text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
# **U-Net (2D)**
---
<font size = 4>U-Net is an encoder-decoder network architecture originally used for image segmentation, first published by [Ronneberger *et al.*](https://arxiv.org/abs/1505.04597). The first half of the U-Net architecture is a downsampling convolutional neural network which acts as a feature extra... | github_jupyter |
Some random sanity checks and scratchpads worth keeping around.
```
import jax.numpy as jnp
from scipy import signal
import numpy as np
import time
from jaxdsp import processor_graph
from jaxdsp.processors import fir_filter, iir_filter, clip, delay_line, biquad_lowpass, lowpass_feedback_comb_filter as lbcf, allpass_f... | github_jupyter |
```
%matplotlib inline
from cosmodc2.sdss_colors import load_umachine_processed_sdss_catalog
sdss = load_umachine_processed_sdss_catalog()
print(sdss.keys())
import os
from astropy.table import Table
# MDPL2-based mock
dirname = "/Users/aphearin/work/random/0331"
basename = "cutmock_1e9.hdf5"
fname = os.path.join(dir... | github_jupyter |
```
!pip install git+https://github.com/AlpacaDB/backlight
import os
import numpy as np
import pandas as pd
import backlight
```
# Generate example dummy data
```
np.random.seed(0)
# market data
if not os.path.exists("example_market.csv"):
idx = pd.date_range("2018-04-01 00:00:00", "2018-06-30 23:59:59", freq="1... | github_jupyter |
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc"><ul class="toc-item"><li><span><a href="#Load-data" data-toc-modified-id="Load-data-1"><span class="toc-item-num">1 </span>Load data</a></span></li><li><span><a href="#All-patients" data-toc-modified-id="All-patients-2"><span class="toc... | github_jupyter |
# Lateral Movement
The adversary is trying to move through your environment.
Lateral Movement consists of techniques that adversaries use to enter and control remote systems on a network. Following through on their primary objective often requires exploring the network to find their target and subsequently gaining acc... | github_jupyter |
```
import json
import pandas as pd
import operator
with open('../docs/data/dams.geojson') as f:
in_json = json.load(f)
in_ftrs = in_json['features']
ftr1 = in_ftrs[0]
prop1 = ftr1['properties']
var_names = prop1.keys()
types = {}
vals = {}
to_skip = ['Url_Address','NID_ID','key']
for v in prop1:
if v not i... | github_jupyter |
```
%matplotlib inline
%config InlineBackend.figure_formats = {'png', 'retina'}
data_key = pd.read_csv('key.csv')
data_key = data_key[data_key['station_nbr'] != 5]
data_weather = pd.read_csv('weather.csv')
data_weather = data_weather[data_weather['station_nbr'] != 5] ## Station 5번 제거한 나머지
data_train = pd.read_csv('tra... | github_jupyter |
```
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import sys
from tqdm import tqdm
sys.path.insert(0,'..')
%matplotlib inline
from dataset import Dataset
from models import CNP
from train import Trainer
from utils.dataset_utils import (load_data,train_test_split, make_features)
from types impo... | github_jupyter |
# <img style="float: left; padding-right: 10px; width: 45px" src="https://raw.githubusercontent.com/Harvard-IACS/2018-CS109A/master/content/styles/iacs.png"> CS-109B Introduction to Data Science
## Lab 5: Convolutional Neural Networks
**Harvard University**<br>
**Spring 2020**<br>
**Instructors:** Mark Glickman, Pavlo... | github_jupyter |
```
# default_exp naive_bayes
#hide
from nbdev.showdoc import *
# all_flag
```
# Naive Bayes Classifier
> Summary: Naive Bayes, Text classification, Sentiment analysis, bag-of-words, BOW
