util functions
```
#export
import json
from copy import deepcopy
import numpy as np
from PIL import Image
from icevision.core.mask import EncodedRLEs, MaskArray
from pycocotools import mask as mask_utils
```
## Test data setup
```
import icedata
from icevis... | github_jupyter |
```
import tensorflow as tf
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
#building all kinds of evaluating parameters
from sklearn.model_selection import train_test_split
from sklearn.metrics import classification_report, accuracy_score
from sklearn.metrics impor... | github_jupyter |
<hr style="height:2px;">
# Demo: Neural network training for joint denoising and surface projection of *Drosophila melanogaster* wing
This notebook demonstrates training a CARE model for a 3D → 2D denoising+projection task, assuming that training data was already generated via [1_datagen.ipynb](1_datagen.ipynb) and h... | github_jupyter |
# ReGraph tutorial (NetworkX backend)
## Part 1: Rewriting simple graph with attributes
This notebook consists of simple examples of usage of the ReGraph library
```
from regraph import NXGraph, Rule
from regraph import plot_graph, plot_instance, plot_rule
%matplotlib inline
```
### 1. Creating and modifying a gra... | github_jupyter |
###### Reference:
https://finthon.com/learn-cnn-two-tfrecord-read-data/
https://finthon.com/learn-cnn-three-resnet-prediction/
# 匯入圖片資料並輸出成tfrecord檔案
```
import os
from PIL import Image
import tensorflow as tf
'''
設置路徑
# 將需分類之圖片目錄放置Working Directory於之下,Folder以Int作為命名
'''
# 图片路径,两组标签都在该目录下
cwd = r"./OM/"
# tf... | github_jupyter |
# KIC 9651065
```
%run setup.py
t, y = np.loadtxt('../lc/9651065_lc.txt', usecols=(0,1)).T
ms = Maelstrom(t, y, max_peaks=5, fmin=5, fmax=48)
ms.first_look()
period_guess = 300
a_guess = 200
time, flux = ms.time, ms.flux
freq = ms.freq
weights = ms.get_weights(norm=False)
pg = ms.period_search()
periods = np.linspace... | github_jupyter |
Zipline Beginner Tutorial
=========================
Basics
------
Zipline is an open-source algorithmic trading simulator written in Python.
The source can be found at: https://github.com/quantopian/zipline
Some benefits include:
* Realistic: slippage, transaction costs, order delays.
* Stream-based: Process each ... | github_jupyter |
# Derived Fields and Profiles
One of the most powerful features in yt is the ability to create derived fields that act and look exactly like fields that exist on disk. This means that they will be generated on demand and can be used anywhere a field that exists on disk would be used. Additionally, you can create the... | github_jupyter |
# One time pad
In the previous lesson we performed an attack over the Monoalphabetic cipher where the attacker (Charlie) only knew that Alice and Bob were communicating in english and that they were using this concrete cipher. Therefore the ciphertext is leaking information. Can we find a cipher whose ciphertext doesn... | github_jupyter |
# Anna KaRNNa
In this notebook, I'll build a character-wise RNN trained on Anna Karenina, one of my all-time favorite books. It'll be able to generate new text based on the text from the book.
This network is based off of Andrej Karpathy's [post on RNNs](http://karpathy.github.io/2015/05/21/rnn-effectiveness/) and [i... | github_jupyter |
# Circuit Translation
In this notebook we will introduce a tool of `sqwalk` that is useful to decompose (or translate) an unitary transormation (in our case the one generated by the walker's Hamiltonian) into a series of gates that can be simulated or even run on quantum hardware. The decomposition method is based on ... | github_jupyter |
# Analysis of Consumer Healthcare Costs
>Project submission for Applied Statistics course as a part of the PGP-AIML programme
***
#### Author:
>Abhinav Kimothi
#### Project Description:
>With the rising healthcare costs, it becomes imperative for a medical insurance provider to carefully analyze the costs viz-a-viz ... | github_jupyter |
# Preparing for Your Proposal
## Which client/dataset did you select and why?
Client 3: SportsStats (Olympics Dataset - 120 years of data)
SportsStats is a sports analysis firm partnering with local news and elite personal trainers to provide “interesting” insights to help their partners. Insights could be pattern... | github_jupyter |
<a href="https://colab.research.google.com/github/kentokura/ox_2x2_retrograde_analysis/blob/main/ox2x2/makeAllState.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
ノードの3状態:
- 未発見
unsolved, solvedのいずれにもnext_nodeが存在しない
- 未訪問
unsolvedに存在する
... | github_jupyter |
# Boston Housing Prices Dataset
## Contents
0. [Introduction](#intro)
1. [Pre-processing and Splitting Data](#split)
2. [Models for median price predictions](#model)
3. [Stacked model](#stack)
## Introduction <a class="anchor" id="intro"></a>
This notebook illustrates the use of the `Stacker` to conveniently stack... | github_jupyter |
