text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
```
#default_exp foundation
#export
from fastcore.imports import *
from fastcore.basics import *
from functools import lru_cache
from contextlib import contextmanager
from copy import copy
from configparser import ConfigParser
import random,pickle,inspect
from fastcore.test import *
from nbdev.showdoc import *
from fastcore.nb_imports import *
```
# Foundation
> The `L` class and helpers for it
## Foundational Functions
```
#export
@contextmanager
def working_directory(path):
"Change working directory to `path` and return to previous on exit."
prev_cwd = Path.cwd()
os.chdir(path)
try: yield
finally: os.chdir(prev_cwd)
#export
def add_docs(cls, cls_doc=None, **docs):
"Copy values from `docs` to `cls` docstrings, and confirm all public methods are documented"
if cls_doc is not None: cls.__doc__ = cls_doc
for k,v in docs.items():
f = getattr(cls,k)
if hasattr(f,'__func__'): f = f.__func__ # required for class methods
f.__doc__ = v
# List of public callables without docstring
nodoc = [c for n,c in vars(cls).items() if callable(c)
and not n.startswith('_') and c.__doc__ is None]
assert not nodoc, f"Missing docs: {nodoc}"
assert cls.__doc__ is not None, f"Missing class docs: {cls}"
```
`add_docs` allows you to add docstrings to a class and its associated methods. This function allows you to group docstrings together seperate from your code, which enables you to define one-line functions as well as organize your code more succintly. We believe this confers a number of benefits which we discuss in [our style guide](https://docs.fast.ai/dev/style.html).
Suppose you have the following undocumented class:
```
class T:
def foo(self): pass
def bar(self): pass
```
You can add documentation to this class like so:
```
add_docs(T, cls_doc="A docstring for the class.",
foo="The foo method.",
bar="The bar method.")
```
Now, docstrings will appear as expected:
```
test_eq(T.__doc__, "A docstring for the class.")
test_eq(T.foo.__doc__, "The foo method.")
test_eq(T.bar.__doc__, "The bar method.")
```
`add_docs` also validates that all of your public methods contain a docstring. If one of your methods is not documented, it will raise an error:
```
class T:
def foo(self): pass
def bar(self): pass
f=lambda: add_docs(T, "A docstring for the class.", foo="The foo method.")
test_fail(f, contains="Missing docs")
#hide
class _T:
def f(self): pass
@classmethod
def g(cls): pass
add_docs(_T, "a", f="f", g="g")
test_eq(_T.__doc__, "a")
test_eq(_T.f.__doc__, "f")
test_eq(_T.g.__doc__, "g")
#export
def docs(cls):
"Decorator version of `add_docs`, using `_docs` dict"
add_docs(cls, **cls._docs)
return cls
```
Instead of using `add_docs`, you can use the decorator `docs` as shown below. Note that the docstring for the class can be set with the argument `cls_doc`:
```
@docs
class _T:
def f(self): pass
def g(cls): pass
_docs = dict(cls_doc="The class docstring",
f="The docstring for method f.",
g="A different docstring for method g.")
test_eq(_T.__doc__, "The class docstring")
test_eq(_T.f.__doc__, "The docstring for method f.")
test_eq(_T.g.__doc__, "A different docstring for method g.")
```
For either the `docs` decorator or the `add_docs` function, you can still define your docstrings in the normal way. Below we set the docstring for the class as usual, but define the method docstrings through the `_docs` attribute:
```
@docs
class _T:
"The class docstring"
def f(self): pass
_docs = dict(f="The docstring for method f.")
test_eq(_T.__doc__, "The class docstring")
test_eq(_T.f.__doc__, "The docstring for method f.")
show_doc(is_iter)
assert is_iter([1])
assert not is_iter(array(1))
assert is_iter(array([1,2]))
assert (o for o in range(3))
# export
def coll_repr(c, max_n=10):
"String repr of up to `max_n` items of (possibly lazy) collection `c`"
return f'(#{len(c)}) [' + ','.join(itertools.islice(map(repr,c), max_n)) + (
'...' if len(c)>max_n else '') + ']'
```
`coll_repr` is used to provide a more informative [`__repr__`](https://stackoverflow.com/questions/1984162/purpose-of-pythons-repr) about list-like objects. `coll_repr` and is used by `L` to build a `__repr__` that displays the length of a list in addition to a preview of a list.
Below is an example of the `__repr__` string created for a list of 1000 elements:
```
test_eq(coll_repr(range(1000)), '(#1000) [0,1,2,3,4,5,6,7,8,9...]')
test_eq(coll_repr(range(1000), 5), '(#1000) [0,1,2,3,4...]')
test_eq(coll_repr(range(10), 5), '(#10) [0,1,2,3,4...]')
test_eq(coll_repr(range(5), 5), '(#5) [0,1,2,3,4]')
```
We can set the option `max_n` to optionally preview a specified number of items instead of the default:
```
test_eq(coll_repr(range(1000), max_n=5), '(#1000) [0,1,2,3,4...]')
# export
def is_bool(x):
"Check whether `x` is a bool or None"
return isinstance(x,(bool,NoneType)) or risinstance('bool_', x)
# export
def mask2idxs(mask):
"Convert bool mask or index list to index `L`"
if isinstance(mask,slice): return mask
mask = list(mask)
if len(mask)==0: return []
it = mask[0]
if hasattr(it,'item'): it = it.item()
if is_bool(it): return [i for i,m in enumerate(mask) if m]
return [int(i) for i in mask]
test_eq(mask2idxs([False,True,False,True]), [1,3])
test_eq(mask2idxs(array([False,True,False,True])), [1,3])
test_eq(mask2idxs(array([1,2,3])), [1,2,3])
#export
def cycle(o):
"Like `itertools.cycle` except creates list of `None`s if `o` is empty"
o = listify(o)
return itertools.cycle(o) if o is not None and len(o) > 0 else itertools.cycle([None])
test_eq(itertools.islice(cycle([1,2,3]),5), [1,2,3,1,2])
test_eq(itertools.islice(cycle([]),3), [None]*3)
test_eq(itertools.islice(cycle(None),3), [None]*3)
test_eq(itertools.islice(cycle(1),3), [1,1,1])
#export
def zip_cycle(x, *args):
"Like `itertools.zip_longest` but `cycle`s through elements of all but first argument"
return zip(x, *map(cycle,args))
test_eq(zip_cycle([1,2,3,4],list('abc')), [(1, 'a'), (2, 'b'), (3, 'c'), (4, 'a')])
#export
def is_indexer(idx):
"Test whether `idx` will index a single item in a list"
return isinstance(idx,int) or not getattr(idx,'ndim',1)
```
You can, for example index a single item in a list with an integer or a 0-dimensional numpy array:
```
assert is_indexer(1)
assert is_indexer(np.array(1))
```
However, you cannot index into single item in a list with another list or a numpy array with ndim > 0.
```
assert not is_indexer([1, 2])
assert not is_indexer(np.array([[1, 2], [3, 4]]))
```
## `L` helpers
```
#export
class CollBase:
"Base class for composing a list of `items`"
def __init__(self, items): self.items = items
def __len__(self): return len(self.items)
def __getitem__(self, k): return self.items[list(k) if isinstance(k,CollBase) else k]
def __setitem__(self, k, v): self.items[list(k) if isinstance(k,CollBase) else k] = v
def __delitem__(self, i): del(self.items[i])
def __repr__(self): return self.items.__repr__()
def __iter__(self): return self.items.__iter__()
```
`ColBase` is a base class that emulates the functionality of a python `list`:
```
class _T(CollBase): pass
l = _T([1,2,3,4,5])
test_eq(len(l), 5) # __len__
test_eq(l[-1], 5); test_eq(l[0], 1) #__getitem__
l[2] = 100; test_eq(l[2], 100) # __set_item__
del l[0]; test_eq(len(l), 4) # __delitem__
test_eq(str(l), '[2, 100, 4, 5]') # __repr__
```
## L -
```
#export
class _L_Meta(type):
def __call__(cls, x=None, *args, **kwargs):
if not args and not kwargs and x is not None and isinstance(x,cls): return x
return super().__call__(x, *args, **kwargs)
#export
class L(GetAttr, CollBase, metaclass=_L_Meta):
"Behaves like a list of `items` but can also index with list of indices or masks"
_default='items'
def __init__(self, items=None, *rest, use_list=False, match=None):
if (use_list is not None) or not is_array(items):
items = listify(items, *rest, use_list=use_list, match=match)
super().__init__(items)
@property
def _xtra(self): return None
def _new(self, items, *args, **kwargs): return type(self)(items, *args, use_list=None, **kwargs)
def __getitem__(self, idx): return self._get(idx) if is_indexer(idx) else L(self._get(idx), use_list=None)
def copy(self): return self._new(self.items.copy())
def _get(self, i):
if is_indexer(i) or isinstance(i,slice): return getattr(self.items,'iloc',self.items)[i]
i = mask2idxs(i)
return (self.items.iloc[list(i)] if hasattr(self.items,'iloc')
else self.items.__array__()[(i,)] if hasattr(self.items,'__array__')
else [self.items[i_] for i_ in i])
def __setitem__(self, idx, o):
"Set `idx` (can be list of indices, or mask, or int) items to `o` (which is broadcast if not iterable)"
if isinstance(idx, int): self.items[idx] = o
else:
idx = idx if isinstance(idx,L) else listify(idx)
if not is_iter(o): o = [o]*len(idx)
for i,o_ in zip(idx,o): self.items[i] = o_
def __eq__(self,b):
if risinstance('ndarray', b): return array_equal(b, self)
if isinstance(b, (str,dict)): return False
return all_equal(b,self)
def sorted(self, key=None, reverse=False): return self._new(sorted_ex(self, key=key, reverse=reverse))
def __iter__(self): return iter(self.items.itertuples() if hasattr(self.items,'iloc') else self.items)
def __contains__(self,b): return b in self.items
def __reversed__(self): return self._new(reversed(self.items))
def __invert__(self): return self._new(not i for i in self)
def __repr__(self): return repr(self.items)
def _repr_pretty_(self, p, cycle):
p.text('...' if cycle else repr(self.items) if is_array(self.items) else coll_repr(self))
def __mul__ (a,b): return a._new(a.items*b)
def __add__ (a,b): return a._new(a.items+listify(b))
def __radd__(a,b): return a._new(b)+a
def __addi__(a,b):
a.items += list(b)
return a
@classmethod
def split(cls, s, sep=None, maxsplit=-1): return cls(s.split(sep,maxsplit))
@classmethod
def range(cls, a, b=None, step=None): return cls(range_of(a, b=b, step=step))
def map(self, f, *args, gen=False, **kwargs): return self._new(map_ex(self, f, *args, gen=gen, **kwargs))
def argwhere(self, f, negate=False, **kwargs): return self._new(argwhere(self, f, negate, **kwargs))
def filter(self, f=noop, negate=False, gen=False, **kwargs):
return self._new(filter_ex(self, f=f, negate=negate, gen=gen, **kwargs))
def enumerate(self): return L(enumerate(self))
def renumerate(self): return L(renumerate(self))
def unique(self, sort=False, bidir=False, start=None): return L(uniqueify(self, sort=sort, bidir=bidir, start=start))
def val2idx(self): return val2idx(self)
def cycle(self): return cycle(self)
def map_dict(self, f=noop, *args, gen=False, **kwargs): return {k:f(k, *args,**kwargs) for k in self}
def map_first(self, f=noop, g=noop, *args, **kwargs):
return first(self.map(f, *args, gen=False, **kwargs), g)
def itemgot(self, *idxs):
x = self
for idx in idxs: x = x.map(itemgetter(idx))
return x
def attrgot(self, k, default=None):
return self.map(lambda o: o.get(k,default) if isinstance(o, dict) else nested_attr(o,k,default))
def starmap(self, f, *args, **kwargs): return self._new(itertools.starmap(partial(f,*args,**kwargs), self))
def zip(self, cycled=False): return self._new((zip_cycle if cycled else zip)(*self))
def zipwith(self, *rest, cycled=False): return self._new([self, *rest]).zip(cycled=cycled)
def map_zip(self, f, *args, cycled=False, **kwargs): return self.zip(cycled=cycled).starmap(f, *args, **kwargs)
def map_zipwith(self, f, *rest, cycled=False, **kwargs): return self.zipwith(*rest, cycled=cycled).starmap(f, **kwargs)
def shuffle(self):
it = copy(self.items)
random.shuffle(it)
return self._new(it)
def concat(self): return self._new(itertools.chain.from_iterable(self.map(L)))
def reduce(self, f, initial=None): return reduce(f, self) if initial is None else reduce(f, self, initial)
def sum(self): return self.reduce(operator.add)
def product(self): return self.reduce(operator.mul)
def setattrs(self, attr, val): [setattr(o,attr,val) for o in self]
#export
add_docs(L,
__getitem__="Retrieve `idx` (can be list of indices, or mask, or int) items",
range="Class Method: Same as `range`, but returns `L`. Can pass collection for `a`, to use `len(a)`",
split="Class Method: Same as `str.split`, but returns an `L`",
copy="Same as `list.copy`, but returns an `L`",
sorted="New `L` sorted by `key`. If key is str use `attrgetter`; if int use `itemgetter`",
unique="Unique items, in stable order",
val2idx="Dict from value to index",
filter="Create new `L` filtered by predicate `f`, passing `args` and `kwargs` to `f`",
argwhere="Like `filter`, but return indices for matching items",
map="Create new `L` with `f` applied to all `items`, passing `args` and `kwargs` to `f`",
map_first="First element of `map_filter`",
map_dict="Like `map`, but creates a dict from `items` to function results",
starmap="Like `map`, but use `itertools.starmap`",
itemgot="Create new `L` with item `idx` of all `items`",
attrgot="Create new `L` with attr `k` (or value `k` for dicts) of all `items`.",
cycle="Same as `itertools.cycle`",
enumerate="Same as `enumerate`",
renumerate="Same as `renumerate`",
zip="Create new `L` with `zip(*items)`",
zipwith="Create new `L` with `self` zip with each of `*rest`",
map_zip="Combine `zip` and `starmap`",
map_zipwith="Combine `zipwith` and `starmap`",
concat="Concatenate all elements of list",
shuffle="Same as `random.shuffle`, but not inplace",
reduce="Wrapper for `functools.reduce`",
sum="Sum of the items",
product="Product of the items",
setattrs="Call `setattr` on all items"
)
#export
#hide
# Here we are fixing the signature of L. What happens is that the __call__ method on the MetaClass of L shadows the __init__
# giving the wrong signature (https://stackoverflow.com/questions/49740290/call-from-metaclass-shadows-signature-of-init).
def _f(items=None, *rest, use_list=False, match=None): ...
L.__signature__ = inspect.signature(_f)
#export
Sequence.register(L);
```
`L` is a drop in replacement for a python `list`. Inspired by [NumPy](http://www.numpy.org/), `L`, supports advanced indexing and has additional methods (outlined below) that provide additional functionality and encourage simple expressive code. For example, the code below takes a list of pairs, selects the second item of each pair, takes its absolute value, filters items greater than 4, and adds them up:
```
from fastcore.utils import gt
d = dict(a=1,b=-5,d=6,e=9).items()
test_eq(L(d).itemgot(1).map(abs).filter(gt(4)).sum(), 20) # abs(-5) + abs(6) + abs(9) = 20; 1 was filtered out.
```
Read [this overview section](https://fastcore.fast.ai/#L) for a quick tutorial of `L`, as well as background on the name.
You can create an `L` from an existing iterable (e.g. a list, range, etc) and access or modify it with an int list/tuple index, mask, int, or slice. All `list` methods can also be used with `L`.
```
t = L(range(12))
test_eq(t, list(range(12)))
test_ne(t, list(range(11)))
t.reverse()
test_eq(t[0], 11)
t[3] = "h"
test_eq(t[3], "h")
t[3,5] = ("j","k")
test_eq(t[3,5], ["j","k"])
test_eq(t, L(t))
test_eq(L(L(1,2),[3,4]), ([1,2],[3,4]))
t
```
Any `L` is a `Sequence` so you can use it with methods like `random.sample`:
```
assert isinstance(t, Sequence)
import random
random.sample(t, 3)
#hide
# test set items with L of collections
x = L([[1,2,3], [4,5], [6,7]])
x[0] = [1,2]
test_eq(x, L([[1,2], [4,5], [6,7]]))
```
There are optimized indexers for arrays, tensors, and DataFrames.
```
arr = np.arange(9).reshape(3,3)
t = L(arr, use_list=None)
test_eq(t[1,2], arr[[1,2]])
import pandas as pd
df = pd.DataFrame({'a':[1,2,3]})
t = L(df, use_list=None)
test_eq(t[1,2], L(pd.DataFrame({'a':[2,3]}, index=[1,2]), use_list=None))
```
You can also modify an `L` with `append`, `+`, and `*`.
```
t = L()
test_eq(t, [])
t.append(1)
test_eq(t, [1])
t += [3,2]
test_eq(t, [1,3,2])
t = t + [4]
test_eq(t, [1,3,2,4])
t = 5 + t
test_eq(t, [5,1,3,2,4])
test_eq(L(1,2,3), [1,2,3])
test_eq(L(1,2,3), L(1,2,3))
t = L(1)*5
t = t.map(operator.neg)
test_eq(t,[-1]*5)
test_eq(~L([True,False,False]), L([False,True,True]))
t = L(range(4))
test_eq(zip(t, L(1).cycle()), zip(range(4),(1,1,1,1)))
t = L.range(100)
test_shuffled(t,t.shuffle())
def _f(x,a=0): return x+a
t = L(1)*5
test_eq(t.map(_f), t)
test_eq(t.map(_f,1), [2]*5)
test_eq(t.map(_f,a=2), [3]*5)
```
An `L` can be constructed from anything iterable, although tensors and arrays will not be iterated over on construction, unless you pass `use_list` to the constructor.
```
test_eq(L([1,2,3]),[1,2,3])
test_eq(L(L([1,2,3])),[1,2,3])
test_ne(L([1,2,3]),[1,2,])
test_eq(L('abc'),['abc'])
test_eq(L(range(0,3)),[0,1,2])
test_eq(L(o for o in range(0,3)),[0,1,2])
test_eq(L(array(0)),[array(0)])
test_eq(L([array(0),array(1)]),[array(0),array(1)])
test_eq(L(array([0.,1.1]))[0],array([0.,1.1]))
test_eq(L(array([0.,1.1]), use_list=True), [array(0.),array(1.1)]) # `use_list=True` to unwrap arrays/arrays
```
If `match` is not `None` then the created list is same len as `match`, either by:
- If `len(items)==1` then `items` is replicated,
- Otherwise an error is raised if `match` and `items` are not already the same size.
```
test_eq(L(1,match=[1,2,3]),[1,1,1])
test_eq(L([1,2],match=[2,3]),[1,2])
test_fail(lambda: L([1,2],match=[1,2,3]))
```
If you create an `L` from an existing `L` then you'll get back the original object (since `L` uses the `NewChkMeta` metaclass).
```
test_is(L(t), t)
```
An `L` is considred equal to a list if they have the same elements. It's never considered equal to a `str` a `set` or a `dict` even if they have the same elements/keys.
```
test_eq(L(['a', 'b']), ['a', 'b'])
test_ne(L(['a', 'b']), 'ab')
test_ne(L(['a', 'b']), {'a':1, 'b':2})
```
### `L` Methods
```
show_doc(L.__getitem__)
t = L(range(12))
test_eq(t[1,2], [1,2]) # implicit tuple
test_eq(t[[1,2]], [1,2]) # list
test_eq(t[:3], [0,1,2]) # slice
test_eq(t[[False]*11 + [True]], [11]) # mask
test_eq(t[array(3)], 3)
show_doc(L.__setitem__)
t[4,6] = 0
test_eq(t[4,6], [0,0])
t[4,6] = [1,2]
test_eq(t[4,6], [1,2])
show_doc(L.unique)
test_eq(L(4,1,2,3,4,4).unique(), [4,1,2,3])
show_doc(L.val2idx)
test_eq(L(1,2,3).val2idx(), {3:2,1:0,2:1})
show_doc(L.filter)
list(t)
test_eq(t.filter(lambda o:o<5), [0,1,2,3,1,2])
test_eq(t.filter(lambda o:o<5, negate=True), [5,7,8,9,10,11])
show_doc(L.argwhere)
test_eq(t.argwhere(lambda o:o<5), [0,1,2,3,4,6])
show_doc(L.map)
test_eq(L.range(4).map(operator.neg), [0,-1,-2,-3])
```
If `f` is a string then it is treated as a format string to create the mapping:
```
test_eq(L.range(4).map('#{}#'), ['#0#','#1#','#2#','#3#'])
```
If `f` is a dictionary (or anything supporting `__getitem__`) then it is indexed to create the mapping:
```
test_eq(L.range(4).map(list('abcd')), list('abcd'))
```
You can also pass the same `arg` params that `bind` accepts:
```
def f(a=None,b=None): return b
test_eq(L.range(4).map(f, b=arg0), range(4))
show_doc(L.map_dict)
test_eq(L(range(1,5)).map_dict(), {1:1, 2:2, 3:3, 4:4})
test_eq(L(range(1,5)).map_dict(operator.neg), {1:-1, 2:-2, 3:-3, 4:-4})
show_doc(L.zip)
t = L([[1,2,3],'abc'])
test_eq(t.zip(), [(1, 'a'),(2, 'b'),(3, 'c')])
t = L([[1,2,3,4],['a','b','c']])
test_eq(t.zip(cycled=True ), [(1, 'a'),(2, 'b'),(3, 'c'),(4, 'a')])
test_eq(t.zip(cycled=False), [(1, 'a'),(2, 'b'),(3, 'c')])
show_doc(L.map_zip)
t = L([1,2,3],[2,3,4])
test_eq(t.map_zip(operator.mul), [2,6,12])
show_doc(L.zipwith)
b = [[0],[1],[2,2]]
t = L([1,2,3]).zipwith(b)
test_eq(t, [(1,[0]), (2,[1]), (3,[2,2])])
show_doc(L.map_zipwith)
test_eq(L(1,2,3).map_zipwith(operator.mul, [2,3,4]), [2,6,12])
show_doc(L.itemgot)
test_eq(t.itemgot(1), b)
show_doc(L.attrgot)
# Example when items are not a dict
a = [SimpleNamespace(a=3,b=4),SimpleNamespace(a=1,b=2)]
test_eq(L(a).attrgot('b'), [4,2])
#Example of when items are a dict
b =[{'id': 15, 'name': 'nbdev'}, {'id': 17, 'name': 'fastcore'}]
test_eq(L(b).attrgot('id'), [15, 17])
show_doc(L.sorted)
test_eq(L(a).sorted('a').attrgot('b'), [2,4])
show_doc(L.split)
test_eq(L.split('a b c'), list('abc'))
show_doc(L.range)
test_eq_type(L.range([1,1,1]), L(range(3)))
test_eq_type(L.range(5,2,2), L(range(5,2,2)))
show_doc(L.concat)
test_eq(L([0,1,2,3],4,L(5,6)).concat(), range(7))
show_doc(L.copy)
t = L([0,1,2,3],4,L(5,6)).copy()
test_eq(t.concat(), range(7))
show_doc(L.map_first)
t = L(0,1,2,3)
test_eq(t.map_first(lambda o:o*2 if o>2 else None), 6)
show_doc(L.setattrs)
t = L(SimpleNamespace(),SimpleNamespace())
t.setattrs('foo', 'bar')
test_eq(t.attrgot('foo'), ['bar','bar'])
```
## Config -
```
#export
def save_config_file(file, d, **kwargs):
"Write settings dict to a new config file, or overwrite the existing one."
config = ConfigParser(**kwargs)
config['DEFAULT'] = d
config.write(open(file, 'w'))
#export
def read_config_file(file, **kwargs):
config = ConfigParser(**kwargs)
config.read(file)
return config
```
Config files are saved and read using Python's `configparser.ConfigParser`, inside the `DEFAULT` section.
```
_d = dict(user='fastai', lib_name='fastcore', some_path='test')
try:
save_config_file('tmp.ini', _d)
res = read_config_file('tmp.ini')
finally: os.unlink('tmp.ini')
test_eq(res['DEFAULT'], _d)
#export
def _add_new_defaults(cfg, file, **kwargs):
for k,v in kwargs.items():
if cfg.get(k, None) is None:
cfg[k] = v
save_config_file(file, cfg)
#export
@lru_cache(maxsize=None)
class Config:
"Reading and writing `settings.ini`"
def __init__(self, cfg_name='settings.ini'):
cfg_path = Path.cwd()
while cfg_path != cfg_path.parent and not (cfg_path/cfg_name).exists(): cfg_path = cfg_path.parent
self.config_path,self.config_file = cfg_path,cfg_path/cfg_name
assert self.config_file.exists(), f"Could not find {cfg_name}"
self.d = read_config_file(self.config_file)['DEFAULT']
_add_new_defaults(self.d, self.config_file,
host="github", doc_host="https://%(user)s.github.io", doc_baseurl="/%(lib_name)s/")
def __setitem__(self,k,v): self.d[k] = str(v)
def __contains__(self,k): return k in self.d
def save(self): save_config_file(self.config_file,self.d)
def __getattr__(self,k): return stop(AttributeError(k)) if k=='d' or k not in self.d else self.get(k)
def get(self,k,default=None): return self.d.get(k, default)
def path(self,k,default=None):
v = self.get(k, default)
return v if v is None else self.config_path/v
```
`Config` searches parent directories for a config file, and provides direct access to the 'DEFAULT' section. Keys ending in `_path` are converted to paths in the config file's directory.
```
try:
save_config_file('../tmp.ini', _d)
cfg = Config('tmp.ini')
finally: os.unlink('../tmp.ini')
test_eq(cfg.user,'fastai')
test_eq(cfg.doc_baseurl,'/fastcore/')
test_eq(cfg.get('some_path'), 'test')
test_eq(cfg.path('some_path'), Path('../test').resolve())
test_eq(cfg.get('foo','bar'),'bar')
```
# Export -
```
#hide
from nbdev.export import notebook2script
notebook2script()
```
| github_jupyter |
```
import tensorflow as tf
import numpy as np
import mvc_env
p = 4; initialization_stddev=1e-3; n_mlp_layers = 1; T = 2; n_nodes = 5
x = tf.placeholder(dtype=tf.float32, shape=[None, n_nodes])
adj = tf.placeholder(tf.float32, [None, n_nodes, n_nodes])
w = tf.placeholder(tf.float32, [None, n_nodes, n_nodes])
with tf.variable_scope('Q_func', reuse=False):
with tf.variable_scope('thetas'):
theta1 = tf.Variable(tf.random_normal([p], stddev=initialization_stddev), name='theta1')
theta2 = tf.Variable(tf.random_normal([p, p], stddev=initialization_stddev), name='theta2')
theta3 = tf.Variable(tf.random_normal([p, p], stddev=initialization_stddev), name='theta3')
theta4 = tf.Variable(tf.random_normal([p], stddev=initialization_stddev), name='theta4')
theta5 = tf.Variable(tf.random_normal([2 * p], stddev=initialization_stddev), name='theta5')
theta6 = tf.Variable(tf.random_normal([p, p], stddev=initialization_stddev), name='theta6')
theta7 = tf.Variable(tf.random_normal([p, p], stddev=initialization_stddev), name='theta7')
with tf.variable_scope('MLP', reuse=False):
Ws_MLP = []; bs_MLP = []
for i in range(n_mlp_layers):
Ws_MLP.append(tf.Variable(tf.random_normal([p, p], stddev=initialization_stddev),
name='W_MLP_' + str(i)))
bs_MLP.append(tf.Variable(tf.random_normal([p], stddev=initialization_stddev),
name='b_MLP_' + str(i)))
# Define the mus
# Initial mu
# Loop over t
for t in range(T):
# First part of mu
mu_part1 = tf.einsum('iv,k->ivk', x, theta1)
# Second part of mu
if t != 0:
mu_part2 = tf.einsum('kl,ivk->ivl', theta2, tf.einsum('ivu,iuk->ivk', adj, mu))
# Add some non linear transformations of the pooled neighbors' embeddings
with tf.variable_scope('MLP', reuse=False):
for i in range(n_mlp_layers):
mu_part2 = tf.nn.relu(tf.einsum('kl,ivk->ivl', Ws_MLP[i],
mu_part2) + bs_MLP[i])
# Third part of mu
mu_part3_0 = tf.einsum('ikvu->ikv', tf.nn.relu(tf.einsum('k,ivu->ikvu', theta4, w)))
mu_part3_1 = tf.einsum('kl,ilv->ivk', theta3, mu_part3_0)
# All all of the parts of mu and apply ReLui
if t != 0:
mu = tf.nn.relu(tf.add(mu_part1 + mu_part2, mu_part3_1, name='mu_' + str(t)))
else:
mu = tf.nn.relu(tf.add(mu_part1, mu_part3_1, name='mu_' + str(t)))
# Define the Qs
Q_part1 = tf.einsum('kl,ivk->ivl', theta6, tf.einsum('ivu,iuk->ivk', adj, mu))
Q_part2 = tf.einsum('kl,ivk->ivl', theta7, mu)
out = tf.identity(tf.einsum('k,ivk->iv', theta5,
tf.nn.relu(tf.concat([Q_part1, Q_part2], axis=2))),
name='Q')
env = mvc_env.MVC_env(5); env.reset()
sess = tf.InteractiveSession(); tf.global_variables_initializer().run()
sess.run(mu, feed_dict={x: np.array([1, 0, 0, 0, 0])[None],
adj: env.adjacency_matrix[None],
w: env.weight_matrix[None]})
a = sess.run(theta3)
sess.run(theta4)
sess.run(tf.einsum('ikvu->ikv', tf.nn.relu(tf.einsum('k,ivu->ikvu', theta4, w))),
feed_dict={w: env.weight_matrix[None]})
b = sess.run(tf.nn.relu(tf.einsum('k,ivu->ikv', theta4, w)),
feed_dict={w: env.weight_matrix[None]})[0, :, 3]
a.dot(b)
mu3_t0 = sess.run(tf.einsum('kl,ilv->ivk', theta3, tf.einsum('ikvu->ikv', tf.nn.relu(tf.einsum('k,ivu->ikvu', theta4, w)))),
feed_dict={w: env.weight_matrix[None]})
mu3_t0
mu_1_t0 = sess.run(tf.einsum('iv,k->ivk', x, theta1),
feed_dict={x: np.array([1, 0, 0, 0, 0])[None],
adj: env.adjacency_matrix[None],
w: env.weight_matrix[None]})
mu_1_t0
mu_t0 = sess.run(tf.nn.relu(mu_1_t0 + mu3_t0))
mu_t0
```
| github_jupyter |
# NQL Gridworld Pathfollowing
# Setup
```
%tensorflow_version 2.x
!test -d language_repo || git clone https://github.com/google-research/language.git language_repo
%cd /content/language_repo/language/nql
!pip install .
import random
import os
import numpy as np
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers, models, regularizers
from tensorflow.keras.optimizers import schedules
from tensorflow.keras.preprocessing import text
from tensorflow.keras.models import Sequential
import nql
print(tf.__version__)
# Makes things as deterministic as possible for reproducibility
seed = 1234 #@param{type: "integer"}
tf.random.set_seed(seed)
random.seed(seed)
```
## Build the gridworld
```
class GridWorld():
"""A simple grid world for testing NQL algorithms.
The world is square grid with every cell connected to its cardinal direction
neighbors, with the exception of several holes which are inescapable."""
size = None
context = None
words = ['go', 'top', 'left', 'right', 'bottom', 'center', 'up', 'down',
'then']
# A 2D dictionary mapping a cell and a valid move to another cell.
# Ex: _cell_move_cell['cell_2_3']['up'] == 'cell_1_3'
_cell_move_cell = {}
# _cell_paths[start_cell][num_moves][end_cell] is a list of efficient paths
# starting at start_cell, taking num_moves, and ending at end_cell
_cell_paths = {}
# _cell_dists[start_cell][dist] is a set of all of the cells exactly dist away
# from start cell
_cell_dists = {}
def __init__(self, size, n_holes):
if size ** 2 < n_holes:
raise ValueError(
'size ^ 2 (%d) must be great than n_holes (%d).'
% (size ** 2, n_holes))
self.size = size
self.holes = set()
# Remove cells
while len(self.holes) < n_holes:
r = random.randrange(size)
c = random.randrange(size)
self.holes.add((r,c))
self.context = nql.NeuralQueryContext()
self.context.declare_relation('up','place_t','place_t')
self.context.declare_relation('down','place_t','place_t')
self.context.declare_relation('left','place_t','place_t')
self.context.declare_relation('right','place_t','place_t')
kg_lines = []
for (r,c) in [(r,c) for r in range(self.size) for c in range(self.size)]:
if (r,c) in self.holes:
continue
self._cell_move_cell[self._cell(r,c)] = {}
def connect_to_cell(dest_r, dest_c, dir):
dest = self._cell(dest_r, dest_c)
self._cell_move_cell[self._cell(r,c)][dir] = dest
kg_lines.append('%s\t%s\t%s' % (dir, self._cell(r,c), dest))
if(r > 0):
connect_to_cell(r-1, c, 'up')
if(r < self.size-1):
connect_to_cell(r+1, c, 'down')
if(c > 0):
connect_to_cell(r, c-1, 'left')
if(c < self.size-1):
connect_to_cell(r, c+1, 'right')
self.context.load_kg(lines=kg_lines, freeze=True)
self.context.construct_relation_group('dir_g', 'place_t', 'place_t')
def _cell(self, i, j):
if (i, j) in self.holes:
return 'hole_%d_%d' % (i+1, j+1)
return 'cell_%d_%d' % (i+1, j+1)
# Output: [(query, starting_cell, ending_cell, num moves)]
def generate_examples(self, num, possible_moves=[1,2,3]):
# Extends all paths from start_cell by 1 step
def _extend_paths(start_cell):
cell_paths = self._cell_paths[start_cell]
cell_dists = self._cell_dists[start_cell]
prev_path_len = len(cell_dists) - 1
cell_dists.append(set())
cell_paths.append({})
if not len(cell_dists[prev_path_len]):
# We're already longer than the longest path possible from this cell.
return
seen_cells = set(
[cell for cells_at_dist in cell_dists for cell in cells_at_dist])
for cell in cell_dists[prev_path_len]:
prev_paths = cell_paths[prev_path_len][cell]
if cell not in self._cell_move_cell:
continue # This is a hole
for (dir, next_cell) in self._cell_move_cell[cell].items():
if next_cell not in seen_cells: # This is an efficient path
if next_cell not in cell_dists[prev_path_len+1]:
# This is the first time we found this cell
cell_dists[prev_path_len+1].add(next_cell)
cell_paths[prev_path_len+1][next_cell] = []
for path in prev_paths:
if prev_path_len == 0:
new_path = 'go ' + dir
else:
new_path = path + ' then ' + dir
cell_paths[prev_path_len+1][next_cell].append(new_path)
def generate_example(num_moves):
example = 'go '
while True:
start_row = random.randrange(self.size)
start_col = random.randrange(self.size)
starting_cell = self._cell(start_row, start_col)
if not starting_cell in self._cell_move_cell:
# Starting cell is a hole. Try again.
continue
if not starting_cell in self._cell_paths:
# We've never started from this cell before.
self._cell_paths[starting_cell] = [{starting_cell: ['']}]
self._cell_dists[starting_cell] = [{starting_cell}]
while len(self._cell_paths[starting_cell]) <= num_moves:
_extend_paths(starting_cell)
possible_ending_cells = list(self._cell_paths[starting_cell][num_moves])
if not possible_ending_cells:
# No paths num_moves long from this cell. Start over.
continue
ending_cell = random.choice(possible_ending_cells)
path = random.choice(
self._cell_paths[starting_cell][num_moves][ending_cell])
return (path, starting_cell, ending_cell)
examples = [None] * num
while True:
for i in range(num):
# Each element of possible_moves is equally likely to be selected
num_moves = random.choice(possible_moves)
example = ''
while not example:
example, starting_cell, ending_cell = generate_example(num_moves)
if examples[i] is None:
examples[i] = ({
"input_text": example,
"start": tf.squeeze(self.context.one(starting_cell, 'place_t').tf),
"start_name": starting_cell,
"end_name": ending_cell,
}, tf.squeeze(self.context.one(ending_cell, 'place_t').tf))
yield examples[i]
grid_size = 10#@param {type: "integer"}
p_holes = 0.25 #@param {type: "number"}
grid_holes = grid_size ** 2 * p_holes
max_moves = 10#@param {type: "integer"}
env = GridWorld(grid_size, grid_holes)
print("Holes:", env.holes)
from itertools import islice
print("Examples:", list(islice(env.generate_examples(4), 3)))
nqc = env.context
dataset_size = 100000 #@param{type: "integer"}
train_gen = env.generate_examples(num=dataset_size, possible_moves=range(1, max_moves+1))
test_gen = env.generate_examples(int(dataset_size / 10))
max_seq_len = max(len(x["input_text"].split(" ")) for (x, _) in islice(train_gen, 1000))
print("max_seq_len is %d" % max_seq_len)
assert max_seq_len >= 2
assert max_seq_len < 50
train_dataset = tf.data.Dataset.from_generator(
lambda: (x for x in train_gen),
output_types=(
{ "input_text": tf.string,
"start": tf.float32,
"start_name": tf.string,
"end_name": tf.string,
}, tf.float32),
output_shapes=(
{ "input_text": tf.TensorShape([]),
"start": tf.TensorShape([nqc.get_max_id('place_t')]),
"start_name": tf.TensorShape([]),
"end_name": tf.TensorShape([]),
}, tf.TensorShape([nqc.get_max_id('place_t')])),
)
text_dataset = train_dataset.map(lambda x,_: x["input_text"])
for i in train_dataset.take(2):
print(repr(i))
for i in text_dataset.take(2):
print(repr(i))
print(repr(train_dataset))
print(repr(train_dataset.element_spec))
print(repr(text_dataset))
print(repr(text_dataset.element_spec))
```
## Example
```
name = nqc.get_entity_name(grid_size * 2, 'place_t')
print(name)
cell = nqc.one(name, 'place_t')
print(cell.eval())
cell = nqc.one(name, 'place_t').follow('right')
print(cell.eval())
cell = nqc.one(name, 'place_t').follow('right').follow('up')
print(cell.eval())
```
# Multiple Moves: Solve a problem with multiple possible templates
## RNN Model
```
#from tensorflow.compat.v2.keras.layers.experimental.preprocessing import TextVectorization
from tensorflow.keras.layers.experimental.preprocessing import TextVectorization
class NqlFollowLayer(layers.Layer):
def __init__(self, context, **kwargs):
self.context = context
super(NqlFollowLayer, self).__init__(**kwargs)
def build(self, input_shape):
assert isinstance(input_shape, list)
super(NqlFollowLayer, self).build(input_shape)
def call(self, x):
assert isinstance(x, list)
place_tf, dir_tf = x
place_nql = self.context.as_nql(place_tf, "place_t")
assert isinstance(place_nql, nql.NeuralQueryExpression)
dir_nql = self.context.as_nql(dir_tf, "dir_g")
assert isinstance(dir_nql, nql.NeuralQueryExpression)
new_place_nql = place_nql.follow(dir_nql)
return new_place_nql.tf
def compute_output_shape(self, input_shape):
return self.context.get_max_id("place_t")
def get_config(self):
config = super(NqlFollowLayer, self).get_config()
config['context'] = self.context
return config
def build_model(nqc, layer_width, embedding_dim, max_num_moves, text_dataset, num_layers=1, dropout=0.0, l1=0.0001, l2=0.0001):
# Transforms text input into int sequences of length max_seq_len
vectorize_layer = TextVectorization(max_tokens=len(env.words),
output_mode="int",
output_sequence_length=max_seq_len,
name="VectorizationLayer",
)
vectorize_layer.adapt(text_dataset)
# Build the encoder
text_input = layers.Input(shape=(1,), dtype=tf.string, name="input_text")
vec_input = vectorize_layer(text_input)
# Account for the OOV token in this layer
embedding_layer = layers.Embedding(input_dim=len(env.words)+1,
output_dim=embedding_dim,
mask_zero=True,
embeddings_regularizer=regularizers.l1_l2(l1=l1, l2=l2),
name="TextEmbedding")
enc_emb = embedding_layer(vec_input)
# Use an LSTM layer here to process the whole input sequence
encoder_lstm = layers.LSTM(layer_width,
recurrent_dropout=dropout,
kernel_regularizer=regularizers.l1_l2(l1=l1, l2=l2),
recurrent_regularizer=regularizers.l1_l2(l1=l1, l2=l2),
bias_regularizer=regularizers.l1_l2(l1=l1, l2=l2),
return_state=True,
name="EncoderLstm"
)
enc_out, enc_state_h, enc_state_c = encoder_lstm(enc_emb)
# Build the decoder
decoder_lstm = layers.LSTM(layer_width,
recurrent_dropout=dropout,
kernel_regularizer=regularizers.l1_l2(l1=l1, l2=l2),
recurrent_regularizer=regularizers.l1_l2(l1=l1, l2=l2),
bias_regularizer=regularizers.l1_l2(l1=l1, l2=l2),
return_sequences=True,
name="DecoderLstm"
)
decoder_move_model = Sequential(name="DecoderMoveModel")
for i in range(num_layers-1):
decoder_move_model.add(layers.Dense(layer_width,
kernel_regularizer=regularizers.l1_l2(l1=l1, l2=l2),
bias_regularizer=regularizers.l1_l2(l1=l1, l2=l2),
))
decoder_move_model.add(layers.Dense(
nqc.get_max_id("dir_g"), activation="softmax"
))
decoder_prob_model = Sequential(name="DecoderProbModel")
for i in range(num_layers-1):
decoder_prob_model.add(layers.Dense(layer_width,
kernel_regularizer=regularizers.l1_l2(l1=l1, l2=l2),
bias_regularizer=regularizers.l1_l2(l1=l1, l2=l2),
))
decoder_prob_model.add(layers.Dense(1, activation="sigmoid"))
initial_place = layers.Input(shape=(nqc.get_max_id("place_t"),), name="start")
# Initial state of the decoder is the final state of the encoder
dec_states = [enc_state_h, enc_state_c]
cur_place = initial_place
nql_layer = NqlFollowLayer(nqc, dynamic=True)
# This should get turned into a tensor of the right shape the first time it's
# updated.
prob_remaining = 1.0
final_place = tf.zeros(shape=(nqc.get_max_id('place_t'),))
decoder_lstm_outs = decoder_lstm(
tf.ones(shape=(1, max_num_moves, layer_width)),
initial_state=dec_states)
for i in range(max_num_moves):
move_out = decoder_move_model(decoder_lstm_outs[:,i,:])
prob_out = decoder_prob_model(decoder_lstm_outs[:,i,:])
## Use the output move to update the current place
cur_place = nql_layer([cur_place, move_out])
prob_stopping = prob_remaining * prob_out
prob_remaining = prob_remaining - prob_stopping
## Update final output place
final_place = final_place + (prob_stopping * cur_place)
# Build the final model that goes from text to a place
model = models.Model(inputs=[text_input, initial_place], outputs=final_place)
return model
#@title Model Params { form-width: "25%" }
layer_width = 128 #@param{type: "integer"}
embedding_dim = 4#@param{type: "integer"}
num_layers = 3#@param{type: "integer"}
l1 = 0.00002 #@param{type: "number"}
l2 = 0.00002 #@param{type: "number"}
dropout = 0.1 #@param{type: "number"}
# For setting up the TextVectorization layer
text_dataset_sample = text_dataset.take(1024).batch(1024)
model = build_model(nqc,
layer_width=layer_width,
embedding_dim=embedding_dim,
max_num_moves=max_moves,
text_dataset=text_dataset_sample,
dropout=dropout,
l1=l1,
l2=l2,
num_layers=num_layers)
batch_size = 1024#@param{type: "integer"}
learning_rate = 0.0025 #@param{type: "number"}
clip_norm = 2.0 #@param{type: "number"}
decay = 0.005 #@param{type: "number"}
optimizer = tf.keras.optimizers.Nadam(learning_rate=learning_rate,
clipnorm=clip_norm,
decay=decay,
)
model.compile(optimizer=optimizer,
loss="categorical_crossentropy",
metrics=["accuracy"],
# run_eagerly=True, # TODO(rofer): Figure out if this is only needed on CPUs
)
batched_dataset = train_dataset.batch(batch_size)
callbacks = [
keras.callbacks.TerminateOnNaN(),
]
n_epochs = 100#@param{type: "integer"}
history = model.fit(batched_dataset,
epochs=n_epochs,
steps_per_epoch=50,
callbacks=callbacks)
```
## Evaluation
## Accuracy by # of moves
```
# Accuracy by number of moves
num_batches = 3 #@param{type: "integer"}
num_samples = num_batches * batch_size
for moves in range(1, max_moves + 1):
print("For %d moves:" % moves)
one_move_dataset = tf.data.Dataset.from_generator(
lambda: (x for x in env.generate_examples(
num=num_samples, possible_moves=[moves]
)),
output_types=(
{ "input_text": tf.string,
"start": tf.float32,
"start_name": tf.string,
"end_name": tf.string,
}, tf.float32),
output_shapes=(
{ "input_text": tf.TensorShape([]),
"start": tf.TensorShape([nqc.get_max_id('place_t')]),
"start_name": tf.TensorShape([]),
"end_name": tf.TensorShape([]),
}, tf.TensorShape([nqc.get_max_id('place_t')])),
)
model.evaluate(one_move_dataset.take(num_samples).batch(batch_size))
print("")
```
## Live Evaluation
```
query = "go right then down" #@param{type: "string"}
start = "cell_3_3" #@param{type: "string"}
nql_start = nqc.one(start, "place_t")
if "<UNK>" in nql_start.eval():
print("%s is not a valid starting point. It's probably a hole or a typo." %
start)
else:
example_dataset = tf.data.Dataset.from_tensors({
"input_text": query,
"start": tf.squeeze(nql_start.tf),
})
pred = model.predict(example_dataset.batch(1))
print(pred.shape)
pred_nql = nqc.as_nql(pred, "place_t")
print(pred_nql.eval(as_top=3))
```
| github_jupyter |
```
import nltk
from nltk.tokenize import word_tokenize
from nltk.tag import pos_tag
from nltk.chunk import ne_chunk
sentence = 'European authorities fined Google a record $5.1 billion on Wednesday for abusing its power in the mobile phone market and ordered the company to alter its practices'
ne_tree = ne_chunk(pos_tag(word_tokenize(sentence)))
print(ne_tree)
ex = 'European authorities fined Google a record $5.1 billion on Wednesday for abusing its power in the mobile phone market and ordered the company to alter its practices'
def preprocess(sent):
sent = nltk.word_tokenize(sent)
sent = nltk.pos_tag(sent)
return sent
sent = preprocess(ex)
sent
pattern = 'NP: {<DT>?<JJ>*<NN>}'
cp = nltk.RegexpParser(pattern)
cs = cp.parse(sent)
print(cs)
NPChunker = nltk.RegexpParser(pattern)
result = NPChunker.parse(sent)
result.draw()
import Tkinter
from nltk.chunk import conlltags2tree, tree2conlltags
from pprint import pprint
iob_tagged = tree2conlltags(cs)
pprint(iob_tagged)
import spacy
from spacy import displacy
from collections import Counter
import en_core_web_sm
nlp = en_core_web_sm.load()
from pprint import pprint
doc = nlp('European authorities fined Google a record $5.1 billion on Wednesday for abusing its power in the mobile phone market and ordered the company to alter its practices')
pprint([(X.text, X.label_) for X in doc.ents])
pprint([(X, X.ent_iob_, X.ent_type_) for X in doc])
from bs4 import BeautifulSoup
import requests
import re
def url_to_string(url):
res = requests.get(url)
html = res.text
soup = BeautifulSoup(html, 'html5lib')
for script in soup(["script", "style", 'aside']):
script.extract()
return " ".join(re.split(r'[\n\t]+', soup.get_text()))
ny_bb = url_to_string('https://www.nytimes.com/2018/08/13/us/politics/peter-strzok-fired-fbi.html?hp&action=click&pgtype=Homepage&clickSource=story-heading&module=first-column-region®ion=top-news&WT.nav=top-news')
article = nlp(ny_bb)
len(article.ents)
labels = [x.label_ for x in article.ents]
Counter(labels)
items = [x.text for x in article.ents]
Counter(items).most_common(3)
sentences = [x for x in article.sents]
print(sentences[20])
displacy.render(nlp(str(sentences[20])), jupyter=True, style='ent')
sentence_spans = list(doc.sents)
displacy.serve(sentence_spans, style="dep")
displacy.render(nlp(str(sentences[20])), style='dep', jupyter = True, options = {'distance': 120})
[(x.orth_,x.pos_, x.lemma_) for x in [y
for y
in nlp(str(sentences[20]))
if not y.is_stop and y.pos_ != 'PUNCT']]
dict([(str(x), x.label_) for x in nlp(str(sentences[20])).ents])
print([(x, x.ent_iob_, x.ent_type_) for x in sentences[20]])
displacy.render(article, jupyter=True, style='ent')
```
| github_jupyter |
# Project: Train a Quadcopter How to Fly
Design an agent that can fly a quadcopter, and then train it using a reinforcement learning algorithm of your choice! Try to apply the techniques you have learnt, but also feel free to come up with innovative ideas and test them.

## Instructions
> **Note**: If you haven't done so already, follow the steps in this repo's README to install ROS, and ensure that the simulator is running and correctly connecting to ROS.
When you are ready to start coding, take a look at the `quad_controller_rl/src/` (source) directory to better understand the structure. Here are some of the salient items:
- `src/`: Contains all the source code for the project.
- `quad_controller_rl/`: This is the root of the Python package you'll be working in.
- ...
- `tasks/`: Define your tasks (environments) in this sub-directory.
- `__init__.py`: When you define a new task, you'll have to import it here.
- `base_task.py`: Generic base class for all tasks, with documentation.
- `takeoff.py`: This is the first task, already defined for you, and set to run by default.
- ...
- `agents/`: Develop your reinforcement learning agents here.
- `__init__.py`: When you define a new agent, you'll have to import it here, just like tasks.
- `base_agent.py`: Generic base class for all agents, with documentation.
- `policy_search.py`: A sample agent has been provided here, and is set to run by default.
- ...
### Tasks
Open up the base class for tasks, `BaseTask`, defined in `tasks/base_task.py`:
```python
class BaseTask:
"""Generic base class for reinforcement learning tasks."""
def __init__(self):
"""Define state and action spaces, initialize other task parameters."""
pass
def set_agent(self, agent):
"""Set an agent to carry out this task; to be called from update."""
self.agent = agent
def reset(self):
"""Reset task and return initial condition."""
raise NotImplementedError
def update(self, timestamp, pose, angular_velocity, linear_acceleration):
"""Process current data, call agent, return action and done flag."""
raise NotImplementedError
```
All tasks must inherit from this class to function properly. You will need to override the `reset()` and `update()` methods when defining a task, otherwise you will get `NotImplementedError`'s. Besides these two, you should define the state (observation) space and the action space for the task in the constructor, `__init__()`, and initialize any other variables you may need to run the task.
Now compare this with the first concrete task `Takeoff`, defined in `tasks/takeoff.py`:
```python
class Takeoff(BaseTask):
"""Simple task where the goal is to lift off the ground and reach a target height."""
...
```
In `__init__()`, notice how the state and action spaces are defined using [OpenAI Gym spaces](https://gym.openai.com/docs/#spaces), like [`Box`](https://github.com/openai/gym/blob/master/gym/spaces/box.py). These objects provide a clean and powerful interface for agents to explore. For instance, they can inspect the dimensionality of a space (`shape`), ask for the limits (`high` and `low`), or even sample a bunch of observations using the `sample()` method, before beginning to interact with the environment. We also set a time limit (`max_duration`) for each episode here, and the height (`target_z`) that the quadcopter needs to reach for a successful takeoff.
The `reset()` method is meant to give you a chance to reset/initialize any variables you need in order to prepare for the next episode. You do not need to call it yourself; it will be invoked externally. And yes, it will be called once before each episode, including the very first one. Here `Takeoff` doesn't have any episode variables to initialize, but it must return a valid _initial condition_ for the task, which is a tuple consisting of a [`Pose`](http://docs.ros.org/api/geometry_msgs/html/msg/Pose.html) and [`Twist`](http://docs.ros.org/api/geometry_msgs/html/msg/Twist.html) object. These are ROS message types used to convey the pose (position, orientation) and velocity (linear, angular) you want the quadcopter to have at the beginning of an episode. You may choose to supply the same initial values every time, or change it a little bit, e.g. `Takeoff` drops the quadcopter off from a small height with a bit of randomness.
> **Tip**: Slightly randomized initial conditions can help the agent explore the state space faster.
Finally, the `update()` method is perhaps the most important. This is where you define the dynamics of the task and engage the agent. It is called by a ROS process periodically (roughly 30 times a second, by default), with current data from the simulation. A number of arguments are available: `timestamp` (you can use this to check for timeout, or compute velocities), `pose` (position, orientation of the quadcopter), `angular_velocity`, and `linear_acceleration`. You do not have to include all these variables in every task, e.g. `Takeoff` only uses pose information, and even that requires a 7-element state vector.
Once you have prepared the state you want to pass on to your agent, you will need to compute the reward, and check whether the episode is complete (e.g. agent crossed the time limit, or reached a certain height). Note that these two things (`reward` and `done`) are based on actions that the agent took in the past. When you are writing your own agents, you have to be mindful of this.
Now you can pass in the `state`, `reward` and `done` values to the agent's `step()` method and expect an action vector back that matches the action space that you have defined, in this case a `Box(6,)`. After checking that the action vector is non-empty, and clamping it to the space limits, you have to convert it into a ROS `Wrench` message. The first 3 elements of the action vector are interpreted as force in x, y, z directions, and the remaining 3 elements convey the torque to be applied around those axes, respectively.
Return the `Wrench` object (or `None` if you don't want to take any action) and the `done` flag from your `update()` method (note that when `done` is `True`, the `Wrench` object is ignored, so you can return `None` instead). This will be passed back to the simulation as a control command, and will affect the quadcopter's pose, orientation, velocity, etc. You will be able to gauge the effect when the `update()` method is called in the next time step.
### Agents
Reinforcement learning agents are defined in a similar way. Open up the generic agent class, `BaseAgent`, defined in `agents/base_agent.py`, and the sample agent `RandomPolicySearch` defined in `agents/policy_search.py`. They are actually even simpler to define - you only need to implement the `step()` method that is discussed above. It needs to consume `state` (vector), `reward` (scalar value) and `done` (boolean), and produce an `action` (vector). The state and action vectors must match the respective space indicated by the task. And that's it!
Well, that's just to get things working correctly! The sample agent given `RandomPolicySearch` uses a very simplistic linear policy to directly compute the action vector as a dot product of the state vector and a matrix of weights. Then, it randomly perturbs the parameters by adding some Gaussian noise, to produce a different policy. Based on the average reward obtained in each episode ("score"), it keeps track of the best set of parameters found so far, how the score is changing, and accordingly tweaks a scaling factor to widen or tighten the noise.
```
%%html
<div style="width: 100%; text-align: center;">
<h3>Teach a Quadcopter How to Tumble</h3>
<video poster="images/quadcopter_tumble.png" width="640" controls muted>
<source src="images/quadcopter_tumble.mp4" type="video/mp4" />
<p>Video: Quadcopter tumbling, trying to get off the ground</p>
</video>
</div>
```
Obviously, this agent performs very poorly on the task. It does manage to move the quadcopter, which is good, but instead of a stable takeoff, it often leads to dizzying cartwheels and somersaults! And that's where you come in - your first _task_ is to design a better agent for this takeoff task. Instead of messing with the sample agent, create new file in the `agents/` directory, say `policy_gradients.py`, and define your own agent in it. Remember to inherit from the base agent class, e.g.:
```python
class DDPG(BaseAgent):
...
```
You can borrow whatever you need from the sample agent, including ideas on how you might modularize your code (using helper methods like `act()`, `learn()`, `reset_episode_vars()`, etc.).
> **Note**: This setup may look similar to the common OpenAI Gym paradigm, but there is one small yet important difference. Instead of the agent calling a method on the environment (to execute an action and obtain the resulting state, reward and done value), here it is the task that is calling a method on the agent (`step()`). If you plan to store experience tuples for learning, you will need to cache the last state ($S_{t-1}$) and last action taken ($A_{t-1}$), then in the next time step when you get the new state ($S_t$) and reward ($R_t$), you can store them along with the `done` flag ($\left\langle S_{t-1}, A_{t-1}, R_t, S_t, \mathrm{done?}\right\rangle$).
When an episode ends, the agent receives one last call to the `step()` method with `done` set to `True` - this is your chance to perform any cleanup/reset/batch-learning (note that no reset method is called on an agent externally). The action returned on this last call is ignored, so you may safely return `None`. The next call would be the beginning of a new episode.
One last thing - in order to run your agent, you will have to edit `agents/__init__.py` and import your agent class in it, e.g.:
```python
from quad_controller_rl.agents.policy_gradients import DDPG
```
Then, while launching ROS, you will need to specify this class name on the commandline/terminal:
```bash
roslaunch quad_controller_rl rl_controller.launch agent:=DDPG
```
Okay, now the first task is cut out for you - follow the instructions below to implement an agent that learns to take off from the ground. For the remaining tasks, you get to define the tasks as well as the agents! Use the `Takeoff` task as a guide, and refer to the `BaseTask` docstrings for the different methods you need to override. Use some debug print statements to understand the flow of control better. And just like creating new agents, new tasks must inherit `BaseTask`, they need be imported into `tasks/__init__.py`, and specified on the commandline when running:
```bash
roslaunch quad_controller_rl rl_controller.launch task:=Hover agent:=DDPG
```
> **Tip**: You typically need to launch ROS and then run the simulator manually. But you can automate that process by either copying/symlinking your simulator to `quad_controller_rl/sim/DroneSim` (`DroneSim` must be an executable/link to one), or by specifying it on the command line, as follows:
>
> ```bash
> roslaunch quad_controller_rl rl_controller.launch task:=Hover agent:=DDPG sim:=<full path>
> ```
## Task 1: Takeoff
### Implement takeoff agent
Train an agent to successfully lift off from the ground and reach a certain threshold height. Develop your agent in a file under `agents/` as described above, implementing at least the `step()` method, and any other supporting methods that might be necessary. You may use any reinforcement learning algorithm of your choice (note that the action space consists of continuous variables, so that may somewhat limit your choices).
The task has already been defined (in `tasks/takeoff.py`), which you should not edit. The default target height (Z-axis value) to reach is 10 units above the ground. And the reward function is essentially the negative absolute distance from that set point (upto some threshold). An episode ends when the quadcopter reaches the target height (x and y values, orientation, velocity, etc. are ignored), or when the maximum duration is crossed (5 seconds). See `Takeoff.update()` for more details, including episode bonus/penalty.
As you develop your agent, it's important to keep an eye on how it's performing. Build in a mechanism to log/save the total rewards obtained in each episode to file. Once you are satisfied with your agent's performance, return to this notebook to plot episode rewards, and answer the questions below.
### Plot episode rewards
Plot the total rewards obtained in each episode, either from a single run, or averaged over multiple runs.
```
# TODO: Read and plot episode rewards
%matplotlib inline
import pandas as pd
#df_stats = pd.read_csv('../out/task01/stats_2018-02-14_08-06-12.csv')
df_stats = pd.read_csv('../out/task01/stats_2018-02-20_11-28-13.csv')
df_stats[['total_reward']].plot(title="Episode Rewards")
%%html
<div style="width: 100%; text-align: center;">
<h3>Teach a Quadcopter How to Takeoff</h3>
<video poster="images/poster_01_takeoff.png" width="816" controls muted>
<source src="images/01_takeoff.mp4" type="video/mp4" />
<p>Video: Solution - Quadcopter Takeoff</p>
</video>
</div>
```
**Q**: What algorithm did you use? Briefly discuss why you chose it for this task.
**A**: The algorithm used was Deep Deterministic Policy Gradients (DDPG). This algorithm was chosen because of the continuous state and action spaces. This is actually an actor-critic method but the idea is similar. Alternatively, DQN (Deep Q-Network) could be used but the state and action spaces would have to converted to discrete space, an extra overhead I wanted to avoid.
**Q**: Using the episode rewards plot, discuss how the agent learned over time.
- Was it an easy task to learn or hard?
- Was there a gradual learning curve, or an aha moment?
- How good was the final performance of the agent? (e.g. mean rewards over the last 10 episodes)
**A**:
- Was it an easy task to learn or hard?
I think the answer to the question is dependant on the context. Once the setup was done, the model architecture was set, and the hyper-parameters were set, the task was not learned at all until around the episodes in the mid 100. All of a sudden, it learned the task.
However, what took up the most time was not the task itself but the overhead of getting a virtual machine of linux to run on my windows machine and get the network to communicate. I had to change BIOS settings to get this to work. It took 3 days to just get up and running. It would also crash all the time and use up all my memory. I think it took a week to even start the first task of take off. Then finding a good architecture (before it crashed) was challenging. In the end, having 3 layers of around 32 or 64 nodes worked. Did not need to user regularizers, dropout, or batch normalization.
- Was there a gradual learning curve, or an aha moment?
It was an aha momemnt. It didn't learn anything then it all of sudden learned around episode in the mid 100.
- How good was the final performance of the agent? (e.g. mean rewards over the last 10 episodes)
I would think they are good given the reward system used. I think they were good because the consistently stayed within the same range which was approximately -600. Please note the a reward system is arbitrarily set so good or bad depends on what the designer setup. This rewards system used negative rewards so the less negative the better. As you can see from my plot, it learned consistently around episode 200 but I let it run for 1000 episodes and the results were still consistent.
## Task 2: Hover
### Implement hover agent
Now, your agent must take off and hover at the specified set point (say, 10 units above the ground). Same as before, you will need to create an agent and implement the `step()` method (and any other supporting methods) to apply your reinforcement learning algorithm. You may use the same agent as before, if you think your implementation is robust, and try to train it on the new task. But then remember to store your previous model weights/parameters, in case your results were worth keeping.
### States and rewards
Even if you can use the same agent, you will need to create a new task, which will allow you to change the state representation you pass in, how you verify when the episode has ended (the quadcopter needs to hover for at least a few seconds), etc. In this hover task, you may want to pass in the target height as part of the state (otherwise how would the agent know where you want it to go?). You may also need to revisit how rewards are computed. You can do all this in a new task file, e.g. `tasks/hover.py` (remember to follow the steps outlined above to create a new task):
```python
class Hover(BaseTask):
...
```
**Q**: Did you change the state representation or reward function? If so, please explain below what worked best for you, and why you chose that scheme. Include short code snippet(s) if needed.
**A**: Yes, I did change the state representation and reward function. I included velocity in the z-axis as a state input. Howere, the action space stayed the same. The reward system changed, it was a negative weighted sum of the error from the target height, target orientation, and the target velocity. I put the weight for orientation to 0.0 because orientation was not used. I biased more the weight for the position at 0.7 and 0.3 for the weight of the velocity. The more away from the target the actual value is the bigger the number. The sum is negative for a negative rewards (punishment.) So the more close to zero the less punishment. There is a punishment if the error for position is too high. There is a positive reward if the time lapsed surpassed the set duration.
### Implementation notes
**Q**: Discuss your implementation below briefly, using the following questions as a guide:
- What algorithm(s) did you try? What worked best for you?
- What was your final choice of hyperparameters (such as $\alpha$, $\gamma$, $\epsilon$, etc.)?
- What neural network architecture did you use (if any)? Specify layers, sizes, activation functions, etc.
**A**:
- What algorithm(s) did you try? What worked best for you?
The algorithm used was again Deep Deterministic Policy Gradients (DDPG). In the end about 2 or 3 layers with about 4 to 8 nodes per layer worked out. I think it worked out because I limited the state and action space to just one dimension in position and ignoring rotation. I actually solved the first task again with all 3 position axis but it took so long (and crashed all the time) I would never be able to finish the assignment. I added dropout and batch normalization because in general I find those work. I added saving out model weights early because it would keep crashing. I did not have to change the hyperparameters, just the model architecture and state and reward system.
- What was your final choice of hyperparameters (such as αα , γγ , ϵϵ , etc.)?
I left the hyperameters as they initially were in the Takeoff task. I changed the model architecture and the state and reward system to get the results I was looking for. I would like to point out that I was stuck on this task for about 2 weeks but then Udacity added a 12.Troubleshooting section. There I saw that you were allowed to change the initial height to the target height. Before, I was trying to train it to takeoff to target height and then hover. It would keep crashing around 200 episodes or 1000 episodes so I couldn't really train it even if I wanted to.
- What neural network architecture did you use (if any)? Specify layers, sizes, activation functions, etc.
Th neural network architecture used was about 2 or 3 layers with about 4 to 8 nodes per layer for both the actor and the critic. I also added dropout and batch normalization. I found using less nodes and in general dropout and batch normalization helped out. Activation functions remained rectified linear units (relu) for all hidden layers. I did try sigmoid for all the hidden layers, I did not notice any difference visually or in the plot so I went back to relu.
### Plot episode rewards
As before, plot the episode rewards, either from a single run, or averaged over multiple runs. Comment on any changes in learning behavior.
```
# TODO: Read and plot episode rewards
%matplotlib inline
import pandas as pd
# stats_2018-02-19_15-18-18.csv - Started to work around episode 650
# stats_2018-02-20_10-32-16 - Should have been good around episode 350 but overrode the file.
# This should work stats_2018-02-20_11-02-40.csv.
df_stats = pd.read_csv('../out/task02/stats_2018-02-20_12-43-46.csv')
#df_stats = pd.read_csv('../out/task02/stats_2018-02-20_08-13-41.csv')
df_stats[['total_reward']].plot(title="Episode Rewards")
%%html
<div style="width: 100%; text-align: center;">
<h3>Teach a Quadcopter How to Hover</h3>
<video poster="images/poster_02_hover.png" width="816" controls muted>
<source src="images/02_hover.mp4" type="video/mp4" />
<p>Video: Solution - Quadcopter Hover</p>
</video>
</div>
```
## Task 3: Landing
What goes up, must come down! But safely!
### Implement landing agent
This time, you will need to edit the starting state of the quadcopter to place it at a position above the ground (at least 10 units). And change the reward function to make the agent learn to settle down _gently_. Again, create a new task for this (e.g. `Landing` in `tasks/landing.py`), and implement the changes. Note that you will have to modify the `reset()` method to return a position in the air, perhaps with some upward velocity to mimic a recent takeoff.
Once you're satisfied with your task definition, create another agent or repurpose an existing one to learn this task. This might be a good chance to try out a different approach or algorithm.
### Initial condition, states and rewards
**Q**: How did you change the initial condition (starting state), state representation and/or reward function? Please explain below what worked best for you, and why you chose that scheme. Were you able to build in a reward mechanism for landing gently?
**A**:
The initial condition was changed to start at the target height. State representation is the same as the Hover task, position z and velocity z. The reward function changed. It is similar in idea as the hover reward system except that I changed it to be a positive reward system and exponential. I only use position z and velocity z. It gets rewards by having the value close to the target. This is done by subtracting from 1 the difference and then taking the power (in this case 2). For example if the position is at the target, the difference would be 0. Subtracting 0 from 1 would make it 1 and then squaring it. So you get more points for having a difference closer to zero. I also gave a bonus for landing gently. If the velocity was too high, I gave a negative reward (punishment,) and end the episode. This way, the quadcopter cannot finish the episode if it is moving too fast.
### Implementation notes
**Q**: Discuss your implementation below briefly, using the same questions as before to guide you.
**A**:
-What algorithm(s) did you try? What worked best for you?
I still used DDPG again. I just changed the reward system. The model architecture was the same as hover.
-What was your final choice of hyperparameters (such as αα , γγ , ϵϵ , etc.)?
I used the same hyperparameters as the hover and the takeoff. I just changed the architecture.
-What neural network architecture did you use (if any)? Specify layers, sizes, activation functions, etc.
I kept the same network architecture as hover.
Th neural network architecture used was about 2 or 3 layers with about 4 to 8 nodes per layer for both the actor and the critic. I also added dropout and batch normalization. I found using less nodes and in general dropout and batch normalization helped out. Activation functions remained rectified linear units (relu) for all hidden layers.
### Plot episode rewards
As before, plot the episode rewards, either from a single run, or averaged over multiple runs. This task is a little different from the previous ones, since you're starting in the air. Was it harder to learn? Why/why not?
```
# TODO: Read and plot episode rewards
%matplotlib inline
import pandas as pd
df_stats = pd.read_csv('../out/task03/stats_2018-02-20_21-03-45.csv')
df_stats[['total_reward']].plot(title="Episode Rewards")
%%html
<div style="width: 100%; text-align: center;">
<h3>Teach a Quadcopter How to Land</h3>
<video poster="images/poster_03_land.png" width="816" controls muted>
<source src="images/03_land.mp4" type="video/mp4" />
<p>Video: Solution - Quadcopter Land</p>
</video>
</div>
```
## Task 4: Combined
In order to design a complete flying system, you will need to incorporate all these basic behaviors into a single agent.
### Setup end-to-end task
The end-to-end task we are considering here is simply to takeoff, hover in-place for some duration, and then land. Time to create another task! But think about how you might go about it. Should it be one meta-task that activates appropriate sub-tasks, one at a time? Or would a single combined task with something like waypoints be easier to implement? There is no right or wrong way here - experiment and find out what works best (and then come back to answer the following).
**Q**: What setup did you ultimately go with for this combined task? Explain briefly.
**A**:
### Implement combined agent
Using your end-to-end task, implement the combined agent so that it learns to takeoff (at least 10 units above ground), hover (again, at least 10 units above ground), and gently come back to ground level.
### Combination scheme and implementation notes
Just like the task itself, it's up to you whether you want to train three separate (sub-)agents, or a single agent for the complete end-to-end task.
**Q**: What did you end up doing? What challenges did you face, and how did you resolve them? Discuss any other implementation notes below.
**A**:
What I ended up doing was created an agent for task 4 called task04_ddpg_agent.py with a class called Task04_DDPG. I also created a task called combined.py with a class called Combined. How these worked is by using instances of the other tasks object from classes and swapped them depending on the mode. The swapping used a modulus
self.mode %= 3
and the possible modes are 0, 1, 2. Where mode 0 is task takeoff, mode 1 is task hover, and mode 2 is land. The task would just keep cycling if the simulator is running.
For the agent, in the constructor __init__, the 3 task objects and 3 agents were instantiated and assigned to member variables. The instance for the task is used to instantiate the appropriate instance of the corresponding agent. This was important because numpy shapes and member variables would not match up if they were mixed and matched. The model weights for each tasks were trained and preloaded from the other tasks. You can look at the different model weights and plots generated for this project in the 'out' folder under its corresponding task.s
self.task_takeoff = takeoff_b.TakeoffB()
self.task_hover = hover_b.HoverB()
self.task_land = land_b.LandB()
self.o_task01_agent = task01_ddpg_agent_b.Task01_DDPG(self.task_takeoff)
self.o_task02_agent = task02_ddpg_agent_b.Task02_DDPG(self.task_hover)
self.o_task03_agent = task03_ddpg_agent_b.Task03_DDPG(self.task_land)
Then to know which agent instance to use, a member variable would be set. Depending on the mode, this member
variable would be updated.
# Current agent
self.o_current_agent = self.o_task01_agent
For the agents all the same methods exists so they would be called, the instance method would be called and returned.
def act(self, states):
return self.o_current_agent.act(states)
To know when to swap instances for agents, the 'done' variable was checked during the step() method.
def step(self, state, reward, done):
self.total_reward += reward
if done:
# DEBUG
#
print("\n\nDEBUG - done have been called. self.mode: {}\n\n".format(self.mode))
# Go to the mode
self.mode += 1
#
# Cycle mode back to the beginning
self.mode %= 3
if self.mode == 0:
self.o_current_agent = self.o_task01_agent
self.task = self.task_takeoff
if self.mode == 1:
self.o_current_agent = self.o_task02_agent
self.task =self.task_hover
if self.mode == 2:
self.o_current_agent = self.o_task03_agent
self.task = self.task_land
done = False
# If we cycle back to take off, we count it as an episode
# and reset reward.
if self.mode == 0:
# Write episode stats
self.write_stats([self.episode_num, self.total_reward])
self.reset_episode_vars()
self.episode_num += 1
return self.o_current_agent.step(state, reward, done)
NOTE: self.task was not used in the end because I instantiated the agents with their correct task instances in the __init__, but I kept it there.
This alone almost worked. When 'done' is True, .reset() method for a task is called. However, I do not want the position or the angular or linear velocity to be reset. I created a new member variable to all of the tasks called
self.current_condition
In the task Combine class, in the update() method takes the current condition and set it as the member variable for the current task
def update(self, timestamp, pose, angular_velocity, linear_acceleration):
# Save current condition
position = np.array([pose.position.x, pose.position.y, pose.position.z])
self.current_condition = Pose(
position=Point(*position),
orientation=Quaternion(0.0, 0.0, 0.0, 1.0),
), Twist(
linear=linear_acceleration,
angular=angular_velocity
)
# Update the tasks current condition.
self.o_current_task.current_condition = self.current_condition
When 'done' is set the True in the Combine class method .update(), the mode will swap the current task it will also
set the newly set current condition to be what it is now and not reset it to a height of zero or target height and still have the same angular and linear velocity.
if done:
self.mode += 1
self.mode %= 3
if self.mode == 0:
self.o_current_task = self.task_takeoff
if self.mode == 1:
self.o_current_task = self.task_hover
if self.mode == 2:
self.o_current_task = self.task_land
done = False
# Update the tasks current condition.
self.o_current_task.current_condition = self.current_condition
If you let the simulation run, it will just loop.
The reward is given by a combine specific episode. This is anytime the mode loops back to 0. The rewards are added up and written to the .csv.
I was going to freeze all the layers for the models to prevent training, but decided not to.
<B>NOTE: It seems that for all the simulations (Takeoff, Hover, Land, Combined) the first episode is useless. The second episode and beyond works. I don't know how to prevent this. I gather that this is expected behaviour.</B>
### Plot episode rewards
As before, plot the episode rewards, either from a single run, or averaged over multiple runs.
```
# TODO: Read and plot episode rewards
%matplotlib inline
import pandas as pd
df_stats = pd.read_csv('../out/task04/stats_2018-02-21_14-07-26.csv')
df_stats[['total_reward']].plot(title="Episode Rewards")
%%html
<div style="width: 100%; text-align: center;">
<h3>Teach a Quadcopter How to combined Takeoff, Hover, and Land</h3>
<video poster="images/poster_04_combined.png" width="816" controls muted>
<source src="images/04_combined.mp4" type="video/mp4" />
<p>Video: Solution - Quadcopter combine Takeoff, Hover, and Land</p>
</video>
</div>
```
## Reflections
**Q**: Briefly summarize your experience working on this project. You can use the following prompts for ideas.
- What was the hardest part of the project? (e.g. getting started, running ROS, plotting, specific task, etc.)
- How did you approach each task and choose an appropriate algorithm/implementation for it?
- Did you find anything interesting in how the quadcopter or your agent behaved?
**A**:
- What was the hardest part of the project? (e.g. getting started, running ROS, plotting, specific task, etc.)
Getting setup was by far the hardest thing about this project. I could not get it run in the cloud. It used up all my computer's resources. I had to change BIOS settings. I had to use 2 different operating systems. The simulation crashed. Then the tasks were challenging themselves because there were no examples. Luckily, there were a few dedicated people trying to complete the assignment in Slack where I can bounce ideas off of.
Unfortunately, I had a lot of prior commitment so it was hard to give the dedicated extra time to get this assignment done. I had to stay up until 3 or 4 am in the morning and a lot of the times the simulations were worthless. I also wasn't sure if I was even on the right track.
Also, the office hours I have on my calendar were for general deep learning questions. There were very few office hours specifically for this assignment. this made it hard as well.
- How did you approach each task and choose an appropriate algorithm/implementation for it?
I stuck with DDPG mainly because the problem was continuous. The lectures taught with discrete space, but DDPG was the only one explained with continuous space. I just stuck with it. I know other people solved it using DQN. I did not want the extra overhead of converting continuous space to discrete space. I had enough hoops to jump through.
- Did you find anything interesting in how the quadcopter or your agent behaved?
The implementation I did seemed too abrupt. I would rather a smoother transition. I would also like to know what are waypoints and how to use them. I assume waypoints may solve the abrupt problem. I also used different model architecture and state spaces for the tasks. I figured if I was able to use the same model architecture and state space for all the tasks, I would just have to update the weights of the model and transitions between tasks would be smoother. The code would be easier to write as well instead of swapping current tasks and current agents.
### To run all 4 simulation
<B>NOTE: It seems that for all the simulations (Takeoff, Hover, Land, Combined) the first episode is useless. The second episode and beyond works. I don't know how to prevent this. I gather that this is expected behaviour.</B>
```bash
roslaunch quad_controller_rl rl_controller.launch task:=Takeoff agent:=Task01_DDPG
roslaunch quad_controller_rl rl_controller.launch task:=Hover agent:=Task02_DDPG
roslaunch quad_controller_rl rl_controller.launch task:=Land agent:=Task03_DDPG
roslaunch quad_controller_rl rl_controller.launch task:=Combined agent:=Task04_DDPG
```
| github_jupyter |
<a href="https://colab.research.google.com/github/parekhakhil/pyImageSearch/blob/main/1405_opencv_haar_cascades_zip.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>