## What is Naive Bayes Method?
Naive Bayes technique is a supervised method. It is a probabilistic learning method for classifyin... | github_jupyter |
# Table of Contents
<div class="toc" style="margin-top: 1em;"><ul class="toc-item" id="toc-level0"><li><span><a href="http://localhost:8889/notebooks/nn_postprocessing/discrete_crps_test.ipynb#Sebastians-example" data-toc-modified-id="Sebastians-example-1"><span class="toc-item-num">1 </span>Sebastians exam... | github_jupyter |
# Regularization
Welcome to the second assignment of this week. Deep Learning models have so much flexibility and capacity that **overfitting can be a serious problem**, if the training dataset is not big enough. Sure it does well on the training set, but the learned network **doesn't generalize to new examples** that... | github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
from scipy import stats
import pymc3 as pm
import arviz as az
```
# Preparation of results for business: Newpaper Sales
We will illustrate how to prepare results for a business audience using ArviZ. The motivating example we'll use is a classic example in Industr... | github_jupyter |
<a href="https://colab.research.google.com/github/rlworkgroup/garage/blob/master/examples/jupyter/trpo_gym_tf_cartpole.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
This is a jupyter notebook demonstrating usage of [garage](https://github.com/rlwo... | github_jupyter |
#### Outline
- for each dataset:
- load dataset;
- for each network:
- load network
- project 1000 test dataset samples
- save to metric dataframe
```
# reload packages
%load_ext autoreload
%autoreload 2
```
### Choose GPU (this may not be needed on your computer)
```
%env CUDA_DEV... | github_jupyter |
# Analyze consensus motif
The third output from the computational pipeline is a fasta file of the best predicted promoter for each input sequence. For more details about how robust these predictions are, see Section 2 of `inspect_BioProspector_results.ipynb`.
Given a fasta file of best predictions from a given set of... | github_jupyter |
```
import sys
sys.path.append('../../')
import os
import dill
import numpy as np
import scipy as sc
import random as rand
from sklearn import preprocessing, linear_model
import matplotlib.pyplot as plt
from core.controllers import ConstantController
from koopman_core.dynamics import LinearLiftedDynamics, BilinearLi... | github_jupyter |
```
import numpy as np
import librosa
import glob
import os
from random import randint
import torch
import torch.nn as nn
from torch.utils import data
import torch.optim as optim
from torch.utils.data import DataLoader
from torch.utils.data import sampler
import matplotlib.pyplot as plt
%matplotlib inline
import impor... | github_jupyter |
## Lab 1: Tensor Manipulation
First Author: Seungjae Ryan Lee (seungjaeryanlee at gmail dot com)
Second Author: Ki Hyun Kim (nlp.with.deep.learning at gmail dot com)
<div class="alert alert-warning">
NOTE: This corresponds to <a href="https://www.youtube.com/watch?v=ZYX0FaqUeN4&t=23s&list=PLlMkM4tgfjnLSOjrEJN31gZ... | github_jupyter |
### Introduction
This notebook provides an example for how to use the PAKKR library in a training and validation pipeline using Fisher's iris dataset.
### Setup
Install the packages required for this example
```
%pip install numpy pandas scikit-learn
from typing import Callable, Dict, NamedTuple, List, Union, Tuple
... | github_jupyter |
# Class 5 Lab: Databases and ETL
## Objectives
- Configure Google Cloud SQL Databases
- Discover Database Security Options
- Connect to a MySQL DB via Python
- Generate UUIDs in Python
- Normalize API request payload
- Insert API request payloads into DB tables
## Requirements
In order to follow along, the following... | github_jupyter |
<a href="https://colab.research.google.com/github/Harrow-Enigma/TeamEngima-ProjectEco-AI/blob/main/Project_Eco_AI_Beta_Testing.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Project ECO AI Beta Testing Model
Copyright 2021 YIDING SONG
Licensed ... | github_jupyter |
```
#Gerekli kütüphaneler
import pandas as pd
import numpy as np
import requests
from bs4 import BeautifulSoup
#Gerekli listeler
url_list = []
prices_list = []
propTitles = []
propValues = []
#Özelliklerin çekilmesi
for i in range(1,2): #2 yerine sayfa sayısı gelmeli
url = "https://www.trendyol.com/cep-telefonu-x-c... | github_jupyter |
# CIFAR10 CNN Classification
Note: This notebook is desinged to run with Python3 and GPU runtime.
