text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
04 .Differential expression using DESeq2
========================================
The analysis process includes three main steps, namely normalization, dispersion estimation and test for differential expression.
```
library(phyloseq)
library(ggplot2)
library(scales)
library(gridExtra)
suppressPackageStartupMessages(library(DESeq2))
#rds <- readRDS("../../../../../data/tmp/microbiome/quant/qiime2/silva/physeq.rds")
rds <- readRDS(snakemake@input[[1]])
df <- data.frame(as(sample_data(rds), "matrix"))
group <- factor(df[,snakemake@params$condition])
cat("Condition: ", snakemake@params$condition, "\n")
cat("Summary:\n")
print(summary(group))
if (!is.null(snakemake@params$ref_level)){
cat("Setting", snakemake@params$ref_level, "as reference level", "\n")
group <- relevel(group, ref=snakemake@params$ref_level)
}
df[,"Sample_Group"] <- group
sample_data(rds) <- df
block <- NULL
if (!is.null(snakemake@params$block)){
print(cat("\nBlocking factor given as: ", snakemake@params$block, "\n"))
block <- factor(df[,snakemake@params$block])
print(summary(block))
df[,snakemake@params$block] <- block
sample_data(rds) <- df
}
if (!is.null(snakemake@params$taxrank)){
cat("Taxrank: ", snakemake@params$taxrank, "\n")
if (require(speedyseq)){
rds <- speedyseq::tax_glom(rds, taxrank=snakemake@params$taxrank)
} else{
rds <- tax_glom(rds, taxrank=snakemake@params$taxrank)
}
} else{
cat("Taxrank: None, Data is not aggregated. Each feature is an ASV (specific sequence)", "\n")
}
rds
#if (!is.null(snakemake@params$independent_filtering)){
# if (snakemake@params$independent_filtering == TRUE){
# rds <- filter_taxa(rds, function(x) sum(x>0) > 10, TRUE)
# }
#}
counts <- as(otu_table(rds), "matrix")
head(counts)
df <- data.frame(as(sample_data(rds), "matrix"))
fun_summary=function(x){
out=c(quantile(x,c(0,0.25,0.5),type=1),mean(x),quantile(x,c(0.75,1),type=1))
names(out)=c("Min.","1st Qu.","Median","Mean","3rd Qu.","Max.")
return(round(out,0))
}
t(apply(counts, 2, fun_summary))
if (utils::packageVersion("ggplot2") >= "3.3.0") expand_scale <- expansion
barplotTotal <- function(counts, group, col=c("lightblue","orange","MediumVioletRed","SpringGreen"), outfile=TRUE){
if (outfile) png(filename="figures/barplotTotal.png", width=min(3600, 1800+800*ncol(counts)/10), height=1800, res=300)
d <- data.frame(tc=colSums(counts)/1e3, sample=factor(colnames(counts), colnames(counts)), group)
print(ggplot(d, aes(x=.data$sample, y=.data$tc, fill=.data$group)) +
geom_bar(stat="identity", show.legend=TRUE) +
labs(fill="") +
scale_fill_manual(values=col) +
xlab("Samples") +
ylab("Total read count (thousands)") +
scale_y_continuous(expand=expand_scale(mult=c(0.01, 0.05))) +
ggtitle("Total read count per sample (million)") +
theme(axis.text.x=element_text(angle=90, hjust=1, vjust=0.5)))
if (outfile) dev.off()
}
barplotTotal(counts, df$Sample_Group, outfile=FALSE)
gm_mean = function(x, na.rm=TRUE){
exp(sum(log(x[x > 0]), na.rm=na.rm) / length(x))
}
if (!is.null(snakemake@params$block)){
Sample_Group <- group
des <- model.matrix(~block + Sample_Group)
dds = phyloseq_to_deseq2(rds, design=des)
} else{
dds = phyloseq_to_deseq2(rds, design=~Sample_Group)
}
geoMeans = apply(counts(dds), 1, gm_mean)
dds = estimateSizeFactors(dds, geoMeans=geoMeans)
de = DESeq(dds)
countsBoxplots <- function(object, group, col = c("lightblue","orange","MediumVioletRed","SpringGreen"), outfile=TRUE){
if (class(object)=="DESeqDataSet"){
counts <- counts(object)
#counts <- removeNull(counts)
norm.counts <- counts(object, normalized=TRUE)
#norm.counts <- removeNull(norm.counts)
} else{
counts <- object$counts
#counts <- removeNull(counts)
tmm <- object$samples$norm.factors
N <- colSums(object$counts)
f <- tmm * N/mean(tmm * N)
norm.counts <- scale(object$counts, center=FALSE, scale=f)
#norm.counts <- removeNull(norm.counts)
}
if (outfile) png(filename="figures/countsBoxplots.png", width=2*min(2200, 1800+800*ncol(norm.counts)/10), height=1800, res=300)
d <- stack(as.data.frame(counts))
d$group <- rep(group, each=nrow(counts))
p1 <- ggplot(d) +
geom_boxplot(aes(x=.data$ind, y=.data$values+1, fill=.data$group), show.legend=TRUE) +
labs(fill="") +
scale_y_continuous(trans = log10_trans(),
breaks = trans_breaks("log10", function(x) 10^x),
labels = trans_format("log10", math_format(~10^.x))) +
scale_fill_manual(values=col) +
xlab("Samples") +
ylab("Raw counts") +
ggtitle("Raw counts distribution") +
theme(axis.text.x=element_text(angle=90, hjust=1, vjust=0.5))
d <- stack(as.data.frame(norm.counts))
d$group <- rep(group, each=nrow(norm.counts))
p2 <- ggplot(d) +
geom_boxplot(aes(x=.data$ind, y=.data$values+1, fill=.data$group), show.legend=TRUE) +
labs(fill="") +
scale_y_continuous(trans = log10_trans(),
breaks = trans_breaks("log10", function(x) 10^x),
labels = trans_format("log10", math_format(~10^.x))) +
scale_fill_manual(values=col) +
xlab("Samples") +
ylab("Normalized counts") +
ggtitle("Normalized counts distribution") +
theme(axis.text.x=element_text(angle=90, hjust=1, vjust=0.5))
grid.arrange(p1, p2, nrow=2, ncol=1)
if (outfile) dev.off()
}
countsBoxplots(de, df$Sample_Group, outfile=FALSE)
densityPlot <- function(counts, group, col=c("lightblue","orange","MediumVioletRed","SpringGreen"), outfile=TRUE){
if (outfile) png(filename="figures/densplot.png", width=2000, height=1800, res=300)
#counts <- removeNull(counts)
d <- stack(data.frame(counts))
d$group <- rep(group, each=nrow(counts))
print(ggplot(d, aes(x=.data$values+1)) +
stat_density(aes(group=.data$ind, color=.data$group), position="identity", geom="line", show.legend=TRUE) +
scale_x_continuous(trans = log10_trans(),
breaks = trans_breaks("log10", function(x) 10^x),
labels = trans_format("log10", math_format(~10^.x))) +
labs(color="") +
scale_colour_manual(values=col) +
xlab("Raw counts") +
ylab("Density") +
ggtitle("Density of counts distribution"))
if (outfile) dev.off()
}
densityPlot(counts, df$Sample_Group, outfile=FALSE)
majSequences <- function(rds, n=3, group, taxrank="Species", col=c("lightblue","orange","MediumVioletRed","SpringGreen"), outfile=TRUE){
counts <- as(otu_table(rds), "matrix")
seqnames <- apply(counts, 2, function(x){x <- sort(x, decreasing=TRUE); names(x)[1:n]})
seqnames <- unique(unlist(as.character(seqnames)))
sum <- apply(counts,2,sum)
counts <- counts[seqnames,]
sum <- matrix(sum,nrow(counts),ncol(counts),byrow=TRUE)
p <- round(100*counts/sum,digits=3)
if (outfile) png(filename="figures/majSeq.png",width=min(3600,1800+800*ncol(counts)/10),height=1800,res=300)
maj <- apply(p, 2, max)
seqname <- rownames(p)[apply(p, 2, which.max)]
TAX <- data.frame(as(tax_table(rds), "matrix"))[seqname,]
taxname <- as.character(TAX[,taxrank])
taxname[is.na(taxname)] <- as.character(TAX[is.na(taxname),"Genus"])
d <- data.frame(maj=maj, sample=factor(names(maj), levels=names(maj)), group, seqname=seqname)
print(ggplot(d, aes(x=.data$sample, y=.data$maj, fill=.data$group)) +
geom_bar(stat="identity", show.legend=TRUE) +
labs(fill="") +
scale_fill_manual(values=col) +
xlab("Samples") +
ylab("Percentage of reads") +
scale_y_continuous(expand=expand_scale(mult=c(0.01, 0.05))) +
ggtitle("Percentage of reads from most expressed sequence") +
theme(axis.text.x=element_text(angle=90, hjust=1, vjust=0.5)) +
geom_text(aes(y=0.8*maj, label=taxname), color="black", size=2.5, angle=90, fontface="bold"))
if (outfile) dev.off()
return(invisible(p))
}
majSequences(rds, n=1, df$Sample_Group, outfile=FALSE)
dispersionsPlot <- function(dds, outfile=TRUE){
if (outfile) png(filename="figures/dispersionsPlot.png", width=3600, height=1800, res=300)
# dispersions plot
d <- as.data.frame(mcols(dds)[,c("baseMean", "dispGeneEst", "dispFit", "dispersion")])
d <- d[which(d$baseMean > 0),]
d <- data.frame(baseMean=rep(d$baseMean, 3),
value=c(d$dispGeneEst, d$dispersion, d$dispFit),
variable=factor(rep(c("dispGeneEst", "dispersion", "dispFit"), each=nrow(d)),
levels=c("dispGeneEst", "dispersion", "dispFit")))
p1 <- ggplot(d, aes(x=.data$baseMean, y=.data$value, colour=.data$variable)) +
geom_point(size=0.1) +
scale_x_continuous(trans = log10_trans(),
breaks = trans_breaks("log10", function(x) 10^x),
labels = trans_format("log10", math_format())) +
scale_y_continuous(trans = log10_trans(),
breaks = trans_breaks("log10", function(x) 10^x),
labels = trans_format("log10", math_format())) +
ylab("Dispersion") +
xlab("Mean of normalized counts") +
scale_colour_manual(
values=c("Black", "#377eb8", "#e41a1c"),
breaks=c("dispGeneEst", "dispersion", "dispFit"),
labels=c("Estimate", "Final", "Fit"),
name="") +
guides(colour = guide_legend(override.aes = list(size=2))) +
ggtitle("Dispersions")
# diagnostic of log normality
disp <- mcols(dds)$dispGeneEst
disp <- disp[!is.na(disp)]
disp <- disp[disp>1e-8]
disp <- log(disp)
mean.disp <- mean(disp,na.rm=TRUE)
sd.disp <- sd(disp,na.rm=TRUE)
d <- data.frame(disp)
p2 <- ggplot(data=d, aes(x=.data$disp)) +
geom_histogram(bins=80, aes(y=.data$..density..)) +
scale_y_continuous(expand=expand_scale(mult=c(0.01, 0.05))) +
xlab("Feature dispersion estimate") +
ylab("Density") +
ggtitle("log-normality dispersion diagnostic") +
stat_function(fun = dnorm, args = list(mean = mean.disp, sd = sd.disp))
grid.arrange(p1, p2, layout_matrix=matrix(c(1, 1, 1, 1, 1, 2, 2, 2, 2), nrow=1))
if (outfile) dev.off()
}
dispersionsPlot(de, outfile=FALSE)
##CONTRASTS <- list()
##df <- colData(de)
##condition <- df[,snakemake@params$condition]
##if (snakemake@params$test == "ALLvsREF"){
## ref <- snakemake@params$ref_level
## for (test in levels(condition)){
## if (test != ref) CONTRASTS[[paste0(test, "__vs__", ref)]] <- list(snakemake@params$condition, test, ref)
## }
##} else{
## for (comp in combn(nlevels(condition), 2, simplify=FALSE)){
## ref <- levels(condition)[comp[1]]
## test <- levels(condition)[comp[2]]
## CONTRASTS[[paste0(test, "__vs__", ref)]] <- list(snakemake@params$condition, test, ref)
## }
## }
nDiffTotal <- function(complete, alpha=0.05){
nDiffTotal <- matrix(NA,ncol=4,nrow=length(complete),dimnames=list(names(complete),c("Test vs Ref", "# down","# up","# total")))
for (name in names(complete)){
complete.name <- complete[[name]]
if (!is.null(complete.name$betaConv)){
nDiffTotal[name,2:3]=c(nrow(complete.name[which(complete.name$padj <= alpha & complete.name$betaConv & complete.name$log2FoldChange<=0),]),
nrow(complete.name[which(complete.name$padj <= alpha & complete.name$betaConv & complete.name$log2FoldChange>=0),]))
} else{
nDiffTotal[name,2:3]=c(nrow(complete.name[which(complete.name$padj <= alpha & complete.name$log2FoldChange<=0),]),
nrow(complete.name[which(complete.name$padj <= alpha & complete.name$log2FoldChange>=0),]))
}
}
nDiffTotal[,4] <- nDiffTotal[,2] + nDiffTotal[,3]
nDiffTotal[,1] <- gsub("_"," ",rownames(nDiffTotal))
rownames(nDiffTotal) <- NULL
return(nDiffTotal)
}
complete <- list()
#for (contrast in names(CONTRASTS)){
# res = results(de, cooksCutoff=FALSE, contrast=CONTRASTS[[contrast]], alpha=snakemake@params$alpha)
# tab = cbind(as(res, "data.frame"), as(tax_table(rds)[rownames(res), ], "matrix"))
# tab = tab[order(tab$padj, na.last=TRUE), ]
# fn <- file.path(snakemake@output[[1]], paste0(snakemake@params$taxrank, "_", contrast, ".txt"))
# dir.create(snakemake@output[[1]], showWarnings=FALSE, recursive=TRUE)
# write.table(tab, file=fn, sep="\t")
# complete[[contrast]] <- tab
#}
#nDiffTotal(complete)
rawpHist <- function(complete, outfile=TRUE){
ncol <- min(2, length(complete))
nrow <- ceiling(length(complete)/ncol)
if (outfile) png(filename="figures/rawpHist.png", width=cairoSizeWrapper(1800*ncol), height=cairoSizeWrapper(1800*nrow), res=300)
p <- list()
for (name in names(complete)){
complete.name <- complete[[name]]
complete.name <- complete.name[which(!is.na(complete.name$pvalue)),]
p[[name]] <- ggplot(data=complete.name, aes(x=.data$pvalue)) +
geom_histogram(binwidth=0.02) +
scale_y_continuous(expand=expand_scale(mult=c(0.01, 0.05))) +
xlab("Raw p-value") +
ylab("Frequency") +
ggtitle(paste0("Distribution of raw p-values - ", gsub("_"," ",name)))
}
tmpfun <- function(...) grid.arrange(..., nrow=nrow, ncol=ncol)
do.call(tmpfun, p)
if (outfile) dev.off()
}
#rawpHist(complete, outfile=FALSE)
volcanoPlot <- function(complete, alpha=0.05, outfile=TRUE, padjlim=NULL){
ncol <- min(2, length(complete))
nrow <- ceiling(length(complete)/ncol)
if (outfile) png(filename="figures/volcanoPlot.png", width=cairoSizeWrapper(1800*ncol), height=cairoSizeWrapper(1800*nrow), res=300)
p <- list()
for (name in names(complete)){
complete.name <- complete[[name]]
complete.name$padj[which(complete.name$padj==0)] <- .Machine$double.xmin
complete.name <- complete.name[which(!is.na(complete.name$padj)),]
complete.name$DE <- factor(ifelse(complete.name$padj <= alpha, "yes", "no"), levels=c("no", "yes"))
if (is.null(padjlim)) padjlim.name <- quantile(complete.name$padj, probs=0.01, na.rm=TRUE) else padjlim.name <- padjlim
complete.name$outfield <- factor(ifelse(complete.name$padj < padjlim.name, "top", "in"), levels=c("in", "top"))
complete.name$padj[which(complete.name$padj < padjlim.name)] <- padjlim.name
reverselog_trans <- function(base = exp(1)) {
trans <- function(x) -log(x, base)
inv <- function(x) base^(-x)
trans_new(paste0("reverselog-", format(base)), trans, inv,
log_breaks(base = base),
domain = c(.Machine$double.xmin, Inf))
}
p[[name]] <- ggplot(data=complete.name,
aes(x=.data$log2FoldChange, y=.data$padj, color=.data$DE, shape=.data$outfield)) +
geom_point(show.legend=FALSE, alpha=0.5) +
scale_y_continuous(trans = reverselog_trans(10),
breaks = trans_breaks("log10", function(x) 10^x),
labels = trans_format("log10", math_format(~10^.x))) +
scale_colour_manual(values=c("no"="black", "yes"="red"), drop=FALSE) +
scale_shape_manual(values=c("in"=16, "top"=17), drop=FALSE) +
xlab(expression(log[2]~fold~change)) +
ylab("Adjusted P-value") +
ggtitle(paste0("Volcano plot - ", gsub("_", " ", name)))
}
tmpfun <- function(...) grid.arrange(..., nrow=nrow, ncol=ncol)
do.call(tmpfun, p)
if (outfile) dev.off()
}
#volcanoPlot(complete, outfile=FALSE)
MAPlot <- function(complete, alpha=0.05, outfile=TRUE, log2FClim=NULL){
ncol <- min(2, length(complete))
nrow <- ceiling(length(complete)/ncol)
if (outfile) png(filename="figures/MAPlot.png", width=cairoSizeWrapper(1800*ncol), height=cairoSizeWrapper(1800*nrow), res=300)
p <- list()
for (name in names(complete)){
complete.name <- complete[[name]]
complete.name <- complete.name[which(complete.name$baseMean>0),]
complete.name$padj <- ifelse(is.na(complete.name$padj), 1, complete.name$padj)
complete.name$DE <- factor(ifelse(complete.name$padj <= alpha, "yes", "no"), levels=c("no", "yes"))
py <- complete.name$log2FoldChange
if (is.null(log2FClim)) ymax <- quantile(abs(py[is.finite(py)]), probs=0.99) else ymax <- log2FClim
complete.name$log2FoldChange[which(py > ymax)] <- ymax
complete.name$log2FoldChange[which(py < -ymax)] <- -ymax
complete.name$outfield <- factor(ifelse(py > ymax, "top", ifelse(py < -ymax, "bottom", "in")),
levels=c("bottom", "in", "top"))
p[[name]] <- ggplot(data=complete.name,
aes(x=.data$baseMean, y=.data$log2FoldChange, color=.data$DE, fill=.data$DE, shape=.data$outfield)) +
scale_x_continuous(trans = log10_trans(),
breaks = trans_breaks("log10", function(x) 10^x),
labels = trans_format("log10", math_format(~10^.x))) +
geom_point(show.legend=FALSE, alpha=0.5, size=0.8) +
scale_colour_manual(values=c("no"="black", "yes"="red"), drop=FALSE) +
scale_shape_manual(values=c("bottom"=25, "in"=21, "top"=24), drop=FALSE) +
scale_fill_manual(values=c("no"="black", "yes"="red"), drop=FALSE) +
scale_y_continuous(expand=expand_scale(mult=c(0.03, 0.03))) +
xlab("Mean of normalized counts") +
ylab(expression(log[2]~fold~change)) +
ggtitle(paste0("MA-plot - ", gsub("_"," ",name)))
}
tmpfun <- function(...) grid.arrange(..., nrow=nrow, ncol=ncol)
do.call(tmpfun, p)
if (outfile) dev.off()
}
#MAPlot(complete, outfile=FALSE)
all_sign <- c()
#for (name in names(complete)){
# complete.name <- complete[[name]]
# complete.name$padj <- ifelse(is.na(complete.name$padj), 1, complete.name$padj)
# ids <- rownames(complete.name)[complete.name$padj <= snakemake@params$alpha]
#all_sign <- c(all_sign, ids)
# }
#all_sign <- unique(all_sign)
if (length(all_sign) > 1){
gpac <- prune_taxa(all_sign, rds)
gpac <- filter_taxa(gpac, function(x) sum(x>0) > 1, TRUE)
gpac <- prune_samples(sample_sums(gpac) > 1, gpac)
plot_heatmap(gpac, "NMDS", "bray", snakemake@params$condition, snakemake@params$taxrank, low="#66CCFF", high="#000033", na.value="white")
}
```
Hetmap of features found significant in analysis
| github_jupyter |
```
import json
with open("../out/202006222159_spanishfn.json") as fp:
data = json.load(fp)
from scipy.stats import rankdata
def rank_transform(orig):
data = np.copy(orig)
indices = [i for i, s in enumerate(data) if s > 0]
norm = rankdata([data[i] for i in indices], "max") / len(indices)
for i, s in zip(indices, norm):
data[i] = s
return data
import numpy as np
import pandas as pd
scaled_rank = [
(rank_transform(np.array(x["data"]).flatten()), x["id"])
for x in data["alignments"]
]
```
# Score techniques correlations
```
m = np.matrix([X for X, label in scaled_rank]).T
df = pd.DataFrame(m, columns=[label for X, label in scaled_rank])
df.corr()
indices = data["indices"]
en_len = len(indices[0])
l2_len = len(indices[1])
sums = np.sum([X for X, label in scaled_rank], axis=0).reshape(en_len, l2_len)
```
# Best alignment pairs scored by all techniques
```
sum_idx = [(idx[0], idx[1], score) for idx, score in np.ndenumerate(sums)]
sorted_idx = sorted(sum_idx, key=lambda x: -x[2])
printed = 0
for x in sorted_idx:
en_frm = data["frames"][indices[0][x[0]]]
l2_frm = data["frames"][indices[1][x[1]]]
if en_frm["name"] != l2_frm["name"] and (len(en_frm["LUs"]) > 0 and len(l2_frm["LUs"]) > 0):
score = '{:.3f}'.format(x[2]/10)
print(f'{score}: {(en_frm["name"]+"("+en_frm["gid"]+")").ljust(40)} {l2_frm["name"]} ({l2_frm["gid"]})')
printed += 1
printed += 1
if printed == 50:
break
```
# Worst alignment pairs compared to baseline
The baseline in this case is name matching. This list sorts alignemnt pairs by the difference of their aggregated score and the name_matching score and only prints frame pairs with the same name (baseline).
```
try:
baseline = next(np.array(x["data"]) for x in data["alignments"] if x["id"] == "name_matching")
except:
baseline = next(np.array(x["data"]) for x in data["alignments"] if x["id"] == "id_matching")
sum_idx = [(idx[0], idx[1], score) for idx, score in np.ndenumerate(baseline - (sums / 10))]
sorted_idx = [x for x in sorted(sum_idx, key=lambda x: -x[2]) if x[2] > 0]
printed = 0
for x in sorted_idx:
en_frm = data["frames"][indices[0][x[0]]]
l2_frm = data["frames"][indices[1][x[1]]]
if len(en_frm["LUs"]) > 0 and len(l2_frm["LUs"]) > 0:
score = '{:.3f}'.format(1-x[2])
print(f'{score}: {(en_frm["name"]+"("+en_frm["gid"]+")").ljust(40)} {l2_frm["name"]} ({l2_frm["gid"]})')
printed += 1
if printed == 50:
break
import networkx as nx
def min_matching(scores):
G = nx.Graph()
G.add_nodes_from(indices[0])
G.add_nodes_from(indices[1])
edge_matrix = scores.reshape(en_len, l2_len)
G.add_edges_from([
(indices[0][idx[0]], indices[1][idx[1]], { "weight": 1-score })
for idx, score in np.ndenumerate(edge_matrix)
])
m = nx.bipartite.minimum_weight_full_matching(G, top_nodes=indices[0])
return {k:v for k, v in m.items() if k.endswith(".en")}
import time
matchings = []
for X, label in scaled_rank:
start = time.time()
matchings.append(min_matching(X))
print(f'Computed matching for {label}')
print("--- %s seconds ---" % (time.time() - start))
votes = {}
for m in matchings[1:]:
for k,v in m.items():
if (k, v) in votes:
votes[(k, v)] += 1
else:
votes[(k, v)] = 1
len(matchings)-1
sorted_idx = sorted([(k[0], k[1], v) for k, v in votes.items()], key=lambda x: -x[2])
printed = 0
for x in sorted_idx:
if x[2] <= 3:
continue
en_frm = data["frames"][x[0]]
l2_frm = data["frames"][x[1]]
if en_frm["name"] != l2_frm["name"]:
score = '{:d}'.format(x[2])
print(f'{score}: {(en_frm["name"]+"("+en_frm["gid"]+")").ljust(40)} {l2_frm["name"]} ({l2_frm["gid"]})')
printed += 1
if printed == 100:
break
```
| github_jupyter |
# Quantum Teleportation
This notebook demonstrates quantum teleportation. We first use Qiskit's built-in simulators to test our quantum circuit, and then try it out on a real quantum computer.
## Contents
1. [Overview](#overview)
2. [The Quantum Teleportation Protocol](#how)
3. [Simulating the Teleportation Protocol](#simulating)
3.1 [How will we Test this Result on a Real Quantum Computer?](#testing)
3.2 [Using the Statevector Simulator](#simulating-sv)
3.3 [Using the QASM Simulator](#simulating-qs)
4. [Teleportation on a Real Quantum Computer](#real_qc)
4.1 [IBM hardware and Deferred Measurement](#deferred-measurement)
4.2 [Executing](#executing)
4. [References](#references)
## 1. Overview <a id='overview'></a>
Alice wants to send quantum information to Bob. Specifically, suppose she wants to send the qubit state
$\vert\psi\rangle = \alpha\vert0\rangle + \beta\vert1\rangle$.
This entails passing on information about $\alpha$ and $\beta$ to Bob.
There exists a theorem in quantum mechanics which states that you cannot simply make an exact copy of an unknown quantum state. This is known as the no-cloning theorem. As a result of this we can see that Alice can't simply generate a copy of $\vert\psi\rangle$ and give the copy to Bob. We can only copy classical states (not superpositions).
However, by taking advantage of two classical bits and an entangled qubit pair, Alice can transfer her state $\vert\psi\rangle$ to Bob. We call this teleportation because, at the end, Bob will have $\vert\psi\rangle$ and Alice won't anymore.
## 2. The Quantum Teleportation Protocol <a id='how'></a>
To transfer a quantum bit, Alice and Bob must use a third party (Telamon) to send them an entangled qubit pair. Alice then performs some operations on her qubit, sends the results to Bob over a classical communication channel, and Bob then performs some operations on his end to receive Alice’s qubit.

We will describe the steps on a quantum circuit below. Here, no qubits are actually ‘sent’, you’ll just have to imagine that part!
First we set up our session:
```
# Do the necessary imports
import numpy as np
from qiskit import QuantumCircuit, QuantumRegister, ClassicalRegister, execute, BasicAer, IBMQ
from qiskit.visualization import plot_histogram, plot_bloch_multivector
from qiskit.extensions import Initialize
from qiskit_textbook.tools import random_state, array_to_latex
```
and create our quantum circuit:
```
qr = QuantumRegister(3) # Protocol uses 3 qubits
crz = ClassicalRegister(1) # and 2 classical bits
crx = ClassicalRegister(1) # in 2 different registers
teleportation_circuit = QuantumCircuit(qr, crz, crx)
```
#### Step 1
A third party, Telamon, creates an entangled pair of qubits and gives one to Bob and one to Alice.
The pair Telamon creates is a special pair called a Bell pair. In quantum circuit language, the way to create a Bell pair between two qubits is to first transfer one of them to the X-basis ($|+\rangle$ and $|-\rangle$) using a Hadamard gate, and then to apply a CNOT gate onto the other qubit controlled by the one in the X-basis.
```
def create_bell_pair(qc, a, b):
"""Creates a bell pair in qc using qubits a & b"""
qc.h(a) # Put qubit a into state |+>
qc.cx(a,b) # CNOT with a as control and b as target
## SETUP
# Protocol uses 3 qubits and 2 classical bits in 2 different registers
qr = QuantumRegister(3)
crz, crx = ClassicalRegister(1), ClassicalRegister(1)
teleportation_circuit = QuantumCircuit(qr, crz, crx)
## STEP 1
# In our case, Telamon entangles qubits q1 and q2
# Let's apply this to our circuit:
create_bell_pair(teleportation_circuit, 1, 2)
# And view the circuit so far:
teleportation_circuit.draw()
```
Let's say Alice owns $q_1$ and Bob owns $q_2$ after they part ways.
#### Step 2
Alice applies a CNOT gate to $q_1$, controlled by $\vert\psi\rangle$ (the qubit she is trying to send Bob). Then Alice applies a Hadamard gate to $|\psi\rangle$. In our quantum circuit, the qubit ($|\psi\rangle$) Alice is trying to send is $q_0$:
```
def alice_gates(qc, psi, a):
qc.cx(psi, a)
qc.h(psi)
## SETUP
# Protocol uses 3 qubits and 2 classical bits in 2 different registers
qr = QuantumRegister(3)
crz, crx = ClassicalRegister(1), ClassicalRegister(1)
teleportation_circuit = QuantumCircuit(qr, crz, crx)
## STEP 1
create_bell_pair(teleportation_circuit, 1, 2)
## STEP 2
teleportation_circuit.barrier() # Use barrier to separate steps
alice_gates(teleportation_circuit, 0, 1)
teleportation_circuit.draw()
```
#### Step 3
Next, Alice applies a measurement to both qubits that she owns, $q_1$ and $\vert\psi\rangle$, and stores this result in two classical bits. She then sends these two bits to Bob.
```
def measure_and_send(qc, a, b):
"""Measures qubits a & b and 'sends' the results to Bob"""
qc.barrier()
qc.measure(a,0)
qc.measure(b,1)
## SETUP
# Protocol uses 3 qubits and 2 classical bits in 2 different registers
qr = QuantumRegister(3)
crz, crx = ClassicalRegister(1), ClassicalRegister(1)
teleportation_circuit = QuantumCircuit(qr, crz, crx)
## STEP 1
create_bell_pair(teleportation_circuit, 1, 2)
## STEP 2
teleportation_circuit.barrier() # Use barrier to separate steps
alice_gates(teleportation_circuit, 0, 1)
## STEP 3
measure_and_send(teleportation_circuit, 0 ,1)
teleportation_circuit.draw()
```
#### Step 4
Bob, who already has the qubit $q_2$, then applies the following gates depending on the state of the classical bits:
00 $\rightarrow$ Do nothing
01 $\rightarrow$ Apply $X$ gate
10 $\rightarrow$ Apply $Z$ gate
11 $\rightarrow$ Apply $ZX$ gate
(*Note that this transfer of information is purely classical*.)
```
# This function takes a QuantumCircuit (qc), integer (qubit)
# and ClassicalRegisters (crz & crx) to decide which gates to apply
def bob_gates(qc, qubit, crz, crx):
# Here we use c_if to control our gates with a classical
# bit instead of a qubit
qc.x(qubit).c_if(crx, 1) # Apply gates if the registers
qc.z(qubit).c_if(crz, 1) # are in the state '1'
## SETUP
# Protocol uses 3 qubits and 2 classical bits in 2 different registers
qr = QuantumRegister(3)
crz, crx = ClassicalRegister(1), ClassicalRegister(1)
teleportation_circuit = QuantumCircuit(qr, crz, crx)
## STEP 1
create_bell_pair(teleportation_circuit, 1, 2)
## STEP 2
teleportation_circuit.barrier() # Use barrier to separate steps
alice_gates(teleportation_circuit, 0, 1)
## STEP 3
measure_and_send(teleportation_circuit, 0 ,1)
## STEP 4
teleportation_circuit.barrier() # Use barrier to separate steps
bob_gates(teleportation_circuit, 2, crz, crx)
teleportation_circuit.draw()
```
And voila! At the end of this protocol, Alice's qubit has now teleported to Bob.
## 3. Simulating the Teleportation Protocol <a id='simulating'></a>
### 3.1 How Will We Test the Protocol on a Quantum Computer? <a id='testing'></a>
In this notebook, we will initialise Alice's qubit in a random state $\vert\psi\rangle$ (`psi`). This state will be created using an `Initialize` gate on $|q_0\rangle$. In this chapter we use the function `random_state` to choose `psi` for us, but feel free to set `psi` to any qubit state you want.
```
# Create random 1-qubit state
psi = random_state(1)
# Display it nicely
array_to_latex(psi, pretext="|\\psi\\rangle =")
# Show it on a Bloch sphere
plot_bloch_multivector(psi)
```
Let's create our initialisation gate to create $|\psi\rangle$ from the state $|0\rangle$:
```
init_gate = Initialize(psi)
init_gate.label = "init"
```
If the quantum teleportation circuit works, then at the end of the circuit the qubit $|q_2\rangle$ will be in this state. We will check this using the statevector simulator.
### 3.2 Using the Statevector Simulator <a id='simulating-sv'></a>
We can use the statevector simulator to verify our qubit has been teleported.
```
## SETUP
qr = QuantumRegister(3) # Protocol uses 3 qubits
crz = ClassicalRegister(1) # and 2 classical registers
crx = ClassicalRegister(1)
qc = QuantumCircuit(qr, crz, crx)
## STEP 0
# First, let's initialise Alice's q0
qc.append(init_gate, [0])
qc.barrier()
## STEP 1
# Now begins the teleportation protocol
create_bell_pair(qc, 1, 2)
qc.barrier()
## STEP 2
# Send q1 to Alice and q2 to Bob
alice_gates(qc, 0, 1)
## STEP 3
# Alice then sends her classical bits to Bob
measure_and_send(qc, 0, 1)
## STEP 4
# Bob decodes qubits
bob_gates(qc, 2, crz, crx)
# Display the circuit
qc.draw()
```
At the time of writing, there is a rendering issue with the `Initialise` gate in the image above, but the circuit operates just fine. We can see below, using our statevector simulator, that the state of $|q_2\rangle$ is the same as the state $|\psi\rangle$ we created above, while the states of $|q_0\rangle$ and $|q_1\rangle$ have been collapsed to either $|0\rangle$ or $|1\rangle$. The state $|\psi\rangle$ has been teleported from qubit 0 to qubit 2.
```
backend = BasicAer.get_backend('statevector_simulator')
out_vector = execute(qc, backend).result().get_statevector()
plot_bloch_multivector(out_vector)
```
You can run this cell a few times to make sure. You may notice that the qubits 0 & 1 change states, but qubit 2 is always in the state $|\psi\rangle$.
### 3.3 Using the QASM Simulator <a id='simulating-qs'></a>
Quantum teleportation is designed to send qubits between two parties. We do not have the hardware to demonstrate this, but we can demonstrate that the gates perform the correct transformations on a single quantum chip. Here we use the QASM simulator to simulate how we might test our protocol.
On a real quantum computer, we would not be able to sample the statevector, so if we wanted to check our teleportation circuit is working, we need to do things slightly differently. You will remember that we used `Initialize` to turn our $|0\rangle$ qubit into the state $|\psi\rangle$:
$$ |0\rangle \xrightarrow{\text{Initialise}} |\psi\rangle $$
Since all quantum gates are reversible, we can find the inverse of Initialise using:
```
inverse_init_gate = init_gate.gates_to_uncompute()
```
This operation has the property:
$$ |\psi\rangle \xrightarrow{\text{Inverse Initialise}} |0\rangle $$
To prove the qubit $|q_0\rangle$ has been teleported to $|q_2\rangle$, if we do this inverse initialisation on $|q_2\rangle$, we expect to measure $|0\rangle$ with certainty. We do this in the circuit below:
```
## SETUP
qr = QuantumRegister(3) # Protocol uses 3 qubits
crz = ClassicalRegister(1) # and 2 classical registers
crx = ClassicalRegister(1)
qc = QuantumCircuit(qr, crz, crx)
## STEP 0
# First, let's initialise Alice's q0
qc.append(init_gate, [0])
qc.barrier()
## STEP 1
# Now begins the teleportation protocol
create_bell_pair(qc, 1, 2)
qc.barrier()
## STEP 2
# Send q1 to Alice and q2 to Bob
alice_gates(qc, 0, 1)
## STEP 3
# Alice then sends her classical bits to Bob
measure_and_send(qc, 0, 1)
## STEP 4
# Bob decodes qubits
bob_gates(qc, 2, crz, crx)
## STEP 5
# reverse the initialisation process
qc.append(inverse_init_gate, [2])
# Display the circuit
qc.draw()
```
Again, there is a rendering issue with the `inverse_init_gate` (called 'disentangler' on the circuit diagram), but we can clearly see the gate appearing in the image. Finally, we measure the third qubit and store the result in the third classical bit:
```
# Need to add a new ClassicalRegister
# to see the result
cr_result = ClassicalRegister(1)
qc.add_register(cr_result)
qc.measure(2,2)
qc.draw()
```
and we run our experiment:
```
backend = BasicAer.get_backend('qasm_simulator')
counts = execute(qc, backend, shots=1024).result().get_counts()
plot_histogram(counts)
```
We can see we have a 100% chance of measuring $q_2$ (the leftmost bit in the string) in the state $|0\rangle$. This is the expected result, and indicates the teleportation protocol has worked properly.
## 4. Teleportation on a Real Quantum Computer <a id='real_qc'></a>
### 4.1 IBM hardware and Deferred Measurement <a id='deferred-measurement'></a>
The IBM quantum computers currently do not support instructions after measurements, meaning we cannot run the quantum teleportation in its current form on real hardware. Fortunately, this does not limit our ability to perform any computations due to the _deferred measurement principle_ discussed in chapter 4.4 of [1]. The principle states that any measurement can be postponed until the end of the circuit, i.e. we can move all the measurements to the end, and we should see the same results.

Any benefits of measuring early are hardware related: If we can measure early, we may be able to reuse qubits, or reduce the amount of time our qubits are in their fragile superposition. In this example, the early measurement in quantum teleportation would have allowed us to transmit a qubit state without a direct quantum communication channel.
While moving the gates allows us to demonstrate the "teleportation" circuit on real hardware, it should be noted that the benefit of the teleportation process (transferring quantum states via classical channels) is lost.
Let us re-write the `bob_gates` function to `new_bob_gates`:
```
def new_bob_gates(qc, a, b, c):
qc.cz(a, c)
qc.cx(b, c)
```
And create our new circuit:
```
qc = QuantumCircuit(3,1)
# First, let's initialise Alice's q0
qc.append(init_gate, [0])
qc.barrier()
# Now begins the teleportation protocol
create_bell_pair(qc, 1, 2)
qc.barrier()
# Send q1 to Alice and q2 to Bob
alice_gates(qc, 0, 1)
qc.barrier()
# Alice sends classical bits to Bob
new_bob_gates(qc, 0, 1, 2)
# We undo the initialisation process
qc.append(inverse_init_gate, [2])
# See the results, we only care about the state of qubit 2
qc.measure(2,0)
# View the results:
qc.draw()
```
### 4.2 Executing <a id='executing'></a>
```
# First, see what devices we are allowed to use by loading our saved accounts
IBMQ.load_account()
provider = IBMQ.get_provider(hub='ibm-q')
provider.backends()
# get the least-busy backend at IBM and run the quantum circuit there
from qiskit.providers.ibmq import least_busy
backend = least_busy(provider.backends(filters=lambda b: b.configuration().n_qubits >= 3 and
not b.configuration().simulator and b.status().operational==True))
job_exp = execute(qc, backend=backend, shots=8192)
# Get the results and display them
exp_result = job_exp.result()
exp_measurement_result = exp_result.get_counts(qc)
print(exp_measurement_result)
plot_histogram(exp_measurement_result)
```
As we see here, there are a few results in which we measured $|1\rangle$. These arise due to errors in the gates and the qubits. In contrast, our simulator in the earlier part of the notebook had zero errors in its gates, and allowed error-free teleportation.
```
error_rate_percent = sum([exp_measurement_result[result] for result in exp_measurement_result.keys() if result[0]=='1']) \
* 100./ sum(list(exp_measurement_result.values()))
print("The experimental error rate : ", error_rate_percent, "%")
```
## 5. References <a id='references'></a>
[1] M. Nielsen and I. Chuang, Quantum Computation and Quantum Information, Cambridge Series on Information and the Natural Sciences (Cambridge University Press, Cambridge, 2000).
```
import qiskit
qiskit.__qiskit_version__
```
| github_jupyter |
# Model viewer
Quickly view results of previously run models in Jupyter Notebook. Results and parameters can also be viewed in the directory itself, but this notebook provides a quick way to either (1) view all data from a single run in one place and (2) compare the same file across multiple runs. It does require some familiarity with how the output files are named.
```
import glob
import json
import os
import pprint as pp
import matplotlib.pyplot as plt
from PIL import Image
from ea_drought_burn.config import DATA_DIR
# Set working directory to the earthpy data directory
os.chdir(os.path.join(DATA_DIR, "woolsey-fire"))
def view_output(val):
"""View output from one or more models
Parameters
----------
val: str
a model id or filename
Returns
-------
None
"""
return view_file(val) if val[-4] == "." else view_model(val)
def view_file(filename):
"""View a single output across all models
Parameters
----------
filename: str
the filename to view
Returns
-------
None
"""
path = os.path.join("outputs", "models", "*")
ext = os.path.splitext(filename)[-1].lower()
for fp in sorted(glob.iglob(os.path.join(path, filename))):
if ext == ".json":
print(f"{fp}\n")
with open(fp) as f:
pp.pprint(json.load(f), sort_dicts=False)
print("-" * 80)
elif ext == ".png":
im = Image.open(fp)
fig, ax = plt.subplots(figsize=(20, 20))
ax.imshow(im, interpolation=None)
plt.axis("off")
elif ext == ".txt":
print(f"{fp}\n")
with open(fp) as f:
print(f.read())
print("-" * 80)
def view_model(model_id):
"""View the results of a saved model
Parameters
----------
model_id: str
the id of the model to view
Returns
-------
None
"""
path = os.path.join("outputs", "models", model_id)
# Show classification report
for fp in sorted(glob.iglob(os.path.join(path, "*.txt"))):
print("Classification report\n")
with open(fp) as f:
print(f.read())
print("-" * 80)
# Show params and results as pretty-printed dicts
for fp in sorted(glob.iglob(os.path.join(path, "*.json"))):
print(f"{os.path.basename(fp)}\n")
with open(fp) as f:
pp.pprint(json.load(f), sort_dicts=False)
print("-" * 80)
# Show all saved images
for fp in sorted(glob.iglob(os.path.join(path, "*.png"))):
im = Image.open(fp)
fig, ax = plt.subplots(figsize=(20, 20))
ax.imshow(im, interpolation=None)
plt.axis("off")
# List completed models
model_dir = os.path.join("outputs", "models")
models = []
for dirname in os.listdir(model_dir):
if os.path.isdir(os.path.join(model_dir, dirname)):
models.append(dirname)
# Sort by run time
models.sort(key=lambda fn: fn.split("_")[-1])
models
# View model output
view_output(models[-1])
```
| github_jupyter |
# Bayesian Survival Analysis
Copyright 2017 Allen Downey
MIT License: https://opensource.org/licenses/MIT
```
from __future__ import print_function, division
%matplotlib inline
import warnings
warnings.filterwarnings('ignore')
import numpy as np
import pandas as pd
import thinkbayes2
import thinkplot
```
## Survival analysis
Suppose that you are an auto insurance company interested in the time between collisions for a particular driver. If the probability of a collision is roughly constant over time, the time between collisions will follow an exponential distribution.
Here's an example with parameter $\lambda = 0.5$ collisions / year.
```
from thinkbayes2 import MakeExponentialPmf
pmf = MakeExponentialPmf(lam=0.5, high=30)
thinkplot.Pdf(pmf)
thinkplot.Config(xlabel='Lifetime', ylabel='PMF')
```
For the exponential distribution, the mean and standard deviation are $1/\lambda$.
In this case they are only approximate because we truncated the distribution.
```
pmf.Mean(), pmf.Std()
```
From the PMF, we can compute the CDF.
```
cdf = pmf.MakeCdf()
thinkplot.Cdf(cdf)
thinkplot.Config(xlabel='Lifetime', ylabel='CDF')
```
And from the CDF, we can compute the survival function, which is the complement of the CDF.
$SF(x) = Prob\{X > x\} = 1 - Prob\{X \le x\} = 1 - CDF(x)$
```
from survival import MakeSurvivalFromCdf
sf = MakeSurvivalFromCdf(cdf)
thinkplot.Plot(sf)
thinkplot.Config(xlabel='Lifetime', ylabel='Survival function')
```
From the survival function we can get the hazard function, which is the probability of a collision at $x$, given no collision prior to $x$.
```
hf = sf.MakeHazardFunction()
thinkplot.Plot(hf)
thinkplot.Config(xlabel='Lifetime', ylabel='Hazard function')
```
If the distribution is truly exponential, the hazard function is constant for all $x$.
In this case it goes to 1 at the end, again because we truncated the distribution.
**Exercise:** Go back and increase the value of `high`, and confirm that the hazard function is a constant until we approach the point where we cut off the distribution.
## Remaining lifetime
Given the survival function, we can compute the distribution of remaining lifetime, conditioned on current age. The following function computes the mean remaining lifetime for a range of ages.
```
def RemainingLifetime(sf):
"""Computes remaining lifetime as a function of age.
sf: survival function
returns: Series that maps from age to remaining lifetime
"""
pmf = sf.MakePmf()
d = {}
for t in sorted(pmf.Values()):
pmf[t] = 0
if pmf.Total():
pmf.Normalize()
d[t] = pmf.Mean() - t
return pd.Series(d)
```
And here's what it looks like for the exponential survival function.
```
mean_rem_life = RemainingLifetime(sf)
thinkplot.Plot(mean_rem_life)
thinkplot.Config(xlabel='Lifetime', ylabel='Survival function')
```
The mean time until a collision is pretty much constant, until we approach the point where we truncate the distribution.
## The Weibull distribution
The Weibull distribution is a generalization of the exponential distribution that takes an additional "shape" parameter, `k`.
When `k=1`, the Weibull is an exponential distribution. Other values of `k` yield survival curves with different shapes, and hazard functions that increase, decrease, or both. So the Weibull family can capture a wide range of survival patterns.
```
from thinkbayes2 import MakeWeibullPmf
pmf = MakeWeibullPmf(lam=2.0, k=1.5, high=30)
thinkplot.Pdf(pmf)
thinkplot.Config(xlabel='Lifetime', ylabel='PMF')
```
**Exercise**: In the previous section, replace the exponential distribution with a Weibull distribituion and run the analysis again. What can you infer about the values of the parameters and the behavior of the hazard function and remaining lifetime?
## Bayesian survival analysis
Suppose you are the manager of a large building with many light fixtures. To figure out how often you will need to replace lightbulbs, you install 10 bulbs and measure the time until they fail.
To generate some fake data, I'll choose a Weibull distribution and generate a random sample (let's suppose it's in years):
```
def SampleWeibull(lam, k, n=1):
return np.random.weibull(k, size=n) * lam
data = SampleWeibull(lam=2, k=1.5, n=10)
data
```
**Exercise:** Write a class called `LightBulb` that inherits from `Suite` and provides a `Likelihood` function that takes an observed lifespan as data and a tuple, `(lam, k)`, as a hypothesis. It should return a likelihood proportional to the probability of the observed lifespan in a Weibull distribution with the given parameters.
Test your method by creating a `LightBulb` object with an appropriate prior and update it with the data above.
Plot the posterior distributions of `lam` and `k`. As the sample size increases, does the posterior distribution converge on the values of `lam` and `k` used to generate the sample?
```
# Hint
from thinkbayes2 import Suite, Joint, EvalWeibullPdf
class LightBulb(Suite, Joint):
def Likelihood(self, data, hypo):
lam, k = hypo
x = data
like = 1
return like
# Solution goes here
from itertools import product
lams = np.linspace(0.001, 6, 101)
ks = np.linspace(0.001, 8, 101)
suite = LightBulb(product(lams, ks))
suite.UpdateSet(data)
thinkplot.Contour(suite)
pmf_lam = suite.Marginal(0)
thinkplot.Pdf(pmf_lam)
pmf_lam.Mean()
pmf_k = suite.Marginal(1)
thinkplot.Pdf(pmf_k)
pmf_k.Mean()
```
**Exercise:** Go back and run this analysis again with `n=20` and see if the posterior distributions seem to be converging on the actual parameters.
## Censored data
**Exercise:** Now suppose that instead of observing a complete lifespan, you observe a lightbulb that has operated for 1 year and is still working. Write another version of `LightBulb` that takes data in this form and performs an update.
```
# Hint
from thinkbayes2 import EvalWeibullCdf
class LightBulb2(Suite, Joint):
def Likelihood(self, data, hypo):
lam, k = hypo
x = data
like = 1
return like
# Solution goes here
from itertools import product
lams = np.linspace(0.001, 10, 101)
ks = np.linspace(0.001, 10, 101)
suite = LightBulb2(product(lams, ks))
suite.Update(1)
thinkplot.Contour(suite)
pmf_lam = suite.Marginal(0)
thinkplot.Pdf(pmf_lam)
pmf_lam.Mean()
pmf_k = suite.Marginal(1)
thinkplot.Pdf(pmf_k)
pmf_k.Mean()
```
Note: based on this data alone, we can rule out some small values of `lam` and `k`, but we can't rule out large values. Without more data or a more informative prior, the results are not useful.
To see why, try increasing the upper bounds in the prior distribition.
## Left censored data
**Exercise:** Suppose you install a light bulb and then you don't check on it for a year, but when you come back, you find that it has burned out. Extend `LightBulb` to handle this kind of data, too.
```
# Hint
class LightBulb3(Suite, Joint):
def Likelihood(self, data, hypo):
lam, k = hypo
x = data
like = 1
return like
# Solution goes here
from itertools import product
lams = np.linspace(0.001, 20, 101)
ks = np.linspace(0.001, 20, 101)
suite = LightBulb3(product(lams, ks))
suite.Update(1)
thinkplot.Contour(suite)
pmf_lam = suite.Marginal(0)
thinkplot.Pdf(pmf_lam)
pmf_lam.Mean()
pmf_k = suite.Marginal(1)
thinkplot.Pdf(pmf_k)
pmf_k.Mean()
```
This example has some of the same problems as the previous one. Based on this data alone, we can't pin down the parameters much.
## Pulling it together
**Exercise:** Suppose you have 15 lightbulbs installed at different times over a 10 year period. When you observe them, some have died and some are still working. Write a version of `LightBulb` that takes data in the form of a `(flag, x)` tuple, where:
1. If `flag` is `eq`, it means that `x` is the actual lifespan of a bulb that has died.
2. If `flag` is `gt`, it means that `x` is the current age of a bulb that is still working, so it is a lower bound on the lifespan.
3. If `flag` is `lt`, it means that `x` is the elapsed time between installation and the first time the bulb is seen broken, so it is an upper bound on the lifespan.
To help you test, I will generate some fake data.
First, I'll generate a Pandas DataFrame with random start times and lifespans. The columns are:
* `start`: time when the bulb was installed
* `lifespan`: lifespan of the bulb in years
* `end`: time when bulb died or will die
* `age_t`: age of the bulb at t=10
```
import pandas as pd
lam = 2
k = 1.5
n = 15
t_end = 10
starts = np.random.uniform(0, t_end, n)
lifespans = SampleWeibull(lam, k, n)
df = pd.DataFrame({'start': starts, 'lifespan': lifespans})
df['end'] = df.start + df.lifespan
df['age_t'] = t_end - df.start
df.head()
```
Now I'll process the DataFrame to generate data in the form we want for the update.
```
data = []
for i, row in df.iterrows():
if row.end < t_end:
data.append(('eq', row.lifespan))
else:
data.append(('gt', row.age_t))
for pair in data:
print(pair)
# Hint
class LightBulb4(Suite, Joint):
def Likelihood(self, data, hypo):
lam, k = hypo
flag, x = data
like = 1
return like
# Solution goes here
from itertools import product
lams = np.linspace(0.001, 10, 101)
ks = np.linspace(0.001, 10, 101)
suite = LightBulb4(product(lams, ks))
suite.UpdateSet(data)
thinkplot.Contour(suite)
pmf_lam = suite.Marginal(0)
thinkplot.Pdf(pmf_lam)
pmf_lam.Mean()
pmf_k = suite.Marginal(1)
thinkplot.Pdf(pmf_k)
pmf_k.Mean()
```
## Prediction
Suppose we know that, for a particular kind of lightbulb in a particular location, the distribution of lifespans is well modeled by a Weibull distribution with `lam=2` and `k=1.5`. If we install `n=100` lightbulbs and come back one year later, what is the distribution of `c`, the number of lightbulbs that have burned out?
The probability that any given bulb has burned out comes from the CDF of the distribution.
```
lam = 2
k = 1.5
p = EvalWeibullCdf(1, lam, k)
p
```
The number of bulbs that have burned out is distributed Binom(n, p).
```
from thinkbayes2 import MakeBinomialPmf
n = 100
pmf_c = MakeBinomialPmf(n, p)
thinkplot.Pdf(pmf_c)
```
Or we can approximate the distribution with a random sample.
```
n = 100
sample = np.random.binomial(n, p, 1000)
pdf_c = thinkbayes2.EstimatedPdf(sample)
thinkplot.Pdf(pdf_c)
np.mean(sample), np.std(sample)
```
**Exercise:** Now suppose that `lam` and `k` are not known precisely, but we have a `LightBulb` object that represents the joint posterior distribution of the parameters after seeing some data. Compute the posterior predictive distribution for `c`, the number of bulbs burned out after one year.
```
# Solution goes here
# Solution goes here
# Solution goes here
# Solution goes here
```
| github_jupyter |
# Imports
```
import numpy as np
import sklearn.metrics
from sklearn import linear_model
from sklearn.datasets import load_breast_cancer
```
# Load Data
"Breast Cancer" is a tiny dataset for binary classification
```
features, targets = load_breast_cancer(return_X_y=True)
print('Features')
print('shape:', features.shape)
print('data:')
print(features)
print('Targets')
print('shape:', targets.shape)
print('data:')
print(targets)
```
# Model
Create super simple logistic classifier
```
model = linear_model.LogisticRegression(solver='liblinear')
model.fit(features, targets)
```
Predicted outputs
```
predictions = model.predict(features)
print(predictions)
```
# Metrics
**Confusion Matrix**
```python
' Confusion matrix layout'
' PREDICTED LABEL'
' 0 1 '
'TRUE 0' [[ TN FP ]
'LABEL 1' [ FN TP ]]
```
Confusion matrix in sklearn
```
cm = sklearn.metrics.confusion_matrix(targets, predictions)
print('Confusion Matrix:')
print(cm)
```
Confusion matrix in pure numpy
```
def confusion_matrix(y_true, y_pred, result=None, nb_classes=None, norm='none'):
"""Compute confusion matrix. Works with NumPy and PyTorch tensors seamlessly"""
assert y_true.shape == y_pred.shape
if nb_classes==None:
nb_classes = int(max(y_true.max(), y_pred.max())) + 1
if result is None:
confusion_matrix = np.zeros((nb_classes, nb_classes), dtype=np.long)
else:
confusion_matrix = result
for true_class_idx in range(nb_classes):
y_pred_for_class = y_pred[y_true==true_class_idx]
for pred_class_idx in range(nb_classes):
tmp = (y_pred_for_class==pred_class_idx).sum()
confusion_matrix[true_class_idx, pred_class_idx] = tmp
if norm == 'none':
return confusion_matrix # return raw
elif norm == 'row':
return confusion_matrix / confusion_matrix.sum(axis=1, keepdims=True) # rows sum to 1
elif norm == 'col':
return confusion_matrix / confusion_matrix.sum(axis=0, keepdims=True) # cols sum to 1
else:
raise ValueError('norm must be "none", "row" or "col"')
cm = confusion_matrix(targets, predictions)
print(cm)
```
Confusion matrix manually for 2-class problem
```
pred_for_neg = predictions[targets==0] # predictions for class #1
pred_for_pos = predictions[targets==1] # predictions for class #2
TN = np.sum(pred_for_neg==0)
FP = np.sum(pred_for_neg==1)
FN = np.sum(pred_for_pos==0)
TP = np.sum(pred_for_pos==1)
cm = np.array([[TN, FP],
[FN, TP]])
print(cm)
```
Per class classification accuracy
```
cm_true = cm / cm.sum(axis=1, keepdims=True)
print(cm_true)
```
Per class accuracy for true classes only
```
cm_true.diagonal()
```
**Precision and Recall**
In sklearn
```
print('Accuracy: ', sklearn.metrics.accuracy_score(targets, predictions))
print('Precision: ', sklearn.metrics.precision_score(targets, predictions))
print('Recall: ', sklearn.metrics.recall_score(targets, predictions))
```
In numpy
```
# each cm row is actual class
assert cm.shape == (2, 2)
(TN, FP) = cm[0]
(FN, TP) = cm[1]
print('Accuracy: ', (TP+TN) / np.sum(cm))
print('Precision:', TP / (TP+FP))
print('Recall: ', TP / (TP+FN))
```
And manually from confusion matrix
```
print('Accuracy: ', cm.trace() / cm.sum() )
```
| github_jupyter |
# Sample Survey Bihar Election 2021 EDA
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
```
### Load the dataset into a pandas dataframe. Name the variable as “survey”.
```
survey=pd.read_excel('Sample Survey.xlsx',sheet_name='Data')
survey.head()
```
### How many samples were collected on each day.
```
survey['collection_date'].value_counts()
```
### What proportion of the total respondents were aged less than 45.
```
survey['age'].replace('24ko',24,inplace=True)
survey['age']= survey['age'].astype('int')
print('There are {} % of people in survey less than age 45'.format(len(survey[survey['age']<45])/len(survey)*100))
```
### Create a new column in the dataframe “age_group”. This column should contain the age group the respondent belongs to. The age groups are 18-25, 25-40, 40-55 and 55+.
```
ml1=list(range(18,26))
mydict1=dict.fromkeys(ml1,'18-25')
ml2=list(range(25,41))
mydict2=dict.fromkeys(ml2,'25-40')
ml3=list(range(40,56))
mydict3=dict.fromkeys(ml3,'40-55')
ml4=list(range(56,100))
mydict4=dict.fromkeys(ml4,'55+')
mydict1.update(mydict2)
mydict1.update(mydict3)
mydict1.update(mydict4)
survey['age_group']=survey['age'].map(mydict1)
survey.head()
```
### How many samples were collected for each age-group? Which age-group had the most samples.
```
survey['age_group'].value_counts()
```
### What proportion of the respondents had opted for the RJD party in both the Vote_Now and the Past_Vote questions.
```
x6=len(survey[(survey['Vote_Now']=='RJD') & (survey['Past_Vote']=='RJD')])/len(survey)
print('There are {} % of respondent for RJD both in Vote_Now & Past_Vote'.format(x6*100))
```
### For each day of sample collection, determine the proportion of respondents who were fully satisfied with the performance of the CM. So if there were a total of 1000 samples on day 1 and 300 out of those said they were fully satisfied, then our answer for that day would be 0.3.
```
survey.groupby(by='collection_date')[['CM_satisfaction']].apply(lambda x:np.sum(x=='Fully Satisfied')/len(x)*100)
```
### In a similar fashion create a day-wise proportion of respondents that opted fully dissatisfied with their MLA. Create a line plot of the result with date on x-axis and proportions on the y-axis.
```
x8=pd.DataFrame(survey.groupby(by='collection_date')[['MLA_satisfaction']].apply(lambda x:np.sum(x=='Fully Dissatisfied')/len(x)*100))
x8
import datetime
x8.index.date
x8.plot()
```
### Create a pivot-table (or crosstab) with index as Past_Vote, Column as Vote_Now and cell values as the count of samples.
```
pd.pivot_table(data=survey,index='Past_Vote',columns='Vote_Now',aggfunc='count',values='response_id')
```
### Repeat the above question with the cell values as the sum of “weight”.
```
pd.pivot_table(data=survey,index='Past_Vote',columns='Vote_Now',aggfunc='sum',values='weight')
```
### Create a dataframe by performing a group by over age_group and calculate the count of total samples under each age_group.
```
x11=pd.DataFrame(survey.groupby(by='age_group')[['response_id']].agg('count'))
x11
```
### Create a dataframe by performing a group by over age_group and finding the count of total samples for each age_group that opted for the JD(U) party in Vote_Now.
```
x12=pd.DataFrame(survey.groupby(by='age_group')[['Vote_Now']].apply(lambda x:np.sum(x=='JD(U)')))
x12
```
### Join/Merge the two dataframes from questions 12 and 13 with the common column as age_group.
```
df13=pd.merge(x11,x12,on='age_group')
df13
### end
```
| github_jupyter |
```
# !wget http://s3-ap-southeast-1.amazonaws.com/huseinhouse-storage/bert-bahasa/bert-bahasa-base.tar.gz
# !tar -zxf bert-bahasa-base.tar.gz
from tqdm import tqdm
import json
import bert
from bert import run_classifier
from bert import optimization
from bert import tokenization
from bert import modeling
import numpy as np
import tensorflow as tf
with open('sentiment.json') as fopen:
x = json.load(fopen)
texts = x['x']
labels = x['y']
MAX_SEQ_LENGTH = 100
!ls bert-bahasa-base
import sentencepiece as spm
from prepro_utils import preprocess_text, encode_ids, encode_pieces
sp_model = spm.SentencePieceProcessor()
sp_model.Load('bert-bahasa-base/sp10m.cased.v4.model')
with open('bert-bahasa-base/sp10m.cased.v4.vocab') as fopen:
v = fopen.read().split('\n')[:-1]
v = [i.split('\t') for i in v]
v = {i[0]: i[1] for i in v}
class Tokenizer:
def __init__(self, v):
self.vocab = v
pass
def tokenize(self, string):
return encode_pieces(sp_model, string, return_unicode=False, sample=False)
def convert_tokens_to_ids(self, tokens):
return [sp_model.PieceToId(piece) for piece in tokens]
def convert_ids_to_tokens(self, ids):
return [sp_model.IdToPiece(i) for i in ids]
tokenizer = Tokenizer(v)
BERT_INIT_CHKPNT = 'bert-bahasa-base/model.ckpt'
BERT_CONFIG = 'bert-bahasa-base/bert_config.json'
input_ids, input_masks, segment_ids = [], [], []
for text in tqdm(texts):
tokens_a = tokenizer.tokenize(text)
if len(tokens_a) > MAX_SEQ_LENGTH - 2:
tokens_a = tokens_a[:(MAX_SEQ_LENGTH - 2)]
tokens = ["<cls>"] + tokens_a + ["<sep>"]
segment_id = [0] * len(tokens)
input_id = tokenizer.convert_tokens_to_ids(tokens)
input_mask = [1] * len(input_id)
padding = [0] * (MAX_SEQ_LENGTH - len(input_id))
input_id += padding
input_mask += padding
segment_id += padding
input_ids.append(input_id)
input_masks.append(input_mask)
segment_ids.append(segment_id)
bert_config = modeling.BertConfig.from_json_file(BERT_CONFIG)
epoch = 10
batch_size = 60
warmup_proportion = 0.1
num_train_steps = int(len(texts) / batch_size * epoch)
num_warmup_steps = int(num_train_steps * warmup_proportion)
class Model:
def __init__(
self,
dimension_output,
learning_rate = 2e-5,
):
self.X = tf.placeholder(tf.int32, [None, None])
self.Y = tf.placeholder(tf.int32, [None])
model = modeling.BertModel(
config=bert_config,
is_training=False,
input_ids=self.X,
use_one_hot_embeddings=False)
output_layer = model.get_sequence_output()
self.logits_seq = tf.layers.dense(output_layer, dimension_output)
self.logits_seq = tf.identity(self.logits_seq, name = 'logits_seq')
self.logits = self.logits_seq[:, 0]
self.logits = tf.identity(self.logits, name = 'logits')
self.cost = tf.reduce_mean(
tf.nn.sparse_softmax_cross_entropy_with_logits(
logits = self.logits, labels = self.Y
)
)
self.optimizer = optimization.create_optimizer(self.cost, learning_rate,
num_train_steps, num_warmup_steps, False)
correct_pred = tf.equal(
tf.argmax(self.logits, 1, output_type = tf.int32), self.Y
)
self.accuracy = tf.reduce_mean(tf.cast(correct_pred, tf.float32))
dimension_output = 2
learning_rate = 2e-5
tf.reset_default_graph()
sess = tf.InteractiveSession()
model = Model(
dimension_output,
learning_rate
)
sess.run(tf.global_variables_initializer())
var_lists = tf.get_collection(tf.GraphKeys.TRAINABLE_VARIABLES, scope = 'bert')
saver = tf.train.Saver(var_list = var_lists)
saver.restore(sess, BERT_INIT_CHKPNT)
from sklearn.model_selection import train_test_split
train_input_ids, test_input_ids, train_Y, test_Y = train_test_split(
input_ids, labels, test_size = 0.2
)
import time
EARLY_STOPPING, CURRENT_CHECKPOINT, CURRENT_ACC, EPOCH = 3, 0, 0, 0
while True:
lasttime = time.time()
if CURRENT_CHECKPOINT == EARLY_STOPPING:
print('break epoch:%d\n' % (EPOCH))
break
train_acc, train_loss, test_acc, test_loss = 0, 0, 0, 0
pbar = tqdm(
range(0, len(train_input_ids), batch_size), desc = 'train minibatch loop'
)
for i in pbar:
index = min(i + batch_size, len(train_input_ids))
batch_x = train_input_ids[i: index]
batch_y = train_Y[i: index]
acc, cost, _ = sess.run(
[model.accuracy, model.cost, model.optimizer],
feed_dict = {
model.Y: batch_y,
model.X: batch_x,
},
)
assert not np.isnan(cost)
train_loss += cost
train_acc += acc
pbar.set_postfix(cost = cost, accuracy = acc)
pbar = tqdm(range(0, len(test_input_ids), batch_size), desc = 'test minibatch loop')
for i in pbar:
index = min(i + batch_size, len(test_input_ids))
batch_x = test_input_ids[i: index]
batch_y = test_Y[i: index]
acc, cost = sess.run(
[model.accuracy, model.cost],
feed_dict = {
model.Y: batch_y,
model.X: batch_x,
},
)
test_loss += cost
test_acc += acc
pbar.set_postfix(cost = cost, accuracy = acc)
train_loss /= len(train_input_ids) / batch_size
train_acc /= len(train_input_ids) / batch_size
test_loss /= len(test_input_ids) / batch_size
test_acc /= len(test_input_ids) / batch_size
if test_acc > CURRENT_ACC:
print(
'epoch: %d, pass acc: %f, current acc: %f'
% (EPOCH, CURRENT_ACC, test_acc)
)
CURRENT_ACC = test_acc
CURRENT_CHECKPOINT = 0
else:
CURRENT_CHECKPOINT += 1
print('time taken:', time.time() - lasttime)
print(
'epoch: %d, training loss: %f, training acc: %f, valid loss: %f, valid acc: %f\n'
% (EPOCH, train_loss, train_acc, test_loss, test_acc)
)
EPOCH += 1
real_Y, predict_Y = [], []
pbar = tqdm(
range(0, len(test_input_ids), batch_size), desc = 'validation minibatch loop'
)
for i in pbar:
index = min(i + batch_size, len(test_input_ids))
batch_x = test_input_ids[i: index]
batch_y = test_Y[i: index]
predict_Y += np.argmax(sess.run(model.logits,
feed_dict = {
model.Y: batch_y,
model.X: batch_x,
},
), 1, ).tolist()
real_Y += batch_y
from sklearn import metrics
print(
metrics.classification_report(
real_Y, predict_Y, target_names = ['negative', 'positive'], digits = 6
)
)
strings = ','.join(
[
n.name
for n in tf.get_default_graph().as_graph_def().node
if ('Variable' in n.op
or 'Placeholder' in n.name
or 'logits' in n.name
or 'alphas' in n.name
or 'self/Softmax' in n.name)
and 'adam' not in n.name
and 'beta' not in n.name
and 'global_step' not in n.name
]
)
strings.split(',')
saver = tf.train.Saver(tf.trainable_variables())
saver.save(sess, 'bert-base-sentiment/model.ckpt')
def freeze_graph(model_dir, output_node_names):
if not tf.gfile.Exists(model_dir):
raise AssertionError(
"Export directory doesn't exists. Please specify an export "
'directory: %s' % model_dir
)
checkpoint = tf.train.get_checkpoint_state(model_dir)
input_checkpoint = checkpoint.model_checkpoint_path
absolute_model_dir = '/'.join(input_checkpoint.split('/')[:-1])
output_graph = absolute_model_dir + '/frozen_model.pb'
clear_devices = True
with tf.Session(graph = tf.Graph()) as sess:
saver = tf.train.import_meta_graph(
input_checkpoint + '.meta', clear_devices = clear_devices
)
saver.restore(sess, input_checkpoint)
output_graph_def = tf.graph_util.convert_variables_to_constants(
sess,
tf.get_default_graph().as_graph_def(),
output_node_names.split(','),
)
with tf.gfile.GFile(output_graph, 'wb') as f:
f.write(output_graph_def.SerializeToString())
print('%d ops in the final graph.' % len(output_graph_def.node))
freeze_graph('bert-base-sentiment', strings)
import boto3
bucketName = 'huseinhouse-storage'
Key = 'bert-base-sentiment/frozen_model.pb'
outPutname = "v27/sentiment/bert-base-sentiment.pb"
s3 = boto3.client('s3',
aws_access_key_id='',
aws_secret_access_key='')
s3.upload_file(Key,bucketName,outPutname)
```
| github_jupyter |
```
%matplotlib inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
heart_df = pd.read_csv("data/heart-disease.csv")
heart_df.head() # classification dataset - supervised learning
```
## 1. Tuning hyperparameters by hand
so far we've worked with training and test datasets.
You train a model on a training set and evaluate it on a test dataset.
But hyperparameter tuning introduces a thrid set, **a validation set.**
Now the process becomes, **train a model on the training data, (try to) improve its hyperparameters on the validation set and evaluate it on the test set.**
```
from sklearn.ensemble import RandomForestClassifier
clf = RandomForestClassifier()
clf.get_params()
```
The parameters we are going to adjust (check documentation for definition)
* **max_depth** - the maximum depth of the tree
* **max_features** - the number of features to consider when looking for the best split
* **min_samples_leaf** - the minimum number of samples required to be at a leaf node
* **min_samples_split**
* **n_estimators** - the number of trees in the forest
```
# From 100 samples
# Train - 70, Validation - 15, Test - 15
```
#### Create an evaluation function for models
```
def evaluate_preds(y_true,y_preds):
"""
Performs evaluation comparison on y_true labels vs. y_pred labels
on a classification model.
"""
accuracy = accuracy_score(y_true,y_preds)
precision = precision_score(y_true,y_preds)
recall = recall_score(y_true,y_preds)
f1 = f1_score(y_true,y_preds)
metric_dict = {
"accuracy":round(accuracy,2),
"precision":round(precision,2),
"recall":round(recall,2),
"f1":round(f1,2)
} # A dictionary that stores the results of the evaluation metrics
print(f"Acc: {accuracy * 100:.2f}%")
print(f"Precision: {precision:.2f}")
print(f"Recall: {recall:.2f}")
print(f"F1 score: {f1:.2f}")
return metric_dict
len(heart_df)
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score, precision_score, recall_score, f1_score
from sklearn.ensemble import RandomForestClassifier
np.random.seed(42) # Results are reproducable
# Shuffle the data
heart_df_shuffle = heart_df.sample(frac=1)
# Split into X and y
X = heart_df_shuffle.drop("target",axis=1)
y = heart_df_shuffle["target"]
# Split the data into train, validation and test splits
# train - 70%, validation - 15%, test - 15%
train_split = round(0.7 * len(heart_df_shuffle)) # 70%
valid_split = round(train_split + 0.15 * len(heart_df_shuffle)) # index + next 15% of data
# [from:to]
X_train,y_train = X[:train_split],y[:train_split]
X_valid,y_valid = X[train_split:valid_split],y[train_split:valid_split]
X_test,y_test = X[valid_split:],y[valid_split:]
# len(X_train),len(X_valid),len(X_test)
# Train the model
clf = RandomForestClassifier() # instantiates with base line parameters
clf.fit(X_train, y_train)
# Make baseline predictions (on valid set)
y_preds = clf.predict(X_valid) # tune model on valid set
# Evaluate the classifier on validation set
baseline_metrics = evaluate_preds(y_valid, y_preds)
baseline_metrics
```
Beautiful, now let's try and improve the results.
We'll change 1 of the hyperparameters, n_estimators to 100 and see if it improves on the validation set.
```
np.random.seed(42)
# Create a second classifier with different hyperparameters
clf_2 = RandomForestClassifier(n_estimators=100) # adjusting n_estimators
clf_2.fit(X_train, y_train)
# Make predictions
y_preds_2 = clf_2.predict(X_valid)
# Evaluate the 2nd classifier
clf_2_metrics = evaluate_preds(y_valid, y_preds_2)
clf_2_metrics
# Different models on same data
```
How about we try another parameter?
Wait...
Building new models with new hyperparameters each time (by hand) is taking a lot of time.
Is there a better way?
Ans) **RandomizedSearchCV** provided by Sklearn
| github_jupyter |
# 6장. 알고리즘 체인과 파이프라인
*아래 링크를 통해 이 노트북을 주피터 노트북 뷰어(nbviewer.org)로 보거나 구글 코랩(colab.research.google.com)에서 실행할 수 있습니다.*
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://nbviewer.org/github/rickiepark/intro_ml_with_python_2nd_revised/blob/main/06-algorithm-chains-and-pipelines.ipynb"><img src="https://jupyter.org/assets/share.png" width="60" />주피터 노트북 뷰어로 보기</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/rickiepark/intro_ml_with_python_2nd_revised/blob/main/06-algorithm-chains-and-pipelines.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />구글 코랩(Colab)에서 실행하기</a>
</td>
</table>
<b><font size=2>이 노트북은 맷플롯립 그래프에 한글을 쓰기 위해 나눔 폰트를 사용합니다. 컴퓨터에 나눔 폰트가 없다면 설치해 주세요.<br><br><font color='red'>주의: 코랩에서 실행하는 경우 아래 셀을 실행하고 ⌘+M . 또는 Ctrl+M . 을 눌러 런타임을 재시작한 다음 처음부터 다시 실행해 주세요.</font></b>
```
# 노트북이 코랩에서 실행 중인지 체크합니다.
import os
import sys
if 'google.colab' in sys.modules and not os.path.isdir('mglearn'):
# 사이킷런 최신 버전을 설치합니다.
!pip install -q --upgrade scikit-learn
# mglearn을 다운받고 압축을 풉니다.
!wget -q -O mglearn.tar.gz https://bit.ly/mglearn-tar-gz
!tar -xzf mglearn.tar.gz
# 나눔 폰트를 설치합니다.
!sudo apt-get -qq -y install fonts-nanum
import matplotlib.font_manager as fm
fm._rebuild()
import sklearn
from preamble import *
import matplotlib
# 나눔 폰트를 사용합니다.
matplotlib.rc('font', family='NanumBarunGothic')
matplotlib.rcParams['axes.unicode_minus'] = False
from sklearn.svm import SVC
from sklearn.datasets import load_breast_cancer
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import MinMaxScaler
# 데이터 적재와 분할
cancer = load_breast_cancer()
X_train, X_test, y_train, y_test = train_test_split(
cancer.data, cancer.target, random_state=0)
# 훈련 데이터의 최솟값, 최댓값을 계산합니다
scaler = MinMaxScaler().fit(X_train)
# 훈련 데이터의 스케일을 조정합니다
X_train_scaled = scaler.transform(X_train)
svm = SVC()
# 스케일 조정된 훈련데이터에 SVM을 학습시킵니다
svm.fit(X_train_scaled, y_train)
# 테스트 데이터의 스케일을 조정하고 점수를 계산합니다
X_test_scaled = scaler.transform(X_test)
print("테스트 점수: {:.2f}".format(svm.score(X_test_scaled, y_test)))
```
## 6.1 데이터 전처리와 매개변수 선택
```
from sklearn.model_selection import GridSearchCV
# 이 코드는 예를 위한 것입니다. 실제로 사용하지 마세요.
param_grid = {'C': [0.001, 0.01, 0.1, 1, 10, 100],
'gamma': [0.001, 0.01, 0.1, 1, 10, 100]}
grid = GridSearchCV(SVC(), param_grid=param_grid, cv=5)
grid.fit(X_train_scaled, y_train)
print("최상의 교차 검증 정확도: {:.2f}".format(grid.best_score_))
print("테스트 점수: {:.2f}".format(grid.score(X_test_scaled, y_test)))
print("최적의 매개변수: ", grid.best_params_)
mglearn.plots.plot_improper_processing()
```
## 6.2 파이프라인 구축하기
```
from sklearn.pipeline import Pipeline
pipe = Pipeline([("scaler", MinMaxScaler()), ("svm", SVC())])
pipe.fit(X_train, y_train)
print("테스트 점수: {:.2f}".format(pipe.score(X_test, y_test)))
```
## 6.3 그리드 서치에 파이프라인 적용하기
```
param_grid = {'svm__C': [0.001, 0.01, 0.1, 1, 10, 100],
'svm__gamma': [0.001, 0.01, 0.1, 1, 10, 100]}
grid = GridSearchCV(pipe, param_grid=param_grid, cv=5)
grid.fit(X_train, y_train)
print("최상의 교차 검증 정확도: {:.2f}".format(grid.best_score_))
print("테스트 세트 점수: {:.2f}".format(grid.score(X_test, y_test)))
print("최적의 매개변수:", grid.best_params_)
mglearn.plots.plot_proper_processing()
rnd = np.random.RandomState(seed=0)
X = rnd.normal(size=(100, 10000))
y = rnd.normal(size=(100,))
from sklearn.feature_selection import SelectPercentile, f_regression
select = SelectPercentile(score_func=f_regression, percentile=5).fit(X, y)
X_selected = select.transform(X)
print("X_selected.shape:", X_selected.shape)
from sklearn.model_selection import cross_val_score
from sklearn.linear_model import Ridge
print("교차 검증 점수 (릿지): {:.2f}".format(
np.mean(cross_val_score(Ridge(), X_selected, y, cv=5))))
pipe = Pipeline([("select", SelectPercentile(score_func=f_regression,
percentile=5)),
("ridge", Ridge())])
print("교차 검증 점수 (파이프라인): {:.2f}".format(
np.mean(cross_val_score(pipe, X, y, cv=5))))
```
## 6.4 파이프라인 인터페이스
```
def fit(self, X, y):
X_transformed = X
for name, estimator in self.steps[:-1]:
# 마지막 단계를 빼고 fit과 transform을 반복합니다
X_transformed = estimator.fit_transform(X_transformed, y)
# 마지막 단계 fit을 호출합니다
self.steps[-1][1].fit(X_transformed, y)
return self
def predict(self, X):
X_transformed = X
for step in self.steps[:-1]:
# 마지막 단계를 빼고 transform을 반복합니다
X_transformed = step[1].transform(X_transformed)
# 마지막 단계 predict을 호출합니다
return self.steps[-1][1].predict(X_transformed)
```
#### 파이프라인 그리기
```
from sklearn import set_config
set_config(display='diagram')
pipe
```
### 6.4.1 `make_pipleline`을 사용한 파이프라인 생성
```
from sklearn.pipeline import make_pipeline
# 표준적인 방법
pipe_long = Pipeline([("scaler", MinMaxScaler()), ("svm", SVC(C=100))])
# 간소화된 방법
pipe_short = make_pipeline(MinMaxScaler(), SVC(C=100))
print("파이프라인 단계:\n", pipe_short.steps)
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
pipe = make_pipeline(StandardScaler(), PCA(n_components=2), StandardScaler())
print("파이프라인 단계:\n", pipe.steps)
```
### 6.4.2 단계 속성에 접근하기
```
# cancer 데이터셋에 앞서 만든 파이프라인을 적용합니다
pipe.fit(cancer.data)
# "pca" 단계의 두 개 주성분을 추출합니다
components = pipe.named_steps["pca"].components_
print("components.shape:", components.shape)
```
### 6.4.3 그리드 서치 안의 파이프라인의 속성에 접근하기
```
from sklearn.linear_model import LogisticRegression
pipe = make_pipeline(StandardScaler(), LogisticRegression(max_iter=1000))
param_grid = {'logisticregression__C': [0.01, 0.1, 1, 10, 100]}
X_train, X_test, y_train, y_test = train_test_split(
cancer.data, cancer.target, random_state=4)
grid = GridSearchCV(pipe, param_grid, cv=5)
grid.fit(X_train, y_train)
print("최상의 모델:\n", grid.best_estimator_)
print("로지스틱 회귀 단계:\n",
grid.best_estimator_.named_steps["logisticregression"])
print("로지스틱 회귀 계수:\n",
grid.best_estimator_.named_steps["logisticregression"].coef_)
```
## 6.5 전처리와 모델의 매개변수를 위한 그리드 서치
```
# 보스턴 주택 데이터셋이 1.0 버전에 deprecated 되었고 1.2 버전에서 삭제됩니다.
# 경고 메시지를 피하기 위해 다음 코드를 추가합니다.
import warnings
warnings.filterwarnings("ignore", category=FutureWarning)
from sklearn.datasets import load_boston
boston = load_boston()
X_train, X_test, y_train, y_test = train_test_split(boston.data, boston.target,
random_state=0)
from sklearn.preprocessing import PolynomialFeatures
pipe = make_pipeline(
StandardScaler(),
PolynomialFeatures(),
Ridge())
param_grid = {'polynomialfeatures__degree': [1, 2, 3],
'ridge__alpha': [0.001, 0.01, 0.1, 1, 10, 100]}
grid = GridSearchCV(pipe, param_grid=param_grid, cv=5, n_jobs=-1)
grid.fit(X_train, y_train)
mglearn.tools.heatmap(grid.cv_results_['mean_test_score'].reshape(3, -1),
xlabel="ridge__alpha", ylabel="polynomialfeatures__degree",
xticklabels=param_grid['ridge__alpha'],
yticklabels=param_grid['polynomialfeatures__degree'], vmin=0)
plt.show() # 책에는 없음
print("최적의 매개변수:", grid.best_params_)
print("테스트 세트 점수: {:.2f}".format(grid.score(X_test, y_test)))
param_grid = {'ridge__alpha': [0.001, 0.01, 0.1, 1, 10, 100]}
pipe = make_pipeline(StandardScaler(), Ridge())
grid = GridSearchCV(pipe, param_grid, cv=5)
grid.fit(X_train, y_train)
print("다항 특성이 없을 때 점수: {:.2f}".format(grid.score(X_test, y_test)))
```
## 6.6 모델 선택을 위한 그리드 서치
```
pipe = Pipeline([('preprocessing', StandardScaler()), ('classifier', SVC())])
from sklearn.ensemble import RandomForestClassifier
param_grid = [
{'classifier': [SVC()], 'preprocessing': [StandardScaler()],
'classifier__gamma': [0.001, 0.01, 0.1, 1, 10, 100],
'classifier__C': [0.001, 0.01, 0.1, 1, 10, 100]},
{'classifier': [RandomForestClassifier(n_estimators=100)],
'preprocessing': [None], 'classifier__max_features': [1, 2, 3]}]
X_train, X_test, y_train, y_test = train_test_split(
cancer.data, cancer.target, random_state=0)
grid = GridSearchCV(pipe, param_grid, cv=5)
grid.fit(X_train, y_train)
print("최적의 매개변수:\n{}\n".format(grid.best_params_))
print("최상의 교차 검증 점수: {:.2f}".format(grid.best_score_))
print("테스트 세트 점수: {:.2f}".format(grid.score(X_test, y_test)))
```
### 6.6.1 중복 계산 피하기
```
pipe = Pipeline([('preprocessing', StandardScaler()), ('classifier', SVC())],
memory="cache_folder")
```
## 6.7 요약 및 정리
| github_jupyter |
# Mixture Density Network for Regression
```
import nbloader,os,warnings
warnings.filterwarnings("ignore")
import numpy as np
import matplotlib.pyplot as plt
import scipy.io as sio
import tensorflow as tf
import tensorflow.contrib.slim as slim
from sklearn.utils import shuffle
from util import gpusession,create_gradient_clipping,data4reg,plot_1dRegData,print_n_txt
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
if __name__ == "__main__":
print ("TensorFlow version is [%s]."%(tf.__version__))
```
### Define MDN Class
```
class mdn_reg_class(object):
def __init__(self,_name='MDN',_xdim=1,_ydim=1,_hdims=[64,64],_sigmax=0
,_kmix=5,_actv=tf.nn.relu,_bn=slim.batch_norm
,_l2_reg_coef=1e-5,_GPU_ID=0,_VERBOSE=True):
self.name = _name
self.xdim = _xdim
self.ydim = _ydim
self.hdims = _hdims
self.sigmax = _sigmax
self.kmix = _kmix
self.actv = _actv
self.bn = _bn # slim.batch_norm / None
self.l2_reg_coef = _l2_reg_coef # L2 regularizer
self.GPU_ID = _GPU_ID
self.VERBOSE = _VERBOSE
with tf.device('/device:GPU:%d'%(self.GPU_ID)):
# Build model
self.build_model()
# Build graph
self.build_graph()
# Check parameters
self.check_params()
def build_model(self):
self.x = tf.placeholder(dtype=tf.float32,shape=[None,self.xdim]) # Input [N x xdim]
self.t = tf.placeholder(dtype=tf.float32,shape=[None,self.ydim]) # Output [N x ydim]
self.kp = tf.placeholder(dtype=tf.float32,shape=[]) # Keep probability
self.lr = tf.placeholder(dtype=tf.float32,shape=[]) # Learning rate
self.is_training = tf.placeholder(dtype=tf.bool,shape=[]) # Training flag
# Initializers
self.fully_init = tf.random_normal_initializer(stddev=0.01)
self.bias_init = tf.constant_initializer(0.)
self.bn_init = {'beta': tf.constant_initializer(0.),
'gamma': tf.random_normal_initializer(1., 0.01)}
self.bn_params = {'is_training':self.is_training,'decay':0.9,'epsilon':1e-5,
'param_initializers':self.bn_init,'updates_collections':None}
# Build graph
with tf.variable_scope(self.name,reuse=False) as scope:
with slim.arg_scope([slim.fully_connected],activation_fn=self.actv,
weights_initializer=self.fully_init,biases_initializer=self.bias_init,
normalizer_fn=self.bn,normalizer_params=self.bn_params,
weights_regularizer=None):
_net = self.x
for h_idx in range(len(self.hdims)): # Loop over hidden layers
_hdim = self.hdims[h_idx]
_net = slim.fully_connected(_net,_hdim,scope='lin'+str(h_idx))
_net = slim.dropout(_net,keep_prob=self.kp,is_training=self.is_training
,scope='dr'+str(h_idx))
self.feat = _net # [N x Q]
# Class allocation probability
self.pi_logits = slim.fully_connected(self.feat,self.kmix,scope='pi_logits')
self.pi = tf.nn.softmax(self.pi_logits,dim=1)
# means (data x dim x mixture)
self._mu = slim.linear(self.feat,self.kmix*self.ydim,scope='mu_flatten'
,biases_initializer=tf.random_uniform_initializer(minval=-2,maxval=+2))
self.mu = tf.reshape(self._mu,shape=[-1,self.ydim,self.kmix])
# varainces (data x dim x mixture)
self.sigma_logits = slim.fully_connected(self.feat,self.kmix*self.ydim,scope='sigma_logits')
if self.sigmax == 0:
self._sigma =tf.exp(self.sigma_logits)
else:
self._sigma = self.sigmax*tf.nn.sigmoid(self.sigma_logits)
# _sigma = tf.exp(_sigma_logits)
self.sigma = tf.reshape(self._sigma,shape=[-1,self.ydim,self.kmix]) # [N x D x K]
def build_graph(self):
y = self.t
pi = self.pi
mu = self.mu
sigma = self.sigma
yrepeat = tf.tile(y[:,:,tf.newaxis],[1,1,self.kmix]) # (N x D x K)
self.quadratics = -0.5*tf.reduce_sum(((yrepeat-mu)/sigma)**2,axis=1) # (N x K)
self.logdet = -0.5*tf.reduce_sum(tf.log(sigma),axis=1) # (N x K)
self.logconstant = - 0.5*self.ydim*tf.log(2*np.pi) # (1)
self.logpi = tf.log(pi) # (N x K)
self.exponents = self.quadratics + self.logdet + self.logconstant + self.logpi
self.logprobs = tf.reduce_logsumexp(self.exponents,axis=1) # (N)
self.gmm_prob = tf.exp(self.logprobs) # (N)
self.gmm_nll = -tf.reduce_mean(self.logprobs) # (1)
# Weight decay
# _g_vars = tf.global_variables()
_g_vars = tf.trainable_variables()
self.c_vars = [var for var in _g_vars if '%s/'%(self.name) in var.name]
self.l2_reg = self.l2_reg_coef*tf.reduce_sum(tf.stack([tf.nn.l2_loss(v) for v in self.c_vars])) # [1]
# Loss and optimizer
self.loss_total = self.gmm_nll + self.l2_reg
USE_ADAM = False
if USE_ADAM:
self.optm = tf.train.AdamOptimizer(learning_rate=self.lr
, beta1=0.9, beta2=0.999, epsilon=1e-0).minimize(self.loss_total)
else:
self.optm = tf.train.MomentumOptimizer(learning_rate=self.lr
,momentum=0.0).minimize(self.loss_total)
# Compute regression loss
_N = tf.shape(self.x)[0] # Number of data
maxIdx = tf.argmax(input=pi,axis=1, output_type=tf.int32) # Argmax Index [N]
# maxIdx = 0*tf.ones_like(maxIdx) # <== ???
coords = tf.stack([tf.transpose(gv) for gv in tf.meshgrid(tf.range(_N),tf.range(self.ydim))] +
[tf.reshape(tf.tile(maxIdx[:,tf.newaxis],[1,self.ydim]),shape=(_N,self.ydim))]
,axis=2) # [N x D x 3]
self.mu_bar = tf.gather_nd(mu,coords) # [N x D]
self.fit_mse = tf.reduce_sum(tf.pow(self.mu_bar-self.t, 2))/(tf.cast(_N,tf.float32)) # Fitting error (mse)
# Check parameters
def check_params(self):
_g_vars = tf.global_variables()
self.g_vars = [var for var in _g_vars if '%s/'%(self.name) in var.name]
if self.VERBOSE:
print ("==== Global Variables ====")
for i in range(len(self.g_vars)):
w_name = self.g_vars[i].name
w_shape = self.g_vars[i].get_shape().as_list()
if self.VERBOSE:
print (" [%02d] Name:[%s] Shape:[%s]" % (i,w_name,w_shape))
# Sampler
def sampler(self,_sess,_x,n_samples=10,_USE_ARGMAX=False,_MU_ONLY=False):
pi, mu, sigma = _sess.run([self.pi, self.mu, self.sigma],
feed_dict={self.x:_x,self.kp:1.0,self.is_training:False})
n_points = _x.shape[0]
_y_sampled = np.zeros([n_points,self.ydim,n_samples])
for i in range(n_points):
for j in range(n_samples):
if _USE_ARGMAX: # Use the most likely mixture
k = np.argmax(pi[i,:])
else:
k = np.random.choice(self.kmix,p=pi[i,:])
if _MU_ONLY: # Plot mu only
_y_sampled[i,:,j] = mu[i,:,k]
else: # Sample from Gaussian
_y_sampled[i,:,j] = mu[i,:,k] + np.random.randn(1,self.ydim)*np.sqrt(sigma[i,:,k])
return _y_sampled
# Save
def save(self,_sess,_savename=None):
""" Save name """
if _savename==None:
_savename='../net/net_%s.npz'%(self.name)
""" Get global variables """
self.g_wnames,self.g_wvals,self.g_wshapes = [],[],[]
for i in range(len(self.g_vars)):
curr_wname = self.g_vars[i].name
curr_wvar = [v for v in tf.global_variables() if v.name==curr_wname][0]
curr_wval = _sess.run(curr_wvar)
curr_wval_sqz = curr_wval.squeeze()
self.g_wnames.append(curr_wname)
self.g_wvals.append(curr_wval_sqz)
self.g_wshapes.append(curr_wval.shape)
""" Save """
np.savez(_savename,g_wnames=self.g_wnames,g_wvals=self.g_wvals,g_wshapes=self.g_wshapes)
if self.VERBOSE:
print ("[%s] Saved. Size is [%.4f]MB" %
(_savename,os.path.getsize(_savename)/1000./1000.))
# Save
def save_final(self,_sess,_savename=None):
""" Save name """
if _savename==None:
_savename='../net/net_%s_final.npz'%(self.name)
""" Get global variables """
self.g_wnames,self.g_wvals,self.g_wshapes = [],[],[]
for i in range(len(self.g_vars)):
curr_wname = self.g_vars[i].name
curr_wvar = [v for v in tf.global_variables() if v.name==curr_wname][0]
curr_wval = _sess.run(curr_wvar)
curr_wval_sqz = curr_wval.squeeze()
self.g_wnames.append(curr_wname)
self.g_wvals.append(curr_wval_sqz)
self.g_wshapes.append(curr_wval.shape)
""" Save """
np.savez(_savename,g_wnames=self.g_wnames,g_wvals=self.g_wvals,g_wshapes=self.g_wshapes)
print ("[%s] Saved. Size is [%.4f]MB" %
(_savename,os.path.getsize(_savename)/1000./1000.))
# Restore
def restore(self,_sess,_loadname=None):
if _loadname==None:
_loadname='../net/net_%s_final.npz'%(self.name)
l = np.load(_loadname)
g_wnames = l['g_wnames']
g_wvals = l['g_wvals']
g_wshapes = l['g_wshapes']
for widx,wname in enumerate(g_wnames):
curr_wvar = [v for v in tf.global_variables() if v.name==wname][0]
_sess.run(tf.assign(curr_wvar,g_wvals[widx].reshape(g_wshapes[widx])))
if self.VERBOSE:
print ("Weight restored from [%s] Size is [%.4f]MB" %
(_loadname,os.path.getsize(_loadname)/1000./1000.))
# Save to mat file
def save2mat(self,_xdata='',_ydata='',_yref=''):
# Save weights to mat file so that MATLAB can use it.
npzPath = '../net/net_%s.npz'%(self.name)
l = np.load(npzPath)
g_wnames = l['g_wnames']
g_wvals = l['g_wvals']
g_wshapes = l['g_wshapes']
D = {}
for widx,wname in enumerate(g_wnames):
cName = wname.replace(':0','')
cName = cName.replace(self.name+'/','')
cName = cName.replace('/','_')
cVal = g_wvals[widx].reshape(g_wshapes[widx])
D[cName] = cVal
# Do not print out..
# if self.VERBOSE: print ("name is [%s] shape is %s."%(cName,cVal.shape,))
# Save data
if _xdata!='': D['xdata']=_xdata
if _ydata!='': D['ydata']=_ydata
if _yref!='': D['yref']=_yref
# Save dictionary D to the mat file
matPath = '../data/net_%s.mat'%(self.name)
sio.savemat(matPath,D)
if self.VERBOSE: print ("[%s] saved."%(matPath))
# Save to mat file
def save2mat_final(self,_xdata='',_ydata='',_yref=''):
# Save weights to mat file so that MATLAB can use it.
npzPath = '../net/net_%s_final.npz'%(self.name)
l = np.load(npzPath)
g_wnames = l['g_wnames']
g_wvals = l['g_wvals']
g_wshapes = l['g_wshapes']
D = {}
for widx,wname in enumerate(g_wnames):
cName = wname.replace(':0','')
cName = cName.replace(self.name+'/','')
cName = cName.replace('/','_')
cVal = g_wvals[widx].reshape(g_wshapes[widx])
D[cName] = cVal
if self.VERBOSE: print ("name is [%s] shape is %s."%(cName,cVal.shape,))
# Save data
if _xdata!='': D['xdata']=_xdata
if _ydata!='': D['ydata']=_ydata
if _yref!='': D['yref']=_yref
# Save dictionary D to the mat file
matPath = '../data/net_%s_final.mat'%(self.name)
sio.savemat(matPath,D)
print ("[%s] Saved. Size is [%.4f]MB" %
(matPath,os.path.getsize(matPath)/1000./1000.))
# Train
def train(self,_sess,_x,_y,_yref='',_lr=1e-3,_batchSize=512,_maxEpoch=1e4,_kp=1.0
,_LR_SCHEDULE=True
,_PRINT_EVERY=20,_PLOT_EVERY=20
,_SAVE_TXT=True,_SAVE_BEST_NET=True,_SAVE_FINAL=True):
# Reference training data
_x_train,_y_train = _x,_y
# Iterate
if _PRINT_EVERY == 0: print_period = 0
else: print_period = _maxEpoch//_PRINT_EVERY
if _PLOT_EVERY == 0: plot_period = 0
else: plot_period = _maxEpoch//_PLOT_EVERY
maxIter = max(_x_train.shape[0]//_batchSize, 1)
bestLossVal = np.inf
if _SAVE_TXT:
txtName = ('../res/res_%s.txt'%(self.name));f = open(txtName,'w') # Open txt file
print_n_txt(_f=f,_chars='Text name: '+txtName,_DO_PRINT=True)
for epoch in range((int)(_maxEpoch)+1): # For every epoch
_x_train,_y_train = shuffle(_x_train,_y_train)
for iter in range(maxIter): # For every iteration
start,end = iter*_batchSize,(iter+1)*_batchSize
if _LR_SCHEDULE:
if epoch < 0.5*_maxEpoch:
lr_use = _lr
elif epoch < 0.75*_maxEpoch:
lr_use = _lr/5.
else:
lr_use = _lr/10.
else:
lr_use = _lr
feeds = {self.x:_x_train[start:end,:],self.t:_y_train[start:end,:]
,self.kp:_kp,self.lr:lr_use,self.is_training:True}
# Optimize
_sess.run(self.optm,feeds)
# Track the Best result
BEST_FLAG = False
check_period = _maxEpoch//100
if (epoch%check_period)==0:
feeds = {self.x:_x,self.t:_y,self.kp:1.0,self.is_training:False}
opers = [self.loss_total,self.gmm_nll,self.l2_reg]
lossVal,gmm_nll,l2_reg = _sess.run(opers,feeds)
if (lossVal < bestLossVal) & (epoch >= 3):
bestLossVal = lossVal
BEST_FLAG = True
if _SAVE_BEST_NET:
self.save(_sess) # Save the current best model
self.save2mat(_xdata=_x,_ydata=_y,_yref=_yref)
# Print current result
if (print_period!=0) and ((epoch%print_period)==0 or (epoch==(_maxEpoch-1))): # Print
feeds = {self.x:_x,self.t:_y,self.kp:1.0,self.is_training:False}
opers = [self.loss_total,self.gmm_nll,self.l2_reg]
lossVal,gmm_nll,l2_reg = _sess.run(opers,feeds)
if _SAVE_TXT:
strTemp = ("[%d/%d] loss:%.3f(gmm:%.3f+l2:%.3f) bestLoss:%.3f"
%(epoch,_maxEpoch,lossVal,gmm_nll,l2_reg,bestLossVal))
print_n_txt(_f=f,_chars=strTemp,_DO_PRINT=self.VERBOSE)
else:
if self.VERBOSE:
print ("[%d/%d] loss:%.3f(gmm:%.3f+l2:%.3f) bestLoss:%.3f"
%(epoch,_maxEpoch,lossVal,gmm_nll,l2_reg,bestLossVal))
# Plot current result
if (plot_period!=0) and ((epoch%plot_period)==0 or (epoch==(_maxEpoch-1))): # Plot
feeds = {self.x:_x,self.t:_y,self.kp:1.0,self.is_training:False}
opers = [self.loss_total,self.gmm_nll,self.l2_reg]
lossVal,gmm_nll,l2_reg = _sess.run(opers,feeds)
# Plot sampled outputs
nSample = 3
ytest = self.sampler(_sess=_sess,_x=_x,n_samples=nSample
,_USE_ARGMAX=False,_MU_ONLY=False)
x_plot,y_plot = _x[:,0],_y[:,0] # Traning data
plt.figure(figsize=(8,4));
plt.axis([np.min(x_plot),np.max(x_plot),np.min(y_plot)-0.1,np.max(y_plot)+0.1])
if _yref != '': plt.plot(x_plot,_yref[:,0],'r.') # Plot reference
plt.plot(x_plot,y_plot,'k.') # Plot training data
for i in range(nSample):
plt.plot(_x,ytest[:,0,i],'b.')
plt.title("[%d/%d] name:[%s] lossVal:[%.3e]"%(epoch,_maxEpoch,self.name,lossVal))
# Plot most-likely mean function
ytest = self.sampler(_sess=_sess,_x=_x,n_samples=1
,_USE_ARGMAX=True,_MU_ONLY=True)
x_plot,y_plot = _x[:,0],_y[:,0] # Traning data
plt.figure(figsize=(8,4));
plt.axis([np.min(x_plot),np.max(x_plot),np.min(y_plot)-0.1,np.max(y_plot)+0.1])
if _yref != '': plt.plot(x_plot,_yref[:,0],'r.') # Plot reference
plt.plot(x_plot,y_plot,'k.') # Plot training data
plt.plot(_x,ytest[:,0,0],'b-')
plt.title("[%d/%d] name:[%s] lossVal:[%.3e]"%(epoch,_maxEpoch,self.name,lossVal))
plt.show()
# Save fianl weights
if _SAVE_FINAL:
self.save_final(_sess)
self.save2mat_final(_xdata=_x,_ydata=_y,_yref=_yref)
# Test
def test(self,_sess,_xdata,_ydata,_yref,_xtest
,_titleStr,_PLOT_TRAIN=True,_PLOT_RES=True,_SAVE_FIG=False):
nSample = 1
ytest = self.sampler(_sess=_sess,_x=_xtest,n_samples=nSample
,_USE_ARGMAX=True,_MU_ONLY=True)
# Plot
if _PLOT_TRAIN:
plt.figure(figsize=(6,4))
plt.axis([np.min(_xdata),np.max(_xdata),np.min(_ydata),np.max(_ydata)])
plt.plot(_xdata,_ydata,'k.')
plt.xlabel('Input',fontsize=13);plt.ylabel('Output',fontsize=13)
plt.title('Training Data for a Regression Task',fontsize=16);
if _SAVE_FIG:
plt.savefig('../fig/fig_%s_data.png'%(self.name)); plt.show()
else:
plt.show()
# Plot
if _PLOT_RES:
fig = plt.figure(figsize=(6,4))
plt.axis([np.min(_xdata),np.max(_xdata),np.min(_ydata),np.max(_ydata)])
ht,=plt.plot(_xdata,_yref,'r.');
hd,=plt.plot(_xdata,_ydata,'k.')
for i in range(nSample):
hf,=plt.plot(_xtest,ytest[:,0,i],'b-')
plt.xlabel('Input',fontsize=13);plt.ylabel('Output',fontsize=13)
plt.title('%s'%(_titleStr),fontsize=16)
plt.legend([ht,hd,hf],['Target function','Training data','Fitting result']
,fontsize=15,loc='upper left')
if _SAVE_FIG:
plt.savefig('../fig/fig_%s_res.png'%(self.name)); plt.show()
else:
plt.show()
if __name__ == "__main__":
print ("mdn_reg_class defined.")
```
### Train MDN for Regression
```
if __name__ == "__main__":
# Training data
dataType = 'cosexp' # ['cosexp','linear','step']
oRate = 0.4
measVar = 1e-8
x,y,t=data4reg(_type=dataType,_n=1000,_oRange=[-1.0,+3.0],_oRate=oRate,measVar=measVar)
xtest = np.linspace(start=-3,stop=3,num=1000).reshape((-1,1))
# plot_1dRegData(_x=x,_y=y,_t=t,_type='Training data [%s] function'%(dataType),_figSize=(8,4))
# Make graph
tf.reset_default_graph(); sess = gpusession()
tf.set_random_seed(0); np.random.seed(0)
MDN = mdn_reg_class(_name='MDN_%s_oRate%02d_var%.1e'%(dataType,oRate*100,measVar)
,_xdim=1,_ydim=1,_hdims=[32,32],_sigmax=1
,_kmix=5,_actv=tf.nn.relu,_bn=slim.batch_norm
,_l2_reg_coef=1e-5,_GPU_ID=0,_VERBOSE=False)
sess.run(tf.global_variables_initializer()) # Initialize variables
# Train
DO_TRAIN = True
if DO_TRAIN:
MDN.train(_sess=sess,_x=x,_y=y,_yref=t
,_lr=1e-3,_batchSize=256,_maxEpoch=1e4,_kp=1.0
,_LR_SCHEDULE=True
,_PRINT_EVERY=10,_PLOT_EVERY=10
,_SAVE_TXT=True,_SAVE_BEST_NET=True)
print ("Train done.")
else:
MDN.restore(sess)
print ("Network restored.")
# Test
MDN.test(_sess=sess,_xdata=x,_ydata=y,_yref=t,_xtest=xtest
,_titleStr='Final Best Result'
,_PLOT_TRAIN=True,_PLOT_RES=True,_SAVE_FIG=True)
```
| github_jupyter |
# Ensembles notebook
<a href="https://mybinder.org/v2/gh/tinkoff-ai/etna/master?filepath=examples/ensembles.ipynb">
<img src="https://mybinder.org/badge_logo.svg" align='left'>
</a>
This notebook contains the simple examples of using the ensemble models with ETNA library.
**Table of Contents**
* [Load Dataset](#chapter1)
* [Build Pipelines](#chapter2)
* [Ensembles](#chapter3)
* [VotingEnsemble](#section_3_1)
* [StackingEnsamble](#section_3_2)
* [Results](#section_3_3)
```
import warnings
warnings.filterwarnings("ignore")
```
## 1. Load Dataset <a class="anchor" id="chapter1"></a>
In this notebook we will work with the dataset contains only one segment with monthly wine sales. Working process with the dataset containing more segments will be absolutely the same.
```
import pandas as pd
from etna.datasets import TSDataset
original_df = pd.read_csv("data/monthly-australian-wine-sales.csv")
original_df["timestamp"] = pd.to_datetime(original_df["month"])
original_df["target"] = original_df["sales"]
original_df.drop(columns=["month", "sales"], inplace=True)
original_df["segment"] = "main"
original_df.head()
df = TSDataset.to_dataset(original_df)
ts = TSDataset(df=df, freq="MS")
ts.plot()
```
## 2. Build Pipelines <a class="anchor" id="chapter2"></a>
Given the sales' history, we want to select the best model(pipeline) to forecast future sales.
```
from etna.pipeline import Pipeline
from etna.models import NaiveModel, SeasonalMovingAverageModel, CatBoostModelMultiSegment
from etna.transforms import LagTransform
from etna.metrics import MAE, MSE, SMAPE, MAPE
HORIZON = 3
N_FOLDS = 5
```
Let's build four pipelines using the different models
```
naive_pipeline = Pipeline(model=NaiveModel(lag=12), transforms=[], horizon=HORIZON)
seasonalma_pipeline = Pipeline(
model=SeasonalMovingAverageModel(window=5, seasonality=12), transforms=[], horizon=HORIZON
)
catboost_pipeline = Pipeline(
model=CatBoostModelMultiSegment(),
transforms=[LagTransform(lags=[6, 7, 8, 9, 10, 11, 12], in_column="target")],
horizon=HORIZON,
)
pipeline_names = ["naive", "moving average", "catboost"]
pipelines = [naive_pipeline, seasonalma_pipeline, catboost_pipeline]
```
And evaluate their performance on the backtest
```
metrics = []
for pipeline in pipelines:
metrics.append(
pipeline.backtest(
ts=ts, metrics=[MAE(), MSE(), SMAPE(), MAPE()], n_folds=N_FOLDS, aggregate_metrics=True, n_jobs=5
)[0].iloc[:, 1:]
)
metrics = pd.concat(metrics)
metrics.index = pipeline_names
metrics
```
## 3. Ensembles <a class="anchor" id="chapter3"></a>
To improve the performance of the individual models, we can try to make ensembles out of them. Our library contains two ensembling methods, which we will try on now.
### 3.1 VotingEnsemble<a class="anchor" id="section_3_1"></a>
`VotingEnsemble` forecasts future values with weighted averaging of it's `pipelines` forecasts.
```
from etna.ensembles import VotingEnsemble
```
By default, `VotingEnsemble` uses **uniform** weights for the pipelines' forecasts. However, you can specify the weights manually using the `weights` parameter. The higher weight the more you trust the base model.
*Note*: The `weights` are automatically normalized.
```
voting_ensemble = VotingEnsemble(pipelines=pipelines, weights=[1, 9, 4], n_jobs=4)
voting_ensamble_metrics = voting_ensemble.backtest(
ts=ts, metrics=[MAE(), MSE(), SMAPE(), MAPE()], n_folds=N_FOLDS, aggregate_metrics=True, n_jobs=2
)[0].iloc[:, 1:]
voting_ensamble_metrics.index = ["voting ensemble"]
voting_ensamble_metrics
```
### 3.2 StackingEnsemble<a class="anchor" id="section_3_2"></a>
`StackingEnsemble` forecasts future using the metamodel to combine the forecasts of it's `pipelines`.
```
from etna.ensembles import StackingEnsemble
```
By default, `StackingEnsemble` uses only the pipelines' forecasts as features for the `final_model`. However, you can specify the additional features using the `features_to_use` parameter. The following values are possible:
+ **None** - use only the pipelines' forecasts(default)
+ **List[str]** - use the pipelines' forecasts + features from the list
+ **"all"** - use all the available features
*Note:* It is possible to use only the features available for the base models.
```
stacking_ensemble_unfeatured = StackingEnsemble(pipelines=pipelines, n_folds=10, n_jobs=4)
stacking_ensamble_metrics = stacking_ensemble_unfeatured.backtest(
ts=ts, metrics=[MAE(), MSE(), SMAPE(), MAPE()], n_folds=N_FOLDS, aggregate_metrics=True, n_jobs=2
)[0].iloc[:, 1:]
stacking_ensamble_metrics.index = ["stacking ensemble"]
stacking_ensamble_metrics
```
In addition, it is also possible to specify the `final_model`. You can use any regression model with the sklearn interface for this purpose.
### 3.3 Results<a class="anchor" id="section_3_3"></a>
Finally, let's take a look at the results of our experiments
```
metrics = pd.concat(
[
metrics,
voting_ensamble_metrics,
stacking_ensamble_metrics
]
)
metrics
```
| github_jupyter |
# Calculation of the entropy for sources with and without memory
## Introduction
This tutorial will get you familiar with the calculation of the entropy associated with a given source. We start by recalling some definitions and fundamental results from the [Shannon's information theory](http://people.math.harvard.edu/~ctm/home/text/others/shannon/entropy/entropy.pdf). We will then move to consider some practical examples, focusing on sources generating grey scale natural images (given that the whole training is about image and video).
## Preliminary remarks
Consider a discrete source of information $S$ which emits symbols $a_i$ from an *alphabet* $A$ and according to a Probability Mass Function (PMF) $p_S$:
$$
\large
S = \{A, p_S\},
$$
where, $A$ is the set of symbols $\{a_0, a_1,\ldots,a_N\}$ and $p_S: A \rightarrow [0, 1]$. In other words, the source $S$ emits symbols from $A$ and the $i$-th symbol $a_i$ has a probability of being emitted $p_S(a_i) = p_i$. The number of symbols emitted by a source characterises its arity: sources emitting two symbols are *binary*, three symbols *ternary*, $n$ symbols *n-ary* and so on. For each symbol $a_i\in A$, its **information content** ($I(a_i)$) is defined as:
$$
\large
I(a_i) = \log_2\left(\frac{1}{p_S(s_i)}\right),\quad[bits]
$$
This information content can be thought as the minimum amount of bits required to encode $a_i$. Note that symbols less likely to appear will require more bits and viceversa. From the information content, the entropy of source $S$ is defined as the *expected* information content:
$$
\large
H(S) = E[I(a_i)] = \sum_{i=0}^N -p_S(a_i)\cdot\log_2(p_S(a_i))\quad[bits/symbol]
$$
The entropy measures the *average* number of bits needed to code the symbols emitted by $S$. The Shannon's noiseless source coding theorem, tells you that such an *average* number of bits is also the *lowest* possibly achievable by any coding scheme one can come up with. Another consideration that follows from the definition of entropy is the fact that source whose PMF is uniform will lead to an entropy equal to $\log_2(|A|)$, where $|\cdot|$ denotes the set's cardinality.
The source $S$ considered so far is said to be memoryless since each symbol is emitted irrespective to the symbols emitted earlier. Conversely, sources emitting symbols based on their previous output are said to be *with memory*. Images can be considered as instances emitted by a source with memory. In fact, the intensity value of adjacent pixels is likely to be correlated and indeed simple models (e.g. first order Auto-Regressive (AR(1)) stochastic processes) are often used to study the performance limit of some coding tools such as frequency transforms (e.g. by computing the transform gain). If a source has memory, then the amount of information needed to transmit a given symbol can be reduced based on the symbols already transmitted. Recalling again the example of images, knowing the value of pixels located at even positions (in raster scan order) can help to infer the value of the adjacent pixels located at the odd positions: this is the fundamental principle behind *predictive coding*. For sources with memory, one can compute the entropy associated with the transmission of symbols in light of what has been done in the past. To accomplish this task we will use the concept of *conditional entropy*. Consider two sources of information $X$ and $Y$ with conditional probability mass function $p_{Y|X}$, the conditional entropy $H(Y|X)$ can be computed as:
$$
\large
H(Y|X) = -\sum_{i,j}p(x_i,y_j)\cdot log_2(p_{Y|X}(y_i|x_j))\quad[bits/symbol],
$$
where $p(x_i, y_i)$ denotes the *joint* probability. We now have all ingredients ready to write some toy examples and put *the theory in practice*.
## Example 1: Calculation of the entropy of a memoryless source
As mentioned at the beginning of this tutorial, we will consider a particular source of information that generates grey scale natural images whereas each pixel is represented with 8 bits per pixel \[bpp\]. In particular, we will compute the entropy of such a source assuming that among the many images generated, the well known [*cameraman.tif*](https://homepages.cae.wisc.edu/~ece533/images/cameraman.tif) sample is one example. This will allow us to estimate the source's PMF which is the key ingredient to compute the Shannon's entropy. We start by loading the required Python packages (i.e. `cv2` and `numpy`) and then read the image in memory.
```
import cv2
import numpy as np
image = cv2.imread('../input-data/cameraman.tif', cv2.IMREAD_UNCHANGED)
```
We can visualise the image by using the function `imshow` from the the `matplotlib` package.
```
import matplotlib.pyplot as plt
plt.figure(figsize=(7, 7))
plt.imshow(image, cmap='gray')
plt.axis('off')
plt.title(f"Cameraman grey scale image with size {image.shape[0]}x{image.shape[1]}");
```
As mentioned above, we'll now estimate the source's PMF by computing the normalised frequency of each one of the 256 grey levels from the image's pixels. To do so we will use the `histogram` function from `numpy` which returns the normalised frequency of each one of the 256 gray levels over all images pixels. More information about this function is available [here](https://docs.scipy.org/doc/numpy/reference/generated/numpy.histogram.html)
```
p, bins = np.histogram(image, 256, [0, 255], density=True)
plt.bar(range(len(p)), p)
plt.xlabel('Gray level value', fontsize=16)
plt.ylabel('Normalised frequency', fontsize=16)
plt.grid()
plt.title('Normalised histogram for the cameraman.tif image', fontsize=16);
```
We note from the code cell above that the function `histogram` from `numpy` has the parameter `density` set to true. This will tell the function to return the normalised histogram. If not set, it will return the absolute histogram instead. We also note from the graph obtained above that the histogram is *bi-modal*, i.e. it has two peaks: these are associated with the man's black coat and the grey background. This result should also immediately suggest that the Shannon's entropy won't equate to 8 bits per pixels as the image pixels are currently represented. In other words, we can surely find a code which will require less than 8 bpp to represent this image. The question is now how many bits do we then need? The answer will be given by the computation of the entropy.
```
index_valid = np.where(p != 0)
H = -np.sum(np.multiply(p[index_valid], np.log2(p[index_valid])))
print(f"Estimated entropy for souce generating gray scale images: {H:.2f} bits per pixel [bpp]")
```
We first note that the value of the entropy confirms our observation above, i.e. that more bits can be squeezed to represent the image pixels given the bi-modal nature of the PMF estimated. Please also note the emphasis on the word *estimated*: we are estimating the PMF of our source from one instance (i.e. the *cameraman.tif* image), which we've assumed to be a fairly accurate representative of the data generated. In practice, a more accurate estimate of the PMF would be obtained, had more images been available.
In terms of the code used to compute the entropy, it is worth noting the use of `where` from `numpy` which returns an array with elements from $p$ where the boolean condition is `true`. Such check is required to purge from $p$ all those gray levels associated with zero frequency, since for them the logarithm operation is undefined.
## Example 2: Calculation of the entropy for a source with memory
We now want to go a step further and ask ourselves whether we can reduce more the number of bits required to represent image pixels by considering some sort of data correlation. We remember that if two sources $X$ and $Y$ are correlated, we can reduce the number of bits required to transmit (say) $Y$ by knowing $X$. More precisely, we can transmit the residual information associated with $Y$ which cannot be inferred from $X$. In this example, we'll consider again our initial source which generates grey scale images such as the *cameraman.tif*. To look for data correlation, let's derive from the original source two additional ones as follows:
* Source $X$ is the source generating all values of pixels located at even indexes in a raster scan order.
* Source $Y$ is the source generating all values of pixels located at odd indexes in a raster scan order.
Our original information source is generating grey scale *natural* images so one would expect that the value of pixels lying on even indexes is quite correlated with that of its adjacent odd indexes counterpart. To provide more evidence to such statement, we can plot a scatter diagram of a limited set of values for $X$ and $Y$ extracted from *cameraman.tif*.
```
image = image.flatten()
X = image[:image.size-1:2]
Y = image[1:image.size:2]
plt.plot(X[0:1000], Y[0:1000], 'o')
plt.xlabel('Pixel values for source X', fontsize=16)
plt.ylabel('Pixel values for source Y', fontsize=16)
plt.grid()
```
As may be noted, pairs of values from $X$ and $Y$ are aligned along a 45 degree straight line. Pairs would have been spread throughout the whole 2D scatter plot, had the image been associated with random noise (i.e. each pixel value is uncorrelated with the others). We notice few outliers in the scatter plot: these are associated with pixels lying at the image edges where a sudden transition of the values is likely to happen.
To measure how much information one needs to transmit for source $Y$ assuming that $X$ is somewhat known, we need to compute the conditional entropy $H(Y|X)$. We remember that the formula is as follows:
$$
\large
H(Y|X) = -\sum_{i,j}p(x_i,y_j)\cdot log_2(p_{Y|X}(y_i|x_j))
$$
From the formula above we note that we need two ingredients: the joint probability mass function $p(X,Y)$ and the conditional one $p(Y|X)$. The joint probability can easily be computed by considering all possible values source $X$ and $Y$ can emit: $[0, 255]$ in our case. Then, for each pair of values $(x_i, y_j)$ we can count how many times source $X$ has emitted value $x_i$ and source $Y$ value $y_j$. Accordingly, we declare a 2D array with size $256\times 256$ and initialise all values to zero. Then we jointly loop through all pixel values associated with $X$ and $Y$ and increment the cell in such a 2D array indexed by the value of the pair $(x_i, y_j)$. The following code cell implements this processing.
```
p_joint = np.zeros((256, 256))
for i in range(len(X)):
p_joint[X[i], Y[i]] += 1
p_joint = np.divide(p_joint, len(X))
```
The last statement in the previous code cell normalises the frequency of all pairs so to have an estimate of the joint pdf. To compute the conditional probability, we remember the formula from statistics at Uni:
$$
\large
P(Y|X) = \frac{P(X,Y)}{P(X)}
$$
where $P(X)$ is the marginal probability for source $X$. Given the joint probability $P(X,Y)$, the marginal for $X$ is simply the sum of all $p(x_i,y_j)$ where the value $x_i$ is fixed. We note that this corresponds to add up all elements from $P(X,Y)$ along a given row ($X$ represents the rows in the 2D array of joint probability we just built). Adding elements along a given direction in a 2D array is easily obtained via function `sum` from `numpy`.
```
p_marginal = np.sum(p_joint, 1)
p_marginal_ext = np.tile(p_marginal, (256, 1))
```
We note from the code above that the `sum` function will return a 1D array with 256 cells. However, from the definition of conditional probability given above, we need to divide each row of $P(X,Y)$ by the corresponding cell in $P(X)$ (i.e. the 1D array). This can be done using a for loop (not very efficient) or by element wise division where the 1D array associated with $P(X)$ is extended to a 2D using the function `tile` from `numpy` which simply extents an array along a given dimension. The conditional probability $P(Y|X)$ can now be computed as shown in the following code cell.
```
index_valid = np.where(p_marginal_ext != 0)
p_cond = np.zeros((256, 256))
p_cond[index_valid] = np.divide(p_joint[index_valid], p_marginal_ext[index_valid])
```
We can now compute the conditional entropy, using the formula we recalled in the preliminary remarks section:
```
index_valid = np.where(p_cond != 0)
H_cond = -np.sum(np.multiply(p_joint[index_valid], np.log2(p_cond[index_valid])))
print(f"Conditional entropy value H(Y|X): {H_cond:.2f} bpp")
```
We can now finally appreciate the reduction of bits required to transmit pixels generated by source $Y$ if the value for pixels generated by source $X$ is known.
## Concluding remarks
We shall wrap up this tutorial with some thoughts and considerations. The first one is on the last example, in particular on how one could realised a more practical coding scheme which jointly encodes the pixels values of pixels. A possible workflow could be as follows:
* By scanning the image pixels in raster scan order, consider all pairs of pixels at even and odd indexes and compute their joint histogram (i.e. use the values of the 2D array `p_joint` without dividing them by the total of pairs considered).
* Derive a Huffman code for these pairs where the most likely to appear pairs will be coded with shorter codewords. You can implement your own version of the Huffman's algorithm or use any of the Python's packages out there (e.g. the [Huffman](https://pypi.org/project/huffman/) one from `pypi`).
* Write the Huffman table in the bitstream along with codeword associated with each pair of pixels. The decoder would simply read the Huffman table and then start parsing the bitstream: whenever a codeword is matched, the corresponding pair of pixel values would be written in the buffer associated with the decoded image
The second remark worth sharing is about the value of the entropy associated with the *cameraman.tif* image. It was 7.03 bits per pixel. Despite this being less than 8, any coding scheme using an integer number of bits (Huffman coding is one of those) would still use 8 bits anyway. A coding scheme such as arithmetic coding could archieve such a theoretical limit at the cost of increased complexity.
As a third remark, we shall observe that this tutorial was about the fundamentals results of the Shannon's information theory, thus we focused on entropy and its conditional variant which gave us the tip that for images, where a degree of spatial correlation exists among pixels, better coding schemes can be devised. We didn't appraised another classical and simple coding scheme: ***run length encoding***. We should expected that also this one could do a good job in reducing the bits pixels.
Finally, on a more general level, we also observe that some of the considerations made here also hold for different types of signal. As an example audio samples show a high degree of correlation along the temporal dimension. Accordingly, we would expect that the conditional entropy will yield to lower values than the entropy of the memoreless version of the audio source.
| github_jupyter |
<a href="https://colab.research.google.com/github/clemencia/ML4PPGF_UERJ/blob/master/correlations.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
**Valores esperados, médias e variância**
**Valor esperado** ou média de x:
$\mu = E[x] = \int_{-\infty}^{\infty} x f(x) dx$
**Variância:**
$\mu = E[(x - \mu)^{2}] = \int_{-\infty}^{+\infty} a(x) f(x) dx \Rightarrow V = E[(x - \mu)^{2}] = E[x^{2}] - \mu^{2}$
**Desvio padrão** de x:
$\sigma_x = \sqrt{E[(x - \mu)^{2}]}$
**Covariância:**
$V_{xy}= E[(x - \mu_{x})(y - \mu_y)] = \int_{-\infty}^{+\infty} \int_{-\infty}^{+\infty} (x - \mu_{x})(y - \mu_y) f(x,y) dxdy \\ \Rightarrow V_{xy}= E[xy] - \mu_x \mu_y$
**Coeficiente de correlação:**
$\rho_{xy} = \frac{V_{xy}}{\sigma_x\sigma_y}$
Para variáveis independentes,
$V_{xy} = 0$
No exemplo abaixo, plotamos duas variáveis **não correlacionadas** e calculamos o coeficiente de correlação.
```
import matplotlib.pyplot as plt
import random
import numpy as np
# random seed for reproducibility
##random.seed(9823767454)
# lists of random numbers:
mu = 4.
sigma = 1.
x = [random.gauss(mu, sigma) for i in range(10000)]
y = [random.gauss(mu, sigma) for i in range(10000)]
# convert list to numpy array for easy manipulation
x = np.array(x)
y = np.array(y)
plt.plot(x,y,'o')
plt.xlabel('x')
plt.ylabel('y')
x_mean = x.mean()
y_mean = y.mean()
print("media de x = ", x_mean)
print("media de y = ", y_mean)
corr_coef = np.corrcoef(x,y)
print("correlation coeficient (off-diagonal) = ", corr_coef)
#covariance = np.cov(x,y)
#print('covariance = ', covariance)
# correlation coefficient 'by hand':
xy = x*y
covariance = xy.mean() - (x_mean*y_mean)
pearson_coef = covariance/(np.std(x)*np.std(y))
print("pearson coefficient = ", pearson_coef)
```
No exemplo abaixo, as variáveis estão **correlacionadas**. Podemos ver como o valor do coeficiente de correlação muda.
```
import matplotlib.pyplot as plt
import random
import numpy as np
# random seed for reproducibility
##random.seed(9823767454)
# lists of random numbers:
mu = 4.
sigma = .15
x = [random.gauss(mu, sigma) for i in range(10000)]
y = [i*random.gauss(mu, sigma) for i in x]
# convert list to numpy array for easy manipulation
x = np.array(x)
y = np.array(y)
plt.plot(x,y,'o')
plt.xlabel('x')
plt.ylabel('y')
x_mean = x.mean()
y_mean = y.mean()
print("media de x = ", x_mean)
print("media de y = ", y_mean)
corr_coef = np.corrcoef(x,y)
print("correlation coeficient (off-diagonal) = ", corr_coef)
#covariance = np.cov(x,y)
#print('covariance = ', covariance)
# correlation coefficient 'by hand':
xy = x*y
covariance = xy.mean() - (x_mean*y_mean)
pearson_coef = covariance/(np.std(x)*np.std(y))
print("pearson coefficient = ", pearson_coef)
```
No exemplo abaixo, as duas variáveis estão **totalmente correlacionadas**
```
import matplotlib.pyplot as plt
import random
import numpy as np
# random seed for reproducibility
##random.seed(9823767454)
# lists of random numbers:
mu = 4.
sigma = 1.
x = [random.gauss(mu, sigma) for i in range(10000)]
y = [i for i in x]
# convert list to numpy array for easy manipulation
x = np.array(x)
y = np.array(y)
plt.plot(x,y,'o')
plt.xlabel('x')
plt.ylabel('y')
x_mean = x.mean()
y_mean = y.mean()
print("media de x = ", x_mean)
print("media de y = ", y_mean)
corr_coef = np.corrcoef(x,y)
print("correlation coeficient (off-diagonal) = ", corr_coef)
#covariance = np.cov(x,y)
#print('covariance = ', covariance)
# correlation coefficient 'by hand':
xy = x*y
covariance = xy.mean() - (x_mean*y_mean)
pearson_coef = covariance/(np.std(x)*np.std(y))
print("pearson coefficient = ", pearson_coef)
```
| github_jupyter |
# Tutorial NlOpt
## Зачем это нужно?
В современных компетенциях инженерных или научных специальностей всё чаще приходится сталкиваться с теми или иными задачами требующими оптимизации функции.
В общем смысле под оптимизацией понимают поиск экстремума исследуемой функции.
$$f(x,y) \rightarrow max(min)$$
Заметим, что в случае простейших школьных функций одной переменной достаточно всего лишь приравнять производную от этой функции к нулю и решить полученное уравнение.
Но в более серьёзных задачах, где функции могут уже зависеть от нескольких переменных, такой метод может стать невозможным. Заметим также, что в зависимости от задачи
и самой функции, требуется применять разный алгоритм оптимизации.
К сожалению, использование внутренних команд питон может быть недостаточно для решения поставленной проблемы. В помощь этому приведем туториал по основам использования
мультиязычной и мультиплатформенной библиотеки NlOpt https://nlopt.readthedocs.io/en/latest/. В ней реализовано большое количество различных алгоритмов оптимизации число которых растёт, благодаря поддержке со стороны разработчиков. В качестве примера разберём несколько алгоритмов для оптимизации функции Химмельблау на языке питон. Будет приведён готовый программный код, который сразу можно будет использовать на практике.
> В туториале опущен вопрос по установке модуля на компьютер. С этим необходимо будет справиться самостоятельно.
## Сам процесс написания кода
Вид функции Химмельблау
$$f(x,y)=(x^2+y-11)^2+(x+y^2-7)^2$$
Вначале введём наши модули
```
import nlopt
from numpy import *
```
Вторым этапом распишем функцию *myfunc*
Для этого в строках *grad[0]* и *grad[1]* записываются частные производные функции от первой и второй переменных соответственно. *myfunc* возвращает саму функцию Химмельблау.
```
def myfunc(x, grad):
if grad.size > 0:
grad[0] = 4.0*x[0]*(x[0]**2+x[1]-11.0)+2.0*(x[0]+x[1]**2-7.0)
grad[1] = 2.0*(x[0]**2+x[1]-11.0)+4.0*x[1]*(x[0]+x[1]**2-7.0)
return ((x[0]**2+x[1]-11.0)**2+(x[0]+x[1]**2-7.0)**2)
```
Затем выбираем сам алгоритм оптимизации. На сайте представлен полный список всех алгоритмов и их описание. Со списком можно ознакомиться по ссылке https://nlopt.readthedocs.io/en/latest/NLopt_Algorithms/ Обращаем внимание, что **2** означает количество переменных от которых зависит исследуемая функция.
```
opt = nlopt.opt(nlopt.LN_BOBYQA, 2)
```
Дальше по пунктам:
- Запускам нашу функцию
- Задаём точность
- Выбираем начальную точку
- Задаём переменную, которая и будет равна оптимальному значению функции
- Вывод полученных результатов
```
opt.set_min_objective(myfunc)
opt.set_xtol_rel(1e-6)
x= opt.optimize([ 12.5, 1.5])
minf = opt.last_optimum_value()
print ("optimum at ", x[0], x[1])
print ("minimum value = ", minf)
```
На этом всё. В дальнейших уроках разберём оптимизацию с ограничением.
| github_jupyter |
# Vessels making voyages
The `voyages` table contains top level information about a voyage from one port to another, including when and where the voyage started and ended, and which vessel was involved in the voyage. You can use this information to identify which vessels made a voyage from one port to another in some time range:
```
from descarteslabs.vektorius import vector
from ipyleaflet import Map, GeoJSON
import warnings
warnings.filterwarnings('ignore')
ports = vector.table("ports")
voyages = vector.table("voyages")
# find geometry of start/end ports that we care about
origin_port_name = "PORT DE SALVADOR"
dest_port_name = "PORTO DE SUAPE"
ports = ports.filter(ports.port_name.isin([origin_port_name, dest_port_name]))
port_geoms = ports[ports.port_name, ports.port_geom].execute()
start_port_geom = port_geoms[port_geoms.port_name == origin_port_name].port_geom.iloc[0]
end_port_geom = port_geoms[port_geoms.port_name == dest_port_name].port_geom.iloc[0]
# because we store Geographies as geodesics, we need to force these literal shapes
# to be treated as such
start_port_geom.crs = "EPSG:4326"
end_port_geom.crs = "EPSG:4326"
# find voyages that start or end within timeframe
start_date = "2018-01-01"
end_date = "2018-12-31"
dt_filter = (voyages.departure.between(start_date, end_date) |
voyages.arrival.between(start_date, end_date))
# voyage should be between the two ports you care about
spatial_filter = (voyages.origin.intersects(start_port_geom) &
voyages.destination.intersects(end_port_geom))
voyages = voyages.filter(dt_filter & spatial_filter)
df = voyages[voyages.mmsi, voyages.linestring].execute()
df
```
Now we'll put this on a map to see what the voyages look like:
```
# create a map with ipyleaflet
m = Map(center=(15.453680224345835, -23.466796875000004), zoom=3)
# __geo_interface__ allows us to get the GeoJSON FeatureCollection
# representation of the GeoSeries
geo_json = GeoJSON(data=df.linestring.__geo_interface__)
m.add_layer(geo_json)
m
```
You can also estimate how much cargo in kg was transported from one port to another in each month. You can use functions to alter column data to achieve this analysis before fetching any of the data locally:
```
vessels = vector.table("vessels")
joined = voyages.inner_join(vessels, vessels.mmsi == voyages.mmsi)
capacity = joined[
# get vessel capacity in kg
(vessels.capacity.dwt * 1000).name("vessel_capacity_in_kg"),
# get the departure month for nicer grouping
voyages.departure.month().name("departure_month")
]
grouped = capacity.group_by(capacity.departure_month)
grouped = grouped.aggregate(capacity.vessel_capacity_in_kg
.sum()
.name("est_cargo_weight_in_kg"))
grouped = grouped.sort_by(grouped.departure_month)
df = grouped.execute()
df
```
| github_jupyter |
# The Lasso
Modified from the github repo: https://github.com/JWarmenhoven/ISLR-python which is based on the book by James et al. Intro to Statistical Learning.
```
# %load ../standard_import.txt
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.preprocessing import scale
from sklearn.model_selection import LeaveOneOut
from sklearn.linear_model import LinearRegression, lars_path, Lasso, LassoCV
%matplotlib inline
n=100
p=1000
X = np.random.randn(n,p)
X = scale(X)
sprob = 0.02
Sbool = np.random.rand(p) < sprob
s = np.sum(Sbool)
print("Number of non-zero's: {}".format(s))
mu = 100.
beta = np.zeros(p)
beta[Sbool] = mu * np.random.randn(s)
eps = np.random.randn(n)
y = X.dot(beta) + eps
larper = lars_path(X,y,method="lasso")
S = set(np.where(Sbool)[0])
for j in S:
_ = plt.plot(larper[0],larper[2][j,:],'r')
for j in set(range(p)) - S:
_ = plt.plot(larper[0],larper[2][j,:],'k',linewidth=.5)
_ = plt.title('Lasso path for simulated data')
_ = plt.xlabel('lambda')
_ = plt.ylabel('Coef')
```
# Hitters dataset
Let's load the dataset from the previous lab.
```
# In R, I exported the dataset from package 'ISLR' to a csv file.
df = pd.read_csv('../data/Hitters.csv', index_col=0).dropna()
df.index.name = 'Player'
df.info()
df.head()
dummies = pd.get_dummies(df[['League', 'Division', 'NewLeague']])
dummies.info()
print(dummies.head())
y = df.Salary
# Drop the column with the independent variable (Salary), and columns for which we created dummy variables
X_ = df.drop(['Salary', 'League', 'Division', 'NewLeague'], axis=1).astype('float64')
# Define the feature set X.
X = pd.concat([X_, dummies[['League_N', 'Division_W', 'NewLeague_N']]], axis=1)
X.info()
X.head(5)
```
__Exercise__ Compare the previous methods to the Lasso on this dataset. Tune $\lambda$ and compare the LOO risk to other methods (ridge, forward selection, etc.)
The following is a fast implementation of the lasso path cross-validated using LOO.
```
loo = LeaveOneOut()
looiter = loo.split(X)
hitlasso = LassoCV(cv=looiter)
hitlasso.fit(X,y)
print("The selected lambda value is {:.2f}".format(hitlasso.alpha_))
```
The following is the fitted coefficient vector for this chosen lambda.
```
hitlasso.coef_
np.mean(hitlasso.mse_path_[hitlasso.alphas_ == hitlasso.alpha_])
```
The above is the MSE for the selected model. The best performance for ridge regression was roughly 120,000, so this does not outperform ridge. We can also compare this to the selected model from forward stagewise regression:
```
[-0.21830515, 0.38154135, 0. , 0. , 0. ,
0.16139123, 0. , 0. , 0. , 0. ,
0.09994524, 0.56696569, -0.16872682, 0.16924078, 0. ,
0. , 0. , -0.19429699, 0. ]
```
This is not exactly the same model with differences in the inclusion or exclusion of AtBat, HmRun, Runs, RBI, Years, CHmRun, Errors, League_N, Division_W, NewLeague_N
```
bforw = [-0.21830515, 0.38154135, 0. , 0. , 0. ,
0.16139123, 0. , 0. , 0. , 0. ,
0.09994524, 0.56696569, -0.16872682, 0.16924078, 0. ,
0. , 0. , -0.19429699, 0. ]
print(", ".join(X.columns[(hitlasso.coef_ != 0.) != (bforw != 0.)]))
```
| github_jupyter |
```
import pandas as pd
from googleapiclient.discovery import build
from googleapiclient.errors import HttpError
from oauth2client.tools import argparser
target = "아디다스 슈퍼스타"
DEVELOPER_KEY = "AIzaSyAnEEAKE50qxf5lHbsucDiMNayh9aFUj5g"
YOUTUBE_API_SERVICE_NAME="youtube"
YOUTUBE_API_VERSION="v3"
youtube = build(YOUTUBE_API_SERVICE_NAME,YOUTUBE_API_VERSION,developerKey=DEVELOPER_KEY)
# 유튜브 검색하는 함수
def surfing_target(target):
search_response = youtube.search().list(
q = target, # 검색어
publishedAfter = '2020-01-01T00:00:00Z', # 특정 일시 이후 영상만 검색
order = "viewCount", # 조회수 기준으로 정렬
part = "snippet", # 기본값
maxResults = 10, # 최대 10개의 검색어 가져오기
type = 'video', # 비디오 형태만 가져오기
regionCode = 'KR' # 검색 국가 : 한국
).execute()
# https://developers.google.com/youtube/v3/docs/search/list?hl=ko
search_title_list = []
search_id_list = []
search_date_list = []
search_channel_list = []
search_description_list = []
for i in range(len(search_response['items'])):
search_title_list.append(search_response['items'][i]['snippet']['title'])
search_id_list.append(search_response['items'][i]['id']['videoId'])
search_date_list.append(search_response['items'][i]['snippet']['publishTime'])
search_channel_list.append(search_response['items'][i]['snippet']['channelTitle'])
search_description_list.append(search_response['items'][i]['snippet']['description'])
dataset = pd.DataFrame({'title':search_title_list, 'id':search_id_list, 'date':search_date_list, 'channel':search_channel_list, 'description':search_description_list})
return dataset
dataset = surfing_target(target)
dataset
# 댓글 긁어오는 함수
def get_comments(video_id):
comments = list()
response = youtube.commentThreads().list(part='snippet,replies', videoId=video_id, maxResults=100).execute()
# textDisplay: 댓글의 내용
# authorDisplayName: 댓글 작성자
# publishedAt: 댓글 작성 시간
# likeCount: 좋아요 수
while response:
for item in response['items']:
comment = item['snippet']['topLevelComment']['snippet']
comments.append([comment['textDisplay'], comment['authorDisplayName'], comment['publishedAt'], comment['likeCount']])
if item['snippet']['totalReplyCount'] > 0:
for reply_item in item['replies']['comments']:
reply = reply_item['snippet']
comments.append([reply['textDisplay'], reply['authorDisplayName'], reply['publishedAt'], reply['likeCount']])
if 'nextPageToken' in response:
response = youtube.commentThreads().list(part='snippet,replies', videoId=video_id, pageToken=response['nextPageToken'], maxResults=100).execute()
else:
break
df = pd.DataFrame(data=comments, columns=['comment', 'author', 'date', 'num_likes'])
# df.to_excel('temp.xlsx', index=None)
return df
df_list = [0 for i in range(len(dataset))]
for i in range(len(dataset)):
df_list[i] = get_comments(dataset['id'][i])
df_list[0]
temp_df = pd.concat([tmp for tmp in df_list], ignore_index=True)
temp_df.columns = df_list[0].columns.tolist()
temp_df
temp_df.comment.values
```
| github_jupyter |
# Stacking LSTM Layers
-----------------
Here we implement an LSTM model on all a data set of Shakespeare works. We will stack multiple LSTM models for a more accurate representation of Shakespearean language. We will also use characters instead of words.
```
import os
import re
import string
import requests
import numpy as np
import collections
import random
import pickle
import matplotlib.pyplot as plt
import tensorflow as tf
from tensorflow.python.framework import ops
ops.reset_default_graph()
```
Start a computational graph session.
```
sess = tf.Session()
```
Set RNN Parameters
```
num_layers = 3 # Number of RNN layers stacked
min_word_freq = 5 # Trim the less frequent words off
rnn_size = 128 # RNN Model size, has to equal embedding size
epochs = 10 # Number of epochs to cycle through data
batch_size = 100 # Train on this many examples at once
learning_rate = 0.0005 # Learning rate
training_seq_len = 50 # how long of a word group to consider
save_every = 500 # How often to save model checkpoints
eval_every = 50 # How often to evaluate the test sentences
prime_texts = ['thou art more', 'to be or not to', 'wherefore art thou']
```
Download/store Shakespeare data
```
data_dir = 'temp'
data_file = 'shakespeare.txt'
model_path = 'shakespeare_model'
full_model_dir = os.path.join(data_dir, model_path)
```
Declare the punctuation and then create the model and data directories
```
# Declare punctuation to remove, everything except hyphens and apostrophes
punctuation = string.punctuation
punctuation = ''.join([x for x in punctuation if x not in ['-', "'"]])
# Make Model Directory
if not os.path.exists(full_model_dir):
os.makedirs(full_model_dir)
# Make data directory
if not os.path.exists(data_dir):
os.makedirs(data_dir)
```
Load the Shakespeare Data
```
print('Loading Shakespeare Data')
# Check if file is downloaded.
if not os.path.isfile(os.path.join(data_dir, data_file)):
print('Not found, downloading Shakespeare texts from www.gutenberg.org')
shakespeare_url = 'http://www.gutenberg.org/cache/epub/100/pg100.txt'
# Get Shakespeare text
response = requests.get(shakespeare_url)
shakespeare_file = response.content
# Decode binary into string
s_text = shakespeare_file.decode('utf-8')
# Drop first few descriptive paragraphs.
s_text = s_text[7675:]
# Remove newlines
s_text = s_text.replace('\r\n', '')
s_text = s_text.replace('\n', '')
# Write to file
with open(os.path.join(data_dir, data_file), 'w') as out_conn:
out_conn.write(s_text)
else:
# If file has been saved, load from that file
with open(os.path.join(data_dir, data_file), 'r') as file_conn:
s_text = file_conn.read().replace('\n', '')
print('Done Loading Data.')
```
Clean and split the text data.
```
# Clean text
print('Cleaning Text')
s_text = re.sub(r'[{}]'.format(punctuation), ' ', s_text)
s_text = re.sub('\s+', ' ', s_text ).strip().lower()
# Split up by characters
char_list = list(s_text)
```
Build word vocabulary function and transform the text.
```
def build_vocab(characters):
character_counts = collections.Counter(characters)
# Create vocab --> index mapping
chars = character_counts.keys()
vocab_to_ix_dict = {key:(ix+1) for ix, key in enumerate(chars)}
# Add unknown key --> 0 index
vocab_to_ix_dict['unknown']=0
# Create index --> vocab mapping
ix_to_vocab_dict = {val:key for key,val in vocab_to_ix_dict.items()}
return(ix_to_vocab_dict, vocab_to_ix_dict)
# Build Shakespeare vocabulary
print('Building Shakespeare Vocab by Characters')
ix2vocab, vocab2ix = build_vocab(char_list)
vocab_size = len(ix2vocab)
print('Vocabulary Length = {}'.format(vocab_size))
# Sanity Check
assert(len(ix2vocab) == len(vocab2ix))
```
Convert text to word vectors
```
s_text_ix = []
for x in char_list:
try:
s_text_ix.append(vocab2ix[x])
except:
s_text_ix.append(0)
s_text_ix = np.array(s_text_ix)
```
Define LSTM RNN Model Class
```
class LSTM_Model():
def __init__(self, rnn_size, num_layers, batch_size, learning_rate,
training_seq_len, vocab_size, infer_sample=False):
self.rnn_size = rnn_size
self.num_layers = num_layers
self.vocab_size = vocab_size
self.infer_sample = infer_sample
self.learning_rate = learning_rate
if infer_sample:
self.batch_size = 1
self.training_seq_len = 1
else:
self.batch_size = batch_size
self.training_seq_len = training_seq_len
self.lstm_cell = tf.contrib.rnn.BasicLSTMCell(rnn_size)
self.lstm_cell = tf.contrib.rnn.MultiRNNCell([self.lstm_cell for _ in range(self.num_layers)])
self.initial_state = self.lstm_cell.zero_state(self.batch_size, tf.float32)
self.x_data = tf.placeholder(tf.int32, [self.batch_size, self.training_seq_len])
self.y_output = tf.placeholder(tf.int32, [self.batch_size, self.training_seq_len])
with tf.variable_scope('lstm_vars'):
# Softmax Output Weights
W = tf.get_variable('W', [self.rnn_size, self.vocab_size], tf.float32, tf.random_normal_initializer())
b = tf.get_variable('b', [self.vocab_size], tf.float32, tf.constant_initializer(0.0))
# Define Embedding
embedding_mat = tf.get_variable('embedding_mat', [self.vocab_size, self.rnn_size],
tf.float32, tf.random_normal_initializer())
embedding_output = tf.nn.embedding_lookup(embedding_mat, self.x_data)
rnn_inputs = tf.split(axis=1, num_or_size_splits=self.training_seq_len, value=embedding_output)
rnn_inputs_trimmed = [tf.squeeze(x, [1]) for x in rnn_inputs]
decoder = tf.contrib.legacy_seq2seq.rnn_decoder
outputs, last_state = decoder(rnn_inputs_trimmed,
self.initial_state,
self.lstm_cell)
# RNN outputs
output = tf.reshape(tf.concat(axis=1, values=outputs), [-1, rnn_size])
# Logits and output
self.logit_output = tf.matmul(output, W) + b
self.model_output = tf.nn.softmax(self.logit_output)
loss_fun = tf.contrib.legacy_seq2seq.sequence_loss_by_example
loss = loss_fun([self.logit_output],[tf.reshape(self.y_output, [-1])],
[tf.ones([self.batch_size * self.training_seq_len])],
self.vocab_size)
self.cost = tf.reduce_sum(loss) / (self.batch_size * self.training_seq_len)
self.final_state = last_state
gradients, _ = tf.clip_by_global_norm(tf.gradients(self.cost, tf.trainable_variables()), 4.5)
optimizer = tf.train.AdamOptimizer(self.learning_rate)
self.train_op = optimizer.apply_gradients(zip(gradients, tf.trainable_variables()))
def sample(self, sess, words=ix2vocab, vocab=vocab2ix, num=20, prime_text='thou art'):
state = sess.run(self.lstm_cell.zero_state(1, tf.float32))
char_list = list(prime_text)
for char in char_list[:-1]:
x = np.zeros((1, 1))
x[0, 0] = vocab[char]
feed_dict = {self.x_data: x, self.initial_state:state}
[state] = sess.run([self.final_state], feed_dict=feed_dict)
out_sentence = prime_text
char = char_list[-1]
for n in range(num):
x = np.zeros((1, 1))
x[0, 0] = vocab[char]
feed_dict = {self.x_data: x, self.initial_state:state}
[model_output, state] = sess.run([self.model_output, self.final_state], feed_dict=feed_dict)
sample = np.argmax(model_output[0])
if sample == 0:
break
char = words[sample]
out_sentence = out_sentence + char
return(out_sentence)
```
Initialize the LSTM Model
```
lstm_model = LSTM_Model(rnn_size, num_layers, batch_size, learning_rate,
training_seq_len, vocab_size)
# Tell TensorFlow we are reusing the scope for the testing
with tf.variable_scope(tf.get_variable_scope(), reuse=True):
test_lstm_model = LSTM_Model(rnn_size,num_layers, batch_size, learning_rate,
training_seq_len, vocab_size, infer_sample=True)
```
Create model saver
```
saver = tf.train.Saver(tf.global_variables())
```
Create batches for each epoch
```
num_batches = int(len(s_text_ix)/(batch_size * training_seq_len)) + 1
# Split up text indices into subarrays, of equal size
batches = np.array_split(s_text_ix, num_batches)
# Reshape each split into [batch_size, training_seq_len]
batches = [np.resize(x, [batch_size, training_seq_len]) for x in batches]
```
Initialize all variables and train the model!
