text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
<a href="https://colab.research.google.com/github/jyjoon001/KoBART-KorQuAD/blob/main/AIC4010_OLD_ENG.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
! pip install datasets transformers
! pip install kobart-transformers
from google.colab import drive
drive.mount('/content/drive')
%cd /content/drive/MyDrive
# !mkdir squad_v1_kor
# !wget https://korquad.github.io/dataset/KorQuAD_v1.0_train.json -O squad_v1_kor/KorQuAD_v1.0_train.json
# !wget https://korquad.github.io/dataset/KorQuAD_v1.0_dev.json -O squad_v1_kor/KorQuAD_v1.0_dev.json
!mkdir squad
!wget https://rajpurkar.github.io/SQuAD-explorer/dataset/train-v1.1.json -O squad/train-v1.1.json
!wget https://rajpurkar.github.io/SQuAD-explorer/dataset/dev-v1.1.json -O squad/dev-v1.1.json
import json
from pathlib import Path
def read_squad(path):
path = Path(path)
with open(path, 'rb') as f:
squad_dict = json.load(f)
contexts = []
questions = []
answers = []
for group in squad_dict['data']:
for passage in group['paragraphs']:
context = passage['context']
for qa in passage['qas']:
question = qa['question']
for answer in qa['answers']:
contexts.append(context)
questions.append(question)
answers.append(answer)
return contexts, questions, answers
train_contexts, train_questions, train_answers = read_squad('squad/train-v1.1.json')
val_contexts, val_questions, val_answers = read_squad('squad/dev-v1.1.json')
def add_end_idx(answers, contexts):
for answer, context in zip(answers, contexts):
gold_text = answer['text']
start_idx = answer['answer_start']
end_idx = start_idx + len(gold_text)
# sometimes squad answers are off by a character or two – fix this
if context[start_idx:end_idx] == gold_text:
answer['answer_end'] = end_idx
elif context[start_idx-1:end_idx-1] == gold_text:
answer['answer_start'] = start_idx - 1
answer['answer_end'] = end_idx - 1 # When the gold label is off by one character
elif context[start_idx-2:end_idx-2] == gold_text:
answer['answer_start'] = start_idx - 2
answer['answer_end'] = end_idx - 2 # When the gold label is off by two characters
add_end_idx(train_answers, train_contexts)
add_end_idx(val_answers, val_contexts)
from transformers import DistilBertTokenizerFast
tokenizer = DistilBertTokenizerFast.from_pretrained('distilbert-base-uncased')
train_encodings = tokenizer(train_contexts, train_questions, truncation=True, padding=True)
val_encodings = tokenizer(val_contexts, val_questions, truncation=True, padding=True)
def add_token_positions(encodings, answers):
start_positions = []
end_positions = []
for i in range(len(answers)):
start_positions.append(encodings.char_to_token(i, answers[i]['answer_start']))
end_positions.append(encodings.char_to_token(i, answers[i]['answer_end'] - 1))
# if start position is None, the answer passage has been truncated
if start_positions[-1] is None:
start_positions[-1] = tokenizer.model_max_length
if end_positions[-1] is None:
end_positions[-1] = tokenizer.model_max_length
encodings.update({'start_positions': start_positions, 'end_positions': end_positions})
add_token_positions(train_encodings, train_answers)
add_token_positions(val_encodings, val_answers)
import torch
class SquadDataset(torch.utils.data.Dataset):
def __init__(self, encodings):
self.encodings = encodings
def __getitem__(self, idx):
return {key: torch.tensor(val[idx]) for key, val in self.encodings.items()}
def __len__(self):
return len(self.encodings.input_ids)
train_dataset = SquadDataset(train_encodings)
val_dataset = SquadDataset(val_encodings)
train_dataset[0]
from transformers import DistilBertForQuestionAnswering
model = DistilBertForQuestionAnswering.from_pretrained("distilbert-base-uncased")
from transformers import Trainer, TrainingArguments
training_args = TrainingArguments(
output_dir='./results', # output directory
num_train_epochs=3, # total number of training epochs
per_device_train_batch_size=16, # batch size per device during training
per_device_eval_batch_size=32, # batch size for evaluation
warmup_steps=500, # number of warmup steps for learning rate scheduler
weight_decay=0.01, # strength of weight decay
logging_dir='./logs', # directory for storing logs
logging_steps=10,
)
trainer = Trainer(
model=model, # the instantiated 🤗 Transformers model to be trained
args=training_args, # training arguments, defined above
train_dataset=train_dataset, # training dataset
eval_dataset=val_dataset # evaluation dataset
)
trainer.train()
trainer.save_model("test-squad-trained2")
import torch
for batch in trainer.get_eval_dataloader():
break
batch = {k: v.to(trainer.args.device) for k, v in batch.items()}
with torch.no_grad():
output = trainer.model(**batch)
output.keys()
output.start_logits.shape, output.end_logits.shape
output.start_logits.argmax(dim=-1), output.end_logits.argmax(dim=-1)
n_best_size = 20
import numpy as np
start_logits = output.start_logits[0].cpu().numpy()
end_logits = output.end_logits[0].cpu().numpy()
# Gather the indices the best start/end logits:
start_indexes = np.argsort(start_logits)[-1 : -n_best_size - 1 : -1].tolist()
end_indexes = np.argsort(end_logits)[-1 : -n_best_size - 1 : -1].tolist()
valid_answers = []
for start_index in start_indexes:
for end_index in end_indexes:
if start_index <= end_index: # We need to refine that test to check the answer is inside the context
valid_answers.append(
{
"score": start_logits[start_index] + end_logits[end_index],
"text": "" # We need to find a way to get back the original substring corresponding to the answer in the context
}
)
def prepare_validation_features(examples):
# Tokenize our examples with truncation and maybe padding, but keep the overflows using a stride. This results
# in one example possible giving several features when a context is long, each of those features having a
# context that overlaps a bit the context of the previous feature.
tokenized_examples = tokenizer(
examples["question" if pad_on_right else "context"],
examples["context" if pad_on_right else "question"],
truncation="only_second" if pad_on_right else "only_first",
max_length=max_length,
stride=doc_stride,
return_overflowing_tokens=True,
return_offsets_mapping=True,
padding="max_length",
)
# Since one example might give us several features if it has a long context, we need a map from a feature to
# its corresponding example. This key gives us just that.
sample_mapping = tokenized_examples.pop("overflow_to_sample_mapping")
# We keep the example_id that gave us this feature and we will store the offset mappings.
tokenized_examples["example_id"] = []
for i in range(len(tokenized_examples["input_ids"])):
# Grab the sequence corresponding to that example (to know what is the context and what is the question).
sequence_ids = tokenized_examples.sequence_ids(i)
context_index = 1 if pad_on_right else 0
# One example can give several spans, this is the index of the example containing this span of text.
sample_index = sample_mapping[i]
tokenized_examples["example_id"].append(examples["id"][sample_index])
# Set to None the offset_mapping that are not part of the context so it's easy to determine if a token
# position is part of the context or not.
tokenized_examples["offset_mapping"][i] = [
(o if sequence_ids[k] == context_index else None)
for k, o in enumerate(tokenized_examples["offset_mapping"][i])
]
return tokenized_examples
raw_predictions = trainer.predict(val_dataset)
max_answer_length = 30
start_logits = output.start_logits[0].cpu().numpy()
end_logits = output.end_logits[0].cpu().numpy()
offset_mapping = validation_features[0]["offset_mapping"]
# The first feature comes from the first example. For the more general case, we will need to be match the example_id to
# an example index
context = datasets["validation"][0]["context"]
# Gather the indices the best start/end logits:
start_indexes = np.argsort(start_logits)[-1 : -n_best_size - 1 : -1].tolist()
end_indexes = np.argsort(end_logits)[-1 : -n_best_size - 1 : -1].tolist()
valid_answers = []
for start_index in start_indexes:
for end_index in end_indexes:
# Don't consider out-of-scope answers, either because the indices are out of bounds or correspond
# to part of the input_ids that are not in the context.
if (
start_index >= len(offset_mapping)
or end_index >= len(offset_mapping)
or offset_mapping[start_index] is None
or offset_mapping[end_index] is None
):
continue
# Don't consider answers with a length that is either < 0 or > max_answer_length.
if end_index < start_index or end_index - start_index + 1 > max_answer_length:
continue
if start_index <= end_index: # We need to refine that test to check the answer is inside the context
start_char = offset_mapping[start_index][0]
end_char = offset_mapping[end_index][1]
valid_answers.append(
{
"score": start_logits[start_index] + end_logits[end_index],
"text": context[start_char: end_char]
}
)
valid_answers = sorted(valid_answers, key=lambda x: x["score"], reverse=True)[:n_best_size]
valid_answers
```
| github_jupyter |
# 影像分類
*電腦視覺*認知服務提供實用的預建模型以便使用影像工作,但是您需要經常訓練自己的電腦視覺模型。例如,假設 Northwind Traders 零售公司想要建立自動化結帳系統,該系統能夠依據相機在付款臺所拍攝的影像來識別客戶想要購買的雜貨店商品。為此,您需要訓練分類模型,該模型可以分類影像以識別正在購買的商品。

在 Azure 中,您可以使用***自訂視覺***認知服務在現有影像的基礎上來訓練影像分類模型。可以建立影像分類解決方案的元素有兩種。首先,您必須使用現有影像來訓練模型以識別不同的類別。然後,當模型訓練完成時,您必須將其發佈為能夠由應用程式使用的服務。
## 建立自訂視覺資源
若要使用自訂視覺服務,您需要可以用來*訓練*模型的 Azure 資源,以及可以*發佈*以供應用程式使用的資源。一項 (或者兩項) 工作的資源可以是一般性**認知服務**資源,或特定的**自訂視覺**資源。您可以將相同的認知服務資源用於這些工作中的每一項工作,或者您可以將不同的資源 (位於同一區域) 用於每項工作,以便分別管理成本。
使用以下指示來建立全新**自訂視覺**資源。
1. 在新的瀏覽器索引標籤中,透過 [https://portal.azure.com](https://portal.azure.com) 開啟 Azure 入口網站,並使用與您的 Azure 訂用帳戶關聯的 Microsoft 帳戶登入。
2. 選取 **[+ 建立資源]** 按鈕,搜尋*自訂視覺*,並建立包含以下設定的**自訂視覺**資源:
- **建立選項**: 兩個
- **訂用帳戶**: *您的 Azure 訂用帳戶*
- **資源群組**: *選取或建立具有唯一名稱的資源群組*
- **名稱**: *輸入唯一名稱*
- **訓練位置**: *選擇任一可用區域*
- **訓練定價層**: F0
- **預測位置**: *與訓練資源位於同一區域*
- **預測定價層**: F0
> **備註**: 若您的訂用帳戶中已經有 F0 自訂視覺服務,請為這一個選取 **[S0]**。
3. 等待資源建立,注意建置兩個自訂視覺資源,一個用於訓練,另一個用於預測。您可以透過導覽到建立它們的資源群組來檢視這些資源。
## 建立自訂視覺專案
要訓練物件偵測模型,您需要以訓練資源為基礎建立自訂視覺專案。為此,您將使用自訂視覺入口網站。
1. 從 https://aka.ms/fruit-images 中下載並擷取訓練影像。**注意:** 作為臨時解決辦法,如果您無法存取訓練影像,請前往 https://www.github.com,然後前往 https://aka.ms/fruit-images。
2. 在其它瀏覽器索引標籤中,透過 [https://customvision.ai](https://customvision.ai) 開啟自訂視覺入口網站。若出現提示,請使用與您的 Azure 訂用帳戶關聯的 Microsoft 帳戶登入並同意服務條款。
3. 在自訂視覺入口網站,建立一個包含以下設定的新專案:
- **名稱**: Grocery Checkout
- **描述**: 雜貨店影像分類
- **資源**: *先前建立的自訂視覺資源*
- **專案類型**: 分類
- **分類類型**: 多類別 (每個影像一個標記)
- **網域**: 食物
4. 按一下 **\[+\] [新增影像]**,並選取先前已擷取的**蘋果**資料夾中的所有檔案。然後上傳影像檔案,指定*蘋果*標記,如下所示:

5. 重複上一個步驟以便上傳**香蕉**資料夾中帶有*香蕉*標記的影像,以及**柳橙**資料夾中帶有*柳橙*標記的影像。
6. 瀏覽您已經在自訂視覺專案中上傳的影像,每個類別應該有 15 個影像,如下所示:

7. 在自訂視覺專案中,影像的上方,按一下 **[訓練]** 以使用已標記的影像訓練分類模型。選取 **[快速訓練]**選項,然後等待訓練反覆運算完成 (這可能需要一分鐘左右)。
8. 當模型反覆運算已經完成訓練,檢閱*精確度*、*重新叫用*和 *AP*效能計量,這些計量測量分類模型的預測正確性,各項計量應該都較高。
## 測試模型
在發佈此模型的反覆運算以供應用程式使用之前,您應該對其進行測試。
1.在效能計量上方,按一下 **[快速測試]**。
2. 在 **[影像 URL]** 方塊中,鍵入`https://aka.ms/apple-image` 並按一下 ➔
3. 檢視您的模型退回的預測,*蘋果*的可能性分數應該是最高的,如下所示:

4. 關閉 **[快速測試]**視窗。
## 發佈並使用影像分類模型
現在,您可以準備發佈已訓練的模型,還可以從用戶端應用程式中使用該模型。
9. 按一下 **[🗸 發佈]** 以發佈包含以下設定的已訓練模型:
- **模型名稱**: groceries
- **預測資源**: *先前已建立的預測資源*。
### (!)簽入
您是否使用了相同的模型名稱:**groceries**?
10. 發佈後,在 **[效能]** 頁面右上方按一下設定 *(⚙)* 圖示,以檢視專案設定。然後,在 **[一般]** 下 (在左側),複製**專案識別碼**。向下捲動並將其貼上到步驟 13 下面的程式碼儲存格中,取代 **YOUR_PROJECT_ID**。

> _**備註**:如果您在本次練習開始時使用了*認知服務*資源,而不是建立**自訂視覺**資源,則可以從專案設定的右側複製其金鑰和端點,將其貼上到下面的程式碼儲存格中,然後執行該資源以查看結果。否則,請繼續完成以下步驟以獲取自訂視覺預測資源的金鑰和端點。_
11. 在 **[專案設定]** 頁面的左上方,按一下*專案資源庫*(👁) 圖示以退回到自訂視覺入口網站首頁,現在您的專案已在其中列出。
12. 在自訂視覺入口網站首頁的右上方,按一下*設定*(⚙) 圖示以檢視自訂視覺服務的設定。然後,在**資源**下方,展開您**的預測**資源 (<u>而非</u>訓練資源) 並將其**金鑰**和**端點**值複製到步驟 13 下面的程式碼儲存格,取代 **YOUR_KEY** 和 **YOUR_ENDPOINT**。
### (!)簽入
若您使用的是**自訂視覺**資源,您是否使用*了預測*資源 (<u>而非</u> 訓練資源)?

13. 透過按一下 **[執行儲存格]**(▷) 按鈕 (位於儲存格左側) 執行下面的程式碼儲存格來為您的專案識別碼、金鑰和端點值設定變數。
```
project_id = 'YOUR_PROJECT_ID'
cv_key = 'YOUR_KEY'
cv_endpoint = 'YOUR_ENDPOINT'
model_name = 'groceries' # this must match the model name you set when publishing your model iteration (it's case-sensitive)!
print('Ready to predict using model {} in project {}'.format(model_name, project_id))
```
現在,您可以將金鑰和端點與自訂視覺用戶端一起使用,以連接到您的自訂視覺分類模型。
執行以下程式碼儲存格以便使用已發佈的模型分類測試影像的選取範圍。
> **備註**:請勿過於擔心程式碼的詳細資料。程式碼使用適用於 Python 的電腦視覺 SDK 來獲取針對 /data/image-classification/test-fruit 資料夾中的每個影像之類別預測。
```
from azure.cognitiveservices.vision.customvision.prediction import CustomVisionPredictionClient
from msrest.authentication import ApiKeyCredentials
import matplotlib.pyplot as plt
from PIL import Image
import os
%matplotlib inline
# Get the test images from the data/vision/test folder
test_folder = os.path.join('data', 'image-classification', 'test-fruit')
test_images = os.listdir(test_folder)
# Create an instance of the prediction service
credentials = ApiKeyCredentials(in_headers={"Prediction-key": cv_key})
custom_vision_client = CustomVisionPredictionClient(endpoint=cv_endpoint, credentials=credentials)
# Create a figure to display the results
fig = plt.figure(figsize=(16, 8))
# Get the images and show the predicted classes for each one
print('Classifying images in {} ...'.format(test_folder))
for i in range(len(test_images)):
# Open the image, and use the custom vision model to classify it
image_contents = open(os.path.join(test_folder, test_images[i]), "rb")
classification = custom_vision_client.classify_image(project_id, model_name, image_contents.read())
# The results include a prediction for each tag, in descending order of probability - get the first one
prediction = classification.predictions[0].tag_name
# Display the image with its predicted class
img = Image.open(os.path.join(test_folder, test_images[i]))
a=fig.add_subplot(len(test_images)/3, 3,i+1)
a.axis('off')
imgplot = plt.imshow(img)
a.set_title(prediction)
plt.show()
```
希望您的影像分類模型已正確識別了影像中的雜貨店。
## 了解更多資訊
相比於我們在本練習中已探索的功能,自訂視覺服務可以提供更多功能。例如,您亦可使用自訂視覺服務來建立物件偵測模型**;其不僅可以分類影像中的物件,還能識別在影像中顯示物件位置的周框方塊**。
若要深入了解自訂視覺認知服務,請檢視[自訂視覺文件](https://docs.microsoft.com/azure/cognitive-services/custom-vision-service/home)
| github_jupyter |
```
import requests,pprint,json,datetime,time
from pymongo import MongoClient
from secrets import *
# Sign up for free API key at https://openweathermap.org/appid
```
# 3.1 Acquring data from an API
```
city='london'
requestString=u'https://api.openweathermap.org/data/2.5/weather?q=%s&APPID=%s' % (city,key)
res=requests.get(requestString)
res.reason
pprint.pprint(res.json())
def getData(city):
requestString=u'https://api.openweathermap.org/data/2.5/weather?q=%s&APPID=%s' % (city,key)
res=requests.get(requestString)
return res
res=getData('La paz')
res
```
# 3.2 Ingesting data into MongoDB
## Create DB and collection
```
client=MongoClient('localhost')
db=client.packt
weatherCollection=db.weather
res=weatherCollection.insert_one(res.json())
```
## Get list of cities
```
!wget http://bulk.openweathermap.org/sample/city.list.json.gz
!gunzip city.list.json.gz
!head city.list.json
with open('data/city.list.json','r') as inFile:
citiesJson=json.loads(inFile.read())
citiesJson[0]
type(citiesJson)
```
## Limit to Chilean cities
```
citiesJsonCL=filter(lambda x:x[u'country']==u'CL',citiesJson)
len(citiesJsonCL)
cities=map(lambda x:x['name'],citiesJsonCL)
ids=map(lambda x:x['id'],citiesJsonCL)
res=getData(cities[0])
res
```
## Cycle through cities
```
for i,name in zip(ids,cities):
res=getData(name)
if not res.status_code==200:
print 'Error grabbing data for %s' % name
print res.reason
else:
try:
weatherCollection.insert_one(res.json())
except e:
print 'Error inserting into DB' % e
print '(City %s)' % name
time.sleep(1)
# Sleep so we dont thrash the API
def getTimestamp(dt):
return pd.datetime.fromtimestamp(dt)
def getDate(tstamp):
dt=datetime.datetime.fromtimestamp(tstamp)
return dt.strftime('%b %d - %H:%m')
```
# 3.3 Querying MongoDB for useful information
```
cur=weatherCollection.find()
sortedCur=cur.sort('main.temp')
for doc in sortedCur.limit(10):
try:
print doc['name']
print doc['main']['temp']
except:
print 'Error: missing name/temp'
cur.count()
```
## Find the max recorded temperatures over time per station
```
pipeline=[]
pipeline.append({'$group':{'_id':'$name','maxTemp':{'$max':'$main.temp'}}})
# Group measurements by city name, extract maximum recorded tmep for each
pipeline.append({'$limit':10})
# Limit results to first 10 cities
cur=weatherCollection.aggregate(pipeline=pipeline)
for d in cur:
print d['_id'],d['maxTemp']
```
## Get datetime of max temp per station
```
pipeline=[]
pipeline.append({'$match':{'name':{'$exists':True}}})
# Filter out dirty rows
pipeline.append({'$sort':{'name':1,'main.temp':-1}})
# Sort by name (esc) and temperature (asc)
pipeline.append({'$group':{'_id':'$name','maxTemp':{'$first':'$main.temp'},'date':{'$first':'$dt'}}})
# Group by name, grab maximum temperature and date of temperature
# Sorted by temp so grab first document from each group
pipeline.append({'$limit':10})
# limit to first 10 cities
cur=weatherCollection.aggregate(pipeline=pipeline)
for d in cur:
print getDate(d['date']),d['_id'],d['maxTemp']
print ''
```
## Do sanity check for one city
```
for d in weatherCollection.find({'name':'Caletones'}):
print getDate(d['dt']),'',d['main']['temp']
```
| github_jupyter |
# Introduction
*This jupyter notebook is part of a [collection of notebooks](../index.ipynb) on various topics of Digital Signal Processing.
## Digital Signal Processing
A digital signal is a discrete representation of the information conveyed by the signal. The information itself may be discrete by nature, as numbers or letters, or it may be continuous as most physical quantities. In many cases a digital signal has been derived by [sampling](https://en.wikipedia.org/wiki/Sampling_(signal_processing%29) and [quantization](https://en.wikipedia.org/wiki/Quantization_(signal_processing%29) of an analog signal captured by a sensor from the real world.

The process of sampling and quantization is technically realized by an [analog-to-digital converter](https://en.wikipedia.org/wiki/Analog-to-digital_converter) (ADC). An example for a digital signal is the sound pressure produced by a human speaker picked up by a microphone whose electrical signal is fed into an ADC after amplification.
[Digital Signal Processing](https://en.wikipedia.org/wiki/Digital_signal_processing) (DSP) refers to the digital processing of signals. The field covers the mathematics, algorithms, techniques and hardware to analyze, manipulate and generate digital signals. This may involve linear as well as nonlinear processing which is performed on the signals itself or their representations in other domains. For instance processing in the frequency or spectral domain. The processing is realized either directly in hardware or more common these days by software on general purpose or specialized ([Digital Signal Processor](https://en.wikipedia.org/wiki/Digital_signal_processor)) microprocessors. By use of a [digital-to-analog converter](https://en.wikipedia.org/wiki/Digital-to-analog_converter) (DAC) digital signals may be converted back into the analog domain. For instance to feed a loudspeaker or control a display. A typical signal processing chain is depicted below.

The history of DSP is directly coupled to the spread of microprocessors and computers. While many foundations may be found in the field of numerical mathematics before, specialized theory for the processing of digital signals has emerged in the 1950s. In its early stage, DSP was only applied to a few critical applications due to limited availability and high cost of microprocessors. However, with widespread availability of highly integrated circuits at reasonable prices the usage of DSP techniques has exploded since the 1980s. Nowadays, they can be found in almost any non-commercial and commercial device. Application areas include but are not limited to the processing of audio/image/video signals, communications engineering, radar/sonar/seismics, instrumentation, control systems, simulations and medicine.
A number of benefits make the usage of digital in contrast to analog signal processing very attractive:
* **robustness against external influences**: Analog systems show typically some gradual dependency on environmental conditions. For instance the analog elements of a circuit change their values when the ambient temperature changes. Digital systems generate the same results as long as the logic circuits/microprocessors are used within their technical specifications.
* **long-term stability**: The elements of analog systems change their values gradually when aging. Digital systems do not show gradual changes until they malfunction.
* **flexibility of implementations**: DSP implemented by means of software offers a high degree of flexibility which is hard to realize by analog circuits. For instance branching and looping.
* **extended possibilities**: Besides traditional signal processing techniques (e.g. filtering) the digital processing of signals may involve algorithms from numerical mathematics or machine learning, just to name a few.
In summary, at less technical effort the reproducibility of DSP is higher compared to analog signal processing. DSP system are typically cheaper compared to their analog counterparts. This is due to the tremendous efforts that have been spend in the last decades in the development and production of digital circuits.
## Exercises and Computational Examples
The theory discussed in this course is accompanied by computational examples and exercises. They aim at improving the understanding and show typical applications. Most of the basis DSP techniques can be implemented quite straightforward in software. Various programming languages and software environments are used for this purpose. Quite common is the use of MATLAB® and Simulink® from [MathWorks®](https://www.mathworks.com), [GNU Octave](https://www.gnu.org/software/octave) and [Python](https://www.python.org/).
Here interactive Python ([IPython](https://ipython.org/)) scripts are used which are directly embedded in [Jupyter](http://jupyter.org/) notebooks. The scripts and their results can be viewed straightforward as static snapshots. However, interactive use is highly recommended. For instance by rerunning the examples after a change of parameters. The exercises point to cases of interest and include also sample solutions. Feel motivated to extend the examples and write your own algorithms. If you are not familiar with IPython and Jupyter please take a look at the various tutorials available, for instance this [video](https://www.youtube.com/watch?v=HW29067qVWk). The examples base on various Python toolboxes/modules for convenience and performance. The basic functionality for most signal processing applications is provided by [`numpy`](http://www.numpy.org/) and [`scipy.signal`](https://docs.scipy.org/doc/scipy/reference/signal.html). For visualization purposes [`matplotlib`](https://matplotlib.org/) is used.
## Contents
An understanding of the underlying mechanisms and the limitations of basic DSP techniques is essential for the design of more complex algorithms. This course covers the fundamentals of DSP. A focus is laid on the discussion of generic building blocks in contrast to more specific complex algorithms. These blocks are addressed by a detailed mathematical treatise as well as practical examples and exercises. The discussion of the mathematical background is important to understand the principles of operation and derive properties in a more general manner. An outlook to practical applications is given whenever possible.
The materials start off with a discussion of the limitations of **spectral analysis of deterministic signals**. These are essentially the limitations of the discrete Fourier transform and play therefore a prominent role in many fields of DSP. Practical signals are often of stochastic nature. The foundations of **random signals and their processing by linear time-invariant systems** is the next topic. It is not sensible to describe random signals by their amplitude values, instead statistical measures are introduced which characterize average properties. The **quantization** of sampled signals is required to derive a digital signal suitable for DSP. In general, quantization is a non-linear process from which the amplitude continuous signal cannot be recovered exactly. In order to quantify the resulting deviations, a statistical analysis of the quantization error is presented for various signal classes. The filtering of signals is a basic task in DSP. The **realization of non-recursive and recursive filters** is therefore discussed in detail. Amongst others this covers practically relevant aspects like computationally efficient algorithms and the effects of quantization. For the **design of digital filters** various techniques are introduced in the last section.
## Prerequisites
The materials assume that the reader is already well familiarized with the theory of signals and linear time-invariant systems. In particular
* continuous signals and systems,
* Fourier and Laplace transform,
* sampling of signals,
* discrete signals and systems,
* discrete-time Fourier transform (DTFT), discrete Fourier transform (DFT) and $z$-transform.
| github_jupyter |
Simple Stock Analysis
Stock analysis is the evaluation or examination of the stock market. There are many trading tools to use to analysis stocks such as fundamental and technical analysis. Fundamental analysis is more focused on data from the financial statements, economic reports, and company assets. Technical analysis is based on the study of the past of historical price to predict the future price movement. However, this tutorial is not to get rich quick. Therefore, do not use your money to trade based on this stock analysis. Please do not use this method to invest with your money and I am not responsible for you loss.
Simple stock is a basic stock analysis tutorial. There are 7 parts in this tutorial.
1. Import Libraries
2. Get data from Yahoo
3. Analysis Data
4. Understand the Data based on Statistics
5. Calculate Prices
6. Plot Charts
7. Calculate Holding Period Return
I. Import Libraries
```
# Libaries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
import warnings
warnings.filterwarnings("ignore")
import fix_yahoo_finance as yf
yf.pdr_override()
```
II. Get Data from Yahoo!
This section we will pull the data from the website in Yahoo. We will be using the company of Apple and the symbol is 'AAPL'. Also, we will have a starting date and ending date.
```
stock = 'AAPL'
start = '2015-01-01'
end = '2017-01-01'
df = yf.download(stock, start, end)
```
III. Analysis Data
```
df.head() # the first 5 rows
df.tail() # the last 5 rows
df.shape # (rows, columns)
df.columns # Shows names of columns
df.dtypes # Shows data types
df.info() # Shows information about DataFrame
df.describe() # Shows summary statistics based on stock data
```
IV. Understand the Data based on Statistics
We will be using "Adj. Closing" price to find the minimum, maximum, average and standard deviation prices. The reason we are using "Adj. Closing" because is mostly use for historical returns. Also, the Adjusting Prices is change where the stock was accounts for the dividend and splits. However, the "Closing" price was not including with dividend and splits.
```
# Use only Adj. Closing
# Find the minimum
df['Adj Close'].min()
# Find the maximum
df['Adj Close'].max()
# Find the average
df['Adj Close'].mean()
# Find the standard deviation
df['Adj Close'].std()
```
V. Calculate the Prices
This section, we will be calculating the daily returns, log returns, and other technical indicators such as RSI(Relative Strength Index), MA(Moving Average), SMA(Simple Moving Averga), EMA(Exponential Moving Average), and VWAP(Voume Weighted Average Price). Also, we will calculate drawdowns.
```
# Daily Returns
# Formula: (Today Price / Yesterday Price) - 1
df['Daily_Returns'] = df['Adj Close'].shift(1) / df['Adj Close'] - 1
df['Daily_Returns'].head()
# Another way of calculating Daily Returns in simple way
DR = df['Adj Close'].pct_change(1) # 1 is for "One Day" in the past
DR.head()
# Log Returns
# Formula: log(Today Price/ Yesterday Price)
df['Log_Returns'] = np.log(df['Adj Close']) - np.log(df['Adj Close'].shift(1))
```
In this part of this section, we will be using the library of technical analysis. This packages has many different types of technical indicators. However, it does not have every single technical indicators.
We do not need to do calculation since the library has done it for us.
https://mrjbq7.github.io/ta-lib/doc_index.html
```
import talib as ta
# Creating Indicators
n=30 # number of periods
# RSI(Relative Strength Index)
# RSI is technical analysis indicator
# https://www.investopedia.com/terms/r/rsi.asp
df['RSI']=ta.RSI(np.array(df['Adj Close'].shift(1)), timeperiod=n)
# MA(Moving Average)
# https://www.investopedia.com/terms/m/movingaverage.asp
df['MA']=ta.MA(np.array(df['Adj Close'].shift(1)), timeperiod=n, matype=0)
# SMA(Simple Moving Average)
# https://www.investopedia.com/terms/s/sma.asp
df['SMA']=ta.SMA(np.array(df['Adj Close'].shift(1)))
# EMA(Exponential Moving Average)
# https://www.investopedia.com/terms/e/ema.asp
df['EMA']=ta.EMA(np.array(df['Adj Close'].shift(1)), timeperiod=n)
# Volume Weighted Average Price - VWAP
# http://stockcharts.com/school/doku.php?id=chart_school:technical_indicators:vwap_intraday
df['VWAP'] = round(np.cumsum(df['Volume']*(df['High']+df['Low'])/2) / np.cumsum(df['Volume']), 2)
df.head()
# Drawdown
# Drawdown shows the decline price since the stock began trading
# https://www.investopedia.com/terms/d/drawdown.asp
# There are 252 trading day in a year
window = 252
# Calculate the maximum drawdown
# Use the min_period of 1 (1 is the least valid observations) for the first 252 day in the data
Maximum_Drawdown = df['Adj Close'].rolling(window, min_periods=1).max()
Daily_Drawdown = df['Adj Close']/Maximum_Drawdown - 1.0
# Calculate the negative drawdown
Negative_Drawdown = Daily_Drawdown.rolling(window, min_periods=1).min()
```
VI. Plot Charts
```
# Plot Simple Line Chart
# Plot Adj Close
plt.figure(figsize=(16,10))
df['Adj Close'].plot(grid=True)
plt.title("Stock Adj Close Price", fontsize=18, fontweight='bold')
plt.xlabel("Date", fontsize=12)
plt.ylabel("Price",fontsize=12)
plt.show()
# Plot High, Low, Adj Close
df[['High', 'Low', 'Adj Close']].plot(figsize=(16,10), grid=True)
plt.title("Stock Adj Close Price", fontsize=18, fontweight='bold')
plt.xlabel("Date", fontsize=12)
plt.ylabel("Price", fontsize=12)
plt.show()
# Plot Daily Returns
df['Daily_Returns'].plot(figsize=(12,6))
plt.title("Daily Returns",fontsize=18, fontweight='bold')
plt.xlabel("Date", fontsize=12)
plt.ylabel("Price", fontsize=12)
plt.show()
# Plot Log Returns
df['Log_Returns'].plot(figsize=(12,6))
plt.title("Log Returns", fontsize=18, fontweight='bold')
plt.xlabel("Date", fontsize=12)
plt.ylabel("Price", fontsize=12)
plt.show()
# Histogram of Daily Returns
# Histogram is distribution of numerical data and has a rectangle whose area is prportional to the frequency of a variable.
plt.figure(figsize=(16,10))
plt.hist(df['Daily_Returns'].dropna(), bins=100, label='Daily Returns data') # Drop NaN
plt.title("Histogram of Daily Returns", fontsize=18, fontweight='bold')
plt.axvline(df['Daily_Returns'].mean(), color='r', linestyle='dashed', linewidth=2) # Shows the average line
plt.xlabel("Date", fontsize=12)
plt.ylabel("Daily Returns", fontsize=12)
plt.show()
# Plot Drawdown
plt.figure(figsize=(16,10))
Daily_Drawdown.plot()
Negative_Drawdown.plot(color='r',grid=True)
plt.title("Maximum Drawdown", fontsize=18, fontweight='bold')
plt.xlabel("Date", fontsize=12)
plt.ylabel("Price", fontsize=12)
plt.show()
```
VII. Holding Period Return(HPR)
Holding period return (HPR) is the rate of return on an individual stocks or portfolio over the whole period during the time it was held and it a measurement of investment performance.
```
# https://www.investopedia.com/exam-guide/series-65/quantitative-methods/holding-period-return.asp
# Formula: (Ending Value of Investment + Dividend - Beginning Value of Investment) / Beginning Value of Investment
# To get dividend in Yahoo!
DIV = yf.download(stock, start, end, actions=True)['Dividends']
# See how much dividends and splits was given during the time period
DIV
# Add all the dividend
Total_Dividend = DIV.sum()
Total_Dividend
# You invest beginning 2015 and sold it end of 2017
HPR = (df['Adj Close'][502] + Total_Dividend - df['Adj Close'][0]) / df['Adj Close'][0]
HPR
# You can use round for 4 decimal points
print('Holding Period Return: ', str(round(HPR,4)*100)+"%")
```
We going to pick another stocks that is Microsoft and we will compare it to Apple.
```
MSFT = yf.download('MSFT', start, end)['Adj Close'] # Use Adj Close only
MSFT_DIV = yf.download('MSFT', start, end, actions=True)['Dividends']
MSFT.head() # Shows only Date and Adj Close
MSFT_DIV # Shows how much dividend was given
MSFT_Dividend = MSFT_DIV.sum()
MSFT_Dividend
# You invest beginning 2015 and sold it end of 2017
MSFT_HPR = (MSFT[502] + Total_Dividend - MSFT[0]) / MSFT[0]
MSFT_HPR
# You can use round for 4 decimal points
print('Apple Holding Period Return: ', str(round(HPR,4)*100)+"%")
print('Microsoft Holding Period Return: ', str(round(MSFT_HPR,4)*100)+"%")
```
In the conclusion, we use 2 stocks to compare holding period return. Therefore, Microsoft had higher holding period return than Apple. Therefore, I would invest in Microsoft based on the stock analysis. However, if you comparing 2 stocks or 2 portfolio. You would pick the ones with the highest rate of return.
| github_jupyter |
---
_You are currently looking at **version 1.0** of this notebook. To download notebooks and datafiles, as well as get help on Jupyter notebooks in the Coursera platform, visit the [Jupyter Notebook FAQ](https://www.coursera.org/learn/python-data-analysis/resources/0dhYG) course resource._
---
# Merging Dataframes
```
import pandas as pd
df = pd.DataFrame([{'Name': 'Chris', 'Item Purchased': 'Sponge', 'Cost': 22.50},
{'Name': 'Kevyn', 'Item Purchased': 'Kitty Litter', 'Cost': 2.50},
{'Name': 'Filip', 'Item Purchased': 'Spoon', 'Cost': 5.00}],
index=['Store 1', 'Store 1', 'Store 2'])
df
df['Date'] = ['December 1', 'January 1', 'mid-May']
df
df['Delivered'] = True
df
df['Feedback'] = ['Positive', None, 'Negative']
df
adf = df.reset_index()
#adf
adf['Date'] = pd.Series({0: 'December 1', 2: 'mid-May'})
adf
staff_df = pd.DataFrame([{'Name': 'Kelly', 'Role': 'Director of HR'},
{'Name': 'Sally', 'Role': 'Course liasion'},
{'Name': 'James', 'Role': 'Grader'}])
staff_df = staff_df.set_index('Name')
student_df = pd.DataFrame([{'Name': 'James', 'School': 'Business'},
{'Name': 'Mike', 'School': 'Law'},
{'Name': 'Sally', 'School': 'Engineering'}])
student_df = student_df.set_index('Name')
print(staff_df.head())
print()
print(student_df.head())
pd.merge(staff_df, student_df, how='outer', left_index=True, right_index=True)
pd.merge(staff_df, student_df, how='inner', left_index=True, right_index=True)
pd.merge(staff_df, student_df, how='left', left_index=True, right_index=True)
pd.merge(staff_df, student_df, how='right', left_index=True, right_index=True)
staff_df = staff_df.reset_index()
student_df = student_df.reset_index()
pd.merge(staff_df, student_df, how='left', left_on='Name', right_on='Name')
staff_df = pd.DataFrame([{'Name': 'Kelly', 'Role': 'Director of HR', 'Location': 'State Street'},
{'Name': 'Sally', 'Role': 'Course liasion', 'Location': 'Washington Avenue'},
{'Name': 'James', 'Role': 'Grader', 'Location': 'Washington Avenue'}])
student_df = pd.DataFrame([{'Name': 'James', 'School': 'Business', 'Location': '1024 Billiard Avenue'},
{'Name': 'Mike', 'School': 'Law', 'Location': 'Fraternity House #22'},
{'Name': 'Sally', 'School': 'Engineering', 'Location': '512 Wilson Crescent'}])
pd.merge(staff_df, student_df, how='left', left_on='Name', right_on='Name')
staff_df = pd.DataFrame([{'First Name': 'Kelly', 'Last Name': 'Desjardins', 'Role': 'Director of HR'},
{'First Name': 'Sally', 'Last Name': 'Brooks', 'Role': 'Course liasion'},
{'First Name': 'James', 'Last Name': 'Wilde', 'Role': 'Grader'}])
student_df = pd.DataFrame([{'First Name': 'James', 'Last Name': 'Hammond', 'School': 'Business'},
{'First Name': 'Mike', 'Last Name': 'Smith', 'School': 'Law'},
{'First Name': 'Sally', 'Last Name': 'Brooks', 'School': 'Engineering'}])
staff_df
student_df
pd.merge(staff_df, student_df, how='inner', left_on=['First Name','Last Name'], right_on=['First Name','Last Name'])
```
# Idiomatic Pandas: Making Code Pandorable
```
import pandas as pd
df = pd.read_csv('census.csv')
df
(df.where(df['SUMLEV']==50)
.dropna()
.set_index(['STNAME','CTYNAME'])
.rename(columns={'ESTIMATESBASE2010': 'Estimates Base 2010'}))
df = df[df['SUMLEV']==50]
df.set_index(['STNAME','CTYNAME'], inplace=True)
df.rename(columns={'ESTIMATESBASE2010': 'Estimates Base 2010'})
import numpy as np
def min_max(row):
data = row[['POPESTIMATE2010',
'POPESTIMATE2011',
'POPESTIMATE2012',
'POPESTIMATE2013',
'POPESTIMATE2014',
'POPESTIMATE2015']]
return pd.Series({'min': np.min(data), 'max': np.max(data)})
df.apply(min_max, axis=1)
import numpy as np
def min_max(row):
data = row[['POPESTIMATE2010',
'POPESTIMATE2011',
'POPESTIMATE2012',
'POPESTIMATE2013',
'POPESTIMATE2014',
'POPESTIMATE2015']]
row['max'] = np.max(data)
row['min'] = np.min(data)
return row
df.apply(min_max, axis=1)
rows = ['POPESTIMATE2010',
'POPESTIMATE2011',
'POPESTIMATE2012',
'POPESTIMATE2013',
'POPESTIMATE2014',
'POPESTIMATE2015']
df.apply(lambda x: np.max(x[rows]), axis=1)
```
# Group by
```
import pandas as pd
import numpy as np
df = pd.read_csv('census.csv')
df = df[df['SUMLEV']==50]
df
%%timeit -n 10
for state in df['STNAME'].unique():
avg = np.average(df.where(df['STNAME']==state).dropna()['CENSUS2010POP'])
print('Counties in state ' + state + ' have an average population of ' + str(avg))
%%timeit -n 10
for group, frame in df.groupby('STNAME'):
avg = np.average(frame['CENSUS2010POP'])
print('Counties in state ' + group + ' have an average population of ' + str(avg))
df.head()
df = df.set_index('STNAME')
def fun(item):
if item[0]<'M':
return 0
if item[0]<'Q':
return 1
return 2
for group, frame in df.groupby(fun):
print('There are ' + str(len(frame)) + ' records in group ' + str(group) + ' for processing.')
df = pd.read_csv('census.csv')
df = df[df['SUMLEV']==50]
df.groupby('STNAME').agg({'CENSUS2010POP': np.average})
print(type(df.groupby(level=0)['POPESTIMATE2010','POPESTIMATE2011']))
print(type(df.groupby(level=0)['POPESTIMATE2010']))
(df.set_index('STNAME').groupby(level=0)['CENSUS2010POP']
.agg({'avg': np.average, 'sum': np.sum}))
(df.set_index('STNAME').groupby(level=0)['POPESTIMATE2010','POPESTIMATE2011']
.agg({'avg': np.average, 'sum': np.sum}))
(df.set_index('STNAME').groupby(level=0)['POPESTIMATE2010','POPESTIMATE2011']
.agg({'POPESTIMATE2010': np.average, 'POPESTIMATE2011': np.sum}))
```
# Scales
```
df = pd.DataFrame(['A+', 'A', 'A-', 'B+', 'B', 'B-', 'C+', 'C', 'C-', 'D+', 'D'],
index=['excellent', 'excellent', 'excellent', 'good', 'good', 'good', 'ok', 'ok', 'ok', 'poor', 'poor'])
df.rename(columns={0: 'Grades'}, inplace=True)
df
df['Grades'].astype('category').head()
grades = df['Grades'].astype('category',
categories=['D', 'D+', 'C-', 'C', 'C+', 'B-', 'B', 'B+', 'A-', 'A', 'A+'],
ordered=True)
grades.head()
grades > 'C'
df = pd.read_csv('census.csv')
df = df[df['SUMLEV']==50]
df = df.set_index('STNAME').groupby(level=0)['CENSUS2010POP'].agg({'avg': np.average})
pd.cut(df['avg'],10)
```
# Pivot Tables
```
#http://open.canada.ca/data/en/dataset/98f1a129-f628-4ce4-b24d-6f16bf24dd64
df = pd.read_csv('cars.csv')
df.head()
df.pivot_table(values='(kW)', index='YEAR', columns='Make', aggfunc=np.mean)
df.pivot_table(values='(kW)', index='YEAR', columns='Make', aggfunc=[np.mean,np.min], margins=True)
```
# Date Functionality in Pandas
```
import pandas as pd
import numpy as np
```
### Timestamp
```
pd.Timestamp('9/1/2016 10:05AM')
```
### Period
```
pd.Period('1/2016')
pd.Period('3/5/2016')
```
### DatetimeIndex
```
t1 = pd.Series(list('abc'), [pd.Timestamp('2016-09-01'), pd.Timestamp('2016-09-02'), pd.Timestamp('2016-09-03')])
t1
type(t1.index)
```
### PeriodIndex
```
t2 = pd.Series(list('def'), [pd.Period('2016-09'), pd.Period('2016-10'), pd.Period('2016-11')])
t2
type(t2.index)
```
### Converting to Datetime
```
d1 = ['2 June 2013', 'Aug 29, 2014', '2015-06-26', '7/12/16']
ts3 = pd.DataFrame(np.random.randint(10, 100, (4,2)), index=d1, columns=list('ab'))
ts3
ts3.index = pd.to_datetime(ts3.index)
ts3
pd.to_datetime('4.7.12', dayfirst=True)
```
### Timedeltas
```
pd.Timestamp('9/3/2016')-pd.Timestamp('9/1/2016')
pd.Timestamp('9/2/2016 8:10AM') + pd.Timedelta('12D 3H')
```
### Working with Dates in a Dataframe
```
dates = pd.date_range('10-01-2016', periods=9, freq='2W-SUN')
dates
df = pd.DataFrame({'Count 1': 100 + np.random.randint(-5, 10, 9).cumsum(),
'Count 2': 120 + np.random.randint(-5, 10, 9)}, index=dates)
df
df.index.weekday_name
df.diff()
df.resample('M').mean()
df['2017']
df['2016-12']
df['2016-12':]
df.asfreq('W', method='ffill')
import matplotlib.pyplot as plt
%matplotlib inline
df.plot()
```
| github_jupyter |
# Introduction to the Insight Toolkit (ITK)
### Learning Objectives
* Learn how to **run** cells in **a Jupyter Notebook**
* Run a segmentation example that demonstrates **ITK**'s ability to provide **insight into images**
* Understand the **purpose and capabilities** of the toolkit
## Jupyter Notebooks
These are [Jupyter Notebooks](https://jupyter.org/), an open-source web application that allows you to create and share documents that contain live code, equations, visualizations and narrative text.
To run cells in the notebook, press *shift + enter*.
For more information, see the [Notebook Help](https://nbviewer.jupyter.org/github/ipython/ipython/blob/3.x/examples/Notebook/Index.ipynb)
## Insight Into Images
```
import itk
from itkwidgets import view
file_name = 'data/brainweb165a10f17.mha'
image = itk.imread(file_name, itk.ctype('float'))
view(image, slicing_planes=True, gradient_opacity=0.8, ui_collapsed=True)
# Smooth the image
smoothed = itk.curvature_flow_image_filter(image,
number_of_iterations=6,
time_step=0.005)
view(smoothed, slicing_planes=True, gradient_opacity=0.8, ui_collapsed=True)
# Segment the white matter with a 3D region-growing algorithm
confidence_connected = itk.ConfidenceConnectedImageFilter.New(smoothed)
confidence_connected.SetMultiplier(2.5)
confidence_connected.SetNumberOfIterations(5)
confidence_connected.SetInitialNeighborhoodRadius(2)
confidence_connected.SetReplaceValue(255)
confidence_connected.AddSeed([118, 133, 92])
confidence_connected.AddSeed([63, 135, 94])
confidence_connected.AddSeed([63, 157, 90])
confidence_connected.AddSeed([111, 150, 90])
confidence_connected.AddSeed([111, 50, 88])
confidence_connected.Update()
view(confidence_connected, ui_collapsed=True, cmap='BuPu', shadow=False, annotations=False)
```
## History of ITK
### Insight into Images

http://itk.org
### History
In 1999, the US National Institute of Health’s (NIH) National Library of Medicine (NLM) started a project to support the Visible Human Project.


### Goals
* Collect best-of-the-best image analysis algorithms for reproducible science.
* Provide a software resource for teaching medical image analysis algorithms.
* Establish a foundation for future research.
* Support commercial applications.
* Create conventions for future work.
* Grow a self-sustaining community of software users and developers.
### Continued Development
* Development has progressed since 1999
* Contributions from over 267 developers
* Over 1.6 million lines of code
* Downloaded over 500,000 times

ITK contributors locations for the 4.8 and 4.9 releases.
## Features
### N-Dimensional Image Filtering

### Filtering Algorithms Classes
* Fast marching methods
* Convolution
* Image gradient
* Denoising
* Thresholding
* Mathematical morphology
* Smoothing
* Image features
* Image statistics
* Bias correction
* Image grid operations
* ....
### Segmentation
"Image segmentation is the process of partitioning a digital image into multiple segments, i.e. sets of pixels. The goal of segmentation is to simplify and/or change the representation of an image into something that is more meaningful and easier to analyze."
Source: [Wikipedia](https://en.wikipedia.org/wiki/Image_segmentation)
### Segmentation

### Segmentation

### Segmentation

### Segmentation

### Segmentation

### Segmentation

### Registration
"Image registration is the process of transforming different sets of data into one coordinate system. [...] Registration is necessary in order to be able to compare or integrate the data obtained from these different measurements."
Source: [Wikipedia](https://en.wikipedia.org/wiki/Image_registration)
### Registration

### Other Data Structures
* Mesh's
* Transforms's
* SpatialObject's
* Path's
* LabelMap's


### ITK Resources
* ITK Software Guide: https://www.itk.org/ItkSoftwareGuide.pdf
* Discourse Discussion: https://discourse.itk.org
* Sphinx Examples: https://www.itk.org/ITKExamples
* `Examples/` directory in the ITK repository
* Wiki Examples: http://www.itk.org/Wiki/ITK/Examples
* Doxygen: http://www.itk.org/Doxygen/html/index.html
* Kitware: https://www.kitware.com/
### Kitware, Inc


- Collaborative, Open Source, Scientific Software Development
- Carrboro, NC, near University of North Carolina-Chapel Hill, in the Research Triangle
### Enjoy ITK!
| github_jupyter |
# Feature: "Jaccard with WHQ" (@dasolmar)
Based on the kernel [XGB with whq jaccard](https://www.kaggle.com/dasolmar/xgb-with-whq-jaccard/) by David Solis.
## Imports
This utility package imports `numpy`, `pandas`, `matplotlib` and a helper `kg` module into the root namespace.
```
from pygoose import *
```
NLTK tools
```
import nltk
from collections import Counter
from nltk.corpus import stopwords
nltk.download('stopwords')
```
## Config
Automatically discover the paths to various data folders and compose the project structure.
```
project = kg.Project.discover()
```
Identifier for storing these features on disk and referring to them later.
```
feature_list_id = '3rdparty_dasolmar_whq'
```
## Read data
Original question sets.
```
df_train = pd.read_csv(project.data_dir + 'train.csv').fillna('')
df_test = pd.read_csv(project.data_dir + 'test.csv').fillna('')
```
NLTK built-in stopwords.
```
stops = set(stopwords.words("english"))
```
## Build features
```
# If a word appears only once, we ignore it completely (likely a typo)
# Epsilon defines a smoothing constant, which makes the effect of extremely rare words smaller
def get_weight(count, eps=10000, min_count=2):
return 0 if count < min_count else 1 / (count + eps)
def add_word_count(x, df, word):
x['das_q1_' + word] = df['question1'].apply(lambda x: (word in str(x).lower())*1)
x['das_q2_' + word] = df['question2'].apply(lambda x: (word in str(x).lower())*1)
x['das_' + word + '_both'] = x['das_q1_' + word] * x['das_q2_' + word]
train_qs = pd.Series(df_train['question1'].tolist() + df_train['question2'].tolist()).astype(str)
words = (" ".join(train_qs)).lower().split()
counts = Counter(words)
weights = {word: get_weight(count) for word, count in counts.items()}
def word_shares(row):
q1_list = str(row['question1']).lower().split()
q1 = set(q1_list)
q1words = q1.difference(stops)
if len(q1words) == 0:
return '0:0:0:0:0:0:0:0'
q2_list = str(row['question2']).lower().split()
q2 = set(q2_list)
q2words = q2.difference(stops)
if len(q2words) == 0:
return '0:0:0:0:0:0:0:0'
words_hamming = sum(1 for i in zip(q1_list, q2_list) if i[0]==i[1])/max(len(q1_list), len(q2_list))
q1stops = q1.intersection(stops)
q2stops = q2.intersection(stops)
q1_2gram = set([i for i in zip(q1_list, q1_list[1:])])
q2_2gram = set([i for i in zip(q2_list, q2_list[1:])])
shared_2gram = q1_2gram.intersection(q2_2gram)
shared_words = q1words.intersection(q2words)
shared_weights = [weights.get(w, 0) for w in shared_words]
q1_weights = [weights.get(w, 0) for w in q1words]
q2_weights = [weights.get(w, 0) for w in q2words]
total_weights = q1_weights + q1_weights
R1 = np.sum(shared_weights) / np.sum(total_weights) #tfidf share
R2 = len(shared_words) / (len(q1words) + len(q2words) - len(shared_words)) #count share
R31 = len(q1stops) / len(q1words) #stops in q1
R32 = len(q2stops) / len(q2words) #stops in q2
Rcosine_denominator = (np.sqrt(np.dot(q1_weights,q1_weights))*np.sqrt(np.dot(q2_weights,q2_weights)))
Rcosine = np.dot(shared_weights, shared_weights)/Rcosine_denominator
if len(q1_2gram) + len(q2_2gram) == 0:
R2gram = 0
else:
R2gram = len(shared_2gram) / (len(q1_2gram) + len(q2_2gram))
return '{}:{}:{}:{}:{}:{}:{}:{}'.format(R1, R2, len(shared_words), R31, R32, R2gram, Rcosine, words_hamming)
df = pd.concat([df_train, df_test])
df['word_shares'] = df.apply(word_shares, axis=1, raw=True)
x = pd.DataFrame()
x['das_word_match'] = df['word_shares'].apply(lambda x: float(x.split(':')[0]))
x['das_word_match_2root'] = np.sqrt(x['das_word_match'])
x['das_tfidf_word_match'] = df['word_shares'].apply(lambda x: float(x.split(':')[1]))
x['das_shared_count'] = df['word_shares'].apply(lambda x: float(x.split(':')[2]))
x['das_stops1_ratio'] = df['word_shares'].apply(lambda x: float(x.split(':')[3]))
x['das_stops2_ratio'] = df['word_shares'].apply(lambda x: float(x.split(':')[4]))
x['das_shared_2gram'] = df['word_shares'].apply(lambda x: float(x.split(':')[5]))
x['das_cosine'] = df['word_shares'].apply(lambda x: float(x.split(':')[6]))
x['das_words_hamming'] = df['word_shares'].apply(lambda x: float(x.split(':')[7]))
x['das_diff_stops_r'] = np.abs(x['das_stops1_ratio'] - x['das_stops2_ratio'])
x['das_len_q1'] = df['question1'].apply(lambda x: len(str(x)))
x['das_len_q2'] = df['question2'].apply(lambda x: len(str(x)))
x['das_diff_len'] = np.abs(x['das_len_q1'] - x['das_len_q2'])
x['das_caps_count_q1'] = df['question1'].apply(lambda x:sum(1 for i in str(x) if i.isupper()))
x['das_caps_count_q2'] = df['question2'].apply(lambda x:sum(1 for i in str(x) if i.isupper()))
x['das_diff_caps'] = np.abs(x['das_caps_count_q1'] - x['das_caps_count_q2'])
x['das_len_char_q1'] = df['question1'].apply(lambda x: len(str(x).replace(' ', '')))
x['das_len_char_q2'] = df['question2'].apply(lambda x: len(str(x).replace(' ', '')))
x['das_diff_len_char'] = np.abs(x['das_len_char_q1'] - x['das_len_char_q2'])
x['das_len_word_q1'] = df['question1'].apply(lambda x: len(str(x).split()))
x['das_len_word_q2'] = df['question2'].apply(lambda x: len(str(x).split()))
x['das_diff_len_word'] = np.abs(x['das_len_word_q1'] - x['das_len_word_q2'])
x['das_avg_word_len1'] = x['das_len_char_q1'] / x['das_len_word_q1']
x['das_avg_word_len2'] = x['das_len_char_q2'] / x['das_len_word_q2']
x['das_diff_avg_word'] = np.abs(x['das_avg_word_len1'] - x['das_avg_word_len2'])
# x['exactly_same'] = (df['question1'] == df['question2']).astype(int)
# x['duplicated'] = df.duplicated(['question1','question2']).astype(int)
whq_words = ['how', 'what', 'which', 'who', 'where', 'when', 'why']
for whq in whq_words:
add_word_count(x, df, whq)
whq_columns_q1 = ['das_q1_' + whq for whq in whq_words]
whq_columns_q2 = ['das_q2_' + whq for whq in whq_words]
x['whq_count_q1'] = x[whq_columns_q1].sum(axis=1)
x['whq_count_q2'] = x[whq_columns_q2].sum(axis=1)
x['whq_count_diff'] = np.abs(x['whq_count_q1'] - x['whq_count_q2'])
feature_names = list(x.columns.values)
print("Features: {}".format(feature_names))
X_train = x[:df_train.shape[0]].values
X_test = x[df_train.shape[0]:].values
```
## Save features
```
project.save_features(X_train, X_test, feature_names, feature_list_id)
```
| github_jupyter |
# Classification
```
# import necessary libraries
import pandas as pd # for data input/output and processing
import numpy as np # for matrix calculations
from sklearn.linear_model import LogisticRegression # An already implemented version of logistic regression
from sklearn.preprocessing import PolynomialFeatures # To transform our features to the corresponding polynomial version
from sklearn.metrics import accuracy_score, log_loss, zero_one_loss # a way to evaluate how erroneous our model is
import matplotlib.pyplot as plt # For visualizations
plt.rcParams.update({'font.size': 20})
```
## Data Preprocessing
Preprocessing refers to some data manipulation techniques that are used to make sure that our data carry important information in the format that we expect it to.
Here we are going to use a small dataset of only 89 samples. These correspond to meteorological observations from the [Finnish Meteorological Institute (FMI)](https://en.ilmatieteenlaitos.fi/download-observations).
```
# First load the dataset into a pandas DataFrame
df = pd.read_csv("fmi_nov_dec_2021.csv")
print("Number of data points:", len(df))
df.head(2) # print the first 2 rows
```
### Datapoints and Features vs. Labels
- **Datapoint**: A datapoint is a single row of our dataset that represents an object with specific properties.
- **Label**: Some property (i.e. a column of our dataset) that is a quantity of interest.
- **Features**: All of our dataset's columns that are not the target. It represents the amount of information that we think we need in order to predict the label.
**FMI Example:**
- Our goal is to predict whether there is going to be a high or a low temperature.
- We will label as **high**, all temperatures above 5 degrees. The rest will be considered **low** temperatures.
- To make classification easier, the class **high** will be represented with the label **1** and the class **low** with **0**.
- We will **consider** the `day` and the `minimum temperature` as possible features.
- This way we get:
- **Datapoint**: Represents whether we had a high or low temperature at a given day and minimum temperature.
- **Label**: Temperature class - High or Low (1 or 0).
- **Features**: `Day`, `Minimum Temperature`
```
feature_columns = ["d", "Minimum temperature (degC)"]
label_column = ["Maximum temperature (degC)"]
# Keep only relevant columns
df = df[feature_columns+label_column]
# Rename the columns to make access easier
df.columns = ['d', 'min_temp', 'max_temp']
df.head(2)
```
We now need to process the `Maximum temperature (degC)` column and convert all values above $5$ to **1** and the rest to **0**.
```
# <new column> <`row` is a single entry of the df.max_temp Series>
df['label'] = df.max_temp.apply(lambda max_temp: 1 if max_temp > 5 else 0)
df.head(2)
# Drop the max_temp column since we no longer need it
df.drop(columns=['max_temp'], inplace=True)
# Drop NaN (Not A Number) values
df.dropna(inplace=True)
df.head(2)
```
Now let's visualize the correlation of the remaining columns with the target variable. For this, we can simply make a scatter plot, as follows:
```
# def plot_model(X, y,
fig=plt.figure(figsize=(16, 7))
fig.suptitle("Relation between the features and the maximum temperature")
plt.subplot(1, 2, 1)
plt.scatter(df.d, df.label, c="blue")
plt.xlabel("Day")
plt.ylabel("Max Temperature Classification")
plt.subplot(1, 2, 2)
plt.scatter(df.min_temp, df.label, c="red")
plt.xlabel("Min Temperature")
plt.ylabel("Max Temperature Classification")
fig.tight_layout() # Separate the two subplots
plt.show()
```
The `d` column seems to have a very noisy relationship with the label. On the other hand, high `min_temp` values seem to correspond to high temperatures, while lower `min_temp` values lead to lower maximum temperatures.
For this reason we are going to completely ignore the `d` column and move on with the `min_temp` as our only feature. Using 1 feature also allows for better visualizations in a 2d space (e.g. if we had two features, then we would need 3 axes in total, and 3d plots are not as intuitive as 2d plots. If we had 3 features, then visualization would be impossible).
```
df.drop(columns=['d'], inplace=True)
print(len(df))
df.head(2)
# Let's separate features from labels
labels = df.label.values # .values convert to a numpy array
features = df.drop(columns=['label']).values
```
## Logistic Regression
> Logistic regression is a ML method that allows to classify data points **according to two categories**. Thus, logistic regression is a **binary classification method** that can be applied to data points characterized by feature vectors $x \in \mathbf{R}^n$ (feature space $X = \mathbf{R}^n$) and binary labels $y$.
> These binary labels take on values from a label space that contains two different label values. Each of these two label values represents one of the two categories to which the data points can belong.
> Note that logistic regression uses the same hypothesis space as linear regression.
> We can always obtain a predicted label $\hat{y} ∈ {0, 1}$ by comparing hypothesis value $h(x)$ with a threshold. A data point with features $x$, is classified as $\hat{y} = 1 \text{ if } h(x) ≥ 0 \text{ and } \hat{y}=0 \text{ for } h(x) < 0$.
### Loss Function
Logistic regression tries to minimize the empirical risk (average logistic loss):
$$
\hat{L}(w|D) \stackrel{(*)}{=} \frac{1}{m} \sum_{i=1}^m log\left( 1+ exp\left( -y^{(i)} w^T x^{(i)}\right) \right)
$$
Our goal is to find the optimal vector $\hat{w}$ which minimizes the loss.
*(\*)The function $h$ is a linear combination of the input features: $h(x^{(i)}) = w_0 + \sum_{j=1}^{n} w_i x^{(i)}_j = w^T x^{(i)}$*
```
# Create a logistic regression model
lr = LogisticRegression()
# Fit the model to our data in order to get the most suitable parameters
lr = lr.fit(features, labels)
```
## Evaluation
- We are using the average logistic loss as a loss function which we try to minimize. The goal is to get its values as close to $0$ as possible.
- Even though "average logistic loss" is a very good loss function, it is not very intuitive and it cannot be used to compare different models.
- Instead, we opt to use the 0/1 loss, which is the opposite of the more commonly known "accuracy" ($0/1$ $loss = 1-accuracy$).
- Accuracy can be calculated as the ratio of correctly classified examples against the total number of examples.
```
# Calculate the mean square error on the training set
predictions = lr.predict(features)
zero_one_loss(predictions, labels) # ideally this would be 0 which would mean that the number of incorrect predictions is close to 0
log_loss(predictions, labels) # this doesn't really tell us anything.
plt.figure(figsize=(12,7))
# How good does the model fit our data?
plt.scatter(features, labels, c="blue", label="Datapoints")
# an increasing vector in the [min, max] range of the features
X_line = np.linspace(features.min(), features.max(), len(features)).reshape(-1, 1) # needs to be 2d and so we reshape
predictions = lr.predict_proba(X_line)[:, 1]
# plt.plot(X_line, predictions, c="red", label="Model")
plt.scatter(X_line, predictions, c='red', label="Predictions")
plt.xlabel("Minimum Temperature")
plt.ylabel("Maximum Temperature")
plt.title("Linear Model (min_temp -> max_temp)")
plt.legend()
plt.show()
```
## Increasing Model Size
Include terms of the feature values, raised to some power $r$ for some $r>1$:
$
\hat{h}(x) = w_0 + w_1 x + w_1 x^2 + ... + w_{r+1} x^r
$
To find the best value of $r$ we usually have to search through a lot of values, as follows:
```
def get_poly_predictor(features, labels, degree, **kwargs):
poly_features = PolynomialFeatures(degree=degree, include_bias=False)
# transform features so as to include their polynomial version.
# E.g. feas_new will now also include x^2, ..., x^{degree}
feas_new = poly_features.fit_transform(features)
lr = LogisticRegression(**kwargs)
lr.fit(feas_new, labels)
model_preds = lr.predict(feas_new)
zo = zero_one_loss(model_preds, labels)
ll = log_loss(model_preds, labels)
return zo, ll
# Try out different degrees and print the corresponding MSEs
for r in range(2, 8):
zo, ll = get_poly_predictor(features, labels, degree=r, max_iter=500)
print(f"Degree={r} -> 0/1 Loss={zo} -> Log Loss={ll}")
```
The errors decrease by a lot as we increase the degree. Does that mean that our model is better?
Let's make a scatter of the models for a degree of 2, 4, 6 and 7:
```
def poly_visualize(
features,
labels,
degrees=[2, 4, 6, 7],
colors=['blue', 'green', 'cyan', 'purple'],
**log_reg_kwargs,
):
fig = plt.figure(figsize=(22, 18))
wrong_poly_colors = ['gold', 'teal', 'tomato', 'firebrick', 'orchid']
for i, r in enumerate(degrees):
# ===================================================================
# Fit model
# ===================================================================
poly_features = PolynomialFeatures(degree=r, include_bias=False)
feas_new = poly_features.fit_transform(features)
lr = LogisticRegression(**log_reg_kwargs)
lr.fit(feas_new, labels)
# ===================================================================
# Scatter plot
# ===================================================================
plt.subplot(2, 2, i+1)
# How good does the model fit our data?
plt.scatter(features, labels, c=colors[i], label="Datapoints")
# an increasing vector in the [min, max] range of the features
X_line = np.linspace(features.min(), features.max(), 400).reshape(-1, 1) # needs to be 2d and so we reshape
predictions = lr.predict_proba(poly_features.transform(X_line))[:, 1]
plt.plot(X_line, predictions, c="red", label="Best Fitted Model", linewidth=7.0)
plt.xlabel("Minimum Temperature")
plt.ylabel("Maximum Temperature Classification")
zo = zero_one_loss(lr.predict(feas_new), labels)
plt.title(f"Degree {r} - 0/1 Loss={round(zo, 2)}")
plt.legend()
plt.show()
poly_visualize(features, labels, degrees=[2, 4], colors=["blue", "green"], max_iter=500)
poly_visualize(features, labels, degrees=[6, 7], colors=["cyan", "purple"], max_iter=500)
```
| github_jupyter |
# Wide vs. Long Format Data
## About the data
In this notebook, we will be using daily temperature data from the [National Centers for Environmental Information (NCEI) API](https://www.ncdc.noaa.gov/cdo-web/webservices/v2). We will use the Global Historical Climatology Network - Daily (GHCND) data set for the Boonton 1 station (GHCND:USC00280907); see the documentation [here](https://www1.ncdc.noaa.gov/pub/data/cdo/documentation/GHCND_documentation.pdf).
*Note: The NCEI is part of the National Oceanic and Atmospheric Administration (NOAA) and, as you can see from the URL for the API, this resource was created when the NCEI was called the NCDC. Should the URL for this resource change in the future, you can search for the NCEI weather API to find the updated one.*
```
import matplotlib.pyplot as plt
import pandas as pd
wide_df = pd.read_csv('data/wide_data.csv', parse_dates=['date'])
long_df = pd.read_csv(
'data/long_data.csv',
usecols=['date', 'datatype', 'value'],
parse_dates=['date']
)[['date', 'datatype', 'value']] # sort columns
```
## Wide format
Our variables each have their own column:
```
wide_df.head(6)
```
Describing all the columns is easy:
```
wide_df.describe(include='all')
```
Easy to graph with `pandas` (covered in [chapter 5](https://github.com/stefmolin/Hands-On-Data-Analysis-with-Pandas/tree/master/ch_05)):
```
wide_df.plot(
kind='line', y=['TMAX', 'TMIN', 'TOBS'], x='date',
title='Temperature in NYC in October 2018',
figsize=(15, 5)
).set_ylabel('Temperature in Celsius')
plt.show()
```
## Long format
Our variable names are now in the `datatype` column and their values are in the `value` column. We now have 3 rows for each date, since we have 3 different `datatypes`:
```
long_df.head(6)
```
Since we have many rows for the same date, using `describe()` is not trivial anymore:
```
long_df.describe(include='all')
```
Plotting long format data in `pandas` can be rather tricky. Instead we use `seaborn` (covered in [chapter 6](https://github.com/stefmolin/Hands-On-Data-Analysis-with-Pandas/blob/master/ch_06/1-introduction_to_seaborn.ipynb)):
```
import seaborn as sns
sns.set(rc={'figure.figsize':(15, 5)}, style='white')
ax = sns.lineplot(data=long_df, hue='datatype', y='value', x='date')
ax.set_ylabel('Temperature in Celsius')
ax.set_title('Temperature in NYC in October 2018')
plt.show()
```
With long data and `seaborn`, we can easily facet our plots:
```
sns.set(rc={'figure.figsize':(20, 10)}, style='white', font_scale=2)
g = sns.FacetGrid(long_df, col="datatype", height=10)
g = g.map(plt.plot, "date", "value")
g.set_titles(size=25)
g.set_xticklabels(rotation=45)
plt.show()
```
| github_jupyter |
# Exploratory Data Analysis
First pass at analysing the output of blasting AML RNA-Seq data against the three tryptase sequences from Jonathon.
```
import pysam
import glob
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
def get_files(folder):
return glob.glob('{folder}/*.filtered.bam'.format(folder=folder))
list_of_aml_results = get_files('../aml_results/')
list_of_control_results = get_files('../control_results/')
aml_file_names = [x.split('/')[len(x.split('/'))-1] for x in list_of_aml_results]
control_file_names = [x.split('/')[len(x.split('/'))-1] for x in list_of_control_results]
df_aml = pd.DataFrame(index=aml_file_names)
df_control = pd.DataFrame(index=control_file_names)
df_aml['file_location'] = list_of_aml_results
df_control['file_location'] = list_of_control_results
df_aml['source'] = 'AML'
df_control['source'] = 'control'
df_aml.head()
df_control.head()
df = df_aml.append(df_control)
df.count()
def get_read_count(df):
return int(pysam.view("-c", df['file_location']))
df['alignment_count'] = df.apply(get_read_count, axis=1)
df.head()
def get_transcript_count(df, transcript_name):
"""
Return the count of the transcript i.e how many of each of the three are in there.
Note - Does not look at the quality of the alignment.
"""
sam_file_location = df['file_location']
samfile = pysam.AlignmentFile(sam_file_location, "rb")
count = 0
for read in samfile:
if read is not None and read.reference_name == transcript_name:
count = count + 1
return count
df['alpha_wt_count'] = df.apply(get_transcript_count, axis=1, args=['Alpha_GEX_64k_HEX'])
df['alpha_dup_count'] = df.apply(get_transcript_count, axis=1, args=['Alpha_GEX_79k_dup_FAM'])
df['beta_count'] = df.apply(get_transcript_count, axis=1, args=['BETA_new_GEX_FAM'])
df.head()
def get_zero_edit_distance_count(df, transcript_name):
"""
For a goven transcript e.g. Alpha_GEX_64k_HEX get how many of the matches have an NM tag of zero
i.e. the match was exact.
"""
sam_file_location = df['file_location']
samfile = pysam.AlignmentFile(sam_file_location, "rb")
count = 0
for read in samfile:
if read is not None and read.reference_name == transcript_name:
edit_distance = read.get_tag('NM')
if edit_distance == 0:
count = count + 1
return count
df['alpha_wt_zero_edit_count'] = df.apply(get_zero_edit_distance_count, axis=1, args=['Alpha_GEX_64k_HEX'])
df['alpha_dup_zero_edit_count'] = df.apply(get_zero_edit_distance_count, axis=1, args=['Alpha_GEX_79k_dup_FAM'])
df['beta_zero_edit_count'] = df.apply(get_zero_edit_distance_count, axis=1, args=['BETA_new_GEX_FAM'])
df.head()
def get_transcript_read_count_filtered(df, transcript_name, start, end):
"""
Count hits which cover the bit of the reference we are interested in
"""
sam_file_location = df['file_location']
samfile = pysam.AlignmentFile(sam_file_location, "rb")
iter = samfile.fetch(transcript_name, start, end)
count =0
for read in iter:
if read.reference_start <=3 and read.reference_end >= 41:
count = count +1
return count
df['alpha_read_covers_snps_count'] = df.apply(get_transcript_read_count_filtered,
axis=1,
args=['Alpha_GEX_64k_HEX', 0,45])
df['alpha_dup_read_covers_snps_count'] = df.apply(get_transcript_read_count_filtered,
axis=1,
args=['Alpha_GEX_79k_dup_FAM', 0,45])
df['beta_read_covers_snps_count'] = df.apply(get_transcript_read_count_filtered,
axis=1,
args=['BETA_new_GEX_FAM', 0,45])
df.sort_values(by='alpha_dup_read_covers_snps_count', ascending=False).head()
def get_transcript_read_count_filtered_exact(df, transcript_name, start, end):
"""
Count hits which cover the bit of the reference we are interested in and which are exact.
That is do they cross position 0 - 44 of the transcript ( given by transcipt_name) and \
have an edit distance from the reference of 0.
Reads which cross this location will cross all four SNPs needed to tell the 3 transcripts apart.
"""
sam_file_location = df['file_location']
samfile = pysam.AlignmentFile(sam_file_location, "rb")
iter = samfile.fetch(transcript_name, start, end)
count =0
for read in iter:
if read.reference_start <=3 and read.reference_end >= 41:
edit_distance = read.get_tag('NM')
if edit_distance == 0:
count = count + 1
return count
df['alpha_read_covers_snps_count_exact'] = df.apply(get_transcript_read_count_filtered_exact,
axis=1,
args=['Alpha_GEX_64k_HEX', 0,45])
df['alpha_dup_read_covers_snps_count_exact'] = df.apply(get_transcript_read_count_filtered_exact,
axis=1,
args=['Alpha_GEX_79k_dup_FAM', 0,45])
df['beta_read_covers_snps_count_exact'] = df.apply(get_transcript_read_count_filtered_exact,
axis=1,
args=['BETA_new_GEX_FAM', 0,45])
# Quick check to see if any such reads exist
df.sort_values(by='alpha_dup_read_covers_snps_count_exact', ascending=False).head()
```
## Sanity Check
Print out some of the reads. I then did a manual BLAST and alignment to check the code was working as expected
```
def get_transcript_read_count_filtered_exact_print(sam_file_location, transcript_name, start, end):
"""
Same as get_transcript_read_count_filtered_exact() except we just print out the reads instead of \
counting them.
"""
samfile = pysam.AlignmentFile(sam_file_location, "rb")
iter = samfile.fetch(transcript_name, start, end)
for read in iter:
if read.reference_start <=3 and read.reference_end >= 41:
edit_distance = read.get_tag('NM')
if edit_distance == 0:
print (read)
#Pick a random bam file with some read alignments in this area
get_transcript_read_count_filtered_exact_print('../aml_results/aml_aabspliced_SRR5626188.sam.sorted.bam',
'Alpha_GEX_64k_HEX',
0,
45 )
df.describe()
aml_total_hits = df[df['source']=='AML'].sum(axis=0)
aml_total_hits
control_total_hits = df[df['source']=='control'].sum(axis=0)
control_total_hits
df.head()
def get_bases_at_pos(df, transcript, pos):
"""
For a given position in the reference what are the bases in the reads which align at \
that position.
NB - only counts bases that are properly aligned (see same flag 163) so the number of bases \
may be less than the count of reads aligned fully over the area.
"""
sam_file_location = df['file_location']
samfile = pysam.AlignmentFile(sam_file_location, "rb")
iter = samfile.pileup(contig=transcript)
for column in iter:
#print (column.reference_name, column.reference_pos)
if column.reference_pos == pos:
base_list = []
for read in column.pileups:
if read.is_del == 0:
base_list.append(read.alignment.query_sequence[read.query_position])
#print(read)
#print (read.query_position)
#print (read.alignment.query_sequence[read.query_position])
else:
base_list.append('_')
return ''.join(base_list)
df['alpha_wt_pos3_pileup'] = df.apply(get_bases_at_pos, axis=1, args=['Alpha_GEX_64k_HEX', 3])
df['alpha_dp_pos3_pileup'] = df.apply(get_bases_at_pos, axis=1, args=['Alpha_GEX_79k_dup_FAM', 3])
df['beta_pos3_pileup'] = df.apply(get_bases_at_pos, axis=1, args=['BETA_new_GEX_FAM', 3])
df.head(5)
df['alpha_wt_pos25_pileup'] = df.apply(get_bases_at_pos, axis=1, args=['Alpha_GEX_64k_HEX', 25])
df['alpha_dp_pos25_pileup'] = df.apply(get_bases_at_pos, axis=1, args=['Alpha_GEX_79k_dup_FAM', 25])
df['beta_pos25_pileup'] = df.apply(get_bases_at_pos, axis=1, args=['BETA_new_GEX_FAM', 25])
df['alpha_wt_pos40_pileup'] = df.apply(get_bases_at_pos, axis=1, args=['Alpha_GEX_64k_HEX', 40])
df['alpha_dp_pos40_pileup'] = df.apply(get_bases_at_pos, axis=1, args=['Alpha_GEX_79k_dup_FAM', 40])
df['beta_pos40_pileup'] = df.apply(get_bases_at_pos, axis=1, args=['BETA_new_GEX_FAM', 40])
df['alpha_wt_pos41_pileup'] = df.apply(get_bases_at_pos, axis=1, args=['Alpha_GEX_64k_HEX', 41])
df['alpha_dp_pos41_pileup'] = df.apply(get_bases_at_pos, axis=1, args=['Alpha_GEX_79k_dup_FAM', 41])
df['beta_pos41_pileup'] = df.apply(get_bases_at_pos, axis=1, args=['BETA_new_GEX_FAM', 41])
df.head(50)
df[['file_location', 'source','alpha_wt_pos3_pileup',
'alpha_dp_pos3_pileup', 'alpha_wt_pos25_pileup',
'alpha_dp_pos25_pileup', 'alpha_wt_pos40_pileup',
'alpha_dp_pos40_pileup', 'alpha_wt_pos41_pileup',
'alpha_dp_pos41_pileup']].head(50)
def combine_pileups(df,columns):
final_string = ''
for column in columns:
if df[column] is not None:
final_string = final_string + df[column]
return final_string
df['pos3_pileup'] = df.apply(combine_pileups, axis=1, args=[['alpha_wt_pos3_pileup','alpha_dp_pos3_pileup']])
df['pos25_pileup'] = df.apply(combine_pileups, axis=1, args=[['alpha_wt_pos25_pileup','alpha_dp_pos25_pileup']])
df['pos40_pileup'] = df.apply(combine_pileups, axis=1, args=[['alpha_wt_pos40_pileup','alpha_dp_pos40_pileup']])
df['pos41_pileup'] = df.apply(combine_pileups, axis=1, args=[['alpha_wt_pos41_pileup','alpha_wt_pos41_pileup']])
df[['file_location', 'source','pos3_pileup','pos25_pileup','pos40_pileup', 'pos41_pileup']].tail()
def max_base(df, column):
base_dict = {}
for base in df[column]:
if base not in base_dict:
base_dict[base]=1
else:
base_dict[base] = base_dict[base] +1
total_count = sum(base_dict.values()
return base_dict
df['pos3_call'] = df.apply(max_base, axis=1, args=['pos3_pileup'])
df['pos25_call'] = df.apply(max_base, axis=1, args=['pos25_pileup'])
df['pos40_call'] = df.apply(max_base, axis=1, args=['pos40_pileup'])
df['pos41_call'] = df.apply(max_base, axis=1, args=['pos41_pileup'])
df[['file_location', 'source','pos3_call','pos25_call','pos40_call','pos41_call']].head()
```
| github_jupyter |
# DIET PROBLEM - PYOMO
*Zuria Bauer Hartwig and David Senén García Hurtado* ( [CAChemE](http://cacheme.org))
Original Problem: [Linear and Integer Programming](https://www.coursera.org/course/linearprogramming) (Coursera Course) - University of Colorado Boulder & University of Colorado System
Based on the Examples from the Optimization Course = [Taller-Optimizacion-Python-Pyomo](https://github.com/CAChemE/Taller-Optimizacion-Python-Pyomo) from [CAChemE.org](http://cacheme.org/optimizacion-programacion-matematica-con-python-pyomo/)
[Neos Guide - The Diet Problem](http://www.neos-guide.org/content/diet-problem)
### SUMMARY
The goal of the diet problem is to select a set of foods we can get at McDonaldsx,
that will satisfy a set of daily nutritional requirement at minimum cost.
The problem is formulated as a linear program where the objective is to minimize cost
and the constraints are to satisfy the specified nutritional requirements.
The diet problem constraints typically regulate the number of calories,
the calories from Fat, Total Fat, Cholesterol, Sodium,
Carbohydrates, Fiber, Sugar, Proteine, Vitamine A and C,
Calcium and Sodium in the diet.
While the mathematical formulation is simple,
the solution may not be palatable!
The nutritional requirements can be met without regard for taste or variety,
so consider the output before digging into a meal from an "optimal" menu!
#### 1. Import
```
# Loading pyomo
from pyomo.environ import *
infinity = float('inf')
```
#### 2. Data
Let's get the data:
- http://nutrition.mcdonalds.com/getnutrition/nutritionfacts.pdf
- http://www.fastfoodmenuprices.com/mcdonalds-prices/
```
#Data
!cat data1.dat
#To show the data we can use !cat (or !type for windows) commands
```
### Solving
#### 3. Model
```
# Creation of a Concrete Model
model = AbstractModel()
```
#### 4. Sets
```
#DEFINE SETS
# Products
model.F = Set()
# Nutrients
model.N = Set()
```
#### 5. Parameters
```
# DEFINE PARAMETERS
# Calories
model.Cal = Param(model.F, within = PositiveReals)
# Cost
model.a = Param(model.F, model.N, within = NonNegativeReals)
# Max and Min Cost
model.Cmin = Param(model.N, within = NonNegativeReals, default = 0.0)
model.Cmax = Param(model.N, within = NonNegativeReals, default = infinity)
```
#### 6. Variables
```
# Number of servings
model.x = Var(model.F, within = NonNegativeIntegers)
```
#### 7. Objective
```
# MAXIMIZE z(calories)
def calories(model):
return sum(model.Cal[i] * model.x[i] for i in model.F)
model.calories = Objective(rule=calories, sense=maximize)
```
#### 8. Constrains
```
#CONSTRAINS
#Max
def cost_max(model, j):
value = sum(model.a[i,j] * model.x[i] for i in model.F)
return value <= model.Cmax[j]
model.cost_limit_max = Constraint(model.N, rule=cost_max)
#Min
def cost_min(model, j):
value = sum(model.a[i,j] * model.x[i] for i in model.F)
return model.Cmin[j] <= value
model.cost_limit_min = Constraint(model.N, rule=cost_min)
```
#### 9. Solution
```
!cat code1.py
#Get our Solution:
!pyomo solve --solver=glpk code1.py data1.dat
```
#### 10. Results
```
#Results
!cat results.yml
```
##### Results
- Bacon Buffalo Ranch McChickenx = 1€
- Hamburguerx = 1.29€
| github_jupyter |
```
import requests
import geopandas as gpd
```
# How to Read, Filter, and Convert a Shapefile to .geojson
Many public entities provide [shapefiles](https://en.wikipedia.org/wiki/Shapefile) to represent the borders of geographic areas. The shapefiles are often provided for larger geographic areas than we're interested in using. This notebook shows you how to start from a larger shapefile (Illinois), filter the data for a smaller region (Cook County), and export the data as a [`.geojson`](https://en.wikipedia.org/wiki/GeoJSON) file, an increasingly common and open source data format for geographic representation.
First, we start by downloading the Illinois shapefile from the Census website.
```
il_shp_url = 'https://www2.census.gov/geo/tiger/GENZ2010/gz_2010_17_140_00_500k.zip'
req = requests.get(il_shp_url)
file_path = './gz_2010_17_140_00_500k.zip'
with open(file_path, 'wb') as f:
f.write(req.content)
```
Unzip the downloaded shapefile data.
```
! unzip -o ./gz_2010_17_140_00_500k.zip -d ./gz_2010_17_140_00_500k
```
Next, using [`geopandas.read_file()`](http://geopandas.org/reference/geopandas.read_file.html), we read in the shapefile and plot it.
```
il_shp = gpd.read_file('./gz_2010_17_140_00_500k/gz_2010_17_140_00_500k.shp')
il_shp.plot(figsize=(8, 8))
```
You'll notice that Illinois looks a little distorted. That's because we need to assign to it a different [CRS (Coordinate Reference System)](https://en.wikipedia.org/wiki/Spatial_reference_system). Once we've assigned it an appropriate CRS, it will not appear distorted.
```
il_shp = il_shp.to_crs(epsg=3528)
il_shp.plot(figsize=(8,8))
```
In addition to being able to plot the shapefile data, it is also representable in table form as shown below.
```
il_shp.head()
```
In order to get the subset of tracts for Cook County, we'll need to filter the dataset for the rows where **COUNTY** is **031**.
```
cook_county_tracts = il_shp[il_shp['COUNTY'] == '031']
cook_county_tracts.plot(figsize = (8,8))
```
Finally, we can export the resulting filtered data to a `.geojson` file.
```
cook_county_tracts.to_file('./cook_county_tract_level.geojson', driver='GeoJSON')
```
**NOTE**: If the above command fails, you might be using `geopandas < 0.5`. You can either upgrade or use the following command before writing to file.
```
from shapely.geometry import MultiPolygon, Polygon
cook_county_tracts.set_geometry(cook_county_tracts.geometry.apply(lambda x : MultiPolygon([x]) if x.type == "Polygon" else x), inplace = True)
```
| github_jupyter |
# CrossCorrelation
This Tutorial is intended to give a demostration of How to make a CrossCorrelation Object in Stingray Library.
```
import numpy as np
from stingray import Lightcurve
from stingray.crosscorrelation import CrossCorrelation
import matplotlib.pyplot as plt
import matplotlib.font_manager as font_manager
%matplotlib inline
font_prop = font_manager.FontProperties(size=16)
```
# CrossCorrelation Example
# 1. Create two light curves
There are two ways to create a Lightcurve.<br>
1) Using an array of time stamps and an array of counts.<br>
2) From the Photon Arrival times.
In this example, Lightcurve is created using arrays of time stamps and counts.
Generate an array of relative timestamps that's 10 seconds long, with dt = 0.03125 s, and make two signals in units of counts. The signal is a sine wave with amplitude = 300 cts/s, frequency = 2 Hz, phase offset of pi/2 radians, and mean = 1000 cts/s. We then add Poisson noise to the light curve.
```
dt = 0.03125 # seconds
exposure = 10. # seconds
freq = 1 # Hz
times = np.arange(0, exposure, dt) # seconds
signal_1 = 300 * np.sin(2.*np.pi*freq*times) + 1000 # counts/s
signal_2 = 300 * np.sin(2.*np.pi*freq*times + np.pi/2) + 1000 # counts/s
noisy_1 = np.random.poisson(signal_1*dt) # counts
noisy_2 = np.random.poisson(signal_2*dt) # counts
```
Now let's turn noisy_1 and noisy_2 into Lightcurve objects. This way we have two Lightcurves to calculate CrossCorrelation.
```
lc1 = Lightcurve(times, noisy_1)
lc2 = Lightcurve(times, noisy_2)
len(lc1)
fig, ax = plt.subplots(1,1,figsize=(10,6))
ax.plot(lc1.time, lc1.counts, lw=2, color='blue')
ax.plot(lc1.time, lc2.counts, lw=2, color='red')
ax.set_xlabel("Time (s)", fontproperties=font_prop)
ax.set_ylabel("Counts (cts)", fontproperties=font_prop)
ax.tick_params(axis='x', labelsize=16)
ax.tick_params(axis='y', labelsize=16)
ax.tick_params(which='major', width=1.5, length=7)
ax.tick_params(which='minor', width=1.5, length=4)
plt.show()
```
# 2. Create a CrossCorrelation Object from two Light curves created above
To create a CrossCorrelation Object from LightCurves, simply pass both Lightvurves created above into the CrossCorrelation.
```
cr = CrossCorrelation(lc1, lc2)
```
Now, Cross Correlation values are stored in attribute corr, which is called below.
```
cr.corr[:10]
# Time Resolution for Cross Correlation is same as that of each of the Lightcurves
cr.dt
```
# 3. Plot Cross Correlation for Different lags
To visulaize correlation for different values of time lags, simply call plot function on cs.
```
cr.plot(labels = ['Time Lags (seconds)','Correlation'])
```
Given the Phase offset of pi/2 between two lightcurves created above, and freq=1 Hz, `time_shift` should be close to 0.25 sec. Small error is due to time resolution.
```
cr.time_shift #seconds
```
## Modes of Correlation
You can also specify an optional argument on modes of cross-correlation. <br>
There are three modes : 1) same 2) valid 3) full
Visit following ink on more details on mode : https://docs.scipy.org/doc/scipy-0.18.1/reference/generated/scipy.signal.correlate.html
Default mode is 'same' and it gives output equal to the size of larger lightcurve and is most common in astronomy. You can see mode of your CrossCorrelation by calling mode attribute on the object.
```
cr.mode
```
The number of data points in corr and largest lightcurve are same in this mode.
```
cr.n
```
Creating CrossCorrelation with full mode now using same data as above.
```
cr1 = CrossCorrelation(lc1, lc2, mode = 'full')
cr1.plot()
cr1.mode
```
Full mode does a full cross-correlation.
```
cr1.n
```
## Another Example
You can also create CrossCorrelation Object by using Cross Correlation data. This can be useful in some cases when you have correlation data and want to calculate time shift for max. correlation. You need to specify time resolution for correlation(default value of 1.0 seconds is used otherwise).
```
cs = CrossCorrelation()
cs.corr = np.array([ 660, 1790, 3026, 4019, 5164, 6647, 8105, 7023, 6012, 5162])
time_shift, time_lags, n = cs.cal_timeshift(dt=0.5)
time_shift
cs.plot( ['Time Lags (seconds)','Correlation'])
```
## Yet another Example with longer Lingcurve
I will be using same lightcurves as in the example above but with much longer duration and shorter lags.<br>
Both Lightcurves are chosen to be more or less same with a certain phase shift to demonstrate Correlation in a better way.
Again Generating two signals this time without poission noise so that time lag can be demonstrated. For noisy lightcurves, accurate calculation requires interpolation.
```
dt = 0.0001 # seconds
exposure = 50. # seconds
freq = 1 # Hz
times = np.arange(0, exposure, dt) # seconds
signal_1 = 300 * np.sin(2.*np.pi*freq*times) + 1000 * dt # counts/s
signal_2 = 200 * np.sin(2.*np.pi*freq*times + np.pi/2) + 900 * dt # counts/s
```
Converting noisy signals into Lightcurves.
```
lc1 = Lightcurve(times, signal_1)
lc2 = Lightcurve(times, signal_2)
len(lc1)
fig, ax = plt.subplots(1,1,figsize=(10,6))
ax.plot(lc1.time, lc1.counts, lw=2, color='blue')
ax.plot(lc1.time, lc2.counts, lw=2, color='red')
ax.set_xlabel("Time (s)", fontproperties=font_prop)
ax.set_ylabel("Counts (cts)", fontproperties=font_prop)
ax.tick_params(axis='x', labelsize=16)
ax.tick_params(axis='y', labelsize=16)
ax.tick_params(which='major', width=1.5, length=7)
ax.tick_params(which='minor', width=1.5, length=4)
plt.show()
```
Now, creating CrossCorrelation Object by passing lc1 and lc2 into the constructor.
```
cs = CrossCorrelation(lc1, lc2)
print('Done')
cs.corr[:50]
# Time Resolution for Cross Correlation is same as that of each of the Lightcurves
cs.dt
cs.plot( ['Time Lags (seconds)','Correlation'])
cs.time_shift #seconds
```
`time_shift` is very close to 0.25 sec, in this case.
## AutoCorrelation
Stingray has also separate class for AutoCorrelation. AutoCorrealtion is similar to crosscorrelation but involves only One Lightcurve.i.e. Correlation of Lightcurve with itself.
AutoCorrelation is part of `stingray.crosscorrelation` module. Following line imports AutoCorrelation.
```
from stingray.crosscorrelation import AutoCorrelation
```
To create `AutoCorrelation` object, simply pass lightcurve into AutoCorrelation Constructor.<br> Using same Lighrcurve created above to demonstrate `AutoCorrelation`.
```
lc = lc1
ac = AutoCorrelation(lc)
ac.n
ac.corr[:10]
ac.time_lags
```
`time_Shift` for `AutoCorrelation` is always zero. Since signals are maximally correlated at zero lag.
```
ac.time_shift
ac.plot()
```
# Another Example
Another example is demonstrated using a `Lightcurve` with Poisson Noise.
```
dt = 0.001 # seconds
exposure = 20. # seconds
freq = 1 # Hz
times = np.arange(0, exposure, dt) # seconds
signal_1 = 300 * np.sin(2.*np.pi*freq*times) + 1000 # counts/s
noisy_1 = np.random.poisson(signal_1*dt) # counts
lc = Lightcurve(times, noisy_1)
```
`AutoCorrelation` also supports `{full,same,valid}` modes similar to `CrossCorrelation`
```
ac = AutoCorrelation(lc, mode = 'full')
ac.corr
ac.time_lags
ac.time_shift
ac.plot()
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import random
import json
import time
import datetime
import os
from torch.utils.data import TensorDataset, random_split
from torch.utils.data import DataLoader, RandomSampler, SequentialSampler
from transformers import BertForSequenceClassification, AdamW, BertConfig
import torch
from transformers import BertTokenizer
from transformers import get_linear_schedule_with_warmup
ADDITIONAL_SPECIAL_TOKENS = ["<e1>", "</e1>", "<e2>", "</e2>"]
data_fname = './data/train_relation_candidates.csv'
df = pd.read_csv(data_fname)
print('train data size: ', len(df))
df.head()
df.type.value_counts()
label_map_dic = {
'Association': 0,
'Positive_Correlation': 1,
'Negative_Correlation': 2,
'Bind': 3,
'Cotreatment': 4,
'Comparison':5,
'Drug_Interaction':6,
'Conversion':7,
'Negative':8
}
df['label'] = df['type'].map(label_map_dic)
df.head()
# Instantiate the Bert tokenizer
# biobert
pretrained_fname = 'dmis-lab/biobert-base-cased-v1.1'
tokenizer = BertTokenizer.from_pretrained(pretrained_fname, do_lower_case=False)
tokenizer.add_special_tokens({"additional_special_tokens": ADDITIONAL_SPECIAL_TOKENS})
print('information about tokenizer: \n', tokenizer)
def get_inputs(df):
input_ids = []
attention_masks = []
labels = []
#e_mask = []
e1_mask_list = []
e2_mask_list = []
MAX_LENGTH = 128
# For every sentence...
for i,row in df.iterrows():
sent = row['sentences'] + str('[SEP]') + row['entity_1_mention'] + str('[SEP]') + row['entity_2_mention']
#sent = sent[:124]
labels.append(int(row['label']))
encoded_dict = tokenizer.encode_plus(
sent, # Sentence to encode.
add_special_tokens = True, # Add '[CLS]' and '[SEP]'
max_length = MAX_LENGTH, # Pad & truncate all sentences.
pad_to_max_length = True,
return_attention_mask = True, # Construct attn. masks.
return_tensors = 'pt', # Return pytorch tensors.
)
# Add the encoded sentence to the list.
input_ids.append(encoded_dict['input_ids'])
# And its attention mask (simply differentiates padding from non-padding).
attention_masks.append(encoded_dict['attention_mask'])
#print(encoded_dict['attention_mask'])
# e1 mask, e2 mask
e1_mask = torch.zeros_like(encoded_dict['attention_mask'])
e2_mask = torch.zeros_like(encoded_dict['attention_mask'])
tokens_a = tokenizer.tokenize(sent)
e11_p = tokens_a.index('<e1>') # the start position of entity1
e12_p = tokens_a.index('</e1>') # the end position of entity1
e21_p = tokens_a.index('<e2>') # the start position of entity2
e22_p = tokens_a.index('</e2>') # the end position of entity2
for i in range(e11_p, e12_p + 1):
if i < 128:
e1_mask[0][i] = 1
else:
continue
for i in range(e21_p, e22_p + 1):
if i < 128:
e2_mask[0][i] = 1
else:
continue
#print(e1_mask)
e1_mask_list.append(e1_mask)
e2_mask_list.append(e2_mask)
#print(attention_masks)
#print(e1_mask_list)
# Convert the lists into tensors.
input_ids = torch.cat(input_ids, dim=0)
attention_masks = torch.cat(attention_masks, dim=0)
labels = torch.tensor(labels)
e1_mask_list = torch.cat(e1_mask_list, dim=0)
e2_mask_list = torch.cat(e2_mask_list, dim=0)
return input_ids,attention_masks,labels,e1_mask_list,e2_mask_list
kfold = 0
train_df = df[df.kfold != kfold].reset_index(drop=True)
valid_df = df[df.kfold == kfold].reset_index(drop=True)
train_input_ids, train_attention_masks, train_labels, train_e1_mask, train_e2_mask = get_inputs(train_df)
valid_input_ids, valid_attention_masks, valid_labels, valid_e1_mask, valid_e2_mask = get_inputs(valid_df)
train_dataset = TensorDataset(train_input_ids, train_attention_masks, train_labels,train_e1_mask, train_e2_mask)
val_dataset = TensorDataset(valid_input_ids, valid_attention_masks, valid_labels,valid_e1_mask, valid_e2_mask)
# The DataLoader needs to know our batch size for training, so we specify it
# here. For fine-tuning BERT on a specific task, the authors recommend a batch
# size of 16 or 32.
batch_size = 32
# Create the DataLoaders for our training and validation sets.
# We'll take training samples in random order.
train_dataloader = DataLoader(
train_dataset, # The training samples.
sampler = RandomSampler(train_dataset), # Select batches randomly
batch_size = batch_size # Trains with this batch size.
)
# For validation the order doesn't matter, so we'll just read them sequentially.
validation_dataloader = DataLoader(
val_dataset, # The validation samples.
sampler = SequentialSampler(val_dataset), # Pull out batches sequentially.
batch_size = batch_size # Evaluate with this batch size.
)
model = BertForSequenceClassification.from_pretrained(
'dmis-lab/biobert-base-cased-v1.1',
num_labels = len(label_map_dic), # The number of output labels. 2 for binary classification.
output_attentions = False,
output_hidden_states = False)
# after add special token, we should resize model size
model.resize_token_embeddings(len(tokenizer))
# Tell pytorch to run this model on the GPU.
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# send model to device
model.to(device);
optimizer = AdamW(model.parameters(),
lr = 5e-5, # args.learning_rate - default is 5e-5
eps = 1e-8 # args.adam_epsilon - default is 1e-8.
)
# Number of training epochs. The BERT authors recommend between 2 and 4.
# We chose to run for 2,I have already seen that the model starts overfitting beyound 2 epochs
epochs = 10
# Total number of training steps is [number of batches] x [number of epochs].
total_steps = len(train_dataloader) * epochs
# Create the learning rate scheduler.
scheduler = get_linear_schedule_with_warmup(optimizer,
num_warmup_steps = 0, # Default value in run_glue.py
num_training_steps = total_steps)
# Function to calculate the accuracy of our predictions vs labels
def flat_accuracy(preds, labels):
pred_flat = np.argmax(preds, axis=1).flatten()
labels_flat = labels.flatten()
return np.sum(pred_flat == labels_flat) / len(labels_flat)
def format_time(elapsed):
'''
Takes a time in seconds and retur`````
ns a string hh:mm:ss
'''
# Round to the nearest second.
elapsed_rounded = int(round((elapsed)))
# Format as hh:mm:ss
return str(datetime.timedelta(seconds=elapsed_rounded))
# Set the seed value all over the place to make this reproducible.
seed_val = 42
random.seed(seed_val)
np.random.seed(seed_val)
torch.manual_seed(seed_val)
torch.cuda.manual_seed_all(seed_val)
# store a number of quantities such as training and validation loss,validation accuracy, and timings.
training_stats = []
# Measure the total training time for the whole run.
total_t0 = time.time()
# For each epoch...
for epoch_i in range(0, epochs):
# training
print('======== Epoch {:} / {:} ========'.format(epoch_i + 1, epochs))
print('Training...')
# Measure how long the training epoch takes.
t0 = time.time()
# Reset the total loss for this epoch.
total_train_loss = 0
# Put the model into training mode.
model.train()
# For each batch of training data...
for step, batch in enumerate(train_dataloader):
# Progress update every 40 batches.
if step % 40 == 0 and not step == 0:
# Calculate elapsed time in minutes.
elapsed = format_time(time.time() - t0)
# Report progress.
print(' Batch {:>5,} of {:>5,}. Elapsed: {:}.'.format(step, len(train_dataloader), elapsed))
# Unpack this training batch from our dataloader andcopy each tensor to the GPU
# `batch` contains three pytorch tensors:
# [0]: input ids
# [1]: attention masks
# [2]: labels
b_input_ids = batch[0].to(device)
b_input_mask = batch[1].to(device)
b_labels = batch[2].to(device)
# Always clear any previously calculated gradients before performing abackward pass.
model.zero_grad()
# Perform a forward pass (evaluate the model on this training batch).
outputs = model(b_input_ids,
token_type_ids=None,
attention_mask=b_input_mask,
labels=b_labels)
# Accumulate the training loss over all of the batches
loss= outputs[0]
logits = outputs[1]
total_train_loss += loss.item()
# Perform a backward pass to calculate the gradients.
loss.backward()
# Clip the norm of the gradients to 1.0.
# This is to help prevent the "exploding gradients" problem.
torch.nn.utils.clip_grad_norm_(model.parameters(), 1.0)
# Update parameters and take a step using the computed gradient.
optimizer.step()
# Update the learning rate.
scheduler.step()
# Calculate the average loss over all of the batches.
avg_train_loss = total_train_loss / len(train_dataloader)
# Measure how long this epoch took.
training_time = format_time(time.time() - t0)
print("")
print(" Average training loss: {0:.4f}".format(avg_train_loss))
print(" Training epoch took: {:}".format(training_time))
# ========================================
# Save model of every epoch
output_dir = './outputs/biobert_models/'
# Create output directory if needed
if not os.path.exists(output_dir):
os.makedirs(output_dir)
models_dir = output_dir + 'model_epoch_' + str(epoch_i + 1)
# Create output directory if needed
if not os.path.exists(models_dir):
os.makedirs(models_dir)
print("Saving model to %s" % models_dir)
# Save a trained model, configuration and tokenizer using `save_pretrained()`.
# They can then be reloaded using `from_pretrained()`
model_to_save = model.module if hasattr(model, 'module') else model # Take care of distributed/parallel training
model_to_save.save_pretrained(models_dir)
tokenizer.save_pretrained(models_dir)
# ========================================
# Validation
# ========================================
# After the completion of each training epoch, measure our performance on
# our validation set.
print("")
print("Running Validation...")
t0 = time.time()
# Put the model in evaluation mode--the dropout layers behave differently
# during evaluation.
model.eval()
# Tracking variables
total_eval_accuracy = 0.0
total_eval_loss = 0.0
nb_eval_steps = 0.0
# Evaluate data for one epoch
for batch in validation_dataloader:
b_input_ids = batch[0].to(device)
b_input_mask = batch[1].to(device)
b_labels = batch[2].to(device)
with torch.no_grad():
outputs= model(b_input_ids,
token_type_ids=None,
attention_mask=b_input_mask,
labels=b_labels)
# Accumulate the validation loss.
loss= outputs[0]
logits = outputs[1]
total_eval_loss += loss.item()
# Move logits and labels to CPU
logits = logits.detach().cpu().numpy()
label_ids = b_labels.to('cpu').numpy()
# Calculate the accuracy for this batch of test sentences, and
# accumulate it over all batches.
total_eval_accuracy += flat_accuracy(logits, label_ids)
# Report the final accuracy for this validation run.
avg_val_accuracy = total_eval_accuracy / len(validation_dataloader)
print(" Accuracy: {0:.4f}".format(avg_val_accuracy))
# Calculate the average loss over all of the batches.
avg_val_loss = total_eval_loss / len(validation_dataloader)
# Measure how long the validation run took.
validation_time = format_time(time.time() - t0)
print(" Validation Loss: {0:.4f}".format(avg_val_loss))
print(" Validation took: {:}".format(validation_time))
# Record all statistics from this epoch.
training_stats.append(
{
'epoch': epoch_i + 1,
'Training Loss': avg_train_loss,
'Valid. Loss': avg_val_loss,
'Valid. Accur.': avg_val_accuracy,
'Training Time': training_time,
'Validation Time': validation_time
}
)
print("")
print("Training complete!")
print("Total training took {:} (h:mm:ss)".format(format_time(time.time()-total_t0)))
import os
# Display floats with two decimal places.
pd.set_option('precision', 4)
# Create a DataFrame from our training statistics.
df_stats = pd.DataFrame(data=training_stats)
# Use the 'epoch' as the row index.
df_stats = df_stats.set_index('epoch')
output_dir = './outputs/biobert_models/'
# # Create output directory if needed
# if not os.path.exists(output_dir):
# os.makedirs(output_dir)
df_stats.to_csv(output_dir + 'df_train_stats.csv',index=None)
# according df_stats, pick the best performace model
df_stats
```
| github_jupyter |
# Language Translation
In this project, you’re going to take a peek into the realm of neural network machine translation. You’ll be training a sequence to sequence model on a dataset of English and French sentences that can translate new sentences from English to French.
## Get the Data
Since translating the whole language of English to French will take lots of time to train, we have provided you with a small portion of the English corpus.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
import helper
import problem_unittests as tests
source_path = 'data/small_vocab_en'
target_path = 'data/small_vocab_fr'
source_text = helper.load_data(source_path)
target_text = helper.load_data(target_path)
```
## Explore the Data
Play around with view_sentence_range to view different parts of the data.
```
view_sentence_range = (0, 10)
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
import numpy as np
print('Dataset Stats')
print('Roughly the number of unique words: {}'.format(len({word: None for word in source_text.split()})))
sentences = source_text.split('\n')
word_counts = [len(sentence.split()) for sentence in sentences]
print('Number of sentences: {}'.format(len(sentences)))
print('Average number of words in a sentence: {}'.format(np.average(word_counts)))
print()
print('English sentences {} to {}:'.format(*view_sentence_range))
print('\n'.join(source_text.split('\n')[view_sentence_range[0]:view_sentence_range[1]]))
print()
print('French sentences {} to {}:'.format(*view_sentence_range))
print('\n'.join(target_text.split('\n')[view_sentence_range[0]:view_sentence_range[1]]))
```
## Implement Preprocessing Function
### Text to Word Ids
As you did with other RNNs, you must turn the text into a number so the computer can understand it. In the function `text_to_ids()`, you'll turn `source_text` and `target_text` from words to ids. However, you need to add the `<EOS>` word id at the end of each sentence from `target_text`. This will help the neural network predict when the sentence should end.
You can get the `<EOS>` word id by doing:
```python
target_vocab_to_int['<EOS>']
```
You can get other word ids using `source_vocab_to_int` and `target_vocab_to_int`.
```
def text_to_ids(source_text, target_text, source_vocab_to_int, target_vocab_to_int):
"""
Convert source and target text to proper word ids
:param source_text: String that contains all the source text.
:param target_text: String that contains all the target text.
:param source_vocab_to_int: Dictionary to go from the source words to an id
:param target_vocab_to_int: Dictionary to go from the target words to an id
:return: A tuple of lists (source_id_text, target_id_text)
"""
# TODO: Implement Function
return None, None
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_text_to_ids(text_to_ids)
```
### Preprocess all the data and save it
Running the code cell below will preprocess all the data and save it to file.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
helper.preprocess_and_save_data(source_path, target_path, text_to_ids)
```
# Check Point
This is your first checkpoint. If you ever decide to come back to this notebook or have to restart the notebook, you can start from here. The preprocessed data has been saved to disk.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
import numpy as np
import helper
(source_int_text, target_int_text), (source_vocab_to_int, target_vocab_to_int), _ = helper.load_preprocess()
```
### Check the Version of TensorFlow and Access to GPU
This will check to make sure you have the correct version of TensorFlow and access to a GPU
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
from distutils.version import LooseVersion
import warnings
import tensorflow as tf
# Check TensorFlow Version
assert LooseVersion(tf.__version__) in [LooseVersion('1.0.0'), LooseVersion('1.0.1')], 'This project requires TensorFlow version 1.0 You are using {}'.format(tf.__version__)
print('TensorFlow Version: {}'.format(tf.__version__))
# Check for a GPU
if not tf.test.gpu_device_name():
warnings.warn('No GPU found. Please use a GPU to train your neural network.')
else:
print('Default GPU Device: {}'.format(tf.test.gpu_device_name()))
```
## Build the Neural Network
You'll build the components necessary to build a Sequence-to-Sequence model by implementing the following functions below:
- `model_inputs`
- `process_decoding_input`
- `encoding_layer`
- `decoding_layer_train`
- `decoding_layer_infer`
- `decoding_layer`
- `seq2seq_model`
### Input
Implement the `model_inputs()` function to create TF Placeholders for the Neural Network. It should create the following placeholders:
- Input text placeholder named "input" using the TF Placeholder name parameter with rank 2.
- Targets placeholder with rank 2.
- Learning rate placeholder with rank 0.
- Keep probability placeholder named "keep_prob" using the TF Placeholder name parameter with rank 0.
Return the placeholders in the following the tuple (Input, Targets, Learing Rate, Keep Probability)
```
def model_inputs():
"""
Create TF Placeholders for input, targets, and learning rate.
:return: Tuple (input, targets, learning rate, keep probability)
"""
# TODO: Implement Function
return None, None, None, None
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_model_inputs(model_inputs)
```
### Process Decoding Input
Implement `process_decoding_input` using TensorFlow to remove the last word id from each batch in `target_data` and concat the GO ID to the beginning of each batch.
```
def process_decoding_input(target_data, target_vocab_to_int, batch_size):
"""
Preprocess target data for decoding
:param target_data: Target Placeholder
:param target_vocab_to_int: Dictionary to go from the target words to an id
:param batch_size: Batch Size
:return: Preprocessed target data
"""
# TODO: Implement Function
return None
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_process_decoding_input(process_decoding_input)
```
### Encoding
Implement `encoding_layer()` to create a Encoder RNN layer using [`tf.nn.dynamic_rnn()`](https://www.tensorflow.org/api_docs/python/tf/nn/dynamic_rnn).
```
def encoding_layer(rnn_inputs, rnn_size, num_layers, keep_prob):
"""
Create encoding layer
:param rnn_inputs: Inputs for the RNN
:param rnn_size: RNN Size
:param num_layers: Number of layers
:param keep_prob: Dropout keep probability
:return: RNN state
"""
# TODO: Implement Function
return None
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_encoding_layer(encoding_layer)
```
### Decoding - Training
Create training logits using [`tf.contrib.seq2seq.simple_decoder_fn_train()`](https://www.tensorflow.org/versions/r1.0/api_docs/python/tf/contrib/seq2seq/simple_decoder_fn_train) and [`tf.contrib.seq2seq.dynamic_rnn_decoder()`](https://www.tensorflow.org/versions/r1.0/api_docs/python/tf/contrib/seq2seq/dynamic_rnn_decoder). Apply the `output_fn` to the [`tf.contrib.seq2seq.dynamic_rnn_decoder()`](https://www.tensorflow.org/versions/r1.0/api_docs/python/tf/contrib/seq2seq/dynamic_rnn_decoder) outputs.
```
def decoding_layer_train(encoder_state, dec_cell, dec_embed_input, sequence_length, decoding_scope,
output_fn, keep_prob):
"""
Create a decoding layer for training
:param encoder_state: Encoder State
:param dec_cell: Decoder RNN Cell
:param dec_embed_input: Decoder embedded input
:param sequence_length: Sequence Length
:param decoding_scope: TenorFlow Variable Scope for decoding
:param output_fn: Function to apply the output layer
:param keep_prob: Dropout keep probability
:return: Train Logits
"""
# TODO: Implement Function
return None
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_decoding_layer_train(decoding_layer_train)
```
### Decoding - Inference
Create inference logits using [`tf.contrib.seq2seq.simple_decoder_fn_inference()`](https://www.tensorflow.org/versions/r1.0/api_docs/python/tf/contrib/seq2seq/simple_decoder_fn_inference) and [`tf.contrib.seq2seq.dynamic_rnn_decoder()`](https://www.tensorflow.org/versions/r1.0/api_docs/python/tf/contrib/seq2seq/dynamic_rnn_decoder).
```
def decoding_layer_infer(encoder_state, dec_cell, dec_embeddings, start_of_sequence_id, end_of_sequence_id,
maximum_length, vocab_size, decoding_scope, output_fn, keep_prob):
"""
Create a decoding layer for inference
:param encoder_state: Encoder state
:param dec_cell: Decoder RNN Cell
:param dec_embeddings: Decoder embeddings
:param start_of_sequence_id: GO ID
:param end_of_sequence_id: EOS Id
:param maximum_length: The maximum allowed time steps to decode
:param vocab_size: Size of vocabulary
:param decoding_scope: TensorFlow Variable Scope for decoding
:param output_fn: Function to apply the output layer
:param keep_prob: Dropout keep probability
:return: Inference Logits
"""
# TODO: Implement Function
return None
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_decoding_layer_infer(decoding_layer_infer)
```
### Build the Decoding Layer
Implement `decoding_layer()` to create a Decoder RNN layer.
- Create RNN cell for decoding using `rnn_size` and `num_layers`.
- Create the output fuction using [`lambda`](https://docs.python.org/3/tutorial/controlflow.html#lambda-expressions) to transform it's input, logits, to class logits.
- Use the your `decoding_layer_train(encoder_state, dec_cell, dec_embed_input, sequence_length, decoding_scope, output_fn, keep_prob)` function to get the training logits.
- Use your `decoding_layer_infer(encoder_state, dec_cell, dec_embeddings, start_of_sequence_id, end_of_sequence_id, maximum_length, vocab_size, decoding_scope, output_fn, keep_prob)` function to get the inference logits.
Note: You'll need to use [tf.variable_scope](https://www.tensorflow.org/api_docs/python/tf/variable_scope) to share variables between training and inference.
```
def decoding_layer(dec_embed_input, dec_embeddings, encoder_state, vocab_size, sequence_length, rnn_size,
num_layers, target_vocab_to_int, keep_prob):
"""
Create decoding layer
:param dec_embed_input: Decoder embedded input
:param dec_embeddings: Decoder embeddings
:param encoder_state: The encoded state
:param vocab_size: Size of vocabulary
:param sequence_length: Sequence Length
:param rnn_size: RNN Size
:param num_layers: Number of layers
:param target_vocab_to_int: Dictionary to go from the target words to an id
:param keep_prob: Dropout keep probability
:return: Tuple of (Training Logits, Inference Logits)
"""
# TODO: Implement Function
return None, None
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_decoding_layer(decoding_layer)
```
### Build the Neural Network
Apply the functions you implemented above to:
- Apply embedding to the input data for the encoder.
- Encode the input using your `encoding_layer(rnn_inputs, rnn_size, num_layers, keep_prob)`.
- Process target data using your `process_decoding_input(target_data, target_vocab_to_int, batch_size)` function.
- Apply embedding to the target data for the decoder.
- Decode the encoded input using your `decoding_layer(dec_embed_input, dec_embeddings, encoder_state, vocab_size, sequence_length, rnn_size, num_layers, target_vocab_to_int, keep_prob)`.
```
def seq2seq_model(input_data, target_data, keep_prob, batch_size, sequence_length, source_vocab_size, target_vocab_size,
enc_embedding_size, dec_embedding_size, rnn_size, num_layers, target_vocab_to_int):
"""
Build the Sequence-to-Sequence part of the neural network
:param input_data: Input placeholder
:param target_data: Target placeholder
:param keep_prob: Dropout keep probability placeholder
:param batch_size: Batch Size
:param sequence_length: Sequence Length
:param source_vocab_size: Source vocabulary size
:param target_vocab_size: Target vocabulary size
:param enc_embedding_size: Decoder embedding size
:param dec_embedding_size: Encoder embedding size
:param rnn_size: RNN Size
:param num_layers: Number of layers
:param target_vocab_to_int: Dictionary to go from the target words to an id
:return: Tuple of (Training Logits, Inference Logits)
"""
# TODO: Implement Function
return None
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_seq2seq_model(seq2seq_model)
```
## Neural Network Training
### Hyperparameters
Tune the following parameters:
- Set `epochs` to the number of epochs.
- Set `batch_size` to the batch size.
- Set `rnn_size` to the size of the RNNs.
- Set `num_layers` to the number of layers.
- Set `encoding_embedding_size` to the size of the embedding for the encoder.
- Set `decoding_embedding_size` to the size of the embedding for the decoder.
- Set `learning_rate` to the learning rate.
- Set `keep_probability` to the Dropout keep probability
```
# Number of Epochs
epochs = None
# Batch Size
batch_size = None
# RNN Size
rnn_size = None
# Number of Layers
num_layers = None
# Embedding Size
encoding_embedding_size = None
decoding_embedding_size = None
# Learning Rate
learning_rate = None
# Dropout Keep Probability
keep_probability = None
```
### Build the Graph
Build the graph using the neural network you implemented.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
save_path = 'checkpoints/dev'
(source_int_text, target_int_text), (source_vocab_to_int, target_vocab_to_int), _ = helper.load_preprocess()
max_source_sentence_length = max([len(sentence) for sentence in source_int_text])
train_graph = tf.Graph()
with train_graph.as_default():
input_data, targets, lr, keep_prob = model_inputs()
sequence_length = tf.placeholder_with_default(max_source_sentence_length, None, name='sequence_length')
input_shape = tf.shape(input_data)
train_logits, inference_logits = seq2seq_model(
tf.reverse(input_data, [-1]), targets, keep_prob, batch_size, sequence_length, len(source_vocab_to_int), len(target_vocab_to_int),
encoding_embedding_size, decoding_embedding_size, rnn_size, num_layers, target_vocab_to_int)
tf.identity(inference_logits, 'logits')
with tf.name_scope("optimization"):
# Loss function
cost = tf.contrib.seq2seq.sequence_loss(
train_logits,
targets,
tf.ones([input_shape[0], sequence_length]))
# Optimizer
optimizer = tf.train.AdamOptimizer(lr)
# Gradient Clipping
gradients = optimizer.compute_gradients(cost)
capped_gradients = [(tf.clip_by_value(grad, -1., 1.), var) for grad, var in gradients if grad is not None]
train_op = optimizer.apply_gradients(capped_gradients)
```
### Train
Train the neural network on the preprocessed data. If you have a hard time getting a good loss, check the forums to see if anyone is having the same problem.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
import time
def get_accuracy(target, logits):
"""
Calculate accuracy
"""
max_seq = max(target.shape[1], logits.shape[1])
if max_seq - target.shape[1]:
target = np.pad(
target,
[(0,0),(0,max_seq - target.shape[1])],
'constant')
if max_seq - logits.shape[1]:
logits = np.pad(
logits,
[(0,0),(0,max_seq - logits.shape[1]), (0,0)],
'constant')
return np.mean(np.equal(target, np.argmax(logits, 2)))
train_source = source_int_text[batch_size:]
train_target = target_int_text[batch_size:]
valid_source = helper.pad_sentence_batch(source_int_text[:batch_size])
valid_target = helper.pad_sentence_batch(target_int_text[:batch_size])
with tf.Session(graph=train_graph) as sess:
sess.run(tf.global_variables_initializer())
for epoch_i in range(epochs):
for batch_i, (source_batch, target_batch) in enumerate(
helper.batch_data(train_source, train_target, batch_size)):
start_time = time.time()
_, loss = sess.run(
[train_op, cost],
{input_data: source_batch,
targets: target_batch,
lr: learning_rate,
sequence_length: target_batch.shape[1],
keep_prob: keep_probability})
batch_train_logits = sess.run(
inference_logits,
{input_data: source_batch, keep_prob: 1.0})
batch_valid_logits = sess.run(
inference_logits,
{input_data: valid_source, keep_prob: 1.0})
train_acc = get_accuracy(target_batch, batch_train_logits)
valid_acc = get_accuracy(np.array(valid_target), batch_valid_logits)
end_time = time.time()
print('Epoch {:>3} Batch {:>4}/{} - Train Accuracy: {:>6.3f}, Validation Accuracy: {:>6.3f}, Loss: {:>6.3f}'
.format(epoch_i, batch_i, len(source_int_text) // batch_size, train_acc, valid_acc, loss))
# Save Model
saver = tf.train.Saver()
saver.save(sess, save_path)
print('Model Trained and Saved')
```
### Save Parameters
Save the `batch_size` and `save_path` parameters for inference.
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
# Save parameters for checkpoint
helper.save_params(save_path)
```
# Checkpoint
```
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
import tensorflow as tf
import numpy as np
import helper
import problem_unittests as tests
_, (source_vocab_to_int, target_vocab_to_int), (source_int_to_vocab, target_int_to_vocab) = helper.load_preprocess()
load_path = helper.load_params()
```
## Sentence to Sequence
To feed a sentence into the model for translation, you first need to preprocess it. Implement the function `sentence_to_seq()` to preprocess new sentences.
- Convert the sentence to lowercase
- Convert words into ids using `vocab_to_int`
- Convert words not in the vocabulary, to the `<UNK>` word id.
```
def sentence_to_seq(sentence, vocab_to_int):
"""
Convert a sentence to a sequence of ids
:param sentence: String
:param vocab_to_int: Dictionary to go from the words to an id
:return: List of word ids
"""
# TODO: Implement Function
return None
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_sentence_to_seq(sentence_to_seq)
```
## Translate
This will translate `translate_sentence` from English to French.
```
translate_sentence = 'he saw a old yellow truck .'
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
translate_sentence = sentence_to_seq(translate_sentence, source_vocab_to_int)
loaded_graph = tf.Graph()
with tf.Session(graph=loaded_graph) as sess:
# Load saved model
loader = tf.train.import_meta_graph(load_path + '.meta')
loader.restore(sess, load_path)
input_data = loaded_graph.get_tensor_by_name('input:0')
logits = loaded_graph.get_tensor_by_name('logits:0')
keep_prob = loaded_graph.get_tensor_by_name('keep_prob:0')
translate_logits = sess.run(logits, {input_data: [translate_sentence], keep_prob: 1.0})[0]
print('Input')
print(' Word Ids: {}'.format([i for i in translate_sentence]))
print(' English Words: {}'.format([source_int_to_vocab[i] for i in translate_sentence]))
print('\nPrediction')
print(' Word Ids: {}'.format([i for i in np.argmax(translate_logits, 1)]))
print(' French Words: {}'.format([target_int_to_vocab[i] for i in np.argmax(translate_logits, 1)]))
```
## Imperfect Translation
You might notice that some sentences translate better than others. Since the dataset you're using only has a vocabulary of 227 English words of the thousands that you use, you're only going to see good results using these words. Additionally, the translations in this data set were made by Google translate, so the translations themselves aren't particularly good. (We apologize to the French speakers out there!) Thankfully, for this project, you don't need a perfect translation. However, if you want to create a better translation model, you'll need better data.
You can train on the [WMT10 French-English corpus](http://www.statmt.org/wmt10/training-giga-fren.tar). This dataset has more vocabulary and richer in topics discussed. However, this will take you days to train, so make sure you've a GPU and the neural network is performing well on dataset we provided. Just make sure you play with the WMT10 corpus after you've submitted this project.