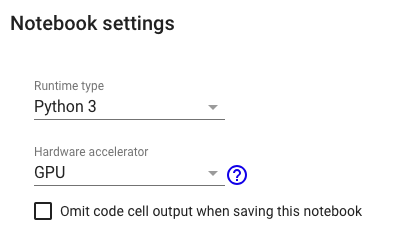
This notebook uses TensorFlow 2.x.
```
%tensorflow_version 2.x
```
####[CCC-01]
Import modules and ... | github_jupyter |
# About the data
The 20 newsgroups dataset comprises around 18000 newsgroups posts on 20 topics. The classification problem is to identify the newsgroup a post was summited to, given the text of the post.
There are a few versions of this dataset from different sources online. Below, we use the version within scikit-l... | github_jupyter |
```
import OnePy as op
%matplotlib inline
```
# Cleaner介绍
```
from OnePy.sys_module.base_cleaner import CleanerBase
class SMA(CleanerBase):
"""
编写自己的cleaner,只需自己创建一个CleanerBase的子类,然后覆盖calculate方法
默认提供data字典, key 为 ticker_frequency 的形式,比如 000001_D
self.data 内又是以open,high,low,close,volume为键值的字典,
每... | github_jupyter |
```
import numpy as np
np.random.seed(1)
# grAdapt
import grAdapt
from grAdapt.space.datatype import Float, Integer
from grAdapt.optimizer import AMSGrad, Adam, AMSGradBisection
from grAdapt.surrogate import GPRSlidingWindow, NoModel, NoGradient
from grAdapt.models import Sequential
# sklearn
# Import datasets, class... | github_jupyter |
```
#
# Convolution Neural Network Image classifier
#
# @author becxer
# @email becxer87@gmail.com
# @reference https://github.com/sjchoi86/Tensorflow-101
#
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
%matplotlib inline
print ("packages are loaded")
# Load npz data
npz_path = "images/... | github_jupyter |
# Building Dense Vectors Using Transformers
We will be using the [`sentence-transformers/stsb-distilbert-base`](https://huggingface.co/sentence-transformers/stsb-distilbert-base) model to build our dense vectors.
```
from transformers import AutoTokenizer, AutoModel
import torch
```
First we initialize our model and... | github_jupyter |
# 2 Dead reckoning
*Dead reckoning* is a means of navigation that does not rely on external observations. Instead, a robot’s position is estimated by summing its incremental movements relative to a known starting point.
Estimates of the distance traversed are usually obtained from measuring how many times the wheels ... | github_jupyter |
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed u... | github_jupyter |
# Best-practices for Cloud-Optimized Geotiffs
**Part 4. Dask GatewayCluster**
Unlike LocalCluster, a Dask GatewayCluster gives us the ability to dynamically increase our CPU and RAM across many machines! This is extremely powerful, because now we can load very big datasets into RAM for efficient calculations. There i... | github_jupyter |
# 적층 양방향 LSTM 감성 분류기
이 노트북에서 *적층* 양방향 LSTM을 사용해 감성에 따라 IMDB 영화 리뷰를 분류합니다.