```
# Initialize all variables
init = tf.global_variables_initializer()
sess.run(init)
# Train model
train_loss = []
iteration_count = 1
for epoch in range(epochs):
# Shuffle word indices
random.shuffle(batches)
# Create targets from shuffled batches
targets = [np.roll(x, -1, axis=1) for x in batches]
# Run a through one epoch
print('Starting Epoch #{} of {}.'.format(epoch+1, epochs))
# Reset initial LSTM state every epoch
state = sess.run(lstm_model.initial_state)
for ix, batch in enumerate(batches):
training_dict = {lstm_model.x_data: batch, lstm_model.y_output: targets[ix]}
# We need to update initial state for each RNN cell:
for i, (c, h) in enumerate(lstm_model.initial_state):
training_dict[c] = state[i].c
training_dict[h] = state[i].h
temp_loss, state, _ = sess.run([lstm_model.cost, lstm_model.final_state, lstm_model.train_op],
feed_dict=training_dict)
train_loss.append(temp_loss)
# Print status every 10 gens
if iteration_count % 10 == 0:
summary_nums = (iteration_count, epoch+1, ix+1, num_batches+1, temp_loss)
print('Iteration: {}, Epoch: {}, Batch: {} out of {}, Loss: {:.2f}'.format(*summary_nums))
# Save the model and the vocab
if iteration_count % save_every == 0:
# Save model
model_file_name = os.path.join(full_model_dir, 'model')
saver.save(sess, model_file_name, global_step = iteration_count)
print('Model Saved To: {}'.format(model_file_name))
# Save vocabulary
dictionary_file = os.path.join(full_model_dir, 'vocab.pkl')
with open(dictionary_file, 'wb') as dict_file_conn:
pickle.dump([vocab2ix, ix2vocab], dict_file_conn)
if iteration_count % eval_every == 0:
for sample in prime_texts:
print(test_lstm_model.sample(sess, ix2vocab, vocab2ix, num=10, prime_text=sample))
iteration_count += 1
```
Plot loss over time
```
plt.plot(train_loss, 'k-')
plt.title('Sequence to Sequence Loss')
plt.xlabel('Generation')
plt.ylabel('Loss')
plt.show()
```
| github_jupyter |
#### Verification Alignment
A forecast is verified by comparing a set of initializations at a given lead to
observations over some window of time. However, there are a few ways to decide *which*
initializations or verification window to use in this alignment.
One must pass the keyword ``alignment=...`` to the hindcast `.verify()` method to set the behavior for aligning forecasts with the verification product. Note that the alignment decision only matters for [hindcast experiments](terminology.html#simulation-design). [Perfect-model experiments](terminology.html#simulation-design) are perfectly time-aligned by design, equating to our `same_inits` keyword.
The available keywords for hindcast alignment are:
* `'same_inits'`: Use a common set of initializations that verify
across all leads. This ensures that there is no bias in the result due to the state
of the system for the given initializations.
* `'same_verifs'`: Use a common verification window across all leads. This ensures
that there is no bias in the result due to the observational period being verified
against.
* `'maximize'`: Use all available initializations at each lead that verify against
the observations provided. This changes both the set of initializations and the
verification window used at each lead.
```
# linting
%load_ext nb_black
%load_ext lab_black
from climpred import HindcastEnsemble
from climpred.tutorial import load_dataset
from esmtools.stats import rm_trend
import matplotlib.pyplot as plt
plt.style.use("fivethirtyeight")
%matplotlib inline
import numpy as np
import warnings
# Supress datetime warnings for this page.
warnings.filterwarnings("ignore")
def create_hindcast_object():
"""Loads in example data from CESM-DPLE and ERSST observations and detrends."""
hind = load_dataset("CESM-DP-SST")["SST"]
verif = load_dataset("ERSST")["SST"]
# Bias-correct over same period as CESM-DPLE.
verif = verif - verif.sel(time=slice(1964, 2014)).mean("time")
# Remove linear trend.
hind_dt = rm_trend(hind, dim="init").rename("SST")
verif_dt = rm_trend(verif, dim="time").rename("SST")
# Create `HindcastEnsemble` object from `climpred`.
hindcast = HindcastEnsemble(hind)
hindcast = hindcast.add_observations(verif)
hindcast_dt = HindcastEnsemble(hind_dt)
hindcast_dt = hindcast_dt.add_observations(verif_dt)
return hindcast, hindcast_dt
hindcast, hindcast_dt = create_hindcast_object()
```
The user can simply change the alignment strategy by passing in the keyword `alignment=...`. Note that the choice of alignment strategy changes the lead-dependent metric results.
```
f, axs = plt.subplots(ncols=2, figsize=(12, 4), sharex=True)
for alignment in ["same_inits", "same_verifs", "maximize"]:
hindcast.verify(metric="acc", comparison="e2o", dim="init", alignment=alignment)[
"SST"
].plot(label=alignment, ax=axs[0])
hindcast_dt.verify(metric="acc", comparison="e2o", dim="init", alignment=alignment)[
"SST"
].plot(label=alignment, ax=axs[1])
axs[0].legend()
axs[1].legend()
axs[0].set(
ylabel="anomaly\ncorrelation coefficient",
xlabel="lead year",
xticks=np.arange(1, 11),
title="SST with trend",
)
axs[1].set(
ylabel="anomaly\ncorrelation coefficient", xlabel="lead year", title="detrended SST"
)
f.suptitle("Verification with Different Alignment Methods", fontsize=14, weight="bold")
plt.subplots_adjust(top=0.85)
plt.show()
```
These alignment keywords also extend to reference forecasts (e.g. `reference='persistence'`), which uses the identical set of initializations (and alignment strategy) in its computation. Below, the dashed lines represent the persistence forecast for the given alignment strategy, while the solid lines denote the initialized anomaly correlation coefficient (as in the above plots).
```
COLORS = ["#008FD5", "#FC4F30", "#E5AE38"]
f, axs = plt.subplots()
for alignment, color in zip(["same_inits", "same_verifs", "maximize"], COLORS):
result = hindcast_dt.verify(
metric="acc",
reference="persistence",
comparison="e2o",
dim="init",
alignment=alignment,
)
result.sel(skill="initialized").SST.plot(label=alignment, color=color)
result.sel(skill="persistence").SST.plot(linestyle="--", color=color, lw=3)
axs.set(
ylabel="anomaly\ncorrelation coefficient",
xlabel="lead year",
xticks=np.arange(1, 11),
title="Detrended SST Verification with Persistence",
)
plt.legend()
plt.show()
```
We'll be using the same example data as above. `climpred` will be aligning the following initialization and verification dates:
```
print(f"initialization dates: \n{hindcast.get_initialized().init.to_index()}")
print(f"verification dates: \n{hindcast.get_observations().time.to_index()}")
```
We use the standard python library `logging` to log the initializations and verification dates used in alignment at each lead. The user can check these logs to ensure that the expected initializations and verification dates are being retained. See the logging section on this page for more details.
```
import logging
# Print log to screen with initializations and verification dates.
logger = logging.getLogger()
logger.setLevel(logging.INFO)
```
## Same Verification Dates
`alignment='same_verifs'`
The `same_verifs` alignment finds a set of verification dates that can be verified against over all leads. It also requires that the verification data have an observation at each initialization being retained. This is so that the reference forecast, such as persistence, uses an identical set of initializations in deriving its forecast. Notice in the logger output that a common set of verification dates spanning 1965-2015 are used, while the initialization window slides one year at each lead.
**References**:
1. Boer, George J., et al. "The decadal climate prediction project (DCPP) contribution to CMIP6." Geoscientific Model Development (Online) 9.10 (2016). [https://doi.org/10.5194/gmd-9-3751-2016]
2. Hawkins, Ed, et al. "The interpretation and use of biases in decadal climate predictions." Journal of climate 27.8 (2014): 2931-2947. [https://doi.org/10.1175/JCLI-D-13-00473.1]
3. Smith, Doug M., Rosie Eade, and Holger Pohlmann. "A comparison of full-field and anomaly initialization for seasonal to decadal climate prediction." Climate dynamics 41.11-12 (2013): 3325-3338. [https://doi.org/10.1007/s00382-013-1683-2]
```
skill = hindcast.verify(
metric="acc", comparison="e2o", dim="init", alignment="same_verifs"
)
```
Here, we include a figure of a simpler alignment case with annual initializations from 1990 through 2000 and three lead years. We verify this hypothetical initialized ensemble against a product that spans 1995 through 2002.
Two conditions must be met when selecting the verification window:
1. There must be a union between the initialization dates and verification dates. This
is represented by the black vertical lines in the top panel below, which leave out
1990-1994 initializations since there aren't observations before 1995. This logic
exists so that any reference forecasts
(e.g. a persistence forecast) use an identical set of initializations as the
initialized forecast.
2. A given verification time must exist across all leads. This is to ensure that at each
lead, the entire set of chosen verification dates can be verified against. This is
represented by diagonals in the top panel below (and the dashed black lines).
Without the first stipulation, this would set the verification window at 1995-2001.
This leaves us with a verification window of [1998, 1999, 2000, 2001] which can be verified against across all leads (and have a complimentary persistence forecast with the same set of initializations used at each lead).

## Same Initializations
`alignment='same_inits'`
The `same_inits` alignment finds a set of initializations that can verify over all leads. It also requires that the verification data have an observation at each initialization being retained. This is so that the reference forecast, such as persistence, uses an identical set of initializations in deriving its forecast. Notice in the logger output that a common set of initializations spanning 1955-2005 are used, while the verification window slides one year at each lead.
```
skill = hindcast.verify(
metric="acc", comparison="e2o", dim="init", alignment="same_inits"
)
```
Here, we include a figure of a simpler alignment case with annual initializations from 1990 through 2000 and three lead years. We verify this hypothetical initialized ensemble against a product that spans 1995 through 2002.
Two conditions must be met to retain the initializations for verification:
1. There must be an observation in the verification data for the given initialization.
In combination with (1), initializations 1990 through 1994 are left out. This logic
exists so that any reference forecast (e.g. a persistence forecast) use an identical set of initializations as the
initialized forecast.
2. All forecasted times (i.e., initialization + lead year) for a given initialization
must be contained in the verification data. Schematically, this means that there must
be a union between a column in the top panel and the time series in the bottom panel.
The 2000 initialization below is left out since the verification data does not
contain 2003.
This leaves us with initializations [1995, 1996, ..., 1999] which can verify against the observations at all three lead years.

## Maximize Degrees of Freedom
`alignment='maximize'`
The `maximize` alignment verifies against every available observation at each lead. This means that both the initializations and verification dates could be different at each lead. It also requires that the verification data have an observation at each initialization being retained. This is so that the reference forecast, such as persistence, uses an identical set of initializations in deriving its forecast.
Notice in the logger output that the initialization window shrinks from 1955-2014 (N=60) at lead year 1 to 1955-2005 (N=51) at lead year 10. Similarly, the verification window spans 1956-2015 at lead year 1 and 1965-2015 at lead year 10. However, using the other two alignment strategies (`same_verifs` and `same_inits`), there is a fixed N=51 to ensure constant initializations or verification dates, while the number of samples is extended to as high as 60 with this alignment strategy.
**References**:
1. Yeager, S. G., et al. "Predicting near-term changes in the Earth System: A large ensemble of initialized decadal prediction simulations using the Community Earth System Model." Bulletin of the American Meteorological Society 99.9 (2018): 1867-1886. [https://doi.org/10.1175/BAMS-D-17-0098.1]
```
skill = hindcast.verify(
metric="acc", comparison="e2o", dim="init", alignment="maximize"
)
```
Here, we include a figure of a simpler alignment case with annual initializations from 1990 through 2000 and three lead years. We verify this hypothetical initialized ensemble against a product that spans 1995 through 2002.
Two conditions must be met when selecting initializations/verifications at each lead:
1. There must be a union between the initialization dates and verification dates. This
is represented by the black vertical lines in the top panel below, which leave out
1990-1994 initializations since there aren't observations before 1995. This logic
exists so that any reference forecasts
(e.g. a persistence forecast) use an identical set of initializations as the
initialized forecast.
2. The selected initializations must verify with the provided observations for the given lead.
This is shown by the hatching in the figure below. The 2000 initialization is left out
at lead year 3 since there is no observation for 2003.
This leaves us with the following alignment:
* LY1 initializations: [1995, 1996, 1997, 1998, 1999, 2000]
* LY2 initializations: [1995, 1996, 1997, 1998, 1999, 2000]
* LY3 initializations: [1995, 1996, 1997, 1998, 1999]

## Logging
``climpred`` uses the standard library ``logging`` to store the initializations and verification dates used at each lead for a given computation. This is used internally for testing, but more importantly, can be activated by the user so they can be sure of how computations are being done.
To see the log interactively, e.g. while working in Jupyter notebooks or on the command line use the following:
```
import logging
logger = logging.getLogger()
logger.setLevel(logging.INFO)
skill = hindcast.verify(
metric="acc", comparison="e2o", dim="init", alignment="same_verifs"
)
```
The `INFO` level reports the minimum and maximum bounds for initializations and verification dates. To see every single initialization and verification date used, set the level to `DEBUG`.
```
logger.setLevel(logging.DEBUG)
skill = hindcast.isel(lead=slice(0, 2)).verify(
metric="acc", comparison="e2o", dim="init", alignment="same_verifs"
)
```
One can also save out the log to a file.
```
logger = logging.getLogger()
logger.setLevel(logging.INFO)
fh = logging.FileHandler("hindcast.out")
logger.addHandler(fh)
skill = hindcast.verify(
metric="acc", comparison="e2o", dim="init", alignment="same_verifs"
)
skill = hindcast.verify(
metric="acc", comparison="e2o", dim="init", alignment="same_verifs"
)
!cat hindcast.out
!rm hindcast.out
```
| github_jupyter |
In this notebook, we explore the learning curve for the toxic spans detector
```
from transformers import RobertaTokenizer, RobertaForTokenClassification
from transformers import BertTokenizer, BertForTokenClassification
from transformers import AutoTokenizer, AutoModelForTokenClassification
import torch
import numpy as np
import pandas as pd
import os
os.environ['CUDA_VISIBLE_DEVICES'] = '5'
device = torch.device('cuda:0')
model_name = 'roberta-base' #roberta-base
tokenizer = AutoTokenizer.from_pretrained(model_name)
# model = AutoModelForTokenClassification.from_pretrained(model_name)
```
```
inputs = tokenizer("Hello, my dog is cute", return_tensors="pt")
labels = torch.tensor([1] * inputs["input_ids"].size(1)).unsqueeze(0) # Batch size 1
outputs = model(**inputs, labels=labels)
```
# Create labels for tagging
```
import os
import numpy as np
import pandas as pd
from ast import literal_eval
import re
import nltk
import matplotlib.pyplot as plt
from nltk.tokenize import word_tokenize
path = 'data/'
trial = pd.read_csv(path + 'tsd_trial.csv')
train = pd.read_csv(path + 'tsd_train.csv')
# final_test = pd.read_csv(path + 'tsd_test.csv')
final_test = pd.read_csv(path + 'tsd_test_gt.csv')
train['spans'] = train.spans.apply(literal_eval)
trial['spans'] = trial.spans.apply(literal_eval)
final_test['spans'] = final_test.spans.apply(literal_eval)
trial.shape, train.shape, final_test.shape
print(len(set(trial.text).intersection(set(train.text))))
print(len(set(final_test.text).intersection(set(train.text))))
print((train.spans.apply(len) == 0).mean())
print((trial.spans.apply(len) == 0).mean())
import spans_utils
from importlib import reload
reload(spans_utils)
from spans_utils import display_spans, spans2labels, labels2spans
display_spans(trial.spans[0], trial.text[0])
display_spans(trial.spans[0], trial.text[0])
from tqdm.auto import tqdm, trange
train_labels = [spans2labels(row.text, row.spans, tokenizer) for i, row in tqdm(train.iterrows())]
trial_labels = [spans2labels(row.text, row.spans, tokenizer) for i, row in tqdm(trial.iterrows())]
train['labels'] = train_labels
trial['labels'] = trial_labels
class SpansDataset(torch.utils.data.Dataset):
def __init__(self, encodings, labels=None):
self.encodings = encodings
self.labels = labels
def __getitem__(self, idx):
item = {key: val[idx] for key, val in self.encodings.items()}
if self.labels is not None:
item['labels'] = self.labels[idx]
return item
def __len__(self):
return len(self.encodings['input_ids'])
train_dataset = SpansDataset(tokenizer(train.text.tolist()), train_labels)
eval_dataset = SpansDataset(tokenizer(trial.text.tolist()), trial_labels)
indices = np.arange(train.shape[0])
np.random.seed(1)
np.random.shuffle(indices)
indices
train_sets = {
k: SpansDataset(tokenizer(train.text.iloc[indices[:k]].tolist()), [train_labels[i] for i in indices[:k]])
for k in [10, 30, 100, 300, 1000, 3000, 10000]
}
train_corpora = {
k: (train.text.iloc[indices[:k]].tolist(), [train_labels[i] for i in indices[:k]])
for k in [10, 30, 100, 300, 1000, 3000, 10000]
}
final_test_dataset = SpansDataset(tokenizer(final_test.text.tolist()))
from transformers import DataCollatorForTokenClassification
data_collator = DataCollatorForTokenClassification(tokenizer, padding=True)
import numpy as np
from semeval2021 import f1
```
# Train a single-task model
https://github.com/huggingface/notebooks/blob/master/examples/token_classification.ipynb
https://huggingface.co/transformers/custom_datasets.html
```
from transformers import Trainer, TrainingArguments, EarlyStoppingCallback
from transformers.file_utils import cached_property
from typing import Tuple
class TrAr(TrainingArguments):
@cached_property
def _setup_devices(self):
return device
torch.cuda.set_device(device)
from datetime import datetime
datetime.now()
import gc
def cleanup():
gc.collect()
torch.cuda.empty_cache()
for k in [10, 30, 100, 300, 1000, 3000, 10000]:
print(f'Training model on {k} examples')
print(datetime.now())
model = AutoModelForTokenClassification.from_pretrained(model_name)
cleanup()
model.to(device);
# first part
for param in model.roberta.parameters():
param.requires_grad = False
MODEL_NAME = f'./models_scale/roberta_single_{k}'
training_args = TrAr(
output_dir=MODEL_NAME, # output directory
overwrite_output_dir=True,
num_train_epochs=100_000, # total # of training epochs
per_device_train_batch_size=8, # batch size per device during training
per_device_eval_batch_size=8, # batch size for evaluation
warmup_steps=3000, # number of warmup steps for learning rate scheduler
weight_decay=1e-8, # strength of weight decay
learning_rate=1e-3,
logging_dir='./logs2', # directory for storing logs
logging_steps=50,
eval_steps=50,
evaluation_strategy='steps',
save_total_limit=1,
load_best_model_at_end=True,
)
trainer = Trainer(
model=model, # the instantiated 🤗 Transformers model to be trained
args=training_args, # training arguments, defined above
train_dataset=train_sets[k], # training dataset
eval_dataset=eval_dataset, # evaluation dataset
data_collator=data_collator,
tokenizer=tokenizer,
callbacks=[EarlyStoppingCallback(early_stopping_patience=5, early_stopping_threshold=0)]
)
trainer.train()
model.save_pretrained(MODEL_NAME + '_head')
loss_head = trainer.evaluate()['eval_loss']
for param in model.parameters():
param.requires_grad = True
print(f'Tuning model on {k} examples')
training_args = TrAr(
output_dir=MODEL_NAME, # output directory
overwrite_output_dir=True,
num_train_epochs=100_000, # total # of training epochs
per_device_train_batch_size=8, # batch size per device during training
per_device_eval_batch_size=8, # batch size for evaluation
warmup_steps=3000, # number of warmup steps for learning rate scheduler
weight_decay=1e-8, # strength of weight decay
learning_rate=1e-5,
logging_dir='./logs2', # directory for storing logs
logging_steps=50,
eval_steps=50,
evaluation_strategy='steps',
save_total_limit=1,
load_best_model_at_end=True,
)
trainer = Trainer(
model=model, # the instantiated 🤗 Transformers model to be trained
args=training_args, # training arguments, defined above
train_dataset=train_sets[k], # training dataset
eval_dataset=eval_dataset, # evaluation dataset
data_collator=data_collator,
tokenizer=tokenizer,
callbacks=[EarlyStoppingCallback(early_stopping_patience=5, early_stopping_threshold=0)]
)
trainer.train()
loss_full = trainer.evaluate()['eval_loss']
model.save_pretrained(MODEL_NAME + '_full')
cleanup()
print(f'Tuning model on {k} examples completed: losses {loss_head} -> {loss_full}')
print(datetime.now())
```
### evaluate
```
scores_50 = {
'head': [],
'full': []
}
scores_max = {
'head': [],
'full': []
}
for k in [10, 30, 100, 300, 1000, 3000, 10000]:
for mode in ['head', 'full']:
MODEL_NAME = f'./models_scale/roberta_single_{k}_{mode}'
model = AutoModelForTokenClassification.from_pretrained(MODEL_NAME)
cleanup()
model.to(device);
model.eval();
training_args = TrAr(
output_dir='tmp',
per_device_eval_batch_size=64, # batch size for evaluation
)
trainer = Trainer(
model=model, # the instantiated 🤗 Transformers model to be trained
args=training_args, # training arguments, defined above
train_dataset=train_sets[k], # training dataset
data_collator=data_collator,
tokenizer=tokenizer,
)
pred = trainer.predict(final_test_dataset)
scores = []
for threshold in [0.1, 0.15, 0.2, 0.25, 0.3, 0.35, 0.4, 0.45, 0.5, 0.55, 0.6, 0.65, 0.7, 0.75, 0.8, 0.85, 0.9]:
preds = []
for text, pr in zip(final_test.text, pred.predictions):
proba = np.exp(pr[pr[:, 0]!=-100])
proba /= proba.sum(axis=1, keepdims=True)
labels = (proba[:, 1] >= threshold).astype(int).tolist()
preds.append(labels2spans(text, labels, tokenizer))
score = np.mean([f1(p, y) for p, y in zip(preds, final_test.spans)])
scores.append(score)
if threshold == 0.5:
score_50 = score
score_max = max(scores)
print(k, mode, score_50, score_max)
scores_50[mode].append(score_50)
scores_max[mode].append(score_max)
print(scores_50)
# scores_50 = {'head': [0.41317980330200765, 0.39298222111786046, 0.4651392154783845, 0.5054483376082518, 0.5754306834078432, 0.5775647779757941, 0.5790846863257059], 'full': [0.4694123313296014, 0.46052868203784997, 0.49794774709892325, 0.5500632412719044, 0.6098243106751826, 0.6543757786629506, 0.6416565127682992]}
print(scores_max)
# scores_max = {'head': [0.43610746759966934, 0.42549428253200916, 0.48549169897463534, 0.5434453554105662, 0.5754306834078432, 0.5891508198890186, 0.5920799374256265], 'full': [0.4698994695689647, 0.46052868203784997, 0.5033596644709653, 0.56008830890506, 0.6106484939446047, 0.6681581737239815, 0.6606782917283851]}
import matplotlib.pyplot as plt
%matplotlib inline
xs = np.array([10, 30, 100, 300, 1000, 3000, 7939])
plt.plot(xs, scores_50['head'])
plt.plot(xs, scores_50['full'])
plt.legend(['head', 'full'])
plt.xscale('log');
plt.plot(xs, scores_max['head'], marker='*')
plt.plot(xs, scores_max['full'], marker='o')
plt.legend(['train only head', 'train full model'])
#plt.scatter(xs, scores_max['head'])
#plt.scatter(xs, scores_max['full'])
plt.xscale('log');
plt.xlabel('training set size');
plt.ylabel('F1 score on the test set');
plt.title('learning curve of a vanilla tagger');
```
Result: the learning curve is steep, but not very steep really.
```
sm2 = [0.4698994695689647, 0.46052868203784997, 0.5033596644709653, 0.56008830890506, 0.6106484939446047, 0.6681581737239815, 0.6769273 ]
plt.plot(xs, scores_max['head'], marker='*')
plt.plot(xs, sm2, marker='o')
plt.legend(['train only head', 'train full model'])
#plt.scatter(xs, scores_max['head'])
#plt.scatter(xs, scores_max['full'])
plt.xscale('log');
plt.xlabel('training set size');
plt.ylabel('F1 score on the test set');
plt.title('learning curve of a vanilla tagger');
```
# Fine-tune the head-first classifier
```
for k in [10, 30, 100, 300, 1000, 3000, 10000]:
print(f'Training model on {k} examples')
print(datetime.now())
model = AutoModelForTokenClassification.from_pretrained('models/roberta_clf_proba')
cleanup()
model.to(device);
# first part
for param in model.roberta.parameters():
param.requires_grad = False
MODEL_NAME = f'./models_scale_ft/roberta_single_{k}'
training_args = TrAr(
output_dir=MODEL_NAME, # output directory
overwrite_output_dir=True,
num_train_epochs=100_000, # total # of training epochs
per_device_train_batch_size=8, # batch size per device during training
per_device_eval_batch_size=8, # batch size for evaluation
warmup_steps=3000, # number of warmup steps for learning rate scheduler
weight_decay=1e-8, # strength of weight decay
learning_rate=1e-3,
logging_dir='./logs2', # directory for storing logs
logging_steps=50,
eval_steps=50,
evaluation_strategy='steps',
save_total_limit=1,
load_best_model_at_end=True,
)
trainer = Trainer(
model=model, # the instantiated 🤗 Transformers model to be trained
args=training_args, # training arguments, defined above
train_dataset=train_sets[k], # training dataset
eval_dataset=eval_dataset, # evaluation dataset
data_collator=data_collator,
tokenizer=tokenizer,
callbacks=[EarlyStoppingCallback(early_stopping_patience=5, early_stopping_threshold=0)]
)
trainer.train()
model.save_pretrained(MODEL_NAME + '_head')
loss_head = trainer.evaluate()['eval_loss']
for param in model.parameters():
param.requires_grad = True
print(f'Tuning model on {k} examples')
training_args = TrAr(
output_dir=MODEL_NAME, # output directory
overwrite_output_dir=True,
num_train_epochs=100_000, # total # of training epochs
per_device_train_batch_size=8, # batch size per device during training
per_device_eval_batch_size=8, # batch size for evaluation
warmup_steps=3000, # number of warmup steps for learning rate scheduler
weight_decay=1e-8, # strength of weight decay
learning_rate=1e-5,
logging_dir='./logs2', # directory for storing logs
logging_steps=50,
eval_steps=50,
evaluation_strategy='steps',
save_total_limit=1,
load_best_model_at_end=True,
)
trainer = Trainer(
model=model, # the instantiated 🤗 Transformers model to be trained
args=training_args, # training arguments, defined above
train_dataset=train_sets[k], # training dataset
eval_dataset=eval_dataset, # evaluation dataset
data_collator=data_collator,
tokenizer=tokenizer,
callbacks=[EarlyStoppingCallback(early_stopping_patience=5, early_stopping_threshold=0)]
)
trainer.train()
loss_full = trainer.evaluate()['eval_loss']
model.save_pretrained(MODEL_NAME + '_full')
cleanup()
print(f'Tuning model on {k} examples completed: losses {loss_head} -> {loss_full}')
print(datetime.now())
```
## Evaluate
```
scores_50_ft = {
'head': [],
'full': []
}
scores_max_ft = {
'head': [],
'full': []
}
for k in [10, 30, 100, 300, 1000, 3000, 10000]:
for mode in ['head', 'full']:
MODEL_NAME = f'./models_scale_ft/roberta_single_{k}_{mode}'
model = AutoModelForTokenClassification.from_pretrained(MODEL_NAME)
cleanup()
model.to(device);
model.eval();
training_args = TrAr(
output_dir='tmp',
per_device_eval_batch_size=64, # batch size for evaluation
)
trainer = Trainer(
model=model, # the instantiated 🤗 Transformers model to be trained
args=training_args, # training arguments, defined above
train_dataset=train_sets[k], # training dataset
data_collator=data_collator,
tokenizer=tokenizer,
)
pred = trainer.predict(final_test_dataset)
scores = []
for threshold in [0.1, 0.15, 0.2, 0.25, 0.3, 0.35, 0.4, 0.45, 0.5, 0.55, 0.6, 0.65, 0.7, 0.75, 0.8, 0.85, 0.9]:
preds = []
for text, pr in zip(final_test.text, pred.predictions):
proba = np.exp(pr[pr[:, 0]!=-100])
proba /= proba.sum(axis=1, keepdims=True)
labels = (proba[:, 1] >= threshold).astype(int).tolist()
preds.append(labels2spans(text, labels, tokenizer))
score = np.mean([f1(p, y) for p, y in zip(preds, final_test.spans)])
scores.append(score)
if threshold == 0.5:
score_50 = score
score_max = max(scores)
print(k, mode, score_50, score_max)
scores_50_ft[mode].append(score_50)
scores_max_ft[mode].append(score_max)
print(scores_50_ft)
#{'head': [0.20996358946778593, 0.19665258203054256, 0.19426943388289686, 0.2249463028813179, 0.2711240067635228, 0.22743156478529175, 0.20691776409080218], 'full': [0.4558391877431416, 0.43972790342989826, 0.5722858622974533, 0.6344381401889334, 0.6527472992730478, 0.6517859231684686, 0.6577082315501016]}
print(scores_max_ft)
# {'head': [0.24778191428625018, 0.24015520569320026, 0.38145677664626454, 0.5484361004401123, 0.5619014200066833, 0.6100396795764801, 0.5795356023564205], 'full': [0.48412770986043197, 0.4410907253856781, 0.5722858622974533, 0.6352267779114283, 0.6586461703712368, 0.657493664504611, 0.6685484505957388]}
plt.plot(xs, scores_max_ft['head'], marker='*')
plt.plot(xs, scores_max_ft['full'], marker='o')
plt.legend(['train only head', 'train full model'])
#plt.scatter(xs, scores_max['head'])
#plt.scatter(xs, scores_max['full'])
plt.xscale('log');
plt.xlabel('training set size');
plt.ylabel('F1 score on the test set');
plt.title('learning curve of a tagger transferred from a classifier');
sm2 = [0.4698994695689647, 0.46052868203784997, 0.5033596644709653, 0.56008830890506, 0.6106484939446047, 0.6681581737239815, 0.6769273 ]
#fig = plt.figure(figsize=(3, 3))
plt.plot(xs, sm2, marker='o')
plt.plot(xs, scores_max_ft['full'], marker='*')
plt.legend(['tagger only', 'classifier -> tagger'])
#plt.scatter(xs, scores_max['head'])
#plt.scatter(xs, scores_max['full'])
plt.xscale('log');
plt.xlabel('training set size');
plt.ylabel('F1 score on the test set');
plt.title('learning curves of a vanilla and transferred taggers');
sm2 = [0.4698994695689647, 0.46052868203784997, 0.5033596644709653, 0.56008830890506, 0.6106484939446047, 0.6681581737239815, 0.6769273 ]
fig, ax = plt.subplots(figsize=(3,3))
plt.plot(xs, scores_max['full'], marker='o')
plt.plot(xs, scores_max_ft['full'], marker='*')
plt.legend(['standard tagger', 'tagging classifier'])
#plt.scatter(xs, scores_max['head'])
#plt.scatter(xs, scores_max['full'])
plt.xscale('log');
plt.xlabel('training set size');
plt.ylabel('F1 score on the test set');
#plt.title('learning curves of a vanilla and transferred taggers');
#ax.xaxis.set_minor_locator(mpl.ticker.MultipleLocator(500));
#ax.yaxis.set_minor_locator(mpl.ticker.MultipleLocator(500));
plt.savefig('images/curve.png', dpi=300, transparent=False, bbox_inches='tight')
sm2 = [0.4698994695689647, 0.46052868203784997, 0.5033596644709653, 0.56008830890506, 0.6106484939446047, 0.6681581737239815, 0.6769273 ]
fig, ax = plt.subplots(figsize=(3,3))
plt.plot(xs, scores_50['full'], marker='o')
plt.plot(xs, scores_50_ft['full'], marker='*')
plt.legend(['RoBERTa tagger', 'tagging classifier'])
#plt.scatter(xs, scores_max['head'])
#plt.scatter(xs, scores_max['full'])
plt.xscale('log');
plt.xlabel('training set size');
plt.ylabel('F1 score on the test set');
#plt.title('learning curves of a vanilla and transferred taggers');
#ax.xaxis.set_minor_locator(mpl.ticker.MultipleLocator(500));
#ax.yaxis.set_minor_locator(mpl.ticker.MultipleLocator(500));
plt.savefig('images/curve.png', dpi=300, transparent=False, bbox_inches='tight')
```
# Toxic vocabulary size
```
def get_substrings(character_ids, text, inverse=False):
""" extract contiguous spans from text given by character_ids"""
if inverse:
character_ids =
if not character_ids:
return []
prev = -100
w = []
result = []
for idx in character_ids:
if prev >=0 and idx > prev + 1:
result.append(''.join(w))
w = []
if idx < len(text):
w.append(text[idx])
prev = idx
if w:
result.append(''.join(w))
return result
print(get_substrings(train.spans[0], train.text[0]))
print()
train_ordered = train.iloc[indices]
familiar = []
vocab_sizes = []
vocab = set()
for i, row in train_ordered.iterrows():
subs = get_substrings(row.spans, row.text.lower())
if subs:
words = [w for s in subs for w in s.strip().split() if w]
familiar.append(np.mean([w in vocab for w in words]))
vocab.update(words)
vocab_sizes.append(len(vocab))
import matplotlib as mpl
fig, ax = plt.subplots(figsize=(3,3))
pd.Series(vocab_sizes).plot();
# plt.plot([0, len(vocab_sizes)], [0, len(vocab)])
# plt.title('Proportion of toxic vocabulary over time');
plt.xlabel('size of training set');
plt.ylabel('size of toxic vocabulary');
ax.xaxis.set_minor_locator(mpl.ticker.MultipleLocator(500));
ax.yaxis.set_minor_locator(mpl.ticker.MultipleLocator(500));
plt.savefig('images/vocab.png', dpi=300, transparent=False, bbox_inches='tight')
s = pd.Series(familiar).rolling(500, min_periods=1).mean()
print(s.max())
s.plot();
plt.title('proportion of toxic words which are familiar')
np.mean(familiar[-500:])
import nltk
sentlen = pd.Series([len(nltk.sent_tokenize(t)) for t in train.text])
sentlen.describe()
sentlen.median()
sentlen.value_counts()
```
| github_jupyter |
# DecisionTreeRegressor with Normalize
This Code template is for regression analysis using simple DecisionTreeRegressor based on the Classification and Regression Trees algorithm along with Normalize Feature Scaling technique.
### Required Packages
```
import warnings
import numpy as np
import pandas as pd
import seaborn as se
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.tree import DecisionTreeRegressor,plot_tree
from sklearn.metrics import r2_score,mean_squared_error,mean_absolute_error
from sklearn.preprocessing import Normalizer
warnings.filterwarnings('ignore')
```
#### Initialization
Filepath of CSV file
```
file_path=""
```
List of features which are required for model training .
```
features=[]
```
Target feature for prediction.
```
target=''
```
### Data Fetching
Pandas is an open-source, BSD-licensed library providing high-performance, easy-to-use data manipulation and data analysis tools.
We will use panda's library to read the CSV file using its storage path.And we use the head function to display the initial row or entry.
```
df=pd.read_csv(file_path);
df.head()
```
### Feature Selections
It is the process of reducing the number of input variables when developing a predictive model. Used to reduce the number of input variables to both reduce the computational cost of modelling and, in some cases, to improve the performance of the model.
We will assign all the required input features to X and target/outcome to Y.
```
X=df[features]
Y=df[target]
```
### Data Preprocessing
Since the majority of the machine learning models in the Sklearn library doesn't handle string category data and Null value, we have to explicitly remove or replace null values. The below snippet have functions, which removes the null value if any exists. And convert the string classes data in the datasets by encoding them to integer classes.
```
def NullClearner(df):
if(isinstance(df, pd.Series) and (df.dtype in ["float64","int64"])):
df.fillna(df.mean(),inplace=True)
return df
elif(isinstance(df, pd.Series)):
df.fillna(df.mode()[0],inplace=True)
return df
else:return df
def EncodeX(df):
return pd.get_dummies(df)
```
Calling preprocessing functions on the feature and target set.
```
x=X.columns.to_list()
for i in x:
X[i]=NullClearner(X[i])
X=EncodeX(X)
Y=NullClearner(Y)
X.head()
```
#### Correlation Map
In order to check the correlation between the features, we will plot a correlation matrix. It is effective in summarizing a large amount of data where the goal is to see patterns.
```
f,ax = plt.subplots(figsize=(18, 18))
matrix = np.triu(X.corr())
se.heatmap(X.corr(), annot=True, linewidths=.5, fmt= '.1f',ax=ax, mask=matrix)#plotting correlation matrix
plt.show()
```
### Data Rescaling
#### Normalizer:
Normalize samples individually to unit norm.
Each sample (i.e. each row of the data matrix) with at least one non zero component is rescaled independently of other samples so that its norm (l1 or l2) equals one.
This transformer is able to work both with dense numpy arrays and scipy.sparse matrix (use CSR format if you want to avoid the burden of a copy / conversion).
Scaling inputs to unit norms is a common operation for text classification or clustering for instance. For instance the dot product of two l2-normalized TF-IDF vectors is the cosine similarity of the vectors and is the base similarity metric for the Vector Space Model commonly used by the Information Retrieval community.
```
X_scaled=Normalizer().fit_transform(X)
X_scaled=pd.DataFrame(data = X_scaled,columns = X.columns)
X_scaled.head()
```
### Data Splitting
The train-test split is a procedure for evaluating the performance of an algorithm. The procedure involves taking a dataset and dividing it into two subsets. The first subset is utilized to fit/train the model. The second subset is used for prediction. The main motive is to estimate the performance of the model on new data.
```
x_train,x_test,y_train,y_test=train_test_split(X_scaled,Y,test_size=0.2,random_state=123)
```
### Model
Decision tree is the most powerful and popular tool for classification and prediction. A Decision tree is a flowchart like tree structure, where each internal node denotes a test on an attribute, each branch represents an outcome of the test, and each leaf node holds a outcome label.
Decision trees can also be applied to regression problems, using the DecisionTreeRegressor class.
As in the classification setting, the fit method will take as argument arrays X and y, only that in this case y is expected to have floating point values instead of integer values
#### Model Tuning Parameter
> - criterion -> The function to measure the quality of a split. Supported criteria are “mse” for the mean squared error, which is equal to variance reduction as feature selection criterion and minimizes the L2 loss using the mean of each terminal node, “friedman_mse”, which uses mean squared error with Friedman’s improvement score for potential splits, “mae” for the mean absolute error, which minimizes the L1 loss using the median of each terminal node, and “poisson” which uses reduction in Poisson deviance to find splits.
> - max_depth -> The maximum depth of the tree. If None, then nodes are expanded until all leaves are pure or until all leaves contain less than min_samples_split samples.
> - max_leaf -> Grow a tree with max_leaf_nodes in best-first fashion. Best nodes are defined as relative reduction in impurity. If None then unlimited number of leaf nodes.
> - max_features -> The number of features to consider when looking for the best split: **{auto , sqrt, log2}**
```
model = DecisionTreeRegressor(random_state=123)
model = model.fit(x_train,y_train)
```
#### Model Accuracy
We will use the trained model to make a prediction on the test set.Then use the predicted value for measuring the accuracy of our model.
The model score function computes the accuracy, either the fraction or the count of correct predictions.
> **score**: The **score** function returns the coefficient of determination <code>R<sup>2</sup></code> of the prediction.
```
print("Accuracy on test: {:.2f} %".format(model.score(x_test, y_test)*100))
```
> **r2_score**: The **r2_score** function computes the percentage variablility explained by our model, either the fraction or the count of correct predictions.
> **mae**: The **mean abosolute error** function calculates the amount of total error(absolute average distance between the real data and the predicted data) by our model.
> **mse**: The **mean squared error** function squares the error(penalizes the model for large errors) by our model.
```
y_pred = model.predict(x_test)
print("R2 Score: {:.2f}%".format(r2_score(y_test,y_pred)*100))
print("Mean Absolute Error {:.2f}".format(mean_absolute_error(y_test,y_pred)))
print("Mean Squared Error {:.2f}".format(mean_squared_error(y_test,y_pred)))
```
#### Feature Importances
The Feature importance refers to techniques that assign a score to features based on how useful they are for making the prediction.
```
plt.figure(figsize=(8,6))
n_features = len(X.columns)
plt.barh(range(n_features), model.feature_importances_, align='center')
plt.yticks(np.arange(n_features), X.columns)
plt.xlabel("Feature importance")
plt.ylabel("Feature")
plt.ylim(-1, n_features)
```
#### Prediction Plot
First, we make use of a plot to plot the actual observations, with x_train on the x-axis and y_train on the y-axis.
For the regression line, we will use x_train on the x-axis and then the predictions of the x_train observations on the y-axis.
```
n=len(x_test) if len(x_test)<20 else 20
plt.figure(figsize=(14,10))
plt.plot(range(n),y_test[0:n], color = "green")
plt.plot(range(n),model.predict(x_test[0:n]), color = "red")
plt.legend(["Actual","prediction"])
plt.title("Predicted vs True Value")
plt.xlabel("Record number")
plt.ylabel(target)
plt.show()
```
#### Prediction Plot
Tree Plot
Plot a decision tree.The visualization is fit automatically to the size of the axis. Use the figsize or dpi arguments of plt.figure to control the size of the rendering.
```
fig, axes = plt.subplots(nrows = 1,ncols = 1,figsize = (3,3), dpi=400)
cls_target = [str(x) for x in pd.unique(y_train)]
cls_target.sort()
plot_tree(model,feature_names = X.columns, class_names=cls_target,filled = True)
fig.savefig('./tree.png')
```
#### Creator:Shreepad Nade , Github: [Profile](https://github.com/shreepad-nade)
| github_jupyter |
```
from IPython.core.display import HTML, display
display(HTML("<style>.container { width:80% !important; }</style>"))
display(HTML("<style>div.output_scroll { height: 44em; }</style>"))
%%capture
# install popmon (if not installed yet)
import sys
!"{sys.executable}" -m pip install popmon
import pandas as pd
import popmon
from popmon import resources
```
# Data generation
Let's first load some data!
```
df = pd.read_csv(
resources.data("flight_delays.csv.gz"), index_col=0, parse_dates=["DATE"]
)
```
# Simple report
Now we can go ahead and generate our first report!
```
df.pm_stability_report(time_axis="DATE")
```
If you inspect the report in the above example, you can see that for example for the maximum `departure_delay` on 2015-08-22 was more extreme than expected.
The time axis is a bit weird now (split into 40 bins of 9 days each), but fortunately we can specify that ourselves using the `time_width` parameter!
We'll also set the `time_offset`, which we set equal to the first data in the document (otherwise we may end up with the first bin containing only half a week of data).
Finally, for the remaining examples, we'll use `extended_report=False` in order to keep the size of the notebook somewhat limited.
```
df.pm_stability_report(
time_axis="DATE", time_width="1w", time_offset="2015-07-02", extended_report=False
)
```
Finally, we could make the thresholds used in the traffic lights more stringent.
For example, we could show the yellow traffic light for deviations bigger than 7 standard deviations, and the red traffic light for deviations bigger than 10 standard deviations.
```
df.pm_stability_report(
time_axis="DATE",
time_width="1w",
time_offset="2015-07-02",
extended_report=False,
pull_rules={"*_pull": [10, 7, -7, -10]},
)
```
There are quite a few more parameters in `pm_stability_report()`, for example to select which features to use (e.g. `features=['x']`), or how to bin the different features (`bin_specs={'x': {'bin_width': 1, 'bin_offset': 0}}`).
We suggest that you check them out on your own!
Have a look at the documentation for `popmon.pipeline.report.df_stability_report()` (which corresponds to `df.pm_stability_report()`).
# What about Spark DataFrames?
No problem! We can easily perform the same steps on a Spark DataFrame. One important thing to note there is that we need to include two jar files (used to create the histograms using Histogrammar) when we create our Spark session.
These will be automatically downloaded the first time you run this command.
```
# download histogrammar jar files if not already installed, used for histogramming of spark dataframe
try:
from pyspark.sql import SparkSession
from pyspark import __version__ as pyspark_version
pyspark_installed = True
except ImportError:
print("pyspark needs to be installed for this example")
pyspark_installed = False
if pyspark_installed:
scala = '2.12' if int(pyspark_version[0]) >= 3 else '2.11'
hist_jar = f'io.github.histogrammar:histogrammar_{scala}:1.0.20'
hist_spark_jar = f'io.github.histogrammar:histogrammar-sparksql_{scala}:1.0.20'
spark = SparkSession.builder.config(
"spark.jars.packages", f'{hist_spark_jar},{hist_jar}'
).getOrCreate()
sdf = spark.createDataFrame(df)
sdf.pm_stability_report(
time_axis="DATE",
time_width="1w",
time_offset="2015-07-02",
extended_report=False,
)
```
# Using other reference types
## Using an external reference
Let's go back to Pandas again! (While all of this functionality also works on Spark DataFrames, it's just faster to illustrate it with Pandas.)
What if we want to compare our DataFrame to another DataFrame?
For example, because we trained a machine learning model on another DataFrame (which we'll call the reference data) and we want to monitor whether the new data (the current DataFrame) comes from a similar distribution?
We can do that by specifying an external reference DataFrame.
```
df_ref = pd.read_csv(
resources.data("flight_delays_reference.csv.gz"), index_col=0, parse_dates=["DATE"]
)
df.pm_stability_report(
time_axis="DATE",
time_width="1w",
time_offset="2015-07-02",
extended_report=False,
reference_type="external",
reference=df_ref,
)
```
## Using an expanding reference
We can also use an expanding reference, which for each time slot uses all preceding time slots as a reference.
```
df.pm_stability_report(
time_axis="DATE",
time_width="1w",
time_offset="2015-07-02",
extended_report=False,
reference_type="expanding",
)
```
## Using a rolling window reference
And finally, we can use a rolling window reference. Here we can play with some additional parameters: shift and window.
We'll set the window parameter to 5.
```
df.pm_stability_report(
time_axis="DATE",
time_width="1w",
time_offset="2015-07-02",
extended_report=False,
reference_type="rolling",
window=5,
)
```
# Plotting the individual histograms
Sometimes, when you're diving into alerts from the report, you may want to plot some individual histograms.
Fortunately, you can! Let's first have a look at how these histograms are stored.
```
report = df.pm_stability_report(
time_axis="DATE", time_width="1w", time_offset="2015-07-02"
)
split_hists = report.datastore["split_hists"]["DEPARTURE_DELAY"]
split_hists
```
Here we see the histograms for each time slot. Let us focus on the first time slot and plot the corresponding histogram.
```
split_hist = split_hists.query("date == '2015-07-05 12:00:00'")
split_hist.histogram[0].plot.matplotlib()
```
And let's also plot the corresponding reference histogram.
```
split_hist.histogram_ref[0].plot.matplotlib()
```
# Saving the report and the histograms to disk
If you run popmon regularly on the same dataset, you may want to store the report and the histograms to disk, so you can keep track of the alerts and easily inspect the histograms if anything goes wrong.
```
# As HTML report
report.to_file("report.html")
# Alternatively, as serialized Python object
# import pickle
# with open("report.pkl", "wb") as f:
# pickle.dump(report, f)
```
# Tuning parameters after generating the report
If you want to tune parameters after you've created the report, you can do so easily using `report.regenerate()`
```
report.regenerate(
last_n=0,
skip_first_n=0,
skip_last_n=0,
plot_hist_n=2,
skip_empty_plots=True,
report_filepath=None,
store_key="html_report",
sections_key="report_sections",
)
```
# Building your own pipelines
The `stability_report()` interface covers many use cases, but if you need more flexibility, you can define your own custom pipeline. We provide an example here!
```
from popmon.hist.hist_splitter import HistSplitter
from popmon.analysis.profiling import HistProfiler
from popmon.pipeline.report import StabilityReport
from popmon.base import Pipeline
from popmon.visualization import SectionGenerator, ReportGenerator
monitoring_rules = {
"*_pull": [7, 4, -4, -7],
"*_zscore": [7, 4, -4, -7],
"[!p]*_unknown_labels": [0.5, 0.5, 0, 0],
}
datastore = {}
datastore["hists"] = df.pm_make_histograms(
time_axis="DATE", time_width="1w", time_offset="2015-07-02"
)
modules = [
HistSplitter(read_key="hists", store_key="split_hists", feature_begins_with="DATE"),
HistProfiler(read_key="split_hists", store_key="profiles"),
SectionGenerator(
section_name="Profiles", read_key="profiles", store_key="report_sections"
),
ReportGenerator(read_key="report_sections", store_key="html_report"),
]
pipeline = Pipeline(modules)
stability_report = StabilityReport()
stability_report.transform(pipeline.transform(datastore))
stability_report
```
The above makes a very simple report, containing only the profiles (and no comparisons, traffic lights or alerts). The next examples shows how you can add the comparisons!
```
from popmon.analysis.comparison.hist_comparer import ReferenceHistComparer
datastore = {}
datastore["hists"] = df.pm_make_histograms(
time_axis="DATE", time_width="1w", time_offset="2015-07-02"
)
modules = [
HistSplitter(read_key="hists", store_key="split_hists", feature_begins_with="DATE"),
HistProfiler(read_key="split_hists", store_key="profiles"),
ReferenceHistComparer(
reference_key="split_hists",
assign_to_key="split_hists",
store_key="comparisons",
),
SectionGenerator(
section_name="Profiles", read_key="profiles", store_key="report_sections"
),
SectionGenerator(
section_name="Comparisons", read_key="comparisons", store_key="report_sections"
),
ReportGenerator(read_key="report_sections", store_key="html_report"),
]
pipeline = Pipeline(modules)
stability_report = StabilityReport()
stability_report.transform(pipeline.transform(datastore))
stability_report
```
If you're interested in more complex examples, check the code in `popmon.pipeline.report_pipelines`.
Using the custom pipelines it becomes relatively easy to include new profiles and new comparisons.
If you do, be sure to let us know! You may be able to make a pull request and add it to the package.
| github_jupyter |
## packages
```
import tensorflow as tf
from tensorflow import keras
import tensorflow_probability as tfp
from tensorflow.keras import layers
from tensorflow.keras.models import load_model
from sklearn.metrics import mean_squared_error
from sklearn.preprocessing import RobustScaler
from tensorflow.keras.preprocessing.sequence import TimeseriesGenerator
import talib as ta
import pandas as pd
import numpy as np
import os
import gc
import traceback
import mysql.connector
import ast
with open('mysql_configure') as f:
mysqlkeys = f.read()
mysqlkeys = ast.literal_eval(mysqlkeys)
mydb = mysql.connector.connect(**mysqlkeys)
version_num = 1
DEBUG=True #load feature added data from crypto-lstm-data
############################################configure
##use GPU if no TPU
DEVICE = "TPU"
SEED = 42
EPOCHS = 10
N_ASSETS = 14 #14 assets
WINDOW_SIZE = 15 #15mins
BATCH_SIZE = 128
PCT_VALIDATION = 10 # last 10% of the data are used as validation set
print(f'N_ASSETS = {N_ASSETS}, WINDOW_SIZE ={WINDOW_SIZE}, BATCH_SIZE = {BATCH_SIZE},EPOCHS = {EPOCHS},PCT_VALIDATION={PCT_VALIDATION}')
if DEVICE == "TPU":
print("connecting to TPU...")
try:
tpu = tf.distribute.cluster_resolver.TPUClusterResolver()
print('Running on TPU ', tpu.master())
except ValueError:
tpu = None
if tpu:
try:
print("initializing TPU ...")
tf.config.experimental_connect_to_cluster(tpu)
tf.tpu.experimental.initialize_tpu_system(tpu)
strategy = tf.distribute.experimental.TPUStrategy(tpu)
print("TPU initialized")
except: print("failed to initialize TPU")
else: DEVICE = "GPU"
if DEVICE != "TPU": strategy = tf.distribute.get_strategy()
if DEVICE == "GPU": print("Num GPUs Available: ", len(tf.config.experimental.list_physical_devices('GPU')))
AUTO = tf.data.experimental.AUTOTUNE
REPLICAS = strategy.num_replicas_in_sync
```
## Data
```
#add name
data_path = './data'
# assets = pd.read_csv(data_path+'/asset_details.csv')
assets = pd.read_sql("select * from asset_details", mydb)
assets_names = dict(zip(assets.Asset_ID, assets.Asset_Name))
#the order of the cryptos collected at same timestamp.
mod_order = pd.read_csv(data_path+'/supplemental_train.csv').Asset_ID[:N_ASSETS]
assets_order = dict((t,i) for i,t in enumerate(mod_order))
if DEBUG:
print('read data from crypto-lstm-data')
train = pd.read_feather('./data/lstm/train_1.ftr')
else:
df_train = pd.read_feather('./data/new_data3.ftr')
train = df_train[df_train.timestamp > df_train.timestamp.quantile(0.9)]
del df_train
gc.collect()
train = train.set_index("timestamp")
print(f"Loaded data range {train.index.values.astype('datetime64[s]')[[0,-1]]}")
train.dropna(axis=0,inplace=True)
train.shape
```
## Add Features
```
fpara_dict={'std_lr_15': 30, 'std_Mkt_lrt_15': 10, 'std_Crypto_Index': 30,
'rsi': 30, 'adx': 50, 'macd_sig': 15, 'macd_s': 10, 'macd_l': 60, 'lrtn': 50,
'fastk2': 10, 'fastk1': 15, 'beta_s': '6h', 'beta_l': '2d', 'vol_sum': 15}
def log_return(series, periods=5):
return np.log(series).diff(periods)
def beta_resid(df, window):
num, unit = int(window[:-1]),window[-1]
if unit == 'h':
width = 60*num
elif unit == 'd':
width = 60*24*num
b = ((ta.MULT(df.Mkt_lrt_15,df.lr_15).rolling(width).mean())/ \
(ta.MULT(df.Mkt_lrt_15,df.Mkt_lrt_15).rolling(width).mean())).rename(f"beta_{window}")
b = b.replace([np.nan,np.inf,-np.inf], 0)
resids = ta.SUB(df.lr_15, ta.MULT(b, df.Mkt_lrt_15)).rename(f"lr_15_resid_{window}")
return pd.concat([b, resids],axis=1)
def lag_features(df,fastk1,fastk2,adx,macd_s,macd_l,macd_sig,rsi,vol_sum,std_Crypto_Index,std_lr_15,std_Mkt_lrt_15,**kwargs):
####TECH indicators
df['slowK'], df['slowD'] = ta.STOCH(df.High, df.Low, df.Close,
fastk_period=fastk1, slowk_period=int(3*fastk1/5), slowd_period=int(3*fastk1/5),
slowk_matype=0, slowd_matype=0)
df['fastK'], df['fastD'] = ta.STOCHF(df.High, df.Low, df.Close,
fastk_period=fastk2, fastd_period=int(3*fastk2/5),
fastd_matype=0)
df[f'rsi_{rsi}'] = ta.RSI(df['Close'], timeperiod=rsi)
df[f'macd_{macd_s}_{macd_l}'],df[f'macd_signal_{macd_sig}'], df['macd_hist'] = \
ta.MACD(df['Close'],fastperiod=macd_s, slowperiod=macd_l, signalperiod=macd_sig)
df[f'adx_{adx}'] = ta.ADX(df['High'], df['Low'],df['Close'], timeperiod=adx)#Average Directional Movement Index
df[f'vol_sum_{vol_sum}'] = ta.SMA(df['Volume'],vol_sum)*vol_sum
####std volatility
df[f'std_lr_15_{std_lr_15}'] = ta.STDDEV(df.lr_15,timeperiod=std_lr_15, nbdev=1)
df[f'std_Mkt_lrt_15_{std_Mkt_lrt_15}'] = ta.STDDEV(df.Mkt_lrt_15,timeperiod=std_Mkt_lrt_15, nbdev=1)
df[f'std_Crypto_Index_{std_Crypto_Index}'] = ta.STDDEV(df.Crypto_Index,timeperiod=std_Crypto_Index, nbdev=1)
def get_features(df_feat, fpara_dict):
pd.options.mode.chained_assignment = None # default='warn'
df_feat[[f"beta_{fpara_dict['beta_s']}",f"lr_15_resid_{fpara_dict['beta_s']}"]] = beta_resid(df_feat, window = fpara_dict['beta_s'])
df_feat[[f"beta_{fpara_dict['beta_l']}",f"lr_15_resid_{fpara_dict['beta_l']}"]] = beta_resid(df_feat, window = fpara_dict['beta_l'])
df_feat[f"lrtn_index_{fpara_dict['lrtn']}"] = log_return(df_feat.Crypto_Index, fpara_dict['lrtn'])
lag_features(df_feat, **fpara_dict)
return df_feat
print('Adding features!')
if DEBUG:
pass
else:
train = train.groupby('Asset_ID').apply(lambda x: get_features(x,fpara_dict))
train.tail()
train.columns
train.head()
```
## Timestamp consistent and Reordering
```
#################### missing timestamp fillin and 14 assets all timestamp
ind = train.index.unique()
def reindex(df):
df = df.reindex(range(ind[0],ind[-1]+60,60),method='nearest')
df = df.fillna(method="ffill").fillna(method="bfill")
return df
#all aassets has same timestamps with missing fillin
train = train.groupby('Asset_ID').apply(reindex).reset_index(0, drop=True).sort_index()
# Memory saving function credit to https://www.kaggle.com/gemartin/load-data-reduce-memory-usage
def reduce_mem_usage(df):
start_mem = df.memory_usage().sum() / 1024**2
print('Memory usage of dataframe is {:.2f} MB'.format(start_mem))
for col in df.columns:
col_type = df[col].dtype.name
if col_type not in ['object', 'category', 'datetime64[ns, UTC]', 'datetime64[ns]']:
c_min = df[col].min()
c_max = df[col].max()
if str(col_type)[:3] == 'int':
if c_min > np.iinfo(np.int8).min and c_max < np.iinfo(np.int8).max:
df[col] = df[col].astype(np.int16)
elif c_min > np.iinfo(np.int16).min and c_max < np.iinfo(np.int16).max:
df[col] = df[col].astype(np.int16)
elif c_min > np.iinfo(np.int32).min and c_max < np.iinfo(np.int32).max:
df[col] = df[col].astype(np.int32)
elif c_min > np.iinfo(np.int64).min and c_max < np.iinfo(np.int64).max:
df[col] = df[col].astype(np.int64)
else:
if c_min > np.finfo(np.float16).min and c_max < np.finfo(np.float16).max:
df[col] = df[col].astype(np.float16)
elif c_min > np.finfo(np.float32).min and c_max < np.finfo(np.float32).max:
df[col] = df[col].astype(np.float32)
else:
df[col] = df[col].astype(np.float64)
end_mem = df.memory_usage().sum() / 1024**2
print('Memory usage after optimization is: {:.2f} MB'.format(end_mem))
print('Decreased by {:.1f}%'.format(100 * (start_mem - end_mem) / start_mem))
return df
train = reduce_mem_usage(train)
gc.collect()
train.head(14)
train.set_index('Asset_ID',append=True, inplace=True)
train = train.reindex(list(zip(train.index.get_level_values('timestamp'),
list(mod_order)*int(train.shape[0]/14))) )
train.tail(14)
mod_order
train.iloc[-14*16,:]
train = train.iloc[:-14*16,:]##fake target
```
## Data format for LSTM
```
#####################################make 3D-numpy arrays for train and targets
targets = train['Target'].to_numpy().reshape(-1, N_ASSETS)
features = ['Count', 'Open', 'High', 'Low', 'Close', 'Volume',
'lr_15', 'Mkt_lrt_15', 'Crypto_Index', 'beta_6h',
'lr_15_resid_6h', 'beta_2d', 'lr_15_resid_2d', 'lrtn_index_50', 'slowK',
'slowD', 'fastK', 'fastD', 'rsi_30', 'macd_10_60', 'macd_signal_15',
'macd_hist', 'adx_50', 'vol_sum_15', 'std_lr_15_30',
'std_Mkt_lrt_15_10', 'std_Crypto_Index_30']
train = train[features].values
train = train.reshape(-1, N_ASSETS, train.shape[-1])
assert targets.shape[0] == train.shape[0], "wrong data shape"
print(targets.shape[0] == train.shape[0])
PCT_VALIDATION = 10 # last 10% of the data are used as validation set
X_train, X_test = train[:-len(train)//PCT_VALIDATION], train[-len(train)//PCT_VALIDATION:]
y_train, y_test = targets[:-len(train)//PCT_VALIDATION], targets[-len(train)//PCT_VALIDATION:]
del train,targets
gc.collect()
######################################################windowing
class sample_generator(keras.utils.Sequence):
def __init__(self, x_set, y_set, batch_size, length):
#(train, targets) arrays tuple
self.x, self.y = x_set, y_set
self.batch_size = batch_size
self.length = length#window size
self.size = len(x_set)#unique timestamp size
def __len__(self):
#number of batchs
return int(np.ceil(len(self.x) / float(self.batch_size)))
def __getitem__(self, idx):
#(train, targets) tuple
batch_x=[]
batch_y=[]
for i in range(self.batch_size):
start_ind = self.batch_size*idx + i
end_ind = start_ind + self.length
if end_ind <= self.size:
batch_x.append(self.x[start_ind : end_ind])
batch_y.append(self.y[end_ind -1])
return np.array(batch_x), np.array(batch_y)
#######################################windowing
train_generator = sample_generator(X_train, y_train, length = WINDOW_SIZE, batch_size = BATCH_SIZE)
val_generator = sample_generator(X_test, y_test, length = WINDOW_SIZE, batch_size = BATCH_SIZE)
```
## LSTM model design
- Lambda layer needed for assets separation;
- Masking layer. Generated records (filled gaps) has zeros as features values, so they are not used in the computations. (masking layer affects saved model to be correctly reloaded)
- Our sequence model architecture(lstm)
- Concatenation layer
- Dense layer
```
########################custom metrics
# def MaxCorrelation(y_true,y_pred):
# return -tf.math.abs(tfp.stats.correlation(y_pred,y_true, sample_axis=None, event_axis=None))
def Correlation(y_true,y_pred):
return tf.math.abs(tfp.stats.correlation(y_pred,y_true, sample_axis=None, event_axis=None))
##########################custom loss func
# def masked_mse(y_true, y_pred):
# mask = tf.math.not_equal(y_true, 0.)
# y_true_masked = tf.boolean_mask(y_true, mask)
# y_pred_masked = tf.boolean_mask(y_pred, mask)
# return tf.keras.losses.mean_squared_error(y_true = y_true_masked, y_pred = y_pred_masked)
# def masked_mae(y_true, y_pred):
# mask = tf.math.not_equal(y_true, 0.)
# y_true_masked = tf.boolean_mask(y_true, mask)
# y_pred_masked = tf.boolean_mask(y_pred, mask)
# return tf.keras.losses.mean_absolute_error(y_true = y_true_masked, y_pred = y_pred_masked)
def masked_cosine(y_true, y_pred):
'''
2.x uses the negative similarity because it's used as a loss function
masking layer cause problem to reload the saved model. Consider the nonmasked below.
'''
mask = tf.math.not_equal(y_true, 0.)
y_true_masked = tf.boolean_mask(y_true, mask)
y_pred_masked = tf.boolean_mask(y_pred, mask)
return tf.keras.losses.cosine_similarity(y_true_masked, y_pred_masked)
def nonmasked_cosine(y_true, y_pred):
'''
2.x uses the negative similarity because it's used as a loss function
'''
return tf.keras.losses.cosine_similarity(y_true, y_pred)
############################define lstm layer
def get_squence_model(x):
x = layers.LSTM(units=32, return_sequences=True)(x)
return x
############################compile model
def get_model(n_assets = 14, trainshape=(15,14,12)):
#Keras tensor
x_input = keras.Input(shape=trainshape)
## parallel sequence model branches
branch_outputs = []
for i in range(n_assets):
# Slicing the ith asset: x_input into x of the lambda function
a = layers.Lambda(lambda x: x[:,:, i])(x_input) #lambda layer
#a = layers.Masking(mask_value = 0., )(a) #masking layer bad for reloading saved model
#a = get_squence_model(a)#lstm layer
a = layers.LSTM(units=32, return_sequences=True)(a)
a = layers.GlobalAvgPool1D()(a)#globalavgpool layer
branch_outputs.append(a)
x = layers.Concatenate()(branch_outputs)
x = layers.Dense(units = 128)(x)#dense layer 0
out = layers.Dense(units = n_assets)(x)#dense layer 1
model = keras.Model(inputs=x_input, outputs=out)
model.compile(optimizer = tf.keras.optimizers.Adam(learning_rate=1e-3),
loss = nonmasked_cosine, metrics=[Correlation])
return model
#######################################model compile enable GPU/TPU
model = get_model(n_assets=N_ASSETS,
trainshape=(train_generator[0][0].shape[1],
N_ASSETS,
train_generator[0][0].shape[-1]))
model.summary()
tf.keras.utils.plot_model(model, show_shapes=True,to_file=f"./pic/lstm-kaggle{version_num}.png")
```
## Start model fitting
```
########################################model fit
tf.random.set_seed(0)
##early stop if 7 epoches no improvement on loss value of the val dataset
estop = keras.callbacks.EarlyStopping(monitor = 'val_loss', patience = 7, verbose = 0,
mode = 'min',restore_best_weights = True)
##how the learning rate of your optimizer changes over time
scheduler = keras.optimizers.schedules.ExponentialDecay(initial_learning_rate=1e-3,
decay_steps=(0.5 * len(X_train) / BATCH_SIZE),
decay_rate = 1e-3)
lr = keras.callbacks.LearningRateScheduler(scheduler, verbose = 1)
del X_train
del X_test
del y_train
del y_test
gc.collect()
print("start model fitting")
history = model.fit(train_generator, validation_data = (val_generator),
epochs = EPOCHS, callbacks = [lr, estop])
print('finished model fitting')
model.save(f"./trainedNN/kaggleLSTM_{version_num}")
```
## model performance
| github_jupyter |
## Flexible models
This toolbox can handle models with fitted model parts. In this demo we will see how this is done.
First we need some imports:
```
import numpy as np
import matplotlib.pyplot as plt
import rsatoolbox
```
As a first step lets generate a few random RDMs, which will serve as our data. We generate 10 RDMs for 5 conditions measured in 20 channels. We will attempt to model these RDMs with a model which takes the first two of these RDMs and is allowed to linearly weight these two. In real applications these model RDMs would of course be generated based on some model or based on some other data than the one we want to fit to.
```
data = np.random.rand(10, 5, 20)
datasets = []
for i_dat in range(data.shape[0]):
dataset = rsatoolbox.data.Dataset(data[i_dat], descriptors={'subj':i_dat})
datasets.append(dataset)
rdms = rsatoolbox.rdm.calc_rdm(datasets, method='euclidean')
model = rsatoolbox.model.ModelWeighted('testModel', rdms[[0,1]])
```
This model object contains the RDMs we gave it as a basis. Its primary two functions are to predict an RDM given a parameter vector $\theta$ and to fit the model, i.e. to find a parameter vector $\theta$ to approximate some data-rdms as closely as possible.
### Predicting RDMs
The model object has two different predict functions: predict and predict_rdm. 'predict' produces a vectorized form of the RDM as a numpy array, which is the minimal representation for computations. 'predict_rdm' produces a typical RDMs object containing the descriptors from the RDMs object passed originally to generate the model object.
Our model object takes 2 parameters as input, which are the two weights for the two rdms. As we see below for $\theta=[1,0]$ and $\theta=[0,1]$ it will return the original rdms, $\theta=[1,1]$ will return the sum of the rdms.
```
# first predicting only the vectors:
print('Predicting with theta = [1,0], should return the first rdm, which is:')
print(rdms[0].get_vectors())
print('The output of the model is:')
print(model.predict([1,0]))
print('Which is indeed identical')
print('')
print('Predicting with theta = [0,1], should return the second rdm, which is:')
print(rdms[1].get_vectors())
print('The output of the model is:')
print(model.predict([0,1]))
print('Which is indeed identical')
print('')
print('Predicting with theta = [1,1], should return the sum of the first two rdms, which is:')
print(rdms[0].get_vectors() + rdms[1].get_vectors())
print('The output of the model is:')
print(model.predict([1,1]))
print('Which is indeed identical')
# Predicting full rdm objects works the same. Here this illustrated only for the [1,0] case:
print('Predicting with theta = [1,0], should return the first rdm, which is:')
print(rdms[0])
print('The output of the model is:')
print(model.predict_rdm([1,0]))
print('Which is the same RDM, but lost the rdm_descriptors as it is now formally a combination of the models RDMs')
```
### Fitting data
To fit data with our model, we use the fitting functions also provided in rsatoolbox.Model. These functions take a model
and some data rdms as input and return an optimal theta value. For a weighted sum model as we created here there are a couple of different fitting algorithms available. Let's try them out:
```
# Let's start with a general optimisation function:
theta_opt = rsatoolbox.model.fit_optimize(model, rdms)
print('Theta based on optimization:')
print(theta_opt)
# As we are dealing with a linear weighting model, we can achieve the same fit using
# fit_regress, which uses the linear algebra solution to the fitting problem,
# which will usually be much faster
theta_regress = rsatoolbox.model.fit_regress(model, rdms)
print('Theta based on fit_regress:')
print(theta_regress)
```
These parameter values are always quite similar, but not identical due to inaccuracy of the optimization.
We can use these parameter values and see that the parameter values we found indeed produce very similar rdms:
```
# get rdm prediction with the fitted parameters:
rdm_opt = model.predict_rdm(theta_opt)
rdm_regress = model.predict_rdm(theta_opt)
# show rdms:
rsatoolbox.vis.show_rdm(rdm_opt)
rsatoolbox.vis.show_rdm(rdm_regress)
```
The model object also has a default fitter attached, which allows us to fit the model using model.fit(data)
```
theta_model = model.fit(rdms)
print(theta_model)
print('the used fitting function was:')
print(model.default_fitter)
```
In this case the default fitting function (`model.default_fitter`) is `fit_optimize`, such that the call we just performed results in exactly the same computations as `rsatoolbox.model.fit_optimize(model, rdms)` did above.
### Optimization criterion
The most important additional input the fitting functions take is `method`, which specifies which comparison method between RDMs is optimized. So far we left this at the default which is the cosine similarity. If we want to optimize a different measure, like the correlation we only need to pass the name:
```
# to maximize the correlation instead we can use either of the following:
theta_corr_regress = rsatoolbox.model.fit_regress(model, rdms, method='corr')
print(theta_corr_regress)
theta_corr_opt = rsatoolbox.model.fit_optimize(model, rdms, method='corr')
print(theta_corr_opt)
theta_corr_model = model.fit(rdms, method='corr')
print(theta_corr_model)
```
Note that these values are very similar to each other, but differ from the ones we got above when optimizing the cosine similarity.
As we now know, what criterion was optimized we can also evaluate this criterion to check that the found parameters are indeed better than others. For example, the parameters found to optimize the correlation should yield a higher average correlation to the data rdms, but a lower cosine similarity than the parameters optimized for the cosine similarity.
```
rdm_cosine = model.predict_rdm(theta_regress)
rdm_corr = model.predict_rdm(theta_corr_regress)
print('The average correlation for the correlation parameters is:')
print(np.mean(rsatoolbox.rdm.compare(rdm_corr, rdms, 'corr')))
print('The average correlation for the cosine similarity parameters is:')
print(np.mean(rsatoolbox.rdm.compare(rdm_cosine, rdms, 'corr')))
print('The average cosine similarity for the correlation parameters is:')
print(np.mean(rsatoolbox.rdm.compare(rdm_corr, rdms, 'cosine')))
print('The average cosine similarity for the cosine similarity parameters is:')
print(np.mean(rsatoolbox.rdm.compare(rdm_cosine, rdms, 'cosine')))
```
## Crossvalidation
## Bootstrap-wrapped crossvalidation
## Other model types
| github_jupyter |
# 1-异常检测
## note:
* [covariance matrix](http://docs.scipy.org/doc/numpy/reference/generated/numpy.cov.html)
* [multivariate_normal](http://docs.scipy.org/doc/numpy/reference/generated/numpy.random.multivariate_normal.html)
* [seaborn bivariate kernel density estimate](https://stanford.edu/~mwaskom/software/seaborn/generated/seaborn.kdeplot.html#seaborn.kdeplot)
```
import matplotlib.pyplot as plt
import seaborn as sns
sns.set(context='notebook',style='white',palette=sns.color_palette("RdBu"))
import numpy as np
import pandas as pd
import scipy.io as scio
from scipy import stats
from sklearn.model_selection import train_test_split
```
You want to divide data into 3 set.
1. Training set
2. Cross Validation set
3. Test set.
You shouldn't be doing prediction using training data or Validation data as it does in the exercise.
```
mat = scio.loadmat('data/ex8data1.mat')
mat.keys()
X = mat.get('X')
```
divide original validation data into validation and test set
```
Xval, Xtest, yval, ytest = train_test_split(mat.get('Xval'),
mat.get('yval').ravel(),
test_size=0.5)
sns.regplot(x = 'Latency', y = 'Throughput',
data=pd.DataFrame(X, columns=['Latency', 'Throughput']),
fit_reg=False,
scatter_kws={"s":20,
"alpha":0.5})
plt.show()
```
## estimate multivariate Gaussian parameters 𝜇 and 𝜎2
> according to data, X1, and X2 is not independent
```
mu = X.mean(axis=0)
print(mu, '\n')
cov = np.cov(X.T)
print(cov)
# example of creating 2d grid to calculate probability density
np.dstack(np.mgrid[0:3,0:3])
# create multi-var Gaussian model
multi_normal = stats.multivariate_normal(mu, cov)
# create a grid
x, y = np.mgrid[0:30:0.01, 0:30:0.01]
pos = np.dstack((x, y))
fig, ax = plt.subplots()
# plot probability density
ax.contourf(x, y, multi_normal.pdf(pos), cmap='Blues')
# plot original data points
sns.regplot(x = 'Latency', y = 'Throughput',
data=pd.DataFrame(X, columns=['Latency', 'Throughput']),
fit_reg=False,
ax=ax,
scatter_kws={"s":10,
"alpha":0.4})
plt.show()
```
## select threshold 𝜖
```
def select_threshold(X, Xval, yval):
"""use CV data to find the best epsilon
Returns:
e: best epsilon with the highest f-score
f-score: such best f-score
"""
# create multivariate model using training data
mu = X.mean(axis=0)
cov = np.cov(X.T)
multi_normal = stats.multivariate_normal(mu, cov)
# this is key, use CV data for fine tuning hyper parameters
pval = multi_normal.pdf(Xval)
# set up epsilon candidates
epsilon = np.linspace(np.min(pval), np.max(pval), num=10000)
# calculate f-score
fs = []
for e in epsilon:
y_pred = (pval <= e).astype('int')
fs.append(f1_score(yval, y_pred))
# find the best f-score
argmax_fs = np.argmax(fs)
return epsilon[argmax_fs], fs[argmax_fs]
from sklearn.metrics import f1_score, classification_report
e, fs = select_threshold(X, Xval, yval)
print('Best epsilon: {}\nBest F-score on validation data: {}'.format(e, fs))
```
## visualize prediction of Xval using learned 𝜖
```
def select_threshold(X, Xval, yval):
"""use CV data to find the best epsilon
Returns:
e: best epsilon with the highest f-score
f-score: such best f-score
"""
# create multivariate model using training data
mu = X.mean(axis=0)
cov = np.cov(X.T)
multi_normal = stats.multivariate_normal(mu, cov)
# this is key, use CV data for fine tuning hyper parameters
pval = multi_normal.pdf(Xval)
# set up epsilon candidates
epsilon = np.linspace(np.min(pval), np.max(pval), num=10000)
# calculate f-score
fs = []
for e in epsilon:
y_pred = (pval <= e).astype('int')
fs.append(f1_score(yval, y_pred))
# find the best f-score
argmax_fs = np.argmax(fs)
return epsilon[argmax_fs], fs[argmax_fs]
def predict(X, Xval, e, Xtest, ytest):
"""with optimal epsilon, combine X, Xval and predict Xtest
Returns:
multi_normal: multivariate normal model
y_pred: prediction of test data
"""
Xdata = np.concatenate((X, Xval), axis=0)
mu = Xdata.mean(axis=0)
cov = np.cov(Xdata.T)
multi_normal = stats.multivariate_normal(mu, cov)
# calculate probability of test data
pval = multi_normal.pdf(Xtest)
y_pred = (pval <= e).astype('int')
print(classification_report(ytest, y_pred))
return multi_normal, y_pred
multi_normal, y_pred = predict(X, Xval, e, Xtest, ytest)
# construct test DataFrame
data = pd.DataFrame(Xtest, columns=['Latency', 'Throughput'])
data['y_pred'] = y_pred
# create a grid for graphing
x, y = np.mgrid[0:30:0.01, 0:30:0.01]
pos = np.dstack((x, y))
fig, ax = plt.subplots()
# plot probability density
ax.contourf(x, y, multi_normal.pdf(pos), cmap='Blues')
# plot original Xval points
sns.regplot(x = 'Latency', y = 'Throughput',
data=data,
fit_reg=False,
ax=ax,
scatter_kws={"s":10,
"alpha":0.4})
# mark the predicted anamoly of CV data. We should have a test set for this...
anamoly_data = data[data['y_pred']==1]
ax.scatter(anamoly_data['Latency'], anamoly_data['Throughput'], marker='x', s=50)
plt.show()
```
## high dimension data
```
mat = scio.loadmat('data/ex8data2.mat')
X = mat.get('X')
Xval, Xtest, yval, ytest = train_test_split(mat.get('Xval'),
mat.get('yval').ravel(),
test_size=0.5)
e, fs = select_threshold(X, Xval, yval)
print('Best epsilon: {}\nBest F-score on validation data: {}'.format(e, fs))
multi_normal, y_pred = predict(X, Xval, e, Xtest, ytest)
print('find {} anamolies'.format(y_pred.sum()))
```
# 2-推荐系统
```
movies_mat = scio.loadmat('data/ex8_movies.mat')
Y, R = movies_mat.get('Y'), movies_mat.get('R')
Y.shape, R.shape
m, u = Y.shape
# m: how many movies
# u: how many users
n = 10 # how many features for a movie
param_mat = scio.loadmat('./data/ex8_movieParams.mat')
theta, X = param_mat.get('Theta'), param_mat.get('X')
theta.shape, X.shape
def serialize(X, theta):
"""serialize 2 matrix
"""
# X (movie, feature), (1682, 10): movie features
# theta (user, feature), (943, 10): user preference
return np.concatenate((X.ravel(), theta.ravel()))
def deserialize(param, n_movie, n_user, n_features):
"""into ndarray of X(1682, 10), theta(943, 10)"""
return param[:n_movie * n_features].reshape(n_movie, n_features), \
param[n_movie * n_features:].reshape(n_user, n_features)
# recommendation fn
def cost(param, Y, R, n_features):
"""compute cost for every r(i, j)=1
Args:
param: serialized X, theta
Y (movie, user), (1682, 943): (movie, user) rating
R (movie, user), (1682, 943): (movie, user) has rating
"""
# theta (user, feature), (943, 10): user preference
# X (movie, feature), (1682, 10): movie features
n_movie, n_user = Y.shape
X, theta = deserialize(param, n_movie, n_user, n_features)
inner = np.multiply(X @ theta.T - Y, R)
return np.power(inner, 2).sum() / 2
def gradient(param, Y, R, n_features):
# theta (user, feature), (943, 10): user preference
# X (movie, feature), (1682, 10): movie features
n_movies, n_user = Y.shape
X, theta = deserialize(param, n_movies, n_user, n_features)
inner = np.multiply(X @ theta.T - Y, R) # (1682, 943)
# X_grad (1682, 10)
X_grad = inner @ theta
# theta_grad (943, 10)
theta_grad = inner.T @ X
# roll them together and return
return serialize(X_grad, theta_grad)
def regularized_cost(param, Y, R, n_features, l=1):
reg_term = np.power(param, 2).sum() * (l / 2)
return cost(param, Y, R, n_features) + reg_term
def regularized_gradient(param, Y, R, n_features, l=1):
grad = gradient(param, Y, R, n_features)
reg_term = l * param
return grad + reg_term
# use subset of data to calculate the cost as in pdf...
users = 4
movies = 5
features = 3
X_sub = X[:movies, :features]
theta_sub = theta[:users, :features]
Y_sub = Y[:movies, :users]
R_sub = R[:movies, :users]
param_sub = serialize(X_sub, theta_sub)
cost(param_sub, Y_sub, R_sub, features)
param = serialize(X, theta) # total real params
cost(serialize(X, theta), Y, R, 10) # this is real total cost
n_movie, n_user = Y.shape
X_grad, theta_grad = deserialize(gradient(param, Y, R, 10),
n_movie, n_user, 10)
assert X_grad.shape == X.shape
assert theta_grad.shape == theta.shape
```
## regularized cost
```
# in the ex8_confi.m, lambda = 1.5, and it's using sub data set
regularized_cost(param_sub, Y_sub, R_sub, features, l=1.5)
regularized_cost(param, Y, R, 10, l=1) # total regularized cost
```
## regularized gradient
```
n_movie, n_user = Y.shape
X_grad, theta_grad = deserialize(regularized_gradient(param, Y, R, 10),
n_movie, n_user, 10)
assert X_grad.shape == X.shape
assert theta_grad.shape == theta.shape
```
## parse movie_id.txt
```
movie_list = []
with open('./data/movie_ids.txt', encoding='latin-1') as f:
for line in f:
tokens = line.strip().split(' ')
movie_list.append(' '.join(tokens[1:]))
movie_list = np.array(movie_list)
```
## reproduce my ratings
```
ratings = np.zeros(1682)
ratings[0] = 4
ratings[6] = 3
ratings[11] = 5
ratings[53] = 4
ratings[63] = 5
ratings[65] = 3
ratings[68] = 5
ratings[97] = 2
ratings[182] = 4
ratings[225] = 5
ratings[354] = 5
```
## prepare data
```
Y, R = movies_mat.get('Y'), movies_mat.get('R')
Y = np.insert(Y, 0, ratings, axis=1) # now I become user 0
Y.shape
R = np.insert(R, 0, ratings != 0, axis=1)
R.shape
n_features = 50
n_movie, n_user = Y.shape
l = 10
X = np.random.standard_normal((n_movie, n_features))
theta = np.random.standard_normal((n_user, n_features))
X.shape, theta.shape
param = serialize(X, theta)
Y_norm = Y - Y.mean() # 这不对吧? 这难道不是每一行减去每一行的均值?
Y_norm.mean()
```
## training
```
import scipy.optimize as opt
res = opt.minimize(fun=regularized_cost,
x0=param,
args=(Y_norm, R, n_features, l),
method='TNC',
jac=regularized_gradient)
#这里很慢
res
X_trained, theta_trained = deserialize(res.x, n_movie, n_user, n_features)
X_trained.shape, theta_trained.shape
prediction = X_trained @ theta_trained.T
my_preds = prediction[:, 0] + Y.mean()
idx = np.argsort(my_preds)[::-1] # Descending order
idx.shape
# top ten idx
my_preds[idx][:10]
for m in movie_list[idx][:10]:
print(m)
```
| github_jupyter |
## Filtering and Annotation Tutorial
### Filter
You can filter the rows of a table with [Table.filter](https://hail.is/docs/0.2/hail.Table.html#hail.Table.filter). This returns a table of those rows for which the expression evaluates to `True`.
```
import hail as hl
hl.utils.get_movie_lens('data/')
users = hl.read_table('data/users.ht')
users.filter(users.occupation == 'programmer').count()
```
We can also express this query in multiple ways using [aggregations](https://hail.is/docs/0.2/tutorials/04-aggregation.html):
```
users.aggregate(hl.agg.filter(users.occupation == 'programmer', hl.agg.count()))
users.aggregate(hl.agg.counter(users.occupation == 'programmer'))[True]
```
### Annotate
You can add new fields to a table with [annotate](https://hail.is/docs/0.2/hail.Table.html#hail.Table.annotate). As an example, let's create a new column called `cleaned_occupation` that replaces missing entries in the `occupation` field labeled as 'other' with 'none.'
```
missing_occupations = hl.set(['other', 'none'])
t = users.annotate(
cleaned_occupation = hl.if_else(missing_occupations.contains(users.occupation),
hl.missing('str'),
users.occupation))
t.show()
```
Compare this to what we had before:
```
users.show()
```
Note: `annotate` is functional: it doesn't mutate `users`, but returns a new table. This is also true of `filter`. In fact, all operations in Hail are functional.
```
users.describe()
```
### Select and Transmute
There are two other annotate methods: [select](https://hail.is/docs/0.2/hail.Table.html#hail.Table.select) and [transmute](https://hail.is/docs/0.2/hail.Table.html#hail.Table.transmute). `select` allows you to create new tables from old ones by selecting existing fields, or creating new ones.
First, let's extract the `sex` and `occupation` fields:
```
users.select(users.sex, users.occupation).show()
```
We can also create a new field that stores the age relative to the average. Note that new fields *must* be assigned a name (in this case `mean_shifted_age`):
```
mean_age = round(users.aggregate(hl.agg.stats(users.age)).mean)
users.select(users.sex, users.occupation, mean_shifted_age = users.age - mean_age).show()
```
`transmute` replaces any fields mentioned on the right-hand side with the new fields, but leaves unmentioned fields unchanged. `transmute` is useful for transforming data into a new form. Compare the following two snippts of code. The second is identical to the first with `transmute` replacing `select.`
```
missing_occupations = hl.set(['other', 'none'])
t = users.select(
cleaned_occupation = hl.if_else(missing_occupations.contains(users.occupation),
hl.missing('str'),
users.occupation))
t.show()
missing_occupations = hl.set(['other', 'none'])
t = users.transmute(
cleaned_occupation = hl.if_else(missing_occupations.contains(users.occupation),
hl.missing('str'),
users.occupation))
t.show()
```
### Global Fields
Finally, you can add global fields with [annotate_globals](https://hail.is/docs/0.2/hail.Table.html#hail.Table.annotate_globals). Globals are useful for storing metadata about a dataset or storing small data structures like sets and maps.
```
t = users.annotate_globals(cohort = 5, cloudable = hl.set(['sample1', 'sample10', 'sample15']))
t.describe()
t.cloudable
hl.eval(t.cloudable)
```
### Exercises
- [Z-score normalize](https://en.wikipedia.org/wiki/Standard_score) the age field of `users`.
- Convert `zip` to an integer. Hint: Not all zipcodes are US zipcodes! Use [hl.int32](https://hail.is/docs/0.2/functions/constructors.html#hail.expr.functions.int32) to convert a string to an integer. Use [StringExpression.matches](https://hail.is/docs/0.2/expressions.html#hail.expr.expressions.StringExpression.matches) to see if a string matches a regular expression.
| github_jupyter |
# Logistic Regression
---
Lets first import required libraries:
```
import pandas as pd
import numpy as np
from sklearn import preprocessing
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import classification_report, confusion_matrix, jaccard_score, log_loss
import itertools
import matplotlib.pyplot as plt
%matplotlib inline
```
## Customer churn with Logistic Regression
A telecommunications company is concerned about the number of customers leaving their land-line business for cable competitors. They need to understand who is leaving. Imagine that you are an analyst at this company and you have to find out who is leaving and why.
<h2 id="about_dataset">About the dataset</h2>
We will use a telecommunications dataset for predicting customer churn. This is a historical customer dataset where each row represents one customer. The data is relatively easy to understand, and you may uncover insights you can use immediately. Typically it is less expensive to keep customers than acquire new ones, so the focus of this analysis is to predict the customers who will stay with the company.
This data set provides information to help you predict what behavior will help you to retain customers. You can analyze all relevant customer data and develop focused customer retention programs.
The dataset includes information about:
- Customers who left within the last month – the column is called Churn
- Services that each customer has signed up for – phone, multiple lines, internet, online security, online backup, device protection, tech support, and streaming TV and movies
- Customer account information – how long they had been a customer, contract, payment method, paperless billing, monthly charges, and total charges
- Demographic info about customers – gender, age range, and if they have partners and dependents
Telco Churn is a hypothetical data file that concerns a telecommunications company's efforts to reduce turnover in its customer base. Each case corresponds to a separate customer and it records various demographic and service usage information. Before you can work with the data, you must use the URL to get the ChurnData.csv.
### Load Data From CSV File
```
churn_df = pd.read_csv("ChurnData.csv")
churn_df.head()
```
<h2 id="preprocessing">Data pre-processing and selection</h2>
Lets select some features for the modeling. Also we change the target data type to be integer, as it is a requirement by the skitlearn algorithm:
```
churn_df = churn_df[['tenure', 'age', 'address', 'income', 'ed', 'employ', 'equip', 'callcard', 'wireless','churn']]
churn_df['churn'] = churn_df['churn'].astype('int')
churn_df.head()
```
How many rows and columns are in this dataset in total? What are the name of columns?
```
print(churn_df.shape)
print(churn_df.head(0))
```
Lets define X, and y for our dataset:
```
X = np.asarray(churn_df[['tenure', 'age', 'address', 'income', 'ed', 'employ', 'equip']])
X[0:5]
y = np.asarray(churn_df['churn'])
y [0:5]
```
Also, we normalize the dataset:
```
X = preprocessing.StandardScaler().fit(X).transform(X)
X[0:5]
```
## Train/Test dataset
Okay, we split our dataset into train and test set:
```
X_train, X_test, y_train, y_test = train_test_split( X, y, test_size=0.2, random_state=4)
print ('Train set:', X_train.shape, y_train.shape)
print ('Test set:', X_test.shape, y_test.shape)
```
<h2 id="modeling">Modeling (Logistic Regression with Scikit-learn)</h2>
Lets build our model using __LogisticRegression__ from Scikit-learn package. This function implements logistic regression and can use different numerical optimizers to find parameters, including ‘newton-cg’, ‘lbfgs’, ‘liblinear’, ‘sag’, ‘saga’ solvers. You can find extensive information about the pros and cons of these optimizers if you search it in internet.
The version of Logistic Regression in Scikit-learn, support regularization. Regularization is a technique used to solve the overfitting problem in machine learning models.
__C__ parameter indicates __inverse of regularization strength__ which must be a positive float. Smaller values specify stronger regularization.
Now lets fit our model with train set:
```
LR = LogisticRegression(C=0.01, solver='liblinear').fit(X_train,y_train)
LR
```
Now we can predict using our test set:
```
yhat = LR.predict(X_test)
print(yhat)
print(y_test)
```
__predict_proba__ returns estimates for all classes, ordered by the label of classes. So, the first column is the probability of class 1, P(Y=1|X), and second column is probability of class 0, P(Y=0|X):
```
yhat_prob = LR.predict_proba(X_test)
```
<h2 id="evaluation">Evaluation</h2>
### jaccard index
Lets try jaccard index for accuracy evaluation. we can define jaccard as the size of the intersection divided by the size of the union of two label sets. If the entire set of predicted labels for a sample strictly match with the true set of labels, then the subset accuracy is 1.0; otherwise it is 0.0.
```
jaccard_score(y_test, yhat)
```
### confusion matrix
Another way of looking at accuracy of classifier is to look at __confusion matrix__.
```
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
print(cm)
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
print(confusion_matrix(y_test, yhat, labels=[1,0]))
# Compute confusion matrix
cnf_matrix = confusion_matrix(y_test, yhat, labels=[1,0])
np.set_printoptions(precision=2)
# Plot non-normalized confusion matrix
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=['churn=1','churn=0'],normalize= False, title='Confusion matrix')
```
Look at first row. The first row is for customers whose actual churn value in test set is 1.
As you can calculate, out of 40 customers, the churn value of 15 of them is 1.
And out of these 15, the classifier correctly predicted 6 of them as 1, and 9 of them as 0.
It means, for 6 customers, the actual churn value were 1 in test set, and classifier also correctly predicted those as 1. However, while the actual label of 9 customers were 1, the classifier predicted those as 0, which is not very good. We can consider it as error of the model for first row.
What about the customers with churn value 0? Lets look at the second row.
It looks like there were 25 customers whom their churn value were 0.
The classifier correctly predicted 24 of them as 0, and one of them wrongly as 1. So, it has done a good job in predicting the customers with churn value 0. A good thing about confusion matrix is that shows the model’s ability to correctly predict or separate the classes. In specific case of binary classifier, such as this example, we can interpret these numbers as the count of true positives, false positives, true negatives, and false negatives.
```
print (classification_report(y_test, yhat))
```
Based on the count of each section, we can calculate precision and recall of each label:
- __Precision__ is a measure of the accuracy provided that a class label has been predicted. It is defined by: precision = TP / (TP + FP)
- __Recall__ is true positive rate. It is defined as: Recall = TP / (TP + FN)
So, we can calculate precision and recall of each class.
__F1 score:__
Now we are in the position to calculate the F1 scores for each label based on the precision and recall of that label.
The F1 score is the harmonic average of the precision and recall, where an F1 score reaches its best value at 1 (perfect precision and recall) and worst at 0. It is a good way to show that a classifer has a good value for both recall and precision.
And finally, we can tell the average accuracy for this classifier is the average of the F1-score for both labels, which is 0.72 in our case.
### log loss
Now, lets try __log loss__ for evaluation. In logistic regression, the output can be the probability of customer churn is yes (or equals to 1). This probability is a value between 0 and 1.
Log loss( Logarithmic loss) measures the performance of a classifier where the predicted output is a probability value between 0 and 1.
```
log_loss(y_test, yhat_prob)
```
## Thanks for Reading :)
Created by [Saeed Aghabozorgi](https://www.linkedin.com/in/saeedaghabozorgi/) and modified by [Tarun Kamboj](https://www.linkedin.com/in/kambojtarun/).
| github_jupyter |
# Linear regression as a statistical estimation problem
### Dr. Tirthajyoti Sarkar, Fremont, CA 94536
---
This notebook demonstrates linear regression as a statistical estimation problem. We will see how to do the following as part of a linear regression modeling,
- Compute statistical properties like standard error, t-statistic, and p-values of the regression coefficients
- Compute F-statistic of the overall regression and infer from the F-test
- Compute $R^2$ and Adjusted-$R^2$ from first principle/definition
- Plot residuals vs. fitted and residuals vs. predictors to check assumptions of linearity and homoscedasticity
- Histogram and Q-Q plot of the standardized residuals to check Normality assumption
The dataset is taken from UCI ML repository. We chose the slump test dataset.
<br>Here is the link: https://archive.ics.uci.edu/ml/datasets/Concrete+Slump+Test
We will use a new Python library for these analyses: `MLR`.
<br>You can read the documentation of this library here: https://mlr.readthedocs.io/en/latest/
```
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
```
### Read the dataset
```
df = pd.read_csv("Data/slump_test.csv",sep=',')
df.drop('No',axis=1,inplace=True)
df.head()
df.shape
```
### Import `MyLinearRegression` from MLR and fit
To fit data from a Pandas DataFrame, just pass on the list of the names of the columns you want to use as predictors, and the name of the response variable column as a string.
```
from mlr.MLR import MyLinearRegression as mlr
m = mlr()
predictors = list(df.columns[:7])
print(predictors)
response = 'Compressive Strength (28-day)(Mpa)'
m.fit_dataframe(X=predictors,y=response,dataframe=df)
```
### Print all the coefficients and the intercept
```
m.coef_
m.intercept_
```
### Print metrics
```
print ("R-squared: ",m.r_squared())
print ("Adjusted R-squared: ",m.adj_r_squared())
print("MSE: ",m.mse())
```
### All metrics at once!
```
m.print_metrics()
```
### Compute $R^2$ and adjusted-$R^2$ from their definition
$R^2$ is a basic metric which tells you about that **how much variance is been explained by the model**. It is calculated by taking the ratio of sum of errors squared (SSR) and total sum of squares (SST).
Now,
$$ SST = SSR + SSE $$
Therefore,
$$ R^2 = \frac{SSR}{SST} = 1 - \frac{SSE}{SST} $$
In a multivariate linear regression, if you keep on adding new variables, the $R^2$ value will always increase irrespective of the variable significance. What adjusted-$R^2$ does is to adjust for these variable additions (penalize the addition of additional variables) from only those variables whose addition in the model which are significant. While doing a multivariate linear regression, we should always look at the adjusted-$R^2$ square.
$$ Adj.R^2 = 1 - \frac{SSE/(n-p-1)}{SST/(n-1)} $$
where $n$ = total number of observation/sample, $p$ = number of predictor variables in the model
```
n = df.shape[0]
p = df.shape[1]-3
r2 = 1-(m.sse()/m.sst())
adjr2 = 1 - (m.sse()/m.sst())*((n-1)/(n-p-1))
print("R^2 from first principles:",round(r2,4))
print("Adjusted-R^2 from first principles:",round(adjr2,4))
```
### AIC and BIC
The [Akaike information criterion (AIC)](https://en.wikipedia.org/wiki/Akaike_information_criterion) is an estimator of the relative quality of statistical models for a given set of data. Given a collection of models for the data, AIC estimates the quality of each model, relative to each of the other models. Thus, AIC provides a means for model selection. AIC was formulated by the statistician Hirotugu Akaike.
AIC is founded on **information theory**. When a statistical model is used to represent the process that generated the data, the representation will almost never be exact; so some information will be lost by using the model to represent the process. AIC estimates the relative amount of information lost by a given model: the less information a model loses, the higher the quality of that model.
BIC stands of **Bayesian information criterion** and follows the same idea as AIC, with only a slight change in the exact formula.
### Residuals plots
```
m.fitted_vs_residual()
m.fitted_vs_features()
```
### Histogram and Q-Q plot of the standardized residuals
From the shape of the histogram, we can judge if the residuals (estimate of original error terms) follow a Normal distribution. Q-Q plot also gives this information. If the Q-Q plot deviates too much from the theoretical 45 degree line, then Normality assumption is violated.
```
m.histogram_resid()
m.qqplot_resid()
```
### F-test of overall significance
```
m.ftest()
```
### Standard errors, t-statistic, p-values
We get all the standard error, t-statistic, and p-values at once. Based on this list, we can say that following predictors do not have statistical significance in the current model,
- Slag
- SP
- Fine Aggr
Why? Because their p-values are > 0.05 and therefore we cannot reject the NULL hypothesis that their true (population) values are actually zero. Note, the first row in the methods corresponds to the intercept of the model, so we show the index from 1 while considering the actual predictors.
```
print("Standard errors:",m.std_err())
print()
print("t-test values:",m.tvalues())
print()
print("P-values:",m.pvalues())
for i in range(7):
print(f"Predictor: {df.columns[i]}, Standard error: {m.std_err()[i+1]}, t-statistic: {m.tvalues()[i+1]}, p-value: {m.pvalues()[i+1]}")
print()
```
### We can print the confidence interval of the regression coefficients directly
Note carefully, the 2nd, 5th, and 7th rows have ranges which include zero. That means the 95% confidence interval include zero i.e. we cannot reject the NULL hypothesis that these coefficients are actually zero, and therefore have no impact on the response variable. The first row corresponds to the intercept of the model, so we show the index from 1 while considering the actual predictors.
These are the same variables - Slag, SP, and Fine Aggr., which showed p-values > 0.05.
```
m.conf_int()[1:]
```
### If we change the statistical significance level to 0.01 from 0.05, then two more variables show range including zero
```
m.conf_int(alpha=0.01)[1:]
```
### Now, we can build a model removing those three variables who showed p-val > 0.05
```
m2 = mlr()
predictors = ['Cement', 'Fly ash', 'Water', 'Coarse Aggr.']
m2.fit_dataframe(X=predictors,y=response,dataframe=df)
print("Metrics of the old (full) model\n"+"-"*40)
m.print_metrics()
print("Metrics of the new (smaller) model\n"+"-"*40)
m2.print_metrics()
```
### We can also plot something called Cook's distance plot to see if there is any outliers in the data
```
m.cook_distance()
```
### We can plot the full pairwise scatterplots
```
m.pairplot()
```
### You can also use Seaborn library for visualization like pairplots and correlation heatmaps
```
import seaborn as sns
sns.pairplot(data=df[df.columns[:7]])
corr = np.corrcoef(df[df.columns[:7]],rowvar=False)
plt.figure(figsize=(10,10))
sns.heatmap(data=corr,linewidths=1,annot=True)
```
| github_jupyter |
## Tercera parte pandas
- Operaciones con fechas
- Combinar dataframes
- Reacomodar datos
```
%pylab inline
import pandas as pd
# Cargar nuestra base de datos de elencos
elenco = pd.read_csv('data/cast.csv', encoding='utf-8')
elenco.head()
# Ahora tambien cargaremos datos de otra base de datos
#
fecha_lanz = pd.read_csv('data/release_dates.csv', encoding='utf-8')
fecha_lanz.head()
# Que tal si cargamos esta base de datos pero la columna date la convertirmos a un
# formato datetime que tiene metodos para hacer operaciones sobre fechas.. Mejor!
fecha_lanz = pd.read_csv('data/release_dates.csv' , parse_dates=['date'], infer_datetime_format=True)
fecha_lanz.head()
# Podemos acceder a los metodos del namespace datetime, en pandas "dt"
fl = fecha_lanz
fl.date.dt.year
# Con esto que podemos hacer un listado categorizando por algun elemento de la
# fecha como por ejemplo:
# En que meses se lanzan mas comunmente peliculas que contienen la palabra
# "horror"
fl = fecha_lanz
fl = fl[(fl.title.str.contains('horror',case=False)) & (fl.country == 'USA')]
fl.date.dt.month.value_counts().sort_index().plot(kind='bar')
# Exploremos la funcionalidad para unir dataframes con el metodo "merge"
e = elenco
e = e[e.name == "Denzel Washington"]
e.head()
# Nos interesa conocer la fecha de lanzamiento de las peliculas en la que ah participado
# Denzel Washington.. Pero esa información esta en otro dataframe!!
fl = fecha_lanz
fl.head()
# Es realmente sencillo con pandas y su poderoso comando "merge"
e = elenco
e = e[e.name == "Denzel Washington"]
e = e.merge(fecha_lanz)
e
# Pero como funciona?
# Empata para ambas dimensiones --> keys e indices las coincidencias y
# agrega las entradas que no estan disponibles en el primer dataframe
# Por default nos funciono, pero el comando es muy flexible.
# Mas información: http://pandas.pydata.org/pandas-docs/stable/merging.html
```
## Otro concepto: Pivot
```
# Pivot permite el reacomodo de los valores del dataframe, moviendo una columna como
# indice, otra como llave de datos y otra para los valores
# Ejemplo fechas de lanzamiento de peliculas Harry Potter en Mexico, USA y Canada
fl = fecha_lanz
fl = fl[fl.title.str.startswith("Harry Potter")]
fl = fl[(fl.country == "Mexico") | (fl.country == "USA") | (fl.country == "Canada")]
fl
# Reacomodemos los datos,
# : title como indice
# : country como columnas
# : date como valores
fl = fecha_lanz
fl = fl[fl.title.str.startswith("Harry Potter")]
fl = fl[(fl.country == "Mexico") | (fl.country == "USA") | (fl.country == "Canada")]
fl = fl.pivot('title','country','date')
fl
# Extra
#fl.apply(lambda x : x.dt.month)
```
## Ejercicios
```
# Del periodo 2010 al presente en cuantas peliculas ah participado
# la actriz ""
# En que fechas fue lanzada cada pelicula de "Salma Hayek"
# dentro de un periodo de 1990 al presente en Mexico
fl = fecha_lanz
fl = fl[fl.country == 'USA']
e = elenco
e = e[e.name == 'Salma Hayek']
e = e[(e.year > 1990)]
e = e.merge(fl)
#e
# Una nueva nueva columna con los valores del mes
e['month'] = e.date.dt.month
e
# Graficado
e.month.value_counts().sort_index().plot(kind='bar',ylim=[0,10])
# En que dia de la semana se han lanzado en USA las peliculas de
# donde aparece Tom Cuise.. Incluir un grafico
fl = fecha_lanz
fl = fl[fl.country == 'USA']
e = elenco
e = e[ (e.name == 'Tom Cruise') ]
wd = e.merge(fl).date.dt.dayofweek.value_counts().sort_index()
wd
wd = wd.rename({4:'Friday', 3:'Thursday', 2:'Wednesday'})
wd.plot(kind='bar')
# Que peliculas han sido lanzadas en la mayor cantidad de
# paises , incluir en un grafico las 5 mayores
fl = fecha_lanz
fl = fl.groupby(['title','year'])
fl.size().sort_values(ascending=False)[:5].plot(kind='barh')
# Quien tiende a ser el actor si clasificado (n > 1) que mas a
# participado en peliculas donde el protagonista es
# Leonardo DiCaprio
e = elenco
hf = e[(e.name == 'Leonardo DiCaprio') & (e.n == 1)]
er = e[(e.n > 1)]
mt = hf.merge(er,on=['title','year'])
mt.groupby(['name_x','name_y']).size().sort_values(ascending=False)
```
| github_jupyter |
# Initialization
Welcome to the first assignment of "Improving Deep Neural Networks".
Training your neural network requires specifying an initial value of the weights. A well chosen initialization method will help learning.
If you completed the previous course of this specialization, you probably followed our instructions for weight initialization, and it has worked out so far. But how do you choose the initialization for a new neural network? In this notebook, you will see how different initializations lead to different results.
A well chosen initialization can:
- Speed up the convergence of gradient descent
- Increase the odds of gradient descent converging to a lower training (and generalization) error
To get started, run the following cell to load the packages and the planar dataset you will try to classify.
```
import numpy as np
import matplotlib.pyplot as plt
import sklearn
import sklearn.datasets
from init_utils import sigmoid, relu, compute_loss, forward_propagation, backward_propagation
from init_utils import update_parameters, predict, load_dataset, plot_decision_boundary, predict_dec
%matplotlib inline
plt.rcParams['figure.figsize'] = (7.0, 4.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
# load image dataset: blue/red dots in circles
train_X, train_Y, test_X, test_Y = load_dataset()
```
You would like a classifier to separate the blue dots from the red dots.
## 1 - Neural Network model
You will use a 3-layer neural network (already implemented for you). Here are the initialization methods you will experiment with:
- *Zeros initialization* -- setting `initialization = "zeros"` in the input argument.
- *Random initialization* -- setting `initialization = "random"` in the input argument. This initializes the weights to large random values.
- *He initialization* -- setting `initialization = "he"` in the input argument. This initializes the weights to random values scaled according to a paper by He et al., 2015.
**Instructions**: Please quickly read over the code below, and run it. In the next part you will implement the three initialization methods that this `model()` calls.
```
def model(X, Y, learning_rate = 0.01, num_iterations = 15000, print_cost = True, initialization = "he"):
"""
Implements a three-layer neural network: LINEAR->RELU->LINEAR->RELU->LINEAR->SIGMOID.
Arguments:
X -- input data, of shape (2, number of examples)
Y -- true "label" vector (containing 0 for red dots; 1 for blue dots), of shape (1, number of examples)
learning_rate -- learning rate for gradient descent
num_iterations -- number of iterations to run gradient descent
print_cost -- if True, print the cost every 1000 iterations
initialization -- flag to choose which initialization to use ("zeros","random" or "he")
Returns:
parameters -- parameters learnt by the model
"""
grads = {}
costs = [] # to keep track of the loss
m = X.shape[1] # number of examples
layers_dims = [X.shape[0], 10, 5, 1]
# Initialize parameters dictionary.
if initialization == "zeros":
parameters = initialize_parameters_zeros(layers_dims)
elif initialization == "random":
parameters = initialize_parameters_random(layers_dims)
elif initialization == "he":
parameters = initialize_parameters_he(layers_dims)
# Loop (gradient descent)
for i in range(0, num_iterations):
# Forward propagation: LINEAR -> RELU -> LINEAR -> RELU -> LINEAR -> SIGMOID.
a3, cache = forward_propagation(X, parameters)
# Loss
cost = compute_loss(a3, Y)
# Backward propagation.
grads = backward_propagation(X, Y, cache)
# Update parameters.
parameters = update_parameters(parameters, grads, learning_rate)
# Print the loss every 1000 iterations
if print_cost and i % 1000 == 0:
print("Cost after iteration {}: {}".format(i, cost))
costs.append(cost)
# plot the loss
plt.plot(costs)
plt.ylabel('cost')
plt.xlabel('iterations (per hundreds)')
plt.title("Learning rate =" + str(learning_rate))
plt.show()
return parameters
```
## 2 - Zero initialization
There are two types of parameters to initialize in a neural network:
- the weight matrices $(W^{[1]}, W^{[2]}, W^{[3]}, ..., W^{[L-1]}, W^{[L]})$
- the bias vectors $(b^{[1]}, b^{[2]}, b^{[3]}, ..., b^{[L-1]}, b^{[L]})$
**Exercise**: Implement the following function to initialize all parameters to zeros. You'll see later that this does not work well since it fails to "break symmetry", but lets try it anyway and see what happens. Use np.zeros((..,..)) with the correct shapes.
```
# GRADED FUNCTION: initialize_parameters_zeros
def initialize_parameters_zeros(layers_dims):
"""
Arguments:
layer_dims -- python array (list) containing the size of each layer.
Returns:
parameters -- python dictionary containing your parameters "W1", "b1", ..., "WL", "bL":
W1 -- weight matrix of shape (layers_dims[1], layers_dims[0])
b1 -- bias vector of shape (layers_dims[1], 1)
...
WL -- weight matrix of shape (layers_dims[L], layers_dims[L-1])
bL -- bias vector of shape (layers_dims[L], 1)
"""
parameters = {}
L = len(layers_dims) # number of layers in the network
for l in range(1, L):
### START CODE HERE ### (≈ 2 lines of code)
parameters['W' + str(l)] = np.zeros((layers_dims[l], layers_dims[l-1]))
parameters['b' + str(l)] = np.zeros((layers_dims[l], 1))
### END CODE HERE ###
return parameters
parameters = initialize_parameters_zeros([3,2,1])
print("W1 = " + str(parameters["W1"]))
print("b1 = " + str(parameters["b1"]))
print("W2 = " + str(parameters["W2"]))
print("b2 = " + str(parameters["b2"]))
```
**Expected Output**:
<table>
<tr>
<td>
**W1**
</td>
<td>
[[ 0. 0. 0.]
[ 0. 0. 0.]]
</td>
</tr>
<tr>
<td>
**b1**
</td>
<td>
[[ 0.]
[ 0.]]
</td>
</tr>
<tr>
<td>
**W2**
</td>
<td>
[[ 0. 0.]]
</td>
</tr>
<tr>
<td>
**b2**
</td>
<td>
[[ 0.]]
</td>
</tr>
</table>
Run the following code to train your model on 15,000 iterations using zeros initialization.
```
parameters = model(train_X, train_Y, initialization = "zeros")
print ("On the train set:")
predictions_train = predict(train_X, train_Y, parameters)
print ("On the test set:")
predictions_test = predict(test_X, test_Y, parameters)
```
The performance is really bad, and the cost does not really decrease, and the algorithm performs no better than random guessing. Why? Lets look at the details of the predictions and the decision boundary:
```
print ("predictions_train = " + str(predictions_train))
print ("predictions_test = " + str(predictions_test))
plt.title("Model with Zeros initialization")
axes = plt.gca()
axes.set_xlim([-1.5,1.5])
axes.set_ylim([-1.5,1.5])
plot_decision_boundary(lambda x: predict_dec(parameters, x.T), train_X, train_Y)
```
The model is predicting 0 for every example.
In general, initializing all the weights to zero results in the network failing to break symmetry. This means that every neuron in each layer will learn the same thing, and you might as well be training a neural network with $n^{[l]}=1$ for every layer, and the network is no more powerful than a linear classifier such as logistic regression.
<font color='blue'>
**What you should remember**:
- The weights $W^{[l]}$ should be initialized randomly to break symmetry.
- It is however okay to initialize the biases $b^{[l]}$ to zeros. Symmetry is still broken so long as $W^{[l]}$ is initialized randomly.
## 3 - Random initialization
To break symmetry, lets intialize the weights randomly. Following random initialization, each neuron can then proceed to learn a different function of its inputs. In this exercise, you will see what happens if the weights are intialized randomly, but to very large values.
**Exercise**: Implement the following function to initialize your weights to large random values (scaled by \*10) and your biases to zeros. Use `np.random.randn(..,..) * 10` for weights and `np.zeros((.., ..))` for biases. We are using a fixed `np.random.seed(..)` to make sure your "random" weights match ours, so don't worry if running several times your code gives you always the same initial values for the parameters.
```
# GRADED FUNCTION: initialize_parameters_random
def initialize_parameters_random(layers_dims):
"""
Arguments:
layer_dims -- python array (list) containing the size of each layer.
Returns:
parameters -- python dictionary containing your parameters "W1", "b1", ..., "WL", "bL":
W1 -- weight matrix of shape (layers_dims[1], layers_dims[0])
b1 -- bias vector of shape (layers_dims[1], 1)
...
WL -- weight matrix of shape (layers_dims[L], layers_dims[L-1])
bL -- bias vector of shape (layers_dims[L], 1)
"""
np.random.seed(3) # This seed makes sure your "random" numbers will be the as ours
parameters = {}
L = len(layers_dims) # integer representing the number of layers
for l in range(1, L):
### START CODE HERE ### (≈ 2 lines of code)
parameters['W' + str(l)] = np.random.randn(layers_dims[l], layers_dims[l-1]) * 10
parameters['b' + str(l)] = np.zeros((layers_dims[l], 1))
### END CODE HERE ###
return parameters
parameters = initialize_parameters_random([3, 2, 1])
print("W1 = " + str(parameters["W1"]))
print("b1 = " + str(parameters["b1"]))
print("W2 = " + str(parameters["W2"]))
print("b2 = " + str(parameters["b2"]))
```
**Expected Output**:
<table>
<tr>
<td>
**W1**
</td>
<td>
[[ 17.88628473 4.36509851 0.96497468]
[-18.63492703 -2.77388203 -3.54758979]]
</td>
</tr>
<tr>
<td>
**b1**
</td>
<td>
[[ 0.]
[ 0.]]
</td>
</tr>
<tr>
<td>
**W2**
</td>
<td>
[[-0.82741481 -6.27000677]]
</td>
</tr>
<tr>
<td>
**b2**
</td>
<td>
[[ 0.]]
</td>
</tr>
</table>
Run the following code to train your model on 15,000 iterations using random initialization.
```
parameters = model(train_X, train_Y, initialization = "random")
print ("On the train set:")
predictions_train = predict(train_X, train_Y, parameters)
print ("On the test set:")
predictions_test = predict(test_X, test_Y, parameters)
```
If you see "inf" as the cost after the iteration 0, this is because of numerical roundoff; a more numerically sophisticated implementation would fix this. But this isn't worth worrying about for our purposes.
Anyway, it looks like you have broken symmetry, and this gives better results. than before. The model is no longer outputting all 0s.
```
print (predictions_train)
print (predictions_test)
plt.title("Model with large random initialization")
axes = plt.gca()
axes.set_xlim([-1.5,1.5])
axes.set_ylim([-1.5,1.5])
plot_decision_boundary(lambda x: predict_dec(parameters, x.T), train_X, train_Y)
```
**Observations**:
- The cost starts very high. This is because with large random-valued weights, the last activation (sigmoid) outputs results that are very close to 0 or 1 for some examples, and when it gets that example wrong it incurs a very high loss for that example. Indeed, when $\log(a^{[3]}) = \log(0)$, the loss goes to infinity.
- Poor initialization can lead to vanishing/exploding gradients, which also slows down the optimization algorithm.
- If you train this network longer you will see better results, but initializing with overly large random numbers slows down the optimization.
<font color='blue'>
**In summary**:
- Initializing weights to very large random values does not work well.
- Hopefully intializing with small random values does better. The important question is: how small should be these random values be? Lets find out in the next part!
## 4 - He initialization
Finally, try "He Initialization"; this is named for the first author of He et al., 2015. (If you have heard of "Xavier initialization", this is similar except Xavier initialization uses a scaling factor for the weights $W^{[l]}$ of `sqrt(1./layers_dims[l-1])` where He initialization would use `sqrt(2./layers_dims[l-1])`.)
**Exercise**: Implement the following function to initialize your parameters with He initialization.
**Hint**: This function is similar to the previous `initialize_parameters_random(...)`. The only difference is that instead of multiplying `np.random.randn(..,..)` by 10, you will multiply it by $\sqrt{\frac{2}{\text{dimension of the previous layer}}}$, which is what He initialization recommends for layers with a ReLU activation.
```
# GRADED FUNCTION: initialize_parameters_he
def initialize_parameters_he(layers_dims):
"""
Arguments:
layer_dims -- python array (list) containing the size of each layer.
Returns:
parameters -- python dictionary containing your parameters "W1", "b1", ..., "WL", "bL":
W1 -- weight matrix of shape (layers_dims[1], layers_dims[0])
b1 -- bias vector of shape (layers_dims[1], 1)
...
WL -- weight matrix of shape (layers_dims[L], layers_dims[L-1])
bL -- bias vector of shape (layers_dims[L], 1)
"""
np.random.seed(3)
parameters = {}
L = len(layers_dims) - 1 # integer representing the number of layers
for l in range(1, L + 1):
### START CODE HERE ### (≈ 2 lines of code)
parameters['W' + str(l)] = np.random.randn(layers_dims[l], layers_dims[l-1]) * np.sqrt(2 / layers_dims[l - 1])
parameters['b' + str(l)] = np.zeros((layers_dims[l], 1))
### END CODE HERE ###
return parameters
parameters = initialize_parameters_he([2, 4, 1])
print("W1 = " + str(parameters["W1"]))
print("b1 = " + str(parameters["b1"]))
print("W2 = " + str(parameters["W2"]))
print("b2 = " + str(parameters["b2"]))
```
**Expected Output**:
<table>
<tr>
<td>
**W1**
</td>
<td>
[[ 1.78862847 0.43650985]
[ 0.09649747 -1.8634927 ]
[-0.2773882 -0.35475898]
[-0.08274148 -0.62700068]]
</td>
</tr>
<tr>
<td>
**b1**
</td>
<td>
[[ 0.]
[ 0.]
[ 0.]
[ 0.]]
</td>
</tr>
<tr>
<td>
**W2**
</td>
<td>
[[-0.03098412 -0.33744411 -0.92904268 0.62552248]]
</td>
</tr>
<tr>
<td>
**b2**
</td>
<td>
[[ 0.]]
</td>
</tr>
</table>
Run the following code to train your model on 15,000 iterations using He initialization.
```
parameters = model(train_X, train_Y, initialization = "he")
print ("On the train set:")
predictions_train = predict(train_X, train_Y, parameters)
print ("On the test set:")
predictions_test = predict(test_X, test_Y, parameters)
plt.title("Model with He initialization")
axes = plt.gca()
axes.set_xlim([-1.5,1.5])
axes.set_ylim([-1.5,1.5])
plot_decision_boundary(lambda x: predict_dec(parameters, x.T), train_X, train_Y)
```
**Observations**:
- The model with He initialization separates the blue and the red dots very well in a small number of iterations.
## 5 - Conclusions
You have seen three different types of initializations. For the same number of iterations and same hyperparameters the comparison is:
<table>
<tr>
<td>
**Model**
</td>
<td>
**Train accuracy**
</td>
<td>
**Problem/Comment**
</td>
</tr>
<td>
3-layer NN with zeros initialization
</td>
<td>
50%
</td>
<td>
fails to break symmetry
</td>
<tr>
<td>
3-layer NN with large random initialization
</td>
<td>
83%
</td>
<td>
too large weights
</td>
</tr>
<tr>
<td>
3-layer NN with He initialization
</td>
<td>
99%
</td>
<td>
recommended method
</td>
</tr>
</table>
<font color='blue'>
**What you should remember from this notebook**:
- Different initializations lead to different results
- Random initialization is used to break symmetry and make sure different hidden units can learn different things
- Don't intialize to values that are too large
- He initialization works well for networks with ReLU activations.
| github_jupyter |
```
import os, sys
import paddle
sys.path.append('/workspace/fnet_paddle/PaddleNLP')
from paddlenlp.datasets import load_dataset
test_ds = load_dataset("glue", name="cola", splits=("test"))
len(test_ds)
def convert_example(example,
tokenizer,
max_seq_length=512,
is_test=False):
text = example["sentence"]
text_pair = None
encoded_inputs = tokenizer(
text=text, text_pair=text_pair, max_seq_len=max_seq_length)
input_ids = encoded_inputs["input_ids"]
token_type_ids = encoded_inputs["token_type_ids"]
if is_test:
return input_ids, token_type_ids
label = np.array([example["labels"]], dtype="int64")
return input_ids, token_type_ids, label
def create_dataloader(dataset,
mode='train',
batch_size=1,
batchify_fn=None,
trans_fn=None):
if trans_fn:
dataset = dataset.map(trans_fn)
shuffle = True if mode == 'train' else False
if mode == 'train':
batch_sampler = paddle.io.DistributedBatchSampler(
dataset, batch_size=batch_size, shuffle=shuffle)
else:
batch_sampler = paddle.io.BatchSampler(
dataset, batch_size=batch_size, shuffle=shuffle)
return paddle.io.DataLoader(
dataset=dataset,
batch_sampler=batch_sampler,
collate_fn=batchify_fn,
return_list=True)
import argparse
import os
import paddle
import paddle.nn.functional as F
import paddlenlp as ppnlp
from paddlenlp.data import Tuple, Pad
from functools import partial
parser = argparse.ArgumentParser()
parser.add_argument("--params_path", type=str, required=False, default="checkpoints/model_900/model_state.pdparams", help="The path to model parameters to be loaded.")
parser.add_argument("--max_seq_length", type=int, default=128, help="The maximum total input sequence length after tokenization. "
"Sequences longer than this will be truncated, sequences shorter will be padded.")
parser.add_argument("--batch_size", type=int, default=32, help="Batch size per GPU/CPU for training.")
parser.add_argument('--device', choices=['cpu', 'gpu', 'xpu', 'npu'], default="gpu", help="Select which device to train model, defaults to gpu.")
args = parser.parse_args([])
args.params_path = 'checkpoints/model_400/model_state.pdparams'
fnet = ppnlp.transformers.FNetModel.from_pretrained('pretrained_model/paddle/large')
model = ppnlp.transformers.FNetForSequenceClassification(fnet, num_classes=len(test_ds.label_list))
tokenizer = ppnlp.transformers.FNetTokenizer.from_pretrained('fnet-large')
if args.params_path and os.path.isfile(args.params_path):
state_dict = paddle.load(args.params_path)
model.set_dict(state_dict)
print("Loaded parameters from %s" % args.params_path)
trans_func = partial(
convert_example,
tokenizer=tokenizer,
max_seq_length=args.max_seq_length,
is_test=True)
batchify_fn = lambda samples, fn=Tuple(
Pad(axis=0, pad_val=tokenizer.pad_token_id), # input
Pad(axis=0, pad_val=tokenizer.pad_token_type_id), # segment
): [data for data in fn(samples)]
test_data_loader = create_dataloader(
test_ds,
mode='test',
batch_size=args.batch_size,
batchify_fn=batchify_fn,
trans_fn=trans_func)
results = []
model.eval()
for batch in test_data_loader:
input_ids, token_type_ids = batch
logits = model(input_ids, token_type_ids)
probs = F.softmax(logits, axis=1)
idx = paddle.argmax(probs, axis=1).numpy()
idx = idx.tolist()
results.extend(idx)
results[:3]
import pandas as pd
res_df = pd.DataFrame()
res_df['prediction'] = results
res_df.index.name = 'index'
res_df.sample(5)
(res_df['prediction'] == 0).sum()
res_df.to_csv('CoLA.tsv', sep='\t')
```
| github_jupyter |
```
# get current timestamp for proper documentation of testing and validation results
from datetime import datetime
currentTime = str(datetime.now())
model_save_name = 'causal_classifier_' + currentTime + '.bin'
#path = F"/content/gdrive/My Drive/Causality Classification/"
```
## Setup
Load the transformers library from HuggingFace.
```
import sys, os
!{sys.executable} -m pip install --user watermark
!{sys.executable} -m pip install --user transformers
!{sys.executable} -m pip install --user spacy
# Make sure that python can load the required packages
# we need to specify the location for which the python packages will be installed
sys.path.insert(0, os.path.expanduser('~/.local/lib/python3.6/site-packages'))
!python3 -m spacy download en_core_web_sm
# Use Pos Tagging ?
use_pos_tags = False
import transformers
from transformers import BertModel, BertTokenizer, AdamW, get_linear_schedule_with_warmup
import torch
import numpy as np
import pandas as pd
import seaborn as sns
from pylab import rcParams
import matplotlib.pyplot as plt
from matplotlib import rc
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix, classification_report
from collections import defaultdict
from textwrap import wrap
from torch import nn, optim
from torch.utils.data import Dataset, DataLoader
import torch.nn.functional as F
# packages for adding pos tags
import spacy
%matplotlib inline
%config InlineBackend.figure_format='retina'
sns.set(style='whitegrid', palette='muted', font_scale=1.2)
HAPPY_COLORS_PALETTE = ["#01BEFE", "#FFDD00", "#FF7D00", "#FF006D", "#ADFF02", "#8F00FF"]
sns.set_palette(sns.color_palette(HAPPY_COLORS_PALETTE))
rcParams['figure.figsize'] = 12, 8
RANDOM_SEED = 42
np.random.seed(RANDOM_SEED)
torch.manual_seed(RANDOM_SEED)
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
device
```
## Data Exploration
Import Causal Sentences from our annotation platform.
```
df = pd.read_csv("annotation_causal.csv", encoding='utf-8')
# As the sentences are labeled by multiple annotators, we need to merge all labels into one colum.
# looks like = 1,1,1,1,1...
df['Label'] = df[df.columns[1:7]].apply(
lambda x: ','.join(x.dropna().astype(str)),
axis=1
)
# After that we just need to use the first label of the merged colum.
# Different labeling decisions are already discussed in our labeling process.
for index, row in df.iterrows():
df.at[index,'Label'] = int(float(row['Label'][0:3]))
# to make further processing easier we can drop unncessary columns
df.drop(df.columns[[1,2,3,4,5,6]], axis=1, inplace=True)
df.head()
df.shape
# Check for class imbalance
import seaborn as sns
import matplotlib.pyplot as plt
sns.countplot(df.Label)
plt.xlabel('Categories with 0 = non causal and 1 = causal');
class_names = ['not causal', 'causal']
ax = sns.countplot(df.Label)
plt.xlabel('Causal categories')
ax.set_xticklabels(class_names);
# Class count
count_class_non_causal, count_class_causal = df.Label.value_counts()
# Divide by class
df_class_non_causal = df[df['Label'] == 0]
df_class_causal = df[df['Label'] == 1]
# undersample "non causal" class
df_class_non_causal_under = df_class_non_causal.sample(count_class_causal)
df_undersampled = pd.concat([df_class_non_causal_under, df_class_causal], axis=0)
print('Random under-sampling:')
print(df_undersampled.Label.value_counts())
df_undersampled.Label.value_counts().plot(kind='bar', title='Count (target)');
df = df_undersampled
# add POS Tags to sentences
import en_core_web_sm
if use_pos_tags == True:
#nlp = spacy.load('en_core_web_sm')
nlp = en_core_web_sm.load()
for index, row in df.iterrows():
doc = nlp(row['Sentence'])
new_sentence = ""
for token in doc:
new_sentence += token.text + "_" + token.dep_ + " "
df.at[index,'Sentence'] = new_sentence
PRE_TRAINED_MODEL_NAME = 'bert-base-cased'
tokenizer = BertTokenizer.from_pretrained(PRE_TRAINED_MODEL_NAME)
print(df.shape)
sample_txt = 'if_DET the_ADJ if sun_NOUN is_VERB shining, I will go to the beach characteristically._.'
encoding = tokenizer.encode_plus(
sample_txt,
max_length=62,
add_special_tokens=True, # Add '[CLS]' and '[SEP]'
return_token_type_ids=False,
pad_to_max_length=True,
return_attention_mask=True,
return_tensors='pt', # Return PyTorch tensors
truncation=True
)
print(tokenizer.unk_token_id)
print(tokenizer.tokenize(sample_txt))
encoding.keys()
encoding.items
```
### Choosing Sequence Length
```
token_lens = []
for txt in df.Sentence:
tokens = tokenizer.encode(txt, max_length=512, truncation=True)
token_lens.append(len(tokens))
sns.distplot(token_lens, kde=False)
plt.xlim([0, 512]);
plt.xlabel('Token count');
plt.ylabel('REQ count');
MAX_LEN = 384
class CausalDataset(Dataset):
def __init__(self, sentences, targets, tokenizer, max_len):
self.sentences = sentences
self.targets = targets
self.tokenizer = tokenizer
self.max_len = max_len
def __len__(self):
return len(self.sentences)
def __getitem__(self, item):
sentence = str(self.sentences[item])
target = self.targets[item]
encoding = self.tokenizer.encode_plus(
sentence,
add_special_tokens=True,
max_length=self.max_len,
return_token_type_ids=False,
pad_to_max_length=True,
return_attention_mask=True,
return_tensors='pt',
truncation=True
)
return {
'requirement_text': sentence,
'input_ids': encoding['input_ids'].flatten(),
'attention_mask': encoding['attention_mask'].flatten(),
'targets': torch.tensor(target, dtype=torch.long)
}
df_train, df_test = train_test_split(df, test_size=0.2, random_state=RANDOM_SEED)
df_val, df_test = train_test_split(df_test, test_size=0.5, random_state=RANDOM_SEED)
```
We also need to create a couple of data loaders. Here's a helper function to do it:
```
def create_data_loader(df, tokenizer, max_len, batch_size):
ds = CausalDataset(
sentences=df.Sentence.to_numpy(),
targets=df.Label.to_numpy(),
tokenizer=tokenizer,
max_len=max_len
)
return DataLoader(
ds,
batch_size=batch_size,
num_workers=4
)
BATCH_SIZE = 16
train_data_loader = create_data_loader(df_train, tokenizer, MAX_LEN, BATCH_SIZE)
val_data_loader = create_data_loader(df_val, tokenizer, MAX_LEN, BATCH_SIZE)
test_data_loader = create_data_loader(df_test, tokenizer, MAX_LEN, BATCH_SIZE)
```
## Causality Classification of Requirements
```
bert_model = BertModel.from_pretrained(PRE_TRAINED_MODEL_NAME)
class CausalClassifier(nn.Module):
def __init__(self, n_classes):
super(CausalClassifier, self).__init__()
self.bert = BertModel.from_pretrained(PRE_TRAINED_MODEL_NAME)
self.drop = nn.Dropout(p=0.3)
self.out = nn.Linear(self.bert.config.hidden_size, n_classes)
def forward(self, input_ids, attention_mask):
_, pooled_output = self.bert(
input_ids=input_ids,
attention_mask=attention_mask
)
output = self.drop(pooled_output)
return self.out(output)
model = CausalClassifier(len(class_names))
model = model.to(device)
```
### Training
```
EPOCHS = 20
LEARNING_RATE = 2e-5
optimizer = AdamW(model.parameters(), lr=LEARNING_RATE, correct_bias=False)
total_steps = len(train_data_loader) * EPOCHS
scheduler = get_linear_schedule_with_warmup(
optimizer,
num_warmup_steps=0,
num_training_steps=total_steps
)
loss_fn = nn.CrossEntropyLoss().to(device)
def train_epoch(
model,
data_loader,
loss_fn,
optimizer,
device,
scheduler,
n_examples
):
model = model.train()
losses = []
correct_predictions = 0
for d in data_loader:
input_ids = d["input_ids"].to(device)
attention_mask = d["attention_mask"].to(device)
targets = d["targets"].to(device)
outputs = model(
input_ids=input_ids,
attention_mask=attention_mask
)
_, preds = torch.max(outputs, dim=1)
loss = loss_fn(outputs, targets)
correct_predictions += torch.sum(preds == targets)
losses.append(loss.item())
loss.backward()
nn.utils.clip_grad_norm_(model.parameters(), max_norm=1.0)
optimizer.step()
scheduler.step()
optimizer.zero_grad()
return correct_predictions.double() / n_examples, np.mean(losses)
def eval_model(model, data_loader, loss_fn, device, n_examples):
model = model.eval()
losses = []
correct_predictions = 0
with torch.no_grad():
for d in data_loader:
input_ids = d["input_ids"].to(device)
attention_mask = d["attention_mask"].to(device)
targets = d["targets"].to(device)
outputs = model(
input_ids=input_ids,
attention_mask=attention_mask
)
_, preds = torch.max(outputs, dim=1)
loss = loss_fn(outputs, targets)
correct_predictions += torch.sum(preds == targets)
losses.append(loss.item())
return correct_predictions.double() / n_examples, np.mean(losses)
%%time
history = defaultdict(list)
best_accuracy = 0
for epoch in range(EPOCHS):
print(f'Epoch {epoch + 1}/{EPOCHS}')
print('-' * 10)
train_acc, train_loss = train_epoch(
model,
train_data_loader,
loss_fn,
optimizer,
device,
scheduler,
len(df_train)
)
print(f'Train loss {train_loss} accuracy {train_acc}')
val_acc, val_loss = eval_model(
model,
val_data_loader,
loss_fn,
device,
len(df_val)
)
print(f'Val loss {val_loss} accuracy {val_acc}')
print()
history['epoch'].append(epoch)
history['batch_size'].append(BATCH_SIZE)
history['learning_rate'].append(LEARNING_RATE)
history['train_acc'].append(train_acc.item())
history['train_loss'].append(train_loss)
history['val_acc'].append(val_acc.item())
history['val_loss'].append(val_loss)
if val_acc > best_accuracy:
torch.save(model.state_dict(), model_save_name)
best_accuracy = val_acc
pd.DataFrame(history).to_csv('training_results_' + currentTime + '.csv', mode='a')
```
## Evaluation
```
test_acc, _ = eval_model(
model,
test_data_loader,
loss_fn,
device,
len(df_test)
)
text_file = open("test_acc.txt", "a")
n = text_file.write('Test Accuracy of the best model (highest val accuracy): ' + str(test_acc.item()) + "\n")
text_file.close()
print(test_acc.item())
def get_predictions(model, data_loader):
model = model.eval()
requirement_texts = []
predictions = []
prediction_probs = []
real_values = []
with torch.no_grad():
for d in data_loader:
texts = d["requirement_text"]
input_ids = d["input_ids"].to(device)
attention_mask = d["attention_mask"].to(device)
targets = d["targets"].to(device)
outputs = model(
input_ids=input_ids,
attention_mask=attention_mask
)
_, preds = torch.max(outputs, dim=1)
probs = F.softmax(outputs, dim=1)
requirement_texts.extend(texts)
predictions.extend(preds)
prediction_probs.extend(probs)
real_values.extend(targets)
predictions = torch.stack(predictions).cpu()
prediction_probs = torch.stack(prediction_probs).cpu()
real_values = torch.stack(real_values).cpu()
return requirement_texts, predictions, prediction_probs, real_values
```
This is similar to the evaluation function, except that we're storing the text of the reviews and the predicted probabilities (by applying the softmax on the model outputs):
```
y_requirement_texts, y_pred, y_pred_probs, y_test = get_predictions(
model,
test_data_loader
)
len(y_requirement_texts)
report = classification_report(y_test, y_pred, target_names=class_names, output_dict=True)
df_report = pd.DataFrame(report).transpose()
df_report.to_csv("classification_report_BERT.csv")
print(classification_report(y_test, y_pred, target_names=class_names))
def show_confusion_matrix(confusion_matrix):
hmap = sns.heatmap(confusion_matrix, annot=True, fmt="d", cmap="Blues")
hmap.yaxis.set_ticklabels(hmap.yaxis.get_ticklabels(), rotation=0, ha='right')
hmap.xaxis.set_ticklabels(hmap.xaxis.get_ticklabels(), rotation=30, ha='right')
plt.ylabel('True causal category')
plt.xlabel('Predicted causal category');
cm = confusion_matrix(y_test, y_pred)
df_cm = pd.DataFrame(cm, index=class_names, columns=class_names)
show_confusion_matrix(df_cm)
df_cm.to_csv("confusion_matrix.csv")
idx = 2
sentence_text = y_requirement_texts[idx]
true_label = y_test[idx]
pred_df = pd.DataFrame({
'class_names': class_names,
'values': y_pred_probs[idx]
})
# calculate AUC Score on test set
from sklearn.metrics import roc_auc_score
roc_auc_score(y_test, y_pred)
text_file = open("test_acc.txt", "a")
n = text_file.write('AUC Score of the best model (highest val accuracy): ' + str(roc_auc_score(y_test, y_pred)) + "\n")
text_file.close()
print("\n".join(wrap(sentence_text)))
print()
print(f'True sentiment: {class_names[true_label]}')
sns.barplot(x='values', y='class_names', data=pred_df, orient='h')
plt.ylabel('sentiment')
plt.xlabel('probability')
plt.xlim([0, 1]);
# lets test the normal regex approach on the test data set
# we iterate over all sentences and mark a sentence as causal in case it contains one of the cue phrases
# we define a list of cue phrase which we want to check
cue_phrases = ['if ', 'when ', 'because ', 'since ', 'therefore ', 'hence ', 'given ', 'where ',
'whose ', 'in order to ', 'in the case of ', 'due to ', 'needed ', 'require ', 'required ' 'during ',
'in case of ', 'while ', 'thus ', 'as ', 'except ', 'forced by ', 'only for ', 'within ', 'after ', 'whenever ', 'which ', 'before ',
'allows ', 'allow ', 'unless ', 'prior to', 'as long as ', 'depending on ', 'depends on ', 'result in ', 'increases ',
'lead to ', 'thereby ', 'cause ', 'in the event ', 'once ', 'in such cases ', 'throughout ', 'improve ', 'to that end ', 'to this end']
# sentences to lower case
y_requirement_texts = [entry.lower() for entry in y_requirement_texts]
y_pred_regex = pd.DataFrame(columns=['Sentence', 'Label'])
for row in y_requirement_texts:
if any(substring in row for substring in cue_phrases):
new_row = {'Sentence': row, 'Label': 1}
y_pred_regex = y_pred_regex.append(new_row, ignore_index=True)
else:
new_row = {'Sentence': row, 'Label': 0}
y_pred_regex = y_pred_regex.append(new_row, ignore_index=True)
from sklearn.preprocessing import LabelEncoder
Encoder = LabelEncoder()
y_pred_regex = Encoder.fit_transform(y_pred_regex['Label'])
report_regex = classification_report(y_test, y_pred_regex, target_names=class_names, output_dict=True)
df_report_regex = pd.DataFrame(report_regex).transpose()
df_report_regex.to_csv("classification_report_regex.csv")
print(classification_report(y_test, y_pred_regex))
```
| github_jupyter |
# Interactions from the literature
```
%pylab inline
%config InlineBackend.figure_format = 'retina'
import json
import numpy as np
studies = [ { 'name' : 'Gopher, Lice',
'type' : 'parasitism',
'host' : 'data/gopher-louse/gopher.tree',
'guest': 'data/gopher-louse/lice.tree',
'links': 'data/gopher-louse/links.csv' },
{ 'name' : 'Sedge, Smut',
'type' : 'parasitism',
'host' : 'data/sedge-smut/host.tree',
'guest': 'data/sedge-smut/guest.tree',
'links': 'data/sedge-smut/links.csv' },
{ 'name' : 'Fish, Worm',
'type' : 'parasitism',
'host' : 'data/fish-worm/host.tree',
'guest': 'data/fish-worm/guest.tree',
'links': 'data/fish-worm/links.csv' },
{ 'name' : 'beeh',
'type' : 'frugivory',
'host' : 'data/plant-pollinators/beeh/plant.tree',
'guest': 'data/plant-pollinators/beeh/animal.tree',
'links': 'data/plant-pollinators/beeh/beeh_links.csv' },
{ 'name' : 'arr1',
'type' : 'pollination',
'host' : 'data/plant-pollinators/arr1/plant.tree',
'guest': 'data/plant-pollinators/arr1/animal.tree',
'links': 'data/plant-pollinators/arr1/arr1_links.csv' },
{ 'name' : 'arr2',
'type' : 'pollination',
'host' : 'data/plant-pollinators/arr2/plant.tree',
'guest': 'data/plant-pollinators/arr2/animal.tree',
'links': 'data/plant-pollinators/arr2/arr2_links.csv' },
{ 'name' : 'arr3',
'type' : 'pollination',
'host' : 'data/plant-pollinators/arr3/plant.tree',
'guest': 'data/plant-pollinators/arr3/animal.tree',
'links': 'data/plant-pollinators/arr3/arr3_links.csv' },
{ 'name' : 'bair',
'type' : 'frugivory',
'host' : 'data/plant-pollinators/bair/plant.tree',
'guest': 'data/plant-pollinators/bair/animal.tree',
'links': 'data/plant-pollinators/bair/bair_links.csv' },
{ 'name' : 'cacg',
'type' : 'frugivory',
'host' : 'data/plant-pollinators/cacg/plant.tree',
'guest': 'data/plant-pollinators/cacg/animal.tree',
'links': 'data/plant-pollinators/cacg/cacg_links.csv' },
{ 'name' : 'caco',
'type' : 'frugivory',
'host' : 'data/plant-pollinators/caco/plant.tree',
'guest': 'data/plant-pollinators/caco/animal.tree',
'links': 'data/plant-pollinators/caco/caco_links.csv' },
{ 'name' : 'caci',
'type' : 'frugivory',
'host' : 'data/plant-pollinators/caci/plant.tree',
'guest': 'data/plant-pollinators/caci/animal.tree',
'links': 'data/plant-pollinators/caci/caci_links.csv' },
{ 'name' : 'cafr',
'type' : 'frugivory',
'host' : 'data/plant-pollinators/cafr/plant.tree',
'guest': 'data/plant-pollinators/cafr/animal.tree',
'links': 'data/plant-pollinators/cafr/cafr_links.csv' },
{ 'name' : 'cllo',
'type' : 'pollination',
'host' : 'data/plant-pollinators/cllo/plant.tree',
'guest': 'data/plant-pollinators/cllo/animal.tree',
'links': 'data/plant-pollinators/cllo/cllo_links.csv' },
{ 'name' : 'crom',
'type' : 'frugivory',
'host' : 'data/plant-pollinators/crom/plant.tree',
'guest': 'data/plant-pollinators/crom/animal.tree',
'links': 'data/plant-pollinators/crom/crom_links.csv' },
{ 'name' : 'dihi',
'type' : 'pollination',
'host' : 'data/plant-pollinators/dihi/plant.tree',
'guest': 'data/plant-pollinators/dihi/animal.tree',
'links': 'data/plant-pollinators/dihi/dihi_links.csv' },
{ 'name' : 'dish',
'type' : 'pollination',
'host' : 'data/plant-pollinators/dish/plant.tree',
'guest': 'data/plant-pollinators/dish/animal.tree',
'links': 'data/plant-pollinators/dish/dish_links.csv' },
{ 'name' : 'dupo',
'type' : 'pollination',
'host' : 'data/plant-pollinators/dupo/plant.tree',
'guest': 'data/plant-pollinators/dupo/animal.tree',
'links': 'data/plant-pollinators/dupo/dupo_links.csv' },
{ 'name' : 'eol',
'type' : 'pollination',
'host' : 'data/plant-pollinators/eol/plant.tree',
'guest': 'data/plant-pollinators/eol/animal.tree',
'links': 'data/plant-pollinators/eol/eol_links.csv' },
{ 'name' : 'eolz',
'type' : 'pollination',
'host' : 'data/plant-pollinators/eolz/plant.tree',
'guest': 'data/plant-pollinators/eolz/animal.tree',
'links': 'data/plant-pollinators/eolz/eolz_links.csv' },
{ 'name' : 'eski',
'type' : 'pollination',
'host' : 'data/plant-pollinators/eski/plant.tree',
'guest': 'data/plant-pollinators/eski/animal.tree',
'links': 'data/plant-pollinators/eski/eski_links.csv' },
{ 'name' : 'fros',
'type' : 'frugivory',
'host' : 'data/plant-pollinators/fros/plant.tree',
'guest': 'data/plant-pollinators/fros/animal.tree',
'links': 'data/plant-pollinators/fros/fros_links.csv' },
{ 'name' : 'gen1',
'type' : 'frugivory',
'host' : 'data/plant-pollinators/gen1/plant.tree',
'guest': 'data/plant-pollinators/gen1/animal.tree',
'links': 'data/plant-pollinators/gen1/gen1_links.csv' },
{ 'name' : 'gen2',
'type' : 'frugivory',
'host' : 'data/plant-pollinators/gen2/plant.tree',
'guest': 'data/plant-pollinators/gen2/animal.tree',
'links': 'data/plant-pollinators/gen2/gen2_links.csv' },
{ 'name' : 'hamm',
'type' : 'frugivory',
'host' : 'data/plant-pollinators/hamm/plant.tree',
'guest': 'data/plant-pollinators/hamm/animal.tree',
'links': 'data/plant-pollinators/hamm/hamm_links.csv' },
{ 'name' : 'herr',
'type' : 'pollination',
'host' : 'data/plant-pollinators/herr/plant.tree',
'guest': 'data/plant-pollinators/herr/animal.tree',
'links': 'data/plant-pollinators/herr/herr_links.csv' },
{ 'name' : 'hock',
'type' : 'pollination',
'host' : 'data/plant-pollinators/hock/plant.tree',
'guest': 'data/plant-pollinators/hock/animal.tree',
'links': 'data/plant-pollinators/hock/hock_links.csv' },
{ 'name' : 'hrat',
'type' : 'frugivory',
'host' : 'data/plant-pollinators/hrat/plant.tree',
'guest': 'data/plant-pollinators/hrat/animal.tree',
'links': 'data/plant-pollinators/hrat/hrat_links.csv' },
{ 'name' : 'inpk',
'type' : 'pollination',
'host' : 'data/plant-pollinators/inpk/plant.tree',
'guest': 'data/plant-pollinators/inpk/animal.tree',
'links': 'data/plant-pollinators/inpk/inpk_links.csv' },
{ 'name' : 'kant',
'type' : 'frugivory',
'host' : 'data/plant-pollinators/kant/plant.tree',
'guest': 'data/plant-pollinators/kant/animal.tree',
'links': 'data/plant-pollinators/kant/kant_links.csv' },
{ 'name' : 'kevn',
'type' : 'pollination',
'host' : 'data/plant-pollinators/kevn/plant.tree',
'guest': 'data/plant-pollinators/kevn/animal.tree',
'links': 'data/plant-pollinators/kevn/kevn_links.csv' },
{ 'name' : 'lope',
'type' : 'frugivory',
'host' : 'data/plant-pollinators/lope/plant.tree',
'guest': 'data/plant-pollinators/lope/animal.tree',
'links': 'data/plant-pollinators/lope/lope_links.csv' },
{ 'name' : 'mack',
'type' : 'frugivory',
'host' : 'data/plant-pollinators/mack/plant.tree',
'guest': 'data/plant-pollinators/mack/animal.tree',
'links': 'data/plant-pollinators/mack/mack_links.csv' },
{ 'name' : 'med1',
'type' : 'pollination',
'host' : 'data/plant-pollinators/med1/plant.tree',
'guest': 'data/plant-pollinators/med1/animal.tree',
'links': 'data/plant-pollinators/med1/med1_links.csv' },
{ 'name' : 'med2',
'type' : 'pollination',
'host' : 'data/plant-pollinators/med2/plant.tree',
'guest': 'data/plant-pollinators/med2/animal.tree',
'links': 'data/plant-pollinators/med2/med2_links.csv' },
{ 'name' : 'memm',
'type' : 'pollination',
'host' : 'data/plant-pollinators/med2/plant.tree',
'guest': 'data/plant-pollinators/med2/animal.tree',
'links': 'data/plant-pollinators/med2/med2_links.csv' },
{ 'name' : 'moma',
'type' : 'pollination',
'host' : 'data/plant-pollinators/moma/plant.tree',
'guest': 'data/plant-pollinators/moma/animal.tree',
'links': 'data/plant-pollinators/moma/moma_links.csv' },
{ 'name' : 'mont',
'type' : 'frugivory',
'host' : 'data/plant-pollinators/mont/plant.tree',
'guest': 'data/plant-pollinators/mont/animal.tree',
'links': 'data/plant-pollinators/mont/mont_links.csv' },
{ 'name' : 'mott',
'type' : 'pollination',
'host' : 'data/plant-pollinators/mott/plant.tree',
'guest': 'data/plant-pollinators/mott/animal.tree',
'links': 'data/plant-pollinators/mott/mott_links.csv' },
{ 'name' : 'mull',
'type' : 'pollination',
'host' : 'data/plant-pollinators/mull/plant.tree',
'guest': 'data/plant-pollinators/mull/animal.tree',
'links': 'data/plant-pollinators/mull/mull_links.csv' },
{ 'name' : 'ncor',
'type' : 'frugivory',
'host' : 'data/plant-pollinators/ncor/plant.tree',
'guest': 'data/plant-pollinators/ncor/animal.tree',
'links': 'data/plant-pollinators/ncor/ncor_links.csv' },
{ 'name' : 'nnog',
'type' : 'frugivory',
'host' : 'data/plant-pollinators/nnog/plant.tree',
'guest': 'data/plant-pollinators/nnog/animal.tree',
'links': 'data/plant-pollinators/nnog/nnog_links.csv' },
{ 'name' : 'olau',
'type' : 'pollination',
'host' : 'data/plant-pollinators/olau/plant.tree',
'guest': 'data/plant-pollinators/olau/animal.tree',
'links': 'data/plant-pollinators/olau/olau_links.csv' },
{ 'name' : 'perc',
'type' : 'pollination',
'host' : 'data/plant-pollinators/perc/plant.tree',
'guest': 'data/plant-pollinators/perc/animal.tree',
'links': 'data/plant-pollinators/perc/perc_links.csv' },
{ 'name' : 'rabr',
'type' : 'pollination',
'host' : 'data/plant-pollinators/rabr/plant.tree',
'guest': 'data/plant-pollinators/rabr/animal.tree',
'links': 'data/plant-pollinators/rabr/rabr_links.csv' },
{ 'name' : 'rmrz',
'type' : 'pollination',
'host' : 'data/plant-pollinators/rmrz/plant.tree',
'guest': 'data/plant-pollinators/rmrz/animal.tree',
'links': 'data/plant-pollinators/rmrz/rmrz_links.csv' },
{ 'name' : 'sapf',
'type' : 'frugivory',
'host' : 'data/plant-pollinators/sapf/plant.tree',
'guest': 'data/plant-pollinators/sapf/animal.tree',
'links': 'data/plant-pollinators/sapf/sapf_links.csv' },
{ 'name' : 'schm',
'type' : 'pollination',
'host' : 'data/plant-pollinators/schm/plant.tree',
'guest': 'data/plant-pollinators/schm/animal.tree',
'links': 'data/plant-pollinators/schm/schm_links.csv' },
{ 'name' : 'smal',
'type' : 'pollination',
'host' : 'data/plant-pollinators/smal/plant.tree',
'guest': 'data/plant-pollinators/smal/animal.tree',
'links': 'data/plant-pollinators/smal/smal_links.csv' },
{ 'name' : 'snow',
'type' : 'frugivory',
'host' : 'data/plant-pollinators/snow/plant.tree',
'guest': 'data/plant-pollinators/snow/animal.tree',
'links': 'data/plant-pollinators/snow/snow_links.csv' },
{ 'name' : 'wes',
'type' : 'frugivory',
'host' : 'data/plant-pollinators/wes/plant.tree',
'guest': 'data/plant-pollinators/wes/animal.tree',
'links': 'data/plant-pollinators/wes/wes_links.csv' },
{ 'name' : 'wyth',
'type' : 'frugivory',
'host' : 'data/plant-pollinators/wyth/plant.tree',
'guest': 'data/plant-pollinators/wyth/animal.tree',
'links': 'data/plant-pollinators/wyth/wyth_links.csv' },
#{ 'name' : 'lopez',
# 'type' : 'pollination',
# 'host' : 'data/plant-pollinators/lopez-fig-wasp/host.tree',
# 'guest': 'data/plant-pollinators/lopez-fig-wasp/guest.tree',
# 'links': 'data/plant-pollinators/lopez-fig-wasp/lopez-fig-wasp_links.csv' },
]
null_studies = []
for i in xrange(50) :
name = 'null' + str(i)
path = 'data/simulated/null/' + name
s = { 'name' : name,
'type' : 'null',
'host' : path + '/host.tree',
'guest' : path + '/guest.tree',
'links' : path + '/links.csv' }
null_studies.append(s)
perfect_studies = []
for i in xrange(50) :
name = 'perfect' + str(i)
path = 'data/simulated/perfect/' + name
s = { 'name' : name,
'type' : 'perfect',
'host' : path + '/host.tree',
'guest' : path + '/guest.tree',
'links' : path + '/links.csv' }
perfect_studies.append(s)
with open( 'studies.json', 'w' ) as f :
json.dump( studies + null_studies + perfect_studies, f )
```
## Simulated interactions
This notebook generates simulated datasets of phylogenies of interacting
groups of species.