# OpenCV Haar Cascades
### by [PyImageSearch.com](http://www.pyimagesearch.com)
## Welcome to **[PyImageSearch Plus](http://pyimg.co/plus)** Jupyter Notebooks!
This notebook is associated with the [OpenCV Haar Cascades](https://www.pyimagesearch.com/2021/04/12/opencv-haar-cascades/) blog post published on 2021-04-12.
Only the code for the blog post is here. Most codeblocks have a 1:1 relationship with what you find in the blog post with two exceptions: (1) Python classes are not separate files as they are typically organized with PyImageSearch projects, and (2) Command Line Argument parsing is replaced with an `args` dictionary that you can manipulate as needed.
We recommend that you execute (press ▶️) the code block-by-block, as-is, before adjusting parameters and `args` inputs. Once you've verified that the code is working, you are welcome to hack with it and learn from manipulating inputs, settings, and parameters. For more information on using Jupyter and Colab, please refer to these resources:
* [Jupyter Notebook User Interface](https://jupyter-notebook.readthedocs.io/en/stable/notebook.html#notebook-user-interface)
* [Overview of Google Colaboratory Features](https://colab.research.google.com/notebooks/basic_features_overview.ipynb)
As a reminder, these PyImageSearch Plus Jupyter Notebooks are not for sharing; please refer to the **Copyright** directly below and **Code License Agreement** in the last cell of this notebook.
Happy hacking!
*Adrian*
<hr>
***Copyright:*** *The contents of this Jupyter Notebook, unless otherwise indicated, are Copyright 2021 Adrian Rosebrock, PyimageSearch.com. All rights reserved. Content like this is made possible by the time invested by the authors. If you received this Jupyter Notebook and did not purchase it, please consider making future content possible by joining PyImageSearch Plus at http://pyimg.co/plus/ today.*
### Download the code zip file
```
!wget https://pyimagesearch-code-downloads.s3-us-west-2.amazonaws.com/opencv-haar-cascades/opencv-haar-cascades.zip
!unzip -qq opencv-haar-cascades.zip
%cd opencv-haar-cascades
```
## Blog Post Code
### Import Packages
```
# import the necessary packages
from imutils.video import VideoStream
import argparse
import imutils
import time
import cv2
import os
```
### Implementing OpenCV Haar Cascade object detection (face, eyes, and mouth)
```
# first, let's get a video which we can use for haar cascade object detection
!wget https://colab-notebook-videos.s3-us-west-2.amazonaws.com/guitar.mp4
# construct the argument parser and parse the arguments
#ap = argparse.ArgumentParser()
#ap.add_argument("-c", "--cascades", type=str, default="cascades",
# help="path to input directory containing haar cascades")
#args = vars(ap.parse_args())
# since we are using Jupyter Notebooks we can replace our argument
# parsing code with *hard coded* arguments and values
args = {
"cascades": "cascades",
"input": "guitar.mp4",
"output": "output.avi"
}
# initialize a dictionary that maps the name of the haar cascades to
# their filenames
detectorPaths = {
"face": "haarcascade_frontalface_default.xml",
"eyes": "haarcascade_eye.xml",
"smile": "haarcascade_smile.xml",
}
# initialize a dictionary to store our haar cascade detectors
print("[INFO] loading haar cascades...")
detectors = {}
# loop over our detector paths
for (name, path) in detectorPaths.items():
# load the haar cascade from disk and store it in the detectors
# dictionary
path = os.path.sep.join([args["cascades"], path])
detectors[name] = cv2.CascadeClassifier(path)
# grab a reference to the video file and initialize pointer to output
# video file
print("[INFO] opening video file...")
vs = cv2.VideoCapture(args["input"])
writer = None
# loop over the frames from the video stream
while True:
# grab the next frame
frame = vs.read()[1]
# if we did not grab a frame then we have reached the end of the
# video
if frame is None:
break
# rotate the image
frame = imutils.rotate(frame, -90)
# resize the frame and convert it to grayscale
frame = imutils.resize(frame, width=500)
gray = cv2.cvtColor(frame, cv2.COLOR_BGR2GRAY)
# perform face detection using the appropriate haar cascade
faceRects = detectors["face"].detectMultiScale(
gray, scaleFactor=1.05, minNeighbors=5, minSize=(30, 30),
flags=cv2.CASCADE_SCALE_IMAGE)
# loop over the face bounding boxes
for (fX, fY, fW, fH) in faceRects:
# extract the face ROI
faceROI = gray[fY:fY+ fH, fX:fX + fW]
# apply eyes detection to the face ROI
eyeRects = detectors["eyes"].detectMultiScale(
faceROI, scaleFactor=1.1, minNeighbors=10,
minSize=(15, 15), flags=cv2.CASCADE_SCALE_IMAGE)
# apply smile detection to the face ROI
smileRects = detectors["smile"].detectMultiScale(
faceROI, scaleFactor=1.1, minNeighbors=10,
minSize=(15, 15), flags=cv2.CASCADE_SCALE_IMAGE)
# loop over the eye bounding boxes
for (eX, eY, eW, eH) in eyeRects:
# draw the eye bounding box
ptA = (fX + eX, fY + eY)
ptB = (fX + eX + eW, fY + eY + eH)
cv2.rectangle(frame, ptA, ptB, (0, 0, 255), 2)
# loop over the smile bounding boxes
for (sX, sY, sW, sH) in smileRects:
# draw the smile bounding box
ptA = (fX + sX, fY + sY)
ptB = (fX + sX + sW, fY + sY + sH)
cv2.rectangle(frame, ptA, ptB, (255, 0, 0), 2)
# draw the face bounding box on the frame
cv2.rectangle(frame, (fX, fY), (fX + fW, fY + fH),
(0, 255, 0), 2)
# if the video writer is None *AND* we are supposed to write
# the output video to disk initialize the writer
if writer is None and args["output"] is not None:
fourcc = cv2.VideoWriter_fourcc(*"MJPG")
writer = cv2.VideoWriter(args["output"], fourcc, 20,
(frame.shape[1], frame.shape[0]), True)
# if the writer is not None, write the frame to disk
if writer is not None:
writer.write(frame)
# do a bit of cleanup
vs.release()
# check to see if the video writer point needs to be released
if writer is not None:
writer.release()
```
Note that the above code block may take time to execute. If you are interested to view the video within Colab just execute the following code blocks. Note that it may be time-consuming.
Our output video is produced in `.avi` format. First, we need to convert it to `.mp4` format.
```
!ffmpeg -i output.avi output.mp4
#@title Display video inline
from IPython.display import HTML
from base64 import b64encode
mp4 = open("output.mp4", "rb").read()
dataURL = "data:video/mp4;base64," + b64encode(mp4).decode()
HTML("""
<video width=400 controls>
<source src="%s" type="video/mp4">
</video>
""" % dataURL)
```
This code is referred from [this StackOverflow thread](https://stackoverflow.com/a/57378660/7636462).
For a detailed walkthrough of the concepts and code, be sure to refer to the full tutorial, [*OpenCV Haar Cascades*](https://www.pyimagesearch.com/2021/04/12/opencv-haar-cascades/) blog post published on 2021-04-12.
# Code License Agreement
```
Copyright (c) 2021 PyImageSearch.com
SIMPLE VERSION
Feel free to use this code for your own projects, whether they are
purely educational, for fun, or for profit. THE EXCEPTION BEING if
you are developing a course, book, or other educational product.
Under *NO CIRCUMSTANCE* may you use this code for your own paid
educational or self-promotional ventures without written consent
from Adrian Rosebrock and PyImageSearch.com.
LONGER, FORMAL VERSION
Permission is hereby granted, free of charge, to any person obtaining
a copy of this software and associated documentation files
(the "Software"), to deal in the Software without restriction,
including without limitation the rights to use, copy, modify, merge,
publish, distribute, sublicense, and/or sell copies of the Software,
and to permit persons to whom the Software is furnished to do so,
subject to the following conditions:
The above copyright notice and this permission notice shall be
included in all copies or substantial portions of the Software.
Notwithstanding the foregoing, you may not use, copy, modify, merge,
publish, distribute, sublicense, create a derivative work, and/or
sell copies of the Software in any work that is designed, intended,
or marketed for pedagogical or instructional purposes related to
programming, coding, application development, or information
technology. Permission for such use, copying, modification, and
merger, publication, distribution, sub-licensing, creation of
derivative works, or sale is expressly withheld.
THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND,
EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES
OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND
NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS
BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN
ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN
CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE
SOFTWARE.
```
| github_jupyter |
```
import altair as alt
import pandas as pd
from vega_datasets import data
```
# 1. Color the lines of a slopegraph, based on one of the fields, if the slope is positive.
Source: https://github.com/altair-viz/altair/issues/2178 and https://altair-viz.github.io/gallery/slope_graph.html.
```
source = data.barley()
alt.Chart(source).mark_line().encode(
x="year:O",
y="median_yield:Q",
color=alt.condition("datum.slope > 0", "site:N", alt.value("lightgray")),
).transform_aggregate(
median_yield="median(yield)", groupby=["year", "site"]
).transform_pivot(
"year", value="median_yield", groupby=["site"]
).transform_calculate(
slope=alt.datum["1932"] - alt.datum["1931"]
).transform_fold(
["1931", "1932"], as_=["year", "median_yield"]
)
```
# 2. Layered count histogram and cumulative histogram.
Source: https://vega.github.io/vega-lite/examples/layer_cumulative_histogram.html.
```
movies = data.movies()
movies.head()
base = (
alt.Chart(movies)
.encode(
x=alt.X("bin_IMDB_Rating:Q", axis=alt.Axis(title="IMDB Rating")),
x2=alt.X2("bin_IMDB_Rating_end:Q"),
)
.transform_bin("bin_IMDB_Rating", field="IMDB Rating", bin=True)
.transform_aggregate(
groupby=["bin_IMDB_Rating", "bin_IMDB_Rating_end"], count="count()",
)
.transform_filter("datum.bin_IMDB_Rating != null")
.transform_window(
cumulative_count="sum(count)", sort=[alt.SortField("bin_IMDB_Rating")]
)
)
count_hist = base.mark_bar(color="yellow", opacity=0.5).encode(y=alt.Y("count:Q"))
cum_hist = base.mark_bar().encode(y=alt.Y("cumulative_count:Q"))
cum_hist + count_hist
```
# 3. Chart title with the same style as the axis title or axis labels (via built-in guide styles).
Source: https://altair-viz.github.io/user_guide/generated/core/altair.TitleParams.html and https://vega.github.io/vega-lite/docs/config.html#guide-config.
```
source = alt.pd.DataFrame(
{
"a": ["A", "B", "C", "D", "E", "F", "G", "H", "I"],
"b": [28, 55, 43, 91, 81, 53, 19, 87, 52],
}
)
default = (
alt.Chart(source, title=alt.TitleParams("Bar Chart", style="group-title"))
.mark_bar()
.encode(x="a", y="b")
)
axis_title = (
alt.Chart(source, title=alt.TitleParams("Bar Chart", style="guide-title"))
.mark_bar()
.encode(x="a", y="b")
)
axis_labels = (
alt.Chart(source, title=alt.TitleParams("Bar Chart", style="guide-label"))
.mark_bar()
.encode(x="a", y="b")
)
default | axis_title | axis_labels
```
# 4. Use the `pointer` cursor to identify a clickable mark, that is, a mark with an associated selection.
Source: https://altair-viz.github.io/user_guide/generated/core/altair.Cursor.html and https://www.w3schools.com/cssref/pr_class_cursor.asp.
**Note**: It currently does not work in JupyterLab (more info [here](https://github.com/altair-viz/altair/issues/535)).
```
selector = alt.selection_single()
alt.Chart(data.cars.url).mark_rect(cursor="pointer").encode(
x="Cylinders:O",
y="Origin:N",
color=alt.condition(selector, "count()", alt.value("lightgray")),
).properties(width=300, height=180).add_selection(selector)
```
| github_jupyter |
<!--NOTEBOOK_HEADER-->
*This notebook contains course material from [CBE40455](https://jckantor.github.io/CBE40455) by
Jeffrey Kantor (jeff at nd.edu); the content is available [on Github](https://github.com/jckantor/CBE40455.git).
The text is released under the [CC-BY-NC-ND-4.0 license](https://creativecommons.org/licenses/by-nc-nd/4.0/legalcode),
and code is released under the [MIT license](https://opensource.org/licenses/MIT).*
<!--NAVIGATION-->
< [Points after Touchdown Decision](http://nbviewer.jupyter.org/github/jckantor/CBE40455/blob/master/notebooks/06.05-Points-after-Touchdown-Decision.ipynb) | [Contents](toc.ipynb) | [Measuring Return](http://nbviewer.jupyter.org/github/jckantor/CBE40455/blob/master/notebooks/07.01-Measuring-Return.ipynb) ><p><a href="https://colab.research.google.com/github/jckantor/CBE40455/blob/master/notebooks/07.00-Risk-and-Diversification.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open in Google Colaboratory"></a><p><a href="https://raw.githubusercontent.com/jckantor/CBE40455/master/notebooks/07.00-Risk-and-Diversification.ipynb"><img align="left" src="https://img.shields.io/badge/Github-Download-blue.svg" alt="Download" title="Download Notebook"></a>
# Risk and Diversification
## Popular books
There is a popular literature on the role of randomness in financial and engineering analysis. This is a small selection along with links to related material and commentaries.
* Poundstone, William. Fortune's Formula: The untold story of the scientific betting system that beat the casinos and wall street. Hill and Wang, 2010. https://www.amazon.com/Fortunes-Formula-Scientific-Betting-Casinos-ebook/dp/B000SBTWNC
* MacLean, Leonard C., et al. "How Does the Fortune’s Formula Kelly CapitalGrowth Model Perform?." The Journal of Portfolio Management 37.4 (2011): 96-111. http://hari.seshadri.com/docs/kelly-betting/kelly1.pdf
* Ziemba, Bill. "Fortune's Formula." Wilmott 2018.94 (2018): 32-37. https://alphazadvisors.com/wp-content/uploads/2018/06/Ziemba-2018-Wilmott_Fortunes_Formula.pdf
* Wójtowicz, Michał. "A counterexample to the Fortune’s Formula investing method." Revista de la Real Academia de Ciencias Exactas, Físicas y Naturales. Serie A. Matemáticas 113.2 (2019): 749-767. https://link.springer.com/article/10.1007/s13398-018-0508-x
* Taleb, Nassim Nicholas. The black swan: The impact of the highly improbable. Vol. 2. Random house, 2007. https://www.amazon.com/Black-Swan-Improbable-Robustness-Fragility/dp/081297381X
* Taleb, Nassim Nicholas. "Statistical Consequences of Fat Tails: Real World Preasymptotics, Epistemology, and Applications." arXiv preprint arXiv:2001.10488 (2020). https://arxiv.org/pdf/2001.10488.pdf
* Thorp, Edward O. A man for all markets: From Las Vegas to wall street, how i beat the dealer and the market. Random House Incorporated, 2017. https://www.amazon.com/Man-All-Markets-Street-Dealer/dp/0812979907
<!--NAVIGATION-->
< [Points after Touchdown Decision](http://nbviewer.jupyter.org/github/jckantor/CBE40455/blob/master/notebooks/06.05-Points-after-Touchdown-Decision.ipynb) | [Contents](toc.ipynb) | [Measuring Return](http://nbviewer.jupyter.org/github/jckantor/CBE40455/blob/master/notebooks/07.01-Measuring-Return.ipynb) ><p><a href="https://colab.research.google.com/github/jckantor/CBE40455/blob/master/notebooks/07.00-Risk-and-Diversification.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open in Google Colaboratory"></a><p><a href="https://raw.githubusercontent.com/jckantor/CBE40455/master/notebooks/07.00-Risk-and-Diversification.ipynb"><img align="left" src="https://img.shields.io/badge/Github-Download-blue.svg" alt="Download" title="Download Notebook"></a>
| github_jupyter |
```
!date
```
#### Python [conda env: wrs-plots]
```
import pandas as pd
import pandas.io.sql as sqlio
import psycopg2
import datetime
x = datetime.datetime.now() - datetime.timedelta(days=10)
start, end = x.replace(day=1).strftime('%Y-%m-%d'), x.replace(day=30).strftime('%Y-%m-%d')
start, end
conn = psycopg2.connect("host='{}' port={} dbname='{}' user={} password={}".format(host, port, dbname, username, pwd))
sql = '''
select count(*) n_scenes,
case when split_part(s.name, '_', 2) = ''
then right(left(s.name,6),3)
else left(split_part(s.name, '_', 3), 3) end as path,
case when split_part(s.name, '_', 2) = ''
then right(left(s.name,9),3)
else right(split_part(s.name, '_', 3), 3) end as row
from ordering_scene s
left join ordering_order o on o.id=s.order_id
where
o.order_date::date >= '{0}'
and o.order_date::date <= '{1}'
-- and s.download_size > 0
and s.sensor_type = 'landsat'
group by path, row
;
'''.format(start, end)
dat = sqlio.read_sql_query(sql, conn)
print(dat['n_scenes'].max(), dat['n_scenes'].sum())
dat.head()
conn = None
```
---
```
import shapely
import shapely.ops
import geopandas as gp
def load_wrs(filename='wrs2_asc_desc/wrs2_asc_desc.shp'):
wrs = gp.GeoDataFrame.from_file(filename)
# PATH, ROW, geometry
return wrs
FEATURES = load_wrs()
%pylab inline
import matplotlib as mpl
from mpl_toolkits.basemap import Basemap
from matplotlib.collections import PatchCollection
import numpy as np
#from descartes import PolygonPatch
from matplotlib.patches import Polygon
from IPython.display import Image
def get_poly_wrs(path, row, features=FEATURES, facecolor='w'):
prid = '{}_{}'.format(path, row)
ix = (features['PATH'] == path) & (features['ROW'] == row)
geom = features[ix]
#geom = [f for f in features if prid == f.name]
if (len(geom) != 1):
print(prid)
print(geom)
raise AssertionError()
poly = geom.geometry.values[0]
if poly.geom_type == 'Polygon':
lons, lats = poly.exterior.coords.xy
elif poly.geom_type == 'MultiPolygon':
lons = []; lats = []
for subpoly in poly:
ln, la = subpoly.exterior.coords.xy
lons += ln; lats+=la
return lons, lats
def plot_poly(lons, lats, mm, **kwargs):
# facecolor='red', edgecolor='red', alpha=0.1
x, y = mm( lons, lats )
xy = zip(x,y)
patch = Polygon( xy, **kwargs )
return patch
FIGSIZE=(10,6)
def make_basemap(path_rows_alpha):
water = 'white'
earth = 'grey'
#font = {'size' : 20}
#mpl.rc('font', **font)
RED = '#f03b20'
fig, ax = plt.subplots(figsize=FIGSIZE)
mm = Basemap(llcrnrlon=-180,llcrnrlat=-60,urcrnrlon=180,urcrnrlat=90, projection='mill')
#mm = Basemap(projection='mill',lon_0=0)
mm.drawmapboundary()
coast = mm.drawcoastlines()
continents = mm.fillcontinents(color=earth, lake_color=water)
bound= mm.drawmapboundary(fill_color=water)
countries = mm.drawcountries()
merid = mm.drawmeridians(np.arange(-180, 180, 60), labels=[False, False, False, True])
parall = mm.drawparallels(np.arange(-80, 80, 20), labels=[True, False, False, False])
ax = plt.gca()
for path, row, alpha in path_rows_alpha:
lons, lats = get_poly_wrs(path, row)
delta_lons = abs(max(lons) - min(lons))
if delta_lons > 180:
#print('PATH {} ROW {} | MINLON {} MAXLON {}'.format(path, row, min(lons), max(lons)))
def p_dateline(x):
if x > 0:
return x-360
return x
lons = map(p_dateline, lons)
patch = plot_poly(lons, lats, mm, facecolor='red', edgecolor='red', alpha=alpha )
ax.add_patch(patch)
```
# Show geographic regions of products
```
MINALPHA=0.035
MAXALPHA=1.0
MINVALUES=dat['n_scenes'].min()
MAXVALUES=dat['n_scenes'].max()
def get_alpha(x, b, a, mmin, mmax):
# b = max desired
# a = min desired
return a + (((b-a)*(x-mmin))/(mmax-mmin))
dat['path'] = dat['path'].astype(int)
dat['row'] = dat['row'].astype(int)
dat['alpha'] = dat['n_scenes'].apply(lambda v: get_alpha(v, MAXALPHA, MINALPHA, MINVALUES, MAXVALUES))
dat = dat.sort_values(by='alpha')
path_rows_alpha = dat[['path', 'row', 'alpha']].values
dat.tail()
def fake_cb():
mmin, mmax = (MINVALUES, MAXVALUES)
step = 100
mymap = mpl.colors.LinearSegmentedColormap.from_list('mycolors',['white','red'])
# Using contourf to provide my colorbar info, then clearing the figure
Z = [[0,0],[0,0]]
plt.subplots(figsize=FIGSIZE, facecolor='w')
levels = range(mmin,mmax+step,step)
CS3 = plt.contourf(Z, levels, cmap=mymap)
plt.clf()
return CS3
cb = fake_cb()
make_basemap(path_rows_alpha)
plt.title('Landsat Scenes (path/row) Ordered\nALL USERS: {} - {}'.format(start, end), fontsize=14)
cbar = plt.colorbar(cb); cbar.ax.set_title(' Scenes',weight='bold', fontsize=14); cbar.ax.tick_params(labelsize=12)
pltfname = '/tmp/paths_rows_ordered_ALL.png'
print(pltfname)
plt.savefig(pltfname, bbox_inches='tight')
```
| github_jupyter |
<a href="https://colab.research.google.com/github/cltl/python-for-text-analysis/blob/colab/Chapters-colab/Chapter_22_Sentiment_analysis_with_VADER.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
%%capture
!wget https://github.com/cltl/python-for-text-analysis/raw/master/zips/Data.zip
!wget https://github.com/cltl/python-for-text-analysis/raw/master/zips/images.zip
!wget https://github.com/cltl/python-for-text-analysis/raw/master/zips/Extra_Material.zip
!unzip Data.zip -d ../
!unzip images.zip -d ./
!unzip Extra_Material.zip -d ../
!rm Data.zip
!rm Extra_Material.zip
!rm images.zip
```
# Chapter 22 - Sentiment analysis using VADER
In this notebook, we focus on sentiment analysis, which is the task of determining whether a text expresses a negative, neutral, or positive opinion. We introduce how to work with [VADER](https://github.com/cjhutto/vaderSentiment) as part of the NLTK to perform sentiment analysis. Given a sentence, e.g., "I like Python", VADER will predict a sentiment score on a scale from -1 to 1. The goal of this notebook is to show you how to work with VADER. One of the learning goals of the accompanying assignment is to gain insight into VADER by reading blogs about the system.
### at the end of this notebook, you will:
* have VADER installed on your computer
* be able to load the VADER model
* be able to apply the VADER model on new sentences:
* with and without lemmatization
* with providing VADER with certain parts of speech, e.g., providing the adjectives from a sentence as input to VADER
### If you want to learn more about this chapter, you might find the following links useful:
* [blog on sentiment analysis](https://towardsdatascience.com/quick-introduction-to-sentiment-analysis-74bd3dfb536c)
* [GitHub repository](https://github.com/cjhutto/vaderSentiment)
* [this blog](http://t-redactyl.io/blog/2017/04/using-vader-to-handle-sentiment-analysis-with-social-media-text.html)
If you have questions about this chapter, please contact us (cltl.python.course@gmail.com).
## 1. Downloading VADER package
Please run the following commands to download VADER to your computer.
```
import nltk
# You only need to run this cell once.
# After that, you can comment it out.
nltk.download('vader_lexicon', quiet=False)
```
To verify that the download was successful, you can run the following command.
```
from nltk.sentiment import vader
```
## 2. Load VADER model
The model can be loaded in the following way.
```
from nltk.sentiment.vader import SentimentIntensityAnalyzer
vader_model = SentimentIntensityAnalyzer()
```
We will use the following three sentences:
```
sentences = ["Here are my sentences.",
"It's a nice day.",
"It's a rainy day."]
```
The next for loop assigns a sentiment score from VADER to **each sentence**.
```
for sent in sentences:
scores = vader_model.polarity_scores(sent)
print()
print('INPUT SENTENCE', sent)
print('VADER OUTPUT', scores)
```
VADER provides a dictionary containing four ratings, i.e., keys, for each sentence.
The sentence is rated on how negative (key *neg*), positive (key *pos*), and neutral (key *neu*), it is.
Also, there is a *compound* key that combines the values of the keys *neg*, *pos*, and *neu* into one single score, i.e., the *compound* key. The compound value ranges from -1, i.e., very negative, to 1, i.e., very positive. You can read more about the VADER system on [this blog](http://t-redactyl.io/blog/2017/04/using-vader-to-handle-sentiment-analysis-with-social-media-text.html).
## 3. Using spaCy to manipulate the input to VADER
In the examples in Section 2, VADER always takes into account each token, i.e., word, in the sentence to arrive at its sentiment prediction. In this section,
we are going to use spaCy to manipulate the input to VADER. This is one way to gain insight into how language systems work, i.e., by manipulating the input and inspecting the output.
We use spaCy as a tool to manipulate the input to VADER.
Please first install spaCy by following the instructions from **Chapter 19 - More about Natural Language Processing Tools (spaCy) -- Section 2.1 Installing and loading spaCy**
```
import spacy
nlp = spacy.load('en') # en_core_web_sm
```
The next function defines an API to process texts (textual_unit) using different settings. This function operates on texts and assumes spaCy is loaded with the corresponding language model as we just did. Take a little time to analyze the function, which uses certain spaCy token properties to process the text in different ways and returns the VADER sentiment.
```
def run_vader(nlp,
textual_unit,
lemmatize=False,
parts_of_speech_to_consider=set(),
verbose=0):
"""
Run VADER on a sentence from spacy
:param nlp: spaCy model
:param str textual unit: a textual unit, e.g., sentence, sentences (one string)
(by looping over doc.sents)
:param bool lemmatize: If True, provide lemmas to VADER instead of words
:param set parts_of_speech_to_consider:
-empty set -> all parts of speech are provided
-non-empty set: only these parts of speech are considered
:param int verbose: if set to 1, information is printed
about input and output
:rtype: dict
:return: vader output dict
"""
doc = nlp(textual_unit)
input_to_vader = []
for sent in doc.sents:
for token in sent:
if verbose >= 2:
print(token, token.pos_)
to_add = token.text
if lemmatize:
to_add = token.lemma_
if to_add == '-PRON-':
to_add = token.text
if parts_of_speech_to_consider:
if token.pos_ in parts_of_speech_to_consider:
input_to_vader.append(to_add)
else:
input_to_vader.append(to_add)
scores = vader_model.polarity_scores(' '.join(input_to_vader))
if verbose >= 1:
print()
print('INPUT SENTENCE', sent)
print('INPUT TO VADER', input_to_vader)
print('VADER OUTPUT', scores)
return scores
```
We can now use various API settings to experiment with processing text in different ways.
### 3.1 Lemmatization
The first setting is to lemmatize the provided sentence. If you want to know more about lemmas, you can read [this blog](https://www.retresco.de/en/encyclopedia/lemma/). If you want the function to print more information, you can set the keyword parameter **verbose** to 1.
```
sentences = ["Here are my sentences.",
"It's a nice day.",
"It's a rainy day."]
prediction = run_vader(nlp, sentences[1], lemmatize=False, verbose=1)
print(prediction)
prediction = run_vader(nlp, sentences[1], lemmatize=True, verbose=1)
print(prediction)
```
Perhaps you are surprised to see that there is no difference in the output! This is useful information for you to try to understand how the system works! Perhaps there are sentences for which it does matter. Feel free to experiment with other sentences.
### 3.2 Filter on part of speech
You can also filter on the part of speech, i.e., we let VADER make a prediction by only considering the nouns, verbs, or adjectives. The manipulation of the input to VADER allows you to gain insight how the system works.
Only Nouns:
```
run_vader(nlp,
sentences[1],
lemmatize=True,
parts_of_speech_to_consider={'NOUN'},
verbose=1)
```
Please note that in this case, VADER only considers *day* to predict the sentiment score and ignores all other words. Do you agree with the assessment that *day* is neutral? I hope you have a great day!
Only verbs:
```
run_vader(nlp,
sentences[1],
lemmatize=True,
parts_of_speech_to_consider={'VERB'},
verbose=1)
```
This is even more interesting. The part of speech label *VERB* is not applied to any of the tokens (*'s* is labeled as auxiliary and not with the label VERB). We have not provided VADER with input at all!
Let's also try adjectives:
```
run_vader(nlp,
sentences[1],
lemmatize=True,
parts_of_speech_to_consider={'ADJ'},
verbose=1)
```
Very interesting! By only considering adjectives, i.e., *nice*, VADER predicts that the sentence is very positive! I hope that you start to get an understanding of how VADER works.
```
```
| github_jupyter |
# Preámbulo
```
# Módulo de economía ambiental #
import EcoAmb as EA
# Gráficos y estilo #
import matplotlib.pyplot as plt
import matplotlib.patches as mpl_patches
from mpl_toolkits.mplot3d import Axes3D # perspectiva 3d
from matplotlib import cm # mapas de colores
plt.style.use('seaborn-whitegrid') # estilo de gráficos
# Módulo de arreglos y vectores
import numpy as np
# Gráficos interactivos #
from ipywidgets import interact, fixed # Para hacer gráficos interactivos
# Ignorar advertencias
import warnings
warnings.filterwarnings("ignore")
# Recargar módulo automáticamente
%load_ext autoreload
%autoreload 2
```
# Preferencias
## Preferencias fuertemente convexas
Función Cobb-Douglas
$u(x,y) = x^{\alpha}y^{1-\alpha}$
```
import EcoAmb as EA
def My_U1(x,y, r = 0.5):
return x**r *y**(1-r)
C1 = EA.Consumer(My_U1)
#C1.PlotDemanda(p2 = 3, I =7, r1 = 0.9)
C1.Indiff_Map_Plot(np.array([1,2,3]))
#C1.Indiff_Map_Plot(np.array([3]))
#C1.TMGS(c1 = 9, u = 3)
```
$dy/dx$ = -Umg_x/Umg_y
si aumento en $dx$ el consumo de $x$, ¿cuánto disminuyo $y$ para quedar en el mismo nivel de utilidad?
$dy/dx = -10 $
Función de utilidad cuasi-lineal
$u(x,y) = x^r+ y$
con $r \in [0,1[$
```
import EcoAmb as EA
def My_U2(x,y, r = 0.5):
return y + x**r
C2 = EA.Consumer(My_U2)
C2.Indiff_Map_Plot(np.array([1,2,3,4,5,6]))
#C2.PlotDemanda( p2 = 4.5, I = 7)
```
## Preferencias cónvexas
Utilidad lineal
$u(x,y) = x + y$
```
def My_U3(x,y):
return x+y
C3 = EA.Consumer(My_U3)
C3.Indiff_Map_Plot(np.array([1,2,3,4,5,6]))
#C3.PlotDemanda( p2 = 4.5, I = 7)
```
## Preferencias no convexas
$u(x,y) = x^2 + y^2$
```
def My_U4(x,y):
return x**2 + y**2
C4 = EA.Consumer(My_U4)
C4.Indiff_Map_Plot(np.array([1,6,12,24,64]))
#C4.Indiff_Map_Plot(np.array([6]))
#C4.TMGS(c1 = 2.2, u = 6)
```
$u(x,y) = \log(x) + y^2$
```
def My_U5(x,y):
return np.log(x) + y**2
C5 = EA.Consumer(My_U5)
C5.Indiff_Map_Plot(np.array([1,2,3,4,5,6]))
```
# Demanda
## Planteamiento y gráfica
El problema que resuelve cada consumidor es
$\max_{c_1,c_2} u(x,y)$ sujeto a $p_1 c_1 + p_2 c_2 \leq I$
En este caso la función de utilidad corresponde a una Cobb-Douglas
$u(x,y) = x^{\alpha}\cdot y^{1-\alpha}$
```
import EcoAmb as EA
def My_U1(x,y, r = 0.5):
return x**r *y**(1-r)
def My_U5(x,y):
return np.log(x) + y**2
C1 = EA.Consumer(My_U1)
C5 = EA.Consumer(My_U5)
interact(C1.Opt_consumo_Plot,
p1 = (1,6,0.1), p2 = (1,2,0.1), I = (1,10,0.1))
#interact(C1.Opt_consumo_Plot, p1 = fixed(1),p2 = (C1.Pmin, C1.Pmax, 1) , I = (4,10,1))
interact(C1.PlotDemanda,p2=(1,3,0.1), I = (1,10,0.5))
def My_U3(x,y):
return x+y
C3 = EA.Consumer(My_U3)
#interact(C3.Opt_consumo_Plot, p1 = fixed(1),p2 = (C1.Pmin, C1.Pmax, 1) , I = (4,10,1))
interact(C3.PlotDemanda,p2=(1,3,0.1), I = (1,10,0.5))
def My_U5(x,y):
return np.log(x) + y**2
C5 = EA.Consumer(My_U5)
#C5.Indiff_Map_Plot(np.array([1,2,3,4,5,6]))
interact(C5.PlotDemanda,p2=(1,3,0.1), I = (1,10,0.5))
```
## Resolviendo analíticamente
Recta presupuestaria:
$p_x x + p_y y = R$
$y = -(p_x/p_y) x + R/p_y$
$\mathcal{L}(x,y,\lambda) = u(x,y) + \lambda (R-p_x x -p_y y)$
* Derivamos con respecto a x e y
$\frac{\partial }{\partial x} \mathcal{L} = \frac{\partial}{\partial x} u -\lambda p_x = 0$
$\frac{\partial }{\partial y} \mathcal{L} = \frac{\partial}{\partial y} u -\lambda p_y = 0$
* Despejamos $\lambda$ en cada ecuación
$\lambda = \frac{\partial}{\partial x}( u(x,y) )/ p_x$
$\lambda = \frac{\partial}{\partial y}( u(x,y) )/ p_y$
* Igualamos
$\frac{\partial}{\partial x}( u(x,y) )/ p_x = \frac{\partial}{\partial y}( u(x,y) )/ p_y$
$\frac{\mbox{Umg}_x}{p_x} = \frac{\mbox{Umg}_y}{p_y}$
## Un ejemplo
Encontremos la demanda de $x$ e $y$.
1. Primer paso: Encontrar $\mbox{Umg}_x$ y $\mbox{Umg}_y$
$\mbox{Umg}_x = \frac{\partial u(x,y)}{\partial x} = \alpha x^{\alpha-1}y^{1-\alpha} = \alpha (y/x)^{1-\alpha}$
$\mbox{Umg}_y = \frac{\partial u(x,y)}{\partial y} = (1- \alpha) x^{\alpha}y^{-\alpha} = (1-\alpha )(x/y)^{\alpha}$
2. Segundo paso: Reemplazamos en la condición de optimalidad $\frac{\mbox{Umg}_x}{p_x} = \frac{\mbox{Umg}_y}{p_y}$
$\frac{\mbox{Umg}_x}{\mbox{Umg}_y} = \frac{ \alpha x^{\alpha-1}y^{1-\alpha}}{(1- \alpha) x^{\alpha}y^{-\alpha}} $
$\frac{\mbox{Umg}_x}{\mbox{Umg}_y} = \frac{\alpha}{1-\alpha} (y/x)^{1-\alpha } (y/x) ^{\alpha} = \frac{\alpha}{1-\alpha} (y/x)$
$\frac{\alpha}{1-\alpha}x ^{-1} y ^{1} = \frac{\alpha}{1-\alpha}\frac{y}{x}$
3. Igualamos la TMGS al ratio de precios
$\frac{\mbox{Umg}_x}{\mbox{Umg}_y} = \frac{\alpha}{1-\alpha}\frac{y}{x} = \frac{p_x}{p_y}$
4. Despejamos una variable
$y = x (p_x/p_y)(1-\alpha)/\alpha$
5. Reemplazamos en la restricción presupuestaria $x p_x + y p_y = R$
$ R/p_y - x (p_x/p_y) = x (p_x/p_y)\frac{1-\alpha}{\alpha} $
6. Despejamos x
$x^* = \frac{R \alpha}{p_x }$
$y^* = \frac{R(1-\alpha)}{p_y}$
```
C1.PlotDemanda(p2 = 3, I =7)
```
# Pruebas varias
```
def My_U(x,y, r = 0.5):
return x**r *y**(1-r)
def My_U2(x,y):
return x+y
def My_U5(x,y):
return np.log(x) + y**2
def My_U6(x,y, r = 0.5):
return y + x**r
C1 = EA.Consumer(My_U)
C1.PlotDemanda(p2 = 3, I =7, r1 = 0.9)
C1.Indiff_Map_Plot(np.array([0.5,1,2,3]))
#interact(Ramon.PlotDemanda, p2 = (Ramon.Pmin, Ramon.Pmax, 1) , I = (4,10,1), r1 = (0.00,.9,0.01))
C2 = EA.Consumer(U = My_U6)
#interact(Lorca.PlotDemanda, p2 = (Pmin, Pmax, 1) , I = (4,10,1))
C2.Indiff_Map_Plot(np.array([1,2,3,4,5,6]))
C2.PlotDemanda( p2 = 4.5, I = 7)
C3 = EA.Consumer(My_U2)
C3.Indiff_Map_Plot(np.array([1,2,3,4,5,6]))
C3.PlotDemanda( p2 = 4.5, I = 7)
```
| github_jupyter |
```
import pandas as pd
import os
from pybedtools import BedTool
import seaborn as sns
import numpy as np
%matplotlib inline
```
Compare sgRNAs designed with AlleleAnalyzer for WTC to platinum sgRNAs from Scott & Zhang 2017.
Make BED file of first 5 exons of *PCSK9* using GENCODE release 19 (hg19) as done in that paper.
```
# load gene annotations
pcsk9_annots = pd.read_csv('/pollard/data/genetics/GENCODE/release19/pcsk9_annots.gtf',
sep='\t', header=None, names=[
'seqname',
'source',
'feature',
'start',
'end',
'score',
'strand',
'frame',
'attribute'
])
pcsk9_exons = pcsk9_annots.query('(feature == "exon")').copy()
pcsk9_exons['exon_number'] = pcsk9_exons['attribute'].str.split('exon_number').str[1].str.split(';').str[0]
pcsk9_exons['level'] = pcsk9_exons['attribute'].str.split('level').str[1].str.split(';').str[0]
pcsk9_exons['exon_number'] = pcsk9_exons['exon_number'].astype(int)
pcsk9_exons['level'] = pcsk9_exons['level'].astype(int)
pcsk9_exons['transcript_type'] = pcsk9_exons['attribute'].str.split('transcript_type "').str[1].str.split('";').str[0]
pcsk9_first_five_exons = pcsk9_exons.query('(0 <= exon_number <= 5) and (level < 3) and (transcript_type == "protein_coding")')
exon_ids = []
for ix, row in pcsk9_first_five_exons.iterrows():
exon_ids.append(row['attribute'].split('exon_id "')[1].split('";')[0])
# make BED file of exons
pcsk9_first_five_exons_bed = pcsk9_first_five_exons[['seqname', 'start', 'end', 'exon_number']].copy()
pcsk9_first_five_exons_bed.to_csv('/pollard/home/kathleen/projects/AlleleAnalyzer/manuscript_analyses/sgRNA_comparison_scott_zhang/dat/pcsk9_first_five_exons.bed',
sep='\t', header=None, index=None)
sel_genes_spcas9 = pd.read_csv('171101_NM_Supplementary_Information/SpCas9_platinum_targets-select_genes.csv',
skiprows=1)
pcsk9_spcas9_platinum = sel_genes_spcas9.query('transcript == "PCSK9-001"').copy()
pcsk9_spcas9_platinum_first_five = pcsk9_spcas9_platinum[pcsk9_spcas9_platinum['exon id'].isin(exon_ids)]
pcsk9_spcas9_platinum_first_five.columns
pcsk9_spcas9_platinum_first_five.head()
pcsk9_spcas9_platinum_first_five['PAM orient.'].unique()
# make BED file for PCSK9 platinum sgRNAs
pcsk9_platinum_first_five_bed = pcsk9_spcas9_platinum_first_five[['chrm', 'start', 'end', 'seq.']]
pcsk9_platinum_first_five_bed.to_csv('/pollard/home/kathleen/projects/AlleleAnalyzer/manuscript_analyses/sgRNA_comparison_scott_zhang/dat/pcsk9_platinum_targets_first_five_exons.bed',
sep='\t', header=None, index=None)
len(pcsk9_platinum_first_five_bed)
```
Verified that platinum sgRNAs overlap the exons I've identified via IGV.
Reformat variants:
`python ~/projects/AlleleAnalyzer/preprocessing/generate_gens_dfs/get_gens_df.py /pollard/home/kathleen/conklin_wt_seq_data/wtc_wgs_data/phased_yin/wtc_PASS_hg19.phased.vcf.gz pcsk9_first_five_exons.bed pcsk9_first_five_exons_gens --bed -v`
Annotate variants:
`python ~/projects/AlleleAnalyzer/preprocessing/annotate_variants/annot_variants.py -v pcsk9_first_five_exons_gens.h5 SpCas9 /pollard/data/projects/AlleleAnalyzer_data/pam_sites_hg19/ /pollard/data/vertebrate_genomes/human/hg19/hg19/hg19.fa pcsk9_first_five_exons_annots --guide_len=20`
Design sgRNAs for these first 5 exons in WTC using AlleleAnalyzer with CRISPOR specificity scoring.
`python ~/projects/AlleleAnalyzer/scripts/gen_sgRNAs.py /pollard/home/kathleen/conklin_wt_seq_data/wtc_wgs_data/phased_yin/wtc_PASS_hg19.phased.vcf.gz /pollard/home/kathleen/projects/AlleleAnalyzer/manuscript_analyses/sgRNA_comparison_scott_zhang/dat/pcsk9_first_five_exons_annots.h5 /pollard/home/kathleen/projects/AlleleAnalyzer/manuscript_analyses/sgRNA_comparison_scott_zhang/dat/pcsk9_first_five_exons.bed /pollard/data/projects/AlleleAnalyzer_data/pam_sites_hg19/ /pollard/data/vertebrate_genomes/human/hg19/hg19/hg19.fa /pollard/home/kathleen/projects/AlleleAnalyzer/manuscript_analyses/sgRNA_comparison_scott_zhang/dat/pcsk9_first_five_wtc SpCas9 20 -v --hom --bed --crispor=hg19`
Make BED for IGV
`python ~/projects/AlleleAnalyzer/scripts/make_pretty_igv.py pcsk9_first_five_wtc.tsv pcsk9_first_five_wtc_igv pcsk9_first_five_wtc`
```
aa_sgrnas_pcsk9_first_five = pd.read_csv('/pollard/home/kathleen/projects/AlleleAnalyzer/manuscript_analyses/sgRNA_comparison_scott_zhang/dat/pcsk9_first_five_wtc.tsv',
sep='\t')
aa_sgrnas_pcsk9_first_five['chrom'] = 'chr' + aa_sgrnas_pcsk9_first_five['chrom'].astype(str)
aa_sgrnas_pcsk9_first_five.head()
chroms = []
starts = []
stops = []
ids = []
scores = []
for ix, row in aa_sgrnas_pcsk9_first_five.iterrows():
if row['strand'] == 'positive':
chroms.append('chr1')
starts.append(row['start'])
stops.append(row['stop'] + 3)
ids.append(ix)
scores.append(row['scores_ref'])
else:
chroms.append('chr1')
starts.append(row['start'] - 3)
stops.append(row['stop'])
ids.append(ix)
scores.append(row['scores_ref'])
new_aa_bed_df = pd.DataFrame({
'chrom': chroms,
'start': starts,
'stop': stops,
'score': scores
}).drop_duplicates()
# new_aa_bed_df['size'] = new_aa_bed_df['stop'] - new_aa_bed_df['start']
new_aa_bed = BedTool.from_dataframe(new_aa_bed_df)
len(aa_sgrnas_pcsk9_first_five)
aa_sgrnas_pcsk9_first_five.head()
len(aa_sgrnas_pcsk9_first_five.query('scores_ref > 50'))
platinum_bed = BedTool('/pollard/home/kathleen/projects/AlleleAnalyzer/manuscript_analyses/sgRNA_comparison_scott_zhang/dat/pcsk9_platinum_targets_first_five_exons.bed')
# aa_bed = BedTool.from_dataframe(aa_sgrnas_pcsk9_first_five.sort_values(by='scores_ref',
# ascending=False)[['chrom','start','stop']].head(65))
aa_bed = BedTool.from_dataframe(aa_sgrnas_pcsk9_first_five[['chrom','start','stop','gRNAs','scores_ref']])
aa_bed.head()
len(platinum_bed.intersect(new_aa_bed, wb=True, f=1.0, u=True).to_dataframe())
aa_platinum = new_aa_bed.intersect(platinum_bed, u=True, f=1.0).to_dataframe().drop_duplicates()
len(aa_platinum)
aa_platinum['platinum'] = True
aa_platinum.to_csv('dat/new_aa_plat.bed',
sep='\t', index=False, header=False)
new_aa_bed_df.head()
aa_platinum.columns = ['chrom','start','stop','score','platinum']
aa_platinum.head()
aa_bed_df = new_aa_bed_df.merge(aa_platinum, how='outer', left_on=['chrom','start','stop','score'],
right_on=['chrom','start','stop','score']).replace(np.nan, False)
aa_bed_df
len(aa_bed_df.query('platinum'))
sns.swarmplot(x='platinum', y='score', data=aa_bed_df)
aa_bed_df.query('platinum')['score'].mean()
aa_bed_df.query('~platinum')['score'].mean()
pcsk9_spcas9_platinum_first_five.head()
platinum_beddf = platinum_bed.to_dataframe()
platinum_beddf.head()
platinum_bed_starts = platinum_beddf['start'].to_list()
platinum_bed_ends = platinum_beddf['end'].to_list()
len(aa_platinum.name.unique())
aa_sgrnas_pcsk9_first_five['platinum'] = aa_sgrnas_pcsk9_first_five['gRNAs'].isin(aa_platinum['name'])
aa_sgrnas_pcsk9_first_five = aa_sgrnas_pcsk9_first_five.query('(start in @platinum_bed_starts) or (stop in @platinum_bed_ends)').copy()
len(aa_sgrnas_pcsk9_first_five)
len(aa_sgrnas_pcsk9_first_five.query('platinum'))
len(platinum_bed)
aa_sgrnas_pcsk9_first_five.head()
len(platinum_bed)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/tbeucler/2022_ML_Earth_Env_Sci/blob/main/Lab_Notebooks/S4_1_Dimensionality.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
##**Chapter 8 – Dimensionality Reduction**
<img src='https://unils-my.sharepoint.com/:i:/g/personal/tom_beucler_unil_ch/EX7KlNGWYypLnH_53OnJR6oBjfgb_gCZ4gmnOeR68a6zMA?download=1'>
<center> Caption: <i>Denise diagnoses an overheated CPU at our data center in The Dalles, Oregon. <br> For more than a decade, we have built some of the world's most efficient servers.</i> <br> Photo from the <a href='https://www.google.com/about/datacenters/gallery/'>Google Data Center gallery</a> </center>
*Our world is increasingly filled with data from all sorts of sources, including environmental data. Can we reduce the data to a reduced, meaningful space to save on computation time and increase explainability?*
This notebook will be used in the lab session for week 4 of the course, covers Chapters 8 of Géron, and builds on the [notebooks made available on _Github_](https://github.com/ageron/handson-ml2).
Need a reminder of last week's labs? Click [_here_](https://colab.research.google.com/github/tbeucler/2022_ML_Earth_Env_Sci/blob/main/Lab_Notebooks/Week_3_Decision_Trees_Random_Forests_SVMs.ipynb) to go to notebook for week 3 of the course.
##Setup
First, let's import a few common modules, ensure MatplotLib plots figures inline and prepare a function to save the figures. We also check that Python 3.5 or later is installed (although Python 2.x may work, it is deprecated so we strongly recommend you use Python 3 instead), as well as Scikit-Learn ≥0.20.
```
# Python ≥3.5 is required
import sys
assert sys.version_info >= (3, 5)
# Scikit-Learn ≥0.20 is required
import sklearn
assert sklearn.__version__ >= "0.20"
# Common imports
import numpy as np
import os
# to make this notebook's output stable across runs
rnd_seed = 42
rnd_gen = np.random.default_rng(rnd_seed)
# To plot pretty figures
%matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rc('axes', labelsize=14)
mpl.rc('xtick', labelsize=12)
mpl.rc('ytick', labelsize=12)
# Where to save the figures
PROJECT_ROOT_DIR = "."
CHAPTER_ID = "dim_reduction"
IMAGES_PATH = os.path.join(PROJECT_ROOT_DIR, "images", CHAPTER_ID)
os.makedirs(IMAGES_PATH, exist_ok=True)
def save_fig(fig_id, tight_layout=True, fig_extension="png", resolution=300):
path = os.path.join(IMAGES_PATH, fig_id + "." + fig_extension)
print("Saving figure", fig_id)
if tight_layout:
plt.tight_layout()
plt.savefig(path, format=fig_extension, dpi=resolution)
```
## Dimensionality Reduction using PCA
This week we'll be looking at how to reduce the dimensionality of a large dataset in order to improve our classifying algorithm's performance! With that in mind, let's being the exercise by loading the MNIST dataset.
###**Q1) Load the input features and truth variable into X and y, then split the data into a training and test dataset using scikit's train_test_split method. Use *test_size=0.15*, and remember to set the random state to *rnd_seed!***
*Hint 1: The `'data'` and `'target'` keys for mnist will return X and y.*
*Hint 2: [Here's the documentation](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.train_test_split.html) for train/test split.*
```
# Load the mnist dataset
from sklearn.datasets import fetch_openml
mnist = fetch_openml('mnist_784', version=1, as_frame=False)
# Load X and y
X =
y =
# Import the train/test split function from sklearn
# Split the dataset
X_train, X_test, y_train, y_test =
```
We now once again have a training and testing dataset with which to work with. Let's try training a random forest tree classifier on it. You've had experience with them before, so let's have you import the `RandomForestClassifier` from sklearn and instantiate it.
###**Q2) Import the `RandomForestClassifier` model from sklearn. Then, instantiate it with 100 estimators and set the random state to `*rnd_seed!*`**
*Hint 1: [Here's the documentation](https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.RandomForestClassifier.html) for `RandomForestClassifier`*
*Hint 2: [Here's the documentation](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.train_test_split.html) for train/test split.*
*Hint 3: If you're still confused about **instantiation**, there's a [blurb on wikipedia](https://en.wikipedia.org/wiki/Instance_(computer_science)) describing it in the context of computer science.*
```
# Complete the code
from sklearn.______ import _______
rnd_clf = _____(______=______, #Number of estimators
______=______) #Random State
```
We're now going to measure how quickly the algorithm is fitted to the mnist dataset! To do this, we'll have to import the `time` library. With it, we'll be able to get a timestamp immediately before and after we fit the algorithm, and we'll get the time by calculating the difference.
###**Q3) Import the time library and calculate how long it takes to fit the `RandomForestClassifier` model.**
*Hint 1: [Here's the documentation](https://docs.python.org/3/library/time.html#time.time) to the function used for getting timestamps*
*Hint 2: [Here's the documentation](https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.RandomForestClassifier.html#sklearn.ensemble.RandomForestClassifier.fit) for the fitting method used in `RandomForestClassifier`.*
```
import time
t0 = _____._____() # Load the timestamp before running
rnd_clf.___(_____, _____) # Fit the model with the training data
t1 = _____._____() # Load the timestamp after running
train_t_rf = t1-t0
print(f"Training took {train_t_rf:.2f}s")
```
We care about more than just how long we took to trian the model, however! Let's get an accuracy score for our model.
###**Q4) Get an accuracy score for the predictions from the RandomForestClassifier**
*Hint 1: [Here is the documentation](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.accuracy_score.html) for the `accuracy_score` metric in sklearn.*
*Hint 2: [Here is the documentation](https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.RandomForestClassifier.html#sklearn.ensemble.RandomForestClassifier.predict) for the predict method in `RandomForestClassifier`*
```
from _____._____ import _____ # Import the accuracy score metric
# Get a set of predictions from the random forest classifier
y_pred = _____._____(_____) # Get a set of predictions from the test set
rf_accuracy = accuracy_score(_____, _____) # Feed in the truth and predictions
print(f"RF Model Accuracy: {rf_accuracy:.2%}")
```
Let's try doing the same with with a logistic regression algorithm to see how it compares.
###**Q5) Repeat Q2-4 with a logistic regression algorithm using sklearn's `LogisticRegression` class. Hyperparameters: `multi_class='multinomial'` and `solver='lbfgs'`**
*Hint 1: [Here is the documentation](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LogisticRegression.html) for the `LogisticRegression` class.
```
from _____._____ import _____
log_clf = _____(_____="multinomial", #Multiclass
_____="lbfgs", Solver
_____=42) #Random State
t0 = time.time() # Timestamp before training
log_clf.fit(_____, _____) # Fit the model with the training data
t1 = time.time() # Timestamp after training
train_t_log = t1-t0
print(f"Training took {train_t_log:.2f}s")
# Get a set of predictions from the logistric regression classifier
y_pred = _____._____(_____) # Get a set of predictions from the test set
log_accuracy = accuracy_score(_____, _____) # Feed in the truth and predictions
print(f"Log Model Accuracy: {log_accuracy:.2%}")
```
Up to now, everything that we've done are things we've done in previous labs - but now we'll get to try out some algorithms useful for reducing dimensionality! Let's use principal component analysis. Here, we'll reduce the space using enough axes to explain over 95% of the variability in the data...
###**Q6) Import scikit's implementation of `PCA` and fit it to the training dataset so that 95% of the variability is explained.**
*Hint 1: [Here is the documentation](https://scikit-learn.org/stable/modules/generated/sklearn.decomposition.PCA.html) for scikit's `PCA` class.*
*Hint 2: [Here is the documentation](https://scikit-learn.org/stable/modules/generated/sklearn.decomposition.PCA.html#sklearn.decomposition.PCA.fit_transform) for scikit's `.fit_transform()` method.*
```
from _____._____ import _____ # Importing PCA
pca = PCA(_____=_____) # Set number of components to explain 95% of variability
X_train_reduced = pca._____(____) # Fit-transform the training data
X_test_reduced = pca._____(____) # Transform the test data (!!No fitting!!)
```
###**Q7) Repeat Q3 & Q4 using the *reduced* `X_train` dataset instead of `X_train`.**
```
# Complete the code
t0 = _____._____() # Load the timestamp before running
rnd_clf.___(_____, _____) # Fit the model with the reduced training data
t1 = _____._____() # Load the timestamp after running
train_t_rf = t1-t0
print(f"Training took {train_t_rf:.2f}s")
# Get a set of predictions from the random forest classifier
y_pred = _____._____(_____) # Get predictions from the reduced test set
red_rf_accuracy = accuracy_score(_____, _____) # Feed in the truth and predictions
print(f"RF Model Accuracy on reduced dataset: {red_rf_accuracy:.2%}")
```
###**Q8) Repeat Q5 using the *reduced* X_train dataset instead of X_train.**
```
#Complete the code
t0 = time.time() # Timestamp before training
log_clf.fit(_____, _____) # Fit the model with the reduced training data
t1 = time.time() # Timestamp after training
train_t_log = t1-t0
print(f"Training took {train_t_log:.2f}s")
# Get a set of predictions from the logistric regression classifier
y_pred = _____._____(_____) # Get a set of predictions from the test set
log_accuracy = accuracy_score(_____, _____) # Feed in the truth and predictions
print(f"Log Model Accuracy on reduced training data: {log_accuracy:.2%}")
```
You can now compare how well the random forest classifier and logistic regression classifier performed on both the full dataset and the reduced dataset. What were you able to observe?
Write your comments on the performance of the algorithms in this box, if you'd like 😀
(Double click to activate editing mode)
| github_jupyter |
## Lesson 15 - Pandas Time Series
### Readings
* McKinney: [Chapter 11. Time Series](http://proquest.safaribooksonline.com/book/programming/python/9781491957653/time-series/timeseries_html)
### Table of Contents
* [Examples](#examples)
* [Timeseries classes](#classes)
* [Timestamps vs. Periods (time spans)](#timestamps-vs-periods)
* [Converting to Timestamps](#coverting-to-timestamps)
* [Generating ranges of timestamps](#ranges-of-timestamps)
* [DatetimeIndex](#datetimeindex)
* [DateOffset objects](#dateoffset-objects)
* [Time series-related instance methods](#instance-methods)
* [Time span representation](#time-spans)
* [Time zone handling](#time-zones)
*Adapted from Pandas documentation (esoteric/financial applications left out).*
In working with time series data, we will frequently seek to:
* generate sequences of fixed-frequency dates and time spans
* conform or convert time series to a particular frequency
* compute “relative” dates based on various non-standard time increments (e.g. 5 business days before the last business day of the year), or “roll” dates forward or backward
Pandas provides a relatively compact and self-contained set of tools for performing the above tasks.
```
# modules used in this tutorial
import pandas as pd
import numpy as np
import pytz
import dateutil
# set max row and random seed
pd.set_option("display.max_rows", 10)
np.random.seed(12)
```
<a id="examples"></a>
## Examples
Create a range of dates:
```
rng = pd.date_range('1/1/2011', periods=72, freq='H')
rng[:5]
len(rng)
```
Index Pandas objects with dates:
```
ts = pd.Series(np.random.randn(len(rng)), index=rng)
ts.head()
```
Change frequency and fill gaps:
```
# to 45 minute frequency and forward fill
converted = ts.asfreq('45Min', method='pad')
converted.head()
```
Frequency conversion and resampling of a time series:
```
ts.resample('D').mean()
ts.resample('T').mean().dropna()
```
Find difference between dates:
```
pd.Timestamp('11/19/18') - pd.Timestamp('1/1/18')
pd.Period('11/19/18') - pd.Period('1/1/18')
```
<a id="classes"></a>
## Timeseries classes
The following table shows the type of time-related classes pandas can handle and how to create them:
Class | Remarks | How to create
------|---------|--------------
`Timestamp` | Represents a single time stamp | `to_datetime`, `Timestamp`
`DatetimeIndex` | Index (~array) of `Timestamp` | `to_datetime`, `date_range`, `DatetimeIndex`
`Period` | Represents a single time span | `Period`
`PeriodIndex` | Index (~array) of `Period` | `period_range`, `PeriodIndex`
<a id="timestamps-vs-periods"></a>
## Timestamps vs. Periods (time spans)
Time-stamped data is the most basic type of timeseries data that associates values with points in time. For pandas objects it means using the points in time.
```
pd.datetime(2012, 5, 1)
pd.Timestamp(pd.datetime(2012, 5, 1))
pd.Timestamp('2012-05-01')
```
However, in many cases it is more natural to associate things like change variables with a time span instead. The span represented by Period can be specified explicitly, or inferred from datetime string format.
For example:
```
pd.Period('2011-01')
pd.Period('2012-05', freq='D')
```
Timestamp and Period can be the index. Lists of Timestamp and Period are automatically coerced to DatetimeIndex and PeriodIndex respectively.
```
dates = [pd.Timestamp('2012-05-01'), pd.Timestamp('2012-05-02'), pd.Timestamp('2012-05-03')]
ts = pd.Series(np.random.randn(3), dates)
ts
ts.index
type(ts.index)
periods = [pd.Period('2012-01'), pd.Period('2012-02'), pd.Period('2012-03')]
ts = pd.Series(np.random.randn(3), periods)
ts
ts.index
type(ts.index)
```
Pandas allows you to capture both representations and convert between them. Under the hood, Pandas represents timestamps using instances of Timestamp and sequences of timestamps using instances of DatetimeIndex. For regular time spans, Pandas uses Period objects for scalar values and PeriodIndex for sequences of spans.
<a id="coverting-to-timestamps"></a>
## Converting to Timestamps
To convert a Series or list-like object of date-like objects e.g. strings, epochs, or a mixture, you can use the to_datetime function. When passed a Series, this returns a Series (with the same index), while a list-like is converted to a DatetimeIndex:
```
pd.to_datetime(pd.Series(['Jul 31, 2009', '2010-01-10', None]))
pd.to_datetime(['2005/11/23', '2010.12.31'])
```
If you use dates which start with the day first (i.e. European style), you can pass the dayfirst flag:
```
pd.to_datetime(['04-01-2012']) # default behavior is dayfirst=False
pd.to_datetime(['04-01-2012'], dayfirst=False)
pd.to_datetime(['04-01-2012'], dayfirst=True)
```
**Warning:** If a date can’t be parsed with the day being first it will be parsed as if dayfirst were False:
```
pd.to_datetime(['04-13-2012'], dayfirst=True)
```
**Warning:** Even with a list of dates that are clearly European (day first), each date is processed separately. You must specify `dayfirst=True` (or a format argument) to get the desired behaviour:
```
pd.to_datetime(['10/1/18', '11/1/18', '12/1/18', '13/1/18', '14/1/18'])
pd.to_datetime(['10/1/18', '11/1/18', '12/1/18', '13/1/18', '14/1/18'], dayfirst=True)
```
**Note:** Specifying a format argument will potentially speed up the conversion considerably and on versions later than 0.13.0 explicitly specifying a format string of ‘%Y%m%d’ takes a faster path still.
If you pass a single string to to_datetime, it returns single Timestamp. Also, Timestamp can accept the string input. Note that Timestamp doesn’t accept string parsing option like dayfirst or format: use to_datetime if these are required.
The first option below would work with a list of dates, but the second would not.
```
pd.to_datetime('11/12/2010', format='%d/%m/%Y')
pd.to_datetime('11/12/2010', format='%m/%d/%Y')
pd.to_datetime('11/12/2010')
pd.Timestamp('2010-11-12')
```
### Invalid data
In version 0.17.0, the default for to_datetime is now errors='raise', rather than errors='ignore'. This means that invalid parsing will raise rather that return the original input as in previous versions.
Pass errors='coerce' to convert invalid data to NaT (not a time):
```
# don't convert anything and return the original input when unparseable
pd.to_datetime(['2009/07/31', 'asd'], errors='ignore')
# return NaT for input when unparseable
pd.to_datetime(['2009/07/31', 'asd'], errors='coerce')
```
### Epoch timestamps
It’s also possible to convert integer or float epoch times. The default unit for these is nanoseconds (since these are how Timestamps are stored). However, often epochs are stored in another unit which can be specified:
Typical epoch stored units (nanoseconds):
```
pd.to_datetime(0)
pd.to_datetime(1)
pd.to_datetime(3e9)
pd.to_datetime([1349720105, 1349806505, 1349892905, 1349979305, 1350065705], unit='s')
pd.to_datetime([1349720105100, 1349720105200, 1349720105300, 1349720105400, 1349720105500 ], unit='ms')
```
**Note:** Epoch times will be rounded to the nearest nanosecond.
<a id="ranges-of-timestamps"></a>
## Generating ranges of timestamps
To generate an index with time stamps, you can use either the DatetimeIndex or Index constructor and pass in a list of datetime objects
### The slow way
```
dates = [pd.datetime(2012, 5, 1), pd.datetime(2012, 5, 2), pd.datetime(2012, 5, 3)]
index = pd.DatetimeIndex(dates)
index # Note the frequency information
index = pd.Index(dates)
index # Automatically converted to DatetimeIndex
```
### The quick way - date_range by periods or start-end
Practically, this becomes very cumbersome because we often need a very long index with a large number of timestamps.
If we need timestamps on a regular frequency, we can use the pandas function date_range to create timestamp indexes. (A similar function and bdate_range does the same thing for business days; the 'b' stands for 'business'.)
Information needed to specify date ranges:
* start, number of periods, AND frequency
* start, end, AND frequency
Frequency defaults to 'D' for date_range and 'B' for bdate_range.
```
# freq='M' uses last day of month by default
index = pd.date_range('2018-3-1', periods=1000, freq='M')
index
# freq='B' is business days
index = pd.date_range('2018-3-1', periods=1000, freq='B')
index
# default is freq='D' for date_range
index = pd.date_range('2018-3-1', periods=1000)
index
# default is freq='B' for bdate_range
index = pd.bdate_range('2018-3-1', periods=1000)
index
```
### Commonly used of offset aliases
Alias | Description
------|------------
D | day frequency
B | business day frequency
W | weekly frequency
BW | business week end frequency
M | month end frequency
BM | business month end frequency
MS | month start frequency
A | year end frequency
BA | business year end frequency
AS | year start frequency
H | hourly frequency
T, min | minutely frequency
S | secondly frequency
L, ms | milliseonds
U, us | microseconds
N | nanoseconds
```
t1 = pd.datetime(2011, 1, 1)
t2 = pd.datetime(2012, 1, 1)
rng = pd.date_range(t1, t2, freq='D')
rng
pd.date_range(t1, t2, freq='W-SUN')
pd.date_range(t1, t2, freq='B')
pd.date_range(t1, t2, freq='BM')
pd.bdate_range(end=t2, periods=20)
pd.bdate_range(start=t1, periods=20)
```
<a id="datetimeindex"></a>
## DatetimeIndex
One of the main uses for DatetimeIndex is as an index for Pandas objects. The DatetimeIndex class contains many timeseries related optimizations:
* A large range of dates for various offsets are pre-computed and cached under the hood in order to make generating subsequent date ranges very fast (just have to grab a slice)
* Fast shifting using the shift and tshift method on pandas objects
* Unioning of overlapping DatetimeIndex objects with the same frequency is very fast (important for fast data alignment)
* Quick access to date fields via properties such as year, month, etc.
* Regularization functions like snap and very fast asof logic
DatetimeIndex objects has all the basic functionality of regular Index objects and a smorgasbord of advanced timeseries-specific methods for easy frequency processing.
See also documentation for *Reindexing methods*.
**Note:** While Pandas does not force you to have a sorted date index, some of these methods may have unexpected or incorrect behavior if the dates are unsorted. So please be careful.
DatetimeIndex can be used like a regular index and offers all of its intelligent functionality like selection, slicing, etc.
```
start = pd.datetime(2011, 1, 1)
end = pd.datetime(2012, 1, 1)
rng = pd.date_range(start, end, freq='BM')
ts = pd.Series(np.random.randn(len(rng)), index=rng)
ts.index
ts[:5].index
ts[::2].index
```
### Slicing revisited
In Pandas, **label-based slicing is inclusive**, whereas **index-based slicing is NOT inclusive** (like index-based slicing in Python generally).
```
x = pd.Series([0.0, 0.1, 0.2], index=['a', 'b', 'c'])
x
x[0:2]
x['a':'c']
```
DatetimeIndex-based slicing is effectively label-based, so it is inclusive. It will not generate any dates outside of those dates if specified.
### DatetimeIndex partial string 'smart' indexing
You can pass in dates and strings that parse to dates as indexing parameters:
```
ts
ts['1/31/2011']
ts[pd.datetime(2011, 12, 25):]
ts['10/1/2011':'12/31/2011']
```
To provide convenience for accessing longer time series, you can also pass in the year or year and month as strings:
```
ts['2011']
ts['2011-6']
```
If we only want timestamps that match a certain day of the month (e.g., 31):
```
x = pd.Timestamp('10/31/2011')
x
x.day
ts[[(x.day == 31) for x in ts.index]]
ts[ts.index.day == 31]
```
This type of slicing will work on a DataFrame with a DateTimeIndex as well. Since the partial string selection is a form of label slicing, the endpoints will be included. This would include matching times on an included date. Here’s an example:
```
dft = pd.DataFrame(np.random.randn(100000,1), columns=['A'],
index=pd.date_range('20130101', periods=100000, freq='T'))
dft.shape
dft
dft['2013']
```
This starts on the very first time in the month, and includes the last date & time for the month:
```
dft['2013-1':'2013-2']
```
This specifies a stop time that includes all of the times on the last day:
```
dft['2013-1':'2013-2-28']
```
This specifies an exact stop time (and is not the same as the above):
```
dft['2013-1':'2013-2-28 00:00:00']
```
We are stopping on the included end-point as it is part of the index:
```
dft['2013-1-15':'2013-1-15 12:30:00']
```
To select a single row, use .loc:
```
dft.loc['2013-1-15 12:30:00']
```
### Datetime indexing
Indexing a DateTimeIndex with a partial string depends on the “accuracy” of the period, in other words how specific the interval is in relation to the frequency of the index. In contrast, indexing with datetime objects is exact, because the objects have exact meaning. These also follow the semantics of including both endpoints.
These datetime objects are specific hours, minutes, and seconds even though they were not explicitly specified (they are 0).
```
dft[pd.datetime(2013, 1, 1):pd.datetime(2013, 2, 28)]
```
With no defaults:
```
dft[pd.datetime(2013, 1, 1, 10, 12, 0):pd.datetime(2013, 2, 28, 10, 12, 0)]
```
### Truncating & fancy indexing
A truncate convenience function is provided that is equivalent to slicing:
```
ts.truncate(before='10/31/2011', after='12/31/2011')
```
Even complicated fancy indexing that breaks the DatetimeIndex’s frequency regularity will result in a DatetimeIndex (but frequency is lost):
```
ts[[0, 2, 6]].index
```
### Time/Date Components
There are several time/date properties that one can access from Timestamp or a collection of timestamps like a DateTimeIndex.
Property | Description
---------|------------
year | The year of the datetime
month | The month of the datetime
day | The days of the datetime
hour | The hour of the datetime
minute | The minutes of the datetime
second | The seconds of the datetime
microsecond | The microseconds of the datetime
nanosecond | The nanoseconds of the datetime
date | Returns datetime.date
time | Returns datetime.time
dayofyear | The ordinal day of year
weekofyear | The week ordinal of the year
week | The week ordinal of the year
dayofweek | The day of the week with Monday=0, Sunday=6
weekday | The day of the week with Monday=0, Sunday=6
quarter | Quarter of the date: Jan=Mar = 1, Apr-Jun = 2, etc.
days_in_month | The number of days in the month of the datetime
is_month_start | Logical indicating if first day of month (defined by frequency)
is_month_end | Logical indicating if last day of month (defined by frequency)
is_quarter_start | Logical indicating if first day of quarter (defined by frequency)
is_quarter_end | Logical indicating if last day of quarter (defined by frequency)
is_year_start | Logical indicating if first day of year (defined by frequency)
is_year_end | Logical indicating if last day of year (defined by frequency)
Furthermore, if you have a Series with datetimelike values, then you can access these properties via the .dt accessor, see the docs.
```
ts
ts.index.month
```
<a id="dateoffset-objects"></a>
## DateOffset objects
In the preceding examples, we created DatetimeIndex objects at various frequencies by passing in frequency strings like 'M', 'W', and 'BM' to the freq keyword. Under the hood, these frequency strings are being translated into an instance of pandas DateOffset, which represents a regular frequency increment. Specific offset logic like “month”, “business day”, or “one hour” is represented in its various subclasses.
Class name | Description
-----------|------------
DateOffset | Generic offset class, defaults to 1 calendar day
BDay | business day (weekday)
CDay | custom business day (experimental)
Week | one week, optionally anchored on a day of the week
WeekOfMonth | the x-th day of the y-th week of each month
LastWeekOfMonth | the x-th day of the last week of each month
MonthEnd | calendar month end
MonthBegin | calendar month begin
BMonthEnd | business month end
BMonthBegin | business month begin
CBMonthEnd | custom business month end
CBMonthBegin | custom business month begin
QuarterEnd | calendar quarter end
QuarterBegin | calendar quarter begin
BQuarterEnd | business quarter end
BQuarterBegin | business quarter begin
FY5253Quarter | retail (aka 52-53 week) quarter
YearEnd | calendar year end
YearBegin | calendar year begin
BYearEnd | business year end
BYearBegin | business year begin
FY5253 | retail (aka 52-53 week) year
BusinessHour | business hour
Hour | one hour
Minute | one minute
Second | one second
Milli | one millisecond
Micro | one microsecond
Nano | one nanosecond
Basic function of DateOffset:
```
d = pd.datetime(2018, 2, 27, 9, 0)
d
pd.Timestamp(d)
d + pd.tseries.offsets.DateOffset(months=2, days=1)
```
The key features of a DateOffset object are:
* it can be added / subtracted to/from a datetime object to obtain a shifted date
* it can be multiplied by an integer (positive or negative) so that the increment will be applied multiple times
* it has rollforward and rollback methods for moving a date forward or backward to the next or previous “offset date”
Subclasses of DateOffset define the apply function which dictates custom date increment logic, such as adding business days:
class BDay(DateOffset):
"""DateOffset increments between business days"""
def apply(self, other):
...
```
d - 5 * pd.tseries.offsets.BDay()
d + pd.tseries.offsets.BMonthEnd()
```
The rollforward and rollback methods do exactly what you would expect:
```
d
offset = pd.tseries.offsets.BMonthEnd()
offset.rollforward(d)
offset.rollback(d)
```
If you expect to use these functions, explore the pandas.tseries.offsets module and the various docstrings for the classes.
These operations (apply, rollforward and rollback) preserves time (hour, minute, etc) information by default. To reset time, use normalize=True keyword when creating the offset instance. If normalize=True, result is normalized after the function is applied.
```
day = pd.tseries.offsets.Day()
day.apply(pd.Timestamp('2014-01-01 09:00'))
day = pd.tseries.offsets.Day(normalize=True)
day.apply(pd.Timestamp('2014-01-01 09:00'))
hour = pd.tseries.offsets.Hour()
hour.apply(pd.Timestamp('2014-01-01 22:00'))
pd.Timestamp('2014-01-01 23:00:00')
hour = pd.tseries.offsets.Hour(normalize=True)
hour.apply(pd.Timestamp('2014-01-01 22:00'))
hour.apply(pd.Timestamp('2014-01-01 23:00'))
```
### Parametric offsets
Some of the offsets can be “parameterized” when created to result in different behaviors. For example, the Week offset for generating weekly data accepts a weekday parameter which results in the generated dates always lying on a particular day of the week:
```
d
d + pd.tseries.offsets.Week()
d + pd.tseries.offsets.Week(weekday=4)
(d + pd.tseries.offsets.Week(weekday=4)).weekday()
d - pd.tseries.offsets.Week()
```
normalize option will be effective for addition and subtraction.
```
d + pd.tseries.offsets.Week(normalize=True)
d - pd.tseries.offsets.Week(normalize=True)
```
Another example is parameterizing YearEnd with the specific ending month:
```
d + pd.tseries.offsets.YearEnd()
d + pd.tseries.offsets.YearEnd(month=6)
```
### Using offsets with Series / DatetimeIndex
Offsets can be used with either a Series or DatetimeIndex to apply the offset to each element.
```
rng = pd.date_range('2012-01-01', '2012-01-03')
s = pd.Series(rng)
rng
rng + pd.tseries.offsets.DateOffset(months=2)
s + pd.tseries.offsets.DateOffset(months=2)
s - pd.tseries.offsets.DateOffset(months=2)
```
If the offset class maps directly to a Timedelta (Day, Hour, Minute, Second, Micro, Milli, Nano) it can be used exactly like a Timedelta - see the Timedelta section for more examples.
```
s - pd.tseries.offsets.Day(2)
td = s - pd.Series(pd.date_range('2011-12-29', '2011-12-31'))
td
td + pd.tseries.offsets.Minute(15)
```
Note that some offsets (such as BQuarterEnd) do not have a vectorized implementation. They can still be used but may calculate signficantly slower and will raise a PerformanceWarning.
```
rng + pd.tseries.offsets.BQuarterEnd()
```
### Offset Aliases
A number of string aliases are given to useful common time series frequencies. We will refer to these aliases as offset aliases (referred to as time rules prior to v0.8.0).
Alias | Description
------|------------
B | business day frequency
C | custom business day frequency (experimental)
D | calendar day frequency
W | weekly frequency
M | month end frequency
BM | business month end frequency
CBM | custom business month end frequency
MS | month start frequency
BMS | business month start frequency
CBMS | custom business month start frequency
Q | quarter end frequency
BQ | business quarter endfrequency
QS | quarter start frequency
BQS | business quarter start frequency
A | year end frequency
BA | business year end frequency
AS | year start frequency
BAS | business year start frequency
BH | business hour frequency
H | hourly frequency
T, min | minutely frequency
S | secondly frequency
L, ms | milliseonds
U, us | microseconds
N | nanoseconds
### Combining Aliases
As we have seen previously, the alias and the offset instance are fungible in most functions:
```
pd.date_range(start, periods=5, freq='B')
pd.date_range(start, periods=5, freq=pd.tseries.offsets.BDay())
```
You can combine together day and intraday offsets:
```
pd.date_range(start, periods=10, freq='2h20min')
pd.date_range(start, periods=10, freq='1D10U')
```
<a id="instance-methods"></a>
## Time series-related instance methods
### Shifting / lagging
One may want to shift or lag the values in a time series back and forward in time. The method for this is shift, which is available on all of the pandas objects.
```
ts = pd.Series(['a', 'b', 'c', 'd', 'e'], index=pd.date_range(start='3/1/18', periods=5))
ts
ts.shift(3)
```
Notes: If freq is specified then the index values are shifted but the data
is not realigned. That is, use freq if you would like to extend the
index when shifting and preserve the original data.
```
ts.shift(3, freq='D')
ts.shift(3, freq='M')
```
The same thing can be accomplisehd with the tshift convenience method that changes all the dates in the index by a specified number of offsets, rather than changing the alignment of the data and the index:
```
ts.tshift(3)
```
Note that with tshift (and with shift+freq), the leading entry is no longer NaN because the data is not being realigned.
### Frequency conversion
The primary function for changing frequencies is the asfreq function. For a DatetimeIndex, this is a convenient wrapper around reindex which generates a date_range and calls reindex.
```
dr = pd.date_range('1/1/2010', periods=3, freq='3B')
ts = pd.Series(np.random.randn(3), index=dr)
ts
ts.asfreq('B')
```
asfreq provides a further convenience so you can specify an interpolation method (e.g. 'pad', i.e. forward fill) for any gaps that may appear after the frequency conversion.
```
ts.asfreq(pd.tseries.offsets.BDay(), method='pad')
```
### Filling forward / backward
Related to asfreq and reindex is the fillna function, which can be used with any Pandas object.
```
ts.asfreq('B').fillna(0)
```
<a id="time-spans"></a>
## Time span representation
Regular intervals of time are represented by Period objects in pandas while sequences of Period objects are collected in a PeriodIndex, which can be created with the convenience function period_range.
### Period
A Period represents a span of time (e.g., a day, a month, a quarter, etc). You can specify the span via freq keyword using a frequency alias like below. Because freq represents a span of Period, it cannot be negative like "-3D". ('A-DEC' is annual frequency, anchored end of December; same as 'A'.)
```
pd.Period('2012', freq='A')
pd.Period('2012-1-1', freq='D')
pd.Period('2012-1-1 19:00', freq='H')
pd.Period('2012-1-1 19:00', freq='5H')
```
Adding and subtracting integers from periods shifts the period by its own frequency. Arithmetic is not allowed between Period with different freq (span).
```
p = pd.Period('2012', freq='A')
p + 1
p - 3
p = pd.Period('2012-01', freq='2M')
p + 2
p - 1
```
If Period freq is daily or higher (D, H, T, S, L, U, N), offsets and timedelta-like can be added if the result can have the same freq.
```
p = pd.Period('2014-07-01 09:00', freq='H')
p + pd.tseries.offsets.Hour(2)
p + pd.tseries.offsets.timedelta(minutes=120)
p + np.timedelta64(7200, 's')
```
If Period has other freqs, only the same offsets can be added.
```
p = pd.Period('2014-07', freq='M')
p + pd.tseries.offsets.MonthEnd(3)
pd.Period('2012', freq='A') - pd.Period('2002', freq='A')
```
### Periods vs. Timestamps
Notice the difference between Timestamps and Periods in determining differences in times.
```
pd.datetime.today() - pd.Timestamp('1/1/1970')
pd.Period(pd.datetime.today(), 'D') - pd.Period('1/1/1970', 'D')
pd.Period(pd.datetime.today(), 'A') - pd.Period('1/1/1970', 'A')
t = pd.datetime.today() - pd.Timestamp('1/1/1970')
t
t.days
```
### PeriodIndex and period_range
Regular sequences of Period objects can be collected in a PeriodIndex, which can be constructed using the period_range convenience function:
```
prng = pd.period_range('1/1/2011', '1/1/2012', freq='M')
prng
```
The PeriodIndex constructor can also be used directly:
```
pd.PeriodIndex(['2011-1', '2011-2', '2011-3'], freq='M')
```
Passing multiplied frequency outputs a sequence of Period which has multiplied span.
```
pd.PeriodIndex(start='2014-01', freq='3M', periods=4)
```
Just like DatetimeIndex, a PeriodIndex can also be used to index pandas objects:
```
ps = pd.Series(np.random.randn(len(prng)), prng)
ps
```
PeriodIndex supports addition and subtraction with the same rule as Period.
```
idx = pd.period_range('2014-07-01 09:00', periods=5, freq='H')
idx
idx + pd.tseries.offsets.Hour(2)
idx = pd.period_range('2014-07', periods=5, freq='M')
idx
idx + pd.tseries.offsets.MonthEnd(3)
```
### PeriodIndex partial string indexing
You can pass in dates and strings to Series and DataFrame with PeriodIndex, in the same manner as DatetimeIndex. For details, refer to DatetimeIndex Partial String Indexing.
```
ps
ps['2011-01']
ps[pd.datetime(2011, 12, 25):]
ps['10/31/2011':'12/31/2011']
```
Passing a string representing a lower frequency than PeriodIndex returns partial sliced data.
```
ps['2011']
dfp = pd.DataFrame(np.random.randn(600,1), columns=['A'], index=pd.period_range('2013-01-01 9:00',
periods=600, freq='T'))
dfp
dfp['2013-01-01 10H']
```
As with DatetimeIndex, the endpoints will be included in the result. The example below slices data starting from 10:00 to 11:59.
```
dfp['2013-01-01 10H':'2013-01-01 11H']
```
### Frequency conversion and resampling with PeriodIndex
The frequency of Period and PeriodIndex can be converted via the asfreq method. Let’s start with the fiscal year 2011, ending in December:
```
p = pd.Period('2011', freq='A-DEC')
p
```
We can convert it to a monthly frequency. Using the how parameter, we can specify whether to return the starting or ending month:
```
p.asfreq('M', how='start')
p.asfreq('M', how='end')
```
### Converting between representations
Timestamped data can be converted to PeriodIndex-ed data using to_period and vice-versa using to_timestamp:
```
rng = pd.date_range('1/1/2012', periods=5, freq='M')
ts = pd.Series(np.random.randn(len(rng)), index=rng)
ts
ts.index
ps = ts.to_period()
ps
ps.index
ps.to_timestamp()
ps.to_timestamp('D', how='start')
ps.to_timestamp('D', how='end')
```
<a id="time-zones"></a>
## Time zone handling
Pandas provides rich support for working with timestamps in different time zones using pytz and dateutil libraries. dateutil support is new in 0.14.1 and currently only supported for fixed offset and tzfile zones. The default library is pytz. Support for dateutil is provided for compatibility with other applications e.g. if you use dateutil in other python packages.
### Working with time zones
By default, pandas objects are time zone unaware:
```
rng = pd.date_range('3/6/2012 00:00', periods=15, freq='D')
rng.tz is None
```
To supply the time zone, you can use the tz keyword to date_range and other functions. There are two options for time zone formats:
* pytz - pytz brings the Olson tz database into Python. This library allows accurate and cross platform timezone calculations using Python 2.4 or higher. You can find a list of common (and less common) time zones using from pytz import common_timezones, all_timezones. For UTC, there is a special case tzutc.
* dateutil - The dateutil module provides powerful extensions to the datetime module available in the Python standard library. It uses the OS timezones so there isn’t a fixed list available. For common zones, the names are the same as pytz. Dateutil time zone strings are distinguished from pytz time zones by starting with "dateutil/".
```
# pytz
rng_pytz = pd.date_range('3/6/2012 00:00', periods=10, freq='D', tz='Europe/London')
rng_pytz.tz
# dateutil
rng_dateutil = pd.date_range('3/6/2012 00:00', periods=10, freq='D', tz='dateutil/Europe/London')
rng_dateutil.tz
# dateutil - utc special case
rng_utc = pd.date_range('3/6/2012 00:00', periods=10, freq='D', tz=dateutil.tz.tzutc())
rng_utc.tz
```
Note that the UTC timezone is a special case in dateutil and should be constructed explicitly as an instance of dateutil.tz.tzutc. You can also construct other timezones explicitly first, which gives you more control over which time zone is used:
```
# pytz
tz_pytz = pytz.timezone('Europe/London')
rng_pytz = pd.date_range('3/6/2012 00:00', periods=10, freq='D', tz=tz_pytz)
rng_pytz.tz == tz_pytz
# dateutil
tz_dateutil = dateutil.tz.gettz('Europe/London')
rng_dateutil = pd.date_range('3/6/2012 00:00', periods=10, freq='D', tz=tz_dateutil)
rng_dateutil.tz == tz_dateutil
```
Timestamps, like Python’s datetime.datetime object can be either time zone naive or time zone aware. Naive time series and DatetimeIndex objects can be localized using tz_localize:
```
ts = pd.Series(np.random.randn(len(rng)), rng)
ts
ts_utc = ts.tz_localize('UTC')
ts_utc
```
Again, you can explicitly construct the timezone object first. You can use the tz_convert method to convert pandas objects to convert tz-aware data to another time zone:
```
ts_utc.tz_convert('US/Eastern')
```
Warnings:
* Be wary of conversions between libraries. For some zones pytz and dateutil have different definitions of the zone. This is more of a problem for unusual timezones than for ‘standard’ zones like US/Eastern.
* Be aware that a timezone definition across versions of timezone libraries may not be considered equal. This may cause problems when working with stored data that is localized using one version and operated on with a different version. See here for how to handle such a situation.
* It is incorrect to pass a timezone directly into the datetime.datetime constructor (e.g., datetime.datetime(2011, 1, 1, tz=timezone('US/Eastern')). Instead, the datetime needs to be localized using the the localize method on the timezone.
Under the hood, all timestamps are stored in UTC. Scalar values from a DatetimeIndex with a time zone will have their fields (day, hour, minute) localized to the time zone. However, timestamps with the same UTC value are still considered to be equal even if they are in different time zones:
```
rng_eastern = rng_utc.tz_convert('US/Eastern')
rng_berlin = rng_utc.tz_convert('Europe/Berlin')
rng_eastern[5]
rng_berlin[5]
rng_eastern[5] == rng_berlin[5]
```
Like Series, DataFrame, and DatetimeIndex, Timestamps can be converted to other time zones using tz_convert:
```
rng_eastern[5]
rng_berlin[5]
rng_eastern[5].tz_convert('Europe/Berlin')
```
Localization of Timestamps functions just like DatetimeIndex and Series:
```
rng[5]
rng[5].tz_localize('Asia/Shanghai')
```
Operations between Series in different time zones will yield UTC Series, aligning the data on the UTC timestamps:
```
eastern = ts_utc.tz_convert('US/Eastern')
berlin = ts_utc.tz_convert('Europe/Berlin')
result = eastern + berlin
result
result.index
```
### Other time zones
List all the time zones available from pytz:
```
for tz in pytz.all_timezones:
print(tz)
```
For example, Berlin is GMT+1 (flip the sign):
```
rng.tz_localize('Europe/Berlin')
rng.tz_localize('Etc/GMT-1')
```
We can check timezones by converting the current time 'now' to another time zone:
```
pd.Timestamp('now')
pd.Timestamp('now', tz='US/Eastern')
pd.Timestamp('now', tz='UTC')
pd.Timestamp('now', tz='Asia/Riyadh')
pd.Timestamp('now', tz='Etc/GMT-3')
```
### Example: Day of week corresponding to a future date
```
def future_day_of_week(date, years_offset):
# dictionary to rename day of week
day_dict = {0: 'Monday', 1: 'Tuesday', 2: 'Wednesday', 3: 'Thursday',
4: 'Friday', 5: 'Saturday', 6: 'Sunday'}
# covert date to timestamp
ts = pd.Timestamp(date)
# offset date by X years
ts_offset = ts + pd.tseries.offsets.DateOffset(years=years_offset)
# print with strftime to make date pretty
print('%s in %s years will be %s which is a %s.' % (
ts.strftime('%Y-%m-%d'),
years_offset,
ts_offset.strftime('%Y-%m-%d'),
day_dict[ts_offset.dayofweek]))
future_day_of_week('1/1/00', 21)
# days from today until a given day
pd.Period('12/25/18', 'D') - pd.Period(pd.datetime.today(), 'D')
```
| github_jupyter |
# 实战 Kaggle 比赛:预测房价
实现几个函数来方便下载数据
```
import sys
sys.path.append('..')
import hashlib
import os
import tarfile
import zipfile
import requests
DATA_HUB = dict()
DATA_URL = 'http://d2l-data.s3-accelerate.amazonaws.com/'
def download(name, cache_dir=os.path.join('..', 'data')):
"""下载一个DATA_HUB中的文件,返回本地文件名。"""
assert name in DATA_HUB, f"{name} 不存在于 {DATA_HUB}."
url, sha1_hash = DATA_HUB[name]
os.makedirs(cache_dir, exist_ok=True)
fname = os.path.join(cache_dir, url.split('/')[-1])
if os.path.exists(fname):
sha1 = hashlib.sha1()
with open(fname, 'rb') as f:
while True:
data = f.read(1048576)
if not data:
break
sha1.update(data)
if sha1.hexdigest() == sha1_hash:
return fname
print(f'正在从{url}下载{fname}...')
r = requests.get(url, stream=True, verify=True)
with open(fname, 'wb') as f:
f.write(r.content)
return fname
def download_extract(name, folder=None):
"""下载并解压zip/tar文件。"""
fname = download(name)
base_dir = os.path.dirname(fname)
data_dir, ext = os.path.splitext(fname)
if ext == '.zip':
fp = zipfile.ZipFile(fname, 'r')
elif ext in ('.tar', '.gz'):
fp = tarfile.open(fname, 'r')
else:
assert False, '只有zip/tar文件可以被解压缩。'
fp.extractall(base_dir)
return os.path.join(base_dir, folder) if folder else data_dir
def download_all():
"""下载DATA_HUB中的所有文件。"""
for name in DATA_HUB:
download(name)
```
使用`pandas`读入并处理数据
```
%matplotlib inline
import numpy as np
import pandas as pd
import mindspore
from mindspore import nn
import mindspore.ops as ops
import mindspore.numpy as mnp
from d2l import mindspore as d2l
from mindspore import Tensor
DATA_HUB['kaggle_house_train'] = (
DATA_URL + 'kaggle_house_pred_train.csv',
'585e9cc93e70b39160e7921475f9bcd7d31219ce')
DATA_HUB['kaggle_house_test'] = (
DATA_URL + 'kaggle_house_pred_test.csv',
'fa19780a7b011d9b009e8bff8e99922a8ee2eb90')
train_data = pd.read_csv(download('kaggle_house_train'))
test_data = pd.read_csv(download('kaggle_house_test'))
print(train_data.shape)
print(test_data.shape)
```
前四个和最后两个特征,以及相应标签
```
print(train_data.iloc[0:4, [0, 1, 2, 3, -3, -2, -1]])
```
在每个样本中,第一个特征是ID,
我们将其从数据集中删除
```
all_features = pd.concat((train_data.iloc[:, 1:-1], test_data.iloc[:, 1:]))
```
将所有缺失的值替换为相应特征的平均值。
通过将特征重新缩放到零均值和单位方差来标准化数据
```
#若无法获得测试数据,则可根据训练数据计算均值和标准差
numeric_features = all_features.dtypes[all_features.dtypes != 'object'].index
all_features[numeric_features] = all_features[numeric_features].apply(
lambda x: (x - x.mean()) / (x.std()))
all_features[numeric_features] = all_features[numeric_features].fillna(0)
```
处理离散值。
我们用独热编码替换它们
```
all_features = pd.get_dummies(all_features, dummy_na=True)
all_features.shape
```
从`pandas`格式中提取NumPy格式
```
n_train = train_data.shape[0]
train_features = all_features[:n_train].values
test_features = all_features[n_train:].values
train_labels = train_data.SalePrice.values.reshape(-1, 1)
```
训练
```
mse_loss = nn.MSELoss()
in_features = train_features.shape[1]
def get_net():
net = nn.SequentialCell([nn.Dense(in_features,1)])
return net
```
我们更关心相对误差$\frac{y - \hat{y}}{y}$,
解决这个问题的一种方法是用价格预测的对数来衡量差异
```
def log_rmse(net, features, labels):
features = Tensor(features, mindspore.float32)
labels = Tensor(labels, mindspore.float32)
clipped_preds = ops.clip_by_value(net(features), Tensor(1, mindspore.float32), Tensor(float('inf'), mindspore.float32))
rmse = mnp.sqrt(mse_loss(mnp.log(clipped_preds),
mnp.log(labels)))
return rmse.asnumpy()
```
我们的训练函数将借助Adam优化器
```
def train(net, train_features, train_labels, test_features, test_labels,
num_epochs, learning_rate, weight_decay, batch_size):
train_ls, test_ls = [], []
train_dataset = d2l.load_array((train_features, train_labels), batch_size)
optim = nn.Adam(net.trainable_params(), learning_rate = learning_rate, weight_decay = weight_decay)
net_with_mse_loss = nn.WithLossCell(net, mse_loss)
train = nn.TrainOneStepCell(net_with_mse_loss, optim)
for epoch in range(num_epochs):
for X, y in train_dataset.create_tuple_iterator():
l = train(X, y)
train_ls.append(log_rmse(net, train_features, train_labels))
if test_labels is not None:
test_ls.append(log_rmse(net, test_features, test_labels))
return train_ls, test_ls
```
K折交叉验证
```
def get_k_fold_data(k, i, X, y):
assert k > 1
fold_size = X.shape[0] // k
X_train, y_train = None, None
for j in range(k):
idx = slice(j * fold_size, (j + 1) * fold_size)
X_part, y_part = X[idx, :], y[idx]
if j == i:
X_valid, y_valid = X_part, y_part
elif X_train is None:
X_train, y_train = X_part, y_part
else:
X_train = np.concatenate([X_train, X_part], 0)
y_train = np.concatenate([y_train, y_part], 0)
return X_train, y_train, X_valid, y_valid
```
返回训练和验证误差的平均值
```
def k_fold(k, X_train, y_train, num_epochs, learning_rate, weight_decay,
batch_size):
train_l_sum, valid_l_sum = 0, 0
for i in range(k):
data = get_k_fold_data(k, i, X_train, y_train)
net = get_net()
train_ls, valid_ls = train(net, *data, num_epochs, learning_rate,
weight_decay, batch_size)
train_l_sum += train_ls[-1]
valid_l_sum += valid_ls[-1]
if i == 0:
d2l.plot(list(range(1, num_epochs + 1)), [train_ls, valid_ls],
xlabel='epoch', ylabel='rmse', xlim=[1, num_epochs],
legend=['train', 'valid'], yscale='log')
print(f'fold {i + 1}, train log rmse {float(train_ls[-1]):f}, '
f'valid log rmse {float(valid_ls[-1]):f}')
return train_l_sum / k, valid_l_sum / k
```
模型选择
```
k, num_epochs, lr, weight_decay, batch_size = 5, 100, 5, 0, 64
train_l, valid_l = k_fold(k, train_features, train_labels, num_epochs, lr,
weight_decay, batch_size)
print(f'{k}-折验证: 平均训练log rmse: {float(train_l):f}, '
f'平均验证log rmse: {float(valid_l):f}')
```
提交你的Kaggle预测
```
def train_and_pred(train_features, test_feature, train_labels, test_data,
num_epochs, lr, weight_decay, batch_size):
net = get_net()
train_ls, _ = train(net, train_features, train_labels, None, None,
num_epochs, lr, weight_decay, batch_size)
d2l.plot(np.arange(1, num_epochs + 1), [train_ls], xlabel='epoch',
ylabel='log rmse', xlim=[1, num_epochs], yscale='log')
print(f'train log rmse {float(train_ls[-1]):f}')
preds = net(Tensor(test_features, mindspore.float32)).asnumpy()
test_data['SalePrice'] = pd.Series(preds.reshape(1, -1)[0])
submission = pd.concat([test_data['Id'], test_data['SalePrice']], axis=1)
submission.to_csv('../data/submission.csv', index=False)
train_and_pred(train_features, test_features, train_labels, test_data,
num_epochs, lr, weight_decay, batch_size)
```
| github_jupyter |
## Dependencies
```
import os
import cv2
import shutil
import random
import warnings
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.utils import class_weight
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix, cohen_kappa_score
from keras import backend as K
from keras.models import Model
from keras.utils import to_categorical
from keras import optimizers, applications
from keras.preprocessing.image import ImageDataGenerator
from keras.callbacks import EarlyStopping, ReduceLROnPlateau, Callback, LearningRateScheduler
from keras.layers import Dense, Dropout, GlobalAveragePooling2D, Input
# Set seeds to make the experiment more reproducible.
from tensorflow import set_random_seed
def seed_everything(seed=0):
random.seed(seed)
os.environ['PYTHONHASHSEED'] = str(seed)
np.random.seed(seed)
set_random_seed(0)
seed = 0
seed_everything(seed)
%matplotlib inline
sns.set(style="whitegrid")
warnings.filterwarnings("ignore")
```
## Load data
```
hold_out_set = pd.read_csv('../input/aptos-data-split/hold-out.csv')
X_train = hold_out_set[hold_out_set['set'] == 'train']
X_val = hold_out_set[hold_out_set['set'] == 'validation']
test = pd.read_csv('../input/aptos2019-blindness-detection/test.csv')
print('Number of train samples: ', X_train.shape[0])
print('Number of validation samples: ', X_val.shape[0])
print('Number of test samples: ', test.shape[0])
# Preprocecss data
X_train["id_code"] = X_train["id_code"].apply(lambda x: x + ".png")
X_val["id_code"] = X_val["id_code"].apply(lambda x: x + ".png")
test["id_code"] = test["id_code"].apply(lambda x: x + ".png")
X_train['diagnosis'] = X_train['diagnosis']
X_val['diagnosis'] = X_val['diagnosis']
display(X_train.head())
```
# Model parameters
```
# Model parameters
BATCH_SIZE = 32
EPOCHS = 40
WARMUP_EPOCHS = 5
LEARNING_RATE = 1e-4
WARMUP_LEARNING_RATE = 1e-3
HEIGHT = 224
WIDTH = 224
CHANNELS = 3
ES_PATIENCE = 5
RLROP_PATIENCE = 3
DECAY_DROP = 0.5
```
# Pre-procecess images
```
train_base_path = '../input/aptos2019-blindness-detection/train_images/'
test_base_path = '../input/aptos2019-blindness-detection/test_images/'
train_dest_path = 'base_dir/train_images/'
validation_dest_path = 'base_dir/validation_images/'
test_dest_path = 'base_dir/test_images/'
# Making sure directories don't exist
if os.path.exists(train_dest_path):
shutil.rmtree(train_dest_path)
if os.path.exists(validation_dest_path):
shutil.rmtree(validation_dest_path)
if os.path.exists(test_dest_path):
shutil.rmtree(test_dest_path)
# Creating train, validation and test directories
os.makedirs(train_dest_path)
os.makedirs(validation_dest_path)
os.makedirs(test_dest_path)
def crop_image(img, tol=7):
if img.ndim ==2:
mask = img>tol
return img[np.ix_(mask.any(1),mask.any(0))]
elif img.ndim==3:
gray_img = cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
mask = gray_img>tol
check_shape = img[:,:,0][np.ix_(mask.any(1),mask.any(0))].shape[0]
if (check_shape == 0): # image is too dark so that we crop out everything,
return img # return original image
else:
img1=img[:,:,0][np.ix_(mask.any(1),mask.any(0))]
img2=img[:,:,1][np.ix_(mask.any(1),mask.any(0))]
img3=img[:,:,2][np.ix_(mask.any(1),mask.any(0))]
img = np.stack([img1,img2,img3],axis=-1)
return img
def circle_crop(img):
img = crop_image(img)
height, width, depth = img.shape
largest_side = np.max((height, width))
img = cv2.resize(img, (largest_side, largest_side))
height, width, depth = img.shape
x = width//2
y = height//2
r = np.amin((x, y))
circle_img = np.zeros((height, width), np.uint8)
cv2.circle(circle_img, (x, y), int(r), 1, thickness=-1)
img = cv2.bitwise_and(img, img, mask=circle_img)
img = crop_image(img)
return img
def preprocess_image(base_path, save_path, image_id, HEIGHT, WIDTH, sigmaX=10):
image = cv2.imread(base_path + image_id)
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
image = circle_crop(image)
image = cv2.resize(image, (HEIGHT, WIDTH))
image = cv2.addWeighted(image, 4, cv2.GaussianBlur(image, (0,0), sigmaX), -4 , 128)
cv2.imwrite(save_path + image_id, image)
# Pre-procecss train set
for i, image_id in enumerate(X_train['id_code']):
preprocess_image(train_base_path, train_dest_path, image_id, HEIGHT, WIDTH)
# Pre-procecss validation set
for i, image_id in enumerate(X_val['id_code']):
preprocess_image(train_base_path, validation_dest_path, image_id, HEIGHT, WIDTH)
# Pre-procecss test set
for i, image_id in enumerate(test['id_code']):
preprocess_image(test_base_path, test_dest_path, image_id, HEIGHT, WIDTH)
```
# Data generator
```
datagen=ImageDataGenerator(rescale=1./255,
rotation_range=360,
horizontal_flip=True,
vertical_flip=True)
train_generator=datagen.flow_from_dataframe(
dataframe=X_train,
directory=train_dest_path,
x_col="id_code",
y_col="diagnosis",
class_mode="raw",
batch_size=BATCH_SIZE,
target_size=(HEIGHT, WIDTH),
seed=seed)
valid_generator=datagen.flow_from_dataframe(
dataframe=X_val,
directory=validation_dest_path,
x_col="id_code",
y_col="diagnosis",
class_mode="raw",
batch_size=BATCH_SIZE,
target_size=(HEIGHT, WIDTH),
seed=seed)
test_generator=datagen.flow_from_dataframe(
dataframe=test,
directory=test_dest_path,
x_col="id_code",
batch_size=1,
class_mode=None,
shuffle=False,
target_size=(HEIGHT, WIDTH),
seed=seed)
```
# Model
```
def create_model(input_shape):
input_tensor = Input(shape=input_shape)
base_model = applications.MobileNet(weights=None,
include_top=False,
input_tensor=input_tensor)
base_model.load_weights('../input/keras-notop/mobilenet_1_0_224_tf_no_top.h5')
x = GlobalAveragePooling2D()(base_model.output)
x = Dropout(0.5)(x)
x = Dense(2048, activation='relu')(x)
x = Dropout(0.5)(x)
final_output = Dense(1, activation='linear', name='final_output')(x)
model = Model(input_tensor, final_output)
return model
```
# Train top layers
```
model = create_model(input_shape=(HEIGHT, WIDTH, CHANNELS))
for layer in model.layers:
layer.trainable = False
for i in range(-5, 0):
model.layers[i].trainable = True
metric_list = ["accuracy"]
optimizer = optimizers.Adam(lr=WARMUP_LEARNING_RATE)
model.compile(optimizer=optimizer, loss='mean_squared_error', metrics=metric_list)
model.summary()
STEP_SIZE_TRAIN = train_generator.n//train_generator.batch_size
STEP_SIZE_VALID = valid_generator.n//valid_generator.batch_size
history_warmup = model.fit_generator(generator=train_generator,
steps_per_epoch=STEP_SIZE_TRAIN,
validation_data=valid_generator,
validation_steps=STEP_SIZE_VALID,
epochs=WARMUP_EPOCHS,
verbose=1).history
```
# Fine-tune the complete model (1st step)
```
for layer in model.layers:
layer.trainable = True
es = EarlyStopping(monitor='val_loss', mode='min', patience=ES_PATIENCE, restore_best_weights=True, verbose=1)
rlrop = ReduceLROnPlateau(monitor='val_loss', mode='min', patience=RLROP_PATIENCE, factor=DECAY_DROP, min_lr=1e-6, verbose=1)
callback_list = [es, rlrop]
optimizer = optimizers.Adam(lr=LEARNING_RATE)
model.compile(optimizer=optimizer, loss='mean_squared_error', metrics=metric_list)
model.summary()
history_finetunning = model.fit_generator(generator=train_generator,
steps_per_epoch=STEP_SIZE_TRAIN,
validation_data=valid_generator,
validation_steps=STEP_SIZE_VALID,
epochs=int(EPOCHS*0.8),
callbacks=callback_list,
verbose=1).history
```
# Fine-tune the complete model (2nd step)
```
optimizer = optimizers.SGD(lr=LEARNING_RATE, momentum=0.9, nesterov=True)
model.compile(optimizer=optimizer, loss='mean_squared_error', metrics=metric_list)
history_finetunning_2 = model.fit_generator(generator=train_generator,
steps_per_epoch=STEP_SIZE_TRAIN,
validation_data=valid_generator,
validation_steps=STEP_SIZE_VALID,
epochs=int(EPOCHS*0.2),
callbacks=callback_list,
verbose=1).history
```
# Model loss graph
```
history = {'loss': history_finetunning['loss'] + history_finetunning_2['loss'],
'val_loss': history_finetunning['val_loss'] + history_finetunning_2['val_loss'],
'acc': history_finetunning['acc'] + history_finetunning_2['acc'],
'val_acc': history_finetunning['val_acc'] + history_finetunning_2['val_acc']}
sns.set_style("whitegrid")
fig, (ax1, ax2) = plt.subplots(2, 1, sharex='col', figsize=(20, 14))
ax1.plot(history['loss'], label='Train loss')
ax1.plot(history['val_loss'], label='Validation loss')
ax1.legend(loc='best')
ax1.set_title('Loss')
ax2.plot(history['acc'], label='Train accuracy')
ax2.plot(history['val_acc'], label='Validation accuracy')
ax2.legend(loc='best')
ax2.set_title('Accuracy')
plt.xlabel('Epochs')
sns.despine()
plt.show()
# Create empty arays to keep the predictions and labels
df_preds = pd.DataFrame(columns=['label', 'pred', 'set'])
train_generator.reset()
valid_generator.reset()
# Add train predictions and labels
for i in range(STEP_SIZE_TRAIN + 1):
im, lbl = next(train_generator)
preds = model.predict(im, batch_size=train_generator.batch_size)
for index in range(len(preds)):
df_preds.loc[len(df_preds)] = [lbl[index], preds[index][0], 'train']
# Add validation predictions and labels
for i in range(STEP_SIZE_VALID + 1):
im, lbl = next(valid_generator)
preds = model.predict(im, batch_size=valid_generator.batch_size)
for index in range(len(preds)):
df_preds.loc[len(df_preds)] = [lbl[index], preds[index][0], 'validation']
df_preds['label'] = df_preds['label'].astype('int')
```
# Threshold optimization
```
def classify(x):
if x < 0.5:
return 0
elif x < 1.5:
return 1
elif x < 2.5:
return 2
elif x < 3.5:
return 3
return 4
def classify_opt(x):
if x <= (0 + best_thr_0):
return 0
elif x <= (1 + best_thr_1):
return 1
elif x <= (2 + best_thr_2):
return 2
elif x <= (3 + best_thr_3):
return 3
return 4
def find_best_threshold(df, label, label_col='label', pred_col='pred', do_plot=True):
score = []
thrs = np.arange(0, 1, 0.01)
for thr in thrs:
preds_thr = [label if ((pred >= label and pred < label+1) and (pred < (label+thr))) else classify(pred) for pred in df[pred_col]]
score.append(cohen_kappa_score(df[label_col].astype('int'), preds_thr))
score = np.array(score)
pm = score.argmax()
best_thr, best_score = thrs[pm], score[pm].item()
print('Label %s: thr=%.2f, Kappa=%.3f' % (label, best_thr, best_score))
plt.rcParams["figure.figsize"] = (20, 5)
plt.plot(thrs, score)
plt.vlines(x=best_thr, ymin=score.min(), ymax=score.max())
plt.show()
return best_thr
# Best threshold for label 3
best_thr_3 = find_best_threshold(df_preds, 3)
# Best threshold for label 2
best_thr_2 = find_best_threshold(df_preds, 2)
# Best threshold for label 1
best_thr_1 = find_best_threshold(df_preds, 1)
# Best threshold for label 0
best_thr_0 = find_best_threshold(df_preds, 0)
# Classify predictions
df_preds['predictions'] = df_preds['pred'].apply(lambda x: classify(x))
# Apply optimized thresholds to the predictions
df_preds['predictions_opt'] = df_preds['pred'].apply(lambda x: classify_opt(x))
train_preds = df_preds[df_preds['set'] == 'train']
validation_preds = df_preds[df_preds['set'] == 'validation']
```
# Model Evaluation
## Confusion Matrix
### Original thresholds
```
labels = ['0 - No DR', '1 - Mild', '2 - Moderate', '3 - Severe', '4 - Proliferative DR']
def plot_confusion_matrix(train, validation, labels=labels):
train_labels, train_preds = train
validation_labels, validation_preds = validation
fig, (ax1, ax2) = plt.subplots(1, 2, sharex='col', figsize=(24, 7))
train_cnf_matrix = confusion_matrix(train_labels, train_preds)
validation_cnf_matrix = confusion_matrix(validation_labels, validation_preds)
train_cnf_matrix_norm = train_cnf_matrix.astype('float') / train_cnf_matrix.sum(axis=1)[:, np.newaxis]
validation_cnf_matrix_norm = validation_cnf_matrix.astype('float') / validation_cnf_matrix.sum(axis=1)[:, np.newaxis]
train_df_cm = pd.DataFrame(train_cnf_matrix_norm, index=labels, columns=labels)
validation_df_cm = pd.DataFrame(validation_cnf_matrix_norm, index=labels, columns=labels)
sns.heatmap(train_df_cm, annot=True, fmt='.2f', cmap="Blues",ax=ax1).set_title('Train')
sns.heatmap(validation_df_cm, annot=True, fmt='.2f', cmap=sns.cubehelix_palette(8),ax=ax2).set_title('Validation')
plt.show()
plot_confusion_matrix((train_preds['label'], train_preds['predictions']), (validation_preds['label'], validation_preds['predictions']))
```
### Optimized thresholds
```
plot_confusion_matrix((train_preds['label'], train_preds['predictions_opt']), (validation_preds['label'], validation_preds['predictions_opt']))
```
## Quadratic Weighted Kappa
```
def evaluate_model(train, validation):
train_labels, train_preds = train
validation_labels, validation_preds = validation
print("Train Cohen Kappa score: %.3f" % cohen_kappa_score(train_preds, train_labels, weights='quadratic'))
print("Validation Cohen Kappa score: %.3f" % cohen_kappa_score(validation_preds, validation_labels, weights='quadratic'))
print("Complete set Cohen Kappa score: %.3f" % cohen_kappa_score(np.append(train_preds, validation_preds), np.append(train_labels, validation_labels), weights='quadratic'))
print(" Original thresholds")
evaluate_model((train_preds['label'], train_preds['predictions']), (validation_preds['label'], validation_preds['predictions']))
print(" Optimized thresholds")
evaluate_model((train_preds['label'], train_preds['predictions_opt']), (validation_preds['label'], validation_preds['predictions_opt']))
```
## Apply model to test set and output predictions
```
def apply_tta(model, generator, steps=5):
step_size = generator.n//generator.batch_size
preds_tta = []
for i in range(steps):
generator.reset()
preds = model.predict_generator(generator, steps=step_size)
preds_tta.append(preds)
return np.mean(preds_tta, axis=0)
preds = apply_tta(model, test_generator)
predictions = [classify(x) for x in preds]
predictions_opt = [classify_opt(x) for x in preds]
results = pd.DataFrame({'id_code':test['id_code'], 'diagnosis':predictions})
results['id_code'] = results['id_code'].map(lambda x: str(x)[:-4])
results_opt = pd.DataFrame({'id_code':test['id_code'], 'diagnosis':predictions_opt})
results_opt['id_code'] = results_opt['id_code'].map(lambda x: str(x)[:-4])
# Cleaning created directories
if os.path.exists(train_dest_path):
shutil.rmtree(train_dest_path)
if os.path.exists(validation_dest_path):
shutil.rmtree(validation_dest_path)
if os.path.exists(test_dest_path):
shutil.rmtree(test_dest_path)
```
# Predictions class distribution
```
fig, (ax1, ax2) = plt.subplots(1, 2, sharex='col', figsize=(24, 8.7))
sns.countplot(x="diagnosis", data=results, palette="GnBu_d", ax=ax1).set_title('Test')
sns.countplot(x="diagnosis", data=results_opt, palette="GnBu_d", ax=ax2).set_title('Test optimized')
sns.despine()
plt.show()
val_kappa = cohen_kappa_score(validation_preds['label'], validation_preds['predictions'], weights='quadratic')
val_opt_kappa = cohen_kappa_score(validation_preds['label'], validation_preds['predictions_opt'], weights='quadratic')
results_name = 'submission.csv'
results_opt_name = 'submission_opt.csv'
# if val_kappa > val_opt_kappa:
# results_name = 'submission.csv'
# results_opt_name = 'submission_opt.csv'
# else:
# results_name = 'submission_norm.csv'
# results_opt_name = 'submission.csv'
results.to_csv(results_name, index=False)
display(results.head())
results_opt.to_csv(results_opt_name, index=False)
display(results_opt.head())
```
| github_jupyter |
```
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib
import matplotlib_venn
import tikzplotlib
plt.style.use("ggplot")
df1 = pd.read_csv('User_ProgEx_Loc.csv')
x = df1.loc[:, "Where The Users Gained Programming Experience"]
print(x)
y = df1.loc[:, "Number of Users"]
label = ['school', 'none', 'both', 'outside school']
fig = plt.figure()
ax1 = fig.add_subplot(111)
ax1.bar(x, y)
ax1.set_xticklabels(label)
ax1.set_xlabel("Where The Users Gained Programming Experience")
ax1.set_ylabel("Number of Users")
tikzplotlib.save("fig1.tex")
df = pd.read_csv('User_ProgExp.csv')
x = df.loc[:, "Languages The Users Had Experience With"]
y = df.loc[:, "Number of Users"]
label = ['Scratch', 'Python', 'Java', 'C++', 'Others']
fig = plt.figure()
ax = fig.add_subplot(111)
ax.bar(x, y, color=(0.1, 0.7, 0.3))
ax.set_xticklabels(label)
ax.set_xlabel("Languages The Users Had Experience With")
ax.set_ylabel("Number of Users")
ax.set_facecolor('xkcd:white')
plt.grid(color='grey', linestyle='-.', linewidth=0.7)
tikzplotlib.save("fig2.tex")
from matplotlib_venn import venn3, venn3_circles
fig = plt.figure()
ax = fig.add_subplot(111)
# depict venn diagram
v = venn3(subsets=(17, 1, 5, 10, 0, 0, 0), set_colors=((0.1, 0.7, 0.1), (0.1, 0.1, 0.7), 'r'),
set_labels=('In School', 'Outside School', 'None'))
# add outline
venn3_circles(subsets=(17, 1, 5, 10, 0, 0, 0),
linewidth=1.5)
plt.show()
tikzplotlib.save("fig3.tex")
df = pd.read_csv('Users_Grade.csv')
x = df.loc[:, "Grade"]
y = df.loc[:, "Number of Users"]
label = ['6', '7', '8', 'Unknown']
fig = plt.figure()
ax = fig.add_subplot(111)
ax.barh(x, y, color=(0.1, 0.7, 0.3))
ax.set_yticklabels(label)
ax.set_ylabel("Grade")
ax.set_xlabel("Number of Users")
ax.set_facecolor('xkcd:white')
plt.grid(color='grey', linestyle='-.', linewidth=0.7)
tikzplotlib.save("fig5.tex")
df = pd.read_csv('Prog_Level.csv')
x = df.loc[:, "Number of Commands"]
y1 = df.loc[:, "Level"]
y2 = df.loc[:, "Legend"]
label = ['0.0', '0.1', '0.2', '0.3', '1.1', '1.2', '1.3', '2.1', '2.2', '2.3', '3.1', '3.2', '3.3']
fig = plt.figure()
ax = fig.add_subplot(111)
# x = ['A', 'B', 'C', 'D']
# y1 = [10, 20, 10, 30]
# y2 = [20, 25, 15, 25]
# # plot bars in stack manner
# plt.bar(x, y1, color='r')
# plt.bar(x, y2, bottom=y1, color='b')
ax.barh(y1, x, color=(0.1, 0.7, 0.3))
# ax.bar(y2, x, bottom=y1, color='b')
ax.set_yticklabels(label)
ax.set_ylabel("Grade")
ax.set_xlabel("Number of Users")
ax.set_facecolor('xkcd:white')
plt.grid(color='grey', linestyle='-.', linewidth=0.7)
plt.show()
tikzplotlib.save("fig6.tex")
df = pd.read_csv('Users_Gender.csv')
x = df.loc[:, "Gender"]
y = df.loc[:, "Number of Users"]
label = ['Male', 'Female']
fig = plt.figure()
ax = fig.add_subplot(111)
ax.bar(x, y, width=0.3, color=(0.1, 0.7, 0.3))
ax.set_xticklabels(label)
ax.set_ylabel("Gender")
ax.set_xlabel("Number of Users")
ax.set_facecolor('xkcd:white')
plt.grid(color='grey', linestyle='-.', linewidth=0.7)
tikzplotlib.save("fig4.tex")
df = pd.read_csv('User_ProgExp.csv')
x = df.loc[:, "Languages The Users Had Experience With"]
y = df.loc[:, "Number of Users"]
label = ['Scratch', 'Python', 'Others', 'C++', 'Java']
fig = plt.figure()
ax = fig.add_subplot(111)
ax.pie(y, labels = label, colors=[(0.1, 0.7, 0.3), (0.1, 0.3, 0.7), 'yellow', (0.7, 0.1, 0.2), (0.1, 0.7, 0.8)])
plt.show()
tikzplotlib.save("fig2.tex")
```
| github_jupyter |
# How to Efficiently Read BigQuery Data from TensorFlow 2.3
## Learning Objectives
* Build a benchmark model.
* Find the breakoff point for Keras.
* Training a TensorFlow/Keras model that reads from BigQuery.
* Load TensorFlow model into BigQuery.
## Introduction
In this notebook, you learn
["How to efficiently read BigQuery data from TensorFlow 2.x"](https://medium.com/@lakshmanok/how-to-read-bigquery-data-from-tensorflow-2-0-efficiently-9234b69165c8)
The example problem is to find credit card fraud from the dataset published in:
<i>
Andrea Dal Pozzolo, Olivier Caelen, Reid A. Johnson and Gianluca Bontempi. Calibrating Probability with Undersampling for Unbalanced Classification. In Symposium on Computational Intelligence and Data Mining (CIDM), IEEE, 2015
</i>
and available in BigQuery at <pre>bigquery-public-data.ml_datasets.ulb_fraud_detection</pre>
Each learning objective will correspond to a __#TODO__ in this student lab notebook -- try to complete this notebook first and then review the [solution notebook](../solutions/bigquery_tensorflow.ipynb).
## Build a benchmark model
In order to compare things, we will do a simple logistic regression in BigQuery ML.
Note that we are using all the columns in the dataset as predictors (except for the Time and Class columns).
The Time column is used to split the dataset 80:20 with the first 80% used for training and the last 20% used for evaluation.
We will also have BigQuery ML automatically balance the weights.
Because the Amount column has a huge range, we take the log of it in preprocessing.
```
%%bash
# create output dataset
bq mk advdata
%%bigquery
CREATE OR REPLACE MODEL advdata.ulb_fraud_detection
TRANSFORM(
* EXCEPT(Amount),
SAFE.LOG(Amount) AS log_amount
)
OPTIONS(
INPUT_LABEL_COLS=['class'],
AUTO_CLASS_WEIGHTS = TRUE,
DATA_SPLIT_METHOD='seq',
DATA_SPLIT_COL='Time',
MODEL_TYPE='logistic_reg'
) AS
SELECT
*
FROM `bigquery-public-data.ml_datasets.ulb_fraud_detection`
%%bigquery
# Use the ML.EVALUATE function to evaluate model metrics
SELECT * FROM TODO: ___________(MODEL advdata.ulb_fraud_detection)
%%bigquery
SELECT predicted_class_probs, Class
# The ML.PREDICT function is used to predict outcomes using the model
FROM TODO: ___________( MODEL advdata.ulb_fraud_detection,
(SELECT * FROM `bigquery-public-data.ml_datasets.ulb_fraud_detection` WHERE Time = 85285.0)
)
```
## Find the breakoff point for Keras
When we do the training in Keras & TensorFlow, we need to find the place to split the dataset and how to weight the imbalanced data.
(BigQuery ML did that for us because we specified 'seq' as the split method and auto_class_weights to be True).
```
%%bigquery
WITH counts AS (
SELECT
APPROX_QUANTILES(Time, 5)[OFFSET(4)] AS train_cutoff
, COUNTIF(CLASS > 0) AS pos
, COUNTIF(CLASS = 0) AS neg
FROM `bigquery-public-data`.ml_datasets.ulb_fraud_detection
)
SELECT
train_cutoff
, SAFE.LOG(SAFE_DIVIDE(pos,neg)) AS output_bias
, 0.5*SAFE_DIVIDE(pos + neg, pos) AS weight_pos
, 0.5*SAFE_DIVIDE(pos + neg, neg) AS weight_neg
FROM TODO: ___________ # Table Name
```
The time cutoff is 144803 and the Keras model's output bias needs to be set at -6.36
The class weights need to be 289.4 and 0.5
## Training a TensorFlow/Keras model that reads from BigQuery
Create the dataset from BigQuery
```
# import necessary libraries
import tensorflow as tf
from tensorflow.python.framework import dtypes
from tensorflow_io.bigquery import BigQueryClient
from tensorflow_io.bigquery import BigQueryReadSession
def features_and_labels(features):
label = features.pop('Class') # this is what we will train for
return features, label
def read_dataset(client, row_restriction, batch_size=2048):
GCP_PROJECT_ID='qwiklabs-gcp-03-5b2f0816822f' # CHANGE
COL_NAMES = ['Time', 'Amount', 'Class'] + ['V{}'.format(i) for i in range(1,29)]