## Submitting This Project
When submitting this project, make sure to run all the cells before saving the notebook. Save the notebook file as "dlnd_language_translation.ipynb" and save it as a HTML file under "File" -> "Download as". Include the "helper.py" and "problem_unittests.py" files in your submission.
| github_jupyter |
<h4>
<ul>
<li><strong><span style="font-family: 'Trebuchet MS', Helvetica, sans-serif;">OBJECTIVE:</span></strong></li>
</ul>
<p><span style="font-family: 'Trebuchet MS', Helvetica, sans-serif;">Use the text data to build simple feed-forward Neural Nets and benchmark against the base ML models.</span></p>
</h4>
```
# imports
import os
import math
import random
import warnings
from time import time
from pathlib import Path
import pandas as pd, numpy as np
from pprint import pprint
import matplotlib.pyplot as plt
import seaborn as sns
from tqdm import tqdm
from collections import defaultdict, Counter
from sklearn.preprocessing import LabelEncoder
from wordcloud import WordCloud, STOPWORDS
import tensorflow
tqdm.pandas()
warnings.filterwarnings('ignore')
warnings.simplefilter(action='ignore', category=FutureWarning)
%matplotlib inline
# reproducibility
seed = 7
random.seed(seed)
tensorflow.random.set_seed(seed)
```
<h3><ul>
<li><strong><span style="font-family: 'Trebuchet MS', Helvetica, sans-serif;">Import & Analyse the data.</span></strong></li>
</ul></h3>
```
dataset = pd.read_excel('./data/cleaned_data.xlsx')
dataset.sample(10)
dataset.isna().sum()
dataset[dataset.isna().any(axis=1)].to_csv('./data/missing_keywords.csv')
dataset[dataset.isna().any(axis=1)] # check rows with missing values
le = LabelEncoder()
dataset['group_code'] = le.fit_transform(dataset.group)
dataset.info()
le.classes_
X = np.array(dataset.cleaned_short_description)
y = np.array(dataset.group_code)
X.shape, y.shape
from keras.utils import np_utils
y_dummy_coded = np_utils.to_categorical(y)
y[0], y_dummy_coded[0]
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y_dummy_coded, test_size=.2, random_state=seed) # splits are stratified by y
X_train.shape, X_test.shape, y_train.shape, y_test.shape
X_train[0], y_train[0] # check sample
# TODO: Check the distributions of groups in training and testing sets, i.e, if they vary too much
# stratify by y if required during splits
# or data augmentation to upsample minority classes to balance the group distributions
```
<h3><ul>
<li><strong><span style="font-family: 'Trebuchet MS', Helvetica, sans-serif;">Tokenize and pad sequences</span></strong></li>
</ul></h3>
```
# define params
NUM_WORDS = 20000
EMBEDDING_DIM = 300
MAX_LEN = 100 # dataset['word_length'].max()
MAX_LEN
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
tokenizer = Tokenizer(num_words=NUM_WORDS)
tokenizer.fit_on_texts(X_train)
X_train_tokens = tokenizer.texts_to_sequences(X_train)
X_test_tokens = tokenizer.texts_to_sequences(X_test)
X_train_tokens[0], X_test_tokens[0]
y_train[0], y_test[0]
# pad sequences to cut longer texts to a uniform length and pad the sentences that are shorter than that with <PAD> token
# using just 20 words from each headline will severely limit the information that is
# available to the model and affect performance although the training will be faster
X_train_padded = pad_sequences(X_train_tokens,
padding='post',
truncating='post',
maxlen=MAX_LEN)
X_test_padded = pad_sequences(X_test_tokens,
padding='post',
truncating='post',
maxlen=MAX_LEN)
print(f'X train: {X_train_padded.shape}\nX test: {X_test_padded.shape}')
pprint(X_train_padded[0], compact=True)
WORD_TO_INDEX = tokenizer.word_index
# pprint(WORD_TO_INDEX, compact=True)
pprint(list(WORD_TO_INDEX.keys())[:100], compact=True)
VOCAB_SIZE = len(WORD_TO_INDEX) + 1
VOCAB_SIZE
# https://www.tensorflow.org/api_docs/python/tf/keras/preprocessing/sequence/pad_sequences
def retrieve_description_feat(x, mapping=WORD_TO_INDEX) -> str:
# increment 3
mapping = {k:(v + 3) for k, v in mapping.items()}
mapping['<PAD>'] = 0
mapping['<START>'] = 1
mapping['<UNK>'] = 2
inv_mapping = {v: k for k, v in mapping.items()}
return str(" ".join(inv_mapping.get(i, '<NA>') for i in x))
retrieve_description_feat(X_test_padded[7])
```
<h3><ul>
<li><strong><span style="font-family: 'Trebuchet MS', Helvetica, sans-serif;">GloVe Embeddings</span></strong></li>
</ul></h3>
```
EMBEDDING_DIM
def get_embedding_matrix(embedding_dim=EMBEDDING_DIM):
embeddings = defaultdict()
if embedding_dim == 200:
file_path = f'./data/glove.6B.{embedding_dim}d.txt'
elif embedding_dim == 300:
file_path = f'./data/glove.840B.{embedding_dim}d.txt'
for l in open(file_path, encoding='utf-8'):
word = l.split(" ")[0]
embeddings[word] = np.asarray(l.split(" ")[1:], dtype='float32')
embeddings = dict(embeddings)
# create a weight matrix for words in training docs
embedding_matrix = np.zeros((NUM_WORDS, embedding_dim))
for word, idx in WORD_TO_INDEX.items():
embedding_vector = embeddings.get(word)
if embedding_vector is not None:
embedding_matrix[idx] = embedding_vector
return embedding_matrix
# use pre-trained glove embedding matrix to initialize weights in our model
# embedding_matrix = get_embedding_matrix()
# embedding_matrix.shape
```
<h4><ul>
<li><strong><span style="font-family: 'Trebuchet MS', Helvetica, sans-serif;">Simple Feed-Forward Neural Net</span></strong></li>
</ul></h4>
```
# !pip install livelossplot
from tensorflow.python.keras.models import Sequential
from sklearn.metrics import accuracy_score, confusion_matrix
from tensorflow.keras.regularizers import l2
from tensorflow.keras.constraints import max_norm, unit_norm
from tensorflow.python.keras.callbacks import LambdaCallback, EarlyStopping, ReduceLROnPlateau
from tensorflow.keras.layers import Flatten, Dense, Activation, BatchNormalization, Dropout, Embedding, LSTM, MaxPooling1D, Conv1D
NUM_CLASSES = len(le.classes_)
VOCAB_SIZE, MAX_LEN, EMBEDDING_DIM, NUM_CLASSES
# define model
model1 = Sequential([
Embedding(input_dim=VOCAB_SIZE, output_dim=EMBEDDING_DIM, input_length=MAX_LEN),
Flatten(),
Dense(1024, activation = 'relu'),
Dense(1024, activation = 'relu'),
Dense(128, activation = 'relu'),
Dense(NUM_CLASSES, activation = 'softmax')
])
model1.compile(
loss = 'categorical_crossentropy',
optimizer = 'adam',
metrics = ['accuracy']
)
# Define Callbacks and a few helper functions
# simplify the training log
simple_log = LambdaCallback(
on_epoch_end = lambda e, l: print(f" ~| Epoch: {e+1} | Validation Loss: {l['val_loss']:.5f}", end =" >|> \n" ))
# early stopping
early_stop = EarlyStopping(monitor='val_loss',
min_delta=0,
patience=7,
verbose=0,
restore_best_weights=True)
# learning rate reduction
lr_reduce_on_plateau = ReduceLROnPlateau(monitor='val_loss',
patience=4,
verbose=1,
factor=0.4,
min_lr=0.00001)
def plot_learning_curve(hist):
sns.set()
plt.figure(figsize=(5,5))
train = hist.history['loss']
val = hist.history['val_loss']
epochs_run = range(1,len(train) + 1)
sns.lineplot(epochs_run, train, marker = 'o', color = 'coral', label = 'Training Loss')
sns.lineplot(epochs_run, val, marker = '>', color = 'green', label = 'Validation Loss')
plt.title("Loss vs. Epochs", fontsize = 20)
plt.legend()
plt.show()
# !pip install nvidia-smi
!nvidia-smi
import gc
gc.collect()
X_train[0]
X_train.shape, y_train.shape, X_test.shape, y_test.shape
EPOCHS = 200
try:
print("Training on GPU:")
with tensorflow.device("gpu:0"): # train on gpu
h1 = model1.fit(
X_train_padded, y_train,
validation_split = 0.2, # do not use the test data for validation to prevent data leakage, we only use to test at the model in the end
epochs = EPOCHS,
callbacks = [simple_log, early_stop, lr_reduce_on_plateau],
verbose = False)
except Exception as e:
print(e)
print("\nTraining on CPU:")
h1 = model1.fit(
X_train_padded, y_train,
validation_split = 0.2, # do not use the test data for validation to prevent data leakage, we only use to test at the model in the end
epochs = EPOCHS,
callbacks = [simple_log, early_stop, lr_reduce_on_plateau],
verbose = False)
print("\nTraining Done.")
plot_learning_curve(h1)
loss, acc = model1.evaluate(X_test_padded, y_test)
print("Testing Loss: ", loss*100)
print("Testing Accuracy: ", acc*100)
```
<h4><ul>
<span style="font-family: 'Trebuchet MS', Helvetica, sans-serif;">
<li>This model is clearly overfitting, we will add regularization to the next iteration
</span></ul></h4>
```
# define model
model2 = Sequential([
Embedding(input_dim=VOCAB_SIZE, output_dim=EMBEDDING_DIM, input_length=MAX_LEN),
Flatten(),
Dense(256, activation = 'relu'),
BatchNormalization(),
Dense(256, activation = 'relu'),
BatchNormalization(),
Dense(NUM_CLASSES, activation = 'softmax')
])
model2.compile(
loss = 'categorical_crossentropy',
optimizer = 'adam',
metrics = ['accuracy']
)
EPOCHS = 200
try:
print("Training on GPU:")
with tensorflow.device("gpu:0"): # train on gpu
h2 = model2.fit(
X_train_padded, y_train,
validation_split = 0.2, # do not use the test data for validation to prevent data leakage, we only use to test at the model in the end
epochs = EPOCHS,
callbacks = [simple_log, early_stop, lr_reduce_on_plateau],
verbose = False)
except Exception as e:
print(e)
print("Training on CPU:")
h2 = model2.fit(
X_train_padded, y_train,
validation_split = 0.2, # do not use the test data for validation to prevent data leakage, we only use to test at the model in the end
epochs = EPOCHS,
callbacks = [simple_log, early_stop, lr_reduce_on_plateau],
verbose = False)
print("\nTraining Done.")
plot_learning_curve(h2)
loss, acc = model2.evaluate(X_test_padded, y_test)
print("Testing Loss: ", loss*100)
print("Testing Accuracy: ", acc*100)
# define model
model3 = Sequential([
Embedding(input_dim=VOCAB_SIZE, output_dim=EMBEDDING_DIM, input_length=MAX_LEN),
Flatten(),
Dense(20, activation = 'relu'),
Dropout(0.4),
Dense(NUM_CLASSES, activation = 'softmax')
])
model3.compile(
loss = 'categorical_crossentropy',
optimizer = 'adam',
metrics = ['accuracy']
)
EPOCHS = 200
try:
print("Training on GPU:")
with tensorflow.device("gpu:0"): # train on gpu
h3 = model3.fit(
X_train_padded, y_train,
validation_split = 0.2, # do not use the test data for validation to prevent data leakage, we only use to test at the model in the end
epochs = EPOCHS,
callbacks = [simple_log, early_stop, lr_reduce_on_plateau],
verbose = False)
except Exception as e:
print(e)
print("Training on CPU:")
h3 = model3.fit(
X_train_padded, y_train,
validation_split = 0.2, # do not use the test data for validation to prevent data leakage, we only use to test at the model in the end
epochs = EPOCHS,
callbacks = [simple_log, early_stop, lr_reduce_on_plateau],
verbose = False)
print("\nTraining Done.")
plot_learning_curve(h3)
loss, acc = model3.evaluate(X_test_padded, y_test)
print("Testing Accuracy: ", acc*100)
```
<h4><ul>
<li><strong><span style="font-family: 'Trebuchet MS', Helvetica, sans-serif;">LSTM</span></strong></li>
</ul></h4>
```
# define model
model4 = Sequential([
Embedding(input_dim=VOCAB_SIZE, output_dim=EMBEDDING_DIM, input_length=MAX_LEN),
LSTM(128),
Dropout(0.4),
Dense(NUM_CLASSES, activation = 'softmax')
])
model4.compile(
loss = 'categorical_crossentropy',
optimizer = 'adam',
metrics = ['accuracy']
)
EPOCHS = 50
try:
print("Training on GPU:")
with tensorflow.device("gpu:0"): # train on gpu
h4 = model4.fit(
X_train_padded, y_train,
validation_split = 0.2, # do not use the test data for validation to prevent data leakage, we only use to test at the model in the end
epochs = EPOCHS,
callbacks = [simple_log, early_stop, lr_reduce_on_plateau],
verbose = False)
except Exception as e:
print(e)
print("Training on CPU:")
h4 = model4.fit(
X_train_padded, y_train,
validation_split = 0.2, # do not use the test data for validation to prevent data leakage, we only use to test at the model in the end
epochs = EPOCHS,
callbacks = [simple_log, early_stop, lr_reduce_on_plateau],
verbose = False)
print("\nTraining Done.")
plot_learning_curve(h4)
loss, acc = model4.evaluate(X_test_padded, y_test)
print("Testing Accuracy: ", acc*100)
model5 = Sequential([
Embedding(input_dim=VOCAB_SIZE, output_dim=256, input_length=MAX_LEN),
Dropout(0.25),
Conv1D(256, 5, padding = 'same', activation = 'relu', strides = 1),
Conv1D(256, 5, padding = 'same', activation = 'relu', strides = 1),
MaxPooling1D(pool_size = 2),
Conv1D(64, 5, padding = 'same', activation = 'relu', strides = 1),
MaxPooling1D(pool_size = 2),
LSTM(75),
Dense(NUM_CLASSES, activation = 'softmax')
])
model5.compile(
loss = 'categorical_crossentropy',
optimizer = 'adam',
metrics = ['accuracy']
)
EPOCHS = 20
if tensorflow.test.is_gpu_available():
print("Training on GPU:")
with tensorflow.device("gpu:0"): # train on gpu
h5 = model5.fit(
X_train_padded, y_train,
validation_split = 0.2, # do not use the test data for validation to prevent data leakage, we only use to test at the model in the end
epochs = EPOCHS,
callbacks = [simple_log, early_stop, lr_reduce_on_plateau],
verbose = False)
else:
print("Training on CPU:")
h5 = model5.fit(
X_train_padded, y_train,
validation_split = 0.2, # do not use the test data for validation to prevent data leakage, we only use to test at the model in the end
epochs = EPOCHS,
callbacks = [simple_log, early_stop, lr_reduce_on_plateau],
verbose = False)
print("\nTraining Done.")
plot_learning_curve(h5)
loss, acc = model5.evaluate(X_test_padded, y_test)
print("Testing Accuracy: ", acc*100)
```
<h4><ul>
<li><strong><span style="font-family: 'Trebuchet MS', Helvetica, sans-serif;">Use TfIdf vectors instead of Embedding Layer + Feature Selection</span></strong></li>
</ul></h4>
```
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.model_selection import train_test_split
# using 75-25 split instead of 50-50 split as we need more data to train neural nets
X_train_vec, X_test_vec, y_train_vec, y_test_vec = train_test_split(X, y, test_size=0.2, random_state=seed)
print(f"Train dataset shape: {X_train_vec.shape}, \nTest dataset shape: {X_test_vec.shape}")
NGRAM_RANGE = (1, 2)
TOP_K = 20000
TOKEN_MODE = 'word'
MIN_DOC_FREQ = 2
kwargs = {
'ngram_range' : NGRAM_RANGE,
'dtype' : 'int32',
'strip_accents' : 'unicode',
'decode_error' : 'replace',
'analyzer' : TOKEN_MODE,
'min_df' : MIN_DOC_FREQ
}
vectorizer = TfidfVectorizer(**kwargs)
X_train_vec = vectorizer.fit_transform(X_train_vec)
X_test_vec = vectorizer.transform(X_test_vec)
print(f"Train dataset shape: {X_train_vec.shape}, \nTest dataset shape: {X_test_vec.shape}")
from sklearn.feature_selection import SelectKBest, f_classif
# Select best k features, with feature importance measured by f_classif
# Set k as 20000 or (if number of ngrams is less) number of ngrams
selector = SelectKBest(f_classif, k=min(TOP_K, X_train_vec.shape[1]))
selector.fit(X_train_vec, y_train_vec)
X_train_vec = selector.transform(X_train_vec).astype('float32')
X_test_vec = selector.transform(X_test_vec).astype('float32')
X_train_vec = X_train_vec.toarray()
X_test_vec = X_test_vec.toarray()
print(f"Train dataset shape: {X_train.shape}, \nTest dataset shape: {X_test.shape}")
model6 = Sequential([
Dense(64, activation='relu', input_shape=X_train_vec.shape[1:]),
Dropout(0.2),
Dense(16, activation='relu'),
Dropout(0.2),
Dense(NUM_CLASSES, activation='softmax')
])
model6.compile(
loss = 'categorical_crossentropy',
optimizer = 'adam',
metrics = ['accuracy']
)
EPOCHS = 20
try:
print("Training on GPU:")
with tensorflow.device("gpu:0"): # train on gpu
h6 = model6.fit(
X_train_vec, y_train,
validation_split = 0.2, # do not use the test data for validation to prevent data leakage, we only use to test at the model in the end
epochs = EPOCHS,
callbacks = [simple_log, early_stop],
verbose = False)
except Exception:
print("Training on CPU:")
h6 = model6.fit(
X_train_vec, y_train,
validation_split = 0.2, # do not use the test data for validation to prevent data leakage, we only use to test at the model in the end
epochs = EPOCHS,
callbacks = [simple_log, early_stop],
verbose = False)
print("\nTraining Done.")
plot_learning_curve(h6)
loss, acc = model6.evaluate(X_test_vec, y_test)
print("Testing Accuracy: ", acc*100)
```
<h3><ul>
<li><strong><span style="font-family: 'Trebuchet MS', Helvetica, sans-serif;">Metrics: </span></strong></li>
</ul></h3>
<table style="width:100%">
<tr>
<th>Model</th>
<th>Test Accuracy</th>
</tr>
<tr>
<td>Simple Feed-Forward Net using Embedding Layer</td>
<td></td>
</tr>
<tr>
<td>Feed-Forward + Batch Norm</td>
<td></td>
</tr>
<tr>
<td>Feed-Forward + Dropout</td>
<td></td>
</tr>
<tr>
<td>LSTM</td>
<td></td>
</tr>
<tr>
<td>Convolution Blocks (Dimensionality Reduction) + LSTM</td>
<td></td>
</tr>
<tr>
<td>TfIdf Vectors + Better train-test split + Feature Selection + Feed-forward Neural Net</td>
<td>63</td>
</tr>
</table>
<h4><ul>
<li><strong><span style="font-family: 'Trebuchet MS', Helvetica, sans-serif;"></span></strong></li>
</ul></h4>
| github_jupyter |
```
import pandas as pd
# データの読み込み
reserve_tb = pd.read_csv('data/reserve.csv', encoding='UTF-8')
```
# 第3章 集約
## 3-1 データ数,種類数の算出
最も基本的な集約処理として,データ数のカウントがあります.これは,対象となるデータのレコード数(行数)をカウントする処理です.この他にもよく利用するカウント処理として,データのユニークカウントがあります.ユニークカウントとは,対象となるデータから同じ値のレコードを排除した後にレコード数をカウントする処理です.つまり,データの値の種類をカウントします.
### Q:カウントとユニークカウント
対象のデータセットは,ホテルの予約レコードです.\
予約テーブルから,ホテルごとに予約件数と予約したことがある顧客数を算出しましょう.\
\
ヒント:agg関数で集約処理をまとめて指定するやりやすいです
```
# agg関数を利用して、集約処理をまとめて指定
# reserve_idを対象にcount関数を適用
# customer_idを対象にnunique関数を適用
result_1 = reserve_tb \
.groupby('hotel_id') \
.agg({'reserve_id': 'count', 'customer_id': 'nunique'})
# reset_index関数によって、列番号を振り直す(inplace=Trueなので、直接resultを更新)
result_1.reset_index(inplace=True)
result_1.columns = ['hotel_id', 'rsv_cnt', 'cus_cnt']
```
## 3-2 合計値の算出
### Q:合計値
対象のデータセットはホテルの予約レコードです.\
予約テーブルから,ホテルごとの宿泊人数別の合計予約金額を算出しましょう.
```
# 集約単位をhotel_idとpeople_numの組み合わせを指定
# 集約したデータからtotal_priceを取り出し、sum関数に適用することで売上合計金額を算出
result_2 = reserve_tb \
.groupby(['hotel_id', 'people_num'])['total_price'] \
.sum().reset_index()
# 売上合計金額の列名がtotal_priceになっているので、price_sumに変更
result_2.rename(columns={'total_price': 'price_sum'}, inplace=True)
```
## 3-3 極値,代表値の算出
### Q:代表値
対象のデータセットはホテルの予約レコードです.
予約テーブルから,ホテルごとの予約金額の最大値,最小値,平均値,中央値,20パーセンタイル値を算出しましょう.
```
# total_priceを対象にmax/min/mean/median関数を適用
# Pythonのラムダ式をagg関数の集約処理に指定
# ラムダ式にはnumpy.percentileを指定しパーセントタイル値を算出(パーセントは20指定)
import numpy as np
result_3 = reserve_tb \
.groupby('hotel_id') \
.agg({'total_price': ['max', 'min', 'mean', 'median',
lambda x: np.percentile(x, q=20)]}) \
.reset_index()
result_3.columns = ['hotel_id', 'price_max', 'price_min', 'price_mean',
'price_median', 'price_20per']
```
## 3-4 ばらつき具合の算出
### Q:分散値と標準偏差
対象のデータセットはホテルの予約レコードです.\
予約テーブルから,各ホテルの予約金額の分散値と標準偏差値を算出しましょう.\
ただし,予約が1件しかない場合は,分散値と標準偏差値を0としましょう.
```
# total_priceに対して、var関数とstd関数を適用し、分散値と標準偏差値を算出
result_4 = reserve_tb \
.groupby('hotel_id') \
.agg({'total_price': ['var', 'std']}).reset_index()
result_4.columns = ['hotel_id', 'price_var', 'price_std']
# データ数が1件だったときは、分散値と標準偏差値がnaになっているので、0に置き換え
result_4.fillna(0, inplace=True)
```
## 3-5 最頻値の算出
最頻値は最も多く出現しているあたいのことです.\
数値でも,カテゴリ値に変換することによって最頻値を利用できます.\
例えば,数値を四捨五入で整数かしたり,100ごとのレンジで値をカテゴリ化して,最頻値を利用します.
### Q:最頻値
対象のデータセットは,ホテルの予約レコードです.予約テーブルの予約金額を1000単位にカテゴリ化して最頻値を算出しましょう.
```
# round関数で四捨五入した後に、mode関数で最頻値を算出
reserve_tb['total_price'].round(-3).mode()
```
## 3-6 順位の算出
対象データを絞る際に順位を利用したり,複雑な時系列の結合をする際に時間順に順位づけし,結合条件に利用することもできます.\
順位づけする際には,計算こづとに注意する必要があります.順位づけには並び替えを実地する必要があり,データ数が多いと計算コストが跳ね上がってしまうからです.\
順位付けをする計算は,Window関数を利用すると柑橘かつ計算パフォーマンスよくかけます.Window関数は集約関数の1つですが,通常の集約関数とは違う点があります.それは,行を集約せず,集約した値を計算してから各行に付与するという点です.
### Q:時系列に番号を付与
対象のデータセットは,ホテルの予約レコードです.\
予約テーブルを利用して,顧客ごとに予約日時の順位を古い順につけましょう.\
同じ予約日時の場合は,データ行の読み込み順に小さな順位をつけましょう.
```
# rank関数で並び替えるために、データ型を文字列からtimestamp型に変換
# (「第10章 日時型」で解説)
reserve_tb['reserve_datetime'] = pd.to_datetime(
reserve_tb['reserve_datetime'], format='%Y-%m-%d %H:%M:%S'
)
# log_noを新たな列として追加
# 集約単位の指定はgroup_byを利用
# 顧客ごとにまとめたreserve_datetimeを生成し、rank関数によって順位を生成
# ascendingをTrueにすることで昇順に設定(Falseだと降順に設定)
reserve_tb['log_no'] = reserve_tb \
.groupby('customer_id')['reserve_datetime'] \
.rank(ascending=True, method='first')
```
### Q:ランキング
対象のデータセットは,ホテルの予約レコードです.\
予約テーブルを利用して,ホテルごとの予約数に順位付けしましょう.\
同じ予約数の場合は,同予約数の全ホテルに最小の順位をつけましょう
```
# 予約回数を計算(「3-1 データ数、種類数の算出」の例題を参照)
rsv_cnt_tb = reserve_tb.groupby('hotel_id').size().reset_index()
rsv_cnt_tb.columns = ['hotel_id', 'rsv_cnt']
# 予約回数をもとに順位を計算
# ascendingをFalseにすることで降順に指定
# methodをminに指定し、同じ値の場合は取り得る最小順位に指定
rsv_cnt_tb['rsv_cnt_rank'] = rsv_cnt_tb['rsv_cnt'] \
.rank(ascending=False, method='min')
# 必要のないrsv_cntの列を削除
rsv_cnt_tb.drop('rsv_cnt', axis=1, inplace=True)
```
| github_jupyter |
# Maryland schools star ratings map
By [Christine Zhang](mailto:czhang@baltsun.com)
This notebook appends latitude and longitude coordinates for schools in Maryland for mapping purposes.
The map can be found embedded in Baltimore Sun stories [here](https://www.baltimoresun.com/news/maryland/education/k-12/bs-md-star-rating-release-20181203-story.html) and [here](https://www.baltimoresun.com/news/maryland/education/k-12/bs-md-star-ratings-key-takeaways-20181204-story.html).
Geographical information for schools comes from National Center for Education Statistics 2016-17 [Education Demographic and Geographic Estimates (EDGE)](https://nces.ed.gov/programs/edge/Geographic/SchoolLocations).
## How we did it
### Import R data analysis libraries and read in star ratings data
```
suppressMessages(library('tidyverse'))
suppressMessages(library('stringr'))
suppressMessages(library('janitor'))
```
Read in the scores data.
```
scores <- suppressMessages(read_csv('input/accountability_schools_download_file.csv', na = 'na') %>% clean_names())
```
Schools in the star ratings data are uniquely identified by a combination of the `lea_number` and `school_number`.
```
glimpse(scores)
```
### Read in the EDGE data, which provides coordinates for schools nationwide
```
edge <- read.csv('input/EDGE_GEOCODE_PUBLICSCH_1617.csv', stringsAsFactors = F,
colClasses = c('NCESSCH' = 'character')) %>% clean_names()
```
Schools in the EDGE data are uniquely idenfified by the 12-digit `ncessch` number (it's irrelevant for MD schools, but we specify `colClasses = c('NCESSCH' = 'character')` so that `R` will not drop the leading zero.
```
glimpse(edge)
```
### Read in the school directory data
We can't directly match up schools in the star ratings data with schools in the EDGE data because they are have different identifiers. Here we read in the Maryland school directory from the Maryland State Department of Education [website](http://reportcard.msde.maryland.gov/). This file provides the a way to link the two datasets.
```
directory <- suppressMessages(read.csv('input/School_Directory_2018.csv',
colClasses = c('LEA.Number' = 'character',
'School.Number' = 'character',
'NCES.Number' = 'character')) %>% clean_names())
```
Schools are identified by `lea_number` and `school_number`.
```
glimpse(directory)
```
### Merge `scores` with `directory` to get the NCES id for each school
We can merge the `scores` and the `directory` dataframes on the `lea_number` and `school_number` columns. However, we first need to add a leading zero to `lea_number` and `school_number` in the `scores` dataframe. We can do this using `str_pad()`.
```
scores$school_number <- str_pad(scores$school_number, 4, pad = '0')
scores$lea_number <- str_pad(scores$lea_number, 2, pad = '0')
```
We will call the merged dataframe `scores.nces`.
```
scores.nces <- merge(scores, directory %>% select(-lea_name, -school_name),
by = c('lea_number', 'school_number'), all.x = T)
```
### Merge `scores.nces` with `edge` to get the geographical coordinates for each school
We can merge the `scores.nces` and the `edge` dataframes on the `nces_number` (from `scores.nces`) and `ncessch` (from `edge`). This is the 12-digit NCES id for each school. We will call the merged dataframe `scores.geo`.
```
scores.geo <- merge(scores.nces, edge,
by.x = 'nces_number',
by.y = 'ncessch', all.x = T,
suffixes = c('_msde', '_nces'))
```
Note: there are three schools that do no have coordinates provided by EDGE.
```
scores.geo %>% filter(is.na(lat))
```
We can add in the coordinates for these schools manually.
```
added <- suppressMessages(read_csv('input/addresses_add.csv'))
scores.geo.added <- merge(scores.geo, added, by = c('lea_number', 'school_number'), all.x = T)
scores.geo.added <- scores.geo.added %>% mutate(lat = ifelse(is.na(lat.x), lat.y, lat.x),
lon = ifelse(is.na(lon.x), lon.y, lon.x),
address = ifelse(is.na(address.x), as.character(address.y), as.character(address.x)),
city_msde = ifelse(is.na(city_msde.x), as.character(city_msde.y), as.character(city_msde.x)))
scores.geo.added <- scores.geo.added %>% select(lea_number,
lea_name,
school_number,
school_name = school_name.x,
nces_number,
number_academic_year,
star_rating,
total_earned_points_percent,
percentile_rank_elementary,
percentile_rank_middle,
percentile_rank_high,
address,
city = city_msde,
lat,
lon)
head(scores.geo.added)
```
Write to `output/` folder
```
write_csv(scores.geo.added, 'output/scores_clean.csv')
```
| github_jupyter |
```
#libs for reading data
import xarray as xr
import numpy as np
import matplotlib.pyplot as plt
import intake
import dask
#libs for dask gateway
from dask_gateway import Gateway
from dask.distributed import Client
```
### Start a cluster, a group of computers that will work together.
(A cluster is the key to big data analysis on on Cloud.)
- This will set up a [dask kubernetes](https://docs.dask.org/en/latest/setup/kubernetes.html) cluster for your analysis and give you a path that you can paste into the top of the Dask dashboard to visualize parts of your cluster.
- You don't need to paste the link below into the Dask dashboard for this to work, but it will help you visualize progress.
- Try 20 workers to start (during the tutorial) but you can increase to speed things up later
```
gateway = Gateway()
cluster = gateway.new_cluster()
cluster.adapt(minimum=1, maximum=20)
client = Client(cluster)
cluster
```
** ☝️ Don’t forget to click the link above or copy it to the Dask dashboard  on the left to view the scheduler dashboard! **
### Initialize Dataset
Here we load the dataset from the zarr store. Note that this very large dataset (273 GB) initializes nearly instantly, and we can see the full list of variables and coordinates.
### Examine Metadata
For those unfamiliar with this dataset, the variable metadata is very helpful for understanding what the variables actually represent
Printing the dataset will show you the dimensions, coordinates, and data variables with clickable icons at the end that show more metadata and size.
```
%%time
cat_pangeo = intake.open_catalog("https://raw.githubusercontent.com/pangeo-data/pangeo-datastore/master/intake-catalogs/master.yaml")
ds_ccmp = cat_pangeo.atmosphere.nasa_ccmp_wind_vectors.to_dask()
ds_ccmp['wspd'] = np.sqrt(ds_ccmp.uwnd**2 + ds_ccmp.vwnd**2)
ds_ccmp
```
# time series plot
```
%%time
ds_ccmp.sel(latitude=slice(0,50),longitude=slice(180,210)).mean({'latitude','longitude'}).wspd.plot()
```
# year average plot
```
%%time
ds_ccmp.sel(time=slice('2000-01-01','2000-12-31')).mean({'time'}).wspd.plot()
```
# hovmoller type plot
```
%%time
ds_ccmp.sel(latitude=0.125,longitude=slice(120,275)).wspd.plot(vmin=3,vmax=15,cmap='magma')
```
# read data NOT using intake
```
%%time
import gcsfs
zstore = 'gs://pangeo-nasa-ccmp/zarr'
fs = gcsfs.GCSFileSystem(requester_pays=True)
ds = xr.open_zarr(fs.get_mapper(zstore), consolidated=True)
ds['wspd'] = np.sqrt(ds.uwnd**2 + ds.vwnd**2)
ds
```
| github_jupyter |
# 机器学习工程师纳米学位
## 机器学习基础
## 项目 0: 预测泰坦尼克号乘客生还率
1912年,泰坦尼克号在第一次航行中就与冰山相撞沉没,导致了大部分乘客和船员身亡。在这个入门项目中,我们将探索部分泰坦尼克号旅客名单,来确定哪些特征可以最好地预测一个人是否会生还。为了完成这个项目,你将需要实现几个基于条件的预测并回答下面的问题。我们将根据代码的完成度和对问题的解答来对你提交的项目的进行评估。
> **提示**:这样的文字将会指导你如何使用 iPython Notebook 来完成项目。
点击[这里](https://github.com/udacity/machine-learning/blob/master/projects/titanic_survival_exploration/titanic_survival_exploration.ipynb)查看本文件的英文版本。
### 了解数据
当我们开始处理泰坦尼克号乘客数据时,会先导入我们需要的功能模块以及将数据加载到 `pandas` DataFrame。运行下面区域中的代码加载数据,并使用 `.head()` 函数显示前几项乘客数据。
> **提示**:你可以通过单击代码区域,然后使用键盘快捷键 **Shift+Enter** 或 **Shift+ Return** 来运行代码。或者在选择代码后使用**播放**(run cell)按钮执行代码。像这样的 MarkDown 文本可以通过双击编辑,并使用这些相同的快捷键保存。[Markdown](http://daringfireball.net/projects/markdown/syntax) 允许你编写易读的纯文本并且可以转换为 HTML。
```
# 检查你的Python版本
from sys import version_info
if version_info.major != 2 and version_info.minor != 7:
raise Exception('请使用Python 2.7来完成此项目')
import numpy as np
import pandas as pd
# 数据可视化代码
from titanic_visualizations import survival_stats
from IPython.display import display
%matplotlib inline
# 加载数据集
in_file = 'titanic_data.csv'
full_data = pd.read_csv(in_file)
# 显示数据列表中的前几项乘客数据
display(full_data.head())
```
从泰坦尼克号的数据样本中,我们可以看到船上每位旅客的特征
- **Survived**:是否存活(0代表否,1代表是)
- **Pclass**:社会阶级(1代表上层阶级,2代表中层阶级,3代表底层阶级)
- **Name**:船上乘客的名字
- **Sex**:船上乘客的性别
- **Age**:船上乘客的年龄(可能存在 `NaN`)
- **SibSp**:乘客在船上的兄弟姐妹和配偶的数量
- **Parch**:乘客在船上的父母以及小孩的数量
- **Ticket**:乘客船票的编号
- **Fare**:乘客为船票支付的费用
- **Cabin**:乘客所在船舱的编号(可能存在 `NaN`)
- **Embarked**:乘客上船的港口(C 代表从 Cherbourg 登船,Q 代表从 Queenstown 登船,S 代表从 Southampton 登船)
因为我们感兴趣的是每个乘客或船员是否在事故中活了下来。可以将 **Survived** 这一特征从这个数据集移除,并且用一个单独的变量 `outcomes` 来存储。它也做为我们要预测的目标。
运行该代码,从数据集中移除 **Survived** 这个特征,并将它存储在变量 `outcomes` 中。
```
# 从数据集中移除 'Survived' 这个特征,并将它存储在一个新的变量中。
outcomes = full_data['Survived']
data = full_data.drop('Survived', axis = 1)
# 显示已移除 'Survived' 特征的数据集
display(data.head())
```
这个例子展示了如何将泰坦尼克号的 **Survived** 数据从 DataFrame 移除。注意到 `data`(乘客数据)和 `outcomes` (是否存活)现在已经匹配好。这意味着对于任何乘客的 `data.loc[i]` 都有对应的存活的结果 `outcome[i]`。
### 计算准确率
为了验证我们预测的结果,我们需要一个标准来给我们的预测打分。因为我们最感兴趣的是我们预测的**准确率**,既正确预测乘客存活的比例。运行下面的代码来创建我们的 `accuracy_score` 函数以对前五名乘客的预测来做测试。
**思考题**:在前五个乘客中,如果我们预测他们全部都存活,你觉得我们预测的准确率是多少?
*60%*
```
def accuracy_score(truth, pred):
""" 返回 pred 相对于 truth 的准确率 """
# 确保预测的数量与结果的数量一致
if len(truth) == len(pred):
# 计算预测准确率(百分比)
return "Predictions have an accuracy of {:.2f}%.".format((truth == pred).mean()*100)
else:
return "Number of predictions does not match number of outcomes!"
# 测试 'accuracy_score' 函数
predictions = pd.Series(np.ones(5, dtype = int)) #五个预测全部为1,既存活
print accuracy_score(outcomes[:5], predictions)
```
> **提示**:如果你保存 iPython Notebook,代码运行的输出也将被保存。但是,一旦你重新打开项目,你的工作区将会被重置。请确保每次都从上次离开的地方运行代码来重新生成变量和函数。
### 最简单的预测
如果我们要预测泰坦尼克号上的乘客是否存活,但是我们又对他们一无所知,那么最好的预测就是船上的人无一幸免。这是因为,我们可以假定当船沉没的时候大多数乘客都遇难了。下面的 `predictions_0` 函数就预测船上的乘客全部遇难。
```
def predictions_0(data):
""" 不考虑任何特征,预测所有人都无法生还 """
predictions = []
for _, passenger in data.iterrows():
# 预测 'passenger' 的生还率
predictions.append(0)
# 返回预测结果
return pd.Series(predictions)
# 进行预测
predictions = predictions_0(data)
```
**问题1**:对比真实的泰坦尼克号的数据,如果我们做一个所有乘客都没有存活的预测,这个预测的准确率能达到多少?
**回答**: *61.62%*
**提示**:运行下面的代码来查看预测的准确率。
```
print accuracy_score(outcomes, predictions)
```
### 考虑一个特征进行预测
我们可以使用 `survival_stats` 函数来看看 **Sex** 这一特征对乘客的存活率有多大影响。这个函数定义在名为 `titanic_visualizations.py` 的 Python 脚本文件中,我们的项目提供了这个文件。传递给函数的前两个参数分别是泰坦尼克号的乘客数据和乘客的 生还结果。第三个参数表明我们会依据哪个特征来绘制图形。
运行下面的代码绘制出依据乘客性别计算存活率的柱形图。
```
survival_stats(data, outcomes, 'Sex')
```
观察泰坦尼克号上乘客存活的数据统计,我们可以发现大部分男性乘客在船沉没的时候都遇难了。相反的,大部分女性乘客都在事故中**生还**。让我们以此改进先前的预测:如果乘客是男性,那么我们就预测他们遇难;如果乘客是女性,那么我们预测他们在事故中活了下来。
将下面的代码补充完整,让函数可以进行正确预测。
**提示**:您可以用访问 dictionary(字典)的方法来访问船上乘客的每个特征对应的值。例如, `passenger['Sex']` 返回乘客的性别。
```
def predictions_1(data):
""" 只考虑一个特征,如果是女性则生还 """
predictions = []
for _, passenger in data.iterrows():
# TODO 1
# 移除下方的 'pass' 声明
# 输入你自己的预测条件
if passenger['Sex'] == 'female':
predictions.append(1)
else:
predictions.append(0)
# 返回预测结果
return pd.Series(predictions)
# 进行预测
predictions = predictions_1(data)
```
**问题2**:当我们预测船上女性乘客全部存活,而剩下的人全部遇难,那么我们预测的准确率会达到多少?
**回答**: *78.68%*
**提示**:你需要在下面添加一个代码区域,实现代码并运行来计算准确率。
```
print accuracy_score(outcomes, predictions)
```
### 考虑两个特征进行预测
仅仅使用乘客性别(Sex)这一特征,我们预测的准确性就有了明显的提高。现在再看一下使用额外的特征能否更进一步提升我们的预测准确度。例如,综合考虑所有在泰坦尼克号上的男性乘客:我们是否找到这些乘客中的一个子集,他们的存活概率较高。让我们再次使用 `survival_stats` 函数来看看每位男性乘客的年龄(Age)。这一次,我们将使用第四个参数来限定柱形图中只有男性乘客。
运行下面这段代码,把男性基于年龄的生存结果绘制出来。
```
survival_stats(data, outcomes, 'Age', ["Sex == 'male'"])
```
仔细观察泰坦尼克号存活的数据统计,在船沉没的时候,大部分小于10岁的男孩都活着,而大多数10岁以上的男性都随着船的沉没而**遇难**。让我们继续在先前预测的基础上构建:如果乘客是女性,那么我们就预测她们全部存活;如果乘客是男性并且小于10岁,我们也会预测他们全部存活;所有其它我们就预测他们都没有幸存。
将下面缺失的代码补充完整,让我们的函数可以实现预测。
**提示**: 您可以用之前 `predictions_1` 的代码作为开始来修改代码,实现新的预测函数。
```
def predictions_2(data):
""" 考虑两个特征:
- 如果是女性则生还
- 如果是男性并且小于10岁则生还 """
predictions = []
for _, passenger in data.iterrows():
# TODO 2
# 移除下方的 'pass' 声明
# 输入你自己的预测条件
if passenger['Sex'] == 'female':
predictions.append(1)
elif passenger['Sex'] == 'male' and passenger['Age'] < 10:
predictions.append(1)
else:
predictions.append(0)
# 返回预测结果
return pd.Series(predictions)
# 进行预测
predictions = predictions_2(data)
#print(predictions)
```
**问题3**:当预测所有女性以及小于10岁的男性都存活的时候,预测的准确率会达到多少?
**回答**: *79.53%*
**提示**:你需要在下面添加一个代码区域,实现代码并运行来计算准确率。
```
print accuracy_score(outcomes, predictions)
```
### 你自己的预测模型
添加年龄(Age)特征与性别(Sex)的结合比单独使用性别(Sex)也提高了不少准确度。现在该你来做预测了:找到一系列的特征和条件来对数据进行划分,使得预测结果提高到80%以上。这可能需要多个特性和多个层次的条件语句才会成功。你可以在不同的条件下多次使用相同的特征。**Pclass**,**Sex**,**Age**,**SibSp** 和 **Parch** 是建议尝试使用的特征。
使用 `survival_stats` 函数来观测泰坦尼克号上乘客存活的数据统计。
**提示:** 要使用多个过滤条件,把每一个条件放在一个列表里作为最后一个参数传递进去。例如: `["Sex == 'male'", "Age < 18"]`
#### 观察特征 为 female,Age,Pclass
##### Pclass == 1,10岁以下的女性乘客都遇难了,我可以准确预测这部分乘客遇难,然后10岁以上的女性乘客我假设他们全部生还
```
survival_stats(data, outcomes, 'Age',["Sex == 'female'","Pclass == 1"])
survival_stats(data, outcomes, 'Age',["Sex == 'female'","Pclass == 2"])
```
##### Pclass == 3,40岁到50岁之间的女性乘客都遇难了,我可以准确预测这部分乘客遇难,然后小于40岁,大于50岁的女性乘客我假设他们全部生还
```
survival_stats(data, outcomes, 'Age',["Sex == 'female'","Pclass == 3"])
```
#### 观察特征为 male,Age,Pclass
##### Pclass == 1,10岁以下的男性乘客都生还了,我可以准确预测这部分乘客生还
##### Pclass == 1,30到40岁大部分的的男性乘客都生还了,我可以假设这部分乘客生还,然后其他年龄段的男性乘客我假设他们全部遇难
```
survival_stats(data, outcomes, 'Age',["Sex == 'male'","Pclass == 1"])
```
##### Pclass == 2,10岁以下的男性乘客都生还了,我可以准确预测这部分乘客生还,然后10岁以上的男性乘客我假设他们全部遇难
```
survival_stats(data, outcomes, 'Age',["Sex == 'male'","Pclass == 2"])
survival_stats(data, outcomes, 'Age',["Sex == 'male'","Pclass == 3"])
```
当查看和研究了图形化的泰坦尼克号上乘客的数据统计后,请补全下面这段代码中缺失的部分,使得函数可以返回你的预测。
在到达最终的预测模型前请确保记录你尝试过的各种特征和条件。
**提示:** 您可以用之前 `predictions_2` 的代码作为开始来修改代码,实现新的预测函数。
```
def predictions_3(data):
""" 考虑多个特征,准确率至少达到80% """
predictions = []
for _, passenger in data.iterrows():
# TODO 3
# 移除下方的 'pass' 声明
# 输入你自己的预测条件
if passenger['Sex'] == 'female':
if passenger['Pclass'] == 1 :
if passenger['Age'] < 10:
predictions.append(0)
else:
predictions.append(1)
elif passenger['Pclass'] == 3:
if passenger['Age'] < 50 and passenger['Age'] > 40:
predictions.append(0)
else:
predictions.append(1)
else:
predictions.append(1)
elif passenger['Sex'] == 'male':
if passenger['Pclass'] == 1 :
if passenger['Age'] < 10:
predictions.append(1)
elif passenger['Age'] < 40 and passenger['Age'] > 30:
predictions.append(1)
else:
predictions.append(0)
elif passenger['Pclass'] == 2:
if passenger['Age'] < 10:
predictions.append(1)
else:
predictions.append(0)
else:
predictions.append(0)
# 返回预测结果
return pd.Series(predictions)
# 进行预测
predictions = predictions_3(data)
```
**问题4**:请描述你实现80%准确度的预测模型所经历的步骤。您观察过哪些特征?某些特性是否比其他特征更有帮助?你用了什么条件来预测生还结果?你最终的预测的准确率是多少?
**回答**:
1. 根据之前的分析:运行代码survival_stats(data, outcomes, 'Sex')观察, 大部分男性乘客遇难,大部分女性乘客生还,据此把数据分为2组来观察,分别为男性乘客组合女性乘客组
2. 女性乘客组观察目标:找女性乘客遇难的条件, 我尝试过的条件有:
* 'Age',["Sex == 'female'","Pclass == 1"]
* 'Age',["Sex == 'female'","Pclass == 2"]
* 'Age',["Sex == 'female'","Pclass == 3"]
* 等等还有许多其他尝试
3. 通过仔细分析survival_stats生成的直方图可以发现:
* Pclass == 1,10岁以下的女性乘客都遇难了,我可以准确预测这部分乘客遇难,然后10岁以上的女性乘客我假设他们全部生还
* Pclass == 3,40岁到50岁之间的女性乘客都遇难了,我可以准确预测这部分乘客遇难,然后小于40岁,大于50岁的女性乘客我假设他们全部生还
* 然后其他条件下的女性乘客我假设他们全部生还
4. 男性乘客组观察目标:找男性乘客生还的条件,我尝试的条件有:
* 'Age',["Sex == 'male'","Pclass == 1"]
* 'Age',["Sex == 'male'","Pclass == 2"]
* 'Age',["Sex == 'male'","Pclass == 3"]
* 等等还有许多其他尝试
5. 通过仔细分析survival_stats生成的直方图可以发现:
* Pclass == 1,10岁以下的男性乘客都生还了,我可以准确预测这部分乘客生还
* Pclass == 1,30到40岁大部分的的男性乘客都生还了,我可以假设这部分乘客生还,然后其他年龄段的男性乘客我假设他们全部遇难
* Pclass == 2,10岁以下的男性乘客都生还了,我可以准确预测这部分乘客生还,然后10岁以上的男性乘客我假设他们全部遇难
* 然后其他条件下的男性乘客我假设他们全部遇难
最终准确率:**81.14%**
**提示**:你需要在下面添加一个代码区域,实现代码并运行来计算准确率。
```
print accuracy_score(outcomes, predictions)
```
### 结论
经过了数次对数据的探索和分类,你创建了一个预测泰坦尼克号乘客存活率的有用的算法。在这个项目中你手动地实现了一个简单的机器学习模型——决策树(*decision tree*)。决策树每次按照一个特征把数据分割成越来越小的群组(被称为 *nodes*)。每次数据的一个子集被分出来,如果分割后新子集之间的相似度比分割前更高(包含近似的标签),我们的预测也就更加准确。电脑来帮助我们做这件事会比手动做更彻底,更精确。[这个链接](http://www.r2d3.us/visual-intro-to-machine-learning-part-1/)提供了另一个使用决策树做机器学习入门的例子。
决策树是许多**监督学习**算法中的一种。在监督学习中,我们关心的是使用数据的特征并根据数据的结果标签进行预测或建模。也就是说,每一组数据都有一个真正的结果值,不论是像泰坦尼克号生存数据集一样的标签,或者是连续的房价预测。
**问题5**:想象一个真实世界中应用监督学习的场景,你期望预测的结果是什么?举出两个在这个场景中能够帮助你进行预测的数据集中的特征。
**回答**: *某网上书店记录了用户的年龄,性别,查询的书目类别等数据,我希望通过这些数据预测用户登录可能想买什么类别的书,从而在网店的首页向该用户推荐合适的书籍。
年龄,性别等特征可以帮助预测
比如 通过数据分析,发现女性用户20-30岁之间,买情世界名著比较多,既可以推荐这类
通过数据分析,发现男性用户25-35之间,买推理类书较多,既可以推荐这类书籍*
> **注意**: 当你写完了所有**5个问题,3个TODO**。你就可以把你的 iPython Notebook 导出成 HTML 文件。你可以在菜单栏,这样导出**File -> Download as -> HTML (.html)** 把这个 HTML 和这个 iPython notebook 一起做为你的作业提交。
---
翻译:毛礼建 | 校译:黄强 | 审译:曹晨巍
| github_jupyter |
# Rigid rods
This example demonstrates how to constrain particles in rigid bodies.
## Initialize
Import the hoomd python package and the md component to execute MD simulations.
```
from __future__ import division
import hoomd
import hoomd.md
```
Initialize the execution context to control where HOOMD will execute the simulation. When no command line options are provided, HOOMD will auto-select a GPU if it exists, or run on the CPU.
```
hoomd.context.initialize("");
```
The center of mass of each rigid body must be present in the system as a particle. The type assigned to this particle must be unique to all such center of mass particles for rigid bodies with the same geometry. The position and orientation of the center of mass particle, together with a definition of the local environment of the body define where all constituent particles should be placed. Constituent particles are also particles in the system.
First, create a simple lattice of the center of mass particles. These particles define the mass and moment of inertia of the rigid body.
```
uc = hoomd.lattice.unitcell(N = 1,
a1 = [10.8, 0, 0],
a2 = [0, 1.2, 0],
a3 = [0, 0, 1.2],
dimensions = 3,
position = [[0,0,0]],
type_name = ['R'],
mass = [1.0],
moment_inertia = [[0,
1/12*1.0*8**2,
1/12*1.0*8**2]],
orientation = [[1, 0, 0, 0]]);
system = hoomd.init.create_lattice(unitcell=uc, n=[2,18,18]);
```
This only creates the center of mass particles (type *R*).
Examine how the system configuration: [ex_render](ex_render.py) is a helper script that builds animated gifs from trajectory files and system snapshots. It is part of the [hoomd-examples](https://github.com/glotzerlab/hoomd-examples) repository and designed only to render these examples.
```
import ex_render
ex_render.render_sphere_frame(system.take_snapshot())
```
``md.constrain.rigid`` applies the rigid constraint to all the constituent particles, and can also create the constituent particles in the system.
Add the particle type for the constituent particles. Constituents may not share a type with any rigid central particle.
```
system.particles.types.add('A');
```
Define each rigid body type in the local coordinate system of the body. This example creates rods 9 particles long. The center of mass particle is at ``(0,0,0)`` and the constituent particles extend to either side.
Recall that the moment of inertia was specified as a length 3 vector [Ix, Iy, Iz]. The positions of the particles in the local reference frame must be such that the moment of inertia tensor is diagonal. The actual values specified need not assume anything about the body, such as point particles - HOOMD takes the moment of inertia as given and uses it directly in the equations of motion. Above, these bodies are initialized as if they are ideal line segments of length 8.
```
rigid = hoomd.md.constrain.rigid();
rigid.set_param('R',
types=['A']*8,
positions=[(-4,0,0),(-3,0,0),(-2,0,0),(-1,0,0),
(1,0,0),(2,0,0),(3,0,0),(4,0,0)]);
```
Instruct the rigid constraint to create the constituent particles.
```
rigid.create_bodies()
```
Now, the rods are complete.
```
import ex_render
ex_render.render_sphere_frame(system.take_snapshot())
```
## Define the potential energy
The WCA potential has the same functional form as Lennard-Jones, but $r_\mathrm{cut}=2^{1/6}\sigma$ and is shifted up to 0 at the cutoff.
Choose the neighbor list acceleration structure to find neighboring particles efficiently. In systems with only one cutoff length, the cell method performs best.
```
nl = hoomd.md.nlist.cell()
```
Define the functional form of the pair interaction and evaluate using the given neighbor list acceleration structure.
```
lj = hoomd.md.pair.lj(r_cut=2**(1/6), nlist=nl)
lj.set_params(mode='shift')
```
Specify pair potential parameters for every pair of types in the simulation. The particle types *R* and *A* exist in the system for the rigid bodies, but both should interact identically. The *R* particle in the center of the rod is a part of the rod.
```
lj.pair_coeff.set(['R', 'A'], ['R', 'A'], epsilon=1.0, sigma=1.0)
```
## Select integrator
The integrator defines the equations of motion that govern the system of particles, given the current configuration of the particles and the net force from all potentials. The standard integration mode in HOOMD allows different integrators to apply to different groups of particles with the same step size $dt$.
```
hoomd.md.integrate.mode_standard(dt=0.005);
```
Apply the Langevin equations of motion to all the rigid center of mass particles. $kT$ defines the temperature of the system in energy units and *seed* defines the seed for the random number generator. If there were non-rigid particles in this system, the same integrator could apply to both. Do **not** apply an integrator to constituent particles.
```
rigid = hoomd.group.rigid_center();
hoomd.md.integrate.langevin(group=rigid, kT=1.0, seed=42);
```
## Write output
Periodically log the potential energy of the system to a text file.
```
hoomd.analyze.log(filename="log-output.log",
quantities=['potential_energy',
'translational_kinetic_energy',
'rotational_kinetic_energy'],
period=100,
overwrite=True);
```
Periodically write the particle configurations to a gsd file.
```
hoomd.dump.gsd("trajectory.gsd",
period=2e3,
group=hoomd.group.all(),
overwrite=True);
```
## Run the simulation
Take 10,000 steps forward in time.
```
hoomd.run(1e4);
```
## Examine the output
Use matplotlib to plot the potential energy vs time step.
```
import numpy
from matplotlib import pyplot
%matplotlib inline
data = numpy.genfromtxt(fname='log-output.log', skip_header=True);
pyplot.figure(figsize=(4,2.2), dpi=140);
pyplot.plot(data[:,0], data[:,1]);
pyplot.xlabel('time step');
pyplot.ylabel('potential_energy');
```
Notice that the average rotational kinetic energy is 2/3 of the translational. This is consistent with the equipartition theorem: the rods have 3 translational degrees of freedom but only 2 rotational ones. HOOMD determined the number of rotational degrees of freedom from the moment of inertia tensor provided during initialization. The 0 moment of inertia about *x* notified HOOMD that the bodies should not rotate about that axis.
```
pyplot.figure(figsize=(4,2.2), dpi=140);
pyplot.plot(data[:,0], data[:,2]);
pyplot.plot(data[:,0], data[:,3]);
pyplot.xlabel('time step');
pyplot.ylabel('kinetic_energy');
pyplot.legend(['translational', 'rotational'], loc='lower right');
ex_render.display_movie(ex_render.render_sphere_frame, 'trajectory.gsd');
```
| github_jupyter |
# Training a Dense Neural Network
The handwritten digit recognition is a classification problem. We will start with the simplest possible approach for image classification - a fully-connected neural network (which is also called a *perceptron*). We use `pytorchcv` helper to load all data we have talked about in the previous unit.
```
import torch
import torch.nn as nn
import torchvision
import matplotlib.pyplot as plt
import pytorchcv
pytorchcv.load_mnist()
```
## Fully-Connected Dense Neural Network
A simplest **neural network** in PyTorch consists of a number of **layers**. The simplest network would include just one fully-connected layer, which is called **Linear** layer, with 784 inputs (one input for each pixel of the input image) and 10 outputs (one output for each class).

As we discussed above, the dimension of our digit images is $1\times28\times28$. Because the input dimension of fully-connected layer is 784, we need to insert another layer into the network, called **Flatten**, to change tensor shape from $1\times28\times28$ to $784$.
We want $n$-th output of the network to return the probability of the input digit being equal to $n$. Because output of fully-connected layer is not normalized in any way, it cannot be regarded as probablility. To turn it into a probability, we apply another layer, called **Softmax**.
> In PyTorch, it is more convenient to use **LogSoftmax** function, which will additionally compute logarithms of output probabilities. To turn the output vector into the actual probablities, we need to take **torch.exp** of the ouput.
Thus, the architecture of our network can be represented by the following sequence of layers:

It can be defined in PyTorch in the following way, using `Sequential` syntax:
```
net = nn.Sequential(
nn.Flatten(),
nn.Linear(784,10), # 784 inputs, 10 outputs
nn.LogSoftmax())
```
## Training the Network
A network defined in this manner can take any digit as input, and produce a vector of probabilities as an output. Let's see how this network performs, but giving it a digit from our dataset:
```
print('Digit to be predicted: ',data_train[0][1])
torch.exp(net(data_train[0][0]))
```
As you can see, the network predicts almost equal probablilites for each digit, because it is not trained. To train the network, we need to present it with our training data.
To do so, we will need to chunk original dataset into **batches** of a certain size, let's say 64. An object called **DataLoader** can do it for us automatically:
```
train_loader = torch.utils.data.DataLoader(data_train,batch_size=64)
test_loader = torch.utils.data.DataLoader(data_test,batch_size=64) # we can use larger batch size for testing
```
The process of training looks like this:
1. We take a minibatch from the input dataset, which consists of input data (features) and expected result (label).
2. We calculate the result that network predicts on this minibatch.
3. The difference between this result and expected result is calculated using a special function called **loss function**
4. We calculate the gradients of this loss function with respect to model weights (parameters), which are then used to adjust the weights to optimize the preformance of the network. The amount of adjustment is controlled by a parameter called **learning rate**, and the details of optimization alogrithm are defined in the **optimizer** object.
5. We repeat those steps until the whole dataset is used.
Complete pass through the dataset is called **an epoch**. Here is a function that performs one epoch training:
```
def train_epoch(net,dataloader,lr=0.01,optimizer=None,loss_fn = nn.NLLLoss()):
optimizer = optimizer or torch.optim.Adam(net.parameters(),lr=lr)
net.train()
total_loss,acc,count = 0,0,0
for features,labels in dataloader:
optimizer.zero_grad()
out = net(features)
loss = loss_fn(out,labels) #cross_entropy(out,labels)
loss.backward()
optimizer.step()
total_loss+=loss
_,predicted = torch.max(out,1)
acc+=(predicted==labels).sum()
count+=len(labels)
return total_loss.item()/count, acc.item()/count
train_epoch(net,train_loader)
```
This function is pretty generic, which will allow us to use it later in all our examples. It takes the following parameters:
* **Neural network** itself
* **DataLoader**, which defines the data to train on
* **Loss Function**, which is a function that measures the difference between expected result and the one produced by the network. In most of the classification tasks `NLLLoss` is used, so we will make it a default.
* **Optimizer**, which defined an *optimization algorithm*. The most traditional algorithm is *stochastic gradient descent*, but we will use more advanced version called **Adam** by default.
* **Learning rate** defines the speed at which network learns. During learning, we show the same data multiple times, and each time weights are adjusted. If learning rate is too high, new values will overwrite the knowledge from the old ones, and the network would behave badly. Too small learning rate results in very slow learning.
Here is what we do when training:
* Switch the network to training mode (`net.train()`)
* Go over all batches in the dataset, and for each batch do the following:
- compute predictions made by the network on this batch (`out`)
- compute `loss`, which is the discrepancy between predicted and expected values
- try to minimize the loss by adjusting weights of the network (`optimizer.step()`)
- compute the number of correctly predicted cases (**accuracy**)
The function calculates and returns the average loss per one data item, and training accuracy (percentage of cases guessed correctly). By observing this loss during training we can see whether network actually learns something.
It is also important to control the accuracy on the test dataset (also called **validation accuracy**). A rich neural network with a lot of parameters can approximate any training dataset very well, but it may poorly generalize to other data. That's why in most cases we set aside part of our data, and then periodically check how well the model performs on them. Here is the function to evaluate the network on test dataset:
```
def validate(net, dataloader,loss_fn=nn.NLLLoss()):
net.eval()
count,acc,loss = 0,0,0
with torch.no_grad():
for features,labels in dataloader:
out = net(features)
loss += loss_fn(out,labels)
pred = torch.max(out,1)[1]
acc += (pred==labels).sum()
count += len(labels)
return loss.item()/count, acc.item()/count
validate(net,test_loader)
```
In most of the cases, we train the model for several epochs, observing training and validation accuracy. Normally, training accuracy will tend to increase, while validation accuracy may start to decrease at some point. That would be an indication of **overfitting**, and we probably need to stop training.
Below is the training function that can be used to perform both training and validation. It prints the training and validation accuracy for each epoch, and also returns the history that can be used to plot the loss and accuracy on the graph.
```
def train(net,train_loader,test_loader,optimizer=None,lr=0.01,epochs=10,loss_fn=nn.NLLLoss()):
optimizer = optimizer or torch.optim.Adam(net.parameters(),lr=lr)
res = { 'train_loss' : [], 'train_acc': [], 'val_loss': [], 'val_acc': []}
for ep in range(epochs):
tl,ta = train_epoch(net,train_loader,optimizer=optimizer,lr=lr,loss_fn=loss_fn)
vl,va = validate(net,test_loader,loss_fn=loss_fn)
print(f"Epoch {ep:2}, Train acc={ta:.3f}, Val acc={va:.3f}, Train loss={tl:.3f}, Val loss={vl:.3f}")
res['train_loss'].append(tl)
res['train_acc'].append(ta)
res['val_loss'].append(vl)
res['val_acc'].append(va)
return res
# Re-initialize the network to start from scratch
net = nn.Sequential(
nn.Flatten(),
nn.Linear(784,10), # 784 inputs, 10 outputs
nn.LogSoftmax())
hist = train(net,train_loader,test_loader,epochs=5)
```
This function prints diagnostic messages with accuracy on training and valiadation data obtained on each epoch, and also returns this data as a dictionary (so-called **history**). We can then visualize this data to make sense of what goes on:
```
plt.figure(figsize=(15,5))
plt.subplot(121)
plt.plot(hist['train_acc'], label='Training acc')
plt.plot(hist['val_acc'], label='Validation acc')
plt.legend()
plt.subplot(122)
plt.plot(hist['train_loss'], label='Training loss')
plt.plot(hist['val_loss'], label='Validation loss')
plt.legend()
```
What we can observe in most of the cases is that training accuracy always increases (which corresponds to the network learning to classify our training data better and better), while validation accuracy may start to fall. This would indicate **overfitting**, and that is the point where we need to stop training.
## Visualizing Network Weights
It may be interesting to see if weights of a neural network make any sense. In most of the cases, when network is more complex than just one layer, it can be a difficult thing to do; however, in our case classification of a digit happpens by multiplying the initial image by a weight matrix.
Let's denote a weight tensor as `weight_tensor`, which will have a dimension of 784x10. This tensor can be obtained by calling `net.parameters()` method of our neural network. In this example, if we want to see if our number if 0 or not, we will multiply input digit by `weight_tensor[0]`, and pass the result through softmax normalization to get the answer. So we may expect that weight tensor elements might somehow resemble the average shape of the digit they classify:
```
weight_tensor = next(net.parameters())
fig,ax = plt.subplots(1,10,figsize=(15,4))
for i,x in enumerate(weight_tensor):
ax[i].imshow(x.view(28,28).detach())
```
## Takeaway
Training a neural netowork in PyTorch can be programmed with a training loop. It may seem like a compicated process, but in real life we need to write it once, and we can then re-use this training code later in unchanged form.
We can see that single-layer dense neural network shows relatively good performance, but we definitely want to get higher than 91% on accuracy! In the next unit, we will try to use multi-level perceptrons.
| github_jupyter |
```
%autosave 60
%load_ext autoreload
%autoreload 2
%matplotlib inline
import json
import os
import pickle
from collections import Counter, OrderedDict
from copy import deepcopy
from pathlib import Path
from typing import Any, Dict, List, Optional, Tuple, Union, cast
import cv2
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import PIL.Image as pil_img
import seaborn as sns
import sklearn as skl
import torch
import torch.nn as nn
from IPython.display import Image, display
from matplotlib.patches import Rectangle
from matplotlib_inline.backend_inline import set_matplotlib_formats
from torch.nn import functional as F
from tqdm.contrib import tenumerate, tmap, tzip
from tqdm.contrib.bells import tqdm, trange
pd.set_option("display.max_colwidth", None)
pd.set_option("display.max_columns", 15)
pd.set_option("display.max_rows", 50)
# Suitable default display for floats
pd.options.display.float_format = "{:,.2f}".format
plt.rcParams["figure.figsize"] = (12, 10)
# This one is optional -- change graphs to SVG only use if you don't have a
# lot of points/lines in your graphs. Can also just use ['retina'] if you
# don't want SVG.
%config InlineBackend.figure_formats = ["retina"]
set_matplotlib_formats("pdf", "png")
def get_sims(a: torch.tensor, b: torch.tensor, batch_size=1000):
sims = {i: (0, -1) for i in range(a.shape[0])}
for j in tqdm(range(0, b.shape[0], batch_size)):
batch_sims = a @ b[j : j + batch_size].T
batch_sims = torch.from_numpy(batch_sims)
values, idxs = batch_sims.max(dim=-1)
# Append an index and sim every iteration
for i in range(a.shape[0]):
offset = j + idxs[i].item()
if values[i] > sims[i][0]:
sims[i] = (values[i].item(), offset)
return sims
def normalize_rows(mat: torch.tensor) -> None:
for i in range(len(mat)):
mat[i] /= mat[i].norm(p=2, dim=-1, keepdim=True)
return mat
```
---
## Geoguessr In-game Frames -> GPT-J Text Lookup
```
split = "test"
# Load query image embeddings:
image_embs = pickle.load(
open(
f"/shared/gbiamby/geo/models/clip_ft/vit-b32/geoframes_clip_samples_fixed_window_{split}_img.pkl",
"rb",
)
)
text_embs = pickle.load(
open(f"/shared/gbiamby/geo/models/clip_ft/vit-b32/gptj_clues_text.pkl", "rb")
)
# load target captions:
gpt_caps = json.load(open("/shared/g-luo/geoguessr/data/data/guidebook/kb/v3/cleaned_clues.json"))["clues"]
# # Append embeddings
# for caption in gpt_caps:
# caption["clip_emb"] = image_embs[caption["file_path"]]
t_image_embs = normalize_rows(
torch.stack([torch.tensor(emb) for emb in image_embs.values()]).to("cuda")
)
t_text_embs = normalize_rows(
torch.stack([torch.tensor(emb) for emb in text_embs.values()]).to("cuda")
)
print(f"image_embs.shape: {t_image_embs.shape}, text_emb.shape: {t_text_embs.shape}")
sims = torch.mm(t_image_embs, t_text_embs.T)
max_sim_scores, max_sim_idxs = sims.max(dim=1)
texts_unique = list(text_embs.keys())
img_to_text_sims = {}
for i, img_path in enumerate(image_embs.keys()):
img_to_text_sims[img_path] = {
"best_match_text": texts_unique[max_sim_idxs[i]],
"best_sim_score": max_sim_scores[i].tolist(),
"best_sim_idx": max_sim_idxs[i].tolist(),
"sims_all": sims[i].tolist(),
"file_path": img_path,
}
print(t_image_embs.shape, t_text_embs.shape, sims.shape)
# list(img_to_text_sims.items())[:2]
from IPython.core.display import HTML, Markdown
def show_samples(img_to_text_sims, n_samples: int = 20):
df_random = np.random.choice(range(len(img_to_text_sims)), n_samples, replace=False)
for i in df_random:
img_row = deepcopy(img_to_text_sims[i])
del img_row["sims_all"]
print("=" * 180)
# print(img_row.keys())
# print(img_row)
display(pd.DataFrame({k: [v] for k, v in img_row.items()}).T)
img = pil_img.open(img_row["file_path"])
img.thumbnail((1080, 640), pil_img.NEAREST)
display(img)
print("\n")
```
### Choose Images w/ Highest Img/Text Similarity Scores
```
show_samples(
sorted(img_to_text_sims.values(), key=lambda x: x["best_sim_score"], reverse=True)[:100],
n_samples=20,
)
```
### Choose Random Images, show best match
```
show_samples(list(img_to_text_sims.values()), n_samples=20)
```
---
| github_jupyter |
# Results
In this notebook we will learn the following about the Results class returned a Benchmark evaluation:
1. **Result Creation**: How to create Results from a Benchmark and Learners.
2. **Result Plotting**: How to create plots of the learners that were evaluted by a benchmark
3. **Result Saving/Loading**: How to save and load results to and from disk
4. **Result to Pandas**: How to turn Result into a collection of Pandas dataframes for more in-depth analysis
## Result Creation
**Results** are created whenever Learners are evaluated on a Benchmark. Below is a simple example of this:
```
from coba.simulations import ValidationSimulation
from coba.learners import RandomLearner, VowpalLearner
from coba.benchmarks import Benchmark
result = Benchmark([ValidationSimulation()]).evaluate([RandomLearner(), VowpalLearner()])
```
## Result Plotting
The **Result** class is capable of creating two separate plots: 1) `plot_learners` and `plot_shuffles`.
### Plot Learners
Perhaps the most straightforward question after running a Benchmark is asking which Learners performed best.
To answer this question the **Result** has a `plot_learners` method.
```
from coba.simulations import ValidationSimulation
from coba.learners import RandomLearner, VowpalLearner
from coba.benchmarks import Benchmark
result = Benchmark([ValidationSimulation()], shuffle=[1,2,3,4]).evaluate([RandomLearner(), VowpalLearner()])
result.plot_learners()
```
The `plot_learners` method has many arguments which allow customization of the plot.
```python
"""
Args:
source_pattern: The pattern to match when determining which simulations to include in the plot. The "source"
matched against is either the "source" column in the simulations table or the first item in the list in
the simulation 'pipes' column. The simulations can be seen most easily by Result.simulations.to_pandas().
learner_pattern: The pattern to match against the 'full_name' column in learners to determine which learners
to include in the plot. In the case of multiple matches only the last match is kept. The learners table in
Result can be examined via result.learners.to_pandas().
span: In general this indicates how many previous evaluations to average together. In practice this works
identically to ewm span value in the Pandas API. Additionally, if span equals None then all previous
rewards are averaged together and that value is plotted. Compare this to span = 1 WHERE only the current
reward is plotted for each interaction.
start: Determines at which interaction the plot will start at. If start is greater than 1 we assume start is
an interaction index. If start is less than 1 we assume start is the percent of interactions to skip
before starting the plot.
err_every: Determines frequency of bars indicating the standard deviation of the population should be drawn.
Standard deviation gives a sense of how well the plotted average represents the underlying distribution.
Standard deviation is most valuable when plotting against multiple simulations. If plotting against a single
simulation standard error may be a more useful indicator of confidence. The value for sd_every should be
between 0 to 1 and will determine how frequently the standard deviation bars are drawn.
err_type: Determines what the error bars are. Valid types are `None`, 'se', and 'sd'. If err_type is None then
plot will use SEM when there is only one source simulation otherwise it will use SD. Otherwise plot will
display the standard error of the mean for 'se' and the standard deviation for 'sd'.
"""
```
For example, we can examine learner performance at the end of the Benchmark and compare smoothed reward to their instantaneous reward.
```
result.plot_learners(start=0.8, end=1.0, span=1 , err_every=0 )
result.plot_learners(start=0.8, end=1.0, span=40, err_every=0.1, err_type='se')
```
### Plot Shuffles
In addition to plotting average Learner performance we can also zoom in on a single learner and superimpose its average performance on top of each of the individual shuffles that we included in our Benchmark. In our current results we have 4 shuffle seeds so there are four different performance results which are averaged together.
The `plot_shuffles` command has all the same arguments are `plot_learners`. Additionally, `plot_shuffles` assumes there's only one source of shuffles and one learner. For this reason you should pass in a source pattern and a learner pattern if there are multiple learners or simulation sources contained in the result.
```
result.plot_shuffles(learner_pattern="vw")
```
## Result Saving/Loading
Saving and restoring a **Result** is the easiest way to archive and share results with others.
Saving is done via Benchmark `evaluation`. The following code block will save the Result of the benchmark evaluation in result.log
```
from coba.simulations import ValidationSimulation
from coba.learners import RandomLearner, VowpalLearner
from coba.benchmarks import Benchmark
simulations = [ValidationSimulation() ]
learners = [RandomLearner(), VowpalLearner()]
result = Benchmark(simulations, shuffle=[1,2,3]).evaluate(learners, "result.log")
```
After a result has been saved it can be restored using `Result.from_file`.
```
from coba.benchmarks import Result
Result.from_file('result.log').plot_learners()
```
## Result To Pandas
Finally, if the plotting functionality just shared isn't enough it is possible to extract all the data stored in Result into dataframes.
```
from coba.benchmarks import Result
Result.from_file('result.log').simulations.to_pandas()
from coba.benchmarks import Result
Result.from_file('result.log').learners.to_pandas()
from coba.benchmarks import Result
Result.from_file('result.log').interactions.to_pandas()
```
| github_jupyter |
# Single Stepping `logictools` Generators
This notebook will show how to use single stepping mode in `logictools` generators. In this example the pattern generator is used to implement a simple 4-bit up-counter, we will be able to single step the clock and verify the counter operation. The output is verifired using the waveforms captured by the trace analyzer as well as the patterns on the on-board LEDs.
We use the boolean generator to transfer the pattern generator signals to the on-board LEDs. It implements 4 buffer functions that pass the input values received from the pattern generator to the on-board LEDs. we observe the patterns on the on-board LEDs by single stepping the clock at desired intervals.
Single stepping is supported in all the `logictools` generators.
### Step 1: Download the `logictools` overlay
```
from pynq.overlays.logictools import LogicToolsOverlay
logictools_olay = LogicToolsOverlay('logictools.bit')
```
### Step 2: Create WaveJSON waveform
The pattern to be generated is specified in the waveJSON format
The pattern is applied to the Arduino interface, pins **D0**, **D1**, **D2** and **D3** are set to generate a 4-bit count.
The Waveform class is used to display the specified waveform.
```
from pynq.lib.logictools import Waveform
up_counter = {'signal': [
['stimulus',
{'name': 'bit0', 'pin': 'D0', 'wave': 'lh' * 8},
{'name': 'bit1', 'pin': 'D1', 'wave': 'l.h.' * 4},
{'name': 'bit2', 'pin': 'D2', 'wave': 'l...h...' * 2},
{'name': 'bit3', 'pin': 'D3', 'wave': 'l.......h.......'}],
{},
['analysis',
{'name': 'bit0_loopback', 'pin': 'D0'},
{'name': 'bit1_loopback', 'pin': 'D1'},
{'name': 'bit2_loopback', 'pin': 'D2'},
{'name': 'bit3_loopback', 'pin': 'D3'}]],
'foot': {'tock': 1},
'head': {'text': 'up_counter'}}
waveform = Waveform(up_counter)
waveform.display()
```
**Note:** Since there are no captured samples at this moment, the analysis group will be empty.
### Step 3: Instantiate and setup the pattern generator and trace analyzer objects
Users can choose whether to use the trace analyzer by calling the `trace()` method.
The analyzer can be set to trace a specific number of samples using, `num_analyzer_samples` argument.
```
pattern_generator = logictools_olay.pattern_generator
pattern_generator.trace(num_analyzer_samples=16)
pattern_generator.setup(up_counter,
stimulus_group_name='stimulus',
analysis_group_name='analysis')
```
### Step 4: Setup the boolean generator
```
boolean_generator = logictools_olay.boolean_generator
functions = {'Buffer 1': 'LD3 = D16',
'Buffer 2': 'LD2 = D17',
'Buffer 3': 'LD1 = D18',
'Buffer 4': 'LD0 = D19'}
boolean_generator.setup(functions)
boolean_generator.expressions
```
### Step 5: Instantiate logictools controller to run both the instantiated generators simultaneously
```
logictools_controller = logictools_olay.logictools_controller
```
__Set the loopback connections using jumper wires on the Arduino Interface__

* __Output pins D0, D1, D2 and D3 are connected to pins D19, D18, D17 and D16 respectively__
* __Internal Loopback pins D0, D1, D2 and D3 are observed using the trace analyzer as shown below__
* __After setup, the pattern generator should be ready to run__
* __Patterns on pins D19, D18, D17 and D16 are observed on the on-board LEDs__
**Note:** Make sure all other pins are disconnected.
### Step 6: Step and display waveform
The `step()` method is used to single step the pattern, `show_waveform()` method is used to display the waveforms.
__`Step 4 cycles using a 1 second delay loop. Observe the pattern on the on-board LEDs`__
```
from time import sleep
for _ in range(4):
logictools_controller.step([boolean_generator, pattern_generator])
sleep(1)
pattern_generator.show_waveform()
```
__`Step an additional 11 cycles using a 1 second delay loop. Observe the pattern on the on-board LEDs`__
```
from time import sleep
for _ in range(11):
logictools_controller.step([boolean_generator, pattern_generator])
sleep(1)
pattern_generator.show_waveform()
```
__`Step 1 additional cycle to reach the maximum count value. Observe the pattern on the on-board LEDs`__
```
logictools_controller.step([boolean_generator, pattern_generator])
pattern_generator.show_waveform()
```
### Step 7: Stop the generators
Calling `stop()` will clear the logic values on output pins; however, the waveform will be recorded locally in the pattern generator instance.
```
pattern_generator.stop()
boolean_generator.stop()
```
### Step 8: Reset the generators
Calling `reset()` will reset generator instances.
```
pattern_generator.reset()
boolean_generator.reset()
```
| github_jupyter |
```
"""
File : lineNumberDriversOverTime.ipynb
Plot : line plot of the average number of drivers harboured in a tumour section with respect to the tumour size
Input :
SourceData_ExtendedData_Fig8c.xlsx,
"""
import pandas as pd
import matplotlib.pyplot as plt
import os
path_to_all_source_data = "../Source data for figures"
def read_and_plot(
sheet_names,
driver_advantage_types
):
def config_font_size(figsize):
params = {'legend.fontsize': 6,
'figure.figsize': figsize,
'axes.labelsize': 8,
'axes.titlesize': 8,
'xtick.labelsize':6,
'ytick.labelsize':6,
'pdf.fonttype':42}
plt.rcParams.update(params)
for sheet_name, driver_advantage_type in zip(
excelfile.sheet_names, driver_advantage_types
):
data = pd.read_excel(
excelfile,
sheet_name=sheet_name
)
print(sheet_name)
dict_results_per_sim_pdriver6e_4_Surf = {
simid : data.loc[
data.SimID == simid
]
for simid in data.loc[
data.Condition == 'Surface'
].SimID.unique()
}
dict_results_per_sim_pdriver6e_4_Volu = {
simid : data.loc[
data.SimID == simid
]
for simid in data.loc[
data.Condition == 'Volume'
].SimID.unique()
}
fig = plt.figure(dpi=300)
ax = fig.add_axes([0.2,0.2,0.75,0.75])
config_font_size(figsize = (2.75,3))
pdriver = 6e-4
ax.set_xlabel('Diameter of tumour slice (mm)')
ax.set_ylabel('Average number of drivers in tumour voxels')
ax.set_xlim([0,150])
# surface
for sim in dict_results_per_sim_pdriver6e_4_Surf.keys():
results_per_sim = dict_results_per_sim_pdriver6e_4_Surf[sim]
ax.plot(results_per_sim.SliceDiameter, results_per_sim.DriverNumberMean,
marker='o', linewidth = 0.2, markersize=0.5, color = "red")
# volume
for sim in dict_results_per_sim_pdriver6e_4_Volu.keys():
results_per_sim = dict_results_per_sim_pdriver6e_4_Volu[sim]
ax.plot(results_per_sim.SliceDiameter, results_per_sim.DriverNumberMean,
marker='o', linewidth = 0.2, markersize=0.5, color = "blue")
plt.show()
```
# Extended Data Figure 8c
```
path_to_excelfile = os.path.join(
path_to_all_source_data,
"Source_Data_Extended_Data_Figure_8",
"SourceData_ExtendedData_Fig8c.xlsx"
)
excelfile = pd.ExcelFile(path_to_excelfile)
excelfile.sheet_names
read_and_plot(
excelfile.sheet_names,
['saturated', 'additive', 'additive']
)
```
| github_jupyter |
# Breast Cancer Prediction
_**Predict Breast Cancer using SageMaker's Linear-Learner with features derived from images of Breast Mass**_
---
---
## Contents
1. [Background](#Background)
1. [Setup](#Setup)
1. [Data](#Data)
1. [Train](#Train)
1. [Host](#Host)
1. [Predict](#Predict)
1. [Extensions](#Extensions)
---
## Background
This notebook illustrates how one can use SageMaker's algorithms for solving applications which require `linear models` for prediction. For this illustration, we have taken an example for breast cancer prediction using UCI'S breast cancer diagnostic data set available at https://archive.ics.uci.edu/ml/datasets/Breast+Cancer+Wisconsin+%28Diagnostic%29. The data set is also available on Kaggle at https://www.kaggle.com/uciml/breast-cancer-wisconsin-data. The purpose here is to use this data set to build a predictve model of whether a breast mass image indicates benign or malignant tumor. The data set will be used to illustrate
* Basic setup for using SageMaker.
* converting datasets to protobuf format used by the Amazon SageMaker algorithms and uploading to S3.
* Training SageMaker's linear learner on the data set.
* Hosting the trained model.
* Scoring using the trained model.
---
## Setup
Let's start by specifying:
* The SageMaker role arn used to give learning and hosting access to your data. The snippet below will use the same role used by your SageMaker notebook instance, if you're using other. Otherwise, specify the full ARN of a role with the SageMakerFullAccess policy attached.
* The S3 bucket that you want to use for training and storing model objects.
```
import os
import boto3
import re
from sagemaker import get_execution_role
role = get_execution_role()
bucket = '<your_s3_bucket_name_here>'# enter your s3 bucket where you will copy data and model artifacts
prefix = 'sagemaker/breast_cancer_prediction' # place to upload training files within the bucket
```
Now we'll import the Python libraries we'll need.
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import io
import time
import json
import sagemaker.amazon.common as smac
```
---
## Data
Data Source: https://archive.ics.uci.edu/ml/machine-learning-databases/breast-cancer-wisconsin/wdbc.data
https://www.kaggle.com/uciml/breast-cancer-wisconsin-data
Let's download the data and save it in the local folder with the name data.csv and take a look at it.
```
data = pd.read_csv('https://archive.ics.uci.edu/ml/machine-learning-databases/breast-cancer-wisconsin/wdbc.data', header = None)
# specify columns extracted from wbdc.names
data.columns = ["id","diagnosis","radius_mean","texture_mean","perimeter_mean","area_mean","smoothness_mean",
"compactness_mean","concavity_mean","concave points_mean","symmetry_mean","fractal_dimension_mean",
"radius_se","texture_se","perimeter_se","area_se","smoothness_se","compactness_se","concavity_se",
"concave points_se","symmetry_se","fractal_dimension_se","radius_worst","texture_worst",
"perimeter_worst","area_worst","smoothness_worst","compactness_worst","concavity_worst",
"concave points_worst","symmetry_worst","fractal_dimension_worst"]
# save the data
data.to_csv("data.csv", sep=',', index=False)
# print the shape of the data file
print(data.shape)
# show the top few rows
display(data.head())
# describe the data object
display(data.describe())
# we will also summarize the categorical field diganosis
display(data.diagnosis.value_counts())
```
#### Key observations:
* Data has 569 observations and 32 columns.
* First field is 'id'.
* Second field, 'diagnosis', is an indicator of the actual diagnosis ('M' = Malignant; 'B' = Benign).
* There are 30 other numeric features available for prediction.
## Create Features and Labels
#### Split the data into 80% training, 10% validation and 10% testing.
```
rand_split = np.random.rand(len(data))
train_list = rand_split < 0.8
val_list = (rand_split >= 0.8) & (rand_split < 0.9)
test_list = rand_split >= 0.9
data_train = data[train_list]
data_val = data[val_list]
data_test = data[test_list]
train_y = ((data_train.iloc[:,1] == 'M') +0).as_matrix();
train_X = data_train.iloc[:,2:].as_matrix();
val_y = ((data_val.iloc[:,1] == 'M') +0).as_matrix();
val_X = data_val.iloc[:,2:].as_matrix();
test_y = ((data_test.iloc[:,1] == 'M') +0).as_matrix();
test_X = data_test.iloc[:,2:].as_matrix();
```
Now, we'll convert the datasets to the recordIO-wrapped protobuf format used by the Amazon SageMaker algorithms, and then upload this data to S3. We'll start with training data.
```
train_file = 'linear_train.data'
f = io.BytesIO()
smac.write_numpy_to_dense_tensor(f, train_X.astype('float32'), train_y.astype('float32'))
f.seek(0)
boto3.Session().resource('s3').Bucket(bucket).Object(os.path.join(prefix, 'train', train_file)).upload_fileobj(f)
```
Next we'll convert and upload the validation dataset.
```
validation_file = 'linear_validation.data'
f = io.BytesIO()
smac.write_numpy_to_dense_tensor(f, val_X.astype('float32'), val_y.astype('float32'))
f.seek(0)
boto3.Session().resource('s3').Bucket(bucket).Object(os.path.join(prefix, 'validation', train_file)).upload_fileobj(f)
```
---
## Train
Now we can begin to specify our linear model. Amazon SageMaker's Linear Learner actually fits many models in parallel, each with slightly different hyperparameters, and then returns the one with the best fit. This functionality is automatically enabled. We can influence this using parameters like:
- `num_models` to increase to total number of models run. The specified parameters will always be one of those models, but the algorithm also chooses models with nearby parameter values in order to find a solution nearby that may be more optimal. In this case, we're going to use the max of 32.
- `loss` which controls how we penalize mistakes in our model estimates. For this case, let's use absolute loss as we haven't spent much time cleaning the data, and absolute loss will be less sensitive to outliers.
- `wd` or `l1` which control regularization. Regularization can prevent model overfitting by preventing our estimates from becoming too finely tuned to the training data, which can actually hurt generalizability. In this case, we'll leave these parameters as their default "auto" though.
### Specify container images used for training and hosting SageMaker's linear-learner
```
# See 'Algorithms Provided by Amazon SageMaker: Common Parameters' in the SageMaker documentation for an explanation of these values.
containers = {'us-west-2': '174872318107.dkr.ecr.us-west-2.amazonaws.com/linear-learner:latest',
'us-east-1': '382416733822.dkr.ecr.us-east-1.amazonaws.com/linear-learner:latest',
'us-east-2': '404615174143.dkr.ecr.us-east-2.amazonaws.com/linear-learner:latest',
'eu-west-1': '438346466558.dkr.ecr.eu-west-1.amazonaws.com/linear-learner:latest'}
linear_job = 'linear-' + time.strftime("%Y-%m-%d-%H-%M-%S", time.gmtime())
print("Job name is:", linear_job)
linear_training_params = {
"RoleArn": role,
"TrainingJobName": linear_job,
"AlgorithmSpecification": {
"TrainingImage": containers[boto3.Session().region_name],
"TrainingInputMode": "File"
},
"ResourceConfig": {
"InstanceCount": 1,
"InstanceType": "ml.c4.2xlarge",
"VolumeSizeInGB": 10
},
"InputDataConfig": [
{
"ChannelName": "train",
"DataSource": {
"S3DataSource": {
"S3DataType": "S3Prefix",
"S3Uri": "s3://{}/{}/train/".format(bucket, prefix),
"S3DataDistributionType": "ShardedByS3Key"
}
},
"CompressionType": "None",
"RecordWrapperType": "None"
},
{
"ChannelName": "validation",
"DataSource": {
"S3DataSource": {
"S3DataType": "S3Prefix",
"S3Uri": "s3://{}/{}/validation/".format(bucket, prefix),
"S3DataDistributionType": "FullyReplicated"
}
},
"CompressionType": "None",
"RecordWrapperType": "None"
}
],
"OutputDataConfig": {
"S3OutputPath": "s3://{}/{}/".format(bucket, prefix)
},
"HyperParameters": {
"feature_dim": "30",
"mini_batch_size": "100",
"predictor_type": "regressor",
"epochs": "10",
"num_models": "32",
"loss": "absolute_loss"
},
"StoppingCondition": {
"MaxRuntimeInSeconds": 60 * 60
}
}
```
Now let's kick off our training job in SageMaker's distributed, managed training, using the parameters we just created. Because training is managed, we don't have to wait for our job to finish to continue, but for this case, let's use boto3's 'training_job_completed_or_stopped' waiter so we can ensure that the job has been started.
```
%%time
region = boto3.Session().region_name
sm = boto3.client('sagemaker')
sm.create_training_job(**linear_training_params)
status = sm.describe_training_job(TrainingJobName=linear_job)['TrainingJobStatus']
print(status)
sm.get_waiter('training_job_completed_or_stopped').wait(TrainingJobName=linear_job)
if status == 'Failed':
message = sm.describe_training_job(TrainingJobName=linear_job)['FailureReason']
print('Training failed with the following error: {}'.format(message))
raise Exception('Training job failed')
```
---
## Host
Now that we've trained the linear algorithm on our data, let's setup a model which can later be hosted. We will:
1. Point to the scoring container
1. Point to the model.tar.gz that came from training
1. Create the hosting model
```
linear_hosting_container = {
'Image': containers[boto3.Session().region_name],
'ModelDataUrl': sm.describe_training_job(TrainingJobName=linear_job)['ModelArtifacts']['S3ModelArtifacts']
}
create_model_response = sm.create_model(
ModelName=linear_job,
ExecutionRoleArn=role,
PrimaryContainer=linear_hosting_container)
print(create_model_response['ModelArn'])
```
Once we've setup a model, we can configure what our hosting endpoints should be. Here we specify:
1. EC2 instance type to use for hosting
1. Initial number of instances
1. Our hosting model name
```
linear_endpoint_config = 'linear-endpoint-config-' + time.strftime("%Y-%m-%d-%H-%M-%S", time.gmtime())
print(linear_endpoint_config)
create_endpoint_config_response = sm.create_endpoint_config(
EndpointConfigName=linear_endpoint_config,
ProductionVariants=[{
'InstanceType': 'ml.m4.xlarge',
'InitialInstanceCount': 1,
'ModelName': linear_job,
'VariantName': 'AllTraffic'}])
print("Endpoint Config Arn: " + create_endpoint_config_response['EndpointConfigArn'])
```
Now that we've specified how our endpoint should be configured, we can create them. This can be done in the background, but for now let's run a loop that updates us on the status of the endpoints so that we know when they are ready for use.
```
%%time
linear_endpoint = 'linear-endpoint-' + time.strftime("%Y%m%d%H%M", time.gmtime())
print(linear_endpoint)
create_endpoint_response = sm.create_endpoint(
EndpointName=linear_endpoint,
EndpointConfigName=linear_endpoint_config)
print(create_endpoint_response['EndpointArn'])
resp = sm.describe_endpoint(EndpointName=linear_endpoint)
status = resp['EndpointStatus']
print("Status: " + status)
sm.get_waiter('endpoint_in_service').wait(EndpointName=linear_endpoint)
resp = sm.describe_endpoint(EndpointName=linear_endpoint)
status = resp['EndpointStatus']
print("Arn: " + resp['EndpointArn'])
print("Status: " + status)
if status != 'InService':
raise Exception('Endpoint creation did not succeed')
```
## Predict
### Predict on Test Data
Now that we have our hosted endpoint, we can generate statistical predictions from it. Let's predict on our test dataset to understand how accurate our model is.
There are many metrics to measure classification accuracy. Common examples include include:
- Precision
- Recall
- F1 measure
- Area under the ROC curve - AUC
- Total Classification Accuracy
- Mean Absolute Error
For our example, we'll keep things simple and use total classification accuracy as our metric of choice. We will also evaluate Mean Absolute Error (MAE) as the linear-learner has been optimized using this metric, not necessarily because it is a relevant metric from an application point of view. We'll compare the performance of the linear-learner against a naive benchmark prediction which uses majority class observed in the training data set for prediction on the test data.
### Function to convert an array to a csv
```
def np2csv(arr):
csv = io.BytesIO()
np.savetxt(csv, arr, delimiter=',', fmt='%g')
return csv.getvalue().decode().rstrip()
```
Next, we'll invoke the endpoint to get predictions.
```
runtime= boto3.client('runtime.sagemaker')
payload = np2csv(test_X)
response = runtime.invoke_endpoint(EndpointName=linear_endpoint,
ContentType='text/csv',
Body=payload)
result = json.loads(response['Body'].read().decode())
test_pred = np.array([r['score'] for r in result['predictions']])
```
Let's compare linear learner based mean absolute prediction errors from a baseline prediction which uses majority class to predict every instance.
```
test_mae_linear = np.mean(np.abs(test_y - test_pred))
test_mae_baseline = np.mean(np.abs(test_y - np.median(train_y))) ## training median as baseline predictor
print("Test MAE Baseline :", round(test_mae_baseline, 3))
print("Test MAE Linear:", round(test_mae_linear,3))
```
Let's compare predictive accuracy using a classification threshold of 0.5 for the predicted and compare against the majority class prediction from training data set
```
test_pred_class = (test_pred > 0.5)+0;
test_pred_baseline = np.repeat(np.median(train_y), len(test_y))
prediction_accuracy = np.mean((test_y == test_pred_class))*100
baseline_accuracy = np.mean((test_y == test_pred_baseline))*100
print("Prediction Accuracy:", round(prediction_accuracy,1), "%")
print("Baseline Accuracy:", round(baseline_accuracy,1), "%")
```
###### Run the cell below to delete endpoint once you are done.
```
sm.delete_endpoint(EndpointName=linear_endpoint)
```
---
## Extensions
- Our linear model does a good job of predicting breast cancer and has an overall accuracy of close to 92%. We can re-run the model with different values of the hyper-parameters, loss functions etc and see if we get improved prediction. Re-running the model with further tweaks to these hyperparameters may provide more accurate out-of-sample predictions.
- We also did not do much feature engineering. We can create additional features by considering cross-product/intreaction of multiple features, squaring or raising higher powers of the features to induce non-linear effects, etc. If we expand the features using non-linear terms and interactions, we can then tweak the regulaization parameter to optimize the expanded model and hence generate improved forecasts.
- As a further extension, we can use many of non-linear models available through SageMaker such as XGBoost, MXNet etc.
| github_jupyter |
<!DOCTYPE html>
<html>
<body>
<div align="center">
<h3>Prepared by Asif Bhat</h3>
<h1>Data Visualization With Plotly (Part - 1)</h1>
</div>
</body>
</html>
```
import numpy as np
import pandas as pd
import plotly.graph_objects as go
import plotly.offline as po
from plotly.offline import download_plotlyjs, init_notebook_mode, plot, iplot
import matplotlib.pyplot as plt
import dash
import plotly.express as px
import random
import plotly.figure_factory as ff
```
# Loading Datasets
```
pokemon = pd.read_csv("C:/Users/DELL/Documents/GitHub/Public/Data-Visualization/Plotly/Datasets/pokemon_updated.csv")
pokemon.head(10)
stdperf = pd.read_csv("C:/Users/DELL/Documents/GitHub/Public/Data-Visualization/Plotly/Datasets/studentp.csv")
stdperf.head(10)
corona = pd.read_csv('C:/Users/DELL/Documents/GitHub/Public/COVID-19/covid/data/countries-aggregated.csv' ,
index_col='Date' , parse_dates=True)
corona.head(10)
spotify = pd.read_csv("C:/Users/DELL/Documents/GitHub/Public/Data-Visualization/Plotly/Datasets/spotify.csv" , index_col="Date")
spotify.head(10)
housing = pd.read_csv('C:/Users/DELL/Documents/GitHub/Data-Visualization/housing.csv')
housing.tail()
insurance = pd.read_csv('C:/Users/DELL/Documents/GitHub/Data-Visualization/insurance.csv')
insurance.head(10)
employment = pd.read_excel("C:/Users/DELL/Documents/GitHub/Public/Data-Visualization/Plotly/Datasets/unemployment.xlsx")
employment.head(10)
helpdesk = pd.read_csv("C:/Users/DELL/Documents/GitHub/Public/Data-Visualization/Plotly/Datasets/helpdesk.csv")
helpdesk.head(10)
fish= pd.read_csv("C:/Users/DELL/Documents/GitHub/Public/Data-Visualization/Plotly/Datasets/Fish.csv")
fish.head(10)
exercise = pd.read_csv("C:/Users/DELL/Documents/GitHub/Public/Data-Visualization/Plotly/Datasets/exercise.csv")
exercise.head(10)
suicide = pd.read_csv("C:/Users/DELL/Documents/GitHub/Public/Data-Visualization/Plotly/Datasets/suicide.csv")
suicide.head(10)
canada = pd.read_csv("C:/Users/DELL/Documents/GitHub/Public/Data-Visualization/Plotly/Datasets/canada.csv")
canada.head()
canada.columns
canada.drop(columns=['AREA' , 'DEV', 'DevName' , 'REG', 'Type', 'Coverage' , 'AreaName', 'RegName' ], inplace=True)
canada.head()
canada.rename(columns={'OdName':'Country'} , inplace=True)
canada.set_index(canada.Country,inplace=True)
canada.head()
canada2 = canada.copy()
canada2.head()
canada.index.name=None
canada.head()
del canada['Country']
canada.head()
canada = canada.transpose()
canada.head()
```
# Sankey Diagram
```
#Simple Sankey Diagram
fig = go.Figure(
go.Sankey(
node = {
"label": ["India", "USA", "China", "Pakistan", "Bangladesh", "Mexico"],
},
link = {
"source": [0, 1, 2, 3, 4, 0, 2, 5],
"target": [1, 2, 3, 4, 5, 3, 5, 3],
"value": [300, 400, 200, 450, 700, 200,150, 200]
}
)
)
fig.show()
#Simple Sankey Diagram
fig = go.Figure(
go.Sankey(
node = dict(
thickness = 40, # Changing thickness of nodes
color = "lightgreen", # Changing color of the node
line = dict(color = "red", width = 0.5), # Changing line color
label = ["India", "USA", "China", "Pakistan", "Bangladesh", "Mexico"],
),
link = {
"source": [0, 1, 2, 3, 4, 0, 2, 5],
"target": [1, 2, 3, 4, 5, 3, 5, 3],
"value": [300, 400, 200, 450, 550, 200,150, 200]
}
)
)
fig.show()
#Simple Sankey Diagram
fig = go.Figure(
go.Sankey(
node = {
"label": ["Married: NO", "Married: Yes",
"Pet: No", "Pet: Yes",
"Happy: Yes", "Happy: No"],
"color" : px.colors.qualitative.Set3 # Node color
},
link = dict(
source = [0, 0, 1, 1, 2, 2, 3, 5],
target = [2, 3, 2, 3, 5, 4, 4, 3],
value = [200, 300, 400, 600, 150, 350,700],
color = px.colors.qualitative.Set2 # Color of links
)
)
)
fig.show()
```
# END
| github_jupyter |
```
from __future__ import print_function, absolute_import
import gc
import sys
from torch.autograd import Variable
import torch
import torch.nn as nn
import torch.optim as optim
import torch.optim.lr_scheduler
torch.backends.cudnn.benchmark=True
import time
import torchvision.transforms as transforms
import torchvision.models as models
import numpy as np
import random
import cv2
from PIL import Image
# test CUDA available
import os
os.environ["CUDA_VISIBLE_DEVICES"] = "0"
print('CUDA enable: ', torch.cuda.is_available())
# import dataset from ./lib/dataset.py
import lib.dataset as dataset
from models.__init__ import load_model
from lib.config import cfg
from lib.solver import Solver
# Hyperparameter
BASE_LR = cfg.TRAIN.DEFAULT_LEARNING_RATE
EPOCH_DECAY = 10 # number of epochs after which the Learning rate is decayed exponentially.
DECAY_WEIGHT = cfg.TRAIN.WEIGHT_DECAY
cfg.CONST.BATCH_SIZE = 16
class AverageMeter(object):
"""Computes and stores the average and current value"""
def __init__(self):
self.reset()
def reset(self):
self.val = 0
self.avg = 0
self.sum = 0
self.count = 0
def update(self, val, n=1):
self.val = val
self.sum += val * n
self.count += n
self.avg = self.sum / self.count
def save_checkpoint(state, is_best, filename='checkpoint.pth.tar'):
torch.save(state, filename + '_latest.pth.tar')
if is_best:
shutil.copyfile(filename + '_latest.pth.tar', filename + '_best.pth.tar')
def accuracy(output, target, topk=(1,)):
"""Computes the precision@k for the specified values of k"""
maxk = max(topk)
batch_size = target.size(0)
_, pred = output.topk(maxk, 1, True, True)
pred = pred.t()
correct = pred.eq(target.view(1, -1).expand_as(pred))
res = []
for k in topk:
correct_k = correct[:k].view(-1).float().sum(0)
res.append(correct_k.mul_(100.0 / batch_size))
return res
# training hyperparameters
batch_size = cfg.CONST.BATCH_SIZE
train_val_ratio = cfg.TRAIN.DATASET_PORTION[1]
ren_len = dataset.ren_dataset.__len__()
vox_len = dataset.vox_dataset.__len__()
print(ren_len,vox_len)
dict_ren1 = dataset.ren_dataset.class_to_idx
list_ren = [[]]*(vox_len+1)
for (path, idx) in dataset.ren_dataset.samples:
list_ren[idx] = list(set(list_ren[idx]))
list_ren[idx].append(path)
dict_vox = {idx:path for (path, idx) in dataset.vox_dataset.samples}
# print(list_ren[202][2])
# print(dataset.center_crop(Image.open(list_ren[202][2])))
# print(dict_vox[202])
# This function changes the learning rate over the training model.
def exp_lr_scheduler(optimizer, epoch, init_lr=BASE_LR, lr_decay_epoch=EPOCH_DECAY):
"""Decay learning rate by a factor of DECAY_WEIGHT every lr_decay_epoch epochs."""
lr = init_lr * (DECAY_WEIGHT**(epoch // lr_decay_epoch))
if epoch % lr_decay_epoch == 0:
print('LR is set to {}'.format(lr))
for param_group in optimizer.param_groups:
param_group['lr'] = lr
return optimizer
def testDataInput():
NetClass = load_model(cfg.CONST.NETWORK_CLASS)
# print('Network definition: \n')
net = NetClass()
# print(net)
# start an epoch
# slice training and validation index
rand_idx = np.random.permutation(np.arange(vox_len))
thr = int(train_val_ratio*len(rand_idx))
train_idx = rand_idx[:thr]
val_idx = rand_idx[thr:]
batch_size = 4
max_num_views = 5
dict_vox = {idx:path for (path, idx) in dataset.vox_dataset.samples}
for i in range(thr//batch_size):
# for each batch
num_views = random.randint(2,max_num_views)
idx = train_idx[i*batch_size: (i+1)*batch_size]
voxel_loader = dataset.get_vox_data_loaders(idx)
label_list = []
for it, (labels, model_ids) in enumerate(voxel_loader):
batch_image = []
for model_id in model_ids:
image_list = []
image_ids = np.random.choice(cfg.TRAIN.NUM_RENDERING, num_views)
# print(image_ids)
for n_view in range(num_views):
image_list.append(dataset.center_crop(Image.open(list_ren[(model_id).item()][image_ids[n_view]]))[:3])
image_1 = torch.stack(image_list , dim=0)
batch_image.append(image_1)
# print(image_1.shape)
batch_image = torch.stack(batch_image,dim=0)
batch_image = batch_image.transpose(1,0)
# batch_image = batch_image.transpose(4,2)
# batch_image = batch_image.transpose(4,3)
labels0 = (labels < 1)
batch_voxel = torch.stack((labels0.float(),labels.float()),dim=0)
batch_voxel = batch_voxel.transpose(1,0)
inputs=Variable(batch_image)
labels=Variable(batch_voxel)
inputs = inputs.to(device)
labels = labels.to(device)
print(inputs.shape)
print(inputs[0].shape)
print(labels.shape)
# test mode
if i ==3:
break
# test mode end
def run():
# Parameters
num_epochs = 10
output_period = 100
batch_size = cfg.CONST.BATCH_SIZE
# setup the device for running
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
print(device)
NetClass = load_model(cfg.CONST.NETWORK_CLASS)
model = NetClass().to(device)
criterion = nn.CrossEntropyLoss().to(device)
optimizer = optim.Adam(model.parameters(), lr=cfg.TRAIN.DEFAULT_LEARNING_RATE,weight_decay=cfg.TRAIN.WEIGHT_DECAY)
top1trset,top5trset = [],[]
top1set,top5set = [],[]
epoch = 1
while epoch <= num_epochs:
batch_time = AverageMeter()
data_time = AverageMeter()
losses = AverageMeter()
running_loss = 0.0
optimizer = exp_lr_scheduler(optimizer, epoch)
for param_group in optimizer.param_groups:
print('Current learning rate: ' + str(param_group['lr']))
model.train()
# start an epoch
# slice training and validation index
rand_idx = np.random.permutation(np.arange(vox_len))
thr = int(train_val_ratio*len(rand_idx))
train_idx = rand_idx[:thr]
val_idx = rand_idx[thr:]
max_num_views = 5
dict_vox = {idx:path for (path, idx) in dataset.vox_dataset.samples}
num_train_batches = thr//batch_size
for i in range(num_train_batches):
# for each batch
num_views = random.randint(2,max_num_views)
idx = train_idx[i*batch_size: (i+1)*batch_size]
# print(idx)
voxel_loader = dataset.get_vox_data_loaders(idx)
label_list = []
for it, (labels, model_ids) in enumerate(voxel_loader):
batch_image = []
for model_id in model_ids:
image_list = []
image_ids = np.random.choice(cfg.TRAIN.NUM_RENDERING, num_views)
# print(image_ids)
for n_view in range(num_views):
image_list.append(dataset.center_crop(Image.open(list_ren[(model_id).item()][image_ids[n_view]]))[:3])
image_1 = torch.stack(image_list , dim=0)
batch_image.append(image_1)
# print(image_1.shape)
batch_image = torch.stack(batch_image,dim=0)
batch_image = batch_image.transpose(1,0)
labels0 = (labels < 1)
batch_voxel = torch.stack((labels0.float(),labels.float()),dim=0)
batch_voxel = batch_voxel.transpose(1,0)
inputs=Variable(batch_image)
labels=Variable(labels)
inputs = inputs.to(device)
labels = labels.to(device).long()
outputs = model(inputs,test=True)
# print('outputs[0].shape: ',outputs[0].shape)
# print('labels.shape: ',labels.shape)
loss = criterion(outputs[0], labels)
# measure accuracy and record loss
# prec1 = accuracy(outputs[0].data, labels, topk=(1,))
losses.update(loss.data.item(), inputs.size(0))
# top1.update(prec1.item(), inputs.size(0))
optimizer.zero_grad()
loss.backward()
optimizer.step()
running_loss += loss.item()
if i % output_period == 0:
print('[%d:%.2f] loss: %.3f' % (
epoch, i*1.0/num_train_batches,
running_loss/output_period
))
running_loss = 0.0
gc.collect()
gc.collect()
# save after every epoch
torch.save(model.state_dict(), "models/model.%d" % epoch)
model.eval()
batch_size_val = batch_size
for i in range((len(rand_idx)-thr)//batch_size_val):
# for each batch
num_views = 1
idx = val_idx[i*batch_size_val: (i+1)*batch_size_val]
voxel_loader = dataset.get_vox_data_loaders(idx)
label_list = []
for it, (labels, model_ids) in enumerate(voxel_loader):
batch_image = []
for model_id in model_ids:
image_list = []
image_ids = np.random.choice(cfg.TRAIN.NUM_RENDERING, num_views)
for n_view in range(num_views):
image_list.append(dataset.center_crop(Image.open(list_ren[(model_id).item()][image_ids[n_view]]))[:3])
image_1 = torch.stack(image_list , dim=0)
batch_image.append(image_1)
batch_image = torch.stack(batch_image,dim=0)
batch_image = batch_image.transpose(1,0)
labels0 = (labels < 1)
batch_voxel = torch.stack((labels0.float(),labels.float()),dim=0)
batch_voxel = batch_voxel.transpose(1,0)
inputs=Variable(batch_image)
labels=Variable(labels)
inputs = inputs.to(device)
labels = labels.to(device).long()
outputs = model(inputs)
loss = criterion(outputs[0], labels)
# measure accuracy and record loss
# prec1 = accuracy(outputs[0].data, labels, topk=(1,))
losses.update(loss.data.item(), inputs.size(0))
# top1.update(prec1.item(), inputs.size(0))
print('test loss = '+ losses.avg)
gc.collect()
epoch += 1
print('Starting training')
BASE_LR = cfg.TRAIN.DEFAULT_LEARNING_RATE
run()
print('Training terminated')
```
| github_jupyter |
# Desenvolvendo um Modelo
<p> Nesta seção, iremos desenvolver vários modelos que irão prever o preço do carro usando as suas características. Esta é apenas uma estimativa, mas deve nos dar uma ideia objetiva de quanto o carro deve custar. </p>
Algumas perguntas que podemos fazer:
<ul>
<li> eu sei se o revendedor está oferecendo um valor justo pela minha troca? </li>
<li> eu sei se valorizo meu carro? </li>
</ul>
<p> Geralmente usamos um <b> modelo </b> para nos ajudar a prever observações futuras a partir dos dados que temos. </p>
<p> Um modelo nos ajudará a entender a relação exata entre as diferentes variáveis e como essas variáveis são usadas para prever o resultado. </p>
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
df = pd.read_csv('clean_auto_df.csv')
df.head()
```
<h3>Regressão Linear e Regressão Linear Múltipla </h3>
<h4> Regressão Linear </h4>
<p> Um exemplo de modelo que usaremos é a</p>
<b> Regressão Linear Simples </b>.
<br>
<p> A regressão linear simples é um método para nos ajudar a entender a relação entre duas variáveis: </p>
<ul>
<li> O preditor / variável independente (X) </li>
<li> A resposta / variável dependente (que queremos prever) (Y) </li>
</ul>
<p> O resultado da regressão linear é uma <b>função linear </b> que prevê a variável de resposta (dependente) como uma função da variável preditora (independente). </p>
$$
Y: Response \ Variable\\
X: Predictor \ Variables
$$
<b>Linear function:</b>
$$
Yhat = a + b X
$$
<ul>
<li> a refere-se à <b> interceptação </b> da linha de regressão 0, em outras palavras: o valor de Y quando X é 0 </li>
<li> b refere-se à <b> inclinação </b> da linha de regressão, em outras palavras: o valor com o qual Y muda quando X aumenta em 1 unidade </li>
</ul>
```
# Carregando os módulos da regressão linear
from sklearn.linear_model import LinearRegression
# Criando um objeto da classe de regressão linear
lm = LinearRegression()
lm
```
<h4>A variável city-mpg pode nos ajudar a prever o preço do carro?</h4
Para este exemplo, queremos ver como a city-mpg pode nos ajudar a prever o preço do carro.
Usando regressão linear simples, criaremos uma função linear com "city-mpg" como variável preditora e o "price" como variável de resposta.
```
X = df[['city-mpg']]
Y = df['price']
lm.fit(X,Y)
# vai utilizar o intercepto (coeficiente angular)
#
Yhat=lm.predict(X)
Yhat[0:5]
# o valor de interceptação
lm.intercept_
# o valor de inclinação
lm.coef_
```
Com esses valores, se utilizarmos a função:
$$
Yhat = a + b X
$$
Teremos:
price = 38423.31 - 821.73 * city-mpg
<h4>Testando com outra variável:</h4>
```
lm1 = LinearRegression()
lm1.fit(df[['engine-size']], df[['price']])
# Inclinação
lm1.coef_
# Interceptação
lm1.intercept_
```
##### Utilizando a função:
Yhat= a + b*X
##### temos:
price=-7963.34 + 166.86*engine-size
<h4> Regressão Linear Múltipla</h4>
<p> E se quisermos prever o preço do carro usando mais de uma variável? </p>
<p> Se quisermos usar mais variáveis em nosso modelo para prever o preço do carro, podemos usar <b> Regressão linear múltipla </b>.
A Regressão Linear Múltipla é muito semelhante à Regressão Linear Simples, mas este método é usado para explicar a relação entre uma variável de resposta contínua (dependente) e <b> duas ou mais </b> variáveis preditoras (independentes).
A maioria dos modelos de regressão do mundo real envolve vários preditores. Ilustraremos a estrutura usando quatro variáveis preditoras, mas esses resultados podem ser generalizados para qualquer número inteiro: </p>
$$
Y: Response \ Variable\\
X_1 :Predictor\ Variable \ 1\\
X_2: Predictor\ Variable \ 2\\
X_3: Predictor\ Variable \ 3\\
X_4: Predictor\ Variable \ 4\\
$$
$$
a: intercept\\
b_1 :coefficients \ of\ Variable \ 1\\
b_2: coefficients \ of\ Variable \ 2\\
b_3: coefficients \ of\ Variable \ 3\\
b_4: coefficients \ of\ Variable \ 4\\
$$
A equação é dada por:
$$
Yhat = a + b_1 X_1 + b_2 X_2 + b_3 X_3 + b_4 X_4
$$
<p>Analisamos em outros notebooks que algumas variáveis podem ser boas para prever o preço do carro:</p>
<ul>
<li>Horsepower</li>
<li>Curb-weight</li>
<li>Engine-size</li>
<li>city-mpg</li>
</ul>
Vamos criar um modelo com essas variaveis como nossas variaveis preditoras
```
Z = df[['horsepower', 'curb-weight', 'engine-size', 'city-mpg']]
lm.fit(Z, df['price'])
# valor do intercepto
lm.intercept_
# valor do coeficiente
lm.coef_
```
Se substituirmos a equação pelos valores que obtivemos, temos:
$$
Yhat = a + b_1 X_1 + b_2 X_2 + b_3 X_3 + b_4 X_4
$$
<b>Price</b> = -15678.742628061467 + 52.65851272 x <b>horsepower</b> + 4.69878948 x <b>curb-weight</b> + 81.95906216 x <b>engine-size</b> + 33.58258185 x <b>city-mpg</b>
<h3>Avaliação de modelo usando visualização </h3>
```
import seaborn as sns
%matplotlib inline
```
<h3>Regression Plot</h3>
<p> Quando se trata de regressão linear simples, uma excelente maneira de visualizar o ajuste do nosso modelo é usando <b> gráficos de regressão </b>. </p>
<p> Este gráfico mostrará uma combinação de pontos de dados dispersos (um <b> gráfico de dispersão </b>), bem como a linha de <b> regressão linear </b> ajustada passando pelos dados. Isso nos dará uma estimativa razoável da relação entre as duas variáveis, a força da correlação, bem como a direção (correlação positiva ou negativa). </p>
```
width = 12
height = 10
plt.figure(figsize=(width, height))
sns.regplot(x="city-mpg", y="price", data=df)
plt.ylim(0,)
```
<p> Podemos ver neste gráfico que o preço está negativamente correlacionado com a variavel city-mpg, uma vez que a inclinação da regressão é negativa.
Uma coisa a ter em mente ao examinar um gráfico de regressão é prestar atenção em como os pontos de dados estão espalhados ao redor da linha de regressão. Isso lhe dará uma boa indicação da variação dos dados e se um modelo linear seria o melhor ajuste ou não. Se os dados estiverem muito distantes da linha, este modelo linear pode não ser o melhor modelo para esses dados. Vamos comparar este gráfico com o gráfico de regressão de "peak-rpm". </p>
```
plt.figure(figsize=(width, height))
sns.regplot(x="peak-rpm", y="price", data=df)
plt.ylim(0,)
```
<p> Comparando o gráfico de regressão de "peak-rpm" e "city-mpg", vemos que os pontos de "city-mpg" estão muito mais próximos da linha gerada e na diminuição média. Os pontos para "peak-rpm" estão mais espalhados ao redor da linha prevista e é muito mais difícil determinar se os pontos estão diminuindo ou aumentando à medida que a "city-mpg" aumenta. </p>
<h3> Gráfico residual </h3>
<p> Uma boa maneira de visualizar a variação dos dados é usar um gráfico residual. </p>
<p> O que é um <b> residual </b>? </p>
<p> A diferença entre o valor observado (y) e o valor previsto (Yhat) é chamada de residual (e). Quando olhamos para um gráfico de regressão, o residual é a distância do ponto de dados até a linha de regressão ajustada. </p>
<p> Então, o que é um <b> gráfico residual </b>? </p>
<p> Um gráfico residual é um gráfico que mostra os resíduos no eixo vertical y e a variável independente no eixo horizontal x. </p>
<p> Em que prestamos atenção quando olhamos para um lote residual? </p>
<p> Nós olhamos a propagação dos resíduos: </p>
<p> - Se os pontos em um gráfico residual estiverem <b> espalhados aleatoriamente em torno do eixo x </b>, então um <b> modelo linear é apropriado </b> para os dados. Por que é que? Resíduos espalhados aleatoriamente significam que a variância é constante e, portanto, o modelo linear é um bom ajuste para esses dados. </p>
```
width = 12
height = 10
plt.figure(figsize=(width, height))
sns.residplot(df['city-mpg'], df['price'])
plt.show()
```
<i> O que este gráfico está nos dizendo? </i>
<p> Podemos ver neste gráfico residual que os resíduos não estão espalhados aleatoriamente em torno do eixo x, o que nos leva a acreditar que talvez um modelo não linear seja mais apropriado para esses dados. </p>
<h3>Multiple Linear Regression</h3>
<p> Como podemos visualizar um modelo de regressão linear múltipla? Isso fica um pouco mais complicado porque você não pode visualizá-lo com regressão ou gráfico residual. </p>
<p> Uma maneira de ver o ajuste do modelo é olhando para o <b> gráfico de distribuição </b>: podemos olhar para a distribuição dos valores ajustados que resultam do modelo e compará-la com a distribuição de os valores reais. </p>
```
Y_hat = lm.predict(Z)
plt.figure(figsize=(width, height))
ax1 = sns.distplot(df['price'], hist=False, color="r", label="Actual Value")
sns.distplot(Y_hat, hist=False, color="b", label="Fitted Values" , ax=ax1)
plt.title('Actual vs Fitted Values for Price')
plt.xlabel('Price (in dollars)')
plt.ylabel('Proportion of Cars')
plt.show()
plt.close()
```
<p> Podemos ver que os valores ajustados estão razoavelmente próximos dos valores reais, uma vez que as duas distribuições se sobrepõem um pouco. No entanto, definitivamente há espaço para melhorias. </p>
<h2>Regressão polinomial e pipelines </h2>
<p> <b> Regressão polinomial </b> é um caso particular do modelo de regressão linear geral ou modelos de regressão linear múltipla. </p>
<p> Obtemos relacionamentos não lineares elevando ao quadrado ou definindo termos de ordem superior das variáveis preditoras. </p>
<p> Existem diferentes ordens de regressão polinomial: </p>
<center><b>Quadratic - 2nd order</b></center>
$$
Yhat = a + b_1 X +b_2 X^2
$$
<center><b>Cubic - 3rd order</b></center>
$$
Yhat = a + b_1 X +b_2 X^2 +b_3 X^3\\\\
$$
<center><b>Higher order</b>:</center>
$$
Y = a + b_1 X +b_2 X^2 +b_3 X^3 ....\\\\
$$
<p> Vimos anteriormente que um modelo linear não fornece o melhor ajuste ao usar city-mpg como variável preditora. Vamos ver se podemos tentar ajustar um modelo polinomial aos dados. </p>
```
# plotando a função
def PlotPolly(model, independent_variable, dependent_variabble, Name):
x_new = np.linspace(15, 55, 100)
y_new = model(x_new)
plt.plot(independent_variable, dependent_variabble, '.', x_new, y_new, '-')
plt.title('Polynomial Fit - Price ~ Length')
ax = plt.gca()
ax.set_facecolor((0.898, 0.898, 0.898))
fig = plt.gcf()
plt.xlabel(Name)
plt.ylabel('Preço dos carros')
plt.show()
plt.close()
x = df['city-mpg']
y = df['price']
# utilizando uma regressão polinomial de 3a ordem (cubic)
f = np.polyfit(x, y, 3)
p = np.poly1d(f)
print(p)
PlotPolly(p, x, y, 'city-mpg')
np.polyfit(x, y, 3)
```
<p> Já podemos ver na plotagem que este modelo polinomial tem um desempenho melhor do que o modelo linear. Isso ocorre porque a função polinomial gerada "atinge" mais os pontos de dados. </p>
```
# repetindo uma regressão polinomial agora na 11a ordem (cubic)
f = np.polyfit(x, y, 11)
p = np.poly1d(f)
print(p)
PlotPolly(p,x,y, 'Highway MPG')
```
<p> A expressão analítica para função polinomial multivariada é complicada. Por exemplo, a expressão para um polinômio de segunda ordem (grau = 2) com duas variáveis é dada por: </p>
$$
Yhat = a + b_1 X_1 +b_2 X_2 +b_3 X_1 X_2+b_4 X_1^2+b_5 X_2^2
$$
Podemos realizar uma transformação polinomial em vários recursos. Primeiro, importamos o módulo:
```
from sklearn.preprocessing import PolynomialFeatures
#criando um objeto com grau 2
pr=PolynomialFeatures(degree=2)
pr
```
<h2>Pipeline</h2>
<p> Os pipelines de dados simplificam as etapas de processamento dos dados. Usamos o módulo <b> Pipeline </b> para criar um pipeline. Também usamos <b> StandardScaler </b> como uma etapa em nosso pipeline. </p>
```
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
```
Criamos o pipeline, criando uma lista de tuplas incluindo o nome do modelo ou estimador e seu construtor correspondente.
```
# inserimos a lista como um argumento para o construtor do pipeline
Input=[('scale',StandardScaler()), ('polynomial', PolynomialFeatures(include_bias=False)), ('model',LinearRegression())]
pipe=Pipeline(Input)
pipe
# Podemos normalizar os dados, realizar uma transformação e ajustar o modelo simultaneamente.
pipe.fit(Z,y)
# Da mesma forma, podemos normalizar os dados, realizar uma transformação e produzir uma previsão simultaneamente
ypipe=pipe.predict(Z)
ypipe[0:4]
```
<h2>Medidas para avaliação</h2>
<p> Ao avaliar nossos modelos, não queremos apenas visualizar os resultados, mas também uma medida quantitativa para determinar a precisão do modelo. </p>
<p> Duas medidas muito importantes que costumam ser usadas em estatísticas para determinar a precisão de um modelo são: </p>
<ul>
<li> <b> R ^ 2 / R-squared (R ao quadrado)</b> </li>
<li> <b> Mean Squared Error (Erro quadrático médio) (MSE) </b> </li>
</ul>
<b> R-squared </b>
<p> R-squared (R ao quadrado), também conhecido como coeficiente de determinação, é uma medida para indicar a proximidade dos dados com a linha de regressão ajustada. </p>
<p> O valor do R-squared é a porcentagem de variação da variável de resposta (y) que é explicada por um modelo linear. </p>
<b> Erro quadrático médio (MSE) </b>
<p> O erro quadrático médio mede a média dos quadrados dos erros, ou seja, a diferença entre o valor real (y) e o valor estimado (ŷ). </p>
<h3>Model 1: Simple Linear Regression</h3>
```
# Vamos calcular o R^2
lm.fit(X, Y)
# R^2
print('The R-square is: ', lm.score(X, Y))
```
Podemos dizer que ~ 47,13% da variação do preço é explicada por este modelo linear simples.
```
# Vamos calcular o MSE
# Podemos prever a saída, ou seja, "yhat" usando o método de previsão, onde X é a variável de entrada:
Yhat=lm.predict(X)
print('A saída dos primeiros quatro valores previstos é: ', Yhat[0:4])
```
vamos importar a função <b> mean_squared_error </b> do módulo <b> metrics </b>
```
from sklearn.metrics import mean_squared_error
# comparamos os resultados previstos com os resultados reais
mse = mean_squared_error(df['price'], Yhat)
print('O erro quadrático médio do preço e valor previsto é: ', mse)
```
<h3>Model 2: Multiple Linear Regression</h3>
```
# fit the model
lm.fit(Z, df['price'])
# R^2
print('R-square: ', lm.score(Z, df['price']))
```
Podemos dizer que ~ 80,896% da variação de preço é explicada por esta regressão linear múltipla "multi_fit".
```
# MSE
Y_predict_multifit = lm.predict(Z)
print('O erro quadrático médio de preço e valor previsto usando multifit é', \
mean_squared_error(df['price'], Y_predict_multifit))
```
<h3>Model 3: Polynomial Fit</h3>
```
from sklearn.metrics import r2_score
r_squared = r2_score(y, p(x))
print('R-square: ', r_squared)
mean_squared_error(df['price'], p(x))
```
<h2> Previsão e tomada de decisão </h2>
<h3> Previsão </h3>
<p> Treinamos o modelo usando o método <b> fit </b>. Agora vamos usar o método <b> predict </b> para produzir uma previsão. Vamos importar <b> pyplot </b> para plotagem; também usaremos algumas funções do numpy. </p>
```
import matplotlib.pyplot as plt
import numpy as np
%matplotlib inline
# novo input
new_input=np.arange(1, 100, 1).reshape(-1, 1)
# fit
lm.fit(X, Y)
lm
# criando uma previsão
yhat=lm.predict(new_input)
yhat[0:5]
# plotando o resultado
plt.plot(new_input, yhat)
plt.show()
```
<h3> Tomada de decisão: Determinando um bom ajuste do modelo </h3>
<p> Agora que visualizamos os diferentes modelos e geramos os valores R-quadrado e MSE para os ajustes, como determinamos um bom ajuste do modelo?
<ul>
<li> <i> O que é um bom valor de R ao quadrado? </i> </li>
</ul>
</p>
<p> Ao comparar modelos, <b> o modelo com o valor de R quadrado mais alto é um ajuste melhor </b> para os dados.
<ul>
<li> <i> O que é um bom MSE? </i> </li>
</ul>
</p>
<p> Ao comparar modelos, <b> o modelo com o menor valor de MSE é um ajuste melhor </b> para os dados. </p>
<h4> Vamos dar uma olhada nos valores dos diferentes modelos. </h4>
<p> Regressão linear simples: usando city-mpg como variável preditora de preço.
<ul>
<li> R ao quadrado: 0,49659118843391759 </li>
<li> MSE: 3,16 x10 ^ 7 </li>
</ul>
</p>
<p> Regressão linear múltipla: usando horsepower, engine-size, tamanho do motor e city-mpg como variáveis preditoras de preço.
<ul>
<li> R ao quadrado: 0,80896354913783497 </li>
<li> MSE: 1,2 x10 ^ 7 </li>
</ul>
</p>
<p> Ajuste polinomial: usando city-mpg como uma variável preditora de preço.
<ul>
<li> R ao quadrado: 0,6741946663906514 </li>
<li> MSE: 2,05 x 10 ^ 7 </li>
</ul>
</p>
<h3> Modelo de regressão linear simples (SLR) vs modelo de regressão linear múltipla (MLR) </h3>
<p> Normalmente, quanto mais variáveis você tiver, melhor será a previsão do seu modelo, mas nem sempre isso é verdade. Às vezes, você pode não ter dados suficientes, pode ter problemas numéricos ou muitas das variáveis podem não ser úteis e ou mesmo atuar como ruído. Como resultado, você deve sempre verificar o MSE e R ^ 2. </p>
<p> Assim, para poder comparar os resultados dos modelos MLR vs SLR, olhamos para uma combinação de R-quadrado e MSE para tirar a melhor conclusão sobre o ajuste do modelo.
<ul>
<li> <b> MSE </b> O MSE de SLR é 3,16x10 ^ 7, enquanto MLR tem um MSE de 1,2 x10 ^ 7. O MSE do MLR é muito menor. </li>
<li> <b> R ao quadrado </b>: neste caso, também podemos ver que há uma grande diferença entre o R ao quadrado do SLR e o R ao quadrado do MLR. O R-quadrado do SLR (~ 0,497) é muito pequeno em comparação com o R-quadrado do MLR (~ 0,809). </li>
</ul>
</p>
Este R-quadrado em combinação com o MSE mostra que o MLR parece ser o modelo de melhor ajuste neste caso, em comparação com o SLR.
<h3> Modelo Linear Simples (SLR) vs Ajuste Polinomial </h3>
<ul>
<li> <b> MSE </b>: podemos ver que o ajuste polinomial derrubou o MSE, já que este MSE é menor que o do SLR</li>
<li> <b> R ao quadrado </b>: o R ao quadrado do Polyfit é maior do que o R ao quadrado do SLR, então o ajuste polinomial também aumentou o R ao quadrado.</li >
</ul>
<p> Como o ajuste polinomial resultou em um MSE inferior e um R-quadrado mais alto, podemos concluir que este foi um modelo de ajuste melhor do que a regressão linear simples para prever o preço com city-mpg como variável preditora. </p>
<h3> Regressão Linear Múltipla (MLR) vs Ajuste Polinomial </h3>
<ul>
<li> <b> MSE </b>: O MSE para o MLR é menor do que o MSE para o ajuste polinomial. </li>
<li> <b> R ao quadrado </b>: o R ao quadrado do MLR também é muito maior do que para o ajuste polinomial. </li>
</ul>
<h2>Conclusão:</h2>
<p> Comparando esses três modelos, concluímos que <b> o modelo MLR é o melhor modelo </b> para ser capaz de prever o preço de nosso conjunto de dados. Esse resultado faz sentido, pois temos 27 variáveis no total e sabemos que mais de uma dessas variáveis são preditores potenciais do preço final do carro. </p>
# É isso!
### Este é apenas um exemplo de aplicação de um modelo, com pandas.
Este notebook faz parte de uma série de notebooks com conteúdos extraídos de cursos dos quais participei como aluno, ouvinte, professor, monitor... Reunidos para consulta futura e compartilhamento de idéias, soluções e conhecimento!
### Muito obrigado pela sua leitura!
<h4>Anderson Cordeiro</h4>
Você pode encontrar mais conteúdo no meu Medium<br> ou então entrar em contato comigo :D
<a href="https://www.linkedin.com/in/andercordeiro/" target="_blank">[LinkedIn]</a>
<a href="https://medium.com/@andcordeiro" target="_blank">[Medium]</a>
| github_jupyter |

# Array
In this notebook, we will learn about arrays as a data structure using python.
### Definition
An array is a collection of items stored at contiguous memory locations. The idea is to store multiple items of the same datatype together. This makes it easier to calculate the position of each element by simply adding an offset to a base value, i.e., the memory location of the first element of the array (generally denoted by the name of the array).
> Arrays are defined as the collection of similar type of data items stored at contiguous memory locations.
> Arrays have a fixed datatype, i.e., an integer array can only store integer values, unlike "list" in python which can store all datatypes.
> Arrays are the derived data type in C programming language which can store the primitive type of data such as int, char, double, float, etc.
> Arrays technically have fixed size i.e, if an array has capacity of storing 10 elements, it cannot store above 10 elements. But in python, arrays are flexible, i.e size of arrays can be increased or decreased, unlike Java or C++.
> Arrays are the derived data type in C programming language which can store the primitive type of data such as int, char, double, float, etc.
For example, if we want to store the marks of a student in 6 subjects, then we don't need to define different variable for the marks in different subject. instead of that, we can define an array which can store the marks in each subject at a the contiguous memory locations.
The above image can be looked as a top-level view of a staircase where you are at the base of the staircase. Each element can be uniquely identified by their index in the array (in a similar way as you could identify your friends by the step on which they were on in the above example).
### Implementing Array In Python
Arrays in python are stored in the array module, which we will import using the import statement.
```
from array import *
```
The above line will import array and its functions.
Further, you will have to initialize array as follows:
```
arr = array('i',[1,2,3,4,5])
print(arr)
#Try it out yourself in this block!
```
Now you must be wondering what does 'i' stand for?
Well the 'i' in this case denotes the datatype of array which is signed integer.
There are various types of datatypes and the following table mentions datatypes and their typecodes.

Go through the table above, and try to create and print array using various data types.
```
#Explore here!
```
# Operations on Array
### Iterating through an array
```
from array import *
arr = array('i',[1,2,3,4,5])
for x in arr:
print(x)
```
Task: Multiply every element of array with 2
```
#Perform task here:
```
### Inserting in an array
```
arr = array('i',[1,2,3])
arr.append(4) #this inserts 4 in array
print(arr)
```
### Deleting from an array
```
arr = array('i',[1,2,3,4,5])
arr.pop() #this deletes last element of the array
print(arr)
```
### Taking user input in an array
```
arr = array('i',[])
for x in range(5):
a = int(input("Enter value here: "))
arr.append(a)
print(arr)
```
Task: Take array as input from the user and print square of each element
```
#Perform task here
```
| github_jupyter |
# Implementation of [Complex Networks Classification with Convolutional Neural Network](https://arxiv.org/pdf/1802.00539.pdf)
```
# disable GPU support for TF
import os
os.environ['CUDA_VISIBLE_DEVICES'] = '-1'
import networkx as nx
from karateclub import DeepWalk
import warnings
import matplotlib.pyplot as plt
from matplotlib import rc
from sklearn.decomposition import PCA
import numpy as np
import io
import cv2
from tqdm import tqdm
import pickle
import datetime
import tensorflow as tf
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Conv2D, Flatten, Dropout, MaxPooling2D
rc('font', **{'family': 'sans-serif', 'sans-serif': ['DejaVu Sans'], 'size': 13})
warnings.filterwarnings("ignore")
```
### Converting graph into image
1. Create graph embedding (using e.g. DeepWalk)
2. Use PCA to reduce the dimension of node representation into 2-d space
3. Rasterize the 2-d scatter plot into 48x48 grid. **In my implementation: convert matplotlib.plot into image and then create numpy array as input to the CNNs**
4. Count the number of nodes in each grid as the pixel grayscale
5. Input to CNN :)

```
def draw_network(g: nx.Graph, scale_size=True, node_size=40, edge_alpha=0.3):
plt.figure(figsize=(8, 8))
pos = nx.kamada_kawai_layout(g)
if scale_size:
d = [node_size * size for size in dict(g.degree).values()]
else:
d = [node_size] * len(g)
nx.draw_networkx(g, pos=pos, with_labels=False, node_size=1, alpha=0)
nx.draw_networkx_edges(g, pos=pos, alpha=edge_alpha)
nx.draw_networkx_nodes(g, pos=pos, node_size=d, node_color='darkorange', edgecolors='black')
plt.axis('off')
ba = nx.barabasi_albert_graph(200, m=3)
draw_network(ba, node_size=10, edge_alpha=0.1)
# plt.savefig('ba_network.png', dpi=300, bbox_layout=True)
def deep_walk(g, walk_number=10, walk_length=80, dimensions=128):
model = DeepWalk(walk_number=walk_number, walk_length=walk_length, dimensions=dimensions)
model.fit(g)
return model.get_embedding()
def get_pca_components(embeddings, n_components=2):
pca = PCA(n_components=n_components)
pca.fit(embeddings)
return pca.components_
def draw_embeddings(pca_components):
plt.scatter(pca_components[0, :], pca_components[1, :])
plt.title("2D embeddings of network")
def get_img_from_fig(fig, dpi=200, new_size=(48, 48)):
buf = io.BytesIO()
fig.savefig(buf, format="png", dpi=dpi)
buf.seek(0)
img_arr = np.frombuffer(buf.getvalue(), dtype=np.uint8)
buf.close()
img = cv2.imdecode(img_arr, 1)
img = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
img = cv2.resize(img, new_size)
return img
def rgb2gray(rgb):
return np.dot(rgb[...,:3], [0.2989, 0.5870, 0.1140])
def get_image_from_embeddings(pca_components, input_size=(48, 48)):
"""
Based on: https://stackoverflow.com/a/58641662/9511702
"""
fig = plt.figure(figsize=(6, 6))
fig.tight_layout(pad=0)
plt.scatter(pca_components[0, :], pca_components[1, :], alpha=0.3)
plt.axis('off')
plt.xticks([], [])
image = get_img_from_fig(fig, new_size=input_size)
image_greyscale = rgb2gray(image)
image_greyscale = np.reshape(image_greyscale, newshape=(image_greyscale.shape[0], image_greyscale.shape[1], 1))
plt.close()
return image_greyscale
def plot_network_image(image):
plt.figure(figsize=(8, 8))
plt.imshow(image[:, :, 0], cmap=plt.get_cmap('gray'))
plt.axis('off')
a = get_pca_components(deep_walk(ba).T)
a.shape
res = get_image_from_embeddings(a, input_size=(28, 28))
plot_network_image(1 - res)
# plt.savefig('ba_network_after.png', dpi=300, bbox_layout=True)
```
| **Intput BA network** | **After pipeline** |
|:-: |:-: |
| |  |
```
def create_synthetic_input(size, k, network_type, input_size=(100, 100)):
if network_type == 'BA':
network = nx.barabasi_albert_graph(size, m=(int)(k / 2))
elif network_type == 'WS':
network = nx.watts_strogatz_graph(size, k, p=0.1)
else:
print('Wrong network type. Available `BA`, `WS`.')
embedding = get_pca_components(deep_walk(network).T)
return network_type, get_image_from_embeddings(embedding, input_size)
def save_networks(file, networks):
with open('data/' + file + '.pickle', 'wb') as f:
pickle.dump(networks, f)
def load_networks(file):
with open(file, 'rb') as f:
return pickle.load(f)
N = 1000
k = 6
LENGTH = 800
ba_networks = []
ws_networks = []
for n in tqdm(range(LENGTH)):
ba = create_synthetic_input(N, k, network_type='BA', input_size=(28, 28))
ws = create_synthetic_input(N, k, network_type='WS', input_size=(28, 28))
ba_networks.append(ba)
ws_networks.append(ws)
if (n + 1) % 10 == 0:
save_networks(f'smaller_images_bigger_network/ws_networks_N={N}_k={k}_length={LENGTH}_n={n+1}_size=28x28', ws_networks)
save_networks(f'smaller_images_bigger_network/ba_networks_N={N}_k={k}_length={LENGTH}_n={n+1}_size=28x28', ba_networks)
ws_networks = []
ba_networks = []
```
### Split train/test data
```
import glob
def load_network_in_parts(path):
network = []
for net in glob.glob(path):
network += load_networks(net)
return network
ba_networks = load_network_in_parts('data/smaller_images_bigger_network/ba*')
ws_networks = load_network_in_parts('data/smaller_images_bigger_network/ws*')
len(ba_networks), len(ws_networks)
ba_label, ba_data = zip(*ba_networks)
ws_label, ws_data = zip(*ws_networks)
label = np.array(list(ba_label + ws_label))
network_data = np.array(list(ba_data + ws_data))
# Convert label to 0 (BA) - 1 (WS)
label = np.array([0 if l == 'BA' else 1 for l in list(label)])
network_data /= 255
network_data = 1 - network_data
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(network_data, label, test_size=0.1, random_state=32)
len(X_train), len(X_test), X_train.shape
```
### Create CNN
```
IMG_HEIGHT = 28 # 100
IMG_WIDTH = 28 # 100
model = Sequential([
Conv2D(64, 3, activation='relu', input_shape=(IMG_HEIGHT, IMG_WIDTH, 1)),
MaxPooling2D(),
Conv2D(32, 3, activation='relu'),
MaxPooling2D(),
Dropout(0.3),
Conv2D(16, 2, activation='relu'),
MaxPooling2D(),
Flatten(),
Dropout(0.3),
Dense(256, activation='relu'),
Dense(1, activation='sigmoid')
])
model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy', 'AUC'])
model.summary()
tf.keras.utils.plot_model(model, show_shapes=True)
%load_ext tensorboard
log_dir = "logs\\fit\\" + datetime.datetime.now().strftime("%Y%m%d-%H%M%S")
tensorboard_callback = tf.keras.callbacks.TensorBoard(log_dir=log_dir, histogram_freq=1)
!rm -rf logs/fit/*
early_stopping_callback = tf.keras.callbacks.EarlyStopping(monitor='val_loss', patience=3)
EPOCHS = 100
BATCH_SIZE = 128
history = model.fit(X_train,
y_train,
epochs=EPOCHS,
validation_data=(X_test, y_test),
batch_size=BATCH_SIZE,
verbose=2,
callbacks=[early_stopping_callback])
# callbacks=[tensorboard_callback])
%tensorboard --logdir logs/fit
results = model.evaluate(X_test, y_test, batch_size=128, verbose=2)
print("test loss, test acc, test AUC:", results, '\n')
print("Generate predictions for 3 samples")
predictions = model.predict(X_test[:10])
print("predictions shape:", predictions.shape)
(predictions.flatten() > 0.5).astype('int'), y_test[:10]
plt.figure(figsize=(15, 4))
plt.subplot(131)
plt.plot(history.history['loss'], label='train')
plt.plot(history.history['val_loss'], label='test')
plt.ylabel('Loss')
plt.xlabel('epoch')
plt.legend()
plt.subplot(132)
plt.plot(history.history['accuracy'], label='train')
plt.plot(history.history['val_accuracy'], label='test')
plt.ylabel('Accuracy')
plt.xlabel('epoch')
plt.legend()
plt.subplot(133)
plt.plot(history.history['AUC'], label='train')
plt.plot(history.history['val_AUC'], label='test')
plt.ylabel('AUC')
plt.xlabel('epoch')
plt.legend()
plt.tight_layout()
```
| github_jupyter |
<img align="left" style="padding-right:10px;" width="150" src="https://upload.wikimedia.org/wikipedia/commons/thumb/6/6c/Star_Wars_Logo.svg/320px-Star_Wars_Logo.svg.png" />
*elaborado por Ferran Carrascosa Mallafrè.*
__[Abre en Colab](https://colab.research.google.com/github/griu/init_python_b1/blob/master/Ejercicios_Python_I.ipynb)__
# Preparación del entorno
Padawan! Cuando inicies sesión en Colab, prepara el entorno ejecutando el siguiente código.