[](https://colab.research.google.com/github/rickiepark/dl-illustrated/blob/master/notebooks/11-7.stacked_bi_lstm_sentiment_classifier.ipynb)
#### 라이브러리 적재
```
from tens... | github_jupyter |
```
import pandas as pd
df = pd.read_csv('/Users/pbhagwat/DEV/CohortAnalysis/Cohort-Analysis/Data/Telco-Customer-Churn.csv')
pd.set_option('display.max_columns', 100)
df.head()
dummies = pd.get_dummies(
df[['gender', 'SeniorCitizen', 'Partner', 'Dependents', 'tenure', 'PhoneService', 'MultipleLines',
... | github_jupyter |
# Import Libraries
```
#from __future__ import print_function
from pandas import read_csv
from pandas import DataFrame
from pandas import concat
from datetime import datetime
from matplotlib import pyplot
from math import sqrt
from numpy import concatenate
from sklearn.preprocessing import MinMaxScaler
from sklearn.pr... | github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
from scipy import signal
```
FT algorithm receives a trajectory, apply its filters to find the appropriate cycles, and outputs the full set of cyclic components. There are two algorithms:
- the Discrete Fourier Transform (DFT) which requires $O(n^2)$ operations (... | github_jupyter |
# cyBERT: a flexible log parser based on the BERT language model
## Table of Contents
* Introduction
* Generating Labeled Logs
* Subword Tokenization
* Data Loading
* Fine-tuning pretrained BERT
* Model Evaluation
* Parsing with cyBERT
## Introduction
One of the most arduous tasks of any security operation (and equa... | github_jupyter |
```
# Imports
from datetime import datetime, timedelta
from Database import db
import numpy as np
import pickle
import os
import re
import matplotlib.pyplot as plt
from tqdm import tqdm_notebook
from keras.optimizers import RMSprop
from keras.models import Sequential, load_model, Model
from keras.preprocessing.te... | github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/GetStarted/05_map_function.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" h... | github_jupyter |
```
%load_ext autoreload
%autoreload 2
import sys
sys.path.append("..")
from optimus import Optimus
# Create optimus
op = Optimus("dask", verbose = True)
```
# Mysql
```
# !pip install mysqlclient
# Put your db credentials here
db = op.connect(
driver="mysql",
host="165.227.196.70",
database= "optimus"... | github_jupyter |
# Migrating from PyTorch Lightning
[PyTorch Lightning](https://www.pytorchlightning.ai/) is a popular and very well designed framework for training deep learning models. If you are interested in trying our efficient algorithms and using the Composer trainer, the below is a quick guide on how to adapt your models.
If ... | github_jupyter |
```
import matplotlib.pyplot as plt
from matplotlib.ticker import MaxNLocator
import numpy as np
algos_labels = ['CBS', 'CBS+PC', 'CBS+DS', 'CBS+H']
def represent_scatter(min_agents, max_agents, results, ylabel, title, ax):
ax.xaxis.set_major_locator(MaxNLocator(integer=True))
agents = range(min_agents, max_age... | github_jupyter |
# 4c. Improving the training loop
Now that we are able to compute the loss for our training data, we are able to train the model with the same couple of steps that we have encountered at the end of [**Notebook 2**](../2_Tensors/2b_Tensors_features_Solution.ipynb).
We will take this as a starting point to introduce th... | github_jupyter |
This is the notebook associated with the blog post titled Interactive Explainable Machine Learning with SAS Viya, Streamlit and Docker
Install SWAT if you haven't done so already. Import the required modules
```
#!pip install swat
from swat import CAS, options
import pandas as pd
import numpy as np
```
Connect to CA... | github_jupyter |
```
%matplotlib inline
%run notebook_setup
```
# Interpolation with PyMC3
## A 1D example
To start, we'll do a simple 1D example where we have a model evaluated at control points and we interpolate between them to estimate the model value.
```
import numpy as np
import matplotlib.pyplot as plt
import exoplanet as ... | github_jupyter |
```
#Basics
import pandas as pd
import numpy as np
#sklearn
from sklearn.model_selection import train_test_split,cross_val_score,GridSearchCV
from sklearn.linear_model import LogisticRegression
from sklearn.svm import SVC
from sklearn.ensemble import RandomForestRegressor,RandomForestClassifier
from sklearn.metrics im... | github_jupyter |
# Módulo 2: Scraping con Selenium
## LATAM Airlines
<a href="https://www.latam.com/es_ar/"><img src="https://i.pinimg.com/originals/dd/52/74/dd5274702d1382d696caeb6e0f6980c5.png" width="420"></img></a>
<br>
Vamos a scrapear el sitio de Latam para averiguar datos de vuelos en funcion el origen y destino, fecha y cabin... | github_jupyter |
# Data setup
```
#Uploading Dataset
from google.colab import files
uploaded = files.upload()
# ignore the error
pip install -U numpy pandas scikit-learn
import os
import glob
import datetime
from collections import defaultdict
import pandas as pd
from sklearn.model_selection import train_test_split, StratifiedKFold
f... | github_jupyter |
##### Copyright 2018 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or ... | github_jupyter |
# GCP Dataflow Component Sample
A Kubeflow Pipeline component that prepares data by submitting an Apache Beam job (authored in Python) to Cloud Dataflow for execution. The Python Beam code is run with Cloud Dataflow Runner.