COL_TYPES = [dtypes.float64, dtypes.float64, dtypes.int64] + [dtypes.float64 for i in range(1,29)]
DATASET_GCP_PROJECT_ID, DATASET_ID, TABLE_ID, = 'bigquery-public-data.ml_datasets.ulb_fraud_detection'.split('.')
bqsession = client.read_session(
"projects/" + GCP_PROJECT_ID,
DATASET_GCP_PROJECT_ID, TABLE_ID, DATASET_ID,
COL_NAMES, COL_TYPES,
requested_streams=2,
row_restriction=row_restriction)
dataset = bqsession.parallel_read_rows()
return dataset.prefetch(1).map(features_and_labels).shuffle(batch_size*10).batch(batch_size)
client = BigQueryClient()
temp_df = TODO: ___________(client, 'Time <= 144803', 2) # Function Name
for row in temp_df:
print(row)
break
train_df = read_dataset(client, 'Time <= 144803', 2048)
eval_df = read_dataset(client, 'Time > 144803', 2048)
```
Create Keras model
```
metrics = [
tf.keras.metrics.BinaryAccuracy(name='accuracy'),
tf.keras.metrics.Precision(name='precision'),
tf.keras.metrics.Recall(name='recall'),
tf.keras.metrics.AUC(name='roc_auc'),
]
# create inputs, and pass them into appropriate types of feature columns (here, everything is numeric)
inputs = {
'V{}'.format(i) : tf.keras.layers.Input(name='V{}'.format(i), shape=(), dtype='float64') for i in range(1, 29)
}
inputs['Amount'] = tf.keras.layers.Input(name='Amount', shape=(), dtype='float64')
input_fc = [tf.feature_column.numeric_column(colname) for colname in inputs.keys()]
# transformations. only the Amount is transformed
transformed = inputs.copy()
transformed['Amount'] = tf.keras.layers.Lambda(
lambda x: tf.math.log(tf.math.maximum(x, 0.01)), name='log_amount')(inputs['Amount'])
input_layer = tf.keras.layers.DenseFeatures(input_fc, name='inputs')(transformed)
# Deep learning model
d1 = tf.keras.layers.Dense(16, activation='relu', name='d1')(input_layer)
d2 = tf.keras.layers.Dropout(0.25, name='d2')(d1)
d3 = tf.keras.layers.Dense(16, activation='relu', name='d3')(d2)
output = tf.keras.layers.Dense(1, activation='sigmoid', name='d4', bias_initializer=tf.keras.initializers.Constant())(d3)
model = tf.keras.Model(inputs, output)
model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=metrics)
tf.keras.utils.plot_model(model, rankdir='LR')
class_weight = {0: 0.5, 1: 289.4}
# Trains the model for a fixed number of epochs
history = TODO: _________(train_df, validation_data=eval_df, epochs=20, class_weight=class_weight)
import matplotlib.pyplot as plt
plt.plot(history.history['val_roc_auc']);
plt.xlabel('Epoch');
plt.ylabel('AUC');
```
## Load TensorFlow model into BigQuery
Now that we have trained a TensorFlow model off BigQuery data ...
let's load the model into BigQuery and use it for batch prediction!
```
BUCKET='<your-bucket>' # CHANGE TO SOMETHING THAT YOU OWN
model.save('gs://{}/bqexample/export'.format(BUCKET))
%%bigquery
CREATE OR REPLACE MODEL advdata.keras_fraud_detection
OPTIONS(model_type='tensorflow', model_path='gs://qwiklabs-gcp-03-5b2f0816822f/bqexample/export/*')
```
Now predict with this model (the reason it's called 'd4' is because the output node of my Keras model was called 'd4').
To get probabilities, etc. we'd have to add the corresponding outputs to the Keras model.
```
%%bigquery
SELECT d4, Class
FROM ML.PREDICT( MODEL advdata.keras_fraud_detection,
(SELECT * FROM `bigquery-public-data.ml_datasets.ulb_fraud_detection` WHERE Time = 85285.0)
)
```
Copyright 2021 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
| github_jupyter |
## Kats 203 Time Series Features
This tutorial will introduce `TsFeatures` in Kats, which allows you to extract meaningful features from time series. The table of contents for Kats 203 is as follows:
1. Basic Usage with a Single Time Series
2. Applications with Multiple Time Series
2.1 Largest Seasonal Component
2.2 Highest Entropy ("Least Predictable")
2.3 Cluster Similar Time Series
3. Out-in/out features for calculation
3.1 Opting-out features
3.2 Opting-in features
**Note:** We provide two types of tutorial notebooks
- **Kats 101**, basic data structure and functionalities in Kats
- **Kats 20x**, advanced topics, including advanced forecasting techniques, advanced detection algorithms, `TsFeatures`, meta-learning, etc.
```
%%capture
# For Google Colab:
!pip install kats
import sys
sys.path.append("../")
import numpy as np
import pandas as pd
import pprint
import matplotlib.pyplot as plt
from sklearn.decomposition import PCA
from kats.consts import TimeSeriesData
from statsmodels.tsa.seasonal import STL
from kats.utils.simulator import Simulator
from sklearn.preprocessing import StandardScaler
from kats.tsfeatures.tsfeatures import TsFeatures
import warnings
warnings.simplefilter(action='ignore')
```
For the purposes of this tutorial, we are going to use the `Simulator` to generate a list of 30 different `TimeSeriesData` objects called `ts_list`. It contains 10 time series simulated from the ARIMA model, 10 time series with trend shifts, and 10 time series with level shifts.
```
sim = Simulator(n=90, freq="D", start = "2021-01-01") # simulate 90 days of data
random_seed = 100
# generate 10 TimeSeriesData with arima_sim
np.random.seed(random_seed) # setting numpy seed
arima_sim_list = [sim.arima_sim(ar=[0.1, 0.05], ma = [0.04, 0.1], d = 1) for _ in range(10)]
# generate 10 TimeSeriesData with trend shifts
trend_sim_list = [
sim.trend_shift_sim(
cp_arr = [30, 60, 75],
trend_arr=[3, 15, 2, 8],
intercept=30,
noise=50,
seasonal_period=7,
seasonal_magnitude=np.random.uniform(10, 100),
random_seed=random_seed
) for _ in range(10)
]
# generate 10 TimeSeriesData with level shifts
level_shift_list = [
sim.level_shift_sim(
cp_arr = [30, 60, 75],
level_arr=[1.35, 1.05, 1.35, 1.2],
noise=0.05,
seasonal_period=7,
seasonal_magnitude=np.random.uniform(0.1, 1.0),
random_seed=random_seed
) for _ in range(10)
]
ts_list = arima_sim_list + trend_sim_list + level_shift_list
```
# 1. Basic Usage with a Single Time Series
We begin by introducing the basic usage of `TsFeatures`. For the purposes of this example, we will use the simulator to simulate a bunch of different time series from the ARIMA model.
`TsFeatures` currently can only process one time series a time, so let's start by taking a look at the first time series we generated above.
```
ts = ts_list[0]
# plot the time series
ts.plot(cols=['value'])
plt.xticks(rotation = 45)
plt.show()
```
Extracting the basic features from the time series using `TsFeatures` is straightforward. We can do so as follows.
```
# Step 1. initiate TsFeatures
model = TsFeatures()
# Step 2. use .transform() method, and apply on the target time series data
output_features = model.transform(ts)
output_features
```
The dictionary above shows 40 features, which are the features that we calculate by default. There are 28 additional features that we support in `TsFeatures`, and users can select which features they would like to include in their calculations using the `selected_features` argument. We will see an example of this later.
## 2. Applications with Multiple Time Series
Now we will look at each of the time series in `ts_list` and use `TsFeatures` to calculate the features for each of them
```
model = TsFeatures()
output_features = [model.transform(ts) for ts in ts_list] # loop through time series data and perform transformation
```
We can view the results as a `DataFrame` as follows, which has one row for each time series in `ts_list` and each column represents a different feature.
```
df_features = pd.DataFrame(output_features) # converting to dataframe
df_features.head()
```
Now we will take a look at some of the applications of the features we have just calculated for each each of the time series in `TsFeatures`.
### 2.1 Largest Seasonal Component
We will demonstrate leveraging `TsFeatures` for finding the time series data with the highest seasonality strength among the simulated time series data.
```
# finding the index of the time series sample with the highest seasonality strength
index_target_ts = df_features['seasonality_strength'].argmax()
target_ts = ts_list[index_target_ts]
# Plot the time series
target_ts.plot(cols=['value'])
plt.xticks(rotation = 45)
plt.show()
```
Now let's take a look at the STL decomposition of this time series to see its seasonal component. We use the [`STL` module from `statsmodels`](https://www.statsmodels.org/devel/generated/statsmodels.tsa.seasonal.STL.html).
```
stl = STL(target_ts.value.values, period=7)
res = stl.fit()
plt.plot(
pd.to_datetime(target_ts.time.values),
res.seasonal
)
plt.xticks(rotation = 90);
plt.title(f'Seasonal component - variance: {np.round(np.var(res.seasonal), 2)}');
```
Now we repeat the process for the time series in `ts_list` with the smallest seasonal component. We see that the variance of the seasonal components is much smaller in this case.
```
# finding the index of the time series sample with the smallest seasonality strength
index_target_ts = df_features['seasonality_strength'].argmin()
target_ts = ts_list[index_target_ts].to_dataframe()
# Do an STL decomposition and plot the results
stl = STL(target_ts.value.values, period=7)
res = stl.fit()
plt.plot(
pd.to_datetime(target_ts.time.values),
res.seasonal
)
plt.xticks(rotation = 45);
plt.title(f'Seasonal component - variance: {np.round(np.var(res.seasonal), 2)}');
```
### 2.2 Highest Entropy ("Least Predictable")
We can intuitively understand entropy as the measure of the disorder of a time series. In general, a time series with higher entropy will be harder to forecast. Here, we show how to use `TsFeatures` to find the time series in `ts_list` with the highest entropy.
```
# TODO: Re-run these cells once we have fixed the definition of entropy in TsFeatures.
# find the index of the time series sample with the highest entropy
index_target_ts = df_features['entropy'].argmax()
target_ts = ts_list[index_target_ts]
# Plot the time series
target_ts.plot(cols=['value'])
plt.xticks(rotation = 45)
plt.show()
```
Let's compare the above figure with the time series data with the lowest entropy identified by `TsFeatures`.
```
# find the index of the time series sample with the lowest entropy
index_target_ts = df_features['entropy'].argmin()
target_ts = ts_list[index_target_ts]
# Plot the time series
target_ts.plot(cols=['value'])
plt.xticks(rotation = 45)
plt.show()
```
As we can see from the figures above, this second plot shows a time series with a clear change point and two distinct trends, suggesting it is easier to forecast than the first time series.
### 2.3 Cluster Similar Time Series
Here we are going to use the features we get from `TsFeatures` to try to cluster the time series.
Let's perform a dimension reduction on the simulated time series data, and visualize to see if there's clear pattern of clusters. In this example, we pick 5 features to use for each time series, and then we use `PCA` (combined with `StandardScaler`) from `sklearn` to project this representation into two dimensions.
```
# performing dimension reduction on the time series samples
ls_features = ['lumpiness', 'entropy', 'seasonality_strength', 'stability', 'level_shift_size']
df_dataset = df_features[ls_features]
x_2d = PCA(n_components=2).fit_transform(X=StandardScaler().fit_transform(df_dataset[ls_features].values))
df_dataset['pca_component_1'] = x_2d[:,0]
df_dataset['pca_component_2'] = x_2d[:,1]
```
Now we can plot the results below. We have color-coded the different types of simulated time series (ARIMA, trend shift, level shift) and we can see that series of the same type are closer to each other.
```
# Plot the PCA projections of each time series
plt.figure(figsize = (15,8))
# Plot ARIMA time series in red
ax = df_dataset.iloc[0:10].plot(x='pca_component_1', y='pca_component_2', kind='scatter', color='red')
# Plot trend shift time series in green
df_dataset.iloc[10:20].plot(x='pca_component_1', y='pca_component_2', kind='scatter', color='green', ax=ax)
# Plot level shift time series in yellow
df_dataset.iloc[20:].plot(x='pca_component_1', y='pca_component_2', kind='scatter', color='yellow', ax=ax)
plt.title('Visualization of the dimension reduced time series samples')
plt.legend(['ARIMA', 'Trend Shift', 'Level Shift'])
plt.show()
```
## 3. Out-in/out features for calculation
In `TsFeatures`, you can choose which features and feature groups you would like to calculate and which you would like to skip.
### 3.1 Opting-out features
Let's start by returning to our initial example, where we calculated the first time series in `ts_list`. This calculates the 40 features that we calculate by default.
```
ts = ts_list[0]
TsFeatures().transform(ts)
```
We can omit a single feature, like `seasonality_strength`, by passing `seasonality_strength=False` into `TsFeatures` like such.
```
TsFeatures(seasonality_strength=False).transform(ts)
```
The `seasonality_strength` feature belongs to the 'stl_features' feature group. Each of the 68 features that we currently support in `TsFeatures` are arranged into 14 feature groups. Below is the dictionary displaying the mapping feature feature groups and features.
```
feature_group_mapping = TsFeatures().feature_group_mapping
pprint.pprint(feature_group_mapping)
```
We can opt to skip the calculation of an entire feature group like `stl_features` the same way we skipped the calculation of `seasonality_strength` in the example above.
```
TsFeatures(stl_features=False).transform(ts)
```
We can also opt-out combinations of features and feature groups like su
```
tsf = TsFeatures(length = False, mean = False, stl_features=False)
tsf.transform(ts)
```
### 3.2 Opting-in features
We can use the `selected_features` argument in `TsFeatures` to specify which features and feature groups we would like to include in our calculations. When we use this argument, a default feature will not be included unless that feature or its group is explicited included in `selected_features`. Similarly, a feature not included by default can be included by including it or its group in `selected_features`.
Here is an example where we specify a list of features to calculate.
```
TsFeatures(selected_features = [
'mean',
'var',
'entropy',
'lumpiness',
'hurst',
'trend_strength',
'y_acf1',
'firstmin_ac',
'holt_alpha',
'nowcast_roc',
'bocp_num',
'seasonality_mag',
]).transform(ts)
```
Here is an example where we specify a list of features groups to calculate.
```
TsFeatures(selected_features = [
'statistics',
'acfpacf_features',
'nowcasting',
'time',
]).transform(ts)
```
We can specify a combination of features and feature groups to calculate:
```
TsFeatures(selected_features = [
'statistics',
'acfpacf_features',
'nowcasting',
'seasonality_strength',
]).transform(ts)
```
Lastly, we can mix up the opt-in and opt-out to calculate majority of the features among some feature groups, while opting-out some of the features within these feature groups that are opt-in.
We can also opt-out specific features from the feature groups we include in `selected_features`. Here is an example where we calculate all of the features in the `statistics` group except for the `mean`.
```
TsFeatures(selected_features = ['statistics'], mean = False).transform(ts)
```
If we include the name of a specific feature in `selected_features` and then try to opt-out that same feature or its feature group, we will get an error. For example, the inverse of our previous example doesn't work.
```
# THIS DOES NOT WORK
# TsFeatures(selected_features = ['mean'], statitics = False).transform(ts)
# THIS ALSO DOES NOT WORK
# TsFeatures(selected_features = ['mean'], mean = False).transform(ts)
```
## Conclusion
In this tutorial, we've demonstrated basic functions of `TsFeatures`, along with the demonstration of some of the interesting applications. We hope you've enjoyed the tutorial and look forward to using `TsFeatures` in the future!
| github_jupyter |
# Script based external simulators
In this notebook, we demonstrate the usage of the script based external simulators, summary statistics, and distance functions features.
These allow to use near-arbitrary programing languages and output for pyABC. The main concept is that all communication happens via the file system. This comes at the cost of a considerable overhead, making this feature not optimal for models with a low simulation time. For more expensive models, the overhead should be negligible.
This notebook is similar to the using_R notebook, but circumvents usage of the rpy2 package.
```
# install if not done yet
!pip install pyabc --quiet
import pyabc
import pyabc.external
```
Here, we define model, summary statistics and distance. Note that if possible, alternatively FileIdSumStat can be used to read in the summary statistics directly to python and then use a python based distance function.
```
model = pyabc.external.ExternalModel(executable="Rscript", file="rmodel/model.r")
sumstat = pyabc.external.ExternalSumStat(executable="Rscript", file="rmodel/sumstat.r")
distance = pyabc.external.ExternalDistance(executable="Rscript", file="rmodel/distance.r")
dummy_sumstat = pyabc.external.create_sum_stat() # can also use a real file here
```
```
pars = {'meanX': 3, 'meanY': 4}
mf = model(pars)
print(mf)
sf = sumstat(mf)
print(sf)
distance(sf, dummy_sumstat)
from pyabc import Distribution, RV, ABCSMC
prior = Distribution(meanX=RV("uniform", 0, 10),
meanY=RV("uniform", 0, 10))
abc = ABCSMC(model, prior, distance,
summary_statistics=sumstat,
population_size=20)
import os
from tempfile import gettempdir
db = "sqlite:///" + os.path.join(gettempdir(), "test.db")
abc.new(db, dummy_sumstat)
history = abc.run(minimum_epsilon=0.9, max_nr_populations=4)
```
Note the significantly longer runtimes compared to using rpy2. This is because the simulation time of this model is very short, such that repeatedly accessing the file system creates a notable overhead. For more expensive models, this overhead however becomes less notable. Still, if applicable, more efficient ways of communication between model and simulator are preferable.
```
from pyabc.visualization import plot_kde_2d
for t in range(history.n_populations):
df, w = abc.history.get_distribution(0, t)
ax = plot_kde_2d(df, w, "meanX", "meanY",
xmin=0, xmax=10,
ymin=0, ymax=10,
numx=100, numy=100)
ax.scatter([4], [8],
edgecolor="black",
facecolor="white",
label="Observation");
ax.legend();
ax.set_title("PDF t={}".format(t))
```
| github_jupyter |
This demo shows how to use integrated gradients in graph convolutional networks to obtain accurate importance estimations for both the nodes and edges. The notebook consists of three parts:
- setting up the node classification problem for Cora citation network
- training and evaluating a GCN model for node classification
- calculating node and edge importances for model's predictions of query ("target") nodes
<a name="refs"></a>
**References**
[1] Axiomatic Attribution for Deep Networks. M. Sundararajan, A. Taly, and Q. Yan.
Proceedings of the 34th International Conference on Machine Learning, Sydney, Australia, PMLR 70, 2017
([link](https://arxiv.org/pdf/1703.01365.pdf)).
[2] Adversarial Examples on Graph Data: Deep Insights into Attack and Defense. H. Wu, C. Wang, Y. Tyshetskiy, A. Docherty, K. Lu, and L. Zhu. arXiv: 1903.01610 ([link](https://arxiv.org/abs/1903.01610)).
```
import networkx as nx
import pandas as pd
import numpy as np
from scipy import stats
import os
import time
import stellargraph as sg
from stellargraph.mapper import FullBatchNodeGenerator
from stellargraph.layer import GCN
import keras
from keras import layers, optimizers, losses, metrics, Model, regularizers
from sklearn import preprocessing, feature_extraction, model_selection
from copy import deepcopy
import matplotlib.pyplot as plt
%matplotlib inline
```
### Loading the CORA network
**Downloading the CORA dataset:**
The dataset used in this demo can be downloaded from https://linqs-data.soe.ucsc.edu/public/lbc/cora.tgz
The following is the description of the dataset:
> The Cora dataset consists of 2708 scientific publications classified into one of seven classes.
> The citation network consists of 5429 links. Each publication in the dataset is described by a
> 0/1-valued word vector indicating the absence/presence of the corresponding word from the dictionary.
> The dictionary consists of 1433 unique words. The README file in the dataset provides more details.
Download and unzip the cora.tgz file to a location on your computer and set the `data_dir` variable to
point to the location of the dataset (the directory containing "cora.cites" and "cora.content").
```
data_dir = os.path.expanduser("~/data/cora")
```
Load the graph from edgelist (in `cited-paper` <- `citing-paper` order)
```
edgelist = pd.read_csv(os.path.join(data_dir, "cora.cites"), sep='\t', header=None, names=["target", "source"])
edgelist["label"] = "cites"
Gnx = nx.from_pandas_edgelist(edgelist, edge_attr="label")
nx.set_node_attributes(Gnx, "paper", "label")
feature_names = ["w_{}".format(ii) for ii in range(1433)]
column_names = feature_names + ["subject"]
node_data = pd.read_csv(os.path.join(data_dir, "cora.content"), sep='\t', header=None, names=column_names)
```
The adjacency matrix is ordered by the graph nodes from G.nodes(). To make computations easier, let's reindex the node data to maintain the same ordering.
```
graph_nodes = list(Gnx.nodes())
node_data = node_data.loc[graph_nodes]
```
### Splitting the data
For machine learning we want to take a subset of the nodes for training, and use the rest for validation and testing. We'll use scikit-learn again to do this.
Here we're taking 140 node labels for training, 500 for validation, and the rest for testing.
```
train_data, test_data = model_selection.train_test_split(node_data,
train_size=140,
test_size=None,
stratify=node_data['subject'])
val_data, test_data = model_selection.train_test_split(test_data,
train_size=500,
test_size=None,
stratify=test_data['subject'])
```
### Converting to numeric arrays
For our categorical target, we will use one-hot vectors that will be fed into a soft-max Keras layer during training. To do this conversion ...
We now do the same for the node attributes we want to use to predict the subject. These are the feature vectors that the Keras model will use as input. The CORA dataset contains attributes 'w_x' that correspond to words found in that publication. If a word occurs more than once in a publication the relevant attribute will be set to one, otherwise it will be zero.
```
target_encoding = feature_extraction.DictVectorizer(sparse=False)
feature_names = ["w_{}".format(ii) for ii in range(1433)]
column_names = feature_names + ["subject"]
train_targets = target_encoding.fit_transform(train_data[["subject"]].to_dict('records'))
val_targets = target_encoding.transform(val_data[["subject"]].to_dict('records'))
test_targets = target_encoding.transform(test_data[["subject"]].to_dict('records'))
node_ids = node_data.index
all_targets = target_encoding.transform(
node_data[["subject"]].to_dict("records")
)
node_features = node_data[feature_names]
```
### Creating the GCN model in Keras
Now create a StellarGraph object from the NetworkX graph and the node features and targets. To feed data from the graph to the Keras model we need a generator. Since GCN is a full-batch model, we use the `FullBatchNodeGenerator` class.
```
G = sg.StellarGraph(Gnx, node_features=node_features)
generator = FullBatchNodeGenerator(G, sparse=True)
```
For training we map only the training nodes returned from our splitter and the target values.
```
train_gen = generator.flow(train_data.index, train_targets)
```
Now we can specify our machine learning model: tn this example we use two GCN layers with 16-dimensional hidden node features at each layer with ELU activation functions.
```
layer_sizes = [16, 16]
gcn = GCN(layer_sizes=layer_sizes,
activations=["elu", "elu"],
generator=generator,
dropout=0.3,
kernel_regularizer=regularizers.l2(5e-4))
# Expose the input and output tensors of the GCN model for node prediction, via GCN.node_model() method:
x_inp, x_out = gcn.node_model()
# Snap the final estimator layer to x_out
x_out = layers.Dense(units=train_targets.shape[1], activation="softmax")(x_out)
```
### Training the model
Now let's create the actual Keras model with the input tensors `x_inp` and output tensors being the predictions `x_out` from the final dense layer
```
model = keras.Model(inputs=x_inp, outputs=x_out)
model.compile(
optimizer=optimizers.Adam(lr=0.01),# decay=0.001),
loss=losses.categorical_crossentropy,
metrics=[metrics.categorical_accuracy],
)
```
Train the model, keeping track of its loss and accuracy on the training set, and its generalisation performance on the validation set (we need to create another generator over the validation data for this)
```
val_gen = generator.flow(val_data.index, val_targets)
```
Train the model
```
history = model.fit_generator(train_gen,
shuffle=False,
epochs=20,
verbose=2,
validation_data=val_gen)
import matplotlib.pyplot as plt
%matplotlib inline
def remove_prefix(text, prefix):
return text[text.startswith(prefix) and len(prefix):]
def plot_history(history):
metrics = sorted(set([remove_prefix(m, "val_") for m in list(history.history.keys())]))
for m in metrics:
# summarize history for metric m
plt.plot(history.history[m])
plt.plot(history.history['val_' + m])
plt.title(m)
plt.ylabel(m)
plt.xlabel('epoch')
plt.legend(['train', 'validation'], loc='best')
plt.show()
plot_history(history)
```
Evaluate the trained model on the test set
```
test_gen = generator.flow(test_data.index, test_targets)
test_metrics = model.evaluate_generator(test_gen)
print("\nTest Set Metrics:")
for name, val in zip(model.metrics_names, test_metrics):
print("\t{}: {:0.4f}".format(name, val))
```
# Node and link importance via saliency maps
In order to understand why a selected node is predicted as a certain class we want to find the node feature importance, total node importance, and link importance for nodes and edges in the selected node's neighbourhood (ego-net). These importances give information about the effect of changes in the node's features and its neighbourhood on the prediction of the node, specifically:
#### Node feature importance:
Given the selected node $t$ and the model's prediction $s(c)$ for class $c$. The feature importance can be calculated for each node $v$ in the selcted node's ego-net where the importance of feature $f$ for node $v$ is the change predicted score $s(c)$ for the selected node when the feature $f$ of node $v$ is perturbed.
#### Total node importance:
This is defined as the sum of the feature importances for node $v$ for all features. Nodes with high importance (positive or negative) affect the prediction for the selected node more than links with low importance.
#### Link importance:
This is defined as the change in the selected node's predicted score $s(c)$ if the link $e=(u, v)$ is removed from the graph. Links with high importance (positive or negative) affect the prediction for the selected node more than links with low importance.
Node and link importances can be used to assess the role of nodes and links in model's predictions for the node(s) of interest (the selected node). For datasets like CORA-ML, the features and edges are binary, vanilla gradients may not perform well so we use integrated gradients [[1]](#refs) to compute them.
Another interesting application of node and link importances is to identify model vulnerabilities to attacks via perturbing node features and graph structure (see [[2]](#refs)).
To investigate these importances we use the StellarGraph `saliency_maps` routines:
```
from stellargraph.utils.saliency_maps import IntegratedGradients, GradientSaliency
```
Select the target node whose prediction is to be interpreted
```
target_idx = 7 # index of the selected node in G.nodes()
target_nid = graph_nodes[target_idx] # Node ID of the selected node
y_true = all_targets[target_idx] # true class of the target node
all_gen = generator.flow(graph_nodes)
y_pred = model.predict_generator(all_gen)[0, target_idx]
class_of_interest = np.argmax(y_pred)
print("Selected node id: {}, \nTrue label: {}, \nPredicted scores: {}".format(target_nid, y_true, y_pred.round(2)))
```
Get the node feature importance by using integrated gradients
```
int_grad_saliency = IntegratedGradients(model, train_gen)
```
For the parameters of `get_node_importance` method, `X` and `A` are the feature and adjacency matrices, respectively. If `sparse` option is enabled, `A` will be the non-zero values of the adjacency matrix with `A_index` being the indices. `target_idx` is the node of interest, and `class_of_interest` is set as the predicted label of the node. `steps` indicates the number of steps used to approximate the integration in integrated gradients calculation. A larger value of `steps` gives better approximation, at the cost of higher computational overhead.
```
integrated_node_importance = int_grad_saliency.get_node_importance(
target_idx, class_of_interest, steps=50
)
integrated_node_importance.shape
print('\nintegrated_node_importance', integrated_node_importance.round(2))
print('integrate_node_importance.shape = {}'.format(integrated_node_importance.shape))
print('integrated self-importance of target node {}: {}'.format(target_nid, integrated_node_importance[target_idx].round(2)))
```
Check that number of non-zero node importance values is less or equal the number of nodes in target node's K-hop ego net (where K is the number of GCN layers in the model)
```
G_ego = nx.ego_graph(G, target_nid, radius=len(gcn.activations))
print('Number of nodes in the ego graph: {}'.format(len(G_ego.nodes())))
print("Number of non-zero elements in integrated_node_importance: {}".format(np.count_nonzero(integrated_node_importance)))
```
We now compute the link importance using integrated gradients [[1]](#refs). Integrated gradients are obtained by cumulating the gradients along the path between the baseline (all-zero graph) and the state of the graph. They provide better sensitivity for the graphs with binary features and edges compared with the vanilla gradients.
```
integrate_link_importance = int_grad_saliency.get_integrated_link_masks(target_idx, class_of_interest, steps=50)
integrate_link_importance_dense = np.array(integrate_link_importance.todense())
print('integrate_link_importance.shape = {}'.format(integrate_link_importance.shape))
print("Number of non-zero elements in integrate_link_importance: {}".format(np.count_nonzero(integrate_link_importance.todense())))
```
We can now find the nodes that have the highest importance to the prediction of the selected node:
```
sorted_indices = np.argsort(integrate_link_importance_dense.flatten())
N = len(graph_nodes)
integrated_link_importance_rank = [(k//N, k%N) for k in sorted_indices[::-1]]
topk = 10
#integrate_link_importance = integrate_link_importance_dense
print('Top {} most important links by integrated gradients are:\n {}'.format(topk, integrated_link_importance_rank[:topk]))
#Set the labels as an attribute for the nodes in the graph. The labels are used to color the nodes in different classes.
nx.set_node_attributes(
G_ego,
values={x[0]:{'subject': x[1]} for x in node_data['subject'].items()}
)
```
In the following, we plot the link and node importance (computed by integrated gradients) of the nodes within the ego graph of the target node.
For nodes, the shape of the node indicates the positive/negative importance the node has. 'round' nodes have positive importance while 'diamond' nodes have negative importance. The size of the node indicates the value of the importance, e.g., a large diamond node has higher negative importance.
For links, the color of the link indicates the positive/negative importance the link has. 'red' links have positive importance while 'blue' links have negative importance. The width of the link indicates the value of the importance, e.g., a thicker blue link has higher negative importance.
```
integrated_node_importance.max()
integrate_link_importance.max()
node_size_factor = 1e2
link_width_factor = 2
nodes = list(G_ego.nodes())
colors = pd.DataFrame([v[1]['subject'] for v in G_ego.nodes(data=True)],
index=nodes,
columns=['subject'])
colors = np.argmax(target_encoding.transform(colors.to_dict('records')), axis=1) + 1
fig, ax = plt.subplots(1, 1, figsize=(15, 10));
pos = nx.spring_layout(G_ego)
# Draw ego as large and red
node_sizes = [integrated_node_importance[graph_nodes.index(k)] for k in nodes]
node_shapes = ['o' if w > 0 else 'd' for w in node_sizes]
positive_colors, negative_colors = [], []
positive_node_sizes, negative_node_sizes = [], []
positive_nodes, negative_nodes = [], []
node_size_scale = node_size_factor/np.max(node_sizes)
for k in range(len(nodes)):
if nodes[k] == target_idx:
continue
if node_shapes[k] == 'o':
positive_colors.append(colors[k])
positive_nodes.append(nodes[k])
positive_node_sizes.append(node_size_scale*node_sizes[k])
else:
negative_colors.append(colors[k])
negative_nodes.append(nodes[k])
negative_node_sizes.append(node_size_scale*abs(node_sizes[k]))
# Plot the ego network with the node importances
cmap = plt.get_cmap('jet', np.max(colors)-np.min(colors)+1)
nc = nx.draw_networkx_nodes(G_ego, pos, nodelist=positive_nodes, node_color=positive_colors, cmap=cmap,
node_size=positive_node_sizes, with_labels=False,
vmin=np.min(colors)-0.5, vmax=np.max(colors)+0.5, node_shape='o')
nc = nx.draw_networkx_nodes(G_ego, pos, nodelist=negative_nodes, node_color=negative_colors, cmap=cmap,
node_size=negative_node_sizes, with_labels=False,
vmin=np.min(colors)-0.5, vmax=np.max(colors)+0.5, node_shape='d')
# Draw the target node as a large star colored by its true subject
nx.draw_networkx_nodes(G_ego, pos, nodelist=[target_nid], node_size=50*abs(node_sizes[nodes.index(target_nid)]),
node_shape='*', node_color=[colors[nodes.index(target_nid)]], cmap=cmap,
vmin=np.min(colors)-0.5, vmax=np.max(colors)+0.5, label="Target")
# Draw the edges with the edge importances
edges = G_ego.edges()
weights = [integrate_link_importance[graph_nodes.index(u),graph_nodes.index(v)] for u,v in edges]
edge_colors = ['red' if w > 0 else 'blue' for w in weights]
weights = link_width_factor*np.abs(weights)/np.max(weights)
ec = nx.draw_networkx_edges(G_ego, pos, edge_color=edge_colors, width=weights)
plt.legend()
plt.colorbar(nc, ticks=np.arange(np.min(colors),np.max(colors)+1))
plt.axis('off')
plt.show()
```
We then remove the node or edge in the ego graph one by one and check how the prediction changes. By doing so, we can obtain the ground truth importance of the nodes and edges. Comparing the following figure and the above one can show the effectiveness of integrated gradients as the importance approximations are relatively consistent with the ground truth.
```
(X, _, A_index, A), _ = train_gen[0]
X_bk = deepcopy(X)
A_bk = deepcopy(A)
selected_nodes = np.array([[target_idx]], dtype='int32')
nodes = [graph_nodes.index(v) for v in G_ego.nodes()]
edges = [(graph_nodes.index(u),graph_nodes.index(v)) for u,v in G_ego.edges()]
clean_prediction = model.predict([X, selected_nodes, A_index, A]).squeeze()
predict_label = np.argmax(clean_prediction)
groud_truth_node_importance = np.zeros((N,))
for node in nodes:
#we set all the features of the node to zero to check the ground truth node importance.
X_perturb = deepcopy(X_bk)
X_perturb[:, node, :] = 0
predict_after_perturb = model.predict([X_perturb, selected_nodes, A_index, A]).squeeze()
groud_truth_node_importance[node] = clean_prediction[predict_label] - predict_after_perturb[predict_label]
node_shapes = ['o' if groud_truth_node_importance[k] > 0 else 'd' for k in range(len(nodes))]
positive_colors, negative_colors = [], []
positive_node_sizes, negative_node_sizes = [], []
positive_nodes, negative_nodes = [], []
#node_size_scale is used for better visulization of nodes
node_size_scale = node_size_factor/max(groud_truth_node_importance)
for k in range(len(node_shapes)):
if nodes[k] == target_idx:
continue
if node_shapes[k] == 'o':
positive_colors.append(colors[k])
positive_nodes.append(graph_nodes[nodes[k]])
positive_node_sizes.append(node_size_scale*groud_truth_node_importance[nodes[k]])
else:
negative_colors.append(colors[k])
negative_nodes.append(graph_nodes[nodes[k]])
negative_node_sizes.append(node_size_scale*abs(groud_truth_node_importance[nodes[k]]))
X = deepcopy(X_bk)
groud_truth_edge_importance = np.zeros((N,N))
G_edge_indices = [(A_index[0, k, 0], A_index[0, k, 1]) for k in range(A.shape[1])]
for edge in edges:
edge_index = G_edge_indices.index((edge[0], edge[1]))
origin_val = A[0, edge_index]
A[0, edge_index] = 0
#we set the weight of a given edge to zero to check the ground truth link importance
predict_after_perturb = model.predict([X, selected_nodes, A_index, A]).squeeze()
groud_truth_edge_importance[edge[0], edge[1]] = (predict_after_perturb[predict_label]
- clean_prediction[predict_label])/(0 - 1)
A[0, edge_index] = origin_val
fig, ax = plt.subplots(1, 1, figsize=(15, 10));
cmap = plt.get_cmap('jet', np.max(colors)-np.min(colors)+1)
# Draw the target node as a large star colored by its true subject
nx.draw_networkx_nodes(G_ego, pos, nodelist=[target_nid], node_size=50*abs(node_sizes[nodes.index(target_idx)]),
node_color=[colors[nodes.index(target_idx)]], cmap=cmap,
node_shape='*', vmin=np.min(colors)-0.5, vmax=np.max(colors)+0.5, label="Target")
# Draw the ego net
nc = nx.draw_networkx_nodes(G_ego, pos, nodelist=positive_nodes, node_color=positive_colors, cmap=cmap,
node_size=positive_node_sizes, with_labels=False,
vmin=np.min(colors)-0.5, vmax=np.max(colors)+0.5, node_shape='o')
nc = nx.draw_networkx_nodes(G_ego, pos, nodelist=negative_nodes, node_color=negative_colors, cmap=cmap,
node_size=negative_node_sizes, with_labels=False,
vmin=np.min(colors)-0.5, vmax=np.max(colors)+0.5, node_shape='d')
edges = G_ego.edges()
weights = [groud_truth_edge_importance[graph_nodes.index(u),graph_nodes.index(v)] for u,v in edges]
edge_colors = ['red' if w > 0 else 'blue' for w in weights]
weights = link_width_factor*np.abs(weights)/np.max(weights)
ec = nx.draw_networkx_edges(G_ego, pos, edge_color=edge_colors, width=weights)
plt.legend()
plt.colorbar(nc, ticks=np.arange(np.min(colors),np.max(colors)+1))
plt.axis('off')
plt.show()
```
By comparing the above two figures, one can see that the integrated gradients are quite consistent with the brute-force approach. The main benefit of using integrated gradients is scalability. The gradient operations are very efficient to compute on deep learning frameworks with the parallism provided by GPUs. Also, integrated gradients can give the importance of individual node features, for all nodes in the graph. Achieving this by brute-force approch is often non-trivial.
| github_jupyter |
# Tutorial 14: Multiagent
This tutorial covers the implementation of multiagent experiments in Flow. It assumes some level of knowledge or experience in writing custom environments and running experiments with RLlib. The rest of the tutorial is organized as follows. Section 1 describes the procedure through which custom environments can be augmented to generate multiagent environments. Then, section 2 walks you through an example of running a multiagent environment
in RLlib.
本教程介绍了流中多代理实验的实现。它假定您具有编写自定义环境和使用RLlib运行实验方面的一定知识或经验。本教程的其余部分组织如下。第1节描述了通过扩展自定义环境来生成多代理环境的过程。然后,第2节将介绍一个运行多代理环境的示例
在RLlib。
## 1. Creating a Multiagent Environment Class 创建一个多代理环境类
In this part we will be setting up steps to create a multiagent environment. We begin by importing the abstract multi-agent evironment class.
在本部分中,我们将设置创建多代理环境的步骤。我们首先导入抽象的多代理环境类。
```
# import the base Multi-agent environment
from flow.envs.multiagent.base import MultiEnv
```
In multiagent experiments, the agent can either share a policy ("shared policy") or have different policies ("non-shared policy"). In the following subsections, we describe the two.
在多代理实验中,代理可以共享一个策略(“共享策略”),也可以拥有不同的策略(“非共享策略”)。在下面的小节中,我们将介绍这两种方法。
### 1.1 Shared policies 共享策略
In the multi-agent environment with a shared policy, different agents will use the same policy.
在具有共享策略的多代理环境中,不同的代理将使用相同的策略。
We define the environment class, and inherit properties from the Multi-agent version of base env.
我们定义了environment类,并从基本环境的多代理版本中继承属性。
```
class SharedMultiAgentEnv(MultiEnv):
pass
```
This environment will provide the interface for running and modifying the multiagent experiment. Using this class, we are able to start the simulation (e.g. in SUMO), provide a network to specify a configuration and controllers, perform simulation steps, and reset the simulation to an initial configuration.
该环境将提供运行和修改多代理实验的接口。使用这个类,我们可以启动模拟(例如在SUMO中),提供一个网络来指定配置和控制器,执行模拟步骤,并将模拟重置为初始配置。
For the multi-agent experiments, certain functions of the `MultiEnv` will be changed according to the agents. Some functions will be defined according to a *single* agent, while the other functions will be defined according to *all* agents.
在多主体实验中,“MultiEnv”的某些功能会根据主体的不同而改变。一些函数将根据*单个*代理定义,而其他函数将根据*所有*代理定义。
In the follwing functions, observation space and action space will be defined for a *single* agent (not all agents):
在下面的功能中,将定义一个*单个* agent(不是所有agent)的观察空间和行动空间:
* **observation_space**
* **action_space**
For instance, in a multiagent traffic light grid, if each agents is considered as a single intersection controlling the traffic lights of the intersection, the observation space can be define as *normalized* velocities and distance to a *single* intersection for nearby vehicles, that is defined for every intersection.
例如,在一个多智能体交通信号灯网格中,如果每个智能体都被视为一个控制该交叉口交通灯的单个交叉口,那么对于附近车辆,观测空间可以定义为*归一化*速度和到*单个*交叉口的距离,即为每个交叉口定义。
```
def observation_space(self):
"""State space that is partially observed.
Velocities and distance to intersections for nearby
vehicles ('num_observed') from each direction.
"""
tl_box = Box(
low=0.,
high=1,
shape=(2 * 4 * self.num_observed),
dtype=np.float32)
return tl_box
```
The action space can be defined for a *single* intersection as follows
可以为一个*单个*交集定义操作空间,如下所示
```
def action_space(self):
"""See class definition."""
if self.discrete:
# each intersection is an agent, and the action is simply 0 or 1.
# - 0 means no-change in the traffic light
# - 1 means switch the direction
return Discrete(2)
else:
return Box(low=0, high=1, shape=(1,), dtype=np.float32)
```
Conversely, the following functions (including their return values) will be defined to take into account *all* agents:
相反,下面的函数(包括它们的返回值)将被定义为考虑*所有*代理:
* **apply_rl_actions**
* **get_state**
* **compute_reward**
Instead of calculating actions, state, and reward for a single agent, in these functions, the ctions, state, and reward will be calculated for all the agents in the system. To do so, we create a dictionary with agent ids as keys and different parameters (actions, state, and reward ) as vaules. For example, in the following `_apply_rl_actions` function, based on the action of intersections (0 or 1), the state of the intersections' traffic lights will be changed.
在这些函数中,将计算系统中所有代理的动作、状态和奖励,而不是计算单个代理的动作、状态和奖励。为此,我们创建了一个字典,其中代理id作为键,不同的参数(动作、状态和奖励)作为变量。例如,在下面的‘_apply_rl_actions’函数中,根据交叉口(0或1)的动作,将改变交叉口交通灯的状态。
```
class SharedMultiAgentEnv(MultiEnv):
def _apply_rl_actions(self, rl_actions):
for agent_name in rl_actions:
action = rl_actions[agent_name]
# check if the action space is discrete
# check if our timer has exceeded the yellow phase, meaning it
# should switch to red
if self.currently_yellow[tl_num] == 1: # currently yellow
self.last_change[tl_num] += self.sim_step
if self.last_change[tl_num] >= self.min_switch_time: # check if our timer has exceeded the yellow phase, meaning it
# should switch to red
if self.direction[tl_num] == 0:
self.k.traffic_light.set_state(
node_id='center{}'.format(tl_num),
state="GrGr")
else:
self.k.traffic_light.set_state(
node_id='center{}'.format(tl_num),
state='rGrG')
self.currently_yellow[tl_num] = 0
else:
if action:
if self.direction[tl_num] == 0:
self.k.traffic_light.set_state(
node_id='center{}'.format(tl_num),
state='yryr')
else:
self.k.traffic_light.set_state(
node_id='center{}'.format(tl_num),
state='ryry')
self.last_change[tl_num] = 0.0
self.direction[tl_num] = not self.direction[tl_num]
self.currently_yellow[tl_num] = 1
```
Similarly, the `get_state` and `compute_reward` methods support the dictionary structure and add the observation and reward, respectively, as a value for each correpsonding key, that is agent id.
类似地,' get_state '和' compute_reward '方法支持字典结构,并分别为每个correpsonding键(即代理id)添加观察值和奖赏值。
```
class SharedMultiAgentEnv(MultiEnv):
def get_state(self):
"""Observations for each intersection
:return: dictionary which contains agent-wise observations as follows:
- For the self.num_observed number of vehicles closest and incomingsp
towards traffic light agent, gives the vehicle velocity and distance to
intersection.
"""
# Normalization factors
max_speed = max(
self.k.network.speed_limit(edge)
for edge in self.k.network.get_edge_list())
max_dist = max(grid_array["short_length"], grid_array["long_length"],
grid_array["inner_length"])
# Observed vehicle information
speeds = []
dist_to_intersec = []
for _, edges in self.network.node_mapping:
local_speeds = []
local_dists_to_intersec = []
# .... More code here (removed for simplicity of example)
# ....
speeds.append(local_speeds)
dist_to_intersec.append(local_dists_to_intersec)
obs = {}
for agent_id in self.k.traffic_light.get_ids():
# .... More code here (removed for simplicity of example)
# ....
observation = np.array(np.concatenate(speeds, dist_to_intersec))
obs.update({agent_id: observation})
return obs
def compute_reward(self, rl_actions, **kwargs):
if rl_actions is None:
return {}
if self.env_params.evaluate:
rew = -rewards.min_delay_unscaled(self)
else:
rew = -rewards.min_delay_unscaled(self) \
+ rewards.penalize_standstill(self, gain=0.2)
# each agent receives reward normalized by number of lights
rew /= self.num_traffic_lights
rews = {}
for rl_id in rl_actions.keys():
rews[rl_id] = rew
return rews
```
### 1.2 Non-shared policies 非共享策略
In the multi-agent environment with a non-shared policy, different agents will use different policies. In what follows we will see the two agents in a ring road using two different policies, 'adversary' and 'av' (non-adversary).
在具有非共享策略的多代理环境中,不同的代理将使用不同的策略。在接下来的内容中,我们将看到这两个代理在一个环形路上使用了两种不同的策略,“对手”和“av”(非对手)。
Similarly to the shared policies, observation space and action space will be defined for a *single* agent (not all agents):
与共享策略类似,将为一个*单个*代理(不是所有代理)定义观察空间和操作空间:
* **observation_space**
* **action_space**
And, the following functions (including their return values) will be defined to take into account *all* agents::
并且,以下函数(包括它们的返回值)将被定义为考虑*所有*代理::
* **apply_rl_actions**
* **get_state**
* **compute_reward**
\* Note that, when observation space and action space will be defined for a single agent, it means that all agents should have the same dimension (i.e. space) of observation and action, even when their policise are not the same.
请注意,当为单个agent定义了观察空间和行动空间时,这意味着所有agent都应该具有相同的观察和行动维度(即空间),即使它们的策略并不相同。
Let us start with defining `apply_rl_actions` function. In order to make it work for a non-shared policy multi-agent ring road, we define `rl_actions` as a combinations of each policy actions plus the `perturb_weight`.
让我们从定义' apply_rl_actions '函数开始。为了使它适用于非共享策略多代理环路,我们将“rl_actions”定义为每个策略操作加上“扰动权值”的组合。
```
class NonSharedMultiAgentEnv(MultiEnv):
def _apply_rl_actions(self, rl_actions):
# the names of all autonomous (RL) vehicles in the network
agent_ids = [
veh_id for veh_id in self.sorted_ids
if veh_id in self.k.vehicle.get_rl_ids()
]
# define different actions for different multi-agents
av_action = rl_actions['av']
adv_action = rl_actions['adversary']
perturb_weight = self.env_params.additional_params['perturb_weight']
rl_action = av_action + perturb_weight * adv_action
# use the base environment method to convert actions into accelerations for the rl vehicles
self.k.vehicle.apply_acceleration(agent_ids, rl_action)
```
In the `get_state` method, we define the state for each of the agents. Remember, the sate of the agents can be different. For the purpose of this example and simplicity, we define the state of the adversary and non-adversary agent to be the same.
在“get_state”方法中,我们为每个代理定义状态。记住,代理的状态可以是不同的。为了这个示例和简单起见,我们将对手和非对手代理的状态定义为相同的。
In the `compute_reward` method, the agents receive opposing speed rewards. The reward of the adversary agent is more when the speed of the vehicles is small, while the non-adversary agent tries to increase the speeds of the vehicles.
在“compute_reward”方法中,代理收到相反的速度奖励。当车辆的速度较小时,敌手的奖励较多,而非敌手则试图提高车辆的速度。
```
class NonSharedMultiAgentEnv(MultiEnv):
def get_state(self, **kwargs):
state = np.array([[
self.k.vehicle.get_speed(veh_id) / self.k.network.max_speed(),
self.k.vehicle.get_x_by_id(veh_id) / self.k.network.length()
] for veh_id in self.sorted_ids])
state = np.ndarray.flatten(state)
return {'av': state, 'adversary': state}
def compute_reward(self, rl_actions, **kwargs):
if self.env_params.evaluate:
reward = np.mean(self.k.vehicle.get_speed(
self.k.vehicle.get_ids()))
return {'av': reward, 'adversary': -reward}
else:
reward = rewards.desired_velocity(self, fail=kwargs['fail'])
return {'av': reward, 'adversary': -reward}
```
## 2. Running Multiagent Environment in RLlib 在RLlib中运行多代理环境
When running the experiment that uses a multiagent environment, we specify certain parameters in the `flow_params` dictionary.
在运行使用多代理环境的实验时,我们在“flow_params”字典中指定某些参数。
Similar to any other experiments, the following snippets of codes will be inserted into a blank python file (e.g. `new_multiagent_experiment.py`, and should be saved under `flow/examples/exp_configs/rl/multiagent/` directory. (all the basic imports and initialization of variables are omitted in this example for brevity)
与其他实验类似,下面的代码片段将被插入到一个空白的python文件中(例如“new_multiagent_experiment。,应该保存在“flow/examples/exp_configs/rl/multiagent/”目录下。(为了简单起见,本例中省略了所有基本的变量导入和初始化)
```
from flow.envs.multiagent import MultiWaveAttenuationPOEnv
from flow.networks import MultiRingNetwork
from flow.core.params import SumoParams, EnvParams, NetParams, VehicleParams, InitialConfig
from flow.controllers import ContinuousRouter, IDMController, RLController
# time horizon of a single rollout
HORIZON = 3000
# Number of rings
NUM_RINGS = 1
vehicles = VehicleParams()
for i in range(NUM_RINGS):
vehicles.add(
veh_id='human_{}'.format(i),
acceleration_controller=(IDMController, {
'noise': 0.2
}),
routing_controller=(ContinuousRouter, {}),
num_vehicles=21)
vehicles.add(
veh_id='rl_{}'.format(i),
acceleration_controller=(RLController, {}),
routing_controller=(ContinuousRouter, {}),
num_vehicles=1)
flow_params = dict(
# name of the experiment
exp_tag='multiagent_ring_road',
# name of the flow environment the experiment is running on
env_name=MultiWaveAttenuationPOEnv,
# name of the network class the experiment is running on
network=MultiRingNetwork,
# simulator that is used by the experiment
simulator='traci',
# sumo-related parameters (see flow.core.params.SumoParams)
sim=SumoParams(
sim_step=0.1,
render=False,
),
# environment related parameters (see flow.core.params.EnvParams)
env=EnvParams(
horizon=HORIZON,
warmup_steps=750,
additional_params={
'max_accel': 1,
'max_decel': 1,
'ring_length': [230, 230],
'target_velocity': 4
},
),
# network-related parameters
net=NetParams(
additional_params={
'length': 230,
'lanes': 1,
'speed_limit': 30,
'resolution': 40,
'num_rings': NUM_RINGS
},
),
# vehicles to be placed in the network at the start of a rollout
veh=vehicles,
# parameters specifying the positioning of vehicles upon initialization/
# reset
initial=InitialConfig(bunching=20.0, spacing='custom'),
)
```
Then we run the following code to create the environment
```
from flow.utils.registry import make_create_env
from ray.tune.registry import register_env
create_env, env_name = make_create_env(params=flow_params, version=0)
# Register as rllib env
register_env(env_name, create_env)
test_env = create_env()
obs_space = test_env.observation_space
act_space = test_env.action_space
```
### 2.1 Shared policies 共享策略
When we run a shared-policy multiagent experiment, we refer to the same policy for each agent. In the example below the agents will use 'av' policy.
当我们运行共享策略多代理实验时,我们为每个代理引用相同的策略。在下面的示例中,代理将使用“av”策略。
```
from ray.rllib.agents.ppo.ppo_policy import PPOTFPolicy
def gen_policy():
"""Generate a policy in RLlib."""
return PPOTFPolicy, obs_space, act_space, {}
# Setup PG with an ensemble of `num_policies` different policy graphs
POLICY_GRAPHS = {'av': gen_policy()}
def policy_mapping_fn(_):
"""Map a policy in RLlib."""
return 'av'
POLICIES_TO_TRAIN = ['av']
```
### 2.2 Non-shared policies 非共享策略
When we run the non-shared multiagent experiment, we refer to different policies for each agent. In the example below, the policy graph will have two policies, 'adversary' and 'av' (non-adversary).
当我们运行非共享多代理实验时,我们引用每个代理的不同策略。在下面的示例中,策略图将有两个策略,“对手”和“av”(非对手)。
```
def gen_policy():
"""Generate a policy in RLlib."""
return PPOTFPolicy, obs_space, act_space, {}
# Setup PG with an ensemble of `num_policies` different policy graphs
POLICY_GRAPHS = {'av': gen_policy(), 'adversary': gen_policy()}
def policy_mapping_fn(agent_id):
"""Map a policy in RLlib."""
return agent_id
```
Lastly, just like any other experiments, we run our code using `train_rllib.py` as follows:
最后,与其他实验一样,我们使用' train_rllib运行代码。py”如下:
python flow/examples/train_rllib.py new_multiagent_experiment.py
| github_jupyter |
## Licence
```
Copyright 2017 Despoina Antonakaki despoina@ics.forth.gr
Licensed under the Apache License, Version 2.0 (the "License");
you may not use this file except in compliance with the License.
You may obtain a copy of the License at
http://www.apache.org/licenses/LICENSE-2.0
Unless required by applicable law or agreed to in writing, software
distributed under the License is distributed on an "AS IS" BASIS,
WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
See the License for the specific language governing permissions and
limitations under the License.
```
## A note on data sources
### Twitter data
* Original tweets are not available due to Twitter's [Developer Agreement & Policy](https://dev.twitter.com/overview/terms/agreement-and-policy). For specific information please read [this paragraph](https://dev.twitter.com/overview/terms/agreement-and-policy#f-be-a-good-partner-to-twitter).
### SentiStrength Lexicon
This study introduces and makes use of a new Greek [SentiStrength](http://sentistrength.wlv.ac.uk/) lexicon. This lexicon combines terms from three different lexicons:
* The Greek lexicon that is available on SentiStrength's website. More information regarding non-English SentiStrength Lexicons are here: http://sentistrength.wlv.ac.uk/#Non-English. The file that contains this lexicon is available on this zip: http://sentistrength.wlv.ac.uk/9LanguageSetsPoor.zip. After downloading and extracting, the file is located at /greek/EmotionLookupTable.txt
* A Greek sentiment lexicon that was created with the support of the EC-funded FP7 Project [SocialSensor](http://socialsensor.eu/). The location of the lexicon is here: https://github.com/MKLab-ITI/greek-sentiment-lexicon . Please note that this Lexicon is available under the terms of the Apache License.
* A custom lexicon with terms from the political domain of this analysis.
This new lexicon is NOT necessary for the reproduction of the analysis presented here since we provide the files that contain the combined sentiment for each tweet (according to the new Lexicon). Nevertheless we do plan to make this Lexicon available from the official site of SentiStrength in the near future. When this happens we will also update this notebook.
## Useful imports
```
import os
import re # Regular Expressions
import json
import time
import codecs # To be able to read unicode files
import datetime
import itertools
from collections import Counter
%matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
import seaborn
import numpy as np
import pandas as pd
# https://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.mannwhitneyu.html
# https://en.wikipedia.org/wiki/Mann%E2%80%93Whitney_U_test
from scipy.stats import mannwhitneyu
```
## Fetch data
Data are available on the following repository: https://figshare.com/articles/Social_media_analysis_during_political_turbulence_DATA/5492443
Please make note that Original tweets are not available due to [Twitter's Terms and Conditions](https://dev.twitter.com/overview/terms/agreement-and-policy).
```
data_dir = '../../referendum_elections/' # Change this to the desired data location
# Original tweets are not available due to Twitter's Terms and Conditions.
# https://dev.twitter.com/overview/terms/agreement-and-policy
data = {
'entities_from_hashtags' : {
#'url' : 'https://www.dropbox.com/s/u0annzjim9q07yk/cutdown_entities_from_hashtags_v20160420_2.txt?dl=1',
'basename': 'cutdown_entities_from_hashtags_v20160420_2.txt',
},
'entities_from_hashtags_old_1': {
#'url': 'https://www.dropbox.com/s/349ct9k6q5oifp6/hashtags_for_entities_v20160405.txt?dl=1',
'basename': 'hashtags_for_entities_v20160405.txt',
},
'entities_from_text': {
#'url' : 'https://www.dropbox.com/s/t0l1d079feexifz/cutdown_entities_from_text_v20160419.txt?dl=1',
'basename': 'cutdown_entities_from_text_v20160419.txt',
},
'sarcasm': {
#'url': 'https://www.dropbox.com/s/hn5sb394edwbsfz/ola_text_classified.txt?dl=1',
'basename': 'ola_text_classified.txt',
},
'elections_tweets': {
#'url': None, # Not available due to Twitter's Terms and Conditions
'basename': 'ht_sorted_unique.json',
},
'referendum_tweets': {
#'url': None, # Not available due to Twitter's Terms and Conditions
'basename': 'ht_common_final_greek_sorted_reversed.json',
},
'elections_sentiment': {
#'url': 'https://www.dropbox.com/s/psneegp3kuq9vgu/ht_sorted_unique_with_SENTIMENT_20160419.txt?dl=1',
'basename': 'ht_sorted_unique_with_SENTIMENT_20160419.txt',
},
'referendum_sentiment': {
#'url': 'https://www.dropbox.com/s/p7rt5e5f6snn069/ht_common_final_greek_sorted_reversed_with_SENTIMENT_20160419.txt?dl=1',
'basename': 'ht_common_final_greek_sorted_reversed_with_SENTIMENT_20160419.txt',
},
}
```
### Make sure that data exist
```
for data_item, data_entry in data.iteritems():
#Check if data exist
data_filename= data_filename = os.path.join(data_dir, data_entry['basename'])
data[data_item]['filename'] = data_filename
if os.path.exists(data_filename):
print "{} --> {} exists".format(data_item, data_filename)
else:
print "WARNING!! FILENAME: {} does not exist!".format(data_filename)
```
## Preprocess Greek text
```
greek_punctuation_replacement = [[u'\u03ac', u'\u03b1'], [u'\u03ad', u'\u03b5'], [u'\u03ae', u'\u03b7'], [u'\u03af', u'\u03b9'], [u'\u03ca', u'\u03b9'], [u'\u0390', u'\u03b9'], [u'\u03cc', u'\u03bf'], [u'\u03cd', u'\u03c5'], [u'\u03cb', u'\u03c5'], [u'\u03b0', u'\u03c5'], [u'\u03ce', u'\u03c9']]
```
This looks like this:
```
for original, replacement in greek_punctuation_replacement:
print original, replacement
greek_punctuation = [x[0] for x in greek_punctuation_replacement]
sigma_teliko = (u'\u03c2', u'\u03c3') # (u'ς', u'σ')
def remove_greek_punctuation(text):
'''
This is function removes greek punctuation
! NOT TEXT! (sigmal teliko)
'''
ret = []
for x in text:
x_lower = x.lower()
if x_lower in greek_punctuation:
ret += [greek_punctuation_replacement[greek_punctuation.index(x_lower)][1]]
else:
ret += [x_lower]
# Fix sigma teliko
if len(ret) > 0:
if ret[-1] == sigma_teliko[0]:
ret[-1] = sigma_teliko[1]
return u''.join(ret)
```
For example:
```
print remove_greek_punctuation(u"Ευχάριστος")
def remove_punctuation(text):
punctuation = u''',./;'\][=-§`<>?:"|}{+_±~!@$%^&*()'''
return u''.join([x for x in text if not x in punctuation])
```
## Load entities
```
def load_entities(filename):
'''
Reads a filename with entities
'''
ret = {} # Item to return
print 'Opening entities filename:', filename
with codecs.open(filename, encoding='utf8') as f:
for l in f:
ls = l.replace(os.linesep, '').split() # Split entries
ls = [remove_greek_punctuation(x) for x in ls] # normalize words
#Some words may be present as hashtags. So we remove the '#' in the front
ls = [x[1:] if x[0] == '#' else x for x in ls]
# The first column is the entity name
main_entity = ls[0]
# The rest columns are synonyms of this entity
for entity in ls[1:]:
ret[entity] = ret.get(entity, set()) | {main_entity} # Create a set of all main_entities for each entity
return ret
```
Let's create a single variable that contains all entites (from text and from hashtags)
```
entities = {
'hashtags' : load_entities(data['entities_from_hashtags']['filename']),
'text': load_entities(data['entities_from_text']['filename'])
}
all_entities = set([y for x in entities['hashtags'].values() for y in x]) | set([y for x in entities['text'].values() for y in x])
print 'All entities: ', len(all_entities)
```
## YES/NO entities for referendum
```
def load_yes_no_entities():
#Load older version of hastags
ht = {}
with codecs.open(data['entities_from_hashtags_old_1']['filename'], encoding='utf8') as f:
print 'Using:', f.name
for l in f:
ls = l.replace(os.linesep, '').split()
if ls[0][0] == '#':
ht[ls[0][1:]] = ls[1:] # Remove #
else:
ht[ls[0]] = ls[1:] # DO NOT Remove #
ht2 = {}
ht2['yes'] = list(set(ht['voteYes'] + ht['YesToEurope']))
ht2['no'] = list(set(ht['voteNo'] + ht['NoToEurope']))
del ht
ht = ht2
return ht
yes_no_entities = load_yes_no_entities()
```
## Load sarcasm
Sarcasm is in a JSON file with the following format:
```
{"text": "#debate #enikos #debate2015 #ERTdebate2015 #ekloges2015_round2 #alphaekloges #ekloges \u039c\u03b5\u03b3\u03ba\u03b1 #mega https://t.co/ufUG57w037", "percentage": 3, "score": 0.069323771015285293}
```
```
def load_sarcasm():
ret = {} # Item to return
with open(data['sarcasm']['filename']) as f:
for l in f:
j = json.loads(l) # Read JSON line
# We construct a dictionary were the key is the text of the tweet!
ret[j['text']] = {'score': j['score'], 'percentage' : j['percentage']}
print "{} TWEETS READ".format(len(ret))
return ret
sarcasm = load_sarcasm()
```
We need a function that will adjust the sentiment of a tweet according to the sarcasm percentage
```
def correct_sarcasm(sentiment, sarcasm_percentage):
ret = sentiment
if sarcasm_percentage>0.0: # Correct
if sentiment > 0.0:
sentiment_value = (ret-1) / 4.0
elif sentiment <0.0:
sentiment_value = (-ret-1) / 4.0
else:
raise Exception('This should not happen 491266')
#Penalize sentiment
sentiment_value = sentiment_value - ((sarcasm_percentage/100.0) * sentiment_value)
#Extend to previous range
sentiment_value = 1 + (sentiment_value * 4.0)
#Assign right sign
if sentiment < 0:
sentiment_value = -sentiment_value
ret = sentiment_value
return ret
```
## Read Sentiment
Example of entry for sentiment file:
```
612662791447687169 η εε αγωνιά αν θα πείσει τον τσίπρα κι αυτός τον συριζα το ζητημα πρέπει να είναι να πείσουν εμάς οι πολιτες να αποφασίσουν #dimopsifisma 1 -2 -1
```
```
def sentiment_generator(filename, print_problem=False):
print 'Opening sentiment filename:', filename
parse_errors = 0
c = 0 # Line counter
with open(filename) as f:
for l in f:
c += 1
ls = l.decode('utf8').split()
text = ' '.join(ls[1:-3]) # Text is the concatenation of all columns from 2 to -3
#Create the object that we will yield
to_yield = {
'sum' : int(ls[-1]),
'neg' : int(ls[-2]),
'pos' : int(ls[-3]),
'text' : text,
'hashtags' : re.findall(r'#(\w+)', text, re.UNICODE),
}
try:
this_id = int(ls[0])
except:
parse_errors += 1
continue # Ignore lines with parse error
to_yield['id'] = this_id
if this_id == 1: # for some reason this happens...
parse_errors += 1
if print_problem:
print 'Problem:'
print l
print 'Line:', c
continue # Ignore these lines
yield to_yield
```
## Read Tweets
Create a "tweet generator" from the files
```
def tweet_generator(filename):
print 'Reading json tweets from:', filename
with open(filename) as f:
for l in f:
lj = json.loads(l)
yield lj
```
Create useful functions for reading data from Twitter JSON objects
```
def tweet_get_text(j):
return j['text']
def tweet_get_id(j):
return j['id']
def tweet_get_user(j):
return j['user']['screen_name']
def tweet_get_hashtags(j):
return [x['text'] for x in j['entities']['hashtags']]
def tweet_get_date_text(j):
return j['created_at']
def tweet_get_date_epochs(j):
return time.mktime(time.strptime(j['created_at'], '%a %b %d %H:%M:%S +0000 %Y'))
def nice_print_tweet(j):
print json.dumps(j, indent=4)
def get_retweet_status(status):
if u'retweeted_status' in status:
return status[u'retweeted_status']
return None
def is_retweet(status):
# print '------------', status[u'retweeted'], get_retweet_status(status)
# return status[u'retweeted']
return not get_retweet_status(status) is None
def is_reply(status):
return not status['in_reply_to_status_id'] is None
def get_retweet_text(status):
if is_retweet(status):
return status[u'retweeted_status'][u'text']
return None
def get_original_text(status):
'''
Get the "original" text of the tweet if this a retweet
Otherwise return the normal text
'''
text = get_retweet_text(status)
if text is None:
text = tweet_get_text(status)
return text
def is_greek_text(text):
'''
a "greek" tweet should contain at least one greek character
'''
greek_letters = '\xce\xb1\xce\xb2\xce\xb3\xce\xb4\xce\xb5\xce\xb6\xce\xb7\xce\xb8\xce\xb9\xce\xba' \
'\xce\xbb\xce\xbc\xce\xbd\xce\xbe\xce\xbf\xcf\x80\xcf\x81\xcf\x83\xcf\x84\xcf\x85\xcf\x86\xcf\x87' \
'\xcf\x88\xcf\x89\xce\x91\xce\x92\xce\x93\xce\x94\xce\x95\xce\x96\xce\x97\xce\x98\xce\x99\xce\x9a' \
'\xce\x9b\xce\x9c\xce\x9d\xce\x9e\xce\x9f\xce\xa0\xce\xa1\xce\xa3\xce\xa4\xce\xa5\xce\xa6\xce\xa7' \
'\xce\xa8\xce\xa9'.decode('utf-8')
for x in text:
if x in greek_letters:
return True
return False
def convert_unix_to_human(unix_time, string_time='%a %H:%M'):
'''
Convert the unix epoch to a human readable format (in Greece)
%d/%m == 15/4
'''
offset = 60.0 * 60.0 * 3 # Three hours.
# This is the time difference between UTC (Twitter's time) and Greece.
# This the the +2 hours difference of EET plus 1 hour for daylight savings time (applied in Greece)
# Both Referendum and Elections happened during daylight saving time
# This is to make sure that when date is plotted, it is aligned to "Greek" time.
return time.strftime(string_time, time.localtime(unix_time + offset))
def get_json(input_filename):
print 'Reading json tweets from:', input_filename
with open(input_filename) as f:
for l in f:
lj = json.loads(l)
yield lj
def print_now_2():
'''
Print current time
'''
return time.strftime("%a, %d %b %Y %H:%M:%S +0000", time.gmtime())
```
## Merge Tweets and Sentiment
Create a "merged" generator for tweets and sentiment
```
def merge_tweets_sentiment(kind):
'''
kind is either 'elections' or 'referendum'
'''
accepted = ['elections', 'referendum']
if not kind in accepted:
raise Exception('kind parameter should be one of {}'.format(accepted))
tweets = '{}_tweets'.format(kind)
sentiment = '{}_sentiment'.format(kind)
missed_replies = 0 # See comments below
not_greek = 0 # See comments below
c = 0 # Line counter
for j, s in itertools.izip(
tweet_generator(data[tweets]['filename']),
sentiment_generator(data[sentiment]['filename'])):
c += 1
#Print some progress stats..
if c % 10000 == 0:
print 'Lines:', c
id_ = tweet_get_id(j) # This id of the tweet (id is reserved in python)
# These two generator should be in sync
assert id_ == s['id']
#Get all usable elements
text = tweet_get_text(j)
original_text = get_original_text(j)
text_splitted = text.split()
text_splitted = [remove_greek_punctuation(remove_punctuation(x)) for x in text_splitted]
hashtags = tweet_get_hashtags(j)
hashtags = [remove_greek_punctuation(x) for x in hashtags]
date_epochs = tweet_get_date_epochs(j)
username = tweet_get_user(j)
#Get text entities
text_entities = [list(entities['text'][w]) for w in text_splitted if w in entities['text']]
#Flat text entities and take single entries
text_entities = list(set([y for x in text_entities for y in x]))
#Get hashtag entities
hashtag_entites = [list(entities['hashtags'][w]) for w in hashtags if w in entities['hashtags']]
#Flat hashtag entities and take single entries
hashtag_entites = list(set([y for x in hashtag_entites for y in x]))
hashtag_entites = [x for x in hashtag_entites if x not in text_entities] # Remove hashtag entities that are text entities
# For some reply tweets we do not have sarcasm information
if is_reply(j):
if not original_text in sarcasm:
missed_replies += 1
continue
# Ignore tweets that are not Greek!!
# Especially in referendum (worldwide interest!), there are many english/foreign tweets
# Nevertheless all out lexicons (entities, ...) are tailored for greek text analysis
# Referendum tweets dataset contains only greek
if not is_greek_text(original_text):
not_greek += 1
continue
# Create a dictionary that contains info from tweets + sentiment + sarcasm
tweet_item = {
'id' : id_,
'ht' : hashtags,
'date' : date_epochs,
'sn' : s['neg'],
'sp' : s['pos'],
's0' : s['sum'],
'un' : username,
'sarcasm' : sarcasm[original_text],
'hashtag_entites' : hashtag_entites,
'text_entities' : text_entities,
'original_text' : original_text,
}
# Yield it!
yield tweet_item
# Print errors
print 'Missed replies:', missed_replies
print 'Not greek tweets:', not_greek
```
## Plotting helper functions
Useful function to convert the labels of ticks from epoch to human understandable dates
```
def fix_time_ticks(ax, fontsize=None, string_time='%a %H:%M', apply_index=lambda x : x[1]):
'''
Make time ticks from unix time
string_time='%e %b') # 4 Jul
'''
ticks = ax.get_xticks().tolist()
print ticks
new_ticks = [convert_unix_to_human(x, string_time) for x in ticks]
new_ticks = map(apply_index, enumerate(new_ticks))
print new_ticks
if fontsize is None:
ax.set_xticklabels(new_ticks)
else:
print 'Using fontsize:', fontsize
ax.set_xticklabels(new_ticks, fontsize=fontsize)
for tick in ax.xaxis.get_major_ticks(): # http://stackoverflow.com/questions/6390393/matplotlib-make-tick-labels-font-size-smaller
tick.label.set_fontsize(fontsize)
def save_figure(output_filename, formats=['eps', 'png']):
for ext in formats:
output_filename_tmp = output_filename + '.' + ext
print 'Output filename:', output_filename_tmp
plt.savefig(output_filename_tmp)
```
## Examine correlation between twitter volume and YES/NO vote
Reviewer's comment:
One of the major complaints of Syriza members before the referendum was that the media
were strongly favoring the "YES" voices and suppressing the "NO". It would be interesting
to see if there are statistically significant differences in how polarized are the Tweets
coming from regular citizens (without obvious political affiliation) and the Tweets coming
from various media outlets and journalists, and measure whether that complaint can be
substantiated by analyzing the Tweets. That's obviously a suggestion for future work.
Create functions to assign "YES" or "NO" vote
```
def yes_no_vote_1(username):
'''
Is yes/no only iff she has has yes/no votes and no opposite vote
'''
entities = referendum_user_stats[username]['e'].keys()
if ('voteyes' in entities) and (not 'voteno' in entities):
return (1,0)
elif (not 'voteyes' in entities) and ('voteno' in entities):
return (0,1)
return (0,0)
def yes_no_vote_2(username):
'''
Examine the yes/no hashtags (instead of entities)
'''
y_voter = referendum_user_stats[username]['y_voter']
n_voter = referendum_user_stats[username]['n_voter']
if y_voter > 0 and n_voter == 0:
return (1,0)
elif y_voter == 0 and n_voter > 0:
return (0,1)
return (0,0)
```
Function to measure user's vote:
```
def correlation_volume_yes_no_vote(n_kind, vote_func=None):
'''
n_kind = 'n' # All entities tweets.
n_kind = 'n_yesno' # Only yes/no twetts
'''
if not vote_func:
vote_func = yes_no_vote_1
yes_voters = []
no_voters = []
for username in referendum_user_stats:
n = referendum_user_stats[username]['n']
n_yes = referendum_user_stats[username]['e'].get('voteyes', {'n': 0})["n"]
n_no = referendum_user_stats[username]['e'].get('voteno', {'n': 0})["n"]
yes_voter, no_voter = vote_func(username)
if yes_voter > no_voter:
if n_kind == 'n':
yes_voters.append(n)
elif n_kind == 'n_yesno':
yes_voters.append(n_yes)
elif no_voter > yes_voter:
if n_kind == 'n':
no_voters.append(n)
elif n_kind == 'n_yesno':
no_voters.append(n_no)
return yes_voters, no_voters
def correlation_volume_yes_no_vote_statistical():
for n_kind in ['n', 'n_yesno']:
yes_voters, no_voters = correlation_volume_yes_no_vote(n_kind)
print 'KIND:', n_kind
print 'YES VOTERS: {} NO VOTERS: {}'.format(len(yes_voters), len(no_voters))
print 'YES AVERAGE VALUE: {} NO AVERAGE VALUE: {}'.format(np.average(yes_voters), np.average(no_voters))
print 'Mann–Whitney U test: {}'.format(mannwhitneyu(yes_voters, no_voters))
correlation_volume_yes_no_vote_statistical()
```
## Who are the top "YES" twitteres?
```
def top_yes_voters():
yes_voters, no_voters = correlation_volume_yes_no_vote('n')
yes_voters_sorted = sorted(referendum_user_stats, key=lambda x : 0 )
a = sorted(referendum_user_stats, key=lambda x : referendum_user_stats[x]['e']['voteyes']['n'] if yes_no_vote_1(x)[0]==1 else 0, reverse=True)
a[0:20]
```
## How YES/NO users changed sentiment
Reviewer's comment:
Another suggestion for future work would be to analyze how polarized/divided
the public was during the time before the referendum and how that changed right after
the vote and Tsipras' announcement that was perceived by many as changing the "NO" vote
to a "YES
```
def per_user_analysis(kind):
filename = '{}_user_stats.json'.format(kind)
if os.path.exists(filename):
print 'Filename: {} exists. Delete it to regenerate it'.format(filename)
with open(filename) as f:
return json.load(f)
user_stats = {}
c=0
for item in merge_tweets_sentiment(kind):
c += 1
username = item['un']
#Initialize stats
if not username in user_stats:
user_stats[username] = {
'n': 0, # Number of tweets
's_s': 0.0, # sarcasm score
's_p': 0.0, # Sarcasm percentage
'sp': 0.0, # Sum of positive sentiment
'sn': 0.0, # Sum of negative sentiment
's0': 0.0, # Sum of neutral sentiment
'sp_c': 0.0, # Sum of sarcasm corrected positive sentiment
'sn_c': 0.0, # Sum of sarcasm corrected negative sentiment
'e': {}, # Entities stats,
'y_voter': 0, # Number of yes tweets
'n_voter': 0, # Number of no tweets
}
user_stats[username]['n'] += 1
user_stats[username]['s_s'] += item['sarcasm']['score']
user_stats[username]['s_p'] += item['sarcasm']['percentage']
user_stats[username]['sp'] += item['sp']
user_stats[username]['sp_c'] += correct_sarcasm(item['sp'], item['sarcasm']['percentage'])
user_stats[username]['sn_c'] += correct_sarcasm(item['sn'], item['sarcasm']['percentage'])
user_stats[username]['sn'] += item['sn']
user_stats[username]['s0'] += item['s0']
#store per entity per username stats
for entity in item['hashtag_entites'] + item['text_entities']:
#Intialize
if not entity in user_stats[username]['e']:
user_stats[username]['e'][entity] = {
'n': 0,
's_s': 0.0,
's_p': 0.0,
'sp': 0.0,
'sn': 0.0,
'sp_c': 0.0,
'sn_c': 0.0,
's0': 0.0,
}
user_stats[username]['e'][entity]['n'] += 1
user_stats[username]['e'][entity]['s_s'] += item['sarcasm']['score']
user_stats[username]['e'][entity]['s_p'] += item['sarcasm']['percentage']
user_stats[username]['e'][entity]['sp'] += item['sp']
user_stats[username]['e'][entity]['sp_c'] += correct_sarcasm(item['sp'], item['sarcasm']['percentage'])
user_stats[username]['e'][entity]['sn'] += item['sn']
user_stats[username]['e'][entity]['sn_c'] += correct_sarcasm(item['sn'], item['sarcasm']['percentage'])
user_stats[username]['e'][entity]['s0'] += item['s0']
# Store referendum voting info
item_hashtags = item['ht']
yes_voter = len([x for x in item_hashtags if x in yes_no_entities['yes']]) > 0
no_voter = len([x for x in item_hashtags if x in yes_no_entities['no']]) > 0
user_stats[username]['y_voter'] += 1 if yes_voter else 0
user_stats[username]['n_voter'] += 1 if no_voter else 0
#Save file
with open(filename, 'w') as f:
json.dump(user_stats, f, indent=4)
print 'Created filename:', filename
return user_stats
elections_user_stats = per_user_analysis('elections')
referendum_user_stats = per_user_analysis('referendum')
def compare_emotions_before_after_referendum_for_NO_voters(yes_no, voting_f):
'''
voting_f is one of [yes_no_vote_1, yes_no_vote_2]
'''
compare_entities = all_entities
results = {}
for election_voter in elections_user_stats:
if not election_voter in referendum_user_stats:
continue # This user did was not present in referendum
v = voting_f(election_voter)
if yes_no == 'no':
if v[0] >= v[1]:
continue # This is either a YES voter or didn't vote
elif yes_no == 'yes':
if v[0] <= v[1]:
continue #This is either a NO voter or didn't vote
for entity in compare_entities:
if entity in referendum_user_stats[election_voter]['e'] and entity in elections_user_stats[election_voter]['e']:
pass
else:
continue #This entity is not present in both election and referendum stats
if entity not in results:
results[entity] = {
'r': {'s_p': [], 's_s': [], 'sp': [], 'sn': [], 'sp_c':[], 'sn_c':[], 's0': [], 'n': 0},
'e': {'s_p': [], 's_s': [], 'sp': [], 'sn': [], 'sp_c':[], 'sn_c':[], 's0': [], 'n': 0},
}
n_r = referendum_user_stats[election_voter]['e'][entity]['n']
results[entity]['r']['s_p'].append(referendum_user_stats[election_voter]['e'][entity]['s_p'] / n_r)
results[entity]['r']['s_s'].append(referendum_user_stats[election_voter]['e'][entity]['s_s'] / n_r)
results[entity]['r']['sp'].append(referendum_user_stats[election_voter]['e'][entity]['sp'] / n_r)
results[entity]['r']['sp_c'].append(referendum_user_stats[election_voter]['e'][entity]['sp_c'] / n_r)
results[entity]['r']['sn'].append(referendum_user_stats[election_voter]['e'][entity]['sn'] / n_r)
results[entity]['r']['sn_c'].append(referendum_user_stats[election_voter]['e'][entity]['sn_c'] / n_r)
results[entity]['r']['s0'].append(referendum_user_stats[election_voter]['e'][entity]['s0'] / n_r)
results[entity]['r']['n'] += n_r
n_e = elections_user_stats[election_voter]['e'][entity]['n']
results[entity]['e']['s_p'].append(elections_user_stats[election_voter]['e'][entity]['s_p'] / n_e)
results[entity]['e']['s_s'].append(elections_user_stats[election_voter]['e'][entity]['s_s'] / n_e)
results[entity]['e']['sp'].append(elections_user_stats[election_voter]['e'][entity]['sp'] / n_e)
results[entity]['e']['sp_c'].append(elections_user_stats[election_voter]['e'][entity]['sp_c'] / n_e)
results[entity]['e']['sn'].append(elections_user_stats[election_voter]['e'][entity]['sn'] / n_e)
results[entity]['e']['sn_c'].append(elections_user_stats[election_voter]['e'][entity]['sn_c'] / n_e)
results[entity]['e']['s0'].append(elections_user_stats[election_voter]['e'][entity]['s0'] / n_e)
results[entity]['e']['n'] += n_e
#Normalize
#for entity in compare_entities:
# for k in ['r', 'e']:
# for stat in ['s_p', 's_s', 'sp', 'sn', 's0']:
# if entity in results:
# results[entity][k][stat] = results[entity][k][stat] / results[entity][k]['n']
return results
def compare_emotions_before_after_referendum_for_NO_voters_stat(yes_no, voting_f, stats):
results = compare_emotions_before_after_referendum_for_NO_voters(yes_no, voting_f)
ret = {}
for entity in all_entities:
if not entity in results:
continue
#if results[entity]['r']['n'] < 20 or results[entity]['e']['n'] < 20:
# continue # Not enough data for Mann–Whitney U test
#for stat in ['s_p', 's_s', 'sp', 'sn', 's0']:
#for stat in ['sp', 'sn']:
#for stat in ['sp_c', 'sn_c']:
for stat in stats:
r_stat = results[entity]['r'][stat]
e_stat = results[entity]['e'][stat]
if len(r_stat)<20 or len(e_stat)<20:
continue # Not enough data for Mann–Whitney U test
u_stat, p_value = mannwhitneyu(r_stat, e_stat)
if p_value < 0.001:
r_average = np.average(r_stat)
e_average = np.average(e_stat)
print 'Entity: {} STAT: {} p_value: {} r_av: {} e_av:{}'.format(entity, stat, p_value, r_average, e_average)
if not entity in ret:
ret[entity] = {}
if not stat in ret[entity]:
ret[entity][stat] = {}
ret[entity][stat] = {
'p': p_value,
'r_av': r_average,
'e_av': e_average,
}
return ret
beautified_labels = {
'syriza': 'SYRIZA',
'greece': 'Greece',
'greekgovernment': 'Government',
'alexistsipras': 'Tsipras',
'newdemocracy': 'ND',
'hellenicparliament': 'Parliament',
'greekdebt': 'Debt',
'massmedia': 'Media',
'voteno': 'NO',
'europe': 'Europe',
}
def compare_emotions_before_after_referendum_for_NO_voters_stat_plot(yes_no, voting_f, stats, ax=None):
results = compare_emotions_before_after_referendum_for_NO_voters_stat(yes_no, voting_f, stats)
plot_ylabel = False
if not ax:
plot_ylabel = True
fig, ax = plt.subplots()
#ax = plt.subplot(111)
labels = []
for e_index, entity in enumerate(results.keys()):
#for stat in ['sp', 'sn']:
#for stat in ['sp_c', 'sn_c']:
for stat_i, stat in enumerate(stats):
if stat in results[entity]:
p_from = results[entity][stat]['r_av']
p_to = results[entity][stat]['e_av']
nice_entity = beautified_labels.get(entity, entity)
if not nice_entity in labels:
labels.append(nice_entity)
if stat_i == 0: # positive
color = 'blue'
elif stat_i == 1: # negative
color = 'red'
else:
assert False
#print p_from, p_to
#print p_from, p_to-p_from
#ax.plot([e_index, e_index], [p_from, p_to], c=color)
#ax.arrow( 5, 8, 0.0, -2, fc="k", ec="k", head_width=0.05, head_length=0.5 )
ax.arrow(e_index, p_from, 0.0, p_to-p_from, fc=color, ec=color, head_width=0.4, head_length=0.2, width=0.2)
#ax.annotate("", xy=(1.2, 0.2), xycoords=ax, xytext=(0.8, 0.8), textcoords=ax, arrowprops=dict(arrowstyle="->",connectionstyle="arc3"),)
leg_1, = ax.plot([], [], '-', c='blue', )
leg_2, = ax.plot([], [], '-', c='red', )
labels_x = range(-1, len(labels)+1)
labels = [''] + labels + ['']
plt.xticks(labels_x, labels, rotation='vertical')
#plt.xticks(labels_x, labels, rotation=45)
ax.set_xlim((-1, len(labels)-2))
ax.set_ylim((-3, 2))
ax.set_xlabel('{} "Voters"'.format(yes_no.upper()))
if plot_ylabel:
if yes_no == 'no':
plt.legend([leg_1, leg_2], ["Positive Sentiment", "Negative Sentiment"], loc=1, bbox_to_anchor=(0.5, 1, 0.2, 0))
else:
plt.legend([leg_1, leg_2], ["Positive Sentiment", "Negative Sentiment"], loc=1)
if plot_ylabel:
ax.set_ylabel(u'Variation of Referentum \u2192 Elections Sentiment')
#plt.savefig("plot20_{}.eps".format(yes_no))
```
## Figure 10. Change of sentiment between Referendum and Elections
```
def compare_emotions_before_after_referendum_for_NO_voters_stat_plot_combinations():
for yes_no in ['yes', 'no']:
for vote_f in [yes_no_vote_1, yes_no_vote_2]:
for stats in [['sp', 'sn'], ['sp_c', 'sn_c']]:
print '{} {} {}'.format(yes_no, vote_f.__name__, stats)
compare_emotions_before_after_referendum_for_NO_voters_stat_plot(yes_no, vote_f, stats)
#compare_emotions_before_after_referendum_for_NO_voters_stat_plot_combinations()
def compare_emotions_before_after_referendum_for_NO_voters_stat_plot_two():
plt.figure(1)
ax1 = plt.subplot(1,2,1)
compare_emotions_before_after_referendum_for_NO_voters_stat_plot('yes', yes_no_vote_2, ['sp', 'sn'], ax1)
ax1.set_ylabel(u'Referentum \u2192 Elections')
leg_1, = ax1.plot([], [], '-', c='blue', )
leg_2, = ax1.plot([], [], '-', c='red', )
#ax2 = plt.subplot(1,2,2, sharey=ax1)
ax2 = plt.subplot(1,2,2)
compare_emotions_before_after_referendum_for_NO_voters_stat_plot('no', yes_no_vote_2, ['sp', 'sn'], ax2)
ax2.get_yaxis().set_ticklabels([])
plt.legend([leg_1, leg_2], ["Positive Sentiment", "Negative Sentiment"], loc=1, bbox_to_anchor=(-0.5, 0.6, 0.2, 0))
#plt.suptitle(u'Change of sentiment between Referendum and Elections') # PLOS
plt.tight_layout()
plt.savefig('figure10.eps')
plt.savefig('plot21.eps')
compare_emotions_before_after_referendum_for_NO_voters_stat_plot_two()
```
## Figure 3. Variation of YES/NO percentage overtime
Reviewer's comment:
Plots and axis labelling to be improved (data are from 25 or 27 of June? In Fig 3 why 30 Jun is missing?).
```
def variation_YES_NO_percentage():
#Load older version of hastags
#ht = yes_no_entities()
ht = yes_no_entities
users_ht = {}
start_time = None
duration = 60 * 60 # One hour
to_plot = []
fn_1 = 'plot_variation_YES_NO_percentage.json'
if os.path.exists(fn_1):
print ("Loading file:", fn_1)
with open(fn_1) as f:
to_plot = json.load(f)
else:
for item in merge_tweets_sentiment('referendum'):
this_time = item['date']
#Remove tweets BEFORE the announcement of the referendum
#Interesting very few tweets exist that predicted(?) that a referendum will be announced.
if this_time < 1435288026.0:
continue
#item_entities = set(item['hashtag_entites']) | set(item['text_entities'])
item_hashtags = item['ht']
username = item['un']
if not username in users_ht:
users_ht[username] = {'yes': 0, 'no': 0, 'both': 0, 'none': 0}
#Every <duration> we collect user stats
if not start_time:
start_time = this_time
elif this_time - start_time > duration:
start_time = this_time
yes_users = len(filter(lambda x : x['yes'] > 0 and x['no'] == 0, users_ht.values()))
no_users = len(filter(lambda x : x['yes'] == 0 and x['no'] > 0, users_ht.values()))
yes_users_multi = len(filter(lambda x : x['yes'] > x['no'], users_ht.values()))
no_users_multi = len(filter(lambda x : x['no'] > x['yes'], users_ht.values()))
to_plot.append([start_time, {'yes_users': yes_users, 'no_users': no_users, 'yes_users_multi':yes_users_multi, 'no_users_multi':no_users_multi}])
#yes_voter = 'voteyes' in item_entities
yes_voter = len([x for x in item_hashtags if x in ht['yes']]) > 0
no_voter = len([x for x in item_hashtags if x in ht['no']]) > 0
if not yes_voter and not no_voter:
users_ht[username]['none'] += 1
elif yes_voter and no_voter:
users_ht[username]['both'] += 1
elif yes_voter and not no_voter:
users_ht[username]['yes'] += 1
elif not yes_voter and no_voter:
users_ht[username]['no'] += 1
with open(fn_1, 'w') as f:
print 'Saving..'
json.dump(to_plot, f, indent=4)
print ('Created file:', fn_1)
# PLOTTING
fig, ax = plt.subplots()
#start_from = 1.43539e9
#start_from = 1.43510e9
start_from = 0
yes_key = 'yes_users'
no_key = 'no_users'
perc = [(100.0 * float(x[1][yes_key])/float(x[1][yes_key] + x[1][no_key]) if x[1][yes_key] + x[1][no_key] > 0 else 0.0 ) for x in to_plot if x[0] > start_from]
X = [x[0] for x in to_plot if x[0] > start_from]
leg_yes, = ax.plot(X, [x[1][yes_key] for x in to_plot if x[0] > start_from], '-b')
leg_no, = ax.plot(X, [x[1][no_key] for x in to_plot if x[0] > start_from], '-r')
ax2 = ax.twinx()
perc_to_plot = [(x,y) for x, y in zip(X, perc) if x > 1435395600]
#leg_perc, = ax2.plot(X, perc, '--k')
leg_perc, = ax2.plot([x[0] for x in perc_to_plot], [x[1] for x in perc_to_plot], '--k')
ax2.plot([X[0]-100000, X[-1]+100000], [38.7, 38.7], '-g', linewidth=6, alpha=0.6, color='0.2')
ax2.text(1.43557e9, 39.6, 'Result: YES=38.6%', fontsize=12)
#ax.set_title('Variation of YES percentage') # IN PLOS
ax.set_xlabel('Date (2015)')
ax.set_ylabel('Number of Users')
ax2.set_ylabel('Percentage of "YES"')
ax2.set_ylim((0.0, 70.0))
#ax.set_xlim((X[0]-10000, X[-1] + 10000))
ax.set_xlim((X[0]-20000, X[-1] + 10000))
ax2.annotate('Capital Controls', xy=(1.43558e9, 25.0), xytext=(1.43535e9, 15.0),
arrowprops=dict(facecolor='0.8',
#shrink=0.05,
),
bbox=dict(boxstyle="round", fc="0.8"),
)
#fix_time_ticks(ax, string_time='%e %b')
epochs = []
#labels = []
for day,month in ((25, 6), (26,6), (27,6), (28,6), (29,6), (30,6), (1,7), (2,7), (3,7), (4,7), (5,7)):
e = float(datetime.datetime(2015, month, day, 12, 0, 0).strftime('%s'))
epochs.append(e)
# .strftime('%s')
ax.set_xticks(epochs)
fix_time_ticks(ax, string_time='%e %b', fontsize=7)
plt.legend([leg_yes, leg_no, leg_perc], ['YES users', 'NO users', 'YES percentage'], loc=2)
plt.savefig('figure3')
plt.savefig('figure3.eps')
plt.savefig('plot_variation_YES_NO_percentage.eps')
ax
variation_YES_NO_percentage()
```
## Sarcasm percentages
```
def plot_22():
# https://stackoverflow.com/questions/24575869/read-file-and-plot-cdf-in-python
referendum_sarcasm = [item['sarcasm']['percentage'] for item in merge_tweets_sentiment('referendum')]
sorted_ref_data = np.sort(referendum_sarcasm)
elections_sarcasm = [item['sarcasm']['percentage'] for item in merge_tweets_sentiment('elections')]
sorted_ele_data = np.sort(elections_sarcasm)
print "Percentage of ref tweets with sarcasm > 20:"
print len(sorted_ref_data[sorted_ref_data>20])/float(len(sorted_ref_data)-1)
print "Percentage of ele tweets with sarcasm > 20:"
print len(sorted_ele_data[sorted_ele_data>20])/float(len(sorted_ele_data)-1)
fig, ax = plt.subplots()
yvals_ref=100.0 * np.arange(len(sorted_ref_data))/float(len(sorted_ref_data)-1)
yvals_ele=100.0 * np.arange(len(sorted_ele_data))/float(len(sorted_ele_data)-1)
leg_ref, = ax.plot(sorted_ref_data,yvals_ref, c='b')
leg_ele, = ax.plot(sorted_ele_data,yvals_ele, c='r')
plt.legend([leg_ref, leg_ele], ['Referendum', 'Elections'], loc=2)
ax.set_xlabel('Sarcasm Percentage (<0 not sarcastic, >0 sarcastic)')
ax.set_ylabel('Cumulative Percent')
ax.set_title('Cumulative Distribution Function for Sarcasm')
plt.savefig('plot_22.eps')
plt.savefig('plot_22.png')
ax
plot_22()
```
**Caption of the figure above**: Cumulative Distribution Function of Sarcasm for Referendum (blue) and Elections (red) tweets. Each tweet has a sarcasm percentage value assigned by the sarcasm detection algorithm. Tweets with negative percentage are not sarcastic whereas tweets with positive percentage are sarcastic. T he percentage, indicates the confidence of the algorithm. Overall approximately 50% of the tweets had a positive sarcasm value.
## Show examples of tweets from various categories
```
yes_set = set(map(unicode.lower, yes_no_entities['yes']))
no_set = set(map(unicode.lower, yes_no_entities['no']))
def item_entities(item):
return set(item['hashtag_entites'] + item['text_entities'])
def is_yes_item(item):
return not (not (item_entities(item) & yes_set)) # the not (not..) part is to convert fast from set to boolean
def is_no_item(item):
return not (not (item_entities(item) & no_set))
def example_of_tweets(n, kind, filter_):
shown = []
item_c = 0
for item in merge_tweets_sentiment(kind):
item_c += 1
text = item['original_text']
if filter_(item):
if not text in shown:
shown.append(text)
print text
if len(shown) > n:
break
```
10 "YES" tweets from referendum:
```
example_of_tweets(10, 'referendum', lambda item : is_yes_item(item) )
```
10 "YES" tweets from referendum:
```
example_of_tweets(10, 'referendum', lambda item : is_yes_item(item) )
```
10 election tweets with a sarcasm level between 0% and 20%
```
example_of_tweets(10, 'elections', lambda item : 0<item['sarcasm']['percentage']<20 ) # a<b<c is valid in python!
```
10 election tweets with a sarcasm level above 40%
```
example_of_tweets(10, 'elections', lambda item : item['sarcasm']['percentage']>40 )
```
## Figure 1. Frequency of tweets per hour
```
def figure_1():
min_time = 1435342648.0
final_max_time = 1436101199.0 # 16.59 Sunday 5 July
if True:
#created_at_gre = [get_created_at_unix(x) for x in status_generator('ht_greferendum_FINAL_greek.json')]
#created_at_dim = [get_created_at_unix(x) for x in status_generator('ht_dimopsifisma_FINAL_greek.json')]
created_at_gre = []
created_at_dim = []
for item in merge_tweets_sentiment('referendum'):
hashtags = item['ht']
date_epochs = item['date']
if u'greferendum' in hashtags:
created_at_gre.append(date_epochs)
if u'dimopsifisma' in hashtags:
created_at_dim.append(date_epochs)
filename = 'plot_9.json'
print 'Saving:', filename
with open(filename, 'w') as f:
json.dump([created_at_gre, created_at_dim], f)
if True:
filename = 'plot_9.json'
print 'Opening:', filename
with open(filename) as f:
data = json.load(f)
created_at_gre, created_at_dim = data
if True:
step = 60.0 # 1 minute
bins = int((final_max_time - min_time) / step)
print 'Bins:', bins
fig, ax = plt.subplots()
#sns.distplot(created_at_gre, norm_hist=False, kde=False, bins=bins, label='YES users', color='blue')
n, bins, patches = ax.hist(created_at_gre, bins = bins, histtype='step', color='green',) # , alpha=0.5)
n, bins, patches = ax.hist(created_at_dim, bins = bins, histtype='step', color='magenta',) #, alpha=0.5)
ax.set_xlim((min_time, final_max_time))
fix_time_ticks(ax, string_time='%d/%m')
leg_gre, = ax.plot([], [], '-', color='green', linewidth=3 )
leg_dim, = ax.plot([], [], '-', color='magenta', linewidth=3 )
plt.legend([leg_gre, leg_dim], ['#greferendum', '#dimopsifisma'])
#ax.set_ylim((0, 140)) # 300
ax.set_ylim((0, 250)) # 300
ax.set_xlabel('Date (2015)')
ax.set_ylabel('Tweets per hour')
#ax.set_title('Frequency of referendum tweets') # ADD TITLE IN PLOS ARTICLE
save_figure('figure1')
#save_figure('plot_9')
ax
figure_1()
```
## Figure 2. Frequency of YES/NO tweets in Referendum
```
def figure_2(figure_to_plot):
'''
PLOT 1
YES / NO Tendencies
Hastags:
#voteYes
#voteNo
#YesToEurope
#NoToEurope
Peak 1:
http://www.theguardian.com/business/live/2015/jun/29/greek-crisis-stock-markets-slide-capital-controls-banks-closed-live
https://twitter.com/ThePressProject/status/615574647443193856/video/1 29 June 2015
Peak 2:
#ΟΧΙ (NO) is not just a slogan.
NO is a decisive step toward a better deal. #Greece #dimopsifisma #Greferendum 735
#ΟΧΙ / NO does not mean breaking w/Europe, but returning to the Europe of values.
#ΟΧΙ / NO means: strong pressure.
#Greece #dimopsifisma 691
Είναι η ευθύνη προς τους γονείς μας, προς τα παιδιά μας,προς τους εαυτούς μας.
Είναι το χρέος μας απέναντι στην ιστορία #dimopsifisma #OXI 203
Peak 3:
https://twitter.com/vanpotamianos/status/617054822924218369/photo/1 3 July 2015
https://www.rt.com/news/271597-greece-athens-referendum-rallies/
YES and NO
'''
# 'entities_from_hashtags' : '/Users/despoina/Dropbox/Despoina/hashtags_for_entities/hashtags_for_entities_v20160405.txt',
# 'entities_from_text' : '/Users/despoina/Dropbox/Despoina/entities_from_text/entities_from_text_v20160415.txt',
ht = {}
with codecs.open(data['entities_from_hashtags_old_1']['filename'], encoding='utf8') as f:
print 'Using:', f.name
for l in f:
ls = l.replace(os.linesep, '').split()
if ls[0][0] == '#':
ht[ls[0][1:]] = ls[1:] # Remove #
else:
ht[ls[0]] = ls[1:] # DO NOT Remove #
ht2 = {}
ht2['yes'] = list(set(ht['voteYes'] + ht['YesToEurope']))
ht2['no'] = list(set(ht['voteNo'] + ht['NoToEurope']))
# ht2['yes'] = list(set(ht['voteYes'] + ht['YesEurope']))
# ht2['no'] = list(set(ht['voteNo'] + ht['NoEurope']))
# ht2['yes'] = list(set(ht['voteyes'] + ht['yestoeurope']))
# ht2['no'] = list(set(ht['voteno'] + ht['notoEurope']))
del ht
ht = ht2
c = 0
start_time = None
duration = 60 * 60 # One hour
use_only_originals = False
originals = {}
to_plot_1 = []
to_plot_2 = []
picks = [
[(1.43558e9, 1.43560e09), {}],
[(1.43574e9, 1.43576e09), {}],
[(1.43594e9, 1.43596e09), {}],
]
'''
2 RT @tsipras_eu: #ΟΧΙ (NO) is not just a slogan. NO is a decisive step toward a better deal. #Greece #dimopsifisma #Greferendum
2 RT @tsipras_eu: #ΟΧΙ / NO does not mean breaking w/Europe, but returning to the Europe of values.
#ΟΧΙ / NO means: strong pressure.
#Greece…
'''
users_ht = {}
no_greek = 0
if True: # Builds to_plot_1 , to_plot_2 data
for j in get_json(data['referendum_tweets']['filename']):
c += 1
original_text = get_original_text(j)
if use_only_originals:
if original_text in originals:
continue
originals[original_text] = None
if c % 10000 == 0:
print c
hashtags = tweet_get_hashtags(j)
#print hashtags
this_time = tweet_get_date_epochs(j)
text = tweet_get_text(j)
if not is_greek_text(text):
not_greek += 1
continue
username = tweet_get_user(j)
if not username in users_ht:
users_ht[username] = {'yes': 0, 'no': 0, 'both': 0, 'none': 0}
if this_time < 1435288026.0:
continue
#Check picks
for p_index, p in enumerate(picks):
if p[0][0] <= this_time <= p[0][1]:
original_text = get_original_text(j)
#print p_index+1, original_text
if not original_text in p[1]:
p[1][original_text] = 0
p[1][original_text] += 1
if not start_time:
this_bucket = {'yes':0, 'no': 0, 'both': 0, 'none': 0}
start_time = this_time
elif this_time - start_time > duration:
to_plot_1.append([start_time, this_bucket])
this_bucket = {'yes':0, 'no': 0, 'both': 0, 'none': 0}
start_time = this_time
yes_users = len([None for v in users_ht.values() if v['yes'] > 0 and v['no'] == 0])
no_users = len([None for v in users_ht.values() if v['yes'] == 0 and v['no'] > 0])
yes_users_multi = len([None for v in users_ht.values() if v['yes'] > v['no']])
no_users_multi = len([None for v in users_ht.values() if v['yes'] < v['no']])
to_plot_2.append([start_time, {'yes_users': yes_users, 'no_users': no_users, 'yes_users_multi':yes_users_multi, 'no_users_multi':no_users_multi}])
yes_hts = len([x for x in hashtags if x in ht['yes']])
no_hts = len([x for x in hashtags if x in ht['no']])
if yes_hts == 0 and no_hts == 0:
this_bucket['none'] += 1
users_ht[username]['none'] += 1
elif yes_hts > 0 and no_hts > 0:
this_bucket['both'] += 1
users_ht[username]['both'] += 1
elif yes_hts > 0 and no_hts == 0:
this_bucket['yes'] += 1
users_ht[username]['yes'] += 1
elif yes_hts == 0 and no_hts > 0:
this_bucket['no'] += 1
users_ht[username]['no'] += 1
else:
raise Exception('This should not happen')
#print yes_hts, no_hts
#print this_time
#if c > 50000:
# break
if False: # Print picks text
for p_index, p in enumerate(picks):
s = sorted(list(p[1].iteritems()), key=lambda x:x[1], reverse=True)
print p_index
for x in range(10):
print s[x][0], s[x][1]
if True:
#Save plot data
suffix = '_originals' if use_only_originals else ''
with open('plot_1%s.json' % (suffix), 'w') as f:
f.write(json.dumps(to_plot_1) + '\n')
print 'Saved: plot_1%s.json' % (suffix)
with open('plot_2%s.json' % (suffix), 'w') as f:
f.write(json.dumps(to_plot_2) + '\n')
print 'Saved: plot_2%s.json' % (suffix)
if False:
with open('plot_1.json') as f:
to_plot_1 = json.load(f)
if False:
with open('plot_2.json') as f:
to_plot_2 = json.load(f)
print 'NO GREEK TWEETS:', no_greek
fig, ax = plt.subplots()
#YES
if figure_to_plot == 2: # Make plot_1
leg_yes, = ax.plot([x[0] for x in to_plot_1], [x[1]['yes'] for x in to_plot_1], '-b')
leg_no, = ax.plot([x[0] for x in to_plot_1], [x[1]['no'] for x in to_plot_1], '-r')
#leg_none, = ax.plot([x[0] for x in to_plot_1], [x[1]['none'] for x in to_plot_1], '-k')
plt.legend([leg_yes, leg_no], ['YES Tweets', 'NO Tweets'])
ax.set_xlabel('Date (2015)')
ax.set_ylabel('Number of Tweets per hour')
# ax.set_title('Frequency of YES/NO tweets in Referendum') # PLOS SET TITLE
ax.annotate('29 June 2015\nfirst NO demonstration', xy=(1.43558e9, 900), xytext=(1.43530e9, 1300),
arrowprops=dict(facecolor='black', shrink=0.05, ),
)
ax.annotate('retweets of @atsipras\nNO promoting posts', xy=(1.43575e9, 1950), xytext=(1.43545e9, 2100),
arrowprops=dict(facecolor='black', shrink=0.05, ),
)
ax.annotate('3 July 2015\nBig YES and NO demonstrations', xy=(1.43593e9, 2550), xytext=(1.43550e9, 2600),
arrowprops=dict(facecolor='black', shrink=0.05, ),
)
fix_time_ticks(ax, string_time='%e %b')
save_figure('figure2')
#save_figure('plot_1')
ax
figure_2(2)
```
## Figure 4. Frequency of elections tweets
```
def figure_4():
'''
PLOT 5
Sunday, 20 September
'''
# 'entities_from_hashtags' : '/Users/despoina/Dropbox/Despoina/hashtags_for_entities/hashtags_for_entities_v20160405.txt',
# 'entities_from_text' : '/Users/despoina/Dropbox/Despoina/entities_from_text/entities_from_text_v20160415.txt',
ht = {}
with codecs.open(data['entities_from_hashtags']['filename'], encoding='utf8') as f:
for l in f:
ls = l.replace(os.linesep, '').split()
ht[ls[0][1:]] = ls[1:]
ht2 = {}
#ht2['yes'] = list(set(ht['voteYes'] + ht['YesToEurope']))
#ht2['no'] = list(set(ht['voteNo'] + ht['NoToEurope']))
ht2['SYRIZA'] = list(set(ht['Syriza']))
ht2['ND'] = list(set(ht['NewDemocracy']))
ht2['TSIPRAS'] = list(set(ht['AlexisTsipras']))
ht2['MEIMARAKIS'] = list(set(ht['VangelisMeimarakis']))
ht2['ANEL'] = list(set(ht['ANEL']))
ht2['PopularUnity'] = list(set(ht['PopularUnity']))
ht2['TheRiver'] = list(set(ht['TheRiver']))
ht2['PASOK'] = list(set(ht['PanhellenicSocialistMovement']))
ht2['GoldenDawn'] = list(set(ht['GoldenDawn']))
ht2['CommunistPartyOfGreece'] = list(set(ht['CommunistPartyOfGreece']))
del ht
ht = ht2
rivals = [('SYRIZA', 'ND'), ('TSIPRAS', 'MEIMARAKIS')]
c = 0
start_time = None
#duration = 60 * 60 # One hour
duration = 60 * 60 * 24 # 12 hours
to_plot_5 = []
to_plot_6 = []
if False:
picks = [
[(1.43558e9, 1.43560e09), {}],
[(1.43574e9, 1.43576e09), {}],
[(1.43594e9, 1.43596e09), {}],
]
'''
2 RT @tsipras_eu: #ΟΧΙ (NO) is not just a slogan. NO is a decisive step toward a better deal. #Greece #dimopsifisma #Greferendum
2 RT @tsipras_eu: #ΟΧΙ / NO does not mean breaking w/Europe, but returning to the Europe of values.
#ΟΧΙ / NO means: strong pressure.
#Greece…
'''
users_ht = {}
not_greek = 0
def empty_dict():
ret = {}
for rival_index, rival in enumerate(rivals):
for rival_party in rival:
ret[rival_party] = 0
ret['both_%i' % rival_index] = 0
ret['none_%i' % rival_index] = 0
return ret
if True: # Builds to_plot_1 , to_plot_2 data
for j in get_json(data['elections_tweets']['filename']):
c += 1
if c % 10000 == 0:
print c, tweet_get_date_text(j)
hashtags = tweet_get_hashtags(j)
#print hashtags
this_time = tweet_get_date_epochs(j)
text = tweet_get_text(j)
if not is_greek_text(text):
not_greek += 1
continue
username = tweet_get_user(j)
if not username in users_ht:
users_ht[username] = empty_dict()
# users_ht[username] = {'nd': 0, 'syriza': 0, 'both': 0, 'none': 0, 'tsipras':0, 'meimarakis': 0: 'none_tm': 0, 'both_tm': 0}
if this_time < 1440000000.0:
continue
if False: #Check picks
for p_index, p in enumerate(picks):
if p[0][0] <= this_time <= p[0][1]:
original_text = get_original_text(j)
#print p_index+1, original_text
if not original_text in p[1]:
p[1][original_text] = 0
p[1][original_text] += 1
if not start_time:
this_bucket = empty_dict()
#this_bucket = {'nd':0, 'syriza': 0, 'both': 0, 'none': 0, 'tsipras':0, 'meimarakis': 0: 'none_tm': 0, 'both_tm': 0}
start_time = this_time
elif this_time - start_time > duration:
to_plot_5.append([start_time, this_bucket])
this_bucket = empty_dict()
#this_bucket = {'nd':0, 'syriza': 0, 'both': 0, 'none': 0, 'tsipras':0, 'meimarakis': 0: 'none_tm': 0, 'both_tm': 0}
start_time = this_time
#Count users
dict_to_add = {}
for rival_index, rival in enumerate(rivals):
for rival_party in rival:
dict_to_add[rival_party + '_users'] = len([None for v in users_ht.values() if v[rival_party] > 0 and sum([v[x] for x in rival if x != rival_party]) == 0])
dict_to_add[rival_party + '_multi'] = len([None for v in users_ht.values() if v[rival_party] > sum([v[x] for x in rival if x != rival_party])])
to_plot_6.append([start_time, dict_to_add])
# yes_users = len([None for v in users_ht.values() if v['nd'] > 0 and v['syriza'] == 0])
# no_users = len([None for v in users_ht.values() if v['nd'] == 0 and v['syriza'] > 0])
# meimarakis_users = len([None for v in users_ht.values() if v['meimarakis'] > 0 and v['tsipras'] == 0])
# tsipras_users = len([None for v in users_ht.values() if v['meimarakis'] == 0 and v['tsipras'] > 0])
# yes_users_multi = len([None for v in users_ht.values() if v['nd'] > v['syriza']])
# no_users_multi = len([None for v in users_ht.values() if v['nd'] < v['syriza']])
# meimarakis_users_multi = len([None for v in users_ht.values() if v['meimarakis'] > v['tsipras']])
# tsipras_users_multi = len([None for v in users_ht.values() if v['meimarakis'] < v['tsipras']])
# to_plot_6.append([start_time,
# {'nd_users': yes_users,
# 'syriza_users': no_users,
# 'nd_users_multi':yes_users_multi,
# 'syriza_users_multi':no_users_multi,
# 'meimarakis_users':meimarakis_users,
# 'tsipras_users':tsipras_users,
# 'meimarakis_users_multi': meimarakis_users_multi,
# 'tsipras_users_multi': tsipras_users_multi,
# }])
#Count tweets
for rival_index, rival in enumerate(rivals):
rival_dict = {}
selected_party = None
for rival_party in rival:
rival_dict[rival_party] = len([None for x in hashtags if x in ht[rival_party]])
if rival_dict[rival_party] > 0:
selected_party = rival_party
if all(rival_dict.values()):
this_bucket['both_%i' % rival_index] += 1
users_ht[username]['both_%i' % rival_index] += 1
elif not any(rival_dict.values()):
this_bucket['none_%i' % rival_index] += 1
users_ht[username]['none_%i' % rival_index] += 1
else:
if selected_party is None:
raise Exception('This should not happen 5691')
else:
this_bucket[selected_party] += 1
users_ht[username][selected_party] += 1
# yes_hts = len([x for x in hashtags if x in ht['ND']])
# no_hts = len([x for x in hashtags if x in ht['SYRIZA']])
# meimarakis_hts = len([x for x in hashtags if x in ht['MEIMARAKIS']])
# tsipras_hts = len([x for x in hashtags if x in ht['TSIPRAS']])
# if yes_hts == 0 and no_hts == 0:
# this_bucket['none'] += 1
# users_ht[username]['none'] += 1
# elif yes_hts > 0 and no_hts > 0:
# this_bucket['both'] += 1
# users_ht[username]['both'] += 1
# elif yes_hts > 0 and no_hts == 0:
# this_bucket['nd'] += 1
# users_ht[username]['nd'] += 1
# elif yes_hts == 0 and no_hts > 0:
# this_bucket['syriza'] += 1
# users_ht[username]['syriza'] += 1
# else:
# raise Exception('This should not happen')
#print yes_hts, no_hts
#print this_time
#if c > 50000:
# break
if False: # Print picks text
for p_index, p in enumerate(picks):
s = sorted(list(p[1].iteritems()), key=lambda x:x[1], reverse=True)
print p_index
for x in range(10):
print s[x][0], s[x][1]
if False:
with open('plot_5.json') as f:
to_plot_5 = json.load(f)
if False:
with open('plot_6.json') as f:
to_plot_6 = json.load(f)
print 'NO GREEK TWEETS:', not_greek
fig, ax = plt.subplots()
#YES
if True: # Make plot_1
leg_yes, = ax.plot([x[0] for x in to_plot_5], [x[1]['ND'] for x in to_plot_5], '-b')
leg_no, = ax.plot([x[0] for x in to_plot_5], [x[1]['SYRIZA'] for x in to_plot_5], '-r')
leg_meimarakis, = ax.plot([x[0] for x in to_plot_5], [x[1]['MEIMARAKIS'] for x in to_plot_5], '--b')
leg_tsipras, = ax.plot([x[0] for x in to_plot_5], [x[1]['TSIPRAS'] for x in to_plot_5], '--r')
#leg_none, = ax.plot([x[0] for x in to_plot_1], [x[1]['none'] for x in to_plot_1], '-k')
plt.legend([leg_yes, leg_no, leg_meimarakis, leg_tsipras], ['ND Tweets', 'SYRIZA Tweets', ' Meimarakis (ND leader) tweets', 'Tsipras (SYRIZA leader) tweets'])
ax.set_xlabel('Date (2015)')
ax.set_ylabel('Number of Tweets per day')
#ax.set_title('Frequency of Election tweets') # PLOS
ax.set_xlim(to_plot_5[0][0], to_plot_5[-1][0])
# ax.annotate('29 June 2015\nfirst NO demonstration', xy=(1.43558e9, 900), xytext=(1.43530e9, 1300),
# arrowprops=dict(facecolor='black', shrink=0.05, ),
# )
# ax.annotate('retweets of @atsipras\nNO promoting posts', xy=(1.43575e9, 1950), xytext=(1.43545e9, 2100),
# arrowprops=dict(facecolor='black', shrink=0.05, ),
# )
# ax.annotate('3 July 2015\nBig YES and NO demonstrations', xy=(1.43593e9, 2550), xytext=(1.43550e9, 2600),
# arrowprops=dict(facecolor='black', shrink=0.05, ),
# )
fix_time_ticks(ax, string_time='%e %b')
#Save plot data
with open('plot_5.json', 'w') as f:
f.write(json.dumps(to_plot_5) + '\n')
print 'Saved: plot_5.json'
save_figure('figure4')
save_figure('plot_5')
if False: # Make plot_2
#start_from = 1.43539e9
if False:
yes_key = 'yes_users_multi'
no_key = 'no_users_multi'
if True:
yes_key = 'yes_users'
no_key = 'no_users'
perc = [(100.0 * float(x[1][yes_key])/float(x[1][yes_key] + x[1][no_key]) if x[1][yes_key] + x[1][no_key] > 0 else 0.0 ) for x in to_plot_2 if x[0] > start_from]
#X = [x[0] for x in to_plot_2 if x[0] > start_from]
X = [x[0] for x in to_plot_6]
print '\n'.join(map(str, perc[-10:]))
leg_yes, = ax.plot(X, [x[1][yes_key] for x in to_plot_2 if x[0] > start_from], '-b')
leg_no, = ax.plot(X, [x[1][no_key] for x in to_plot_2 if x[0] > start_from], '-r')
ax2 = ax.twinx()
leg_perc, = ax2.plot(X, perc, '--k')
ax2.plot([X[0]-100000, X[-1]+100000], [38.7, 38.7], '-g', linewidth=6, alpha=0.6, color='0.2')
ax2.text(1.43557e9, 39.6, 'Result: YES=38.6%', fontsize=12)
ax.set_title('Variation of YES percentage')
ax.set_xlabel('Date (2015)')
ax.set_ylabel('Number of Users')
ax2.set_ylabel('Percentage of "YES"')
ax2.set_ylim((0.0, 70.0))
ax.set_xlim((X[0]-10000, X[-1] + 10000))
ax2.annotate('Capital Controls', xy=(1.43558e9, 25.0), xytext=(1.43540e9, 20.0),
arrowprops=dict(facecolor='0.8',
#shrink=0.05,
),
bbox=dict(boxstyle="round", fc="0.8"),
)
fix_time_ticks(ax, string_time='%e %b')
#fix_time_ticks(ax)
plt.legend([leg_yes, leg_no, leg_perc], ['YES users', 'NO users', 'YES percentage'], loc=2)
with open('plot_6.json', 'w') as f:
f.write(json.dumps(to_plot_6) + '\n')
print 'Saved: plot_6.json'
save_figure('plot_6')
ax
figure_4()
```
## Figure 5. Entities cooccurrence in Referendum
```
def plot_distances(
input_filename,
output_filename,
distance_fun = lambda x : x,
line_ignore = lambda x : False,
color_fun = lambda t : 'white',
):
'''
input_filename should have lines TERM1 TERM2 DISTANCE
'''
import pygraphviz as PG
G_gv = PG.AGraph()
G_gv.node_attr['style']='filled'
G_gv.node_attr['fillcolor']='white'
added_nodes = {}
with open(input_filename) as input_f:
for l in input_f:
ls = l.replace(os.linesep, '').split('__')
term_1 = ls[0]
term_2 = ls[1]
raw_distance = float(ls[2])
if line_ignore(term_1, term_2, raw_distance):
#print 'Ignored:', ls
continue
term_1_dec = term_1.decode('utf-8')
term_2_dec = term_2.decode('utf-8')
distance = distance_fun(term_1_dec, term_2_dec, raw_distance)
#term_1.encode('ascii', 'ignore')
#term_2.decode('utf-8')
#term_2.encode('ascii', 'ignore')
#G_gv.add_edge(term_1.encode('ascii', 'ignore'), term_2.encode('ascii', 'ignore'), len=distance)
#G_gv.add_edge(term_1.encode('utf-8'), term_2.encode('utf-8'), len=distance)
if term_1_dec not in added_nodes:
added_nodes[term_1_dec] = None
G_gv.add_node(term_1_dec, fillcolor=color_fun(term_1))
if term_2_dec not in added_nodes:
added_nodes[term_2_dec] = None
G_gv.add_node(term_2_dec, fillcolor=color_fun(term_2))
G_gv.add_edge(term_1_dec, term_2_dec, len=distance)
print 'Saving..'
G_gv.draw( input_filename + '.png', format='png', prog='neato')
G_gv.draw( input_filename + '.plain', format='plain', prog='neato')
G_gv.draw( input_filename + '.eps', format='eps', prog='neato')
G_gv.draw( output_filename + '.eps', format='eps', prog='neato')
G_gv.draw( output_filename + '.png', format='png', prog='neato')
print 'Saved:', input_filename + '.png'
print 'Saved:', input_filename + '.plain'
print 'Saved:', input_filename + '.eps'
print 'Saved:', output_filename + '.eps'
print 'Saved:', output_filename + '.png'
def figure_5():
kind = 'referendum'
occurences = {}
if True: # Build occurences
for item in merge_tweets_sentiment(kind):
hashtag_entites = item['hashtag_entites']
text_entities = item['text_entities']
all_entities = sorted(list(set(hashtag_entites + text_entities)))
if len(all_entities) < 2:
continue
for e1, e2 in itertools.combinations(all_entities, 2):
key = e1 + '__' + e2
if not key in occurences:
occurences[key] = 0
occurences[key] += 1
with open('occurences_%s.json' % (kind), 'w') as f:
json.dump(occurences, f)
print 'Saved: occurences_%s.json' % (kind)
if True:
nice = {
'greekdebt': 'debt',
'communistpartyofgreece': 'Communist Party',
'massmedia': 'media',
'greekgovernment': 'Government',
'antonissamaras': 'Antonis Samaras',
'hellenicparliament': 'parliament',
'theriver' : 'The River',
'stavrostheodorakis': 'Stavros Theodorakis',
'angelamerkel': 'Angela Merkel',
'alexistsipras': 'Alexis Tsipras',
'jeanclaudejuncker': 'Jean-Claude Juncker',
'capitalcontrols': 'Capital Controls',
'syriza': 'Syriza',
'greekprimeminister': 'Prime Minister',
'germany': 'Germany',
'imf': 'IMF',
'europe': 'Europe',
'voteno': 'NO',
'voteyes': 'YES',
'eurozone': 'Eurozone',
'yeseurope': 'YesEurope',
}
print 'Reading: occurences_%s.json' % (kind)
with open('occurences_%s.json' % (kind)) as f:
occurences = json.load(f)
#Sorting
occurences_sorted = sorted(occurences.iteritems(), key=lambda x:x[1], reverse=True)
filename = 'occurences_%s_sorted.txt' % (kind)
print 'Saving to:', filename
with codecs.open(filename, encoding='utf8', mode='w') as f:
for occ in occurences_sorted:
occ_s = occ[0].split('__')
for occ_s_i, s in enumerate(occ_s):
if s in nice:
occ_s[occ_s_i] = nice[s]
if 'greece' in occ_s:
continue
to_print = occ_s + [unicode(occ[1])]
f.write('__'.join(to_print) + '\n')
if True:
plot_distances(
input_filename = 'occurences_%s_sorted.txt' % (kind),
output_filename = 'figure5',
#distance_fun = lambda e1,e2,d : np.log(13883+10-d),
distance_fun = lambda e1,e2,d : np.log(13883+10-d) * 0.3,
line_ignore = lambda e1,e2,d : d<500,
#color_fun = lambda t : 'red' if t == 'voteno' else ('blue' if t == 'voteyes' else 'white')
color_fun = lambda t : '#FF7F50' if t == 'NO' else ('#5CB3FF' if t == 'YES' else 'white')
)
# OLD : ht_co_occurence.png
figure_5()
```