There are two kinds of simulated data : Null data sets, consisting of two randomly generated
trees with random links between leafs, and "perfect" data sets, consisting of identical trees
with leaves linked in a bijection.
```
## Null datasets
import pandas as pd
import dendropy
import os
N_min, N_max = 10, 200
for i in xrange(50) :
N = np.random.randint( N_min, N_max )
M = np.random.randint( N_min, N_max )
L = np.random.randint( 0.5*( N+M ), 4.0*( N+M ) )
host_taxa = [ 'host_' + str(x) for x in xrange(N) ]
guest_taxa = [ 'guest_' + str(x) for x in xrange(M) ]
ht = dendropy.simulate.treesim.birth_death_tree( 1.0, 0.5,
#birth_rate_sd=0.5, death_rate_sd=0.5,
ntax=len(host_taxa) )
gt = dendropy.simulate.treesim.birth_death_tree( 1.0, 0.5,
#birth_rate_sd=0.5, death_rate_sd=0.5,
ntax=len(guest_taxa) )
for leaf,name in zip(ht.leaf_node_iter(),host_taxa) :
leaf.taxon.label = name
for leaf,name in zip(gt.leaf_node_iter(),guest_taxa) :
leaf.taxon.label = name
lm = np.zeros( (N,M) )
for j in xrange( L ) :
lm[ np.random.randint(N), np.random.randint(M) ] = 1
lmdf = pd.DataFrame( lm, columns=guest_taxa, index=host_taxa, dtype=int )
path = 'simulated/null/null' + str(i) +'/'
if not os.path.exists( path ) :
os.mkdir( path )
ht.write( file=open( path + 'host.tree', 'w'), schema='newick' )
gt.write( file=open( path + 'guest.tree', 'w'), schema='newick' )
lmdf.to_csv( path + 'links.csv' )
## "Perfect" datasets
import numpy as np
import pandas as pd
import dendropy
import os
N_min, N_max = 10, 200
for i in xrange(50) :
N = np.random.randint( N_min, N_max )
path = 'simulated/perfect/perfect' + str(i) +'/'
if not os.path.exists( path ) :
os.mkdir( path )
host_taxa = [ 'host_' + str(x) for x in xrange(N) ]
guest_taxa = [ 'guest_' + str(x) for x in xrange(N) ]
t = dendropy.simulate.treesim.birth_death_tree( 1.0, 0.5,
#birth_rate_sd=0.5, death_rate_sd=0.5,
ntax=len(host_taxa) )
for leaf,name in zip(t.leaf_node_iter(),host_taxa) :
leaf.taxon.label = name
t.write( file=open( path + 'host.tree', 'w'), schema='newick' )
for leaf,name in zip(t.leaf_node_iter(),guest_taxa) :
leaf.taxon.label = name
t.write( file=open( path + 'guest.tree', 'w'), schema='newick' )
lm = np.zeros( (N,N) )
for j in xrange(N) :
lm[ j, j ] = 1
lmdf = pd.DataFrame( lm, columns=guest_taxa, index=host_taxa, dtype=int )
lmdf.to_csv( path + 'links.csv' )
```
## Data integrity
Most of the time, datasets from the literature are broken in some way.
Here are some useful routines for checking their integrity and fixing errors.
```
from SuchTree import SuchTree, SuchLinkedTrees
import pandas as pd
import seaborn
name = 'snow'
T1 = SuchTree( 'plant-pollinators/' + name + '/plant.tree' )
T2 = SuchTree( 'plant-pollinators/' + name + '/animal.tree' )
links = pd.DataFrame.from_csv( 'plant-pollinators/' + name + '/' + name + '_links.csv' )
links = links.loc[ T1.leafs.keys() ]
links = links[ T2.leafs.keys() ]
SLT = SuchLinkedTrees( T1, T2, links )
links = pd.DataFrame.from_csv( 'plant-pollinators/' + name + '/' + name + '_links.csv' )
set( T2.leafs.keys() ) ^ set(links.columns)
#set( T2.leafs.keys() )
# flip link matrix
links.T.to_csv( 'plant-pollinators/' + name + '/' + name + '_links.csv' )
# underscores for spaces
links.index = [ s.replace(' ','_') for s in links.index ]
links.columns = [ s.replace(' ','_') for s in links.columns ]
links.to_csv( 'plant-pollinators/' + name + '/' + name + '_links.csv' )
seaborn.heatmap( SLT.adjacency() )
#seaborn.kdeplot( SLT.spectrum() )
```
| github_jupyter |
# Rigid-body transformations in three-dimensions
> Marcos Duarte
> Laboratory of Biomechanics and Motor Control ([http://demotu.org/](http://demotu.org/))
> Federal University of ABC, Brazil
The kinematics of a rigid body is completely described by its pose, i.e., its position and orientation in space (and the corresponding changes, translation and rotation). In a three-dimensional space, at least three coordinates and three angles are necessary to describe the pose of the rigid body, totalizing six degrees of freedom for a rigid body.
In motion analysis, to describe a translation and rotation of a rigid body with respect to a coordinate system, typically we attach another coordinate system to the rigid body and determine a transformation between these two coordinate systems.
A transformation is any function mapping a set to another set. For the description of the kinematics of rigid bodies, we are interested only in what is called rigid or Euclidean transformations (denoted as SE(3) for the three-dimensional space) because they preserve the distance between every pair of points of the body (which is considered rigid by definition). Translations and rotations are examples of rigid transformations (a reflection is also an example of rigid transformation but this changes the right-hand axis convention to a left hand, which usually is not of interest). In turn, rigid transformations are examples of [affine transformations](https://en.wikipedia.org/wiki/Affine_transformation). Examples of other affine transformations are shear and scaling transformations (which preserves angles but not lengths).
We will follow the same rationale as in the notebook [Rigid-body transformations in a plane (2D)](http://nbviewer.ipython.org/github/BMClab/bmc/blob/master/notebooks/Transformation2D.ipynb) and we will skip the fundamental concepts already covered there. So, you if haven't done yet, you should read that notebook before continuing here.
## Translation
A pure three-dimensional translation of a rigid body (or a coordinate system attached to it) in relation to other rigid body (with other coordinate system) is illustrated in the figure below.
<br>
<figure><img src='./../images/translation3D.png' alt='translation 3D'/> <figcaption><center><i>Figure. A point in three-dimensional space represented in two coordinate systems, with one coordinate system translated.</i></center></figcaption> </figure>
The position of point $\mathbf{P}$ originally described in the $xyz$ (local) coordinate system but now described in the $\mathbf{XYZ}$ (Global) coordinate system in vector form is:
$$ \mathbf{P_G} = \mathbf{L_G} + \mathbf{P_l} $$
Or in terms of its components:
$$ \begin{array}{}
\mathbf{P_X} =& \mathbf{L_X} + \mathbf{P}_x \\
\mathbf{P_Y} =& \mathbf{L_Y} + \mathbf{P}_y \\
\mathbf{P_Z} =& \mathbf{L_Z} + \mathbf{P}_z
\end{array} $$
And in matrix form:
$$
\begin{bmatrix}
\mathbf{P_X} \\
\mathbf{P_Y} \\
\mathbf{P_Z}
\end{bmatrix} =
\begin{bmatrix}
\mathbf{L_X} \\
\mathbf{L_Y} \\
\mathbf{L_Z}
\end{bmatrix} +
\begin{bmatrix}
\mathbf{P}_x \\
\mathbf{P}_y \\
\mathbf{P}_z
\end{bmatrix}
$$
From classical mechanics, this is an example of [Galilean transformation](http://en.wikipedia.org/wiki/Galilean_transformation).
Let's use Python to compute some numeric examples:
```
# Import the necessary libraries
import numpy as np
# suppress scientific notation for small numbers:
np.set_printoptions(precision=4, suppress=True)
```
For example, if the local coordinate system is translated by $\mathbf{L_G}=[1, 2, 3]$ in relation to the Global coordinate system, a point with coordinates $\mathbf{P_l}=[4, 5, 6]$ at the local coordinate system will have the position $\mathbf{P_G}=[5, 7, 9]$ at the Global coordinate system:
```
LG = np.array([1, 2, 3]) # Numpy array
Pl = np.array([4, 5, 6])
PG = LG + Pl
PG
```
This operation also works if we have more than one point (NumPy try to guess how to handle vectors with different dimensions):
```
Pl = np.array([[1, 2, 3], [4, 5, 6], [7, 8, 9]]) # 2D array with 3 rows and 2 columns
PG = LG + Pl
PG
```
## Rotation
A pure three-dimensional rotation of a $xyz$ (local) coordinate system in relation to other $\mathbf{XYZ}$ (Global) coordinate system and the position of a point in these two coordinate systems are illustrated in the next figure (remember that this is equivalent to describing a rotation between two rigid bodies).
<br>
<figure><img src='./../images/rotation3D.png' alt='rotation 3D'/> <figcaption><center><i>A point in three-dimensional space represented in two coordinate systems, with one system rotated.</i></center></figcaption> </figure>
In analogy to the rotation in two dimensions, we can calculate the rotation matrix that describes the rotation of the $xyz$ (local) coordinate system in relation to the $\mathbf{XYZ}$ (Global) coordinate system using the direction cosines between the axes of the two coordinate systems:
$$ \mathbf{R_{Gl}} = \begin{bmatrix}
\cos\mathbf{X}x & \cos\mathbf{X}y & \cos\mathbf{X}z \\
\cos\mathbf{Y}x & \cos\mathbf{Y}y & \cos\mathbf{Y}z \\
\cos\mathbf{Z}x & \cos\mathbf{Z}y & \cos\mathbf{Z}z
\end{bmatrix} $$
Note however that for rotations around more than one axis, these angles will not lie in the main planes ($\mathbf{XY, YZ, ZX}$) of the $\mathbf{XYZ}$ coordinate system, as illustrated in the figure below for the direction angles of the $y$ axis only. Thus, the determination of these angles by simple inspection, as we have done for the two-dimensional case, would not be simple.
<br>
<figure>
<img src='./../images/directioncosine3D.png' width=260 alt='direction angles 3D'/> <figcaption><center><i>Figure. Definition of direction angles for the $y$ axis of the local coordinate system in relation to the $\mathbf{XYZ}$ Global coordinate system.</i></center></figcaption>
</figure>
Note that the nine angles shown in the matrix above for the direction cosines are obviously redundant since only three angles are necessary to describe the orientation of a rigid body in the three-dimensional space.
An important characteristic of angles in the three-dimensional space is that angles cannot be treated as vectors: the result of a sequence of rotations of a rigid body around different axes depends on the order of the rotations, as illustrated in the next figure.
<br>
<figure>
<img src='./../images/rotationsseqs2.png' alt='rotations'/><figcaption><i>Figure. The result of a sequence of rotations around different axes of a coordinate system depends on the order of the rotations. In the first example (first row), the rotations are around a Global (fixed) coordinate system. In the second example (second row), the rotations are around a local (rotating) coordinate system.</i></figcaption>
</figure>
Let's focus now on how to understand rotations in the three-dimensional space, looking at the rotations between coordinate systems (or between rigid bodies). Later we will apply what we have learned to describe the position of a point in these different coordinate systems.
### Euler angles
There are different ways to describe a three-dimensional rotation of a rigid body (or of a coordinate system). The most straightforward solution would probably be to use a [spherical coordinate system](http://nbviewer.ipython.org/github/demotu/BMC/blob/master/notebooks/ReferenceFrame.ipynb#Spherical-coordinate-system), but spherical coordinates would be difficult to give an anatomical or clinical interpretation. A solution that has been often employed in biomechanics to handle rotations in the three-dimensional space is to use Euler angles. Under certain conditions, Euler angles can have an anatomical interpretation, but this representation also has some caveats. Let's see the Euler angles now.
[Leonhard Euler](https://en.wikipedia.org/wiki/Leonhard_Euler) in the XVIII century showed that two three-dimensional coordinate systems with a common origin can be related by a sequence of up to three elemental rotations about the axes of the local coordinate system, where no two successive rotations may be about the same axis, which now are known as [Euler (or Eulerian) angles](http://en.wikipedia.org/wiki/Euler_angles).
#### Elemental rotations
First, let's see rotations around a fixed Global coordinate system as we did for the two-dimensional case. The next figure illustrates elemental rotations of the local coordinate system around each axis of the fixed Global coordinate system.
<br>
<figure>
<img src='./../images/rotations.png' alt='rotations'/> <figcaption><center><i>Figure. Elemental rotations of the $xyz$ coordinate system around each axis, $\mathbf{X}$, $\mathbf{Y}$, and $\mathbf{Z}$, of the fixed $\mathbf{XYZ}$ coordinate system. Note that for better clarity, the axis around where the rotation occurs is shown perpendicular to this page for each elemental rotation.</i></center></figcaption>
</figure>
#### Rotations around the fixed coordinate system
The rotation matrices for the elemental rotations around each axis of the fixed $\mathbf{XYZ}$ coordinate system (rotations of the local coordinate system in relation to the Global coordinate system) are shown next.
Around $\mathbf{X}$ axis:
<span class="notranslate">
$$ \mathbf{R_{Gl,\,X}} =
\begin{bmatrix}
1 & 0 & 0 \\
0 & \cos\alpha & -\sin\alpha \\
0 & \sin\alpha & \cos\alpha
\end{bmatrix} $$
</span>
Around $\mathbf{Y}$ axis:
$$ \mathbf{R_{Gl,\,Y}} =
\begin{bmatrix}
\cos\beta & 0 & \sin\beta \\
0 & 1 & 0 \\
-\sin\beta & 0 & \cos\beta
\end{bmatrix} $$
Around $\mathbf{Z}$ axis:
$$ \mathbf{R_{Gl,\,Z}} =
\begin{bmatrix}
\cos\gamma & -\sin\gamma & 0\\
\sin\gamma & \cos\gamma & 0 \\
0 & 0 & 1
\end{bmatrix} $$
These matrices are the rotation matrices for the case of two-dimensional coordinate systems plus the corresponding terms for the third axes of the local and Global coordinate systems, which are parallel.
To understand why the terms for the third axes are 1's or 0's, for instance, remember they represent the cosine directors. The cosines between $\mathbf{X}x$, $\mathbf{Y}y$, and $\mathbf{Z}z$ for the elemental rotations around respectively the $\mathbf{X}$, $\mathbf{Y}$, and $\mathbf{Z}$ axes are all 1 because $\mathbf{X}x$, $\mathbf{Y}y$, and $\mathbf{Z}z$ are parallel ($\cos 0^o$). The cosines of the other elements are zero because the axis around where each rotation occurs is perpendicular to the other axes of the coordinate systems ($\cos 90^o$).
#### Rotations around the local coordinate system
The rotation matrices for the elemental rotations this time around each axis of the $xyz$ coordinate system (rotations of the Global coordinate system in relation to the local coordinate system), similarly to the two-dimensional case, are simply the transpose of the above matrices as shown next.
Around $x$ axis:
$$ \mathbf{R}_{\mathbf{lG},\,x} =
\begin{bmatrix}
1 & 0 & 0 \\
0 & \cos\alpha & \sin\alpha \\
0 & -\sin\alpha & \cos\alpha
\end{bmatrix} $$
Around $y$ axis:
$$ \mathbf{R}_{\mathbf{lG},\,y} =
\begin{bmatrix}
\cos\beta & 0 & -\sin\beta \\
0 & 1 & 0 \\
\sin\beta & 0 & \cos\beta
\end{bmatrix} $$
Around $z$ axis:
$$ \mathbf{R}_{\mathbf{lG},\,z} =
\begin{bmatrix}
\cos\gamma & \sin\gamma & 0\\
-\sin\gamma & \cos\gamma & 0 \\
0 & 0 & 1
\end{bmatrix} $$
Notice this is equivalent to instead of rotating the local coordinate system by $\alpha, \beta, \gamma$ in relation to axes of the Global coordinate system, to rotate the Global coordinate system by $-\alpha, -\beta, -\gamma$ in relation to the axes of the local coordinate system; remember that $\cos(-\:\cdot)=\cos(\cdot)$ and $\sin(-\:\cdot)=-\sin(\cdot)$.
The fact that we chose to rotate the local coordinate system by a counterclockwise (positive) angle in relation to the Global coordinate system is just a matter of convention.
#### Sequence of elemental rotations
Consider now a sequence of elemental rotations around the $\mathbf{X}$, $\mathbf{Y}$, and $\mathbf{Z}$ axes of the fixed $\mathbf{XYZ}$ coordinate system illustrated in the next figure.
<br>
<figure><img src='./../images/rotations_XYZ.png' alt='rotations'/> <figcaption><center><i>Figure. Sequence of elemental rotations of the $xyz$ coordinate system around each axis, $\mathbf{X}$, $\mathbf{Y}$, $\mathbf{Z}$, of the fixed $\mathbf{XYZ}$ coordinate system.</i></center></figcaption> </figure>
This sequence of elemental rotations (each one of the local coordinate system with respect to the fixed Global coordinate system) is mathematically represented by a multiplication between the rotation matrices:
$$ \begin{array}{l l}
\mathbf{R_{Gl,\;XYZ}} & = \mathbf{R_{Z}} \mathbf{R_{Y}} \mathbf{R_{X}} \\
\\
& = \begin{bmatrix}
\cos\gamma & -\sin\gamma & 0\\
\sin\gamma & \cos\gamma & 0 \\
0 & 0 & 1
\end{bmatrix}
\begin{bmatrix}
\cos\beta & 0 & \sin\beta \\
0 & 1 & 0 \\
-\sin\beta & 0 & \cos\beta
\end{bmatrix}
\begin{bmatrix}
1 & 0 & 0 \\
0 & \cos\alpha & -sin\alpha \\
0 & \sin\alpha & cos\alpha
\end{bmatrix} \\
\\
& =
\begin{bmatrix}
\cos\beta\:\cos\gamma \;&\;
\sin\alpha\:\sin\beta\:cos\gamma-\cos\alpha\:\sin\gamma \;&\;
\cos\alpha\:\sin\beta\:cos\gamma+\sin\alpha\:\sin\gamma \;\;\; \\
\cos\beta\:\sin\gamma \;&\;
\sin\alpha\:\sin\beta\:sin\gamma+\cos\alpha\:\cos\gamma \;&\;
\cos\alpha\:\sin\beta\:sin\gamma-\sin\alpha\:\cos\gamma \;\;\; \\
-\sin\beta \;&\; \sin\alpha\:\cos\beta \;&\; \cos\alpha\:\cos\beta \;\;\;
\end{bmatrix}
\end{array} $$
Note that the order of the matrices.
We can check this matrix multiplication using [Sympy](http://sympy.org/en/index.html):
```
#import the necessary libraries
from IPython.core.display import Math, display
import sympy as sym
cos, sin = sym.cos, sym.sin
a, b, g = sym.symbols('alpha, beta, gamma')
# Elemental rotation matrices of xyz in relation to XYZ:
RX = sym.Matrix([[1, 0, 0], [0, cos(a), -sin(a)], [0, sin(a), cos(a)]])
RY = sym.Matrix([[cos(b), 0, sin(b)], [0, 1, 0], [-sin(b), 0, cos(b)]])
RZ = sym.Matrix([[cos(g), -sin(g), 0], [sin(g), cos(g), 0], [0, 0, 1]])
# Rotation matrix of xyz in relation to XYZ:
RXYZ = RZ*RY*RX
display(Math(sym.latex(r'\mathbf{R_{Gl,\,XYZ}}=') + sym.latex(RXYZ, mat_str='matrix')))
```
For instance, we can calculate the numerical rotation matrix for these sequential elemental rotations by $90^o$ around $\mathbf{X,Y,Z}$:
```
R = sym.lambdify((a, b, g), RXYZ, 'numpy')
R = R(np.pi/2, np.pi/2, np.pi/2)
display(Math(r'\mathbf{R_{Gl,\,XYZ\,}}(90^o, 90^o, 90^o) =' + \
sym.latex(sym.Matrix(R).n(chop=True, prec=3))))
```
Examining the matrix above and the correspondent previous figure, one can see they agree: the rotated $x$ axis (first column of the above matrix) has value -1 in the $\mathbf{Z}$ direction $[0,0,-1]$, the rotated $y$ axis (second column) is at the $\mathbf{Y}$ direction $[0,1,0]$, and the rotated $z$ axis (third column) is at the $\mathbf{X}$ direction $[1,0,0]$.
We also can calculate the sequence of elemental rotations around the $x$, $y$, $z$ axes of the rotating $xyz$ coordinate system illustrated in the next figure.
<br>
<figure>
<img src='./../images/rotations_xyz2.png' alt='rotations'/> <figcaption><center><i>Figure. Sequence of elemental rotations of a second $xyz$ local coordinate system around each axis, $x$, $y$, $z$, of the rotating $xyz$ coordinate system.</i></center></figcaption>
</figure>
Likewise, this sequence of elemental rotations (each one of the local coordinate system with respect to the rotating local coordinate system) is mathematically represented by a multiplication between the rotation matrices (which are the inverse of the matrices for the rotations around $\mathbf{X,Y,Z}$ as we saw earlier):
$$ \begin{array}{l l}
\mathbf{R}_{\mathbf{lG},\,xyz} & = \mathbf{R_{z}} \mathbf{R_{y}} \mathbf{R_{x}} \\
\\
& = \begin{bmatrix}
\cos\gamma & \sin\gamma & 0\\
-\sin\gamma & \cos\gamma & 0 \\
0 & 0 & 1
\end{bmatrix}
\begin{bmatrix}
\cos\beta & 0 & -\sin\beta \\
0 & 1 & 0 \\
sin\beta & 0 & \cos\beta
\end{bmatrix}
\begin{bmatrix}
1 & 0 & 0 \\
0 & \cos\alpha & \sin\alpha \\
0 & -\sin\alpha & \cos\alpha
\end{bmatrix} \\
\\
& =
\begin{bmatrix}
\cos\beta\:\cos\gamma \;&\;
\sin\alpha\:\sin\beta\:\cos\gamma+\cos\alpha\:\sin\gamma \;&\;
\cos\alpha\:\sin\beta\:\cos\gamma-\sin\alpha\:\sin\gamma \;\;\; \\
-\cos\beta\:\sin\gamma \;&\;
-\sin\alpha\:\sin\beta\:\sin\gamma+\cos\alpha\:\cos\gamma \;&\;
\cos\alpha\:\sin\beta\:\sin\gamma+\sin\alpha\:\cos\gamma \;\;\; \\
\sin\beta \;&\; -\sin\alpha\:\cos\beta \;&\; \cos\alpha\:\cos\beta \;\;\;
\end{bmatrix}
\end{array} $$
As before, the order of the matrices is from right to left.
Once again, we can check this matrix multiplication using [Sympy](http://sympy.org/en/index.html):
```
a, b, g = sym.symbols('alpha, beta, gamma')
# Elemental rotation matrices of xyz (local):
Rx = sym.Matrix([[1, 0, 0], [0, cos(a), sin(a)], [0, -sin(a), cos(a)]])
Ry = sym.Matrix([[cos(b), 0, -sin(b)], [0, 1, 0], [sin(b), 0, cos(b)]])
Rz = sym.Matrix([[cos(g), sin(g), 0], [-sin(g), cos(g), 0], [0, 0, 1]])
# Rotation matrix of xyz' in relation to xyz:
Rxyz = Rz*Ry*Rx
Math(sym.latex(r'\mathbf{R}_{\mathbf{lG},\,xyz}=') + sym.latex(Rxyz, mat_str='matrix'))
```
For instance, let's calculate the numerical rotation matrix for these sequential elemental rotations by $90^o$ around $x,y,z$:
```
R = sym.lambdify((a, b, g), Rxyz, 'numpy')
R = R(np.pi/2, np.pi/2, np.pi/2)
display(Math(r'\mathbf{R}_{\mathbf{lG},\,xyz\,}(90^o, 90^o, 90^o) =' + \
sym.latex(sym.Matrix(R).n(chop=True, prec=3))))
```
Once again, let's compare the above matrix and the correspondent previous figure to see if it makes sense. But remember that this matrix is the Global-to-local rotation matrix, $\mathbf{R}_{\mathbf{lG},\,xyz}$, where the coordinates of the local basis' versors are rows, not columns, in this matrix. With this detail in mind, one can see that the previous figure and matrix also agree: the rotated $x$ axis (first row of the above matrix) is at the $\mathbf{Z}$ direction $[0,0,1]$, the rotated $y$ axis (second row) is at the $\mathbf{-Y}$ direction $[0,-1,0]$, and the rotated $z$ axis (third row) is at the $\mathbf{X}$ direction $[1,0,0]$.
In fact, this example didn't serve to distinguish versors as rows or columns because the $\mathbf{R}_{\mathbf{lG},\,xyz}$ matrix above is symmetric!
Let's look on the resultant matrix for the example above after only the first two rotations, $\mathbf{R}_{\mathbf{lG},\,xy}$ to understand this difference:
```
Rxy = Ry*Rx
R = sym.lambdify((a, b), Rxy, 'numpy')
R = R(np.pi/2, np.pi/2)
display(Math(r'\mathbf{R}_{\mathbf{lG},\,xy\,}(90^o, 90^o) =' + \
sym.latex(sym.Matrix(R).n(chop=True, prec=3))))
```
Comparing this matrix with the third plot in the figure, we see that the coordinates of versor $x$ in the Global coordinate system are $[0,1,0]$, i.e., local axis $x$ is aligned with Global axis $Y$, and this versor is indeed the first row, not first column, of the matrix above. Confer the other two rows.
What are then in the columns of the local-to-Global rotation matrix?
The columns are the coordinates of Global basis' versors in the local coordinate system! For example, the first column of the matrix above is the coordinates of $X$, which is aligned with $z$: $[0,0,1]$.
#### Rotations in a coordinate system is equivalent to minus rotations in the other coordinate system
Remember that we saw for the elemental rotations that it's equivalent to instead of rotating the local coordinate system, $xyz$, by $\alpha, \beta, \gamma$ in relation to axes of the Global coordinate system, to rotate the Global coordinate system, $\mathbf{XYZ}$, by $-\alpha, -\beta, -\gamma$ in relation to the axes of the local coordinate system. The same property applies to a sequence of rotations: rotations of $xyz$ in relation to $\mathbf{XYZ}$ by $\alpha, \beta, \gamma$ result in the same matrix as rotations of $\mathbf{XYZ}$ in relation to $xyz$ by $-\alpha, -\beta, -\gamma$:
$$ \begin{array}{l l}
\mathbf{R_{Gl,\,XYZ\,}}(\alpha,\beta,\gamma) & = \mathbf{R_{Gl,\,Z}}(\gamma)\, \mathbf{R_{Gl,\,Y}}(\beta)\, \mathbf{R_{Gl,\,X}}(\alpha) \\
& = \mathbf{R}_{\mathbf{lG},\,z\,}(-\gamma)\, \mathbf{R}_{\mathbf{lG},\,y\,}(-\beta)\, \mathbf{R}_{\mathbf{lG},\,x\,}(-\alpha) \\
& = \mathbf{R}_{\mathbf{lG},\,xyz\,}(-\alpha,-\beta,-\gamma)
\end{array}
$$
Confer that by examining the $\mathbf{R_{Gl,\,XYZ}}$ and $\mathbf{R}_{\mathbf{lG},\,xyz}$ matrices above.
Let's verify this property with Sympy:
```
RXYZ = RZ*RY*RX
# Rotation matrix of xyz in relation to XYZ:
display(Math(sym.latex(r'\mathbf{R_{Gl,\,XYZ\,}}(\alpha,\beta,\gamma) =')))
display(Math(sym.latex(RXYZ, mat_str='matrix')))
# Elemental rotation matrices of XYZ in relation to xyz and negate all angles:
Rx_neg = sym.Matrix([[1, 0, 0], [0, cos(-a), -sin(-a)], [0, sin(-a), cos(-a)]]).T
Ry_neg = sym.Matrix([[cos(-b), 0, sin(-b)], [0, 1, 0], [-sin(-b), 0, cos(-b)]]).T
Rz_neg = sym.Matrix([[cos(-g), -sin(-g), 0], [sin(-g), cos(-g), 0], [0, 0, 1]]).T
# Rotation matrix of XYZ in relation to xyz:
Rxyz_neg = Rz_neg*Ry_neg*Rx_neg
display(Math(sym.latex(r'\mathbf{R}_{\mathbf{lG},\,xyz\,}(-\alpha,-\beta,-\gamma) =')))
display(Math(sym.latex(Rxyz_neg, mat_str='matrix')))
# Check that the two matrices are equal:
display(Math(sym.latex(r'\mathbf{R_{Gl,\,XYZ\,}}(\alpha,\beta,\gamma) \;==\;' + \
r'\mathbf{R}_{\mathbf{lG},\,xyz\,}(-\alpha,-\beta,-\gamma)')))
RXYZ == Rxyz_neg
```
#### Rotations in a coordinate system is the transpose of inverse order of rotations in the other coordinate system
There is another property of the rotation matrices for the different coordinate systems: the rotation matrix, for example from the Global to the local coordinate system for the $xyz$ sequence, is just the transpose of the rotation matrix for the inverse operation (from the local to the Global coordinate system) of the inverse sequence ($\mathbf{ZYX}$) and vice-versa:
$$ \begin{array}{l l}
\mathbf{R}_{\mathbf{lG},\,xyz}(\alpha,\beta,\gamma) & = \mathbf{R}_{\mathbf{lG},\,z\,} \mathbf{R}_{\mathbf{lG},\,y\,} \mathbf{R}_{\mathbf{lG},\,x} \\
& = \mathbf{R_{Gl,\,Z\,}^{-1}} \mathbf{R_{Gl,\,Y\,}^{-1}} \mathbf{R_{Gl,\,X\,}^{-1}} \\
& = \mathbf{R_{Gl,\,Z\,}^{T}} \mathbf{R_{Gl,\,Y\,}^{T}} \mathbf{R_{Gl,\,X\,}^{T}} \\
& = (\mathbf{R_{Gl,\,X\,}} \mathbf{R_{Gl,\,Y\,}} \mathbf{R_{Gl,\,Z}})^\mathbf{T} \\
& = \mathbf{R_{Gl,\,ZYX\,}^{T}}(\gamma,\beta,\alpha)
\end{array}
$$
Where we used the properties that the inverse of the rotation matrix (which is orthonormal) is its transpose and that the transpose of a product of matrices is equal to the product of their transposes in reverse order.
Let's verify this property with Sympy:
```
RZYX = RX*RY*RZ
Rxyz = Rz*Ry*Rx
display(Math(sym.latex(r'\mathbf{R_{Gl,\,ZYX\,}^T}=') + sym.latex(RZYX.T, mat_str='matrix')))
display(Math(sym.latex(r'\mathbf{R}_{\mathbf{lG},\,xyz\,}(\alpha,\beta,\gamma) \,==\,' + \
r'\mathbf{R_{Gl,\,ZYX\,}^T}(\gamma,\beta,\alpha)')))
Rxyz == RZYX.T
```
#### Sequence of rotations of a Vector
We saw in the notebook [Rigid-body transformations in a plane (2D)](http://nbviewer.jupyter.org/github/BMClab/bmc/blob/master/notebooks/Transformation2D.ipynb#Rotation-of-a-Vector) that the rotation matrix can also be used to rotate a vector (in fact, a point, image, solid, etc.) by a given angle around an axis of the coordinate system. Let's investigate that for the 3D case using the example earlier where a book was rotated in different orders and around the Global and local coordinate systems.
Before any rotation, the point shown in that figure as a round black dot on the spine of the book has coordinates $\mathbf{P}=[0, 1, 2]$ (the book has thickness 0, width 1, and height 2).
After the first sequence of rotations shown in the figure (rotated around $X$ and $Y$ by $90^0$ each time), $\mathbf{P}$ has coordinates $\mathbf{P}=[1, -2, 0]$ in the global coordinate system. Let's verify that:
```
P = np.array([[0, 1, 2]]).T
RXY = RY*RX
R = sym.lambdify((a, b), RXY, 'numpy')
R = R(np.pi/2, np.pi/2)
P1 = np.dot(R, P)
print('P1 =', P1.T)
```
As expected.
The reader is invited to deduce the position of point $\mathbf{P}$ after the inverse order of rotations, but still around the Global coordinate system.
Although we are performing vector rotation, where we don't need the concept of transformation between coordinate systems, in the example above we used the local-to-Global rotation matrix, $\mathbf{R_{Gl}}$. As we saw in the notebook for the 2D transformation, when we use this matrix, it performs a counter-clockwise (positive) rotation.
If we want to rotate the vector in the clockwise (negative) direction, we can use the very same rotation matrix entering a negative angle or we can use the inverse rotation matrix, the Global-to-local rotation matrix, $\mathbf{R_{lG}}$ and a positive (negative of negative) angle, because $\mathbf{R_{Gl}}(\alpha) = \mathbf{R_{lG}}(-\alpha)$, but bear in mind that even in this latter case we are rotating around the Global coordinate system!
Consider now that we want to deduce algebraically the position of the point $\mathbf{P}$ after the rotations around the local coordinate system as shown in the second set of examples in the figure with the sequence of book rotations. The point has the same initial position, $\mathbf{P}=[0, 1, 2]$, and after the rotations around $x$ and $y$ by $90^0$ each time, what is the position of this point?
It's implicit in this question that the new desired position is in the Global coordinate system because the local coordinate system rotates with the book and the point never changes its position in the local coordinate system. So, by inspection of the figure, the new position of the point is $\mathbf{P1}=[2, 0, 1]$.
Let's naively try to deduce this position by repeating the steps as before:
```
Rxy = Ry*Rx
R = sym.lambdify((a, b), Rxy, 'numpy')
R = R(np.pi/2, np.pi/2)
P1 = np.dot(R, P)
print('P1 =', P1.T)
```
The wrong answer.
The problem is that we defined the rotation of a vector using the local-to-Global rotation matrix. One correction solution for this problem is to continuing using the multiplication of the Global-to-local rotation matrices, $\mathbf{R}_{xy} = \mathbf{R}_y\,\mathbf{R}_x$, transpose $\mathbf{R}_{xy}$ to get the Global-to-local coordinate system, $\mathbf{R_{XY}}=\mathbf{R^T}_{xy}$, and then rotate the vector using this matrix:
```
Rxy = Ry*Rx
RXY = Rxy.T
R = sym.lambdify((a, b), RXY, 'numpy')
R = R(np.pi/2, np.pi/2)
P1 = np.dot(R, P)
print('P1 =', P1.T)
```
The correct answer.
Another solution is to understand that when using the Global-to-local rotation matrix, counter-clockwise rotations (as performed with the book the figure) are negative, not positive, and that when dealing with rotations with the Global-to-local rotation matrix the order of matrix multiplication is inverted, for example, it should be $\mathbf{R\_}_{xyz} = \mathbf{R}_x\,\mathbf{R}_y\,\mathbf{R}_z$ (an added underscore to remind us this is not the convention adopted here).
```
R_xy = Rx*Ry
R = sym.lambdify((a, b), R_xy, 'numpy')
R = R(-np.pi/2, -np.pi/2)
P1 = np.dot(R, P)
print('P1 =', P1.T)
```
The correct answer.
The reader is invited to deduce the position of point $\mathbf{P}$ after the inverse order of rotations, around the local coordinate system.
In fact, you will find elsewhere texts about rotations in 3D adopting this latter convention as the standard, i.e., they introduce the Global-to-local rotation matrix and describe sequence of rotations algebraically as matrix multiplication in the direct order, $\mathbf{R\_}_{xyz} = \mathbf{R}_x\,\mathbf{R}_y\,\mathbf{R}_z$, the inverse we have done in this text. It's all a matter of convention, just that.
#### The 12 different sequences of Euler angles
The Euler angles are defined in terms of rotations around a rotating local coordinate system. As we saw for the sequence of rotations around $x, y, z$, the axes of the local rotated coordinate system are not fixed in space because after the first elemental rotation, the other two axes rotate.
Other sequences of rotations could be produced without combining axes of the two different coordinate systems (Global and local) for the definition of the rotation axes. There is a total of 12 different sequences of three elemental rotations that are valid and may be used for describing the rotation of a coordinate system with respect to another coordinate system:
$$ xyz \quad xzy \quad yzx \quad yxz \quad zxy \quad zyx $$
$$ xyx \quad xzx \quad yzy \quad yxy \quad zxz \quad zyz $$
The first six sequences (first row) are all around different axes, they are usually referred as Cardan or Tait–Bryan angles. The other six sequences (second row) have the first and third rotations around the same axis, but keep in mind that the axis for the third rotation is not at the same place anymore because it changed its orientation after the second rotation. The sequences with repeated axes are known as proper or classic Euler angles.
Which order to use it is a matter of convention, but because the order affects the results, it's fundamental to follow a convention and report it. In Engineering Mechanics (including Biomechanics), the $xyz$ order is more common; in Physics the $zxz$ order is more common (but the letters chosen to refer to the axes are arbitrary, what matters is the directions they represent). In Biomechanics, the order for the Cardan angles is most often based on the angle of most interest or of most reliable measurement. Accordingly, the axis of flexion/extension is typically selected as the first axis, the axis for abduction/adduction is the second, and the axis for internal/external rotation is the last one. We will see about this order later. The $zyx$ order is commonly used to describe the orientation of a ship or aircraft and the rotations are known as the nautical angles: yaw, pitch and roll, respectively (see next figure).
<br>
<figure><img src='https://upload.wikimedia.org/wikipedia/commons/thumb/1/16/Yaw_Axis.svg/319px-Yaw_Axis.svg.png' alt='translation and rotation 3D'/> <figcaption><center><i>Figure. The principal axes of an aircraft and the names for the rotations around these axes (<a href="https://en.wikipedia.org/wiki/Euler_angles">image from Wikipedia</a>).</i></center></figcaption> </figure>
If instead of rotations around the rotating local coordinate system we perform rotations around the fixed Global coordinate system, we will have other 12 different sequences of three elemental rotations, these are called simply rotation angles. So, in total there are 24 possible different sequences of three elemental rotations, but the 24 orders are not independent; with the 12 different sequences of Euler angles at the local coordinate system we can obtain the other 12 sequences at the Global coordinate system.
The Python function `euler_rotmat.py` (code at the end of this text) determines the rotation matrix in algebraic form for any of the 24 different sequences (and sequences with only one or two axes can be inputed). This function also determines the rotation matrix in numeric form if a list of up to three angles are inputed.
For instance, the rotation matrix in algebraic form for the $zxz$ order of Euler angles at the local coordinate system and the correspondent rotation matrix in numeric form after three elemental rotations by $90^o$ each are:
```
import sys
sys.path.insert(1, r'./../functions')
from euler_rotmat import euler_rotmat
Ra, Rn = euler_rotmat(order='zxz', frame='local', angles=[90, 90, 90])
```
#### Line of nodes
The second axis of rotation in the rotating coordinate system is also referred as the nodal axis or line of nodes; this axis coincides with the intersection of two perpendicular planes, one from each Global (fixed) and local (rotating) coordinate systems. The figure below shows an example of rotations and the nodal axis for the $xyz$ sequence of the Cardan angles.
<div class='center-align'><figure><img src='./../images/Node.png' alt='rotations'/> <figcaption><center><i>Figure. First row: example of rotations for the $xyz$ sequence of the Cardan angles. The Global (fixed) $XYZ$ coordinate system is shown in green, the local (rotating) $xyz$ coordinate system is shown in blue. The nodal axis (<b>N</b>, shown in red) is defined by the intersection of the $YZ$ and $xy$ planes and all rotations can be described in relation to this nodal axis or to a perpendicular axis to it. Second row: starting from no rotation, the local coordinate system is rotated by $\alpha$ around the $x$ axis, then by $\beta$ around the rotated $y$ axis, and finally by $\gamma$ around the twice rotated $z$ axis. Note that the line of nodes coincides with the $y$ axis for the second rotation. </i></center></figcaption> </figure></div>
#### Determination of the Euler angles
Once a convention is adopted, the corresponding three Euler angles of rotation can be found.
For example, for the $\mathbf{R}_{xyz}$ rotation matrix:
```
R = euler_rotmat(order='xyz', frame='local')
```
The corresponding Cardan angles for the `xyz` sequence can be given by:
$$ \begin{array}{}
\alpha = \arctan\left(\dfrac{\sin(\alpha)}{\cos(\alpha)}\right) = \arctan\left(\dfrac{-\mathbf{R}_{21}}{\;\;\;\mathbf{R}_{22}}\right) \\
\\
\beta = \arctan\left(\dfrac{\sin(\beta)}{\cos(\beta)}\right) = \arctan\left(\dfrac{\mathbf{R}_{20}}{\sqrt{\mathbf{R}_{00}^2+\mathbf{R}_{10}^2}}\right) \\
\\
\gamma = \arctan\left(\dfrac{\sin(\gamma)}{\cos(\gamma)}\right) = \arctan\left(\dfrac{-\mathbf{R}_{10}}{\;\;\;\mathbf{R}_{00}}\right)
\end{array} $$
Note that we prefer to use the mathematical function `arctan` rather than simply `arcsin` because the latter cannot for example distinguish $45^o$ from $135^o$ and also for better numerical accuracy. See the text [Angular kinematics in a plane (2D)](https://nbviewer.jupyter.org/github/BMClab/bmc/blob/master/notebooks/KinematicsAngular2D.ipynb) for more on these issues.
And here is a Python function to compute the Euler angles of rotations from the Global to the local coordinate system for the $xyz$ Cardan sequence:
```
def euler_angles_from_rot_xyz(rot_matrix, unit='deg'):
""" Compute Euler angles from rotation matrix in the xyz sequence."""
import numpy as np
R = np.array(rot_matrix, copy=False).astype(np.float64)[:3, :3]
angles = np.zeros(3)
angles[0] = np.arctan2(-R[2, 1], R[2, 2])
angles[1] = np.arctan2( R[2, 0], np.sqrt(R[0, 0]**2 + R[1, 0]**2))
angles[2] = np.arctan2(-R[1, 0], R[0, 0])
if unit[:3].lower() == 'deg': # convert from rad to degree
angles = np.rad2deg(angles)
return angles
```
For instance, consider sequential rotations of 45$^o$ around $x,y,z$. The resultant rotation matrix is:
```
Ra, Rn = euler_rotmat(order='xyz', frame='local', angles=[45, 45, 45], showA=False)
```
Let's check that calculating back the Cardan angles from this rotation matrix using the `euler_angles_from_rot_xyz()` function:
```
euler_angles_from_rot_xyz(Rn, unit='deg')
```
We could implement a function to calculate the Euler angles for any of the 12 sequences (in fact, plus another 12 sequences if we consider all the rotations from and to the two coordinate systems), but this is tedious. There is a smarter solution using the concept of [quaternion](http://en.wikipedia.org/wiki/Quaternion), but we wont see that now.
Let's see a problem with using Euler angles known as gimbal lock.
### Gimbal lock
[Gimbal lock](http://en.wikipedia.org/wiki/Gimbal_lock) is the loss of one degree of freedom in a three-dimensional coordinate system that occurs when an axis of rotation is placed parallel with another previous axis of rotation and two of the three rotations will be around the same direction given a certain convention of the Euler angles. This "locks" the system into rotations in a degenerate two-dimensional space. The system is not really locked in the sense it can't be moved or reach the other degree of freedom, but it will need an extra rotation for that.
For instance, let's look at the $zxz$ sequence of rotations by the angles $\alpha, \beta, \gamma$:
$$ \begin{array}{l l}
\mathbf{R}_{zxz} & = \mathbf{R_{z}} \mathbf{R_{x}} \mathbf{R_{z}} \\
\\
& =
\begin{bmatrix}
\cos\gamma & \sin\gamma & 0\\
-\sin\gamma & \cos\gamma & 0 \\
0 & 0 & 1
\end{bmatrix}
\begin{bmatrix}
1 & 0 & 0 \\
0 & \cos\beta & \sin\beta \\
0 & -\sin\beta & \cos\beta
\end{bmatrix}
\begin{bmatrix}
\cos\alpha & \sin\alpha & 0\\
-\sin\alpha & \cos\alpha & 0 \\
0 & 0 & 1
\end{bmatrix}
\end{array} $$
Which results in:
```
a, b, g = sym.symbols('alpha, beta, gamma')
# Elemental rotation matrices of xyz (local):
Rz = sym.Matrix([[cos(a), sin(a), 0], [-sin(a), cos(a), 0], [0, 0, 1]])
Rx = sym.Matrix([[1, 0, 0], [0, cos(b), sin(b)], [0, -sin(b), cos(b)]])
Rz2 = sym.Matrix([[cos(g), sin(g), 0], [-sin(g), cos(g), 0], [0, 0, 1]])
# Rotation matrix for the zxz sequence:
Rzxz = Rz2*Rx*Rz
Math(sym.latex(r'\mathbf{R}_{zxz}=') + sym.latex(Rzxz, mat_str='matrix'))
```
Let's examine what happens with this rotation matrix when the rotation around the second axis ($x$) by $\beta$ is zero:
$$ \begin{array}{l l}
\mathbf{R}_{zxz}(\alpha, \beta=0, \gamma) =
\begin{bmatrix}
\cos\gamma & \sin\gamma & 0\\
-\sin\gamma & \cos\gamma & 0 \\
0 & 0 & 1
\end{bmatrix}
\begin{bmatrix}
1 & 0 & 0 \\
0 & 1 & 0 \\
0 & 0 & 1
\end{bmatrix}
\begin{bmatrix}
\cos\alpha & \sin\alpha & 0\\
-\sin\alpha & \cos\alpha & 0 \\
0 & 0 & 1
\end{bmatrix}
\end{array} $$
The second matrix is the identity matrix and has no effect on the product of the matrices, which will be:
```
Rzxz = Rz2*Rz
Math(sym.latex(r'\mathbf{R}_{xyz}(\alpha, \beta=0, \gamma)=') + \
sym.latex(Rzxz, mat_str='matrix'))
```
Which simplifies to:
```
Rzxz = sym.simplify(Rzxz)
Math(sym.latex(r'\mathbf{R}_{xyz}(\alpha, \beta=0, \gamma)=') + \
sym.latex(Rzxz, mat_str='matrix'))
```
Despite different values of $\alpha$ and $\gamma$ the result is a single rotation around the $z$ axis given by the sum $\alpha+\gamma$. In this case, of the three degrees of freedom one was lost (the other degree of freedom was set by $\beta=0$). For movement analysis, this means for example that one angle will be undetermined because everything we know is the sum of the two angles obtained from the rotation matrix. We can set the unknown angle to zero but this is arbitrary.
In fact, we already dealt with another example of gimbal lock when we looked at the $xyz$ sequence with rotations by $90^o$. See the figure representing these rotations again and perceive that the first and third rotations were around the same axis because the second rotation was by $90^o$. Let's do the matrix multiplication replacing only the second angle by $90^o$ (and let's use the `euler_rotmat.py`:
```
Ra, Rn = euler_rotmat(order='xyz', frame='local', angles=[None, 90., None], showA=False)
```
Once again, one degree of freedom was lost and we will not be able to uniquely determine the three angles for the given rotation matrix and sequence.
Possible solutions to avoid the gimbal lock are: choose a different sequence; do not rotate the system by the angle that puts the system in gimbal lock (in the examples above, avoid $\beta=90^o$); or add an extra fourth parameter in the description of the rotation angles.
But if we have a physical system where we measure or specify exactly three Euler angles in a fixed sequence to describe or control it, and we can't avoid the system to assume certain angles, then we might have to say "Houston, we have a problem".
A famous situation where such a problem occurred was during the Apollo 13 mission. This is an actual conversation between crew and mission control during the Apollo 13 mission (Corke, 2011):
>`Mission clock: 02 08 12 47`
**Flight**: *Go, Guidance.*
**Guido**: *He’s getting close to gimbal lock there.*
**Flight**: *Roger. CapCom, recommend he bring up C3, C4, B3, B4, C1 and C2 thrusters, and advise he’s getting close to gimbal lock.*
**CapCom**: *Roger.*
*Of note, it was not a gimbal lock that caused the accident with the the Apollo 13 mission, the problem was an oxygen tank explosion.*
## Determination of the rotation matrix
A typical way to determine the rotation matrix for a rigid body in biomechanics is to use motion analysis to measure the position of at least three non-collinear markers placed on the rigid body, and then calculate a basis with these positions, analogue to what we have described in the notebook [Rigid-body transformations in a plane (2D)](http://nbviewer.ipython.org/github/BMClab/bmc/blob/master/notebooks/Transformation2D.ipynb).
### Basis
If we have the position of three markers: **m1**, **m2**, **m3**, a basis (formed by three orthogonal versors) can be found as:
- First axis, **v1**, the vector **m2-m1**;
- Second axis, **v2**, the cross product between the vectors **v1** and **m3-m1**;
- Third axis, **v3**, the cross product between the vectors **v1** and **v2**.
Then, each of these vectors are normalized resulting in three orthogonal versors.
For example, given the positions m1 = [1,0,0], m2 = [0,1,0], m3 = [0,0,1], a basis can be found:
```
m1 = np.array([1, 0, 0])
m2 = np.array([0, 1, 0])
m3 = np.array([0, 0, 1])
v1 = m2 - m1
v2 = np.cross(v1, m3 - m1)
v3 = np.cross(v1, v2)
print('Versors:')
v1 = v1/np.linalg.norm(v1)
print('v1 =', v1)
v2 = v2/np.linalg.norm(v2)
print('v2 =', v2)
v3 = v3/np.linalg.norm(v3)
print('v3 =', v3)
print('\nNorm of each versor:\n',
np.linalg.norm(np.cross(v1, v2)),
np.linalg.norm(np.cross(v1, v3)),
np.linalg.norm(np.cross(v2, v3)))
```
Remember from the text [Rigid-body transformations in a plane (2D)](http://nbviewer.ipython.org/github/BMClab/bmc/blob/master/notebooks/Transformation2D.ipynb) that the versors of this basis are the columns of the $\mathbf{R_{Gl}}$ and the rows of the $\mathbf{R_{lG}}$ rotation matrices, for instance:
```
RlG = np.array([v1, v2, v3])
print('Rotation matrix from Global to local coordinate system:\n', RlG)
```
And the corresponding angles of rotation using the $xyz$ sequence are:
```
euler_angles_from_rot_xyz(RlG)
```
These angles don't mean anything now because they are angles of the axes of the arbitrary basis we computed. In biomechanics, if we want an anatomical interpretation of the coordinate system orientation, we define the versors of the basis oriented with anatomical axes (e.g., for the shoulder, one versor would be aligned with the long axis of the upper arm).
We will see how to perform this computation later. Now we will combine translation and rotation in a single transformation.
## Translation and Rotation
Consider the case where the local coordinate system is translated and rotated in relation to the Global coordinate system as illustrated in the next figure.
<br>
<figure><img src='./../images/transrot3D.png' alt='translation and rotation 3D'/> <figcaption><center><i>Figure. A point in three-dimensional space represented in two coordinate systems, with one system translated and rotated.</i></center></figcaption> </figure>
The position of point $\mathbf{P}$ originally described in the local coordinate system, but now described in the Global coordinate system in vector form is:
$$ \mathbf{P_G} = \mathbf{L_G} + \mathbf{R_{Gl}}\mathbf{P_l} $$
This means that we first *disrotate* the local coordinate system and then correct for the translation between the two coordinate systems. Note that we can't invert this order: the point position is expressed in the local coordinate system and we can't add this vector to another vector expressed in the Global coordinate system, first we have to convert the vectors to the same coordinate system.
If now we want to find the position of a point at the local coordinate system given its position in the Global coordinate system, the rotation matrix and the translation vector, we have to invert the expression above:
$$ \begin{array}{l l}
\mathbf{P_G} = \mathbf{L_G} + \mathbf{R_{Gl}}\mathbf{P_l} \implies \\
\\
\mathbf{R_{Gl}^{-1}}\cdot\mathbf{P_G} = \mathbf{R_{Gl}^{-1}}\left(\mathbf{L_G} + \mathbf{R_{Gl}}\mathbf{P_l}\right) \implies \\
\\
\mathbf{R_{Gl}^{-1}}\mathbf{P_G} = \mathbf{R_{Gl}^{-1}}\mathbf{L_G} + \mathbf{R_{Gl}^{-1}}\mathbf{R_{Gl}}\mathbf{P_l} \implies \\
\\
\mathbf{P_l} = \mathbf{R_{Gl}^{-1}}\left(\mathbf{P_G}-\mathbf{L_G}\right) = \mathbf{R_{Gl}^T}\left(\mathbf{P_G}-\mathbf{L_G}\right) \;\;\;\;\; \text{or} \;\;\;\;\; \mathbf{P_l} = \mathbf{R_{lG}}\left(\mathbf{P_G}-\mathbf{L_G}\right)
\end{array} $$
The expression above indicates that to perform the inverse operation, to go from the Global to the local coordinate system, we first translate and then rotate the coordinate system.
### Transformation matrix
It is possible to combine the translation and rotation operations in only one matrix, called the transformation matrix:
$$ \begin{bmatrix}
\mathbf{P_X} \\
\mathbf{P_Y} \\
\mathbf{P_Z} \\
1
\end{bmatrix} =
\begin{bmatrix}
. & . & . & \mathbf{L_{X}} \\
. & \mathbf{R_{Gl}} & . & \mathbf{L_{Y}} \\
. & . & . & \mathbf{L_{Z}} \\
0 & 0 & 0 & 1
\end{bmatrix}
\begin{bmatrix}
\mathbf{P}_x \\
\mathbf{P}_y \\
\mathbf{P}_z \\
1
\end{bmatrix} $$
Or simply:
$$ \mathbf{P_G} = \mathbf{T_{Gl}}\mathbf{P_l} $$
Remember that in general the transformation matrix is not orthonormal, i.e., its inverse is not equal to its transpose.
The inverse operation, to express the position at the local coordinate system in terms of the Global reference system, is:
$$ \mathbf{P_l} = \mathbf{T_{Gl}^{-1}}\mathbf{P_G} $$
And in matrix form:
$$ \begin{bmatrix}
\mathbf{P_x} \\
\mathbf{P_y} \\
\mathbf{P_z} \\
1
\end{bmatrix} =
\begin{bmatrix}
\cdot & \cdot & \cdot & \cdot \\
\cdot & \mathbf{R^{-1}_{Gl}} & \cdot & -\mathbf{R^{-1}_{Gl}}\:\mathbf{L_G} \\
\cdot & \cdot & \cdot & \cdot \\
0 & 0 & 0 & 1
\end{bmatrix}
\begin{bmatrix}
\mathbf{P_X} \\
\mathbf{P_Y} \\
\mathbf{P_Z} \\
1
\end{bmatrix} $$
### Example with actual motion analysis data
*The data for this example is taken from page 183 of David Winter's book.*
Consider the following marker positions placed on a leg (described in the laboratory coordinate system with coordinates $x, y, z$ in cm, the $x$ axis points forward and the $y$ axes points upward): lateral malleolus (**lm** = [2.92, 10.10, 18.85]), medial malleolus (**mm** = [2.71, 10.22, 26.52]), fibular head (**fh** = [5.05, 41.90, 15.41]), and medial condyle (**mc** = [8.29, 41.88, 26.52]). Define the ankle joint center as the centroid between the **lm** and **mm** markers and the knee joint center as the centroid between the **fh** and **mc** markers. An anatomical coordinate system for the leg can be defined as: the quasi-vertical axis ($y$) passes through the ankle and knee joint centers; a temporary medio-lateral axis ($z$) passes through the two markers on the malleolus, an anterior-posterior as the cross product between the two former calculated orthogonal axes, and the origin at the ankle joint center.
a) Calculate the anatomical coordinate system for the leg as described above.
b) Calculate the rotation matrix and the translation vector for the transformation from the anatomical to the laboratory coordinate system.
c) Calculate the position of each marker and of each joint center at the anatomical coordinate system.
d) Calculate the Cardan angles using the $zxy$ sequence for the orientation of the leg with respect to the laboratory (but remember that the letters chosen to refer to axes are arbitrary, what matters is the directions they represent).
```
# calculation of the joint centers
mm = np.array([2.71, 10.22, 26.52])
lm = np.array([2.92, 10.10, 18.85])
fh = np.array([5.05, 41.90, 15.41])
mc = np.array([8.29, 41.88, 26.52])
ajc = (mm + lm)/2
kjc = (fh + mc)/2
print('Poition of the ankle joint center:', ajc)
print('Poition of the knee joint center:', kjc)
# calculation of the anatomical coordinate system axes (basis)
y = kjc - ajc
x = np.cross(y, mm - lm)
z = np.cross(x, y)
print('Versors:')
x = x/np.linalg.norm(x)
y = y/np.linalg.norm(y)
z = z/np.linalg.norm(z)
print('x =', x)
print('y =', y)
print('z =', z)
Oleg = ajc
print('\nOrigin =', Oleg)
# Rotation matrices
RGl = np.array([x, y , z]).T
print('Rotation matrix from the anatomical to the laboratory coordinate system:\n', RGl)
RlG = RGl.T
print('\nRotation matrix from the laboratory to the anatomical coordinate system:\n', RlG)
# Translational vector
OG = np.array([0, 0, 0]) # Laboratory coordinate system origin
LG = Oleg - OG
print('Translational vector from the anatomical to the laboratory coordinate system:\n', LG)
```
To get the coordinates from the laboratory (global) coordinate system to the anatomical (local) coordinate system:
$$ \mathbf{P_l} = \mathbf{R_{lG}}\left(\mathbf{P_G}-\mathbf{L_G}\right) $$
```
# position of each marker and of each joint center at the anatomical coordinate system
mml = np.dot(RlG, (mm - LG)) # equivalent to the algebraic expression RlG*(mm - LG).T
lml = np.dot(RlG, (lm - LG))
fhl = np.dot(RlG, (fh - LG))
mcl = np.dot(RlG, (mc - LG))
ajcl = np.dot(RlG, (ajc - LG))
kjcl = np.dot(RlG, (kjc - LG))
print('Coordinates of mm in the anatomical system:\n', mml)
print('Coordinates of lm in the anatomical system:\n', lml)
print('Coordinates of fh in the anatomical system:\n', fhl)
print('Coordinates of mc in the anatomical system:\n', mcl)
print('Coordinates of kjc in the anatomical system:\n', kjcl)
print('Coordinates of ajc in the anatomical system (origin):\n', ajcl)
```
## Problems
1. For the example about how the order of rotations of a rigid body affects the orientation shown in a figure above, deduce the rotation matrices for each of the 4 cases shown in the figure. For the first two cases, deduce the rotation matrices from the global to the local coordinate system and for the other two examples, deduce the rotation matrices from the local to the global coordinate system.
2. Consider the data from problem 7 in the notebook [Frame of reference](http://nbviewer.ipython.org/github/BMClab/bmc/blob/master/notebooks/ReferenceFrame.ipynb) where the following anatomical landmark positions are given (units in meters): RASIS=[0.5,0.8,0.4], LASIS=[0.55,0.78,0.1], RPSIS=[0.3,0.85,0.2], and LPSIS=[0.29,0.78,0.3]. Deduce the rotation matrices for the global to anatomical coordinate system and for the anatomical to global coordinate system.
3. For the data from the last example, calculate the Cardan angles using the $zxy$ sequence for the orientation of the leg with respect to the laboratory (but remember that the letters chosen to refer to axes are arbitrary, what matters is the directions they represent).
## References
- Corke P (2011) [Robotics, Vision and Control: Fundamental Algorithms in MATLAB](http://www.petercorke.com/RVC/). Springer-Verlag Berlin.
- Robertson G, Caldwell G, Hamill J, Kamen G (2013) [Research Methods in Biomechanics](http://books.google.com.br/books?id=gRn8AAAAQBAJ). 2nd Edition. Human Kinetics.
- [Maths - Euler Angles](http://www.euclideanspace.com/maths/geometry/rotations/euler/).
- Murray RM, Li Z, Sastry SS (1994) [A Mathematical Introduction to Robotic Manipulation](http://www.cds.caltech.edu/~murray/mlswiki/index.php/Main_Page). Boca Raton, CRC Press.
- Ruina A, Rudra P (2013) [Introduction to Statics and Dynamics](http://ruina.tam.cornell.edu/Book/index.html). Oxford University Press.
- Siciliano B, Sciavicco L, Villani L, Oriolo G (2009) [Robotics - Modelling, Planning and Control](http://books.google.com.br/books/about/Robotics.html?hl=pt-BR&id=jPCAFmE-logC). Springer-Verlag London.
- Winter DA (2009) [Biomechanics and motor control of human movement](http://books.google.com.br/books?id=_bFHL08IWfwC). 4 ed. Hoboken, USA: Wiley.
- Zatsiorsky VM (1997) [Kinematics of Human Motion](http://books.google.com.br/books/about/Kinematics_of_Human_Motion.html?id=Pql_xXdbrMcC&redir_esc=y). Champaign, Human Kinetics.
## Function `euler_rotmatrix.py`
```
# %load ./../functions/euler_rotmat.py
#!/usr/bin/env python
"""Euler rotation matrix given sequence, frame, and angles."""
from __future__ import division, print_function
__author__ = 'Marcos Duarte, https://github.com/demotu/BMC'
__version__ = 'euler_rotmat.py v.1 2014/03/10'
def euler_rotmat(order='xyz', frame='local', angles=None, unit='deg',
str_symbols=None, showA=True, showN=True):
"""Euler rotation matrix given sequence, frame, and angles.
This function calculates the algebraic rotation matrix (3x3) for a given
sequence ('order' argument) of up to three elemental rotations of a given
coordinate system ('frame' argument) around another coordinate system, the
Euler (or Eulerian) angles [1]_.
This function also calculates the numerical values of the rotation matrix
when numerical values for the angles are inputed for each rotation axis.
Use None as value if the rotation angle for the particular axis is unknown.
The symbols for the angles are: alpha, beta, and gamma for the first,
second, and third rotations, respectively.
The matrix product is calulated from right to left and in the specified
sequence for the Euler angles. The first letter will be the first rotation.
The function will print and return the algebraic rotation matrix and the
numerical rotation matrix if angles were inputed.
Parameters
----------
order : string, optional (default = 'xyz')
Sequence for the Euler angles, any combination of the letters
x, y, and z with 1 to 3 letters is accepted to denote the
elemental rotations. The first letter will be the first rotation.
frame : string, optional (default = 'local')
Coordinate system for which the rotations are calculated.
Valid values are 'local' or 'global'.
angles : list, array, or bool, optional (default = None)
Numeric values of the rotation angles ordered as the 'order'
parameter. Enter None for a rotation whith unknown value.
unit : str, optional (default = 'deg')
Unit of the input angles.
str_symbols : list of strings, optional (default = None)
New symbols for the angles, for instance, ['theta', 'phi', 'psi']
showA : bool, optional (default = True)
True (1) displays the Algebraic rotation matrix in rich format.
False (0) to not display.
showN : bool, optional (default = True)
True (1) displays the Numeric rotation matrix in rich format.
False (0) to not display.
Returns
-------
R : Matrix Sympy object
Rotation matrix (3x3) in algebraic format.
Rn : Numpy array or Matrix Sympy object (only if angles are inputed)
Numeric rotation matrix (if values for all angles were inputed) or
a algebraic matrix with some of the algebraic angles substituted
by the corresponding inputed numeric values.
Notes
-----
This code uses Sympy, the Python library for symbolic mathematics, to
calculate the algebraic rotation matrix and shows this matrix in latex form
possibly for using with the IPython Notebook, see [1]_.
References
----------
.. [1] http://nbviewer.ipython.org/github/duartexyz/BMC/blob/master/Transformation3D.ipynb
Examples
--------
>>> # import function
>>> from euler_rotmat import euler_rotmat
>>> # Default options: xyz sequence, local frame and show matrix
>>> R = euler_rotmat()
>>> # XYZ sequence (around global (fixed) coordinate system)
>>> R = euler_rotmat(frame='global')
>>> # Enter numeric values for all angles and show both matrices
>>> R, Rn = euler_rotmat(angles=[90, 90, 90])
>>> # show what is returned
>>> euler_rotmat(angles=[90, 90, 90])
>>> # show only the rotation matrix for the elemental rotation at x axis
>>> R = euler_rotmat(order='x')
>>> # zxz sequence and numeric value for only one angle
>>> R, Rn = euler_rotmat(order='zxz', angles=[None, 0, None])
>>> # input values in radians:
>>> import numpy as np
>>> R, Rn = euler_rotmat(order='zxz', angles=[None, np.pi, None], unit='rad')
>>> # shows only the numeric matrix
>>> R, Rn = euler_rotmat(order='zxz', angles=[90, 0, None], showA='False')
>>> # Change the angles' symbols
>>> R = euler_rotmat(order='zxz', str_symbols=['theta', 'phi', 'psi'])
>>> # Negativate the angles' symbols
>>> R = euler_rotmat(order='zxz', str_symbols=['-theta', '-phi', '-psi'])
>>> # all algebraic matrices for all possible sequences for the local frame
>>> s=['xyz','xzy','yzx','yxz','zxy','zyx','xyx','xzx','yzy','yxy','zxz','zyz']
>>> for seq in s: R = euler_rotmat(order=seq)
>>> # all algebraic matrices for all possible sequences for the global frame
>>> for seq in s: R = euler_rotmat(order=seq, frame='global')
"""
import numpy as np
import sympy as sym
try:
from IPython.core.display import Math, display
ipython = True
except:
ipython = False
angles = np.asarray(np.atleast_1d(angles), dtype=np.float64)
if ~np.isnan(angles).all():
if len(order) != angles.size:
raise ValueError("Parameters 'order' and 'angles' (when " +
"different from None) must have the same size.")
x, y, z = sym.symbols('x, y, z')
sig = [1, 1, 1]
if str_symbols is None:
a, b, g = sym.symbols('alpha, beta, gamma')
else:
s = str_symbols
if s[0][0] == '-': s[0] = s[0][1:]; sig[0] = -1
if s[1][0] == '-': s[1] = s[1][1:]; sig[1] = -1
if s[2][0] == '-': s[2] = s[2][1:]; sig[2] = -1
a, b, g = sym.symbols(s)
var = {'x': x, 'y': y, 'z': z, 0: a, 1: b, 2: g}
# Elemental rotation matrices for xyz (local)
cos, sin = sym.cos, sym.sin
Rx = sym.Matrix([[1, 0, 0], [0, cos(x), sin(x)], [0, -sin(x), cos(x)]])
Ry = sym.Matrix([[cos(y), 0, -sin(y)], [0, 1, 0], [sin(y), 0, cos(y)]])
Rz = sym.Matrix([[cos(z), sin(z), 0], [-sin(z), cos(z), 0], [0, 0, 1]])
if frame.lower() == 'global':
Rs = {'x': Rx.T, 'y': Ry.T, 'z': Rz.T}
order = order.upper()
else:
Rs = {'x': Rx, 'y': Ry, 'z': Rz}
order = order.lower()
R = Rn = sym.Matrix(sym.Identity(3))
str1 = r'\mathbf{R}_{%s}( ' %frame # last space needed for order=''
#str2 = [r'\%s'%var[0], r'\%s'%var[1], r'\%s'%var[2]]
str2 = [1, 1, 1]
for i in range(len(order)):
Ri = Rs[order[i].lower()].subs(var[order[i].lower()], sig[i] * var[i])
R = Ri * R
if sig[i] > 0:
str2[i] = '%s:%s' %(order[i], sym.latex(var[i]))
else:
str2[i] = '%s:-%s' %(order[i], sym.latex(var[i]))
str1 = str1 + str2[i] + ','
if ~np.isnan(angles).all() and ~np.isnan(angles[i]):
if unit[:3].lower() == 'deg':
angles[i] = np.deg2rad(angles[i])
Rn = Ri.subs(var[i], angles[i]) * Rn
#Rn = sym.lambdify(var[i], Ri, 'numpy')(angles[i]) * Rn
str2[i] = str2[i] + '=%.0f^o' %np.around(np.rad2deg(angles[i]), 0)
else:
Rn = Ri * Rn
Rn = sym.simplify(Rn) # for trigonometric relations
try:
# nsimplify only works if there are symbols
Rn2 = sym.latex(sym.nsimplify(Rn, tolerance=1e-8).n(chop=True, prec=4))
except:
Rn2 = sym.latex(Rn.n(chop=True, prec=4))
# there are no symbols, pass it as Numpy array
Rn = np.asarray(Rn)
if showA and ipython:
display(Math(str1[:-1] + ') =' + sym.latex(R, mat_str='matrix')))
if showN and ~np.isnan(angles).all() and ipython:
str2 = ',\;'.join(str2[:angles.size])
display(Math(r'\mathbf{R}_{%s}(%s)=%s' %(frame, str2, Rn2)))
if np.isnan(angles).all():
return R
else:
return R, Rn
```
| github_jupyter |
There are two main functions
* decision_function
* predict_proba
Most of classifiers have at least one of them, and many have both
```
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.model_selection import train_test_split
from sklearn.datasets import make_circles
import numpy as np
import matplotlib.pyplot as plt
import mglearn
%matplotlib inline
X, y = make_circles(noise=0.25, factor=0.5, random_state=1)
# we rename the classes "blue" and "red" for illustration purposes
y_named = np.array(["blue", "red"])[y]
# We can call train_test_split with arbitrarily many arrays;
# all will be split in a consistent manner
X_train, X_test, y_train_named, y_test_named, y_train, y_test = train_test_split(X,
y_named,
y,
random_state=0)
# build the gradient boosting model
gbrt = GradientBoostingClassifier(random_state=0)
gbrt.fit(X_train, y_train_named)
```
Uncertaintly in Binary classification case
```
print("Shape of probabilities: ", gbrt.predict_proba(X_test).shape)
# show the first few entries of predict_proba
print("Predicted probabilities:\n", gbrt.predict_proba(X_test[:6]))
fig, axes = plt.subplots(1, 2, figsize=(13, 5))
mglearn.tools.plot_2d_separator(
gbrt, X, ax=axes[0], alpha=.4, fill=True, cm=mglearn.cm2)
scores_image = mglearn.tools.plot_2d_scores(
gbrt, X, ax=axes[1], alpha=.5, cm=mglearn.ReBl, function='predict_proba')
for ax in axes:
# plot training and test points
mglearn.discrete_scatter(X_test[:, 0], X_test[:, 1], y_test,
markers='^', ax=ax)
mglearn.discrete_scatter(X_train[:, 0], X_train[:, 1], y_train,
markers='o', ax=ax)
ax.set_xlabel("Feature 0")
ax.set_ylabel("Feature 1")
# don't want a transparent colorbar
cbar = plt.colorbar(scores_image, ax=axes.tolist())
cbar.set_alpha(1)
cbar.draw_all()
axes[0].legend(["Test class 0", "Test class 1", "Train class 0",
"Train class 1"], ncol=4, loc=(.1, 1.1))
```
Uncertaintly in Multi class classification
```
from sklearn.datasets import load_iris
iris = load_iris()
X_train, X_test, y_train, y_test = train_test_split(iris.data, iris.target, random_state=42)
gbrt = GradientBoostingClassifier(learning_rate=0.01, random_state=0)
gbrt.fit(X_train, y_train)
# show the first few entries of predict_proba
print("Predicted probabilities:\n{}".format(gbrt.predict_proba(X_test)[:6]))
# show that sums across rows are one
print("Sums: {}".format(gbrt.predict_proba(X_test)[:6].sum(axis=1)))
print("Argmax of predicted probabilities:\n{}".format(np.argmax(gbrt.predict_proba(X_test), axis=1)))
print("Predictions:\n{}".format(gbrt.predict(X_test)))
from sklearn.linear_model import LogisticRegression
logreg = LogisticRegression()
# represent each target by its class name in the iris dataset
named_target = iris.target_names[y_train]
logreg.fit(X_train, named_target)
print("unique classes in training data: {}".format(logreg.classes_))
print("predictions: {}".format(logreg.predict(X_test)[:10]))
argmax_dec_func = np.argmax(logreg.decision_function(X_test), axis=1)
print("argmax of decision function: {}".format(argmax_dec_func[:10]))
print("argmax combined with classes_: {}".format(logreg.classes_[argmax_dec_func][:10]))
```
| github_jupyter |
```
from google.colab import drive
drive.mount('/content/drive')
import torch
import torch.nn as nn
class DepthwiseConv2d(nn.Module):
def __init__(self, in_channels, out_channels, kernel_size, stride, padding=1):
super(DepthwiseConv2d, self).__init__()
self.depthwiseconv = nn.Sequential(
nn.Conv2d(in_channels, out_channels, kernel_size, stride=stride, padding=padding),
nn.BatchNorm2d(out_channels),
nn.ReLU(inplace=True),
)
def forward(self, x):
return self.depthwiseconv(x)
class PointwiseConv2d(nn.Module):
def __init__(self, in_channels, out_channels, kernel_size, stride, padding=0):
super(PointwiseConv2d, self).__init__()
self.pointwiseconv = nn.Sequential(
nn.Conv2d(in_channels, out_channels, kernel_size, stride=stride, padding=0),
nn.BatchNorm2d(out_channels),
nn.ReLU(inplace=True),
)
def forward(self, x):
return self.pointwiseconv(x)
class MobileNetv1(nn.Module):
def __init__(self, num_classes=100):
super(MobileNetv1, self).__init__()
self.conv1 = nn.Sequential(
nn.Conv2d(3, 32, kernel_size=3, stride=2, padding=1),
nn.BatchNorm2d(32),
nn.ReLU(inplace=True),
)
self.features = nn.Sequential(
DepthwiseConv2d(32, 32, kernel_size=3, stride=1),
PointwiseConv2d(32, 64, kernel_size=1, stride=1),
DepthwiseConv2d(64, 64, kernel_size=3, stride=2),
PointwiseConv2d(64, 128, kernel_size=1, stride=1),
DepthwiseConv2d(128, 128, kernel_size=3, stride=1),
PointwiseConv2d(128, 128, kernel_size=1, stride=1),
DepthwiseConv2d(128, 128, kernel_size=3, stride=2),
PointwiseConv2d(128, 256, kernel_size=1, stride=1),
DepthwiseConv2d(256, 256, kernel_size=3, stride=1),
PointwiseConv2d(256, 256, kernel_size=1, stride=1),
DepthwiseConv2d(256, 256, kernel_size=3, stride=2),
PointwiseConv2d(256, 512, kernel_size=1, stride=1),
DepthwiseConv2d(512, 512, kernel_size=3, stride=1),
PointwiseConv2d(512, 512, kernel_size=1, stride=1),
DepthwiseConv2d(512, 512, kernel_size=3, stride=1),
PointwiseConv2d(512, 512, kernel_size=1, stride=1),
DepthwiseConv2d(512, 512, kernel_size=3, stride=1),
PointwiseConv2d(512, 512, kernel_size=1, stride=1),
DepthwiseConv2d(512, 512, kernel_size=3, stride=1),
PointwiseConv2d(512, 512, kernel_size=1, stride=1),
DepthwiseConv2d(512, 512, kernel_size=3, stride=1),
PointwiseConv2d(512, 512, kernel_size=1, stride=1),
DepthwiseConv2d(512, 512, kernel_size=3, stride=2),
PointwiseConv2d(512, 1024, kernel_size=1, stride=1),
DepthwiseConv2d(1024, 1024, kernel_size=3, stride=2),
PointwiseConv2d(1024, 1024, kernel_size=1, stride=1),
)
self.avgpool = nn.AvgPool2d(2)
self.fc = nn.Linear(1024, num_classes)
for m in self.modules():
if isinstance(m, nn.Conv2d):
nn.init.kaiming_normal_(m.weight, mode='fan_out', nonlinearity='relu')
if m.bias is not None:
nn.init.constant_(m.bias, 0)
elif isinstance(m, nn.BatchNorm2d):
nn.init.constant_(m.weight, 1)
nn.init.constant_(m.bias, 0)
elif isinstance(m, nn.Linear):
nn.init.normal_(m.weight, 0, 0.01)
nn.init.constant_(m.bias, 0)
def forward(self, x):
x = self.conv1(x)
x = self.features(x)
#x = self.avgpool(x)
x = x.view(x.size(0), -1)
x = self.fc(x)
return x
def mobilenetv1():
return MobileNetv1()
```
config.py
```
import easydict
def config():
cfg = easydict.EasyDict({
"arch": "mobilenetv1_20200415",
"dataset": "cifar100",
"batch_size": 256,
"epochs": 200,
"learning_rate": 0.1,
"weight_decay": 0,
"momentum": 0.9,
"nesterov": True,
"print_freq": 50,
"ckpt": "/content/drive/My Drive/MLVC/Baseline/checkpoint/mobilnetv1_20200415/",
"results_dir": "./results/",
"resume": False,
"evaluate": False,
"cuda": True,
"gpuids": [0],
"colab": True,
})
cfg.gpuids = list(map(int, cfg.gpuids))
model = mobilenetv1()
if cfg.arch == "mobilenetv1_20200415":
model = mobilenetv1()
#elif cfg.arch == "resnet-cifar":
# model = resnet.resnet20()
#elif cfg.arch == "vgg-cifar-binary":
# model = vgg_bnn.vgg11()
#elif cfg.arch == "resnet-cifar-dorefa":
# model = resnet_dorefanet.resnet20()
return cfg, model
```
utility.py
```
import torch
import time
import shutil
import pathlib
from collections import OrderedDict
def load_model(model, ckpt_file, args):
if args.cuda:
checkpoint = torch.load(
ckpt_file, map_location=lambda storage, loc: storage.cuda(args.gpuids[0])
)
try:
model.load_state_dict(checkpoint["model"])
except: # noqa
model.module.load_state_dict(checkpoint["model"])
else:
checkpoint = torch.load(ckpt_file, map_location=lambda storage, loc: storage)
try:
model.load_state_dict(checkpoint["model"])
except: # noqa
# create new OrderedDict that does not contain `module.`
new_state_dict = OrderedDict()
for k, v in checkpoint["model"].items():
if k[:7] == "module.":
name = k[7:] # remove `module.`
else:
name = k[:]
new_state_dict[name] = v
model.load_state_dict(new_state_dict)
return checkpoint
def save_model(state, epoch, is_best, args):
dir_ckpt = pathlib.Path("/content/drive/My Drive/MLVC/Baseline/checkpoint/mobilnetv1_20200415/")
dir_path = dir_ckpt / args.dataset
dir_path.mkdir(parents=True, exist_ok=True)
model_file = dir_path / "ckpt_epoch_{}.pth".format(epoch)
torch.save(state, model_file)
if is_best:
shutil.copyfile(model_file, dir_path / "ckpt_best.pth")
class AverageMeter(object):
"""Computes and stores the average and current value"""
def __init__(self, name, fmt=":f"):
self.name = name
self.fmt = fmt
self.reset()
def reset(self):
self.val = 0
self.avg = 0
self.sum = 0
self.count = 0
def update(self, val, n=1):
self.val = val
self.sum += val * n
self.count += n
self.avg = self.sum / self.count
def __str__(self):
fmtstr = "{name} {val" + self.fmt + "} ({avg" + self.fmt + "})"
return fmtstr.format(**self.__dict__)
class ProgressMeter(object):
def __init__(self, num_batches, *meters, prefix=""):
self.batch_fmtstr = self._get_batch_fmtstr(num_batches)
self.meters = meters
self.prefix = prefix
def print(self, batch):
entries = [self.prefix + self.batch_fmtstr.format(batch)]
entries += [str(meter) for meter in self.meters]
print("\t".join(entries))
def _get_batch_fmtstr(self, num_batches):
num_digits = len(str(num_batches // 1))
fmt = "{:" + str(num_digits) + "d}"
return "[" + fmt + "/" + fmt.format(num_batches) + "]"
def adjust_learning_rate(optimizer, epoch, lr):
"""Sets the learning rate, decayed rate of 0.1 every epoch"""
#if epoch >= 60:
# lr = 0.01
#if epoch >= 120:
# lr = 0.001
#if epoch >= 160:
# lr = 0.0001
for param_group in optimizer.param_groups:
param_group["lr"] = lr
return lr
def accuracy(output, target, topk=(1,)):
"""Computes the accuracy over the k top predictions for the specified values of k"""
with torch.no_grad():
maxk = max(topk)
batch_size = target.size(0)
_, pred = output.topk(maxk, 1, True, True)
pred = pred.t()
correct = pred.eq(target.view(1, -1).expand_as(pred))
res = []
for k in topk:
correct_k = correct[:k].view(-1).float().sum(0, keepdim=True)
res.append(correct_k.mul_(100.0 / batch_size))
return res
def print_reults(start_time, train_time, validate_time, start_epoch, epochs):
avg_train_time = train_time / (epochs - start_epoch)
avg_valid_time = validate_time / (epochs - start_epoch)
total_train_time = train_time + validate_time
print(
"====> average training time per epoch: {:,}m {:.2f}s".format(
int(avg_train_time // 60), avg_train_time % 60
)
)
print(
"====> average validation time per epoch: {:,}m {:.2f}s".format(
int(avg_valid_time // 60), avg_valid_time % 60
)
)
print(
"====> training time: {}h {}m {:.2f}s".format(
int(train_time // 3600), int((train_time % 3600) // 60), train_time % 60
)
)
print(
"====> validation time: {}h {}m {:.2f}s".format(
int(validate_time // 3600),
int((validate_time % 3600) // 60),
validate_time % 60,
)
)
print(
"====> total training time: {}h {}m {:.2f}s".format(
int(total_train_time // 3600),
int((total_train_time % 3600) // 60),
total_train_time % 60,
)
)
elapsed_time = time.time() - start_time
print(
"====> total time: {}h {}m {:.2f}s".format(
int(elapsed_time // 3600), int((elapsed_time % 3600) // 60), elapsed_time % 60
)
)
```
data_loader.py
```
import torch
import torchvision.transforms as transforms
from torchvision import datasets
def dataloader(dataset, batch_size):
train_dataset, val_dataset = load_cifar10()
if dataset == "cifar100":
train_dataset, val_dataset = load_cifar100()
# Data loader
train_loader = torch.utils.data.DataLoader(
dataset=train_dataset, batch_size=batch_size, shuffle=True
)
val_loader = torch.utils.data.DataLoader(
dataset=val_dataset, batch_size=batch_size, shuffle=False
)
return train_loader, val_loader
def load_cifar10():
# CIFAR-10 dataset
normalize = transforms.Normalize(
mean=[0.485, 0.456, 0.406], std=[0.229, 0.224, 0.225]
)
train_dataset = datasets.CIFAR10(
root="../../data/",
train=True,
transform=transforms.Compose(
[
transforms.Pad(4),
transforms.RandomHorizontalFlip(),
transforms.RandomCrop(32),
transforms.ToTensor(),
normalize,
]
),
download=True,
)
val_dataset = datasets.CIFAR10(
root="../../data/",
train=False,
transform=transforms.Compose([transforms.ToTensor(), normalize]),
)
return train_dataset, val_dataset
def load_cifar100():
# CIFAR-100 dataset
normalize = transforms.Normalize(
mean=[0.4914, 0.4822, 0.4465], std=[0.2023, 0.1994, 0.2010]
)
train_dataset = datasets.CIFAR100(
root="../../data/",
train=True,
transform=transforms.Compose(
[
transforms.RandomCrop(32),
transforms.Pad(4),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
normalize,
]
),
download=True,
)
val_dataset = datasets.CIFAR100(
root="../../data/",
train=False,
transform=transforms.Compose([transforms.ToTensor(), normalize]),
)
return train_dataset, val_dataset
```
main.py
```
import torch
import torch.nn as nn
import torch.optim as optim
import torch.backends.cudnn as cudnn
import time
import pathlib
from os.path import isfile
import pandas as pd
def main():
global args, start_epoch, best_acc1
args, model = config()
print("Model: {}".format(args.arch))
if args.cuda and not torch.cuda.is_available():
raise Exception("No GPU found, please run without --cuda")
criterion = nn.CrossEntropyLoss()
optimizer = optim.SGD(
model.parameters(),
lr=args.learning_rate,
weight_decay=args.weight_decay,
momentum=args.momentum,
nesterov=args.nesterov,
)
best_acc1 = 0
start_epoch = 0
if args.cuda:
torch.cuda.set_device(args.gpuids[0])
with torch.cuda.device(args.gpuids[0]):
model = model.cuda()
criterion = criterion.cuda()
model = nn.DataParallel(
model, device_ids=args.gpuids, output_device=args.gpuids[0]
)
cudnn.benchmark = True
# checkpoint file
ckpt_dir = pathlib.Path(args.ckpt)
ckpt_file = ckpt_dir / args.dataset / args.ckpt
# for resuming training
if args.resume:
retrain(ckpt_file, model, optimizer)
# Data loading
print("\n==> Load data..")
train_loader, val_loader = dataloader(args.dataset, args.batch_size)
# initiailizae
train_time, validate_time = 0.0, 0.0
avgloss_train = 0.0
acc1_train, acc5_train, acc1_valid, acc5_valid = 0.0, 0.0, 0.0, 0.0
is_best = False
# result lists
result_epoch, result_lr, result_train_avgtime, result_train_avgloss = [], [], [], []
result_train_avgtop1acc, result_train_avgtop5acc = [], []
result_val_avgtime, result_val_avgtop1acc, result_val_avgtop5acc = [], [], []
# train...
lr = args.learning_rate
curr_lr = lr
for epoch in range(start_epoch, args.epochs):
curr_lr = adjust_learning_rate(optimizer, epoch, lr)
print("\n==> Epoch: {}, lr = {}".format(epoch, optimizer.param_groups[0]["lr"]))
# train for one epoch
train_time, acc1_train, acc5_train, avgloss_train = train_epoch(
train_time,
acc1_train,
acc5_train,
avgloss_train,
train_loader,
epoch,
model,
criterion,
optimizer,
)
# evaluate on validation set
validate_time, acc1_valid, acc5_valid = validation_epoch(
validate_time, acc1_valid, acc5_valid, val_loader, model, criterion
)
# remember best Acc@1 and save checkpoint
is_best = save_model_data(
is_best, best_acc1, acc1_valid, epoch, model, optimizer, args
)
result_epoch.append(epoch)
result_lr.append(curr_lr)
result_train_avgtime.append(train_time)
result_train_avgloss.append(avgloss_train)
result_train_avgtop1acc.append(acc1_train.item())
result_train_avgtop5acc.append(acc5_train.item())
result_val_avgtop1acc.append(acc1_valid.item())
result_val_avgtop5acc.append(acc5_valid.item())
df = pd.DataFrame({
'Epoch': result_epoch,
'Learning rate': result_lr,
'Training avg loss': result_train_avgloss,
'Training avg top1 acc': result_train_avgtop1acc,
'Training avg top5 acc': result_train_avgtop5acc,
'Test avg top1 acc': result_val_avgtop1acc,
'Test avg top5 acc': result_val_avgtop5acc,
})
if args.colab:
df.to_csv('/content/drive/My Drive/MLVC/Baseline/results/{}_result.csv'.format(args.arch))
else:
df.to_csv('./results/{}_result.csv'.format(args.arch))
print_results(train_time, validate_time)
def retrain(ckpt_file, model, optimizer):
if isfile(ckpt_file):
print("\n==> Loading Checkpoint '{}'".format(args.ckpt))
checkpoint = load_model(model, ckpt_file, args)
start_epoch = checkpoint["epoch"]
optimizer.load_state_dict(checkpoint["optimizer"])
print("==> Loaded Checkpoint '{}' (epoch {})".format(args.ckpt, start_epoch))
else:
print("==> no checkpoint found '{}'".format(args.ckpt))
return
def train_epoch(
train_time, acc1_train, acc5_train, avgloss_train, train_loader, epoch, model, criterion, optimizer
):
print("===> [ Training ]")
start_time = time.time()
acc1_train, acc5_train, avgloss_train = train(
train_loader, epoch=epoch, model=model, criterion=criterion, optimizer=optimizer
)
elapsed_time = time.time() - start_time
train_time += elapsed_time
print("====> {:.2f} seconds to train this epoch\n".format(elapsed_time))
return train_time, acc1_train, acc5_train, avgloss_train
def validation_epoch(
validate_time, acc1_valid, acc5_valid, val_loader, model, criterion
):
print("===> [ Validation ]")
start_time = time.time()
acc1_valid, acc5_valid, avgloss_valid = validate(val_loader, model, criterion)
elapsed_time = time.time() - start_time
validate_time += elapsed_time
print("====> {:.2f} seconds to validate this epoch\n".format(elapsed_time))
return validate_time, acc1_valid, acc5_valid
def save_model_data(is_best, best_acc1, acc1_valid, epoch, model, optimizer, args):
is_best = acc1_valid > best_acc1
best_acc1 = max(acc1_valid, best_acc1)
state = {
"epoch": epoch + 1,
"model": model.state_dict(),
"optimizer": optimizer.state_dict(),
}
if (epoch + 1) % 20 == 0:
save_model(state, epoch, is_best, args)
return is_best
def train(train_loader, **kwargs):
epoch = kwargs.get("epoch")
model = kwargs.get("model")
criterion = kwargs.get("criterion")
optimizer = kwargs.get("optimizer")
batch_time = AverageMeter("Time", ":6.3f")
data_time = AverageMeter("Data", ":6.3f")
losses = AverageMeter("Loss", ":.4e")
top1 = AverageMeter("Acc@1", ":6.2f")
top5 = AverageMeter("Acc@5", ":6.2f")
progress = ProgressMeter(
len(train_loader),
batch_time,
data_time,
losses,
top1,
top5,
prefix="Epoch: [{}]".format(epoch),
)
# switch to train mode
model.train()
end = time.time()
running_loss = 0.0
for i, (input, target) in enumerate(train_loader):
# measure data loading time
data_time.update(time.time() - end)
if args.cuda:
input = input.cuda(non_blocking=True)
target = target.cuda(non_blocking=True)
# compute output
output = model(input)
loss = criterion(output, target)
# measure accuracy and record loss
acc1, acc5 = accuracy(output, target, topk=(1, 5))
losses.update(loss.item(), input.size(0))
top1.update(acc1[0], input.size(0))
top5.update(acc5[0], input.size(0))
# compute gradient and do SGD step.
optimizer.zero_grad()
loss.backward()
optimizer.step()
# measure elapsed time
batch_time.update(time.time() - end)
running_loss += loss.item()
if i % args.print_freq == 0:
progress.print(i)
end = time.time()
print(
"====> Acc@1 {top1.avg:.3f} Acc@5 {top5.avg:.3f}".format(top1=top1, top5=top5)
)
epoch_loss = running_loss / len(train_loader)
print("====> Epoch loss {:.3f}".format(epoch_loss))
return top1.avg, top5.avg, epoch_loss
def validate(val_loader, model, criterion):
batch_time = AverageMeter("Time", ":6.3f")
losses = AverageMeter("Loss", ":.4e")
top1 = AverageMeter("Acc@1", ":6.2f")
top5 = AverageMeter("Acc@5", ":6.2f")
progress = ProgressMeter(
len(val_loader), batch_time, losses, top1, top5, prefix="Test: "
)
# switch to evaluate mode
model.eval()
total_loss = 0.0
with torch.no_grad():
end = time.time()
for i, (input, target) in enumerate(val_loader):
if args.cuda:
input = input.cuda(non_blocking=True)
target = target.cuda(non_blocking=True)
# compute output
output = model(input)
loss = criterion(output, target)
# measure accuracy and record loss
acc1, acc5 = accuracy(output, target, topk=(1, 5))
losses.update(loss.item(), input.size(0))
top1.update(acc1[0], input.size(0))
top5.update(acc5[0], input.size(0))
# measure elapsed time
batch_time.update(time.time() - end)
total_loss += loss.item()
if i % args.print_freq == 0:
progress.print(i)
end = time.time()
print(
"====> Acc@1 {top1.avg:.3f} Acc@5 {top5.avg:.3f}".format(
top1=top1, top5=top5
)
)
total_loss = total_loss / len(val_loader)
return top1.avg, top5.avg, loss.item()
def print_results(train_time, validate_time):
avg_train_time = train_time / (args.epochs - start_epoch)
avg_valid_time = validate_time / (args.epochs - start_epoch)
total_train_time = train_time + validate_time
print(
"====> average training time per epoch: {:,}m {:.2f}s".format(
int(avg_train_time // 60), avg_train_time % 60
)
)
print(
"====> average validation time per epoch: {:,}m {:.2f}s".format(
int(avg_valid_time // 60), avg_valid_time % 60
)
)
print(
"====> training time: {}h {}m {:.2f}s".format(
int(train_time // 3600), int((train_time % 3600) // 60), train_time % 60
)
)
print(
"====> validation time: {}h {}m {:.2f}s".format(
int(validate_time // 3600),
int((validate_time % 3600) // 60),
validate_time % 60,
)
)
print(
"====> total training time: {}h {}m {:.2f}s".format(
int(total_train_time // 3600),
int((total_train_time % 3600) // 60),
total_train_time % 60,
)
)
if __name__ == "__main__":
start_time = time.time()
main()
elapsed_time = time.time() - start_time
print(
"====> total time: {}h {}m {:.2f}s".format(
int(elapsed_time // 3600),
int((elapsed_time % 3600) // 60),
elapsed_time % 60,
)
)
```
| github_jupyter |
```
import os
import csv
import sys
import scipy.optimize as opt
import scipy.stats as stat
from operator import itemgetter
import random
import numpy as np
import numpy.ma as ma
import numpy.linalg as la
pi = np.pi
sin = np.sin
cos = np.cos
def fillin2(data):
"""
Fills in blanks of arrays without shifting frames by the starting frame. Compare to fillin.
Input: trajectory dataset from MOSAIC tracking software read into a numpy array
Output: modified numpy array with missing frames filled in.
"""
shap = int(max(data[:, 1])) + 1
shape1 = int(min(data[:, 1]))
newshap = shap - shape1
filledin = np.zeros((newshap, 5))
filledin[0, :] = data[0, :]
count = 0
new = 0
other = 0
tot = 0
for num in range(1, newshap):
if data[num-new, 1]-data[num-new-1, 1] == 1:
count = count + 1
filledin[num, :] = data[num-new, :]
elif data[num - new, 1]-data[num - new - 1, 1] > 1:
new = new + 1
iba = int(data[num - new+1, 1]-data[num - new, 1])
togoin = data[num - new]
togoin[1] = togoin[1] + 1
filledin[num, :] = togoin
# dataset[2] = np.insert(dataset[2], [num + new - 2], togoin, axis=0)
else:
other = other + 1
tot = count + new + other
return filledin
def MSD_iteration(folder, name, cut, totvids, conversion, frames):
"""
Cleans up data for MSD analysis from csv files. Outputs in form of
dictionaries.
"""
trajectory = dict()
tots = dict() # Total particles in each video
newtots = dict() # Cumulative total particles.
newtots[0] = 0
tlen = dict()
tlength = dict()
tlength[0] = 0
for num in range(1, totvids + 1):
trajectory[num] = np.genfromtxt(folder+'Traj_{}_{}.tif.csv'.format(name, num), delimiter=",")
trajectory[num] = np.delete(trajectory[num], 0, 1)
tots[num] = trajectory[num][-1, 0].astype(np.int64)
newtots[num] = newtots[num-1] + tots[num]
tlen[num] = trajectory[num].shape[0]
tlength[num] = tlength[num-1] + tlen[num]
placeholder = np.zeros((tlength[totvids], 11))
for num in range(1, totvids + 1):
placeholder[tlength[num-1]:tlength[num], :] = trajectory[num]
placeholder[tlength[num-1]:tlength[num], 0] = placeholder[tlength[num-1]:tlength[num], 0] + newtots[num-1]
dataset = dict()
rawdataset = np.zeros(placeholder.shape)
particles = placeholder[:, 0]
total = int(max(particles))
total1 = total + 1
rawdataset = placeholder[:, :]
fixed = np.zeros(placeholder.shape)
fixed[:, 0:2] = rawdataset[:, 0:2]
fixed[:, 2:4] = conversion[0] * rawdataset[:, 2:4]
fixed[:, 4] = conversion[2] * rawdataset[:, 4]
x = np.zeros((frames, total1 - 1))
y = np.zeros((frames, total1 - 1))
xs = np.zeros((frames, total1 - 1))
ys = np.zeros((frames, total1 - 1))
nones = 0
cutoff = cut
for num in range(1, total1):
hold = np.where(particles == num)
itindex = hold[0]
min1 = min(itindex)
max1 = max(itindex)
if max1 - min1 < cutoff:
nones = nones + 1
else:
holdplease = fillin2(fixed[min1:max1+1, 0:5])
x[int(holdplease[0, 1]):int(holdplease[-1, 1])+1, num - nones - 1] = holdplease[:, 2]
y[int(holdplease[0, 1]):int(holdplease[-1, 1])+1, num - nones - 1] = holdplease[:, 3]
xs[0:int(holdplease[-1, 1])+1-int(holdplease[0, 1]), num - nones - 1] = holdplease[:, 2]
ys[0:int(holdplease[-1, 1])+1-int(holdplease[0, 1]), num - nones - 1] = holdplease[:, 3]
total1 = total1 - nones - 1
x_m = x[:, :total1-1]
y_m = y[:, :total1-1]
xs_m = xs[:, :total1-1]
ys_m = ys[:, :total1-1]
return total1, xs_m, ys_m, x_m, y_m
def vectorized_MMSD_calcs(frames, total1, xs_m, ys_m, x_m, y_m, frame_m):
SM1x = np.zeros((frames, total1-1))
SM1y = np.zeros((frames, total1-1))
SM2xy = np.zeros((frames, total1-1))
xs_m = ma.masked_equal(xs_m, 0)
ys_m = ma.masked_equal(ys_m, 0)
x_m = ma.masked_equal(x_m, 0)
y_m = ma.masked_equal(y_m, 0)
geoM1x = np.zeros(frame_m)
geoM1y = np.zeros(frame_m)
for frame in range(1, frame_m):
bx = xs_m[frame, :]
cx = xs_m[:-frame, :]
Mx = (bx - cx)**2
Mxa = np.mean(Mx, axis=0)
# Mxab = np.mean(np.log(Mxa), axis=0)
# geoM1x[frame] = Mxab
by = ys_m[frame, :]
cy = ys_m[:-frame, :]
My = (by - cy)**2
Mya = np.mean(My, axis=0)
# Myab = np.mean(np.log(Mya), axis=0)
# geoM1y[frame] = Myab
SM1x[frame, :] = Mxa
SM1y[frame, :] = Mya
geoM2xy = np.mean(ma.log(SM1x+SM1y), axis=1)
gSEM = stat.sem(ma.log(SM1x+SM1y), axis=1)
SM2xy = SM1x + SM1y
return geoM2xy, gSEM, SM1x, SM1y, SM2xy
folder = "./"
path = "./geoM2xy_{sample_name}.csv"
frames = 651
SD_frames = [10, 20, 50, 80]
conversion = (0.16, 100.02, 1) # (0.3, 3.95, 1)
to_frame = 120
dimension = "2D"
time_to_calculate = 1
base = "in_agarose"
base_name = "RED"
test_bins = np.linspace(0, 75, 76)
# name = 'RED_KO_PEG_P1_S1_cortex'
cut = 1
totvids = 2
frame_m = 651 # atm I can't go lower than the actual value.
parameters = {}
parameters["channels"] = ["RED"]
parameters["surface functionalities"] = ["nPEG"]
parameters["slices"] = ["S1", "S2"]
parameters["videos"] = [1, 2]
parameters["replicates"] = [1, 2]
channels = parameters["channels"]
surface_functionalities = parameters["surface functionalities"]
slices = parameters["slices"]
videos = parameters["videos"]
replicates = parameters["replicates"]
geoM2xy = {}
gSEM = {}
SM1x = {}
SM1y = {}
SM2xy = {}
for channel in channels:
for surface_functionality in surface_functionalities:
slice_counter = 0
for slic in slices:
for video in videos:
sample_name = "{}_{}_{}_{}_{}".format(channel, surface_functionality, '37C_pH72', slic, video)
DIR = folder
total1, xs, ys, x, y = MSD_iteration(DIR, sample_name, cut, totvids, conversion, frame_m)
geoM2xy[sample_name], gSEM[sample_name], SM1x[sample_name], SM1y[sample_name],\
SM2xy[sample_name] = vectorized_MMSD_calcs(frames, total1, xs, ys, x, y, frame_m)
np.savetxt(DIR+'geoM2xy_{}.csv'.format(sample_name), geoM2xy[sample_name], delimiter=',')
np.savetxt(DIR+'gSEM_{}.csv'.format(sample_name), gSEM[sample_name], delimiter=',')
np.savetxt(DIR+'SM2xy_{}.csv'.format(sample_name), SM2xy[sample_name], delimiter=',')
slice_counter = slice_counter + 1
SM2xy1 = SM2xy['RED_nPEG_37C_pH72_S1_1']
geoM2xy1 = np.mean(ma.log(SM2xy1+SM2xy1), axis=1)
geoM2xy1 = stat.sem(ma.log(SM2xy1+SM2xy1), axis=1)
geoM2xy
geoM2xy[sample_name]
```
| github_jupyter |
```
import pandas as pd
import numpy as np
data=pd.read_csv("/home/jay/Desktop/Cricket/Final/FinalTrainingDataset.csv")
data
X=data.iloc[:,1:14].values
y=data.iloc[:,14].values
y
X.shape
from sklearn.model_selection import train_test_split
X_train,X_test,y_train,y_test=train_test_split(X,y,test_size=0.20,random_state=0)
```
# Multiple Linear Regression
```
from sklearn.linear_model import LinearRegression
Lregressor=LinearRegression()
Lregressor.fit(X_train,y_train)
score = Lregressor.score(X_train, y_train)
score2 = Lregressor.score(X_test, y_test)
print("Training set accuracy: ", '%.3f'%(score))
print("Test set accuracy: ", '%.3f'%(score2))
```
# Random Forest Regression
```
from sklearn.ensemble import RandomForestClassifier
rf = RandomForestClassifier(n_estimators=100,random_state=0)
rf.fit(X_train, y_train)
score = rf.score(X_train, y_train)
score2 = rf.score(X_test, y_test)
print("Training set accuracy: ", '%.3f'%(score))
print("Test set accuracy: ", '%.3f'%(score2))
```
# As the Accuracy is coming better in Multiple Linear regression is better so we are using that model to predict auction_values
```
y_pred=Lregressor.predict(X_test)
y_pred
y_test
feature_cols=['Matches','Runs','HS','Bat_Avg','Bowl_Avg','Avg_Diff','Wickets','Centuries','Fifers','Catches','Bat_Str','Bowl_Eco','Ind_Abroad']
X_trainPred=data[feature_cols]
y_TrainPred=Lregressor.predict(X_trainPred)
y_TrainPred
dd=pd.DataFrame({'Predicted_Price':y_TrainPred})
DataTrainResult=pd.concat([data,dd],axis=1)
DataTrainResult
# Here we can see the Actual Price and the Predicted Price
Vis=DataTrainResult.iloc[:15,[0,14,15]]
%matplotlib inline
plot1= Vis.plot.bar(x='Player',y={'Price','Predicted_Price'},title='Actual Price .vs. Predicted Price')
```
# Now we'll apply the model created on the Predict Dataset
```
predict=pd.read_csv("/home/jay/Desktop/Cricket/Final/Predict.csv")
predict
predict=predict.rename(columns={'Ind/Abr':'Ind_Abroad','Bat_str':'Bat_Str'})
X_finalPred=predict[feature_cols].values
y_Finalpred=Lregressor.predict(X_finalPred)
y_Finalpred
dd=pd.DataFrame({'Predicted':y_Finalpred})
predict=pd.concat([predict,dd],axis=1)
predict.nlargest(5, ['Predicted'])
X.shape
```
# Feature Selection using Backward Elimination and Adj. R Sq value
```
import statsmodels.api as sm
import statsmodels.formula.api as smf
X=np.append(arr = np.ones((50,1)).astype(int) , values = X ,axis=1)
X.shape
X_opt = X[:, [0,1,2,3,4,5,6,7,8,9,10,11,12,13]]
regressor_ols=sm.OLS(endog=y,exog=X_opt).fit()
regressor_ols.summary()
X_opt = X[:, [0,1,2,3,6,7,8,9,10,11,12,13]]
regressor_ols=sm.OLS(endog=y,exog=X_opt).fit()
regressor_ols.summary()
X_opt = X[:, [0,1,2,3,6,7,8,9,10,11,13]]
regressor_ols=sm.OLS(endog=y,exog=X_opt).fit()
regressor_ols.summary()
X_opt = X[:, [0,1,2,3,6,7,9,10,11,13]]
regressor_ols=sm.OLS(endog=y,exog=X_opt).fit()
regressor_ols.summary()
X_opt = X[:, [0,1,2,6,7,9,10,11,13]]
regressor_ols=sm.OLS(endog=y,exog=X_opt).fit()
regressor_ols.summary()
X_opt = X[:, [0,1,2,7,9,10,11,13]]
regressor_ols=sm.OLS(endog=y,exog=X_opt).fit()
regressor_ols.summary()
X_opt = X[:, [0,2,7,9,10,11,13]]
regressor_ols=sm.OLS(endog=y,exog=X_opt).fit()
regressor_ols.summary()
X_opt2 = X[:, [0,2,7,10,11,13]]
regressor_ols=sm.OLS(endog=y,exog=X_opt2).fit()
regressor_ols.summary()
```
# As in above summary when we remove x3, Adj.R Sq value goes from 0.811 to 0.803 so we shouldn't remove x3
```
X_opt = X[:, [0,2,7,9,10,11,13]]
```
```
FeaturedData= data.iloc[:,[0,2,7,9,10,11,13,14]]
FeaturedData
X=FeaturedData.iloc[:,1:7].values
y=FeaturedData.iloc[:,7].values
from sklearn.model_selection import train_test_split
X_train,X_test,y_train,y_test=train_test_split(X,y,test_size=0.20,random_state=0)
from sklearn.linear_model import LinearRegression
Lregressor=LinearRegression()
Lregressor.fit(X_train,y_train)
score = Lregressor.score(X_train, y_train)
score2 = Lregressor.score(X_test, y_test)
print("Training set accuracy: ", '%.3f'%(score))
print("Test set accuracy: ", '%.3f'%(score2))
from sklearn.ensemble import RandomForestClassifier
rf = RandomForestClassifier(n_estimators=5,random_state=0)
rf.fit(X_train, y_train)
score = rf.score(X_train, y_train)
score2 = rf.score(X_test, y_test)
print("Training set accuracy: ", '%.3f'%(score))
print("Test set accuracy: ", '%.3f'%(score2))
predict=pd.read_csv("/home/jay/Desktop/Cricket/Final/Predict.csv")
predict
predict=predict.rename(columns={'Ind/Abr':'Ind_Abroad','Bat_str':'Bat_Str'})
FeaturedPredict=predict.iloc[:,[0,2,7,9,10,11,13]]
FeatureSelection_cols=['Runs','Wickets','Fifers','Catches','Bat_Str','Ind_Abroad']
X_Fpred=FeaturedPredict[FeatureSelection_cols].values
y_LinearPred=Lregressor.predict(X_Fpred)
y_LinearPred
y_RFPred=rf.predict(X_Fpred)
y_RFPred
ddlj=pd.DataFrame({'LinearRegression_Price':y_LinearPred,'RandomForest_Price':y_RFPred})
FeaturedPredict=pd.concat([FeaturedPredict,ddlj],axis=1)
FeaturedPredict.head()
Vis_FP=FeaturedPredict.iloc[:,[0,7,8]]
%matplotlib inline
plot1= Vis_FP.plot.line(x='Player',y={'LinearRegression_Price','RandomForest_Price'},title='LinearRegression .vs. RandomForest')
```
# Decision Tree
```
from sklearn.tree import DecisionTreeClassifier
dtc=DecisionTreeClassifier(criterion='entropy',max_depth=None,splitter='best')
dtc=dtc.fit(X_train,y_train)
from sklearn.externals.six import StringIO
from IPython.display import Image
from sklearn.tree import export_graphviz
import pydotplus as pdp
dot_data=StringIO()
export_graphviz(dtc,out_file=dot_data,filled=True,rounded=True,special_characters=True,feature_names=FeatureSelection_cols)
graph=pdp.graph_from_dot_data(dot_data.getvalue())
graph.write_png('AuctionDecisionTree.png')
```
# K-Means Clustering
```
Data1=FeaturedPredict.iloc[:,1:8]
Data2=FeaturedData.iloc[:,1:]
Data1.to_csv('Data1.csv')
Data2.to_csv('Data2.csv')
DataTotalFinal=pd.read_csv('/home/jay/DataTotalFinal.csv')
DataTotalFinal
DTF=DataTotalFinal.iloc[:,:-1]
```
# PCA
```
from sklearn.decomposition import PCA
pca=PCA(.95)
pca.fit(DTF)
DTF.shape
DF=pca.fit_transform(DTF)
DF.shape
DF
z=DataTotalFinal.iloc[:,[6]].values
x=np.concatenate((DF,z),axis=1)
import matplotlib.pyplot as plt
from sklearn.cluster import KMeans
wcss=[]
for i in range(1,11):
kmeans=KMeans(n_clusters=i,init='k-means++',max_iter=300,n_init=10,random_state=0)
kmeans.fit(x)
wcss.append(kmeans.inertia_)
plt.plot(range(1, 11), wcss)
plt.title('The Elbow Method')
plt.show()
kmeans=KMeans(n_clusters=4,init='k-means++',max_iter=300,n_init=10,random_state=0)
y_kmeans=kmeans.fit_predict(x)
y_kmeans
plt.figure(figsize=(15, 8))
plt.scatter(x[y_kmeans==3, 0],x[y_kmeans==3, 1],s=100,c='yellow',label='Superstars')
plt.scatter(x[y_kmeans==1, 0],x[y_kmeans==1, 1],s=100,c='blue',label='Match-Winners')
plt.scatter(x[y_kmeans==0, 0],x[y_kmeans==0, 1],s=100,c='red',label='Average-Players')
plt.scatter(x[y_kmeans==2, 0],x[y_kmeans==2, 1],s=100,c='green',label='Good-Players')
plt.scatter(kmeans.cluster_centers_[:, 0],kmeans.cluster_centers_[:,1],s=200,c='black',label='Centroids')
plt.title('Division of Players')
plt.legend()
plt.show()
```
| github_jupyter |
# 作業 : (Kaggle)鐵達尼生存預測
***
- 分數以網站評分結果為準, 請同學實際將提交檔(*.csv)上傳試試看
https://www.kaggle.com/c/titanic/submit
# [作業目標]
- 試著模仿範例寫法, 在鐵達尼生存預測中, 觀查堆疊泛化 (Stacking) 的寫法與效果
# [作業重點]
- 完成堆疊泛化的寫作, 看看提交結果, 想想看 : 分類與回歸的堆疊泛化, 是不是也與混合泛化一樣有所不同呢?(In[14])
如果可能不同, 應該怎麼改寫會有較好的結果?
- Hint : 請參考 mlxtrend 官方網站 StackingClassifier 的頁面說明 : Using Probabilities as Meta-Features
http://rasbt.github.io/mlxtend/user_guide/classifier/StackingClassifier/
```
# 做完特徵工程前的所有準備 (與前範例相同)
import pandas as pd
import numpy as np
import copy, time
import warnings
warnings.filterwarnings('ignore')
from IPython.display import display
from sklearn.preprocessing import MinMaxScaler
from sklearn.model_selection import cross_val_score
from sklearn.linear_model import LogisticRegression
from sklearn.preprocessing import LabelEncoder
data_path = 'data/'
df_train = pd.read_csv(data_path + 'titanic_train.csv')
df_test = pd.read_csv(data_path + 'titanic_test.csv')
train_Y = df_train['Survived']
ids = df_test['PassengerId']
df_train = df_train.drop(['PassengerId', 'Survived'] , axis=1)
df_test = df_test.drop(['PassengerId'] , axis=1)
df = pd.concat([df_train,df_test])
df.head()
# 檢查 DataFrame 空缺值的狀態
def na_check(df_data):
data_na = (df_data.isnull().sum() / len(df_data)) * 100
data_na = data_na.drop(data_na[data_na == 0].index).sort_values(ascending=False)
missing_data = pd.DataFrame({'Missing Ratio' :data_na})
display(missing_data.head(10))
na_check(df)
# 以下 In[3]~In[10] 只是鐵達尼預測中的一組特徵工程, 並以此組特徵工程跑參數, 若更換其他特徵工程, In[10]的參數需要重新跑
# Sex : 直接轉男 0 女 1
df["Sex"] = df["Sex"].map({"male": 0, "female":1})
# Fare : 用 log 去偏態, 0 則直接取 0
df["Fare"] = df["Fare"].map(lambda i: np.log(i) if i > 0 else 0)
# Age : 缺值用中位數補
df["Age"] = df["Age"].fillna(df['Age'].median())
# Title 的 特徵工程 : 將各種頭銜按照類型分類, 最後取 One Hot
df_title = [i.split(",")[1].split(".")[0].strip() for i in df["Name"]]
df["Title"] = pd.Series(df_title)
df["Title"] = df["Title"].replace(['Lady', 'the Countess','Countess','Capt', 'Col','Don', 'Dr', 'Major', 'Rev', 'Sir', 'Jonkheer', 'Dona'], 'Rare')
df["Title"] = df["Title"].map({"Master":0, "Miss":1, "Ms" : 1 , "Mme":1, "Mlle":1, "Mrs":1, "Mr":2, "Rare":3})
df["Title"] = df["Title"].astype(int)
df = pd.get_dummies(df, columns = ["Title"])
# 新建:家庭大小 (Fsize)特徵, 並依照大小分別建獨立欄位
df["Fsize"] = df["SibSp"] + df["Parch"] + 1
df['Single'] = df['Fsize'].map(lambda s: 1 if s == 1 else 0)
df['SmallF'] = df['Fsize'].map(lambda s: 1 if s == 2 else 0)
df['MedF'] = df['Fsize'].map(lambda s: 1 if 3 <= s <= 4 else 0)
df['LargeF'] = df['Fsize'].map(lambda s: 1 if s >= 5 else 0)
# Ticket : 如果不只是數字-取第一個空白之前的字串(去除'.'與'/'), 如果只是數字-設為'X', 最後再取 One Hot
Ticket = []
for i in list(df.Ticket):
if not i.isdigit() :
Ticket.append(i.replace(".","").replace("/","").strip().split(' ')[0])
else:
Ticket.append("X")
df["Ticket"] = Ticket
df = pd.get_dummies(df, columns = ["Ticket"], prefix="T")
# Cabib 依照第一碼分類, 再取 One Hot
df["Cabin"] = pd.Series([i[0] if not pd.isnull(i) else 'X' for i in df['Cabin'] ])
df = pd.get_dummies(df, columns = ["Cabin"], prefix="Cabin")
# Embarked, Pclass 取 One Hot
df = pd.get_dummies(df, columns = ["Embarked"], prefix="Em")
df["Pclass"] = df["Pclass"].astype("category")
df = pd.get_dummies(df, columns = ["Pclass"], prefix="Pc")
# 捨棄 Name 欄位
df.drop(labels = ["Name"], axis = 1, inplace = True)
na_check(df)
df.head()
# 將資料最大最小化
df = MinMaxScaler().fit_transform(df)
# 將前述轉換完畢資料 df , 重新切成 train_X, test_X
train_num = train_Y.shape[0]
train_X = df[:train_num]
test_X = df[train_num:]
# 使用三種模型 : 邏輯斯迴歸 / 梯度提升機 / 隨機森林, 參數使用 Random Search 尋找
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import GradientBoostingClassifier, RandomForestClassifier
lr = LogisticRegression(tol=0.001, penalty='l2', fit_intercept=True, C=1.0)
gdbt = GradientBoostingClassifier(tol=100, subsample=0.75, n_estimators=250, max_features=20,
max_depth=6, learning_rate=0.03)
rf = RandomForestClassifier(n_estimators=100, min_samples_split=2, min_samples_leaf=1,
max_features='sqrt', max_depth=6, bootstrap=True)
# 線性迴歸預測檔 (結果有部分隨機, 請以 Kaggle 計算的得分為準, 以下模型同理)
lr.fit(train_X, train_Y)
lr_pred = lr.predict_proba(test_X)[:,1]
sub = pd.DataFrame({'PassengerId': ids, 'Survived': lr_pred})
sub['Survived'] = sub['Survived'].map(lambda x:1 if x>0.5 else 0)
sub.to_csv('titanic_lr.csv', index=False)
# 梯度提升機預測檔
gdbt.fit(train_X, train_Y)
gdbt_pred = gdbt.predict_proba(test_X)[:,1]
sub = pd.DataFrame({'PassengerId': ids, 'Survived': gdbt_pred})
sub['Survived'] = sub['Survived'].map(lambda x:1 if x>0.5 else 0)
sub.to_csv('titanic_gdbt.csv', index=False)
# 隨機森林預測檔
rf.fit(train_X, train_Y)
rf_pred = rf.predict_proba(test_X)[:,1]
sub = pd.DataFrame({'PassengerId': ids, 'Survived': rf_pred})
sub['Survived'] = sub['Survived'].map(lambda x:1 if x>0.5 else 0)
sub.to_csv('titanic_rf.csv', index=False)
```
# 作業
* 分類預測的集成泛化, 也與回歸的很不一樣
既然分類的 Blending 要變成機率, 才比較容易集成,
那麼分類的 Stacking 要讓第一層的模型輸出機率當特徵, 應該要怎麼寫呢?
```
from mlxtend.classifier import StackingClassifier
meta_estimator = GradientBoostingClassifier(tol=100, subsample=0.70, n_estimators=50,
max_features='sqrt', max_depth=4, learning_rate=0.3)
"""
Your Code Here
"""
stacking = StackingClassifier(classifiers=[lr, gdbt, rf], meta_classifier=meta_estimator, use_probas=True, average_probas=True)
stacking.fit(train_X, train_Y)
stacking_pred = stacking.predict(test_X)
sub = pd.DataFrame({'PassengerId': ids, 'Survived': stacking_pred})
sub.to_csv('titanic_stacking.csv', index=False)
```
| github_jupyter |
# SLU09 - Linear Algebra & NumPy, Part 1
### Learning Notebook 1/2
In this notebook we will be covering the following:
- **Vectors**: definition, transpose, norm, multiplication by a scalar and addition, linear combinations, linear independence and dot product;
- **Introduction to NumPy arrays:** vectors and numpy arrays, shape, dimension, basic vector operations using arrays;
### Imports
```
# numpy is the package we're going to learn about
# it is a widespread convention to import numpy using the alias np
# this convention makes your code more readable, so do use it
import numpy as np
# auxiliary stuff
import utils
```
<br>
<img src="media/ds_without_linalg.jpg" width="580"/>
<br>
---
First of all, welcome to Week 6!! You probably know a lot about programming in Python at this point, so you should definitely feel proud of yourself.
However, to become a good data professional, you also need Mathematics. Just about any machine learning algorithm you might use will depend on linear algebra, calculus and statistics. Although you don't need to know all that goes on inside that black box, you do need to have an idea of what kind of input it takes and what it does to it, what are its **underlying assumptions** and **limitations**...
Simply feeding data into a machine learning algorithm to get predictions, will leave you in the same situation of a cat owner who puts his cat into the washing machine because its purpose is to clean...
##### *Don't be that cat owner...*
<img src="./media/cat_washing_machine.jpg" width="300"/>
---
#### Reading the matrix form of a machine learning algorithm:
By the end of SLUs 09 and 10, you'll be familiar with all the linear algebra you need to read the matrix form solution to the *multiple linear regression algorithm*, the most popular starting point for machine learning students:
$$ \mathbf{\beta} = (X^TX)^{-1}(X^T\mathbf{y})$$
That's a lot of Maths!
---
### ⚠️ Survival Guidelines for Week 6 ⚠️
---
**1 - Do not rush it**: It takes time to understand linear algebra, so take your time;
**2 - Follow the sequence of the learning materials**: don't jump sections - each one builds on the previous ones;
**3 - Try to solve the (ungraded) *pen and paper exercises***: they will pop through the learning notebook, to help you self-check your learning;
**4 - If you need help or have some feedback, do reach out**: [*There is no such thing as a dumb question*](https://en.wikipedia.org/wiki/No_such_thing_as_a_stupid_question). If you don't believe me, [believe him](https://en.wikipedia.org/wiki/Carl_Sagan).
**Last but not least**: That is a lot of linear algebra for 1 week. The most important thing here is that you get some intuition on these concepts, understand the rules of the game and make sure you know what you can and cannot do. You can always take some time with these concepts, after week 6 is over.
That's all. Ready? Let's go!
---
## 0. What is Linear Algebra?
[Linear algebra](https://en.wikipedia.org/wiki/Linear_algebra) is "the branch of mathematics concerning linear equations, linear functions and their representations in vector spaces and through matrices."! Cool, right?
Well, if you've never heard about Linear Algebra before, you probably have no idea what this means. Simplistically, this means that [linear algebra is about using](https://machinelearningmastery.com/gentle-introduction-linear-algebra/) "arithmetic on columns of numbers called vectors and arrays of numbers called matrices, to create new columns and arrays of numbers."
Besides the fact that we just made [Gauss](https://en.wikipedia.org/wiki/Carl_Friedrich_Gauss) turn in his grave, this was a much easier concept to grasp.
### How much Linear Algebra do I need?
Probably the **very least** you should be able to understand is:
- Scalar multiplication, addition and multiplication of vectors and matrices (tables of numbers), matrix inverse, and their properties;
- The intuition behind concepts such as linear combinations and linear independence;
- Understand systems of linear equations and the concept of eigenvalues and eigenvectors.
We will go through all of these concepts (*what?!*) in **SLUs 09 and 10**, starting from the very basics, and learn how to apply them using a library called [NumPy](https://numpy.org/).
---
<img src="./media/one_does_not_simply.jpg"/>
---
## 1. Vectors
Let's begin with a very basic example. Consider the kitten below, walking timidly at a speed of $0.1$ m/s, in order to reach his food bowl:
<img src="./media/kitten_walking.png" width="600"/>
- The value $0.1$ corresponds to a **magnitude** which tells us how fast the kitten is walking;
- The kitten is walking in a straight line in order to reach the food bowl, so we can easily define his **direction**.
The velocity of the kitten is a quantity **defined by both a magnitude and a direction**, which we call the velocity **vector** of the kitten, $\mathbf{v} = [0.1]$ m/s. We could draw this 1D (1-dimensional) vector in a 1D [coordinate system](https://en.wikipedia.org/wiki/Coordinate_system), as follows:
<img src="./media/kitten_vector.PNG" width="200"/>
---
Our kitten has now eaten all his food, and he's staring at you, asking for a refill:
<img src="./media/kitten_stopped.png" width="600"/>
He's not going to move until you feed him, but we could still represent his zero velocity by the zero vector $\mathbf{v} = [0]$ m/s.
---
We can actually define vectors in various dimensions.
A **2-dimensional vector** belongs to the 2-dimensional real coordinate space, $\mathbb{R}^2$, and we can plot it using a Cartesian coordinate system.
<img src="./media/vector_2d.png" width="240"/>
The vector $[-1, 2]$ has its tail located at the origin of the x-y plane, $(0,0)$, and its tip (head) located at the point $(-1, 2)$. To go from tail to tip, we walk 1 step leftwards (x-coordinate = -1) and 2 steps upwards (y-coordinate = 2). Note that, in linear algebra, we root the vector at the origin of the coordinate system.
Imagine a cyclist climbing a hill at 6 km/h (it's a steep climb!). We could draw the velocity vector $\mathbf{v}$ of the cyclist in the xy-plane:
<img src="./media/cyclist.png" width="320"/>
> 📝 **Pen and paper exercise 1**: Grab a pen and a piece of paper and draw the vectors $[-1, 2]$, $[2, -1]$ and $[1, 2]$ on the xy-plane. Notice that they all have the same *magnitude* (length) but different *direction*. We'll see how to determine this length in section 1.2.
A **3-dimensional vector** belongs to the 3-dimensional real coordinate space, $\mathbb{R}^3$. We can draw it on the xyz coordinate system, using the same logic as for the xy-plane. To get from the tail to the tip of the vector, for the first component (x coordinate) you would walk parallelly to the yz [plane](https://en.wikipedia.org/wiki/Plane_(geometry)), for the second (y coordinate) you would walk parallelly to the xz plane, and for the third element of the vector (z coordinate), you would walk parallelly to the xy plane. You can play with your own 3-D vectors in this applet: https://www.intmath.com/vectors/3d-space-interactive-applet.php.
What about a **4-dimensional vector**?
<img src="./media/brain_says_no.jpg" width="380"/><br>
The human brain is not able to *visualize* more than 3 dimensions, although it's possible to overcome this limitation with the help of your own imagination, brilliant [interactive visualizations](https://ciechanow.ski/tesseract/) or the smoothing voice of [Carl Sagan](https://vimeo.com/199561184).... Hey, don't be sad. It doesn't really matter whether or not you can see in 4D. In linear algebra you can extend properties of vectors and matrices, and the operations between them, to any number of dimensions.
An $m$-dimensional vector belongs to a [real coordinate space](https://en.wikipedia.org/wiki/Real_coordinate_space) of $m$ dimensions, denoted by $\mathbb{R}^m$, where we have the set of all different $m$-dimensional vectors.
### 1.1 Vector definition
<a name="vector_def"></a>
<div class="alert alert-block alert-info">
An <b>$m$-dimensional vector $\mathbf{x}$</b> is an ordered list of $m$ scalars represented as $\mathbf{x} = \left[x_1, x_2,..., x_m\right]$, $x_i \in \mathbb{R}$. It has a <b>magnitude</b> and a <b>direction</b>.
</div>
**Equality of vectors**: Two vectors $\mathbf{u}$ and $\mathbf{v}$ are equal only if they have the same magnitude and the same direction. This is the same as saying two vectors are equal if the ordered lists which represent them are equal, element-wise.
- **Geometrically** speaking, a vector is an arrow pointing in space, with a given *magnitude* (length), and a *direction*, describing where the arrow points to.
- **Numerically** speaking, you can think of a vector as an ordered list of scalars (real numbers).
$x_i \in \mathbb{R}$ means that each scalar $x_i$ in the vector belongs ($\in$) to the set of all real numbers ($\mathbb{R}$). $m$ belongs to the set of all positive integer numbers, $m \in \mathbb{Z}^+$. Also note that when describing vectors we usually use square brackets `[]` and **not** round brackets `()`, although these might be used somewhere else.
Vectors are usually represented by bold lowercase letters and scalars by a non-bold lowercase letter. However, you might find different notations. For example, arrow over lowercase letter, $\overrightarrow{v}$, or both arrow and uppercase letter, like the [force vector](https://en.wikipedia.org/wiki/Force) $\overrightarrow{F}$.
#### 1.1.1 Vector representations and the transpose
We can represent **the same** vector in several ways. For example, we can represent a given *4-dimensional vector* as:
* an ordered list, $\left[0,\; -1,\; 2.6,\; \sqrt{3}\right]$,
<br>
<br>
* a **row vector**,
$
\begin{bmatrix}
0 & -1 & 2.6 & \sqrt{3}\\
\end{bmatrix}
$,
* or its **transpose**, a **column vector**,
$
\begin{bmatrix}
0 & -1 & 2.6 & \sqrt{3}\\
\end{bmatrix}^T =
\begin{bmatrix}
0 \\
-1 \\
2.6 \\
\sqrt{3} \\
\end{bmatrix}
$.
The relevance of the type of representation we use will become evident when we introduce matrices. For now, just know that the row representation of a vector is called the **transpose** of its column representation, and vice versa.
### 1.2 Vector norm
The **magnitude** of a 2-dimensional vector, also called the **norm** or the **length**, can be determined by the [Pythagorean theorem](https://en.wikipedia.org/wiki/Pythagorean_theorem), which says that "In a right angled triangle, the square of the hypotenuse is equal to the sum of the squares of the other two sides".
On the xy-plane below, the dashed lines represent the two other sides of a right angled triangle, and the hypothenuse corresponds to the length of the vector:
<br>
<img src="./media/vector_2d.png" width="240"/>
We can represent the vector on the image by an ordered list: $\mathbf{a} = [a_1, a_2]$, with components $a_1 = -1$ ($x$ coordinate) and $a_2 = 2$ ($y$ coordinate).
Hence, we'll have a right-angled triangle with sides length equal to $1$ and $2$, and an unknown hypotenuse length, which corresponds to the norm of the vector.
Let's use the Pythagorean theorem to **find the norm of $\mathbf{a}$**, $\| a\|$:
$$\| a\|^2 = a_1^2 + a_2^2$$
$$\| a\| = \sqrt{a_1^2 + a_2^2} = \sqrt{(-1)^2 + (2)^2} = \sqrt{5} $$
You can actually use this formula with any $m$-dimensional vector. (*It's a kind of magic...* 🎵)
<div class="alert alert-block alert-info">
The <b>norm of an $m$-dimensional vector</b> $\mathbf{x} = \left[x_1, x_2, ..., x_m\right]$, $x_i\in \mathbb{R}$, also known as the magnitude or length, is defined as $\|\mathbf{x}\| = \sqrt{x_1^2 + x_2^2 + ... + x_m^2}$.
</div>
> 📝 **Pen and paper exercise 2**: Calculate the norm of the vectors $[-1, 2]$, $[2, -1]$ and $[1, 2]$, which you've drawn in the last exercise. You should find that they all have the same norm, as expected. Find one more vector with the same norm as them.
>
> **Notice that** there is an infinite number of 2-D vectors with the same norm (length).
---
### Ever asked yourself...
<br>
<img src="./media/straightforward.jpg" width="380"/>
[*bad pun source*](https://math.stackexchange.com/questions/62789/what-does-linear-mean-in-linear-algebra#comment146861_62789)
Now to the [serious answer](https://math.stackexchange.com/questions/62789/what-does-linear-mean-in-linear-algebra/62791#62791): linear algebra is "linear" because it's about linear functions. Remember $y = mx + b$, the equation for the line?
```
# run this cell and check the plots below
utils.plot_school_functions()
```
In linear algebra we deal with linear functions (*deal with it!*). This means that all transformations we do on our data are based on linear relations, just like the line on the first plot. We don't need to worry about polynomial functions, exponentials, or other evil sourceries. Only simple, beautiful, linear magic. 😍😃
Simple, yet powerful.
---
### 1.3 Vector operations: multiplication by scalar and addition
#### 1.3.1 Multiplying a vector by a scalar
What happens with a vector if we multiply it by a scalar?
Consider the vector $\mathbf{u}=\begin{bmatrix}1\\2\\\end{bmatrix}$. On the image below you can see several vectors that result from multiplying the vector $\mathbf{u}$ by different scalars.
<br><img src="./media/vector_scaling.png" width="400"/>
**Multiplying $\mathbf{u}$ by -1:** $\hspace{5cm}\mathbf{v}=-1\cdot \mathbf{u}=-1\cdot\begin{bmatrix}1\\2\\\end{bmatrix}$ $=\begin{bmatrix}-1\times 1\\-1\times 2\\\end{bmatrix}=$ $\begin{bmatrix}-1\\-2\\\end{bmatrix}$
Multiplying a 2D vector by $-1$ causes it to rotate $180^{\circ}$ ($\pi$ radians) around the origin. Its *magnitude* (norm) remains the same, but the *direction* changes.
**Multiplying $\mathbf{u}$ by 0:**$\hspace{5cm} 0 \cdot \mathbf{u}=0\cdot\begin{bmatrix}1\\2\\\end{bmatrix} = $ $\begin{bmatrix}0\\0\\\end{bmatrix}$
Multiplying any vector by $0$ results in a vector with the same dimension, where all components are zero (the zero vector).
**Multiplying $\mathbf{u}$ by 2**$:\hspace{5cm} \mathbf{w}=2\cdot \mathbf{u}=2\cdot\begin{bmatrix}1\\2\\\end{bmatrix}$ $= \begin{bmatrix}2\\4\\\end{bmatrix}$
Multiplying a vector by a positive scalar increases its *magnitude* but does not affect its *direction*.
**Multiplying $\mathbf{u}$ by 1:**$\hspace{5cm}
1\cdot\mathbf{u}=1\cdot\begin{bmatrix}1\\2\\\end{bmatrix} = $ $\begin{bmatrix}1\times 1\\1\times 2\\\end{bmatrix} = \begin{bmatrix}1\\2\\\end{bmatrix} = \mathbf{u}$
Multiplying any vector by the scalar 1 does not change the vector (**identity property**).
> Note that the dot symbol $\cdot $ in the expressions above denotes multiplication, however as we'll see in a few sections, when it is written between two vectors it means dot product instead.
> 📝 **Pen and paper exercise 3**: Multiply the vector $\mathbf{u}$ by the scalar -2. What happens to its *magnitude* and *direction*?
---
#### 1.3.2 Addition and subtraction
To add two $m$-dimensional vectors, we simply add the corresponding components from each vector.
For example, we can add vectors $\mathbf{u} = \begin{bmatrix} 1\\ 2\\\end{bmatrix}$ and $\mathbf{v} = \begin{bmatrix} 3\\ 1\\\end{bmatrix}$ as follows: $\hspace{.2cm} \mathbf{w} = \mathbf{u} + \mathbf{v} = \begin{bmatrix} 1\\ 2\\\end{bmatrix} + $ $\begin{bmatrix} 3\\ 1\\\end{bmatrix} = $ $\begin{bmatrix} 1 + 3\\ 2 + 1\\\end{bmatrix} = $ $\begin{bmatrix} 4\\ 3\\\end{bmatrix}$
**Geometrical interpretation:**
<img src="./media/vector_addition.png" width="340"/>
The vectors $\mathbf{u}$, $\mathbf{v}$ and $\mathbf{w}$ are plotted on the image above. Notice that $\mathbf{b}$ is equivalent to the vector $\mathbf{v}$ (same magnitude and same direction), the only difference being that it is not rooted at the origin. The same happens with vector $\mathbf{a}$ in relation to vector $\mathbf{u}$.
Vector $\mathbf{u}$ takes us from point $(0,0)$ to point $(1,2)$. After placing the tail of vector $\mathbf{b}$ at the tip of vector $\mathbf{u}$, we walk 3 steps rightwards (x-coordinate of $\mathbf{v}$ = 3) and 1 step upwards (y-coordinate of $\mathbf{v}$ = 1), getting to the tip of $\mathbf{w}$, the result of adding $\mathbf{u}$ and $\mathbf{v}$.
**Vector addition is commutative:**
Notice that we could also start at vector $\mathbf{v}$, add vector $\mathbf{a}$ (the equivalent of vector $\mathbf{u}$), and we would still get the vector $\mathbf{w}$ as a result. This means that the **addition between vectors is commutative**. The same applies to any two or more $m$-dimensional vectors added together.
> ❗ You **cannot** add vectors with different dimensions!!
>
> For example, if you tried to add $[1, 2]$ with $[1, 2, 3]$, you would have no corresponding component on the first vector to add to the third component of the second vector.
> 📝 **Pen and paper exercise 4**: Grab a pen and paper and draw the vector $\mathbf{x} = \mathbf{u} - \mathbf{v}$ on the xy-plane:
> - Multiply the vector $\mathbf{v}$ by the scalar $-1$ and draw the resulting vector, $\mathbf{-v}$;
> - Add $\mathbf{-v}$ to vector $\mathbf{u}$;
**Properties of vector addition and scalar multiplication (for any $m$-dimensional vectors):**
$\;\;\text{1. }\;\; \mathbf{u} + \mathbf{v} = \mathbf{v} + \mathbf{u}$
$\;\;\text{2. }\;\; \mathbf{u} + \mathbf{0} = \mathbf{u}$
$\;\;\text{3. }\;\; c\left(\mathbf{u} + \mathbf{v}\right) = c\mathbf{u} + c\mathbf{v},\hspace{.2cm} c\in \mathbb{R}$
$\;\;\text{4. }\;\; \left(cd\right)\mathbf{u} = c\left(d\mathbf{u}\right),\hspace{.2cm} c,d \in \mathbb{R}$
$\;\;\text{5. }\;\; \mathbf{u} + (\mathbf{v} + \mathbf{w}) = (\mathbf{u} + \mathbf{v}) + \mathbf{w}$
$\;\;\text{6. }\;\; \mathbf{u} + (-\mathbf{u}) = \mathbf{0}$
$\;\;\text{7. }\;\; (c + d) \mathbf{u} = c \mathbf{u} + d \mathbf{u}$
$\;\;\text{8. }\;\; 1\mathbf{u} = \mathbf{u}$
> 📝 **Pen and paper exercise 5 (this one is for the skeptical minds)**: Choose 2 of the properties above and check their veracity.
>
> You can use, for example, the vectors $\mathbf{u} = \begin{bmatrix}1\\ 2\end{bmatrix},\;\;$
> $\mathbf{v} = \begin{bmatrix}2\\ 4\end{bmatrix},\;\;$
> $\mathbf{w} = \begin{bmatrix}0\\ -1\end{bmatrix}\;\;$
> and the scalars (real numbers) $c=-0.2$ and $d=\frac{1}{4}$.
If you "find out" any of the rules is wrong, I'm sorry but you probably made some error on the arithmetics along the way.
***Trust me, linear algebra will never fail you. Never. Ever.***
<img src="./media/we_will_see.gif"/>
### 1.4 Linear combinations and linear independence
Everytime we scale vectors and add them together, we're performing a **linear combination**. This is what it looks like for 2 $m$-dimensional vectors:
$$c_1 \cdot \begin{bmatrix} u_1\\ u_2\\ ... \\ u_m\\\end{bmatrix}+c_2\cdot\begin{bmatrix}v_1\\v_2\\\dots\\v_m\\\end{bmatrix}=\begin{bmatrix}c_1\cdot u_1 + c_2\cdot v_1\\ c_1\cdot u_2 + c_2\cdot v_2\\ ... \\ c_1\cdot u_m + c_2\cdot v_m\\\end{bmatrix},\hspace{.2cm} c_i\in \mathbb{R}$$
In the *pen and paper* exercise **4**, you basically performed a linear combination between vectors using scalars $1$ and $-1$.
---
We can have a linear combination of $n$ vectors, as follows:
$$c_1\cdot \mathbf{x_1} + c_2\cdot \mathbf{x_2} + ... + c_n\cdot \mathbf{x_n},\hspace{.2cm} c_i\in \mathbb{R},\hspace{.2cm} \mathbf{x_i}\in\mathbb{R}^m$$
Note that $\mathbf{x_1}, \mathbf{x_2},..., \mathbf{x_n}$ are **not vector components but actual vectors** (bold lowercase letter).
---
#### Example
For $\mathbf{v_1} = \begin{bmatrix}1\\-1\end{bmatrix}$, $\mathbf{v_2} = \begin{bmatrix}2\\2\end{bmatrix}$ and $\mathbf{w} = \begin{bmatrix}4\\0\end{bmatrix}$, we have $w = 2\cdot \mathbf{v_1} + 1\cdot\mathbf{v_2}$. Thus $\mathbf{w}$ can be written as a linear combination of $\mathbf{v_1}$ and $\mathbf{v_2}$.
---
**Linear (in)dependence**
If we have two vectors $\mathbf{a}$ and $\mathbf{b}$, and $\mathbf{b}$ can be expressed as $c\cdot \mathbf{a}, c\in \mathbb{R}$, we say that $\mathbf{a}$ and $\mathbf{b}$ are **linearly dependent**, or collinear. If either one cannot be expressed as a linear combination of the other, then we say they are **linearly independent**, or non-collinear.
---
For example, $[2, 1]$ and $[4, 2]$ are linearly dependent, but $[2, 1]$ and $[0, 1]$ are linearly independent.
---
Generally, a set of vectors is said to be [linearly dependent](https://en.wikipedia.org/wiki/Linear_independence) if at least one of the vectors in the set can be defined as a linear combination of the others.
> 📌 **Tip**: The concept of linear dependence is extremely important in data science!
<img src="./media/noted.gif" width="500"/>
### 1.5 Representing all vectors in space
<img src="./media/linear_combinations.png" width="400"/>
In the image above we can see the resulting vectors of 4 distinct linear combinations of vectors $\mathbf{u}=[1,2]$ and $\mathbf{v}=[3,1]$, namely:
- $1\cdot \mathbf{u} + 1\cdot \mathbf{v}$
- $2\cdot \mathbf{u} + 1\cdot \mathbf{v}$
- $-1\cdot \mathbf{u} + 1\cdot \mathbf{v}$
- $1\cdot \mathbf{u} + (-1)\cdot \mathbf{v}$
Actually, if you had the time (*infinite time*) to plot all possible linear combinations of vectors $\mathbf{u}$ and $\mathbf{v}$, you would fill the entire xy-plane, and get **all the 2-dimensional vectors, this is, all the vectors in $\mathbb{R}^2$**.
**But only because** these 2 2D vectors are **linearly independent**!!
---
We could **not** create the set of all the 2-dimensional vectors if our vectors were **linearly dependent**.
If you don't believe me, write some linear combinations ($c\mathbf{u} + d\mathbf{v}$) for the collinear vectors $\mathbf{u}=[1,2]$ and $\mathbf{v} = [2,4]$, using any scalars you wish.
Now try not to get [stuck on the line](https://www.theguardian.com/politics/video/2012/aug/01/boris-johnson-stuck-zip-wire-video)...