## Model - Infinite DPM - Chinese Restaurant Mixture Model (CRPMM)
#### Dirichlet mixture model where number of clusters is learned.
ref = reference sequence
$N$ = number of reads
$K$ = number of clusters/components
$L$ = genome length (number of positions)
alphabet = {A, C, G, T, -}
no-mutation rate: $\ga... | github_jupyter |
```
import pandas as pd
import datetime
import matplotlib.pyplot as plt
all_o3_df = pd.read_csv("./all_years_o3.csv")
#turn date column elements into datetime objects
all_o3_df["Date"] = pd.to_datetime(all_o3_df["Date"])
all_o3_df = all_o3_df.set_index("Date")
all_pm25_df = pd.read_csv("./all_years_pm25.csv")
#turn d... | github_jupyter |
# Inverse Kinematics tutorial
we'll demonstrate inverse kinematics on a baxter robot
## Setup
```
import numpy as np
from pykin.robots.bimanual import Bimanual
from pykin.kinematics.transform import Transform
from pykin.utils import plot_utils as plt
from pykin.utils.transform_utils import compute_pose_error
file_pa... | github_jupyter |
# Imports
```
import numpy as np
import pandas as pd
import glob
import re
from bs4 import BeautifulSoup
import nltk
nltk.download('punkt')
from nltk.tokenize import word_tokenize as wt
nltk.download('stopwords')
from nltk.corpus import stopwords
```
# Load data
```
def load_reviews(path, columns=["filename", 'rev... | github_jupyter |
# Nonstationary Temporal Matrix Factorization
Taking into account both seasonal differencing and first-order differencing.
```
import numpy as np
def compute_mape(var, var_hat):
return np.sum(np.abs(var - var_hat) / var) / var.shape[0]
def compute_rmse(var, var_hat):
return np.sqrt(np.sum((var - var_hat) **... | github_jupyter |
# FTE/BTE Experiment for MNIST & Fashion-MNIST
As an extension of the FTE/BTE experiments demonstrated on the CIFAR and food-101 datasets, we now look to examine the performance of progressive learning algorithms on the MNIST and fashion-MNIST datasets.
Due to their similarity in structure, both containing 60,000 tr... | github_jupyter |
<a href="https://colab.research.google.com/github/j3nguyen/jupyter_notebooks/blob/master/Optimizing_a_Library_Collection.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
#Stocking a digital library using combinatorial optimization
## Background
Supp... | github_jupyter |
# ---------------------------------------------------------------
# python best courses https://courses.tanpham.org/
# ---------------------------------------------------------------
# 100 numpy exercises
This is a collection of exercises that have been collected in the numpy mailing list, on stack overflow and in th... | github_jupyter |
# Feature Engineering
Feature engineering is an answer to the question, "How can I make the most of the data I have?"
Let's get started, then. How does one do feature engineering?
I'll assume you're familiar with pandas and the decision tree pipeline that we're using for this project. That's the algorithm we're goin... | github_jupyter |
```
# default_exp image.color_palette
# hide
from nbdev.showdoc import *
# hide
%reload_ext autoreload
%autoreload 2
```
# Color Palettes
> Tools for generating color palettes of various data-sets.
```
# export
def pascal_voc_palette(num_cls=None):
"""
Generates the PASCAL Visual Object Classes (PASCAL VOC) ... | github_jupyter |
```
%load_ext autoreload
%autoreload 2
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
```
## Introduction
In order to get you familiar with graph ideas,
I have deliberately chosen to steer away from
the more pedantic matters
of loading graph data to and from disk.
That said, the following scenario ... | github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/Algorithms/center_pivot_irrigation_detector.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a t... | github_jupyter |
# M² Real Examples
**Scott Prahl**
**Mar 2021**
This notebook demonstrates what happens when the ISO 11146 guidelines are violated.
---
*If* `` laserbeamsize `` *is not installed, uncomment the following cell (i.e., delete the initial #) and execute it with* `` shift-enter ``. *Afterwards, you may need to restart ... | github_jupyter |
# Introduction to Numpy
This is a NumPy cheat sheet that is created in the Treehouse course [Introduction to NumPy](https://teamtreehouse.com/library/introduction-to-numpy)
```
import matplotlib.pyplot as plt
import numpy as np
np.__version__
```
## Differences between lists and NumPy Arrays
* An array's size is immu... | github_jupyter |
# Energy terms and energy equation
There are several different energy terms that are implemented in `micromagneticmodel`. Here, we will provide a short list of them, together with some basic properties.
## Energy terms
### 1. Exchange energy
The main parameter required for the exchange energy is the exchange energy... | github_jupyter |
# Edafa on ImageNet dataset
This notebook shows an example on how to use Edafa to obtain better results on **classification task**. We use [ImageNet](http://www.image-net.org/) dataset which has **1000 classes**. We use *pytorch* and pretrained weights of AlexNet. At the end we compare results of the same model with a... | github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or ... | github_jupyter |
# What's this PyTorch business?
You've written a lot of code in this assignment to provide a whole host of neural network functionality. Dropout, Batch Norm, and 2D convolutions are some of the workhorses of deep learning in computer vision. You've also worked hard to make your code efficient and vectorized.