## Figure 9. Temporal variation of sentiment
```
def plot_sentiment(kind=None, posneg=None, ax=None, save=True, xlabel=None, ylabel=None, xticks=True, xtickfontsize=17, xtickapply=lambda x:x[1]):
'''
Sentiment over time
'''
if not kind:
kind='referendum'
#kind = 'elections'
to_plot_3 = {}
originals = {}
def correct_sarcasm(sentiment, sarcasm_percentage):
ret = sentiment
if sarcasm_percentage>0.0: # Correct
if sentiment > 0.0:
sentiment_value = (ret-1) / 4.0
elif sentiment <0.0:
sentiment_value = (-ret-1) / 4.0
else:
raise Exception('This should not happen 491266')
#Penalize sentiment
sentiment_value = sentiment_value - ((sarcasm_percentage/100.0) * sentiment_value)
#Extend to previous range
sentiment_value = 1 + (sentiment_value * 4.0)
#Assign right sign
if sentiment < 0:
sentiment_value = -sentiment_value
ret = sentiment_value
return ret
def add_sentiment(entity, all_entities, this_time, sp, sn, s0, is_original, sarcasm_percentage):
if entity in all_entities:
if sp > 1:
if not entity in to_plot_3:
to_plot_3[entity] = {'sp': [], 'sn': []}
to_plot_3[entity]['sp'].append((this_time, correct_sarcasm(sp, sarcasm_percentage), sp, is_original, sarcasm_percentage))
if sn < -1:
if not entity in to_plot_3:
to_plot_3[entity] = {'sp': [], 'sn': [], 'sp_nc': [], 'sn_nc':[], 's0': [] }
to_plot_3[entity]['sn'].append((this_time, correct_sarcasm(sn, sarcasm_percentage), sn, is_original, sarcasm_percentage))
if True:
for item in merge_tweets_sentiment(kind):
this_time = item['date']
sp = item['sp']
sn = item['sn']
s0 = item['s0']
original_text = item['original_text']
if this_time < 1435288026.0:
continue
if sp == 1 and sn == -1:
continue
# 634360327572951040
text_entities = item['text_entities']
#to_print.append(len(text_entities))
hashtag_entites = item['hashtag_entites']
sarcasm_percentage = item['sarcasm']['percentage']
all_entities = list(set(text_entities+hashtag_entites))
is_original = 1 if original_text in originals else 0
originals[original_text] = None
for x in all_entities:
add_sentiment(x, all_entities, this_time, sp, sn, s0, is_original, sarcasm_percentage)
if True:
with open('plot_3_%s.json' % (kind), 'w') as f:
json.dump(to_plot_3, f)
print 'Saved plot_3_%s.json' % (kind)
if True:
if kind == 'referendum':
with open('plot_3_referendum.json') as f: # mv plot_3.json plot_3_referendum.json
to_plot_3 = json.load(f)
print 'Opening: plot_3_referendum.json'
elif kind == 'elections':
with open('plot_3_elections.json') as f: # mv plot_3.json plot_3_elections.json
to_plot_3 = json.load(f)
print 'Opening: plot_3_elections.json'
if False: # Make CSV with sentiments for each entity
for kind in ['elections', 'referendum']:
with open('plot_3_%s.json' % kind) as f:
data = json.load(f)
for sentiment, key in [('positive', 'sp'), ('negative', 'sn')]:
for e in data:
print 'Kind:', kind, 'Entity:', e, 'Sentiment:', sentiment
filename = 'sentiments__%s_%s_%s.csv' % (kind, e, sentiment)
print 'Filename:', filename
with open(filename, 'w') as f:
for entry in data[e][key]:
to_print = ','.join(map(str, entry))
f.write(to_print + '\n')
if False:
# Which are the most common entities
f = [(x, max(len(y['sp']), len(y['sn']))) for x,y in to_plot_3.iteritems()]
fs = sorted(f, key=lambda x : x[1], reverse=True)
for x in fs:
print x
'''
REFERENDUM:
(u'voteno', 42699)
(u'greece', 35298)
(u'hellenicparliament', 11479)
(u'alexistsipras', 11474)
(u'europe', 10777)
(u'voteyes', 8588)
(u'syriza', 7473)
(u'yeseurope', 6562)
(u'bank', 6449)
(u'greekprimeminister', 6095)
(u'eurozone', 5801)
(u'greekgovernment', 5689)
(u'state', 5293)
(u'capitalcontrols', 4994)
(u'massmedia', 3529)
(u'jeanclaudejuncker', 3094)
(u'communistpartyofgreece', 3036)
'''
'''
ELECTIONS:
(u'alexistsipras', 12173)
(u'syriza', 8256)
(u'greece', 6613)
(u'vangelismeimarakis', 3526)
(u'communistpartyofgreece', 3281)
(u'state', 3044)
(u'anel', 2800)
(u'panagiotislafazanis', 2770)
(u'popularunity', 2761)
(u'newdemocracy', 2633)
(u'zoekonstantopoulou', 2427)
(u'greekgovernment', 2207)
(u'panhellenicsocialistmovement', 2053)
(u'theriver', 1903)
'''
minx = []
maxx = []
def plot_sentiment(entity, s, color, axes, linestyle = '-'):
if entity == 'Tsipras':
entity = 'AlexisTsipras'
if entity == 'Troika':
entity = 'EuropeanTroika'
if entity == 'Communist Party':
entity = 'communistpartyofgreece'
if entity == 'New Democracy':
entity = 'newdemocracy'
if entity == 'Government':
entity = 'greekgovernment'
if entity == 'PASOK':
entity = 'panhellenicsocialistmovement'
if entity == 'Varoufakis':
entity = 'YanisVaroufakis'
if entity == u'Schäuble':
entity = 'WolfgangSchauble'
if entity == 'Capital Controls':
entity = 'CapitalControls'
if entity == 'Debt':
entity = 'GreekDebt'
entity = entity.lower()
X = [x[0] for x in to_plot_3[entity][s]]
if s == 'sp':
Y = [x[1] for x in to_plot_3[entity][s]]
elif s == 'sn':
Y = [-x[1] for x in to_plot_3[entity][s]]
else:
raise Exception('sdgdfg34234')
#data = zip(X, Y)[0:100]
data = zip(X, Y)
df = pd.DataFrame(data, columns=["Date", "Y"])
print 'Entity: %s Data points: %i' % (entity, len(data))
print print_now_2(), 'Before regression..'
#axes.set(ylim=(1.6, 2.8))
g = seaborn.regplot(x="Date", y='Y', data=df, ax=axes,
lowess=True,
truncate=False,
#order = 2,
#x_ci = None,
#scatter_kws={'s':2, 'color':'k'},
line_kws={'color': color, 'linestyle': linestyle}, # 'alpha': line_alpha, 'linestyle': linestyle, 'linewidth': 1}
scatter=False,
)
#axes.set(ylim=(1.6, 2.8))
#leg, = plt.plot([], [], color= color, linestyle= linestyle)
leg, = plt.plot([], [], '.', color= color)
#print g
#print dir(g)
print print_now_2(), 'After regression'
#print X, Y
minx.append(min(X))
maxx.append(max(X))
return leg
#matplotlib.rcParams.update({'font.size': 22})
if True: # Plot
if not posneg:
#posneg = 'positive'
posneg = 'negative'
try:
seaborn.set_style("ticks")
except NameError as e:
print 'WARNING: SEABORN IS NOT IMPORTED!!!!'
if not ax:
fig, ax = plt.subplots()
if kind == 'referendum':
e_list = [
('Tsipras', 'blue'),
('hellenicparliament', 'red'),
('europe', 'green'),
('bank', 'm'),
('eurozone', 'y'),
]
elif kind == 'elections': # Elections
e_list = [
('Tsipras', 'blue'),
('syriza', 'red'),
('greece', 'green'),
('vangelismeimarakis', 'm'),
('communistpartyofgreece', 'y'),
]
#ax2 = ax.twinx()
legends = []
for e, e_color in e_list:
if posneg == 'positive':
leg = plot_sentiment(e, 'sp', e_color, ax)
elif posneg == 'negative':
leg = plot_sentiment(e, 'sn', e_color, ax)
legends.append(leg)
#plot_sentiment(e, 'sn', e_color, ax2, linestyle='--')
x_min = min(minx)
x_max = max(maxx)
ax.set_xlim((x_min, x_max))
if kind == 'referendum':
if posneg == 'positive':
ax.set(ylim=(1.6, 2.8))
elif posneg == 'negative':
ax.set(ylim=(1.74, 2.16))
elif kind == 'elections':
pass
#ax.set_ylim((1.0, 4.5))
#ax.set_xlabel('Date (2015)', fontsize=17)
#ax.set_xlabel('Date (2015)')
ax.set_xlabel('')
if posneg == 'positive':
if not ylabel:
ylabel = 'Positive Sentiment'
ax.set_ylabel(ylabel, fontsize=17)
elif posneg == 'negative':
if not ylabel:
ylabel = 'Negative Sentiment'
ax.set_ylabel(ylabel, fontsize=17)
#ax.set_title('Variation of Positive Sentiment in %s' % (kind))
print 'X LIMITS:', ax.get_xlim()
print 'Y LIMITS:', ax.get_ylim()
if kind == 'referendum':
if posneg == 'positive':
if not xlabel:
xlabel = 'Variation of Positive Sentiment in %s' % (kind)
ax.text(1435443938.0, 2.85, xlabel, fontsize=17 )
if save:
plt.legend(legends, ['Tsipras', 'Parliament', 'Europe', 'Bank', 'Eurozone'], fontsize=17) #
else:
plt.legend(legends, ['Tsipras', 'Parliament', 'Europe', 'Bank', 'Eurozone'], bbox_to_anchor=(-.05, -0.22, 1.0, -.138), loc=3, ncol=3, mode="expand", borderaxespad=0., ) # fontsize=17 $ (0., 1.02, 1., .102)
elif posneg == 'negative':
if not xlabel:
xlabel = 'Variation of Negative Sentiment in %s' % (kind)
ax.text(1435443938.0, 2.18, xlabel, fontsize=17 )
if save:
plt.legend(legends, ['Tsipras', 'Parliament', 'Europe', 'Bank', 'Eurozone'], bbox_to_anchor=(1.0, 1.03), fontsize=17)
if kind == 'elections':
if posneg == 'positive':
if not xlabel:
xlabel = 'Variation of Positive Sentiment in %s' % (kind)
ax.text(1440300000.0, 2.74, xlabel, fontsize=17 )
if save:
plt.legend(legends, ['Tsipras', 'Syriza', 'Greece', 'Meimarakis', 'Communist party'], loc=4, fontsize=17) # fontsize=17
else:
plt.legend(legends, ['Tsipras', 'Syriza', 'Greece', 'Meimarakis', 'Communism'], bbox_to_anchor=(-.05, -0.22, 1.1, -0.138), loc=3, ncol=3, mode="expand", borderaxespad=0.,) # fontsize=17
elif posneg == 'negative':
if not xlabel:
xlabel = 'Variation of Negative Sentiment in %s' % (kind)
if save:
ax.text(1440300000.0, 2.65, xlabel, fontsize=17 )
else:
pass
#plt.legend(legends, ['Tsipras', 'Syriza', 'Greece', 'Meimarakis', 'Communist party'], loc=2, bbox_to_anchor=(0.03, 0.93), fontsize=17) # fontsize=17
if xticks:
fix_time_ticks(ax, string_time='%e %b', fontsize=xtickfontsize, apply_index=xtickapply)
else:
fix_time_ticks(ax, string_time='', fontsize=xtickfontsize)
#ax2.set_ylabel('Negative Sentiment')
if save:
# plot_3b_ has smaller fonts
save_figure('plot_3c_%s_%s' % (posneg, kind))
plt.show()
#a=1/0
if False: # Plot
seaborn.set_style("ticks")
fig, ax = plt.subplots()
# ax.set_ylim((-3.0, +3.0))
#plot_sentiment('voteno', 'sn', color='b')
if kind == 'referendum':
e_list = [
#'EuropeanUnion',
['Tsipras', -2.05, -1.75, 0, -0.05, 0, +50000],
#['memorandum', -3, -1],
['Debt', -2.02, -1.9, -0.5, -0.04, 0, +50000],
['Capital Controls', -2.2, -1.5, 0, 0, -0.22, -50000],
['Europe', -2.1, -1.91, 0, +0.02, 0, 0],
[u'Schäuble', -2, -1.8, 0, -0.04, 0, -50000],
['Varoufakis', -2.2, -1.85, -0.5, -0.03, -0.1, 0],
['Economy', -2.0, -1.7, +0.02, 0.03, -0.25, 0],
#['Economy', -2.1, -1.9, 0.0, 0.0, -0.0, 0],
['Troika', -2.1, -1.8, 0, -0.03, 0, +50000],
'agreement',
'AngelaMerkel',
'JeroenDijsselbloem',
'IMF'
]
elif kind == 'elections': # Elections
e_list = [
['AlexisTsipras', -2.1, -1.6, 0, -0.05, 0, 0],
['Syriza', -2.2, -1.8, 0, 0, 0, 0],
['Greece', -2.2, -1.9, 0, -0.07, 0, 0],
['VangelisMeimarakis', -1.9, -1.8, 0, -0.01, 0, -500000],
['Communist Party', -2.6, -2.3, 0, -0.13, 0, -130000],
['New Democracy', -2.4, -1.9, 0, -0.08, 0, -100000],
['Government', -1.95, -1.8, 0, -0.02, 0, 0],
['PASOK', -2.5, -1.95, 0, -0.1, 0, 0],
]
all_axes = []
height = 0.1
for e_index, e_set in enumerate(e_list[:8]):
#for e_index, e_set in enumerate([e_list[1]]):
e = e_set[0]
e_min = e_set[1]
e_max = e_set[2]
boost_2_y_down = e_set[3] # From_y
move_text_y = e_set[4]
boost_2_y_up = e_set[5] # To_y
move_text_x = e_set[6]
ax_new = fig.add_axes([0.09, 0.1 + (height*e_index), 0.8, height])
##########
if True:
#ax_new2 = ax_new.twinx()
plot_sentiment(e, 'sp', 'b', ax_new)
ax_new.set_xticks([])
ax_new_ylim = ax_new.get_ylim()
ax_new_ylim = (ax_new_ylim[0] + boost_2_y_down, ax_new_ylim[1] + boost_2_y_up)
ax_new.set_ylim(ax_new_ylim)
ax_new.yaxis.set_ticks(np.arange(ax_new_ylim[0] ,ax_new_ylim[1], (ax_new_ylim[1]-ax_new_ylim[0])/3.0 ))
ax_new_labels = [ '%.2f' % (float(x)) if i else '' for i, x in enumerate(ax_new.get_yticks().tolist())]
ax_new.set_yticklabels(ax_new_labels)
##########
if True:
ax_new2 = ax_new.twinx()
ax_new2.set_ylim((e_min, e_max))
plot_sentiment(e, 'sn', 'r', ax_new2)
ax_new2.yaxis.set_ticks(np.arange(e_min, e_max, (e_max-e_min)/3.0 ))
ax_new2_labels = [ '%.2f' % (float(x)) if i else '' for i, x in enumerate(ax_new2.get_yticks().tolist())]
ax_new2.set_yticklabels(ax_new2_labels)
ax_new2.set_xticks([])
#ax_new.set_yticks([-2.5,-1.5])
if e_index != 0:
ax_new2.spines['bottom'].set_visible(False)
if e_index != 0:
ax_new.spines['bottom'].set_visible(False)
#ax_new.spines['left'].set_visible(False)
all_axes.append(ax_new)
all_axes.append(ax_new2)
#ax_new.text(1435960000 + move_text_x, (e_min + e_max)/2.0 + move_text_y , e, verticalalignment='bottom', bbox={'facecolor':'white', 'alpha':0.1})
if kind == 'referendum':
ax_new2.text(1435960000 + move_text_x, (e_min + e_max)/2.0 + move_text_y , e, verticalalignment='bottom')
elif kind == 'elections':
ax_new2.text(1442200000 + move_text_x, (e_min + e_max)/2.0 + move_text_y , e, verticalalignment='bottom')
if kind == 'referendum':
step = 100000
elif kind == 'elections':
step = 300000
rr = range(step/2 + int(min(minx)), int(max(maxx)), step)
print 'Minimum:', min(minx)
print 'Maximum:', max(maxx)
for a_index, a in enumerate(all_axes):
a.set_xlim(min(minx), max(maxx))
a.set_ylabel('')
a.set_xlabel('')
#ax.set_ylim((-2.0, +2.0))
if a_index == 0:
a.set_xticks(rr)
fix_time_ticks(a, string_time='%e %b')
elif a_index % 2 == 0:
a.set_xticks(rr)
ticks_a = a.get_xticks().tolist()
ticks_a = ['' for x in ticks_a]
#ax_new.set_yticklabels(new_ticks)
a.set_xticklabels(ticks_a)
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.spines['bottom'].set_visible(False)
ax.spines['left'].set_visible(False)
ax.set_xticks([])
ax.set_yticks([])
#ax.plot([0,1], [0,1], '-')
#ax.set_xlabel('Date (2015)')
ax.text(-0.15, 0.8, 'Positive Sentiment (Blue)', rotation='vertical',)
ax.text(1.08, 0.8, 'Negative Sentiment (Red)', rotation='vertical',)
if kind == 'referendum':
ax.text(0.2, 1.05, 'Variation of Sentiment in Referendum tweets')
elif kind == 'elections':
ax.text(0.2, 1.05, 'Variation of Sentiment in Elections tweets')
print print_now_2(), 'Finished'
# fix_time_ticks(ax, string_time='%e %b')
if kind == 'referendum':
save_figure('plot_3_referendum')
elif kind == 'elections':
save_figure('plot_3_elections')
plt.show()
def figure_9():
'''
- Figures containing multiple panels/subfigures must be combined into one image file before submission. https://mail.google.com/mail/u/0/#inbox/156382863ee888af
cp plot_3d.eps GitHub/phd/latex/referendum_plos_one/figures/
'''
current_plot = 0
for posneg in ['positive', 'negative']:
for kind in ['referendum', 'elections']:
current_plot += 1
ax = plt.subplot(2, 2, current_plot)
xtickapply = lambda x : x[1]
if current_plot == 1:
xlabel = 'Referendum'
ylabel = 'Positive'
xticks = False
elif current_plot == 2:
xlabel = 'Elections'
ylabel = ' '
xticks = False
elif current_plot == 3:
xlabel = ' '
ylabel = 'Negative'
xticks = True
xtickapply = lambda x : x[1] if not x[0]%2 else ''
elif current_plot == 4:
xlabel = ' '
ylabel = ' '
xticks = True
plot_sentiment(kind=kind, posneg=posneg, ax=ax, save=False, xlabel=xlabel, ylabel=ylabel, xticks=xticks, xtickfontsize=9, xtickapply=xtickapply)
plt.tick_params(axis='x', which='both', top='off', )
if current_plot in [1,2]:
plt.tick_params(axis='x', which='both', bottom='off', )
save_figure('figure9')
save_figure('plot_3d')
ax
figure_9()
```
## LDA Topic Model
The position of the clusters for LDA clustering depend on the random seed. Different runs will generate the same number of clusters and same distances. Nevertheless, the absolute locations of the clusters will differ. The functions below, plot the results from a specific LDA run. The files with the results of these runs are ``ldasave_referendum.html`` and ``ldasave_elections.html``. These files exist in this repository.
```
class My_LDA():
def __init__(self):
pass
#@timer
def fit_LDA_model(self, corpus, id2word, num_topics=50, passes=10):
from gensim import models
print 'Fitting the LDA model..'
lda = models.ldamodel.LdaModel(corpus=corpus, id2word=id2word, num_topics=num_topics, passes=passes)
return lda
def visualize_LDA_model(self, lda, corpus, dictionary, output_filename):
import pyLDAvis.gensim as gensimvis
import pyLDAvis
print 'Visualizing LDA model..'
vis_data = gensimvis.prepare(lda, corpus, dictionary)
#with open('ldasave.html', 'w') as f:
with open(output_filename, 'w') as f:
pyLDAvis.save_html(vis_data, f)
print 'Created:', output_filename
def LDA(self, list_of_docs, num_topics=50, no_above=0.5, output_filename='ldasave.html'):
dictionary, corpus = self.build_dictionary(list_of_docs, no_above=no_above)
lda = self.fit_LDA_model(corpus, dictionary, num_topics=num_topics)
self.visualize_LDA_model(lda, corpus, dictionary, output_filename)
def build_dictionary(self, docs, no_below=5, no_above=0.5):
from gensim.corpora import Dictionary, MmCorpus
print 'Building dictionary..'
dictionary = Dictionary(docs)
dictionary.compactify()
dictionary.filter_extremes(no_below=no_below, no_above=no_above, keep_n=None)
dictionary.compactify()
print 'Building corpus..'
corpus = [dictionary.doc2bow(doc) for doc in docs]
return dictionary, corpus
def do_LDA(figure_to_plot, output_html):
'''
LDA for referendum
LDA referendum
ldasave_1.html --> No sarcasm filtering & remove voteyes, voteno
ldasave_2.html --> Sarcasm > 5% & remove voteyes, voteno
ldasave_3.html --> Sarcasm filtering, add voteyes, add noteno
'''
if figure_to_plot == 10:
kind = 'referendum'
elif figure_to_plot == 11:
kind = 'elections'
else:
raise Exception('Unknown value for figure_to_plot parameter')
c = 0
sentiment_entities = {}
entity_collection = []
for item in merge_tweets_sentiment(kind):
#print item['sarcasm']['percentage'], item['sarcasm']['score']
c += 1
this_time = item['date']
if this_time < 1435288026.0:
continue
#this_entities = []
if False:
for level in [1,2]:
for entity_set in item[str(level)]:
for entity_name in entity_set[1]:
if entity_name[0] == '#':
entity_name = entity_name[1:]
#if entity_name in ['referendum', 'dimopsifisma', 'voteYes', 'voteNo']:
# continue
if entity_name in ['referendum', 'dimopsifisma']:
continue
#if True: # Remove sarcasm
# if item['sarcasm']['percentage'] > 5.0:
# continue
this_entities.append(discretize_sarcasm(item['sarcasm']['percentage']))
if not entity_name in this_entities:
this_entities.append(entity_name)
if True:
if item['sarcasm']['percentage'] > 5.0:
continue
if True:
text_entities = item['text_entities']
hashtag_entites = item['hashtag_entites']
all_entities = list(set(text_entities + hashtag_entites))
if False:
all_entities = [x for x in all_entities if not x in ['referendum', 'dimopsifisma', 'voteyes', 'voteno']]
if True:
all_entities = [x for x in all_entities if not x in ['referendum', 'dimopsifisma']]
if kind == 'referendum':
if (('voteyes' in all_entities) or ('yeseurope' in all_entities)) and (('voteno' in all_entities) or ('noeurope' in all_entities)):
continue
if len(all_entities) > 1:
#print c, ' '.join(all_entities)
if True:
#Store sentiment entities
if item['sp'] != 1:
for x in all_entities:
if not x in sentiment_entities:
sentiment_entities[x] = {'sp': [], 'sn':[]}
sentiment_entities[x]['sp'].append( (item['sp']-1)/float(4) )
if item['sn'] != -1:
for x in all_entities:
if not x in sentiment_entities:
sentiment_entities[x] = {'sp': [], 'sn': []}
sentiment_entities[x]['sn'].append( (-item['sn']-1)/float(4))
entity_collection.append(all_entities)
if True:
print 'Saving sentiment entities'
for x in sentiment_entities:
if sentiment_entities[x]['sp']:
sentiment_entities[x]['sp'] = np.mean(sentiment_entities[x]['sp'])
if sentiment_entities[x]['sn']:
sentiment_entities[x]['sn'] = np.mean(sentiment_entities[x]['sn'])
with open('sentiment_entities_%s.json' % (kind), 'w') as f:
f.write(json.dumps(sentiment_entities))
print 'Saved: sentiment_entities.json'
#a=1/0
print print_now_2(), 'Performing LDA..'
lda = My_LDA()
lda.LDA(entity_collection, num_topics=5, output_filename='ldasave_{}_{}.html'.format(kind, output_html)) # , no_above=0.7)
print print_now_2(), 'Done LDA'
do_LDA(10, 'run1')
do_LDA(11, 'run1')
def figure_11(figure_to_plot):
'''
process ldasave.html
Plot circles
'''
import matplotlib.patches as mpatches
from matplotlib.collections import PatchCollection
if figure_to_plot == 10:
kind = 'referendum'
elif figure_to_plot == 11:
kind = 'elections'
else:
raise Exception('Invalid value in figure_to_plot parameter')
add_colorbars = False
add_sentiment_color_in_cicles = False
voteno_x_offset = 0.18
remove_axes = True
with open('ldasave_%s.html' % (kind)) as f:
for l in f:
if 'var ldavis_' in l:
l2 = re.sub(r'var ldavis_el[\d]+_data = ', '', l)
l2 = l2.replace(';', '')
d = json.loads(l2)
#print json.dumps(d, indent=4)
X = d['mdsDat']['x']
Y = d['mdsDat']['y']
freq = d['mdsDat']['Freq']
topic_label = d['mdsDat']['topics']
#print X,Y,freq
topic = d['token.table']['Topic']
fr = d['token.table']['Freq']
term = d['token.table']['Term']
#print len(topic), len(fr), len(term)
counts_name = d['tinfo']['Term']
counts_freq = d['tinfo']['Freq']
assert len(counts_name) == len(counts_freq)
term_freq = dict(zip(counts_name, counts_freq))
with open('sentiment_entities_%s.json' % (kind)) as f:
sentiment_entities = json.load(f)
print 'Opening: sentiment_entities_%s.json' % (kind)
topic_color = {}
for x, y, z in zip(topic, fr, term):
if not z in sentiment_entities:
continue
if not x in topic_color:
topic_color[x] = {'sp': [], 'sn': [], 'sum': 0}
sp = sentiment_entities[z]['sp']
sn = sentiment_entities[z]['sn']
this_counts = term_freq[z] * y
topic_color[x]['sp'].append((sp if type(sp) is float else 0.0, this_counts))
topic_color[x]['sn'].append((sn if type(sn) is float else 0.0, this_counts))
topic_color[x]['sum'] += this_counts
for x in topic_color:
#print '========================'
#print topic_color[x]['sp']
#print '========================'
topic_color[x]['final_sp'] = sum([ (y[1]/float(topic_color[x]['sum'])) * y[0] for y in topic_color[x]['sp']])
topic_color[x]['final_sn'] = sum([ (y[1]/float(topic_color[x]['sum'])) * y[0] for y in topic_color[x]['sn']])
print 'Topic:', x, 'sp:', topic_color[x]['final_sp'], 'sn:', topic_color[x]['final_sn']
min_sp = min([topic_color[x]['final_sp'] for x in topic_color])
max_sp = max([topic_color[x]['final_sp'] for x in topic_color])
min_sn = min([topic_color[x]['final_sn'] for x in topic_color])
max_sn = max([topic_color[x]['final_sn'] for x in topic_color])
for x in topic_color:
topic_color[x]['final_sp'] = (((topic_color[x]['final_sp'] - min_sp) * (0.9 - 0.1)) / (max_sp - min_sp)) + 0.1
topic_color[x]['final_sn'] = (((topic_color[x]['final_sn'] - min_sn) * (0.9 - 0.1)) / (max_sn - min_sn)) + 0.1
seaborn.set_style("white")
#seaborn.set_context(rc = {'patch.linewidth': 0.0})
#seaborn.despine()
fig, ax = plt.subplots()
#fig = plt.figure(dpi=600)
#ax = fig.gca()
patches = []
# ax.set_xlim((-0.6, 0.8))
cb_pos_x = 0.19
cb_pos_y = 0.18
cb_width = 0.02
cb_height = 0.2
cb_fz = 8
cmap_left = plt.cm.Blues
cax_left = fig.add_axes([cb_pos_x, cb_pos_y, cb_width, cb_height])
norm_left = matplotlib.colors.Normalize(vmin=0.0, vmax=1.0)
cb_left = matplotlib.colorbar.ColorbarBase(cax_left, cmap=cmap_left, norm=norm_left, spacing='proportional', ticks=[0, 1])
cb_left.ax.yaxis.set_ticks_position('left')
# cb_left.ax.set_yticklabels(['Week\nPositive\nSentiment', 'Strong\nPositive\nSentiment'], cb_fz)
cmap_right = plt.cm.Reds
cax_right = fig.add_axes([cb_pos_x+cb_width, cb_pos_y, cb_width, cb_height])
norm_right = matplotlib.colors.Normalize(vmin=0.0, vmax=1.0)
cb_right = matplotlib.colorbar.ColorbarBase(cax_right, cmap=cmap_right, norm=norm_right, spacing='proportional', ticks=[0, 1])
# cb_right.ax.yaxis.set_ticks_position('right')
# cb_right.ax.set_yticklabels(['Week\nNegative\nSentiment', 'Strong\nNegative\nSentiment'], cb_fz)
if not add_colorbars: # http://stackoverflow.com/questions/5263034/remove-colorbar-from-figure-in-matplotlib
fig.delaxes(cax_left)
fig.delaxes(cax_right)
def add_semisircle(x,y,radius, color_sp, color_sn):
if add_sentiment_color_in_cicles:
colorVal_left = cb_left.to_rgba(color_sp)
colorVal_right = cb_right.to_rgba(color_sn)
#Left
wedge = mpatches.Wedge((x, y), radius, 90, 90+180)
collection = PatchCollection([wedge], alpha=0.7, color=colorVal_left)
ax.add_collection(collection)
#Right
wedge = mpatches.Wedge((x, y), radius, 90+180, 90)
collection = PatchCollection([wedge], alpha=0.7, color=colorVal_right)
ax.add_collection(collection)
else:
wedge = mpatches.Wedge((x, y), radius, 0, 360)
collection = PatchCollection([wedge], alpha=0.7, color='k')
ax.add_collection(collection)
for x,y,f,t in zip(X,Y,freq,topic_label):
print x,y
add_semisircle(x,y,f/300.0, topic_color[t]['final_sp'], topic_color[t]['final_sn'])
#dd_semisircle(x,y,f/300.0, 0.1, 0.9) # 0.1 = black, 0.9 = white
#ax.add_collection(collection)
ax.axis('equal')
#ax.axis('off')
# ax.spines['top'].set_visible(False)
# ax.spines['right'].set_visible(False)
# if not plot_bottom_spine:
# ax.spines['bottom'].set_visible(False)
# if not plot_left_spine:
# ax.spines['left'].set_visible(False)
def scale(old_value, old_min, old_max, new_min, new_max):
return (((old_value - old_min) * (new_max - new_min)) / (old_max - old_min)) + new_min
def weighted_sentiment_entity(entity, posneg):
# Referendum: 2.1 - 3.0 ----- 2.0 - 2.8
# Elections: 2.09 - 2.94 ----- 2.12 - 3.03
if kind == 'referendum':
old_min_p = 2.1
old_max_p = 3.0
old_min_n = 2.0
old_max_n = 2.8
elif kind == 'elections':
old_min_p = 2.05
old_max_p = 2.97
old_min_n = 2.09
old_max_n = 3.06
new_min = 0.01
new_max = 0.99
if posneg == 'sp':
s = 1 + (sentiment_entities[entity][posneg] * 4.0)
return scale(s, old_min_p, old_max_p, new_min, new_max)
elif posneg == 'sn':
s = 1 + (sentiment_entities[entity][posneg] * 4.0)
return scale(s, old_min_n, old_max_n, new_min, new_max)
else:
raise Exception('234EDF#$23323')
def text_sentiment_entites(e):
e = e.lower()
sp = 1 + (sentiment_entities[e]['sp']*4.0)
sn = 1 + (sentiment_entities[e]['sn']*4.0)
#return '%.2f/%.2f' % (sp, sn)
return ''
#text_x_offset = 0.2
# \u2588 --> block
def add_rectangle_sentiment(e, r_x, r_y, order):
r_length = 0.1
r_length2 = r_length / 2.0
if kind == 'referendum':
r_height = 0.02
r_y_order = r_y - ((order-1) * 0.027)
elif kind == 'elections':
r_height = 0.024
r_y_order = r_y - ((order-1) * 0.032)
rect = mpatches.Rectangle((r_x, r_y_order), r_length, r_height, )
collection = PatchCollection([rect], facecolor='white')
ax.add_collection(collection)
rect = mpatches.Rectangle((r_x + r_length2 - (weighted_sentiment_entity(e, 'sp') * r_length2), r_y_order), weighted_sentiment_entity(e, 'sp') * r_length2, r_height, )
collection = PatchCollection([rect], facecolor='blue')
ax.add_collection(collection)
rect = mpatches.Rectangle((r_x + r_length2, r_y_order), weighted_sentiment_entity(e, 'sn') * r_length2, r_height, )
collection = PatchCollection([rect], facecolor='red')
ax.add_collection(collection)
if kind == 'referendum':
text_x_offset = 0.04
ax.annotate(u'Germany %s\nVaroufakis %s\nTroika %s\nEurogroup %s' %
(
text_sentiment_entites('germany'),
text_sentiment_entites('yanisvaroufakis'),
text_sentiment_entites('europeantroika'),
text_sentiment_entites('eurogroup'),
),
xy=(-0.379463285475, 0.0592180667832), xytext=(-0.42 + text_x_offset, 0.13),
#arrowprops=dict(facecolor='black', shrink=0.05, ),
horizontalalignment='right',
)
# Add rectangle indicator for entities
# class matplotlib.patches.Rectangle(xy, width, height, angle=0.0, **kwargs)
r_x = -0.36
r_y = 0.21
add_rectangle_sentiment('germany', r_x = r_x, r_y = r_y, order=1)
add_rectangle_sentiment('yanisvaroufakis', r_x = r_x, r_y = r_y, order=2)
add_rectangle_sentiment('europeantroika', r_x = r_x, r_y = r_y, order=3)
add_rectangle_sentiment('eurogroup', r_x = r_x, r_y = r_y, order=4)
text_x_offset = 0.15
#ax.annotate('Prime Minister %s\nGovernment %s\nParliament %s\nMinisters %s' % (
ax.annotate('Prime Minister %s\nGovernment %s\nParliament %s\nGreece %s' % (
text_sentiment_entites('greekprimeminister'),
text_sentiment_entites('greekgovernment'),
text_sentiment_entites('hellenicparliament'),
#text_sentiment_entites('minister'),
text_sentiment_entites('greece'),
),
xy=(-0.379463285475, 0.0592180667832), xytext=(0.25 + text_x_offset, 0.29),
#arrowprops=dict(facecolor='black', shrink=0.05, ),
horizontalalignment='right',
)
# Add rectangle indicator for entities
# class matplotlib.patches.Rectangle(xy, width, height, angle=0.0, **kwargs)
r_x = 0.42
r_y = 0.37
add_rectangle_sentiment('greekprimeminister', r_x = r_x, r_y = r_y, order=1)
add_rectangle_sentiment('greekgovernment', r_x = r_x, r_y = r_y, order=2)
add_rectangle_sentiment('hellenicparliament', r_x = r_x, r_y = r_y, order=3)
add_rectangle_sentiment('greece', r_x = r_x, r_y = r_y, order=4)
# ax.annotate('VoteNo\nFreedom\nFar Right\nFar Left', xy=(-0.379463285475, 0.0592180667832), xytext=(0.17, 0.0),
#arrowprops=dict(facecolor='black', shrink=0.05, ),
# )
text_x_offset = 0.15
#ax.annotate('VoteNo %s\nFreedom %s\nFar Right %s\nFar Left %s' % (
#ax.annotate('VoteNo %s\nFreedom %s\nConservatives %s\nCommunists %s' % (
ax.annotate('Freedom %s\nConservatives %s\nCommunists %s\nGreece %s' % (
#text_sentiment_entites('voteno'),
text_sentiment_entites('freedom'),
text_sentiment_entites('anel'),
text_sentiment_entites('communistpartyofgreece'),
text_sentiment_entites('greece'),
),
xy=(-0.379463285475, 0.0592180667832), xytext=(-0.01 + voteno_x_offset + text_x_offset, -0.05),
#arrowprops=dict(facecolor='black', shrink=0.05, ),
horizontalalignment='right',
)
r_x = 0.34
r_y = 0.03
add_rectangle_sentiment('freedom', r_x = r_x, r_y = r_y, order=1)
add_rectangle_sentiment('anel', r_x = r_x, r_y = r_y, order=2)
add_rectangle_sentiment('communistpartyofgreece', r_x = r_x, r_y = r_y, order=3)
add_rectangle_sentiment('greece', r_x = r_x, r_y = r_y, order=4)
#VoteNo in the center
ax.annotate('#VoteNo',
xy=(-0.379463285475, 0.0592180667832),
xytext=(-0.02, -0.042),
style='italic',
color='white',
#bbox={'facecolor':'white', 'alpha':0.5, 'pad':1}
)
text_x_offset = 0.16
ax.annotate('Alexis Tsipras %s\nSYRIZA %s\nEconomy %s' % (
text_sentiment_entites('alexistsipras'),
text_sentiment_entites('syriza'),
text_sentiment_entites('economy'),
),
xy=(0.14, -0.14),
xytext=(0.27 + text_x_offset, -0.188),
# arrowprops=dict(facecolor='black', shrink=0.05, ),
horizontalalignment='right',
)
ax.arrow(0.27, -0.15, -0.09, +0.00, color='black')
r_x = 0.45
r_y = -0.137
add_rectangle_sentiment('alexistsipras', r_x = r_x, r_y = r_y, order=1)
add_rectangle_sentiment('syriza', r_x = r_x, r_y = r_y, order=2)
add_rectangle_sentiment('economy', r_x = r_x, r_y = r_y, order=3)
text_x_offset = -0.1
#ax.annotate('VoteYes %s\nDebt %s\nCapital Controls %s\nEU %s' % (
ax.annotate('Bank %s\nDebt %s\nCapital Controls %s\nEU %s' % (
#text_sentiment_entites('voteyes'),
text_sentiment_entites('bank'),
text_sentiment_entites('greekdebt'),
text_sentiment_entites('capitalcontrols'),
text_sentiment_entites('europeanunion'),
),
xy=(-0.379463285475, 0.0592180667832), xytext=(-0.08 + text_x_offset , -0.26),
horizontalalignment='right',
#arrowprops=dict(facecolor='black', shrink=0.05, ),
)
r_x = -0.16
r_y = -0.18
add_rectangle_sentiment('bank', r_x = r_x, r_y = r_y, order=1)
add_rectangle_sentiment('greekdebt', r_x = r_x, r_y = r_y, order=2)
add_rectangle_sentiment('capitalcontrols', r_x = r_x, r_y = r_y, order=3)
add_rectangle_sentiment('europeanunion', r_x = r_x, r_y = r_y, order=4)
#VoteNo in the center
ax.annotate('#VoteYes',
xy=(-0.379463285475, 0.0592180667832),
xytext=(+0.038, -0.23),
style='italic',
color='white',
#bbox={'facecolor':'white', 'alpha':0.5, 'pad':1}
)
ax.arrow(-0.06, -0.21, +0.05, +0.00, color='black')
leg_p, = ax.plot([], [], '-', color='b', linewidth=3)
leg_n, = ax.plot([], [], '-', color='r', linewidth=3)
#cmap = plt.cm.gray
if not add_colorbars:
ax.set_xlim((-0.40, 0.5))
ax.set_xlabel('Principal Component 1')
ax.set_ylabel('Principal Component 2')
#ax.set_title('Referendum LDA Topic Model') # PLOS
print 'Y Limits:', ax.get_ylim()
# ax.set_ylim((-0.4, 0.5))
plt.legend([leg_p, leg_n], ['Positive Sentiment', 'Negative Sentiment'], loc=2)
elif kind == 'elections':
bar_distance_x = 0.00
bar_distance_y = 0.095
x = 0.0
y = 0.23
ax.annotate(u'Syriza %s\nTsipras %s\nERT %s\nCorruption %s' %
(
text_sentiment_entites('syriza'),
text_sentiment_entites('alexistsipras'),
text_sentiment_entites('hellenicbroadcastingcorporation'),
text_sentiment_entites('corruption'),
),
xy=(0.0, 0.0), xytext=(x, y),
#arrowprops=dict(facecolor='black', shrink=0.05, ),
horizontalalignment='right',
)
# Add rectangle indicator for entities
# class matplotlib.patches.Rectangle(xy, width, height, angle=0.0, **kwargs)
r_x = x + bar_distance_x
r_y = y + bar_distance_y
add_rectangle_sentiment('syriza', r_x = r_x, r_y = r_y, order=1)
add_rectangle_sentiment('alexistsipras', r_x = r_x, r_y = r_y, order=2)
add_rectangle_sentiment('hellenicbroadcastingcorporation', r_x = r_x, r_y = r_y, order=3)
add_rectangle_sentiment('corruption', r_x = r_x, r_y = r_y, order=4)
######################################
x = 0.08
y = 0.02
ax.annotate(u'Konstantopoulou %s\nGovernment %s\nPASOK %s\nParliament %s' %
(
text_sentiment_entites('zoekonstantopoulou'),
text_sentiment_entites('greekgovernment'),
text_sentiment_entites('panhellenicsocialistmovement'),
text_sentiment_entites('hellenicparliament'),
),
xy=(0.0, 0.0), xytext=(x, y),
#arrowprops=dict(facecolor='black', shrink=0.05, ),
horizontalalignment='right',
)
r_x = x + bar_distance_x
r_y = y + bar_distance_y
add_rectangle_sentiment('zoekonstantopoulou', r_x = r_x, r_y = r_y, order=1)
add_rectangle_sentiment('greekgovernment', r_x = r_x, r_y = r_y, order=2)
add_rectangle_sentiment('panhellenicsocialistmovement', r_x = r_x, r_y = r_y, order=3)
add_rectangle_sentiment('hellenicparliament', r_x = r_x, r_y = r_y, order=4)
######################################
x = 0.0
y = -0.46
ax.annotate(u'Communism %s\nLAE %s\nLafazanis %s\nKoutsoumpas %s' %
(
text_sentiment_entites('communistpartyofgreece'),
text_sentiment_entites('popularunity'),
text_sentiment_entites('panagiotislafazanis'),
text_sentiment_entites('dimitriskoutsoumpas'),
),
xy=(0.0, 0.0), xytext=(x, y),
#arrowprops=dict(facecolor='black', shrink=0.05, ),
horizontalalignment='right',
)
r_x = x + bar_distance_x
r_y = y + bar_distance_y
add_rectangle_sentiment('communistpartyofgreece', r_x = r_x, r_y = r_y, order=1)
add_rectangle_sentiment('popularunity', r_x = r_x, r_y = r_y, order=2)
add_rectangle_sentiment('panagiotislafazanis', r_x = r_x, r_y = r_y, order=3)
add_rectangle_sentiment('dimitriskoutsoumpas', r_x = r_x, r_y = r_y, order=4)
#############################################
x = -0.5
y = 0.15
ax.annotate(u'New Democracy %s\nGreece %s\nMeimarakis %s\nThe River %s' %
(
text_sentiment_entites('newdemocracy'),
text_sentiment_entites('greece'),
text_sentiment_entites('vangelismeimarakis'),
text_sentiment_entites('theriver'),
),
xy=(-0.36948, 0.0502176), xytext=(x, y),
arrowprops=dict(facecolor='black', arrowstyle="->", connectionstyle='arc3,rad=0.3', linewidth=1),
horizontalalignment='right',
)
r_x = x + bar_distance_x
r_y = y + bar_distance_y
add_rectangle_sentiment('newdemocracy', r_x = r_x, r_y = r_y, order=1)
add_rectangle_sentiment('greece', r_x = r_x, r_y = r_y, order=2)
add_rectangle_sentiment('vangelismeimarakis', r_x = r_x, r_y = r_y, order=3)
add_rectangle_sentiment('theriver', r_x = r_x, r_y = r_y, order=4)
############################################
x = -0.5
y = -0.26
ax.annotate(u'Conservatives %s\nGreece %s\nState %s\nDebt %s' %
(
text_sentiment_entites('anel'),
text_sentiment_entites('greece'),
text_sentiment_entites('state'),
text_sentiment_entites('greekdebt'),
),
xy=(-0.371287, -0.0317222), xytext=(x, y),
arrowprops=dict(facecolor='black', arrowstyle="->", connectionstyle='arc3,rad=-0.3', linewidth=1),
horizontalalignment='right',
)
r_x = x + bar_distance_x
r_y = y + bar_distance_y
add_rectangle_sentiment('anel', r_x = r_x, r_y = r_y, order=1)
add_rectangle_sentiment('greece', r_x = r_x, r_y = r_y, order=2)
add_rectangle_sentiment('state', r_x = r_x, r_y = r_y, order=3)
add_rectangle_sentiment('greekdebt', r_x = r_x, r_y = r_y, order=4)
leg_p, = ax.plot([], [], '-', color='b', linewidth=3)
leg_n, = ax.plot([], [], '-', color='r', linewidth=3)
#ax.set_title('Elections LDA Topic Model') # PLOS
plt.legend([leg_p, leg_n], ['Positive Sentiment', 'Negative Sentiment'], loc=3)
if remove_axes:
plt.axis('off')
plt.tight_layout()
#save_figure('plot_4', formats=['png', 'pdf'])
save_figure('plot_4_%s' % (kind), formats=['png', 'pdf', 'eps', 'tiff'])
plt.savefig('plot_4_%s_300dpi.tiff' % (kind), format='tiff', dpi=300) # https://stackoverflow.com/questions/12192661/matplotlib-increase-resolution-to-see-details
if kind == 'referendum':
print 'Saving figure11.pdf for publication'
#ax.set_rasterized(True) # https://stackoverflow.com/questions/19638773/matplotlib-plots-lose-transparency-when-saving-as-ps-eps
fig.savefig('figure11.pdf', dpi=300)
elif kind == 'elections':
print 'Saving figure12.pdf for publication'
fig.savefig('figure12.pdf', dpi=300)
ax
```
## Figure 12. LDA topic model for Referendum
```
figure_11(10)
```
There is a known problem about generating transparent eps files. More information: https://stackoverflow.com/questions/19638773/matplotlib-plots-lose-transparency-when-saving-as-ps-eps The best solution seems to be to generate .pdf files and use the pdftops command:
```
!pdftops -eps -r 300 figure11.pdf figure11.eps
figure_11(11)
!pdftops -eps -r 300 figure12.pdf figure12.eps
```
| github_jupyter |
# Building your Deep Neural Network: Step by Step
Welcome to your week 4 assignment (part 1 of 2)! You have previously trained a 2-layer Neural Network (with a single hidden layer). This week, you will build a deep neural network, with as many layers as you want!
- In this notebook, you will implement all the functions required to build a deep neural network.
- In the next assignment, you will use these functions to build a deep neural network for image classification.
**After this assignment you will be able to:**
- Use non-linear units like ReLU to improve your model
- Build a deeper neural network (with more than 1 hidden layer)
- Implement an easy-to-use neural network class
**Notation**:
- Superscript $[l]$ denotes a quantity associated with the $l^{th}$ layer.
- Example: $a^{[L]}$ is the $L^{th}$ layer activation. $W^{[L]}$ and $b^{[L]}$ are the $L^{th}$ layer parameters.
- Superscript $(i)$ denotes a quantity associated with the $i^{th}$ example.
- Example: $x^{(i)}$ is the $i^{th}$ training example.
- Lowerscript $i$ denotes the $i^{th}$ entry of a vector.
- Example: $a^{[l]}_i$ denotes the $i^{th}$ entry of the $l^{th}$ layer's activations).
Let's get started!
## 1 - Packages
Let's first import all the packages that you will need during this assignment.
- [numpy](www.numpy.org) is the main package for scientific computing with Python.
- [matplotlib](http://matplotlib.org) is a library to plot graphs in Python.
- dnn_utils provides some necessary functions for this notebook.
- testCases provides some test cases to assess the correctness of your functions
- np.random.seed(1) is used to keep all the random function calls consistent. It will help us grade your work. Please don't change the seed.
```
import numpy as np
import h5py
import matplotlib.pyplot as plt
from testCases_v3 import *
from dnn_utils_v2 import sigmoid, sigmoid_backward, relu, relu_backward
%matplotlib inline
plt.rcParams['figure.figsize'] = (5.0, 4.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
%load_ext autoreload
%autoreload 2
np.random.seed(1)
```
## 2 - Outline of the Assignment
To build your neural network, you will be implementing several "helper functions". These helper functions will be used in the next assignment to build a two-layer neural network and an L-layer neural network. Each small helper function you will implement will have detailed instructions that will walk you through the necessary steps. Here is an outline of this assignment, you will:
- Initialize the parameters for a two-layer network and for an $L$-layer neural network.
- Implement the forward propagation module (shown in purple in the figure below).
- Complete the LINEAR part of a layer's forward propagation step (resulting in $Z^{[l]}$).
- We give you the ACTIVATION function (relu/sigmoid).
- Combine the previous two steps into a new [LINEAR->ACTIVATION] forward function.
- Stack the [LINEAR->RELU] forward function L-1 time (for layers 1 through L-1) and add a [LINEAR->SIGMOID] at the end (for the final layer $L$). This gives you a new L_model_forward function.
- Compute the loss.
- Implement the backward propagation module (denoted in red in the figure below).
- Complete the LINEAR part of a layer's backward propagation step.
- We give you the gradient of the ACTIVATE function (relu_backward/sigmoid_backward)
- Combine the previous two steps into a new [LINEAR->ACTIVATION] backward function.
- Stack [LINEAR->RELU] backward L-1 times and add [LINEAR->SIGMOID] backward in a new L_model_backward function
- Finally update the parameters.
<img src="images/final outline.png" style="width:800px;height:500px;">
<caption><center> **Figure 1**</center></caption><br>
**Note** that for every forward function, there is a corresponding backward function. That is why at every step of your forward module you will be storing some values in a cache. The cached values are useful for computing gradients. In the backpropagation module you will then use the cache to calculate the gradients. This assignment will show you exactly how to carry out each of these steps.
## 3 - Initialization
You will write two helper functions that will initialize the parameters for your model. The first function will be used to initialize parameters for a two layer model. The second one will generalize this initialization process to $L$ layers.
### 3.1 - 2-layer Neural Network
**Exercise**: Create and initialize the parameters of the 2-layer neural network.
**Instructions**:
- The model's structure is: *LINEAR -> RELU -> LINEAR -> SIGMOID*.
- Use random initialization for the weight matrices. Use `np.random.randn(shape)*0.01` with the correct shape.
- Use zero initialization for the biases. Use `np.zeros(shape)`.
```
# GRADED FUNCTION: initialize_parameters
def initialize_parameters(n_x, n_h, n_y):
"""
Argument:
n_x -- size of the input layer
n_h -- size of the hidden layer
n_y -- size of the output layer
Returns:
parameters -- python dictionary containing your parameters:
W1 -- weight matrix of shape (n_h, n_x)
b1 -- bias vector of shape (n_h, 1)
W2 -- weight matrix of shape (n_y, n_h)
b2 -- bias vector of shape (n_y, 1)
"""
np.random.seed(1)
### START CODE HERE ### (≈ 4 lines of code)
W1 = None
b1 = None
W2 = None
b2 = None
### END CODE HERE ###
assert(W1.shape == (n_h, n_x))
assert(b1.shape == (n_h, 1))
assert(W2.shape == (n_y, n_h))
assert(b2.shape == (n_y, 1))
parameters = {"W1": W1,
"b1": b1,
"W2": W2,
"b2": b2}
return parameters
parameters = initialize_parameters(3,2,1)
print("W1 = " + str(parameters["W1"]))
print("b1 = " + str(parameters["b1"]))
print("W2 = " + str(parameters["W2"]))
print("b2 = " + str(parameters["b2"]))
```
**Expected output**:
<table style="width:80%">
<tr>
<td> **W1** </td>
<td> [[ 0.01624345 -0.00611756 -0.00528172]
[-0.01072969 0.00865408 -0.02301539]] </td>
</tr>
<tr>
<td> **b1**</td>
<td>[[ 0.]
[ 0.]]</td>
</tr>
<tr>
<td>**W2**</td>
<td> [[ 0.01744812 -0.00761207]]</td>
</tr>
<tr>
<td> **b2** </td>
<td> [[ 0.]] </td>
</tr>
</table>
### 3.2 - L-layer Neural Network
The initialization for a deeper L-layer neural network is more complicated because there are many more weight matrices and bias vectors. When completing the `initialize_parameters_deep`, you should make sure that your dimensions match between each layer. Recall that $n^{[l]}$ is the number of units in layer $l$. Thus for example if the size of our input $X$ is $(12288, 209)$ (with $m=209$ examples) then:
<table style="width:100%">
<tr>
<td> </td>
<td> **Shape of W** </td>
<td> **Shape of b** </td>
<td> **Activation** </td>
<td> **Shape of Activation** </td>
<tr>
<tr>
<td> **Layer 1** </td>
<td> $(n^{[1]},12288)$ </td>
<td> $(n^{[1]},1)$ </td>
<td> $Z^{[1]} = W^{[1]} X + b^{[1]} $ </td>
<td> $(n^{[1]},209)$ </td>
<tr>
<tr>
<td> **Layer 2** </td>
<td> $(n^{[2]}, n^{[1]})$ </td>
<td> $(n^{[2]},1)$ </td>
<td>$Z^{[2]} = W^{[2]} A^{[1]} + b^{[2]}$ </td>
<td> $(n^{[2]}, 209)$ </td>
<tr>
<tr>
<td> $\vdots$ </td>
<td> $\vdots$ </td>
<td> $\vdots$ </td>
<td> $\vdots$</td>
<td> $\vdots$ </td>
<tr>
<tr>
<td> **Layer L-1** </td>
<td> $(n^{[L-1]}, n^{[L-2]})$ </td>
<td> $(n^{[L-1]}, 1)$ </td>
<td>$Z^{[L-1]} = W^{[L-1]} A^{[L-2]} + b^{[L-1]}$ </td>
<td> $(n^{[L-1]}, 209)$ </td>
<tr>
<tr>
<td> **Layer L** </td>
<td> $(n^{[L]}, n^{[L-1]})$ </td>
<td> $(n^{[L]}, 1)$ </td>
<td> $Z^{[L]} = W^{[L]} A^{[L-1]} + b^{[L]}$</td>
<td> $(n^{[L]}, 209)$ </td>
<tr>
</table>
Remember that when we compute $W X + b$ in python, it carries out broadcasting. For example, if:
$$ W = \begin{bmatrix}
j & k & l\\
m & n & o \\
p & q & r
\end{bmatrix}\;\;\; X = \begin{bmatrix}
a & b & c\\
d & e & f \\
g & h & i
\end{bmatrix} \;\;\; b =\begin{bmatrix}
s \\
t \\
u
\end{bmatrix}\tag{2}$$
Then $WX + b$ will be:
$$ WX + b = \begin{bmatrix}
(ja + kd + lg) + s & (jb + ke + lh) + s & (jc + kf + li)+ s\\
(ma + nd + og) + t & (mb + ne + oh) + t & (mc + nf + oi) + t\\
(pa + qd + rg) + u & (pb + qe + rh) + u & (pc + qf + ri)+ u
\end{bmatrix}\tag{3} $$
**Exercise**: Implement initialization for an L-layer Neural Network.
**Instructions**:
- The model's structure is *[LINEAR -> RELU] $ \times$ (L-1) -> LINEAR -> SIGMOID*. I.e., it has $L-1$ layers using a ReLU activation function followed by an output layer with a sigmoid activation function.
- Use random initialization for the weight matrices. Use `np.random.rand(shape) * 0.01`.
- Use zeros initialization for the biases. Use `np.zeros(shape)`.
- We will store $n^{[l]}$, the number of units in different layers, in a variable `layer_dims`. For example, the `layer_dims` for the "Planar Data classification model" from last week would have been [2,4,1]: There were two inputs, one hidden layer with 4 hidden units, and an output layer with 1 output unit. Thus means `W1`'s shape was (4,2), `b1` was (4,1), `W2` was (1,4) and `b2` was (1,1). Now you will generalize this to $L$ layers!
- Here is the implementation for $L=1$ (one layer neural network). It should inspire you to implement the general case (L-layer neural network).
```python
if L == 1:
parameters["W" + str(L)] = np.random.randn(layer_dims[1], layer_dims[0]) * 0.01
parameters["b" + str(L)] = np.zeros((layer_dims[1], 1))
```
```
# GRADED FUNCTION: initialize_parameters_deep
def initialize_parameters_deep(layer_dims):
"""
Arguments:
layer_dims -- python array (list) containing the dimensions of each layer in our network
Returns:
parameters -- python dictionary containing your parameters "W1", "b1", ..., "WL", "bL":
Wl -- weight matrix of shape (layer_dims[l], layer_dims[l-1])
bl -- bias vector of shape (layer_dims[l], 1)
"""
np.random.seed(3)
parameters = {}
L = len(layer_dims) # number of layers in the network
for l in range(1, L):
### START CODE HERE ### (≈ 2 lines of code)
parameters['W' + str(l)] = None
parameters['b' + str(l)] = None
### END CODE HERE ###
assert(parameters['W' + str(l)].shape == (layer_dims[l], layer_dims[l-1]))
assert(parameters['b' + str(l)].shape == (layer_dims[l], 1))
return parameters
parameters = initialize_parameters_deep([5,4,3])
print("W1 = " + str(parameters["W1"]))
print("b1 = " + str(parameters["b1"]))
print("W2 = " + str(parameters["W2"]))
print("b2 = " + str(parameters["b2"]))
```
**Expected output**:
<table style="width:80%">
<tr>
<td> **W1** </td>
<td>[[ 0.01788628 0.0043651 0.00096497 -0.01863493 -0.00277388]
[-0.00354759 -0.00082741 -0.00627001 -0.00043818 -0.00477218]
[-0.01313865 0.00884622 0.00881318 0.01709573 0.00050034]
[-0.00404677 -0.0054536 -0.01546477 0.00982367 -0.01101068]]</td>
</tr>
<tr>
<td>**b1** </td>
<td>[[ 0.]
[ 0.]
[ 0.]
[ 0.]]</td>
</tr>
<tr>
<td>**W2** </td>
<td>[[-0.01185047 -0.0020565 0.01486148 0.00236716]
[-0.01023785 -0.00712993 0.00625245 -0.00160513]
[-0.00768836 -0.00230031 0.00745056 0.01976111]]</td>
</tr>
<tr>
<td>**b2** </td>
<td>[[ 0.]
[ 0.]
[ 0.]]</td>
</tr>
</table>
## 4 - Forward propagation module
### 4.1 - Linear Forward
Now that you have initialized your parameters, you will do the forward propagation module. You will start by implementing some basic functions that you will use later when implementing the model. You will complete three functions in this order:
- LINEAR
- LINEAR -> ACTIVATION where ACTIVATION will be either ReLU or Sigmoid.
- [LINEAR -> RELU] $\times$ (L-1) -> LINEAR -> SIGMOID (whole model)
The linear forward module (vectorized over all the examples) computes the following equations:
$$Z^{[l]} = W^{[l]}A^{[l-1]} +b^{[l]}\tag{4}$$
where $A^{[0]} = X$.
**Exercise**: Build the linear part of forward propagation.
**Reminder**:
The mathematical representation of this unit is $Z^{[l]} = W^{[l]}A^{[l-1]} +b^{[l]}$. You may also find `np.dot()` useful. If your dimensions don't match, printing `W.shape` may help.
```
# GRADED FUNCTION: linear_forward
def linear_forward(A, W, b):
"""
Implement the linear part of a layer's forward propagation.
Arguments:
A -- activations from previous layer (or input data): (size of previous layer, number of examples)
W -- weights matrix: numpy array of shape (size of current layer, size of previous layer)
b -- bias vector, numpy array of shape (size of the current layer, 1)
Returns:
Z -- the input of the activation function, also called pre-activation parameter
cache -- a python dictionary containing "A", "W" and "b" ; stored for computing the backward pass efficiently
"""
### START CODE HERE ### (≈ 1 line of code)
Z = None
### END CODE HERE ###
assert(Z.shape == (W.shape[0], A.shape[1]))
cache = (A, W, b)
return Z, cache
A, W, b = linear_forward_test_case()
Z, linear_cache = linear_forward(A, W, b)
print("Z = " + str(Z))
```
**Expected output**:
<table style="width:35%">
<tr>
<td> **Z** </td>
<td> [[ 3.26295337 -1.23429987]] </td>
</tr>
</table>
### 4.2 - Linear-Activation Forward
In this notebook, you will use two activation functions:
- **Sigmoid**: $\sigma(Z) = \sigma(W A + b) = \frac{1}{ 1 + e^{-(W A + b)}}$. We have provided you with the `sigmoid` function. This function returns **two** items: the activation value "`a`" and a "`cache`" that contains "`Z`" (it's what we will feed in to the corresponding backward function). To use it you could just call:
``` python
A, activation_cache = sigmoid(Z)
```
- **ReLU**: The mathematical formula for ReLu is $A = RELU(Z) = max(0, Z)$. We have provided you with the `relu` function. This function returns **two** items: the activation value "`A`" and a "`cache`" that contains "`Z`" (it's what we will feed in to the corresponding backward function). To use it you could just call:
``` python
A, activation_cache = relu(Z)
```
For more convenience, you are going to group two functions (Linear and Activation) into one function (LINEAR->ACTIVATION). Hence, you will implement a function that does the LINEAR forward step followed by an ACTIVATION forward step.
**Exercise**: Implement the forward propagation of the *LINEAR->ACTIVATION* layer. Mathematical relation is: $A^{[l]} = g(Z^{[l]}) = g(W^{[l]}A^{[l-1]} +b^{[l]})$ where the activation "g" can be sigmoid() or relu(). Use linear_forward() and the correct activation function.
```
# GRADED FUNCTION: linear_activation_forward
def linear_activation_forward(A_prev, W, b, activation):
"""
Implement the forward propagation for the LINEAR->ACTIVATION layer
Arguments:
A_prev -- activations from previous layer (or input data): (size of previous layer, number of examples)
W -- weights matrix: numpy array of shape (size of current layer, size of previous layer)
b -- bias vector, numpy array of shape (size of the current layer, 1)
activation -- the activation to be used in this layer, stored as a text string: "sigmoid" or "relu"
Returns:
A -- the output of the activation function, also called the post-activation value
cache -- a python dictionary containing "linear_cache" and "activation_cache";
stored for computing the backward pass efficiently
"""
if activation == "sigmoid":
# Inputs: "A_prev, W, b". Outputs: "A, activation_cache".
### START CODE HERE ### (≈ 2 lines of code)
Z, linear_cache = None
A, activation_cache = None
### END CODE HERE ###
elif activation == "relu":
# Inputs: "A_prev, W, b". Outputs: "A, activation_cache".
### START CODE HERE ### (≈ 2 lines of code)
Z, linear_cache = None
A, activation_cache = None
### END CODE HERE ###
assert (A.shape == (W.shape[0], A_prev.shape[1]))
cache = (linear_cache, activation_cache)
return A, cache
A_prev, W, b = linear_activation_forward_test_case()
A, linear_activation_cache = linear_activation_forward(A_prev, W, b, activation = "sigmoid")
print("With sigmoid: A = " + str(A))
A, linear_activation_cache = linear_activation_forward(A_prev, W, b, activation = "relu")
print("With ReLU: A = " + str(A))
```
**Expected output**:
<table style="width:35%">
<tr>
<td> **With sigmoid: A ** </td>
<td > [[ 0.96890023 0.11013289]]</td>
</tr>
<tr>
<td> **With ReLU: A ** </td>
<td > [[ 3.43896131 0. ]]</td>
</tr>
</table>
**Note**: In deep learning, the "[LINEAR->ACTIVATION]" computation is counted as a single layer in the neural network, not two layers.
### d) L-Layer Model
For even more convenience when implementing the $L$-layer Neural Net, you will need a function that replicates the previous one (`linear_activation_forward` with RELU) $L-1$ times, then follows that with one `linear_activation_forward` with SIGMOID.
<img src="images/model_architecture_kiank.png" style="width:600px;height:300px;">
<caption><center> **Figure 2** : *[LINEAR -> RELU] $\times$ (L-1) -> LINEAR -> SIGMOID* model</center></caption><br>
**Exercise**: Implement the forward propagation of the above model.
**Instruction**: In the code below, the variable `AL` will denote $A^{[L]} = \sigma(Z^{[L]}) = \sigma(W^{[L]} A^{[L-1]} + b^{[L]})$. (This is sometimes also called `Yhat`, i.e., this is $\hat{Y}$.)
**Tips**:
- Use the functions you had previously written
- Use a for loop to replicate [LINEAR->RELU] (L-1) times
- Don't forget to keep track of the caches in the "caches" list. To add a new value `c` to a `list`, you can use `list.append(c)`.
```
# GRADED FUNCTION: L_model_forward
def L_model_forward(X, parameters):
"""
Implement forward propagation for the [LINEAR->RELU]*(L-1)->LINEAR->SIGMOID computation
Arguments:
X -- data, numpy array of shape (input size, number of examples)
parameters -- output of initialize_parameters_deep()
Returns:
AL -- last post-activation value
caches -- list of caches containing:
every cache of linear_relu_forward() (there are L-1 of them, indexed from 0 to L-2)
the cache of linear_sigmoid_forward() (there is one, indexed L-1)
"""
caches = []
A = X
L = len(parameters) // 2 # number of layers in the neural network
# Implement [LINEAR -> RELU]*(L-1). Add "cache" to the "caches" list.
for l in range(1, L):
A_prev = A
### START CODE HERE ### (≈ 2 lines of code)
A, cache = None
### END CODE HERE ###
# Implement LINEAR -> SIGMOID. Add "cache" to the "caches" list.
### START CODE HERE ### (≈ 2 lines of code)
AL, cache = None
### END CODE HERE ###
assert(AL.shape == (1,X.shape[1]))
return AL, caches
X, parameters = L_model_forward_test_case_2hidden()
AL, caches = L_model_forward(X, parameters)
print("AL = " + str(AL))
print("Length of caches list = " + str(len(caches)))
```
<table style="width:50%">
<tr>
<td> **AL** </td>
<td > [[ 0.03921668 0.70498921 0.19734387 0.04728177]]</td>
</tr>
<tr>
<td> **Length of caches list ** </td>
<td > 3 </td>
</tr>
</table>
Great! Now you have a full forward propagation that takes the input X and outputs a row vector $A^{[L]}$ containing your predictions. It also records all intermediate values in "caches". Using $A^{[L]}$, you can compute the cost of your predictions.
## 5 - Cost function
Now you will implement forward and backward propagation. You need to compute the cost, because you want to check if your model is actually learning.
**Exercise**: Compute the cross-entropy cost $J$, using the following formula: $$-\frac{1}{m} \sum\limits_{i = 1}^{m} (y^{(i)}\log\left(a^{[L] (i)}\right) + (1-y^{(i)})\log\left(1- a^{[L](i)}\right)) \tag{7}$$
```
# GRADED FUNCTION: compute_cost
def compute_cost(AL, Y):
"""
Implement the cost function defined by equation (7).
Arguments:
AL -- probability vector corresponding to your label predictions, shape (1, number of examples)
Y -- true "label" vector (for example: containing 0 if non-cat, 1 if cat), shape (1, number of examples)
Returns:
cost -- cross-entropy cost
"""
m = Y.shape[1]
# Compute loss from aL and y.
### START CODE HERE ### (≈ 1 lines of code)
cost = None
### END CODE HERE ###
cost = np.squeeze(cost) # To make sure your cost's shape is what we expect (e.g. this turns [[17]] into 17).
assert(cost.shape == ())
return cost
Y, AL = compute_cost_test_case()
print("cost = " + str(compute_cost(AL, Y)))
```
**Expected Output**:
<table>
<tr>
<td>**cost** </td>
<td> 0.41493159961539694</td>
</tr>
</table>
## 6 - Backward propagation module
Just like with forward propagation, you will implement helper functions for backpropagation. Remember that back propagation is used to calculate the gradient of the loss function with respect to the parameters.
**Reminder**:
<img src="images/backprop_kiank.png" style="width:650px;height:250px;">
<caption><center> **Figure 3** : Forward and Backward propagation for *LINEAR->RELU->LINEAR->SIGMOID* <br> *The purple blocks represent the forward propagation, and the red blocks represent the backward propagation.* </center></caption>
<!--
For those of you who are expert in calculus (you don't need to be to do this assignment), the chain rule of calculus can be used to derive the derivative of the loss $\mathcal{L}$ with respect to $z^{[1]}$ in a 2-layer network as follows:
$$\frac{d \mathcal{L}(a^{[2]},y)}{{dz^{[1]}}} = \frac{d\mathcal{L}(a^{[2]},y)}{{da^{[2]}}}\frac{{da^{[2]}}}{{dz^{[2]}}}\frac{{dz^{[2]}}}{{da^{[1]}}}\frac{{da^{[1]}}}{{dz^{[1]}}} \tag{8} $$
In order to calculate the gradient $dW^{[1]} = \frac{\partial L}{\partial W^{[1]}}$, you use the previous chain rule and you do $dW^{[1]} = dz^{[1]} \times \frac{\partial z^{[1]} }{\partial W^{[1]}}$. During the backpropagation, at each step you multiply your current gradient by the gradient corresponding to the specific layer to get the gradient you wanted.
Equivalently, in order to calculate the gradient $db^{[1]} = \frac{\partial L}{\partial b^{[1]}}$, you use the previous chain rule and you do $db^{[1]} = dz^{[1]} \times \frac{\partial z^{[1]} }{\partial b^{[1]}}$.
This is why we talk about **backpropagation**.
!-->
Now, similar to forward propagation, you are going to build the backward propagation in three steps:
- LINEAR backward
- LINEAR -> ACTIVATION backward where ACTIVATION computes the derivative of either the ReLU or sigmoid activation
- [LINEAR -> RELU] $\times$ (L-1) -> LINEAR -> SIGMOID backward (whole model)
### 6.1 - Linear backward
For layer $l$, the linear part is: $Z^{[l]} = W^{[l]} A^{[l-1]} + b^{[l]}$ (followed by an activation).
Suppose you have already calculated the derivative $dZ^{[l]} = \frac{\partial \mathcal{L} }{\partial Z^{[l]}}$. You want to get $(dW^{[l]}, db^{[l]} dA^{[l-1]})$.
<img src="images/linearback_kiank.png" style="width:250px;height:300px;">
<caption><center> **Figure 4** </center></caption>
The three outputs $(dW^{[l]}, db^{[l]}, dA^{[l]})$ are computed using the input $dZ^{[l]}$.Here are the formulas you need:
$$ dW^{[l]} = \frac{\partial \mathcal{L} }{\partial W^{[l]}} = \frac{1}{m} dZ^{[l]} A^{[l-1] T} \tag{8}$$
$$ db^{[l]} = \frac{\partial \mathcal{L} }{\partial b^{[l]}} = \frac{1}{m} \sum_{i = 1}^{m} dZ^{[l](i)}\tag{9}$$
$$ dA^{[l-1]} = \frac{\partial \mathcal{L} }{\partial A^{[l-1]}} = W^{[l] T} dZ^{[l]} \tag{10}$$
**Exercise**: Use the 3 formulas above to implement linear_backward().
```
# GRADED FUNCTION: linear_backward
def linear_backward(dZ, cache):
"""
Implement the linear portion of backward propagation for a single layer (layer l)
Arguments:
dZ -- Gradient of the cost with respect to the linear output (of current layer l)
cache -- tuple of values (A_prev, W, b) coming from the forward propagation in the current layer
Returns:
dA_prev -- Gradient of the cost with respect to the activation (of the previous layer l-1), same shape as A_prev
dW -- Gradient of the cost with respect to W (current layer l), same shape as W
db -- Gradient of the cost with respect to b (current layer l), same shape as b
"""
A_prev, W, b = cache
m = A_prev.shape[1]
### START CODE HERE ### (≈ 3 lines of code)
dW = None
db = None
dA_prev = None
### END CODE HERE ###
assert (dA_prev.shape == A_prev.shape)
assert (dW.shape == W.shape)
assert (db.shape == b.shape)
return dA_prev, dW, db
# Set up some test inputs
dZ, linear_cache = linear_backward_test_case()
dA_prev, dW, db = linear_backward(dZ, linear_cache)
print ("dA_prev = "+ str(dA_prev))
print ("dW = " + str(dW))
print ("db = " + str(db))
```
**Expected Output**:
<table style="width:90%">
<tr>
<td> **dA_prev** </td>
<td > [[ 0.51822968 -0.19517421]
[-0.40506361 0.15255393]
[ 2.37496825 -0.89445391]] </td>
</tr>
<tr>
<td> **dW** </td>
<td > [[-0.10076895 1.40685096 1.64992505]] </td>
</tr>
<tr>
<td> **db** </td>
<td> [[ 0.50629448]] </td>
</tr>
</table>
### 6.2 - Linear-Activation backward
Next, you will create a function that merges the two helper functions: **`linear_backward`** and the backward step for the activation **`linear_activation_backward`**.
To help you implement `linear_activation_backward`, we provided two backward functions:
- **`sigmoid_backward`**: Implements the backward propagation for SIGMOID unit. You can call it as follows:
```python
dZ = sigmoid_backward(dA, activation_cache)
```
- **`relu_backward`**: Implements the backward propagation for RELU unit. You can call it as follows:
```python
dZ = relu_backward(dA, activation_cache)
```
If $g(.)$ is the activation function,
`sigmoid_backward` and `relu_backward` compute $$dZ^{[l]} = dA^{[l]} * g'(Z^{[l]}) \tag{11}$$.
**Exercise**: Implement the backpropagation for the *LINEAR->ACTIVATION* layer.
```
# GRADED FUNCTION: linear_activation_backward
def linear_activation_backward(dA, cache, activation):
"""
Implement the backward propagation for the LINEAR->ACTIVATION layer.
Arguments:
dA -- post-activation gradient for current layer l
cache -- tuple of values (linear_cache, activation_cache) we store for computing backward propagation efficiently
activation -- the activation to be used in this layer, stored as a text string: "sigmoid" or "relu"
Returns:
dA_prev -- Gradient of the cost with respect to the activation (of the previous layer l-1), same shape as A_prev
dW -- Gradient of the cost with respect to W (current layer l), same shape as W
db -- Gradient of the cost with respect to b (current layer l), same shape as b
"""
linear_cache, activation_cache = cache
if activation == "relu":
### START CODE HERE ### (≈ 2 lines of code)
dZ = None
dA_prev, dW, db = None
### END CODE HERE ###
elif activation == "sigmoid":
### START CODE HERE ### (≈ 2 lines of code)
dZ = None
dA_prev, dW, db = None
### END CODE HERE ###
return dA_prev, dW, db
AL, linear_activation_cache = linear_activation_backward_test_case()
dA_prev, dW, db = linear_activation_backward(AL, linear_activation_cache, activation = "sigmoid")
print ("sigmoid:")
print ("dA_prev = "+ str(dA_prev))
print ("dW = " + str(dW))
print ("db = " + str(db) + "\n")
dA_prev, dW, db = linear_activation_backward(AL, linear_activation_cache, activation = "relu")
print ("relu:")
print ("dA_prev = "+ str(dA_prev))
print ("dW = " + str(dW))
print ("db = " + str(db))
```
**Expected output with sigmoid:**
<table style="width:100%">
<tr>
<td > dA_prev </td>
<td >[[ 0.11017994 0.01105339]
[ 0.09466817 0.00949723]
[-0.05743092 -0.00576154]] </td>
</tr>
<tr>
<td > dW </td>
<td > [[ 0.10266786 0.09778551 -0.01968084]] </td>
</tr>
<tr>
<td > db </td>
<td > [[-0.05729622]] </td>
</tr>
</table>
**Expected output with relu:**
<table style="width:100%">
<tr>
<td > dA_prev </td>
<td > [[ 0.44090989 0. ]
[ 0.37883606 0. ]
[-0.2298228 0. ]] </td>
</tr>
<tr>
<td > dW </td>
<td > [[ 0.44513824 0.37371418 -0.10478989]] </td>
</tr>
<tr>
<td > db </td>
<td > [[-0.20837892]] </td>
</tr>
</table>
### 6.3 - L-Model Backward
Now you will implement the backward function for the whole network. Recall that when you implemented the `L_model_forward` function, at each iteration, you stored a cache which contains (X,W,b, and z). In the back propagation module, you will use those variables to compute the gradients. Therefore, in the `L_model_backward` function, you will iterate through all the hidden layers backward, starting from layer $L$. On each step, you will use the cached values for layer $l$ to backpropagate through layer $l$. Figure 5 below shows the backward pass.
<img src="images/mn_backward.png" style="width:450px;height:300px;">
<caption><center> **Figure 5** : Backward pass </center></caption>
** Initializing backpropagation**:
To backpropagate through this network, we know that the output is,
$A^{[L]} = \sigma(Z^{[L]})$. Your code thus needs to compute `dAL` $= \frac{\partial \mathcal{L}}{\partial A^{[L]}}$.
To do so, use this formula (derived using calculus which you don't need in-depth knowledge of):
```python
dAL = - (np.divide(Y, AL) - np.divide(1 - Y, 1 - AL)) # derivative of cost with respect to AL
```
You can then use this post-activation gradient `dAL` to keep going backward. As seen in Figure 5, you can now feed in `dAL` into the LINEAR->SIGMOID backward function you implemented (which will use the cached values stored by the L_model_forward function). After that, you will have to use a `for` loop to iterate through all the other layers using the LINEAR->RELU backward function. You should store each dA, dW, and db in the grads dictionary. To do so, use this formula :
$$grads["dW" + str(l)] = dW^{[l]}\tag{15} $$
For example, for $l=3$ this would store $dW^{[l]}$ in `grads["dW3"]`.
**Exercise**: Implement backpropagation for the *[LINEAR->RELU] $\times$ (L-1) -> LINEAR -> SIGMOID* model.
```
# GRADED FUNCTION: L_model_backward
def L_model_backward(AL, Y, caches):
"""
Implement the backward propagation for the [LINEAR->RELU] * (L-1) -> LINEAR -> SIGMOID group
Arguments:
AL -- probability vector, output of the forward propagation (L_model_forward())
Y -- true "label" vector (containing 0 if non-cat, 1 if cat)
caches -- list of caches containing:
every cache of linear_activation_forward() with "relu" (it's caches[l], for l in range(L-1) i.e l = 0...L-2)
the cache of linear_activation_forward() with "sigmoid" (it's caches[L-1])
Returns:
grads -- A dictionary with the gradients
grads["dA" + str(l)] = ...
grads["dW" + str(l)] = ...
grads["db" + str(l)] = ...
"""
grads = {}
L = len(caches) # the number of layers
m = AL.shape[1]
Y = Y.reshape(AL.shape) # after this line, Y is the same shape as AL
# Initializing the backpropagation
### START CODE HERE ### (1 line of code)
dAL = None
### END CODE HERE ###
# Lth layer (SIGMOID -> LINEAR) gradients. Inputs: "AL, Y, caches". Outputs: "grads["dAL"], grads["dWL"], grads["dbL"]
### START CODE HERE ### (approx. 2 lines)
current_cache = None
grads["dA" + str(L)], grads["dW" + str(L)], grads["db" + str(L)] = None
### END CODE HERE ###
for l in reversed(range(L-1)):
# lth layer: (RELU -> LINEAR) gradients.
# Inputs: "grads["dA" + str(l + 2)], caches". Outputs: "grads["dA" + str(l + 1)] , grads["dW" + str(l + 1)] , grads["db" + str(l + 1)]
### START CODE HERE ### (approx. 5 lines)
current_cache = None
dA_prev_temp, dW_temp, db_temp = None
grads["dA" + str(l + 1)] = None
grads["dW" + str(l + 1)] = None
grads["db" + str(l + 1)] = None
### END CODE HERE ###
return grads
AL, Y_assess, caches = L_model_backward_test_case()
grads = L_model_backward(AL, Y_assess, caches)
print_grads(grads)
```
**Expected Output**
<table style="width:60%">
<tr>
<td > dW1 </td>
<td > [[ 0.41010002 0.07807203 0.13798444 0.10502167]
[ 0. 0. 0. 0. ]
[ 0.05283652 0.01005865 0.01777766 0.0135308 ]] </td>
</tr>
<tr>
<td > db1 </td>
<td > [[-0.22007063]
[ 0. ]
[-0.02835349]] </td>
</tr>
<tr>
<td > dA1 </td>
<td > [[ 0.12913162 -0.44014127]
[-0.14175655 0.48317296]
[ 0.01663708 -0.05670698]] </td>
</tr>
</table>
### 6.4 - Update Parameters
In this section you will update the parameters of the model, using gradient descent:
$$ W^{[l]} = W^{[l]} - \alpha \text{ } dW^{[l]} \tag{16}$$
$$ b^{[l]} = b^{[l]} - \alpha \text{ } db^{[l]} \tag{17}$$
where $\alpha$ is the learning rate. After computing the updated parameters, store them in the parameters dictionary.
**Exercise**: Implement `update_parameters()` to update your parameters using gradient descent.
**Instructions**:
Update parameters using gradient descent on every $W^{[l]}$ and $b^{[l]}$ for $l = 1, 2, ..., L$.
```
# GRADED FUNCTION: update_parameters
def update_parameters(parameters, grads, learning_rate):
"""
Update parameters using gradient descent
Arguments:
parameters -- python dictionary containing your parameters
grads -- python dictionary containing your gradients, output of L_model_backward
Returns:
parameters -- python dictionary containing your updated parameters
parameters["W" + str(l)] = ...
parameters["b" + str(l)] = ...
"""
L = len(parameters) // 2 # number of layers in the neural network
# Update rule for each parameter. Use a for loop.
### START CODE HERE ### (≈ 3 lines of code)
### END CODE HERE ###
return parameters
parameters, grads = update_parameters_test_case()
parameters = update_parameters(parameters, grads, 0.1)
print ("W1 = "+ str(parameters["W1"]))
print ("b1 = "+ str(parameters["b1"]))
print ("W2 = "+ str(parameters["W2"]))
print ("b2 = "+ str(parameters["b2"]))
```
**Expected Output**:
<table style="width:100%">
<tr>
<td > W1 </td>
<td > [[-0.59562069 -0.09991781 -2.14584584 1.82662008]
[-1.76569676 -0.80627147 0.51115557 -1.18258802]
[-1.0535704 -0.86128581 0.68284052 2.20374577]] </td>
</tr>
<tr>
<td > b1 </td>
<td > [[-0.04659241]
[-1.28888275]
[ 0.53405496]] </td>
</tr>
<tr>
<td > W2 </td>
<td > [[-0.55569196 0.0354055 1.32964895]]</td>
</tr>
<tr>
<td > b2 </td>
<td > [[-0.84610769]] </td>
</tr>
</table>
## 7 - Conclusion
Congrats on implementing all the functions required for building a deep neural network!
We know it was a long assignment but going forward it will only get better. The next part of the assignment is easier.
In the next assignment you will put all these together to build two models:
- A two-layer neural network
- An L-layer neural network
You will in fact use these models to classify cat vs non-cat images!
| github_jupyter |
# Dask GLM
[`dask-glm`](https://github.com/dask/dask-glm) is a library for fitting generalized linear models on large datasets.
The heart of the project is the set of optimization routines that work on either NumPy or dask arrays.
See [these](https://mrocklin.github.com/blog/work/2017/03/22/dask-glm-1) [two](http://matthewrocklin.com/blog/work/2017/04/19/dask-glm-2) blogposts describing how dask-glm works internally.
This notebook is shows an example of the higher-level scikit-learn style API built on top of these optimization routines.
```
import os
import s3fs
import pandas as pd
import dask.array as da
import dask.dataframe as dd
from distributed import Client
from dask import persist, compute
from dask_glm.estimators import LogisticRegression
```
We'll setup a [`distributed.Client`](http://distributed.readthedocs.io/en/latest/api.html#distributed.client.Client) locally. In the real world you could connect to a cluster of dask-workers.
```
client = Client()
```
For demonstration, we'll use the perennial NYC taxi cab dataset.
Since I'm just running things on my laptop, we'll just grab the first month's worth of data.
```
if not os.path.exists('trip.csv'):
s3 = S3FileSystem(anon=True)
s3.get("dask-data/nyc-taxi/2015/yellow_tripdata_2015-01.csv", "trip.csv")
ddf = dd.read_csv("trip.csv")
ddf = ddf.repartition(npartitions=8)
```
I happen to know that some of the values in this dataset are suspect, so let's drop them.
Scikit-learn doesn't support filtering observations inside a pipeline ([yet](https://github.com/scikit-learn/scikit-learn/issues/3855)), so we'll do this before anything else.
```
# these filter out less than 1% of the observations
ddf = ddf[(ddf.trip_distance < 20) &
(ddf.fare_amount < 150)]
ddf = ddf.repartition(npartitions=8)
```
Now, we'll split our DataFrame into a train and test set, and select our feature matrix and target column (whether the passenger tipped).
```
df_train, df_test = ddf.random_split([0.8, 0.2], random_state=2)
columns = ['VendorID', 'passenger_count', 'trip_distance', 'payment_type', 'fare_amount']
X_train, y_train = df_train[columns], df_train['tip_amount'] > 0
X_test, y_test = df_test[columns], df_test['tip_amount'] > 0
X_train, y_train, X_test, y_test = persist(
X_train, y_train, X_test, y_test
)
X_train.head()
y_train.head()
print(f"{len(X_train):,d} observations")
```
With our training data in hand, we fit our logistic regression.
Nothing here should be surprising to those familiar with `scikit-learn`.
```
%%time
# this is a *dask-glm* LogisticRegresion, not scikit-learn
lm = LogisticRegression(fit_intercept=False)
lm.fit(X_train.values, y_train.values)
```
Again, following the lead of scikit-learn we can measure the performance of the estimator on the training dataset using the `.score` method.
For LogisticRegression this is the mean accuracy score (what percent of the predicted matched the actual).
```
%%time
lm.score(X_train.values, y_train.values).compute()
```
and on the test dataset:
```
%%time
lm.score(X_test.values, y_test.values).compute()
```
## Pipelines
The bulk of my time "doing data science" is data cleaning and pre-processing.
Actually fitting an estimator or making predictions is a relatively small proportion of the work.
You could manually do all your data-processing tasks as a sequence of function calls starting with the raw data.
Or, you could use [scikit-learn's `Pipeline`](http://scikit-learn.org/stable/modules/pipeline.html) to accomplish this and then some.
`Pipeline`s offer a few advantages over the manual solution.
First, your entire modeling process from raw data to final output is in a self-contained object. No more wondering "did I remember to scale this version of my model?" It's there in the `Pipeline` for you to check.
Second, `Pipeline`s combine well with scikit-learn's model selection utilties, specifically `GridSearchCV` and `RandomizedSearchCV`. You're able to search over hyperparameters of the pipeline stages, just like you would for an estimator.
Third, `Pipeline`s help prevent leaking information from your test and validation sets to your training set.
A common mistake is to compute some pre-processing statistic on the *entire* dataset (before you've train-test split) rather than just the training set. For example, you might normalize a column by the average of all the observations.
These types of errors can lead you overestimate the performance of your model on new observations.
Since dask-glm follows the scikit-learn API, we can reuse scikit-learn's `Pipeline` machinery, *with a few caveats.*
Many of the tranformers built into scikit-learn will validate their inputs. As part of this,
array-like things are cast to numpy arrays. Since dask-arrays are array-like they are converted
and things "work", but this might not be ideal when your dataset doesn't fit in memory.
Second, some things are just fundamentally hard to do on large datasets.
For example, naively dummy-encoding a dataset requires a full scan of the data to determine the set of unique values per categorical column.
When your dataset fits in memory, this isn't a huge deal. But when it's scattered across a cluster, this could become
a bottleneck.
If you know the set of possible values *ahead* of time, you can do much better.
You can encode the categorical columns as pandas `Categoricals`, and then convert with `get_dummies`, without having to do an expensive full-scan, just to compute the set of unique values.
We'll do that on the `VendorID` and `payment_type` columnms.
```
from sklearn.base import TransformerMixin, BaseEstimator
from sklearn.pipeline import make_pipeline
```
First let's write a little transformer to convert columns to `Categoricals`.
If you aren't familar with scikit-learn transformers, the basic idea is that the transformer must implement two methods: `.fit` and `.tranform`.
`.fit` is called during training.
It learns something about the data and records it on `self`.
Then `.transform` uses what's learned during `.fit` to transform the feature matrix somehow.
A `Pipeline` is simply a chain of transformers, each one is `fit` on some data, and passes the output of `.transform` onto the next step; the final output is an `Estimator`, like `LogisticRegression`.
```
class CategoricalEncoder(BaseEstimator, TransformerMixin):
"""Encode `categories` as pandas `Categorical`
Parameters
----------
categories : Dict[str, list]
Mapping from column name to list of possible values
"""
def __init__(self, categories):
self.categories = categories
def fit(self, X, y=None):
# "stateless" transformer. Don't have anything to learn here
return self
def transform(self, X, y=None):
X = X.copy()
for column, categories in self.categories.items():
X[column] = X[column].astype('category').cat.set_categories(categories)
return X
```
We'll also want a daskified version of scikit-learn's `StandardScaler`, that won't eagerly
convert a dask.array to a numpy array (N.B. the scikit-learn version has more features and error handling, but this will work for now).
```
class StandardScaler(BaseEstimator, TransformerMixin):
def __init__(self, columns=None, with_mean=True, with_std=True):
self.columns = columns
self.with_mean = with_mean
self.with_std = with_std
def fit(self, X, y=None):
if self.columns is None:
self.columns_ = X.columns
else:
self.columns_ = self.columns
if self.with_mean:
self.mean_ = X[self.columns_].mean(0)
if self.with_std:
self.scale_ = X[self.columns_].std(0)
return self
def transform(self, X, y=None):
X = X.copy()
if self.with_mean:
X[self.columns_] = X[self.columns_] - self.mean_
if self.with_std:
X[self.columns_] = X[self.columns_] / self.scale_
return X.values
```
Finally, I've written a dummy encoder transformer that converts categoricals
to dummy-encoded interger columns. The full implementation is a bit long for a blog post, but you can see it [here](https://github.com/TomAugspurger/sktransformers/blob/master/sktransformers/preprocessing.py#L77).
```
from dummy_encoder import DummyEncoder
pipe = make_pipeline(
CategoricalEncoder({"VendorID": [1, 2],
"payment_type": [1, 2, 3, 4, 5]}),
DummyEncoder(),
StandardScaler(columns=['passenger_count', 'trip_distance', 'fare_amount']),
LogisticRegression(fit_intercept=False)
)
```
So that's our pipeline.
We can go ahead and fit it just like before, passing in the raw data.
```
%%time
pipe.fit(X_train, y_train.values)
```
And we can score it as well. The `Pipeline` ensures that all of the nescessary transformations take place before calling the estimator's `score` method.
```
pipe.score(X_train, y_train.values).compute()
pipe.score(X_test, y_test.values).compute()
```
## Grid Search
As explained earlier, Pipelines and grid search go hand-in-hand.
Let's run a quick example with [dask-searchcv](http://dask-searchcv.readthedocs.io/en/latest/).
```
from sklearn.model_selection import GridSearchCV
import dask_searchcv as dcv
```
We'll search over two hyperparameters
1. Whether or not to standardize the variance of each column in `StandardScaler`
2. The strength of the regularization in `LogisticRegression`
This involves fitting many models, one for each combination of paramters.
dask-searchcv is smart enough to know that early stages in the pipeline (like the categorical and dummy encoding) are shared among all the combinations, and so only fits them once.
```
param_grid = {
'standardscaler__with_std': [True, False],
'logisticregression__lamduh': [.001, .01, .1, 1],
}
pipe = make_pipeline(
CategoricalEncoder({"VendorID": [1, 2],
"payment_type": [1, 2, 3, 4, 5]}),
DummyEncoder(),
StandardScaler(columns=['passenger_count', 'trip_distance', 'fare_amount']),
LogisticRegression(fit_intercept=False)
)
gs = dcv.GridSearchCV(pipe, param_grid)
%%time
gs.fit(X_train, y_train.values)
```
Now we have access to the usual attributes like `cv_results_` learned by the grid search object:
```
pd.DataFrame(gs.cv_results_)
```
And we can do our usual checks on model fit for the training set:
```
gs.score(X_train, y_train.values).compute()
```
And the test set:
```
gs.score(X_test, y_test.values).compute()
```
Hopefully your reaction to everything here is somewhere between a nodding head and a yawn.
If you're familiar with scikit-learn, everything here should look pretty routine.
It's the same API you know and love, scaled out to larger datasets thanks to dask-glm.
| github_jupyter |
# Introduction
In this tutorial, we will learn how to implement different policies and temporal difference learning algorithms, as well as a hybrid (model building) algorithm, then compare their performance to the dynamic programming algorithms we implemented in the first tutorial.
# Notebook setup
## Instructions
- Import numpy, scipy and matplotlib
- Configure inline plots
```
% matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
from pylab import *
```
# Tutorial
## Learning algorithms and policies
__Learning algorithms__:
*Sarsa (on-policy)*
\begin{align}
Q(s_t,a_t) \leftarrow Q(s_t,a_t) + \alpha \big(r_t + \gamma Q(s_{t+1},a_{t+1}) - Q(s_t,a_t)\big)
\end{align}
with temporal discount rate $\gamma$ and learning rate $\alpha$.
*Q-learning (off-policy)*
\begin{align}
Q(s_t,a_t) \leftarrow Q(s_t,a_t) + \alpha \big(r_t + \gamma\max_\limits{a} Q(s_{t+1},a_{t+1}) - Q(s_t,a_t)\big)
\end{align}
__Policies__:
*Epsilon-greedy*
\begin{align}
P(a_t|s_t) = \epsilon \frac{1}{N_a} + (1-\epsilon)1[a_t =\max_\limits{a}Q(a_t,s_t)]
\end{align}
*Softmax*
\begin{align}
P(a_t|s_t) = \frac{\exp(Q(a_t,s_t)/\tau)}{\Sigma_{i=1}^n \exp(Q(i)/\tau)}
\end{align}
Please familiarize yourself with the code below, as it will help your write your own code to solve the exercises.
```
# Import definitions of the environments.
import RL_worlds as worlds
# Import helper functions for plotting.
from plot_util import *
def init_state(params):
"""
Initialize the state at the beginning of an episode.
Args:
params: a dictionary containing the default parameters.
Returns:
an integer corresponding to the initial state.
"""
if params['environment'].name == 'windy_cliff_grid':
return 0
elif params['environment'].name == 'n_armed_bandit':
return 0
elif params['environment'].name == 'cheese_world':
return 0
elif params['environment'].name == 'cliff_world':
return 0
elif params['environment'].name == 'quentins_world':
return 54
def update_state(state, action, params):
"""
State transition based on world, action and current state.
Args:
state: integer corresponding to the current state.
action: integer corresponding to the action taken.
params: a dictionary containing the default parameters.
Returns:
an integer corresponding to the next state;
an integer corresponding to the reward received.
"""
next_state, reward = params['environment'].get_outcome(state, action)
return next_state, reward
def call_policy(state, value, params):
"""
Call a policy to choose actions, given current state and value function.
Args:
state: integer corresponding to the current state.
value: a matrix indexed by state and action.
params: a dictionary containing the default parameters.
Returns:
an integer corresponding action chosen according to the policy.
"""
# multiple options for policy
if params['policy'] == 'epsilon_greedy':
return epsilon_greedy(state, value, params)
elif params['policy'] == 'softmax':
return softmax(state, value, params)
else: # random policy (if policy not recognized, choose randomly)
return randint(params['environment'].n_actions)
def update_value(prev_state, action, reward, state, value, params):
"""
Update the value function.
Args:
prev_state: an integer corresponding to the previous state.
action: an integer correspoding to action taken.
reward: a float corresponding to the reward received.
state: an integer corresponding to the current state;
should be None if the episode ended.
value: a matrix indexed by state and action.
params: a dictionary containing the default parameters.
Returns:
the updated value function (matrix indexed by state and action).
"""
if params['learning_rule'] == 'q_learning':
# off policy learning
return q_learning(prev_state, action, reward, state, value, params)
elif params['learning_rule'] == 'sarsa':
# on policy learning
return sarsa(prev_state, action, reward, state, value, params)
else:
print('Learning rule not recognized')
def default_params(environment):
"""
Define the default parameters.
Args:
environment: an object corresponding to the environment.
Returns:
a dictionary containing the default parameters, where the keys
are strings (parameter names).
"""
params = dict()
params['environment'] = environment
params['alpha'] = 0.1 # learning rate
params['beta'] = 10 # inverse temperature
params['policy'] = 'epsilon_greedy'
params['epsilon'] = 0.05 # epsilon-greedy policy
params['learning_rule'] = 'q_learning'
params['epsilon_decay'] = 0.9
if environment.name == 'windy_cliff_grid':
params['gamma'] = 0.6 # temporal discount factor
elif environment.name == 'n_armed_bandit':
params['gamma'] = 0.9 # temporal discount factor
elif environment.name == 'cliff_world':
params['gamma'] = 1.0 # no discounting
elif environment.name == 'cheese_world':
params['gamma'] = 0.5 # temporal discount factor
elif environment.name == 'quentins_world':
params['gamma'] = 0.9 # temporal discount factor
return params
```
## Exercise 1: Decision Policies
1. Write an epsilon-greedy policy function.
2. Write a softmax policy function.
Tip: Both functions should take the current state, the value function and default parameters as input and return an action.
```
def epsilon_greedy(state, value, params):
"""
Epsilon-greedy policy: selects the maximum value action with probabilty
(1-epsilon) and selects randomly with epsilon probability.
Args:
state: an integer corresponding to the current state.
value: a matrix indexed by state and action.
params: a dictionary containing the default parameters.
Returns:
action: an integer corresponding action chosen according to the policy.
"""
value_now = value[state,:]
if rand() > params['epsilon']:
action = where(value_now == max(value_now))[0][0]
else:
# We deal with ties by selecting randomly.
action = randint(len(value_now))
return action
def softmax(state, value, params):
"""
Softmax policy: selects action probabilistically depending on the value.
Args:
state: an integer corresponding to the current state.
value: a matrix indexed by state and action.
params: a dictionary containing the default parameters.
Returns:
an integer corresponding to the action chosen according to the policy.
"""
value_now = value[state,:]
prob = exp(value_now * params['beta']) # beta is the inverse temperature
prob = prob / sum(prob) # normalize
cum_prob = cumsum(prob) # cummulation summation
action = where(cum_prob > rand())[0][0]
return action
```
## Exercise 2: Learning Algorithms
1. Write a Q-learning (off-policy) algorithm.
2. Modify your Q-learning algorithm to obtain a Sarsa (on-policy) algorithm.
Tip: Both functions should take the previous state, action taken, reward received, value function, current state and default parameters and return the updated value function.
```
def q_learning(prev_state, action, reward, state, value, params):
"""
Q-learning: updates the value function and returns it.
Args:
prev_state: an integer corresponding to the previous state.
action: an integer corresponding to the action taken.
reward: a float corresponding to the reward received.
state: an integer corresponding to the current state.
value: a matrix indexed by state and action.
params: a dictionary containing the default parameters.
Returns:
the updated value function (matrix indexed by state and action).
"""
# maximum value at current state
if state == None:
max_value = 0
else:
max_value = max(value[state,:])
# value of previous state-action pair
prev_value = value[prev_state, action]
# reward prediction error
delta = reward + params['gamma'] * max_value - prev_value # gamma is the temporal discount factor
# update value of previous state-action pair
value[prev_state, action] = prev_value + params['alpha'] * delta # alpha: learning rate
return value
def sarsa(prev_state, action, reward, state, value, params):
"""
Sarsa: updates the value function and returns it.
Args:
prev_state: an integer corresponding to the previous state.
action: an integer correspoding to action taken.
reward: a float corresponding to the reward received.
state: an integer corresponding to the current state.
value: a matrix indexed by state and action.
params: a dictionary containing the default parameters.
Returns:
the updated value function (matrix indexed by state and action).
"""
# select the expected value at current state based on our
# policy by sampling from it
if state == None:
policy_value = 0
else:
# sample action from the policy for the next state
policy_action = call_policy(state, value, params)
# get the value based on the action sampled from the policy
policy_value = value[state, policy_action]
# value of previous state-action pair
prev_value = value[prev_state, action]
# reward prediction error
delta = reward + params['gamma'] * policy_value - prev_value # gamma is the temporal discount factor
# update value of previous state-action pair
value[prev_state, action] = prev_value + params['alpha'] * delta # alpha: learning rate
return value
```
## Exercise 3
1. Write code that allows you to select a world, a learning algorithm and a decision policy. Run 500 episodes (visits to the world) with learning across episodes. Make sure to set a maximum number of steps per episode (e.g. 1000). Use the functions provided in the plot_util module to:
- Plot the value associated with each action at each state;
- Plot the action corresponding to the maximum value at each state;
- Plot the maximum value in each state;
- Plot the total reward obtained in each episode.
2. Experiment with different values for the parameters:
- Pick a range for the learning rate $\alpha$ and look at how the results change.
- Pick a range for the inverse temperature $\beta$ (using a softmax policy) and look at how the results change.
- Pick a range for $\epsilon$ (using an $\epsilon$-greedy policy) and look at how the results change.
- Pick a range for the temporal discount factor $\gamma$ and look at how the results change.
3. Explore the cliff world with an $\epsilon$-greedy policy (try $\epsilon$=0.1) comparing the performance of Q-learning (off-policy) and Sarsa (on-policy). What differences do you notice? What do these differences tell us about on- and off-policy learning?
To make sure that your algorithms have been implemented correctly, compare your results to the ones shown below.
Cliff world using Q-learning and an $\epsilon$-greedy policy with $\epsilon$=0.1 and $\alpha$=0.3:
<img src="fig/tutorial1_ex3_qlearning_values.png",height="300",width="300",align="left">
<img src="fig/tutorial1_ex3_qlearning_actions.png",height="300",width="300">
<img src="fig/tutorial1_ex3_qlearning_maxval.png",height="300",width="300",align="left">
<img src="fig/tutorial1_ex3_qlearning_rewards.png",height="300",width="300">
Quentin's world using Sarsa and a softmax policy with $\beta$=10 and $\alpha$=0.4:
<img src="fig/tutorial1_ex3_sarsa_values.png",height="300",width="300",align="left">
<img src="fig/tutorial1_ex3_sarsa_actions.png",height="300",width="300">
<img src="fig/tutorial1_ex3_sarsa_maxval.png",height="300",width="300",align="left">
<img src="fig/tutorial1_ex3_sarsa_rewards.png",height="300",width="300">
```
def run_learning(value, params, n_episodes, max_steps):
"""
Args:
value: a matrix indexed by state and action.
params: a dictionary containing the default parameters.
n_episodes: integer, number of episodes to run.
max_steps: integer, maximum number of steps to take in each episode.
Returns:
a dictionary where the keys are integers (episode numbers)
and the values are integers (total rewards per episode);
the updated value function (matrix indexed by state and action).
"""
reward_sums = np.zeros(n_episodes)
# Loop over episodes
for episode in xrange(n_episodes):
state = init_state(params) # initialize state
step = 0
reward_sum = 0
# Make sure to break after max number of steps
while step < max_steps:
action = call_policy(state, value, params) # get action from policy
next_state, reward = update_state(state, action, params) # update state based on action
value = update_value(state, action, reward, next_state, value, params) # update value function
state = next_state
reward_sum += reward # sum rewards obtained
step += 1
if next_state == None:
break # episode ends
reward_sums[episode] = reward_sum
return reward_sums, value
# Choose a world
# env = worlds.n_armed_bandit()
# env = worlds.cheese_world()
# env = worlds.cliff_world()
# env = worlds.quentins_world()
env = worlds.windy_cliff_grid()
# Default parameters
params = default_params(environment=env)
# Decision-maker
# Choose a policy, a learning rule and parameter values
params['learning_rule'] = 'sarsa'
params['policy'] = 'epsilon_greedy'
params['epsilon'] = 0.01
params['alpha'] = 0.5
params['beta'] = 10
params['gamma'] = 0.8
# Episodes/trials
n_episodes = 1000
max_steps = 1000
# Initialization
# Start with uniform value function
value = np.ones((env.n_states, env.n_actions))
# Run learning
reward_sums, value = run_learning(value, params, n_episodes, max_steps)
fig = plot_state_action_values(env, value)
fig = plot_quiver_max_action(env, value)
fig = plot_heatmap_max_val(env, value)
fig = plot_rewards(n_episodes, reward_sums, average_range=10)
```
## Exercise 4: Dyna-Q
1. Implement the Dyna-Q algorithm for a deterministic environment.
Tip: The function should take the default parameters, an integer k for the number random updates, and a threshold for the stopping criterium as input and return a value function and deterministic model of the environment.
```
def dyna_q(value, T, R, params, k=100, n_iter=1000):
"""
Dyna-Q algorithm.
Args:
params: a dictionary containing the default parameters.
k: int, number of random updates per iteration.
n_iter: int, number of iterations.
Returns:
the value function (matrix indexed by state and action);
the transition matrix (matrix indexed by state and action);
the reward matrix (matrix indexed by state and action).
"""
env = params['environment']
observed_pairs = []
# Randomly sample an initial state
state = np.random.randint(env.n_states)
for _ in xrange(n_iter):
# Select an action according to the current value function
action = call_policy(state, value, params)
# Add state-action pair to the list of observed pairs
observed_pairs.append((state, action))
# Obtain next state and reward
next_state, reward = update_state(state, action, params)
# Value of previous state-action pair
prev_value = value[state, action]
# Maximum value at the new state
if next_state == None:
max_value = 0
else:
max_value = max(value[next_state,:])
# Reward prediction error
delta = reward + params['gamma'] * max_value - prev_value # gamma is the temporal discount factor
# Update the value function
value[state, action] = prev_value + params['alpha'] * delta # alpha is the learning rate
# Update the model
T[state,action] = next_it_state = next_state
R[state,action] = reward
# Perform k additional updates at random
for _ in xrange(k):
# Randomly sample a state-action pair
state, action = observed_pairs[np.random.randint(len(observed_pairs))]
# Obtain next state and reward according to the current model
next_state = T[state, action] if np.isfinite(T[state, action]) else None
reward = R[state, action]
# Value of previous state-action pair
prev_value = value[state, action]
# Maximum value at the new state
if next_state == None:
max_value = 0
else:
max_value = max(value[int(next_state),:])
# Reward prediction error
delta = reward + params['gamma'] * max_value - prev_value # gamma is the temporal discount factor
# Update the value function
value[state, action] = prev_value + params['alpha'] * delta # alpha is the learning rate
# Update the state for the next iteration
# If next state is None, we've reached the end of the MDP, so start over from random state
if next_it_state == None:
state = np.random.randint(env.n_states)
else:
state = next_it_state
return value, T, R
```
## Exercise 5
1. Write code that allows you to test the performance of Dyna-Q for a selected world and a selected learning rule. Run 500 episodes (visits to the world) with learning across episodes. Make sure to set a maximum number of steps per episode (e.g. 1000). Use the functions provided in the plot_util module to:
- Plot the value associated with each action at each state;
- Plot the action corresponding to the maximum value at each state;
- Plot the maximum value in each state;
- Plot the total reward obtained in each episode.
2. Experiment with different values for the parameters:
- Pick a range for the learning rate $\alpha$ and look at how the results change.
- Pick a range for the temporal discount factor $\gamma$ and look at how the results change.
- Pick a range for k (number of random updates performed in Dyna-Q) and look at how the results change.
3. Compare these results with those obtained for TD learning in the previous tutorial, as well as the dynamic programming algorithms from this tutorial.
To make sure that your algorithm has been implemented correctly, compare your results to the ones shown below.
Windy cliff grid using Dyna-Q and an $\epsilon$-greedy policy with $\epsilon$=0.05, $\alpha$=0.5 and $\gamma$=0.8:
<img src="fig/tutorial2_ex5_dynaq_values.png",height="300",width="300",align="left">
<img src="fig/tutorial2_ex5_dynaq_actions.png",height="300",width="300">
<img src="fig/tutorial2_ex5_dynaq_maxval.png",height="300",width="300",align="left">
<img src="fig/tutorial2_ex5_dynaq_rewards.png",height="300",width="300">
```
def run_dyna_q(value, params, n_episodes, max_steps):
"""
Args:
value: a matrix indexed by state and action.
params: a dictionary containing the default parameters.
n_episodes: integer, number of episodes to run.
max_steps: integer, maximum number of steps to take in each episode.
Returns:
a dictionary where the keys are integers (episode numbers)
and the values are integers (total rewards per episode);
the updated value function (matrix indexed by state and action).
"""
reward_sums = np.zeros(n_episodes)
T = np.zeros((env.n_states, env.n_actions))
R = np.zeros((env.n_states, env.n_actions))
# Loop over episodes
for episode in xrange(n_episodes):
state = init_state(params) # initialize state
step = 0
reward_sum = 0
value, T, R = dyna_q(value, T, R, params)
# Make sure to break after max number of steps
while step < max_steps:
action = call_policy(state, value, params) # get action from policy
next_state, reward = update_state(state, action, params) # update state based on action
reward_sum += reward # sum rewards obtained
if next_state is None:
break # episode ends
state = next_state
step += 1
reward_sums[episode] = reward_sum
return reward_sums, value
# Choose a world
# env = worlds.cliff_world()
# env = worlds.quentins_world()
env = worlds.windy_cliff_grid()
# Default parameters
params = default_params(environment=env)
# Decision-maker
# Choose a policy and parameter values
params['policy'] = 'epsilon_greedy'
params['epsilon'] = 0.05
params['alpha'] = 0.5
params['gamma'] = 0.8
params['beta'] = 3
# Episodes/trials
n_episodes = 500
max_steps = 1000
# Initialization
# Start with uniform value function
value = np.ones((env.n_states, env.n_actions))
# Run learning
reward_sums, value = run_dyna_q(value, params, n_episodes, max_steps)
fig = plot_state_action_values(env, value)
fig = plot_quiver_max_action(env, value)
fig = plot_heatmap_max_val(env, value)
fig = plot_rewards(n_episodes, reward_sums, average_range=10)
```
## Exercise 6 (Optional)
1. We will now compare Q-learning and Dyna-Q to see how each of these algorithms responds to a change in the structure of the environment. We will use a modified version of the windy cliff grid, windy_cliff_grid_2, which is the same as the original except for the location of the doors leading to the two windy rooms.
- For each algorithm, run 500 episodes in the original world. Then, switch to the modified world and run 500 more episodes.
- You can use an $\epsilon$-greedy policy with $\epsilon$=0.05 throughout.
- After both sets of episodes, plot the value of each action at each state, the action of maximum value at each state, the maximum value at each state and the total reward obtained in each episode (plot the accumulated rewards only for the 500 episodes in the modified world).
- What do you notice about the difference in each algorithm's performance once the environment has changed?
```
def update_value(prev_state, action, reward, state, value, params):
"""
Update the value function.
Args:
prev_state: an integer corresponding to the previous state.