<img src="./media/stuck_line.png" width="400"/>
Wrapping up:
- in a **2D** space, we need **2 and only 2 linearly independent vectors** to define all other 2-dimensional vectors as linear combinations of these 2 vectors;
- in a **3D** space, we need **3 and only 3 linearly independent vectors** to define all other 3-dimensional vectors as linear combinations of these 3 vectors;
- and so on and so forth.
This also means that, for example, if you define 3 vectors in 2-dimensional space, any one of them will be a linear combination of the other two.
Linear algebra is a minimalist and does not like to have more than the essential: if you can use only 2 vectors to represent a 2D space, why waste a 3rd one?
### 1.6 Dot product
#### 1.6.1 Definition of the dot product
We already know how to multiply vectors by scalars and add vectors together. But can we multiply one vector by another? Yes we can! Actually, we can do it in [several ways](https://en.wikipedia.org/wiki/Multiplication_of_vectors), however, let's just focus on the most simple one: the **dot product**, also known as the **scalar product**, because the result is a scalar.
<div class="alert alert-block alert-info">
The <b>dot product</b> of two $m$-dimensional vectors $\mathbf{u}=[u_1, u_2, ..., u_m]$ and $\mathbf{v}=[v_1, v_2, ..., v_m]$ is a <b>scalar</b> given by:
$$\mathbf{u}\cdot \mathbf{v} = u_1 v_1 + u_2 v_2 + ... + u_m v_m$$
</div>
Consider the vectors $\mathbf{a} = [1, 2, 0]$ and $\mathbf{b} = [-1, 4, -2]$. The dot product between $\mathbf{a}$ and $\mathbf{b}$ is:
$$\mathbf{a}\cdot \mathbf{b} = 1\times (-1) + 2\times 4 + 0\times (-2) = -1 + 8 + 0 = 7$$
---
You might also find the dot product of two vectors written as $\mathbf{u} \cdot \mathbf{v} = \|u\|\|v\|\cos{\theta} = \|v\|\|u\|\cos{\theta}$.
This means that $\mathbf{u} \cdot \mathbf{v}$ is the magnitude of $\mathbf{v}$ times the magnitude of the component of $\mathbf{u}$ that points along $\mathbf{v}$, namely $\|u\|\cos{\theta}$ (projection of $\mathbf{u}$ onto $\mathbf{v}$):
<img src="./media/projection_u_onto_v.png" width="400"/>
We can therefore determine the angle between any two *non zero* vectors by using the relation: $\;\;\;\;\mathbf{u} \cdot \mathbf{v} = \|u\|\|v\|\cos{\theta} \iff \cos{\theta} = \frac{\mathbf{u} \cdot \mathbf{v}} {\|u\|\|v\|}$
> 📝 **Pen and paper exercise 6 (just for the adventurous!!)**: Find the angle $\theta$ (in degrees or radians) between the vectors $\mathbf{u} = \begin{bmatrix}1\\ 0\end{bmatrix}$ and $\mathbf{v} = \begin{bmatrix}2\\ 2\sqrt{3}\end{bmatrix}$ using the formula $\mathbf{u} \cdot \mathbf{v} = \|u\|\|v\|\cos{\theta}$.
> - Calculate $\mathbf{u}\cdot\mathbf{v}$ using the [dot product formula](#1.6-Dot-product);
> - Find $\|u\|\|v\|$ using the formula for the [norm of a vector](#1.2-Vector-norm).
>
> You get **3 ⭐️s** if you solve this using a calculator and **5 ⭐️s** if you can solve it without a calculator (you may use the [unit circle](https://en.wikipedia.org/wiki/Unit_circle) below).
<img src="./media/unit_circle.png" width="300"/>
Once more, all of this applies to any two vectors in any $m$-dimensional real space.
#### 1.6.2 Properties of the dot product
You don't need to memorize them all, just know they exist.
$\;\;\;\;\;\text{1. }\;\; \mathbf{u} \cdot \mathbf{u} = \|\mathbf{u}\|^2$
$\;\;\;\;\;\text{2. }\;\; \mathbf{0} \cdot \mathbf{u} = \mathbf{0}$
$\;\;\;\;\;\text{3. }\;\; \mathbf{u} \cdot \mathbf{v} = \mathbf{v} \cdot \mathbf{u}$
$\;\;\;\;\;\text{4. }\;\; (c \mathbf{u}) \cdot \mathbf{v} = c (\mathbf{u} \cdot \mathbf{v})$
$\;\;\;\;\;\text{5. }\;\; \mathbf{u} \cdot \mathbf{v} = \|\mathbf{u}\|\|\mathbf{v}\|\cos{\theta}$
$\;\;\;\;\;\text{6. }\;\; \mathbf{u} \cdot (\mathbf{v} + \mathbf{w}) = \mathbf{u} \cdot \mathbf{v} + \mathbf{u} \cdot \mathbf{w}$
Remember that $c$ is a scalar (non-bold lowercase letter) and $\mathbf{u}$, $\mathbf{v}$ and $\mathbf{w}$ represent vectors (bold lowercase letters). Also, we have here a special vector, the **zero vector**, where all elements are equal to zero, which we denote by $\mathbf{0}$.
If you're skeptical about dot product properties, check this [video](https://www.youtube.com/watch?v=rVQ3G9epCjw).
### 1.7 Orthogonal vectors
<div class="alert alert-block alert-info">
Two vectors $\mathbf{u}$ and $\mathbf{v}$ are said to be <b>orthogonal</b> if their dot product is equal to zero: $\;\;\mathbf{u}\cdot \mathbf{v} = \mathbf{0}$
</div>
If we think about the formula $\mathbf{u} \cdot \mathbf{v} = \|v\|\|u\|\cos{\theta}$, we see that $\|u\|\cos{\theta}$ (projection of vector $\mathbf{u}$ onto $\mathbf{v}$) fits in a point at the tail of $\mathbf{v}$, having magnitude zero:
<img src="./media/orthogonal_vectors.PNG" width="200"/>
> 📝 **Pen and paper exercise 7**: Determine the dot product between vectors $[1,0]$ and $[0,-2]$. Are they orthogonal?
---
### 1.8 Vectors recap
1. Vectors can be represented as **ordered lists of scalars**; vectors have both **magnitude and direction**;
2. The **transpose** of the row vector is a column vector, and vice-versa;
3. The **norm**/magnitude/length of an $m$-dimensional vector $\mathbf{x}$ is given by $\| \mathbf{x}\| = \sqrt{x_1^2 + x_2^2 + ... + x_m^2}$;
4. Everytime we scale vectors and/or add them together, we're performing **linear combinations**;
5. We can represent the set of all $m$-dimensional vectors using linear combinations of $m$ linearly independent vectors (also $m$-dimensional);
6. Several properties of addition and multiplication by scalars are generalizable for vectors, such as commutativity, associativity and distributivity;
7. The **dot product** between two vectors, $\mathbf{u} \cdot \mathbf{v}$, can be defined as $\mathbf{u} \cdot \mathbf{v} = \|u\|\|v\|\cos{\theta}$, where $\theta$ refers to the angle made by $\mathbf{u}$ and $\mathbf{v}$, or $\mathbf{u} \cdot \mathbf{v} = u_1 v_1 + u_2 v_2 + ... + u_m v_m.$
---
Break time!
Look through your window and watch the world outside. You were blind and now you see: everywhere there are vectors, everywhere there is linear algebra...
<img src="./media/pause_time.gif" width="420"/>
See you soon!
---
## 2. Introduction to NumPy arrays
No old school pen and paper on this section!! Time to put Python and your machine to work. 💻🐍
### 2.1 The NumPy package
You might have noticed we have imported a package at the beginning of the notebook, which goes by the name of ``numpy``:
```python
# it is a widespread convention to import numpy using the alias np
# this convention makes your code more readable, so do use it
import numpy as np
```
`numpy` is commonly imported with the alias `np`. This means that every time we instantiate a new object from NumPy, call a NumPy function or use a module from it, we'll use `np` instead of `numpy`. I know it's fun to be different, and make up your own aliases, but it's better to be readable than to be laughable.
[NumPy](https://numpy.org/) is the fundamental package for scientific computing with Python. Among many other amazing possibilities, it allows us to work efficiently with vectors and matrices, performing lots of linear algebra operations.
In the `requirements.txt` file, you can see that we are using version 1.18 of NumPy. A detailed reference documentation of the functions and classes contained in this package is available in the [NumPy reference](https://numpy.org/doc/1.18/reference/index.html) webpage. You can also download the [pdf version](https://numpy.org/doc/1.18/numpy-ref.pdf) here.
#### 2.1.1 Why NumPy?
You might remember lists from SLU02. You already know that you can represent a vector by an ordered list, as follows:
```
u = [2, -1, 0, 2, 0.4, 3, 6, 0, 1] # 9-dimensional vector
print("Length of the list representing vector u (not the same as the length of the vector!):", len(u))
```
Using Python lists and for loops, we could implement some basic operations. At first, you might think this is a reasonable approach for small tasks such as linear combinations of vectors (recall [section 1.4](#1.4-Linear-combinations-and-linear-independence)). But is it?
Let's run a simulation to compare using Python lists with using NumPy (do not worry about the code behind this).
We'll see how long it would take to compute a simple linear combination of the form $2\mathbf{u} + 2\mathbf{v}$ between two vectors of length $10^6$, using NumPy *versus* Python lists.
**Creating a linear combination using NumPy**
```
# perform linear combination using NumPy magic
numpy_duration = utils.lincomb_numpy()
print("Using NumPy arrays we took {:.2} seconds to perform a linear combination.".format(numpy_duration))
```
**Creating a linear combination using Python lists and `for` loops**
```
# perform linear combination using Python lists and for loops
python_duration = utils.lincomb_lists()
print("Using Python lists we took {:.2} seconds to perform a linear combination.".format(python_duration))
```
How much faster was NumPy?...
```
print("Python lists approach was {} times SLOWER than NumPy!!".format(int(python_duration/numpy_duration)))
```
#### [Ain't nobody got time for that!!](https://www.youtube.com/watch?v=bFEoMO0pc7k&feature=youtu.be&t=10)
Other than being much faster, NumPy is also awesome because:
- It uses less memory to store the same amount of data
- It interfaces with libraries you'll often use, such as Pandas, Scikit-learn, Matplotlib and many others;
- It supports a great variety of numerical types;
- It has a comprehensive list of [functions, modules and objects](https://numpy.org/doc/1.18/reference/index.html) useful for linear algebra.
By the way, now is the time to start getting comfortable reading documentation. You can either:
- refer to documentation pages (for example, you could consult the [documentation webpage for version 1.18.1 of NumPy](https://numpy.org/doc/1.18/) to check its functionalities);
- access the docstring using `?` inside a jupyter cell.
```
# write the name of the function followed by a quotation mark
# The docstring with basic information on the function
# should appear at the lower part of your browser.
print?
```
<img src="./media/racoon.png"/>
You can close the pager with the docstring when you're done.
### 2.2 The `ndarray`
The main object in NumPy is the [NumPy array](https://numpy.org/doc/1.18/reference/arrays.ndarray.html), or simply `ndarray`. An ndarray is a collection of items, all of the same size and type. You can think of arrays as tables (2 dimensions), but you can actually have arrays of 3, 4 or 5 dimensions, and so on. Here we're deal mainly with ndarrays of 1 or 2 dimensions, where the items stored are numbers.
#### 2.2.1 Create an `ndarray`
Let's create an array to represent a vector, using [np.array()](https://numpy.org/doc/1.18/reference/generated/numpy.array.html):
```
u = [0, 1, 2] # this is a Python list representing a vector
a = np.array([0, 1, 2]) # create a NumPy array object using the Python list above
a
```
You can also create a table of numbers (a matrix) from a list of lists, using `np.array()`:
```
# notice we input 1 list with 3 lists in it
b = np.array([[0, 1, 2], # 1st row
[3, 4, 5], # 2nd row
[6, 7, 8]]) # 3rd row
print(b) # you can use Python's built-in function print() to print an array
```
#### 2.2.2 Dimensions and shape of an array
If you are to manipulate arrays correctly, you definitely need to know what **shape** and **dimension** mean in terms of NumPy arrays.
**Number of array dimensions**
Let's start by checking the array dimensions of `a`, which represents a 3-dimensional vector, using the attribute **`ndim`**:
```
a = np.array([0, 1, 2]) # a 3-dimensional row vector
a.ndim # number of array dimensions
```
Wait, what?! Our array has 1 dimension?... But our vector is 3-dimensional!... What's happening?
Remember lists of lists? Well, you can think of arrays in a similar way. You would use two brackets to access a single value in a list of lists, as follows:
```
nested_list = [[2, 4, 6],
[8, 10, 12]]
# the first index will access the first list -> nested_list[0]
# the second index will access the second element in the first list -> nested_list[0][2]
nested_list[0][2]
```
You could also access the first list inside the list as follows:
```
nested_list[0]
```
Just as you can nest a list inside another list, which is inside another list, and so on... so too can you create as many *axes* (dimensions) in arrays as you wish. You could access the first axis (*axis 0*) with the first pair of brackets `[]`, the second axis (*axis 1*) with the second pair of brackets `[]`, and so on.
<img src="./media/array_axes.png" width="500"/>
---
Don't worry if this seems too confusing right now. We'll check some examples to see how this works.
A 2D array (2 dimensions) has 2 axes. You can think of it as a table of numbers (matrix):
```
# you can think of b as a table (matrix) represented by a numpy array
b = np.array(
[[0, 1, 2],
[3, 4, 5],
[6, 7, 8]]
)
b.ndim # b is a 2D array
```
We can access the first row in the table like this:
```
b[0] # access 1st row in b
```
We can access the second element of the first row in the table as follows:
```
b[0][1] # access 2nd element of the 1st row in b
```
We can't access a third dimension because the array is 2D. Thus, the following command will throw an `IndexError` (which we'll catch, because we're awesome):
```
# trying to access an element in the 3rd axis of a 2D array does not compute
try:
b[0][2][0]
except IndexError as e:
print("IndexError:", e)
```
We can represent a column vector with a 2D array (2 axes):
```
a = np.array([[0],
[1],
[2]]) # a 2D numpy array, a 3-dimensional COLUMN vector
a.ndim # number of array dimensions
```
Notice the difference between a 1D array:
```
np.array([0, 1, 2])
```
and a 2D array with the same elements:
```
np.array([[0, 1, 2]])
```
In the 2D array, we have one extra outside square bracket (just like in a nested list).
**Shape of an array**
The other attribute you should understand is the **shape** of the array. An array's shape is a *tuple of integers* which indicates the size of the array in each dimension (axis). Hence, for a table (matrix) with $m$ rows and $n$ columns, the shape will be $(m, n)$.
The length of the shape tuple corresponds to the number of axes, given by `.ndim`, as we just saw.
```
a = np.array([[0],
[1],
[2]]) # same vector as in the last code cell
a.shape # shape of the array (number of elements in axis 0, number of elements in axis 1)
```
Above, we see that axis 0 has size 3 (3 rows in the column vector) and axis 1 has size 1 (1 column).
Luckily for us, we wont need more than 2 dimensions to represent vectors and matrices.
---
Notice that the underlying class which creates the NumPy array is the class `numpy.ndarray`. However, it is advisable to construct arrays using its built-in functions, such as `array`, `zeros` or `ones`.
For simplicity, we'll refer to NumPy arrays and arrays interchangeably throughout this notebook (note that these are **not** the Python [`array.array`](https://www.tutorialspoint.com/python/python_arrays.htm) objects).
```Python
if (student.question == "What is a class?") or (student.question == "What is an object?"):
print("How dare you?! You go review SLU07!")
```
You can check a quick explanation of `ndarrays` [here](https://www.tutorialspoint.com/numpy/numpy_ndarray_object.htm).
---
### 2.3 Vectors and linear algebra using NumPy
Time to have some fun! Let's put all the knowledge we gathered about vectors to use.
#### 2.3.1 Vector transpose
Remember learning that the transpose of a row vector is a column vector and vice-versa? NumPy has the transpose implemented as an attribute of arrays.
If we start with a 4-dimensional column vector, represented by an array of shape `(4, 1)`, we'll have `2` axes (`.ndim` = 2):
```
a = np.array([[0],
[1],
[2],
[3]]) # a 4-dimensional column vector
print("a:\n", a, "\n")
print("a.shape:", a.shape)
print("a.ndim:", a.ndim)
```
Getting the attribute `.T` (for transpose) of the array will return a row vector represented by a 2D array, just as we expected:
```
a_T = a.T # the transpose of a
print("a_T:", a_T, "\n")
print("a_T.shape:", a_T.shape)
print("a_T.ndim:", a.ndim)
```
**What would happen if you used a 1D array?**
If we try to transpose an array with only 1 dimension (thus the tuple shape has only 1 element), we get exactly the same shape!!
```
print("shape of 1D array: ", np.array([0,1,2,3]).shape)
print("shape of the transpose of 1D array: ", np.array([0,1,2,3]).T.shape)
```
For a 1D array, we cannot get the transpose of our vector using `.T`!
---
#### `reshape()`
To **get the transpose of a vector represented by a 1D array in NumPy**, we would need to **first reshape** the array to 2D, using `.reshape()`, with argument `(1, -1)`, to get a **row vector**, or `(-1, 1)` to get a **column vector**.
The `reshape` method allows you to reshape an array of data to any given shape.
For example, for the 1-dimensional array below, `array_1d`, with 6 elements (shape = `(6,)`)...
```
array_1d = np.array([0, 1, 2, 3, 4, 5]) # 1-dimensional array, with 6 elements, shape is (6, )
print(array_1d)
print("shape: ", array_1d.shape)
```
...we can reshape it to a 2-dimensional array with the **same** 6 elements, displayed in a 2-dimensional array of shape `(3, 2)`:
```
array_reshaped = array_1d.reshape((3, 2)) # reshape to a table!
print(array_reshaped)
print("shape: ", array_reshaped.shape)
```
We can't however reshape to a shape which is not compatible with the number of elements we have (say, for example, `(4, 2)`):
```
# we can't reshape to a table with 8 entries 'cause we only have 6 elements in the array!
# we'll just catch that ValueError, that NumPy will throw at us, flawlessly
try:
array_1d.reshape((4, 2))
except ValueError as e:
print("ValueError:", e)
```
---
Let's now consider a 1-dimensional array representing a 4-dimensional vector:
```
a = np.array([0,1,2,3]) # vector represented by a 1D array
a.shape # shape of array a
```
We can convert this 1D array to a 2D array row vector using `reshape()` with argument `(1, -1)`:
```
a_row = a.reshape((1, -1)) # use reshape to get a 2D array representation of a row vector
print("a_row:\n", a_row, "\n")
print("a_row.shape:", a_row.shape)
```
We could also convert it directly to a column vector using `reshape()` with argument `(-1, 1)`:
```
a_column = a.reshape((-1, 1)) # use reshape to get a 2D array representation of a column vector
print("a_column:\n", a_column, "\n")
print("a_column.shape:", a_column.shape)
```
Because we now have our vector in a 2D array, we could use the transpose attribute safely:
```
# same result
print(a_column.T)
```
> 📌 **Tip**: Errors due to incompatible NumPy shapes and dimensions are a very common issue when using libraries which build on the `ndarray` object, such as some data science libraries you'll learn about. But since you're now aware of the difference between dimension and shape in ndarrays, you'll be much quicker to stop and debug such issues later on!
---
NumPy also has a module called [`numpy.linalg`](https://numpy.org/doc/1.18/reference/routines.linalg.html), which is entirely dedicated to linear algebra operations. How amazing is that?
#### 2.3.2 Vector norm using [`numpy.linalg.norm()`](https://numpy.org/doc/1.18/reference/generated/numpy.linalg.norm.html)
[Just like most humans](https://vimeo.com/379750591), vectors always conform to the norm (recall section [1.2 Vector norm](#1.2-Vector-norm)):
$$\|\mathbf{u}\| = \sqrt{u_1^2 + u_2^2 + ... + u_m^2} = \sqrt{\sum_{i}^{m} u_i^2}$$
```
np.linalg.norm?
```
Ok, from the docstring we see that `np.linalg.norm` expects an array. Let's determine the norm of the vector $\begin{bmatrix}-1\\ 2\end{bmatrix}$, as we did in section [1.2 Vector norm](#1.2-Vector-norm), using the Pythagorean theorem:
```
a = np.array([[-1],
[2]])
np.linalg.norm(a)
```
Which is actually the square root of $5$:
```
# np.sqrt() computes the square root value of its input
np.sqrt(5)
```
#### 2.3.3 Vector operations (multiplication by scalars and addition)
Remember linear combinations being about multiplying vectors by scalars and adding them together?
$$c\; \mathbf{u} + d\; \mathbf{v}$$
```
# let's create two ndarrays representing 4-dimensional vectors
u = np.array([1, 0, 1, 1])
v = np.array([1, -2, 0, 1])
# print vectors to check your lucky numbers
print(f"u = {u}")
print(f"v = {v}\n")
```
We can use the [numeric operators](https://docs.python.org/3/library/stdtypes.html#numeric-types-int-float-complex) we already know with NumPy.
```
# multiplication by a scalar
-(1/2) * u
# addition
u + v
# linear combination
(-(1/2) * u) + (1 * v)
```
#### 2.3.4 Dot product
Finally, the [dot product](#Properties-of-the-dot-product):
$$\mathbf{u} \cdot \mathbf{v} = u_1 v_1 + ... + u_m v_m$$
Let's use [`numpy.dot`](https://numpy.org/doc/1.18/reference/generated/numpy.dot.html?highlight=dot%20product) to determine the dot product of two vectors!
```
# create two vectors using numpy arrays
u = np.array([-1, 2, 2]) # row vector, 1D array
v = np.array([-2, 1, 1]) # row vector, 1D array
# determine the dot product between vectors u and v
np.dot(u, v)
# create two vectors using numpy arrays
u = np.array([-1, 2, 2]).reshape((1, 3)) # row vector, 2D array
v = np.array([-2, 1, 1]) # row vector, 1D array
# determine the dot product between vectors u and v
np.dot(u, v)
# let's check those results "by hand", because we love linear algebra!
((-1) * (-2)) + (2 * 1) + (2 * 1)
```
---
Great job! You already know a lot about vectors, their meaning, how to work with them, the super important concept of linear independence, NumPy arrays...
This calls for a break before our next journey into the world of **matrices**!
Breathe in... Breathe out... Let the **magnitude** of all this knowledge flow through you. You're just like a vector now, pointing in the **direction** of success.
<img src="./media/breathe_in.gif"/>
---
## Wrapping up
What we've learned so far:
- what are vectors, their properties, linear combinations and linear independence;
- the `ndarray` and vector operations in NumPy.
```
Student: -"What's next?"
Instructor: -"Matrices!! Head over to Learning Notebook 2 - Matrices and NumPy!"
```
---
| github_jupyter |
```
# from google.colab import drive
# drive.mount('/content/drive')
# path = "/content/drive/MyDrive/Research/cods_comad_plots/sdc_task/mnist/"
import torch.nn as nn
import torch.nn.functional as F
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import torch
import torchvision
import torchvision.transforms as transforms
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
from matplotlib import pyplot as plt
import copy
# Ignore warnings
import warnings
warnings.filterwarnings("ignore")
torch.backends.cudnn.deterministic = True
torch.backends.cudnn.benchmark = False
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5), (0.5))])
trainset = torchvision.datasets.MNIST(root='./data', train=True, download=True, transform=transform)
testset = torchvision.datasets.MNIST(root='./data', train=False, download=True, transform=transform)
classes = ('zero','one','two','three','four','five','six','seven','eight','nine')
foreground_classes = {'zero','one'}
fg_used = '01'
fg1, fg2 = 0,1
all_classes = {'zero','one','two','three','four','five','six','seven','eight','nine'}
background_classes = all_classes - foreground_classes
background_classes
trainloader = torch.utils.data.DataLoader(trainset, batch_size=10, shuffle = False)
testloader = torch.utils.data.DataLoader(testset, batch_size=10, shuffle = False)
dataiter = iter(trainloader)
background_data=[]
background_label=[]
foreground_data=[]
foreground_label=[]
batch_size=10
for i in range(6000):
images, labels = dataiter.next()
for j in range(batch_size):
if(classes[labels[j]] in background_classes):
img = images[j].tolist()
background_data.append(img)
background_label.append(labels[j])
else:
img = images[j].tolist()
foreground_data.append(img)
foreground_label.append(labels[j])
foreground_data = torch.tensor(foreground_data)
foreground_label = torch.tensor(foreground_label)
background_data = torch.tensor(background_data)
background_label = torch.tensor(background_label)
def imshow(img):
img = img / 2 + 0.5 # unnormalize
npimg = img#.numpy()
plt.imshow(np.reshape(npimg, (28,28)))
plt.show()
foreground_data.shape, foreground_label.shape, background_data.shape, background_label.shape
val, idx = torch.max(background_data, dim=0, keepdims= True,)
# torch.abs(val)
mean_bg = torch.mean(background_data, dim=0, keepdims= True)
std_bg, _ = torch.max(background_data, dim=0, keepdims= True)
mean_bg.shape, std_bg.shape
foreground_data = (foreground_data - mean_bg) / std_bg
background_data = (background_data - mean_bg) / torch.abs(std_bg)
foreground_data.shape, foreground_label.shape, background_data.shape, background_label.shape
torch.sum(torch.isnan(foreground_data)), torch.sum(torch.isnan(background_data))
imshow(foreground_data[0])
imshow(background_data[0])
```
## generating CIN train and test data
```
m = 5
desired_num = 30000
np.random.seed(0)
bg_idx = np.random.randint(0,47335,m-1)
fg_idx = np.random.randint(0,12665)
bg_idx, fg_idx
for i in background_data[bg_idx]:
imshow(i)
imshow(torch.sum(background_data[bg_idx], axis = 0))
imshow(foreground_data[fg_idx])
tr_data = ( torch.sum(background_data[bg_idx], axis = 0) + foreground_data[fg_idx] )/m
tr_data.shape
imshow(tr_data)
foreground_label[fg_idx]
train_images =[] # list of mosaic images, each mosaic image is saved as list of 9 images
train_label=[] # label of mosaic image = foreground class present in that mosaic
for i in range(desired_num):
np.random.seed(i)
bg_idx = np.random.randint(0,47335,m-1)
fg_idx = np.random.randint(0,12665)
tr_data = ( torch.sum(background_data[bg_idx], axis = 0) + foreground_data[fg_idx] ) / m
label = (foreground_label[fg_idx].item())
train_images.append(tr_data)
train_label.append(label)
train_images = torch.stack(train_images)
train_images.shape, len(train_label)
imshow(train_images[0])
test_images =[] # list of mosaic images, each mosaic image is saved as list of 9 images
test_label=[] # label of mosaic image = foreground class present in that mosaic
for i in range(10000):
np.random.seed(i)
fg_idx = np.random.randint(0,12665)
tr_data = ( foreground_data[fg_idx] ) / m
label = (foreground_label[fg_idx].item())
test_images.append(tr_data)
test_label.append(label)
test_images = torch.stack(test_images)
test_images.shape, len(test_label)
imshow(test_images[0])
torch.sum(torch.isnan(train_images)), torch.sum(torch.isnan(test_images))
np.unique(train_label), np.unique(test_label)
```
## creating dataloader
```
class CIN_Dataset(Dataset):
"""CIN_Dataset dataset."""
def __init__(self, list_of_images, labels):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.image = list_of_images
self.label = labels
def __len__(self):
return len(self.label)
def __getitem__(self, idx):
return self.image[idx] , self.label[idx]
batch = 250
train_data = CIN_Dataset(train_images, train_label)
train_loader = DataLoader( train_data, batch_size= batch , shuffle=True)
test_data = CIN_Dataset( test_images , test_label)
test_loader = DataLoader( test_data, batch_size= batch , shuffle=False)
train_loader.dataset.image.shape, test_loader.dataset.image.shape
```
## model
```
class Classification(nn.Module):
def __init__(self):
super(Classification, self).__init__()
self.fc1 = nn.Linear(28*28, 2)
torch.nn.init.xavier_normal_(self.fc1.weight)
torch.nn.init.zeros_(self.fc1.bias)
def forward(self, x):
x = x.view(-1, 28*28)
x = self.fc1(x)
return x
```
## training
```
torch.manual_seed(12)
classify = Classification().double()
classify = classify.to("cuda")
import torch.optim as optim
criterion = nn.CrossEntropyLoss()
optimizer_classify = optim.Adam(classify.parameters(), lr=0.001 ) #, momentum=0.9)
correct = 0
total = 0
count = 0
flag = 1
with torch.no_grad():
for data in train_loader:
inputs, labels = data
inputs = inputs.double()
inputs, labels = inputs.to("cuda"),labels.to("cuda")
outputs = classify(inputs)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the %d train images: %f %%' % ( desired_num , 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
correct = 0
total = 0
count = 0
flag = 1
with torch.no_grad():
for data in test_loader:
inputs, labels = data
inputs = inputs.double()
inputs, labels = inputs.to("cuda"),labels.to("cuda")
outputs = classify(inputs)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the %d train images: %f %%' % ( 10000 , 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
nos_epochs = 200
tr_loss = []
for epoch in range(nos_epochs): # loop over the dataset multiple times
epoch_loss = []
cnt=0
iteration = desired_num // batch
running_loss = 0
#training data set
for i, data in enumerate(train_loader):
inputs, labels = data
inputs = inputs.double()
inputs, labels = inputs.to("cuda"),labels.to("cuda")
inputs = inputs.double()
# zero the parameter gradients
optimizer_classify.zero_grad()
outputs = classify(inputs)
_, predicted = torch.max(outputs.data, 1)
# print(outputs)
# print(outputs.shape,labels.shape , torch.argmax(outputs, dim=1))
loss = criterion(outputs, labels)
loss.backward()
optimizer_classify.step()
running_loss += loss.item()
mini = 20
if cnt % mini == mini-1: # print every 40 mini-batches
# print('[%d, %5d] loss: %.3f' %(epoch + 1, cnt + 1, running_loss / mini))
epoch_loss.append(running_loss/mini)
running_loss = 0.0
cnt=cnt+1
tr_loss.append(np.mean(epoch_loss))
if(np.mean(epoch_loss) <= 0.001):
break;
else:
print('[Epoch : %d] loss: %.3f' %(epoch + 1, np.mean(epoch_loss) ))
print('Finished Training')
plt.plot(tr_loss)
correct = 0
total = 0
count = 0
flag = 1
with torch.no_grad():
for data in train_loader:
inputs, labels = data
inputs = inputs.double()
inputs, labels = inputs.to("cuda"),labels.to("cuda")
outputs = classify(inputs)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the %d train images: %f %%' % ( desired_num , 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
correct = 0
total = 0
count = 0
flag = 1
with torch.no_grad():
for data in test_loader:
inputs, labels = data
inputs = inputs.double()
inputs, labels = inputs.to("cuda"),labels.to("cuda")
outputs = classify(inputs)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the %d train images: %f %%' % ( 10000 , 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
```
| github_jupyter |
## Notebook 1:
```
### Notebook 1
### Data set 1 (Viburnum)
### Language: Bash
### Data Location: NCBI SRA PRJNA299402 & PRJNA299407
%%bash
## make a new directory for this analysis
mkdir -p empirical_1/
mkdir -p empirical_1/halfrun
mkdir -p empirical_1/fullrun
## import Python libraries
import pandas as pd
import numpy as np
import ipyparallel
import urllib2
import glob
import os
```
### Download the sequence data
Sequence data for this study is archived on the NCBI sequence read archive (SRA). The data were run in two separate Illumina runs, but are combined under a single project id number.
+ Project SRA: SRP055977
+ Project number: PRJNA277574
+ Biosample numbers: SAMN03394519 -- SAMN03394561
+ Runs: SRR1915524 -- SRR1915566
+ The barcodes file is in the github repository for this [project]().
The library contains 95 samples. We uploaded the two demultiplexed samples for each individual separately, so each sample has 2 files. Below we examine just the first library (the "half" data set) and then both libraries combined (the "full" data set). We analyze on 64 samples since the remaining samples are replicate individuals within species that are part of a separate project.
You can download the data set using the script below:
```
%%bash
## get the data from Dryad
for run in $(seq 24 28);
do
wget -q -r -nH --cut-dirs=9 \
ftp://ftp-trace.ncbi.nlm.nih.gov/\
sra/sra-instant/reads/ByRun/sra/SRR/\
SRR191/SRR19155$run/SRR19155$run".sra";
done
%%bash
## convert sra files to fastq using fastq-dump tool
fastq-dump *.sra
## IPython code
## This reads in a table mapping sample names to SRA numbers
## that is hosted on github
## open table from github url
url = "https://raw.githubusercontent.com/"+\
"dereneaton/virentes/master/SraRunTable.txt"
intable = urllib2.urlopen(url)
## make name xfer dictionary
DF = pd.read_table(intable, sep="\t")
D = {DF.Run_s[i]:DF.Library_Name_s[i] for i in DF.index}
## change file names and move to fastq dir/
for fname in glob.glob("*.fastq"):
os.rename(fname, "analysis_pyrad/fastq/"+\
D[fname.replace(".fastq",".fq")])
```
### Create a set with reads concatenated from both technical replicates of each sample
```
%%bash
mkdir -p fastq_combined
## IPython code that makes a bash call w/ (!)
## get all the data from the two libraries and concatenate it
lib1tax = glob.glob("/home/deren/Documents/Vib_Lib1/fastq_Lib1/*.gz")
lib2tax = glob.glob("/home/deren/Documents/Vib_Lib1/fastq_Lib2/*.gz")
## names had to be modified to match
taxa = [i.split("/")[-1].split("_", 1)[1] for i in lib1tax]
for tax in taxa:
! cat /home/deren/Documents/Vib_Lib1/fastq_Lib1/Lib1_$tax \
/home/deren/Documents/Vib_Lib1/fastq_Lib2/Lib2_$tax \
> /home/deren/Documents/Vib_Lib1/fastq_combined/$tax
```
## Make a params file
```
%%bash
pyrad --version
%%bash
## create a new default params file
rm params.txt
pyrad -n
%%bash
## substitute new parameters into file
sed -i '/## 1. /c\empirical_1/halfrun ## 1. working directory ' params.txt
sed -i '/## 6. /c\TGCAG ## 6. cutters ' params.txt
sed -i '/## 7. /c\30 ## 7. N processors ' params.txt
sed -i '/## 9. /c\6 ## 9. NQual ' params.txt
sed -i '/## 10./c\.85 ## 10. clust threshold ' params.txt
sed -i '/## 12./c\4 ## 12. MinCov ' params.txt
sed -i '/## 13./c\10 ## 13. maxSH ' params.txt
sed -i '/## 14./c\empirical_1_half_m4 ## 14. output name ' params.txt
sed -i '/## 18./c\/home/deren/Documents/Vib_Lib1/fastq_Lib1/*.fastq ## 18. data location ' params.txt
sed -i '/## 29./c\2,2 ## 29. trim overhang ' params.txt
sed -i '/## 30./c\p,n,s,a ## 30. output formats ' params.txt
cat params.txt
%%bash
pyrad -p params.txt -s 234567 >> log.txt 2>&1
%%bash
sed -i '/## 12./c\2 ## 12. MinCov ' params.txt
sed -i '/## 14./c\empirical_1_half_m2 ## 14. output name ' params.txt
%%bash
pyrad -p params.txt -s 7 >> log.txt 2>&1
```
### Assemble the full data set
Added the 'a' option to output formats to build an ".alleles" file which will be used later for mrbayes/bucky analyses.
```
%%bash
## substitute new parameters into file
sed -i '/## 1. /c\empirical_1/fullrun ## 1. working directory ' params.txt
sed -i '/## 6. /c\TGCAG ## 6. cutters ' params.txt
sed -i '/## 7. /c\30 ## 7. N processors ' params.txt
sed -i '/## 9. /c\6 ## 9. NQual ' params.txt
sed -i '/## 10./c\.85 ## 10. clust threshold ' params.txt
sed -i '/## 12./c\4 ## 12. MinCov ' params.txt
sed -i '/## 13./c\10 ## 13. maxSH ' params.txt
sed -i '/## 14./c\empirical_1_full_m4 ## 14. output name ' params.txt
sed -i '/## 18./c\/home/deren/Documents/Vib_Lib1/fastq_combined/*.fastq ## 18. data location ' params.txt
sed -i '/## 29./c\2,2 ## 29. trim overhang ' params.txt
sed -i '/## 30./c\p,n,s,a ## 30. output formats ' params.txt
%%bash
pyrad -p params.txt -s 234567 >> log.txt 2>&1
%%bash
sed -i '/## 12./c\2 ## 12. MinCov ' params.txt
sed -i '/## 14./c\empirical_1_full_m2 ## 14. output name ' params.txt
%%bash
pyrad -p params.txt -s 7 >> log.txt 2>&1
```
## Results
We are interested in the relationship between the amount of input (raw) data between any two samples, the average coverage they recover when clustered together, and the phylogenetic distances separating samples.
### Raw data amounts (1 sequence lane)
The average number of raw reads per sample is 1.37M.
```
## read in the data
s2dat = pd.read_table("empirical_1/halfrun/stats/s2.rawedit.txt", header=0, nrows=66)
## print summary stats
print s2dat["passed.total"].describe()
## find which sample has the most raw data
maxraw = s2dat["passed.total"].max()
print "\nmost raw data in sample:"
print s2dat['sample '][s2dat['passed.total']==maxraw]
```
### Raw data amounts (2 sequence lanes)
The average nreads now is 2.74M
```
## read in the data
s2dat = pd.read_table("empirical_1/fullrun/stats/s2.rawedit.txt", header=0, nrows=66)
## print summary stats
print s2dat["passed.total"].describe()
## find which sample has the most raw data
maxraw = s2dat["passed.total"].max()
print "\nmost raw data in sample:"
print s2dat['sample '][s2dat['passed.total']==maxraw]
```
### Look at distributions of coverage
pyrad v.3.0.63 outputs depth information for each sample which I read in here and plot. First let's ask which sample has the highest depth of coverage. The std here is the std in means across samples. The std of depths within individuals is much higher.
```
## read in the s3 results
s3dat = pd.read_table("empirical_1/halfrun/stats/s3.clusters.txt", header=0, nrows=66)
## print summary stats
print "summary of means\n=================="
print s3dat['dpt.me'].describe()
## print summary stats
print "\nsummary of std\n=================="
print s3dat['dpt.sd'].describe()
## print summary stats
print "\nsummary of proportion lowdepth\n=================="
print pd.Series(1-s3dat['d>5.tot']/s3dat["total"]).describe()
## find which sample has the greatest depth of retained loci
maxdepth = s3dat["d>5.tot"].max()
print "\nhighest coverage in sample:"
print s3dat['taxa'][s3dat['d>5.tot']==maxdepth]
## read in the s3 results
s3dat = pd.read_table("empirical_1/fullrun/stats/s3.clusters.txt", header=0, nrows=66)
## print summary stats
print "summary of means\n=================="
print s3dat['dpt.me'].describe()
## print summary stats
print "\nsummary of std\n=================="
print s3dat['dpt.sd'].describe()
## print summary stats
print "\nsummary of proportion hidepth\n=================="
print pd.Series(1-s3dat['d>5.tot']/s3dat["total"]).describe()
## find which sample has the greatest depth of retained loci
max_hiprop = (s3dat["d>5.tot"]/s3dat["total"]).max()
print "\nhighest coverage in sample:"
print s3dat['taxa'][s3dat['d>5.tot']/s3dat["total"]==max_hiprop]
## print mean and std of coverage for the highest coverage sample
with open("empirical_1/fullrun/clust.85/lantanoides_D15_Beartown_2.depths", 'rb') as indat:
depths = np.array(indat.read().strip().split(","), dtype=int)
print depths.mean(), depths.std()
import toyplot
import toyplot.svg
import numpy as np
## read in the depth information for this sample
with open("empirical_1/fullrun/clust.85/lantanoides_D15_Beartown_2.depths", 'rb') as indat:
depths = np.array(indat.read().strip().split(","), dtype=int)
## make a barplot in Toyplot
canvas = toyplot.Canvas(width=350, height=300)
axes = canvas.axes(xlabel="Depth of coverage (N reads)",
ylabel="N loci",
label="dataset1/sample=sulcatum_D9_MEX_003")
## select the loci with depth > 5 (kept)
keeps = depths[depths>5]
## plot kept and discarded loci
edat = np.histogram(depths, range(30)) # density=True)
kdat = np.histogram(keeps, range(30)) #, density=True)
axes.bars(edat)
axes.bars(kdat)
#toyplot.svg.render(canvas, "empirical_1_full_depthplot.svg")
cat empirical_1/halfrun/stats/empirical_1_half_m4.stats
```
#### get average number of loci per sample
```
import numpy as np
indat = open("empirical_1/halfrun/stats/empirical_1_half_m4.stats").readlines()
counts = [int(i.strip().split("\t")[1]) for i in indat[8:73]]
print np.mean(counts)
print np.std(counts)
```
#### get average number of samples with data for a locus
```
import numpy as np
import itertools
indat = open("empirical_1/halfrun/stats/empirical_1_half_m4.stats").readlines()
counts = [i.strip().split("\t") for i in indat[81:142]]
#print counts
ntax = [int(i[0]) for i in counts]
ncounts = [int(i[1]) for i in counts]
tots = list(itertools.chain(*[[i]*n for i,n in zip(ntax, ncounts)]))
print np.mean(tots)
print np.std(tots)
cat empirical_1/fullrun/stats/empirical_1_full_m4.stats
import numpy as np
indat = open("empirical_1/fullrun/stats/empirical_1_full_m4.stats").readlines()
counts = [int(i.strip().split("\t")[1]) for i in indat[8:73]]
print np.mean(counts)
print np.std(counts)
import numpy as np
import itertools
indat = open("empirical_1/fullrun/stats/empirical_1_full_m4.stats").readlines()
counts = [i.strip().split("\t") for i in indat[81:140]]
#print counts
ntax = [int(i[0]) for i in counts]
ncounts = [int(i[1]) for i in counts]
tots = list(itertools.chain(*[[i]*n for i,n in zip(ntax, ncounts)]))
print np.mean(tots)
print np.std(tots)
```
## Infer an ML phylogeny
```
%%bash
## raxml argumement w/ ...
raxmlHPC-PTHREADS-AVX -f a -m GTRGAMMA -N 100 -x 12345 -p 12345 -T 35 \
-w /home/deren/Documents/RADmissing/empirical_1/halfrun \
-n empirical_1_halfrun -s empirical_1/halfrun/outfiles/empirical_1_half_m4.phy \
-o "Lib1_clemensiae_DRY6_PWS_2135"
%%bash
raxmlHPC-PTHREADS-AVX -f a -m GTRGAMMA -N 100 -x 12345 -p 12345 -T 35 \
-w /home/deren/Documents/RADmissing/empirical_1/fullrun \
-n empirical_1_fullrun -s empirical_1/fullrun/outfiles/empirical_1_full_m4.phy \
-o "clemensiae_DRY6_PWS_2135"
%%bash
head -n 20 empirical_1/halfrun/RAxML_info.empirical_1_half_m4
%%bash
head -n 20 empirical_1/fullrun/RAxML_info.empirical_1_full_m4
```
### Plot the tree in R using `ape`
```
%load_ext rpy2.ipython
%%R -w 600 -h 1000
library(ape)
tre_half <- read.tree("empirical_1/halfrun/RAxML_bipartitions.empirical_1_halfrun")
#rtre <- root(tre, "Lib1_clemensiae_DRY6_PWS_2135", resolve.root=T)
#rtre <- root(rtre, "Lib1_clemensiae_DRY6_PWS_2135", resolve.root=T)
ltre_half <- ladderize(tre_half)
plot(ltre_half, cex=0.8, edge.width=2)
nodelabels(ltre_half$node.label)
%%R -w 600 -h 1000
library(ape)
svg("outtree.svg", height=11, width=8)
tre_full <- read.tree("empirical_1/fullrun/RAxML_bipartitions.empirical_1_fullrun")
#rtre <- root(tre, "Lib1_clemensiae_DRY6_PWS_2135", resolve.root=T)
#rtre <- root(rtre, "Lib1_clemensiae_DRY6_PWS_2135", resolve.root=T)
ltre_full <- ladderize(tre_full)
plot(ltre_full, cex=0.8, edge.width=3)
#nodelabels(ltre_full$node.label)
dev.off()
```
## BUCKY -- write mrbayes nexus blocks for each locus
The functions `nexmake` and `subsample` are used to split the .loci file into individual nexus files for each locus within a new directory. Each nexus file is given a mrbayes command to run. Then we run the bucky tool `mbsum` to summarize the mrbayes output, and finally run `bucky` to infer concordance trees from the posterior distributions of trees across all loci.
Loci are selected on the basis that they have coverage across all tips of the selected subtree and that they contain at least 1 SNP.
```
def nexmake(taxadict, loc, nexdir, trim):
outloc = open(nexdir+"/"+str(loc)+".nex", 'w')
header = """
#NEXUS
begin data;
dimensions ntax={} nchar={};
format datatype=dna interleave=yes missing=N gap=-;
matrix
""".format(len(taxadict), len(taxadict.values()[0]))
outloc.write(header)
for tax, seq in taxadict.items():
outloc.write("{}{}{}\n"\
.format(tax[trim:trim+9],
" "*(10-len(tax[0:9])),
"".join(seq)))
mbstring = """
;
end;
begin mrbayes;
set autoclose=yes nowarn=yes;
lset nst=6 rates=gamma;
mcmc ngen=2200000 samplefreq=2000;
sump burnin=200000;
sumt burnin=200000;
end;
"""
outloc.write(mbstring)
outloc.close()
def unstruct(amb):
" returns bases from ambiguity code"
D = {"R":["G","A"],
"K":["G","T"],
"S":["G","C"],
"Y":["T","C"],
"W":["T","A"],
"M":["C","A"]}
if amb in D:
return D.get(amb)
else:
return [amb,amb]
def resolveambig(subseq):
N = []
for col in subseq:
N.append([unstruct(i)[np.random.binomial(1, 0.5)] for i in col])
return np.array(N)
def newPIS(seqsamp):
counts = [Counter(col) for col in seqsamp.T if not ("-" in col or "N" in col)]
pis = [i.most_common(2)[1][1] > 1 for i in counts if len(i.most_common(2))>1]
if sum(pis) >= 2:
return sum(pis)
else:
return 0
def parseloci(iloci, taxadict, nexdir, trim=0):
nloc = 0
## create subsampled data set
for loc in iloci:
## if all tip samples have data in this locus
names = [line.split()[0] for line in loc.split("\n")[:-1]]
## check that locus has required samples for each subtree
if all([i in names for i in taxadict.values()]):
seqs = np.array([list(line.split()[1]) for line in loc.split("\n")[:-1]])
seqsamp = seqs[[names.index(tax) for tax in taxadict.values()]]
seqsamp = resolveambig(seqsamp)
pis = newPIS(seqsamp)
if pis:
nloc += 1
## remove invariable columns given this subsampling
keep = []
for n, col in enumerate(seqsamp.T):
if all([i not in ["N","-"] for i in col]):
keep.append(n)
subseq = seqsamp.T[keep].T
## write to a nexus file
nexdict = dict(zip(taxadict.keys(), [i.tostring() for i in subseq]))
nexmake(nexdict, nloc, nexdir, trim)
print nloc, 'loci kept'
```
#### Modify line endings of loci string for easier parsing
```
def getloci(locifile):
## parse the loci file by new line characters
locifile = open(locifile)
lines = locifile.readlines()
## add "|" to end of lines that contain "|"
for idx in range(len(lines)):
if "|" in lines[idx]:
lines[idx] = lines[idx].strip()+"|\n"
## join lines back together into one large string
locistr = "".join(lines)
## break string into loci at the "|\n" character
loci = locistr.split("|\n")[:-1]
## how many loci?
print len(loci), "loci"
return loci
## run on both files
loci_full = getloci("empirical_1/fullrun/outfiles/empirical_1_full_m4.loci")
loci_half = getloci("empirical_1/halfrun/outfiles/empirical_1_half_m4.loci")
```
### Make nexus files
```
parseloci(loci_full[:], deep_dict_f, "deep_dict_full", 0)
parseloci(loci_half[:], deep_dict_h, "deep_dict_half", 0)
#parseloci(loci[:], shallow_dict, "shallow_dict", 0)
## create a parallel client
ipclient = ipyparallel.Client()
lview = ipclient.load_balanced_view()
## call function across all engines
def mrbayes(infile):
import subprocess
cmd = "mb %s" % infile
subprocess.check_call(cmd, shell=True)
## submit all nexus files to run mb
allnex = glob.glob("deep_dict_full/*.nex")
for nex in allnex:
lview.apply(mrbayes, nex)
ipclient.wait_interactive()
```
### Summarize posteriors with `mbsum`
```
def mbsum(nexdir, nloci):
import subprocess
## combine trees from the two replicate runs
for n in range(1, nloci+1):
cmd = "mbsum -n 101 -o {}{}.in {}{}.nex.run1.t {}{}.nex.run2.t".\
format(nexdir, n, nexdir, n, nexdir, n)
subprocess.check_call(cmd, shell=True)
```
### Run Bucky to infer concordance factors
```
import os
import numpy as np
from collections import Counter
def subsample(infile, requires, outgroup, nexdir, trim):
""" sample n taxa from infile to create nex file"""
## counter
loc = 0
## create output directory
if not os.path.exists(nexdir):
os.mkdir(nexdir)
## input .alleles file
loci = open(infile, 'r').read().strip().split("//")
## create a dictionary of {names:seqs}
for locus in xrange(len(loci)):
locnex = [""]*len(requires)
for line in loci[locus].strip().split("\n"):
tax = line.split()[0]
seq = line.split()[-1]
if ">" in tax:
if tax in requires:
locnex[requires.index(tax)] = seq
## if all tips
if len([i for i in locnex if i]) == len(requires):
## if locus is variable
## count how many times each char occurs in each site
ccs = [Counter(i) for i in np.array([list(i) for i in locnex]).T]
## remove N and - characters and the first most occurring base
for i in ccs:
del i['-']
del i['N']
if i:
del i[i.most_common()[0][0]]
## is anything left occuring more than once (minor allele=ma)?
ma = max([max(i.values()) if i else 0 for i in ccs])
if ma > 1:
nexmake(requires, locnex, loc, outgroup, nexdir, trim)
loc += 1
return loc
```
### Subtree 1 (Oreinodentinus) (full data set)
```
## inputs
requires = [">triphyllum_D13_PWS_1783_0",
">jamesonii_D12_PWS_1636_0",
">sulcatum_D9_MEX_003_0",
">acutifolium_DRY3_MEX_006_0",
">dentatum_ELS4_0",
">recognitum_AA_1471_83B_0"]
outgroup = ""
infile = "empirical_1/fullrun/outfiles/empirical_1_full_m4.alleles"
nexdir = "nex_files1"
## run function
nloci = subsample(infile, requires, outgroup, nexdir, trim=1)
print nloci
```
### Subtree 1 (Oreinodentinus) (half data set)
```
## inputs
requires = [">Lib1_triphyllum_D13_PWS_1783_0",
">Lib1_jamesonii_D12_PWS_1636_0",
">Lib1_sulcatum_D9_MEX_003_0",
">Lib1_acutifolium_DRY3_MEX_006_0",
">Lib1_dentatum_ELS4_0",
">Lib1_recognitum_AA_1471_83B_0"]
outgroup = ""
infile = "empirical_1/halfrun/outfiles/empirical_1_half_m4.alleles"
nexdir = "nex_files2"
## run function
nloci = subsample(infile, requires, outgroup, nexdir, trim=6)
print nloci
```
### Subtree 2 (Urceolata) (full data set)
```
## inputs
requires = [">clemensiae_DRY6_PWS_2135_0",
">tinus_D33_WC_277_0",
">taiwanianum_TW1_KFC_1952_0",
">lantanoides_D15_Beartown_2_0",
">amplificatum_D3_SAN_156003_0",
">lutescens_D35_PWS_2077_0",
">lentago_ELS85_0",
">dentatum_ELS4_0"]
outgroup = ""
infile = "empirical_1/fullrun/outfiles/empirical_1_full_m4.alleles"
nexdir = "nex_files5"
## run function
nloci = subsample(infile, requires, outgroup, nexdir, trim=1)
print nloci
```
### Subtree 2 (Urceolata) (half data set)
```
## inputs
requires = [">Lib1_clemensiae_DRY6_PWS_2135_0",
">Lib1_tinus_D33_WC_277_0",
">Lib1_taiwanianum_TW1_KFC_1952_0",
">Lib1_lantanoides_D15_Beartown_2_0",
">Lib1_amplificatum_D3_SAN_156003_0",
">Lib1_lutescens_D35_PWS_2077_0",
">Lib1_lentago_ELS85_0",
">Lib1_dentatum_ELS4_0"]
outgroup = ""
infile = "empirical_1/halfrun/outfiles/empirical_1_half_m4.alleles"
nexdir = "nex_files6"
## run function
nloci = subsample(infile, requires, outgroup, nexdir, trim=6)
print nloci
```
### Run mrbayes on all nex files
```
import ipyparallel
import subprocess
import glob
## create a parallel client
ipclient = ipyparallel.Client()
lview = ipclient.load_balanced_view()
## call function across all engines
def mrbayes(infile):
import subprocess
cmd = "mb %s" % infile
subprocess.check_call(cmd, shell=True)
## run on the full data set
res = lview.map_async(mrbayes, glob.glob("nex_files1/*"))
_ = res.get()
## run on the half data set
res = lview.map_async(mrbayes, glob.glob("nex_files2/*"))
_ = res.get()
## run on the half data set
res = lview.map_async(mrbayes, glob.glob("nex_files3/*"))
_ = res.get()
## run on the half data set
res = lview.map_async(mrbayes, glob.glob("nex_files4/*"))
_ = res.get()
## run on the half data set
res = lview.map_async(mrbayes, glob.glob("nex_files5/*"))
_ = res.get()
## run on the half data set
res = lview.map_async(mrbayes, glob.glob("nex_files6/*"))
_ = res.get()
```
### Run mbsum to summarize the results
```
import os
import subprocess
def mbsum(nexdir, nloci):
## create dir for bucky input files
insdir = os.path.join(nexdir, "ins")
if not os.path.exists(insdir):
os.mkdir(insdir)
## combine trees from the two replicate runs
for n in range(nloci):
cmd = "mbsum -n 101 -o {}/{}.in {}{}.nex.run1.t {}{}.nex.run2.t".\
format(insdir, n, nexdir, n, nexdir, n)
subprocess.check_call(cmd, shell=True)
#mbsum("nex_files1/", 3300)
#mbsum("nex_files2/", 364)
#mbsum("nex_files3/", 1692)
#mbsum("nex_files4/", 169)
mbsum("nex_files5/", 1203)
mbsum("nex_files6/", 106)
```
### Run Bucky
```
args = []
for insdir in ["nex_files5/ins", "nex_files6/ins"]:
## independence test
args.append("bucky --use-independence-prior -k 4 -n 500000 \
-o {}/BUCKY.ind {}/*.in".format(insdir, insdir))
## alpha at three levels
for alpha in [0.1, 1, 10]:
args.append("bucky -a {} -k 4 -n 500000 -c 4 -o {}/BUCKY.{} {}/*.in".\
format(alpha, insdir, alpha, insdir))
def bucky(arg):
import subprocess
subprocess.check_call(arg, shell=True)
return arg
res = lview.map_async(bucky, args)
res.get()
```
#### Cleanup
```
del lbview
ipclient.close()
```
### check out the results
```
%%bash
head -n 40 nex_files1/ins/BUCKY.0.1.concordance
%%bash
head -n 40 nex_files1/ins/BUCKY.1.concordance
%%bash
head -n 40 nex_files2/ins/BUCKY.1.concordance
! head -n 45 nex_files3/ins/BUCKY.0.1.concordance
```
### FINAL BUCKY RESULTS (DEEP_SCALE)
```
! head -n 45 nex_files4/ins/BUCKY.0.1.concordance
! head -n 45 nex_files5/ins/BUCKY.0.1.concordance
! head -n 45 nex_files6/ins/BUCKY.0.1.concordance
```
### Get missing data percentatge for m2 data sets
For this I start raxml to get the info and then quit. Kind of lazy but simpler than calculating it myself.
```
%%bash
## raxml argumement w/ ...
raxmlHPC-PTHREADS-AVX -f a -m GTRGAMMA -N 100 -x 12345 -p 12345 -T 20 \
-w /home/deren/Documents/RADmissing/empirical_1/fullrun \
-n empirical_1_full_m2 -s empirical_1/fullrun/outfiles/empirical_1_m2.phy
%%bash
head -n 20 empirical_1/fullrun/RAxML_info.empirical_1_full_m2
```
### Get average phylo dist (GTRgamma dist)
```
%%R
mean(cophenetic.phylo(ltre))
```
| github_jupyter |
# Class Session 10 - Date Hubs and Party Hubs
## Comparing the histograms of local clustering coefficients of date hubs and party hubs
In this class, we will analyze the protein-protein interaction network for two classes of yeast proteins, "date hubs" and "party hubs" as defined by Han et al. in their 2004 study of protein-interaction networks and gene expression (Han et al., Nature, v430, p88, 2004). The authors of that study claimed that there is no difference in the local clustering density, between "date hubs" and "party hubs". We will put this to the test. We for each of the "date hub" and "party hub" proteins, we will compute its local clustering coefficient (*C<sub>i</sub>*) in the protein-protein interaction network. We will then histogram the *C<sub>i</sub>*) values for the two sets of hubs, so that we can compare the distributions of local clustering coefficients for "date hubs" and "party hubs". We will use a statistical test (Kolmogorov-Smirnov) to compare the two distributions of *C<sub>i</sub>* values.
To get started, we load the python packages that we will require:
```
import igraph
import numpy
import pandas
import matplotlib.pyplot
```
The `scipy` package doesn't import the `stats` module automatically. so we have to force its import using `from`
Next, we'll load the file of hub types `shared/han_hub_data.txt` (which is a two-column TSV file in which the first column is the protein name and the second column contains the string `date` or `party` for each row; the first row of the file contains the column headers), using our old friend `pandas.read_csv`. This file has a header so pass `header=0` to `read_csv`.
Let's take a peek at the structure of the `hub_data` data frame, using `head` and `shape`. Here's what it should look like:
Next, let's load the file of yeat protein-protein interaction network edges `shared/han_network_edges.txt` (which is a two-column file, with first column is the first protein in the interacting pair, and the second column is the second protein in the interacting pair).This file has a header so pass `header=0` to `read_csv`.
Let's take a peek at the data frame `edge_df`, using `head` and `shape`:
make an undirected igraph `Graph` from the edgelist data; show summary data on the graph as a sanity-check
It will be convenient to let `igraph` compute the local clustering coefficients. So, we'll want to make an undirected igraph `igraph.Graph` object from the edgelist data, using our old friend `igraph.Graph.TupleList`:
```
ppi_graph =
```
As always, we'll use `igraph.Graph.summary` to sanity check the `Graph` object:
Generate a list of the names of the proteins in the order of the proteins' corresponding vertices in the igraph `Graph` object, using the method `vs` and then indexing into the resulting object using `["name"]`.
```
graph_vertices =
```
Let's take a look at the first ten proteins in the resulting list
Make a dataframe containing the protein names (as column "Protein") using `pandas.DataFrame` and `pandas.Series`. Set the column name to `Protein` using the `columns` attribute on the data frame. Define a new column `order` that will contain the list of IDs in `graph_vertices_df.index`.
```
graph_vertices_df =
graph_vertices_df.columns =
graph_vertices_df["order"] =
```
Let's take a peek at this data frame:
```
graph_vertices_df.head()
```
Let's use the `pandas.DataFrame.merge` method on the `graph_vertices_df` object to pull in the hub type (date or party) for vertices that are hubs, by passing `hub_data` to `merge`. Don't forget to specify `how='outer'` and `on="Protein"`:
```
graph_vertices_df_merged =
graph_vertices_df_merged =
```
Having merged the hub type information into `graph_vertices_df`, let's take a peek at it using `head` and `shape`:
NOTE: a `NaN` in the `HubType` column means that the protein is not a hub.
Let's pull out the `HubType` column as a numpy array, using column indexing (`["HubType"]`) and then `values.tolist()`:
```
vertex_types_np =
```
Let's take a peek at this `numpy.array` that we have just created:
```
vertex_types_np
```
Use `numpy.where` in order to find the index numbers of the proteins that are "date hubs" and that are "party hubs":
```
date_hub_inds =
party_hub_inds =
```
Use the `igraph.Graph.transitivity_local_undirected` function in igraph to compute the local clustering coefficients for every vertex in the graph. Make a `numpy.array` from the resulting list of Ci values:
```
ci_values =
ci_values_np =
```
Let's take a peek at the `ci_values_np` array that you have just created. What are the `nan` values, and what do they signify? Is this normal?
Make a `numpy.array` of the Ci values of the date hubs (`ci_values_date_hubs`) and the Ci values of the party hubs (`ci_values_party_hubs`)
```
ci_values_date_hubs =
ci_values_party_hubs =
```
Plot the histograms of the local clustering coefficients of the "date hubs" and the "party hubs". Use `matplotlib.pyplot.hist`. Use alpha blending so that the two overlapping histograms can be plotted together.
Do these histograms look the same to you? Let's test it with a Kolmogorov-Smirnov test, using the function `scipy.stats.ks_2samp`.
| github_jupyter |
```
import autoreg
import GPy
import numpy as np
from matplotlib import pyplot as plt
from __future__ import print_function
%matplotlib inline
from autoreg.benchmark import tasks
# Function to compute root mean square error:
def comp_RMSE(a,b):
return np.sqrt(np.square(a.flatten()-b.flatten()).mean())
# Define class for normalization
class Normalize(object):
def __init__(self, data, name, norm_name):
self.data_mean = data.mean(axis=0)
self.data_std = data.std(axis=0)
self.normalization_computed = True
setattr(self, name, data)
setattr(self, norm_name, (data-self.data_mean) / self.data_std )
def normalize(self, data, name, norm_name):
if hasattr(self,norm_name):
raise ValueError("This normalization name already exist, choose another one")
setattr(self, name, data )
setattr(self, norm_name, (data-self.data_mean) / self.data_std )
def denormalize(self, data):
return data*self.data_std + self.data_mean
trainned_models_folder_name = "/Users/grigoral/work/code/RGP/examples/identif_trainded"
task_name = 'Actuator'
# task names:
# Actuator, Ballbeam, Drive, Gas_furnace, Flutter, Dryer, Tank,
# IdentificationExample1..5
task = getattr( tasks, task_name)
task = task()
task.load_data()
print("Data OUT train shape: ", task.data_out_train.shape)
print("Data IN train shape: ", task.data_in_train.shape)
print("Data OUT test shape: ", task.data_out_test.shape)
print("Data IN test shape: ", task.data_in_test.shape)
```
### Normalize training and test data:
```
normalize = False
in_data = Normalize(task.data_in_train,'in_train','in_train_norm' )
out_data = Normalize(task.data_out_train,'out_train','out_train_norm' )
in_data.normalize(task.data_in_test, 'in_test','in_test_norm')
out_data.normalize(task.data_out_test, 'out_test','out_test_norm')
if normalize:
out_train = out_data.out_train_norm #out_data.out_train
in_train = in_data.in_train_norm # in_data.in_train
out_test = out_data.out_test_norm #out_data.out_test
in_test = in_data.in_test_norm #in_data.in_test
else:
out_train = out_data.out_train #out_data.out_train
in_train = in_data.in_train # in_data.in_train
out_test = out_data.out_test #out_data.out_test
in_test = in_data.in_test #in_data.in_test
print("Training OUT mean: ", out_train.mean(0));
print("Training OUT std: ", out_train.std(0))
print("")
print("Test OUT mean: ", out_test.mean(0));
print("Test OUT std: ", out_test.std(0))
print("")
print("Training IN mean: ", in_train.mean(0));
print("Training IN std: ", in_train.std(0))
print("")
print("Test IN mean: ", in_test.mean(0));
print("Test IN std: ", in_test.std(0))
```
### Plot training and test data:
```
# Plot training:
fig1 = plt.figure(1,figsize=(20,8))
fig1.suptitle('Training data')
ax1 = plt.subplot(1,2,1)
ax1.plot(out_train)
ax1.set_title('Data OUT training')
ax2 = plt.subplot(1,2,2)
ax2.plot(in_train)
ax2.set_title('Data IN training')
fig2 = plt.figure(2,figsize=(20,8))
fig2.suptitle('Test data')
ax3 = plt.subplot(1,2,1)
ax3.plot(out_test)
ax3.set_title('Data OUT test')
ax4 = plt.subplot(1,2,2)
ax4.plot(in_test)
ax4.set_title('Data IN test')
del ax1, ax2, ax3, ax4
```
### Model definition:
```
Q = 100 # 200 # Inducing points num
win_in = task.win_in # 20
win_out = task.win_out # 20
use_controls = True
back_cstr = False
inference_method = None
# 1 layer:
wins = [0, win_out] # 0-th is output layer
nDims = [out_train.shape[1],1]
# 2 layers:
# wins = [0, win_out, win_out]
# nDims = [out_train.shape[1],1,1]
MLP_dims = [300,200]
print("Input window: ", win_in)
print("Output window: ", win_out)
m = autoreg.DeepAutoreg(wins, out_train, U=in_train, U_win=win_in,
num_inducing=Q, back_cstr=back_cstr, MLP_dims=MLP_dims, nDims=nDims,
init='Y', # how to initialize hidden states means
X_variance=0.05, # how to initialize hidden states variances
#inference_method=inference_method, # Inference method
# 1 layer:
kernels=[GPy.kern.RBF(win_out,ARD=True,inv_l=True),
GPy.kern.RBF(win_in + win_out,ARD=True,inv_l=True)] )
# 2 layers:
# kernels=[GPy.kern.RBF(win_out,ARD=True,inv_l=True),
# GPy.kern.RBF(win_out+win_out,ARD=True,inv_l=True),
# GPy.kern.RBF(win_out+win_in,ARD=True,inv_l=True)])
#m = autoreg.DeepAutoreg([0,win_out],out_train, U=in_train, U_win=win_in,X_variance=0.01,
# num_inducing=50)
# pattern for model name: #task_name, inf_meth=?, wins=layers, Q = ?, backcstr=?,MLP_dims=?, nDims=
model_file_name = '%s--inf_meth=%s--wins=%s--Q=%i--backcstr=%i--nDims=%s' % (task.name,
'reg' if inference_method is None else inference_method, str(wins), Q, back_cstr, str(nDims))
if back_cstr == True:
model_file_name += '--MLP_dims=%s' % (MLP_dims,)
print('Model file name: ', model_file_name)
print(m)
```
### Model initialization:
```
# Here layer numbers are different than in initialization. 0-th layer is the top one
for i in range(m.nLayers):
m.layers[i].kern.variance = 0.1
m.layers[i].kern.inv_l[:] = np.mean( 1./((m.layers[i].X.mean.values.max(0)-m.layers[i].X.mean.values.min(0))/np.sqrt(2.)) )
m.layers[i].likelihood.variance[:] = 0.01*out_train.var()
m.layers[i].kern.variance.fix(warning=False)
m.layers[i].likelihood.fix(warning=False)
print(m)
print(m.layer_1.kern.inv_l)
print(m.layer_0.kern.inv_l)
print( np.mean(1./((m.layer_1.X.mean.values.max(0)-m.layer_1.X.mean.values.min(0))/np.sqrt(2.))) )
# Plot initialization of hidden layer:
def plot_hidden_states(fig_no, layer, layer_start_point=None, layer_end_point=None,
data_start_point=None, data_end_point=None):
if layer_start_point is None: layer_start_point=0;
if layer_end_point is None: layer_end_point = len(layer.mean)
if data_start_point is None: data_start_point=0;
if data_end_point is None: layer_end_point = len(out_train)
data = out_train[data_start_point:data_end_point]
layer_means = layer.mean[layer_start_point:layer_end_point]
layer_vars = layer.variance[layer_start_point:layer_end_point]
fig4 = plt.figure(fig_no,figsize=(10,8))
ax1 = plt.subplot(1,1,1)
fig4.suptitle('Hidden layer plotting')
ax1.plot(out_train[data_start_point:data_end_point], label="Orig data Train_out", color = 'b')
ax1.plot( layer_means, label = 'pred mean', color = 'r' )
ax1.plot( layer_means +\
2*np.sqrt( layer_vars ), label = 'pred var', color='r', linestyle='--' )
ax1.plot( layer_means -\
2*np.sqrt( layer_vars ), label = 'pred var', color='r', linestyle='--' )
ax1.legend(loc=4)
ax1.set_title('Hidden layer vs Training data')
del ax1
plot_hidden_states(5,m.layer_1.qX_0)
#plot_hidden_states(6,m.layer_2.qX_0)
```
### Model training:
```
#init_runs = 50 if out_train.shape[0]<1000 else 100
init_runs = 50
print("Init runs: ", init_runs)
m.optimize('lbfgs',messages=1,max_iters=init_runs)
for i in range(m.nLayers):
m.layers[i].kern.variance.constrain_positive(warning=False)
m.layers[i].likelihood.constrain_positive(warning=False)
m.optimize('lbfgs',messages=1,max_iters=10000)
print(m)
#m.optimize('scg',messages=1,max_iters=1000)
print(m)
```
### Look at trained parameters
```
if hasattr(m, 'layer_1'):
print("Layer 1: ")
print("States means (min and max), shapes: ", m.layer_1.qX_0.mean.min(),
m.layer_1.qX_0.mean.max(), m.layer_1.qX_0.mean.shape)
print("States variances (min and max), shapes: ", m.layer_1.qX_0.variance.min(),
m.layer_1.qX_0.variance.max(), m.layer_1.qX_0.mean.shape)
print("Inverse langthscales (min and max), shapes: ", m.layer_1.rbf.inv_lengthscale.min(),
m.layer_1.rbf.inv_lengthscale.max(), m.layer_1.rbf.inv_lengthscale.shape )
if hasattr(m, 'layer_0'):
print("")
print("Layer 0 (output): ")
print("Inverse langthscales (min and max), shapes: ", m.layer_0.rbf.inv_lengthscale.min(),
m.layer_0.rbf.inv_lengthscale.max(), m.layer_0.rbf.inv_lengthscale.shape )
print(m.layer_0.rbf.inv_lengthscale)
print(m.layer_1.rbf.inv_lengthscale)
```
### Analyze and plot model on test data:
```
# Free-run on the train data
# initialize to last part of trained latent states
#init_Xs = [None, m.layer_1.qX_0[0:win_out]] # init_Xs for train prediction
# initialize to zeros
init_Xs = None
predictions_train = m.freerun(init_Xs = init_Xs, U=in_train, m_match=True)
# initialize to last part of trainig latent states
#init_Xs = [None, m.layer_1.qX_0[-win_out:] ] # init_Xs for test prediction
#U_test = np.vstack( (in_train[-win_in:], in_test) )
# initialize to zeros
init_Xs = None
U_test = in_test
# Free-run on the test data
predictions_test = m.freerun(init_Xs = init_Xs, U=U_test, m_match=True)
del init_Xs, U_test
# Plot predictions
def plot_predictions(fig_no,posterior_train, posterior_test=None, layer_no = None):
"""
Plots the output data along with posterior of the layer.
Used for plotting the hidden states or
layer_no: int or Normal posterior
plot states of this layer (0-th is output). There is also some logic about compting
the MSE, and aligning with actual data.
"""
if layer_no is None: #default
layer_no = 1
if posterior_test is None:
no_test_data = True
else:
no_test_data = False
if isinstance(posterior_train, list):
layer_in_list = len(predictions_train)-1-layer_no # standard layer no (like in printing the model)
predictions_train_layer = predictions_train[layer_in_list]
else:
predictions_train_layer = posterior_train
if not no_test_data:
if isinstance(posterior_test, list):
predictions_test_layer = predictions_test[layer_in_list]
else:
predictions_test_layer = posterior_test
# Aligning the data ->
# training of test data can be longer than leyer data because of the initial window.
if out_train.shape[0] > predictions_train_layer.mean.shape[0]:
out_train_tmp = out_train[win_out:]
else:
out_train_tmp = out_train
if not no_test_data:
if out_test.shape[0] > predictions_test_layer.mean.shape[0]:
out_test_tmp = out_test[win_out:]
else:
out_test_tmp = out_test
# Aligning the data <-
if layer_no == 0:
# Not anymore! Compute RMSE ignoring first output values of length "win_out"
train_rmse = [comp_RMSE(predictions_train_layer.mean,
out_train_tmp)]
print("Train overall RMSE: ", str(train_rmse))
if not no_test_data:
# Compute RMSE ignoring first output values of length "win_out"
test_rmse = [comp_RMSE(predictions_test_layer.mean,
out_test_tmp)]
print("Test overall RMSE: ", str(test_rmse))
# Plot predictions:
if not no_test_data:
fig5 = plt.figure(10,figsize=(20,8))
else:
fig5 = plt.figure(10,figsize=(10,8))
fig5.suptitle('Predictions on Training and Test data')
if not no_test_data:
ax1 = plt.subplot(1,2,1)
else:
ax1 = plt.subplot(1,1,1)
ax1.plot(out_train_tmp, label="Train_out", color = 'b')
ax1.plot( predictions_train_layer.mean, label = 'pred mean', color = 'r' )
ax1.plot( predictions_train_layer.mean +\
2*np.sqrt( predictions_train_layer.variance ), label = 'pred var', color='r', linestyle='--' )
ax1.plot( predictions_train_layer.mean -\
2*np.sqrt( predictions_train_layer.variance ), label = 'pred var', color='r', linestyle='--' )
ax1.legend(loc=4)
ax1.set_title('Predictions on Train')
if not no_test_data:
ax2 = plt.subplot(1,2,2)
ax2.plot(out_test_tmp, label="Test_out", color = 'b')
ax2.plot( predictions_test_layer.mean, label = 'pred mean', color = 'r' )
#ax2.plot( predictions_test_layer.mean +\
# 2*np.sqrt( predictions_test_layer.variance ), label = 'pred var', color='r', linestyle='--' )
#ax2.plot( predictions_test_layer.mean -\
# 2*np.sqrt( predictions_test_layer.variance ), label = 'pred var', color='r', linestyle='--' )
ax2.legend(loc=4)
ax2.set_title('Predictions on Test')
del ax2
del ax1
plot_predictions(7,predictions_train, predictions_test , layer_no = 0)
plot_predictions(7,predictions_train, None , layer_no = 1)
comp_RMSE(np.zeros( (len(out_train[20:]),1) ), out_train[20:] )
out_train[20:].mean(0)
plot_hidden_states(8,m.layer_1.qX_0)
#plot_hidden_states(9,m.layer_2.qX_0)
```
| github_jupyter |
[](http://rpi.analyticsdojo.com)
<center><h1>Train Test Splits with Python</h1></center>
<center><h3><a href = 'http://rpi.analyticsdojo.com'>rpi.analyticsdojo.com</a></h3></center>
```
#Let's get rid of some imports
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
```
Training and Testing Data
=====================================
To evaluate how well our supervised models generalize, we can split our data into a training and a test set.
- It is common to see `X` as the feature of independent variables and `y` as the dv or label.
```
from sklearn.datasets import load_iris
#Iris is available from the sklearn package
iris = load_iris()
X, y = iris.data, iris.target
X
```
Thinking about how machine learning is normally performed, the idea of a train/test split makes sense. Real world systems train on the data they have, and as other data comes in (from customers, sensors, or other sources) the classifier that was trained must predict on fundamentally *new* data. We can simulate this during training using a train/test split - the test data is a simulation of "future data" which will come into the system during production.
Specifically for iris, the 150 labels in iris are sorted, which means that if we split the data using a proportional split, this will result in fudamentally altered class distributions. For instance, if we'd perform a common 2/3 training data and 1/3 test data split, our training dataset will only consists of flower classes 0 and 1 (Setosa and Versicolor), and our test set will only contain samples with class label 2 (Virginica flowers).
Under the assumption that all samples are independent of each other (in contrast time series data), we want to **randomly shuffle the dataset before we split the dataset** as illustrated above.
```
y
```
### Shuffling Dataset
- Now we need to split the data into training and testing.
- Luckily, this is a common pattern in machine learning and scikit-learn has a pre-built function to split data into training and testing sets for you.
- Here, we use 50% of the data as training, and 50% testing.
- 80% and 20% is another common split, but there are no hard and fast rules.
- The most important thing is to fairly evaluate your system on data it *has not* seen during training!
```
#Import Module
from sklearn.model_selection import train_test_split
train_X, test_X, train_y, test_y = train_test_split(X, y,
train_size=0.5,
test_size=0.5,
random_state=122)
print("Labels for training and testing data")
print(train_y)
print(test_y)
```
---
### Stratified Splitting
- Especially for relatively small datasets, it's better to stratify the split.
- Stratification means that we maintain the original class proportion of the dataset in the test and training sets.
- For example, after we randomly split the dataset as shown in the previous code example, we have the following class proportions in percent:
```
print('All:', np.bincount(y) / float(len(y)) * 100.0)
print('Training:', np.bincount(train_y) / float(len(train_y)) * 100.0)
print('Test:', np.bincount(test_y) / float(len(test_y)) * 100.0)
```
So, in order to stratify the split, we can pass the label array as an additional option to the `train_test_split` function:
```
train_X, test_X, train_y, test_y = train_test_split(X, y,
train_size=0.5,
test_size=0.5,
random_state=123,
stratify=y)
print('All:', np.bincount(y) / float(len(y)) * 100.0)
print('Training:', np.bincount(train_y) / float(len(train_y)) * 100.0)
print('Test:', np.bincount(test_y) / float(len(test_y)) * 100.0)
```
---
| github_jupyter |
# MNIST SVD Classification
Follows Chapter 11 of Matrix Methods in Data Mining and Pattern Recognition by Lars Elden,
with added dimensionality reduction visualization
#### Author: Daniel Yan
#### Email: daniel.yan@vanderbilt.edu
```
from keras.datasets import mnist
from matplotlib import pyplot as plt
import numpy as np
```
# Load Data
Load in Keras dataset
```
# Load in mnist
(x_train, y_train), (x_test, y_test) = mnist.load_data()
# Reshape to each image to a row vector and column vector
x_train_rowvector = np.reshape(x_train, (-1, 28*28))
x_train_colvector = np.copy(x_train_rowvector).T
x_test_rowvector = np.reshape(x_test, (-1, 28*28))
x_test_colvector = np.copy(x_test_rowvector).T
# Take small sample of 2000 training images
x_train_colvector_sample2000 = x_train_colvector[:, :2000]
y_train_sample2000 = y_train[:2000]
# Take small sample of 200 testing images
x_test_colvector_sample200 = x_test_colvector[:, :200]
y_test_sample200 = y_test[:200]
```
# Visualize Examples
```
# Visualize a few samples
for i in range(5):
print("Label: ", y_train[i])
image = x_train_colvector[:, i]
plt.imshow(image.reshape(28, 28), cmap="Greys")
plt.show()
plt.close()
```
# PCA Visualization
Credits: https://towardsdatascience.com/pca-and-svd-explained-with-numpy-5d13b0d2a4d8
```
# Calculate the covariance matrix
covariance = np.cov(x_train_colvector_sample2000)
# Calculate the eigenvalues and the eigenvectors for the covariance matrix
eigenvalues, eigenvectors = np.linalg.eig(covariance)
# Get the real part of the eigenvalues and eigenvectors only
eigenvalues = np.real(eigenvalues)
eigenvectors = np.real(eigenvectors)
# Project original data onto eigenvectors
pca = np.dot(x_train_colvector_sample2000.T, eigenvectors)
# Get only the first two columns for the first two principal components
pca = pca[:, 0:2]
```
Sort by label
```
pca_list= [0] * 10
y_list = [0] * 10
for i in range(10):
pca_list[i] = (pca[y_train_sample2000 == i])
y_list[i] = (y_train_sample2000[y_train_sample2000 == i])
```
Plot each label separately on graph
```
COLORS = ["red", "blue", "green", "yellow", "darkviolet",
"maroon", "greenyellow", "hotpink", "black", "cyan"]
fig, ax = plt.subplots()
for i in range(10):
# Get the pca array corresponding to the current label
pca_current_label = pca_list[i]
ax.scatter(pca_current_label[:, 0], pca_current_label[:, 1],
c=COLORS[i], label=str(i))
ax.legend()
plt.show()
```
Calculate and plot the mean for each digit in PCA coordinates
```
pca_mean_list = [0] * 10
for i in range(10):
pca_mean_list[i] = np.mean(pca_list[i], axis=0)
COLORS = ["red", "blue", "green", "yellow", "darkviolet",
"maroon", "greenyellow", "hotpink", "black", "cyan"]
fig, ax = plt.subplots()
for i in range(10):
# Get the pca array corresponding to the current label
pca_current_label = pca_mean_list[i]
ax.scatter(pca_current_label[0], pca_current_label[1],
c=COLORS[i], label=str(i))
ax.legend()
plt.show()
```
# SVD Visualization
Compare the PCA visualization with SVD dimensionality reduction
Calculate SVD and use dimensionality reduction to get down to 2 coordinates
```
# Calculate u, s, v
u, s, v = np.linalg.svd(x_train_colvector_sample2000, full_matrices=False)
# Set all singular values greater than the first two to 0
for i in range(2, s.shape[0]):
s[i] = 0
# Calculate the reduced dimensions with svd
svd_cords = np.diag(s) @ v
```
Sort by label
```
svd_list= [0] * 10
for i in range(10):
svd_list[i] = (svd_cords.T[y_train_sample2000 == i])
```
Plot the SVD coordinates
```
COLORS = ["red", "blue", "green", "yellow", "darkviolet",
"maroon", "greenyellow", "hotpink", "black", "cyan"]
fig, ax = plt.subplots()
for i in range(10):
# Get the pca array corresponding to the current label
svd_current_label = svd_list[i]
ax.scatter(svd_current_label[:, 0], svd_current_label[:, 1],
c=COLORS[i], label=str(i))
ax.legend()
plt.show()
```
Calculate and plot the mean for each digit in SVD coordinates
```
svd_mean_list = [0] * 10
for i in range(10):
svd_mean_list[i] = np.mean(svd_list[i], axis=0)
COLORS = ["red", "blue", "green", "yellow", "darkviolet",
"maroon", "greenyellow", "hotpink", "black", "cyan"]
fig, ax = plt.subplots()
for i in range(10):
# Get the pca array corresponding to the current label
svd_current_label = svd_mean_list[i]
ax.scatter(svd_current_label[0], svd_current_label[1],
c=COLORS[i], label=str(i))
ax.legend()
plt.show()
```
# Sorting Training Digits By Label
Sort the training images by label
```
x_list= [0] * 10
y_list = [0] * 10
for i in range(10):
# Get x and y values in each label by the coordinate in the list
x_list[i] = (x_train_colvector[:, y_train == i])
y_list[i] = (y_train[y_train == i])
```
# Mean Clustering Classification
Calculate the Mean Image for Each Digit
```
means_list = [0] * 10
for i in range(10):
means_list[i] = np.mean(x_list[i], axis=1)
```
Visualize the Mean Image for Each Digit
```
for i in range(10):
print("Mean Image for Digit", i)
image = means_list[i]
# Show singular image
plt.imshow(image.reshape(28, 28), cmap="Greys")
plt.show()
plt.close()
```
Classify Each Unknown Digit by the Mean Image
```
# Create vector for y predictions
y_pred = np.zeros(len(y_test_sample200))
# Iterate through all the testing images and make a prediction
for i in range(len(y_pred)):
# Get the unknown digit
x = x_test_colvector_sample200[:, i]
# Calculate the residual of the digit to each of the mean digits
residuals = np.zeros(10)
for j in range(10):
# Calculate residual,
residuals[j] = np.linalg.norm(means_list[j] - x, ord=2)
# Find the minimum residual and store as prediction
y_pred[i] = np.argmin(residuals)
```
Calculate the accuracy score
```
correct = np.where(y_pred == y_test_sample200, 1, 0)
print("Accuracy For Mean Digit: ", np.sum(correct) / len(correct))
```
# SVD Singular Images Visualization
Compute Top 3 Singular Images for each digit and visualize
```
# Iterate through all the digits
for i in range(10):
print("#################################################################")
print("#################################################################")
print("Visualizing Singular Images for " + str(i))
print("#################################################################")
print("#################################################################")
# Calculate the SVD for that digit
u, s, v = np.linalg.svd(x_list[i], full_matrices=False)
# Visualize the first three singular images
for j in range(3):
print("Visualizing Singular Image Number " + str(j + 1))
# Get the singular image
image = u[:, j]
# Show singular image
plt.imshow(image.reshape(28, 28), cmap="Greys")
plt.show()
plt.close()
```
# SVD Singular Image Classification
Compute the Singular Value Decomposition for each digit
```
u_list = [0] * 10
s_list = [0] * 10
v_list = [0] * 10
# Iterate through each digit
for i in range(10):
# Calculate the SVD for that digit
u_list[i], s_list[i], v_list[i] = np.linalg.svd(x_list[i], full_matrices=False)
```
Calculate the Accuracy for Different Number of Singular Images
```
# Store predictions and accuracy at different number of singular images used
acc_list = [0] * 5
pred_list = [0] * 5
# Use only the first k basis image for classification
for k in range(5):
# List to store the values of uk @ uk.T to get the singular images sum
uk_ukt_list = [0] * 10
# Iterate through all digits and calculate uk @ uk.T for that digit
for i in range(10):
uk = np.zeros((784, 784))
uk[:,0:k+1] = u_list[i][:, 0:k+1]
uk_ukt_list[i] = uk @ uk.T
# Iterate through the testing images and get the prediction for each image
# Initialize predictions to 0
y_pred = np.zeros(len(y_test_sample200))
# Iterate through all the testing images
for i in range(len(y_pred)):
# Get the unknown digit
x = x_test_colvector_sample200[:, i]
# Calculate the residual of the digit to each of the singular bases
residuals = np.zeros(10)
# Iterate through the 10 singular bases
for j in range(10):
# Calculate residual, which is norm of (I - uk @ uk.T) @ z
residuals[j] = np.linalg.norm((np.identity(28*28) - uk_ukt_list[j]) @ x, ord=2)
# Find the minimum residual and store that as the predicted digit
y_pred[i] = np.argmin(residuals)
# Store all the predictions for this threshold
pred_list[k] = y_pred
# Calculate and store the accuracy for this threshold
correct = np.where(y_pred == y_test_sample200, 1, 0)
accuracy = np.sum(correct) / len(correct)
print("Accuracy with", k + 1, "singular images: ", accuracy)
acc_list[k] = accuracy
```
# Confusion Matrix Visualization at Each Number of Singular Images
Visualize a confusion matrix at each number of singular images
```
import seaborn
from sklearn import metrics
for k in range(5):
print("Confusion Matrix at", k + 1, "singular images")
# Use scikit-learn to calculate confusion matrix
confusion_matrix = metrics.confusion_matrix(y_test_sample200, pred_list[k], normalize="true")
# Use seaborn to plot heatmap
axes = seaborn.heatmap(confusion_matrix, annot=True)
axes.set(xlabel="Predicted Label", ylabel="Actual Label", title="Confusion Matrix")
# Save as image and show plot.
plt.show()
plt.close()
```
| github_jupyter |
The goal of this notebook is to verify that you can load the checkpointed model from it's github repo and run it on a few test image samples and verify that the whole inference pipeline works.
```
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:100% !important; }</style>"))
```
First, the imports:
```
%matplotlib inline
import sys
import numpy as np
import cv2 as cv
import tensorflow as tf
from models import resnet as resnet
import matplotlib.pyplot as plt
import pandas as pd
import os
```
Helper functions
```
def _load_dictionary(dict_file):
dictionary = dict()
with open(dict_file, 'r') as lines:
for line in lines:
sp = line.rstrip('\n').split('\t')
idx, name = sp[0], sp[1]
dictionary[idx] = name
return dictionary
# Load the labels:
# dictionary = _load_dictionary("ml2020_dictionary.txt")
# I generated these a priori
def preprocess(img):
rawH = float(img.shape[0])
rawW = float(img.shape[1])
newH = 256.0
newW = 256.0
test_crop = 224.0
if rawH <= rawW:
newW = (rawW/rawH) * newH
else:
newH = (rawH/rawW) * newW
img = cv.resize(img, (int(newW), int(newH)))
img = img[int((newH-test_crop)/2):int((newH-test_crop)/2)+int(test_crop),int((newW-test_crop)/2):int((newW-test_crop)/2)+int(test_crop)]
img = ((img/255.0) - 0.5) * 2.0
img = img[...,::-1]
return img
```
Model declaration and weight restoration
```
images = tf.placeholder(dtype=tf.float32, shape=[None, 224, 224, 3])
net = resnet.ResNet(images, is_training=False)
net.build_model()
logit = net.logit
prob = tf.nn.softmax(logit)
prob_topk, pred_topk = tf.nn.top_k(prob, k=20)
# restore model
config = tf.ConfigProto()
config.gpu_options.allow_growth = True
config.log_device_placement=False
sess = tf.Session(config=config)
saver = tf.train.Saver(tf.global_variables())
saver.restore(sess, "./checkpoints/model.ckpt")
print('Architecture details \n')
print(f'N_class:{net.num_classes},Stages: {net.stages}, N_filters: {net.filters}')
```
In case you want generate the dictionary of labels on the spot:
```
url_ml='https://raw.githubusercontent.com/Tencent/tencent-ml-images/master/data/dictionary_and_semantic_hierarchy.txt'
df_ml=pd.read_csv(url_ml,delimiter=' ')
print(df_ml.shape)
df_ml.head()
dictionary_ml = {}
N_class=df_ml.shape[0]
keys = range(N_class)
values = list(df_ml.loc[:,'category name'].values)
from tqdm.notebook import tqdm
for i in keys:
dictionary_ml[i] = values[i]
# print(dictionary_ml)
# Manual selection
test_dir=os.getcwd()+'/ml_test/'
fig=plt.figure(figsize=(15,10))
for im_ind,im in enumerate(os.listdir(test_dir)):
im=test_dir+im
raw_img = cv.imread(im)
img = preprocess(raw_img)
logits, probs_topk, preds_topk = sess.run([logit, prob_topk, pred_topk], {images:np.expand_dims(img, axis=0)})
preds_topk = np.squeeze(preds_topk)
# print(preds_topk)
names_topk = [dictionary_ml[i] for i in preds_topk]
ax = fig.add_subplot(2,4,im_ind+ 1)
ax.imshow(raw_img[...,::-1])
plt.axis('Off')
predictions = []
for i, pred in enumerate(preds_topk[0:10]):
predictions.append('%d %s: %.3f' % (pred, names_topk[i], probs_topk[0][i]))
ax.set_title('\n'.join(predictions),fontsize=8)
file_name=im.split('/')[-1]
ax.text(0.5,-0.1,f'File: {file_name}',ha="center",
transform=ax.transAxes)
# plt.tight_layout()
```
| github_jupyter |
# Neural Network for Regression
In the previous homework you implemented a linear regression network. In this exercise, we will solve the same problem with a neural network instead, to leverage the power of Deep Learning.
We will implement our neural networks using a modular approach. For each layer we will implement a `forward` and a `backward` function. The `forward` function will receive inputs, weights, and other parameters and will return both an output and a `cache` object storing data needed for the backward pass, like this:
```python
def layer_forward(x, w):
""" Receive inputs x and weights w """
# Do some computations ...
z = # ... some intermediate value
# Do some more computations ...
out = # the output
cache = (x, w, z, out) # Values we need to compute gradients
return out, cache
```
The backward pass will receive upstream derivatives and the `cache` object, and will return gradients with respect to the inputs and weights, like this:
```python
def layer_backward(dout, cache):
"""
Receive derivative of loss with respect to outputs and cache,
and compute derivative with respect to inputs.
"""
# Unpack cache values
x, w, z, out = cache
# Use values in cache to compute derivatives
dx = # Derivative of loss with respect to x
dw = # Derivative of loss with respect to w
return dx, dw
```
After implementing a bunch of layers this way, we will be able to easily combine them to build networks with different architectures.
```
# As usual, a bit of setup
from exercise_code.data.csv_dataset import CSVDataset
from exercise_code.data.csv_dataset import FeatureSelectorAndNormalizationTransform
from exercise_code.data.dataloader import DataLoader
import matplotlib.pyplot as plt
import numpy as np
import os
import pandas as pd
import seaborn as sns
pd.options.mode.chained_assignment = None # default='warn'
%matplotlib inline
%load_ext autoreload
%autoreload 2
```
# 1. Load your data
We apply the same dataloading and preprocessing steps as in the previous exercise.
```
target_column = 'SalePrice'
i2dl_exercises_path = os.path.dirname(os.path.abspath(os.getcwd()))
root_path = os.path.join(i2dl_exercises_path, "datasets", 'housing')
housing_file_path = os.path.join(root_path, "housing_train.csv")
download_url = 'https://cdn3.vision.in.tum.de/~dl4cv/housing_train.zip'
# Always make sure this line was run at least once before trying to
# access the data manually, as the data is downloaded in the
# constructor of CSVDataset.
train_dataset = CSVDataset(target_column=target_column, root=root_path, download_url=download_url, mode="train")
df = train_dataset.df
target_column = 'SalePrice'
# Select only 2 features to keep plus the target column.
selected_columns = ['GrLivArea','GarageArea', target_column]
mn, mx, mean = df.min(), df.max(), df.mean()
column_stats = {}
for column in selected_columns:
crt_col_stats = {'min' : mn[column],
'max' : mx[column],
'mean': mean[column]}
column_stats[column] = crt_col_stats
transform = FeatureSelectorAndNormalizationTransform(column_stats, target_column)
def rescale(data, key = "SalePrice", column_stats = column_stats):
""" Rescales input series y"""
mx = column_stats[key]["max"]
mn = column_stats[key]["min"]
return data * (mx - mn) + mn
train_dataset = CSVDataset(mode="train", target_column=target_column, root=root_path, download_url=download_url, transform=transform)
val_dataset = CSVDataset(mode="val", target_column=target_column, root=root_path, download_url=download_url, transform=transform)
test_dataset = CSVDataset(mode="test", target_column=target_column, root=root_path, download_url=download_url, transform=transform)
print("Number of training samples:", len(train_dataset))
print("Number of validation samples:", len(val_dataset))
print("Number of test samples:", len(test_dataset))
```
# 2. Build your Model
Now we want to build our model. But let's first construct the building blocks we want to use. We will define the forward and backward pass for an affine layer and a Sigmoid activation function
## 2.1 Affine Layer
Open the file `exercise_code/networks/layer.py` and implement the `affine_forward` and the `affine_backward` function. Remember, and affine layer computes a function of
$$\mathbf{z} = \mathbf{W} \mathbf{x}$$
To check the correctness of your implementation, we will again use numeric gradient checking:
$$ \frac {df(x)}{dx} = \frac{f(x+h) - f(x-h)}{2h} $$
Once you are done you can test your implementaion by running the following:
```
a=np.random.rand(20,20,8,6,4)
a.shape
s=a.shape[:2]
s
if a.ndim>2 : s = s+ (-1,)
a.shape[:2] + (-1,)
s
b=a.reshape(s)
# Test the affine function
from exercise_code.tests.layer_tests import *
print(AffineLayerTest()())
```
## 2.2 Sigmoid layer:
Implement the forward pass for the sigmoid activation function in the `sigmoid_forward` function and the backward pass in `sigmoid_backward`.
$$ y = \sigma(z) = \frac{1}{1+\mathrm{exp}(-z)}, $$
Test your implementation using the following:
```
# Test the sigmoid function
print(SigmoidTest()())
```
## 2.3 Two-layer regression network
Now that you have all the necessary building block, let's build your first neural network.
Open the file `exercise_code/networks/regression_net.py` and complete the implementation of the `RegressionNet` class. Specifically, you again need co complete the `forward` and `backward` functions. You can run the cell below to test your implementation.
```
from exercise_code.tests.regression_net_tests import test_regression_net
from exercise_code.networks.regression_net import RegressionNet
test_regression_net(RegressionNet)
```
# 3. Optimizer & Loss Function
We have now implemented:
- [x] A dataloader
- [x] A loss function
- [x] A model
- [ ] An optimizer
- [ ] A loss function
The only things missing in out Deep Learning pipeline are an optimizer and a loss function. Since you already implemented SGD and MSE in last weeks' exercise, we will give them to you this time. Have a look at their implementations in `exercise_code/networks/optimizer.py` and `exercise_code/networks/loss.py`.
```
from exercise_code.networks.optimizer import SGD
from exercise_code.networks.loss import MSE, L1
```
# 4. Solver
Now that we have everything together, let's update our solver from exercise_04 and finally start training our model.
Open the file `exercise_code/solver.py` and read through it to familiarize yourself with the API. In the `train` and `_step` functions, you can see all of the components you implemented in the last exercises working together. Now, run the solver to train your model.
We provide you with a default set of hyperparameters here as hyperparameter search is not the scope of this exercise. However, you can still play around with those values and see how the training performance changes. Especially the `std` which is the standard deviation of the gaussian distribution used to initialize the weights of your model is very sensitive.
```
from exercise_code.networks.regression_net import RegressionNet
from exercise_code.solver import Solver
batch_size = 4
lr = 1e-3
hidden_size = 100
std = 1.
epochs = 20
model = RegressionNet(input_size=2, hidden_size=hidden_size, std=std)
train_dataloader = DataLoader(train_dataset, batch_size=batch_size)
val_dataloader = DataLoader(val_dataset, batch_size=batch_size)
test_dataloader = DataLoader(test_dataset, batch_size=batch_size)
solver = Solver(model, train_dataloader, val_dataloader, learning_rate=lr, loss_func=MSE(), optimizer=SGD)
# add test data to test before training
X_test = [test_dataset[i]['features'] for i in range((len(test_dataset)))]
X_test = np.stack(X_test, axis=0)
y_test = [test_dataset[i]['target'] for i in range((len(test_dataset)))]
y_test = np.stack(y_test, axis=0)
y_out = solver.get_dataset_prediction(test_dataloader)
l1_loss = L1()
mse_loss = MSE()
print("L1 loss on test set BEFORE training: {:,.0f}".format(l1_loss(rescale(y_out), rescale(y_test))[0].mean() ))
print("MSE loss on test set BEFORE training: {:,.0f}".format(mse_loss(rescale(y_out), rescale(y_test))[0].mean() ))
solver.train(epochs=epochs)
y_out, _ = model(X_test)
l1_loss = L1()
mse_loss = MSE()
print("L1 loss on test set AFTER training: {:,.0f}".format(l1_loss(rescale(y_out), rescale(y_test))[0].mean() ))
print("MSE loss on test set AFTER training: {:,.0f}".format(mse_loss(rescale(y_out), rescale(y_test))[0].mean() ))
# # Run this cell to visualize your training and validation loss and your prediction
y_out = solver.get_dataset_prediction(test_dataloader)
plt.title('Loss curves')
plt.plot(solver.train_loss_history, '-', label='train')
plt.plot(solver.val_loss_history, '-', label='val')
plt.legend(loc='lower right')
plt.xlabel('Iteration')
plt.show()
if np.shape(X_test)[1]==1:
plt.scatter(X_test, y_test, label = "Ground Truth")
inds = X_test.argsort(0).flatten()
plt.plot(X_test[inds], y_out[inds], color='r', label = "Prediction")
plt.legend()
plt.show()
else:
from mpl_toolkits.mplot3d import Axes3D
fig = plt.figure()
ax = plt.axes(projection='3d')
first_feature = X_test[:, 0]
second_feature = X_test[:, 1]
salePrice = y_test[:, 0]
salePricePred = y_out[:, 0]
ax.plot_trisurf(first_feature, second_feature, salePricePred, linewidth=0, antialiased=True,color ="red")
ax.scatter(first_feature, second_feature, salePrice)
ax.set_xlabel(selected_columns[0])
ax.set_ylabel(selected_columns[1])
ax.set_zlabel(selected_columns[2])
plt.tight_layout()
plt.show()
```
## Save the model for submission
Simply save your objects using the following cell. This will save them to a pickle file `models/two_layer_regression.p`.
```
from exercise_code.tests import save_pickle
save_pickle(
data_dict={
"Regression_Net": RegressionNet
},
file_name="two_layer_regression.p"
)
```
# Submission Goals
- Goal: Successfully implement the forward and backward pass of a two layer regression neural network
- Test cases:
1. Does `forward()` and `backward()` of your 2 layer regression neural net return the correct value and data type?
- Reachable points [0, 100]: 0 if not implemented, 100 if all tests passed, 50 per passed test
- Threshold to clear exercise: 100
- Submission start: __May 22, 2020 12.00__
- Submission deadline : __May 27, 2020 23.59__
- You can make multiple submission uptil the deadline. Your __best submission__ will be considered for bonus
| github_jupyter |
```
# import packages
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import featuretools as ft
import lightgbm as lgb
%matplotlib inline
import seaborn as sns
import math
import pickle
import os, sys, gc, warnings, random, datetime
RSEED = 50
```
## Load Data
```
# Load training data
df_train_transac = pd.read_csv('./data/train_transaction.csv')
df_train_identity = pd.read_csv('./data/train_identity.csv')
df_train = pd.merge(df_train_transac,df_train_identity,on='TransactionID',how='left')
# Load test data
df_test_transac = pd.read_csv('./data/test_transaction.csv')
df_test_identity = pd.read_csv('./data/test_identity.csv')
df_test = pd.merge(df_test_transac,df_test_identity,on='TransactionID',how='left')
# combine train and test
df_total = df_train.append(df_test,sort=False)
df_total.tail()
```
# Feature Engineer
```
def clean_id31(df):
df['id_31'] = df['id_31'].str.replace("([0-9\.])", "")
df['id_31'][df['id_31'].str.contains('chrome', regex=False)==True] = 'chrome'
df['id_31'][df['id_31'].str.contains('Samsung', regex=False)==True] = 'Samsung'
df['id_31'][df['id_31'].str.contains('samsung', regex=False)==True] = 'Samsung'
df['id_31'][df['id_31'].str.contains('firefox', regex=False)==True] = 'firefox'
df['id_31'][df['id_31'].str.contains('safari', regex=False)==True] = 'safari'
df['id_31'][df['id_31'].str.contains('opera', regex=False)==True] = 'opera'
df['id_31'] = df['id_31'].str.replace(" ", "")
df.loc[df['id_31'].str.contains('Generic/Android', na=False), 'id_31'] = 'Android'
df.loc[df['id_31'].str.contains('androidbrowser', na=False), 'id_31'] = 'Android'
df.loc[df['id_31'].str.contains('androidwebview', na=False), 'id_31'] = 'Android'
df.loc[df['id_31'].str.contains('android', na=False), 'id_31'] = 'Android'
df.loc[df['id_31'].str.contains('chromium', na=False), 'id_31'] = 'chrome'
df.loc[df['id_31'].str.contains('google', na=False), 'id_31'] = 'chrome'
df.loc[df['id_31'].str.contains('googlesearchapplication', na=False), 'id_31'] = 'chrome'
df.loc[df['id_31'].str.contains('iefordesktop', na=False), 'id_31'] = 'ie'
df.loc[df['id_31'].str.contains('iefortablet', na=False), 'id_31'] = 'ie'
df.loc[df.id_31.isin(df.id_31.value_counts()[df.id_31.value_counts() < 20].index), 'id_31'] = "rare"
return df
def label_encoder(df, categorical_columns=None):
"""Encode categorical values as integers (0,1,2,3...) with pandas.factorize. """
# if categorical_colunms are not given than treat object as categorical features
if not categorical_columns:
categorical_columns = [col for col in df.columns if df[col].dtype == 'object']
for col in categorical_columns:
df[col], uniques = pd.factorize(df[col])
return df, categorical_columns
def make_dow_feature(df, offset=0, tname='TransactionDT'):
"""
Creates a day of the week feature, encoded as 0-6.
Parameters:
-----------
df : pd.DataFrame
df to manipulate.
offset : float (default=0)
offset (in days) to shift the start/end of a day.
tname : str
Name of the time column in df.
"""
# found a good offset is 0.58
days = df[tname] / (3600*24)
encoded_days = np.floor(days-1+offset) % 7
return encoded_days
def make_hour_feature(df, tname='TransactionDT'):
"""
Creates an hour of the day feature, encoded as 0-23.
Parameters:
-----------
df : pd.DataFrame
df to manipulate.
tname : str
Name of the time column in df.
"""
hours = df[tname] / (3600)
encoded_hours = np.floor(hours) % 24
return encoded_hours
def make_pdc_amt_ratio(df):
df_product_aveAmt = df.groupby(['ProductCD'])['TransactionAmt'].agg(['mean'])
df_product_aveAmt.reset_index(inplace=True)
df_ratio = pd.merge(df[['TransactionID','ProductCD',
'TransactionAmt','isFraud']],
df_product_aveAmt,on='ProductCD',how='left')
return df_ratio['TransactionAmt']/df_ratio['mean']
def make_card_id(df):
'''
cards_cols= ['card1', 'card2', 'card3', 'card5']
for card in cards_cols:
if '1' in card:
df['card_id']= df[card].map(str)
else :
df['card_id']+= ' '+df[card].map(str)
'''
cards_cols= ['card2','card3','card4','card5','card6','addr1',
'C1','C2','C3','C4','C5','C6','C7','C8','C9','C10','C11']
df['card_id'] = df['card1'].astype(str)
for card in cards_cols:
df['card_id']+= '_'+df[card].astype(str)
return df['card_id']
def high_missing_cols(df,threshold):
"""return features with high missing rate"""
rm_cols = [col for col in df.columns
if df[col].isnull().mean() > threshold]
return rm_cols
```
# Confirmed Features
```
# proved important feature
df_total = clean_id31(df_total)
# proved important feature
# clean Pemail
df_total['P_email'] = df_total['P_emaildomain'].str.split('.',expand=True)[0]
#df_total['P_email_suffix'] = df_total['P_emaildomain'].map(lambda x: str(x).split('.')[-1])
#df_total['P_email_suffix'] = df_total['P_email_suffix'].map(lambda x: x if str(x) not in us_emails else 'us')
df_total.drop('P_emaildomain',axis=1,inplace = True)
# proved important feature
# clean R_emaildomain
df_total['R_email'] = df_total['R_emaildomain'].str.split('.',expand=True)[0]
#df_total['R_email_suffix'] = df_total['R_emaildomain'].map(lambda x: str(x).split('.')[-1])
#df_total['R_email_suffix'] = df_total['R_email_suffix'].map(lambda x: x if str(x) not in us_emails else 'us')
df_total.drop('R_emaildomain',axis=1,inplace = True)
# proved important feature
df_total['screen_width'] = df_total['id_33'].str.split('x', expand=True)[0]
df_total['screen_height'] = df_total['id_33'].str.split('x', expand=True)[1]
df_total.drop('id_33',axis=1,inplace = True)
# proved important feature
df_total['pdc_amt_ratio'] = df_total['TransactionAmt']/df_total.groupby('ProductCD')['TransactionAmt'].transform('mean')
for col in ['card1','addr1']:
df_total[col+'_fq_enc'] = df_total.groupby([col])['TransactionID'].transform('count')
# 识别持卡人
df_total['card_id'] = make_card_id(df_total)
START_DATE = datetime.datetime.strptime('2017-11-30', '%Y-%m-%d')
df_total['DT'] = df_total['TransactionDT'].apply(lambda x: (START_DATE + datetime.timedelta(seconds = x)))
df_total['DT_hour'] = df_total['DT'].dt.hour
df_total['DT_day'] = df_total['DT'].dt.day
df_total['DT_month'] = df_total['DT'].dt.month
df_total['DT_year'] = df_total['DT'].dt.year
# 该持卡人当前一天刷卡次数
df_total['card_mv_day_fq'] = df_total.groupby(['card_id','DT_day','DT_month','DT_year'])['TransactionID'].transform('count')
for col in ['DeviceType']:
df_total['card_'+col+'_fq'] = df_total.groupby(['card_id',col])['TransactionID'].transform('count')
```
# Test Features
```
df_total['pdc_hour_Amt_mean'] = df_total.groupby(['ProductCD','DT_hour','DT_day','DT_month','DT_year'])['TransactionAmt'].transform('mean')
df_total['pdc_hour_Amt_ratio'] = df_total['TransactionAmt']/df_total['pdc_hour_Amt_mean']
df_total['pdc_month_Amt_mean'] = df_total.groupby(['ProductCD','DT_month','DT_year'])['TransactionAmt'].transform('mean')
df_total['pdc_month_Amt_ratio'] = df_total['TransactionAmt']/df_total['pdc_month_Amt_mean']
# 持卡人在某种类下的统计 'DeviceInfo','screen_width'
for col in ['id_30','id_31']:
df_total['card_'+col+'_fq'] = df_total.groupby(['card_id',col])['TransactionID'].transform('count')
# 不同产品不同地区消费均值比
df_total['pdc_addr_Amt_ratio'] = df_total['TransactionAmt']/df_total.groupby(['ProductCD','addr1'])['TransactionAmt'].transform('mean')
# 不同地区消费均值比
df_total['addr_Amt_ratio'] = df_total['TransactionAmt']/df_total.groupby(['addr1'])['TransactionAmt'].transform('mean')
# 不同设备类别消费均值比
df_total['dev_Amt_ratio'] = df_total['TransactionAmt']/df_total.groupby(['DeviceType'])['TransactionAmt'].transform('mean')
df_total['pemail_Amt_ratio'] = df_total['TransactionAmt']/df_total.groupby(['P_email'])['TransactionAmt'].transform('mean')
# 刷卡人历史不同时段刷卡次数
#df_total['card_hour_fq'] = df_total.groupby(['card_id','DT_hour'])['TransactionID'].transform('count')
# 当前时段刷卡次数
#df_total['card_this_hour_fq'] = df_total.groupby(['card_id','DT_hour','DT_day','DT_month','DT_year'])['TransactionID'].transform('count')
# 持卡人当前小时刷卡次数比历史该时段次数
#df_total['card_hour_ratio'] = df_total['card_this_hour_fq']/df_total['card_hour_fq']
# 不同时段产品购买数量
#df_total['pdc_hour_cnt'] = df_total.groupby(['ProductCD','DT_hour'])['TransactionID'].transform('count')
# 不同产品不同时段购买均值
#df_total['pdc_hour_Amt_mean'] = df_total.groupby(['card_id','ProductCD','DT_hour'])['TransactionAmt'].transform('mean')
# 持卡人距离均值
#df_total['card_dist1_mean'] = df_total['dist1']/df_total.groupby(['card_id'])['dist1'].transform('mean')
#df_total['card_dist2_mean'] = df_total['dist2']/df_total.groupby(['card_id'])['dist2'].transform('mean')
# 产品品种使用设备统计
#for col in ['DeviceType','DeviceInfo','screen_width','id_30','id_31']:
# df_total['pdc_'+col+'_fq'] = df_total.groupby(['ProductCD',col])['TransactionID'].transform('count')
# 设备组合出现次数统计
#df_total['DeviceType_DeviceInfo_fq'] = df_total.groupby(['DeviceType','DeviceInfo'])['TransactionID'].transform('count')
#df_total['DeviceType_screen_width_fq'] = df_total.groupby(['DeviceType','screen_width'])['TransactionID'].transform('count')
#df_total['DeviceType_id_30_fq'] = df_total.groupby(['DeviceType','id_30'])['TransactionID'].transform('count')
#df_total['DeviceType_id_31_fq'] = df_total.groupby(['DeviceType','id_31'])['TransactionID'].transform('count')
#df_total['DeviceInfo_id_31_fq'] = df_total.groupby(['DeviceInfo','id_31'])['TransactionID'].transform('count')
#df_total['DeviceInfo_id_30_fq'] = df_total.groupby(['DeviceInfo','id_30'])['TransactionID'].transform('count')
#df_total['DeviceInfo_screen_width_fq'] = df_total.groupby(['DeviceInfo','screen_width'])['TransactionID'].transform('count')
#df_total['DeviceInfo_screen_width_fq'] = df_total.groupby(['DeviceInfo','screen_width'])['TransactionID'].transform('count')
#df_total['screen_width_id_30_fq'] = df_total.groupby(['id_30','screen_width'])['TransactionID'].transform('count')
#df_total['screen_width_id_31_fq'] = df_total.groupby(['id_31','screen_width'])['TransactionID'].transform('count')
#df_total['id_30_id_31_fq'] = df_total.groupby(['id_31','id_30'])['TransactionID'].transform('count')
#类别下的单纯计数
# useless 'id_30',,'card2','card3','card5','P_email','R_email','addr2',
# 'dist1','dist2','D1','D2','D3','D4','D5','id_31'
#for col in ['D6','D7','D8','D9','DeviceInfo','screen_width']:
# df_total[col+'_fq_enc'] = df_total.groupby([col])['TransactionID'].transform('count')
#for col in ['DT_hour','DT_day','DT_month','card2','card1',
# 'card3','card4','card5','card6']:#,
# df_total[col+'_Amt_ratio'] = df_total['TransactionAmt']/df_total.groupby([col])['TransactionAmt'].transform('mean')
#df_total['DeviceInfo'] = df_total['DeviceInfo'].fillna('unknown_device').str.lower()
#df_total['DeviceInfo'] = df_total['DeviceInfo'].apply(lambda x: ''.join([i for i in x if i.isalpha()]))
#df_total['DeviceInfo_version'] = df_total['DeviceInfo'].apply(lambda x: ''.join([i for i in x if i.isnumeric()]))
# D9是小时记录D9中的NA信息同时保留做出来的小时变量
#df_total['D9'] = np.where(df_total['D9'].isna(),0,1)
#df_total['id_30'] = df_total['id_30'].fillna('unknown_device').str.lower()
#df_total['id_30'] = df_total['id_30'].apply(lambda x: ''.join([i for i in x if i.isalpha()]))
#df['id_30_version'] = df['id_30'].apply(lambda x: ''.join([i for i in x if i.isnumeric()]))
#i_cols = ['M1','M2','M3','M5','M6','M7','M8','M9']
#df_total['M_sum'] = df_total[i_cols].sum(axis=1).astype(np.int8)
#df_total['M_na'] = df_total[i_cols].isna().sum(axis=1).astype(np.int8)
# 该持卡人当前一小时总计刷卡次数
#df_total['card_hour_fq'] = df_total.groupby(['card_id','DT_hour','DT_day','DT_month','DT_year'])['TransactionID'].transform('count')
# 该持卡人当前一小时总刷卡次数比平均次数
#df_total['card_hour_fq_ratio'] = df_total['card_mv_hour_fq']/df_total.groupby('card_id')['card_mv_hour_fq'].transform('mean')
# 该持卡人每小时消费总额
#df_total['card_hour_Amt'] = df_total.groupby(['card_id','DT_hour','DT_day','DT_month','DT_year'])['TransactionAmt'].transform('sum')
# 该持卡人当前小时消费额比平均小时消费额
#df_total['card_hour_Amt_ratio'] = df_total['card_hour_Amt']/df_total.groupby('card_id')['card_hour_Amt'].transform('mean')
# 该持卡人当前一天刷卡次数比平均次数
#df_total['card_mv_day_fq_ratio'] = df_total['card_mv_day_fq']/df_total.groupby('card_id')['card_mv_day_fq'].transform('mean')
# 该持卡人当日消费总额
#df_total['card_day_Amt'] = df_total.groupby(['card_id','DT_day','DT_month','DT_year'])['TransactionAmt'].transform('sum')
# 该持卡人当日消费总额比平均消费额
#df_total['card_day_Amt_ratio'] = df_total['card_day_Amt']/df_total.groupby('card_id')['card_day_Amt'].transform('mean')
#,'ProductCD','M4','remail_fraud_rate',
# 'R_email','card6' boost but less than card4
#for col in ['P_email','card4']:
# df_total[col+'_fraud_rate'] = df_total.groupby([col])['isFraud'].transform('mean')
# proved important feature
#df_total['hour'] = make_hour_feature(df_total)
#df_total['hour_Amt'] = df_total['TransactionAmt']/df_total.groupby('hour')['TransactionAmt'].transform('mean')
#df_total['card_TAmt_ratio'] = df_total['TransactionAmt']/df_total.groupby('card_id')['TransactionAmt'].transform('mean')
#df_total['card_hour_fq'] = df_total.groupby(['card_id','hour'])['TransactionID'].transform('count')
#df_total['card_pdc_cnt'] = df_total.groupby(['card_id','ProductCD'])['TransactionID'].transform('count')
#df_total['dow'] = make_day_feature(df_total, offset=0.58)
#df_total['pdc_amt_std_ratio'] = df_total['TransactionAmt']/df_total.groupby('ProductCD')['TransactionAmt'].transform('std')
#df_total['id_30_OS'] = df_total['id_30'].str.split(' ',expand=True)[0]
#df_total['id_30_version'] = df_total['id_30'].str.split(' ',expand=True)[1]
#df_total['is_win8_vista'] = (df_total.id_30_OS == 'Windows')&((df_total.id_30_version == '8')| (df_total.id_30_version == 'Vista'))
#df_total['is_windows_otheros'] = (df_total.DeviceInfo == 'Windows')&((df_total.id_30_OS == 'Linux')| (df_total.id_30_OS == 'other'))
#df_total.drop('id_30',axis=1,inplace = True)
# # proved important feature
#df_total['pdc_D1_ratio'] = df_total['D1']/df_total.groupby('ProductCD')['D1'].transform('mean')
# proved important feature
#df_total['pdc_D2_ratio'] = df_total['D2']/df_total.groupby('ProductCD')['D2'].transform('mean')
#df_total['pdc_D3_ratio'] = df_total['D3']/df_total.groupby('ProductCD')['D3'].transform('mean')
#df_total['pdc_D4_ratio'] = df_total['D4']/df_total.groupby('ProductCD')['D4'].transform('mean')
# proved important feature
#df_total['pdc_D1_std_ratio'] = df_total['D1']/df_total.groupby('ProductCD')['D1'].transform('std')
# proved important feature
#df_total['pdc_D2_std_ratio'] = df_total['D2']/df_total.groupby('ProductCD')['D2'].transform('std')
# not so important
#df_total['pdc_D3_std_ratio'] = df_total['D3']/df_total.groupby('ProductCD')['D3'].transform('std')
#df_total['pdc_D4_std_ratio'] = df_total['D4']/df_total.groupby('ProductCD')['D4'].transform('std')
# proved important feature
#df_total['card_TAmt_std_ratio'] = df_total['TransactionAmt'] / df_total.groupby(['card_id'])['TransactionAmt'].transform('std')
#df_total['card_freq_pdc'] = df_total.groupby('card_id')['ProductCD'].transform(lambda x:x.value_counts().index[0])
#df_total['is_card_freq_pdc'] = (df_total.ProductCD == df_total.card_freq_pdc)
#df_total.drop(['card_freq_pdc'],axis=1,inplace=True)
#df_total['card_freq_addr1'] = df_total.groupby('card_id')['addr1'].transform(lambda x: x.value_counts(dropna=False).index[0])
#df_total['is_card_freq_addr1'] = (df_total.addr1 == df_total.card_freq_addr1)
#df_total.drop(['card_freq_addr1'],axis=1,inplace=True)
#df_total['card1_count'] = df_total['card1'].map(df_total['card1'].value_counts(dropna=False)).head()
#df_total['card_id_02_mean'] = df_total['id_02'] / df_total.groupby(['card_id'])['id_02'].transform('mean')
#df_total['card_id_02_std'] = df_total['id_02'] / df_total.groupby(['card_id'])['id_02'].transform('std')
# not so important
#df_total['card_D1_mean'] = df_total['D1'] / df_total.groupby(['card_id'])['D1'].transform('mean')
# not so important
#df_total['card_D2_mean'] = df_total['D2'] / df_total.groupby(['card_id'])['D2'].transform('mean')
#df_total['card_D3_mean'] = df_total['D3'] / df_total.groupby(['card_id'])['D3'].transform('mean')
#df_total['card_D4_mean'] = df_total['D4'] / df_total.groupby(['card_id'])['D4'].transform('mean')
# proved important feature
#df_total['card_D15_mean'] = df_total['D15'] / df_total.groupby(['card_id'])['D15'].transform('mean')
#df_total['card_D1_std'] = df_total['D1'] / df_total.groupby(['card_id'])['D1'].transform('std')
# not so important
#df_total['card_D2_std'] = df_total['D2'] / df_total.groupby(['card_id'])['D2'].transform('std')
#df_total['card_D3_std'] = df_total['D3'] / df_total.groupby(['card_id'])['D3'].transform('std')
#df_total['card_D4_std'] = df_total['D4'] / df_total.groupby(['card_id'])['D4'].transform('std')
# proved important feature
#df_total['card_D15_std'] = df_total['D15'] / df_total.groupby(['card_id'])['D15'].transform('std')
#df_total['addr1_D15_mean'] = df_total['D15'] / df_total.groupby(['addr1'])['D15'].transform('mean')
# proved important feature
#df_total['addr1_D15_std'] = df_total['D15'] / df_total.groupby(['addr1'])['D15'].transform('std')
# proved important feature
# decimal part of the transaction amount
#df_total['TransactionAmt_decimal'] = ((df_total['TransactionAmt'] - df_total['TransactionAmt'].astype(int)) * 1000).astype(int)
#df_total['Device_name'] = df_total['DeviceInfo'].str.split('/', expand=True)[0]
#df_total['Device_version'] = df_total['DeviceInfo'].str.split('/', expand=True)[1]
#df_total.drop('DeviceInfo',axis=1,inplace = True)
#df_total.loc[df_total['Device_name'].str.contains('SM', na=False), 'Device_name'] = 'Samsung'
#df_total.loc[df_total['Device_name'].str.contains('SAMSUNG', na=False), 'Device_name'] = 'Samsung'
#df_total.loc[df_total['Device_name'].str.contains('GT-', na=False), 'Device_name'] = 'Samsung'
#df_total.loc[df_total['Device_name'].str.contains('Moto G', na=False), 'Device_name'] = 'Motorola'
#df_total.loc[df_total['Device_name'].str.contains('Moto', na=False), 'Device_name'] = 'Motorola'
#df_total.loc[df_total['Device_name'].str.contains('moto', na=False), 'Device_name'] = 'Motorola'
#df_total.loc[df_total['Device_name'].str.contains('LG-', na=False), 'Device_name'] = 'LG'
#df_total.loc[df_total['Device_name'].str.contains('rv:', na=False), 'Device_name'] = 'RV'
#df_total.loc[df_total['Device_name'].str.contains('HUAWEI', na=False), 'Device_name'] = 'Huawei'
#df_total.loc[df_total['Device_name'].str.contains('ALE-', na=False), 'Device_name'] = 'Huawei'
#df_total.loc[df_total['Device_name'].str.contains('-L', na=False), 'Device_name'] = 'Huawei'
#df_total.loc[df_total['Device_name'].str.contains('Blade', na=False), 'Device_name'] = 'ZTE'
#df_total.loc[df_total['Device_name'].str.contains('BLADE', na=False), 'Device_name'] = 'ZTE'
#df_total.loc[df_total['Device_name'].str.contains('XT', na=False), 'Device_name'] = 'Sony'
#df_total.loc[df_total.Device_name.isin(df_total.Device_name.value_counts()[df_total.Device_name.value_counts() < 200].index), 'Device_name'] = "Others"
# not so important
#df_total['card_freq_Device'] = df_total.groupby('card_id')['Device_name'].transform(lambda x: x.value_counts(dropna=False).index[0])
#df_total['is_card_freq_Device'] = (df_total.Device_name == df_total.card_freq_Device)
#df_total['is_wide'] = df_total['screen_width'] > df_total['screen_height']
#df_total['is_long'] = df_total['screen_width'] < df_total['screen_height']
#df_total['is_zero'] = (df_total['screen_width'] == 0)
# this feature lead to over fitting
df_total.drop('card_id',axis=1,inplace=True)
#df_total.drop('hour',axis=1,inplace=True)
df_total.drop('DT',axis=1,inplace=True)
df_total.drop('DT_hour',axis=1,inplace=True)
df_total.drop('DT_day',axis=1,inplace=True)
df_total.drop('DT_month',axis=1,inplace=True)
df_total.drop('DT_year',axis=1,inplace=True)
#df_total.drop(['card_freq_Device'],axis=1,inplace=True)
```
https://www.kaggle.com/davidcairuz/feature-engineering-lightgbm-corrected
# Remove Features
```
high_miss_cols = high_missing_cols(df_total,0.9)
one_value_cols = [col for col in df_total.columns
if df_total[col].nunique() <= 1]
big_top_value_cols = [col for col in df_total.columns
if df_total[col].value_counts(dropna=False, normalize=True).values[0] > 0.9]
cols_to_drop = list(set(high_miss_cols + one_value_cols + big_top_value_cols))
len(cols_to_drop)
df_total.drop(cols_to_drop, axis=1, inplace=True)
```
# Deal With NA
```
#cols_to_fill = list(set(df_total.columns)-set(['isFraud']))
#for col in cols_to_fill:
# df_total[col].fillna(-999, inplace=True)
#df_total.tail()
```
# Encoder Categorical
```
df_total_final,colname = label_encoder(df_total, categorical_columns=None)
df_total_final.shape
```
# Save Final Features
```
#df_total_final.to_csv('./data/features476.csv', index = False)
with open('./data/features450.pickle', 'wb') as handle:
pickle.dump(df_total, handle, protocol=pickle.HIGHEST_PROTOCOL)
```
| github_jupyter |
# Transfer Learning
A Convolutional Neural Network (CNN) for image classification is made up of multiple layers that extract features, such as edges, corners, etc; and then use a final fully-connected layer to classify objects based on these features. You can visualize this like this:
<table>
<tr><td rowspan=2 style='border: 1px solid black;'>⇒</td><td style='border: 1px solid black;'>Convolutional Layer</td><td style='border: 1px solid black;'>Pooling Layer</td><td style='border: 1px solid black;'>Convolutional Layer</td><td style='border: 1px solid black;'>Pooling Layer</td><td style='border: 1px solid black;'>Fully Connected Layer</td><td rowspan=2 style='border: 1px solid black;'>⇒</td></tr>
<tr><td colspan=4 style='border: 1px solid black; text-align:center;'>Feature Extraction</td><td style='border: 1px solid black; text-align:center;'>Classification</td></tr>
</table>
*Transfer Learning* is a technique where you can take an existing trained model and re-use its feature extraction layers, replacing its final classification layer with a fully-connected layer trained on your own custom images. With this technique, your model benefits from the feature extraction training that was performed on the base model (which may have been based on a larger training dataset than you have access to) to build a classification model for your own specific set of object classes.
How does this help? Well, think of it this way. Suppose you take a professional tennis player and a complete beginner, and try to teach them both how to play raquetball. It's reasonable to assume that the professional tennis player will be easier to train, because many of the underlying skills involved in raquetball are already learned. Similarly, a pre-trained CNN model may be easier to train to classify specific set of objects because it's already learned how to identify the features of common objects, such as edges and corners. Fundamentally, a pre-trained model can be a great way to produce an effective classifier even when you have limited data with which to train it.
In this notebook, we'll see how to implement transfer learning for a classification model using PyTorch.
## Install and import libraries
First, let's install and import the PyTorch libraries we're going to use.
```
!pip install torch==1.6.0+cpu torchvision==0.7.0+cpu -f https://download.pytorch.org/whl/torch_stable.html
# Import PyTorch libraries
import torch
import torchvision
import torchvision.transforms as transforms
import torch.nn as nn
import torch.optim as optim
from torch.autograd import Variable
import torch.nn.functional as F
# Other libraries we'll use
import numpy as np
import os
import matplotlib.pyplot as plt
%matplotlib inline
print("Libraries imported - ready to use PyTorch", torch.__version__)
```
## Prepare the base model
To use transfer learning, we need a base model from which we can use the trained feature extraction layers. The ***resnet*** model is an CNN-based image classifier that has been pre-trained using a huge dataset containing a large number of images of 1000 classes of object, so let's download it and take a look at its layers.
```
# Load the model (download if not already present)
model = torchvision.models.resnet34(pretrained=True)
print(model)
```
## Prepare the image data
The pretrained model has many layers, starting with a convolutional layer that starts the feature extraction process from image data, and ending with a fully-connected linear layer that maps the extracted features to 1000 class labels.
For feature extraction to work with our own images, we need to ensure that the image data we use the train our prediction layer has the same number of features (pixel values) as the images originally used to train the feaure extraction layers. The model does not explicitly give this size, but the first convolutional layer applies by a 7x7 kernel with a stride of 2x2 and results in 64 feature values, so the original size must be 64 x (7 ÷ 2), which is 224.
PyTorch includes functions for loading and transforming data. We'll use these to create an iterative loader for training data, and a second iterative loader for test data (which we'll use to validate the trained model). The loaders will transform the image data to match the format used to train the original resnet CNN model, convert the image data into *tensors* (which are the core data structure used in PyTorch), and normalize them.
Run the following cell to define the data loaders and list the classes for our images.
```
# Function to ingest data using training and test loaders
def load_dataset(data_path):
# Resize to 256 x 256, then center-crop to 224x224 (to match the resnet image size)
transformation = transforms.Compose([
transforms.Resize(256),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize(mean=[0.5, 0.5, 0.5], std=[0.5, 0.5, 0.5])
])
# Load all of the images, transforming them
full_dataset = torchvision.datasets.ImageFolder(
root=data_path,
transform=transformation
)
# Split into training (70%) and testing (30%) datasets)
train_size = int(0.7 * len(full_dataset))
test_size = len(full_dataset) - train_size
train_dataset, test_dataset = torch.utils.data.random_split(full_dataset, [train_size, test_size])
# define a loader for the training data we can iterate through in 30-image batches
train_loader = torch.utils.data.DataLoader(
train_dataset,
batch_size=30,
num_workers=0,
shuffle=False
)
# define a loader for the testing data we can iterate through in 30-image batches
test_loader = torch.utils.data.DataLoader(
test_dataset,
batch_size=30,
num_workers=0,
shuffle=False
)
return train_loader, test_loader
# Now load the images from the shapes folder
import os
data_path = 'data/shapes/'
# Get the iterative dataloaders for test and training data
train_loader, test_loader = load_dataset(data_path)
# Get the class names
classes = os.listdir(data_path)
classes.sort()
print('class names:', classes)
```
## Create a prediction layer
We downloaded the complete *resnet* model including its final **fc** linear layer. This fully-connected linear layer takes 512 inputs (the extracted features) and produces 1000 outputs (class predictions based on the original training image classes). We need to replace this layer with one that takes the same number of inputs (so we can use the same number of extracted features), but produces a prediction for each of our image classes.
We also need to freeze the feature extraction layers to retain the trained weights. Then when we train the model using our images, only the final prediction layer will learn new weight and bias values - the pre-trained weights already learned for feature extraction will remain the same.
```
# Set the existing feature extraction layers to read-only
for param in model.parameters():
param.requires_grad = False
# Replace the prediction layer
num_ftrs = model.fc.in_features
model.fc = nn.Linear(num_ftrs, len(classes))
# Now print the full model, which will include the feature extraction layers of the base model and our prediction layer
print(model)
```
## Train the model
With the layers of the CNN defined, we're ready to train it using our image data. The weights used in the feature extraction layers from the base resnet model will not be changed by training, only the final linear layer that maps the features to our shape classes will be trained.
```
def train(model, device, train_loader, optimizer, epoch):
# Set the model to training mode
model.train()
train_loss = 0
print("Epoch:", epoch)
# Process the images in batches
for batch_idx, (data, target) in enumerate(train_loader):
# Use the CPU or GPU as appropriate
data, target = data.to(device), target.to(device)
# Reset the optimizer
optimizer.zero_grad()
# Push the data forward through the model layers
output = model(data)
# Get the loss
loss = loss_criteria(output, target)
# Keep a running total
train_loss += loss.item()
# Backpropagate
loss.backward()
optimizer.step()
# Print metrics for every 10 batches so we see some progress
if batch_idx % 10 == 0:
print('Training set [{}/{} ({:.0f}%)] Loss: {:.6f}'.format(
batch_idx * len(data), len(train_loader.dataset),
100. * batch_idx / len(train_loader), loss.item()))
# return average loss for the epoch
avg_loss = train_loss / (batch_idx+1)
print('Training set: Average loss: {:.6f}'.format(avg_loss))
return avg_loss
def test(model, device, test_loader):
# Switch the model to evaluation mode (so we don't backpropagate or drop)
model.eval()
test_loss = 0
correct = 0
with torch.no_grad():
batch_count = 0
for data, target in test_loader:
batch_count += 1
data, target = data.to(device), target.to(device)
# Get the predicted classes for this batch
output = model(data)
# Calculate the loss for this batch
test_loss += loss_criteria(output, target).item()
# Calculate the accuracy for this batch
_, predicted = torch.max(output.data, 1)
correct += torch.sum(target==predicted).item()
# Calculate the average loss and total accuracy for this epoch
avg_loss = test_loss/batch_count
print('Validation set: Average loss: {:.6f}, Accuracy: {}/{} ({:.0f}%)\n'.format(
avg_loss, correct, len(test_loader.dataset),
100. * correct / len(test_loader.dataset)))
# return average loss for the epoch
return avg_loss
# Now use the train and test functions to train and test the model
device = "cpu"
if (torch.cuda.is_available()):
# if GPU available, use cuda (on a cpu, training will take a considerable length of time!)
device = "cuda"
print('Training on', device)
# Create an instance of the model class and allocate it to the device
model = model.to(device)
# Use an "Adam" optimizer to adjust weights
# (see https://pytorch.org/docs/stable/optim.html#algorithms for details of supported algorithms)
optimizer = optim.Adam(model.parameters(), lr=0.001)
# Specify the loss criteria
loss_criteria = nn.CrossEntropyLoss()
# Track metrics in these arrays
epoch_nums = []
training_loss = []
validation_loss = []
# Train over 3 epochs (in a real scenario, you'd likely use many more)
epochs = 3
for epoch in range(1, epochs + 1):
train_loss = train(model, device, train_loader, optimizer, epoch)
test_loss = test(model, device, test_loader)
epoch_nums.append(epoch)
training_loss.append(train_loss)
validation_loss.append(test_loss)
```
## View the loss history
We tracked average training and validation loss for each epoch. We can plot these to verify that the loss reduced over the training process and to detect *over-fitting* (which is indicated by a continued drop in training loss after validation loss has levelled out or started to increase).
```
%matplotlib inline
from matplotlib import pyplot as plt
plt.plot(epoch_nums, training_loss)
plt.plot(epoch_nums, validation_loss)
plt.xlabel('epoch')
plt.ylabel('loss')
plt.legend(['training', 'validation'], loc='upper right')
plt.show()
```
## Evaluate model performance
We can see the final accuracy based on the test data, but typically we'll want to explore performance metrics in a little more depth. Let's plot a confusion matrix to see how well the model is predicting each class.
```
#Pytorch doesn't have a built-in confusion matrix metric, so we'll use SciKit-Learn
from sklearn.metrics import confusion_matrix
import matplotlib.pyplot as plt
%matplotlib inline
# Set the model to evaluate mode
model.eval()
# Get predictions for the test data and convert to numpy arrays for use with SciKit-Learn
print("Getting predictions from test set...")
truelabels = []
predictions = []
for data, target in test_loader:
for label in target.cpu().data.numpy():
truelabels.append(label)
for prediction in model.cpu()(data).data.numpy().argmax(1):
predictions.append(prediction)
# Plot the confusion matrix
cm = confusion_matrix(truelabels, predictions)
plt.imshow(cm, interpolation="nearest", cmap=plt.cm.Blues)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
plt.xlabel("Actual Shape")
plt.ylabel("Predicted Shape")
plt.show()
```
## Use the trained model
Now that we've trained the model, we can use it to predict the class of an image.
```
# Function to create a random image (of a square, circle, or triangle)
def create_image (size, shape):
from random import randint
import numpy as np
from PIL import Image, ImageDraw
xy1 = randint(10,40)
xy2 = randint(60,100)
col = (randint(0,200), randint(0,200), randint(0,200))
img = Image.new("RGB", size, (255, 255, 255))
draw = ImageDraw.Draw(img)
if shape == 'circle':
draw.ellipse([(xy1,xy1), (xy2,xy2)], fill=col)
elif shape == 'triangle':
draw.polygon([(xy1,xy1), (xy2,xy2), (xy2,xy1)], fill=col)
else: # square
draw.rectangle([(xy1,xy1), (xy2,xy2)], fill=col)
del draw
return img
# Function to predict the class of an image
def predict_image(classifier, image):
import numpy
# Set the classifer model to evaluation mode
classifier.eval()
# Apply the same transformations as we did for the training images
transformation = transforms.Compose([
transforms.Resize(256),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize(mean=[0.5, 0.5, 0.5], std=[0.5, 0.5, 0.5])
])
# Preprocess the image
image_tensor = transformation(image).float()
# Add an extra batch dimension since pytorch treats all inputs as batches
image_tensor = image_tensor.unsqueeze_(0)
# Turn the input into a Variable
input_features = Variable(image_tensor)
# Predict the class of the image
output = classifier(input_features)
index = output.data.numpy().argmax()
return index
# Now let's try it with a new image
from random import randint
from PIL import Image
import os, shutil
# Create a random test image
shape = classes[randint(0, len(classes)-1)]
img = create_image ((128,128), shape)
# Display the image
plt.imshow(img)
index = predict_image(model, img)
print(classes[index])
```
## Learn more
* [PyTorch Documentation](https://pytorch.org/tutorials/beginner/transfer_learning_tutorial.html)
| github_jupyter |
```
var = 3
print(var)
var = 7
var
arr1 = []
type(arr1)
arr2 = [1,2,3,4,5]
type(arr2)
len(arr2)
dir(arr1)
print(arr1)
arr1.append(3)
arr1
arr1.append(4)
arr1
arr1.append(5)
arr1.insert(3,2)
arr1
dir(arr1.insert)
arr3 = [1,3,4,'Winner','Emeto',4,6,4]
arr3
arr3.count(4)
arr3.index(3)
def hi():
print('Hello Fellows!')
hi()
class Hi():
def __init__(self):
pass
def hi(self):
value = 'Hello Fellow!'
return value
do = Hi()
print(do.hi())
value = do.hi()
print(value)
dic = {}
type(dic)
dic1 = {1:'Winner', 2:'Emeto'}
dir(dic1)
dic1[1]
dic1[2]
dic1.keys()
con = list()
con
dic1[3] = 'Mercy'
dic1
dic1.values()
dic1.keys()
dic1.items()
dic1.pop(1)
dic1
```
### Build a calculator app that sums, multiply, and divides two numbers
Functional Requirement:
Add two numbers;
Multipy two numbers;
Divides two numbers
```
data_bank = {}
num1 = float(input('Enter First Number: '))
num2 = float(input('Enter Second Number: '))
def add(num1,num2):
result = num1+num2
data_bank['Add'] = result
return result
add(num1,num2)
def mul(num1,num2):
result = num1*num2
data_bank['Mul'] = result
return result
mul(num1,num2)
def div(num1,num2):
result = num1/num2
data_bank['Div'] = result
return result
div(num1,num2)
data_bank
import time
name = input('Enter your name: ')
def execute():
def add(num1,num2):
result = num1+num2
data_bank['Add'] = result
return result
def mul(num1,num2):
result = num1*num2
data_bank['Mul'] = result
return result
def div(num1,num2):
result = num1/num2
data_bank['Div'] = result
return result
add(num1,num2)
mul(num1,num2)
div(num1,num2)
def main(name):
print('Hi '+name+', welcome to my calculator app! ')
print('==========================================')
num1 = float(input('Enter First Number: '))
num2 = float(input('Enter Second Number: '))
execute()
print('Calculating valuse....')
time.sleep(10)
print('Values after calculations is: ',data_bank)
main(name)
for key in data_bank.keys():
print(key)
for key in data_bank.values():
print(key)
data_bank.items()
for key,value in data_bank.items():
print(key+' '+str(value))
counter = 5
while counter >= 1:
print('True')
counter -= 1
if 0:
print(True)
else:
print(False)
age = int(input('Enter your age: '))
if age == 18:
print('Welcome')
elif age > 18:
print('You are welcome')
elif age >= 100:
print('Please you are too old for this!')
else:
print('Please get out of here, you are not old enough')
import this
def hello():
print('Hello World!!')
hello()
print(hello())
def hello():
result = 'Hello World'
print(result)
return result
hello()
print(hello())
var = hello()
print(var)
def add():
num1 = 2
num2 = 3
result = num1 + num2
return num1,num2,result
num1,num2,result = add()
print(result)
def add(num1,num2):
result = num1 + num2
return num1,num2,result
add(5,7)
num1,num2,result = add(5,7)
num1
num2
result
```
| github_jupyter |
# [Sensor name]
:::{eval-rst}
:opticon:`tag`
:badge:`[Environment],badge-primary`
:badge:`Sensors,badge-secondary`
:::
## Context
### Purpose
*Describe the purpose of the use case.*
### Sensor description
*Describe the main features of the sensor e.g. variables.*
### Highlights
*Provide 3-5 bullet points that convey the use case’s core procedures. Each bullet point must have a maximum of 85 characters, including spaces.*
* Highlight 1
* Highlight 2
### Contributions
#### Notebook
Author (role), Affiliation, GitHub alias
#### Dataset originator/creator
Institution/Community/Individual (affiliation)
#### Dataset authors
Institution/Community/Individual (affiliation)
#### Dataset documentation
```{bibliography}
:style: plain
:list: bullet
:filter: topic % "replace by the `topic` entry linked to the publication(s) in the `_bibliography/references.bib` file"
```
:::{note}
*Optional: add credits or acknowledgements to data providers or authors of code snippets*
:::
## Install and load libraries
*For installation, add only libraries not listed in the [environment.yml](https://github.com/alan-turing-institute/environmental-ds-book/blob/master/environment.yml) file, but required by the notebook. Libraries can be installed in silent mode e.g. `pip -q install <package_name>`*
*For loading libraries, order them according to their role e.g. libraries to manipulate folders i.e. os (first), handle data i.e. numpy, xarray (second), visualisation e.g. holoviews (third), etc. The cell below contains two libraries, `os` and `warning` which are common among the notebooks. Don't remove them.*
```
import os
import warnings
warnings.filterwarnings(action='ignore')
```
## Set project structure
*The cell below creates a separate folder to save the notebook outputs. This facilitates the reader to inspect inputs/outputs stored within a defined destination folder. Change `<replace-by-notebook-filename>` with your notebook identifier.*
```
notebook_folder = '../sensors/<replace-by-notebook-filename>'
if not os.path.exists(notebook_folder):
os.makedirs(notebook_folder)
```
## Load data
*Load full dataset from original or mirror sources. If the license of the dataset permits, we suggest creating sample data (preprocessed) for the notebook stored in a data repository e.g. Zenodo.*
## Visualisation
*Create a visual narrative of the dataset! We suggest exploring libraries suited for interactive plotting e.g. Holoviews, Panel, Bokeh.*
## Summary
*Provide 3-5 bullet points summarising the main aspects of the dataset and tools covered in the notebook.*
* Sentence 1 e.g. `tool-name` to perform...
* Sentence 2 e.g. `tool-name` to perform...
## Additional information
**Dataset**: Type here details of dataset(s) version.
**License**: The code in this notebook is licensed under the MIT License. The Environmental Data Science book is licensed under the Creative Commons by Attribution 4.0 license. See further details [here](https://github.com/alan-turing-institute/environmental-ds-book/blob/master/LICENSE.md).
**Contact**: If you have any suggestion or report an issue with this notebook, feel free to [create an issue](https://github.com/alan-turing-institute/environmental-ds-book/issues/new/choose) or send a direct message to [environmental.ds.book@gmail.com](mailto:environmental.ds.book@gmail.com).
```
from datetime import date
print(f'Last tested: {date.today()}')
```
| github_jupyter |
```
%pylab inline
from simqso.sqgrids import *
from simqso import sqbase
from simqso.sqmodels import QLF_McGreer_2013
# set up a luminosity-redshift grid
M = AbsMagVar(UniformSampler(-30,-25),restWave=1450)
z = RedshiftVar(UniformSampler(1,5))
MzGrid = QsoSimGrid([M,z],(4,3),2,seed=12345)
scatter(MzGrid.z,MzGrid.absMag,)
xlabel('z')
ylabel('absolute mag');
# set up a flux-redshift grid with Lya EW as a third dimension
m = AppMagVar(UniformSampler(17,22),'SDSS-i')
lya = GaussianLineEqWidthVar(UniformSampler(0,2),'LyaEW',1215.67,10.)
mzlyaGrid = QsoSimGrid([m,z,lya],(4,3,2),2,seed=12345)
scatter(mzlyaGrid.z,mzlyaGrid.appMag,c=mzlyaGrid.LyaEW)
cb = colorbar()
xlabel('z')
ylabel('apparent mag')
cb.set_label('log EW(Lya)')
# Generate m,z points by sampling from a model QLF
cosmo = QLF_McGreer_2013.cosmo
kcorr = sqbase.ContinuumKCorr('SDSS-i',1450)
qlfGrid = generateQlfPoints(QLF_McGreer_2013,
(17,22),(4.5,5.3),kcorr,
skyArea=100,
qlfseed=12345,gridseed=67890)
scatter(qlfGrid.z,qlfGrid.appMag)
xlabel('z')
ylabel('apparent mag')
# test the canonical values for power law continuum slopes in FUV/NUV
contVar = BrokenPowerLawContinuumVar([GaussianSampler(-1.5,0.3),
GaussianSampler(-0.5,0.3)],[1215.],
seed=12345)
contVals = contVar(1000)
_ = hist(contVals[:,0],alpha=0.8)
_ = hist(contVals[:,1],alpha=0.8)
# Add gaussian emission lines
g = GaussianSampler
lya = [g(1215.7,0.5),g(80,20),g(7,2)]
civ = [g(1550.0,1.5),g(20,10),g(10,2)]
emLinesVar = GaussianEmissionLinesTemplateVar([lya,civ],seed=12345)
emLinesVals = emLinesVar(1000)
figure(figsize=(14,4))
for k in range(3):
subplot(1,3,k+1)
for j in range(2):
hist(emLinesVals[:,j,k],alpha=0.8)
# add gaussian lines to the qlf grid
qlfGrid.addVar(emLinesVar)
print(qlfGrid)
scatter(qlfGrid.z,qlfGrid.appMag,c=qlfGrid.emLines[:,0,1])
cb = colorbar()
xlabel('z')
ylabel('apparent mag')
cb.set_label('EW(Lya)')
# Now use the BOSS DR9 emission line template, including the Baldwin Effect
lineTemplate = generateBEffEmissionLines(qlfGrid.absMag,seed=12345)
lines = lineTemplate()
lines.shape
scatter(qlfGrid.z,qlfGrid.appMag,c=lines[:,13,1])
cb = colorbar()
xlabel('z')
ylabel('apparent mag')
cb.set_label('EW(CIV)')
vdblines = generateVdBCompositeEmLines(minEW=0)
vdblines = vdblines(qlfGrid.nObj)
wave = arange(3000,1e4,0.5)
z = 2.0
def quick_spec(emlines):
lineWave,eqWidth,sigma = emlines.T * (1+z)
spec = np.zeros_like(wave)
A = eqWidth/(np.sqrt(2*np.pi)*sigma)
twosig2 = 2*sigma**2
nsig = np.sqrt(-2*np.log(1e-3/A))
nsig = (nsig*np.array([[-1.],[1]])).T
for i in range(lineWave.shape[0]):
i1,i2 = np.searchsorted(wave,lineWave[i]+nsig[i]*sigma[i])
if i2 != i1:
lineprofile = A[i]*np.exp(-(wave[i1:i2]-lineWave[i])**2
/ twosig2[i])
spec[i1:i2] += lineprofile
return spec
figure(figsize=(12,4))
plot(wave/(1+z),quick_spec(lines[0]),label='BOSS DR9')
plot(wave/(1+z),quick_spec(vdblines[0]),label='Vanden Berk composite')
legend()
```
| github_jupyter |
# <div align="center">Credit Fraud Detector</div>
---------------------------------------------------------------------
you can find the kernel link below:
> ###### [ Kaggle](https://www.kaggle.com/janiobachmann/credit-fraud-dealing-with-imbalanced-datasets)
## Introduction
In this kernel we will use various predictive models to see how accurate they are in detecting whether a transaction is a normal payment or a fraud. As described in the dataset, the features are scaled and the names of the features are not shown due to privacy reasons. Nevertheless, we can still analyze some important aspects of the dataset. Let's start!