## Intended use
Use this component to run a Python Beam code to submit a Cloud Dataflow job as... | github_jupyter |
<small><small><i>
All the IPython Notebooks in this lecture series by Dr. Milan Parmar are available @ **[GitHub](https://github.com/milaan9/05_Python_Files)**
</i></small></small>
# Python Directory and Files Management
In this class, you'll learn about file and directory management in Python, i.e. creating a direct... | github_jupyter |
# Machine Learning with PySpark - Introduction
> Spark is a framework for working with Big Data. In this chapter you'll cover some background about Spark and Machine Learning. You'll then find out how to connect to Spark using Python and load CSV data.
You'll learn about them in this chapter. This is the Summary of le... | github_jupyter |
<a href="https://colab.research.google.com/github/jan-kreischer/UZH_ML4NLP/blob/main/Project-01/index_jan.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Exercise 01 - Linear Classification
## Dependencies
```
!pip install demoji
!pip install go... | github_jupyter |
# PyNNDescent Performance
How fast is PyNNDescent for approximate nearest neighbor search? How does it compare with other approximate nearest neighbor search algorithms and implementations? To answer these kinds of questions we'll make use of the [ann-benchmarks](https://github.com/erikbern/ann-benchmarks) suite of to... | github_jupyter |
# Mask R-CNN - Train on NewShapes Dataset
### Notes from implementation
This notebook shows how to train Mask R-CNN on your own dataset. To keep things simple we use a synthetic dataset of shapes (squares, triangles, and circles) which enables fast training. You'd still need a GPU, though, because the network backbon... | github_jupyter |
# Momentum and AdaGrad
Presented during ML reading group, 2019-11-5.
Author: Ioana Plajer, ioana.plajer@unitbv.ro
```
#%matplotlib notebook
%matplotlib inline
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
print(f'Numpy version: {np.__version__}'... | github_jupyter |
```
import os
import cv2
import math
import warnings
import numpy as np
import pandas as pd
import seaborn as sns
import tensorflow as tf
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix, fbeta_score
from keras import optimizers
from keras... | github_jupyter |
# Integral Calculus
:label:`sec_integral_calculus`
Differentiation only makes up half of the content of a traditional calculus education. The other pillar, integration, starts out seeming a rather disjoint question, "What is the area underneath this curve?" While seemingly unrelated, integration is tightly intertwin... | github_jupyter |
# Latent Dirichlet Allocation for Text Data
In this assignment you will
* apply standard preprocessing techniques on Wikipedia text data
* use GraphLab Create to fit a Latent Dirichlet allocation (LDA) model
* explore and interpret the results, including topic keywords and topic assignments for documents
Recall that... | github_jupyter |
<center>
<img src="https://gitlab.com/ibm/skills-network/courses/placeholder101/-/raw/master/labs/module%201/images/IDSNlogo.png" width="300" alt="cognitiveclass.ai logo" />
</center>
# **SpaceX Falcon 9 first stage Landing Prediction**
# Lab 1: Collecting the data
Estimated time needed: **45** minutes
In thi... | github_jupyter |
```
# python packages pd
import numpy as np
import matplotlib.pyplot as plt
import sys
import os
import inspect
import tensorflow as tf
import keras
from keras.models import Sequential
from keras.layers import Dense, Embedding, LSTM, SpatialDropout1D, Bidirectional, Activation
from keras.layers import CuDNNLSTM
from k... | github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/Image/band_math.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https:... | github_jupyter |
# Class 4: Plotting with Matplotlib
Matplotlib is a powerful plotting module that is part of Python's standard library. The website for matplotlib is at http://matplotlib.org/. And you can find a bunch of examples at the following two locations: http://matplotlib.org/examples/index.html and http://matplotlib.org/galle... | github_jupyter |
# Multiclass Partition Explainer: Emotion Data Example
This notebook demonstrates how to use the partition explainer for multiclass scenario with text data and visualize feature attributions towards individual classes. For computing shap values for a multiclass scenario, it uses the partition explainer over the text d... | github_jupyter |
```
from WenShuan import WenShuan
from bs4 import BeautifulSoup
import re
from matplotlib import pyplot as plt
%matplotlib inline
%config InlineBackend.figure_format = "retina"
```
# Organize WenShuan into Text and Comment Tuples
In this notebook, we would try to split texts and commentaries in WenShuan into a list ... | github_jupyter |
# `AStream` Online Training
**[THIS IS WORK IN PROGRESS]**
This notebook performs online training of the **appearance stream parent model** on the **car-shadow** sequence, so make sure you've run the [`AStream` Offline Training](astream_offline_training.ipynb) notebook before running this one.