action: an integer correspoding to action taken.
reward: a float corresponding to the reward received.
state: an integer corresponding to the current state;
should be None if the episode ended.
value: a matrix indexed by state and action.
params: a dictionary containing the default parameters.
Returns:
the updated value function (matrix indexed by state and action).
"""
if params['learning_rule'] == 'q_learning':
# off policy learning
return q_learning(prev_state, action, reward, state, value, params)
elif params['learning_rule'] == 'sarsa':
# on policy learning
return sarsa(prev_state, action, reward, state, value, params)
else:
print('Learning rule not recognized')
def q_learning(prev_state, action, reward, state, value, params):
"""
Q-learning: updates the value function and returns it.
Args:
prev_state: an integer corresponding to the previous state.
action: an integer corresponding to the action taken.
reward: a float corresponding to the reward received.
state: an integer corresponding to the current state.
value: a matrix indexed by state and action.
params: a dictionary containing the default parameters.
Returns:
the updated value function (matrix indexed by state and action).
"""
# maximum value at current state
if state == None:
max_value = 0
else:
max_value = max(value[state,:])
# value of previous state-action pair
prev_value = value[prev_state, action]
# reward prediction error
delta = reward + params['gamma'] * max_value - prev_value # gamma is the temporal discount factor
# update value of previous state-action pair
value[prev_state, action] = prev_value + params['alpha'] * delta # alpha: learning rate
return value
def run_learning(value, params, n_episodes, max_steps):
"""
Args:
value: a matrix indexed by state and action.
params: a dictionary containing the default parameters.
n_episodes: integer, number of episodes to run.
max_steps: integer, maximum number of steps to take in each episode.
Returns:
a dictionary where the keys are integers (episode numbers)
and the values are integers (total rewards per episode);
the updated value function (matrix indexed by state and action).
"""
reward_sums = np.zeros(n_episodes)
# Loop over episodes
for episode in xrange(n_episodes):
state = init_state(params) # initialize state
step = 0
reward_sum = 0
# Make sure to break after max number of steps
while step < max_steps:
action = call_policy(state, value, params) # get action from policy
next_state, reward = update_state(state, action, params) # update state based on action
value = update_value(state, action, reward, next_state, value, params) # update value function
state = next_state
reward_sum += reward # sum rewards obtained
step += 1
if next_state == None:
break # episode ends
reward_sums[episode] = reward_sum
return reward_sums, value
env = worlds.windy_cliff_grid()
# Default parameters
params = default_params(environment=env)
# Decision-maker
params['learning_rule'] = 'q_learning'
params['policy'] = 'epsilon_greedy'
params['epsilon'] = 0.05
params['alpha'] = 0.5
params['gamma'] = 0.8
# Episodes/trials
n_episodes = 500
max_steps = 1000
# Initialization
# Start with uniform value functions
value_q = np.ones((env.n_states, env.n_actions))
value_dyna = np.ones((env.n_states, env.n_actions))
# Run Q-learning
reward_sums, value_q = run_learning(value_q, params, n_episodes, max_steps)
# Run Dyna-Q
reward_sums, value_dyna = run_dyna_q(value_dyna, params, n_episodes, max_steps)
# Change the environment.
env = worlds.windy_cliff_grid_2()
params = default_params(environment=env)
# Run Q-learning
reward_sums, value_q = run_learning(value_q, params, n_episodes, max_steps)
fig = plot_state_action_values(env, value_q)
fig = plot_quiver_max_action(env, value_q)
fig = plot_heatmap_max_val(env, value_q)
fig = plot_rewards(n_episodes, reward_sums, average_range=10)
# Run Dyna-Q
reward_sums, value_dyna = run_dyna_q(value_dyna, params, n_episodes, max_steps)
fig = plot_state_action_values(env, value_dyna)
fig = plot_quiver_max_action(env, value_dyna)
fig = plot_heatmap_max_val(env, value_dyna)
fig = plot_rewards(n_episodes, reward_sums, average_range=10)
```
| github_jupyter |
```
# File Contains: Python code containing closed-form solutions for the valuation of European Options,
# American Options, Asian Options, Spread Options, Heat Rate Options, and Implied Volatility
#
# This document demonstrates a Python implementation of some option models described in books written by Davis
# Edwards: "Energy Trading and Investing", "Risk Management in Trading", "Energy Investing Demystified".
#
# for backward compatability with Python 2.7
from __future__ import division
# import necessary libaries
import unittest
import math
import numpy as np
from scipy.stats import norm
from scipy.stats import mvn
# Developer can toggle _DEBUG to True for more messages
# normally this is set to False
_DEBUG = False
```
MIT License
Copyright (c) 2017 Davis William Edwards
Permission is hereby granted, free of charge, to any person obtaining a copy
of this software and associated documentation files (the "Software"), to deal
in the Software without restriction, including without limitation the rights
to use, copy, modify, merge, publish, distribute, sublicense, and/or sell
copies of the Software, and to permit persons to whom the Software is
furnished to do so, subject to the following conditions:
The above copyright notice and this permission notice shall be included in all
copies or substantial portions of the Software.
THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE
AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM,
OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE
SOFTWARE.
# Closed Form Option Pricing Formulas
## Generalized Black Scholes (GBS) and similar models
** ChangeLog: **
* 1/1/2017 Davis Edwards, Created GBS and Asian option formulas
* 3/2/2017 Davis Edwards, added TeX formulas to describe calculations
* 4/9/2017 Davis Edwards, added spread option (Kirk's approximation)
* 5/10/2017 Davis Edwards, added graphics for sensitivity analysis
* 5/18/2017 Davis Edwards, added Bjerksund-Stensland (2002) approximation for American Options
* 5/19/2017 Davis Edwards, added implied volatility calculations
* 6/7/2017 Davis Edwards, expanded sensitivity tests for American option approximation.
* 6/21/2017 Davis Edwards, added documentation for Bjerksund-Stensland models
* 7/21/2017 Davis Edwards, refactored all of the functions to match the parameter order to Haug's "The Complete Guide to Option Pricing Formulas".
** TO DO List **
1. Since the Asian Option valuation uses an approximation, need to determine the range of acceptable stresses that can be applied to the volatility input
2. Sub-class the custom assertions in this module to work with "unittest"
3. Update the greek calculations for American Options - currently the Greeks are approximated by the greeks from GBS model.
4. Add a bibliography referencing the academic papers used as sources
5. Finish writing documentation formulas for Close-Form approximation for American Options
6. Refactor the order of parameters for the function calls to replicate the order of parameters in academic literature
-------------------------
## Purpose:
(Why these models exist)
The software in this model is intended to price particular types of financial products called "options". These are a common type of financial product and fall into the category of "financial derivative". This documentation assumes that the reader is already familiar with options terminology. The models are largely variations of the Black Scholes Merton option framework (collectively called "Black Scholes Genre" or "Generalized Black Scholes") that are used to price European options (options that can only be exercised at one point in time). This library also includes approximations to value American options (options that can be exercised prior to the expiration date) and implied volatility calculators.
Pricing Formulas
1. BlackScholes() Stock Options (no dividend yield)
2. Merton() Assets with continuous dividend yield (Index Options)
3. Black76() Commodity Options
4. GK() FX Options (Garman-Kohlhagen)
5. Asian76() Asian Options on Commodities
6. Kirks76() Spread Options (Kirk's Approximation)
7. American() American options
8. American76() American Commodity Options
Implied Volatility Formulas
9. EuroImpliedVol Implied volatility calculator for European options
10. EuroImpliedVol76 Implied volatiltity calculator for European commodity options
11. AmerImpliedVol Implied volatiltity calculator for American options
11. AmerImpliedVol76 Implied volatility calculator for American commodity options
Note:
In honor of the Black76 model, the 76() on the end of functions indicates a commodity option.
-------------------------
## Scope
(Where this model is to be used):
This model is built to price financial option contracts on a wide variety of financial commodities. These options are widely used and represent the benchmark to which other (more complicated) models are compared. While those more complicated models may outperform these models in specific areas, outperformance is relatively uncommon. By an large, these models have taken on all challengers and remain the de-facto industry standard.
## Theory:
### Generalized Black Scholes
Black Scholes genre option models widely used to value European options. The original “Black Scholes” model was published in 1973 for non-dividend paying stocks. This created a revolution in quantitative finance and opened up option trading to the general population. Since that time, a wide variety of extensions to the original Black Scholes model have been created. Collectively, these are referred to as "Black Scholes genre” option models. Modifications of the formula are used to price other financial instruments like dividend paying stocks, commodity futures, and FX forwards. Mathematically, these formulas are nearly identical. The primary difference between these models is whether the asset has a carrying cost (if the asset has a cost or benefit associated with holding it) and how the asset gets present valued. To illustrate this relationship, a “generalized” form of the Black Scholes equation is shown below.
The Black Scholes model is based on number of assumptions about how financial markets operate. Black Scholes style models assume:
1. **Arbitrage Free Markets**. Black Scholes formulas assume that traders try to maximize their personal profits and don’t allow arbitrage opportunities (riskless opportunities to make a profit) to persist.
2. **Frictionless, Continuous Markets**. This assumption of frictionless markets assumes that it is possible to buy and sell any amount of the underlying at any time without transaction costs.
3. **Risk Free Rates**. It is possible to borrow and lend money at a risk-free interest rate
4. **Log-normally Distributed Price Movements**. Prices are log-normally distributed and described by Geometric Brownian Motion
5. **Constant Volatility**. The Black Scholes genre options formulas assume that volatility is constant across the life of the option contract.
In practice, these assumptions are not particularly limiting. The primary limitation imposed by these models is that it is possible to (reasonably) describe the dispersion of prices at some point in the future in a mathematical equation.
In the traditional Black Scholes model intended to price stock options, the underlying assumption is that the stock is traded at its present value and that prices will follow a random walk diffusion style process over time. Prices are assumed to start at the spot price and, on the average, to drift upwards over time at the risk free rate. The “Merton” formula modifies the basic Black Scholes equation by introducing an additional term to incorporate dividends or holding costs. The Black 76 formula modifies the assumption so that the underlying starts at some forward price rather than a spot price. A fourth variation, the Garman Kohlhagen model, is used to value foreign exchange (FX) options. In the GK model, each currency in the currency pair is discounted based on its own interest rate.
1. **Black Scholes (Stocks)**. In the traditional Black Scholes model, the option is based on common stock - an instrument that is traded at its present value. The stock price does not get present valued – it starts at its present value (a ‘spot price’) and drifts upwards over time at the risk free rate.
2. **Merton (Stocks with continuous dividend yield)**. The Merton model is a variation of the Black Scholes model for assets that pay dividends to shareholders. Dividends reduce the value of the option because the option owner does not own the right to dividends until the option is exercised.
3. **Black 76 (Commodity Futures)**. The Black 76 model is for an option where the underlying commodity is traded based on a future price rather than a spot price. Instead of dealing with a spot price that drifts upwards at the risk free rate, this model deals with a forward price that needs to be present valued.
4. **Garman-Kohlhagen (FX Futures)**. The Garman Kohlhagen model is used to value foreign exchange (FX) options. In the GK model, each currency in the currency pair is discounted based on its own interest rate.
An important concept of Black Scholes models is that the actual way that the underlying asset drifts over time isn't important to the valuation. Since European options can only be exercised when the contract expires, it is only the distribution of possible prices on that date that matters - the path that the underlying took to that point doesn't affect the value of the option. This is why the primary limitation of the model is being able to describe the dispersion of prices at some point in the future, not that the dispersion process is simplistic.
The generalized Black-Scholes formula can found below (see *Figure 1 – Generalized Black Scholes Formula*). While these formulas may look complicated at first glance, most of the terms can be found as part of an options contract or are prices readily available in the market. The only term that is difficult to calculate is the implied volatility (σ). Implied volatility is typically calculated using prices of other options that have recently been traded.
>*Call Price*
>\begin{equation}
C = Fe^{(b-r)T} N(D_1) - Xe^{-rT} N(D_2)
\end{equation}
>*Put Price*
>\begin{equation}
P = Xe^{-rT} N(-D_2) - Fe^{(b-r)T} N(-D_1)
\end{equation}
>*with the following intermediate calculations*
>\begin{equation}
D_1 = \frac{ln\frac{F}{X} + (b+\frac{V^2}{2})T}{V*\sqrt{T}}
\end{equation}
>\begin{equation}
D_2 = D_1 - V\sqrt{T}
\end{equation}
>*and the following inputs*
>| Symbol | Meaning |
>|--------------|---------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------|
>| F or S | **Underlying Price**. The price of the underlying asset on the valuation date. S is used commonly used to represent a spot price, F a forward price |
>| X | **Strike Price**. The strike, or exercise, price of the option. |
>| T | **Time to expiration**. The time to expiration in years. This can be calculated by comparing the time between the expiration date and the valuation date. T = (t_1 - t_0)/365 |
>| t_0 | **Valuation Date**. The date on which the option is being valued. For example, it might be today’s date if the option we being valued today. |
>| t_1 | **Expiration Date**. The date on which the option must be exercised. |
>| V | **Volatility**. The volatility of the underlying security. This factor usually cannot be directly observed in the market. It is most often calculated by looking at the prices for recent option transactions and back-solving a Black Scholes style equation to find the volatility that would result in the observed price. This is commonly abbreviated with the greek letter sigma,σ, although V is used here for consistency with the code below. |
>| q | **Continuous Yield**. Used in the Merton model, this is the continuous yield of the underlying security. Option holders are typically not paid dividends or other payments until they exercise the option. As a result, this factor decreases the value of an option. |
>| r | **Risk Free Rate**. This is expected return on a risk-free investment. This is commonly a approximated by the yield on a low-risk government bond or the rate that large banks borrow between themselves (LIBOR). The rate depends on tenor of the cash flow. For example, a 10-year risk-free bond is likely to have a different rate than a 20-year risk-free bond.[DE1] |
>| rf | **Foreign Risk Free Rate**. Used in the Garman Kohlhagen model, this is the risk free rate of the foreign currency. Each currency will have a risk free rate. |
>*Figure 1 - Generalized Black Scholes Formula*
The correction term, b, varies by formula – it differentiates the various Black Scholes formula from one another (see *Figure 2 - GBS Cost of Carry Adjustments*). The cost of carry refers to the cost of “carrying” or holding a position. For example, holding a bond may result in earnings from interest, holding a stock may result in stock dividends, or the like. Those payments are made to the owner of the underlying asset and not the owner of the option. As a result, they reduce the value of the option.
>| | Model | Cost of Carry (b) |
>|----|------------------|-------------------|
>| 1. | BlackScholes | b = r |
>| 2. | Merton | b = r - q |
>| 3. | Black 1976 | b = 0 |
>| 4. | Garman Kohlhagen | b = r - rf |
>| 5. | Asian | b = 0, modified V |
>*Figure 2 - GBS Cost of Carry Adjustment*
### Asian Volatility Adjustment
An Asian option is an option whose payoff is calculated using the average price of the underlying over some period of time rather than the price on the expiration date. As a result, Asian options are also called average price options. The reason that traders use Asian options is that averaging a settlement price over a period of time reduces the affect of manipulation or unusual price movements on the expiration date on the value of the option. As a result, Asian options are often found on strategically important commodities, like crude oil or in markets with intermittent trading.
The average of a set of random numbers (prices in this case) will have a lower dispersion (a lower volatility) than the dispersion of prices observed on any single day. As a result, the implied volatility used to price Asian options will usually be slightly lower than the implied volatility on a comparable European option. From a mathematical perspective, valuing an Asian option is slightly complicated since the average of a set of lognormal distributions is not itself lognormally distributed. However, a reasonably good approximation of the correct answer is not too difficult to obtain.
In the case of Asian options on futures, it is possible to use a modified Black-76 formula that replaces the implied volatility term with an adjusted implied volatility of the average price. As long as the first day of the averaging period is in the future, the following formula can be used to value Asian options (see *Figure 3 – Asian Option Formula*).
>*Asian Adjusted Volatility*
\begin{equation}
V_a = \sqrt{\frac{ln(M)}{T}}
\end{equation}
>*with the intermediate calculation*
\begin{equation}
M = \frac{2e^{V^2T} - 2e^{V^2T}[1+V^2(T-t)]}{V^4(T-t)^2}
\end{equation}
>| Symbol | Meaning |
|--------|-----------------------------------------------------------------------------------------------------------------|
| Va | **Asian Adjusted Volatility**, This will replace the volatility (V) term in the GBS equations shown previously. |
| T | **Time to expiration**. The time to expiration of the option (measured in years). |
| t | **Time to start of averaging period**. The time to the start of the averaging period (measured in years). |
>*Figure 3 - Asian Option Formula*
### Spread Option (Kirk's Approximation) Calculation
Spread options are based on the spread between two commodity prices. They are commonly used to model physical investments as "real options" or to mark-to-market contracts that hedge physical assets. For example, a natural gas fueled electrical generation unit can be used to convert fuel (natural gas) into electricity. Whenever this conversion is profitable, it would be rational to operate the unit. This type of conversion is readily modeled by a spread option. When the spread of (electricity prices - fuel costs) is greater than the conversion cost, then the unit would operate. In this example, the conversion cost, which might be called the *Variable Operations and Maintenance* or VOM for a generation unit, would represent the strike price.
Analytic formulas similar to the Black Scholes equation are commonly used to value commodity spread options. One such formula is called *Kirk’s approximation*. While an exact closed form solution does not exist to value spread options, approximate solutions can give reasonably accurate results. Kirk’s approximation uses a Black Scholes style framework to analyze the joint distribution that results from the ratio of two log-normal distributions.
In a Black Scholes equation, the distribution of price returns is assumed to be normally distributed on the expiration date. Kirk’s approximation builds on the Black Scholes framework by taking advantage of the fact that the ratio of two log-normal distributions is approximately normally distributed. By modeling a ratio of two prices rather than the spread between the prices, Kirk’s approximation can use the same formulas designed for options based on a single underlying. In other words, Kirk’s approximation uses an algebraic transformation to fit the spread option into the Black Scholes framework.
The payoff of a spread option is show in *Figure 4 - Spread Option Payoff*.
>\begin{equation}
C = max[F_1 - F_2 - X, 0]
\end{equation}
>\begin{equation}
P = max[X - (F_1 - F_2), 0]
\end{equation}
>where
>| Symbol | Meaning |
|--------|----------------------------------------------------|
| F_1 | **Price of Asset 1**, The prices of the first asset. |
| F_2 | **Price of Asset 2**. The price of the second asset. |
>*Figure 4 - Spread Option Payoff*
This can be algebraically manipulated as shown in *Figure 5 - Spread Option Payoff, Manipulated*.
>\begin{equation}
C = max \biggl[\frac{F_1}{F_2+X}-1,0 \biggr](F_2 + X)
\end{equation}
>\begin{equation}
P = max \biggl[1-\frac{F_1}{F_2+X},0 \biggr](F_2 + X)
\end{equation}
>*Figure 5 - Spread Option Payoff, Manipulated*
This allows Kirk’s approximation to model the distribution of the spread as the ratio of the price of asset 1 over the price of asset 2 plus the strike price. This ratio can then be converted into a formula very similar to the Generalized Black Scholes formulas. In fact, this is the Black Scholes formula shown above with the addition of a (F_2 + X) term (See *Figure 6 – Kirk’s Approximation Ratio*).
>*Ratio of prices*
>\begin{equation}
F = \frac{F_1}{F_2 + X}
\end{equation}
>The ratio implies that the option is profitable to exercise (*in the money*) whenever the ratio of prices (F in the formula above) is greater than 1. This occurs the cost of the finished product (F_1) exceeds total cost of the raw materials (F_2) and the conversion cost (X). This requires a modification to the Call/Put Price formulas and to the D_1 formula. Because the option is in the money when F>1, the "strike" price used in inner square brackets of the Call/Put Price formulas and the D1 formula is set to 1.
>*Spread Option Call Price*
>\begin{equation}
C = (F_2 + X)\biggl[Fe^{(b-r)T} N(D_1) - e^{-rT} N(D_2)\biggr]
\end{equation}
>*Spread Option Put Price*
>\begin{equation}
P = (F_2 + X)\biggl[e^{-rT} N(-D_2) - Fe^{(b-r)T} N(-D_1)\biggr]
\end{equation}
>\begin{equation}
D_1 = \frac{ln(F) + (b+\frac{V^2}{2})T}{V*\sqrt{T}}
\end{equation}
>\begin{equation}
D_2 = D_1 - V\sqrt{T}
\end{equation}
>*Figure 6- Kirk's Approximation Ratio*
The key complexity is determining the appropriate volatility that needs to be used in the equation. The “approximation” which defines Kirk’s approximation is the assumption that the ratio of two log-normal distributions is normally distributed. That assumption makes it possible to estimate the volatility needed for the modified Black Scholes style equation. (See *Figure 7 - Kirk's Approximation (Volatility)*).
>\begin{equation}
V = \sqrt{ V_1^{2}+ \biggl[V_2\frac{F_2}{F_2+X}\biggr]^2 - 2ρ V_1 V_2 \frac{F_2}{F_2+X} }
\end{equation}
>| Symbol | Meaning |
>|--------------|---------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------|
>| V | **Volatility**. The Kirk's approximation volatility that will be placed into the formula shown in Figure 6 |
>| V1 | **Volatility of Asset 1**. The strike, or exercise, price of the option. |
>| V2 | **Volatility of Asset 2**. The volatility of the second asset |
>| ρ | **Correlation**. The correlation between price of asset 1 and the price of asset 2. |
>*Figure 7- Kirk's Approximation (Volatility)*
A second complexity is that the prices of two assets (F1 and F2) have to be in the same units. For example, in a heat rate option, the option represents the ability to convert fuel (natural gas) into electricity. The price of the first asset, electricity, might be quoted in US dollars per megawatt-hour or USD/MWH. However, the price of the second asset might be quoted in USD/MMBTU. To use the approximation, it is necessary to convert the price of the second asset into the units of the first asset (See *Example 1 - a Heat Rate Option*). This conversion rate will typically be specified as part of the contract.
>Example: A 10 MMBTU/MWH heat rate call option
>* F1 = price of electricity = USD 35/MWH
>* F2* = price of natural gas = USD 3.40/MMBTU; *This is not the price to plug into the model!*
>* V1 = volatility of electricity forward prices = 35%
>* V2 = volatility of natural gas forward price = 35%
>* Rho = correlation between electricity and natural gas forward prices = 90%
>* VOM = variable operation and maintenance cost (the conversion cost) = USD 3/MWH
>Before being placed into a spread option model, the price of natural gas would need to
>be converted into the correct units.
>* F2 = Heat Rate * Fuel Cost = (10 MMBTU/MWH)(USD 3.40/MMBTU) = USD 34/MWH
>The strike price would be set equal to the conversion cost
>* X = VOM costs = USD 3/MWH
> *Example 1 - a Heat Rate Call Option*
Another important consideration (not discussed in this write-up) is that volatility and correlation need to be matched to the tenor of the underlying assets. This means that it is necessary to measure the volatility of forward prices rather than spot prices. It may also be necessary to match the volatility and correlation to the correct month. For example, power prices in August may behave very differently than power prices in October or May in certain regions.
Like any model, spread options are subject to the "garbage in = garbage out" problem. However, the relative complexity of modeling commodity prices (the typical underlying for spread options) makes calibrating inputs a key part of the model.
### American Options
American options differ from European options because they can be exercised at any time. If there is a possibility that it will be more profitable to exercise the option than sell it, an American option will have more value than a corresponding European option. Early exercise typically occurs only when an option is *in the money*. If an option is out of the money, there is usually no reason to exercise early - it would be better to sell the option (in the case of a put option, to sell the option and the underlying asset).
The decision of whether to exercise early is primarily a question of interest rates and carrying costs. Carrying costs, or *cost of carry*, is a term that means an intermediate cash flows that is the result of holding an asset. For example, dividends on stocks are a postive cost of carry (owning the asset gives the owner a cash flow). A commodity might have a negative cost of carry. For example, a commodity that requires its owner to pay for storage would cause the owner of the physical commodity to pay cash to hold the asset. (**Note:** Commodity options are typically written on forwards or futures which have zero cost of carry instead of the actual underlying commodity). Cost of carry is cash flow that affects the owner of the underlying commodity and not the owner of the option. For example, when a stock pays a dividend, the owner of a call option does not receive the dividend - just the owner of the stock. For the perspective for the owner of a call option on a stock, the cost of carry will be the interest received from holding cash (r) less any dividends paid to owners of the stock (q).
Since an option has some value (the *extrinsic value*) that would be given up by exercising the option, exercising an option prior to maturity is a trade off between the option's extrinsic value (the remaining optionality) and the relative benefit of holding cash (time value of money) versus the benefit of holding the asset (carrying costs).
The early exercise feature of **American equity put options** may have value when:
* The cost of carry on the asset is low - preferably zero or negative.
* Interest rates are high
* The option is in the money
The early exercise feature of **American equity call options** may have value when:
* The cost of carry on the asset is positive
* Interest rates are low or negative
* The option is in the money
With commodities, things are a slightly different. There is typically no cost of carry since the underlying is a forward or a futures contract. It does not cost any money to enter an at-the-money commodity forward, swap, or futures contract. Also, these contracts don't have any intermediate cash flows. As a result, the primary benefit of early exercise is to get cash immediately (exercising an in-the-money option) rather than cash in the future. In high interest rate environements, the money recieved immediately from immediate execution may exceed the extrinsic value of the contract. This is due to strike price not being present valued in immediate execution (it is specified in the contract and a fixed number) but the payoff of a European option is discounted (forward price - strike price).
The overall result is that early exercise is fairly uncommon for most commodity options. Typically, it only occurs when interest rates are high. Generally, interest rates have to be higher than 15%-20% for American commodity options to differ substantially in value from European options with the same terms.
The early exercise feature of **American commodity options** has value when:
* Interest rates are high
* Volatility is low (this makes selling the option less of a good option)
* The option is in the money
There is no exact closed-form solution for American options. However, there are many approximations that are reasonably close to prices produced by open-form solutions (like binomial tree models). Two models are shown below, both created by Bjerksund and Stensland. The first was produced in 1993 and the second in 2002. The second model is a refinement of the first model, adding more complexity, in exchange for better accuracy.
#### Put-Call Parity
Because of Put/Call parity, it is possible to use a call valuation formula to calculate the value of a put option.
>\begin{equation}
P(S,X,T,r,b,V) = C(X,S,T,r-b,-b,V)
\end{equation}
or using the order of parameters used in this library:
>\begin{equation}
P(X,S,T,b,r,V) = C(S,X,T,-b,r-b,V)
\end{equation}
#### BjerksundStensland 1993 (BS1993)
There is no closed form solution for American options, and there are multiple people who have developed closed-form approximations to value American options. This is one such approximation. However, this approximation is no longer in active use by the public interface. It is primarily included as a secondary test on the BS2002 calculation. This function uses a numerical approximation to estimate the value of an American option. It does this by estimating a early exercise boundary and analytically estimating the probability of hitting that boundary. This uses the same inputs as a Generalized Black Scholes model:
FS = Forward or spot price (abbreviated FS in code, F in formulas below)
X = Strike Price
T = time to expiration
r = risk free rate
b = cost of carry
V = volatility
_Intermediate Calculations_. To be consistent with the Bjerksund Stensland paper, this write-up uses similar notation. Please note that both a capital B (B_0 and B_Infinity), a lower case b, and the greek symbol Beta are all being used. B_0 and B_infinity represent that optimal exercise boundaries in edge cases (for call options where T=0 and T=infinity respectively), lower case b is the cost of carry (passed in as an input), and Beta is an intermediate calculations.
>\begin{array}{lcl}
\beta & = & (0.5 - \frac{b}{V^2}) + \sqrt{(\frac{b}{V^2} - 0.5)^2 + 2 \frac{r}{V^2}} \\
B_\infty & = & \frac{\beta}{\beta-1} X \\
B_0 & = & max\biggl[X, (\frac{r}{r-b}) X\biggr] \\
h_1 & = & - b T + 2 V \sqrt{T} \frac{B_0}{B_\infty-B_0} \\
\end{array}
_Calculate the Early Exercise Boundary (i)_. The lower case i represents the early exercise boundary. Alpha is an intermediate calculation.
>\begin{array}{lcl}
i & = & B_0 + (B_\infty-B_0)(1 - e^{h_1} ) \\
\alpha & = & (i-X) i^{-\beta}
\end{array}
Check for immediate exercise_.
>\begin{equation}
if F >= i, then Value = F - X
\end{equation}
If no immediate exercise, approximate the early exercise price.
>\begin{eqnarray}
Value & = & \alpha * F^\beta \\
& - & \alpha * \phi(F,T,\beta,i,i,r,b,V) \\
& + & \phi(F,T,1,i,i,r,b,V) \\
& - & \phi(F,T,1,X,i,r,b,V) \\
& - & X * \phi(F,T,0,i,i,r,b,V) \\
& + & X * \phi(F,T,0,X,i,r,b,V)
\end{eqnarray}
_Compare to European Value_. Due to the approximation, it is sometime possible to get a value slightly smaller than the European value. If so, set the value equal to the European value estimated using Generalized Black Scholes.
>\begin{equation}
Value_{BS1993} = Max \biggl[ Value, Value_{GBS} \biggr]
\end{equation}
#### Bjerksund Stensland 2002 (BS2002)
source: https://www.researchgate.net/publication/228801918
FS = Forward or spot price (abbreviated FS in code, F in formulas below)
X = Strike Price
T = time to expiration
r = risk free rate
b = cost of carry
V = volatility
#### Psi
Psi is an intermediate calculation used by the Bjerksund Stensland 2002 approximation.
\begin{equation}
\psi(F, t_2, \gamma, H, I_2, I_1, t_1, r, b, V)
\end{equation}
_Intermediate calculations_.
The Psi function has a large number of intermediate calculations. For clarity, these are loosely organized into groups with each group used to simplify the next set of intermediate calculations.
>\begin{array}{lcl}
A_1 & = & V \ln(t_1) \\
A_2 & = & V \ln(t_2) \\
B_1 & = & \biggl[ b+(\gamma-0.5) V^2 \biggr] t_1 \\
B_2 & = & \biggl[ b+(\gamma-0.5) V^2 \biggr] t_2
\end{array}
More Intermediate calculations
>\begin{array}{lcl}
d_1 & = & \frac{ln(\frac{F}{I_1}) + B_1}{A_1} \\
d_2 & = & \frac{ln(\frac{I_2^2}{F I_1}) + B_1}{A_1} \\
d_3 & = & \frac{ln(\frac{F}{I_1}) - B_1}{A_1} \\
d_4 & = & \frac{ln(\frac{I_2^2}{F I_1}) - B_1}{A_1} \\
e_1 & = & \frac{ln(\frac{F}{H}) + B_2}{A_2} \\
e_2 & = & \frac{ln(\frac{I_2^2}{F H}) + B_2}{A_2} \\
e_3 & = & \frac{ln(\frac{I_1^2}{F H}) + B_2}{A_2} \\
e_4 & = & \frac{ln(\frac{F I_1^2}{H I_2^2}) + B_2}{A_2}
\end{array}
Even More Intermediate calculations
>\begin{array}{lcl}
\tau & = & \sqrt{\frac{t_1}{t_2}} \\
\lambda & = & -r+\gamma b+\frac{\gamma}{2} (\gamma-1) V^2 \\
\kappa & = & \frac{2b}{V^2} +(2 \gamma - 1)
\end{array}
_The calculation of Psi_.
This is the actual calculation of the Psi function. In the function below, M() represents the cumulative bivariate normal distribution (described a couple of paragraphs below this section). The abbreviation M() is used instead of CBND() in this section to make the equation a bit more readable and to match the naming convention used in Haug's book "The Complete Guide to Option Pricing Formulas".
>\begin{eqnarray}
\psi & = & e^{\lambda t_2} F^\gamma M(-d_1, -e_1, \tau) \\
& - & \frac{I_2}{F}^\kappa M(-d_2, -e_2, \tau) \\
& - & \frac{I_1}{F}^\kappa M(-d_3, -e_3, -\tau) \\
& + & \frac{I_1}{I_2}^\kappa M(-d_4, -e_4, -\tau))
\end{eqnarray}
#### Phi
Phi is an intermediate calculation used by both the Bjerksun Stensland 1993 and 2002 approximations. Many of the parameters are the same as the GBS model.
\begin{equation}
\phi(FS, T, \gamma, h, I, r, b, V)
\end{equation}
FS = Forward or spot price (abbreviated FS in code, F in formulas below).
T = time to expiration.
I = trigger price (as calculated in either BS1993 or BS2002 formulas
gamma = modifier to T, calculated in BS1993 or BS2002 formula
r = risk free rate.
b = cost of carry.
V = volatility.
Internally, the Phi() function is implemented as follows:
>\begin{equation}
d_1 = -\frac{ln(\frac{F}{h}) + \biggl[b+(\gamma-0.5) V^2 \biggr] T}{V \sqrt{T}}
\end{equation}
>\begin{equation}
d_2 = d_1 - 2 \frac{ln(I/F)}{V \sqrt(T)}
\end{equation}
>\begin{equation}
\lambda = -r+\gamma b+0.5 \gamma (\gamma-1) V^2
\end{equation}
>\begin{equation}
\kappa = \frac{2b}{V^2}+(2\gamma-1)
\end{equation}
>\begin{equation}
\phi = e^{\lambda T} F^{\gamma} \biggl[ N(d_1)-\frac{I}{F}^{\kappa} N(d_2) \biggr]
\end{equation}
##### Normal Cumulative Density Function (N)
This is the normal cumulative density function. It can be found described in a variety of statistical textbooks and/or wikipedia. It is part of the standard scipy.stats distribution and imported using the "from scipy.stats import norm" command.
Example:
\begin{equation}
N(d_1)
\end{equation}
#### Cumulative bivariate normal distribution (CBND)
The bivariate normal density function (BNDF) is given below (See *Figure 8 - Bivariate Normal Density Function (BNDF)*):
>\begin{equation}
BNDF(x, y) = \frac{1}{2 \pi \sqrt{1-p^2}} exp \biggl[-\frac{x^2-2pxy+y^2}{2(1-p^2)}\biggr]
\end{equation}
>*Figure 8. Bivariate Normal Density Function (BNDF)*
This can be integrated over x and y to calculate the joint probability that x < a and y < b. This is called the cumulative bivariate normal distribution (CBND) (See *Figure 9 - Cumulative Bivariate Normal Distribution (CBND))*:
>\begin{equation}
CBND(a, b, p) = \frac{1}{2 \pi \sqrt{1-p^2}} \int_{-\infty}^{a} \int_{-\infty}^{b} exp \biggl[-\frac{x^2-2pxy+y^2}{2(1-p^2)}\biggr] d_x d_y
\end{equation}
>*Figure 9. Cumulative Bivariate Normal Distribution (CBND)*
Where
* x = the first variable
* y = the second variable
* a = upper bound for first variable
* b = upper bound for second variable
* p = correlation between first and second variables
There is no closed-form solution for this equation. However, several approximations have been developed and are included in the numpy library distributed with Anaconda. The Genz 2004 model was chosen for implementation. Alternative models include those developed by Drezner and Wesolowsky (1990) and Drezner (1978). The Genz model improves these other model by going to an accuracy of 14 decimal points (from approximately 8 decimal points and 6 decimal points respectively).
-------------------------
## Limitations:
These functions have been tested for accuracy within an allowable range of inputs (see "Model Input" section below). However, general modeling advice applies to the use of the model. These models depend on a number of assumptions. In plain English, these models assume that the distribution of future prices can be described by variables like implied volatility. To get good results from the model, the model should only be used with reliable inputs.
The following limitations are also in effect:
1. The Asian Option approximation shouldn't be used for Asian options that are into the Asian option calculation period.
2. The American and American76 approximations break down when r < -20%. The limits are set wider in this example for testing purposes, but production code should probably limit interest rates to values between -20% and 100%. In practice, negative interest rates should be extremely rare.
3. No greeks are produced for spread options
4. These models assume a constant volatility term structure. This has no effect on European options. However, options that are likely to be exercise early (certain American options) and Asian options may be more affected.
-------------------------
## Model Inputs
This section describes the function calls an inputs needed to call this model:
These functions encapsulate the most commonly encountered option pricing formulas. These function primarily figure out the cost-of-carry term (b) and then call the generic version of the function. All of these functions return an array containg the premium and the greeks.
#### Public Functions in the Library
Pricing Formulas:
1. BlackScholes (OptionType, X, FS, T, r, V)
2. Merton (OptionType, X, FS, T, r, q, V)
3. Black76 (OptionType, X, FS, T, r, V)
4. GK (OptionType, X, FS, T, b, r, rf, V)
5. Asian76 (OptionType, X, FS, T, TA, r, V)
6. Kirks76
7. American (OptionType, X, FS, T, r, q, V)
8. American76 (OptionType, X, FS, T, r, V)
Implied Volatility Formulas
9. GBS_ImpliedVol(OptionType, X, FS, T, r, q, CP)
10. GBS_ImpliedVol76(OptionType, X, FS, T, r, CP)
11. American_ImpliedVol(OptionType, X, FS, T, r, q, CP)
11. American_ImpliedVol76(OptionType, X, FS, T, r, CP)
#### Inputs used by all models
| **Parameter** | **Description** |
|---------------|------------------------------------------------------------------------------------------------------------------------------------------------------|
| option_type | **Put/Call Indicator** Single character, "c" indicates a call; "p" a put|
| fs | **Price of Underlying** FS is generically used, but for specific models, the following abbreviations may be used: F = Forward Price, S = Spot Price) |
| x | **Strike Price** |
| t | **Time to Maturity** This is in years (1.0 = 1 year, 0.5 = six months, etc)|
| r | **Risk Free Interest Rate** Interest rates (0.10 = 10% interest rate |
| v | **Implied Volatility** Annualized implied volatility (1=100% annual volatility, 0.34 = 34% annual volatility|
#### Inputs used by some models
| **Parameter** | **Description** |
|---------------|------------------------------------------------------------------------------------------------------------------------------------------------------|
| b | **Cost of Carry** This is only found in internal implementations, but is identical to the cost of carry (b) term commonly found in academic option pricing literature|
| q | **Continuous Dividend** Used in Merton and American models; Internally, this is converted into cost of carry, b, with formula b = r-q |
| rf | **Foreign Interest Rate** Only used GK model; this functions similarly to q |
| t_a | **Asian Start** Used for Asian options; This is the time that starts the averaging period (TA=0 means that averaging starts immediately). As TA approaches T, the Asian value should become very close to the Black76 Value |
| cp | **Option Price** Used in the implied vol calculations; This is the price of the call or put observed in the market |
#### Outputs
All of the option pricing functions return an array. The first element of the array is the value of the option, the other elements are the greeks which measure the sensitivity of the option to changes in inputs. The greeks are used primarily for risk-management purposes.
| **Output** | **Description** |
|------------|-------------------------------------------------------------------------------------------------------------------|
| [0] | **Value** |
| [1] | **Delta** Sensitivity of Value to changes in price |
| [2] | **Gamma** Sensitivity of Delta to changes in price |
| [3] | **Theta** Sensitivity of Value to changes in time to expiration (annualized). To get a daily Theta, divide by 365 |
| [4] | **Vega** Sensitivity of Value to changes in Volatility |
| [5] | **Rho** Sensitivity of Value to changes in risk-free rates. |
The implied volatility functions return a single value (the implied volatility).
#### Acceptable Range for inputs
All of the inputs are bounded. While many of these functions will work with inputs outside of these bounds, they haven't been tested and are generally believed to be uncommon. The pricer will return an exception to the caller if an out-of-bounds input is used. If that was a valid input, the code below will need to be modified to allow wider inputs and the testing section updated to test that the models work under the widened inputs.
```
# This class contains the limits on inputs for GBS models
# It is not intended to be part of this module's public interface
class _GBS_Limits:
# An GBS model will return an error if an out-of-bound input is input
MAX32 = 2147483248.0
MIN_T = 1.0 / 1000.0 # requires some time left before expiration
MIN_X = 0.01
MIN_FS = 0.01
# Volatility smaller than 0.5% causes American Options calculations
# to fail (Number to large errors).
# GBS() should be OK with any positive number. Since vols less
# than 0.5% are expected to be extremely rare, and most likely bad inputs,
# _gbs() is assigned this limit too
MIN_V = 0.005
MAX_T = 100
MAX_X = MAX32
MAX_FS = MAX32
# Asian Option limits
# maximum TA is time to expiration for the option
MIN_TA = 0
# This model will work with higher values for b, r, and V. However, such values are extremely uncommon.
# To catch some common errors, interest rates and volatility is capped to 100%
# This reason for 1 (100%) is mostly to cause the library to throw an exceptions
# if a value like 15% is entered as 15 rather than 0.15)
MIN_b = -1
MIN_r = -1
MAX_b = 1
MAX_r = 1
MAX_V = 1
```
------------------------
## Model Implementation
These functions encapsulate a generic version of the pricing formulas. They are primarily intended to be called by the other functions within this libary. The following functions will have a fixed interface so that they can be called directly for academic applicaitons that use the cost-of-carry (b) notation:
_GBS() A generalized European option model
_American() A generalized American option model
_GBS_ImpliedVol() A generalized European option implied vol calculator
_American_ImpliedVol() A generalized American option implied vol calculator
The other functions in this libary are called by the four main functions and are not expected to be interface safe (the implementation and interface may change over time).
### Implementation: European Options
These functions implement the generalized Black Scholes (GBS) formula for European options. The main function is _gbs().
```
# ------------------------------
# This function verifies that the Call/Put indicator is correctly entered
def _test_option_type(option_type):
if (option_type != "c") and (option_type != "p"):
raise GBS_InputError("Invalid Input option_type ({0}). Acceptable value are: c, p".format(option_type))
# ------------------------------
# This function makes sure inputs are OK
# It throws an exception if there is a failure
def _gbs_test_inputs(option_type, fs, x, t, r, b, v):
# -----------
# Test inputs are reasonable
_test_option_type(option_type)
if (x < _GBS_Limits.MIN_X) or (x > _GBS_Limits.MAX_X):
raise GBS_InputError(
"Invalid Input Strike Price (X). Acceptable range for inputs is {1} to {2}".format(x, _GBS_Limits.MIN_X,
_GBS_Limits.MAX_X))
if (fs < _GBS_Limits.MIN_FS) or (fs > _GBS_Limits.MAX_FS):
raise GBS_InputError(
"Invalid Input Forward/Spot Price (FS). Acceptable range for inputs is {1} to {2}".format(fs,
_GBS_Limits.MIN_FS,
_GBS_Limits.MAX_FS))
if (t < _GBS_Limits.MIN_T) or (t > _GBS_Limits.MAX_T):
raise GBS_InputError(
"Invalid Input Time (T = {0}). Acceptable range for inputs is {1} to {2}".format(t, _GBS_Limits.MIN_T,
_GBS_Limits.MAX_T))
if (b < _GBS_Limits.MIN_b) or (b > _GBS_Limits.MAX_b):
raise GBS_InputError(
"Invalid Input Cost of Carry (b = {0}). Acceptable range for inputs is {1} to {2}".format(b,
_GBS_Limits.MIN_b,
_GBS_Limits.MAX_b))
if (r < _GBS_Limits.MIN_r) or (r > _GBS_Limits.MAX_r):
raise GBS_InputError(
"Invalid Input Risk Free Rate (r = {0}). Acceptable range for inputs is {1} to {2}".format(r,
_GBS_Limits.MIN_r,
_GBS_Limits.MAX_r))
if (v < _GBS_Limits.MIN_V) or (v > _GBS_Limits.MAX_V):
raise GBS_InputError(
"Invalid Input Implied Volatility (V = {0}). Acceptable range for inputs is {1} to {2}".format(v,
_GBS_Limits.MIN_V,
_GBS_Limits.MAX_V))
# The primary class for calculating Generalized Black Scholes option prices and deltas
# It is not intended to be part of this module's public interface
# Inputs: option_type = "p" or "c", fs = price of underlying, x = strike, t = time to expiration, r = risk free rate
# b = cost of carry, v = implied volatility
# Outputs: value, delta, gamma, theta, vega, rho
def _gbs(option_type, fs, x, t, r, b, v):
_debug("Debugging Information: _gbs()")
# -----------
# Test Inputs (throwing an exception on failure)
_gbs_test_inputs(option_type, fs, x, t, r, b, v)
# -----------
# Create preliminary calculations
t__sqrt = math.sqrt(t)
d1 = (math.log(fs / x) + (b + (v * v) / 2) * t) / (v * t__sqrt)
d2 = d1 - v * t__sqrt
if option_type == "c":
# it's a call
_debug(" Call Option")
value = fs * math.exp((b - r) * t) * norm.cdf(d1) - x * math.exp(-r * t) * norm.cdf(d2)
delta = math.exp((b - r) * t) * norm.cdf(d1)
gamma = math.exp((b - r) * t) * norm.pdf(d1) / (fs * v * t__sqrt)
theta = -(fs * v * math.exp((b - r) * t) * norm.pdf(d1)) / (2 * t__sqrt) - (b - r) * fs * math.exp(
(b - r) * t) * norm.cdf(d1) - r * x * math.exp(-r * t) * norm.cdf(d2)
vega = math.exp((b - r) * t) * fs * t__sqrt * norm.pdf(d1)
rho = x * t * math.exp(-r * t) * norm.cdf(d2)
else:
# it's a put
_debug(" Put Option")
value = x * math.exp(-r * t) * norm.cdf(-d2) - (fs * math.exp((b - r) * t) * norm.cdf(-d1))
delta = -math.exp((b - r) * t) * norm.cdf(-d1)
gamma = math.exp((b - r) * t) * norm.pdf(d1) / (fs * v * t__sqrt)
theta = -(fs * v * math.exp((b - r) * t) * norm.pdf(d1)) / (2 * t__sqrt) + (b - r) * fs * math.exp(
(b - r) * t) * norm.cdf(-d1) + r * x * math.exp(-r * t) * norm.cdf(-d2)
vega = math.exp((b - r) * t) * fs * t__sqrt * norm.pdf(d1)
rho = -x * t * math.exp(-r * t) * norm.cdf(-d2)
_debug(" d1= {0}\n d2 = {1}".format(d1, d2))
_debug(" delta = {0}\n gamma = {1}\n theta = {2}\n vega = {3}\n rho={4}".format(delta, gamma,
theta, vega,
rho))
return value, delta, gamma, theta, vega, rho
```
### Implementation: American Options
This section contains the code necessary to price American options. The main function is _American(). The other functions are called from the main function.
```
# -----------
# Generalized American Option Pricer
# This is a wrapper to check inputs and route to the current "best" American option model
def _american_option(option_type, fs, x, t, r, b, v):
# -----------
# Test Inputs (throwing an exception on failure)
_debug("Debugging Information: _american_option()")
_gbs_test_inputs(option_type, fs, x, t, r, b, v)
# -----------
if option_type == "c":
# Call Option
_debug(" Call Option")
return _bjerksund_stensland_2002(fs, x, t, r, b, v)
else:
# Put Option
_debug(" Put Option")
# Using the put-call transformation: P(X, FS, T, r, b, V) = C(FS, X, T, -b, r-b, V)
# WARNING - When reconciling this code back to the B&S paper, the order of variables is different
put__x = fs
put_fs = x
put_b = -b
put_r = r - b
# pass updated values into the Call Valuation formula
return _bjerksund_stensland_2002(put_fs, put__x, t, put_r, put_b, v)
# -----------
# American Call Option (Bjerksund Stensland 1993 approximation)
# This is primarily here for testing purposes; 2002 model has superseded this one
def _bjerksund_stensland_1993(fs, x, t, r, b, v):
# -----------
# initialize output
# using GBS greeks (TO DO: update greek calculations)
my_output = _gbs("c", fs, x, t, r, b, v)
e_value = my_output[0]
delta = my_output[1]
gamma = my_output[2]
theta = my_output[3]
vega = my_output[4]
rho = my_output[5]
# debugging for calculations
_debug("-----")
_debug("Debug Information: _Bjerksund_Stensland_1993())")
# if b >= r, it is never optimal to exercise before maturity
# so we can return the GBS value
if b >= r:
_debug(" b >= r, early exercise never optimal, returning GBS value")
return e_value, delta, gamma, theta, vega, rho
# Intermediate Calculations
v2 = v ** 2
sqrt_t = math.sqrt(t)
beta = (0.5 - b / v2) + math.sqrt(((b / v2 - 0.5) ** 2) + 2 * r / v2)
b_infinity = (beta / (beta - 1)) * x
b_zero = max(x, (r / (r - b)) * x)
h1 = -(b * t + 2 * v * sqrt_t) * (b_zero / (b_infinity - b_zero))
i = b_zero + (b_infinity - b_zero) * (1 - math.exp(h1))
alpha = (i - x) * (i ** (-beta))
# debugging for calculations
_debug(" b = {0}".format(b))
_debug(" v2 = {0}".format(v2))
_debug(" beta = {0}".format(beta))
_debug(" b_infinity = {0}".format(b_infinity))
_debug(" b_zero = {0}".format(b_zero))
_debug(" h1 = {0}".format(h1))
_debug(" i = {0}".format(i))
_debug(" alpha = {0}".format(alpha))
# Check for immediate exercise
if fs >= i:
_debug(" Immediate Exercise")
value = fs - x
else:
_debug(" American Exercise")
value = (alpha * (fs ** beta)
- alpha * _phi(fs, t, beta, i, i, r, b, v)
+ _phi(fs, t, 1, i, i, r, b, v)
- _phi(fs, t, 1, x, i, r, b, v)
- x * _phi(fs, t, 0, i, i, r, b, v)
+ x * _phi(fs, t, 0, x, i, r, b, v))
# The approximation can break down in boundary conditions
# make sure the value is at least equal to the European value
value = max(value, e_value)
return value, delta, gamma, theta, vega, rho
# -----------
# American Call Option (Bjerksund Stensland 2002 approximation)
def _bjerksund_stensland_2002(fs, x, t, r, b, v):
# -----------
# initialize output
# using GBS greeks (TO DO: update greek calculations)
my_output = _gbs("c", fs, x, t, r, b, v)
e_value = my_output[0]
delta = my_output[1]
gamma = my_output[2]
theta = my_output[3]
vega = my_output[4]
rho = my_output[5]
# debugging for calculations
_debug("-----")
_debug("Debug Information: _Bjerksund_Stensland_2002())")
# If b >= r, it is never optimal to exercise before maturity
# so we can return the GBS value
if b >= r:
_debug(" Returning GBS value")
return e_value, delta, gamma, theta, vega, rho
# -----------
# Create preliminary calculations
v2 = v ** 2
t1 = 0.5 * (math.sqrt(5) - 1) * t
t2 = t
beta_inside = ((b / v2 - 0.5) ** 2) + 2 * r / v2
# forcing the inside of the sqrt to be a positive number
beta_inside = abs(beta_inside)
beta = (0.5 - b / v2) + math.sqrt(beta_inside)
b_infinity = (beta / (beta - 1)) * x
b_zero = max(x, (r / (r - b)) * x)
h1 = -(b * t1 + 2 * v * math.sqrt(t1)) * ((x ** 2) / ((b_infinity - b_zero) * b_zero))
h2 = -(b * t2 + 2 * v * math.sqrt(t2)) * ((x ** 2) / ((b_infinity - b_zero) * b_zero))
i1 = b_zero + (b_infinity - b_zero) * (1 - math.exp(h1))
i2 = b_zero + (b_infinity - b_zero) * (1 - math.exp(h2))
alpha1 = (i1 - x) * (i1 ** (-beta))
alpha2 = (i2 - x) * (i2 ** (-beta))
# debugging for calculations
_debug(" t1 = {0}".format(t1))
_debug(" beta = {0}".format(beta))
_debug(" b_infinity = {0}".format(b_infinity))
_debug(" b_zero = {0}".format(b_zero))
_debug(" h1 = {0}".format(h1))
_debug(" h2 = {0}".format(h2))
_debug(" i1 = {0}".format(i1))
_debug(" i2 = {0}".format(i2))
_debug(" alpha1 = {0}".format(alpha1))
_debug(" alpha2 = {0}".format(alpha2))
# check for immediate exercise
if fs >= i2:
value = fs - x
else:
# Perform the main calculation
value = (alpha2 * (fs ** beta)
- alpha2 * _phi(fs, t1, beta, i2, i2, r, b, v)
+ _phi(fs, t1, 1, i2, i2, r, b, v)
- _phi(fs, t1, 1, i1, i2, r, b, v)
- x * _phi(fs, t1, 0, i2, i2, r, b, v)
+ x * _phi(fs, t1, 0, i1, i2, r, b, v)
+ alpha1 * _phi(fs, t1, beta, i1, i2, r, b, v)
- alpha1 * _psi(fs, t2, beta, i1, i2, i1, t1, r, b, v)
+ _psi(fs, t2, 1, i1, i2, i1, t1, r, b, v)
- _psi(fs, t2, 1, x, i2, i1, t1, r, b, v)
- x * _psi(fs, t2, 0, i1, i2, i1, t1, r, b, v)
+ x * _psi(fs, t2, 0, x, i2, i1, t1, r, b, v))
# in boundary conditions, this approximation can break down
# Make sure option value is greater than or equal to European value
value = max(value, e_value)
# -----------
# Return Data
return value, delta, gamma, theta, vega, rho
# ---------------------------
# American Option Intermediate Calculations
# -----------
# The Psi() function used by _Bjerksund_Stensland_2002 model
def _psi(fs, t2, gamma, h, i2, i1, t1, r, b, v):
vsqrt_t1 = v * math.sqrt(t1)
vsqrt_t2 = v * math.sqrt(t2)
bgamma_t1 = (b + (gamma - 0.5) * (v ** 2)) * t1
bgamma_t2 = (b + (gamma - 0.5) * (v ** 2)) * t2
d1 = (math.log(fs / i1) + bgamma_t1) / vsqrt_t1
d3 = (math.log(fs / i1) - bgamma_t1) / vsqrt_t1
d2 = (math.log((i2 ** 2) / (fs * i1)) + bgamma_t1) / vsqrt_t1
d4 = (math.log((i2 ** 2) / (fs * i1)) - bgamma_t1) / vsqrt_t1
e1 = (math.log(fs / h) + bgamma_t2) / vsqrt_t2
e2 = (math.log((i2 ** 2) / (fs * h)) + bgamma_t2) / vsqrt_t2
e3 = (math.log((i1 ** 2) / (fs * h)) + bgamma_t2) / vsqrt_t2
e4 = (math.log((fs * (i1 ** 2)) / (h * (i2 ** 2))) + bgamma_t2) / vsqrt_t2
tau = math.sqrt(t1 / t2)
lambda1 = (-r + gamma * b + 0.5 * gamma * (gamma - 1) * (v ** 2))
kappa = (2 * b) / (v ** 2) + (2 * gamma - 1)
psi = math.exp(lambda1 * t2) * (fs ** gamma) * (_cbnd(-d1, -e1, tau)
- ((i2 / fs) ** kappa) * _cbnd(-d2, -e2, tau)
- ((i1 / fs) ** kappa) * _cbnd(-d3, -e3, -tau)
+ ((i1 / i2) ** kappa) * _cbnd(-d4, -e4, -tau))
return psi
# -----------
# The Phi() function used by _Bjerksund_Stensland_2002 model and the _Bjerksund_Stensland_1993 model
def _phi(fs, t, gamma, h, i, r, b, v):
d1 = -(math.log(fs / h) + (b + (gamma - 0.5) * (v ** 2)) * t) / (v * math.sqrt(t))
d2 = d1 - 2 * math.log(i / fs) / (v * math.sqrt(t))
lambda1 = (-r + gamma * b + 0.5 * gamma * (gamma - 1) * (v ** 2))
kappa = (2 * b) / (v ** 2) + (2 * gamma - 1)
phi = math.exp(lambda1 * t) * (fs ** gamma) * (norm.cdf(d1) - ((i / fs) ** kappa) * norm.cdf(d2))
_debug("-----")
_debug("Debug info for: _phi()")
_debug(" d1={0}".format(d1))
_debug(" d2={0}".format(d2))
_debug(" lambda={0}".format(lambda1))
_debug(" kappa={0}".format(kappa))
_debug(" phi={0}".format(phi))
return phi
# -----------
# Cumulative Bivariate Normal Distribution
# Primarily called by Psi() function, part of the _Bjerksund_Stensland_2002 model
def _cbnd(a, b, rho):
# This distribution uses the Genz multi-variate normal distribution
# code found as part of the standard SciPy distribution
lower = np.array([0, 0])
upper = np.array([a, b])
infin = np.array([0, 0])
correl = rho
error, value, inform = mvn.mvndst(lower, upper, infin, correl)
return value
```
### Implementation: Implied Vol
This section implements implied volatility calculations. It contains 3 main models:
1. **At-the-Money approximation.** This is a very fast approximation for implied volatility. It is used to estimate a starting point for the search functions.
2. **Newton-Raphson Search.** This is a fast implied volatility search that can be used when there is a reliable estimate of Vega (i.e., European options)
3. **Bisection Search.** An implied volatility search (not quite as fast as a Newton search) that can be used where there is no reliable Vega estimate (i.e., American options).
```
# ----------
# Inputs (not all functions use all inputs)
# fs = forward/spot price
# x = Strike
# t = Time (in years)
# r = risk free rate
# b = cost of carry
# cp = Call or Put price
# precision = (optional) precision at stopping point
# max_steps = (optional) maximum number of steps
# ----------
# Approximate Implied Volatility
#
# This function is used to choose a starting point for the
# search functions (Newton and bisection searches).
# Brenner & Subrahmanyam (1988), Feinstein (1988)
def _approx_implied_vol(option_type, fs, x, t, r, b, cp):
_test_option_type(option_type)
ebrt = math.exp((b - r) * t)
ert = math.exp(-r * t)
a = math.sqrt(2 * math.pi) / (fs * ebrt + x * ert)
if option_type == "c":
payoff = fs * ebrt - x * ert
else:
payoff = x * ert - fs * ebrt
b = cp - payoff / 2
c = (payoff ** 2) / math.pi
v = (a * (b + math.sqrt(b ** 2 + c))) / math.sqrt(t)
return v
# ----------
# Find the Implied Volatility of an European (GBS) Option given a price
# using Newton-Raphson method for greater speed since Vega is available
def _gbs_implied_vol(option_type, fs, x, t, r, b, cp, precision=.00001, max_steps=100):
return _newton_implied_vol(_gbs, option_type, x, fs, t, b, r, cp, precision, max_steps)
# ----------
# Find the Implied Volatility of an American Option given a price
# Using bisection method since Vega is difficult to estimate for Americans
def _american_implied_vol(option_type, fs, x, t, r, b, cp, precision=.00001, max_steps=100):
return _bisection_implied_vol(_american_option, option_type, fs, x, t, r, b, cp, precision, max_steps)
# ----------
# Calculate Implied Volatility with a Newton Raphson search
def _newton_implied_vol(val_fn, option_type, x, fs, t, b, r, cp, precision=.00001, max_steps=100):
# make sure a valid option type was entered
_test_option_type(option_type)
# Estimate starting Vol, making sure it is allowable range
v = _approx_implied_vol(option_type, fs, x, t, r, b, cp)
v = max(_GBS_Limits.MIN_V, v)
v = min(_GBS_Limits.MAX_V, v)
# Calculate the value at the estimated vol
value, delta, gamma, theta, vega, rho = val_fn(option_type, fs, x, t, r, b, v)
min_diff = abs(cp - value)
_debug("-----")
_debug("Debug info for: _Newton_ImpliedVol()")
_debug(" Vinitial={0}".format(v))
# Newton-Raphson Search
countr = 0
while precision <= abs(cp - value) <= min_diff and countr < max_steps:
v = v - (value - cp) / vega
if (v > _GBS_Limits.MAX_V) or (v < _GBS_Limits.MIN_V):
_debug(" Volatility out of bounds")
break
value, delta, gamma, theta, vega, rho = val_fn(option_type, fs, x, t, r, b, v)
min_diff = min(abs(cp - value), min_diff)
# keep track of how many loops
countr += 1
_debug(" IVOL STEP {0}. v={1}".format(countr, v))
# check if function converged and return a value
if abs(cp - value) < precision:
# the search function converged
return v
else:
# if the search function didn't converge, try a bisection search
return _bisection_implied_vol(val_fn, option_type, fs, x, t, r, b, cp, precision, max_steps)
# ----------
# Find the Implied Volatility using a Bisection search
def _bisection_implied_vol(val_fn, option_type, fs, x, t, r, b, cp, precision=.00001, max_steps=100):
_debug("-----")
_debug("Debug info for: _bisection_implied_vol()")
# Estimate Upper and Lower bounds on volatility
# Assume American Implied vol is within +/- 50% of the GBS Implied Vol
v_mid = _approx_implied_vol(option_type, fs, x, t, r, b, cp)
if (v_mid <= _GBS_Limits.MIN_V) or (v_mid >= _GBS_Limits.MAX_V):
# if the volatility estimate is out of bounds, search entire allowed vol space
v_low = _GBS_Limits.MIN_V
v_high = _GBS_Limits.MAX_V
v_mid = (v_low + v_high) / 2
else:
# reduce the size of the vol space
v_low = max(_GBS_Limits.MIN_V, v_mid * .5)
v_high = min(_GBS_Limits.MAX_V, v_mid * 1.5)
# Estimate the high/low bounds on price
cp_mid = val_fn(option_type, fs, x, t, r, b, v_mid)[0]
# initialize bisection loop
current_step = 0
diff = abs(cp - cp_mid)
_debug(" American IVOL starting conditions: CP={0} cp_mid={1}".format(cp, cp_mid))
_debug(" IVOL {0}. V[{1},{2},{3}]".format(current_step, v_low, v_mid, v_high))
# Keep bisection volatility until correct price is found
while (diff > precision) and (current_step < max_steps):
current_step += 1
# Cut the search area in half
if cp_mid < cp:
v_low = v_mid
else:
v_high = v_mid
cp_low = val_fn(option_type, fs, x, t, r, b, v_low)[0]
cp_high = val_fn(option_type, fs, x, t, r, b, v_high)[0]
v_mid = v_low + (cp - cp_low) * (v_high - v_low) / (cp_high - cp_low)
v_mid = max(_GBS_Limits.MIN_V, v_mid) # enforce high/low bounds
v_mid = min(_GBS_Limits.MAX_V, v_mid) # enforce high/low bounds
cp_mid = val_fn(option_type, fs, x, t, r, b, v_mid)[0]
diff = abs(cp - cp_mid)
_debug(" IVOL {0}. V[{1},{2},{3}]".format(current_step, v_low, v_mid, v_high))
# return output
if abs(cp - cp_mid) < precision:
return v_mid
else:
raise GBS_CalculationError(
"Implied Vol did not converge. Best Guess={0}, Price diff={1}, Required Precision={2}".format(v_mid, diff,
precision))
```
--------------------
### Public Interface for valuation functions
This section encapsulates the functions that user will call to value certain options. These function primarily figure out the cost-of-carry term (b) and then call the generic version of the function (like _GBS() or _American). All of these functions return an array containg the premium and the greeks.
```
# This is the public interface for European Options
# Each call does a little bit of processing and then calls the calculations located in the _gbs module
# Inputs:
# option_type = "p" or "c"
# fs = price of underlying
# x = strike
# t = time to expiration
# v = implied volatility
# r = risk free rate
# q = dividend payment
# b = cost of carry
# Outputs:
# value = price of the option
# delta = first derivative of value with respect to price of underlying
# gamma = second derivative of value w.r.t price of underlying
# theta = first derivative of value w.r.t. time to expiration
# vega = first derivative of value w.r.t. implied volatility
# rho = first derivative of value w.r.t. risk free rates
# ---------------------------
# Black Scholes: stock Options (no dividend yield)
def black_scholes(option_type, fs, x, t, r, v):
b = r
return _gbs(option_type, fs, x, t, r, b, v)
# ---------------------------
# Merton Model: Stocks Index, stocks with a continuous dividend yields
def merton(option_type, fs, x, t, r, q, v):
b = r - q
return _gbs(option_type, fs, x, t, r, b, v)
# ---------------------------
# Commodities
def black_76(option_type, fs, x, t, r, v):
b = 0
return _gbs(option_type, fs, x, t, r, b, v)
# ---------------------------
# FX Options
def garman_kohlhagen(option_type, fs, x, t, r, rf, v):
b = r - rf
return _gbs(option_type, fs, x, t, r, b, v)
# ---------------------------
# Average Price option on commodities
def asian_76(option_type, fs, x, t, t_a, r, v):
# Check that TA is reasonable
if (t_a < _GBS_Limits.MIN_TA) or (t_a > t):
raise GBS_InputError(
"Invalid Input Averaging Time (TA = {0}). Acceptable range for inputs is {1} to <T".format(t_a,
_GBS_Limits.MIN_TA))
# Approximation to value Asian options on commodities
b = 0
if t_a == t:
# if there is no averaging period, this is just Black Scholes
v_a = v
else:
# Approximate the volatility
m = (2 * math.exp((v ** 2) * t) - 2 * math.exp((v ** 2) * t_a) * (1 + (v ** 2) * (t - t_a))) / (
(v ** 4) * ((t - t_a) ** 2))
v_a = math.sqrt(math.log(m) / t)
# Finally, have the GBS function do the calculation
return _gbs(option_type, fs, x, t, r, b, v_a)
# ---------------------------
# Spread Option formula
def kirks_76(option_type, f1, f2, x, t, r, v1, v2, corr):
# create the modifications to the GBS formula to handle spread options
b = 0
fs = f1 / (f2 + x)
f_temp = f2 / (f2 + x)
v = math.sqrt((v1 ** 2) + ((v2 * f_temp) ** 2) - (2 * corr * v1 * v2 * f_temp))
my_values = _gbs(option_type, fs, 1.0, t, r, b, v)
# Have the GBS function return a value
return my_values[0] * (f2 + x), 0, 0, 0, 0, 0
# ---------------------------
# American Options (stock style, set q=0 for non-dividend paying options)
def american(option_type, fs, x, t, r, q, v):
b = r - q
return _american_option(option_type, fs, x, t, r, b, v)
# ---------------------------
# Commodities
def american_76(option_type, fs, x, t, r, v):
b = 0
return _american_option(option_type, fs, x, t, r, b, v)
```
### Public Interface for implied Volatility Functions
```
# Inputs:
# option_type = "p" or "c"
# fs = price of underlying
# x = strike
# t = time to expiration
# v = implied volatility
# r = risk free rate
# q = dividend payment
# b = cost of carry
# Outputs:
# value = price of the option
# delta = first derivative of value with respect to price of underlying
# gamma = second derivative of value w.r.t price of underlying
# theta = first derivative of value w.r.t. time to expiration
# vega = first derivative of value w.r.t. implied volatility
# rho = first derivative of value w.r.t. risk free rates
def euro_implied_vol(option_type, fs, x, t, r, q, cp):
b = r - q
return _gbs_implied_vol(option_type, fs, x, t, r, b, cp)
def euro_implied_vol_76(option_type, fs, x, t, r, cp):
b = 0
return _gbs_implied_vol(option_type, fs, x, t, r, b, cp)
def amer_implied_vol(option_type, fs, x, t, r, q, cp):
b = r - q
return _american_implied_vol(option_type, fs, x, t, r, b, cp)
def amer_implied_vol_76(option_type, fs, x, t, r, cp):
b = 0
return _american_implied_vol(option_type, fs, x, t, r, b, cp)
```
### Implementation: Helper Functions
These functions aren't part of the main code but serve as utility function mostly used for debugging
```
# ---------------------------
# Helper Function for Debugging
# Prints a message if running code from this module and _DEBUG is set to true
# otherwise, do nothing
def _debug(debug_input):
if (__name__ is "__main__") and (_DEBUG is True):
print(debug_input)
# This class defines the Exception that gets thrown when invalid input is placed into the GBS function
class GBS_InputError(Exception):
def __init__(self, mismatch):
Exception.__init__(self, mismatch)
# This class defines the Exception that gets thrown when there is a calculation error
class GBS_CalculationError(Exception):
def __init__(self, mismatch):
Exception.__init__(self, mismatch)
# This function tests that two floating point numbers are the same
# Numbers less than 1 million are considered the same if they are within .000001 of each other
# Numbers larger than 1 million are considered the same if they are within .0001% of each other
# User can override the default precision if necessary
def assert_close(value_a, value_b, precision=.000001):
my_precision = precision
if (value_a < 1000000.0) and (value_b < 1000000.0):
my_diff = abs(value_a - value_b)
my_diff_type = "Difference"
else:
my_diff = abs((value_a - value_b) / value_a)
my_diff_type = "Percent Difference"
_debug("Comparing {0} and {1}. Difference is {2}, Difference Type is {3}".format(value_a, value_b, my_diff,
my_diff_type))
if my_diff < my_precision:
my_result = True
else:
my_result = False
if (__name__ is "__main__") and (my_result is False):
print(" FAILED TEST. Comparing {0} and {1}. Difference is {2}, Difference Type is {3}".format(value_a, value_b,
my_diff,
my_diff_type))
else:
print(".")
return my_result
```
## Unit Testing
This will print out a "." if the test is successful or an error message if the test fails
```
if __name__ == "__main__":
print ("=====================================")
print ("American Options Intermediate Functions")
print ("=====================================")
# ---------------------------
# unit tests for _psi()
# _psi(FS, t2, gamma, H, I2, I1, t1, r, b, V):
print("Testing _psi (American Option Intermediate Calculation)")
assert_close(_psi(fs=120, t2=3, gamma=1, h=375, i2=375, i1=300, t1=1, r=.05, b=0.03, v=0.1), 112.87159814023171)
assert_close(_psi(fs=125, t2=2, gamma=1, h=100, i2=100, i1=75, t1=1, r=.05, b=0.03, v=0.1), 1.7805459905819128)
# ---------------------------
# unit tests for _phi()
print("Testing _phi (American Option Intermediate Calculation)")
# _phi(FS, T, gamma, h, I, r, b, V):
assert_close(_phi(fs=120, t=3, gamma=4.51339343051624, h=151.696096685711, i=151.696096685711, r=.02, b=-0.03, v=0.14),
1102886677.05955)
assert_close(_phi(fs=125, t=3, gamma=1, h=374.061664206768, i=374.061664206768, r=.05, b=0.03, v=0.14),
117.714544103477)
# ---------------------------
# unit tests for _CBND
print("Testing _CBND (Cumulative Binomial Normal Distribution)")
assert_close(_cbnd(0, 0, 0), 0.25)
assert_close(_cbnd(0, 0, -0.5), 0.16666666666666669)
assert_close(_cbnd(-0.5, 0, 0), 0.15426876936299347)
assert_close(_cbnd(0, -0.5, 0), 0.15426876936299347)
assert_close(_cbnd(0, -0.99999999, -0.99999999), 0.0)
assert_close(_cbnd(0.000001, -0.99999999, -0.99999999), 0.0)
assert_close(_cbnd(0, 0, 0.5), 0.3333333333333333)
assert_close(_cbnd(0.5, 0, 0), 0.3457312306370065)
assert_close(_cbnd(0, 0.5, 0), 0.3457312306370065)
assert_close(_cbnd(0, 0.99999999, 0.99999999), 0.5)
assert_close(_cbnd(0.000001, 0.99999999, 0.99999999), 0.5000003989422803)
# ---------------------------
# Testing American Options
if __name__ == "__main__":
print("=====================================")
print("American Options Testing")
print("=====================================")
print("testing _Bjerksund_Stensland_2002()")
# _american_option(option_type, X, FS, T, b, r, V)
assert_close(_bjerksund_stensland_2002(fs=90, x=100, t=0.5, r=0.1, b=0, v=0.15)[0], 0.8099, precision=.001)
assert_close(_bjerksund_stensland_2002(fs=100, x=100, t=0.5, r=0.1, b=0, v=0.25)[0], 6.7661, precision=.001)
assert_close(_bjerksund_stensland_2002(fs=110, x=100, t=0.5, r=0.1, b=0, v=0.35)[0], 15.5137, precision=.001)
assert_close(_bjerksund_stensland_2002(fs=100, x=90, t=0.5, r=.1, b=0, v=0.15)[0], 10.5400, precision=.001)
assert_close(_bjerksund_stensland_2002(fs=100, x=100, t=0.5, r=.1, b=0, v=0.25)[0], 6.7661, precision=.001)
assert_close(_bjerksund_stensland_2002(fs=100, x=110, t=0.5, r=.1, b=0, v=0.35)[0], 5.8374, precision=.001)
print("testing _Bjerksund_Stensland_1993()")
# Prices for 1993 model slightly different than those presented in Haug's Complete Guide to Option Pricing Formulas
# Possibly due to those results being based on older CBND calculation?
assert_close(_bjerksund_stensland_1993(fs=90, x=100, t=0.5, r=0.1, b=0, v=0.15)[0], 0.8089, precision=.001)
assert_close(_bjerksund_stensland_1993(fs=100, x=100, t=0.5, r=0.1, b=0, v=0.25)[0], 6.757, precision=.001)
assert_close(_bjerksund_stensland_1993(fs=110, x=100, t=0.5, r=0.1, b=0, v=0.35)[0], 15.4998, precision=.001)
print("testing _american_option()")
assert_close(_american_option("p", fs=90, x=100, t=0.5, r=0.1, b=0, v=0.15)[0], 10.5400, precision=.001)
assert_close(_american_option("p", fs=100, x=100, t=0.5, r=0.1, b=0, v=0.25)[0], 6.7661, precision=.001)
assert_close(_american_option("p", fs=110, x=100, t=0.5, r=0.1, b=0, v=0.35)[0], 5.8374, precision=.001)
assert_close(_american_option('c', fs=100, x=95, t=0.00273972602739726, r=0.000751040922831883, b=0, v=0.2)[0], 5.0,
precision=.01)
assert_close(_american_option('c', fs=42, x=40, t=0.75, r=0.04, b=-0.04, v=0.35)[0], 5.28, precision=.01)
assert_close(_american_option('c', fs=90, x=100, t=0.1, r=0.10, b=0, v=0.15)[0], 0.02, precision=.01)
print("Testing that American valuation works for integer inputs")
assert_close(_american_option('c', fs=100, x=100, t=1, r=0, b=0, v=0.35)[0], 13.892, precision=.001)
assert_close(_american_option('p', fs=100, x=100, t=1, r=0, b=0, v=0.35)[0], 13.892, precision=.001)
print("Testing valuation works at minimum/maximum values for T")
assert_close(_american_option('c', 100, 100, 0.00396825396825397, 0.000771332656950173, 0, 0.15)[0], 0.3769,
precision=.001)
assert_close(_american_option('p', 100, 100, 0.00396825396825397, 0.000771332656950173, 0, 0.15)[0], 0.3769,
precision=.001)
assert_close(_american_option('c', 100, 100, 100, 0.042033868311581, 0, 0.15)[0], 18.61206, precision=.001)
assert_close(_american_option('p', 100, 100, 100, 0.042033868311581, 0, 0.15)[0], 18.61206, precision=.001)
print("Testing valuation works at minimum/maximum values for X")
assert_close(_american_option('c', 100, 0.01, 1, 0.00330252458693489, 0, 0.15)[0], 99.99, precision=.001)
assert_close(_american_option('p', 100, 0.01, 1, 0.00330252458693489, 0, 0.15)[0], 0, precision=.001)
assert_close(_american_option('c', 100, 2147483248, 1, 0.00330252458693489, 0, 0.15)[0], 0, precision=.001)
assert_close(_american_option('p', 100, 2147483248, 1, 0.00330252458693489, 0, 0.15)[0], 2147483148, precision=.001)
print("Testing valuation works at minimum/maximum values for F/S")
assert_close(_american_option('c', 0.01, 100, 1, 0.00330252458693489, 0, 0.15)[0], 0, precision=.001)
assert_close(_american_option('p', 0.01, 100, 1, 0.00330252458693489, 0, 0.15)[0], 99.99, precision=.001)
assert_close(_american_option('c', 2147483248, 100, 1, 0.00330252458693489, 0, 0.15)[0], 2147483148, precision=.001)
assert_close(_american_option('p', 2147483248, 100, 1, 0.00330252458693489, 0, 0.15)[0], 0, precision=.001)
print("Testing valuation works at minimum/maximum values for b")
assert_close(_american_option('c', 100, 100, 1, 0, -1, 0.15)[0], 0.0, precision=.001)
assert_close(_american_option('p', 100, 100, 1, 0, -1, 0.15)[0], 63.2121, precision=.001)
assert_close(_american_option('c', 100, 100, 1, 0, 1, 0.15)[0], 171.8282, precision=.001)
assert_close(_american_option('p', 100, 100, 1, 0, 1, 0.15)[0], 0.0, precision=.001)
print("Testing valuation works at minimum/maximum values for r")
assert_close(_american_option('c', 100, 100, 1, -1, 0, 0.15)[0], 16.25133, precision=.001)
assert_close(_american_option('p', 100, 100, 1, -1, 0, 0.15)[0], 16.25133, precision=.001)
assert_close(_american_option('c', 100, 100, 1, 1, 0, 0.15)[0], 3.6014, precision=.001)
assert_close(_american_option('p', 100, 100, 1, 1, 0, 0.15)[0], 3.6014, precision=.001)
print("Testing valuation works at minimum/maximum values for V")
assert_close(_american_option('c', 100, 100, 1, 0.05, 0, 0.005)[0], 0.1916, precision=.001)
assert_close(_american_option('p', 100, 100, 1, 0.05, 0, 0.005)[0], 0.1916, precision=.001)
assert_close(_american_option('c', 100, 100, 1, 0.05, 0, 1)[0], 36.4860, precision=.001)
assert_close(_american_option('p', 100, 100, 1, 0.05, 0, 1)[0], 36.4860, precision=.001)
# ---------------------------
# Testing European Options
if __name__ == "__main__":
print("=====================================")
print("Generalized Black Scholes (GBS) Testing")
print("=====================================")
print("testing GBS Premium")
assert_close(_gbs('c', 100, 95, 0.00273972602739726, 0.000751040922831883, 0, 0.2)[0], 4.99998980469552)
assert_close(_gbs('c', 92.45, 107.5, 0.0876712328767123, 0.00192960198828152, 0, 0.3)[0], 0.162619795863781)
assert_close(_gbs('c', 93.0766666666667, 107.75, 0.164383561643836, 0.00266390125346286, 0, 0.2878)[0],
0.584588840095316)
assert_close(_gbs('c', 93.5333333333333, 107.75, 0.249315068493151, 0.00319934651984034, 0, 0.2907)[0],
1.27026849732877)
assert_close(_gbs('c', 93.8733333333333, 107.75, 0.331506849315069, 0.00350934592318849, 0, 0.2929)[0],
1.97015685523537)
assert_close(_gbs('c', 94.1166666666667, 107.75, 0.416438356164384, 0.00367360967852615, 0, 0.2919)[0],
2.61731599547608)
assert_close(_gbs('p', 94.2666666666667, 107.75, 0.498630136986301, 0.00372609838856132, 0, 0.2888)[0],
16.6074587545269)
assert_close(_gbs('p', 94.3666666666667, 107.75, 0.583561643835616, 0.00370681407974257, 0, 0.2923)[0],
17.1686196701434)
assert_close(_gbs('p', 94.44, 107.75, 0.668493150684932, 0.00364163303865433, 0, 0.2908)[0], 17.6038273793172)
assert_close(_gbs('p', 94.4933333333333, 107.75, 0.750684931506849, 0.00355604221290591, 0, 0.2919)[0],
18.0870982577296)
assert_close(_gbs('p', 94.49, 107.75, 0.835616438356164, 0.00346100468320478, 0, 0.2901)[0], 18.5149895730975)
assert_close(_gbs('p', 94.39, 107.75, 0.917808219178082, 0.00337464630758452, 0, 0.2876)[0], 18.9397688539483)
print("Testing that valuation works for integer inputs")
assert_close(_gbs('c', fs=100, x=95, t=1, r=1, b=0, v=1)[0], 14.6711476484)
assert_close(_gbs('p', fs=100, x=95, t=1, r=1, b=0, v=1)[0], 12.8317504425)
print("Testing valuation works at minimum/maximum values for T")
assert_close(_gbs('c', 100, 100, 0.00396825396825397, 0.000771332656950173, 0, 0.15)[0], 0.376962465712609)
assert_close(_gbs('p', 100, 100, 0.00396825396825397, 0.000771332656950173, 0, 0.15)[0], 0.376962465712609)
assert_close(_gbs('c', 100, 100, 100, 0.042033868311581, 0, 0.15)[0], 0.817104022604705)
assert_close(_gbs('p', 100, 100, 100, 0.042033868311581, 0, 0.15)[0], 0.817104022604705)
print("Testing valuation works at minimum/maximum values for X")
assert_close(_gbs('c', 100, 0.01, 1, 0.00330252458693489, 0, 0.15)[0], 99.660325245681)
assert_close(_gbs('p', 100, 0.01, 1, 0.00330252458693489, 0, 0.15)[0], 0)
assert_close(_gbs('c', 100, 2147483248, 1, 0.00330252458693489, 0, 0.15)[0], 0)
assert_close(_gbs('p', 100, 2147483248, 1, 0.00330252458693489, 0, 0.15)[0], 2140402730.16601)
print("Testing valuation works at minimum/maximum values for F/S")
assert_close(_gbs('c', 0.01, 100, 1, 0.00330252458693489, 0, 0.15)[0], 0)
assert_close(_gbs('p', 0.01, 100, 1, 0.00330252458693489, 0, 0.15)[0], 99.660325245681)
assert_close(_gbs('c', 2147483248, 100, 1, 0.00330252458693489, 0, 0.15)[0], 2140402730.16601)
assert_close(_gbs('p', 2147483248, 100, 1, 0.00330252458693489, 0, 0.15)[0], 0)
print("Testing valuation works at minimum/maximum values for b")
assert_close(_gbs('c', 100, 100, 1, 0.05, -1, 0.15)[0], 1.62505648981223E-11)
assert_close(_gbs('p', 100, 100, 1, 0.05, -1, 0.15)[0], 60.1291675389721)
assert_close(_gbs('c', 100, 100, 1, 0.05, 1, 0.15)[0], 163.448023481557)
assert_close(_gbs('p', 100, 100, 1, 0.05, 1, 0.15)[0], 4.4173615264761E-11)
print("Testing valuation works at minimum/maximum values for r")
assert_close(_gbs('c', 100, 100, 1, -1, 0, 0.15)[0], 16.2513262267156)
assert_close(_gbs('p', 100, 100, 1, -1, 0, 0.15)[0], 16.2513262267156)
assert_close(_gbs('c', 100, 100, 1, 1, 0, 0.15)[0], 2.19937783786316)
assert_close(_gbs('p', 100, 100, 1, 1, 0, 0.15)[0], 2.19937783786316)
print("Testing valuation works at minimum/maximum values for V")
assert_close(_gbs('c', 100, 100, 1, 0.05, 0, 0.005)[0], 0.189742620249)
assert_close(_gbs('p', 100, 100, 1, 0.05, 0, 0.005)[0], 0.189742620249)
assert_close(_gbs('c', 100, 100, 1, 0.05, 0, 1)[0], 36.424945370234)
assert_close(_gbs('p', 100, 100, 1, 0.05, 0, 1)[0], 36.424945370234)
print("Checking that Greeks work for calls")
assert_close(_gbs('c', 100, 100, 1, 0.05, 0, 0.15)[0], 5.68695251984796)
assert_close(_gbs('c', 100, 100, 1, 0.05, 0, 0.15)[1], 0.50404947485)
assert_close(_gbs('c', 100, 100, 1, 0.05, 0, 0.15)[2], 0.025227988795588)
assert_close(_gbs('c', 100, 100, 1, 0.05, 0, 0.15)[3], -2.55380111351125)
assert_close(_gbs('c', 100, 100, 2, 0.05, 0.05, 0.25)[4], 50.7636345571413)
assert_close(_gbs('c', 100, 100, 1, 0.05, 0, 0.15)[5], 44.7179949651117)
print("Checking that Greeks work for puts")
assert_close(_gbs('p', 100, 100, 1, 0.05, 0, 0.15)[0], 5.68695251984796)
assert_close(_gbs('p', 100, 100, 1, 0.05, 0, 0.15)[1], -0.447179949651)
assert_close(_gbs('p', 100, 100, 1, 0.05, 0, 0.15)[2], 0.025227988795588)
assert_close(_gbs('p', 100, 100, 1, 0.05, 0, 0.15)[3], -2.55380111351125)
assert_close(_gbs('p', 100, 100, 2, 0.05, 0.05, 0.25)[4], 50.7636345571413)
assert_close(_gbs('p', 100, 100, 1, 0.05, 0, 0.15)[5], -50.4049474849597)
# ---------------------------
# Testing Implied Volatility
if __name__ == "__main__":
print("=====================================")
print("Implied Volatility Testing")
print("=====================================")
print("For options far away from ATM or those very near to expiry, volatility")
print("doesn't have a major effect on the price. When large changes in vol result in")
print("price changes less than the minimum precision, it is very difficult to test implied vol")
print("=====================================")
print ("testing at-the-money approximation")
assert_close(_approx_implied_vol(option_type="c", fs=100, x=100, t=1, r=.05, b=0, cp=5),0.131757)
assert_close(_approx_implied_vol(option_type="c", fs=59, x=60, t=0.25, r=.067, b=0.067, cp=2.82),0.239753)
print("testing GBS Implied Vol")
assert_close(_gbs_implied_vol('c', 92.45, 107.5, 0.0876712328767123, 0.00192960198828152, 0, 0.162619795863781),0.3)
assert_close(_gbs_implied_vol('c', 93.0766666666667, 107.75, 0.164383561643836, 0.00266390125346286, 0, 0.584588840095316),0.2878)
assert_close(_gbs_implied_vol('c', 93.5333333333333, 107.75, 0.249315068493151, 0.00319934651984034, 0, 1.27026849732877),0.2907)
assert_close(_gbs_implied_vol('c', 93.8733333333333, 107.75, 0.331506849315069, 0.00350934592318849, 0, 1.97015685523537),0.2929)
assert_close(_gbs_implied_vol('c', 94.1166666666667, 107.75, 0.416438356164384, 0.00367360967852615, 0, 2.61731599547608),0.2919)
assert_close(_gbs_implied_vol('p', 94.2666666666667, 107.75, 0.498630136986301, 0.00372609838856132, 0, 16.6074587545269),0.2888)
assert_close(_gbs_implied_vol('p', 94.3666666666667, 107.75, 0.583561643835616, 0.00370681407974257, 0, 17.1686196701434),0.2923)
assert_close(_gbs_implied_vol('p', 94.44, 107.75, 0.668493150684932, 0.00364163303865433, 0, 17.6038273793172),0.2908)
assert_close(_gbs_implied_vol('p', 94.4933333333333, 107.75, 0.750684931506849, 0.00355604221290591, 0, 18.0870982577296),0.2919)
assert_close(_gbs_implied_vol('p', 94.39, 107.75, 0.917808219178082, 0.00337464630758452, 0, 18.9397688539483),0.2876)
print("Testing that GBS implied vol works for integer inputs")
assert_close(_gbs_implied_vol('c', fs=100, x=95, t=1, r=1, b=0, cp=14.6711476484), 1)
assert_close(_gbs_implied_vol('p', fs=100, x=95, t=1, r=1, b=0, cp=12.8317504425), 1)
print("testing American Option implied volatility")
assert_close(_american_implied_vol("p", fs=90, x=100, t=0.5, r=0.1, b=0, cp=10.54), 0.15, precision=0.01)
assert_close(_american_implied_vol("p", fs=100, x=100, t=0.5, r=0.1, b=0, cp=6.7661), 0.25, precision=0.0001)
assert_close(_american_implied_vol("p", fs=110, x=100, t=0.5, r=0.1, b=0, cp=5.8374), 0.35, precision=0.0001)
assert_close(_american_implied_vol('c', fs=42, x=40, t=0.75, r=0.04, b=-0.04, cp=5.28), 0.35, precision=0.01)
assert_close(_american_implied_vol('c', fs=90, x=100, t=0.1, r=0.10, b=0, cp=0.02), 0.15, precision=0.01)
print("Testing that American implied volatility works for integer inputs")
assert_close(_american_implied_vol('c', fs=100, x=100, t=1, r=0, b=0, cp=13.892), 0.35, precision=0.01)
assert_close(_american_implied_vol('p', fs=100, x=100, t=1, r=0, b=0, cp=13.892), 0.35, precision=0.01)
# ---------------------------
# Testing the external interface
if __name__ == "__main__":
print("=====================================")
print("External Interface Testing")
print("=====================================")
# BlackScholes(option_type, X, FS, T, r, V)
print("Testing: GBS.BlackScholes")
assert_close(black_scholes('c', 102, 100, 2, 0.05, 0.25)[0], 20.02128028)
assert_close(black_scholes('p', 102, 100, 2, 0.05, 0.25)[0], 8.50502208)
# Merton(option_type, X, FS, T, r, q, V)
print("Testing: GBS.Merton")
assert_close(merton('c', 102, 100, 2, 0.05, 0.01, 0.25)[0], 18.63371484)
assert_close(merton('p', 102, 100, 2, 0.05, 0.01, 0.25)[0], 9.13719197)
# Black76(option_type, X, FS, T, r, V)
print("Testing: GBS.Black76")
assert_close(black_76('c', 102, 100, 2, 0.05, 0.25)[0], 13.74803567)
assert_close(black_76('p', 102, 100, 2, 0.05, 0.25)[0], 11.93836083)
# garman_kohlhagen(option_type, X, FS, T, b, r, rf, V)
print("Testing: GBS.garman_kohlhagen")
assert_close(garman_kohlhagen('c', 102, 100, 2, 0.05, 0.01, 0.25)[0], 18.63371484)
assert_close(garman_kohlhagen('p', 102, 100, 2, 0.05, 0.01, 0.25)[0], 9.13719197)
# Asian76(option_type, X, FS, T, TA, r, V):
print("Testing: Asian76")
assert_close(asian_76('c', 102, 100, 2, 1.9, 0.05, 0.25)[0], 13.53508930)
assert_close(asian_76('p', 102, 100, 2, 1.9, 0.05, 0.25)[0], 11.72541446)
# Kirks76(option_type, X, F1, F2, T, r, V1, V2, corr)
print("Testing: Kirks")
assert_close(kirks_76("c", f1=37.384913362, f2=42.1774, x=3.0, t=0.043055556, r=0, v1=0.608063, v2=0.608063, corr=.8)[0],0.007649192)
assert_close(kirks_76("p", f1=37.384913362, f2=42.1774, x=3.0, t=0.043055556, r=0, v1=0.608063, v2=0.608063, corr=.8)[0],7.80013583)
# ---------------------------
# Testing the external interface
if __name__ == "__main__":
print("=====================================")
print("External Interface Testing")
print("=====================================")
# BlackScholes(option_type, X, FS, T, r, V)
print("Testing: GBS.BlackScholes")
assert_close(black_scholes('c', 102, 100, 2, 0.05, 0.25)[0], 20.02128028)
assert_close(black_scholes('p', 102, 100, 2, 0.05, 0.25)[0], 8.50502208)
# Merton(option_type, X, FS, T, r, q, V)
print("Testing: GBS.Merton")
assert_close(merton('c', 102, 100, 2, 0.05, 0.01, 0.25)[0], 18.63371484)
assert_close(merton('p', 102, 100, 2, 0.05, 0.01, 0.25)[0], 9.13719197)
# Black76(option_type, X, FS, T, r, V)
print("Testing: GBS.Black76")
assert_close(black_76('c', 102, 100, 2, 0.05, 0.25)[0], 13.74803567)
assert_close(black_76('p', 102, 100, 2, 0.05, 0.25)[0], 11.93836083)
# garman_kohlhagen(option_type, X, FS, T, b, r, rf, V)
print("Testing: GBS.garman_kohlhagen")
assert_close(garman_kohlhagen('c', 102, 100, 2, 0.05, 0.01, 0.25)[0], 18.63371484)
assert_close(garman_kohlhagen('p', 102, 100, 2, 0.05, 0.01, 0.25)[0], 9.13719197)
# Asian76(option_type, X, FS, T, TA, r, V):
print("Testing: Asian76")
assert_close(asian_76('c', 102, 100, 2, 1.9, 0.05, 0.25)[0], 13.53508930)
assert_close(asian_76('p', 102, 100, 2, 1.9, 0.05, 0.25)[0], 11.72541446)
# Kirks76(option_type, X, F1, F2, T, r, V1, V2, corr)
print("Testing: Kirks")
assert_close(
kirks_76("c", f1=37.384913362, f2=42.1774, x=3.0, t=0.043055556, r=0, v1=0.608063, v2=0.608063, corr=.8)[0],
0.007649192)
assert_close(
kirks_76("p", f1=37.384913362, f2=42.1774, x=3.0, t=0.043055556, r=0, v1=0.608063, v2=0.608063, corr=.8)[0],
7.80013583)
```
## Benchmarking
This section benchmarks the output against output from a 3rd party option pricing libraries described in the book "The Complete Guide to Option Pricing Formulas" by Esper Haug.
*Haug, Esper. The Complete Guide to Option Pricing Formulas. McGraw-Hill 1997, pages 10-15*
Indexes for GBS Functions:
* [0] Value
* [1] Delta
* [2] Gamma
* [3] Theta (annualized, divide by 365 to get daily theta)
* [4] Vega
* [5] Rho
```
# ------------------
# Benchmarking against other option models
if __name__ == "__main__":
print("=====================================")
print("Selected Comparison to 3rd party models")
print("=====================================")
print("Testing GBS.BlackScholes")
assert_close(black_scholes('c', fs=60, x=65, t=0.25, r=0.08, v=0.30)[0], 2.13336844492)
print("Testing GBS.Merton")
assert_close(merton('p', fs=100, x=95, t=0.5, r=0.10, q=0.05, v=0.20)[0], 2.46478764676)
print("Testing GBS.Black76")
assert_close(black_76('c', fs=19, x=19, t=0.75, r=0.10, v=0.28)[0], 1.70105072524)
print("Testing GBS.garman_kohlhagen")
assert_close(garman_kohlhagen('c', fs=1.56, x=1.60, t=0.5, r=0.06, rf=0.08, v=0.12)[0], 0.0290992531494)
print("Testing Delta")
assert_close(black_76('c', fs=105, x=100, t=0.5, r=0.10, v=0.36)[1], 0.5946287)
assert_close(black_76('p', fs=105, x=100, t=0.5, r=0.10, v=0.36)[1], -0.356601)
print("Testing Gamma")
assert_close(black_scholes('c', fs=55, x=60, t=0.75, r=0.10, v=0.30)[2], 0.0278211604769)
assert_close(black_scholes('p', fs=55, x=60, t=0.75, r=0.10, v=0.30)[2], 0.0278211604769)
print("Testing Theta")
assert_close(merton('p', fs=430, x=405, t=0.0833, r=0.07, q=0.05, v=0.20)[3], -31.1923670565)
print("Testing Vega")
assert_close(black_scholes('c', fs=55, x=60, t=0.75, r=0.10, v=0.30)[4], 18.9357773496)
assert_close(black_scholes('p', fs=55, x=60, t=0.75, r=0.10, v=0.30)[4], 18.9357773496)
print("Testing Rho")
assert_close(black_scholes('c', fs=72, x=75, t=1, r=0.09, v=0.19)[5], 38.7325050173)
```
| github_jupyter |
# Data 620 Assignment: Document Classification
Jithendra Seneviratne, Sheryl Piechocki
July 3, 2020
### Import Modules and Libraries for Analysis
```
import string
import re
from nltk.corpus import wordnet
from nltk.tokenize import sent_tokenize, word_tokenize
from nltk.stem import WordNetLemmatizer
from nltk.tag import pos_tag
from nltk.corpus import stopwords
from sklearn.linear_model import LogisticRegression
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.metrics import confusion_matrix
from sklearn.model_selection import train_test_split
import pandas as pd
```
### Load Text
We'll be using text from a series of BBC articles found in [Kaggle](https://www.kaggle.com/yufengdev/bbc-fulltext-and-category?select=bbc-text.csv), categorized as one of the following:
* Tech
* Business
* Sport
* Entertainment
* Politics
The corpus consists of 2225 articles.
```
text_df = pd.read_csv('bbc-text.csv')
text_df.shape
text_df.head()
```
### Create list of stopwords
```
stop = stopwords.words('english') + ['mr',
'mrs',
'miss',
'say',
'have',
'might',
'thought',
'would',
'could',
'make',
'much',
'dear',
'must',
'know',
'one',
'good',
'every',
'towards',
'give',
'dr',
'none',
'go',
'come',
'upon',
'get',
'see',
'like',
'appear',
'sometimes',
'the',
'and',
'a',
'be',
'i',
'of',
'to',
'have',
'in',
'he',
'that',
'you',
'it',
'his',
'my',
'with',
'for',
'on',
'say',
'but',
'me',
'at',
'we',
'all',
'not',
'this',
'by',
'him',
'one',
'there',
'now',
'man',
'so',
'do',
'out',
'they',
'go',
'well',
'from',
'come',
'if',
'like',
'up',
'see',
'no',
'when',
'put',
'take',
'begin',
'two',
'three',
'u',
'still',
'last',
'never',
'always',
'thing',
'tell']
```
### Create Clean and Lemmatize Functions
```
def get_wordnet_pos(treebank_tag):
if treebank_tag.startswith('J'):
return wordnet.ADJ
elif treebank_tag.startswith('V'):
return wordnet.VERB
elif treebank_tag.startswith('N'):
return wordnet.NOUN
elif treebank_tag.startswith('R'):
return wordnet.ADV
else:
return 'n'
def lemmatize_word(word):
lemmatizer = WordNetLemmatizer()
try:
tag = get_wordnet_pos(pos_tag([word])[0][1])
return lemmatizer.lemmatize(word, pos=tag)
except:
pass
def clean_doc(doc):
line= re.sub('[%s]' % re.escape(string.punctuation), '', doc)
line = re.sub('[^a-zA-Z\ ]', '', line)
line = line.lower()
line = line.split()
line = ' '.join([lemmatize_word(x) for x in line if lemmatize_word(x) not in stop])
return line
```
### Apply cleaning and lemmatizing functions to corpus
We clean the corpus by removing punctuation, lemmatizing the words, and removing stop words.
```
text_df['cleaned_text'] = text_df['text'].apply(lambda x: clean_doc(x))
text_df.head()
```
### Function to vectorize corpus using TFIDF
The term frequency-inverse document frequency measure places importance on terms that are more frequent in a document, but are not frequent in all documents.
```
def vectorize_text(df,
maxdf=.5,
mindf=5,
ngram_range=(1, 1),
stop_words=stop):
tfidf_vectorizer = TfidfVectorizer(ngram_range=ngram_range,
max_df=maxdf,
min_df=mindf,
stop_words=stop_words)
tfidf = tfidf_vectorizer.fit_transform(text_df['cleaned_text'])
tfidf_feature_names = tfidf_vectorizer.get_feature_names()
df_transform = pd.DataFrame(tfidf.toarray())
df_transform.columns = tfidf_feature_names
df_transform['category'] = text_df['category']
return df_transform
```
### Apply transformation to corpus
```
df_transform = vectorize_text(text_df)
df_transform.head()
```
### Create logistic classifier function
```
def logistic_classifier(df):
X = df.drop(labels=['category'],
axis=1)
y = df['category']
lr = LogisticRegression(penalty='l2',
dual=False,
tol=.0001,
C=1,
)
X_train, X_test, y_train, y_test = train_test_split(X,
y,
test_size=.3)
lr.fit(X_train,y_train)
y_pred = lr.predict(X_test)
print('Test Accuracy','{:.1%}'.format(lr.score(X_test, y_test)))
print('Train Accuracy','{:.1%}'.format(lr.score(X_train, y_train)))
print('Confusion Matrix')
print(confusion_matrix(y_test, y_pred))
```
### Classify documents
```
logistic_classifier(df_transform)
```
Our model seems to have done very well, classifying test data with high accuracy. Let's see if we can do better by changing the ngram range.
### Change N-Gram Range
```
df_transform = vectorize_text(text_df,
ngram_range=(1, 2))
logistic_classifier(df_transform)
```
As we can see, changing the number of ngrams from a max of one word to two words has helped let's try changing it to three
```
df_transform = vectorize_text(text_df,
ngram_range=(1, 3))
logistic_classifier(df_transform)
```
As we can see, now the model is overfit
### Conclusions
| github_jupyter |
```
from laika.lib.coordinates import ecef2geodetic, geodetic2ecef
%pylab inline
import numpy as np
import seaborn as sns
# this is the main component of laika, and once initialized can
# immediately provide information about sattelites and signal delays
from laika import AstroDog
constellations = ['GPS', 'GLONASS']
dog = AstroDog(valid_const=constellations)
# For example if we want the position and speed of sattelite 7 (a GPS sat)
# at the start of January 7th 2018. Laika's custom GPSTime object is used throughout
# and can be initialized from python's datetime.
from datetime import datetime
from laika.gps_time import GPSTime
time = GPSTime.from_datetime(datetime(2018,1,7))
# We use RINEX3 PRNs to identify satellites
sat_prn = 'G07'
sat_pos, sat_vel, sat_clock_err, sat_clock_drift = dog.get_sat_info(sat_prn, time)
print("Sattelite's position in ecef(m) : \n", sat_pos, '\n')
print("Sattelite's velocity in ecef(m/s) : \n", sat_vel, '\n')
print("Sattelite's clock error(s) : \n", sat_clock_err, '\n\n')
# we can also get the pseudorange delay (tropo delay + iono delay + DCB correction)
# in the San Francisco area
receiver_position = [-2702584.60036925, -4325039.45362552, 3817393.16034817]
delay = dog.get_delay(sat_prn, time, receiver_position)
print("Sattelite's delay correction (m) in San Fransisco \n", delay)
# We can use these helpers to plot the orbits of sattelites
# by plotting the sattelite positions over the course
# of 12 hours, which is the approximate orbital period
# of GNSS sattelites.
from collections import defaultdict
import PIL
import matplotlib.pyplot as plt
import numpy as np
import matplotlib.patches as mpatches
from mpl_toolkits.mplot3d import Axes3D
from laika.constants import EARTH_RADIUS, EARTH_ROTATION_RATE
from laika.helpers import get_constellation
from laika.lib.orientation import rot_from_euler
# We start by plotting the world
# load bluemarble with PIL
bm = PIL.Image.open('bluemarble.jpg')
# it's big, so I'll rescale it, convert to array, and divide by 256 to get RGB values that matplotlib accept
bm = np.array(bm.resize([d//5 for d in bm.size]))/256.
lons = np.linspace(-180, 180, bm.shape[1]) * np.pi/180
lats = np.linspace(-90, 90, bm.shape[0])[::-1] * np.pi/180
fig = plt.figure(figsize=(14,14))
ax = fig.add_subplot(111, projection='3d')
x = EARTH_RADIUS*np.outer(np.cos(lons), np.cos(lats)).T
y = EARTH_RADIUS*np.outer(np.sin(lons), np.cos(lats)).T
z = EARTH_RADIUS*np.outer(np.ones(np.size(lons)), np.sin(lats)).T
ax.plot_surface(x, y, z, rstride=4, cstride=4, facecolors = bm)
ax.set_xlim(-2.5e7, 2.5e7)
ax.set_ylim(-2.5e7, 2.5e7)
ax.set_zlim(-2.5e7, 2.5e7)
# Now we get all the sattelite positions
# over a 12 hour period and plot them
sat_positions = defaultdict(lambda: [])
colors = {
'GPS': 'b',
'GLONASS': 'r'
}
for i in range(49):
dt = i*15*60
all_sat_info = dog.get_all_sat_info(time + dt)
for sat_prn, sat_info in all_sat_info.items():
positions = sat_positions[sat_prn]
theta = EARTH_ROTATION_RATE * dt
rot_matrix = rot_from_euler((0,0,theta))
positions.append(rot_matrix.dot(sat_info[0]))
for sat_prn, positions in sat_positions.items():
if len(positions) < 6:
continue
ax.plot([p[0] for p in positions],
[p[1] for p in positions],
[p[2] for p in positions],
color=colors[get_constellation(sat_prn)])
patches = [mpatches.Patch(color='r', label='GLONASS orbits'),
mpatches.Patch(color='b', label='GPS orbits')]
plt.legend(handles=patches, fontsize=14)
plt.title('GPS and GLONASS orbits', fontsize=25)
plt.show()
import laika.raw_gnss as raw
import laika.helpers as helpers
# this example data is the from the example segment
# of the comma2k19 dataset (research.comma.ai)
# example data contains an array of raw GNSS observables
# that were recorded during a minute of highway driving of
# a car, this array format can be used to create Laika's
# GNSSMeasurent object which can be processed with astrodog
# to then be analysed or used for position estimated.
with open('example_data/raw_gnss_ublox/value', 'br') as f:
example_data = np.load(f)
measurements = [raw.normal_meas_from_array(m_arr) for m_arr in example_data]
# lets limit this to GPS sattelite for the sake of simplicity
measurements = [m for m in measurements if helpers.get_constellation(m.prn) == 'GPS']
# we organize the measurements by epoch and by sattelite for easy plotting
measurements_by_epoch = raw.group_measurements_by_epoch(measurements)
measurements_by_sattelite = raw.group_measurements_by_sat(measurements)
# we can plot the amount of sattelites in view at each epoch
figsize(10,10)
plot([len(epoch) for epoch in measurements_by_epoch])
title('amount of sattelites in view', fontsize=25)
# we can plot the prr(pseudorange-rate) measured from the doppler shift
# towards every sattelite, we use the RINEX 3 convention to save
# the observables, so for doppler shifts on L1 its D1C.
figsize(10,10)
for prn in measurements_by_sattelite:
plot([m.recv_time.tow for m in measurements_by_sattelite[prn]],
[m.observables['D1C'] for m in measurements_by_sattelite[prn]],
label=prn)
legend()
title('PRR to for every sattelite (m/s)', fontsize=25)
from laika.constants import GPS_L1, SPEED_OF_LIGHT