## Our Goals:
* Understand the little distribution of the "little" data that was provided to us.
* Create a 50/50 sub-dataframe ratio of "Fraud" and "Non-Fraud" transactions. (NearMiss Algorithm)
* Determine the Classifiers we are going to use and decide which one has a higher accuracy.
* Create a Neural Network and compare the accuracy to our best classifier.
* Understand common mistaked made with imbalanced datasets.
## Outline:
I. Understanding our data
a) Gather Sense of our data
II. Preprocessing
a) Scaling and Distributing
b) Splitting the Data
III. Random UnderSampling and Oversampling
a) Distributing and Correlating
b) Anomaly Detection
c) Dimensionality Reduction and Clustering (t-SNE)
d) Classifiers
e) A Deeper Look into Logistic Regression
f) Oversampling with SMOTE
IV. Testing
a) Testing with Logistic Regression
b) Neural Networks Testing (Undersampling vs Oversampling)
## Correcting Previous Mistakes from Imbalanced Datasets:
* Never test on the oversampled or undersampled dataset.
* If we want to implement cross validation, remember to oversample or undersample your training data during cross-validation, not before!
* Don't use accuracy score as a metric with imbalanced datasets (will be usually high and misleading), instead use f1-score, precision/recall score or confusion matrix
## References:
* Hands on Machine Learning with Scikit-Learn & TensorFlow by Aurélien Géron (O'Reilly). CopyRight 2017 Aurélien Géron
* Machine Learning - Over-& Undersampling - Python/ Scikit/ Scikit-Imblearn by Coding-Maniac
* auprc, 5-fold c-v, and resampling methods by Jeremy Lane (Kaggle Notebook)
# <div align="center">Gather Sense of Our Data:</div>
---------------------------------------------------------------------
The first thing we must do is gather a basic sense of our data. Remember, except for the transaction and amount we dont know what the other columns are (due to privacy reasons). The only thing we know, is that those columns that are unknown have been scaled already.
## Summary:
* The transaction amount is relatively small. The mean of all the mounts made is approximately USD 88.
* There are no "Null" values, so we don't have to work on ways to replace values.
* Most of the transactions were Non-Fraud (99.83%) of the time, while Fraud transactions occurs (017%) of the time in the dataframe.
## Feature Technicalities:
* PCA Transformation: The description of the data says that all the features went through a PCA transformation (Dimensionality Reduction technique) (Except for time and amount).
* Scaling: Keep in mind that in order to implement a PCA transformation features need to be previously scaled. (In this case, all the V features have been scaled or at least that is what we are assuming the people that develop the dataset did.)
```
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load in
# Imported Libraries
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import tensorflow as tf
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.manifold import TSNE
from sklearn.decomposition import PCA, TruncatedSVD
import matplotlib.patches as mpatches
import time
# Classifier Libraries
from sklearn.linear_model import LogisticRegression
from sklearn.svm import SVC
from sklearn.neighbors import KNeighborsClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
import collections
# Other Libraries
from imblearn.datasets import fetch_datasets
from sklearn.model_selection import train_test_split
from sklearn.pipeline import make_pipeline
from imblearn.pipeline import make_pipeline as imbalanced_make_pipeline
from imblearn.over_sampling import SMOTE
from imblearn.under_sampling import NearMiss
from imblearn.metrics import classification_report_imbalanced
from sklearn.metrics import precision_score, recall_score, f1_score, roc_auc_score, accuracy_score, classification_report
from collections import Counter
from sklearn.model_selection import KFold, StratifiedKFold
import warnings
warnings.filterwarnings("ignore")
df = pd.read_csv('input/creditcard.csv')
df.head()
df.describe()
# Good No Null Values!
df.isnull().sum().max()
df.columns
# The classes are heavily skewed we need to solve this issue later.
print('No Frauds', round(df['Class'].value_counts()[0]/len(df) * 100,2), '% of the dataset')
print('Frauds', round(df['Class'].value_counts()[1]/len(df) * 100,2), '% of the dataset')
```
***Note:*** Notice how imbalanced is our original dataset! Most of the transactions are non-fraud. If we use this dataframe as the base for our predictive models and analysis we might get a lot of errors and our algorithms will probably overfit since it will "assume" that most transactions are not fraud. But we don't want our model to assume, we want our model to detect patterns that give signs of fraud!
```
colors = ["#0101DF", "#DF0101"]
sns.countplot('Class', data=df, palette=colors)
plt.title('Class Distributions \n (0: No Fraud || 1: Fraud)', fontsize=14);
```
***Distributions:*** By seeing the distributions we can have an idea how skewed are these features, we can also see further distributions of the other features. There are techniques that can help the distributions be less skewed which will be implemented in this notebook in the future.
```
fig, ax = plt.subplots(1, 2, figsize=(18,4))
amount_val = df['Amount'].values
time_val = df['Time'].values
sns.distplot(amount_val, ax=ax[0], color='r')
ax[0].set_title('Distribution of Transaction Amount', fontsize=14)
ax[0].set_xlim([min(amount_val), max(amount_val)])
sns.distplot(time_val, ax=ax[1], color='b')
ax[1].set_title('Distribution of Transaction Time', fontsize=14)
ax[1].set_xlim([min(time_val), max(time_val)])
plt.show();
```
## Scaling and Distributing
In this phase of our kernel, we will first ***scale the columns comprise of Time and Amount*** . Time and amount should be scaled as the other columns. On the other hand, we need to also create a sub sample of the dataframe in order to have an equal amount of Fraud and Non-Fraud cases, helping our algorithms better understand patterns that determines whether a transaction is a fraud or not.
## What is a sub-Sample?
In this scenario, our subsample will be a dataframe with a 50/50 ratio of fraud and non-fraud transactions. Meaning our sub-sample will have the same amount of fraud and non fraud transactions.
## Why do we create a sub-Sample?
In the beginning of this notebook we saw that the original dataframe was heavily imbalanced! Using the original dataframe will cause the following issues:
* Overfitting: Our classification models will assume that in most cases there are no frauds! What we want for our model is to be certain when a fraud occurs.
* Wrong Correlations: Although we don't know what the "V" features stand for, it will be useful to understand how each of this features influence the result (Fraud or No Fraud) by having an imbalance dataframe we are not able to see the true correlations between the class and features.
## Summary:
* Scaled amount and scaled time are the columns with scaled values.
* There are 492 cases of fraud in our dataset so we can randomly get 492 cases of non-fraud to create our new sub dataframe.
* We concat the 492 cases of fraud and non fraud, creating a new sub-sample.
```
# Since most of our data has already been scaled we should scale the columns that are left to scale (Amount and Time)
from sklearn.preprocessing import StandardScaler, RobustScaler
# RobustScaler is less prone to outliers.
std_scaler = StandardScaler()
rob_scaler = RobustScaler()
df['scaled_amount'] = rob_scaler.fit_transform(df['Amount'].values.reshape(-1,1))
df['scaled_time'] = rob_scaler.fit_transform(df['Time'].values.reshape(-1,1))
df.drop(['Time','Amount'], axis=1, inplace=True)
scaled_amount = df['scaled_amount']
scaled_time = df['scaled_time']
df.drop(['scaled_amount', 'scaled_time'], axis=1, inplace=True)
df.insert(0, 'scaled_amount', scaled_amount)
df.insert(1, 'scaled_time', scaled_time)
# Amount and Time are Scaled!
df.head()
```
## Splitting the Data (Original DataFrame)
Before proceeding with the Random UnderSampling technique we have to separate the orginal dataframe. Why? for testing purposes, remember although we are splitting the data when implementing Random UnderSampling or OverSampling techniques, we want to test our models on the original testing set not on the testing set created by either of these techniques. The main goal is to fit the model either with the dataframes that were undersample and oversample (in order for our models to detect the patterns), and test it on the original testing set.
```
from sklearn.model_selection import train_test_split
from sklearn.model_selection import StratifiedShuffleSplit
print('No Frauds', round(df['Class'].value_counts()[0]/len(df) * 100,2), '% of the dataset')
print('Frauds', round(df['Class'].value_counts()[1]/len(df) * 100,2), '% of the dataset')
X = df.drop('Class', axis=1)
y = df['Class']
sss = StratifiedKFold(n_splits=5, random_state=None, shuffle=False)
for train_index, test_index in sss.split(X, y):
print("Train:", train_index, "Test:", test_index)
original_Xtrain, original_Xtest = X.iloc[train_index], X.iloc[test_index]
original_ytrain, original_ytest = y.iloc[train_index], y.iloc[test_index]
# We already have X_train and y_train for undersample data thats why I am using original to distinguish and to not overwrite these variables.
# original_Xtrain, original_Xtest, original_ytrain, original_ytest = train_test_split(X, y, test_size=0.2, random_state=42)
# Check the Distribution of the labels
# Turn into an array
original_Xtrain = original_Xtrain.values
original_Xtest = original_Xtest.values
original_ytrain = original_ytrain.values
original_ytest = original_ytest.values
# See if both the train and test label distribution are similarly distributed
train_unique_label, train_counts_label = np.unique(original_ytrain, return_counts=True)
test_unique_label, test_counts_label = np.unique(original_ytest, return_counts=True)
print('-' * 100)
print('Label Distributions: \n')
print(train_counts_label/ len(original_ytrain))
print(test_counts_label/ len(original_ytest))
```
## Random Under-Sampling:
In this phase of the project we will implement "Random Under Sampling" which basically consists of removing data in order to have a more balanced dataset and thus avoiding our models to overfitting.
## Steps:
* The first thing we have to do is determine how imbalanced is our class (use "value_counts()" on the class column to determine the amount for each label)
* Once we determine how many instances are considered fraud transactions (Fraud = "1") , we should bring the non-fraud transactions to the same amount as fraud transactions (assuming we want a 50/50 ratio), this will be equivalent to 492 cases of fraud and 492 cases of non-fraud transactions.
* After implementing this technique, we have a sub-sample of our dataframe with a 50/50 ratio with regards to our classes. Then the next step we will implement is to shuffle the data to see if our models can maintain a certain accuracy everytime we run this script.
***Note:*** The main issue with "Random Under-Sampling" is that we run the risk that our classification models will not perform as accurate as we would like to since there is a great deal of information loss (bringing 492 non-fraud transaction from 284,315 non-fraud transaction)
```
# Since our classes are highly skewed we should make them equivalent in order to have a normal distribution of the classes.
# Lets shuffle the data before creating the subsamples
df = df.sample(frac=1)
# amount of fraud classes 492 rows.
fraud_df = df.loc[df['Class'] == 1]
non_fraud_df = df.loc[df['Class'] == 0][:492]
normal_distributed_df = pd.concat([fraud_df, non_fraud_df])
# Shuffle dataframe rows
new_df = normal_distributed_df.sample(frac=1, random_state=42)
new_df.head()
```
## Equally Distributing and Correlating:
Now that we have our dataframe correctly balanced, we can go further with our analysis and data preprocessing.
```
print('Distribution of the Classes in the subsample dataset')
print(new_df['Class'].value_counts()/len(new_df))
sns.countplot('Class', data=new_df, palette=colors)
plt.title('Equally Distributed Classes', fontsize=14)
plt.show()
```
## Correlation Matrices
Correlation matrices are the essence of understanding our data. We want to know if there are features that influence heavily in whether a specific transaction is a fraud. However, it is important that we use the correct dataframe (subsample) in order for us to see which features have a high positive or negative correlation with regards to fraud transactions.
## Summary and Explanation:
* Negative Correlations: V17, V14, V12 and V10 are negatively correlated. Notice how the lower these values are, the more likely the end result will be a fraud transaction.
* Positive Correlations: V2, V4, V11, and V19 are positively correlated. Notice how the higher these values are, the more likely the end result will be a fraud transaction.
* BoxPlots: We will use boxplots to have a better understanding of the distribution of these features in fradulent and non fradulent transactions.
***Note:*** We have to make sure we use the subsample in our correlation matrix or else our correlation matrix will be affected by the high imbalance between our classes. This occurs due to the high class imbalance in the original dataframe.
```
# Make sure we use the subsample in our correlation
f, (ax1, ax2) = plt.subplots(2, 1, figsize=(24,20))
# Entire DataFrame
corr = df.corr()
sns.heatmap(corr, cmap='coolwarm_r', annot_kws={'size':20}, ax=ax1)
ax1.set_title("Imbalanced Correlation Matrix \n (don't use for reference)", fontsize=14)
sub_sample_corr = new_df.corr()
sns.heatmap(sub_sample_corr, cmap='coolwarm_r', annot_kws={'size':20}, ax=ax2)
ax2.set_title('SubSample Correlation Matrix \n (use for reference)', fontsize=14)
plt.show()
f, axes = plt.subplots(ncols=4, figsize=(20,4))
# Negative Correlations with our Class (The lower our feature value the more likely it will be a fraud transaction)
sns.boxplot(x="Class", y="V17", data=new_df, palette=colors, ax=axes[0])
axes[0].set_title('V17 vs Class Negative Correlation')
sns.boxplot(x="Class", y="V14", data=new_df, palette=colors, ax=axes[1])
axes[1].set_title('V14 vs Class Negative Correlation')
sns.boxplot(x="Class", y="V12", data=new_df, palette=colors, ax=axes[2])
axes[2].set_title('V12 vs Class Negative Correlation')
sns.boxplot(x="Class", y="V10", data=new_df, palette=colors, ax=axes[3])
axes[3].set_title('V10 vs Class Negative Correlation')
plt.show()
f, axes = plt.subplots(ncols=4, figsize=(20,4))
# Positive correlations (The higher the feature the probability increases that it will be a fraud transaction)
sns.boxplot(x="Class", y="V11", data=new_df, palette=colors, ax=axes[0])
axes[0].set_title('V11 vs Class Positive Correlation')
sns.boxplot(x="Class", y="V4", data=new_df, palette=colors, ax=axes[1])
axes[1].set_title('V4 vs Class Positive Correlation')
sns.boxplot(x="Class", y="V2", data=new_df, palette=colors, ax=axes[2])
axes[2].set_title('V2 vs Class Positive Correlation')
sns.boxplot(x="Class", y="V19", data=new_df, palette=colors, ax=axes[3])
axes[3].set_title('V19 vs Class Positive Correlation')
plt.show()
```
## Anomaly Detection:
Our main aim in this section is to remove "extreme outliers" from features that have a high correlation with our classes. This will have a positive impact on the accuracy of our models.
## Interquartile Range Method:
* Interquartile Range (IQR): We calculate this by the difference between the 75th percentile and 25th percentile. Our aim is to create a threshold beyond the 75th and 25th percentile that in case some instance pass this threshold the instance will be deleted.
* Boxplots: Besides easily seeing the 25th and 75th percentiles (both end of the squares) it is also easy to see extreme outliers (points beyond the lower and higher extreme).
## Outlier Removal Tradeoff:
* We have to be careful as to how far do we want the threshold for removing outliers. We determine the threshold by multiplying a number (ex: 1.5) by the (Interquartile Range). The higher this threshold is, the less outliers will detect (multiplying by a higher number ex: 3), and the lower this threshold is the more outliers it will detect.
* The Tradeoff: The lower the threshold the more outliers it will remove however, we want to focus more on "extreme outliers" rather than just outliers. Why? because we might run the risk of information loss which will cause our models to have a lower accuracy. You can play with this threshold and see how it affects the accuracy of our classification models.
## Summary:
* Visualize Distributions: We first start by visualizing the distribution of the feature we are going to use to eliminate some of the outliers. V14 is the only feature that has a Gaussian distribution compared to features V12 and V10.
* Determining the threshold: After we decide which number we will use to multiply with the iqr (the lower more outliers removed), we will proceed in determining the upper and lower thresholds by substrating q25 - threshold (lower extreme threshold) and adding q75 + threshold (upper extreme threshold).
* Conditional Dropping: Lastly, we create a conditional dropping stating that if the "threshold" is exceeded in both extremes, the instances will be removed.
* Boxplot Representation: Visualize through the boxplot that the number of "extreme outliers" have been reduced to a considerable amount.
***Note:*** After implementing outlier reduction our accuracy has been improved by over 3%! Some outliers can distort the accuracy of our models but remember, we have to avoid an extreme amount of information loss or else our model runs the risk of underfitting.
```
from scipy.stats import norm
f, (ax1, ax2, ax3) = plt.subplots(1,3, figsize=(20, 6))
v14_fraud_dist = new_df['V14'].loc[new_df['Class'] == 1].values
sns.distplot(v14_fraud_dist,ax=ax1, fit=norm, color='#FB8861')
ax1.set_title('V14 Distribution \n (Fraud Transactions)', fontsize=14)
v12_fraud_dist = new_df['V12'].loc[new_df['Class'] == 1].values
sns.distplot(v12_fraud_dist,ax=ax2, fit=norm, color='#56F9BB')
ax2.set_title('V12 Distribution \n (Fraud Transactions)', fontsize=14)
v10_fraud_dist = new_df['V10'].loc[new_df['Class'] == 1].values
sns.distplot(v10_fraud_dist,ax=ax3, fit=norm, color='#C5B3F9')
ax3.set_title('V10 Distribution \n (Fraud Transactions)', fontsize=14)
plt.show()
# # -----> V14 Removing Outliers (Highest Negative Correlated with Labels)
v14_fraud = new_df['V14'].loc[new_df['Class'] == 1].values
q25, q75 = np.percentile(v14_fraud, 25), np.percentile(v14_fraud, 75)
print('Quartile 25: {} | Quartile 75: {}'.format(q25, q75))
v14_iqr = q75 - q25
print('iqr: {}'.format(v14_iqr))
v14_cut_off = v14_iqr * 1.5
v14_lower, v14_upper = q25 - v14_cut_off, q75 + v14_cut_off
print('Cut Off: {}'.format(v14_cut_off))
print('V14 Lower: {}'.format(v14_lower))
print('V14 Upper: {}'.format(v14_upper))
outliers = [x for x in v14_fraud if x < v14_lower or x > v14_upper]
print('Feature V14 Outliers for Fraud Cases: {}'.format(len(outliers)))
print('V10 outliers:{}'.format(outliers))
new_df = new_df.drop(new_df[(new_df['V14'] > v14_upper) | (new_df['V14'] < v14_lower)].index)
print('----' * 44)
# -----> V12 removing outliers from fraud transactions
v12_fraud = new_df['V12'].loc[new_df['Class'] == 1].values
q25, q75 = np.percentile(v12_fraud, 25), np.percentile(v12_fraud, 75)
v12_iqr = q75 - q25
v12_cut_off = v12_iqr * 1.5
v12_lower, v12_upper = q25 - v12_cut_off, q75 + v12_cut_off
print('V12 Lower: {}'.format(v12_lower))
print('V12 Upper: {}'.format(v12_upper))
outliers = [x for x in v12_fraud if x < v12_lower or x > v12_upper]
print('V12 outliers: {}'.format(outliers))
print('Feature V12 Outliers for Fraud Cases: {}'.format(len(outliers)))
new_df = new_df.drop(new_df[(new_df['V12'] > v12_upper) | (new_df['V12'] < v12_lower)].index)
print('Number of Instances after outliers removal: {}'.format(len(new_df)))
print('----' * 44)
# Removing outliers V10 Feature
v10_fraud = new_df['V10'].loc[new_df['Class'] == 1].values
q25, q75 = np.percentile(v10_fraud, 25), np.percentile(v10_fraud, 75)
v10_iqr = q75 - q25
v10_cut_off = v10_iqr * 1.5
v10_lower, v10_upper = q25 - v10_cut_off, q75 + v10_cut_off
print('V10 Lower: {}'.format(v10_lower))
print('V10 Upper: {}'.format(v10_upper))
outliers = [x for x in v10_fraud if x < v10_lower or x > v10_upper]
print('V10 outliers: {}'.format(outliers))
print('Feature V10 Outliers for Fraud Cases: {}'.format(len(outliers)))
new_df = new_df.drop(new_df[(new_df['V10'] > v10_upper) | (new_df['V10'] < v10_lower)].index)
print('Number of Instances after outliers removal: {}'.format(len(new_df)))
f,(ax1, ax2, ax3) = plt.subplots(1, 3, figsize=(20,6))
colors = ['#B3F9C5', '#f9c5b3']
# Boxplots with outliers removed
# Feature V14
sns.boxplot(x="Class", y="V14", data=new_df,ax=ax1, palette=colors)
ax1.set_title("V14 Feature \n Reduction of outliers", fontsize=14)
ax1.annotate('Fewer extreme \n outliers', xy=(0.98, -17.5), xytext=(0, -12),
arrowprops=dict(facecolor='black'),
fontsize=14)
# Feature 12
sns.boxplot(x="Class", y="V12", data=new_df, ax=ax2, palette=colors)
ax2.set_title("V12 Feature \n Reduction of outliers", fontsize=14)
ax2.annotate('Fewer extreme \n outliers', xy=(0.98, -17.3), xytext=(0, -12),
arrowprops=dict(facecolor='black'),
fontsize=14)
# Feature V10
sns.boxplot(x="Class", y="V10", data=new_df, ax=ax3, palette=colors)
ax3.set_title("V10 Feature \n Reduction of outliers", fontsize=14)
ax3.annotate('Fewer extreme \n outliers', xy=(0.95, -16.5), xytext=(0, -12),
arrowprops=dict(facecolor='black'),
fontsize=14)
plt.show()
```
# Dimensionality Reduction and Clustering:
## Understanding t-SNE:
In order to understand this algorithm you have to understand the following terms:
* Euclidean Distance
* Conditional Probability
* Normal and T-Distribution Plots
***Note:*** If you want a simple instructive video look at StatQuest: t-SNE, Clearly Explained by Joshua Starmer
## Summary:
* t-SNE algorithm can pretty accurately cluster the cases that were fraud and non-fraud in our dataset.
* Although the subsample is pretty small, the t-SNE algorithm is able to detect clusters pretty accurately in every scenario (I shuffle the dataset before running t-SNE)
* This gives us an indication that further predictive models will perform pretty well in separating fraud cases from non-fraud cases.
```
# New_df is from the random undersample data (fewer instances)
X = new_df.drop('Class', axis=1)
y = new_df['Class']
# T-SNE Implementation
t0 = time.time()
X_reduced_tsne = TSNE(n_components=2, random_state=42).fit_transform(X.values)
t1 = time.time()
print("T-SNE took {:.2} s".format(t1 - t0))
# PCA Implementation
t0 = time.time()
X_reduced_pca = PCA(n_components=2, random_state=42).fit_transform(X.values)
t1 = time.time()
print("PCA took {:.2} s".format(t1 - t0))
# TruncatedSVD
t0 = time.time()
X_reduced_svd = TruncatedSVD(n_components=2, algorithm='randomized', random_state=42).fit_transform(X.values)
t1 = time.time()
print("Truncated SVD took {:.2} s".format(t1 - t0))
f, (ax1, ax2, ax3) = plt.subplots(1, 3, figsize=(24,6))
# labels = ['No Fraud', 'Fraud']
f.suptitle('Clusters using Dimensionality Reduction', fontsize=14)
blue_patch = mpatches.Patch(color='#0A0AFF', label='No Fraud')
red_patch = mpatches.Patch(color='#AF0000', label='Fraud')
# t-SNE scatter plot
ax1.scatter(X_reduced_tsne[:,0], X_reduced_tsne[:,1], c=(y == 0), cmap='coolwarm', label='No Fraud', linewidths=2)
ax1.scatter(X_reduced_tsne[:,0], X_reduced_tsne[:,1], c=(y == 1), cmap='coolwarm', label='Fraud', linewidths=2)
ax1.set_title('t-SNE', fontsize=14)
ax1.grid(True)
ax1.legend(handles=[blue_patch, red_patch])
# PCA scatter plot
ax2.scatter(X_reduced_pca[:,0], X_reduced_pca[:,1], c=(y == 0), cmap='coolwarm', label='No Fraud', linewidths=2)
ax2.scatter(X_reduced_pca[:,0], X_reduced_pca[:,1], c=(y == 1), cmap='coolwarm', label='Fraud', linewidths=2)
ax2.set_title('PCA', fontsize=14)
ax2.grid(True)
ax2.legend(handles=[blue_patch, red_patch])
# TruncatedSVD scatter plot
ax3.scatter(X_reduced_svd[:,0], X_reduced_svd[:,1], c=(y == 0), cmap='coolwarm', label='No Fraud', linewidths=2)
ax3.scatter(X_reduced_svd[:,0], X_reduced_svd[:,1], c=(y == 1), cmap='coolwarm', label='Fraud', linewidths=2)
ax3.set_title('Truncated SVD', fontsize=14)
ax3.grid(True)
ax3.legend(handles=[blue_patch, red_patch])
plt.show()
```
# Classifiers (UnderSampling):
In this section we will train four types of classifiers and decide which classifier will be more effective in detecting fraud transactions. Before we have to split our data into training and testing sets and separate the features from the labels.
## Summary:
* Logistic Regression classifier is more accurate than the other three classifiers in most cases. (We will further analyze Logistic Regression)
* GridSearchCV is used to determine the paremeters that gives the best predictive score for the classifiers.
* Logistic Regression has the best Receiving Operating Characteristic score (ROC), meaning that LogisticRegression pretty accurately separates fraud and non-fraud transactions.
## Learning Curves:
* The wider the gap between the training score and the cross validation score, the more likely your model is overfitting (high variance).
* If the score is low in both training and cross-validation sets this is an indication that our model is underfitting (high bias)
* Logistic Regression Classifier shows the best score in both training and cross-validating sets.
```
# Undersampling before cross validating (prone to overfit)
X = new_df.drop('Class', axis=1)
y = new_df['Class']
# Our data is already scaled we should split our training and test sets
from sklearn.model_selection import train_test_split
# This is explicitly used for undersampling.
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42)
# Turn the values into an array for feeding the classification algorithms.
X_train = X_train.values
X_test = X_test.values
y_train = y_train.values
y_test = y_test.values
# Let's implement simple classifiers
classifiers = {
"LogisiticRegression": LogisticRegression(),
"KNearest": KNeighborsClassifier(),
"Support Vector Classifier": SVC(),
"DecisionTreeClassifier": DecisionTreeClassifier()
}
# Wow our scores are getting even high scores even when applying cross validation.
from sklearn.model_selection import cross_val_score
for key, classifier in classifiers.items():
classifier.fit(X_train, y_train)
training_score = cross_val_score(classifier, X_train, y_train, cv=5)
print("Classifiers: ", classifier.__class__.__name__, "Has a training score of", round(training_score.mean(), 2) * 100, "% accuracy score")
# Use GridSearchCV to find the best parameters.
from sklearn.model_selection import GridSearchCV
# Logistic Regression
log_reg_params = {"penalty": ['l1', 'l2'], 'C': [0.001, 0.01, 0.1, 1, 10, 100, 1000]}
grid_log_reg = GridSearchCV(LogisticRegression(), log_reg_params)
grid_log_reg.fit(X_train, y_train)
# We automatically get the logistic regression with the best parameters.
log_reg = grid_log_reg.best_estimator_
knears_params = {"n_neighbors": list(range(2,5,1)), 'algorithm': ['auto', 'ball_tree', 'kd_tree', 'brute']}
grid_knears = GridSearchCV(KNeighborsClassifier(), knears_params)
grid_knears.fit(X_train, y_train)
# KNears best estimator
knears_neighbors = grid_knears.best_estimator_
# Support Vector Classifier
svc_params = {'C': [0.5, 0.7, 0.9, 1], 'kernel': ['rbf', 'poly', 'sigmoid', 'linear']}
grid_svc = GridSearchCV(SVC(), svc_params)
grid_svc.fit(X_train, y_train)
# SVC best estimator
svc = grid_svc.best_estimator_
# DecisionTree Classifier
tree_params = {"criterion": ["gini", "entropy"], "max_depth": list(range(2,4,1)),
"min_samples_leaf": list(range(5,7,1))}
grid_tree = GridSearchCV(DecisionTreeClassifier(), tree_params)
grid_tree.fit(X_train, y_train)
# tree best estimator
tree_clf = grid_tree.best_estimator_
# Overfitting Case
log_reg_score = cross_val_score(log_reg, X_train, y_train, cv=5)
print('Logistic Regression Cross Validation Score: ', round(log_reg_score.mean() * 100, 2).astype(str) + '%')
knears_score = cross_val_score(knears_neighbors, X_train, y_train, cv=5)
print('Knears Neighbors Cross Validation Score', round(knears_score.mean() * 100, 2).astype(str) + '%')
svc_score = cross_val_score(svc, X_train, y_train, cv=5)
print('Support Vector Classifier Cross Validation Score', round(svc_score.mean() * 100, 2).astype(str) + '%')
tree_score = cross_val_score(tree_clf, X_train, y_train, cv=5)
print('DecisionTree Classifier Cross Validation Score', round(tree_score.mean() * 100, 2).astype(str) + '%')
# We will undersample during cross validating
undersample_X = df.drop('Class', axis=1)
undersample_y = df['Class']
for train_index, test_index in sss.split(undersample_X, undersample_y):
print("Train:", train_index, "Test:", test_index)
undersample_Xtrain, undersample_Xtest = undersample_X.iloc[train_index], undersample_X.iloc[test_index]
undersample_ytrain, undersample_ytest = undersample_y.iloc[train_index], undersample_y.iloc[test_index]
undersample_Xtrain = undersample_Xtrain.values
undersample_Xtest = undersample_Xtest.values
undersample_ytrain = undersample_ytrain.values
undersample_ytest = undersample_ytest.values
undersample_accuracy = []
undersample_precision = []
undersample_recall = []
undersample_f1 = []
undersample_auc = []
# Implementing NearMiss Technique
# Distribution of NearMiss (Just to see how it distributes the labels we won't use these variables)
X_nearmiss, y_nearmiss = NearMiss().fit_sample(undersample_X.values, undersample_y.values)
print('NearMiss Label Distribution: {}'.format(Counter(y_nearmiss)))
# Cross Validating the right way
for train, test in sss.split(undersample_Xtrain, undersample_ytrain):
undersample_pipeline = imbalanced_make_pipeline(NearMiss(sampling_strategy='majority'), log_reg) # SMOTE happens during Cross Validation not before..
undersample_model = undersample_pipeline.fit(undersample_Xtrain[train], undersample_ytrain[train])
undersample_prediction = undersample_model.predict(undersample_Xtrain[test])
undersample_accuracy.append(undersample_pipeline.score(original_Xtrain[test], original_ytrain[test]))
undersample_precision.append(precision_score(original_ytrain[test], undersample_prediction))
undersample_recall.append(recall_score(original_ytrain[test], undersample_prediction))
undersample_f1.append(f1_score(original_ytrain[test], undersample_prediction))
undersample_auc.append(roc_auc_score(original_ytrain[test], undersample_prediction))
# Let's Plot LogisticRegression Learning Curve
from sklearn.model_selection import ShuffleSplit
from sklearn.model_selection import learning_curve
def plot_learning_curve(estimator1, estimator2, estimator3, estimator4, X, y, ylim=None, cv=None,
n_jobs=1, train_sizes=np.linspace(.1, 1.0, 5)):
f, ((ax1, ax2), (ax3, ax4)) = plt.subplots(2,2, figsize=(20,14), sharey=True)
if ylim is not None:
plt.ylim(*ylim)
# First Estimator
train_sizes, train_scores, test_scores = learning_curve(
estimator1, X, y, cv=cv, n_jobs=n_jobs, train_sizes=train_sizes)
train_scores_mean = np.mean(train_scores, axis=1)
train_scores_std = np.std(train_scores, axis=1)
test_scores_mean = np.mean(test_scores, axis=1)
test_scores_std = np.std(test_scores, axis=1)
ax1.fill_between(train_sizes, train_scores_mean - train_scores_std,
train_scores_mean + train_scores_std, alpha=0.1,
color="#ff9124")
ax1.fill_between(train_sizes, test_scores_mean - test_scores_std,
test_scores_mean + test_scores_std, alpha=0.1, color="#2492ff")
ax1.plot(train_sizes, train_scores_mean, 'o-', color="#ff9124",
label="Training score")
ax1.plot(train_sizes, test_scores_mean, 'o-', color="#2492ff",
label="Cross-validation score")
ax1.set_title("Logistic Regression Learning Curve", fontsize=14)
ax1.set_xlabel('Training size (m)')
ax1.set_ylabel('Score')
ax1.grid(True)
ax1.legend(loc="best")
# Second Estimator
train_sizes, train_scores, test_scores = learning_curve(
estimator2, X, y, cv=cv, n_jobs=n_jobs, train_sizes=train_sizes)
train_scores_mean = np.mean(train_scores, axis=1)
train_scores_std = np.std(train_scores, axis=1)
test_scores_mean = np.mean(test_scores, axis=1)
test_scores_std = np.std(test_scores, axis=1)
ax2.fill_between(train_sizes, train_scores_mean - train_scores_std,
train_scores_mean + train_scores_std, alpha=0.1,
color="#ff9124")
ax2.fill_between(train_sizes, test_scores_mean - test_scores_std,
test_scores_mean + test_scores_std, alpha=0.1, color="#2492ff")
ax2.plot(train_sizes, train_scores_mean, 'o-', color="#ff9124",
label="Training score")
ax2.plot(train_sizes, test_scores_mean, 'o-', color="#2492ff",
label="Cross-validation score")
ax2.set_title("Knears Neighbors Learning Curve", fontsize=14)
ax2.set_xlabel('Training size (m)')
ax2.set_ylabel('Score')
ax2.grid(True)
ax2.legend(loc="best")
# Third Estimator
train_sizes, train_scores, test_scores = learning_curve(
estimator3, X, y, cv=cv, n_jobs=n_jobs, train_sizes=train_sizes)
train_scores_mean = np.mean(train_scores, axis=1)
train_scores_std = np.std(train_scores, axis=1)
test_scores_mean = np.mean(test_scores, axis=1)
test_scores_std = np.std(test_scores, axis=1)
ax3.fill_between(train_sizes, train_scores_mean - train_scores_std,
train_scores_mean + train_scores_std, alpha=0.1,
color="#ff9124")
ax3.fill_between(train_sizes, test_scores_mean - test_scores_std,
test_scores_mean + test_scores_std, alpha=0.1, color="#2492ff")
ax3.plot(train_sizes, train_scores_mean, 'o-', color="#ff9124",
label="Training score")
ax3.plot(train_sizes, test_scores_mean, 'o-', color="#2492ff",
label="Cross-validation score")
ax3.set_title("Support Vector Classifier \n Learning Curve", fontsize=14)
ax3.set_xlabel('Training size (m)')
ax3.set_ylabel('Score')
ax3.grid(True)
ax3.legend(loc="best")
# Fourth Estimator
train_sizes, train_scores, test_scores = learning_curve(
estimator4, X, y, cv=cv, n_jobs=n_jobs, train_sizes=train_sizes)
train_scores_mean = np.mean(train_scores, axis=1)
train_scores_std = np.std(train_scores, axis=1)
test_scores_mean = np.mean(test_scores, axis=1)
test_scores_std = np.std(test_scores, axis=1)
ax4.fill_between(train_sizes, train_scores_mean - train_scores_std,
train_scores_mean + train_scores_std, alpha=0.1,
color="#ff9124")
ax4.fill_between(train_sizes, test_scores_mean - test_scores_std,
test_scores_mean + test_scores_std, alpha=0.1, color="#2492ff")
ax4.plot(train_sizes, train_scores_mean, 'o-', color="#ff9124",
label="Training score")
ax4.plot(train_sizes, test_scores_mean, 'o-', color="#2492ff",
label="Cross-validation score")
ax4.set_title("Decision Tree Classifier \n Learning Curve", fontsize=14)
ax4.set_xlabel('Training size (m)')
ax4.set_ylabel('Score')
ax4.grid(True)
ax4.legend(loc="best")
return plt
cv = ShuffleSplit(n_splits=100, test_size=0.2, random_state=42)
plot_learning_curve(log_reg, knears_neighbors, svc, tree_clf, X_train, y_train, (0.87, 1.01), cv=cv, n_jobs=4)
from sklearn.metrics import roc_curve
from sklearn.model_selection import cross_val_predict
# Create a DataFrame with all the scores and the classifiers names.
log_reg_pred = cross_val_predict(log_reg, X_train, y_train, cv=5,
method="decision_function")
knears_pred = cross_val_predict(knears_neighbors, X_train, y_train, cv=5)
svc_pred = cross_val_predict(svc, X_train, y_train, cv=5,
method="decision_function")
tree_pred = cross_val_predict(tree_clf, X_train, y_train, cv=5)
from sklearn.metrics import roc_auc_score
print('Logistic Regression: ', roc_auc_score(y_train, log_reg_pred))
print('KNears Neighbors: ', roc_auc_score(y_train, knears_pred))
print('Support Vector Classifier: ', roc_auc_score(y_train, svc_pred))
print('Decision Tree Classifier: ', roc_auc_score(y_train, tree_pred))
log_fpr, log_tpr, log_thresold = roc_curve(y_train, log_reg_pred)
knear_fpr, knear_tpr, knear_threshold = roc_curve(y_train, knears_pred)
svc_fpr, svc_tpr, svc_threshold = roc_curve(y_train, svc_pred)
tree_fpr, tree_tpr, tree_threshold = roc_curve(y_train, tree_pred)
def graph_roc_curve_multiple(log_fpr, log_tpr, knear_fpr, knear_tpr, svc_fpr, svc_tpr, tree_fpr, tree_tpr):
plt.figure(figsize=(16,8))
plt.title('ROC Curve \n Top 4 Classifiers', fontsize=18)
plt.plot(log_fpr, log_tpr, label='Logistic Regression Classifier Score: {:.4f}'.format(roc_auc_score(y_train, log_reg_pred)))
plt.plot(knear_fpr, knear_tpr, label='KNears Neighbors Classifier Score: {:.4f}'.format(roc_auc_score(y_train, knears_pred)))
plt.plot(svc_fpr, svc_tpr, label='Support Vector Classifier Score: {:.4f}'.format(roc_auc_score(y_train, svc_pred)))
plt.plot(tree_fpr, tree_tpr, label='Decision Tree Classifier Score: {:.4f}'.format(roc_auc_score(y_train, tree_pred)))
plt.plot([0, 1], [0, 1], 'k--')
plt.axis([-0.01, 1, 0, 1])
plt.xlabel('False Positive Rate', fontsize=16)
plt.ylabel('True Positive Rate', fontsize=16)
plt.annotate('Minimum ROC Score of 50% \n (This is the minimum score to get)', xy=(0.5, 0.5), xytext=(0.6, 0.3),
arrowprops=dict(facecolor='#6E726D', shrink=0.05),
)
plt.legend()
graph_roc_curve_multiple(log_fpr, log_tpr, knear_fpr, knear_tpr, svc_fpr, svc_tpr, tree_fpr, tree_tpr)
plt.show()
```
# A Deeper Look into LogisticRegression:
In this section we will dive a deeper look into the logistic regression classifier.
## Terms:
* True Positives: Correctly Classified Fraud Transactions
* False Positives: Incorrectly Classified Fraud Transactions
* True Negative: Correctly Classified Non-Fraud Transactions
* False Negative: Incorrectly Classified Non-Fraud Transactions
* Precision: True Positives/(True Positives + False Positives)
* Recall: True Positives/(True Positives + False Negatives)
* Precision as the name says, says how precise (how sure) is our model in detecting fraud transactions while recall is the amount of fraud cases our model is able to detect.
* Precision/Recall Tradeoff: The more precise (selective) our model is, the less cases it will detect. Example: Assuming that our model has a precision of 95%, Let's say there are only 5 fraud cases in which the model is 95% precise or more that these are fraud cases. Then let's say there are 5 more cases that our model considers 90% to be a fraud case, if we lower the precision there are more cases that our model will be able to detect.
## Summary:
Precision starts to descend between 0.90 and 0.92 nevertheless, our precision score is still pretty high and still we have a descent recall score.
```
def logistic_roc_curve(log_fpr, log_tpr):
plt.figure(figsize=(12,8))
plt.title('Logistic Regression ROC Curve', fontsize=16)
plt.plot(log_fpr, log_tpr, 'b-', linewidth=2)
plt.plot([0, 1], [0, 1], 'r--')
plt.xlabel('False Positive Rate', fontsize=16)
plt.ylabel('True Positive Rate', fontsize=16)
plt.axis([-0.01,1,0,1])
logistic_roc_curve(log_fpr, log_tpr)
plt.show()
from sklearn.metrics import precision_recall_curve
precision, recall, threshold = precision_recall_curve(y_train, log_reg_pred)
from sklearn.metrics import recall_score, precision_score, f1_score, accuracy_score
y_pred = log_reg.predict(X_train)
# Overfitting Case
print('---' * 45)
print('Overfitting: \n')
print('Recall Score: {:.2f}'.format(recall_score(y_train, y_pred)))
print('Precision Score: {:.2f}'.format(precision_score(y_train, y_pred)))
print('F1 Score: {:.2f}'.format(f1_score(y_train, y_pred)))
print('Accuracy Score: {:.2f}'.format(accuracy_score(y_train, y_pred)))
print('---' * 45)
# How it should look like
print('---' * 45)
print('How it should be:\n')
print("Accuracy Score: {:.2f}".format(np.mean(undersample_accuracy)))
print("Precision Score: {:.2f}".format(np.mean(undersample_precision)))
print("Recall Score: {:.2f}".format(np.mean(undersample_recall)))
print("F1 Score: {:.2f}".format(np.mean(undersample_f1)))
print('---' * 45)
undersample_y_score = log_reg.decision_function(original_Xtest)
from sklearn.metrics import average_precision_score
undersample_average_precision = average_precision_score(original_ytest, undersample_y_score)
print('Average precision-recall score: {0:0.2f}'.format(
undersample_average_precision))
from sklearn.metrics import precision_recall_curve
import matplotlib.pyplot as plt
fig = plt.figure(figsize=(12,6))
precision, recall, _ = precision_recall_curve(original_ytest, undersample_y_score)
plt.step(recall, precision, color='#004a93', alpha=0.2,
where='post')
plt.fill_between(recall, precision, step='post', alpha=0.2,
color='#48a6ff')
plt.xlabel('Recall')
plt.ylabel('Precision')
plt.ylim([0.0, 1.05])
plt.xlim([0.0, 1.0])
plt.title('UnderSampling Precision-Recall curve: \n Average Precision-Recall Score ={0:0.2f}'.format(
undersample_average_precision), fontsize=16)
```
# SMOTE Technique (Over-Sampling):
<img src="https://raw.githubusercontent.com/rikunert/SMOTE_visualisation/master/SMOTE_R_visualisation_3.png", width=800> SMOTE stands for Synthetic Minority Over-sampling Technique. Unlike Random UnderSampling, SMOTE creates new synthetic points in order to have an equal balance of the classes. This is another alternative for solving the "class imbalance problems".
## Understanding SMOTE:
Solving the Class Imbalance: SMOTE creates synthetic points from the minority class in order to reach an equal balance between the minority and majority class.
Location of the synthetic points: SMOTE picks the distance between the closest neighbors of the minority class, in between these distances it creates synthetic points.
Final Effect: More information is retained since we didn't have to delete any rows unlike in random undersampling.
Accuracy || Time Tradeoff: Although it is likely that SMOTE will be more accurate than random under-sampling, it will take more time to train since no rows are eliminated as previously stated.
## Overfitting during Cross Validation:
In our undersample analysis I want to show you a common mistake I made that I want to share with all of you. It is simple, if you want to undersample or oversample your data you should not do it before cross validating. Why because you will be directly influencing the validation set before implementing cross-validation causing a "data leakage" problem. In the following section you will see amazing precision and recall scores but in reality our data is overfitting!
***Wrong Way***
<img src="asset/1.jpg" />
As mentioned previously, if we get the minority class ("Fraud) in our case, and create the synthetic points before cross validating we have a certain influence on the "validation set" of the cross validation process. Remember how cross validation works, let's assume we are splitting the data into 5 batches, 4/5 of the dataset will be the training set while 1/5 will be the validation set. The test set should not be touched! For that reason, we have to do the creation of synthetic datapoints "during" cross-validation and not before, just like below:
***Right Way***
<img src="asset/1.jpg" />
As you see above, SMOTE occurs "during" cross validation and not "prior" to the cross validation process. Synthetic data are created only for the training set without affecting the validation set.
```
from imblearn.over_sampling import SMOTE
from sklearn.model_selection import train_test_split, RandomizedSearchCV
print('Length of X (train): {} | Length of y (train): {}'.format(len(original_Xtrain), len(original_ytrain)))
print('Length of X (test): {} | Length of y (test): {}'.format(len(original_Xtest), len(original_ytest)))
# List to append the score and then find the average
accuracy_lst = []
precision_lst = []
recall_lst = []
f1_lst = []
auc_lst = []
# Classifier with optimal parameters
# log_reg_sm = grid_log_reg.best_estimator_
log_reg_sm = LogisticRegression()
rand_log_reg = RandomizedSearchCV(LogisticRegression(), log_reg_params, n_iter=4)
# Implementing SMOTE Technique
# Cross Validating the right way
# Parameters
log_reg_params = {"penalty": ['l1', 'l2'], 'C': [0.001, 0.01, 0.1, 1, 10, 100, 1000]}
for train, test in sss.split(original_Xtrain, original_ytrain):
pipeline = imbalanced_make_pipeline(SMOTE(sampling_strategy='minority'), rand_log_reg) # SMOTE happens during Cross Validation not before..
model = pipeline.fit(original_Xtrain[train], original_ytrain[train])
best_est = rand_log_reg.best_estimator_
prediction = best_est.predict(original_Xtrain[test])
accuracy_lst.append(pipeline.score(original_Xtrain[test], original_ytrain[test]))
precision_lst.append(precision_score(original_ytrain[test], prediction))
recall_lst.append(recall_score(original_ytrain[test], prediction))
f1_lst.append(f1_score(original_ytrain[test], prediction))
auc_lst.append(roc_auc_score(original_ytrain[test], prediction))
print('---' * 45)
print('')
print("accuracy: {}".format(np.mean(accuracy_lst)))
print("precision: {}".format(np.mean(precision_lst)))
print("recall: {}".format(np.mean(recall_lst)))
print("f1: {}".format(np.mean(f1_lst)))
print('---' * 45)
labels = ['No Fraud', 'Fraud']
smote_prediction = best_est.predict(original_Xtest)
print(classification_report(original_ytest, smote_prediction, target_names=labels))
y_score = best_est.decision_function(original_Xtest)
average_precision = average_precision_score(original_ytest, y_score)
print('Average precision-recall score: {0:0.2f}'.format(
average_precision))
fig = plt.figure(figsize=(12,6))
precision, recall, _ = precision_recall_curve(original_ytest, y_score)
plt.step(recall, precision, color='r', alpha=0.2,
where='post')
plt.fill_between(recall, precision, step='post', alpha=0.2,
color='#F59B00')
plt.xlabel('Recall')
plt.ylabel('Precision')
plt.ylim([0.0, 1.05])
plt.xlim([0.0, 1.0])
plt.title('OverSampling Precision-Recall curve: \n Average Precision-Recall Score ={0:0.2f}'.format(
average_precision), fontsize=16)
# SMOTE Technique (OverSampling) After splitting and Cross Validating
sm = SMOTE(ratio='minority', random_state=42)
# Xsm_train, ysm_train = sm.fit_sample(X_train, y_train)
# This will be the data were we are going to
Xsm_train, ysm_train = sm.fit_sample(original_Xtrain, original_ytrain)
# We Improve the score by 2% points approximately
# Implement GridSearchCV and the other models.
# Logistic Regression
t0 = time.time()
log_reg_sm = grid_log_reg.best_estimator_
log_reg_sm.fit(Xsm_train, ysm_train)
t1 = time.time()
print("Fitting oversample data took :{} sec".format(t1 - t0))
```
# Test Data with Logistic Regression:
## Confusion Matrix:
* Positive/Negative: Type of Class (label) ["No", "Yes"] True/False: Correctly or Incorrectly classified by the model.
* True Negatives (Top-Left Square): This is the number of correctly classifications of the "No" (No Fraud Detected) class or potenial clients that are not willing to suscribe a term deposit.
* False Negatives (Top-Right Square): This is the number of incorrectly classifications of the "No"(No Fraud Detected) class or potential clients that are not willing to suscribe a term depositt.
* False Positives (Bottom-Left Square): This is the number of incorrectly classifications of the "Yes" (Fraud Detected) class or potential clients that are willing to suscribe a term deposit.
* True Positives (Bottom-Right Square): This is the number of correctly classifications of the "Yes" (Fraud Detected) class or potenial clients that are willing to suscribe a term deposit.
## Summary:
* Random UnderSampling: We will evaluate the final performance of the classification models in the random undersampling subset. Keep in mind that this is not the data from the original dataframe.
* Classification Models: The models that performed the best were logistic regression and support vector classifier (SVM)
```
from sklearn.metrics import confusion_matrix
# Logistic Regression fitted using SMOTE technique
y_pred_log_reg = log_reg_sm.predict(X_test)
# Other models fitted with UnderSampling
y_pred_knear = knears_neighbors.predict(X_test)
y_pred_svc = svc.predict(X_test)
y_pred_tree = tree_clf.predict(X_test)
log_reg_cf = confusion_matrix(y_test, y_pred_log_reg)
kneighbors_cf = confusion_matrix(y_test, y_pred_knear)
svc_cf = confusion_matrix(y_test, y_pred_svc)
tree_cf = confusion_matrix(y_test, y_pred_tree)
fig, ax = plt.subplots(2, 2,figsize=(22,12))
sns.heatmap(log_reg_cf, ax=ax[0][0], annot=True, cmap=plt.cm.copper)
ax[0, 0].set_title("Logistic Regression \n Confusion Matrix", fontsize=14)
ax[0, 0].set_xticklabels(['', ''], fontsize=14, rotation=90)
ax[0, 0].set_yticklabels(['', ''], fontsize=14, rotation=360)
sns.heatmap(kneighbors_cf, ax=ax[0][1], annot=True, cmap=plt.cm.copper)
ax[0][1].set_title("KNearsNeighbors \n Confusion Matrix", fontsize=14)
ax[0][1].set_xticklabels(['', ''], fontsize=14, rotation=90)
ax[0][1].set_yticklabels(['', ''], fontsize=14, rotation=360)
sns.heatmap(svc_cf, ax=ax[1][0], annot=True, cmap=plt.cm.copper)
ax[1][0].set_title("Suppor Vector Classifier \n Confusion Matrix", fontsize=14)
ax[1][0].set_xticklabels(['', ''], fontsize=14, rotation=90)
ax[1][0].set_yticklabels(['', ''], fontsize=14, rotation=360)
sns.heatmap(tree_cf, ax=ax[1][1], annot=True, cmap=plt.cm.copper)
ax[1][1].set_title("DecisionTree Classifier \n Confusion Matrix", fontsize=14)
ax[1][1].set_xticklabels(['', ''], fontsize=14, rotation=90)
ax[1][1].set_yticklabels(['', ''], fontsize=14, rotation=360)
plt.show()
from sklearn.metrics import classification_report
print('Logistic Regression:')
print(classification_report(y_test, y_pred_log_reg))
print('KNears Neighbors:')
print(classification_report(y_test, y_pred_knear))
print('Support Vector Classifier:')
print(classification_report(y_test, y_pred_svc))
print('Support Vector Classifier:')
print(classification_report(y_test, y_pred_tree))
# Final Score in the test set of logistic regression
from sklearn.metrics import accuracy_score
# Logistic Regression with Under-Sampling
y_pred = log_reg.predict(X_test)
undersample_score = accuracy_score(y_test, y_pred)
# Logistic Regression with SMOTE Technique (Better accuracy with SMOTE t)
y_pred_sm = best_est.predict(original_Xtest)
oversample_score = accuracy_score(original_ytest, y_pred_sm)
d = {'Technique': ['Random UnderSampling', 'Oversampling (SMOTE)'], 'Score': [undersample_score, oversample_score]}
final_df = pd.DataFrame(data=d)
# Move column
score = final_df['Score']
final_df.drop('Score', axis=1, inplace=True)
final_df.insert(1, 'Score', score)
# Note how high is accuracy score it can be misleading!
final_df
```
# Neural Networks Testing Random UnderSampling Data vs OverSampling (SMOTE):
In this section we will implement a simple Neural Network (with one hidden layer) in order to see which of the two logistic regressions models we implemented in the (undersample or oversample(SMOTE)) has a better accuracy for detecting fraud and non-fraud transactions.
## Our Main Goal:
Our main goal is to explore how our simple neural network behaves in both the random undersample and oversample dataframes and see whether they can predict accuractely both non-fraud and fraud cases. Why not only focus on fraud? Imagine you were a cardholder and after you purchased an item your card gets blocked because the bank's algorithm thought your purchase was a fraud. That's why we shouldn't emphasize only in detecting fraud cases but we should also emphasize correctly categorizing non-fraud transactions.
## The Confusion Matrix:
Here is again, how the confusion matrix works:
* Upper Left Square: The amount of correctly classified by our model of no fraud transactions.
* Upper Right Square: The amount of incorrectly classified transactions as fraud cases, but the actual label is no fraud .
* Lower Left Square: The amount of incorrectly classified transactions as no fraud cases, but the actual label is fraud .
* Lower Right Square: The amount of correctly classified by our model of fraud transactions.
## Summary (Keras || Random UnderSampling):
* Dataset: In this final phase of testing we will fit this model in both the random undersampled subset and oversampled dataset (SMOTE) in order to predict the final result using the original dataframe testing data.
* Neural Network Structure: As stated previously, this will be a simple model composed of one input layer (where the number of nodes equals the number of features) plus bias node, one hidden layer with 32 nodes and one output node composed of two possible results 0 or 1 (No fraud or fraud).
* Other characteristics: The learning rate will be 0.001, the optimizer we will use is the AdamOptimizer, the activation function that is used in this scenario is "Relu" and for the final outputs we will use sparse categorical cross entropy, which gives the probability whether an instance case is no fraud or fraud (The prediction will pick the highest probability between the two.)
```
import keras
from keras import backend as K
from keras.models import Sequential
from keras.layers import Activation
from keras.layers.core import Dense
from keras.optimizers import Adam
from keras.metrics import categorical_crossentropy
n_inputs = X_train.shape[1]
undersample_model = Sequential([
Dense(n_inputs, input_shape=(n_inputs, ), activation='relu'),
Dense(32, activation='relu'),
Dense(2, activation='softmax')
])
undersample_model.summary()
undersample_model.compile(Adam(lr=0.001), loss='sparse_categorical_crossentropy', metrics=['accuracy'])
undersample_model.fit(X_train, y_train, validation_split=0.2, batch_size=25, epochs=20, shuffle=True, verbose=2)
undersample_predictions = undersample_model.predict(original_Xtest, batch_size=200, verbose=0)
undersample_fraud_predictions = undersample_model.predict_classes(original_Xtest, batch_size=200, verbose=0)
import itertools
# Create a confusion matrix
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
print(cm)
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title, fontsize=14)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
undersample_cm = confusion_matrix(original_ytest, undersample_fraud_predictions)
actual_cm = confusion_matrix(original_ytest, original_ytest)
labels = ['No Fraud', 'Fraud']
fig = plt.figure(figsize=(16,8))
fig.add_subplot(221)
plot_confusion_matrix(undersample_cm, labels, title="Random UnderSample \n Confusion Matrix", cmap=plt.cm.Reds)
fig.add_subplot(222)
plot_confusion_matrix(actual_cm, labels, title="Confusion Matrix \n (with 100% accuracy)", cmap=plt.cm.Greens)
```
# Keras || OverSampling (SMOTE):
```
n_inputs = Xsm_train.shape[1]
oversample_model = Sequential([
Dense(n_inputs, input_shape=(n_inputs, ), activation='relu'),
Dense(32, activation='relu'),
Dense(2, activation='softmax')
])
oversample_model.compile(Adam(lr=0.001), loss='sparse_categorical_crossentropy', metrics=['accuracy'])
oversample_model.fit(Xsm_train, ysm_train, validation_split=0.2, batch_size=300, epochs=20, shuffle=True, verbose=2)
oversample_predictions = oversample_model.predict(original_Xtest, batch_size=200, verbose=0)
oversample_fraud_predictions = oversample_model.predict_classes(original_Xtest, batch_size=200, verbose=0)
oversample_smote = confusion_matrix(original_ytest, oversample_fraud_predictions)
actual_cm = confusion_matrix(original_ytest, original_ytest)
labels = ['No Fraud', 'Fraud']
fig = plt.figure(figsize=(16,8))
fig.add_subplot(221)
plot_confusion_matrix(oversample_smote, labels, title="OverSample (SMOTE) \n Confusion Matrix", cmap=plt.cm.Oranges)
fig.add_subplot(222)
plot_confusion_matrix(actual_cm, labels, title="Confusion Matrix \n (with 100% accuracy)", cmap=plt.cm.Greens)
```
# Conclusion:
Implementing SMOTE on our imbalanced dataset helped us with the imbalance of our labels (more no fraud than fraud transactions). Nevertheless, I still have to state that sometimes the neural network on the oversampled dataset predicts less correct fraud transactions than our model using the undersample dataset. However, remember that the removal of outliers was implemented only on the random undersample dataset and not on the oversampled one. Also, in our undersample data our model is unable to detect for a large number of cases non fraud transactions correctly and instead, misclassifies those non fraud transactions as fraud cases. Imagine that people that were making regular purchases got their card blocked due to the reason that our model classified that transaction as a fraud transaction, this will be a huge disadvantage for the financial institution. The number of customer complaints and customer disatisfaction will increase. The next step of this analysis will be to do an outlier removal on our oversample dataset and see if our accuracy in the test set improves.
| github_jupyter |
This short example show how to get data from FMI Open Data multipointcoverage format. The format is used in INSPIRE specifications and is somewhat complex. Anyway, it's the most efficient way to get large amounts of data.
Here we fetch all observations from Finland during two days.
This example is for "old" format WFS2. You may try to use new WFS3 beta service as well. It's available in: http://beta.fmi.fi/data/3/wfs/sofp/
```
import requests
import datetime as dt
import xml.etree.ElementTree as ET
import numpy as np
import re
```
Required functions to get param names. Param keys are in the response document but longer names along with other metadata need to be fetched separately.
```
def get_param_names(url):
""" Get parameters metadata"""
req = requests.get(url)
params = {}
if req.status_code == 200:
xmlstring = req.content
tree = ET.ElementTree(ET.fromstring(xmlstring))
for p in tree.iter(tag='{http://inspire.ec.europa.eu/schemas/omop/2.9}ObservableProperty'):
params[p.get('{http://www.opengis.net/gml/3.2}id')] = p.find('{http://inspire.ec.europa.eu/schemas/omop/2.9}label').text
return params
def get_params(tree):
""" Get parameters from response xml tree """
retParams = []
for el in tree.iter(tag='{http://www.opengis.net/om/2.0}observedProperty'):
url = el.get('{http://www.w3.org/1999/xlink}href')
params = re.findall(r"(?<=param=).*,.*(?=&)", url)[0].split(',')
param_names = get_param_names(url)
for p in params:
retParams.append('{} ({})'.format(param_names[p], p))
return retParams
```
Positions are in the separate element. Positions are listed as lat, lon, timestamp.
```
def get_positions(tree):
"""
Function to get times and coordinates from multipointcoverage answer
"""
positions = []
for el in tree.iter(tag='{http://www.opengis.net/gmlcov/1.0}positions'):
pos = el.text.split()
i = 0
while len(pos) > 0:
lat = float(pos.pop(0))
lon = float(pos.pop(0))
timestamp = int(pos.pop(0))
positions.append([lat,lon,timestamp])
return np.array(positions)
```
Get data. For longer periods we have to fetch data in the loop
```
url = 'http://opendata.fmi.fi/wfs'
starttime = dt.datetime.strptime('2010-01-01', "%Y-%m-%d")
endtime = dt.datetime.strptime('2010-01-03', "%Y-%m-%d")
daystep = 1
start = starttime
end = start + dt.timedelta(days=daystep)
if end > endtime: end = endtime
while end <= endtime and start < end:
startStr = start.strftime('%Y-%m-%d')
endStr = end.strftime('%Y-%m-%d')
# Get data
payload = {
'request': 'getFeature',
'storedquery_id': 'fmi::observations::weather::multipointcoverage',
'bbox': '19,59,35,75',
'starttime': startStr,
'endtime': endStr,
}
r = requests.get(url, params=payload)
# Construct XML tree
tree = ET.ElementTree(ET.fromstring(r.content))
# Get geospatial and temporal positions of data elements
positions = get_positions(tree)
# Extract data from XML tree
d = []
for el in tree.iter(tag='{http://www.opengis.net/gml/3.2}doubleOrNilReasonTupleList'):
for pos in el.text.strip().split("\n"):
d.append(pos.strip().split(' '))
# Assign data values to positions
junk = np.append(positions, np.array(d), axis=1)
try:
data = np.append(data, junk, axis=0)
except NameError:
data = junk
print('Time interval {} - {} provided {} rows'.format(startStr, endStr, junk.shape[0]))
start = end
end = start + dt.timedelta(days=daystep)
if end > endtime: end = endtime
print('Done fetching data. Final dimensions of the result: {}'.format(data.shape))
```
Get params from the last XML tree element (they don't change over time)
```
params = get_params(tree)
```
Finally you can do whatever you want with the data. Here we just print some example.
```
print('Params: {}'.format(params))
print(data[0:2])
```
| github_jupyter |
# HPDM097: Foundations of combinatorial optimisation for routing and scheduling problems in health
Many healthcare systems manage assets or workforce that they need to deploy geographically. One example, is a community nursing team. These are teams of highly skilled nurses that must visit patients in their own home. Another example, is patient transport services where a fleet of non-emergency ambulances pick up patients from their own home and transport them to outpatient appointments in a clinical setting. These problems are highly complex. For example, in the community nursing example, patients will have a variety of conditions, treatments may be time dependent (for example, insulin injections), nurses will have mixed skills and staffing will vary over time.
---
# The Travelling Nurse Problem
For simplicity you will first consider a single asset that has to visit patients in their own home and ignore the complex constraints described above. We will frame this problem as the famous **Travelling Salesperson (or Nurse!) Problem (TSP).**
**By the end of this section you will have learnt how to:**
* represent a routing and scheduling problem in a form suitable for solution by an optimisation algorithm
* solve small instances of the Travelling Salesman Problem (TSP) using a brute force approach
* solve and obtain good solutions to larger TSP problem by applying hill climbing algorithms in combination with stochastic algorithms
* understand and apply a more intelligent hill climbing approach called Iterated Local Search
> Please use the conda environment `hds_logistics` when running this workbook. You will also need to run this workbook in the same directory as `metapy`. This is a small python package that contains the code to solve the TSP.
# Imports
```
import numpy as np
import matplotlib.pyplot as plt
import time
```
# `metapy` package imports
```
import metapy.tsp.tsp_io as io
import metapy.tsp.euclidean as e
from metapy.tsp.init_solutions import TSPPopulationGenerator
from metapy.tsp.objective import SimpleTSPObjective, OptimisedSimpleTSPObjective
from metapy.tsp.bruteforce import BruteForceSolver, RandomSearch
from metapy.local_search.ils import (IteratedLocalSearch,
HigherQualityHomeBase,
RandomHomeBase,
EpsilonGreedyHomeBase,
AnnealingEpsilonGreedyHomeBase,
TempFastCoolingSchedule,
DoubleBridgePertubation,
TabuDoubleBridgeTweak)
from metapy.local_search.hill_climbing import (HillClimber,
TweakTwoOpt,
SimpleTweak,
HillClimberRandomRestarts)
from metapy.evolutionary.evolutionary import (EvolutionaryAlgorithm,
MuLambdaEvolutionStrategy,
MuPlusLambdaEvolutionStrategy,
GeneticAlgorithmStrategy,
ElitistGeneticAlgorithmStrategy,
TwoOptMutator, TwoCityMutator,
TruncationSelector,
TournamentSelector,
PartiallyMappedCrossover)
```
# Load the data.
In this notebook, you will work with the famous **st70** problem from [TSPLib](http://elib.zib.de/pub/mp-testdata/tsp/tsplib/tsplib.html).
> You will move onto a real health service dataset in **part 2** where you will work with the a more complex variant of this problem for routing and scheduling with multiple health service assets.
The data is located in `data/st70.tsp`. The data format from TSPLib contains both metadata and 2D coordinates of 'cities'. The files therefore need some minor preprocessing before they are usable.
> For efficiency you will work mainly with `numpy`. It of course possible to use `pandas` for this type of problem, but you will pay a heavy price in terms of execution time!
```
#load file
file_path = "data/st70.tsp"
#number of rows in the file that are meta_data
md_rows = 6
#read the coordinates
cities = io.read_coordinates(file_path, md_rows)
#read the meta data
meta = io.read_meta_data(file_path, md_rows)
#should be an numpy.ndarray
print(type(cities))
# should be 70 cities
print(cities.shape)
#print first 2 coordinate pairs
print(cities[:2])
print("st70 meta data")
print(meta)
```
The meta data confirms that problem is Euclidean 2D. This means that we need to calculate the euclidean distance between points.
```
#example of calculating a single euclidean distance
e.euclidean_distance(cities[0], cities[1])
from decimal import Decimal, ROUND_HALF_UP
def gen_matrix(cities, as_integer=False):
"""
Creates a numpy array of euclidian distances between 2 sets of
cities
Parameters:
----------
points: numpy.array
coordinate pairs
as_integers: bool, optional (default=False)
If true then round to nearest int.
Behaviour: 1.5 -> 2
1.2 -> 1
1.8 -> 2
Returns:
-------
np.ndarray
Matrix of city to city costs
"""
size = len(cities)
matrix = np.zeros(shape=(size, size))
row = 0
col = 0
for city1 in cities:
col = 0
for city2 in cities:
distance = e.euclidean_distance(city1, city2)
if as_integer:
distance = int(Decimal(distance).quantize(0, ROUND_HALF_UP))
matrix[row, col] = distance
col+=1
row +=1
return matrix
#generate matrix
matrix = gen_matrix(cities, as_integer=True)
file_out = 'data/st70_matrix.csv'
#output city matrix - to validate and use for manual calcs etc.
np.savetxt(file_out, matrix, delimiter=",")
matrix.shape
```
# Representation
While you develop your code it is recommended that you work with a small tour. This means that you can find the optimal solution by enumerating all solutions and check that your algorithm is working.
Representation is straightforward in TSP. It is recommended that create a `np.ndarray` as a vector of city indexes. For example in a TSP problem with 8 cities.
```
#create ordered list of cities to visit
tour = np.arange(8)
tour
```
> Remember that the TSP is a loop. You need to remember this when calculating the tour length
# Calculating the length of a tour
To calculate the length of a tour you can use either `SimpleTSPObjective` or `OptimisedSimpleTSPObjective`. For larger problems (e.g. a 70 city) problem you should find that `OptimisedSimpleTSPObjective` offers an efficiency boost (it runs quicker). But for smaller problems the overhead to set up the optimised approach means that `SimpleTSPObjective` is more efficient!
The code below illustrates how to create each type of objective and how to use them to cost a tour. If you are interested try changing the tour size (up to a max of 70) and executing the code. It will report an average runtime.
```python
#create a tour with 8 cities.
tour = np.arange(8)
```
```
#create a tour
rng = np.random.default_rng(seed=42)
tour = np.arange(8)
rng.shuffle(tour)
tour
#create an instance of an objective object and cost a tour.
objective = SimpleTSPObjective(matrix)
objective.evaluate(tour)
objective.evaluate(tour)
#create an instance of an optimised objective function
objective2 = OptimisedSimpleTSPObjective(matrix)
objective2.evaluate(tour)
```
The following code run the `evaluate` method multiple times and reports average execution speed.
This will vary by the system you are using and by the size of the problem instance.
```
%timeit objective.evaluate(tour)
%timeit objective2.evaluate(tour)
```
# Visualising a tour
A simple way to visualise a tour is to use matplotlib. The function `plot_tour` below has been provided to help you visualise a single tour.
Run the code below. It should be easy to see that this isn't a very sensible tour if your objective is to simply minimise travel distance!
```
def plot_tour(tour, cities, figsize=(6,4)):
'''
Plots a tour. Each city visited is
labelled in order. Red point is the initial city.
Params:
------
tour: np.ndarray
ordered vector representing tour e.g. [1, 4, 2, 3]
cities: np.ndarray
matrix representing city coordinates
figsize: tuple, optional (default = (6,3))
tuple of ints for figure size
Returns
-------
tuple of matplotlib figure, and axis
'''
tour_length = len(tour)
fig, ax = plt.subplots(1, 1, figsize=figsize)
#plot points
ax.plot(cities[:tour_length][:, 0],
cities[:tour_length][:, 1],'bo')
#plot lines
for j in range(len(tour)-1):
city_1 = tour[j]
city_2 = tour[j+1]
#lookup coordinates
coords_1 = cities[city_1]
coords_2 = cities[city_2]
coords = np.vstack([coords_1, coords_2])
#plot lines
ax.plot(coords[:,0], coords[:,1], 'g-')
#show order in tour
ax.text(coords_1[0] + 0.8, coords_1[1] + 0.8, str(j))
#add in loop back colour code in red...
city_1 = tour[-1]
city_2 = tour[0]
coords_1 = cities[city_1]
coords_2 = cities[city_2]
coords = np.vstack([coords_1, coords_2])
ax.text(coords_1[0] + 0.8, coords_1[1] + 0.8, str(tour_length-1))
ax.plot(coords[:,0], coords[:,1], 'r--')
return fig, ax
#example visualising a tour
rng = np.random.default_rng(seed=42)
tour = np.arange(8)
rng.shuffle(tour)
#plot the tour
fig, ax = plot_tour(tour, cities)
```
# Enumerating all solutions
You can enumerate all solutions of a **small** TSP using the `metapy.tsp.bruteforce.BruteForceSolver` class. The code below creates a `solver` passes in a initial solution (a tour) and a `objective` and then runs the solver.
The function `print_output` has been provided so that you can quickly output the results of the solver.
```
def print_output(solver):
'''
Utility function for printing formatted output of a solver
Params:
-------
solver: object
Solver class that has .best_solutions, .best_cost attributes
'''
print("\nbest solutions:\t{0}".format(len(solver.best_solutions)))
print("best cost:\t{0}".format(solver.best_cost))
print("best solutions:")
[print(s) for s in solver.best_solutions]
#create a tour - there is NO need to randomise for bruteforce
tour = np.arange(8)
#create the objective
objective = SimpleTSPObjective(matrix)
#create the brute force solver
solver = BruteForceSolver(tour, objective)
#run the solver (should be quick below tour of length 10)
print("Enumerating all solutions...")
solver.solve()
print("\n** BRUTEFORCE OUTPUT ***")
#this should find two optimal solutions! (the reverse of each other)
print_output(solver)
#now visualise the result of solution 1
fig, ax = plot_tour(solver.best_solutions[0], cities)
#now visualise the results of solution 2 (the reverse of 1)
fig, ax = plot_tour(solver.best_solutions[1], cities)
```
# Exercise 1: Solving a 9 city TSP
**Task**:
* Use a brute force approach to solve a 9 city TSP from the st70 dataset
* Plot the results
```
# your code here...
```
# A basic optimisation method: random search
Instead of a brute force enumeration we could have solved the small TSP problem using a **global optimisation algorithm**. These algorithms do not get stuck in 'local optima' and will find the optimum solution **if run for long enough**. That is a big **IF**!
The simplest method is **random search**. This makes *b* shuffles of the tour where *b* is an fixed iteration budget or the number of iterations that can be complete in a specified time limit.
> Random search is straightforward to implement yourself. It is a loop with a if statement checking for new best solutions.
You can also use `metapy.trp.bruteforce.RandomSearch` to conduct a random search on the TSP.
```python
#note max_iter is an optional parameter with default value of 1000
solver = RandomSearch(tour, objective, max_iter=1000)
```
# Exercise 2: Setting a benchmark with random search
**Task**:
* Apply random search to the 9 city problem in the st70 dataset. Use a max_iter budget of 1000.
* Compare the result to the optimal solution obtained in exercise 1.
* Set a benchmark for solving the 70 city problem - apply random search to the full 70 city problem
**Hints:**
* When using random search with the 9 city problem you may want to complete multiple runs to get a feel for its performance.
```
# your code here ...
```
# Using a hill-climbing approach
When working in logistics it is likely that you will need to employ some form of simple hill-climbing algorithm. These are very simple algorithms that iteratively test neighbouring solutions to see if they find any improvement. This **local search** approach is often very successful at finding reasonably good solutions to a routing and scheduling problem. You will see that you can easily out perform random search. However, hill climbers do suffer from getting stuck in a **local optimum** and you can often do better by employing a more sophisticated algorithm.
**However,** you might be surprised at how useful hill-climbers turn out to be when used in combination with other approaches. Here you will first experiment with a simple first improvement hill climber and then use it to **clean up** the solution produced by a evolutionary strategy and **combine** the framework into random search followed by hill climbing. One of the key benefits of hill climbers is that they are relatively fast (because they are simple). You can even set a time limit to get some of the benefit of local search without greatly extending the execution time of your algorithm.
> Although this the approach is called Hill-Climbing in the TSP you are **descending** a hill to the find the shortest route. The algorithm is the same, but you are maximising -1*objective (or alternatively $\dfrac{1}{objective}$).
# Exercise 3: Simple versus 2-Opt tweaks
Hill-Climbing works by iteratively **tweaking** a solution to search for better neighbouring solutions. `metapy` provides two relatively straightforward tweak operators. `SimpleTweak` swaps the position of two cities at a time while `TweakTwoOpt` reverses a section of the route between two cities. Generally speaking `TweakTwoOpt` will produce better solutions, but it is worth considering a `SimpleTweak` approach when **cleaning up** the output of another algorithm. You could also try both!
You create the tweak operators as follows:
```python
operator1 = SimpleTweak()
operator2 = TweakTwoOpt()
```
Each tweak operator provides a `tweak(tour, index1, index2)` method. **Note that the change to tour happens in place**
```python
tour = np.arange(10)
tweaker = SimpleTweak()
#swap cities at index 1 and index 2.
tweaker.tweak(tour, 1, 2)
```
**Task**:
* Create a numpy vector representing a tour of 10 cities
* Perform a simple tweak of cities in elements 5 and 9
* Perform a 2-opt tweak between cities 1 and 4
* Print out the updated tour.
```
#your code here...
```
# Exercise 4: Hill-Climbing
You have been provided with a simple hill climber class in `metapy`. The code below demonstrates how to create a hill-climbing object and run the algorithm.
**Task:**
* Read the code below and check your understand it.
* Run the code below and check if the hill climber is better or worse than random search.
* Modify the code below so that you pass a random initial solution to the hill climber.
**Hints**:
* a random initial solution is just a **shuffled** numpy array.
```
#Basic First Improvement Hill Climber
#create a tour (full 70 cities)
tour = np.arange(70)
###########################################
# MODIFY CODE HERE TO SHUFFLE tour
#
###########################################
#create TSP objective
objective = SimpleTSPObjective(matrix)
#create Hill climbing algorithm
solver = HillClimber(objective=objective,
init_solution=tour,
tweaker=TweakTwoOpt(),
maximisation=False)
#run the local search
solver.solve()
#output results
print("\n** Hill Climber First Improvement OUTPUT ***")
print("best cost:\t{0}".format(solver.best_cost))
print("best solutions:")
print(solver.best_solutions)
fig, ax = plot_tour(solver.best_solutions[0], cities, figsize=(12,9))
```
# Exercise 5: Using an evolutionary algorithm followed by hill climbing
You will now experiment with using a hill climber to **clean up** the solution provided by a $(\mu, \lambda)$ evolutionary strategy. It is often useful to make a few small computationally cheap tweaks to the solution provided by a more complex algorithm to gain additional performance.
The code below has been set up for you to run an evolutionary strategy against the st70 problem.
**Task:**
* Read and run the code. Does the EA beat the basic hill climber and random search? You may want to try this a few times or tune parameters.
* The final line of code assigns the EAs solution to `interim_solution`. Create a `HillClimber` and pass in `interim_solution` as its initial solution.
* Try the `SimpleTweak()` operator.
* Output the hill climbers results and plot the route.
**Hints**:
* The EA will take a few seconds to run. If you use new Jupyter cells for your hill climbing you can run each algorithm separately.
* Remember the EA is stochastic. Feel free to run it a few times to see how hill climbing can help. It may not help every time.
```
%%time
#Evolutionary Algorithm - (mu, lambda) strategy for TSP
mu = 10
_lambda = 200
#full tour
tour = np.arange(70)
###########################################################
# Create objective
# if you are finding EA a bit slow try OptimisedSimpleTSPObjective
# its experimental so be warned!
objective = SimpleTSPObjective(matrix)
#objective = OptimisedSimpleTSPObjective(matrix)
###########################################################
#create initial TSP population
init = TSPPopulationGenerator(tour)
#(Mu, Lambda) strategy using 2-Opt mutation
strategy = MuLambdaEvolutionStrategy(mu, _lambda, TwoOptMutator())
#EA
solver = EvolutionaryAlgorithm(init, objective,_lambda, strategy,
maximisation=False, generations=1000)
#run the EA
print("\nRunning (mu, lambda) evolutionary alg...")
solver.solve()
#output EA results
print("\n** (mu, LAMBDA) OUTPUT ***")
print("best cost:\t{0}".format(solver.best_fitness))
print("best solutions:")
print(solver.best_solution)
fig, ax = plot_tour(solver.best_solution, cities, figsize=(12,9))
interim_solution = solver.best_solution
#################################################
#
# Modification here: pass interim_solution to a hill climber
# or use a new Jupyter cell.
#
#################################################
```
# Exercise 6: Hill Climbing with Random Restarts
Hill-Climbing algorithms may provide a different local optima dependent on the initial solution it is provided. One option is therefore to combine Random Search and Hill Climbing into a general (and still rather dumb) algorithm called Hill-Climbing with Random Restarts. Effectively it runs Hill-Climbing multiple times with a new starting point each time. The algorithm picks the best solution either as it executes or after it has completed.
**Task:**
* The code below allows you to run a `HillClimber` multiple times each time with a random initial solution.
* Execute the code - how does it compare with the other procedures tested?
* Options:
* Try `SimpleTweak()` instead of `TweakTwoOpt()`
* Try a different `random_seed` or drop it?
* Try a higher `max_iter` (remember this increased runtime!)
```
# Hill Climbing with random restarts
tour = np.arange(70)
objective = SimpleTSPObjective(matrix)
# basic first improvement hill climber
localsearch = HillClimber(objective, tour, TweakTwoOpt(),
maximisation=False)
#random restarts (multiple runs with random init solution)
solver = HillClimberRandomRestarts(objective, localsearch, tour,
maxiter=20, random_seed=101)
print("\nRunning Hill-Climbing with Random Restarts...")
solver.solve()
print("\n** Hill Climbing with Random Restarts OUTPUT ***")
print("best cost:\t{0}".format(solver.best_cost))
print("best solutions:")
print(solver.best_solutions[0])
fig, ax = plot_tour(solver.best_solutions[0], cities, figsize=(12,9))
```
# Exercise 7: Iterated Local Search
A more sophisticated version of Hill-Climbing with random restarts is **Iterated Local Search** or **ILS** for short.
Instead of randomly restarting ILS defaults to a **homebase**. A large tweak operation (called a perturbation!) is applied to the homebase and this is then used an the initial solution for Hill-Climbing. There are a few more parameters/operators to tune with ILS and most important are deciding when to change homebase and what perturbation operator to use. ILS can get quite creative in practice, but the pseudo code below gives the general appearance of the algorithm.
```
function iterated_local_search(init_solution)
best = copy(init_solution)
home = copy(init_solution)
candidate = copy(init_solution)
history = [home]
while time_remains
candidate = local_search(candidate)
if quality(candidate) > quality(best)
best = copy(candidate)
end
home = update_home(home, candidate)
candidate, history = perturb(home, history)
end
return best
end
```
In the algorithm `history` is not always used. If implemented, it essentially gives the algorithm a memory (or tabu list). It contains a list of previous initial solutions used in hill climbing and prevents the algorithm from repeating itself. It usual to have a fixed sized memory (another hyper-parameter to tune!)
The function `update_home` returns a homebase for perturbing. Three simple implementations are a **random walk** and **greedy** and **epsilon greedy**. A random walk uses the last local optima returned from hill climbing. Greedy only accepts a new home base if the new local optima is better than the current homebase. Finally, epsilon greedy takes a random walk epsilon of the time and acts greedy 1 - epsilon of the time. A neat variation on epsilon greedy is to initially allow a lot of exploration and gradually decrease epsilon. However, you will need more iterations (and longer execution time!) to get this to work in practice (but it may return better solutions to large problems).
The function `perturb` is essentially a tweak operator and hence tends to be problem specific. In routing and scheduling problems (and particularly the TSP) a good operator is called the Double Bridge Tweak. This breaks the tour into four parts, reverses and recombines. You could combine with a tabu list if felt it was necessary.
**Task:**
* The code below illustrates how to use an implementation of ILS provided in `metapy`
* Your task to to experiment with ILS and compare how it performs to basic hill climbing with random restarts.
* There are several options you can use to experiment. Uncomment the lines of code to explore the different approaches.
* updating the home base
* perturbing the home base
* the number of iterations of ILS
```
#Iterated Local Search Template
#multiple runs of Hill Climbing with intelligent initial conditions
#random intial solution of size 70
tour = np.arange(70)
np.random.shuffle(tour)
##################################################################
#objective function
objective = SimpleTSPObjective(matrix)
objective = OptimisedSimpleTSPObjective(matrix)
###################################################################
#create the general hill climber with two opt swaps
localsearch = HillClimber(objective, tour,
TweakTwoOpt(),
maximisation=False)
####################################################################
#OPTIONS FOR UPDATING HOMEBASE
#UNCOMMENT THE OPTION YOU WOULD LIKE TO USE.
homebase_accept = EpsilonGreedyHomeBase(epsilon=0.3) #epsilon greedy
#homebase_accept = HigherQualityHomeBase() #greedy method
#homebase_accept = RandomHomeBase() # random walk
#homebase_accept = AnnealingEpsilonGreedyHomeBase(maxiter_per_temp=20,
# verbose=True)
####################################################################
####################################################################
#OPTIONS FOR PERTURBING HOMEBASE (medium to large tweak to homebase)
#UNCOMMENT THE OPTION YOU WOULD LIKE TO USE.
perturb = DoubleBridgePertubation() #no memory
#perturb = TabuDoubleBridgeTweak(tabu_size=10, init_solution=tour) #with tabu
######################################################################
#create the ILS solver
#set verbose=False to suppress output of each iteration.
solver = IteratedLocalSearch(localsearch,
accept=homebase_accept,
perturb=perturb,
verbose=True)
######################################################################
# NO. ITERATIONS OF ILS.
# This is a good parameter to experiment with. Try more than 30.
n = 30
######################################################################
print(f"\nRunning {n} iterations...")
solver.run(n)
print("\n** ILS RESULTS ***")
print("best cost:\t{0}".format(solver.best_cost))
print("best solution:")
print(solver.best_solutions[0])
fig, ax = plot_tour(solver.best_solutions[0], cities, figsize=(12,9))
```
# Optional Exercise 8: Good solutions
**Task:**
* The tours below represent 'good', but not optimal solutions to the st70 problem.
* Can you improve on them? Either by using them as initial solutions in a hill-climbing / iterated local search algorithm or by tuning an evolutionary strategy?
* If you beat them then do tell!
**Hints**
* You can see the cost of each tour by calling `objective.evaluate()`
```
# cost = 688
objective = SimpleTSPObjective(matrix)
tour1 = np.array([45, 26, 67, 43, 29, 19, 13, 27, 48, 54, 25, 7, 2, 31, 41, 17, 3,
1, 6, 18, 23, 14, 56, 62, 65, 21, 22, 37, 58, 34, 68, 30, 69, 12,
28, 35, 0, 15, 46, 36, 57, 49, 50, 64, 63, 10, 55, 66, 47, 53, 61,
33, 20, 11, 32, 59, 51, 9, 4, 52, 5, 40, 42, 16, 8, 39, 60, 38,
44, 24])
objective.evaluate(tour1)
fig, ax = plot_tour(tour1, cities, figsize=(12,9))
#cost = 683
tour2 = np.array([68, 30, 34, 69, 12, 28, 35, 0, 22, 15, 46, 36, 57, 49, 52, 4, 9,
51, 59, 50, 64, 63, 10, 55, 66, 47, 53, 61, 32, 11, 33, 20, 16, 42,
8, 39, 60, 38, 44, 24, 45, 26, 67, 43, 29, 19, 13, 27, 48, 54, 25,
7, 2, 31, 41, 40, 5, 17, 3, 1, 6, 18, 23, 14, 56, 62, 65, 21,
37, 58])
objective.evaluate(tour2)
fig, ax = plot_tour(tour2, cities, figsize=(12,9))
#cost = 686
tour3 = np.array([65, 56, 14, 23, 18, 6, 1, 3, 17, 41, 31, 2, 7, 25, 54, 48, 27,
13, 19, 29, 43, 67, 26, 45, 24, 44, 38, 60, 39, 8, 16, 42, 40, 5,
52, 4, 9, 51, 59, 11, 33, 20, 32, 61, 53, 66, 47, 10, 63, 64, 55,
50, 49, 57, 36, 46, 15, 0, 35, 22, 37, 12, 28, 69, 30, 68, 34, 21,
58, 62])
objective.evaluate(tour3)
fig, ax = plot_tour(tour3, cities, figsize=(12,9))
```
| github_jupyter |
# Cox model
```
import warnings
import arviz as az
import numpy as np
import pymc3 as pm
import scipy as sp
import theano.tensor as tt
from pymc3 import (
NUTS,
Gamma,
Metropolis,
Model,
Normal,
Poisson,
find_MAP,
sample,
starting,
)
from theano import function as fn
from theano import printing
print(f"Running on PyMC3 v{pm.__version__}")
warnings.filterwarnings("ignore")
%config InlineBackend.figure_format = 'retina'
az.style.use("arviz-darkgrid")
```
Here is the original model, implemented in BUGS:
```R
model
{
# Set up data
for(i in 1:Nsubj) {
for(j in 1:T) {
# risk set = 1 if obs.t >= t
Y[i,j] <- step(obs.t[i] - t[j] + eps)
# counting process jump = 1 if obs.t in [ t[j], t[j+1] )
# i.e. if t[j] <= obs.t < t[j+1]
dN[i, j] <- Y[i, j] * step(t[j + 1] - obs.t[i] - eps) * FAIL[i]
}
}
# Model
for(j in 1:T) {
for(i in 1:Nsubj) {
dN[i, j] ~ dpois(Idt[i, j]) # Likelihood
Idt[i, j] <- Y[i, j] * exp(beta[1]*pscenter[i] + beta[2]*
hhcenter[i] + beta[3]*ncomact[i] + beta[4]*rleader[i] + beta[5]*dleader[i] + beta[6]*inter1[i] + beta[7]*inter2[i]) * dL0[j] # Intensity
}
dL0[j] ~ dgamma(mu[j], c)
mu[j] <- dL0.star[j] * c # prior mean hazard
}
c ~ dgamma(0.0001, 0.00001)
r ~ dgamma(0.001, 0.0001)
for (j in 1 : T) { dL0.star[j] <- r * (t[j + 1] - t[j]) }
# next line indicates number of covariates and is for the corresponding betas
for(i in 1:7) {beta[i] ~ dnorm(0.0,0.00001)}
}
```
```
# fmt: off
dta = dict(T=73, Nsubj=430, eps=0.0, t=[1, 21, 85, 128, 129, 148, 178, 204,
206, 210, 211, 212, 225, 238, 241,
248, 259, 273, 275, 281, 286, 289,
301, 302, 303, 304, 313, 317, 323,
344, 345, 349, 350, 351, 355, 356,
359, 364, 385, 386, 389, 390, 391,
392, 394, 395, 396, 397, 398, 399,
400, 406, 415, 416, 426, 427, 434,
435, 437, 441, 447, 448, 449, 450,
451, 453, 455, 456, 458, 459, 460,
461, 462, 463],
obs_t = [460, 313, 435, 350, 435, 350, 350, 460, 460, 448, 225, 225, 396, 435, 396, 396, 453, 396, 456, 397, 397, 396, 395, 275, 449, 395, 395, 462, 302, 302, 458, 461, 396, 241, 389, 458, 304, 304, 395, 395, 364, 460, 415, 463, 396, 459, 441, 435, 396, 458, 437, 396, 356, 356, 396, 455, 396, 462, 399, 400, 350, 350, 395, 395, 441, 355, 85, 458, 128, 396, 386, 386, 386, 462, 458, 390, 390, 396, 396, 396, 427, 458, 395, 275, 275, 395, 359, 395, 395, 441, 395, 463, 178, 275, 463, 396, 396, 259, 396, 396, 458, 441, 396, 463, 396, 463, 435, 396, 437, 396, 398, 463, 460, 462, 460, 460, 210, 396, 435, 458, 385, 323, 323, 359, 396, 396, 460, 238, 441, 450, 392, 458, 396, 458, 396, 396, 462, 435, 396, 394, 396, 435, 458, 1, 395, 395, 451, 462, 458, 462, 396, 286, 396, 349, 449, 462, 455, 21, 463, 461, 461, 456, 435, 396, 460, 462, 462, 435, 435, 460, 386, 396, 458, 386, 461, 441, 435, 435, 463, 456, 396, 275, 460, 406, 460, 406, 317, 406, 461, 396, 359, 458, 463, 435, 462, 458, 396, 396, 273, 396, 435, 281, 275, 396, 447, 225, 447, 396, 435, 416, 396, 248, 396, 435, 435, 396, 461, 385, 396, 458, 458, 396, 461, 396, 448, 396, 396, 460, 455, 456, 463, 462, 458, 463, 396, 462, 395, 456, 396, 463, 396, 435, 459, 396, 396, 396, 395, 435, 455, 395, 461, 344, 396, 395, 396, 317, 396, 395, 426, 461, 396, 289, 441, 395, 396, 458, 396, 396, 435, 396, 395, 396, 441, 345, 396, 359, 435, 435, 396, 396, 395, 458, 461, 458, 212, 301, 458, 456, 395, 396, 395, 435, 396, 396, 303, 458, 460, 400, 396, 462, 359, 458, 396, 206, 441, 396, 458, 396, 462, 396, 396, 275, 396, 395, 435, 435, 462, 225, 458, 462, 396, 396, 289, 396, 303, 455, 400, 400, 359, 461, 396, 462, 460, 463, 463, 463, 204, 435, 435, 396, 396, 396, 463, 458, 396, 455, 435, 396, 396, 463, 396, 461, 463, 460, 441, 460, 435, 435, 460, 455, 460, 395, 460, 460, 460, 435, 449, 463, 462, 129, 391, 396, 391, 391, 434, 356, 462, 396, 349, 225, 396, 435, 461, 391, 391, 351, 211, 461, 212, 434, 148, 356, 458, 456, 455, 435, 463, 463, 462, 435, 463, 437, 460, 396, 406, 451, 460, 435, 396, 460, 455, 396, 398, 456, 458, 396, 456, 449, 396, 128, 396, 462, 463, 396, 396, 396, 435, 460, 396, 458],
FAIL= [1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1],
pscenter= [
-.01434325, -.01460965, .01322687, .00971885, -.03223412, -.01113493, -.01359567, -.03357866, -.0387039, -.0553269, -.03238896, -.07464545, -.07325128, -.07062459, -.07464545, -.07032613, -.0703005, .00965232, -.01408955, .00577483, -.00219072, -.00084567, .01643198, .06509522, .06824313, .07300876, .07300876, .01394272, .06824313, .02063087, .00383186, -.02573045, -.02410864, -.02272752, .05120398, -.00997729, -.00550709, -.02062663, -.03077685, -.01688493, .01035959, .01149963, .01149963, .01149963, .01149963, .01149963, .01149963, .01149963, .01149963, .01149963, .01149963, .01149963, .01149963, .01149963, .01149963, .0034338, .0376236, .00733331, .01520069, .03832785, .03832785, -.02622275, -.02622275, -.02622275, -.01492678, -.02897806, -.02897806, -.02897806, -.02847666, -.031893, -.03919478, -.04224754, -.04743705, -.0510477, -.031893, -.01129093, .01706207, .00193999, -.01503116, .003101, -.00083466, .02395027, -.07952866, -.08559135, -.07251801, -.06586029, -.08432532, -.0613939, -.081205, -.07540084, -.08488011, -.08488011, -.08488011, -.07492433, -.08907269, -.09451609, -.05301854, -.08980743, -.0771635, -.0771635, -.08650947, -.07856082, -.0771635, -.08204606, -.08178245, -.05263504, -.05355574, -.05109092, -.04696729, -.04696729, -.04696729, -.05257489, -.05303248, -.05348096, -.04983674, -.04699414, .00584956, -.00792241, -.01719816, -.02138029, -.01576016, -.04274812, -.04014061, .0471441, .0471441, .0471441, .0471441, .0471441, .0471441, .0471441, .04233112, .0471441, .04233112, .050568, .07388823, .0493324, .04512087, .03205975, .02913185, .06010427, .05324252, .06973204, .05579907, .01212243, .07962459, .05054695, .06672142, .14026688, .01734403, .06078221, .06543709, .06438115, .20126908, -.03138622, -.02180659, .01637333, -.02415774, .01828684, .03106104, .04268495, .01897239, .01591935, -.02367065, -.0619156, -.06403028, -.06851645, -.04821694, -.03889525, -.05023452, -.05013452, -.01557191, -.01171948, -.01362136, -.01174715, -.02707938, -.02634164, -.02634164, -.02634164, -.00692153, -.02381614, -.00890537, -.00611669, -.00894752, -.03551984, -.0252678, -.01513384, -.01016569, -.03551984, -.03773227, -.01978032, .06803483, .06706496, .10551275, .15091534, .03092981, .06556855, .10781559, .12671031, .0936299, .09362991, .09362991, .08294538, .09362991, .09362991, .09362991, .01177025, .02610553, .03546937, .03546937, .03546937, .034415, -.00305626, .04973665, .05103208, .07546701, .05306436, .00824125, .01961115, .01202359, -.02919447, -.01016712, .01756074, -.04035511, -.04753104, -.04463152, -.04845615, -.05010044, .00031411, -.07911871, -.08799869, -.07980882, -.09393142, -.08000018, -.07666632, -.07817401, -.07444922, -.07226554, -.08216553, -.0777643, -.07752042, -.05767992, -.04727952, -.03774814, -.06870384, -.05999847, -.05947695, .02989959, .04627543, .02772475, .02883079, .03642944, .02871235, .04148949, .04240279, .07747082, .07626323, .04268012, .03225577, .06468724, -.05140995, -.05399637, -.05351515, .07302427, .02432223, .0490674, .0490674, .0490674, .0490674, .09013112, .10476315, .10476315, .10476315, .10476315, .10476315, .10476315, .10476315, .10476315, .10476315, .10476315, .10476315, .10476315, .10476315, .07008056, .08666077, .01546215, .01667466, .03417671, .05253941, .04293926, .01496588, .02692172, -.03827151, .04809769, .08742411, .04533176, .01455173, .01831875, .02710811, .09834951, .09952456, .06993483, .02945534, .038731, .1181948, .04435538, .04435538, -.02357505, .05824019, .05820741, -.02357505, .09324722, .15534712, .07207468, .04692869, -.03490683, -.04404809, -.05054474, -.05325826, -.0474724, -.04905931, .01068221, .02879751, .00852646, .02693032, .01835589, .02989959, .02989959, .02989959, .04976377, .04439012, .03397319, .02989959, .02989959, .05468828, .04463226, .05886378, .06311052, .02989959, .04595331, .04203459, .01231324, -.01399783, .04595331, .00145386, .04601278, .06459354, -.0007196, .00012216, -.07614055, -.08435525, -.07957162, -.10299519, -.08156988, -.08225659, -.07449063, -.00210284, -.00797183, -.025355, -.01258251, -.04372031, -.03985972, -.03545086, -.03384566, -.04025533, -.07523724, -.05947702, -.061286, -.07666647, -.07663169, -.05902354, -.07652324, -.07645561, -.06258684, -.09604834, -.08813326, -.03292062, -.07848112, -.08239502, -.08316891, -.07244316, -.075417, -.07652324, -.07922532, -.08755959, -.08583414, -.07450142, -.08066016, -.06057205, -.07652324, -.06249051, -.08781742, -.086076, -.07652324, -.07696518, -.0618688, -.06073988, -.06524737, -.04419825, -.04489509, -.04390368, -.04358438, -.04489509, -.04520512, -.04187583, -.03653955, -.03973426, -.03753508, -.03569439, -.06789339, .06689456, .05526327, .05139003, .02641841, .04891529, .07078697, .06862645, .06832582, .04104258, -.00120631, .01947345, .04891779, .04891779, .03561932, .02576244, .03158225, .03608047, .08685057, .04632537, .06841581, -.02899643],
hhcenter= [ -.78348798, -.63418788, -.91218799, -.98388809, -.23518796, .11481193, -1.415588, -1.2535881, -.55738801, -.88128799, -1.109488, .05721192, -1.045788, -.30888793, .29651192, -.36688802, -.50058788, .02271203, -.59088796, -.04198809, .50561196, -.07418796, .98481184, .78921205, .09431199, -.06488796, 2.1662121, .08891205, 1.4004121, 1.316112, 1.9362121, 2.0107121, 1.150712, .31951192, -.23918791, -.1562881, -.9575879, -.07728811, .29641202, 1.2273121, 1.7717118, 1.5764117, .14181189, .72131211, 1.279212, .68241197, -.72808808, -.00488802, -.23938794, -1.000788, .55081207, -.52348799, 1.780612, -.35888812, .36481193, 1.5480118, -.03078791, 1.389112, .30211189, .70901209, -.16668792, 1.435812, .47001198, 2.0838118, 1.1673121, .18461208, -.30608794, 1.4470119, .23301201, -.58458799, .44011191, -.61948794, -.41388795, .263212, .66171199, .92451197, .78081208, .90991193, 1.6920118, 1.334012, 1.2101121, .41591194, -.48498794, -.73278803, -1.093588, .09911207, -.93418807, -.46908805, .0205119, .0535119, -.14228792, -.55708808, -.45498797, -.54008788, -.30998799, -.10958811, -.0960879, -.01338812, -.88168806, -.51788801, .36801198, .46621206, .13271193, -.11208793, -.76768798, -.54508799, -1.2773881, .16641192, .95871216, -.48238799, 1.6281118, -.18848796, -.49718806, -.41348812, -.31628796, -.59528798, -.11718794, -.57058805, -.59488791, -.21248789, -.65658802, -.56298798, -.52698797, -.65758795, -.04988809, .55341202, -.76328796, .254612, 1.3500118, -.54958791, 1.665812, .14671211, 1.963912, .29161194, -.56838793, 1.9371119, .90991193, -.39558789, .39521196, -.55208796, -.05268808, -.77368802, -.45428798, .05841212, -.45308802, -.12458798, .01431207, -.28228804, .79281193, -.26358792, -.54738802, -.38158795, -.54118794, -.72828788, -.58128804, .355912, -.24078794, -1.0384881, -.75038809, -.41018793, -.43538806, -1.566388, -.53388804, -.28388807, -1.2348881, -.69028801, -1.620088, -.78128809, -.54648799, -.92738789, .11871199, .26851204, .61571199, .82891208, 1.1985121, 1.012012, 1.0602121, -.02988811, .79301196, .67731196, .43991187, .9404121, .5254119, 1.0365119, 1.6220121, .61671191, -.50318807, 2.6073117, .02361206, -.60438794, -.79278797, -.18108793, -.48178813, -.44038793, -.22628804, -.07398792, .519512, .40211204, .582012, 1.830512, .80441195, .58801204, -.56368798, -1.5451881, .45991209, -.23448797, -.36918804, 1.3247118, .19541197, -.20818801, 1.163012, -.78228801, -.6048879, -.575288, 1.3241119, .0147119, -.76518792, -.37478802, -.35508797, -.90038794, -1.250888, -.46608803, -.98488802, -1.5185881, -.90908808, -1.048188, -.90138787, -.77278799, -1.248988, -.34448811, -.61628789, .38531187, -.51728791, -.00878807, -.60078806, -.45358798, .46301201, -.22048803, -.71518797, -.76478809, -.75028795, -.4952881, .01731209, -.83718795, .57951194, .54291207, .45341209, .16941194, 1.054112, .61721212, 2.2717118, 1.1593118, 2.0280118, .92281204, 1.0100121, -.1866879, 2.6503119, 2.3914118, -.19948788, -.36418793, -.9259879, -.71058792, -.1104879, .16971211, 1.474812, 1.9360118, 2.5344119, 2.0171118, 1.9387121, .55071193, -.03918811, .20681195, .40421203, -.75518793, -.45678803, -1.0271881, .77211195, 1.146812, -1.147788, -1.565588, -.34888789, 1.303812, 1.952312, 1.639112, .07731203, .25901201, -.45608804, -.5028879, .03641204, -.03808804, .38571194, .31831196, -.17648788, -.44528791, -.55918807, -.53108805, .39721206, -.06328794, -.34038803, -.05988808, -.89548796, -.03518792, .045512, -.1859879, -.039288, -.82568806, .01431207, .40091208, -.2531881, .030412, -.31918809, -.54958791, -.79078788, .36691192, -.324388, -1.0082881, -1.232188, -.53248805, -.23678799, -.89188808, .25111201, -.6766879, -.3565881, -.61228794, -.21078797, -1.0343881, -.58358806, -.15588804, -.39238808, -.67818803, -.19498797, 1.099412, 1.2767119, -.64068788, -.50678796, -.64058799, -.86918801, 1.4048119, -.59648794, .23331194, .68371207, .11251191, -.17128797, .17081194, -.44218799, -.48708794, .09591202, .20131211, -.20108791, -.02158805, -.48188803, -.3012881, -.55008787, -1.146188, -.82128805, -.87638801, -.54488796, -.60288805, -1.003088, -.25078794, -.14818807, -.14738794, -.80938786, -.85988802, -.90188807, -.94998807, -.75718802, -.37418792, -.66708797, 1.0981121, 1.1441121, .47381189, -.12958808, -.34358808, -.84328789, -.33498809, -.98088807, -.6903879, -1.284988, -.80838794, -.91838807, -.81848806, -.34488794, -.83438796, .12971191, .99381214, -.91608804, -.31808802, -.01018806, .98171192, -.91638798, -1.043988, -1.0103881, 1.451612, -.01528808, .02441196, -.41458794, .25691202, .18601207, -.815988, -.02908798, -.59088796, -.35608789, .79691201, 1.8123121, -.98588794, 1.548912, 2.3653121, -.09238812, .96741205, .05891208, -.15618797, -.5660879, -.28338811, -.10088798, 1.1663117, .21981196, .07151202, -.009088, -.49578807, .15441208, -.44488809, -.2677879, -.54388803, -.25468799, .68631202, -.88128799, -.84628791, -1.2549881, -.36198804],
ncomact= [ 0, 0, 0, 0, 0, 1, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 1, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 1, 1, 1, 1, 0, 0, 0, 0, 1, 0, 1, 0, 0, 0, 1, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 0, 0, 0, 1, 1, 1, 0, 0, 0, 0, 0, 1, 0, 1, 1, 0, 1, 0, 0, 0, 1, 0, 0, 0, 1, 1, 0, 0, 0, 0, 0, 0, 1, 1, 0, 0, 1, 0, 0, 0, 1, 1, 0, 0, 1, 1, 1, 1, 1, 0, 1, 1, 0, 1, 1, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 1, 0, 0, 0, 0, 0, 1, 0, 0, 0, 1, 1, 1, 1, 1, 0, 0, 0, 0, 1, 0, 0, 1, 1, 0, 1, 0, 0, 1, 1, 0, 0, 1, 0, 0, 0, 0, 0, 1, 0, 1, 0, 0, 0, 0, 1, 1, 0, 0, 0, 1, 0, 0, 0, 1, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 1, 0, 0, 1, 0, 1, 1, 0, 0, 0, 0, 0, 0, 0, 1, 1, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 1, 1, 0, 0, 1, 1, 0, 1, 1, 0, 1, 0, 1, 1, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 1, 1, 0, 1, 0, 1, 0, 0, 0, 1, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 1, 0, 0, 0, 1, 0, 1, 1, 0, 0, 0, 1, 0, 1, 0, 0, 0, 0, 0, 1, 0, 0, 0, 1, 0, 0, 1, 0, 1, 1, 0, 0, 0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 1, 0, 1, 0, 0, 0, 1, 1, 1, 1, 1, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 1, 0, 0, 0, 0, 1, 0, 0, 0, 0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 1, 0, 0, 1, 0, 1, 0, 0, 0, 1, 0, 1, 0, 1, 0, 1, 1, 0, 0, 1, 0, 0, 0, 0, 1],
rleader= [ 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0],
dleader= [ 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 1, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0],
inter1= [ -.01434325, -.01460965, 0, 0, 0, -.01113493, 0, 0, 0, -.0553269, -.03238896, 0, 0, -.07062459, -.07464545, -.07032613, 0, 0, -.01408955, 0, -.00219072, 0, 0, 0, 0, 0, .07300876, .01394272, 0, 0, 0, 0, 0, 0, .05120398, 0, -.00550709, -.02062663, -.03077685, -.01688493, 0, .01149963, 0, .01149963, .01149963, 0, 0, 0, 0, 0, 0, 0, 0, 0, .01149963, .0034338, .0376236, .00733331, 0, .03832785, .03832785, -.02622275, -.02622275, -.02622275, -.01492678, 0, 0, -.02897806, -.02847666, 0, 0, -.04224754, -.04743705, -.0510477, -.031893, 0, 0, 0, -.01503116, .003101, -.00083466, .02395027, -.07952866, 0, 0, -.06586029, 0, -.0613939, -.081205, -.07540084, -.08488011, -.08488011, 0, -.07492433, -.08907269, -.09451609, 0, -.08980743, 0, -.0771635, 0, 0, -.0771635, -.08204606, 0, -.05263504, 0, -.05109092, -.04696729, 0, -.04696729, 0, -.05303248, -.05348096, 0, 0, .00584956, -.00792241, -.01719816, 0, -.01576016, 0, -.04014061, 0, 0, 0, 0, 0, .0471441, 0, .04233112, 0, .04233112, 0, 0, .0493324, .04512087, .03205975, .02913185, 0, .05324252, 0, 0, 0, 0, .05054695, 0, .14026688, .01734403, .06078221, 0, 0, 0, -.03138622, 0, .01637333, 0, 0, 0, 0, .01897239, .01591935, 0, -.0619156, 0, -.06851645, 0, -.03889525, -.05023452, -.05013452, 0, 0, -.01362136, 0, 0, -.02634164, 0, 0, 0, 0, -.00890537, -.00611669, 0, 0, 0, -.01513384, 0, -.03551984, 0, -.01978032, 0, .06706496, .10551275, 0, .03092981, .06556855, 0, 0, 0, .09362991, 0, 0, 0, 0, 0, 0, .02610553, .03546937, 0, 0, .034415, 0, 0, 0, .07546701, 0, 0, 0, 0, -.02919447, -.01016712, 0, 0, 0, 0, -.04845615, -.05010044, 0, 0, 0, 0, 0, 0, -.07666632, 0, 0, -.07226554, -.08216553, -.0777643, 0, 0, -.04727952, 0, -.06870384, -.05999847, 0, 0, 0, .02772475, .02883079, .03642944, 0, .04148949, 0, 0, 0, .04268012, .03225577, 0, -.05140995, -.05399637, 0, 0, .02432223, 0, .0490674, .0490674, .0490674, 0, 0, 0, 0, 0, 0, 0, 0, .10476315, 0, 0, 0, 0, 0, .07008056, 0, 0, .01667466, 0, .05253941, .04293926, 0, .02692172, 0, 0, .08742411, .04533176, 0, .01831875, 0, .09834951, .09952456, 0, .02945534, .038731, 0, .04435538, 0, -.02357505, 0, 0, -.02357505, .09324722, 0, 0, 0, -.03490683, 0, -.05054474, 0, -.0474724, -.04905931, 0, .02879751, 0, 0, 0, 0, 0, 0, 0, .04439012, 0, .02989959, .02989959, .05468828, .04463226, 0, 0, 0, 0, 0, .01231324, -.01399783, .04595331, .00145386, 0, .06459354, -.0007196, 0, -.07614055, -.08435525, 0, -.10299519, 0, 0, 0, -.00210284, -.00797183, 0, 0, 0, 0, -.03545086, 0, 0, 0, 0, -.061286, -.07666647, 0, -.05902354, -.07652324, -.07645561, 0, 0, 0, -.03292062, 0, 0, 0, 0, -.075417, 0, -.07922532, 0, -.08583414, -.07450142, -.08066016, 0, 0, -.06249051, 0, 0, 0, 0, -.0618688, 0, -.06524737, -.04419825, -.04489509, 0, 0, 0, -.04520512, -.04187583, 0, 0, -.03753508, 0, 0, 0, 0, 0, 0, 0, 0, .06862645, 0, 0, -.00120631, .01947345, 0, 0, .03561932, 0, .03158225, .03608047, 0, 0, 0, -.02899643],
inter2= [-.78348798, -.63418788, 0, 0, 0, .11481193, 0, 0, 0, -.88128799, -1.109488, 0, 0, -.30888793, .29651192, -.36688802, 0, 0, -.59088796, 0, .50561196, 0, 0, 0, 0, 0, 2.1662121, .08891205, 0, 0, 0, 0, 0, 0, -.23918791, 0, -.9575879, -.07728811, .29641202, 1.2273121, 0, 1.5764117, 0, .72131211, 1.279212, 0, 0, 0, 0, 0, 0, 0, 0, 0, .36481193, 1.5480118, -.03078791, 1.389112, 0, .70901209, -.16668792, 1.435812, .47001198, 2.0838118, 1.1673121, 0, 0, 1.4470119, .23301201, 0, 0, -.61948794, -.41388795, .263212, .66171199, 0, 0, 0, 1.6920118, 1.334012, 1.2101121, .41591194, -.48498794, 0, 0, .09911207, 0, -.46908805, .0205119, .0535119, -.14228792, -.55708808, 0, -.54008788, -.30998799, -.10958811, 0, -.01338812, 0, -.51788801, 0, 0, .13271193, -.11208793, 0, -.54508799, 0, .16641192, .95871216, 0, 1.6281118, 0, -.49718806, -.41348812, 0, 0, -.11718794, -.57058805, -.59488791, 0, -.65658802, 0, -.52698797, 0, 0, 0, 0, 0, 1.3500118, 0, 1.665812, 0, 1.963912, 0, 0, 1.9371119, .90991193, -.39558789, .39521196, 0, -.05268808, 0, 0, 0, 0, -.12458798, 0, -.28228804, .79281193, -.26358792, 0, 0, 0, -.72828788, 0, .355912, 0, 0, 0, 0, -.43538806, -1.566388, 0, -.28388807, 0, -.69028801, 0, -.78128809, -.54648799, -.92738789, 0, 0, .61571199, 0, 0, 1.012012, 0, 0, 0, 0, .43991187, .9404121, 0, 0, 0, .61671191, 0, 2.6073117, 0, -.60438794, 0, -.18108793, -.48178813, 0, -.22628804, -.07398792, 0, 0, 0, 1.830512, 0, 0, 0, 0, 0, 0, -.36918804, 1.3247118, 0, 0, 1.163012, 0, 0, 0, 1.3241119, 0, 0, 0, 0, -.90038794, -1.250888, 0, 0, 0, 0, -1.048188, -.90138787, 0, 0, 0, 0, 0, 0, -.00878807, 0, 0, .46301201, -.22048803, -.71518797, 0, 0, -.4952881, 0, -.83718795, .57951194, 0, 0, 0, 1.054112, .61721212, 2.2717118, 0, 2.0280118, 0, 0, 0, 2.6503119, 2.3914118, 0, -.36418793, -.9259879, 0, 0, .16971211, 0, 1.9360118, 2.5344119, 2.0171118, 0, 0, 0, 0, 0, 0, 0, 0, .77211195, 0, 0, 0, 0, 0, 1.952312, 0, 0, .25901201, 0, -.5028879, .03641204, 0, .38571194, 0, 0, -.44528791, -.55918807, 0, .39721206, 0, -.34038803, -.05988808, 0, -.03518792, .045512, 0, -.039288, 0, .01431207, 0, 0, .030412, -.31918809, 0, 0, 0, -.324388, 0, -1.232188, 0, -.23678799, -.89188808, 0, -.6766879, 0, 0, 0, 0, 0, 0, 0, -.67818803, 0, 1.099412, 1.2767119, -.64068788, -.50678796, 0, 0, 0, 0, 0, .68371207, .11251191, -.17128797, .17081194, 0, -.48708794, .09591202, 0, -.20108791, -.02158805, 0, -.3012881, 0, 0, 0, -.87638801, -.54488796, 0, 0, 0, 0, -.14738794, 0, 0, 0, 0, -.75718802, -.37418792, 0, 1.0981121, 1.1441121, .47381189, 0, 0, 0, -.33498809, 0, 0, 0, 0, -.91838807, 0, -.34488794, 0, .12971191, .99381214, -.91608804, 0, 0, .98171192, 0, 0, 0, 0, -.01528808, 0, -.41458794, .25691202, .18601207, 0, 0, 0, -.35608789, .79691201, 0, 0, 1.548912, 0, 0, 0, 0, 0, 0, 0, 0, 1.1663117, 0, 0, -.009088, -.49578807, 0, 0, -.2677879, 0, -.25468799, .68631202, 0, 0, 0, -.36198804])
#fmt: off
def load_data_cox(dta):
array = lambda x : np.array(dta[x], dtype=float)
t = array('t')
obs_t = array('obs_t')
pscenter = array('pscenter')
hhcenter = array('hhcenter')
ncomact = array('ncomact')
rleader = array('rleader')
dleader = array('dleader')
inter1 = array('inter1')
inter2 = array('inter2')
fail = array('FAIL')
return (t, obs_t, pscenter, hhcenter, ncomact,
rleader, dleader, inter1, inter2, fail)
(t, obs_t, pscenter, hhcenter, ncomact, rleader,
dleader, inter1, inter2, fail) = load_data_cox(dta)
X = np.array([pscenter, hhcenter, ncomact, rleader, dleader, inter1, inter2])
X.shape
with Model() as model:
T = len(t) - 1
nsubj = len(obs_t)
# risk set equals one if obs_t >= t
Y = np.array([[int(obs >= time) for time in t] for obs in obs_t])
# counting process. jump = 1 if obs_t \in [t[j], t[j+1])
dN = np.array([[Y[i,j]*int(t[j+1] >= obs_t[i])*fail[i] for j in range(T)] for i in
range(nsubj)])
c = Gamma('c', .0001, .00001)
r = Gamma('r', .001, .0001)
dL0_star = r*np.diff(t)
# prior mean hazard
mu = dL0_star * c
dL0 = Gamma('dL0', mu, c, shape=T)
beta = Normal('beta', np.zeros(7),
np.ones(7)*100, shape=7)
linear_model = tt.exp(tt.dot(X.T, beta))
idt = Y[:, :-1] * tt.outer(linear_model, dL0)
dn_like = Poisson('dn_like', idt, observed=dN)
with model:
trace = sample(2000, n_init=10000, init='advi_map')
az.plot_trace(trace, var_names=['c', 'r']);
az.plot_forest(trace, var_names=['beta']);
%load_ext watermark
%watermark -n -u -v -iv -w
```
| github_jupyter |
## ------- >--------- >----------PLAYSTORE ANALYSIS USING PYTHON-------- >----------- >--------- ##
# BY :
# ARAVINTH.S
# BE - COMP SCIENCE ENGINEERING
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
ps = pd.read_csv("plst.csv")
ps.head()
ps.shape
ps.size
ps.info()
ps.describe(include = 'all')
ps.isnull().sum()
```
# Task 1 : Data clean up – Missing value treatment
## a. Drop records where rating is missing since rating is our target/study variable
```
ps.dropna(subset = ['Rating'],axis = 0,how = 'any',inplace = True)
ps['Rating'].isnull().sum()
```
## b. Check the null values for the Android Ver column.
## i. Are all 3 records having the same problem?
```
ps.loc[ps['Android Ver'].isnull()]
```
YES all 3 record has same problem
## ii. Drop the 3rd record i.e. record for “Life Made WIFI …”
```
ps.drop([10472],inplace = True)
ps.loc[ps['Android Ver'].isnull()]
```
# iii. Replace remaining missing values with the mode
```
ps['Android Ver'].fillna(ps["Android Ver"].mode()[0],inplace = True)
```
# c. Current ver – replace with most common value
```
ps['Current Ver'].fillna(ps["Current Ver"].mode(),inplace = True)
```
# 2. Data clean up – correcting the data types
# a. Which all variables need to be brought to numeric types?
Price and installs are need to brought to numeric type
# b. Price variable – remove $ sign and convert to float
```
ps['Price'] = ps['Price'].replace('[\$,]','',regex = True).astype(float)
```
# c. Installs – remove ‘,’ and ‘+’ sign, convert to integer
```
install = []
for i in ps['Installs']:
install.append(i.replace(',','').replace('+','').strip())
ps['Installs'] = install
ps['Installs'] = ps['Installs'].astype(int)
```
# d. Convert all other identified columns to numeric
All other columns are in numeric
# 3. Sanity checks – check for the following and handle accordingly
# a. Avg. rating should be between 1 and 5, as only these values are allowed on the play store.
# i. Are there any such records? Drop if so
```
ps.loc[ps.Rating < 1] & ps.loc[ps.Rating > 5]
```
There is no such record found.
# b. Reviews should not be more than installs as only those who installed can review the app.
# i. Are there any such records? Drop if so.
```
temp = ps.loc[ps['Reviews'].astype(int) > ps['Installs'].astype(int)].index
temp
```
There are 7 rows where reviews more than installs.
```
ps.drop(temp,inplace = True)
ps.loc[ps['Reviews'].astype(int) > ps['Installs'].astype(int)]
```
Now we deleted all the seven records.
# 4. Identify and handle outliers –
# a. Price column
# i. Make suitable plot to identify outliers in price
```
plt.figure(figsize = (5,5))
plt.boxplot(ps['Price'])
plt.show()
```
There are some outliers in price column
# ii. Do you expect apps on the play store to cost $200? Check out these cases
```
ps.loc[ps['Price'] == 200]
```
There is no app on the playstore to cost $200
# iii. After dropping the useless records, make the suitable plot again to identify outliers
```
plt.boxplot(ps['Price'] <300)
plt.show()
```
# iv. Limit data to records with price < $30
```
ps["Price"] = ps['Price'] < 30
```
# b. Reviews column
# i. Make suitable plot
```
sns.displot(ps['Reviews'],bins = 30,kde = False)
plt.show()
```
# ii. Limit data to apps with < 1 Million reviews
```
lr = ps.loc[ps['Reviews'].astype(int) > 1000000].index
ps.drop(labels = lr,inplace = True )
lr.value_counts().sum()
```
# c. Installs
# i. What is the 95th percentile of the installs?
```
print("95 percentile of installs :",np.percentile(ps['Installs'],95))
percentile = ps.Installs.quantile(0.95)
percentile
```
# ii. Drop records having a value more than the 95th percentile
```
ints = ps.loc[ps['Installs'].astype(int) > ps.Installs.quantile(0.95)].index
ints.value_counts().sum()
ps.drop(labels = ints , inplace = True)
```
# Data analysis to answer business questions
# 5. What is the distribution of ratings like? (use Seaborn) More skewed towards higher/lower values?
# a. How do you explain this?
```
sns.displot(ps['Rating'])
plt.show()
print(ps["Rating"].skew())
print(ps['Rating'].mean())
print(ps['Rating'].median())
```
# b. What is the implication of this on your analysis?
```
ps['Rating'].mode()
```
since MODE >= MEDIAN > MEAN,the distribution of rating is negatively skewed and more skewed towards lower values
# 6. What are the top Content Rating values?
# a. Are there any values with very few records?
```
ps['Content Rating'].value_counts()
```
YES Adults only 18+ and unrated have very few records
# b. If yes, drop those as they won’t help in the analysis
```
cr = []
for i in ps['Content Rating']:
cr.append(i.replace('Adults only 18+','NaN').replace('Unrated','NaN'))
ps['Content Rating'] = cr
temp = ps[ps['Content Rating'] == 'NaN'].index
ps.drop(labels = temp , inplace = True)
ps['Content Rating'].value_counts()
```
# 7. Effect of size on rating
# a. Make a joinplot to understand the effect of size on rating
```
sns.jointplot(y = 'Size',x = 'Rating',data = ps,kind = 'scatter')
plt.show()
```
# b. Do you see any patterns?
```
YES we see pattern between Rating and Size and they are correlated with each other.
```
# c. How do you explain the pattern?
Generally on increasing Rating, Size of App also increases. But this is not always true ie. for higher Rating, their is constant Size.Here we can see size of app increases when the rating of app increase so it is highly positively correlated.
# 8. Effect of price on rating
# a. Make a jointplot (with regression line)
```
sns.jointplot(x = 'Rating',y = 'Price',data = ps,kind = 'reg')
plt.show()
```
# b. What pattern do you see?
Here increasing the price rating constantly stands above four.
# c. How do you explain the pattern?
Since increasing the price,rating almost constantly above the four so it has poor highly positively correlation between Price and Rating
```
ps.corr()
```
# d. Replot the data, this time with only records with price > 0
```
temp = ps.loc[ps["Price"] > 0]
sns.jointplot(x = 'Rating',y = 'Price',data = temp,kind = 'reg')
plt.show()
```
# e. Does the pattern change?
yes,there is slight changes in the pattern after removing 0 from records.
# f. What is your overall inference on the effect of price on the rating
Generally increasing in price does not have significant effect on rating,Here increasing in price increases the rating and it is almost constant after four.
# 9. Look at all the numeric interactions together –
# a. Make a pairplort with the colulmns - 'Reviews', 'Size', 'Rating', 'Price'
```
sns.pairplot(ps,vars = ['Rating','Reviews','Size','Price'])
plt.show()
```
# 10. Rating vs. content rating
# a. Make a bar plot displaying the rating for each content rating
```
ps.groupby(['Content Rating'])['Rating'].count().plot.bar(color="darkgreen")
plt.show()
```
# b. Which metric would you use? Mean? Median? Some other quantile?
```
plt.boxplot(ps.Rating)
plt.show
```
we can there some outliers in rating so we can go for median because in case of outliers median is the best measure of central tendancy
# c. Choose the right metric and plot
```
ps.groupby(['Content Rating'])['Rating'].median().plot.barh(color="red")
plt.show()
```
# 11. Content rating vs. size vs. rating – 3 variables at a time
# a. Create 5 buckets (20% records in each) based on Size
```
bins =[0,20000,40000,60000,80000,100000]
ps['Bucket size'] = pd.cut(ps['Size'],bins,labels = ['0-20k','20-40k','40-60k','60-80k','80-100k'])
pd.pivot_table(ps,values = 'Rating',index = 'Bucket size',columns = 'Content Rating')
```
# b. By Content Rating vs. Size buckets, get the rating (20th percentile) for each combination
```
temp = pd.pivot_table(ps,values = 'Rating',index = 'Bucket size',columns = 'Content Rating',aggfunc = lambda x : np.quantile(x,0.2))
temp
```
# c. Make a heatmap of this
# i. Annotated
```
sns.heatmap(data = temp,annot = True,linewidths = 5,linecolor = 'black')
plt.show()
```
# ii. Greens color map
```
sns.heatmap(data = temp,annot = True,linewidths = 5,linecolor = 'black',cmap = 'Greens')
plt.show()
```
# d. What’s your inference? Are lighter apps preferred in all categories? Heavier? Some?
Based on analysis,we saw that lighter app are not preferred in all categories.App with size between 60k-80k and 80k-100k are highly rated in all categories so we can conclude heavier appa are preferred in all categories.
| github_jupyter |
# Example Seldon Core Deployments using Helm with Istio
Prequisites
* [Install istio](https://istio.io/latest/docs/setup/getting-started/#download)
## Setup Cluster and Ingress
Use the setup notebook to [Setup Cluster](https://docs.seldon.io/projects/seldon-core/en/latest/examples/seldon_core_setup.html#Setup-Cluster) with [Istio Ingress](https://docs.seldon.io/projects/seldon-core/en/latest/examples/seldon_core_setup.html#Istio). Instructions [also online](https://docs.seldon.io/projects/seldon-core/en/latest/examples/seldon_core_setup.html).
```
!kubectl create namespace seldon
!kubectl config set-context $(kubectl config current-context) --namespace=seldon
```
## Configure Istio
For this example we will create the default istio gateway for seldon which needs to be called `seldon-gateway`. You can supply your own gateway by adding to your SeldonDeployments resources the annotation `seldon.io/istio-gateway` with values the name of your istio gateway.
Create a gateway for our istio-ingress
```
%%writefile resources/seldon-gateway.yaml
apiVersion: networking.istio.io/v1alpha3
kind: Gateway
metadata:
name: seldon-gateway
namespace: istio-system
spec:
selector:
istio: ingressgateway # use istio default controller
servers:
- port:
number: 80
name: http
protocol: HTTP
hosts:
- "*"
!kubectl create -f resources/seldon-gateway.yaml -n istio-system
```
Ensure the istio ingress gatewaty is port-forwarded to localhost:8004
* Istio: `kubectl port-forward $(kubectl get pods -l istio=ingressgateway -n istio-system -o jsonpath='{.items[0].metadata.name}') -n istio-system 8004:8080`
```
ISTIO_GATEWAY="localhost:8004"
VERSION=!cat ../version.txt
VERSION=VERSION[0]
VERSION
```
## Start Seldon Core
Use the setup notebook to [Install Seldon Core](https://docs.seldon.io/projects/seldon-core/en/latest/examples/seldon_core_setup.html#Install-Seldon-Core) with Istio Ingress. Instructions [also online](https://docs.seldon.io/projects/seldon-core/en/latest/examples/seldon_core_setup.html).
## Serve Single Model
```
!helm install mymodel ../helm-charts/seldon-single-model --set model.image=seldonio/mock_classifier:$VERSION
!helm template mymodel ../helm-charts/seldon-single-model --set model.image=seldonio/mock_classifier:$VERSION | pygmentize -l json
!kubectl rollout status deploy/mymodel-default-0-model
```
### Get predictions
```
from seldon_core.seldon_client import SeldonClient
sc = SeldonClient(deployment_name="mymodel",namespace="seldon",gateway_endpoint=ISTIO_GATEWAY)
```
#### REST Request
```
r = sc.predict(gateway="istio",transport="rest")
assert(r.success==True)
print(r)
```
## gRPC Request
```
r = sc.predict(gateway="istio",transport="grpc")
assert(r.success==True)
print(r)
!helm delete mymodel
```
| github_jupyter |
<a href="https://colab.research.google.com/github/agemagician/CodeTrans/blob/main/prediction/transfer%20learning%20fine-tuning/source%20code%20summarization/csharp/small_model.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
**<h3>Summarize the csharp source code using codeTrans transfer learning finetuning model</h3>**
<h4>You can make free prediction online through this
<a href="https://huggingface.co/SEBIS/code_trans_t5_small_source_code_summarization_csharp_transfer_learning_finetune">Link</a></h4> (When using the prediction online, you need to parse and tokenize the code first.)
**1. Load necessry libraries including huggingface transformers**
```
!pip install -q transformers sentencepiece
from transformers import AutoTokenizer, AutoModelWithLMHead, SummarizationPipeline
```
**2. Load the token classification pipeline and load it into the GPU if avilabile**
```
pipeline = SummarizationPipeline(
model=AutoModelWithLMHead.from_pretrained("SEBIS/code_trans_t5_small_source_code_summarization_csharp_transfer_learning_finetune"),
tokenizer=AutoTokenizer.from_pretrained("SEBIS/code_trans_t5_small_source_code_summarization_csharp_transfer_learning_finetune", skip_special_tokens=True),
device=0
)
```
**3 Give the code for summarization, parse and tokenize it**
```
code = "public static DateTime ParseUnixDateTime(double unixTime)\n {\n var dt= new DateTime(1970, 1, 1, 0, 0, 0, 0, System.DateTimeKind.Utc);\n dt= dt.AddSeconds(unixTimeStamp).ToLocalTime();\n return dt;\n }" #@param {type:"raw"}
!pip install antlr4-python3-runtime==4.5.2
import antlr4
!wget https://www.dropbox.com/s/o87gk1jxf8645eu/CSharp4Lexer.py?dl=1 -O CSharp4Lexer.py
from CSharp4Lexer import CSharp4Lexer
def csTokenizer(line):
l = line.replace('\\n', '\n')
parsedVersion = []
stream = antlr4.InputStream(l)
lexer = CSharp4Lexer(stream)
toks = antlr4.CommonTokenStream(lexer)
toks.fetch(500)
identifiers = {}
identCount = 0
for token in toks.tokens:
if token.type == 109:
parsedVersion.append("CODE_INTEGER")
elif token.type == 111:
parsedVersion.append("CODE_REAL")
elif token.type == 112:
parsedVersion.append("CODE_CHAR")
elif token.type == 113:
parsedVersion.append("CODE_STRING")
elif token.type == 9 or token.type == 7 or token.type == 6 or token.type == 4 or token.type == 8 or token.type == 5: # whitespace and comments and newline
pass
else:
parsedVersion.append(str(token.text))
parsedVersion.remove('<EOF>')
return ' '.join(parsedVersion)
tokenized_code = csTokenizer(code)
print("code after tokenization: " + tokenized_code)
```
**4. Make Prediction**
```
pipeline([tokenized_code])
```
| github_jupyter |
# Spam Filter using Naive Bayes Classifier
```
import os
print(os.listdir("../input"))
```
**Import libraries**
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
```
**Read csv file**
```
df = pd.read_csv('../input/spam.csv', encoding='latin-1')[['v1', 'v2']]
```
Viewing after renaming the columns
```
df.columns = ['label', 'message']
df.head()
```
View the lable statistics
```
df.groupby('label').describe()
```
View the counts of ham and spam present in label column
```
sns.countplot(data=df, x='label')
```
**Steps**
1. Clean and Normalize text
2. Convert text into vectors (BoW) we use TfIdf
3. Train and test Classifier
**Cleaning steps**
<br>
It will be done in following steps:
<br>
1. Remove punctuations
2. Remove all stopwords
3. Apply [stemming](https://en.wikipedia.org/wiki/Stemming) (get the stem of the word).
** Write a method to return normailzed text in form of tokens (lemmas)**
```
import string
from nltk.corpus import stopwords
from nltk import PorterStemmer as Stemmer
def pre_process(text):
text = text.lower()
text = ''.join([t for t in text if t not in string.punctuation])
text = [t for t in text.split() if t not in stopwords.words('english')] # All words other tha stopwrods and punctuations in lowercase
st = Stemmer()
text = [st.stem(t) for t in text]
return text
pre_process('It\'s holiday lads :D. Mount is playing very well!!!')
# Test with our dataset
df['message'][:21].apply(pre_process)
```
Refer scikitlearn for details on TfIDf

**Fit and transform SMS corpus**
```
from sklearn.feature_extraction.text import TfidfVectorizer
tfidf= TfidfVectorizer(analyzer=pre_process)
data = tfidf.fit_transform(df['message'])
message=df.iloc[2]
print(tfidf.transform(df.iloc[2]))
```
**Having messages in form of vectors, we are ready to train our classifier. <br>We will use Naive Bayes which is well known classifier while working with text data.
<br>Before that we will use pipeline feature of sklearn to create a pipeline of TfidfVectorizer followed by Classifier.**
<br>Input will be message passed to first stage TfidfVectorizer which will transform it and pass it to Naive Bayes Classifier to get output label
```
from sklearn.base import TransformerMixin
class DenseTransformer(TransformerMixin):
def fit(self, X, y=None, **fit_params):
return self
def transform(self, X, y=None, **fit_params):
return X.todense()
from sklearn.pipeline import Pipeline
from sklearn.naive_bayes import GaussianNB, MultinomialNB
spam_filter = Pipeline([
('vectorizer', TfidfVectorizer(analyzer=pre_process)),
('to_dense', DenseTransformer()),
('classifier', GaussianNB())
])
```
train test split
```
from sklearn.model_selection import train_test_split
x=df['message']
y=df['label']
x=x.values
y=y.values
x_train, x_test, y_train, y_test = train_test_split(x,y, test_size=0.20, random_state = 21) #Pareto principle
```
**Train spam_filter**
```
spam_filter.fit(x_train, y_train)
```
**Predict for test cases**
```
predictions = spam_filter.predict(x_test)
count = 0
for i in range(len(y_test)):
if y_test[i] != predictions[i]:
count += 1
print('Total number of test cases', len(y_test))
print('Number of wrong of predictions', count)
```
**Check for wrong predictions that were classified as ham**
```
#x_test[y_test != predictions]
```
**Use classification report to get more details**
```
from sklearn.metrics import classification_report
from sklearn.metrics import accuracy_score
print(classification_report(predictions, y_test))
accuracy_score(y_test, predictions)
```
Function to predict whether passed message is ham or spam
```
def detect_spam(s):
return spam_filter.predict([s])[0]
detect_spam('Your cash-balance is currently 500 pounds - to maximize your cash-in now, send COLLECT to 83600.')
from sklearn.metrics import confusion_matrix
confusion_matrix(predictions,y_test)
```
| github_jupyter |
# Setup
```
library(ggplot2)
library(cowplot)
library(ranger)
library(Metrics)
library(latex2exp)
library(reshape2)
library(akima)
library(pander)
```
# Generate Model
Following is an example of how to generate the prediction model using the Random Forest Model with AIWC metrics and experimental runtimes of the Extended Benchmark Suite.
```
load("./data/intermediate/full_dat.Rda")
#manually typecast integer variables
int.inds <- c(2,6,7,8,9,10,11,12,13,16,17,29,30,34,37)
#standardise numeric variables
num.inds <- lapply(full_dat, is.numeric)
num.inds <- as.logical(num.inds)
for (i in seq(along = num.inds)){
feature.name = names(full_dat[i])
ifelse((any(i == int.inds) || num.inds[i] == 'FALSE' || feature.name == "kernel_time"), next, full_dat[i] <- scale(full_dat[i]))
} #end i loop
#use 20% of data for training
sampled_indices <- sample(seq_len(nrow(full_dat)), size = round(nrow(full_dat)*0.2))
train_dat <- full_dat[sampled_indices, ]
test_dat <- full_dat[-sampled_indices, ]
#remove certain variables unavailable during real-world training
train_dat = subset(train_dat, select = -size)
train_dat = subset(train_dat, select = -application)
train_dat = subset(train_dat, select = -kernel)
train_dat = subset(train_dat, select = -total_time)
#build the model
rgd.aiwc <- ranger(log(kernel_time)~.,
data = train_dat,
num.trees = 505,
mtry = 30,
min.node.size = 9,
importance = "impurity",
splitrule = 'variance',
respect.unordered.factors = 'order')
#make the predictions
rgd.pred <- predict(rgd.aiwc,type='response',data=test_dat)
```
# Making Scheduler Predictions with the Model
This Section takes AIWC metrics and makes specific predictions.
A comparison to the experimental runtimes for the selected kernel, device and size is presented. Feel free to change the `k`, `d` and `s` variables to see changes and accuracies in the predictions. This is a prototype of what could be used in smart accelerator selections for heterogenous node scheduling. Any new data can be added by loading the new kernels AIWC metrics and selecting a device name.
```
print("Select the following to evaluate the model")
#query dataframe parameters
#names(test_dat)
#query unique kernels -- to make a prediction with the model
print("kernels:")
unique(test_dat$kernel)
#query unique devices
print("devices")
unique(test_dat$device)
k = "needle_opencl_shared_2"
d = "titanx"
s = "tiny"
y <- subset(test_dat,kernel==k&device==d&size==s)
# Note the exponent is used as the inverse to the natural logarithm which was used to add stability in the model.
p <- exp(mean(predict(rgd.aiwc,type='response',data=y)$predictions))
m <- mean(y$kernel_time)
print(paste("The mean predicted time (ms) of the ",s,k," on the ",d," is ",p," the measured time is ",m))
```
The remainder of the Jupyter Artefact shows how the figures and results were prepared for the [paper](https://www.researchgate.net/publication/328418090_OpenCL_Performance_Prediction_using_Architecture-Independent_Features).
# Constructing the Performance Model
The R programming language was used to analyse the data, construct the model and analyse the results. In particular, the ranger package by Wright and Ziegler was used for the development of the regression model. The ranger package provides computationally efficient implementations of the Random Forest model which performs recursive partitioning of high dimensional data.
The ranger function accepts three main parameters, each of which influences the fit of the model to the data. In optimizing the model, we searched over a range of values for each parameter including:
* num.trees, the number of trees grown in the random forest: over the range of 10 − 10, 000 by 500
* mtry, the number of features tried to possibly split within each node: ranges from 1 − 34, where 34 is the maximum number of input features available from AIWC,
* min.node.size, the minimal node size per tree: ranges from 1 − 50, where 50 is the number of observations per sample.
Given the size of the data set, it was not computationally viable to perform an exhaustive search of the entire 3-dimensional range of parameters. Auto-tuning to determine the suitability of these parameters has been performed by Ließ et al. to determine the optimal value of mtry given a fixed num.trees. Instead, to enable an efficient search of all variables at once, we used Flexible Global Optimization with Simulated-Annealing, in particular, the variant found in the R package optimization by Husmann, Lange and Spiegel. The simulated-annealing method both reduces the risk of getting trapped in a local minimum and is able to deal with irregular and complex parameter spaces as well as with non-continuous and sophisticated loss functions. In this setting, it is desirable to minimise the out-of-bag prediction error of the resultant fitted model, by simultaneously changing the parameters (num.trees, mtry and min.node.size). The optim_sa function allows defining the search space of interest, a starting position, the magnitude of the steps according to the relative change in temperature and the wrapper around the ranger function (which parses the 3 parameters and returns a cost function — the predicted error). It allows for an approximate global minimum to be detected with significantly fewer iterations than an exhaustive grid search.
## Full coverage of min.node.size with fixed tuning parameters: num.trees = 300 and mtry = 30.
```
#this was generated in ./analysis_tools/variation_in_min.node.size.R.
#and stored the results in "./data/intermediate/variation_in_min.node.size.Rdf" as the data.frame z
load('./data/intermediate/variation_in_min.node.size.Rdf')
z$y <- z$y * 100
tplot <- ggplot(dat=z,aes(x=x,y=y)) + geom_line() + labs(x="min.node.size",y="prediction error (%)") + ylim(0,max(z$y)) + theme(text = element_text(size=20),axis.text.x = element_text(size=15),axis.text.y= element_text(size=15))
print(tplot)
```
This Figure shows the relationship between out-of-bag predic- tion error and min.node.size, with the num.trees = 300 and mtry = 30 parameters fixed. In general, the min.node.size has the smallest prediction error for values less than 15 and variation in prediction error is similar throughout this range. As such, the selection to fix min.node.size = 9 was made to reduce the search-space in the remainder of the tuning work. We assume conditional (relative) independence between min.node.size and the other variables.
```
# this was generated by steering the starting positions of the optim_sa function to all corners of the search space and adding the log trace for each location, then randomly selecting 8 internal starting locations to sample internal detail.
x <- read.table('./data/intermediate/full_scan_random_sampled_heatmap.Rtable',header=TRUE, sep=" ")
y <- interp(x=x$mtry,
y=x$num.trees,
z=x$prediction.error*100,
xo=seq(1,34),
yo=seq(1,10000),
duplicate="mean")
d2 <- melt(y$z, na.rm = TRUE)
names(d2) <- c("mtry", "num.trees", "prediction.error")
ggplot(data=d2, aes(x=mtry,y=num.trees,fill=prediction.error)) + geom_tile(aes(fill=prediction.error)) + scale_fill_gradient(low="white", high="black") + theme(panel.grid=element_blank()) + scale_x_continuous(breaks = c(1, 5, 10, 15, 20, 25, 30, 34), expand = c(0,0)) + scale_y_continuous(breaks=c(1,2500,5000,7500,10000), expand=c(0,0)) + labs(fill="prediction\nerror (%)\n") + theme(text = element_text(size=20), axis.text.x = element_text(size=15), axis.text.y = element_text(size=15))
```
This Figure shows how the prediction error of the random- forest ranger model changes over a wide range of values for the two remaining tuning parameters, mtry and num.trees. Full coverage was achieved by selecting starting locations in each of the 4 outer-most points of the search space, along with 8 random internal points — to avoid missing out on some critical internal structure. For each combination of parameter values, the optim_sa function was allowed to execute until a global minimum was found. At each step of optimization a full trace was collected, where all parameters and the corresponding out-of-bag prediction error value were logged to a file. This file was finally loaded, the points interpolated using the R package akima, without extrapolation between points, using the mean values for duplication between points. The generated heatmap is shown in the Figure.
A lower out-of-bag prediction error is better. For values of mtry above 25, there is good model fit irrespective of the number of trees. For lower values of mtry, fit varies significantly with different values of num.trees. The worst fit was for a model with a value of 1 num.trees, and 1 for mtry, which had the highest out-of-bag prediction error at 194%. In general, the average prediction error across all choices of parameters is very low at 16%. Given these results, the final ranger model should use a small value for num.trees and a large value for mtry, with the added benefit that such a model can be computed faster given a smaller number of trees.
# Choosing Model Parameters
The selected model should be able to accurately predict execution times for a previously unseen kernel over the full range of accelerators. To show this, the model must not be over-fitted, that is to say, the random forest model parameters should not be tuned to the particular set of kernels in the training data, but should generate equally good fits if trained on any other reasonable selection of kernels.
We evaluated how robust the selection of model parameters is to the choice of kernel by repeatedly retraining the model on a set of kernels, each time removing a different kernel.
For each selection of kernels, optima_sa was run from the same starting location – num.trees=500, mtry=32 – and the final optimal values were recorded. min.node.size was fixed at 9.
The optimal – and final – parameters for each omitted kernel are presented in the Table.
```
# this was generated in ../analysis_tools/split_and_fit_study.R where each kernel is omitted one by one and the generated model is evaluated for both optimal tuning parameters using the optimization function and final r-squared and out-of-bag prediction error values
x <- read.table("./data/intermediate/tuning-variations.Rtable",header=TRUE,sep=' ')
# drop min.node.size results
x <- subset(x, select=-c(min.node.size,r.squared))
# make a percentage
x$prediction.error <- round(x$prediction.error * 100,1)
# split the really long value:
x$kernel.omitted <- as.character(x$kernel.omitted)
#x$kernel.omitted[x$kernel.omitted == "calc_potential_single_step_dev"] <- "calc_potential_single\n_step_dev"
x$kernel.omitted[x$kernel.omitted == "calc_potential_single_step_dev"] <- "calc_potential_single_step"
x$kernel.omitted <- as.factor(x$kernel.omitted)
panderOptions('table.split.table', Inf)
panderOptions('digits', 2)
set.caption('Optimal tuning parameters from the same starting location for all models omitting each individual kernel.\\label{tab:optimal-tuning-parameters}')
#names(x) <- c('Kernel omitted','num.trees','mtry','min.node.size','prediction error','R-squared')
names(x) <- c('Kernel omitted','num.trees','mtry','prediction error (%)')
pander(x)
```
# Prediction error across all benchmarks for models trained with varying numbers of kernels
```
#<!-- see ../analysis_tools/suitable_kernel_counts.R for implementation -->
results <- read.table('./data/intermediate/rmse_vs_kernel_count.Rtable',header=TRUE, sep=" ")
ggplot(dat=results,aes(x=number.of.kernels,y=mae)) + geom_line() + labs(x="number of kernels",y="mean absolute error") + ylim(0,max(results$mae)) + theme(text = element_text(size=20),axis.text.x = element_text(size=15),axis.text.y = element_text(size=15))
```
The results presented in the Figure show the mean absolute error of models trained on varying numbers of kernels. As expected, the model fit improves with increasing number of kernels. In particular, larger improvements occur with each new kernel early in the series and tapers off as a new kernel is added to an already large number of kernels. The gradient is still significant until the largest number of samples examined (k = 37) suggesting that the model could benefit from additional training data. However, the model proposed is a proof of concept and suggests that a general purpose model is attainable and may not require many more kernels.
# Evaluation -- Predicted vs. measured execution time for all kernels
```
#generated with ../analysis_tools/sample_generator.R
load("./data/intermediate/full_dat.Rda")
#manually typecast integer variables
int.inds <- c(2,6,7,8,9,10,11,12,13,16,17,29,30,34,37)
#standardise numeric variables
num.inds <- lapply(full_dat, is.numeric)
num.inds <- as.logical(num.inds)
for (i in seq(along = num.inds)){
feature.name = names(full_dat[i])
ifelse((any(i == int.inds) || num.inds[i] == 'FALSE' || feature.name == "kernel_time"), next, full_dat[i] <- scale(full_dat[i]))
} #end i loop
#use 20% of data for training
sampled_indices <- sample(seq_len(nrow(full_dat)), size = round(nrow(full_dat)*0.2))
train_dat <- full_dat[sampled_indices, ]
test_dat <- full_dat[-sampled_indices, ]
#remove certain variables unavailable during real-world training
train_dat = subset(train_dat, select = -size)
train_dat = subset(train_dat, select = -application)
train_dat = subset(train_dat, select = -kernel)
train_dat = subset(train_dat, select = -total_time)
#build the model
rgd.aiwc <- ranger(log(kernel_time)~.,
data = train_dat,
num.trees = 505,
mtry = 30,
min.node.size = 9,
importance = "impurity",
splitrule = 'variance',
respect.unordered.factors = 'order')
#make the predictions
rgd.pred <- predict(rgd.aiwc,type='response',data=test_dat)
#plot the error
z <- data.frame(predicted=rgd.pred$predictions,
actual=log(test_dat$kernel_time),
device=test_dat$device,
size=test_dat$size,
kernel=test_dat$kernel)
#plot the difference in log(times)
ggplot(data=z,aes(x=actual,y=predicted,colour=size)) + geom_point() + labs(x=TeX("log(measured $\\mu$s)"),y=TeX("log(predicted $\\mu$s)"))+ theme(text = element_text(size=20),axis.text.x = element_text(size=15),axis.text.y = element_text(size=15))
#plot the absolute difference in times -- but I'd prefer to show this in the next section
#ggplot(data=z,aes(x=exp(actual),y=exp(predicted),colour=size)) + geom_point() + labs(x="predicted (us)",y="measured (us)")
#back up the predictions for the remainder of the paper
oz <- z
```
This Figure presents the measured kernel execution times against the predicted execution times from the trained model. Each point represents a single combination of kernel and problem size. The plot shows a strong linear correlation indicating a good model fit. Under-predictions typically occur on four kernels over the medium and large problem sizes, while over-predictions occur on the tiny and small problem sizes. However, these outliers are visually over-represented in this figure as the final mean absolute error is low, at ~0.11.