```
if 'google.colab' in str(get_ipython()):
!git clone https://github.com/griu/init_python_b1.git /content/init_python_b1
!git -C /content/init_python_b1 pull
%cd /content/init_python_b1
```
# Ejercicio 1
Para realizar el ejercicio cargamos los datos de *Especies* en STARWARS SWAPI y las librerías principales.
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns; sns.set() # para el estilo de graficos
entidades = ['planets','starships','vehicles','people','species']
entidades_df = {x: pd.read_pickle('www/' + x + '_df.pkl') for x in entidades}
# Datos principales
people_df = entidades_df['people'][["height","mass","birth_year","gender","homeworld"]].dropna()
people_df
```
## Ejercicio 1.1.
Construye un gráfico de dispersión de los personajes donde se visualice: la altura (height), el peso (mass), la edad en años BBY (birth_year) y el género (gender). Para ello utiliza la función `sns.scatterplot()` de la librería seaborn. Aprovecha todos los parámetros: `x`, `y`, `size`, `hue` y `style` (consulta la ayuda de la función [.scatterplot()](https://seaborn.pydata.org/generated/seaborn.scatterplot.html).
```
# Solución:
```
## Ejercicio 1.2.
Sobre el gráfico del ejercicio 1.1:
- Pon título al gráfico y a los ejes x e y.
- Modifica los límites del eje y para que aparezcan sólo personajes de menos de 150 Kg de peso.
- Sitúa en el gráfico los nombres de "Darth Vader" y "Anakin Skywalker". ¿Cómo es posible tengan un peso y altura tan distintos si eran la misma persona?
```
# Solución:
```
## Ejercicio 1.3.
Utiliza las *list comprehension* para calcular el cuadrado de los valores positivos de la siguiente lista:
Muestra el resultado por pantalla.
```
val = [5, 6, -1, 2, -3, -7, 9, 1]
# Solución:
```
## Ejercicio 1.4.
Construye un diccionario donde se identifique, mediante claves y valores, las siguientes características del personaje Yoda: "nombre", "altura", "peso", "edad" y "genero". Utiliza los datos de people_df.
Muestra el diccionario por pantalla.
```
# Solución:
```
## Ejercicio 1.5.
Calcula, a partir de los vectores numpy de altura y peso, definidos a continuación, el [índice de masa corporal (IMC)](https://es.wikipedia.org/wiki/%C3%8Dndice_de_masa_corporal) de los personajes de star wars contenidos en people_df:
$IMC = \frac{peso}{altura^{2}}$ donde altura está en metros y el peso en kg.
Muestra los datos por pantalla.
```
# Solución:
```
## Ejercicio 1.6.
A partir del IMC que has calculado en el ejercicio 1.5. Construye un panel con dos histogramas:
- Un histograma con toda la muestra
- Un histograma seleccionando los valores con un IMC inferior a 100.
```
# Solución:
```
## Ejercicio 1.7.
A partir del vector 1 y 2 que se definen a continuación contesta las siguientes preguntas:
- Calcula el shape, ndim, size del vector1 y vector2
- Explica cual es la diferencia entre vector1 y vector2 a partir de los que hayas observado
```
vector1 = np.hstack([altura,peso])
vector2 = np.vstack([altura,peso])
# Solución:
```
## Ejercicio 1.8.
Crea una copia de people_df llamada personajes_df y renombra las columnas con su traducción al castellano.
Muestra los 5 primeros registros del nuevo data frame con `.head()`.
```
# Solución:
```
## Ejercicio 1.9.
Haz el mismo cálculo de IMC que has hecho en 1.5. pero directamente sobre el objeto personajes_df.
Ordena el data frame de mayor a menor IMC y muestra el nombre e IMC de los personajes con IMC por encima de 30.
> ¿Sabias que IMC por encima de 30 se considera obeso?
```
# Solución:
```
## Ejercicio 1.10.
Inserta un valor faltante en los valores de IMC que sean superiores a 100 y dibuja el histograma del IMC transformado con `.plot.hist()`.
```
# Solución:
```
| github_jupyter |
```
import re
import json
from os import path
import pandas as pd
import numpy as np
from Bio import SeqIO
from Bio.Seq import Seq
from Bio.SeqRecord import SeqRecord
from Bio import AlignIO
from Bio.Align import MultipleSeqAlignment
from Bio.Align import AlignInfo
def edit_fasta_dates(cov, gene, separate_lineages=False):
input_file_alignment = '../beast/'+str(cov)+'/'+str(gene)+'/aligned_'+str(cov)+'_'+str(gene)+'.fasta'
sequences = []
with open(input_file_alignment, "r") as aligned_handle:
for virus in SeqIO.parse(aligned_handle, "fasta"):
if separate_lineages:
clade_df = separate_clades(cov, gene)
for lineage in ['A', 'B']:
if virus.id in clade_df[clade_df['clade']==lineage]['strain'].tolist():
year = virus.id.split('/')[-1]
year_check = re.compile('\d{4}')
if year_check.match(year):
sequences.append(SeqRecord(virus.seq, id=virus.id, description=virus.description))
else:
year = virus.id.split('/')[-1]
year_check = re.compile('\d{4}')
if year_check.match(year):
sequences.append(SeqRecord(virus.seq, id=virus.id, description=virus.description))
if separate_lineages:
for lineage in ['a', 'b']:
output_file_alignment = '../beast/'+str(cov)+str(lineage)+'/'+str(gene)+'/aligned_'+str(cov)+str(lineage)+'_'+str(gene)+'.fasta'
SeqIO.write(sequences, output_file_alignment, "fasta")
else:
SeqIO.write(sequences, input_file_alignment, "fasta")
covs= ['oc43']
genes = ['rdrp', 'spike', 's1', 's2']
for cov in covs:
for gene in genes:
edit_fasta_dates(cov, gene, separate_lineages=True)
covs= ['229e', 'nl63', 'oc43']
genes = ['rdrp', 'spike', 's1', 's2']
for cov in covs:
for gene in genes:
edit_fasta_dates(cov, gene)
#split oc43 into clades
def separate_clades(cov, gene):
if path.exists('../'+str(cov)+'/results/clades_'+str(gene)+'.json'):
clade_file = '../'+str(cov)+'/results/clades_'+str(gene)+'.json'
else:
clade_file = '../'+str(cov)+'/results/clades_full.json'
clade_lists = []
with open(clade_file, "r") as clade_handle:
clades = json.load(clade_handle)
for node, v in clades['nodes'].items():
if 'NODE' not in node:
clade_lists.append({'clade':v['clade_membership'],
'strain':node})
clade_df = pd.DataFrame(clade_lists)
return clade_df
```
| github_jupyter |
# Talks markdown generator for academicpages
Takes a TSV of talks with metadata and converts them for use with [academicpages.github.io](academicpages.github.io). This is an interactive Jupyter notebook ([see more info here](http://jupyter-notebook-beginner-guide.readthedocs.io/en/latest/what_is_jupyter.html)). The core python code is also in `talks.py`. Run either from the `markdown_generator` folder after replacing `talks.tsv` with one containing your data.
TODO: Make this work with BibTex and other databases, rather than Stuart's non-standard TSV format and citation style.
```
import pandas as pd
import os
```
## Data format
The TSV needs to have the following columns: title, type, url_slug, venue, date, location, talk_url, description, with a header at the top. Many of these fields can be blank, but the columns must be in the TSV.
- Fields that cannot be blank: `title`, `url_slug`, `date`. All else can be blank. `type` defaults to "Talk"
- `date` must be formatted as YYYY-MM-DD.
- `url_slug` will be the descriptive part of the .md file and the permalink URL for the page about the paper.
- The .md file will be `YYYY-MM-DD-[url_slug].md` and the permalink will be `https://[yourdomain]/talks/YYYY-MM-DD-[url_slug]`
- The combination of `url_slug` and `date` must be unique, as it will be the basis for your filenames
This is how the raw file looks (it doesn't look pretty, use a spreadsheet or other program to edit and create).
```
!cat talks.tsv
```
## Import TSV
Pandas makes this easy with the read_csv function. We are using a TSV, so we specify the separator as a tab, or `\t`.
I found it important to put this data in a tab-separated values format, because there are a lot of commas in this kind of data and comma-separated values can get messed up. However, you can modify the import statement, as pandas also has read_excel(), read_json(), and others.
```
talks = pd.read_csv("talks.tsv", sep="\t", header=0)
talks
```
## Escape special characters
YAML is very picky about how it takes a valid string, so we are replacing single and double quotes (and ampersands) with their HTML encoded equivilents. This makes them look not so readable in raw format, but they are parsed and rendered nicely.
```
html_escape_table = {
"&": "&",
'"': """,
"'": "'"
}
def html_escape(text):
if type(text) is str:
return "".join(html_escape_table.get(c,c) for c in text)
else:
return "False"
```
## Creating the markdown files
This is where the heavy lifting is done. This loops through all the rows in the TSV dataframe, then starts to concatentate a big string (```md```) that contains the markdown for each type. It does the YAML metadata first, then does the description for the individual page.
```
loc_dict = {}
for row, item in talks.iterrows():
md_filename = str(item.date) + "-" + item.url_slug + ".md"
html_filename = str(item.date) + "-" + item.url_slug
year = item.date[:4]
md = "---\ntitle: \"" + item.title + '"\n'
md += "collection: talks" + "\n"
if len(str(item.type)) > 3:
md += 'type: "' + item.type + '"\n'
else:
md += 'type: "Talk"\n'
md += "permalink: /talks/" + html_filename + "\n"
if len(str(item.venue)) > 3:
md += 'venue: "' + item.venue + '"\n'
if len(str(item.location)) > 3:
md += "date: " + str(item.date) + "\n"
if len(str(item.location)) > 3:
md += 'location: "' + str(item.location) + '"\n'
md += "---\n"
if len(str(item.talk_url)) > 3:
md += "\n[More information here](" + item.talk_url + ")\n"
if len(str(item.description)) > 3:
md += "\n" + html_escape(item.description) + "\n"
md_filename = os.path.basename(md_filename)
#print(md)
with open("../_talks/" + md_filename, 'w') as f:
f.write(md)
```
These files are in the talks directory, one directory below where we're working from.
```
!ls ../_talks
!cat ../_talks/2013-03-01-tutorial-1.md
```
| github_jupyter |
# Distributed AMFabric
```
import os
from dotenv import load_dotenv
# setup configuration
#
load_dotenv()
config = {'db_name': os.getenv("DB_NAME"),
'db_username': os.getenv("DB_USERNAME"),
'db_password': os.getenv("DB_PASSWORD"),
'db_system': os.getenv("DB_SYSTEM"),
'db_config_file_path': os.getenv("DB_CONFIG_PATH"),
'db_queries_file_path': os.getenv("DB_QUERIES_PATH"),
'fabric_graph' : 'HSG',
'scheduler_address': os.getenv("DASK_SCHEDULER")}
```
### Instantiate Distributed AMFabric object
```
from src.distributed_amfabric import DistributedAMFabric
amf = DistributedAMFabric(config=config)
```
### Create some simple training data use an SDR
```
from src.sdr import SDR
sdr_1 = SDR()
sdr_1.set_item(source_node = ('rainbow_trade', '*'),
edge=('has_rgb', 'r'),
target_node=('rgb', 'r'),
probability=1.0,
numeric=125,
numeric_min=0,
numeric_max=255
)
sdr_1.sdr
```
### Create an area in the fabric to train
```
# the length of the short term memeory - defines number of neurons in a neuro_column
#
short_term_memory = 1
# percentage margin around the previous highest high and lowest low of the matrix profile when looking for anomalies and motifs
#
mp_threshold = 0.15
# the strategy for connecting neuro_columns
#
structure = 'star'
# edges with probability below this threshold will be assumed to be zero and deleted
#
prune_threshold = 0.001
# a seed to ensure we can repeat our experiments
#
random_seed = 221166
# create a new amfabric area
#
amf.create_distributed_fabric(fabric_uid='test',
short_term_memory=short_term_memory,
mp_threshold=mp_threshold,
structure=structure,
prune_threshold=prune_threshold,
random_seed=random_seed,
restore=True)
```
### Search for the neuro_column most similar to the SDRs in short term memory
```
search_por = amf.search_for_bmu(fabric_uid='test', sdr=sdr_1, ref_id=1)
search_por
search_por = search_por.result()
search_por
```
### Learn the current SDRs in short term memory, using the results from the previous search
```
learn_por = amf.learn(fabric_uid='test', search_por=search_por, persist=True)
learn_por
learn_por = learn_por.result()
learn_por
```
### Query the fabric - basically a search for the most similar neuro_column to the SDRs in the query
```
q_result = amf.query(fabric_uid='test', sdr=sdr_1)
q_result
q_result = q_result.result()
q_result
```
### Decode the fabric
```
decode_result = amf.decode_fabric(fabric_uid='test', all_details=True,community_sdr=True)
decode_result
decode_result = decode_result.result()
decode_result
```
| github_jupyter |
```
import os
os.environ['CUDA_VISIBLE_DEVICES'] = ""
import geomloss
import matplotlib.pyplot as plt
import numpy as np
import ot
import pandas as pd
import scipy.stats as st
import seaborn
import torch
cpu = torch.device('cpu')
```
$X \sim \sum_i w_i, \delta_{X_i}$ where the $X_i$'s are i.i.d. $U([-2, 2])$ and $w_i \propto f(X_i)$ where $f$ is the pdf of a normal distribution $ \mathcal{N}(1, 1)$
```
randomState = np.random.RandomState(42)
N = 500
X = randomState.uniform(-2, 2, N)
loc=0.5
scale=1.
weights = st.norm.pdf(X, loc=loc, scale=scale)
weights /= weights.sum()
uniform_weights = np.full(N, 1/N)
```
Illustration of the regularised OT problem:
```
exactTransport = ot.bregman.empirical_sinkhorn(X[:, None], X[:, None], 1e-2, weights, uniform_weights)
# sadly the Earth moving distance doesn't work
torch_uniform_weights = torch.from_numpy(uniform_weights).to(cpu)
torch_X = torch.from_numpy(X[:, None]).to(cpu)
torch_weights = torch.from_numpy(weights).to(cpu)
def transport_from_potentials(x, f, g, w, eps):
C = (x.T - x) ** 2 / 2.
FG = f.T + g
T = torch.exp((FG - C)/eps**2) * w.unsqueeze(1)
return T.T @ x, (T.T / N ).sum(axis=1)
epsilons = [0.01, 0.05, 0.1, 0.5]
approximateTransports = []
for eps in epsilons:
biasedSampleLoss = geomloss.SamplesLoss(reach=None, potentials=True, debias=False, scaling=0.9, blur=eps)
alpha, beta = biasedSampleLoss(torch_weights, torch_X, torch_uniform_weights, torch_X)
approximateTransports.append(transport_from_potentials(torch_X, alpha, beta, torch_weights, eps))
linspace = np.sort(np.random.uniform(-3, 3, 10000))
theoretical_unweighted = st.norm.pdf(linspace, loc=loc, scale=scale)
fig, axes = plt.subplots(ncols=2, figsize=(12, 5))
weighted_sample_kde = st.kde.gaussian_kde(X, weights=weights)
exact_transport_kde = st.kde.gaussian_kde((exactTransport.T * N) @ X)
_ = axes[0].plot(linspace, weighted_sample_kde(linspace), label='weighted sample KDE')
_ = axes[0].plot(linspace, exact_transport_kde(linspace), label='unweighted transport KDE')
dataframeWeights = []
for eps, plan in zip(epsilons, approximateTransports):
approx_transport_kde = st.kde.gaussian_kde(plan[0].detach().numpy().squeeze(), weights=plan[1].detach().numpy())
dataframeWeights.append(pd.Series(data=plan[1].detach().numpy(), name=eps))
_ = axes[1].plot(linspace, approx_transport_kde(linspace), label=f'eps: {eps}')
_ = axes[1].plot(linspace, weighted_sample_kde(linspace), label='weighted sample KDE')
_ = axes[0].legend(loc='upper left')
_ = axes[1].legend(loc='upper left')
fig.savefig( 'KDEIllustration.png')
print((pd.concat(dataframeWeights, axis=1).describe(percentiles=[0.1, 0.5, 0.9]) * 1e3).to_latex(float_format='{0:.3f}'.format))
```
| github_jupyter |
```
import networkx as nx
import random
import math
import csv
import datetime
from sklearn import svm
from sklearn.tree import DecisionTreeClassifier
from sklearn.metrics import accuracy_score
from sklearn.metrics import f1_score
import multiprocessing as mp
from sklearn import svm
from sklearn.tree import DecisionTreeClassifier
from sklearn.metrics import average_precision_score
from sklearn.metrics import precision_score
from sklearn.preprocessing import normalize
import numpy as np
from sklearn import linear_model
from sklearn.metrics import log_loss
from sklearn.neural_network import MLPClassifier
from sklearn.preprocessing import StandardScaler
from sklearn.linear_model import LogisticRegression
def CommonNeighbors(u, v, g):
u_neighbors = set(g.neighbors(u))
v_neighbors = set(g.neighbors(v))
return len(u_neighbors.intersection(v_neighbors))
def common_neighbors(g, edges):
result = []
for edge in edges:
node_one, node_two = edge[0], edge[1]
num_common_neighbors = 0
try:
neighbors_one, neighbors_two = g.neighbors(node_one), g.neighbors(node_two)
for neighbor in neighbors_one:
if neighbor in neighbors_two:
num_common_neighbors += 1
result.append((node_one, node_two, num_common_neighbors))
except:
pass
return result
def AdamicAdar(u, v, g):
u_neighbors = set(g.neighbors(u))
v_neighbors = set(g.neighbors(v))
aa = 0
for i in u_neighbors.intersection(v_neighbors):
aa += 1 / math.log(len(g.neighbors(i)))
return aa
def ResourceAllocation(u, v, g):
u_neighbors = set(g.neighbors(u))
v_neighbors = set(g.neighbors(v))
ra = 0
for i in u_neighbors.intersection(v_neighbors):
ra += 1 / float(len(g.neighbors(i)))
return ra
def JaccardCoefficient(u, v, g):
u_neighbors = set(g.neighbors(u))
v_neighbors = set(g.neighbors(v))
return len(u_neighbors.intersection(v_neighbors)) / float(len(u_neighbors.union(v_neighbors)))
def PreferentialAttachment(u, v, g):
return len(g.neighbors(u))*len(g.neighbors(v))
def AllFeatures(u,v,g1, g2):
'''
the change of features in two consecutive sub graphs
'''
try:
cn = CommonNeighbors(u, v, g2)
aa = AdamicAdar(u, v, g2)
ra = ResourceAllocation(u, v, g2)
jc = JaccardCoefficient(u, v, g2)
pa = PreferentialAttachment(u, v, g2)
delta_cn = cn - CommonNeighbors(u, v, g1)
delta_aa = aa - AdamicAdar(u, v, g1)
delta_ra = ra - ResourceAllocation(u, v, g1)
delta_jc = jc - JaccardCoefficient(u, v, g1)
delta_pa = pa - PreferentialAttachment(u, v, g1)
return {"cn":cn, "aa": aa, "ra":ra, "jc":jc, "pa":pa,
"delta_cn": delta_cn, "delta_aa": delta_aa, "delta_ra": delta_ra,
"delta_jc": delta_jc, "delta_pa": delta_pa}
except:
pass
feature_set = [common_neighbors,
nx.resource_allocation_index,
nx.jaccard_coefficient,
nx.adamic_adar_index,
nx.preferential_attachment
]
def produce_fake_edge(g, neg_g,num_test_edges):
i = 0
while i < num_test_edges:
edge = random.sample(g.nodes(), 2)
try:
shortest_path = nx.shortest_path_length(g,source=edge[0],target=edge[1])
if shortest_path >= 2:
neg_g.add_edge(edge[0],edge[1], positive="False")
i += 1
except:
pass
def create_graph_from_file(filename):
print("----------------build graph--------------------")
f = open(filename, "rb")
g = nx.read_edgelist(f)
return g
def sample_extraction(g, pos_num, neg_num, neg_mode, neg_distance=2, delete=1):
"""
:param g: the graph
:param pos_num: the number of positive samples
:param neg_num: the number of negative samples
:param neg_distance: the distance between two nodes in negative samples
:param delete: if delete ==0, don't delete positive edges from graph
:return: pos_sample is a list of positive edges, neg_sample is a list of negative edges
"""
print("----------------extract positive samples--------------------")
# randomly select pos_num as test edges
pos_sample = random.sample(g.edges(), pos_num)
sample_g = nx.Graph()
sample_g.add_edges_from(pos_sample, positive="True")
nx.write_edgelist(sample_g, r"data\sample_positive_" +str(pos_num)+ ".txt", data=['positive'])
# adding non-existing edges
print("----------------extract negative samples--------------------")
i = 0
neg_g = nx.Graph()
produce_fake_edge(g,neg_g,neg_num)
nx.write_edgelist(neg_g, r"data\sample_negative_" +str(neg_num)+ ".txt", data=["positive"])
neg_sample = neg_g.edges()
neg_g.add_edges_from(pos_sample,positive="True")
nx.write_edgelist(neg_g, r"data\sample_combine_" +str(pos_num + neg_num)+ ".txt", data=["positive"])
# remove the positive sample edges, the rest is the training set
if delete == 0:
return pos_sample, neg_sample
else:
g.remove_edges_from(pos_sample)
nx.write_edgelist(g, r"data\training.txt", data=False)
return pos_sample, neg_sample
def feature_extraction(g, pos_sample, neg_sample, feature_name, model="single", combine_num=5):
data = []
if model == "single":
print ("-----extract feature:", feature_name.__name__, "----------")
preds = feature_name(g, pos_sample)
feature = [feature_name.__name__] + [i[2] for i in preds]
label = ["label"] + ["Pos" for i in range(len(feature))]
preds = feature_name(g, neg_sample)
feature1 = [i[2] for i in preds]
feature = feature + feature1
label = label + ["Neg" for i in range(len(feature1))]
data = [feature, label]
data = transpose(data)
print("----------write the feature to file---------------")
write_data_to_file(data, r"data\features_" + model + "_" + feature_name.__name__ + ".csv")
else:
label = ["label"] + ["1" for i in range(len(pos_sample))] + ["0" for i in range(len(neg_sample))]
for j in feature_name:
print ("-----extract feature:", j.__name__, "----------")
preds = j(g, pos_sample)
feature = [j.__name__] + [i[2] for i in preds]
preds = j(g, neg_sample)
feature = feature + [i[2] for i in preds]
data.append(feature)
data.append(label)
data = transpose(data)
print("----------write the features to file---------------")
write_data_to_file(data, r"data\features_" + model + "_" + str(combine_num) + ".csv")
return data
def write_data_to_file(data, filename):
csvfile = open(filename, "w")
writer = csv.writer(csvfile)
for i in data:
writer.writerow(i)
csvfile.close()
def transpose(data):
return [list(i) for i in zip(*data)]
def main(filename=r"data\facebook_combined.txt", pos_num=0.1, neg_num=0.1, model="combined", combine_num=1,
feature_name=common_neighbors, neg_mode="hard"):
if combine_num==2:
pos_num=0.08
neg_num=0.08
if combine_num==3:
pos_num=0.1
neg_num=0.1
g = create_graph_from_file(filename)
num_edges = g.number_of_edges()
pos_num = int(num_edges * pos_num)
neg_num = int(num_edges * neg_num)
pos_sample, neg_sample = sample_extraction(g, pos_num, neg_num,neg_mode)
train_data = feature_extraction(g, pos_sample, neg_sample, feature_name, model, combine_num)
#______________________Entry Point________________________
#Fn: Name of data set you want to run this code for, and cn is a integer for that dataset(any integer will work but different for each dataset)
#By default it is set to Twitter Data Set
#The project was run using Facebook and Twitter dataset but it works with any social network dataset from http://snap.stanford.edu/data/
#Following Scoring Methods are used to construct feature Set----
#common_neighbors,resource_allocation_index, jaccard_coefficient, adamic_adar_index, preferential_attachment
#SVM ANN and Logistic Regresssion is used for classificaion
fn=r"data\facebook_combined.txt";
cn=3;
#Run this line to genrate feature Set
main(filename=fn,model="combined",combine_num=cn, feature_name=feature_set, neg_mode="easy")
r=np.loadtxt(open(r"data\features_combined_"+str(cn)+".csv", "rb"), delimiter=",", skiprows=1);
l,b=r.shape;
np.random.shuffle(r);
train_l=int(0.75*l)
X_train=r[0:train_l,0:b-1]
Y_train=r[0:train_l,b-1]
test = r[train_l:l,:]
X_test=test[:,0:b-1]
Y_test=test[:,b-1]
X_train = normalize(X_train, axis=0, norm='max')
X_test = normalize(X_test, axis=0, norm='max')
scaler = StandardScaler()
scaler.fit(X_train)
X_train = scaler.transform(X_train)
X_test = scaler.transform(X_test)
len(r)
def mySvm(training, training_labels, testing, testing_labels):
#Support Vector Machine
start = datetime.datetime.now()
clf = svm.SVC()
clf.fit(training, training_labels)
print ("+++++++++ Finishing training the SVM classifier ++++++++++++")
result = clf.predict(testing)
print ("SVM accuracy:", accuracy_score(testing_labels, result))
#keep the time
finish = datetime.datetime.now()
print ((finish-start).seconds)
return result
#Run this to for SVM classification
svmres=mySvm(X_train,Y_train,X_test,Y_test)
def logistic(training, training_labels, testing, testing_labels):
clf = LogisticRegression(random_state=0, solver='lbfgs',multi_class='ovr')
start = datetime.datetime.now()
clf.fit(training, training_labels)
result=clf.predict(testing)
print ("+++++++++ Finishing training the Linear classifier ++++++++++++")
print ("Linear accuracy:", accuracy_score(testing_labels, result))
#keep the time
finish = datetime.datetime.now()
print ((finish-start).seconds)
return result
#Run this for Logistic Regression
logres=logistic(X_train,Y_train,X_test,Y_test)
def ANN(training, training_labels, testing, testing_labels):
clf = MLPClassifier(solver='adam', alpha=1e-5,hidden_layer_sizes=(12,9), random_state=1, max_iter=1000)
start = datetime.datetime.now()
clf.fit(training, training_labels)
print ("+++++++++ Finishing training the ANN classifier ++++++++++++")
result = clf.predict(testing)
print ("ANN accuracy:", accuracy_score(testing_labels, result))
#keep the time
finish = datetime.datetime.now()
print ((finish-start).seconds)
return result
# Run this for ANN classification
annres=ANN(X_train,Y_train,X_test,Y_test)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/souravgopal25/DeepLearnigNanoDegree/blob/master/cifar10_cnn_augmentation.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Convolutional Neural Networks
---
In this notebook, we train a **CNN** to classify images from the CIFAR-10 database.
The images in this database are small color images that fall into one of ten classes
### Test for [CUDA](http://pytorch.org/docs/stable/cuda.html)
Since these are larger (32x32x3) images, it may prove useful to speed up your training time by using a GPU. CUDA is a parallel computing platform and CUDA Tensors are the same as typical Tensors, only they utilize GPU's for computation.
```
import torch
import numpy as np
# check if CUDA is available
train_on_gpu = torch.cuda.is_available()
if not train_on_gpu:
print('CUDA is not available. Training on CPU ...')
else:
print('CUDA is available! Training on GPU ...')
```
---
## Load and Augment the [Data](http://pytorch.org/docs/stable/torchvision/datasets.html)
Downloading may take a minute. We load in the training and test data, split the training data into a training and validation set, then create DataLoaders for each of these sets of data.
#### Augmentation
In this cell, we perform some simple [data augmentation](https://medium.com/nanonets/how-to-use-deep-learning-when-you-have-limited-data-part-2-data-augmentation-c26971dc8ced) by randomly flipping and rotating the given image data. We do this by defining a torchvision `transform`, and you can learn about all the transforms that are used to pre-process and augment data, [here](https://pytorch.org/docs/stable/torchvision/transforms.html).
#### TODO: Look at the [transformation documentation](https://pytorch.org/docs/stable/torchvision/transforms.html); add more augmentation transforms, and see how your model performs.
This type of data augmentation should add some positional variety to these images, so that when we train a model on this data, it will be robust in the face of geometric changes (i.e. it will recognize a ship, no matter which direction it is facing). It's recommended that you choose one or two transforms.
```
from torchvision import datasets
import torchvision.transforms as transforms
from torch.utils.data.sampler import SubsetRandomSampler
# number of subprocesses to use for data loading
num_workers = 0
# how many samples per batch to load
batch_size = 20
# percentage of training set to use as validation
valid_size = 0.2
# convert data to a normalized torch.FloatTensor
transform = transforms.Compose([
transforms.RandomHorizontalFlip(), # randomly flip and rotate
transforms.RandomRotation(10),
transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))
])
# choose the training and test datasets
train_data = datasets.CIFAR10('data', train=True,
download=True, transform=transform)
test_data = datasets.CIFAR10('data', train=False,
download=True, transform=transform)
# obtain training indices that will be used for validation
num_train = len(train_data)
indices = list(range(num_train))
np.random.shuffle(indices)
split = int(np.floor(valid_size * num_train))
train_idx, valid_idx = indices[split:], indices[:split]
# define samplers for obtaining training and validation batches
train_sampler = SubsetRandomSampler(train_idx)
valid_sampler = SubsetRandomSampler(valid_idx)
# prepare data loaders (combine dataset and sampler)
train_loader = torch.utils.data.DataLoader(train_data, batch_size=batch_size,
sampler=train_sampler, num_workers=num_workers)
valid_loader = torch.utils.data.DataLoader(train_data, batch_size=batch_size,
sampler=valid_sampler, num_workers=num_workers)
test_loader = torch.utils.data.DataLoader(test_data, batch_size=batch_size,
num_workers=num_workers)
# specify the image classes
classes = ['airplane', 'automobile', 'bird', 'cat', 'deer',
'dog', 'frog', 'horse', 'ship', 'truck']
```
### Visualize a Batch of Training Data
```
import matplotlib.pyplot as plt
%matplotlib inline
# helper function to un-normalize and display an image
def imshow(img):
img = img / 2 + 0.5 # unnormalize
plt.imshow(np.transpose(img, (1, 2, 0))) # convert from Tensor image
# obtain one batch of training images
dataiter = iter(train_loader)
images, labels = dataiter.next()
images = images.numpy() # convert images to numpy for display
# plot the images in the batch, along with the corresponding labels
fig = plt.figure(figsize=(25, 4))
# display 20 images
for idx in np.arange(20):
ax = fig.add_subplot(2, 20/2, idx+1, xticks=[], yticks=[])
imshow(images[idx])
ax.set_title(classes[labels[idx]])
```
### View an Image in More Detail
Here, we look at the normalized red, green, and blue (RGB) color channels as three separate, grayscale intensity images.
```
rgb_img = np.squeeze(images[3])
channels = ['red channel', 'green channel', 'blue channel']
fig = plt.figure(figsize = (36, 36))
for idx in np.arange(rgb_img.shape[0]):
ax = fig.add_subplot(1, 3, idx + 1)
img = rgb_img[idx]
ax.imshow(img, cmap='gray')
ax.set_title(channels[idx])
width, height = img.shape
thresh = img.max()/2.5
for x in range(width):
for y in range(height):
val = round(img[x][y],2) if img[x][y] !=0 else 0
ax.annotate(str(val), xy=(y,x),
horizontalalignment='center',
verticalalignment='center', size=8,
color='white' if img[x][y]<thresh else 'black')
```
---
## Define the Network [Architecture](http://pytorch.org/docs/stable/nn.html)
This time, you'll define a CNN architecture. Instead of an MLP, which used linear, fully-connected layers, you'll use the following:
* [Convolutional layers](https://pytorch.org/docs/stable/nn.html#conv2d), which can be thought of as stack of filtered images.
* [Maxpooling layers](https://pytorch.org/docs/stable/nn.html#maxpool2d), which reduce the x-y size of an input, keeping only the most _active_ pixels from the previous layer.
* The usual Linear + Dropout layers to avoid overfitting and produce a 10-dim output.
A network with 2 convolutional layers is shown in the image below and in the code, and you've been given starter code with one convolutional and one maxpooling layer.
#### TODO: Define a model with multiple convolutional layers, and define the feedforward metwork behavior.
The more convolutional layers you include, the more complex patterns in color and shape a model can detect. It's suggested that your final model include 2 or 3 convolutional layers as well as linear layers + dropout in between to avoid overfitting.
It's good practice to look at existing research and implementations of related models as a starting point for defining your own models. You may find it useful to look at [this PyTorch classification example](https://github.com/pytorch/tutorials/blob/master/beginner_source/blitz/cifar10_tutorial.py) or [this, more complex Keras example](https://github.com/keras-team/keras/blob/master/examples/cifar10_cnn.py) to help decide on a final structure.
#### Output volume for a convolutional layer
To compute the output size of a given convolutional layer we can perform the following calculation (taken from [Stanford's cs231n course](http://cs231n.github.io/convolutional-networks/#layers)):
> We can compute the spatial size of the output volume as a function of the input volume size (W), the kernel/filter size (F), the stride with which they are applied (S), and the amount of zero padding used (P) on the border. The correct formula for calculating how many neurons define the output_W is given by `(W−F+2P)/S+1`.
For example for a 7x7 input and a 3x3 filter with stride 1 and pad 0 we would get a 5x5 output. With stride 2 we would get a 3x3 output.
```
import torch.nn as nn
import torch.nn.functional as F
# define the CNN architecture
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
# convolutional layer (sees 32x32x3 image tensor)
self.conv1 = nn.Conv2d(3, 16, 3, padding=1)
# convolutional layer (sees 16x16x16 tensor)
self.conv2 = nn.Conv2d(16, 32, 3, padding=1)
# convolutional layer (sees 8x8x32 tensor)
self.conv3 = nn.Conv2d(32, 64, 3, padding=1)
# max pooling layer
self.pool = nn.MaxPool2d(2, 2)
# linear layer (64 * 4 * 4 -> 500)
self.fc1 = nn.Linear(64 * 4 * 4, 500)
# linear layer (500 -> 10)
self.fc2 = nn.Linear(500, 10)
# dropout layer (p=0.25)
self.dropout = nn.Dropout(0.25)
def forward(self, x):
# add sequence of convolutional and max pooling layers
x = self.pool(F.relu(self.conv1(x)))
x = self.pool(F.relu(self.conv2(x)))
x = self.pool(F.relu(self.conv3(x)))
# flatten image input
x = x.view(-1, 64 * 4 * 4)
# add dropout layer
x = self.dropout(x)
# add 1st hidden layer, with relu activation function
x = F.relu(self.fc1(x))
# add dropout layer
x = self.dropout(x)
# add 2nd hidden layer, with relu activation function
x = self.fc2(x)
return x
# create a complete CNN
model = Net()
print(model)
# move tensors to GPU if CUDA is available
if train_on_gpu:
model.cuda()
```
### Specify [Loss Function](http://pytorch.org/docs/stable/nn.html#loss-functions) and [Optimizer](http://pytorch.org/docs/stable/optim.html)
Decide on a loss and optimization function that is best suited for this classification task. The linked code examples from above, may be a good starting point; [this PyTorch classification example](https://github.com/pytorch/tutorials/blob/master/beginner_source/blitz/cifar10_tutorial.py) or [this, more complex Keras example](https://github.com/keras-team/keras/blob/master/examples/cifar10_cnn.py). Pay close attention to the value for **learning rate** as this value determines how your model converges to a small error.
#### TODO: Define the loss and optimizer and see how these choices change the loss over time.
```
import torch.optim as optim
# specify loss function (categorical cross-entropy)
criterion = nn.CrossEntropyLoss()
# specify optimizer
optimizer = optim.SGD(model.parameters(), lr=0.01)
```
---
## Train the Network
Remember to look at how the training and validation loss decreases over time; if the validation loss ever increases it indicates possible overfitting.
```
# number of epochs to train the model
n_epochs = 30
valid_loss_min = np.Inf # track change in validation loss
for epoch in range(1, n_epochs+1):
# keep track of training and validation loss
train_loss = 0.0
valid_loss = 0.0
###################
# train the model #
###################
model.train()
for batch_idx, (data, target) in enumerate(train_loader):
# move tensors to GPU if CUDA is available
if train_on_gpu:
data, target = data.cuda(), target.cuda()
# clear the gradients of all optimized variables
optimizer.zero_grad()
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the batch loss
loss = criterion(output, target)
# backward pass: compute gradient of the loss with respect to model parameters
loss.backward()
# perform a single optimization step (parameter update)
optimizer.step()
# update training loss
train_loss += loss.item()*data.size(0)
######################
# validate the model #
######################
model.eval()
for batch_idx, (data, target) in enumerate(valid_loader):
# move tensors to GPU if CUDA is available
if train_on_gpu:
data, target = data.cuda(), target.cuda()
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the batch loss
loss = criterion(output, target)
# update average validation loss
valid_loss += loss.item()*data.size(0)
# calculate average losses
train_loss = train_loss/len(train_loader.dataset)
valid_loss = valid_loss/len(valid_loader.dataset)
# print training/validation statistics
print('Epoch: {} \tTraining Loss: {:.6f} \tValidation Loss: {:.6f}'.format(
epoch, train_loss, valid_loss))
# save model if validation loss has decreased
if valid_loss <= valid_loss_min:
print('Validation loss decreased ({:.6f} --> {:.6f}). Saving model ...'.format(
valid_loss_min,
valid_loss))
torch.save(model.state_dict(), 'model_augmented.pt')
valid_loss_min = valid_loss
```
### Load the Model with the Lowest Validation Loss
```
model.load_state_dict(torch.load('model_augmented.pt'))
```
---
## Test the Trained Network
Test your trained model on previously unseen data! A "good" result will be a CNN that gets around 70% (or more, try your best!) accuracy on these test images.
```
# track test loss
test_loss = 0.0
class_correct = list(0. for i in range(10))
class_total = list(0. for i in range(10))
model.eval()
# iterate over test data
for batch_idx, (data, target) in enumerate(test_loader):
# move tensors to GPU if CUDA is available
if train_on_gpu:
data, target = data.cuda(), target.cuda()
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the batch loss
loss = criterion(output, target)
# update test loss
test_loss += loss.item()*data.size(0)
# convert output probabilities to predicted class
_, pred = torch.max(output, 1)
# compare predictions to true label
correct_tensor = pred.eq(target.data.view_as(pred))
correct = np.squeeze(correct_tensor.numpy()) if not train_on_gpu else np.squeeze(correct_tensor.cpu().numpy())
# calculate test accuracy for each object class
for i in range(batch_size):
label = target.data[i]
class_correct[label] += correct[i].item()
class_total[label] += 1
# average test loss
test_loss = test_loss/len(test_loader.dataset)
print('Test Loss: {:.6f}\n'.format(test_loss))
for i in range(10):
if class_total[i] > 0:
print('Test Accuracy of %5s: %2d%% (%2d/%2d)' % (
classes[i], 100 * class_correct[i] / class_total[i],
np.sum(class_correct[i]), np.sum(class_total[i])))
else:
print('Test Accuracy of %5s: N/A (no training examples)' % (classes[i]))
print('\nTest Accuracy (Overall): %2d%% (%2d/%2d)' % (
100. * np.sum(class_correct) / np.sum(class_total),
np.sum(class_correct), np.sum(class_total)))
```
| github_jupyter |
# SLU02 Command Line & Text Editor - Exercise Notebook
***
```
# For evaluation purposes, please import this package:
import hashlib
```
### Exercise 1: Where is home?
<img src="./assets/tortoise_home.png" width="400"/>
Imagine that you have a Linux user called `tortoise`, what is the correct path for this user's *home directory*?
* Option A: */root*
* Option B: */tortoise*
* Option C: */root/tortoise*
* Option D: */home/tortoise*
```
# exercise_1 =
# YOUR CODE HERE
raise NotImplementedError()
ex_1 = '50c9e8d5fc98727b4bbc93cf5d64a68db647f04f'
assert isinstance(ex_1, str), "The variable should be a string."
assert ex_1 == hashlib.sha1(bytes(exercise_1, encoding='utf8')).hexdigest(), "The value of the option is incorrect."
```
### Exercise 2.1: Changing directory
Now, for our same `tortoise` user, if our current working directory is `/var/lib/systemd/`, how do we go this user's *home directory* ?
* Option A: pwd
* Option B: cd /home
* Option C: cd ~
* Option D: cd tortoise
```
# exercise_2_1 =
# YOUR CODE HERE
raise NotImplementedError()
ex_2_1 = '484d52691fcadbfabec5a318d1cf9692c7f293cbc8c1d5f22b2d839b'
assert isinstance(ex_2_1,str), "The variable should be a string."
assert ex_2_1 == hashlib.sha224(bytes(exercise_2_1, encoding='utf8')).hexdigest(), "The value of the option is incorrect."
```
### Exercise 2.2: Changing directory
Now, for our same `tortoise` user,
if our current working directory is `/var/lib/systemd/` and we run
`cd ~/../..`,
what will our current working directory be?
* Option A: /home/tortoise
* Option B: /home/tortoise/home/tortoise
* Option C: ~
* Option D: /
```
# exercise_2_2 =
# YOUR CODE HERE
raise NotImplementedError()
ex_2_2 = '7497e88b8c64062edfb2f30b9c5f845552b841d1fbae97ab3f224f74'
assert isinstance(ex_2_2, str), "The variable should be a string."
assert ex_2_2 == hashlib.sha224(bytes(exercise_2_2, encoding='utf8')).hexdigest(), "The value of the option is incorrect."
```
### Exercise 3: Rename
How do we rename a non empty folder from `tortoise` to `tortoise-new`?
- Option A: *rename tortoise tortoise-new*
- Option B: *rn tortoise tortoise-new*
- Option C: *mv tortoise tortoise-new*
- Option D: *mv -r tortoise tortoise-new*
```
# exercise_3 =
# YOUR CODE HERE
raise NotImplementedError()
ex_3 = '484d52691fcadbfabec5a318d1cf9692c7f293cbc8c1d5f22b2d839b'
assert isinstance(ex_3, str), "The variable should be a string."
assert ex_3 == hashlib.sha224(bytes(exercise_3, encoding='utf8')).hexdigest(), "The value of the option is incorrect."
```
### Exercise 4: Copy
`tortoise` is not happy. This user needs to copy the `baby-tortoise` folder inside the `tortoise-family` folder.
What command should we use?
- Option A: *mv baby-tortoise tortoise-family*
- Option B: *cp baby-tortoise tortoise-family*
- Option C: *cp -a baby-tortoise tortoise-family*
- Option D: *cp -r baby-tortoise tortoise-family*
```
# exercise_4 =
# YOUR CODE HERE
raise NotImplementedError()
ex_4 = '7497e88b8c64062edfb2f30b9c5f845552b841d1fbae97ab3f224f74'
assert isinstance(ex_4, str), "The variable should be a string."
assert ex_4 == hashlib.sha224(bytes(exercise_4, encoding='utf8')).hexdigest(), "The value of the option is incorrect."
```
### Exercise 5: Create Directories and files
<img src="./assets/directories.png" width="400"/>
Create the Directory structure in the image above, including the files.
were you able to do it?
* Option A: YES
* Option B: NO, and I'll ask some question on Slack to help me figure this out!
```
# exercise_5 =
# YOUR CODE HERE
raise NotImplementedError()
ex_5 = '5cfe2cddbb9940fb4d8505e25ea77e763a0077693dbb01b1a6aa94f2'
assert isinstance(ex_5, str), "The variable should be a string."
assert ex_5 == hashlib.sha224(bytes(exercise_5, encoding='utf8')).hexdigest(), "The value of the option is incorrect."
```
### Exercise 6: Remove
<img src="./assets/tortoise.jpg" width="400"/>
Before you start questioning whether you have a weird fixation over tortoises, you might want to remove some directories and files.
What is the correct command to remove the `tortoisephilia` folder (which has some tortoise pictures)?
* Option A: `drop tortoisephilia`
* Option B: `rm tortoisephilia/*`
* Option C: `remove tortoisephilia`
* Option D: `rm -r tortoisephilia`
```
# exercise_6 =
# YOUR CODE HERE
raise NotImplementedError()
ex_6 = '7497e88b8c64062edfb2f30b9c5f845552b841d1fbae97ab3f224f74'
assert isinstance(ex_6, str), "The variable should be a string."
assert ex_6 == hashlib.sha224(bytes(exercise_6, encoding='utf8')).hexdigest(), "The value of the option is incorrect."
```
### Exercise 7:
Our `tortoise` created a `tortoise_codes.py` file with some Python code (as all good tortoises do), now he needs to check what is stored in the file in a simple and easy way.
What is the correct command for that procedure?
- Option A: *cat tortoise_codes.py*
- Option B: *read tortoise_codes.py*
- Option C: *ls tortoise_codes.py*
- Option D: *touch tortoise_codes.py*
```
# exercise_7 =
# YOUR CODE HERE
raise NotImplementedError()
ex_7 = 'ad14aaf25020bef2fd4e3eb5ec0c50272cdfd66074b0ed037c9a11254321aac0729985374beeaa5b80a504d048be1864'
assert isinstance(ex_7,str), "The variable should be a string."
assert ex_7 == hashlib.sha384(bytes(exercise_7, encoding='utf8')).hexdigest(), "The value of the option is incorrect."
```
### Exercise 8: Visual Studio Code (VS Code)
Managing to install and set VS Code up is part of this SLU, so if you have any problems please do contact an instructor!
It is very important that you get it running properly.
- Option A: Rock On! Everything is working fine :)
- Option B: NO, I am still struggling with it. It's this week's SLU instructor's fault!!!
```
# YOUR CODE HERE
raise NotImplementedError()
# exercise_8 =
ex_8 = '559aead08264d5795d3909718cdd05abd49572e84fe55590eef31a88a08fdffd'
assert isinstance(ex_8,str), "The variable should be a string."
assert ex_8 == hashlib.sha256(bytes(exercise_8, encoding='utf8')).hexdigest(), "The value of the option is incorrect."
```
### Exercise 9: Running basic Python programs
To finish this SLU in style:
1. create a folder in the location of your choice
1. open that folder in VS Code
1. in VS Code, create a Python file that prints your name, and 2 other files that print 2 other names
1. activate your virtual environment
1. run those Python files and check that the names are printed!
Were you able to do this?
- Option A: No, I've been stuck on this for more than 15 min! I need to ask for help on Slack!
- Option B: Yes, I did it!
```
# YOUR CODE HERE
raise NotImplementedError()
# exercise_9 =
ex_9 = 'df7e70e5021544f4834bbee64a9e3789febc4be81470df629cad6ddb03320a5c'
assert isinstance(ex_9,str), "The variable should be a string."
assert ex_9 == hashlib.sha256(bytes(exercise_9, encoding='utf8')).hexdigest(), "The value of the option is incorrect."
```
# Submit your work!
To submit your work, [get your slack id](https://moshfeu.medium.com/how-to-find-my-member-id-in-slack-workspace-d4bba942e38c) and fill it in the `slack_id` variable.
Example: `slack_id = "UTS63FC02"`
```
# YOUR CODE HERE
raise NotImplementedError()
# slack_id =
from submit import submit
assert slack_id is not None
submit(slack_id=slack_id, learning_unit=2)
```
| github_jupyter |
# Day 3. Interactive Controls
* [IPython Widgets](#1)
* [Bokeh](#2)
```
from IPython.display import IFrame
from IPython.display import YouTubeVideo
from ipywidgets import interact, interactive, fixed
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
%matplotlib inline
import pandas as pd
from cryptography.fernet import Fernet
import base64
def encrypt(string, key):
keygen = lambda x : base64.urlsafe_b64encode(x.encode() + b' '*(32 - len(x)))
cipher = Fernet(keygen(key))
return cipher.encrypt(string.encode())
def decrypt(string, key):
keygen = lambda x : base64.urlsafe_b64encode(x.encode() + b' '*(32 - len(x)))
cipher = Fernet(keygen(key))
return print(cipher.decrypt(string.encode()).decode())
```
<a id='1'></a>
# IPython Widgets
```
IFrame(src='https://api.kaltura.nordu.net/p/310/sp/31000/embedIframeJs/uiconf_id/23449977/partner_id/310?iframeembed=true&playerId=kaltura_player&entry_id=0_18ipnucr&flashvars[streamerType]=auto&flashvars[localizationCode]=en&flashvars[leadWithHTML5]=true&flashvars[sideBarContainer.plugin]=true&flashvars[sideBarContainer.position]=left&flashvars[sideBarContainer.clickToClose]=true&flashvars[chapters.plugin]=true&flashvars[chapters.layout]=vertical&flashvars[chapters.thumbnailRotator]=false&flashvars[streamSelector.plugin]=true&flashvars[EmbedPlayer.SpinnerTarget]=videoHolder&flashvars[dualScreen.plugin]=true&&wid=0_m97dwa52', width=608, height=402)
```
## Task 1
Create an interactive function that returns the number of words in a text. The user interface should consist of a text area. Hint: use [split](https://docs.python.org/3/library/stdtypes.html#str.split).
```
answer = 'gAAAAABcBE3nc21nV9chE1Qim80hFg5iHLw0-5jQcV1gtsFX4MN9_by8PWa7XcN80CGvShXFmh33AOaB6YCuTmTnNDGJmjQQJD4ip_bRIpdrRYSQTngImF4o4bi2Wv5fOLhk1dparYfliwieqb-yVi6b2jkTDhim6Ot9ipQ2hXgv9S_qFwweovdbN5VKr9zHgFCsrXDjQPRJ5WGrb7Ye55-1UhtkF2gHUvKhjIgxvzKI07qWfKTamLLVXns96RtSK6a6-gVQWlDS'
decrypt(answer,'Skurup')
@interact(text='hello')
def f(text):
print(text)
@interact(text='',)
def f(text):
n = 0
if len(text)>0:
n = len( text.split(' ') )
print('Number of words:',n)
```
## Task 2
Create an interactive function that returns the number of words and the number of characters in a text. The user interface should consist of a dropdown menu and a text area. Hint: use [split](https://docs.python.org/3/library/stdtypes.html#str.split) and str.replace().
```
answer = 'gAAAAABcBFWm9q_kCXaTRs_seAM7b8XtKB182n80Hojlw-eyY0eqK-W9ZukhIeWf_4cP1XNEZ2273RH0Rcp3q59ex-sYl6kafEeJV0cJnIqnzt3h0BKYVFHH_g5qPhOFPT9Kf3sLf4ZX_pUU-jELD_7MTtI2yMeBhA9aTKg55ylQOOKb-YG6X9uD80VF1VdqUX2QpT0HJIAxaotyuTU_l5OCsr4ZbjbjNROXy-zU9UgfmMlBxvSH02wD-VJzWY0PB5ZkgK-oi0mJI5NhN7tWnauQ3Upn2S7AhlDqEjTHgfrMT2J8aV2PLgU3efkO_njxrZDATJuipghPaVbpPG-a5p-vz6IW_oseWsjQAUyAj1mWWZwAYSTbQrzd1Hz2o4ChxrrseY-vMW3JLA8oAZNsrmftmYxaIBhBMFXRcJ12yrVNV5RrBo6MnaqacI29Pnqb5vVkRH42QeQgkvcpQLPzWb8zzJDBPDXYXg=='
decrypt(answer,)
```
## Task 3
Make an interactive plot of the following function with two sliding bars to control $\nu$ and $\Gamma$
\begin{equation}
I(t) = \cos{(2 \pi \nu t)} \exp{(-t 2 \pi \Gamma)}
\end{equation}
1. Define a function of $\nu$ and $\Gamma$ that plots $I(t)$
2. Use interactive to generate the sliders coupled to the parameters $\nu$ and $\Gamma$
3. Modify the description of the children (the two sliders)
Complete the following code
```.py
def plot_cos_decay(freq=1,gamma=.2):
def cos_decay(time,freq,gamma):
cos = np.cos(2*np.pi*freq*time)
exp = np.exp(-time*2*np.pi*gamma)
return exp*cos
time = np.arange(0,10,.001)
fig = plt.figure(figsize=(3.5,4))
ax = plt.axes()
ax.plot(,,color=plt.get_cmap('tab10')(3), lw=1)
ax.set_ylabel('$I(t)$',fontsize=12)
ax.set_xlabel(r'Time',fontsize=12)
ax.set_xlim(0,10)
interactive_plot = interactive( )
interactive_plot.children[0].description=r'$f$' # slider
interactive_plot```
```
answer = 'gAAAAABcBvmULdmxrFtMQ1AVeXU44jtvp8aF3jqh8G4ot7SRg6STnut7NJ7zKy57iriNHxyaHG0P6Ut8z5W5sC3QSvVLfh4C63Bg2xv2OLZJjjqUNsPRNv_KiAK1qU1y8qBJYAY_zvkLQcQyEozLf0bq_0DRpwIEmM_8Q6PTpq_K51D5rM2cDxloCh1xE_Lqysuo9PAclHsIbYR6aJVvoK2s8vD42Sx5O5QMNvxssbYgXvU2KZCazLYu9Y9Ks35E0reqmu77OORCQZjJKFpc_g7YfTj_nv3w7jeW43I17s8xENkGgcQtsXAM1UO-B6w8MtaNGtszv1uPaYVjJ-APl6nCaChqSCNLBFXLoxuGjOuLF5txCzRf8SiDRgHgEIx6Ni74AYRx8hftB53jGBwwNV12MjVis5l7gL2rH_6DOL3ltJiGobNUsZLoos0VKsNvZn49rnO7Z_fqWonq0gK8UfzKzceSJcMpufmDOH4KZW4wCB6iJSp0lzNQXTUxRt4_V99ky2DNrb7ReHeU-phWt1-vEeTD1o1GwT2NYSKAXEFIhjRszNkWwbIxLMxLOqlDKZhO-YcvG5KObDbIzKIsI3Sth0HLhKK4CPE0ZPL7uPWh6eQv-KmsvV_NMungPShD9185P5uYdbqZ-23BOdBtwCZdBWi7H-GgvmG_NpINSQ_jeLv3qgkNgEbQvbOkrzRDCYxWfneVH7z3U3sSU3zK_Qlnjheo4SaByUWGtRXRZsJgFDHrhoFPpAxh88ulpGjXUBhtBHc_ypfX6NGNxjPjpV1_vWYV4mookP-xgkSziaHCc1biCBT0K3N3WcCCDUj6Imy1ll3aT-eaMeA0rrr3OvLcNYobt08iC6JqX-XoJ62twDNJUfZOsuIzHdwVjq3q3p8pGcadfO1Vz81sFp3NabqoyZVof79UwW0AUChsYFvpqjwrz4xuA8Jftynyq8_n4anKwyUqen3BD3xt1epnWCl6QYMd5y9CDA=='
decrypt(answer,)
```
## Task 4
Add an inset in the previous plot showing the Fourier Transform of the decaying cosine
\begin{equation}
I(\nu) = \int_0^\infty \mathrm{d}t \cos{(2 \pi \nu t)}\; I(t)
\end{equation}
The plot in the inset should also be coupled to the sliders.
<img src="figs/FT.png" width="400" align="left"/>
```
answer = 'gAAAAABcBvkEo2TnSlwfLSy7cy53MY3K7wBcVtKvQEww8lWVRrra5u4TWq7ro9o1IoBRq1V-CVPp5QIrBlFWv7XHClZtUkk14YzW9ChEKponzkQo7-L9xXUPSEMlqckftQAAxiF5FBz1Od4azI7bYwLFQb7RZSCG2MS82x1OEXKv9BfSp3yfN_x-gb5Cnl8QsCTlkun8nBJv5j6xIL-dLpfJU4IO5Pwifcsu_eJj4QDYlVNwB3u_6mWvQ5rWox2LTt8eJHJkz9GbGFxGylGhApKyPsbItb7KDrCxRgW4gh1w0IwEInF8vNbkUwlQoXjhpb0jL6f2cx4we_mVz_OfuePnW9R-gET0JXm7R9hW4yuSR9_r0cSSIZyNREqJDA4cLqAnkhEVupw6LSvG926oQp7dYtuyEUYmq1OgIQ2jW6CYFPSwYecTN-nxVdBnUgXnwOWiKK52-O9tQ46U167LOt78vVa4hJm5cKiJIjQtV1LzHuT4rC8nTAghUxKZLustFRKt72s3Hx7w5D2XuIkyig152nzWoGGJyoRC-kwS9y5-MfB5a1OMAdriTZIAjfbrYcBNZA9X41poZIdCrhwRgvxP_I3HC4Y8jgFHohN4wb0PwFhmrKFC5r1OaqqLpA4HMyCkywNPa9FgXhzGx0P8Fzy6pzJztmzeSPoBiZmCqxs3P0PUfi07IesCcXqE_-AizVUpkG6iZyP7V_INEd15niD9aY1g_kg8evb0qAf8UN57CBNs7A8naLPJ4P0n3xtazkLL84CGIAfQx6ivZv1HclHs92ogo3Ayv_eE4RNcUdq0vxO0761-wcO6RFfyNkKK5MnzWsb-a8tGMZUmgk04klLZ9XUbjNEPnRFWVQSapDfWkzsRNfTMHC-ajJ7QOlNaS13EWRou3QvhyvNcwxhw-oQ4pMpvkMa0liBfp1l8YI6X8r0HI-hyunfF7seHwbkkqry4FlojP8lIHuyqlcaqvYuZec5kU12Gm7Mync-4X1bXBjU0VE3qvZBJqHMHr_zs00Z_qXSXZkeyL1CqvXgzXLpomSYiSVqzh7Qzbo378uafjH_uCJBE_g_3zDgy2VCg4tB2tgbVCZ7GqGsCctLr70KQhy0i2StHUYUtTTug1z4CSdC852IRx_8VlREuDSBVlJas0vMLKJo2asUrcZwb62WEwH_3ru0nJLjJ8m_CK97mMmXnxkvQL_QZZGGReJRxI7rNZakI8g6xE3zKw4xSzgSZ0piFlv30-_IxuDnwOXpKXihv_M8CmJKAWLdslIhSK0eX67BuIcrfoQ0v2cmhVH252GK0Z6NeC3bspLFJzdIvG5a9vM9ZEDaEE7MHx-jaRNIBrjSWM1j7ivTohT6rXXLTVqGDUadsv8Yk1nbzFLxNpvl5gJ6CAUiKNIfxbY5FgIEEfvJOgofStaP8zGyOdpIwxTJv-kDwGeSfzpVEaLmiAlmadbst1mzsbpKHQr1BTRHtH2c9FeehuoSR85-Y5qjVNSRgAR5aXzqLTDATjO7T6uXQutHXP530-COsA0PlMoivnglelE-j9ghFntUPAsfrIr5m5pJGL5tX_xTUlMI5WJkC851RHGHXF_MY2KxnLWmDvSP0S38rucob7Hpq6RUqpZKzocKCmwaVLPaYdYYyBvGSI4T_09M9YJ_ZufaS4Fbab11M_1gFKn1cnsF7igRBULMHrhUkbV9l73jldb-Cbice9DXGUsa61xGbWvrj5a-KXZKfh7jJ09lZe5I8SaWozRM22MQDaUoG7I9QzesfZ_-fYb9QUJiSe256JK7jiBKddNc4fQF5G-QqY3DJ7bXLbH0C73R4t9qQI20VNRruPLwo2q9fccHgbP2VbHzB5Vv2MzeQoX2gM2lpz_7Wje7D1zPVmiKTXhHAI9tDZSrSnJo8EDCwCn1I1JE-7ijdy8GKRS_IVKnz'
decrypt(answer,)
```
<a id='2'></a>
# Bokeh
```
YouTubeVideo('oLU5eIO7b84')
```
## Task 5
Create an animated plot showing the data on CO$_2$ release from the [ICOS data portal](https://www.icos-cp.eu/) in Hohenpeißenberg (HPB), Hyltemossa (HTM), and Norunda (NOR).<br>
In the following cells, some steps are already implemented:
1. We read the data sets into `pandas` DataFrames
2. We average the emissions over years, days, and minutes to obtain the average emission throughout a day, hour by hour, for the different months
3. We concatenate the DataFrames
4. We create a static Bokeh plot with a tooltip inspection tool. The stations are represented by circles on a map. The radius of the circle is proportional to the CO$_2$ emission.
Now let's add 2 slider widgets to control the month and the hour, following the approach shown in this [video lecture](https://youtu.be/oLU5eIO7b84).
```
!cat data/ICOS_ATC_L2_L2-2018.1_NOR_32.0_462.CO2 | grep "LATITUDE"
!cat data/ICOS_ATC_L2_L2-2018.1_NOR_32.0_462.CO2 | grep "LONGITUDE"
!cat data/ICOS_ATC_L2_L2-2018.1_HPB_93.0_382.CO2 | grep "LATITUDE"
!cat data/ICOS_ATC_L2_L2-2018.1_HPB_93.0_382.CO2 | grep "LONGITUDE"
!cat data/ICOS_ATC_L2_L2-2018.1_HTM_70.0_461.CO2 | grep "LATITUDE"
!cat data/ICOS_ATC_L2_L2-2018.1_HTM_70.0_461.CO2 | grep "LONGITUDE"
data_hpb = pd.read_csv('data/ICOS_ATC_L2_L2-2018.1_HPB_93.0_382.CO2',header=0,skiprows=38,sep=';')
data_hpb[data_hpb.co2<0] = np.nan
data_hpb.head()
data_hpb = data_hpb[['Year','Month','Day','Hour','Minute','co2','SamplingHeight']]
data_hpb.set_index(['Year','Month','Day','Hour','Minute'],inplace=True)
data_hpb = data_hpb.groupby(['Month','Hour'],level=(1,3)).mean()
data_hpb.reset_index(inplace=True)
data_hpb['co2'] /= 10
data_hpb['lat'] = 47.8011
data_hpb['lon'] = 11.0246
data_hpb['station'] = 'Hohenpeißenberg'
data_hpb.head()
data_nor = pd.read_csv('data/ICOS_ATC_L2_L2-2018.1_NOR_32.0_462.CO2',header=0,skiprows=38,sep=';')
data_nor[data_nor.co2<0] = np.nan
data_nor = data_nor[['Year','Month','Day','Hour','Minute','co2','SamplingHeight']]
data_nor.set_index(['Year','Month','Day','Hour','Minute'],inplace=True)
data_nor = data_nor.groupby(['Month','Hour'],level=(1,3)).mean()
data_nor.reset_index(inplace=True)
data_nor['co2'] /= 10
data_nor['lat'] = 60.0864
data_nor['lon'] = 17.4794
data_nor['station'] = 'Norunda'
data_nor.head()
data_htm = pd.read_csv('data/ICOS_ATC_L2_L2-2018.1_HTM_70.0_461.CO2',header=0,skiprows=38,sep=';')
data_htm[data_htm.co2<0] = np.nan
data_htm = data_htm[['Year','Month','Day','Hour','Minute','co2','SamplingHeight']]
data_htm.set_index(['Year','Month','Day','Hour','Minute'],inplace=True)
data_htm = data_htm.groupby(['Month','Hour'],level=(1,3)).mean()
data_htm.reset_index(inplace=True)
data_htm['co2'] /= 10
data_htm['lat'] = 56.0976
data_htm['lon'] = 13.4189
data_htm['station'] = 'Hyltemossa'
data_htm.head()
df = pd.concat([data_nor,data_hpb,data_htm])
df.set_index(['Month','Hour'],inplace=True)
df.sort_index(inplace=True)
df.head()
def make_data(df,month,hour):
data = df.loc[month,hour].copy()
return data.reset_index()
make_data(df,1,4)
import geopandas as gpd
from bokeh.plotting import figure
from bokeh.io import show, output_notebook
from bokeh.palettes import Category10
from bokeh.models import GeoJSONDataSource, ColumnDataSource, HoverTool
from bokeh.transform import factor_cmap
from bokeh.models.widgets import Slider
from bokeh.layouts import column, row, WidgetBox
source = ColumnDataSource( data = make_data(df,1,4) )
location = { 'HPB': {'lat':47.8011,'lon':11.0246},
'NOR': {'lat':60.0864,'lon':17.4794} }
europe = gpd.read_file(gpd.datasets.get_path('naturalearth_lowres'))
geo_source = GeoJSONDataSource(geojson=europe.to_json())
p = figure(title="CO\u2082 Release in Europe", x_range=(-30,60), y_range=(30,85),
tools='pan,reset')
p.patches('xs', 'ys', fill_alpha=0,
line_color=Category10[10][7], line_width=1.5, source=geo_source)
glyph = p.scatter('lon','lat',
source = source, size = 'co2', fill_alpha = .5, hover_fill_alpha = 1.0,
marker = 'circle',
legend_label = 'station',
color = factor_cmap( 'station', 'Category10_5', source.to_df().station.unique() ),
hover_line_color = factor_cmap( 'station', 'Category10_5', source.to_df().station.unique() ),
hover_fill_color = factor_cmap( 'station', 'Category10_5', source.to_df().station.unique() ) )
glyph_hover = HoverTool(renderers=[glyph],
tooltips=[('Longitude','@lon E'), ('Latitude','@lat N'), ('Sation','@station'),
('Month','@Month'), ('Hour','@Hour'), ('CO2','@co2'),
('Sampling Height','@SamplingHeight')])
p.add_tools(glyph_hover)
p.title.align = 'center'
p.legend.location = 'top_left'
output_notebook()
show(p)
answer = 'gAAAAABcBvoKW3LiJnuzl-rUkXnWXDyKdCa1TtA-6Zp5eHeGiSp42LcPzqyChSRfFel5HeESU-z8_OLoZp2MFw8VAOWoCNSkIacf758tpY_HOggr6j8rYC9e1pTfzus0HXEsdOBReP58YQ8eLw63dR-la2nC4TcRXezGAQ2IKOGav8WM11nMlkhNXPV126xh4jsBT0nRxmNUaQejldIJC_q4WPDsWuQAHl3b5e-sraepPnmkONRTq4VqlSrFpd62CBsmCo8KbOuCRXMjBHv_xCmt34htey7DQ65WnNauZqOMLpPze_6d58jjA3hzFth9oqEspzsP5J_VrK55vqdB8LZM1dnu7g97QEVWWe2XsKVpXEa2TXPUqACN4AKlhfSy-zfS78v5qWzgFZLscMHa2ifgFZGgPXoWVRhcxLq5AL644559F8g1jCAGnXNaYmMXoUZ6xZCdMFliYiFuL7V8dhEtaHawX441rIEtoIb2XS0ZK0lPTav5eIHhceQLuTm4lCmppAXvkLkrJnkBzlom3bYPZb00whL4nyMN39c5el4TNo5Sf-v-3GZl4uSWubhh5pPE3opFA8NC444CBvU9VBn2ZklTyeziByeKhdCOYHlKXECXWKRqAEK0ScBuZOTFNm83Kj18XcXmEt5-v2vjMGfM-rVSAcPyBNU3gOKhnFYNbtNgwL69F3ehIBQlkH5rGkgvilbmlgcJx0hPS3aVyFybxN9xGUlFcSs8DmDdGF5hxAs4vZWNgy_PjAXNeW-hdMUGY3wIU3JPC96DNIBTyMQV_BtzGVgpNm4kJ8M7ljNq563pVnf0q3vXwPSYnFUS6-aL0GarW47U2MQzCg8VerMKMR5-WTfNcz0euna78dg7hb_ZqeJgmESiL0gqFQdHNMriGMNJeLfd-sMlgI1WBLV9TSeFqbbOoar0IjzzzmoRmla8WFrSULbK0zMGz_DdiQOLQCWoEBRVDkwcVPmobdYaS4xy2DE2w_rBQOkCM68k-sNqg04_hZ652Pq7KnOAG1BwL3D4Pz73ZaE-uDL1vbnr6auI7dsxcPgJX3_b0pIqIjwcPwd0iaoU2SdEVU2OpGBnY6KsSl5MrwX7P-dYVhWwJ4FfYP59l7HfTJrw7t03jkA399adGWF70wA7v-2bHHzCTf66wgCnFmQtLL8ukmrkImiAzCMA3af47hq8GPKbFK74UsnwyjabLv26N0WZcuc2P3Tib-qMtc0ebuHTXWFtmio0RSDCjKVOehakMwoeIKfIAWZAoYfAg5NFQWa8Xw8ybsHvwCrDwt4j7qb-hdoyIQ2cMEFfXXx4ZcL-GRJy6jlwyP0C90JjvWv8ono8ffdhuuSD_ylCHCt0c58UxibJXUkvHzgA40rlWRH7Pmh6iw1zeG5lkxkHZFt6AaKpxNhtu0WDsHLUxqMoa1PrjJjjP1VwjVRKpeaIWMui9j-Wg3gQB0Bt9b7iqvgPW8R5OLAXOm_84E8ozBFn1DWdJaMyutqgh3SsHiL_bOlWeJfm9-G_YeVaXgjsj4sSAf01VzBxIe_bdgY44OUiobZ5aoF0O4Ya7AVMhXPeJHi8f4GVeacI7KJAiGFH3Vmm_X-cgM8axMPthxtC7vBYgw_a4TOikWYewPh5uU9f4mjs4rKS9xC0H8tjp5ICrDhdYFWqviVnnuunSWR9bmbRKYVIsmIJHYZLZpci6bOz8jKVyxjp4YgD9-3IYe5EPy_6uF9XbNZesyeJkeY_q5P1qt2iigQFi8hD88lt4nvFjV5yXdGwfeyLIm0YaS_SE9mAu2NnQgejRKrcsmahQAvYUy64DD38ktJevblcNRy90xMd6GXHr69ZPVz65QsZAroqHy_GXjjQhJpRS-aajoL6LcolcFvrvzq46Hv_JIscJscnPHkqwWbKgS1QLD4HdUC_VVK_9GmfbjgI8Y05hOEPF2c_0Nhm1-F3dkTBrIXLgHnd88FHHU9feobDBNerd9_sqQctTE7d2FHxrcvNZRVlhHZiFyFuer1cz29keRSJm0Gp035WFhieYECzaM-_7NMbRkUdwAf3yxzUjNy1WKyehC-ZawcXwsyTOVvMugfHZ4olSImjGvVKsd7Ho5oTip2fZEv9ic2GdGFxnzP3bhpBhp4fLj8QeDEKYkeGNmDumo-yPJo_NVr30-rNUdeNDIQtOKB8rvv-HKFsz8t_nvctKEMOpe4Iq0nI1QadWCQtkyUAG7kIfsby6jtgANtyLjddZ5HZAvpbpros4jr3fanKxr4Qzr8kDzOw09oid1it8SetOWnBM9YzEbHf2-Rm1ZFhkAfjaf0_wSGBeMXinAr66Gk11pTmCOHqOFzSZRcE6-bGkSBJXe4gf17EY9Ot8S2CjDNqSO9s1eDSh5Xrw_SbPq9OCYDk58CY1RihwEr6O6u_tdpB-i-PEaPUc1te-Sk9dOSLrRE97K-AHjjW1Zxb8lEL4k2d9-NQUUg0jACC7eYACWOgoUwnoq3QrYR-lbhoM0EZSLIEjuP5fhPQWMpVUQ3iaXTtl-j1cfXjDT8_bPqE4oD2IdO_MIaiAV0NsBjM9s5MR_w2ssJ9vexRnaQ92k7pL4OTjMQ4Cgu0IriH7roFIil1UQAaH-gZFWnpKYJrD6zIAQrdgZ_Cb2LNSwgcybA7SFfzFbAOitp5foBGlHr1UtGdlN30SmKQ105mLeBHE3zFdqP4SazKCL1Gyo6XNGyS_n_Qj-yYl66GLdegIBNtQMY6apVCpAHoSCzAXzEHw2ARW3LIZMjjeJn5UFlmAyuW7elMGV-rKSdxlszOXAryXBgl3GIbiYXAnZ_LL04EqXydNaDdtNL4r6_cQnMw3ykTLC6HW997onodrnWuVRswO6KR_0kJ97KUwPULxs1Q8ANcMoV4MsLqDsDPJBi1txw0PJKfzONAun4Wbbj5w-NDn3yckR7cakFieDcWqblAMdAoyFw-Cx7JZyuqWXaQQh6PzD3k2FOIQoH8qUegVabq7iW1bZt_xgv6HxGixf_u_RvQ3ViL8nsL_SiPGBTqC_sjVKlXkO9JJZd3DBOWgt-tG7ft99QacBpfz3n6L6j7c759G7uIspqCRD2PvpXcykk9MRPMzBPe7uzFkir4z6I61wnwFaxurKo6Sps_DHGAQZuYC3PemsWh9yNjVruTV0ZNTwlG-PgEbGNfI_eiIt7K0Mw6BjCY7co5SyPPP6KSU64dLPjjaO--0F1Uc-7mzdjy5PyfalG4FnPymXnhLg6_vHjtLgtw1Vh_jL2Bwda6hc_Z-zU2ZXvsAe1dkvgEdhJ5W0MSzHPi3qo8YLgpVw5UAwTWqzI-INJpMqh7CZOkNP31qLI7x1gpV2hWGM-UzW3FGuJdKMPpwEE5ppDlbXgM3zscGiNYTqVlpl58MKoh7OCFFGhxM1aaX4tLIEonRCEH-UapKpqU-mArBIFM0AHy5zLeHHal72jBpQC4OlTf4F8cY_sFCf52SjcayIAR4wlvI6yQKDOSTWS1nEl5JrAuWzR4RIBVtuT_ohT_Gw-8i1CzocFLi2e_CWoHWBvrkAcOI9RiyiidmhyaJrQdfmSdpEFC-Kv5tlTuKxypxP5Rwi4vCh-rvYAOpu_mucOhzKEshoTXunlfarmsJiFk5neSEuZ20LmsVi_Nt7S9HNwgM5RA-k5xxxm-sLzMLCFWwAuHU9vZjhH5s44So3xyGhssI9uyGCJC4QA0M41ihZUOmcQKeEighxgXvVmY'
decrypt(answer,)
```
| github_jupyter |
```
#export
from typing import List, Iterator, Any, NewType, TypeVar, Generic
import k1lib.cli as cli; from numbers import Number
import k1lib, itertools, copy, torch; import numpy as np
__all__ = ["BaseCli", "Table", "T", "fastF",
"serial", "oneToMany", "manyToMany", "mtmS"]
#export
settings = k1lib.Settings()
atomic = k1lib.Settings()
settings.add("atomic", atomic, "classes/types that are considered atomic and specified cli tools should never try to iterate over them")
settings.add("defaultDelim", "\t", "default delimiter used in-between columns when creating tables. Defaulted to tab character.")
settings.add("defaultIndent", " ", "default indent used for displaying nested structures")
settings.add("strict", False, "turning it on can help you debug stuff, but could also be a pain to work with")
settings.add("inf", float("inf"), "infinity definition for many clis. Here because you might want to temporarily not loop things infinitely")
k1lib.settings.add("cli", settings, "from k1lib.cli module")
#export
def patchDefaultDelim(st:str):
"""
:param s:
- if not None, returns self
- else returns the default delimiter in :attr:`~k1lib.settings`"""
return settings.defaultDelim if st is None else st
def patchDefaultIndent(st:str):
"""
:param s:
- if not None, returns self
- else returns the default indent character in :attr:`~k1lib.settings`"""
return settings.defaultIndent if st is None else st
#export
T = TypeVar("T")
"""Generic type variable"""
class _MetaType(type):
def __getitem__(self, generic):
d = {"__args__": generic, "_n": self._n, "__doc__": self.__doc__}
return _MetaType(self._n, (), d)
def __repr__(self):
def main(self):
def trueName(o):
if isinstance(o, _MetaType): return main(o)
try: return o.__name__
except: return f"{o}"
if hasattr(self, "__args__"):
if isinstance(self.__args__, tuple):
return f"{self._n}[{', '.join([trueName(e) for e in self.__args__])}]"
else: return f"{self._n}[{trueName(self.__args__)}]"
return self._n
return main(self)
def newTypeHint(name, docs=""):
"""Creates a new type hint that can be sliced and yet still looks fine
in sphinx. Crudely written by my poorly understood idea of Python's
metaclasses. Seriously, this shit is bonkers, read over it https://stackoverflow.com/questions/100003/what-are-metaclasses-in-python
Example::
Table = newTypeHint("Table", "some docs")
Table[int] # prints out as "Table[int]", and sphinx fell for it too
Table[Table[str], float] # prints out as "Table[Table[str], float]"
"""
return _MetaType(name, (), {"_n": name, "__doc__": docs})
#Table = newTypeHint("Table", """Essentially just Iterator[List[T]]. This class is just here so that I can generate the docs with nicely formatted types like "Table[str]".""")
#Table = NewType("Table", List)
class Table(Generic[T]):
"""Essentially just Iterator[List[T]]. This class is just here so that I can generate the docs with nicely formatted types like "Table[str]"."""
pass
Table._name = "Table"
#Table.__module__ = "cli"
class Row(list):
"""Not really used currently. Just here for potential future feature"""
pass
#export
class BaseCli:
"""A base class for all the cli stuff. You can definitely create new cli tools that
have the same feel without extending from this class, but advanced stream operations
(like ``+``, ``&``, ``.all()``, ``|``) won't work.
At the moment, you don't have to call super().__init__() and super().__ror__(),
as __init__'s only job right now is to solidify any :class:`~k1lib.cli.modifier.op`
passed to it, and __ror__ does nothing."""
def __init__(self, fs=[]):
"""Not expected to be instantiated by the end user.
:param fs: if functions inside here is actually a :class:`~k1lib.cli.modifier.op`,
then solidifies it (make it not absorb __call__ anymore)"""
[f.op_solidify() for f in fs if isinstance(f, cli.op)]
def __and__(self, cli:"BaseCli") -> "oneToMany":
"""Duplicates input stream to multiple joined clis.
Example::
# returns [[5], [0, 1, 2, 3, 4]]
range(5) | (shape() & iden()) | deref()
Kinda like :class:`~k1lib.cli.modifier.apply`. There're just multiple ways of doing
this. This I think, is more intuitive, and :class:`~k1lib.cli.modifier.apply` is more
for lambdas and columns mode. Performances are pretty much identical."""
if isinstance(self, oneToMany): return self._after(cli)
if isinstance(cli, oneToMany): return cli._before(self)
return oneToMany(self, cli)
def __add__(self, cli:"BaseCli") -> "mtmS":
"""Parallel pass multiple streams to multiple clis."""
if isinstance(self, mtmS): return self._after(cli)
if isinstance(cli, mtmS): return cli._before(self)
return mtmS(self, cli)
def all(self, n:int=1) -> "BaseCli":
"""Applies this cli to all incoming streams.
:param n: how many times should I chain ``.all()``?"""
s = self
for i in range(n): s = manyToMany(s)
return s
def __or__(self, cli) -> "serial":
"""Joins clis end-to-end"""
if isinstance(self, serial): return self._after(cli)
if isinstance(cli, serial): return cli._before(self)
return serial(self, cli)
def __ror__(self, it): return NotImplemented
def f(self) -> Table[Table[int]]:
"""Creates a normal function :math:`f(x)` which is equivalent to
``x | self``."""
return lambda it: self.__ror__(it)
def __lt__(self, it):
"""Default backup join symbol `>`, in case `it` implements __ror__()"""
return self.__ror__(it)
def __call__(self, it, *args):
"""Another way to do ``it | cli``. If multiple arguments are fed, then the
argument list is passed to cli instead of just the first element. Example::
@applyS
def f(it):
return it
f(2) # returns 2
f(2, 3) # returns [2, 3]"""
if len(args) == 0: return self.__ror__(it)
else: return self.__ror__([it, *args])
#export
def fastF(c):
"""Tries to figure out what's going on, is it a normal function, or an applyS,
or a BaseCli, etc., and return a really fast function for execution. Example::
# both returns 16, fastF returns "lambda x: x**2", so it's really fast
fastF(op()**2)(4)
fastF(applyS(lambda x: x**2))(4)"""
if isinstance(c, cli.op): return c.ab_fastF()
if isinstance(c, cli.applyS): return fastF(c.f)
if isinstance(c, BaseCli): return c.__ror__
return c
assert fastF(cli.op() ** 2)(4) == 16
#export
class serial(BaseCli):
def __init__(self, *clis:List[BaseCli]):
"""Merges clis into 1, feeding end to end. Used in chaining clis
together without a prime iterator. Meaning, without this, stuff like this
fails to run::
[1, 2] | a() | b() # runs
c = a() | b(); [1, 2] | c # doesn't run if this class doesn't exist"""
super().__init__(fs=clis); self.clis = list(clis); self._cache()
def _cache(self):
self._hasTrace = any(isinstance(c, cli.trace) for c in self.clis)
self._cliCs = [fastF(c) for c in self.clis]; return self
def __ror__(self, it:Iterator[Any]) -> Iterator[Any]:
if self._hasTrace: # slower, but tracable
for cli in self.clis: it = it | cli
else: # faster, but not tracable
for cli in self._cliCs: it = cli(it)
return it
def _before(self, c): self.clis = [c] + self.clis; return self._cache()
def _after(self, c): self.clis = self.clis + [c]; return self._cache()
@cli.applyS
def f(it): return it
assert f(2) == 2
assert f(2, 3) == [2, 3]
assert range(5) | (cli.shape() & cli.iden()) | cli.deref() == [[5], [0, 1, 2, 3, 4]]
assert isinstance([np.random.randn(2, 3, 4)] | (cli.item() | serial(cli.shape())), tuple)
# examples in cli rst docs
assert cli.shape()(np.random.randn(2, 3, 5)) == (2, 3, 5)
assert [np.random.randn(2, 3, 5)] | (cli.item() | cli.shape()) == (2, 3, 5)
#export
atomic.add("baseAnd", (Number, np.number, str, dict, bool, bytes, list, tuple, torch.Tensor), "used by BaseCli.__and__")
def _iterable(it):
try: iter(it); return True
except: return False
class oneToMany(BaseCli):
def __init__(self, *clis:List[BaseCli]):
"""Duplicates 1 stream into multiple streams, each for a cli in the
list. Used in the "a & b" joining operator. See also: :meth:`BaseCli.__and__`"""
super().__init__(fs=clis); self.clis = clis; self._cache()
def __ror__(self, it:Iterator[Any]) -> Iterator[Iterator[Any]]:
if isinstance(it, atomic.baseAnd) or not _iterable(it):
for cli in self._cliCs: yield cli(it)
else:
its = itertools.tee(it, len(self.clis))
for cli, it in zip(self._cliCs, its): yield cli(it)
def _cache(self): self._cliCs = [fastF(c) for c in self.clis]; return self
def _before(self, c): self.clis = [c] + self.clis; return self._cache()
def _after(self, c): self.clis = self.clis + [c]; return self._cache()
#export
class manyToMany(BaseCli):
def __init__(self, cli:BaseCli):
"""Applies multiple streams to a single cli. Used in the :meth:`BaseCli.all`
operator."""
super().__init__(fs=[cli]); self.cli = cli; self._cliC = fastF(cli)
def __ror__(self, it:Iterator[Iterator[Any]]) -> Iterator[Iterator[Any]]:
f = self._cliC; return (f(s) for s in it)
#export
class mtmS(BaseCli):
def __init__(self, *clis:List[BaseCli]):
"""Applies multiple streams to multiple clis independently. Used in
the "a + b" joining operator. See also: :meth:`BaseCli.__add__`.
Weird name is actually a shorthand for "many to many specific"."""
super().__init__(fs=clis); self.clis = list(clis); self._cache()
def _cache(self): self._cliCs = [fastF(c) for c in self.clis]; return self
def _before(self, c): self.clis = [c] + self.clis; return self._cache()
def _after(self, c): self.clis = self.clis + [c]; return self._cache()
def __ror__(self, its:Iterator[Any]) -> Iterator[Any]:
for cli, it in zip(self._cliCs, its): yield cli(it)
@staticmethod
def f(f, i:int, n:int=100):
"""Convenience method, so
that this::
mtmS(iden(), op()**2, iden(), iden(), iden())
# also the same as this btw:
(iden() + op()**2 + iden() + iden() + iden())
is the same as this::
mtmS.single(op()**2, 1, 5)
Example::
# returns [5, 36, 7, 8, 9]
range(5, 10) | mtmS.single(op()**2, 1, 5) | deref()
:param i: where should I put the function?
:param n: how many clis in total? Defaulted to 100"""
return mtmS(*([cli.iden()]*i + [f] + [cli.iden()]*(n-i-1)))
assert len((cli.iden() + cli.op()**2 + cli.iden() + cli.iden() + cli.iden()).clis) == 5
assert len(mtmS(cli.iden(), cli.op()**2, cli.iden(), cli.iden(), cli.iden()).clis) == 5
assert range(5, 10) | mtmS.f(cli.op()**2, 1, 5) | cli.deref() == [5, 36, 7, 8, 9]
!../../export.py cli/init
```
| github_jupyter |
###### Text provided under a Creative Commons Attribution license, CC-BY. All code is made available under the FSF-approved MIT license. (c) Daniel Koehn based on Jupyter notebooks by Marc Spiegelman [Dynamical Systems APMA 4101](https://github.com/mspieg/dynamical-systems) and Kyle Mandli from his course [Introduction to numerical methods](https://github.com/mandli/intro-numerical-methods), notebook style sheet by L.A. Barba, N.C. Clementi [Engineering Computations](https://github.com/engineersCode)
```
# Execute this cell to load the notebook's style sheet, then ignore it
from IPython.core.display import HTML
css_file = '../style/custom.css'
HTML(open(css_file, "r").read())
```
# Simplified Convection Problem: The Lorenz Equations
The Lorenz Equations are a 3-D dynamical system that is a simplified model of Rayleigh-Benard thermal convection. They are derived and described in detail in Edward Lorenz' 1963 paper [Deterministic Nonperiodic Flow](http://journals.ametsoc.org/doi/pdf/10.1175/1520-0469%281963%29020%3C0130%3ADNF%3E2.0.CO%3B2) in the Journal of Atmospheric Science.
Here we will just sketch out the key points of the derivation. A more complete derivation can be found [here](https://www.math.uni-hamburg.de/home/lauterbach/scripts/seminar03/prill.pdf)
The key idea is that the Lorenz Equations result from a severely truncated spectral approximation to the 2-D equations for incompressible thermal convection in stream-function/vorticity form. These equations govern the flow of a buouyant incompressible fluid with a temperature dependent density in a layer of depth $h$, that is heated from below and cooled from the top.
## Governing Equations
The full coupled set of scaled PDE's describe the coupling of incompressible 2D Navier Stokes flow with an advection-diffusion equation for temperature, and can be written in dimensionless form as,
$$
\frac{1}{\mathrm{Pr}}\left[ \frac{\partial \omega}{\partial t} + \vec{v}\cdot\nabla\omega\right] = \nabla^2\omega + \mathrm{Ra}\frac{\partial T}{\partial x}
$$
$$
\nabla^2 \psi = -\omega
$$
$$
\frac{\partial T}{\partial t} + \vec{v}\cdot\nabla T = \nabla^2 T
$$
where
$$
\vec{v}=(u,0,w) = \nabla\times\psi\hat{j}=(-\frac{\partial\psi}{\partial z}, 0, \frac{\partial\psi}{\partial x})
$$
is the fluid velocity field (which in this form is exactly incompressible with $\nabla\cdot\vec{v}=0$). $\psi$ is the "Streamfunction" whose contours are tangent to the fluid trajectories at all times. The vorticity,
$$
\omega = \bf{\vec{\omega} \cdot \hat{j}} = (\nabla\times\vec{v}) \cdot \hat{j}
$$
measures the local rate of rotation of the fluid, and is driven by horizontal variations in temperature (actually density). $\hat{j}=(0,1,0)^T$ denotes the unit vector in y-direction.
Boundary conditions for temperature are $T=1$ on the bottom of the layer and $T=0$ on the top. In the absence of any fluid motion ($\omega=\vec{v}=0$), the temperature field is just a steady conductive ramp with
$$
T = 1 - z
$$
Thus we can also solve for the perturbation $\theta(x,z,t)$ away from this steady state by substituting
$$
T = 1 - z + \theta(x,z,t)
$$
into the energy equation to solve for the perturbed temperature using
$$
\frac{\partial \theta}{\partial t} + \vec{v}\cdot\nabla \theta = \nabla^2\theta + w
$$
### Parameters
In dimensionless form, these equations have two important dimensionless numbers that control the structure and behavior of the convection.
**The Prandtl Number**
The first is the "Prandtl Number", $\mathrm{Pr} = \frac{\nu}{\kappa}$ which is the ratio of the fluid viscosity $\nu$ to the thermal diffusivitiy $\kappa$. Since both vorticity and temperature both obey advection diffusion equations (and viscosity acts to diffuse momentum/vorticity), the Prandtl number is a measure of whether momemtum or energy is more dissipative.
**The Rayleigh Number**
The second key parameter is the Rayleigh number
$$
\mathrm{Ra} = \frac{g\alpha(T_1 - T_0)h^3}{\nu\kappa}
$$
which measures the balance of forces that drive convection (i.e. gravity, or temperature differences), to those that damp convection such as viscosity and thermal diffusivity. Systems with large Rayleigh numbers are prone to vigorous convection. However, it was shown by Rayleigh, that there is a critical value of the Rayleigh Number $\mathrm{Ra}_c$ below which there is no convection. This value depends on the size of the convection cell and boundary conditions for stress on the fluid, however, for the simplest case of a layer with no-slip top and bottom boundary conditions and cell with aspect ratio $a=h/L$ (with $h$ the layer depth and $L$ the width of the convection cell), then the critical Ra number is
$$
\mathrm{Ra}_c = \pi^4 (1 + a^2)^3/a^2
$$
which has a minimum value for $a_{min}^2=1/2$.
```
# load libraries
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
from pylab import rcParams
from pylab import rcParams
from matplotlib import rc
# define font size
FSize = 18
font = {'color': 'black',
'weight': 'normal',
'size': FSize}
mpl.rc('xtick', labelsize=FSize)
mpl.rc('ytick', labelsize=FSize)
# Show Ra vs a
a = np.linspace(0.01,2.)
Rc = np.pi**4*(1. + a**2)**3/a**2
plt.figure()
plt.semilogy(a,Rc)
amin = np.sqrt(1./2.)
Rcmin = np.pi**4*(1. + amin**2)**3/amin**2
plt.semilogy(amin,Rcmin,'ro')
plt.xlabel('a', fontdict=font)
plt.ylabel('Ra$_c$', fontdict=font)
plt.title('Critical Rayleigh Number', fontdict=font)
plt.show()
```
## Spectral decomposition
Next, we expand the streamfunction and temperature fields in terms of a highly truncated Fourier Series where the streamfunction contains one cellular mode
$$
\psi(x,z,t) = X(t)\sin(a\pi x)\sin(\pi z)
$$
and temperature has two modes
$$
\theta(x,z,t) = Y(t)\cos(a\pi x)\sin(\pi z) - Z(t)\sin(2\pi z)
$$
Here, $X(t)$, $Y(t)$ and $Z(t)$ are the time dependent amplitudes of each mode. Defining
\begin{equation}
\begin{split}
\phi(x,z) &= \sin(a\pi x)\sin(\pi z)\\
\theta_0(x,z) &= \cos(a\pi x)\sin(\pi z)\\
\theta_1(z) &= -\sin(2\pi z)
\end{split}
\notag
\end{equation}
streamfunction and temperature fields simplify to:
\begin{equation}
\begin{split}
\psi(x,z,t) &= X(t) \phi(x,z)\\
\theta(x,z,t) &= Y(t) \theta_0(x,z) + Z(t) \theta_1(z)\\
\end{split}
\notag
\end{equation}
Using $a = a_{min} = \sqrt{0.5}$ for the minimum critical Rayleigh number $Ra_c^{min}$, the spatial components of each mode looks like
```
a = np.sqrt(0.5)
x0 = np.linspace(0,1./a)
z0 = np.linspace(0.,1.)
x,z = np.meshgrid(x0,z0)
psi = np.sin(a*np.pi*x)*np.sin(np.pi*z)
theta0 = np.cos(a*np.pi*x)*np.sin(np.pi*z)
theta1 = -np.sin(2.*np.pi*z)
plt.figure()
plt.figure(figsize=(10,8))
plt.subplot(2,2,1)
plt.contourf(x,z,psi)
plt.title(r'$\Phi = sin(a \pi x)sin(\pi z)$', fontdict=font)
plt.gca().set_aspect('equal')
plt.subplot(2,2,3)
plt.contourf(x,z,theta0)
plt.title(r'$\theta_0 = cos(a \pi x)sin(\pi z)$', fontdict=font)
plt.gca().set_aspect('equal')
plt.subplot(2,2,4)
plt.contourf(x,z,theta1)
plt.title(r'$\theta_1 = -sin(2 \pi z)$', fontdict=font)
plt.gca().set_aspect('equal')
plt.tight_layout()
plt.show()
```
For our initial conditions $X(0) = 2$, $Y(0) = 3$, $Z(0) = 4$, the streamfunction $\psi(x,z,t)$ and temperature field $\theta(x,z,t)$
\begin{equation}
\begin{split}
\psi(x,z,t) &= X(t)\sin(a\pi x)\sin(\pi z)\\
\theta(x,z,t) & = Y(t)\cos(a\pi x)\sin(\pi z) - Z(t)\sin(2\pi z)\\
\end{split}
\notag
\end{equation}
become
\begin{equation}
\begin{split}
\psi(x,z,0) &= 2\sin(a\pi x)\sin(\pi z)\\
\theta(x,z,0) &= 3\cos(a\pi x)\sin(\pi z) - 4\sin(2\pi z)\\
\end{split}
\notag
\end{equation}
```
# Define figure size
plt.figure()
plt.figure(figsize=(20,20))
# Initial Streamfunction psi and velocity field
plt.subplot(1,2,1)
plt.contourf(x,z,2.*psi,cmap='viridis_r')
# Velocity field
U = - 2. * np.pi * np.sin(a*np.pi*x) * np.cos(np.pi*z)
V = 2. * a * np.pi * np.cos(a*np.pi*x) * np.sin(np.pi*z)
plt.quiver(x,z,U,V)
plt.gca().set_aspect('equal')
plt.title('Streamfunction $\psi$', fontdict=font)
# Initial temperature field
plt.subplot(1,2,2)
plt.contourf(x,z,3*theta0 + 4*theta1,cmap='magma')
plt.gca().set_aspect('equal')
plt.title(r'Temperature $\theta$', fontdict=font)
plt.tight_layout()
plt.show()
```
## Lorenz equations
Inserting the spectral decomposition solutions into the momentum and energy conservation equations lead to the **Lorenz equations**
\begin{equation}
\begin{split}
\frac{\partial X}{\partial t} &= \sigma( Y - X)\\
\frac{\partial Y}{\partial t} &= rX - Y - XZ \\
\frac{\partial Z}{\partial t} &= XY -b Z
\end{split}
\notag
\end{equation}
where $\sigma$ denotes the "Prandtl number", $r = \mathrm{Ra}/\mathrm{Ra}_c$ is a scaled "Rayleigh number" and $b$ is a parameter that is related to the the aspect ratio of a convecting cell in the original derivation.
## What we learned:
- The basic concept of the Lorenz equations to approximate the governing equations for thermal convection, described by the incompressible 2D Navier Stokes flow with an advection-diffusion equation for temperature.
| github_jupyter |
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
import tensorflow as tf
print(tf.__version__)
# EXPECTED OUTPUT
# 2.0.0-beta1 (or later)
#### salimt
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
from tensorflow import keras
def plot_series(time, series, format="-", start=0, end=None):
plt.plot(time[start:end], series[start:end], format)
plt.xlabel("Time")
plt.ylabel("Value")
plt.grid(True)
def trend(time, slope=0):
return slope * time
def seasonal_pattern(season_time):
"""Just an arbitrary pattern, you can change it if you wish"""
return np.where(season_time < 0.1,
np.cos(season_time * 7 * np.pi),
1 / np.exp(5 * season_time))
def seasonality(time, period, amplitude=1, phase=0):
"""Repeats the same pattern at each period"""
season_time = ((time + phase) % period) / period
return amplitude * seasonal_pattern(season_time)
def noise(time, noise_level=1, seed=None):
rnd = np.random.RandomState(seed)
return rnd.randn(len(time)) * noise_level
time = np.arange(4 * 365 + 1, dtype="float32")
baseline = 10
series = trend(time, 0.1)
baseline = 10
amplitude = 40
slope = 0.01
noise_level = 2
# Create the series
series = baseline + trend(time, slope) + seasonality(time, period=365, amplitude=amplitude)
# Update with noise
series += noise(time, noise_level, seed=42)
plt.figure(figsize=(10, 6))
plot_series(time, series)
plt.show()
# EXPECTED OUTPUT
# Chart as in the screencast. First should have 5 distinctive 'peaks'
```
Now that we have the time series, let's split it so we can start forecasting
```
split_time = 980
time_train = time[:split_time]
x_train = series[:split_time]
time_valid = time[split_time:]
x_valid = series[split_time:]
plt.figure(figsize=(10, 6))
plot_series(time_train, x_train)
plt.show()
plt.figure(figsize=(10, 6))
plot_series(time_valid, x_valid)
plt.show()
# EXPECTED OUTPUT
# Chart WITH 4 PEAKS between 50 and 65 and 3 troughs between -12 and 0
# Chart with 2 Peaks, first at slightly above 60, last at a little more than that, should also have a single trough at about 0
```
# Naive Forecast
```
naive_forecast = series[split_time - 1:-1]
plt.figure(figsize=(10, 6))
plot_series(time_valid, x_valid)
plot_series(time_valid, naive_forecast)
# Expected output: Chart similar to above, but with forecast overlay
```
Let's zoom in on the start of the validation period:
```
plt.figure(figsize=(10, 6))
plot_series(time_valid, x_valid)
plot_series(time_valid, naive_forecast)
# EXPECTED - Chart with X-Axis from 1100-1250 and Y Axes with series value and projections. Projections should be time stepped 1 unit 'after' series
```
Now let's compute the mean squared error and the mean absolute error between the forecasts and the predictions in the validation period:
```
print(keras.metrics.mean_squared_error(x_valid, naive_forecast).numpy())
print(keras.metrics.mean_absolute_error(x_valid, naive_forecast).numpy())
# Expected Output
# 19.578304
# 2.6011968
```
That's our baseline, now let's try a moving average:
```
def moving_average_forecast(series, window_size):
"""Forecasts the mean of the last few values.
If window_size=1, then this is equivalent to naive forecast"""
forecast = []
for time in range(len(series) - window_size):
forecast.append(series[time:time + window_size].mean())
return np.array(forecast)
moving_avg = moving_average_forecast(series, 30)[split_time - 30:]
plt.figure(figsize=(10, 6))
plot_series(time_valid, x_valid)
plot_series(time_valid, moving_avg)
# EXPECTED OUTPUT
# CHart with time series from 1100->1450+ on X
# Time series plotted
# Moving average plotted over it
print(keras.metrics.mean_squared_error(x_valid, moving_avg).numpy())
print(keras.metrics.mean_absolute_error(x_valid, moving_avg).numpy())
# EXPECTED OUTPUT
# 65.786224
# 4.3040023
diff_series = (series[365:] - series[:-365])
diff_time = time[365:]
plt.figure(figsize=(10, 6))
plot_series(diff_time, diff_series)
plt.show()
# EXPECETED OUTPUT: CHart with diffs
```
Great, the trend and seasonality seem to be gone, so now we can use the moving average:
```
diff_moving_avg = moving_average_forecast(diff_series, 50)[split_time - 365 - 50:]
plt.figure(figsize=(10, 6))
plot_series(time_valid, diff_series[split_time - 365:])
plot_series(time_valid, diff_moving_avg)
plt.show()
# Expected output. Diff chart from 1100->1450 +
# Overlaid with moving average
```
Now let's bring back the trend and seasonality by adding the past values from t – 365:
```
diff_moving_avg_plus_past = series[split_time - 365:-365] + diff_moving_avg
plt.figure(figsize=(10, 6))
plot_series(time_valid, x_valid)
plot_series(time_valid, diff_moving_avg_plus_past)
plt.show()
# Expected output: Chart from 1100->1450+ on X. Same chart as earlier for time series, but projection overlaid looks close in value to it
print(keras.metrics.mean_squared_error(x_valid, diff_moving_avg_plus_past).numpy())
print(keras.metrics.mean_absolute_error(x_valid, diff_moving_avg_plus_past).numpy())
# EXPECTED OUTPUT
# 8.498155
# 2.327179
```
Better than naive forecast, good. However the forecasts look a bit too random, because we're just adding past values, which were noisy. Let's use a moving averaging on past values to remove some of the noise:
```
diff_moving_avg_plus_smooth_past = moving_average_forecast(series[split_time - 370:-360], 10) + diff_moving_avg
plt.figure(figsize=(10, 6))
plot_series(time_valid, x_valid)
plot_series(time_valid, diff_moving_avg_plus_smooth_past)
plt.show()
# EXPECTED OUTPUT:
# Similar chart to above, but the overlaid projections are much smoother
print(keras.metrics.mean_squared_error(x_valid, diff_moving_avg_plus_smooth_past).numpy())
print(keras.metrics.mean_absolute_error(x_valid, diff_moving_avg_plus_smooth_past).numpy())
# EXPECTED OUTPUT
# 12.527958
# 2.2034433
```
| github_jupyter |
##### Copyright 2020 The TensorFlow Authors. [Licensed under the Apache License, Version 2.0](#scrollTo=ByZjmtFgB_Y5).
```
%install '.package(url: "https://github.com/tensorflow/swift-models", .branch("tensorflow-0.10"))' Datasets ImageClassificationModels
print("\u{001B}[2J")
// #@title Licensed under the Apache License, Version 2.0 (the "License"); { display-mode: "form" }
// Licensed under the Apache License, Version 2.0 (the "License");
// you may not use this file except in compliance with the License.
// You may obtain a copy of the License at
//
// https://www.apache.org/licenses/LICENSE-2.0
//
// Unless required by applicable law or agreed to in writing, software
// distributed under the License is distributed on an "AS IS" BASIS,
// WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
// See the License for the specific language governing permissions and
// limitations under the License.
```
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/swift/tutorials/introducing_x10"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/swift/blob/master/docs/site/tutorials/introducing_x10.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/swift/blob/master/docs/site/tutorials/introducing_x10.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
</table>
# Introducing X10
By default, Swift For TensorFlow performs tensor operations using eager dispatch. This allows for rapid iteration, but isn't the most performant option for training machine learning models.
The [X10 tensor library](https://github.com/tensorflow/swift-apis/blob/master/Documentation/X10/API_GUIDE.md) adds a high-performance backend to Swift for TensorFlow, leveraging tensor tracing and the [XLA compiler](https://www.tensorflow.org/xla). This tutorial will introduce X10 and guide you through the process of updating a training loop to run on GPUs or TPUs.
## Eager vs. X10 tensors
Accelerated calculations in Swift for TensorFlow are performed through the Tensor type. Tensors can participate in a wide variety of operations, and are the fundamental building blocks of machine learning models.
By default, a Tensor uses eager execution to perform calculations on an operation-by-operation basis. Each Tensor has an associated Device that describes what hardware it is attached to and what backend is used for it.
```
import TensorFlow
import Foundation
let eagerTensor1 = Tensor([0.0, 1.0, 2.0])
let eagerTensor2 = Tensor([1.5, 2.5, 3.5])
let eagerTensorSum = eagerTensor1 + eagerTensor2
eagerTensorSum
eagerTensor1.device
```
If you are running this notebook on a GPU-enabled instance, you should see that hardware reflected in the device description above. The eager runtime does not have support for TPUs, so if you are using one of them as an accelerator you will see the CPU being used as a hardware target.
When creating a Tensor, the default eager mode device can be overridden by specifying an alternative. This is how you opt-in to performing calculations using the X10 backend.
```
let x10Tensor1 = Tensor([0.0, 1.0, 2.0], on: Device.defaultXLA)
let x10Tensor2 = Tensor([1.5, 2.5, 3.5], on: Device.defaultXLA)
let x10TensorSum = x10Tensor1 + x10Tensor2
x10TensorSum
x10Tensor1.device
```
If you're running this in a GPU-enabled instance, you should see that accelerator listed in the X10 tensor's device. Unlike for eager execution, if you are running this in a TPU-enabled instance, you should now see that calculations are using that device. X10 is how you take advantage of TPUs within Swift for TensorFlow.
The default eager and X10 devices will attempt to use the first accelerator on the system. If you have GPUs attached, the will use the first available GPU. If TPUs are present, X10 will use the first TPU core by default. If no accelerator is found or supported, the default device will fall back to the CPU.
Beyond the default eager and XLA devices, you can provide specific hardware and backend targets in a Device:
```
// let tpu1 = Device(kind: .TPU, ordinal: 1, backend: .XLA)
// let tpuTensor1 = Tensor([0.0, 1.0, 2.0], on: tpu1)
```
## Training an eager-mode model
Let's take a look at how you'd set up and train a model using the default eager execution mode. In this example, we'll be using the simple LeNet-5 model from the [swift-models repository](https://github.com/tensorflow/swift-models) and the MNIST handwritten digit classification dataset.
First, we'll set up and download the MNIST dataset.
```
import Datasets
let epochCount = 5
let batchSize = 128
let dataset = MNIST(batchSize: batchSize)
```
Next, we will configure the model and optimizer.
```
import ImageClassificationModels
var eagerModel = LeNet()
var eagerOptimizer = SGD(for: eagerModel, learningRate: 0.1)
```
Now, we will implement basic progress tracking and reporting. All intermediate statistics are kept as tensors on the same device where training is run and `scalarized()` is called only during reporting. This will be especially important later when using X10, because it avoids unnecessary materialization of lazy tensors.
```
struct Statistics {
var correctGuessCount = Tensor<Int32>(0, on: Device.default)
var totalGuessCount = Tensor<Int32>(0, on: Device.default)
var totalLoss = Tensor<Float>(0, on: Device.default)
var batches: Int = 0
var accuracy: Float {
Float(correctGuessCount.scalarized()) / Float(totalGuessCount.scalarized()) * 100
}
var averageLoss: Float { totalLoss.scalarized() / Float(batches) }
init(on device: Device = Device.default) {
correctGuessCount = Tensor<Int32>(0, on: device)
totalGuessCount = Tensor<Int32>(0, on: device)
totalLoss = Tensor<Float>(0, on: device)
}
mutating func update(logits: Tensor<Float>, labels: Tensor<Int32>, loss: Tensor<Float>) {
let correct = logits.argmax(squeezingAxis: 1) .== labels
correctGuessCount += Tensor<Int32>(correct).sum()
totalGuessCount += Int32(labels.shape[0])
totalLoss += loss
batches += 1
}
}
```
Finally, we'll run the model through a training loop for five epochs.
```
print("Beginning training...")
for (epoch, batches) in dataset.training.prefix(epochCount).enumerated() {
let start = Date()
var trainStats = Statistics()
var testStats = Statistics()
Context.local.learningPhase = .training
for batch in batches {
let (images, labels) = (batch.data, batch.label)
let 𝛁model = TensorFlow.gradient(at: eagerModel) { eagerModel -> Tensor<Float> in
let ŷ = eagerModel(images)
let loss = softmaxCrossEntropy(logits: ŷ, labels: labels)
trainStats.update(logits: ŷ, labels: labels, loss: loss)
return loss
}
eagerOptimizer.update(&eagerModel, along: 𝛁model)
}
Context.local.learningPhase = .inference
for batch in dataset.validation {
let (images, labels) = (batch.data, batch.label)
let ŷ = eagerModel(images)
let loss = softmaxCrossEntropy(logits: ŷ, labels: labels)
testStats.update(logits: ŷ, labels: labels, loss: loss)
}
print(
"""
[Epoch \(epoch)] \
Training Loss: \(String(format: "%.3f", trainStats.averageLoss)), \
Training Accuracy: \(trainStats.correctGuessCount)/\(trainStats.totalGuessCount) \
(\(String(format: "%.1f", trainStats.accuracy))%), \
Test Loss: \(String(format: "%.3f", testStats.averageLoss)), \
Test Accuracy: \(testStats.correctGuessCount)/\(testStats.totalGuessCount) \
(\(String(format: "%.1f", testStats.accuracy))%) \
seconds per epoch: \(String(format: "%.1f", Date().timeIntervalSince(start)))
""")
}
```
As you can see, the model trained as we would expect, and its accuracy against the validation set increased each epoch. This is how Swift for TensorFlow models are defined and run using eager execution, now let's see what modifications need to be made to take advantage of X10.
## Training an X10 model
Datasets, models, and optimizers contain tensors that are initialized on the default eager execution device. To work with X10, we'll need to move these tensors to an X10 device.
```
let device = Device.defaultXLA
device
```
For the datasets, we'll do that at the point in which batches are processed in the training loop, so we can re-use the dataset from the eager execution model.
In the case of the model and optimizer, we'll initialize them with their internal tensors on the eager execution device, then move them over to the X10 device.
```
var x10Model = LeNet()
x10Model.move(to: device)
var x10Optimizer = SGD(for: x10Model, learningRate: 0.1)
x10Optimizer = SGD(copying: x10Optimizer, to: device)
```
The modifications needed for the training loop come at a few specific points. First, we'll need to move the batches of training data over to the X10 device. This is done via `Tensor(copying:to:)` when each batch is retrieved.
The next change is to indicate where to cut off the traces during the training loop. X10 works by tracing through the tensor calculations needed in your code and just-in-time compiling an optimized representation of that trace. In the case of a training loop, you’re repeating the same operation over and over again, an ideal section to trace, compile, and re-use.
In the absence of code that explicitly requests a value from a Tensor (these usually stand out as `.scalars` or `.scalarized()` calls), X10 will attempt to compile all loop iterations together. To prevent this, and cut the trace at a specific point, we place an explicit `LazyTensorBarrier()` after the optimizer updates the model weights and after the loss and accuracy are obtained during validation. This creates two reused traces: each step in the training loop and each batch of inference during validation.
These modifications result in the following training loop.
```
print("Beginning training...")
for (epoch, batches) in dataset.training.prefix(epochCount).enumerated() {
let start = Date()
var trainStats = Statistics(on: device)
var testStats = Statistics(on: device)
Context.local.learningPhase = .training
for batch in batches {
let (eagerImages, eagerLabels) = (batch.data, batch.label)
let images = Tensor(copying: eagerImages, to: device)
let labels = Tensor(copying: eagerLabels, to: device)
let 𝛁model = TensorFlow.gradient(at: x10Model) { x10Model -> Tensor<Float> in
let ŷ = x10Model(images)
let loss = softmaxCrossEntropy(logits: ŷ, labels: labels)
trainStats.update(logits: ŷ, labels: labels, loss: loss)
return loss
}
x10Optimizer.update(&x10Model, along: 𝛁model)
LazyTensorBarrier()
}
Context.local.learningPhase = .inference
for batch in dataset.validation {
let (eagerImages, eagerLabels) = (batch.data, batch.label)
let images = Tensor(copying: eagerImages, to: device)
let labels = Tensor(copying: eagerLabels, to: device)
let ŷ = x10Model(images)
let loss = softmaxCrossEntropy(logits: ŷ, labels: labels)
LazyTensorBarrier()
testStats.update(logits: ŷ, labels: labels, loss: loss)
}
print(
"""
[Epoch \(epoch)] \
Training Loss: \(String(format: "%.3f", trainStats.averageLoss)), \
Training Accuracy: \(trainStats.correctGuessCount)/\(trainStats.totalGuessCount) \
(\(String(format: "%.1f", trainStats.accuracy))%), \
Test Loss: \(String(format: "%.3f", testStats.averageLoss)), \
Test Accuracy: \(testStats.correctGuessCount)/\(testStats.totalGuessCount) \
(\(String(format: "%.1f", testStats.accuracy))%) \
seconds per epoch: \(String(format: "%.1f", Date().timeIntervalSince(start)))
""")
}
```
Training of the model using the X10 backend should have proceeded in the same manner as the eager execution model did before. You may have noticed a delay before the first batch and at the end of the first epoch, due to the just-in-time compilation of the unique traces at those points. If you're running this with an accelerator attached, you should have seen the training after that point proceeding faster than with eager mode.
There is a tradeoff of initial trace compilation time vs. faster throughput, but in most machine learning models the increase in throughput from repeated operations should more than offset compilation overhead. In practice, we've seen an over 4X improvement in throughput with X10 in some training cases.
As has been stated before, using X10 now makes it not only possible but easy to work with TPUs, unlocking that whole class of accelerators for your Swift for TensorFlow models.
| github_jupyter |
# Introduction
Quandl provides financial data.
Data is queried through Quandl's RESTful API. To use it you need to register for an API key.
```
# Installation of the pip package in the current Jupyter kernel
import sys
#!{sys.executable} -m pip install quandl
import quandl
import config as cfg
import altair as alt
# Authentication
quandl.ApiConfig.api_key = cfg.QUANDL['API_KEY']
```
# Time-Series
```
# Getting WTI Crude Oil Price from the US Dept. of Energy dataset
data_oil = quandl.get('EIA/PET_RWTC_D')
```
### Time-series parameters
You can slice, transform and otherwise customize your time-series dataset prior to download by appending various optional parameters to your query.
```
# Getting data wiht limit
data_oil = quandl.get('EIA/PET_RWTC_D', limit = 100)
# Getting data with start and end dates
data_oil = quandl.get('EIA/PET_RWTC_D', start_date="2001-12-31", end_date="2021-05-03")
# Collapsing data
data_oil = quandl.get('EIA/PET_RWTC_D', start_date="2001-12-31", end_date="2021-05-03", collapse = 'annual')
data_oil.head()
# Plotting using Altair
# Since 'Date' is an index in the DataFrame, reset_index() needs to be run
crude_oil = alt.Chart(data_oil.reset_index()).mark_line().encode(x = 'Date', y='Value')
crude_oil
```
### Transformations
```
data_gdp = quandl.get("FRED/GDP", transformation="rdiff")
data_gdp
alt.Chart(data_gdp.reset_index()).mark_bar().encode(
x = 'Date:T',
y = 'Value:Q',
color=alt.condition(
alt.datum.Value > 0,
alt.value("steelblue"), # The positive color
alt.value("orange") # The negative color
)
).properties(width = 600)
```
### Downloading a dataset
```
# downloads a zip file, 2.7 MB for 'MX' dataset
quandl.bulkdownload('MX')
quandl.get('MX/BAXG2020')
```
### Examples
```
# variables
five_years = '2016-05-03'
two_years = '2019-05-03'
end_date = '2021-05-03'
# dataset example 1
datasets_ex1 = {
'Nasdaq Composite': 'NASDAQOMX/COMP',
'Nasdaq Index': 'NASDAQOMX/NDX',
'Nasdaq Small Cap': 'NASDAQOMX/NQUSS',
'Nasdaq Mid Cap': 'NASDAQOMX/NQUSM',
'Nasdaq Large Cap': 'NASDAQOMX/NQUSL',
'Michigan Consumer Sentiment Index': 'UMICH/SOC1',
}
data_ex1 = {}
for k, v in datasets_ex1.items():
data_ex1[k] = quandl.get(v, start_date = five_years, end_date = end_date)
data_ex1['Michigan Consumer Sentiment Index'].head()
chart_nasdaq = alt.Chart(data_ex1['Nasdaq Composite'].reset_index()).mark_line(stroke = '#f4a261').encode(
x = 'Trade Date', y='Index Value')#.configure_mark(color = '#264653')
chart_consumer_sent = alt.Chart(data_ex1['Michigan Consumer Sentiment Index'].reset_index()).mark_line(stroke = '#2a9d8f').encode( x = 'Date', y='Index')#.configure_mark(color = '#e76f51')
alt.layer(chart_nasdaq, chart_consumer_sent).resolve_scale(y = 'independent').properties(
title = 'NASDAQ Composite vs Consumer Sentiment'
)
datasets_ex2 = {
'Lumber Futures': 'CHRIS/CME_LB1',
'Gold Futures': 'CHRIS/CME_GC5',
'Crude Oil Futures': 'CHRIS/CME_CL11',
}
data_ex2 = {}
for k, v in datasets_ex2.items():
data_ex2[k] = quandl.get(v, start_date = two_years, end_date = end_date, transformation="normalize")
chart_lumber = alt.Chart(data_ex2['Lumber Futures'].reset_index()).mark_line(stroke = '#264653').encode(
x = 'Date:T', y='Open')#.configure_mark(color = '#264653')
chart_gold = alt.Chart(data_ex2['Gold Futures'].reset_index()).mark_line(stroke = '#e9c46a').encode(
x = 'Date:T', y='Open')#.configure_mark(color = '#264653')
chart_oil = alt.Chart(data_ex2['Crude Oil Futures'].reset_index()).mark_line(stroke = '#e76f51').encode(
x = 'Date:T', y='Open')#.configure_mark(color = '#264653')
alt.layer(chart_lumber, chart_gold, chart_oil).properties(
title = 'Normalized Gold, Lumber, Oil Futures'
)
```
# Tables
Tables are used to show unsorted data types (strings, numbers, dates, etc.). They can be filtered by diffrent fields allowing for more customization.
```
# Downloading the Noika data table
data = quandl.get_table('MER/F1', compnumber="39102", paginate=True)
data.head()
# You can also download specific columns from tables
# In this example we will pull the report date, indicator, and amount columns
data3 = quandl.get_table('MER/F1',compnumber="39102", qopts={"columns":["reporttype", "statement"]}, paginate=True)
data3
# Another usefull example is getting data for a certain time frame
#in this example we will pull the data for nokia again but this time only see the data from January 01 2010 to December 31 2011
data4 = quandl.get_table('MER/F1',compnumber="39102",qopts={"columns":['reportdate',"reporttype", "statement"]},reportdate={ 'gte': '2010-01-01', 'lte': '2011-12-31' }, paginate=True)
data4
# If you have the table with the data you would like, you can export that data to your desktop
# We will now save the table to our desktop
quandl.export_table('MER/F1',compnumber="39102",qopts={"columns":['reportdate',"reporttype", "statement"]},reportdate={ 'gte': '2010-01-01', 'lte': '2011-12-31' },filename='/Users/shahnilpunja/Downloads/quandl/Nokia.zip')
```
| github_jupyter |
# Introduction to Recurrent Neural Networks
(c) Deniz Yuret, 2019
* Objectives: learn about RNNs, the RNN layer, compare with MLP on a tagging task.
* Prerequisites: [MLP models](40.mlp.ipynb)
* New functions:
[RNN](http://denizyuret.github.io/Knet.jl/latest/reference/#Knet.RNN),
[adam](http://denizyuret.github.io/Knet.jl/latest/reference/#Knet.adam)
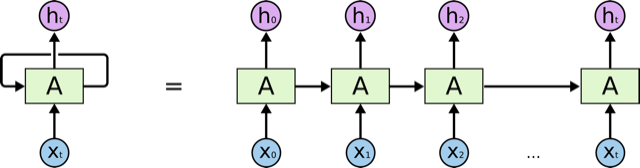([image
source](http://colah.github.io/posts/2015-08-Understanding-LSTMs))
In this notebook we will see how to implement a recurrent neural network (RNN) in Knet. In RNNs, connections between units form a directed cycle, which allows them to keep a persistent state over time. This gives them the ability to process sequences of arbitrary length one element at a time, while keeping track of what happened at previous elements. One can view the current state of the RNN as a representation for the sequence processed so far.
We will build a part-of-speech tagger using a large annotated corpus of English. We will represent words with numeric vectors appropriate as inputs to a neural network. These word vectors will be initialized randomly and learned during training just like other model parameters. We will compare three network architectures: (1) an MLP which tags each word independently of its neighbors, (2) a simple RNN that can represent the neighboring words to the left, (3) a bidirectional RNN that can represent both left and right contexts. As can be expected 1 < 2 < 3 in performance. More surprisingly, the three models are very similar to each other: we will see their model diagrams are identical except for the horizontal connections that carry information across the sequence.
```
# Setup display width, load packages, import symbols
ENV["COLUMNS"] = 72
using Pkg; for p in ("Knet","Plots"); haskey(Pkg.installed(),p) || Pkg.add(p); end
using Random: shuffle!
using Base.Iterators: flatten
using Knet: Knet, AutoGrad, param, param0, mat, RNN, relu, Data, adam, progress, nll, zeroone
```
## The Brown Corpus
To introduce recurrent neural networks (RNNs) we will train a part-of-speech tagger using the [Brown Corpus](https://en.wikipedia.org/wiki/Brown_Corpus). We will train three models: a MLP, a unidirectional RNN, a bidirectional RNN and observe significant performance differences.
```
include(Knet.dir("data/nltk.jl"))
(data,words,tags) = brown()
println("The Brown Corpus has $(length(data)) sentences, $(sum(length(p[1]) for p in data)) tokens, with a word vocabulary of $(length(words)) and a tag vocabulary of $(length(tags)).")
```
`data` is an array of `(w,t)` pairs each representing a sentence, where `w` is a sequence of word ids, and `t` is a sequence of tag ids. `words` and `tags` contain the strings for the ids.
```
println.(summary.((data,words,tags)));
```
Here is what the first sentence looks like with ids and with strings:
```
(w,t) = first(data)
display(permutedims(Int[w t]))
display(permutedims([words[w] tags[t]]))
```
## Chain of layers
```
# Let's define a chain of layers
struct Chain
layers
Chain(layers...) = new(layers)
end
(c::Chain)(x) = (for l in c.layers; x = l(x); end; x)
(c::Chain)(x,y) = nll(c(x),y)
```
## Dense layers
```
# Redefine dense layer (See mlp.ipynb):
struct Dense; w; b; f; end
Dense(i::Int,o::Int,f=identity) = Dense(param(o,i), param0(o), f)
(d::Dense)(x) = d.f.(d.w * mat(x,dims=1) .+ d.b)
```
## Word Embeddings
`data` has each sentence tokenized into an array of words and each word mapped to a `UInt16` id. To use these words as inputs to a neural network we further map each word to a Float32 vector. We will keep these vectors in the columns of a size (X,V) matrix where X is the embedding dimension and V is the vocabulary size. The vectors will be initialized randomly, and trained just like any other network parameter. Let's define an embedding layer for this purpose:
```
struct Embed; w; end
Embed(vocabsize::Int,embedsize::Int) = Embed(param(embedsize,vocabsize))
(e::Embed)(x) = e.w[:,x]
```
This is what the words, word ids and embeddings for a sentence looks like: (note the identical id and embedding for the 2nd and 5th words)
```
embedlayer = Embed(length(words),8)
(w,t) = data[52855]
display(permutedims(words[w]))
display(permutedims(Int.(w)))
display(embedlayer(w))
```
## RNN layers
```
@doc RNN
```
## The three taggers: MLP, RNN, biRNN
## Tagger0 (MLP)
This is what Tagger0 looks like. Every tag is predicted independently. The prediction of each tag only depends on the corresponding word.
<img src="https://docs.google.com/drawings/d/e/2PACX-1vTfV4-TB0KwjDbFKpj3rL0tfeApEh9XXaDJ1OF3emNVAmc_-hvgqpEBuA_K0FsNuxymZrv3ztScXxqF/pub?w=378&h=336"/>
## Tagger1 (RNN)
In Tagger1, the RNN layer takes its previous output as an additional input. The prediction of each tag is based on words to the left.
<img src="https://docs.google.com/drawings/d/e/2PACX-1vTaizzCISuSxihPCjndr7xMVwklsrefi9zn7ZArCvsR8fb5V4DGKtusyIzn3Ujp3QbAJgUz1WSlLvIJ/pub?w=548&h=339"/>
## Tagger2 (biRNN)
In Tagger2 there are two RNNs: the forward RNN reads the sequence from left to right, the backward RNN reads it from right to left. The prediction of each tag is dependent on all the words in the sentence.
<img src="https://docs.google.com/drawings/d/e/2PACX-1vQawvnCj6odRF2oakF_TgXd8gLxSsfQP8-2ZdBdEIpfgIyPq0Zp_EF6zcFJf6JlGhfiKQvdVyg-Weq2/pub?w=566&h=335"/>
```
Tagger0(vocab,embed,hidden,output)= # MLP Tagger
Chain(Embed(vocab,embed),Dense(embed,hidden,relu),Dense(hidden,output))
Tagger1(vocab,embed,hidden,output)= # RNN Tagger
Chain(Embed(vocab,embed),RNN(embed,hidden,rnnType=:relu),Dense(hidden,output))
Tagger2(vocab,embed,hidden,output)= # biRNN Tagger
Chain(Embed(vocab,embed),RNN(embed,hidden,rnnType=:relu,bidirectional=true),Dense(2hidden,output));
```
## Sequence Minibatching
Minibatching is a bit more complicated with sequences compared to simple classification problems, this section can be skipped on a first reading. In addition to the input and minibatch sizes, there is also the time dimension to consider. To keep things simple we will concatenate all sentences into one big sequence, then split this sequence into equal sized chunks. The input to the tagger will be size (B,T) where B is the minibatch size, and T is the chunk size. The input to the RNN layer will be size (X,B,T) where X is the embedding size.
```
BATCHSIZE = 64
SEQLENGTH = 32;
function seqbatch(x,y,B,T)
N = length(x) ÷ B
x = permutedims(reshape(x[1:N*B],N,B))
y = permutedims(reshape(y[1:N*B],N,B))
d = []; for i in 0:T:N-T
push!(d, (x[:,i+1:i+T], y[:,i+1:i+T]))
end
return d
end
allw = vcat((x->x[1]).(data)...)
allt = vcat((x->x[2]).(data)...)
d = seqbatch(allw, allt, BATCHSIZE, SEQLENGTH);
```
This may be a bit more clear if we look at an example minibatch:
```
(x,y) = first(d)
words[x]
```
## Embedding a minibatch
Julia indexing allows us to get the embeddings for this minibatch in one go as an (X,B,T) array where X is the embedding size, B is the minibatch size, and T is the subsequence length.
```
embedlayer = Embed(length(words),128)
summary(embedlayer(x))
```
## Experiments
```
# shuffle and split minibatches into train and test portions
shuffle!(d)
dtst = d[1:10]
dtrn = d[11:end]
length.((dtrn,dtst))
# For running experiments we will use the Adam algorithm which typically converges faster than SGD.
function trainresults(file,maker,savemodel)
if (print("Train from scratch? "); readline()[1]=='y')
model = maker()
takeevery(n,itr) = (x for (i,x) in enumerate(itr) if i % n == 1)
results = ((nll(model,dtst), zeroone(model,dtst))
for x in takeevery(100, progress(adam(model,repeat(dtrn,5)))))
results = reshape(collect(Float32,flatten(results)),(2,:))
Knet.save(file,"model",(savemodel ? model : nothing),"results",results)
Knet.gc() # To save gpu memory
else
isfile(file) || download("http://people.csail.mit.edu/deniz/models/tutorial/$file",file)
model,results = Knet.load(file,"model","results")
end
println(minimum(results,dims=2))
return model,results
end
VOCABSIZE = length(words)
EMBEDSIZE = 128
HIDDENSIZE = 128
OUTPUTSIZE = length(tags);
# 2.35e-01 100.00%┣┫ 2780/2780 [00:13/00:13, 216.36i/s] [0.295007; 0.0972656]
t0maker() = Tagger0(VOCABSIZE,EMBEDSIZE,HIDDENSIZE,OUTPUTSIZE)
(t0,r0) = trainresults("tagger113a.jld2",t0maker,false);
# 1.49e-01 100.00%┣┫ 2780/2780 [00:19/00:19, 142.58i/s] [0.21358; 0.0616211]
t1maker() = Tagger1(VOCABSIZE,EMBEDSIZE,HIDDENSIZE,OUTPUTSIZE)
(t1,r1) = trainresults("tagger113b.jld2",t1maker,false);
# 9.37e-02 100.00%┣┫ 2780/2780 [00:25/00:25, 109.77i/s] [0.156669; 0.044043]
t2maker() = Tagger2(VOCABSIZE,EMBEDSIZE,HIDDENSIZE,OUTPUTSIZE)
(t2,r2) = trainresults("tagger113c.jld2",t2maker,true);
using Plots; default(fmt=:png,ls=:auto,ymirror=true)
plot([r0[2,:], r1[2,:], r2[2,:]]; xlabel="x100 updates", ylabel="error",
ylim=(0,0.15), yticks=0:0.01:0.15, labels=["MLP","RNN","biRNN"])
plot([r0[1,:], r1[1,:], r2[1,:]]; xlabel="x100 updates", ylabel="loss",
ylim=(0,.5), yticks=0:0.1:.5, labels=["MLP","RNN","biRNN"])
```
## Playground
Below, you can type and tag your own sentences:
```
wdict=Dict{String,UInt16}(); for (i,w) in enumerate(words); wdict[w]=i; end
unk = UInt16(length(words))
wid(w) = get(wdict,w,unk)
function tag(tagger,s::String)
w = permutedims(split(s))
t = tags[(x->x[1]).(argmax(Array(tagger(wid.(w))),dims=1))]
vcat(w,t)
end
tag(t2,readline())
```
| github_jupyter |
```
# Copyright 2019 Google Inc. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# Install Pipeline SDK - This only needs to be ran once in the enviroment.
!pip3 install kfp --upgrade
!pip3 install tensorflow==1.14 --upgrade
```
## KubeFlow Pipelines Serving Component
In this notebook, we will demo:
* Saving a Keras model in a format compatible with TF Serving
* Creating a pipeline to serve a trained model within a KubeFlow cluster
Reference documentation:
* https://www.tensorflow.org/tfx/serving/architecture
* https://www.tensorflow.org/beta/guide/keras/saving_and_serializing
* https://www.kubeflow.org/docs/components/serving/tfserving_new/
### Setup
```
# Set your output and project. !!!Must Do before you can proceed!!!
project = 'Your-Gcp-Project-ID' #'Your-GCP-Project-ID'
model_name = 'model-name' # Model name matching TF_serve naming requirements
experiment_name = 'serving_component'
import time
ts = int(time.time())
model_version = str(ts) # Here we use timestamp as version to avoid conflict
output = 'Your-Gcs-Path' # A GCS bucket for asset outputs
KUBEFLOW_DEPLOYER_IMAGE = 'gcr.io/ml-pipeline/ml-pipeline-kubeflow-deployer:57d9f7f1cfd458e945d297957621716062d89a49'
model_path = '%s/%s' % (output,model_name)
model_version_path = '%s/%s/%s' % (output,model_name,model_version)
#Get or create an experiment and submit a pipeline run
import kfp
client = kfp.Client()
try:
experiment = client.get_experiment(experiment_name=experiment_name)
except:
experiment = client.create_experiment(experiment_name)
```
### Load a Keras Model
Loading a pretrained Keras model to use as an example.
```
import tensorflow as tf
model = tf.keras.applications.NASNetMobile(input_shape=None,
include_top=True,
weights='imagenet',
input_tensor=None,
pooling=None,
classes=1000)
```
### Saved the Model for TF-Serve
Save the model using keras export_saved_model function. Note that specifically for TF-Serve the output directory should be structure as model_name/model_version/saved_model.
```
tf.keras.experimental.export_saved_model(model, model_version_path)
```
### Create a pipeline using KFP TF-Serve component
```
def kubeflow_deploy_op():
return dsl.ContainerOp(
name = 'deploy',
image = KUBEFLOW_DEPLOYER_IMAGE,
arguments = [
'--model-export-path', model_path,
'--server-name', model_name,
]
)
import kfp
import kfp.dsl as dsl
from kfp.gcp import use_gcp_secret
# The pipeline definition
@dsl.pipeline(
name='Sample model deployer',
description='Sample for deploying models using KFP model serving component'
)
def model_server():
deploy = kubeflow_deploy_op().apply(use_gcp_secret('user-gcp-sa'))
pipeline_func = model_server
pipeline_filename = pipeline_func.__name__ + '.pipeline.zip'
import kfp.compiler as compiler
compiler.Compiler().compile(pipeline_func, pipeline_filename)
#Specify pipeline argument values
arguments = {}
#Submit a pipeline run
run_name = pipeline_func.__name__ + ' run'
run_result = client.run_pipeline(experiment.id, run_name, pipeline_filename, arguments)
#This link leads to the run information page.