The online training o... | github_jupyter |
This notebook was prepared by [Donne Martin](https://github.com/donnemartin). Source and license info is on [GitHub](https://github.com/donnemartin/interactive-coding-challenges).
# Solution Notebook
## Problem: Given sorted arrays A, B, merge B into A in sorted order.
* [Constraints](#Constraints)
* [Test Cases](#T... | github_jupyter |
# Named Entity Recognition (NER) With SpaCy
We will be performing NER on threads from the **Investing** subreddit, but first let's test SpaCy for named entity recognition (NER) using an example from */r/investing*.
```
import spacy
from spacy import displacy
!python -m spacy download en_core_web_sm
nlp = spacy.load('... | github_jupyter |
# Named Entity Recognition (NER)
In this, you will learn to build a model for Named Entity Recognition (NER) task with Trax.
# Introduction
We first start by defining named entity recognition (NER). NER is a subtask of information extraction that locates and classifies named entities in a text. The named entities co... | github_jupyter |
**Authors:** Jozef Hanč, Martina Hančová <br>
[Faculty of Science](https://www.upjs.sk/en/faculty-of-science/?prefferedLang=EN) *P. J. Šafárik University in Košice, Slovakia* <br>
email: [jozef.hanc@upjs.sk](mailto:jozef.hanc@upjs.sk)
***
**<font size=6 color=brown> Introduction</font>**
**<font size=4> Scholarly ... | github_jupyter |
---
layout: page
title: Intervalos de Confiança
nav_order: 9
---
[<img src="./colab_favicon_small.png" style="float: right;">](https://colab.research.google.com/github/icd-ufmg/icd-ufmg.github.io/blob/master/_lessons/09-ics.ipynb)
# Intervalos de Confiança
{: .no_toc .mb-2 }
Conceito base para pesquisas estatísticas... | github_jupyter |
The InfiniteHMM class is capable of reading a GROMACS trajectory file and converting the xy coordinates to radial coordinates with respect to the pore centers. This is all done in the __init__ function. This notebook outlines how the radial coordinates are calculated.
```
import hdphmm
import mdtraj as md
```
First, ... | github_jupyter |
## Import modules. Remember it is always good practice to do this at the beginning of a notebook.
If you don't have seaborn, you can install it with conda install seaborn
```
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
```
### Use notebook magic to render matplotlib figures inline with ... | github_jupyter |
# ORF recognition by CNN
Use variable number of bases between START and STOP. Thus, ncRNA will have its STOP out-of-frame or too close to the START, and pcRNA will have its STOP in-frame and far from the START.
```
import time
t = time.time()
time.strftime('%Y-%m-%d %H:%M:%S %Z', time.localtime(t))
PC_SEQUENCES=3200... | github_jupyter |
# Introduction to Python & Jupyter Notebooks
In this class, we will rely on Python as our main tool for data science. We will be running in Python in Jupyter Notebooks. Most of you are at home in Python, and will only have to spend a few moments here. I have planned for three scenarios
1. **You don't know anything ab... | github_jupyter |
# Generative Adversarial Network
In this notebook, we'll be building a generative adversarial network (GAN) trained on the MNIST dataset. From this, we'll be able to generate new handwritten digits!