# Making Predictions -- Error in predicted execution time for each kernel invocation over four problem sizes
```
construct_percent_matrix <- function(data){
x <- data
m <- length(unique(x$kernel))
n <- length(unique(x$device))
y <- matrix(nrow=m,ncol=n,dimnames = list(unique(x$kernel),unique(x$device)))
m_c <- 1
for (i in unique(x$kernel)){
n_c <- 1
for (j in unique(x$device)){
pred <- mean(subset(x,kernel==i&device==j)$predicted)
meas <- mean(subset(x,kernel==i&device==j)$actual)
y[m_c,n_c] <- abs(pred - meas) / meas * 100
n_c <- n_c + 1
}
m_c <- m_c + 1
}
return (y)
}
z <- oz
total_percent <- data.frame()
for(sz in c('tiny','small','medium','large')){
x <- subset(z,size==sz)
hm_z <- construct_percent_matrix(x)
my <- melt(hm_z)
total_percent <- rbind(total_percent,my)
#dirty hack to get plots the same size -- some label lengths are different so just get one label in the bottom plots to be as long
#library(stringi)
#max(stri_length(total_percent$Var1))
##30 is the longest string
my$Var1 <- as.character(my$Var1)
if(sz %in% c('tiny','medium'))
my$Var1[which(my$Var1=="needle_opencl_shared_2")] <- " needle_opencl_shared_2"
else
my$Var1[which(my$Var1=="needle_opencl_shared_2")] <- "needle_opencl_shared_2 "
my$Var1 <- as.factor(my$Var1)
assign(paste('my_',sz,sep=''),my)
}
for(sz in c('tiny','small','medium','large')){
my <- eval(parse(text=paste('my_',sz,sep='')))
p <- ggplot(data=my,aes(y=Var1,x=Var2,fill=value,label=value))
p <- p + geom_tile()
p <- p + scale_fill_continuous(high = "black", low = "white",limits=c(0, max(total_percent$value)))
p <- p + labs(x='device',y='kernel',fill=TeX('prediction error ($\\%$)'))
p <- p + theme(legend.position = "none",
axis.text.x = element_text(size = 8),
axis.text.y = element_text(size = 8,angle=45),
axis.title.x = element_text(size=10),
axis.title.y = element_text(size=10))
if(sz == 'tiny')
p <- p + scale_x_discrete(position = 'top') + theme(axis.text.x = element_text(angle = 45, hjust = 0, vjust=0))
if(sz == 'small')
p <- p + scale_y_discrete(position = 'right') + scale_x_discrete(position = 'top') + theme(axis.text.x = element_text(angle = 45, hjust = 0, vjust=0))
if(sz == 'medium')
p <- p + scale_y_discrete(position = 'left') + scale_x_discrete(position = 'bottom') + theme(axis.text.x = element_text(angle = 45, hjust = 1, vjust=1))
if(sz == 'large')
p <- p + scale_y_discrete(position = 'right') + scale_x_discrete(position = 'bottom') + theme(axis.text.x = element_text(angle = 45, hjust = 1, vjust=1))
assign(paste('stacked_',sz,sep=''),p)
}
legend_generic <- get_legend(stacked_tiny + theme(legend.title=element_text(face="bold",size=10),
legend.text=element_text(size=10),
legend.position="bottom",
legend.justification="right",
legend.direction="horizontal"))
plots <- align_plots(stacked_tiny ,
stacked_small ,
stacked_medium,
stacked_large)
#align="hv",axis="tblr")
xoff <- .22 # relative x position of label, within one plot
yoff <- .98 # relative y position of label, within one plot
x <- plot_grid(plot_grid(plots[[1]],plots[[2]],ncol=2,align="h")+draw_plot_label(label=c("Tiny", "Small"),
x=c(0.22,0.55),
y=rep(1-(1-yoff),2),
hjust=.5, vjust=.5,
size=14),
plot_grid(plots[[3]],plots[[4]],ncol=2,align="h")+draw_plot_label(label=c(" Medium", "Large"),
x=c(0.22,0.55),
y=c(0.985,0.985),
hjust=.5, vjust=.5,
size=14),
legend_generic,
rel_heights=c(1,1,.065),
nrow=3)
print(x)
print(paste("Mean prediction error:",mean(total_percent$value),"%"))
print(paste("Average prediction accuracy = ",100-mean(total_percent$value),"%"))
print(paste("min execution time mispredictions (us): ",round(min(full_dat$kernel_time)* round(mean(total_percent$value),1))))
print(paste("min execution time mispredictions (ms): ",round(max(full_dat$kernel_time)* round(mean(total_percent$value),1))*10**-3))
```
In this section, we examine differences in accuracy of predicted execution times between different kernels, which is of importance if the predictions are to be used in a scheduling setting.
The four heat maps presented in the Figure show the difference between mean predicted and measured kernel execution times as a percentage of the measured time. Thus, they depict the relative error in prediction – lighter indicates a smaller error. Four different problem sizes are presented: tiny in the top-left, small in the top-right, medium bottom-left, large bottom-right.
In general, we see highly accurate predictions which on average differ from the measured experimental run- times by 1.1%, which correspond to actual execution time mispredictions of 8 μs to 1 secs according to problem size.
The init_alpha_dev kernel is the worst predicted kernel over both the tiny and small problem sizes, with mean misprediction at 7.3%. However, this kernel is only run once per application run – it is used in the initialization of the Hidden Markov Model – and as such there are fewer response variables available for model training.
# The benefits of this approach -- Mean measured kernel execution times compared against mean predicted kernel execution times to perform a selection of kernels on large problem sizes across 15 accelerator devices.
```
#generate predictions
motivating_df <- data.frame()
#NOTE: be sure to only use kernels with results for the specified problem size
featured_kernels <- c('srad_cuda_2',
'kmeansPoint',
'lud_diagonal',
# 'fftRadix16Kernel',
'needle_opencl_shared_1')
featured_size <- c('large')
for(k in featured_kernels){
for(d in unique(full_dat$device)){
for(s in featured_size){
zx <- subset(test_dat, kernel==k & device == d & size == s)
rgd.pred <- predict(rgd.aiwc,type='response',data=zx)
zy <- subset(full_dat, kernel==k & device == d & size == s)
motivating_df <- rbind(motivating_df,
data.frame('kernel'=k,
'device'=d,
'size'=s,
'measured'=mean(zy$kernel_time)*0.001,
'predicted'=mean(exp(rgd.pred$predictions))*0.001))
}
}
}
square <- 22
diamond<- 23
p <- ggplot(motivating_df,aes(x=device,y=measured,colour=kernel,shape=NA)) +
geom_point(shape=square) +
geom_point(aes(x=device,y=predicted,colour=kernel),shape=diamond, size=3) +
theme(axis.text.x=element_text(angle=45,hjust=1)) +
scale_y_log10(breaks=c(0.0001, 0.10, 1, 10),labels=c("100 ns",TeX("100 $\\mu s$"),"1 ms", "10 ms")) +
labs(y="log(execution time)",x="device",colour="kernel",shape="type")
print(p)
```
To demonstrate the utility of the trained model to guide scheduling choices, we focus on the accuracy of performance time prediction of individual kernels over all devices. The model performance in terms of real execution times is presented for four randomly selected kernels in the Figure. The shape denotes the type of execution time data point, a square indicates the mean measured time, and the diamond indicates the mean predicted time. Thus, a perfect prediction occurs where the measured time – square – fits perfectly within the predicted – diamond – as seen in the legend.
The purpose of showing these results is to highlight the set- ting in which they could be used – on the supercomputing node. In this instance, it is expected a node to be composed of any combination of the 15 devices presented in the Figure 6. Thus, to be able to advise a scheduler which device to use to execute a kernel, the model must be able to correctly predict on which of a given pair of devices the kernel will run fastest. For any selected pair of devices, if the relative ordering of the measured and predicted execution times is different, the scheduler would choose the wrong device. In almost all cases, the relative order is preserved using our model. In other words, our model will correctly predict the fastest device in all cases – with one exception, the kmeansPoint kernel. For this kernel, the predicted time of the fiji-furyx is lower than the hawaii-r9-290x, however the measured times between the two shows the furyx completing the task in a shorter time. For all other device pairs, the relative order for the kmeansPoint kernel is correct. Additionally, the lud_diagonal kernel suffers from systematic under-prediction of execution times on AMD GPU devices, however the relative ordering is still correct. As such, the proposed model provides sufficiently accurate execution time predictions to be useful for scheduling to heterogeneous compute devices on supercomputers.
# Relative Performance to Gamma GLM
The remainder of the artefact is added to present a baseline comparison to the random forest model.
For this comparison we show the same usage with a Generalized Linear Model using gamma family variance and a log link function.
We show the poor predictive accuracy -- the high magnitude of error, and wide prediction intervals -- and the mean low quality prediction results presented alongside a prediction matrix.
```
load("./data/intermediate/full_dat.Rda")
#manually typecast integer variables
int.inds <- c(2,6,7,8,9,10,11,12,13,16,17,29,30,34,37)
#use 20% of data for training
sampled_indices <- sample(seq_len(nrow(full_dat)), size = round(nrow(full_dat)*0.2))
train_dat <- full_dat[sampled_indices, ]
test_dat <- full_dat[-sampled_indices, ]
#remove certain variables unavailable during real-world training
train_dat = subset(train_dat, select = -size)
train_dat = subset(train_dat, select = -application)
train_dat = subset(train_dat, select = -kernel)
train_dat = subset(train_dat, select = -total_time)
glm.aiwc <- glm(log(kernel_time) ~ . ,data=subset(train_dat,select=-c(invocation)),family=Gamma(link = "log"),control = list(maxit = 1000))
estimates <- predict(glm.aiwc,test_dat, type = 'response')
glm.pred <- estimates
#plot the error
z <- data.frame(predicted=glm.pred,
actual=log(test_dat$kernel_time),
device=test_dat$device,
size=test_dat$size,
kernel=test_dat$kernel)
#plot the difference in log(times)
ggplot(data=z,aes(x=actual,y=predicted,colour=size)) + geom_point() + labs(x=TeX("log(measured $\\mu$s)"),y=TeX("log(predicted $\\mu$s)"))+ theme(text = element_text(size=20),axis.text.x = element_text(size=15),axis.text.y = element_text(size=15))
for (i in seq(1,10)){
print(paste("predicted: ",estimates[i]," measured:", log(test_dat[i,]$kernel_time)))
}
```
We see poor prediction results -- for instance we see a much wider range of ground-truth values in the sample of measured execution times relatively to the averaged linear range in predicted kernel times.
### Distribution of Prediction Errors
```
actual = log(test_dat$kernel_time)
predicted = glm.pred
pred_error = actual - predicted
summary(exp(pred_error))
res = data.frame(
actual = actual,
predicted = predicted,
error = pred_error,
experror = exp(pred_error)
)
ggplot(data=res,aes(x=error)) + geom_density() + labs(x=TeX("Error Magnitude (log_{e})"),y=TeX("% of distribution"))
```
The model has a predictive accuracy of 84%, but mispredictions can be out by 483 micro-seconds.
### Prediction Intervals
The magnitude of the error in predictions is large, and it is difficult to understand how the factors impact on prediction error, for this reason, difficult to plot/visualise.
We now present the lower and upper prediction bounds by fixing all variables -- we selected the kmeansPoint kernel on the GTX1080Ti GPU -- and vary the problem size.
In practice any other variable could be fixed and varied, but this is only done to show the large uncertainity of the prediction intervals across problem size and that the GLM is a poor choice of model.
#### Subset the data
```
# Plot marginals for fixing all but one variable and choose a few levels.
marginal_test_dat <- subset(test_dat, kernel == "kmeansPoint" & device == "gtx1080ti")
```
#### Generate the predictions and the upper and lower prediction intervals
```
glm.aiwc <- glm(log(kernel_time) ~ . ,data=subset(train_dat,select=-c(invocation)),family=Gamma(link = "log"),control = list(maxit = 100))
glm.pred <- predict(glm.aiwc,newdata = marginal_test_dat,type="response")
piv <- predict(glm.aiwc,newdata = marginal_test_dat, interval="prediction", level=.95, se.fit=TRUE)
z <- data.frame(predicted=glm.pred,
actual=marginal_test_dat$kernel_time,
device=marginal_test_dat$device,
size=marginal_test_dat$size,
kernel=marginal_test_dat$kernel,
lpb=(piv$fit-piv$residual.scale*piv$se.fit),
upb=(piv$fit+piv$residual.scale*piv$se.fit))
```
#### Plot the predictions with upper and lower prediction intervals
```
ggplot(data=z,aes(x=size,y=predicted)) + geom_point() +
labs(x=TeX("problem size"),y=TeX("log predicted times ($\\mu$s)")) +
theme(text = element_text(size=20),axis.text.x = element_text(size=15),axis.text.y = element_text(size=15))+
geom_errorbar(aes(ymin = predicted-lpb, ymax = predicted+upb),) + expand_limits(y = 1)
```
## Predictions with GLM -- Error in predicted execution time for each kernel invocation over four problem sizes
This section shows the average prediction accuracy if GLM were used as the proposed model -- instead of the random forest.
#### Generate the predictions
```
glm.aiwc <- glm(log(kernel_time) ~ . ,data=subset(train_dat,select=-c(invocation)),family=Gamma(link = "log"),control = list(maxit = 100))
glm.pred <- predict(glm.aiwc,newdata = test_dat,type="response")
z <- data.frame(predicted=glm.pred,
actual=log(test_dat$kernel_time),
device=test_dat$device,
size=test_dat$size,
kernel=test_dat$kernel)
construct_percent_matrix <- function(data){
x <- data
m <- length(unique(x$kernel))
n <- length(unique(x$device))
y <- matrix(nrow=m,ncol=n,dimnames = list(unique(x$kernel),unique(x$device)))
m_c <- 1
for (i in unique(x$kernel)){
n_c <- 1
for (j in unique(x$device)){
pred <- mean(subset(x,kernel==i&device==j)$predicted)
meas <- mean(subset(x,kernel==i&device==j)$actual)
y[m_c,n_c] <- abs(pred - meas) / meas * 100
n_c <- n_c + 1
}
m_c <- m_c + 1
}
return (y)
}
total_percent <- data.frame()
for(sz in c('tiny','small','medium','large')){
x <- subset(z,size==sz)
hm_z <- construct_percent_matrix(x)
my <- melt(hm_z)
total_percent <- rbind(total_percent,my)
#dirty hack to get plots the same size -- some label lengths are different so just get one label in the bottom plots to be as long
#library(stringi)
#max(stri_length(total_percent$Var1))
##30 is the longest string
my$Var1 <- as.character(my$Var1)
if(sz %in% c('tiny','medium'))
my$Var1[which(my$Var1=="needle_opencl_shared_2")] <- " needle_opencl_shared_2"
else
my$Var1[which(my$Var1=="needle_opencl_shared_2")] <- "needle_opencl_shared_2 "
my$Var1 <- as.factor(my$Var1)
assign(paste('my_',sz,sep=''),my)
}
for(sz in c('tiny','small','medium','large')){
my <- eval(parse(text=paste('my_',sz,sep='')))
p <- ggplot(data=my,aes(y=Var1,x=Var2,fill=value,label=value))
p <- p + geom_tile()
p <- p + scale_fill_continuous(high = "black", low = "white")
p <- p + labs(x='device',y='kernel',fill=TeX('prediction error ($\\%$)'))
p <- p + theme(legend.position = "none",
axis.text.x = element_text(size = 8),
axis.text.y = element_text(size = 8,angle=45),
axis.title.x = element_text(size=10),
axis.title.y = element_text(size=10))
if(sz == 'tiny')
p <- p + scale_x_discrete(position = 'top') + theme(axis.text.x = element_text(angle = 45, hjust = 0, vjust=0))
if(sz == 'small')
p <- p + scale_y_discrete(position = 'right') + scale_x_discrete(position = 'top') + theme(axis.text.x = element_text(angle = 45, hjust = 0, vjust=0))
if(sz == 'medium')
p <- p + scale_y_discrete(position = 'left') + scale_x_discrete(position = 'bottom') + theme(axis.text.x = element_text(angle = 45, hjust = 1, vjust=1))
if(sz == 'large')
p <- p + scale_y_discrete(position = 'right') + scale_x_discrete(position = 'bottom') + theme(axis.text.x = element_text(angle = 45, hjust = 1, vjust=1))
assign(paste('stacked_',sz,sep=''),p)
}
legend_generic <- get_legend(stacked_tiny + theme(legend.title=element_text(face="bold",size=10),
legend.text=element_text(size=10),
legend.position="bottom",
legend.justification="right",
legend.direction="horizontal"))
plots <- align_plots(stacked_tiny ,
stacked_small ,
stacked_medium,
stacked_large)
#align="hv",axis="tblr")
xoff <- .22 # relative x position of label, within one plot
yoff <- .98 # relative y position of label, within one plot
x <- plot_grid(plot_grid(plots[[1]],plots[[2]],ncol=2,align="h")+draw_plot_label(label=c("Tiny", "Small"),
x=c(0.22,0.55),
y=rep(1-(1-yoff),2),
hjust=.5, vjust=.5,
size=14),
plot_grid(plots[[3]],plots[[4]],ncol=2,align="h")+draw_plot_label(label=c(" Medium", "Large"),
x=c(0.22,0.55),
y=c(0.985,0.985),
hjust=.5, vjust=.5,
size=14),
legend_generic,
rel_heights=c(1,1,.065),
nrow=3)
options(repr.plot.width=8, repr.plot.height=16)
print(x)
print(paste("Mean prediction error:",mean(total_percent$value),"%"))
print(paste("Average prediction accuracy = ",100-mean(total_percent$value),"%"))
print(paste("min execution time mispredictions (us): ",round(min(full_dat$kernel_time)* round(mean(total_percent$value),1))))
print(paste("min execution time mispredictions (s): ",round(max(full_dat$kernel_time)* round(mean(total_percent$value),1))*10**-6))
```
We initially fitted the GLM, it was not included in the original paper due to the poor prediction results — the mean error of prediction times was 2.9080 micro-seconds but this error could be much greater and up to 451.102 microseconds. This level of error was unacceptable and it is suggested that the GLM was unable to capture the complicated statistical relationships between AIWC features and execution time.
This motivated the use of non-parametric models — such as the random forest — which we have demonstrated is more suitable for the data.
| github_jupyter |
Note! For a most up to date version of this notebook, make sure you copy from:
[](https://colab.research.google.com/drive/1wTMIrJhYsQdq_u7ROOkf0Lu_fsX5Mu8a)
## Configs and Hyperparameters
Support a variety of models, you can find more pretrained model from [Tensorflow detection model zoo: COCO-trained models](https://github.com/tensorflow/models/blob/master/research/object_detection/g3doc/detection_model_zoo.md#coco-trained-models), as well as their pipline config files in [object_detection/samples/configs/](https://github.com/tensorflow/models/tree/master/research/object_detection/samples/configs).
```
# If you forked the repo, you can replace the link.
repo_url = 'https://github.com/roboflow-ai/tensorflow-object-detection-faster-rcnn'
# Number of training steps - 1000 will train very quickly, but more steps will increase accuracy.
num_steps = 10000 # 200000 to improve
# Number of evaluation steps.
num_eval_steps = 50
MODELS_CONFIG = {
'ssd_mobilenet_v2': {
'model_name': 'ssd_mobilenet_v2_coco_2018_03_29',
'pipeline_file': 'ssd_mobilenet_v2_coco.config',
'batch_size': 12
},
'faster_rcnn_inception_v2': {
'model_name': 'faster_rcnn_inception_v2_coco_2018_01_28',
'pipeline_file': 'faster_rcnn_inception_v2_pets.config',
'batch_size': 12
},
'rfcn_resnet101': {
'model_name': 'rfcn_resnet101_coco_2018_01_28',
'pipeline_file': 'rfcn_resnet101_pets.config',
'batch_size': 8
},
}
# Pick the model you want to use
# Select a model in `MODELS_CONFIG`.
selected_model = 'ssd_mobilenet_v2'
# Name of the object detection model to use.
MODEL = MODELS_CONFIG[selected_model]['model_name']
# Name of the pipline file in tensorflow object detection API.
pipeline_file = MODELS_CONFIG[selected_model]['pipeline_file']
# Training batch size fits in Colabe's Tesla K80 GPU memory for selected model.
batch_size = MODELS_CONFIG[selected_model]['batch_size']
# use TF 1.x for Object Detection APIs as they are not ported to TF 2.0 yet
%tensorflow_version 1.x
```
## Clone the `tensorflow-object-detection` repository or your fork.
```
import os
%cd /content
repo_dir_path = os.path.abspath(os.path.join('.', os.path.basename(repo_url)))
!git clone {repo_url}
%cd {repo_dir_path}
!git pull
```
## Install required packages
```
%cd /content
!git clone --quiet https://github.com/tensorflow/models.git
!pip install tf_slim
!apt-get install -qq protobuf-compiler python-pil python-lxml python-tk
!pip install -q Cython contextlib2 pillow lxml matplotlib
!pip install -q pycocotools
%cd /content/models/research
!protoc object_detection/protos/*.proto --python_out=.
import os
os.environ['PYTHONPATH'] += ':/content/models/research/:/content/models/research/slim/'
!python object_detection/builders/model_builder_test.py
```
## Prepare `tfrecord` files
Roboflow automatically creates our TFRecord and label_map files that we need!
**Generating your own TFRecords the only step you need to change for your own custom dataset.**
Because we need one TFRecord file for our training data, and one TFRecord file for our test data, we'll create two separate datasets in Roboflow and generate one set of TFRecords for each.
To create a dataset in Roboflow and generate TFRecords, follow [this step-by-step guide](https://blog.roboflow.ai/getting-started-with-roboflow/).
```
%cd /content/tensorflow-object-detection-faster-rcnn/data
!curl -L "https://app.roboflow.com/ds/robins" > roboflow.zip; unzip roboflow.zip; rm roboflow.zip
# training set
%ls train
# test set
%ls test
# NOTE: Update these TFRecord names from "cells" and "cells_label_map" to your files!
test_record_fname = '/content/tensorflow-object-detection-faster-rcnn/data/test/fire.tfrecord'
train_record_fname = '/content/tensorflow-object-detection-faster-rcnn/data/train/fire.tfrecord'
label_map_pbtxt_fname = '/content/tensorflow-object-detection-faster-rcnn/data/train/fire_label_map.pbtxt'
```
## Download base model
```
%cd /content/models/research
import os
import shutil
import glob
import urllib.request
import tarfile
MODEL_FILE = MODEL + '.tar.gz'
DOWNLOAD_BASE = 'http://download.tensorflow.org/models/object_detection/'
DEST_DIR = '/content/models/research/pretrained_model'
if not (os.path.exists(MODEL_FILE)):
urllib.request.urlretrieve(DOWNLOAD_BASE + MODEL_FILE, MODEL_FILE)
tar = tarfile.open(MODEL_FILE)
tar.extractall()
tar.close()
os.remove(MODEL_FILE)
if (os.path.exists(DEST_DIR)):
shutil.rmtree(DEST_DIR)
os.rename(MODEL, DEST_DIR)
!echo {DEST_DIR}
!ls -alh {DEST_DIR}
fine_tune_checkpoint = os.path.join(DEST_DIR, "model.ckpt")
fine_tune_checkpoint
```
## Configuring a Training Pipeline
```
import os
pipeline_fname = os.path.join('/content/models/research/object_detection/samples/configs/', pipeline_file)
assert os.path.isfile(pipeline_fname), '`{}` not exist'.format(pipeline_fname)
def get_num_classes(pbtxt_fname):
from object_detection.utils import label_map_util
label_map = label_map_util.load_labelmap(pbtxt_fname)
categories = label_map_util.convert_label_map_to_categories(
label_map, max_num_classes=90, use_display_name=True)
category_index = label_map_util.create_category_index(categories)
return len(category_index.keys())
import re
num_classes = get_num_classes(label_map_pbtxt_fname)
with open(pipeline_fname) as f:
s = f.read()
with open(pipeline_fname, 'w') as f:
# fine_tune_checkpoint
s = re.sub('fine_tune_checkpoint: ".*?"',
'fine_tune_checkpoint: "{}"'.format(fine_tune_checkpoint), s)
# tfrecord files train and test.
s = re.sub(
'(input_path: ".*?)(train.record)(.*?")', 'input_path: "{}"'.format(train_record_fname), s)
s = re.sub(
'(input_path: ".*?)(val.record)(.*?")', 'input_path: "{}"'.format(test_record_fname), s)
# label_map_path
s = re.sub(
'label_map_path: ".*?"', 'label_map_path: "{}"'.format(label_map_pbtxt_fname), s)
# Set training batch_size.
s = re.sub('batch_size: [0-9]+',
'batch_size: {}'.format(batch_size), s)
# Set training steps, num_steps
s = re.sub('num_steps: [0-9]+',
'num_steps: {}'.format(num_steps), s)
# Set number of classes num_classes.
s = re.sub('num_classes: [0-9]+',
'num_classes: {}'.format(num_classes), s)
f.write(s)
!cat {pipeline_fname}
model_dir = 'training/'
# Optionally remove content in output model directory to fresh start.
!rm -rf {model_dir}
os.makedirs(model_dir, exist_ok=True)
```
## Run Tensorboard(Optional)
```
!wget https://bin.equinox.io/c/4VmDzA7iaHb/ngrok-stable-linux-amd64.zip
!unzip -o ngrok-stable-linux-amd64.zip
LOG_DIR = model_dir
get_ipython().system_raw(
'tensorboard --logdir {} --host 0.0.0.0 --port 6006 &'
.format(LOG_DIR)
)
get_ipython().system_raw('./ngrok http 6006 &')
```
### Get Tensorboard link
```
! curl -s http://localhost:4040/api/tunnels | python3 -c \
"import sys, json; print(json.load(sys.stdin)['tunnels'][0]['public_url'])"
```
## Train the model
```
!python /content/models/research/object_detection/model_main.py \
--pipeline_config_path={pipeline_fname} \
--model_dir={model_dir} \
--alsologtostderr \
--num_train_steps={num_steps} \
--num_eval_steps={num_eval_steps}
!ls {model_dir}
```
## Exporting a Trained Inference Graph
Once your training job is complete, you need to extract the newly trained inference graph, which will be later used to perform the object detection. This can be done as follows:
```
import re
import numpy as np
output_directory = './fine_tuned_model'
lst = os.listdir(model_dir)
lst = [l for l in lst if 'model.ckpt-' in l and '.meta' in l]
steps=np.array([int(re.findall('\d+', l)[0]) for l in lst])
last_model = lst[steps.argmax()].replace('.meta', '')
last_model_path = os.path.join(model_dir, last_model)
print(last_model_path)
!python /content/models/research/object_detection/export_inference_graph.py \
--input_type=image_tensor \
--pipeline_config_path={pipeline_fname} \
--output_directory={output_directory} \
--trained_checkpoint_prefix={last_model_path}
!ls {output_directory}
```
## Download the model `.pb` file
```
import os
pb_fname = os.path.join(os.path.abspath(output_directory), "frozen_inference_graph.pb")
assert os.path.isfile(pb_fname), '`{}` not exist'.format(pb_fname)
!ls -alh {pb_fname}
```
### Option2 : Download the `.pb` file directly to your local file system
This method may not be stable when downloading large files like the model `.pb` file. Try **option 1** instead if not working.
```
from google.colab import files
files.download(pb_fname)
```
### OPTIONAL: Download the `label_map.pbtxt` file
```
from google.colab import files
files.download(label_map_pbtxt_fname)
```
### OPTIONAL: Download the modified pipline file
If you plan to use OpenVINO toolkit to convert the `.pb` file to inference faster on Intel's hardware (CPU/GPU, Movidius, etc.)
```
files.download(pipeline_fname)
# !tar cfz fine_tuned_model.tar.gz fine_tuned_model
# from google.colab import files
# files.download('fine_tuned_model.tar.gz')
%cd /content
ls
```
upload test image via ui
## Run inference test
Test with images in repository `tensorflow-object-detection/test` directory.
**To test with your own images, you need to place your images inside the `test` directory in this Colab notebook!** More on this below.
```
import os
import glob
# Path to frozen detection graph. This is the actual model that is used for the object detection.
PATH_TO_CKPT = pb_fname
# List of the strings that is used to add correct label for each box.
PATH_TO_LABELS = label_map_pbtxt_fname
# If you want to test the code with your images, just add images files to the PATH_TO_TEST_IMAGES_DIR.
PATH_TO_TEST_IMAGES_DIR = os.path.join(repo_dir_path, "test")
assert os.path.isfile(pb_fname)
assert os.path.isfile(PATH_TO_LABELS)
TEST_IMAGE_PATHS = '/content/pan-fire.jpg'
assert len(TEST_IMAGE_PATHS) > 0, 'No image found in `{}`.'.format(PATH_TO_TEST_IMAGES_DIR)
print(TEST_IMAGE_PATHS)
%cd /content/models/research/object_detection
import numpy as np
import os
import six.moves.urllib as urllib
import sys
import tarfile
import tensorflow as tf
import zipfile
from collections import defaultdict
from io import StringIO
from matplotlib import pyplot as plt
from PIL import Image
# This is needed since the notebook is stored in the object_detection folder.
sys.path.append("..")
from object_detection.utils import ops as utils_ops
# This is needed to display the images.
%matplotlib inline
from object_detection.utils import label_map_util
from object_detection.utils import visualization_utils as vis_util
detection_graph = tf.Graph()
with detection_graph.as_default():
od_graph_def = tf.GraphDef()
with tf.gfile.GFile(PATH_TO_CKPT, 'rb') as fid:
serialized_graph = fid.read()
od_graph_def.ParseFromString(serialized_graph)
tf.import_graph_def(od_graph_def, name='')
label_map = label_map_util.load_labelmap(PATH_TO_LABELS)
categories = label_map_util.convert_label_map_to_categories(
label_map, max_num_classes=num_classes, use_display_name=True)
category_index = label_map_util.create_category_index(categories)
def load_image_into_numpy_array(image):
(im_width, im_height) = image.size
return np.array(image.getdata()).reshape(
(im_height, im_width, 3)).astype(np.uint8)
# Size, in inches, of the output images.
IMAGE_SIZE = (12, 8)
def run_inference_for_single_image(image, graph):
with graph.as_default():
with tf.Session() as sess:
# Get handles to input and output tensors
ops = tf.get_default_graph().get_operations()
all_tensor_names = {
output.name for op in ops for output in op.outputs}
tensor_dict = {}
for key in [
'num_detections', 'detection_boxes', 'detection_scores',
'detection_classes', 'detection_masks'
]:
tensor_name = key + ':0'
if tensor_name in all_tensor_names:
tensor_dict[key] = tf.get_default_graph().get_tensor_by_name(
tensor_name)
if 'detection_masks' in tensor_dict:
# The following processing is only for single image
detection_boxes = tf.squeeze(
tensor_dict['detection_boxes'], [0])
detection_masks = tf.squeeze(
tensor_dict['detection_masks'], [0])
# Reframe is required to translate mask from box coordinates to image coordinates and fit the image size.
real_num_detection = tf.cast(
tensor_dict['num_detections'][0], tf.int32)
detection_boxes = tf.slice(detection_boxes, [0, 0], [
real_num_detection, -1])
detection_masks = tf.slice(detection_masks, [0, 0, 0], [
real_num_detection, -1, -1])
detection_masks_reframed = utils_ops.reframe_box_masks_to_image_masks(
detection_masks, detection_boxes, image.shape[0], image.shape[1])
detection_masks_reframed = tf.cast(
tf.greater(detection_masks_reframed, 0.5), tf.uint8)
# Follow the convention by adding back the batch dimension
tensor_dict['detection_masks'] = tf.expand_dims(
detection_masks_reframed, 0)
image_tensor = tf.get_default_graph().get_tensor_by_name('image_tensor:0')
# Run inference
output_dict = sess.run(tensor_dict,
feed_dict={image_tensor: np.expand_dims(image, 0)})
# all outputs are float32 numpy arrays, so convert types as appropriate
output_dict['num_detections'] = int(
output_dict['num_detections'][0])
output_dict['detection_classes'] = output_dict[
'detection_classes'][0].astype(np.uint8)
output_dict['detection_boxes'] = output_dict['detection_boxes'][0]
output_dict['detection_scores'] = output_dict['detection_scores'][0]
if 'detection_masks' in output_dict:
output_dict['detection_masks'] = output_dict['detection_masks'][0]
return output_dict
# Output images not showing? Run this cell again, and try the cell above
# This is needed to display the images.
%matplotlib inline
image_path = TEST_IMAGE_PATHS
image = Image.open(image_path)
# the array based representation of the image will be used later in order to prepare the
# result image with boxes and labels on it.
image_np = load_image_into_numpy_array(image)
# Expand dimensions since the model expects images to have shape: [1, None, None, 3]
image_np_expanded = np.expand_dims(image_np, axis=0)
# Actual detection.
output_dict = run_inference_for_single_image(image_np, detection_graph)
# Visualization of the results of a detection.
vis_util.visualize_boxes_and_labels_on_image_array(
image_np,
output_dict['detection_boxes'],
output_dict['detection_classes'],
output_dict['detection_scores'],
category_index,
instance_masks=output_dict.get('detection_masks'),
use_normalized_coordinates=True,
line_thickness=8)
plt.figure(figsize=IMAGE_SIZE)
plt.imshow(image_np)
### Adding your own images to tensorflow-object-detection/data
def upload_files():
from google.colab import files
uploaded = files.upload()
for k, v in uploaded.items():
open(k, 'wb').write(v)
return list(uploaded.keys())
# navigate to correct folder
%cd /content/tensorflow-object-detection/test
# call function to upload
upload_files()
```
| github_jupyter |
# Attentional Networks in Computer Vision
Prepared by Comp411 Teaching Unit (TA Can Küçüksözen) in the context of Computer Vision with Deep Learning Course. Do not hesitate to ask in case you have any questions, contact me at: ckucuksozen19@ku.edu.tr
Up until this point, we have worked with deep fully-connected networks, convolutional networks and recurrent networks using them to explore different optimization strategies and network architectures. Fully-connected networks are a good testbed for experimentation because they are very computationally efficient, on the other hand, most successful image processing methods use convolutional networks. However recent state-of-the-art results on computer vision realm are acquired using Attentional layers and Transformer architectures.
First you will implement several layer types that are used in fully attentional networks. You will then use these layers to train an Attentional Image Classification network, specifically a smaller version of Vision Transformer (VIT) on the CIFAR-10 dataset. The original paper can be accessed via the following link: https://arxiv.org/pdf/2010.11929.pdf
# Part I. Preparation
First, we load the CIFAR-10 dataset. This might take a couple minutes the first time you do it, but the files should stay cached after that.
In previous parts of the assignment we had to write our own code to download the CIFAR-10 dataset, preprocess it, and iterate through it in minibatches; PyTorch provides convenient tools to automate this process for us.
```
import torch
import torch.nn as nn
import torch.optim as optim
from torch.utils.data import DataLoader
from torch.utils.data import sampler
import torch.nn.functional as F
from torch.autograd import Variable
import torchvision.datasets as dset
import torchvision.transforms as T
import os
os.environ['KMP_DUPLICATE_LIB_OK']='True'
import numpy as np
NUM_TRAIN = 49000
# The torchvision.transforms package provides tools for preprocessing data
# and for performing data augmentation; here we set up a transform to
# preprocess the data by subtracting the mean RGB value and dividing by the
# standard deviation of each RGB value; we've hardcoded the mean and std.
transform = T.Compose([
T.ToTensor(),
T.Normalize((0.4914, 0.4822, 0.4465), (0.2023, 0.1994, 0.2010))
])
# We set up a Dataset object for each split (train / val / test); Datasets load
# training examples one at a time, so we wrap each Dataset in a DataLoader which
# iterates through the Dataset and forms minibatches. We divide the CIFAR-10
# training set into train and val sets by passing a Sampler object to the
# DataLoader telling how it should sample from the underlying Dataset.
cifar10_train = dset.CIFAR10('./comp411/datasets', train=True, download=True,
transform=transform)
loader_train = DataLoader(cifar10_train, batch_size=64,
sampler=sampler.SubsetRandomSampler(range(NUM_TRAIN)))
cifar10_val = dset.CIFAR10('./comp411/datasets', train=True, download=True,
transform=transform)
loader_val = DataLoader(cifar10_val, batch_size=64,
sampler=sampler.SubsetRandomSampler(range(NUM_TRAIN, 50000)))
cifar10_test = dset.CIFAR10('./comp411/datasets', train=False, download=True,
transform=transform)
loader_test = DataLoader(cifar10_test, batch_size=64)
```
You have an option to **use GPU by setting the flag to True below**. It is not necessary to use GPU for this assignment. Note that if your computer does not have CUDA enabled, `torch.cuda.is_available()` will return False and this notebook will fallback to CPU mode.
The global variables `dtype` and `device` will control the data types throughout this assignment.
```
USE_GPU = True
dtype = torch.float32 # we will be using float throughout this tutorial
if USE_GPU and torch.cuda.is_available():
device = torch.device('cuda')
else:
device = torch.device('cpu')
# Constant to control how frequently we print train loss
print_every = 100
print('using device:', device)
```
# Part II. Barebones Transformers: Self-Attentional Layer
Here you will complete the implementation of the Pytorch nn.module `SelfAttention`, which will perform the forward pass of a self-attentional layer. Our implementation of the SelfAttentional layer will include three distinct fully connected layers which will be responsible of:
1. A fully connected layer, `W_Q`, which will be used to project our input into `queries`
2. A fully connected layer, `W_K`, which will be used to project our input into `keys`
3. A fully connected layer, `W_V`, which will be used to project our input into `values`
After defining such three fully connected layers, and obtain our `queries, keys, and values` variables at the beginning of our forward pass, the following operations should be carried out in order to complete the attentional layer implementation.
1. Seperate each of `query, key, and value` projections into their respective heads. In other words, split the feature vector dimension of each matrix into necessarry number of chunks.
2. Compute the `Attention Scores` between each pair of sequence elements via conducting a scaled dot product operation between every pair of `queries` and `keys`. Note that `Attention Scores` matrix should have the size of `[# of queries , # of keys]`
3. Calculate the `Attention Weights` of each query by applying the non-linear `Softmax` normalization accross the `keys` dimension of the `Attention Scores` matrix.
4. Obtain the output combination of `values` by matrix multiplying `Attention Weights` with `values`
5. Reassemble heads into one flat vector and return the output.
**HINT**: For a more detailed explanation of the self attentional layer, examine the Appendix A of the original ViT manuscript here: https://arxiv.org/pdf/2010.11929.pdf
```
class SelfAttention(nn.Module):
def __init__(self, input_dims, head_dims=128, num_heads=2, bias=False):
super(SelfAttention, self).__init__()
## initialize module's instance variables
self.input_dims = input_dims
self.head_dims = head_dims
self.num_heads = num_heads
self.proj_dims = head_dims * num_heads
## Declare module's parameters
self.W_Q = nn.Linear(input_dims, self.proj_dims,bias=bias)
self.W_K = nn.Linear(input_dims, self.proj_dims,bias=bias)
self.W_V = nn.Linear(input_dims, self.proj_dims,bias=bias)
self.W_O = nn.Linear(self.proj_dims,self.proj_dims,bias=bias)
self.init_weights()
def init_weights(self):
for m in self.modules():
if isinstance(m, nn.Linear):
nn.init.xavier_normal_(m.weight.data)
if m.bias is not None:
m.bias.data.fill_(0.1)
def forward(self, x):
## Input of shape, [B, N, D] where:
## - B denotes the batch size
## - N denotes number of sequence elements. I.e. the number of patches + the class token
## - D corresponds to model dimensionality
b,n,d = x.shape
## Construct queries,keys,values
q_ = self.W_Q(x)
k_ = self.W_K(x)
v_ = self.W_V(x)
## Seperate q,k,v into their corresponding heads,
## After this operation each q,k,v will have the shape: [B,H,N,D//H] where
## - B denotes the batch size
## - H denotes number of heads
## - N denotes number of sequence elements. I.e. the number of patches + the class token
## - D//H corresponds to per head dimensionality
q, k, v = map(lambda z: torch.reshape(z, (b,n,self.num_heads,self.head_dims)).permute(0,2,1,3), [q_,k_,v_])
attn_out = None
#########################################################################################
# TODO: Complete the forward pass of the SelfAttention layer, follow the comments below #
#########################################################################################
# *****START OF YOUR CODE (DO NOT DELETE/MODIFY THIS LINE)*****
## Compute attention logits. Note that this operation is implemented as a
## batched matrix multiplication between q and k, the output is scaled by 1/(D//H)^(1/2)
## inputs are queries and keys that are both of size [B,H,N,D//H]
## Output Attention logits should have the size: [B,H,N,N]
intermdeiary = torch.matmul(q,k.transpose(-2,-1)) * (1/((d//self.num_heads)**(1/2)))
## Compute attention Weights. Recall that this operation is conducted as a
## Softmax Normalization across the keys dimension.
## Hint: You can apply the Softmax operation across the final dimension
# attn_weights = nn.softmax(intermediary,dim=-1)
attn_weights = intermdeiary.softmax(dim=-1)
## Compute attention output values. Bear in mind that this operation is applied as a
## batched matrix multiplication between the Attention Weights matrix and
## the values tensor. After computing output values, the output should be reshaped
## Inputs are Attention Weights with size [B, H, N, N], values with size [B, H, N, D//H]
## Output should be of size [B, N, D]
## Hint: you should use torch.matmul, torch.permute, torch.reshape in that order
## (or any other equivalent torch operations)
scores = torch.matmul(attn_weights,v)
out = scores.reshape(b,n,self.proj_dims)
## Compute output feature map. This operation is just passing the concatenated attention
## output that we have just obtained through a final projection layer W_O.
## Both the input and the output should be of size [B, N, D]
attn_out = self.W_O(out)
# *****END OF YOUR CODE (DO NOT DELETE/MODIFY THIS LINE)*****
################################################################################
# END OF YOUR CODE
################################################################################
return attn_out
```
After defining the forward pass of the Self-Attentional Layer above, run the following cell to test your implementation.
When you run this function, output should have shape (64, 16, 64).
```
def test_self_attn_layer():
x = torch.zeros((64, 16, 32), dtype=dtype) # minibatch size 64, sequence elements size 16, feature channels size 32
layer = SelfAttention(32,64,4)
out = layer(x)
print(out.size()) # you should see [64,16,256]
test_self_attn_layer()
```
# Part III. Barebones Transformers: Transformer Encoder Block
Here you will complete the implementation of the Pytorch nn.module `TransformerBlock`, which will perform the forward pass of a Transfomer Encoder Block. You can refer to Figure 1 of the original manuscript of ViT from this link: https://arxiv.org/pdf/2010.11929.pdf in order to get yourself familiar with the architecture.
```
## Implementation of a two layer GELU activated Fully Connected Network is provided for you below:
class MLP(nn.Module):
def __init__(self, input_dims, hidden_dims, output_dims, bias=True):
super().__init__()
self.fc_1 = nn.Linear(input_dims, hidden_dims, bias=bias)
self.fc_2 = nn.Linear(hidden_dims, output_dims, bias=bias)
self.init_weights()
def init_weights(self):
for m in self.modules():
if isinstance(m, nn.Linear):
nn.init.xavier_normal_(m.weight.data)
if m.bias is not None:
m.bias.data.fill_(0.1)
def forward(self, x):
o = F.gelu(self.fc_1(x))
o = self.fc_2(o)
return o
## Build from scratch a TransformerBlock Module. Note that the architecture of this
## module follows a simple computational pipeline:
## input --> layernorm --> SelfAttention --> skip connection
## --> layernorm --> MLP ---> skip connection ---> output
## Note that the TransformerBlock module works on a single hidden dimension hidden_dims,
## in order to faciliate skip connections with ease. Be careful about the input arguments
## to the SelfAttention block.
class TransformerBlock(nn.Module):
def __init__(self, hidden_dims, num_heads=4, bias=False):
super(TransformerBlock, self).__init__()
###############################################################
# TODO: Complete the consturctor of TransformerBlock module #
###############################################################
# *****START OF YOUR CODE (DO NOT DELETE/MODIFY THIS LINE)****
self.layerNorm_1 = nn.LayerNorm(hidden_dims)
self.selfAttention = SelfAttention(hidden_dims, hidden_dims//num_heads, num_heads, bias=bias)
self.layerNorm_2 = nn.LayerNorm(hidden_dims)
self.fullyConnected = MLP(hidden_dims,hidden_dims,hidden_dims,bias=bias)
# *****END OF YOUR CODE (DO NOT DELETE/MODIFY THIS LINE)*****
###################################################################
# END OF YOUR CODE #
###################################################################
def forward(self, x):
##############################################################
# TODO: Complete the forward of TransformerBlock module #
##############################################################
# *****START OF YOUR CODE (DO NOT DELETE/MODIFY THIS LINE)****
normalized_1 = self.layerNorm_1(x)
attn_out = self.selfAttention(normalized_1)
skipped_1 = attn_out + x
normalized_2 = self.layerNorm_2(skipped_1)
fullyConnected_out = self.fullyConnected(normalized_2)
skipped_2 = fullyConnected_out + x
return skipped_2
# *****END OF YOUR CODE (DO NOT DELETE/MODIFY THIS LINE)*****
###################################################################
# END OF YOUR CODE #
###################################################################
```
After defining the forward pass of the Transformer Block Layer above, run the following cell to test your implementation.
When you run this function, output should have shape (64, 16, 64).
```
def test_transfomerblock_layer():
x = torch.zeros((64, 16, 128), dtype=dtype) # minibatch size 64, sequence elements size 16, feature channels size 128
layer = TransformerBlock(128,4) # hidden dims size 128, heads size 4
out = layer(x)
print(out.size()) # you should see [64,16,128]
test_transfomerblock_layer()
```
# Part IV The Vision Transformer (ViT)
The final implementation for the Pytorch nn.module `ViT` is given to you below, which will perform the forward pass of the Vision Transformer. Study it and get yourself familiar with the API.
```
class ViT(nn.Module):
def __init__(self, hidden_dims, input_dims=3, output_dims=10, num_trans_layers = 4, num_heads=4, image_k=32, patch_k=4, bias=False):
super(ViT, self).__init__()
## initialize module's instance variables
self.hidden_dims = hidden_dims
self.input_dims = input_dims
self.output_dims = output_dims
self.num_trans_layers = num_trans_layers
self.num_heads = num_heads
self.image_k = image_k
self.patch_k = patch_k
self.image_height = self.image_width = image_k
self.patch_height = self.patch_width = patch_k
assert self.image_height % self.patch_height == 0 and self.image_width % self.patch_width == 0,\
'Image size must be divisible by the patch size.'
self.num_patches = (self.image_height // self.patch_height) * (self.image_width // self.patch_width)
self.patch_flat_len = self.patch_height * self.patch_width
## Declare module's parameters
## ViT's flattened patch embedding projection:
self.linear_embed = nn.Linear(self.input_dims*self.patch_flat_len, self.hidden_dims)
## Learnable positional embeddings, an embedding is learned for each patch location and the class token
self.pos_embedding = nn.Parameter(torch.randn(1, self.num_patches + 1, self.hidden_dims))
## Learnable classt token and its index among attention sequence elements.
self.cls_token = nn.Parameter(torch.randn(1,1,self.hidden_dims))
self.cls_index = torch.LongTensor([0])
## Declare cascaded Transformer blocks:
transformer_encoder_list = []
for _ in range(self.num_trans_layers):
transformer_encoder_list.append(TransformerBlock(self.hidden_dims, self.num_heads, bias))
self.transformer_encoder = nn.Sequential(*transformer_encoder_list)
## Declare the output mlp:
self.out_mlp = MLP(self.hidden_dims, self.hidden_dims, self.output_dims)
def unfold(self, x, f = 7, st = 4, p = 0):
## Create sliding window pathes using nn.Functional.unfold
## Input dimensions: [B,D,H,W] where
## --B : input batch size
## --D : input channels
## --H, W: input height and width
## Output dimensions: [B,N,H*W,D]
## --N : number of patches, decided according to sliding window kernel size (f),
## sliding window stride and padding.
b,d,h,w = x.shape
x_unf = F.unfold(x, (f,f), stride=st, padding=p)
x_unf = torch.reshape(x_unf.permute(0,2,1), (b,-1,d,f*f)).transpose(-1,-2)
n = x_unf.size(1)
return x_unf,n
def forward(self, x):
b = x.size(0)
## create sliding window patches from the input image
x_patches,n = self.unfold(x, self.patch_height, self.patch_height, 0)
## flatten each patch into a 1d vector: i.e. 3x4x4 image patch turned into 1x1x48
x_patch_flat = torch.reshape(x_patches, (b,n,-1))
## linearly embed each flattened patch
x_embed = self.linear_embed(x_patch_flat)
## retrieve class token
cls_tokens = self.cls_token.repeat(b,1,1)
## concatanate class token to input patches
xcls_embed = torch.cat([cls_tokens, x_embed], dim=-2)
## add positional embedding to input patches + class token
xcls_pos_embed = xcls_embed + self.pos_embedding
## pass through the transformer encoder
trans_out = self.transformer_encoder(xcls_pos_embed)
## select the class token
out_cls_token = torch.index_select(trans_out, -2, self.cls_index.to(trans_out.device))
## create output
out = self.out_mlp(out_cls_token)
return out.squeeze(-2)
```
After defining the forward pass of the ViT above, run the following cell to test your implementation.
When you run this function, output should have shape (64, 16, 64).
```
def test_vit():
x = torch.zeros((64, 3, 32, 32), dtype=dtype) # minibatch size 64, image size 3,32,32
model = ViT(hidden_dims=128, input_dims=3, output_dims=10, num_trans_layers = 4, num_heads=4, image_k=32, patch_k=4)
out = model(x)
print(out.size()) # you should see [64,10]
test_vit()
```
# Part V. Train the ViT
### Check Accuracy
Given any minibatch of input data and desired targets, we can check the classification accuracy of a neural network.
The check_batch_accuracy function is provided for you below:
```
def check_batch_accuracy(out, target,eps=1e-7):
b, c = out.shape
with torch.no_grad():
_, pred = out.max(-1)
correct = np.sum(np.equal(pred.cpu().numpy(), target.cpu().numpy()))
return correct, np.float(correct) / (b)
```
### Training Loop
As we have already seen in the Second Assignment, in our PyTorch based training loops, we use an Optimizer object from the `torch.optim` package, which abstract the notion of an optimization algorithm and provides implementations of most of the algorithms commonly used to optimize neural networks.
```
def train(network, optimizer, trainloader):
"""
Train a model on CIFAR-10 using the PyTorch Module API for a single epoch
Inputs:
- network: A PyTorch Module giving the model to train.
- optimizer: An Optimizer object we will use to train the model
- trainloader: Iterable DataLoader object that fetches the minibatches
Returns: overall training accuracy for the epoch
"""
print('\nEpoch: %d' % epoch)
network.train() # put model to training mode
network = network.to(device=device) # move the model parameters to CPU/GPU
train_loss = 0
correct = 0
total = 0
for batch_idx, (inputs, targets) in enumerate(trainloader):
inputs, targets = Variable(inputs.to(device)), targets.to(device) # move to device, e.g. GPU
outputs = network(inputs)
loss = F.cross_entropy(outputs, targets)
# Zero out all of the gradients for the variables which the optimizer
# will update.
optimizer.zero_grad()
# This is the backwards pass: compute the gradient of the loss with
# respect to each parameter of the model.
loss.backward()
# Actually update the parameters of the model using the gradients
# computed by the backwards pass.
optimizer.step()
loss = loss.detach()
train_loss += loss.item()
correct_p, _ = check_batch_accuracy(outputs, targets)
correct += correct_p
total += targets.size(0)
print('Loss: %.3f | Acc: %.3f%% (%d/%d)'
% (train_loss/(batch_idx+1), 100.*correct/total, correct, total))
return 100.*correct/total
```
### Evaluation Loop
We have also prepared a Evaluation loop in order to determine our networks capabilities in terms of classification accuracy on a given dataset, either the training, or the validation split
```
def evaluate(network, evalloader):
"""
Evaluate a model on CIFAR-10 using the PyTorch Module API for a single epoch
Inputs:
- network: A PyTorch Module giving the model to train.
- evalloader: Iterable DataLoader object that fetches the minibatches
Returns: overall evaluation accuracy for the epoch
"""
network.eval() # put model to evaluation mode
network = network.to(device=device) # move the model parameters to CPU/GPU
eval_loss = 0
correct = 0
total = 0
print('\n---- Evaluation in process ----')
with torch.no_grad():
for batch_idx, (inputs, targets) in enumerate(evalloader):
inputs, targets = inputs.to(device), targets.to(device) # move to device, e.g. GPU
outputs = network(inputs)
loss = F.cross_entropy(outputs, targets)
eval_loss += loss.item()
correct_p, _ = check_batch_accuracy(outputs, targets)
correct += correct_p
total += targets.size(0)
print('Loss: %.3f | Acc: %.3f%% (%d/%d)'
% (eval_loss/(batch_idx+1), 100.*correct/total, correct, total))
return 100.*correct/total
```
### Overfit a ViT
Now we are ready to run the training loop. A nice trick is to train your model with just a few training samples in order to see if your implementation is actually bug free.
Simply pass the input size, hidden layer size, and number of classes (i.e. output size) to the constructor of `ViT`.
You also need to define an optimizer that tracks all the learnable parameters inside `ViT`. We prefer to use `Adam` optimizer for this part.
You should be able to overfit small datasets, which will result in very high training accuracy and comparatively low validation accuracy.
```
sample_idx_tr = torch.randperm(len(cifar10_train))[:100]
sample_idx_val = torch.randperm(len(cifar10_train))[-100:]
trainset_sub = torch.utils.data.Subset(cifar10_train, sample_idx_tr)
valset_sub = torch.utils.data.Subset(cifar10_train, sample_idx_val)
print("For overfitting experiments, the subset of the dataset that is used has {} sample images".format(len(trainset_sub)))
batch_size_sub = 25
trainloader_sub = torch.utils.data.DataLoader(trainset_sub, batch_size=batch_size_sub, shuffle=True)
valloader_sub = torch.utils.data.DataLoader(valset_sub, batch_size=batch_size_sub, shuffle=False)
print('==> Data ready, batchsize = {}'.format(batch_size_sub))
learning_rate = 0.001
input_dims = 3
hidden_dims = 128
output_dims=10
num_trans_layers = 4
num_heads=4
image_k=32
patch_k=4
model = None
optimizer = None
################################################################################
# TODO: Instantiate your ViT model and a corresponding optimizer #
################################################################################
# *****START OF YOUR CODE (DO NOT DELETE/MODIFY THIS LINE)*****
model = ViT(hidden_dims, input_dims, output_dims, num_trans_layers, num_heads, image_k, patch_k)
optimizer = torch.optim.Adam(model.parameters(), lr=learning_rate)
# *****END OF YOUR CODE (DO NOT DELETE/MODIFY THIS LINE)*****
################################################################################
# END OF YOUR CODE
################################################################################
tr_accs=[]
eval_accs=[]
for epoch in range(15):
tr_acc = train(model, optimizer, trainloader_sub)
print('Epoch {} of training is completed, Training accuracy for this epoch is {}'\
.format(epoch, tr_acc))
eval_acc = evaluate(model, valloader_sub)
print('Evaluation of Epoch {} is completed, Validation accuracy for this epoch is {}'\
.format(epoch, eval_acc))
tr_accs.append(tr_acc)
eval_accs.append(eval_acc)
print("\nFinal train set accuracy is {}".format(tr_accs[-1]))
print("Final val set accuracy is {}".format(eval_accs[-1]))
```
## Train the net
By training the four-layer ViT network for three epochs, with untuned hyperparameters that are initialized as below, you should achieve greater than 50% accuracy both on the training set and the test set:
```
learning_rate = 0.001
input_dims = 3
hidden_dims = 128
output_dims=10
num_trans_layers = 4
num_heads=4
image_k=32
patch_k=4
network = None
optimizer = None
################################################################################
# TODO: Instantiate your ViT model and a corresponding optimizer #
################################################################################
# *****START OF YOUR CODE (DO NOT DELETE/MODIFY THIS LINE)*****
network = ViT(hidden_dims, input_dims, output_dims, num_trans_layers, num_heads, image_k, patch_k)
optimizer = torch.optim.Adam(network.parameters(), lr=learning_rate)
# *****END OF YOUR CODE (DO NOT DELETE/MODIFY THIS LINE)*****
################################################################################
# END OF YOUR CODE
################################################################################
tr_accs=[]
test_accs=[]
for epoch in range(3):
tr_acc = train(network, optimizer, loader_train)
print('Epoch {} of training is completed, Training accuracy for this epoch is {}'\
.format(epoch, tr_acc))
test_acc = evaluate(network, loader_test)
print('Evaluation of Epoch {} is completed, Test accuracy for this epoch is {}'\
.format(epoch, test_acc))
tr_accs.append(tr_acc)
test_accs.append(test_acc)
print("\nFinal train set accuracy is {}".format(tr_accs[-1]))
print("Final test set accuracy is {}".format(test_accs[-1]))
```
| github_jupyter |
# German Company Registry IDs
## Introduction
The function `clean_de_handelsregisternummer()` cleans a column containing German company registry id (handelsregisternummer) strings, and standardizes them in a given format. The function `validate_de_handelsregisternummer()` validates either a single handelsregisternummer strings, a column of handelsregisternummer strings or a DataFrame of handelsregisternummer strings, returning `True` if the value is valid, and `False` otherwise.
handelsregisternummer strings can be converted to the following formats via the `output_format` parameter:
* `compact`: only number strings without any seperators or whitespace, like "Aachen HRA 11223"
* `standard`: handelsregisternummer strings with proper whitespace in the proper places. Note that in the case of handelsregisternummer, the compact format is the same as the standard one.
Invalid parsing is handled with the `errors` parameter:
* `coerce` (default): invalid parsing will be set to NaN
* `ignore`: invalid parsing will return the input
* `raise`: invalid parsing will raise an exception
The following sections demonstrate the functionality of `clean_de_handelsregisternummer()` and `validate_de_handelsregisternummer()`.
### An example dataset containing handelsregisternummer strings
```
import pandas as pd
import numpy as np
df = pd.DataFrame(
{
"handelsregisternummer": [
'Aachen HRA 11223',
'Aachen HRC 44123',
'BE 428759497',
'BE431150351',
"002 724 334",
"hello",
np.nan,
"NULL",
],
"address": [
"123 Pine Ave.",
"main st",
"1234 west main heights 57033",
"apt 1 789 s maple rd manhattan",
"robie house, 789 north main street",
"1111 S Figueroa St, Los Angeles, CA 90015",
"(staples center) 1111 S Figueroa St, Los Angeles",
"hello",
]
}
)
df
```
## 1. Default `clean_de_handelsregisternummer`
By default, `clean_de_handelsregisternummer` will clean handelsregisternummer strings and output them in the standard format with proper separators.
```
from dataprep.clean import clean_de_handelsregisternummer
clean_de_handelsregisternummer(df, column = "handelsregisternummer")
```
## 2. Output formats
This section demonstrates the output parameter.
### `standard` (default)
```
clean_de_handelsregisternummer(df, column = "handelsregisternummer", output_format="standard")
```
### `compact`
```
clean_de_handelsregisternummer(df, column = "handelsregisternummer", output_format="compact")
```
## 3. `inplace` parameter
This deletes the given column from the returned DataFrame.
A new column containing cleaned handelsregisternummer strings is added with a title in the format `"{original title}_clean"`.
```
clean_de_handelsregisternummer(df, column="handelsregisternummer", inplace=True)
```
## 4. `errors` parameter
### `coerce` (default)
```
clean_de_handelsregisternummer(df, "handelsregisternummer", errors="coerce")
```
### `ignore`
```
clean_de_handelsregisternummer(df, "handelsregisternummer", errors="ignore")
```
## 4. `validate_de_handelsregisternummer()`
`validate_de_handelsregisternummer()` returns `True` when the input is a valid handelsregisternummer. Otherwise it returns `False`.
The input of `validate_de_handelsregisternummer()` can be a string, a Pandas DataSeries, a Dask DataSeries, a Pandas DataFrame and a dask DataFrame.
When the input is a string, a Pandas DataSeries or a Dask DataSeries, user doesn't need to specify a column name to be validated.
When the input is a Pandas DataFrame or a dask DataFrame, user can both specify or not specify a column name to be validated. If user specify the column name, `validate_de_handelsregisternummer()` only returns the validation result for the specified column. If user doesn't specify the column name, `validate_de_handelsregisternummer()` returns the validation result for the whole DataFrame.
```
from dataprep.clean import validate_de_handelsregisternummer
print(validate_de_handelsregisternummer('Aachen HRA 11223'))
print(validate_de_handelsregisternummer('Aachen HRC 44123'))
print(validate_de_handelsregisternummer('BE 428759497'))
print(validate_de_handelsregisternummer('BE431150351'))
print(validate_de_handelsregisternummer("004085616"))
print(validate_de_handelsregisternummer("hello"))
print(validate_de_handelsregisternummer(np.nan))
print(validate_de_handelsregisternummer("NULL"))
```
### Series
```
validate_de_handelsregisternummer(df["handelsregisternummer"])
```
### DataFrame + Specify Column
```
validate_de_handelsregisternummer(df, column="handelsregisternummer")
```
### Only DataFrame
```
validate_de_handelsregisternummer(df)
```
| github_jupyter |
```
import sys
sys.path.append("/home/sean/pench")
sys.path.append("/network/lustre/iss01/home/adrien.martel")
import os
# os.environ["CUDA_VISIBLE_DEVICES"]="1"
# !git clone https://github.com/vlawhern/arl-eegmodels.git
from eegmodels.EEGModels import EEGNet, ShallowConvNet, DeepConvNet
from myModels import dualLSTM, singleLSTM
import tensorflow as tf
from tensorflow import keras
tf.enable_eager_execution()
from threading import Thread
from math import sqrt
from numpy import concatenate
from matplotlib import pyplot
from pandas import read_csv
from pandas import DataFrame
from pandas import concat
from sklearn.preprocessing import MinMaxScaler
from sklearn.preprocessing import LabelEncoder
from sklearn.metrics import mean_squared_error
from sklearn.preprocessing import normalize
import math
import threading
import pickle
import numpy as np
from tensorflow.python.client import device_lib
from tensorflow.keras.utils import to_categorical
from tensorflow import keras.backend as K
# import keras
# from tqdm.keras import TqdmCallback
print(device_lib.list_local_devices()) # list of DeviceAttributes
# %gui qt
import numpy as np
# import mne
import pickle
import os
import matplotlib
import matplotlib.pyplot as plt
from multiprocessing import Pool, Queue
import multiprocessing
# tf.enable_eager_execution()
from collections import deque
from tensorflow.keras.backend import set_session
config = tf.ConfigProto()
config.gpu_options.allow_growth = True # dynamically grow the memory used on the GPU
config.log_device_placement = True # to log device placement (on which device the operation ran)
sess = tf.Session(config=config)
set_session(sess)
def randomize(a, b, c):
# Generate the permutation index array.
permutation = np.random.permutation(a.shape[0])
# Shuffle the arrays by giving the permutation in the square brackets.
shuffled_a = a[permutation]
shuffled_b = b[permutation]
shuffled_c = c[permutation]
return shuffled_a, shuffled_b, shuffled_c
baseFolder='one/'
baseFolder='/network/lustre/iss01/home/adrien.martel/data/MW/'
files=[f for f in os.listdir(baseFolder) if not f.startswith('.')]
def createData(file):
data=pickle.load(open(baseFolder+file, 'rb'))
sfreq=512
features=[]
flipFeatures=[]
labels=[]
for i in range(numClasses):
for k in range(len(data[i])):
labels.append(i)
features.append(data[i][k])
flipFeatures.append([np.transpose(data[i][k])])
labels=np.array(labels)
features=np.array(features)
flipFeatures=np.array(flipFeatures)
labels, features, flipFeatures = randomize(labels, features, flipFeatures)
labels = to_categorical(labels, num_classes=numClasses)
return [features, flipFeatures, labels]
sam=2560
chans=62
numClasses=2
models = [
[EEGNet(nb_classes=numClasses, Chans=chans, Samples=sam), True, 'EEGNet-V1'],
[ShallowConvNet(nb_classes=numClasses, Chans=chans, Samples =sam), True, 'ShallowConvNet-V1'],
[DeepConvNet(nb_classes=numClasses, Chans=chans, Samples=sam), True, 'DeepConvNet-V1'],
[singleLSTM(clas=numClasses, sam=sam, chans=chans), False, 'singleLSTM-V1'],
[dualLSTM(clas=numClasses, sam=sam, chans=chans), False, 'dualLSTM-V1'],
]
def createWork(n):
arc=inps[n][0]
file=inps[n][1]
features, flipFeatures, labels = createData(file)
if arc[1]:
train_X = np.array(flipFeatures[0:int(7*len(labels)/10)])
test_X = np.array(flipFeatures[int(7*len(labels)/10):-1])
else:
train_X = np.array(features[0:int(7*len(labels)/10)])
test_X = np.array(features[int(7*len(labels)/10):-1])
train_y = np.array(labels[0:int(7*len(labels)/10)])
test_y = np.array(labels[int(7*len(labels)/10):-1])
print("Putted", file, out.empty())
# out.put([arc[0], train_X, test_X, train_y, test_y, file, arc[2]])
print(file, train_X.shape, test_X.shape, train_y.shape, test_y.shape)
# print(out.empty())
return [arc[0], train_X, test_X, train_y, test_y, file, arc[2]]
inps=[]
for model in models:
try:
os.mkdir(model[2])
except:
print("probably exists")
for file in files:
inps.append([model, file])
manager = multiprocessing.Manager()
out = manager.Queue()
# p = Pool(20)
# master=p.map(createWork, list(range(2)))
# master=p.map(createWork, list(range(len(inps))))
gpus=4
out.empty()
# out = Queue()
# out.queue = queue.deque(master)
[out.put(i) for i in list(range(len(inps)))]
out.empty()
def doWork(i):
# i = args[0]
# out = args[1]
# i=1
os.environ["CUDA_VISIBLE_DEVICES"]=str(i)
while not out.empty():
with tf.Session(config=tf.ConfigProto(allow_soft_placement=True, device_count = {'GPU': i}), graph=tf.Graph()) as sess:
K.set_session(sess)
# dat=out.get()
# print("not empty")
n=out.get()
dat= createWork(n)
model=dat[0]
train_X=dat[1]
test_X=dat[2]
train_y=dat[3]
test_y=dat[4]
file=dat[5]
folder=dat[6]
# print('processed')
# sgd = keras.optimizers.SGD(learning_rate=0.015, momentum=0.0, nesterov=False)
# adam = keras.optimizers.Adam(learning_rate=0.001, beta_1=0.9, beta_2=0.999, amsgrad=False)
print('Done getting data')
# sgd = keras.optimizers.SGD()
adam = tf.train.AdamOptimizer(
learning_rate=0.001, beta1=0.9, beta2=0.999, epsilon=1e-08, use_locking=False,
name='Adam'
)
print('Compiling model')
# break
model.compile(loss='categorical_crossentropy', optimizer=adam, metrics=['accuracy'])
# fit network
history = model.fit(train_X, train_y, epochs=10, batch_size=2, validation_data=(test_X, test_y), verbose=0, shuffle=True)
# plot history
print(history.history.keys())
pyplot.figure(figsize=(25,10), dpi=250)
pyplot.plot(history.history['loss'], label='train')
pyplot.plot(history.history['val_loss'], label='test')
pyplot.plot(history.history['accuracy'], label='accuracy')
pyplot.plot(history.history['val_accuracy'], label='test accuracy')
pyplot.legend()
pyplot.savefig(folder+'/'+file + '.png')
pickle.dump(history, open(folder+'/'+file+'-hist.p', "wb"))
model.save(folder+'/'+file+'.h5')
print('done')
workers=[]
for i in range(gpus):
workers.append(Thread(target = doWork, args=(i,)))
for worker in workers:
worker.start()
# s = Pool(2)
# master=s.map(doWork, [(x, out) for x in range(gpus)])
worker.join()
```
| github_jupyter |
# Home 4: Build a seq2seq model for machine translation.
### Name: [Your-Name?]
### Task: Translate English to [what-language?]
## 0. You will do the following:
1. Read and run my code.
2. Complete the code in Section 1.1 and Section 4.2.
* Translation English to **German** is not acceptable!!! Try another language.
3. **Make improvements.** Directly modify the code in Section 3. Do at least one of the followings. By doing more, you will get up to 2 bonus scores to the total.
* Bi-LSTM instead of LSTM
* Multi-task learning (e.g., both English to French and English to Spanish)
* Attention
4. Evaluate the translation using the BLEU score.
* Optional. Up to 2 bonus scores to the total.
5. Convert the notebook to .HTML file.
* The HTML file must contain the code and the output after execution.
6. Put the .HTML file in your own Github repo.
7. Submit the link to the HTML file to Canvas
* E.g., https://github.com/wangshusen/CS583A-2019Spring/blob/master/homework/HM4/seq2seq.html
### Hint:
To implement ```Bi-LSTM```, you will need the following code to build the encoder; the decoder won't be much different.
```
from keras.layers import Bidirectional, Concatenate
encoder_bilstm = Bidirectional(LSTM(latent_dim, return_state=True,
dropout=0.5, name='encoder_lstm'))
_, forward_h, forward_c, backward_h, backward_c = encoder_bilstm(encoder_inputs)
state_h = Concatenate()([forward_h, backward_h])
state_c = Concatenate()([forward_c, backward_c])
```
### Hint:
To implement multi-task training, you can refer to ```Section 7.1.3 Multi-output models``` of the textbook, ```Deep Learning with Python```.
## 1. Data preparation
1. Download data (e.g., "deu-eng.zip") from http://www.manythings.org/anki/
2. Unzip the .ZIP file.
3. Put the .TXT file (e.g., "deu.txt") in the directory "./Data/".
### 1.1. Load and clean text
```
import re
import string
from unicodedata import normalize
import numpy
# load doc into memory
def load_doc(filename):
# open the file as read only
file = open(filename, mode='rt', encoding='utf-8')
# read all text
text = file.read()
# close the file
file.close()
return text
# split a loaded document into sentences
def to_pairs(doc):
lines = doc.strip().split('\n')
pairs = [line.split('\t') for line in lines]
return pairs
def clean_data(lines):
cleaned = list()
# prepare regex for char filtering
re_print = re.compile('[^%s]' % re.escape(string.printable))
# prepare translation table for removing punctuation
table = str.maketrans('', '', string.punctuation)
for pair in lines:
clean_pair = list()
for line in pair:
# normalize unicode characters
line = normalize('NFD', line).encode('ascii', 'ignore')
line = line.decode('UTF-8')
# tokenize on white space
line = line.split()
# convert to lowercase
line = [word.lower() for word in line]
# remove punctuation from each token
line = [word.translate(table) for word in line]
# remove non-printable chars form each token
line = [re_print.sub('', w) for w in line]
# remove tokens with numbers in them
line = [word for word in line if word.isalpha()]
# store as string
clean_pair.append(' '.join(line))
cleaned.append(clean_pair)
return numpy.array(cleaned)
```
#### Fill the following blanks:
```
# e.g., filename = 'Data/deu.txt'
filename = <what is your file name?>
# e.g., n_train = 20000
n_train = <how many sentences are you going to use for training?>
# load dataset
doc = load_doc(filename)
# split into Language1-Language2 pairs
pairs = to_pairs(doc)
# clean sentences
clean_pairs = clean_data(pairs)[0:n_train, :]
for i in range(3000, 3010):
print('[' + clean_pairs[i, 0] + '] => [' + clean_pairs[i, 1] + ']')
input_texts = clean_pairs[:, 0]
target_texts = ['\t' + text + '\n' for text in clean_pairs[:, 1]]
print('Length of input_texts: ' + str(input_texts.shape))
print('Length of target_texts: ' + str(input_texts.shape))
max_encoder_seq_length = max(len(line) for line in input_texts)
max_decoder_seq_length = max(len(line) for line in target_texts)
print('max length of input sentences: %d' % (max_encoder_seq_length))
print('max length of target sentences: %d' % (max_decoder_seq_length))
```
**Remark:** To this end, you have two lists of sentences: input_texts and target_texts
## 2. Text processing
### 2.1. Convert texts to sequences
- Input: A list of $n$ sentences (with max length $t$).
- It is represented by a $n\times t$ matrix after the tokenization and zero-padding.
```
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
# encode and pad sequences
def text2sequences(max_len, lines):
tokenizer = Tokenizer(char_level=True, filters='')
tokenizer.fit_on_texts(lines)
seqs = tokenizer.texts_to_sequences(lines)
seqs_pad = pad_sequences(seqs, maxlen=max_len, padding='post')
return seqs_pad, tokenizer.word_index
encoder_input_seq, input_token_index = text2sequences(max_encoder_seq_length,
input_texts)
decoder_input_seq, target_token_index = text2sequences(max_decoder_seq_length,
target_texts)
print('shape of encoder_input_seq: ' + str(encoder_input_seq.shape))
print('shape of input_token_index: ' + str(len(input_token_index)))
print('shape of decoder_input_seq: ' + str(decoder_input_seq.shape))
print('shape of target_token_index: ' + str(len(target_token_index)))
num_encoder_tokens = len(input_token_index) + 1
num_decoder_tokens = len(target_token_index) + 1
print('num_encoder_tokens: ' + str(num_encoder_tokens))
print('num_decoder_tokens: ' + str(num_decoder_tokens))
```
**Remark:** To this end, the input language and target language texts are converted to 2 matrices.
- Their number of rows are both n_train.
- Their number of columns are respective max_encoder_seq_length and max_decoder_seq_length.
The followings print a sentence and its representation as a sequence.
```
target_texts[100]
decoder_input_seq[100, :]
```
## 2.2. One-hot encode
- Input: A list of $n$ sentences (with max length $t$).
- It is represented by a $n\times t$ matrix after the tokenization and zero-padding.
- It is represented by a $n\times t \times v$ tensor ($t$ is the number of unique chars) after the one-hot encoding.
```
from keras.utils import to_categorical
# one hot encode target sequence
def onehot_encode(sequences, max_len, vocab_size):
n = len(sequences)
data = numpy.zeros((n, max_len, vocab_size))
for i in range(n):
data[i, :, :] = to_categorical(sequences[i], num_classes=vocab_size)
return data
encoder_input_data = onehot_encode(encoder_input_seq, max_encoder_seq_length, num_encoder_tokens)
decoder_input_data = onehot_encode(decoder_input_seq, max_decoder_seq_length, num_decoder_tokens)
decoder_target_seq = numpy.zeros(decoder_input_seq.shape)
decoder_target_seq[:, 0:-1] = decoder_input_seq[:, 1:]
decoder_target_data = onehot_encode(decoder_target_seq,
max_decoder_seq_length,
num_decoder_tokens)
print(encoder_input_data.shape)
print(decoder_input_data.shape)
```
## 3. Build the networks (for training)
- Build encoder, decoder, and connect the two modules to get "model".
- Fit the model on the bilingual data to train the parameters in the encoder and decoder.
### 3.1. Encoder network
- Input: one-hot encode of the input language
- Return:
-- output (all the hidden states $h_1, \cdots , h_t$) are always discarded
-- the final hidden state $h_t$
-- the final conveyor belt $c_t$
```
from keras.layers import Input, LSTM
from keras.models import Model
latent_dim = 256
# inputs of the encoder network
encoder_inputs = Input(shape=(None, num_encoder_tokens),
name='encoder_inputs')
# set the LSTM layer
encoder_lstm = LSTM(latent_dim, return_state=True,
dropout=0.5, name='encoder_lstm')
_, state_h, state_c = encoder_lstm(encoder_inputs)
# build the encoder network model
encoder_model = Model(inputs=encoder_inputs,
outputs=[state_h, state_c],
name='encoder')
```
Print a summary and save the encoder network structure to "./encoder.pdf"
```
from IPython.display import SVG
from keras.utils.vis_utils import model_to_dot, plot_model
SVG(model_to_dot(encoder_model, show_shapes=False).create(prog='dot', format='svg'))
plot_model(
model=encoder_model, show_shapes=False,
to_file='encoder.pdf'
)
encoder_model.summary()
```
### 3.2. Decoder network
- Inputs:
-- one-hot encode of the target language
-- The initial hidden state $h_t$
-- The initial conveyor belt $c_t$
- Return:
-- output (all the hidden states) $h_1, \cdots , h_t$
-- the final hidden state $h_t$ (discarded in the training and used in the prediction)
-- the final conveyor belt $c_t$ (discarded in the training and used in the prediction)
```
from keras.layers import Input, LSTM, Dense
from keras.models import Model
# inputs of the decoder network
decoder_input_h = Input(shape=(latent_dim,), name='decoder_input_h')
decoder_input_c = Input(shape=(latent_dim,), name='decoder_input_c')
decoder_input_x = Input(shape=(None, num_decoder_tokens), name='decoder_input_x')
# set the LSTM layer
decoder_lstm = LSTM(latent_dim, return_sequences=True,
return_state=True, dropout=0.5, name='decoder_lstm')
decoder_lstm_outputs, state_h, state_c = decoder_lstm(decoder_input_x,
initial_state=[decoder_input_h, decoder_input_c])
# set the dense layer
decoder_dense = Dense(num_decoder_tokens, activation='softmax', name='decoder_dense')
decoder_outputs = decoder_dense(decoder_lstm_outputs)
# build the decoder network model
decoder_model = Model(inputs=[decoder_input_x, decoder_input_h, decoder_input_c],
outputs=[decoder_outputs, state_h, state_c],
name='decoder')
```
Print a summary and save the encoder network structure to "./decoder.pdf"
```
from IPython.display import SVG
from keras.utils.vis_utils import model_to_dot, plot_model
SVG(model_to_dot(decoder_model, show_shapes=False).create(prog='dot', format='svg'))
plot_model(
model=decoder_model, show_shapes=False,
to_file='decoder.pdf'
)
decoder_model.summary()
```
### 3.3. Connect the encoder and decoder
```
# input layers
encoder_input_x = Input(shape=(None, num_encoder_tokens), name='encoder_input_x')
decoder_input_x = Input(shape=(None, num_decoder_tokens), name='decoder_input_x')
# connect encoder to decoder
encoder_final_states = encoder_model([encoder_input_x])
decoder_lstm_output, _, _ = decoder_lstm(decoder_input_x, initial_state=encoder_final_states)
decoder_pred = decoder_dense(decoder_lstm_output)
model = Model(inputs=[encoder_input_x, decoder_input_x],
outputs=decoder_pred,
name='model_training')
print(state_h)
print(decoder_input_h)
from IPython.display import SVG
from keras.utils.vis_utils import model_to_dot, plot_model
SVG(model_to_dot(model, show_shapes=False).create(prog='dot', format='svg'))
plot_model(
model=model, show_shapes=False,
to_file='model_training.pdf'
)
model.summary()
```
### 3.5. Fit the model on the bilingual dataset
- encoder_input_data: one-hot encode of the input language
- decoder_input_data: one-hot encode of the input language
- decoder_target_data: labels (left shift of decoder_input_data)
- tune the hyper-parameters
- stop when the validation loss stop decreasing.
```
print('shape of encoder_input_data' + str(encoder_input_data.shape))
print('shape of decoder_input_data' + str(decoder_input_data.shape))
print('shape of decoder_target_data' + str(decoder_target_data.shape))
model.compile(optimizer='rmsprop', loss='categorical_crossentropy')
model.fit([encoder_input_data, decoder_input_data], # training data
decoder_target_data, # labels (left shift of the target sequences)
batch_size=64, epochs=50, validation_split=0.2)
model.save('seq2seq.h5')
```
## 4. Make predictions
### 4.1. Translate English to XXX
1. Encoder read a sentence (source language) and output its final states, $h_t$ and $c_t$.
2. Take the [star] sign "\t" and the final state $h_t$ and $c_t$ as input and run the decoder.
3. Get the new states and predicted probability distribution.
4. sample a char from the predicted probability distribution
5. take the sampled char and the new states as input and repeat the process (stop if reach the [stop] sign "\n").
```
# Reverse-lookup token index to decode sequences back to something readable.
reverse_input_char_index = dict((i, char) for char, i in input_token_index.items())
reverse_target_char_index = dict((i, char) for char, i in target_token_index.items())
def decode_sequence(input_seq):
states_value = encoder_model.predict(input_seq)
target_seq = numpy.zeros((1, 1, num_decoder_tokens))
target_seq[0, 0, target_token_index['\t']] = 1.
stop_condition = False
decoded_sentence = ''
while not stop_condition:
output_tokens, h, c = decoder_model.predict([target_seq] + states_value)
# this line of code is greedy selection
# try to use multinomial sampling instead (with temperature)
sampled_token_index = numpy.argmax(output_tokens[0, -1, :])
sampled_char = reverse_target_char_index[sampled_token_index]
decoded_sentence += sampled_char
if (sampled_char == '\n' or
len(decoded_sentence) > max_decoder_seq_length):
stop_condition = True
target_seq = numpy.zeros((1, 1, num_decoder_tokens))
target_seq[0, 0, sampled_token_index] = 1.
states_value = [h, c]
return decoded_sentence
for seq_index in range(2100, 2120):
# Take one sequence (part of the training set)
# for trying out decoding.
input_seq = encoder_input_data[seq_index: seq_index + 1]
decoded_sentence = decode_sequence(input_seq)
print('-')
print('English: ', input_texts[seq_index])
print('German (true): ', target_texts[seq_index][1:-1])
print('German (pred): ', decoded_sentence[0:-1])
```
### 4.2. Translate an English sentence to the target language
1. Tokenization
2. One-hot encode
3. Translate
```
input_sentence = 'why is that'
input_sequence = <do tokenization...>
input_x = <do one-hot encode...>
translated_sentence = <do translation...>
print('source sentence is: ' + input_sentence)
print('translated sentence is: ' + translated_sentence)
```
## 5. Evaluate the translation using BLEU score
Reference:
- https://machinelearningmastery.com/calculate-bleu-score-for-text-python/
- https://en.wikipedia.org/wiki/BLEU
**Hint:** Randomly partition the dataset to training, validation, and test. Evaluate the BLEU score using the test set.
| github_jupyter |
###### Content under Creative Commons Attribution license CC-BY 4.0, code under MIT license (c)2014 L.A. Barba, G.F. Forsyth.
# Relax and hold steady
Ready for more relaxing? This is the third lesson of **Module 5** of the course, exploring solutions to elliptic PDEs.
In [Lesson 1](http://nbviewer.ipython.org/github/numerical-mooc/numerical-mooc/blob/master/lessons/05_relax/05_01_2D.Laplace.Equation.ipynb) and [Lesson 2](http://nbviewer.ipython.org/github/numerical-mooc/numerical-mooc/blob/master/lessons/05_relax/05_02_2D.Poisson.Equation.ipynb) of this module we used the Jacobi method (a relaxation scheme) to iteratively find solutions to Laplace and Poisson equations.
And it worked, so why are we still talking about it? Because the Jacobi method is slow, very slow to converge. It might not have seemed that way in the first two notebooks because we were using small grids, but we did need more than 3,000 iterations to reach the exit criterion while solving the Poisson equation on a $41\times 41$ grid.
You can confirm this below: using `nx,ny=` $128$ on the Laplace problem of Lesson 1, the Jacobi method requires nearly *20,000* iterations before we reach $10^{-8}$ for the L2-norm of the difference between two iterates. That's a *lot* of iterations!
Now, consider this application: an incompressible Navier-Stokes solver has to ensure that the velocity field is divergence-free at every timestep. One of the most common ways to ensure this is to solve a Poisson equation for the pressure field. In fact, the pressure Poisson equation is responsible for the majority of the computational expense of an incompressible Navier-Stokes solver. Imagine having to do 20,000 Jacobi iterations for *every* time step in a fluid-flow problem with many thousands or perhaps millions of grid points!
The Jacobi method is the slowest of all relaxation schemes, so let's learn how to improve on it. In this lesson, we'll study the Gauss-Seidel method—twice as fast as Jacobi, in theory—and the successive over-relaxation (SOR) method. We also have some neat Python tricks lined up for you to get to the solution even faster. Let's go!
### Test problem
Let's use the same example problem as in [Lesson 1](./05_01_2D.Laplace.Equation.ipynb): Laplace's equation with boundary conditions
\begin{equation}
\begin{gathered}
p=0 \text{ at } x=0\\
\frac{\partial p}{\partial x} = 0 \text{ at } x = L\\
p = 0 \text{ at }y = 0 \\
p = \sin \left( \frac{\frac{3}{2}\pi x}{L} \right) \text{ at } y = H
\end{gathered}
\end{equation}
We import our favorite Python libraries, and also some custom functions that we wrote in [Lesson 1](./05_01_2D.Laplace.Equation.ipynb), which we have saved in a 'helper' Python file for re-use.
```
import numpy
from matplotlib import pyplot, cm
from mpl_toolkits.mplot3d import Axes3D
%matplotlib inline
from matplotlib import rcParams
rcParams['font.family'] = 'serif'
rcParams['font.size'] = 16
from laplace_helper import p_analytical, plot_3D, L2_rel_error
```
We now have the analytical solution in the array `p_analytical`, and we have the functions `plot_3D` and `L2_rel_error` in our namespace. If you can't remember how they work, just use `help()` and take advantage of the docstrings. It's a good habit to always write docstrings in your functions, and now you see why!
In this notebook, we are going to use larger grids than before, to better illustrate the speed increases we achieve with different iterative methods. Let's create a $128\times128$ grid and initialize.
```
nx = 128
ny = 128
L = 5
H = 5
x = numpy.linspace(0,L,nx)
y = numpy.linspace(0,H,ny)
dx = L/(nx-1)
dy = H/(ny-1)
p0 = numpy.zeros((ny, nx))
p0[-1,:] = numpy.sin(1.5*numpy.pi*x/x[-1])
```
We said above that the Jacobi method takes nearly 20,000 iterations before it satisfies our exit criterion of $10^{-8}$ (L2-norm difference between two consecutive iterations). You'll just have to confirm that now. Have a seat!
```
def laplace2d(p, l2_target):
'''Solves the Laplace equation using the Jacobi method
with a 5-point stencil
Parameters:
----------
p: 2D array of float
Initial potential distribution
l2_target: float
Stopping criterion
Returns:
-------
p: 2D array of float
Potential distribution after relaxation
'''
l2norm = 1
pn = numpy.empty_like(p)
iterations = 0
while l2norm > l2_target:
pn = p.copy()
p[1:-1,1:-1] = .25 * (pn[1:-1,2:] + pn[1:-1,:-2] +\
pn[2:,1:-1] + pn[:-2,1:-1])
##Neumann B.C. along x = L
p[1:-1,-1] = .25 * (2*pn[1:-1,-2] + pn[2:,-1] + pn[:-2, -1])
l2norm = numpy.sqrt(numpy.sum((p - pn)**2)/numpy.sum(pn**2))
iterations += 1
return p, iterations
l2_target = 1e-8
p, iterations = laplace2d(p0.copy(), l2_target)
print ("Jacobi method took {} iterations at tolerance {}".\
format(iterations, l2_target))
```
Would we lie to you? 19,993 iterations before we reach the exit criterion of $10^{-8}$. Yikes!
We can also time how long the Jacobi method takes using the `%%timeit` cell-magic. Go make some tea, because this can take a while—the `%%timeit` magic runs the function a few times and then averages their runtimes to give a more accurate result.
- - -
##### Notes
1. When using `%%timeit`, the return values of a function (`p` and `iterations` in this case) *won't* be saved.
2. We document our timings below, but your timings can vary quite a lot, depending on your hardware. In fact, you may not even see the same trends (some recent hardware can play some fancy tricks with optimizations that you have no control over).
- - -
With those caveats, let's give it a shot:
```
%%timeit
laplace2d(p0.copy(), l2_target)
```
The printed result above (and others to come later) is from a mid-2007 Mac Pro, powered by two 3-GHz quad-core Intel Xeon X5364 (Clovertown). We tried also on more modern machines, and get conflicting results—like the Gauss-Seidel method being slightly slower than Jacobi, even though it required fewer iterations. Don't get too hung up on this: the hardware optimizations applied by more modern CPUs are varied and make a big difference sometimes.
Meanwhile, let's check the overall accuracy of the numerical calculation by comparing it to the analytical solution.
```
pan = p_analytical(x,y)
L2_rel_error(p,pan)
```
That's a pretty small error. Let's assume it is good enough and focus on speeding up the process.
## Gauss-Seidel
You will recall from [Lesson 1](./2D_Laplace_Equation.ipynb) that a single Jacobi iteration is written as:
\begin{equation}
p^{k+1}_{i,j} = \frac{1}{4} \left(p^{k}_{i,j-1} + p^k_{i,j+1} + p^{k}_{i-1,j} + p^k_{i+1,j} \right)
\end{equation}
The Gauss-Seidel method is a simple tweak to this idea: use updated values of the solution as soon as they are available, instead of waiting for the values in the whole grid to be updated.
If you imagine that we progress through the grid points in the order shown by the arrow in Figure 1, then you can see that the updated values $p^{k+1}_{i-1,j}$ and $p^{k+1}_{i,j-1}$ can be used to calculate $p^{k+1}_{i,j}$.
<img src="./figures/solvepath.svg" width=350>
#### Figure 1. Assumed order of updates on a grid.
The iteration formula for Gauss-Seidel is thus:
\begin{equation}
p^{k+1}_{i,j} = \frac{1}{4} \left(p^{k+1}_{i,j-1} + p^k_{i,j+1} + p^{k+1}_{i-1,j} + p^k_{i+1,j} \right)
\end{equation}
There's now a problem for the Python implementation. You can no longer use NumPy's array operations to evaluate the solution updates. Since Gauss-Seidel requires using values immediately after they're updated, we have to abandon our beloved array operations and return to nested `for` loops. Ugh.
We don't like it, but if it saves us a bunch of time, then we can manage. But does it?
Here's a function to compute the Gauss-Seidel updates using a double loop.
```
def laplace2d_gauss_seidel(p, nx, ny, l2_target):
iterations = 0
iter_diff = l2_target+1 #init iter_diff to be larger than l2_target
while iter_diff > l2_target:
pn = p.copy()
iter_diff = 0.0
for j in range(1,ny-1):
for i in range(1,nx-1):
p[j,i] = .25 * (p[j,i-1] + p[j,i+1] + p[j-1,i] + p[j+1,i])
iter_diff += (p[j,i] - pn[j,i])**2
#Neumann 2nd-order BC
for j in range(1,ny-1):
p[j,-1] = .25 * (2*p[j,-2] + p[j+1,-1] + p[j-1, -1])
iter_diff = numpy.sqrt(iter_diff/numpy.sum(pn**2))
iterations += 1
return p, iterations
```
We would then run this with the following function call:
```Python
p, iterations = laplace2d_gauss_seidel(p,1e-8)
```
<br>
But **don't do it**. We did it so that you don't have to!
The solution of our test problem with the Gauss-Seidel method required several thousand fewer iterations than the Jacobi method, but it took nearly *10 minutes* to run on our machine.
##### What happened?
If you think back to the far off days when you first learned about array operations, you might recall that we discovered that NumPy array operations could drastically improve code performance compared with nested `for` loops. NumPy operations are written in C and pre-compiled, so they are *much* faster than vanilla Python.
But the Jacobi method is not algorithmically optimal, giving slow convergence. We want to take advantage of the faster-converging iterative methods, yet unpacking the array operations into nested loops destroys performance. *What can we do?*
## Use Numba!
[Numba](http://numba.pydata.org) is an open-source optimizing compiler for Python. It works by reading Python functions that you give it, and generating a compiled version for you—also called Just-In-Time (JIT) compilation. You can then use the function at performance levels that are close to what you can get with compiled languages (like C, C++ and fortran).
It can massively speed up performance, especially when dealing with loops. Plus, it's pretty easy to use. Like we overheard at a conference: [*Numba is a Big Deal.*](http://twitter.com/lorenaabarba/status/625383941453656065)
##### Caveat
We encourage everyone following the course to use the [Anaconda Python](https://www.continuum.io/downloads) distribution because it's well put-together and simple to use. If you *haven't* been using Anaconda, that's fine, but let us **strongly** suggest that you take the plunge now. Numba is great and easy to use, but it is **not** easy to install without help. Those of you using Anaconda can install it by running <br><br>
`conda install numba`<br><br>
If you *really* don't want to use Anaconda, you will have to [compile all of Numba's dependencies](https://pypi.python.org/pypi/numba).
- - -
### Intro to Numba
Let's dive in! Numba is great and easy to use. We're going to first walk you through a simple example to give you a taste of Numba's abilities.
After installing Numba (see above), we can use it by adding a line to `import numba` and another to `import autojit` (more on this in a bit).
```
import numba
from numba import jit
```
You tell Numba which functions you want to accelerate by using a [Python decorator](http://www.learnpython.org/en/Decorators), a special type of command that tells the Python interpreter to modify a callable object (like a function). For example, let's write a quick function to calculate the $n^{\text{th}}$ number in the Fibonacci sequence:
```
def fib_it(n):
a = 1
b = 1
for i in range(n-2):
a, b = b, a+b
return b
```
There are several faster ways to program the Fibonacci sequence, but that's not a concern right now (but if you're curious, [check them out](http://mathworld.wolfram.com/BinetsFibonacciNumberFormula.html)). Let's use `%%timeit` and see how long this simple function takes to find the 500,000-th Fibonacci number.
```
%%timeit
fib_it(500000)
```
Now let's try Numba! Just add the `@jit` decorator above the function name and let's see what happens!
```
@jit
def fib_it(n):
a = 1
b = 1
for i in range(n-2):
a, b = b, a+b
return b
%%timeit
fib_it(500000)
```
*Holy cow!* In our machine, that's more than 8,000 times faster!
That warning from `%%timeit` is due to the compilation overhead for Numba. The very first time that it executes the function, it has to compile it, then it caches that code for reuse without extra compiling. That's the 'Just-In-Time' bit. You'll see it disappear if we run `%%timeit` again.
```
%%timeit
fib_it(500000)
```
We would agree if you think that this is a rather artificial example, but the speed-up is very impressive indeed. Just adding the one-word decorator!
##### Running in `nopython` mode
Numba is very clever, but it can't optimize everything. When it can't, rather than failing to run, it will fall back to the regular Python, resulting in poor performance again. This can be confusing and frustrating, since you might not know ahead of time which bits of code will speed up and which bits won't.
To avoid this particular annoyance, you can tell Numba to use `nopython` mode. In this case, your code will simply fail if the "jitted" function can't be optimized. It's simply an option to give you "fast or nothing."
Use `nopython` mode by adding the following line above the function that you want to JIT-compile:
```Python
@jit(nopython=True)
```
- - -
##### Numba version check
In these examples, we are using the latest (as of publication) version of Numba: 0.22.1. Make sure to upgrade or some of the code examples below may not run.
- - -
```
print(numba.__version__)
```
## Back to Jacobi
We want to compare the performance of different iterative methods under the same conditions. Because the Gauss-Seidel method forces us to unpack the array operations into nested loops (which are very slow in Python), we use Numba to get the code to perform well. Thus, we need to write a new Jacobi method using for-loops and Numba (instead of NumPy), so we can make meaningful comparisons.
Let's write a "jitted" Jacobi with loops.
```
@jit(nopython=True)
def laplace2d_jacobi(p, pn, l2_target):
'''Solves the Laplace equation using the Jacobi method
with a 5-point stencil
Parameters:
----------
p: 2D array of float
Initial potential distribution
pn: 2D array of float
Allocated array for previous potential distribution
l2_target: float
Stopping criterion
Returns:
-------
p: 2D array of float
Potential distribution after relaxation
'''
iterations = 0
iter_diff = l2_target+1 #init iter_diff to be larger than l2_target
denominator = 0.0
ny, nx = p.shape
l2_diff = numpy.zeros(20000)
while iter_diff > l2_target:
for j in range(ny):
for i in range(nx):
pn[j,i] = p[j,i]
iter_diff = 0.0
denominator = 0.0
for j in range(1,ny-1):
for i in range(1,nx-1):
p[j,i] = .25 * (pn[j,i-1] + pn[j,i+1] + pn[j-1,i] + pn[j+1,i])
#Neumann 2nd-order BC
for j in range(1,ny-1):
p[j,-1] = .25 * (2*pn[j,-2] + pn[j+1,-1] + pn[j-1, -1])
for j in range(ny):
for i in range(nx):
iter_diff += (p[j,i] - pn[j,i])**2
denominator += (pn[j,i]*pn[j,i])
iter_diff /= denominator
iter_diff = iter_diff**0.5
l2_diff[iterations] = iter_diff
iterations += 1
return p, iterations, l2_diff
p, iterations, l2_diffJ = laplace2d_jacobi(p0.copy(), p0.copy(), 1e-8)
print("Numba Jacobi method took {} iterations at tolerance {}".format(iterations, l2_target))
%%timeit
laplace2d_jacobi(p0.copy(), p0.copy(), 1e-8)
```
In our old machine, that's faster than the NumPy version of Jacobi, but on some newer machines it might not be. Don't obsess over this: there is much hardware black magic that we cannot control.
Remember that NumPy is a highly optimized library. The fact that we can get competitive execution times with this JIT-compiled code is kind of amazing. Plus(!) now we get to try out those techniques that aren't possible with NumPy array operations.
##### Note
We're also saving the history of the L2-norm of the difference between consecutive iterations. We'll take a look at that once we have a few more methods to compare.
- - -
##### Another Note
Why did we use
```Python
l2_diff = numpy.zeros(20000)
```
Where did the `20000` come from?
We cheated a little bit. Numba doesn't handle _mutable_ objects well in `nopython` mode, which means we can't use a *list* and append each iteration's value of the L2-norm. So we need to define an array big enough to hold all of them and we know from the first run that Jacobi converges in fewer than 20,000 iterations.
- - -
##### Challenge task
It is possible to get a good estimate of the number of iterations needed by the Jacobi method to reduce the initial error by a factor $10^{-m}$, for given $m$. The formula depends on the largest eigenvalue of the coefficient matrix, which is known for the discrete Poisson problem on a square domain. See Parviz Moin, *"Fundamentals of Engineering Numerical Analysis"* (2nd ed., pp.141–143).
* Find the estimated number of iterations to reduce the initial error by $10^{-8}$ when using the grids listed below, in the section on grid convergence, with $11$, $21$, $41$ and $81$ grid points on each coordinate axis.
## Back to Gauss-Seidel
If you recall, the reason we got into this Numba sidetrack was to try out Gauss-Seidel and compare the performance with Jacobi. Recall from above that the formula for Gauss-Seidel is as follows:
\begin{equation}
p^{k+1}_{i,j} = \frac{1}{4} \left(p^{k+1}_{i,j-1} + p^k_{i,j+1} + p^{k+1}_{i-1,j} + p^k_{i+1,j} \right)
\end{equation}
We only need to slightly tweak the Jacobi function to get one for Gauss-Seidel. Instead of updating `p` in terms of `pn`, we just update `p` using `p`!
```
@jit(nopython=True)
def laplace2d_gauss_seidel(p, pn, l2_target):
'''Solves the Laplace equation using Gauss-Seidel method
with a 5-point stencil
Parameters:
----------
p: 2D array of float
Initial potential distribution
pn: 2D array of float
Allocated array for previous potential distribution
l2_target: float
Stopping criterion
Returns:
-------
p: 2D array of float
Potential distribution after relaxation
'''
iterations = 0
iter_diff = l2_target + 1 #initialize iter_diff to be larger than l2_target
denominator = 0.0
ny, nx = p.shape
l2_diff = numpy.zeros(20000)
while iter_diff > l2_target:
for j in range(ny):
for i in range(nx):
pn[j,i] = p[j,i]
iter_diff = 0.0
denominator = 0.0
for j in range(1,ny-1):
for i in range(1,nx-1):
p[j,i] = .25 * (p[j,i-1] + p[j,i+1] + p[j-1,i] + p[j+1,i])
#Neumann 2nd-order BC
for j in range(1,ny-1):
p[j,-1] = .25 * (2*p[j,-2] + p[j+1,-1] + p[j-1, -1])
for j in range(ny):
for i in range(nx):
iter_diff += (p[j,i] - pn[j,i])**2
denominator += (pn[j,i]*pn[j,i])
iter_diff /= denominator
iter_diff = iter_diff**0.5
l2_diff[iterations] = iter_diff
iterations += 1
return p, iterations, l2_diff
p, iterations, l2_diffGS = laplace2d_gauss_seidel(p0.copy(), p0.copy(), 1e-8)
print("Numba Gauss-Seidel method took {} iterations at tolerance {}".format(iterations, l2_target))
```
Cool! Using the most recently updated values of the solution in the Gauss-Seidel method saved 6,000 iterations! Now we can see how much faster than Jacobi this is, because both methods are implemented the same way:
```
%%timeit
laplace2d_gauss_seidel(p0.copy(), p0.copy(), 1e-8)
```
We get some speed-up over the Numba version of Jacobi, but not a lot. And you may see quite different results—on some of the machines we tried, we could still not beat the NumPy version of Jacobi. This can be confusing, and hard to explain without getting into the nitty grity of hardware optimizations.
Don't lose hope! We have another trick up our sleeve!
## Successive Over-Relaxation (SOR)
Successive over-relaxation is able to improve on the Gauss-Seidel method by using in the update a linear combination of the previous and the current solution, as follows:
\begin{equation}
p^{k+1}_{i,j} = (1 - \omega)p^k_{i,j} + \frac{\omega}{4} \left(p^{k+1}_{i,j-1} + p^k_{i,j+1} + p^{k+1}_{i-1,j} + p^k_{i+1,j} \right)
\end{equation}
The relaxation parameter $\omega$ will determine how much faster SOR will be than Gauss-Seidel. SOR iterations are only stable for $0 < \omega < 2$. Note that for $\omega = 1$, SOR reduces to the Gauss-Seidel method.
If $\omega < 1$, that is technically an "under-relaxation" and it will be slower than Gauss-Seidel.
If $\omega > 1$, that's the over-relaxation and it should converge faster than Gauss-Seidel.
Let's write a function for SOR iterations of the Laplace equation, using Numba to get high performance.
```
@jit(nopython=True)
def laplace2d_SOR(p, pn, l2_target, omega):
'''Solves the Laplace equation using SOR with a 5-point stencil
Parameters:
----------
p: 2D array of float
Initial potential distribution
pn: 2D array of float
Allocated array for previous potential distribution
l2_target: float
Stopping criterion
omega: float
Relaxation parameter
Returns:
-------
p: 2D array of float
Potential distribution after relaxation
'''
iterations = 0
iter_diff = l2_target + 1 #initialize iter_diff to be larger than l2_target
denominator = 0.0
ny, nx = p.shape
l2_diff = numpy.zeros(20000)
while iter_diff > l2_target:
for j in range(ny):
for i in range(nx):
pn[j,i] = p[j,i]
iter_diff = 0.0
denominator = 0.0
for j in range(1,ny-1):
for i in range(1,nx-1):
p[j,i] = (1-omega)*p[j,i] + omega*.25 * (p[j,i-1] + p[j,i+1] + p[j-1,i] + p[j+1,i])
#Neumann 2nd-order BC
for j in range(1,ny-1):
p[j,-1] = .25 * (2*p[j,-2] + p[j+1,-1] + p[j-1, -1])
for j in range(ny):
for i in range(nx):
iter_diff += (p[j,i] - pn[j,i])**2
denominator += (pn[j,i]*pn[j,i])
iter_diff /= denominator
iter_diff = iter_diff**0.5
l2_diff[iterations] = iter_diff
iterations += 1
return p, iterations, l2_diff
```
That wasn't too bad at all. Let's try this out first with $\omega = 1$ and check that it matches the Gauss-Seidel results from above.
```
l2_target = 1e-8
omega = 1
p, iterations, l2_diffSOR = laplace2d_SOR(p0.copy(), p0.copy(), l2_target, omega)
print("Numba SOR method took {} iterations\
at tolerance {} with omega = {}".format(iterations, l2_target, omega))
```
We have the exact same number of iterations as Gauss-Seidel. That's a good sign that things are working as expected.
Now let's try to over-relax the solution and see what happens. To start, let's try $\omega = 1.5$.
```
l2_target = 1e-8
omega = 1.5
p, iterations, l2_diffSOR = laplace2d_SOR(p0.copy(), p0.copy(), l2_target, omega)
print("Numba SOR method took {} iterations\
at tolerance {} with omega = {}".format(iterations, l2_target, omega))
```
Wow! That really did the trick! We dropped from 13939 iterations down to 7108. Now we're really cooking! Let's try `%%timeit` on SOR.
```
%%timeit
laplace2d_SOR(p0.copy(), p0.copy(), l2_target, omega)
```
Things continue to speed up. But we can do even better!
### Tuned SOR
Above, we picked $\omega=1.5$ arbitrarily, but we would like to over-relax the solution as much as possible without introducing instability, as that will result in the fewest number of iterations.
For square domains, it turns out that the ideal factor $\omega$ can be computed as a function of the number of nodes in one direction, e.g., `nx`.
\begin{equation}
\omega \approx \frac{2}{1+\frac{\pi}{nx}}
\end{equation}
This is not some arbitrary formula, but its derivation lies outside the scope of this course. (If you're curious and have some serious math chops, you can check out Reference 3 for more information). For now, let's try it out and see how it works.
```
l2_target = 1e-8
omega = 2./(1 + numpy.pi/nx)
p, iterations, l2_diffSORopt = laplace2d_SOR(p0.copy(), p0.copy(), l2_target, omega)
print("Numba SOR method took {} iterations\
at tolerance {} with omega = {:.4f}".format(iterations, l2_target, omega))
```
Wow! That's *very* fast. Also, $\omega$ is very close to the upper limit of 2. SOR tends to work fastest when $\omega$ approaches 2, but don't be tempted to push it. Set $\omega = 2$ and the walls will come crumbling down.
Let's see what `%%timeit` has for us now.
```
%%timeit
laplace2d_SOR(p0.copy(), p0.copy(), l2_target, omega)
```
Regardless of the hardware in which we tried this, the tuned SOR gave *big* speed-ups, compared to the Jacobi method (whether implemented with NumPy or Numba). Now you know why we told you at the end of [Lesson 1](http://nbviewer.ipython.org/github/numerical-mooc/numerical-mooc/blob/master/lessons/05_relax/05_01_2D.Laplace.Equation.ipynb) that the Jacobi method is the *worst* iterative solver and almost never used.
Just to convince ourselves that everything is OK, let's check the error after the 1,110 iterations of tuned SOR:
```
L2_rel_error(p,pan)
```
Looking very good, indeed.
We didn't explain it in any detail, but notice the very interesting implication of Equation $(5)$: the ideal relaxation factor is a function of the grid size.
Also keep in mind that the formula only works for square domains with uniform grids. If your problem has an irregular geometry, you will need to find a good value of $\omega$ by numerical experiments.
## Decay of the difference between iterates
In the [Poisson Equation notebook](./05_02_2D.Poisson.Equation.ipynb), we noticed how the norm of the difference between consecutive iterations first dropped quite fast, then settled for a more moderate decay rate. With Gauss-Seidel, SOR and tuned SOR, we reduced the number of iterations required to reach the stopping criterion. Let's see how that reflects on the time history of the difference between consecutive solutions.
```
pyplot.figure(figsize=(8,8))
pyplot.xlabel(r'iterations', fontsize=18)
pyplot.ylabel(r'$L_2$-norm', fontsize=18)
pyplot.semilogy(numpy.trim_zeros(l2_diffJ,'b'),
'k-', lw=2, label='Jacobi')
pyplot.semilogy(numpy.trim_zeros(l2_diffGS,'b'),
'k--', lw=2, label='Gauss-Seidel')
pyplot.semilogy(numpy.trim_zeros(l2_diffSOR,'b'),
'g-', lw=2, label='SOR')
pyplot.semilogy(numpy.trim_zeros(l2_diffSORopt,'b'),
'g--', lw=2, label='Optimized SOR')
pyplot.legend(fontsize=16);
```
The Jacobi method starts out with very fast convergence, but then it settles into a slower rate. Gauss-Seidel shows a faster rate in the first few thousand iterations, but it seems to be slowing down towards the end. SOR is a lot faster to converge, though, and optimized SOR just plunges down!
## References
1. [Gonsalves, Richard J. Computational Physics I. State University of New York, Buffalo: (2011): Section 3.1 ](http://www.physics.buffalo.edu/phy410-505/2011/index.html)
2. Moin, Parviz, "Fundamentals of Engineering Numerical Analysis," Cambridge University Press, 2nd edition (2010).
3. Young, David M. "A bound for the optimum relaxation factor for the successive overrelaxation method." Numerische Mathematik 16.5 (1971): 408-413.
```
from IPython.core.display import HTML
css_file = '../../styles/numericalmoocstyle.css'
HTML(open(css_file, "r").read())
```
| github_jupyter |
# Day 0 Practical: Churn for Bank Customers
Welcome to the first practical session of the SPAI Advanced Machine Learning Workshop. In this practical, you will experience the full workflow of building a simple classifier to predict whether does a customer decides to leave the bank*(also known as churning)* given the features of the customer.
## Data Dictionary
| Features | Descriptions |
| :-- | :--- |
| RowNumber| corresponds to the record (row) number and has no effect on the output.|
| CustomerId|contains random values and has no effect on customer leaving the bank.|
|Surname | the surname of a customer has no impact on their decision to leave the bank.|
|CreditScore|can have an effect on customer churn, since a customer with a higher credit score is less likely to leave the bank.|
|Geography|a customer’s location can affect their decision to leave the bank.|
|Gender|it’s interesting to explore whether gender plays a role in a customer leaving the bank.|
|Age|this is certainly relevant, since older customers are less likely to leave their bank than younger ones.|
|Tenure|refers to the number of years that the customer has been a client of the bank. Normally, older clients are more loyal and less likely to leave a bank.|
|Balance|also a very good indicator of customer churn, as people with a higher balance in their accounts are less likely to leave the bank compared to those with lower balances.|
|NumOfProducts|refers to the number of products that a customer has purchased through the bank.|
|HasCrCard|denotes whether or not a customer has a credit card. This column is also relevant, since people with a credit card are less likely to leave the bank.|
|IsActiveMember|active customers are less likely to leave the bank.|
|EstimatedSalary|as with balance, people with lower salaries are more likely to leave the bank compared to those with higher salaries. |
|Exited|whether or not the customer left the bank.|
# Exercise 1️⃣
# Import Libraries
Before we starting injesting the data, it is always a good practice to first import all libraries that you will be using for the project. In this notebook we will be using some common Machine Learning libraries as listed below:
- `pandas` : Widely used for data analysis and manipulation.
- `numpy` : Provides high-performance mathematical, algebraic and transformation for multi-dimensional array and matrix data structures.
- `matplotlib.pyplot` : Data visualisation and plotting libraries
- `seaborn` : Data visualisation and plotting libraries
- `sklearn` : Simple and efficient tools for predictive modeling
- `yellowbrick` : Visualization library for evaluating models
## Task
1. Import `pandas`, `numpy` and `matplotlib.pyplot`, `seaborn`, `sklearn` libraries with their commonly-used convention/accronym.
```
'''
Hint:
1. Make use of "import ___ as ___" to import the libraries stated above with their accronym (pd, np, plt)
'''
### Write your code here ###
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import sklearn
### End ###
```
# Exercise 2️⃣
# Data Ingestion
Let us begin the journey by first importing the data from the `.csv` file from the kaggle website. For the sake of this exercise, the dataset have been uploaded to the Github Repository and is accessible via the link below.
## Task
1. Read the `.csv` files provided in the link below and assign it in a variable called `bank_df`
2. Show the first five row of the `bank_df` dataset
Churn for Bank Customers Link :
```
https://raw.githubusercontent.com/SPAI-Team/Advanced-ML-Workshop-2021/main/Datasets/churn.csv
```
```
'''
Hint:
1. Make use of pd.read_csv() method and parse the url as the parameter into the method and save the outcome in variable called bank_df.
2. Use .head() method to show the first five row of the dataset.
'''
### Write your code here ###
bank_df = pd.read_csv("https://raw.githubusercontent.com/SPAI-Team/Advanced-ML-Workshop-2021/main/Datasets/churn.csv")
bank_df.head()
### End ###
```
# Exploratory Data Analysis
In our attempt to better understand the dataset, we will be performing EDA by using `pandas` library for data manipulation and `matplotlib` & `seaborn` libraries for visualisation.
For the sake of the time in this workshop, we will be showing you the EDA process and code. If you are unclear about any of the methods used for EDA, do refer back to the Tutorial and Practical notebook from [SPAI Beginner Machine Learning Bootcamp 2021](https://github.com/Tien-Cheng/ML-Bootcamp-2021/tree/main/Beginner%20Machine%20Learning%20Bootcamp).
The following are a few questions that we attempt to investigate from our dataset:
1. What columns do we have and what are their datatypes?
2. How many records and features do we have in this dataset?
3. Is there any null values in the dataset?
4. What is the numerical summary for numerical features?
5. How is the data distribution for numerical features?
6. How is the data distribution for categorical features?
```
bank_df.info() # Shows the name and datatypes of the columns in the dataset
bank_df.shape # Show the number of records and number of features
bank_df.isnull().sum() # Show the number of null values in each columns
bank_df.describe().T # Show the numerical summaries
# Histogram of numerical columns
bank_df.hist(figsize=(12, 12))
plt.show()
# Count Plot on Geography
bank_df['Geography'].value_counts().plot(kind='barh')
plt.show()
# Count Plot on Gender
bank_df['Gender'].value_counts().plot(kind='barh')
plt.show()
```
# Data Preprocessing
Let us import the functions that we will be using for data preprocessing and data splitting from `sklearn` library.
```
# Sklearn Functions
# Preprocessing
from sklearn.preprocessing import StandardScaler, OneHotEncoder, OrdinalEncoder
from sklearn.impute import SimpleImputer
# Data Partition
from sklearn.model_selection import train_test_split
# Modelling
from sklearn.neighbors import KNeighborsClassifier
# Model Evaluation
from sklearn.metrics import f1_score
```
## Exercise 3️⃣
## Dropping Unnecessary Columns
Before we start splitting our data, let us first discard some columns that are not helpful for our prediction task. Those features are dropped as knowing the customer *'RowNumber', 'CustomerId' & 'Surname'* would not be useful for us to determine whether does the customer decides to churn.
### Task
1. Drop `['RowNumber', 'CustomerId', 'Surname']` from our original dataframe and assign it to variable `df_drop`.
```
'''
Hint:
1. Make use of .drop() methods and use columns parameter to specify the columns to drop
'''
### Write your code here ###
df_drop = bank_df.drop(columns = ['RowNumber', 'CustomerId', 'Surname'])
### End ###
df_drop
```
## Dropping Missing Values
Since there are no null values and anomalies in the dataset, we will not drop any rows and do any imputation to the value. In practice, you should always treat the null values by either dropping or imputation since most machine learning model couldn't handle missing values by default.
## Exercise 4️⃣
## Data Splitting : Train Test Split
To allow us to better gauge the performance of the model, we will be performing train-test-split towards our `df_drop` so that we can fit our model with the train set and evaluate it using the test set.
### Task
1. Identify and extract features and target variable, and save it to variable `X` and `y` accordingly
2. Make use of `train_test_split` function to split a train and test set with `test_size = 0.3` and `random_state = 42`
```
'''
Hint:
1. Make use of .drop( columns = <target_variable_name> ) to remove target variable and keep only features
2. Make use of column slicing with [ <target_variable_name> ] to extract target variable
'''
### Write your code here ###
X = df_drop.drop(columns = "Exited")
y = df_drop["Exited"]
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.3, random_state = 42)
### End ###
X_train
```
## Exercise 5️⃣
## Handling Numerical Features (Feature Scaling)
In this dataset, there are several continuous numerical features like *['CreditScore', 'Age', 'Balance', 'EstimatedSalary']* and discrete numerical features like *['Tenure', 'NumOfProducts']*
Since the discrete numerical features are encoded nicely without the need of further manipulation, the only preprocessing steps we could take is to perform **Feature Scaling** to the continuous numerical features. We will be using `StandardScaler` from `sklearn.preprocessing` to complete the task below.
### Task
1. Create variable with list of numerical features to perform standard scaling `['CreditScore', 'Age', 'Balance', 'EstimatedSalary']` and store it in variable `num_col`
2. Initialise `StandardScaler()` class imported from sklearn library and save the instance in variable `scaler`
3. Perform fit transform with the feature scaler to the numerical columns in `X_train` and replace the result to the numerical columns in `X_train_scaled`
4. Make use of the same feature scaler and transform the numerical columns in `X_test` and replace the result to the numerical columns in `X_test_scaled`
```
'''
Hint:
1. Define a list named num_col with all the numerical columns mentioned above
2. Initiate the StandardScaler() object and store it in variable scaler
3. Use .fit_transform() from scaler to X_train[num_col] and stored the result in X_train_scaled[num_col]
4. Use .trasform() from scaler to X_test[num_col] and stored the result in X_test_scaled[num_col]
'''
X_train_scaled = X_train.copy()
X_test_scaled = X_test.copy()
### Write your code here ###
num_col = ['CreditScore', 'Age', 'Balance', 'EstimatedSalary']
scaler = StandardScaler()
X_train_scaled[num_col] = scaler.fit_transform(X_train[num_col])
X_test_scaled[num_col] = scaler.transform(X_test[num_col])
### End ###
X_train_scaled[num_col]
```
## Exercise 6️⃣
## Handling Categorical Features (Nominal Encoding)
For nominal data, we need to convert value into 1 or 0 values to the column. This is to allow our model to better understand the categorical variable since it could not interpret a string value without any feature encoding. In this practical, we will be using `pd.get_dummies` to perform the nominal encoding but in subsequent days, you will be exposed to using `OneHotEncoder` from `sklearn.preprocessing` in a Pipeline.
### Task
1. List out all Normial Categorical Column `['Geography','Gender']` and store it in variable `cat_col`
2. Create dummy variable for `X_train_scaled` with `.get_dummies()` method by parsing parameter `(X_train_scaled, columns=cat_col, drop_first=True)` and assign it to `X_train_onehot`
3. Create dummy variable for `X_test_scaled` with `.get_dummies()` method by parsing parameter `(X_test_scaled, columns=cat_col, drop_first=True)` and assign it to `X_test_onehot`
```
'''
Hint:
1. Define a list named cat_col with all the nominal categorical columns mentioned above
2. Use pd.get_dummies() with parameters mentioned above and store the outcome in X_train_onehot
3. Use pd.get_dummeis() with parameters mentioned above and store the outcome in X_test_onehot
'''
### Write your code here ###
cat_col = ['Geography','Gender']
X_train_onehot = pd.get_dummies(X_train_scaled, columns=cat_col, drop_first=True)
X_test_onehot = pd.get_dummies(X_test_scaled, columns=cat_col, drop_first=True)
### End ###
X_train_onehot
```
# Exercise 7️⃣
# Model Building and Evaluation
Our data is now ready to be parse into our machine learning model. Let us continue to build a simple K-Nearest Neighbour model and predict whether does a customer decides to leave the bank.
5 Steps of Model Fitting and Prediction with Scikit-Learn Library:
- Import model class from Sklearn library
- Initiate the class with hyperparameters and store the instance at a variable
- Call .fit() method to train the model by parsing X_train and y_train
- Call .predict() method with the trained model to predict features, X_test
- Evaluate the performance of model by comparing the prediction with ground truth, y_test
## Task
1. Initiate the `KNeighborsClassifier` class with `n_neighbors=5` as hyperparameter and save it as variable `knn`
1. Fit the model with `X_train_onehot` and `y_train`
1. Predict the label of `X_test_onehot` and save it as variable `prediction`
1. Compare `prediction` and `y_test` with `f1_score` function from `sklearn.metrics` and save the result at variable `f1`.
```
### Write your code here ###
knn = KNeighborsClassifier(n_neighbors=5)
knn.fit(X_train_onehot, y_train)
prediction = knn.predict(X_test_onehot)
f1 = f1_score(y_test, prediction)
### End ###
print(f"F1 Score for {KNeighborsClassifier.__name__} : {f1}") # Print out the score
```
# Conclusion
Congratulations! You have completed Practical Notebook for Day 0 of SPAI Advanced Machine Learning Workshop. If you have any doubts or require any clarification, feel free to approach us us through our [Instagram](https://www.instagram.com/spai.sp/) or [Discord Server](https://discord.gg/zPYJMGfQFa)*(remember to verify yourself)* or revise the material from Beginner Machine Learning Bootcamp.
Meeting Recordings
- OneDrive : https://ichatspedu.sharepoint.com/:f:/t/SPAI/Er059biGhKJGlPpqMNcWGxIBX7ynLf5fi6jCEO3suhijYg?e=XG3E8U
Lesson Material:
- Google Drive : https://drive.google.com/drive/u/1/folders/1jfDncbT7Vl7X8U-SYs0djJ3i449uWLGz
- GitHub Repository : https://github.com/SPAI-Team/ML-Bootcamp-2021/tree/main/Beginner%20Machine%20Learning%20Bootcamp
---
> ### Feel Free to Join the Subsequent **SPAI Machine Learning Competition** to earn **valuable Prize💰 and Experience🏆** if you have not done so!🥳🥳
[ML Comp Sign Up Link](https://docs.google.com/forms/d/e/1FAIpQLSchLDXDAY0LqM6fuRDyQwdRNbVT4FYrgDtqthEIfYpFvpWMAg/viewform) *(If it has not been closed)*
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/NAIP/loop_FeatureCollection.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/NAIP/loop_FeatureCollection.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/NAIP/loop_FeatureCollection.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API and geemap
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geemap](https://github.com/giswqs/geemap). The **geemap** Python package is built upon the [ipyleaflet](https://github.com/jupyter-widgets/ipyleaflet) and [folium](https://github.com/python-visualization/folium) packages and implements several methods for interacting with Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, and `Map.centerObject()`.
The following script checks if the geemap package has been installed. If not, it will install geemap, which automatically installs its [dependencies](https://github.com/giswqs/geemap#dependencies), including earthengine-api, folium, and ipyleaflet.
**Important note**: A key difference between folium and ipyleaflet is that ipyleaflet is built upon ipywidgets and allows bidirectional communication between the front-end and the backend enabling the use of the map to capture user input, while folium is meant for displaying static data only ([source](https://blog.jupyter.org/interactive-gis-in-jupyter-with-ipyleaflet-52f9657fa7a)). Note that [Google Colab](https://colab.research.google.com/) currently does not support ipyleaflet ([source](https://github.com/googlecolab/colabtools/issues/60#issuecomment-596225619)). Therefore, if you are using geemap with Google Colab, you should use [`import geemap.eefolium`](https://github.com/giswqs/geemap/blob/master/geemap/eefolium.py). If you are using geemap with [binder](https://mybinder.org/) or a local Jupyter notebook server, you can use [`import geemap`](https://github.com/giswqs/geemap/blob/master/geemap/geemap.py), which provides more functionalities for capturing user input (e.g., mouse-clicking and moving).
```
# Installs geemap package
import subprocess
try:
import geemap
except ImportError:
print('geemap package not installed. Installing ...')
subprocess.check_call(["python", '-m', 'pip', 'install', 'geemap'])
# Checks whether this notebook is running on Google Colab
try:
import google.colab
import geemap.eefolium as geemap
except:
import geemap
# Authenticates and initializes Earth Engine
import ee
try:
ee.Initialize()
except Exception as e:
ee.Authenticate()
ee.Initialize()
```
## Create an interactive map
The default basemap is `Google Maps`. [Additional basemaps](https://github.com/giswqs/geemap/blob/master/geemap/basemaps.py) can be added using the `Map.add_basemap()` function.
```
Map = geemap.Map(center=[40,-100], zoom=4)
Map
```
## Add Earth Engine Python script
```
# Add Earth Engine dataset
year = 2015
collection = ee.ImageCollection('USDA/NAIP/DOQQ')
startTime = ee.Date(str(year) + '-01-01')
endTime = ee.Date(str(year) + '-12-31')
# year = startTime.get('year').getInfo()
# print(year)
fromFT = ee.FeatureCollection('ft:1CLldB-ULPyULBT2mxoRNv7enckVF0gCQoD2oH7XP')
# count = fromFT.size().getInfo()
# print(count)
polys = fromFT.geometry()
centroid = polys.centroid()
lng, lat = centroid.getInfo()['coordinates']
# print("lng = {}, lat = {}".format(lng, lat))
values = fromFT.reduceColumns(ee.Reducer.toList(2), ['system:index', 'name']).getInfo()['list']
# print(values)
Map.setCenter(lng, lat, 10)
def subsetNAIP(img_col, startTime, endTime, fc):
img = img_col.filterDate(startTime, endTime).filterBounds(fc).mosaic().clip(fc)
return img
def calNDWI(image):
"""A function to compute NDWI."""
ndwi = image.normalizedDifference(['G', 'N'])
ndwiViz = {'min': 0, 'max': 1, 'palette': ['00FFFF', '0000FF']}
ndwiMasked = ndwi.updateMask(ndwi.gte(0.2))
ndwi_bin = ndwiMasked.gt(0)
patch_size = ndwi_bin.connectedPixelCount(500, True)
large_patches = patch_size.eq(500)
large_patches = large_patches.updateMask(large_patches)
opened = large_patches.focal_min(1).focal_max(1)
return opened
def rasterToVector(img, fc):
vec = img.reduceToVectors(geometry=fc, eightConnected=True, maxPixels=59568116121, crs=img.projection(), scale=1)
return vec
def exportToDrive(vec, filename):
taskParams = {
'driveFolder': 'image',
'fileFormat': 'KML'
}
task = ee.batch.Export.table(vec, filename, taskParams)
task.start()
vis = {'bands': ['N', 'R', 'G']}
for (id, name) in values:
watershed = fromFT.filter(ee.Filter.eq('system:index', str(id)))
filename = "Y" + str(year) + "_" + str(id) + "_" + str(name).replace(" ", "_")
print(filename)
image = subsetNAIP(collection, startTime, endTime, watershed)
ndwi = calNDWI(image)
vector = rasterToVector(ndwi, watershed)
exportToDrive(vector, filename)
# Map.addLayer(image, vis)
# Map.addLayer(vector)
# for i in range(2, 2 + count):
# watershed = fromFT.filter(ee.Filter.eq('system:index', str(i)))
# re = fc.filterBounds(watershed)
# task = ee.batch.Export.table(re, 'watershed-' + str(i), taskParams)
# task.start()
#
#
#
# lng_lat = ee.Geometry.Point(lng, lat)
# naip = collection.filterBounds(polys)
# naip_2015 = naip.filterDate('2015-01-01', '2015-12-31')
# ppr = naip_2015.mosaic()
#
# count = naip_2015.size().getInfo()
# print("Count: ", count)
#
# # print(naip_2015.size().getInfo())
# vis = {'bands': ['N', 'R', 'G']}
# Map.setCenter(lng, lat, 12)
# Map.addLayer(ppr,vis)
# # Map.addLayer(polys)
#
# def NDWI(image):
# """A function to compute NDWI."""
# ndwi = image.normalizedDifference(['G', 'N'])
# ndwiViz = {'min': 0, 'max': 1, 'palette': ['00FFFF', '0000FF']}
# ndwiMasked = ndwi.updateMask(ndwi.gte(0.05))
# ndwi_bin = ndwiMasked.gt(0)
# patch_size = ndwi_bin.connectedPixelCount(500, True)
# large_patches = patch_size.eq(500)
# large_patches = large_patches.updateMask(large_patches)
# opened = large_patches.focal_min(1).focal_max(1)
# return opened
#
# ndwi_collection = naip_2015.map(NDWI)
# # Map.addLayer(ndwi_collection)
# # print(ndwi_collection.getInfo())
#
# # downConfig = {'scale': 10, "maxPixels": 1.0E13, 'driveFolder': 'image'} # scale means resolution.
# # img_lst = ndwi_collection.toList(100)
# #
# # taskParams = {
# # 'driveFolder': 'image',
# # 'driveFileNamePrefix': 'ndwi',
# # 'fileFormat': 'KML'
# # }
# #
# # for i in range(0, count):
# # image = ee.Image(img_lst.get(i))
# # name = image.get('system:index').getInfo()
# # print(name)
# # # task = ee.batch.Export.image(image, "ndwi2-" + name, downConfig)
# # # task.start()
#
# mosaic = ndwi_collection.mosaic().clip(polys)
# fc = mosaic.reduceToVectors(eightConnected=True, maxPixels=59568116121, crs=mosaic.projection(), scale=1)
# # Map.addLayer(fc)
# taskParams = {
# 'driveFolder': 'image',
# 'driveFileNamePrefix': 'water',
# 'fileFormat': 'KML'
# }
#
# count = fromFT.size().getInfo()
# Map.setCenter(lng, lat, 10)
#
# for i in range(2, 2 + count):
# watershed = fromFT.filter(ee.Filter.eq('system:index', str(i)))
# re = fc.filterBounds(watershed)
# # task = ee.batch.Export.table(re, 'watershed-' + str(i), taskParams)
# # task.start()
# # Map.addLayer(fc)
#
#
# # lpc = fromFT.filter(ee.Filter.eq('name', 'Little Pipestem Creek'))
```
## Display Earth Engine data layers
```
Map.addLayerControl() # This line is not needed for ipyleaflet-based Map.
Map
```
| github_jupyter |
# Aplicação: cores PANTONE
## Leitura de arquivos _json_
```
import os, json
# diretório base
base = '../database/pantone-colors/'
for fi in os.listdir(base):
n,e = os.path.splitext(fi)
if e == '.json':
with open(os.path.join(base,fi), 'r') as f:
# define variáveis dinamicamente
exec(f'{n} = {json.load(f)}')
# verifica se variáveis foram criadas
%who
```
### Desempacotar sequencia em variáveis separadas
```
# no. elementos da lista
len(pantone_coated)
# cada elemento é um dict
pantone_coated[0:3],pantone_metallic[-4:-1]
# desempacotamento apenas de valores
pname,hexcolor = pantone_coated[0].values()
```
- Atualizar dicts com códigos de cores rgb
```
# função para converter hex para rgb
def hex2rgb(hexcolor):
h = hexcolor.lstrip('#')
rgb = tuple(int(h[i:i+2],16) for i in (0,2,4))
return rgb
# atualiza dict com rgb
for c in range(len(pantone_coated)):
d = pantone_coated[c]
pname,hexcolor = d.values()
rgb = hex2rgb(hexcolor)
pantone_coated[c].update({'rgb':rgb})
for c in range(len(pantone_metallic)):
d = pantone_metallic[c]
pname,hexcolor = d.values()
rgb = hex2rgb(hexcolor)
pantone_metallic[c].update({'rgb':rgb})
```
**Exercício:** Defina uma função `update_colors(clist)` que recebe um dict `clist` de cores Pantone e o atualiza para conter a chave `'rgb'`.
## Desempacotar elementos em iteráveis de comprimento arbitrário
Suponha que o dict estivesse organizado de tal forma que o nome e os códigos HEX e RGB de uma certa cor estivessem sequenciados em uma tuple ou list:
```
ex = pantone_coated[0]
ex = ex['pantone'],ex['hex'], ex['rgb']
ex
```
O desempacotamento das cores poderia ser feito por meio de uma _expressão estrela_ (_star expression_):
```
# desempacota por star expression e ignora primeiro item
_,*pcolors = ex
pcolors
```
Note que usamos 2 variáveis para desempacotar uma tuple de 3 elementos. Outros casos em que _star expressions_ são úteis envolvem dados que possuem mais de um valor atribuível a um mesmo registro (p.ex.: 1 pessoa com 2 números de telefone), ou quando se quer quebrar iteráveis em comprimentos arbitrários. Por exemplo:
```{hint}
Use um _underscore_ (`_`) para representar uma variável que, devido ao desempacotamento, você considere "indesejada".
```
```
_,*m4,_ = [3,4,8,12,16,10]
m4 # múltiplos de 4
_,_,_,*m5 = [4,8,12,5,10]
m5 # múltiplos de 5
*m6,_,_ = [6,12,5,10]
m6 # múltiplos de 6
# 2 star expressions não são permitidas
*m6,*m5 = [6,12,5,10]
```
```{note}
Ao usar uma _star expression_, certifique-se que o número de variáveis usadas no desempacotamento é consistente com os seus objetivos.
```
## Localizar os maiores ou menores N itens em uma coleção
No espaço vetorial RGB, as cores variam do vetor (0,0,0), branco, ao vetor (255,255,255), preto. Em um cubo unitário, podemos trabalhar com valores "normalizados". Para isso, basta dividir todos os valores nas tuplas por 255, de modo que o maior vetor seja (1,1,1).
Em linhas gerais, o espaço de cores RGB é uma região do espaço tridimensional limitada definida por:
$$V = \{(r,g,b) \ : \ 0 \leq r,g,b \leq 1\}$$
Assim, vamos construir um dict com cores normalizadas para utilizarmos mais à frente.
```
# indexa cores
c_pc = {i:c['rgb'] for i,c in enumerate(pantone_coated)}
# normaliza
for i,v in c_pc.items():
r,g,b = v
c_pc[i] = (r/255,g/255,b/255)
```
Suponhamos que nosso interesse seja saber quais são as _N_ maiores ou menores pigmentações de vermelho em uma fatia do dict `pantone_coated`. Vejamos duas formas de fazer isso:
- Usando `sorted`:
```
# pigmentações
r = [i[0] for i in c_pc.values()][1200:1211]
# 10 pigmentações
N = 10
# ordena
r1 = sorted(r)
# ordena em modo reverso
r2 = sorted(r,reverse=True)
# menores
print(r1[:N],'\n')
# maiores
print(r2[:N])
```
- Usando o módulo `heapq`
```
import heapq
# N menores
r3 = heapq.nsmallest(N,r)
print(r3,'\n')
# N maiores
r4 = heapq.nlargest(N,r)
print(r4)
```
Entretanto, é útil pensar em como buscar pelas tonalidades de cor mais claras ou mais escuras ordenando-as pelos valores das pigmentações individualmente usando `sorted` e _key functions_.
A seguir, usamos permutações de $(r,g,b)$ para mostrar como podemos fazer múltiplas ordenações.
```
from operator import itemgetter
# ordena por R, depois G, então B
sorted_rgb = sorted(c_pc.values(),key=itemgetter(0,1,2),reverse=True)
# ordena por G, depois B, então R
sorted_gbr = sorted(c_pc.values(),key=itemgetter(1,2,0),reverse=True)
# ordena por B, depois R, então G
sorted_brg = sorted(c_pc.values(),key=itemgetter(2,0,1),reverse=True)
# ordena por R, depois B, então G
sorted_rbg = sorted(c_pc.values(),key=itemgetter(0,2,1))
# ordena por G, depois R, então B
sorted_grb = sorted(c_pc.values(),key=itemgetter(1,0,2))
# ordena por B, depois G, então R
sorted_bgr = sorted(c_pc.values(),key=itemgetter(2,1,0))
```
Agora, plotamos paletas com as _k_ primeiras cores dos dicts anteriores:
```
import matplotlib.pyplot as plt
def plotKColors(d,k,t):
'''
Plota paleta simples de k primeiras cores
do dict d com título t.
'''
_, ax = plt.subplots(1,k,figsize=(10,1),facecolor=None)
for i in range(k):
ax[i].plot(0.5,0.5,'s',ms=20,c=d[i])
ax[i].axis('off')
plt.title(t)
# k = 30
plotKColors(sorted_rgb,30,'RGB claro -> escuro')
plotKColors(sorted_gbr,30,'GBR claro -> escuro')
plotKColors(sorted_brg,30,'BRG claro -> escuro')
plotKColors(sorted_rbg,30,'RBG escuro -> claro')
plotKColors(sorted_grb,30,'GRB escuro -> claro')
plotKColors(sorted_bgr,30,'BGR escuro -> claro')
```
| github_jupyter |
<div class="contentcontainer med left" style="margin-left: -50px;">
<dl class="dl-horizontal">
<dt>Title</dt> <dd> RGB Element</dd>
<dt>Dependencies</dt> <dd>Matplotlib</dd>
<dt>Backends</dt>
<dd><a href='./RGB.ipynb'>Matplotlib</a></dd>
<dd><a href='../bokeh/RGB.ipynb'>Bokeh</a></dd>
<dd><a href='../plotly/RGB.ipynb'>Plotly</a></dd>
</dl>
</div>
```
import numpy as np
import holoviews as hv
from holoviews import opts
hv.extension('matplotlib')
```
``RGB`` represents a regularly spaced 2D grid of an underlying continuous space of RGB(A) (red, green, blue and alpha) color space values. The definition of the grid closely matches the semantics of an Image and in the simplest case the grid may be specified as a ``NxMx3`` or ``NxMx4`` array of values along with a bounds, but it may also be defined through explicit and regularly spaced x/y-coordinate arrays. The two most basic supported constructors of an ``RGB`` element therefore include:
RGB((X, Y, R, G, B))
where ``X`` is a 1D array of shape ``M``, ``Y`` is a 1D array of shape ``N`` and ``R``/``G``/``B`` are 2D array of shape ``NxM``, or equivalently:
RGB(Z, bounds=(x0, y0, x1, y1))
where Z is a 3D array of stacked R/G/B arrays with shape NxMx3/4 and the bounds define the (left, bottom, right, top) edges of the four corners of the grid. Other gridded formats which support declaring of explicit x/y-coordinate arrays such as xarray are also supported. See the [Gridded Datasets](../../../user_guide/09-Gridded_Datasets.ipynb) user guide for all the other accepted data formats.
One of the simplest ways of creating an ``RGB`` element is to load an image file (such as PNG) off disk, using the ``load_image`` classmethod:
```
hv.RGB.load_image('../assets/penguins.png')
```
If you have ``PIL`` or [``pillow``](https://python-pillow.org) installed, you can also pass in a PIL Image as long as you convert it to Numpy arrays first:
```
from PIL import Image
hv.RGB(np.array(Image.open('../assets/penguins.png')))
```
This Numpy-based method for constructing an ``RGB`` can be used to stack up arbitrary 2D arrays into a color image:
```
x,y = np.mgrid[-50:51, -50:51] * 0.1
r = 0.5*np.sin(np.pi +3*x**2+y**2)+0.5
g = 0.5*np.sin(x**2+2*y**2)+0.5
b = 0.5*np.sin(np.pi/2+x**2+y**2)+0.5
hv.RGB(np.dstack([r,g,b]))
```
You can see how the RGB object is created from the original channels:
```
opts.defaults(opts.Image(cmap='gray'))
hv.Image(r,label="R") + hv.Image(g,label="G") + hv.Image(b,label="B")
```
``RGB`` also supports an optional alpha channel, which will be used as a mask revealing or hiding any ``Element``s it is overlaid on top of:
```
mask = 0.5*np.sin(0.2*(x**2+y**2))+0.5
rgba = hv.RGB(np.dstack([r,g,b,mask]))
bg = hv.Image(0.5*np.cos(x*3)+0.5, label="Background") * hv.VLine(x=0,label="Background")
overlay = (bg*rgba).relabel("RGBA Overlay")
bg + hv.Image(mask,label="Mask") + overlay
```
One additional way to create RGB objects is via the separate [ImaGen](https://github.com/pyviz-topics/imagen) library, which creates parameterized streams of images for experiments, simulations, or machine-learning applications.
For full documentation and the available style and plot options, use ``hv.help(hv.RGB).``
| github_jupyter |
## pyHail MESH Animation
This code utilizes the pyHAIL package to plot MESH, or the "maximum expected size of hail", grid the plots, and then create an animation with the plots.
```
from __future__ import print_function
import warnings
import warnings
warnings.filterwarnings('ignore')
"""
MESH sub-module of pyhail
Contains the single pol MESH retrieval for gridded radar data.
Required reflectivity and temperature data.
Joshua Soderholm - 15 June 2018
"""
import os
import netCDF4
import numpy as np
import pyart
import pyhail as ph
from pyhail import common
from pyhail import mesh
from pylab import *
import pyart, boto3, tempfile, os, shutil, datetime, matplotlib
import cartopy.crs as ccrs
import cartopy.feature as cfeature
import pyart
import matplotlib.pyplot as plt
import matplotlib
%matplotlib inline
from time import time
from datetime import datetime
from dateutil import tz
import os
import cartopy.crs as ccrs
import matplotlib.colors as colors
import cartopy.io.shapereader as shpreader
from skewt import SkewT
import glob
from glob import glob
from botocore.handlers import disable_signing
from matplotlib.animation import FuncAnimation
# from cpol_processing import processing as cpol_prc
from pyhail import hsda, hdr, mesh, common
# Obtaining radar scans...
def get_radar_scan(station='KLOT', date=None, key_index=-20):
'''
Function will pull the latest radar scan from any radar site using
Amazon S3.
----------
Station = Four letter NEXRAD identifier
Example: 'KEPZ'
Date = default is none for current date, else enter date in format "YYYY/MM/DD"
Ex: date ='2013/11/17
Key_index = Number of keys you want pulled from most recent scan.
Ex: key_index = -15 would pull ht most recent 15 scans
'''
# Creating a bucket and a client to be able to pull data from AWS and setting it as unsigned
bucket = 'noaa-nexrad-level2'
s3 = boto3.resource('s3')
s3.meta.client.meta.events.register('choose-signer.s3.*', disable_signing)
# Connects the bucket create above with radar data
aws_radar = s3.Bucket(bucket)
# Setting the date and time to current...
# This will allow for allow the current date's radar scands to be pulled
if date == None:
target_string = datetime.datetime.utcnow().strftime('%Y/%m/%d/'+station)
else:
target_string = date+'/'+station
for obj in aws_radar.objects.filter(Prefix= target_string):
'{0}:{1}'.format(aws_radar.name, obj.key)
my_list_of_keys = [this_object.key for this_object in aws_radar.objects.filter(Prefix= target_string)]
keys = my_list_of_keys[key_index:]
for key in keys:
if 'MDM' in key:
keys.remove(key)
print(keys)
return aws_radar, keys
aws_radar, keys = get_radar_scan(station='KLOT', date='2019/05/27', key_index=-400)
out_path_dir = 'home/amedendorp/Desktop/april182013'
nk = keys[175:210] #:210
nk
localfile = tempfile.NamedTemporaryFile()
aws_radar.download_file(keys[0], localfile.name)
radar = pyart.io.read(localfile.name)
radar.fields.keys()
# Turning the data into grid data and saving it to a folder...
# If the grids are already created, there is no need to run this code block again.
def get_grid(aws_radar, nk):
localfile = tempfile.NamedTemporaryFile()
aws_radar.download_file(nk, localfile.name)
radar = pyart.io.read(localfile.name)
# Create rainfall rate field
# Mask out last 10 gates of each ray, this removes the "ring" around the radar.
radar.fields['reflectivity']['data'][:, -10:] = np.ma.masked
gatefilter = pyart.filters.GateFilter(radar)
gatefilter.exclude_transition()
gatefilter.exclude_masked('reflectivity')
grid = pyart.map.grid_from_radars(
(radar, ), grid_shape=(16, 300, 300),
grid_limits=((0, 15000), (-123000.0, 123000.0), (-123000.0, 123000.0)),
fields=['reflectivity'], weighting_function='Barnes2',
gridding_algo='map_gates_to_grid',
h_factor=0., nb=0.6, bsp=1., min_radius=500., gatefilters=(gatefilter, ))
del radar
return grid
for num,key in enumerate(nk):
print('saving grid', num)
grid = get_grid(aws_radar, key)
name = os.path.join('/home/amedendorp/Desktop/MESH/MESH_grid_' + str(num).zfill(3) + '.nc')
pyart.io.write_grid(name, grid)
del grid
# If the code encounters a .tar file or any other unknown file, it will stop running.
# Every grid created before that will be preserved.
from glob import glob
files = glob('/home/amedendorp/Desktop/MESH/MESH_grid_*')
files.sort()
reader = shpreader.Reader('/home/amedendorp/Downloads/countyl010g_shp_nt00964/countyl010g.shp')
counties = list(reader.geometries())
COUNTIES = cfeature.ShapelyFeature(counties, ccrs.PlateCarree())
# This code was created using a modified version of pyART. The only thing that will change versus default
# pyART is the thickness and color of the lat and lon lines, and the county and state outlines.
def rr_animation(nframe):
plt.clf()
nfile = files[nframe]
radar = pyart.io.read_grid(nfile)
# Converting the default UTC time to local time...
# Converts to 24-hour time. No AM or PM.
utc = netCDF4.num2date(radar.time['data'][0],
radar.time['units'])
print(str(utc))
z = datetime.strptime(str(utc), '%Y-%m-%d %H:%M:%S.%f')
from_zone = tz.tzutc()
to_zone = tz.tzlocal()
z = z.replace(tzinfo=from_zone)
central = z.astimezone(to_zone)
t = datetime.strftime(central, '%Y-%m-%dT%H:%M:%S.%f')
title = ('KLOT ' + str(radar.z['data'][0]/1000) + ' km ' + t + ' \n'
+ ' Maximum Expected Size of Hail')
hail = mesh.main(grid=radar, ref_name='reflectivity',
snd_input='/home/amedendorp/Desktop/Sounding.nc',
sonde_temp='temp', sonde_height='height',
out_ffn=nfile)
projection = ccrs.PlateCarree()
ax = plt.axes(projection=projection)
# Plot site locations...
ANL_lon, ANL_lat = -87.981810, 41.713969
NW_lon, NW_lat = -87.675885, 42.057888
Naperville_lon, Naperville_lat = -88.181798, 41.738107
IBP_lon, IBP_lat = -87.687151, 41.606367
plt.plot([ANL_lon], [ANL_lat], color='black', marker= '.')
plt.plot([NW_lon], [NW_lat], color='black', marker= '.')
plt.plot([Naperville_lon], [Naperville_lat], color='black', marker= '.')
plt.plot([IBP_lon], [IBP_lat], color='black', marker= '.')
# Plot names of sites:
plt.text(ANL_lon + 0.01, ANL_lat - 0., 'ANL', horizontalalignment='left')
plt.text(NW_lon - 0.01, NW_lat - 0, 'Northwestern', horizontalalignment='right')
plt.text(Naperville_lon - 0.01, Naperville_lat + 0.01, 'Naperville', horizontalalignment='left')
plt.text(IBP_lon - 0.01, IBP_lat + 0.01, 'IBP', horizontalalignment='left')
display = pyart.graph.GridMapDisplay(hail)
display.plot_grid('MESH', level= 0, lat_lines=np.arange(41, 43, .5),
lon_lines=np.arange(-89, -86.5, .5), cmap='hot_r', vmax=55, vmin=0)
plt.rcParams.update({'axes.titlesize': '18'})
del radar, display
ax.add_feature(COUNTIES, facecolor='none', edgecolor='gray')
ax.add_feature(cfeature.LAKES, zorder=.5)
fig = plt.figure(figsize=[12,7])
# Match the frames to the amount of grids
sat_anim = FuncAnimation(fig, rr_animation, frames=34)
sat_anim.save('/home/amedendorp/Desktop/pyhailanimtest2.gif',
writer='imagemagick', fps=3)
plt.close
```
| github_jupyter |
# MinPy (MXNet NumPy)
*"Everybody loves NumPy."*
In this tutorial, we present MinPy -- a NumPy-like package based on MXNet. NumPy is a well-known python package widely used in scientific computing, statistics and machine learning. It supports a wide range of tensor operators and is very friendly to machine learning beginners. Here, we will walk with you through a simple example of *logistic regression* using NumPy. After that we will illustrate how MinPy utilizes GPUs to speed up your machine learning algorithm with few (or no) changes in your original NumPy code. We hope by following this direction, we could solve the ultimate trade-off that every machine learning tools are facing with: *how to achieve both efficiency and flexibility?*
```
# A bit of setup, just ignore this cell.
from __future__ import absolute_import
from __future__ import print_function
import matplotlib.pyplot as plt
# For auto-reloading external modules.
%load_ext autoreload
%autoreload 2
%matplotlib inline
# Set default size of plots.
plt.rcParams['figure.figsize'] = (8.0, 6.0)
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
```
*Before start:* MinPy is still a work in progress, so you may encounter problems when playing with it. You could always try *Kernel->Restart & Clear Output* and go through the tutorial again.