#Note: There is a bug in JupyterLab that modifies the URL and makes the link stop working
```
| github_jupyter |
# Clustering
Perform data clustering (try several methods for this purpose, at least 3) and check if there are any segments of borrowers, use appropriate methods for the optimal number of clusters (40 points)
```
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
import seaborn as sns
#importing the PCA scaling library
from sklearn.decomposition import PCA
from sklearn.decomposition import IncrementalPCA
from sklearn.linear_model import Ridge, Lasso
# Import KNN Regressor machine learning library
from sklearn.neighbors import KNeighborsRegressor
from sklearn import metrics
# Import stats from scipy
from scipy import stats
# Import zscore for scaling
from scipy.stats import zscore
#importing the metrics
from sklearn.metrics import mean_squared_error,mean_absolute_error,r2_score
from sklearn import preprocessing
# importing the Polynomial features
from sklearn.preprocessing import PolynomialFeatures
#importing kmeans clustering library
from sklearn.cluster import KMeans
from sklearn.utils import resample
from sklearn.datasets import make_classification
from numpy import where
from sklearn.metrics import silhouette_score, silhouette_samples
dummies_loan_status = pd.read_csv('dummies_loan_status.csv')
pd.set_option('display.max_colwidth', None)
pd.set_option('display.max_row', None)
dummies_loan_status.head()
dummies_loan_status.drop('Unnamed: 0', axis = 1, inplace = True)
dummies_loan_status.shape
# define dataset
X = dummies_loan_status.drop('loan_status', axis = 1)
y = dummies_loan_status.loan_status
X.shape, y.shape
from sklearn.decomposition import PCA
pca = PCA(n_components=2, svd_solver = 'full')
df_pca = pca.fit_transform(X)
loan_pca = pd.DataFrame(df_pca, columns=['c1', 'c2'], index=X.index)
loan_pca.head()
sns.distplot(loan_pca.c1).set(title = 'Density graph c1')
plt.show()
sns.distplot(loan_pca.c2).set(title = 'Density graph c2')
plt.show()
for col in loan_pca:
if loan_pca[col].min() <= 0:
loan_pca[col] = loan_pca[col] + np.abs(loan_pca[col].min()) + 1
loan_pca = np.log(loan_pca)
q1 = loan_pca.quantile(0.25)
q3 = loan_pca.quantile(0.75)
iqr = q3 - q1
low_boundary = (q1 - 1.5 * iqr)
upp_boundary = (q3 + 1.5 * iqr)
num_of_outliers_L = (loan_pca[iqr.index] < low_boundary).sum()
num_of_outliers_U = (loan_pca[iqr.index] > upp_boundary).sum()
outliers = pd.DataFrame({'lower_boundary':low_boundary, 'upper_boundary':upp_boundary,'num_of_outliers__lower_boundary':num_of_outliers_L, 'num_of_outliers__upper_boundary':num_of_outliers_U})
outliers
for row in outliers.iterrows():
loan_pca = loan_pca[(loan_pca[row[0]] >= row[1]['lower_boundary']) & (loan_pca[row[0]] <= row[1]['upper_boundary'])]
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
scaler.fit(loan_pca)
loan_pca_std = scaler.transform(loan_pca)
loan_pca_df = pd.DataFrame(data=loan_pca_std, index=loan_pca.index, columns=loan_pca.columns)
sns.distplot(loan_pca_df.c1).set(title = 'Density graph c1')
plt.show()
sns.distplot(loan_pca_df.c2).set(title = 'Density graph c2')
plt.show()
loan_pca_df.agg(['mean', 'std', 'max', 'min']).round(2)
loan_pca = loan_pca_df.sample(1000, random_state=42)
sns.lmplot('c1', 'c2', data = loan_pca, fit_reg=False).set(title = 'Scatterplot')
plt.show()
from sklearn.cluster import AgglomerativeClustering
from scipy.cluster.hierarchy import dendrogram, linkage, fcluster
model_AggCl = AgglomerativeClustering(linkage='ward', affinity='euclidean', n_clusters=3).fit(loan_pca)
loan_pca['AggCl'] = model_AggCl.labels_
silhouette_score(loan_pca[['c1', 'c2']], loan_pca['AggCl']).round(4)
sns.lmplot('c1', 'c2', data = loan_pca, hue = 'AggCl', fit_reg=False).set(title='Groups visualisation')
plt.show()
model_link = linkage(loan_pca.iloc[:,0:2], method = 'ward', metric = 'euclidean')
fig = plt.figure(figsize=(20, 10))
dn = dendrogram(model_link)
plt.show()
clusters = fcluster(model_link, 3, criterion='maxclust')
loan_pca['linkage'] = clusters
sns.lmplot('c1', 'c2', data = loan_pca, fit_reg=False, hue = 'linkage')
plt.show()
silhouette_score(loan_pca[['c1', 'c2']], loan_pca['linkage']).round(4)
from sklearn.cluster import DBSCAN
model_DBSCAN = DBSCAN(eps = 0.3, min_samples=30, leaf_size=60).fit(loan_pca.iloc[:,0:2])
loan_pca['DBSCAN'] = model_DBSCAN.labels_
sns.lmplot('c1', 'c2', data = loan_pca, fit_reg=False, hue = 'DBSCAN')
plt.show()
silhouette_score(loan_pca[['c1', 'c2']], loan_pca['DBSCAN']).round(4)
ssd = [] # Sum of squared distances
range_n_clusters = [2, 3, 4, 5, 6, 7, 8]
for num_clusters in range_n_clusters:
kmeans = KMeans(n_clusters=num_clusters, max_iter=1000)
kmeans.fit(loan_pca.iloc[:,0:2])
ssd.append(kmeans.inertia_) #Sum of squared distances
# plot the SSDs for each n_clusters
# ssd
plt.plot(ssd)
range_n_clusters = [2, 3, 4, 5, 6, 7, 8]
for num_clusters in range_n_clusters:
# intialise kmeans
kmeans = KMeans(n_clusters=num_clusters, max_iter=1000)
kmeans.fit(loan_pca.iloc[:,0:2])
cluster_labels = kmeans.labels_
# silhouette score
silhouette_avg = silhouette_score(loan_pca.iloc[:,0:2], cluster_labels)
print("For n_clusters={0}, the silhouette score is {1}".format(num_clusters, silhouette_avg))
model_km = KMeans(n_clusters=4, max_iter=1000, random_state=42)
model_km.fit(loan_pca.iloc[:,0:2])
loan_pca['KMeans'] = model_km.labels_
sns.lmplot('c1', 'c2', data = loan_pca, fit_reg=False, hue = 'KMeans')
plt.scatter(model_km.cluster_centers_[:, 0], model_km.cluster_centers_[:, 1], c='black', s=100, alpha=0.5)
plt.show()
silhouette_score(loan_pca[['c1', 'c2']], loan_pca['KMeans']).round(4)
loan_pca
```
As we can see above 4 different models created from 3 to 4 clusters. As I worked on the 1000 observation sample, I'm going to choose one (KMeans) and elaborate it with whole dataset in a new notebook
| github_jupyter |
```
%matplotlib notebook
import os
import numpy as np
import cv2
import imgaug as ia
import keras.backend as K
from sklearn.model_selection import train_test_split
import imgaug.augmenters as iaa
from imgaug.augmentables.kps import KeypointsOnImage
from sklearn.utils import class_weight
from keras.layers import GlobalAveragePooling2D, Dense, Dropout, Flatten, Input, Conv2D, multiply, LocallyConnected2D, Lambda, AvgPool2D
from keras.models import Sequential,Model
from keras.layers import ReLU, Dense, Conv2D, Flatten,Dropout, MaxPooling2D, GlobalAveragePooling2D, LeakyReLU, Activation, BatchNormalization, Input, merge, Softmax
import matplotlib.pyplot as plt
from keras.utils import to_categorical
import tensorflow as tf
from keras.callbacks import EarlyStopping, ModelCheckpoint, ReduceLROnPlateau
import random
from keras.constraints import max_norm
from keras.optimizers import Adam
from keras.applications.inception_v3 import InceptionV3
from sklearn.metrics import multilabel_confusion_matrix
from sklearn.metrics import precision_recall_curve
import matplotlib.pyplot as plt
from inspect import signature
from sklearn.metrics import average_precision_score,mean_squared_error
from keras.applications.inception_v3 import InceptionV3,preprocess_input
font = cv2.FONT_HERSHEY_SIMPLEX
def loadData(noBatchSamples,batchIdx,testData="test_data.txt",rootPathCentreLabel="./obj/labels",rootPathCroppedImages = "./obj/images"):
"""This function loads data from the directory of labels, it works with the yolo data format.
Args:
noBatchSamples (int) : Number of samples per batch
batchIdx (int) : Batch number
duplicativeFactor (int) : Number of times to oversample rare date
rareData (str) : Filenames of rare data samples
rootPathCentreLabel (str) : Directory with labels in yolo format
rootPathCroppedImages (str) : Directory with images, image name and label name should be same eg: 1.jpg 1.txt
Returns:
list: Images in list
list: Targets
list: File names
"""
y_train=[]
X_train=[]
name=[]
f = open(testData)
lines = f.readlines()
lines = [x.strip() for x in lines]
f.close()
for ind,element in enumerate(os.listdir(rootPathCentreLabel)):
if ind >= noBatchSamples*batchIdx and ind<=noBatchSamples*(batchIdx+1):
with open(os.path.join(rootPathCentreLabel,element)) as fin:
y = [(0,0),(0,0)]
# print(os.path.join(rootPathCroppedImages,element.replace(".txt",".jpg")))
img = cv2.imread(os.path.join(rootPathCroppedImages,element.replace(".txt",".jpg")),cv2.IMREAD_COLOR)
img = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
for line in fin:
line=line.split(" ")
if line[0]=="0" :
pixel_y_pos =int(float(line[2]))
y[0]=(int(float(line[1])*100),int(float(line[2])*2000))
elif line[0]=="1" :
pixel_y_pos =int(float(line[2]))
if float(line[2])<1:
y[1]=(int(float(line[1])*100),int(float(line[2])*2000))
elif line[0]=="2" :
pixel_y_pos =int(float(line[2]))
if float(line[2])<1:
y[1]=(int(float(line[1])*100),int(float(line[2])*2000))
if element in lines:
y_train.append(y)
X_train.append(img)
name.append(element)
else:
pass
combined = list(zip(X_train, y_train,name))
random.Random(23).shuffle(combined)
X_train[:], y_train[:],name[:] = zip(*combined)
X_train=np.array(X_train)
print(len(X_train))
return X_train,y_train,name #np.zeros((128, 32, 32, 3), dtype=np.uint8) + (batch_idx % 255)
def key2Target(keypoints,name):
"""This function converts keypoints returned after data augmentation to numpy arrays.
Args:
keypoints (imgaug.augmentables.kps.KeypointsOnImage) : Keypoints on the image
name (list) : File names
Returns:
list: Images in list
list: Targets
list: File names
"""
numred=0
y_test_regression=[]
y_test_categorical = []
for i,k in enumerate(keypoints):
y=np.zeros((2))
y_class=np.zeros((2))
if k[0][1]<=700 and (k[1][1]<=700 or k[1][1]>=1800): # Red line:False && Blue line:False
y[0]=0
y[1]=0
y_class[0]=0
y_class[1]=0
elif k[0][1]>=700 and (k[1][1]<=700 or k[1][1]>=1800): # Red line:False && Blue line:True
y[0]=k[0][1]/2000
y[1]=0
y_class[0]=1
y_class[1]=0
elif (k[1][1]>=700 and k[1][1]<=1900) and k[0][1]>=700: # Red line:True && Blue line:True
numred+=1
y[0]=k[0][1]/2000
y[1]=k[1][1]/2000
y_class[0]=1
y_class[1]=1
y_test_regression.append(np.array(y))
y_test_categorical.append(np.array(y_class))
return np.array(y_test_regression),np.array(y_test_categorical)
def returnAugmentationObj(percentageOfChance=0.):
"""This function returns an augementation pipeline which can be used to augment training data.
Args:
percentageOfChance (float) : Percentage of chance , eg: if it is 0.5, 50% of the images will go through the pipeline
Returns:
:class:`imgaug.augmenters.meta.Sequential` : Image augmentor
"""
sometimes = lambda aug: iaa.Sometimes(percentageOfChance, aug)
# Define our sequence of augmentation steps that will be applied to every image
# All augmenters with per_channel=0.5 will sample one value _per image_
# in 50% of all cases. In all other cases they will sample new values
# _per channel_.
seq = iaa.Sequential(
[
sometimes(iaa.Affine(
translate_percent={"x": (-0.05, 0.05), "y": (-0.03, 0.03)} # translate by -x to +x percent (per axis)
))
])
return seq
def lossReg(y_true,y_pred):
"""Custom loss function to penalize A type virus versus B type written for keras.
"""
mask=K.ones_like(y_true)
l=K.square(y_pred-y_true)
penalty = tf.constant([10.0])
mask =tf.add(penalty,tf.to_float (tf.math.logical_or(tf.math.logical_and(tf.math.greater(y_true[:,0],y_true[:,1]),tf.math.less(y_pred[:,0],y_pred[:,1])),tf.math.logical_and(tf.math.less(y_true[:,0],y_true[:,1]),tf.math.greater(y_pred[:,0],y_pred[:,1])))))
mask = tf.stack([K.ones_like(y_true[:,0]),mask],axis=1)
return K.mean(tf.math.multiply(l,mask),axis=-1)
def returnModel(loadWeights,weightsFile="./red_blue_cust.hdf5"):
"""This function returns a keras model.
Args:
loadWeights (bool) : Load weights specified in the weightsFile param
weightsFile (str) : Path to weights
Returns:
:class:`keras.model.Model` : Neural Network
"""
x = Input(shape=(500, 100,3))
conv1=Conv2D(8, (3,3), padding='valid')(x)
batchnorm1 = BatchNormalization()(conv1)
act1 = ReLU()(batchnorm1)
conv2=Conv2D(8, (3,3), padding='valid')(act1)
batchnorm2 = BatchNormalization()(conv2)
act2 = ReLU()(batchnorm2)
maxpool2 = MaxPooling2D((2,2))(act2)
conv3=Conv2D(16, (3,3), padding='valid')(maxpool2)
batchnorm3 = BatchNormalization()(conv3)
act3 = ReLU()(batchnorm3)
conv4=Conv2D(16, (3,3), padding='valid')(act3)
batchnorm4 = BatchNormalization()(conv4)
act4 = ReLU()(batchnorm4)
maxpool3 = MaxPooling2D((2,2))(act4)
flat1 = Flatten()(maxpool3)
D1 = Dense(256)(flat1)
batchnorm5 = BatchNormalization()(D1)
act5 = ReLU()(batchnorm5)
D2 = Dense(128,kernel_constraint=max_norm(2))(act5)
batchnorm6 = BatchNormalization()(D2)
act6 = ReLU()(batchnorm6)
D_soft = Dense(2)(act6)
batchnorm7 = BatchNormalization()(D_soft)
out1 = Activation('sigmoid',name="cat_kash")(batchnorm7)
D_sigmoid = Dense(2)(act6)
batchnorm8 = BatchNormalization()(D_sigmoid)
out2 = Activation('sigmoid',name="reg_kash")(batchnorm8)
model = Model(inputs=x, outputs=[out1,out2])
if (loadWeights):
model.load_weights(weightsFile,by_name=True)
return model
def modelTransferLearning(loadWeights,weightsFile="./red_blue_transf.hdf5"):
base_model = InceptionV3(weights='imagenet', include_top=False)
x = base_model.output
x = GlobalAveragePooling2D()(x)
D1 = Dense(256)(x)
batchnorm5 = BatchNormalization()(D1)
act5 = ReLU()(batchnorm5)
D2 = Dense(128)(act5)
batchnorm6 = BatchNormalization()(D2)
act6 = ReLU()(batchnorm6)
D_soft = Dense(2)(act6)
batchnorm7 = BatchNormalization()(D_soft)
out1 = Activation('sigmoid',name="cat_kash")(batchnorm7)
D_sigmoid = Dense(2)(act6)
batchnorm8 = BatchNormalization()(D_sigmoid)
out2 = Activation('sigmoid',name="reg_kash")(batchnorm8)
model = Model(inputs=base_model.input, outputs=[out1,out2])
for layer in model.layers[:249]:
layer.trainable = False
for layer in model.layers[249:]:
layer.trainable = True
# model.summary()
if (loadWeights):
model.load_weights(weightsFile,by_name=True)
return model
config = tf.ConfigProto( device_count = {'GPU': 0 , 'CPU': 4} ) # Correctly put number of GPU and CPU
sess = tf.Session(config=config)
with sess:
useTransferLearning = False
SeqAug = returnAugmentationObj()
print("loading model...")
if useTransferLearning:
model = modelTransferLearning(True,"red_blue_transf.hdf5")
else:
model = returnModel(True,"red_blue_cust.hdf5")
filepath="weights-latest_model_YCrCb_test.hdf5" # Name and path of weights to save
checkpoint = ModelCheckpoint(filepath, monitor='val_loss', verbose=1, save_best_only=True, mode='min') # Checkpoint call back to save best model on validation set
lrd=ReduceLROnPlateau(monitor='val_loss', factor=0.9, patience=10, verbose=1, mode='auto', min_delta=0.00001, cooldown=5, min_lr=0.00000000000000000001) # Callback to control learning rate on plateau condition
print("loading data...")
X_train,y_train,names = loadData(650,0) # Load all data in one batch, 3 since sample data has only 3
# X_train, X_test, y_train, y_test = train_test_split(X, Y, test_size=0., random_state=42) # Split data into training and testing
callbacks_list = [checkpoint,lrd]
optimizer = Adam(lr=0.0009, beta_1=0.9, beta_2=0.999, epsilon=None, decay=0.0, amsgrad=True) # Optimizer used to train
model.compile(optimizer=optimizer, loss={"cat_kash":"binary_crossentropy","reg_kash":lossReg}, metrics={"cat_kash":'accuracy',"reg_kash":"mse"}) # Compile model for training
K.set_learning_phase(0)
xx_tr=[]
yy_reg_tr=[]
yy_cat_tr=[]
images_aug_tr, keypoints_aug_tr = SeqAug(images=X_train,keypoints=y_train)
tar_train_reg,tar_train_cat=key2Target(keypoints_aug_tr,names)
for ind,im in enumerate(images_aug_tr):
im=im[1000:1500,:,:] # Crop out only test area
if(useTransferLearning):
xx_tr.append(preprocess_input(im))
else:
im = im/255.0
im = np.array(im,dtype=np.float32)
yuv_im = cv2.cvtColor(im, cv2.COLOR_RGB2YCrCb)
xx_tr.append(yuv_im)
for ii in tar_train_reg:
yy_reg_tr.append(ii)
for ind,ii in enumerate(tar_train_cat):
if False: # Set to true to save train images augmentated
images_aug_tr[ind] = cv2.putText(images_aug_tr[ind],str(tar_train_cat[ind]),(0,20), font, 0.5,(255,0,0),2,cv2.LINE_AA)
images_aug_tr[ind] = cv2.circle(images_aug_tr[ind],(50,int(keypoints_aug_tr[ind][0][1])),5, (0,0,255), 5)
try:
images_aug_tr[ind] = cv2.circle(images_aug_tr[ind],(50,int(keypoints_aug_tr[ind][1][1])),5, (255,0,0), 5)
except:
pass
images_aug_tr[ind] = cv2.cvtColor(images_aug_tr[ind], cv2.COLOR_RGB2BGR)
cv2.imwrite("./sample/train/"+str(ind)+".jpg",images_aug_tr[ind][1000:1500,:,:])
yy_cat_tr.append(ii)
xxx=np.array(xx_tr)
yyy_reg=np.array(yy_reg_tr)
yyy_cat=np.array(yy_cat_tr)
predictions=model.predict(xxx)
y_test = yyy_cat[:,0]
y_score = predictions[0][:,0]
precision, recall, _ = precision_recall_curve(y_test, y_score)
average_precision1 = average_precision_score(y_test, y_score)
# In matplotlib < 1.5, plt.fill_between does not have a 'step' argument
step_kwargs = ({'step': 'post'}
if 'step' in signature(plt.fill_between).parameters
else {})
plt.step(recall, precision, color='b', alpha=0.2,
where='post',label="Blue line")
plt.fill_between(recall, precision, alpha=0.2, color='b', **step_kwargs)
plt.xlabel('Recall')
plt.ylabel('Precision')
plt.ylim([0.0, 1.05])
plt.xlim([0.0, 1.0])
y_test = yyy_cat[:,1]
y_score = predictions[0][:,1]
precision, recall, _ = precision_recall_curve(y_test, y_score)
average_precision2 = average_precision_score(y_test, y_score)
# In matplotlib < 1.5, plt.fill_between does not have a 'step' argument
step_kwargs = ({'step': 'post'}
if 'step' in signature(plt.fill_between).parameters
else {})
plt.step(recall, precision, color='r', alpha=0.2,
where='post',label="Red line")
plt.fill_between(recall, precision, alpha=0.2, color='r', **step_kwargs)
plt.xlabel('Recall')
plt.ylabel('Precision')
plt.ylim([0.0, 1.05])
plt.xlim([0.0, 1.0])
print(average_precision2)
plt.title('2-class Precision-Recall curve: AP for blue={0:0.2f} AP for red={1:0.2f}'.format(
average_precision1,average_precision2))
plt.legend()
print("Mean squared Error:",mean_squared_error(yyy_reg, predictions[1]))
0.0189923
0.024536721439542924
print(len(name),len(predictions[1]))
%matplotlib inline
ims_path="./obj/images"
label_path="./obj/labels"
label_path="./obj/labels"
for index,name in enumerate(names):
basename = name.replace(".txt","")
try:
img_path = os.path.join(ims_path,basename+".jpg")
img = cv2.imread(img_path)
img[0]
except:
img_path = os.path.join(ims_path,basename+".jpeg")
img = cv2.imread(img_path)
img[0]
# img = cv2.putText(images_aug_te[ind],str(tar_test_cat[ind]),(0,20), font, 0.5,(255,0,0),2,cv2.LINE_AA)
if predictions[0][index,0]>0.5:
img = cv2.circle(img,(50,int(predictions[1][index,0]*2000)),5, (255,0,0), 5)
if predictions[0][index,1]>0.5:
img = cv2.circle(img,(50,int(predictions[1][index,1]*2000)),5, (0,0,255), 5)
cv2.imwrite("./sample/test/"+basename+".jpg",img)
import flasker
import cv2
servObj = flasker.FluServer()
img = cv2.imread("D:/source/repos/audere/new_images_sarvesh/FluA/Morning/IMG_1578.jpg")
flasker.runPipeline(img,servObj)
import glob
root_dir = "D:/source/repos/audere/new_images_sarvesh/"
y_truth =[]
y_pred = []
failed_images = []
for filename in glob.iglob(root_dir + '**/*.jpg', recursive=True):
img = cv2.imread(filename)
tmp_pred=flasker.runPipeline(img,servObj)
y_pred.append(tmp_pred)
if "Negative" in filename:
y_truth.append(0)
if tmp_pred != 0:
failed_images.append(filename)
elif "FluA+B" in filename:
y_truth.append(3)
if tmp_pred != 3:
failed_images.append(filename)
elif "FluA" in filename:
y_truth.append(1)
if tmp_pred != 1:
failed_images.append(filename)
elif "FluB" in filename:
y_truth.append(2)
if tmp_pred != 2:
failed_images.append(filename)
print(filename)
```
| github_jupyter |
# Summarizing Data
- [Download the lecture notes](https://philchodrow.github.io/PIC16A/content/pd/pd_4.ipynb).
In this lecture, we'll discuss how to descriptively *summarize* data. Descriptive data summarization is one of the fundamental processes of exploratory data analysis. The `pandas` package offers us a powerful suite of tools for creating summaries.
```
import pandas as pd
import numpy as np
penguins = pd.read_csv("palmer_penguins.csv")
cols = ["Species", "Region", "Island", "Culmen Length (mm)", "Culmen Depth (mm)"]
penguins = penguins[cols]
# shorten the species name
penguins["Species"] = penguins["Species"].str.split().str.get(0)
penguins.head()
```
## Simple Aggregation
Because the columns of a data frame behave a lot like `numpy` arrays, we can use standard methods to compute summary statistics. Here are a few examples.
```
x = penguins["Culmen Length (mm)"]
x
np.sum(x) # note: NaNs are ignored by default
x.sum() # also works
x.mean(), x.std() # mean and standard deviation
(x > 40).sum() # number of penguins with culmens longer than 40 mm
```
It's also possible to aggregate the entire data frame at once, in which case `pandas` will attempt to apply the specified function to each column for which this is possible. When passing a numerical aggregation function, non-numeric columns are ignored.
```
penguins.count() # excludes NA values, works for text columns
penguins.mean() # ignores all the text columns
# a bit counterintuitive: in text columns, returns the last
# value alphabetically
penguins.max()
```
It is technically possible to aggregate across columns (rather than rows) in `pandas`; however, doing so usually violates the [*tidy data* principles](https://cran.r-project.org/web/packages/tidyr/vignettes/tidy-data.html) and is not recommended.
We've already seen `describe()`, a convenience function for calculating numerical summary statistics.
```
penguins.describe()
```
## Split-Apply-Combine
One of the fundamental tasks in exploratory data analysis is to summarize your data **by group**. In our penguins data, for example, a very natural thing to do is to compute summary statistics **by species**, or perhaps by habitat (or both!). We can contextualize this task in three stages:
1. **Split** the data data frame into pieces, one for each species.
2. **Apply** an aggregation function to each piece, yielding a single number.
3. **Combine** the results into a new data frame.
This pattern is so common that the phrase "split-apply-combine" now appears in many texts on data analysis. This phrase was originally coined by Hadley Wickham, who is famous for developing many of the modern tools for data analysis in the `R` programming language.
<figure class="image" style="width:50%">
<img src="https://jakevdp.github.io/PythonDataScienceHandbook/figures/03.08-split-apply-combine.png" alt="Left: A single dataframe is split into three pieces. Middle: The data within each piece is summed. Right: the resulting sums are combined, resulting in a new data frame with one sum for each piece.">
<figcaption><i>split-apply-combine. Image credit: Jake VanderPlas, in the Python Data Science Handbook</i></figcaption>
</figure>
Python lets us easily perform split-apply-combine operations using the `groupby()` method of data frames.
```
penguins.groupby("Species")
```
We can think of the result of `groupby()` as a special "view" of the data frame, such that any aggregation functions used will by applied to each of the individual "groups" (i.e. species). As before, numerical aggregation functions will drop text columns.
```
penguins.groupby("Species").mean()
```
We now have a pleasant summary of the mean culmen (bill) measurements for each species. It is now clear, for example, that Adelie penguins have much shorter bills than Chinstrap and Gentoo penguins.
If you only want to show summaries for certain columns, just pass those in list form as an index to the `groupby` object:
```
# note the double brackets
penguins.groupby("Species")[["Culmen Length (mm)"]].mean()
```
While it's useful to compute a single set of summary statistics like this, it's often more useful to apply multiple aggregation functions simultaneously. The `aggregate()` method allows us to pass multiple functions, all of which will be applied and represented as new columns. For example, a common format for measurements is the mean $\pm$ the standard deviation. We can easily compute both quantities simultaneously, per penguin species:
```
penguins.groupby("Species").aggregate([np.mean, np.std])
```
It's also possible to group by multiple columns -- just pass a list of column names to `groupby`:
```
summary = penguins.groupby(["Species", "Island"]).aggregate([np.mean, np.std])
summary
```
## Hierarchical Indexing
Complex data summary tables like the one above are useful and powerful, but they also pose an important problem -- how can we extract the data from these summaries? For example, how can I get the mean bill length for Chinstrap penguins on Dream island? To extract this kind of data, we need to use *hierarchical indexing*, in which we pass multiple keys to the `.loc` attribute. After passing all the row indices, we need to use `.loc` again to get at the column indices.
```
chinstrap_dream = summary.loc["Chinstrap", "Dream"]
chinstrap_dream
# mean culmen length of chinstrap penguins on Dream Island
chinstrap_dream.loc["Culmen Length (mm)", "mean"]
```
| github_jupyter |
```
import os
import numpy as np
import pyvista as pv
from morphomatics.geom import Surface
from morphomatics.manifold import FundamentalCoords, PointDistributionModel, util
from morphomatics.stats import StatisticalShapeModel
import torch
import torch.nn as nn
os.environ["KMP_DUPLICATE_LIB_OK"]="TRUE"
# max number of objects (for debugging and dev)
nObjects=1000
dataPath = 'adni_hippos_hackathon/'
# load all data
meshes = []
labels = []
for (dirpath, dirnames, filenames) in os.walk(dataPath+"AD"):
for file in filenames:
if file[-3:] == "obj" and nObjects > len(meshes):
path = os.sep.join([dirpath, file])
mesh = pv.read(path)
meshes.append(mesh)
labels.append(0)
for (dirpath, dirnames, filenames) in os.walk(dataPath+"CN"):
for file in filenames:
if file[-3:] == "obj" and nObjects > len(meshes):
path = os.sep.join([dirpath, file])
load = mesh_straight = pv.read(path)
meshes.append(load)
labels.append(1)
# to Surface type
as_surface = lambda mesh: Surface(mesh.points, mesh.faces.reshape(-1, 4)[:, 1:])
surfaces = [as_surface(m) for m in meshes]
# construct model
SSM = StatisticalShapeModel(lambda ref: FundamentalCoords(ref)) # replace me with PointDistributionModel
SSM.construct(surfaces)
coeffs = SSM.coeffs
#print shape
print("Coeffs Shape: " + str(len(coeffs)) + " " + str(len(coeffs[0])))
coeffs = np.array(coeffs)
coeffs = torch.from_numpy(coeffs).float()
labels = torch.from_numpy(np.array(labels)).float()
x_test = coeffs[54:60]
y_test = labels[54:60]
x_train = coeffs[:54]
y_train = labels[:54]
x_train_norm = coeffs[60:114]
y_train_norm = labels[60:114]
x_test_norm = coeffs[114:]
y_test_norm = labels[114:]
print(y_train_norm)
from FrEIA.framework import InputNode, OutputNode, Node, ReversibleGraphNet, ConditionNode
from FrEIA.modules import GLOWCouplingBlock
device = 'cuda'
DIMENSION = 119
neural_net = nn.Sequential(nn.Linear(DIMENSION,256),nn.ReLU(),
nn.Linear(256,256),nn.ReLU(),
nn.Linear(256,1)).to(device)
neural_net2 = nn.Sequential(nn.Linear(DIMENSION,256),nn.ReLU(),
nn.Linear(256,256),nn.ReLU(),
nn.Linear(256,1)).to(device)
optimizer = torch.optim.Adam(neural_net.parameters(), lr = 1e-3)
optimizer2 = torch.optim.Adam(neural_net2.parameters(), lr = 1e-3)
batch_size = 16
def create_INN(num_layers, sub_net_size,dimension=119):
def subnet_fc(c_in, c_out):
return nn.Sequential(nn.Linear(c_in, sub_net_size), nn.ReLU(),
nn.Linear(sub_net_size, sub_net_size), nn.ReLU(),
nn.Linear(sub_net_size, c_out))
nodes = [InputNode(dimension, name='input')]
for k in range(num_layers):
nodes.append(Node(nodes[-1],
GLOWCouplingBlock,
{'subnet_constructor':subnet_fc, 'clamp':1.4},
name=F'coupling_{k}'))
nodes.append(OutputNode(nodes[-1], name='output'))
model = ReversibleGraphNet(nodes, verbose=False).to(device)
return model
def langevin_step(x, stepsize, neural_net, lang_steps):
log_det = torch.zeros((x.shape[0], 1), device = device)
beta=1.
for i in range(lang_steps):
x = x.requires_grad_(True)
eta = torch.randn_like(x, device = device)
out = neural_net(x)
grad_x = torch.autograd.grad(out.sum(), x,create_graph=True)[0]
y = x - stepsize * grad_x + np.sqrt(2*stepsize/beta) * eta
x = y
return x
def train(x_train,neural_net, optimizer):
perm = torch.randperm(len(x_train))[:batch_size]
xs = x_train[perm].to(device)
optimizer.zero_grad()
loss = torch.mean(neural_net(xs))
mcmc_samples = langevin_step(torch.randn(batch_size,DIMENSION, device = device), 1e-4, neural_net, lang_steps = 300)
loss += -1*torch.mean(neural_net(mcmc_samples))
loss.backward()
optimizer.step()
return loss
def val_step():
xs = x_val.to(device)
loss = torch.mean(neural_net(xs))
mcmc_samples = langevin_step(torch.randn(len(xs),DIMENSION, device = device), 1e-4, neural_net, lang_steps = 300)
loss += -1*torch.mean(neural_net(mcmc_samples))
return loss
for i in range(1000):
loss = train(x_train,neural_net, optimizer)
print(loss)
for i in range(1000):
loss = train(x_train_norm,neural_net2, optimizer2)
print(loss)
def estimate_norm_constant(neural_net):
INN = create_INN(4,256)
opti_INN = torch.optim.Adam(INN.parameters(),lr = 1e-3)
for i in range(2000):
z = torch.randn(32, DIMENSION, device = device)
out, jac = INN(z)
loss = torch.mean(neural_net(out))-torch.mean(jac)
opti_INN.zero_grad()
loss.backward()
opti_INN.step()
z = torch.randn(64, DIMENSION, device = device)
out, jac = INN(z)
loss = torch.mean(neural_net(out))- torch.mean(jac)
return loss
norm1 = estimate_norm_constant(neural_net)
norm2 = estimate_norm_constant(neural_net2)
print(norm1)
print(norm2)
probs_alz = neural_net(x_test.to(device))+norm1
probs_alz2 = neural_net2(x_test.to(device))+norm2
print(probs_alz)
print(probs_alz2)
probs1 = neural_net(x_test_norm.to(device))+norm1
probs2 = neural_net2(x_test_norm.to(device))+norm2
print(probs1)
print(probs2)
```
| github_jupyter |
# Artificial and synthetic datasets
In this notebook, instead of using and synthesizing *real* datasets, we first create *artifical* datasets and then analyze the outputs. This way, we can engineer some required features in the artifical dataset.
```
# solve issue with autocomplete
%config Completer.use_jedi = False
%load_ext autoreload
%autoreload 2
%matplotlib inline
from warnings import simplefilter
# ignore all future warnings
simplefilter(action='ignore', category=FutureWarning)
import copy
from datetime import datetime
from joblib import load as jload
import matplotlib.pyplot as plt
import matplotlib.pylab as pl
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import GradientBoostingClassifier
from privgem import tabular_ppgm
from privgem import tabular_patectgan
from privgem import tabular_metrics
from privgem import tabular_utils
from privgem import tabular_artificial
from privgem import rbo_metric
from utils import train_save_pate_models
# For reproducibility
np.random.seed(1364)
```
## Create an artificial data
```
n_samples = 10000
# Classes
n_classes = 2
class_weights = [0.5, 0.5]
n_clusters_per_class = 1
# Features
n_features=5
n_informative=5
n_redundant=0
n_repeated=0
# Control "noise"
flip_y=0.1
class_sep=1.0
# number of categorical columns and their bins
n_categorical=5
n_categorical_bins=[5, 5, 5, 5, 5]
X, y, categories = \
tabular_artificial.make_table(n_samples=n_samples,
n_classes=n_classes,
class_weights=class_weights,
n_clusters_per_class=n_clusters_per_class,
n_features=n_features,
n_informative=n_informative,
n_redundant=n_redundant,
n_repeated=n_repeated,
n_categorical=n_categorical,
n_categorical_bins=n_categorical_bins,
flip_y=flip_y,
class_sep=class_sep)
```
## Utility of original/artifical dataset
```
# extract numerical and categorical columns
num_columns, cat_columns = tabular_utils.extract_col_names_by_type(X)
orig_corr_matrix = tabular_metrics.compute_associations(X, cat_columns)
# create a pipeline
custom_pipe = tabular_metrics.create_pipeline(num_columns, cat_columns,
categories=categories,
inp_classifer=RandomForestClassifier())
#inp_classifer=GradientBoostingClassifier())
# split the data
test_size=0.3
X_train, X_test, y_train, y_test = \
train_test_split(X, y, test_size=test_size, shuffle=True)
X_train.head()
f1_orig, auc_orig, roc_auc_orig, f_orig_built, f_orig_perm, f_orig_shap = \
tabular_metrics.performance_classification(X_train, y_train,
X_test, y_test,
model_imp=custom_pipe,
pipe_classifier_name="classifier")
print(f"F1: {f1_orig:.3f}\n"\
f"AUC: {auc_orig:.3f}\n"\
f"ROC-AUC: {roc_auc_orig:.3f}\n"\
f"Features (built): {f_orig_built}\n"\
f"Features (perm) : {f_orig_perm}\n"\
f"Features (shap) : {f_orig_shap}")
# Sort the features
sorted_f_orig_built, _ = \
tabular_utils.sort_feature_vector(f_orig_built, X_train.columns.to_list())
sorted_f_orig_perm, _ = \
tabular_utils.sort_feature_vector(f_orig_perm, X_train.columns.to_list())
sorted_f_orig_shap, sorted_f_orig_shap_val = \
tabular_utils.sort_feature_vector(f_orig_shap, X_train.columns.to_list())
```
## Shuffle columns, independently
```
def shuffle(df):
df_shuffled = copy.deepcopy(df)
for indx in range(df_shuffled.shape[1]):
df_shuffled.iloc[:, indx] = np.random.permutation(df_shuffled.iloc[:, indx])
return df_shuffled
X_train_shuffled = shuffle(X_train)
X_test_shuffled = shuffle(X_test)
f1_shuffled, auc_shuffled, roc_auc_shuffled, f_shuffled_built, f_shuffled_perm, f_shuffled_shap = \
tabular_metrics.performance_classification(X_train_shuffled, y_train,
X_test_shuffled, y_test,
model_imp=custom_pipe,
pipe_classifier_name="classifier")
print(f"F1: {f1_shuffled:.3f}\n"\
f"AUC: {auc_shuffled:.3f}\n"\
f"ROC-AUC: {roc_auc_shuffled:.3f}\n"\
f"Features (built): {f_shuffled_built}\n"\
f"Features (perm) : {f_shuffled_perm}\n"\
f"Features (shap) : {f_shuffled_shap}")
# Sort the features
sorted_f_shuffled_built, _ = \
tabular_utils.sort_feature_vector(f_shuffled_built, X_train.columns.to_list())
sorted_f_shuffled_perm, _ = \
tabular_utils.sort_feature_vector(f_shuffled_perm, X_train.columns.to_list())
sorted_f_shuffled_shap, sorted_f_shuffled_shap_val = \
tabular_utils.sort_feature_vector(f_shuffled_shap, X_train.columns.to_list())
sorted_f_orig_shap
sorted_f_shuffled_shap
# cosine similarity between original and shuffled dataset
cosine_sim_measure_shuffled = \
tabular_metrics.cosine_sim(f_shuffled_shap, f_orig_shap)
kl_div_measure_shuffled = tabular_metrics.kl_div(f_shuffled_shap, f_orig_shap)
# RBO
rbo_shuffled = rbo_metric(sorted_f_orig_shap, sorted_f_shuffled_shap)
```
## Choose a synthesizer
```
synthesizer_method = "pgm"
```
## Synthesize using PATE-CTGAN
```
if synthesizer_method == "pate-ctgan":
list_eps = [0.4, 1, 10]
#list_nm = [4.2e-5, 1.05e-4, 9e-4]
list_nm = [4.2e-4, 1.05e-3, 9e-3]
list_mo = [1000, 100, 100]
list_save_log = [
"./pate_00_40/patectgan_training.csv",
"./pate_01_00/patectgan_training.csv",
"./pate_10_00/patectgan_training.csv",
]
list_save_model = [
"./pate_00_40/model.pkl",
"./pate_01_00/model.pkl",
"./pate_10_00/model.pkl",
]
batch_size = 64
device = "default" # or "default" or "cpu" or "cuda:1"
discrete_columns = cat_columns + ["label"]
if synthesizer_method == "pate-ctgan":
# prepare data
Xy = X_train.copy()
Xy["label"] = y_train
Xy[cat_columns] = Xy[cat_columns].astype("int")
if synthesizer_method == "pate-ctgan":
from parhugin import multiFunc
myproc = multiFunc(num_req_p=4)
for i in range(len(list_eps)):
myproc.add_job(target_func=train_save_pate_models,
target_args=(Xy,
discrete_columns,
list_eps[i],
batch_size,
list_nm[i],
list_mo[i],
list_save_log[i],
device,
list_save_model[i])
)
if synthesizer_method == "pate-ctgan":
myproc.run_jobs()
if synthesizer_method == "pate-ctgan":
list_models = []
for i in range(len(list_save_model)):
list_models.append(jload(list_save_model[i]))
# plot the results
tabular_utils.plot_log_patectgan(filename=list_save_log[0])
```
## Synthesize using PGM
```
if synthesizer_method == "pgm":
num_iters = 5000
list_eps = [0.005, 0.01, 0.1, 0.4, 1, 4.0, 10]
#num_iters = 100
#list_eps = [0.005, 1, 10]
delta = 1e-5
```
## Synthesize the artificial data
```
t1 = datetime.now()
target_var = "label"
rbo_p = 0.6
# Number of runs per epsilon
ensemble_runs = 1
# prepare data
Xy = X_train.copy()
Xy["label"] = y_train
Xy[cat_columns] = Xy[cat_columns].astype("int")
# --- collect results of ensemble runs
ensemble_roc_auc = {}
ensemble_f1 = {}
ensemble_cosine = {}
ensemble_rbo = {}
ensemble_rbo_corr = {}
ensemble_kl = {}
ensemble_syn_features = {}
for eps in list_eps:
print(f"--- EPS: {eps}")
for er in range(ensemble_runs):
t_eps_1 = datetime.now()
if synthesizer_method == "pgm":
# train a PGM model
pgm = tabular_ppgm(target_variable=target_var,
target_epsilon=eps,
target_delta=delta)
pgm.train(Xy, iters=num_iters)
# generate synthetic output
if synthesizer_method == "pgm":
synth_pd = pgm.generate(num_rows=len(Xy))
elif synthesizer_method == "pate-ctgan":
synth_pd = list_models[i].sample(len(Xy))
# utility of synthetic data
Xsyn_train = synth_pd.drop(columns=[target_var]).astype("str")
ysyn_train = synth_pd[target_var].to_list()
f1_tmp, auc_tmp, roc_auc_tmp, f_syn_built, f_syn_perm, f_syn_shap = \
tabular_metrics.performance_classification(Xsyn_train, ysyn_train,
X_test, y_test,
model_imp=custom_pipe,
pipe_classifier_name="classifier")
# Sort the features
sorted_f_syn_built, _ = \
tabular_utils.sort_feature_vector(f_syn_built, X_train.columns.to_list())
sorted_f_syn_perm, _ = \
tabular_utils.sort_feature_vector(f_syn_perm, X_train.columns.to_list())
sorted_f_syn_shap, sorted_f_syn_shap_val = \
tabular_utils.sort_feature_vector(f_syn_shap, X_train.columns.to_list())
# cosine similarity between original and synthetic dataset
cosine_sim_measure = \
tabular_metrics.cosine_sim(f_syn_shap, f_orig_shap)
kl_div_measure = tabular_metrics.kl_div(f_syn_shap, f_orig_shap)
# RBO
rbo = rbo_metric(sorted_f_orig_shap, sorted_f_syn_shap)
if not eps in ensemble_roc_auc.keys():
ensemble_roc_auc[eps] = []
ensemble_f1[eps] = []
ensemble_cosine[eps] = []
ensemble_rbo[eps] = []
ensemble_rbo_corr[eps] = []
ensemble_kl[eps] = []
ensemble_syn_features[eps] = []
# collect results
ensemble_roc_auc[eps].append(roc_auc_tmp)
ensemble_f1[eps].append(f1_tmp)
ensemble_cosine[eps].append(cosine_sim_measure)
ensemble_rbo[eps].append(rbo.rbo(p=rbo_p))
ensemble_rbo_corr[eps].append(
rbo.correlated_rank_similarity(p=rbo_p,
correlation_matrix=orig_corr_matrix))
ensemble_kl[eps].append(kl_div_measure)
ensemble_syn_features[eps].append(f_syn_shap)
print(f"Time for this iteration, {er}, eps: {eps}: {datetime.now() - t_eps_1}")
print(f"Total time: {datetime.now() - t1}")
```
## Plot the results
```
%matplotlib inline
plt.figure(figsize=(7, 5))
mean2plot = []
std2plot = []
for one_eps in list_eps:
mean2plot.append(np.mean(ensemble_roc_auc[one_eps]))
std2plot.append(np.std(ensemble_roc_auc[one_eps]))
plt.errorbar(list_eps, mean2plot, std2plot,
capsize=5, elinewidth=2,
lw=3, marker="o", c="k")
plt.axhline(roc_auc_orig,
ls="--", c="r",
label="original")
plt.axhline(roc_auc_shuffled,
ls="--", c="blue",
label="shuffled")
plt.xlabel("$\epsilon$", size=20)
plt.ylabel("ROC-AUC", size=20)
plt.xscale("log")
plt.xticks(size=16)
plt.yticks(size=16)
plt.legend(loc='center left',
bbox_to_anchor=(1, 0.5),
fontsize=16)
plt.grid()
plt.show()
%matplotlib inline
plt.figure(figsize=(7, 5))
mean2plot = []
std2plot = []
for one_eps in list_eps:
mean2plot.append(np.mean(ensemble_f1[one_eps]))
std2plot.append(np.std(ensemble_f1[one_eps]))
plt.errorbar(list_eps, mean2plot, std2plot,
capsize=5, elinewidth=2,
lw=3, marker="o", c="k")
plt.axhline(f1_orig,
ls="--", c="r",
label="original")
plt.axhline(f1_shuffled,
ls="--", c="blue",
label="shuffled")
plt.xlabel("$\epsilon$", size=20)
plt.ylabel("F1", size=20)
plt.xscale("log")
plt.xticks(size=16)
plt.yticks(size=16)
plt.legend(loc='center left',
bbox_to_anchor=(1, 0.5),
fontsize=16)
plt.grid()
plt.show()
%matplotlib inline
plt.figure(figsize=(7, 5))
# --- RBO
mean2plot = []
std2plot = []
for one_eps in list_eps:
mean2plot.append(np.mean(ensemble_rbo[one_eps]))
std2plot.append(np.std(ensemble_rbo[one_eps]))
plt.errorbar(list_eps, mean2plot, std2plot,
lw=3, marker="o", c="k",
capsize=5, elinewidth=2,
label="RBO")
# --- RBO corr
mean2plot = []
std2plot = []
for one_eps in list_eps:
mean2plot.append(np.mean(ensemble_rbo_corr[one_eps]))
std2plot.append(np.std(ensemble_rbo_corr[one_eps]))
plt.errorbar(np.array(list_eps), mean2plot, std2plot,
lw=3, marker="o", c="gray", ls="--",
capsize=5, elinewidth=2,
label="RBOcorr", zorder=100)
plt.axhline(1,
ls="--", c="r",
label="skyline")
plt.axhline(rbo_shuffled.rbo(p=rbo_p),
ls="--", c="blue",
label="RBO, shuffled")
plt.xlabel("$\epsilon$", size=20)
plt.ylabel("RBO score", size=20)
plt.xscale("log")
plt.xticks(size=16)
plt.yticks(size=16)
plt.legend(loc='center left',
bbox_to_anchor=(1, 0.5),
fontsize=16)
plt.grid()
plt.show()
# print(sorted_f_syn_shap)
# print(sorted_f_orig_shap)
# f_orig_shap, f_shuffled_shap
# from scipy import spatial
# 1. - spatial.distance.cosine(f_orig_shap, f_shuffled_shap)
plt.figure(figsize=(10, 10))
plt.subplot(2, 1, 1)
# original features
plt.plot(f_orig_shap, c="r",
lw=2, ls="--", marker="o",
label="original")
# shuffled features
plt.plot(f_shuffled_shap, c="b",
lw=2, ls="--", marker="o",
label="shuffled")
# synthetic features
#colors = pl.cm.viridis_r(np.linspace(0.0,1,len(list_eps)))
colors = pl.cm.viridis_r(np.linspace(0.05,1,len(list_eps)))
for i, one_eps in enumerate(list_eps):
mean2plot = np.mean(ensemble_syn_features[one_eps], axis=0)
std2plot = np.std(ensemble_syn_features[one_eps], axis=0)
plt.plot(range(len(mean2plot)), mean2plot,
c=colors[i], lw=1.,
marker="o", label=f"$\epsilon$:{one_eps}")
plt.xlabel("Features", size=20)
plt.ylabel("Score", size=20)
list_features = X_train.columns.to_list()
plt.xticks(range(len(list_features)), list_features,
size=16)
plt.yticks(size=16)
plt.legend(loc='center left',
bbox_to_anchor=(1, 0.5),
fontsize=16)
plt.grid()
plt.subplot(2, 1, 2)
# original features
plt.plot(f_orig_shap, c="r",
lw=2, ls="--", marker="o",
label="original")
# shuffled features
plt.plot(f_shuffled_shap, c="b",
lw=2, ls="--", marker="o",
label="shuffled")
# synthetic features
#colors = pl.cm.viridis_r(np.linspace(0.0,1,len(list_eps)))
colors = pl.cm.viridis_r(np.linspace(0.05,1,len(list_eps)))
for i, one_eps in enumerate(list_eps):
mean2plot = np.mean(ensemble_syn_features[one_eps], axis=0)
std2plot = np.std(ensemble_syn_features[one_eps], axis=0)
plt.errorbar(range(len(mean2plot)), mean2plot, std2plot,
c=colors[i], lw=1.,
capsize=5, elinewidth=2,
marker="o", label=f"$\epsilon$:{one_eps}")
plt.xlabel("Features", size=20)
plt.ylabel("Score", size=20)
list_features = X_train.columns.to_list()
plt.xticks(range(len(list_features)), list_features,
size=16)
plt.yticks(size=16)
plt.legend(loc='center left',
bbox_to_anchor=(1, 0.5),
fontsize=16)
plt.grid()
plt.show()
%matplotlib inline
plt.figure(figsize=(7, 5))
mean2plot = []
std2plot = []
for one_eps in list_eps:
mean2plot.append(np.mean(ensemble_cosine[one_eps]))
std2plot.append(np.std(ensemble_cosine[one_eps]))
plt.errorbar(list_eps, mean2plot, std2plot,
capsize=5, elinewidth=2,
lw=3, marker="o", c="k")
plt.axhline(1,
ls="--", c="r",
label="original")
plt.axhline(cosine_sim_measure_shuffled,
ls="--", c="blue",
label="shuffled")
plt.xlabel("$\epsilon$", size=20)
plt.ylabel("Cosine sim", size=20)
plt.xscale("log")
plt.xticks(size=16)
plt.yticks(size=16)
plt.legend(loc='center left',
bbox_to_anchor=(1, 0.5),
fontsize=16)
plt.grid()
plt.show()
np.array(ensemble_kl[0.005])
%matplotlib inline
plt.figure(figsize=(7, 5))
mean2plot = []
std2plot = []
for one_eps in list_eps:
ensemble_kl_arr = np.array(ensemble_kl[one_eps])
ensemble_kl_arr[ensemble_kl_arr == np.inf] = 200
mean2plot.append(np.mean(ensemble_kl_arr))
std2plot.append(np.std(ensemble_kl_arr))
plt.errorbar(list_eps, mean2plot, std2plot,
capsize=5, elinewidth=2,
lw=3, marker="o", c="k")
plt.axhline(0,
ls="--", c="r",
label="original")
plt.axhline(kl_div_measure_shuffled,
ls="--", c="blue",
label="shuffled")
plt.xlabel("$\epsilon$", size=20)
plt.ylabel("KL-divergence", size=20)
plt.xscale("log")
plt.xticks(size=16)
plt.yticks(size=16)
plt.legend(loc='center left',
bbox_to_anchor=(1, 0.5),
fontsize=16)
plt.grid()
plt.show()
```
## Why cosine sim. between `f_shuffled_shap` and `f_orig_shap` is so high?
```
# Range of values for elements in random vectors
min_val = 0
max_val = 1000
# Repetition
num_iter = 1000000
# list_dims = [ 2, 3, 4, 5, 6, 7, 8, 9, 10,
# 20, 30, 40, 50, 60, 70, 80, 90, 100,
# 200, 300, 400, 500, 600, 700, 800, 900, 1000]
list_dims = [2] + list(range(5, 105, 5))
# --- list to collect results
dims = []
# cosine
cs_means = []
cs_stds = []
# L2
l2_means = []
l2_stds = []
# j specifies the dimension of random vectors
for j in list_dims:
print(j, end=" ")
repetitions = range(num_iter)
tmp_cs_dists = []
tmp_l2_dists = []
x_used = []
for i in repetitions:
v1 = np.random.uniform(min_val, max_val, j)
# Weight some components?
# v1[:int(len(v1)/2)] *= 10
# v2 is a permutation of v1
v2 = np.random.permutation(v1)
# v2 = np.random.uniform(min_val, max_val, j)
cs_sim = tabular_metrics.cosine_sim(v1, v2)
l2_dist = tabular_metrics.L2_norm_dist(v1, v2)
if isinstance(cs_sim, float) and isinstance(l2_dist, float):
tmp_cs_dists.append(cs_sim)
tmp_l2_dists.append(l2_dist)
x_used.append(i)
tmp_cs_dists = np.array(tmp_cs_dists)
tmp_l2_dists = np.array(tmp_l2_dists)
tmp_noninf_cs_dists = tmp_cs_dists[tmp_cs_dists != np.inf]
curr_cs_mean = np.mean(tmp_noninf_cs_dists)
curr_cs_std = np.std(tmp_noninf_cs_dists)
tmp_noninf_l2_dists = tmp_l2_dists[tmp_l2_dists != np.inf]
curr_l2_mean = np.mean(tmp_noninf_l2_dists)
curr_l2_std = np.std(tmp_noninf_l2_dists)
dims.append(j)
cs_means.append(curr_cs_mean)
cs_stds.append(curr_cs_std)
l2_means.append(curr_l2_mean)
l2_stds.append(curr_l2_std)
#plt.scatter(x_act, dists, alpha=0.01, s=10)
%matplotlib inline
plt.figure(figsize=(7, 5))
plt.errorbar(dims, cs_means, cs_stds,
c="k", linestyle='None', marker='o')
plt.xlabel("Dimension", size=20)
plt.ylabel("Cosine sim", size=20)
plt.xticks(size=16)
plt.yticks(size=16)
plt.ylim(0, 1)
plt.xlim(0, 100)
# plt.xscale("log")
plt.grid()
plt.show()
plt.scatter(x_used, tmp_cs_dists, alpha=0.01, c='k')
plt.ylim(0, 1)
```
| github_jupyter |
### (1) 物体検出の分野にはどういった手法が存在したか。
-> Selective serach, CPMC, MCG, sliding window
Widely used object proposal methods include those based on grouping super-pixels (e.g., Selective Search, CPMC, MCG) and those based on sliding windows (e.g., objectness in windows, EdgeBoxes).
### (2) Fasterとあるが、どういった仕組みで高速化したのか。
-> RPNをconvolutional networkで構築した
Our observation is that the convolutional feature maps used by region-based detectors, like Fast R-CNN, can also be used for generating region proposals. On top of these convolutional features, we construct an RPN by adding a few additional convolutional layers that simultaneously regress region bounds adn objectness scores at each location on a regular grid. The RPN is thus a kind of fully convolutional network and can be trained ene_to_end specifically for the task for generating detection proposals.
### (3) One-Stageの手法とTwo-Stageの手法はどう違うのか。
-> object proposalsの前にpreprocessingがあるかないか
a. Two stage detection frameworks, which include a preprocessing step for generating object proposals;
b. One stage detection frameworks, or region proposal free frame-works, having a single proposed method which does not separate the process of the detection proposal.
(https://arxiv.org/pdf/1809.02165.pdf)
### (4) RPNとは何か。
-> object boundingとobjectness scoreを生成するところ
An RPN is a fully convolutional network that simultaneously predicts object bounds and objectness scores at each position.
### (5) RoIプーリングとは何か。
-> RoIを固定の形に変えるためのPooling層
The RoI pooling layer uses max pooling to convert the features inside any valid region of interest into a small fea- ture map with a fixed spatial extent of H × W (e.g., 7 × 7), where H and W are layer hyper-parameters that are inde- pendent of any particular RoI. (https://www.cv-foundation.org/openaccess/content_iccv_2015/papers/Girshick_Fast_R-CNN_ICCV_2015_paper.pdf)
### (6) Anchorのサイズはどうするのが適切か。
-> いろいろなscaleとaspect ratioのものを使用すると良い
We show by experiments the effects of this scheme for addressing multiple scales and sizes (Table 8).
Because of this multi-scale design based on anchors, we can simply use the convolutional features computed on a single-scale image, as is also done by the Fast R-CNN detector. The design of multi-scale anchors is a key component for sharing features without extra cost for addressing scales.
### (7) 何というデータセットを使い、先行研究に比べどういった指標値が得られているか。
-> PASCAL VOC test setで評価を行った。Selective Searchと比較して計算コストが減少し、精度も向上した。
SS has an mAP of 58.7% and EB has an mAP of 58.6% under the Fast R-CNN framework. RPN with Fast R-CNN achieves competitive results, with
8 an mAP of 59.9% while using up to 300 proposals .
Using RPN yields a much faster detection system than using either SS or EB because of shared convolutional computations; the fewer proposals also reduce the region-wise fully-connected layers’ cost (Table 5).
| github_jupyter |
```
## Get dependencies ##
import string
import math
import sys
import matplotlib.pyplot as plt
import matplotlib
import seaborn as sn
sys.path.append('..')
from GIR import *
import scipy as sp
import pickle
import time
import scipy as sp
from scipy import ndimage
from scipy import signal
import os
import statsmodels.api as sm
from mpl_toolkits.axes_grid1.inset_locator import inset_axes
import glob
import requests
import ftplib
import PyPDF2
import io
import cmocean
import multiprocessing
import xarray as xr
import numpy as np
import pandas as pd
import requests
import xml.etree.ElementTree as ET
import zarr
import gcsfs
import multiprocessing
def esgf_search(server="https://esgf-node.llnl.gov/esg-search/search",
files_type="OPENDAP", local_node=True, project="CMIP6",
verbose=False,url_verbose=False, format="application%2Fsolr%2Bjson",
use_csrf=False, **search):
client = requests.session()
payload = search
payload["project"] = project
payload["type"]= "File"
if local_node:
payload["distrib"] = "false"
if use_csrf:
client.get(server)
if 'csrftoken' in client.cookies:
# Django 1.6 and up
csrftoken = client.cookies['csrftoken']
else:
# older versions
csrftoken = client.cookies['csrf']
payload["csrfmiddlewaretoken"] = csrftoken
payload["format"] = format
offset = 0
numFound = 10000
all_files = []
files_type = files_type.upper()
while offset < numFound:
payload["offset"] = offset
url_keys = []
for k in payload:
url_keys += ["{}={}".format(k, payload[k])]
url = "{}/?{}".format(server, "&".join(url_keys))
if url_verbose:
print(url)
r = client.get(url)
r.raise_for_status()
resp = r.json()["response"]
numFound = int(resp["numFound"])
resp = resp["docs"]
offset += len(resp)
for d in resp:
if verbose:
for k in d:
print("{}: {}".format(k,d[k]))
url = d["url"]
for f in d["url"]:
sp = f.split("|")
if sp[-1] == files_type:
all_files.append(sp[0].split(".html")[0])
return sorted(all_files)
def get_annual_CMIP6_data_esgf(activity, table, variable, experiment, institution, source, member):
# eg activity='CMIP', table='Amon', variable='tas', experiment='historical', institution="NCAR", source="CESM2", member="r10i1p1f1"
result = esgf_search(activity_id=activity, table_id=table, variable_id=variable, experiment_id=experiment,institution_id=institution, source_id=source, member_id=member)
if not result:
print('No results for this request')
return None
# select results with only the latest datestamp:
latest = sorted([x.split('/')[15] for x in result])[-1]
result = [x for x in result if x.split('/')[15]==latest]
# remove duplicate results
result_1 = []
for item in result:
if item.split('/')[-1] in [x.split('/')[-1] for x in result_1]:
continue
else:
result_1 += [item]
ds = xr.open_mfdataset(result_1, combine='by_coords')
files_area = esgf_search(variable_id='areacella', activity_id=activity, institution_id=institution, source_id=source)
if not files_area:
print('No areacella for this request')
return None
ds_area = xr.open_dataset(files_area[0])
coords = list(ds[variable].coords.keys())
if 'lat' in coords:
dims = ['lat','lon']
else:
dims = ['latitude','longitude']
total_area = ds_area.areacella.sum(dim=dims)
ta_timeseries = (ds[variable] * ds_area.areacella).sum(dim=dims) / total_area
return ta_timeseries.groupby('time.year').mean('time').to_pandas().rename(institution+'_'+source+'_'+member)
gs_stores = pd.read_csv('gs://cmip6/cmip6-zarr-consolidated-stores.csv')
gcs = gcsfs.GCSFileSystem(token='anon')
gs_stores.loc[:,'ism'] = gs_stores.loc[:,'institution_id'] + '_' + gs_stores.loc[:,'source_id'] + '_' + gs_stores.loc[:,'member_id']
def get_annual_CMIP6_data_gstore(activity, table, variable, experiment, institution, source, member):
# eg activity='CMIP', table='Amon', variable='tas', experiment='historical', institution="NCAR", source="CESM2", member="r10i1p1f1"
try:
query = gs_stores.query("activity_id==\'"+activity+"\' & table_id==\'"+table+"\' & variable_id==\'"+variable+"\' & experiment_id==\'"+experiment+"\' & institution_id==\'"+institution+"\' & source_id==\'"+source+"\' & member_id==\'"+member+"\'")
if query.empty:
print('No results for this request')
return None
# create a mutable-mapping-style interface to the store
mapper = gcs.get_mapper(query.zstore.values[0])
# open it using xarray and zarr
ds = xr.open_zarr(mapper, consolidated=True)
dims = list(ds[variable].dims)
dims.remove('time')
lat_dim_name = [x for x in dims if 'lat' in x][0]
weights = np.cos(np.deg2rad(ds[variable][lat_dim_name]))
weights.name = "weights"
ta_timeseries = ds[variable].weighted(weights).mean(dims)
return ta_timeseries.groupby('time.year').mean('time').to_pandas().rename(institution+'_'+source+'_'+member+'_'+variable+'_'+experiment)
except:
print('retrieval failed')
return None
onepct_info = gs_stores.loc[(gs_stores.experiment_id=='1pctCO2')&(gs_stores.variable_id.isin(['tas','rlut','rsut','rsdt']))&(gs_stores.table_id=='Amon')]
onepct_data=[]
for index,row in onepct_info.iterrows():
print('getting '+row.loc['ism']+'_'+row.loc['variable_id'])
onepct_data += [get_annual_CMIP6_data_gstore(row.loc['activity_id'], row.loc['table_id'], row.loc['variable_id'], row.loc['experiment_id'], row.loc['institution_id'], row.loc['source_id'], row.loc['member_id'])]
pd.concat(onepct_data,axis=1).to_csv('./cmip6_data/onepct.csv')
abrupt_info = gs_stores.loc[(gs_stores.experiment_id=='abrupt-4xCO2')&(gs_stores.variable_id.isin(['tas','rlut','rsut','rsdt']))&(gs_stores.table_id=='Amon')]
abrupt_data=[]
for index,row in abrupt_info.iterrows():
print('getting '+row.loc['ism']+'_'+row.loc['variable_id'])
abrupt_data += [get_annual_CMIP6_data_gstore(row.loc['activity_id'], row.loc['table_id'], row.loc['variable_id'], row.loc['experiment_id'], row.loc['institution_id'], row.loc['source_id'], row.loc['member_id'])]
pd.concat(abrupt_data,axis=1).to_csv('./cmip6_data/abrupt-4xCO2.csv')
# piControl_info = gs_stores.loc[(gs_stores.experiment_id=='piControl')&(gs_stores.variable_id.isin(['tas','rlut','rsut','rsdt']))&(gs_stores.table_id=='Amon')]
# piControl_data=[]
for index,row in piControl_info.loc[26752:].iterrows():
print('getting '+row.loc['ism']+'_'+row.loc['variable_id'])
piControl_data += [get_annual_CMIP6_data_gstore(row.loc['activity_id'], row.loc['table_id'], row.loc['variable_id'], row.loc['experiment_id'], row.loc['institution_id'], row.loc['source_id'], row.loc['member_id'])]
pd.concat(piControl_data,axis=1)#.to_csv('./cmip6_data/piControl.csv')
# ssp_list = ['ssp370', 'ssp126', 'ssp245', 'ssp585', 'ssp119', 'ssp434', 'ssp460', 'ssp534-over']
# ssp_info = gs_stores.loc[(gs_stores.experiment_id.isin(ssp_list))&(gs_stores.variable_id=='tas')&(gs_stores.table_id=='Amon')]
# ssp_data=[]
for index,row in ssp_info.iterrows():
print('getting '+row.loc['ism']+'_'+row.loc['variable_id'])
ssp_data += [get_annual_CMIP6_data_gstore(row.loc['activity_id'], row.loc['table_id'], row.loc['variable_id'], row.loc['experiment_id'], row.loc['institution_id'], row.loc['source_id'], row.loc['member_id'])]
pd.concat(ssp_data,axis=1).to_csv('./cmip6_data/ssp_tas.csv')
# hist_info = gs_stores.loc[(gs_stores.experiment_id=='historical')&(gs_stores.variable_id=='tas')&(gs_stores.table_id=='Amon')]
# hist_data=[]
for index,row in hist_info.iloc[[i for i,x in enumerate(hist_data) if x is None]].iterrows():
print('getting '+row.loc['ism']+'_'+row.loc['variable_id'])
hist_data += [get_annual_CMIP6_data_gstore(row.loc['activity_id'], row.loc['table_id'], row.loc['variable_id'], row.loc['experiment_id'], row.loc['institution_id'], row.loc['source_id'], row.loc['member_id'])]
pd.concat(hist_data,axis=1).to_csv('./cmip6_data/hist_tas.csv')
def get_annual_CMIP6_data_gstore_co2(activity, table, variable, experiment, institution, source, member):
# eg activity='CMIP', table='Amon', variable='tas', experiment='historical', institution="NCAR", source="CESM2", member="r10i1p1f1"
try:
query = gs_stores.query("activity_id==\'"+activity+"\' & table_id==\'"+table+"\' & variable_id==\'"+variable+"\' & experiment_id==\'"+experiment+"\' & institution_id==\'"+institution+"\' & source_id==\'"+source+"\' & member_id==\'"+member+"\'")
if query.empty:
print('No results for this request')
return None
# create a mutable-mapping-style interface to the store
mapper = gcs.get_mapper(query.zstore.values[0])
# open it using xarray and zarr
ds = xr.open_zarr(mapper, consolidated=True)
dims = list(ds[variable].dims)
dims.remove('time')
lat_dim_name = [x for x in dims if 'lat' in x][0]
weights = np.cos(np.deg2rad(ds[variable][lat_dim_name]))
weights.name = "weights"
ta_timeseries = ds[variable].isel(plev=0).weighted(weights).mean(dims)
return ta_timeseries.groupby('time.year').mean('time').to_pandas().rename(institution+'_'+source+'_'+member+'_'+variable+'_'+experiment)
except:
print('retrieval failed')
return None
co2_info = gs_stores.loc[(gs_stores.experiment_id.isin(['abrupt-4xCO2','1pctCO2','historical','piControl']))&(gs_stores.variable_id=='co2')&(gs_stores.table_id=='Amon')]
co2_data=[]
for index,row in co2_info.iterrows():
print('getting '+row.loc['ism']+'_'+row.loc['variable_id'])
co2_data += [get_annual_CMIP6_data_gstore_co2(row.loc['activity_id'], row.loc['table_id'], row.loc['variable_id'], row.loc['experiment_id'], row.loc['institution_id'], row.loc['source_id'], row.loc['member_id'])]
pd.concat(co2_data,axis=1).to_csv('./cmip6_data/co2.csv')
def get_annual_CMIP6_data_gstore_co2mass(activity, table, variable, experiment, institution, source, member):
# eg activity='CMIP', table='Amon', variable='tas', experiment='historical', institution="NCAR", source="CESM2", member="r10i1p1f1"
try:
query = gs_stores.query("activity_id==\'"+activity+"\' & table_id==\'"+table+"\' & variable_id==\'"+variable+"\' & experiment_id==\'"+experiment+"\' & institution_id==\'"+institution+"\' & source_id==\'"+source+"\' & member_id==\'"+member+"\'")
if query.empty:
print('No results for this request')
return None
# create a mutable-mapping-style interface to the store
mapper = gcs.get_mapper(query.zstore.values[0])
# open it using xarray and zarr
ds = xr.open_zarr(mapper, consolidated=True)
return ds[variable].groupby('time.year').mean('time').to_pandas().rename(institution+'_'+source+'_'+member+'_'+variable+'_'+experiment)
except:
print('retrieval failed')
return None
co2mass_info = gs_stores.loc[(gs_stores.experiment_id.isin(['abrupt-4xCO2','1pctCO2','historical','piControl']))&(gs_stores.variable_id=='co2mass')&(gs_stores.table_id=='Amon')]
co2mass_data=[]
for index,row in co2mass_info.iterrows():
print('getting '+row.loc['ism']+'_'+row.loc['variable_id'])
co2mass_data += [get_annual_CMIP6_data_gstore_co2mass(row.loc['activity_id'], row.loc['table_id'], row.loc['variable_id'], row.loc['experiment_id'], row.loc['institution_id'], row.loc['source_id'], row.loc['member_id'])]
pd.concat(co2mass_data,axis=1).to_csv('./cmip6_data/co2mass.csv')
nbp_info = gs_stores.loc[(gs_stores.experiment_id.isin(['esm-ssp585', 'esm-hist', 'historical', 'ssp585', 'ssp245']))&(gs_stores.variable_id=='nbp')&(gs_stores.table_id=='Lmon')]
nbp_data=[]
for index,row in nbp_info.iterrows():
print('getting '+row.loc['ism']+'_'+row.loc['variable_id'])
nbp_data += [get_annual_CMIP6_data_gstore(row.loc['activity_id'], row.loc['table_id'], row.loc['variable_id'], row.loc['experiment_id'], row.loc['institution_id'], row.loc['source_id'], row.loc['member_id'])]
pd.concat(nbp_data,axis=1).to_csv('./cmip6_data/nbp.csv')
```
### Getting 1pct nbp / nep / fgco2 / tas data from esgf
```
def get_annual_CMIP6_data_esgf(activity, table, variable, experiment, institution, source, member):
# eg activity='CMIP', table='Amon', variable='tas', experiment='historical', institution="NCAR", source="CESM2", member="r10i1p1f1"
try:
result = esgf_search(activity_id=activity, table_id=table, variable_id=variable, experiment_id=experiment,institution_id=institution, source_id=source, member_id=member)
if not result:
print('No results for this request')
return None
# select results with only the latest datestamp:
latest = sorted([x.split('/')[15] for x in result])[-1]
result = [x for x in result if x.split('/')[15]==latest]
# remove duplicate results
result_1 = []
for item in result:
if item.split('/')[-1] in [x.split('/')[-1] for x in result_1]:
continue
else:
result_1 += [item]
ds = xr.open_mfdataset(result_1, combine='by_coords')
dims = list(ds[variable].dims)
dims.remove('time')
lat_dim_name = [x for x in dims if 'lat' in x][0]
weights = np.cos(np.deg2rad(ds[variable][lat_dim_name]))
weights.name = "weights"
ta_timeseries = ds[variable].weighted(weights).mean(dims)
return ta_timeseries.groupby('time.year').mean('time').to_pandas().rename(institution+'_'+source+'_'+member+'_'+variable+'_'+experiment)
except:
print('retrieval failed')
return None
pd.Series(['v20190306','v20200429']).sort_values()
search_results = esgf_search(table_id='Lmon', variable_id='nbp',experiment_id='1pctCO2-bgc')
search_df = pd.DataFrame([x.split('/')[-10:-1] for x in search_results],columns=['activity','institution','source','experiment','member','table','variable','grid','date'])
chosen_indices = []
for i,row in search_df.iterrows():
duplicates = search_df.query("activity==\'"+row.loc['activity']+"\' & table==\'"+row.loc['table']+"\' & variable==\'"+row.loc['variable']+"\' & experiment==\'"+row.loc['experiment']+"\' & institution==\'"+row.loc['institution']+"\' & source==\'"+row.loc['source']+"\' & member==\'"+row.loc['member']+"\' & grid==\'"+row.loc['grid']+"\'")
duplicate_dates = duplicates.loc[:,'date'].sort_values()
latest_date = duplicate_dates.iloc[-1]
if row.loc['date'] == latest_date:
chosen_indices += [i]
else:
continue
search_df = search_df.loc[chosen_indices].drop_duplicates()
experiments = ['1pctCO2','1pctCO2-bgc','1pctCO2-rad']
variables = ['nep','nbp','fgco2','fco2nat','tas']
tables = dict(zip(variables,['Emon','Lmon','Omon','Amon','Amon']))
search_dfs = []
for experiment in experiments:
for variable in variables:
search_results = esgf_search(table_id=tables[variable], variable_id=variable,experiment_id=experiment)
search_df = pd.DataFrame([x.split('/')[-10:-1] for x in search_results],columns=['activity','institution','source','experiment','member','table','variable','grid','date'])
chosen_indices = []
for i,row in search_df.iterrows():
duplicates = search_df.query("activity==\'"+row.loc['activity']+"\' & table==\'"+row.loc['table']+"\' & variable==\'"+row.loc['variable']+"\' & experiment==\'"+row.loc['experiment']+"\' & institution==\'"+row.loc['institution']+"\' & source==\'"+row.loc['source']+"\' & member==\'"+row.loc['member']+"\' & grid==\'"+row.loc['grid']+"\'")
duplicate_dates = duplicates.loc[:,'date'].sort_values()
latest_date = duplicate_dates.iloc[-1]
if row.loc['date'] == latest_date:
chosen_indices += [i]
else:
continue
search_df = search_df.loc[chosen_indices].drop_duplicates()
search_dfs += [search_df]
search_df = pd.concat(search_dfs,axis=0)
# filter search results by date
chosen_models = search_df.loc[(search_df.experiment.isin(['1pctCO2-bgc','1pctCO2-rad']))&(search_df.variable.isin(['fco2nat','fgco2']))].source.unique()
search_df.loc[(search_df.experiment.isin(['1pctCO2-bgc','1pctCO2-rad']))].source.unique()
# get area weights for each model
def get_model_area(source,area_var='areacella'):
try:
files_area = esgf_search(variable_id=area_var, source_id=source)
ds_area = xr.open_dataset(files_area[0])
return ds_area
except:
print('retrieval failed')
return None
model_areacella = {}
for model in chosen_models:
print('getting '+model+' areacella')
model_areacella[model] = get_model_areacella(model)
model_areacello = {}
for model in chosen_models:
print('getting '+model+' areacello')
model_areacello[model] = get_model_area(model,'areacello')
## switches:
## if gr available, get that
hist_info = gs_stores.loc[(gs_stores.experiment_id=='historical')&(gs_stores.variable_id=='tas')&(gs_stores.table_id=='Amon')]
hist_data = []
for index,row in hist_info.iterrows():
hist_data += [get_annual_CMIP6_data_gstore(row.loc['activity_id'], row.loc['table_id'], row.loc['variable_id'], row.loc['experiment_id'], row.loc['institution_id'], row.loc['source_id'], row.loc['member_id'])]
pd.concat(hist_data,axis=1).to_csv('./cmip6_data/historical_tas.csv')
def get_annual_CMIP6_data_gstore_nbp(activity, table, variable, experiment, institution, source, member):
# eg activity='CMIP', table='Amon', variable='tas', experiment='historical', institution="NCAR", source="CESM2", member="r10i1p1f1"
area_query = gs_stores.query("variable_id=='areacella' & source_id==\'"+source+"\'")
if area_query.empty:
files_area = esgf_search(variable_id='areacella', activity_id=activity, institution_id=institution, source_id=source)
if not files_area:
print('No areacella for this request')
return None
ds_area = xr.open_dataset(files_area[0])
else:
ds_area = xr.open_zarr(gcs.get_mapper(area_query.zstore.values[0]), consolidated=True)
query = gs_stores.query("activity_id==\'"+activity+"\' & table_id==\'"+table+"\' & variable_id==\'"+variable+"\' & experiment_id==\'"+experiment+"\' & institution_id==\'"+institution+"\' & source_id==\'"+source+"\' & member_id==\'"+member+"\'")
if query.empty:
print('No results for this request')
return None
# create a mutable-mapping-style interface to the store
mapper = gcs.get_mapper(query.zstore.values[0])
# open it using xarray and zarr
ds = xr.open_zarr(mapper, consolidated=True)
if source=='E3SM-1-1' and variable=='tas' and experiment=='piControl':
ds = xr.open_mfdataset(esgf_search(activity_id=activity, table_id=table, variable_id=variable, experiment_id=experiment, institution_id=institution, source_id=source, member_id=member)[7:],combine='by_coords')
coords = list(ds[variable].coords.keys())
if 'latitude' in coords:
dims = ['latitude','longitude']
_dims = ['lat','lon']
else:
dims = ['lat','lon']
_dims = ['latitude','longitude']
if not dims[0] in list(ds_area['areacella'].coords.keys()):
ds_area = ds_area.rename(dict(zip(_dims,dims)))
total_area = ds_area.areacella.sum(dim=dims)
ta_timeseries = (ds[variable] * ds_area.areacella).sum(dim=dims)
return ta_timeseries.groupby('time.year').mean('time').to_pandas().rename(institution+'_'+source+'_'+member+'_'+experiment)
nbp_cmip6=gs_stores.loc[(gs_stores.variable_id=='nbp')&(gs_stores.table_id=='Lmon')]
# nbp_data = []
# for index,row in nbp_cmip6.iterrows():
# nbp_data += [get_annual_CMIP6_data_gstore_nbp(row.loc['activity_id'], row.loc['table_id'], row.loc['variable_id'], row.loc['experiment_id'], row.loc['institution_id'], row.loc['source_id'], row.loc['member_id'])]
pd.concat(nbp_data,axis=1).to_csv('./cmip6_data/nbp.csv')
def get_annual_CMIP6_data_gstore_fgco2(activity, table, variable, experiment, institution, source, member):
# eg activity='CMIP', table='Amon', variable='tas', experiment='historical', institution="NCAR", source="CESM2", member="r10i1p1f1"
area_query = gs_stores.query("variable_id=='areacello' & source_id==\'"+source+"\'")
if area_query.empty:
files_area = esgf_search(variable_id='areacello', activity_id=activity, institution_id=institution, source_id=source)
if not files_area:
print('No areacello for this request')
return None
ds_area = xr.open_dataset(files_area[0])
else:
ds_area = xr.open_zarr(gcs.get_mapper(area_query.zstore.values[0]), consolidated=True)
query = gs_stores.query("activity_id==\'"+activity+"\' & table_id==\'"+table+"\' & variable_id==\'"+variable+"\' & experiment_id==\'"+experiment+"\' & institution_id==\'"+institution+"\' & source_id==\'"+source+"\' & member_id==\'"+member+"\'")
if query.empty:
print('No results for this request')
return None
# create a mutable-mapping-style interface to the store
mapper = gcs.get_mapper(query.zstore.values[0])
# open it using xarray and zarr
ds = xr.open_zarr(mapper, consolidated=True)
if source=='E3SM-1-1' and variable=='tas' and experiment=='piControl':
ds = xr.open_mfdataset(esgf_search(activity_id=activity, table_id=table, variable_id=variable, experiment_id=experiment, institution_id=institution, source_id=source, member_id=member)[7:],combine='by_coords')
dims = list(ds[variable].dims)
dims.remove('time')
# total_area = ds_area.areacello.sum(dim=dims)
ta_timeseries = (ds[variable] * ds_area.areacello).sum(dim=dims)
return ta_timeseries.groupby('time.year').mean('time').to_pandas().rename(institution+'_'+source+'_'+member+'_'+experiment)
gs_stores['isme'] = gs_stores['ism']+'_'+gs_stores['experiment_id']
fgco2_cmip6=gs_stores.loc[(gs_stores.variable_id=='fgco2')&(gs_stores.table_id=='Omon')&(gs_stores.grid_label=='gn')&(gs_stores.isme.isin(nbp_cmip6.isme))]
fgco2_data = []
for index,row in fgco2_cmip6.loc[237786:].iterrows():
fgco2_data += [get_annual_CMIP6_data_gstore_fgco2(row.loc['activity_id'], row.loc['table_id'], row.loc['variable_id'], row.loc['experiment_id'], row.loc['institution_id'], row.loc['source_id'], row.loc['member_id'])]
activity, table, variable, experiment, institution, source, member = row.loc['activity_id'], row.loc['table_id'], row.loc['variable_id'], row.loc['experiment_id'], row.loc['institution_id'], row.loc['source_id'], row.loc['member_id']
area_query = gs_stores.query("variable_id=='areacello' & source_id==\'"+source+"\'")
if area_query.empty:
files_area = esgf_search(variable_id='areacello', activity_id=activity, institution_id=institution, source_id=source)
if not files_area:
print('No areacello for this request')
# return None
ds_area = xr.open_dataset(files_area[0])
else:
ds_area = xr.open_zarr(gcs.get_mapper(area_query.zstore.values[0]), consolidated=True)
query = gs_stores.query("activity_id==\'"+activity+"\' & table_id==\'"+table+"\' & variable_id==\'"+variable+"\' & experiment_id==\'"+experiment+"\' & institution_id==\'"+institution+"\' & source_id==\'"+source+"\' & member_id==\'"+member+"\'")
if query.empty:
print('No results for this request')
# return None
# create a mutable-mapping-style interface to the store
mapper = gcs.get_mapper(query.zstore.values[0])
# open it using xarray and zarr
ds = xr.open_zarr(mapper, consolidated=True)
if source=='E3SM-1-1' and variable=='tas' and experiment=='piControl':
ds = xr.open_mfdataset(esgf_search(activity_id=activity, table_id=table, variable_id=variable, experiment_id=experiment, institution_id=institution, source_id=source, member_id=member)[7:],combine='by_coords')
dims = list(ds[variable].dims)
dims.remove('time')
total_area = ds_area.areacello.sum(dim=dims)
ta_timeseries = (ds[variable] * ds_area.areacello).sum(dim=dims)
pd.concat(fgco2_data,axis=1).to_csv('./cmip6_data/fgco2.csv')
abrupt_4x_ism = gs_stores.loc[(gs_stores.experiment_id=='abrupt-4xCO2')&(gs_stores.variable_id.isin(['tas','rlut','rsdt','rsut']))]
abrupt_4x_ism = list(set([x for x in abrupt_4x_ism.ism if abrupt_4x_ism.loc[abrupt_4x_ism.ism==x].shape[0]>=4]))
onepct_ism = gs_stores.loc[(gs_stores.experiment_id=='1pctCO2')&(gs_stores.variable_id.isin(['tas','rlut','rsdt','rsut']))]
onepct_ism = list(set([x for x in onepct_ism.ism if onepct_ism.loc[onepct_ism.ism==x].shape[0]>=4]))
piControl_ism = gs_stores.loc[(gs_stores.experiment_id=='piControl')&(gs_stores.variable_id=='tas')]
piControl_ism = list(set(piControl_ism.ism))
areacella_s_gs = [x.split('_')[1] for x in list(set(gs_stores.loc[(gs_stores.variable_id=='areacella')].ism))]
areacella_list = esgf_search(activity_id='CMIP', variable_id='areacella')
areacella_list_nodupl = []
for item in areacella_list:
if item.split('/')[-1] in [x.split('/')[-1] for x in areacella_list_nodupl]:
continue
else:
areacella_list_nodupl += [item]
areacella_ism_list = list(set([x.split('/')[8]+'_'+x.split('/')[9]+'_'+x.split('/')[11] for x in areacella_list_nodupl]))
areacella_s_esgf = list(set([x.split('_')[1] for x in areacella_ism_list]))
areacella_s_all = list(set(areacella_s_gs).union(areacella_s_esgf))
abrupt_4x_ism_areacella_exist = [x for x in abrupt_4x_ism if x.split('_')[1] in areacella_s_all]
onepct_ism_areacella_exist = [x for x in onepct_ism if x.split('_')[1] in areacella_s_all]
piControl_ism_areacella_exist = [x for x in piControl_ism if x.split('_')[1] in areacella_s_all]
def get_cmip6_data_gs(ism,var,exp):
print('getting '+ism+' '+var)
ism_split = ism.split('_')
_out = get_annual_CMIP6_data_gstore('CMIP', 'Amon', var, exp, ism_split[0], ism_split[1], ism_split[2])
print('got '+ism)
return _out
onepct_tas_df_list = []
for ism in onepct_ism_areacella_exist:
onepct_tas_df_list += [get_cmip6_data_gs(ism,'tas','1pctCO2')]
onepct_rlut_df_list = []
for ism in onepct_ism_areacella_exist:
onepct_rlut_df_list += [get_cmip6_data_gs(ism,'rlut','1pctCO2')]
onepct_rsut_df_list = []
for ism in onepct_ism_areacella_exist:
onepct_rsut_df_list += [get_cmip6_data_gs(ism,'rsut','1pctCO2')]
onepct_rsdt_df_list = []
for ism in onepct_ism_areacella_exist:
onepct_rsdt_df_list += [get_cmip6_data_gs(ism,'rsdt','1pctCO2')]
def get_annual_CMIP6_data_gstore_co2mass(activity, table, variable, experiment, institution, source, member):
# eg activity='CMIP', table='Amon', variable='tas', experiment='historical', institution="NCAR", source="CESM2", member="r10i1p1f1"
query = gs_stores.query("activity_id==\'"+activity+"\' & table_id==\'"+table+"\' & variable_id==\'"+variable+"\' & experiment_id==\'"+experiment+"\' & institution_id==\'"+institution+"\' & source_id==\'"+source+"\' & member_id==\'"+member+"\'")
if query.empty:
print('No results for this request')
return None
# create a mutable-mapping-style interface to the store
mapper = gcs.get_mapper(query.zstore.values[0])
# open it using xarray and zarr
ds = xr.open_zarr(mapper, consolidated=True)
return ds[variable].groupby('time.year').mean('time').to_pandas().rename(institution+'_'+source+'_'+member)
onepct_co2mass_ism = gs_stores.loc[(gs_stores.experiment_id=='1pctCO2')&(gs_stores.variable_id=='co2mass')].ism
def get_cmip6_data_gs_co2mass(ism,var,exp):
print('getting '+ism+' '+var)
ism_split = ism.split('_')
_out = get_annual_CMIP6_data_gstore_co2mass('CMIP', 'Amon', var, exp, ism_split[0], ism_split[1], ism_split[2])
print('got '+ism)
return _out
onepct_co2mass_df_list = []
for ism in onepct_co2mass_ism:
onepct_co2mass_df_list += [get_cmip6_data_gs_co2mass(ism,'co2mass','1pctCO2')]
onepct_co2_esgf_results = esgf_search(activity_id='CMIP', table_id='Amon', variable_id='co2mass', experiment_id='1pctCO2')
onepct_co2_esgf_ism = [x.split('/')[8]+'_'+x.split('/')[9]+'_'+x.split('/')[11] for x in onepct_co2_esgf_results]
onepct_co2_esgf_ism = [x for x in onepct_co2_esgf_ism if not x in list(onepct_co2mass_ism)]
def get_annual_CMIP6_data_esgf_co2mass(activity, table, variable, experiment, institution, source, member):
# eg activity='CMIP', table='Amon', variable='tas', experiment='historical', institution="NCAR", source="CESM2", member="r10i1p1f1"
result = esgf_search(activity_id=activity, table_id=table, variable_id=variable, experiment_id=experiment,institution_id=institution, source_id=source, member_id=member)
if not result:
print('No results for this request')
return None
# select results with only the latest datestamp:
latest = sorted([x.split('/')[15] for x in result])[-1]
result = [x for x in result if x.split('/')[15]==latest]
# remove duplicate results
result_1 = []
for item in result:
if item.split('/')[-1] in [x.split('/')[-1] for x in result_1]:
continue
else:
result_1 += [item]
ds = xr.open_mfdataset(result_1, combine='by_coords')
return ds[variable].groupby('time.year').mean('time').to_pandas().rename(institution+'_'+source+'_'+member)
for x in onepct_co2_esgf_ism:
onepct_co2mass_df_list += [get_annual_CMIP6_data_esgf_co2mass('CMIP','Amon','co2mass','1pctCO2',x.split('_')[0],x.split('_')[1],x.split('_')[2])]
def get_annual_CMIP6_data_gstore_co2(activity, table, variable, experiment, institution, source, member):
# eg activity='CMIP', table='Amon', variable='tas', experiment='historical', institution="NCAR", source="CESM2", member="r10i1p1f1"
query = gs_stores.query("activity_id==\'"+activity+"\' & table_id==\'"+table+"\' & variable_id==\'"+variable+"\' & experiment_id==\'"+experiment+"\' & institution_id==\'"+institution+"\' & source_id==\'"+source+"\' & member_id==\'"+member+"\'")
if query.empty:
print('No results for this request')
return None
# create a mutable-mapping-style interface to the store
mapper = gcs.get_mapper(query.zstore.values[0])
# open it using xarray and zarr
ds = xr.open_zarr(mapper, consolidated=True)
area_query = gs_stores.query("variable_id=='areacella' & source_id==\'"+source+"\'")
if area_query.empty:
files_area = esgf_search(variable_id='areacella', activity_id=activity, institution_id=institution, source_id=source)
if not files_area:
print('No areacella for this request')
return None
ds_area = xr.open_dataset(files_area[0])
else:
ds_area = xr.open_zarr(gcs.get_mapper(area_query.zstore.values[0]), consolidated=True)
coords = list(ds[variable].coords.keys())
if 'lat' in coords:
dims = ['lat','lon']
else:
dims = ['latitude','longitude']
total_area = ds_area.areacella.sum(dim=dims)
ta_timeseries = (ds[variable].isel(plev=0) * ds_area.areacella).sum(dim=dims) / total_area
return ta_timeseries.groupby('time.year').mean('time').to_pandas().rename(institution+'_'+source+'_'+member)
onepct_co2_ism = gs_stores.loc[(gs_stores.experiment_id=='1pctCO2')&(gs_stores.variable_id=='co2')].ism
def get_cmip6_data_gs_co2(ism,var,exp):
print('getting '+ism+' '+var)
ism_split = ism.split('_')
_out = get_annual_CMIP6_data_gstore_co2('CMIP', 'Amon', var, exp, ism_split[0], ism_split[1], ism_split[2])
print('got '+ism)
return _out
onepct_co2_ism_areacella_exist = [x for x in onepct_co2_ism if x.split('_')[1] in areacella_s_all]
onepct_co2_df_list = []
for ism in onepct_co2_ism_areacella_exist:
onepct_co2_df_list += [get_cmip6_data_gs_co2(ism,'co2','1pctCO2')]
def get_annual_CMIP6_data_esgf_co2(activity, table, variable, experiment, institution, source, member):
# eg activity='CMIP', table='Amon', variable='tas', experiment='historical', institution="NCAR", source="CESM2", member="r10i1p1f1"
result = esgf_search(activity_id=activity, table_id=table, variable_id=variable, experiment_id=experiment,institution_id=institution, source_id=source, member_id=member)
if not result:
print('No results for this request')
return None
# select results with only the latest datestamp:
latest = sorted([x.split('/')[15] for x in result])[-1]
result = [x for x in result if x.split('/')[15]==latest]
# remove duplicate results
result_1 = []
for item in result:
if item.split('/')[-1] in [x.split('/')[-1] for x in result_1]:
continue
else:
result_1 += [item]
ds = xr.open_mfdataset(result_1, combine='by_coords')
files_area = esgf_search(variable_id='areacella', activity_id=activity, institution_id=institution, source_id=source)
if not files_area:
print('No areacella for this request')
return None
ds_area = xr.open_dataset(files_area[0])
coords = list(ds[variable].coords.keys())
if 'lat' in coords:
dims = ['lat','lon']
else:
dims = ['latitude','longitude']
total_area = ds_area.areacella.sum(dim=dims)
ta_timeseries = (ds[variable].isel(plev=0) * ds_area.areacella).sum(dim=dims) / total_area
return ta_timeseries.groupby('time.year').mean('time').to_pandas().rename(institution+'_'+source+'_'+member)
onepct_co2_esgf_results = esgf_search(activity_id='CMIP', table_id='Amon', variable_id='co2', experiment_id='1pctCO2')
onepct_co2_esgf_ism = [x.split('/')[8]+'_'+x.split('/')[9]+'_'+x.split('/')[11] for x in onepct_co2_esgf_results]
for ism in [x for x in set(onepct_co2_esgf_ism) if not x in onepct_co2_ism_areacella_exist]:
print('getting '+ism)
onepct_co2_df_list += [get_annual_CMIP6_data_esgf_co2('CMIP', 'Amon', 'co2', '1pctCO2', ism.split('_')[0], ism.split('_')[1], ism.split('_')[2])]
pd.concat(onepct_co2_df_list,axis=1).to_csv('./cmip6_data/onepct_co2.csv')
# piControl_df_list = []
for ism in piControl_ism_areacella_exist:
piControl_df_list += [get_cmip6_data_gs(ism,'tas','piControl')]
ESM3_picontrol = xr.open_mfdataset(esgf_search(activity_id='CMIP', table_id='Amon', variable_id='tas', experiment_id='piControl', institution_id='E3SM-Project', source_id='E3SM-1-0', member_id='r1i1p1f1'),combine='by_coords')
# ds_area = xr.open_dataset(files_area[0])
# coords = list(ds[variable].coords.keys())
# if 'lat' in coords:
# dims = ['lat','lon']
# else:
# dims = ['latitude','longitude']
# total_area = ds_area.areacella.sum(dim=dims)
# ta_timeseries = (ds[variable].isel(plev=0) * ds_area.areacella).sum(dim=dims) / total_area
# return ta_timeseries.groupby('time.year').mean('time').to_pandas().rename(institution+'_'+source+'_'+member)
ESM3_area = xr.open_dataset(esgf_search(activity_id='CMIP', variable_id='areacella',source_id='E3SM-1-0')[-1])
dims=['lat','lon']
total_area = ESM3_area.areacella.sum(dim=dims)
ta_timeseries = (ESM3_picontrol['tas'] * ESM3_area.areacella).sum(dim=dims) / total_area
ESM3_picontrol_tas = ta_timeseries.groupby('time.year').mean('time').to_pandas().rename('E3SM-Project_E3SM-1-0_r1i1p1f1')
ESM3_picontrol_tas.plot()
piControl_rlut_df_list = []
for ism in piControl_ism_areacella_exist:
piControl_rlut_df_list += [get_cmip6_data_gs(ism,'rlut','piControl')]
piControl_rsut_df_list = []
for ism in piControl_ism_areacella_exist:
piControl_rsut_df_list += [get_cmip6_data_gs(ism,'rsut','piControl')]
piControl_rsdt_df_list = []
for ism in piControl_ism_areacella_exist:
piControl_rsdt_df_list += [get_cmip6_data_gs(ism,'rsdt','piControl')]
abrutp4x_tas_df_list = []
for ism in abrupt_4x_ism_areacella_exist:
abrutp4x_tas_df_list += [get_cmip6_data_gs(ism,'tas','abrupt-4xCO2')]
abrutp4x_rlut_df_list = []
for ism in abrupt_4x_ism_areacella_exist:
abrutp4x_rlut_df_list += [get_cmip6_data_gs(ism,'rlut','abrupt-4xCO2')]
abrutp4x_rsut_df_list = []
for ism in abrupt_4x_ism_areacella_exist:
abrutp4x_rsut_df_list += [get_cmip6_data_gs(ism,'rsut','abrupt-4xCO2')]
abrutp4x_rsdt_df_list = []
for ism in abrupt_4x_ism_areacella_exist:
abrutp4x_rsdt_df_list += [get_cmip6_data_gs(ism,'rsdt','abrupt-4xCO2')]
abrutp4x_rsdt_df = pd.concat(abrutp4x_rsdt_df_list,axis=1)
abrutp4x_rsut_df = pd.concat(abrutp4x_rsut_df_list,axis=1)
abrutp4x_rlut_df = pd.concat(abrutp4x_rlut_df_list,axis=1)
abrutp4x_tas_df = pd.concat(abrutp4x_tas_df_list,axis=1)
piControl_tas_df = pd.concat(piControl_df_list,axis=1)
piControl_rsdt_df = pd.concat(piControl_rsdt_df_list,axis=1)
piControl_rsut_df = pd.concat(piControl_rsut_df_list,axis=1)
piControl_rlut_df = pd.concat(piControl_rlut_df_list,axis=1)
onepct_tas_df = pd.concat(onepct_tas_df_list,axis=1)
onepct_rlut_df = pd.concat(onepct_rlut_df_list,axis=1)
onepct_rsut_df = pd.concat(onepct_rsut_df_list,axis=1)
onepct_rsdt_df = pd.concat(onepct_rsdt_df_list,axis=1)
onepct_co2mass_df = pd.concat(onepct_co2mass_df_list,axis=1)
# onepct_co2mass_df.to_csv('./cmip6_data/onepct_co2mass.csv')
# onepct_tas_df.to_csv('./cmip6_data/onepct_tas.csv')
# onepct_rlut_df.to_csv('./cmip6_data/onepct_rlut.csv')
# onepct_rsut_df.to_csv('./cmip6_data/onepct_rsut.csv')
# onepct_rsdt_df.to_csv('./cmip6_data/onepct_rsdt.csv')
# piControl_rlut_df.to_csv('./cmip6_data/piControl_rlut.csv')
# piControl_rsut_df.to_csv('./cmip6_data/piControl_rsut.csv')
# piControl_rsdt_df.to_csv('./cmip6_data/piControl_rsdt.csv')
# abrutp4x_rsdt_df.to_csv('./cmip6_data/abrupt-4xCO2_rsdt.csv')
# abrutp4x_rsut_df.to_csv('./cmip6_data/abrupt-4xCO2_rsut.csv')
# abrutp4x_rlut_df.to_csv('./cmip6_data/abrupt-4xCO2_rlut.csv')
# abrutp4x_tas_df.to_csv('./cmip6_data/abrupt-4xCO2_tas.csv')
# piControl_tas_df.to_csv('./cmip6_data/piControl_tas.csv')
def get_annual_CMIP6_data_esgf(activity, table, variable, experiment, institution, source, member):
# eg activity='CMIP', table='Amon', variable='tas', experiment='historical', institution="NCAR", source="CESM2", member="r10i1p1f1"
result = esgf_search(activity_id=activity, table_id=table, variable_id=variable, experiment_id=experiment,institution_id=institution, source_id=source, member_id=member)
if not result:
print('No results for this request')
return None
# select results with only the latest datestamp:
latest = sorted([x.split('/')[15] for x in result])[-1]
result = [x for x in result if x.split('/')[15]==latest]
# remove duplicate results
result_1 = []
for item in result:
if item.split('/')[-1] in [x.split('/')[-1] for x in result_1]:
continue
else:
result_1 += [item]
ds = xr.open_mfdataset(result_1, combine='by_coords')
files_area = esgf_search(variable_id='areacella', activity_id=activity, institution_id=institution, source_id=source)
if not files_area:
print('No areacella for this request')
return None
ds_area = xr.open_dataset(files_area[0])
coords = list(ds[variable].coords.keys())
if 'lat' in coords:
dims = ['lat','lon']
else:
dims = ['latitude','longitude']
total_area = ds_area.areacella.sum(dim=dims)
ta_timeseries = (ds[variable] * ds_area.areacella).sum(dim=dims) / total_area
return ta_timeseries.groupby('time.year').mean('time').to_pandas().rename(institution+'_'+source+'_'+member)
def get_annual_CMIP6_data_esgf_multivar(activity, table, variables, experiment, institution, source, member):
# eg activity='CMIP', table='Amon', variable='tas', experiment='historical', institution="NCAR", source="CESM2", member="r10i1p1f1"
result = esgf_search(activity_id=activity, table_id=table, experiment_id=experiment,institution_id=institution, source_id=source, member_id=member)
result = [x for x in result if x.split('/')[13] in variables]
if not result:
print('No results for this request')
return None
# select results with only the latest datestamp:
# latest = sorted([x.split('/')[15] for x in result])[-1]
# result = [x for x in result if x.split('/')[15]==latest]
# remove duplicate results
result_1 = []
for item in result:
if item.split('/')[-1] in [x.split('/')[-1] for x in result_1]:
continue
else:
result_1 += [item]
ds = xr.open_mfdataset(result_1, combine='by_coords')
files_area = esgf_search(variable_id='areacella', activity_id=activity, institution_id=institution, source_id=source)
if not files_area:
print('No areacella for this request')
return None
ds_area = xr.open_dataset(files_area[0])
coords = list(ds[variables].coords.keys())
if 'lat' in coords:
dims = ['lat','lon']
else:
dims = ['latitude','longitude']
total_area = ds_area.areacella.sum(dim=dims)
ta_timeseries = (ds[variables] * ds_area.areacella).sum(dim=dims) / total_area
_out = ta_timeseries.groupby('time.year').mean('time').to_dataframe()[variables]
return pd.concat([_out],axis=1,keys=[institution+'_'+source+'_'+member])
piControl_list = esgf_search(activity_id='CMIP', table_id='Amon', variable_id='tas', experiment_id='piControl')
piControl_list_nodupl = []
for item in piControl_list:
if item.split('/')[-1] in [x.split('/')[-1] for x in piControl_list_nodupl]:
continue
else:
piControl_list_nodupl += [item]
abrupt4x_list = esgf_search(activity_id='CMIP', table_id='Amon', variable_id='tas', experiment_id='abrupt-4xCO2')
abrupt4x_list_nodupl = []
for item in abrupt4x_list:
if item.split('/')[-1] in [x.split('/')[-1] for x in abrupt4x_list_nodupl]:
continue
else:
abrupt4x_list_nodupl += [item]
areacella_list = esgf_search(activity_id='CMIP', variable_id='areacella')
areacella_list_nodupl = []
for item in areacella_list:
if item.split('/')[-1] in [x.split('/')[-1] for x in areacella_list_nodupl]:
continue
else:
areacella_list_nodupl += [item]
abrupt4x_ism_list = list(set([x.split('/')[8]+'_'+x.split('/')[9]+'_'+x.split('/')[11] for x in abrupt4x_list_nodupl]))
piControl_ism_list = list(set([x.split('/')[8]+'_'+x.split('/')[9]+'_'+x.split('/')[11] for x in piControl_list_nodupl]))
areacella_ism_list = list(set([x.split('/')[8]+'_'+x.split('/')[9]+'_'+x.split('/')[11] for x in areacella_list_nodupl]))
areacella_s_list = list(set([x.split('_')[1] for x in areacella_ism_list]))
piControl_ism_areacella_exist = [x for x in piControl_ism_list if x.split('_')[1] in areacella_s_list]
abrupt4x_ism_areacella_exist = [x for x in abrupt4x_ism_list if x.split('_')[1] in areacella_s_list]
abrupt4x_tas_df = pd.read_csv('./cmip6_data/abrupt-4xCO2_tas.csv',index_col=0)
esgf_abrupt4x_list = [x for x in abrupt4x_ism_areacella_exist if not x in abrupt4x_tas_df.columns]
piControl_tas_df = pd.read_csv('./cmip6_data/piControl_tas.csv',index_col=0)
esgf_piControl_list = [x for x in piControl_ism_areacella_exist if not x in piControl_tas_df.columns]
def get_CMIP6_data(ism,exp='abrupt-4xCO2',var='tas',multivar=False):
ism_split = ism.split('_')
if multivar:
_out = get_annual_CMIP6_data_esgf_multivar('CMIP', 'Amon', var, exp, ism_split[0], ism_split[1], ism_split[2])
else:
_out = get_annual_CMIP6_data_esgf('CMIP', 'Amon', var, exp, ism_split[0], ism_split[1], ism_split[2])
print(ism+' complete')
return _out
# abrupt4x_df_list_esgf = []
for x in esgf_abrupt4x_list:
abrupt4x_df_list_esgf += [get_CMIP6_data(x,'abrupt-4xCO2',['tas','rlut','rsut','rsdt'],True)]
# P1=multiprocessing.Pool(processes=8)
# abrupt4xCO2_df = P1.starmap(get_CMIP6_data,[(x,'abrupt-4xCO2',['tas','rlut','rsut','rsdt'],True) for x in abrupt4x_ism_areacella_exist])
# P1.close
piControl_df_list_esgf = []
for x in piControl_ism_areacella_exist:
piControl_df_list_esgf += [get_CMIP6_data(x,'piControl','tas')]
# P1=multiprocessing.Pool(processes=8)
# piControl_df = P1.starmap(get_CMIP6_data,[(x,'piControl','tas',False) for x in piControl_ism_areacella_exist])
# P1.close
```
| github_jupyter |
```
import pandas as pd
import numpy as np
```
First it loads the spreadsheet we assembled at eLife's hackathon (https://docs.google.com/spreadsheets/d/1Pr5q3xG1SwjuZ-GhbNxWgQXk57aQfbnkuonw1OmOHSY/edit?usp=sharing)
```
data = pd.read_csv("data/essential_oil_instrument_list_elife2019 - sheet 1.csv")
data.head()
```
We want to get only the rows that are not done, have entries for manufacturer and instument type and not have been assigned a number
```
data_filtered = data.query('done=="no"')
data_filtered = data_filtered[data_filtered["Q-number for manufacturer"].notnull()]
data_filtered = data_filtered[data_filtered["Q-number for instrument types"].notnull()]
data_filtered = data_filtered[data_filtered["Q-number for instruments"].isnull()]
print(data_filtered)
```
Now we just extract row-wise the information. Then, we need to copy and paste it to quickstatements (https://tools.wmflabs.org/quickstatements/#/batch).
```
for index, row in data_filtered.iterrows():
print("CREATE\n" +
'LAST|Len|' + '"' + row['instrument'] + '"\n' +
'LAST|Den|' + '"'+ row['instrument type'] + " manufactured by " + row['manufacturer'] + '"\n' +
"LAST|P31|" + row['Q-number for instrument types'] + '\n' +
"LAST|P178|" + row['Q-number for manufacturer'] )
```
This has to be manually copied and pasted in quickstatements (https://tools.wmflabs.org/quickstatements/#/batch).
Notice that you need to have 100+ edits in wikidata to be allowed to do batches in quickstatements.
We also would like to add the information of which projects used which equiment. For this we will use the property P4510, (describes a project that uses). This a tiny bit more complicated, as we need to get the Q-value for the paper from its PMC id.
For that, we will use the wikidata integrator module, from the SuLab:
https://github.com/SuLab/WikidataIntegrator
```
from wikidataintegrator import wdi_helpers
# Note: It might take some time depending on internet connection speed.
for index, row in data_filtered.iterrows():
pmcid = row['article where it was mentioned (P4510)'][3:]
a = wdi_helpers.PublicationHelper(pmcid, id_type="pmcid", source="europepmc").get_or_create(None)
paper_q_value = a[0]
print("CREATE\n" +
'LAST|Len|' + '"' + row['instrument'] + '"\n' +
'LAST|Den|' + '"'+ row['instrument type'] + " manufactured by " + row['manufacturer'] + '"\n' +
"LAST|P31|" + row['Q-number for instrument types'] + '\n' +
"LAST|P178|" + row['Q-number for manufacturer'] + '\n' +
paper_q_value + "|P4510|" + "LAST")
```
This will only work in quickstatements if the item has not been created already. As I had already created the item for "MEGCs QP-2010 Plus" (Q67204908),
for now, I will manually add a quickstatements of the sort:
Q28554682|P4510|Q67204908
| github_jupyter |
```
%matplotlib inline
```
# Model Building Part 3
Code for building the models
_Author: Jimmy Charité_
_Email: jimmy.charite@gmail.com_
Following up with part one and two, I will combine the bag of words and the macro-level text summary statistics approach
```
import os
import pandas as pd
import numpy as np
import scipy as sp
import seaborn as sns
import matplotlib.pyplot as plt
import json
from IPython.display import Image
from IPython.core.display import HTML
retval=os.chdir("..")
clean_data=pd.read_pickle('./clean_data/clean_data.pkl')
clean_data.head()
kept_cols=['helpful', 'num_sents', 'num_words', 'readability', 'neg_senti',
'pos_senti', 'neu_senti', 'comp_senti', 'text_lemma',]
```
## Training and Testing Split
```
my_rand_state=0
test_size=0.25
from sklearn.model_selection import train_test_split
X = (clean_data[kept_cols].iloc[:,1:]).as_matrix()
y = (clean_data[kept_cols].iloc[:,0]).tolist()
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=test_size,
random_state=my_rand_state)
```
## Scaling
```
from sklearn.preprocessing import StandardScaler
std_scale=StandardScaler()
```
## Text
```
from sklearn.feature_extraction.text import TfidfVectorizer
#set max_features to minimize training time
#also, I didn't apply LDA-based dimensionality reduction
tfidf=TfidfVectorizer(lowercase=False,max_features=200)
```
## Custom Feature Separator
```
from sklearn.base import BaseEstimator, TransformerMixin
class ExtractText(BaseEstimator, TransformerMixin):
"""
Separates the features by numerical and text
"""
def __init__(self, text,n_text=-1):
self.text = text
self.n_text=n_text
def fit(self, X, y=None):
return self
def transform(self, X, y=None):
if(self.text):
return X[:,self.n_text]
else:
return X[:,:self.n_text]
from sklearn.pipeline import FeatureUnion
```
## Classification Models
```
from sklearn.naive_bayes import GaussianNB
from sklearn.discriminant_analysis import QuadraticDiscriminantAnalysis
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
```
Although tuning is not necessary for Naive Bayes, I pass the default parameters of those models to GridSearchCV anyway so that I can do a direct pair-wise comparison with the other models across the different steps of cross-validation.
In the interest of time, I didn't use the SVM classifier.
```
nb_clf=GaussianNB()
priors=[None]
qda_clf=QuadraticDiscriminantAnalysis()
reg_param=[0.0, 0.25, 0.5, 0.75]
log_clf=LogisticRegression(penalty='l2')
C=[0.001 , 0.01, 10, 100,1000]
rf_clf=RandomForestClassifier()
n_estimators=[100,200]
max_features=[.1,.3,.5]
class_weight=['balanced']
class_weight.extend([{1: w} for w in [1, 2, 10]])
```
## Creating Pipelines
```
from imblearn import pipeline #needed if mixing imblearn with sklearn classes
from sklearn.model_selection import GridSearchCV
from sklearn.model_selection import StratifiedKFold
```
I plan on using imblearn classes for later iterations so I use it's pipeline in the beginning for convenience
```
n_jobs=4
n_folds=10
skfold = StratifiedKFold(n_splits=n_folds,random_state=my_rand_state, shuffle=False)
class DenseTransformer(BaseEstimator, TransformerMixin):
def transform(self, X, y=None, **fit_params):
return X.todense()
def fit_transform(self, X, y=None, **fit_params):
self.fit(X, y, **fit_params)
return self.transform(X)
def fit(self, X, y=None, **fit_params):
return self
```
#### Main Feature Union
```
ft_union=FeatureUnion(transformer_list=[('text_pipe',pipeline.Pipeline([('extract',ExtractText(text=True)),
('tfidf',tfidf),
('to_dense', DenseTransformer())])),
('numb_pipe',pipeline.Pipeline([('extract',ExtractText(text=False)),
('scale',std_scale)]))])
```
#### Naive Bayes Estimators
```
nb_clf_b = pipeline.Pipeline(steps=[('union',ft_union),('clf',nb_clf)])
nb_clf_est_b = GridSearchCV(estimator=nb_clf_b,cv=skfold,
scoring='roc_auc',n_jobs=n_jobs,
param_grid=dict(clf__priors=priors))
```
#### QDA Estimators
```
qda_clf_b = pipeline.Pipeline(steps=[('union',ft_union),('clf',qda_clf)])
qda_clf_est_b = GridSearchCV(estimator=qda_clf_b,cv=skfold,
scoring='roc_auc',n_jobs=n_jobs,
param_grid=dict(clf__reg_param=reg_param))
```
#### Logistic Estimators
```
log_clf_b = pipeline.Pipeline(steps=[('union',ft_union),('clf',log_clf)])
log_clf_est_b = GridSearchCV(estimator=log_clf_b,cv=skfold,
scoring='roc_auc',n_jobs=n_jobs,
param_grid=dict(clf__C=C,
clf__class_weight=class_weight))
```
#### Random Forest Estimators
```
rf_clf_b = pipeline.Pipeline(steps=[('union',ft_union),('clf',rf_clf)])
rf_clf_est_b = GridSearchCV(estimator=rf_clf_b,cv=skfold,
scoring='roc_auc',n_jobs=n_jobs,
param_grid=dict(clf__n_estimators=n_estimators,
clf__max_features=max_features,
clf__class_weight=class_weight))
```
## Fitting Estimators
```
from sklearn.externals import joblib
```
Basic Estimators: no bag of words or PCA
```
nb_clf_est_b.fit(X_train,y_train)
joblib.dump(nb_clf_est_b, './other_output/merged/nb_clf_est_b.pkl')
qda_clf_est_b.fit(X_train,y_train)
joblib.dump(qda_clf_est_b, './other_output/merged/qda_clf_est_b.pkl')
log_clf_est_b.fit(X_train,y_train)
joblib.dump(log_clf_est_b, './other_output/merged/log_clf_est_b.pkl')
rf_clf_est_b.fit(X_train,y_train)
joblib.dump(rf_clf_est_b, './other_output/merged/rf_clf_est_b.pkl')
```
## Testing Estimators
```
from sklearn.metrics import roc_curve, auc
nb_clf_est_b=joblib.load('./other_output/merged/nb_clf_est_b.pkl')
qda_clf_est_b=joblib.load('./other_output/merged/qda_clf_est_b.pkl')
log_clf_est_b=joblib.load('./other_output/merged/log_clf_est_b.pkl')
rf_clf_est_b=joblib.load('./other_output/merged/rf_clf_est_b.pkl')
nb_fpr, nb_tpr, _ = roc_curve(y_test,
nb_clf_est_b.predict_proba(X_test)[:,1])
nb_roc_auc = auc(nb_fpr, nb_tpr)
qda_fpr, qda_tpr, _ = roc_curve(y_test,
qda_clf_est_b.predict_proba(X_test)[:,1])
qda_roc_auc = auc(qda_fpr, qda_tpr)
log_fpr, log_tpr, _ = roc_curve(y_test,
log_clf_est_b.predict_proba(X_test)[:,1])
log_roc_auc = auc(log_fpr, log_tpr)
rf_fpr, rf_tpr, _ = roc_curve(y_test,
rf_clf_est_b.predict_proba(X_test)[:,1])
rf_roc_auc = auc(rf_fpr, rf_tpr)
plt.plot(nb_fpr, nb_tpr, color='cyan', linestyle='--',
label='NB (area = %0.2f)' % nb_roc_auc, lw=2)
plt.plot(qda_fpr, qda_tpr, color='indigo', linestyle='--',
label='QDA (area = %0.2f)' % qda_roc_auc, lw=2)
plt.plot(log_fpr, log_tpr, color='seagreen', linestyle='--',
label='LOG (area = %0.2f)' % log_roc_auc, lw=2)
plt.plot(rf_fpr, rf_tpr, color='blue', linestyle='--',
label='RF (area = %0.2f)' % rf_roc_auc, lw=2)
plt.plot([0, 1], [0, 1], linestyle='--', lw=2, color='k',
label='Luck')
plt.xlim([-0.05, 1.05])
plt.ylim([-0.05, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('ROC Curves of Basic Models Using Bag of Words and Macro-Text Stats')
plt.legend(loc="lower right")
plt.savefig('./plots/ROC_Basic_BOW_MERGED.png', bbox_inches='tight')
plt.show()
```
Closer look at the variability of the best model
```
from scipy.stats import sem
len(y_test) #the sample is large enough that we can get away with 5% draws
y_test=np.array(y_test)
X_test=np.array(X_test)
#initialize
n_bootstraps = 2000
rng_seed = 1
sample_percent=0.05
min_index=0
max_index=len(y_test)-1
draw_size=int(len(y_test)*sample_percent)
bootstrapped_scores = []
rng = np.random.RandomState(rng_seed)
for i in range(n_bootstraps):
# bootstrap by sampling with replacement on
indices = rng.random_integers(min_index, max_index, draw_size)
#calculate ROC from
rf_fpr, rf_tpr, _ = roc_curve(y_test[indices],
rf_clf_est_b.predict_proba(X_test[indices,:])[:,1])
rf_roc_auc = auc(rf_fpr, rf_tpr)
#save
bootstrapped_scores.append(rf_roc_auc)
import pickle
with open('./other_output/merged/rf_bootstrapped_scores.pkl', 'wb') as f:
pickle.dump(bootstrapped_scores, f)
plt.hist(bootstrapped_scores, bins=50)
plt.title('Histogram of Bootstrapped AUC ROC of\nRandom Forest Model Using Bag of Words and Macro-Text Stats')
plt.savefig('./plots/ROC_Histogram_Basic_BOW_MERGED.png', bbox_inches='tight')
plt.show()
pd.DataFrame({'auc':bootstrapped_scores}).auc.describe()
```
Importing bootstrapped scores from parts 1 & 2 for side-by-side comparison
```
with open('./other_output/rf_bootstrapped_scores.pkl', 'rb') as f:
bootstrapped_scores_macrotext = pickle.load(f)
with open('./other_output/bow/rf_bootstrapped_scores.pkl', 'rb') as f:
bootstrapped_scores_bow = pickle.load(f)
bs_auc=pd.DataFrame({'Macro-Text Only':bootstrapped_scores_macrotext,
'Bag of Words Only':bootstrapped_scores_bow,
'Combined':bootstrapped_scores})
bs_auc=bs_auc[['Macro-Text Only','Bag of Words Only','Combined']]
bs_auc.describe()
pl_axes=bs_auc.boxplot(return_type='axes')
x=pl_axes.axes.set_ylabel('AUC ROC')
x=pl_axes.axes.set_title('Comparison of Bootstrapped AUC ROC of Random Forest Model\nUsing Macro-Text Stats, Bag of Words, and Combined Approaches')
plt.savefig('./plots/BoxPlot_ROC_MacText_BOX.png', bbox_inches='tight')
plt.show()
```
Interestingly, but as expected, combining the marco-text and bag of words approaches improves the minimum, mean, and maximum AUC ROC of the random forest model.
| github_jupyter |
### using textrank to get key sentences
```
# 출력이 너무 길어지지 않게하기 위해 찍지 않도록 했으나
# 실제 학습 할 때는 아래 두 줄을 주석처리 하는 것을 권장한다.
import warnings
warnings.filterwarnings('ignore')
import pandas as pd
import numpy as np
print(pd.__version__)
print(np.__version__)
# pandas 설정: 모든 row와 모든 column을 출력하게 하고, 각 column 별 출력되는 글자수는 50자 이내
pd.set_option('display.max_rows', None, 'display.max_columns', None, 'display.max_colwidth', 50)
# 위의 설정으로 간단하게 보는 df_specific
df_specific = pd.read_csv("./data_wrangled/df_specific_wrangle.csv", encoding="utf-8")
df_specific.head(2)
# 위의 설정으로 간단하게 보는 df_specific
df_specific.tail(2)
print(df_specific.columns)
print(df_specific.shape)
# pandas 설정: 각 column 별 (혹은 한 cell에서) 출력되는 글자수 제한을 없애기
pd.set_option('display.max_colwidth', None)
import re
# removing other letters except for korean letters from the string data
def remove_junk(str_data):
str_data = str_data.replace("\\\\n", "")
str_data = str_data.replace("\\n", "")
str_data = str_data.replace("\n", "")
str_data = str_data.replace("\\", "")
str_data = str_data.replace("\\t", "")
str_data = str_data.replace("NaN", "")
# print(str_data) makes an error. too much of data
# 한글, english 빼고 전부 날리기
# str_data = re.sub('[^가-힣ㄱ-ㅎㅏ-ㅣa-zA-Z|0-9|.,]', ' ', str_data)
# 한글 빼고 다 날리기
str_data = re.sub('[^가-힣ㄱ-ㅎㅏ]', ' ', str_data)
# replace multiple spaces into single space
str_data = ' '.join(str_data.split())
return str_data
def access_univ_info(dataframe, column, univ_code):
df_row = dataframe.loc[dataframe["대학코드"] == univ_code]
str_univ_info = df_row[column].to_string(index=False).lstrip()
str_univ_info = remove_junk(str_univ_info)
return str_univ_info
# removing junks only from the string data
def remove_junk_only(str_data):
str_data = str_data.replace("\\\\n", "")
str_data = str_data.replace("\\n", "")
str_data = str_data.replace("\n", "")
str_data = str_data.replace("\\", "")
str_data = str_data.replace("\\t", "")
str_data = str_data.replace("NaN", "")
# replace multiple spaces into single space
str_data = ' '.join(str_data.split())
return str_data
# accessing university info of given column (= accessing one cell)
def access_univ_info_raw(dataframe, column, univ_code):
df_row = dataframe.loc[dataframe["대학코드"] == univ_code]
str_univ_info = df_row[column].to_string(index=False).lstrip()
str_univ_info = remove_junk_only(str_univ_info)
return str_univ_info
# collecting all column values into one string
def column_to_string(dataframe, column_name: str):
str_corpus = dataframe[column_name].to_string(index=False).lstrip()
str_corpus = remove_junk_only(str_corpus)
return str_corpus
column_data = "gen_info"
univ_code = "DK000003"
str_example = access_univ_info_raw(df_specific, column_data, univ_code)
str_example[:100]
str_example[:2500]
# collecting all column values into one string
corpus = column_to_string(df_specific, column_data)
print("글자수:", len(corpus))
corpus[:100]
from textrankr import TextRank
def yield_summary(df_specific, column_data, univ_code, sentence_no):
string_data = access_univ_info_raw(df_specific, column_data, univ_code)
textrank = TextRank(string_data)
# print(textrank.summarize()) # gives you some text
summarized_list = textrank.summarize(sentence_no, verbose=False)
summary_text_md = ""
for sentence in summarized_list:
summary_line = \
f"""* {sentence}
"""
summary_text_md += summary_line
return summary_text_md
column_data = "etc_info"
univ_code = "CN000016"
print(yield_summary(df_specific, column_data, univ_code, 5))
from textrankr import TextRank
column_data = "gen_info"
univ_code = "DK000003"
str_example = access_univ_info(df_specific, column_data, univ_code)
str_example = str_example.replace("니다", "니다.")
textrank = TextRank(str_example)
print(textrank.summarize()) # gives you some text
textrank.summarize(10, verbose=False) # up to 3 sentences, returned as list
from textrankr import TextRank
column_data = "gen_info"
univ_code = "DK000003"
str_example = access_univ_info_raw(df_specific, column_data, univ_code)
textrank = TextRank(str_example)
print(textrank.summarize()) # gives you some text
textrank.summarize(10, verbose=False) # up to 3 sentences, returned as list
from textrankr import TextRank
column_data = "gen_info"
univ_code = "DK000003"
str_example = access_univ_info_raw(df_specific, column_data, univ_code)
textrank = TextRank(str_example)
print(textrank.summarize()) # gives you some text
textrank.summarize(10, verbose=False) # up to 3 sentences, returned as list
from textrankr import TextRank
column_data = "gen_info"
univ_code = "JP000017"
str_example = access_univ_info_raw(df_specific, column_data, univ_code)
textrank = TextRank(str_example)
print(textrank.summarize()) # gives you some text
print(textrank.summarize(10, verbose=False)) # up to 3 sentences, returned as list
from textrankr import TextRank
column_data = "gen_info"
univ_code = "CN000016"
str_example = access_univ_info_raw(df_specific, column_data, univ_code)
textrank = TextRank(str_example)
print(textrank.summarize()) # gives you some text
print(textrank.summarize(10, verbose=False)) # up to 3 sentences, returned as list
```
| github_jupyter |
# Predicting handwriting number with mnist
The model of mnist is generated using numpy/keras on Python.
```python
from tensorflow.contrib.keras.api.keras.datasets import mnist
from tensorflow.contrib.keras.api.keras.models import Sequential
from tensorflow.contrib.keras.api.keras.layers import Dense, Dropout, Activation
from tensorflow.contrib.keras.api.keras.optimizers import Adam
from tensorflow.contrib.keras.api.keras.utils import to_categorical
import tensorflow.contrib.lite as lite
(x_train, y_train), (x_test, y_test) = mnist.load_data()
x_train = x_train.reshape(60000, 784)
x_test = x_test.reshape(10000, 784)
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
x_train /= 255
x_test /= 255
y_train = to_categorical(y_train, 10)
y_test = to_categorical(y_test, 10)
model = Sequential()
model.add(Dense(512, input_shape=(784,)))
model.add(Activation("relu"))
model.add(Dropout(0.2))
model.add(Dense(512))
model.add(Activation("relu"))
model.add(Dropout(0.2))
model.add(Dense(10))
model.add(Activation("softmax"))
model.compile(loss="categorical_crossentropy", optimizer=Adam(), metrics=['accuracy'])
model.fit(x_train,y_train, batch_size=128, epochs=20, verbose=1, validation_data=(x_test, y_test))
model.save('mnist_model.h5')
converter = lite.TFLiteConverter.from_keras_model_file('mnist_model.h5')
tflite_model = converter.convert()
open('mnist_model.tflite', 'wb').write(tflite_model)
```
At the first, import packages.
```
import (
"fmt"
"image"
_ "image/png"
"log"
"os"
)
```
And go-tflite
```
import "github.com/mattn/go-tflite"
```
Load tflite model file that is generated by Python.
```
model := tflite.NewModelFromFile("mnist_model.tflite")
if model == nil {
log.Fatal("cannot load model")
}
```
Now you can create interpreter of TensorFlow Lite. The second argument is InterpreterOptions. If you want to specify thread number of the interpreter, modify options that created by NewInterpreterOptions.
```go
options := tflite.NewInterpreterOptions()
options.SetNumThread(4)
defer options.Delete()
interpreter := tflite.NewInterpreter(model, options)
```
```
interpreter := tflite.NewInterpreter(model, nil)
```
Allocate tensors according to the model.
```
if status := interpreter.AllocateTensors(); status != tflite.OK {
log.Fatal("cannot allocate tensors")
}
```
Then, load an image file

```
f, err := os.Open("3.png")
if err != nil {
log.Fatal(err)
}
img, _, err := image.Decode(f)
if err != nil {
log.Fatal(err)
}
```
Mnist images are cropped as 28x28 images. So the input image must be resized.
```
import "github.com/nfnt/resize"
resized := resize.Resize(28, 28, img, resize.NearestNeighbor)
```
You can update this slice directly. It can use built-in function copy().
```
in := interpreter.GetInputTensor(0).Float32s()
for y := 0; y < 28; y++ {
for x := 0; x < 28; x++ {
r, g, b, _ := resized.At(x, y).RGBA()
in[y*28+x] = (float32(b) + float32(g) + float32(r)) / 3.0 / 65535.0
}
}
```
Then, call `Invoke()`
```
if status := interpreter.Invoke(); status != tflite.OK {
log.Fatal("invoke failed")
}
```
Now you can access output tensor.
```
out := interpreter.GetOutputTensor(0).Float32s()
```
`out` is slice of float32 numbers.
```
out
```
The wanted number is encodeded as offset that Python encoded this number as 1 hot. Then, add function to get index of item which is largest number.
```
func top(a []float32) int {
t := 0
m := float32(0)
for i, e := range a {
if i == 0 || e > m {
m = e
t = i
}
}
return t
}
```
Predicted number is...
```
fmt.Sprintf("The number written on the image is: %v", top(out))
```
| github_jupyter |
```
!wget https://hgdownload.soe.ucsc.edu/goldenPath/hg38/bigZips/hg38.fa.gz
!wget https://hgdownload.soe.ucsc.edu/goldenPath/felCat9/bigZips/felCat9.fa.gz
!gunzip hg38.fa.gz
!gunzip felCat9.fa.gz
!pip install biopython==1.72
import Bio
import numpy as np
from tqdm import tqdm
from Bio import SeqIO, Seq
from Bio.Alphabet import IUPAC
hg38_genome_file = '/content/hg38.fa'
h38 = SeqIO.index(hg38_genome_file,'fasta', alphabet=IUPAC.unambiguous_dna)
h38_chromosomes = [k for k in h38.keys()]
dm4R6_genome_file = '/content/felCat9.fa'
dm4R6 = SeqIO.index(dm4R6_genome_file,'fasta', alphabet=IUPAC.unambiguous_dna)
dm4R6_chromosomes = [k for k in dm4R6.keys()]
h38_chromosomes
def get_random_sequence(genome, chromosomes, l, N) :
"""
Get a random sequence of nucleotides from the genome. It will
be a sequence of N words of l letters.