# Or more intrestingly we can compare the measured carrier phase
# to a prediction of the carrier phase based on the previous carrier phase
# measurement incremented by the pseudorange_rate * timedelta
# With perfect measurements we expect this always to be around 0,
# we can see that this is often the case, but not always, indicating
# bad measurements
figsize(10,10)
for prn in measurements_by_sattelite:
predicted_vs_measured = []
for k in range(len(measurements_by_sattelite[prn])-1):
prev_meas = measurements_by_sattelite[prn][k-1]
current_meas = measurements_by_sattelite[prn][k]
avg_prr = np.mean([prev_meas.observables['D1C'], current_meas.observables['D1C']])
time_delta = (current_meas.recv_time - prev_meas.recv_time)
cp_increment = time_delta * avg_prr
cp_increment_cycles = cp_increment * (GPS_L1 / SPEED_OF_LIGHT)
cycle_meas_diff = current_meas.observables['L1C'] - prev_meas.observables['L1C']
predicted_vs_measured.append((cp_increment_cycles - cycle_meas_diff-.5)%1 - .5)
plot([m.recv_time.tow for m in measurements_by_sattelite[prn][1:]],
predicted_vs_measured,
label=prn)
legend()
title('Measured CP - predicted CP (L1 cycles)', fontsize=25)
# now we can do some fun stuff like get some basic position
# and speed estimates with Laika's simple WLS. First we process
# and correct measurement groups with our astrodog and then solve
# the GNSS trilateration problem
# to correct the measurements we need a receiver position estimate,
# this can be off by ~1000m, in this case we can get it by solving
# for the uncorrected measurements first.
pos_solutions, vel_solutions = [], []
corrected_measurements_by_epoch = []
for meas_epoch in measurements_by_epoch[::10]:
processed = raw.process_measurements(meas_epoch, dog)
est_pos = raw.calc_pos_fix(processed)[0][:3]
corrected = raw.correct_measurements(meas_epoch, est_pos, dog)
corrected_measurements_by_epoch.append(corrected)
pos_solutions.append(raw.calc_pos_fix(corrected))
# you need an estimate position to calculate a velocity fix
vel_solutions.append(raw.calc_vel_fix(corrected, pos_solutions[-1][0]))
# we can plot the ecef positions
figsize(10,10)
figure()
title('ECEF position estimate (m)', fontsize=25)
plot([sol[0][:3] for sol in pos_solutions])
# and the resiudal distribution of the WLS solutions
figure()
title('Residuals of pseudoranges in positions solutions (m)', fontsize=25)
sns.distplot(np.concatenate([sol[1] for sol in pos_solutions]), bins=arange(-5,5,.5),kde=False)
xlim(-5,5)
# we can plot the ecef velocities
figure()
figsize(10,10)
title('ECEF velocity estimate (m/s)', fontsize=25)
plot([sol[0][:3] for sol in vel_solutions])
# and the resiudal distribution of the WLS solutions
figure()
figsize(10,10)
title('Residuals of pseudorange-rates in velocity solutions (m/s)', fontsize=25)
sns.distplot(np.concatenate([sol[1] for sol in vel_solutions]), bins=arange(-2,2,.05),kde=False)
xlim(-2,2)
# The above residual plots give us an idea of how accurate our
# pseudoranges are that are used for the solution, but what does that
# actually mean for the accuracy of our position estimate? Lets look
# at the geometry of the situation before we procceed.
# lets plot the world again
# load bluemarble with PIL
bm = PIL.Image.open('bluemarble.jpg')
# it's big, so I'll rescale it, convert to array, and divide by 256 to get RGB values that matplotlib accept
bm = np.array(bm.resize([d//5 for d in bm.size]))//256.
# coordinates of the image - don't know if this is entirely accurate, but probably close
lons = np.linspace(-180, 180, bm.shape[1]) * np.pi/180
lats = np.linspace(-90, 90, bm.shape[0])[::-1] * np.pi/180
# repeat code from one of the examples linked to in the question, except for specifying facecolors:
fig = plt.figure(figsize=(14,14))
ax = fig.add_subplot(111, projection='3d')
x = EARTH_RADIUS*np.outer(np.cos(lons), np.cos(lats)).T
y = EARTH_RADIUS*np.outer(np.sin(lons), np.cos(lats)).T
z = EARTH_RADIUS*np.outer(np.ones(np.size(lons)), np.sin(lats)).T
ax.plot_surface(x, y, z, rstride=4, cstride=4, facecolors = bm)
ax.set_xlim(-2.5e7, 1e7)
ax.set_ylim(-2.5e7, 1e7)
ax.set_zlim(-1e7, 2.5e7)
# Lets plot the situation for the first epoch,
# it does not change significantly over our
# minute of example data.
receiver_pos = pos_solutions[0][0]
ax.scatter([receiver_pos[0]],
[receiver_pos[1]],
[receiver_pos[2]],
color='w',
#markersize=15,
marker='+')
#sattelite_positions
sat_positions = []
for m in corrected_measurements_by_epoch[0]:
sat_positions.append(m.sat_pos)
ax.plot([receiver_pos[0], m.sat_pos[0]],
[receiver_pos[1], m.sat_pos[1]],
[receiver_pos[2], m.sat_pos[2]],
color='b')
sat_positions = np.array(sat_positions)
ax.scatter(sat_positions[:,0],
sat_positions[:,1],
sat_positions[:,2],
color='k',
marker='^',
linewidths=6,
alpha=1)
ax.view_init(20,150)
plt.show()
from laika.lib.coordinates import LocalCoord
converter = LocalCoord.from_ecef(receiver_position)
sat_positions_ned = converter.ecef2ned(sat_positions)
azimuths_radians = np.arctan2(sat_positions_ned[:,0], sat_positions_ned[:,1])
elevation_radians = np.arctan2(-sat_positions_ned[:,2], np.linalg.norm(sat_positions_ned[:,:2],axis=1))
fig = plt.figure()
ax = fig.add_subplot(111, polar=True)
ax.scatter(azimuths_radians, elevation_radians*180/np.pi, marker='^', color='k', linewidth=6)
ax.set_yticks(range(0, 90+10, 10)) # Define the yticks
ax.set_theta_direction(-1)
ax.set_theta_zero_location('N')
yLabel = ['90', '', '', '60', '', '', '30', '', '', '']
ax.set_yticklabels(yLabel)
ax.set_title('Sattelite aximuth and elevation as seen by receiver', fontsize=25)
plt.show()
# The above plot highlights an important consideration of GNSS position
# estimation. That is, the concept of DOP (https://en.wikipedia.org/wiki/Dilution_of_precision_(navigation)).
# In short, it means that if the elevations and azimuths of the sattelites in view are simial, the DOP is high.
# A high DOP means that small errors in the pseudoranges can cause large errors in the computed position.
from laika.raw_gnss import get_DOP
print('GDOP of this epoch is ', get_DOP(receiver_position, sat_positions))
# This data clearly has a very good DOP
# By comparing the 4 sattelites with the highest elevation
# to those 4 with the lowest we can see how different sattelite
# positions can lead to very precisions
sat_positions_sorted = sat_positions[argsort(elevation_radians)]
print('GDOP of 4 lowest elevation sattelites of this epoch is ', get_DOP(receiver_position, sat_positions_sorted[-4:]))
print('GDOP of 4 highest elevation sattelites of this epoch is ', get_DOP(receiver_position, sat_positions_sorted[:4]))
```
| github_jupyter |
**Executed:** Sat Mar 18 01:17:33 2017
**Duration:** 4 seconds.
**Autogenerated from:** [./param-learning-template.ipynb](./param-learning-template.ipynb)
```
# default values of file paths, assuming quantum_fog is working dir
in_bif = 'examples_cbnets/earthquake.bif'
in_dot = 'examples_cbnets/earthquake.dot'
in_csv = 'learning/training_data_c/earthquake.csv'
qfog_path = None
# Cell inserted during automated execution.
in_bif = 'examples_cbnets/earthquake.bif'
in_dot = 'examples_cbnets/earthquake.dot'
in_csv = 'learning/training_data_c/earthquake.csv'
qfog_path = '/home/jupyter/Notebooks/Quantum/quantum-fog'
```
**Parameter Learning Template**
# Table of Contents
<p><div class="lev1 toc-item"><a href="#Learned-Network-Parameters" data-toc-modified-id="Learned-Network-Parameters-1"><span class="toc-item-num">1 </span>Learned Network Parameters</a></div><div class="lev2 toc-item"><a href="#QFog" data-toc-modified-id="QFog-11"><span class="toc-item-num">1.1 </span>QFog</a></div><div class="lev2 toc-item"><a href="#bnlearn" data-toc-modified-id="bnlearn-12"><span class="toc-item-num">1.2 </span>bnlearn</a></div><div class="lev2 toc-item"><a href="#Summary-of-Running-Times" data-toc-modified-id="Summary-of-Running-Times-13"><span class="toc-item-num">1.3 </span>Summary of Running Times</a></div>
```
import pandas as pd
import numpy as np
from graphviz import Source
import warnings
warnings.filterwarnings("ignore", module="rpy2")
import rpy2
%load_ext rpy2.ipython
%R library("bnlearn");
%R library("Rgraphviz");
import os
import sys
if not qfog_path:
os.chdir('../../')
else:
os.chdir(qfog_path)
cwd = os.getcwd()
sys.path.insert(0,cwd)
print("cwd=", cwd)
from learning.NetParamsLner import *
```
Read in_csv, create Pandas Dataframe with it, push dataframe into R
```
states_df = pd.read_csv(in_csv)
states_df.head()
states_df.tail()
%Rpush states_df
%R str(states_df)
```
For bnlearn, data.frame columns cannot be int type, must be changed to factor.
data.frame is a list so can use lapply
```
%R states_df[] <- lapply(states_df, factor)
%R str(states_df)
```
# Learned Network Parameters
## QFog
```
# emp = empirical, learned
is_quantum = False
bnet = BayesNet.read_bif(in_bif, is_quantum)
bnet_emp = BayesNet.read_dot(in_dot)
vtx_to_states = bnet.get_vtx_to_state_names()
bnet_emp.import_nd_state_names(vtx_to_states)
lnr = NetParamsLner(is_quantum, bnet_emp, states_df)
%%capture qfog_params_time
%time lnr.learn_all_bnet_pots()
print(qfog_params_time)
lnr.compare_true_and_emp_pots(bnet, bnet_emp)
```
## bnlearn
```
%Rpush in_bif
%R bn.fit = read.bif(in_bif)
%R str(bn.fit)
%R bn = bn.net(bn.fit)
%R str(bn)
%%capture bnlearn_params_time
%time %R rfit = bn.fit(bn, data = states_df);
print(bnlearn_params_time)
%R str(rfit)
%%R
j = 0
for( nd in nodes(bn.fit)){
j = j + 1
cat('----------------', nd[1], '\n')
cat('true:\n')
print(coef(bn.fit)[j])
cat('empirical:\n')
print(coef(rfit)[j])
}
```
## Summary of Running Times
```
print("QFog:\n", qfog_params_time)
print("bnlearn:\n", bnlearn_params_time)
```
| github_jupyter |
# Introduction to NumPy
NumPy is one of the two most important libraries in Python for data science, along with pandas. NumPy is a crucial library for effectively loading, storing, and manipulating in-memory data in Python, which will be at the heart of what you do with data science in Python.
Datasets come from a huge range of sources and in a wide range of formats, such as text documents, images, sound clips, numerical measurements, and nearly anything else. Despite this variety, however, the start of data science is to think of all data fundamentally as arrays of numbers.
For example, the words in documents can be represented as the numbers that encode letters in computers or even the frequency of particular words in a collection of documents. Digital images can be thought of as two-dimensional arrays of numbers representing pixel brightness or color. Sound files can be represented as one-dimensional arrays of frequency versus time. However, no matter what form our data takes, to analyze it, our first step will be to transform it into arrays of numbers--which is where NumPy comes in (and pandas down the road).
NumPy is short for *Numerical Python*, and it provides an efficient means of storing and operating on dense data buffers in Python. Array-oriented computing in Python goes back to 1995 with the Numeric library. Scientific programming in Python took off over the next 10 years, but the collections of libraries splintered. The NumPy project began in 2005 as a means of bringing the Numeric and NumArray projects together around a single array-based framework.
Some examples in this section are drawn from the *Python Data Science Handbook* by Jake VanderPlas (content available [on GitHub](https://github.com/jakevdp/PythonDataScienceHandbook)) and *Python for Data Analysis* by Wes McKinney. Text from the *Python Data Science Handbook* is released under the [CC-BY-NC-ND license](https://creativecommons.org/licenses/by-nc-nd/3.0/us/legalcode); code is released under the [MIT license](https://opensource.org/licenses/MIT).
Let's get started exploring NumPy! Our first step will be to import NumPy using `np` as an alias:
```
import numpy as np
```
Get used to this convention; it's a common convention in Python, and it's the way we will use and refer to NumPy throughout the rest of this course.
## Built-In Help
There's a lot to learn about NumPy, and it can be tough to remember it all the first time through. Don't worry! IPython, the underlying program that enables notebooks like this one to interact with Python, has you covered.
First off, IPython gives you the ability to quickly explore the contents of a package like NumPy by using the tab-completion feature. So, if you want to see all the functions available with NumPy, type this:
```ipython
In [2]: np.<TAB>
```
When you do so, a drop-down menu will appear next to the `np.`
### Exercise
```
# Place your cursor after the period and press <TAB>:
np.
```
From the drop-down menu, you can select any function to run. Better still, you can select any function and view the built-in help documentation for it. For example, to view the documentation for the NumPy `add()` function, you can run this code:
```ipython
In [3]: np.add?
```
Try this with a few different functions. Remember, these functions are just like ones you wrote in Section 2; the documentation will help explain what parameters you can (or should) provide the function, in addition to what output you can expect.
### Exercise
```
# Replace 'add' below with a few different NumPy function names and look over the documentation:
np.add?
```
For more detailed documentation, along with additional tutorials and other resources, visit [www.numpy.org](http://www.numpy.org).
Now that you know how to quickly get help while you are working on your own, let's return to storing data in arrays.
## NumPy arrays: a specialized data structure for analysis
> **Learning goal:** By the end of this subsection, you should have a basic understanding of what NumPy arrays are and how they differ from the other Python data structures you have studied thus far.
We started the discussion in this section by noting that data science starts by representing data as arrays of numbers.
"Wait!" you might be thinking. "Can't we just use Python lists for that?"
Depending on the data, yes, you could (and you will use lists as a part of working with data in Python). But to see what we might want to use a specialized data structure for, let's look a little more closely at lists.
### Lists in Python
Python lists can hold just one kind of object. Let's use one to create a list of just integers:
```
myList = list(range(10))
myList
```
Remember list comprehension? We can use it to probe the data types of items in a list:
```
[type(item) for item in myList]
```
Of course, a really handy feature of Python lists is that they can hold heterogeneous types of data in a single list object:
```
myList2 = [True, "2", 3.0, 4]
[type(item) for item in myList2]
```
However, this flexibility comes at a price. Each item in a list is really a separate Python object (the list is an object itself, but mostly it is an object that serves as a container for the memory pointers to the constituent objects). That means that each item in a list must contain its own type info, reference count, and other information. This information can become expensive in terms of memory and performance if we are dealing with hundreds of thousands or millions of items in a list. Moreover, for many uses in data science, our arrays just store a single type of data (such as integers or floats), which means that the object-related information for items in such an array would be redundant. It can be much more efficient to store data in a fixed-type array.
<img align="left" style="padding-right:10px;" src="https://raw.githubusercontent.com/microsoft/computerscience/master/Educator%20Resources/Reactor%20Workshops/Data%20Science/Track%201/Graphics/Sec3_array_vs_list.png">
Enter the fixed-type, NumPy-style array.
### Fixed-type arrays in Python
At the level of implementation by the computer, the `ndarray` that is part of the NumPy package contains a single pointer to one contiguous block of data. This is efficient, both memory-wise and computationally. Better still, NumPy provides efficient *operations* on data stored in `ndarray` objects.
(Note that we will use “array,” “NumPy array,” and “ndarray” interchangeably throughout this section to refer to the ndarray object.)
#### Creating NumPy arrays method 1: using Python lists
There are multiple ways to create arrays in NumPy. Let's start by using our familiar Python lists. We will use the `np.array()` function to do this (remember, we imported NumPy as '`np`'):
```
# Create an integer array:
np.array([1, 4, 2, 5, 3])
```
Remember that, unlike Python lists, NumPy constrains arrays to contain a single type. So, if data types fed into a NumPy array do not match, NumPy will attempt to *upcast* them if possible. To see what we mean, here NumPy upcasts integers to floats:
```
np.array([3.14, 4, 2, 3])
```
### Exercise
```
# What happens if you construct an array using a list that contains a combination of integers, floats, and strings?
```
If you want to explicitly set the data type of your array when you create it, you can use the `dtype` keyword:
```
np.array([1, 2, 3, 4], dtype='float32')
```
### Exercise
```
# Try this using a different dtype.
# Remember that you can always refer to the documentation with the command np.array.
```
Most usefully for many applications in data science, NumPy arrays can explicitly be multidimensional (like matrices or tensors). Here's one way of creating a multidimensional array using a list of lists:
```
# nested lists result in multi-dimensional arrays
np.array([range(i, i + 3) for i in [2, 4, 6]])
```
The inner lists in a list of lists are treated as rows of the two-dimensional array you created.
#### Creating NumPy arrays method 2: building from scratch
In practice, it is often more efficient to create arrays from scratch using functions built into NumPy, particularly for larger arrays. Here are a few examples; these examples will help introduce you to several useful NumPy functions.
```
# Create an integer array of length 10 filled with zeros
np.zeros(10, dtype=int)
# Create a 3x5 floating-point array filled with ones
np.ones((3, 5), dtype=float)
# Create a 3x5 array filled with 3.14
# The first number in the tuple gives the number of rows
# The second number in the tuple sets the number of columns
np.full((3, 5), 3.14)
# Create an array filled with a linear sequence
# Starting at 0, ending at 20, stepping by 2
# (this is similar to the built-in Python range() function)
np.arange(0, 20, 2)
# Create an array of five values evenly spaced between 0 and 1
np.linspace(0, 1, 5)
# Create a 3x3 array of uniformly distributed
# random values between 0 and 1
np.random.random((3, 3))
# Create a 3x3 array of normally distributed random values
# with mean 0 and standard deviation 1
np.random.normal(0, 1, (3, 3))
# Create a 3x3 array of random integers in the interval [0, 10)
np.random.randint(0, 10, (3, 3))
# Create a 3x3 identity matrix
np.eye(3)
# Create an uninitialized array of three integers
# The values will be whatever happens to already exist at that memory location
np.empty(3)
```
Now take a couple of minutes to go back and play with these code snippets, changing the parameters. These functions are the essence of creating NumPy arrays and you will want to become comfortable with them.
Below is a table listing several of the array-creation functions in NumPy.
| Function | Description |
|:--------------|:------------|
| `array` | Converts input data (list, tuple, array, or other sequence type) to an ndarray either |
| | by inferring a dtype or explicitly specifying a dtype. Copies the input data by default. |
| `asarray` | Converts input to ndarray, but does not copy if the input is already an ndarray. |
| `arange` | Similar to the built-in `range()` function but returns an ndarray instead of a list. |
| `ones`, `ones_like` | Produces an array of all 1s with the given shape and dtype. |
| | `ones_like` takes another array and produces a ones-array of the same shape and dtype. |
| `zeros`, `zeros_like` | Similar to `ones` and `ones_like` but producing arrays of 0s instead. |
| `empty`, `empty_like` | Creates new arrays by allocating new memory, but does not populate with any values
| | like `ones` and `zeros`. |
| `full`, `full_like` | Produces an array of the given shape and dtype with all values set to the indicated “fill value.” |
| | `full_like` takes another array and produces a a filled array of the same shape and dtype. |
| `eye`, `identity` | Create a square $N \times N$ identity matrix (1s on the diagonal and 0s elsewhere) |
### NumPy data types
The standard NumPy data types are listed in the following table. Note that when constructing an array, they can be specified using a string:
```python
np.zeros(8, dtype='int16')
```
Or they can be specified directly using the NumPy object:
```python
np.zeros(8, dtype=np.int16)
```
| Data type | Description |
|:--------------|:------------|
| ``bool_`` | Boolean (True or False) stored as a byte |
| ``int_`` | Default integer type (same as C ``long``; normally either ``int64`` or ``int32``)|
| ``intc`` | Identical to C ``int`` (normally ``int32`` or ``int64``)|
| ``intp`` | Integer used for indexing (same as C ``ssize_t``; normally either ``int32`` or ``int64``)|
| ``int8`` | Byte (-128 to 127)|
| ``int16`` | Integer (-32768 to 32767)|
| ``int32`` | Integer (-2147483648 to 2147483647)|
| ``int64`` | Integer (-9223372036854775808 to 9223372036854775807)|
| ``uint8`` | Unsigned integer (0 to 255)|
| ``uint16`` | Unsigned integer (0 to 65535)|
| ``uint32`` | Unsigned integer (0 to 4294967295)|
| ``uint64`` | Unsigned integer (0 to 18446744073709551615)|
| ``float_`` | Shorthand for ``float64``.|
| ``float16`` | Half-precision float: sign bit, 5 bits exponent, 10 bits mantissa|
| ``float32`` | Single-precision float: sign bit, 8 bits exponent, 23 bits mantissa|
| ``float64`` | Double-precision float: sign bit, 11 bits exponent, 52 bits mantissa|
| ``complex_`` | Shorthand for ``complex128``.|
| ``complex64`` | Complex number, represented by two 32-bit floats|
| ``complex128``| Complex number, represented by two 64-bit floats|
If these data types seem similar to those in C, that's because NumPy is built in C.
> **Takeaway:** NumPy arrays are a data structure similar to Python lists that provide high performance when storing and working on large amounts of homogeneous data--precisely the kind of data that you will encounter frequently in data science. NumPy arrays support many data types beyond those discussed in this course. With that said, however, don’t worry about memorizing all the NumPy dtypes. **It’s often just necessary to care about the general kind of data you’re dealing with: floating point, integer, Boolean, string, or general Python object.**
## Working with NumPy arrays: the basics
> **Learning goal:** By the end of this subsection, you should be comfortable working with NumPy arrays in basic ways.
Now that you know how to create arrays in NumPy, you need to get comfortable manipulating them for two reasons. First, you will work with NumPy arrays as part of your exploration of data science. Second, our other important Python data science tool, pandas, is actually built around NumPy. Getting good at working with NumPy arrays will pay dividends in the next section (Section 4) and beyond: NumPy arrays are the building blocks for the `Series` and `DataFrame` data structures in the Python pandas library and you will use them *a lot* in data science. To get comfortable with array manipulation, we will cover five specifics:
- **Arrays attributes**: Assessing the size, shape, and data types of arrays
- **Indexing arrays**: Getting and setting the value of individual array elements
- **Slicing arrays**: Getting and setting smaller subarrays within a larger array
- **Reshaping arrays**: Changing the shape of a given array
- **Joining and splitting arrays**: Combining multiple arrays into one and splitting one array into multiple arrays
### Array attributes
First, let's look at some array attributes. We'll start by defining three arrays filled with random numbers: one one-dimensional, another two-dimensional, and the last three-dimensional. Because we will be using NumPy's random number generator, we will set a *seed* value to ensure that you get the same random arrays each time you run this code:
```
import numpy as np
np.random.seed(0) # seed for reproducibility
a1 = np.random.randint(10, size=6) # One-dimensional array
a2 = np.random.randint(10, size=(3, 4)) # Two-dimensional array
a3 = np.random.randint(10, size=(3, 4, 5)) # Three-dimensional array
```
Each array has attributes ``ndim`` (the number of dimensions of an array), ``shape`` (the size of each dimension of an array), and ``size`` (the total number of elements in an array).
### Exercise:
```
# Change the values in this code snippet to look at the attributes for a1, a2, and a3:
print("a3 ndim: ", a3.ndim)
print("a3 shape:", a3.shape)
print("a3 size: ", a3.size)
```
Another useful array attribute is the `dtype`, which we already encountered earlier in this section as a means of determining the type of data in an array:
```
print("dtype:", a3.dtype)
```
### Exercise:
```
# Explore the dtype for the other arrays.
# What dtypes do you predict them to have?
print("dtype:", a3.dtype)
```
### Indexing arrays
Indexing in NumPy is similar to indexing lists in standard Python. In fact, indices in one-dimensional arrays work exactly as they do with Python lists:
```
a1
a1[0]
a1[4]
```
As with regular Python lists, to index from the end of the array you can use negative indices:
```
a1[-1]
a1[-2]
```
### Exercise:
```
# Do multidimensional NumPy arrays work like Python lists of lists?
# Try a few combinations like a2[1][1] or a3[0][2][1] and see what comes back
```
You might have noticed that we can treat multidimensional arrays like lists of lists. But a more common means of accessing items in multidimensional arrays is to use a comma-separated tuple of indices.
(Yes, we realize that these comma-separated tuples use square brackets rather than the parentheses the name might suggest, but they are nevertheless referred to as tuples.)
```
a2
a2[0, 0]
a2[2, 0]
a2[2, -1]
```
You can also modify values by use of this same comma-separated index notation:
```
a2[0, 0] = 12
a2
```
Remember, once defined, NumPy arrays have a fixed data type. So, if you attempt to insert a float into an integer array, the value will be silently truncated.
```
a1[0] = 3.14159
a1
```
### Exercise:
```
# What happens if you try to insert a string into a1?
# Hint: try both a string like '3' and one like 'three'
```
### Slicing arrays
Similar to how you can use square brackets to access individual array elements, you can also use them to access subarrays. You do this with the *slice* notation, marked by the colon (`:`) character. NumPy slicing syntax follows that of the standard Python list; so, to access a slice of an array `a`, use this notation:
``` python
a[start:stop:step]
```
If any of these are unspecified, they default to the values ``start=0``, ``stop=``*``size of dimension``*, ``step=1``.
Let's take a look at accessing subarrays in one dimension and in multiple dimensions.
#### One-dimensional slices
```
a = np.arange(10)
a
a[:5] # first five elements
a[5:] # elements after index 5
a[4:7] # middle sub-array
a[::2] # every other element
a[1::2] # every other element, starting at index 1
```
### Exercise:
```
# How would you access the *last* five elements of array a?
# How about every other element of the last five elements of a?
# Hint: Think back to list indexing in Python
```
Be careful when using negative values for ``step``. When ``step`` has a negative value, the defaults for ``start`` and ``stop`` are swapped and you can use this functionality to reverse an array:
```
a[::-1] # all elements, reversed
a[5::-2] # reversed every other from index 5
```
### Exercise:
```
# How can you create a slice that contains every third element of a
# descending from the second-to-last element to the second element of a?
```
#### Multidimensional slices
Multidimensional slices use the same slice notation of one-dimensional subarrays mixed with the comma-separated notation of multidimensional arrays. Some examples will help illustrate this.
```
a2
a2[:2, :3] # two rows, three columns
a2[:3, ::2] # all rows, every other column
```
Finally, subarray dimensions can even be reversed together:
```
a2[::-1, ::-1]
```
### Exercise:
```
# Now try to show 2 rows and 4 columns with every other element?
```
#### Accessing array rows and columns
One thing you will often need to do in manipulating data is accessing a single row or column in an array. You can do this through a combination of indexing and slicing--specifically by using an empty slice marked by a single colon (``:``). Again, some examples will help illustrate this.
```
print(a2[:, 0]) # first column of x2
print(a2[0, :]) # first row of x2
```
In the case of row access, the empty slice can be omitted for a more compact syntax:
```
print(a2[0]) # equivalent to a2[0, :]
```
### Exercise:
```
# How would you access the third column of a3?
# How about the third row of a3?
```
#### Slices are no-copy views
It's important to know that slicing produces *views* of array data, not *copies*. This is a **huge** difference between NumPy array slicing and Python list slicing. With Python lists, slices are only shallow copies of lists; if you modify a copy, it doesn't affect the parent list. When you modify a NumPy subarray, you modify the original list. Be careful: this can have ramifications when you are trying to work with a small part of a large dataset and you don’t want to change the whole thing. Let's look more closely.
```
print(a2)
```
Extract a $2 \times 2$ subarray from `a2`:
```
a2_sub = a2[:2, :2]
print(a2_sub)
```
Now modify this subarray:
```
a2_sub[0, 0] = 99
print(a2_sub)
```
`a2` is now modified as well:
```
print(a2)
```
### Exercise:
```
# Now try reversing the column and row order of a2_sub
# Does a2 look the way you expected it would after that manipulation?
```
The fact that slicing produces views rather than copies is useful for data science work. As you work with large datasets, you will often find that it is easier to access and manipulate pieces of those datasets rather than copying them entirely.
#### Copying arrays
Instead of just creating views, sometimes it is necessary to copy the data in one array to another. When you need to do this, use the `copy()` method:
```
a2_sub_copy = a2[:2, :2].copy()
print(a2_sub_copy)
```
If we now modify this subarray, the original array is not touched:
```
a2_sub_copy[0, 0] = 42
print(a2_sub_copy)
print(a2)
```
### Reshaping arrays
Another way you will need to manipulate arrays is by reshaping them. This involves changing the number and size of dimensions of an array. This kind of manipulation can be important in getting your data to meet the expectations of machine learning programs or APIs.
The most flexible way of doing this kind of manipulation is with the `reshape` method. For example, if you want to put the numbers 1 through 9 in a $3 \times 3$ grid, you can do the following:
```
grid = np.arange(1, 10).reshape((3, 3))
print(grid)
```
Another common manipulation you will do in data science is converting one-dimensional arrays into two-dimensional row or column matrices. This can be a common necessity when doing linear algebra for machine learning. While you can do this by means of the `reshape` method, an easier way is to use the `newaxis` keyword in a slice operation:
```
a = np.array([1, 2, 3])
# row vector via reshape
a.reshape((1, 3))
# row vector via newaxis
a[np.newaxis, :]
# column vector via reshape
a.reshape((3, 1))
# column vector via newaxis
a[:, np.newaxis]
```
You will see this type of transformation a lot in the remainder of this course.
### Joining and splitting arrays
Another common data-manipulation need in data science is combining multiple datasets. First learning how to do this with NumPy arrays will help you in the next section (Section 4) when we do this with more complex data structures. You may often also need to split a single array into multiple arrays.
#### Joining arrays
To join arrays in NumPy, you will most often use `np.concatenate`, which is the method we will cover here. In the future, if you find yourself needing to specifically join arrays in mixed dimensions (a rarer case), read the documentation on `np.vstack`, `np.hstack`, and `np.dstack`.
##### `np.concatenate()`
`np.concatenate` takes a tuple or list of arrays as its first argument:
```
a = np.array([1, 2, 3])
b = np.array([3, 2, 1])
np.concatenate([a, b])
```
You can also concatenate more than two arrays at once:
```
c = [99, 99, 99]
print(np.concatenate([a, b, c]))
```
`np.concatenate` can also be used for two-dimensional arrays:
```
grid = np.array([[1, 2, 3],
[4, 5, 6]])
# concatenate along the first axis, which is the default
np.concatenate([grid, grid])
```
### Exercise:
```
# Recall that axes are zero-indexed in NumPy.
# What do you predict np.concatenate([grid, grid], axis=1) will produce?
```
#### Splitting arrays
To split arrays into multiple smaller arrays, you can use the functions ``np.split``, ``np.hsplit``, ``np.vsplit``, and ``np.dsplit``. As above, we will only cover the most commonly-used function (`np.split`) in this course.
##### `np.split()`
Let's first examine the case of a one-dimensional array:
```
a = [1, 2, 3, 99, 99, 3, 2, 1]
a1, a2, a3 = np.split(a, [3, 5])
print(a1, a2, a3)
```
Notice that *N* split-points produces to *N + 1* subarrays. In this case, it has formed the subarray `a2` with `a[3]` and `a[4]` (the element just before position 5 [remember how Python indexing goes], the second input in the tuple) as elements. `a1` and `a3` pick up the leftover portions from the original array `a`.
### Exercise:
```
grid = np.arange(16).reshape((4, 4))
grid
# What does np.split(grid, [1, 2]) produce?
# What about np.split(grid, [1, 2], axis=1)?
```
> **Takeaway:** Manipulating datasets is a fundamental part of preparing data for analysis. The skills you learned and practiced here will form building blocks for the most sophisticated data manipulation you will learn in later sections in this course.
## Fancy indexing
So far, we have explored how to access and modify portions of arrays using simple indices like `arr[0]`) and slices like `arr[:5]`. Now it is time for fancy indexing, in which we pass an array of indices to an array to access or modify multiple array elements at the same time.
Let's try it out:
```
rand = np.random.RandomState(42)
arr = rand.randint(100, size=10)
print(arr)
```
Suppose you need to access three different elements. Using the tools you currently have, your code might look something like this:
```
[arr[3], arr[7], arr[2]]
```
With fancy indexing, you can pass a single list or array of indices to do the same thing:
```
ind = [3, 7, 4]
arr[ind]
```
Another useful aspect of fancy indexing is that the shape of the output array reflects the shape of the *index arrays* you supply, rather than the shape of the array you are accessing. This is handy because there will be many times in a data scientist's life when they want to grab data from an array in a particular manner, such as to pass it to a machine learning API. Let's examine this property with an example:
```
ind = np.array([[3, 7],
[4, 5]])
arr[ind]
```
`arr` is a one-dimensional array, but `ind`, your index array, is a $2 \times 2$ array, and that is the shape the results comes back in.
### Exercise:
```
# What happens when your index array is bigger than the target array?
# Hint: you could use a large one-dimensional array or something fancier like ind = np.arange(0, 12).reshape((6, 2))
```
Fancy indexing also works in multiple dimensions:
```
arr2 = np.arange(12).reshape((3, 4))
arr2
```
As with standard indexing, the first index refers to the row and the second to the column:
```
row = np.array([0, 1, 2])
col = np.array([2, 1, 3])
arr2[row, col]
```
What did you actually get as your final result? The first value in the result array is `arr2[0, 2]`, the second one is `arr2[1, 1]`, and the third one is `arr2[2, 3]`.
The pairing of indices in fancy indexing follows all the same broadcasting rules we covered earlier. Thus, if you combine a column vector and a row vector within the indices, you get a two-dimensional result:
```
arr2[row[:, np.newaxis], col]
```
Here, each row value is matched with each column vector, exactly as we saw in broadcasting of arithmetic operations.
### Exercise:
```
# Now try broadcasting this on your own.
# What do you get with row[:, np.newaxis] * col?
# Or row[:, np.newaxis] * row? col[:, np.newaxis] * row?
# What about col[:, np.newaxis] * row?
# Hint: think back to the broadcast rules
```
**The big takeaway:** It is always important to remember that fancy indexing returns values reflected by the *broadcasted shape of the indices*, and not the shape of the array being indexed.
### Combined indexing
You can also combine fancy indexing with the other indexing schemes you have learned. Consider `arr2` again:
```
print(arr2)
```
Now combine fancy and simple indices:
```
arr2[2, [2, 0, 1]]
```
What did you get back? The elements at positions 2, 0, and 1 of row 2 (the third row).
You can also combine fancy indexing with slicing:
```
arr2[1:, [2, 0, 1]]
```
Again, consider what you got back as output: the elements at positions 2, 0, and 1 of each row after the first one (so the second and third rows).
Of course, you can also combine fancy indexing with masking:
```
mask = np.array([1, 0, 1, 0], dtype=bool)
arr2[row[:, np.newaxis], mask]
```
### Modifying values using fancy indexing
Fancy indexing is, of course, not just for accessing parts of an array, but also for modifying parts of an array:
```
ind = np.arange(10)
arr = np.array([2, 1, 8, 4])
ind[arr] = 99
print(ind)
```
You can also use a ufunc here and subtract 10 from each element of the array:
```
ind[arr] -= 10
print(ind)
```
Be cautious when using repeated indices with operations like these. They might not always produce the results you expect. For example:
```
ind = np.zeros(10)
ind[[0, 0]] = [4, 6]
print(ind)
```
Where did the 4 go? The result of this operation is to first assign `ind[0] = 4`, followed by `ind[0] = 6`. So the result is that `ind[0]` contains the value 6.
But not every operation repeats the way you think it should:
```
arr = [2, 3, 3, 4, 4, 4]
ind[arr] += 1
ind
```
We might have expected that `ind[3]` would contain the value 2 and `ind[4]` would contain the value 3. After all, that is how many times each index is repeated. So what happened?
This happened because `ind[arr] += 1` is shorthand for `ind[arr] = ind[arr] + 1`. `ind[arr] + 1` is evaluated, and then the result is assigned to the indices in `ind`. So, similar to the previous example, this is not augmentation that happens multiple times, but an assignment, which can lead to potentially counterintuitive results.
But what if you want an operation to repeat? To do this, use the `at()` method of ufuncs:
```
ind = np.zeros(10)
np.add.at(ind, arr, 1)
print(ind)
```
### Exercise:
```
# What does np.subtract.at(ind, arr, 1) give you?
# Play around with some of the other ufuncs we have seen.
```
> **Takeaway:** Fancy indexing enables you to select and manipulate several array members at once. This type of programmatic data manipulation is common in data science; often what you want to do with your data you want to do on several data points at once.
## Sorting arrays
So far we have just worried about accessing and modifying NumPy arrays. Another huge thing you will need to do as a data scientist is sort array data. Sorting is often an important means of teasing out the structure in data (such as outlying data points).
Although you could use Python's built-in `sort` and `sorted` functions, they will not work nearly as efficiently as NumPy's `np.sort` function.
`np.sort` returns a sorted version of an array without modifying the input:
```
a = np.array([2, 1, 4, 3, 5])
np.sort(a)
```
To sort the array in-place, use the `sort` method directly on arrays:
```
a.sort()
print(a)
```
A related function is `argsort`, which returns the *indices* of the sorted elements rather than the elements themselves:
```
a = np.array([2, 1, 4, 3, 5])
b = np.argsort(a)
print(b)
```
The first element of this result gives the index of the smallest element, the second value gives the index of the second smallest, and so on. These indices can then be used (via fancy indexing) to reconstruct the sorted array:
```
a[b]
```
### Sorting along rows or columns
A useful feature of NumPy's sorting algorithms is the ability to sort along specific rows or columns of a multidimensional array using the `axis` argument. For example:
```
rand = np.random.RandomState(42)
table = rand.randint(0, 10, (4, 6))
print(table)
# Sort each column of the table
np.sort(table, axis=0)
# Sort each row of the table
np.sort(table, axis=1)
```
Bear in mind that this treats each row or column as an independent array; any relationships between the row or column values will be lost doing this kind of sorting.
## Partial sorting: partitioning
Sometimes you don't need to sort an entire array, you just need to find the *k* smallest values in the array (often when looking at the distance of data points from one another). NumPy supplies this functionality through the `np.partition` function. `np.partition` takes an array and a number *k*; the result is a new array with the smallest *k* values to the left of the partition and the remaining values to the right (in arbitrary order):
```
arr = np.array([7, 2, 3, 1, 6, 5, 4])
np.partition(arr, 3)
```
Note that the first three values in the resulting array are the three smallest in the array, and the remaining array positions contain the remaining values. Within the two partitions, the elements have arbitrary order.
Similarly to sorting, we can partition along an arbitrary axis of a multidimensional array:
```
np.partition(table, 2, axis=1)
```
The result is an array where the first two slots in each row contain the smallest values from that row, with the remaining values filling the remaining slots.
Finally, just as there is an `np.argsort` that computes indices of the sort, there is an `np.argpartition` that computes indices of the partition. We'll see this in action in the following section when we discuss pandas.
> **Takeaway:** Sorting your data is a fundamental means of exploring and answering questions about it. The sorting algorithms in NumPy provide you with a fast, computationally efficient way of doing this on large amounts of data and with fine grain control.
## Efficient computation on NumPy arrays: Universal functions
> **Learning goal:** By the end of this subsection, you should have a basic understanding of what NumPy universal functions are and how (and why) to use them.
Some of the properties that make Python great to work with for data science (its dynamic, interpreted nature, for example) can also make it slow. This is particularly true with looping. These small performance hits can add up to minutes (or longer) when dealing with truly huge datasets.
When we first examined loops in Introduction to Python, you probably didn't notice any delay: the loops were short enough that Python’s relatively slow looping wasn’t an issue. Consider this function, which calculates the reciprocal for an array of numbers:
```
import numpy as np
np.random.seed(0)
def compute_reciprocals(values):
output = np.empty(len(values))
for i in range(len(values)):
output[i] = 1.0 / values[i]
return output
values = np.random.randint(1, 10, size=5)
compute_reciprocals(values)
```
Running this loop, it was probably difficult to even discern that execution wasn't instantaneous.
But let’s try it on a much larger array. To empirically do this, we'll time this with IPython's `%timeit` magic command.
```
big_array = np.random.randint(1, 100, size=1000000)
%timeit compute_reciprocals(big_array)
```
You certainly noticed that delay. The slowness of this looping becomes noticeable when we repeat many small operations many times.
The performance bottleneck is not the operations themselves, but the type-checking and function dispatches that Python performs on each cycle of the loop. In the case of the `compute_reciprocals` function above, each time Python computes the reciprocal, it first examines the object's type and does a dynamic lookup of the correct function to use for that type. Such is life with interpreted code. However, were we working with compiled code instead (such as in C), the object-type specification would be known before the code executes, and the result could be computed much more efficiently. This is where NumPy universal functions come into play.
### Ufuncs
Universal functions in NumPy (often shortened to *ufuncs*) provide a statically-typed, compiled function for many operations that we will need to run when manipulating and analyzing data.
Let's examine what this means in practice. Let's find the reciprocals of `big_array` again, this time using a built-in NumPy division ufunc on the array:
```
%timeit (1.0 / big_array)
```
That’s orders of magnitude better.
Ufuncs can be used between a scalar and an array and between arrays of arbitrary dimensions.
Computations vectorized by ufuncs are almost always more efficient than doing the same computation using Python loops. This is especially true on large arrays. When possible, try to use ufuncs when operating on NumPy arrays, rather than using ordinary Python loops.
Ufuncs come in two flavors: *unary ufuncs*, which use a single input, and *binary ufuncs*, which operate on two inputs. The common ufuncs we'll look at here encompass both kinds.
#### Array arithmetic
Many NumPy ufuncs use Python's native arithmetic operators, so you can use the standard addition, subtraction, multiplication, and division operators that we covered in Section 1:
```
a = np.arange(4)
print("a =", a)
print("a + 5 =", a + 5)
print("a - 5 =", a - 5)
print("a * 2 =", a * 2)
print("a / 2 =", a / 2)
print("a // 2 =", a // 2) # floor division
```
There are also ufuncs for negation, exponentiation, and the modulo operation:
```
print("-a = ", -a)
print("a ** 2 = ", a ** 2)
print("a % 2 = ", a % 2)
```
You can also combine these ufuncs using the standard order of operations:
```
-(0.5*a + 1) ** 2
```
The Python operators are not actually the ufuncs, but are rather wrappers around functions built into NumPy. So the `+` operator is actually a wrapper for the `add` function:
```
np.add(a, 2)
```
Here is a cheat sheet for the equivalencies between Python operators and NumPy ufuncs:
| Operator | Equivalent ufunc | Description |
|:--------------|:--------------------|:--------------------------------------|
|``+`` |``np.add`` |Addition (e.g., ``1 + 1 = 2``) |
|``-`` |``np.subtract`` |Subtraction (e.g., ``3 - 2 = 1``) |
|``-`` |``np.negative`` |Unary negation (e.g., ``-2``) |
|``*`` |``np.multiply`` |Multiplication (e.g., ``2 * 3 = 6``) |
|``/`` |``np.divide`` |Division (e.g., ``3 / 2 = 1.5``) |
|``//`` |``np.floor_divide`` |Floor division (e.g., ``3 // 2 = 1``) |
|``**`` |``np.power`` |Exponentiation (e.g., ``2 ** 3 = 8``) |
|``%`` |``np.mod`` |Modulus/remainder (e.g., ``9 % 4 = 1``)|
Python Boolean operators also work; we will explore those later in this section.
#### Absolute value
NumPy also understands Python's built-in absolute value function:
```
a = np.array([-2, -1, 0, 1, 2])
abs(a)
```
This corresponds to the NumPy ufunc `np.absolute` (which is also available under the alias `np.abs`):
```
np.absolute(a)
np.abs(a)
```
#### Exponents and logarithms
You will need to use exponents and logarithms a lot in data science; these are some of the most common data transformations for machine learning and statistical work.
```
a = [1, 2, 3]
print("a =", a)
print("e^a =", np.exp(a))
print("2^a =", np.exp2(a))
print("3^a =", np.power(3, a))
```
The basic `np.log` gives the natural logarithm. If you need to compute base-2 or base-10 logarithms, NumPy also provides those:
```
a = [1, 2, 4, 10]
print("a =", a)
print("ln(a) =", np.log(a))
print("log2(a) =", np.log2(a))
print("log10(a) =", np.log10(a))
```
There are also some specialized versions of these ufuncs to help maintain precision when dealing with very small inputs:
```
a = [0, 0.001, 0.01, 0.1]
print("exp(a) - 1 =", np.expm1(a))
print("log(1 + a) =", np.log1p(a))
```
These functions give more precise values than if you were to use the raw `np.log` or `np.exp` on very small values of `a`.
#### Specialized ufuncs
NumPy has many other ufuncs. Another source for specialized and obscure ufuncs is the submodule `scipy.special`. If you need to compute some specialized mathematical or statistical function on your data, chances are it is implemented in `scipy.special`.
```
from scipy import special
# Gamma functions (generalized factorials) and related functions
a = [1, 5, 10]
print("gamma(a) =", special.gamma(a))
print("ln|gamma(a)| =", special.gammaln(a))
print("beta(a, 2) =", special.beta(a, 2))
```
> **Takeaway:** Universal functions in NumPy provide you with computational functions that are faster than regular Python functions, particularly when working on large datasets that are common in data science. This speed is important because it can make you more efficient as a data scientist and it makes a broader range of inquiries into your data tractable in terms of time and computational resources.
## Aggregations
> **Learning goal:** By the end of this subsection, you should be comfortable aggregating data in NumPy.
One of the first things you will find yourself doing with most datasets is computing the summary statistics for the data to get a general overview of your data before exploring it further. These summary statistics include the mean and standard deviation, in addition to other aggregates, such as the sum, product, median, minimum and maximum, or quantiles of the data.
NumPy has fast built-in aggregation functions for working on arrays that are the subject of this subsection.
### Summing the values of an array
You can use the built-in Python `sum` function to sum up the values in an array.
```
import numpy as np
myList = np.random.random(100)
sum(myList)
```
If you guessed that there is also a built-in NumPy function for this, you guessed correctly:
```
np.sum(myList)
```
And if you guessed that the NumPy version is faster, you are doubly correct:
```
large_array = np.random.rand(1000000)
%timeit sum(large_array)
%timeit np.sum(large_array)
```
For all their similarity, bear in mind that `sum` and `np.sum` are not identical. Their optional arguments have different meanings and `np.sum` is aware of multiple array dimensions.
### Minimum and maximum
Just as Python has built-in `min` and `max` functions, NumPy has similar, vectorized versions:
```
np.min(large_array), np.max(large_array)
```
You can also use `min`, `max`, and `sum` (and several other NumPy aggregates) as methods of the array object itself:
```
print(large_array.min(), large_array.max(), large_array.sum())
```
### Multidimensional aggregates
Because you will often treat the rows and columns of two-dimensional arrays differently (treating columns as variables and rows as observations of those variables, for example), it can often be desirable to aggregate array data along a row or column. Let's consider a two-dimensional array:
```
md = np.random.random((3, 4))
print(md)
```
Unless you specify otherwise, each NumPy aggregation function will compute the aggregate for the entire array. Hence:
```
md.sum()
```
Aggregation functions take an additional argument specifying the *axis* along which to compute the aggregation. For example, we can find the minimum value within each column by specifying `axis=0`:
```
md.min(axis=0)
```
### Exercise:
```
# What do you get when you try md.max(axis=1)?
```
Remember that the `axis` keyword specifies the *dimension of the array that is to be collapsed*, not the dimension that will be returned. Thus specifying `axis=0` means that the first axis will be the one collapsed. For two-dimensional arrays, this means that values within each column will be aggregated.
### Other aggregation functions
The table below lists other aggregation functions in NumPy. Most NumPy aggregates have a '`NaN`-safe' version, which computes the result while ignoring missing values marked by the `NaN` value.
|Function Name | NaN-safe Version | Description |
|:------------------|:--------------------|:----------------------------------------------|
| ``np.sum`` | ``np.nansum`` | Compute sum of elements |
| ``np.prod`` | ``np.nanprod`` | Compute product of elements |
| ``np.mean`` | ``np.nanmean`` | Compute mean of elements |
| ``np.std`` | ``np.nanstd`` | Compute standard deviation |
| ``np.var`` | ``np.nanvar`` | Compute variance |
| ``np.min`` | ``np.nanmin`` | Find minimum value |
| ``np.max`` | ``np.nanmax`` | Find maximum value |
| ``np.argmin`` | ``np.nanargmin`` | Find index of minimum value |
| ``np.argmax`` | ``np.nanargmax`` | Find index of maximum value |
| ``np.median`` | ``np.nanmedian`` | Compute median of elements |
| ``np.percentile`` | ``np.nanpercentile``| Compute rank-based statistics of elements |
| ``np.any`` | N/A | Evaluate whether any elements are true |
| ``np.all`` | N/A | Evaluate whether all elements are true |
We will see these aggregates often throughout the rest of the course.
> **Takeaway:** Aggregation is the primary means you will use to explore data--not just when using NumPy, but particularly in conjunction with pandas, the Python library you will learn about in the next section, which builds off of NumPy and thus off of everything you have learned thus far.
## Computation on arrays with broadcasting
> **Learning goal:** By the end of this subsection, you should have a basic understanding of how broadcasting works in NumPy (and why NumPy uses it).
Another means of vectorizing operations is to use NumPy's *broadcasting* functionality: creating rules for applying binary ufuncs like addition, subtraction, or multiplication on arrays of different sizes.
Before, when we performed binary operations on arrays of the same size, those operations were performed on an element-by-element basis.
```
first_array = np.array([3, 6, 8, 1])
second_array = np.array([4, 5, 7, 2])
first_array + second_array
```
Broadcasting enables you to perform these types of binary operations on arrays of different sizes. Thus, you could just as easily add a scalar (which is really just a zero-dimensional array) to an array:
```
first_array + 5
```
Similarly, you can add a one-dimensional array to a two-dimensional array:
```
one_dim_array = np.ones((1))
one_dim_array
two_dim_array = np.ones((2, 2))
two_dim_array
one_dim_array + two_dim_array
```
So far, so easy. But you can use broadcasting on arrays in more complicated ways. Consider this example:
```
horizontal_array = np.arange(3)
vertical_array = np.arange(3)[:, np.newaxis]
print(horizontal_array)
print(vertical_array)
horizontal_array + vertical_array
```
### Rules of broadcasting
Broadcasting follows a set of rules to determine the interaction between the two arrays:
- **Rule 1**: If the two arrays differ in their number of dimensions, the shape of the one with fewer dimensions is *padded* with ones on its leading (left) side.
- **Rule 2**: If the shape of the two arrays does not match in any dimension, the array with shape equal to 1 in that dimension is stretched to match the other shape.
- **Rule 3**: If, in any dimension, the sizes disagree and neither is equal to 1, NumPy raises an error.
Let's see these rules in action to better understand them.
#### Broadcasting example 1
Let's look at adding a two-dimensional array to a one-dimensional array:
```
two_dim_array = np.ones((2, 3))
one_dim_array = np.arange(3)
```
Let's consider an operation on these two arrays. The shape of the arrays are:
- `two_dim_array.shape = (2, 3)`
- `one_dim_array.shape = (3,)`
We see by rule 1 that the array `one_dim_array` has fewer dimensions, so we pad it on the left with ones:
- `two_dim_array.shape -> (2, 3)`
- `one_dim_array.shape -> (1, 3)`
By rule 2, we now see that the first dimension disagrees, so we stretch this dimension to match:
- `two_dim_array.shape -> (2, 3)`
- `one_dim_array.shape -> (2, 3)`
The shapes match, and we see that the final shape will be `(2, 3)`:
```
two_dim_array + one_dim_array
```
### Exercise:
```
# Flip this around. Try adding these with two_dim_array = np.ones((3, 2))
# and one_dim_array = np.arange(3)[:, np.newaxis].
# What do you get?
```
#### Broadcasting example 2
Let's examine what happens when both arrays need to be broadcast:
```
vertical_array = np.arange(3).reshape((3, 1))
horizontal_array = np.arange(3)
```
Again, we'll start by writing out the shape of the arrays:
- `vertical_array.shape = (3, 1)`
- `horizontal_array.shape = (3,)`
Rule 1 says we must pad the shape of `horizontal_array ` with ones:
- `vertical_array.shape -> (3, 1)`
- `horizontal_array.shape -> (1, 3)`
And rule 2 tells us that we upgrade each of these to match the corresponding size of the other array:
- `vertical_array.shape -> (3, 3)`
- `horizontal_array.shape -> (3, 3)`
Because the result matches, these shapes are compatible. We can see this here:
```
vertical_array + horizontal_array
```
#### Broadcasting example 3
Here's what happens with incompatible arrays:
```
M = np.ones((3, 2))
i = np.arange(3)
```
This is a slightly different situation than in the first example: the matrix ``M`` is transposed.
How does this affect the calculation? The shape of the arrays are:
- ``M.shape = (3, 2)``
- ``i.shape = (3,)``
Again, rule 1 tells us that we must pad the shape of ``i`` with ones:
- ``M.shape -> (3, 2)``
- ``i.shape -> (1, 3)``
By rule 2, the first dimension of ``i`` is stretched to match that of ``M``:
- ``M.shape -> (3, 2)``
- ``i.shape -> (3, 3)``
Now we hit Rule 3: the final shapes do not match and the two arrays are incompatible:
```
M + i
```
### Broadcasting in practice
Ufuncs enable you to avoid using slow Python loops; broadcasting builds on that.
A common data practice is to *center* an array of data. For example, if we have an array of 10 observations, each of which consists of three values (called features in this context), we might want to center that data so that we have the differences from the mean rather than the raw data itself. Doing this can help us better compare the different values.
We'll store this in a $10 \times 3$ array:
```
T = np.random.random((10, 3))
T
```
Now compute the mean of each feature using the ``mean`` aggregate across the first dimension:
```
Tmean = T.mean(0)
Tmean
```
Finally, center ``T`` by subtracting the mean. (This is a broadcasting operation).
```
T_centered = T - Tmean
T_centered
```
This is not just faster, but easier than writing a loop to do this.
> **Takeaway:** The data you will work with in data science invariably comes in different shapes and sizes (at least in terms of the arrays in which you work with that data). The broadcasting functionality in NumPy enables you to use binary functions on irregularly fitting data in a predictable way.
## Comparisons, masks, and Boolean logic in NumPy
> **Learning goal:** By the end of this subsection, you should be comfortable with and understand how to use Boolean masking in NumPy to answer basic questions about your data.
*Masking* is when you want to manipulate, count, or extract values in an array based on a criterion. For example, counting the values in an array greater than a certain value is an example of masking. Boolean masking is often the most efficient way to accomplish these tasks in NumPy and it plays a large part in cleaning and otherwise preparing data for analysis (see Section 5).
### Example: Counting Rainy Days
Let's see masking in practice by examining the monthly rainfall statistics for Seattle. The data is in a CSV file from data.gov. To load the data, we will use pandas, which we will formally introduce in Section 4.
```
import numpy as np
import pandas as pd
# Use pandas to extract rainfall as a NumPy array
rainfall_2003 = pd.read_csv('Data/Observed_Monthly_Rain_Gauge_Accumulations_-_Oct_2002_to_May_2017.csv')['RG01'][ 2:14].values
rainfall_2003
```
Let’s break down what we just did in the code cell above. The rainfall data contains monthly rainfall totals from several rain gauges around the city of Seattle; we selected the first one. From that gauge, we then selected the relevant months for the first full calendar year in the dataset: 2003. That range of months started at the third row of the CSV file (remember, Python zero-indexes!) and ran through the thirteenth row, hence `2:14]`.
You now have an array containing 12 values, each of which records the monthly rainfall in inches from January to December 2003.
Commonly in data science, you will want to take a quick first exploratory look at the data. In this case, a bar chart is a good way to do this. To generate this bar chart, we will use Matplotlib, another important data science tool that we will introduce formally later in the course. (This also brings up another widely used Python convention you should adopt: `import matplotlib.pyplot as plt`.)
```
%matplotlib inline
import matplotlib.pyplot as plt
plt.bar(np.arange(1, len(rainfall_2003) + 1), rainfall_2003)
```
To briefly interpret the code snippet above, we passed two parameters to the bar function in pyplot: the first defining the index for the x-axis and the second defining the data to use for the bars (the y-axis). To create the index, we use the NumPy function `arange` to create a sequence of numbers (this is the same `arange` we encountered earlier in this section). We know that the length of our array is 12, but it can be a good habit to programmatically pass the length of an array in case it changes or you don’t know it with specificity. We also added 1 to both the start and the end of the `arange` to accommodate for Python zero-indexing (because there is no “month-zero” in the calendar).
Looking at the chart above (and as residents can attest), Seattle can have lovely, sunny summers. However, this is only a first glimpse of the data. There are still several questions we would like to answer: how many months did it rain or what was the average precipitation in those months? We would use masking to answer those questions. (We will also return to this example dataset to demonstrate concepts throughout the rest of this section). Before we dive deeper into explaining what masking is, we should briefly touch on comparison operators in NumPy.
### Comparison operators as ufuncs
In addition to the computational operators such as ufuncs that we have already encountered, NumPy also implements comparison operators such as `<` (less than) and `>` (greater than) as element-wise ufuncs. The standard Python comparison operations are available:
```
simple_array = np.array([1, 2, 3, 4, 5])
simple_array < 2 # less than
simple_array >= 4 # greater than or equal
simple_array == 2 # equal
```
It is also possible to do an element-wise comparison of two arrays, and to include compound expressions:
```
(2 * simple_array) == (simple_array ** 2)
```
As with the arithmetic operators, these comparison operators are wrappers for the NumPy ufuncs: when you write ``x < 3``, NumPy actually uses ``np.less(x, 3)``. Here is a summary of the comparison operators and their equivalent ufuncs:
| Operator | Equivalent ufunc || Operator | Equivalent ufunc |
|:--------------|:--------------------||:--------------|:--------------------|
|``==`` |``np.equal`` ||``!=`` |``np.not_equal`` |
|``<`` |``np.less`` ||``<=`` |``np.less_equal`` |
|``>`` |``np.greater`` ||``>=`` |``np.greater_equal`` |
Just like the arithmetic ufuncs, the comparison ufuncs work on arrays of any size and shape.
```
rand = np.random.RandomState(0)
two_dim_array = rand.randint(10, size=(3, 4))
two_dim_array
two_dim_array < 6
```
The result is a Boolean array, and NumPy provides a number of straightforward patterns for working with these Boolean results.
## Working with Boolean arrays
Given a Boolean array, there are a host of useful operations you can do.
We'll work with `two_dim_array`, the two-dimensional array we created earlier.
```
print(two_dim_array)
```
### Counting entries
To count the number of ``True`` entries in a Boolean array, ``np.count_nonzero`` is useful:
```
# how many values less than 6?
np.count_nonzero(two_dim_array < 6)
```
We see that there are eight array entries that are less than 6.
Another way to get this information is to use ``np.sum``; in this case, ``False`` is interpreted as ``0``, and ``True`` is interpreted as ``1``:
```
np.sum(two_dim_array < 5)
```
The benefit of `sum()` is that, like other NumPy aggregation functions, this summation can be done along rows or columns:
```
# how many values less than 5 in each row?
np.sum(two_dim_array < 5, axis=1)
```
This counts the number of values less than 5 in each row of the matrix.
If we're interested in quickly checking whether any or all the values are true, we can use (you guessed it) ``np.any`` or ``np.all``:
```
# Are there any values less than zero?
np.any(two_dim_array < 0)
```
### Exercise:
```
# Now check to see if all values less than 10?
# Hint: use np.all()
```
``np.all`` and ``np.any`` can be used along particular axis as well. For example:
```
# are all values in each row less than 7?
np.all(two_dim_array < 7, axis=1)
```
Here, all the elements in the first and third rows are less than 7, while this is not the case for the second row.
**A reminder:** Python has built-in `sum()`, `any()`, and `all()` functions. These have a different syntax than the NumPy versions, and, in particular, will fail or produce unintended results when used on multidimensional arrays. Be sure that you are using `np.sum()`, `np.any()`, and `np.all()` for these examples.
### Boolean operators
We've already seen how we might count, say, all months with rain less than four inches, or all months with more than two inches of rain. But what if we want to know all months with rain less than four inches and greater than one inch? This is accomplished through Python's *bitwise logic operators*, `&`, `|`, `^`, and `~`. Like with the standard arithmetic operators, NumPy overloads these as ufuncs which work element-wise on (usually Boolean) arrays.
For example, we can address this compound question as follows:
```
np.sum((rainfall_2003 > 0.5) & (rainfall_2003 < 1))
```
So we see that there are two months with rainfall between 0.5 and 1.0 inches.
Note that the parentheses here are important because of operator-precedence rules. With parentheses removed, this expression would be evaluated as follows, which results in an error:
```
rainfall_2003 > (0.5 & rainfall_2003) < 1
```
Using the equivalence of *A AND B and NOT (NOT A OR NOT B)* (which you might remember if you've taken an introductory logic course), we can compute the same result in a different manner:
```
np.sum(~((rainfall_2003 <= 0.5) | (rainfall_2003 >= 1)))
```
Combining comparison operators and Boolean operators on arrays can lead to a wide range of efficient logical operations.
The following table summarizes the bitwise Boolean operators and their equivalent ufuncs:
| Operator | Equivalent ufunc || Operator | Equivalent ufunc |
|:--------------|:--------------------||:--------------|:--------------------|
|``&`` |``np.bitwise_and`` ||| |``np.bitwise_or`` |
|``^`` |``np.bitwise_xor`` ||``~`` |``np.bitwise_not`` |
Using these tools, you can start to answer the questions we listed above about the Seattle rainfall data. Here are some examples of results we can compute when combining masking with aggregations:
```
print("Number of months without rain:", np.sum(rainfall_2003 == 0))
print("Number of months with rain: ", np.sum(rainfall_2003 != 0))
print("Months with more than 1 inch: ", np.sum(rainfall_2003 > 1))
print("Rainy months with < 1 inch: ", np.sum((rainfall_2003 > 0) &
(rainfall_2003 < 1)))
```
## Boolean arrays as masks
In the prior section, we looked at aggregates computed directly on Boolean arrays.
A more powerful pattern is to use Boolean arrays as masks, to select particular subsets of the data themselves.
Returning to our `two_dim_array` array from before, suppose we want an array of all values in the array that are less than 5:
```
two_dim_array
```
You can obtain a Boolean array for this condition easily:
```
two_dim_array < 5
```
Now, to *select* these values from the array, you can simply index on this Boolean array. This is the *masking* operation:
```
two_dim_array[two_dim_array < 5]
```
What is returned is a one-dimensional array filled with the values that meet your condition. Put another way, these are the values in positions at which the mask array is ``True``.
You can use masking as a way to compute some relevant statistics on the Seattle rain data:
```
# Construct a mask of all rainy months
rainy = (rainfall_2003 > 0)
# Construct a mask of all summer months (June through September)
months = np.arange(1, 13)
summer = (months > 5) & (months < 10)
print("Median precip in rainy months in 2003 (inches): ",
np.median(rainfall_2003[rainy]))
print("Median precip in summer months in 2003 (inches): ",
np.median(rainfall_2003[summer]))
print("Maximum precip in summer months in 2003 (inches): ",
np.max(rainfall_2003[summer]))
print("Median precip in non-summer rainy months (inches):",
np.median(rainfall_2003[rainy & ~summer]))
```
> **Takeaway:** By combining Boolean operations, masking operations, and aggregates, you can quickly answer questions similar to those we posed about the Seattle rainfall data about any dataset. Operations like these will form the basis for the data exploration and preparation for analysis that will be our primary concerns in Sections 4 and 5.
| github_jupyter |
<a href="https://colab.research.google.com/github/nnuncert/nnuncert/blob/master/notebooks/DNNC_uci.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Git + Repo Installs
```
!git clone https://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@github.com/nnuncert/nnuncert
%cd nnuncert
!pip install -r requirements.txt
```
# Imports
```
# general imports
import numpy as np
import numexpr as ne
import tensorflow as tf
import matplotlib.pyplot as plt
# thesis code
import nnuncert
from nnuncert.models import make_model, type2name
from nnuncert.utils.traintest import TrainTestSplit
from nnuncert.app.uci import UCI_DATASETS, load_uci
from nnuncert.utils.dist import Dist
```
# UCI
## Load a dataset
```
# show all available datasets
UCI_DATASETS
# load boston and look at ddata
uci = load_uci("boston")
# must be given proper directory where data .csv files are stored
uci.get_data("data/uci")
# prepare data by hot encoding categoricals
uci.prepare_run(drop_first=True)
# look at boston data
uci.data.data.head()
# create train / test split (10 % test ratio)
# we standardize the categroical features to be zero mean and unit variance
# split has attributes such as 'x_train', 'x_test', 'y_train', 'y_test'
split = uci.make_train_test_split(ratio=0.1)
# you can get id for train and test sets with
# split.train_id or split.train_id
# i.e., see training data as DataFrame
split.data_train
```
## Fit model
```
# estimate KDE for response
dist = Dist._from_values(uci.data.y, method=uci.dist_method, **uci.dist_kwargs)
y0 = np.linspace(uci.data.y.min(), uci.data.y.max(), 100)
_ = plt.hist(uci.data.y, bins=50, density=True, color="lightgrey")
plt.plot(y0, dist.pdf(y0))
# handle general settings
arch = [[50, "relu", 0]] # list of hidden layer description (size, act. func, dropout rate)
epochs = 40
verbose = 0
learning_rate = 0.01
# get input shape from x_train
input_shape = split.x_train.shape[1]
# make model and compile
model = make_model("DNNC-R", input_shape, arch)
model.compile(optimizer=tf.keras.optimizers.Adam(learning_rate=learning_rate),
metrics=["mae", "mse"])
# fit to x_train, y_train
# specify kwargs for DNNC by passing arguments into dnnc_kwargs
# example parameters: J = [burn-in, samples], theta, tau2start
# see nnuncert/models/dnnc/_dnnc.py for details
model.fit(split.x_train, split.y_train, epochs=epochs, verbose=verbose, dist=dist, dnnc_kwargs={})
# get predictions for training and test features
pred_train = model.make_prediction(split.x_train)
pred_test = model.make_prediction(split.x_test)
```
## Evaluate performance for run
```
from nnuncert.app.uci import UCIRun
# get scores
scores = UCIRun(pred_train, split.y_train, pred_test, split.y_test,
model="PNN-E", dataset="boston")
# scores has attributes:
# 'rmse_train', 'rmse_test', 'log_score_train', 'log_score_test', 'crps_train',
# 'crps_test', 'picp_train', 'picp_test', 'mpiw_train', 'mpiw_test'
print("RMSE: \t\t", scores.rmse_test, "\nLog Score: \t", scores.log_score_test)
```
## Plot
```
from nnuncert.utils.indexing import index_to_rowcol
pred = pred_test
fig, ax = plt.subplots(2, 4, figsize=(14, 6))
# where to evaluate density
y0 = np.linspace(5, 50, 100)
# plot predictive densities (choose 8 randomly)
# randomly shuffled in train/test anyway
for i in range(8):
r, c = index_to_rowcol(i, 4)
ax_ = ax[r, c]
ax_.plot(y0, pred.pdfi(i, y0))
```
# DNNC attributes
```
# access DNNC sampling details with the object model.dnnc
# e.g., samples betas:
model.dnnc.betahat
# or acceptance rates for tau2:
model.dnnc.tau2accs
# Expected values, variance, etc
# pred_test.dens.Ey, pred_test.dens.Vary, pred_test.dens.lpy
pred_test.dens.Vary
```
| github_jupyter |
# Perceptron with Quantile Transformer
This Code template is for the Classification task using simple Perceptron. Which is a simple classification algorithm suitable for large scale learning and feature transformation technique Quantile Transformer in a pipeline.
### Required Packages
```
!pip install imblearn
import warnings
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as se
from imblearn.over_sampling import RandomOverSampler
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import train_test_split
from sklearn.linear_model import Perceptron
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import QuantileTransformer
from sklearn.metrics import classification_report,plot_confusion_matrix
warnings.filterwarnings('ignore')
```
### Initialization
Filepath of CSV file
```
#filepath
file_path= ""
```
List of features which are required for model training .
```
#x_values
features=[]
```
Target feature for prediction.
```
#y_value
target=''
```
### Data Fetching
Pandas is an open-source, BSD-licensed library providing high-performance, easy-to-use data manipulation and data analysis tools.
We will use panda's library to read the CSV file using its storage path.And we use the head function to display the initial row or entry.
```
df=pd.read_csv(file_path)
df.head()
```
### Feature Selections
It is the process of reducing the number of input variables when developing a predictive model. Used to reduce the number of input variables to both reduce the computational cost of modelling and, in some cases, to improve the performance of the model.
We will assign all the required input features to X and target/outcome to Y.
```
X = df[features]
Y = df[target]
```
### Data Preprocessing
Since the majority of the machine learning models in the Sklearn library doesn't handle string category data and Null value, we have to explicitly remove or replace null values. The below snippet have functions, which removes the null value if any exists. And convert the string classes data in the datasets by encoding them to integer classes.
```
def NullClearner(df):
if(isinstance(df, pd.Series) and (df.dtype in ["float64","int64"])):
df.fillna(df.mean(),inplace=True)
return df
elif(isinstance(df, pd.Series)):
df.fillna(df.mode()[0],inplace=True)
return df
else:return df
def EncodeX(df):
return pd.get_dummies(df)
def EncodeY(df):
if len(df.unique())<=2:
return df
else:
un_EncodedT=np.sort(pd.unique(df), axis=-1, kind='mergesort')
df=LabelEncoder().fit_transform(df)
EncodedT=[xi for xi in range(len(un_EncodedT))]
print("Encoded Target: {} to {}".format(un_EncodedT,EncodedT))
return df
x=X.columns.to_list()
for i in x:
X[i]=NullClearner(X[i])
X=EncodeX(X)
Y=EncodeY(NullClearner(Y))
X.head()
```
#### Correlation Map
In order to check the correlation between the features, we will plot a correlation matrix. It is effective in summarizing a large amount of data where the goal is to see patterns.
```
f,ax = plt.subplots(figsize=(18, 18))
matrix = np.triu(X.corr())
se.heatmap(X.corr(), annot=True, linewidths=.5, fmt= '.1f',ax=ax, mask=matrix)
plt.show()
```
#### Distribution Of Target Variable
```
plt.figure(figsize = (10,6))
se.countplot(Y)
```
### Data Splitting
The train-test split is a procedure for evaluating the performance of an algorithm. The procedure involves taking a dataset and dividing it into two subsets. The first subset is utilized to fit/train the model. The second subset is used for prediction. The main motive is to estimate the performance of the model on new data.
```
x_train,x_test,y_train,y_test=train_test_split(X,Y,test_size=0.2,random_state=123)
```
#### Handling Target Imbalance
The challenge of working with imbalanced datasets is that most machine learning techniques will ignore, and in turn have poor performance on, the minority class, although typically it is performance on the minority class that is most important.
One approach to addressing imbalanced datasets is to oversample the minority class. The simplest approach involves duplicating examples in the minority class.We will perform overspampling using imblearn library.
```
x_train,y_train = RandomOverSampler(random_state=123).fit_resample(x_train, y_train)
```
### Model
the perceptron is an algorithm for supervised learning of binary classifiers.
The algorithm learns the weights for the input signals in order to draw a linear decision boundary.This enables you to distinguish between the two linearly separable classes +1 and -1.
#### Model Tuning Parameters
> **penalty** ->The penalty (aka regularization term) to be used. {‘l2’,’l1’,’elasticnet’}
> **alpha** -> Constant that multiplies the regularization term if regularization is used.
> **l1_ratio** -> The Elastic Net mixing parameter, with 0 <= l1_ratio <= 1. l1_ratio=0 corresponds to L2 penalty, l1_ratio=1 to L1. Only used if penalty='elasticnet'.
> **tol** -> The stopping criterion. If it is not None, the iterations will stop when (loss > previous_loss - tol).
> **early_stopping**-> Whether to use early stopping to terminate training when validation. score is not improving. If set to True, it will automatically set aside a stratified fraction of training data as validation and terminate training when validation score is not improving by at least tol for n_iter_no_change consecutive epochs.
> **validation_fraction** -> The proportion of training data to set aside as validation set for early stopping. Must be between 0 and 1. Only used if early_stopping is True.
> **n_iter_no_change** -> Number of iterations with no improvement to wait before early stopping.
Feature Transformation
This method transforms the features to follow a uniform or a normal distribution. Therefore, for a given feature, this transformation tends to spread out the most frequent values. It also reduces the impact of (marginal) outliers: this is therefore a robust preprocessing scheme.
The transformation is applied on each feature independently. First an estimate of the cumulative distribution function of a feature is used to map the original values to a uniform distribution. The obtained values are then mapped to the desired output distribution using the associated quantile function. Features values of new/unseen data that fall below or above the fitted range will be mapped to the bounds of the output distribution. Note that this transform is non-linear. It may distort linear correlations between variables measured at the same scale but renders variables measured at different scales more directly comparable.
Refer [API](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.QuantileTransformer.html) for the parameters
```
# Build Model here
model=make_pipeline(QuantileTransformer(),Perceptron(random_state=123))
model.fit(x_train, y_train)
```
#### Model Accuracy
score() method return the mean accuracy on the given test data and labels.
In multi-label classification, this is the subset accuracy which is a harsh metric since you require for each sample that each label set be correctly predicted.
```
print("Accuracy score {:.2f} %\n".format(model.score(x_test,y_test)*100))
```
#### Confusion Matrix
A confusion matrix is utilized to understand the performance of the classification model or algorithm in machine learning for a given test set where results are known.
```
plot_confusion_matrix(model,x_test,y_test,cmap=plt.cm.Blues)
```
#### Classification Report
A Classification report is used to measure the quality of predictions from a classification algorithm. How many predictions are True, how many are False.
* **where**:
- Precision:- Accuracy of positive predictions.
- Recall:- Fraction of positives that were correctly identified.
- f1-score:- percent of positive predictions were correct
- support:- Support is the number of actual occurrences of the class in the specified dataset.
```
print(classification_report(y_test,model.predict(x_test)))
```
#### Creator: Akshar Nerkar , Github: [Profile](https://github.com/Akshar777)
| github_jupyter |
Final Project-
AIM: To simulate safe navigation of the robot
Steps:
1. Make a custom grid
2. Give Start and Goal pose
3. Perform Astar algorithms to get the global path planning
4. Get the trajectory of poses(x,y,theta) using x,y values from astar path planning
5. Implement the DWA (local planner + tracker) to track the robot
6. Add collison avoidance by the help of Lidar sensor on the robot to avoid any dynamic obstacle
```
import numpy as np
import time
import toml
import networkx as nx
import matplotlib.pyplot as plt
from matplotlib.animation import FuncAnimation
from IPython.display import Image
import cv2
from PIL import Image
%matplotlib inline
```
Make a custom grid and add obstacle
```
#custom grid
#choose resolution and dimensions
grid_res = 0.05
grid_span = 50 # square circuit dimensions in m
#calculate grid_shape from grid
# dimensions have to be integers
grid_shape = (np.array([grid_span]*2)/grid_res).astype('int')
#Initialize
grid_data = np.zeros(grid_shape)
#Create rectangular obstacles in world co-ordinates
#xmin, xmax, ymin, ymax
obstacles = np.array([[25, 26, 10, 40],
[2, 8, 16, 20]])
for obs in obstacles:
# calculate obstacles extent in pixel coords
xmin, xmax, ymin, ymax = (obs/grid_res).astype('int')
# mark them as occupied
grid_data[xmin:xmax ,ymin:ymax ] = 1.0
#calculate the extents
x1, y1 = 0, 0
x2, y2 = grid_span, grid_span
#imshow() from prior cell
plt.figure()
plt.imshow(1-grid_data.T, origin='lower', cmap=plt.cm.gray, extent=[x1,x2,y1,y2] )
```
Eulcidean function for calculating distance between two points
```
# write the Euclidean function that takes in the
# node x, y and compute the distance
def euclidean(node1, node2):
x1, y1 = node1
x2, y2 = node2
return np.sqrt((x1-x2)**2+(y1-y2)**2)
```
NetwrokX initialization of Graph
```
#initialize graph
grid_size=grid_shape
G=nx.grid_2d_graph(*grid_size)
deleted_nodes = 0 # counter to keep track of deleted nodes
#loop to remove nodes
for i in range(grid_size[0]):
for j in range(grid_size[1]):
if grid_data[i,j]==1:
G.remove_node((i,j))
deleted_nodes+=1
print(f"removed {deleted_nodes} nodes")
print(f"number of occupied cells in grid {np.sum(grid_data)}")
```
Give Start and Goal pose as needed[User should input this]
```
start_pose=(5/grid_res,10/grid_res)
goal_pose=(5/grid_res,25/grid_res)
```
Astar implementation in NetworkX
```
nx.set_edge_attributes(G, {e: 1 for e in G.edges()}, "cost")
astar_path = nx.astar_path(G, start_pose, goal_pose, heuristic=euclidean, weight="cost")
astar_path
```
Plotting the astar path on the grid
```
fig, ax = plt.subplots(figsize=(12,12))
ax.imshow(grid_data, cmap=plt.cm.Dark2)
ax.scatter(start_pose[1],start_pose[0], marker = "+", color = "yellow", s = 50)
ax.scatter(goal_pose[1],goal_pose[0], marker = "+", color = "red", s = 50)
for s in astar_path[1:]:
ax.plot(s[1], s[0],'y+')
```
Converting the (x,y) from astar to route commands
```
commands=[]
cor=1
for i in range (len(astar_path)-2):
x1,y1=astar_path[i]
x2,y2=astar_path[i+1]
x3,y3=astar_path[i+2]
eval = cor*((x1-x2)*(y3-y2)-(x3-x2)*(y2-y1))
if (eval != 0):
cor*=-1
if eval >0:
## left turn +90
commands.append(("turn",90))
else:
commands.append(("turn",-90))
else:
commands.append(("straight",euclidean( (x2,y2), (x1,y1)) ))
print(commands)
```
Converting the route command into trajectory of poses(x,y,theta) and smooth planning with the help of cubic spiral fucntion varitaion of theta
```
v = 1
dt = 0.1
num_st_pts = int(v/dt)
num_pts = 8
def cubic_spiral(theta_i, theta_f, n=10):
x = np.linspace(0, 1, num=n)
#-2*x**3 + 3*x**2
return (theta_f-theta_i)*(-2*x**3 + 3*x**2) + theta_i
def straight(dist, curr_pose, n=num_st_pts):
# the straight-line may be along x or y axis
#curr_theta will determine the orientation
x0, y0, t0 = curr_pose
xf, yf = x0 + dist*np.cos(t0), y0 + dist*np.sin(t0)
x = (xf - x0) * np.linspace(0, 1, n) + x0
y = (yf - y0) * np.linspace(0, 1, n) + y0
return x, y, t0*np.ones_like(x)
def turn(change, curr_pose, n=num_pts):
# adjust scaling constant for desired turn radius
x0, y0, t0 = curr_pose
theta = cubic_spiral(t0, t0 + np.deg2rad(change), n)
x= x0 + np.cumsum(v*np.cos(theta)*dt)
y= y0 + np.cumsum(v*np.sin(theta)*dt)
return x, y, theta
def generate_trajectory(route, init_pose = (start_pose[0], start_pose[1], np.pi/2)):
curr_pose = init_pose
func = {'straight': straight, 'turn': turn}
x, y, t = np.array([]), np.array([]),np.array([])
for manoeuvre, command in route:
px, py, pt = func[manoeuvre](command, curr_pose)
curr_pose = px[-1],py[-1],pt[-1]
x = np.concatenate([x, px])
y = np.concatenate([y, py])
t = np.concatenate([t, pt])
return np.vstack([x, y, t])
```
Plotting the final trajectory poses
```
x, y, t = generate_trajectory(commands)
plt.figure()
plt.plot(x, y)
plt.grid()
print(t)
ref_path=generate_trajectory(commands)
print(ref_path)
```
Configuring the parameters for DWA implementation
```
config_params = toml.load("config.toml")['params']
print(config_params)
locals().update(config_params)
print(dt, V_MAX)
v_min, v_max = 0.0, 0.2
w_min, w_max = -0.1, 0.1
vs = np.linspace(v_min, v_max, num=11)
ws = np.linspace(w_min, w_max, num=11)
cmd = np.transpose([np.tile(vs, len(ws)), np.repeat(ws, len(vs))])
print(vs)
```
Making circles to resemble the robot footprint
```
w=0.8
l=1.2
mule_extents = np.array([[-w/2,-l/2],
[w/2, -l/2],
[w/2, l/2],
[-w/2, l/2],
[-w/2,-l/2]])
r = 0.5
l = 0.4
circles = [(0, 0, r), (0, l, r), (0, -l, r)]
plt.figure()
#plot rectangle or just the 4 vertices
for vertex in mule_extents:
p, q = vertex
plt.plot(p, q, 'b.')
for v1, v2 in zip(mule_extents[:-1],mule_extents[1:]):
p1, q1 = v1
p2, q2 = v2
plt.plot((p1, p2), (q1,q2), 'k-')
ax = plt.gca()
for x,y,rad in circles:
ax.add_patch(plt.Circle((x,y), rad, fill=False))
```
Circle collision check function
```
grid_res = 0.05
def circle_collision_check(grid_data, local_traj):
xmax, ymax = grid_shape
all_x = np.arange(xmax)
all_y = np.arange(ymax)
X, Y = np.meshgrid(all_x, all_y)
for xl, yl, tl in local_traj:
rot = np.array([[np.sin(tl), -np.cos(tl)],[np.cos(tl), np.sin(tl)]])
for xc, yc, rc in circles:
xc_rot, yc_rot = rot @ np.array([xc, yc]) + np.array([xl, yl])
xc_pix, yc_pix = int(xc_rot/grid_res), int(yc_rot/ grid_res)
rc_pix = (rc/ grid_res)
inside_circle = ((X-xc_pix)**2 +(Y-yc_pix)**2 - rc_pix**2 < 0)
occupied_pt = grid_data[X, Y] == 1
if np.sum(np.multiply( inside_circle, occupied_pt)):
return True
return False
```
Lidar sensors class definition
```
class Lidar(object):
def __init__(self, nbeams=7, fov=60, max_dist=5.0, sampling_pts=20):
#nbeams = number of sensing beams
#FOV = field-of-view of Lidar/ coverage in degrees
#max_dist = maximum distance Lidar can sense
#sampling_pts = num pts on a given beam for obstacle check
self.beam_angles = np.deg2rad(np.linspace(-fov/2+np.pi/2,fov/2+np.pi/2,num=nbeams))
self.line_sampler = max_dist * np.linspace(0, 1, num=sampling_pts)
def set_env(self, grid, grid_res=0.05):
#2-D occupancy grid and grid_resolution
self.grid = grid_data
self.grid_res = grid_res
def sense_obstacles(self, pose):
xc, yc, theta = pose[-1][0], pose[-1][1], pose[-1][2]
beam_data = []
for b in self.beam_angles:
direction = np.array([np.cos(theta+b), np.sin(theta+b)])
for d in self.line_sampler:
beam_x, beam_y = np.array([xc, yc]) + d * direction
i, j = int(beam_x), int(beam_y)
if self.grid[i][j] == 1:
break
beam_data.append(d)
return beam_data
```
DWA fucntions definiton
```
def simulate_unicycle(pose, v,w, N=1, dt=0.1):
x, y, t = pose[-1][0], pose[-1][1], pose[-1][2]
poses = []
for _ in range(N):
x += v*np.cos(t)*dt
y += v*np.sin(t)*dt
t += w*dt
# Keep theta bounded between [-pi, pi]
t = np.arctan2(np.sin(t), np.cos(t))
poses.append([x,y,t])
return np.array(poses)
def command_window(v, w, dt=0.1):
"""Returns acceptable v,w commands given current v,w"""
# velocity can be (0, V_MAX)
# ACC_MAX = max linear acceleration
v_max = min(V_MAX, v + ACC_MAX*dt)
v_min = max(0, v - ACC_MAX*dt)
# omega can be (-W_MAX, W_MAX)
#W_DOT_MAX = max angular acceleration
epsilon = 1e-6
w_max = min(W_MAX, w + W_DOT_MAX*dt)
w_min = max(-W_MAX, w - W_DOT_MAX*dt)
#generate quantized range for v and omega
vs = np.linspace(v_min, v_max, num=11)
ws = np.linspace(w_min, w_max, num=21)
#cartesian product of [vs] and [ws]
#remember there are 0 velocity entries which have to be discarded eventually
commands = np.transpose([np.tile(vs, len(ws)), np.repeat(ws, len(vs))])
#calculate kappa for the set of commands
kappa = commands[:,1]/(commands[:,0]+epsilon)
#returning only commands < max curvature
return commands[(kappa < K_MAX) & (commands[:, 0] != 0)]
def track(ref_path, pose, v, w, dt=0.1):
commands = command_window(v, w, dt)
#initialize path cost
best_cost, best_command = np.inf, None
for i, (v, w) in enumerate(commands):
local_path = simulate_unicycle(pose,v, w, pred_horizon) #Number of steps = prediction horizon
w_obs=0
#if circle_collision_check(grid_data, local_path): #ignore colliding paths
# print("local path has a collision")
# continue
#lidar = Lidar(max_dist=5)
#lidar.set_env(grid_data)
#dist = lidar.sense_obstacles(pose=pose)
#print(f"Beam distances are {dist}")
#for i in range(len(dist)):
# if dist[i]<5.0:
# w_obs=100
# else:
# w_obs=0
#calculate cross-track error
#can use a simplistic definition of
#how close is the last pose in local path from the ref path
cte = euclidean(local_path[-1][:2], ref_path[-1][:2])
#other cost functions are possible
#can modify collision checker to give distance to closest obstacle
cost = w_cte*cte + w_speed*(V_MAX - v)**2 + w_obs
#check if there is a better candidate
if cost < best_cost:
best_cost, best_command = cost, (v, w)
if best_command:
return best_command
else:
return [0, 0]
```
Function call for DWA
```
ref_path=generate_trajectory(commands)
print(ref_path)
pred_horizon=2
start_pose_ = np.array([start_pose[0],start_pose[1], np.pi/2])
ref_path = ref_path.T
pose = []
pose.append(start_pose_)
logs = []
path_index = 0
v, w = 0.0, 0.0
while path_index < len(ref_path)-1:
t0 = time.time()
local_ref_path = ref_path[path_index:path_index+pred_horizon]
# update path_index using current pose and local_ref_path
if np.sqrt((local_ref_path[-1][0]-pose[-1][0])**2 + (local_ref_path[-1][1]-pose[-1][1])**2) < goal_threshold*100:
path_index += pred_horizon
# get next command
v, w = track(local_ref_path, pose, v, w)
#simulate vehicle for 1 step
# remember the function now returns a trajectory, not a single pose
pose = simulate_unicycle(pose, v, w)
print(pose)
#update logs
logs.append([*pose, v, w])
t1 = time.time() #simplest way to time-profile your code
print(f"idx:{path_index}, v:{v:0.3f}, w:{w:0.3f}, time:{(t1-t0) * 1000:0.1f}ms")
poses
poses = np.array(logs)[:,:3]
plt.figure()
plt.axes().set_aspect('equal', 'datalim')
plt.plot(ref_path[:,0], ref_path[:,1], '.', c='y')
for s in range(len(poses[:,0])):
plt.plot(poses[:,0][s][0],poses[:,0][s][1],'r+')
plt.show()
```
Plotting the tracking of robot path
```
fig, ax = plt.subplots(figsize=(12,12))
ax.imshow(grid_data, cmap=plt.cm.Dark2)
ax.scatter(start_pose[1],start_pose[0], marker = "+", color = "yellow", s = 700)
ax.scatter(goal_pose[1],goal_pose[0], marker = "+", color = "red", s = 700)
for s in range(len(poses[:,0])):
ax.plot(poses[:,0][s][1],poses[:,0][s][0],'r+')
```
| github_jupyter |
# Classify Movie Reviews using Embeddings
```
!pip install tensorflow==2.0.0.beta0
#this version of numpy is required to avoid an error related to numpy defaulting to not allowing pickle files
!pip install numpy==1.16.2
import tensorflow as tf
from tensorflow.keras import datasets, preprocessing, models, layers
import numpy as np
```
## Import IMDB Reviews
```
imdb = datasets.imdb
#Let's work with a dictionary of 20,000 words
NUM_WORDS = 20000
#load IMDB dataset as lists of integers
(train_data, train_labels), (test_data, test_labels) = imdb.load_data(num_words=NUM_WORDS, )
```
## Get Word Index To See Words
```
word_index = imdb.get_word_index()
# The first indices are reserved
word_index = {k: (v+3) for k, v in word_index.items()}
word_index['<PAD>'] = 0
word_index['<START>'] = 1
word_index['<UNK>'] = 2
word_index['<UNUSED>'] = 3
reverse_word_index = dict([(value, key) for (key, value) in word_index.items()])
```
## Inspect First Review
```
print(' '.join([reverse_word_index[i] for i in train_data[0]]))
```
## Was this a positive review?
```
print(train_labels[0])
```
## Note: Reviews have variable length
```
print('Len 0: {}, Len 1: {}, Len 2: {}'.format(len(train_data[0]), len(train_data[1]), len(train_data[2])))
```
Variable lenght is fixed by truncating after a certain number of words. For reviews that are less than the number of words we are cutting off, we pad.
```
LEN_WORDS = 300
train_data = preprocessing.sequence.pad_sequences(train_data, maxlen=LEN_WORDS)
test_data = preprocessing.sequence.pad_sequences(test_data, maxlen=LEN_WORDS)
print('Len 0: {}, Len 1: {}, Len 2: {}'.format(len(train_data[0]), len(train_data[1]), len(train_data[2])))
print(train_data.shape)
```
## Sequential Model with Dense Layers
```
dense_model = models.Sequential([
layers.Dense(300, input_shape=(300,), activation='relu'),
layers.Dense(300, activation='relu'),
layers.Dense(300, activation='relu'),
layers.Dense(300, activation='relu'),
layers.Dense(1, activation='sigmoid')
])
print(dense_model.summary())
dense_model.compile(optimizer='adam', loss='binary_crossentropy', metrics=['accuracy'])
dense_model.fit(train_data, train_labels, epochs=10)
```
## Introduce Embeddings
```
DIMENSION = 16
e_model = models.Sequential([
layers.Embedding(NUM_WORDS, DIMENSION, input_length=LEN_WORDS),
layers.GlobalAveragePooling1D(),
layers.Dense(1, activation='sigmoid')
])
print(e_model.summary())
e_model.compile(optimizer='adam', loss='binary_crossentropy', metrics=['accuracy'])
e_model.fit(train_data, train_labels, epochs=10)
```
| github_jupyter |
```
import astropy.io.fits as pf
import os
import sys
import numpy as np
import datetime
import pyds9
ds = pyds9.DS9()
def check_header_refpoint(hdr):
"""
Check a FITS header to see if it does not have the expected data table
columns for telescope aspect.
Require the following keywords:
TELESCOP = 'NuSTAR'
INSTRUME = 'FPM?' # ? is 'A' or 'B'
EXTNAME = 'DET?_REFPOINT' # ? is a digit 1 to 4
HDUCLAS1 = 'TEMPORALDATA'
NAXIS = 2
Require the following fields:
TTYPE1 = 'TIME'
TUNIT1 = 's'
TTYPE2 = 'X_DET1'
TUNIT2 = 'pixel'
TTYPE3 = 'Y_DET1'
TUNIT3 = 'pixel'
"""
try:
if not (
hdr['TELESCOP'] == 'NuSTAR' and
(hdr['INSTRUME'] == 'FPMA' or hdr['INSTRUME'] == 'FPMB') and
(hdr['EXTNAME'] in
['DET%d_REFPOINT' % i for i in (1, 2, 3, 4)]) and
hdr['HDUCLAS1'] == 'TEMPORALDATA' and
hdr['NAXIS'] == 2 and
hdr['TTYPE1'] == 'TIME' and hdr['TUNIT1'] == 's' and
(hdr['TTYPE2'] in
['X_DET%d' % i for i in (1, 2, 3, 4)]) and
hdr['TUNIT2'] == 'pixel' and
(hdr['TTYPE3'] in
['Y_DET%d' % i for i in (1, 2, 3, 4)]) and
hdr['TUNIT3'] == 'pixel'
):
return False
except KeyError:
return False
return True
def check_header_gti(hdr):
"""
Check FITS header to see if it has the expected data table columns for GTI extension.
Require the following keywords:
TELESCOP = 'NuSTAR'
EXTNAME = 'STDGTI' # ? is a digit 1 to 4
NAXIS = 2
Require the following fields:
TTYPE1 = 'START'
TUNIT1 = 's' or 'sec'
TTYPE2 = 'STOP'
TUNIT2 = 's' or 'sec'
"""
try:
if not (
hdr['TELESCOP'] == 'NuSTAR' and
hdr['EXTNAME'] == 'STDGTI' and
hdr['NAXIS'] == 2 and
hdr['TTYPE1'] == 'START' and
hdr['TUNIT1'] in ('s', 'sec') and
hdr['TTYPE2'] == 'STOP' and
hdr['TUNIT2'] in ('s', 'sec')
):
return False
except KeyError:
return False
return True
os.chdir('/Users/qw/astro/nustar/IC342_X1/90201039002/event_cl')
os.getcwd()
check_header_refpoint(detB1fh[1].header)
refpointfile = 'nu90201039002B_det1.fits'
detB1fh = pf.open(refpointfile)
detB1fh.info()
refpointext = None
for ext in detB1fh:
if check_header_refpoint(ext.header):
refpointext = ext
break
if refpointext is None:
print('No aspect info in the specified file %s.' % refpointfile)
else:
print('Found extension %s.' % refpointext.header['EXTNAME'])
refpointext.data.columns
refpointext.data
gtifile = 'nu90201039002B01_gti.fits'
gtiBfh = pf.open(gtifile)
gtiBfh.info()
gtiext = None
for ext in gtiBfh:
if check_header_gti(ext.header):
gtiext = ext
break
if gtiext is None:
print('No GTI info in the specified file %s.' % gtifile)
else:
print('Found extension %s.' % gtiext.header['EXTNAME'])
gtiext.data[:10]
gtiarr = np.sort(gtiext.data, order='START', kind='mergesort')
gtiarr[:10]
refpointarr = np.sort(refpointext.data, order='TIME', kind='mergesort')
refpointarr[:10][0]
i_refpoint = 0
n_refpoint = len(refpointarr)
i_gti = 0
n_gti = len(gtiarr)
coords = []
dt = []
print('Processing %d aimpoints...' % n_refpoint)
while i_refpoint < (n_refpoint - 1) and i_gti < n_gti:
# Ref pointings have a single time while GTI intervals have two.
# Original logic in projobs.pro considers pointing time against [start, stop),
# i.e. >= for start time, < for stop.
if refpointarr[i_refpoint][0] < gtiarr[i_gti][0]:
# Pointing is before current interval, go to next pointing
i_refpoint += 1
continue
if refpointarr[i_refpoint][0] >= gtiarr[i_gti][1]:
# Pointing is after current interval, go to next interval
i_gti += 1
continue
# Otherwise there is some overlap. Add this pointing only if it is entirely within the GTI interval (original behavior).
coords.append([refpointarr[i_refpoint][1], refpointarr[i_refpoint][2]])
dt.append(refpointarr[i_refpoint + 1][0] - refpointarr[i_refpoint][0])
i_refpoint += 1
print('%d aimpoints (%f / %f s)' % (
len(coords), np.sum(dt),
np.sum(gtiarr.field('STOP') - gtiarr.field('START'))))
asphistimg = np.zeros((1000, 1000), dtype=np.float64)
x_min, y_min, x_max, y_max = 999, 999, 0, 0
for i in range(len(dt)):
x, y = int(np.floor(coords[i][0])), int(np.floor(coords[i][1]))
asphistimg[y, x] += dt[i]
if y < y_min:
y_min = y
elif y > y_max:
y_max = y
if x < x_min:
x_min = x
elif x > x_max:
x_max = x
ds.set_np2arr(asphistimg[y_min:y_max+1, x_min:x_max+1])
print('Image subspace: ', x_min, x_max, y_min, y_max)
np.sum(asphistimg[y_min:y_max+1, x_min:x_max+1])
np.sum(asphistimg)
aspecthistfh = pf.HDUList(
pf.PrimaryHDU(asphistimg[y_min:y_max+1, x_min:x_max+1])
)
aspecthistfh[0].header['EXTNAME'] = 'ASPECT_HISTOGRAM'
aspecthistfh[0].header['X_OFF'] = (x_min, 'x offset in pixels')
aspecthistfh[0].header['Y_OFF'] = (y_min, 'y offset in pixels')
aspecthistfh[0].header['EXPOSURE'] = (np.float32(np.sum(asphistimg[y_min:y_max+1, x_min:x_max+1])), 'seconds, total exposure time')
aspecthistfh[0].header['COMMENT'] = 'Add the x/y offset to image coordinates to recover aimpoint in detector coordinates.'
aspecthistfh[0].header['DATE'] = (datetime.datetime.utcnow().strftime('%Y-%m-%dT%H:%M:%S'), 'File creation date (YYYY-MM-DDThh:mm:ss UTC)')
aspecthistfh[0].header['HISTORY'] = 'Aspect histogram image created by filtering %s using %s.' % (
refpointfile, gtifile
)
for keyword in ('TELESCOP', 'INSTRUME', 'OBS_ID', 'OBJECT', 'TARG_ID',
'RA_OBJ', 'DEC_OBJ', 'RA_NOM', 'DEC_NOM', 'RA_PNT', 'DEC_PNT', 'TIMESYS',
'MJDREFI', 'MJDREFF', 'CLOCKAPP', 'TSTART', 'TSTOP', 'DATE-OBS', 'DATE-END'):
if keyword in refpointext.header:
aspecthistfh[0].header[keyword] = (refpointext.header[keyword], refpointext.header.comments[keyword])
aspecthistfh[0].header
aspecthistfh.writeto('aspecthistB.fits')
refpointext.header
```
| github_jupyter |
     