GANs were [first reported on](https://arxiv.org/abs/1406.2661) in 2014 from Ian Goodfellow and others in Yoshua Bengio'... | github_jupyter |
```
import os
import sys
import random
import numpy as np
import pandas as pd
import torch
from torch.utils.data import Dataset, DataLoader
!pip install transformers
from transformers import BertTokenizer
from transformers import BertForSequenceClassification
from transformers import BertConfig
!pip install -U spacy[cu... | github_jupyter |
## Identifiability Test of Linear VAE on Synthetic Dataset
```
%load_ext autoreload
%autoreload 2
import torch
import torch.nn.functional as F
from torch.utils.data import DataLoader, random_split
import leap
import numpy as np
from leap.datasets.sim_dataset import SimulationDataset
from leap.modules.linear_vae import... | github_jupyter |
```
import sys
sys.path.append('..')
import torch
import pandas as pd
import numpy as np
import pickle
import argparse
import networkx as nx
from collections import Counter
from torch_geometric.utils import dense_to_sparse, degree
import matplotlib.pyplot as plt
from src.gcn import GCNSynthetic
from src.utils.utils imp... | github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import scipy.stats as ss
import tensorflow as tf
import math
import random
import tensorflow as tf
from tensorflow.keras import backend as K
from tensorflow.keras.utils import get_custom_objects
from tensorflow.keras.layers import Activation
df=... | github_jupyter |
### 98. Validate Binary Search Tree
#### Content
<p>Given the <code>root</code> of a binary tree, <em>determine if it is a valid binary search tree (BST)</em>.</p>
<p>A <strong>valid BST</strong> is defined as follows:</p>
<ul>
<li>The left subtree of a node contains only nodes with keys <strong>less than</strong> ... | github_jupyter |
# Rethinking Statistics course in Stan - Week 5
Lecture 9: Conditional Manatees
- [Video](https://www.youtube.com/watch?v=QhHfo6-Bx8o)
- [Slides](https://speakerdeck.com/rmcelreath/l09-statistical-rethinking-winter-2019)
Lecture 10: Markov Chain Monte Carlo
- [Video](https://youtu.be/v-j0UmWf3Us)
- [Slides](https:/... | github_jupyter |
```
top_directory = '/Users/iaincarmichael/Dropbox/Research/law/law-net/'
from __future__ import division
import os
import sys
import time
from math import *
import copy
import cPickle as pickle
# data
import numpy as np
import pandas as pd
# viz
import matplotlib.pyplot as plt
# graph
import igraph as ig
# NLP... | github_jupyter |
# EODAG as STAC client
## STAC API
EODAG can perform an item search over a STAC compliant API. Found STAC items are returned as [EOProduct](../../api_reference/eoproduct.rst#eodag.api.product._product.EOProduct) objects with STAC metadata mapped to OGC OpenSearch Extension for Earth Observation.
EODAG comes with alre... | github_jupyter |
## **MoroccoAI Data Challenge (Edition 001)**
This notebook walks through The prcoccess of detecting plates from images using our 2 Fast-RCNN models that were trained on Plate Detection and Moroccan Plate Charachter Detection, and the post-processing that followed the predection.
<br>
### **Overview**
In Morocco, t... | github_jupyter |
# Gene enrichment analysis
```
from pymodulon.enrichment import *
from pymodulon.example_data import load_ecoli_data, trn
ica_data = load_ecoli_data()
```
## General functions
To perform a basic enrichment test between two gene sets, use the ``compute_enrichment`` function.