"""
# first, pick a random chromosome
chromosome = genome[np.random.choice(chromosomes)]
# then pick a random spot in the chromosome
found = False
chromosome_size = len(chromosome.seq)
while not found :
start = np.random.choice(chromosome_size)
myseq = chromosome.seq[start:start+l*N]
# here we take care that there are no 'N's in the sequence,
# and that the length of the sequence is good
if 'N' not in myseq and len(myseq)==l*N :
found = True
s = str(myseq).upper()
return [s[0+i:l+i] for i in range(0, l*N, l)]
def sequence_encoder(sequence, mapping) :
l = len(sequence)
return np.sum([4**(l-i-1)*mapping[sequence[i]] for i in range(l)])
l = 4
N = 10
s = get_random_sequence(dm4R6, dm4R6_chromosomes, l, N)
mapping = {
'A' : 0,
'T' : 1,
'C' : 2,
'G' : 3
}
sequence_encoder('GCC', mapping)
ndata = 100000
l = 7
N = 80
human_fname = '/content/human.dataset'
droso_fname = '/content/cat.dataset'
fh = open(human_fname, 'w')
fd = open(droso_fname, 'w')
for n in tqdm(range(ndata)) :
human_seq = get_random_sequence(h38, h38_chromosomes, l, N)
droso_seq = get_random_sequence(dm4R6, dm4R6_chromosomes, l, N)
for i in range(N) :
fh.write('%d\t'%(sequence_encoder(human_seq[i], mapping)))
fd.write('%d\t'%(sequence_encoder(droso_seq[i], mapping)))
fh.write('\n')
fd.write('\n')
fh.close()
fd.close()
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Embedding
from tensorflow.keras.layers import LSTM
from tensorflow.keras.utils import to_categorical
from keras.callbacks import ModelCheckpoint
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.lines import Line2D
!grep -v '^>' /content/hg38.fa >/content/hg38.txt
f = open("/content/hg38.txt", "r")
print(f.readline())
for x in f:
print(x)
f = open("/content/hg38.txt", "r")
li=[]
for x in f:
li.append(x)
li
list3=[]
def convertOpposite(str):
ln = len(str)
# Conversion according to ASCII values
for i in range(ln):
if str[i] >= 'A' and str[i] <= 'Z':
# Convert lowercase to uppercase
str[i] = chr(ord(str[i]) + 32)
# Driver code
if __name__ == "__main__":
#f = open("/content/hg38.txt", "r")
#str=list(f)
for i in range(len(list2)):
for x in list2:
lower=convertOpposite(x)
list3.append(lower)
#= list2[i]
#convertOpposite(str1)
#str1 = ''.join(str1)
#print(str1)
f = open("/content/hg38.txt", "r")
str1=list(f)
human="""'taaccctaaccctaaccctaaccctaaccctaaccctaaccctaacccta\n',
'accctaaccctaaccctaaccctaaccctaaccctaaccctaaccctaac\n',
'cctaacccaaccctaaccctaaccctaaccctaaccctaaccctaacccc\n',
'taaccctaaccctaaccctaaccctaacctaaccctaaccctaaccctaa\n',
'ccctaaccctaaccctaaccctaaccctaacccctaaccctaaccctaaa\n',
'ccctaaaccctaaccctaaccctaaccctaaccctaaccccaaccccaac\n',
'cccaaccccaaccccaaccccaaccctaacccctaaccctaaccctaacc\n',
'ctaccctaaccctaaccctaaccctaaccctaaccctaacccctaacccc\n',
'taaccctaaccctaaccctaaccctaaccctaaccctaacccctaaccct\n',
'aaccctaaccctaaccctcgcggtaccctcagccggcccgcccgcccggg\n',
'tctgacctgaggagaactgtgctccgccttcagagtaccaccgaaatctg\n',
'tgcagaggacaacgcagctccgccctcgcggtgctctccgggtctgtgct\n',
'gaggagaacgcaactccgccgttgcaaaggcgcgccgcgccggcgcaggc\n',
'gcagagaggcgcgccgcgccggcgcaggcgcagagaggcgcgccgcgccg\n',
'gcgcaggcgcagagaggcgcgccgcgccggcgcaggcgcagagaggcgcg\n',
'ccgcgccggcgcaggcgcagagaggcgcgccgcgccggcgcaggcgcaga\n',
'cacatgctagcgcgtcggggtggaggcgtggcgcaggcgcagagaggcgc\n',
'gccgcgccggcgcaggcgcagagacacatgctaccgcgtccaggggtgga\n',
'ggcgtggcgcaggcgcagagaggcgcaccgcgccggcgcaggcgcagaga\n',
'cacatgctagcgcgtccaggggtggaggcgtggcgcaggcgcagagacgc\n',
'aagcctacgggcgggggttgggggggcgtgtgttgcaggagcaaagtcgc\n',
'acggcgccgggctggggcggggggagggtggcgccgtgcacgcgcagaaa\n',
'ctcacgtcacggtggcgcggcgcagagacgggtagaacctcagtaatccg\n',
'aaaagccgggatcgaccgccccttgcttgcagccgggcactacaggaccc\n',
'gcttgctcacggtgctgtgccagggcgccccctgctggcgactagggcaa\n',
'ctgcagggctctcttgcttagagtggtggccagcgccccctgctggcgcc\n',
'ggggcactgcagggccctcttgcttactgtatagtggtggcacgccgcct\n',
'gctggcagctagggacattgcagggtcctcttgctcaaggtgtagtggca\n',
'gcacgcccacctgctggcagctggggacactgccgggccctcttgctCCA\n',
'ACAGTACTGGCGGATTATAGGGAAACACCCGGAGCATATGCTGTTTGGTC\n',
'TCAGtagactcctaaatatgggattcctgggtttaaaagtaaaaaataaa\n',
'tatgtttaatttgtgaactgattaccatcagaattgtactgttctgtatc\n',
'ccaccagcaatgtctaggaatgcctgtttctccacaaagtgtttactttt\n',
'ggatttttgccagtctaacaggtgaAGccctggagattcttattagtgat\n',
'ttgggctggggcctggccatgtgtatttttttaaatttccactgatgatt\n',
'ttgctgcatggccggtgttgagaatgactgCGCAAATTTGCCGGATTTCC\n',
'TTTGCTGTTCCTGCATGTAGTTTAAACGAGATTGCCAGCACCGGGTATCA\n',
'TTCACCATTTTTCTTTTCGTTAACTTGCCGTCAGCCTTTTCTTTGACCTC\n',
'TTCTTTCTGTTCATGTGTATTTGCTGTCTCTTAGCCCAGACTTCCCGTGT\n',
'CCTTTCCACCGGGCCTTTGAGAGGTCACAGGGTCTTGATGCTGTGGTCTT\n',
'CATCTGCAGGTGTCTGACTTCCAGCAACTGCTGGCCTGTGCCAGGGTGCA\n',
'AGCTGAGCACTGGAGTGGAGTTTTCCTGTGGAGAGGAGCCATGCCTAGAG\n',
'TGGGATGGGCCATTGTTCATCTTCTGGCCCCTGTTGTCTGCATGTAACTT\n',
'AATACCACAACCAGGCATAGGGGAAAGATTGGAGGAAAGATGAGTGAGAG\n',
'CATCAACTTCTCTCACAACCTAGGCCAGTAAGTAGTGCTTGTGCTCATCT\n',
'CCTTGGCTGTGATACGTGGCCGGCCCTCGCTCCAGCAGCTGGACCCCTAC\n',
'CTGCCGTCTGCTGCCATCGGAGCCCAAAGCCGGGCTGTGACTGCTCAGAC\n',
'CAGCCGGCTGGAGGGAGGGGCTCAGCAGGTCTGGCTTTGGCCCTGGGAGA\n',
'GCAGGTGGAAGATCAGGCAGGCCATCGCTGCCACAGAACCCAGTGGATTG\n',
'GCCTAGGTGGGATCTCTGAGCTCAACAAGCCCTCTCTGGGTGGTAGGTGC\n',
'AGAGACGGGAGGGGCAGAGCCGCAGGCACAGCCAAGAGGGCTGAAGAAAT\n',
'GGTAGAACGGAGCAGCTGGTGATGTGTGGGCCCACCGGCCCCAGGCTCCT\n',
'GTCTCCCCCCAGGTGTGTGGTGATGCCAGGCATGCCCTTCCCCAGCATCA\n',
'GGTCTCCAGAGCTGCAGAAGACGACGGCCGACTTGGATCACACTCTTGTG\n',
'AGTGTCCCCAGTGTTGCAGAGGTGAGAGGAGAGTAGACAGTGAGTGGGAG\n',
'TGGCGTCGCCCCTAGGGCTCTACGGGGCCGGCGTCTCCTGTCTCCTGGAG\n',
'AGGCTTCGATGCCCCTCCACACCCTCTTGATCTTCCCTGTGATGTCATCT\n',
'GGAGCCCTGCTGCTTGCGGTGGCCTATAAAGCCTCCTAGTCTGGCTCCAA\n',
'GGCCTGGCAGAGTCTTTCCCAGGGAAAGCTACAAGCAGCAAACAGTCTGC\n',
'ATGGGTCATCCCCTTCACTCCCAGCTCAGAGCCCAGGCCAGGGGCCCCCA\n',
'AGAAAGGCTCTGGTGGAGAACCTGTGCATGAAGGCTGTCAACCAGTCCAT\n',
'AGGCAAGCCTGGCTGCCTCCAGCTGGGTCGACAGACAGGGGCTGGAGAAG\n',
'GGGAGAAGAGGAAAGTGAGGTTGCCTGCCCTGTCTCCTACCTGAGGCTGA\n',
'GGAAGGAGAAGGGGATGCACTGTTGGGGAGGCAGCTGTAACTCAAAGCCT\n',
'TAGCCTCTGTTCCCACGAAGGCAGGGCCATCAGGCACCAAAGGGATTCTG\n',
'CCAGCATAGTGCTCCTGGACCAGTGATACACCCGGCACCCTGTCCTGGAC\n',
'ACGCTGTTGGCCTGGATCTGAGCCCTGGTGGAGGTCAAAGCCACCTTTGG\n',
'TTCTGCCATTGCTGCTGTGTGGAAGTTCACTCCTGCCTTTTCCTTTCCCT\n',
'AGAGCCTCCACCACCCCGAGATCACATTTCTCACTGCCTTTTGTCTGCCC\n',
'AGTTTCACCAGAAGTAGGCCTCTTCCTGACAGGCAGCTGCACCACTGCCT\n',
'GGCGCTGTGCCCTTCCTTTGCTCTGCCCGCTGGAGACGGTGTTTGTCATG\n',
'GGCCTGGTCTGCAGGGATCCTGCTACAAAGGTGAAACCCAGGAGAGTGTG\n',
'GAGTCCAGAGTGTTGCCAGGACCCAGGCACAGGCATTAGTGCCCGTTGGA\n',
'GAAAACAGGGGAATCCCGAAGAAATGGTGGGTCCTGGCCATCCGTGAGAT\n',
'CTTCCCAGGGCAGCTCCCCTCTGTGGAATCCAATCTGTCTTCCATCCTGC\n',
'GTGGCCGAGGGCCAGGCTTCTCACTGGGCCTCTGCAGGAGGCTGCCATTT\n',
'GTCCTGCCCACCTTCTTAGAAGCGAGACGGAGCAGACCCATCTGCTACTG\n',
'CCCTTTCTATAATAACTAAAGTTAGCTGCCCTGGACTATTCACCCCCTAG\n',
'TCTCAATTTAAGAAGATCCCCATGGCCACAGGGCCCCTGCCTGGGGGCTT\n',
'GTCACCTCCCCCACCTTCTTCCTGAGTCATTCCTGCAGCCTTGCTCCCTA\n',
'ACCTGCCCCACAGCCTTGCCTGGATTTCTATCTCCCTGGCTTGGTGCCAG\n',
'TTCCTCCAAGTCGATGGCACCTCCCTCCCTCTCAACCACTTGAGCAAACT\n',
'CCAAGACATCTTCTACCCCAACACCAGCAATTGTGCCAAGGGCCATTAGG\n',
'CTCTCAGCATGACTATTTTTAGAGACCCCGTGTCTGTCACTGAAACCTTT\n',
'TTTGTGGGAGACTATTCCTCCCATCTGCAACAGCTGCCCCTGCTGACTGC\n',
'CCTTCTCTCCTCCCTCTCATCCCAGAGAAACAGGTCAGCTGGGAGCTTCT\n',
'GCCCCCACTGCCTAGGGACCAACAGGGGCAGGAGGCAGTCACTGACCCCG\n',
'AGACGTTTGCATCCTGCACAGCTAGAGATCCTTTATTAAAAGCACACTGT\n',
'TGGTTTCTGCTCAGTTCTTTATTGATTGGTGTGCCGTTTTCTCTGGAAGC\n',
'CTCTTAAGAACACAGTGGCGCAGGCTGGGTGGAGCCGTCCCCCCATGGAG\n',
'CACAGGCAGACAGAAGTCCCCGCCCCAGCTGTGTGGCCTCAAGCCAGCCT\n',
'TCCGCTCCTTGAAGCTGGTCTCCACACAGTGCTGGTTCCGTCACCCCCTC\n',
'CCAAGGAAGTAGGTCTGAGCAGCTTGTCCTGGCTGTGTCCATGTCAGAGC\n',
'AACGGCCCAAGTCTGGGTCTGGGGGGGAAGGTGTCATGGAGCCCCCTACG\n',
'ATTCCCAGTCGTCCTCGTCCTCCTCTGCCTGTGGCTGCTGCGGTGGCGGC\n',
'AGAGGAGGGATGGAGTCTGACACGCGGGCAAAGGCTCCTCCGGGCCCCTC\n',
'ACCAGCCCCAGGTCCTTTCCCAGAGATGCCTGGAGGGAAAAGGCTGAGTG\n',
'AGGGTGGTTGGTGGGAAACCCTGGTTCCCCCAGCCCCCGGAGACTTAAAT\n',
'ACAGGAAGAAAAAGGCAGGACAGAATTACAAGGTGCTGGCCCAGGGCGGG\n',
'CAGCGGCCCTGCCTCCTACCCTTGCGCCTCATGACCAGCTTGTTGAAGAG\n',
'ATCCGACATCAAGTGCCCACCTTGGCTCGTGGCTCTCACTGCAACGGGAA\n',
'AGCCACAGACTGGGGTGAAGAGTTCAGTCACATGCGACCGGTGACTCCCT\n',
'GTCCCCACCCCCATGACACTCCCCAGCCCTCCAAGGCCACTGTGTTTCCC\n',
'AGTTAGCTCAGAGCCTCAGTCGATCCCTGACCCAGCACCGGGCACTGATG\n',
'AGACAGCGGCTGTTTGAGGAGCCACCTCCCAGCCACCTCGGGGCCAGGGC\n',
'CAGGGTGTGCAGCAccactgtacaatggggaaactggcccagagaggtga\n',
'ggcagcttgcctggggtcacagagcaaggcaaaagcagcgctgggtacaa\n',
'gctcaAAACCATAGTGCCCAGGGCACTGCCGCTGCAGGCGCAGGCATCGC\n',
'ATCACACCAGTGTCTGCGTTCACAGCAGGCATCATCAGTAGCCTCCAGAG\n',
'GCCTCAGGTCCAGTCTCTAAAAATATCTCAGGAGGCTGCAGTGGCTGACC\n',
'ATTGCCTTGGACCGCTCTTGGCAGTCGAAGAAGATTCTCCTGTCAGTTTG\n',
'AGCTGGGTGAGCTTAGAGAGGAAAGCTCCACTATGGCTCCCAAACCAGGA\n',
'AGGAGCCATAGCCCAGGCAGGAGGGCTGAGGACCTCTGGTGGCGGCCCAG\n',
'GGCTTCCAGCATGTGCCCTAGGGGAAGCAGGGGCCAGCTGGCAAGAGCAG\n',
'GGGGTGGGCAGAAAGCACCCGGTGGACTCAGGGCTGGAGGGGAGGAGGCG\n',
'ATCTTGCCCAAGGCCCTCCGACTGCAAGCTCCAGGGCCCGCTCACCTtgc\n',
'tcctgctccttctgctgctgcttctccagctttcgctccttcatgctgcG\n',
'CAGCTTGGCCTTGCCGATGCCCCCAGCTTGGCGGATGGACTCTAGCAGAG\n',
'TGGCCAGCCACCGGAGGGGTCAACCACTTCCCTGGGAGCTCCCTGGACTG\n',
'GAGCCGGGAGGTGGGGAACAGGGCAAGGAGGAAAGGCTGCTCAGGCAGGG\n',
'CTGGGGAAGCTTACTGTGTCCAAGAGCCTGCTGGGAGGGAAGTCACCTCC\n',
'CCTCAAACGAGGAGCCCTGCGCTGGGGAGGCCGGACCTTTGGAGACTGTG\n',
'TGTGGGGGCCTGGGCACTGACTTCTGCAACCACCTGAGCGCGGGCATCCT\n',
'GTGTGCAGATACTCCCTGCTTCCTCTCTAGCCCCCACCCTGCAGAGCTGG\n',
'ACCCCTGAGCTAGCCATGCTCTGACAGTCTCAGTTGCACACACGAGCCAG\n',
'CAGAGGGGTTTTGTGCCACTTCTGGATGCTAGGGTTACACTGGGAGACAC\n',
'AGCAGTGAAGCTGAAATGAAAAATGTGTTGCTGTAGTTTGTTATTAGACC\n',
'CCTTCTTTCCATTGGTTTAATTAGGAATGGGGAACCCAGAGCCTCACTTG\n',
'TTCAGGCTCCCTCTGCCCTAGAAGTGAGAAGTCCAGAGCTCTACAGTTTG\n',
'AAAACCACTATTTTATGAACCAAGTAGAACAAGATATTTGAAATGGAAAC\n',
'TATTCAAAAAATTGAGAATTTCTGACCACTTAACAAACCCACAGAAAATC\n',
'CACCCGAGTGCACTGAGCACGCCAGAAATCAGGTGGCCTCAAAGAGCTGC\n',
'TCCCACCTGAAGGAGACGCGCTGCTGCTGCTGTCGTCCTGCCTGGCGCCT\n',
'TGGCCTACAGGGGCCGCGGTTGAGGGTGGGAGTGGGGGTGCACTGGCCAG\n',
'CACCTCAGGAGCtgggggtggtggtgggggcggtgggggtggtgTTAGTA\n',
'CCCCATCTTGTAGGTCTGAAACACAAAGTGTGGGGTGTCTAGGGAAGAAG\n',
'GTGTGTGACCAGGGAGGTCCCCGGCCCAGCTCCCATCCCAGAACCCAGCT\n',
'CACCTACCTTGAGAGGCTCGGCTACCTCAGTGTGGAAGGTGGGCAGTTCT\n',
'GGAATGGTGCCAGGGGCAGAGGGGGCAATGCCGGGGCCCAGGTCGGCAAT\n',
'GTACATGAGGTCGTTGGCAATGCCGGGCAGGTCAGGCAGGTAGGATGGAA\n',
'CATCAATCTCAGGCACCTGGCCCAGGTCTGGCACATAGAAGTAGTTCTCT\n',
'GGGACCTGCAAGATTAGGCAGGGACATGTGAGAGGTGACAGGGACCTGCA\n',
'GGGGCAGCCAACAAGACCTTGTGTGCACCTCCCATGGGTGGAATAAGGGG\n',
'CCCAACAGCCTTGACTGGAGAGGAGCTCTGGCAAGGCCCTGGGCCACTGC\n',
'ACCTGTCTCCACCTCTGTCCCACCCCTCCCACCTGCTGTTCCAGCTGCTC\n',
'TCTCTTGCTGATGGACAAGGGGGCATCAAACAGCTTCTCCTCTGTCTCTG\n',
'CCCCCAGCATCACATGGGTCTTTGTTACAGCACCAGCCAGGGGGTCCAGG\n',
'AAGACATACTTCTTCTACCTACAGAGGCGACATGGGGGTCAGGCAAGCTG\n',
'ACACCCGCTGTCCTGAGCCCATGTTCCTCTCCCACATCATCAGGGGCACA\n',
'GCGTGCACTGTGGGGTCCCAGGCCTCCCGAGCCGAGCCACCCGTCACCCC\n',
'CTGGCTCCTGGCCTATGTGCTGTACCTGTGTCTGATGCCCTGGGTCCCCA\n',
'CTAAGCCAGGCCGGGCCTCCCGCCCACACCCCTCGGCCCTGCCCTCTGGC\n',
'CATACAGGTTCTCGGTGGTGTTGAAGAGCAGCAAGGAGCTGACAGAGCTG\n',
'ATGTTGCTGGGAAGACCCCCAAGTCCCTCTTCTGCATCGTCCTCGGGCTC\n',
'CGGCTTGGTGCTCACGCACACAGGAAAGTCCTTCAGCTTCTCCTGAGAGG\n',
'GCCAGGATGGCCAAGGGATGGTGAATATTTGGTGCTGGGCCTAATCAGCT\n',
'GCCATCCCATCCCAGTCAGCCTCCTCTGGGGGACAGAACCCTATGGTGGC\n',
'CCCGGCTCCTCCCCAGTATCCAGTCCTCCTGGTGTGTGACAGGCTATATG\n',
'CGCGGCCAGCAGACCTGCAGGGCCCGCTCGTCCAGGGGGCGGTGCTTGCT\n',
'CTGGATCCTGTGGCGGGGGCGTCTCTGCAGGCCAGGGTCCTGGGCGCCCG\n',
'TGAAGATGGAGCCATATTCCTGCAGGCGCCCTGGAGCAGGGTACTTGGCA\n',
'CTGGAGAACACCTGTGGACACAGGGACAAGTCTGAGGGGGCCCCAAGAGG\n',
'CTCAGAGGGCTAGGATTGCTTGGCAGGAGAGGGTGGAGTTGGAAGCCTGG\n',
'GCGAGAAGAAAGCTCAAGGTACAGGTGGGCAGCAGGGCAGAGACTGGGCA\n',
'GCCTCAGAGGCACGGGGAAATGGAGGGACTGCCCAGTAGCCTCAGGACAC\n',
'AGGGGTATGGGGACTACCTTGATGGCCTTCTTGCTGCCCTTGATCTTCTC\n',
'AATCTTGGCCTGGGCCAAGGAGACCTTCTCTCCAATGGCCTGCACCTGGC\n',
'TCCGGCTCTGCTCTACCTGCTGGGAGATCCTGCCATGGAGAAGATCACAG\n',
'AGGCTGGGCTGCTCCCCACCCTCTGCACACCTCCTGCTTCTAACAGCAGA\n',
'GCTGCCAGGCCAGGCCCTCAGGCAAGGGCTCTGAAGTCAGGGTCACCTAC\n',
'TTGCCAGGGCCGATCTTGGTGCCATCCAGGGGGCCTCTACAAGGATAATC\n',
'TGACCTGCAGGGTCGAGGAGTTGACGGTGCTGAGTTCCCTGCACTCTCAG\n',
'TAGGGACAGGCCCTATGCTGCCACCTGTACATGCTATCTGAAGGACAGCC\n',
'TCCAGGGCACACAGAGGATGGTATTTACACATGCACACATGGCTACTGAT\n',
'GGGGCAAGCACTTCACAACCCCTCATGATCACGTGCAGCAGACAATGTGG\n',
'CCTCTGCAGAGGGGGAACGGAGACCGGAGGCTGAGACTGGCAAGGCTGGA\n',
'CCTGAGTGTCGTCACCTAAATTCAGACGGGGAACTGCCCCTGCACATACT\n',
'GAACGGCTCACTGAGCAAACCCCGAGTCCCGACCACCGCCTCAGTGTGGT\n',
'CTAGCTcctcacctgcttccatcctccctggtgcggggtgggcccagtga\n',
'tatcagctgcctgctgttccccagatgtgccaagtgcattcttgtgtgct\n',
'tgcatctcatggaacgccatttccccagacatccctgtggctggctccTG\n',
'ATGCCCGAGGCCCAAGTGTCTGATGCTTTAAGGCACATCACCCCACTCAT\n',
'GCTTTTCCATGTTCTTTGGCCGCAGCAAGGCCGCTCTCACTGCAAAGTTA\n',
'ACTCTGATGCGTGTGTAACACAACATCCTCCTCCCAGTCGCCCCTGTAGC\n',
'TCCCCTACCTCCAAGAGCCCAGCCCTTGCCCACAGGGCCACACTCCACGT\n',
'GCAGAGCAGCCTCAGCACTCACCGGGCACGAGCGAGCCTGTGTGGTGCGC\n',
'AGGGATGAGAAGGCAGAGGCGCGACTGGGGTTCATGAGGAAGGGCAGGAG\n',
'GAGGGTGTGGGATGGTGGAGGGGTTTGAGAAGGCAGAGGCGCGACTGGGG\n',
'TTCATGAGGAAAGGGAGGGGGAGGATGTGGGATGGTGGAGGGGCTGCAGA\n',
'CTCTGGGCTAGGGAAAGCTGGGATGTCTCTAAAGGTTGGAATGAATGGCC\n',
'TAGAATCCGACCCAATAAGCCAAAGCCACTTCCACCAACGTTAGAAGGCC\n',
'TTGGCCCCCAGAGAGCCAATTTCACAATCCAGAAGTCCCCGTGCCCTAAA\n',
'GGGTCTGCCCTGATTACTCCTGGCTCCTTGTGTGCAGGGGGCTCAGGCAT\n',
'GGCAGGGCTGGGAGTACCAGCAGGCACTCAAGCGGCTTAAGTGTTCCATG\n',
'ACAGACTGGTATGAAGGTGGCCACAATTCAGAAAGAAAAAAGAAGAGCAC\n',
'CATCTCCTTCCAGTGAGGAAGCGGGACCACCACCCAGCGTGTGCTCCATC\n',
'TTTTCTGGCTGGGGAGAGGCCTTCATCTGCTGTAAAGGGTCCTCCAGCAC\n',
'AAGCTGTCTTAATTGACCCTAGTTCCCAGGGCAGCCTCGTTCTGCCTTGG\n',
'GTGCTGACACGACCTTCGGTAGGTGCATAAGCTCTGCATTCGAGGTCCAC\n',
'AGGGGCAGTGGGAGGGAACTGagactggggagggacaaaggctgctctgt\n',
'cctggtgctcccacaaaggagaagggctgatcactcaaagttgcgaacac\n',
'caagctcaacaatgagccctggaaaatttctggaatggattattaaacag\n',
'agagtctgtaagcacttagaaaaggccgcggtgagtcccaggggccagca\n',
'ctgctcgaaatgtacagcatttctctttgtaacaggattattagcctgct\n',
'gtgcccggggaaaacatgcagcacagtgcatctcgagtcagcaggatttt\n',
'gacggcttctaacaaaatcttgtagacaagatggagctatgggggttgga\n',
'ggagagaacatataggaaaaatcagagccaaatgaaccacagccccaaag\n',
'ggcacagttgaacaatggactgattccagccttgcacggagggatctggc\n',
'agagtCCATCCAGTTCATTCAACACCTGGTTAGAAAACTGGGGCCAGCAC\n',
'ACAGGGGAAGGGTAAGCTGGTTTCATGATCGAATCAAGGCTCAGACAATT\n',
'TTTAAAGGCCAGAGGGTAGACTGCAATCACcaagatgaaatttacaagga\n',
'acaaatgtgaagcccaacatttaggttttaaaaatcaagcgtataaatac\n',
'agaaggtggagggaacttgctttagacacagttcaggtgaagaaagacct\n',
'ggaaacttctgttaactataagctcagtaGGGGCTAAAAGCATGTTAATC\n',
'GGCATAAAAAGGCAATGAGATCTTAGGGCACACAGCTCCCCGCCCCTCTT\n',
'CTGCCCTTCATCCTTCTTTCAATCAGCAGGGACCGTGCACTCTCTTGGAG\n',
'CCACCACAGAAAACAGAGGTGCATCCAGCACCACAGAAAACAGAGCCACC\n',
'ACAGAAAACAGAGGGTGACTGTCATCCCCTCCAGTCTCTGCACACTCCCA\n',
'GCTGCAGCAGAGCAGGAGGAGAGAGCACAGCCTGCAATGCTAATTTGCCA\n',
'GGAGCTCACCTGCCTGCGTCACTGGGCACAGACGCCAGTGAGGCCAGAGG\n',
'CCGGGCTGTGCTGGGGCCTGAGCCGGGTGGTGGGGAGAGAGTCTCTCCCC\n',
'TGCCCCTGTCTCTTCCGTGCAGGAGGAGCATGTTTAAGGGGACGGGTTCA\n',
'AAGCTGGTCACATCCCCACCGAAAAAGCCCATGGACAACGAAAAGCCCAC\n',
'TAGCTTGTCCAGTGCCACAGGAGGGGCAAGTGGAGGAGGAGAGGTGGCGG\n',
'TGCTCCCCACTCCACTGCCAGTCGTCACTGGCTCTCCCTTCCCTTCATCC\n',
'TCGTTCCCTATCTGTCACCATTTCCTGTCGTCGTTTCCTCTGAATGTCTC\n',
'ACCCTGCCCTCCCTGCTTGCAAGTCCCCTGTCTGTAGCCTCACCCCTGTC\n',
'GTATCCTGACTACAATAACAGCTTCTGGGTGTCCCTGGCATCCACTCTCT\n',
'CTCCCTTCTTGTCCCTTCCGTGACGGATGCCTGAGGAACCTTCCCCAAAC\n',
'TCTTCTGTCCCATCCCTGCCCTGCTCAAAATCCAATCACAGCTCCCTAAC\n',
'ACGCCTGAATCAACTTGAAGTCCTGTCTTGAGTAATCCGTGGGCCCTAAC\n',
'TCACTCATCCCAACTCTTCACTCACTGCCCTGCCCCACACCCTGCCAGGG\n',
'AGCCTCCCGTGGCACCGTGGGGACACAAAGGAACCAGGGCAAAGCTCCCT\n',
'CAGCCCCATTCAAAGAGGCCTGGCCCACAGGCTCACGGAAAGTCAGCCTC\n',
'TCATGCCCCGAGAGCTGAGTGCAAGGGAGAGGCAGCGCTGTCTGTGCTTC\n',
'CCATGCAGAAGCACCCCCCTCCCACCCCTGTGCAGGCCGGCCTTCGCGGC\n',
'AGACCACCATACACCACGTTCCAAGCCACACTGAGGCCTCCCTCCAAGCC\n',
'TGCAGCCCCCATTTCCAGACCCTGCCAGGGCAACCTGCATATCCACCTCC\n',
'CTACCCTGCCCCCCTCTTCCAGGAGTCTGCCCTATGTGGAGTAAGCACgt\n',
'ggttttcctcttcagcaactatttcctttttactcaagcaatggccccat\n',
'ttcccttggggaatccatctctctcgcaggcttagtcccagagcttcagg\n',
'tggggctgcccacagagctcctcagTCTAAGCCAAGTGGTGTGTCATAGT\n',
'CCCCTGGCCCCATTAATGGATTCTGGGATAGACATGAGGACCAAGCCAGG\n',
'TGGGATGAGTGAGTGTGGCTTCTGGAGGAAGTGGGGACACAGGACAGCAT\n',
'TCTTTCCTGCTGGACCTGACCCTGTGTCATGTCACCTTGCTACCACGAGA\n',
'GCATGGCCTGTCTGGGAATGCAGCCAGACCCAAAGAAGCAAACTGACATG\n',
'GAAGGAAAGCAAAACCAGGCCCTGAGGACATCATTTTAGCCCTTACTCCG\n',
'AAGGCTGCTCTACTGATTGGTTAATTTTTGCTTAGCTTGGTCTGGGGAGT\n',
'TCTGACAGGCGTGCCACCAATTCTTACCGATTTCTCTCCACTCTAGACCC\n',
'TGAGAAGCCCACGCGGTTCATGCTAGCAATTAACAATCAATCTCGCCCTA\n',
'TGTGTTCCCATTCCAGCCTCTAGGACACAGTGGCAGCCACATAATTGGTA\n',
'TCTCTTAAGGTCCAGCACGAGGTGGAGCACATGGTGGAGAGACAGATGCA\n',
'GTGACCTGGAACCCAGGAGTGAGGGAGCCAGGACTCAGGCCCAAGGCTCC\n',
'TGAGAGGCATCTGGCCCTCCCTGCGCTGTGCCAGCAGCTTGGAGAACCCA\n',
'CACTCAATGAACGCAGCACTCCACTACCCAGGAAATGCCTTCCTGCCCTC\n',
'TCCTCATCCCATCCCTGGGCAGGGGACATGCAACTGTCTACAAGGTGCCA\n',
'AGTACCAGGACAGGAAAGGAAAGACGCCAAAAATCCAGCGCTGCCCTCAG\n',
'AGAAGGGCAACCACGCAGTCCCCATCTTGGCAAGGAAACACAATTTCCGA\n',
'GGGAATGGTTTTGGCCTCCATTCTAAGTGCTGGACATGGGGTGGCCATAA\n',
'TCTGGAGCTGATGGCTCTTAAAGACCTGCATCCTCTTCCCTAGGTGTCCC\n',
'TCGGGCACATTTAGCACAAAGATAAGCACAAAAGGTGCATCCAGCACTTT\n',
'GTTACTATTGGTGGCAGGTTTATGAATGGCAACCAAAGGCAGTGTACGGG\n',
'TCAAGATTATCAACAGGGAagagatagcatttcctgaaggcttcctaggt\n',
'gccaggcactgttccattcctttgcatgttttgattaatttaatatttaa\n',
'aataattctaccaggaagctaccattattaccacaacttcacaaatgaga\n',
'acaccgaggcttagaggggttgggttgcccaaggttacagaggaagaaaa\n',
'caggggagctggatctgagccaaggcatcaactccaaggtaacccctcag\n',
'tcacttcactgtgtgtcccctGGTTACTGGGACATTCTTGACAAACTCGG\n',
'GGCAAGCCGGTGAGTCAGTGGGGGAGGACTTTCAGGAAGAGGTGGGTTCC\n',
'CAGTTGGTGACAGAAGAGGAGGCTGCAAAGTGAAGGAGCAGGGGCTCCAG\n',
'GTCTGGCGACAACCAGGGAAGGGACAGGGCAGGGATGGCTTGGACCACGA\n',
'GAGGCACCTGAGTCAGGCAGTCACATACTTCCCACTGGGGTCTACCATGT\n',
'GAGGCATGGTGTGGGATCCTGGGAAGGAGACCAAGCCTCATTTCAGTTTG\n',
'CTTATGGCCAAAGACAGGACCTGTGTACCCGACAACCCCTGGGACCTTTA\n',
'CCAAAAAAAGAGCAAACACCATTCACTCACTCATGTTAGATAAACACTGA\n',
'GTGAAGTCACTGGAGCCCAAGGACTGTGCGAGGTCAGCACTGCCAATACA\n',
'AGAagctgcagccctccagctcgcctccctcaatggccactccgtgctcc\n',
'agccatgctggcttccttttaggtcctccacctccaggctgtagttcatg\n',
'tgcttctttctggaatgttcttcccaacctacccactcaaccctcagact\n',
'ttaccataaatgtcatttcctcacgtctgccttccctgacctgagaccaa\n',
'gccaggcttcccatgacgagcctcacagtaccccatctCCCCTGAACAGA\n',
'TGCAGTAATAACCTACATAACCCGGGGCCATGATCTAtggctttgaatcc\n',
'tggctctgtcactaggccaggtctctcagcccttctgtgcctcagtttcc\n',
'tcatctataaaatgagatgacggcagtgcctgctcatgaagtgtgagtta\n',
'atgcactcaaatcaatggttgtgcacggtttatatgaatattagtgatta\n',
'CAAAATATTATCAATAGACCTTGTCACAACTGTTATTGAAGAACtaatca\n',
'tctattgcttatttaggtctttctctcctgccagaatgtgcgctccaggt\n',
'ggagaggtatgttgccttatccgtggctggatatatagagattcccacac\n',
'tgccttgcacacgagcactgctgggtaaatatttgttggctgcaggaaAA\n',
'CGTGAAGGAATAGGCCCTCCAATGGGAGGAAAAGCATGAGTTGTGAGAGC\n',
'AGAGCCACCACAGGAAACCAGGAGGCTAAGTGGGGTGGAAGGGAGTGAGC\n',
'TCTCGGACTCCCAGGAGTAAAAGCTTCCAAGTTGGGCTCTCACTTCAGCC\n',
'CCTCCCACACAGGGAAGCCAGATGGGTTCCCCAGGACCGGGATTCCCCAA\n',
'GGGGGCTGCTCCCAGAGGGTGTGTTGCTGGGATTGCCCAGGACAGGGATG\n',
'GCCCTCTCATCAGGTGGGGGTGAGTGGCAGCACCCACCTGCTGAAGATGT\n',
'CTCCAGAGACCTTCTGCAGGTACTGCAGGGCATCCGCCATCTGCTGGACG\n',
'GCCTCCTCTCGCCGCAGGTCTGGCTGGATGAAGGGCACGGCATAGGTCTG\n',
'ACCTGCCAGGGAGTGCTGCATCCTCACAGGAGTCATGGTGCCTGTGGGTC\n',
'GGAGCCGGAGCGTCAGAGCCACCCACGACCACCGGCACGCCCCCACCACA\n',
'GGGCAGCGTGGTGTTGAGACAACACAGCCCTCATCCCAACTATGCACATA\n',
'GCTTCAGCCTGCACAGATAGGGGAGTAGGGGACAGAGCATTTGCTGAGAG\n',
'GCCAGGAGCGCATAGATGGGACTCTGCTGATGCCTGCTGAGTGAATGAGG\n',
'GAAAGGGCAGGGCCCGGGACTGGGGAATCTGTAGGGTCAATGGAGGAGTT\n',
'CAGAGAAGGTGCAACATTTCTGACCCCCTACAAGGTGCTTGCTACCTGCC\n',
'AGGCACCCTTTCCATACCTTGTCTCAGTTCAGCTCCCCACCTTGGATAAA\n',
'CAAGAAACCTTGGTTGCAGAGGAAAAAAGAGGCTGGAAACAAAGGGGTAG\n',
'AAATGGGGTAGCAGGGGAGATTGCCTGATCAACTGCCAAATGGTACACAG\n',
'TTCTGGAAAAGCACAAAAAATGTGCACACACGGGTTCTTCCCACTTTAAC\n',
'CCCTGAGGAATCTGAGGCCTGCTCCTGAAACAGACTGGGCAGTGGCTAGT\n',
'GACTCTAGGTATAGGAGTATCCAGCCCTGCTCACCCAGGCTAGAGCTTAG\n',
'GGGGACAAGAGGAAAGAGGTGCCTGTGGGGGTGGAGGACAGGAAGGAAAA\n',
'ACACTCCTGGAATTGCAAAGTGAGGGCAGAGTCTATTTATATTGGGTTTA\n',
'ATTAACTCCTCTCCCTGGTGCCACTAAAGCAGCAATCACACTGCAGACAG\n',
'CACTGATTTGATTGGCAAGAGATGCACCAGGCAGAATATTAAGGGACCAG\n',
'GCCCCTATAAATAGGCCTAATCACAGCCCCTCACTGGAAAATGGTAAGGA\n',
'AGACATTAATCAGGCCTGGCACTGTGCCCTAGACCTGCTCCCCTAGGCAC\n',
'TACAGTGGGGCCCTTGGTTGCAACACAAGTAGGTAGGGATGGATGAGTGT\n',
'GGCATGAAGGGCCTAGGAGATTTCACTTGGGTTTAAAATGCTGTGACCTT\n',
'GAGTAAGTTGCCGTCTCTGAATCTGATCCTTTCGATTTCCCATTCTCCAA\n',
'ACTGAGAACTAGCACTGCTGAGACGTGGTTATTTCCAATAATAATTTGTA\n',
'TATTTTACATAACGCACCACACCAACATCTTCACCCAGTTGGAGCCTACT\n',
'CCTTTGCTCCCGCTGCTGGCTTCCCCAGCCCTCCCTTCTGCCCTCCTCAG\n',
'GCCAGCACTTTTCAGTGAGTTCCTCCTTTGCATACAGGCTTTCCAGATCT\n',
'GTACTTGCCTTGAATACTCATCAGAGCCCAGGAGTTACTCCTCACCTCCC\n',
'ACTTATTTTTCCTCCCATCAAATAACTAAAGCATGGCCAGCTGATGCCCA\n',
'GCCAACTGAGAAACCCAACCCTCTGAGACCAGCACACCCCTTTCAAGCAT\n',
'GTTCCTCCCTCCCCTTCTTTGTATTTATACTGATGCAAGTTTGCTGGCTG\n',
'TCCTAacttatttctgtgcctcagttctcccatatgtaagatcacaaagg\n',
'gggtaaagatgcAAGATATTTCCTGTGCACATCTTCAGATGAATTTCTTG\n',
'TTAGTGTGTGTGTGTTTGCTCACACATATGCGTGAAAGAAGAGTACATAC\n',
'ACAGATCTCCTCAAAAAGGAGGCAGCAAGCCCGTTCAAGAATGGGACTGA\n',
'ATACACCTGATGAGTGGTTTACTTTCTGTCTGcaaacatctactgatcat\n',
'ctgttaggtgcaggccatgatcacaacaaagacgaataagacactacact\n',
'agccagggagagtctcaaaaacaactaaactcaaattaaattcattctac\n',
'tccagtcatgggtacaaagctaaggagtgacaaatccctcttggagttag\n',
'gggagtcaggaaaaagctcttagcagaatgtgtgcctctcggccgggcgc\n',
'agcggctcacgcctgtaatcccagcactttgggaggcgaaggcaggcaga\n',
'tcacctgaggtcgggagttcgagaccagtctgaccaacatggtgaaactc\n',
'catctctactaaaaatacaaaattagccaggcgtggtggtgcatgcctgt\n',
'aatccccgctactcgggaggctgaggaaggagaatcacttgaaccaggaa\n',
'ggtggaggttgcagtgtgccaagatcgcgccatggcactccagcctaggc\n',
'aacgagggtgaaccaggtccaggaagaaggtgcaaagacagcattccagg\n',
'taaaagaaacagcttgaacaaaaagtgtgtaggggaaCCGCAAGCGGTCT\n',
'TGAGTGCTGAGGGTACAATCATCCTTGGGGAAGTACTAGAAGAAAGAATG\n',
'ATAAACAGAGGCCAGTTTGTTAAAAACACTCAAAATTAAAGCTAGGAGTT\n',
'TGGACTTGTGGCAGGAATgaaatccttagacctgtgctgtccaatatggt\n',
'agccaccaggcacatgcagccactgagcacttgaaatgtggatagtctga\n',
'attgagatgtgccataagtgtaaaatatgcaccaaatttcaaaggctaga\n',
'aaaaaagaatgtaaaatatcttattattttatattgattacgtgctaaaa\n',
'taaccatatttgggatatactggattttaaaaatatatcactaatttcat\n',
'ctgtttctttttacttttAGAAATCACATATGTGACTTAAATATTTCTTT\n',
'TCTTTTTCTTTCCTCTCACTCAGCGTCCTGTGATTCCAAAGAAATGAGTC\n',
'TCTGCTGTTTTTGGGCAGCAGATATCCTAGAATGGACTCTGACCTAAGCA\n',
'TCAAAATTAATCATCATAACGTTATCATTTTATGGCCCCTTCTTCCTATA\n',
'TCTGGTAGCTTTTAAATGATGACCATGTAGATAATCTTTATTGTCCCTCT\n',
'TTCAGCAGACGGTATTTTCTTATGCTACAGTATGACTGCTAATAATACCT\n',
'ACACATGTTAGAACCATTCTGACTCCTCAAGAatctcatttaactcttat\n',
'tatcagtgaatttatcatcatcccctattttacataaggaaatggggtta\n',
'gaaagaccaaataacattttttcaacatcaaaacactagcttgagatcaa\n',
'gcccagacttggatctgtcgtctgaattccaagctttttgttatttattg\n',
'atatgttttgttgtTTTCATGCAATAATGCAAATCTTAGCCCAAACATTT\n',
'TGTTAGTAGTACCAACTGTAAGTCACCTTATCTTCATACTTTGTCTTTAT\n',
'GTAAACCTAAATTAGATCTGTTTTTGATACTGAGGGAAAAACAAGGGAAT\n',
'ctaacactaaccagcccgtagtgtgtggtcaacactttcgttactttagt\n',
'atacatcaccccaattgtttgtcttcaccacacactttggagttaggtag\n',
'tagtatctatttttacaaataagaaaacccaggcacaaaggggttgatta\n',
'gcAATTATCTTTTGAAAAGCCTGTAGTTGCTCATCTGAAGAAGTGACGGA\n',
'CCACCTCTTATTTAGTGGACAGACAGTAACTAGTTGAGAAGACAGGGGAT\n',
'TTTGTTGGCGGAAAAAAAAATTTATCAAAAGTCGTCTTCTATCAGGGAGT\n',
'TTTATGAGAAACCCTAGCTCCTCAGTTCCACAGTGGGTAACTGTAATTCA\n',
'TTCTAGGTCTGCGATATTTCCTGCCTATCCATTTTGTTAACTCTTCAATG\n',
'CATTCCACAAATACCTAAGTATTCTTTAATAATGGTGGTTTTTTTTTTTT\n',
'TTTGCATCTATGAAGTTTTTTCAAATTCTTTTTAAGTGACAAAACTTGTA\n',
'CATGTGTATCGCTCAATATTTCTAGTCGACAGCACTGCTTTCGAGAATGT\n',
'AAACCGTGCACTCCCAGGAAAATGCAGACACAGCACGCCTCTTTGGGACC\n',
'GCGGTTTATACTTTCGAAGTGCTCGGAGCCCTTCCTCCAGACCGTTCTCC\n',
'CACACCCCGCTCCAGGGTCTCTCCCGGAGTTACAAGCCTCGCTGTAGGCC\n',
'CCGGGAACCCAACGCGGTGTCAGAGAAGTGGGGTCCCCTACGAGGGACCA\n',
'GGAGCTCCGGGCGGGCAGCAGCTGCGGAAGAGCCGCGCGAGGCTTCCCAG\n',
'AACCCGGCAGGGGCGGGAAGACGCAGGAGTGGGGAGGCGGAACCGGGACC\n',
'CCGCAGAGCCCGGGTCCCTGCGCCCCACAAGCCTTGGCTTCCCTGCTAGG\n',
'GCCGGGCAAGGCCGGGTGCAGGGCGCGGCTCCAGGGAGGAAGCTCCGGGG\n',
'CGAGCCCAAGACGCCTCCCGGGCGGTCGGGGCCCAGCGGCGGCGTTCGCA\n',
'GTGGAGCCGGGCACCGGGCAGCGGCCGCGGAACACCAGCTTGGCGCAGGC\n',
'TTCTCGGTCAGGAACGGTCCCGGGCCTCCCGCCCGCCTCCCTCCAGCCCC\n',
'TCCGGGTCCCCTACTTCGCCCCGCCAGGCCCCCACGACCCTACTTCCCGC\n',
'GGCCCCGGACGCCTCCTCACCTGCGAGCCGCCCTCCCGGAAGCTCCCGCC\n',
'GCCGCTTCCGCTCTGCCGGAGCCGCTGGGTCCTAGCCCCGCCGCCCCCAG\n',
'TCCGCCCGCGCCTCCGGGTCCTAACGCCGCCGCTCGCCCTCCACTGCGCC\n',
'CTCCCCGAGCGCGGCTCCAGGACCCCGTCGACCCGGAGCGCTGTCCTGTC\n',
'GGGCCGAGTCGCGGGCCTGGGCACGGAACTCACGCTCACTCCGAGCTCCC\n',
'GACGTGCACACGGCTCCCATGCGTTGTCTTCCGAGCGTCAGGCCGCCCCT\n',
'ACCCGTGCTTTCTGCTCTGCAGACCCTCTTCCTAGACCTCCGTCCTTTGT\n',
'CCCATCGCTGCCTTCCCCTCAAGCTCAGGGCCAAGCTGTCCGCCAACCTC\n',
'GGCTCCTCCGGGCAGCCCTCGCCCGGGGTGCGCCCCGGGGCAGGaccccc\n',
'agcccacgcccagggcccgcccctgccctccagccctacgccTTGACCCG\n',
'CTTTCCTGCGTCTCTCAGCCTACCTGACCTTGTCTTTACCTCTGTGGGCA\n',
'GCTCCCTTGTGATCTGCTTAGTTCCCACCCCCCTTTAAGAATTCAATAGA\n',
'Gaagccagacgcaaaactacagatatcgtatgagtccagttttgtgaagt\n',
'gcctagaatagtcaaaattcacagagacagaagcagtggtcgccaggaat\n',
'ggggaagcaaggcggagttgggcagctcgtgttcaatgggtagagtttca\n',
'ggctggggtgatggaagggtgctggaaatgagtggtagtgatggcggcac\n',
'aacagtgtgaatctacttaatcccactgaactgtatgctgaaaaatggtt\n',
'tagacggtgaattttaggttatgtatgttttaccacaatttttaaaaaGC\n',
'TAGTGAAAAGCTGGTAAAAAGAAAGAAAAGAGGCTTTTTTAAAAAGTTAA\n',
'ATATATAAAAAGAGCATCATCAGTCCAAAGTCCAGCAGTTGTCCCTCCTG\n',
'GAATCCGTTGGCTTGCCTCCGGCATTTTTGGCCCTTGCCTTTtagggttg\n',
'ccagattaaaagacaggatgcccagctagtttgaattttagataaacaac\n',
'gaataatttcgtagcataaatatgtcccaagcttagtttgggacatactt\n',
'atgctaaaaaacattattggttgtttatctgagattcagaattaagcatt\n',
'ttatattttatttgctgcctctggccaccctaCTCTCTTCCTAACACTCT\n',
'CTCCCTCTCCCAGTTTTGTCCGCCTTCCCTGCCTCCTCTTCTGGGGGAGT\n',
'TAGATCGAGTTGTAACAAGAACATGCCACTGTCTCGCTGGCTGCAGCGTG\n',
'TGGTCCCCTTACCAGAGGTAAAGAAGAGATGGATCTCCACTCAtgttgta\n',
'gacagaatgtttatgtcctctccaaatgcttatgttgaaaccctaacccc\n',
'taatgtgatggtatgtggagatgggcctttggtaggtaattacggttaga\n',
'tgaggtcatggggtggggccctcattatagatctggtaagaaaagagaGC\n',
'ATTGtctctgtgtctccctctctctctctctctctctctctcatttctct\n',
'ctatctcatttctctctctctcgctatctcatttttctctctctctcttt\n',
'ctctcctctgtcttttcccaccaagtgaggatgcgaagagaaggtggctg\n',
'tctgcaaaccaggaagagagccctcaccgggaacccgtccagctgccacc\n',
'ttgaacttggacttccaagcctccagaactgtgagggataaatgtatgat\n',
'tttaaagtcgcccagtgtgtggtattttgttTTGACTAATACAACCTGAA\n',
'AACATTTTCCCCTCACTCCACCTGAGCAATATCTGAGTGGCTTAAGGTAC\n',
'TCAGGACACAACAAAGGAGAAATGTCCCATGCACAAGGTGCACCCATGCC\n',
'TGGGTAAAGCAGCCTGGCACAGAGGGAAGCACACAGGCTCAGggatctgc\n',
'tattcattctttgtgtgaccctgggcaagccatgaatggagcttcagtca\n',
'ccccatttgtaatgggatttaattgtgcttgccctgcctccttttgaggg\n',
'ctgtagagaaaagatgtcaaagtattttgtaatctggctgggcgtggtgg\n',
'ctcatgcctgtaatcctagcactttggtaggctgacgcgagaggactgct\n',
'tgagcccaagagtttgagatcagcctgggcaatattgtgagattccatct\n',
'ctacaaaaataaaataaaatagccagtcatggtgtcacacacctgtagtc\n',
'ccagctacatgggaggctgaggcgggaggatcacttgagcttgggagatc\n',
'gaggctgcagtgagctatgattgtaccactgcactccaggctgggcgaca\n',
'gagagagaccctgtctcagaaaaaaaaaaaaaagtactttgtaatctgta\n',
'aggtTTATTTCAACACACACAAAAAAAGTGTATATGCTCCACGATGCCTG\n',
'TGAATATACACACACACCACATCATATACCAAGCCTGGCTGTGTCTTCTC\n',
'ACAAATGCACTGCTAGGCACCACCCCCAGTTCTAGAATCACACCAGCCAG\n',
'TTCACCCTCCAGATGGTTCACCCTCAACTTCATAAAAGTTCCCTACCTAA\n',
'TCTACTGACAGGCTCATCCCCGACCTAATTTTAAAGATTTCCTAGGAGCT\n',
'GCAGTGGGAATCCTGGACCTCAGCCTGGACAAAGAACAGCTGCAGGTCAT\n',
'TCTCATGTGTGGACACAGAAGCTCTGCCTGCCTTTGCTGGCCAGCTGGGC\n',
'TGAGCGGGCCTGGGAATTAAGGCTGCAGGGTTGGTCCCAGGCAGTCTTGC\n',
'TGAAGCTTGCCACATCCCCCAGCCTCCTGGATTTGCCAGGATCCAAGAGC\n',
'ATGGACTTTAGGAATTCCTGGTGGAGGAGTGAAGAAAATGTGACAGGGTG\n',
'TCCTAAGCCCCGATCTACAGGAAGAAAACTGGAAATAAGACTGAGGACTT\n',
'AGTTTAAGATGTTCCTACTCAGCCTCTAGCTTTTGTGCTACAGTTCTGGG\n',
'AACAGACTCCTCTCTCCTGAAAACCACTTCCCTCCGCAGCATTAGATTTC\n',
'ACCAAGATGTCTTGCTTGTGGGAAAGACTTCCAAGGATGCCTGGAGAGAG\n',
'GAGGATGGAAATGTCCTGCTCTCTAAACAGATAGACAGATGCAGCCAGAC\n',
'AGAAAATAGTTTATCTTGCTGAGGTTTCTAATGTATTTGAAAGAGGCCTG\n',
'GGTCTAGAAGTCTACCCAGAGGGCTCTGTGTTGTGCACGCAAAGATAAGA\n',
'ACCTTCCCTGTGGGAGTTCCAGAGCCAGTTTTCATAAACACCCATCGGTG\n',
'ACTGTGTTCAGAGTGAGTTCACACCATCCTGACCTGCCCTGAGTTAGACC\n',
'TTACATGGTCTTCCTCCTCTAGGAAGCCTCTGCAGCCCAGGAACCTCCCC\n',
'TTATCGGAAATGAACAGCATTTGAAGCTTCACCAGACAGACCAGACAGCT\n',
'TAGCCCTCGTGTTGTGCCATGTGGGTTGTTCTCTGAGAGGcaggagagca\n',
'tagtggttactaggaagggaaggactttgggactagactgcctcggctgg\n',
'agtcctctttctgcttcatagccacgtgatcctaggcatgttacctgtgc\n',
'ctcagttttcactctgtcaatatgtaataactgaatctgtctttgtggtg\n',
'aggattcagtgagttaacatatttgaagtgcttaaaaATGAGGCTTGtgt\n',
'ccatagattaatgagtgaatacacaaatggtgatatggacatacagtgga\n',
'gtattagtcataaaaaggaaggcagagctgatccatggcaccatgtgaca\n',
'gaacctcaaaagcattaggttaagtggaagaagccagacacaggtcacct\n',
'attgtgtaattccatttataggaaatatacagaatatgtaaatccgtgga\n',
'gaaagaaagccgatttccaggggctaaggggaggggagaatgggaagtgg\n',
'ctgcttcatgggtacaaggtttcattttgagctgatgaaaatgttttgga\n',
'actacatagagatagtgttggcacaacatggtgaatgtactgaatgccac\n',
'tgattgttcaatttaaaatggtcaaacttatatgaatttcacctccatta\n',
'aaaaaaAAAAAAAAGgaccagatgtggttgctcacacccataatcccaac\n',
'actttggaaAAAGGTGAAAGTTTTTTTTtctttttttttttatatactta\n',
'agttctagggtacatgtgcataatgtgcaggttggatacatagatatgcg\n',
'tgtgccatgttggtttgctgcacccatcaacttgtcatttacattaggta\n',
'tttcttctaatgctatccctcccccagccccccacccactgacaggcccc\n',
'agtgtatgatgttctctgccccatgtccaagcgttctcattgttcaattc\n',
'ccacctgtgagtgagaacatgcagtgtttggttttctgtctttgtgatag\n',
'tttgctcagaatgatggtttccagcttcatccatgtccctgcaaaggaca\n',
'tgaactcatcctttttaatggctgcatagtatcccatggtatatatgtgc\n',
'cacattctcttaatccagtctgtcattgatggacatttgggttggttcaa\n',
'agtctttgctattgtgaatactgccacaataaacatacatgtgcatgtgt\n',
'ctttatagtagcacgatttataatcctttgggtatatacccTAAGACctg\n',
'ggacgcatttaaagcagtgtgtaaagagacatttatagcactaaatgccc\n',
'acaagagaCCTCTGCCTGAGAACGTGGGTTTCAGCCTAAGAGTTGTAATA\n',
'TGTGTGCCCATTCACAGGTGCTGCATCAGAGTCCCAGGTGGGAAGAAGGC\n',
'AAGCATACACAAAAATGGTAAAAGGCAGAAAGGAGCCCAGTCTCGTTCTT\n',
'TTTAAGAAGTTTTCCTAAGAATCTCCACCCAGCGACTTGCTCTCACATCT\n',
'TCTTGGCCAGCACTGGACCACACAACTCCTTCTAGATACAGAGGAGTCCT\n',
'AGGATTCTATGAGAAAGAAGGGGAGGGTGGGCAAAGGGCAGCCAGCTGTG\n',
'CAGCATCTGCTGGAGACACCTAACCCTTGGTGGAGGGGTTGTGGTGCTGG\n',
'gagaaggctttctggacggtgtgacagcagagataaacttaaaggccaag\n',
'taggagttaccctggtgaagcagggcagggttacaagcattccagcaaca\n',
'tgaagcagcaGGAGtgttttaattaaaagaaggcagttgctgtaaccaac\n',
'tataaacaaataaaggcttaaacacaatggaagtttatttctcactaagg\n',
'gaacatccaaatccatgatactttaagtcagggacccaggttcctcccat\n',
'ctatggttctgccatcactaatctgggtcttccacaattgccgtgctcct\n',
'tggaggtgggaagagcaggcggaggacacgtgggaggttttagggacaag\n',
'cctggaggcagcatgcgtcactcccatgcagagtccattggccaatgctg\n',
'gctccgatggccacatctcactgcaggggcagctgggaaatacagtctgg\n',
'ctgtctacccaggaggaagagCAGCCAGTTTCTGCTGCTGATGATCAGGA\n',
'GGTGGAGAAAATGTTCAGTCAGGCAGGGAGTGGGAATAGACAAGACCACA\n',
'AGCAGCTTGGTGCCTCTGAAAGGGAGAGGGGTGGAGGGGAGACTAGAGAG\n',
'GTGGGTAGGAATACTGGATTCCACTGACCACGTGCTGGATGTCACGCTTA\n',
'GCCCTCCTGCTCTGTGCCGGGTTAGGCACCTGGTGTTTTACGTACATAAT\n',
'CTCAATTCTGTGAGGGCATCCGACCTGTGGGAAAAGAGCTGTTTGTTTCA\n',
'AATGCCAGTCCTGCTTcctaacaagtgtttagagcttaatcgtgttcaaa\n',
'atacatatacaatgtttaatacttacaagaatttggtggggaaaatatta\n',
'ccatctttcccttttgtgattggagaaaaatgaggctttgaagggtttaa\n',
'gaacttgcccaaggtcggccaggtgcagtggctcatgtctataatcccaa\n',
'cactttgggaggctgaggtgggaggatcgcttgaggccaggagttcaaga\n',
'ccagcctgagcaacatagtgagactttgtctctataaaaaataaataaaT\n',
'AAATAAAAACAACTTGTCCAAGGTCAGACAGGCAGCCTCTTAGTAAGCAC\n',
'ACATATCCTCTATATTATACTACCTCTCATGGAGGATCTCCTGTGTTCTA\n',
'CAAATAGTCTGGACTTGAGCCAGAATGTGTTATAATCCTGGGATCACGGC\n',
'CAGTGGGCTTAGAAGAAGCCATCTCTTTCTCATGCCAAGATGAGGCTCCC\n',
'CCAGATTTGCTCAGACTTACCTATAGTCAGCAGCATCGGGGGTCAGGAAA\n',
'GACTTCACGAAGCCATAAATGCATCCTTCTCGGGGCAGCACCTGGCTCTC\n',
'CCAGGTGAGAGAGGACTCCATTTTCACAGGCAGGCGTGGGAGCTTCAGCA\n',
'CCCATCTCTGGGCCCAGAATGACCCACTGGAGACCTTACAGCTCTCCTGT\n',
'CACCCCCAATTCCTGCCCCCTCTGCAGCCTTGGAGGAGAATGGAGCTGAA\n',
'GGGCCTGCCCTCTGTAGGGTGAGAAAGGGAGGCTAAAGCCTGGTGCCCAC\n',
'TGCCCTGGCTGCTCCGCATTGCAGGAGCTGCGCCCTTCCTTTCCTGGCAC\n',
'AGGGTCCACAGCCCCGAAACCCCGTTGTGTGGGAGCTGGGCACAGGGCAG\n',
'CAGGACTAATCCTTGGAACAGCTCAGGGAGGATTATCCCAGCCACTGTCA\n',
'GCAGCGGTGCAGCTGGCTCATTCCCATATAGGGGGAGGCCAGAGCCAGGG\n',
'GCCTGCCACAAGTTGGAAGGCTGGGGAAGGGGAGGCCAGCAGAGGTGTCC\n',
'TGGCTGTGGGTGGCTCTGAGGGGGCTCTCAGGGGTGGGGCTAAATCTCAG\n',
'GGGCAGGATTATGTAAATCAAACCAATTCTAGCCACAGATTTAAAGTTTG\n',
'GAAAAAAAAAAAAACCCAGCCTGGCGGAAAGAATTTAAATTATAAAAACT\n',
'TAGAAGTATGGAATGTGAAATCATCCTGTAGGTGCTTATTTAACAACGAA\n',
'ATCATCCCGACACAATGAGCCATATGTGAAAAGTCATCCTTCCCCAACAC\n',
'ATCCCCCAACAGGCACTCCTCAAGCCTCTCCCACCCAAGTGCTGGCATCC\n',
'TCCCTGTCCTGCTTCACCTGAGACACCCCTTGTCTCATTAGACATGCAAC\n',
'TACGGGAGGGGTGACAGGAAGACAAGACACTATTTCCTCAGGCCCAGTTT\n',
'GGTGTGGGGAGAAAGCCTCCTGATCCTGAAAGCAAGAATTTGACCAGAGC\n',
'AGAAGTAATCAGTATGCAGATTGATTCTGTGGTATGTTAATGTTTATGCA\n',
'TAGATTATGAGGACCAGGTGAAAAGTGGGCCAGGGGAGCCAGATGTGTGT\n',
'GTGAGTCATGGGTGGCTGAGATGAGGACAGGAGGGAAACTGGTTTGGAGG\n',
'GTGCTGGCGATGGGGTGGGGGTGCCAGGAGGAAGGGAGGCTAGTTGTTTG\n',
'AATGTCTGCATGAAAAAGCGGACGACAGCGGGGTCTGGGTGAATTCGGGC\n',
'AACCATTTGGACCGTGGAGAAAACTGCCTGCGTGCGGCTGAGGACCTGCA\n',
'CTATTAATTTGTTTTTTAGCTAAGGCAAAGATAAATATAAAAACtgatac\n',
'tccacccagttaccagaaaacatttaggtatgtgtgagacaacttgggta\n',
'tgtgaacctaccttttcaatgtaaattcagtgaaatctaagtgcagatcc\n',
'catatttccaataaaaaggtaacatccaaactcagatgtcctatgagtat\n',
'aaaatacacaaagatcttctggacttagtatgaaaagggatttttttttt\n',
'gtcaggtacctcactagttatttttaaaataggattgcatgttgaaatga\n',
'taatcttttggatatattgggttaaataaatttattattaaagttaattt\n',
'cacttaaaaatgtttaatgtagctactagaaattttaaaattaagcatgt\n',
'tgctcaccttatgtttctattggacggctctCTCTAGATACAAAGGCTGC\n',
'CAAGAGGGACCTCACTCTAGCTTCAGGGAGAAGAGAGGAATTAGCAAGGC\n',
'CAAGCAGAGGCTCCTGAGGGCAGGGCCAAGGGCGGCTTGGTGGGGTGGGG\n',
'ATGGGATGCACAGAGATAACTCCAACCCTTAAGAAGGTGTTTCCTAGAGC\n',
'AGGCTGTGACCTGTCAGTTTATATACTGAGGCTTAGGAGCCTCTTGGATG\n',
'CCCCCAGATCTGCACCCCTGAATTGCCCTGTGCCCCTGCCGTCTTTGTTC\n',
'CTGTGCTGGCATAGTGGTCTCACCTCCGGCAGtatcaccaccactgggca\n',
'caagcttctccagcacagcaactgtgtcttatttctccttgtactcccag\n',
'tgttcacaccatgctgcactcacagaagactcttcgttgatattttgtgg\n',
'acagagagaatGCCTGTGAGAGTGGGCTGAAGTGTGCGTTGGGCTCCAGA\n',
'GACCTTAAGGAGGGGAGACCAGGTCCTGAGTAAAGTTGAAGGGGAGGGGC\n',
'TGAGTCCTGCTAGCCAGGAGTCTCATCCCCTGGGGAAGTTCCAGGGACCC\n',
'CTCAGAAGTGCAAGGGGACGGTGTTAGTGTTAGTCCAGTAACACAGCCCA\n',
'GAGCCTGCcttccacgtgggtttgacaggagcctcctaactgctcttctg\n',
'cttccatttttgccccttcagtctattctcaacagggaagccagaggcat\n',
'ccttaaccatgtcagatcatgtggctcctcagctcaaagccTCATCTCAG\n',
'AGGAAAGCTCTGGTCCCTTAGAAATGGCCCAAGTGGTGACAGACAGACTC\n',
'TAAGGtgagcagactgttgctagatatctgggctcggaggactcgccact\n',
'gctcaaaggcagtgaggattttcgcactagaagctggaggacagggatcc\n',
'ttgttaggtaggagcagaaagcttagaaaagtggtctcctgcagttacgt\n',
'ggcaaacacatcatgtaagtgataaattgggtatgcagttgaggagattt\n',
'ccaagtaaaatgttgaggatgctgcctggtttcttcttactgcttataat\n',
'atagtgtgagagaagagagataaattgagaaagagactggtttttaaact\n',
'gttaaaattgaatcaggacttgatgattttgaaaattgtcagtctcccca\n',
'catggaaaaagatgctgaaattaacaaatggcttctgagcatgtggcata\n',
'gggtgtaactgtacagtcttttgtgattatgcataaagatcaaaggatgg\n',
'gagtagcaatgagtcacacagaggtctgttgcaagagattacaagggtgt\n',
'accatgcagaacctctccaccaaaccttagggcccttgggaagcttcagt\n',
'gagttaccctgggggccatcttggcaggagctgaaggtagaaaggtagag\n',
'tttatctctaaaagattcatgggtatggctcttgacaaatcgactatgag\n',
'ccccaccgaaacccacagaggacaggcaaagggtttgggaaagctgtttc\n',
'acccacagtgctggcagattggtctgtaggggacagagtgcaaaatgaaa\n',
'gaagactgtcagagaccccaaactctgctgtcaagaagaaggctgataaa\n',
'actacttggctgcaaacacgtggatctttcgtgagaaaagaaggatgacc\n',
'cagaggcagaagcccagaaggcagagccaagagacatggaatcttcccac\n',
'atcttaaaacctgtttagggaacaccagcatctgtccagctggatttcag\n',
'aaccaccattccttcatccttcccctgctgcctctttctgaacagcaatg\n',
'tctcaagctttacccaccattgtgtgttgcatatgtagggggcagatagc\n',
'ttgtatctttagttttccagatcagaggaacatccaaagaaatctgttct\n',
'acacctaaacccgatttagatgagattcgggactgtgagcatgaagggat\n',
'ctcaagaggggtgaatgtgttttgcatgcacaagggacaggagtcttggg\n',
'gacagaggacaggctgtggtggcagatactaaggtgacccccacaacccc\n',
'cacctctgccattcacacccttgaataatccccttctctggttgtaagca\n',
'gaacctgtggcttgcttatgaaggaggcggtatatatgtgattcatgtac\n',
'tgatcatattgtataagatcactggctggatgcagtggctcgtgcctgta\n',
'atcccaacactttgggaggctgaggcgggtggatcacctgaggtcaggag\n',
'ttcgagaccaggctggccaacatggcaaaaccccgcctctactaaaaata\n',
'caaaaattagccaggcatagtggtgcacgcctgtaatcacagctactcaa\n',
'gaggctgaagcaggagaattgcttgaactcaggaggtggaggtggcagtg\n',
'agccaagatcgtgccactgcactccagcctcagtgacagagcgagactct\n',
'gtctcaaaaaataaataaataaaatgttaagatcataacctgtctttctg\n',
'gggactctctcttgacgcctttgaagaagcaggctgccatgttgcaagct\n',
'gcctcatggaggggatcagctgcgaggagctaagagccccctccagtcga\n',
'tgctcaccaggaagctgaggtcttgtgtccagcaccctgcatagaactga\n',
'atgctgccatgtgagcttggaagcagagccatccacacagctgagcccta\n',
'gatgagaacccagtgctggctgacaccctgatggcaccttacagaggacc\n',
'agttaggctgtgccaactcctgacctgcagaagctggggaacactgggtc\n',
'gtatttgcagctgctggatttgtgggaatttgtcacacagcaatTGGGAG\n',
'TCACACAgcctgtgacgccccaacaatccacacctcctgcatctccctgc\n',
'cttcacttcctagcacactgccctgactccctctgccgcagccacgctgg\n',
'ccctctgctgttcttcgaagccaccagggctgcattggctcccagccttt\n',
'gctctcactgctttctcctcctagagagcccttcctgcatgtatatgttt\n',
'gactcactcccttgcctccttcagacttgtacttaaaaatctcagtaagc\n',
'atttccctggctacccttttaaaaattgcaacccacttccatccccatcc\n',
'ccaacatgccatatttcctttcttctTCttccttcttccttttttttttt\n',
'ttttttttgacacaggttctctgtcacccagcctggagtgcagtgacatg\n',
'atctcggctcactgcaacctctgcctcccCAGGCAagaaaaggggaggat\n',
'gccaataaaggatgcattgatttgtatttactacagtggacatcaagggc\n',
'acattcttgctgtggccatcaagagactgtataaattctatgacttgtag\n',
'ttgtcccacttaagaaacaaagaagctgTGCATTTCTTTACTGGTCTAGA\n',
'GCTGCTCTAGGGCATTTTCTCTACAGCAATTCTAGGTTTCCCCACCTTGT\n',
'GAGTTTAGCTTTTTCTATATTCAAAGAAAAGTCCTCAGCCAGAGATTCTC\n',
'AGGAGCTTATAGAACAATCCAAACTCTTGGGAATATTAAGTGGAGAGGGG\n',
'TACGTGCAAGACACCAACAGCACTAGAAACAGTCCACATCTTTCCATGCG\n',
'TGGAGGAGTTTATGCTCTATGTGAGTTCACTCCATCATTAATTCTTCAAA\n',
'CACAAGAGTGTTAAAGGAACAAGAGTTAATGGGTCCTGTCATTACACTTG\n',
'TTCCCAGGATGACATTCTTCATCTTCCTCTTCTACAACCTGTTCTATATT\n',
'CCCCTCATGTTTATCCAGTGCTTCTGCTAGTCTAGTTCACTTCCAAAGAC\n',
'CCATGATTACCATGGCCCTGTCAGGCTGTAATTGCTGCAATTTCCAATTT\n',
'ACAATTGTCATCATCTATGGTTGATAAAGGTATAGCAATATTTctatttc\n',
'ctcatgataatgaaggtcaattacaactgccagtataataacttatttct\n',
'ttgtctgccaacctacatacacaaggaagccaaaatgacagggagctact\n',
'aaaactttattcttattggaatgcttactatgtacccagaagaagcattc\n',
'tccctactccagcagagcttaatgctgtaggtccaggaagctcaaattct\n',
'ccaagggagttttagtgagaggagccactctcaccctctgcccttggttt\n',
'acaaacctgtatattctaggacccaatatcttacaatgtccattggttca\n',
'aagtataacatgttaaagcacagagccccaactctgaaaagtaccatccc\n',
'taaattggcatttagttgcacctttatatccacctttaaaagaaatatct\n',
'tttaatgttctatcagactgatagattctgtttaatatagtatattatag\n',
'caccagtggatcatttggttgtatgcatattattgtaccttctctgctac\n',
'aaaatatattcctttgtcctaaggtgtgttacaaagaacattaggcattc\n',
'tatgcatctttggatagtttaatggccaagacattgatggcaggagagtc\n',
'aaagccacaggtggaaaacacatttatcccagtaagaacaaattgctatt\n',
'cttccactgtagagagggtaaacaatgtgccattacgttgccaattgaat\n',
'gcctcaatcatgtcaagggctgaacatctatgactgtttctgaaaggtca\n',
'aacattcaacagaggctgtagctagaactgccttaatgataagagatcat\n',
'gctgaattacccatgcaaaaccttaatacttgacacttatcactacttta\n',
'ttcaagagcctattgtgcaagcaTAAGTGGCTGAGTCAGGTTCTCAACTC\n',
'TGCTCATTAATACTATGCTTGGAGTATACAGTAAGATAAGAAACATAAAT\n',
'AAGAAGTGTACATTTGTTTcttcctgttttcttctggctattggatcaat\n',
'tacatcccatcttaagctgacccctgtgtaattaatcaatatccgtttta\n',
'agcagcaatccatagttgtgcagaaattagaaaactgacccacacagaaa\n',
'aactAATTGTGAGAACCAATATTATACTAAATTCATTTGACAATTCTCAG\n',
'CAAAGTGCTGGGTTGATCTCTATTTACGCTTTTCTTAAACACACAAAATA\n',
'CAAAAGTTAACCCATATGGAATGCAATGGAGGAAATCAATGACATATCAG\n',
'ATCTAGAAACTAATCAATTAGCAATCAGGAAGGAGTTGTGGTAGGAAGTC\n',
'TGTGCTGTTGAATGTACACTAATCAATGATTCCTTAAATTATTCACAATA\n',
'AAAAAAAAGATTAGAATAGTTTTTTTAAAAAAAAAGCCCAGAAACTAATC\n',
'TAAGTTTTGTCTGGTAATAAAGGTATATTTTCAAAAGAGAGGTAAATAGA\n',
'TCCACATACTGTGGAGGGAATAAAATACTTTTTGAAAAACAAACAACAAG\n',
'TTGGATTTTTAGACACATAGAAATTGAATATGTACATTTATAAATATTTT\n',
'TGGATTGAACTATTTCAAAATTATACCATAAAATAACTTGTAAAAATGTA\n',
'GGCAAAATGTATATAATTATGGCATGAGGTATGCAACTTTAGGCAAGGAA\n',
'GCAAAAGCAGAAACCATGAAAAAAGTCTAAATTTTACCATATTGAATTTA\n',
'AATTTTCAAAAACAAAAATAAAGACAAAGTGGGAAAAATATGTATGCTTC\n',
'ATGTGTGACAAGCCACTGATACCTATTAAATATGAAGAATATTATAAATC\n',
'ATATCAATAACCACAACATTCAAGCTGTCAGTTTGAATAGACaatgtaaa\n',
'tgacaaaactacatactcaacaagataacagcaaaccagcttcgacagca\n',
'cgttaaaggggtcatacaacataatcgagtagaatttatctctgagatgc\n',
'aagaatggttcaaaatatggaaaccaataaatgtgatatgccacactaac\n',
'agaataaaaaataaaaatcatattatcatctcaatagatgcagaaaaagc\n',
'attaacaaaagtaaacattctttcataataagacatcagataaaacaaat\n',
'taggaatagaaggaatgtaccgcaacacaataaaggccatatataacaag\n',
'cccacagctaacatcataatagtaaaatcatcacactggtaaaaaaaatg\n',
'aaagcttttcctctaaggtcagaaataatataaaggttcccactcttgct\n',
'atttctattccatatcgtactaaaagtcctagccaggacaattagacaaa\n',
'ataaaaataaaaacacccaaattggaaagatagaagcaaacttttctgtt\n',
'tacagataacataatcttatatgtagaaaccccttaaaacttcagcaaaa\n',
'aaaaaaaaaaaactacagagctagtaaattcagtgaagttgcagaataca\n',
'aaatcaacatacaaaaatcagtagtgtctctatacactaataaggactta\n',
'acagagaaagaagttaagaaaacaataccactaacaatagaatccaaaaa\n',
'ataaaatacttaggaataaattttaccaaacatctgtacactaaaaacta\n',
'taaaacattgaaaaaagaagttgaataagacacatataaatagaaagcta\n',
'tctcatgttaatagattagaaaaagtaatattgttaagatgtcctcacta\n',
'cttaaagcaatttatagatctaatgcatttattgcaatctcttcaaaatc\n',
'ccaaaggtatttttgacagaaataaaaaaaaaattctaaaatatgcatga\n',
'aaccacaaaagactgtgaatagctaaagcaatcttgagcaagatgaacaa\n',
'cactggaagcatcacactaccttatttcaaaatctactacaaagctatag\n',
'tgatcaaagcaacatgatactgtcataaaaacacacagataaacctatgg\n',
'aatggaataaagagcacagaaataagtccacacatttacattcaattgat\n',
'tttcaacaacaatgtcaagaagacaatggggaaaagacaatctcttcaat\n',
'aaatgatgctggaaaaactatatatccacatgcagaagaatgcagttgaa\n',
'tcctgatttcataccatatgcaaaattcaactggaaatggattaaataca\n',
'aatttaaaacatgaaatggtataactattagaacaaaacatagaaaatat\n',
'tcttcctgacattggtttgggccatcatttttctgatatgactctaaaag\n',
'cacaggcaaaaaaagaaaaaatagacaaatgagactatgccaaattaaaa\n',
'aatttctaacaacaaaagaaacgatcaatagagtgaaaaagataacctct\n',
'tgaatgggagaaatatttgcaaactactcatccaaccggggattgatatc\n',
'cagaatatacaagtaacacaaatatgtcaaaagtaaaataaataaataaa\n',
'taaataaataaataaattaaataaattatttaaaaatcggcagaggacag\n',
'gaatagacatttctcaggagacaacatacaaagggccacagatacatcaa\n',
'aaaatgctcaacatcactatttgtcagggaagtactaattaaaaccaaaa\n',
'tgagatgtcccctcaaacctgttagaatggctattatcaaaaagatgaaa\n',
'gatagcaactatcagagaggatgatagaaaagggaacccttgcatcatgt\n',
'acaaattaaaaatagaactatcacatgatccaagaatcctacttctgggt\n',
'atatagccaaaggaattgaaatcaatatgtcaaagggatatctgcactcc\n',
'tatgttattgcagcatgttcacaatggccaagatatagaatcaacctaac\n',
'tgttcatagacagatgaatggataaatgaaatgtgatatggaaaattatt\n',
'cagccttaaaaacagtaggaaattctgtcatttgagacaacgtggatgaa\n',
'cctagaggacattaagctaagtgaaataagctagacacagaaagacaaat\n',
'attgcatgatctcacttagaatctaaaaaatctgaactcatagaagcaga\n',
'gaatagtatgatggttactagggttatctggcagggagaggatgaggaaa\n',
'tgggacattgttaataaaaggaaaaaaattcaattagtaggattacattc\n',
'aggggacccaatatacgacatgttgactgtaattaataatgtattgtatg\n',
'cttgaaaattgctaatacagtatattgtaaatgttaatatgaggtaatat\n',
'atgtgttaattaacttgatttattcattcaacaacatacacatatattaa\n',
'aacatcacactgtattccacaaatatatataatttttgtcaattaaaaaa\n',
'taaTTTTTAAAAATGAGAAACAAAAAAGCTGACATTTTCAGATTAAAAAA\n',
'ATTATACAGAAGAATTAATTCATTAAAGTAAAAACAAATGTGGGAAAATG\n',
'GTTTTTAAATATAATTTAAACCAAATTTAAAATAAGcatataaagactat\n',
'ggacaaaacaagaaatccaaataaaaaataaacatatgaagaatattcaa\n',
'actcactttttatcaaagaaatgtaaattttaaaataTAGCATTGCTATT\n',
'GTGTTTTCATAAATAATAATATATCATGGATGAGCCTGTGAGGAAACAGA\n',
'CACTCATACTCTGCAAAGCAATGACTAAgataattatgtcagatcatgaa\n',
'ttacgttaattagcttgatggtggtcactgtttcacgataaatatacata\n',
'tgtatcaaaacatcacattacacaccataaagatatataacttgttatCA\n',
'AAAAGAAATATAGCAGTTAAAATTTAAAATTTTTAAAAAACGTCTTTTTG\n',
'AGGTTCGTACCTCACTTAAGTCACACTGTTCAAAATATTCATGCACTCAT\n',
'TTCTCTCATTCATGTGTTAATGTACAGGGTACGGGCCACTATAAATTCCT\n',
'TCAGCAACTGGAAAGGAAACTTTATGTACTGAGTGCTCAGAGTTGTATTA\n',
'ACTTTTTTTTTTTTTtgagcagcagcaagatttattgtgaagagtgaaag\n',
'aacaaagcttccacagtgtggaaggggacccgagcggtttgccCAGTTGT\n',
'ATTAACTTCTAATTCAACACTTTAAGATTCTTAGCATTATTGCAGACAAC\n',
'ATcagcttcacaagtgtgtgtcctgtgcagttgaacaagatcccacactt\n',
'aaaaggatcctacactttttttaatgctctgctgtttctgccttgaaatt\n',
'cttaacaatttttttaaccaaagtcctcacaaattcagtttacattagcc\n',
'ctgcaatcatgtagacatcctgATTCCAGACAATGTGTCTGGAGGCAGGG\n',
'TTTACAGGACTTCAAGAACCTTACCTTCTCAACTTTCATCTGCATCTTTA\n',
'CTCCCAACTATATATGAAGATGATGAAGATAGATATGGATGGTGCTTCTA\n',
'CCATACCCTCTTCCTCTGCCAAACTTCCTTGATCTAGGATAAggtcagta\n',
'aacttcttccgtaaaaggccaaaagtaaatattataggctctacaggccc\n',
'tagagtgtctgtcataactactcaactcttattgtagcataaaaactgtc\n',
'aacagacaatacagaaacaaatgagtgtgactgggttccagtgaaacttt\n',
'atttacaaaagatttgtcccatgagtcaaatttaccacctccAGATCTAG\n',
'AGAAACAGTTTTGAGCCCTTTTATTTTGCTCAACAGTTAAGCATGGCTCC\n',
'ATGTCCCTTATATTTAGTCAGAACTCGGTATGTTTTAAGGAAAGAATGGT\n',
'TACACGAAGACATACATTCATTCATTTATACAACACATTTTCAGTGTTGA\n',
'ATGATAAATTTTGGAATAGTTAACAGATGATAAAAGTGTTGTTTTCAGTC\n',
'ATCCCTATCCAATGAAGTAAAAAAAAAAGTGTTGAATGGGAAGAAATCAA\n',
'GAATAGTTATACGAATATCACCATTGCATTAAAGCTCTCTTCCTTGTTTC\n',
'TAAAAGAATATCTTGACACACATTAAGCTCACTGACCCCCACACCATGAA\n',
'TGAGGGCATCTTCAACAATGGTGGATGACGTCTTAGTTTCCCTCAACTCA\n',
'GTTAATCTAAGTAAGCTCATGGTATCACTTTCCTGTCCTAGAGGGAACAT\n',
'ATTTCCTGCATTTTTCTTTTTTTCCTTACTTTCCATCACCAAGTAACTCT\n',
'TCTGATATTTTTTCTCTTGAGAAAATTAATATGACTCATAGATCTGGTTC\n',
'CCAAGAGAAATCAATGGAGGCCTGGTTACAAGGATCTAAGAAGCATCAAT\n',
'GGGTCACTAACATCTAGTGGTACTAATTAACTCTGTTAATCATTGGGAAG\n',
'AAAATGTATATATACTTTTGTCTTGGAGCTGATTCTACTAGAAAGCAGAA\n',
'ATCAAAATGATCAGTTTCCCAGTGTCACTACTGCACACCCTGGAACAGAA\n',
'CAGGTAGGTCAGAAAAACGCTCCCAAAGTTTAGCAATGTCAAGGCAATCT\n',
'CTCTCTTCTTACATTTCCCTTCAACCTTCTATCTCCTCCACTTTTCTGTT\n',
'TTCCTCCTATCTCCAATTATTTCAATCCTCAGAGCATTATTCTTACAATC\n',
'TTAATCACTAAATTATATTACACCCGTTAAAGGAGAGATTTCTAAATGCA\n',
'TTGACATTTGTACTGTCTCTCTTTGGAGAATTAGTATTATAAGGATCTGT\n',
'TATCTCTTGTCACCTTCCTTATGTCATATGATATGTCACATTTCCCACTG\n',
'CGGAGACCAAACATGTTCACATCGTGTGCGTTCCATTTTCCTAATGGAAA\n',
'GTGGGGGGAAGTGATTTTCTGTCCTCATATAGAGAATGCTGGGGCCATTC\n',
'CCTCTGTATGCCATATTTGATAAAGCATTTGATAATCTTAGTCAATGCCT\n',
'GGGCCAAGAATTAAAGGGGTAATTATCAGAATGAAAATGGTTTAATGAAA\n',
'CTGTGTCTATCAGTTCTGAAAAGGGCCTCTATCACAATGAACTAAGGTAG\n',
'TTATGAATAGAGCTAAaacttaggcaacaccatcctggacataggaacgg\n',
'gcaaagatttcatgacaaagacacggaaaccaatcacaacaaaagcaaaa\n',
'attgagaagtggaatctaataaaacaatagcttctgcacagcaaaagaag\n',
'ctaccaacaaagtaaacagacaacctacagaatgggagaaaatatttgcc\n',
'aactgtaagtctgacaaaaatctaatatctggcagctataaggaacttaa\n',
'atttacaagacaaaaacaaccccattaaaaagtgggcaaagaacatgaat\n',
'agacactctcaaaagaagatatacatatggttaacaagcatatgaaaaaa\n',
'aagctcaatatactgagcattagagaaatgcaaatcaaaaccatattgag\n',
'atatcatctcataccaggcagaatggctattattaaaaagtcaaaaataa\n',
'cagatatcggtgaggttacagagaaaagggaacacttatacactgttggt\n',
'gggactgtaaattatttcaaccattgtggaaagcagtatgggatggcgat\n',
'tcctcaaaaagccaaaaacagaactatcattcaacccagcaattccatta\n',
'ctgggtatatacccagaagaatataaatcgttctaccataaagacgcatg\n',
'catgagaatgttcattgcagcactactcacaatagcagagacatggaatc\n',
'aacttaaatgcccatcagtaacagactggataaagaaagtgtggtacaga\n',
'tacaccgtggattactatgcagccataaaaaagaacaagatcatgtcttt\n',
'gacaggaacatggatggagctggaggctactatccttagcaagctaaggc\n',
'aggaacagaaatccaaataccgcatgttctcacttatgagcgtgagataa\n',
'atgatgagaacttgtaaacacaaagaaggaaacaacaggcagtggggtct\n',
'acttgaggacgacgggaagagggagaggagcagaaaagataactactgac\n',
'taccgggcgctacctgggggatgaaacaatctgtacaacgaacccccagg\n',
'acatgagtttacctatgtaacaaaccttcacgtgtacccccgaacctaaa\n',
'ataaaagtcaaaaagaaaAAGAAAAAAAGAAAAATCCATGCATATGATAC\n',
'ATCAGTTAACAAGGCACTGGTGAAATTAATTTTAAGTATTATTGTCTCTT\n',
'TGTGTTTTTGGTCTCAGAAAAGTTACGATTTCCCTTAGTTCCTTAGGGCA\n',
'GAGAGAATCTTCAATCACTGAAGTCAGGAGACACACATTCTATCTGATTT\n',
'TCTACATTATCTGTTTGAAAAGGTTACCCACTTATTAGTGTTAAAGCCAA\n',
'GATATCCAGCAAGGATAGCAACCAACTCTTAAGGTACTCTCCCTTAGGAG\n',
'GATTCCTGATTCTTTAATGTTTTCTaaaaaagcaaaacaaacaaacaaac\n',
'aaaacaaaacaCTAAATGTTTTCTCTTTCAACTTATTTGAATACACTCTT\n',
'TTCTCACTGCTCTGAGCATGAATTCAATATTTCAGGGCAAACTAACTGAA\n',
'TGTTAGAACCAACTCCTGATAAGTCTTGAACAAAAGATAGGATCCTCTAT\n',
"""
for s in "\n," :
human = human.replace(human, "")
print(human)
lower=convertOpposite(list2)
human="""
"""
list2=[ 'taaccctaaccctaaccctaaccctaaccctaaccctaaccctaacccta\n',
'accctaaccctaaccctaaccctaaccctaaccctaaccctaaccctaac\n',
'cctaacccaaccctaaccctaaccctaaccctaaccctaaccctaacccc\n',
'taaccctaaccctaaccctaaccctaacctaaccctaaccctaaccctaa\n',
'ccctaaccctaaccctaaccctaaccctaacccctaaccctaaccctaaa\n',
'ccctaaaccctaaccctaaccctaaccctaaccctaaccccaaccccaac\n',
'cccaaccccaaccccaaccccaaccctaacccctaaccctaaccctaacc\n',
'ctaccctaaccctaaccctaaccctaaccctaaccctaacccctaacccc\n',
'taaccctaaccctaaccctaaccctaaccctaaccctaacccctaaccct\n',
'aaccctaaccctaaccctcgcggtaccctcagccggcccgcccgcccggg\n',
'tctgacctgaggagaactgtgctccgccttcagagtaccaccgaaatctg\n',
'tgcagaggacaacgcagctccgccctcgcggtgctctccgggtctgtgct\n',
'gaggagaacgcaactccgccgttgcaaaggcgcgccgcgccggcgcaggc\n',
'gcagagaggcgcgccgcgccggcgcaggcgcagagaggcgcgccgcgccg\n',
'gcgcaggcgcagagaggcgcgccgcgccggcgcaggcgcagagaggcgcg\n',
'ccgcgccggcgcaggcgcagagaggcgcgccgcgccggcgcaggcgcaga\n',
'cacatgctagcgcgtcggggtggaggcgtggcgcaggcgcagagaggcgc\n',
'gccgcgccggcgcaggcgcagagacacatgctaccgcgtccaggggtgga\n',
'ggcgtggcgcaggcgcagagaggcgcaccgcgccggcgcaggcgcagaga\n',
'cacatgctagcgcgtccaggggtggaggcgtggcgcaggcgcagagacgc\n',
'aagcctacgggcgggggttgggggggcgtgtgttgcaggagcaaagtcgc\n',
'acggcgccgggctggggcggggggagggtggcgccgtgcacgcgcagaaa\n',
'ctcacgtcacggtggcgcggcgcagagacgggtagaacctcagtaatccg\n',
'aaaagccgggatcgaccgccccttgcttgcagccgggcactacaggaccc\n',
'gcttgctcacggtgctgtgccagggcgccccctgctggcgactagggcaa\n',
'ctgcagggctctcttgcttagagtggtggccagcgccccctgctggcgcc\n',
'ggggcactgcagggccctcttgcttactgtatagtggtggcacgccgcct\n',
'gctggcagctagggacattgcagggtcctcttgctcaaggtgtagtggca\n',
'gcacgcccacctgctggcagctggggacactgccgggccctcttgctCCA\n',
'ACAGTACTGGCGGATTATAGGGAAACACCCGGAGCATATGCTGTTTGGTC\n',
'TCAGtagactcctaaatatgggattcctgggtttaaaagtaaaaaataaa\n',
'tatgtttaatttgtgaactgattaccatcagaattgtactgttctgtatc\n',
'ccaccagcaatgtctaggaatgcctgtttctccacaaagtgtttactttt\n',
'ggatttttgccagtctaacaggtgaAGccctggagattcttattagtgat\n',
'ttgggctggggcctggccatgtgtatttttttaaatttccactgatgatt\n',
'ttgctgcatggccggtgttgagaatgactgCGCAAATTTGCCGGATTTCC\n',
'TTTGCTGTTCCTGCATGTAGTTTAAACGAGATTGCCAGCACCGGGTATCA\n',
'TTCACCATTTTTCTTTTCGTTAACTTGCCGTCAGCCTTTTCTTTGACCTC\n',
'TTCTTTCTGTTCATGTGTATTTGCTGTCTCTTAGCCCAGACTTCCCGTGT\n',
'CCTTTCCACCGGGCCTTTGAGAGGTCACAGGGTCTTGATGCTGTGGTCTT\n',
'CATCTGCAGGTGTCTGACTTCCAGCAACTGCTGGCCTGTGCCAGGGTGCA\n',
'AGCTGAGCACTGGAGTGGAGTTTTCCTGTGGAGAGGAGCCATGCCTAGAG\n',
'TGGGATGGGCCATTGTTCATCTTCTGGCCCCTGTTGTCTGCATGTAACTT\n',
'AATACCACAACCAGGCATAGGGGAAAGATTGGAGGAAAGATGAGTGAGAG\n',
'CATCAACTTCTCTCACAACCTAGGCCAGTAAGTAGTGCTTGTGCTCATCT\n',
'CCTTGGCTGTGATACGTGGCCGGCCCTCGCTCCAGCAGCTGGACCCCTAC\n',
'CTGCCGTCTGCTGCCATCGGAGCCCAAAGCCGGGCTGTGACTGCTCAGAC\n',
'CAGCCGGCTGGAGGGAGGGGCTCAGCAGGTCTGGCTTTGGCCCTGGGAGA\n',
'GCAGGTGGAAGATCAGGCAGGCCATCGCTGCCACAGAACCCAGTGGATTG\n',
'GCCTAGGTGGGATCTCTGAGCTCAACAAGCCCTCTCTGGGTGGTAGGTGC\n',
'AGAGACGGGAGGGGCAGAGCCGCAGGCACAGCCAAGAGGGCTGAAGAAAT\n',
'GGTAGAACGGAGCAGCTGGTGATGTGTGGGCCCACCGGCCCCAGGCTCCT\n',
'GTCTCCCCCCAGGTGTGTGGTGATGCCAGGCATGCCCTTCCCCAGCATCA\n',
'GGTCTCCAGAGCTGCAGAAGACGACGGCCGACTTGGATCACACTCTTGTG\n',
'AGTGTCCCCAGTGTTGCAGAGGTGAGAGGAGAGTAGACAGTGAGTGGGAG\n',
'TGGCGTCGCCCCTAGGGCTCTACGGGGCCGGCGTCTCCTGTCTCCTGGAG\n',
'AGGCTTCGATGCCCCTCCACACCCTCTTGATCTTCCCTGTGATGTCATCT\n',
'GGAGCCCTGCTGCTTGCGGTGGCCTATAAAGCCTCCTAGTCTGGCTCCAA\n',
'GGCCTGGCAGAGTCTTTCCCAGGGAAAGCTACAAGCAGCAAACAGTCTGC\n',
'ATGGGTCATCCCCTTCACTCCCAGCTCAGAGCCCAGGCCAGGGGCCCCCA\n',
'AGAAAGGCTCTGGTGGAGAACCTGTGCATGAAGGCTGTCAACCAGTCCAT\n',
'AGGCAAGCCTGGCTGCCTCCAGCTGGGTCGACAGACAGGGGCTGGAGAAG\n',
'GGGAGAAGAGGAAAGTGAGGTTGCCTGCCCTGTCTCCTACCTGAGGCTGA\n',
'GGAAGGAGAAGGGGATGCACTGTTGGGGAGGCAGCTGTAACTCAAAGCCT\n',
'TAGCCTCTGTTCCCACGAAGGCAGGGCCATCAGGCACCAAAGGGATTCTG\n',
'CCAGCATAGTGCTCCTGGACCAGTGATACACCCGGCACCCTGTCCTGGAC\n',
'ACGCTGTTGGCCTGGATCTGAGCCCTGGTGGAGGTCAAAGCCACCTTTGG\n',
'TTCTGCCATTGCTGCTGTGTGGAAGTTCACTCCTGCCTTTTCCTTTCCCT\n',
'AGAGCCTCCACCACCCCGAGATCACATTTCTCACTGCCTTTTGTCTGCCC\n',
'AGTTTCACCAGAAGTAGGCCTCTTCCTGACAGGCAGCTGCACCACTGCCT\n',
'GGCGCTGTGCCCTTCCTTTGCTCTGCCCGCTGGAGACGGTGTTTGTCATG\n',
'GGCCTGGTCTGCAGGGATCCTGCTACAAAGGTGAAACCCAGGAGAGTGTG\n',
'GAGTCCAGAGTGTTGCCAGGACCCAGGCACAGGCATTAGTGCCCGTTGGA\n',
'GAAAACAGGGGAATCCCGAAGAAATGGTGGGTCCTGGCCATCCGTGAGAT\n',
'CTTCCCAGGGCAGCTCCCCTCTGTGGAATCCAATCTGTCTTCCATCCTGC\n',
'GTGGCCGAGGGCCAGGCTTCTCACTGGGCCTCTGCAGGAGGCTGCCATTT\n',
'GTCCTGCCCACCTTCTTAGAAGCGAGACGGAGCAGACCCATCTGCTACTG\n',
'CCCTTTCTATAATAACTAAAGTTAGCTGCCCTGGACTATTCACCCCCTAG\n',
'TCTCAATTTAAGAAGATCCCCATGGCCACAGGGCCCCTGCCTGGGGGCTT\n',
'GTCACCTCCCCCACCTTCTTCCTGAGTCATTCCTGCAGCCTTGCTCCCTA\n',
'ACCTGCCCCACAGCCTTGCCTGGATTTCTATCTCCCTGGCTTGGTGCCAG\n',
'TTCCTCCAAGTCGATGGCACCTCCCTCCCTCTCAACCACTTGAGCAAACT\n',
'CCAAGACATCTTCTACCCCAACACCAGCAATTGTGCCAAGGGCCATTAGG\n',
'CTCTCAGCATGACTATTTTTAGAGACCCCGTGTCTGTCACTGAAACCTTT\n',
'TTTGTGGGAGACTATTCCTCCCATCTGCAACAGCTGCCCCTGCTGACTGC\n',
'CCTTCTCTCCTCCCTCTCATCCCAGAGAAACAGGTCAGCTGGGAGCTTCT\n',
'GCCCCCACTGCCTAGGGACCAACAGGGGCAGGAGGCAGTCACTGACCCCG\n',
'AGACGTTTGCATCCTGCACAGCTAGAGATCCTTTATTAAAAGCACACTGT\n',
'TGGTTTCTGCTCAGTTCTTTATTGATTGGTGTGCCGTTTTCTCTGGAAGC\n',
'CTCTTAAGAACACAGTGGCGCAGGCTGGGTGGAGCCGTCCCCCCATGGAG\n',
'CACAGGCAGACAGAAGTCCCCGCCCCAGCTGTGTGGCCTCAAGCCAGCCT\n',
'TCCGCTCCTTGAAGCTGGTCTCCACACAGTGCTGGTTCCGTCACCCCCTC\n',
'CCAAGGAAGTAGGTCTGAGCAGCTTGTCCTGGCTGTGTCCATGTCAGAGC\n',
'AACGGCCCAAGTCTGGGTCTGGGGGGGAAGGTGTCATGGAGCCCCCTACG\n',
'ATTCCCAGTCGTCCTCGTCCTCCTCTGCCTGTGGCTGCTGCGGTGGCGGC\n',
'AGAGGAGGGATGGAGTCTGACACGCGGGCAAAGGCTCCTCCGGGCCCCTC\n',
'ACCAGCCCCAGGTCCTTTCCCAGAGATGCCTGGAGGGAAAAGGCTGAGTG\n',
'AGGGTGGTTGGTGGGAAACCCTGGTTCCCCCAGCCCCCGGAGACTTAAAT\n',
'ACAGGAAGAAAAAGGCAGGACAGAATTACAAGGTGCTGGCCCAGGGCGGG\n',
'CAGCGGCCCTGCCTCCTACCCTTGCGCCTCATGACCAGCTTGTTGAAGAG\n',
'ATCCGACATCAAGTGCCCACCTTGGCTCGTGGCTCTCACTGCAACGGGAA\n',
'AGCCACAGACTGGGGTGAAGAGTTCAGTCACATGCGACCGGTGACTCCCT\n',
'GTCCCCACCCCCATGACACTCCCCAGCCCTCCAAGGCCACTGTGTTTCCC\n',
'AGTTAGCTCAGAGCCTCAGTCGATCCCTGACCCAGCACCGGGCACTGATG\n',
'AGACAGCGGCTGTTTGAGGAGCCACCTCCCAGCCACCTCGGGGCCAGGGC\n',
'CAGGGTGTGCAGCAccactgtacaatggggaaactggcccagagaggtga\n',
'ggcagcttgcctggggtcacagagcaaggcaaaagcagcgctgggtacaa\n',
'gctcaAAACCATAGTGCCCAGGGCACTGCCGCTGCAGGCGCAGGCATCGC\n',
'ATCACACCAGTGTCTGCGTTCACAGCAGGCATCATCAGTAGCCTCCAGAG\n',
'GCCTCAGGTCCAGTCTCTAAAAATATCTCAGGAGGCTGCAGTGGCTGACC\n',
'ATTGCCTTGGACCGCTCTTGGCAGTCGAAGAAGATTCTCCTGTCAGTTTG\n',
'AGCTGGGTGAGCTTAGAGAGGAAAGCTCCACTATGGCTCCCAAACCAGGA\n',
'AGGAGCCATAGCCCAGGCAGGAGGGCTGAGGACCTCTGGTGGCGGCCCAG\n',
'GGCTTCCAGCATGTGCCCTAGGGGAAGCAGGGGCCAGCTGGCAAGAGCAG\n',
'GGGGTGGGCAGAAAGCACCCGGTGGACTCAGGGCTGGAGGGGAGGAGGCG\n',
'ATCTTGCCCAAGGCCCTCCGACTGCAAGCTCCAGGGCCCGCTCACCTtgc\n',
'tcctgctccttctgctgctgcttctccagctttcgctccttcatgctgcG\n',
'CAGCTTGGCCTTGCCGATGCCCCCAGCTTGGCGGATGGACTCTAGCAGAG\n',
'TGGCCAGCCACCGGAGGGGTCAACCACTTCCCTGGGAGCTCCCTGGACTG\n',
'GAGCCGGGAGGTGGGGAACAGGGCAAGGAGGAAAGGCTGCTCAGGCAGGG\n',
'CTGGGGAAGCTTACTGTGTCCAAGAGCCTGCTGGGAGGGAAGTCACCTCC\n',
'CCTCAAACGAGGAGCCCTGCGCTGGGGAGGCCGGACCTTTGGAGACTGTG\n',
'TGTGGGGGCCTGGGCACTGACTTCTGCAACCACCTGAGCGCGGGCATCCT\n',
'GTGTGCAGATACTCCCTGCTTCCTCTCTAGCCCCCACCCTGCAGAGCTGG\n',
'ACCCCTGAGCTAGCCATGCTCTGACAGTCTCAGTTGCACACACGAGCCAG\n',
'CAGAGGGGTTTTGTGCCACTTCTGGATGCTAGGGTTACACTGGGAGACAC\n',
'AGCAGTGAAGCTGAAATGAAAAATGTGTTGCTGTAGTTTGTTATTAGACC\n',
'CCTTCTTTCCATTGGTTTAATTAGGAATGGGGAACCCAGAGCCTCACTTG\n',
'TTCAGGCTCCCTCTGCCCTAGAAGTGAGAAGTCCAGAGCTCTACAGTTTG\n',
'AAAACCACTATTTTATGAACCAAGTAGAACAAGATATTTGAAATGGAAAC\n',
'TATTCAAAAAATTGAGAATTTCTGACCACTTAACAAACCCACAGAAAATC\n',
'CACCCGAGTGCACTGAGCACGCCAGAAATCAGGTGGCCTCAAAGAGCTGC\n',
'TCCCACCTGAAGGAGACGCGCTGCTGCTGCTGTCGTCCTGCCTGGCGCCT\n',
'TGGCCTACAGGGGCCGCGGTTGAGGGTGGGAGTGGGGGTGCACTGGCCAG\n',
'CACCTCAGGAGCtgggggtggtggtgggggcggtgggggtggtgTTAGTA\n',
'CCCCATCTTGTAGGTCTGAAACACAAAGTGTGGGGTGTCTAGGGAAGAAG\n',
'GTGTGTGACCAGGGAGGTCCCCGGCCCAGCTCCCATCCCAGAACCCAGCT\n',
'CACCTACCTTGAGAGGCTCGGCTACCTCAGTGTGGAAGGTGGGCAGTTCT\n',
'GGAATGGTGCCAGGGGCAGAGGGGGCAATGCCGGGGCCCAGGTCGGCAAT\n',
'GTACATGAGGTCGTTGGCAATGCCGGGCAGGTCAGGCAGGTAGGATGGAA\n',
'CATCAATCTCAGGCACCTGGCCCAGGTCTGGCACATAGAAGTAGTTCTCT\n',
'GGGACCTGCAAGATTAGGCAGGGACATGTGAGAGGTGACAGGGACCTGCA\n',
'GGGGCAGCCAACAAGACCTTGTGTGCACCTCCCATGGGTGGAATAAGGGG\n',
'CCCAACAGCCTTGACTGGAGAGGAGCTCTGGCAAGGCCCTGGGCCACTGC\n',
'ACCTGTCTCCACCTCTGTCCCACCCCTCCCACCTGCTGTTCCAGCTGCTC\n',
'TCTCTTGCTGATGGACAAGGGGGCATCAAACAGCTTCTCCTCTGTCTCTG\n',
'CCCCCAGCATCACATGGGTCTTTGTTACAGCACCAGCCAGGGGGTCCAGG\n',
'AAGACATACTTCTTCTACCTACAGAGGCGACATGGGGGTCAGGCAAGCTG\n',
'ACACCCGCTGTCCTGAGCCCATGTTCCTCTCCCACATCATCAGGGGCACA\n',
'GCGTGCACTGTGGGGTCCCAGGCCTCCCGAGCCGAGCCACCCGTCACCCC\n',
'CTGGCTCCTGGCCTATGTGCTGTACCTGTGTCTGATGCCCTGGGTCCCCA\n',
'CTAAGCCAGGCCGGGCCTCCCGCCCACACCCCTCGGCCCTGCCCTCTGGC\n',
'CATACAGGTTCTCGGTGGTGTTGAAGAGCAGCAAGGAGCTGACAGAGCTG\n',
'ATGTTGCTGGGAAGACCCCCAAGTCCCTCTTCTGCATCGTCCTCGGGCTC\n',
'CGGCTTGGTGCTCACGCACACAGGAAAGTCCTTCAGCTTCTCCTGAGAGG\n',
'GCCAGGATGGCCAAGGGATGGTGAATATTTGGTGCTGGGCCTAATCAGCT\n',
'GCCATCCCATCCCAGTCAGCCTCCTCTGGGGGACAGAACCCTATGGTGGC\n',
'CCCGGCTCCTCCCCAGTATCCAGTCCTCCTGGTGTGTGACAGGCTATATG\n',
'CGCGGCCAGCAGACCTGCAGGGCCCGCTCGTCCAGGGGGCGGTGCTTGCT\n',
'CTGGATCCTGTGGCGGGGGCGTCTCTGCAGGCCAGGGTCCTGGGCGCCCG\n',
'TGAAGATGGAGCCATATTCCTGCAGGCGCCCTGGAGCAGGGTACTTGGCA\n',
'CTGGAGAACACCTGTGGACACAGGGACAAGTCTGAGGGGGCCCCAAGAGG\n',
'CTCAGAGGGCTAGGATTGCTTGGCAGGAGAGGGTGGAGTTGGAAGCCTGG\n',
'GCGAGAAGAAAGCTCAAGGTACAGGTGGGCAGCAGGGCAGAGACTGGGCA\n',
'GCCTCAGAGGCACGGGGAAATGGAGGGACTGCCCAGTAGCCTCAGGACAC\n',
'AGGGGTATGGGGACTACCTTGATGGCCTTCTTGCTGCCCTTGATCTTCTC\n',
'AATCTTGGCCTGGGCCAAGGAGACCTTCTCTCCAATGGCCTGCACCTGGC\n',
'TCCGGCTCTGCTCTACCTGCTGGGAGATCCTGCCATGGAGAAGATCACAG\n',
'AGGCTGGGCTGCTCCCCACCCTCTGCACACCTCCTGCTTCTAACAGCAGA\n',
'GCTGCCAGGCCAGGCCCTCAGGCAAGGGCTCTGAAGTCAGGGTCACCTAC\n',
'TTGCCAGGGCCGATCTTGGTGCCATCCAGGGGGCCTCTACAAGGATAATC\n',
'TGACCTGCAGGGTCGAGGAGTTGACGGTGCTGAGTTCCCTGCACTCTCAG\n',
'TAGGGACAGGCCCTATGCTGCCACCTGTACATGCTATCTGAAGGACAGCC\n',
'TCCAGGGCACACAGAGGATGGTATTTACACATGCACACATGGCTACTGAT\n',
'GGGGCAAGCACTTCACAACCCCTCATGATCACGTGCAGCAGACAATGTGG\n',
'CCTCTGCAGAGGGGGAACGGAGACCGGAGGCTGAGACTGGCAAGGCTGGA\n',
'CCTGAGTGTCGTCACCTAAATTCAGACGGGGAACTGCCCCTGCACATACT\n',
'GAACGGCTCACTGAGCAAACCCCGAGTCCCGACCACCGCCTCAGTGTGGT\n',
'CTAGCTcctcacctgcttccatcctccctggtgcggggtgggcccagtga\n',
'tatcagctgcctgctgttccccagatgtgccaagtgcattcttgtgtgct\n',
'tgcatctcatggaacgccatttccccagacatccctgtggctggctccTG\n',
'ATGCCCGAGGCCCAAGTGTCTGATGCTTTAAGGCACATCACCCCACTCAT\n',
'GCTTTTCCATGTTCTTTGGCCGCAGCAAGGCCGCTCTCACTGCAAAGTTA\n',
'ACTCTGATGCGTGTGTAACACAACATCCTCCTCCCAGTCGCCCCTGTAGC\n',
'TCCCCTACCTCCAAGAGCCCAGCCCTTGCCCACAGGGCCACACTCCACGT\n',
'GCAGAGCAGCCTCAGCACTCACCGGGCACGAGCGAGCCTGTGTGGTGCGC\n',
'AGGGATGAGAAGGCAGAGGCGCGACTGGGGTTCATGAGGAAGGGCAGGAG\n',
'GAGGGTGTGGGATGGTGGAGGGGTTTGAGAAGGCAGAGGCGCGACTGGGG\n',
'TTCATGAGGAAAGGGAGGGGGAGGATGTGGGATGGTGGAGGGGCTGCAGA\n',
'CTCTGGGCTAGGGAAAGCTGGGATGTCTCTAAAGGTTGGAATGAATGGCC\n',
'TAGAATCCGACCCAATAAGCCAAAGCCACTTCCACCAACGTTAGAAGGCC\n',
'TTGGCCCCCAGAGAGCCAATTTCACAATCCAGAAGTCCCCGTGCCCTAAA\n',
'GGGTCTGCCCTGATTACTCCTGGCTCCTTGTGTGCAGGGGGCTCAGGCAT\n',
'GGCAGGGCTGGGAGTACCAGCAGGCACTCAAGCGGCTTAAGTGTTCCATG\n',
'ACAGACTGGTATGAAGGTGGCCACAATTCAGAAAGAAAAAAGAAGAGCAC\n',
'CATCTCCTTCCAGTGAGGAAGCGGGACCACCACCCAGCGTGTGCTCCATC\n',
'TTTTCTGGCTGGGGAGAGGCCTTCATCTGCTGTAAAGGGTCCTCCAGCAC\n',
'AAGCTGTCTTAATTGACCCTAGTTCCCAGGGCAGCCTCGTTCTGCCTTGG\n',
'GTGCTGACACGACCTTCGGTAGGTGCATAAGCTCTGCATTCGAGGTCCAC\n',
'AGGGGCAGTGGGAGGGAACTGagactggggagggacaaaggctgctctgt\n',
'cctggtgctcccacaaaggagaagggctgatcactcaaagttgcgaacac\n',
'caagctcaacaatgagccctggaaaatttctggaatggattattaaacag\n',
'agagtctgtaagcacttagaaaaggccgcggtgagtcccaggggccagca\n',
'ctgctcgaaatgtacagcatttctctttgtaacaggattattagcctgct\n',
'gtgcccggggaaaacatgcagcacagtgcatctcgagtcagcaggatttt\n',
'gacggcttctaacaaaatcttgtagacaagatggagctatgggggttgga\n',
'ggagagaacatataggaaaaatcagagccaaatgaaccacagccccaaag\n',
'ggcacagttgaacaatggactgattccagccttgcacggagggatctggc\n',
'agagtCCATCCAGTTCATTCAACACCTGGTTAGAAAACTGGGGCCAGCAC\n',
'ACAGGGGAAGGGTAAGCTGGTTTCATGATCGAATCAAGGCTCAGACAATT\n',
'TTTAAAGGCCAGAGGGTAGACTGCAATCACcaagatgaaatttacaagga\n',
'acaaatgtgaagcccaacatttaggttttaaaaatcaagcgtataaatac\n',
'agaaggtggagggaacttgctttagacacagttcaggtgaagaaagacct\n',
'ggaaacttctgttaactataagctcagtaGGGGCTAAAAGCATGTTAATC\n',
'GGCATAAAAAGGCAATGAGATCTTAGGGCACACAGCTCCCCGCCCCTCTT\n',
'CTGCCCTTCATCCTTCTTTCAATCAGCAGGGACCGTGCACTCTCTTGGAG\n',
'CCACCACAGAAAACAGAGGTGCATCCAGCACCACAGAAAACAGAGCCACC\n',
'ACAGAAAACAGAGGGTGACTGTCATCCCCTCCAGTCTCTGCACACTCCCA\n',
'GCTGCAGCAGAGCAGGAGGAGAGAGCACAGCCTGCAATGCTAATTTGCCA\n',
'GGAGCTCACCTGCCTGCGTCACTGGGCACAGACGCCAGTGAGGCCAGAGG\n',
'CCGGGCTGTGCTGGGGCCTGAGCCGGGTGGTGGGGAGAGAGTCTCTCCCC\n',
'TGCCCCTGTCTCTTCCGTGCAGGAGGAGCATGTTTAAGGGGACGGGTTCA\n',
'AAGCTGGTCACATCCCCACCGAAAAAGCCCATGGACAACGAAAAGCCCAC\n',
'TAGCTTGTCCAGTGCCACAGGAGGGGCAAGTGGAGGAGGAGAGGTGGCGG\n',
'TGCTCCCCACTCCACTGCCAGTCGTCACTGGCTCTCCCTTCCCTTCATCC\n',
'TCGTTCCCTATCTGTCACCATTTCCTGTCGTCGTTTCCTCTGAATGTCTC\n',
'ACCCTGCCCTCCCTGCTTGCAAGTCCCCTGTCTGTAGCCTCACCCCTGTC\n',
'GTATCCTGACTACAATAACAGCTTCTGGGTGTCCCTGGCATCCACTCTCT\n',
'CTCCCTTCTTGTCCCTTCCGTGACGGATGCCTGAGGAACCTTCCCCAAAC\n',
'TCTTCTGTCCCATCCCTGCCCTGCTCAAAATCCAATCACAGCTCCCTAAC\n',
'ACGCCTGAATCAACTTGAAGTCCTGTCTTGAGTAATCCGTGGGCCCTAAC\n',
'TCACTCATCCCAACTCTTCACTCACTGCCCTGCCCCACACCCTGCCAGGG\n',
'AGCCTCCCGTGGCACCGTGGGGACACAAAGGAACCAGGGCAAAGCTCCCT\n',
'CAGCCCCATTCAAAGAGGCCTGGCCCACAGGCTCACGGAAAGTCAGCCTC\n',
'TCATGCCCCGAGAGCTGAGTGCAAGGGAGAGGCAGCGCTGTCTGTGCTTC\n',
'CCATGCAGAAGCACCCCCCTCCCACCCCTGTGCAGGCCGGCCTTCGCGGC\n',
'AGACCACCATACACCACGTTCCAAGCCACACTGAGGCCTCCCTCCAAGCC\n',
'TGCAGCCCCCATTTCCAGACCCTGCCAGGGCAACCTGCATATCCACCTCC\n',
'CTACCCTGCCCCCCTCTTCCAGGAGTCTGCCCTATGTGGAGTAAGCACgt\n',
'ggttttcctcttcagcaactatttcctttttactcaagcaatggccccat\n',
'ttcccttggggaatccatctctctcgcaggcttagtcccagagcttcagg\n',
'tggggctgcccacagagctcctcagTCTAAGCCAAGTGGTGTGTCATAGT\n',
'CCCCTGGCCCCATTAATGGATTCTGGGATAGACATGAGGACCAAGCCAGG\n',
'TGGGATGAGTGAGTGTGGCTTCTGGAGGAAGTGGGGACACAGGACAGCAT\n',
'TCTTTCCTGCTGGACCTGACCCTGTGTCATGTCACCTTGCTACCACGAGA\n',
'GCATGGCCTGTCTGGGAATGCAGCCAGACCCAAAGAAGCAAACTGACATG\n',
'GAAGGAAAGCAAAACCAGGCCCTGAGGACATCATTTTAGCCCTTACTCCG\n',
'AAGGCTGCTCTACTGATTGGTTAATTTTTGCTTAGCTTGGTCTGGGGAGT\n',
'TCTGACAGGCGTGCCACCAATTCTTACCGATTTCTCTCCACTCTAGACCC\n',
'TGAGAAGCCCACGCGGTTCATGCTAGCAATTAACAATCAATCTCGCCCTA\n',
'TGTGTTCCCATTCCAGCCTCTAGGACACAGTGGCAGCCACATAATTGGTA\n',
'TCTCTTAAGGTCCAGCACGAGGTGGAGCACATGGTGGAGAGACAGATGCA\n',
'GTGACCTGGAACCCAGGAGTGAGGGAGCCAGGACTCAGGCCCAAGGCTCC\n',
'TGAGAGGCATCTGGCCCTCCCTGCGCTGTGCCAGCAGCTTGGAGAACCCA\n',
'CACTCAATGAACGCAGCACTCCACTACCCAGGAAATGCCTTCCTGCCCTC\n',
'TCCTCATCCCATCCCTGGGCAGGGGACATGCAACTGTCTACAAGGTGCCA\n',
'AGTACCAGGACAGGAAAGGAAAGACGCCAAAAATCCAGCGCTGCCCTCAG\n',
'AGAAGGGCAACCACGCAGTCCCCATCTTGGCAAGGAAACACAATTTCCGA\n',
'GGGAATGGTTTTGGCCTCCATTCTAAGTGCTGGACATGGGGTGGCCATAA\n',
'TCTGGAGCTGATGGCTCTTAAAGACCTGCATCCTCTTCCCTAGGTGTCCC\n',
'TCGGGCACATTTAGCACAAAGATAAGCACAAAAGGTGCATCCAGCACTTT\n',
'GTTACTATTGGTGGCAGGTTTATGAATGGCAACCAAAGGCAGTGTACGGG\n',
'TCAAGATTATCAACAGGGAagagatagcatttcctgaaggcttcctaggt\n',
'gccaggcactgttccattcctttgcatgttttgattaatttaatatttaa\n',
'aataattctaccaggaagctaccattattaccacaacttcacaaatgaga\n',
'acaccgaggcttagaggggttgggttgcccaaggttacagaggaagaaaa\n',
'caggggagctggatctgagccaaggcatcaactccaaggtaacccctcag\n',
'tcacttcactgtgtgtcccctGGTTACTGGGACATTCTTGACAAACTCGG\n',
'GGCAAGCCGGTGAGTCAGTGGGGGAGGACTTTCAGGAAGAGGTGGGTTCC\n',
'CAGTTGGTGACAGAAGAGGAGGCTGCAAAGTGAAGGAGCAGGGGCTCCAG\n',
'GTCTGGCGACAACCAGGGAAGGGACAGGGCAGGGATGGCTTGGACCACGA\n',
'GAGGCACCTGAGTCAGGCAGTCACATACTTCCCACTGGGGTCTACCATGT\n',
'GAGGCATGGTGTGGGATCCTGGGAAGGAGACCAAGCCTCATTTCAGTTTG\n',
'CTTATGGCCAAAGACAGGACCTGTGTACCCGACAACCCCTGGGACCTTTA\n',
'CCAAAAAAAGAGCAAACACCATTCACTCACTCATGTTAGATAAACACTGA\n',
'GTGAAGTCACTGGAGCCCAAGGACTGTGCGAGGTCAGCACTGCCAATACA\n',
'AGAagctgcagccctccagctcgcctccctcaatggccactccgtgctcc\n',
'agccatgctggcttccttttaggtcctccacctccaggctgtagttcatg\n',
'tgcttctttctggaatgttcttcccaacctacccactcaaccctcagact\n',
'ttaccataaatgtcatttcctcacgtctgccttccctgacctgagaccaa\n',
'gccaggcttcccatgacgagcctcacagtaccccatctCCCCTGAACAGA\n',
'TGCAGTAATAACCTACATAACCCGGGGCCATGATCTAtggctttgaatcc\n',
'tggctctgtcactaggccaggtctctcagcccttctgtgcctcagtttcc\n',
'tcatctataaaatgagatgacggcagtgcctgctcatgaagtgtgagtta\n',
'atgcactcaaatcaatggttgtgcacggtttatatgaatattagtgatta\n',
'CAAAATATTATCAATAGACCTTGTCACAACTGTTATTGAAGAACtaatca\n',
'tctattgcttatttaggtctttctctcctgccagaatgtgcgctccaggt\n',
'ggagaggtatgttgccttatccgtggctggatatatagagattcccacac\n',
'tgccttgcacacgagcactgctgggtaaatatttgttggctgcaggaaAA\n',
'CGTGAAGGAATAGGCCCTCCAATGGGAGGAAAAGCATGAGTTGTGAGAGC\n',
'AGAGCCACCACAGGAAACCAGGAGGCTAAGTGGGGTGGAAGGGAGTGAGC\n',
'TCTCGGACTCCCAGGAGTAAAAGCTTCCAAGTTGGGCTCTCACTTCAGCC\n',
'CCTCCCACACAGGGAAGCCAGATGGGTTCCCCAGGACCGGGATTCCCCAA\n',
'GGGGGCTGCTCCCAGAGGGTGTGTTGCTGGGATTGCCCAGGACAGGGATG\n',
'GCCCTCTCATCAGGTGGGGGTGAGTGGCAGCACCCACCTGCTGAAGATGT\n',
'CTCCAGAGACCTTCTGCAGGTACTGCAGGGCATCCGCCATCTGCTGGACG\n',
'GCCTCCTCTCGCCGCAGGTCTGGCTGGATGAAGGGCACGGCATAGGTCTG\n',
'ACCTGCCAGGGAGTGCTGCATCCTCACAGGAGTCATGGTGCCTGTGGGTC\n',
'GGAGCCGGAGCGTCAGAGCCACCCACGACCACCGGCACGCCCCCACCACA\n',
'GGGCAGCGTGGTGTTGAGACAACACAGCCCTCATCCCAACTATGCACATA\n',
'GCTTCAGCCTGCACAGATAGGGGAGTAGGGGACAGAGCATTTGCTGAGAG\n',
'GCCAGGAGCGCATAGATGGGACTCTGCTGATGCCTGCTGAGTGAATGAGG\n',
'GAAAGGGCAGGGCCCGGGACTGGGGAATCTGTAGGGTCAATGGAGGAGTT\n',
'CAGAGAAGGTGCAACATTTCTGACCCCCTACAAGGTGCTTGCTACCTGCC\n',
'AGGCACCCTTTCCATACCTTGTCTCAGTTCAGCTCCCCACCTTGGATAAA\n',
'CAAGAAACCTTGGTTGCAGAGGAAAAAAGAGGCTGGAAACAAAGGGGTAG\n',
'AAATGGGGTAGCAGGGGAGATTGCCTGATCAACTGCCAAATGGTACACAG\n',
'TTCTGGAAAAGCACAAAAAATGTGCACACACGGGTTCTTCCCACTTTAAC\n',
'CCCTGAGGAATCTGAGGCCTGCTCCTGAAACAGACTGGGCAGTGGCTAGT\n',
'GACTCTAGGTATAGGAGTATCCAGCCCTGCTCACCCAGGCTAGAGCTTAG\n',
'GGGGACAAGAGGAAAGAGGTGCCTGTGGGGGTGGAGGACAGGAAGGAAAA\n',
'ACACTCCTGGAATTGCAAAGTGAGGGCAGAGTCTATTTATATTGGGTTTA\n',
'ATTAACTCCTCTCCCTGGTGCCACTAAAGCAGCAATCACACTGCAGACAG\n',
'CACTGATTTGATTGGCAAGAGATGCACCAGGCAGAATATTAAGGGACCAG\n',
'GCCCCTATAAATAGGCCTAATCACAGCCCCTCACTGGAAAATGGTAAGGA\n',
'AGACATTAATCAGGCCTGGCACTGTGCCCTAGACCTGCTCCCCTAGGCAC\n',
'TACAGTGGGGCCCTTGGTTGCAACACAAGTAGGTAGGGATGGATGAGTGT\n',
'GGCATGAAGGGCCTAGGAGATTTCACTTGGGTTTAAAATGCTGTGACCTT\n',
'GAGTAAGTTGCCGTCTCTGAATCTGATCCTTTCGATTTCCCATTCTCCAA\n',
'ACTGAGAACTAGCACTGCTGAGACGTGGTTATTTCCAATAATAATTTGTA\n',
'TATTTTACATAACGCACCACACCAACATCTTCACCCAGTTGGAGCCTACT\n',
'CCTTTGCTCCCGCTGCTGGCTTCCCCAGCCCTCCCTTCTGCCCTCCTCAG\n',
'GCCAGCACTTTTCAGTGAGTTCCTCCTTTGCATACAGGCTTTCCAGATCT\n',
'GTACTTGCCTTGAATACTCATCAGAGCCCAGGAGTTACTCCTCACCTCCC\n',
'ACTTATTTTTCCTCCCATCAAATAACTAAAGCATGGCCAGCTGATGCCCA\n',
'GCCAACTGAGAAACCCAACCCTCTGAGACCAGCACACCCCTTTCAAGCAT\n',
'GTTCCTCCCTCCCCTTCTTTGTATTTATACTGATGCAAGTTTGCTGGCTG\n',
'TCCTAacttatttctgtgcctcagttctcccatatgtaagatcacaaagg\n',
'gggtaaagatgcAAGATATTTCCTGTGCACATCTTCAGATGAATTTCTTG\n',
'TTAGTGTGTGTGTGTTTGCTCACACATATGCGTGAAAGAAGAGTACATAC\n',
'ACAGATCTCCTCAAAAAGGAGGCAGCAAGCCCGTTCAAGAATGGGACTGA\n',
'ATACACCTGATGAGTGGTTTACTTTCTGTCTGcaaacatctactgatcat\n',
'ctgttaggtgcaggccatgatcacaacaaagacgaataagacactacact\n',
'agccagggagagtctcaaaaacaactaaactcaaattaaattcattctac\n',
'tccagtcatgggtacaaagctaaggagtgacaaatccctcttggagttag\n',
'gggagtcaggaaaaagctcttagcagaatgtgtgcctctcggccgggcgc\n',
'agcggctcacgcctgtaatcccagcactttgggaggcgaaggcaggcaga\n',
'tcacctgaggtcgggagttcgagaccagtctgaccaacatggtgaaactc\n',
'catctctactaaaaatacaaaattagccaggcgtggtggtgcatgcctgt\n',
'aatccccgctactcgggaggctgaggaaggagaatcacttgaaccaggaa\n',
'ggtggaggttgcagtgtgccaagatcgcgccatggcactccagcctaggc\n',
'aacgagggtgaaccaggtccaggaagaaggtgcaaagacagcattccagg\n',
'taaaagaaacagcttgaacaaaaagtgtgtaggggaaCCGCAAGCGGTCT\n',
'TGAGTGCTGAGGGTACAATCATCCTTGGGGAAGTACTAGAAGAAAGAATG\n',
'ATAAACAGAGGCCAGTTTGTTAAAAACACTCAAAATTAAAGCTAGGAGTT\n',
'TGGACTTGTGGCAGGAATgaaatccttagacctgtgctgtccaatatggt\n',
'agccaccaggcacatgcagccactgagcacttgaaatgtggatagtctga\n',
'attgagatgtgccataagtgtaaaatatgcaccaaatttcaaaggctaga\n',
'aaaaaagaatgtaaaatatcttattattttatattgattacgtgctaaaa\n',
'taaccatatttgggatatactggattttaaaaatatatcactaatttcat\n',
'ctgtttctttttacttttAGAAATCACATATGTGACTTAAATATTTCTTT\n',
'TCTTTTTCTTTCCTCTCACTCAGCGTCCTGTGATTCCAAAGAAATGAGTC\n',
'TCTGCTGTTTTTGGGCAGCAGATATCCTAGAATGGACTCTGACCTAAGCA\n',
'TCAAAATTAATCATCATAACGTTATCATTTTATGGCCCCTTCTTCCTATA\n',
'TCTGGTAGCTTTTAAATGATGACCATGTAGATAATCTTTATTGTCCCTCT\n',
'TTCAGCAGACGGTATTTTCTTATGCTACAGTATGACTGCTAATAATACCT\n',
'ACACATGTTAGAACCATTCTGACTCCTCAAGAatctcatttaactcttat\n',
'tatcagtgaatttatcatcatcccctattttacataaggaaatggggtta\n',
'gaaagaccaaataacattttttcaacatcaaaacactagcttgagatcaa\n',
'gcccagacttggatctgtcgtctgaattccaagctttttgttatttattg\n',
'atatgttttgttgtTTTCATGCAATAATGCAAATCTTAGCCCAAACATTT\n',
'TGTTAGTAGTACCAACTGTAAGTCACCTTATCTTCATACTTTGTCTTTAT\n',
'GTAAACCTAAATTAGATCTGTTTTTGATACTGAGGGAAAAACAAGGGAAT\n',
'ctaacactaaccagcccgtagtgtgtggtcaacactttcgttactttagt\n',
'atacatcaccccaattgtttgtcttcaccacacactttggagttaggtag\n',
'tagtatctatttttacaaataagaaaacccaggcacaaaggggttgatta\n',
'gcAATTATCTTTTGAAAAGCCTGTAGTTGCTCATCTGAAGAAGTGACGGA\n',
'CCACCTCTTATTTAGTGGACAGACAGTAACTAGTTGAGAAGACAGGGGAT\n',
'TTTGTTGGCGGAAAAAAAAATTTATCAAAAGTCGTCTTCTATCAGGGAGT\n',
'TTTATGAGAAACCCTAGCTCCTCAGTTCCACAGTGGGTAACTGTAATTCA\n',
'TTCTAGGTCTGCGATATTTCCTGCCTATCCATTTTGTTAACTCTTCAATG\n',
'CATTCCACAAATACCTAAGTATTCTTTAATAATGGTGGTTTTTTTTTTTT\n',
'TTTGCATCTATGAAGTTTTTTCAAATTCTTTTTAAGTGACAAAACTTGTA\n',
'CATGTGTATCGCTCAATATTTCTAGTCGACAGCACTGCTTTCGAGAATGT\n',
'AAACCGTGCACTCCCAGGAAAATGCAGACACAGCACGCCTCTTTGGGACC\n',
'GCGGTTTATACTTTCGAAGTGCTCGGAGCCCTTCCTCCAGACCGTTCTCC\n',
'CACACCCCGCTCCAGGGTCTCTCCCGGAGTTACAAGCCTCGCTGTAGGCC\n',
'CCGGGAACCCAACGCGGTGTCAGAGAAGTGGGGTCCCCTACGAGGGACCA\n',
'GGAGCTCCGGGCGGGCAGCAGCTGCGGAAGAGCCGCGCGAGGCTTCCCAG\n',
'AACCCGGCAGGGGCGGGAAGACGCAGGAGTGGGGAGGCGGAACCGGGACC\n',
'CCGCAGAGCCCGGGTCCCTGCGCCCCACAAGCCTTGGCTTCCCTGCTAGG\n',
'GCCGGGCAAGGCCGGGTGCAGGGCGCGGCTCCAGGGAGGAAGCTCCGGGG\n',
'CGAGCCCAAGACGCCTCCCGGGCGGTCGGGGCCCAGCGGCGGCGTTCGCA\n',
'GTGGAGCCGGGCACCGGGCAGCGGCCGCGGAACACCAGCTTGGCGCAGGC\n',
'TTCTCGGTCAGGAACGGTCCCGGGCCTCCCGCCCGCCTCCCTCCAGCCCC\n',
'TCCGGGTCCCCTACTTCGCCCCGCCAGGCCCCCACGACCCTACTTCCCGC\n',
'GGCCCCGGACGCCTCCTCACCTGCGAGCCGCCCTCCCGGAAGCTCCCGCC\n',
'GCCGCTTCCGCTCTGCCGGAGCCGCTGGGTCCTAGCCCCGCCGCCCCCAG\n',
'TCCGCCCGCGCCTCCGGGTCCTAACGCCGCCGCTCGCCCTCCACTGCGCC\n',
'CTCCCCGAGCGCGGCTCCAGGACCCCGTCGACCCGGAGCGCTGTCCTGTC\n',
'GGGCCGAGTCGCGGGCCTGGGCACGGAACTCACGCTCACTCCGAGCTCCC\n',
'GACGTGCACACGGCTCCCATGCGTTGTCTTCCGAGCGTCAGGCCGCCCCT\n',
'ACCCGTGCTTTCTGCTCTGCAGACCCTCTTCCTAGACCTCCGTCCTTTGT\n',
'CCCATCGCTGCCTTCCCCTCAAGCTCAGGGCCAAGCTGTCCGCCAACCTC\n',
'GGCTCCTCCGGGCAGCCCTCGCCCGGGGTGCGCCCCGGGGCAGGaccccc\n',
'agcccacgcccagggcccgcccctgccctccagccctacgccTTGACCCG\n',
'CTTTCCTGCGTCTCTCAGCCTACCTGACCTTGTCTTTACCTCTGTGGGCA\n',
'GCTCCCTTGTGATCTGCTTAGTTCCCACCCCCCTTTAAGAATTCAATAGA\n',
'Gaagccagacgcaaaactacagatatcgtatgagtccagttttgtgaagt\n',
'gcctagaatagtcaaaattcacagagacagaagcagtggtcgccaggaat\n',
'ggggaagcaaggcggagttgggcagctcgtgttcaatgggtagagtttca\n',
'ggctggggtgatggaagggtgctggaaatgagtggtagtgatggcggcac\n',
'aacagtgtgaatctacttaatcccactgaactgtatgctgaaaaatggtt\n',
'tagacggtgaattttaggttatgtatgttttaccacaatttttaaaaaGC\n',
'TAGTGAAAAGCTGGTAAAAAGAAAGAAAAGAGGCTTTTTTAAAAAGTTAA\n',
'ATATATAAAAAGAGCATCATCAGTCCAAAGTCCAGCAGTTGTCCCTCCTG\n',
'GAATCCGTTGGCTTGCCTCCGGCATTTTTGGCCCTTGCCTTTtagggttg\n',
'ccagattaaaagacaggatgcccagctagtttgaattttagataaacaac\n',
'gaataatttcgtagcataaatatgtcccaagcttagtttgggacatactt\n',
'atgctaaaaaacattattggttgtttatctgagattcagaattaagcatt\n',
'ttatattttatttgctgcctctggccaccctaCTCTCTTCCTAACACTCT\n',
'CTCCCTCTCCCAGTTTTGTCCGCCTTCCCTGCCTCCTCTTCTGGGGGAGT\n',
'TAGATCGAGTTGTAACAAGAACATGCCACTGTCTCGCTGGCTGCAGCGTG\n',
'TGGTCCCCTTACCAGAGGTAAAGAAGAGATGGATCTCCACTCAtgttgta\n',
'gacagaatgtttatgtcctctccaaatgcttatgttgaaaccctaacccc\n',
'taatgtgatggtatgtggagatgggcctttggtaggtaattacggttaga\n',
'tgaggtcatggggtggggccctcattatagatctggtaagaaaagagaGC\n',
'ATTGtctctgtgtctccctctctctctctctctctctctctcatttctct\n',
'ctatctcatttctctctctctcgctatctcatttttctctctctctcttt\n',
'ctctcctctgtcttttcccaccaagtgaggatgcgaagagaaggtggctg\n',
'tctgcaaaccaggaagagagccctcaccgggaacccgtccagctgccacc\n',
'ttgaacttggacttccaagcctccagaactgtgagggataaatgtatgat\n',
'tttaaagtcgcccagtgtgtggtattttgttTTGACTAATACAACCTGAA\n',
'AACATTTTCCCCTCACTCCACCTGAGCAATATCTGAGTGGCTTAAGGTAC\n',
'TCAGGACACAACAAAGGAGAAATGTCCCATGCACAAGGTGCACCCATGCC\n',
'TGGGTAAAGCAGCCTGGCACAGAGGGAAGCACACAGGCTCAGggatctgc\n',
'tattcattctttgtgtgaccctgggcaagccatgaatggagcttcagtca\n',
'ccccatttgtaatgggatttaattgtgcttgccctgcctccttttgaggg\n',
'ctgtagagaaaagatgtcaaagtattttgtaatctggctgggcgtggtgg\n',
'ctcatgcctgtaatcctagcactttggtaggctgacgcgagaggactgct\n',
'tgagcccaagagtttgagatcagcctgggcaatattgtgagattccatct\n',
'ctacaaaaataaaataaaatagccagtcatggtgtcacacacctgtagtc\n',
'ccagctacatgggaggctgaggcgggaggatcacttgagcttgggagatc\n',
'gaggctgcagtgagctatgattgtaccactgcactccaggctgggcgaca\n',
'gagagagaccctgtctcagaaaaaaaaaaaaaagtactttgtaatctgta\n',
'aggtTTATTTCAACACACACAAAAAAAGTGTATATGCTCCACGATGCCTG\n',
'TGAATATACACACACACCACATCATATACCAAGCCTGGCTGTGTCTTCTC\n',
'ACAAATGCACTGCTAGGCACCACCCCCAGTTCTAGAATCACACCAGCCAG\n',
'TTCACCCTCCAGATGGTTCACCCTCAACTTCATAAAAGTTCCCTACCTAA\n',
'TCTACTGACAGGCTCATCCCCGACCTAATTTTAAAGATTTCCTAGGAGCT\n',
'GCAGTGGGAATCCTGGACCTCAGCCTGGACAAAGAACAGCTGCAGGTCAT\n',
'TCTCATGTGTGGACACAGAAGCTCTGCCTGCCTTTGCTGGCCAGCTGGGC\n',
'TGAGCGGGCCTGGGAATTAAGGCTGCAGGGTTGGTCCCAGGCAGTCTTGC\n',
'TGAAGCTTGCCACATCCCCCAGCCTCCTGGATTTGCCAGGATCCAAGAGC\n',
'ATGGACTTTAGGAATTCCTGGTGGAGGAGTGAAGAAAATGTGACAGGGTG\n',
'TCCTAAGCCCCGATCTACAGGAAGAAAACTGGAAATAAGACTGAGGACTT\n',
'AGTTTAAGATGTTCCTACTCAGCCTCTAGCTTTTGTGCTACAGTTCTGGG\n',
'AACAGACTCCTCTCTCCTGAAAACCACTTCCCTCCGCAGCATTAGATTTC\n',
'ACCAAGATGTCTTGCTTGTGGGAAAGACTTCCAAGGATGCCTGGAGAGAG\n',
'GAGGATGGAAATGTCCTGCTCTCTAAACAGATAGACAGATGCAGCCAGAC\n',
'AGAAAATAGTTTATCTTGCTGAGGTTTCTAATGTATTTGAAAGAGGCCTG\n',
'GGTCTAGAAGTCTACCCAGAGGGCTCTGTGTTGTGCACGCAAAGATAAGA\n',
'ACCTTCCCTGTGGGAGTTCCAGAGCCAGTTTTCATAAACACCCATCGGTG\n',
'ACTGTGTTCAGAGTGAGTTCACACCATCCTGACCTGCCCTGAGTTAGACC\n',
'TTACATGGTCTTCCTCCTCTAGGAAGCCTCTGCAGCCCAGGAACCTCCCC\n',
'TTATCGGAAATGAACAGCATTTGAAGCTTCACCAGACAGACCAGACAGCT\n',
'TAGCCCTCGTGTTGTGCCATGTGGGTTGTTCTCTGAGAGGcaggagagca\n',
'tagtggttactaggaagggaaggactttgggactagactgcctcggctgg\n',
'agtcctctttctgcttcatagccacgtgatcctaggcatgttacctgtgc\n',
'ctcagttttcactctgtcaatatgtaataactgaatctgtctttgtggtg\n',
'aggattcagtgagttaacatatttgaagtgcttaaaaATGAGGCTTGtgt\n',
'ccatagattaatgagtgaatacacaaatggtgatatggacatacagtgga\n',
'gtattagtcataaaaaggaaggcagagctgatccatggcaccatgtgaca\n',
'gaacctcaaaagcattaggttaagtggaagaagccagacacaggtcacct\n',
'attgtgtaattccatttataggaaatatacagaatatgtaaatccgtgga\n',
'gaaagaaagccgatttccaggggctaaggggaggggagaatgggaagtgg\n',
'ctgcttcatgggtacaaggtttcattttgagctgatgaaaatgttttgga\n',
'actacatagagatagtgttggcacaacatggtgaatgtactgaatgccac\n',
'tgattgttcaatttaaaatggtcaaacttatatgaatttcacctccatta\n',
'aaaaaaAAAAAAAAGgaccagatgtggttgctcacacccataatcccaac\n',
'actttggaaAAAGGTGAAAGTTTTTTTTtctttttttttttatatactta\n',
'agttctagggtacatgtgcataatgtgcaggttggatacatagatatgcg\n',
'tgtgccatgttggtttgctgcacccatcaacttgtcatttacattaggta\n',
'tttcttctaatgctatccctcccccagccccccacccactgacaggcccc\n',
'agtgtatgatgttctctgccccatgtccaagcgttctcattgttcaattc\n',
'ccacctgtgagtgagaacatgcagtgtttggttttctgtctttgtgatag\n',
'tttgctcagaatgatggtttccagcttcatccatgtccctgcaaaggaca\n',
'tgaactcatcctttttaatggctgcatagtatcccatggtatatatgtgc\n',
'cacattctcttaatccagtctgtcattgatggacatttgggttggttcaa\n',
'agtctttgctattgtgaatactgccacaataaacatacatgtgcatgtgt\n',
'ctttatagtagcacgatttataatcctttgggtatatacccTAAGACctg\n',
'ggacgcatttaaagcagtgtgtaaagagacatttatagcactaaatgccc\n',
'acaagagaCCTCTGCCTGAGAACGTGGGTTTCAGCCTAAGAGTTGTAATA\n',
'TGTGTGCCCATTCACAGGTGCTGCATCAGAGTCCCAGGTGGGAAGAAGGC\n',
'AAGCATACACAAAAATGGTAAAAGGCAGAAAGGAGCCCAGTCTCGTTCTT\n',
'TTTAAGAAGTTTTCCTAAGAATCTCCACCCAGCGACTTGCTCTCACATCT\n',
'TCTTGGCCAGCACTGGACCACACAACTCCTTCTAGATACAGAGGAGTCCT\n',
'AGGATTCTATGAGAAAGAAGGGGAGGGTGGGCAAAGGGCAGCCAGCTGTG\n',
'CAGCATCTGCTGGAGACACCTAACCCTTGGTGGAGGGGTTGTGGTGCTGG\n',
'gagaaggctttctggacggtgtgacagcagagataaacttaaaggccaag\n',
'taggagttaccctggtgaagcagggcagggttacaagcattccagcaaca\n',
'tgaagcagcaGGAGtgttttaattaaaagaaggcagttgctgtaaccaac\n',
'tataaacaaataaaggcttaaacacaatggaagtttatttctcactaagg\n',
'gaacatccaaatccatgatactttaagtcagggacccaggttcctcccat\n',
'ctatggttctgccatcactaatctgggtcttccacaattgccgtgctcct\n',
'tggaggtgggaagagcaggcggaggacacgtgggaggttttagggacaag\n',
'cctggaggcagcatgcgtcactcccatgcagagtccattggccaatgctg\n',
'gctccgatggccacatctcactgcaggggcagctgggaaatacagtctgg\n',
'ctgtctacccaggaggaagagCAGCCAGTTTCTGCTGCTGATGATCAGGA\n',
'GGTGGAGAAAATGTTCAGTCAGGCAGGGAGTGGGAATAGACAAGACCACA\n',
'AGCAGCTTGGTGCCTCTGAAAGGGAGAGGGGTGGAGGGGAGACTAGAGAG\n',
'GTGGGTAGGAATACTGGATTCCACTGACCACGTGCTGGATGTCACGCTTA\n',
'GCCCTCCTGCTCTGTGCCGGGTTAGGCACCTGGTGTTTTACGTACATAAT\n',
'CTCAATTCTGTGAGGGCATCCGACCTGTGGGAAAAGAGCTGTTTGTTTCA\n',
'AATGCCAGTCCTGCTTcctaacaagtgtttagagcttaatcgtgttcaaa\n',
'atacatatacaatgtttaatacttacaagaatttggtggggaaaatatta\n',
'ccatctttcccttttgtgattggagaaaaatgaggctttgaagggtttaa\n',
'gaacttgcccaaggtcggccaggtgcagtggctcatgtctataatcccaa\n',
'cactttgggaggctgaggtgggaggatcgcttgaggccaggagttcaaga\n',
'ccagcctgagcaacatagtgagactttgtctctataaaaaataaataaaT\n',
'AAATAAAAACAACTTGTCCAAGGTCAGACAGGCAGCCTCTTAGTAAGCAC\n',
'ACATATCCTCTATATTATACTACCTCTCATGGAGGATCTCCTGTGTTCTA\n',
'CAAATAGTCTGGACTTGAGCCAGAATGTGTTATAATCCTGGGATCACGGC\n',
'CAGTGGGCTTAGAAGAAGCCATCTCTTTCTCATGCCAAGATGAGGCTCCC\n',
'CCAGATTTGCTCAGACTTACCTATAGTCAGCAGCATCGGGGGTCAGGAAA\n',
'GACTTCACGAAGCCATAAATGCATCCTTCTCGGGGCAGCACCTGGCTCTC\n',
'CCAGGTGAGAGAGGACTCCATTTTCACAGGCAGGCGTGGGAGCTTCAGCA\n',
'CCCATCTCTGGGCCCAGAATGACCCACTGGAGACCTTACAGCTCTCCTGT\n',
'CACCCCCAATTCCTGCCCCCTCTGCAGCCTTGGAGGAGAATGGAGCTGAA\n',
'GGGCCTGCCCTCTGTAGGGTGAGAAAGGGAGGCTAAAGCCTGGTGCCCAC\n',
'TGCCCTGGCTGCTCCGCATTGCAGGAGCTGCGCCCTTCCTTTCCTGGCAC\n',
'AGGGTCCACAGCCCCGAAACCCCGTTGTGTGGGAGCTGGGCACAGGGCAG\n',
'CAGGACTAATCCTTGGAACAGCTCAGGGAGGATTATCCCAGCCACTGTCA\n',
'GCAGCGGTGCAGCTGGCTCATTCCCATATAGGGGGAGGCCAGAGCCAGGG\n',
'GCCTGCCACAAGTTGGAAGGCTGGGGAAGGGGAGGCCAGCAGAGGTGTCC\n',
'TGGCTGTGGGTGGCTCTGAGGGGGCTCTCAGGGGTGGGGCTAAATCTCAG\n',
'GGGCAGGATTATGTAAATCAAACCAATTCTAGCCACAGATTTAAAGTTTG\n',
'GAAAAAAAAAAAAACCCAGCCTGGCGGAAAGAATTTAAATTATAAAAACT\n',
'TAGAAGTATGGAATGTGAAATCATCCTGTAGGTGCTTATTTAACAACGAA\n',
'ATCATCCCGACACAATGAGCCATATGTGAAAAGTCATCCTTCCCCAACAC\n',
'ATCCCCCAACAGGCACTCCTCAAGCCTCTCCCACCCAAGTGCTGGCATCC\n',
'TCCCTGTCCTGCTTCACCTGAGACACCCCTTGTCTCATTAGACATGCAAC\n',
'TACGGGAGGGGTGACAGGAAGACAAGACACTATTTCCTCAGGCCCAGTTT\n',
'GGTGTGGGGAGAAAGCCTCCTGATCCTGAAAGCAAGAATTTGACCAGAGC\n',
'AGAAGTAATCAGTATGCAGATTGATTCTGTGGTATGTTAATGTTTATGCA\n',
'TAGATTATGAGGACCAGGTGAAAAGTGGGCCAGGGGAGCCAGATGTGTGT\n',
'GTGAGTCATGGGTGGCTGAGATGAGGACAGGAGGGAAACTGGTTTGGAGG\n',
'GTGCTGGCGATGGGGTGGGGGTGCCAGGAGGAAGGGAGGCTAGTTGTTTG\n',
'AATGTCTGCATGAAAAAGCGGACGACAGCGGGGTCTGGGTGAATTCGGGC\n',
'AACCATTTGGACCGTGGAGAAAACTGCCTGCGTGCGGCTGAGGACCTGCA\n',
'CTATTAATTTGTTTTTTAGCTAAGGCAAAGATAAATATAAAAACtgatac\n',
'tccacccagttaccagaaaacatttaggtatgtgtgagacaacttgggta\n',
'tgtgaacctaccttttcaatgtaaattcagtgaaatctaagtgcagatcc\n',
'catatttccaataaaaaggtaacatccaaactcagatgtcctatgagtat\n',
'aaaatacacaaagatcttctggacttagtatgaaaagggatttttttttt\n',
'gtcaggtacctcactagttatttttaaaataggattgcatgttgaaatga\n',
'taatcttttggatatattgggttaaataaatttattattaaagttaattt\n',
'cacttaaaaatgtttaatgtagctactagaaattttaaaattaagcatgt\n',
'tgctcaccttatgtttctattggacggctctCTCTAGATACAAAGGCTGC\n',
'CAAGAGGGACCTCACTCTAGCTTCAGGGAGAAGAGAGGAATTAGCAAGGC\n',
'CAAGCAGAGGCTCCTGAGGGCAGGGCCAAGGGCGGCTTGGTGGGGTGGGG\n',
'ATGGGATGCACAGAGATAACTCCAACCCTTAAGAAGGTGTTTCCTAGAGC\n',
'AGGCTGTGACCTGTCAGTTTATATACTGAGGCTTAGGAGCCTCTTGGATG\n',
'CCCCCAGATCTGCACCCCTGAATTGCCCTGTGCCCCTGCCGTCTTTGTTC\n',
'CTGTGCTGGCATAGTGGTCTCACCTCCGGCAGtatcaccaccactgggca\n',
'caagcttctccagcacagcaactgtgtcttatttctccttgtactcccag\n',
'tgttcacaccatgctgcactcacagaagactcttcgttgatattttgtgg\n',
'acagagagaatGCCTGTGAGAGTGGGCTGAAGTGTGCGTTGGGCTCCAGA\n',
'GACCTTAAGGAGGGGAGACCAGGTCCTGAGTAAAGTTGAAGGGGAGGGGC\n',
'TGAGTCCTGCTAGCCAGGAGTCTCATCCCCTGGGGAAGTTCCAGGGACCC\n',
'CTCAGAAGTGCAAGGGGACGGTGTTAGTGTTAGTCCAGTAACACAGCCCA\n',
'GAGCCTGCcttccacgtgggtttgacaggagcctcctaactgctcttctg\n',
'cttccatttttgccccttcagtctattctcaacagggaagccagaggcat\n',
'ccttaaccatgtcagatcatgtggctcctcagctcaaagccTCATCTCAG\n',
'AGGAAAGCTCTGGTCCCTTAGAAATGGCCCAAGTGGTGACAGACAGACTC\n',
'TAAGGtgagcagactgttgctagatatctgggctcggaggactcgccact\n',
'gctcaaaggcagtgaggattttcgcactagaagctggaggacagggatcc\n',
'ttgttaggtaggagcagaaagcttagaaaagtggtctcctgcagttacgt\n',
'ggcaaacacatcatgtaagtgataaattgggtatgcagttgaggagattt\n',
'ccaagtaaaatgttgaggatgctgcctggtttcttcttactgcttataat\n',
'atagtgtgagagaagagagataaattgagaaagagactggtttttaaact\n',
'gttaaaattgaatcaggacttgatgattttgaaaattgtcagtctcccca\n',
'catggaaaaagatgctgaaattaacaaatggcttctgagcatgtggcata\n',
'gggtgtaactgtacagtcttttgtgattatgcataaagatcaaaggatgg\n',
'gagtagcaatgagtcacacagaggtctgttgcaagagattacaagggtgt\n',
'accatgcagaacctctccaccaaaccttagggcccttgggaagcttcagt\n',
'gagttaccctgggggccatcttggcaggagctgaaggtagaaaggtagag\n',
'tttatctctaaaagattcatgggtatggctcttgacaaatcgactatgag\n',
'ccccaccgaaacccacagaggacaggcaaagggtttgggaaagctgtttc\n',
'acccacagtgctggcagattggtctgtaggggacagagtgcaaaatgaaa\n',
'gaagactgtcagagaccccaaactctgctgtcaagaagaaggctgataaa\n',
'actacttggctgcaaacacgtggatctttcgtgagaaaagaaggatgacc\n',
'cagaggcagaagcccagaaggcagagccaagagacatggaatcttcccac\n',
'atcttaaaacctgtttagggaacaccagcatctgtccagctggatttcag\n',
'aaccaccattccttcatccttcccctgctgcctctttctgaacagcaatg\n',
'tctcaagctttacccaccattgtgtgttgcatatgtagggggcagatagc\n',
'ttgtatctttagttttccagatcagaggaacatccaaagaaatctgttct\n',
'acacctaaacccgatttagatgagattcgggactgtgagcatgaagggat\n',
'ctcaagaggggtgaatgtgttttgcatgcacaagggacaggagtcttggg\n',
'gacagaggacaggctgtggtggcagatactaaggtgacccccacaacccc\n',
'cacctctgccattcacacccttgaataatccccttctctggttgtaagca\n',
'gaacctgtggcttgcttatgaaggaggcggtatatatgtgattcatgtac\n',
'tgatcatattgtataagatcactggctggatgcagtggctcgtgcctgta\n',
'atcccaacactttgggaggctgaggcgggtggatcacctgaggtcaggag\n',
'ttcgagaccaggctggccaacatggcaaaaccccgcctctactaaaaata\n',
'caaaaattagccaggcatagtggtgcacgcctgtaatcacagctactcaa\n',
'gaggctgaagcaggagaattgcttgaactcaggaggtggaggtggcagtg\n',
'agccaagatcgtgccactgcactccagcctcagtgacagagcgagactct\n',
'gtctcaaaaaataaataaataaaatgttaagatcataacctgtctttctg\n',
'gggactctctcttgacgcctttgaagaagcaggctgccatgttgcaagct\n',
'gcctcatggaggggatcagctgcgaggagctaagagccccctccagtcga\n',
'tgctcaccaggaagctgaggtcttgtgtccagcaccctgcatagaactga\n',
'atgctgccatgtgagcttggaagcagagccatccacacagctgagcccta\n',
'gatgagaacccagtgctggctgacaccctgatggcaccttacagaggacc\n',
'agttaggctgtgccaactcctgacctgcagaagctggggaacactgggtc\n',
'gtatttgcagctgctggatttgtgggaatttgtcacacagcaatTGGGAG\n',
'TCACACAgcctgtgacgccccaacaatccacacctcctgcatctccctgc\n',
'cttcacttcctagcacactgccctgactccctctgccgcagccacgctgg\n',
'ccctctgctgttcttcgaagccaccagggctgcattggctcccagccttt\n',
'gctctcactgctttctcctcctagagagcccttcctgcatgtatatgttt\n',
'gactcactcccttgcctccttcagacttgtacttaaaaatctcagtaagc\n',
'atttccctggctacccttttaaaaattgcaacccacttccatccccatcc\n',
'ccaacatgccatatttcctttcttctTCttccttcttccttttttttttt\n',
'ttttttttgacacaggttctctgtcacccagcctggagtgcagtgacatg\n',
'atctcggctcactgcaacctctgcctcccCAGGCAagaaaaggggaggat\n',
'gccaataaaggatgcattgatttgtatttactacagtggacatcaagggc\n',
'acattcttgctgtggccatcaagagactgtataaattctatgacttgtag\n',
'ttgtcccacttaagaaacaaagaagctgTGCATTTCTTTACTGGTCTAGA\n',
'GCTGCTCTAGGGCATTTTCTCTACAGCAATTCTAGGTTTCCCCACCTTGT\n',
'GAGTTTAGCTTTTTCTATATTCAAAGAAAAGTCCTCAGCCAGAGATTCTC\n',
'AGGAGCTTATAGAACAATCCAAACTCTTGGGAATATTAAGTGGAGAGGGG\n',
'TACGTGCAAGACACCAACAGCACTAGAAACAGTCCACATCTTTCCATGCG\n',
'TGGAGGAGTTTATGCTCTATGTGAGTTCACTCCATCATTAATTCTTCAAA\n',
'CACAAGAGTGTTAAAGGAACAAGAGTTAATGGGTCCTGTCATTACACTTG\n',
'TTCCCAGGATGACATTCTTCATCTTCCTCTTCTACAACCTGTTCTATATT\n',
'CCCCTCATGTTTATCCAGTGCTTCTGCTAGTCTAGTTCACTTCCAAAGAC\n',
'CCATGATTACCATGGCCCTGTCAGGCTGTAATTGCTGCAATTTCCAATTT\n',
'ACAATTGTCATCATCTATGGTTGATAAAGGTATAGCAATATTTctatttc\n',
'ctcatgataatgaaggtcaattacaactgccagtataataacttatttct\n',
'ttgtctgccaacctacatacacaaggaagccaaaatgacagggagctact\n',
'aaaactttattcttattggaatgcttactatgtacccagaagaagcattc\n',
'tccctactccagcagagcttaatgctgtaggtccaggaagctcaaattct\n',
'ccaagggagttttagtgagaggagccactctcaccctctgcccttggttt\n',
'acaaacctgtatattctaggacccaatatcttacaatgtccattggttca\n',
'aagtataacatgttaaagcacagagccccaactctgaaaagtaccatccc\n',
'taaattggcatttagttgcacctttatatccacctttaaaagaaatatct\n',
'tttaatgttctatcagactgatagattctgtttaatatagtatattatag\n',
'caccagtggatcatttggttgtatgcatattattgtaccttctctgctac\n',
'aaaatatattcctttgtcctaaggtgtgttacaaagaacattaggcattc\n',
'tatgcatctttggatagtttaatggccaagacattgatggcaggagagtc\n',
'aaagccacaggtggaaaacacatttatcccagtaagaacaaattgctatt\n',
'cttccactgtagagagggtaaacaatgtgccattacgttgccaattgaat\n',
'gcctcaatcatgtcaagggctgaacatctatgactgtttctgaaaggtca\n',
'aacattcaacagaggctgtagctagaactgccttaatgataagagatcat\n',
'gctgaattacccatgcaaaaccttaatacttgacacttatcactacttta\n',
'ttcaagagcctattgtgcaagcaTAAGTGGCTGAGTCAGGTTCTCAACTC\n',
'TGCTCATTAATACTATGCTTGGAGTATACAGTAAGATAAGAAACATAAAT\n',
'AAGAAGTGTACATTTGTTTcttcctgttttcttctggctattggatcaat\n',
'tacatcccatcttaagctgacccctgtgtaattaatcaatatccgtttta\n',
'agcagcaatccatagttgtgcagaaattagaaaactgacccacacagaaa\n',
'aactAATTGTGAGAACCAATATTATACTAAATTCATTTGACAATTCTCAG\n',
'CAAAGTGCTGGGTTGATCTCTATTTACGCTTTTCTTAAACACACAAAATA\n',
'CAAAAGTTAACCCATATGGAATGCAATGGAGGAAATCAATGACATATCAG\n',
'ATCTAGAAACTAATCAATTAGCAATCAGGAAGGAGTTGTGGTAGGAAGTC\n',
'TGTGCTGTTGAATGTACACTAATCAATGATTCCTTAAATTATTCACAATA\n',
'AAAAAAAAGATTAGAATAGTTTTTTTAAAAAAAAAGCCCAGAAACTAATC\n',
'TAAGTTTTGTCTGGTAATAAAGGTATATTTTCAAAAGAGAGGTAAATAGA\n',
'TCCACATACTGTGGAGGGAATAAAATACTTTTTGAAAAACAAACAACAAG\n',
'TTGGATTTTTAGACACATAGAAATTGAATATGTACATTTATAAATATTTT\n',
'TGGATTGAACTATTTCAAAATTATACCATAAAATAACTTGTAAAAATGTA\n',
'GGCAAAATGTATATAATTATGGCATGAGGTATGCAACTTTAGGCAAGGAA\n',
'GCAAAAGCAGAAACCATGAAAAAAGTCTAAATTTTACCATATTGAATTTA\n',
'AATTTTCAAAAACAAAAATAAAGACAAAGTGGGAAAAATATGTATGCTTC\n',
'ATGTGTGACAAGCCACTGATACCTATTAAATATGAAGAATATTATAAATC\n',
'ATATCAATAACCACAACATTCAAGCTGTCAGTTTGAATAGACaatgtaaa\n',
'tgacaaaactacatactcaacaagataacagcaaaccagcttcgacagca\n',
'cgttaaaggggtcatacaacataatcgagtagaatttatctctgagatgc\n',
'aagaatggttcaaaatatggaaaccaataaatgtgatatgccacactaac\n',
'agaataaaaaataaaaatcatattatcatctcaatagatgcagaaaaagc\n',
'attaacaaaagtaaacattctttcataataagacatcagataaaacaaat\n',
'taggaatagaaggaatgtaccgcaacacaataaaggccatatataacaag\n',
'cccacagctaacatcataatagtaaaatcatcacactggtaaaaaaaatg\n',
'aaagcttttcctctaaggtcagaaataatataaaggttcccactcttgct\n',
'atttctattccatatcgtactaaaagtcctagccaggacaattagacaaa\n',
'ataaaaataaaaacacccaaattggaaagatagaagcaaacttttctgtt\n',
'tacagataacataatcttatatgtagaaaccccttaaaacttcagcaaaa\n',
'aaaaaaaaaaaactacagagctagtaaattcagtgaagttgcagaataca\n',
'aaatcaacatacaaaaatcagtagtgtctctatacactaataaggactta\n',
'acagagaaagaagttaagaaaacaataccactaacaatagaatccaaaaa\n',
'ataaaatacttaggaataaattttaccaaacatctgtacactaaaaacta\n',
'taaaacattgaaaaaagaagttgaataagacacatataaatagaaagcta\n',
'tctcatgttaatagattagaaaaagtaatattgttaagatgtcctcacta\n',
'cttaaagcaatttatagatctaatgcatttattgcaatctcttcaaaatc\n',
'ccaaaggtatttttgacagaaataaaaaaaaaattctaaaatatgcatga\n',
'aaccacaaaagactgtgaatagctaaagcaatcttgagcaagatgaacaa\n',
'cactggaagcatcacactaccttatttcaaaatctactacaaagctatag\n',
'tgatcaaagcaacatgatactgtcataaaaacacacagataaacctatgg\n',
'aatggaataaagagcacagaaataagtccacacatttacattcaattgat\n',
'tttcaacaacaatgtcaagaagacaatggggaaaagacaatctcttcaat\n',
'aaatgatgctggaaaaactatatatccacatgcagaagaatgcagttgaa\n',
'tcctgatttcataccatatgcaaaattcaactggaaatggattaaataca\n',
'aatttaaaacatgaaatggtataactattagaacaaaacatagaaaatat\n',
'tcttcctgacattggtttgggccatcatttttctgatatgactctaaaag\n',
'cacaggcaaaaaaagaaaaaatagacaaatgagactatgccaaattaaaa\n',
'aatttctaacaacaaaagaaacgatcaatagagtgaaaaagataacctct\n',
'tgaatgggagaaatatttgcaaactactcatccaaccggggattgatatc\n',
'cagaatatacaagtaacacaaatatgtcaaaagtaaaataaataaataaa\n',
'taaataaataaataaattaaataaattatttaaaaatcggcagaggacag\n',
'gaatagacatttctcaggagacaacatacaaagggccacagatacatcaa\n',
'aaaatgctcaacatcactatttgtcagggaagtactaattaaaaccaaaa\n',
'tgagatgtcccctcaaacctgttagaatggctattatcaaaaagatgaaa\n',
'gatagcaactatcagagaggatgatagaaaagggaacccttgcatcatgt\n',
'acaaattaaaaatagaactatcacatgatccaagaatcctacttctgggt\n',
'atatagccaaaggaattgaaatcaatatgtcaaagggatatctgcactcc\n',
'tatgttattgcagcatgttcacaatggccaagatatagaatcaacctaac\n',
'tgttcatagacagatgaatggataaatgaaatgtgatatggaaaattatt\n',