     
     
     
   
[Home Page](../../Start_Here.ipynb)
     
     
     
     
   
[1]
[2](../diffusion_1d/Diffusion_Problem_Notebook.ipynb)
[3](../spring_mass/Spring_Mass_Problem_Notebook.ipynb)
[4](../chip_2d/Challenge_CFD_Problem_Notebook.ipynb)
     
     
     
     
[Next Notebook](../diffusion_1d/Diffusion_Problem_Notebook.ipynb)
# Introduction to Physics Informed Neural Networks (PINNs)
In this notebook we will see the advantages of Physics Informed modeling over data-driven modeling and will also outline the brief theory about Physics Informed Neural Networks (PINNs).
## PINNs vs Data-Driven Methods
The data-driven methods are very useful in applications where the data is already available to learn from, or in situations where the underlying physics is not known. For such methods, the generalization errors are dependent on the amount of training data. Usually, obtaining the training data from simulations is a resource-hungry process and can limit the amounts of data the model is trained on. This places restrictions on the quality of results obtained from the data-driven methods and it usually takes a large amount of training data and fine-tuning of the model to capture the physics better.
<img src="image_data_driven_cons.png" alt="Drawing" style="width: 700px;"/>
Since for most of the physical systems, we know the underlying governing equations (e.g. Navier Stokes equations for the fluid mechanics problems), we can make the neural networks more cognizant of these physical laws by incorporating these governing equations in training framework. In such cases we can train the models on lesser data, or even train completely without any data from a solver. Also, in such Physics-Informed Neural Networks, treatment of inverse problems and parameterized problems (e.g. problems with parameterized geometry/material properties/boundary conditions) is easy. Due to these advantages, we will now see how to apply PINNs for physics-based problems using the SimNet library.
Let's start with discussing the theory behind the PINNs briefly.
## Neural Network Solver Methodology
In this section we provide a brief introduction to solving differential equations with neural networks. The idea is to use a neural network to approximate the solution to the given differential equation and boundary conditions. We train this neural network by constructing a loss function for how well the neural network is satisfying the differential equation and boundary conditions. If the network is able to minimize this loss function then it will in effect, solve the given differential equation.
To illustrate this idea we will give an example of solving the following problem,
\begin{equation} \label{1d_equation}
\mathbf{P} : \left\{\begin{matrix}
\frac{\delta^2 u}{\delta x^2}(x) = f(x), \\
\\
u(0) = u(1) = 0,
\end{matrix}\right.
\end{equation}
We start by constructing a neural network $u_{net}(x)$. The input to this network is a single value $x \in \mathbb{R}$ and its output is also a single value $u_{net}(x) \in \mathbb{R}$. We suppose that this neural network is infinitely differentiable, $u_{net} \in C^{\infty}$. The typical neural network used is a deep fully connected network where the activation functions are infinitely differentiable.
Next we need to construct a loss function to train this neural network. We easily encode the boundary conditions as a loss in the following way:
\begin{equation}
L_{BC} = u_{net}(0)^2 + u_{net}(1)^2
\end{equation}
For encoding the equations, we need to compute the derivatives of $u_{net}$. Using automatic differentiation we can do so and compute $\frac{\delta^2 u_{net}}{\delta x^2}(x)$. This allows us to write a loss function of the form:
\begin{equation} \label{sumation_loss}
L_{residual} = \frac{1}{N}\sum^{N}_{i=0} \left( \frac{\delta^2 u_{net}}{\delta x^2}(x_i) - f(x_i) \right)^2
\end{equation}
Where the $x_i$'s are a batch of points sampled in the interior, $x_i \in (0, 1)$. Our total loss is then $L = L_{BC} + L_{residual}$. Optimizers such as Adam are used to train this neural network. Given $f(x)=1$, the true solution is $\frac{1}{2}(x-1)x$. Upon solving the problem, you can obtain good agreement between the exact solution and the neural network solution as shown in Figure below.
<img src="single_parabola.png" alt="Drawing" style="width: 500px;"/>
## Parmeterized Problems
One important advantage of a neural network solver over traditional numerical methods is its ability to solve parameterized geometries. To illustrate this concept we solve a parameterized version of the above problem. Suppose we want to know how the solution to this equation changes as we move the position on the boundary condition $u(l)=0$. We can parameterize this position with a variable $l \in [1,2]$ and our equation now has the form,
\begin{equation} \label{1d_equation2}
\mathbf{P} : \left\{\begin{matrix}
\frac{\delta^2 u}{\delta x^2}(x) = f(x), \\
\\
u(0) = u(l) = 0,
\end{matrix}\right.
\end{equation}
To solve this parameterized problem we can have the neural network take $l$ as input, $u_{net}(x,l)$. The losses then take the form,
\begin{equation}
L_{residual} = \int_1^2 \int_0^l \left( \frac{\delta^2 u_{net}}{\delta x^2}(x,l) - f(x) \right)^2 dx dl \approx \left(\int_1^2 \int^l_0 dxdl\right) \frac{1}{N} \sum^{N}_{i=0} \left(\frac{\delta^2 u_{net}}{\delta x^2}(x_i, l_i) - f(x_i)\right)^2
\end{equation}
\begin{equation}
L_{BC} = \int_1^2 (u_{net}(0,l))^2 + (u_{net}(l,l) dl \approx \left(\int_1^2 dl\right) \frac{1}{N} \sum^{N}_{i=0} (u_{net}(0, l_i))^2 + (u_{net}(l_i, l_i))^2
\end{equation}
In figure below we see the solution to the differential equation for various $l$ values after optimizing the network on this loss. While this example problem is overly simplistic, the ability to solve parameterized geometries presents significant industrial value. Instead of performing a single simulation we can solve multiple designs at the same time and for reduced computational cost. More examples of this can be found in *SimNet User Guide Chapter 13*.
<img src="every_parabola.png" alt="Drawing" style="width: 500px;"/>
## Inverse Problems
Another useful application of a neural network solver is solving inverse problems. In an inverse problem, we start with a set of observations and then use those observations to calculate the causal factors that produced them. To illustrate how to solve inverse problems with a neural network solver, we give the example of inverting out the source term $f(x)$ from same equation from above problem. Suppose we are given the solution $u_{true}(x)$ at 100 random points between 0 and 1 and we want to determine the $f(x)$ that is causing it. We can do this by making two neural networks $u_{net}(x)$ and $f_{net}(x)$ to approximate both $u(x)$ and $f(x)$. These networks are then optimized to minimize the following losses;
\begin{equation}
L_{residual} \approx \left(\int^1_0 dx\right) \frac{1}{N} \sum^{N}_{i=0} \left(\frac{\delta^2 u_{net}}{\delta x^2}(x_i, l_i) - f_{net}(x_i)\right)^2
\end{equation}
\begin{equation}
L_{data} = \frac{1}{100} \sum^{100}_{i=0} (u_{net}(x_i) - u_{true}(x_i))^2
\end{equation}
Using the function $u_{true}(x)=\frac{1}{48} (8 x (-1 + x^2) - (3 sin(4 \pi x))/\pi^2)$ the solution for $f(x)$ is $x + sin(4 \pi x)$. We solve this problem and compare the results in Figures below.
Comparison of true solution for $f(x)$ and the function approximated by the NN:
<img src="inverse_parabola.png" alt="Drawing" style="width: 500px;"/>
Comparison of $u_{net}(x)$ and the train points from $u_{true}$:
<img src="inverse_parabola_2.png" alt="Drawing" style="width: 500px;"/>
More examples of solving an inverse problem can be found in the *SimNet User Guide Chapter 12*.
```
from IPython.display import IFrame, display
#filepath = "http://wikipedia.org" # works with websites too!
filepath = "SimNet_v21.06_User_Guide.pdf"
IFrame(filepath, width=700, height=400)
```
# Licensing
This material is released by NVIDIA Corporation under the Creative Commons Attribution 4.0 International (CC BY 4.0)
     
     
     
     
   
[Home Page](../../Start_Here.ipynb)
     
     
     
     
   
[1]
[2](../diffusion_1d/Diffusion_Problem_Notebook.ipynb)
[3](../spring_mass/Spring_Mass_Problem_Notebook.ipynb)
[4](../chip_2d/Challenge_CFD_Problem_Notebook.ipynb)
     
     
     