Optional arguments:
* ``label``: Label fo... | github_jupyter |
```
# !export SPOTIPY_CLIENT_ID='63594c9b2f99411a8cbd18df04851fc4'
# !export SPOTIPY_CLIENT_SECRET='096168b2bd1f4378ae410726955c9ed8'
# !export SPOTIPY_REDIRECT_URI='https://www.google.com/'
# ! SPOTIPY_CLIENT_ID
import os
import sys
import json
import spotipy
import webbrowser
import spotipy.util as util
from json.dec... | github_jupyter |
```
%matplotlib inline
from keras.datasets import mnist
from keras.layers import Input, Dense, Lambda
from keras.models import Model
from keras.objectives import binary_crossentropy
from keras.callbacks import LearningRateScheduler
import numpy as np
import matplotlib.pyplot as plt
import keras.backend as K
import ten... | github_jupyter |
# Exercise- Neural Network
As introduced in the previous section, a neural network is a powerful tool often utilized in machine learning. Because neural networks are, fundamentally, very mathematical, we'll use them to motivate Numpy!
We review the simplest neural network here:

The output of... | github_jupyter |
# 1.4 Data types
`Kiddo explanation 😇: `
We might use many materials like sand, bricks, concrete to construct a house. These are basic and essential needs to have the construction done and each of them have a specific role or usage.
Likewise, we need various data types like string, boolean, integer, dictionary etc.... | github_jupyter |
### This is a common homework assignment for both frameworks
This week's assignment appears to be unusually grandeur, so please read submission/grading guidelines before you upload it for review.
__Submisson__: To ease mutual pain, please submit
- Some kind of readable report with links to your evaluations, gym uploa... | github_jupyter |
# Data Structures
* tuple
* list
* dict
* set
## tuple
A tuple is a one dimensional, fixed-length, immutable sequence.
Create a tuple:
```
tup = (1, 2, 3)
tup
```
Convert to a tuple:
```
list_1 = [1, 2, 3]
type(tuple(list_1))
```
Create a nested tuple:
```
nested_tup = ([1, 2, 3], (4, 5))
nested_tup
```
Acces... | github_jupyter |
# Python Crash Course Exercises
This is an optional exercise to test your understanding of Python Basics. If you find this extremely challenging, then you probably are not ready for the rest of this course yet and don't have enough programming experience to continue. I would suggest you take another course more geare... | github_jupyter |
## Model compression demo
This notebook demonstrates model compression through quantization using TFLite. We trained a ResNet50 mask/no-mask model to demonstrate this, which can be found in ../data/classifier_model_weights/resnet50_classifier.h5. Of course you are free to train your own model using the train-mask-noma... | github_jupyter |
```
# Copyright 2021 Google LLC
# Use of this source code is governed by an MIT-style
# license that can be found in the LICENSE file or at
# https://opensource.org/licenses/MIT.
# Author(s): Kevin P. Murphy (murphyk@gmail.com) and Mahmoud Soliman (mjs@aucegypt.edu)
```
<a href="https://opensource.org/licenses/MIT" t... | github_jupyter |
# Project 1
- **Team Members**: Chika Ozodiegwu, Kelsey Wyatt, Libardo Lambrano, Kurt Pessa

### Data set used::
* https://open-fdoh.hub.arcgis.com/datasets/florida-covid19-case-line-data
##### Dependencies
```
import step1_raw_data_collection as step1
import step2_data_processi... | github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import mglearn
from IPython.display import display
from sklearn.model_selection import train_test_split
%matplotlib inline
mglearn.plots.plot_knn_regression(n_neighbors=1)
mglearn.plots.plot_knn_regression(n_neighbors=3)
# Implementing the knn ... | github_jupyter |
# Randomized Image Sampling for Explanations (RISE)
```
import os
import numpy as np
from matplotlib import pyplot as plt
from skimage.transform import resize
from tqdm import tqdm
```
## Change code below to incorporate your *model* and *input processing*
### Define your model here:
```
from keras.applications.res... | github_jupyter |
<center>
<img src="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-ML0101EN-SkillsNetwork/labs/Module%202/images/IDSNlogo.png" width="300" alt="cognitiveclass.ai logo" />
</center>
# Simple Linear Regression
Estimated time needed: **15** minutes
## Objectives
After ... | github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.