'cagccttaaaaacagtaggaaattctgtcatttgagacaacgtggatgaa\n',
'cctagaggacattaagctaagtgaaataagctagacacagaaagacaaat\n',
'attgcatgatctcacttagaatctaaaaaatctgaactcatagaagcaga\n',
'gaatagtatgatggttactagggttatctggcagggagaggatgaggaaa\n',
'tgggacattgttaataaaaggaaaaaaattcaattagtaggattacattc\n',
'aggggacccaatatacgacatgttgactgtaattaataatgtattgtatg\n',
'cttgaaaattgctaatacagtatattgtaaatgttaatatgaggtaatat\n',
'atgtgttaattaacttgatttattcattcaacaacatacacatatattaa\n',
'aacatcacactgtattccacaaatatatataatttttgtcaattaaaaaa\n',
'taaTTTTTAAAAATGAGAAACAAAAAAGCTGACATTTTCAGATTAAAAAA\n',
'ATTATACAGAAGAATTAATTCATTAAAGTAAAAACAAATGTGGGAAAATG\n',
'GTTTTTAAATATAATTTAAACCAAATTTAAAATAAGcatataaagactat\n',
'ggacaaaacaagaaatccaaataaaaaataaacatatgaagaatattcaa\n',
'actcactttttatcaaagaaatgtaaattttaaaataTAGCATTGCTATT\n',
'GTGTTTTCATAAATAATAATATATCATGGATGAGCCTGTGAGGAAACAGA\n',
'CACTCATACTCTGCAAAGCAATGACTAAgataattatgtcagatcatgaa\n',
'ttacgttaattagcttgatggtggtcactgtttcacgataaatatacata\n',
'tgtatcaaaacatcacattacacaccataaagatatataacttgttatCA\n',
'AAAAGAAATATAGCAGTTAAAATTTAAAATTTTTAAAAAACGTCTTTTTG\n',
'AGGTTCGTACCTCACTTAAGTCACACTGTTCAAAATATTCATGCACTCAT\n',
'TTCTCTCATTCATGTGTTAATGTACAGGGTACGGGCCACTATAAATTCCT\n',
'TCAGCAACTGGAAAGGAAACTTTATGTACTGAGTGCTCAGAGTTGTATTA\n',
'ACTTTTTTTTTTTTTtgagcagcagcaagatttattgtgaagagtgaaag\n',
'aacaaagcttccacagtgtggaaggggacccgagcggtttgccCAGTTGT\n',
'ATTAACTTCTAATTCAACACTTTAAGATTCTTAGCATTATTGCAGACAAC\n',
'ATcagcttcacaagtgtgtgtcctgtgcagttgaacaagatcccacactt\n',
'aaaaggatcctacactttttttaatgctctgctgtttctgccttgaaatt\n',
'cttaacaatttttttaaccaaagtcctcacaaattcagtttacattagcc\n',
'ctgcaatcatgtagacatcctgATTCCAGACAATGTGTCTGGAGGCAGGG\n',
'TTTACAGGACTTCAAGAACCTTACCTTCTCAACTTTCATCTGCATCTTTA\n',
'CTCCCAACTATATATGAAGATGATGAAGATAGATATGGATGGTGCTTCTA\n',
'CCATACCCTCTTCCTCTGCCAAACTTCCTTGATCTAGGATAAggtcagta\n',
'aacttcttccgtaaaaggccaaaagtaaatattataggctctacaggccc\n',
'tagagtgtctgtcataactactcaactcttattgtagcataaaaactgtc\n',
'aacagacaatacagaaacaaatgagtgtgactgggttccagtgaaacttt\n',
'atttacaaaagatttgtcccatgagtcaaatttaccacctccAGATCTAG\n',
'AGAAACAGTTTTGAGCCCTTTTATTTTGCTCAACAGTTAAGCATGGCTCC\n',
'ATGTCCCTTATATTTAGTCAGAACTCGGTATGTTTTAAGGAAAGAATGGT\n',
'TACACGAAGACATACATTCATTCATTTATACAACACATTTTCAGTGTTGA\n',
'ATGATAAATTTTGGAATAGTTAACAGATGATAAAAGTGTTGTTTTCAGTC\n',
'ATCCCTATCCAATGAAGTAAAAAAAAAAGTGTTGAATGGGAAGAAATCAA\n',
'GAATAGTTATACGAATATCACCATTGCATTAAAGCTCTCTTCCTTGTTTC\n',
'TAAAAGAATATCTTGACACACATTAAGCTCACTGACCCCCACACCATGAA\n',
'TGAGGGCATCTTCAACAATGGTGGATGACGTCTTAGTTTCCCTCAACTCA\n',
'GTTAATCTAAGTAAGCTCATGGTATCACTTTCCTGTCCTAGAGGGAACAT\n',
'ATTTCCTGCATTTTTCTTTTTTTCCTTACTTTCCATCACCAAGTAACTCT\n',
'TCTGATATTTTTTCTCTTGAGAAAATTAATATGACTCATAGATCTGGTTC\n',
'CCAAGAGAAATCAATGGAGGCCTGGTTACAAGGATCTAAGAAGCATCAAT\n',
'GGGTCACTAACATCTAGTGGTACTAATTAACTCTGTTAATCATTGGGAAG\n',
'AAAATGTATATATACTTTTGTCTTGGAGCTGATTCTACTAGAAAGCAGAA\n',
'ATCAAAATGATCAGTTTCCCAGTGTCACTACTGCACACCCTGGAACAGAA\n',
'CAGGTAGGTCAGAAAAACGCTCCCAAAGTTTAGCAATGTCAAGGCAATCT\n',
'CTCTCTTCTTACATTTCCCTTCAACCTTCTATCTCCTCCACTTTTCTGTT\n',
'TTCCTCCTATCTCCAATTATTTCAATCCTCAGAGCATTATTCTTACAATC\n',
'TTAATCACTAAATTATATTACACCCGTTAAAGGAGAGATTTCTAAATGCA\n',
'TTGACATTTGTACTGTCTCTCTTTGGAGAATTAGTATTATAAGGATCTGT\n',
'TATCTCTTGTCACCTTCCTTATGTCATATGATATGTCACATTTCCCACTG\n',
'CGGAGACCAAACATGTTCACATCGTGTGCGTTCCATTTTCCTAATGGAAA\n',
'GTGGGGGGAAGTGATTTTCTGTCCTCATATAGAGAATGCTGGGGCCATTC\n',
'CCTCTGTATGCCATATTTGATAAAGCATTTGATAATCTTAGTCAATGCCT\n',
'GGGCCAAGAATTAAAGGGGTAATTATCAGAATGAAAATGGTTTAATGAAA\n',
'CTGTGTCTATCAGTTCTGAAAAGGGCCTCTATCACAATGAACTAAGGTAG\n',
'TTATGAATAGAGCTAAaacttaggcaacaccatcctggacataggaacgg\n',
'gcaaagatttcatgacaaagacacggaaaccaatcacaacaaaagcaaaa\n',
'attgagaagtggaatctaataaaacaatagcttctgcacagcaaaagaag\n',
'ctaccaacaaagtaaacagacaacctacagaatgggagaaaatatttgcc\n',
'aactgtaagtctgacaaaaatctaatatctggcagctataaggaacttaa\n',
'atttacaagacaaaaacaaccccattaaaaagtgggcaaagaacatgaat\n',
'agacactctcaaaagaagatatacatatggttaacaagcatatgaaaaaa\n',
'aagctcaatatactgagcattagagaaatgcaaatcaaaaccatattgag\n',
'atatcatctcataccaggcagaatggctattattaaaaagtcaaaaataa\n',
'cagatatcggtgaggttacagagaaaagggaacacttatacactgttggt\n',
'gggactgtaaattatttcaaccattgtggaaagcagtatgggatggcgat\n',
'tcctcaaaaagccaaaaacagaactatcattcaacccagcaattccatta\n',
'ctgggtatatacccagaagaatataaatcgttctaccataaagacgcatg\n',
'catgagaatgttcattgcagcactactcacaatagcagagacatggaatc\n',
'aacttaaatgcccatcagtaacagactggataaagaaagtgtggtacaga\n',
'tacaccgtggattactatgcagccataaaaaagaacaagatcatgtcttt\n',
'gacaggaacatggatggagctggaggctactatccttagcaagctaaggc\n',
'aggaacagaaatccaaataccgcatgttctcacttatgagcgtgagataa\n',
'atgatgagaacttgtaaacacaaagaaggaaacaacaggcagtggggtct\n',
'acttgaggacgacgggaagagggagaggagcagaaaagataactactgac\n',
'taccgggcgctacctgggggatgaaacaatctgtacaacgaacccccagg\n',
'acatgagtttacctatgtaacaaaccttcacgtgtacccccgaacctaaa\n',
'ataaaagtcaaaaagaaaAAGAAAAAAAGAAAAATCCATGCATATGATAC\n',
'ATCAGTTAACAAGGCACTGGTGAAATTAATTTTAAGTATTATTGTCTCTT\n',
'TGTGTTTTTGGTCTCAGAAAAGTTACGATTTCCCTTAGTTCCTTAGGGCA\n',
'GAGAGAATCTTCAATCACTGAAGTCAGGAGACACACATTCTATCTGATTT\n',
'TCTACATTATCTGTTTGAAAAGGTTACCCACTTATTAGTGTTAAAGCCAA\n',
'GATATCCAGCAAGGATAGCAACCAACTCTTAAGGTACTCTCCCTTAGGAG\n',
'GATTCCTGATTCTTTAATGTTTTCTaaaaaagcaaaacaaacaaacaaac\n',
'aaaacaaaacaCTAAATGTTTTCTCTTTCAACTTATTTGAATACACTCTT\n',
'TTCTCACTGCTCTGAGCATGAATTCAATATTTCAGGGCAAACTAACTGAA\n',
'TGTTAGAACCAACTCCTGATAAGTCTTGAACAAAAGATAGGATCCTCTAT\n',
]
from numpy.random import choice
import numpy as np
import time
import tracemalloc
import gzip
def consume_cpu_ram(n): return np.ones((n, n))
def consume_cpu_ram_128mb(): return consume_cpu_ram(2**12)
z = [consume_cpu_ram_128mb() for i in range(8)] # 1GB
del z
tracemalloc.start()
a1 = consume_cpu_ram_128mb()
a2 = consume_cpu_ram_128mb()
del a1
start = time.time()
sequence = open('/content/hg38.fa','r').read()
base_tallies = []
base_tallies.append(sequence.count('a'))
base_tallies.append(sequence.count('g'))
base_tallies.append(sequence.count('c'))
base_tallies.append(sequence.count('t'))
base_tallies = np.array(base_tallies).astype(float)
sequence_length = float(len(sequence))
probabilities = base_tallies /sequence_length
print(f'Sum of probs: {np.sum(probabilities)}')
print(f'Probs : {probabilities}')
probabilities = probabilities/np.sum(probabilities)
print(f'Prob A: {probabilities[0]:0.3f}')
print(f'Prob G: {probabilities[1]:0.3f}')
print(f'Prob C: {probabilities[2]:0.3f}')
print(f'Prob T: {probabilities[3]:0.3f}')
bases = ['a','g','c','t']
output_sequence_zero = []
for i in range(100):
output_sequence_zero.append(choice(bases,p=probabilities))
log2_probability_zero = 0
for base in sequence:
base_index = bases.index(base)
log2_probability_zero += np.log2(probabilities[base_index])
print(f'Log2Prob for Zeroth Order: {log2_probability_zero}')
transition_probabilities = np.zeros(shape=(4,4))
base_patterns = np.array([
['aa','at','ac','ag'],
['ta','tt','tc','tg'],
['ca','ct','cc','cg'],
['ga','gt','gc','gg']
])
print('First Order Count')
for i in range(len(sequence) - 1):
if (sequence[i] + sequence[i+1]) in base_patterns:
x = np.where(base_patterns==(sequence[i]+sequence[i+1]))[0][0]
y = np.where(base_patterns==(sequence[i]+sequence[i+1]))[1][0]
transition_probabilities[x][y] += 1
print(transition_probabilities)
for row in transition_probabilities:
row /= np.sum(row)
print('Probability 1st')
print(transition_probabilities)
print(np.sum(transition_probabilities))
sequence = sequence.casefold()
print(f'Length of sequence:{len(sequence)}')
string='attaaaggtttataccttcccaggtaacaaaccaaccaactttcgatctcttgtagatct'
print(len(string))
countA=0
countT=0
countG=0
countC=0
for i in range(len(char1)):
if char1[i] =='a':
countA += 1
elif char1[i] =='t':
countT += 1
elif char1[i] =='g':
countG += 1
else :
countC += 1
print("number of a's in string = ",countA)
print("number of t's in string = ",countT)
print("number of g's in string = ",countG)
print("number of c's in string = ",countC)
char1='attaaaggtttataccttcccaggtaacaaaccaaccaactttcgatctcttgtagatct'
len(char1)
covid_dna = """
1 attaaaggtt tataccttcc caggtaacaa accaaccaac tttcgatctc ttgtagatct
61 gttctctaaa cgaactttaa aatctgtgtg gctgtcactc ggctgcatgc ttagtgcact
121 cacgcagtat aattaataac taattactgt cgttgacagg acacgagtaa ctcgtctatc
181 ttctgcaggc tgcttacggt ttcgtccgtg ttgcagccga tcatcagcac atctaggttt
241 cgtccgggtg tgaccgaaag gtaagatgga gagccttgtc cctggtttca acgagaaaac
301 acacgtccaa ctcagtttgc ctgttttaca ggttcgcgac gtgctcgtac gtggctttgg
361 agactccgtg gaggaggtct tatcagaggc acgtcaacat cttaaagatg gcacttgtgg
421 cttagtagaa gttgaaaaag gcgttttgcc tcaacttgaa cagccctatg tgttcatcaa
481 acgttcggat gctcgaactg cacctcatgg tcatgttatg gttgagctgg tagcagaact
541 cgaaggcatt cagtacggtc gtagtggtga gacacttggt gtccttgtcc ctcatgtggg
601 cgaaatacca gtggcttacc gcaaggttct tcttcgtaag aacggtaata aaggagctgg
base_patterns = np.array([
['aa','at','ac','ag'],
['ta','tt','tc','tg'],
['ca','ct','cc','cg'],
['ga','gt','gc','gg']
])
count_A=0
count_T=0
count_G=0
count_C=0
count_TA=0
count_AT=0
for i in range(len(char1)):
if char1[i] =='a':
count_A += 1
elif char1[i] =='t':
count_T += 1
elif char1[i] =='g':
count_G += 1
elif char1[i]=='c' :
count_C += 1
elif char1[i]=='a' and char1[i+1]=='t':
count_AT += 1
elif char1[i]+char1[i+1]=='ta':
count_TA += 1
print("number of a's in string = ",count_A)
print("number of t's in string = ",count_T)
print("number of g's in string = ",count_G)
print("number of c's in string = ",count_C)
print("number of at's in string = ",count_AT)
print("number of ta's in string = ",count_TA)
lis1=['aa','at','ag','ac','tt','ta','tg','tc','gg','ga','gt','gc','cc','ca','ct','cg']
lis2=
lis1[0]
lis1[8]
count_AA=0
for i in range(len(char1)):
conca=char1[i]+char1[i+1]
print(conca)
#if lis1[0] == conca:
# count_AA += 1
#print(count_AA)
for i in range(len(char1)):
r=i+1%2
j=r+1
print(j)
|
```
| github_jupyter |
[Sascha Spors](https://orcid.org/0000-0001-7225-9992),
Professorship Signal Theory and Digital Signal Processing,
[Institute of Communications Engineering (INT)](https://www.int.uni-rostock.de/),
Faculty of Computer Science and Electrical Engineering (IEF),
[University of Rostock, Germany](https://www.uni-rostock.de/en/)
# Tutorial Signals and Systems (Signal- und Systemtheorie)
Summer Semester 2021 (Bachelor Course #24015)
- lecture: https://github.com/spatialaudio/signals-and-systems-lecture
- tutorial: https://github.com/spatialaudio/signals-and-systems-exercises
WIP...
The project is currently under heavy development while adding new material for the summer semester 2021
Feel free to contact lecturer [frank.schultz@uni-rostock.de](https://orcid.org/0000-0002-3010-0294)
## Übung / Exercise 4
# Spectrum from Rect Cosine
```
import numpy as np
import matplotlib.pyplot as plt
def my_sinc(x): # we rather use definition sinc(x) = sin(x)/x, thus:
return np.sinc(x/np.pi)
def plot_rect_cos(T):
# rect(t/T) o-o T sinc(wT/2)
t = np.linspace(-T/2, +T/2, 2**11)
w = np.linspace(-6, 6, 2**11)
A1 = 1
w1 = 1
A2 = 3/2
w2 = 3/2
A3 = 2
w3 = 2
x = A1*np.cos(w1*t) + A2*np.cos(w2*t) + A3*np.cos(w3*t)
X1p = 1/(2*np.pi) * (A1*np.pi) * T*my_sinc((w-w1)*T/2)
X1n = 1/(2*np.pi) * (A1*np.pi) * T*my_sinc((w+w1)*T/2)
X2p = 1/(2*np.pi) * (A2*np.pi) * T*my_sinc((w-w2)*T/2)
X2n = 1/(2*np.pi) * (A2*np.pi) * T*my_sinc((w+w2)*T/2)
X3p = 1/(2*np.pi) * (A3*np.pi) * T*my_sinc((w-w3)*T/2)
X3n = 1/(2*np.pi) * (A3*np.pi) * T*my_sinc((w+w3)*T/2)
plt.figure(figsize=(10, 10))
plt.subplot(3, 1, 1)
plt.plot(t, x, 'C0', lw=3)
plt.plot([-np.ceil(T/2), -T/2], [0, 0], 'C0', lw=3)
plt.plot([T/2, np.ceil(T/2)], [0, 0], 'C0', lw=3)
plt.xlabel(r'$t$ / s')
plt.ylabel('$x(t)$')
plt.xlim(-np.ceil(T/2), +np.ceil(T/2))
plt.grid(True)
plt.subplot(3, 1, 2)
head_length = 0.25
plt.arrow(+w1, 0, 0, A1*np.pi-head_length, color='C0',
width=0.05, head_length=head_length)
plt.arrow(+w2, 0, 0, A2*np.pi-head_length, color='C1',
width=0.05, head_length=head_length)
plt.arrow(+w3, 0, 0, A3*np.pi-head_length, color='C2',
width=0.05, head_length=head_length)
plt.arrow(-w1, 0, 0, A1*np.pi-head_length, color='C3',
width=0.05, head_length=head_length)
plt.arrow(-w2, 0, 0, A2*np.pi-head_length, color='C4',
width=0.05, head_length=head_length)
plt.arrow(-w3, 0, 0, A3*np.pi-head_length, color='C5',
width=0.05, head_length=head_length)
plt.plot(w, X1p+X1n + X2p+X2n + X3p+X3n, 'k')
plt.xlabel(r'$\omega$ / (rad/s)')
plt.ylabel('$X(\mathrm{j}\omega)$')
plt.xticks(np.arange(-6, 7))
plt.xlim(-6, 6)
plt.grid(True)
plt.subplot(3, 1, 3)
plt.plot(
w, X1p, label=r'$\frac{A_1 \pi}{2\pi}\delta(\omega-\omega_1)\ast T \mathrm{sinc}(\frac{\omega T}{2})$')
plt.plot(
w, X2p, label=r'$\frac{A_2 \pi}{2\pi}\delta(\omega-\omega_2)\ast T \mathrm{sinc}(\frac{\omega T}{2})$')
plt.plot(
w, X3p, label=r'$\frac{A_3 \pi}{2\pi}\delta(\omega-\omega_3)\ast T \mathrm{sinc}(\frac{\omega T}{2})$')
plt.plot(
w, X1n, label=r'$\frac{A_1 \pi}{2\pi}\delta(\omega+\omega_1)\ast T \mathrm{sinc}(\frac{\omega T}{2})$')
plt.plot(
w, X2n, label=r'$\frac{A_2 \pi}{2\pi}\delta(\omega+\omega_2)\ast T \mathrm{sinc}(\frac{\omega T}{2})$')
plt.plot(
w, X3n, label=r'$\frac{A_3 \pi}{2\pi}\delta(\omega+\omega_3)\ast T \mathrm{sinc}(\frac{\omega T}{2})$')
plt.xlabel(r'$\omega$ / (rad/s)')
plt.xticks(np.arange(-6, 7))
plt.xlim(-6, 6)
plt.legend()
plt.grid(True)
T = 2*np.pi*20
plot_rect_cos(T)
T = 2*np.pi*10
plot_rect_cos(T)
T = 2*np.pi*2
plot_rect_cos(T)
T = 2*np.pi*1.5
plot_rect_cos(T)
T = 2*np.pi*1
plot_rect_cos(T)
A = 1
T0 = 1
w0 = 2*np.pi/T0
Th = 4.8/1.5*T0
Th = 3.2
print(Th)
# rect(t/T) o-o T sinc(wT/2)
t = np.linspace(-Th/2, +Th/2, 2**11)
w = np.linspace(-2*w0, +2*w0, 2**11)
x = A*np.cos(w0*t)
X1p = 1/(2*np.pi) * (A*np.pi) * Th*my_sinc((w-w0)*Th/2)
X1n = 1/(2*np.pi) * (A*np.pi) * Th*my_sinc((w+w0)*Th/2)
plt.figure(figsize=(10, 10))
plt.subplot(3, 1, 1)
plt.plot(t, x, 'C2', lw=3,
label=r'$\mathrm{rect}(\frac{t}{T_h})\cdot A \cos(\omega_0 t)$')
plt.plot([-np.ceil(Th/2), -Th/2], [0, 0], 'C2', lw=3)
plt.plot([Th/2, np.ceil(Th/2)], [0, 0], 'C2', lw=3)
plt.xlabel(r'$t$ / s')
plt.ylabel('$x(t)$')
plt.title(
r'$A$=%1.1f, $T_0=$%1.1f s$\rightarrow\omega_0 = \frac{2\pi}{T_0}\approx$%2.2f rad/s, $T_h$=%1.1f s' % (A, T0, w0, Th))
plt.xlim(-np.ceil(Th/2), +np.ceil(Th/2))
plt.legend(loc='upper right')
plt.grid(True)
plt.subplot(3, 1, 2)
head_length = 0.25
plt.plot([w0, w0], [0, 1], 'C0', label=r'$\pi\delta(\omega-\omega_0)$')
plt.plot([-w0, -w0], [0, 1], 'C1', label=r'$\pi\delta(\omega+\omega_0)$')
plt.arrow(+w0, 0, 0, A*np.pi-head_length, color='C0',
width=0.05, head_length=head_length)
plt.arrow(-w0, 0, 0, A*np.pi-head_length, color='C1',
width=0.05, head_length=head_length)
plt.plot(w, X1p+X1n, 'k')
plt.text(-w0, np.pi, r'$(\pi)$', color='C1')
plt.text(+w0, np.pi, r'$(\pi)$', color='C0')
plt.xlabel(r'$\omega$ / (rad/s)')
plt.ylabel('$X(\mathrm{j}\omega)$')
if T0 == 1:
plt.xticks(np.arange(-12, 14, 2))
plt.xlim(-12, 12)
if Th == 3.2 and A == 1:
plt.ylim(-1, 4)
plt.legend()
plt.grid(True)
plt.subplot(3, 1, 3)
plt.plot(
w, X1p, label=r'$\frac{1}{2\pi}\pi\delta(\omega-\omega_0)\ast A T_h \mathrm{sinc}(\frac{\omega T_h}{2})$')
plt.plot(
w, X1n, label=r'$\frac{1}{2\pi}\pi\delta(\omega+\omega_0)\ast A T_h \mathrm{sinc}(\frac{\omega T_h}{2})$')
plt.xlabel(r'$\omega$ / (rad/s)')
if T0 == 1:
plt.xticks(np.arange(-12, 14, 2))
plt.xlim(-12, 12)
if Th == 3.2 and A == 1:
plt.yticks(np.arange(-0.4, 2.4, 0.4))
plt.ylim(-0.4, 2)
plt.legend()
plt.grid(True)
plt.savefig('rect_cos_610482EF57_1.pdf')
A = 1
T0 = 1
w0 = 2*np.pi/T0
Th = 4.8/1.5*T0
Th = 3.2
print(Th)
# rect(t/T) o-o T sinc(wT/2)
w = np.linspace(-2*w0, +2*w0, 2**11)
Xrect = A*Th * my_sinc(w*Th/2)
Xmodp = A*Th/2*my_sinc((w-w0)*Th/2)
Xmodn = A*Th/2*my_sinc((w+w0)*Th/2)
plt.figure(figsize=(10, 10*10/16))
plt.plot(
w, Xrect, 'C0', lw=0.5)
plt.plot(
w, Xrect, 'C0:', label=r'$X_1(\mathrm{j}\omega) = A T_h \mathrm{sinc}(\frac{\omega T_h}{2})$')
plt.plot(
w, Xmodp+Xmodn, 'C2', lw=3, label=r'$X(\mathrm{j}\omega) = \frac{A T_h}{2} \mathrm{sinc}(\frac{[\omega-\omega_0] T_h}{2})+\frac{A T_h}{2} \mathrm{sinc}(\frac{[\omega+\omega_0] T_h}{2})$')
plt.plot(
w, Xmodp, 'C1--', lw=2, label=r'$\frac{A T_h}{2} \mathrm{sinc}(\frac{[\omega-\omega_0] T_h}{2})$')
plt.plot(
w, Xmodn, 'C3-.', lw=2, label=r'$\frac{A T_h}{2} \mathrm{sinc}(\frac{[\omega+\omega_0] T_h}{2})$')
plt.xlabel(r'$\omega$ / (rad/s)')
plt.title(
r'$A$=%1.1f, $T_0=$%1.1f s$\rightarrow\omega_0 = \frac{2\pi}{T_0}\approx$%2.2f rad/s, $T_h$=%1.1f s' % (A, T0, w0, Th))
plt.xticks(np.arange(-12, 14, 2))
plt.xlim(-12, 12)
plt.yticks(np.arange(-0.8, 4, 0.4))
plt.ylim(-0.8, 3.6)
plt.legend()
plt.grid(True)
plt.savefig('rect_cos_610482EF57_2.pdf')
```
## Copyright
This tutorial is provided as Open Educational Resource (OER), to be found at
https://github.com/spatialaudio/signals-and-systems-exercises
accompanying the OER lecture
https://github.com/spatialaudio/signals-and-systems-lecture.
Both are licensed under a) the Creative Commons Attribution 4.0 International
License for text and graphics and b) the MIT License for source code.
Please attribute material from the tutorial as *Frank Schultz,
Continuous- and Discrete-Time Signals and Systems - A Tutorial Featuring
Computational Examples, University of Rostock* with
``main file, github URL, commit number and/or version tag, year``.
| github_jupyter |
# Ideal rocket analysis
```
# this line makes figures interactive in Jupyter notebooks
%matplotlib inline
from matplotlib import pyplot as plt
import numpy as np
# we can use this to solve nonlinear/transcendental equations
from scipy.optimize import root_scalar
# this provides access to many physical constants
from scipy import constants
# provides the 1976 US Standard Atmosphere model
from fluids.atmosphere import ATMOSPHERE_1976
# Module used to parse and work with units
from pint import UnitRegistry
ureg = UnitRegistry()
Q_ = ureg.Quantity
# for convenience:
def to_si(quant):
'''Converts a Pint Quantity to magnitude at base SI units.
'''
return quant.to_base_units().magnitude
# these lines are only for helping improve the display
from IPython.display import set_matplotlib_formats
set_matplotlib_formats('pdf', 'png')
plt.rcParams['figure.dpi']= 150
plt.rcParams['savefig.dpi'] = 150
```
## Characteristic velocity: c*
$$
c^* = \frac{p_0 A_t}{\dot{m}} = \sqrt{\frac{R T_0}{\gamma} \left( \frac{\gamma + 1}{2} \right)^{\frac{\gamma+1}{\gamma-1}} } \;,
$$
where $p_0$ and $T_0$ are the combustion chamber pressure and temperature,
$R$ is the specific gas constant ($\mathcal{R}_u / MW$),
and $\gamma$ is the specific heat ratio.
## Thrust coefficient
$$
\begin{align*}
C_F &= \frac{T}{p_0 A_t} = \sqrt{\frac{2\gamma^2}{\gamma-1} \left( \frac{2}{\gamma+1} \right)^{\frac{\gamma+1}{\gamma-1}} \left[1 - \left( \frac{p_e}{p_0} \right)^{\frac{\gamma-1}{\gamma}} \right] } + \frac{A_e}{A_t} \frac{p_e - p_a}{p_0} \\
&= C_F^0 + \Delta C_F \;,
\end{align*}
$$
where $T$ is thrust, $p_e$ is the nozzle exit pressure,
$A_e$ and $A_t$ are the nozzle exit and throat areas, and $p_a$ is the ambient pressure.
## Relationships with thrust, specific impulse, and effective exhaust velocity
Using characteristic velocity ($c^*$) and thrust coefficient ($C_F$),
we can express thrust, specific impulse ($I_{\text{sp}}$),
and effective exhause velocity ($c$):
$$
\begin{align*}
T &= \dot{m} c^* C_F \\
I_{\text{sp}} &= \frac{c^* C_F}{g_0} \\
c &= c^* C_F \;.
\end{align*}
$$
## Area ratio
The nozzle area ratio (exit area to throat area) can be determined directly
from the nozzle pressure ratio:
$$
\frac{A_e}{A_t} = \epsilon = \left( \frac{2}{\gamma+1}\right)^{\frac{1}{\gamma-1}} \left( \frac{p_0}{p_e} \right)^{\frac{1}{\gamma}} \sqrt{ \frac{\gamma-1}{\gamma+1} \left[1 - \left( \frac{p_0}{p_e}\right)^{\frac{1-\gamma}{\gamma}} \right]^{-1} } \;,
$$
where $\frac{p_0}{p_e}$ is the pressure ratio (chamber pressure to nozzle exit pressure).
## Designing rocket nozzles
Rocket scientists (... rocket engineers) have used the above equations for thrust
coefficient, area ratio, and Mach number to design rockets for many years.
For example, the following figures show area ratio vs. pressure ratio, and
thrust coefficient (at optimum expansion conditions, where $p_e = p_a$)
vs. pressure ratio, both for various specific heat ratios:
```
## area ratio as a function of pressure ratio
gammas = [1.1, 1.25, 1.4, 1.7]
pressure_ratios = np.logspace(1, 4, num=50)
labels = [r'$\gamma = 1.1$', '1.25', '1.4', '1.7']
# let's define a function to calculate area ratio based on gamma and the pressure ratio:
def calc_area_ratio(gamma, pressure_ratio):
'''Calculates area ratio based on specific heat ratio and pressure ratio.
pressure ratio: chamber / exit
area ratio: exit / throat
'''
return (
np.power(2 / (gamma + 1), 1/(gamma-1)) *
np.power(pressure_ratio, 1 / gamma) *
np.sqrt((gamma - 1) / (gamma + 1) /
(1 - np.power(pressure_ratio, (1 - gamma)/gamma))
)
)
for gamma, label in zip(gammas, labels):
area_ratios = calc_area_ratio(gamma, pressure_ratios)
plt.plot(pressure_ratios, area_ratios)
plt.text(
0.9*pressure_ratios[-10], 1.01*area_ratios[-10],
label,
horizontalalignment='right', fontsize=8
)
plt.xlim([10, 1e4])
plt.ylim([1, 500])
plt.xlabel(r'Pressure ratio, $\frac{p_0}{p_e}$')
plt.ylabel(r'Area ratio, $\epsilon = \frac{A_e}{A_t}$')
plt.grid(True)
plt.xscale('log')
plt.yscale('log')
plt.title('Area ratio vs. pressure ratio')
plt.show()
# Define a function to calculate thrust coefficient, assuming optimum expansion
# (exit pressure = ambient pressure), based on gamma and pressure ratio:
def calc_thrust_coeff(gamma, pressure_ratio):
''' Calculates thrust coefficient for optimum expansion.
pressure ratio: chamber / exit
area ratio: exit / throat
'''
return np.sqrt(
2 * np.power(gamma, 2) / (gamma - 1) *
np.power(2 / (gamma + 1), (gamma + 1)/(gamma - 1)) *
(1 - np.power(1.0 / pressure_ratio, (gamma - 1)/gamma))
)
# This function returns zero for a given area ratio, pressure ratio, and gamma,
#and is used to numerically calculate pressure ratio given the other two values.
def root_area_ratio(pressure_ratio, gamma, area_ratio):
''' pressure ratio: chamber / exit
area ratio: exit / throat
'''
return area_ratio - calc_area_ratio(gamma, pressure_ratio)
gammas = [1.1, 1.2, 1.3, 1.4]
labels = [r'$\gamma = 1.1$', '1.2', '1.3', '1.4']
area_ratios = [3, 5, 10, 20, 50, 100]
pressure_ratios = np.logspace(1, 4, num=50)
for gamma, label in zip(gammas, labels):
thrust_coeffs = calc_thrust_coeff(gamma, pressure_ratios)
plt.plot(pressure_ratios, thrust_coeffs)
plt.text(
0.9*pressure_ratios[-1], 1.01*thrust_coeffs[-1],
label, horizontalalignment='right'
)
for area_ratio in area_ratios:
pressure_ratios2 = np.zeros(len(gammas))
thrust_coeffs2 = np.zeros(len(gammas))
for idx, gamma in enumerate(gammas):
sol = root_scalar(root_area_ratio, x0=20, x1=100, args=(gamma, area_ratio))
pressure_ratios2[idx] = sol.root
thrust_coeffs2[idx] = calc_thrust_coeff(gamma, sol.root)
plt.plot(pressure_ratios2, thrust_coeffs2, '--')
plt.text(
pressure_ratios2[-1], thrust_coeffs2[-1],
r'$\epsilon =$' + f'{area_ratio}',
horizontalalignment='left', verticalalignment='top', fontsize=8
)
plt.xlim([10, 1e4])
plt.ylim([1.2, 2.3])
plt.xlabel(r'Pressure ratio, $p_0/p_e$')
plt.ylabel(r'Thrust coefficient, $C_F$')
plt.grid(True)
plt.xscale('log')
plt.title('Thrust coefficient vs. pressure ratio, at optimum expansion')
plt.show()
```
## Example: using equations to design optimal rocket nozzle
Using the above equations, design a rocket nozzle optimally for these conditions:
$p_c$ = 70 atm, $p_e$ = 1 atm, and $\gamma$ = 1.2.
Find the nozzle area ratio, and the rocket thrust as a function of nozzle exit area.
We know the pressure ratio:
$$
\frac{p_c}{p_e} = 70 \;,
$$
so we can directly calculate the nozzle area ratio:
$$
\frac{A_e}{A_t} = \epsilon = \left( \frac{2}{\gamma+1}\right)^{\frac{1}{\gamma-1}} \left( \frac{p_0}{p_e} \right)^{\frac{1}{\gamma}} \sqrt{ \frac{\gamma-1}{\gamma+1} \left[1 - \left( \frac{p_0}{p_e}\right)^{\frac{1-\gamma}{\gamma}} \right]^{-1} } \;.
$$ (area_ratio)
```
# set the given constants
chamber_pressure = Q_(70, 'atm')
exit_pressure = Q_(1, 'atm')
gamma = 1.2
pressure_ratio = chamber_pressure / exit_pressure
area_ratio = (
np.power(2 / (gamma + 1), 1/(gamma-1)) *
np.power(pressure_ratio, 1 / gamma) *
np.sqrt((gamma - 1) / (gamma + 1) /
(1 - np.power(pressure_ratio, (1 - gamma)/gamma))
)
)
print(f'Nozzle area ratio: {area_ratio: 3.2f~P}')
```
Since $p_e = p_a$ by design (for an optimal nozzle), we can calculate the thrust coefficient using:
$$
C_F = C_F^0 = \sqrt{\frac{2\gamma^2}{\gamma-1} \left( \frac{2}{\gamma+1} \right)^{\frac{\gamma+1}{\gamma-1}} \left[1 - \left( \frac{p_e}{p_0} \right)^{\frac{\gamma-1}{\gamma}} \right] }
$$ (thrust_coefficient)
and then thrust using:
$$
T = C_F^0 p_c A_e \frac{A_t}{A_e} \;.
$$
```
thrust_coeff = np.sqrt(
2 * np.power(gamma, 2) / (gamma - 1) *
np.power(2 / (gamma + 1), (gamma + 1)/(gamma - 1)) *
(1 - np.power(1.0 / pressure_ratio, (gamma - 1)/gamma))
)
print(f'Thrust coefficient = {thrust_coeff: 3.2f~P}')
thrust_per_area = thrust_coeff * chamber_pressure / area_ratio
print(f'Thrust = ({thrust_per_area.to("kPa"): 5.1f~P} * A_e)')
```
We can now examine the thrust for a range of exit areas:
```
exit_areas = Q_(np.linspace(0.1, 5, num=50), 'm^2')
plt.plot(exit_areas.to('m^2').magnitude, (thrust_per_area * exit_areas).to('kN').magnitude)
plt.xlabel('Nozzle exit area ' + r'(m$^2$)')
plt.ylabel('Thrust (kN)')
plt.grid(True)
plt.show()
```
## Example for design of a rocket
```
# given constants
altitude = Q_(10000, 'm')
c_star = Q_(1500, 'm/s')
gamma = 1.2
MW = Q_(25, 'kg/kmol')
chamber_pressure = Q_(70, 'bar')
thrust_10k = Q_(100000, 'N')
burn_time = Q_(5, 's')
```
First, we need to calculate the nozzle pressure ratio, which requires obtaining the pressure at the rocket's flight altitude.
The 1976 U.S. Standard Atmosphere {cite}`standard_atmosphere_1976` is a model for how pressure, temperature, density, etc., vary
with altitude in the atmosphere, and provides reasonable answers for up to about 86 km.
The [`fluids` package](https://github.com/CalebBell/fluids) provides a convenient interface to this model in Python {cite}`fluids`,
along with other models such as the NRLMSISE-00 model {cite}`Picone2002`, which applies up to 1000 km.
Let's look at how temperature and pressure vary with altitude, using this model:
```
altitudes = np.linspace(0, 86000, num=100)
temperatures = np.zeros(len(altitudes))
pressures = np.zeros(len(altitudes))
for idx, h in enumerate(altitudes):
atm = ATMOSPHERE_1976(h)
temperatures[idx] = atm.T
pressures[idx] = atm.P
fig, axes = plt.subplots(1, 2)
axes[0].plot(temperatures, altitudes/1000)
axes[0].set_xlabel('Temperature (K)')
axes[0].set_ylabel('Altitude (km)')
axes[0].grid(True)
axes[1].plot(pressures/1000, altitudes/1000)
axes[1].set_xlabel('Pressure (kPa)')
axes[1].grid(True)
plt.show()
```
We can use this to obtain the pressure at the flight altitude, and calculate pressure ratio:
$$
\frac{p_c}{p_e}
$$
```
# use model for 1976 US Standard Atmosphere to get pressure at altitude
ten_km = ATMOSPHERE_1976(to_si(altitude))
exit_pressure = Q_(ten_km.P, 'Pa')
print(f'Pressure at {altitude.to("km")}: {exit_pressure: .0f}')
pressure_ratio = chamber_pressure / exit_pressure
print(f'Pressure ratio (p_c/p_e): {pressure_ratio.to_base_units(): .1f}')
```
Next, we can calculate the nozzle area ratio and thrust coefficient (for optimum expansion, since $p_e = p_a$):
$$
\begin{align*}
\frac{A_e}{A_t} &= f \left( \gamma, \frac{p_c}{p_e} \right) \\
C_F &= C_F^0 = g \left( \gamma, \frac{p_c}{p_e} \right) \\
\end{align*}
$$
which are defined in Equations {eq}`area_ratio` and {eq}`thrust_coefficient` above—and we have already written functions to evaluate them!.
```
area_ratio = calc_area_ratio(gamma, pressure_ratio)
print(f'Area ratio: {area_ratio.to_base_units(): .2f~P}')
thrust_coeff = calc_thrust_coeff(gamma, pressure_ratio)
print(f'Thrust coefficient: {thrust_coeff.to_base_units(): .2f~P}')
throat_area = thrust_10k / (thrust_coeff * chamber_pressure)
print(f'Throat area: {throat_area.to("m^2"): .4f~P}')
throat_diameter = 2 * np.sqrt(throat_area / np.pi)
print(f'Throat diameter: {throat_diameter.to("m"): .3f~P}')
exit_area = area_ratio * throat_area
print(f'Exit area: {exit_area.to("m^2"): .2e~P}')
exit_diameter = 2 * np.sqrt(exit_area / np.pi)
print(f'Exit diameter: {exit_diameter.to("m"): .3f~P}')
mass_flow_rate = chamber_pressure * throat_area / c_star
print(f'Mass flow rate: {mass_flow_rate.to("kg/s"): .2f~P}')
specific_impulse = c_star * thrust_coeff / Q_(constants.g, 'm/s^2')
print(f'Specific impulse: {specific_impulse.to("s"): .1f~P}')
propellant_mass = mass_flow_rate * burn_time
print(f'Propellant mass: {propellant_mass.to("kg"): .1f~P}')
# constant.R is given in J/(K mol), need J/(K kmol)
chamber_temperature = (
c_star**2 * gamma * (MW / Q_(constants.R, 'J/(K*mol)')) *
np.power(2 / (gamma+1), (gamma+1)/(gamma-1))
)
print(f'Chamber temperature: {chamber_temperature.to("K"): .1f~P}')
# Thrust at sea level
thrust_coeff_sea_level = thrust_coeff + area_ratio * (
1/pressure_ratio - Q_(1, 'atm')/chamber_pressure
)
thrust_sea_level = thrust_coeff_sea_level * chamber_pressure * throat_area
print(f'Thrust at sea level: {thrust_sea_level.to("kN"): .1f~P}')
```
| github_jupyter |
# Solving Image Recognition problem with better Deep Learning architectures: CNN for Image Recognition
```
# import modules
%pylab inline
import os
import tflearn
import numpy as np
import pandas as pd
import tensorflow as tf
from scipy.misc import imread
from tflearn.layers.estimator import regression
from tflearn.layers.conv import conv_2d, max_pool_2d
from tflearn.layers.core import input_data, dropout, fully_connected
## create dataset
# give path of files
data_dir = 'datasets\mnist'
# load dataset
train = pd.read_csv(os.path.join(data_dir, 'Train', 'train.csv'))
test = pd.read_csv(os.path.join(data_dir, 'Test.csv'))
# load whole dataset
temp = []
for img_name in train.filename:
image_path = os.path.join(data_dir, 'Train', 'Images', 'train', img_name)
img = imread(image_path, mode='RGB')
img = img.astype('float32')
temp.append(img)
train_x = np.stack(temp)
temp = []
for img_name in test.filename:
image_path = os.path.join(data_dir, 'Train', 'Images', 'test', img_name)
img = imread(image_path, mode='RGB')
img = img.astype('float32')
temp.append(img)
test_x = np.stack(temp)
# create validation dataset
split_size = int(train_x.shape[0]*0.7)
train_x, val_x = train_x[:split_size, ...], train_x[split_size:, ...]
train_y, val_y = train.label.values[:split_size], train.label.values[split_size:]
# helper functions
def dense_to_one_hot(labels_dense, num_classes=10):
"""Convert class labels from scalars to one-hot vectors"""
num_labels = labels_dense.shape[0]
index_offset = np.arange(num_labels) * num_classes
labels_one_hot = np.zeros((num_labels, num_classes))
labels_one_hot.flat[index_offset + labels_dense.ravel()] = 1
return labels_one_hot
# one hot encode outputs
train_y = dense_to_one_hot(train_y)
val_y = dense_to_one_hot(val_y)
tf.reset_default_graph()
# Building the model architecture
# Input Layer
model = input_data(shape=[None, 28, 28, 3])
# First Convolution Layer
model = conv_2d(model, 32, 5, activation='relu')
# First Max Pooling
model = max_pool_2d(model, 5)
# Fully Connected Layer
model = fully_connected(model, 10, activation='softmax')
model = regression(model, optimizer='adam', loss='categorical_crossentropy',learning_rate=0.05)
model = tflearn.DNN(model)
model.fit(train_x, train_y, n_epoch=2, batch_size=128, show_metric=True)
model.evaluate(val_x, val_y)
```
## Exercise 1
Play with the hyperparameters and increase the validation accuracy
```
tf.reset_default_graph()
# Building the model architecture
# Input Layer
model = input_data(shape=[None, 28, 28, 3])
# First Convolution Layer
model = conv_2d(model, 32, 5, activation='relu')
# First Max Pooling
model = max_pool_2d(model, 5)
# Fully Connected Layer
model = fully_connected(model, 50, activation='relu')
model = fully_connected(model, 10, activation='softmax')
model = regression(model, optimizer='adam', loss='categorical_crossentropy',learning_rate=0.001)
model = tflearn.DNN(model)
model.fit(train_x, train_y, n_epoch=20, batch_size=128, show_metric=True)
model.evaluate(val_x, val_y)
```
Now similar to simple neural network, let's visualize our output
```
# # Make Prediction
pred = model.predict(test_x)
print(pred)
# To stop potential randomness
seed = 128
rng = np.random.RandomState(seed)
# get predictions
img_name = rng.choice(test.filename)
filepath = os.path.join(data_dir, 'Train', 'Images', 'test', img_name)
img = imread(filepath, flatten=True)
test_index = int(img_name.split('.')[0]) - 49000
print( "Prediction is: ", np.argmax(pred[test_index]))
pylab.imshow(img, cmap='gray')
pylab.axis('off')
pylab.show()
```
## Exercise 2
Play with CNN architecture. You can tune the following parameters
* Number of Convolutional layers
* number of Pooling layers
* Size of kernels in Convolutional layer
* Size of pooling operation in Pooling layer
* Add Regularization layer in the network (Dropout)
```
tf.reset_default_graph()
# Building the model architecture
# Input Layer
model = input_data(shape=[None, 28, 28, 3])
# First Convolution Layer
model = conv_2d(model, 32, 5, activation='relu')
# First Max Pooling
model = max_pool_2d(model, 5)
# second Convolution Layer
model = conv_2d(model, 64, 5, activation = 'relu')
# Second Max Pooling
model = max_pool_2d(model, 5)
# Fully Connected Layer
model = fully_connected(model, 50, activation='relu')
model = dropout(model, 0.9)
model = fully_connected(model, 10, activation='softmax')
model = regression(model, optimizer='adam', loss='categorical_crossentropy',learning_rate=0.001)
model = tflearn.DNN(model)
model.fit(train_x, train_y, n_epoch=100, batch_size=128, show_metric=True)
model.evaluate(val_x, val_y)
pred = model.predict(test_x)
# get predictions
img_name = rng.choice(test.filename)
filepath = os.path.join(data_dir, 'Train', 'Images', 'test', img_name)
img = imread(filepath, flatten=True)
test_index = int(img_name.split('.')[0]) - 49000
print( "Prediction is: ", np.argmax(pred[test_index]))
pylab.imshow(img, cmap='gray')
pylab.axis('off')
pylab.show()
```
## END
| github_jupyter |
```
import pandas as pd
import numpy as np
# Import as tabelas do SNIS
data_sul = pd.read_csv("AgrupamentoDesagregado--2018-05-31--11-43-29_Regiao_Sul.csv", encoding='latin1', delimiter=';', engine='python', skipfooter = 3)
data_sudeste = pd.read_csv("AgrupamentoDesagregado--2018-05-31--12-05-11_Regiao_Sudeste.csv", encoding='latin1', delimiter=';', engine='python', skipfooter = 3)
data_norte = pd.read_csv("AgrupamentoDesagregado--2018-05-31--12-24-50_Regiao_Norte.csv", encoding='latin1', delimiter=';', engine='python', skipfooter = 3)
data_nordeste = pd.read_csv("AgrupamentoDesagregado--2018-05-31--12-49-58_Regiao_Nordeste.csv", encoding='latin1', delimiter=';', engine='python', skipfooter = 3)
data_centro_oeste = pd.read_csv("AgrupamentoDesagregado--2018-05-31--12-54-02_Regiao_centro_Oeste.csv", encoding='latin1', delimiter=';', engine='python', skipfooter = 3)
# Import as tabelas do SNIS
#data_sul = pd.read_csv("AgrupamentoDesagregado--2018-05-31--11-43-29_Regiao_Sul.csv", delimiter=';', engine='python', skipfooter = 3)
#data_sudeste = pd.read_csv("AgrupamentoDesagregado--2018-05-31--12-05-11_Regiao_Sudeste.csv", delimiter=';', engine='python', skipfooter = 3)
#data_norte = pd.read_csv("AgrupamentoDesagregado--2018-05-31--12-24-50_Regiao_Norte.csv", delimiter=';', engine='python', skipfooter = 3)
#data_nordeste = pd.read_csv("AgrupamentoDesagregado--2018-05-31--12-49-58_Regiao_Nordeste.csv", delimiter=';', engine='python', skipfooter = 3)
#data_centro_oeste = pd.read_csv("AgrupamentoDesagregado--2018-05-31--12-54-02_Regiao_centro_Oeste.csv", delimiter=';', engine='python', skipfooter = 3)
df_snis = pd.concat([data_sul, data_sudeste, data_norte, data_nordeste, data_centro_oeste])
display(df_snis.head())
df_snis[['FN026 - Quantidade total de empregados próprios (Empregados)']][df_snis['Município '] == 'Campinas']
df_snis['FN026 - Quantidade total de empregados próprios (Empregados)'].dtypes
df_snis['POP_TOT - População total do município do ano de referência (Fonte: IBGE): (Habitantes)'].dtypes
df_snis.info()
len(df_snis)
# deleta linhas com municipio = ---
municipios_index_sem_id = []
# Selecione os índices dos municipios iguais a --- para remover
municipios_sem_id = df_snis[df_snis['Município '] == '---']
municipios_index_sem_id += municipios_sem_id.index.tolist()
# Remove os municipios
df_snis = df_snis.drop(municipios_index_sem_id).reset_index(drop = True)
len(df_snis)
df_snis['Código do Município '].dtypes
# muda o tipo do Código do Município para int64
df_snis['Código do Município '] = df_snis['Código do Município '].str.replace('.', '')
df_snis['Código do Município '] = pd.to_numeric(df_snis['Código do Município '])
df_snis['Código do Município '].dtypes
df_snis['POP_TOT - População total do município do ano de referência (Fonte: IBGE): (Habitantes)'].dtypes
#Alterando virgulas por pontos e transformando coluna em inteiro
df_snis['POP_TOT - População total do município do ano de referência (Fonte: IBGE): (Habitantes)'] = df_snis['POP_TOT - População total do município do ano de referência (Fonte: IBGE): (Habitantes)'].str.replace('.', '')
df_snis['POP_TOT - População total do município do ano de referência (Fonte: IBGE): (Habitantes)'] = pd.to_numeric(df_snis['POP_TOT - População total do município do ano de referência (Fonte: IBGE): (Habitantes)'])
df_snis['POP_TOT - População total do município do ano de referência (Fonte: IBGE): (Habitantes)'].dtypes
df_snis['POP_TOT - População total do município do ano de referência (Fonte: IBGE): (Habitantes)'].max()
df_snis[df_snis['POP_TOT - População total do município do ano de referência (Fonte: IBGE): (Habitantes)'] == 12038175]
df_snis['FN026 - Quantidade total de empregados próprios (Empregados)'][df_snis['POP_TOT - População total do município do ano de referência (Fonte: IBGE): (Habitantes)'] == 12038175]
df_snis['FN026 - Quantidade total de empregados próprios (Empregados)'].dtypes
# limpar os residuos decimais acima de 3 casas para obter o inteiro corretamente
df_snis['FN026 - Quantidade total de empregados próprios (Empregados)'] = round(df_snis['FN026 - Quantidade total de empregados próprios (Empregados)'],3).astype(str)
df_snis['FN026 - Quantidade total de empregados próprios (Empregados)'].dtypes
df_snis['FN026 - Quantidade total de empregados próprios (Empregados)'][df_snis['POP_TOT - População total do município do ano de referência (Fonte: IBGE): (Habitantes)'] == 12038175]
#Alterando virgulas por pontos e transformando coluna em inteiro
df_snis['FN026 - Quantidade total de empregados próprios (Empregados)'] = df_snis['FN026 - Quantidade total de empregados próprios (Empregados)'].str.replace('.', '')
df_snis['FN026 - Quantidade total de empregados próprios (Empregados)'] = pd.to_numeric(df_snis['FN026 - Quantidade total de empregados próprios (Empregados)'])
df_snis['FN026 - Quantidade total de empregados próprios (Empregados)'].dtypes
df_snis['FN026 - Quantidade total de empregados próprios (Empregados)'][df_snis['Município '] == 'Campinas']
df_snis['FN026 - Quantidade total de empregados próprios (Empregados)'].dtypes
df_snis['FN026 - Quantidade total de empregados próprios (Empregados)'][df_snis['POP_TOT - População total do município do ano de referência (Fonte: IBGE): (Habitantes)'] == 12038175]
#Mapeando novos valores para a coluna Tipos de Despesa
df_snis['Tipo Empresa'] = df_snis['Natureza jurídica '].map({'Autarquia': 'Pública',
'Administração pública direta': 'Pública',
'Sociedade de economia mista com administração privada': 'Pública',
'Empresa privada': 'Privada',
'Sociedade de economia mista com administração pública': 'Pública',
'Organização social': 'Pública',
'Empresa pública': 'Pública'})
#Mapeando novos valores para a coluna Tipos de Despesa
df_snis['Nat Juridica'] = df_snis['Natureza jurídica '].map({'Autarquia': 'Autarquia',
'Administração pública direta': 'APD',
'Sociedade de economia mista com administração privada': 'SEMAPriv',
'Empresa privada': 'Privada',
'Sociedade de economia mista com administração pública': 'SEMAPubl',
'Organização social': 'OS',
'Empresa pública': 'Pública'})
df_snis.groupby(['Tipo Empresa']).groups.keys()
len(df_snis.groupby(['Tipo Empresa']).groups['Privada'])
len(df_snis.groupby(['Tipo Empresa']).groups['Pública'])
#Como o pandas está tratando cada coluna do dataset
df_snis['POP_TOT - População total do município do ano de referência (Fonte: IBGE): (Habitantes)'].dtypes
top100 = df_snis[['Código do Município ',# Cód Município
'Tipo Empresa', # Tp Empresa
'Nat Juridica', # Nat Jurídica
'POP_TOT - População total do município do ano de referência (Fonte: IBGE): (Habitantes)', # Pop Total
# 'IN046 - Índice de esgoto tratado referido à água consumida (percentual)', # Esg Tratado
'IN013 - Índice de perdas faturamento (percentual)', # Perdas Fat
'IN049 - Índice de perdas na distribuição (percentual)', # Perdas Dist
'IN004 - Tarifa média praticada (R$/m³)', # Tarifa méd
# 'FN033 - Investimentos totais realizados pelo prestador de serviços (R$/ano)', # Investimento
'AG005 - Extensão da rede de água (km)', # Ext rede água
'ES004 - Extensão da rede de esgotos (km)', # Ext rede esgotos
'FN026 - Quantidade total de empregados próprios (Empregados)'# Empregados
]]
top100.shape
top100 = top100.dropna()
top100.shape
top100 = top100.sort_values(by='POP_TOT - População total do município do ano de referência (Fonte: IBGE): (Habitantes)',ascending=False)
# foram 110 porque 5 empresas a mais para 5 municipios
top100 = top100[:200]
top100.shape
top100.head()
top100.groupby(['Tipo Empresa']).count()
top100[top100['Código do Município '] == 355030]
#não precisa pois esta dado drop na top100 com todo o conjunto
# top100 = top100.fillna(0)
top100.info()
top100.describe()
#Alterando virgulas por pontos e transformando coluna em inteiro
top100['IN013 - Índice de perdas faturamento (percentual)'] = top100['IN013 - Índice de perdas faturamento (percentual)'].str.replace(',', '.')
top100['IN013 - Índice de perdas faturamento (percentual)'] = pd.to_numeric(top100['IN013 - Índice de perdas faturamento (percentual)'])
#Alterando virgulas por pontos e transformando coluna em inteiro
top100['IN049 - Índice de perdas na distribuição (percentual)'] = top100['IN049 - Índice de perdas na distribuição (percentual)'].str.replace(',', '.')
top100['IN049 - Índice de perdas na distribuição (percentual)'] = pd.to_numeric(top100['IN049 - Índice de perdas na distribuição (percentual)'])
#Alterando virgulas por pontos e transformando coluna em inteiro
top100['IN004 - Tarifa média praticada (R$/m³)'] = top100['IN004 - Tarifa média praticada (R$/m³)'].str.replace(',', '.')
top100['IN004 - Tarifa média praticada (R$/m³)'] = pd.to_numeric(top100['IN004 - Tarifa média praticada (R$/m³)'])
#Alterando virgulas por pontos e transformando coluna em inteiro
top100['AG005 - Extensão da rede de água (km)'] = top100['AG005 - Extensão da rede de água (km)'].str.replace('.', '')
top100['AG005 - Extensão da rede de água (km)'] = top100['AG005 - Extensão da rede de água (km)'].str.replace(',', '.')
top100['AG005 - Extensão da rede de água (km)'] = pd.to_numeric(top100['AG005 - Extensão da rede de água (km)'])
#Alterando virgulas por pontos e transformando coluna em inteiro
top100['ES004 - Extensão da rede de esgotos (km)'] = top100['ES004 - Extensão da rede de esgotos (km)'].str.replace('.', '')
top100['ES004 - Extensão da rede de esgotos (km)'] = top100['ES004 - Extensão da rede de esgotos (km)'].str.replace(',', '.')
top100['ES004 - Extensão da rede de esgotos (km)'] = pd.to_numeric(top100['ES004 - Extensão da rede de esgotos (km)'])
top100.head()
top100.info()
# Municipios repetidos por terem duas empresas
top100[top100.duplicated('Código do Município ')==True]
data_tatra_brasil = pd.read_excel("Tabela_Trata_Brasil.xlsx",sheetname='Tabela Final Filtros', encoding='latin1', skiprows=3)
data_tatra_brasil.head()
novo_trata = data_tatra_brasil[['Código do município', # Cód Município
'Município', # 'Município'
'UF', # UF
'Operador', # Empresa
'Indicador de atendimento urbano de água (%)', # At Água
'Indicador de atendimento urbano de esgoto (%)', # At Esgoto
'Indicador novas ligações de água/ligações faltantes (%)', # Novas Lig Água
'Indicador novas ligações de esgoto/ligações faltantes (%)', # Novas Lig Esgoto
'Indicador evolução nas perdas de faturamento (%)', # Ev Perdas Fat
'Indicador evolução nas perdas de distribuição (%)', # Ev Perdas Dist
'Indicador de esgoto tratado por água consumida (%)', # Esg Tratado
'Investimento 5 anos (Milhões R$)'# Investimento
]]
novo_trata = novo_trata.round({'Indicador de atendimento urbano de água (%)':2,
'Indicador de atendimento urbano de esgoto (%)':2,
'Indicador novas ligações de água/ligações faltantes (%)':2,
'Indicador novas ligações de esgoto/ligações faltantes (%)':2,
'Indicador evolução nas perdas de faturamento (%)':2,
'Indicador evolução nas perdas de distribuição (%)':2,
'Indicador de esgoto tratado por água consumida (%)':2,
'Investimento 5 anos (Milhões R$)':2
})
novo_trata.head()
novo_trata[novo_trata['Município'] == 'São Paulo']
novo_trata.info()
novo_trata.describe()
len(novo_trata['Município'].unique())
data_tatra_brasil['Código do município'].head()
novo_trata['Código do município'].dtypes
novo_trata['Código do município'].dtypes
top100['Código do Município '].dtypes
novo_trata['Código do município'].dtypes
novo_trata[novo_trata['Código do município'] == 330455]
cod_municipios = novo_trata['Código do município']
cod_municipios.head()
top100['Código do Município '].head()
top100['Código do Município '].dtypes
#Visualizando tamanho do dataset
top100.shape
# df_colunas = df.columns
top100_colunas = pd.DataFrame(top100.columns, columns=['colunas'])
trata_colunas = pd.DataFrame(novo_trata.columns, columns=['colunas'])
top100_colunas
trata_colunas
```
- At água - Indicador de atendimento urbano de água (%)
- At esgoto - Indicador de atendimento urbano de esgoto (%)
- Esgoto tratado - Indicador de esgoto tratado por água consumida (%)
- Novas lig água - Indicador novas ligações de água/ligações faltantes (%)
- Novas lig esgoto - Indicador novas ligações de esgoto/ligações faltantes (%)
- Ev perdas fat - Indicador evolução nas perdas de faturamento (%)
- Ev perdas dist - Indicador evolução nas perdas de distribuição (%)
- Perdas fat - Indicador perdas no faturamento 2016 (%)
- Perdas dist - Indicador perdas na distribuição 2016 (%)
- Tarifa méd - Tarifa média (Real/m³)
- Investimento - Investimento
- Ext rede água - Extensão rede de água
- Ext rede esgotos - Extensão rede de esgotos
- N func - N de funcionários
```
#data_tatra_brasil['Código do município']
top100['Código do município'] = top100['Código do Município ']
top100['Código do município'].dtypes
novo_trata['Código do município'].dtypes
type(top100)
type(novo_trata)
novo_trata
tabela_limpa = pd.merge(top100, novo_trata, on='Código do município', how='inner')
tabela_limpa.shape
tabela_limpa.head()
#tabela_limpa = novo_trata.merge(novo_top100, left_on='Código do município', right_on='Código do Município ', how='outer')
tabela_limpa[tabela_limpa.duplicated('Código do município')==True]
len(tabela_limpa)
# Renomear colunas
tabela_limpa.columns = ['Cód Município',
'Tp Empresa',
'Nat Jurídica',
'Pop Total',
'Perdas Fat',
'Perdas Dist',
'Tarifa méd',
'Ext rede água',
'Ext rede esgotos',
'Empregados',
'Código do município',
'Município',
'UF',
'Empresa',
'At Água',
'At Esgoto',
'Novas Lig Água',
'Novas Lig Esgoto',
'Ev Perdas Fat',
'Ev Perdas Dist',
'Esg Tratado',
'Investimento'
]
tabela_limpa.head()
tabela_limpa[tabela_limpa['Município']=='São Paulo']
#Criando novo dataset com os dados limpos
tabela_limpa.to_csv('dados_limpos_saneamento.csv', index=False)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/mengwangk/dl-projects/blob/master/04_06_auto_ml_2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Automated ML - Tuning
```
COLAB = True
DATASET_NAME = '4D.zip'
FEATURE_DATASET_PREFIX = 'feature_matrix_d2_v3'
if COLAB:
!pip install -U imblearn
!rm -rf dl-projects
!git clone https://github.com/mengwangk/dl-projects
if COLAB:
!cp dl-projects/utils* .
!cp dl-projects/preprocess* .
!cp dl-projects/plot* .
%load_ext autoreload
# %reload_ext autoreload
%autoreload 2
%matplotlib inline
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import math
import matplotlib
import sys
from scipy import stats
from collections import Counter
from pathlib import Path
plt.style.use('fivethirtyeight')
sns.set(style="ticks")
import featuretools as ft
import warnings
warnings.filterwarnings('ignore')
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import MinMaxScaler, StandardScaler, RobustScaler
from sklearn.impute import SimpleImputer
from sklearn.metrics import precision_score, recall_score, f1_score, confusion_matrix, roc_auc_score, precision_recall_curve, roc_curve, mean_squared_error, accuracy_score, average_precision_score, classification_report
from sklearn.model_selection import train_test_split, cross_val_score, GridSearchCV, RandomizedSearchCV
from sklearn.decomposition import PCA
import pylab as pl
from collections import Counter
# from skopt import BayesSearchCV
# from skopt.space import Real, Categorical, Integer
# from sklearn.ensemble import RandomForestClassifier
# from scikitplot.plotters import plot_precision_recall_curve
from dateutil.relativedelta import relativedelta
from IPython.display import display
from utils import *
from preprocess import *
import xgboost as xgb
np.set_printoptions(threshold=sys.maxsize)
# The Answer to the Ultimate Question of Life, the Universe, and Everything.
np.random.seed(42)
from utils import feature_selection, plot_feature_importances
from plot import plot_correlation_matrix, plot_labelled_scatter
%aimport
```
## Preparation
```
if COLAB:
from google.colab import drive
drive.mount('/content/gdrive')
GDRIVE_DATASET_FOLDER = Path('gdrive/My Drive/datasets/')
if COLAB:
DATASET_PATH = GDRIVE_DATASET_FOLDER
ORIGIN_DATASET_PATH = Path('dl-projects/datasets')
else:
DATASET_PATH = Path("datasets")
ORIGIN_DATASET_PATH = Path('datasets')
DATASET = DATASET_PATH/f"{FEATURE_DATASET_PREFIX}.ft"
ORIGIN_DATASET = ORIGIN_DATASET_PATH/DATASET_NAME
if COLAB:
!ls -l gdrive/"My Drive"/datasets/ --block-size=M
!ls -l dl-projects/datasets --block-size=M
data = pd.read_feather(DATASET)
origin_data = format_tabular(ORIGIN_DATASET)
data.info()
```
## Exploratory Data Analysis
### View data
```
# Feature matrix
feature_matrix = data.drop(columns=['NumberId', 'month', 'year'])
# Sort data
feature_matrix.sort_values(by=['time', 'MAX(Results.LuckyNo)'], inplace=True)
print('Positive: ' + str(feature_matrix['Label'].value_counts()[0]) + ' which is ', round(feature_matrix['Label'].value_counts()[0]/len(feature_matrix) * 100,2), '% of the dataset')
print('Negative: ' + str(feature_matrix['Label'].value_counts()[1]) + ' which is ', round(feature_matrix['Label'].value_counts()[1]/len(feature_matrix) * 100,2), '% of the dataset')
plt.figure(figsize=(8, 8))
sns.countplot('Label', data=feature_matrix)
feature_matrix.isna().sum().sort_values(ascending=False)
feature_matrix[feature_matrix.isnull().any(axis=1)].head()
```
### Data Cleansing
```
## Fill all NaN with 0
feature_matrix = feature_matrix.fillna(0)
feature_matrix.isna().sum().sort_values(ascending=False)
feature_matrix[feature_matrix.isnull().any(axis=1)].head()
```
### Feature Selection
```
# Feature scaling first??
print(feature_matrix.shape)
feature_matrix.columns
feature_matrix_selection = feature_selection(feature_matrix.drop(columns = ['time', 'TotalStrike', 'Label']))
feature_matrix_selection.shape, feature_matrix_selection.columns
feature_matrix_selection['time'] = feature_matrix['time']
feature_matrix_selection['TotalStrike'] = feature_matrix['TotalStrike']
feature_matrix_selection['Label'] = feature_matrix['Label']
```
### Feature Correlation
```
# Check without feature selection
# corrs = feature_matrix.corr().sort_values('Label')
# corrs['Label'].tail(100)
# Check with feature selection
corrs = feature_matrix_selection.corr().sort_values('Label')
corrs['Label'].tail(20)
```
### Balancing data
```
from imblearn.under_sampling import (RandomUnderSampler,
ClusterCentroids,
TomekLinks,
NeighbourhoodCleaningRule,
AllKNN,
NearMiss)
from imblearn.combine import SMOTETomek
from imblearn.pipeline import make_pipeline
pipeline = Pipeline([('imputer', SimpleImputer(strategy = 'constant', fill_value=0)), ('scaler', StandardScaler())])
def under_sampling_test(feature_data, cut_off_date, ratio=0.8):
y = feature_data.loc[feature_data['time'] < cut_off_date, 'Label']
X = feature_data[feature_data['time'] < cut_off_date].drop(columns = ['Label', 'TotalStrike','time','date'], errors='ignore')
total_count = y.value_counts()
neg_count = y.value_counts()[0]
pos_count = y.value_counts()[1]
target_neg_count = int(pos_count / (1-ratio))
target_ratio = {0: target_neg_count, 1: pos_count}
print(X.shape, y.shape, target_ratio)
# print('Before sampling {}'.format(Counter(y)))
sampler = RandomUnderSampler(sampling_strategy=target_ratio, random_state=42)
X_bal, y_bal = sampler.fit_sample(X, y)
print('Undersampling {}'.format(Counter(y_bal)))
return X_bal, y_bal
# https://stackoverflow.com/questions/52499788/smotetomek-how-to-set-ratio-as-dictionary-for-fixed-balance
def balancing_pipeline_test(feature_data, cut_off_date, ratio=0.8):
y = feature_data.loc[feature_data['time'] < cut_off_date, 'Label']
X = feature_data[feature_data['time'] < cut_off_date].drop(columns = ['Label', 'TotalStrike','time','date'], errors='ignore')
total_count = y.value_counts()
neg_count = y.value_counts()[0]
pos_count = y.value_counts()[1]
target_neg_count = int(pos_count / (1-ratio))
target_ratio = {0: target_neg_count, 1: pos_count}
print('Before sampling {}'.format(Counter(y)))
#sampler = NearMiss(sampling_strategy={0: target_neg_count}, n_jobs=4)
sampler = SMOTETomek(sampling_strategy='auto')
X = pipeline.fit_transform(X)
X_bal, y_bal = sampler.fit_sample(X, y)
print('Over and undersampling {}'.format(Counter(y_bal)))
return X_bal, y_bal
%time X_res, y_res = balancing_pipeline_test(feature_matrix_selection, pd.datetime(2019,6,1))
def plot_this(X_rs,y_rs,method):
# Use principal component to condense the 10 features to 2 features
pca = PCA(n_components=2).fit(X_rs)
pca_2d = pca.transform(X_rs)
# Assign colors
for i in range(0, pca_2d.shape[0]):
if y_rs[i] == 0:
c1 = pl.scatter(pca_2d[i,0],pca_2d[i,1],c='r', marker='o')
elif y_rs[i] == 1:
c2 = pl.scatter(pca_2d[i,0],pca_2d[i,1],c='g', marker='*')
pl.legend([c1, c2], ['Class 1', 'Class 2'])
pl.title(method)
pl.axis([-4, 5, -4, 4]) # x axis (-4,5), y axis (-4,4)
pl.show()
# Plot
plot_this(X_res, y_res, 'SMOTETomek')
def under_sampling(X, y, ratio=0.9):
total_count = y.value_counts()
neg_count = y.value_counts()[0]
pos_count = y.value_counts()[1]
target_neg_count = int(pos_count / (1-ratio))
target_ratio = {0: target_neg_count, 1: pos_count}
#print(X.shape, y.shape, target_ratio)
print('Before sampling {}'.format(Counter(y)))
sampler = RandomUnderSampler(random_state=42, sampling_strategy=target_ratio)
X_bal, y_bal = sampler.fit_sample(X, y)
print('Undersampling {}'.format(Counter(y_bal)))
return X_bal, y_bal
def over_under_sampling(X, y):
pass
```
## Modeling
```
def predict(dt, feature_matrix, sampling = False, return_probs = False):
feature_matrix['date'] = feature_matrix['time']
# Subset labels
test_labels = feature_matrix.loc[feature_matrix['date'] == dt, 'Label']
train_labels = feature_matrix.loc[feature_matrix['date'] < dt, 'Label']
print(f"Size of test labels {len(test_labels)}")
print(f"Size of train labels {len(train_labels)}")
# Features
X_train = feature_matrix[feature_matrix['date'] < dt].drop(columns = ['NumberId', 'time',
'date', 'Label', 'TotalStrike', 'month', 'year', 'index'], errors='ignore')
X_test = feature_matrix[feature_matrix['date'] == dt].drop(columns = ['NumberId', 'time',
'date', 'Label', 'TotalStrike', 'month', 'year', 'index'], errors='ignore')
print(f"Size of X train {len(X_train)}")
print(f"Size of X test {len(X_test)}")
feature_names = list(X_train.columns)
# Impute and scale features
pipeline = Pipeline([('imputer', SimpleImputer(strategy = 'constant', fill_value=0)), ('scaler', StandardScaler())])
# Fit and transform training data
X_train = pipeline.fit_transform(X_train)
X_test = pipeline.transform(X_test)
# Balance the data
if sampling:
X_train, train_labels = under_sampling(X_train, train_labels)
# Labels
y_train = np.array(train_labels).reshape((-1, ))
y_test = np.array(test_labels).reshape((-1, ))
print('Training on {} observations.'.format(len(X_train)))
print('Testing on {} observations.\n'.format(len(X_test)))
# https://xgboost.readthedocs.io/en/latest/parameter.html
# https://xgboost.readthedocs.io/en/latest/tutorials/param_tuning.html
# https://stats.stackexchange.com/questions/224512/reduce-false-positives-with-xgboost
if type(train_labels) == np.ndarray:
hit_ratio = float( len(np.where(train_labels == 0)[0]) / len(np.where(train_labels == 1)[0]) )
else:
hit_ratio = float(train_labels.value_counts()[0]/train_labels.value_counts()[1])
print(f"Hit ratio - {hit_ratio}\n")
# Create the classifier
model = xgb.XGBClassifier(n_jobs=-1,
random_state = 42,
n_estimators=100,
max_depth=3,
objective='binary:logistic',
min_child_weight=1,
scale_pos_weight=hit_ratio
)
# Train
model.fit(X_train, y_train)
# Make predictions
predictions = model.predict(X_test)
probs = model.predict_proba(X_test)[:, 1]
# Total positive
positive = np.where((predictions==1))
print(f'Total predicted to be positive: {len(positive[0])} \n')
# Calculate metrics
rpt = classification_report(y_test, predictions)
cm = confusion_matrix(y_test, predictions)
print('Classification report')
print(rpt)
print(f'Confusion matrix:\n {cm}\n')
# Total predicted matches
print('Predicted matches')
pred = np.where((predictions==1))
print(len(pred[0]), pred[0][0:23])
topN = np.argpartition(probs, -23)[-23:]
print(f'\n{topN}\n') # Top N most high probability numbers
if len(positive[0]) > 0:
# Matching draws
print('Matched draws')
md = np.where((predictions==1) & (y_test==1))
print(f"Count: {len(md[0])}, Index: {md}")
month_data = feature_matrix.loc[feature_matrix['date'] == dt]
numbers = month_data.iloc[md[0]][['MAX(Results.LuckyNo)']]
print('\n\nTop 23 Possibility')
print(origin_data[(origin_data['DrawDate'].dt.year == dt.year) &
(origin_data['DrawDate'].dt.month == dt.month) &
(origin_data['LuckyNo'].isin(topN))].head(23))
print('\n\nFirst 23 Numbers')
print(origin_data[(origin_data['DrawDate'].dt.year == dt.year) &
(origin_data['DrawDate'].dt.month == dt.month) &
(origin_data['LuckyNo'].isin(pred[0][0:23]))].head(23))
print('\n\nAll matched')
print(origin_data[(origin_data['DrawDate'].dt.year == dt.year) &
(origin_data['DrawDate'].dt.month == dt.month) &
(origin_data['LuckyNo'].isin(numbers['MAX(Results.LuckyNo)']))].head(100))
else:
print('No luck this month')
# Feature importances
fi = pd.DataFrame({'feature': feature_names, 'importance': model.feature_importances_})
if return_probs:
return fi, probs
return fi
%time june_2019 = predict(pd.datetime(2019,6,1), feature_matrix_selection)
#normalized_fi = plot_feature_importances(june_2019)
# Loop through from June to Dec
# start_mt = pd.datetime(2019,6,1)
# how_many_mt = 7
# for i in range(how_many_mt):
# month_to_predict = start_mt + relativedelta(months=i)
# print(f"\n{month_to_predict}\n-------------------\n")
# %time predict(month_to_predict, feature_matrix_selection)
```
| github_jupyter |
# 海上风力发电场
等级:入门
## 目的和先决条件
在此示例中,我们将解决如何最小化铺设水下电缆以收集海上风电场产生的电力的成本的问题。我们将构建此问题的混合整数编程(MIP)模型,在Gurobi Python界面中实现此模型,并计算最佳解决方案。
该建模示例处于初级阶段,我们假设您了解Python,并且具有一些有关构建数学优化模型的知识。
**注意:** 您可以通过单击 [此处](https://github.com/Gurobi/modeling-examples/archive/master.zip) 下载包含此示例和其他示例的仓库。为了正确运行此 Jupyter Notebook,您必须具有Gurobi许可证。如果您没有,则可以 *商业用户身份* 申请
[评估许可证](https://www.gurobi.com/downloads/request-an-evaluation-license/?utm_source=Github&utm_medium=website_JupyterME&utm_campaign=CommercialDataScience),或以 *学术用户* 的身份下载
[免费许可证](https://www.gurobi.com/academia/academic-program-and-licenses/?utm_source=Github&utm_medium=website_JupyterME&utm_campaign=AcademicDataScience)。
## 动机
全球气候变化已经对环境产生了明显影响。冰川消退,河流和湖泊上的冰块破裂早于预期,动植物物种受到影响,树木开花快于预期。全球气候变化的潜在未来影响包括更频繁的山货,某些地区更长的干旱时期以及热带风暴的数量,持续时间和强度增加。[1]
缓解气候变化包括限制全球变暖的幅度或速度及其相关影响的行动。缓解气候变化的第一个挑战是消除煤炭,石油和最终天然气的燃烧。这可能是最艰巨的挑战,因为富裕国家的居民吃饭、穿衣、工作、娱乐甚至睡觉都离不开化石燃料。同样,发展中国家的人民也希望且也应该得到同样的生活。目前对于减少对化石燃料的依赖,并没有完美的解决方案(例如,碳中性生物燃料会抬高食品价格并导致森林破坏;核能虽然不会排放温室气体,但会产生放射性废物)。其他替代品还包括植物衍生的塑料,生物柴油和风力发电等。
海上风力发电是利用风力发电场建造在水体中,通常是在海洋中,以收集风能来发电。与陆地相比,海上的风速更高,因此海上风力发电的单位装机容量更高。
将风力涡轮机安装在海上的好处是,沿海地区的风力要强得多,与大陆上的风力不同,下午的海上微风可能很强劲,与人们用电最多的时间相称。海上涡轮机也可以靠近沿海地区(例如大城市)的负载中心,从而无需新的长距离传输线。
## 问题描述
海上风电场是一组放置在海上的风力涡轮机,以利用强大的海上风力。这些强风可以产生更多的电力,但海上风力发电场的安装和运营成本比陆地上的要高。
我们将使用MIP模型来减少建造海上风电场的部分成本。我们将计算出如何铺设连接涡轮机的水下电缆的计划。这些电缆是将涡轮机产生的电力输送到陆地所必需的。我们计算的计划将使安装水下电缆的成本降到最低,同时确保每个涡轮机都与海岸相连,每根电缆都有足够的容量来处理产生的电流。
在我们的例子中,丹麦西海岸正在建设一个风力发电场。海岸上有一座发电站,所有的电力都必须被输送到电网中。风电场中还设有中转站,可以收集来自多个涡轮机的电力,并通过一条电缆将其传送到岸上。
在安装电缆时,我们必须考虑两个因素。首先,在海底铺设电缆是有固定成本的。这种成本与电缆连接的两个站点之间的距离成正比。第二,我们必须考虑有多少电流会通过电缆。承载大电流的连接需要厚电缆。粗电缆比细电缆贵。
该优化问题的目标是以最小的成本铺设连接风电场电网的电缆。
海上风电场优化问题模型是一种更为普遍的优化模型(称为固定费用和流量问题)一个实例。固定收费网络流量问题可以应用于大量的业务问题——例如,在通信和运输网络的规划中。
## Solution Approach
Mathematical programming is a declarative approach where the modeler formulates a mathematical optimization model that captures the key aspects of a complex decision problem. The Gurobi Optimizer solves such models using state-of-the-art mathematics and computer science.
A mathematical optimization model has five components, namely:
* Sets and indices.
* Parameters.
* Decision variables.
* Objective function(s).
* Constraints.
We now present a MIP formulation for the offshore wind farming problem.
## Model Formulation
### Sets and Indices
$G(V,E)$: Graph that represents the wind farm network, where $V$ is the set of vertices and $E$ is the set of edges. The turbines, transfer stations, and power stations are vertices in the set of vertices $V$ of the graph. The set of potential cables are the edges of the graph.
### Parameters
$s_{i} \in \mathbb{R}$: Power supplied at vertex $i \in V$. Since turbines supply power, they are source vertices with $s_{i} > 0$. Transfer stations do not supply or remove power from the network so they have $s_{i} = 0$. The power station on the coast is a sink that remove all power from the wind farm so it has $s_{i} < 0$.
$u_{i,j} \in \mathbb{R}^+ $: Maximum current capacity a cable can handle from vertex $i \in V$ to vertex $j \in V$.
$c_{i,j} \in \mathbb{R}^+$: Cost per unit of current flow going from vertex $i \in V$ to vertex $j \in V$, i.e. the price we must pay to increase the thickness of the cable to handle an increase in current.
$f_{i,j} \in \mathbb{R}^+$: Fixed cost of laying a cable from vertex $i \in V$ to vertex $j \in V$, and is the result of multiplying the distance between vertices by the cost per mile.
### Decision Variables
$install_{i,j} \in \{0, 1 \}$: This variable is equal to 1 if we lay a cable from vertex $i \in V$ to vertex $j \in V$; and 0 otherwise.
$flow_{i,j} \geq 0$: This non-negative continuous variable represents the amount of current flowing from vertex $i \in V$ to vertex $j \in V$.
The goal of this optimization model is to decide which of these potential edges in the graph should be used at a minimum cost.
### Objective Function
- **Total costs**. We want to minimize the total cost to install the cables. The term on the left is the variable costs (i.e. those that vary according to the current in the cable). The term on right is the fixed cost to install the cable.
\begin{equation}
\text{Max} \quad Z = \sum_{(i,j) \in E} c_{i,j} \cdot flow_{i,j} + \sum_{(i,j) \in E} f_{i,j} \cdot install_{i,j}
\tag{0}
\end{equation}
### Constraints
- **Balance**. For each vertex $i \in V$, we want to ensure the conservation of current in the network.
\begin{equation}
\sum_{j:(i,j) \in E} flow_{i,j} - \sum_{j:(j,i) \in E} flow_{j,i} = s_{i} \quad \forall i \in V
\tag{1}
\end{equation}
- **Capacity**. For each edge $(i,j) \in E$, we want to enforce the limits on the maximum current capacity of each cable.
\begin{equation}
0 \leq flow_{i,j} \leq u_{i,j} \cdot install_{i,j} \quad \forall (i,j) \in E
\tag{2}
\end{equation}
## Python Implementation
This example considers three turbines, one transfer station, and two power stations. The current flowing out at each vertex of the wind farm network is presented in the following table. Recall that since turbines supply power their capacity is positive. Transfer stations do not supply or remove power from the network so they have a capacity of zero. The power stations on the coast are sinks that remove all power from the wind farm network so they have demand of power, in this case we use a negative number.
| <i></i> | Capacity in MW |
| --- | --- |
| vertex 1 | 4 |
| vertex 2 | 3 |
| vertex 3 | 2 |
| vertex 4 | 0 |
| vertex 5 | -6 |
| vertex 6 | -3 |
The capacity, flow cost, and fixed cost of each edge in the wind farm network are provided in the following table.
| <i></i> | Capacity in MW | Flow cost in millions of Euros | Fixed cost in millions of Euros|
| --- | --- | --- | --- |
| Edge: (0,4) | 4 | 1 | 1 |
| Edge: (0,3) | 2 | 1 | 1 |
| Edge: (1,3) | 3 | 1 | 1 |
| Edge: (2,5) | 2 | 1 | 1 |
| Edge: (3,4) | 2 | 1 | 1 |
| Edge: (3,5) | 1 | 1 | 1 |
We now import the Gurobi Python Module. Then, we initialize the data structures with the given data.
```
import gurobipy as gp
from gurobipy import GRB
# Parameters
vertices = {0: 4, 1: 3, 2: 2, 3: 0, 4: -6, 5: -3}
edges, cap, flow_cost, fixed_cost = gp.multidict({
(0,4): [4,1,1],
(0,3): [2,1,1],
(1,3): [3,1,1],
(2,5): [2,1,1],
(3,4): [2,1,1],
(3,5): [1,1,1]
})
```
### Model Deployment
We now determine the MIP model for the offshore wind farming problem, by defining the decision variables, constraints, and objective function. Next, we start the optimization process and Gurobi finds the plan to lay cables at the offshore wind farming network that minimizes total costs.
```
# MIP model formulation
m = gp.Model("offshore_wind_farming")
# Add variables
install = m.addVars(edges, vtype=GRB.BINARY, name="Install")
flow = m.addVars(edges, vtype=GRB.CONTINUOUS, name="Flow")
# Add constraints
m.addConstrs((flow.sum(v,'*') - flow.sum('*',v) == supply for v, supply in vertices.items()), name="Flow_conservation")
m.addConstrs((flow[e] <= cap[e]*install[e] for e in edges), name="Install2flow")
# Set objective
m.setObjective(flow.prod(flow_cost) + install.prod(fixed_cost), GRB.MINIMIZE)
m.optimize()
```
## Analysis
The result of the optimization model shows that the minimum total cost value is 17 million Euros. Let's see the solution that achieves that optimal result.
### Cable Installation Plan
This plan determines the layout of cables in the offshore wind farming network.
```
# display which edges in the offshore wind farming network we plan to install.
for origin, end in install.keys():
if (abs(install[origin, end].x) > 0.5):
print(f"Install cable from location {origin + 1} to location {end + 1} in the offshore wind farming network ")
```
### Cable Capacity Plan
This plan determines the current flow capacity in MW of each cable installed.
```
# Current flow capacity of each cable installed
for origin, end in flow.keys():
if (abs(flow[origin, end].x) > 1e-6):
print(f"The capacity of cable installed from location {origin + 1} to location {end + 1} is {flow[origin, end].x} MW ")
```
## 结论
在这个例子中,我们解决了一个海上风电场的问题,我们想把铺设海底电缆收集海上风电场网络产生的电力的成本降到最低。我们学习了如何将问题表述为MIP模型。此外,我们还学习了如何实现MIP模型公式,并使用Gurobi Python API来解决它。
## 参考资料
[1] https://climate.nasa.gov/effects/
[2] https://www.scientificamerican.com/article/10-solutions-for-climate-change/
Copyright © 2020 Gurobi Optimization, LLC
翻译 By Arvin Xu
| github_jupyter |
<a href="https://colab.research.google.com/github/DJCordhose/ml-workshop/blob/master/notebooks/tf2/feature_columns.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Skeleton for Time Series data
* Idea and some code taken from, and also OPSD time series data set explained here: https://www.dataquest.io/blog/tutorial-time-series-analysis-with-pandas/
```
import pandas as pd
print(pd.__version__)
import numpy as np
print(np.__version__)
```
## Standard Case: explicit time stamp
* each entry belongs to one time stamp
* we just have a sequence of events
_let me know if your case looks different_
```
pd.read_csv?
# for local
# url = 'opsd_germany_daily.csv'
# for colab
url = 'https://raw.githubusercontent.com/jenfly/opsd/master/opsd_germany_daily.csv'
time_series_df = pd.read_csv(url,
sep=',',
index_col=0, # you can use the date as the index for pandas
parse_dates=[0]) # where is the time stamp?
import matplotlib.pyplot as plt
%matplotlib inline
axes = time_series_df.plot(figsize=(20, 8), subplots=True)
for ax in axes:
ax.set_ylabel('Daily Totals (GWh)')
time_series_df.shape
time_series_df.describe()
# just january 2017
time_series_df['2017-01']
# exactly what you think it does
time_series_df['2017-01-01': '2017-01-05']
```
### Recurrent Neural Networks
```
days_2016_2017 = time_series_df['2016': '2017']['Consumption'].to_numpy()
days_2016_2017.shape
plt.figure(figsize=(20,8))
plt.plot(days_2016_2017)
# split a univariate sequence into samples
def split_sequence(sequence, n_steps_in, n_steps_out):
X, y = list(), list()
for i in range(len(sequence)):
# find the end of this pattern
end_ix = i + n_steps_in
out_end_ix = end_ix + n_steps_out
# check if we are beyond the sequence
if out_end_ix > len(sequence):
break
# gather input and output parts of the pattern
seq_x, seq_y = sequence[i:end_ix], sequence[end_ix:out_end_ix]
X.append(seq_x)
y.append(seq_y)
return np.array(X), np.array(y)
```
### For each week predict the nexy view days
```
# most important numbers: how many days used for prediction, and how many dayes are being predicted
n_steps_in, n_steps_out = 7, 30
X, Y = split_sequence(days_2016_2017, n_steps_in, n_steps_out)
X.shape, Y.shape
X[0], Y[0]
X[1], Y[1]
# reshape from [samples, timesteps] into [samples, timesteps, features]
n_features = 1
X = X.reshape((X.shape[0], X.shape[1], n_features))
X.shape
# Gives us a well defined version of tensorflow
try:
# %tensorflow_version only exists in Colab.
%tensorflow_version 2.x
except Exception:
pass
import tensorflow as tf
print(tf.__version__)
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras.layers import Dense, LSTM, GRU, SimpleRNN, Bidirectional
from tensorflow.keras.models import Sequential, Model
model = Sequential()
model.add(GRU(100, activation='relu', input_shape=(n_steps_in, n_features)))
# model.add(GRU(100, activation='relu', return_sequences=True, input_shape=(n_steps_in, n_features)))
# model.add(GRU(100, activation='relu'))
model.add(Dense(n_steps_out))
model.compile(optimizer='adam', loss='mse')
model.summary()
%%time
history = model.fit(X, Y, epochs=200, verbose=0)
import matplotlib.pyplot as plt
plt.figure(figsize=(20,8))
plt.yscale('log')
plt.plot(history.history['loss'])
model.evaluate(X, Y, verbose=0)
final_week_2017 = X[-1]
final_week_2017
first_week_2018 = model.predict(final_week_2017.reshape(1, 7, 1))
first_week_2018
plt.figure(figsize=(20,8))
plt.plot(days_2016_2017)
plt.plot(first_week_2018[0], color='r')
known_x = np.arange(len(days_2016_2017))
pred_x = np.arange(len(days_2016_2017), len(days_2016_2017) + n_steps_out)
plt.figure(figsize=(20,8))
plt.plot(known_x, days_2016_2017)
plt.plot(pred_x, first_week_2018[0], color='r')
```
| github_jupyter |
```
#default_exp transform
#export
from fastcore.imports import *
from fastcore.foundation import *
from fastcore.utils import *
from fastcore.dispatch import *
import inspect
from nbdev.showdoc import *
from fastcore.test import *
from fastcore.nb_imports import *
```
# Transforms
> Definition of `Transform` and `Pipeline`
The classes here provide functionality for creating a composition of *partially reversible functions*. By "partially reversible" we mean that a transform can be `decode`d, creating a form suitable for display. This is not necessarily identical to the original form (e.g. a transform that changes a byte tensor to a float tensor does not recreate a byte tensor when decoded, since that may lose precision, and a float tensor can be displayed already).
Classes are also provided and for composing transforms, and mapping them over collections. `Pipeline` is a transform which composes several `Transform`, knowing how to decode them or show an encoded item.
## Transform -
```
#export
_tfm_methods = 'encodes','decodes','setups'
class _TfmDict(dict):
def __setitem__(self,k,v):
if k not in _tfm_methods or not callable(v): return super().__setitem__(k,v)
if k not in self: super().__setitem__(k,TypeDispatch())
self[k].add(v)
#export
class _TfmMeta(type):
def __new__(cls, name, bases, dict):
res = super().__new__(cls, name, bases, dict)
for nm in _tfm_methods:
base_td = [getattr(b,nm,None) for b in bases]
if nm in res.__dict__: getattr(res,nm).bases = base_td
else: setattr(res, nm, TypeDispatch(bases=base_td))
res.__signature__ = inspect.signature(res.__init__)
return res
def __call__(cls, *args, **kwargs):
f = args[0] if args else None
n = getattr(f,'__name__',None)
if callable(f) and n in _tfm_methods:
getattr(cls,n).add(f)
return f
return super().__call__(*args, **kwargs)
@classmethod
def __prepare__(cls, name, bases): return _TfmDict()
#export
def _get_name(o):
if hasattr(o,'__qualname__'): return o.__qualname__
if hasattr(o,'__name__'): return o.__name__
return o.__class__.__name__
#export
def _is_tuple(o): return isinstance(o, tuple) and not hasattr(o, '_fields')
#export
class Transform(metaclass=_TfmMeta):
"Delegates (`__call__`,`decode`,`setup`) to (<code>encodes</code>,<code>decodes</code>,<code>setups</code>) if `split_idx` matches"
split_idx,init_enc,order,train_setup = None,None,0,None
def __init__(self, enc=None, dec=None, split_idx=None, order=None):
self.split_idx = ifnone(split_idx, self.split_idx)
if order is not None: self.order=order
self.init_enc = enc or dec
if not self.init_enc: return
self.encodes,self.decodes,self.setups = TypeDispatch(),TypeDispatch(),TypeDispatch()
if enc:
self.encodes.add(enc)
self.order = getattr(enc,'order',self.order)
if len(type_hints(enc)) > 0: self.input_types = first(type_hints(enc).values())
self._name = _get_name(enc)
if dec: self.decodes.add(dec)
@property
def name(self): return getattr(self, '_name', _get_name(self))
def __call__(self, x, **kwargs): return self._call('encodes', x, **kwargs)
def decode (self, x, **kwargs): return self._call('decodes', x, **kwargs)
def __repr__(self): return f'{self.name}:\nencodes: {self.encodes}decodes: {self.decodes}'
def setup(self, items=None, train_setup=False):
train_setup = train_setup if self.train_setup is None else self.train_setup
return self.setups(getattr(items, 'train', items) if train_setup else items)
def _call(self, fn, x, split_idx=None, **kwargs):
if split_idx!=self.split_idx and self.split_idx is not None: return x
return self._do_call(getattr(self, fn), x, **kwargs)
def _do_call(self, f, x, **kwargs):
if not _is_tuple(x):
if f is None: return x
ret = f.returns(x) if hasattr(f,'returns') else None
return retain_type(f(x, **kwargs), x, ret)
res = tuple(self._do_call(f, x_, **kwargs) for x_ in x)
return retain_type(res, x)
add_docs(Transform, decode="Delegate to <code>decodes</code> to undo transform", setup="Delegate to <code>setups</code> to set up transform")
show_doc(Transform)
```
A `Transform` is the main building block of the fastai data pipelines. In the most general terms a transform can be any function you want to apply to your data, however the `Transform` class provides several mechanisms that make the process of building them easy and flexible.
### The main `Transform` features:
- **Type dispatch** - Type annotations are used to determine if a transform should be applied to the given argument. It also gives an option to provide several implementations and it choses the one to run based on the type. This is useful for example when running both independent and dependent variables through the pipeline where some transforms only make sense for one and not the other. Another usecase is designing a transform that handles different data formats. Note that if a transform takes multiple arguments only the type of the first one is used for dispatch.
- **Handling of tuples** - When a tuple (or a subclass of tuple) of data is passed to a transform it will get applied to each element separately. You can opt out of this behavior by passing a list or an `L`, as only tuples gets this specific behavior. An alternative is to use `ItemTransform` defined below, which will always take the input as a whole.
- **Reversability** - A transform can be made reversible by implementing the <code>decodes</code> method. This is mainly used to turn something like a category which is encoded as a number back into a label understandable by humans for showing purposes. Like the regular call method, the `decode` method that is used to decode will be applied over each element of a tuple separately.
- **Type propagation** - Whenever possible a transform tries to return data of the same type it received. Mainly used to maintain semantics of things like `ArrayImage` which is a thin wrapper of pytorch's `Tensor`. You can opt out of this behavior by adding `->None` return type annotation.
- **Preprocessing** - The `setup` method can be used to perform any one-time calculations to be later used by the transform, for example generating a vocabulary to encode categorical data.
- **Filtering based on the dataset type** - By setting the `split_idx` flag you can make the transform be used only in a specific `DataSource` subset like in training, but not validation.
- **Ordering** - You can set the `order` attribute which the `Pipeline` uses when it needs to merge two lists of transforms.
- **Appending new behavior with decorators** - You can easily extend an existing `Transform` by creating <code>encodes</code> or <code>decodes</code> methods for new data types. You can put those new methods outside the original transform definition and decorate them with the class you wish them patched into. This can be used by the fastai library users to add their own behavior, or multiple modules contributing to the same transform.
### Defining a `Transform`
There are a few ways to create a transform with different ratios of simplicity to flexibility.
- **Extending the `Transform` class** - Use inheritence to implement the methods you want.
- **Passing methods to the constructor** - Instantiate the `Transform` class and pass your functions as `enc` and `dec` arguments.
- **@Transform decorator** - Turn any function into a `Transform` by just adding a decorator - very straightforward if all you need is a single <code>encodes</code> implementation.
- **Passing a function to fastai APIs** - Same as above, but when passing a function to other transform aware classes like `Pipeline` or `TfmdDS` you don't even need a decorator. Your function will get converted to a `Transform` automatically.
A simple way to create a `Transform` is to pass a function to the constructor. In the below example, we pass an anonymous function that does integer division by 2:
```
f = Transform(lambda o:o//2)
```
If you call this transform, it will apply the transformation:
```
test_eq_type(f(2), 1)
```
Another way to define a Transform is to extend the `Transform` class:
```
class A(Transform): pass
```
However, to enable your transform to do something, you have to define an <code>encodes</code> method. Note that we can use the class name as a decorator to add this method to the original class.
```
@A
def encodes(self, x): return x+1
f1 = A()
test_eq(f1(1), 2) # f1(1) is the same as f1.encode(1)
```
In addition to adding an <code>encodes</code> method, we can also add a <code>decodes</code> method. This enables you to call the `decode` method (without an s). For more information about the purpose of <code>decodes</code>, see the discussion about Reversibility in [the above section](#The-main-Transform-features).
Just like with encodes, you can add a <code>decodes</code> method to the original class by using the class name as a decorator:
```
class B(A): pass
@B
def decodes(self, x): return x-1
f2 = B()
test_eq(f2.decode(2), 1)
test_eq(f2(1), 2) # uses A's encode method from the parent class
```
If you do not define an <code>encodes</code> or <code>decodes</code> method the original value will be returned:
```
class _Tst(Transform): pass
f3 = _Tst() # no encodes or decodes method have been defined
test_eq_type(f3.decode(2.0), 2.0)
test_eq_type(f3(2), 2)
```
#### Defining Transforms With A Decorator
`Transform` can be used as a decorator to turn a function into a `Transform`.
```
@Transform
def f(x): return x//2
test_eq_type(f(2), 1)
test_eq_type(f.decode(2.0), 2.0)
@Transform
def f(x): return x*2
test_eq_type(f(2), 4)
test_eq_type(f.decode(2.0), 2.0)
```
#### Typed Dispatch and Transforms
We can also apply different transformations depending on the type of the input passed by using `TypedDispatch`. `TypedDispatch` automatically works with `Transform` when using type hints:
```
class A(Transform): pass
@A
def encodes(self, x:int): return x//2
@A
def encodes(self, x:float): return x+1
```
When we pass in an `int`, this calls the first encodes method:
```
f = A()
test_eq_type(f(3), 1)
```
When we pass in a `float`, this calls the second encodes method:
```
test_eq_type(f(2.), 3.)
```
When we pass in a type that is not specified in <code>encodes</code>, the original value is returned:
```
test_eq(f('a'), 'a')
```
If the type annotation is a tuple, then any type in the tuple will match:
```
class MyClass(int): pass
class A(Transform):
def encodes(self, x:(MyClass,float)): return x/2
def encodes(self, x:(str,list)): return str(x)+'_1'
f = A()
```
The below two examples match the first encodes, with a type of `MyClass` and `float`, respectively:
```
test_eq(f(MyClass(2)), 1.) # input is of type MyClass
test_eq(f(6.0), 3.0) # input is of type float
```
The next two examples match the second `encodes` method, with a type of `str` and `list`, respectively:
```
test_eq(f('a'), 'a_1') # input is of type str
test_eq(f(['a','b','c']), "['a', 'b', 'c']_1") # input is of type list
```
#### Casting Types With Transform
Without any intervention it is easy for operations to change types in Python. For example, `FloatSubclass` (defined below) becomes a `float` after performing multiplication:
```
class FloatSubclass(float): pass
test_eq_type(FloatSubclass(3.0) * 2, 6.0)
```
This behavior is often not desirable when performing transformations on data. Therefore, `Transform` will attempt to cast the output to be of the same type as the input by default. In the below example, the output will be cast to a `FloatSubclass` type to match the type of the input:
```
@Transform
def f(x): return x*2
test_eq_type(f(FloatSubclass(3.0)), FloatSubclass(6.0))
```
We can optionally turn off casting by annotating the transform function with a return type of `None`:
```
@Transform
def f(x)-> None: return x*2 # Same transform as above, but with a -> None annotation
test_eq_type(f(FloatSubclass(3.0)), 6.0) # Casting is turned off because of -> None annotation
```
However, `Transform` will only cast output back to the input type when the input is a subclass of the output. In the below example, the input is of type `FloatSubclass` which is not a subclass of the output which is of type `str`. Therefore, the output doesn't get cast back to `FloatSubclass` and stays as type `str`:
```
@Transform
def f(x): return str(x)
test_eq_type(f(Float(2.)), '2.0')
```
Just like <code>encodes</code>, the <code>decodes</code> method will cast outputs to match the input type in the same way. In the below example, the output of <code>decodes</code> remains of type `MySubclass`:
```
class MySubclass(int): pass
def enc(x): return MySubclass(x+1)
def dec(x): return x-1
f = Transform(enc,dec)
t = f(1) # t is of type MySubclass
test_eq_type(f.decode(t), MySubclass(1)) # the output of decode is cast to MySubclass to match the input type.
```
#### Apply Transforms On Subsets With `split_idx`
You can apply transformations to subsets of data by specifying a `split_idx` property. If a transform has a `split_idx` then it's only applied if the `split_idx` param matches. In the below example, we set `split_idx` equal to `1`:
```
def enc(x): return x+1
def dec(x): return x-1
f = Transform(enc,dec)
f.split_idx = 1
```
The transformations are applied when a matching `split_idx` parameter is passed:
```
test_eq(f(1, split_idx=1),2)
test_eq(f.decode(2, split_idx=1),1)
```
On the other hand, transformations are ignored when the `split_idx` parameter does not match:
```
test_eq(f(1, split_idx=0), 1)
test_eq(f.decode(2, split_idx=0), 2)
```
#### Transforms on Lists
Transform operates on lists as a whole, **not element-wise**:
```
class A(Transform):
def encodes(self, x): return dict(x)
def decodes(self, x): return list(x.items())
f = A()
_inp = [(1,2), (3,4)]
t = f(_inp)
test_eq(t, dict(_inp))
test_eq(f.decodes(t), _inp)
#hide
f.split_idx = 1
test_eq(f(_inp, split_idx=1), dict(_inp))
test_eq(f(_inp, split_idx=0), _inp)
```
If you want a transform to operate on a list elementwise, you must implement this appropriately in the <code>encodes</code> and <code>decodes</code> methods:
```
class AL(Transform): pass
@AL
def encodes(self, x): return [x_+1 for x_ in x]
@AL
def decodes(self, x): return [x_-1 for x_ in x]
f = AL()
t = f([1,2])
test_eq(t, [2,3])
test_eq(f.decode(t), [1,2])
```
#### Transforms on Tuples
Unlike lists, `Transform` operates on tuples element-wise.
```
def neg_int(x): return -x
f = Transform(neg_int)
test_eq(f((1,2,3)), (-1,-2,-3))
```
Transforms will also apply `TypedDispatch` element-wise on tuples when an input type annotation is specified. In the below example, the values `1.0` and `3.0` are ignored because they are of type `float`, not `int`:
```
def neg_int(x:int): return -x
f = Transform(neg_int)
test_eq(f((1.0, 2, 3.0)), (1.0, -2, 3.0))
#hide
test_eq(f((1,)), (-1,))
test_eq(f((1.,)), (1.,))
test_eq(f.decode((1,2)), (1,2))
test_eq(f.input_types, int)
```
Another example of how `Transform` can use `TypedDispatch` with tuples is shown below:
```
class B(Transform): pass
@B
def encodes(self, x:int): return x+1
@B
def encodes(self, x:str): return x+'hello'
@B
def encodes(self, x)->None: return str(x)+'!'
```
If the input is not an `int` or `str`, the third `encodes` method will apply:
```
b = B()
test_eq(b([1]), '[1]!')
test_eq(b([1.0]), '[1.0]!')
```
However, if the input is a tuple, then the appropriate method will apply according to the type of each element in the tuple:
```
test_eq(b(('1',)), ('1hello',))
test_eq(b((1,2)), (2,3))
test_eq(b(('a',1.0)), ('ahello','1.0!'))