For the ... | github_jupyter |
**Chapter 16 – Natural Language Processing with RNNs and Attention**
_This notebook contains all the sample code in chapter 16._
<table align="left">
<td>
<a target="_blank" href="https://colab.research.google.com/github/ageron/handson-ml2/blob/master/16_nlp_with_rnns_and_attention.ipynb"><img src="https://www.... | github_jupyter |
## The Psychology of Growth
The field of positive psychology studies what are the human behaviours that lead to a great life. You can think of it as the intersection between self help books with the academic rigor of statistics. One of the famous findings of positive psychology is the **Growth Mindset**. The idea is t... | github_jupyter |
<a href="https://colab.research.google.com/github/satyajitghana/TSAI-DeepVision-EVA4.0/blob/master/05_CodingDrill/EVA4S5F3.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Import Libraries
```
from __future__ import print_function
import torch
imp... | github_jupyter |
```
%load_ext autoreload
%autoreload 2
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import torch
import random
device = 'cuda' if torch.cuda.is_available() else 'cpu'
import os,sys
opj = os.path.join
from tqdm import tqdm
import acd
from copy import deepcopy
import torchvision.utils as vutils
i... | github_jupyter |
# Autoencoder
```
from keras.layers import Input, Dense
from keras.models import Model
import matplotlib.pyplot as plt
import matplotlib.colors as mcol
from matplotlib import cm
def graph_colors(nx_graph):
#cm1 = mcol.LinearSegmentedColormap.from_list("MyCmapName",["blue","red"])
#cm1 = mcol.Colormap('viridis'... | github_jupyter |
```
import os
import json
import matplotlib.pyplot as plt
import math
data_folder = "../../data/convergence_tests"
def load_summary(path):
with open(data_folder + "/" + path) as file:
return json.load(file)
def load_summaries(directory):
summary_files = [file for file in os.listdir(directory) if f... | github_jupyter |
# GPU
```
gpu_info = !nvidia-smi
gpu_info = '\n'.join(gpu_info)
print(gpu_info)
```
# CFG
```
CONFIG_NAME = 'config06.yml'
TITLE = '06t-efficientnet_b4_ns-512'
! git clone https://github.com/raijin0704/cassava.git
# ====================================================
# CFG
# =====================================... | github_jupyter |
# From Modeling to Evaluation
## Introduction
In this lab, we will continue learning about the data science methodology, and focus on the **Modeling** and **Evaluation** stages.
------------
## Table of Contents
1. [Recap](#0)<br>
2. [Data Modeling](#2)<br>
3. [Model Evaluation](#4)<br>
# Recap <a id="0"></a>
In... | github_jupyter |
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed u... | github_jupyter |
# A modest proposal for dataset
```
from IPython.display import Image
from IPython.core.display import HTML
Image(url= "https://falexwolf.de/img/scanpy/anndata.svg")
```
## Imports
```
import pinot
import numpy as np
import pandas as pd
```
## Munging and flattening stuff.
```
ds = pinot.data.moonshot_mixed()
# ... | github_jupyter |
# Weight Sampling Tutorial
If you want to fine-tune one of the trained original SSD models on your own dataset, chances are that your dataset doesn't have the same number of classes as the trained model you're trying to fine-tune.
This notebook explains a few options for how to deal with this situation. In particular... | github_jupyter |
```
## Importing the libraries
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
#Importing the dataset
dataset = pd.read_csv('E:\Python\data\Wine.csv')
X = dataset.iloc[:, :-1].values
y = dataset.iloc[:, -1].values
#Splitting the dataset into the Training set and Test set
from sklearn.model_selec... | github_jupyter |
# The generic Broker class
```
from abc import abstractmethod
class Broker(object):
def __init__(self, host, port):
self.host = host
self.port = port
self.__price_event_handler = None
self.__order_event_handler = None
self.__position_event_handler = None
@property
def on_price_event(self):
"""
Lis... | github_jupyter |
# Imports
```
import matplotlib.pyplot as plt
import numpy as np
import os
import PIL
import tensorflow as tf
# Keras
from tensorflow import keras
from tensorflow.keras import layers
from tensorflow.keras.models import Sequential
from tensorflow.keras.preprocessing.image import ImageDataGenerator
import pathlib
```... | github_jupyter |
# A brief demonstration of the double spike toolbox
This is a quick guide to the main features of the double spike toolbox for python. The package uses the numpy, scipy, and matplotlib libraries.
```
import doublespike as ds
import matplotlib.pyplot as plt
import numpy as np
```
# IsoData
A python object called `Iso... | github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.