## Multinomial Logistic Regression using NumPy
Logistic regression is a supervised machine learning algorithm that is commonly used in classification. The *supervised* word here means each data sample (represented by a float vector) is *labeled*. We have provided some util functions to fetch some generated data for you.
```
import numpy as np
import numpy.random as random
import time
from util import get_data, plot_data
# Initialize training data.
data, label = get_data()
num_samples = data.shape[0]
num_features = data.shape[1]
num_classes = label.shape[1]
print('Shapes: data {}, label {}'.format(data.shape, label.shape))
print('#samples: {}'.format(num_samples))
print('#features: {}'.format(num_features))
print('#classes: {}'.format(num_classes))
```
In our example, the training data contains 10000 samples. Each sample has 500 features, so the training samples could be represented by a 10000 x 500 matrix. Our goal is to classify each sample into 5 classes. Therefore, each sample comes with a label which is encoded in *one-of-many* format. For example, say sample 0 is of class 1, then the corresponding label vector for sample 0 will be `[0, 1, 0, 0, 0]`. Therefore, label could be represented by a 10000 x 5 matrix as follows:
```
print('Data matrix:')
print(data)
print()
print('Label matrix:')
print(label)
```
We provide a function `plot_data` for you to visualize what the data looks like. It projects each 500-D samples onto their first two dimensions, so each sample is a point in the figure. We distinguish samples of different classes using different colors. Although this is a very simple example, the idea behind is similar to more complicated examples you will encounter in the following tutorials. For example, if this is an *Optical Character Recognition (OCR)* task (like the MNIST example we are going to talk about), you could think about the blue points are the samples of digit 0 and red points are digit 1. Then our goal is to correctly classify some *unknown* samples (test data) into the most likely classes.
```
# Visualize what the ground truth data looks like.
print('Ground truth:')
plot_data(data, label)
```
The model we are using here is called *Multinomial Logistic Regression*, or called *Softmax Regression*. Original logistic regression could only deal with binary classes (either 0 or 1) while multinomial logistic regression allows you to classify samples into multiple classes. The model contains a weight matrix `w` that projects from feature space to class space (so its dimension is $\#feature \times \#class$). It first calculate the probability of each class given the samples:
$$p(y^{(i)}=j \mid x^{(i)};w)= \frac{\exp(w_j^Tx^{(i)})}{\sum_k\exp(w_k^Tx^{(i)})}.$$
Try implement the following `predict` function that corresponds to the above math equation.
```
# Predict the class using logistic regression.
def predict(w, x):
#===========================================================#
# Your code starts here #
#===========================================================#
# Please try compute the estimated probability of each sample.
a = np.exp(np.dot(x, w))
a_sum = np.sum(a, axis=1, keepdims=True)
prob = a / a_sum
#===========================================================#
# Your code ends here #
#===========================================================#
return prob
```
We could then use the above `predict` function to see what a random guess may look like. We first initialize the `weight` matrix using random numbers. Then feed the predict result into the `plot_data` function for visualization. You could see that, the random guess is completely different than the correct labels.
```
# Initialize training weight.
weight = random.randn(num_features, num_classes)
# Visualize the initial guess.
plot_data(data, predict(weight, data))
```
Multinomial logistic regression then tries to minimize the *cross entropy* between the `predict` class probability and the correct probability. More specifically, it tries to minimize the following equation:
$$J(w)=-\frac{1}{m}\left[\sum_{i=1}^{m}\sum_{j=1}^{k}1\{y^{(i)}=j\}\log (p(y^{(i)}=j \mid x^{(i)};w))\right],$$
where $m$ is the number of samples, $1\{\dots\}$ is the indicator function which equals to 1 if the condition holds, otherwise equals to 0.
Try implement the loss calculation in the following code. Note that our label (the correct probability) has already been transformed into *one-of-many* form, so the indicator function here is simply an element-wise multiplication of the label matrix and the logarithmic probability.
To minimize this function, we use a method called **batch gradient descent**. It first calculates the gradient of the weight matrix as follows:
$$\nabla_{w}J(w)=-\frac{1}{m}\sum_{i=1}^{m}\left[x^{(i)}(1\{y^{(i)}=j\}-p(y^{(i)}=j \mid x^{(i)};w))\right],$$
then update the current model along the gradient direction:
$$w\leftarrow w - \alpha \nabla_{w}J(w).$$
We have implemented the gradient computation and the weight update for you.
```
# Using gradient descent to fit the correct classes.
def train(w, x, loops):
for i in range(loops):
prob = predict(w, x)
#===========================================================#
# Your code starts here #
#===========================================================#
# Please try compute the current loss value.
loss = -np.sum(label * np.log(prob)) / num_samples
#===========================================================#
# Your code ends here #
#===========================================================#
if i % 10 == 0:
print('Iter {}, training loss {}'.format(i, loss))
# Calculate gradient of weight.
dy = prob - label
dw = np.dot(data.T, dy) / num_samples
# Update.
w -= 0.1 * dw
```
Once you have finished the above codes, you could now try to optimize it for 100 iterations. You should see the loss value dropping.
```
# Now training it for 100 iterations.
start_time = time.time()
train(weight, data, 100)
print('Training time: {}s'.format(time.time() - start_time))
```
Now you could visualize the prediction with trained model again. It should now be very close to what the ground truth is.
```
# Plot the prediction after training. It should show similar image as the ground truth.
plot_data(data, predict(weight, data))
```
## Multinomial Logistic Regression using MinPy (MXNet NumPy)
### Utilize GPU computation with little (or no) NumPy syntax change
You could see even with such a tiny example, 100 iterations take around 9 seconds. In real world, there are billions of samples and much more features and classes. How to efficiently train such a model? One solution is to use GPU. Our tool, MinPy allows you to use GPU to speed up the algorithm and in the meantime, keep the neat NumPy syntax you just went through.
```
# All you need to do is replace the NumPy namespace with MinPy's.
import minpy.numpy as np
import minpy.numpy.random as random
# Initialize weight matrix (again).
weight = random.randn(num_features, num_classes)
# Now call the same training function.
# Since the namespace is redefined, it will automatically run on GPU.
start_time = time.time()
train(weight, data, 100)
# You should observe a significant speed up (around 3x) to the previous training time.
print('Training time: {}s'.format(time.time() - start_time))
```
### Automatic gradient calculation
Compute gradient is tedious especially for complex neural networks you will encounter in the following tutorials. Minpy is able to compute the gradient automatically given arbitrary loss function. Please implement the following loss function (or paste it from your previous codes). The `grad_and_loss` function takes your defined `train_loss` function and then returns a function which will calculate both the loss value and the gradient of weight. The gradient could then be directly used to update the model weight.
**Quiz:** Try modify the loss function by adding an L2-regularization. The new loss function is as follows:
$$J'(w)=J(w)+\sum_{i}w_i^2.$$
```
from minpy.core import grad_and_loss
# Initialize weight matrix (again).
weight = random.randn(num_features, num_classes)
# Using gradient descent to fit the correct classes.
def train_loss(w, x):
#===========================================================#
# Your code starts here #
#===========================================================#
prob = predict(w, x)
loss = -np.sum(label * np.log(prob)) / num_samples
#===========================================================#
# Your code ends here #
#===========================================================#
return loss
# Calculate gradient function automatically.
grad_function = grad_and_loss(train_loss)
# Now training it for 100 iterations.
start_time = time.time()
for i in range(100):
dw, loss = grad_function(weight, data)
if i % 10 == 0:
print('Iter {}, training loss {}'.format(i, loss))
weight -= 0.1 * dw
print('Training time: {}s'.format(time.time() - start_time))
# Plot the result.
# ATTENTION: When calling an external package implemented by NumPy, you need to explicitly convert MinPy's array to NumPy's array.
plot_data(data, predict(weight, data).asnumpy())
```
### Graceful fallback when GPU operators are missing
NumPy is a giant library with hundreds of operators. Our supported GPU operators are only a subset of them, so it is inevitable that you want to try some functions that are currently missing on GPU side. To solve this problem, MinPy gracefully adopts the NumPy implementation once the operator is missing on GPU side, and handles the memory copies among GPU and CPU for you.
```
# First turn on the logging to know what happens under the hood.
import logging
logging.getLogger('minpy.array').setLevel(logging.DEBUG)
x = np.zeros((10, 20))
# `cosh` is currently missing in MXNet's GPU implementation.
# So it will fallback to use NumPy's CPU implementation,
# but you don't need to worry about the memory copy from GPU -> CPU
y = np.cosh(x)
# `log` has GPU implementation, so it will copy the array from CPU -> GPU.
# Once again, you don't need to worry about it. It is transparent.
z = np.log(y)
# Turn off the logging.
logging.getLogger('minpy.array').setLevel(logging.WARN)
```
### Use predefined MXNet symbol as one operator
Now we solve the problem of missing GPU operators. What if we want to use some efficient special operators for neural network that only have GPU implementation? For example, convolution is very slow in numpy's implementation, while NVIDIA provides very efficient CUDA implementation. Hence, MinPy allows you to directly use MXNet's symbol as one operator, and you could mix MXNet's declarative programming with Minpy's imperative programming in one piece of code. The following example replaces the computation of:
$$p(y^{(i)}=j \mid x^{(i)};w)= \frac{\exp(w_j^Tx^{(i)})}{\sum_j\exp(w_j^Tx^{(i)})}$$
with a predefined `SoftmaxOutput` symbol.
**Quiz: ** After you have gone through the next tutorial about MNIST OCR task. You could try replace the `np.dot` function with the `sym.FullyConnected` symbol. Remember, you could combine multiple symbols into one operator.
```
import mxnet
import mxnet.symbol as sym
from minpy.core import Function
# Define softmax symbol (this part will be covered in the later tutorial).
x_shape = (num_samples, num_classes)
label_shape = (num_samples,)
softmax_symbol = sym.SoftmaxOutput(data=sym.Variable('x'), name='softmax', grad_scale=1.0/num_samples)
# Convert MXNet symbol into a callable function (with corresponding gradient function).
softmax = Function(softmax_symbol, {'x': x_shape, 'softmax_label': label_shape})
# Make `softmax_label`. MXNet's softmax operator does not use one-of-many label format.
softmax_label = np.argmax(label, axis=1)
# Redefine loss function using softmax as one operator.
def train_loss(w, x):
y = np.dot(x, w)
prob = softmax(x=y, softmax_label=softmax_label)
loss = -np.sum(label * np.log(prob)) / num_samples
return loss
# Initialize weight matrix (again).
weight = random.randn(num_features, num_classes)
# Calculate gradient function automatically.
grad_function = grad_and_loss(train_loss)
# Now training it for 100 iterations.
start_time = time.time()
for i in range(100):
dw, loss = grad_function(weight, data)
if i % 10 == 0:
print('Iter {}, training loss {}'.format(i, loss))
weight -= 0.1 * dw
print('Training time: {}s'.format(time.time() - start_time))
```
| github_jupyter |
```
__depends__=[]
__dest__="../results/f8.eps"
```
# Plot Terms in the Two-fluid EBTEL Equations
As part of our derivation of the two-fluid EBTEL equations, we'll plot the different terms of the two-fluid electron energy equation,
$$
\frac{L}{\gamma - 1}\frac{dp_e}{dt} = \psi_{TR} - (\mathcal{R}_C + \mathcal{R}_{TR}) + \frac{L}{\gamma - 1}k_Bn\nu_{ei}(T_i - T_e) + LQ_e.
$$
We want to plot each term as a function of time to show their relative contributions to the evolution of the electron energy.
```
import sys
import os
import subprocess
import numpy as np
import seaborn.apionly as sns
import astropy.constants as const
from matplotlib import ticker
import matplotlib.pyplot as plt
sys.path.append(os.path.join(os.environ['EXP_DIR'],'ebtelPlusPlus/rsp_toolkit/python'))
from xml_io import InputHandler,OutputHandler
%matplotlib inline
plt.rcParams.update({'figure.figsize' : [8,5]})
```
Configure the EBTEL run. We'll use $\tau=200$ s and $H_0=0.1$ erg cm$^{-3}$ s$^{-1}$, $L=40$ Mm, and Spitzer conduction.
```
ih = InputHandler(os.path.join(os.environ['EXP_DIR'],'ebtelPlusPlus','config','ebtel.example.cfg.xml'))
config_dict = ih.lookup_vars()
config_dict['calculate_dem'] = False
config_dict['save_terms'] = True
config_dict['use_flux_limiting'] = True
config_dict['use_adaptive_solver'] = True
config_dict['heating']['partition'] = 1.0
config_dict['heating']['background'] = 3.5e-5
config_dict['heating']['events'] = [
{'event':{'magnitude':0.1,'rise_start':0.0,'rise_end':100.0,'decay_start':100.0,'decay_end':200.0}}
]
config_dict['total_time'] = 5000.0
config_dict['tau'] = 0.1
config_dict['adaptive_solver_error'] = 1.0e-9
config_dict['saturation_limit'] = 1.0
config_dict['c1_cond0'] = 6.0
config_dict['c1_rad0'] = 0.6
config_dict['use_c1_grav_correction'] = True
config_dict['use_c1_loss_correction'] = True
config_dict['output_filename'] = '../results/_tmp_'
oh = OutputHandler(config_dict['output_filename']+'.xml',config_dict)
oh.print_to_xml()
```
Run the model.
```
subprocess.call([os.path.join(os.environ['EXP_DIR'],'ebtelPlusPlus','bin','ebtel++.run'),'-c',oh.output_filename])
```
Load the data.
```
data = np.loadtxt(oh.output_dict['output_filename'])
t = data[:,0]
Te = data[:,1]
Ti = data[:,2]
n = data[:,3]
q = data[:,-1]
data = np.loadtxt(oh.output_dict['output_filename']+'.terms')
fce = data[:,0]
fci = data[:,1]
r3 = data[:,2]
rad = data[:,3]
```
Define a function to calculate the Coulomb collision frequency according to [Braginskii (1965)](http://adsabs.harvard.edu/abs/1965RvPP....1..205B).
```
def calc_nu_ei(n,Te):
c1 = 16.*np.sqrt(np.pi)/3.
c2 = const.e.gauss.value**4/(const.m_e.cgs.value*const.m_p.cgs.value)
c3 = 2.*const.k_B.cgs.value*Te/const.m_e.cgs.value
colLog = 20.
return c1*c2*c3**(-3./2.)*n*colLog
```
Calculate the terms as given in the equation above.
```
delta_terms = []
delta_terms.append(fce/(config_dict['loop_length'])/(1.+Te/Ti))
delta_terms.append(-fci/(config_dict['loop_length'])*(Te/Ti)/(1.+Te/Ti))
delta_terms.append(-(Te/Ti*(r3+1.) + 1.)/(1.+Te/Ti)*n**2*rad)
#delta_terms.append(q)
tmp = np.zeros(len(Te))
for i in range(len(Te)):
tmp[i] = const.k_B.cgs.value/(5./3. - 1.)*n[i]*calc_nu_ei(n[i],Te[i])*(Ti[i] - Te[i])
delta_terms.append(tmp)
```
Make the figure.
```
labels = [r'$\mathrm{e}^{-}$ $\mathrm{thermal}$ $\mathrm{conduction}$',
r'$\mathrm{ion}$ $\mathrm{thermal}$ $\mathrm{conduction}$',
r'$\mathrm{radiation}$',r'$\mathrm{equilibration}$']
fig = plt.figure()
ax = fig.gca()
for i in range(len(delta_terms)):
ax.plot(t,delta_terms[i],color=sns.color_palette('deep')[i],label=labels[i])
ax.plot(t,1.0/(config_dict['loop_length'])*1./(1.+Te/Ti)*(fce + (r3*(config_dict['loop_length'])*(n**2)*rad)-Te/Ti*fci),
linestyle='dotted',color='k',label=r'$\psi_{TR}$')
ax.set_xscale('log')
ax.yaxis.set_major_locator(ticker.MaxNLocator(nbins=4))
ax.set_xlim([1,config_dict['total_time']])
ax.set_xlabel(r'$t$ $\mathrm{(s)}$')
ax.set_ylabel(r'$\Delta\bar{E}_e$ $(\mathrm{erg}$ $\mathrm{cm}^{-3}$ $\mathrm{s}^{-1})$')
ax.legend(loc='best')
plt.savefig(__dest__)
plt.show()
```
| github_jupyter |
```
from bayestuner.tuner import BayesTuner
import numpy as np
import seaborn as sns
from bayestuner.optimizer import DifferentialEvolution
from bayestuner.acquisitionfunc import UCB
import matplotlib.pyplot as plt
import math
import matplotlib.animation as animation
from sklearn.gaussian_process.kernels import ConstantKernel, RBF
from scipy.optimize import differential_evolution
from sklearn.gaussian_process import GaussianProcessRegressor
%matplotlib notebook
def optimizer(obj_func, initial_theta, bounds):
# * 'obj_func' is the objective function to be minimized, which
# takes the hyperparameters theta as parameter and an
# optional flag eval_gradient, which determines if the
# gradient is returned additionally to the function value
# * 'initial_theta': the initial value for theta, which can be
# used by local optimizers
# * 'bounds': the bounds on the values of theta
# Returned are the best found hyperparameters theta and
# the corresponding value of the target function.
res = differential_evolution(lambda theta: obj_func(theta,eval_gradient = False),bounds)
theta_opt = res.x
func_min = res.fun
return theta_opt, func_min
f = lambda x : x*np.sin(x)
#f = lambda x : np.sin(10*x)/(2*x) + (x-1)**4
dim = 5
tuner = BayesTuner(objective = f,
bounds = [(-0,19,0)],
acquisition = lambda i : UCB(i,lambda x : 1.96),
n_iter = 30, init_samples = 5)
tuner.gp = GaussianProcessRegressor(kernel = ConstantKernel()*RBF(),
alpha = 1e-2,
n_restarts_optimizer = 3,
optimizer = optimizer,
normalize_y = True).fit(tuner.past_hyper,tuner.past_evals)
tuner.tune()
fig, ax = plt.subplots()
x = np.linspace(0,19,100)
def update(num):
plt.cla()
plt.ylim(-35,35)
ax.plot(x,f(x),'k',linewidth = 2.5,alpha = 0.8,label = 'Target')
ax.plot(tuner.past_hyper[:10+num],tuner.past_evals[:10+num],'D',color = 'darkviolet',markersize = 5,label = 'Samples')
#ax.plot(tuner.past_hyper[:10+num][-1],tuner.past_evals[:10+num][-1],'X',color = 'red',markersize = 10)
#print(tuner.gps[10+num].predict(x.reshape(100,1)))
mean,std = tuner.gps[num].predict(x.reshape(100,1),return_std = True)
#mean = [y[0] for y in mean]
ucb = [m + 1.96*s for (m,s) in zip(mean,std)]
ucb = [y[0] for y in ucb]
lcb = [m - 1.96*s for (m,s) in zip(mean,std)]
lcb = [y[0] for y in lcb]
print(len(mean) == len(x))
ax.plot(x,mean,'r--',linewidth = 1.5,label = 'Mean')
ax.set_title(f"Number of samples: {len(tuner.past_hyper[:10+num])}")
ax.fill_between(x,lcb,ucb,alpha = 0.5,color = 'pink',label = '95% confidence bound')
ax.axvline(tuner.past_hyper[:10+num][-1],color = 'k',linestyle = '--')
ax.axhline(tuner.past_evals[:10+num][-1],color = 'k',linestyle = '--')
plt.axis('off')
plt.legend(prop={'size': 6})
x = np.linspace(0,19,100)
number_of_frames = 50
anim = animation.FuncAnimation(fig, update, number_of_frames,repeat = True,interval = 360, blit = True)
mean,std = tuner.gps[10].predict(x.reshape(100,1),return_std = True)
ucb = [m + 1.96*s for (m,s) in zip(mean,std)]
lcb = [m - 1.96*s for (m,s) in zip(mean,std)]
x = np.linspace(0,20,100)
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.animation as animation
Gamma=0.0005
q=1.6e-19
m=0.067*9e-31
B=10
Ec=(1.0567e-34)*B/m
fig, ax = plt.subplots()
n = 3 #number of lines
x = np.arange(0, 3.6e-3, 1.7e-5) # x-array, third number is interval here, x is energy
lines = [ax.plot(x, np.e**(-(x-((1.0567e-34)*1*1/m))**2/Gamma**2), zorder=i+3)[0] for i in range(n)]
fills = [ax.fill_between(x,0,(np.e**(-(x-((1.0567e-34)*1*1/m))**2/Gamma**2)), facecolor=lines[i].get_color(), zorder=i+3) for i in range(n)]
def animate(i):
for d, line in enumerate(lines):
p=(d+1)/2.
line.set_ydata(np.e**((-(x-((1.0567e-34)*p*i/m))**2)/Gamma**2))
fills[d].remove()
fills[d] = ax.fill_between(x,0,(np.e**(-(x-((1.0567e-34)*p*i/m))**2/Gamma**2)), facecolor=lines[d].get_color(), zorder=d+3)# update the data
return lines + fills
#Init only required for blitting to give a clean slate.
def init():
for line in lines:
line.set_ydata(np.ma.array(x, mask=True))
return lines
ani = animation.FuncAnimation(fig, animate, np.arange(0, 2.5, .01), init_func=init,
interval=10, blit=True)
#Writer = animation.writers['ffmpeg']
#writer = Writer(fps=20, metadata=dict(artist='Me'), bitrate=1800)
#
#ani.save('QHanimati.mp4', writer=writer)
plt.show()
```
| github_jupyter |
```
## This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python Docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the read-only "../input/" directory
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
from fastai.vision import *
from fastai import *
from fastai.basics import *
from fastai.callback.all import *
from fastai.vision.all import *
from fastai.medical.imaging import *
import pydicom
import pandas as pd
from pathlib import Path
from shutil import copy
import warnings
warnings.filterwarnings("ignore")
# You can write up to 20GB to the current directory (/kaggle/working/) that gets preserved as output when you create a version using "Save & Run All"
# You can also write temporary files to /kaggle/temp/, but they won't be saved outside of the current session
# csv = pd.read_csv('../input/vinbigdata-1024-image-dataset/vinbigdata/train.csv')
# csv.head()
# twoclasscsv = pd.DataFrame(columns=['file', 'label'])
# csvdict = {}
# for index, row in csv.iterrows():
# csvdict[row['image_id']] = (int(row['class_id']) == 14)
# for key in csvdict:
# print(key, csvdict[key])
# break
# for key in csvdict:
# twoclasscsv.loc[len(twoclasscsv)] = [key+'.png', ('Normal' if csvdict[key] else 'Abnormal')]
# twoclasscsv.head()
# twoclasscsv.to_csv("./two-class-train.csv")
df = pd.read_csv('../input/vinbigdata-2-class-csv/two-class-train.csv')
df.drop('Unnamed: 0', inplace=True, axis=1)
df.head()
!mkdir ./Normal/
!mkdir ./Abnormal/
for index, row in df.iterrows():
copy('../input/vinbigdata-1024-image-dataset/vinbigdata/train/'+row['file'], './'+row['label']+'/')
tfms = aug_transforms(do_flip = True, flip_vert = False, mult=2.0)
np.random.seed(42)
data = ImageDataLoaders.from_folder('./',train = "", valid_pct=0.2, item_tfms=Resize(1024), batch_tfms=tfms, bs = 4, num_workers = 4)
Data = DataBlock( blocks=(ImageBlock, CategoryBlock), get_items=get_image_files,
splitter=RandomSplitter(valid_pct=0.2, seed=42), get_y=parent_label, item_tfms=Resize(1024))
dls = Data.dataloaders('./')
learn = cnn_learner(data, resnet34, metrics=error_rate)
learn.fine_tune(0)
interp = ClassificationInterpretation.from_learner(learn)
interp.plot_confusion_matrix()
interp.plot_top_losses(5, nrows=1, figsize = (25,5))
learn.save('export1')
from google.colab import files
learn.unfreeze()
def find_appropriate_lr(model:Learner, lr_diff:int = 15, loss_threshold:float = .05, adjust_value:float = 1, plot:bool = False) -> float:
#Run the Learning Rate Finder
model.lr_find()
#Get loss values and their corresponding gradients, and get lr values
losses = np.array(model.recorder.losses)
assert(lr_diff < len(losses))
loss_grad = np.gradient(losses)
lrs = model.recorder.lrs
#Search for index in gradients where loss is lowest before the loss spike
#Initialize right and left idx using the lr_diff as a spacing unit
#Set the local min lr as -1 to signify if threshold is too low
r_idx = -1
l_idx = r_idx - lr_diff
while (l_idx >= -len(losses)) and (abs(loss_grad[r_idx] - loss_grad[l_idx]) > loss_threshold):
local_min_lr = lrs[l_idx]
r_idx -= 1
l_idx -= 1
lr_to_use = local_min_lr * adjust_value
if plot:
# plots the gradients of the losses in respect to the learning rate change
plt.plot(loss_grad)
plt.plot(len(losses)+l_idx, loss_grad[l_idx],markersize=10,marker='o',color='red')
plt.ylabel("Loss")
plt.xlabel("Index of LRs")
plt.show()
plt.plot(np.log10(lrs), losses)
plt.ylabel("Loss")
plt.xlabel("Log 10 Transform of Learning Rate")
loss_coord = np.interp(np.log10(lr_to_use), np.log10(lrs), losses)
plt.plot(np.log10(lr_to_use), loss_coord, markersize=10,marker='o',color='red')
plt.show()
return lr_to_use
lr = find_appropriate_lr(learn, plot = True)
learn.fit_one_cycle(5, lr=lr)
learn.export()
test_csv = pd.read_csv('../input/vinbigdata-1024-image-dataset/vinbigdata/train.csv')
for index, row in test_csv.iterrows():
cat, tensor, probs = learn.predict(open_image(f"../input/vinbigdata-1024-image-dataset/vinbigdata/test/{row['image_id']}"))
```
| github_jupyter |
```
# Allow us to load `open_cp` without installing
import sys, os.path
sys.path.insert(0, os.path.abspath(".."))
```
# Crime prediction from Hawkes processes
Here we continue to explore the EM algorithm for Hawkes processes, but now concentrating upon:
1. Mohler et al. "Randomized Controlled Field Trials of Predictive Policing". Journal of the American Statistical Association (2015) DOI:10.1080/01621459.2015.1077710
```
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
```
# Simulation of the process in a single cell
```
import open_cp.sources.sepp as source_sepp
process = source_sepp.SelfExcitingPointProcess(
background_sampler = source_sepp.HomogeneousPoissonSampler(rate=0.1),
trigger_sampler = source_sepp.ExponentialDecaySampler(intensity=0.5, exp_rate=10))
events = process.sample(0, 1000)
fig, ax = plt.subplots(figsize=(18,1))
ax.scatter(events, (np.random.random(len(events))-0.5) * 0.03, alpha=.5)
ax.set(xlim=[900, 1000], ylim=[-0.1,0.1])
```
## Model fitting for cells with varying background rate
We'll create 100 cells with varying background rate, but the same $\omega, \theta$. We use our library to perform this simulation.
```
rates = np.random.random(size=100)
simulation = source_sepp.GridHawkesProcess(rates, 0.5, 10)
cells = simulation.sample(0, 1000)
```
To simulate a steady state, we'll discard the first half of time in each cell.
```
for i in range(100):
times = cells[i]
cells[i] = times[times>=500] - 500
```
The number of events in each cell varies quite a lot.
```
min(len(t) for t in cells), max(len(t) for t in cells)
import open_cp.seppexp
def optimise(cells, initial_omega=10, iterations=100, time=500):
omega = initial_omega
theta = .5
mu = np.zeros_like(cells) + 0.5
for _ in range(iterations):
omega, theta, mu = open_cp.seppexp.maximisation(cells, omega, theta, mu, time)
return omega, theta, mu
def optimise_corrected(cells, initial_omega=10, iterations=100, time=500):
omega = initial_omega
theta = .5
mu = np.zeros_like(cells) + 0.5
for _ in range(iterations):
omega, theta, mu = open_cp.seppexp.maximisation_corrected(cells, omega, theta, mu, time)
return omega, theta, mu
omega, theta, mu = optimise(cells)
omega, theta
omegac, thetac, muc = optimise_corrected(cells)
omegac, thetac
def plot(rates, mu, ax, title):
ax.plot([0,1], [0,1], color="red", linewidth=1)
ax.scatter(rates, mu)
ax.set(xlim=[0,1], ylim=[0,np.max(mu)*1.05], xlabel="$\\mu$", ylabel="predicted $\\mu$",
title=title)
fig, ax = plt.subplots(ncols=2, figsize=(16,6))
plot(rates, mu, ax[0], "From EM algorithm")
plot(rates, muc,ax[1], "From EM algorithm with edge corrections")
```
Noting that our initial estimate for every $\mu$ is $0.5$, this is good convergence.
## More extreme parameters
However, if we try a rather smaller value of $\omega$, then the optimisation doesn't find the real parameters, tending to systematically over-estimate the background rate $\mu$ and under-estimate the aftershock rate.
```
rates = np.random.random(size=100)
simulation = source_sepp.GridHawkesProcess(rates, 0.5, .1)
cells = simulation.sample(0, 1000)
for i in range(100):
times = cells[i]
cells[i] = times[times>=500] - 500
omega, theta, mu = optimise(cells, .1, 100)
omega, theta
omegac, thetac, muc = optimise_corrected(cells, .1, 100)
omegac, thetac
fig, ax = plt.subplots(ncols=2, figsize=(16,6))
plot(rates, mu, ax[0], "From EM algorithm")
plot(rates, muc, ax[1], "From EM algorithm with edge corrections")
```
## Sampling the whole process, not just a "steady state"
```
rates = np.random.random(size=100)
simulation = source_sepp.GridHawkesProcess(rates, 0.5, 10)
cells = simulation.sample(0, 1000)
omega, theta, mu = optimise(cells, 1, 100, 1000)
omega, theta
omegac, thetac, muc = optimise_corrected(cells, 1, 100, 1000)
omegac, thetac
fig, ax = plt.subplots(ncols=2, figsize=(16,6))
plot(rates, mu, ax[0], "From EM algorithm")
plot(rates, muc, ax[1], "From EM algorithm with edge corrections")
```
## Taking a smaller sample
```
rates = np.random.random(size=100)
simulation = source_sepp.GridHawkesProcess(rates, 0.5, 10)
cells = simulation.sample(0, 350)
omega, theta, mu = optimise(cells, 1, 100, 350)
omega, theta
omegac, thetac, muc = optimise_corrected(cells, 1, 100, 350)
omegac, thetac
fig, ax = plt.subplots(ncols=2, figsize=(16,6))
plot(rates, mu, ax[0], "From EM algorithm")
plot(rates, muc, ax[1], "From EM algorithm with edge corrections")
```
| github_jupyter |
# Emotion Classification
**Module 1: Introduction**
* Author: [Andrés Mitre](https://github.com/andresmitre), [Center for Research in Mathematics (CIMAT)](http://www.cimat.mx/en) in Zacatecas, México.
For installation, I highly recommend to follow the instructions from [Jeff Heaton](https://sites.wustl.edu/jeffheaton/) on how you want to setup your Python TensorFlow environment:
* [Installing TensorFlow, Keras, and Python in Windows](https://www.youtube.com/watch?v=z0qhKP2liHs&index=5&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN)
* [Installing TensorFlow, Keras, and Python in Mac](https://www.youtube.com/watch?v=RUs_qR1hKds&index=4&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN)
#Project Description
The objective of this work, is the reliable recognition of spontaneous and acted emotions related with learning activities through a classificatory off facial expressions and Galvanic Skin Response (GSR). The induction of spontaneous emotions, were generated by clips from movies that showed a good capacity to induce positive and negative effects, elevated levels of emotional activation and variations in the perception of emotional control. On the other hand, the emotions performed were replicated by the participants imitating a sequence of illustrations referencing the emotions. The classificatory was applied to 23 participants of Hispanic ethnic, participants were told a series of instructions. The experiments in this work were based in emotion classification by facial expressions through a Convolutional Neural Network and Galvanic Skin Response. Once, obtained the raw data of the experiment, it processed to the training of classificatory from deep learning and statistical analysis. The Convolutional Neural Network, presented a validation accuracy of 96.5%, a confusion matrix was made with the images of the data set at random and external data sets. In the confusion matrix of the external data set, the real emotions are confused with others, mainly in happy and boredom, correctly classifying in the stress. In the results of Galvanic Skin Response, percentage change tests were performed between the emotions in which significant different were found in happy and boredom, except in stress. With the support of the Neuronal Convolutional Network and Galvanic Skin Activity, it is possible to have a correct classifier of emotions related to learning activities.
Module |Title
---------|--------------
Module 1 |[Introduction](https://github.com/andresmitre/Emotion_Classification/blob/master/introduction.ipynb)
Module 2 |[Haar Cascade Algorithm](https://github.com/andresmitre/Emotion_Classification/blob/master/Haar_Feature_based_Cascade_Classifiers.ipynb)
Module 3 |[Data acquisition](https://github.com/andresmitre/Emotion_Classification/blob/master/data_acquisition.ipynb)
Module 4 |[Convolutional Neural Network](https://github.com/andresmitre/Emotion_Classification/blob/master/CNN.ipynb)
# Author: Andrés Mitre

A brief summary of my credentials is given here:
* Currently in Master in Software Engineering (in Spanish: Maestría en Ingeniería en Software, MIS) at the [Center for Research in Mathematics (CIMAT)](http://www.cimat.mx/en) in Zacatecas, México
* B.E. in Telecommunications, OS and Electronics, [Autonomous University of Sinaloa (in Spanish: Universidad Autónoma de Sinaloa, UAS)](http://web.uas.edu.mx/web/ingles/index.php) in Sinaloa, México
* Participated in projects in the most recognized research centers of Mexico at [The Center for Scientific Research and Higher Education at Ensenada (in Spanish: Centro de Investigación Científica y de Educación Superior de Ensenada, CICESE)](http://eng.cicese.edu.mx/int/index.php?mod=acd&op=intro) and The [National Institute of Astrophysics, Optics and Electronics (in Spanish: Instituto Nacional de Astrofísica, Óptica y Electrónica, INAOE)](http://www.inaoep.mx/en/)
* Interships at [San Diego State University (SDSU)](https://www.sdsu.edu/) and [Western Institute of Technology and Higher Education (in Spanish: Instituto Tecnológico y de Estudios Superiores de Occidente, ITESO,)](https://www.iteso.mx/en/inicio)
Social media:
* [Linked In](https://www.linkedin.com/in/andres18m/) - My Linked In profile, feel free to connect.
* [Twitter](https://twitter.com/andres18m) - 60% baseball | 20% Education | 20% worthless tweets.
* [Instagram](https://www.instagram.com/andresmitre/) - My loop life.
# Environments Recommended
* [Python Anaconda](https://www.continuum.io/downloads) - Python distribution that includes many data science packages, such as Numpy, Scipy, Scikit-Learn, Pandas, and much more.
* [TensorFlow](https://www.tensorflow.org/) - Google's mathematics package for deep learning.
# What is Deep Learning
“Deep learning methods are representation-learning methods with multiple levels of representation, obtained by composing simple but nonlinear modules that each transform the representation at one level (starting with the raw input) into a represen- tation at a higher, slightly more abstract level. [. . . ] The key aspect of deep learning is that these layers are not designed by human engineers: they are learned from data using a general-purpose learning procedure [[1]](https://www.nature.com/articles/nature14539).
## What is Machine Learning
As regards machines, we might say, very broadly, that a machine learns whenever it changes its structure, program, or data (based on its inputs or in response to external information) in such a manner that its expected future performance improves. Some of these changes, such as the addition of a record to a data base, fall comfortably within the province of other disciplines and are not necessarily better understood for being called learning. But, for example, when the performance of a speech-recognition machine improves after hearing several samples of a person’s speech, we feel quite justified in that case to say that the machine has learned. Machine learning usually refers to the changes in systems that perform tasks associated with artificial intelligence (AI). Such tasks involve recognition, diagnosis, planning, robot control, prediction, etc. The “changes” might be either enhancements to already performing systems orab initio synthesis of new systems [[2]](http://robotics.stanford.edu/~nilsson/MLBOOK.pdf).
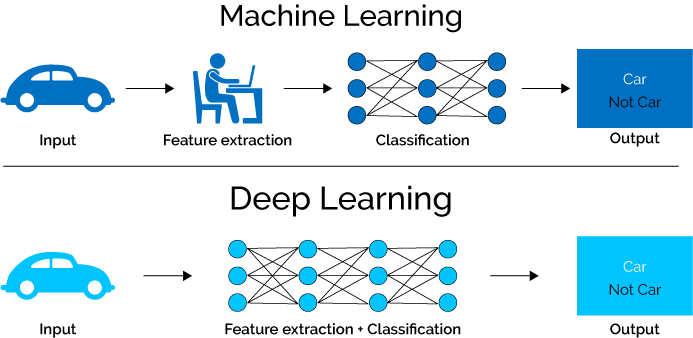
Picture taken from [here](http://digital-nebula.com/2018/01/03/top-terms-you-need-to-know-to-get-started-with-ai/machine-learning-vs-deep-learning/)
##The following packages are needed for this project:
```
conda install scipy
conda install -c trentonoliphant datetime
conda install -c conda-forge opencv
conda install -c mlgill imutils
conda install -c anaconda csvkit
conda install -c conda-forge time
pip install sklearn
pip install pandas
pip install pandas-datareader
pip install matplotlib
pip install pillow
pip install requests
pip install h5py
pip install tensorflow==1.4.0
pip install keras==2.1.2
```
```
# What version of Python do you have?
import keras
import tensorflow as tf
import sys
import sklearn as sk
import pandas as pd
print("Tensor Flow Version: {}".format(tf.__version__))
print("Keras Version: {}".format(keras.__version__))
print()
print("Python {}".format(sys.version))
print('Pandas {}'.format(pd.__version__))
print('Scikit-Learn {}'.format(sk.__version__))
```
| github_jupyter |
# Qiskit Aer: Applying noise to custom unitary gates
The latest version of this notebook is available on https://github.com/Qiskit/qiskit-tutorial.
## Introduction
This notebook shows how to add custom unitary gates to a quantum circuit, and use them for noise simulations in Qiskit Aer.
```
from qiskit import execute, QuantumCircuit, QuantumRegister, ClassicalRegister
from qiskit.quantum_info import Operator, average_gate_fidelity
from qiskit.providers.aer import QasmSimulator
from qiskit.providers.aer.noise import NoiseModel, amplitude_damping_error
from qiskit.tools.visualization import plot_histogram
```
## Creating matrix operators
We can use the `Operator` class in `qiskit.quantum_info` to represent arbitrary matrix operators. If the operator is unitary it can then be added to a quantum circuit and used for simulation on Qiskit Aer.
Lets create two operators below for a CNOT gate and an iSWAP gate:
$$\mbox{CNOT} = \left(\begin{array}
& 1 & 0 & 0 & 0 \\
0 & 0 & 0 & 1 \\
0 & 0 & 1 & 0 \\
0 & 1 & 0 & 0
\end{array}\right), \quad
\mbox{iSWAP} = \left(\begin{array}
& 1 & 0 & 0 & 0 \\
0 & 0 & i & 0 \\
0 & i & 0 & 0 \\
0 & 0 & 0 & 1
\end{array}\right)$$
```
# CNOT matrix operator with qubit-0 as control and qubit-1 as target
cx_op = Operator([[1, 0, 0, 0],
[0, 0, 0, 1],
[0, 0, 1, 0],
[0, 1, 0, 0]])
# iSWAP matrix operator
iswap_op = Operator([[1, 0, 0, 0],
[0, 0, 1j, 0],
[0, 1j, 0, 0],
[0, 0, 0, 1]])
```
**Note:** The matrix is specified with respect to the tensor product $U_{b}\otimes U_{a}$ for qubits specified by list `[a, b]`.
## Using operators in circuits
Let us demonstrate how these can be used in a circuit. We will consider an example of implementing a CNOT gate decomposed in terms of single-qubit gates and the iSWAP gate as follows.
```
# CNOT in terms of iSWAP and single-qubit gates
cx_circ = QuantumCircuit(2)
# Add gates
cx_circ.sdg(1)
cx_circ.h(1)
cx_circ.sdg(0)
cx_circ.unitary(iswap_op, [0, 1], label='iswap')
cx_circ.sdg(0)
cx_circ.h(0)
cx_circ.sdg(0)
cx_circ.unitary(iswap_op, [0, 1], label='iswap')
cx_circ.s(1)
print(cx_circ)
```
Note that we have assigned an optional *label* of `"iswap"` to the unitary when it is inserted. This allows us to identify this unitary in a Qiskit Aer `NoiseModel` so that we can add errors to these custom unitary gates in noisy circuit simulations.
We can confirm this circuit returns the correct output using the `Operator` class as a simulator for the circuit:
```
# Simulate the unitary for the circuit using Operator:
unitary = Operator(cx_circ)
print(unitary)
```
And to confirm the output is correct we can compute the average gate fidelity:
```
f_ave = average_gate_fidelity(cx_op, unitary)
print("Average Gate Fidelity: F = {:f}".format(f_ave))
```
## Creating a custom unitary in a noise model
The Qiskit Aer `QasmSimulator` supports simulation of arbitrary unitary operators directly as specified by the `"unitary"` in the basis gates.
```
'unitary' in QasmSimulator().configuration().basis_gates
```
This allows us to add noise models to arbitrary unitaries in our simulation when we identify them using the optional `label` argument of `QuantumCircuit.unitary`.
We will now do this by creating a `NoiseModel` that includes a quantum error channel on our custom iSWAP gate. For our example we will create a 2-qubit error consisting of two single-qubit amplitude damping channels with different damping parameters. For now we will assume all the other circuit instructions are ideal.
```
# Error parameters
param_q0 = 0.05 # damping parameter for qubit-0
param_q1 = 0.1 # damping parameter for qubit-1
# Construct the error
qerror_q0 = amplitude_damping_error(param_q0)
qerror_q1 = amplitude_damping_error(param_q1)
iswap_error = qerror_q1.tensor(qerror_q0)
# Build the noise model by adding the error to the "iswap" gate
noise_model = NoiseModel()
noise_model.add_all_qubit_quantum_error(iswap_error, 'iswap')
```
Note that when we add an error to a custom label such as `"iswap"` the `NoiseModel` does not know what gate this label is supposed to apply to, so we must manually add the desired gate string to the noise model `basis_gates`. This ensures that the compiler will unroll to the correct basis gates for the noise model simulation. This can done using the `NoiseModel.add_basis_gates` function:
```
noise_model.add_basis_gates(['unitary'])
print(noise_model.basis_gates)
```
By default the basis gates of a noise model are `['cx','id','u3']` plus any standard `QasmSimulator` basis gates that are added to the noise model.
## Simulating a custom unitary noise model
Let us first take our previous CX circuit and add an initial Hadamard gate and final measurement to create a Bell-state preparation circuit that we may simulator on the `QasmSimulator` both for the ideal and noisy case:
```
# Bell state circuit where iSWAPS should be inserted at barrier locations
bell_circ = QuantumCircuit(2, 2, name='bell')
bell_circ.h(0)
bell_circ = bell_circ + cx_circ
bell_circ.measure([0,1], [0,1])
print(bell_circ)
```
### Ideal output
Let's first see the ideal output. Since this generates a Bell-state we expect two peaks for 00 and 11.
```
# Execute on the simulator without noise
job = execute(bell_circ, QasmSimulator(),
basis_gates=noise_model.basis_gates)
ideal_result = job.result()
ideal_counts = ideal_result.get_counts(bell_circ)
plot_histogram(ideal_counts, title='Ideal output for iSWAP bell-state preparation')
```
### Noisy circuit execution
Finally, let's now simulate it with our custom noise model. Since there is a small amplitude damping error on the two-qubit gates we expect small additional peaks for the 01 and 10 outcome probabilities.
```
# Execute on the simulator without noise
job = execute(bell_circ, QasmSimulator(),
basis_gates=noise_model.basis_gates,
noise_model=noise_model)
noise_result = job.result()
noise_counts = noise_result.get_counts(bell_circ)
plot_histogram(noise_counts, title='Noisy output for iSWAP bell-state preparation')
import qiskit.tools.jupyter
%qiskit_version_table
%qiskit_copyright
```
| github_jupyter |
# 🔪 JAX - The Sharp Bits 🔪
*levskaya@ mattjj@*
When walking about the countryside of [Italy](https://iaml.it/blog/jax-intro), the people will not hesitate to tell you that __JAX__ has _"una anima di pura programmazione funzionale"_.
__JAX__ is a language for __expressing__ and __composing__ __transformations__ of numerical programs. As such it needs to control the _unwanted proliferation_ of __side-effects__ in its programs so that analysis and transformation of its computations remain tractable!
This requires us to write code in a _functional_ style with _explicit_ descriptions of how the state of a program changes, which results in __several important differences__ to how you might be used to programming in Numpy, Tensorflow or Pytorch.
Herein we try to cover the most frequent points of trouble that users encounter when starting out in __JAX__.
```
import numpy as onp
from jax import grad, jit
from jax import lax
from jax import random
import jax
import jax.numpy as np
import matplotlib as mpl
from matplotlib import pyplot as plt
from matplotlib import rcParams
rcParams['image.interpolation'] = 'nearest'
rcParams['image.cmap'] = 'viridis'
rcParams['axes.grid'] = False
```
## 🔪 In-Place Updates
In Numpy you're used to doing this:
```
numpy_array = onp.zeros((3,3), dtype=np.float32)
print("original array:")
print(numpy_array)
# In place, mutating update
numpy_array[1, :] = 1.0
print("updated array:")
print(numpy_array)
```
If we try to update a JAX device array in-place, however, we get an __error__! (☉_☉)
```
jax_array = np.zeros((3,3), dtype=np.float32)
# In place update of JAX's array will yield an error!
jax_array[1, :] = 1.0
```
__What gives?!__
Allowing mutation of variables in-place makes program analysis and transformation very difficult. JAX requires a pure functional expression of a numerical program.
Instead, JAX offers the _functional_ update functions: [__index_update__](https://jax.readthedocs.io/en/latest/_autosummary/jax.ops.index_update.html#jax.ops.index_update), [__index_add__](https://jax.readthedocs.io/en/latest/_autosummary/jax.ops.index_add.html#jax.ops.index_add), [__index_min__](https://jax.readthedocs.io/en/latest/_autosummary/jax.ops.index_min.html#jax.ops.index_min), [__index_max__](https://jax.readthedocs.io/en/latest/_autosummary/jax.ops.index_max.html#jax.ops.index_max), and the [__index__](https://jax.readthedocs.io/en/latest/_autosummary/jax.ops.index.html#jax.ops.index) helper.
️⚠️ inside `jit`'d code and `lax.while_loop` or `lax.fori_loop` the __size__ of slices can't be functions of argument _values_ but only functions of argument _shapes_ -- the slice start indices have no such restriction. See the below __Control Flow__ Section for more information on this limitation.
```
from jax.ops import index, index_add, index_update
```
### index_update
If the __input values__ of __index_update__ aren't reused, __jit__-compiled code will perform these operations _in-place_.
```
jax_array = np.zeros((3, 3))
print("original array:")
print(jax_array)
new_jax_array = index_update(jax_array, index[1, :], 1.)
print("old array unchanged:")
print(jax_array)
print("new array:")
print(new_jax_array)
```
### index_add
If the __input values__ of __index_update__ aren't reused, __jit__-compiled code will perform these operations _in-place_.
```
print("original array:")
jax_array = np.ones((5, 6))
print(jax_array)
new_jax_array = index_add(jax_array, index[::2, 3:], 7.)
print("new array post-addition:")
print(new_jax_array)
```
## 🔪 Random Numbers
> _If all scientific papers whose results are in doubt because of bad
> `rand()`s were to disappear from library shelves, there would be a
> gap on each shelf about as big as your fist._ - Numerical Recipes
### RNGs and State
You're used to _stateful_ pseudorandom number generators (PRNGs) from numpy and other libraries, which helpfully hide a lot of details under the hood to give you a ready fountain of pseudorandomness:
```
print(onp.random.random())
print(onp.random.random())
print(onp.random.random())
```
Underneath the hood, numpy uses the [Mersenne Twister](https://en.wikipedia.org/wiki/Mersenne_Twister) PRNG to power its pseudorandom functions. The PRNG has a period of $2^{19937-1}$ and at any point can be described by __624 32bit unsigned ints__ and a __position__ indicating how much of this "entropy" has been used up.
```
onp.random.seed(0)
rng_state = onp.random.get_state()
#print(rng_state)
# --> ('MT19937', array([0, 1, 1812433255, 1900727105, 1208447044,
# 2481403966, 4042607538, 337614300, ... 614 more numbers...,
# 3048484911, 1796872496], dtype=uint32), 624, 0, 0.0)
```
This pseudorandom state vector is automagically updated behind the scenes every time a random number is needed, "consuming" 2 of the uint32s in the Mersenne twister state vector:
```
_ = onp.random.uniform()
rng_state = onp.random.get_state()
#print(rng_state)
# --> ('MT19937', array([2443250962, 1093594115, 1878467924,
# ..., 2648828502, 1678096082], dtype=uint32), 2, 0, 0.0)
# Let's exhaust the entropy in this PRNG statevector
for i in range(311):
_ = onp.random.uniform()
rng_state = onp.random.get_state()
#print(rng_state)
# --> ('MT19937', array([2443250962, 1093594115, 1878467924,
# ..., 2648828502, 1678096082], dtype=uint32), 624, 0, 0.0)
# Next call iterates the RNG state for a new batch of fake "entropy".
_ = onp.random.uniform()
rng_state = onp.random.get_state()
# print(rng_state)
# --> ('MT19937', array([1499117434, 2949980591, 2242547484,
# 4162027047, 3277342478], dtype=uint32), 2, 0, 0.0)
```
The problem with magic PRNG state is that it's hard to reason about how it's being used and updated across different threads, processes, and devices, and it's _very easy_ to screw up when the details of entropy production and consumption are hidden from the end user.
The Mersenne Twister PRNG is also known to have a [number](https://cs.stackexchange.com/a/53475) of problems, it has a large 2.5Kb state size, which leads to problematic [initialization issues](https://dl.acm.org/citation.cfm?id=1276928). It [fails](http://www.pcg-random.org/pdf/toms-oneill-pcg-family-v1.02.pdf) modern BigCrush tests, and is generally slow.
### JAX PRNG
JAX instead implements an _explicit_ PRNG where entropy production and consumption are handled by explicitly passing and iterating PRNG state. JAX uses a modern [Three-fry counter-based PRNG](https://github.com/google/jax/blob/master/design_notes/prng.md) that's __splittable__. That is, its design allows us to __fork__ the PRNG state into new PRNGs for use with parallel stochastic generation.
The random state is described by two unsigned-int32s that we call a __key__:
```
from jax import random
key = random.PRNGKey(0)
key
```
JAX's random functions produce pseudorandom numbers from the PRNG state, but __do not__ change the state!
Reusing the same state will cause __sadness__ and __monotony__, depriving the enduser of __lifegiving chaos__:
```
print(random.normal(key, shape=(1,)))
print(key)
# No no no!
print(random.normal(key, shape=(1,)))
print(key)
```
Instead, we __split__ the PRNG to get usable __subkeys__ every time we need a new pseudorandom number:
```
print("old key", key)
key, subkey = random.split(key)
normal_pseudorandom = random.normal(subkey, shape=(1,))
print(" \---SPLIT --> new key ", key)
print(" \--> new subkey", subkey, "--> normal", normal_pseudorandom)
```
We propagate the __key__ and make new __subkeys__ whenever we need a new random number:
```
print("old key", key)
key, subkey = random.split(key)
normal_pseudorandom = random.normal(subkey, shape=(1,))
print(" \---SPLIT --> new key ", key)
print(" \--> new subkey", subkey, "--> normal", normal_pseudorandom)
```
We can generate more than one __subkey__ at a time:
```
key, *subkeys = random.split(key, 4)
for subkey in subkeys:
print(random.normal(subkey, shape=(1,)))
```
## 🔪 Control Flow
### ✔ python control_flow + autodiff ✔
If you just want to apply `grad` to your python functions, you can use regular python control-flow constructs with no problems, as if you were using [Autograd](https://github.com/hips/autograd) (or Pytorch or TF Eager).
```
def f(x):
if x < 3:
return 3. * x ** 2
else:
return -4 * x
print(grad(f)(2.)) # ok!
print(grad(f)(4.)) # ok!
```
### python control flow + JIT
Using control flow with `jit` is more complicated, and by default it has more constraints.
This works:
```
@jit
def f(x):
for i in range(3):
x = 2 * x
return x
print(f(3))
```
So does this:
```
@jit
def g(x):
y = 0.
for i in range(x.shape[0]):
y = y + x[i]
return y
print(g(np.array([1., 2., 3.])))
```
But this doesn't, at least by default:
```
@jit
def f(x):
if x < 3:
return 3. * x ** 2
else:
return -4 * x
# This will fail!
try:
f(2)
except Exception as e:
print("ERROR:", e)
```
__What gives!?__
When we `jit`-compile a function, we usually want to compile a version of the function that works for many different argument values, so that we can cache and reuse the compiled code. That way we don't have to re-compile on each function evaluation.
For example, if we evaluate an `@jit` function on the array `np.array([1., 2., 3.], np.float32)`, we might want to compile code that we can reuse to evaluate the function on `np.array([4., 5., 6.], np.float32)` to save on compile time.
To get a view of your Python code that is valid for many different argument values, JAX traces it on _abstract values_ that represent sets of possible inputs. There are [multiple different levels of abstraction](https://github.com/google/jax/blob/master/jax/abstract_arrays.py), and different transformations use different abstraction levels.
By default, `jit` traces your code on the `ShapedArray` abstraction level, where each abstract value represents the set of all array values with a fixed shape and dtype. For example, if we trace using the abstract value `ShapedArray((3,), np.float32)`, we get a view of the function that can be reused for any concrete value in the corresponding set of arrays. That means we can save on compile time.
But there's a tradeoff here: if we trace a Python function on a `ShapedArray((), np.float32)` that isn't committed to a specific concrete value, when we hit a line like `if x < 3`, the expression `x < 3` evaluates to an abstract `ShapedArray((), np.bool_)` that represents the set `{True, False}`. When Python attempts to coerce that to a concrete `True` or `False`, we get an error: we don't know which branch to take, and can't continue tracing! The tradeoff is that with higher levels of abstraction we gain a more general view of the Python code (and thus save on re-compilations), but we require more constraints on the Python code to complete the trace.
The good news is that you can control this tradeoff yourself. By having `jit` trace on more refined abstract values, you can relax the traceability constraints. For example, using the `static_argnums` argument to `jit`, we can specify to trace on concrete values of some arguments. Here's that example function again:
```
def f(x):
if x < 3:
return 3. * x ** 2
else:
return -4 * x
f = jit(f, static_argnums=(0,))
print(f(2.))
```
Here's another example, this time involving a loop:
```
def f(x, n):
y = 0.
for i in range(n):
y = y + x[i]
return y
f = jit(f, static_argnums=(1,))
f(np.array([2., 3., 4.]), 2)
```
In effect, the loop gets statically unrolled. JAX can also trace at _higher_ levels of abstraction, like `Unshaped`, but that's not currently the default for any transformation
️⚠️ **functions with argument-__value__ dependent shapes**
These control-flow issues also come up in a more subtle way: numerical functions we want to __jit__ can't specialize the shapes of internal arrays on argument _values_ (specializing on argument __shapes__ is ok). As a trivial example, let's make a function whose output happens to depend on the input variable `length`.
```
def example_fun(length, val):
return np.ones((length,)) * val
# un-jit'd works fine
print(example_fun(5, 4))
bad_example_jit = jit(example_fun)
# this will fail:
try:
print(bad_example_jit(10, 4))
except Exception as e:
print("error!", e)
# static_argnums tells JAX to recompile on changes at these argument positions:
good_example_jit = jit(example_fun, static_argnums=(0,))
# first compile
print(good_example_jit(10, 4))
# recompiles
print(good_example_jit(5, 4))
```
`static_argnums` can be handy if `length` in our example rarely changes, but it would be disastrous if it changed a lot!
Lastly, if your function has global side-effects, JAX's tracer can cause weird things to happen. A common gotcha is trying to print arrays inside __jit__'d functions:
```
@jit
def f(x):
print(x)
y = 2 * x
print(y)
return y
f(2)
```
### Structured control flow primitives
There are more options for control flow in JAX. Say you want to avoid re-compilations but still want to use control flow that's traceable, and that avoids un-rolling large loops. Then you can use these 4 structured control flow primitives:
- `lax.cond` _differentiable_
- `lax.while_loop` __fwd-mode-differentiable__
- `lax.fori_loop` __fwd-mode-differentiable__
- `lax.scan` _differentiable_
#### cond
python equivalent:
```
def cond(pred, true_operand, true_fun, false_operand, false_fun):
if pred:
return true_fun(true_operand)
else:
return false_fun(false_operand)
```
```
from jax import lax
operand = np.array([0.])
lax.cond(True, operand, lambda x: x+1, operand, lambda x: x-1)
# --> array([1.], dtype=float32)
lax.cond(False, operand, lambda x: x+1, operand, lambda x: x-1)
# --> array([-1.], dtype=float32)
```
#### while_loop
python equivalent:
```
def while_loop(cond_fun, body_fun, init_val):
val = init_val
while cond_fun(val):
val = body_fun(val)
return val
```
```
init_val = 0
cond_fun = lambda x: x<10
body_fun = lambda x: x+1
lax.while_loop(cond_fun, body_fun, init_val)
# --> array(10, dtype=int32)
```
#### fori_loop
python equivalent:
```
def fori_loop(start, stop, body_fun, init_val):
val = init_val
for i in range(start, stop):
val = body_fun(i, val)
return val
```
```
init_val = 0
start = 0
stop = 10
body_fun = lambda i,x: x+i
lax.fori_loop(start, stop, body_fun, init_val)
# --> array(45, dtype=int32)
```
#### Summary
$$
\begin{array} {r|rr}
\hline \
\textrm{construct}
& \textrm{jit}
& \textrm{grad} \\
\hline \
\textrm{if} & ❌ & ✔ \\
\textrm{for} & ✔* & ✔\\
\textrm{while} & ✔* & ✔\\
\textrm{lax.cond} & ✔ & ✔\\
\textrm{lax.while_loop} & ✔ & \textrm{fwd}\\
\textrm{lax.fori_loop} & ✔ & \textrm{fwd}\\
\textrm{lax.scan} & ✔ & ✔\\
\hline
\end{array}
$$
<center>$\ast$ = argument-__value__-independent loop condition - unrolls the loop </center>
## 🔪 Convolutions
JAX and XLA offer the very general N-dimensional __conv_general_dilated__ function, but it's not very obvious how to use it. We'll give some examples of the common use-cases. There are also the convenience functions `lax.conv` and `lax.conv_general_padding` for the most common kinds of convolutions.
A survey of the family of convolutional operators, [a guide to convolutional arithmetic](https://arxiv.org/abs/1603.07285) is highly recommended reading!
Let's define a simple diagonal edge kernel:
```
# 2D kernel - HWIO layout
kernel = onp.zeros((3, 3, 3, 3), dtype=np.float32)
kernel += onp.array([[1, 1, 0],
[1, 0,-1],
[0,-1,-1]])[:, :, onp.newaxis, onp.newaxis]
print("Edge Conv kernel:")
plt.imshow(kernel[:, :, 0, 0]);
```
And we'll make a simple synthetic image:
```
# NHWC layout
img = onp.zeros((1, 200, 198, 3), dtype=np.float32)
for k in range(3):
x = 30 + 60*k
y = 20 + 60*k
img[0, x:x+10, y:y+10, k] = 1.0
print("Original Image:")
plt.imshow(img[0]);
```
### lax.conv and lax.conv_with_general_padding
These are the simple convenience functions for convolutions
️⚠️ The convenience `lax.conv`, `lax.conv_with_general_padding` helper function assume __NCHW__ images and __IOHW__ kernels.
```
out = lax.conv(np.transpose(img,[0,3,1,2]), # lhs = NCHW image tensor
np.transpose(kernel,[2,3,0,1]), # rhs = IOHW conv kernel tensor
(1, 1), # window strides
'SAME') # padding mode
print("out shape: ", out.shape)
print("First output channel:")
plt.figure(figsize=(10,10))
plt.imshow(onp.array(out)[0,0,:,:]);
out = lax.conv_with_general_padding(
np.transpose(img,[0,3,1,2]), # lhs = NCHW image tensor
np.transpose(kernel,[2,3,0,1]), # rhs = IOHW conv kernel tensor
(1, 1), # window strides
((2,2),(2,2)), # general padding 2x2
(1,1), # lhs/image dilation
(1,1)) # rhs/kernel dilation
print("out shape: ", out.shape)
print("First output channel:")
plt.figure(figsize=(10,10))
plt.imshow(onp.array(out)[0,0,:,:]);
```
### Dimension Numbers define dimensional layout for conv_general_dilated
The important argument is the 3-tuple of axis layout arguments:
(Input Layout, Kernel Layout, Output Layout)
- __N__ - batch dimension
- __H__ - spatial height
- __W__ - spatial height
- __C__ - channel dimension
- __I__ - kernel _input_ channel dimension
- __O__ - kernel _output_ channel dimension
⚠️ To demonstrate the flexibility of dimension numbers we choose a __NHWC__ image and __HWIO__ kernel convention for `lax.conv_general_dilated` below.
```
dn = lax.conv_dimension_numbers(img.shape, # only ndim matters, not shape
kernel.shape, # only ndim matters, not shape
('NHWC', 'HWIO', 'NHWC')) # the important bit
print(dn)
```
#### SAME padding, no stride, no dilation
```
out = lax.conv_general_dilated(img, # lhs = image tensor
kernel, # rhs = conv kernel tensor
(1,1), # window strides
'SAME', # padding mode
(1,1), # lhs/image dilation
(1,1), # rhs/kernel dilation
dn) # dimension_numbers = lhs, rhs, out dimension permutation
print("out shape: ", out.shape)
print("First output channel:")
plt.figure(figsize=(10,10))
plt.imshow(onp.array(out)[0,:,:,0]);
```
#### VALID padding, no stride, no dilation
```
out = lax.conv_general_dilated(img, # lhs = image tensor
kernel, # rhs = conv kernel tensor
(1,1), # window strides
'VALID', # padding mode
(1,1), # lhs/image dilation
(1,1), # rhs/kernel dilation
dn) # dimension_numbers = lhs, rhs, out dimension permutation
print("out shape: ", out.shape, "DIFFERENT from above!")
print("First output channel:")
plt.figure(figsize=(10,10))
plt.imshow(onp.array(out)[0,:,:,0]);
```
#### SAME padding, 2,2 stride, no dilation
```
out = lax.conv_general_dilated(img, # lhs = image tensor
kernel, # rhs = conv kernel tensor
(2,2), # window strides
'SAME', # padding mode
(1,1), # lhs/image dilation
(1,1), # rhs/kernel dilation
dn) # dimension_numbers = lhs, rhs, out dimension permutation
print("out shape: ", out.shape, " <-- half the size of above")
plt.figure(figsize=(10,10))
print("First output channel:")
plt.imshow(onp.array(out)[0,:,:,0]);
```
#### VALID padding, no stride, rhs kernel dilation ~ Atrous convolution (excessive to illustrate)
```
out = lax.conv_general_dilated(img, # lhs = image tensor
kernel, # rhs = conv kernel tensor
(1,1), # window strides
'VALID', # padding mode
(1,1), # lhs/image dilation
(12,12), # rhs/kernel dilation
dn) # dimension_numbers = lhs, rhs, out dimension permutation
print("out shape: ", out.shape)
plt.figure(figsize=(10,10))
print("First output channel:")
plt.imshow(onp.array(out)[0,:,:,0]);
```
#### VALID padding, no stride, lhs=input dilation ~ Transposed Convolution
```
out = lax.conv_general_dilated(img, # lhs = image tensor
kernel, # rhs = conv kernel tensor
(1,1), # window strides
((0, 0), (0, 0)), # padding mode
(2,2), # lhs/image dilation
(1,1), # rhs/kernel dilation
dn) # dimension_numbers = lhs, rhs, out dimension permutation
print("out shape: ", out.shape, "<-- larger than original!")
plt.figure(figsize=(10,10))
print("First output channel:")
plt.imshow(onp.array(out)[0,:,:,0]);
```
We can use the last to, for instance, implement _transposed convolutions_:
```
# The following is equivalent to tensorflow:
# N,H,W,C = img.shape
# out = tf.nn.conv2d_transpose(img, kernel, (N,2*H,2*W,C), (1,2,2,1))
# transposed conv = 180deg kernel roation plus LHS dilation
# rotate kernel 180deg:
kernel_rot = np.rot90(np.rot90(kernel, axes=(0,1)), axes=(0,1))
# need a custom output padding:
padding = ((2, 1), (2, 1))
out = lax.conv_general_dilated(img, # lhs = image tensor
kernel_rot, # rhs = conv kernel tensor
(1,1), # window strides
padding, # padding mode
(2,2), # lhs/image dilation
(1,1), # rhs/kernel dilation
dn) # dimension_numbers = lhs, rhs, out dimension permutation
print("out shape: ", out.shape, "<-- transposed_conv")
plt.figure(figsize=(10,10))
print("First output channel:")
plt.imshow(onp.array(out)[0,:,:,0]);
```
### 1D Convolutions
You aren't limited to 2D convolutions, a simple 1D demo is below:
```
# 1D kernel - WIO layout
kernel = onp.array([[[1, 0, -1], [-1, 0, 1]],
[[1, 1, 1], [-1, -1, -1]]],
dtype=np.float32).transpose([2,1,0])
# 1D data - NWC layout
data = onp.zeros((1, 200, 2), dtype=np.float32)
for i in range(2):
for k in range(2):
x = 35*i + 30 + 60*k
data[0, x:x+30, k] = 1.0
print("in shapes:", data.shape, kernel.shape)
plt.figure(figsize=(10,5))
plt.plot(data[0]);
dn = lax.conv_dimension_numbers(data.shape, kernel.shape,
('NWC', 'WIO', 'NWC'))
print(dn)
out = lax.conv_general_dilated(data, # lhs = image tensor
kernel, # rhs = conv kernel tensor
(1,), # window strides
'SAME', # padding mode
(1,), # lhs/image dilation
(1,), # rhs/kernel dilation
dn) # dimension_numbers = lhs, rhs, out dimension permutation
print("out shape: ", out.shape)
plt.figure(figsize=(10,5))
plt.plot(out[0]);
```
### 3D Convolutions
```
# Random 3D kernel - HWDIO layout
kernel = onp.array([
[[0, 0, 0], [0, 1, 0], [0, 0, 0]],
[[0, -1, 0], [-1, 0, -1], [0, -1, 0]],
[[0, 0, 0], [0, 1, 0], [0, 0, 0]]],
dtype=np.float32)[:, :, :, onp.newaxis, onp.newaxis]
# 3D data - NHWDC layout
data = onp.zeros((1, 30, 30, 30, 1), dtype=np.float32)
x, y, z = onp.mgrid[0:1:30j, 0:1:30j, 0:1:30j]
data += (onp.sin(2*x*np.pi)*onp.cos(2*y*np.pi)*onp.cos(2*z*np.pi))[None,:,:,:,None]
print("in shapes:", data.shape, kernel.shape)
dn = lax.conv_dimension_numbers(data.shape, kernel.shape,
('NHWDC', 'HWDIO', 'NHWDC'))
print(dn)
out = lax.conv_general_dilated(data, # lhs = image tensor
kernel, # rhs = conv kernel tensor
(1,1,1), # window strides
'SAME', # padding mode
(1,1,1), # lhs/image dilation
(1,1,1), # rhs/kernel dilation
dn) # dimension_numbers
print("out shape: ", out.shape)
# Make some simple 3d density plots:
from mpl_toolkits.mplot3d import Axes3D
def make_alpha(cmap):
my_cmap = cmap(np.arange(cmap.N))
my_cmap[:,-1] = np.linspace(0, 1, cmap.N)**3
return mpl.colors.ListedColormap(my_cmap)
my_cmap = make_alpha(plt.cm.viridis)
fig = plt.figure()
ax = fig.gca(projection='3d')
ax.scatter(x.ravel(), y.ravel(), z.ravel(), c=data.ravel(), cmap=my_cmap)
ax.axis('off')
ax.set_title('input')
fig = plt.figure()
ax = fig.gca(projection='3d')
ax.scatter(x.ravel(), y.ravel(), z.ravel(), c=out.ravel(), cmap=my_cmap)
ax.axis('off')
ax.set_title('3D conv output');
```
## 🔪 NaNs
### Debugging NaNs
If you want to trace where NaNs are occurring in your functions or gradients, you can turn on the NaN-checker by:
- setting the `JAX_DEBUG_NANS=True` environment variable.
- adding `from jax.config import config` and `config.update("jax_debug_nans", True)` near the top of your main file
- adding `from jax.config import config` and `config.parse_flags_with_absl()` to your main file, then set the option using a command-line flag like `--jax_debug_nans=True`.
This will cause computations to error-out immediately on production of a NaN.
⚠️ You shouldn't have the NaN-checker on if you're not debugging, as it can introduce lots of device-host round-trips and performance regressions!
## Double (64bit) precision
At the moment, JAX by default enforces single-precision numbers to mitigate the Numpy API's tendency to aggressively promote operands to `double`. This is the desired behavior for many machine-learning applications, but it may catch you by surprise!
```
x = random.uniform(random.PRNGKey(0), (1000,), dtype=np.float64)
x.dtype
```
To use double-precision numbers, you need to set the `jax_enable_x64` configuration variable __at startup__.
There are a few ways to do this:
1. You can enable 64bit mode by setting the environment variable `JAX_ENABLE_X64=True`.
2. You can manually set the `jax_enable_x64` configuration flag at startup:
```
# again, this only works on startup!
from jax.config import config
config.update("jax_enable_x64", True)
```
3. You can parse command-line flags with `absl.app.run(main)`
```
from jax.config import config
config.config_with_absl()
```
4. If you want JAX to run absl parsing for you, i.e. you don't want to do `absl.app.run(main)`, you can instead use
```
from jax.config import config
if __name__ == '__main__':
# calls config.config_with_absl() *and* runs absl parsing
config.parse_flags_with_absl()
```
Note that #2-#4 work for _any_ of JAX's configuration options.
We can then confirm that `x64` mode is enabled:
```
from jax import numpy as np, random
x = random.uniform(random.PRNGKey(0), (1000,), dtype=np.float64)
x.dtype # --> dtype('float64')
```
### Caveats
⚠️ XLA doesn't support 64-bit convolutions on all backends!
## Fin.
If something's not covered here that has caused you weeping and gnashing of teeth, please let us know and we'll extend these introductory _advisos_!
| github_jupyter |
```
import os
import pyvtk
import numpy as np
import xarray as xr
import matplotlib.pyplot as plt
# The data structure in element-wise output is too complicated for xarray.open_mfdataset.
# Here we open the files as individual datasets and concatenate them on the variable level.
# This code is compatible with parallel netcdf build (single file output)
# load_wave_data=True: read wave data and return numpy.ndarray
# load_wave_data=False: do not read wave data and return xarray.DataArray (use False if data is big)
def read_element_output(data_dir, load_wave_data=True):
################ open files ################
# filenames
nc_fnames = [f for f in os.listdir(data_dir) if 'axisem3d_synthetics.nc' in f]
print('files to open: ', nc_fnames)
# open files
nc_files = []
for nc_fname in nc_fnames:
nc_files.append(xr.open_dataset(data_dir + '/' + nc_fname))
################ variables that are the same in the datasets ################
# read Na grid (all azimuthal dimensions)
na_grid = nc_files[0].data_vars['list_na_grid'].values.astype(int)
# read time
data_time = nc_files[0].data_vars['data_time'].values
################ variables to be concatenated over the datasets ################
# define empty lists of xarray.DataArray objects
xda_list_element_na = []
xda_list_element_coords = []
dict_xda_list_element = {}
dict_xda_data_wave = {}
for nag in na_grid:
dict_xda_list_element[nag] = []
dict_xda_data_wave[nag] = []
# loop over nc files
for nc_file in nc_files:
# append DataArrays
xda_list_element_na.append(nc_file.data_vars['list_element_na'])
xda_list_element_coords.append(nc_file.data_vars['list_element_coords'])
for nag in na_grid:
dict_xda_list_element[nag].append(nc_file.data_vars['list_element__NaG=%d' % nag])
dict_xda_data_wave[nag].append(nc_file.data_vars['data_wave__NaG=%d' % nag])
# concat xarray.DataArray
xda_list_element_na = xr.concat(xda_list_element_na, dim='dim_element')
xda_list_element_coords = xr.concat(xda_list_element_coords, dim='dim_element')
for nag in na_grid:
dict_xda_list_element[nag] = xr.concat(dict_xda_list_element[nag], dim='dim_element__NaG=%d' % nag)
dict_xda_data_wave[nag] = xr.concat(dict_xda_data_wave[nag], dim='dim_element__NaG=%d' % nag)
# read data to numpy.ndarray
list_element_na = xda_list_element_na.values.astype(int)
list_element_coords = xda_list_element_coords.values
dict_list_element = {}
dict_data_wave = {}
for nag in na_grid:
dict_list_element[nag] = dict_xda_list_element[nag].values.astype(int)
if load_wave_data:
dict_data_wave[nag] = dict_xda_data_wave[nag].values
############### return ################
if load_wave_data:
return na_grid, data_time, list_element_na, list_element_coords, dict_list_element, dict_data_wave
else:
return na_grid, data_time, list_element_na, list_element_coords, dict_list_element, dict_xda_data_wave
```
# Ocean floor
## Function to extract waveform
```
# this function extracts waveforms with a given location (x, y)
def get_waveform_xy(xy, na_grid, list_element_na, list_element_coords, dict_data_wave,
channels=None, time_steps=None):
# (x, y) to (s, phi)
s = np.linalg.norm(xy)
phi = np.arctan2(xy[1], xy[0])
# point out of range
if s > np.max(list_element_coords[:, :, 0]):
return None
# deal with default input
if channels is None:
channels = np.arange(dict_data_wave[na_grid[0]].shape[3])
if time_steps is None:
time_steps = np.arange(dict_data_wave[na_grid[0]].shape[4])
########## step 1: inplane interpolation ##########
# find closest element using center coordinate
s_center = list_element_coords[:, 2, 0]
index_element = np.argmin(np.abs(s - s_center))
# find the two GLL points, A and B, between which s is located
s_element = list_element_coords[index_element, :, 0]
index_A = np.argmin(np.abs(s - s_element))
s_element_copy = s_element.copy()
# set s of A to a crazy value to find the second closest point
s_element_copy[index_A] = 1e100
index_B = np.argmin(np.abs(s - s_element_copy))
# interpolation factor at A
factor_A = 1. / (s_element[index_B] - s_element[index_A]) * (s_element[index_B] - s)
factor_B = 1 - factor_A
# na of closest elements
# the FIVE columes are:
# 0 element tag in mesh
# 1 actual nr
# 2 stored nr (or na grid)
# 3 element index in data (local)
# 4 element index in data (global)
element_na = list_element_na[index_element]
# read waveforms at A and B and do inplane interpolation
data_wave_A = dict_data_wave[element_na[2]][element_na[4], :, index_A][:, channels][:, :, time_steps]
data_wave_B = dict_data_wave[element_na[2]][element_na[4], :, index_B][:, channels][:, :, time_steps]
data_wave = data_wave_A * factor_A + data_wave_B * factor_B
########## step 2: Fourier interpolation ##########
# complex type
complex_type = np.complex32 if data_wave.dtype == np.complex64 else np.complex128
# maximum Fourier order
max_Fourier_order = element_na[1] // 2
# initialize result with 0th order
result = data_wave[0].copy()
# add higher orders
for order in np.arange(1, max_Fourier_order + 1):
coeff = np.zeros(result.shape, dtype=complex_type)
# real part
coeff.real = data_wave[order * 2 - 1]
# complex part of Fourier coefficients
if order * 2 < len(data_wave): # check for Nyquist
coeff.imag += data_wave[order * 2]
result += (2. * np.exp(1j * order * phi) * coeff).real
return result
```
## Read
```
# data dir
data_dir = './output/elements/Fourier_coefficients_ocean_floor'
# read
na_grid, data_time, list_element_na, list_element_coords, \
dict_list_element, dict_data_wave = read_element_output(data_dir)
```
## Extract and plot
```
# plot waveform at xy
xy = [2000, 3000]
wave_xy = get_waveform_xy(xy, na_grid, list_element_na, list_element_coords, dict_data_wave)
plt.figure(dpi=200)
for ich, ch in enumerate('RTZ'):
plt.subplot(3, 1, ich + 1)
plt.plot(data_time, wave_xy[ich, :], label=ch, c='C%d' % ich)
if ich < 2:
plt.xticks([])
else:
plt.xlabel('Time (s)')
if ich == 1:
plt.ylabel('Amplitude (m)')
plt.legend(loc='upper left')
plt.show()
```
# Inplane slices
## Read
```
# data dir
data_dir = './output/elements/orthogonal_azimuthal_slices'
# read
na_grid, data_time, list_element_na, list_element_coords, \
dict_list_element, dict_data_wave = read_element_output(data_dir)
```
## Generate animation on each slice
```
# wave dimension to animation
wave_channel = 'Z'
wave_dim = 'RTZ'.index(wave_channel)
# time steps
ntime = len(data_time)
# phi of the slices
phi_slices = np.radians(np.arange(0, 360, 90))
nslice = len(phi_slices)
# GLL coords on elements
nelem = list_element_coords.shape[0]
ngll = list_element_coords.shape[1]
# flattened coords, (s, z)
element_coords_sz = list_element_coords.reshape((nelem * ngll), 2)
# connectivity list, shared by all slices
# with GLL_points_one_edge = [0, 2, 4] in the inparam.output.yaml,
# the GLL point layout on each element is
# 0--1--2
# | | |
# 3--4--5
# | | |
# 6--7--8
connectivity = []
for ielem in np.arange(nelem):
start = ielem * 9
connectivity.append([start + 0, start + 1, start + 4, start + 3])
connectivity.append([start + 1, start + 2, start + 5, start + 4])
connectivity.append([start + 3, start + 4, start + 7, start + 6])
connectivity.append([start + 4, start + 5, start + 8, start + 7])
# loop over slices
for islice, phi in enumerate(phi_slices):
# create vtk folder
vtk_dir = data_dir + '/vtk/slice%d' % islice
os.makedirs(vtk_dir, exist_ok=True)
# vtk mesh
xyz = np.ndarray((nelem * ngll, 3))
xyz[:, 0] = element_coords_sz[:, 0] * np.cos(phi)
xyz[:, 1] = element_coords_sz[:, 0] * np.sin(phi)
xyz[:, 2] = element_coords_sz[:, 1]
vtk_mesh = pyvtk.UnstructuredGrid(list(zip(xyz[:,0], xyz[:,1], xyz[:,2])),
quad=connectivity)
# loop over elements to read wave data
wave = np.ndarray((nelem * ngll, ntime))
for ielem in np.arange(nelem):
wave[(ielem * ngll):(ielem * ngll + ngll), :] = dict_data_wave[nslice][ielem, islice, :, wave_dim, :]
# loop over time to write vtk
for itime in np.arange(ntime):
vtk = pyvtk.VtkData(vtk_mesh, pyvtk.PointData(pyvtk.Scalars(wave[:, itime], name='U' + wave_channel)))
vtk.tofile(vtk_dir + '/' + 'wave%d.vtk' % itime, 'binary')
print('Done time step %d / %d' % (itime + 1, ntime), end='\r')
print('\nDone slice %d' % (islice + 1))
```
| github_jupyter |
# Image Deduplication with FiftyOne
This recipe demonstrates a simple use case of using FiftyOne to detect and
remove duplicate images from your dataset.
## Requirements
This notebook requires the `tensorflow` package:
```
!pip install tensorflow
```
## Download the data
First we download the dataset to disk. The dataset is a 1000 sample subset of
CIFAR-100, a dataset of 32x32 pixel images with one of 100 different
classification labels such as `apple`, `bicycle`, `porcupine`, etc.
```
from image_deduplication_helpers import download_dataset
download_dataset()
```
The above script uses `tensorflow.keras.datasets` to download the dataset, so
you must have [TensorFlow installed](https://www.tensorflow.org/install).
The dataset is organized on disk as follows:
```
/tmp/fiftyone/
└── cifar100_with_duplicates/
├── <classA>/
│ ├── <image1>.jpg
│ ├── <image2>.jpg
│ └── ...
├── <classB>/
│ ├── <image1>.jpg
│ ├── <image2>.jpg
│ └── ...
└── ...
```
As we will soon come to discover, some of these samples are duplicates and we
have no clue which they are!
## Create a dataset
Let's start by importing the FiftyOne library:
```
import fiftyone as fo
```
Let's use a utililty method provided by FiftyOne to load the image
classification dataset from disk:
```
import os
dataset_name = "cifar100_with_duplicates"
dataset_dir = os.path.join("/tmp/fiftyone", dataset_name)
dataset = fo.Dataset.from_dir(
dataset_dir,
fo.types.ImageClassificationDirectoryTree,
name=dataset_name
)
```
## Explore the dataset
We can poke around in the dataset:
```
# Print summary information about the dataset
print(dataset)
# Print a sample
print(dataset.first())
```
Create a view that contains only samples whose ground truth label is
`mountain`:
```
# Used to write view expressions that involve sample fields
from fiftyone import ViewField as F
view = dataset.match(F("ground_truth.label") == "mountain")
# Print summary information about the view
print(view)
# Print the first sample in the view
print(view.first())
```
Create a view with samples sorted by their ground truth labels in reverse
alphabetical order:
```
view = dataset.sort_by("ground_truth.label", reverse=True)
# Print summary information about the view
print(view)
# Print the first sample in the view
print(view.first())
```
## Visualize the dataset
Start browsing the dataset:
```
session = fo.launch_app(dataset=dataset)
```

Narrow your scope to 10 random samples:
```
session.view = dataset.take(10)
```

Click on some some samples in the GUI to select them and access their IDs from
code!
```
# Get the IDs of the currently selected samples in the App
sample_ids = session.selected
```
Create a view that contains your currently selected samples:
```
selected_view = dataset.select(session.selected)
```
Update the App to only show your selected samples:
```
session.view = selected_view
```

## Compute file hashes
Iterate over the samples and compute their file hashes:
```
import fiftyone.core.utils as fou
for sample in dataset:
sample["file_hash"] = fou.compute_filehash(sample.filepath)
sample.save()
print(dataset)
```
We have two ways to visualize this new information.
First, you can view the sample from your Terminal:
```
sample = dataset.first()
print(sample)
```
Or you can refresh the App and toggle on the new `file_hash` field:
```
session.dataset = dataset
```

## Check for duplicates
Now let's use a simple Python statement to locate the duplicate files in the
dataset, i.e., those with the same file hashses:
```
from collections import Counter
filehash_counts = Counter(sample.file_hash for sample in dataset)
dup_filehashes = [k for k, v in filehash_counts.items() if v > 1]
print("Number of duplicate file hashes: %d" % len(dup_filehashes))
```
Now let's create a view that contains only the samples with these duplicate
file hashes:
```
dup_view = (dataset
# Extract samples with duplicate file hashes
.match(F("file_hash").is_in(dup_filehashes))
# Sort by file hash so duplicates will be adjacent
.sort_by("file_hash")
)
print("Number of images that have a duplicate: %d" % len(dup_view))
print("Number of duplicates: %d" % (len(dup_view) - len(dup_filehashes)))
```
Of course, we can always use the App to visualize our work!
```
session.view = dup_view
```

## Delete duplicates
Now let's delete the duplicate samples from the dataset using our `dup_view` to
restrict our attention to known duplicates:
```
print("Length of dataset before: %d" % len(dataset))
_dup_filehashes = set()
for sample in dup_view:
if sample.file_hash not in _dup_filehashes:
_dup_filehashes.add(sample.file_hash)
continue
del dataset[sample.id]
print("Length of dataset after: %d" % len(dataset))
# Verify that the dataset no longer contains any duplicates
print("Number of unique file hashes: %d" % len({s.file_hash for s in dataset}))
```
## Export the deduplicated dataset
Finally, let's export a fresh copy of our now-duplicate-free dataset:
```
EXPORT_DIR = "/tmp/fiftyone/image-deduplication"
dataset.export(label_field="ground_truth", export_dir=EXPORT_DIR)
```
Check out the contents of `/tmp/fiftyone/image-deduplication` on disk to see how the data is
organized.
You can load the deduplicated dataset that you exported back into FiftyOne at
any time as follows:
```
no_dups_dataset = fo.Dataset.from_dir(
EXPORT_DIR,
fo.types.FiftyOneImageClassificationDataset,
name="no_duplicates",
)
print(no_dups_dataset)
```
## Cleanup
You can cleanup the files generated by this recipe by running:
```
!rm -rf /tmp/fiftyone
```
| github_jupyter |
# Comprehensive Guide to Grouping and Aggregating with Pandas
Chris Mofitt. "Comprehensive Guide to Grouping and Aggregating with Pandas". _Practical Business Python_, 9 Nov. 2020, https://pbpython.com/groupby-agg.html.
```
import pandas as pd
import seaborn as sns
df = sns.load_dataset('titanic')
```
## Pandas aggregation options
### List
```
df['fare'].agg(['sum', 'mean'])
```
### Dictionary
```
df.agg({'fare': ['sum', 'mean'],
'sex': ['count']})
```
### Tuple
```
df.agg(fare_sum=('fare', 'sum'),
fare_mean=('fare', 'mean'),
sex_count=('sex', 'count'))
```
## Groupby
### Basic math
```
agg_func_math = {
'fare':
['sum', 'mean', 'median', 'min', 'max', 'std', 'var', 'mad', 'prod']
}
df.groupby(['embark_town']).agg(agg_func_math).round(2)
```
Use describe to run multiple built-in aggregations at once:
```
agg_func_describe = {'fare': ['describe']}
df.groupby(['embark_town']).agg(agg_func_describe).round(2)
```
### Counting
```
agg_func_count = {'embark_town': ['count', 'nunique', 'size']}
df.groupby(['deck']).agg(agg_func_count)
```
### First and last
Select highest and lowest fare by embarked town (need to sort first to have first and last pick max and min values).
```
agg_func_selection = {'fare': ['first', 'last']}
df.sort_values(by=['fare'],
ascending=False).groupby(['embark_town'
]).agg(agg_func_selection)
```
Instead use idxmax and idxmin to select values that correspond to max and min:
```
agg_func_max_min = {'fare': ['idxmax', 'idxmin']}
df.groupby(['embark_town']).agg(agg_func_max_min)
df.loc[[258, 378]]
df.loc[df.groupby('class')['fare'].idxmax()]
```
### Other libraries
```
from scipy.stats import skew, mode
agg_func_stats = {'fare': [skew, mode, pd.Series.mode]}
df.groupby(['embark_town']).agg(agg_func_stats)
```
### Working with text
```
agg_func_text = {'deck': [ 'nunique', mode, set]}
df.groupby(['class']).agg(agg_func_text)
```
### Custom Functions
Calculate the 25th percentile of the data using four approaches.
First, partial function:
```
from functools import partial
# Use partial
q_25 = partial(pd.Series.quantile, q=0.25)
q_25.__name__ = '25%'
# Define a function
def percentile_25(x):
return x.quantile(.25)
# Define a lambda function
lambda_25 = lambda x: x.quantile(.25)
lambda_25.__name__ = 'lambda_25%'
# Use a lambda function inline
agg_func = {
'fare': [q_25, percentile_25, lambda_25, lambda x: x.quantile(.25)]
}
df.groupby(['embark_town']).agg(agg_func).round(2)
```
### Custom function examples
Count number of null values:
```
def count_nulls(s):
return s.size - s.count()
```
Include NaN values in unique counts:
```
def unique_nan(s):
return s.nunique(dropna=False)
```
Summary of all values together:
```
agg_func_custom_count = {
'embark_town': ['count', 'nunique', 'size', unique_nan, count_nulls, set]
}
df.groupby(['deck']).agg(agg_func_custom_count)
```
To calculate the 90th percentile, use quantile:
```
def percentile_90(x):
return x.quantile(.9)
```
For trimmed mean where lowest 10th percent is excluded, use scipy status function:
```
from scipy.stats import trim_mean
def trim_mean_10(x):
return trim_mean(x, 0.1)
```
For largest value, regardless of sort order:
```
def largest(x):
return x.nlargest(1)
```
Incorporate [sparklines](https://pbpython.com/styling-pandas.html):
```
from sparklines import sparklines
import numpy as np
def sparkline_str(x):
bins=np.histogram(x)[0]
sl = ''.join(sparklines(bins))
return sl
```
All put together:
```
agg_func_largest = {
'fare': [percentile_90, trim_mean_10, largest, sparkline_str]
}
df.groupby(['class', 'embark_town']).agg(agg_func_largest)
```
Get total fares for top 10 and bottom 10:
```
def top_10_sum(x):
return x.nlargest(10).sum()
def bottom_10_sum(x):
return x.nsmallest(10).sum()
agg_func_top_bottom_sum = {
'fare': [top_10_sum, bottom_10_sum]
}
df.groupby('class').agg(agg_func_top_bottom_sum)
```
### Custom functions with multiple columns
Use groupby combined with apply:
```
def summary(x):
result = {
'fare_sum': x['fare'].sum(),
'fare_mean': x['fare'].mean(),
'fare_range': x['fare'].max() - x['fare'].min()
}
return pd.Series(result).round(0)
df.groupby(['class']).apply(summary)
```
## Working with group objects
Figure what percentage of total fares sold can be attributed to each embark_town and class combination (using assign and lambda function to add a pct_total column):
```
df.groupby(['embark_town', 'class']).agg({
'fare': 'sum'
}).assign(pct_total=lambda x: x / x.sum())
```
Simpler to use [pd.crosstab](https://pbpython.com/pandas-crosstab.html):
```
pd.crosstab(df['embark_town'],
df['class'],
values=df['fare'],
aggfunc='sum',
normalize=True)
```
Combine agg functions with pivot table:
```
pd.pivot_table(data=df,
index=['embark_town'],
columns=['class'],
aggfunc=agg_func_top_bottom_sum)
```
Show cumulative total of fares by group and aggregate by town and class, then group:
```
fare_group = df.groupby(['embark_town', 'class']).agg({'fare': 'sum'})
fare_group.groupby(level=0).cumsum()
```
Summarize daily sales and convert to cumulative daily and quarterly view (use [pd.Grouper](https://pbpython.com/pandas-grouper-agg.html)).
Here, include total daily sales as well as cumulative quarter amount:
```
sales = pd.read_excel('https://github.com/chris1610/pbpython/blob/master/data/2018_Sales_Total_v2.xlsx?raw=True')
daily_sales = sales.groupby([pd.Grouper(key='date', freq='D')
]).agg(daily_sales=('ext price',
'sum')).reset_index()
daily_sales['quarter_sales'] = daily_sales.groupby(
pd.Grouper(key='date', freq='Q')).agg({'daily_sales': 'cumsum'})
```
Group daily results, then group by quarter and use cumulative sum:
```
sales.groupby([pd.Grouper(key='date', freq='D')
]).agg(daily_sales=('ext price', 'sum')).groupby(
pd.Grouper(freq='Q')).agg({
'daily_sales': 'cumsum'
}).rename(columns={'daily_sales': 'quarterly_sales'})
```
## Flattening Hierarchical Column Indices
```
df.groupby(['embark_town', 'class']).agg({'fare': ['sum', 'mean']}).round(0)
multi_df = df.groupby(['embark_town', 'class'],
as_index=False).agg({'fare': ['sum', 'mean']})
multi_df.columns = [
'_'.join(col).rstrip('_') for col in multi_df.columns.values
]
```
## Subtotals
Add a subtotal using the [sidetable](https://github.com/chris1610/sidetable) package.
```
import sidetable
df.groupby(['class', 'embark_town', 'sex']).agg({'fare': 'sum'}).stb.subtotal()
```
| github_jupyter |
# Tabular Datasets
As we have already discovered, Elements are simple wrappers around your data that provide a semantically meaningful representation. HoloViews can work with a wide variety of data types, but many of them can be categorized as either:
* **Tabular:** Tables of flat columns, or
* **Gridded:** Array-like data on 2-dimensional or N-dimensional grids
These two general data types are explained in detail in the [Tabular Data](../user_guide/07-Tabular_Datasets.ipynb) and [Gridded Data](../user_guide/08-Gridded_Datasets.ipynb) user guides, including all the many supported formats (including Python dictionaries of NumPy arrays, pandas ``DataFrames``, dask ``DataFrames``, and xarray ``DataArrays`` and ``Datasets``).
In this Getting-Started guide we provide a quick overview and introduction to two of the most flexible and powerful formats: columnar **pandas** DataFrames (in this section), and gridded **xarray** Datasets (in the next section).
## Tabular
Tabular data (also called columnar data) is one of the most common, general, and versatile data formats, corresponding to how data is laid out in a spreadsheet. There are many different ways to put data into a tabular format, but for interactive analysis having [**tidy data**](http://www.jeannicholashould.com/tidy-data-in-python.html) provides flexibility and simplicity. For tidy data, the **columns** of the table represent **variables** or **dimensions** and the **rows** represent **observations**. The best way to understand this format is to look at such a dataset:
```
import numpy as np
import pandas as pd
import holoviews as hv
hv.extension('bokeh', 'matplotlib')
diseases = pd.read_csv('../assets/diseases.csv.gz')
diseases.head()
```
This particular dataset was the subject of an excellent piece of visual journalism in the [Wall Street Journal](http://graphics.wsj.com/infectious-diseases-and-vaccines/#b02g20t20w15). The WSJ data details the incidence of various diseases over time, and was downloaded from the [University of Pittsburgh's Project Tycho](http://www.tycho.pitt.edu/). We can see we have 5 data columns, which each correspond either to independent variables that specify a particular measurement ('Year', 'Week', 'State'), or observed/dependent variables reporting what was then actually measured (the 'measles' or 'pertussis' incidence).
Knowing the distinction between those two types of variables is crucial for doing visualizations, but unfortunately the tabular format does not declare this information. Plotting 'Week' against 'State' would not be meaningful, whereas 'measles' for each 'State' (averaging or summing across the other dimensions) would be fine, and there's no way to deduce those constraints from the tabular format. Accordingly, we will first make a HoloViews object called a ``Dataset`` that declares the independent variables (called key dimensions or **kdims** in HoloViews) and dependent variables (called value dimensions or **vdims**) that you want to work with:
```
vdims = [('measles', 'Measles Incidence'), ('pertussis', 'Pertussis Incidence')]
ds = hv.Dataset(diseases, ['Year', 'State'], vdims)
```
Here we've used an optional tuple-based syntax **``(name,label)``** to specify a more meaningful description for the ``vdims``, while using the original short descriptions for the ``kdims``. We haven't yet specified what to do with the ``Week`` dimension, but we are only interested in yearly averages, so let's just tell HoloViews to average over all remaining dimensions:
```
ds = ds.aggregate(function=np.mean)
ds
```
(We'll cover aggregations like ``np.mean`` in detail later, but here the important bit is simply that the ``Week`` dimension can now be ignored.)
The ``repr`` shows us both the ``kdims`` (in square brackets) and the ``vdims`` (in parentheses) of the ``Dataset``. Because it can hold arbitrary combinations of dimensions, a ``Dataset`` is *not* immediately visualizable. There's no single clear mapping from these four dimensions onto a two-dimensional page, hence the textual representation shown above.
To make this data visualizable, we'll need to provide a bit more metadata, by selecting one of the large library of Elements that can help answer the questions we want to ask about the data. Perhaps the most obvious representation of this dataset is as a ``Curve`` displaying the incidence for each year, for each state. We could pull out individual columns one by one from the original dataset, but now that we have declared information about the dimensions, the cleanest approach is to map the dimensions of our ``Dataset`` onto the dimensions of an Element using ``.to``:
```
%%opts Curve [width=600 height=250] {+framewise}
(ds.to(hv.Curve, 'Year', 'measles') + ds.to(hv.Curve, 'Year', 'pertussis')).cols(1)
```
Here we specified two ``Curve`` elements showing measles and pertussis incidence respectively (the vdims), per year (the kdim), and laid them out in a vertical column. You'll notice that even though we specified only the short name for the value dimensions, the plot shows the longer names ("Measles Incidence", "Pertussis Incidence") that we declared on the ``Dataset``.
You'll also notice that we automatically received a dropdown menu to select which ``State`` to view. Each ``Curve`` ignores unused value dimensions, because additional measurements don't affect each other, but HoloViews has to do *something* with every key dimension for every such plot. If the ``State`` (or any other key dimension) isn't somehow plotted or aggregated over, then HoloViews has to leave choosing a value for it to the user, hence the selection widget. Other options for what to do with extra dimensions or just extra data ranges are illustrated below.
### Selecting
One of the most common things we might want to do is to select only a subset of the data. The ``select`` method makes this extremely easy, letting you select a single value, a list of values supplied as a list, or a range of values supplied as a tuple. Here we will use ``select`` to display the measles incidence in four states over one decade. After applying the selection, we use the ``.to`` method as shown earlier, now displaying the data as ``Bars`` indexed by 'Year' and 'State' key dimensions and displaying the 'Measles Incidence' value dimension:
```
%%opts Bars [width=800 height=400 tools=['hover'] xrotation=90 show_legend=False]
states = ['New York', 'New Jersey', 'California', 'Texas']
ds.select(State=states, Year=(1980, 1990)).to(hv.Bars, ['Year', 'State'], 'measles').sort()
```
### Faceting
Above we already saw what happens to key dimensions that we didn't explicitly assign to the Element using the ``.to`` method: they are grouped over, popping up a set of widgets so the user can select the values to show at any one time. However, using widgets is not always the most effective way to view the data, and a ``Dataset`` lets you specify other alternatives using the ``.overlay``, ``.grid`` and ``.layout`` methods. For instance, we can lay out each state separately using ``.grid``:
```
%%opts Curve [width=200] (color='indianred')
grouped = ds.select(State=states, Year=(1930, 2005)).to(hv.Curve, 'Year', 'measles')
grouped.grid('State')
```
Or we can take the same grouped object and ``.overlay`` the individual curves instead of laying them out in a grid:
```
%%opts Curve [width=600] (color=Cycle(values=['indianred', 'slateblue', 'lightseagreen', 'coral']))
grouped.overlay('State')
```
These faceting methods even compose together, meaning that if we had more key dimensions we could ``.overlay`` one dimension, ``.grid`` another and have a widget for any other remaining key dimensions.
### Aggregating
Instead of selecting a subset of the data, another common operation supported by HoloViews is computing aggregates. When we first loaded this dataset, we aggregated over the 'Week' column to compute the mean incidence for every year, thereby reducing our data significantly. The ``aggregate`` method is therefore very useful to compute statistics from our data.
A simple example using our dataset is to compute the mean and standard deviation of the Measles Incidence by ``'Year'``. We can express this simply by passing the key dimensions to aggregate over (in this case just the 'Year') along with a function and optional ``spreadfn`` to compute the statistics we want. The ``spread_fn`` will append the name of the function to the dimension name so we can reference the computed value separately. Once we have computed the aggregate, we can simply cast it to a ``Curve`` and ``ErrorBars``:
```
%%opts Curve [width=600]
agg = ds.aggregate('Year', function=np.mean, spreadfn=np.std)
(hv.Curve(agg) * hv.ErrorBars(agg,vdims=['measles', 'measles_std']).iloc[::2]).redim.range(measles=(0, None))
```
In this way we can summarize a multi-dimensional dataset as something that can be visualized directly, while allowing us to compute arbitrary statistics along a dimension.
## Other data
If you want to know more about working with tabular data, particularly when using datatypes other than pandas, have a look at the [user guide](../user_guide/07-Tabular_Datasets.ipynb). The different interfaces allow you to work with everything from simple NumPy arrays to out-of-core dataframes using dask. Dask dataframes scale to visualizations of billions of rows, when using [datashader](https://anaconda.org/jbednar/holoviews_datashader/notebook) with HoloViews to aggregate the data as needed.
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
import os, sys
sys.path.insert(0, os.path.expandvars('/data/users/$USER/fbsource/fbcode/beanmachine'))
sys.path.insert(1, os.path.expandvars('/data/users/$USER/fbsource/third-party/pypi/flowtorch/0.0.dev2'))
import beanmachine.ppl as bm
import beanmachine.ppl as bm
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
import statsmodels.graphics.gofplots
import scipy.stats as stats
import torch
import torch.distributions as dist
import torch.nn as nn
from beanmachine.ppl.experimental.vi.variational_infer import (
MeanFieldVariationalInference
)
from beanmachine.ppl.distributions import Flat
sns.set_style('darkgrid')
```
# Bayesian Robust Regression
```
n = 100
d = 10
df = 5.0
X_train = torch.randn(n, d)
beta_truth = torch.randn(d+1, 1)
y_train = torch.cat((X_train, torch.ones(n, 1)), -1).mm(beta_truth) \
+ dist.StudentT(df=4.0).sample((n, 1))
@bm.random_variable
def beta():
return dist.Independent(
dist.StudentT(df=4.0*torch.ones(d+1)),
1,
)
# return Flat(shape=(d+1))
@bm.random_variable
def X():
return dist.Normal(0, 1) # dummy
@bm.random_variable
def y():
X_with_ones = torch.cat((X(), torch.ones(X().shape[0], 1)), -1)
b = beta().squeeze()
if b.dim() == 1:
b = b.unsqueeze(0)
mu = b.mm(X_with_ones.T).T
return dist.Independent(
dist.StudentT(df=df, loc=mu, scale=1),
1,
)
def annotate(data, **kws):
coord = data['variable'].unique()
assert len(coord) == 1
ax = plt.gca()
ax.axvline(x=beta_truth[coord[0]], color='r')
vi_dicts_normal = MeanFieldVariationalInference().infer(
queries=[beta()],
observations={
X(): X_train,
y(): y_train,
},
base_dist=dist.Normal,
base_args={
'loc': nn.Parameter(torch.tensor(0.0)),
'scale': nn.Parameter(torch.tensor(1.0)),
},
num_iter=10,
)
vi_dicts_student = MeanFieldVariationalInference().infer(
queries=[beta()],
observations={
X(): X_train,
y(): y_train,
},
base_dist=dist.StudentT,
base_args={
'df': nn.Parameter(torch.tensor(df)),
'loc': nn.Parameter(torch.tensor(0.0)),
'scale': nn.Parameter(torch.tensor(1.0)),
},
num_iter=10,
)
# N = 1000
# g = sns.displot(
# data=pd.concat((
# pd.DataFrame(vi_dicts_normal(beta()).sample((N,)).numpy()).assign(method='vi_normal'),
# pd.DataFrame(vi_dicts_student(beta()).sample((N,)).numpy()).assign(method='vi_student'),
# )).melt(id_vars='method'),
# x='value',
# col='variable',
# col_wrap=3,
# hue='method',
# )
# g.map_dataframe(annotate)
```
Log pointwise predictive density? [link](https://arxiv.org/pdf/1307.5928.pdf), underestimates $E_\pi \log p(\tilde{y} \mid \hat\theta)$ for a given point esitmate $\hat\theta(y)$
```
n_test = 1000
n_posterior = 100
X_test = torch.randn(n_test, d)
y_test = torch.cat((X_test, torch.ones(n_test, 1)), -1).mm(beta_truth) \
+ dist.StudentT(df=4.0).sample((n_test, 1))
for vi_dicts in [
vi_dicts_normal,
vi_dicts_student,
]:
samples = vi_dicts(beta()).sample((n_posterior,))
lppds = torch.stack([
dist.StudentT(df=df).log_prob(y_test - torch.cat((X_test, torch.ones(n_test, 1)), -1).mm(b.unsqueeze(1))).sum() for b in torch.unbind(samples, dim=0)
], dim=0)
print(lppds.mean())
```
QQ plots should show tail behavior
```
n_posterior = int(1e5)
statsmodels.graphics.gofplots.qqplot(
vi_dicts_normal(beta()).sample((n_posterior,))[:,6],
fit=True,
line="45",
dist=stats.t,
distargs=(df,)
)
statsmodels.graphics.gofplots.qqplot(
vi_dicts_student(beta()).sample((n_posterior,))[:,6],
fit=True,
line="45",
dist=stats.t,
distargs=(df,)
)
```
Random Walk MH does poorly here
```
samples = bm.SingleSiteRandomWalk().infer(
queries=[beta()],
observations={
X(): X_train,
y(): y_train,
},
num_samples=1000,
num_chains=1,
)
# g = sns.displot(
# pd.DataFrame(samples[beta()].squeeze().numpy()).melt(),
# col='variable',
# col_wrap=3,
# x='value',
# )
# g.map_dataframe(annotate)
```
| github_jupyter |
```
import pandas as pd
import numpy as np
from tqdm import tqdm_notebook
from sklearn.feature_extraction.text import TfidfTransformer, CountVectorizer, TfidfVectorizer
from sklearn.neighbors import NearestNeighbors
%matplotlib inline
links = pd.read_csv('links.csv')
movies = pd.read_csv('movies.csv')
ratings = pd.read_csv('ratings.csv')
tags = pd.read_csv('tags.csv')
movies.head(10)
def change_string(s):
return ' '.join(s.replace(' ', '').replace('-', '').split('|'))
movie_genres = [change_string(g) for g in movies.genres.values]
movie_genres[:10]
count_vect = CountVectorizer()
X_train_counts = count_vect.fit_transform(movie_genres)
X_train_counts
count_vect.vocabulary_
X_train_counts.toarray()
tfidf_transformer = TfidfTransformer()
X_train_tfidf = tfidf_transformer.fit_transform(X_train_counts)
## Альтернативный способ
tfidf_vectorizer = TfidfVectorizer()
X_train_tfidf = tfidf_vectorizer.fit_transform(movie_genres)
tfidf_vectorizer.vocabulary_
X_train_tfidf.toarray()
neigh = NearestNeighbors(n_neighbors=7, n_jobs=-1, metric='euclidean')
neigh.fit(X_train_tfidf)
test = change_string("Adventure|Comedy|Fantasy|Crime")
X_tfidf2 = tfidf_vectorizer.transform([test])
res = neigh.kneighbors(X_tfidf2, return_distance=True)
res
movies.iloc[res[1][0]]
movies.head()
tags.head()
movies_with_tags = movies.join(tags.set_index('movieId'), on='movieId')
movies_with_tags.head()
movies_with_tags[movies_with_tags.title == 'Toy Story (1995)']
movies_with_tags.tag.unique()
movies_with_tags.dropna(inplace=True)
movies_with_tags.title.unique().shape
tag_strings = []
movies = []
for movie, group in tqdm_notebook(movies_with_tags.groupby('title')):
tag_strings.append(' '.join([str(s).replace(' ', '').replace('-', '') for s in group.tag.values]))
movies.append(movie)
tag_strings[:5]
count_vect = CountVectorizer()
X_train_counts = count_vect.fit_transform(tag_strings)
tfidf_transformer = TfidfTransformer()
X_train_tfidf = tfidf_transformer.fit_transform(X_train_counts)
neigh = NearestNeighbors(n_neighbors=10, n_jobs=-1, metric='manhattan')
neigh.fit(X_train_tfidf)
for i in range(len(movies)):
if 'Magnolia (1999)' == movies[i]:
print(i)
tag_strings[822]
test = change_string('pixar pixar fun')
predict = count_vect.transform([test])
X_tfidf2 = tfidf_transformer.transform(predict)
res = neigh.kneighbors(X_tfidf2, return_distance=True)
res
for i in res[1][0]:
print(movies[i])
movies
movies
tfidfs_on_genre = X_train_tfidf.toarray()
tfidfs_on_genre.shape
for x in range(tfidfs_on_genre.shape[1]):
col_name = 'g{}'.format(x)
movies[col_name] = pd.Series(tfidfs_on_genre[:, x])
movies
def extract_year(s):
try:
return int(s[-5:-1])
except:
return 0
movies['year'] = movies['title'].apply(extract_year)
movies
movies = pd.get_dummies(movies, columns=['year'])
to_train = movies.columns
to_train = [s for i, s in enumerate(movies.columns) if i > 1]
to_train_df = movies[to_train]
to_train_df
```
| github_jupyter |
## Unsupervised Learning
## Project: Creating Customer Segments
## Getting Started
In this project analyzed a dataset containing data on various customers' annual spending amounts (reported in *monetary units*) of diverse product categories for internal structure. One goal of this project is to best describe the variation in the different types of customers that a wholesale distributor interacts with. Doing so would equip the distributor with insight into how to best structure their delivery service to meet the needs of each customer.
The dataset for this project can be found on the [UCI Machine Learning Repository](https://archive.ics.uci.edu/ml/datasets/Wholesale+customers). For the purposes of this project, the features `'Channel'` and `'Region'` will be excluded in the analysis — with focus instead on the six product categories recorded for customers.
Run the code block below to load the wholesale customers dataset, along with a few of the necessary Python libraries required for this project. The dataset loaded successfully if the size of the dataset is reported.
```
# Import libraries necessary for this project
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib as plt
from IPython.display import display # Allows the use of display() for DataFrames
# Import supplementary visualizations code visuals.py
import visuals as vs
# Pretty display for notebooks
%matplotlib inline
# Load the wholesale customers dataset
try:
data = pd.read_csv("customers.csv")
data.drop(['Region', 'Channel'], axis = 1, inplace = True)
print("Wholesale customers dataset has {} samples with {} features each.".format(*data.shape))
except:
print("Dataset could not be loaded. Is the dataset missing?")
```
## Data Exploration
In this section, we will begin exploring the data through visualizations and code to understand how each feature is related to the others. We will observe a statistical description of the dataset, consider the relevance of each feature, and select a few sample data points from the dataset which you will track through the course of this project.
Run the code block below to observe a statistical description of the dataset. Note that the dataset is composed of six important product categories: **'Fresh'**, **'Milk'**, **'Grocery'**, **'Frozen'**, **'Detergents_Paper'**, and **'Delicatessen'**. Consider what each category represents in terms of products you could purchase.
```
# Display a description of the dataset
display(data.describe())
```
### Implementation: Selecting Samples
To get a better understanding of the customers and how their data will transform through the analysis, it would be best to select a few sample data points and explore them in more detail. In the code block below, add **three** indices of your choice to the `indices` list which will represent the customers to track. It is suggested to try different sets of samples until obtain customers that vary significantly from one another.
```
# Select three indices of your choice you wish to sample from the dataset
indices = [64, 128, 256]
# Create a DataFrame of the chosen samples
samples = pd.DataFrame(data.loc[indices], columns = data.keys()).reset_index(drop = True)
print "Chosen samples of wholesale customers dataset:"
display(samples)
# Visualize data on heatmap
plt.rcParams['figure.figsize']=(8,6)
ptl = 100 * data.rank(pct=True).round(decimals=4)
ptl = ptl.iloc[indices]
sns.heatmap(ptl, vmin=1, vmax=100, annot=True, cbar=False, square=True)
```
### Question 1
Consider the total purchase cost of each product category and the statistical description of the dataset above for your sample customers.
*What kind of establishment (customer) could each of the three samples you've chosen represent?*
**Hint:** Examples of establishments include places like markets, cafes, and retailers, among many others. Avoid using names for establishments, such as saying *"McDonalds"* when describing a sample customer as a restaurant.
**Answer:**
Data from indices [64, 128, 256] are aggregated to heatmaps of the percentiles (1-100).
<br>
**[64]** - Restaurant (sushi-shop):<br>
A lot of frozen foods, delicatessen, and detergents (required in big amounts after work with a fish products).
<br><br>
**[128]** - Small grossery / Shop in the farm market:<br>
82th percentile of Milk as well as 44th percentile of Grocery and 56th percentile of Detergents.
<br><br>
**[256]** - Supermarket:<br>
55-62th percentile of Fresh/Milk/Grocery and 17-35th percentile of Frozen food, Detergents, and Delicatessen. It's a typical distribution for average/small Supermarket.
### Implementation: Feature Relevance
One interesting thought to consider is if one (or more) of the six product categories is actually relevant for understanding customer purchasing. That is to say, is it possible to determine whether customers purchasing some amount of one category of products will necessarily purchase some proportional amount of another category of products? We can make this determination quite easily by training a supervised regression learner on a subset of the data with one feature removed, and then score how well that model can predict the removed feature.
In the code block below, you will need to implement the following:
- Assign `new_data` a copy of the data by removing a feature of your choice using the `DataFrame.drop` function.
- Use `sklearn.cross_validation.train_test_split` to split the dataset into training and testing sets.
- Use the removed feature as your target label. Set a `test_size` of `0.25` and set a `random_state`.
- Import a decision tree regressor, set a `random_state`, and fit the learner to the training data.
- Report the prediction score of the testing set using the regressor's `score` function.
```
from sklearn.model_selection import train_test_split
from sklearn.tree import DecisionTreeRegressor
# Make a copy of the DataFrame, using the 'drop' function to drop the given feature
drop_feature = "Grocery"
new_data = data.drop(drop_feature, axis=1)
# Split the data into training and testing sets using the given feature as the target
X_train, X_test, y_train, y_test = train_test_split(new_data, data[drop_feature],
test_size=0.25, random_state=0)
# Create a decision tree regressor and fit it to the training set
regressor = DecisionTreeRegressor(random_state=0)
regressor.fit(X_train, y_train)
# Report the score of the prediction using the testing set
score = regressor.score(X_test, y_test)
print ("Score for '{0}': {1}".format(drop_feature, score))
print ("\nFeature importance:")
print (list(new_data))
print (regressor.feature_importances_)
```
### Question 2
*Which feature did you attempt to predict? What was the reported prediction score? Is this feature necessary for identifying customers' spending habits?*
**Hint:** The coefficient of determination, `R^2`, is scored between 0 and 1, with 1 being a perfect fit. A negative `R^2` implies the model fails to fit the data.
**Answer:**
Feature to predict: 'Grocery' <br>
R^2 score = 0.6 <br>
Follow the 'feature_importances' of the regressor, the 'Grocery' feature is influenced by the 'Detergents_Paper' feature, the 'Grocery' feature is not completely necessary to identify consumer spending habits since it's highly correlated with 'Detergents_Paper'.
### Visualize Feature Distributions
To get a better understanding of the dataset, we can construct a scatter matrix of each of the six product features present in the data. If you found that the feature you attempted to predict above is relevant for identifying a specific customer, then the scatter matrix below may not show any correlation between that feature and the others. Conversely, if you believe that feature is not relevant for identifying a specific customer, the scatter matrix might show a correlation between that feature and another feature in the data. Run the code block below to produce a scatter matrix.
```
# Produce a scatter matrix for each pair of features in the data
pd.plotting.scatter_matrix(data, alpha = 0.3, figsize = (14,8), diagonal = 'kde');
# Plot heatmap for each pair of features in the data
sns.heatmap(data.corr(), annot=True)
```
### Question 3
*Are there any pairs of features which exhibit some degree of correlation? Does this confirm or deny your suspicions about the relevance of the feature you attempted to predict? How is the data for those features distributed?*
**Hint:** Is the data normally distributed? Where do most of the data points lie?
**Answer:**
Correlations:<br>
* 'Grocery - Detergents_Paper': linear correlation with coefficient of 0.92 (heatmap).
* 'Grocery - Milk': linear correlation with coefficient of 0.73 (heatmap).
* 'Milk - Detergents_Paper': linear correlation with coefficient of 0.66 (heatmap).
The data points aren't normally distributed: most of the dataset is skewed in the lower regions (range 0-10000) for most of the products, but Fresh and Groceries are in the range 0-20000.
## Data Preprocessing
In this section, you will preprocess the data to create a better representation of customers by performing a scaling on the data and detecting (and optionally removing) outliers. Preprocessing data is often times a critical step in assuring that results you obtain from your analysis are significant and meaningful.
### Implementation: Feature Scaling
If data is not normally distributed, especially if the mean and median vary significantly (indicating a large skew), it is most [often appropriate](http://econbrowser.com/archives/2014/02/use-of-logarithms-in-economics) to apply a non-linear scaling — particularly for financial data. One way to achieve this scaling is by using a [Box-Cox test](http://scipy.github.io/devdocs/generated/scipy.stats.boxcox.html), which calculates the best power transformation of the data that reduces skewness. A simpler approach which can work in most cases would be applying the natural logarithm.
In the code block below, you will need to implement the following:
- Assign a copy of the data to `log_data` after applying logarithmic scaling. Use the `np.log` function for this.
- Assign a copy of the sample data to `log_samples` after applying logarithmic scaling. Again, use `np.log`.
```
# Scale the data using the natural logarithm
log_data = np.log(data)
# Scale the sample data using the natural logarithm
log_samples = np.log(samples)
# Produce a scatter matrix for each pair of newly-transformed features
pd.plotting.scatter_matrix(log_data, alpha = 0.3, figsize = (14,8), diagonal = 'kde');
sns.heatmap(log_data.corr(), annot=True)
```
### Observation
After applying a natural logarithm scaling to the data, the distribution of each feature should appear much more normal. For any pairs of features you may have identified earlier as being correlated, observe here whether that correlation is still present (and whether it is now stronger or weaker than before).
Run the code below to see how the sample data has changed after having the natural logarithm applied to it.
```
# Display the log-transformed sample data
display(log_samples)
```
### Implementation: Outlier Detection
Detecting outliers in the data is extremely important in the data preprocessing step of any analysis. The presence of outliers can often skew results which take into consideration these data points. There are many "rules of thumb" for what constitutes an outlier in a dataset. Here, we will use [Tukey's Method for identfying outliers](http://datapigtechnologies.com/blog/index.php/highlighting-outliers-in-your-data-with-the-tukey-method/): An *outlier step* is calculated as 1.5 times the interquartile range (IQR). A data point with a feature that is beyond an outlier step outside of the IQR for that feature is considered abnormal.
In the code block below, you will need to implement the following:
- Assign the value of the 25th percentile for the given feature to `Q1`. Use `np.percentile` for this.
- Assign the value of the 75th percentile for the given feature to `Q3`. Again, use `np.percentile`.
- Assign the calculation of an outlier step for the given feature to `step`.
- Optionally remove data points from the dataset by adding indices to the `outliers` list.
**NOTE:** If you choose to remove any outliers, ensure that the sample data does not contain any of these points!
Once you have performed this implementation, the dataset will be stored in the variable `good_data`.
```
# Contains outliers for each feature (with each iteration)
feature_outliers = []
# OPTIONAL: Select the indices for data points you wish to remove
all_outliers = []
# For each feature find the data points with extreme high or low values
for feature in log_data.keys():
# Calculate Q1 (25th percentile of the data) for the given feature
Q1 = np.percentile(log_data[feature], 25.)
# Calculate Q3 (75th percentile of the data) for the given feature
Q3 = np.percentile(log_data[feature], 75.)
# Use the interquartile range to calculate an outlier step
# (1.5 times the interquartile range)
step = 1.5 * (Q3 - Q1)
print ("Outlier step: {0}".format(step))
# Display the outliers
print "Data points considered outliers for the feature '{}':".format(feature)
feature_outliers = log_data[~((log_data[feature] >= Q1 - step) & (log_data[feature] <= Q3 + step))]
display(feature_outliers)
all_outliers += feature_outliers.index.tolist()
print "Number of Outliers (includes duplicates): ", len(all_outliers)
# Find outliers that figure out in 2 or more categories
outliers = pd.Series(all_outliers).value_counts()
outliers = outliers[outliers > 1].index.tolist()
print ("Outliers with 2 or more dublications: {0}".format(outliers))
# Uncomment if you don't want to drop outliers:
#outliers = []
# Remove the outliers, if any were specified
good_data = log_data.drop(log_data.index[outliers]).reset_index(drop = True)
print "New dataset (without Outliers) {} items with {} features each".format(*good_data.shape)
```
### Question 4
*Are there any data points considered outliers for more than one feature based on the definition above? Should these data points be removed from the dataset? If any data points were added to the `outliers` list to be removed, explain why.*
**Answer:**
Outliers for more than one feature:
* **65** - 'Fresh' and 'Frozen'
* **66** - 'Fresh' and 'Delicatessen'
* **75** - 'Grocery' and 'Detergents_Paper'
* **128** - 'Fresh' and 'Delicatessen'
* **154** - 'Milk', 'Delicatessen' and 'Grocery'
It's reasonable to remove outliers with dublication on 2 or more categories from the dataset because they add no valuable information to predictive models, and would only skew the results.
## Feature Transformation
In this section you will use principal component analysis (PCA) to draw conclusions about the underlying structure of the wholesale customer data. Since using PCA on a dataset calculates the dimensions which best maximize variance, we will find which compound combinations of features best describe customers.
### Implementation: PCA
Now that the data has been scaled to a more normal distribution and has had any necessary outliers removed, we can now apply PCA to the `good_data` to discover which dimensions about the data best maximize the variance of features involved. In addition to finding these dimensions, PCA will also report the *explained variance ratio* of each dimension — how much variance within the data is explained by that dimension alone. Note that a component (dimension) from PCA can be considered a new "feature" of the space, however it is a composition of the original features present in the data.
In the code block below, you will need to implement the following:
- Import `sklearn.decomposition.PCA` and assign the results of fitting PCA in six dimensions with `good_data` to `pca`.
- Apply a PCA transformation of `log_samples` using `pca.transform`, and assign the results to `pca_samples`.
```
from sklearn.decomposition import PCA
# Apply PCA by fitting the good data with the same number of dimensions as features
pca = PCA(n_components=6)
pca.fit(good_data)
# Transform log_samples using the PCA fit above
pca_samples = pca.transform(log_samples)
# Generate PCA results plot
pca_results = vs.pca_results(good_data, pca)
```
### Question 5
*How much variance in the data is explained* ***in total*** *by the first and second principal component? What about the first four principal components? Using the visualization provided above, discuss what the first four dimensions best represent in terms of customer spending.*
**Hint:** A positive increase in a specific dimension corresponds with an *increase* of the *positive-weighted* features and a *decrease* of the *negative-weighted* features. The rate of increase or decrease is based on the individual feature weights.
**Answer:**
The first and second principal components accounts for **72.52%** (49.93 + 22.59) of the total variance. The first four principal components account for **92.79%** (49.93 + 22.59 + 10.49 + 9.78) of the total variance.
Each dimension represents different patterns of customer spendings:
* **Dimension 1** - Contains the information represented in 'Grocery', 'Milk' and 'Detergents_Paper' features. These are the most correlated features and could be reduced to a smaller dimension without big loss of information.
* **Dimension 2** - Contains the information represented in 'Fresh', 'Frozen' and 'Delicatessen' features. This makes the 'Dimension 2' component a suitable orthogonal to the 'Dimension 1' component.
* **Dimension 3** - This component capturing the positive impact of 'Frozen' and 'Delicatessen' features on the customer sales and large negative decrease of the 'Fresh' feature. This means that the 'Dimension 3' component tries to account for the lack of correlation between the two features.
* **Dimension 4** - When we transform our data, the values for 'Dimension 4' show us that customers who likely buying more of the positive-weight feature ('Frozen') while buying less of the negative-weight feature ('Delicatessen'). So, if a customer had a high amount of Dimension 4 variance, then we'd know they have an inverse relationship between 'Frozen' and 'Delicatessen'.
### Observation
Run the code below to see how the log-transformed sample data has changed after having a PCA transformation applied to it in six dimensions. Observe the numerical value for the first four dimensions of the sample points. Consider if this is consistent with your initial interpretation of the sample points.
```
# Display sample log-data after having a PCA transformation applied
display(pd.DataFrame(np.round(pca_samples, 4), columns = pca_results.index.values))
```
### Implementation: Dimensionality Reduction
When using principal component analysis, one of the main goals is to reduce the dimensionality of the data — in effect, reducing the complexity of the problem. Dimensionality reduction comes at a cost: Fewer dimensions used implies less of the total variance in the data is being explained. Because of this, the *cumulative explained variance ratio* is extremely important for knowing how many dimensions are necessary for the problem. Additionally, if a signifiant amount of variance is explained by only two or three dimensions, the reduced data can be visualized afterwards.
In the code block below, you will need to implement the following:
- Assign the results of fitting PCA in two dimensions with `good_data` to `pca`.
- Apply a PCA transformation of `good_data` using `pca.transform`, and assign the results to `reduced_data`.
- Apply a PCA transformation of `log_samples` using `pca.transform`, and assign the results to `pca_samples`.
```
# Apply PCA by fitting the good data with only two dimensions
pca = PCA(n_components = 2)
pca.fit(good_data)
# Transform the good data using the PCA fit above
reduced_data = pca.transform(good_data)
# Transform log_samples using the PCA fit above
pca_samples = pca.transform(log_samples)
# Create a DataFrame for the reduced data
reduced_data = pd.DataFrame(reduced_data, columns = ['Dimension 1', 'Dimension 2'])
```
### Observation
Run the code below to see how the log-transformed sample data has changed after having a PCA transformation applied to it using only two dimensions. Observe how the values for the first two dimensions remains unchanged when compared to a PCA transformation in six dimensions.
```
# Display sample log-data after applying PCA transformation in two dimensions
display(pd.DataFrame(np.round(pca_samples, 4), columns = ['Dimension 1', 'Dimension 2']))
```
## Visualizing a Biplot
A biplot is a scatterplot where each data point is represented by its scores along the principal components. The axes are the principal components (in this case `Dimension 1` and `Dimension 2`). In addition, the biplot shows the projection of the original features along the components. A biplot can help us interpret the reduced dimensions of the data, and discover relationships between the principal components and original features.
Run the code cell below to produce a biplot of the reduced-dimension data.
```
# Create a biplot
vs.biplot(good_data, reduced_data, pca)
```
### Observation
Once we have the original feature projections (in red), it is easier to interpret the relative position of each data point in the scatterplot. For instance, a point the lower right corner of the figure will likely correspond to a customer that spends a lot on `'Milk'`, `'Grocery'` and `'Detergents_Paper'`, but not so much on the other product categories.
From the biplot, which of the original features are most strongly correlated with the first component? What about those that are associated with the second component? Do these observations agree with the pca_results plot you obtained earlier?
**Answer:**
* 'Grocery', 'Milk' and 'Detergents_Paper' features are the most correlated to the Dimension 1.
* 'Frozen' and 'Fresh' features are the most correlated to the Dimension 2.
* 'Delicatessen' feature located diagonally to the other components suggesting independance of the feature.
## Clustering
In this section, you will choose to use either a K-Means clustering algorithm or a Gaussian Mixture Model clustering algorithm to identify the various customer segments hidden in the data. You will then recover specific data points from the clusters to understand their significance by transforming them back into their original dimension and scale.
### Question 6
*What are the advantages to using a K-Means clustering algorithm? What are the advantages to using a Gaussian Mixture Model clustering algorithm? Given your observations about the wholesale customer data so far, which of the two algorithms will you use and why?*
**Answer:**
**K-Means Clustering:**
It is an algorithm, which classifies samples based on attributes/features into K number of clusters. Clustering or grouping of samples is done by minimizing the distance between sample and the centroid. i.e. Assign the centroid and optimize the centroid based on the distances from the points to it. This is called as Hard Assignment i.e. We are certain that particular points belong to particular centroid and then based on the least squares distance method, we will optimize the place of the centroid.
Advantages of K-Means:
1. Running Time
2. Better for high dimensional data.
3. Easy to interpret and Implement.
Disadvantages of K-Means:
1. Assumes the clusters as spherical, so does not work efficiently with complex geometrical shaped data(Mostly Non-Linear)
2. Hard Assignment might lead to mis grouping.
**Guassian Mixture:**
Instead of Hard assgning data points to a cluster, if we are uncertain about the data points where they belong or to which group, we use this method. It uses probability of a sample to determine the feasibility of it belonging to a cluster.
Advantages:
1. Does not assume clusters to be of any geometry. Works well with non-linear geometric distributions as well.
2. Does not bias the cluster sizes to have specific structures as does by K-Means (Circular).
Disadvantages:
1. Uses all the components it has access to, so initialization of clusters will be difficult when dimensionality of data is high.
2. Difficult to interpret.
Given scatter plot, a lot of data points are not clearly belong to one particular cluster or another, so it's rational to adopt a Gaussian Mixture Model (GMM) for this task.
Source: [What is the difference between K-means and the mixture model of Gaussian?](https://www.quora.com/What-is-the-difference-between-K-means-and-the-mixture-model-of-Gaussian)
### Implementation: Creating Clusters
Depending on the problem, the number of clusters that you expect to be in the data may already be known. When the number of clusters is not known *a priori*, there is no guarantee that a given number of clusters best segments the data, since it is unclear what structure exists in the data — if any. However, we can quantify the "goodness" of a clustering by calculating each data point's *silhouette coefficient*. The [silhouette coefficient](http://scikit-learn.org/stable/modules/generated/sklearn.metrics.silhouette_score.html) for a data point measures how similar it is to its assigned cluster from -1 (dissimilar) to 1 (similar). Calculating the *mean* silhouette coefficient provides for a simple scoring method of a given clustering.
In the code block below, you will need to implement the following:
- Fit a clustering algorithm to the `reduced_data` and assign it to `clusterer`.
- Predict the cluster for each data point in `reduced_data` using `clusterer.predict` and assign them to `preds`.
- Find the cluster centers using the algorithm's respective attribute and assign them to `centers`.
- Predict the cluster for each sample data point in `pca_samples` and assign them `sample_preds`.
- Import `sklearn.metrics.silhouette_score` and calculate the silhouette score of `reduced_data` against `preds`.
- Assign the silhouette score to `score` and print the result.
```
from sklearn.mixture import GaussianMixture
from sklearn.metrics import silhouette_score
cl_sizes = [2, 3, 4, 5, 6, 8, 10, 12, 14, 17, 20, 23, 26, 30, 35, 40, 50]
best_size = 0
score = 0
for size in cl_sizes:
# Apply your clustering algorithm of choice to the reduced data
clusterer = GaussianMixture(n_components=size)
clusterer.fit(reduced_data)
# Predict the cluster for each data point
cl_preds = clusterer.predict(reduced_data)
# Find the cluster centers (mean parameters for each mixture component)
cl_centers = clusterer.means_
# Predict the cluster for each transformed sample data point
cl_sample_preds = clusterer.predict(pca_samples)
# Calculate the mean silhouette coefficient for the number of clusters chosen
cl_score = silhouette_score(reduced_data, cl_preds)
print "{0:2d} clusters: {1:6f} silhouette scores".format(size, cl_score)
if cl_score > score:
score = cl_score
best_size = size
preds = cl_preds
centers = cl_centers
sample_preds = cl_sample_preds
print ("| Best score: {0:4f} | Cluster size: {1} |".format(score, best_size))
```
### Question 7
*Report the silhouette score for several cluster numbers you tried. Of these, which number of clusters has the best silhouette score?*
**Answer:**
The silhouette scores for various sizes of clusters are displayed above.
A Gaussian Mixture Model with **2 clusters** has the best silhouette scores.
### Cluster Visualization
Once you've chosen the optimal number of clusters for your clustering algorithm using the scoring metric above, you can now visualize the results by executing the code block below. Note that, for experimentation purposes, you are welcome to adjust the number of clusters for your clustering algorithm to see various visualizations. The final visualization provided should, however, correspond with the optimal number of clusters.
```
# Display the results of the clustering from implementation
vs.cluster_results(reduced_data, preds, centers, pca_samples)
```
### Implementation: Data Recovery
Each cluster present in the visualization above has a central point. These centers (or means) are not specifically data points from the data, but rather the *averages* of all the data points predicted in the respective clusters. For the problem of creating customer segments, a cluster's center point corresponds to *the average customer of that segment*. Since the data is currently reduced in dimension and scaled by a logarithm, we can recover the representative customer spending from these data points by applying the inverse transformations.
In the code block below, you will need to implement the following:
- Apply the inverse transform to `centers` using `pca.inverse_transform` and assign the new centers to `log_centers`.
- Apply the inverse function of `np.log` to `log_centers` using `np.exp` and assign the true centers to `true_centers`.
```
# Inverse transform the centers
log_centers = pca.inverse_transform(centers)
# Exponentiate the centers
true_centers = np.exp(log_centers)
# Display the true centers
segments = ['Segment {}'.format(i) for i in range(0,len(centers))]
true_centers = pd.DataFrame(np.round(true_centers), columns = data.keys())
true_centers.index = segments
display(true_centers)
print("Statistical description of the dataset (more info in the Data Explotation section)")
display(samples)
print("[0] - Restaurant (sushi-shop), index=64")
print("[1] - Small grossery / Shop in the farm market, index=128")
print("[2] - Supermarket, index=256")
sns.heatmap((true_centers - data.mean()) / data.std(ddof=0),
square=True, annot=True, cbar=False)
```
### Question 8
Consider the total purchase cost of each product category for the representative data points above, and reference the statistical description of the dataset at the beginning of this project. *What set of establishments could each of the customer segments represent?*
**Hint:** A customer who is assigned to `'Cluster X'` should best identify with the establishments represented by the feature set of `'Segment X'`.
**Answer:**
The customer segments mainly influenced by the 'Segment 0' component which is the purchases of 'Milk', 'Grocery', and 'Detergents_Paper'.
Follow the heatmap, 'Fresh', 'Frozen' and 'Delicatessen' features don't seem to have a strong influence on the cluster/segment membership.
* A customer who is assigned to **Cluster/Segment 0** should best identify with the establishments represented by the feature set of 'Supermarket' (index 256). These customers purchase much higher amout of 'Milk', 'Grocery' and 'Detergent_Paper' comparing with customers from Segment 1.
* A customer who is assigned to **Cluster/Segment 1** should best identify with the establishments represented by the feature set of 'Restaurant (sushi-shop)' (index 64). These customers purchase low amout of 'Milk', 'Grocery' and 'Detergent_Paper'.
### Question 9
*For each sample point, which customer segment from* ***Question 8*** *best represents it? Are the predictions for each sample point consistent with this?*
Run the code block below to find which cluster each sample point is predicted to be.
```
# Display the predictions
for i, pred in enumerate(sample_preds):
print "Sample point", i, "predicted to be in Cluster", pred
```
**Answer:**
* Index 64 and 256 belong to cluster 1 (Supermarket). Model's prediction on Index 64 doesn't agree with my original prediction, but index 256 does.
* Index 128 belongs to cluster 0 (Restaurant (sushi-shop)). Model's prediction on Index 128 doesn't agree with my original prediction.
## Conclusion
In this final section, you will investigate ways that you can make use of the clustered data. First, you will consider how the different groups of customers, the ***customer segments***, may be affected differently by a specific delivery scheme. Next, you will consider how giving a label to each customer (which *segment* that customer belongs to) can provide for additional features about the customer data. Finally, you will compare the ***customer segments*** to a hidden variable present in the data, to see whether the clustering identified certain relationships.
### Question 10
Companies will often run [A/B tests](https://en.wikipedia.org/wiki/A/B_testing) when making small changes to their products or services to determine whether making that change will affect its customers positively or negatively. The wholesale distributor is considering changing its delivery service from currently 5 days a week to 3 days a week. However, the distributor will only make this change in delivery service for customers that react positively. *How can the wholesale distributor use the customer segments to determine which customers, if any, would react positively to the change in delivery service?*
**Hint:** Can we assume the change affects all customers equally? How can we determine which group of customers it affects the most?
**Answer:**
The Company could run A/B tests and generalize customers to some clusters/subsets. Each customer's feedback could be evaluated separately. After this Company could generate a decision whether changing the delivery service is critical to each segment and whether customers are happy with the change.
### Question 11
Additional structure is derived from originally unlabeled data when using clustering techniques. Since each customer has a ***customer segment*** it best identifies with (depending on the clustering algorithm applied), we can consider *'customer segment'* as an **engineered feature** for the data. Assume the wholesale distributor recently acquired ten new customers and each provided estimates for anticipated annual spending of each product category. Knowing these estimates, the wholesale distributor wants to classify each new customer to a ***customer segment*** to determine the most appropriate delivery service.
*How can the wholesale distributor label the new customers using only their estimated product spending and the* ***customer segment*** *data?*
**Hint:** A supervised learner could be used to train on the original customers. What would be the target variable?
**Answer:**
There are **2 customer segments** from the dataset, so, it's possible to use a binary classification of whether the new customers belong to class 0 or 1 (class label will be a target variable in this case). For this purpose could be uses algorithm such as SVM, Decision Trees etc.
### Visualizing Underlying Distributions
At the beginning of this project, it was discussed that the `'Channel'` and `'Region'` features would be excluded from the dataset so that the customer product categories were emphasized in the analysis. By reintroducing the `'Channel'` feature to the dataset, an interesting structure emerges when considering the same PCA dimensionality reduction applied earlier to the original dataset.
Run the code block below to see how each data point is labeled either `'HoReCa'` (Hotel/Restaurant/Cafe) or `'Retail'` the reduced space. In addition, you will find the sample points are circled in the plot, which will identify their labeling.
```
# Display the clustering results based on 'Channel' data
vs.channel_results(reduced_data, outliers, pca_samples)
```
### Question 12
*How well does the clustering algorithm and number of clusters you've chosen compare to this underlying distribution of Hotel/Restaurant/Cafe customers to Retailer customers? Are there customer segments that would be classified as purely 'Retailers' or 'Hotels/Restaurants/Cafes' by this distribution? Would you consider these classifications as consistent with your previous definition of the customer segments?*
**Answer:**
The number of clusters matches the underlying distribution to a very good extent (Split on Dimension 1 lies approximately on 0).
There are a lot of overlaps between 'Hotels' and 'Retail customers' around the point 0,0 (two dimensional space).
The classification of the customer segments generated by the clusters seems accurate with the actual customer segments of the dataset.
| github_jupyter |
```
# PyTorch
import torch
from torch import nn, optim
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, datasets
import torch.nn.functional as F
from torch.optim.lr_scheduler import ReduceLROnPlateau
# PyTorch Lightning
import pytorch_lightning as pl
from pytorch_lightning import Trainer
from pytorch_lightning.loggers import WandbLogger
from pytorch_lightning.callbacks import EarlyStopping, LearningRateMonitor
# ETC
import wandb
import numpy as np
import matplotlib.pyplot as plt
import os
import os.path as osp
import random
import torchmetrics
from PIL import Image
AVAIL_GPUS = min(1, torch.cuda.device_count())
BATCH_SIZE = 256 if AVAIL_GPUS else 64
pl.seed_everything(125)
```
## Load Data (Manually)
```
train_root = './seg_train/'
val_root = './seg_test/'
pred_root = './seg_pred/'
train_root
os.listdir(train_root)
os.listdir(osp.join(train_root, 'buildings'))[0]
train_list = []
train_category = []
for category in os.listdir(train_root):
for file in os.listdir(osp.join(train_root, category)):
train_list.append(osp.join(train_root, category, file))
train_category.append(category)
train_list = np.array(train_list)
train_category = np.array(train_category)
val_list = []
val_category = []
for category in os.listdir(val_root):
for file in os.listdir(osp.join(val_root, category)):
val_list.append(osp.join(val_root, category, file))
val_category.append(category)
val_list = np.array(val_list)
val_category = np.array(val_category)
train_idx = [i for i in range(len(train_list))]
val_idx = [i for i in range(len(val_list))]
random.shuffle(train_idx)
random.shuffle(val_idx)
train_list = train_list[train_idx]
train_category = train_category[train_idx]
val_list = val_list[val_idx]
val_category = val_category[val_idx]
print(len(train_list))
print(len(val_list))
img = Image.open(train_list[0])
plt.imshow(img)
plt.show()
train_category[0]
to_tensor = transforms.ToTensor()
to_tensor(img).shape
```
### One Hot Encoding
```
category = np.unique(train_category)
category
def cat_to_num(cat):
return np.where(category == cat)[0].item()
cat_map = np.vectorize(cat_to_num)
train_target = torch.tensor(cat_map(train_category))
train_target
train_label = F.one_hot(train_target)
train_label
def to_onehot(cat_list):
return F.one_hot(torch.tensor(cat_map(cat_list)))
class IntelImage(Dataset):
def __init__(self, X_url, y_list):
self.X_list = to_tensor(X_list)
self.y_list = to_onehot(y_list)
def __len__(self):
return len(self.X_list)
def __getitem__(self, idx):
return self.X_list[idx], self.y_list[idx]
```
## Load Data (With ImageFolder)
```
df_train = datasets.ImageFolder(root=train_root, transform=transforms.ToTensor())
df_train
dl_train = DataLoader(df_train, shuffle=True)
it = iter(dl_train)
X, y = next(it)
X.shape
y
plt.imshow(X[0].permute([1,2,0]))
plt.show()
print(y)
```
### Calc mean & std
```
# mean = torch.zeros((3,))
# std = torch.zeros((3,))
# for X, _ in dl_train:
# for i in range(3):
# mean[i] += X[:,i,:,:].mean()
# std[i] += X[:,i,:,:].std()
# mean.div_(len(dl_train))
# std.div_(len(dl_train))
mean = torch.tensor([0.4302, 0.4575, 0.4538])
std = torch.tensor([0.2355, 0.2345, 0.2429])
```
## Load Data (With normalization)
```
class ImageTransform():
def __init__(self, mean, std):
self.trsf = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize(mean, std)
])
def __call__(self, img):
return self.trsf(img)
df_train = datasets.ImageFolder(root=train_root, transform=ImageTransform(mean, std))
df_val = datasets.ImageFolder(root=val_root, transform=ImageTransform(mean, std))
df_train
plt.imshow(df_train[0][0].permute([1,2,0]))
plt.show()
for i, (X, y) in enumerate(df_train):
if X.shape != torch.Size([3, 150, 150]):
print(i)
print(X.shape)
plt.imshow(df_train[1637][0].permute([1,2,0]))
plt.show()
class NewImageTransform():
def __init__(self, mean, std):
self.trsf = transforms.Compose([
transforms.Resize((150, 150)),
transforms.ToTensor(),
transforms.Normalize(mean, std)
])
def __call__(self, img):
return self.trsf(img)
```
### Test
```
X_temp = df_train[0][0]
print(X_temp.shape)
X_temp = nn.Conv2d(3, 16, 3, padding=1)(X_temp)
print(X_temp.shape)
X_temp = nn.MaxPool2d(kernel_size=2, stride=2)(X_temp)
print(X_temp.shape)
X_temp = nn.Conv2d(16, 32, 3, padding=1)(X_temp)
print(X_temp.shape)
X_temp = nn.MaxPool2d(kernel_size=2, stride=2)(X_temp)
print(X_temp.shape)
X_temp = nn.Conv2d(32, 64, 3, padding=1)(X_temp)
print(X_temp.shape)
X_temp = nn.MaxPool2d(kernel_size=2, stride=2)(X_temp)
print(X_temp.shape)
X_temp = nn.Conv2d(64, 128, 3, padding=1)(X_temp)
print(X_temp.shape)
X_temp = nn.MaxPool2d(kernel_size=2, stride=2)(X_temp)
print(X_temp.shape)
X_temp = nn.Conv2d(128, 256, 3, padding=1)(X_temp)
print(X_temp.shape)
X_temp = nn.MaxPool2d(kernel_size=2, stride=2)(X_temp)
print(X_temp.shape)
```
## Main - PyTorch Lightning (Basic)
```
class IIC_Basic(pl.LightningModule):
def __init__(self, hparams=None):
super().__init__()
layers = []
in_channels = 3
classes = 6
image_size = 150
cfg = [16, 32, 64, 128, 256]
for v in cfg:
conv2d = nn.Conv2d(in_channels, v, kernel_size=3, padding=1)
relu = nn.ReLU(inplace=True)
maxpool = nn.MaxPool2d(kernel_size=2, stride=2)
layers += [conv2d, relu, maxpool]
in_channels = v
image_size = image_size // 2
layers += [nn.Flatten()]
in_features = cfg[-1] * image_size ** 2
fc_cfg = [256]
for v in fc_cfg:
fc = nn.Linear(in_features, v)
bn = nn.BatchNorm1d(v)
relu = nn.ReLU(inplace=True)
drop = nn.Dropout(0.1)
layers += [fc, bn, relu, drop]
in_features = v
layers += [nn.Linear(in_features, 6)]
self.net = nn.Sequential(*layers)
self.learning_rate = hparams['learning_rate']
self.batch_size = hparams['batch_size']
self.epochs = hparams['epochs']
self.accuracy = torchmetrics.classification.Accuracy()
self.save_hyperparameters(hparams)
def forward(self, x):
return self.net(x)
def training_step(self, batch, batch_idx):
X, y = batch
y_hat = self(X)
loss = F.cross_entropy(y_hat, y)
acc = self.accuracy(y_hat, y)
self.log('train_loss', loss)
self.log('train_acc', acc)
return loss
def validation_step(self, batch, batch_idx):
X, y = batch
y_hat = self(X)
loss = F.cross_entropy(y_hat, y)
acc = self.accuracy(y_hat, y)
self.log('val_loss', loss)
self.log('val_acc', acc)
return loss
def configure_optimizers(self):
optimizer = optim.Adam(self.parameters(), lr=self.learning_rate)
return {
"optimizer": optimizer,
"lr_scheduler": {
"scheduler": ReduceLROnPlateau(
optimizer,
mode="min",
patience=3,
min_lr=1e-6
),
"interval": "epoch",
"monitor": "val_loss",
"strict": True,
}
}
def prepare_data(self):
self.ds_train = datasets.ImageFolder(
root=train_root,
transform=NewImageTransform(mean, std)
)
self.ds_val = datasets.ImageFolder(
root=val_root,
transform=NewImageTransform(mean, std)
)
def train_dataloader(self):
return DataLoader(self.ds_train, batch_size=self.batch_size, shuffle=True)
def val_dataloader(self):
return DataLoader(self.ds_val, batch_size=self.batch_size)
hparams = {
"learning_rate": 1e-3,
"batch_size": BATCH_SIZE,
"epochs": 30
}
model = IIC_Basic(hparams=hparams)
model
wandb_logger = WandbLogger(
project='IIC'
)
trainer = Trainer(
logger=wandb_logger,
max_epochs=hparams["epochs"],
gpus=AVAIL_GPUS,
enable_progress_bar=False,
callbacks=[
LearningRateMonitor(logging_interval="epoch")
]
)
trainer.fit(model)
wandb.finish()
```
| github_jupyter |
```
import os
import numpy as np
import pandas as pd
import cv2
import math
import matplotlib.pyplot as plt
import torch
from PIL import Image
import torchvision.transforms as T
train_data= pd.read_csv("../ELEC576project/sartorius-cell-instance-segmentation/train.csv")
def rotate_image(image, angle):
# Get the image size
image_size = (image.shape[1], image.shape[0])
image_center = tuple(np.array(image_size) / 2)
# Convert the OpenCV 3x2 rotation matrix to 3x3
rot_mat = np.vstack(
[cv2.getRotationMatrix2D(image_center, angle, 1.0), [0, 0, 1]]
)
rot_mat_notranslate = np.matrix(rot_mat[0:2, 0:2])
# Shorthand for below calcs
image_w2 = image_size[0] * 0.5
image_h2 = image_size[1] * 0.5
# Obtain the rotated coordinates of the image corners
rotated_coords = [
(np.array([-image_w2, image_h2]) * rot_mat_notranslate).A[0],
(np.array([ image_w2, image_h2]) * rot_mat_notranslate).A[0],
(np.array([-image_w2, -image_h2]) * rot_mat_notranslate).A[0],
(np.array([ image_w2, -image_h2]) * rot_mat_notranslate).A[0]
]
# Find the size of the new image
x_coords = [pt[0] for pt in rotated_coords]
x_pos = [x for x in x_coords if x > 0]
x_neg = [x for x in x_coords if x < 0]
y_coords = [pt[1] for pt in rotated_coords]
y_pos = [y for y in y_coords if y > 0]
y_neg = [y for y in y_coords if y < 0]
right_bound = max(x_pos)
left_bound = min(x_neg)
top_bound = max(y_pos)
bot_bound = min(y_neg)
new_w = int(abs(right_bound - left_bound))
new_h = int(abs(top_bound - bot_bound))
# We require a translation matrix to keep the image centred
trans_mat = np.matrix([
[1, 0, int(new_w * 0.5 - image_w2)],
[0, 1, int(new_h * 0.5 - image_h2)],
[0, 0, 1]
])
# Compute the tranform for the combined rotation and translation
affine_mat = (np.matrix(trans_mat) * np.matrix(rot_mat))[0:2, :]
# Apply the transform
result = cv2.warpAffine(
image,
affine_mat,
(new_w, new_h),
flags=cv2.INTER_LINEAR
)
return result
def largest_rotated_rect(w, h, angle):
quadrant = int(math.floor(angle / (math.pi / 2))) & 3
sign_alpha = angle if ((quadrant & 1) == 0) else math.pi - angle
alpha = (sign_alpha % math.pi + math.pi) % math.pi
bb_w = w * math.cos(alpha) + h * math.sin(alpha)
bb_h = w * math.sin(alpha) + h * math.cos(alpha)
gamma = math.atan2(bb_w, bb_w) if (w < h) else math.atan2(bb_w, bb_w)
delta = math.pi - alpha - gamma
length = h if (w < h) else w
d = length * math.cos(alpha)
a = d * math.sin(alpha) / math.sin(delta)
y = a * math.cos(gamma)
x = y * math.tan(gamma)
return (
bb_w - 2 * x,
bb_h - 2 * y
)
def crop_around_center(image, width, height):
image_size = (image.shape[1], image.shape[0])
image_center = (int(image_size[0] * 0.5), int(image_size[1] * 0.5))
if(width > image_size[0]):
width = image_size[0]
if(height > image_size[1]):
height = image_size[1]
x1 = int(image_center[0] - width * 0.5)
x2 = int(image_center[0] + width * 0.5)
y1 = int(image_center[1] - height * 0.5)
y2 = int(image_center[1] + height * 0.5)
return image[y1:y2, x1:x2]
def plot_image(tensor):
plt.figure()
plt.imshow(tensor.numpy().transpose(1, 2, 0))
plt.show()
id = "0ba181d412da"
sample_image_df = train_data[train_data['id'] == id]
sample_path = f"../ELEC576project/sartorius-cell-instance-segmentation/train/{sample_image_df['id'].iloc[0]}/{sample_image_df['id'].iloc[0]}.png"
image = cv2.imread(sample_path,cv2.IMREAD_COLOR)
image_height= image.shape[0]
image_width = image.shape[1]
# for i in np.arange(0, 360, 1):
# image_orig = np.copy(image)
# image_rotated = rotate_image(image, i)
# image_rotated_cropped = crop_around_center(
# image_rotated,
# *largest_rotated_rect(
# 256,
# 256,
# math.radians(i)
# )
# )
# img2 = np.zeros(image.shape, np.uint8)
# dst = np.zeros((image_height*2, image_width,3),np.uint8)
# for i in range (image_height):
# for j in range (image_width):
# img2[i,j]=image[image_height-1-i,j]
# for i in range (image_height):
# for j in range (image_width):
# dst[i,j]=image[i,j]
# dst[i+image_height,j]=img2[i,j]
# plt.figure(figsize=(16, 32))
# plt.subplot(3, 1, 1)
# plt.imshow(image_orig)
# plt.axis("off")
# plt.subplot(3, 1, 2)
# plt.imshow(image_rotated)
# plt.axis("off")
# plt.subplot(3, 1, 3)
# plt.imshow(image_rotated_cropped)
# plt.axis("off")
# srcScalar = np.float32([[0.5,0,0],[0,0.5,0]])
# image = cv2.warpAffine(image, srcScalar, (int(image_width*0.5), int(image_height*0.5)))
# image = cv2.imread(sample_path,1)
# image_height= image.shape[0]
# image_width = image.shape[1]
# cv2.line(dst,(0,image_height),(image_width,image_height),(0,0,255))
# cv2.imshow('img', dst)
# cv2.waitKey(0)
# torchvision image reflect
torch.manual_seed(0)
image2 = T.ToPILImage()(image)
image2 = T.Pad(padding=(2,32), padding_mode='reflect')(image2)
image2 = T.ToTensor()(image2)
plot_image(image2)
#revert to original size
image3 = T.ToPILImage()(image2)
image3 = T.RandomCrop(size=(520,704))(image3)
image3 = T.ToTensor()(image3)
plot_image(image3)
def encode_mask_to_rle(mask):
'''
mask: numpy array binary mask
1 - mask
0 - background
Returns encoded run length
'''
pixels = mask.flatten()
pixels = np.concatenate([[0], pixels, [0]])
runs = np.where(pixels[1:] != pixels[:-1])[0] + 1
runs[1::2] -= runs[::2]
return ' '.join(str(x) for x in runs)
example = np.array([1,1,1,1,0,0,0,1,0,1,1,0,0,0,1,1,1])
rle_encoded = encode_mask_to_rle(example)
print(rle_encoded)
```
| github_jupyter |
# Rigid-body transformations in three-dimensions
> Marcos Duarte
> Laboratory of Biomechanics and Motor Control ([http://demotu.org/](http://demotu.org/))
> Federal University of ABC, Brazil
The kinematics of a rigid body is completely described by its pose, i.e., its position and orientation in space (and the corresponding changes, translation and rotation). In a three-dimensional space, at least three coordinates and three angles are necessary to describe the pose of the rigid body, totalizing six degrees of freedom for a rigid body.