#hide
@B
def decodes(self, x:int): return x-1
test_eq(b.decode((2,)), (1,))
test_eq(b.decode(('2',)), ('2',))
assert pickle.loads(pickle.dumps(b))
```
Dispatching over tuples works recursively, by the way:
```
class B(Transform):
def encodes(self, x:int): return x+1
def encodes(self, x:str): return x+'_hello'
def decodes(self, x:int): return x-1
def decodes(self, x:str): return x.replace('_hello', '')
f = B()
start = (1.,(2,'3'))
t = f(start)
test_eq_type(t, (1.,(3,'3_hello')))
test_eq(f.decode(t), start)
```
Dispatching also works with `typing` module type classes, like `numbers.integral`:
```
@Transform
def f(x:numbers.Integral): return x+1
t = f((1,'1',1))
test_eq(t, (2, '1', 2))
#export
class InplaceTransform(Transform):
"A `Transform` that modifies in-place and just returns whatever it's passed"
def _call(self, fn, x, split_idx=None, **kwargs):
super()._call(fn,x,split_idx,**kwargs)
return x
#hide
import pandas as pd
class A(InplaceTransform): pass
@A
def encodes(self, x:pd.Series): x.fillna(10, inplace=True)
f = A()
test_eq_type(f(pd.Series([1,2,None])),pd.Series([1,2,10],dtype=np.float64)) #fillna fills with floats.
# export
class DisplayedTransform(Transform):
"A transform with a `__repr__` that shows its attrs"
@property
def name(self): return f"{super().name} -- {getattr(self,'__stored_args__',{})}"
```
Transforms normally are represented by just their class name and a list of encodes and decodes implementations:
```
class A(Transform): encodes,decodes = noop,noop
f = A()
f
```
A `DisplayedTransform` will in addition show the contents of all attributes listed in the comma-delimited string `self.store_attrs`:
```
class A(DisplayedTransform):
encodes = noop
def __init__(self, a, b=2):
super().__init__()
store_attr()
A(a=1,b=2)
```
#### ItemTransform -
```
#export
class ItemTransform(Transform):
"A transform that always take tuples as items"
_retain = True
def __call__(self, x, **kwargs): return self._call1(x, '__call__', **kwargs)
def decode(self, x, **kwargs): return self._call1(x, 'decode', **kwargs)
def _call1(self, x, name, **kwargs):
if not _is_tuple(x): return getattr(super(), name)(x, **kwargs)
y = getattr(super(), name)(list(x), **kwargs)
if not self._retain: return y
if is_listy(y) and not isinstance(y, tuple): y = tuple(y)
return retain_type(y, x)
```
`ItemTransform` is the class to use to opt out of the default behavior of `Transform`.
```
class AIT(ItemTransform):
def encodes(self, xy): x,y=xy; return (x+y,y)
def decodes(self, xy): x,y=xy; return (x-y,y)
f = AIT()
test_eq(f((1,2)), (3,2))
test_eq(f.decode((3,2)), (1,2))
```
If you pass a special tuple subclass, the usual retain type behavior of `Transform` will keep it:
```
class _T(tuple): pass
x = _T((1,2))
test_eq_type(f(x), _T((3,2)))
#hide
f.split_idx = 0
test_eq_type(f((1,2)), (1,2))
test_eq_type(f((1,2), split_idx=0), (3,2))
test_eq_type(f.decode((1,2)), (1,2))
test_eq_type(f.decode((3,2), split_idx=0), (1,2))
#hide
class Get(ItemTransform):
_retain = False
def encodes(self, x): return x[0]
g = Get()
test_eq(g([1,2,3]), 1)
test_eq(g(L(1,2,3)), 1)
test_eq(g(np.array([1,2,3])), 1)
test_eq_type(g((['a'], ['b', 'c'])), ['a'])
#hide
class A(ItemTransform):
def encodes(self, x): return _T((x,x))
def decodes(self, x): return _T(x)
f = A()
test_eq(type(f.decode((1,1))), _T)
```
### Func -
```
#export
def get_func(t, name, *args, **kwargs):
"Get the `t.name` (potentially partial-ized with `args` and `kwargs`) or `noop` if not defined"
f = getattr(t, name, noop)
return f if not (args or kwargs) else partial(f, *args, **kwargs)
```
This works for any kind of `t` supporting `getattr`, so a class or a module.
```
test_eq(get_func(operator, 'neg', 2)(), -2)
test_eq(get_func(operator.neg, '__call__')(2), -2)
test_eq(get_func(list, 'foobar')([2]), [2])
a = [2,1]
get_func(list, 'sort')(a)
test_eq(a, [1,2])
```
Transforms are built with multiple-dispatch: a given function can have several methods depending on the type of the object received. This is done directly with the `TypeDispatch` module and type-annotation in `Transform`, but you can also use the following class.
```
#export
class Func():
"Basic wrapper around a `name` with `args` and `kwargs` to call on a given type"
def __init__(self, name, *args, **kwargs): self.name,self.args,self.kwargs = name,args,kwargs
def __repr__(self): return f'sig: {self.name}({self.args}, {self.kwargs})'
def _get(self, t): return get_func(t, self.name, *self.args, **self.kwargs)
def __call__(self,t): return mapped(self._get, t)
```
You can call the `Func` object on any module name or type, even a list of types. It will return the corresponding function (with a default to `noop` if nothing is found) or list of functions.
```
test_eq(Func('sqrt')(math), math.sqrt)
#export
class _Sig():
def __getattr__(self,k):
def _inner(*args, **kwargs): return Func(k, *args, **kwargs)
return _inner
Sig = _Sig()
show_doc(Sig, name="Sig")
```
`Sig` is just sugar-syntax to create a `Func` object more easily with the syntax `Sig.name(*args, **kwargs)`.
```
f = Sig.sqrt()
test_eq(f(math), math.sqrt)
```
## Pipeline -
```
#export
def compose_tfms(x, tfms, is_enc=True, reverse=False, **kwargs):
"Apply all `func_nm` attribute of `tfms` on `x`, maybe in `reverse` order"
if reverse: tfms = reversed(tfms)
for f in tfms:
if not is_enc: f = f.decode
x = f(x, **kwargs)
return x
def to_int (x): return Int(x)
def to_float(x): return Float(x)
def double (x): return x*2
def half(x)->None: return x/2
def test_compose(a, b, *fs): test_eq_type(compose_tfms(a, tfms=map(Transform,fs)), b)
test_compose(1, Int(1), to_int)
test_compose(1, Float(1), to_int,to_float)
test_compose(1, Float(2), to_int,to_float,double)
test_compose(2.0, 2.0, to_int,double,half)
class A(Transform):
def encodes(self, x:float): return Float(x+1)
def decodes(self, x): return x-1
tfms = [A(), Transform(math.sqrt)]
t = compose_tfms(3., tfms=tfms)
test_eq_type(t, Float(2.))
test_eq(compose_tfms(t, tfms=tfms, is_enc=False), 1.)
test_eq(compose_tfms(4., tfms=tfms, reverse=True), 3.)
tfms = [A(), Transform(math.sqrt)]
test_eq(compose_tfms((9,3.), tfms=tfms), (3,2.))
#export
def mk_transform(f):
"Convert function `f` to `Transform` if it isn't already one"
f = instantiate(f)
return f if isinstance(f,(Transform,Pipeline)) else Transform(f)
#export
def gather_attrs(o, k, nm):
"Used in __getattr__ to collect all attrs `k` from `self.{nm}`"
if k.startswith('_') or k==nm: raise AttributeError(k)
att = getattr(o,nm)
res = [t for t in att.attrgot(k) if t is not None]
if not res: raise AttributeError(k)
return res[0] if len(res)==1 else L(res)
#export
def gather_attr_names(o, nm):
"Used in __dir__ to collect all attrs `k` from `self.{nm}`"
return L(getattr(o,nm)).map(dir).concat().unique()
#export
class Pipeline:
"A pipeline of composed (for encode/decode) transforms, setup with types"
def __init__(self, funcs=None, split_idx=None):
self.split_idx,self.default = split_idx,None
if funcs is None: funcs = []
if isinstance(funcs, Pipeline): self.fs = funcs.fs
else:
if isinstance(funcs, Transform): funcs = [funcs]
self.fs = L(ifnone(funcs,[noop])).map(mk_transform).sorted(key='order')
for f in self.fs:
name = camel2snake(type(f).__name__)
a = getattr(self,name,None)
if a is not None: f = L(a)+f
setattr(self, name, f)
def setup(self, items=None, train_setup=False):
tfms = self.fs[:]
self.fs.clear()
for t in tfms: self.add(t,items, train_setup)
def add(self,t, items=None, train_setup=False):
t.setup(items, train_setup)
self.fs.append(t)
def __call__(self, o): return compose_tfms(o, tfms=self.fs, split_idx=self.split_idx)
def __repr__(self): return f"Pipeline: {' -> '.join([f.name for f in self.fs if f.name != 'noop'])}"
def __getitem__(self,i): return self.fs[i]
def __setstate__(self,data): self.__dict__.update(data)
def __getattr__(self,k): return gather_attrs(self, k, 'fs')
def __dir__(self): return super().__dir__() + gather_attr_names(self, 'fs')
def decode (self, o, full=True):
if full: return compose_tfms(o, tfms=self.fs, is_enc=False, reverse=True, split_idx=self.split_idx)
#Not full means we decode up to the point the item knows how to show itself.
for f in reversed(self.fs):
if self._is_showable(o): return o
o = f.decode(o, split_idx=self.split_idx)
return o
def show(self, o, ctx=None, **kwargs):
o = self.decode(o, full=False)
o1 = (o,) if not _is_tuple(o) else o
if hasattr(o, 'show'): ctx = o.show(ctx=ctx, **kwargs)
else:
for o_ in o1:
if hasattr(o_, 'show'): ctx = o_.show(ctx=ctx, **kwargs)
return ctx
def _is_showable(self, o):
if hasattr(o, 'show'): return True
if _is_tuple(o): return all(hasattr(o_, 'show') for o_ in o)
return False
add_docs(Pipeline,
__call__="Compose `__call__` of all `fs` on `o`",
decode="Compose `decode` of all `fs` on `o`",
show="Show `o`, a single item from a tuple, decoding as needed",
add="Add transform `t`",
setup="Call each tfm's `setup` in order")
```
`Pipeline` is a wrapper for `compose_tfm`. You can pass instances of `Transform` or regular functions in `funcs`, the `Pipeline` will wrap them all in `Transform` (and instantiate them if needed) during the initialization. It handles the transform `setup` by adding them one at a time and calling setup on each, goes through them in order in `__call__` or `decode` and can `show` an object by applying decoding the transforms up until the point it gets an object that knows how to show itself.
```
# Empty pipeline is noop
pipe = Pipeline()
test_eq(pipe(1), 1)
test_eq(pipe((1,)), (1,))
# Check pickle works
assert pickle.loads(pickle.dumps(pipe))
class IntFloatTfm(Transform):
def encodes(self, x): return Int(x)
def decodes(self, x): return Float(x)
foo=1
int_tfm=IntFloatTfm()
def neg(x): return -x
neg_tfm = Transform(neg, neg)
pipe = Pipeline([neg_tfm, int_tfm])
start = 2.0
t = pipe(start)
test_eq_type(t, Int(-2))
test_eq_type(pipe.decode(t), Float(start))
test_stdout(lambda:pipe.show(t), '-2')
pipe = Pipeline([neg_tfm, int_tfm])
t = pipe(start)
test_stdout(lambda:pipe.show(pipe((1.,2.))), '-1\n-2')
test_eq(pipe.foo, 1)
assert 'foo' in dir(pipe)
assert 'int_float_tfm' in dir(pipe)
```
Transforms are available as attributes named with the snake_case version of the names of their types. Attributes in transforms can be directly accessed as attributes of the pipeline.
```
test_eq(pipe.int_float_tfm, int_tfm)
test_eq(pipe.foo, 1)
pipe = Pipeline([int_tfm, int_tfm])
pipe.int_float_tfm
test_eq(pipe.int_float_tfm[0], int_tfm)
test_eq(pipe.foo, [1,1])
# Check opposite order
pipe = Pipeline([int_tfm,neg_tfm])
t = pipe(start)
test_eq(t, -2)
test_stdout(lambda:pipe.show(t), '-2')
class A(Transform):
def encodes(self, x): return int(x)
def decodes(self, x): return Float(x)
pipe = Pipeline([neg_tfm, A])
t = pipe(start)
test_eq_type(t, -2)
test_eq_type(pipe.decode(t), Float(start))
test_stdout(lambda:pipe.show(t), '-2.0')
s2 = (1,2)
pipe = Pipeline([neg_tfm, A])
t = pipe(s2)
test_eq_type(t, (-1,-2))
test_eq_type(pipe.decode(t), (Float(1.),Float(2.)))
test_stdout(lambda:pipe.show(t), '-1.0\n-2.0')
from PIL import Image
class ArrayImage(ndarray):
_show_args = {'cmap':'viridis'}
def __new__(cls, x, *args, **kwargs):
if isinstance(x,tuple): super().__new__(cls, x, *args, **kwargs)
if args or kwargs: raise RuntimeError('Unknown array init args')
if not isinstance(x,ndarray): x = array(x)
return x.view(cls)
def show(self, ctx=None, figsize=None, **kwargs):
if ctx is None: _,ctx = plt.subplots(figsize=figsize)
ctx.imshow(im, **{**self._show_args, **kwargs})
ctx.axis('off')
return ctx
im = Image.open(TEST_IMAGE)
im_t = ArrayImage(im)
def f1(x:ArrayImage): return -x
def f2(x): return Image.open(x).resize((128,128))
def f3(x:Image.Image): return(ArrayImage(array(x)))
pipe = Pipeline([f2,f3,f1])
t = pipe(TEST_IMAGE)
test_eq(type(t), ArrayImage)
test_eq(t, -array(f3(f2(TEST_IMAGE))))
pipe = Pipeline([f2,f3])
t = pipe(TEST_IMAGE)
ax = pipe.show(t)
test_fig_exists(ax)
#Check filtering is properly applied
add1 = B()
add1.split_idx = 1
pipe = Pipeline([neg_tfm, A(), add1])
test_eq(pipe(start), -2)
pipe.split_idx=1
test_eq(pipe(start), -1)
pipe.split_idx=0
test_eq(pipe(start), -2)
for t in [None, 0, 1]:
pipe.split_idx=t
test_eq(pipe.decode(pipe(start)), start)
test_stdout(lambda: pipe.show(pipe(start)), "-2.0")
def neg(x): return -x
test_eq(type(mk_transform(neg)), Transform)
test_eq(type(mk_transform(math.sqrt)), Transform)
test_eq(type(mk_transform(lambda a:a*2)), Transform)
test_eq(type(mk_transform(Pipeline([neg]))), Pipeline)
```
### Methods
```
#TODO: method examples
show_doc(Pipeline.__call__)
show_doc(Pipeline.decode)
show_doc(Pipeline.setup)
```
During the setup, the `Pipeline` starts with no transform and adds them one at a time, so that during its setup, each transform gets the items processed up to its point and not after.
```
#hide
#Test is with TfmdList
```
## Export -
```
#hide
from nbdev.export import notebook2script
notebook2script()
```
| github_jupyter |
# Using Linux Foundation Delta Lake in Azure Synapse Analytics Spark
Azure Synapse is compatible with Linux Foundation Delta Lake. Delta Lake is an open-source storage layer that brings ACID (atomicity, consistency, isolation, and durability) transactions to Apache Spark and big data workloads.
This notebook provides examples of how to update, merge and delete delta lake tables in Synapse.
## Pre-requisites
In this notebook you will save your tables in Delta Lake format to your workspace's primary storage account. You are required to be a **Blob Storage Contributor** in the ADLS Gen2 account (or folder) you will access.
## Load sample data
First you will load the [public holidays](https://azure.microsoft.com/en-us/services/open-datasets/catalog/public-holidays/) data from last 6 months via Azure Open datasets.
```
from azureml.opendatasets import PublicHolidays
from datetime import datetime
from dateutil import parser
from dateutil.relativedelta import relativedelta
end_date = datetime.today()
start_date = datetime.today() - relativedelta(months=6)
hol = PublicHolidays(start_date=start_date, end_date=end_date)
hol_df = hol.to_spark_dataframe()
display(hol_df)
```
## Write data to the Delta Lake table
```
# Set the strorage path info
# Primary storage info
account_name = '' # fill in your primary storage account name
container_name = '' # fill in your container name
relative_path = '' # fill in your relative folder path
adls_path = 'abfss://%s@%s.dfs.core.windows.net/%s' % (container_name, account_name, relative_path)
print('Primary storage account path: ' + adls_path)
# Delta Lake relative path
delta_relative_path = adls_path + 'delta/holiday/'
print('Delta Lake path: ' + delta_relative_path)
# Filter out indian holidays
hol_df_IN = hol_df[(hol_df.countryRegionCode == "IN")]
hol_df_IN.show(5, truncate = False)
#Let's write the data in the Delta Lake table.
hol_df_IN.write.mode("overwrite").format("delta").partitionBy("holidayName").save(delta_relative_path)
delta_data = spark.read.format("delta").load(delta_relative_path)
delta_data.show()
```
## Overwrite the entire Delta Lake table
```
#Let's overwrite the entire delta file with 1 record
hol_df_JP= hol_df[(hol_df.countryRegionCode == "JP")]
hol_df_JP.write.format("delta").mode("overwrite").save(delta_relative_path)
delta_data = spark.read.format("delta").load(delta_relative_path)
delta_data.show()
```
## Merge new data based on given merge condition
```
# Upsert (merge) the United States' holiday data with Japan's
from delta.tables import *
deltaTable = DeltaTable.forPath(spark,delta_relative_path)
hol_df_US= hol_df[(hol_df.countryRegionCode == "US")]
deltaTable.alias("hol_df_JP").merge(
source = hol_df_US.alias("hol_df_US"),
condition = "hol_df_JP.countryRegionCode = hol_df_US.countryRegionCode"
).whenMatchedUpdate(set =
{}).whenNotMatchedInsert( values =
{
"countryOrRegion" : "hol_df_US.countryOrRegion",
"holidayName" : "hol_df_US.holidayName",
"normalizeHolidayName" : "hol_df_US.normalizeHolidayName",
"isPaidTimeOff":"hol_df_US.isPaidTimeOff",
"countryRegionCode":"hol_df_US.countryRegionCode",
"date":"hol_df_US.date"
}
).execute()
deltaTable.toDF().show()
```
## Update table on the rows that match the given condition
```
# Update column the 'null' value in 'isPaidTimeOff' with 'false'
from pyspark.sql.functions import *
deltaTable.update(
condition = (col("isPaidTimeOff").isNull()),
set = {"isPaidTimeOff": "false"})
deltaTable.toDF().show()
```
## Delete data from the table that match the given condition
```
print("Row count before delete: ")
print(deltaTable.toDF().count())
# Delte data with date later than 2020-01-01
deltaTable.delete ("date > '2020-01-01'")
print("Row count after delete: ")
print(deltaTable.toDF().count())
deltaTable.toDF().show()
```
## Get the operation history of the delta table
```
fullHistoryDF = deltaTable.history()
lastOperationDF = deltaTable.history(1)
print('Full history DF: ')
fullHistoryDF.show(truncate = False)
print('lastOperationDF: ')
lastOperationDF.show(truncate = False)
```
| github_jupyter |
```
import os
import pandas as pd
import numpy as np
from scipy.io import arff
os.system("du -h ./")
#UEA-MTS archive datasets:
"ArticularyWordRecognition",
"AtrialFibrillation",
"BasicMotions",
"CharacterTrajectories",
"Cricket",
"DuckDuckGeese",#5
"EigenWorms",
"Epilepsy",
"ERing",
"EthanolConcentration",
"FaceDetection",#10
"FingerMovements",
"HandMovementDirection",
"Handwriting",
"Heartbeat",
"InsectWingbeat",#15
"JapaneseVowels",
"Libras",
"LSST",
"MotorImagery",
"NATOPS",#20
"PEMS-SF",
"PenDigits",
"PhonemeSpectra",
"RacketSports",
"SelfRegulationSCP1",#25
"SelfRegulationSCP2",
"SpokenArabicDigits",
"StandWalkJump",
"UWaveGestureLibrary"
rep = './'
UEA_MTS_List = [
"PEMS-SF"
]
```
### Convert UEA '.arff' format to SMATE format
```
# convert UEA '.arff' format to meta file and data file containing one single instance
def conv_UEA_SMATE(dataset_names):
for name in dataset_names:
dict = rep + name + '/'
file_train = dict + name + '_TRAIN.arff'
file_test = dict + name + '_TEST.arff'
dict_train_out = dict + 'output_train/'
dict_test_out = dict + 'output_test/'
os.system("mkdir " + dict_train_out)
os.system("mkdir " + dict_test_out)
convert_arff_samples(file_train, dict_train_out)
convert_arff_samples(file_test, dict_test_out)
def convert_arff_samples(file, dict_out):
data = arff.loadarff(file) #load 'arff' files
df = pd.DataFrame(data[0])
#df['input'] = df.iloc[:,0]
df.insert(loc=0, column='d_input', value=df.iloc[:,0]) #standardize the input attribute name
df = df.drop(df.columns[1], axis=1)
df['d_class'] = df.iloc[:,-1] #standardize the class attribute name
df = df.drop(df.columns[-2], axis=1)
df['sample_id'] = df.index # add index as sample_id
df['d_class'] = df['d_class'].apply(lambda x: x.decode("utf-8") ) # convert Byte to String for class column
df_meta = df[['sample_id', 'd_class']]
df_meta.to_csv(dict_out + 'meta_data.csv', index=False, header=None) #save meta_data.csv
for name, sample in df.groupby(['sample_id']):
df_sample = pd.DataFrame(sample['d_input'].values.tolist()).add_prefix('dimension_')
df_sample_conv = pd.DataFrame()
for c in df_sample.columns:
df_sample_conv[[c]] = pd.DataFrame(df_sample[c].values[0].tolist())
df_sample_conv.to_csv(dict_out + str(name) + '.csv', index=False, header=None) #save samples into individual files
conv_UEA_SMATE(UEA_MTS_List)
```
### Convert UEA '.arff' format to NMSU'IJCAI'20 format (i.e., CA-SFCN)
```
# convert UEA '.arff' format to NMSU_IJCAI'20 input format
'''
output: N * (D * L)
'''
def conv_UEA_NMSU(dataset_names):
for name in dataset_names:
dict = rep + name + '/'
file_train = dict + name + '_TRAIN.arff'
file_test = dict + name + '_TEST.arff'
dict_out = 'uea_nmsu' + '/' + dict
os.makedirs(dict_out, exist_ok=True)
X_train, y_train, map_c_l = convert_arff_nmsu(file_train, dict_out, 'train' )
X_test, y_test = convert_arff_nmsu(file_test, dict_out, 'test', map_c_l)
x_row, attr_num, attr_len = X_train.shape
X_train = X_train.reshape(x_row, (attr_num*attr_len))
x_row, attr_num, attr_len = X_test.shape
X_test = X_test.reshape(x_row, (attr_num*attr_len))
file_writingxy(X_train, y_train, dict_out + "train.txt", attr_num)
file_writingxy(X_test, y_test, dict_out + "test.txt", attr_num)
#return X_train
def convert_arff_nmsu(file, dict_out, out_name, map_c_l = {}):
data = arff.loadarff(file) #load 'arff' files
df = pd.DataFrame(data[0])
df.insert(loc=0, column='d_input', value=df.iloc[:,0]) #standardize the input attribute name
df = df.drop(df.columns[1], axis=1)
df['d_class'] = df.iloc[:,-1] #standardize the class attribute name
df = df.drop(df.columns[-2], axis=1)
df['d_class'] = df['d_class'].apply(lambda x: x.decode("utf-8") ) # convert Byte to String for class column
df = df[['d_class', 'd_input']]
def convert_D_L(x):
x_transpose = [[row[i] for row in x] for i in range(len(x[0]))]
x_arr = np.transpose(np.asarray(x_transpose))
return x_arr
df['d_input'] = df['d_input'].map(convert_D_L)
x = df['d_input']
print(x.shape)
x_matrix = np.zeros([len(x), x[0].shape[0], x[0].shape[1]])
print(x_matrix.shape)
for i in range(len(x)):
x_matrix[i,:,:] = x[i]
y_vector = np.array(df['d_class'])
if out_name == 'train':
map_c_l = get_label_map(y_vector)
#print("map_c_l is ", map_c_l)
y_num = np.zeros(y_vector.shape[0])
for idx, y in enumerate(y_vector):
#print("y is ", y)
y_num[idx] = map_c_l[y]
y_num = y_num.reshape(-1, 1)
return np.nan_to_num(x_matrix), np.nan_to_num(y_num), map_c_l
else:
y_num = np.zeros(y_vector.shape[0])
for idx, y in enumerate(y_vector):
y_num[idx] = map_c_l[y]
y_num = y_num.reshape(-1, 1)
return np.nan_to_num(x_matrix), np.nan_to_num(y_num)
def file_writingxy(data_x_matrix, data_y_vector, file_name, attr_num=-1, delimiter=' '):
data_row, data_col = data_x_matrix.shape
with open(file_name, 'w') as f:
if attr_num > 0:
f.write(str(int(attr_num)) + '\n')
for row in range(0, data_row):
row_vector = data_x_matrix[row, :]
row_label = str(int(float(data_y_vector[row])))
row_str = row_label
for index in range(0, data_col):
row_str = row_str + delimiter + str(row_vector[index])
f.write(row_str + '\n')
def get_label_map(y_train):
'''
Input:
- y_train: a vector (n, )
Output:
- mapping_c_l: dict {label: number}, number is in [0, n_class - 1]
'''
No = len(y_train)
classes, counts_cl = np.unique(y_train, return_counts=True)
print("class list is " + str(classes))
mapping_c_l = {} # a mappling between classes and labels
for idx, c in enumerate(list(classes)):
mapping_c_l.update({c: idx})
return mapping_c_l
conv_UEA_NMSU(UEA_MTS_List)
import sys, scipy
print(sys.version)
print (scipy.__version__)
```
| github_jupyter |
## Homework 2
## Practice Linear Regression and Hyperparameter Search
This assignment is aimed to help you get more experience with [linear models](https://scikit-learn.org/stable/modules/linear_model.html) (especially linear regression) and [hyperparameter search](https://scikit-learn.org/stable/model_selection.html) in scikit-learn library.
```
import matplotlib.pyplot as plt
import numpy as np
```
# Today's data
400 fotos of human faces. Each face is a 2d array [64x64] of pixel brightness.
```
from sklearn.datasets import fetch_olivetti_faces
data = fetch_olivetti_faces().images
```
Let's see some faces
```
# this code showcases matplotlib subplots.
fig, ax = plt.subplots(2, 2, figsize=(12, 12))
ax = ax.flatten()
for i in range(4):
ax[i].imshow(data[i], cmap="gray")
plt.show()
```
# Face reconstruction problem
Let's solve the face reconstruction problem: given left halves of facex __(X)__, our algorithm shall predict the right half __(y)__. The idea of this approach is that left face half actually contains quite enough information to reconstruct the right face half (at least partially). Moreover in this task we'll also see, that scikit-learn linear models are capable of predicting multiple targets for a single object example.
Our first step is to slice the photos into X and y using slices.
__Slices in numpy:__
* In regular python, slice looks roughly like this: `a[2:5]` _(select elements from 2 to 5)_
* Numpy allows you to slice N-dimensional arrays along each dimension: [image_index, height, width]
* `data[:10]` - Select first 10 images
* `data[:, :10]` - For all images, select a horizontal stripe 10 pixels high at the top of the image
* `data[10:20, :, -25:-15]` - Take images [10, 11, ..., 19], for each image select a _vetrical stripe_ of width 10 pixels, 15 pixels away from the _right_ side.
__Your task:__
Let's use slices to select all __left image halves as X__ and all __right halves as y__.
```
# select left half of each face as X, right half as Y
X = <Slice left half-images>
y = <Slice right half-images>
# If you did everything right, you're gonna see left half-image and right half-image drawn separately in natural order
plt.subplot(1, 2, 1)
plt.imshow(X[0], cmap="gray")
plt.subplot(1, 2, 2)
plt.imshow(y[0], cmap="gray")
assert (
X.shape == y.shape == (len(data), 64, 32)
), "Please slice exactly the left half-face to X and right half-face to Y"
def glue(left_half, right_half):
# merge photos back together
left_half = left_half.reshape([-1, 64, 32])
right_half = right_half.reshape([-1, 64, 32])
return np.concatenate([left_half, right_half], axis=-1)
# if you did everything right, you're gonna see a valid face
plt.imshow(glue(X, y)[99], cmap="gray")
```
# Machine learning stuff
```
from sklearn.model_selection import train_test_split
X_train, X_test, Y_train, Y_test = train_test_split(
X.reshape([len(X), -1]), y.reshape([len(y), -1]), test_size=0.05, random_state=42
)
print(X_test.shape)
from sklearn.linear_model import LinearRegression
model = LinearRegression()
model.fit(X_train, Y_train)
```
measure [mean squared error](https://en.wikipedia.org/wiki/Mean_squared_error):
$$MSE(\widehat{\theta}) = \mathbf{E}_{\theta}[(\theta - \widehat{\theta})^2] $$
```
from sklearn.metrics import mean_squared_error
mse_train = mean_squared_error(Y_train, model.predict(X_train))
mae_test = mean_squared_error(Y_test, model.predict(X_test))
print(f"Train MSE: {mse_train:.3f}")
print(f"Test MSE: {mse_train:.3f}")
```
---
## Why train error is much smaller than test?
```
# Train predictions
pics = <YOUR CODE> # reconstruct and glue together X and Y for the train dataset
plt.figure(figsize=[16, 12])
for i in range(20):
plt.subplot(4, 5, i + 1)
plt.imshow(pics[i], cmap='gray')
# Test predictions
pics = <YOUR CODE> # reconstruct and glue together X and Y for the test dataset
plt.figure(figsize=[16, 12])
for i in range(20):
plt.subplot(4, 5, i + 1)
plt.imshow(pics[i], cmap='gray')
```
---
Remember regularisation? That is exactly what we need. There are many many linear models in sklearn package, and all of them can be found [here](https://scikit-learn.org/stable/modules/linear_model.html). We will focus on 3 of them: Ridge regression, Lasso and ElasticNet.
Idea of all of them is very simple: Add some penalty to the objective loss function to prevent overfitting.
# Ridge regression
[RidgeRegression](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.Ridge.html) is just a LinearRegression, with l2 regularization - penalized for $ \alpha \cdot \sum _i w_i^2$
Let's train such a model with alpha=0.5
```
from <YOUR CODE> import <YOUR CODE>
ridge = <YOUR CODE>
<YOUR CODE: fit the model on training set>
<YOUR CODE: predict and measure MSE on train and test>
# Test predictions
pics = <YOUR CODE> # reconstruct and glue together X and Y for the test dataset
plt.figure(figsize=[16, 12])
for i in range(20):
plt.subplot(4, 5, i + 1)
plt.imshow(pics[i], cmap='gray')
```
---
# Grid search
Train model with diferent $\alpha$ and find one that has minimal test MSE. It's okay to use loops or any other python stuff here.
```
from sklearn.model_selection import GridSearchCV
def train_and_plot(model, parameter_dict):
"""This function takes a model and parameters
dict as input and plot a graph of MSE loss VS parameter value"""
# use GridSearchCV as before to do grid search
gscv = GridSearchCV(<Your code>)
<Fit your model>
plt.errorbar(gscv.param_grid['alpha'],
gscv.cv_results_['mean_test_score'],
gscv.cv_results_['std_test_score'],
capsize=5, label=model.__str__().split("(")[0])
plt.xscale("log", nonposx='clip')
plt.xlabel("alpha")
plt.ylabel("negative MSE")
plt.grid()
plt.legend()
plt.figure(figsize=(12, 6))
models = <YOUR CODE> # Start from Ridge regression, but feel free to add
# Lasso and ElasticNet. Note that the latter two cannot
# be solved analytically and typically are much slower
# to fit than Ridge regression (so you may want to limit
# the number of grid points).
parameter_dict = <YOUR CODE>
train_and_plot(models, parameter_dict)
```
---
```
# Test predictions
pics = glue(X_test, <predict with your best model>)
plt.figure(figsize=[16, 12])
for i in range(20):
plt.subplot(4, 5, i + 1)
plt.imshow(pics[i], cmap='gray')
from sklearn.linear_model import Lasso, ElasticNet
# Use the code you have just done to do GridSearch for Lasso and/or ElasticNet
# models (if you haven't already). Note that Lasso and ElasticNet are much
# slower to fit, compared to Ridge.
<YOUR CODE>
```
---
## Bonus part
Try using `sklearn.linear_model.SGDRegressor` with `huber` loss in the code above instead of `LinearRegression`. Is it better in this case?
```
<Your code for bonus part>
```
P.S. This assignment is inspired by [YSDA materials](https://github.com/yandexdataschool).
| github_jupyter |
# Estimating The Mortality Rate For COVID-19
> Using Country-Level Covariates To Correct For Testing & Reporting Biases And Estimate a True Mortality Rate.
- author: Joseph Richards
- image: images/corvid-mortality.png
- comments: true
- categories: [MCMC, mortality]
- permalink: /covid-19-mortality-estimation/
- toc: true
```
#hide
# ! pip install pymc3 arviz xlrd
#hide
# Setup and imports
%matplotlib inline
import warnings
warnings.simplefilter('ignore')
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import pymc3 as pm
from IPython.display import display, Markdown
#hide
# constants
ignore_countries = [
'Others',
'Cruise Ship'
]
cpi_country_mapping = {
'United States of America': 'US',
'China': 'Mainland China'
}
wb_country_mapping = {
'United States': 'US',
'Egypt, Arab Rep.': 'Egypt',
'Hong Kong SAR, China': 'Hong Kong',
'Iran, Islamic Rep.': 'Iran',
'China': 'Mainland China',
'Russian Federation': 'Russia',
'Slovak Republic': 'Slovakia',
'Korea, Rep.': 'Korea, South'
}
wb_covariates = [
('SH.XPD.OOPC.CH.ZS',
'healthcare_oop_expenditure'),
('SH.MED.BEDS.ZS',
'hospital_beds'),
('HD.HCI.OVRL',
'hci'),
('SP.POP.65UP.TO.ZS',
'population_perc_over65'),
('SP.RUR.TOTL.ZS',
'population_perc_rural')
]
#hide
# data loading and manipulation
from datetime import datetime
import os
import numpy as np
import pandas as pd
def get_all_data():
'''
Main routine that grabs all COVID and covariate data and
returns them as a single dataframe that contains:
* count of cumulative cases and deaths by country (by today's date)
* days since first case for each country
* CPI gov't transparency index
* World Bank data on population, healthcare, etc. by country
'''
all_covid_data = _get_latest_covid_timeseries()
covid_cases_rollup = _rollup_by_country(all_covid_data['confirmed'])
covid_deaths_rollup = _rollup_by_country(all_covid_data['deaths'])
todays_date = covid_cases_rollup.columns.max()
# Create DataFrame with today's cumulative case and death count, by country
df_out = pd.DataFrame({'cases': covid_cases_rollup[todays_date],
'deaths': covid_deaths_rollup[todays_date]})
_clean_country_list(df_out)
_clean_country_list(covid_cases_rollup)
# Add observed death rate:
df_out['death_rate_observed'] = df_out.apply(
lambda row: row['deaths'] / float(row['cases']),
axis=1)
# Add covariate for days since first case
df_out['days_since_first_case'] = _compute_days_since_first_case(
covid_cases_rollup)
# Add CPI covariate:
_add_cpi_data(df_out)
# Add World Bank covariates:
_add_wb_data(df_out)
# Drop any country w/o covariate data:
num_null = df_out.isnull().sum(axis=1)
to_drop_idx = df_out.index[num_null > 1]
print('Dropping %i/%i countries due to lack of data' %
(len(to_drop_idx), len(df_out)))
df_out.drop(to_drop_idx, axis=0, inplace=True)
return df_out, todays_date
def _get_latest_covid_timeseries():
''' Pull latest time-series data from JHU CSSE database '''
repo = 'https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/'
data_path = 'csse_covid_19_data/csse_covid_19_time_series/'
all_data = {}
for status in ['Confirmed', 'Deaths', 'Recovered']:
file_name = 'time_series_covid19_%s_global' % status
all_data[status] = pd.read_csv(
'%s%s%s' % (repo, data_path, file_name))
return all_data
def _rollup_by_country(df):
'''
Roll up each raw time-series by country, adding up the cases
across the individual states/provinces within the country
:param df: Pandas DataFrame of raw data from CSSE
:return: DataFrame of country counts
'''
gb = df.groupby('Country/Region')
df_rollup = gb.sum()
df_rollup.drop(['Lat', 'Long'], axis=1, inplace=True, errors='ignore')
# Drop dates with all 0 count data
df_rollup.drop(df_rollup.columns[df_rollup.sum(axis=0) == 0],
axis=1,
inplace=True)
# Convert column strings to dates:
idx_as_dt = [datetime.strptime(x, '%m/%d/%y') for x in df_rollup.columns]
df_rollup.columns = idx_as_dt
return df_rollup
def _clean_country_list(df):
''' Clean up input country list in df '''
# handle recent changes in country names:
country_rename = {
'Hong Kong SAR': 'Hong Kong',
'Taiwan*': 'Taiwan',
'Czechia': 'Czech Republic',
'Brunei': 'Brunei Darussalam',
'Iran (Islamic Republic of)': 'Iran',
'Viet Nam': 'Vietnam',
'Russian Federation': 'Russia',
'Republic of Korea': 'South Korea',
'Republic of Moldova': 'Moldova',
'China': 'Mainland China'
}
df.rename(country_rename, axis=0, inplace=True)
df.drop(ignore_countries, axis=0, inplace=True, errors='ignore')
def _compute_days_since_first_case(df_cases):
''' Compute the country-wise days since first confirmed case
:param df_cases: country-wise time-series of confirmed case counts
:return: Series of country-wise days since first case
'''
date_first_case = df_cases[df_cases > 0].idxmin(axis=1)
days_since_first_case = date_first_case.apply(
lambda x: (df_cases.columns.max() - x).days)
# Add 1 month for China, since outbreak started late 2019:
days_since_first_case.loc['Mainland China'] += 30
return days_since_first_case
def _add_cpi_data(df_input):
'''
Add the Government transparency (CPI - corruption perceptions index)
data (by country) as a column in the COVID cases dataframe.
:param df_input: COVID-19 data rolled up country-wise
:return: None, add CPI data to df_input in place
'''
cpi_data = pd.read_excel(
'https://github.com/jwrichar/COVID19-mortality/blob/master/data/CPI2019.xlsx?raw=true',
skiprows=2)
cpi_data.set_index('Country', inplace=True, drop=True)
cpi_data.rename(cpi_country_mapping, axis=0, inplace=True)
# Add CPI score to input df:
df_input['cpi_score_2019'] = cpi_data['CPI score 2019']
def _add_wb_data(df_input):
'''
Add the World Bank data covariates as columns in the COVID cases dataframe.
:param df_input: COVID-19 data rolled up country-wise
:return: None, add World Bank data to df_input in place
'''
wb_data = pd.read_csv(
'https://raw.githubusercontent.com/jwrichar/COVID19-mortality/master/data/world_bank_data.csv',
na_values='..')
for (wb_name, var_name) in wb_covariates:
wb_series = wb_data.loc[wb_data['Series Code'] == wb_name]
wb_series.set_index('Country Name', inplace=True, drop=True)
wb_series.rename(wb_country_mapping, axis=0, inplace=True)
# Add WB data:
df_input[var_name] = _get_most_recent_value(wb_series)
def _get_most_recent_value(wb_series):
'''
Get most recent non-null value for each country in the World Bank
time-series data
'''
ts_data = wb_series[wb_series.columns[3::]]
def _helper(row):
row_nn = row[row.notnull()]
if len(row_nn):
return row_nn[-1]
else:
return np.nan
return ts_data.apply(_helper, axis=1)
#hide
# Load the data (see source/data.py):
df, todays_date = get_all_data()
# Impute NA's column-wise:
df = df.apply(lambda x: x.fillna(x.mean()),axis=0)
```
# Observed mortality rates
```
#collapse-hide
display(Markdown('Data as of %s' % todays_date))
reported_mortality_rate = df['deaths'].sum() / df['cases'].sum()
display(Markdown('Overall reported mortality rate: %.2f%%' % (100.0 * reported_mortality_rate)))
df_highest = df.sort_values('cases', ascending=False).head(15)
mortality_rate = pd.Series(
data=(df_highest['deaths']/df_highest['cases']).values,
index=map(lambda x: '%s (%i cases)' % (x, df_highest.loc[x]['cases']),
df_highest.index))
ax = mortality_rate.plot.bar(
figsize=(14,7), title='Reported Mortality Rate by Country (countries w/ highest case counts)')
ax.axhline(reported_mortality_rate, color='k', ls='--')
plt.show()
```
# Model
Estimate COVID-19 mortality rate, controling for country factors.
```
#hide
import numpy as np
import pymc3 as pm
def initialize_model(df):
# Normalize input covariates in a way that is sensible:
# (1) days since first case: upper
# mu_0 to reflect asymptotic mortality rate months after outbreak
_normalize_col(df, 'days_since_first_case', how='upper')
# (2) CPI score: upper
# mu_0 to reflect scenario in absence of corrupt govts
_normalize_col(df, 'cpi_score_2019', how='upper')
# (3) healthcare OOP spending: mean
# not sure which way this will go
_normalize_col(df, 'healthcare_oop_expenditure', how='mean')
# (4) hospital beds: upper
# more beds, more healthcare and tests
_normalize_col(df, 'hospital_beds', how='mean')
# (5) hci = human capital index: upper
# HCI measures education/health; mu_0 should reflect best scenario
_normalize_col(df, 'hci', how='mean')
# (6) % over 65: mean
# mu_0 to reflect average world demographic
_normalize_col(df, 'population_perc_over65', how='mean')
# (7) % rural: mean
# mu_0 to reflect average world demographic
_normalize_col(df, 'population_perc_rural', how='mean')
n = len(df)
covid_mortality_model = pm.Model()
with covid_mortality_model:
# Priors:
mu_0 = pm.Beta('mu_0', alpha=0.3, beta=10)
sig_0 = pm.Uniform('sig_0', lower=0.0, upper=mu_0 * (1 - mu_0))
beta = pm.Normal('beta', mu=0, sigma=5, shape=7)
sigma = pm.HalfNormal('sigma', sigma=5)
# Model mu from country-wise covariates:
# Apply logit transformation so logistic regression performed
mu_0_logit = np.log(mu_0 / (1 - mu_0))
mu_est = mu_0_logit + \
beta[0] * df['days_since_first_case_normalized'].values + \
beta[1] * df['cpi_score_2019_normalized'].values + \
beta[2] * df['healthcare_oop_expenditure_normalized'].values + \
beta[3] * df['hospital_beds_normalized'].values + \
beta[4] * df['hci_normalized'].values + \
beta[5] * df['population_perc_over65_normalized'].values + \
beta[6] * df['population_perc_rural_normalized'].values
mu_model_logit = pm.Normal('mu_model_logit',
mu=mu_est,
sigma=sigma,
shape=n)
# Transform back to probability space:
mu_model = np.exp(mu_model_logit) / (np.exp(mu_model_logit) + 1)
# tau_i, mortality rate for each country
# Parametrize with (mu, sigma)
# instead of (alpha, beta) to ease interpretability.
tau = pm.Beta('tau', mu=mu_model, sigma=sig_0, shape=n)
# tau = pm.Beta('tau', mu=mu_0, sigma=sig_0, shape=n)
# Binomial likelihood:
d_obs = pm.Binomial('d_obs',
n=df['cases'].values,
p=tau,
observed=df['deaths'].values)
return covid_mortality_model
def _normalize_col(df, colname, how='mean'):
'''
Normalize an input column in one of 3 ways:
* how=mean: unit normal N(0,1)
* how=upper: normalize to [-1, 0] with highest value set to 0
* how=lower: normalize to [0, 1] with lowest value set to 0
Returns df modified in place with extra column added.
'''
colname_new = '%s_normalized' % colname
if how == 'mean':
mu = df[colname].mean()
sig = df[colname].std()
df[colname_new] = (df[colname] - mu) / sig
elif how == 'upper':
maxval = df[colname].max()
minval = df[colname].min()
df[colname_new] = (df[colname] - maxval) / (maxval - minval)
elif how == 'lower':
maxval = df[colname].max()
minval = df[colname].min()
df[colname_new] = (df[colname] - minval) / (maxval - minval)
#hide
# Initialize the model:
mod = initialize_model(df)
# Run MCMC sampler1
with mod:
trace = pm.sample(300, tune=100,
chains=3, cores=2)
#collapse-hide
n_samp = len(trace['mu_0'])
mu0_summary = pm.summary(trace).loc['mu_0']
print("COVID-19 Global Mortality Rate Estimation:")
print("Posterior mean: %0.2f%%" % (100*trace['mu_0'].mean()))
print("Posterior median: %0.2f%%" % (100*np.median(trace['mu_0'])))
lower = np.sort(trace['mu_0'])[int(n_samp*0.025)]
upper = np.sort(trace['mu_0'])[int(n_samp*0.975)]
print("95%% posterior interval: (%0.2f%%, %0.2f%%)" % (100*lower, 100*upper))
prob_lt_reported = sum(trace['mu_0'] < reported_mortality_rate) / len(trace['mu_0'])
print("Probability true rate less than reported rate (%.2f%%) = %.2f%%" %
(100*reported_mortality_rate, 100*prob_lt_reported))
print("")
# Posterior plot for mu0
print('Posterior probability density for COVID-19 mortality rate, controlling for country factors:')
ax = pm.plot_posterior(trace, var_names=['mu_0'], figsize=(18, 8), textsize=18,
credible_interval=0.95, bw=3.0, lw=3, kind='kde',
ref_val=round(reported_mortality_rate, 3))
```
## Magnitude and Significance of Factors
For bias in reported COVID-19 mortality rate
```
#collapse-hide
# Posterior summary for the beta parameters:
beta_summary = pm.summary(trace).head(7)
beta_summary.index = ['days_since_first_case', 'cpi', 'healthcare_oop', 'hospital_beds', 'hci', 'percent_over65', 'percent_rural']
beta_summary.reset_index(drop=False, inplace=True)
err_vals = ((beta_summary['hpd_3%'] - beta_summary['mean']).values,
(beta_summary['hpd_97%'] - beta_summary['mean']).values)
ax = beta_summary.plot(x='index', y='mean', kind='bar', figsize=(14, 7),
title='Posterior Distribution of Beta Parameters',
yerr=err_vals, color='lightgrey',
legend=False, grid=True,
capsize=5)
beta_summary.plot(x='index', y='mean', color='k', marker='o', linestyle='None',
ax=ax, grid=True, legend=False, xlim=plt.gca().get_xlim())
plt.savefig('../images/corvid-mortality.png')
```
# About This Analysis
This analysis was done by [Joseph Richards](https://twitter.com/joeyrichar)
In this project[^3], we attempt to estimate the true mortality rate[^1] for COVID-19 while controlling for country-level covariates[^2][^4] such as:
* age of outbreak in the country
* transparency of the country's government
* access to healthcare
* demographics such as age of population and rural vs. urban
Estimating a mortality rate lower than the overall reported rate likely implies that there has been **significant under-testing and under-reporting of cases globally**.
## Interpretation of Country-Level Parameters
1. days_since_first_case - positive (very statistically significant). As time since outbreak increases, expected mortality rate **increases**, as expected.
2. cpi - negative (statistically significant). As government transparency increases, expected mortality rate **decreases**. This may mean that less transparent governments under-report cases, hence inflating the mortality rate.
3. healthcare avg. out-of-pocket spending - no significant trend.
4. hospital beds per capita - no significant trend.
5. Human Capital Index - no significant trend (slightly negative = mortality rates decrease with increased mobilization of the country)
6. percent over 65 - positive (statistically significant). As population age increases, the mortality rate also **increases**, as expected.
7. percent rural - no significant trend.
[^1]: As of March 10, the **overall reported mortality rate is 3.5%**. However, this figure does not account for **systematic biases in case reporting and testing**. The observed mortality of COVID-19 has varied widely from country to country (as of early March 2020). For instance, as of March 10, mortality rates have ranged from < 0.1% in places like Germany (1100+ cases) to upwards of 5% in Italy (9000+ cases) and 3.9% in China (80k+ cases).
[^2]: The point of our modelling work here is to **try to understand and correct for the country-to-country differences that may cause the observed discrepancies in COVID-19 country-wide mortality rates**. That way we can "undo" those biases and try to **pin down an overall *real* mortality rate**.
[^3]: Full details about the model are available at: https://github.com/jwrichar/COVID19-mortality
[^4]: The affects of these parameters are subject to change as more data are collected.
# Appendix: Model Diagnostics
The following trace plots help to assess the convergence of the MCMC sampler.
```
#hide_input
import arviz as az
az.plot_trace(trace, compact=True);
```
| github_jupyter |
```
videos = """
https://www.youtube.com/watch?v=fAqs-YC8irc
https://www.youtube.com/watch?v=G2oPClnoJpg
https://www.youtube.com/watch?v=unmCwK_9El8
https://www.youtube.com/watch?v=JOL8sFYlBU4
https://www.youtube.com/watch?v=T907QXqWeF0
https://www.youtube.com/watch?v=VjzeR0xgQ8I
https://www.youtube.com/watch?v=BQpcj10O8nA
https://www.youtube.com/watch?v=I_E38TJpU6Q
https://www.youtube.com/watch?v=VH7aC2lbg1c
https://www.youtube.com/watch?v=ZoYg34o8rqI
https://www.youtube.com/watch?v=b_d-Yf-Gzyw
https://www.youtube.com/watch?v=IDg0E93eM2U
https://www.youtube.com/watch?v=N-YoIbr6-1Q
https://www.youtube.com/watch?v=WNIEhpPsfDk
https://www.youtube.com/watch?v=ERPf3TbiK0E
https://www.youtube.com/watch?v=qaGfP5sk7z8
https://www.youtube.com/watch?v=obkefyOHKr4
https://www.youtube.com/watch?v=NtnT4MmSFJw
https://www.youtube.com/watch?v=IxR5I3HS2T8
https://www.youtube.com/watch?v=YSwgyuUO2cU
https://www.youtube.com/watch?v=l6kBSvtuc_8
https://www.youtube.com/watch?v=0xSswYU5ZFE
https://www.youtube.com/watch?v=8IoPsPVxxgs
https://www.youtube.com/watch?v=rx9EW69Qtn8
https://www.youtube.com/watch?v=jZ3_M9TkCX4
https://www.youtube.com/watch?v=ylH-VEO5IbQ
https://www.youtube.com/watch?v=6ZWiTtjSvj8
https://www.youtube.com/watch?v=yXIDru3zM2w
https://www.youtube.com/watch?v=IzL2lwkCDyc
https://www.youtube.com/watch?v=mRCrxLUcVPk
https://www.youtube.com/watch?v=bYjjQpP8FaM
https://www.youtube.com/watch?v=H6W2uNuIm9o
https://www.youtube.com/watch?v=JOI6naLEpMI
https://www.youtube.com/watch?v=oqJny9Qze3k
https://www.youtube.com/watch?v=V_XteZbz5pg
https://www.youtube.com/watch?v=ttmqMaoMdVw&ab_channel=EverydayChinese
https://www.youtube.com/watch?v=s8wA65b8hBs&ab_channel=EazyMandarin
https://www.youtube.com/watch?v=gFE8L6aq1Gs&ab_channel=EazyMandarin
https://www.youtube.com/watch?v=Wt5tmL9yex4&ab_channel=EazyMandarin
https://www.youtube.com/watch?v=jWKfxkF0GVg&ab_channel=EazyMandarin
https://www.youtube.com/watch?v=Wg56kS9gA50&ab_channel=GreatestAudioBooks
https://www.youtube.com/watch?v=cUAlLdwwdYQ&ab_channel=GreatAudioBooksInPublicDomain
https://www.youtube.com/watch?v=U1ywZGocjro&ab_channel=GreatAudioBooksInPublicDomain
https://www.youtube.com/watch?v=DIP-Fgq27f4&ab_channel=GreatAudioBooksInPublicDomain
https://www.youtube.com/watch?v=18zdf5uciVs&ab_channel=%E9%9B%AA%E8%9D%B6
https://www.youtube.com/watch?v=Tmg6oERWyH4
https://www.youtube.com/watch?v=5tH0odCiayo
https://www.youtube.com/watch?v=j18SHgT4n3o
https://www.youtube.com/watch?v=1uZs-xVpfHg
https://www.youtube.com/watch?v=7or77vFZSa4&ab_channel=GreatAudioBooksInPublicDomain
https://www.youtube.com/watch?v=DcPviW7husQ&ab_channel=LowmanWhitfield
https://www.youtube.com/watch?v=NLu0JW6VtGM&ab_channel=EazyMandarin
https://www.youtube.com/watch?v=BieUQR4skTk
https://www.youtube.com/watch?v=Hvh21Ks2HEg
https://www.youtube.com/watch?v=UKuT2XuZ7fM
"""
videos = list(set(filter(None, videos.split('\n'))))
len(videos)
import youtube_dl
import mp
from tqdm import tqdm
def loop(urls):
urls = urls[0]
ydl_opts = {
'format': 'bestaudio/best',
'postprocessors': [{
'key': 'FFmpegExtractAudio',
'preferredcodec': 'mp3',
'preferredquality': '192',
}],
'no-check-certificate': True
}
for i in tqdm(range(len(urls))):
try:
with youtube_dl.YoutubeDL(ydl_opts) as ydl:
ydl.download([urls[i]])
except:
pass
import mp
mp.multiprocessing(videos, loop, cores = 12, returned = False)
!mkdir mandarin
!mv *.mp3 mandarin
```
| github_jupyter |
```
# KNN algorithm (K-Nearest Neighbor) simplified. It can be used for both classification and
# regression predictive problems. For simplicity, IRIS dataset is used to test the code.
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
from sklearn import datasets
from sklearn.metrics import mean_absolute_error as MEA # I imported this but will try to not to use it.
Iris = datasets.load_iris()
print(type(Iris.data))
print(Iris.data.shape)
# check the length of the data, shuffle the data, prepare the test and train sets
# it is normally good to split the data to 70-30 but our KNN does close to 100 percent accuracy in that case
# the data is intentionally split 66-33 to show some errors for overfitting and underfitting
# X is used for the data and y is used for labels
length_of_data = len(Iris.data)
print(length_of_data)
indices = np.random.permutation(len(Iris.data))
X_train = Iris.data[indices[:100]]
y_train = Iris.target[indices[:100]]
X_test = Iris.data[indices[100:]]
y_test = Iris.target[indices[100:]]
# view the data
print(X_train[:5])
print(y_train[:5])
# test the normal distance function to get a scalar distance value in between instances
print(np.linalg.norm(X_train[3] - X_test[1]))
def K_Nearest_Neighbor(test_set, test_labels, training_set, training_labels, n):
results = []
for i in range(len(test_set)):
indices_visited = [] # make a list of indices visited
votes = []
for j in range(n):
min = np.inf
for k in range(len(training_set)):
difference = np.linalg.norm(training_set[k] - test_set[i])
if difference < min and k not in indices_visited:
min = difference
index_at_min_distance = k
the_label_found_at_min_dist = training_labels[k]
indices_visited.append(index_at_min_distance)
votes.append(the_label_found_at_min_dist)
# count the votes
a_dict = {}
for i in votes:
if i in a_dict:
a_dict[i] += 1
else:
a_dict[i] = 1
#print(a_dict)
# find the maximum seen vote
max_v = -np.inf
for k,v in a_dict.items():
if v > max_v:
max_v = v
set_k = int(k)
# the most common vote is the label of the prediction
results.append(set_k)
#print(results, test_labels)
# check the accuracy by comparing our target and actual test labels
accuracy = np.sum(results == test_labels) / len(test_labels)
# print('the actual results are: {}'.format(test_labels))
# print('our labels show: {}'.format(results))
# print('the accuracy is: {}'.format(accuracy*100))
return np.around(100*accuracy, decimals = 2)
# print(K_Nearest_Neighbor(X_test, y_test, X_train, y_train, 1))
all_results = []
for i in range(1,30):
each_result = K_Nearest_Neighbor(X_test, y_test, X_train, y_train, i)
all_results.append(each_result)
print(all_results, len(all_results))
plt.plot(np.arange(len(all_results)), np.array(all_results))
# Conclusion: KNN 8-10 gives the best accuracy for this training and test sets.
# It is slightly hard to see the overfitting
```
| github_jupyter |
```
pp
import sklearn as sns
sns.__version__
```
___
<a href='https://mp.weixin.qq.com/mp/appmsgalbum?__biz=Mzg2OTU4NzI3NQ==&action=getalbum&album_id=1764511202329624577&scene=126#wechat_redirect'> <img src=../../../pic/project_logo.jpg></a>
___
# Matplotlib Overview Lecture
## Introduction
Import the `matplotlib.pyplot` module
```
import matplotlib.pyplot as plt
```
输入下方的代码在notebook中显示图
```
%matplotlib inline
```
That line is only for jupyter notebooks, if you are using another editor, you'll use: **plt.show()** at the end of all your plotting commands to have the figure pop up in another window.
# Basic Example
### Example
x = 0 到 10 的整数
y = x 的平方
```
import numpy as np
x = np.linspace(0, 10, 11)
y = x ** 2
x
y
```
## Basic Matplotlib Commands
创建第一个图(用 Shift+Tab 来检查函数帮助文档!)。
```
plt.plot(x, y, 'r--') # 'r' 红色 -- 代表虚线
plt.xlabel('X 轴')
plt.ylabel('Y 轴')
plt.title('标题')
plt.show()
```
发现好像不能打中文啊,别着急,加上下面的代码就可以了!
```
plt.rcParams['font.sans-serif']=['SimHei'] #用来正常显示中文标签
plt.rcParams['axes.unicode_minus'] = False #用来正常显示负号
plt.plot(x, y, 'r--') # 'r' 红色 -- 代表虚线
plt.xlabel('X 轴')
plt.ylabel('Y 轴')
plt.title('标题')
plt.show()
```
## Creating Multiplots on Same Canvas
如果想在一个画布上画子图咋办,我们用 subplot!
```
# plt.subplot(nrows, ncols, plot_number)
plt.subplot(1,2,1) # 一行两列的第一个子图
plt.plot(x, y, 'r--')
plt.subplot(1,2,2) # 一行两列的第一个子图
plt.plot(y, x, 'g*-');
```
如何更改画布大小?
```
plt.figure(figsize = [12,3]) # 更改画布大小
plt.subplot(1,2,1) # 一行两列的第一个子图
plt.plot(x, y, 'r--')
plt.subplot(1,2,2) # 一行两列的第一个子图
plt.plot(y, x, 'g*-');
```
___
# Matplotlib Object Oriented Method
现在我们已经了解了基础知识,让我们通过更正式的 Matplotlib 面向对象 API 的介绍来分解它。这意味着我们将实例化图形对象,然后从该对象调用方法或属性。
## Introduction to the Object Oriented Method
使用更正式的面向对象方法的主要思想是创建图形对象,然后调用该对象的方法或属性。在处理上面有多个绘图的画布时,这种方法更好。
首先,我们创建一个图形实例。然后我们可以在该图中添加轴:
```
# Create Figure (empty canvas)
fig = plt.figure()
# Add set of axes to figure
axes = fig.add_axes([0.1, 0.1, 0.8, 0.8]) # left, bottom, width, height (range 0 to 1)
# Plot on that set of axes
axes.plot(x, y, 'b')
axes.set_xlabel('Set X Label') # Notice the use of set_ to begin methods
axes.set_ylabel('Set y Label')
axes.set_title('Set Title')
```
代码稍微复杂一点,但优点是我们现在可以完全控制绘图轴的放置位置,并且我们可以轻松地在图中添加多个轴:
```
# Creates blank canvas
fig = plt.figure()
axes1 = fig.add_axes([0.1, 0.1, 0.8, 0.8]) # main axes
axes2 = fig.add_axes([0.2, 0.5, 0.4, 0.3]) # inset axes
# Larger Figure Axes 1
axes1.plot(x, y, 'b')
axes1.set_xlabel('X_label_axes2')
axes1.set_ylabel('Y_label_axes2')
axes1.set_title('Axes 2 Title')
# Insert Figure Axes 2
axes2.plot(y, x, 'r')
axes2.set_xlabel('X_label_axes2')
axes2.set_ylabel('Y_label_axes2')
axes2.set_title('Axes 2 Title');
```
## subplots()
plt.subplots() 对象将充当更自动的【轴】管理器。
基本用例:
```
# Use similar to plt.figure() except use tuple unpacking to grab fig and axes
fig, axes = plt.subplots()
# Now use the axes object to add stuff to plot
axes.plot(x, y, 'r')
axes.set_xlabel('x')
axes.set_ylabel('y')
axes.set_title('title');
```
Then you can specify the number of rows and columns when creating the subplots() object:
```
# Empty canvas of 1 by 2 subplots
fig, axes = plt.subplots(nrows=1, ncols=2)
# Axes is an array of axes to plot on
axes
```
We can iterate through this array:
```
for ax in axes:
ax.plot(x, y, 'b')
ax.set_xlabel('x')
ax.set_ylabel('y')
ax.set_title('title')
# Display the figure object
fig
```
matplolib 的一个常见问题是子图或图形会重叠。
我们可以使用 **fig.tight_layout()** 或 **plt.tight_layout()** 方法,它会自动调整轴在图形画布上的位置,以免内容重叠:
```
fig, axes = plt.subplots(nrows=1, ncols=2)
for ax in axes:
ax.plot(x, y, 'g')
ax.set_xlabel('x')
ax.set_ylabel('y')
ax.set_title('title')
fig
plt.tight_layout()
```
### Figure size, aspect ratio and DPI
Matplotlib 允许在创建 Figure 对象时指定纵横比、DPI 和图形大小。您可以使用 `figsize` 和 `dpi` 关键字参数。
* `figsize` 是以英寸为单位的图形宽度和高度的元组
* `dpi` 是每英寸点数(每英寸像素)。
例如:
```
fig = plt.figure(figsize=(8,4), dpi=100)
```
相同的参数也可以传递给布局管理器,例如 `subplots` 函数:
```
fig, axes = plt.subplots(figsize=(12,3))
axes.plot(x, y, 'r')
axes.set_xlabel('x')
axes.set_ylabel('y')
axes.set_title('title');
```
## Saving figures
Matplotlib can generate high-quality output in a number formats, including PNG, JPG, EPS, SVG, PGF and PDF.
要将图形保存到文件中,我们可以使用 `Figure` 类中的 `savefig` 方法:|
```
fig.savefig("filename.png")
```
Here we can also optionally specify the DPI and choose between different output formats:
```
fig.savefig("filename.png", dpi=200)
```
____
## Legends, labels and titles
现在我们已经介绍了如何创建图形画布和向画布添加轴实例的基础知识,让我们看看如何用标题、轴标签和图例装饰图形。
**Figure titles**
可以为图中的每个轴实例添加标题。要设置标题,请在坐标区实例中使用 `set_title` 方法:
```
ax.set_title("title");
```
**Axis labels**
同样,使用 `set_xlabel` 和 `set_ylabel` 方法,我们可以设置 X 和 Y 轴的标签:
```
ax.set_xlabel("x")
ax.set_ylabel("y");
```
### Legends
在将绘图或其他对象添加到图形时使用 **label="label text"** 关键字参数,然后使用不带参数的 **legend** 方法将图例添加到图形:
```
fig = plt.figure()
ax = fig.add_axes([0,0,1,1])
ax.plot(x, x**2, label="x**2")
ax.plot(x, x**3, label="x**3")
ax.legend()
```
可能图例会和图形重叠!
**legend** 函数采用可选的关键字参数 **loc**,可用于指定要在图中的何处绘制图例。 **loc** 的允许值是可以绘制图例的各个位置的数字代码。有关详细信息,请参阅 [文档页面](http://matplotlib.org/users/legend_guide.html#legend-location)。一些最常见的 **loc** 值是:
```
# Lots of options....
ax.legend(loc=1) # upper right corner
ax.legend(loc=2) # upper left corner
ax.legend(loc=3) # lower left corner
ax.legend(loc=4) # lower right corner
# .. many more options are available
# Most common to choose
ax.legend(loc=1) # let matplotlib decide the optimal location
fig
```
## Setting colors, linewidths, linetypes
Matplotlib 为您提供了*很多*用于自定义颜色、线宽和线型的选项。
### Colors with MatLab like syntax
使用 matplotlib,我们可以通过多种方式定义线条和其他图形元素的颜色。首先,我们可以使用类似 MATLAB 的语法,其中“b”表示蓝色,“g”表示绿色等。还支持用于选择线型的 MATLAB API:例如,“b”。 -' 表示带点的蓝线:
```
# MATLAB style line color and style
fig, ax = plt.subplots()
ax.plot(x, x**2, 'b.-') # blue line with dots
ax.plot(x, x**3, 'g--') # green dashed line
```
### Colors with the color= parameter
我们还可以通过名称或 RGB 十六进制代码定义颜色,并可选择使用 `color` 和 `alpha` 关键字参数提供 alpha 值。 Alpha 表示不透明度。
```
fig, ax = plt.subplots()
ax.plot(x, x+1, color="blue", alpha=0.1) # half-transparant
ax.plot(x, x+2, color="#8B008B") # RGB hex code
ax.plot(x, x+3, color="#FF8C00") # RGB hex code
```
### Line and marker styles
要更改线宽,我们可以使用 `linewidth` 或 `lw` 关键字参数。可以使用 `linestyle` 或 `ls` 关键字参数选择线条样式:
```
fig, ax = plt.subplots(figsize=(12,6))
ax.plot(x, x+1, color="red", linewidth=0.25)
ax.plot(x, x+2, color="red", linewidth=0.50)
ax.plot(x, x+3, color="red", linewidth=1.00)
ax.plot(x, x+4, color="red", linewidth=2.00)
# possible linestype options ‘-‘, ‘–’, ‘-.’, ‘:’, ‘steps’
ax.plot(x, x+5, color="green", lw=3, linestyle='-')
ax.plot(x, x+6, color="green", lw=3, ls='-.')
ax.plot(x, x+7, color="green", lw=3, ls=':')
# custom dash
line, = ax.plot(x, x+8, color="black", lw=1.50)
line.set_dashes([5, 10, 15, 10]) # format: line length, space length, ...
# possible marker symbols: marker = '+', 'o', '*', 's', ',', '.', '1', '2', '3', '4', ...
ax.plot(x, x+ 9, color="blue", lw=3, ls='-', marker='+')
ax.plot(x, x+10, color="blue", lw=3, ls='--', marker='o')
ax.plot(x, x+11, color="blue", lw=3, ls='-', marker='s')
ax.plot(x, x+12, color="blue", lw=3, ls='--', marker='1')
# marker size and color
ax.plot(x, x+13, color="purple", lw=1, ls='-', marker='o', markersize=2)
ax.plot(x, x+14, color="purple", lw=1, ls='-', marker='o', markersize=4)
ax.plot(x, x+15, color="purple", lw=1, ls='-', marker='o', markersize=8, markerfacecolor="red")
ax.plot(x, x+16, color="purple", lw=1, ls='-', marker='s', markersize=8,
markerfacecolor="yellow", markeredgewidth=3, markeredgecolor="green");
```
### Control over axis appearance
在本节中,我们将研究在 matplotlib 图中控制轴大小的属性。
## Plot range
我们可以使用轴对象中的 `set_ylim` 和 `set_xlim` 方法来配置轴的范围,或者使用 `axis('tight')` 来自动获取“紧密拟合”的轴范围:
```
fig, axes = plt.subplots(1, 3, figsize=(12, 4))
axes[0].plot(x, x**2, x, x**3)
axes[0].set_title("default axes ranges")
axes[1].plot(x, x**2, x, x**3)
axes[1].axis('tight')
axes[1].set_title("tight axes")
axes[2].plot(x, x**2, x, x**3)
axes[2].set_ylim([0, 60])
axes[2].set_xlim([2, 5])
axes[2].set_title("custom axes range");
```
# Special Plot Types
我们可以创建许多专门的图,例如条形图、直方图、散点图等等。我们将使用 seaborn(Python 的统计绘图库)实际创建大多数此类绘图。但这里有一些此类图的示例:
```
plt.scatter(x,y)
from random import sample
data = sample(range(1, 1000), 100)
plt.hist(data,bins = 20)
data = [np.random.normal(0, std, 100) for std in range(1, 4)]
# rectangular box plot
plt.boxplot(data,vert=True,patch_artist=True);
```
## Further reading
* http://www.matplotlib.org - The project web page for matplotlib.
* https://github.com/matplotlib/matplotlib - The source code for matplotlib.
* http://matplotlib.org/gallery.html - A large gallery showcaseing various types of plots matplotlib can create. Highly recommended!
* http://www.loria.fr/~rougier/teaching/matplotlib - A good matplotlib tutorial.
* http://scipy-lectures.github.io/matplotlib/matplotlib.html - Another good matplotlib reference.
| github_jupyter |
Now that we have tried some basic **classification** and **regression** algorithms, more complicated algorithms will follow a familiar pattern: given a set of inputs and outputs, can I develop a model that relates the inputs to the outputs, and can I use that model to predict new outputs from a set of new inputs?
In this sense, you could quite easily treat these new regression and classification techniques as black boxes, feed data in and get data out. But don't do that. For a number of reasons. First, that's boring, right? And second, if you don't understand the inner workings of each classifier and regressor, you won't know how to properly tune them (a part of model selection). Here's one example from `scikit-learn` comparing a wide variety of classification techniques:
<img alt="sklearn_classification.png" src="https://github.com/UWDIRECT/UWDIRECT.github.io/blob/master/Wi19_content/DSMCER/images/sklearn_classification.png?raw=true">
OK, let's get some data. For the purposes of this notebook, we're going to create a linearly separable dataset really quick using some `scikit-learn magic`:
```
import numpy as np
import matplotlib.pyplot as plt
from sklearn.datasets import make_classification
from sklearn.model_selection import train_test_split
from matplotlib.colors import ListedColormap
X, y = make_classification(n_features=2, n_redundant=0, n_informative=2,
random_state=1, n_clusters_per_class=1, class_sep=0.7)
#rng = np.random.RandomState(2)
#X += 2 * rng.uniform(size=X.shape)
data = (X, y)
```
Let's do a training-test split:
```
X_train, X_test, y_train, y_test = \
train_test_split(X, y, test_size=.4, random_state=42)
```
Let's plot the data to see what we have.
```
figure, ax = plt.subplots(figsize=(4,4))
cm = plt.cm.RdBu
cm_bright = ListedColormap(['#FF0000', '#0000FF'])
# Plot the training points
ax.scatter(X_train[:, 0], X_train[:, 1], c=y_train, cmap=cm_bright,
edgecolors='k', label='Training')
# Plot the testing points
ax.scatter(X_test[:, 0], X_test[:, 1], c=y_test, cmap=cm_bright, alpha=0.6,
edgecolors='k', label='Test')
#ax.set_xlim([-3,4])
#ax.set_ylim([0,4])
ax.legend()
```
Remember how our KNN classifier worked? It was able to predict the class based on the surrounding K nearest neighbors. An SVM works on a different paradigm. In this case, we're going to try to draw a line that splits the two groups. Let's try applying a simple SVM algorithm to do this for us:
```
from sklearn.svm import SVC
clf = SVC(kernel='linear', C=1)
h=0.02
x_min, x_max = X[:, 0].min() - .5, X[:, 0].max() + .5
y_min, y_max = X[:, 1].min() - .5, X[:, 1].max() + .5
xx, yy = np.meshgrid(np.arange(x_min, x_max, h),
np.arange(y_min, y_max, h))
clf.fit(X_train, y_train)
score = clf.score(X_test, y_test)
figure, ax = plt.subplots(figsize=(4,4))
# Plot the decision boundary. For that, we will assign a color to each
# point in the mesh [x_min, x_max]x[y_min, y_max].
if hasattr(clf, "decision_function"):
Z = clf.decision_function(np.c_[xx.ravel(), yy.ravel()])
else:
Z = clf.predict_proba(np.c_[xx.ravel(), yy.ravel()])[:, 1]
# Put the result into a color plot
Z = Z.reshape(xx.shape)
Zbi = Z > np.median(Z)
ax.contourf(xx, yy, Zbi, cmap=cm, alpha=.1)
# Plot the training points
ax.scatter(X_train[:, 0], X_train[:, 1], c=y_train, cmap=cm_bright,
edgecolors='k', label='Training')
# Plot the testing points
ax.scatter(X_test[:, 0], X_test[:, 1], c=y_test, cmap=cm_bright,
edgecolors='k', alpha=0.6, label='Test')
ax.legend()
```
Luckily, this dataset has two distinct groups, so separating the two with a line is pretty simple. However, that's not always the case. This method would fall apart if there was any overlap. This simple classifier is called the **maximal margin classifier**.
What happens when there is no easy boundary?
```
X, y = make_classification(n_features=2, n_redundant=0, n_informative=2,
random_state=1, n_clusters_per_class=1, class_sep=0.8)
rng = np.random.RandomState(2)
X += 2 * rng.uniform(size=X.shape)
data = (X, y)
X_train, X_test, y_train, y_test = \
train_test_split(X, y, test_size=.4, random_state=42)
figure, ax = plt.subplots(figsize=(4,4))
cm = plt.cm.RdBu
cm_bright = ListedColormap(['#FF0000', '#0000FF'])
# Plot the training points
ax.scatter(X_train[:, 0], X_train[:, 1], c=y_train, cmap=cm_bright,
edgecolors='k', label='Training')
# Plot the testing points
ax.scatter(X_test[:, 0], X_test[:, 1], c=y_test, cmap=cm_bright, alpha=0.6,
edgecolors='k', label='Test')
#ax.set_xlim([-3,4])
#ax.set_ylim([0,4])
ax.legend()
```
The **maximal margin classifier** can't solve this problem anymore. We're now going to introduce a tuning parameter, **C**. Think of C as the "budget" for the number of violations we will allow to the decision boundary. When C is zero, you won't tolerate any violations: all blues must be on one side of the line, and all reds must be on the other.
In this sense, C, is our **bias/variance tradeoff parameter**. Cool, huh?! Similar to K in K-nearest neighbors. Let's try seeing what happens when we adjust C:
```
C_values = [0.00000001, 0.001, 0.1, 1, 10, 100, 1000, 10000]
figure, axes = plt.subplots(nrows=2, ncols=4, figsize=(12, 9))
axes = [item for sublist in axes for item in sublist]
h=0.02
x_min, x_max = X[:, 0].min() - .5, X[:, 0].max() + .5
y_min, y_max = X[:, 1].min() - .5, X[:, 1].max() + .5
xx, yy = np.meshgrid(np.arange(x_min, x_max, h),
np.arange(y_min, y_max, h))
for C, ax in zip(C_values, axes):
clf = SVC(kernel='linear', C=C)
clf.fit(X_train, y_train)
score = clf.score(X_test, y_test)
# Plot the decision boundary. For that, we will assign a color to each
# point in the mesh [x_min, x_max]x[y_min, y_max].
if hasattr(clf, "decision_function"):
Z = clf.decision_function(np.c_[xx.ravel(), yy.ravel()])
else:
Z = clf.predict_proba(np.c_[xx.ravel(), yy.ravel()])[:, 1]
# Put the result into a color plot
Z = Z.reshape(xx.shape)
Zbi = Z > np.median(Z)
ax.contourf(xx, yy, Zbi, cmap=cm, alpha=.1)
# Plot the training points
ax.scatter(X_train[:, 0], X_train[:, 1], c=y_train, cmap=cm_bright,
edgecolors='k', label='Training')
# Plot the testing points
# ax.scatter(X_test[:, 0], X_test[:, 1], c=y_test, cmap=cm_bright,
# edgecolors='k', alpha=0.6, label='Test')
ax.legend()
ax.set_title('C Value: {}'.format(str(C)))
```
But, you might be complaining, this isn't even that exciting! Sure, our predictor works OK-ish. But this is literally just drawing a line. Our KNN predictor did better than this! Is this even a useful method?
**SVMs** really get their power when you take advantage of something called a **kernel**. A kernal takes your original dataset, transoforms it into higher dimensional space, and then draws a hyperplane in that new hyperdimensional space. Such hyperplane in the original predictor space becomes nonlinear. It's a lot of math that we won't go into here. But let's see what happens when we try to classify some data with a radial kernel.
```
from sklearn.datasets import make_circles
data = make_circles(noise=0.2, factor=0.5, random_state=1)
X, y = data
X_train, X_test, y_train, y_test = \
train_test_split(X, y, test_size=.4, random_state=42)
clf = SVC(kernel='linear', C=1)
h=0.02
x_min, x_max = X[:, 0].min() - .5, X[:, 0].max() + .5
y_min, y_max = X[:, 1].min() - .5, X[:, 1].max() + .5
xx, yy = np.meshgrid(np.arange(x_min, x_max, h),
np.arange(y_min, y_max, h))
clf.fit(X_train, y_train)
score = clf.score(X_test, y_test)
figure, ax = plt.subplots(figsize=(4,4))
# Plot the decision boundary. For that, we will assign a color to each
# point in the mesh [x_min, x_max]x[y_min, y_max].
if hasattr(clf, "decision_function"):
Z = clf.decision_function(np.c_[xx.ravel(), yy.ravel()])
else:
Z = clf.predict_proba(np.c_[xx.ravel(), yy.ravel()])[:, 1]
# Put the result into a color plot
Z = Z.reshape(xx.shape)
Zbi = Z > np.median(Z)
ax.contourf(xx, yy, Zbi, cmap=cm, alpha=.1)
# Plot the training points
ax.scatter(X_train[:, 0], X_train[:, 1], c=y_train, cmap=cm_bright,
edgecolors='k', label='Training')
# Plot the testing points
ax.scatter(X_test[:, 0], X_test[:, 1], c=y_test, cmap=cm_bright,
edgecolors='k', alpha=0.6, label='Test')
ax.legend()
```
Yeah, a linear separator isn't going to work here. Let's try a **radial kernel**!
```
clf = SVC(gamma=4, C=0.1)
h=0.02
x_min, x_max = X[:, 0].min() - .5, X[:, 0].max() + .5
y_min, y_max = X[:, 1].min() - .5, X[:, 1].max() + .5
xx, yy = np.meshgrid(np.arange(x_min, x_max, h),
np.arange(y_min, y_max, h))
clf.fit(X_train, y_train)
score = clf.score(X_test, y_test)
figure, ax = plt.subplots(figsize=(4,4))
# Plot the decision boundary. For that, we will assign a color to each
# point in the mesh [x_min, x_max]x[y_min, y_max].
if hasattr(clf, "decision_function"):
Z = clf.decision_function(np.c_[xx.ravel(), yy.ravel()])
else:
Z = clf.predict_proba(np.c_[xx.ravel(), yy.ravel()])[:, 1]
# Put the result into a color plot
Z = Z.reshape(xx.shape)
Zbi = Z > np.median(Z)
ax.contourf(xx, yy, Zbi, cmap=cm, alpha=.1)
# Plot the training points
ax.scatter(X_train[:, 0], X_train[:, 1], c=y_train, cmap=cm_bright,
edgecolors='k', label='Training')
# Plot the testing points
ax.scatter(X_test[:, 0], X_test[:, 1], c=y_test, cmap=cm_bright,
edgecolors='k', alpha=0.6, label='Test')
ax.legend()
```
There are additional kernels that you can try. If you want to build your SVM skills, try some of these additional exercises:
* Calculate the MSE on the above training and test datasets and compare each of the models.
* Look up the types of kernels available, and try using some alternate kernels.
* Compare the KNN classifier to the linear and radial SVMs
## Now we apply these knowledge to the classification problem of atoms ##
Hint: borrow what you learned from the KNN jupyter notebook
```
import pandas as pd
%matplotlib inline
# Add a line to import the SVC/SVM pieces
```
#### Read in the data
* elemental data: [https://raw.githubusercontent.com/UWDIRECT/UWDIRECT.github.io/master/Wi18_content/DSMCER/atomsradii.csv](https://raw.githubusercontent.com/UWDIRECT/UWDIRECT.github.io/master/Wi18_content/DSMCER/atomsradii.csv)
* testing data: [https://raw.githubusercontent.com/UWDIRECT/UWDIRECT.github.io/master/Wi18_content/DSMCER/testing.csv](https://raw.githubusercontent.com/UWDIRECT/UWDIRECT.github.io/master/Wi18_content/DSMCER/testing.csv)
From this article in [Scientific Reports](http://www.nature.com/articles/srep13285)
#### Now, let's make a new classifier object
Start with [LinearSVC](http://scikit-learn.org/stable/modules/generated/sklearn.svm.LinearSVC.html#sklearn.svm.LinearSVC)
### You can use the following function to see how your model is doing:
#### You and your partner should determine:
* Testing error rate
* Training error rate
Grab your code from the L9.MLIntro notebook!
#### With remaining time go through the cell below and look at graphs of the decision boundary vs K.
* See if you can use the graph to determine your **testing** error rate
* Could you also use the graph to determine your **training** error rate? (_open ended_)
This is code to visualize the decision boundary. Fix it up to use your classifier from above or better yet, try a nonlinear kernel and visualize that! Name your classifier `clf.predict` and this should just run.
```
# additional library we will use
from matplotlib.colors import ListedColormap
# just for convenience and similarity with sklearn tutorial
# I am going to assign our X and Y data to specific vectors
# this is not strictly needed and you could use elements df for the whole thing!
X=elements[['rWC','rCh']]
#this is a trick to turn our strings (type of element / class) into unique
#numbers. Play with this in a separate cell and make sure you know wth is
#going on!
levels,labels=pd.factorize(elements.Type)
y=levels
#This determines levelspacing for our color map and the colors themselves
h=0.02
cmap_light = ListedColormap(['#FFAAAA', '#AAFFAA', '#AAAAFF'])
cmap_bold = ListedColormap(['#FF0000', '#00FF00', '#0000FF'])
# in the sklearn tutorial two different weights are compared
# the decision between "uniform" and "distance" determines the probability
# weight. "uniform" is the version presented in class, you can change to
# distance
weights='uniform'
# Straight from the tutorial - quickly read and see if you know what these
# things are going - if you are < 5 min until end then you should skip this part
# Plot the decision boundary. For that, we will assign a color to each
# point in the mesh [x_min, x_max]x[y_min, y_max].
x_min, x_max = elements.rWC.min() - 0.1 , elements.rWC.max() + 0.1
y_min, y_max = elements.rCh.min() - 0.1 , elements.rCh.max() + 0.1
xx, yy = np.meshgrid(np.arange(x_min, x_max, h),
np.arange(y_min, y_max, h))
Z = clf.predict(np.c_[xx.ravel(), yy.ravel()])
# Put the result into a color plot
Z = Z.reshape(xx.shape)
plt.figure(figsize=(4,4));
plt.pcolormesh(xx, yy, Z, cmap=cmap_light)
# Plot also the training points
# This may be the 1st time you have seen how to color points by a 3rd vector
# In this case y ( see c=y in below statement ). This is very useful!
plt.scatter(X.rWC, X.rCh, c=y, cmap=cmap_bold)
# Set limits and lebels
plt.xlim(xx.min(), xx.max())
plt.ylim(yy.min(), yy.max())
plt.xlabel('rWC')
plt.ylabel('rCh')
```
| github_jupyter |
# Varieties of Democracy (vDem)
Pemstein, Daniel, et al. "The V-Dem measurement model: latent variable analysis for cross-national and cross-temporal expert-coded data." V-Dem Working Paper 21 (2018).
Coppedge, Michael, et al. "V-Dem Codebook v11." (2021).
```
import sys
sys.path.append('..')
import os
import pandas as pd
import numpy as np
import time
import torch
from torch.utils.data import DataLoader, TensorDataset, random_split
from pytorch_lightning.callbacks import ModelCheckpoint
from pytorch_lightning import Trainer, seed_everything
from sklearn.ensemble import RandomForestClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.model_selection import StratifiedKFold, train_test_split
from sklearn.feature_selection import mutual_info_classif, chi2
from sklearn.linear_model import LassoCV
import matplotlib.pyplot as plt
import seaborn as sns
from torch_explain.models.explainer import Explainer
from torch_explain.logic.metrics import formula_consistency
from experiments.data.load_datasets import load_vDem
```
## Import v-Dem dataset
```
x, c, y, concept_names = load_vDem()
dataset_xc = TensorDataset(x, c)
dataset_cy = TensorDataset(c, y)
train_size = int(len(dataset_cy) * 0.5)
val_size = (len(dataset_cy) - train_size) // 2
test_size = len(dataset_cy) - train_size - val_size
train_data, val_data, test_data = random_split(dataset_cy, [train_size, val_size, test_size])
train_loader = DataLoader(train_data, batch_size=train_size)
val_loader = DataLoader(val_data, batch_size=val_size)
test_loader = DataLoader(test_data, batch_size=test_size)
n_concepts = next(iter(train_loader))[0].shape[1]
n_classes = 2
print(concept_names)
print(n_concepts)
print(n_classes)
```
## 10-fold cross-validation with explainer network
```
seed_everything(42)
base_dir = f'./results/vdem/explainer'
os.makedirs(base_dir, exist_ok=True)
n_splits = 5
skf = StratifiedKFold(n_splits=n_splits, shuffle=True, random_state=42)
results_list = []
feature_selection = []
explanations = {i: [] for i in range(n_classes)}
for split, (trainval_index, test_index) in enumerate(skf.split(x.cpu().detach().numpy(),
y.argmax(dim=1).cpu().detach().numpy())):
print(f'Split [{split + 1}/{n_splits}]')
x_trainval, x_test = torch.FloatTensor(x[trainval_index]), torch.FloatTensor(x[test_index])
c_trainval, c_test = torch.FloatTensor(c[trainval_index]), torch.FloatTensor(c[test_index])
y_trainval, y_test = torch.FloatTensor(y[trainval_index]), torch.FloatTensor(y[test_index])
x_train, x_val, c_train, c_val, y_train, y_val = train_test_split(x_trainval, c_trainval, y_trainval,
test_size=0.2, random_state=42)
print(f'{len(y_train)}/{len(y_val)}/{len(y_test)}')
# train X->C
train_data_xc = TensorDataset(x_train, c_train)
val_data_xc = TensorDataset(x_val, c_val)
test_data_xc = TensorDataset(x_test, c_test)
train_loader_xc = DataLoader(train_data_xc, batch_size=train_size)
val_loader_xc = DataLoader(val_data_xc, batch_size=val_size)
test_loader_xc = DataLoader(test_data_xc, batch_size=test_size)
checkpoint_callback_xc = ModelCheckpoint(dirpath=base_dir, monitor='val_loss', save_top_k=1)
trainer_xc = Trainer(max_epochs=200, gpus=1, auto_lr_find=True, deterministic=True,
check_val_every_n_epoch=1, default_root_dir=base_dir+'_xc',
weights_save_path=base_dir, callbacks=[checkpoint_callback_xc])
model_xc = Explainer(n_concepts=x.shape[1], n_classes=c.shape[1], l1=0, lr=0.01,
explainer_hidden=[100, 50], temperature=5000, loss=torch.nn.BCEWithLogitsLoss())
trainer_xc.fit(model_xc, train_loader_xc, val_loader_xc)
model_xc.freeze()
c_train_pred = model_xc.model(x_train)
c_val_pred = model_xc.model(x_val)
c_test_pred = model_xc.model(x_test)
# train C->Y
train_data = TensorDataset(c_train_pred, y_train)
val_data = TensorDataset(c_val_pred, y_val)
test_data = TensorDataset(c_test_pred, y_test)
train_loader = DataLoader(train_data, batch_size=train_size)
val_loader = DataLoader(val_data, batch_size=val_size)
test_loader = DataLoader(test_data, batch_size=test_size)
checkpoint_callback = ModelCheckpoint(dirpath=base_dir, monitor='val_loss', save_top_k=1)
trainer = Trainer(max_epochs=200, gpus=1, auto_lr_find=True, deterministic=True,
check_val_every_n_epoch=1, default_root_dir=base_dir,
weights_save_path=base_dir, callbacks=[checkpoint_callback])
model = Explainer(n_concepts=n_concepts, n_classes=n_classes, l1=1e-5, lr=0.01,
explainer_hidden=[20, 20], temperature=5)
start = time.time()
trainer.fit(model, train_loader, val_loader)
model.freeze()
model_results = trainer.test(model, test_dataloaders=test_loader)
for j in range(n_classes):
n_used_concepts = sum(model.model[0].concept_mask[j] > 0.5)
print(f"Extracted concepts: {n_used_concepts}")
results, f = model.explain_class(val_loader, train_loader, test_loader,
topk_explanations=500,
#max_minterm_complexity=7,
concept_names=concept_names)
end = time.time() - start
results['model_accuracy'] = model_results[0]['test_acc']
results['extraction_time'] = end
results_list.append(results)
extracted_concepts = []
all_concepts = model.model[0].concept_mask[0] > 0.5
common_concepts = model.model[0].concept_mask[0] > 0.5
for j in range(n_classes):
n_used_concepts = sum(model.model[0].concept_mask[j] > 0.5)
print(f"Extracted concepts: {n_used_concepts}")
print(f"Explanation: {f[j]['explanation']}")
print(f"Explanation accuracy: {f[j]['explanation_accuracy']}")
explanations[j].append(f[j]['explanation'])
extracted_concepts.append(n_used_concepts)
all_concepts += model.model[0].concept_mask[j] > 0.5
common_concepts *= model.model[0].concept_mask[j] > 0.5
results['extracted_concepts'] = np.mean(extracted_concepts)
results['common_concepts_ratio'] = sum(common_concepts) / sum(all_concepts)
# compare against standard feature selection
i_mutual_info = mutual_info_classif(c_trainval, y_trainval[:, 1])
i_chi2 = chi2(c_trainval, y_trainval[:, 1])[0]
i_chi2[np.isnan(i_chi2)] = 0
lasso = LassoCV(cv=5, random_state=0).fit(c_trainval, y_trainval[:, 1])
i_lasso = np.abs(lasso.coef_)
i_mu = model.model[0].concept_mask[1]
df = pd.DataFrame(np.hstack([
i_mu.numpy(),
i_mutual_info/np.max(i_mutual_info),
i_chi2/np.max(i_chi2),
i_lasso/np.max(i_lasso),
]).T, columns=['feature importance'])
df['method'] = 'explainer'
df.iloc[n_concepts:, 1] = 'MI'
df.iloc[n_concepts*2:, 1] = 'CHI2'
df.iloc[n_concepts*3:, 1] = 'Lasso'
df['feature'] = np.hstack([np.arange(0, n_concepts)] * 4)
feature_selection.append(df)
consistencies = []
for j in range(n_classes):
consistencies.append(formula_consistency(explanations[j]))
explanation_consistency = np.mean(consistencies)
feature_selection = pd.concat(feature_selection, axis=0)
np.hstack([np.arange(0, n_concepts)] * 4).shape
df[df['method']=='explainer'].shape
model_xc.model[0].alpha.shape
next(iter(train_loader_xc))[1].shape
```
## Print results
```
plt.figure(figsize=[4, 8])
ax = sns.barplot(y=feature_selection['feature'], x=feature_selection.iloc[:, 0],
hue=feature_selection['method'], orient='h', errwidth=0.5, errcolor='k')
plt.tight_layout()
plt.savefig(os.path.join(base_dir, 'barplot_vdem.png'))
plt.savefig(os.path.join(base_dir, 'barplot_vdem.pdf'))
plt.show()
plt.figure(figsize=[6, 4])
sns.boxplot(x=feature_selection.iloc[:, 1], y=feature_selection.iloc[:, 0])
plt.tight_layout()
plt.savefig(os.path.join(base_dir, 'boxplot_vdem.png'))
plt.savefig(os.path.join(base_dir, 'boxplot_vdem.pdf'))
plt.show()
results_df = pd.DataFrame(results_list)
results_df['explanation_consistency'] = explanation_consistency
results_df.to_csv(os.path.join(base_dir, 'results_aware_vdem.csv'))
results_df
results_df.mean()
results_df.sem()
```
## Compare with out-of-the-box models
```
dt_scores, rf_scores = [], []
for split, (trainval_index, test_index) in enumerate(skf.split(x.cpu().detach().numpy(), y.argmax(dim=1).cpu().detach().numpy())):
print(f'Split [{split + 1}/{n_splits}]')
x_trainval, x_test = x[trainval_index], x[test_index]
y_trainval, y_test = y[trainval_index].argmax(dim=1), y[test_index].argmax(dim=1)
dt_model = DecisionTreeClassifier(max_depth=5, random_state=split)
dt_model.fit(x_trainval, y_trainval)
dt_scores.append(dt_model.score(x_test, y_test))
rf_model = RandomForestClassifier(random_state=split)
rf_model.fit(x_trainval, y_trainval)
rf_scores.append(rf_model.score(x_test, y_test))
print(f'Random forest scores: {np.mean(rf_scores)} (+/- {np.std(rf_scores)})')
print(f'Decision tree scores: {np.mean(dt_scores)} (+/- {np.std(dt_scores)})')
print(f'Mu net scores (model): {results_df["model_accuracy"].mean()} (+/- {results_df["model_accuracy"].std()})')
print(f'Mu net scores (exp): {results_df["explanation_accuracy"].mean()} (+/- {results_df["explanation_accuracy"].std()})')
```
| github_jupyter |
```
def scope_test():
def do_local():
spam = 'lcoal spam'
def non_local():
nonlocal spam
spam = 'nonlocal spam'
def do_global():
global spam
spam = 'global spam'
spam = 'test spam'
do_local()
print("Spam value after local assignment:", spam)
non_local()
print("Spam value after nonlocal assignment:", spam)
do_global()
print("Spam value after global assignment:", spam)
#spam = 'Global_Initial_Spam'
#print('Initially, the spam value is:', spam)
scope_test()
print('In global, the spam value is:', spam)
```
Note how the local assignment (which is default) didn’t change scope_test’s binding of spam. The nonlocal assignment changed scope_test’s binding of spam, and the global assignment changed the module-level binding. You can also see that there was no previous binding for spam before the global assignment.
```
class MyClass:
'''A simple example class'''
i = 12345
'''second paragraph'''
def f(self):
return 'hello world'
print(MyClass.i)
print(MyClass.f(1))
MyClass.__doc__
class MyClass:
'''A simple example class'''
i = 12345
def f(self):
print('f() func has been executed')
return 'hello world'
#class instantiation automatically invokes __init__() for the newly-created class instance.
def __init__(self):
print('__init__() func has been executed')
x = MyClass()
class Complex:
def __init__(self, realpart, imagpart):
self.r = realpart
self.i = imagpart
x = Complex(3.0, -4.5)
print(x.r, x.i)
x.counter = 1
while x.counter < 10:
x.counter = x.counter * 2
print(x.counter)
del x.counter
x.arbitraryvalue = 1
print(type(x.arbitraryvalue))
print(type(x.__init__))
while x.arbitraryvalue < 10:
x.arbitraryvalue = x.arbitraryvalue * 2
print(x.arbitraryvalue)
del x.arbitraryvalue
class Dog:
kind = 'canine'
# tricks = []
def __init__(self, name):
self.name = name
self.tricks = []
def add_trick(self, trick):
self.tricks.append(trick)
d = Dog('Fido')
e = Dog('Buddy')
print(d.kind+'\n'+d.name)
d.add_trick('roll over')
e.add_trick('play dead')
print(d.tricks)
print(e.tricks)
class Mapping:
def __init__(self, iterable):
self.items_list = []
self.__update(iterable)
def update(self, iterable):
for item in iterable:
self.items_list.append(item)
# private copy of original update() method
__update = update
class MappingSubclass(Mapping):
def update(self, keys, values):
# provides new signature for update()
# but does not break __init__()
for item in zip(keys, values):
self.items_list.append(item)
```
Sometimes it is useful to have a data type similar to the Pascal “record” or C “struct”, bundling together a few named data items. An empty class definition will do nicely:
```
#define an empty class
class Employee:
pass
john = Employee() # create an empty employee record
#fill the fields of the record
john.name = 'John Doe'
john.dept = 'computer lab'
john.salary = 1000
s = 'abc'
it = iter(s)
it
print(next(it))
print(next(it))
print(next(it))
print(next(it))
class Reverse:
'''Iterator for looping over a sequence backwards'''
def __init__(self, data):
self.data = data
self.index = len(data)
def __iter__(self):
return self
def __next__(self):
if self.index == 0:
raise StopIteration
self.index = self.index - 1
return self.data[self.index]
rev = Reverse('spam')
iter(rev)
for char in rev:
print(char)
def reverse(data):
for idx in range(len(data)-1, -1, -1):
yield data[idx]
reverse_result = ''
for char in reverse('golf'):
reverse_result += char
print(reverse_result)
```
| github_jupyter |
<h1>Lists in Python</h1>
<p><strong>Welcome!</strong> This notebook will teach you about the lists in the Python Programming Language. By the end of this lab, you'll know the basics list operations in Python, including indexing, list operations and copy/clone list.</p>
<h2>Table of Contents</h2>
<div class="alert alert-block alert-info" style="margin-top: 20px">
<ul>
<li>
<a href="#dataset">About the Dataset</a>
</li>
<li>
<a href="#list">Lists</a>
<ul>
<li><a href="index">Indexing</a></li>
<li><a href="content">List Content</a></li>
<li><a href="op">List Operations</a></li>
<li><a href="co">Copy and Clone List</a></li>
</ul>
</li>
<li>
<a href="#quiz">Quiz on Lists</a>
</li>
</ul>
<p>
Estimated time needed: <strong>15 min</strong>
</p>
</div>
<hr>
<h2 id="#dataset">About the Dataset</h2>
Imagine you received album recommendations from your friends and compiled all of the recommandations into a table, with specific information about each album.
The table has one row for each movie and several columns:
- **artist** - Name of the artist
- **album** - Name of the album
- **released_year** - Year the album was released
- **length_min_sec** - Length of the album (hours,minutes,seconds)
- **genre** - Genre of the album
- **music_recording_sales_millions** - Music recording sales (millions in USD) on [SONG://DATABASE](http://www.song-database.com/)
- **claimed_sales_millions** - Album's claimed sales (millions in USD) on [SONG://DATABASE](http://www.song-database.com/)
- **date_released** - Date on which the album was released
- **soundtrack** - Indicates if the album is the movie soundtrack (Y) or (N)
- **rating_of_friends** - Indicates the rating from your friends from 1 to 10
<br>
<br>
The dataset can be seen below:
<font size="1">
<table font-size:xx-small style="width:100%">
<tr>
<th>Artist</th>
<th>Album</th>
<th>Released</th>
<th>Length</th>
<th>Genre</th>
<th>Music recording sales (millions)</th>
<th>Claimed sales (millions)</th>
<th>Released</th>
<th>Soundtrack</th>
<th>Rating (friends)</th>
</tr>
<tr>
<td>Michael Jackson</td>
<td>Thriller</td>
<td>1982</td>
<td>00:42:19</td>
<td>Pop, rock, R&B</td>
<td>46</td>
<td>65</td>
<td>30-Nov-82</td>
<td></td>
<td>10.0</td>
</tr>
<tr>
<td>AC/DC</td>
<td>Back in Black</td>
<td>1980</td>
<td>00:42:11</td>
<td>Hard rock</td>
<td>26.1</td>
<td>50</td>
<td>25-Jul-80</td>
<td></td>
<td>8.5</td>
</tr>
<tr>
<td>Pink Floyd</td>
<td>The Dark Side of the Moon</td>
<td>1973</td>
<td>00:42:49</td>
<td>Progressive rock</td>
<td>24.2</td>
<td>45</td>
<td>01-Mar-73</td>
<td></td>
<td>9.5</td>
</tr>
<tr>
<td>Whitney Houston</td>
<td>The Bodyguard</td>
<td>1992</td>
<td>00:57:44</td>
<td>Soundtrack/R&B, soul, pop</td>
<td>26.1</td>
<td>50</td>
<td>25-Jul-80</td>
<td>Y</td>
<td>7.0</td>
</tr>
<tr>
<td>Meat Loaf</td>
<td>Bat Out of Hell</td>
<td>1977</td>
<td>00:46:33</td>
<td>Hard rock, progressive rock</td>
<td>20.6</td>
<td>43</td>
<td>21-Oct-77</td>
<td></td>
<td>7.0</td>
</tr>
<tr>
<td>Eagles</td>
<td>Their Greatest Hits (1971-1975)</td>
<td>1976</td>
<td>00:43:08</td>
<td>Rock, soft rock, folk rock</td>
<td>32.2</td>
<td>42</td>
<td>17-Feb-76</td>
<td></td>
<td>9.5</td>
</tr>
<tr>
<td>Bee Gees</td>
<td>Saturday Night Fever</td>
<td>1977</td>
<td>1:15:54</td>
<td>Disco</td>
<td>20.6</td>
<td>40</td>
<td>15-Nov-77</td>
<td>Y</td>
<td>9.0</td>
</tr>
<tr>
<td>Fleetwood Mac</td>
<td>Rumours</td>
<td>1977</td>
<td>00:40:01</td>
<td>Soft rock</td>
<td>27.9</td>
<td>40</td>
<td>04-Feb-77</td>
<td></td>
<td>9.5</td>
</tr>
</table></font>
<hr>
<h2 id="list">Lists</h2>
<h3 id="index">Indexing</h3>
We are going to take a look at lists in Python. A list is a sequenced collection of different objects such as integers, strings, and other lists as well. The address of each element within a list is called an <b>index</b>. An index is used to access and refer to items within a list.
<img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Chapter%202/Images/ListsIndex.png" width="1000" />
To create a list, type the list within square brackets <b>[ ]</b>, with your content inside the parenthesis and separated by commas. Let’s try it!
```
# Create a list
L = ["Michael Jackson", 10.1, 1982]
L
```
We can use negative and regular indexing with a list :
<img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Chapter%202/Images/ListsNeg.png" width="1000" />
```
# Print the elements on each index
print('the same element using negative and positive indexing:\n Postive:',L[0],
'\n Negative:' , L[-3] )
print('the same element using negative and positive indexing:\n Postive:',L[1],
'\n Negative:' , L[-2] )
print('the same element using negative and positive indexing:\n Postive:',L[2],
'\n Negative:' , L[-1] )
```
<h3 id="content">List Content</h3>
Lists can contain strings, floats, and integers. We can nest other lists, and we can also nest tuples and other data structures. The same indexing conventions apply for nesting:
```
# Sample List
["Michael Jackson", 10.1, 1982, [1, 2], ("A", 1)]
```
<h3 id="op">List Operations</h3>
We can also perform slicing in lists. For example, if we want the last two elements, we use the following command:
```
# Sample List
L = ["Michael Jackson", 10.1,1982,"MJ",1]
L
```
<img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Chapter%202/Images/ListsSlice.png" width="1000">
```
# List slicing
L[3:5]
```
We can use the method <code>extend</code> to add new elements to the list:
```
# Use extend to add elements to list
L = [ "Michael Jackson", 10.2]
L.extend(['pop', 10])
L
```
Another similar method is <code>append</code>. If we apply <code>append</code> instead of <code>extend</code>, we add one element to the list:
```
# Use append to add elements to list
L = [ "Michael Jackson", 10.2]
L.append(['pop', 10])
L
```
Each time we apply a method, the list changes. If we apply <code>extend</code> we add two new elements to the list. The list <code>L</code> is then modified by adding two new elements:
```
# Use extend to add elements to list
L = [ "Michael Jackson", 10.2]
L.extend(['pop', 10])
L
```
If we append the list <code>['a','b']</code> we have one new element consisting of a nested list:
```
# Use append to add elements to list
L.append(['a','b'])
L
```
As lists are mutable, we can change them. For example, we can change the first element as follows:
```
# Change the element based on the index
A = ["disco", 10, 1.2]
print('Before change:', A)
A[0] = 'hard rock'
print('After change:', A)
```
We can also delete an element of a list using the <code>del</code> command:
```
# Delete the element based on the index
print('Before change:', A)
del(A[0])
print('After change:', A)
```
We can convert a string to a list using <code>split</code>. For example, the method <code>split</code> translates every group of characters separated by a space into an element in a list:
```
# Split the string, default is by space
'hard rock'.split()
```
We can use the split function to separate strings on a specific character. We pass the character we would like to split on into the argument, which in this case is a comma. The result is a list, and each element corresponds to a set of characters that have been separated by a comma:
```
# Split the string by comma
'A,B,C,D'.split(',')
```
<h3 id="co">Copy and Clone List</h3>
When we set one variable <b>B</b> equal to <b>A</b>; both <b>A</b> and <b>B</b> are referencing the same list in memory:
```
# Copy (copy by reference) the list A
A = ["hard rock", 10, 1.2]
B = A
print('A:', A)
print('B:', B)
```
<img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Chapter%202/Images/ListsRef.png" width="1000" align="center">
Initially, the value of the first element in <b>B</b> is set as hard rock. If we change the first element in <b>A</b> to <b>banana</b>, we get an unexpected side effect. As <b>A</b> and <b>B</b> are referencing the same list, if we change list <b>A</b>, then list <b>B</b> also changes. If we check the first element of <b>B</b> we get banana instead of hard rock:
```
# Examine the copy by reference
print('B[0]:', B[0])
A[0] = "banana"
print('B[0]:', B[0])
```
This is demonstrated in the following figure:
<img src = "https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Chapter%202/Images/ListsRefGif.gif" width="1000" />
You can clone list **A** by using the following syntax:
```
# Clone (clone by value) the list A
B = A[:]
B
```
Variable **B** references a new copy or clone of the original list; this is demonstrated in the following figure:
<img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Chapter%202/Images/ListsVal.gif" width="1000" />
Now if you change <b>A</b>, <b>B</b> will not change:
```
print('B[0]:', B[0])
A[0] = "hard rock"
print('B[0]:', B[0])
```
<h2 id="quiz">Quiz on List</h2>
Create a list <code>a_list</code>, with the following elements <code>1</code>, <code>hello</code>, <code>[1,2,3]</code> and <code>True</code>.
```
# Write your code below and press Shift+Enter to execute
```
Double-click <b>here</b> for the solution.
<!-- Your answer is below:
a_list = [1, 'hello', [1, 2, 3] , True]
a_list
-->
Find the value stored at index 1 of <code>a_list</code>.
```
# Write your code below and press Shift+Enter to execute
```
Double-click <b>here</b> for the solution.
<!-- Your answer is below:
a_list[1]
-->
Retrieve the elements stored at index 1, 2 and 3 of <code>a_list</code>.
```
# Write your code below and press Shift+Enter to execute
```
Double-click <b>here</b> for the solution.
<!-- Your answer is below:
a_list[1:4]
-->
Concatenate the following lists <code>A = [1, 'a']</code> and <code>B = [2, 1, 'd']</code>:
```
# Write your code below and press Shift+Enter to execute
```
Double-click <b>here</b> for the solution.
<!-- Your answer is below:
A = [1, 'a']
B = [2, 1, 'd']
A + B
-->
| github_jupyter |
## Module/Variable/Data setup
```
%run ../src/python/helpers.py
%matplotlib inline
from numpy import nan, arange
from pandas import read_feather
import seaborn as sns
import ipywidgets as w
from quilt.data.hsiaoyi0504 import aeolus_top5drugs
#VARIABLES
cl = ['atc_1st','atc_2nd','atc_3rd','atc_4th','drug_concept_name']
data = read_feather(aeolus_top5drugs.aeolus_top5drugs._data())
plot_settings()
```
## For a drug class, how many unique adverse reactions are reported?
```
d = dropdown(cl)
d
```
classes in above chosen drug class
```
out= 'outcome_concept_name'
grpd = data.groupby([d.label,out])[out].count().sort_values(ascending=False)
grpd.name = 'n'
grpd2 = grpd.groupby(level=d.label)
series = grpd2.count()
p = plot(series,kind='barh')
q = p.set_title(p.get_ylabel().upper(),rotation=0,weight='bold')
q = p.set_ylabel("")
q = p.set_xlabel("Number of unique adverse reactions",weight="bold")
```
## What are the top adverse reactions in the chosen drug class?
```
ds = dropdown(series.index)
ds
i = w.IntSlider(min=1,max=10,step=1,value=5)
i
```
indices for most frequent ADRs
```
sub = data.query('{0} in @ds.label'.format(d.label)).groupby([d.label,out])[out].count().sort_values(ascending=False).head(i.value)
sub.name= "n"
sub = sub.reset_index()
p = sns.barplot(y=d.label,x='n',hue=out,data=sub,orient='h')
p.legend(bbox_to_anchor=(1.7,1))
q = p.set_ylabel('')
```
## How do the ADRs break down by report year?
```
outs = sub.iloc[:,1].values
sub2 = data.query('({0}==@ds.label) & ({1} in @outs)'.format(d.label,out))
series2 = freqXbyY(sub2,'report_year','id')
p = plot(series2,kind='bar')
q = p.set_title(p.get_ylabel().upper(),rotation=0,weight='bold')
q = p.set_ylabel(p.get_ylabel(),rotation=0)
q = p.set_xlabel("Number of Reports",weight="bold")
```
## subset data by chosen year(s)
```
labels = series2.index
mds = w.SelectMultiple(options = labels ,value = tuple(labels))
mds
```
## How many are reported within these ADRs across the sexes?
```
#plot variables for filtering/wrangling
bars = 'gender_code'
x = 'report_year'
count = 'id'
dat = clean_gender(sub2).query('report_year in @mds.label')
#main
sub_dat = dat[[bars,x,count]]
plot_sub_dat = sub_dat.groupby([bars,x]).count().reset_index(level=bars).pivot(columns=bars)
plot_sub_dat.columns = plot_sub_dat.columns.droplevel(level=0)
plot_sub_dat.plot.bar()
```
## How many are reported within this class across ages?
```
#plot variables for filtering/wrangling
grp = 'age_cat'
#main
dat[[d.label,grp]].groupby([grp]).count().plot.bar()
```
## How many are reported within this class across ages for each sex?
```
#plot variables for filtering/wrangling
bars = 'gender_code'
x = 'age_cat'
count = 'id'
#want to filter dataset for M/F gender and if report year was clicked or selected
sub_dat = clean_gender(dat)[[bars,x,count]]
#main
plot_sub_dat = sub_dat.groupby([bars,x]).count().reset_index(level=bars).pivot(columns=bars)
plot_sub_dat.columns = plot_sub_dat.columns.droplevel(level=0)
plot_sub_dat.plot.bar()
```
| github_jupyter |
# Scraper Development
This notebook shows examples of errors and other dev considerations for the construction or modification of the scraper.
```
import time
import numpy as np
import pandas as pd
from vcue import web
from bs4 import BeautifulSoup
driver = web.launch_driver()
# In Japanese
driver.get('https://www.id.nlbc.go.jp/CattleSearch/search/agreement')
connected = ds.connection_check(driver)
```
Find the last digit given an ID
```
import random_id_generator as rig
str(111425891)+str(rig.get_last_digit(str(111425891)))
soul = '1114258912'
ele = driver.find_elements_by_name('txtIDNO')[0]
ele.send_keys(str(soul))
driver.find_elements_by_name('method:doSearch')[0].click()
```
**Detecting the number of tables on the page will tell you if you found data relevant to the ID you entered**:
```
soup = BeautifulSoup(driver.page_source)
# Detecting "Enable Cookies" is the way to tell if the connection was cut.
lens = [len(list(table.parents)) for table in soup.find_all('table')]
tables = [table for table in soup.find_all('table') if len(list(table.parents))==max(lens)]
```
## <font color=red>Error Examples in Japanese</font>
```
# Sample error with connection/cookies
tables[0].text.strip()
# '1114258912' - doesn't exist '入力された個体識別番号をお確かめくださいますようお願いします'
# You have to check if the list of elements is the right size first
soup.find_all('span', {'class':'nor'})[2].text.strip()
# example error short number, should not appear.
# You have to check that the list of error messages is positive
soup.find_all('ul', {'class':'error_message'})[0].text.strip()
soup.find_all('ul', {'class':'error_message'})[0].text.strip()
```
## <font color=red> Example of Scraper Output</font>
```
dataset
complete_data
```
### Testing List Differences
```
import pandas as pd
import random
data = pd.read_csv('cattle.csv', dtype='O')
data['IDS'] = data['id0'].astype(str)+data['id1'].astype(str)+data['id2'].astype(str)
master_sample_ids = list(master_sample['id'].values)
scraped = random.sample(master_sample_ids,1000)
print(len(master_sample_ids))
print(len(scraped))
%%time
cleaned1 = [i for i in master_sample_ids if i not in scraped]
%%time
seta = set(master_sample_ids)
setb = set(scraped)
cleaned2 = seta.difference(setb)
print(len(cleaned1))
print(len(cleaned2))
assert set(cleaned1) == cleaned2
type(cleaned2)
```
# DEV SPACE
Use the blocks below to develop and test new functions
```
from supply_chain_mapping import supply_chain_data_scraper as scd
from supply_chain_mapping import random_id_generator as rig
# Generate a random ID
import random
def get_random_id():
randidbase = '1' + str(random.randint(10000000, 100000000))
randid = randidbase + str(rig.get_last_digit(randidbase))
return randid
batch_size = 100
complete_data, final_data, failed_data = scd.collect_temporary(
return_data = True,
search_prefix='') # Get all the collected data
collected_ids = list(complete_data['Individual Identification Number']) + list(final_data['個体識別番号'].values) + list(failed_data['ID'].values)
id_batch = []
idbl = len(id_batch)
# Until we've generated a batch of size batch_size
while idbl < batch_size:
# First create a list of ids of the desired length
while len(id_batch) < batch_size:
id_batch.append(get_random_id())
# Remove repeated Ids by turning it into a set and compare the IDs to the list of existing collected IDs
seta = set(id_batch)
setb = set(collected_ids)
id_batch = seta.difference(setb)
id_batch = list(id_batch)
idbl = len(id_batch)
```
| github_jupyter |
## 9.5 Partial-Redundancy Elimination
### 9.5.1
> For the flow graph in Fig. 9.37:
> ```
+-----+
|ENTRY|
+--+--+
|
|
|
+----v----+
| = x+y | B1
+----+----+
|
|
|
+----v----+
+-------> x = | B2
| +----+----+
| |
| |
| |
| +----v----+
+-------+ = x+y | B3
+----+----+
|
|
|
+----v----+
| = x+y | B4
+----+----+
|
|
|
+--v-+
|EXIT|
+----+
```
> a) Compute $anticipated$ for the beginning and end of each block.
> b) Compute $available$ for the beginning and end of each block.
> c) Compute $earliest$ for each block.
> d) Compute $postponable$ for the beginning and end of each block.
> e) Compute $used$ for the beginning and end of each block.
> f) Compute $latest$ for each block.
| Block | $anticipated$ | $available$ | $earliest$ | $postponable$ | $latest$ | $used$ |
|:-------:|:----------------|:--------------|:------------:|:----------------|:----------:|:---------|
| ENTRY | $\begin{array}{rll}\text{OUT}[\text{ENTRY}] &=& \{ x+y \} \end{array}$ | $\begin{array}{rll}\text{OUT}[\text{ENTRY}] &=& \emptyset \end{array}$ | $\emptyset$ | $\begin{array}{rll}\text{OUT}[\text{ENTRY}] &=& \emptyset \end{array}$ | $\emptyset$ | $\begin{array}{rll}\text{OUT}[\text{ENTRY}] &=& \emptyset \end{array}$ |
| $B_1$ | $\begin{array}{rll}\text{IN}[B_1] &=& \{ x+y \} \\ \text{OUT}[B_1] &=& \emptyset \end{array}$ | $\begin{array}{rll}\text{IN}[B_1] &=& \emptyset \\ \text{OUT}[B_1] &=& \{ x+y \} \end{array}$ | $\{ x + y \}$ | $\begin{array}{rll}\text{IN}[B_1] &=& \emptyset \\ \text{OUT}[B_1] &=& \emptyset \end{array}$ | $\{ x + y \}$ | $\begin{array}{rll}\text{IN}[B_1] &=& \emptyset \\ \text{OUT}[B_1] &=& \emptyset \end{array}$ |
| $B_2$ | $\begin{array}{rll}\text{IN}[B_2] &=& \emptyset \\ \text{OUT}[B_2] &=& \{ x+y \} \end{array}$ | $\begin{array}{rll}\text{IN}[B_2] &=& \{ x+y \} \\ \text{OUT}[B_2] &=& \emptyset \end{array}$ | $\emptyset$ | $\begin{array}{rll}\text{IN}[B_2] &=& \emptyset \\ \text{OUT}[B_2] &=& \emptyset \end{array}$ | $\emptyset$ | $\begin{array}{rll}\text{IN}[B_2] &=& \emptyset \\ \text{OUT}[B_2] &=& \emptyset \end{array}$ |
| $B_3$ | $\begin{array}{rll}\text{IN}[B_3] &=& \{ x+y \} \\ \text{OUT}[B_3] &=& \emptyset \end{array}$ | $\begin{array}{rll}\text{IN}[B_3] &=& \emptyset \\ \text{OUT}[B_3] &=& \{ x+y \} \end{array}$ | $\{ x + y \}$ | $\begin{array}{rll}\text{IN}[B_3] &=& \emptyset \\ \text{OUT}[B_3] &=& \emptyset \end{array}$ | $\{ x + y \}$ | $\begin{array}{rll}\text{IN}[B_3] &=& \emptyset \\ \text{OUT}[B_3] &=& \{ x+y \} \end{array}$ |
| $B_4$ | $\begin{array}{rll}\text{IN}[B_4] &=& \{ x+y \} \\ \text{OUT}[B_4] &=& \emptyset \end{array}$ | $\begin{array}{rll}\text{IN}[B_4] &=& \{ x+y \} \\ \text{OUT}[B_4] &=& \{ x+y \} \end{array}$ | $\emptyset$ | $\begin{array}{rll}\text{IN}[B_4] &=& \emptyset \\ \text{OUT}[B_4] &=& \emptyset \end{array}$ | $\emptyset$ | $\begin{array}{rll}\text{IN}[B_4] &=& \{ x+y \}\\ \text{OUT}[B_4] &=& \emptyset \end{array}$ |
| EXIT | $\begin{array}{rll}\text{IN}[\text{EXIT}] &=& \emptyset \end{array}$ | $\begin{array}{rll}\text{IN}[\text{EXIT}] &=& \{ x+y \} \end{array}$ | $\emptyset$ | $\begin{array}{rll}\text{IN}[\text{EXIT}] &=& \emptyset \end{array}$ | $\emptyset$ | $\begin{array}{rll}\text{IN}[\text{EXIT}] &=& \emptyset \end{array}$ |
> g) Introduce temporary variable $t$; show where it is computed and where it is used.
```
+-----+
|ENTRY|
+--+--+
|
|
+----v----+
| = x+y | B1
+----+----+
|
|
|
+----v----+
+-------> x = | B2
| +----+----+
| |
| |
| +----v----+
| | t = x+y |
+-------+ = t | B3
+----+----+
|
|
|
+----v----+
| = t | B4
+----+----+
|
|
|
+--v-+
|EXIT|
+----+
```
| github_jupyter |
# Analyzing Mutations with Trained Models
In this tutorial, we will explore some of Selene's tools for exploring and interpreting sequence predictions.
We are generally interested in models that predict various output labels for an input biological sequence.
For instance, we might want to predict histone marks from the DNA sequence of a human genome.
Beyond recapitulating biological phenomena for known motifs, these models can make accurate predictions for arbitrary sequences.
An *in silico* mutagenesis experiment uses this fact to estimate how mutations in a sequence might change our predictions for it.
In this tutorial, we will use selene's suite of tools for *in silico* mutagenesis to predict the effects of single nucleotide polymorphisms (SNPs).
## Download the data
Download the compressed data from [here](https://zenodo.org/record/1319784):
```sh
wget https://zenodo.org/record/2206957/files/selene_analyzing_mutations_tutorial.tar.gz
```
Extract it and `mv` the files from the extracted directory `selene_analyzing_mutations_tutorial` to the current directory:
```sh
tar -xzvf selene_analyzing_mutations_tutorial.tar.gz
mv selene_analyzing_mutations_tutorial/* .
```
We will analyze the genomic sequences in `sequences.fasta` with an *in silico* mutagenesis experiment.
There should be a file called `example_deeperdeepsea.pth.tar`, containing a trained model weights file (generated when this model was trained using Selene). This model weights file uses the architecture in [`utils/example_model.py`](https://github.com/FunctionLab/selene/blob/master/selene/utils/example_model.py) named DeeperDeepSEA, which we also import in this tutorial. `distinct_features.txt` contains the names of the labels predicted by the model.
Finally, we include the output `*.tsv` files from _in silico_ mutagenesis in case you only want to run parts of the notebook.
## Load the trained model
The model weights file is used to set the weights of the initialized model architecture class, which is `DeeperDeepSEA` wrapped in the `NonStrandSpecific` class
```
%matplotlib inline
import os
import torch
from selene_sdk.utils import DeeperDeepSEA
from selene_sdk.utils import NonStrandSpecific
model_architecture = NonStrandSpecific(DeeperDeepSEA(1000, 919))
```
## *In silico* mutagenesis
In an *in silico* mutagenesis experiment, we have some input sequence $S$ of length $L$ which we would like to mutate.
In the simplest case of *in silico* mutagenesis, we make predict the effects of each individual $3L$ SNPs that are possible in $S$.
The *in silico* mutagenesis tools in selene allow us to do this easily, and can also be used to predict the effects of every pair/triplet/etc of SNPs in our input sequence.
We will now load the list of features from file, and perform an *in silico* mutagenesis experiment on the sequences in `sequences.fasta`.
This will give us predictions for the effects of every possible SNP that could occur in our sequence.
Note that this may take a minute or two if you are not using a GPU.
```
from selene_sdk.predict import AnalyzeSequences
from selene_sdk.utils import load_features_list
features = load_features_list("distinct_features.txt")
analysis = AnalyzeSequences(
model_architecture,
"example_deeperdeepsea.pth.tar",
sequence_length=1000,
features=features,
use_cuda=False)
analysis.in_silico_mutagenesis_from_file("sequences.fasta",
save_data=["abs_diffs", "logits", "predictions"],
output_dir=".",
use_sequence_name=False)
```
There should now be three new files: `0_predictions.tsv`, `0_logits.tsv`, and `0_abs_diffs.tsv`.
They contain the results of the *in silico* mutagenesis experiments on the first record from `sequences.fasta`.
If there was second record in `sequences.fasta`, we would also see `1_predictions.tsv`, `1_logits.tsv`, and `1_abs_diffs.tsv`.
The first of these output files, `0_predictions.tsv`, contains the model predictions for each individual SNP in the *in silico* mutagenesis experiment.
The file `0_logits.tsv` contains the difference between the logits of the reference and SNP predictions.
Consider that we are predicting the $i^{th}$ label for a sequence $S'$ that has been produced by mutating a single base in $S$.
Then $p_i$ is the probability that the non-mutated sequence $S$ is a positive example of the $i^{th}$ feature, and $q_i$ is the probability that the mutated sequence $S'$ is a positive example of the $i^{th}$ feature.
Thus, the corresponding values in `0_logits.tsv` would be be $\mathrm{logit}(q_i) - \mathrm{logit}(p_i)$ where $\mathrm{logit}(x) = \log\left(\frac{x}{1 - x}\right) = \log(x) - \log(1 - x)$.
Finally, the file `0_abs_diffs.tsv` contains the absolute difference between the reference and SNP predictions, i.e. $\left|q_i - p_i\right|$.
## Visualizing *in silico* mutagenesis results
Before we visualize anything, we have to load the predictions for the *in silico* mutagenesis of the first sequence record from `sequences.fasta`. As stated above, these predictions are in `0_predictions.tsv`.
```
from selene_sdk.interpret import ISMResult
ism = ISMResult.from_file("0_predictions.tsv")
```
We now extract the array of predictions for one of the predicted labels, "K562|H3K27ac|None".
```
score_matrix = ism.get_score_matrix_for("K562|H3K27ac|None")[:50,]
```
The structure of the returned `score_matrix` is important, so let's take a moment to discuss it.
For a score matrix $\mathbf{X}$, the element in the $i^{th}$ row and $j^{th}$ column, $x_{i, j}$, is the prediction associated with mutating the $i^{th}$ base in the input sequence to the $j^{th}$ base in $[A, C, G, T]$.
If the $j^{th}$ base in $[A, C, G, T]$ is the same as the reference base, then $x_{i, j}$ will be the predicted value for the reference sequence.
### Plotting heatmaps
We will now plot this matrix as a heatmap, using the `heatmap` method in `selene_sdk.interpret`.
In the example below, we mask the bases that match the reference with cross-hatching.
```
import matplotlib.pyplot as plt
import selene_sdk.interpret
from selene_sdk.sequences import Genome
reference_encoding = Genome.sequence_to_encoding(ism.reference_sequence)[:50,] == 1.
figure, (ax) = plt.subplots(1, 1, figsize=(10, 4))
ax.patch.set(edgecolor="lightgrey", hatch="//")
selene_sdk.interpret.heatmap(score_matrix, mask=reference_encoding, cbar=True, ax=ax, linewidth=0.5)
```
### Plotting sequence logos
In the code below, we've plotted the matrix as a sequence logo using the `sequence_logo` method from `selene_sdk.interpret`.
The vertical ordering of the bases at each position can be controlled with the `order` argument to `sequence_logo`.
With `order="alpha"`, at the $i^{th}$ position along the horizontal axis, the height of the $j^{th}$ letter is the value $x_{i, j}$ from the score matrix.
However, it is more common to order the bases at each position in a sequence logo by the value of their prediction.
This behavior can be achieved with `order="value"`.
```
figure, (ax1, ax2) = plt.subplots(2, 1, figsize=(20, 6))
selene_sdk.interpret.sequence_logo(score_matrix, order="alpha", ax=ax1)
selene_sdk.interpret.sequence_logo(score_matrix, order="value", ax=ax2)
```
### Rescaling predictions
Besides visualization methods, `selene_sdk.interpret` contains tools for rescaling the plotted values.
Although we often want to plot the raw values as we did above, there are other derived values that we are interested in.
The method `rescale_score_matrix` in `selene_sdk.interpret` enables easy calculation of many of these values.
Rescaling with `rescale_score_matrix` can be performed on the individual bases at a loci, or each position in the sequence.
Position-wise scaling is controlled by the `position_scaling` argument to `rescale_score_matrix`, which may be either `"identity"`, `"probability"`, or `"max_effect"`.
The first of these, `"identity"`, performs no position-wise rescaling.
The second, `"probability"`, rescales the values so that the predictions for each base at a given position in the sequence sum to $1$.
Finally, `"max_effect"` scaling is nearly identical to that of `"probability"`, but values are rescaled so that the predictions for each base at a given position in the sequence sum to the difference between the largest and smallest predicted values.
Base-wise scaling is controlled by the `base_scaling` argument to `rescale_score_matrix`, which maybe either `"identity"`, `"probability"`, or `"max_effect"`.
The first, `"identity"`, does not rescale any values.
Since the inputs are assumed to be probabilities to begin with, `"probability"` does not rescale input values either.
Lastly, `"max_effect"` rescales the values so that the predictions for each base at a given position in the sequence is the difference between that prediction and the smallest prediction at that position.
```
rescaled_score_matrix = selene_sdk.interpret.rescale_score_matrix(score_matrix, base_scaling="max_effect")
figure, (ax1, ax2) = plt.subplots(2, 1, figsize=(20, 6))
selene_sdk.interpret.sequence_logo(rescaled_score_matrix, ax=ax1)
selene_sdk.interpret.sequence_logo(score_matrix, ax=ax2)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/PacktPublishing/Hands-On-Computer-Vision-with-PyTorch/blob/master/Chapter14/Siamese_networks.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
!pip install -q torch_snippets
from torch_snippets import *
!wget -q https://www.dropbox.com/s/ua1rr8btkmpqjxh/face-detection.zip
!unzip -q face-detection.zip
device = 'cuda' if torch.cuda.is_available() else 'cpu'
class SiameseNetworkDataset(Dataset):
def __init__(self, folder, transform=None, should_invert=True):
self.folder = folder
self.items = Glob(f'{self.folder}/*/*')
self.transform = transform
def __getitem__(self, ix):
itemA = self.items[ix]
person = fname(parent(itemA))
same_person = randint(2)
if same_person:
itemB = choose(Glob(f'{self.folder}/{person}/*', silent=True))
else:
while True:
itemB = choose(self.items)
if person != fname(parent(itemB)):
break
imgA = read(itemA)
imgB = read(itemB)
if self.transform:
imgA = self.transform(imgA)
imgB = self.transform(imgB)
return imgA, imgB, np.array([1-same_person])
def __len__(self):
return len(self.items)
from torchvision import transforms
trn_tfms = transforms.Compose([
transforms.ToPILImage(),
transforms.RandomHorizontalFlip(),
transforms.RandomAffine(5, (0.01,0.2),
scale=(0.9,1.1)),
transforms.Resize((100,100)),
transforms.ToTensor(),
transforms.Normalize((0.5), (0.5))
])
val_tfms = transforms.Compose([
transforms.ToPILImage(),
transforms.Resize((100,100)),
transforms.ToTensor(),
transforms.Normalize((0.5), (0.5))
])
trn_ds = SiameseNetworkDataset(folder="./data/faces/training/", transform=trn_tfms)
val_ds = SiameseNetworkDataset(folder="./data/faces/testing/", transform=val_tfms)
trn_dl = DataLoader(trn_ds, shuffle=True, batch_size=64)
val_dl = DataLoader(val_ds, shuffle=False, batch_size=64)
def convBlock(ni, no):
return nn.Sequential(
nn.Dropout(0.2),
nn.Conv2d(ni, no, kernel_size=3, padding=1, padding_mode='reflect'),
nn.ReLU(inplace=True),
nn.BatchNorm2d(no),
)
class SiameseNetwork(nn.Module):
def __init__(self):
super(SiameseNetwork, self).__init__()
self.features = nn.Sequential(
convBlock(1,4),
convBlock(4,8),
convBlock(8,8),
nn.Flatten(),
nn.Linear(8*100*100, 500), nn.ReLU(inplace=True),
nn.Linear(500, 500), nn.ReLU(inplace=True),
nn.Linear(500, 5)
)
def forward(self, input1, input2):
output1 = self.features(input1)
output2 = self.features(input2)
return output1, output2
class ContrastiveLoss(torch.nn.Module):
"""
Contrastive loss function.
Based on: http://yann.lecun.com/exdb/publis/pdf/hadsell-chopra-lecun-06.pdf
"""
def __init__(self, margin=2.0):
super(ContrastiveLoss, self).__init__()
self.margin = margin
def forward(self, output1, output2, label):
euclidean_distance = F.pairwise_distance(output1, output2, keepdim = True)
loss_contrastive = torch.mean((1-label) * torch.pow(euclidean_distance, 2) +
(label) * torch.pow(torch.clamp(self.margin - euclidean_distance, min=0.0), 2))
acc = ((euclidean_distance > 0.6) == label).float().mean()
return loss_contrastive, acc
def train_batch(model, data, optimizer, criterion):
imgsA, imgsB, labels = [t.to(device) for t in data]
optimizer.zero_grad()
codesA, codesB = model(imgsA, imgsB)
loss, acc = criterion(codesA, codesB, labels)
loss.backward()
optimizer.step()
return loss.item(), acc.item()
@torch.no_grad()
def validate_batch(model, data, criterion):
imgsA, imgsB, labels = [t.to(device) for t in data]
codesA, codesB = model(imgsA, imgsB)
loss, acc = criterion(codesA, codesB, labels)
return loss.item(), acc.item()
model = SiameseNetwork().to(device)
criterion = ContrastiveLoss()
optimizer = optim.Adam(model.parameters(),lr = 0.001)
n_epochs = 200
log = Report(n_epochs)
for epoch in range(n_epochs):
N = len(trn_dl)
for i, data in enumerate(trn_dl):
loss, acc = train_batch(model, data, optimizer, criterion)
log.record(epoch+(1+i)/N, trn_loss=loss, trn_acc=acc, end='\r')
N = len(val_dl)
for i, data in enumerate(val_dl):
loss, acc = validate_batch(model, data, criterion)
log.record(epoch+(1+i)/N, val_loss=loss, val_acc=acc, end='\r')
if (epoch+1)%20==0: log.report_avgs(epoch+1)
if epoch==10: optimizer = optim.Adam(model.parameters(), lr=0.0005)
log.plot_epochs(['trn_loss', 'val_loss'], log=True, title='Variation in training and validation loss')
log.plot_epochs(['trn_acc', 'val_acc'], title='Variation in training and validation accuracy')
model.eval()
val_dl = DataLoader(val_ds,num_workers=6,batch_size=1,shuffle=True)
dataiter = iter(val_dl)
x0, _, _ = next(dataiter)
for i in range(2):
_, x1, label2 = next(dataiter)
concatenated = torch.cat((x0*0.5+0.5, x1*0.5+0.5),0)
output1,output2 = model(x0.cuda(),x1.cuda())
euclidean_distance = F.pairwise_distance(output1, output2)
output = 'Same Face' if euclidean_distance.item() < 0.6 else 'Different'
show(torchvision.utils.make_grid(concatenated),
title='Dissimilarity: {:.2f}\n{}'.format(euclidean_distance.item(), output))
plt.show()
```
| github_jupyter |
```
%config InlineBackend.figure_format = 'retina'
%load_ext autoreload
%autoreload 1
%aimport ds_tutorial.datasets
%aimport ds_tutorial.transformers
import numpy as np
import pandas as pd
import pickle
from pathlib import Path
```
# Load data
```
data_root = Path.home() / "data" / "tmp"
reuters_dir = data_root / "reuters21578"
reuters_corpus_path = reuters_dir / "corpus.pkl"
reuters = pickle.load(open(reuters_corpus_path, "rb"))
top_ten_ids, top_ten_names = reuters.top_n(n=10)
cache_dir = reuters_dir / "cache"
cat_ids, cat_names = reuters.top_n(n=90)
label_lookup = {k: v for k, v in zip(cat_ids, cat_names)}
label_lookup[6]
topic_lookup = {v: k for k, v in reuters.topics.items()}
```
## Build dataframe
```
df, top_ten_ids, train_labels, test_labels = reuters.build_dataframe()
df.head()
```
# Build feature extraction pipeline
```
from sklearn.pipeline import Pipeline
from sklearn.pipeline import FeatureUnion
from sklearn.preprocessing import Imputer
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import MultiLabelBinarizer
from sklearn.decomposition import TruncatedSVD
from sklearn.metrics import classification_report
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.svm import LinearSVC
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from sklearn.multiclass import OneVsRestClassifier
from sklearn.feature_selection import SelectKBest, chi2
import xgboost as xgb
import lightgbm as lgb
from ds_tutorial.transformers import TextFromColumns, TextStats, ColumnSelector, TextFromColumns2
df_train = df.query("modapte == 'train'")
df_test = df.query("modapte == 'test'")
y_train = MultiLabelBinarizer().fit_transform(df_train.label.values)
y_test = MultiLabelBinarizer().fit_transform(df_test.label.values)
pipeline = Pipeline(
memory=str(cache_dir),
steps=[
("union", FeatureUnion(n_jobs=1, transformer_list=[
("title_stats", Pipeline([
("column", ColumnSelector("title")),
("stats", TextStats()),
#("scaled", StandardScaler()),
])),
("body_stats", Pipeline([
("column", ColumnSelector("body")),
("stats", TextStats()),
#("scaled", StandardScaler()),
])),
("combined_text", Pipeline([
("column", TextFromColumns2()),
("tfidf", TfidfVectorizer()),
])),
])),
# ("feature_selection", SelectKBest(k=20000)),
# ("clf", OneVsRestClassifier(LinearSVC(C=1.5))),
("clf", OneVsRestClassifier(LogisticRegression(C=100))),
# ("clf", OneVsRestClassifier(RandomForestClassifier())),
# ("clf", OneVsRestClassifier(xgb.XGBClassifier())),
])
pipeline.fit(df_train, y_train)
y_pred = pipeline.predict(df_test)
print(classification_report(y_test, y_pred, target_names=top_ten_names, labels=top_ten_ids, digits=3))
#pipeline.fit(df_train, y_train)
#y_pred = pipeline.predict(df_test)
#print(classification_report(y_test, y_pred, target_names=top_ten_names, labels=top_ten_ids, digits=3))
```
# Grid search
```
from sklearn.model_selection import GridSearchCV
```
## linear models
```
param_grid = {
"clf__estimator__C": [1, 100],
"clf": [
OneVsRestClassifier(LinearSVC()),
OneVsRestClassifier(LogisticRegression())
],
}
grid_search = GridSearchCV(pipeline, param_grid=param_grid, verbose=10, scoring="f1_micro")
grid_search.fit(df_train, y_train)
print(grid_search.best_estimator_)
print(grid_search.best_estimator_.steps[-1])
print(grid_search.cv_results_["split1_test_score"])
print(grid_search.cv_results_["split1_test_score"])
y_pred = grid_search.predict(df_test)
print(classification_report(y_test, y_pred, target_names=top_ten_names, labels=top_ten_ids, digits=3))
#grid_search.cv_results_
```
## xgboost
```
import xgboost as xgb
pipeline = Pipeline(
memory=str(cache_dir),
steps=[
("union", FeatureUnion(n_jobs=1, transformer_list=[
("title_stats", Pipeline([
("column", ColumnSelector("title")),
("stats", TextStats()),
("scaled", StandardScaler()),
])),
("body_stats", Pipeline([
("column", ColumnSelector("body")),
("stats", TextStats()),
("scaled", StandardScaler()),
])),
("combined_text", Pipeline([
("column", TextFromColumns2()),
("tfidf", TfidfVectorizer()),
("svd", TruncatedSVD(n_components=300, random_state=2018))
])),
])),
("clf", OneVsRestClassifier(xgb.XGBClassifier(nthread=4, early_stopping_rounds=10)))
])
%%time
pipeline.fit(df_train, y_train)
y_pred = pipeline.predict(df_test)
print(classification_report(y_test, y_pred, target_names=top_ten_names, labels=top_ten_ids, digits=3))
print(pipeline.steps[-1])
xgb_params = {
"nthread": [4],
"objective": ["multi:softprob"],
"learning_rate": [0.05, 0.001],
"max_depth": [10],
"random_state": [2018],
"gamma": [0, 0.1],
"reg_alpha": [0],
"n_estimators": [200],
}
param_grid = {
"clf": [OneVsRestClassifier(xgb.XGBClassifier(early_stopping_rounds=3))]
}
param_grid.update({f"clf__estimator__{k}": v for k, v in xgb_params.items()})
param_grid
%%time
clf = GridSearchCV(pipeline, param_grid, n_jobs=1,
scoring='f1_micro',
verbose=2, refit=True)
clf.fit(df_train, y_train)
y_pred = clf.predict(df_test)
print(classification_report(y_test, y_pred, target_names=top_ten_names, labels=top_ten_ids, digits=3))
clf.best_estimator_.steps[-1]
```
| github_jupyter |
# Basic ETL With Pandas
ETL stands for "Extract Transform Load" and it's a blanket term for a common type of software process. Specifically a process where:
1. You have data in some source format that you need to "extract"
2. The data doesn't perfectly match the format you want the data to be in, so you "transform" it
3. The data needs to be stored once again, so you "load" it into the new storage system or format.
```
# In this case our data starts in a CSV, which is pretty transportable...
import pandas as pd
path_to_ny_sales = 'nyc-property-data/nyc-rolling-sales.csv'
sales_df = pd.read_csv(path_to_ny_sales)
sales_df.head()
```
## Lets perform a few transformations:
1. Replace borough with the actual name of the borough as a string.
2. Remove some columns that we don't care about for whatever reason.
3. Drop rows from numerical columns with non-numerical or otherwise missing data.
4. Add a column that indicates "residential" "commercial" or "mixed use" based on the number of units.
```
# 1: use the map feature to replace values in columns with an alternate
sales_df['BOROUGH'] = sales_df['BOROUGH'].map({
1 : 'Manhattan',
2 : 'Bronx',
3 : 'Brooklyn',
4 : 'Queens',
5 : 'Staten Island',
})
sales_df['BOROUGH']
# 2: Drop some of the columns
columns_to_drop = [
'Unnamed: 0',
'TAX CLASS AT PRESENT',
'BLOCK',
'LOT',
'EASE-MENT',
'BUILDING CLASS AT PRESENT',
'TAX CLASS AT TIME OF SALE',
'BUILDING CLASS AT TIME OF SALE',
'BUILDING CLASS CATEGORY'
]
sales_df = sales_df.drop(columns=columns_to_drop)
sales_df.head()
# 3: Convert columns and drop na values
columns_to_convert = [
'LAND SQUARE FEET',
'GROSS SQUARE FEET',
'SALE PRICE',
'YEAR BUILT'
]
for column_name in columns_to_convert:
sales_df[column_name] = pd.to_numeric(sales_df[column_name], errors='coerce')
sales_df = sales_df[sales_df[column_name].notna()]
sales_df.describe()
# 4: create a column based on data in the other columns
# Specifically, check if this property is residential, commercial, or mixed use
def check_building_type(row):
if row['COMMERCIAL UNITS'] > 0 and row['RESIDENTIAL UNITS'] > 0:
return "MIXED USE"
elif row['COMMERCIAL UNITS'] > 0:
return "COMMERCIAL"
elif row['RESIDENTIAL UNITS']:
return "RESIDENTIAL"
else:
return "UNKNOWN - NO UNITS"
# axis=1 means apply the function to rows
# axis=0 would mean apply the function to the columns
sales_df['BUILDING TYPE'] = sales_df.apply(check_building_type, axis=1)
sales_df.head()
```
# Load
There are alternate mechanisms in Python for loading data into any number of databases and other storage mechanisms. For now, lets just load to a file since we don't assume knowledge of SQL or anything else for this class.
```
# Pandas makes it very easy to write to a CSV just as it does for reading.
# See more options in the docs: https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.to_csv.html
sales_df.to_csv('nyc-property-data/transformed_nyc_housing.csv', index=False)
```
# An Important Note:
ETL jobs ideally will be super easy to run, and potentially even automated in most situations. As a result, Jupyter Notebook is not the most ideal format for this kind of work. In a real world project I would:
1. Write the transformations and confirm they work in a Jupyter notebook during the prototyping phase.
2. Move the transformations to a single script.
3. (probably) Write some tests to ensure the ETL job performs as expected.
The script for step 2 of this transformation is in the code repository with the name:
| github_jupyter |
```
import numpy as np
import pandas as pd
from sklearn.linear_model import LinearRegression, Ridge, Lasso, LogisticRegression
from sklearn.ensemble import RandomForestClassifier, RandomForestRegressor
from sklearn.datasets import load_boston, load_iris, load_wine, load_digits, \
load_breast_cancer, load_diabetes, fetch_mldata
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix, precision_score, recall_score
import matplotlib.pyplot as plt
#%config InlineBackend.figure_format = 'svg'
from sklearn import tree
from dtreeviz.trees import *
```
## Regression
```
df_cars = pd.read_csv("../data/cars.csv")
X = df_cars.drop('MPG', axis=1)
y = df_cars['MPG']
fig, ax = plt.subplots(1,1, figsize=(4,2.5))
t = rtreeviz_univar(ax,
X['WGT'], y,
max_depth=3,
feature_name='Vehicle Weight',
target_name='MPG',
fontsize=10,
colors={'scatter_edge': 'black'})
plt.tight_layout()
plt.show()
rtreeviz_bivar_3D(None,
X[['WGT','ENG']], y,
max_depth=3,
feature_names=['Vehicle Weight', 'Horse Power'],
target_name='MPG',
fontsize=10,
elev=30,
azim=20,
dist=10,
show={'splits','title'},
colors={'tesselation_alpha':.5})
rtreeviz_bivar_heatmap(None,
X[['WGT','ENG']], y,
feature_names=['Vehicle Weight', 'Horse Power'],
max_depth=4,
fontsize=10)
```
## Classification
```
iris = load_iris()
X = iris.data
y = iris.target
len(X), len(y)
feature_names = iris.feature_names
class_names = list(iris.target_names)
print(feature_names)
figsize = (6,2)
X = X[:,2]
ct = ctreeviz_univar(None, X, y, max_depth=1,
feature_name = 'petal length (cm)', class_names=class_names,
target_name='iris',
nbins=40, gtype='barstacked',
show={'splits','title'})
plt.tight_layout()
plt.show()
from dtreeviz.trees import *
wine = load_wine()
X = wine.data
y = wine.target
len(X), len(y)
colors = {'classes':
[None, # 0 classes
None, # 1 class
["#FEFEBB","#a1dab4"], # 2 classes
["#FEFEBB","#D9E6F5",'#a1dab4'], # 3
]
}
feature_names = wine.feature_names
class_names = list(wine.target_names)
X = X[:,[12,6]]
ct = ctreeviz_bivar(None, X, y, max_depth=2,
feature_names = ['proline','flavanoid'], class_names=class_names,
target_name='iris',
show={'splits'},
colors={'scatter_edge': 'black'}
)
plt.tight_layout()
plt.show()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/master/tutorials/W3D3_NetworkCausality/student/W3D3_Tutorial1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Neuromatch Academy 2020: Week 3 Day 3, Tutorial 1
# Causality Day: Interventions
**Content creators**: Ari Benjamin, Tony Liu, Konrad Kording
**Content reviewers**: Mike X Cohen, Madineh Sarvestani, Ella Batty, Michael Waskom
---
#Tutorial Objectives
We list our overall day objectives below, with the sections we will focus on in this notebook in bold:
1. **Master definitions of causality**
2. **Understand that estimating causality is possible**
3. **Learn 4 different methods and understand when they fail**
1. **Perturbations**
2. Correlations
3. Simultaneous fitting/regression
4. Instrumental variables
### Tutorial setting
How do we know if a relationship is causal? What does that mean? And how can we estimate causal relationships within neural data?
The methods we'll learn today are very general and can be applied to all sorts of data, and in many circumstances.
Causal questions are everywhere!
### Tutorial 1 Objectives:
1. Simulate a neural system
2. Understand perturbation as a method of estimating causality
---
# Setup
```
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.axes_grid1 import make_axes_locatable
#@title Figure settings
%config InlineBackend.figure_format = 'retina'
plt.style.use("https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/nma.mplstyle")
#@title Helper functions
def sigmoid(x):
"""
Compute sigmoid nonlinearity element-wise on x
Args:
x (np.ndarray): the numpy data array we want to transform
Returns
(np.ndarray): x with sigmoid nonlinearity applied
"""
return 1 / (1 + np.exp(-x))
def create_connectivity(n_neurons, random_state=42):
"""
Generate our nxn causal connectivity matrix.
Args:
n_neurons (int): the number of neurons in our system.
random_state (int): random seed for reproducibility
Returns:
A (np.ndarray): our 0.1 sparse connectivity matrix
"""
np.random.seed(random_state)
A_0 = np.random.choice([0, 1], size=(n_neurons, n_neurons), p=[0.9, 0.1])
# set the timescale of the dynamical system to about 100 steps
_, s_vals, _ = np.linalg.svd(A_0)
A = A_0 / (1.01 * s_vals[0])
# _, s_val_test, _ = np.linalg.svd(A)
# assert s_val_test[0] < 1, "largest singular value >= 1"
return A
def see_neurons(A, ax):
"""
Visualizes the connectivity matrix.
Args:
A (np.ndarray): the connectivity matrix of shape (n_neurons, n_neurons)
ax (plt.axis): the matplotlib axis to display on
Returns:
Nothing, but visualizes A.
"""
A = A.T # make up for opposite connectivity
n = len(A)
ax.set_aspect('equal')
thetas = np.linspace(0, np.pi * 2, n, endpoint=False)
x, y = np.cos(thetas), np.sin(thetas),
ax.scatter(x, y, c='k', s=150)
# Renormalize
A = A / A.max()
for i in range(n):
for j in range(n):
if A[i, j] > 0:
ax.arrow(x[i], y[i], x[j] - x[i], y[j] - y[i], color='k', alpha=A[i,j],
head_width=.15, width = A[i, j] / 25, shape='right',
length_includes_head=True)
ax.axis('off')
def get_perturbed_connectivity_all_neurons(perturbed_X):
"""
Estimates the connectivity matrix of perturbations through stacked correlations.
Args:
perturbed_X (np.ndarray): the simulated dynamical system X of shape
(n_neurons, timesteps)
Returns:
R (np.ndarray): the estimated connectivity matrix of shape
(n_neurons, n_neurons)
"""
# select perturbations (P) and outcomes (Outs)
# we perturb the system every over time step, hence the 2 in slice notation
P = perturbed_X[:, ::2]
Outs = perturbed_X[:, 1::2]
# stack perturbations and outcomes into a 2n by (timesteps / 2) matrix
S = np.concatenate([P, Outs], axis=0)
# select the perturbation -> outcome block of correlation matrix (upper right)
R = np.corrcoef(S)[:n_neurons, n_neurons:]
return R
def simulate_neurons_perturb(A, timesteps):
"""
Simulates a dynamical system for the specified number of neurons and timesteps,
BUT every other timestep the activity is clamped to a random pattern of 1s and 0s
Args:
A (np.array): the true connectivity matrix
timesteps (int): the number of timesteps to simulate our system.
Returns:
The results of the simulated system.
- X has shape (n_neurons, timeteps)
"""
n_neurons = len(A)
X = np.zeros((n_neurons, timesteps))
for t in range(timesteps - 1):
if t % 2 == 0:
X[:, t] = np.random.choice([0, 1], size=n_neurons)
epsilon = np.random.multivariate_normal(np.zeros(n_neurons), np.eye(n_neurons))
X[:, t + 1] = sigmoid(A.dot(X[:, t]) + epsilon) # we are using helper function sigmoid
return X
def plot_connectivity_matrix(A, ax=None):
"""Plot the (weighted) connectivity matrix A as a heatmap
Args:
A (ndarray): connectivity matrix (n_neurons by n_neurons)
ax: axis on which to display connectivity matrix
"""
if ax is None:
ax = plt.gca()
lim = np.abs(A).max()
im = ax.imshow(A, vmin=-lim, vmax=lim, cmap="coolwarm")
ax.tick_params(labelsize=10)
ax.xaxis.label.set_size(15)
ax.yaxis.label.set_size(15)
cbar = ax.figure.colorbar(im, ax=ax, ticks=[0], shrink=.7)
cbar.ax.set_ylabel("Connectivity Strength", rotation=90,
labelpad= 20,va="bottom")
ax.set(xlabel="Connectivity from", ylabel="Connectivity to")
def plot_connectivity_graph_matrix(A):
"""Plot both connectivity graph and matrix side by side
Args:
A (ndarray): connectivity matrix (n_neurons by n_neurons)
"""
fig, axs = plt.subplots(1, 2, figsize=(10, 5))
see_neurons(A,axs[0]) # we are invoking a helper function that visualizes the connectivity matrix
plot_connectivity_matrix(A)
fig.suptitle("Neuron Connectivity")
plt.show()
def plot_neural_activity(X):
"""Plot first 10 timesteps of neural activity
Args:
X (ndarray): neural activity (n_neurons by timesteps)
"""
f, ax = plt.subplots()
im = ax.imshow(X[:, :10])
divider = make_axes_locatable(ax)
cax1 = divider.append_axes("right", size="5%", pad=0.15)
plt.colorbar(im, cax=cax1)
ax.set(xlabel='Timestep', ylabel='Neuron', title='Simulated Neural Activity')
def plot_true_vs_estimated_connectivity(estimated_connectivity, true_connectivity, selected_neuron=None):
"""Visualize true vs estimated connectivity matrices
Args:
estimated_connectivity (ndarray): estimated connectivity (n_neurons by n_neurons)
true_connectivity (ndarray): ground-truth connectivity (n_neurons by n_neurons)
selected_neuron (int or None): None if plotting all connectivity, otherwise connectivity
from selected_neuron will be shown
"""
fig, axs = plt.subplots(1, 2, figsize=(10, 5))
if selected_neuron is not None:
plot_connectivity_matrix(np.expand_dims(estimated_connectivity, axis=1), ax=axs[0])
plot_connectivity_matrix(true_connectivity[:, [selected_neuron]], ax=axs[1])
axs[0].set_xticks([0])
axs[1].set_xticks([0])
axs[0].set_xticklabels([selected_neuron])
axs[1].set_xticklabels([selected_neuron])
else:
plot_connectivity_matrix(estimated_connectivity, ax=axs[0])
plot_connectivity_matrix(true_connectivity, ax=axs[1])
axs[1].set(title="True connectivity")
axs[0].set(title="Estimated connectivity")
```
---
# Section 1: Defining and estimating causality
```
#@title Video 1: Defining causality
# Insert the ID of the corresponding youtube video
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="yiddT2sMbZM", width=854, height=480, fs=1)
print("Video available at https://youtu.be/" + video.id)
video
```
Let's think carefully about the statement "**A causes B**". To be concrete, let's take two neurons. What does it mean to say that neuron $A$ causes neuron $B$ to fire?
The *interventional* definition of causality says that:
$$
(A \text{ causes } B) \Leftrightarrow ( \text{ If we force }A \text { to be different, then }B\text{ changes})
$$
To determine if $A$ causes $B$ to fire, we can inject current into neuron $A$ and see what happens to $B$.
**A mathematical definition of causality**:
Over many trials, the average causal effect $\delta_{A\to B}$ of neuron $A$ upon neuron $B$ is the average change in neuron $B$'s activity when we set $A=1$ versus when we set $A=0$.
$$
\delta_{A\to B} = \mathbb{E}[B | A=1] - \mathbb{E}[B | A=0]
$$
Note that this is an average effect. While one can get more sophisticated about conditional effects ($A$ only effects $B$ when it's not refractory, perhaps), we will only consider average effects today.
**Relation to a randomized controlled trial (RCT)**:
The logic we just described is the logic of a randomized control trial (RCT). If you randomly give 100 people a drug and 100 people a placebo, the effect is the difference in outcomes.
## Exercise 1: Randomized controlled trial for two neurons
Let's pretend we can perform a randomized controlled trial for two neurons. Our model will have neuron $A$ synapsing on Neuron $B$:
$$B = A + \epsilon$$
where $A$ and $B$ represent the activities of the two neurons and $\epsilon$ is standard normal noise $\epsilon\sim\mathcal{N}(0,1)$.
Our goal is to perturb $A$ and confirm that $B$ changes.
```
def neuron_B(activity_of_A):
"""Model activity of neuron B as neuron A activity + noise
Args:
activity_of_A (ndarray): An array of shape (T,) containing the neural activity of neuron A
Returns:
ndarray: activity of neuron B
"""
noise = np.random.randn(activity_of_A.shape[0])
return activity_of_A + noise
np.random.seed(12)
# Neuron A activity of zeros
A_0 = np.zeros(5000)
# Neuron A activity of ones
A_1 = np.ones(5000)
###########################################################################
## TODO for students: Estimate the causal effect of A upon B
## Use eq above (difference in mean of B when A=0 vs. A=1)
###########################################################################
diff_in_means = ...
#print(diff_in_means)
```
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D3_NetworkCausality/solutions/W3D3_Tutorial1_Solution_9ae3afbe.py)
You should get a difference in means of `0.990719` (so very close to one).
---
# Section 2: Simulating a system of neurons
Can we still estimate causal effects when the neurons are in big networks? This is the main question we will ask today. Let's first create our system, and the rest of today will be spend analyzing it.
```
#@title Video 2: Simulated neural system model
# Insert the ID of the corresponding youtube video
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="oPJz49dAuL8", width=854, height=480, fs=1)
print("Video available at https://youtu.be/" + video.id)
video
```
**Video correction**: the connectivity graph plots and associated explanations in this and other videos show the wrong direction of connectivity (the arrows should be pointing the opposite direction). This has been fixed in the figures below.
## Section 2.1: Our system
This section recaps the system described in Video 2 so may be skipped.
Our system has N interconnected neurons that affect each other over time. Each neuron at time $t+1$ is a function of the activity of the other neurons from the previous time $t$.
Neurons affect each other nonlinearly: each neuron's activity at time $t+1$ consists of a linearly weighted sum of all neural activities at time $t$, with added noise, passed through a nonlinearity:
$$
\vec{x}_{t+1} = \sigma(A\vec{x}_t + \epsilon_t),
$$
- $\vec{x}_t$ is an $n$-dimensional vector representing our $n$-neuron system at timestep $t$
- $\sigma$ is a sigmoid nonlinearity
- $A$ is our $n \times n$ *causal ground truth connectivity matrix* (more on this later)
- $\epsilon_t$ is random noise: $\epsilon_t \sim N(\vec{0}, I_n)$
- $\vec{x}_0$ is initialized to $\vec{0}$
$A$ is a connectivity matrix, so the element $A_{ij}$ represents the causal effect of neuron $i$ on neuron $j$. In our system, neurons will receive connections from only 10% of the whole population on average.
## Section 2.2: Visualize true connectivity
We will create a connectivity matrix between 6 neurons and visualize it in two different ways: as a graph with directional edges between connected neurons and as an image of the connectivity matrix.
*Check your understanding*: do you understand how the left plot relates to the right plot below?
```
#@markdown Execute this cell to visualize connectivity
## Initializes the system
n_neurons = 6
A = create_connectivity(n_neurons) # we are invoking a helper function that generates our nxn causal connectivity matrix.
# Let's plot it!
plot_connectivity_graph_matrix(A)
```
## Exercise 2: System simulation
In this exercise we're going to simulate the system. Please complete the following function so that at every timestep the activity vector $x$ is updated according to:
$$
\vec{x}_{t+1} = \sigma(A\vec{x}_t + \epsilon_t).
$$
```
def simulate_neurons(A, timesteps, random_state=42):
"""Simulates a dynamical system for the specified number of neurons and timesteps.
Args:
A (np.array): the connectivity matrix
timesteps (int): the number of timesteps to simulate our system.
random_state (int): random seed for reproducibility
Returns:
- X has shape (n_neurons, timeteps). A schematic:
___t____t+1___
neuron 0 | 0 1 |
| 1 0 |
neuron i | 0 -> 1 |
| 0 0 |
|___1____0_____|
"""
np.random.seed(random_state)
n_neurons = len(A)
X = np.zeros((n_neurons, timesteps))
for t in range(timesteps - 1):
# Create noise vector
epsilon = np.random.multivariate_normal(np.zeros(n_neurons), np.eye(n_neurons))
########################################################################
## TODO: Fill in the update rule for our dynamical system.
## Fill in function and remove
raise NotImplementedError("Complete simulate_neurons")
########################################################################
# Update activity vector for next step
X[:, t + 1] = sigmoid(...) # we are using helper function sigmoid
return X
# Set simulation length
timesteps = 5000
# Uncomment below to test your function
# Simulate our dynamical system
# X = simulate_neurons(A, timesteps)
# plot_neural_activity(X)
```
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D3_NetworkCausality/solutions/W3D3_Tutorial1_Solution_b2fb6587.py)
*Example output:*
<img alt='Solution hint' align='left' width=557 height=343 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W3D3_NetworkCausality/static/W3D3_Tutorial1_Solution_b2fb6587_0.png>
---
# Section 3: Recovering connectivity through perturbation
```
#@title Video 3: Perturbing systems
# Insert the ID of the corresponding youtube video
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="wOZunGtuqQE", width=854, height=480, fs=1)
print("Video available at https://youtu.be/" + video.id)
video
```
## Section 3.1: Random perturbation in our system of neurons
We want to get the causal effect of each neuron upon each other neuron. The ground truth of the causal effects is the connectivity matrix $A$.
Remember that we would like to calculate:
$$
\delta_{A\to B} = \mathbb{E}[B | A=1] - \mathbb{E}[B | A=0]
$$
We'll do this by randomly setting the system state to 0 or 1 and observing the outcome after one timestep. If we do this $N$ times, the effect of neuron $i$ upon neuron $j$ is:
$$
\delta_{x^i\to x^j} \approx \frac1N \sum_i^N[x_{t+1}^j | x^i_t=1] - \frac1N \sum_i^N[x_{t+1}^j | x^i_t=0]
$$
This is just the average difference of the activity of neuron $j$ in the two conditions.
We are going to calculate the above equation, but imagine it like *intervening* in activity every other timestep.
We will use helper function `simulate_neurons_perturb`. While the rest of the function is the same as the ``simulate_neurons`` function in the previous exercise, every time step we now additionally include:
```
if t % 2 == 0:
X[:,t] = np.random.choice([0,1], size=n_neurons)
```
This means that at every other timestep, every neuron's activity is changed to either 0 or 1.
Pretty serious perturbation, huh? You don't want that going on in your brain.
**Now visually compare the dynamics:** Run this next cell and see if you can spot how the dynamics have changed.
```
# @markdown Execute this cell to visualize perturbed dynamics
timesteps = 5000 # Simulate for 5000 timesteps.
# Simulate our dynamical system for the given amount of time
X_perturbed = simulate_neurons_perturb(A, timesteps)
# Plot our standard versus perturbed dynamics
fig, axs = plt.subplots(1, 2, figsize=(15, 4))
im0 = axs[0].imshow(X[:, :10])
im1 = axs[1].imshow(X_perturbed[:, :10])
# Matplotlib boilerplate code
divider = make_axes_locatable(axs[0])
cax0 = divider.append_axes("right", size="5%", pad=0.15)
plt.colorbar(im0, cax=cax0)
divider = make_axes_locatable(axs[1])
cax1 = divider.append_axes("right", size="5%", pad=0.15)
plt.colorbar(im1, cax=cax1)
axs[0].set_ylabel("Neuron", fontsize=15)
axs[1].set_xlabel("Timestep", fontsize=15)
axs[0].set_xlabel("Timestep", fontsize=15);
axs[0].set_title("Standard dynamics", fontsize=15)
axs[1].set_title("Perturbed dynamics", fontsize=15);
#@title Video 4: Calculating causality
# Insert the ID of the corresponding youtube video
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="EDZtcsIAVGM", width=854, height=480, fs=1)
print("Video available at https://youtu.be/" + video.id)
video
```
## Exercise 3: Using perturbed dynamics to recover connectivity
From the above perturbed dynamics, write a function that recovers the causal effect of a given single neuron (`selected_neuron`) upon all other neurons in the system. Remember from above you're calculating:
$$
\delta_{x^i\to x^j} \approx \frac1N \sum_i^N[x_{t+1}^j | x^i_t=1] - \frac1N \sum_i^N[x_{t+1}^j | x^i_t=0]]
$$
Recall that we perturbed every neuron at every other timestep. Despite perturbing every neuron, in this exercise we are concentrating on computing the causal effect of a single neuron (we will look at all neurons effects on all neurons next). We want to exclusively use the timesteps without perturbation for $x^j_{t+1}$ and the timesteps with perturbation for $x^j_{t}$ in the formulas above. In numpy, indexing occurs as `array[ start_index : end_index : count_by]`. So getting every other element in an array (such as every other timestep) is as easy as `array[::2]`.
```
def get_perturbed_connectivity_from_single_neuron(perturbed_X, selected_neuron):
"""
Computes the connectivity matrix from the selected neuron using differences in means.
Args:
perturbed_X (np.ndarray): the perturbed dynamical system matrix of shape (n_neurons, timesteps)
selected_neuron (int): the index of the neuron we want to estimate connectivity for
Returns:
estimated_connectivity (np.ndarray): estimated connectivity for the selected neuron, of shape (n_neurons,)
"""
# Extract the perturbations of neuron 1 (every other timestep)
neuron_perturbations = perturbed_X[selected_neuron, ::2]
# Extract the observed outcomes of all the neurons (every other timestep)
all_neuron_output = perturbed_X[:, 1::2]
# Initialize estimated connectivity matrix
estimated_connectivity = np.zeros(n_neurons)
# Loop over neurons
for neuron_idx in range(n_neurons):
# Get this output neurons (neuron_idx) activity
this_neuron_output = all_neuron_output[neuron_idx, :]
# Get timesteps where the selected neuron == 0 vs == 1
one_idx = np.argwhere(neuron_perturbations == 1)
zero_idx = np.argwhere(neuron_perturbations == 0)
########################################################################
## TODO: Insert your code here to compute the neuron activation from perturbations.
# Fill out function and remove
raise NotImplementedError("Complete the function get_perturbed_connectivity_single_neuron")
########################################################################
difference_in_means = ...
estimated_connectivity[neuron_idx] = difference_in_means
return estimated_connectivity
# Initialize the system
n_neurons = 6
timesteps = 5000
selected_neuron = 1
# Simulate our perturbed dynamical system
perturbed_X = simulate_neurons_perturb(A, timesteps)
## Uncomment below to test your function
# Measure connectivity of neuron 1
# estimated_connectivity = get_perturbed_connectivity_from_single_neuron(perturbed_X, selected_neuron)
# plot_true_vs_estimated_connectivity(estimated_connectivity, A, selected_neuron)
```
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W3D3_NetworkCausality/solutions/W3D3_Tutorial1_Solution_b51df5f6.py)
*Example output:*
<img alt='Solution hint' align='left' width=486 height=341 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W3D3_NetworkCausality/static/W3D3_Tutorial1_Solution_b51df5f6_0.png>
We can quantify how close our estimated connectivity matrix is to our true connectivity matrix by correlating them. We should see almost perfect correlation between our estimates and the true connectivity - do we?
```
# Correlate true vs estimated connectivity matrix
np.corrcoef(A[:, selected_neuron], estimated_connectivity)[1, 0]
```
**Note on interpreting A**: Strictly speaking, $A$ is not the matrix of causal effects but rather the dynamics matrix. So why compare them like this? The answer is that $A$ and the effect matrix both are $0$ everywhere except where there is a directed connection. So they should have a correlation of $1$ if we estimate the effects correctly. (Their scales, however, are different. This in part because the nonlinearity $\sigma$ squashes the values of $x$ to $[0,1]$.) See the Appendix after Tutorial 2 for more discussion of using correlation as a metric.
## Section 3.2: Measuring how perturbations recover the entire connectivity matrix
Nice job! You just estimated connectivity for a single neuron.
We're now going to use the same strategy for all neurons at once. We provide this helper function `get_perturbed_connectivity_all_neurons`. If you're curious about how this works and have extra time, scroll to the explanation at the bottom.
```
# Parameters
n_neurons = 6
timesteps = 5000
# Generate nxn causal connectivity matrix
A = create_connectivity(n_neurons)
# Simulate perturbed dynamical system
perturbed_X = simulate_neurons_perturb(A, timesteps)
# Get estimated connectivity matrix
R = get_perturbed_connectivity_all_neurons(perturbed_X)
#@markdown Execute this cell to visualize true vs estimated connectivity
# Let's visualize the true connectivity and estimated connectivity together
fig, axs = plt.subplots(1, 2, figsize=(10, 5))
see_neurons(A, axs[0]) # we are invoking a helper function that visualizes the connectivity matrix
plot_connectivity_matrix(A, ax=axs[1])
plt.suptitle("True connectivity matrix A");
plt.show()
fig, axs = plt.subplots(1,2, figsize=(10,5))
see_neurons(R.T,axs[0]) # we are invoking a helper function that visualizes the connectivity matrix
plot_connectivity_matrix(R.T, ax=axs[1])
plt.suptitle("Estimated connectivity matrix R");
```
We can again calculate the correlation coefficient between the elements of the two matrices. As you can see from the cell below, we do a good job recovering the true causality of the system!
```
np.corrcoef(A.transpose().flatten(), R.flatten())[1, 0]
```
---
# Summary
```
#@title Video 5: Summary
# Insert the ID of the corresponding youtube video
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="p3fZW5Woqa4", width=854, height=480, fs=1)
print("Video available at https://youtu.be/" + video.id)
video
```
In this tutorial, we learned about how to define and estimate causality using pertubations. In particular we:
1) Learned how to simulate a system of connected neurons
2) Learned how to estimate the connectivity between neurons by directly perturbing neural activity
---
# Further resources for today
If you are interested in causality after NMA ends, here are some useful texts to consult.
* *Causal Inference for Statistics, Social, and Biomedical Sciences* by Imbens and Rubin
* *Causal Inference: What If* by Hernan and Rubin
* *Mostly Harmless Econometrics* by Angrist and Pischke
* https://www.nature.com/articles/s41562-018-0466-5 for application to neuroscience
---
# Appendix
## Computation of the estimated connectivity matrix
**This is an explanation of what the code is doing in `get_perturbed_connectivity_all_neurons()`**
First, we compute an estimated connectivity matrix $R$. We extract
perturbation matrix $P$ and outcomes matrix $O$:
$$
P = \begin{bmatrix}
\mid & \mid & ... & \mid \\
x_0 & x_2 & ... & x_T \\
\mid & \mid & ... & \mid
\end{bmatrix}_{n \times T/2}
$$
$$
O = \begin{bmatrix}
\mid & \mid & ... & \mid \\
x_1 & x_3 & ... & x_{T-1} \\
\mid & \mid & ... & \mid
\end{bmatrix}_{n \times T/2}
$$
And calculate the correlation of matrix $S$, which is $P$ and $O$ stacked on each other:
$$
S = \begin{bmatrix}
P \\
O
\end{bmatrix}_{2n \times T/2}
$$
We then extract $R$ as the upper right $n \times n$ block of $corr(S)$:
This is because the upper right block corresponds to the estimated perturbation effect on outcomes for each pair of neurons in our system.
This method gives an estimated connectivity matrix that is the proportional to the result you would obtain with differences in means, and differs only in a proportionality constant that depends on the variance of $x$
| github_jupyter |
```
import numpy as np
from numpy.random import Generator, PCG64, SeedSequence
import pandas as pd
import matplotlib.pyplot as plt
import scipy.stats as st
from math import pi
import seaborn as sns
from scipy.special import gamma, logsumexp
from tqdm import tqdm
from joblib import Parallel, delayed
import time
from numba import jit
```
# Bayesian fit and normalizing constant calculation of the exponential distribution
## Conjugate prior
The exponential distribution $\text{Exp}(\delta)$, the likelihood function associated to an iid sample $x=(x_1,\ldots, x_n)$ is
$$
L(x|\delta) = \delta^n\cdot e^{-\delta \sum_{i = 1}^n x_i},\text{ }x>0.
$$
Suppose that the prior distribution over $\delta$ is gamma $\text{Gamma}(a,1/b)$ with pdf
$$
\pi(\delta) = \frac{b^a\delta^{a-1}e^{-b\cdot \delta}}{\Gamma(a)}, \delta >0.
$$
The posterior distribution is then given by
$$
\pi(\delta|x)=\frac{L(x|\delta)\pi(\delta)}{Z(x)} \sim \text{Gamma}\left(n+a, \frac{1}{b+ \sum_{i = 1}^n x_i} \right).
$$
The normalizing constant is equal to
$$
Z(x) = \frac{b^a\Gamma(a+n)}{\Gamma(a)\left(b+\sum_{i = 1}^n x_i\right)^{a+n}}
$$
The following function sample from the posterior distribution.
```
def sample_pos_δ(X, a, b, size):
return(np.random.gamma(shape = len(X) + a, scale=1 / (b + sum(X)), size=size))
```
We illustrate the concentration of the posterior distribution arround the true value as the sample sizes increase from $10$ to $1000$.
```
δ_true = 3
fig, axs = plt.subplots(1, 1, figsize=(2.5, 3.5))
for n in [5, 10, 50]:
X, a, b, size = np.random.gamma(shape = 1, scale= 1 / δ_true, size=n), 0.1, 0.1, 50000
δ_pos = sample_pos_δ(X, a, b, size)
positions = np.linspace(min(δ_pos), max(δ_pos), 1000)
kernel = st.gaussian_kde(δ_pos)
plt.plot(positions, kernel(positions), lw=2, label = "n = "+str(n))
plt.axvline(x = δ_true, c = "black", lw=1, linestyle= "dotted", label = "True value")
plt.legend(fontsize = 6)
plt.yticks([])
plt.xlabel("δ")
sns.despine()
plt.savefig("../Figures/hist_post_sample_exponential_en.pdf")
```
## Metropolis Hasting random walk
We now apply the MH sampling scheme to sample from the posterior distribution. It requires the log ligelihood and also the log probability of the prior distribution
```
# Log likelihood for the MH RW
def logp_exp(X):
def logp(parms):
λ = parms
if λ>0:
return(len(X) * np.log(λ) - sum(X) * λ)
else:
return(-np.inf)
return logp
# Log likelihood for the temperature search
def logp_exp_vect(X):
def logp(parms):
λ = parms
return(len(X) * np.log(λ) - sum(X) * λ)
return logp
def logp_exp_vect(X):
def logp(parms):
λ = parms
return(len(X) * np.log(λ) - sum(X) * λ)
return logp
# Log probability when the parameter is gamma distributed a priori
def gamma_prior(a, b):
def logp_prior(parms):
λ = parms
if λ>0:
return((a - 1) * np.log(λ) - λ * b + a * np.log(b) - np.log(gamma(a)))
else:
return(-np.inf)
return logp_prior
# Function to generate the Markov chain trajectory
def MH_move_exp(n_moves, step_size, log_prob, log_prob_prior, init_parms, γ):
accepted = []
λ = np.array([init_parms])
epsilon = np.random.normal(size=n_moves)
for noise in epsilon:
λ_new = λ[-1] + step_size * noise
old_log_p = γ * log_prob((λ[-1])) + log_prob_prior((λ[-1]))
new_log_p = γ * log_prob((λ_new)) + log_prob_prior((λ_new))
acc = new_log_p - old_log_p
if np.log(np.random.rand()) < acc:
λ = np.append(λ, λ_new)
accepted.append(True)
else:
λ = np.append(λ, np.copy(λ[-1]))
accepted.append(False)
return(λ[1:], accepted)
```
We sample value from the posterior distribution using Metropolis Hasting for different value of the scaling parameter $h$ parameters and return the trace plots.
```
np.random.seed(123)
X, a, b = np.random.gamma(shape = 1, scale= 1 / δ_true, size = 25), 0.001, 0.001
δ_pos = sample_pos_δ(X, a, b, 100000)
positions = np.linspace(min(δ_pos), max(δ_pos), 1000)
kernel = st.gaussian_kde(δ_pos)
num_bins = 50
step_sizes = [0.01, 2.8, 30]
k = 0
for step_size in step_sizes:
n_moves, log_prob, log_prob_prior, init_parms, γ = 10000, logp_exp(X), gamma_prior(a, b), np.array([1]), 1
trace, acc = MH_move_exp(n_moves, step_size, log_prob, log_prob_prior, init_parms, γ)
fig, axs = plt.subplots(1, 2, figsize=(5, 2.5))
axs[0].plot(trace, lw=1)
axs[0].axhline(δ_true, color = "black", linestyle = "dotted")
axs[0].set_xticks([])
axs[1].plot(positions, kernel(positions), lw=3, label = "posterior")
n, bins, patches = axs[1].hist(trace[5000:], num_bins, density=1)
axs[1].axvline(δ_true, color = "black", linestyle = "dotted", label = "true value")
axs[1].set_yticks([])
axs[1].legend(fontsize = 7)
sns.despine()
plt.savefig("../Figures/trace_hist_exponential_"+str(k)+"_en.pdf")
k+=1
```
## Sequential Monte Carlo
The SMC algorithm returns posterior samples and an estimation of the normalizing constant given by (in the exponential model considered)
```
def norm_constant(X, a, b):
return(
a*np.log(b)-(a + len(X)) * np.log(sum(X) + b) + sum(np.log([a+k for k in range(len(X))]))
)
```
The SMC algorithm adapt sequentially the intermediary distribution by either increasing the temperature
```
def temperature_search(λ, W_prev, target, γ_prev, log_prob):
popSize = len(λ)
γ_up, γ_down = 2, γ_prev
while γ_up - γ_down > 1e-6:
γ_new = (γ_up + γ_down) / 2
w = np.exp(log_prob(λ)) ** (γ_new - γ_prev)
if sum(w) == 0 or np.any(np.isnan(w)):
W = 1 / popSize *np.ones(popSize)
ESS = 0
else:
W = w / sum(w)
ESS = 1 / sum(W**2)
# print(γ_new, ESS)
if ESS == target:
break
else:
if ESS < target:
γ_up = γ_new
else:
γ_down = γ_new
if γ_new >= 1:
γ_new = 1
w = np.exp(log_prob(λ)) ** (γ_new - γ_prev)
W = w / sum(w)
ESS = 1 / sum(W**2)
return(γ_new, w, W, int(ESS))
def smc_exp(popSize, a, b,log_prob_vect, log_prob, log_prob_prior, c, ρ,
n_step_max, paralell, n_proc):
# popSize = number of particles
# a,b = hyperparameter of the prior distribution on λ
# c = probability that a particle is moved at least once
# ρ = proportion of the sample of the ESS
# temp_step = size of the step in the search of the right temperature
# The parameter is gamma distributed
λ_prior = st.gamma(a)
# particles initialization
print('sample generation 0')
clouds = [pd.DataFrame({'λ': λ_prior.rvs(popSize) / b,
'w':np.ones(popSize)/ popSize,
'W': np.ones(popSize) / popSize
})]
# Temperature sequence
γ_seq = np.array([0])
# Generation counter
g=0
# We keep on iterating until the temperature reaches 1
while γ_seq[-1] < 1:
# g, ESS, k = g + 1, 1, - err
g = g + 1
print('Sampling particles from generation ' + str(g))
cloud = pd.DataFrame({ 'λ':[], 'w':[], 'W':[]})
# Updating temperature sequence
γ, w, W, ESS = temperature_search(clouds[g-1]['λ'].values,
clouds[g-1]['W'].values,
ρ * popSize,
γ_seq[-1], log_prob_vect)
γ_seq = np.append(γ_seq, γ)
# Updating unormalized weights
cloud['w'] = w
# Updating normalized weights
cloud['W'] = W
step_size = np.sqrt(np.cov(clouds[g-1]['λ'].values
, aweights=W))*2.38
particles = clouds[g-1]['λ'].values[np.random.choice(popSize,
popSize, p = W)]
def move_particle_trial(particle):
λ, accepted = MH_move_exp(1, step_size,
log_prob, log_prob_prior, particle, γ)
return([λ[-1], np.mean(accepted)])
if paralell:
res = np.matrix(Parallel(n_jobs=n_proc)(delayed(move_particle_trial)(i)
for i in particles))
else:
res = np.matrix([move_particle_trial(particle)
for particle in particles])
λ_trial = np.asarray(res[:,0]).flatten()
acc_rate = np.asarray(res[:,1]).flatten()
n_steps = min(n_step_max,
max(2,np.ceil(np.log(1-c) / np.log(1-np.mean(acc_rate)))))
print('Generation: ' + str(g) + " ;temperature: "+str(γ_seq[-1]) +
" ;steps:" + str(n_steps) + " ;stepsize: "+str(step_size)+
" ;ESS: "+str(ESS))
def move_particle(particle):
λ, accepted = MH_move_exp(int(n_steps), step_size,
log_prob, log_prob_prior, particle, γ)
return([λ[-1], np.mean(accepted)])
if paralell:
res = np.matrix(Parallel(n_jobs=n_proc)(delayed(move_particle)(i)
for i in λ_trial))
else:
res = np.matrix([move_particle(particle) for particle in λ_trial])
cloud['λ'] = np.asarray(res[:,0]).flatten()
clouds.append(cloud)
marginal_log_likelihood = sum(np.log(([cloud['w'].mean()
for cloud in clouds[1:g+1]])))
return((clouds[-1].λ.values, marginal_log_likelihood))
popSize, a, b = 2000, 0.001, 0.001
log_prob_vect, log_prob, log_prob_prior = logp_exp_vect(X), logp_exp(X), gamma_prior(a, b)
c, ρ, n_step_max = 0.99, 1/2, 25
paralell, n_proc = True, 4
start = time.time()
trace, log_marg = smc_exp(popSize, a, b, log_prob_vect, log_prob, log_prob_prior, c, ρ, n_step_max, paralell, n_proc)
print(f"Runtime of the program is {time.time() - start}")
print(trace.mean(), log_marg, norm_constant(X, a, b))
δ_pos = sample_pos_δ(X, a, b, popSize)
positions = np.linspace(min(δ_pos), max(δ_pos), 1000)
kernel = st.gaussian_kde(δ_pos)
plt.plot(positions, kernel(positions), lw=2, label = "n = "+str(n))
n, bins, patches = plt.hist(trace, num_bins, density=1)
```
or the sample size
```
def batch_size_search(λ, target, n_prev, X):
n_up, n_down = 2 * len(X), n_prev
while n_up - n_down > 1:
n_new = int(np.ceil((n_up + n_down) / 2))
log_prob = logp_exp_vect(X[n_prev:n_new])
w = np.exp(log_prob(λ))
if sum(w) == 0 or np.any(np.isnan(w)):
W = 1 / popSize *np.ones(popSize)
ESS = 0
else:
W = w / sum(w)
ESS = 1 / sum(W**2)
# print(γ_new, ESS)
if ESS == target:
break
else:
if ESS < target:
n_up = n_new
else:
n_down = n_new
if n_new >= len(X):
n_new = len(X)
log_prob = logp_exp_vect(X[n_prev:n_new])
w = np.exp(log_prob(λ))
W = w / sum(w)
ESS = 1 / sum(W**2)
return(n_new, w, W, int(ESS))
def smc_exp_batch(popSize, a, b, X, log_prob_prior, c, ρ, n_step_max,
paralell, n_proc):
# popSize = number of particles
# a,b = hyperparameter of the prior distribution on λ
# c = probability that a particle is moved at least once
# ρ = proportion of the sample of the ESS
# temp_step = size of the step in the search of the right temperature
# The parameter is gamma distributed
λ_prior = st.gamma(a)
# particles initialization
print('sample generation 0')
clouds = [pd.DataFrame({'λ': λ_prior.rvs(popSize) / b,
'w':np.ones(popSize)/ popSize,
'W': np.ones(popSize) / popSize
})]
# Temperature sequence
n_seq = np.array([0])
# Generation counter
g=0
# We keep on iterating until the temperature reaches 1
while n_seq[-1] < len(X):
# g, ESS, k = g + 1, 1, - err
g = g + 1
print('Sampling particles from generation ' + str(g))
cloud = pd.DataFrame({ 'λ':[], 'w':[], 'W':[]})
# Updating temperature sequence
n, w, W, ESS = batch_size_search(clouds[g-1]['λ'].values,\
ρ * popSize, n_seq[-1], X)
n_seq = np.append(n_seq, n)
# Updating unormalized weights
cloud['w'] = w
# Updating normalized weights
cloud['W'] = W
step_size = np.sqrt(np.cov(clouds[g-1]['λ'].values
, aweights=W))*2.38
particles = clouds[g-1]['λ'].values[np.random.choice(popSize,
popSize, p = W)]
log_prob = logp_exp(X[0:n])
def move_particle_trial(particle):
λ, accepted = MH_move_exp(1, step_size,
log_prob, log_prob_prior, particle, 1)
return([λ[-1], np.mean(accepted)])
if paralell:
res = np.matrix(Parallel(n_jobs=n_proc)(delayed(move_particle_trial)(i)
for i in particles))
else:
res = np.matrix([move_particle_trial(particle)
for particle in particles])
λ_trial = np.asarray(res[:,0]).flatten()
acc_rate = np.asarray(res[:,1]).flatten()
n_steps = min(n_step_max,
max(2,np.ceil(np.log(1-c) / np.log(1-np.mean(acc_rate)))))
print('Generation: ' + str(g) + " ;batch size: "+str(n_seq[-1]) +
" ;steps:" + str(n_steps) + " ;stepsize: "+str(step_size)+
" ;ESS: "+str(ESS))
def move_particle(particle):
λ, accepted = MH_move_exp(int(n_steps), step_size,
log_prob, log_prob_prior, particle, 1)
return([λ[-1], np.mean(accepted)])
if paralell:
res = np.matrix(Parallel(n_jobs=n_proc)(delayed(move_particle)(i)
for i in λ_trial))
else:
res = np.matrix([move_particle(particle) for particle in λ_trial])
cloud['λ'] = np.asarray(res[:,0]).flatten()
clouds.append(cloud)
marginal_log_likelihood = sum(np.log(([cloud['w'].mean()
for cloud in clouds[1:g+1]])))
return((clouds[-1].λ.values, marginal_log_likelihood))
popSize, a, b, log_prob_prior, c, ρ, n_step_max = 20000, 0.01, 0.01, gamma_prior(a, b), 0.99, 1/2, 25
paralell, n_proc = False, 4
start = time.time()
trace, log_marg = smc_exp_batch(popSize, a, b,X, log_prob_prior, c, ρ, n_step_max, paralell, n_proc)
print(f"Runtime of the program is {time.time() - start}")
print(trace.mean(), log_marg, norm_constant(X, a, b))
δ_pos = sample_pos_δ(X, a, b, popSize)
positions = np.linspace(min(δ_pos), max(δ_pos), 1000)
kernel = st.gaussian_kde(δ_pos)
plt.plot(positions, kernel(positions), lw=2, label = "n = "+str(n))
n, bins, patches = plt.hist(trace, num_bins, density=1)
```
## Simulation study
```
np.random.seed(123)
# Data simulation
# iid sample of size 50 drawn the exponential distribution with parameter 1
δ_true, X = 3, np.random.gamma(shape = 1, scale= 1 / δ_true, size=50)
log_prob_vect, log_prob = logp_exp_vect(X), logp_exp(X)
# Gamma prior with parameter a and b
a, b, log_prob_prior = 0.1, 0.1, gamma_prior(a, b)
# SMC algorith setting
c, ρ, n_step_max,paralell, n_proc = 0.99, 1/2, 25, False, 1
# Bridge sampling setting
r_init = 1
res_list = []
for popSize in [500, 2000, 5000]:
print("Nombre de particules = " + str(popSize) )
def compute_log_marg(i):
trace, log_marg_like = smc_exp(popSize, a, b, log_prob_vect, log_prob, log_prob_prior, c, ρ, n_step_max, paralell,
n_proc)
# smc_exp(popSize, a, b, log_prob_vect, log_prob, log_prob_prior, c, ρ, n_step_max, paralell, n_proc)
trace, log_marg_data = smc_exp_batch(popSize, a, b, X, log_prob_prior, c, ρ, n_step_max, paralell, n_proc)
# log_marg_bridge = bridge_log_normalizing_constant(X, a, b,log_prob, log_prob_prior, popSize, r_init)
return(np.array([log_marg_like, log_marg_data]))
res = np.array(Parallel(n_jobs=50)(delayed(compute_log_marg)(i) for i in range(100)))
res_list.append(res)
methods = ["smc simulated anealing", "smc data by batch"]
res = res_list[0]
positions = np.linspace(np.min(res), np.max(res), 1000)
linetypes = ["solid", "dashed"]
for k in range(2):
kernel = st.gaussian_kde(np.asarray(res[:,k]))
plt.plot(positions, kernel(positions), lw=2, label = methods[k], linestyle = linetypes[k])
plt.axvline(x = norm_constant(X, a, b), c = "black", lw=1, linestyle= "dotted", label = "log marginal likelihood")
plt.legend(fontsize = 6, bbox_to_anchor=(1.05, 1), loc='upper left', borderaxespad=0.)
for i in range(2):
res = res_list[i+1]
for k in range(2):
kernel = st.gaussian_kde(np.asarray(res[:,k]))
plt.plot(positions, kernel(positions), lw=2, linestyle = linetypes[k])
plt.axvline(x = norm_constant(X, a, b), c = "black", lw=1, linestyle= "dotted")
plt.legend(bbox_to_anchor=(0., 1.02, 1., .102), loc='lower left',
ncol=2, mode="expand", borderaxespad=0., frameon=False)
plt.text(5.71, 4,'$N = 500$', fontsize = 10)
plt.text(5.71, 8,'$N = 2000$', fontsize = 10)
plt.text(5.71, 12.5,'$N = 5000$', fontsize = 10)
sns.despine()
plt.savefig("../Figures/hist_exponential_normalizing_constant_en.pdf")
plt.show()
res = res_list[0]
stds = np.std(res, axis = 0)
for k in range(2):
res = res_list[k+1]
std_temp = np.std(res, axis = 0)
stds = np.vstack((stds, std_temp))
df_std = pd.DataFrame(stds, index= ["N=500", "N=2000","N=5000"], columns= ["Simulated anealing", "Data by batch"])
print(df_std.to_latex())
```
| github_jupyter |
# A notebook to check daymet data
The purpose of this notebook is to open and check some DayMet data that will be used to locate a point at which I will run the Biome-BGC biogeochemical model. Here we are just using xarray utilities to open, compute some quick stats on, and plot data in the DayMet tile in which Reynolds Creek CZO falls.
## 1. Dependencies and Declarations
The following libraries are required for this Jupyter notebook:
* Numpy - for loading, saving arrays
* xArray - for data management and statistical analysis of large netcdf dataset
* Matplotlib - for plotting result
```
import numpy as np
import xarray as xr
import matplotlib.pyplot as plt
DaymetPath = '/Users/lejoflores/data/daymet/'
prcpName = 'prcp_1980-2018.nc'
tmaxName = 'tmax_1980-2018.nc'
tminName = 'tmin_1980-2018.nc'
```
## 2. Construct the file name
```
prcp_file = DaymetPath + prcpName
print(prcp_file)
```
## 3. Open the dataset as an xarray dataset
```
ds = xr.open_dataset(prcp_file)
print(ds)
```
## 4. Get the latitude and longitude arrays, verify, and plot them
```
lat = ds['lat']
lon = ds['lon']
print(lat.shape)
print(lon.shape)
plt.figure(figsize=(8,6))
plt.imshow(lat)
plt.show()
plt.figure(figsize=(8,6))
plt.imshow(lon)
plt.show()
```
## 5. Group the dataset by year and take the mean through time
```
ds_ann = ds.groupby('time.year').mean('time')
ds_mean_ann = ds_ann.mean('year')
print(ds_mean_ann)
```
## 6. Compute the mean annual precipitation
```
MeanAnnPrcp = ds_mean_ann['prcp'].values
plt.figure(figsize=(16,12))
plt.imshow(MeanAnnPrcp*365.0)
plt.colorbar()
plt.show()
```
## 7. Also get and plot min, max, and coefficient of variation
```
ds_max_ann = ds_ann.max('year')
ds_min_ann = ds_ann.min('year')
MaxAnnPrcp = ds_max_ann['prcp'].values
MinAnnPrcp = ds_min_ann['prcp'].values
plt.figure(figsize=(16,12))
plt.imshow(MaxAnnPrcp*365.0)
plt.colorbar()
plt.show()
plt.figure(figsize=(16,12))
plt.imshow(MinAnnPrcp*365.0)
plt.colorbar()
plt.show()
ds_cv_ann = ds_ann.std('year')/ds_ann.mean('year')
CVAnnPrcp = ds_cv_ann['prcp'].values
plt.figure(figsize=(16,12))
plt.imshow(CVAnnPrcp*100.0)
plt.colorbar()
plt.show()
```
## 8. Do the same for $T_{max}$ and $T_{min}$
```
tmax_file = DaymetPath + tmaxName
tmin_file = DaymetPath + tminName
ds_tmax = xr.open_dataset(tmax_file)
ds_tmin = xr.open_dataset(tmin_file)
ds_tmax_ann = ds_tmax.groupby('time.year').mean('time')
ds_tmin_ann = ds_tmin.groupby('time.year').mean('time')
ds_tmax_mean_ann = ds_tmax_ann.mean('year')
ds_tmin_mean_ann = ds_tmin_ann.mean('year')
MeanAnnTmax = ds_tmax_mean_ann['tmax'].values
MeanAnnTmin = ds_tmin_mean_ann['tmin'].values
plt.figure(figsize=(16,12))
plt.imshow(MeanAnnTmax)
plt.colorbar()
plt.show()
plt.figure(figsize=(16,12))
plt.imshow(MeanAnnTmin)
plt.colorbar()
plt.show()
```
| github_jupyter |
# Neural networks with PyTorch
Deep learning networks tend to be massive with dozens or hundreds of layers, that's where the term "deep" comes from. You can build one of these deep networks using only weight matrices as we did in the previous notebook, but in general it's very cumbersome and difficult to implement. PyTorch has a nice module `nn` that provides a nice way to efficiently build large neural networks.
```
# Import necessary packages
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
import numpy as np
import torch
import helper
import matplotlib.pyplot as plt
```
Now we're going to build a larger network that can solve a (formerly) difficult problem, identifying text in an image. Here we'll use the MNIST dataset which consists of greyscale handwritten digits. Each image is 28x28 pixels, you can see a sample below
<img src='assets/mnist.png'>
Our goal is to build a neural network that can take one of these images and predict the digit in the image.
First up, we need to get our dataset. This is provided through the `torchvision` package. The code below will download the MNIST dataset, then create training and test datasets for us. Don't worry too much about the details here, you'll learn more about this later.
```
### Run this cell
from torchvision import datasets, transforms
# Define a transform to normalize the data
transform = transforms.Compose([transforms.ToTensor(),
transforms.Normalize((0.5,), (0.5,)),
])
# Download and load the training data
trainset = datasets.MNIST('~/.pytorch/MNIST_data/', download=True, train=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=64, shuffle=True)
```
We have the training data loaded into `trainloader` and we make that an iterator with `iter(trainloader)`. Later, we'll use this to loop through the dataset for training, like
```python
for image, label in trainloader:
## do things with images and labels
```
You'll notice I created the `trainloader` with a batch size of 64, and `shuffle=True`. The batch size is the number of images we get in one iteration from the data loader and pass through our network, often called a *batch*. And `shuffle=True` tells it to shuffle the dataset every time we start going through the data loader again. But here I'm just grabbing the first batch so we can check out the data. We can see below that `images` is just a tensor with size `(64, 1, 28, 28)`. So, 64 images per batch, 1 color channel, and 28x28 images.
```
dataiter = iter(trainloader)
images, labels = dataiter.next()
print(type(images))
print(images.shape)
print(labels.shape)
```
This is what one of the images looks like.
```
plt.imshow(images[1].numpy().squeeze(), cmap='Greys_r');
```
First, let's try to build a simple network for this dataset using weight matrices and matrix multiplications. Then, we'll see how to do it using PyTorch's `nn` module which provides a much more convenient and powerful method for defining network architectures.
The networks you've seen so far are called *fully-connected* or *dense* networks. Each unit in one layer is connected to each unit in the next layer. In fully-connected networks, the input to each layer must be a one-dimensional vector (which can be stacked into a 2D tensor as a batch of multiple examples). However, our images are 28x28 2D tensors, so we need to convert them into 1D vectors. Thinking about sizes, we need to convert the batch of images with shape `(64, 1, 28, 28)` to a have a shape of `(64, 784)`, 784 is 28 times 28. This is typically called *flattening*, we flattened the 2D images into 1D vectors.
Previously you built a network with one output unit. Here we need 10 output units, one for each digit. We want our network to predict the digit shown in an image, so what we'll do is calculate probabilities that the image is of any one digit or class. This ends up being a discrete probability distribution over the classes (digits) that tells us the most likely class for the image. That means we need 10 output units for the 10 classes (digits). We'll see how to convert the network output into a probability distribution next.
> **Exercise:** Flatten the batch of images `images`. Then build a multi-layer network with 784 input units, 256 hidden units, and 10 output units using random tensors for the weights and biases. For now, use a sigmoid activation for the hidden layer. Leave the output layer without an activation, we'll add one that gives us a probability distribution next.
```
## Your solution
out = # output of your network, should have shape (64,10)
```
Now we have 10 outputs for our network. We want to pass in an image to our network and get out a probability distribution over the classes that tells us the likely class(es) the image belongs to. Something that looks like this:
<img src='assets/image_distribution.png' width=500px>
Here we see that the probability for each class is roughly the same. This is representing an untrained network, it hasn't seen any data yet so it just returns a uniform distribution with equal probabilities for each class.
To calculate this probability distribution, we often use the [**softmax** function](https://en.wikipedia.org/wiki/Softmax_function). Mathematically this looks like
$$
\Large \sigma(x_i) = \cfrac{e^{x_i}}{\sum_k^K{e^{x_k}}}
$$
What this does is squish each input $x_i$ between 0 and 1 and normalizes the values to give you a proper probability distribution where the probabilites sum up to one.
> **Exercise:** Implement a function `softmax` that performs the softmax calculation and returns probability distributions for each example in the batch. Note that you'll need to pay attention to the shapes when doing this. If you have a tensor `a` with shape `(64, 10)` and a tensor `b` with shape `(64,)`, doing `a/b` will give you an error because PyTorch will try to do the division across the columns (called broadcasting) but you'll get a size mismatch. The way to think about this is for each of the 64 examples, you only want to divide by one value, the sum in the denominator. So you need `b` to have a shape of `(64, 1)`. This way PyTorch will divide the 10 values in each row of `a` by the one value in each row of `b`. Pay attention to how you take the sum as well. You'll need to define the `dim` keyword in `torch.sum`. Setting `dim=0` takes the sum across the rows while `dim=1` takes the sum across the columns.
```
def softmax(x):
## TODO: Implement the softmax function here
# Here, out should be the output of the network in the previous excercise with shape (64,10)
probabilities = softmax(out)
# Does it have the right shape? Should be (64, 10)
print(probabilities.shape)
# Does it sum to 1?
print(probabilities.sum(dim=1))
```
## Building networks with PyTorch
PyTorch provides a module `nn` that makes building networks much simpler. Here I'll show you how to build the same one as above with 784 inputs, 256 hidden units, 10 output units and a softmax output.
```
from torch import nn
class Network(nn.Module):
def __init__(self):
super().__init__()
# Inputs to hidden layer linear transformation
self.hidden = nn.Linear(784, 256)
# Output layer, 10 units - one for each digit
self.output = nn.Linear(256, 10)
# Define sigmoid activation and softmax output
self.sigmoid = nn.Sigmoid()
self.softmax = nn.Softmax(dim=1)
def forward(self, x):
# Pass the input tensor through each of our operations
x = self.hidden(x)
x = self.sigmoid(x)
x = self.output(x)
x = self.softmax(x)
return x
```
Let's go through this bit by bit.
```python
class Network(nn.Module):
```
Here we're inheriting from `nn.Module`. Combined with `super().__init__()` this creates a class that tracks the architecture and provides a lot of useful methods and attributes. It is mandatory to inherit from `nn.Module` when you're creating a class for your network. The name of the class itself can be anything.
```python
self.hidden = nn.Linear(784, 256)
```
This line creates a module for a linear transformation, $x\mathbf{W} + b$, with 784 inputs and 256 outputs and assigns it to `self.hidden`. The module automatically creates the weight and bias tensors which we'll use in the `forward` method. You can access the weight and bias tensors once the network (`net`) is created with `net.hidden.weight` and `net.hidden.bias`.
```python
self.output = nn.Linear(256, 10)
```
Similarly, this creates another linear transformation with 256 inputs and 10 outputs.
```python
self.sigmoid = nn.Sigmoid()
self.softmax = nn.Softmax(dim=1)
```
Here I defined operations for the sigmoid activation and softmax output. Setting `dim=1` in `nn.Softmax(dim=1)` calculates softmax across the columns.
```python
def forward(self, x):
```
PyTorch networks created with `nn.Module` must have a `forward` method defined. It takes in a tensor `x` and passes it through the operations you defined in the `__init__` method.
```python
x = self.hidden(x)
x = self.sigmoid(x)
x = self.output(x)
x = self.softmax(x)
```
Here the input tensor `x` is passed through each operation a reassigned to `x`. We can see that the input tensor goes through the hidden layer, then a sigmoid function, then the output layer, and finally the softmax function. It doesn't matter what you name the variables here, as long as the inputs and outputs of the operations match the network architecture you want to build. The order in which you define things in the `__init__` method doesn't matter, but you'll need to sequence the operations correctly in the `forward` method.
Now we can create a `Network` object.
```
# Create the network and look at it's text representation
model = Network()
model
```
You can define the network somewhat more concisely and clearly using the `torch.nn.functional` module. This is the most common way you'll see networks defined as many operations are simple element-wise functions. We normally import this module as `F`, `import torch.nn.functional as F`.
```
import torch.nn.functional as F
class Network(nn.Module):
def __init__(self):
super().__init__()
# Inputs to hidden layer linear transformation
self.hidden = nn.Linear(784, 256)
# Output layer, 10 units - one for each digit
self.output = nn.Linear(256, 10)
def forward(self, x):
# Hidden layer with sigmoid activation
x = F.sigmoid(self.hidden(x))
# Output layer with softmax activation
x = F.softmax(self.output(x), dim=1)
return x
```
### Activation functions
So far we've only been looking at the softmax activation, but in general any function can be used as an activation function. The only requirement is that for a network to approximate a non-linear function, the activation functions must be non-linear. Here are a few more examples of common activation functions: Tanh (hyperbolic tangent), and ReLU (rectified linear unit).
<img src="assets/activation.png" width=700px>
In practice, the ReLU function is used almost exclusively as the activation function for hidden layers.
### Your Turn to Build a Network
<img src="assets/mlp_mnist.png" width=600px>
> **Exercise:** Create a network with 784 input units, a hidden layer with 128 units and a ReLU activation, then a hidden layer with 64 units and a ReLU activation, and finally an output layer with a softmax activation as shown above. You can use a ReLU activation with the `nn.ReLU` module or `F.relu` function.
```
## Your solution here
```
### Initializing weights and biases
The weights and such are automatically initialized for you, but it's possible to customize how they are initialized. The weights and biases are tensors attached to the layer you defined, you can get them with `model.fc1.weight` for instance.
```
print(model.fc1.weight)
print(model.fc1.bias)
```
For custom initialization, we want to modify these tensors in place. These are actually autograd *Variables*, so we need to get back the actual tensors with `model.fc1.weight.data`. Once we have the tensors, we can fill them with zeros (for biases) or random normal values.
```
# Set biases to all zeros
model.fc1.bias.data.fill_(0)
# sample from random normal with standard dev = 0.01
model.fc1.weight.data.normal_(std=0.01)
```
### Forward pass
Now that we have a network, let's see what happens when we pass in an image.
```
# Grab some data
dataiter = iter(trainloader)
images, labels = dataiter.next()
# Resize images into a 1D vector, new shape is (batch size, color channels, image pixels)
images.resize_(64, 1, 784)
# or images.resize_(images.shape[0], 1, 784) to automatically get batch size
# Forward pass through the network
img_idx = 0
ps = model.forward(images[img_idx,:])
img = images[img_idx]
helper.view_classify(img.view(1, 28, 28), ps)
```
As you can see above, our network has basically no idea what this digit is. It's because we haven't trained it yet, all the weights are random!
### Using `nn.Sequential`
PyTorch provides a convenient way to build networks like this where a tensor is passed sequentially through operations, `nn.Sequential` ([documentation](https://pytorch.org/docs/master/nn.html#torch.nn.Sequential)). Using this to build the equivalent network:
```
# Hyperparameters for our network
input_size = 784
hidden_sizes = [128, 64]
output_size = 10
# Build a feed-forward network
model = nn.Sequential(nn.Linear(input_size, hidden_sizes[0]),
nn.ReLU(),
nn.Linear(hidden_sizes[0], hidden_sizes[1]),
nn.ReLU(),
nn.Linear(hidden_sizes[1], output_size),
nn.Softmax(dim=1))
print(model)
# Forward pass through the network and display output
images, labels = next(iter(trainloader))
images.resize_(images.shape[0], 1, 784)
ps = model.forward(images[0,:])
helper.view_classify(images[0].view(1, 28, 28), ps)
```
Here our model is the same as before: 784 input units, a hidden layer with 128 units, ReLU activation, 64 unit hidden layer, another ReLU, then the output layer with 10 units, and the softmax output.
The operations are available by passing in the appropriate index. For example, if you want to get first Linear operation and look at the weights, you'd use `model[0]`.
```
print(model[0])
model[0].weight
```
You can also pass in an `OrderedDict` to name the individual layers and operations, instead of using incremental integers. Note that dictionary keys must be unique, so _each operation must have a different name_.
```
from collections import OrderedDict
model = nn.Sequential(OrderedDict([
('fc1', nn.Linear(input_size, hidden_sizes[0])),
('relu1', nn.ReLU()),
('fc2', nn.Linear(hidden_sizes[0], hidden_sizes[1])),
('relu2', nn.ReLU()),
('output', nn.Linear(hidden_sizes[1], output_size)),
('softmax', nn.Softmax(dim=1))]))
model
```
Now you can access layers either by integer or the name
```
print(model[0])
print(model.fc1)
```
In the next notebook, we'll see how we can train a neural network to accuractly predict the numbers appearing in the MNIST images.
| github_jupyter |
```
import pathlib
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
import numpy as np
print(tf.__version__)
```
# Auto MPG data
```
# dataset_path = keras.utils.get_file("auto-mpg.data",
# "http://archive.ics.uci.edu/ml/machine-learning-databases/auto-mpg/auto-mpg.data")
# dataset_path
dataset_path = '/Volumes/TimeMachine/data/DR7/eBOSS.ELG.NGC.DR7.table.5.r.npy'
data = np.load(dataset_path, allow_pickle=True).item()
data.keys()
class DATA(object):
def __init__(self, data):
self.x = data['features']
self.y = data['label']
self.p = data['hpind']
self.w = data['fracgood']
def norm_it(self, train_stats):
self.X = (self.x - train_stats['x']['mean'])/train_stats['x']['std']
self.Y = (self.y - train_stats['y']['mean'])/train_stats['y']['std']
# Display training progress by printing a single dot for each completed epoch
class PrintDot(keras.callbacks.Callback):
def on_epoch_end(self, epoch, logs):
if epoch % 10 == 0: print('')
print('.', end='')
#
def plot_history(history):
hist = pd.DataFrame(history.history)
hist['epoch'] = history.epoch
fig, ax = plt.subplots(nrows=2, sharex=True, figsize=(8, 8))
plt.subplots_adjust(hspace=0.0)
# fig 0
ax[0].set_ylabel('Mean Abs Error')
ax[0].plot(hist['epoch'], hist['mae'],
label='Train Error')
ax[0].plot(hist['epoch'], hist['val_mae'],
label = 'Val Error', ls='--')
# fig 1
ax[1].set_xlabel('Epoch')
ax[1].set_ylabel('Mean Square Error')
ax[1].plot(hist['epoch'], hist['mse'],
label='Train Error')
ax[1].plot(hist['epoch'], hist['val_mse'],
label = 'Val Error', ls='--')
ax[0].legend()
plt.show()
train = DATA(data['train']['fold3'])
test = DATA(data['test']['fold3'])
valid = DATA(data['validation']['fold3'])
train
train_stats = {'x':{'mean':np.mean(train.x, axis=0),
'std':np.std(train.x, axis=0, ddof=1)},
'y':{'mean':np.mean(train.y, axis=0),
'std':np.std(train.y, axis=0, ddof=1)}}
train.norm_it(train_stats)
test.norm_it(train_stats)
valid.norm_it(train_stats)
train.X.shape
def build_model(nfeature):
tf.keras.backend.clear_session()
kwargs = dict(kernel_regularizer=keras.regularizers.l2(0.0),
kernel_initializer=keras.initializers.he_normal(seed=123456))
model = keras.Sequential([
layers.Dense(20, activation='relu', input_shape=[nfeature], **kwargs),
layers.Dense(20, activation='relu', **kwargs),
layers.Dense(1, **kwargs)
])
#optimizer = tf.keras.optimizers.RMSprop(0.001)
optimizer = tf.keras.optimizers.Adam(learning_rate=0.001)
model.compile(loss='mse',
optimizer=optimizer,
metrics=['mae', 'mse'])
return model
model = build_model(18)
model.summary()
example_batch = train.X[:10]
example_result = model.predict(example_batch)
example_result
model = build_model(18)
# The patience parameter is the amount of epochs to check for improvement
early_stop = keras.callbacks.EarlyStopping(monitor='val_loss', patience=10, min_delta=1.e-6)
EPOCHS = 1000
history = model.fit(
train.X, train.Y,
epochs=EPOCHS, validation_data=(valid.X, valid.Y), verbose=0,
callbacks=[early_stop, PrintDot()], batch_size=256)
plot_history(history)
loss, mae, mse = model.evaluate(test.X, test.y, verbose=0)
print("Testing set Mean Abs Error : {:5.2f}\nTesting set Mean Square Error : {:5.2f}".format(mae, mse))
pd.DataFrame(history.history).tail()
test_predictions = model.predict(test.X).flatten()
plt.scatter(test.Y, test_predictions, alpha=0.1)
plt.xlabel('True Values')
plt.ylabel('Predictions')
plt.axis('equal')
plt.axis('square')
# plt.xlim([0,plt.xlim()[1]])
# plt.ylim([0,plt.ylim()[1]])
# _ = plt.plot([-100, 100], [-100, 100])
error = test_predictions - test.Y
plt.hist(error, bins=30)
plt.xlabel("Prediction Error")
# plt.yscale('log')
_ = plt.ylabel("Count")
```
| github_jupyter |
# Continuous Bag of Words (CBOW) Text Classifier
The code below implements a continuous bag of words text classifier.
- We tokenize the text, create a vocabulary and encode each piece of text in the dataset
- The lookup allows for extracting embeddings for each tokenized input
- The embedding vectors are added together
- The resulting vector is multiplied with a weight matrix, which is then added a bias vector; this results in scores
- The scores are applied a softmax to generate probabilities which are used for the final classification
The code used in this notebook was inspired by code from the [official repo](https://github.com/neubig/nn4nlp-code) used in the [CMU Neural Networks for NLP class](http://www.phontron.com/class/nn4nlp2021/schedule.html) by [Graham Neubig](http://www.phontron.com/index.php).

```
import torch
import random
import torch.nn as nn
%%capture
# download the files
!wget https://raw.githubusercontent.com/neubig/nn4nlp-code/master/data/classes/dev.txt
!wget https://raw.githubusercontent.com/neubig/nn4nlp-code/master/data/classes/test.txt
!wget https://raw.githubusercontent.com/neubig/nn4nlp-code/master/data/classes/train.txt
# create the data folders
!mkdir data data/classes
!cp dev.txt data/classes
!cp test.txt data/classes
!cp train.txt data/classes
```
## Read and Process Data
```
# function to read in data, process each line and split columns by " ||| "
def read_data(filename):
data = []
with open(filename, 'r') as f:
for line in f:
line = line.lower().strip()
line = line.split(' ||| ')
data.append(line)
return data
train_data = read_data('data/classes/train.txt')
test_data = read_data('data/classes/test.txt')
# creating the word and tag indices
word_to_index = {}
word_to_index["<unk>"] = len(word_to_index) # add <UNK> to dictionary
tag_to_index = {}
# create word to index dictionary and tag to index dictionary from data
def create_dict(data, check_unk=False):
for line in data:
for word in line[1].split(" "):
if check_unk == False:
if word not in word_to_index:
word_to_index[word] = len(word_to_index)
else:
if word not in word_to_index:
word_to_index[word] = word_to_index["<unk>"]
if line[0] not in tag_to_index:
tag_to_index[line[0]] = len(tag_to_index)
create_dict(train_data)
create_dict(test_data, check_unk=True)
# create word and tag tensors from data
def create_tensor(data):
for line in data:
yield([word_to_index[word] for word in line[1].split(" ")], tag_to_index[line[0]])
train_data = list(create_tensor(train_data))
test_data = list(create_tensor(test_data))
number_of_words = len(word_to_index)
number_of_tags = len(tag_to_index)
```
## Model
```
# cpu or gpu
device = "cuda" if torch.cuda.is_available() else "cpu"
# create a simple neural network with embedding layer, bias, and xavier initialization
class CBoW(torch.nn.Module):
def __init__(self, nwords, ntags, emb_size):
super(CBoW, self).__init__()
# layers
self.embedding = torch.nn.Embedding(nwords, emb_size)
self.linear = torch.nn.Linear(emb_size, ntags)
# use xavier initialization for weights
nn.init.xavier_uniform_(self.embedding.weight)
nn.init.xavier_uniform_(self.linear.weight)
def forward(self, x):
emb = self.embedding(x) # seq x emb_size
out = torch.sum(emb, dim=0) # emb_size
out = out.view(1, -1) # reshape to (1, emb_size)
out = self.linear(out) # 1 x ntags
return out
EMB_SIZE = 64
model = CBoW(number_of_words, number_of_tags, EMB_SIZE)
criterion = torch.nn.CrossEntropyLoss()
optimizer = torch.optim.Adam(model.parameters())
type = torch.LongTensor
if torch.cuda.is_available():
model.to(device)
type = torch.cuda.LongTensor
# perform training of the Bow model
for epoch in range(10):
# perform training
model.train()
random.shuffle(train_data)
total_loss = 0.0
train_correct = 0
for sentence, tag in train_data:
sentence = torch.tensor(sentence).type(type)
tag = torch.tensor([tag]).type(type)
output = model(sentence)
predicted = torch.argmax(output.data.detach()).item()
loss = criterion(output, tag)
total_loss += loss.item()
optimizer.zero_grad()
loss.backward()
optimizer.step()
if predicted == tag: train_correct+=1
# perform testing of the model
model.eval()
test_correct = 0
for sentence, tag in test_data:
sentence = torch.tensor(sentence).type(type)
output = model(sentence)
predicted = torch.argmax(output.data.detach()).item()
if predicted == tag: test_correct += 1
# print model performance results
log = f'epoch: {epoch+1} | ' \
f'train loss/sent: {total_loss/len(train_data):.4f} | ' \
f'train accuracy: {train_correct/len(train_data):.4f} | ' \
f'test accuracy: {test_correct/len(test_data):.4f}'
print(log)
```
| github_jupyter |
# 7. Question groups variation analysis
Pretest posttest answers variation analysis
## Table of Contents
[Preparation](#Preparation)
[Functions](#Functions)
# Preparation
```
from pySankey import sankey
%run "../Functions/6. Time analysis.ipynb"
print("7. Question groups variation analysis")
```
# Functions
## Per question analysis
### Interest variation
### Binary analysis
```
def analyseQuestion(allData, q):
pretestScores = allData.loc[answerTemporalities[0] + " " + q, :]
posttestScores = allData.loc[answerTemporalities[1] + " " + q, :]
deltaScores = allData.loc[deltaPrefix + " " + q, :]
print("variation: %0.2f (+/- %0.2f)" % (deltaScores.mean(), deltaScores.std()))
print("from %0.2f (+/- %0.2f) to %0.2f (+/- %0.2f)" % \
(pretestScores.mean(), pretestScores.std(),\
posttestScores.mean(), posttestScores.std(),))
plt.boxplot(deltaScores)
plt.show()
# questionsCoding contains points attributed to each answer
def compareUsingCustomCorrection(gfdf, questions, questionsCoding):
minPotentialScore = 0
maxPotentialScore = 0
for gradingDictionary in questionsCoding:
minPotentialScore += min(gradingDictionary.values())
maxPotentialScore += max(gradingDictionary.values())
minPotentialScore, maxPotentialScore
print("%s < score < %s" % (minPotentialScore, maxPotentialScore))
# split temporalities
gfdfPretest = gfdf[gfdf[QTemporality]==answerTemporalities[0]]
gfdfPretest.index = gfdfPretest[QUserId]
gfdfPostest = gfdf[gfdf[QTemporality]==answerTemporalities[1]]
gfdfPostest.index = gfdfPostest[QUserId]
# only keep relevant questions
gfdfPretest = gfdfPretest.loc[:, questions]
gfdfPostest = gfdfPostest.loc[:, questions]
# code the answers
for (q, c) in zip(questions, questionsCoding):
gfdfPretest[q] = gfdfPretest[q].apply(lambda t: c[t])
gfdfPostest[q] = gfdfPostest[q].apply(lambda t: c[t])
# compute delta
# gfdfDelta = gfdfPostest - gfdfPretest
gfdfResult = gfdfPostest - gfdfPretest
gfdfResult.columns = [deltaPrefix + " " + q for q in questions]
gfdfResult[[answerTemporalities[0] + " " + q for q in questions]] = gfdfPretest
gfdfResult[[answerTemporalities[1] + " " + q for q in questions]] = gfdfPostest
return gfdfResult.T
def getScoresOnQuestionsFromAllData(allData, Qs):
pretestQs = [answerTemporalities[0] + " " + q for q in Qs]
posttestQs = [answerTemporalities[1] + " " + q for q in Qs]
deltaQs = [deltaPrefix + " " + q for q in Qs]
pretestScores = allData.loc[pretestQs, :].sum()
posttestScores = allData.loc[posttestQs, :].sum()
deltaScores = allData.loc[deltaQs, :].sum()
return (pretestScores, posttestScores, deltaScores)
def analyseQuestionGroup(
gfdf,
Qs,
grading,
plotGraphs = True,
printData = True,
saveFiles = False,
title = "",
qualitativeCoding = None,
):
allData = compareUsingCustomCorrection(gfdf, Qs, grading)
(pretestScores, posttestScores, deltaScores) = getScoresOnQuestionsFromAllData(allData, Qs)
questionGroupStem = "question group"
pretestColumn = answerTemporalities[0] + " " + questionGroupStem
posttestColumn = answerTemporalities[1] + " " + questionGroupStem
deltaColumn = deltaPrefix + " " + questionGroupStem
gfdfResult = pd.DataFrame(
data = [deltaScores, pretestScores, posttestScores],
columns = deltaScores.index,
index = [deltaColumn, pretestColumn, posttestColumn],
)
plotPretestPosttestDeltaGfdf(
gfdfResult,
[questionGroupStem],
plotGraphs = plotGraphs,
printData = printData,
saveFiles = saveFiles,
title = title,
qualitativeCoding = qualitativeCoding,
)
```
## Plot
```
def getReorderedByPrefix(prefixes, prefixed):
result = []
for prefix in prefixes:
boolIndex = [s.startswith(prefix) for s in prefixed]
if any(boolIndex):
result += [prefixed[boolIndex.index(True)]]
return result
def plotPretestPosttestDeltaGfdf(allData,
questions,
plotGraphs = True,
printData = True,
saveFiles = False,
title = "",
suffix = "",
fontsize=10,
qualitativeCoding = None):
variationSuffix = ' - variation'
pretestPosttestSuffix = ' - pretest posttest'
sankeySuffix = ' - Sankey'
qualitativeSuffix = ' - qualitative'
# sample size
print("n = " + str(len(allData.columns)))
print()
print()
for q in questions:
deltaScores = allData.loc[deltaPrefix + " " +q ,:]
pretestScores = allData.loc[answerTemporalities[0] + " " + q ,:]
posttestScores = allData.loc[answerTemporalities[1] + " " + q ,:]
if qualitativeCoding != None:
if (not all([(i in qualitativeCoding.keys()) for i in pretestScores.values]))\
or (not all([(i in qualitativeCoding.keys()) for i in posttestScores.values])):
qualitativeCoding = None
if printData:
print(q)
print("variation: %0.2f (+/- %0.2f)" % (deltaScores.mean(), deltaScores.std()))
print("from %0.2f (+/- %0.2f) to %0.2f (+/- %0.2f)" % \
(pretestScores.mean(), pretestScores.std(),\
posttestScores.mean(), posttestScores.std(),))
print(ttest_ind(pretestScores, posttestScores))
if plotGraphs:
#plt.boxplot(deltaScores)
#plt.show()
fig = plt.figure()
ax = plt.subplot(111)
# if pd.isnull(deltaScores).any():
# print("pd.isnull(deltaScores).any(): " + str(deltaScores.index[pd.isnull(deltaScores)]))
plt.hist(deltaScores, bins=int(max(deltaScores) - min(deltaScores) + 1), figure = fig)
#sns.distplot(deltaScores, bins = np.arange(min(deltaScores),max(deltaScores)))
if len(title) == 0:
_title = '"' + q + '"' + variationSuffix + suffix
else:
_title = title + variationSuffix
plt.title(_title)
plt.xlabel("score variation")
plt.ylabel("count")
plt.show()
if saveFiles:
fig.savefig(_title.replace('"', ""))
fig = plt.figure()
ax = plt.subplot(111)
plt.hist(pretestScores, bins=int(max(pretestScores) - min(pretestScores) + 1), label='pretest', alpha=0.5, figure = fig)
plt.hist(posttestScores, bins=int(max(posttestScores) - min(posttestScores) + 1), label='posttest', alpha=0.5, figure = fig)
plt.legend()
if len(title) == 0:
_title = '"' + q + '"' + pretestPosttestSuffix + suffix
else:
_title = title + pretestPosttestSuffix
plt.title(_title)
plt.xlabel("score")
plt.ylabel("count")
plt.show()
if saveFiles:
fig.savefig(_title.replace('"', ""))
classesDF = pd.DataFrame(columns = ['pretest', 'posttest'])
# indexes should be scorePretest->scorePosttest for each such existing pair
# label is then the str(score)
weight = pd.Series()
for userId in pretestScores.index:
changeIndex = "{0:0=2d}".format(int(pretestScores[userId]))+"->{0:0=2d}".format(int(posttestScores[userId]))
if changeIndex in weight.index:
weight[changeIndex] += 1
else:
weight[changeIndex] = 1
if qualitativeCoding != None:
classesDF.loc[changeIndex, 'pretest'] = qualitativeCoding[int(pretestScores[userId])] \
+ " (" + "{0:0=2d}".format(len(pretestScores[pretestScores == pretestScores[userId]])) + ")"
classesDF.loc[changeIndex, 'posttest'] = qualitativeCoding[int(posttestScores[userId])] \
+ " (" + "{0:0=2d}".format(len(posttestScores[posttestScores == posttestScores[userId]])) + ")"
else:
classesDF.loc[changeIndex, 'pretest'] = "{0:0=2d}".format(int(pretestScores[userId])) \
+ " (" + "{0:0=2d}".format(len(pretestScores[pretestScores == pretestScores[userId]])) + ")"
classesDF.loc[changeIndex, 'posttest'] = "{0:0=2d}".format(int(posttestScores[userId])) \
+ " (" + "{0:0=2d}".format(len(posttestScores[posttestScores == posttestScores[userId]])) + ")"
left = classesDF['pretest'].sort_index().values
right = classesDF['posttest'].sort_index().values
leftWeight = weight.sort_index().values.astype(float)
rightWeight = leftWeight
if qualitativeCoding != None:
leftLabels = getReorderedByPrefix(qualitativeCoding.values(), classesDF['pretest'].unique())
rightLabels = getReorderedByPrefix(qualitativeCoding.values(), classesDF['posttest'].unique())
else:
leftLabels = sorted(classesDF['pretest'].unique())
rightLabels = sorted(classesDF['posttest'].unique())
if len(title) == 0:
_title = '"' + q + '"' + sankeySuffix
if qualitativeCoding != None:
_title += qualitativeSuffix
_title += suffix
else:
_title = title + sankeySuffix
if qualitativeCoding != None:
_title += qualitativeSuffix
if saveFiles:
filename = _title.replace('"', "")
else:
filename = None
sankey.sankey(
left=left,
right=right,
leftWeight=leftWeight,
rightWeight=rightWeight,
leftLabels=leftLabels,
rightLabels=rightLabels,
aspect=20,
fontsize=fontsize,
figureName=filename,
title=_title,
)
if printData:
print()
print()
print()
```
## Gradings
```
def getDeviceQuestionsGrading(correctAnsCost,halfCorAnsCost,dontKnoAnsCost,incorreAnsCost,):
r0 = DeviceAnswersPossibleAnswersEN[0]
r1 = DeviceAnswersPossibleAnswersEN[1]
r2 = DeviceAnswersPossibleAnswersEN[2]
r3 = DeviceAnswersPossibleAnswersEN[3]
r4 = DeviceAnswersPossibleAnswersEN[4]
r5 = DeviceAnswersPossibleAnswersEN[5]
QDeviceRbsPconsFlhdcTerCoding = {r0:correctAnsCost, r1:incorreAnsCost, r2:incorreAnsCost, r3:incorreAnsCost, r4:incorreAnsCost, r5:dontKnoAnsCost,}
QDevicePconsRbsFlhdcTerCoding = {r0:incorreAnsCost, r1:halfCorAnsCost, r2:halfCorAnsCost, r3:correctAnsCost, r4:halfCorAnsCost, r5:dontKnoAnsCost,}
QDevicePbadRbsGfpTerCoding = {r0:incorreAnsCost, r1:halfCorAnsCost, r2:correctAnsCost, r3:halfCorAnsCost, r4:halfCorAnsCost, r5:dontKnoAnsCost,}
QDevicePbadGfpRbsTerCoding = {r0:correctAnsCost, r1:incorreAnsCost, r2:incorreAnsCost, r3:incorreAnsCost, r4:incorreAnsCost, r5:dontKnoAnsCost,}
QDeviceGfpRbsPconsTerCoding = {r0:correctAnsCost, r1:incorreAnsCost, r2:incorreAnsCost, r3:incorreAnsCost, r4:incorreAnsCost, r5:dontKnoAnsCost,}
QDevicePconsGfpRbsTerCoding = {r0:correctAnsCost, r1:incorreAnsCost, r2:incorreAnsCost, r3:incorreAnsCost, r4:incorreAnsCost, r5:dontKnoAnsCost,}
QDeviceAmprRbsPconsTerCoding = {r0:correctAnsCost, r1:incorreAnsCost, r2:incorreAnsCost, r3:incorreAnsCost, r4:incorreAnsCost, r5:dontKnoAnsCost,}
QDeviceRbsPconsAmprTerCoding = {r0:correctAnsCost, r1:incorreAnsCost, r2:incorreAnsCost, r3:incorreAnsCost, r4:incorreAnsCost, r5:dontKnoAnsCost,}
return [
QDeviceRbsPconsFlhdcTerCoding,
QDevicePconsRbsFlhdcTerCoding,
QDevicePbadRbsGfpTerCoding,
QDevicePbadGfpRbsTerCoding,
QDeviceGfpRbsPconsTerCoding,
QDevicePconsGfpRbsTerCoding,
QDeviceAmprRbsPconsTerCoding,
QDeviceRbsPconsAmprTerCoding,
]
def getBioBrickFunctionsQuestionsGrading(correctAnsCost,halfCorAnsCost,dontKnoAnsCost,incorreAnsCost,):
r0 = BioBrickAnswersPossibleAnswersEN[0] #"None of these"
r1 = BioBrickAnswersPossibleAnswersEN[1] #"TER"
r2 = BioBrickAnswersPossibleAnswersEN[2] #"PR"
r3 = BioBrickAnswersPossibleAnswersEN[3] #"CDS"
r4 = BioBrickAnswersPossibleAnswersEN[4] #"RBS"
r5 = BioBrickAnswersPossibleAnswersEN[5] #"Plasmid"
r6 = BioBrickAnswersPossibleAnswersEN[6] #"I don't know"
QBBFunctionTERCoding = {r0:incorreAnsCost, r1:correctAnsCost, r2:halfCorAnsCost, r3:halfCorAnsCost, r4:halfCorAnsCost, r5:incorreAnsCost, r6:dontKnoAnsCost,}
QBBFunctionGameCDSCoding = {r0:incorreAnsCost, r1:halfCorAnsCost, r2:halfCorAnsCost, r3:correctAnsCost, r4:halfCorAnsCost, r5:incorreAnsCost, r6:dontKnoAnsCost,}
QBBFunctionBiologyCDSCoding = {r0:incorreAnsCost, r1:halfCorAnsCost, r2:halfCorAnsCost, r3:correctAnsCost, r4:halfCorAnsCost, r5:incorreAnsCost, r6:dontKnoAnsCost,}
QBBFunctionPRCoding = {r0:incorreAnsCost, r1:halfCorAnsCost, r2:correctAnsCost, r3:halfCorAnsCost, r4:halfCorAnsCost, r5:incorreAnsCost, r6:dontKnoAnsCost,}
QBBFunctionRBSCoding = {r0:incorreAnsCost, r1:halfCorAnsCost, r2:halfCorAnsCost, r3:halfCorAnsCost, r4:correctAnsCost, r5:incorreAnsCost, r6:dontKnoAnsCost,}
return [
QBBFunctionTERCoding,
QBBFunctionGameCDSCoding,
QBBFunctionBiologyCDSCoding,
QBBFunctionPRCoding,
QBBFunctionRBSCoding,
]
def getQGenotypePhenotypeGrading(correctAnsCost,halfCorAnsCost,dontKnoAnsCost,incorreAnsCost,):
r0 = QGenotypePhenotypePossibleAnswersEN[0]
r1 = QGenotypePhenotypePossibleAnswersEN[1]
r1bis = 'Gather nanorobots'
r2 = QGenotypePhenotypePossibleAnswersEN[2]
r3 = QGenotypePhenotypePossibleAnswersEN[3]
r4 = QGenotypePhenotypePossibleAnswersEN[4]
# 'Edit the DNA of the bacterium',
# 'Gather nanobots',
# 'Gather nanorobots',
# 'Move the bacterium',
# 'Divide the bacterium',
# "I don't know"
QGenotypePhenotypeGrading = {r0:correctAnsCost, r1:incorreAnsCost, r1bis:incorreAnsCost, r2:halfCorAnsCost, r3:incorreAnsCost, r4:dontKnoAnsCost,}
return [
QGenotypePhenotypeGrading
]
def getInductionQuestionsGrading(correctAnsCost,halfCorAnsCost,dontKnoAnsCost,incorreAnsCost,):
r0 = DeviceAnswersPossibleAnswersEN[0]
r1 = DeviceAnswersPossibleAnswersEN[1]
r2 = DeviceAnswersPossibleAnswersEN[2]
r3 = DeviceAnswersPossibleAnswersEN[3]
r4 = DeviceAnswersPossibleAnswersEN[4]
r5 = QDevicePbadRbsAraTerPossibleAnswersEN[0]
r6 = QDevicePbadRbsAraTerPossibleAnswersEN[1]
r7 = QDevicePbadRbsAraTerPossibleAnswersEN[2]
r8 = QDevicePbadRbsAraTerPossibleAnswersEN[3]
rIDK = DeviceAnswersPossibleAnswersEN[5]
# 'The bricks are not well-ordered',
# 'It generates green fluorescence',
# 'It generates green fluorescence in presence of arabinose inducer',
# 'It makes it possible to move faster',
# 'It generates antibiotic resistance'
QDeviceRbsPconsFlhdcTerGrading = {r0:halfCorAnsCost, r1:halfCorAnsCost, r2:incorreAnsCost, r3:halfCorAnsCost, r4:halfCorAnsCost, rIDK:dontKnoAnsCost,}
QDevicePconsRbsFlhdcTerGrading = {r0:halfCorAnsCost, r1:halfCorAnsCost, r2:incorreAnsCost, r3:halfCorAnsCost, r4:halfCorAnsCost, rIDK:dontKnoAnsCost,}
QDeviceGfpRbsPconsTerGrading = {r0:halfCorAnsCost, r1:halfCorAnsCost, r2:incorreAnsCost, r3:halfCorAnsCost, r4:halfCorAnsCost, rIDK:dontKnoAnsCost,}
QDevicePconsGfpRbsTerGrading = {r0:halfCorAnsCost, r1:halfCorAnsCost, r2:incorreAnsCost, r3:halfCorAnsCost, r4:halfCorAnsCost, rIDK:dontKnoAnsCost,}
QDeviceAmprRbsPconsTerGrading = {r0:halfCorAnsCost, r1:halfCorAnsCost, r2:incorreAnsCost, r3:halfCorAnsCost, r4:halfCorAnsCost, rIDK:dontKnoAnsCost,}
QDeviceRbsPconsAmprTerGrading = {r0:halfCorAnsCost, r1:halfCorAnsCost, r2:incorreAnsCost, r3:halfCorAnsCost, r4:halfCorAnsCost, rIDK:dontKnoAnsCost,}
QDevicePbadRbsGfpTerGrading = {r0:halfCorAnsCost, r1:incorreAnsCost, r2:correctAnsCost, r3:halfCorAnsCost, r4:halfCorAnsCost, rIDK:dontKnoAnsCost,}
QDevicePbadGfpTbsTerGrading = {r0:halfCorAnsCost, r1:halfCorAnsCost, r2:halfCorAnsCost, r3:halfCorAnsCost, r4:halfCorAnsCost, rIDK:dontKnoAnsCost,}
QDevicePbadRbsAraTerGrading = {r5:halfCorAnsCost, r6:halfCorAnsCost, r7:correctAnsCost, r8:incorreAnsCost, rIDK:dontKnoAnsCost,}
return [
QDeviceRbsPconsFlhdcTerGrading,
QDevicePconsRbsFlhdcTerGrading,
QDeviceGfpRbsPconsTerGrading,
QDevicePconsGfpRbsTerGrading,
QDeviceAmprRbsPconsTerGrading,
QDeviceRbsPconsAmprTerGrading,
QDevicePbadRbsGfpTerGrading,
QDevicePbadGfpTbsTerGrading,
QDevicePbadRbsAraTerGrading,
]
def getQuestionsGradingSubset(allQuestions, questionsSubset, grading):
result = []
for i in range(len(questionsSubset)):
result += [allQuestions.index(questionsSubset[i])]
return [grading[i] for i in result]
```
| github_jupyter |
# Bank Loan Approval Prediction using Artificial Neaural Network
In this project, we will build and train a deep neaural network model to predict the likelyhood of a liability customer buying personal loans based on customer features.
```
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras.layers import Dense, Activation, Dropout
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.metrics import Accuracy
import matplotlib.pyplot as plt
bank_df = pd.read_csv("UniversalBank.csv")
bank_df.head()
bank_df.shape
```
- ID: Customer ID
- Age: Customer Age
- Experience: Amount of work experience in years
- Income: Amount of annual income (in thousands)
- Zipcode: Zipcode of where customer lives
- Family: Number of family members
- CCAvg: Average monthly credit card spendings
- Education: Education level (1: Bachelor, 2: Master, 3: Advanced Degree)
- Mortgage: Mortgage of house (in thousands)
- Securities Account: Boolean of whether customer has a securities account
- CD Account: Boolean of whether customer has Certificate of Deposit account
- Online: Boolean of whether customer uses online banking
- CreditCard: Does the customer use credit card issued by the bank?
- Personal Loan: This is the target variable (Binary Classification Problem)
## Exploratory Data Analysis
```
bank_df.info()
bank_df.describe().transpose()
bank_df.isnull().sum()
```
Great, we have no missing values!
```
avg_age = bank_df["Age"].mean()
print ("The average age of this dataset is {:.1f}.".format(avg_age))
percent_cc = sum(bank_df["CreditCard"] == 1)/len(bank_df)
print ("The percentage of customers that own the bank's credit card is {:.2%}.".format(percent_cc))
percent_loan = sum(bank_df["Personal Loan"] == 1)/len(bank_df)
print ("The percentage of customers that took out a personal loan is {:.2%}.".format(percent_loan))
```
## Data Visualization
```
sns.countplot(x=bank_df["Personal Loan"])
plt.show()
sns.countplot(x=bank_df["Education"])
plt.show()
sns.countplot(x=bank_df["CreditCard"])
plt.show()
plt.figure(figsize=(20,10))
sns.countplot(x=bank_df["Age"])
plt.savefig('age.png', facecolor='w', bbox_inches='tight')
plt.show()
# lets look at the distribution of the income
plt.figure(figsize=(15,8))
sns.distplot(bank_df["Income"])
plt.savefig('income.png', facecolor='w', bbox_inches='tight')
plt.show()
# lets create 2 dataframes: one with personal loans and one without personal loans
personal_loans = bank_df[bank_df['Personal Loan'] == 1].copy()
no_personal_loans = bank_df[bank_df['Personal Loan'] == 0].copy()
personal_loans.describe().T
no_personal_loans.describe().T
plt.figure(figsize=(15,8))
sns.distplot(personal_loans["Income"], label='Approved')
sns.distplot(no_personal_loans["Income"], label='Not Approved')
plt.legend()
plt.savefig('approved_not_approved.png', facecolor='w', bbox_inches='tight')
plt.show()
cm = bank_df.corr()
plt.figure(figsize=(20,20))
sns.heatmap(cm, annot=True)
plt.savefig('heatmap.png', facecolor='w', bbox_inches='tight')
plt.show()
# lets look at the distribution of average credit card spending
plt.figure(figsize=(15,8))
sns.distplot(bank_df["CCAvg"])
plt.show()
plt.figure(figsize=(15,8))
sns.distplot(personal_loans["CCAvg"])
sns.distplot(no_personal_loans["CCAvg"])
plt.show()
```
## Data Preparation
```
from tensorflow.keras.utils import to_categorical
X = bank_df.drop(columns=["Personal Loan"])
y = bank_df["Personal Loan"]
y = to_categorical(y)
from sklearn.preprocessing import StandardScaler, MinMaxScaler
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.1)
sc = StandardScaler()
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
X_train.shape, X_test.shape, y_train.shape, y_test.shape
```
## Building a multi-layer neaural network model
```
# sequential model
ann_model = keras.Sequential()
# adding dense layer
ann_model.add(Dense(250, input_dim=13, kernel_initializer='normal', activation='relu'))
ann_model.add(Dropout(0.3))
ann_model.add(Dense(500, activation='relu'))
ann_model.add(Dropout(0.3))
ann_model.add(Dense(500, activation='relu'))
ann_model.add(Dropout(0.3))
ann_model.add(Dense(500, activation='relu'))
ann_model.add(Dropout(0.4))
ann_model.add(Dense(250, activation='linear'))
ann_model.add(Dropout(0.4))
# adding dense layer with softmax activation/output layer
ann_model.add(Dense(2, activation='softmax'))
ann_model.summary()
```
## Compilation and training of deep learning model
```
# custom functions for f1, precision and recall
from keras import backend as K
def recall_m(y_true, y_pred):
true_positives = K.sum(K.round(K.clip(y_true * y_pred, 0, 1)))
possible_positives = K.sum(K.round(K.clip(y_true, 0, 1)))
recall = true_positives / (possible_positives + K.epsilon())
return recall
def precision_m(y_true, y_pred):
true_positives = K.sum(K.round(K.clip(y_true * y_pred, 0, 1)))
predicted_positives = K.sum(K.round(K.clip(y_pred, 0, 1)))
precision = true_positives / (predicted_positives + K.epsilon())
return precision
def f1_m(y_true, y_pred):
precision = precision_m(y_true, y_pred)
recall = recall_m(y_true, y_pred)
return 2*((precision*recall)/(precision+recall+K.epsilon()))
ann_model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=[f1_m]) # metrics=['accuracy']
history = ann_model.fit(X_train, y_train, epochs=20, validation_split=0.2, verbose=1)
# Plot the model performance across epochs
plt.figure(figsize=(15,8))
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('Model Loss')
plt.ylabel('loss')
plt.legend(['train_loss','val_loss'], loc = 'upper right')
plt.savefig('modelloss.png', facecolor='w', bbox_inches='tight')
plt.show()
```
## Evaluating model performance
```
predictions = ann_model.predict(X_test)
predict = []
for i in predictions:
predict.append(np.argmax(i))
from sklearn import metrics
y_test = np.argmax(y_test, axis=1)
f1_test = metrics.f1_score(y_test, predict)
prec = metrics.precision_score(y_test, predict)
rec = metrics.recall_score(y_test, predict)
acc = metrics.accuracy_score(y_test, predict)
print ("F1 Score: {:.4f}.".format(f1_test))
print ("Precision: {:.4f}.".format(prec))
print ("Recall: {:.4f}.".format(rec))
print ("Accuracy: {:.4f}.".format(acc)) # note this is not a good measure of performance for this project as dataset is unbalanced.
conf_mat = metrics.confusion_matrix(y_test, predict)
plt.figure(figsize=(10,8))
sns.heatmap(conf_mat, annot=True, cbar=False)
plt.savefig('conf_matrix.png', facecolor='w', bbox_inches='tight')
plt.show()
print(metrics.classification_report(y_test, predict))
```
| github_jupyter |
# Mask R-CNN Demo
A quick intro to using the pre-trained model to detect and segment objects.
```
import os
import sys
import random
import math
import numpy as np
import skimage.io
import matplotlib
import matplotlib.pyplot as plt
import tensorflow as tf
# Root directory of the project
ROOT_DIR = os.path.abspath("../")
# Import Mask RCNN
sys.path.append(ROOT_DIR) # To find local version of the library
from mrcnn import utils
import mrcnn.model as modellib
from mrcnn import visualize
# Import COCO config
sys.path.append(os.path.join(ROOT_DIR, "samples/coco/")) # To find local version
import coco
%matplotlib inline
# Directory to save logs and trained model
MODEL_DIR = os.path.join(ROOT_DIR, "logs")
# Local path to trained weights file
COCO_MODEL_PATH = os.path.join(ROOT_DIR, "mask_rcnn_coco.h5")
# Download COCO trained weights from Releases if needed
if not os.path.exists(COCO_MODEL_PATH):
utils.download_trained_weights(COCO_MODEL_PATH)
# Directory of images to run detection on
IMAGE_DIR = os.path.join(ROOT_DIR, "images")
```
## Configurations
We'll be using a model trained on the MS-COCO dataset. The configurations of this model are in the ```CocoConfig``` class in ```coco.py```.
For inferencing, modify the configurations a bit to fit the task. To do so, sub-class the ```CocoConfig``` class and override the attributes you need to change.
```
class InferenceConfig(coco.CocoConfig):
# Set batch size to 1 since we'll be running inference on
# one image at a time. Batch size = GPU_COUNT * IMAGES_PER_GPU
GPU_COUNT = 1
IMAGES_PER_GPU = 1
config = InferenceConfig()
config.display()
```
## Create Model and Load Trained Weights
```
# added for tensorboard
sess = tf.Session()
#
# Create model object in inference mode.
model = modellib.MaskRCNN(mode="inference", model_dir=MODEL_DIR, config=config)
# Load weights trained on MS-COCO
model.load_weights(COCO_MODEL_PATH, by_name=True)
# added for tensorboard
tf.summary.FileWriter("./temp_graph", sess.graph)
```
## Class Names
The model classifies objects and returns class IDs, which are integer value that identify each class. Some datasets assign integer values to their classes and some don't. For example, in the MS-COCO dataset, the 'person' class is 1 and 'teddy bear' is 88. The IDs are often sequential, but not always. The COCO dataset, for example, has classes associated with class IDs 70 and 72, but not 71.
To improve consistency, and to support training on data from multiple sources at the same time, our ```Dataset``` class assigns it's own sequential integer IDs to each class. For example, if you load the COCO dataset using our ```Dataset``` class, the 'person' class would get class ID = 1 (just like COCO) and the 'teddy bear' class is 78 (different from COCO). Keep that in mind when mapping class IDs to class names.
To get the list of class names, you'd load the dataset and then use the ```class_names``` property like this.
```
# Load COCO dataset
dataset = coco.CocoDataset()
dataset.load_coco(COCO_DIR, "train")
dataset.prepare()
# Print class names
print(dataset.class_names)
```
We don't want to require you to download the COCO dataset just to run this demo, so we're including the list of class names below. The index of the class name in the list represent its ID (first class is 0, second is 1, third is 2, ...etc.)
```
# COCO Class names
# Index of the class in the list is its ID. For example, to get ID of
# the teddy bear class, use: class_names.index('teddy bear')
class_names = ['BG', 'person', 'bicycle', 'car', 'motorcycle', 'airplane',
'bus', 'train', 'truck', 'boat', 'traffic light',
'fire hydrant', 'stop sign', 'parking meter', 'bench', 'bird',
'cat', 'dog', 'horse', 'sheep', 'cow', 'elephant', 'bear',
'zebra', 'giraffe', 'backpack', 'umbrella', 'handbag', 'tie',
'suitcase', 'frisbee', 'skis', 'snowboard', 'sports ball',
'kite', 'baseball bat', 'baseball glove', 'skateboard',
'surfboard', 'tennis racket', 'bottle', 'wine glass', 'cup',
'fork', 'knife', 'spoon', 'bowl', 'banana', 'apple',
'sandwich', 'orange', 'broccoli', 'carrot', 'hot dog', 'pizza',
'donut', 'cake', 'chair', 'couch', 'potted plant', 'bed',
'dining table', 'toilet', 'tv', 'laptop', 'mouse', 'remote',
'keyboard', 'cell phone', 'microwave', 'oven', 'toaster',
'sink', 'refrigerator', 'book', 'clock', 'vase', 'scissors',
'teddy bear', 'hair drier', 'toothbrush']
```
## Run Object Detection
```
# Load a random image from the images folder
file_names = next(os.walk(IMAGE_DIR))[2]
# image = skimage.io.imread(os.path.join(IMAGE_DIR, random.choice(file_names)))
image = skimage.io.imread('/Users/AsherYartsev/Desktop/daveSnow.jpg')
# Run detection
results = model.detect([image], verbose=1)
# Visualize results
r = results[0]
visualize.display_instances(image, r['rois'], r['masks'], r['class_ids'], class_names, r['scores'])
```
| github_jupyter |
# CHAPTER 5 Getting Started with pandas
这一节终于要开始讲pandas了。闲话不说,直接开始正题。之后的笔记里,这样导入pandas:
```
import pandas as pd
```
另外可以导入Series和DataFrame,因为这两个经常被用到:
```
from pandas import Series, DataFrame
```
# 5.1 Introduction to pandas Data Structures
数据结构其实就是Series和DataFrame。
# 1 Series
这里series我就不翻译成序列了,因为之前的所有笔记里,我都是把sequence翻译成序列的。
series是一个像数组一样的一维序列,并伴有一个数组表示label,叫做index。创建一个series的方法也很简单:
```
obj = pd.Series([4, 7, -5, 3])
obj
```
可以看到,左边表示index,右边表示对应的value。可以通过value和index属性查看:
```
obj.values
obj.index # like range(4)
```
当然我们也可以自己指定index的label:
```
obj2 = pd.Series([4, 7, -5, 3], index=['d', 'b', 'a', 'c'])
obj2
obj2.index
```
可以用index的label来选择:
```
obj2['a']
obj2['d'] = 6
obj2[['c', 'a', 'd']]
```
这里['c', 'a', 'd']其实被当做了索引,尽管这个索引是用string构成的。
使用numpy函数或类似的操作,会保留index-value的关系:
```
obj2[obj2 > 0]
obj2 * 2
import numpy as np
np.exp(obj2)
```
另一种看待series的方法,它是一个长度固定,有顺序的dict,从index映射到value。在很多场景下,可以当做dict来用:
```
'b' in obj2
'e' in obj2
```
还可以直接用现有的dict来创建series:
```
sdata = {'Ohio': 35000, 'Texas': 71000, 'Oregon':16000, 'Utah': 5000}
obj3 = pd.Series(sdata)
obj3
```
series中的index其实就是dict中排好序的keys。我们也可以传入一个自己想要的顺序:
```
states = ['California', 'Ohio', 'Oregon', 'Texas']
obj4 = pd.Series(sdata, index=states)
obj4
```
顺序是按states里来的,但因为没有找到california,所以是NaN。NaN表示缺失数据,用之后我们提到的话就用missing或NA来指代。pandas中的isnull和notnull函数可以用来检测缺失数据:
```
pd.isnull(obj4)
pd.notnull(obj4)
```
series也有对应的方法:
```
obj4.isnull()
```
关于缺失数据,在第七章还会讲得更详细一些。
series中一个有用的特色自动按index label来排序(Data alignment features):
```
obj3
obj4
obj3 + obj4
```
这个Data alignment features(数据对齐特色)和数据库中的join相似。
serice自身和它的index都有一个叫name的属性,这个能和其他pandas的函数进行整合:
```
obj4.name = 'population'
obj4.index.name = 'state'
obj4
```
series的index能被直接更改:
```
obj
obj.index = ['Bob', 'Steve', 'Jeff', 'Ryan']
obj
```
# 2 DataFrame
DataFrame表示一个长方形表格,并包含排好序的列,每一列都可以是不同的数值类型(数字,字符串,布尔值)。DataFrame有行索引和列索引(row index, column index);可以看做是分享所有索引的由series组成的字典。数据是保存在一维以上的区块里的。
(其实我是把dataframe当做excel里的那种表格来用的,这样感觉更直观一些)
构建一个dataframe的方法,用一个dcit,dict里的值是list:
```
data = {'state': ['Ohio', 'Ohio', 'Ohio', 'Nevada', 'Nevada', 'Nevada'],
'year': [2000, 2001, 2002, 2001, 2002, 2003],
'pop': [1.5, 1.7, 3.6, 2.4, 2.9, 3.2]}
frame = pd.DataFrame(data)
frame
```
dataframe也会像series一样,自动给数据赋index, 而列则会按顺序排好。
对于一个较大的DataFrame,用head方法会返回前5行(注:这个函数在数据分析中经常使用,用来查看表格里有什么东西):
```
frame.head()
```
如果指定一列的话,会自动按列排序:
```
pd.DataFrame(data, columns=['year', 'state', 'pop'])
```
如果你导入一个不存在的列名,那么会显示为缺失数据:
```
frame2 = pd.DataFrame(data, columns=['year', 'state', 'pop', 'debt'],
index=['one', 'two', 'three', 'four', 'five', 'six'])
frame2
frame2.columns
```
从DataFrame里提取一列的话会返回series格式,可以以属性或是dict一样的形式来提取:
```
frame2['state']
frame2.year
```
注意:frame2[column]能应对任何列名,但frame2.column的情况下,列名必须是有效的python变量名才行。
返回的series有DataFrame种同样的index,而且name属性也是对应的。
对于行,要用在loc属性里用 位置或名字:
```
frame2.loc['three']
```
列值也能通过赋值改变。比如给debt赋值:
```
frame2['debt'] = 16.5
frame2
frame2['debt'] = np.arange(6.)
frame2
```
如果把list或array赋给column的话,长度必须符合DataFrame的长度。如果把一二series赋给DataFrame,会按DataFrame的index来赋值,不够的地方用缺失数据来表示:
```
val = pd.Series([-1.2, -1.5, -1.7], index=['two', 'four', 'five'])
frame2['debt'] = val
frame2
```
如果列不存在,赋值会创建一个新列。而del也能像删除字典关键字一样,删除列:
```
frame2['eastern'] = frame2.state == 'Ohio'
frame2
```
然后用del删除这一列:
```
del frame2['eastern']
frame2.columns
```
注意:columns返回的是一个view,而不是新建了一个copy。因此,任何对series的改变,会反映在DataFrame上。除非我们用copy方法来新建一个。
另一种常见的格式是dict中的dict:
```
pop = {'Nevada': {2001: 2.4, 2002: 2.9},
'Ohio': {2000: 1.5, 2001: 1.7, 2002: 3.6}}
```
把上面这种嵌套dcit传给DataFrame,pandas会把外层dcit的key当做列,内层key当做行索引:
```
frame3 = pd.DataFrame(pop)
frame3
```
另外DataFrame也可以向numpy数组一样做转置:
```
frame3.T
```
指定index:
```
pd.DataFrame(pop, index=[2001, 2002, 2003])
```
series组成的dict:
```
pdata = {'Ohio': frame3['Ohio'][:-1],
'Nevada': frame3['Nevada'][:2]}
pd.DataFrame(pdata)
```
其他一些可以传递给DataFrame的构造器:

如果DataFrame的index和column有自己的name属性,也会被显示:
```
frame3.index.name = 'year'; frame3.columns.name = 'state'
frame3
```
values属性会返回二维数组:
```
frame3.values
```
如果column有不同的类型,dtype会适应所有的列:
```
frame2.values
```
# 3 Index Objects (索引对象)
pandas的Index Objects (索引对象)负责保存axis labels和其他一些数据(比如axis name或names)。一个数组或其他一个序列标签,只要被用来做构建series或DataFrame,就会被自动转变为index:
```
obj = pd.Series(range(3), index=['a', 'b', 'c'])
index = obj.index
index
index[1:]
```
index object是不可更改的:
```
index[1] = 'd'
```
正因为不可修改,所以data structure中分享index object是很安全的:
```
labels = pd.Index(np.arange(3))
labels
obj2 = pd.Series([1.5, -2.5, 0], index=labels)
obj2
obj2.index is labels
```
index除了想数组,还能像大小一定的set:
```
frame3
frame3.columns
'Ohio' in frame3.columns
2003 in frame3.columns
```
与python里的set不同,pandas的index可以有重复的labels:
```
dup_labels = pd.Index(['foo', 'foo', 'bar', 'bar'])
dup_labels
```
在这种重复的标签中选择的话,会选中所有相同的标签。
Index还有一些方法和属性:

| github_jupyter |
# Grasping an object
For this laboratory, we will use two robots distant from each other of 30cm as depicted in the figure below
<img src="./2_robot_config.png" width="500">
There is 1cm between the end-effector "shell" and the position of the frame as depicted in the figure below
<img src="./foot_close.png" width="300">
The object to grasp is a 15cm wide, 10cm high box that weighs 500g.
<img src="./box.png" width="300">
The goal of the notebook is to find a controller and a sequence of movements that enable to grasp an object, lift it up 15cm and then put it on the ground 10cm closer to robot 2.
```
#setup nice plotting (use widget instead of notebook in the command below if you use jupyter lab)
%matplotlib notebook
# we import useful libraries
import time
import numpy as np
import matplotlib as mp
import matplotlib.pyplot as plt
use_real_robot_computer = False
use_real_robot = False
if use_real_robot_computer:
from nyu_finger import NYUFingerReal
from nyu_finger_sim.nyu_finger_simulator import NYUFingerSimulator
else:
from nyu_finger_simulator import NYUFingerSimulator
import robot_kinematics
# we create a robot simulation
if use_real_robot:
robot1 = NYUFingerReal()
robot1.initialize('enp5s0f1')
robot2 = NYUFingerReal()
robot2.initialize('eno1')
else:
robot1 = NYUFingerSimulator(robotStartPos = [0.,0.,0.])
robot1.add_box(friction=0.5)
# we put the 2nd robot 0.3m away
robot2 = NYUFingerSimulator(robotStartPos = [0.3,0.,0.])
# we reset the simulation to the initial position we want to move
robot1.reset_state(np.array([0.0,0.0,0.3]))
robot2.reset_state(np.array([0.0,0.0,-0.3]))
robot1.step()
robot1.step()
```
The code below implements a sequence of movements towards a list of setpoints (defined in `set_points_rob1` and `set_points_rob2` for each robot). These setpoints for now move the end-effectors towards the object and then move the end-effectors up and down.
The controller is an impedance controller (with stiffness K and damping D - set to fixed values for both robots). Additionally, gravity compensation for the mass of the robot is added to the joint torques.
$$\tau = J_O^T \Big( K (p_{desired} - p_{measured}) + D (\dot{p}_{desired} - \dot{p}_{measured} ) \Big) - g(\theta)$$
## Questions
1. The object weighs 0.5kg, the friction coefficient is $\mu=0.5$ and the stiffness K of the impedance controllers is fixed to $K=150$ in every direction. Additionally, there is 1cm between the finger coordinate frame and the shell of the finger. What distance "inside" the object should the endeffector setpoint be defined for each robot such that it will squeeze the object sufficiently to be able to lift it? (i.e. such that friction can exactly compensate for the mass of the object) Explain your answer.
2. Given your computations, modify the sequence of setpoints such that the robots can lift the object (give a 3mm margin when setting the setpoints inside the object to account for tracking errors and the assumption that the object is static). Plot the motion of the end-effector (desired vs. measured) as a function of time.
3. What happens if you squeeze 1cm more or 1cm less than computed? Is it expected? (plot the results)
4. We would like to additionally compensate for the effect on the robot of the weight of the object when it is lifted. Write a controller that compensates for the weight of the object on both robots when they attempt to lift it (carefull, each robot need to only compensate for half the mass of the robot). Compare its performance with the results of point 2. (remark: the number `current_set_point` can tell you which set_point goal is currently being used in order to know when to start compensating for the object weight).
5. Change the controller and/or sequence of set points in order to put the object down 10cm closer to robot 2. Plot the resulting motions of the end-effectors.
6. (optional) Try any/all of these controllers on the real robots. Save and plot the results.
```
# some parameters for the trajectory generation
traj_time = 0.0 # the timing for each trajectory
current_set_point = 1 #the current goal position
trajectory_duration = 2. #each trajectory is 2 seconds long
# the list of setpoints for the first robot end-effector
set_points_rob1 = [
np.array([0.3,-0.25,0.014]), # the first one does not count
np.array([0.3,-0.25,0.014]),
np.array([0.3+0.065,-0.25,0.014]),
np.array([0.3+0.065,-0.1,0.014]),
np.array([0.3+0.065,-0.25,0.014]),
np.array([0.3,-0.25,0.014]),
]
# the list of setpoints for the second robot end-effector
set_points_rob2 = [
np.array([0.3,-0.25,0.014]), # the first one does not count
np.array([0.3,-0.25,0.014]),
np.array([0.3-0.065,-0.25,0.014]),
np.array([0.3-0.065,-0.1,0.014]),
np.array([0.3-0.065,-0.25,0.014]),
np.array([0.3,-0.25,0.014]),
]
# we simulate until we go through all the setpoints
run_time = (len(set_points_rob1)-1)*trajectory_duration
dt = 0.001
num_steps = int(run_time/dt)
# we create one kinematic object per robot
kin_robot1 = robot_kinematics.robot_kinematics()
kin_robot2 = robot_kinematics.robot_kinematics()
# we create arrays to store information
simulation_time = np.zeros([num_steps]) # will store the running time
# data from the first robot
measured_joint_positions_robot1 = np.zeros([num_steps,3]) # will store the measured position
measured_joint_velocities_robot1 = np.zeros_like(measured_joint_positions_robot1) # will store the measure velocities
desired_torques_robot1 = np.zeros_like(measured_joint_positions_robot1) # will store the commands we send to the robot
foot_measured_position_robot1 = np.zeros([num_steps,3])
foot_measured_velocity_robot1 = np.zeros([num_steps,3])
foot_desired_position_robot1 = np.zeros([num_steps,3])
foot_desired_velocity_robot1 = np.zeros([num_steps,3])
# data from the second robot
measured_joint_positions_robot2 = np.zeros([num_steps,3]) # will store the measured position
measured_joint_velocities_robot2 = np.zeros_like(measured_joint_positions_robot2) # will store the measure velocities
desired_torques_robot2 = np.zeros_like(measured_joint_positions_robot2) # will store the commands we send to the robot
foot_measured_position_robot2 = np.zeros([num_steps,3])
foot_measured_velocity_robot2 = np.zeros([num_steps,3])
foot_desired_position_robot2 = np.zeros([num_steps,3])
foot_desired_velocity_robot2 = np.zeros([num_steps,3])
# now we can enter the main control loop (each loop is 1 control cycle)
for i in range(num_steps):
# get the current time and store it
simulation_time[i] = dt * i
####SENSOR PROCESSING###
# we get the position and velocities of the joints for robot1
q_rob1, dq_rob1 = robot1.get_state()
measured_joint_positions_robot1[i,:] = q_rob1
measured_joint_velocities_robot1[i,:] = dq_rob1
# we get the position and velocities of the joints for robot2
q_rob2, dq_rob2 = robot2.get_state()
measured_joint_positions_robot2[i,:] = q_rob2
measured_joint_velocities_robot2[i,:] = dq_rob2
########################
#### KINEMATICS FUNCTIONS ####
# compute FK, get Jacobian and compute end-effector velocity for robot1
kin_robot1.update_kinematics(q_rob1)
foot_measured_position_robot1[i,:] = kin_robot1.TS_F[0:3,3]
jacobian_rob1 = kin_robot1.orientedJ[3:6,:]
foot_measured_velocity_robot1[i,:] = jacobian_rob1.dot(dq_rob1)
# compute FK, get Jacobian and compute end-effector velocity for robot2
kin_robot2.update_kinematics(q_rob2)
foot_measured_position_robot2[i,:] = kin_robot2.TS_F[0:3,3]
jacobian_rob2 = kin_robot2.orientedJ[3:6,:]
foot_measured_velocity_robot2[i,:] = jacobian_rob2.dot(dq_rob2)
########################
# get the gravity term for each robot (will be used for gravity compensation)
g_rob1 = kin_robot1.getG()
g_rob2 = kin_robot2.getG()
### TRAJECTORY GENERATION ###
# here we compute the desired position and velocity of both end-effectors to go from setpoints to setpoints
# we use the trajectory generator designed in Lab 3
# the function is stored in the package robot_kinematics.py
for f in range(3):
foot_desired_position_robot1[i,f], foot_desired_velocity_robot1[i,f] = robot_kinematics.compute_trajectory(set_points_rob1[current_set_point-1][f], set_points_rob1[current_set_point][f], trajectory_duration, traj_time)
foot_desired_position_robot2[i,f], foot_desired_velocity_robot2[i,f] = robot_kinematics.compute_trajectory(set_points_rob2[current_set_point-1][f], set_points_rob2[current_set_point][f], trajectory_duration, traj_time)
# here we detect if we need to switch to the next setpoint
# or if we can just increment the trajectory timing
if traj_time < trajectory_duration:
traj_time += dt
elif current_set_point < len(set_points_rob1)-1:
traj_time = 0.
current_set_point += 1
########################
### CONTROLLER ###
# we use the same K and D gains for both robots
K = np.diag([150,150,150])
D = np.diag([5,5,5])
# compute the desired end-effector force for robot 1
des_force_rob1 = K.dot(foot_desired_position_robot1[i,:] - foot_measured_position_robot1[i,:])
des_force_rob1 += D.dot(foot_desired_velocity_robot1[i,:] - foot_measured_velocity_robot1[i,:])
# impedance controller with gravity compensation for the joints
desired_torques_robot1[i,:] = (jacobian_rob1.T).dot(des_force_rob1) - g_rob1
# we send the torques to robot 1
robot1.send_joint_torque(desired_torques_robot1[i,:])
# compute the desired end-effector force for robot 2
des_force_rob2 = K.dot(foot_desired_position_robot2[i,:] - foot_measured_position_robot2[i,:])
des_force_rob2 += D.dot(foot_desired_velocity_robot2[i,:] - foot_measured_velocity_robot2[i,:])
# impedance controller with gravity compensation for the joints
desired_torques_robot2[i,:] = (jacobian_rob2.T).dot(des_force_rob2) - g_rob2
# we send the torques to robot 2
robot2.send_joint_torque(desired_torques_robot2[i,:])
########################
# we call step to do one simulation step (only needs to be called once from one robot)
robot1.step()
def plot_foot_trajectory(time, foot_pos, foot_pos_des):
"""
plot the time evolution of the end-effector in 3D vs. the desired trajectories
"""
x_pos = foot_pos[:,0]
y_pos = foot_pos[:,1]
z_pos = foot_pos[:,2]
plt.figure()
plt.subplot(3,1,1)
plt.plot(time, x_pos, time, foot_pos_des[:,0])
plt.ylabel('foot x position [m]')
plt.subplot(3,1,2)
plt.plot(time, y_pos, time, foot_pos_des[:,1])
plt.ylabel('foot y position [m]')
plt.subplot(3,1,3)
plt.plot(time, z_pos, time, foot_pos_des[:,2])
plt.ylabel('foot z position [m]')
plt.xlabel('Time [s]')
plot_foot_trajectory(simulation_time, foot_measured_position_robot1, foot_desired_position_robot1)
plot_foot_trajectory(simulation_time, foot_measured_position_robot2, foot_desired_position_robot2)
```
| github_jupyter |
Bibliography:
* [Stacked Convolutional Auto-Encoders for Hierarchical Feature Extraction](http://people.idsia.ch/~ciresan/data/icann2011.pdf)
```
import torch
import torchvision
from torch import nn, optim
from torch.nn import functional as F
from torch.autograd import Variable
from torch.utils.data import DataLoader, Dataset
from torchvision import transforms, utils
from torchvision import datasets
from torchvision.utils import save_image
import skimage
import math
# import io
# import requests
# from PIL import Image
import numpy as np
import pandas as pd
# import matplotlib.pyplot as plt
import sys
import os
import cae
from helpers import *
from helper_modules import *
from multi_res_cae import *
%load_ext autoreload
%autoreload
%aimport helpers, helper_modules, multi_res_cae
%aimport
# frcae = MultiFullCAE((640,480))
# mrcae = MultiResCAE([640,480])
# Hyper Parameters
# num_epochs = 5
# batch_size = 100
# learning_rate = 0.001
num_epochs = 20
batch_size = 128
learning_rate = 0.0001
%%time
model = MultiFullCAE(in_img_shape=(32,32), channels=1, full_image_resize=(24,24)).cuda()
# model = MultiResCAE(in_img_shape=[32,32], channels=3, conv_layer_feat=[16, 32, 64],
# res_px=[[24, 24], [16, 16], [12, 12]], crop_sizes=[[32, 32], [24,24], [12, 12]],
# # conv_sizes = [(3,5,7), (3,5,7,11), (3,5,7,11)] # this is too much I think
# # conv_sizes=[[1, 3, 5], [1, 3, 5], [1, 3, 5, 7]] # test b
# conv_sizes=[[5, 7, 11], [3, 5, 7, 9], [1, 3, 5]] # test c
# ).cuda()
# model.parameters
%%time
criterion = nn.MSELoss()
#criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.Adam(model.parameters(), lr=learning_rate, weight_decay=1e-5)
def to_img(x):
x = 0.5 * (x + 1)
x = x.clamp(0, 1)
x = x.view(x.size(0), 1, 32, 32)
return x
%%time
transformation = monochrome_preprocess(32,32)
#transformation = fullimage_preprocess(32,32)
#train_loader, test_loader = get_loaders(batch_size, transformation, dataset=datasets.CocoDetection)
train_loader, test_loader = get_loaders(batch_size, transformation)
%%time
for epoch in range(num_epochs):
for i, (img, labels) in enumerate(train_loader):
img = Variable(img).cuda()
# ===================forward=====================
# print("encoding batch of images")
output = model(img)
# print("computing loss")
loss = criterion(output, img)
# ===================backward====================
# print("Backward ")
optimizer.zero_grad()
loss.backward()
optimizer.step()
# ===================log========================
print('epoch [{}/{}], loss:{:.4f}'.format(epoch+1, num_epochs, loss.data))
if epoch % 4 == 0:
pic = to_img(output.cpu().data)
in_pic = to_img(img.cpu().data)
save_image(pic, './fmrcae_results/c_MergingLayer_in-32x32_3-5-7-11_out_image_{}.png'.format(epoch))
save_image(in_pic, './fmrcae_results/c_MergingLayer_in-3-5-7-11_in_image_{}.png'.format(epoch))
# if loss.data[0] < 0.21: #arbitrary number because I saw that it works well enough
# break
#torch.save("fmrcae_in-64x64_32x32_3-5-7-11.pth", model)
#torch.save("mrcae_in-32x32_.pth", model)
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.