In motion analysis, to describe a translation and rotation of a rigid body with respect to a coordinate system, typically we attach another coordinate system to the rigid body and determine a transformation between these two coordinate systems.
A transformation is any function mapping a set to another set. For the description of the kinematics of rigid bodies, we are interested only in what is called rigid or Euclidean transformations (denoted as SE(3) for the three-dimensional space) because they preserve the distance between every pair of points of the body (which is considered rigid by definition). Translations and rotations are examples of rigid transformations (a reflection is also an example of rigid transformation but this changes the right-hand axis convention to a left hand, which usually is not of interest). In turn, rigid transformations are examples of [affine transformations](https://en.wikipedia.org/wiki/Affine_transformation). Examples of other affine transformations are shear and scaling transformations (which preserves angles but not lengths).
We will follow the same rationale as in the notebook [Rigid-body transformations in a plane (2D)](http://nbviewer.ipython.org/github/demotu/BMC/blob/master/notebooks/Transformation2D.ipynb) and we will skip the fundamental concepts already covered there. So, you if haven't done yet, you should read that notebook before continuing here.
## Translation
A pure three-dimensional translation of a rigid body (or a coordinate system attached to it) in relation to other rigid body (with other coordinate system) is illustrated in the figure below.
<br>
<figure><img src='./../images/translation3D.png' alt='translation 3D'/> <figcaption><center><i>Figure. A point in three-dimensional space represented in two coordinate systems, with one coordinate system translated.</i></center></figcaption> </figure>
The position of point $\mathbf{P}$ originally described in the $xyz$ (local) coordinate system but now described in the $\mathbf{XYZ}$ (Global) coordinate system in vector form is:
$$ \mathbf{P_G} = \mathbf{L_G} + \mathbf{P_l} $$
Or in terms of its components:
$$ \begin{array}{}
\mathbf{P_X} =& \mathbf{L_X} + \mathbf{P}_x \\
\mathbf{P_Y} =& \mathbf{L_Y} + \mathbf{P}_y \\
\mathbf{P_Z} =& \mathbf{L_Z} + \mathbf{P}_z
\end{array} $$
And in matrix form:
$$
\begin{bmatrix}
\mathbf{P_X} \\
\mathbf{P_Y} \\
\mathbf{P_Z}
\end{bmatrix} =
\begin{bmatrix}
\mathbf{L_X} \\
\mathbf{L_Y} \\
\mathbf{L_Z}
\end{bmatrix} +
\begin{bmatrix}
\mathbf{P}_x \\
\mathbf{P}_y \\
\mathbf{P}_z
\end{bmatrix}
$$
From classical mechanics, this is an example of [Galilean transformation](http://en.wikipedia.org/wiki/Galilean_transformation).
Let's use Python to compute some numeric examples:
```
# Import the necessary libraries
import numpy as np
# suppress scientific notation for small numbers:
np.set_printoptions(precision=4, suppress=True)
```
For example, if the local coordinate system is translated by $\mathbf{L_G}=[1, 2, 3]$ in relation to the Global coordinate system, a point with coordinates $\mathbf{P_l}=[4, 5, 6]$ at the local coordinate system will have the position $\mathbf{P_G}=[5, 7, 9]$ at the Global coordinate system:
```
LG = np.array([1, 2, 3]) # Numpy array
Pl = np.array([4, 5, 6])
PG = LG + Pl
PG
```
This operation also works if we have more than one point (NumPy try to guess how to handle vectors with different dimensions):
```
Pl = np.array([[1, 2, 3], [4, 5, 6], [7, 8, 9]]) # 2D array with 3 rows and 2 columns
PG = LG + Pl
PG
```
## Rotation
A pure three-dimensional rotation of a $xyz$ (local) coordinate system in relation to other $\mathbf{XYZ}$ (Global) coordinate system and the position of a point in these two coordinate systems are illustrated in the next figure (remember that this is equivalent to describing a rotation between two rigid bodies).
<br>
<figure><img src='./../images/rotation3D.png' alt='rotation 3D'/> <figcaption><center><i>A point in three-dimensional space represented in two coordinate systems, with one system rotated.</i></center></figcaption> </figure>
In analogy to the rotation in two dimensions, we can calculate the rotation matrix that describes the rotation of the $xyz$ (local) coordinate system in relation to the $\mathbf{XYZ}$ (Global) coordinate system using the direction cosines between the axes of the two coordinate systems:
$$ \mathbf{R_{Gl}} = \begin{bmatrix}
\cos\mathbf{X}x & \cos\mathbf{X}y & \cos\mathbf{X}z \\
\cos\mathbf{Y}x & \cos\mathbf{Y}y & \cos\mathbf{Y}z \\
\cos\mathbf{Z}x & \cos\mathbf{Z}y & \cos\mathbf{Z}z
\end{bmatrix} $$
Note however that for rotations around more than one axis, these angles will not lie in the main planes ($\mathbf{XY, YZ, ZX}$) of the $\mathbf{XYZ}$ coordinate system, as illustrated in the figure below for the direction angles of the $y$ axis only. Thus, the determination of these angles by simple inspection, as we have done for the two-dimensional case, would not be simple.
<br>
<figure>
<img src='./../images/directioncosine3D.png' width=260 alt='direction angles 3D'/> <figcaption><center><i>Figure. Definition of direction angles for the $y$ axis of the local coordinate system in relation to the $\mathbf{XYZ}$ Global coordinate system.</i></center></figcaption>
</figure>
Note that the nine angles shown in the matrix above for the direction cosines are obviously redundant since only three angles are necessary to describe the orientation of a rigid body in the three-dimensional space.
An important characteristic of angles in the three-dimensional space is that angles cannot be treated as vectors: the result of a sequence of rotations of a rigid body around different axes depends on the order of the rotations, as illustrated in the next figure.
<br>
<figure>
<img src='./../images/rotationsseqs2.png' alt='rotations'/><figcaption><i>Figure. The result of a sequence of rotations around different axes of a coordinate system depends on the order of the rotations. In the first example (first row), the rotations are around a Global (fixed) coordinate system. In the second example (second row), the rotations are around a local (rotating) coordinate system.</i></figcaption>
</figure>
Let's focus now on how to understand rotations in the three-dimensional space, looking at the rotations between coordinate systems (or between rigid bodies). Later we will apply what we have learned to describe the position of a point in these different coordinate systems.
### Euler angles
There are different ways to describe a three-dimensional rotation of a rigid body (or of a coordinate system). The most straightforward solution would probably be to use a [spherical coordinate system](http://nbviewer.ipython.org/github/demotu/BMC/blob/master/notebooks/ReferenceFrame.ipynb#Spherical-coordinate-system), but spherical coordinates would be difficult to give an anatomical or clinical interpretation. A solution that has been often employed in biomechanics to handle rotations in the three-dimensional space is to use Euler angles. Under certain conditions, Euler angles can have an anatomical interpretation, but this representation also has some caveats. Let's see the Euler angles now.
[Leonhard Euler](https://en.wikipedia.org/wiki/Leonhard_Euler) in the XVIII century showed that two three-dimensional coordinate systems with a common origin can be related by a sequence of up to three elemental rotations about the axes of the local coordinate system, where no two successive rotations may be about the same axis, which now are known as [Euler (or Eulerian) angles](http://en.wikipedia.org/wiki/Euler_angles).
#### Elemental rotations
First, let's see rotations around a fixed Global coordinate system as we did for the two-dimensional case. The next figure illustrates elemental rotations of the local coordinate system around each axis of the fixed Global coordinate system.
<br>
<figure>
<img src='./../images/rotations.png' alt='rotations'/> <figcaption><center><i>Figure. Elemental rotations of the $xyz$ coordinate system around each axis, $\mathbf{X}$, $\mathbf{Y}$, and $\mathbf{Z}$, of the fixed $\mathbf{XYZ}$ coordinate system. Note that for better clarity, the axis around where the rotation occurs is shown perpendicular to this page for each elemental rotation.</i></center></figcaption>
</figure>
#### Rotations around the fixed coordinate system
The rotation matrices for the elemental rotations around each axis of the fixed $\mathbf{XYZ}$ coordinate system (rotations of the local coordinate system in relation to the Global coordinate system) are shown next.
Around $\mathbf{X}$ axis:
$$ \mathbf{R_{Gl,\,X}} =
\begin{bmatrix}
1 & 0 & 0 \\
0 & \cos\alpha & -\sin\alpha \\
0 & \sin\alpha & \cos\alpha
\end{bmatrix} $$
Around $\mathbf{Y}$ axis:
$$ \mathbf{R_{Gl,\,Y}} =
\begin{bmatrix}
\cos\beta & 0 & \sin\beta \\
0 & 1 & 0 \\
-\sin\beta & 0 & \cos\beta
\end{bmatrix} $$
Around $\mathbf{Z}$ axis:
$$ \mathbf{R_{Gl,\,Z}} =
\begin{bmatrix}
\cos\gamma & -\sin\gamma & 0\\
\sin\gamma & \cos\gamma & 0 \\
0 & 0 & 1
\end{bmatrix} $$
These matrices are the rotation matrices for the case of two-dimensional coordinate systems plus the corresponding terms for the third axes of the local and Global coordinate systems, which are parallel.
To understand why the terms for the third axes are 1's or 0's, for instance, remember they represent the cosine directors. The cosines between $\mathbf{X}x$, $\mathbf{Y}y$, and $\mathbf{Z}z$ for the elemental rotations around respectively the $\mathbf{X}$, $\mathbf{Y}$, and $\mathbf{Z}$ axes are all 1 because $\mathbf{X}x$, $\mathbf{Y}y$, and $\mathbf{Z}z$ are parallel ($\cos 0^o$). The cosines of the other elements are zero because the axis around where each rotation occurs is perpendicular to the other axes of the coordinate systems ($\cos 90^o$).
#### Rotations around the local coordinate system
The rotation matrices for the elemental rotations this time around each axis of the $xyz$ coordinate system (rotations of the Global coordinate system in relation to the local coordinate system), similarly to the two-dimensional case, are simply the transpose of the above matrices as shown next.
Around $x$ axis:
$$ \mathbf{R}_{\mathbf{lG},\,x} =
\begin{bmatrix}
1 & 0 & 0 \\
0 & \cos\alpha & \sin\alpha \\
0 & -\sin\alpha & \cos\alpha
\end{bmatrix} $$
Around $y$ axis:
$$ \mathbf{R}_{\mathbf{lG},\,y} =
\begin{bmatrix}
\cos\beta & 0 & -\sin\beta \\
0 & 1 & 0 \\
\sin\beta & 0 & \cos\beta
\end{bmatrix} $$
Around $z$ axis:
$$ \mathbf{R}_{\mathbf{lG},\,z} =
\begin{bmatrix}
\cos\gamma & \sin\gamma & 0\\
-\sin\gamma & \cos\gamma & 0 \\
0 & 0 & 1
\end{bmatrix} $$
Notice this is equivalent to instead of rotating the local coordinate system by $\alpha, \beta, \gamma$ in relation to axes of the Global coordinate system, to rotate the Global coordinate system by $-\alpha, -\beta, -\gamma$ in relation to the axes of the local coordinate system; remember that $\cos(-\:\cdot)=\cos(\cdot)$ and $\sin(-\:\cdot)=-\sin(\cdot)$.
The fact that we chose to rotate the local coordinate system by a counterclockwise (positive) angle in relation to the Global coordinate system is just a matter of convention.
#### Sequence of elemental rotations
Consider now a sequence of elemental rotations around the $\mathbf{X}$, $\mathbf{Y}$, and $\mathbf{Z}$ axes of the fixed $\mathbf{XYZ}$ coordinate system illustrated in the next figure.
<br>
<figure><img src='./../images/rotations_XYZ.png' alt='rotations'/> <figcaption><center><i>Figure. Sequence of elemental rotations of the $xyz$ coordinate system around each axis, $\mathbf{X}$, $\mathbf{Y}$, $\mathbf{Z}$, of the fixed $\mathbf{XYZ}$ coordinate system.</i></center></figcaption> </figure>
This sequence of elemental rotations (each one of the local coordinate system with respect to the fixed Global coordinate system) is mathematically represented by a multiplication between the rotation matrices:
$$ \begin{array}{l l}
\mathbf{R_{Gl,\;XYZ}} & = \mathbf{R_{Z}} \mathbf{R_{Y}} \mathbf{R_{X}} \\
\\
& = \begin{bmatrix}
\cos\gamma & -\sin\gamma & 0\\
\sin\gamma & \cos\gamma & 0 \\
0 & 0 & 1
\end{bmatrix}
\begin{bmatrix}
\cos\beta & 0 & \sin\beta \\
0 & 1 & 0 \\
-\sin\beta & 0 & \cos\beta
\end{bmatrix}
\begin{bmatrix}
1 & 0 & 0 \\
0 & \cos\alpha & -sin\alpha \\
0 & \sin\alpha & cos\alpha
\end{bmatrix} \\
\\
& =
\begin{bmatrix}
\cos\beta\:\cos\gamma \;&\;
\sin\alpha\:\sin\beta\:cos\gamma-\cos\alpha\:\sin\gamma \;&\;
\cos\alpha\:\sin\beta\:cos\gamma+\sin\alpha\:\sin\gamma \;\;\; \\
\cos\beta\:\sin\gamma \;&\;
\sin\alpha\:\sin\beta\:sin\gamma+\cos\alpha\:\cos\gamma \;&\;
\cos\alpha\:\sin\beta\:sin\gamma-\sin\alpha\:\cos\gamma \;\;\; \\
-\sin\beta \;&\; \sin\alpha\:\cos\beta \;&\; \cos\alpha\:\cos\beta \;\;\;
\end{bmatrix}
\end{array} $$
Note that the order of the matrices.
We can check this matrix multiplication using [Sympy](http://sympy.org/en/index.html):
```
#import the necessary libraries
from IPython.core.display import Math, display
import sympy as sym
cos, sin = sym.cos, sym.sin
a, b, g = sym.symbols('alpha, beta, gamma')
# Elemental rotation matrices of xyz in relation to XYZ:
RX = sym.Matrix([[1, 0, 0], [0, cos(a), -sin(a)], [0, sin(a), cos(a)]])
RY = sym.Matrix([[cos(b), 0, sin(b)], [0, 1, 0], [-sin(b), 0, cos(b)]])
RZ = sym.Matrix([[cos(g), -sin(g), 0], [sin(g), cos(g), 0], [0, 0, 1]])
# Rotation matrix of xyz in relation to XYZ:
RXYZ = RZ*RY*RX
display(Math(sym.latex(r'\mathbf{R_{Gl,\,XYZ}}=') + sym.latex(RXYZ, mat_str='matrix')))
```
For instance, we can calculate the numerical rotation matrix for these sequential elemental rotations by $90^o$ around $\mathbf{X,Y,Z}$:
```
R = sym.lambdify((a, b, g), RXYZ, 'numpy')
R = R(np.pi/2, np.pi/2, np.pi/2)
display(Math(r'\mathbf{R_{Gl,\,XYZ\,}}(90^o, 90^o, 90^o) =' + \
sym.latex(sym.Matrix(R).n(chop=True, prec=3))))
```
Examining the matrix above and the correspondent previous figure, one can see they agree: the rotated $x$ axis (first column of the above matrix) has value -1 in the $\mathbf{Z}$ direction $[0,0,-1]$, the rotated $y$ axis (second column) is at the $\mathbf{Y}$ direction $[0,1,0]$, and the rotated $z$ axis (third column) is at the $\mathbf{X}$ direction $[1,0,0]$.
We also can calculate the sequence of elemental rotations around the $x$, $y$, $z$ axes of the rotating $xyz$ coordinate system illustrated in the next figure.
<br>
<figure>
<img src='./../images/rotations_xyz2.png' alt='rotations'/> <figcaption><center><i>Figure. Sequence of elemental rotations of a second $xyz$ local coordinate system around each axis, $x$, $y$, $z$, of the rotating $xyz$ coordinate system.</i></center></figcaption>
</figure>
Likewise, this sequence of elemental rotations (each one of the local coordinate system with respect to the rotating local coordinate system) is mathematically represented by a multiplication between the rotation matrices (which are the inverse of the matrices for the rotations around $\mathbf{X,Y,Z}$ as we saw earlier):
$$ \begin{array}{l l}
\mathbf{R}_{\mathbf{lG},\,xyz} & = \mathbf{R_{z}} \mathbf{R_{y}} \mathbf{R_{x}} \\
\\
& = \begin{bmatrix}
\cos\gamma & \sin\gamma & 0\\
-\sin\gamma & \cos\gamma & 0 \\
0 & 0 & 1
\end{bmatrix}
\begin{bmatrix}
\cos\beta & 0 & -\sin\beta \\
0 & 1 & 0 \\
sin\beta & 0 & \cos\beta
\end{bmatrix}
\begin{bmatrix}
1 & 0 & 0 \\
0 & \cos\alpha & \sin\alpha \\
0 & -\sin\alpha & \cos\alpha
\end{bmatrix} \\
\\
& =
\begin{bmatrix}
\cos\beta\:\cos\gamma \;&\;
\sin\alpha\:\sin\beta\:\cos\gamma+\cos\alpha\:\sin\gamma \;&\;
\cos\alpha\:\sin\beta\:\cos\gamma-\sin\alpha\:\sin\gamma \;\;\; \\
-\cos\beta\:\sin\gamma \;&\;
-\sin\alpha\:\sin\beta\:\sin\gamma+\cos\alpha\:\cos\gamma \;&\;
\cos\alpha\:\sin\beta\:\sin\gamma+\sin\alpha\:\cos\gamma \;\;\; \\
\sin\beta \;&\; -\sin\alpha\:\cos\beta \;&\; \cos\alpha\:\cos\beta \;\;\;
\end{bmatrix}
\end{array} $$
As before, the order of the matrices is from right to left.
Once again, we can check this matrix multiplication using [Sympy](http://sympy.org/en/index.html):
```
a, b, g = sym.symbols('alpha, beta, gamma')
# Elemental rotation matrices of xyz (local):
Rx = sym.Matrix([[1, 0, 0], [0, cos(a), sin(a)], [0, -sin(a), cos(a)]])
Ry = sym.Matrix([[cos(b), 0, -sin(b)], [0, 1, 0], [sin(b), 0, cos(b)]])
Rz = sym.Matrix([[cos(g), sin(g), 0], [-sin(g), cos(g), 0], [0, 0, 1]])
# Rotation matrix of xyz' in relation to xyz:
Rxyz = Rz*Ry*Rx
Math(sym.latex(r'\mathbf{R}_{\mathbf{lG},\,xyz}=') + sym.latex(Rxyz, mat_str='matrix'))
```
For instance, let's calculate the numerical rotation matrix for these sequential elemental rotations by $90^o$ around $x,y,z$:
```
R = sym.lambdify((a, b, g), Rxyz, 'numpy')
R = R(np.pi/2, np.pi/2, np.pi/2)
display(Math(r'\mathbf{R}_{\mathbf{lG},\,xyz\,}(90^o, 90^o, 90^o) =' + \
sym.latex(sym.Matrix(R).n(chop=True, prec=3))))
```
Once again, let's compare the above matrix and the correspondent previous figure to see if it makes sense. But remember that this matrix is the Global-to-local rotation matrix, $\mathbf{R}_{\mathbf{lG},\,xyz}$, where the coordinates of the local basis' versors are rows, not columns, in this matrix. With this detail in mind, one can see that the previous figure and matrix also agree: the rotated $x$ axis (first row of the above matrix) is at the $\mathbf{Z}$ direction $[0,0,1]$, the rotated $y$ axis (second row) is at the $\mathbf{-Y}$ direction $[0,-1,0]$, and the rotated $z$ axis (third row) is at the $\mathbf{X}$ direction $[1,0,0]$.
In fact, this example didn't serve to distinguish versors as rows or columns because the $\mathbf{R}_{\mathbf{lG},\,xyz}$ matrix above is symmetric!
Let's look on the resultant matrix for the example above after only the first two rotations, $\mathbf{R}_{\mathbf{lG},\,xy}$ to understand this difference:
```
Rxy = Ry*Rx
R = sym.lambdify((a, b), Rxy, 'numpy')
R = R(np.pi/2, np.pi/2)
display(Math(r'\mathbf{R}_{\mathbf{lG},\,xy\,}(90^o, 90^o) =' + \
sym.latex(sym.Matrix(R).n(chop=True, prec=3))))
```
Comparing this matrix with the third plot in the figure, we see that the coordinates of versor $x$ in the Global coordinate system are $[0,1,0]$, i.e., local axis $x$ is aligned with Global axis $Y$, and this versor is indeed the first row, not first column, of the matrix above. Confer the other two rows.
What are then in the columns of the local-to-Global rotation matrix?
The columns are the coordinates of Global basis' versors in the local coordinate system! For example, the first column of the matrix above is the coordinates of $X$, which is aligned with $z$: $[0,0,1]$.
#### Rotations in a coordinate system is equivalent to minus rotations in the other coordinate system
Remember that we saw for the elemental rotations that it's equivalent to instead of rotating the local coordinate system, $xyz$, by $\alpha, \beta, \gamma$ in relation to axes of the Global coordinate system, to rotate the Global coordinate system, $\mathbf{XYZ}$, by $-\alpha, -\beta, -\gamma$ in relation to the axes of the local coordinate system. The same property applies to a sequence of rotations: rotations of $xyz$ in relation to $\mathbf{XYZ}$ by $\alpha, \beta, \gamma$ result in the same matrix as rotations of $\mathbf{XYZ}$ in relation to $xyz$ by $-\alpha, -\beta, -\gamma$:
$$ \begin{array}{l l}
\mathbf{R_{Gl,\,XYZ\,}}(\alpha,\beta,\gamma) & = \mathbf{R_{Gl,\,Z}}(\gamma)\, \mathbf{R_{Gl,\,Y}}(\beta)\, \mathbf{R_{Gl,\,X}}(\alpha) \\
& = \mathbf{R}_{\mathbf{lG},\,z\,}(-\gamma)\, \mathbf{R}_{\mathbf{lG},\,y\,}(-\beta)\, \mathbf{R}_{\mathbf{lG},\,x\,}(-\alpha) \\
& = \mathbf{R}_{\mathbf{lG},\,xyz\,}(-\alpha,-\beta,-\gamma)
\end{array}
$$
Confer that by examining the $\mathbf{R_{Gl,\,XYZ}}$ and $\mathbf{R}_{\mathbf{lG},\,xyz}$ matrices above.
Let's verify this property with Sympy:
```
RXYZ = RZ*RY*RX
# Rotation matrix of xyz in relation to XYZ:
display(Math(sym.latex(r'\mathbf{R_{Gl,\,XYZ\,}}(\alpha,\beta,\gamma) =')))
display(Math(sym.latex(RXYZ, mat_str='matrix')))
# Elemental rotation matrices of XYZ in relation to xyz and negate all angles:
Rx_neg = sym.Matrix([[1, 0, 0], [0, cos(-a), -sin(-a)], [0, sin(-a), cos(-a)]]).T
Ry_neg = sym.Matrix([[cos(-b), 0, sin(-b)], [0, 1, 0], [-sin(-b), 0, cos(-b)]]).T
Rz_neg = sym.Matrix([[cos(-g), -sin(-g), 0], [sin(-g), cos(-g), 0], [0, 0, 1]]).T
# Rotation matrix of XYZ in relation to xyz:
Rxyz_neg = Rz_neg*Ry_neg*Rx_neg
display(Math(sym.latex(r'\mathbf{R}_{\mathbf{lG},\,xyz\,}(-\alpha,-\beta,-\gamma) =')))
display(Math(sym.latex(Rxyz_neg, mat_str='matrix')))
# Check that the two matrices are equal:
display(Math(sym.latex(r'\mathbf{R_{Gl,\,XYZ\,}}(\alpha,\beta,\gamma) \;==\;' + \
r'\mathbf{R}_{\mathbf{lG},\,xyz\,}(-\alpha,-\beta,-\gamma)')))
RXYZ == Rxyz_neg
```
#### Rotations in a coordinate system is the transpose of inverse order of rotations in the other coordinate system
There is another property of the rotation matrices for the different coordinate systems: the rotation matrix, for example from the Global to the local coordinate system for the $xyz$ sequence, is just the transpose of the rotation matrix for the inverse operation (from the local to the Global coordinate system) of the inverse sequence ($\mathbf{ZYX}$) and vice-versa:
$$ \begin{array}{l l}
\mathbf{R}_{\mathbf{lG},\,xyz}(\alpha,\beta,\gamma) & = \mathbf{R}_{\mathbf{lG},\,z\,} \mathbf{R}_{\mathbf{lG},\,y\,} \mathbf{R}_{\mathbf{lG},\,x} \\
& = \mathbf{R_{Gl,\,Z\,}^{-1}} \mathbf{R_{Gl,\,Y\,}^{-1}} \mathbf{R_{Gl,\,X\,}^{-1}} \\
& = \mathbf{R_{Gl,\,Z\,}^{T}} \mathbf{R_{Gl,\,Y\,}^{T}} \mathbf{R_{Gl,\,X\,}^{T}} \\
& = (\mathbf{R_{Gl,\,X\,}} \mathbf{R_{Gl,\,Y\,}} \mathbf{R_{Gl,\,Z}})^\mathbf{T} \\
& = \mathbf{R_{Gl,\,ZYX\,}^{T}}(\gamma,\beta,\alpha)
\end{array}
$$
Where we used the properties that the inverse of the rotation matrix (which is orthonormal) is its transpose and that the transpose of a product of matrices is equal to the product of their transposes in reverse order.
Let's verify this property with Sympy:
```
RZYX = RX*RY*RZ
Rxyz = Rz*Ry*Rx
display(Math(sym.latex(r'\mathbf{R_{Gl,\,ZYX\,}^T}=') + sym.latex(RZYX.T, mat_str='matrix')))
display(Math(sym.latex(r'\mathbf{R}_{\mathbf{lG},\,xyz\,}(\alpha,\beta,\gamma) \,==\,' + \
r'\mathbf{R_{Gl,\,ZYX\,}^T}(\gamma,\beta,\alpha)')))
Rxyz == RZYX.T
```
#### Sequence of rotations of a Vector
We saw in the notebook [Rigid-body transformations in a plane (2D)](http://nbviewer.jupyter.org/github/demotu/BMC/blob/master/notebooks/Transformation2D.ipynb#Rotation-of-a-Vector) that the rotation matrix can also be used to rotate a vector (in fact, a point, image, solid, etc.) by a given angle around an axis of the coordinate system. Let's investigate that for the 3D case using the example earlier where a book was rotated in different orders and around the Global and local coordinate systems.
Before any rotation, the point shown in that figure as a round black dot on the spine of the book has coordinates $\mathbf{P}=[0, 1, 2]$ (the book has thickness 0, width 1, and height 2).
After the first sequence of rotations shown in the figure (rotated around $X$ and $Y$ by $90^0$ each time), $\mathbf{P}$ has coordinates $\mathbf{P}=[1, -2, 0]$ in the global coordinate system. Let's verify that:
```
P = np.array([[0, 1, 2]]).T
RXY = RY*RX
R = sym.lambdify((a, b), RXY, 'numpy')
R = R(np.pi/2, np.pi/2)
P1 = np.dot(R, P)
print('P1 =', P1.T)
```
As expected.
The reader is invited to deduce the position of point $\mathbf{P}$ after the inverse order of rotations, but still around the Global coordinate system.
Although we are performing vector rotation, where we don't need the concept of transformation between coordinate systems, in the example above we used the local-to-Global rotation matrix, $\mathbf{R_{Gl}}$. As we saw in the notebook for the 2D transformation, when we use this matrix, it performs a counter-clockwise (positive) rotation.
If we want to rotate the vector in the clockwise (negative) direction, we can use the very same rotation matrix entering a negative angle or we can use the inverse rotation matrix, the Global-to-local rotation matrix, $\mathbf{R_{lG}}$ and a positive (negative of negative) angle, because $\mathbf{R_{Gl}}(\alpha) = \mathbf{R_{lG}}(-\alpha)$, but bear in mind that even in this latter case we are rotating around the Global coordinate system!
Consider now that we want to deduce algebraically the position of the point $\mathbf{P}$ after the rotations around the local coordinate system as shown in the second set of examples in the figure with the sequence of book rotations. The point has the same initial position, $\mathbf{P}=[0, 1, 2]$, and after the rotations around $x$ and $y$ by $90^0$ each time, what is the position of this point?
It's implicit in this question that the new desired position is in the Global coordinate system because the local coordinate system rotates with the book and the point never changes its position in the local coordinate system. So, by inspection of the figure, the new position of the point is $\mathbf{P1}=[2, 0, 1]$.
Let's naively try to deduce this position by repeating the steps as before:
```
Rxy = Ry*Rx
R = sym.lambdify((a, b), Rxy, 'numpy')
R = R(np.pi/2, np.pi/2)
P1 = np.dot(R, P)
print('P1 =', P1.T)
```
The wrong answer.
The problem is that we defined the rotation of a vector using the local-to-Global rotation matrix. One correction solution for this problem is to continuing using the multiplication of the Global-to-local rotation matrices, $\mathbf{R}_{xy} = \mathbf{R}_y\,\mathbf{R}_x$, transpose $\mathbf{R}_{xy}$ to get the Global-to-local coordinate system, $\mathbf{R_{XY}}=\mathbf{R^T}_{xy}$, and then rotate the vector using this matrix:
```
Rxy = Ry*Rx
RXY = Rxy.T
R = sym.lambdify((a, b), RXY, 'numpy')
R = R(np.pi/2, np.pi/2)
P1 = np.dot(R, P)
print('P1 =', P1.T)
```
The correct answer.
Another solution is to understand that when using the Global-to-local rotation matrix, counter-clockwise rotations (as performed with the book the figure) are negative, not positive, and that when dealing with rotations with the Global-to-local rotation matrix the order of matrix multiplication is inverted, for example, it should be $\mathbf{R\_}_{xyz} = \mathbf{R}_x\,\mathbf{R}_y\,\mathbf{R}_z$ (an added underscore to remind us this is not the convention adopted here).
```
R_xy = Rx*Ry
R = sym.lambdify((a, b), R_xy, 'numpy')
R = R(-np.pi/2, -np.pi/2)
P1 = np.dot(R, P)
print('P1 =', P1.T)
```
The correct answer.
The reader is invited to deduce the position of point $\mathbf{P}$ after the inverse order of rotations, around the local coordinate system.
In fact, you will find elsewhere texts about rotations in 3D adopting this latter convention as the standard, i.e., they introduce the Global-to-local rotation matrix and describe sequence of rotations algebraically as matrix multiplication in the direct order, $\mathbf{R\_}_{xyz} = \mathbf{R}_x\,\mathbf{R}_y\,\mathbf{R}_z$, the inverse we have done in this text. It's all a matter of convention, just that.
#### The 12 different sequences of Euler angles
The Euler angles are defined in terms of rotations around a rotating local coordinate system. As we saw for the sequence of rotations around $x, y, z$, the axes of the local rotated coordinate system are not fixed in space because after the first elemental rotation, the other two axes rotate.
Other sequences of rotations could be produced without combining axes of the two different coordinate systems (Global and local) for the definition of the rotation axes. There is a total of 12 different sequences of three elemental rotations that are valid and may be used for describing the rotation of a coordinate system with respect to another coordinate system:
$$ xyz \quad xzy \quad yzx \quad yxz \quad zxy \quad zyx $$
$$ xyx \quad xzx \quad yzy \quad yxy \quad zxz \quad zyz $$
The first six sequences (first row) are all around different axes, they are usually referred as Cardan or Tait–Bryan angles. The other six sequences (second row) have the first and third rotations around the same axis, but keep in mind that the axis for the third rotation is not at the same place anymore because it changed its orientation after the second rotation. The sequences with repeated axes are known as proper or classic Euler angles.
Which order to use it is a matter of convention, but because the order affects the results, it's fundamental to follow a convention and report it. In Engineering Mechanics (including Biomechanics), the $xyz$ order is more common; in Physics the $zxz$ order is more common (but the letters chosen to refer to the axes are arbitrary, what matters is the directions they represent). In Biomechanics, the order for the Cardan angles is most often based on the angle of most interest or of most reliable measurement. Accordingly, the axis of flexion/extension is typically selected as the first axis, the axis for abduction/adduction is the second, and the axis for internal/external rotation is the last one. We will see about this order later. The $zyx$ order is commonly used to describe the orientation of a ship or aircraft and the rotations are known as the nautical angles: yaw, pitch and roll, respectively (see next figure).
<br>
<figure><img src='https://upload.wikimedia.org/wikipedia/commons/thumb/1/16/Yaw_Axis.svg/319px-Yaw_Axis.svg.png' alt='translation and rotation 3D'/> <figcaption><center><i>Figure. The principal axes of an aircraft and the names for the rotations around these axes (<a href="https://en.wikipedia.org/wiki/Euler_angles">image from Wikipedia</a>).</i></center></figcaption> </figure>
If instead of rotations around the rotating local coordinate system we perform rotations around the fixed Global coordinate system, we will have other 12 different sequences of three elemental rotations, these are called simply rotation angles. So, in total there are 24 possible different sequences of three elemental rotations, but the 24 orders are not independent; with the 12 different sequences of Euler angles at the local coordinate system we can obtain the other 12 sequences at the Global coordinate system.
The Python function `euler_rotmat.py` (code at the end of this text) determines the rotation matrix in algebraic form for any of the 24 different sequences (and sequences with only one or two axes can be inputed). This function also determines the rotation matrix in numeric form if a list of up to three angles are inputed.
For instance, the rotation matrix in algebraic form for the $zxz$ order of Euler angles at the local coordinate system and the correspondent rotation matrix in numeric form after three elemental rotations by $90^o$ each are:
```
import sys
sys.path.insert(1, r'./../functions')
from euler_rotmat import euler_rotmat
Ra, Rn = euler_rotmat(order='zxz', frame='local', angles=[90, 90, 90])
```
#### Line of nodes
The second axis of rotation in the rotating coordinate system is also referred as the nodal axis or line of nodes; this axis coincides with the intersection of two perpendicular planes, one from each Global (fixed) and local (rotating) coordinate systems. The figure below shows an example of rotations and the nodal axis for the $xyz$ sequence of the Cardan angles.
<div class='center-align'><figure><img src='./../images/Node.png' alt='rotations'/> <figcaption><center><i>Figure. First row: example of rotations for the $xyz$ sequence of the Cardan angles. The Global (fixed) $XYZ$ coordinate system is shown in green, the local (rotating) $xyz$ coordinate system is shown in blue. The nodal axis (<b>N</b>, shown in red) is defined by the intersection of the $YZ$ and $xy$ planes and all rotations can be described in relation to this nodal axis or to a perpendicular axis to it. Second row: starting from no rotation, the local coordinate system is rotated by $\alpha$ around the $x$ axis, then by $\beta$ around the rotated $y$ axis, and finally by $\gamma$ around the twice rotated $z$ axis. Note that the line of nodes coincides with the $y$ axis for the second rotation. </i></center></figcaption> </figure></div>
#### Determination of the Euler angles
Once a convention is adopted, the corresponding three Euler angles of rotation can be found.
For example, for the $\mathbf{R}_{xyz}$ rotation matrix:
```
R = euler_rotmat(order='xyz', frame='local')
```
The corresponding Cardan angles for the `xyz` sequence can be given by:
$$ \begin{array}{}
\alpha = \arctan\left(\dfrac{\sin(\alpha)}{\cos(\alpha)}\right) = \arctan\left(\dfrac{-\mathbf{R}_{21}}{\;\;\;\mathbf{R}_{22}}\right) \\
\\
\beta = \arctan\left(\dfrac{\sin(\beta)}{\cos(\beta)}\right) = \arctan\left(\dfrac{\mathbf{R}_{20}}{\sqrt{\mathbf{R}_{00}^2+\mathbf{R}_{10}^2}}\right) \\
\\
\gamma = \arctan\left(\dfrac{\sin(\gamma)}{\cos(\gamma)}\right) = \arctan\left(\dfrac{-\mathbf{R}_{10}}{\;\;\;\mathbf{R}_{00}}\right)
\end{array} $$
Note that we prefer to use the mathematical function `arctan` rather than simply `arcsin` because the latter cannot for example distinguish $45^o$ from $135^o$ and also for better numerical accuracy. See the text [Angular kinematics in a plane (2D)](http://nbviewer.ipython.org/github/demotu/BMC/blob/master/notebooks/AngularKinematics2D.ipynb) for more on these issues.
And here is a Python function to compute the Euler angles of rotations from the Global to the local coordinate system for the $xyz$ Cardan sequence:
```
def euler_angles_from_rot_xyz(rot_matrix, unit='deg'):
""" Compute Euler angles from rotation matrix in the xyz sequence."""
import numpy as np
R = np.array(rot_matrix, copy=False).astype(np.float64)[:3, :3]
angles = np.zeros(3)
angles[0] = np.arctan2(-R[2, 1], R[2, 2])
angles[1] = np.arctan2( R[2, 0], np.sqrt(R[0, 0]**2 + R[1, 0]**2))
angles[2] = np.arctan2(-R[1, 0], R[0, 0])
if unit[:3].lower() == 'deg': # convert from rad to degree
angles = np.rad2deg(angles)
return angles
```
For instance, consider sequential rotations of 45$^o$ around $x,y,z$. The resultant rotation matrix is:
```
Ra, Rn = euler_rotmat(order='xyz', frame='local', angles=[45, 45, 45], showA=False)
```
Let's check that calculating back the Cardan angles from this rotation matrix using the `euler_angles_from_rot_xyz()` function:
```
euler_angles_from_rot_xyz(Rn, unit='deg')
```
We could implement a function to calculate the Euler angles for any of the 12 sequences (in fact, plus another 12 sequences if we consider all the rotations from and to the two coordinate systems), but this is tedious. There is a smarter solution using the concept of [quaternion](http://en.wikipedia.org/wiki/Quaternion), but we wont see that now.
Let's see a problem with using Euler angles known as gimbal lock.
### Gimbal lock
[Gimbal lock](http://en.wikipedia.org/wiki/Gimbal_lock) is the loss of one degree of freedom in a three-dimensional coordinate system that occurs when an axis of rotation is placed parallel with another previous axis of rotation and two of the three rotations will be around the same direction given a certain convention of the Euler angles. This "locks" the system into rotations in a degenerate two-dimensional space. The system is not really locked in the sense it can't be moved or reach the other degree of freedom, but it will need an extra rotation for that.
For instance, let's look at the $zxz$ sequence of rotations by the angles $\alpha, \beta, \gamma$:
$$ \begin{array}{l l}
\mathbf{R}_{zxz} & = \mathbf{R_{z}} \mathbf{R_{x}} \mathbf{R_{z}} \\
\\
& =
\begin{bmatrix}
\cos\gamma & \sin\gamma & 0\\
-\sin\gamma & \cos\gamma & 0 \\
0 & 0 & 1
\end{bmatrix}
\begin{bmatrix}
1 & 0 & 0 \\
0 & \cos\beta & \sin\beta \\
0 & -\sin\beta & \cos\beta
\end{bmatrix}
\begin{bmatrix}
\cos\alpha & \sin\alpha & 0\\
-\sin\alpha & \cos\alpha & 0 \\
0 & 0 & 1
\end{bmatrix}
\end{array} $$
Which results in:
```
a, b, g = sym.symbols('alpha, beta, gamma')
# Elemental rotation matrices of xyz (local):
Rz = sym.Matrix([[cos(a), sin(a), 0], [-sin(a), cos(a), 0], [0, 0, 1]])
Rx = sym.Matrix([[1, 0, 0], [0, cos(b), sin(b)], [0, -sin(b), cos(b)]])
Rz2 = sym.Matrix([[cos(g), sin(g), 0], [-sin(g), cos(g), 0], [0, 0, 1]])
# Rotation matrix for the zxz sequence:
Rzxz = Rz2*Rx*Rz
Math(sym.latex(r'\mathbf{R}_{zxz}=') + sym.latex(Rzxz, mat_str='matrix'))
```
Let's examine what happens with this rotation matrix when the rotation around the second axis ($x$) by $\beta$ is zero:
$$ \begin{array}{l l}
\mathbf{R}_{zxz}(\alpha, \beta=0, \gamma) =
\begin{bmatrix}
\cos\gamma & \sin\gamma & 0\\
-\sin\gamma & \cos\gamma & 0 \\
0 & 0 & 1
\end{bmatrix}
\begin{bmatrix}
1 & 0 & 0 \\
0 & 1 & 0 \\
0 & 0 & 1
\end{bmatrix}
\begin{bmatrix}
\cos\alpha & \sin\alpha & 0\\
-\sin\alpha & \cos\alpha & 0 \\
0 & 0 & 1
\end{bmatrix}
\end{array} $$
The second matrix is the identity matrix and has no effect on the product of the matrices, which will be:
```
Rzxz = Rz2*Rz
Math(sym.latex(r'\mathbf{R}_{xyz}(\alpha, \beta=0, \gamma)=') + \
sym.latex(Rzxz, mat_str='matrix'))
```
Which simplifies to:
```
Rzxz = sym.simplify(Rzxz)
Math(sym.latex(r'\mathbf{R}_{xyz}(\alpha, \beta=0, \gamma)=') + \
sym.latex(Rzxz, mat_str='matrix'))
```
Despite different values of $\alpha$ and $\gamma$ the result is a single rotation around the $z$ axis given by the sum $\alpha+\gamma$. In this case, of the three degrees of freedom one was lost (the other degree of freedom was set by $\beta=0$). For movement analysis, this means for example that one angle will be undetermined because everything we know is the sum of the two angles obtained from the rotation matrix. We can set the unknown angle to zero but this is arbitrary.
In fact, we already dealt with another example of gimbal lock when we looked at the $xyz$ sequence with rotations by $90^o$. See the figure representing these rotations again and perceive that the first and third rotations were around the same axis because the second rotation was by $90^o$. Let's do the matrix multiplication replacing only the second angle by $90^o$ (and let's use the `euler_rotmat.py`:
```
Ra, Rn = euler_rotmat(order='xyz', frame='local', angles=[None, 90., None], showA=False)
```
Once again, one degree of freedom was lost and we will not be able to uniquely determine the three angles for the given rotation matrix and sequence.
Possible solutions to avoid the gimbal lock are: choose a different sequence; do not rotate the system by the angle that puts the system in gimbal lock (in the examples above, avoid $\beta=90^o$); or add an extra fourth parameter in the description of the rotation angles.
But if we have a physical system where we measure or specify exactly three Euler angles in a fixed sequence to describe or control it, and we can't avoid the system to assume certain angles, then we might have to say "Houston, we have a problem".
A famous situation where such a problem occurred was during the Apollo 13 mission. This is an actual conversation between crew and mission control during the Apollo 13 mission (Corke, 2011):
>`Mission clock: 02 08 12 47`
**Flight**: *Go, Guidance.*
**Guido**: *He’s getting close to gimbal lock there.*
**Flight**: *Roger. CapCom, recommend he bring up C3, C4, B3, B4, C1 and C2 thrusters, and advise he’s getting close to gimbal lock.*
**CapCom**: *Roger.*
*Of note, it was not a gimbal lock that caused the accident with the the Apollo 13 mission, the problem was an oxygen tank explosion.*
## Determination of the rotation matrix
A typical way to determine the rotation matrix for a rigid body in biomechanics is to use motion analysis to measure the position of at least three non-collinear markers placed on the rigid body, and then calculate a basis with these positions, analogue to what we have described in the notebook [Rigid-body transformations in a plane (2D)](http://nbviewer.ipython.org/github/demotu/BMC/blob/master/notebooks/Transformation2D.ipynb).
### Basis
If we have the position of three markers: **m1**, **m2**, **m3**, a basis (formed by three orthogonal versors) can be found as:
- First axis, **v1**, the vector **m2-m1**;
- Second axis, **v2**, the cross product between the vectors **v1** and **m3-m1**;
- Third axis, **v3**, the cross product between the vectors **v1** and **v2**.
Then, each of these vectors are normalized resulting in three orthogonal versors.
For example, given the positions m1 = [1,0,0], m2 = [0,1,0], m3 = [0,0,1], a basis can be found:
```
m1 = np.array([1, 0, 0])
m2 = np.array([0, 1, 0])
m3 = np.array([0, 0, 1])
v1 = m2 - m1
v2 = np.cross(v1, m3 - m1)
v3 = np.cross(v1, v2)
print('Versors:')
v1 = v1/np.linalg.norm(v1)
print('v1 =', v1)
v2 = v2/np.linalg.norm(v2)
print('v2 =', v2)
v3 = v3/np.linalg.norm(v3)
print('v3 =', v3)
print('\nNorm of each versor:\n',
np.linalg.norm(np.cross(v1, v2)),
np.linalg.norm(np.cross(v1, v3)),
np.linalg.norm(np.cross(v2, v3)))
```
Remember from the text [Rigid-body transformations in a plane (2D)](http://nbviewer.ipython.org/github/demotu/BMC/blob/master/notebooks/Transformation2D.ipynb) that the versors of this basis are the columns of the $\mathbf{R_{Gl}}$ and the rows of the $\mathbf{R_{lG}}$ rotation matrices, for instance:
```
RlG = np.array([v1, v2, v3])
print('Rotation matrix from Global to local coordinate system:\n', RlG)
```
And the corresponding angles of rotation using the $xyz$ sequence are:
```
euler_angles_from_rot_xyz(RlG)
```
These angles don't mean anything now because they are angles of the axes of the arbitrary basis we computed. In biomechanics, if we want an anatomical interpretation of the coordinate system orientation, we define the versors of the basis oriented with anatomical axes (e.g., for the shoulder, one versor would be aligned with the long axis of the upper arm).
We will see how to perform this computation later. Now we will combine translation and rotation in a single transformation.
## Translation and Rotation
Consider the case where the local coordinate system is translated and rotated in relation to the Global coordinate system as illustrated in the next figure.
<br>
<figure><img src='./../images/transrot3D.png' alt='translation and rotation 3D'/> <figcaption><center><i>Figure. A point in three-dimensional space represented in two coordinate systems, with one system translated and rotated.</i></center></figcaption> </figure>
The position of point $\mathbf{P}$ originally described in the local coordinate system, but now described in the Global coordinate system in vector form is:
$$ \mathbf{P_G} = \mathbf{L_G} + \mathbf{R_{Gl}}\mathbf{P_l} $$
This means that we first *disrotate* the local coordinate system and then correct for the translation between the two coordinate systems. Note that we can't invert this order: the point position is expressed in the local coordinate system and we can't add this vector to another vector expressed in the Global coordinate system, first we have to convert the vectors to the same coordinate system.
If now we want to find the position of a point at the local coordinate system given its position in the Global coordinate system, the rotation matrix and the translation vector, we have to invert the expression above:
$$ \begin{array}{l l}
\mathbf{P_G} = \mathbf{L_G} + \mathbf{R_{Gl}}\mathbf{P_l} \implies \\
\\
\mathbf{R_{Gl}^{-1}}\cdot\mathbf{P_G} = \mathbf{R_{Gl}^{-1}}\left(\mathbf{L_G} + \mathbf{R_{Gl}}\mathbf{P_l}\right) \implies \\
\\
\mathbf{R_{Gl}^{-1}}\mathbf{P_G} = \mathbf{R_{Gl}^{-1}}\mathbf{L_G} + \mathbf{R_{Gl}^{-1}}\mathbf{R_{Gl}}\mathbf{P_l} \implies \\
\\
\mathbf{P_l} = \mathbf{R_{Gl}^{-1}}\left(\mathbf{P_G}-\mathbf{L_G}\right) = \mathbf{R_{Gl}^T}\left(\mathbf{P_G}-\mathbf{L_G}\right) \;\;\;\;\; \text{or} \;\;\;\;\; \mathbf{P_l} = \mathbf{R_{lG}}\left(\mathbf{P_G}-\mathbf{L_G}\right)
\end{array} $$
The expression above indicates that to perform the inverse operation, to go from the Global to the local coordinate system, we first translate and then rotate the coordinate system.
### Transformation matrix
It is possible to combine the translation and rotation operations in only one matrix, called the transformation matrix:
$$ \begin{bmatrix}
\mathbf{P_X} \\
\mathbf{P_Y} \\
\mathbf{P_Z} \\
1
\end{bmatrix} =
\begin{bmatrix}
. & . & . & \mathbf{L_{X}} \\
. & \mathbf{R_{Gl}} & . & \mathbf{L_{Y}} \\
. & . & . & \mathbf{L_{Z}} \\
0 & 0 & 0 & 1
\end{bmatrix}
\begin{bmatrix}
\mathbf{P}_x \\
\mathbf{P}_y \\
\mathbf{P}_z \\
1
\end{bmatrix} $$
Or simply:
$$ \mathbf{P_G} = \mathbf{T_{Gl}}\mathbf{P_l} $$
Remember that in general the transformation matrix is not orthonormal, i.e., its inverse is not equal to its transpose.
The inverse operation, to express the position at the local coordinate system in terms of the Global reference system, is:
$$ \mathbf{P_l} = \mathbf{T_{Gl}^{-1}}\mathbf{P_G} $$
And in matrix form:
$$ \begin{bmatrix}
\mathbf{P_x} \\
\mathbf{P_y} \\
\mathbf{P_z} \\
1
\end{bmatrix} =
\begin{bmatrix}
\cdot & \cdot & \cdot & \cdot \\
\cdot & \mathbf{R^{-1}_{Gl}} & \cdot & -\mathbf{R^{-1}_{Gl}}\:\mathbf{L_G} \\
\cdot & \cdot & \cdot & \cdot \\
0 & 0 & 0 & 1
\end{bmatrix}
\begin{bmatrix}
\mathbf{P_X} \\
\mathbf{P_Y} \\
\mathbf{P_Z} \\
1
\end{bmatrix} $$
### Example with actual motion analysis data
*The data for this example is taken from page 183 of David Winter's book.*
Consider the following marker positions placed on a leg (described in the laboratory coordinate system with coordinates $x, y, z$ in cm, the $x$ axis points forward and the $y$ axes points upward): lateral malleolus (**lm** = [2.92, 10.10, 18.85]), medial malleolus (**mm** = [2.71, 10.22, 26.52]), fibular head (**fh** = [5.05, 41.90, 15.41]), and medial condyle (**mc** = [8.29, 41.88, 26.52]). Define the ankle joint center as the centroid between the **lm** and **mm** markers and the knee joint center as the centroid between the **fh** and **mc** markers. An anatomical coordinate system for the leg can be defined as: the quasi-vertical axis ($y$) passes through the ankle and knee joint centers; a temporary medio-lateral axis ($z$) passes through the two markers on the malleolus, an anterior-posterior as the cross product between the two former calculated orthogonal axes, and the origin at the ankle joint center.
a) Calculate the anatomical coordinate system for the leg as described above.
b) Calculate the rotation matrix and the translation vector for the transformation from the anatomical to the laboratory coordinate system.
c) Calculate the position of each marker and of each joint center at the anatomical coordinate system.
d) Calculate the Cardan angles using the $zxy$ sequence for the orientation of the leg with respect to the laboratory (but remember that the letters chosen to refer to axes are arbitrary, what matters is the directions they represent).
```
# calculation of the joint centers
mm = np.array([2.71, 10.22, 26.52])
lm = np.array([2.92, 10.10, 18.85])
fh = np.array([5.05, 41.90, 15.41])
mc = np.array([8.29, 41.88, 26.52])
ajc = (mm + lm)/2
kjc = (fh + mc)/2
print('Poition of the ankle joint center:', ajc)
print('Poition of the knee joint center:', kjc)
# calculation of the anatomical coordinate system axes (basis)
y = kjc - ajc
x = np.cross(y, mm - lm)
z = np.cross(x, y)
print('Versors:')
x = x/np.linalg.norm(x)
y = y/np.linalg.norm(y)
z = z/np.linalg.norm(z)
print('x =', x)
print('y =', y)
print('z =', z)
Oleg = ajc
print('\nOrigin =', Oleg)
# Rotation matrices
RGl = np.array([x, y , z]).T
print('Rotation matrix from the anatomical to the laboratory coordinate system:\n', RGl)
RlG = RGl.T
print('\nRotation matrix from the laboratory to the anatomical coordinate system:\n', RlG)
# Translational vector
OG = np.array([0, 0, 0]) # Laboratory coordinate system origin
LG = Oleg - OG
print('Translational vector from the anatomical to the laboratory coordinate system:\n', LG)
```
To get the coordinates from the laboratory (global) coordinate system to the anatomical (local) coordinate system:
$$ \mathbf{P_l} = \mathbf{R_{lG}}\left(\mathbf{P_G}-\mathbf{L_G}\right) $$
```
# position of each marker and of each joint center at the anatomical coordinate system
mml = np.dot(RlG, (mm - LG)) # equivalent to the algebraic expression RlG*(mm - LG).T
lml = np.dot(RlG, (lm - LG))
fhl = np.dot(RlG, (fh - LG))
mcl = np.dot(RlG, (mc - LG))
ajcl = np.dot(RlG, (ajc - LG))
kjcl = np.dot(RlG, (kjc - LG))
print('Coordinates of mm in the anatomical system:\n', mml)
print('Coordinates of lm in the anatomical system:\n', lml)
print('Coordinates of fh in the anatomical system:\n', fhl)
print('Coordinates of mc in the anatomical system:\n', mcl)
print('Coordinates of kjc in the anatomical system:\n', kjcl)
print('Coordinates of ajc in the anatomical system (origin):\n', ajcl)
```
## Problems
1. For the example about how the order of rotations of a rigid body affects the orientation shown in a figure above, deduce the rotation matrices for each of the 4 cases shown in the figure. For the first two cases, deduce the rotation matrices from the global to the local coordinate system and for the other two examples, deduce the rotation matrices from the local to the global coordinate system.
2. Consider the data from problem 7 in the notebook [Frame of reference](http://nbviewer.ipython.org/github/demotu/BMC/blob/master/notebooks/ReferenceFrame.ipynb) where the following anatomical landmark positions are given (units in meters): RASIS=[0.5,0.8,0.4], LASIS=[0.55,0.78,0.1], RPSIS=[0.3,0.85,0.2], and LPSIS=[0.29,0.78,0.3]. Deduce the rotation matrices for the global to anatomical coordinate system and for the anatomical to global coordinate system.
3. For the data from the last example, calculate the Cardan angles using the $zxy$ sequence for the orientation of the leg with respect to the laboratory (but remember that the letters chosen to refer to axes are arbitrary, what matters is the directions they represent).
## References
- Corke P (2011) [Robotics, Vision and Control: Fundamental Algorithms in MATLAB](http://www.petercorke.com/RVC/). Springer-Verlag Berlin.
- Robertson G, Caldwell G, Hamill J, Kamen G (2013) [Research Methods in Biomechanics](http://books.google.com.br/books?id=gRn8AAAAQBAJ). 2nd Edition. Human Kinetics.
- [Maths - Euler Angles](http://www.euclideanspace.com/maths/geometry/rotations/euler/).
- Murray RM, Li Z, Sastry SS (1994) [A Mathematical Introduction to Robotic Manipulation](http://www.cds.caltech.edu/~murray/mlswiki/index.php/Main_Page). Boca Raton, CRC Press.
- Ruina A, Rudra P (2013) [Introduction to Statics and Dynamics](http://ruina.tam.cornell.edu/Book/index.html). Oxford University Press.
- Siciliano B, Sciavicco L, Villani L, Oriolo G (2009) [Robotics - Modelling, Planning and Control](http://books.google.com.br/books/about/Robotics.html?hl=pt-BR&id=jPCAFmE-logC). Springer-Verlag London.
- Winter DA (2009) [Biomechanics and motor control of human movement](http://books.google.com.br/books?id=_bFHL08IWfwC). 4 ed. Hoboken, USA: Wiley.
- Zatsiorsky VM (1997) [Kinematics of Human Motion](http://books.google.com.br/books/about/Kinematics_of_Human_Motion.html?id=Pql_xXdbrMcC&redir_esc=y). Champaign, Human Kinetics.
## Function `euler_rotmatrix.py`
```
# %load ./../functions/euler_rotmat.py
#!/usr/bin/env python
"""Euler rotation matrix given sequence, frame, and angles."""
from __future__ import division, print_function
__author__ = 'Marcos Duarte, https://github.com/demotu/BMC'
__version__ = 'euler_rotmat.py v.1 2014/03/10'
def euler_rotmat(order='xyz', frame='local', angles=None, unit='deg',
str_symbols=None, showA=True, showN=True):
"""Euler rotation matrix given sequence, frame, and angles.
This function calculates the algebraic rotation matrix (3x3) for a given
sequence ('order' argument) of up to three elemental rotations of a given
coordinate system ('frame' argument) around another coordinate system, the
Euler (or Eulerian) angles [1]_.
This function also calculates the numerical values of the rotation matrix
when numerical values for the angles are inputed for each rotation axis.
Use None as value if the rotation angle for the particular axis is unknown.
The symbols for the angles are: alpha, beta, and gamma for the first,
second, and third rotations, respectively.
The matrix product is calulated from right to left and in the specified
sequence for the Euler angles. The first letter will be the first rotation.
The function will print and return the algebraic rotation matrix and the
numerical rotation matrix if angles were inputed.
Parameters
----------
order : string, optional (default = 'xyz')
Sequence for the Euler angles, any combination of the letters
x, y, and z with 1 to 3 letters is accepted to denote the
elemental rotations. The first letter will be the first rotation.
frame : string, optional (default = 'local')
Coordinate system for which the rotations are calculated.
Valid values are 'local' or 'global'.
angles : list, array, or bool, optional (default = None)
Numeric values of the rotation angles ordered as the 'order'
parameter. Enter None for a rotation whith unknown value.
unit : str, optional (default = 'deg')
Unit of the input angles.
str_symbols : list of strings, optional (default = None)
New symbols for the angles, for instance, ['theta', 'phi', 'psi']
showA : bool, optional (default = True)
True (1) displays the Algebraic rotation matrix in rich format.
False (0) to not display.
showN : bool, optional (default = True)
True (1) displays the Numeric rotation matrix in rich format.
False (0) to not display.
Returns
-------
R : Matrix Sympy object
Rotation matrix (3x3) in algebraic format.
Rn : Numpy array or Matrix Sympy object (only if angles are inputed)
Numeric rotation matrix (if values for all angles were inputed) or
a algebraic matrix with some of the algebraic angles substituted
by the corresponding inputed numeric values.
Notes
-----
This code uses Sympy, the Python library for symbolic mathematics, to
calculate the algebraic rotation matrix and shows this matrix in latex form
possibly for using with the IPython Notebook, see [1]_.
References
----------
.. [1] http://nbviewer.ipython.org/github/duartexyz/BMC/blob/master/Transformation3D.ipynb
Examples
--------
>>> # import function
>>> from euler_rotmat import euler_rotmat
>>> # Default options: xyz sequence, local frame and show matrix
>>> R = euler_rotmat()
>>> # XYZ sequence (around global (fixed) coordinate system)
>>> R = euler_rotmat(frame='global')
>>> # Enter numeric values for all angles and show both matrices
>>> R, Rn = euler_rotmat(angles=[90, 90, 90])
>>> # show what is returned
>>> euler_rotmat(angles=[90, 90, 90])
>>> # show only the rotation matrix for the elemental rotation at x axis
>>> R = euler_rotmat(order='x')
>>> # zxz sequence and numeric value for only one angle
>>> R, Rn = euler_rotmat(order='zxz', angles=[None, 0, None])
>>> # input values in radians:
>>> import numpy as np
>>> R, Rn = euler_rotmat(order='zxz', angles=[None, np.pi, None], unit='rad')
>>> # shows only the numeric matrix
>>> R, Rn = euler_rotmat(order='zxz', angles=[90, 0, None], showA='False')
>>> # Change the angles' symbols
>>> R = euler_rotmat(order='zxz', str_symbols=['theta', 'phi', 'psi'])
>>> # Negativate the angles' symbols
>>> R = euler_rotmat(order='zxz', str_symbols=['-theta', '-phi', '-psi'])
>>> # all algebraic matrices for all possible sequences for the local frame
>>> s=['xyz','xzy','yzx','yxz','zxy','zyx','xyx','xzx','yzy','yxy','zxz','zyz']
>>> for seq in s: R = euler_rotmat(order=seq)
>>> # all algebraic matrices for all possible sequences for the global frame
>>> for seq in s: R = euler_rotmat(order=seq, frame='global')
"""
import numpy as np
import sympy as sym
try:
from IPython.core.display import Math, display
ipython = True
except:
ipython = False
angles = np.asarray(np.atleast_1d(angles), dtype=np.float64)
if ~np.isnan(angles).all():
if len(order) != angles.size:
raise ValueError("Parameters 'order' and 'angles' (when " +
"different from None) must have the same size.")
x, y, z = sym.symbols('x, y, z')
sig = [1, 1, 1]
if str_symbols is None:
a, b, g = sym.symbols('alpha, beta, gamma')
else:
s = str_symbols
if s[0][0] == '-': s[0] = s[0][1:]; sig[0] = -1
if s[1][0] == '-': s[1] = s[1][1:]; sig[1] = -1
if s[2][0] == '-': s[2] = s[2][1:]; sig[2] = -1
a, b, g = sym.symbols(s)
var = {'x': x, 'y': y, 'z': z, 0: a, 1: b, 2: g}
# Elemental rotation matrices for xyz (local)
cos, sin = sym.cos, sym.sin
Rx = sym.Matrix([[1, 0, 0], [0, cos(x), sin(x)], [0, -sin(x), cos(x)]])
Ry = sym.Matrix([[cos(y), 0, -sin(y)], [0, 1, 0], [sin(y), 0, cos(y)]])
Rz = sym.Matrix([[cos(z), sin(z), 0], [-sin(z), cos(z), 0], [0, 0, 1]])
if frame.lower() == 'global':
Rs = {'x': Rx.T, 'y': Ry.T, 'z': Rz.T}
order = order.upper()
else:
Rs = {'x': Rx, 'y': Ry, 'z': Rz}
order = order.lower()
R = Rn = sym.Matrix(sym.Identity(3))
str1 = r'\mathbf{R}_{%s}( ' %frame # last space needed for order=''
#str2 = [r'\%s'%var[0], r'\%s'%var[1], r'\%s'%var[2]]
str2 = [1, 1, 1]
for i in range(len(order)):
Ri = Rs[order[i].lower()].subs(var[order[i].lower()], sig[i] * var[i])
R = Ri * R
if sig[i] > 0:
str2[i] = '%s:%s' %(order[i], sym.latex(var[i]))
else:
str2[i] = '%s:-%s' %(order[i], sym.latex(var[i]))
str1 = str1 + str2[i] + ','
if ~np.isnan(angles).all() and ~np.isnan(angles[i]):
if unit[:3].lower() == 'deg':
angles[i] = np.deg2rad(angles[i])
Rn = Ri.subs(var[i], angles[i]) * Rn
#Rn = sym.lambdify(var[i], Ri, 'numpy')(angles[i]) * Rn
str2[i] = str2[i] + '=%.0f^o' %np.around(np.rad2deg(angles[i]), 0)
else:
Rn = Ri * Rn
Rn = sym.simplify(Rn) # for trigonometric relations
try:
# nsimplify only works if there are symbols
Rn2 = sym.latex(sym.nsimplify(Rn, tolerance=1e-8).n(chop=True, prec=4))
except:
Rn2 = sym.latex(Rn.n(chop=True, prec=4))
# there are no symbols, pass it as Numpy array
Rn = np.asarray(Rn)
if showA and ipython:
display(Math(str1[:-1] + ') =' + sym.latex(R, mat_str='matrix')))
if showN and ~np.isnan(angles).all() and ipython:
str2 = ',\;'.join(str2[:angles.size])
display(Math(r'\mathbf{R}_{%s}(%s)=%s' %(frame, str2, Rn2)))
if np.isnan(angles).all():
return R
else:
return R, Rn
```
| github_jupyter |
# Design of Digital Filters
*This jupyter notebook is part of a [collection of notebooks](../index.ipynb) on various topics of Digital Signal Processing.
## Example: Non-Recursive versus Recursive Filter
In the following example, the characteristics and computational complexity of a non-recursive and a recursive filter are compared for a particular design. Quantization is not considered. In order to design the filters we need to specify the requirements. This is typically done by a *tolerance scheme*. The scheme states the desired frequency response and allowed deviations. This is explained at an example.
We aim at the design of a low-pass filter with
1. unit amplitude with an allowable symmetric deviation of $\delta_\text{p}$ for $|\Omega| < \Omega_\text{p}$
2. an attenuation of $a_\text{s}$ for $|\Omega| > \Omega_\text{s}$
where the indices p and s denote the pass- and stop-band, respectively. The region between the pass-band $\Omega_\text{p}$ and the stop-band $\Omega_\text{s}$ is known as *transition-band*. The phase of the filter is not specified.
The resulting tolerance scheme is illustrated for the design parameters $\Omega_\text{p} = \frac{\pi}{3}$, $\Omega_\text{s} = \frac{\pi}{3} + 0.05$, $\delta_\text{p} = 1.5$ dB and $a_\text{s} = -60$ dB.
```
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
import scipy.signal as sig
def plot_tolerance_scheme(Omp, Oms, d_p, a_s):
Omp = Omp * np.pi
Oms = Oms * np.pi
p = [[0, -d_p], [Omp, -d_p], [Omp, -300], [np.pi, -300], [np.pi, a_s], [Oms, a_s], [Oms, d_p], [0, d_p]]
polygon = mpatches.Polygon(p, closed=True, facecolor='r', alpha=0.3)
plt.gca().add_patch(polygon)
Omp = .3 # normalized corner frequency of pass-band
Oms = .3 + 0.05 # normalized corner frequency of stop-band
d_p = 1.5 # one-sided pass-band ripple in dB
a_s = -60 # stop-band attenuation in dB
plt.figure(figsize = (10, 5))
plot_tolerance_scheme(Omp, Oms, d_p, a_s)
plt.title('Tolerance scheme')
plt.xlabel(r'$\Omega$')
plt.ylabel(r'$|H(e^{j \Omega})|$ in dB')
plt.axis([0, np.pi, -70, 3])
plt.grid();
```
**Exercise**
* What corner frequencies $f_\text{p}$ and $f_\text{s}$ result for a sampling frequency of $f_\text{s} = 48$ kHz?
Solution: It follows that $f_\text{p} = \frac{\Omega_\text{p}}{\pi} \cdot \frac{f_\text{s}}{2} = 8$ kHz and $f_\text{s} = \frac{\Omega_\text{s}}{\pi} \cdot \frac{f_\text{s}}{2} \approx 8.4$ kHz, since the normalized frequency $\Omega = \pi$ corresponds to $\frac{f_\text{s}}{2}$.
The comparison of non-recursive and recursive filters depends heavily on the chosen filter design algorithm. For the design of the non-recursive filter a technique is used which bases on numerical optimization of the filter coefficients with respect to the desired response. The [Remez algorithm](https://en.wikipedia.org/wiki/Remez_algorithm), as implemented in `scipy.signal.remez`, is used for this purpose. The parameters for the algorithm are the corner frequencies of the pass- and stop-band, as well as the desired attenuation in the stop-band. For the recursive filter, a [Chebyshev type II](https://en.wikipedia.org/wiki/Chebyshev_filter) design is used. Here the parameters are the corner frequency and attenuation of the stop-band. The order of both filters has been chosen manually to fit the given tolerance scheme.
```
N = 152 # length of non-recursive filter
M = 13 # order of recursive filter
# design of non-recursive filter
h = sig.remez(N, [0, Omp/2, Oms/2, 1/2], [1, 10**((a_s-5)/20)], weight=[1, 1])
# design of recursive filter
b, a = sig.cheby2(M, -a_s, Oms)
# compute frequency response of filter
Om, Hn = sig.freqz(h, worN=8192)
Om, Hr = sig.freqz(b, a, worN=8192)
# plot frequency response
plt.figure(figsize = (10,5))
plt.plot(Om, 20*np.log10(np.abs(Hn)), 'b-', label=r'non-recursive N=%d'%N)
plt.plot(Om, 20*np.log10(np.abs(Hr)), 'g-', label=r'recursive N=%d'%M)
plot_tolerance_scheme(Omp, Oms, d_p, a_s)
plt.title('Magnitude response')
plt.xlabel(r'$\Omega$')
plt.ylabel(r'$|H(e^{j \Omega})|$ in dB')
plt.legend()
plt.axis([0, np.pi, -70, 3])
plt.grid()
# plot phase
plt.figure(figsize = (10,5))
plt.plot(Om, np.unwrap(np.angle(Hn)), label=r'non-recursive N=%d'%N)
plt.plot(Om, np.unwrap(np.angle(Hr)), label=r'recursive N=%d'%M)
plt.title('Phase response')
plt.xlabel(r'$\Omega$')
plt.ylabel(r'$\varphi(\Omega)$ in rad')
plt.legend(loc=3)
plt.xlim([0, np.pi])
plt.grid()
```
**Exercises**
* How do both designs differ in terms of their magnitude and phase responses?
* Calculate the number of multiplications and additions required to realize the non-recursive filter
* Calculate the number of multiplications and additions required to realize the recursive filter in [transposed direct form II](../recursive_filters/direct_forms.ipynb#Transposed-Direct-Form-II)
* Decrease the corner frequencies and adapt the order of the filters to match the tolerance scheme
Solution: Inspection of the magnitude response $|H(e^{j \Omega})|$ for the designed non-recursive and recursive filters reveals that both fulfill the given tolerance scheme. An obvious difference between both filters is the structure of the magnitude response in the stop-band $\Omega > \Omega_\text{s}$. While the magnitude of the non-recursive filter shows a high number of fluctuations below the desired attenuation, these are much less for the recursive filter. This is a consequence of the different orders of the filters and their respective number of zeros. The non-recursive filter requires $N$ multiplications and $N-1$ additions to compute one output sample, hence 152 multiplications and 151 additions. The recursive filter in transposed direct form II is realized by 7 SOS. Each of the SOS requires 5 multiplications and 4 additions per output sample, resulting in a total of 35 multiplications and 28 additions.
In order to evaluate the computational complexity of both filters, the execution time is measured when filtering a signal $x[k]$ of length $L=10^5$ samples. The non-recursive filter is realized by direct convolution, the recursive filter in transposed direct form II using the respective Python functions.
```
import timeit
reps = 1000 # number of repetitions for timeit
# setup environment for timeit
tsetup = 'import numpy as np; import scipy.signal as sig; from __main__ import h, a, b; x=np.random.normal(size=int(1e5))'
# non-recursive filter
tn = timeit.timeit('np.convolve(x, h, mode="full")', setup=tsetup, number=reps)
# recursive filter
tr = timeit.timeit('sig.lfilter(b, a, x)' , setup=tsetup, number=reps)
# show the results
plt.figure(figsize = (5, 3))
plt.bar(1, tn/reps*1000)
plt.bar(2, tr/reps*1000)
plt.title('Execution time')
plt.xticks([1, 2], ('non-recursive', 'recursive'))
plt.ylabel('time in ms')
plt.grid()
```
**Exercises**
* Do the execution times correspond with the number of algorithmic operations calculated in the previous exercise?
* Estimate the computational load for the filtering of a signal with a sampling rate of 48 kHz
* How could the execution time of the non-recursive filter be decreased?
* Finally, would you prefer the non-recursive or the recursive design for a practical implementation? Consider the numerical complexity, as well as numerical aspects in your decision.
Solution: On general purpose processors, the numerical complexity is mainly determined by the number of multiplications. The ratio of multiplications per output sample for the non-recursive and the recursive filter is given as $\frac{152}{35} \approx 4.3$, the ratio of execution times in above example as $\frac{4.8 \mathrm{ ms}}{1.5 \mathrm{ ms}} \approx 3.2$. The difference between both can be related to the implementation of both methods and their execution on the given hardware. Note that the execution times and their ratio may differ for other environments. The number of samples used in the measurement above relates to a signal with $\frac{10^5}{f_s} \approx 2$ seconds length. The computational load for the non-recursive filter can hence be estimated as $\frac{4.8 \mathrm{ ms}}{2000 \mathrm{ ms}} \approx 2.4 \cdot 10^{-6}$. The execution time for the non-recursive filter may be decreased by using a fast convolution algorithm.
| github_jupyter |
# Visualizing tweets and the Logistic Regression model
**Objectives:** Visualize and interpret the logistic regression model
**Steps:**
* Plot tweets in a scatter plot using their positive and negative sums.
* Plot the output of the logistic regression model in the same plot as a solid line
## Import the required libraries
We will be using [*NLTK*](http://www.nltk.org/howto/twitter.html), an opensource NLP library, for collecting, handling, and processing Twitter data. In this lab, we will use the example dataset that comes alongside with NLTK. This dataset has been manually annotated and serves to establish baselines for models quickly.
So, to start, let's import the required libraries.
```
import nltk # NLP toolbox
from os import getcwd
import pandas as pd # Library for Dataframes
from nltk.corpus import twitter_samples
import matplotlib.pyplot as plt # Library for visualization
import numpy as np # Library for math functions
from utils import process_tweet, build_freqs # Our functions for NLP
```
## Load the NLTK sample dataset
To complete this lab, you need the sample dataset of the previous lab. Here, we assume the files are already available, and we only need to load into Python lists.
```
# select the set of positive and negative tweets
all_positive_tweets = twitter_samples.strings('positive_tweets.json')
all_negative_tweets = twitter_samples.strings('negative_tweets.json')
tweets = all_positive_tweets + all_negative_tweets ## Concatenate the lists.
labels = np.append(np.ones((len(all_positive_tweets),1)), np.zeros((len(all_negative_tweets),1)), axis = 0)
# split the data into two pieces, one for training and one for testing (validation set)
train_pos = all_positive_tweets[:4000]
train_neg = all_negative_tweets[:4000]
train_x = train_pos + train_neg
print("Number of tweets: ", len(train_x))
```
# Load the extracted features
Part of this week's assignment is the creation of the numerical features needed for the Logistic regression model. In order not to interfere with it, we have previously calculated and stored these features in a CSV file for the entire training set.
So, please load these features created for the tweets sample.
```
data = pd.read_csv('logistic_features.csv'); # Load a 3 columns csv file using pandas function
data.head(10) # Print the first 10 data entries
```
Now let us get rid of the data frame to keep only Numpy arrays.
```
# Each feature is labeled as bias, positive and negative
X = data[['bias', 'positive', 'negative']].values # Get only the numerical values of the dataframe
Y = data['sentiment'].values; # Put in Y the corresponding labels or sentiments
print(X.shape) # Print the shape of the X part
print(X) # Print some rows of X
```
## Load a pretrained Logistic Regression model
In the same way, as part of this week's assignment, a Logistic regression model must be trained. The next cell contains the resulting model from such training. Notice that a list of 3 numeric values represents the whole model, that we have called _theta_ $\theta$.
```
theta = [7e-08, 0.0005239, -0.00055517]
```
## Plot the samples in a scatter plot
The vector theta represents a plane that split our feature space into two parts. Samples located over that plane are considered positive, and samples located under that plane are considered negative. Remember that we have a 3D feature space, i.e., each tweet is represented as a vector comprised of three values: `[bias, positive_sum, negative_sum]`, always having `bias = 1`.
If we ignore the bias term, we can plot each tweet in a cartesian plane, using `positive_sum` and `negative_sum`. In the cell below, we do precisely this. Additionally, we color each tweet, depending on its class. Positive tweets will be green and negative tweets will be red.
```
# Plot the samples using columns 1 and 2 of the matrix
fig, ax = plt.subplots(figsize = (8, 8))
colors = ['red', 'green']
# Color based on the sentiment Y
ax.scatter(X[:,1], X[:,2], c=[colors[int(k)] for k in Y], s = 0.1) # Plot a dot for each pair of words
plt.xlabel("Positive")
plt.ylabel("Negative")
```
From the plot, it is evident that the features that we have chosen to represent tweets as numerical vectors allow an almost perfect separation between positive and negative tweets. So you can expect a very high accuracy for this model!
## Plot the model alongside the data
We will draw a gray line to show the cutoff between the positive and negative regions. In other words, the gray line marks the line where $$ z = \theta * x = 0.$$
To draw this line, we have to solve the above equation in terms of one of the independent variables.
$$ z = \theta * x = 0$$
$$ x = [1, pos, neg] $$
$$ z(\theta, x) = \theta_0+ \theta_1 * pos + \theta_2 * neg = 0 $$
$$ neg = (-\theta_0 - \theta_1 * pos) / \theta_2 $$
The red and green lines that point in the direction of the corresponding sentiment are calculated using a perpendicular line to the separation line calculated in the previous equations(neg function). It must point in the same direction as the derivative of the Logit function, but the magnitude may differ. It is only for a visual representation of the model.
$$direction = pos * \theta_2 / \theta_1$$
```
# Equation for the separation plane
# It give a value in the negative axe as a function of a positive value
# f(pos, neg, W) = w0 + w1 * pos + w2 * neg = 0
# s(pos, W) = (w0 - w1 * pos) / w2
def neg(theta, pos):
return (-theta[0] - pos * theta[1]) / theta[2]
# Equation for the direction of the sentiments change
# We don't care about the magnitude of the change. We are only interested
# in the direction. So this direction is just a perpendicular function to the
# separation plane
# df(pos, W) = pos * w2 / w1
def direction(theta, pos):
return pos * theta[2] / theta[1]
```
The green line in the chart points in the direction where z > 0 and the red line points in the direction where z < 0. The direction of these lines are given by the weights $\theta_1$ and $\theta_2$
```
# Plot the samples using columns 1 and 2 of the matrix
fig, ax = plt.subplots(figsize = (8, 8))
colors = ['red', 'green']
# Color base on the sentiment Y
ax.scatter(X[:,1], X[:,2], c=[colors[int(k)] for k in Y], s = 0.1) # Plot a dot for each pair of words
plt.xlabel("Positive")
plt.ylabel("Negative")
# Now lets represent the logistic regression model in this chart.
maxpos = np.max(X[:,1])
print(maxpos)
offset = 5000 # The pos value for the direction vectors origin
# Plot a gray line that divides the 2 areas.
#arrow(x, y, dx, dy, **kwargs): This draws an arrow from (x, y) to (x+dx, y+dy).
ax.plot([0, maxpos], [neg(theta, 0), neg(theta, maxpos)], color = 'gray')
print([0, maxpos], [neg(theta, 0), neg(theta, maxpos)])
# Plot a green line pointing to the positive direction
ax.arrow(offset, neg(theta, offset), offset, direction(theta, offset), head_width=500, head_length=500, fc='g', ec='g')
#fc: 箭头颜色, ec: 线颜色
# Plot a red line pointing to the negative direction
ax.arrow(offset, neg(theta, offset), -offset, -direction(theta, offset), head_width=500, head_length=500, fc='r', ec='r')
print( -offset, -direction(theta, offset))
plt.show()
```
**Note that more critical than the Logistic regression itself, are the features extracted from tweets that allow getting the right results in this exercise.**
That is all, folks. Hopefully, now you understand better what the Logistic regression model represents, and why it works that well for this specific problem.
| github_jupyter |
# Student-t Process
PyMC3 also includes T-process priors. They are a generalization of a Gaussian process prior to the multivariate Student's T distribution. The usage is identical to that of `gp.Latent`, except they require a degrees of freedom parameter when they are specified in the model. For more information, see chapter 9 of [Rasmussen+Williams](http://www.gaussianprocess.org/gpml/), and [Shah et al.](https://arxiv.org/abs/1402.4306).
Note that T processes aren't additive in the same way as GPs, so addition of `TP` objects are not supported.
## Samples from a TP prior
The following code draws samples from a T process prior with 3 degrees of freedom and a Gaussian process, both with the same covariance matrix.
```
import pymc3 as pm
import theano.tensor as tt
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
# set the seed
np.random.seed(1)
n = 100 # The number of data points
X = np.linspace(0, 10, n)[:, None] # The inputs to the GP, they must be arranged as a column vector
# Define the true covariance function and its parameters
ℓ_true = 1.0
η_true = 3.0
cov_func = η_true**2 * pm.gp.cov.Matern52(1, ℓ_true)
# A mean function that is zero everywhere
mean_func = pm.gp.mean.Zero()
# The latent function values are one sample from a multivariate normal
# Note that we have to call `eval()` because PyMC3 built on top of Theano
tp_samples = pm.MvStudentT.dist(mu=mean_func(X).eval(), cov=cov_func(X).eval(), nu=3).random(size=8)
## Plot samples from TP prior
fig = plt.figure(figsize=(12,5)); ax = fig.gca()
ax.plot(X.flatten(), tp_samples.T, lw=3, alpha=0.6);
ax.set_xlabel("X"); ax.set_ylabel("y"); ax.set_title("Samples from TP with DoF=3");
gp_samples = pm.MvNormal.dist(mu=mean_func(X).eval(), cov=cov_func(X).eval()).random(size=8)
fig = plt.figure(figsize=(12,5)); ax = fig.gca()
ax.plot(X.flatten(), gp_samples.T, lw=3, alpha=0.6);
ax.set_xlabel("X"); ax.set_ylabel("y"); ax.set_title("Samples from GP");
```
## Poisson data generated by a T process
For the Poisson rate, we take the square of the function represented by the T process prior.
```
np.random.seed(7)
n = 150 # The number of data points
X = np.linspace(0, 10, n)[:, None] # The inputs to the GP, they must be arranged as a column vector
# Define the true covariance function and its parameters
ℓ_true = 1.0
η_true = 3.0
cov_func = η_true**2 * pm.gp.cov.ExpQuad(1, ℓ_true)
# A mean function that is zero everywhere
mean_func = pm.gp.mean.Zero()
# The latent function values are one sample from a multivariate normal
# Note that we have to call `eval()` because PyMC3 built on top of Theano
f_true = pm.MvStudentT.dist(mu=mean_func(X).eval(), cov=cov_func(X).eval(), nu=3).random(size=1)
y = np.random.poisson(f_true**2)
fig = plt.figure(figsize=(12,5)); ax = fig.gca()
ax.plot(X, f_true**2, "dodgerblue", lw=3, label="True f");
ax.plot(X, y, 'ok', ms=3, label="Data");
ax.set_xlabel("X"); ax.set_ylabel("y"); plt.legend();
with pm.Model() as model:
ℓ = pm.Gamma("ℓ", alpha=2, beta=2)
η = pm.HalfCauchy("η", beta=3)
cov = η**2 * pm.gp.cov.ExpQuad(1, ℓ)
# informative prior on degrees of freedom < 5
ν = pm.Gamma("ν", alpha=2, beta=1)
tp = pm.gp.TP(cov_func=cov, nu=ν)
f = tp.prior("f", X=X)
# adding a small constant seems to help with numerical stability here
y_ = pm.Poisson("y", mu=tt.square(f) + 1e-6, observed=y)
tr = pm.sample(1000)
pm.traceplot(tr, varnames=["ℓ", "ν", "η"], lines={"ℓ": ℓ_true, "η": η_true, "ν": 3});
n_new = 200
X_new = np.linspace(0, 15, n_new)[:,None]
# add the GP conditional to the model, given the new X values
with model:
f_pred = tp.conditional("f_pred", X_new)
# Sample from the GP conditional distribution
with model:
pred_samples = pm.sample_ppc(tr, vars=[f_pred], samples=1000)
fig = plt.figure(figsize=(12,5)); ax = fig.gca()
from pymc3.gp.util import plot_gp_dist
plot_gp_dist(ax, np.square(pred_samples["f_pred"]), X_new);
plt.plot(X, np.square(f_true), "dodgerblue", lw=3, label="True f");
plt.plot(X, y, 'ok', ms=3, alpha=0.5, label="Observed data");
plt.xlabel("X"); plt.ylabel("True f(x)"); plt.ylim([-2, 20])
plt.title("Conditional distribution of f_*, given f"); plt.legend();
```
| github_jupyter |
# GMNS Format Validation for networks stored as CSV files
This notebook demonstrates validation for whether a GMNS network conforms to the schema.
It uses a modified version of [GMNSpy](https://github.com/e-lo/GMNSpy), originally developed by Elizabeth Sall.
The first time you run this notebook after cloning this repo, you may need to run the following commands to update and install the working copy of `gmnspy`:
```
!git submodule update --init --recursive --remote --merge
# if you don't have command-line git, instead download this zip file:
# https://github.com/ianberg-volpe/GMNSpy/archive/refs/heads/hide_output.zip
# and extract the contents of the `GMNSpy-hide_output` folder in that zip archive
# into the folder named `gmnspy` in the same directory as this notebook.
!pip install ./gmnspy
```
## Inputs
GMNSpy takes CSV files as inputs for the network. Place all network files in a single directory.
The validation constraints are checked using a set of JSON files. `gmns.spec.json` provides information about required files and paths to the JSON definition for each table. Each table has its own `.schema.json` file which defines required fields and field constraints. These may be edited to meet a user's specific needs (e.g., to add user-defined fields, or to relax constraints).
## Outputs
Reading a GMNS network using the command below checks the set of files in the `data_directory` against the spec defined by `config`. The script currently performs the following checks:
- Checks whether the required tables, as defined in `gmns.spec.json`, are present.
- Checks each file in the `data_directory` whose name matches one defined in the spec with its associated `.schema.json` file. The following checks are performed:
- whether any required fields are missing (report a FAIL message if so).
- whether any fields outside the spec are present (report a WARN message if so).
- whether the values present in each field have the same datatype (integer, number, boolean, string) as required by the spec (report a FAIL message if so).
- whether any required fields have missing values (report a FAIL message if so).
- whether the primary key field has any duplicate values (report a FAIL message if so).
- whether any values in fields with strict constraints (minimum, maximum, enum) fall outside of those constraints (report a FAIL message if so).
- whether any values in fields with warning constraints (minimum, maximum) fall outside of those constraints (report a WARN message if so).
- Checks the foreign keys specified in each table. The following checks are performed:
- whether the foreign key specified exists on the reference table (report a FAIL message if not).
- whether the foreign key specified has unique values on the reference table (report a FAIL message if not).
- whether all values of the foreign key contained in a source table exist on the reference table (report a FAIL message if not).
```
import gmnspy
out = gmnspy.in_out.read_gmns_network(data_directory = "../Small_Network_Examples/Arlington_Signals", config = "gmnspy/spec/gmns.spec.json")
```
| github_jupyter |
# Census- Employment Status Data
```
import pandas as pd
import requests
#Census Subject Table API for Employment Status data within Unified School Districts in California for 2018
url="https://api.census.gov/data/2016/acs/acs1/subject?get=group(S2301)&for=school%20district%20(unified)&in=state:06"
#Request for HTTP Data from Census API, which is working <Response [200]>
response = requests.get(url)
#Resetting data from API Data for future formatting, lists data in one column
response_json = response.json()
#Places data in a dataframe and drops index column 0 (with headers), which has 117 schools and 702 columns of variables
unifiedschool_df = pd.DataFrame(response_json,columns=response_json[0]).drop(0)
unifiedschool_df
#unifiedschool_df.to_csv("/Users/nataligracia/git/ca-school-enrollment-trend/Unified2016test.csv")
#View School District to view column titles, 2016 data does not include district name
#unifiedschool_df.loc[unifiedschool_df["NAME"]=="Los Angeles Unified School District, California"]
```
## _Format Data_
```
#Census Subject Table Variables for Employment Status data
variableurl = "https://api.census.gov/data/2016/acs/acs1/subject/variables.json"
#Request for HTTP Data from Census API and reset data
variables_json = requests.get(variableurl).json()
#View variable for column title
variables_json["variables"]["S2301_C01_001E"]
#Find and replace all columns with variable titles
new_labels = []
for col in unifiedschool_df.columns:
label = variables_json["variables"].get(col)
if label is not None:
label = label['label']
else:
if col[-2:] == 'EA':
label = variables_json["variables"].get(col[:-1])
label = label['label'] + "||Annotation"
elif col[-1] == 'M':
label = variables_json["variables"].get(col[:-1]+'E')
label = label['label'] + "||MarginOfError"
elif col[-2:] == 'MA':
label = variables_json["variables"].get(col[:-2]+'E')
label = label['label'] + "||MarginOfErrorAnnotation"
new_labels.append(label)
#Find any columns without titles
new_labels
#Change column titles for columns labeled "None"
assert len(new_labels) == len(unifiedschool_df.columns)
#Confirm the number of columns without titles
sum([1 for x in new_labels if x is None])
#Setup new Labels of columns labeled "None"
#new_labels[-283] = 'NAME'
new_labels[-2] = 'STATE'
new_labels[-1] = 'SCHOOL DISTRICT (UNIFIED)'
#Create new labels of columns labeled "None"
unifiedschool_df.columns = new_labels
#Find all columns without "Annotation" in column title
[col for col in unifiedschool_df.columns if "Annotation" not in col]
#Create a new dataframe for data without the columns that have "Annotation" in the title, which is 702 columns
without_annotation = unifiedschool_df[[col for col in unifiedschool_df.columns if "Annotation" not in col]].copy()
without_annotation.head()
#Find all columns without "MarginOfError" in column title
#[col for col in without_annotation_df.columns if "MarginOfError" not in col]
#Create a new dataframe for data without the columns that have "MarginOfError" in the title, which is 142 columns
withoutmarginerror = without_annotation[[col for col in without_annotation.columns if 'MarginOfError' not in col]].copy()
withoutmarginerror.head()
#Find all columns without "Labor Force Participation Rate" in column title
#[col for col in withoutmarginerror.columns if "Labor Force Participation Rate" not in col]
#Create a new dataframe for data without the columns that have "Labor Force Participation Rate" in the title, which is 107 columns
withoutlaborforce = withoutmarginerror[[col for col in withoutmarginerror.columns if 'Labor Force Participation Rate' not in col]].copy()
withoutlaborforce.head()
#Find all columns without "Sex" in column title
#[col for col in withoutlaborforce.columns if "SEX" not in col]
#Create a new dataframe for data without the columns that have "Sex" in the title, which is 89 columns
withoutsex = withoutlaborforce[[col for col in withoutlaborforce.columns if 'SEX' not in col]].copy()
withoutsex.head()
#Find all columns without "Poverty Status" in column title
#[col for col in withoutsex.columns if "POVERTY STATUS" not in col]
#Create a new dataframe for data without the columns that have "Poverty Status" in the title, which is 83 columns
withoutps = withoutsex[[col for col in withoutsex.columns if 'POVERTY STATUS' not in col]].copy()
withoutps.head()
#Find all columns without "Disability Status" in column title
#[col for col in withoutps.columns if "DISABILITY STATUS" not in col]
#Create a new dataframe for data without the columns that have "Disability Status" in the title, which is 80 columns
withoutds = withoutps[[col for col in withoutps.columns if 'DISABILITY STATUS' not in col]].copy()
withoutds.head()
#Find all columns without "Educational Attainment" in column title
#[col for col in withoutds.columns if "EDUCATIONAL ATTAINMENT" not in col]
#Create a new dataframe for data without the columns that have "Educational Attainment" in the title, which is 65 columns
withoutea = withoutds[[col for col in withoutds.columns if 'EDUCATIONAL ATTAINMENT' not in col]].copy()
withoutea.head()
#Find all columns without "Age" in column title
#[col for col in withoutea.columns if "AGE" not in col]
#Create a new dataframe for data without the columns that have "Age" in the title, which is 35 columns
withoutage = withoutea[[col for col in withoutea.columns if 'AGE' not in col]].copy()
withoutage
#Format to rename School District Name
#def cleandistrict(NAME):
#return NAME.replace(", California","")
#Apply formatting condition to School District Name
#withoutage['NAME'].apply(cleandistrict)
#Create new School District name column with formatting titled District
#withoutage['District']= withoutage['NAME'].apply(cleandistrict)
#Find length of Geographt column contents, which is 16
#withoutage['Geography'].apply(len).unique()
#Pull a geography
#geo = "9700000US0622710"
#Find NCESDist ID
#geo.split("US")[1]
#Format to seperate Census Geography code (the state and district ID combined),
#def splitGeo(geo):
#return geo.split("US")[1]
#Apply formatting condition to Geography
#withoutage['Geography'].apply(splitGeo)
#Create new Geography name column with formatting titled NCESDist
#withoutage['NCESDist']= withoutage['Geography'].apply(splitGeo)
#Create new column with combined state and district ID
withoutage['NCESDist'] = withoutage['STATE'].astype(str) + withoutage['SCHOOL DISTRICT (UNIFIED)']
#Add Year column
withoutage['year']= "2016"
withoutage.head()
withoutage.to_csv("/Users/nataligracia/git/ca-school-enrollment-trend/Unified2016.csv")
#Rearrange columns in list
#['NCESDist','District'] + list(without_annotation_df.columns[2:])
#Rearrange columns in dataframe
#final = without_annotation_df[['NCESDist','District','Geography','NAME'] + list(without_annotation_df.columns[2:])]
#final
```
| github_jupyter |
#### New to Plotly?
Plotly's Python library is free and open source! [Get started](https://plotly.com/python/getting-started/) by downloading the client and [reading the primer](https://plotly.com/python/getting-started/).
<br>You can set up Plotly to work in [online](https://plotly.com/python/getting-started/#initialization-for-online-plotting) or [offline](https://plotly.com/python/getting-started/#initialization-for-offline-plotting) mode, or in [jupyter notebooks](https://plotly.com/python/getting-started/#start-plotting-online).
<br>We also have a quick-reference [cheatsheet](https://images.plot.ly/plotly-documentation/images/python_cheat_sheet.pdf) (new!) to help you get started!
#### Version Check
Plotly's python package is updated frequently. Run `pip install plotly --upgrade` to use the latest version.
```
import plotly
plotly.__version__
from plotly.offline import iplot, init_notebook_mode
import plotly.graph_objs as go
import pandas as pd
import numpy as np
import ipywidgets as widgets
```
We'll configure the notebook for use in [offline](https://plotly.com/python/getting-started/#initialization-for-offline-plotting) mode
```
init_notebook_mode(connected=True)
```
#### Parallel Categories Diagram
The parallel categories diagram is a visualization of multi-dimensional categorical data sets. Each variable in the data set is represented by a column of rectangles, where each rectangle corresponds to a discrete value taken on by that variable. The relative heights of the rectangles reflect the relative frequency of occurrence of the corresponding value.
Combinations of category rectangles across dimensions are connected by ribbons, where the height of the ribbon corresponds to the relative frequency of occurrence of the combination of categories in the data set.
#### Basic Parallel Categories Diagram
In this first example, we visualize the hair color, eye color, and sex of a sample of 8 people. Hovering over a category rectangle displays a tooltip with the number of people with that single trait. Hovering over a ribbon in the diagram displays a tooltip with the number of people with a particular combination of the three traits connected by the ribbon.
The dimension labels can be dragged horizontally to reorder the dimensions and the category rectangles can be dragged vertically to reorder the categories within a dimension.
```
parcats = go.Parcats(
dimensions=[
{'label': 'Hair',
'values': ['Black', 'Black', 'Black', 'Brown',
'Brown', 'Brown', 'Red', 'Brown']},
{'label': 'Eye',
'values': ['Brown', 'Brown', 'Brown', 'Brown',
'Brown', 'Blue', 'Blue', 'Blue']},
{'label': 'Sex',
'values': ['Female', 'Female', 'Female', 'Male',
'Female', 'Male', 'Male', 'Male']}]
)
iplot([parcats])
```
#### Basic Parallel Categories Diagram with Counts
If the frequency of occurrence for each combination of attributes is known in advance, this can be specified using the `counts` property
```
parcats = go.Parcats(
dimensions=[
{'label': 'Hair',
'values': ['Black', 'Brown', 'Brown', 'Brown', 'Red']},
{'label': 'Eye',
'values': ['Brown', 'Brown', 'Brown', 'Blue', 'Blue']},
{'label': 'Sex',
'values': ['Female', 'Male', 'Female', 'Male', 'Male']}],
counts=[6, 10, 40, 23, 7]
)
iplot([parcats])
```
#### Mutli-Color Parallel Categories Diagram
The color of the ribbons can be specified with the `line.color` property. Similar to other trace types, this property may be set to an array of numbers, which are then mapped to colors according to the the colorscale specified in the `line.colorscale` property.
Here is an example of visualizing the survival rate of passengers in the titanic dataset, where the ribbons are colored based on survival outcome.
By setting the `hoveron` property to `'color'` and the `hoverinfo` property to `'count+probability'` the tooltips now display count and probability information for each color (survival outcome) per category.
By setting the `arrangement` property to `'freeform'` it is now possible to drag categories horizontally to reorder dimensions as well as vertically to reorder categories within the dimension.
```
titanic_df = pd.read_csv(
"https://raw.githubusercontent.com/plotly/datasets/master/titanic.csv")
# Create dimensions
class_dim = go.parcats.Dimension(
values=titanic_df.Pclass,
categoryorder='category ascending',
label="Class"
)
gender_dim = go.parcats.Dimension(
values=titanic_df.Sex,
label="Gender"
)
survival_dim = go.parcats.Dimension(
values=titanic_df.Survived,
label="Outcome",
categoryarray=[0, 1],
ticktext=['perished', 'survived'],
)
# Create parcats trace
color = titanic_df.Survived;
colorscale = [[0, 'lightsteelblue'], [1, 'mediumseagreen']];
data = [
go.Parcats(
dimensions=[class_dim, gender_dim, survival_dim],
line={'color': color,
'colorscale': colorscale},
hoveron='color',
hoverinfo='count+probability',
labelfont={'size': 18, 'family': 'Times'},
tickfont={'size': 16, 'family': 'Times'},
arrangement='freeform'
)
]
# Display figure
iplot(data)
```
#### Parallel Categories Linked Brushing
This example demonstrates how the `on_selection` and `on_click` callbacks can be used to implement linked brushing between 3 categorical dimensions displayed with a `parcats` trace and 2 continuous dimensions displayed with a `scatter` trace.
This example also sets the `line.shape` property to `hspline` to cause the ribbons to curve between categories.
**Note:** In order for the callback functions to be executed the figure must be a `FigureWidget`, and the figure should display itself. In particular the `plot` and `iplot` functions should not be used.
```
cars_df = pd.read_csv(
'https://raw.githubusercontent.com/plotly/datasets/master/imports-85.csv')
# Build parcats dimensions
categorical_dimensions = [
'body-style',
'drive-wheels',
'fuel-type'
];
dimensions = [
dict(values=cars_df[label], label=label)
for label in categorical_dimensions
]
# Build colorscale
color = np.zeros(len(cars_df), dtype='uint8')
colorscale = [[0, 'gray'], [1, 'firebrick']]
# Build figure as FigureWidget
fig = go.FigureWidget(
data=[
go.Scatter(
x=cars_df.horsepower,
y=cars_df['highway-mpg'],
marker={'color': 'gray'},
mode='markers',
selected={'marker': {'color': 'firebrick'}},
unselected={'marker': {'opacity': 0.3}}),
go.Parcats(
domain={'y': [0, 0.4]},
dimensions=dimensions,
line={
'colorscale': colorscale,
'cmin': 0,
'cmax': 1,
'color': color,
'shape': 'hspline'})
],
layout=go.Layout(
height=800,
xaxis={'title': 'Horsepower'},
yaxis={'title': 'MPG',
'domain': [0.6, 1]},
dragmode='lasso',
hovermode='closest')
)
# Update color callback
def update_color(trace, points, state):
# Update scatter selection
fig.data[0].selectedpoints = points.point_inds
# Update parcats colors
new_color = np.zeros(len(cars_df), dtype='uint8')
new_color[points.point_inds] = 1
fig.data[1].line.color = new_color
# Register callback on scatter selection...
fig.data[0].on_selection(update_color)
# and parcats click
fig.data[1].on_click(update_color)
# Display figure
fig
```
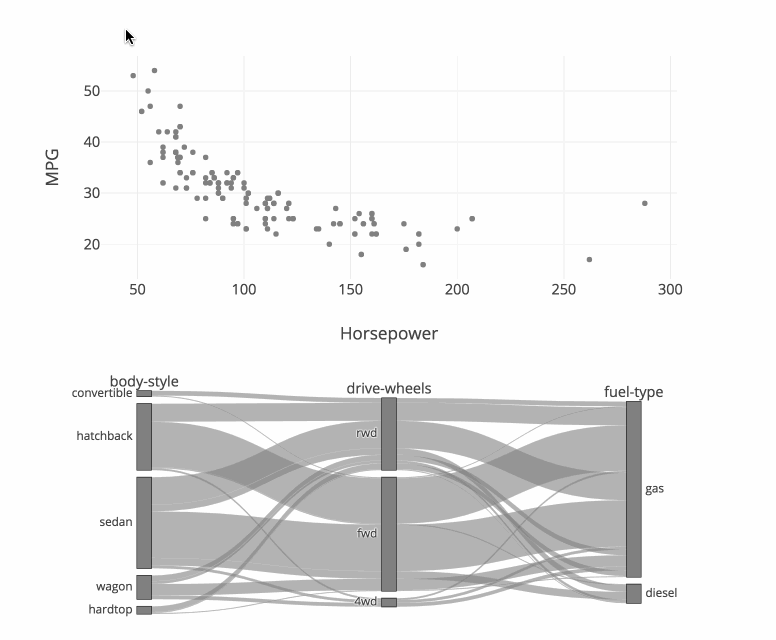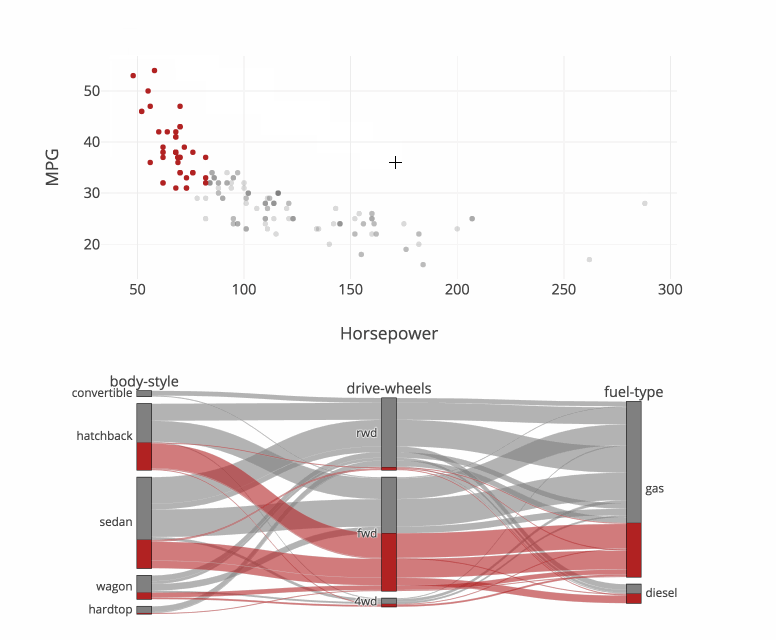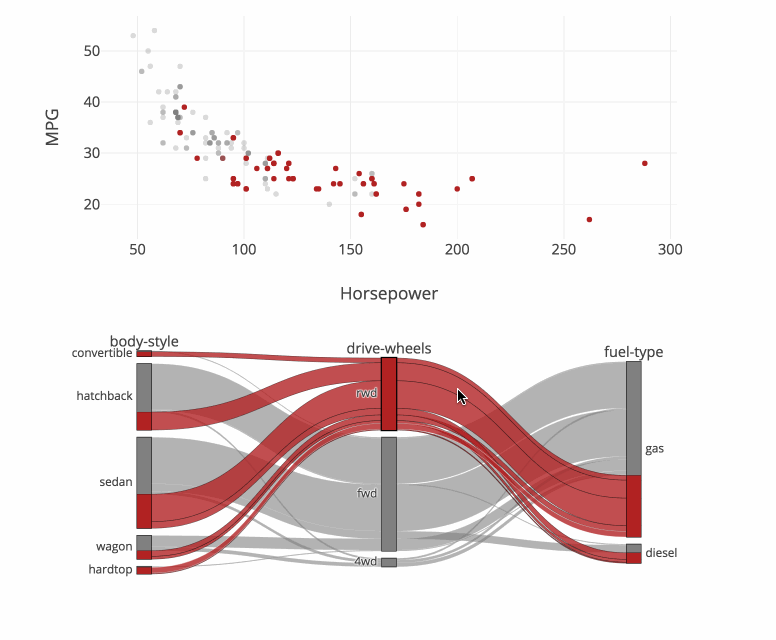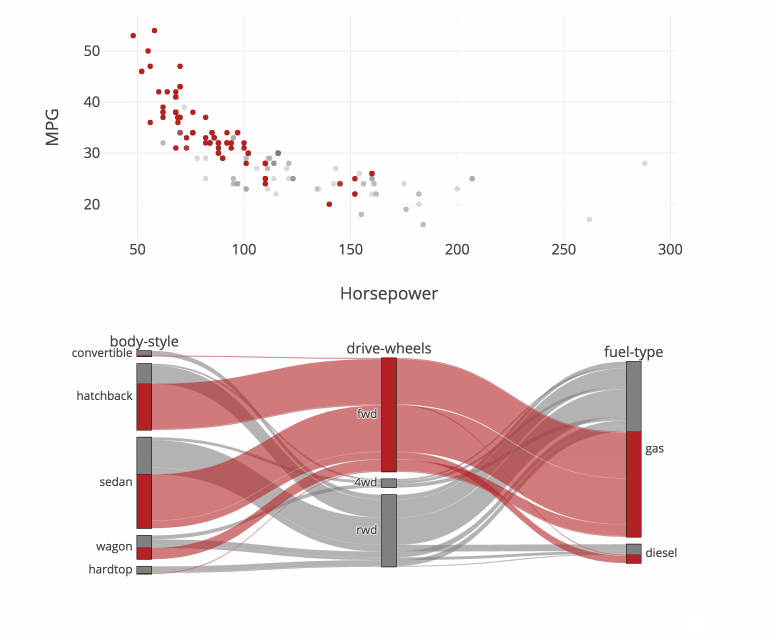
#### Parallel Categories with Multi-Color Linked Brushing
This example extends the previous example to support brushing with multiple colors. The toggle buttons above may be used to select the active color, and this color will be applied when points are selected in the `scatter` trace and when categories or ribbons are clicked in the `parcats` trace.
```
cars_df = pd.read_csv(
'https://raw.githubusercontent.com/plotly/datasets/master/imports-85.csv')
# Build parcats dimensions
categorical_dimensions = [
'body-style',
'drive-wheels',
'fuel-type'
];
dimensions = [
dict(values=cars_df[label], label=label)
for label in categorical_dimensions
]
# Build colorscale
color = np.zeros(len(cars_df), dtype='uint8')
colorscale = [[0, 'gray'], [0.33, 'gray'],
[0.33, 'firebrick'], [0.66, 'firebrick'],
[0.66, 'blue'], [1.0, 'blue']];
cmin = -0.5
cmax = 2.5
# Build figure as FigureWidget
fig = go.FigureWidget(
data=[
go.Scatter(
x=cars_df.horsepower,
y=cars_df['highway-mpg'],
marker={'color': color,
'cmin': cmin,
'cmax': cmax,
'colorscale': colorscale,
'showscale': True,
'colorbar': {'tickvals': [0, 1, 2],
'ticktext': ['None', 'Red', 'Blue']}
},
mode='markers'),
go.Parcats(
domain={'y': [0, 0.4]},
dimensions=dimensions,
line={
'colorscale': colorscale,
'cmin': cmin,
'cmax': cmax,
'color': color,
'shape': 'hspline'})
],
layout=go.Layout(
height=800,
xaxis={'title': 'Horsepower'},
yaxis={'title': 'MPG',
'domain': [0.6, 1]},
dragmode='lasso',
hovermode='closest')
)
# Build color selection widget
color_toggle = widgets.ToggleButtons(
options=['None', 'Red', 'Blue'],
index=1,
description='Brush Color:',
disabled=False,
)
# Update color callback
def update_color(trace, points, state):
# Compute new color array
new_color = np.array(fig.data[0].marker.color)
new_color[points.point_inds] = color_toggle.index
with fig.batch_update():
# Update scatter color
fig.data[0].marker.color = new_color
# Update parcats colors
fig.data[1].line.color = new_color
# Register callback on scatter selection...
fig.data[0].on_selection(update_color)
# and parcats click
fig.data[1].on_click(update_color)
# Display figure
widgets.VBox([color_toggle, fig])
```
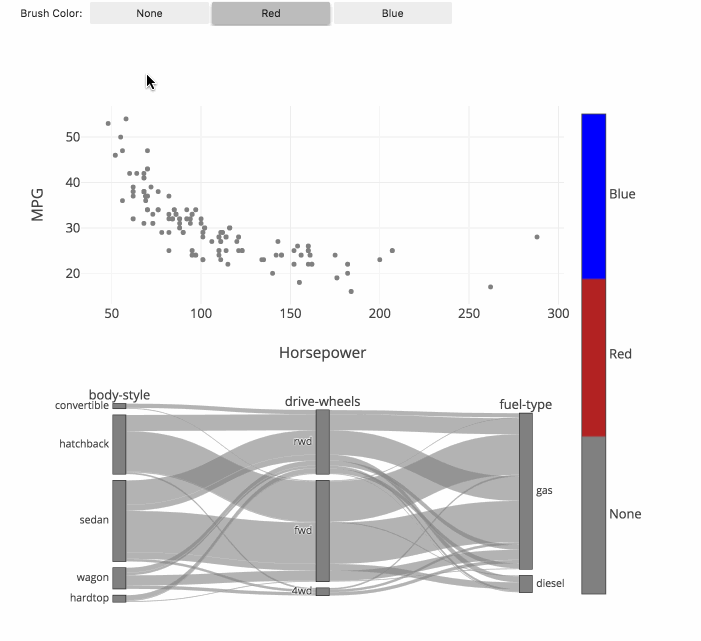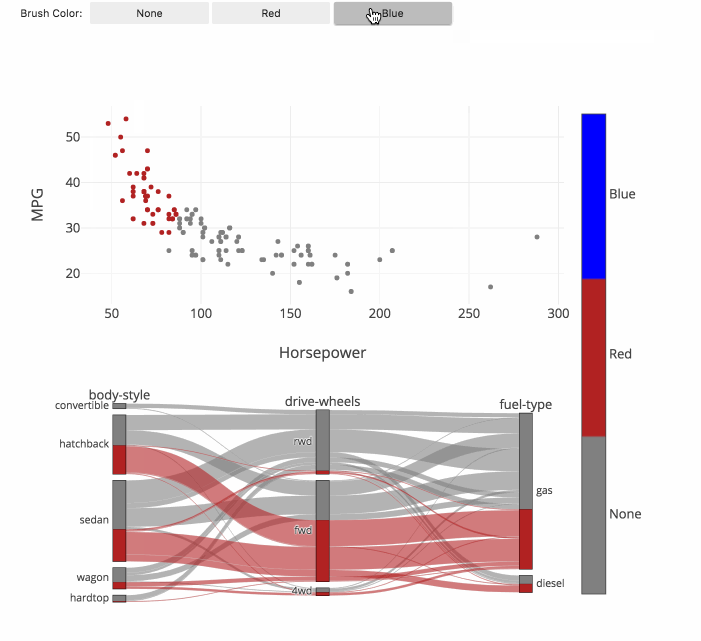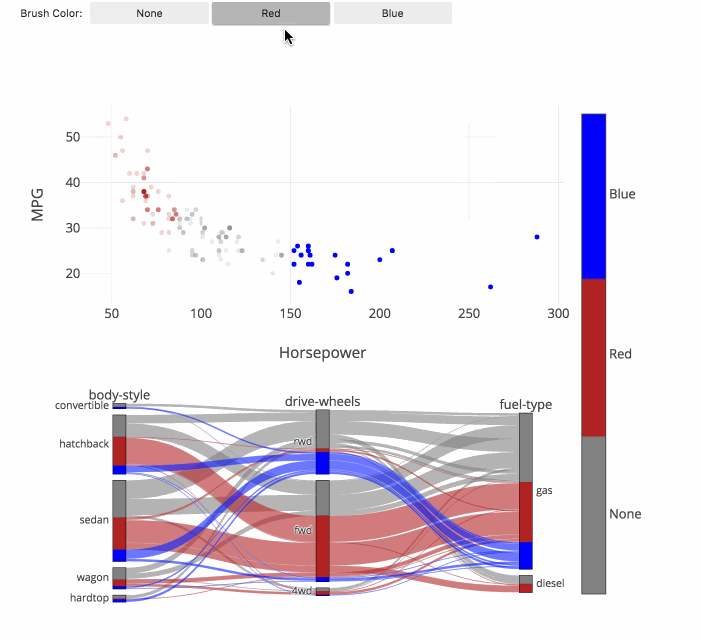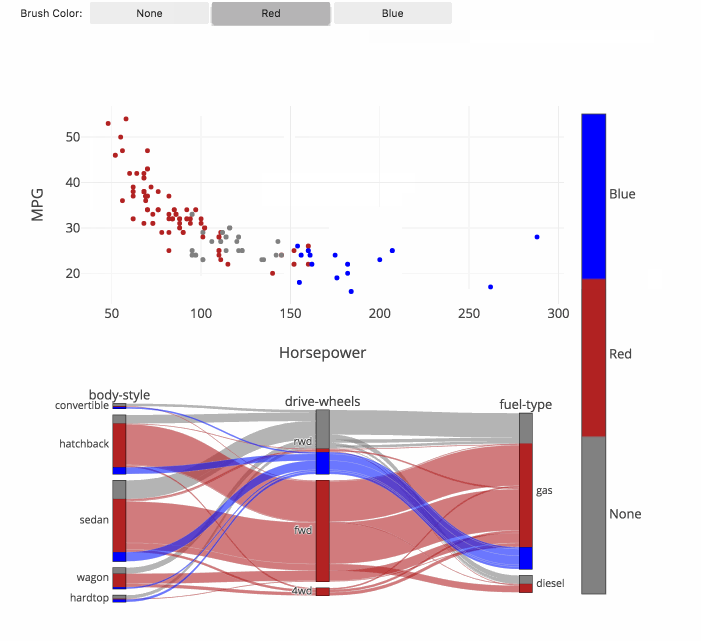
#### Reference
See https://plotly.com/python/reference/#parcats for more information and chart attribute options!
```
from IPython.display import display, HTML
display(HTML('<link href="//fonts.googleapis.com/css?family=Open+Sans:600,400,300,200|Inconsolata|Ubuntu+Mono:400,700" rel="stylesheet" type="text/css" />'))
display(HTML('<link rel="stylesheet" type="text/css" href="http://help.plot.ly/documentation/all_static/css/ipython-notebook-custom.css">'))
! pip install git+https://github.com/plotly/publisher.git --upgrade
import publisher
publisher.publish(
'parcats.ipynb', 'python/parallel-categories-diagram/', 'Parallel Categories Diagram',
'How to make parallel categories diagrams in Python with Plotly.',
title = 'Python Parallel Categories | Plotly',
has_thumbnail='true', thumbnail='thumbnail/parcats.jpg',
language='python',
display_as='statistical', order=10.3,
uses_plotly_offline=True,
ipynb= '~notebook_demo/258')
```
| github_jupyter |
<h1><font color='blue'> 8E and 8F: Finding the Probability P(Y==1|X)</font></h1>
<h2><font color='Geen'> 8E: Implementing Decision Function of SVM RBF Kernel</font></h2>
<font face=' Comic Sans MS' size=3>After we train a kernel SVM model, we will be getting support vectors and their corresponsing coefficients $\alpha_{i}$
Check the documentation for better understanding of these attributes:
https://scikit-learn.org/stable/modules/generated/sklearn.svm.SVC.html
<img src='https://i.imgur.com/K11msU4.png' width=500>
As a part of this assignment you will be implementing the ```decision_function()``` of kernel SVM, here decision_function() means based on the value return by ```decision_function()``` model will classify the data point either as positive or negative
Ex 1: In logistic regression After traning the models with the optimal weights $w$ we get, we will find the value $\frac{1}{1+\exp(-(wx+b))}$, if this value comes out to be < 0.5 we will mark it as negative class, else its positive class
Ex 2: In Linear SVM After traning the models with the optimal weights $w$ we get, we will find the value of $sign(wx+b)$, if this value comes out to be -ve we will mark it as negative class, else its positive class.
Similarly in Kernel SVM After traning the models with the coefficients $\alpha_{i}$ we get, we will find the value of
$sign(\sum_{i=1}^{n}(y_{i}\alpha_{i}K(x_{i},x_{q})) + intercept)$, here $K(x_{i},x_{q})$ is the RBF kernel. If this value comes out to be -ve we will mark $x_{q}$ as negative class, else its positive class.
RBF kernel is defined as: $K(x_{i},x_{q})$ = $exp(-\gamma ||x_{i} - x_{q}||^2)$
For better understanding check this link: https://scikit-learn.org/stable/modules/svm.html#svm-mathematical-formulation
</font>
## Task E
> 1. Split the data into $X_{train}$(60), $X_{cv}$(20), $X_{test}$(20)
> 2. Train $SVC(gamma=0.001, C=100.)$ on the ($X_{train}$, $y_{train}$)
> 3. Get the decision boundry values $f_{cv}$ on the $X_{cv}$ data i.e. ` `$f_{cv}$ ```= decision_function(```$X_{cv}$```)``` <font color='red'>you need to implement this decision_function()</font>
```
import numpy as np
import pandas as pd
from sklearn.datasets import make_classification
from sklearn.model_selection import GridSearchCV
import matplotlib.pyplot as plt
from sklearn.svm import SVC
from sklearn.model_selection import train_test_split
import math
X, y = make_classification(n_samples=5000, n_features=5, n_redundant=2,
n_classes=2, weights=[0.7], class_sep=0.7, random_state=15)
xtrain, xtest, ytrain, ytest = train_test_split(X, y, test_size=0.3, random_state=0)
xcv, xtest,ycv, ytest = train_test_split(xtest, ytest, test_size=0.3, random_state=0)
print(xtrain.shape, ytrain.shape, xtest.shape, ytest.shape)
print(xtest.shape, ytest.shape, xcv.shape, ycv.shape)
clf = SVC(random_state=0, decision_function_shape='ovo')
clf = GridSearchCV(clf, {'C': [0.001, 0.01, 0.1, 1, 10, 100], 'gamma' : [0.001,0.01, 0.1, 1, 10, 100]}, n_jobs=-1, cv=5)
clf = clf.fit(xtrain, ytrain) # set the best parameters
clf.best_estimator_, clf.best_score_
```
### Pseudo code
clf = SVC(gamma=0.001, C=100.)<br>
clf.fit(Xtrain, ytrain)
<font color='green'>def</font> <font color='blue'>decision_function</font>(Xcv, ...): #use appropriate parameters <br>
<font color='green'>for</font> a data point $x_q$ <font color='green'>in</font> Xcv: <br>
<font color='grey'>#write code to implement $(\sum_{i=1}^{\text{all the support vectors}}(y_{i}\alpha_{i}K(x_{i},x_{q})) + intercept)$, here the values $y_i$, $\alpha_{i}$, and $intercept$ can be obtained from the trained model</font><br>
<font color='green'>return</font> <font color='grey'><i># the decision_function output for all the data points in the Xcv</i></font>
fcv = decision_function(Xcv, ...) <i># based on your requirement you can pass any other parameters </i>
<b>Note</b>: Make sure the values you get as fcv, should be equal to outputs of clf.decision_function(Xcv)
```
clf = SVC(random_state=0, gamma=1, C=100, decision_function_shape='ovo')
clf.fit(xtrain, ytrain)
pred = clf.predict(xcv)
clf_dec = clf.decision_function(xcv)
def decision_function(clf, data):
add_intercept = []
for x_q in data:
add_intercept.append(np.sum(clf.dual_coef_ * np.exp(-clf._gamma*np.sum((clf.support_vectors_ - x_q)**2, axis=1))) + clf.intercept_[0])
return add_intercept
fcv = decision_function(clf, xcv)
print(fcv[:5], '\n', clf_dec[:5])
```
<h2><font color='Geen'> 8F: Implementing Platt Scaling to find P(Y==1|X)</font></h2>
Check this <a href='https://drive.google.com/open?id=133odBinMOIVb_rh_GQxxsyMRyW-Zts7a'>PDF</a>
<img src='https://i.imgur.com/CAMnVnh.png'>
```
unique, frequency = np.unique(ytrain, return_counts = True)
count = np.asarray((unique, frequency ))
print(count)
neg, pos = frequency[0], frequency[1]
def target_calib(x):
cal_target = []
for i in x:
if i == 1:
cal_target.append((pos + 1)/(pos + 2))
elif i == 0:
cal_target.append(1 / (neg + 2))
return cal_target
claibrated_target = target_calib(pred.tolist())
```
## TASK F
> 4. Apply SGD algorithm with ($f_{cv}$, $y_{cv}$) and find the weight $W$ intercept $b$ ```Note: here our data is of one dimensional so we will have a one dimensional weight vector i.e W.shape (1,)```
> Note1: Don't forget to change the values of $y_{cv}$ as mentioned in the above image. you will calculate y+, y- based on data points in train data
> Note2: the Sklearn's SGD algorithm doesn't support the real valued outputs, you need to use the code that was done in the `'Logistic Regression with SGD and L2'` Assignment after modifying loss function, and use same parameters that used in that assignment.
<img src='https://i.imgur.com/zKYE9Oc.png'>
if Y[i] is 1, it will be replaced with y+ value else it will replaced with y- value
> 5. For a given data point from $X_{test}$, $P(Y=1|X) = \frac{1}{1+exp(-(W*f_{test}+ b))}$ where ` `$f_{test}$ ```= decision_function(```$X_{test}$```)```, W and b will be learned as metioned in the above step
```
def initialize_weights(dim):
w = np.zeros_like((dim))
b = np.zeros_like((1))
print("Weights-Initialized : ", w.shape)
return w,b
def sigmoid(z):
sigmoid = 1/(1+math.exp(-z))
return sigmoid
def logloss(W, b, X, Y):
N = len(X)
loss=[]
for i in range(N):
z = np.dot(X[i],W) + b
pred = sigmoid(z)
if pred < 0.5:
l = (1-Y[i])*np.log10(1-pred)
loss.append(l)
else:
l = Y[i]*np.log10(pred)
loss.append(l)
loss = (-1 * 1/len(loss) * sum(loss))
return loss
def gradient_dw(x,y,w,b,alpha,N):
dw =x*(y-sigmoid(np.dot(w,x)+b)) - alpha/N * w
return dw
def gradient_db(x,y,w,b):
db =(y-sigmoid(np.dot(w,x)+b))
return db
def pred(w,b, X):
N = len(X.tolist())
predict = []
for i in range(N):
z = np.dot(X[i],w) + b
predict.append(sigmoid(z))
return np.array(predict)
def train(Y_calibrated,fcv,epochs,alpha,eta0):
''' In this function, we will implement logistic regression'''
scale_down_factor = 0.0001
epoch = 1
w, b = initialize_weights(1)
wl = []
bl = []
Lw=np.zeros_like(1)
Lb=0
loss = 0
prev = 0
train_loss = []
test_loss = []
while epoch <= epochs:
y_train_pred = []
y_test_pred = []
np.random.RandomState(seed=2)
for m in range(len(Y_calibrated)):
i = np.random.choice(len(Y_calibrated))
z = np.dot(Y_calibrated[i],w) + b
Lw = gradient_dw(Y_calibrated[i],fcv[i],w,b,alpha,len(Y_calibrated))
Lb = gradient_db(Y_calibrated[i],fcv[i],w,b)
w=(1-(alpha * scale_down_factor/epochs))*w+alpha*Lw
b=b+alpha*Lb
train_loss.append(round(logloss(w,b,Y_calibrated, fcv), 3))
if train_loss[-1] == prev:
break;
else:
prev = train_loss[-1]
print("Epoch: %d, train_Loss: %.3f" %(epoch, train_loss[-1]))
epoch+=1
%matplotlib inline
import matplotlib.pyplot as plt
plt.plot(train_loss, label='train_log_loss')
plt.grid()
plt.legend()
plt.title('Log loss vs epoch')
plt.xlabel('Iterations')
plt.ylabel('log loss')
plt.show()
return w,b
alpha=0.0001
eta0=0.0001
N=len(xcv)
epochs=50
w,b = train(claibrated_target,fcv,epochs,alpha,eta0)
w, b
f_test = decision_function(clf, xtest)
def calibrated_test(ftest, weight, bias):
test_prediction = []
for i in ftest:
z = np.dot(i,weight) + bias
test_prediction.append(sigmoid(z))
return np.array(test_prediction)
test_pred = calibrated_test(f_test, w, b)
print(test_pred[:5])
```
__Note: in the above algorithm, the steps 2, 4 might need hyper parameter tuning, To reduce the complexity of the assignment we are excluding the hyerparameter tuning part, but intrested students can try that__
If any one wants to try other calibration algorithm istonic regression also please check these tutorials
1. http://fa.bianp.net/blog/tag/scikit-learn.html#fn:1
2. https://drive.google.com/open?id=1MzmA7QaP58RDzocB0RBmRiWfl7Co_VJ7
3. https://drive.google.com/open?id=133odBinMOIVb_rh_GQxxsyMRyW-Zts7a
4. https://stat.fandom.com/wiki/Isotonic_regression#Pool_Adjacent_Violators_Algorithm
| github_jupyter |
# Mnist classification pipeline using Sagemaker
The `mnist-classification-pipeline.py` sample runs a pipeline to train a classficiation model using Kmeans with MNIST dataset on Sagemaker.
We will have all required steps here and for other details like how to get source data, please check [documentation](https://github.com/kubeflow/pipelines/tree/master/samples/contrib/aws-samples/mnist-kmeans-sagemaker).
This sample is based on the [Train a Model with a Built-in Algorithm and Deploy it](https://docs.aws.amazon.com/sagemaker/latest/dg/ex1.html).
The sample trains and deploy a model based on the [MNIST dataset](http://www.deeplearning.net/tutorial/gettingstarted.html).
## Prerequisite
1. Create an S3 bucket to store pipeline data
> Note: Be sure to change the HASH variable to random hash and change AWS_REGION before running next cell
> Note: you use us-east-1, please use command `!aws s3 mb s3://$S3_BUCKET --region $AWS_REGION --endpoint-url https://s3.us-east-1.amazonaws.com`
```
import random, string
HASH = ''.join([random.choice(string.ascii_lowercase) for n in range(16)] + [random.choice(string.digits) for n in range(16)])
AWS_REGION = 'us-east-2'
S3_BUCKET = '{}-kubeflow-pipeline-data'.format(HASH)
!aws s3 mb s3://$S3_BUCKET --region $AWS_REGION
!pip install sagemaker
```
2. Copy dataset
> Download and upload `data` and `valid_data.csv` into your S3 bucket.
```
import pickle, gzip, numpy, urllib.request, json
from urllib.parse import urlparse
# Load the dataset
urllib.request.urlretrieve("http://deeplearning.net/data/mnist/mnist.pkl.gz", "mnist.pkl.gz")
with gzip.open('mnist.pkl.gz', 'rb') as f:
train_set, valid_set, test_set = pickle.load(f, encoding='latin1')
# Upload dataset to S3
from sagemaker.amazon.common import write_numpy_to_dense_tensor
import io
import boto3
###################################################################
# This is the only thing that you need to change to run this code
# Give the name of your S3 bucket
bucket = S3_BUCKET
# If you are gonna use the default values of the pipeline then
# give a bucket name which is in us-west-2 region
###################################################################
train_data_key = 'mnist_kmeans_example/train_data'
test_data_key = 'mnist_kmeans_example/test_data'
train_data_location = 's3://{}/{}'.format(bucket, train_data_key)
test_data_location = 's3://{}/{}'.format(bucket, test_data_key)
print('training data will be uploaded to: {}'.format(train_data_location))
print('training data will be uploaded to: {}'.format(test_data_location))
# Convert the training data into the format required by the SageMaker KMeans algorithm
buf = io.BytesIO()
write_numpy_to_dense_tensor(buf, train_set[0], train_set[1])
buf.seek(0)
boto3.resource('s3').Bucket(bucket).Object(train_data_key).upload_fileobj(buf)
# Convert the test data into the format required by the SageMaker KMeans algorithm
write_numpy_to_dense_tensor(buf, test_set[0], test_set[1])
buf.seek(0)
boto3.resource('s3').Bucket(bucket).Object(test_data_key).upload_fileobj(buf)
# Convert the valid data into the format required by the SageMaker KMeans algorithm
numpy.savetxt('valid-data.csv', valid_set[0], delimiter=',', fmt='%g')
s3_client = boto3.client('s3')
input_key = "{}/valid_data.csv".format("mnist_kmeans_example/input")
s3_client.upload_file('valid-data.csv', bucket, input_key)
```
4. Install Kubeflow Pipelines SDK
> You can skip this step if its already installed. You can validate if you have SDK installed by running `!pip show kfp`. The notebook has been tested for kfp v0.1.29 release
```
!pip install https://storage.googleapis.com/ml-pipeline/release/0.1.29/kfp.tar.gz --upgrade
!pip show kfp
```
## Build pipeline
1. Run the following command to load Kubeflow Pipelines SDK
```
import kfp
from kfp import components
from kfp import dsl
from kfp.aws import use_aws_secret
```
2. Load reusable sagemaker components.
```
sagemaker_train_op = components.load_component_from_url('https://raw.githubusercontent.com/kubeflow/pipelines/master/components/aws/sagemaker/train/component.yaml')
sagemaker_model_op = components.load_component_from_url('https://raw.githubusercontent.com/kubeflow/pipelines/master/components/aws/sagemaker/model/component.yaml')
sagemaker_deploy_op = components.load_component_from_url('https://raw.githubusercontent.com/kubeflow/pipelines/master/components/aws/sagemaker/deploy/component.yaml')
sagemaker_batch_transform_op = components.load_component_from_url('https://raw.githubusercontent.com/kubeflow/pipelines/master/components/aws/sagemaker/batch_transform/component.yaml')
sagemaker_hpo_op = components.load_component_from_url('https://raw.githubusercontent.com/kubeflow/pipelines/master/components/aws/sagemaker/hyperparameter_tuning/component.yaml')
```
3. Create pipeline.
We will create Hyperparamater tuning job following by a training job first. Once training job is done, it will persist trained model to S3.
Then a job will be kicked off to create a `Model` manifest in Sagemaker.
With this model, batch transformation job can use it to predict on other datasets, prediction service can create an endpoint using it.
> Note: remember to use pass your **role_arn** to successfully run the job.
> Note: If you use a different region, please replace `us-west-2` with your region.
> Note: ECR Images for k-means algorithm
|Region| ECR Image|
|------|----------|
|us-west-1|632365934929.dkr.ecr.us-west-1.amazonaws.com|
|us-west-2|174872318107.dkr.ecr.us-west-2.amazonaws.com|
|us-east-1|382416733822.dkr.ecr.us-east-1.amazonaws.com|
|us-east-2|404615174143.dkr.ecr.us-east-2.amazonaws.com|
|us-gov-west-1|226302683700.dkr.ecr.us-gov-west-1.amazonaws.com|
|ap-east-1|286214385809.dkr.ecr.ap-east-1.amazonaws.com|
|ap-northeast-1|351501993468.dkr.ecr.ap-northeast-1.amazonaws.com|
|ap-northeast-2|835164637446.dkr.ecr.ap-northeast-2.amazonaws.com|
|ap-south-1|991648021394.dkr.ecr.ap-south-1.amazonaws.com|
|ap-southeast-1|475088953585.dkr.ecr.ap-southeast-1.amazonaws.com|
|ap-southeast-2|712309505854.dkr.ecr.ap-southeast-2.amazonaws.com|
|ca-central-1|469771592824.dkr.ecr.ca-central-1.amazonaws.com|
|eu-central-1|664544806723.dkr.ecr.eu-central-1.amazonaws.com|
|eu-north-1|669576153137.dkr.ecr.eu-north-1.amazonaws.com|
|eu-west-1|438346466558.dkr.ecr.eu-west-1.amazonaws.com|
|eu-west-2|644912444149.dkr.ecr.eu-west-2.amazonaws.com|
|eu-west-3|749696950732.dkr.ecr.eu-west-3.amazonaws.com|
|me-south-1|249704162688.dkr.ecr.me-south-1.amazonaws.com|
|sa-east-1|855470959533.dkr.ecr.sa-east-1.amazonaws.com|
```
# Configure your s3 bucket.
S3_BUCKET = '{}-kubeflow-pipeline-data'.format(HASH)
S3_PIPELINE_PATH='s3://{}/mnist_kmeans_example'.format(S3_BUCKET)
# Configure your Sagemaker execution role.
SAGEMAKER_ROLE_ARN='<Your_SageMaker_Role>'
@dsl.pipeline(
name='MNIST Classification pipeline',
description='MNIST Classification using KMEANS in SageMaker'
)
def mnist_classification(region='us-east-2',
image='404615174143.dkr.ecr.us-east-2.amazonaws.com/kmeans:1',
training_input_mode='File',
hpo_strategy='Bayesian',
hpo_metric_name='test:msd',
hpo_metric_type='Minimize',
hpo_early_stopping_type='Off',
hpo_static_parameters='{"k": "10", "feature_dim": "784"}',
hpo_integer_parameters='[{"Name": "mini_batch_size", "MinValue": "500", "MaxValue": "600"}, {"Name": "extra_center_factor", "MinValue": "10", "MaxValue": "20"}]',
hpo_continuous_parameters='[]',
hpo_categorical_parameters='[{"Name": "init_method", "Values": ["random", "kmeans++"]}]',
hpo_channels='[{"ChannelName": "train", \
"DataSource": { \
"S3DataSource": { \
"S3Uri": "' + S3_PIPELINE_PATH + '/train_data", \
"S3DataType": "S3Prefix", \
"S3DataDistributionType": "FullyReplicated" \
} \
}, \
"ContentType": "", \
"CompressionType": "None", \
"RecordWrapperType": "None", \
"InputMode": "File"}, \
{"ChannelName": "test", \
"DataSource": { \
"S3DataSource": { \
"S3Uri": "' + S3_PIPELINE_PATH + '/test_data", \
"S3DataType": "S3Prefix", \
"S3DataDistributionType": "FullyReplicated" \
} \
}, \
"ContentType": "", \
"CompressionType": "None", \
"RecordWrapperType": "None", \
"InputMode": "File"}]',
hpo_spot_instance='False',
hpo_max_wait_time='3600',
hpo_checkpoint_config='{}',
output_location=S3_PIPELINE_PATH + '/output',
output_encryption_key='',
instance_type='ml.p3.2xlarge',
instance_count='1',
volume_size='50',
hpo_max_num_jobs='9',
hpo_max_parallel_jobs='2',
max_run_time='3600',
endpoint_url='',
network_isolation='True',
traffic_encryption='False',
train_channels='[{"ChannelName": "train", \
"DataSource": { \
"S3DataSource": { \
"S3Uri": "' + S3_PIPELINE_PATH + '/train_data", \
"S3DataType": "S3Prefix", \
"S3DataDistributionType": "FullyReplicated" \
} \
}, \
"ContentType": "", \
"CompressionType": "None", \
"RecordWrapperType": "None", \
"InputMode": "File"}]',
train_spot_instance='False',
train_max_wait_time='3600',
train_checkpoint_config='{}',
batch_transform_instance_type='ml.m4.xlarge',
batch_transform_input=S3_PIPELINE_PATH + '/input',
batch_transform_data_type='S3Prefix',
batch_transform_content_type='text/csv',
batch_transform_compression_type='None',
batch_transform_ouput=S3_PIPELINE_PATH + '/output',
batch_transform_max_concurrent='4',
batch_transform_max_payload='6',
batch_strategy='MultiRecord',
batch_transform_split_type='Line',
role_arn=SAGEMAKER_ROLE_ARN
):
hpo = sagemaker_hpo_op(
region=region,
endpoint_url=endpoint_url,
image=image,
training_input_mode=training_input_mode,
strategy=hpo_strategy,
metric_name=hpo_metric_name,
metric_type=hpo_metric_type,
early_stopping_type=hpo_early_stopping_type,
static_parameters=hpo_static_parameters,
integer_parameters=hpo_integer_parameters,
continuous_parameters=hpo_continuous_parameters,
categorical_parameters=hpo_categorical_parameters,
channels=hpo_channels,
output_location=output_location,
output_encryption_key=output_encryption_key,
instance_type=instance_type,
instance_count=instance_count,
volume_size=volume_size,
max_num_jobs=hpo_max_num_jobs,
max_parallel_jobs=hpo_max_parallel_jobs,
max_run_time=max_run_time,
network_isolation=network_isolation,
traffic_encryption=traffic_encryption,
spot_instance=hpo_spot_instance,
max_wait_time=hpo_max_wait_time,
checkpoint_config=hpo_checkpoint_config,
role=role_arn,
)
#.apply(use_aws_secret('aws-secret', 'AWS_ACCESS_KEY_ID', 'AWS_SECRET_ACCESS_KEY'))
training = sagemaker_train_op(
region=region,
endpoint_url=endpoint_url,
image=image,
training_input_mode=training_input_mode,
hyperparameters=hpo.outputs['best_hyperparameters'],
channels=train_channels,
instance_type=instance_type,
instance_count=instance_count,
volume_size=volume_size,
max_run_time=max_run_time,
model_artifact_path=output_location,
output_encryption_key=output_encryption_key,
network_isolation=network_isolation,
traffic_encryption=traffic_encryption,
spot_instance=train_spot_instance,
max_wait_time=train_max_wait_time,
checkpoint_config=train_checkpoint_config,
role=role_arn,
)
#.apply(use_aws_secret('aws-secret', 'AWS_ACCESS_KEY_ID', 'AWS_SECRET_ACCESS_KEY'))
create_model = sagemaker_model_op(
region=region,
endpoint_url=endpoint_url,
model_name=training.outputs['job_name'],
image=training.outputs['training_image'],
model_artifact_url=training.outputs['model_artifact_url'],
network_isolation=network_isolation,
role=role_arn
)
#.apply(use_aws_secret('aws-secret', 'AWS_ACCESS_KEY_ID', 'AWS_SECRET_ACCESS_KEY'))
prediction = sagemaker_deploy_op(
region=region,
endpoint_url=endpoint_url,
model_name_1=create_model.output,
)
#.apply(use_aws_secret('aws-secret', 'AWS_ACCESS_KEY_ID', 'AWS_SECRET_ACCESS_KEY'))
batch_transform = sagemaker_batch_transform_op(
region=region,
endpoint_url=endpoint_url,
model_name=create_model.output,
instance_type=batch_transform_instance_type,
instance_count=instance_count,
max_concurrent=batch_transform_max_concurrent,
max_payload=batch_transform_max_payload,
batch_strategy=batch_strategy,
input_location=batch_transform_input,
data_type=batch_transform_data_type,
content_type=batch_transform_content_type,
split_type=batch_transform_split_type,
compression_type=batch_transform_compression_type,
output_location=batch_transform_ouput
)
#.apply(use_aws_secret('aws-secret', 'AWS_ACCESS_KEY_ID', 'AWS_SECRET_ACCESS_KEY'))
```
4. Compile your pipeline
```
kfp.compiler.Compiler().compile(mnist_classification, 'mnist-classification-pipeline.zip')
```
5. Deploy your pipeline
```
client = kfp.Client()
aws_experiment = client.create_experiment(name='aws')
my_run = client.run_pipeline(aws_experiment.id, 'mnist-classification-pipeline',
'mnist-classification-pipeline.zip')
```
## Prediction
Open Sagemaker console and find your endpoint name. Please check dataset section to get train_set.
Once your pipeline is done, you can find sagemaker endpoint name and replace `ENDPOINT_NAME` value with your newly created endpoint name.
> Note: make sure to attach `sagemaker:InvokeEndpoint` to the worker node nodegroup that is running this jupyter notebook.
```json
{
"Version": "2012-10-17",
"Statement": [
{
"Effect": "Allow",
"Action": [
"sagemaker:InvokeEndpoint"
],
"Resource": "*"
}
]
}
```
```
!pip install boto3 --user
```
## Find your Endpoint name in AWS Console
Open AWS console and enter Sagemaker service, find the endpoint name as the following picture shows.

```
import pickle, gzip, numpy, urllib.request, json
from urllib.parse import urlparse
import json
import io
import boto3
# Replace the endpoint name with yours.
ENDPOINT_NAME='Endpoint-20190916223205-Y635'
# Load the dataset
urllib.request.urlretrieve("http://deeplearning.net/data/mnist/mnist.pkl.gz", "mnist.pkl.gz")
with gzip.open('mnist.pkl.gz', 'rb') as f:
train_set, valid_set, test_set = pickle.load(f, encoding='latin1')
# Simple function to create a csv from our numpy array
def np2csv(arr):
csv = io.BytesIO()
numpy.savetxt(csv, arr, delimiter=',', fmt='%g')
return csv.getvalue().decode().rstrip()
runtime = boto3.Session(region_name='us-east-2').client('sagemaker-runtime')
payload = np2csv(train_set[0][30:31])
response = runtime.invoke_endpoint(EndpointName=ENDPOINT_NAME,
ContentType='text/csv',
Body=payload)
result = json.loads(response['Body'].read().decode())
print(result)
```
## Clean up
Go to Sagemaker console and delete `endpoint`, `model`.
### Clean up S3 bucket
Delete S3 bucket that was created for this exercise
```
!aws s3 rb s3://$S3_BUCKET --force
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.