     
[Next Notebook](../diffusion_1d/Diffusion_Problem_Notebook.ipynb)
| github_jupyter |
# More on strings
# 1) Introduction
Strings are a special type of list. One can use most of list properties, with the exception that once they are instantiated they can not be modified.
```
animals="python alligator"
print(animals)
print("Look at list length:",len(animals))
print("substring :",animals[0:6]) #
print("substrings :",animals[7:])
print("single element :",animals[1])
animals[1]="3"
```
There are different special characters that we have already seen:
- \n gives a new line
- \t gives a tabulation
- \' will display a ' in the output string
- \" will display a " in the output string
- \\ will display a \ in the output string
```
print("A new line.\nFollowed by a tab \t and many other characters: \', \", \\")
```
When one need to write in a shell over multiple lines, the usage of 3": """ will keep the formatting, for instance:
```
x="""cat
dog
4"""
x
```
## 2) Methods specific to strings:
Here are a few methods specifics to strings:
- .upper(): string in capital.
- .lower(): string in lower cap.
- .capitalize(): capitalize the first letter of a string.
- .split(delimiter): cut the string in different substrings using the delimiter char. Default char is a white space " ".
- .find(str): find and return the position of str in the string.
- .count(str): count the number of occurence of str in the string.
- .replace(str1,str2): replace str1 by str2 in the string.
- .startswith(str): verify if the string starts with str
- .strip(): remove white space and new line at the beginning and end of the string.
The whole list can be accessed using the dir() function.
```
print("upper: good ->","good".upper())
print("lower: GOOD ->","GOOD".lower())
print("capitalize: good ->","good".capitalize())
print("split(): I go to the school ->","I go to the school".split())
print("find(go): I go to the school ->","I go to the school".find("go"))
print("count(o): I go to the school ->","I go to the school".count("o"))
print("replace(go, went): I go to the school ->","I go to the school".replace("go","went"))
print("startswith(I): I go to the school ->","I go to the school".startswith("I"))
print("startswith(go): I go to the school ->","I go to the school".startswith("go"))
print("strip(I): I go to the school \n \n \n -> "," I go to the school \n ".strip())
```
## 3) Extracting numerical values from a string:
A common task in Python consist to read a string of characters that come from a file, for instance, and extracts several information out of it.
For instance let's consider the following string, where we want to extract the different numbers and add them together:
```
val = "3.14 2.71 operator"
lst_val=val.split()
print(lst_val)
summed_val=lst_val[0]+lst_val[1] #This is not correct, since both are strings
print(summed_val)
summed_val=float(lst_val[0])+float(lst_val[1]) #Both numbers needs to be casted in floats.
print(summed_val)
```
## 4) Converting a list of character in a string:
It is possible to easily merge a list of characters in a string, using the .join() method:
```
seq = ["A", "T", "G", "A", "T"]
seq1="-".join(seq)
seq2=" ".join(seq)
seq3="".join(seq)
print(seq)
print(seq1)
print(seq2)
print(seq3)
```
## 5) Exercises:
- From the following list: animals = ["Bird", "Lion", "Tiger", "Porc"] ,with a loop print for each element it's size.
- Define a function that can convert a integer into a string and print it in console.
- Define a function that can receive two integral numbers in string form and compute their sum and then print it in console.
- Define a function that can receive two integral numbers in string form and compute their sum and then print it in console.
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/Image/download.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/Image/download.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://mybinder.org/v2/gh/giswqs/earthengine-py-notebooks/master?filepath=Image/download.ipynb"><img width=58px src="https://mybinder.org/static/images/logo_social.png" />Run in binder</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/Image/download.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API and geemap
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geemap](https://github.com/giswqs/geemap). The **geemap** Python package is built upon the [ipyleaflet](https://github.com/jupyter-widgets/ipyleaflet) and [folium](https://github.com/python-visualization/folium) packages and implements several methods for interacting with Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, and `Map.centerObject()`.
The following script checks if the geemap package has been installed. If not, it will install geemap, which automatically installs its [dependencies](https://github.com/giswqs/geemap#dependencies), including earthengine-api, folium, and ipyleaflet.
**Important note**: A key difference between folium and ipyleaflet is that ipyleaflet is built upon ipywidgets and allows bidirectional communication between the front-end and the backend enabling the use of the map to capture user input, while folium is meant for displaying static data only ([source](https://blog.jupyter.org/interactive-gis-in-jupyter-with-ipyleaflet-52f9657fa7a)). Note that [Google Colab](https://colab.research.google.com/) currently does not support ipyleaflet ([source](https://github.com/googlecolab/colabtools/issues/60#issuecomment-596225619)). Therefore, if you are using geemap with Google Colab, you should use [`import geemap.eefolium`](https://github.com/giswqs/geemap/blob/master/geemap/eefolium.py). If you are using geemap with [binder](https://mybinder.org/) or a local Jupyter notebook server, you can use [`import geemap`](https://github.com/giswqs/geemap/blob/master/geemap/geemap.py), which provides more functionalities for capturing user input (e.g., mouse-clicking and moving).
```
# Installs geemap package
import subprocess
try:
import geemap
except ImportError:
print('geemap package not installed. Installing ...')
subprocess.check_call(["python", '-m', 'pip', 'install', 'geemap'])
# Checks whether this notebook is running on Google Colab
try:
import google.colab
import geemap.eefolium as emap
except:
import geemap as emap
# Authenticates and initializes Earth Engine
import ee
try:
ee.Initialize()
except Exception as e:
ee.Authenticate()
ee.Initialize()
```
## Create an interactive map
The default basemap is `Google Satellite`. [Additional basemaps](https://github.com/giswqs/geemap/blob/master/geemap/geemap.py#L13) can be added using the `Map.add_basemap()` function.
```
Map = emap.Map(center=[40,-100], zoom=4)
Map.add_basemap('ROADMAP') # Add Google Map
Map
```
## Add Earth Engine Python script
```
# Add Earth Engine dataset
# Get a download URL for an image.
image1 = ee.Image('srtm90_v4')
path = image1.getDownloadUrl({
'scale': 30,
'crs': 'EPSG:4326',
'region': '[[-120, 35], [-119, 35], [-119, 34], [-120, 34]]'
})
print(path)
vis_params = {'min': 0, 'max': 3000}
Map.addLayer(image1, vis_params)
```
## Display Earth Engine data layers
```
Map.addLayerControl() # This line is not needed for ipyleaflet-based Map.
Map
```
| github_jupyter |
# Clustering
```
#TODO: remove after development
%load_ext autoreload
%autoreload 2
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns; sns.set()
import pandas as pd
pd.set_option('display.max_columns', 5400)
pd.set_option('max_colwidth', 50)
```
## Loading data
```
#seed_name = 'hair_dryer'
#seed_name = 'video_codec'
#seed_name = 'diesel'
#seed_name = "contact_lens"
#seed_name = "contact_lens_us_c"
seed_name = "3d_printer"
src_dir = "."
import pickle
with open(src_dir + "/data/" + seed_name + "/semantic/tsne.pkl", 'rb') as infile:
df = pickle.load(infile)
print(df.shape)
df.head()
```
### Hierarchical clustering
```
%%time
import scipy.cluster.hierarchy as shc
clustering_features = df[["x", "y"]]
linkage = shc.linkage(clustering_features, method='average', metric='euclidean')
```
Based on this linkage we can plot the dendogram.<br>
The parameter `max_d` gives you the possibility to decide at which `x` value you want to split.
```
max_d = {}
#max_d["hair_dryer"] = {"big": 10, "medium": 3.5, "small": 1} avg incorrect
max_d["hair_dryer"] = {"big": 10, "medium": 6, "small": 3}
max_d["video_codec"] = {"big": 45, "medium": 18, "small": 9.7}
max_d["diesel"] = {"big": 45, "medium": 20, "small": 15}
max_d["contact_lens"] = {"big": 49.5, "medium": 23, "small": 12}
max_d["contact_lens_us_c"] = {"big": 46, "medium": 23, "small": 13} #avg correct weighed
#max_d["contact_lens_us_c"] = {"big": 40, "medium": 24, "small": 13} avg correct
#max_d["contact_lens"] = {"big": 40, "medium": 20, "small": 10} all countries
max_d["3d_printer"] = {"big": 35, "medium": 17, "small": 9}
plt.figure(figsize=(25, 10))
plt.title("Dendogram for dataset " + seed_name)
dend = shc.dendrogram(linkage, p=7, truncate_mode='level', orientation='left', show_leaf_counts=True, show_contracted=True)
for k, v in max_d[seed_name].items():
plt.axvline(x=v, c='k')
from nltk.stem.snowball import SnowballStemmer
from fiz_lernmodule.word2vec import Word2VecReader
def get_similar_words(word2vec_model, stemmer, labels):
augmented = {}
for label in labels:
if len(label.split(" ")) == 1: # word2vec model cannot work with n-grams
augmented[label] = []
similar = word2vec_model.find_similar(label, 10)
stemmed_label = stemmer.stem(label)
for s in similar:
word = s['word']
stemmed_word = stemmer.stem(word)
if stemmed_word != stemmed_label:
augmented[label].append(word)
return augmented
stemmer = SnowballStemmer("english")
w2v_loader = Word2VecReader(src_dir=src_dir)
w2v_model = w2v_loader.load_word_embeddings()
def filter_similar_by_occurence(tokens, augmented):
filtered = {}
for k, v in augmented.items():
filtered[k] = []
for word in v:
contains = tokens.map(lambda x: word in x)
num_contains = contains.values.sum()
if num_contains > len(tokens) / 10:
if word not in augmented.keys():
filtered[k].append(word)
return filtered
from collections import Counter
def collect_counts(col):
counter = Counter()
tokens = [item for sublist in col.values for item in sublist]
for token in tokens:
counter[token] +=1
return counter
%%time
from scipy.cluster.hierarchy import fcluster
from scipy import stats
number_of_key_terms = 15
count_first_n_terms = 10
augment_first_n_terms = 3
total_cluster_centers = pd.DataFrame(columns = ['id', "x", "y", "min_x", "min_y", 'max_x', 'max_y', 'count', 'labels', 'augmented_labels', 'level', 'ids'])
for cluster_size in ["big", "medium", "small"]:
fcluster_pred = fcluster(linkage, max_d[seed_name][cluster_size], criterion='distance')
cluster_indexes = pd.unique(fcluster_pred)
plot_data = pd.DataFrame(columns = ['x', "y", "labels", "count"])
cluster_centers = pd.DataFrame(columns = total_cluster_centers.columns)
for i in range(len(cluster_indexes)):
cluster = df[fcluster_pred == i + 1]
cooccurent = collect_counts(cluster["terms"].apply(lambda x: x[:count_first_n_terms]))
labels = [i[0] for i in cooccurent.most_common(number_of_key_terms)]
similar_words = get_similar_words(w2v_model, stemmer, labels[:augment_first_n_terms])
augmented = filter_similar_by_occurence(cluster["tokens"], similar_words)
ids = pd.unique(cluster.publication_number)
count = len(ids)
without_outliers = cluster[["x", "y"]]
#without_outliers = without_outliers[(np.abs(stats.zscore(without_outliers)) < 3).all(axis=1)]
min_x = np.min(without_outliers.x)
min_y = np.min(without_outliers.y)
max_x = np.max(without_outliers.x)
max_y = np.max(without_outliers.y)
x = np.mean(without_outliers.x)
y = np.mean(without_outliers.y)
d = {"id": [i], "x": [x], "y": [y], "min_x": [min_x], "min_y": [min_y], "max_x": [max_x], "max_y": [max_y], "count": [count],
'labels': [", ".join(labels)], 'level': cluster_size, 'ids': [ids], 'augmented_labels': [augmented]}
cluster_centers = cluster_centers.append(pd.DataFrame(data=d), sort=False)
plot_d = {"x": cluster.x, "y": cluster.y, "count": [1]*cluster.shape[0]}
cluster_plot_data = pd.DataFrame(data = plot_d)
cluster_plot_data["labels"] = ", ".join(labels)
plot_data = plot_data.append(cluster_plot_data, sort=False)
#cluster_centers.reset_index(inplace=True, drop=True)
plot_data_with_centroids = plot_data.append(cluster_centers, sort=False)
plot_data_with_centroids['count'] = pd.to_numeric(plot_data_with_centroids["count"])
fig, ax = plt.subplots(figsize=(12,10))
ax = sns.scatterplot(x="x", y="y", data=plot_data_with_centroids, hue="labels", size="count", sizes=(20, np.max(cluster_centers["count"])))
plt.title("Hierarchical clustering for dataset " + seed_name + " with " + cluster_size + " cluster size.\nMax_d = "
+ str(max_d[seed_name][cluster_size]) + ". Number of clusters: " + str(len(cluster_indexes)))
plt.legend(bbox_to_anchor=(0.5, -len(cluster_indexes)*0.03 - 0.2), loc=8, borderaxespad=0.)
plt.show()
cluster_centers["labels"] = cluster_centers['labels'].map(lambda x: x.split(", "))
#cluster_centers.index.rename("id", inplace=True)
#cluster_centers.reset_index(inplace=True)
total_cluster_centers = total_cluster_centers.append(cluster_centers,sort=False)
total_cluster_centers.to_json(src_dir + "/../demo/data/semantic/" + seed_name + "_clusters.json", orient="records", index=True) #write intermediate results to prevent long wait
print(total_cluster_centers.head())
total_cluster_centers.tail()
```
| github_jupyter |
```
#@title # [minDALL-E github](https://github.com/kakaobrain/minDALL-E)
#@markdown also checkout https://colab.research.google.com/drive/1Gg7-c7LrUTNfQ-Fk-BVNCe9kvedZZsAh?usp=sharing
#@markdown notebook by Annas
#@title check gpu type
#@markdown if you get k80 it would be really slow try again by clicking runtime -> factory reset runtime until you get T4
#@markdown if you can't get T4 it's okay
#@markdown but keep in mind T4 is 9x faster than k80
!nvidia-smi -L
#@title setup run this once
from IPython.display import clear_output
import torch
import torch.nn as nn
torch.cuda.empty_cache()
!pip install rudalle==0.4.0 > /dev/null
!git clone -q https://github.com/kakaobrain/minDALL-E.git
%cd minDALL-E/
!pip install -q tokenizers>=0.10.2
!pip install -q pyflakes>=2.2.0
!pip install -q tqdm>=4.46.0
!pip install -q pytorch-lightning>=1.5
!pip install -q einops
!pip install -q omegaconf
!pip install -q git+https://github.com/openai/CLIP.git
!pip install "torchmetrics<0.7"
clear_output()
#@title run this once also if you want to reload everything you can run this too
import os
from rudalle import get_realesrgan
from rudalle.pipelines import show, super_resolution
import torch
from dalle.utils.utils import set_seed
from dalle.models import Dalle
import clip
import math
torch.cuda.empty_cache()
device = "cuda" if torch.cuda.is_available() else "cpu"
with torch.no_grad():
model = Dalle.from_pretrained("minDALL-E/1.3B")
model.to(device=device)
torch.cuda.empty_cache()
realesrgan = get_realesrgan("x2", device=device)
torch.cuda.empty_cache()
model_clip, preprocess_clip = clip.load("ViT-B/32", device=device)
model_clip.to(device=device)
torch.cuda.empty_cache()
clear_output()
torch.cuda.empty_cache()
import clip
import torch
import numpy as np
from dalle.models import Dalle
from dalle.utils.utils import clip_score
from PIL import Image
#@markdown prompt for the AI
prompt = "avocado smilling. avocado smilling. avocado smilling. avocado smilling. avocado smilling. avocado smilling. avocado smilling. avocado smilling. avocado smilling." #@param {type:"string"}
#@markdown number of images to be generated
num_candidates = 16#@param {type:"integer"}
#@markdown images per batch
batch_size = 8#@param {type:"integer"}
#@markdown number of image to be showed after re-ranking with clip
num_show = 16 #@param {type:"integer"}
#@markdown set this to False to disable realesrgan
use_realesrgan = True #@param {type:"boolean"}
#@markdown if nonzero, limits the sampled tokens to the top k values
top_k = 128 #@param {type:"integer"}
#@markdown if nonzero, limits the sampled tokens to the cumulative probability
top_p = None #@param {type:"raw"}
#@markdown controls the "craziness" of the generation
temperature = 0.7 #@param {type:"number"}
images = []
print(prompt)
for i in range(int(num_candidates / batch_size)):
images.append(model.sampling(prompt=prompt,
top_k=top_k,
top_p=top_p,
softmax_temperature=temperature,
num_candidates=batch_size,
device=device).cpu().numpy())
torch.cuda.empty_cache()
images = np.concatenate(images)
images = np.transpose(images, (0, 2, 3, 1))
if num_candidates > 1:
rank = clip_score(prompt=prompt,
images=images,
model_clip=model_clip,
preprocess_clip=preprocess_clip,
device=device)
torch.cuda.empty_cache()
images = images[rank]
num_show = num_show if num_candidates >= num_show else num_candidates
images = [Image.fromarray((images[i] * 255).astype("uint8")) for i in range(num_show)]
if use_realesrgan:
images = super_resolution(images, realesrgan)
clear_output()
print(prompt)
show(images, int(math.ceil(num_show / math.sqrt(num_show)))) # perfect square when using square numbers by litevex#6982
```
| github_jupyter |
## Tesla and Gasoline
By John Loeber
Notebook released under the Creative Commons Attribution 4.0 License.
---
### Introduction
Is the stock price of Tesla Motors (TSLA) linked to the price of gasoline?
Some people think that cheap gasoline incentivizes the purchase of gas-fuelled cars, lowering demand for Teslas, thus causing a drop in TSLA stock price. I try to find out whether that is true: I investigate the relationship between the price of TSLA and the price of UGA, an ETF tracking the price of gasoline.
### Conclusion
The price of <strong>TSLA roughly follows the price of gasoline, with a lag of about 50 business days.</strong> However, statistical correlations show only very weak results, which is likely due to two reasons:
1. The price of TSLA is not only impacted by gas -- there are other factors: earnings reports, rumours, etc. They cause occasional (smaller) movements in TSLA that are not related to the price of gas.
2. The lag is not necessarily constant -- there's no reason why the offset should always be 50 business days. Sometimes TSLA may take longer to respond, and at other times it may respond more quickly. This makes it hard to fit the time-series to each other in a way that yields a strong correlation.
### Going Further
Use the result, that TSLA roughly follows the price of gasoline with a lag of about 50 business days, to construct a trading strategy. It will be worthwhile to [write an algorithm](https://www.quantopian.com/algorithms) to attempt to trade on this pattern.
To expand further upon this project:
- Test against crude oil and energy ETFs to search for other signals in the data.
- Query the [Quantopian Fundamentals](https://www.quantopian.com/help/fundamentals) and use quarterly sales/revenue data to see how these details correspond to the price of gasoline.
- Consider commodity prices from NYMEX or other potentially more informative datasets.
Have questions? Post to the [community](https://www.quantopian.com/posts) or send us an email: feedback@quantopian.com.
## Investigation
I'll use [Kalman filters](https://www.quantopian.com/posts/quantopian-lecture-series-kalman-filters) to obtain moving averages, both of TSLA and of the ETFs. I overlay these moving average plots and look for relationships between the two (using both the obtained moving averages and the raw data).
```
# Import libraries
from matplotlib import pyplot
from pykalman import KalmanFilter
import numpy
import scipy
import time
import datetime
# Initialize a Kalman Filter.
# Using kf to filter does not change the values of kf, so we don't need to ever reinitialize it.
kf = KalmanFilter(transition_matrices = [1],
observation_matrices = [1],
initial_state_mean = 0,
initial_state_covariance = 1,
observation_covariance=1,
transition_covariance=.01)
# helper functions
# for converting dates to a plottable form
def convert_date(mydate):
# return time.mktime(datetime.datetime.strptime(mydate, "%Y-%m-%d").timetuple())
return datetime.datetime.strptime(mydate, "%Y-%m-%d")
# for grabbing dates and prices for a relevant equity
def get_data(equity_name,trading_start,trading_end='2015-07-20'):
# using today as a default arg.
stock_data = get_pricing(equity_name,
start_date = trading_start,
end_date = trading_end,
fields = ['close_price'],
frequency = 'daily')
stock_data['date'] = stock_data.index
# drop nans. For whatever reason, nans were causing the kf to return a nan array.
stock_data = stock_data.dropna()
# the dates are just those on which the prices were recorded
dates = stock_data['date']
dates = [convert_date(str(x)[:10]) for x in dates]
prices = stock_data['close_price']
return dates, prices
# TSLA started trading on Jun-29-2010.
dates_tsla, scores_tsla = get_data('TSLA','2010-06-29')
# Apply Kalman filter to get a rolling average
scores_tsla_means, _ = kf.filter(scores_tsla.values)
```
I'll plot the daily TSLA price and its Kalman-filtered moving average on it, simply to get a feel for the data.
```
# Use a scatterplot instead of a line plot because a line plot would be far too noisy.
pyplot.scatter(dates_tsla,scores_tsla,c='gray',label='TSLA Price')
pyplot.plot(dates_tsla,scores_tsla_means, c='red', label='TSLA MA')
pyplot.ylabel('TSLA Price')
pyplot.xlabel('Date')
pyplot.ylim([0,300])
pyplot.legend(loc=2)
pyplot.show()
```
This plot raises an important consideration: in 2012, Tesla debuted the Model S and delivered 2650 vehicles. It was only midway through 2013 that Tesla rose to prominence, having dramatically increased their production output. Thus, the period between 2010 and mid-2013 is likely misleading/irrelevant. In this investigation, I will concentrate on data from June 2013 onwards.
### UGA
UGA is the ETF tracking the price of gasoline.
```
# Get UGA data and apply the Kalman filter to get a moving average.
dates_uga, scores_uga = get_data('UGA','2013-06-01')
scores_uga_means, _ = kf.filter(scores_uga.values)
# Get TSLA for June 2013 onwards, and apply the Kalman Filter.
dates_tsla2, scores_tsla2 = get_data('TSLA','2013-06-01')
scores_tsla_means2, _ = kf.filter(scores_tsla2.values)
```
We'll now plot the TSLA price (moving average) and the UGA price (moving average).
```
_, ax1 = pyplot.subplots()
ax1.plot(dates_tsla2,scores_tsla_means2, c='red', label='TSLA MA')
pyplot.xlabel('Date')
pyplot.ylabel('TSLA Price MA')
pyplot.legend(loc=2)
# twinx allows us to use the same plot
ax2 = ax1.twinx()
ax2.plot(dates_uga, scores_uga_means, c='black', label='UGA Price MA')
pyplot.ylabel('UGA Price MA')
pyplot.legend(loc=3)
pyplot.show()
```
You can immediately see a strong, though perhaps lagged correspondence. If you line up the most prominent peaks and troughs in prices, you can see that the two time-series appear to correlate strongly, albeit with a lag of what looks like about two or three months. We'll use a function to find the lag that maximizes correlation.
```
def find_offset(ts1,ts2,window):
""" Finds the offset between two equal-length timeseries that maximizies correlation.
Window is # of days by which we want to left- or right-shift.
N.B. You'll have to adjust the function for negative correlations."""
l = len(ts1)
if l!=len(ts2):
raise Exception("Error! Timeseries lengths not equal!")
max_i_spearman = -1000
max_spearman = -1000
spear_offsets = []
# we try all possible offsets from -window to +window.
# we record the spearman correlation for each offset.
for i in range(window,0,-1):
series1 = ts1[i:]
series2 = ts2[:l-i]
# spearmanr is a correlation test
spear = scipy.stats.spearmanr(series1,series2)[0]
spear_offsets.append(spear)
if spear > max_spearman:
# update best correlation
max_spearman = spear
max_i_spearman = -i
for i in range(0,window):
series1 = ts1[:l-i]
series2 = ts2[i:]
spear = scipy.stats.spearmanr(series1,series2)[0]
spear_offsets.append(spear)
if spear > max_spearman:
max_spearman = spear
max_i_spearman = i
print "Max Spearman:", max_spearman, " At offset: ", max_i_spearman
pyplot.plot(range(-window,window),spear_offsets, c='green', label='Spearman Correlation')
pyplot.xlabel('Offset Size (Number of Business Days)')
pyplot.ylabel('Spearman Correlation')
pyplot.legend(loc=3)
pyplot.show()
print "Kalman-Filtered Smoothed Data"
find_offset(scores_tsla_means2,scores_uga_means,200)
print "Raw Data"
find_offset(scores_tsla2,scores_uga,150)
```
These plots are not promising at all! I want to find a strong positive correlation between TSLA and UGA, but neither in the smoothed nor in the raw data is there a strong, positive correlation. However, I did find that negative offsets of 126 and 50 days are correlation-maximizing, so we'll take a look at the smoothed data with these offsets.
```
# plotting formalities for 126-day offset
d = 126
cseries1 = scores_tsla_means2[d:]
cseries2 = scores_uga_means[:len(scores_tsla_means2)-d]
r = range(len(cseries1))
_, ax1 = pyplot.subplots()
ax1.plot(r, cseries1, c='red', label='TSLA MA')
pyplot.xlabel('Number of Business Days Elapsed')
pyplot.ylabel('TSLA Price MA')
pyplot.legend(loc=2)
ax2 = ax1.twinx()
ax2.plot(r, cseries2, c='black', label='UGA Price MA')
pyplot.ylabel('UGA Price MA')
pyplot.legend(loc=4)
pyplot.title("-126 Day Offset")
pyplot.show()
# plotting for 50-day offset
d = 50
cseries1 = scores_tsla_means2[d:]
cseries2 = scores_uga_means[:len(scores_tsla_means2)-d]
r = range(len(cseries1))
_, ax1 = pyplot.subplots()
ax1.plot(r, cseries1, c='red', label='TSLA MA')
pyplot.xlabel('Number of Business Days Elapsed')
pyplot.ylabel('TSLA Price MA')
pyplot.legend(loc=2)
ax2 = ax1.twinx()
ax2.plot(r, cseries2, c='black', label='UGA Price MA')
pyplot.ylabel('UGA Price MA')
pyplot.legend(loc=4)
pyplot.title("-50 Day Offset")
pyplot.show()
```
It is my opinion that the 50-day offset is a better fit. Not only do the 50-day plots visually appear to correspond more strongly (we can line up all major movements), it is also more plausible that gas price leads Tesla by 50 business days, rather than by 120 business days. However, looking at the 50-day plot, it is nonetheless surprising that the Spearman correlation is so low:
```
print scipy.stats.spearmanr(scores_tsla_means2[d:][250:],scores_uga_means[:len(scores_tsla_means2)-d][250:])
```
A correlation of 0.08 is statistically extremely weak -- basically signifying no relationship. To me, the visual relationship looks stronger than that. I guessed that there could be some noise obscuring the correlation, so I applied a second round of Kalman filters to both datasets. Doing this, I was able to drive the maximum correlation up to 0.20, but that's still not as strong as we would want.
I think the issue is twofold:
1. The <strong>price of TSLA is not only impacted by gas</strong> -- there are obviously other factors, like earnings reports, rumours, etc. This causes occasional movements in price that are not related to the price of gas.
2. The <strong>offset is not necessarily constant</strong> -- in the plot above, you can see that some movements seem not to line up entirely (which, of course, has disastrous consequences for the numerical correlation), and that's to be expected: there's no reason why the offset should always be 50 business days -- sometimes TSLA may take longer to respond, and at other times it may respond more quickly.
In any case, I consider the above plot quite compelling -- I will devote a future investigation to attempting to trade on the lag between TSLA and UGA. I invite you to do the same.
| github_jupyter |
```
import tensorflow as tf
from tools import get_file, get_data_file
from PIL import Image
import numpy as np
import matplotlib.pyplot as plt
detection_graph = tf.Graph()
with detection_graph.as_default():
od_graph_def = tf.GraphDef()
with tf.gfile.GFile(get_file('bin/inference_graph/frozen_inference_graph.pb'), 'rb') as fid:
serialized_graph = fid.read()
od_graph_def.ParseFromString(serialized_graph)
tf.import_graph_def(od_graph_def, name='')
def load_image_into_numpy_array(image):
(im_width, im_height) = image.size
return np.array(image.getdata()).reshape(
(im_height, im_width, 3)).astype(np.uint8)
IMAGE_SIZE = (10, 10)
def run_inference_for_single_image(image, graph):
with graph.as_default():
with tf.Session() as sess:
# Get handles to input and output tensors
ops = tf.get_default_graph().get_operations()
all_tensor_names = {output.name for op in ops for output in op.outputs}
tensor_dict = {}
for key in [
'num_detections', 'detection_boxes', 'detection_scores',
'detection_classes', 'detection_masks'
]:
tensor_name = key + ':0'
if tensor_name in all_tensor_names:
tensor_dict[key] = tf.get_default_graph().get_tensor_by_name(
tensor_name)
if 'detection_masks' in tensor_dict:
# The following processing is only for single image
detection_boxes = tf.squeeze(tensor_dict['detection_boxes'], [0])
detection_masks = tf.squeeze(tensor_dict['detection_masks'], [0])
# Reframe is required to translate mask from box coordinates to image coordinates and fit the image size.
real_num_detection = tf.cast(tensor_dict['num_detections'][0], tf.int32)
detection_boxes = tf.slice(detection_boxes, [0, 0], [real_num_detection, -1])
detection_masks = tf.slice(detection_masks, [0, 0, 0], [real_num_detection, -1, -1])
detection_masks_reframed = utils_ops.reframe_box_masks_to_image_masks(
detection_masks, detection_boxes, image.shape[1], image.shape[2])
detection_masks_reframed = tf.cast(
tf.greater(detection_masks_reframed, 0.5), tf.uint8)
# Follow the convention by adding back the batch dimension
tensor_dict['detection_masks'] = tf.expand_dims(
detection_masks_reframed, 0)
image_tensor = tf.get_default_graph().get_tensor_by_name('image_tensor:0')
# Run inference
output_dict = sess.run(tensor_dict,
feed_dict={image_tensor: image})
# all outputs are float32 numpy arrays, so convert types as appropriate
output_dict['num_detections'] = int(output_dict['num_detections'][0])
output_dict['detection_classes'] = output_dict[
'detection_classes'][0].astype(np.int64)
output_dict['detection_boxes'] = output_dict['detection_boxes'][0]
output_dict['detection_scores'] = output_dict['detection_scores'][0]
if 'detection_masks' in output_dict:
output_dict['detection_masks'] = output_dict['detection_masks'][0]
return output_dict
import sys
print(sys.path)
sys.path.append('/Users/antonioferegrino/Documents/GitHub/salsa-valentina/src/external')
print(sys.path)
from external.object_detection.utils import visualization_utils as vis_util
image = Image.open(get_data_file('external', 'test1.jpg'))
# the array based representation of the image will be used later in order to prepare the
# result image with boxes and labels on it.
image_np = load_image_into_numpy_array(image)
# Expand dimensions since the model expects images to have shape: [1, None, None, 3]
image_np_expanded = np.expand_dims(image_np, axis=0)
# Actual detection.
output_dict = run_inference_for_single_image(image_np_expanded, detection_graph)
# Visualization of the results of a detection.
vis_util.visualize_boxes_and_labels_on_image_array(
image_np,
output_dict['detection_boxes'],
output_dict['detection_classes'],
output_dict['detection_scores'],
{1:{'name':'Valentina'}},
instance_masks=output_dict.get('detection_masks'),
use_normalized_coordinates=True,
line_thickness=8)
plt.figure(figsize=IMAGE_SIZE)
plt.imshow(image_np)
plt.savefig(f'test1.jpg')
plt.show()
```
| github_jupyter |
#### New to Plotly?
Plotly's Python library is free and open source! [Get started](https://plotly.com/python/getting-started/) by downloading the client and [reading the primer](https://plotly.com/python/getting-started/).
<br>You can set up Plotly to work in [online](https://plotly.com/python/getting-started/#initialization-for-online-plotting) or [offline](https://plotly.com/python/getting-started/#initialization-for-offline-plotting) mode, or in [jupyter notebooks](https://plotly.com/python/getting-started/#start-plotting-online).
<br>We also have a quick-reference [cheatsheet](https://images.plot.ly/plotly-documentation/images/python_cheat_sheet.pdf) (new!) to help you get started!
#### Imports
The tutorial below imports [NumPy](http://www.numpy.org/), [Pandas](https://plotly.com/pandas/intro-to-pandas-tutorial/), [SciPy](https://www.scipy.org/) and [PeakUtils](http://pythonhosted.org/PeakUtils/).
```
import plotly.plotly as py
import plotly.graph_objs as go
from plotly.tools import FigureFactory as FF
import numpy as np
import pandas as pd
import scipy
import peakutils
```
#### Import Data
To start detecting peaks, we will import some data on milk production by month:
```
milk_data = pd.read_csv('https://raw.githubusercontent.com/plotly/datasets/master/monthly-milk-production-pounds.csv')
time_series = milk_data['Monthly milk production (pounds per cow)']
time_series = time_series.tolist()
df = milk_data[0:15]
table = FF.create_table(df)
py.iplot(table, filename='milk-production-dataframe')
```
#### Original Plot
```
trace = go.Scatter(
x = [j for j in range(len(time_series))],
y = time_series,
mode = 'lines'
)
data = [trace]
py.iplot(data, filename='milk-production-plot')
```
#### With Peak Detection
We need to find the x-axis indices for the peaks in order to determine where the peaks are located.
```
cb = np.array(time_series)
indices = peakutils.indexes(cb, thres=0.02/max(cb), min_dist=0.1)
trace = go.Scatter(
x=[j for j in range(len(time_series))],
y=time_series,
mode='lines',
name='Original Plot'
)
trace2 = go.Scatter(
x=indices,
y=[time_series[j] for j in indices],
mode='markers',
marker=dict(
size=8,
color='rgb(255,0,0)',
symbol='cross'
),
name='Detected Peaks'
)
data = [trace, trace2]
py.iplot(data, filename='milk-production-plot-with-peaks')
```
#### Only Highest Peaks
We can attempt to set our threshold so that we identify as many of the _highest peaks_ that we can.
```
cb = np.array(time_series)
indices = peakutils.indexes(cb, thres=0.678, min_dist=0.1)
trace = go.Scatter(
x=[j for j in range(len(time_series))],
y=time_series,
mode='lines',
name='Original Plot'
)
trace2 = go.Scatter(
x=indices,
y=[time_series[j] for j in indices],
mode='markers',
marker=dict(
size=8,
color='rgb(255,0,0)',
symbol='cross'
),
name='Detected Peaks'
)
data = [trace, trace2]
py.iplot(data, filename='milk-production-plot-with-higher-peaks')
from IPython.display import display, HTML
display(HTML('<link href="//fonts.googleapis.com/css?family=Open+Sans:600,400,300,200|Inconsolata|Ubuntu+Mono:400,700" rel="stylesheet" type="text/css" />'))
display(HTML('<link rel="stylesheet" type="text/css" href="http://help.plot.ly/documentation/all_static/css/ipython-notebook-custom.css">'))
! pip install git+https://github.com/plotly/publisher.git --upgrade
import publisher
publisher.publish(
'python-Peak-Finding.ipynb', 'python/peak-finding/', 'Peak Finding | plotly',
'Learn how to find peaks and valleys on datasets in Python',
title='Peak Finding in Python | plotly',
name='Peak Finding',
language='python',
page_type='example_index', has_thumbnail='false', display_as='peak-analysis', order=3,
ipynb= '~notebook_demo/120')
```
| github_jupyter |
# Tensorflow: IBACopy
This notebook shows how to use the [IBACopy]() class. It is very flexible and should work with different models.
For classification models, you might want to look at [IBACopyInnvestigate]().
If you can add additional layers to your model, it might also be worth looking at [IBALayer]().
Ensure that `./imagenet` points to your copy of the ImageNet dataset.
You might want to create a symlink:
```
# ! ln -s /path/to/your/imagenet/folder/ imagenet
# select your device
%env CUDA_VISIBLE_DEVICES=1
# reduce tensorflow noise
import warnings
warnings.filterwarnings("ignore")
import sys
import os
from tqdm.notebook import tqdm
import matplotlib.pyplot as plt
import numpy as np
from PIL import Image
import tensorflow.compat.v1 as tf
import keras
import keras.backend as K
from keras.applications.resnet50 import preprocess_input, ResNet50
from keras.applications import VGG16
from skimage.transform import resize
from IBA.utils import plot_saliency_map
from IBA.tensorflow_v1 import IBACopy, TFWelfordEstimator, model_wo_softmax, to_saliency_map
config = tf.ConfigProto()
config.gpu_options.allow_growth=True
sess = tf.Session(config=config)
keras.backend.set_session(sess)
print("TensorFlow version: {}, Keras version: {}".format(
tf.version.VERSION, keras.__version__))
# data loading
def get_imagenet_generator(val_dir, image_size = (224, 224), shuffle=True, batch_size=50):
image_generator = tf.keras.preprocessing.image.ImageDataGenerator(
preprocessing_function=preprocess_input
)
return image_generator.flow_from_directory(
val_dir, shuffle=shuffle, seed=0, batch_size=batch_size, target_size=image_size)
def norm_image(x):
return (x - x.min()) / (x.max() - x.min())
imagenet_dir = "./imagenet"
imagenet_val_dir = os.path.join(imagenet_dir, "validation")
img_batch, target_batch = next(get_imagenet_generator(imagenet_val_dir))
monkey_pil = Image.open("monkeys.jpg").resize((224, 224))
monkey = preprocess_input(np.array(monkey_pil))[None]
monkey_target = 382 # 382: squirrel monkey
# load model
model_softmax = VGG16(weights='imagenet')
# remove the final softmax layer
model = model_wo_softmax(model_softmax)
# select layer after which the bottleneck will be inserted
feat_layer = model.get_layer(name='block4_conv1')
```
Create the Analyzer
```
iba = IBACopy(feat_layer.output, model.output)
```
Double check if model was copied correctly
```
# checks if all variables are equal in the original graph and the copied graph
iba.assert_variables_equal()
iba_logits = iba.predict({model.input: img_batch})
model_logits = model.predict(img_batch)
assert (np.abs(iba_logits - model_logits).mean()) < 1e-5
print(np.abs(iba_logits - model_logits).mean())
```
Fit mean and std of the feature map
```
feed_dict_gen = map(lambda x: {model.input: x[0]},
get_imagenet_generator(imagenet_val_dir))
iba.fit_generator(feed_dict_gen, n_samples=5000)
print("Fitted estimator on {} samples".format(iba._estimator.n_samples()))
# set classification loss
target = iba.set_classification_loss()
# you can also specificy your own loss with :
# iba.set_model_loss(my_loss)
iba.set_default(beta=10)
# get the saliency map
capacity = iba.analyze(
feature_feed_dict={model.input: monkey},
copy_feed_dict={target: np.array([monkey_target])}
)
saliency_map = to_saliency_map(capacity, shape=(224, 224))
saliency_map.shape
K.image_data_format()
plot_saliency_map(saliency_map, img=norm_image(monkey[0]))
```
## Access to internal values
You can access all intermediate values of the optimzation through the `iba.get_report()` method.
To store the intermediate values, you have to call either `iba.collect_all()` or `iba.collect(*var_names)` before running `iba.analyze(..)`.
```
# collect all intermediate tensors
iba.collect_all()
# storing all tensors can slow down the optimization.
# you can also select to store only specific ones:
# iba.collect("alpha", "model_loss")
# to only collect a subset all all tensors
# run the optimization
capacity = iba.analyze(
feature_feed_dict={model.input: monkey},
copy_feed_dict={iba.target: np.array([monkey_target])}
)
# get all saved outputs
report = iba.get_report()
```
`report` is an `OrderedDict` which maps each `iteration` to a dictionray of `{var_name, var_value}`.
The `init` iteration is computed without an optimizer update. Values not changing such as the feature values are only included in the `init` iteration.
The `final` iteration is again computed without an optimizer update.
```
print("iterations:", list(report.keys()))
```
Print all available tensors in the `init` iteration:
```
print("{:<30} {:}".format("name:", "shape"))
print()
for name, val in report['init'].items():
print("{:<30} {:}".format(name + ":", str(val.shape)))
```
### Losses during optimization
```
fig, ax = plt.subplots(1, 2, figsize=(8, 3))
ax[0].set_title("cross entrop loss")
ax[0].plot(list(report.keys()), [it['model_loss'] for it in report.values()])
ax[1].set_title("mean capacity")
ax[1].plot(list(report.keys()), [it['capacity_mean'] for it in report.values()])
```
### Distribution of alpha (pre-softmax) values per iteraton
```
cols = 6
rows = len(report) // cols
fig, axes = plt.subplots(rows, cols, figsize=(2.8*cols, 2.2*rows))
for ax, (it, values) in zip(axes.flatten(), report.items()):
ax.hist(values['alpha'].flatten(), log=True, bins=20)
ax.set_title("iteration: " + str(it))
plt.subplots_adjust(wspace=0.3, hspace=0.5)
fig.suptitle("distribution of alpha (pre-softmax) values per iteraton.", y=1)
plt.show()
```
### Distributiuon of the final capacity
```
plt.hist(report['final']['capacity'].flatten(), bins=20, log=True)
plt.title("Distributiuon of the final capacity")
plt.show()
```
| github_jupyter |
```
# Importing libraries
import json
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from pandasql import sqldf
from sklearn.preprocessing import MinMaxScaler
min_max_scaler = MinMaxScaler()
# To read the datsets on Google Colab
# !pip install -U -q PyDrive
# from pydrive.auth import GoogleAuth
# from pydrive.drive import GoogleDrive
# from google.colab import auth
# from oauth2client.client import GoogleCredentials
# # Authenticate and create the PyDrive client.
# auth.authenticate_user()
# gauth = GoogleAuth()
# gauth.credentials = GoogleCredentials.get_application_default()
# drive = GoogleDrive(gauth)
# file_id = '1V_tKI0Vtd50lzXt77sWLBbpMM2ZCC0hQ'
# downloaded = drive.CreateFile({'id': file_id})
# downloaded.GetContentFile('deliveries.csv')
# file_id1 = '1HQsQxgXgY2KUGNbKv2ZGXPs_fjnvIZwY'
# downloaded1 = drive.CreateFile({'id': file_id1})
# downloaded1.GetContentFile('matches.csv')
```
<h2> PREPROCESSING </h2>
```
df = pd.read_csv('deliveries.csv')
df.columns
df2 = pd.read_csv('matches.csv')
df2.columns
df.head()
df2.head()
# To find which stadiums are repeating with a slightly different name
l = sorted([i for i in df2['venue'].unique()])
print(l)
# Correcting stadium names
df2.replace('M. Chinnaswamy Stadium', 'M Chinnaswamy Stadium', inplace=True)
df2.replace('ACA-VDCA Stadium', 'Dr. Y.S. Rajasekhara Reddy ACA-VDCA Cricket Stadium', inplace=True)
df2.replace('Feroz Shah Kotla Ground', 'Feroz Shah Kotla', inplace=True)
df2.replace('Subrata Roy Sahara Stadium', 'Maharashtra Cricket Association Stadium', inplace=True)
df2.replace('M. A. Chidambaram Stadium', 'MA Chidambaram Stadium, Chepauk', inplace=True)
df2.replace('IS Bindra Stadium', 'Punjab Cricket Association IS Bindra Stadium, Mohali', inplace=True)
df2.replace('Punjab Cricket Association Stadium, Mohali', 'Punjab Cricket Association IS Bindra Stadium, Mohali', inplace=True)
df2.replace('Rajiv Gandhi Intl. Cricket Stadium', 'Rajiv Gandhi International Stadium, Uppal', inplace=True)
df2.replace('Dr. Y.S. Rajasekhara Reddy ACA-VDCA Cricket Stadium', 'Dr YS Rajasekhara Reddy ACA VDCA Cricket Stadium', inplace=True)
# Confirming that there are no null values where there shouldn't be any
df.isnull().sum()
df2.isnull().sum()
# The 4 NaN values in winner is because of No Result (due to rain)
# Correcting the date format as it was different for a few seasons
for i in range(len(df2)):
if df2["season"][i] == 2018 or df2["season"][i] == 2019:
date = df2["date"][i].split("/")
date = "20" + date[2] + "-" + date[1] + "-" + date[0]
df2.loc[i,"date"] = date #updating the format
# Creating the matches dictionary
matches_dict = list()
with open('data.json') as f:
data = json.load(f)
for k in data.keys():
for i in data[k].keys():
winner = data[k][i]["winner"]
date = data[k][i]["date"]
venue = data[k][i]["venue"]
team1 = data[k][i]["team1"]["name"]
team2 = data[k][i]["team2"]["name"]
players1 = data[k][i]["team1"]["players"]
bowler_count_1 = 0
for j in players1:
if 'bowlingStyle' in j.keys():
bowler_count_1 += 1
batsman_count_1 = 11 - bowler_count_1
players1 = data[k][i]["team2"]["players"]
bowler_count_2 = 0
for j in players1:
if 'bowlingStyle' in j.keys():
bowler_count_2 += 1
batsman_count_2 = 11 - bowler_count_2
matches_dict.append({"winner" : " ".join(winner.split(" ")[:-4]), \
"date": date[:10], "venue": venue, \
"team1" : team1, \
"bowler_count_1": bowler_count_1, \
"batsman_count_1": batsman_count_1, \
"team2" : team2, \
"bowler_count_2" : bowler_count_2, \
"batsman_count_2" : batsman_count_2})
# Correcting the venue/stadium names
new_data = pd.DataFrame(matches_dict)
new_data.replace('ACA-VDCA Stadium', 'Dr YS Rajasekhara Reddy ACA VDCA Cricket Stadium', inplace=True)
new_data.replace('DY Patil Stadium', 'Dr DY Patil Sports Academy', inplace=True)
new_data.replace('Feroz Shah Kotla Ground', 'Feroz Shah Kotla', inplace=True)
new_data.replace('HPCA Stadium', 'Himachal Pradesh Cricket Association Stadium', inplace=True)
new_data.replace('IS Bindra Stadium', 'Punjab Cricket Association IS Bindra Stadium, Mohali', inplace=True)
new_data.replace('JSCA International Cricket Stadium', 'JSCA International Stadium Complex', inplace=True)
new_data.replace('Jawaharlal Nehru Stadium', 'Nehru Stadium', inplace=True)
new_data.replace('M. Chinnaswamy Stadium', 'M Chinnaswamy Stadium', inplace=True)
new_data.replace('M. A. Chidambaram Stadium', 'MA Chidambaram Stadium, Chepauk', inplace=True)
new_data.replace("Maharashtra Cricket Association's International Stadium", 'Maharashtra Cricket Association Stadium', inplace=True)
new_data.replace('Rajiv Gandhi Intl. Cricket Stadium', 'Rajiv Gandhi International Stadium, Uppal', inplace=True)
new_data.replace('Sardar Patel Stadium', 'Sardar Patel Stadium, Motera', inplace=True)
new_data.replace('Zayed Cricket Stadium', 'Sheikh Zayed Stadium', inplace=True)
new_data.replace('Shaheed Veer Narayan Singh International Cricket Stadium', 'Shaheed Veer Narayan Singh International Stadium', inplace=True)
new_data.replace('Vidarbha Cricket Association Stadium', 'Vidarbha Cricket Association Stadium, Jamtha', inplace=True)
new_data
# Creating players dataframe
player_details = pd.DataFrame({})
for i in data.keys():
for j in data[i].keys():
y = pd.DataFrame(data[i][j]['team1']['players'])
player_details = pd.concat([player_details, y])
player_details.drop_duplicates(inplace=True)
player_details.reset_index(inplace=True)
player_details
df2[df2.result=='tie']
# Checking number of tied matches for superovers
abc= df[df['is_super_over']==1].groupby(by='match_id').count()
print(abc)
# Number of superovers played (=7, but there were 9 tied matches)
# Further investigation has to be performed, we can see that match ID 11146 and 11342 have is_super_over=0 in inning=3 and 4
df['inning'].unique()
# innings=5 is wrong
df[((df['inning']==3)|(df['inning']==4)|(df['inning']==5))].groupby(by='inning').count()
# There are 8 balls bowled with inning = 5, should be the ones that are missing with is_super_over=1
for i in range(df.shape[0]):
if df.iloc[i, 1]==3:
df.iloc[i, 9]=1 # cleaning: inning=3, is_super_over is made as 1
elif df.iloc[i,1]==5:
df.iloc[i,1]=4 # cleaning: inning=5 is made 4
sorted(list(df2.team1.unique()))
# "Rising Pune Supergiants" and "Rising Pune Supergiant" are treated as separate teams like "Delhi Daredevils" and "Delhi Capitals"
```
<h2> VISUALISATION </h2>
```
# Visualization 1- Head to head analysis of teams
h2h = dict()
def add_to_h2h(team1, team2, winner):
if team1 in h2h.keys(): #if it is the first time, the team name has appeared as the 'home' team
if team2 in h2h[team1]: #if it is the first time, the team name has appeared as the 'away' team
if winner==team1:
h2h[team1][team2][0]+=1
elif winner==team2:
h2h[team1][team2][2]+=1
else:
h2h[team1][team2][1]+=1 #no result matches
else: #adding the first record for the away team (vs the home team)
if winner==team1:
h2h[team1][team2]=[1,0,0]
elif winner==team2:
h2h[team1][team2]=[0,0,1]
else:
h2h[team1][team2]=[0,1,0]
else: #adding the first record for the home team
h2h[team1]=dict()
if winner==team1:
h2h[team1][team2]=[1,0,0]
elif winner==team2:
h2h[team1][team2]=[0,0,1]
else:
h2h[team1][team2]=[0,1,0]
for i in range(df2.shape[0]):
if df2.iloc[i,10] is not None:
add_to_h2h(df2.iloc[i, 4], df2.iloc[i, 5], df2.iloc[i, 10])
add_to_h2h(df2.iloc[i, 5], df2.iloc[i, 4], df2.iloc[i, 10])
else:
add_to_h2h(df2.iloc[i, 4], df2.iloc[i, 5], df2.iloc[i, 10])
def head_to_head(team): #this function displays the given team's head-to-head visualization against all other teams
plt.figure(figsize=(16,10))
teams=sorted(list(df2.team1.unique()))
color=['yellow','midnightblue','royalblue','darkblue','orangered','crimson','darkorange','purple','blue','dodgerblue','fuchsia','mediumvioletred','mediumvioletred','red','orange']
for i in range(len(teams)):
if teams[i]==team: #assigning a color to each team (similar to jersey colour)
team_color=color[i]
break
for team2, team2_color in zip(teams, color):
if team2==team: #skipping the record of a team against itself since it doesn't make sense
continue
if team2 not in h2h[team].keys():
continue
t1,nr,t2 = h2h[team][team2]
t1_ = 100*t1/(t1+nr+t2) #Win percentage for the given team against the team in the iteration
nr_ = 100*nr/(t1+nr+t2) #Percentage of matches between them that ended with No Result
t2_ = 100*t2/(t1+nr+t2) #Loss percentage
plt.barh([team2+'\n'+str(t1)+' '+str(nr)+' '+str(t2)], t1_, color=team_color, height=0.8)
plt.barh([team2+'\n'+str(t1)+' '+str(nr)+' '+str(t2)], nr_, left=t1_, color='grey', height=0.8)
plt.barh([team2+'\n'+str(t1)+' '+str(nr)+' '+str(t2)], t2_, left=(t1_+nr_), color=team2_color, height=0.8)
plt.show()
head_to_head('Royal Challengers Bangalore')
head_to_head('Chennai Super Kings')
# Analysis- Average wickets taken per team over the years
def get_wickets(df, team):
match_id = 0
opponents = None
num_wickets = 0
wickets = dict()
for i in range(len(df)):
if df['bowling_team'][i] == team:
if match_id == 0:
match_id = df['match_id'][i]
opponents = df['batting_team'][i]
if match_id == df['match_id'][i]:
if type(df['player_dismissed'][i]) != float:
num_wickets += 1
else:
if opponents in wickets:
wickets[opponents] += [num_wickets]
else:
wickets[opponents] = [num_wickets]
opponents = df['batting_team'][i]
match_id = df['match_id'][i]
num_wickets = 0
if type(df['player_dismissed'][i]) != float:
num_wickets += 1
for i in wickets:
wickets[i] = round(np.average(wickets[i]),0)
return wickets
output = dict()
for i in df['batting_team'].unique():
row = get_wickets(df, i)
output[i] = row
final_out = pd.DataFrame(output, index = list(df['batting_team'].unique()))
final_out
# Analysis- Average runs scored per team over the years
def get_runs(df, team):
match_id = 0
opponents = None
runs = 0
run_dict = dict()
for i in range(len(df)):
if df['batting_team'][i] == team:
if match_id == 0:
match_id = df['match_id'][i]
opponents = df['bowling_team'][i]
if match_id == df['match_id'][i]:
if type(df['total_runs'][i]) != float:
runs += df['total_runs'][i]
else:
if opponents in run_dict:
run_dict[opponents] += [runs]
else:
run_dict[opponents] = [runs]
opponents = df['bowling_team'][i]
match_id = df['match_id'][i]
runs = 0
if type(df['total_runs'][i]) != float:
runs += df['total_runs'][i]
for i in run_dict:
run_dict[i] = round(np.average(run_dict[i]),0)
return run_dict
output = dict()
for i in df['batting_team'].unique():
row = get_runs(df, i)
output[i] = row
final_out = pd.DataFrame(output, index = list(df['batting_team'].unique()))
final_out
# Toss Analysis- Win percentage on winning toss for each venue counted over groups, grouped by toss decision
def toss_win(df, venue):
match_count = field_count = field_winner_count = bat_count = bat_winner_count = 0
for i in range(len(df)):
if df['venue'][i] == venue:
if df['toss_decision'][i] == 'field':
field_count += 1
if df['toss_winner'][i] == df['winner'][i]:
field_winner_count += 1
if df['toss_decision'][i] == 'bat':
bat_count += 1
if df['toss_winner'][i] == df['winner'][i]:
bat_winner_count += 1
try:
field_winner_prob = round(field_winner_count/field_count, 2)
except:
field_winner_prob = np.nan
try:
bat_winner_prob = round(bat_winner_count/bat_count, 2)
except:
bat_winner_prob = np.nan
return({'bat': bat_winner_prob, 'field' : field_winner_prob})
venues = dict()
for i in df2['venue'].unique():
venues[i] = toss_win(df2, i)
pd.DataFrame(venues).T
# Create a dataset of all players' batting details (those who have batted atleast once) using the ball-by-ball dataset
def get_batting_records(df):
batsmen = {}
match=1 #match count being held to see if the batsman remained not out, and to calculate 100s and 50s as we iterate
for i in range(df.shape[0]):
if df.iloc[i,6] not in batsmen.keys():
batsmen[df.iloc[i,6]]={'temp':0, 'temp2':0, 'Runs':0, 'Balls':0, 'Fifties':0, 'Hundreds':0, 'Fours':0, 'Sixes':0, 'Dismissals':0, 'caught':0, 'bowled':0, 'run out':0, 'lbw':0, 'caught and bowled':0, 'stumped':0, 'retired hurt':0, 'hit wicket':0, 'obstructing the field':0, 'Innings':0}
batsmen[df.iloc[i,6]]['Runs']+=df.iloc[i,15] #df.iloc[i,6] gives the name of the batsmen
batsmen[df.iloc[i,6]]['temp']+=df.iloc[i,15] #2 temporary columns used for calculation of 100s and 50s
if df.iloc[i,15]==4:
batsmen[df.iloc[i,6]]['Fours']+=1
if df.iloc[i,15]==6:
batsmen[df.iloc[i,6]]['Sixes']+=1
if df.iloc[i,10]==0 and df.iloc[i,13]==0:
batsmen[df.iloc[i,6]]['Balls']+=1 #to add 1 ball for each "legal" delivery faced (not a wide or no-ball)
else: #note: wides balls are never considered in a batsman's tally
if df.iloc[i,15]>0 and i+1<df.shape[0] and df.iloc[i,5] != df.iloc[i+1,5]:
batsmen[df.iloc[i,6]]['Balls']+=1 #to account for no-balls
if df.iloc[i,13]>0 and i+1<df.shape[0] and df.iloc[i,5] != df.iloc[i+1,5]:
batsmen[df.iloc[i,6]]['Balls']+=2 #extra if conditions to take care of different format being used to store extras
elif df.iloc[i,13]>0 and i+1<df.shape[0] and df.iloc[i,5] == df.iloc[i+1,5]:
batsmen[df.iloc[i,6]]['Balls']+=1
if df.iloc[i,6]==df.iloc[i,18]:
batsmen[df.iloc[i,6]]['Dismissals']+=1
batsmen[df.iloc[i,6]][df.iloc[i,19]]+=1
if df.iloc[i,0] != batsmen[df.iloc[i,6]]['temp2']: #to check if match has changed, can update 100s, 50s then
if batsmen[df.iloc[i,6]]['temp']>=100:
batsmen[df.iloc[i,6]]['Hundreds']+=1
elif batsmen[df.iloc[i,6]]['temp']>=50:
batsmen[df.iloc[i,6]]['Fifties']+=1
batsmen[df.iloc[i,6]]['temp']=0
batsmen[df.iloc[i,6]]['temp2'] = df.iloc[i,0] #updating the new match number into the temp column for the batsman
batsmen[df.iloc[i,6]]['Innings']+=1
match=df.iloc[i,0]
#After iterating fully, we go through the dict and calculate the various metrics such as strike rate, average
for name in batsmen.keys():
batsmen[name]['Strike Rate']=(100 * batsmen[name]['Runs'] / batsmen[name]['Balls'])
if batsmen[name]['Dismissals']>0:
batsmen[name]['Average']=(batsmen[name]['Runs'] / batsmen[name]['Dismissals'])
batting_records = pd.DataFrame(batsmen)
batting_records = batting_records.T
batting_records.drop(["temp", "temp2"], axis=1, inplace=True)
batting_records.loc[batting_records.Average.isna() == True , "Average"] = batting_records.Runs / batting_records.Innings #A fitting used for NaNs (it is an approximation)
return batting_records
batting_records = get_batting_records(df)
batting_records
# Plotting Dismisssal Analysis
kind=df.groupby('dismissal_kind').count()
t=dict(kind["fielder"][kind.fielder!=0])
plt.bar(t.keys(),t.values())
# Most Caught Dismissal analysis of 10 batsmen dismissal analysis of 10 players in the history of IPL
catch=df[df.dismissal_kind=="caught"].groupby("fielder").count()
f=catch.sort_values(by=["match_id"],ascending=False).head(10)
f1=dict(f["match_id"])
plt.figure(figsize=(20,10))
plt.bar(f1.keys(),f1.values(),width=0.4)
# Most Stumped Dismissal analysis of 10 batsmen in the history of IPL
stump=df[df.dismissal_kind=="stumped"].groupby("fielder").count()
s=stump.sort_values(by=["match_id"],ascending=False).head(10)
s1=dict(s["match_id"])
plt.figure(figsize=(20,10))
plt.bar(s1.keys(),s1.values())
# Create a dataset of all players' bowling details (those who have bowled atleast once) using the ball-by-ball dataset
def get_bowling_records(df):
bowler = {}
match=1 #similar to batting records above, used to calculate if a bowler took 4 or 5 wicket hauls in thematch
for i in range(df.shape[0]):
if df.iloc[i,8] not in bowler.keys():
bowler[df.iloc[i,8]]={'temp':0, 'temp2':0, 'temp3':0, 'temp4':0, 'Runs':0, 'Balls':0, 'Wickets':0, '4+':0, '5+':0, 'Dots':0, 'Maidens':0, 'Wides':0, 'No balls':0, 'caught':0, 'bowled':0, 'lbw':0, 'caught and bowled':0, 'stumped':0, 'hit wicket':0, 'Innings':0}
bowler[df.iloc[i,8]]['Runs']+=(df.iloc[i,15]+df.iloc[i,10]+df.iloc[i,13])
bowler[df.iloc[i,8]]['Wides']+=df.iloc[i,10]
bowler[df.iloc[i,8]]['No balls']+=df.iloc[i,13]
if df.iloc[i,17] == 0:
bowler[df.iloc[i,8]]['Dots']+=1
bowler[df.iloc[i,8]]['temp4']+=1
else:
bowler[df.iloc[i,8]]['temp4']=-200
if df.iloc[i,10]==0 and df.iloc[i,13]==0:
bowler[df.iloc[i,8]]['Balls']+=1
else:
if df.iloc[i,15]>0 and i+1<df.shape[0] and df.iloc[i,5] != df.iloc[i+1,5]:
bowler[df.iloc[i,8]]['Balls']+=1 #accounting for no-balls
if df.iloc[i,13]>0 and i+1<df.shape[0] and df.iloc[i,5] != df.iloc[i+1,5]:
bowler[df.iloc[i,8]]['Balls']+=2 #extra condition due to different formats used in the dataset
elif df.iloc[i,13]>0 and i+1<df.shape[0] and df.iloc[i,5] == df.iloc[i+1,5]:
bowler[df.iloc[i,8]]['Balls']+=1
if df.iloc[i,19] in ['caught', 'bowled', 'lbw', 'caught and bowled', 'stumped', 'hit wicket']:
bowler[df.iloc[i,8]]['Wickets']+=1
bowler[df.iloc[i,8]]['temp']+=1
bowler[df.iloc[i,8]][df.iloc[i,19]]+=1 #adding the type/kind of wicket the bowler has taken
if bowler[df.iloc[i,8]]['temp2'] == df.iloc[i,0] and bowler[df.iloc[i,8]]['temp3'] != df.iloc[i,4]:
if bowler[df.iloc[i,8]]['temp4']==6: #used to check for maidens, end of each over
bowler[df.iloc[i,8]]['Maidens']+=1
bowler[df.iloc[i,8]]['temp4']=0 #set for next over
bowler[df.iloc[i,8]]['temp3']=df.iloc[i,4] #setting new over number
if bowler[df.iloc[i,8]]['temp2'] != df.iloc[i,0]:
if bowler[df.iloc[i,8]]['temp']>=5: #used to check for 5 wicket hauls, end of each match
bowler[df.iloc[i,8]]['5+']+=1
elif bowler[df.iloc[i,8]]['temp']>=4: #4 wicket hauls
bowler[df.iloc[i,8]]['4+']+=1
bowler[df.iloc[i,8]]['temp']=0 #reset for next over
bowler[df.iloc[i,8]]['temp2'] = df.iloc[i,0]
bowler[df.iloc[i,8]]['temp3']=0
bowler[df.iloc[i,8]]['temp4']=0
bowler[df.iloc[i,8]]['Innings']+=1
match=df.iloc[i,0]
#after iterating through the entire ball-by-ball dataset, we iterate through all the bowlers in the dict to caluclate values for various metrics such as average, economy, strike rate
for name in bowler.keys():
bowler[name]['Economy']=(6 * bowler[name]['Runs'] / bowler[name]['Balls'])
if bowler[name]['Wickets']>0:
bowler[name]['Strike_Rate']=(bowler[name]['Balls'] / bowler[name]['Wickets'])
bowler[name]['Average']=(bowler[name]['Runs'] / bowler[name]['Wickets'])
bowling_records = pd.DataFrame(bowler)
bowling_records = bowling_records.T
bowling_records.drop(["temp", "temp2", "temp3", "temp4"], axis=1, inplace=True)
bowling_records.loc[bowling_records.Strike_Rate.isna() == True , "Strike_Rate"] = bowling_records.Balls
bowling_records.loc[bowling_records.Average.isna() == True , "Average"] = 999 #Some fixing for the outliers, using 999 instead of infinity (NaN)
return bowling_records
bowling_records = get_bowling_records(df)
bowling_records
# Checking Bowling economy records
economy=bowling_records.sort_values(by=["Economy"]).head(10)
s1=dict(economy.Economy)
plt.figure(figsize=(20,10))
plt.bar(s1.keys(),s1.values())
'''
strike_rate=batting_records.sort_values(by=['Strike Rate'], ascending=True).head(10)
s2=dict(strike_rate['Strike Rate'])
plt.figure(figsize=(20,10))
plt.bar(s2.keys(),s2.values())
'''
latest_data=pd.merge(df, df2[df2.season>=2018], left_on='match_id', right_on='id', validate='many_to_one')
latest_data.drop(["umpire1","umpire2","umpire3","venue","dl_applied","player_of_match","win_by_wickets","win_by_runs","id"], axis=1, inplace=True)
latest_data.sort_values(by=['match_id', 'inning', 'over', 'ball'],ascending=False)
```
<h2>BATSMAN RATING</h2>
```
latest_batting_records = get_batting_records(latest_data)
def get_overall_batting_score(batting_records):
df = pd.DataFrame(batting_records["Innings"]).rename(columns = {"Innings" : "matches"})
df["matches"] = min_max_scaler.fit_transform(df[["matches"]])
df["achievement_weight"] = 20 * batting_records["Hundreds"] + 10 * batting_records["Fifties"] + 3 * batting_records["Sixes"] + batting_records["Fours"]
df["stat_score"] = df["achievement_weight"]*0.3 + batting_records["Average"]*0.55 + batting_records["Strike Rate"]*0.15
df["overall_score"] = df["stat_score"] * df["matches"]
return df.sort_values(by=["overall_score", "matches", "achievement_weight", "stat_score"])
def get_batsman_rating(batting_records, latest_batting_records):
overall_score = get_overall_batting_score(batting_records)
latest_score = get_overall_batting_score(latest_batting_records)
batsman_rating = pd.merge(overall_score[["overall_score"]], latest_score[["overall_score"]], how="outer", left_index=True, right_index=True, suffixes=["_career", "_latest"]).fillna(0)
batsman_rating["batsman_rating"] = 0.4*batsman_rating["overall_score_career"] + 0.6*batsman_rating["overall_score_latest"]
batsman_rating["batsman_rating"] = min_max_scaler.fit_transform(batsman_rating[["batsman_rating"]]) * 100
return batsman_rating[["batsman_rating"]].sort_values(by=["batsman_rating"], ascending=False)
batsman_rating = get_batsman_rating(batting_records, latest_batting_records)
batsman_rating
```
<h2>BOWLER RATING</h2>
```
latest_bowling_records = get_bowling_records(latest_data)
def get_overall_bowling_score(bowling_records):
df = pd.DataFrame(bowling_records["Innings"]).rename(columns = {"Innings" : "matches"})
df["matches"] = min_max_scaler.fit_transform(df[["matches"]])
df["achievement_weight"] = 5 * bowling_records["Maidens"] + bowling_records["Dots"]
df["wicket_weight"] = 30 * bowling_records["5+"] + 20 * bowling_records["4+"] + 10 * bowling_records["Wickets"]
df["stat_score"] = bowling_records["Average"] + bowling_records["Economy"] + 1 / (1+df["achievement_weight"])
df["overall_score"] = (df["wicket_weight"] * df["matches"]) / df["stat_score"]
return df.sort_values(by=["overall_score", "matches", "achievement_weight", "stat_score"])
def get_bowler_rating(bowling_records, latest_bowling_records):
overall_score = get_overall_bowling_score(bowling_records)
max_overall = overall_score["overall_score"].max()
overall_score["overall_score"] = overall_score["overall_score"] / max_overall
latest_score = get_overall_bowling_score(latest_bowling_records)
max_latest = latest_score["overall_score"].max()
latest_score["overall_score"] = latest_score["overall_score"] / max_latest
bowler_rating = pd.merge(overall_score[["overall_score"]], latest_score[["overall_score"]], how="outer", left_index=True, right_index=True, suffixes=["_career", "_latest"]).fillna(0)
bowler_rating["bowler_rating"] = 0.35*bowler_rating["overall_score_career"] + 0.65*bowler_rating["overall_score_latest"]
bowler_rating["bowler_rating"] = min_max_scaler.fit_transform(bowler_rating[["bowler_rating"]]) * 100
return bowler_rating[["bowler_rating"]].sort_values(by=["bowler_rating"], ascending=False)
bowler_rating = get_bowler_rating(bowling_records, latest_bowling_records)
bowler_rating
```
<h2> PLAYING 11 SELECTION USING DISTANCES </h2>
```
given_team = ["SP Narine", "A Mishra", "DJ Bravo", "RA Jadeja", \
"SL Malinga", "CH Gayle", "AB de Villiers", "MS Dhoni", \
"SK Raina", "V Kohli", "S Gopal", "Yuvraj Singh", "PP Ojha", \
"MP Stoinis", "DA Miller"]
given_team = pd.DataFrame(data={"Players": given_team})
def select_playing_11_euclidian(given_team, batsman_rating, bowler_rating):
temp = pd.merge(given_team, batsman_rating, left_on="Players", right_index=True)
temp = pd.merge(temp, bowler_rating, left_on="Players", right_index=True, how="left").fillna(0).reset_index(drop=True)
top_3_bowlers = temp.sort_values(by=["bowler_rating", "batsman_rating"], ascending=False).head(3)
top_3_batsman = temp.sort_values(by=["batsman_rating", "bowler_rating"], ascending=False).head(3)
selected_team = pd.concat([top_3_batsman, top_3_bowlers]).reset_index(drop=True)
temp2 = temp.apply(lambda x: temp.loc[~x.isin(selected_team[x.name]),x.name]).fillna(0)
euclid_distance = dict()
for i in temp2.index:
dist = ((100 - temp2.batsman_rating[i])**2 + (100 - temp2.bowler_rating[i])**2)**0.5
euclid_distance[temp2["Players"][i]] = dist
rest_5 = pd.DataFrame((list(euclid_distance.items()))).rename(columns = {0:"Players", 1:"Euclid_Distance"}).sort_values(by=["Euclid_Distance"])[["Players"]].head(5)
selected_team = pd.concat([selected_team[["Players"]], rest_5]).reset_index(drop=True)
selected_team = pd.merge(temp, selected_team, how="right")
return selected_team.sort_values(by="batsman_rating", ascending=False).reset_index(drop=True)
select_playing_11_euclidian(given_team, batsman_rating, bowler_rating)
def select_playing_11_city_block(given_team, batsman_rating, bowler_rating):
temp = pd.merge(given_team, batsman_rating, left_on="Players", right_index=True)
temp = pd.merge(temp, bowler_rating, left_on="Players", right_index=True, how="left").fillna(0).reset_index(drop=True)
top_3_bowlers = temp.sort_values(by=["bowler_rating", "batsman_rating"], ascending=False).head(3)
top_3_batsman = temp.sort_values(by=["batsman_rating", "bowler_rating"], ascending=False).head(3)
selected_team = pd.concat([top_3_batsman, top_3_bowlers]).reset_index(drop=True)
temp2 = temp.apply(lambda x: temp.loc[~x.isin(selected_team[x.name]),x.name]).fillna(0)
city_block = dict()
for i in temp2.index:
dist = abs(100 - temp2.batsman_rating[i]) + abs(100 - temp2.bowler_rating[i])
city_block[temp2["Players"][i]] = dist
rest_5 = pd.DataFrame((list(city_block.items()))).rename(columns = {0:"Players", 1:"city_block_distance"}).sort_values(by=["city_block_distance"])[["Players"]].head(5)
selected_team = pd.concat([selected_team[["Players"]], rest_5])
selected_team = pd.merge(temp, selected_team, how="right")
return selected_team.sort_values(by="batsman_rating", ascending=False).reset_index(drop=True)
select_playing_11_city_block(given_team, batsman_rating, bowler_rating)
def select_playing_11_cosine(given_team, batsman_rating, bowler_rating):
temp = pd.merge(given_team, batsman_rating, left_on="Players", right_index=True)
temp = pd.merge(temp, bowler_rating, left_on="Players", right_index=True, how="left").fillna(0).reset_index(drop=True)
top_3_bowlers = temp.sort_values(by=["bowler_rating", "batsman_rating"], ascending=False).head(3)
top_3_batsman = temp.sort_values(by=["batsman_rating", "bowler_rating"], ascending=False).head(3)
selected_team = pd.concat([top_3_batsman, top_3_bowlers]).reset_index(drop=True)
temp2 = temp.apply(lambda x: temp.loc[~x.isin(selected_team[x.name]),x.name]).fillna(0)
cosine_dist = dict()
for i in temp2.index:
bat = temp2.batsman_rating[i]
bowl = temp2.bowler_rating[i]
similarity = (100*bat + 100*bowl)/(((100**2 + bat**2)**0.5) * ((100**2 + bowl**2)**0.5))
cosine_dist[temp2["Players"][i]] = 1 - similarity
rest_5 = pd.DataFrame((list(cosine_dist.items()))).rename(columns = {0:"Players", 1:"Cosine_Distance"}).sort_values(by=["Cosine_Distance"])[["Players"]].head(5)
selected_team = pd.concat([selected_team[["Players"]], rest_5])
selected_team = pd.merge(temp, selected_team, how="right")
return selected_team.sort_values(by="batsman_rating", ascending=False).reset_index(drop=True)
select_playing_11_cosine(given_team, batsman_rating, bowler_rating)
def select_playing_11_chebychev(given_team, batsman_rating, bowler_rating):
temp = pd.merge(given_team, batsman_rating, left_on="Players", right_index=True)
temp = pd.merge(temp, bowler_rating, left_on="Players", right_index=True, how="left").fillna(0).reset_index(drop=True)
top_3_bowlers = temp.sort_values(by=["bowler_rating", "batsman_rating"], ascending=False).head(3)
top_3_batsman = temp.sort_values(by=["batsman_rating", "bowler_rating"], ascending=False).head(3)
selected_team = pd.concat([top_3_batsman, top_3_bowlers]).reset_index(drop=True)
temp2 = temp.apply(lambda x: temp.loc[~x.isin(selected_team[x.name]),x.name]).fillna(0)
chebychev_dist = dict()
for i in temp2.index:
bat = temp2.batsman_rating[i]
bowl = temp2.bowler_rating[i]
chebychev_dist[temp2["Players"][i]] = max(abs(100-bat), abs(100-bowl))
rest_5 = pd.DataFrame((list(chebychev_dist.items()))).rename(columns = {0:"Players", 1:"chebychev_dist"}).sort_values(by=["chebychev_dist"])[["Players"]].head(5)
selected_team = pd.concat([selected_team[["Players"]], rest_5])
selected_team = pd.merge(temp, selected_team, how="right")
return selected_team.sort_values(by="batsman_rating", ascending=False).reset_index(drop=True)
select_playing_11_chebychev(given_team, batsman_rating, bowler_rating)
def select_playing_11_canberra(given_team, batsman_rating, bowler_rating):
temp = pd.merge(given_team, batsman_rating, left_on="Players", right_index=True)
temp = pd.merge(temp, bowler_rating, left_on="Players", right_index=True, how="left").fillna(0).reset_index(drop=True)
top_3_bowlers = temp.sort_values(by=["bowler_rating", "batsman_rating"], ascending=False).head(3)
top_3_batsman = temp.sort_values(by=["batsman_rating", "bowler_rating"], ascending=False).head(3)
selected_team = pd.concat([top_3_batsman, top_3_bowlers]).reset_index(drop=True)
temp2 = temp.apply(lambda x: temp.loc[~x.isin(selected_team[x.name]),x.name]).fillna(0)
canberra_dist = dict()
for i in temp2.index:
bat = temp2.batsman_rating[i]
bowl = temp2.bowler_rating[i]
canberra_dist[temp2["Players"][i]] = abs(bat - bowl)/abs(bat + bowl)
rest_5 = pd.DataFrame((list(canberra_dist.items()))).rename(columns = {0:"Players", 1:"canberra_dist"}).sort_values(by=["canberra_dist"])[["Players"]].head(5)
selected_team = pd.concat([selected_team[["Players"]], rest_5])
selected_team = pd.merge(temp, selected_team, how="right")
return selected_team.sort_values(by="batsman_rating", ascending=False).reset_index(drop=True)
select_playing_11_canberra(given_team, batsman_rating, bowler_rating)
```
<h2> TOSS DECISION </h2>
```
def bat_or_field(match_data, team1, team2, venue):
toss_decision = df2[["team1", "team2", "toss_winner", "toss_decision", "winner", "venue"]]
toss_decision = toss_decision[(toss_decision["team1"] == team1) | (toss_decision["team2"] == team1)]
toss_decision_1 = toss_decision[(toss_decision["team1"] == team2) | (toss_decision["team2"] == team2)]
toss_decision_1 = toss_decision_1[toss_decision_1["toss_winner"] == team1]
toss_decision_1 = toss_decision_1[toss_decision_1["venue"] == venue]
toss_decision_2 = toss_decision[toss_decision["toss_winner"] == team1]
toss_decision_2 = toss_decision_2[toss_decision_2["venue"] == venue]
toss_decision_2 = toss_decision_2[toss_decision_2["winner"] == team1]
try:
bat_1 = toss_decision_1.groupby(["toss_decision", "winner"]).size()["bat"][team1]
except:
bat_1 = 0
try:
field_1 = toss_decision_1.groupby(["toss_decision", "winner"]).size()["field"][team1]
except:
field_1 = 0
try:
bat_2 = toss_decision_2.groupby(["toss_decision", "winner"]).size()["bat"][team1]
except:
bat_2 = 0
try:
field_2 = toss_decision_2.groupby(["toss_decision", "winner"]).size()["field"][team1]
except:
field_2 = 0
bat = 0.6 * bat_1 + 0.4 * bat_2
field = 0.6 * field_1 + 0.4 * field_2
if bat > field:
print("Bat")
else:
print("Field")
team1 = "Royal Challengers Bangalore"
team2 = "Sunrisers Hyderabad"
venue = "M Chinnaswamy Stadium"
bat_or_field(df2, team1, team2, venue)
```
<h2> FINAL TEAM SELECTION</h2>
```
def get_stadium_bias(data, venue):
bat = 0
bowl = 0
count = 0
for i in range(len(data)):
if data["venue"][i] == venue:
count += 1
if data["winner"][i] == data["team1"][i]:
bat += data["batsman_count_1"][i]
bowl += data["bowler_count_1"][i]
else:
bat += data["batsman_count_2"][i]
bowl += data["bowler_count_2"][i]
return (round(bowl/count), round(bat/count))
get_stadium_bias(new_data, 'M Chinnaswamy Stadium')
def latest_team_selecter(given_team, batsman_rating, bowler_rating, data, player_details, venue):
#finding nationality of each player
team_details = list()
for i in range(len(given_team)):
for j in range(len(player_details)):
if set(player_details['shortName'][j]).issuperset(given_team['Players'][i]) or set(given_team['Players'][i]).issuperset(player_details['shortName'][j]):
team_details.append({'Players': given_team['Players'][i], 'Nationality': player_details['nationality'][j]})
break
team_deatils = pd.DataFrame(team_details)
#merging details, batting and bowling rankings
temp = pd.merge(team_deatils, batsman_rating, left_on="Players", right_index=True)
temp = pd.merge(temp, bowler_rating, left_on="Players", right_index=True, how="left").fillna(0).reset_index(drop=True)
#selecting top 6 players(top 3 each)
top_3_bowlers = temp.sort_values(by=["bowler_rating", "batsman_rating"], ascending=False).head(3)
top_3_batsman = temp.sort_values(by=["batsman_rating", "bowler_rating"], ascending=False).head(3)
selected_team = pd.concat([top_3_batsman, top_3_bowlers]).reset_index(drop=True)
#seeing who's not selected yet
temp2 = temp.merge(selected_team.drop_duplicates(), on=['Players', 'Nationality', 'batsman_rating', 'bowler_rating'], how='left', indicator=True)
temp2 = temp2[temp2['_merge'] == 'left_only'].drop(columns='_merge')
#checking if we should have more bowlers than batsmen
bowler_bias, batsman_bias = get_stadium_bias(new_data, venue)
bowlers = 3 if bowler_bias > batsman_bias else 2
batsmen = 3 if bowler_bias < batsman_bias else 2
#accodingly select the remaining 5 players
remaining_bowlers = temp2.sort_values(by="bowler_rating", ascending=False).head(int(bowlers))
remaining_batsmen = temp2.sort_values(by="batsman_rating", ascending=False).head(int(batsmen))
selected_team = pd.merge(selected_team, remaining_batsmen, how="outer")
selected_team = pd.merge(selected_team, remaining_bowlers, how="outer")
selected_team = selected_team.sort_values(by="batsman_rating", ascending=False).reset_index(drop=True)
return selected_team
latest_team_selecter(given_team, batsman_rating, bowler_rating, new_data, player_details, venue="M Chinnaswamy Stadium")
def final_team_selecter(given_team, batsman_rating, bowler_rating, new_data, player_details, venue):
team_details = list()
for i in range(len(given_team)):
for j in range(len(player_details)):
if set(player_details['shortName'][j]).issuperset(given_team['Players'][i]) or set(given_team['Players'][i]).issuperset(player_details['shortName'][j]):
team_details.append({'Players': given_team['Players'][i], 'Nationality': player_details['nationality'][j]})
break
team_deatils = pd.DataFrame(team_details)
#merging details, batting and bowling rankings
temp = pd.merge(team_deatils, batsman_rating, left_on="Players", right_index=True)
temp = pd.merge(temp, bowler_rating, left_on="Players", right_index=True, how="left").fillna(0).reset_index(drop=True)
#selecting top 6 players(top 3 each)
top_3_bowlers = temp.sort_values(by=["bowler_rating", "batsman_rating"], ascending=False).head(3)
top_3_batsman = temp.sort_values(by=["batsman_rating", "bowler_rating"], ascending=False).head(3)
selected_team = pd.concat([top_3_batsman, top_3_bowlers]).reset_index(drop=True)
#seeing who's not selected yet
temp2 = temp.merge(selected_team.drop_duplicates(), on=['Players', 'Nationality', 'batsman_rating', 'bowler_rating'], how='left', indicator=True)
temp2 = temp2[temp2['_merge'] == 'left_only'].drop(columns='_merge')
#using the euclidian distacnce from perfect player to see all rounders
euclid_distance =list()
for i in temp2.index:
dist = ((100 - temp2.batsman_rating[i])**2 + (100 - temp2.bowler_rating[i])**2)**0.5
euclid_distance.append({"Players" : temp2["Players"][i], \
"Nationality" : temp2["Nationality"][i], \
"batsman_rating": temp2["batsman_rating"][i], \
"bowler_rating": temp2["bowler_rating"][i], \
"euclid_distance": dist})
euclid_distance = pd.DataFrame(euclid_distance)
#checking if we should have more bowlers than batsmen
bowler_bias, batsman_bias = get_stadium_bias(new_data, venue)
if bowler_bias >= 7:
bowlers = 4
batsmen = 1
elif batsman_bias >= 7:
bowlers = 1
batsmen = 4
else:
bowlers = bowler_bias - 3
batsmen = batsman_bias - 3
#choosing top 3 all rounders with high bowler/batsman rating respectively
remaining_bowlers = euclid_distance.query("bowler_rating > batsman_rating").sort_values(by="euclid_distance").head(int(bowlers))
remaining_batsmen = euclid_distance.query("bowler_rating < batsman_rating").sort_values(by="euclid_distance").head(int(batsmen))
selected_team = pd.merge(selected_team, remaining_batsmen, how="outer")
selected_team = pd.merge(selected_team, remaining_bowlers, how="outer")
selected_team = selected_team.sort_values(by="batsman_rating", ascending=False).reset_index(drop=True)
return selected_team[["Players", "Nationality", "batsman_rating", "bowler_rating"]]
final_team_selecter(given_team, batsman_rating, bowler_rating, new_data, player_details, venue="M Chinnaswamy Stadium")
```
| github_jupyter |
```
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
import os.path as op
from csv import writer
import math
import cmath
import pickle
import tensorflow as tf
from tensorflow import keras
from keras.models import Model,Sequential,load_model
from keras.layers import Input, Embedding
from keras.layers import Dense, Bidirectional
from keras.layers.recurrent import LSTM
import keras.metrics as metrics
import itertools
from tensorflow.python.keras.utils.data_utils import Sequence
from decimal import Decimal
from keras.layers import Conv1D,MaxPooling1D,Flatten,Dense
A1=np.empty((0,5),dtype='float32')
U1=np.empty((0,7),dtype='float32')
node=['150','149','147','144','142','140','136','61']
mon=['Apr','Mar','Aug','Jun','Jul','Sep','May','Oct']
for j in node:
for i in mon:
inp= pd.read_csv('../../../data_gkv/AT510_Node_'+str(j)+'_'+str(i)+'19_OutputFile.csv',usecols=[1,2,3,15,16])
out= pd.read_csv('../../../data_gkv/AT510_Node_'+str(j)+'_'+str(i)+'19_OutputFile.csv',usecols=[5,6,7,8,17,18,19])
inp=np.array(inp,dtype='float32')
out=np.array(out,dtype='float32')
A1=np.append(A1, inp, axis=0)
U1=np.append(U1, out, axis=0)
print(A1)
print(U1)
from sklearn.decomposition import PCA
import warnings
scaler_obj1=PCA(svd_solver='full')
scaler_obj2=PCA(svd_solver='full')
X1=scaler_obj1.fit_transform(A1)
Y1=scaler_obj2.fit_transform(U1)
warnings.filterwarnings(action='ignore', category=UserWarning)
X1=X1[:,np.newaxis,:]
Y1=Y1[:,np.newaxis,:]
from keras import backend as K
def rmse(y_true, y_pred):
return K.sqrt(K.mean(K.square(y_pred - y_true), axis=-1))
model = Sequential()
model.add(keras.Input(shape=(1,5)))
model.add(tf.keras.layers.GRU(14,activation="tanh",use_bias=True,kernel_initializer="glorot_uniform",bias_initializer="zeros",
kernel_regularizer=keras.regularizers.l1_l2(l1=1e-5, l2=1e-4),
bias_regularizer=keras.regularizers.l2(1e-4),
activity_regularizer=keras.regularizers.l2(1e-5)))
model.add(keras.layers.Dropout(.1))
model.add(Dense(7))
model.add(keras.layers.BatchNormalization(axis=-1,momentum=0.99,epsilon=0.001,center=True,scale=True,
beta_initializer="zeros",gamma_initializer="ones",
moving_mean_initializer="zeros",moving_variance_initializer="ones",trainable=True))
model.add(keras.layers.ReLU())
model.compile(optimizer=keras.optimizers.Adam(learning_rate=1e-5),loss='mse',metrics=['accuracy','mse','mae',rmse])
model.summary()
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(X1, Y1, test_size=0.25, random_state=42)
history2 = model.fit(x_train,y_train,batch_size=256,epochs=50, validation_split=0.1)
model.evaluate(x_test,y_test)
model.evaluate(x_train,y_train)
df1=pd.DataFrame(history2.history['loss'],columns=["Loss"])
df1=df1.join(pd.DataFrame(history2.history["val_loss"],columns=["Val Loss"]))
df1=df1.join(pd.DataFrame(history2.history["accuracy"],columns=['Accuracy']))
df1=df1.join(pd.DataFrame(history2.history["val_accuracy"],columns=['Val Accuracy']))
df1=df1.join(pd.DataFrame(history2.history["mse"],columns=['MSE']))
df1=df1.join(pd.DataFrame(history2.history["val_mse"],columns=['Val MSE']))
df1=df1.join(pd.DataFrame(history2.history["mae"],columns=['MAE']))
df1=df1.join(pd.DataFrame(history2.history["val_mae"],columns=['Val MAE']))
df1=df1.join(pd.DataFrame(history2.history["rmse"],columns=['RMSE']))
df1=df1.join(pd.DataFrame(history2.history["val_mse"],columns=['Val RMSE']))
df1
df1.to_excel("GRU_tanh_mse.xlsx")
model_json = model.to_json()
with open("gru_tanh_mse.json", "w") as json_file:
json_file.write(model_json)
# serialize weights to HDF5
model.save_weights("gru_tanh_mse.h5")
print("Saved model to disk")
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(X1, Y1, test_size=0.25, random_state=42)
from keras.models import model_from_json
json_file = open('gru_tanh_mse.json', 'r')
loaded_model_json = json_file.read()
json_file.close()
loaded_model = model_from_json(loaded_model_json)
# load weights into new model
loaded_model.load_weights("gru_tanh_mse.h5")
print("Loaded model from disk")
loaded_model.compile(optimizer=keras.optimizers.Adam(learning_rate=1e-5),loss='mse',metrics=['accuracy','mse','mae',rmse])
print(loaded_model.evaluate(x_train, y_train, verbose=0))
print(loaded_model.evaluate(x_test,y_test))
print(loaded_model.evaluate(x_train,y_train))
plt.plot(history2.history['loss'])
plt.plot(history2.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
plt.plot(history2.history['accuracy'])
plt.plot(history2.history['val_accuracy'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(X1, Y1, test_size=0.25, random_state=42)
y_test_pred=model.predict(x_test)
y_test_pred
y_test
y_test=y_test[:,0]
from numpy import savetxt
savetxt('gru_tanh_mse_y_test_pred.csv', y_test_pred[:1001], delimiter=',')
from numpy import savetxt
savetxt('gru_tanh_mse_y_test.csv', y_test[:1001], delimiter=',')
#completed
```
| github_jupyter |
```
import numpy as np
import os,sys
sys.path.append('../..')
sys.path.append('../../../RL_lib/Agents/PPO')
sys.path.append('../../../RL_lib/Utils')
%load_ext autoreload
%load_ext autoreload
%autoreload 2
%matplotlib nbagg
import os
print(os.getcwd())
%%html
<style>
.output_wrapper, .output {
height:auto !important;
max-height:1000px; /* your desired max-height here */
}
.output_scroll {
box-shadow:none !important;
webkit-box-shadow:none !important;
}
</style>
```
# Load Policy
```
from env_mdr import Env
from reward_terminal_mdr import Reward
import env_utils as envu
import attitude_utils as attu
from dynamics_model import Dynamics_model
from lander_model import Lander_model
from ic_gen import Landing_icgen
from agent_mdr2 import Agent
from policy_ppo import Policy
from value_function import Value_function
from utils import Mapminmax,Logger,Scaler
from flat_constraint import Flat_constraint
from glideslope_constraint import Glideslope_constraint
from attitude_constraint import Attitude_constraint
from thruster_model import Thruster_model
logger = Logger()
dynamics_model = Dynamics_model()
attitude_parameterization = attu.Quaternion_attitude()
thruster_model = Thruster_model()
thruster_model.max_thrust = 5000
thruster_model.min_thrust = 1000
lander_model = Lander_model(thruster_model, attitude_parameterization=attitude_parameterization,
apf_v0=70, apf_atarg=15., apf_tau2=100.)
lander_model.get_state_agent = lander_model.get_state_agent_tgo_alt
reward_object = Reward(tracking_bias=0.01,tracking_coeff=-0.01, fuel_coeff=-0.05, debug=False, landing_coeff=10.)
glideslope_constraint = Glideslope_constraint(gs_limit=-1.0)
shape_constraint = Flat_constraint()
attitude_constraint = Attitude_constraint(attitude_parameterization,
attitude_penalty=-100,attitude_coeff=-10,
attitude_limit=(10*np.pi, np.pi/2-np.pi/16, np.pi/2-np.pi/16))
env = Env(lander_model,dynamics_model,logger,
reward_object=reward_object,
glideslope_constraint=glideslope_constraint,
shape_constraint=shape_constraint,
attitude_constraint=attitude_constraint,
tf_limit=120.0,print_every=10)
env.ic_gen = Landing_icgen(mass_uncertainty=0.05,
g_uncertainty=(0.0,0.0),
attitude_parameterization=attitude_parameterization,
l_offset=0.,
adapt_apf_v0=True,
inertia_uncertainty_diag=100.0,
inertia_uncertainty_offdiag=10.0,
downrange = (0,2000 , -70, -10),
crossrange = (-1000,1000 , -30,30),
altitude = (2300,2400,-90,-70),
yaw = (-np.pi/8, np.pi/8, 0.0, 0.0) ,
pitch = (np.pi/4-np.pi/8, np.pi/4+np.pi/16, -0.01, 0.01),
roll = (-np.pi/8, np.pi/8, -0.01, 0.01))
env.ic_gen.show()
obs_dim = 12
act_dim = 4
policy = Policy(obs_dim,act_dim,kl_targ=0.001,epochs=20, beta=0.1, shuffle=True, servo_kl=True)
import utils
fname = "optimize_4km"
input_normalizer = utils.load_run(policy,fname)
print(input_normalizer)
```
# Test Policy over 4km
```
policy.test_mode=True
env.ic_gen = Landing_icgen(mass_uncertainty=0.05, g_uncertainty=(0.0,0.0),
adapt_apf_v0=True,
attitude_parameterization=attitude_parameterization,
downrange = (0,2000 , -70, -10),
crossrange = (-1000,1000 , -30,30),
altitude = (2300,2400,-90,-70),
yaw = (-np.pi/8, np.pi/8, 0.0, 0.0) ,
pitch = (np.pi/4-np.pi/8, np.pi/4+np.pi/16, -0.0, 0.0),
roll = (-np.pi/8, np.pi/8, -0.0, 0.0),
noise_u=100*np.ones(3), noise_sd=100*np.ones(3))
env.test_policy_batch(policy,input_normalizer,10000,print_every=100)
policy.test_mode=True
env.ic_gen = Landing_icgen(mass_uncertainty=0.0, g_uncertainty=(0.0,0.0),
adapt_apf_v0=True,
attitude_parameterization=attitude_parameterization,
downrange = (1500,1500 , -70, -70),
crossrange = (-500,500 , -30,-30),
altitude = (2400,2400,-90,-90),
yaw = (-np.pi/8, np.pi/8, 0.0, 0.0) ,
pitch = (np.pi/4-np.pi/8, np.pi/4+np.pi/16, -0.0, 0.0),
roll = (-np.pi/8, np.pi/8, -0.0, 0.0),
noise_u=0*np.ones(3), noise_sd=0*np.ones(3))
env.test_policy_batch(policy,input_normalizer,1,print_every=1)
envu.render_traj(lander_model.trajectory_list[0])
np.save("single_trajectory.npy",lander_model.trajectory_list[0])
policy.close_sess()
```
| github_jupyter |
# OpenCV Face Detection HDMI
In this notebook, opencv face detection will be applied to HDMI input images.
To run all cells in this notebook a HDMI input source and HDMI output monitor are required.
References:
https://github.com/Itseez/opencv/blob/master/data/haarcascades/haarcascade_frontalface_default.xml
https://github.com/Itseez/opencv/blob/master/data/haarcascades/haarcascade_eye.xml
### Step 1: Load the overlay
```
from pynq import Overlay
Overlay("base.bit").download()
```
### Step 2: Initialize HDMI I/O
```
from pynq.drivers import HDMI
from pynq.drivers.video import VMODE_1920x1080
hdmi_out = HDMI('out', video_mode=VMODE_1920x1080)
hdmi_in = HDMI('in', init_timeout=10, frame_list=hdmi_out.frame_list)
hdmi_in.start()
hdmi_out.start()
```
### Step 3:Show input frame using IPython Image
Source: http://www.gardnerproductions.ca/wp-content/uploads/2015/10/bigstock-People-with-different-emotions-102991475.jpg
```
from IPython.display import Image
frame = hdmi_in.frame()
orig_img_path = '/home/xilinx/jupyter_notebooks/examples/data/face_detect.jpg'
frame.save_as_jpeg(orig_img_path)
Image(filename=orig_img_path)
```
### Step 4: Apply the face detection to the input
```
import cv2
import numpy as np
frame = hdmi_in.frame_raw()
np_frame= (np.frombuffer(frame, dtype=np.uint8)).reshape(1080,1920,3)
face_cascade = cv2.CascadeClassifier(
'./data/haarcascade_frontalface_default.xml')
eye_cascade = cv2.CascadeClassifier(
'./data/haarcascade_eye.xml')
gray = cv2.cvtColor(np_frame, cv2.COLOR_BGR2GRAY)
faces = face_cascade.detectMultiScale(gray, 1.3, 5)
for (x,y,w,h) in faces:
cv2.rectangle(np_frame,(x,y),(x+w,y+h),(255,0,0),2)
roi_gray = gray[y:y+h, x:x+w]
roi_color = np_frame[y:y+h, x:x+w]
eyes = eye_cascade.detectMultiScale(roi_gray)
for (ex,ey,ew,eh) in eyes:
cv2.rectangle(roi_color,(ex,ey),(ex+ew,ey+eh),(0,255,0),2)
```
### Step 5: Disconnect HDMI out from HDMI in
The `hdmi_in` will now stream to different frame buffer (no longer connected to `hdmi_out`).
```
hdmi_in.frame_index_next()
```
### Step 6: Show results on HDMI output
Output OpenCV results via HDMI.
```
hdmi_out.frame_raw(bytearray(np_frame.tobytes()))
```
### Step 7: Show results within notebook
Output OpenCV results as JPEG.
```
orig_img_path = '/home/xilinx/jupyter_notebooks/examples/data/face_detect.jpg'
hdmi_out.frame().save_as_jpeg(orig_img_path)
Image(filename=orig_img_path)
```
### Step 7: Release HDMI
```
hdmi_out.stop()
hdmi_in.stop()
del hdmi_in, hdmi_out
```
| github_jupyter |
# Automating DFT Exercises
## Exercise 01: QChem Input/Outputs
Another important DFT code for MP is QChem. VASP uses a plane-wave basis, which makes it very efficient for periodic crystalline systems, but not very efficient for molecules. There are a number of DFT codes that use Gaussian functions to build the basis, such as Gaussian and QChem. Let's begin this example by loading the molecular structure of Ethylene Carbonate
Let's start by loading a `Molecule` object from pymatgen and importing the `ethylene_carbonate.xyz` as a `Molecule` object
```
from pymatgen import Molecule
mol = Molecule.from_file("ethylene_carbonate.xyz")
print(mol)
```
This is an XYZ file, which is a standard format for molecular structures. Several other formats are supported using the openbabel package that can be optionally installed.
For the purpose of this example, we've provided a completed QChem calculation under `QChem_ethylene_carboante`. Let's use pymatgen to read the inputs in this directory.
Use `tab` and `shift`+`tab` to explore the `pymatgen.io.qchem.inputs` module and find something that well let you read a QChem Input.
```
from pymatgen.io.qchem.inputs import QCInput
qcinp = QCInput.from_file("./QChem_etlyene_carbonate/mol.qin.gz")
print(qcinp.molecule)
```
For QChem, the input structure is much simpler as it is all contained in one file, this mol.qin file. The output comes directly from QChem as mostly a single file caled the QCOutput file. We have a corresponding object in pymatgen to read this file.
Let's do the same as above for outputs. Explore the `pymatgen.io.qchem.outputs` module and find something to read a QChem Output
```
from pymatgen.io.qchem.outputs import QCOutput
qcoutput = QCOutput(filename="./QChem_etlyene_carbonate/mol.qout.gz")
```
The data for this is all contained a single `data` attribute which is a dictionary with parsed information. Find the key that will get you the optimized output molecule geometry from the calculation.
```
qcoutput.data.keys()
qcoutput.data["molecule_from_optimized_geometry"]
```
Note that the optimized geoemtry has new coordinates that should be the minimum energy configuration for ethylene carbonate.
## Exercise 2: QChem Input Sets
We also have InputSets for QChem, which act very similarly to VASP. Because the input for QChem is much simpler, these sets just represent a single input file. Let's load the molecule again just incase.
```
from pymatgen import Molecule
mol = Molecule.from_file("ethylene_carbonate.xyz")
print(mol)
```
Explore the `pymatgen.io.qchem.sets` module and find an Input set to "Opt" or optimize the given molecule
```
from pymatgen.io.qchem.sets import OptSet
```
Now load up an input set and print what the QChem Input should look like
```
opt_set = OptSet(molecule=mol)
print(opt_set)
```
Now let's do the same to calculate the frequencies of a given Molecule
```
from pymatgen.io.qchem.sets import FreqSet
freq_set = FreqSet(mol)
print(freq_set)
```
Now inspect the parameters of the frequecny calculation input set using either `help` or `shift`+2x`tab`
```
help(freq_set.__init__)
```
The QChem InputSets just like the VASP InputSets are designed to be flexible for various DFT parameters such as the level of theory and the solvation environment.
Now try changing the DFT Rung and note what changed.
```
freq_set = FreqSet(mol,dft_rung=1)
print(freq_set)
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import parameters as pa
import matplotlib.pyplot as plt
import time
import math
from functools import reduce
from sklearn.preprocessing import MinMaxScaler
import tensorflow as tf
from tensorflow.keras import layers
tf.executing_eagerly()
import os
os.environ['TF_CPP_MIN_LOG_LEVEL'] = '2'
os.environ['TF_ROCM_FUSION_ENABLE'] = '1'
mpf_bct_raw = np.array(pd.read_csv("~/MPF/BCT/BCT.csv"))
daily_r, daily_c = mpf_bct_raw.shape
daily = np.array([s.replace("/", "-") for s in mpf_bct_raw[:,0]]).astype("datetime64")
mpf_bct = mpf_bct_raw[:,1:].astype("float64")
mpf_bct_list = mpf_bct.transpose().tolist()
lstm_size = 128
time_step = 60
batch_size = 512
output_size = 1
#input_size = 1
shift = 1
mpf_bct_returns = np.array([[math.log(col[i+1]/col[i]) if col[i] != 0 else -9999 for i,a in enumerate(col[:-1])] for col in mpf_bct_list], dtype='float32').transpose()
mpf_bct_returns[mpf_bct_returns==-9999.0] = np.nan
def plot_data(data):
plt.figure(figsize=(15,6))
plt.plot(data)
class Attention(tf.keras.Model):
def __init__(self, rnn_units, batch_size, time_step, input_size, output_size):
super(Attention, self).__init__()
self.units = rnn_units
self.batch_size = batch_size
self.time_step = time_step
self.input_step = input_size
self.output_size = output_size
self.lstm = tf.keras.layers.LSTM(self.units,
return_sequences = True,
return_state = True,
stateful = True,
recurrent_initializer='glorot_uniform')
self.Wh = tf.keras.layers.Dense(self.units)
self.Ws = tf.keras.layers.Dense(self.units)
self.Wx = tf.keras.layers.Dense(1)
self.V = tf.keras.layers.Dense(1)
self.O = tf.keras.layers.Dense(self.output_size)
def call(self, x, hidden, state):
# hidden shape (batch, units) to (batch,1,units)
hidden = tf.expand_dims(hidden, 1)
# x shape (batch, time_step, 1)
x = tf.expand_dims(x,0)
score = self.V(tf.nn.tanh(self.Wx(x) + self.Wh(hidden) + self.Ws(state)))
# attention shape = (batch, time_step, 1)
attention_weights = tf.nn.softmax(score, axis = 1)
encoder_outputs, state_h, state_c = self.lstm(x*attention_weights)
output = self.O(state_h)
return attention_weights,state_h, state_c, output
def init_hidden_state(self):
hidden = tf.zeros((self.batch_size, self.units))
state = tf.zeros((self.batch_size, self.time_step, self.units))
return hidden, state
mpf_bct_returns[9,:]
```
| github_jupyter |
Persistent homology examples
* Ripser [paper](https://www.theoj.org/joss-papers/joss.00925/10.21105.joss.00925.pdf) [code](https://github.com/scikit-tda/ripser.py) (fast)
* Dionysus 2 [code](https://mrzv.org/software/dionysus2/) (representative examples)
* Nico's [code](https://github.com/nhchristianson/Math-text-semantic-networks)
* Ann's [code](https://github.com/asizemore/PH_tutorial/blob/master/Tutorial_day1.ipynb)
# Load data
## Load networks
```
%reload_ext autoreload
%autoreload 2
import os,sys
sys.path.insert(1, os.path.join(sys.path[0], '..', 'module'))
topics = [
'anatomy', 'biochemistry', 'cognitive science', 'evolutionary biology',
'genetics', 'immunology', 'molecular biology', 'chemistry', 'biophysics',
'energy', 'optics', 'earth science', 'geology', 'meteorology',
'philosophy of language', 'philosophy of law', 'philosophy of mind',
'philosophy of science', 'economics', 'accounting', 'education',
'linguistics', 'law', 'psychology', 'sociology', 'electronics',
'software engineering', 'robotics',
'calculus', 'geometry', 'abstract algebra',
'Boolean algebra', 'commutative algebra', 'group theory', 'linear algebra',
'number theory', 'dynamical systems and differential equations'
]
import wiki
path_saved = '/Users/harangju/Developer/data/wiki/graphs/dated/'
networks = {}
for topic in topics:
print(topic, end=' ')
networks[topic] = wiki.Net(path_graph=path_saved + topic + '.pickle',
path_barcodes=path_saved + topic + '.barcode')
path_null = '/Users/harangju/Developer/data/wiki/graphs/null-target/'
num_nulls = 10
null_targets = {}
for topic in topics:
print(topic, end=' ')
null_targets[topic] = [None for i in range(num_nulls)]
for i in range(num_nulls):
null_targets[topic][i] = wiki.Net(path_graph=path_null + topic + '-null-' + str(i) + '.pickle',
path_barcodes=path_null + topic + '-null-' + str(i) + '.barcode')
path_null = '/Users/harangju/Developer/data/wiki/graphs/null-year/'
num_nulls = 10
null_years = {}
for topic in topics:
print(topic, end=' ')
null_years[topic] = [None for i in range(num_nulls)]
for i in range(num_nulls):
null_years[topic][i] = wiki.Net(path_graph=path_null + topic + '-null-' + str(i) + '.pickle',
path_barcodes=path_null + topic + '-null-' + str(i) + '.barcode')
```
## Load models
```
simulation = '20200422_1318'
simulation = '20200520_2057'
simulation = '20200820_1919'
base_dir = os.path.join('/', 'Users', 'harangju', 'Developer', 'data', 'wiki', 'simulations')
session_dir = os.path.join(base_dir, simulation)
filenames = sorted(os.listdir(session_dir))
filenames[:3]
filenames[-3:]
model_topics = list(set(
[filename.split('_')[1] for filename in filenames
if filename.split('_')[0]=='model']
))
model_topics[:3]
model_paths = {topic: [os.path.join(session_dir, filename)
for filename in filenames[:-1]
if (filename.split('_')[0]=='model') and (filename.split('_')[1]==topic)]
for topic in model_topics}
{topic: model_paths[topic] for topic in model_topics[:1]}
```
# Compute barcodes
Go to the "Load" sections to load `barcodes` with computed data.
```
import pandas as pd
pd.options.display.max_rows = 12
import dill
barcodes = pd.concat(
[
network.barcodes.assign(topic=topic)\
.assign(type='real')\
.assign(null=0)
for topic, network in networks.items()
] +
[
network.barcodes.assign(topic=topic)\
.assign(type='null_targets')\
.assign(null=i)
for topic, nulls in null_targets.items()
for i, network in enumerate(nulls)
] +
[
network.barcodes.assign(topic=topic)\
.assign(type='null_years')\
.assign(null=i)
for topic, nulls in null_years.items()
for i, network in enumerate(nulls)
] +
[
dill.load(open(path, 'rb'))\
.barcodes\
.assign(topic=topic)\
.assign(type='null_genetic')\
.assign(null=i)
for topic, paths in model_paths.items()
for i, path in enumerate(paths)
],
ignore_index=True, sort=False)
barcodes = barcodes[barcodes.lifetime!=0]
barcodes
```
## Save
```
import pickle
path_analysis = '/Users/harangju/Developer/data/wiki/analysis/'
pickle.dump(
barcodes,
open(
os.path.join(path_analysis, f"barcodes_{simulation}.pickle"),
'wb'
)
)
```
## Load
```
import pickle
path_analysis = '/Users/harangju/Developer/data/wiki/analysis/'
barcodes = pickle.load(
open(
os.path.join(path_analysis, f"barcodes_{simulation}.pickle"),
'rb'
)
)
```
# Plotting functions
```
import os
import numpy as np
import plotly.express as px
import plotly.graph_objects as go
import pandas as pd
# path_fig = '/Users/harangju/Box Sync/Research/my papers/wikipedia/results/'
path_fig = os.path.join(
'/', 'Users', 'harangju', 'Library', 'Mobile Documents', 'com~apple~CloudDocs',
'Documents', 'research', 'wikipedia', 'results'
)
import numpy as np
import pandas as pd
import plotly.offline as po
import plotly.express as px
import plotly.graph_objs as go
import plotly.figure_factory as ff
po.init_notebook_mode(connected=True)
def plot_barcodes(barcodes, title):
fig = go.Figure()
x = {dim: [] for dim in pd.unique(barcodes.dim)}
y = {dim: [] for dim in pd.unique(barcodes.dim)}
deaths = [[],[]]
for i, row in barcodes.iterrows():
dim = row['dim']
birth = row['birth']
death = row['death'] if row['death']!=np.inf else 2050
x[dim].extend([birth,death,None])
y[dim].extend([i,i,None])
if row['death']!=np.inf:
deaths[0].extend([death,None])
deaths[1].extend([i,None])
for dim in pd.unique(barcodes.dim):
fig.add_trace(go.Scatter(x=x[dim], y=y[dim],
mode='lines',
name=f"dim={dim}"))
# fig.add_trace(go.Scatter(x=deaths[0], y=deaths[1], mode='markers',
# marker={'color': 'black', 'size': 1},
# name='deaths'))
fig.update_layout(template='plotly_white',
width=600, height=500,
title_text=f"{title}",
xaxis={'title': 'year', 'range': [0, 2040]},
yaxis={'title': '', 'tickvals': []})
fig.show()
return fig
def plot_persistence_diagram(barcodes):
colors = [mcd.XKCD_COLORS['xkcd:'+c]
for c in ['emerald green', 'tealish', 'peacock blue',
'grey', 'brown', 'red', 'yellow']]
plt.figure(figsize=(10,10))
for dim in set(barcodes['dim']):
data = barcodes.loc[barcodes['dim']==dim]
data.loc[data['death']==np.inf,'death'] = 2030
plt.plot(data['birth'], data['death'], '.')
x = [barcodes['birth'].min(),
barcodes.loc[barcodes['death']!=np.inf,'death'].max()]
print(x)
plt.plot(x, [2030, 2030], '--')
def plot_betti(barcodes, title):
fig = go.Figure()
year_min = int(np.min(barcodes.birth))
year_max = int(np.max(barcodes[barcodes.death!=np.inf].death))
counts = np.zeros((len(barcodes.index), year_max-year_min))
dims = np.zeros(len(barcodes.index))
for i, row in barcodes.iterrows():
dim = row['dim']
birth = int(row['birth'])
death = int(row['death']) if row['death']!=np.inf else year_max
counts[i,birth-year_min:death-year_min] = 1
dims[i] = dim
for dim in pd.unique(barcodes.dim):
betti = np.sum(counts[dims==dim,:], axis=0)
fig.add_trace(go.Scatter(x=np.arange(year_min, year_max) - year_min + 1,
y=betti,
mode='lines',
name=f"dim={dim}"))
fig.update_layout(template='plotly_white',
title_text=f"{title}",
xaxis={'title': 'year',
# 'range': [], #[0,year_max],
'type': 'linear'},
yaxis={'title': 'count',
# 'range': [0,2000],
'type': 'log'})
fig.show()
return fig
```
# Barcodes
```
import os
path_plot = '2 barcodes'
if not os.path.exists(os.path.join(path_fig, path_plot)):
os.mkdir(os.path.join(path_fig, path_plot))
for topic in ['biophysics']:#topics:
plot_barcodes(
networks[topic].barcodes[networks[topic].barcodes.lifetime!=0]\
.reset_index().drop('index', axis=1),
f"Topic: {topic} (empirical)")\
.write_image(os.path.join(path_fig, path_plot, f"{topic}_empirical.pdf"))
# plot_barcodes(null_targets[topic][0].barcodes[null_targets[topic][0].barcodes.lifetime!=0],
# f"Topic: {topic} (target-rewired)")#\
# .write_image(os.path.join(path_fig, path_plot, f"{topic}_target.pdf"))
# plot_barcodes(null_years[topic][0].barcodes[null_years[topic][0].barcodes.lifetime!=0],
# f"Topic: {topic} (year-reordered)")#\
# .write_image(os.path.join(path_fig, path_plot, f"{topic}_year.pdf"))
```
# Betti curves
```
import os
path_plot = '2 betti'
if not os.path.exists(os.path.join(path_fig, path_plot)):
os.mkdir(os.path.join(path_fig, path_plot))
for topic in ['biochemistry']:#topics:
b = networks[topic].barcodes
plot_betti(b[b.lifetime!=0].reset_index(), f"{topic} (empirical)")\
.write_image(os.path.join(path_fig, path_plot, f"{topic}_empirical.pdf"))
b = null_targets[topic][0].barcodes
plot_betti(b[b.lifetime!=0].reset_index(), f"{topic} (target-rewired)")\
# .write_image(os.path.join(path_fig, path_plot, f"{topic}_target.pdf"))
b = null_years[topic][0].barcodes
plot_betti(b[b.lifetime!=0].reset_index(), f"{topic} (year-reordered)")\
# .write_image(os.path.join(path_fig, path_plot, f"{topic}_year.pdf"))
```
# Lifetime (finite)
```
from scipy import stats
lifetime = pd.DataFrame()
for topic in topics:
data = barcodes[barcodes.topic==topic].copy()
data = data[(data.lifetime!=np.inf) & (data.lifetime!=0)]
t_targets, p_targets = stats.ttest_ind(
data[data.type=='real']['lifetime'].values,
data[data.type=='null_targets']['lifetime'].values
)
t_years, p_years = stats.ttest_ind(
data[data.type=='real']['lifetime'].values,
data[data.type=='null_years']['lifetime'].values
)
t_genetic, p_genetic = stats.ttest_ind(
data[data.type=='real']['lifetime'].values,
data[data.type=='null_genetic']['lifetime'].values
)
lifetime = pd.concat(
[lifetime, pd.DataFrame(
[[topic, t_targets, p_targets, t_years, p_years, t_genetic, p_genetic]],
columns=[
'topic', 't (targets)', 'p (targets)',
't (years)', 'p (years)',
't (genetic)', 'p (genetic)'
]
)], ignore_index=True
)
pd.options.display.max_rows = 37
lifetime
barcodes_mean = barcodes[
(barcodes.lifetime!=np.inf) & (barcodes.lifetime!=0)]\
.groupby(['topic', 'type'], as_index=False)\
.mean()\
.drop(['dim','birth','death','null'], axis=1)
barcodes_mean
import os
path_plot = '2 lifetimes'
if not os.path.exists(os.path.join(path_fig, path_plot)):
os.mkdir(os.path.join(path_fig, path_plot))
null_types = [
'null_targets', 'null_genetic'
]
fig = go.Figure()
max_lifetime = np.max(barcodes_mean.lifetime) + 10
fig.add_trace(
go.Scatter(
x=[0,max_lifetime],
y=[0,max_lifetime],
mode='lines',
line=dict(dash='dash'),
name='1:1'
)
)
for null_type in null_types:
fig.add_trace(
go.Scatter(
x=barcodes_mean[barcodes_mean.type==null_type].lifetime,
y=barcodes_mean[barcodes_mean.type=='real'].lifetime,
mode='markers',
name=null_type,
hovertext=barcodes_mean[barcodes_mean.type=='real'].topic
)
)
fig.update_layout(template='plotly_white',
title='Lifetimes (finite)',
width=500, height=500,
xaxis={'title': 'years (null)',
'range': [0,max_lifetime+100],
'dtick': 1000},
yaxis={'title': 'years (real)',
'range': [0,max_lifetime+100],
'scaleanchor': 'x',
'scaleratio': 1,
'dtick': 1000})
fig.show()
fig.write_image(os.path.join(path_fig, path_plot, 'finite.pdf'))
import scipy as sp
for null_type in ['null_targets', 'null_genetic']:
ks, p_ks = sp.stats.ks_2samp(
barcodes[
(barcodes.lifetime!=0) & (barcodes.lifetime!=np.inf) &
(barcodes.type=='real')].lifetime,
barcodes[
(barcodes.lifetime!=0) & (barcodes.lifetime!=np.inf) &
(barcodes.type==null_type)].lifetime,
alternative='two-sided'
)
t, p_t = sp.stats.ttest_ind(
barcodes[
(barcodes.lifetime!=0) & (barcodes.lifetime!=np.inf) &
(barcodes.type=='real')].lifetime,
barcodes[
(barcodes.lifetime!=0) & (barcodes.lifetime!=np.inf) &
(barcodes.type==null_type)].lifetime,
equal_var=True
)
print(null_type, f"ks={ks}, p={p_ks}; t={t}, p={p_t}")
fig = px.violin(
barcodes[(barcodes.lifetime!=0) & (barcodes.type!='null_years')],
x='type', y='lifetime'
)
fig.update_layout(
height=400, width=460,
template='plotly_white',
title_text='Lifetimes (finite)',
xaxis={'title': ''},
yaxis={
'title': 'number',
'type': '-',
}
)
fig.show()
fig.write_image(os.path.join(path_fig, path_plot, 'finite_violin.pdf'))
lifetimes = barcodes[
(barcodes.lifetime!=0) &
(barcodes.lifetime!=np.inf)
]
null_types = [
'real', 'null_targets', 'null_genetic'
]
lifetime_range = np.arange(
np.min(lifetimes.lifetime),
np.max(lifetimes.lifetime)
)
cum_freq = {
null_type: np.zeros(lifetime_range.size)
for null_type in null_types
}
for null_type in null_types:
print(null_type)
for i, lifetime in enumerate(lifetime_range):
cum_freq[null_type][i] = np.sum(
lifetimes[lifetimes.type==null_type].lifetime < lifetime
) / len(lifetimes[lifetimes.type==null_type].index)
fig = go.Figure()
for null_type in null_types:
fig.add_trace(
go.Scatter(
x=lifetime_range,
y=cum_freq[null_type],
name=null_type
)
)
fig.update_layout(
width=400, height=400,
template='plotly_white',
yaxis={'title': 'cumulative frequency'},
xaxis={'title': 'finite lifetime',
'type': 'log'},
legend={'x': 0, 'y':1}
)
fig.show()
fig.write_image(os.path.join(path_fig, path_plot, 'cum_freq_fin.pdf'))
```
# Lifetime (infinite)
```
barcodes[
(barcodes.lifetime==np.inf) &
(barcodes.topic=='biochemistry') &
(barcodes.type=='real')
].shape
import scipy as sp
reals = []
targets = []
years = []
genetics = []
for topic in topics:
reals.append(barcodes[(barcodes.lifetime==np.inf) &
(barcodes.topic==topic) &
(barcodes.type=='real')].shape[0])
targets.append(barcodes[(barcodes.lifetime==np.inf) &
(barcodes.topic==topic) &
(barcodes.type=='null_targets')].shape[0])
years.append(barcodes[(barcodes.lifetime==np.inf) &
(barcodes.topic==topic) &
(barcodes.type=='null_years')].shape[0])
genetics.append(barcodes[(barcodes.lifetime==np.inf) &
(barcodes.topic==topic) &
(barcodes.type=='null_genetic')].shape[0])
t_targets, p_targets = sp.stats.ttest_ind(reals, targets)
t_years, p_years = sp.stats.ttest_ind(reals, years)
t_genetic, p_genetic = sp.stats.ttest_ind(reals, genetics)
t_targets, p_targets, t_years, p_years, t_genetic, p_genetic
ks_targets, p_targets = sp.stats.ks_2samp(reals, targets, alternative='two-sided')
ks_years, p_years = sp.stats.ks_2samp(reals, years, alternative='two-sided')
ks_genetic, p_genetic = sp.stats.ks_2samp(reals, genetics, alternative='two-sided')
ks_targets, p_targets, ks_years, p_years, ks_genetic, p_genetic
import plotly.figure_factory as ff
import os
path_plot = '2 lifetimes'
if not os.path.exists(os.path.join(path_fig, path_plot)):
os.mkdir(os.path.join(path_fig, path_plot))
fig = ff.create_distplot(
[targets, genetics, reals],
['null targets', 'null genetics', 'real'],
bin_size=np.arange(.5, 8) ** 10, #1000,
show_curve=False,
colors=['#2ca02c', '#d62728', '#1f77b4']
)
fig.update_layout(
width=600,
template='plotly_white',
title_text='Lifetimes (infinite)',
xaxis={'title': 'count', 'type': 'log'},
yaxis={'title': 'probability'}
)
fig.show()
fig.write_image(os.path.join(path_fig, path_plot, 'infinite.pdf'))
fig = px.violin(
pd.DataFrame({
'value': reals + targets + genetics,
'type': len(reals)*['real'] + len(targets)*['target']\
+ len(genetics)*['genetic']
}).replace(0, 0.01),
x='type', y='value'
)
fig.update_layout(
height=400, width=460,
template='plotly_white',
title_text='Lifetimes (infinite)',
xaxis={'title': ''},
yaxis={
'title': 'number',
'type': '-',
# 'range': [-1, 6]
}
)
fig.show()
fig.write_image(os.path.join(path_fig, path_plot, 'infinite_violin.pdf'))
inf_lifetimes = {
'real': reals,
'null_targets': targets,
'null_genetic': genetics
}
null_types = [
'real', 'null_targets', 'null_genetic'
]
inf_lifetime_range = np.arange(
0, np.max(reals + targets + genetics)
)
cum_freq = {
null_type: []
for null_type in null_types
}
for null_type in null_types:
print(null_type)
for i, count in enumerate(inf_lifetime_range):
cum_freq[null_type].append(
np.sum(inf_lifetimes[null_type] < count) \
/ len(inf_lifetimes[null_type])
)
fig = go.Figure()
for null_type in null_types:
fig.add_trace(
go.Scatter(
x=inf_lifetime_range,
y=cum_freq[null_type],
name=null_type
)
)
fig.update_layout(
width=400, height=400,
template='plotly_white',
yaxis={'title': 'cumulative frequency'},
xaxis={'title': 'infinite lifetimes',
'type': 'log'},
legend={'x':.5, 'y':.2}
)
fig.show()
fig.write_image(os.path.join(path_fig, path_plot, 'cum_freq_inf.pdf'))
```
# Dimensionality
## Compute
```
pd.options.display.max_rows = 5
counts = barcodes[barcodes.lifetime!=0]\
.assign(count=1)\
.groupby(['type','topic','dim'], as_index=False)['count']\
.sum()\
.sort_values('type', axis=0, ascending=True)
counts
nulls = barcodes[barcodes.lifetime!=0]\
.groupby(['type','topic','dim'], as_index=False)['null'].max()
nulls.null = nulls.null + 1
nulls
nulls = pd.merge(
nulls, counts,
how='left', left_on=['type','topic','dim'],
right_on=['type','topic','dim']
).replace(
['null_targets','null_years', 'null_genetic'],
['targets', 'years', 'genetics']
).sort_values(by='type')
nulls['count'] = nulls['count'] / nulls.null
nulls
```
## Statistics
```
import scipy as sp
dim_stat = pd.DataFrame()
for dim in sorted(pd.unique(nulls.dim)):
nulls_dim = nulls[nulls.dim==dim]
nulls_dim_count_real = nulls_dim[nulls_dim.type=='real']['count'].values
nulls_dim_count_years = nulls_dim[nulls_dim.type=='years']['count'].values
nulls_dim_count_targets = nulls_dim[nulls_dim.type=='targets']['count'].values
nulls_dim_count_genetics = nulls_dim[nulls_dim.type=='genetics']['count'].values
t_years, p_years = sp.stats.ttest_ind(
nulls_dim_count_real, nulls_dim_count_years
)
t_targets, p_targets = sp.stats.ttest_ind(
nulls_dim_count_real, nulls_dim_count_targets
)
t_genetics, p_genetics = sp.stats.ttest_ind(
nulls_dim_count_real, nulls_dim_count_genetics
)
ks_years, ks_p_years = sp.stats.ks_2samp(
nulls_dim_count_real, nulls_dim_count_years, alternative='two-sided'
) if len(nulls_dim_count_years) > 0 else (-1, -1)
ks_targets, ks_p_targets = sp.stats.ks_2samp(
nulls_dim_count_real, nulls_dim_count_targets, alternative='two-sided'
) if len(nulls_dim_count_targets) > 0 else (-1, -1)
ks_genetics, ks_p_genetics = sp.stats.ks_2samp(
nulls_dim_count_real, nulls_dim_count_genetics, alternative='two-sided'
) if len(nulls_dim_count_genetics) > 0 else (-1, -1)
dim_stat = pd.concat(
[
dim_stat,
pd.DataFrame(
[[
dim,
t_years, p_years, t_targets, p_targets, t_genetics, p_genetics,
ks_years, ks_p_years, ks_targets, ks_p_targets, ks_genetics, ks_p_genetics
]],
columns=[
'dim',
't_years', 'p_years', 't_targets', 'p_targets', 't_genetics', 'p_genetics',
'ks_years', 'ks_p_years', 'ks_targets', 'ks_p_targets',
'ks_genetics', 'ks_p_genetics'
]
)
]
)
dim_stat
```
## Plot
```
import os
path_plot = '2 dimensionality'
if not os.path.exists(os.path.join(path_fig, path_plot)):
os.mkdir(os.path.join(path_fig, path_plot))
fig = px.box(
nulls[
(nulls['type']=='real') &
(nulls['type']=='targets') &
(nulls['type']=='genetics')
],
x='dim', y='count', color='type'
)
fig.update_layout(
template='plotly_white',
title_text='Dimensionality',
yaxis={'type': 'log'}
)
fig.update_traces(marker={'size': 4})
fig.show()
fig.write_image(os.path.join(path_fig, path_plot, 'dimensionality.pdf'))
```
## Mean dimensionality
```
barcodes
counts = barcodes[barcodes.lifetime!=0]\
.assign(count=1)\
.groupby(['type', 'topic', 'dim'], as_index=False)['count']\
.sum()
counts
barcodes = barcodes[barcodes.lifetime!=0]
for null_type in ['null_targets', 'null_genetic']:
ks, p_ks = sp.stats.ks_2samp(
barcodes[barcodes.type=='real'].dim,
barcodes[barcodes.type==null_type].dim,
alternative='two-sided'
)
t, p_t = sp.stats.ttest_ind(
barcodes[barcodes.type=='real'].dim,
barcodes[barcodes.type==null_type].dim,
equal_var=True
)
print(null_type, f"ks={ks}, p={p_ks}, t={t}, p={p_t}")
fig = px.violin(
barcodes[barcodes.type!='null_years'], x='type', y='dim'
)
fig.update_layout(
height=400, width=460,
template='plotly_white',
title_text='Dimensionality'
)
fig.show()
fig.write_image(os.path.join(path_fig, path_plot, 'dimensionality_violin.pdf'))
dims = barcodes[(barcodes.lifetime!=0)]
null_types = ['real', 'null_targets', 'null_genetic']
dim_range = np.unique(dims.dim)
cum_freq = {
null_type: []
for null_type in null_types
}
for null_type in null_types:
print(null_type)
for i, dim in enumerate(dim_range):
cum_freq[null_type].append(
np.sum(
dims[dims.type==null_type].dim < dim
) / len(dims[dims.type==null_type].index)
)
fig = go.Figure()
for null_type in null_types:
fig.add_trace(
go.Scatter(
x=dim_range,
y=cum_freq[null_type],
name=null_type,
mode='lines'
)
)
fig.update_layout(
width=400, height=400,
template='plotly_white',
yaxis={'title': 'cumulative frequency'},
xaxis={'title': 'dim'},
legend={'x':.5, 'y':.2}
)
fig.show()
fig.write_image(os.path.join(path_fig, path_plot, 'cum_freq_dim.pdf'))
```
# Lifetime vs Cavity volume
## Compute cavity volume
Useful resource
* [Computational topology](https://books.google.com/books?id=MDXa6gFRZuIC&printsec=frontcover#v=onepage&q=%22persistent%20homology%22&f=true)
* [tutorial](http://pages.cs.wisc.edu/~jerryzhu/pub/cvrghomology.pdf)
```
import pickle
import numpy as np
import gensim.utils as gu
import gensim.matutils as gmat
import sklearn.metrics.pairwise as smp
for i, row in barcodes.iterrows():
sys.stdout.write("\rindex: " + str(i+1) + '/' + str(len(barcodes.index)))
sys.stdout.flush()
nodes = row['homology nodes']
topic = row['topic']
null_index = int(row['null'])
if row['type'] == 'real':
network = networks[topic]
elif row['type'] == 'null_targets':
network = null_targets[topic][null_index]
else:
network = null_years[topic][null_index]
tfidf = network.graph.graph['tfidf']
indices = [network.nodes.index(n) for n in nodes]
centroid = tfidf[:,indices].mean(axis=1) if indices else 0
distances = smp.cosine_distances(X=tfidf[:,indices].transpose(), Y=centroid.transpose())\
if indices else [0]
barcodes.loc[i,'average distance'] = np.mean(distances)
barcodes
```
## Save
```
import pickle
path_analysis = '/Users/harangju/Developer/data/wiki/analysis/'
pickle.dump(barcodes, open(f"{path_analysis}/barcode_volume.pickle",'wb'))
```
## Load
```
import pickle
path_analysis = '/Users/harangju/Developer/data/wiki/analysis/'
barcodes = pickle.load(open(f"{path_analysis}/barcode_volume.pickle",'rb'))
barcodes
```
## Compute regression
```
reg = pd.DataFrame()
for topic in pd.unique(barcodes.topic):
data = barcodes[(barcodes.topic==topic) & (barcodes.type=='real') &
(barcodes.lifetime!=np.inf) & (barcodes.lifetime!=0)]
x = data['average distance'].values
y = data['lifetime'].values
r, p = sp.stats.pearsonr(x, y)
reg = pd.concat([reg, pd.DataFrame([[topic,r,p,data.shape[0]]],
columns=['topic','r','p','n'])],
ignore_index=True)
reg[reg.n>200]
reg[reg.p<0.01]
```
## Plot
```
import scipy as sp
import scipy.stats
path_plot = '2 cavity volume'
if not os.path.exists(os.path.join(path_fig, path_plot)):
os.mkdir(os.path.join(path_fig, path_plot))
for topic in pd.unique(barcodes.topic):
data = barcodes[(barcodes.topic==topic) & (barcodes.type=='real') &
(barcodes.lifetime!=np.inf) & (barcodes.lifetime!=0)]
x = data['average distance'].values
y = data['lifetime'].values
r, p = sp.stats.pearsonr(x, y)
a, b, _, _, _ = sp.stats.linregress(x, y)
fig = go.Figure()
fig.add_trace(go.Scatter(x=x, y=y,
mode='markers',
marker={'size': 4},
showlegend=False))
fig.add_trace(go.Scatter(x=np.linspace(np.min(x),np.max(x)),
y=a*np.linspace(np.min(x),np.max(x))+b,
mode='lines',
showlegend=False))
fig.update_layout(template='plotly_white',
title_text=f"{topic} (r={r:.2f}, p={p:0.1e})",
xaxis={'title': 'average distance to centroid'},
yaxis={'title': 'lifetime'})
fig.show()
fig.write_image(os.path.join(path_fig, path_plot, f"{topic}.pdf"))
```
# Lifetime vs Cavity weights
## Compute
```
import sys
for i, row in barcodes.iterrows():
sys.stdout.write("\rindex: " + str(i+1) + '/' + str(len(barcodes.index)))
sys.stdout.flush()
nodes = row['homology nodes']
topic = row['topic']
null_index = int(row['null'])
if row['type'] == 'real':
network = networks[topic]
elif row['type'] == 'null_targets':
network = null_targets[topic][null_index]
else:
network = null_years[topic][null_index]
subgraph = network.graph.subgraph(nodes)
barcodes.loc[i,'mean edge weights'] = np.mean([subgraph.edges[u,v]['weight']
for u,v in subgraph.edges])
barcodes
```
## Save
```
import pickle
path_analysis = '/Users/harangju/Developer/data/wiki/analysis/'
pickle.dump(barcodes, open(f"{path_analysis}/barcode_volume_weights.pickle",'wb'))
```
## Load
```
import pickle
path_analysis = '/Users/harangju/Developer/data/wiki/analysis/'
barcodes = pickle.load(open(f"{path_analysis}/barcode_volume_weights.pickle",'rb'))
```
## Compute regression
```
import scipy as sp
import scipy.stats
for topic in ['anatomy']:
data = barcodes[(barcodes.topic==topic) & (barcodes.type=='real') &
(barcodes.lifetime!=np.inf) & (barcodes.lifetime!=0)].dropna()
x = data['mean edge weights'].values
y = data['lifetime'].values
r, p = sp.stats.pearsonr(x, y) if len(data['lifetime'])>2 else (0,0)
```
## Plot
```
path_plot = '2 cavity weights'
if not os.path.exists(os.path.join(path_fig, path_plot)):
os.mkdir(os.path.join(path_fig, path_plot))
import scipy as sp
import scipy.stats
for topic in pd.unique(barcodes.topic):
data = barcodes[(barcodes.topic==topic) & (barcodes.type=='real') &
(barcodes.lifetime!=np.inf) & (barcodes.lifetime!=0)].dropna()
x = data['mean edge weights'].values
y = data['lifetime'].values
r, p = sp.stats.pearsonr(x, y)
a, b, _, _, _ = sp.stats.linregress(x, y)
fig = go.Figure()
fig.add_trace(go.Scatter(x=x, y=y,
mode='markers',
marker={'size': 4},
showlegend=False))
fig.add_trace(go.Scatter(x=np.linspace(np.min(x),np.max(x)),
y=a*np.linspace(np.min(x),np.max(x))+b,
mode='lines',
showlegend=False))
fig.update_layout(template='plotly_white',
title_text=f"{topic} (r={r:.2f}, p={p:0.1e})",
xaxis={'title': 'average weights'},
yaxis={'title': 'lifetime'})
fig.show()
fig.write_image(os.path.join(path_fig, path_plot, f"{topic}.pdf"))
```
| github_jupyter |
# 再说几句Python
```
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
```
## 前情提要
上两集中,我们学习了python的基本语法(数据类型、循环、字符串的处理、循环语句、判断语句、列表、怎么建立函数)和numpy的基本功能(array的概念、加减乘除、array的shape、如何得到array的元素、建立mask array等等)。请先回顾之前的内容。
这次补充几个关于python语言的高级用法,为之后讲爬虫和pandas做一点铺垫。
### `map` and `lambda`
仅在定义匿名函数的地方使用这个函数,其他地方用不到,所以就不需要给它取个阿猫阿狗之类的名字了。
```
[i / 256 for i in [55, 125, 34]]
def func(x):
return x + 5
func(4)
map(func, [4, 5, 6, 7])
list(map(func, [4, 5, 6, 7]))
list(map(lambda x: x + 5, [4]))
list(map(lambda x: x + 5, [4, 5, 6, 7]))
provinceName = ["湖北", "浙江", "江西", "安徽"]
def append_sheng(x):
return x + '省'
list(map(append_sheng, provinceName))
provinceFullName = list(map(lambda x: x + '省', provinceName))
provinceFullName
provinceName = ["湖北", "浙江", "江西省", "安徽"]
provinceFullName = list(map(lambda x: x + '省', provinceName))
provinceFullName
provinceFullName = list(map(lambda x: x.rstrip('省') + '省', provinceName))
provinceFullName
```
### `enumerate` and `zip`
```
infection = [27100, 1075, 740, 779]
for prov in provinceFullName:
if prov == '浙江省':
# 浙江有多少人感染???
for i, prov in enumerate(provinceFullName):
if prov == '浙江省':
# 浙江有多少人感染???
print('Infections = {}'.format(infection[i]))
```
用两个数组 provinceFullName 和 infection 来表达疫情,没有很好的结构性,不好一一对应。
```
list(zip(provinceFullName, infection))
```
练习1: print "{city} is in {state}" for each combination.
```
cities = ["Phoenix", "Austin", "San Diego", "New York"]
states = ["Arizona", "Texas", "California", "New York"]
couple = list(zip(cities, states))
for i, city in enumerate(couple):
print(i, city)
couple
['{0} is in {1}'.format(item[0], item[1]) for item in couple]
```
练习2: 湖北每日新增多少确诊病例?
```
hubei_infection = [270, 444, 444, 729, 761, 1423, 1423, 3554, 4586,
5806, 5806, 9074, 9074, 13522, 16678, 19665, 22112, 24953,
27100, 27100] # from Jan 21
len(hubei_infection)
[hubei_infection[i + 1] - num for i, num in enumerate(hubei_infection[:-1])]
```
### 词典 `dict` and `set`
```
NCP_infection = dict(list(zip(provinceFullName, infection)))
NCP_infection
NCP_infection['湖北省']
NCP_infection['安徽省']
NCP_infection['四川省']
NCP_infection.keys()
NCP_infection['北京市'] = 326
NCP_infection
list(NCP_infection.values())
list(NCP_infection.keys())
province_name = ['湖北省', '浙江省', '江西省', '安徽省', '北京市', '北京市', '安徽省']
set(province_name)
list(set(province_name))
np.unique(np.array(province_name), return_counts=True, return_index=True)
```
### `datetime` 每时每刻
```
import datetime
now = datetime.datetime.now()
now
now = now.replace(second=0, microsecond=0)
now
pd.to_datetime(now)
```
怎么把 '2019年10月11日 23:52' 转化机器能看懂的东西?
```
crush_time_str = '2019年10月11日 23:52'
crush_time = datetime.datetime.strptime(crush_time_str, '%Y年%m月%d日 %H:%M')
crush_time
pd.to_datetime(crush_time)
pd.to_datetime(crush_time + datetime.timedelta(days=1))
pd.to_datetime(crush_time + datetime.timedelta(days=365))
```
### 给函数写好注释,做一个负责任的女人
行号
```
def parse_time(time_str, fmt='%Y年%m月%d日 %H:%M', verbose=True): # why not using `format`?
'''
Parse the time given as a string.
Parameters:
time_str (str): the input string, following the given format.
fmt (str): the format of input string. Default is '%Y年%m月%d日 %H:%M'.
verbose (bool): whether print out the parsed time.
Return:
time (pd.Timestamp): parsed time.
'''
import pandas as pd
import datetime
time = pd.to_datetime(datetime.datetime.strptime(time_str, fmt))
if verbose:
print('The time is {}'.format(time))
return time
crush_time = parse_time(crush_time_str, verbose=True)
crush_time
```
### * 和 **
```
def parse_time(time_str, fmt='%Y年%m月%d日 %H:%M', verbose=True, **kwds): # why not using `format`?
'''
Parse the time given as a string.
Parameters:
time_str (str): the input string, following the given format.
fmt (str): the format of input string. Default is '%Y年%m月%d日 %H:%M'.
verbose (bool): whether print out the parsed time.
Return:
time (pd.Timestamp): parsed time.
'''
import pandas as pd
import datetime
time = pd.to_datetime(datetime.datetime.strptime(time_str, fmt))
if 'is_why' in kwds.keys():
print('WHY is comming!!!!!!!')
if verbose:
if 'time_name' in kwds.keys():
print('{0} is {1}'.format(kwds['time_name'], time))
else:
print('Time is {0}'.format(time))
return time
crush_time_str = '2019年10月11日 23:52'
crush_time = parse_time(crush_time_str, verbose=True, time_name='why')
def parse_time(time_str, fmt='%Y年%m月%d日 %H:%M', verbose=True, *args): # why not using `format`?
'''
Parse the time given as a string.
Parameters:
time_str (str): the input string, following the given format.
fmt (str): the format of input string. Default is '%Y年%m月%d日 %H:%M'.
verbose (bool): whether print out the parsed time.
Return:
time (pd.Timestamp): parsed time.
'''
import pandas as pd
import datetime
time = pd.to_datetime(datetime.datetime.strptime(time_str, fmt))
if verbose:
print('{0} is {1}'.format(args, time))
return time
crush_time_str = '2019年10月11日 23:52'
crush_time = parse_time(crush_time_str, '%Y年%m月%d日 %H:%M', True, 'Hahaha')
def multi_sum(*args):
print(args)
s = 0
for item in args:
s += item
return s
multi_sum(1, 2, 3, 4, 5)
```
### assert
```
def parse_time(time_str, fmt='%Y年%m月%d日 %H:%M', verbose=True): # why not using `format`?
'''
Parse the time given as a string.
Parameters:
time_str (str): the input string, following the given format.
fmt (str): the format of input string. Default is '%Y年%m月%d日 %H:%M'.
verbose (bool): whether print out the parsed time.
Return:
time (pd.Timestamp): parsed time.
'''
import pandas as pd
import datetime
assert(isinstance(time_str, str)), '你的数据类型错了!🐧'
time = pd.to_datetime(datetime.datetime.strptime(time_str, fmt))
if verbose:
print('The time is {}'.format(time))
return time
crush_time_str = 2019
crush_time = parse_time(crush_time_str)
```
### 面向对象编程
对象:http://astrojacobli.github.io
🐘🐘
以前写程序的办法都是流程式编程,现在要搞对象(object)。
对象:attributes + methods
class -> object
```
class People:
def __init__(self, name, gender, alias, ):
self.name = name
self.gender = gender
self.alias = alias
self.hunger = 0
self.thirsty = 0
self.health = 100
self.love = 10
def eat(self, amount=1):
if self.hunger - amount >= 0:
self.hunger -= amount
def drink(self, amount=1):
if self.thirsty - amount >= 0:
self.thirsty -= amount
def exercise(self, time=1):
if time > 5:
self.health -= 5 * time
else:
self.health += 10 * time
def meowmeow(self):
self.love += 1
def video_call(self):
self.love += 10
def quarrel(self, others):
assert(isinstance(others, People)), '你不是人!!!'
self.love -= 10
self.health -= 20
self.thirsty += 10
others.love -= 5
others.health -= 10
others.thirsty += 5
def check_healthy(self):
if self.thirsty > 30 or self.hunger > 30:
self.health -= 30
if self.health < 50:
return 'unhealthy'
else:
return 'healthy'
def write_diary(self, stuff=None):
## Basic status
healthy = self.check_healthy()
diary = "I'm {}".format(healthy) + ' today'
diary = "\n".join([diary, stuff])
self.diary = diary
def make_love(self, others):
assert(isinstance(others, People)), '你不是人!!!'
self.love += 100
others.love += 100
babyxuan = People(name='LJX', gender='Male', alias='XX')
babyhuan = People(name='WHY', gender='Female', alias='Huanhuan')
params = 'WHY', 'Female', 'Huanhuan'
babyhuan = People(*params)
babyhuan.__dict__
babyhuan.make_love(babyxuan)
babyxuan.__dict__
babyhuan.__dict__
babyhuan.eat(10)
babyhuan.drink(30)
babyhuan.check_healthy()
babyhuan.write_diary("I don't read some papers today. ")
print(babyhuan.diary)
babyhuan.video_call()
babyhuan.love
babyhuan.__dict__
"I'm unhealthy today".split(' ')
```
其实python里一切东西都是对象。比如字符串有method叫做split.
| github_jupyter |
## TPOT - Auto-ML programming
Following image depicts how TPOT works:
<img src = 'https://res.cloudinary.com/dyd911kmh/image/upload/f_auto,q_auto:best/v1537396029/output_2_0_d7uh0v.png'/>
For more info visit: https://epistasislab.github.io/tpot/examples/
```
# import required modules
import pandas as pd
import numpy as np
data = pd.read_csv('online_shoppers_intention-2.csv')
data.head()
```
## Data prep
It's generally a good idea to randomly **shuffle** the data before starting to avoid any type of ordering in the data.
```
# random data shuffle
data_shuffle=data.iloc[np.random.permutation(len(data))]
data1 = data_shuffle.reset_index(drop=True)
data1.head()
```
- Label cetegorical values and deal with missing values
```
# labeling categorical values
data1.info()
#binning the Month column by quarter(as seen above)
#new column created-month_bin will have months binned by their respective quarters
def Month_bin(Month) :
if Month == 'Jan':
return 1
elif Month == 'Feb':
return 1
elif Month == 'Mar':
return 1
elif Month == 'Apr':
return 2
elif Month == 'May':
return 2
elif Month == 'June':
return 2
elif Month == 'Jul':
return 3
elif Month == 'Aug':
return 3
elif Month == 'Sep':
return 3
elif Month == 'Oct':
return 4
elif Month == 'Nov':
return 4
elif Month == 'Dec':
return 4
data1['Month_bin'] = data1['Month'].apply(Month_bin)
#binning VisitorType
#creating new column--VisitorType_bin
def VisitorType_bin(VisitorType) :
if VisitorType == 'Returning_Visitor':
return 1
elif VisitorType == 'New_Visitor':
return 2
elif VisitorType == 'Other':
return 3
# apply function
data1['VisitorType_bin'] = data1['VisitorType'].apply(VisitorType_bin)
# get dummies
data1 = pd.get_dummies(data1, columns=['VisitorType_bin','Month_bin'])
# convert to bool
data1[['VisitorType_bin_1', 'VisitorType_bin_2', 'VisitorType_bin_3',
'Month_bin_1', 'Month_bin_2', 'Month_bin_3', 'Month_bin_4']] = data1[['VisitorType_bin_1',
'VisitorType_bin_2', 'VisitorType_bin_3','Month_bin_1', 'Month_bin_2', 'Month_bin_3', 'Month_bin_4']].astype(int)
data1 = data1.drop(['Month','VisitorType'], axis = 1)
# tpot doesn't accept bool dtype
data1[['Revenue','Weekend']] = data1[['Revenue','Weekend']].astype(int)
# store target seperetly
target = data1.Revenue.values
# handnling NA values
# assumption exit rates cannot be 0
data1['ExitRates'] = data1['ExitRates'].replace(0,np.NaN)
data1['ExitRates'] = data1['ExitRates'].fillna(data1['ExitRates'].median())
```
## Modeling
- split the DataFrame into a training set and a testing set just like you do while doing any type of machine learning modeling.
- You can do this via sklearn's **cross_validation** **train_test_split**.
```
from sklearn.model_selection import train_test_split
training_indices, testing_indices = train_test_split(data1.index,
stratify = target,
train_size=0.75, test_size=0.25, random_state = 123)
# size of the training set and validation set
training_indices.size, testing_indices.size
```
**tpot** training can take up to several hours to finish BUT there are some hyperparameters than can be adjusted so it does not take forever
- **max_time_mins**: how many minutes TPOT has to optimize the pipeline. If not None, this setting will override the generations parameter and allow TPOT to run until max_time_mins minutes elapse.
- **max_eval_time_mins**: how many minutes TPOT has to evaluate a single pipeline. Setting this parameter to higher values will enable TPOT to evaluate more complex pipelines, but will also allow TPOT to run longer. Use this parameter to help prevent TPOT from wasting time on assessing time-consuming pipelines. The default is 5.
- **early_stop**: how many generations TPOT checks whether there is no improvement in the optimization process. Ends the optimization process if there is no improvement in the given number of generations.
- **n_jobs**: Number of procedures to use in parallel for evaluating pipelines during the TPOT optimization process. Setting n_jobs=-1 will use as many cores as available on the computer. Beware that using multiple methods on the same machine may cause memory issues for large datasets. The default is 1.
- **subsample**: Fraction of training samples that are used during the TPOT optimization process. Must be in the range (0.0, 1.0]. The default is 1.
```
from tpot import TPOTClassifier
tpot = TPOTClassifier(verbosity=2, max_time_mins=30,
max_eval_time_mins=1.2, population_size=30, early_stop=30, n_jobs=-1,scoring="roc_auc")
tpot.fit(data1.drop('Revenue',axis=1).loc[training_indices].values, # X_train
data1.loc[training_indices,'Revenue'].values) # y_train
```
- we trained the model for 30 minutes
- the best pipeline was OneHotEncoder: input_matrix, minimum_fraction=0.25, sparse=False, threshold=10
- best model was random forest with the following hyperparameters
- bootstrap=True, criterion=entropy, max_features=0.4, min_samples_leaf=20, min_samples_split=14, n_estimators=100
One of the key difference here is we use both `X_test` and `y_test` in the code below, since the `.score()` method below combines the __prediction__ and __evaluation__ in the same step.
- our AUC score is 93.06%
```
tpot.score(data1.drop('Revenue',axis=1).loc[testing_indices].values, #X_test
data1.loc[testing_indices, 'Revenue'].values) # y_test
# we can export this pipeline and reuse it
tpot.export('tpot_pipeline.py')
```
| github_jupyter |
<img alt="QuantRocket logo" src="https://www.quantrocket.com/assets/img/notebook-header-logo.png">
© Copyright Quantopian Inc.<br>
© Modifications Copyright QuantRocket LLC<br>
Licensed under the [Creative Commons Attribution 4.0](https://creativecommons.org/licenses/by/4.0/legalcode).
<a href="https://www.quantrocket.com/disclaimer/">Disclaimer</a>
# Residuals Analysis
By Chris Fenaroli and Max Margenot
## Linear Regression
Linear regression is one of our most fundamental modeling techniques. We use it to estimate a linear relationship between a set of independent variables $X_i$ and a dependent outcome variable $y$. Our model takes the form of:
$$ y_i = \beta_{0} 1 + \beta_{i, 1} x_{i, 1} + \dots + \beta_{i, p} x_{i, p} + \epsilon_i = x_i'\beta + \epsilon_i $$
For $i \in \{1, \dots, n\}$, where $n$ is the number of observations. We write this in vector form as:
$$ y = X\beta + \epsilon $$
Where $y$ is a $n \times 1$ vector, $X$ is a $n \times p$ matrix, $\beta$ is a $p \times 1$ vector of coefficients, and $\epsilon$ is a standard normal error term. Typically we call a model with $p = 1$ a simple linear regression and a model with $p > 1$ a multiple linear regression. More background information on regressions can be found in the lectures on [simple linear regression](Lecture12-Linear-Regression.ipynb) and [multiple linear regression](Lecture15-Multiple-Linear-Regression.ipynb).
Whenever we build a model, there will be gaps between what a model predicts and what is observed in the sample. The differences between these values are known as the residuals of the model and can be used to check for some of the basic assumptions that go into the model. The key assumptions to check for are:
* **Linear Fit:** The underlying relationship should be linear
* **Homoscedastic:** The data should have no trend in the variance
* **Independent and Identically Distributed:** The residuals of the regression should be independent and identically distributed (i.i.d.) and show no signs of serial correlation
We can use the residuals to help diagnose whether the relationship we have estimated is real or spurious.
Statistical error is a similar metric associated with regression analysis with one important difference: While residuals quantify the gap between a regression model predictions and the observed sample, statistical error is the difference between a regression model and the unobservable expected value. We use residuals in an attempt to estimate this error.
```
# Import libraries
import numpy as np
import pandas as pd
from statsmodels import regression
import statsmodels.api as sm
import statsmodels.stats.diagnostic as smd
import scipy.stats as stats
import matplotlib.pyplot as plt
import math
```
### Simple Linear Regression
First we'll define a function that performs linear regression and plots the results.
```
def linreg(X,Y):
# Running the linear regression
X = sm.add_constant(X)
model = sm.OLS(Y, X).fit()
B0 = model.params[0]
B1 = model.params[1]
X = X[:, 1]
# Return summary of the regression and plot results
X2 = np.linspace(X.min(), X.max(), 100)
Y_hat = X2 * B1 + B0
plt.scatter(X, Y, alpha=1) # Plot the raw data
plt.plot(X2, Y_hat, 'r', alpha=1); # Add the regression line, colored in red
plt.xlabel('X Value')
plt.ylabel('Y Value')
return model, B0, B1
```
Let's define a toy relationship between $X$ and $Y$ that we can model with a linear regression. Here we define the relationship and construct a model on it, drawing the determined line of best fit with the regression parameters.
```
n = 50
X = np.random.randint(0, 100, n)
epsilon = np.random.normal(0, 1, n)
Y = 10 + 0.5 * X + epsilon
linreg(X,Y)[0];
print("Line of best fit: Y = {0} + {1}*X".format(linreg(X, Y)[1], linreg(X, Y)[2]))
```
This toy example has some generated noise, but all real data will also have noise. This is inherent in sampling from any sort of wild data-generating process. As a result, our line of best fit will never exactly fit the data (which is why it is only "best", not "perfect"). Having a model that fits every single observation that you have is a sure sign of overfitting.
For all fit models, there will be a difference between what the regression model predicts and what was observed, which is where residuals come in.
## Residuals
The definition of a residual is the difference between what is observed in the sample and what is predicted by the regression. For any residual $r_i$, we express this as
$$r_i = Y_i - \hat{Y_i}$$
Where $Y_i$ is the observed $Y$-value and $\hat{Y}_i$ is the predicted Y-value. We plot these differences on the following graph:
```
model, B0, B1 = linreg(X,Y)
residuals = model.resid
plt.errorbar(X,Y,xerr=0,yerr=[residuals,0*residuals],linestyle="None",color='Green');
```
We can pull the residuals directly out of the fit model.
```
residuals = model.resid
print(residuals)
```
### Diagnosing Residuals
Many of the assumptions that are necessary to have a valid linear regression model can be checked by identifying patterns in the residuals of that model. We can make a quick visual check by looking at the residual plot of a given model.
With a residual plot, we look at the predicted values of the model versus the residuals themselves. What we want to see is just a cloud of unrelated points, like so:
```
plt.scatter(model.predict(), residuals);
plt.axhline(0, color='red')
plt.xlabel('Predicted Values');
plt.ylabel('Residuals');
plt.xlim([1,50]);
```
What we want is a fairly random distribution of residuals. The points should form no discernible pattern. This would indicate that a plain linear model is likely a good fit. If we see any sort of trend, this might indicate the presence of autocorrelation or heteroscedasticity in the model.
## Appropriateness of a Linear Model
By looking for patterns in residual plots we can determine whether a linear model is appropriate in the first place. A plain linear regression would not be appropriate for an underlying relationship of the form:
$$Y = \beta_0 + \beta_1 X^2$$
as a linear function would not be able to fully explain the relationship between $X$ and $Y$.
If the relationship is not a good fit for a linear model, the residual plot will show a distinct pattern. In general, a residual plot of a linear regression on a non-linear relationship will show bias and be asymmetrical with respect to residual = 0 line while a residual plot of a linear regression on a linear relationship will be generally symmetrical over the residual = 0 axis.
As an example, let's consider a new relationship between the variables $X$ and $Y$ that incorporates a quadratic term.
```
n = 50
X = np.random.randint(0, 50, n)
epsilon = np.random.normal(0, 1, n)
Y_nonlinear = 10 - X**1.2 + epsilon
model = sm.OLS(Y_nonlinear, sm.add_constant(X)).fit()
B0, B1 = model.params
residuals = model.resid
print('beta_0:', B0)
print('beta_1:', B1)
plt.scatter(model.predict(), residuals);
plt.axhline(0, color='red')
plt.xlabel('Predicted Values');
plt.ylabel('Residuals');
```
The "inverted-U" shape shown by the residuals is a sign that a non-linear model might be a better fit than a linear one.
## Heteroscedasticity
One of the main assumptions behind a linear regression is that the underlying data has a constant variance. If there are some parts of the data with a variance different from another part the data is not appropriate for a linear regression. **Heteroscedasticity** is a term that refers to data with non-constant variance, as opposed to homoscedasticity, when data has constant variance.
Significant heteroscedasticity invalidates linear regression results by biasing the standard error of the model. As a result, we can't trust the outcomes of significance tests and confidence intervals generated from the model and its parameters.
To avoid these consequences it is important to use residual plots to check for heteroscedasticity and adjust if necessary.
As an example of detecting and correcting heteroscedasticity, let's consider yet another relationship between $X$ and $Y$:
```
n = 50
X = np.random.randint(0, 100, n)
epsilon = np.random.normal(0, 1, n)
Y_heteroscedastic = 100 + 2*X + epsilon*X
model = sm.OLS(Y_heteroscedastic, sm.add_constant(X)).fit()
B0, B1 = model.params
residuals = model.resid
plt.scatter(model.predict(), residuals);
plt.axhline(0, color='red')
plt.xlabel('Predicted Values');
plt.ylabel('Residuals');
```
Heteroscedasticity often manifests as this spread, giving us a tapered cloud in one direction or another. As we move along in the $x$-axis, the magnitudes of the residuals are clearly increasing. A linear regression is unable to explain this varying variability and the regression standard errors will be biased.
### Statistical Methods for Detecting Heteroscedasticity
Generally, we want to back up qualitative observations on a residual plot with a quantitative method. The residual plot led us to believe that the data might be heteroscedastic. Let's confirm that result with a statistical test.
A common way to test for the presence of heteroscedasticity is the Breusch-Pagan hypothesis test. It's good to combine the qualitative analysis of a residual plot with the quantitative analysis of at least one hypothesis test. We can add the White test as well, but for now we will use only Breush-Pagan to test our relationship above. A function exists in the `statsmodels` package called `het_breushpagan` that simplifies the computation:
```
breusch_pagan_p = smd.het_breuschpagan(model.resid, model.model.exog)[1]
print(breusch_pagan_p)
if breusch_pagan_p > 0.05:
print("The relationship is not heteroscedastic.")
if breusch_pagan_p < 0.05:
print("The relationship is heteroscedastic.")
```
We set our confidence level at $\alpha = 0.05$, so a Breusch-Pagan p-value below $0.05$ tells us that the relationship is heteroscedastic. For more on hypothesis tests and interpreting p-values, refer to the lecture on hypothesis testing. Using a hypothesis test bears the risk of a false positive or a false negative, which is why it can be good to confirm with additional tests if we are skeptical.
### Adjusting for Heteroscedasticity
If, after creating a residual plot and conducting tests, you believe you have heteroscedasticity, there are a number of methods you can use to attempt to adjust for it. The three we will focus on are differences analysis, log transformations, and Box-Cox transformations.
#### Differences Analysis
A differences analysis involves looking at the first-order differences between adjacent values. With this, we are looking at the changes from period to period of an independent variable rather than looking directly at its values. Often, by looking at the differences instead of the raw values, we can remove heteroscedasticity. We correct for it and can use the ensuing model on the differences.
```
# Finding first-order differences in Y_heteroscedastic
Y_heteroscedastic_diff = np.diff(Y_heteroscedastic)
```
Now that we have stored the first-order differences of `Y_heteroscedastic` in `Y_heteroscedastic_diff` let's repeat the regression and residual plot to see if the heteroscedasticity is still present:
```
model = sm.OLS(Y_heteroscedastic_diff, sm.add_constant(X[1:])).fit()
B0, B1 = model.params
residuals = model.resid
plt.scatter(model.predict(), residuals);
plt.axhline(0, color='red')
plt.xlabel('Predicted Values');
plt.ylabel('Residuals');
breusch_pagan_p = smd.het_breuschpagan(residuals, model.model.exog)[1]
print(breusch_pagan_p)
if breusch_pagan_p > 0.05:
print("The relationship is not heteroscedastic.")
if breusch_pagan_p < 0.05:
print("The relationship is heteroscedastic.")
```
*Note: This new regression was conducted on the differences between data, and therefore the regression output must be back-transformed to reach a prediction in the original scale. Since we regressed the differences, we can add our predicted difference onto the original data to get our estimate:*
$$\hat{Y_i} = Y_{i-1} + \hat{Y}_{diff}$$
#### Logarithmic Transformation
Next, we apply a log transformation to the underlying data. A log transformation will bring residuals closer together and ideally remove heteroscedasticity. In many (though not all) cases, a log transformation is sufficient in stabilizing the variance of a relationship.
```
# Taking the log of the previous data Y_heteroscedastic and saving it in Y_heteroscedastic_log
Y_heteroscedastic_log = np.log(Y_heteroscedastic)
```
Now that we have stored the log transformed version of `Y_heteroscedastic` in `Y_heteroscedastic_log` let's repeat the regression and residual plot to see if the heteroscedasticity is still present:
```
model = sm.OLS(Y_heteroscedastic_log, sm.add_constant(X)).fit()
B0, B1 = model.params
residuals = model.resid
plt.scatter(model.predict(), residuals);
plt.axhline(0, color='red')
plt.xlabel('Predicted Values');
plt.ylabel('Residuals');
# Running and interpreting a Breusch-Pagan test
breusch_pagan_p = smd.het_breuschpagan(residuals, model.model.exog)[1]
print(breusch_pagan_p)
if breusch_pagan_p > 0.05:
print("The relationship is not heteroscedastic.")
if breusch_pagan_p < 0.05:
print("The relationship is heteroscedastic.")
```
*Note: This new regression was conducted on the log of the original data. This means the scale has been altered and the regression estimates will lie on this transformed scale. To bring the estimates back to the original scale, you must back-transform the values using the inverse of the log:*
$$\hat{Y} = e^{\log(\hat{Y})}$$
#### Box-Cox Transformation
Finally, we examine the Box-Cox transformation. The Box-Cox transformation is a powerful method that will work on many types of heteroscedastic relationships. The process works by testing all values of $\lambda$ within the range $[-5, 5]$ to see which makes the output of the following equation closest to being normally distributed:
$$
Y^{(\lambda)} = \begin{cases}
\frac{Y^{\lambda}-1}{\lambda} & : \lambda \neq 0\\ \log{Y} & : \lambda = 0
\end{cases}
$$
The "best" $\lambda$ will be used to transform the series along the above function. Instead of having to do all of this manually, we can simply use the `scipy` function `boxcox`. We use this to adjust $Y$ and hopefully remove heteroscedasticity.
*Note: The Box-Cox transformation can only be used if all the data is positive*
```
# Finding a power transformation adjusted Y_heteroscedastic
Y_heteroscedastic_box_cox = stats.boxcox(Y_heteroscedastic)[0]
```
Now that we have stored the power transformed version of `Y_heteroscedastic` in `Y_heteroscedastic_prime` let's repeat the regression and residual plot to see if the heteroscedasticity is still present:
```
model = sm.OLS(Y_heteroscedastic_box_cox, sm.add_constant(X)).fit()
B0, B1 = model.params
residuals = model.resid
plt.scatter(model.predict(), residuals);
plt.axhline(0, color='red')
plt.xlabel('Predicted Values');
plt.ylabel('Residuals');
# Running and interpreting a Breusch-Pagan test
breusch_pagan_p = smd.het_breuschpagan(residuals, model.model.exog)[1]
print(breusch_pagan_p)
if breusch_pagan_p > 0.05:
print("The relationship is not heteroscedastic.")
if breusch_pagan_p < 0.05:
print("The relationship is heteroscedastic.")
```
*Note: Now that the relationship is not heteroscedastic, a linear regression is appropriate. However, because the data was power transformed, the regression estimates will be on a different scale than the original data. This is why it is important to remember to back-transform results using the inverse of the Box-Cox function:*
$$\hat{Y} = (Y^{(\lambda)}\lambda + 1)^{1/\lambda}$$
### GARCH Modeling
Another approach to dealing with heteroscadasticity is through a GARCH (generalized autoregressive conditional heteroscedasticity) model. More information can be found in the lecture on GARCH modeling.
## Residuals and Autocorrelation
Another assumption behind linear regressions is that the residuals are not autocorrelated. A series is autocorrelated when it is correlated with a delayed version of itself. An example of a potentially autocorrelated time series series would be daily high temperatures. Today's temperature gives you information on tomorrow's temperature with reasonable confidence (i.e. if it is 90 °F today, you can be very confident that it will not be below freezing tomorrow). A series of fair die rolls, however, would not be autocorrelated as seeing one roll gives you no information on what the next might be. Each roll is independent of the last.
In finance, stock prices are usually autocorrelated while stock returns are independent from one day to the next. We represent a time dependency on previous values like so:
$$Y_i = Y_{i-1} + \epsilon$$
If the residuals of a model are autocorrelated, you will be able to make predictions about adjacent residuals. In the case of $Y$, we know the data will be autocorrelated because we can make predictions based on adjacent residuals being close to one another.
```
n = 50
X = np.linspace(0, n, n)
Y_autocorrelated = np.zeros(n)
Y_autocorrelated[0] = 50
for t in range(1, n):
Y_autocorrelated[t] = Y_autocorrelated[t-1] + np.random.normal(0, 1)
# Regressing X and Y_autocorrelated
model = sm.OLS(Y_autocorrelated, sm.add_constant(X)).fit()
B0, B1 = model.params
residuals = model.resid
plt.scatter(model.predict(), residuals);
plt.axhline(0, color='red')
plt.xlabel('Predicted Values');
plt.ylabel('Residuals');
```
Autocorrelation in the residuals in this example is not explicitly obvious, so our check is more to make absolutely certain.
### Statistical Methods for Detecting Autocorrelation
As with all statistical properties, we require a statistical test to ultimately decide whether there is autocorrelation in our residuals or not. To this end, we use a Ljung-Box test.
A Ljung-Box test is used to detect autocorrelation in a time series. The Ljung-Box test examines autocorrelation at all lag intervals below a specified maximum and returns arrays containing the outputs for every tested lag interval.
Let's use the `acorr_ljungbox` function in `statsmodels` to test for autocorrelation in the residuals of our above model. We use a max lag interval of $10$, and see if any of the lags have significant autocorrelation:
```
ljung_box = smd.acorr_ljungbox(residuals, lags=10, return_df=True)
print("Lagrange Multiplier Statistics:", ljung_box.lb_stat.tolist())
print("\nP-values:", ljung_box.lb_pvalue.tolist(), "\n")
if (ljung_box.lb_pvalue < 0.05).any():
print("The residuals are autocorrelated.")
else:
print("The residuals are not autocorrelated.")
```
Because the Ljung-Box test yielded a p-value below $0.05$ for at least one lag interval, we can conclude that the residuals of our model are autocorrelated.
## Adjusting for Autocorrelation
We can adjust for autocorrelation in many of the same ways that we adjust for heteroscedasticity. Let's see if a model on the first-order differences of $Y$ has autocorrelated residuals:
```
# Finding first-order differences in Y_autocorrelated
Y_autocorrelated_diff = np.diff(Y_autocorrelated)
model = sm.OLS(Y_autocorrelated_diff, sm.add_constant(X[1:])).fit()
B0, B1 = model.params
residuals = model.resid
plt.scatter(model.predict(), residuals);
plt.axhline(0, color='red')
plt.xlabel('Predicted Values');
plt.ylabel('Residuals');
# Running and interpreting a Ljung-Box test
ljung_box = smd.acorr_ljungbox(residuals, lags=10, return_df=True)
print("P-values:", ljung_box.lb_pvalue.tolist(), "\n")
if (ljung_box.lb_pvalue < 0.05).any():
print("The residuals are autocorrelated.")
else:
print("The residuals are not autocorrelated.")
```
*Note: This new regression was conducted on the differences between data, and therefore the regression output must be back-transformed to reach a prediction in the original scale. Since we regressed the differences, we can add our predicted difference onto the original data to get our estimate:*
$$\hat{Y_i} = Y_{i-1} + \hat{Y_{diff}}$$
We can also perform a log transformation, if we so choose. This process is identical to the one we performed on the heteroscedastic data up above, so we will leave it out this time.
## Example: Market Beta Calculation
Let's calculate the market beta between AAPL and SPY using a simple linear regression, and then conduct a residual analysis on the regression to ensure the validity of our results. To regress AAPL and SPY, we will focus on their returns, not their price, and set SPY returns as our independent variable and AAPL returns as our outcome variable. The regression will give us a line of best fit:
$$\hat{r_{AAPL}} = \hat{\beta_0} + \hat{\beta_1}r_{SPY}$$
The slope of the regression line $\hat{\beta_1}$ will represent our market beta, as for every $r$ percent change in the returns of SPY, the predicted returns of AAPL will change by $\hat{\beta_1}$.
Let's start by conducting the regression the returns of the two assets.
```
from quantrocket.master import get_securities
from quantrocket import get_prices
securities = get_securities(symbols=["AAPL", "SPY"], vendors='usstock')
start = '2017-01-01'
end = '2018-01-01'
closes = get_prices("usstock-free-1min", data_frequency="daily", sids=securities.index.tolist(), fields='Close', start_date=start, end_date=end).loc['Close']
sids_to_symbols = securities.Symbol.to_dict()
closes = closes.rename(columns=sids_to_symbols)
asset = closes['AAPL']
benchmark = closes['SPY']
# We have to take the percent changes to get to returns
# Get rid of the first (0th) element because it is NAN
r_a = asset.pct_change()[1:].values
r_b = benchmark.pct_change()[1:].values
# Regressing the benchmark b and asset a
r_b = sm.add_constant(r_b)
model = sm.OLS(r_a, r_b).fit()
r_b = r_b[:, 1]
B0, B1 = model.params
# Plotting the regression
A_hat = (B1*r_b + B0)
plt.scatter(r_b, r_a, alpha=1) # Plot the raw data
plt.plot(r_b, A_hat, 'r', alpha=1); # Add the regression line, colored in red
plt.xlabel('AAPL Returns')
plt.ylabel('SPY Returns')
# Print our result
print("Estimated AAPL Beta:", B1)
# Calculating the residuals
residuals = model.resid
```
Our regression yielded an estimated market beta of 1.36; according to the regression, for every 1% in return we see from the SPY, we should see 1.36% from AAPL.
Now that we have the regression results and residuals, we can conduct our residual analysis. Our first step will be to plot the residuals and look for any red flags:
```
plt.scatter(model.predict(), residuals);
plt.axhline(0, color='red')
plt.xlabel('AAPL Returns');
plt.ylabel('Residuals');
```
By simply observing the distribution of residuals, it does not seem as if there are any abnormalities. The distribution is relatively random and no patterns can be observed (the clustering around the origin is a result of the nature of returns to cluster around 0 and is not a red flag). Our qualitative conclusion is that the data is homoscedastic and not autocorrelated and therefore satisfies the assumptions for linear regression.
## Breusch-Pagan Heteroscedasticity Test
Our qualitative assessment of the residual plot is nicely supplemented with a couple statistical tests. Let's begin by testing for heteroscedasticity using a Breusch-Pagan test. Using the `het_breuschpagan` function from the statsmodels package:
```
bp_test = smd.het_breuschpagan(residuals, model.model.exog)
print("Lagrange Multiplier Statistic:", bp_test[0])
print("P-value:", bp_test[1])
print("f-value:", bp_test[2])
print("f_p-value:", bp_test[3], "\n")
if bp_test[1] > 0.05:
print("The relationship is not heteroscedastic.")
if bp_test[1] < 0.05:
print("The relationship is heteroscedastic.")
```
Because the P-value is greater than 0.05, we do not have enough evidence to reject the null hypothesis that the relationship is homoscedastic. This result matches up with our qualitative conclusion.
## Ljung-Box Autocorrelation Test
Let's also check for autocorrelation quantitatively using a Ljung-Box test. Using the `acorr_ljungbox` function from the statsmodels package and the default maximum lag:
```
ljung_box = smd.acorr_ljungbox(r_a, lags=1, return_df=True)
print("P-Values:", ljung_box.lb_pvalue.tolist(), "\n")
if (ljung_box.lb_pvalue < 0.05).any():
print("The residuals are autocorrelated.")
else:
print("The residuals are not autocorrelated.")
```
Because the Ljung-Box test yielded p-values above 0.05 for all lags, we can conclude that the residuals are not autocorrelated. This result matches up with our qualitative conclusion.
After having visually assessed the residual plot of the regression and then backing it up using statistical tests, we can conclude that the data satisfies the main assumptions and the linear model is valid.
## References
* "Analysis of Financial Time Series", by Ruey Tsay
---
**Next Lecture:** [Dangers of Overfitting](Lecture19-Dangers-of-Overfitting.ipynb)
[Back to Introduction](Introduction.ipynb)
---
*This presentation is for informational purposes only and does not constitute an offer to sell, a solicitation to buy, or a recommendation for any security; nor does it constitute an offer to provide investment advisory or other services by Quantopian, Inc. ("Quantopian") or QuantRocket LLC ("QuantRocket"). Nothing contained herein constitutes investment advice or offers any opinion with respect to the suitability of any security, and any views expressed herein should not be taken as advice to buy, sell, or hold any security or as an endorsement of any security or company. In preparing the information contained herein, neither Quantopian nor QuantRocket has taken into account the investment needs, objectives, and financial circumstances of any particular investor. Any views expressed and data illustrated herein were prepared based upon information believed to be reliable at the time of publication. Neither Quantopian nor QuantRocket makes any guarantees as to their accuracy or completeness. All information is subject to change and may quickly become unreliable for various reasons, including changes in market conditions or economic circumstances.*
| github_jupyter |
```
from sklearn.tree import DecisionTreeClassifier
import pandas as pd
clf = DecisionTreeClassifier()
```
# Particles
```
import numpy as np, random, math
class Particle:
def __init__(self, size):
self.position = np.array([random.choice((0, 1)) for _ in range(size)])
self.velocity = np.array([random.uniform(0, 1) for _ in range(size)])
self.best = 0
self.currBest = 0
self.currBestPosition = self.position
self.inertiaWeight = random.uniform(0, 1)
def update_velocity(self, c1, c2, particleBestPosition, vmax = 4, vmin = -4):
self.velocity = np.array([self.calculate_velocity(v, c1, c2, px, pbx, x) for v, px, x, pbx in zip(self.velocity, self.position, self.currBestPosition, particleBestPosition)])
for i in range(len(self.velocity)):
if self.velocity[i] > vmax :
self.velocity[i] = vmax
elif self.velocity[i] < vmin :
self.velocity[i] = vmin
def update_position(self):
self.position = np.array([(1 if self.sigmoid(v) > random.uniform(0, 1) else 0) for v in self.velocity])
def calculate_velocity(self, v0, c1, c2, px, pbx, x):
return self.inertiaWeight * v0 + c1 * random.uniform(0, 1) * (-(px - pbx)) + c2 * random.uniform(0, 1) * (-(px - x))
def sigmoid(self, v):
if v < 0:
return 1 - (1 / (1 + math.exp(-v)))
return 1 / (1 + math.exp(-v))
# def calculate_best(self, train, test):
# pos = self.position.astype(bool)
# # tfidf = TFIDF(train["Review"])
# # tfidf.weights = tfidf.remove_zero_tfidf(tfidf.weights, 0.5)
# # tfidf.termIndex = {key:val for i, (key, val) in enumerate(tfidf.termIndex.items()) if pos[i] == True}
# # print(f"Selected attributes: {len(tfidf.termIndex)}")
# clf = C45(tfidf, train)
# clf.train()
# self.best = clf.score(tfidf, test)
# return self.best
def calculate_best(self, xtrain,ytrain, xtest, ytest):
pos = self.position.astype(bool)
x_train=dataFrame(pos,xtrain)
y_train=ytrain.values
x_test=dataFrame(pos,xtest)
y_test=ytest.values
#clf = DecisionTreeClassifier()
clf.fit(x_train,y_train)
self.best = clf.score(x_test,y_test)
return self.best
def tent_map(self):
if self.inertiaWeight < 0.7:
self.inertiaWeight = self.inertiaWeight / 0.7
else:
self.inertiaWeight = (10 / 3) * (self.inertiaWeight * (1 - self.inertiaWeight))
return self.inertiaWeight
def __repr__(self):
return '<%s.%s object at %s>' % (
self.__class__.__module__,
self.__class__.__name__,
hex(id(self))
)
```
# PSO
```
import random
class PSO:
def __init__(self, particleSize, populationSize, numIteration, c1, c2, target):
self.particleSize = particleSize
self.populationSize = populationSize
self.numIteration = numIteration
self.c1 = c1
self.c2 = c2
self.target = target
self.particles = [Particle(self.particleSize) for _ in range(self.populationSize)]
self.iterationBest = []
def exec(self, xtrain, ytrain, xtest, ytest):
for _ in range(self.numIteration):
for i in range(self.populationSize):
print(self.particles[i].position)
b = self.particles[i].calculate_best(xtrain, ytrain, xtest, ytest)
print(f"Iter-{_} Particle-{i} best: {b}")
self.particles[i].tent_map()
self.particles = sorted(self.particles, key=lambda particle: particle.best, reverse=True)
self.iterationBest.append(self.particles[0])
#print("party123",self.particles[0].currBestPosition)
print(f"Target: {self.target}")
print(f"Iteration {_} best: {self.particles[0].best}")
if self.particles[0].best > self.target:
return self.particles[0]
for i in range(self.populationSize):
self.particles[i].update_velocity(self.c1, self.c2, self.particles[0].position)
self.particles[i].update_position()
self.iterationBest = sorted(self.iterationBest, key=lambda particle: particle.best, reverse=True)
#scoring(self.iterationBest[0].position)
#scoring(self.iterationBest[0].currBestPosition)
return self.iterationBest[0]
#Creating Data Frame from the selected Features(Genes)
def dataFrame(pos,df):
df=df.loc[:,pos==True]
return df.values
#Driver Function of the Optimisation Model
def optimize_model(xtrain,ytrain, xtest, ytest):
results = []
particleSize = xtrain.shape[1]
popSize=xtrain.shape[0]
numIteration=10
c1=2
c2=2
target=0.98
pso = PSO(particleSize, popSize, numIteration, c1, c2,target)
bestParticle = pso.exec(xtrain,ytrain, xtest, ytest)
results.append(bestParticle)
return bestParticle
xtrain=pd.read_csv('xtrain.csv',delimiter=',')
ytrain=pd.read_csv('ytrain.csv',delimiter=',')
xtest=pd.read_csv('xtest.csv',delimiter=',')
ytest=pd.read_csv('ytest.csv',delimiter=',')
xtrain.shape
#xtest.head()
#ytrain
xtrain.shape,xtest.shape,ytrain.shape,ytest.shape
xtrain=xtrain.iloc[:,1:]
xtest=xtest.iloc[:,1:]
xtrain.shape,xtest.shape
ytrain=ytrain['cancer']
ytrain = ytrain.replace({'ALL':0,'AML':1})
ytest=ytest['cancer']
ytest = ytest.replace({'ALL':0,'AML':1})
xtrain.shape,xtest.shape,ytrain.shape,ytest.shape
resu=optimize_model(xtrain,ytrain, xtest, ytest)
# print("hello",resu.currBestPosition)
# print("hello",resu.position)
pos = resu.position.astype(bool)
x_train=dataFrame(pos,xtrain)
x_test=dataFrame(pos,xtest)
y_train=ytrain.values
y_test=ytest.values
print(x_train.shape,x_test.shape,y_train.shape,y_test.shape)
#clf = DecisionTreeClassifier()
clf.fit(x_train,y_train)
sc = clf.score(x_test,y_test)
sc
#note-currentBestposition is optimal as it's value is ranging from 58% to 67% but in position it
#is varying in vast range from 47% to 67%
pos = resu.currBestPosition.astype(bool)
x_train=dataFrame(pos,xtrain)
x_test=dataFrame(pos,xtest)
y_train=ytrain.values
y_test=ytest.values
print(x_train.shape,x_test.shape,y_train.shape,y_test.shape)
#clf = DecisionTreeClassifier()
clf.fit(x_train,y_train)
sc = clf.score(x_test,y_test)
sc
```
| github_jupyter |
Deep Learning Models -- A collection of various deep learning architectures, models, and tips for TensorFlow and PyTorch in Jupyter Notebooks.
- Author: Sebastian Raschka
- GitHub Repository: https://github.com/rasbt/deeplearning-models
```
%load_ext watermark
%watermark -a 'Sebastian Raschka' -v -p tensorflow
```
# Model Zoo -- Multilayer Perceptron
### Low-level Implementation
```
import tensorflow as tf
from tensorflow.examples.tutorials.mnist import input_data
##########################
### DATASET
##########################
mnist = input_data.read_data_sets("./", one_hot=True)
##########################
### SETTINGS
##########################
# Hyperparameters
learning_rate = 0.1
training_epochs = 10
batch_size = 64
# Architecture
n_hidden_1 = 128
n_hidden_2 = 256
n_input = 784
n_classes = 10
##########################
### GRAPH DEFINITION
##########################
g = tf.Graph()
with g.as_default():
# Input data
tf_x = tf.placeholder(tf.float32, [None, n_input], name='features')
tf_y = tf.placeholder(tf.float32, [None, n_classes], name='targets')
# Model parameters
weights = {
'h1': tf.Variable(tf.truncated_normal([n_input, n_hidden_1], stddev=0.1)),
'h2': tf.Variable(tf.truncated_normal([n_hidden_1, n_hidden_2], stddev=0.1)),
'out': tf.Variable(tf.truncated_normal([n_hidden_2, n_classes], stddev=0.1))
}
biases = {
'b1': tf.Variable(tf.zeros([n_hidden_1])),
'b2': tf.Variable(tf.zeros([n_hidden_2])),
'out': tf.Variable(tf.zeros([n_classes]))
}
# Multilayer perceptron
layer_1 = tf.add(tf.matmul(tf_x, weights['h1']), biases['b1'])
layer_1 = tf.nn.relu(layer_1)
layer_2 = tf.add(tf.matmul(layer_1, weights['h2']), biases['b2'])
layer_2 = tf.nn.relu(layer_2)
out_layer = tf.matmul(layer_2, weights['out']) + biases['out']
# Loss and optimizer
loss = tf.nn.softmax_cross_entropy_with_logits(logits=out_layer, labels=tf_y)
cost = tf.reduce_mean(loss, name='cost')
optimizer = tf.train.GradientDescentOptimizer(learning_rate=learning_rate)
train = optimizer.minimize(cost, name='train')
# Prediction
correct_prediction = tf.equal(tf.argmax(tf_y, 1), tf.argmax(out_layer, 1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32), name='accuracy')
##########################
### TRAINING & EVALUATION
##########################
with tf.Session(graph=g) as sess:
sess.run(tf.global_variables_initializer())
for epoch in range(training_epochs):
avg_cost = 0.
total_batch = mnist.train.num_examples // batch_size
for i in range(total_batch):
batch_x, batch_y = mnist.train.next_batch(batch_size)
_, c = sess.run(['train', 'cost:0'], feed_dict={'features:0': batch_x,
'targets:0': batch_y})
avg_cost += c
train_acc = sess.run('accuracy:0', feed_dict={'features:0': mnist.train.images,
'targets:0': mnist.train.labels})
valid_acc = sess.run('accuracy:0', feed_dict={'features:0': mnist.validation.images,
'targets:0': mnist.validation.labels})
print("Epoch: %03d | AvgCost: %.3f" % (epoch + 1, avg_cost / (i + 1)), end="")
print(" | Train/Valid ACC: %.3f/%.3f" % (train_acc, valid_acc))
test_acc = sess.run(accuracy, feed_dict={'features:0': mnist.test.images,
'targets:0': mnist.test.labels})
print('Test ACC: %.3f' % test_acc)
```
### tensorflow.layers Abstraction
```
import tensorflow as tf
from tensorflow.examples.tutorials.mnist import input_data
##########################
### DATASET
##########################
mnist = input_data.read_data_sets("./", one_hot=True)
##########################
### SETTINGS
##########################
# Hyperparameters
learning_rate = 0.1
training_epochs = 10
batch_size = 64
# Architecture
n_hidden_1 = 128
n_hidden_2 = 256
n_input = 784
n_classes = 10
##########################
### GRAPH DEFINITION
##########################
g = tf.Graph()
with g.as_default():
# Input data
tf_x = tf.placeholder(tf.float32, [None, n_input], name='features')
tf_y = tf.placeholder(tf.float32, [None, n_classes], name='targets')
# Multilayer perceptron
layer_1 = tf.layers.dense(tf_x, n_hidden_1, activation=tf.nn.relu,
kernel_initializer=tf.truncated_normal_initializer(stddev=0.1))
layer_2 = tf.layers.dense(layer_1, n_hidden_2, activation=tf.nn.relu,
kernel_initializer=tf.truncated_normal_initializer(stddev=0.1))
out_layer = tf.layers.dense(layer_2, n_classes, activation=None)
# Loss and optimizer
loss = tf.nn.softmax_cross_entropy_with_logits(logits=out_layer, labels=tf_y)
cost = tf.reduce_mean(loss, name='cost')
optimizer = tf.train.GradientDescentOptimizer(learning_rate=learning_rate)
train = optimizer.minimize(cost, name='train')
# Prediction
correct_prediction = tf.equal(tf.argmax(tf_y, 1), tf.argmax(out_layer, 1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32), name='accuracy')
##########################
### TRAINING & EVALUATION
##########################
with tf.Session(graph=g) as sess:
sess.run(tf.global_variables_initializer())
for epoch in range(training_epochs):
avg_cost = 0.
total_batch = mnist.train.num_examples // batch_size
for i in range(total_batch):
batch_x, batch_y = mnist.train.next_batch(batch_size)
_, c = sess.run(['train', 'cost:0'], feed_dict={'features:0': batch_x,
'targets:0': batch_y})
avg_cost += c
train_acc = sess.run('accuracy:0', feed_dict={'features:0': mnist.train.images,
'targets:0': mnist.train.labels})
valid_acc = sess.run('accuracy:0', feed_dict={'features:0': mnist.validation.images,
'targets:0': mnist.validation.labels})
print("Epoch: %03d | AvgCost: %.3f" % (epoch + 1, avg_cost / (i + 1)), end="")
print(" | Train/Valid ACC: %.3f/%.3f" % (train_acc, valid_acc))
test_acc = sess.run('accuracy:0', feed_dict={'features:0': mnist.test.images,
'targets:0': mnist.test.labels})
print('Test ACC: %.3f' % test_acc)
```
| github_jupyter |
# 集合
Python 中set与dict类似,也是一组key的集合,但不存储value。由于key不能重复,所以,在set中,没有重复的key。
所以没有重复的元素
注意,key为不可变类型,即可哈希的值。
```
num = {}
print(type(num)) # <class 'dict'>
num = {1, 2, 3, 4}
print(type(num)) # <class 'set'>
```
## 1. 集合的创建
- 先创建对象再加入元素。
- 在创建**空集合**的时候只能使用s = set(),因为s = {}创建的是空字典。
```
basket = set()
basket.add('apple')
basket.add('banana')
print(basket) # {'banana', 'apple'}
```
直接把一堆元素用花括号括起来{元素1, 元素2, ..., 元素n}。
重复元素在set中会被自动被过滤。
```
basket = {'apple', 'orange', 'apple', 'pear', 'orange', 'banana'}
print(basket) # {'banana', 'apple', 'pear', 'orange'}
```
- 使用set(value)工厂函数,把列表或元组转换成集合。
```
a = set('abracadabra')
print(a)
# {'r', 'b', 'd', 'c', 'a'}
b = set(("Google", "Lsgogroup", "Taobao", "Taobao"))
print(b)
# {'Taobao', 'Lsgogroup', 'Google'}
c = set(["Google", "Lsgogroup", "Taobao", "Google"])
print(c)
# {'Taobao', 'Lsgogroup', 'Google'}
```
- 去掉列表中重复的元素
```
lst = [0, 1, 2, 3, 4, 5, 5, 3, 1]
temp = []
for item in lst:
if item not in temp:
temp.append(item)
print(temp) # [0, 1, 2, 3, 4, 5]
a = set(lst)
print(list(a)) # [0, 1, 2, 3, 4, 5]
```
从结果发现集合的两个特点:**无序 (unordered) 和唯一 (unique)**。
由于 set 存储的是无序集合,所以我们不可以为集合创建索引或执行切片(slice)操作,也没有键(keys)可用来获取集合中元素的值,但是可以判断一个元素是否在集合中。
## 2. 访问集合中的值
- 可以使用len()內建函数得到集合的大小。
```
s = set(['Google', 'Baidu', 'Taobao'])
print(len(s)) # 3
```
- 可以使用for把集合中的数据一个个读取出来。
```
s = set(['Google', 'Baidu', 'Taobao'])
for item in s:
print(item)
# Baidu
# Google
# Taobao
```
- 可以通过in或not in判断一个元素是否在集合中已经存在
```
s = set(['Google', 'Baidu', 'Taobao'])
print('Taobao' in s) # True
print('Facebook' not in s) # True
```
## 3. 集合的内置方法
- set.add(elmnt)用于给集合添加元素,如果添加的元素在集合中已存在,则不执行任何操作。
```
fruits = {"apple", "banana", "cherry"}
fruits.add("orange")
print(fruits)
# {'orange', 'cherry', 'banana', 'apple'}
fruits.add("apple")
print(fruits)
# {'orange', 'cherry', 'banana', 'apple'}
```
- set.update(set)用于修改当前集合,可以添加**新的元素或集合**到当前集合中,如果添加的元素在集合中已存在,则该元素只会出现一次,重复的会忽略。
```
x = {"apple", "banana", "cherry"}
y = {"google", "baidu", "apple"}
x.update(y)
print(x)
# {'cherry', 'banana', 'apple', 'google', 'baidu'}
y.update(["lsgo", "dreamtech"])
print(y)
# {'lsgo', 'baidu', 'dreamtech', 'apple', 'google'}
```
- set.remove(item) 用于移除集合中的指定元素。如果元素不存在,则会发生错误。
```
fruits = {"apple", "banana", "cherry"}
fruits.remove("banana")
print(fruits) # {'apple', 'cherry'}
```
- set.discard(value) 用于移除指定的集合元素。**remove() 方法在移除一个不存在的元素时会发生错误,而 discard() 方法不会。**
```
fruits = {"apple", "banana", "cherry"}
fruits.discard("banana")
print(fruits) # {'apple', 'cherry'}
```
- set.pop() 用于**随机**移除一个元素。和在字典中的操作一样,随机选择
```
fruits = {"apple", "banana", "cherry"}
x = fruits.pop()
print(fruits) # {'cherry', 'apple'}
print(x) # banana
```
由于 set 是无序和无重复元素的集合,所以两个或多个 set 可以做数学意义上的集合操作。
- set.intersection(set1, set2) 返回两个集合的交集。
- set1 & set2 返回两个集合的交集。
- set.intersection_update(set1, set2) 交集,在原始的集合上移除不重叠的元素。
```
a = set('abracadabra')
b = set('alacazam')
print(a) # {'r', 'a', 'c', 'b', 'd'}
print(b) # {'c', 'a', 'l', 'm', 'z'}
c = a.intersection(b)
print(c) # {'a', 'c'}
print(a & b) # {'c', 'a'}
print(a) # {'a', 'r', 'c', 'b', 'd'}
a.intersection_update(b)
print(a) # {'a', 'c'}
```
- set.union(set1, set2) 返回两个集合的并集。
- set1 | set2 返回两个集合的并集。
```
a = set('abracadabra')
b = set('alacazam')
print(a) # {'r', 'a', 'c', 'b', 'd'}
print(b) # {'c', 'a', 'l', 'm', 'z'}
print(a | b)
# {'l', 'd', 'm', 'b', 'a', 'r', 'z', 'c'}
c = a.union(b)
print(c)
# {'c', 'a', 'd', 'm', 'r', 'b', 'z', 'l'}
```
- set.difference(set) 返回集合的差集。
- set1 - set2 返回集合的差集。
- set.difference_update(set) 集合的差集,直接在原来的集合中移除元素,没有返回值。
```
a = set('abracadabra')
b = set('alacazam')
print(a) # {'r', 'a', 'c', 'b', 'd'}
print(b) # {'c', 'a', 'l', 'm', 'z'}
c = a.difference(b)
print(c) # {'b', 'd', 'r'}
print(a - b) # {'d', 'b', 'r'}
print(a) # {'r', 'd', 'c', 'a', 'b'}
a.difference_update(b)
print(a) # {'d', 'r', 'b'}
```
- set.symmetric_difference(set)返回集合的异或。
- set1 ^ set2 返回集合的异或。
- set.symmetric_difference_update(set)移除当前集合中在另外一个指定集合相同的元素,并将另外一个指定集合中不同的元素插入到当前集合中。
```
a = set('abracadabra')
b = set('alacazam')
print(a) # {'r', 'a', 'c', 'b', 'd'}
print(b) # {'c', 'a', 'l', 'm', 'z'}
c = a.symmetric_difference(b)
print(c) # {'m', 'r', 'l', 'b', 'z', 'd'}
print(a ^ b) # {'m', 'r', 'l', 'b', 'z', 'd'}
print(a) # {'r', 'd', 'c', 'a', 'b'}
a.symmetric_difference_update(b)
print(a) # {'r', 'b', 'm', 'l', 'z', 'd'}
```
- set.issubset(set)判断集合是不是被其他集合包含,如果是则返回 True,否则返回 False。
- set1 <= set2 判断集合是不是被其他集合包含,如果是则返回 True,否则返回 False。
```
x = {"a", "b", "c"}
y = {"f", "e", "d", "c", "b", "a"}
z = x.issubset(y)
print(z) # True
print(x <= y) # True
x = {"a", "b", "c"}
y = {"f", "e", "d", "c", "b"}
z = x.issubset(y)
print(z) # False
print(x <= y) # False
```
- set.issuperset(set)用于判断集合是不是包含其他集合,如果是则返回 True,否则返回 False。
- set1 >= set2 判断集合是不是包含其他集合,如果是则返回 True,否则返回 False。
```
x = {"f", "e", "d", "c", "b", "a"}
y = {"a", "b", "c"}
z = x.issuperset(y)
print(z) # True
print(x >= y) # True
x = {"f", "e", "d", "c", "b"}
y = {"a", "b", "c"}
z = x.issuperset(y)
print(z) # False
print(x >= y) # False
```
set.isdisjoint(set) 用于判断两个集合是不是不相交,如果是返回 True,否则返回 False。
```
x = {"f", "e", "d", "c", "b"}
y = {"a", "b", "c"}
z = x.isdisjoint(y)
print(z) # False
x = {"f", "e", "d", "m", "g"}
y = {"a", "b", "c"}
z = x.isdisjoint(y)
print(z) # True
```
## 4. 集合的转换
```
se = set(range(4))
li = list(se)
tu = tuple(se)
print(se, type(se)) # {0, 1, 2, 3} <class 'set'>
print(li, type(li)) # [0, 1, 2, 3] <class 'list'>
print(tu, type(tu)) # (0, 1, 2, 3) <class 'tuple'>
```
## 5. 不可变集合
Python 提供了不能改变元素的集合的实现版本,即不能增加或删除元素,类型名叫frozenset。
需要注意的是frozenset仍然可以进行集合操作,只是不能用带有update的方法。
frozenset([iterable]) 返回一个冻结的集合,冻结后集合不能再添加或删除任何元素。
```
a = frozenset(range(10)) # 生成一个新的不可变集合
print(a)
# frozenset({0, 1, 2, 3, 4, 5, 6, 7, 8, 9})
b = frozenset('lsgogroup')
print(b)
# frozenset({'g', 's', 'p', 'r', 'u', 'o', 'l'})
```
## 练习题
1. 怎么表示只包含⼀个数字1的元组。
```
tuple1 = (1,)
tuple1
```
2. 创建一个空集合,增加 {‘x’,‘y’,‘z’} 三个元素。
```
s = set()
s.add('x')
s.add('y')
s.add('z')
s
```
3. 列表['A', 'B', 'A', 'B']去重。
```
list1 = ['A', 'B', 'A', 'B']
s1= set(list1)
s1
list(s1)
```
4. 求两个集合{6, 7, 8},{7, 8, 9}中不重复的元素(差集指的是两个集合交集外的部分)。
```
a = {6,7,8}
b = {7,8,9}
list1 = []
list1.append(a.difference(b))
list1.append(b.difference(a))
list1
```
5. 求{'A', 'B', 'C'}中元素在 {'B', 'C', 'D'}中出现的次数。
```
c = {'A', 'B', 'C'}
d = {'B', 'C', 'D'}
e = c.intersection(d)
len(e)
```
| github_jupyter |
```
# What version of Python do you have?
import sys
import tensorflow.keras
import pandas as pd
import sklearn as sk
import tensorflow as tf
print(f"Tensor Flow Version: {tf.__version__}")
print(f"Keras Version: {tensorflow.keras.__version__}")
print()
print(f"Python {sys.version}")
print(f"Pandas {pd.__version__}")
print(f"Scikit-Learn {sk.__version__}")
gpu = len(tf.config.list_physical_devices('GPU'))>0
print("GPU is", "available" if gpu else "NOT AVAILABLE")
import pyarrow.feather as feather
import matplotlib.pyplot as plt
import numpy as np
# loading in the training data
path = '/Users/asgnxt/mne-miniconda/mne_data/train_val_16/train_16/eyesclosed_train.feather'
df_closed = feather.read_feather(path)
path = '/Users/asgnxt/mne-miniconda/mne_data/train_val_16/train_16/eyesopen_train.feather'
df_open = feather.read_feather(path)
path = '/Users/asgnxt/mne-miniconda/mne_data/train_val_16/train_16/mathematic_train.feather'
df_math = feather.read_feather(path)
path = '/Users/asgnxt/mne-miniconda/mne_data/train_val_16/train_16/memory_train.feather'
df_memory = feather.read_feather(path)
path = '/Users/asgnxt/mne-miniconda/mne_data/train_val_16/train_16/music_train.feather'
df_music = feather.read_feather(path)
# is data normalized?
print(df_closed.iloc[:, :-1].mean())
print(df_closed.iloc[:, :-1].std())
```
Apparently mean and sd of df_closed are 0 at this precision level!
```
# loading in the validation data
path = '/Users/asgnxt/mne-miniconda/mne_data/train_val_16/val_16/eyesclosed_val.feather'
df_closed_val = feather.read_feather(path)
path = '/Users/asgnxt/mne-miniconda/mne_data/train_val_16/val_16/eyesopen_val.feather'
df_open_val = feather.read_feather(path)
path = '/Users/asgnxt/mne-miniconda/mne_data/train_val_16/val_16/mathematic_val.feather'
df_math_val = feather.read_feather(path)
path = '/Users/asgnxt/mne-miniconda/mne_data/train_val_16/val_16/memory_val.feather'
df_memory_val = feather.read_feather(path)
path = '/Users/asgnxt/mne-miniconda/mne_data/train_val_16/val_16/music_val.feather'
df_music_val = feather.read_feather(path)
# determine the number of samples in each dataframe
print(df_closed.shape)
print(df_open.shape)
print(df_math.shape)
print(df_memory.shape)
print(df_music.shape)
print(df_closed_val.shape)
print(df_open_val.shape)
print(df_math_val.shape)
print(df_memory_val.shape)
print(df_music_val.shape)
```
The training data is either 7076 or 7259 rows x 30000 columns. Given there are 61 channels of EEG, there are 7076 / 61 = 116 / 119 distinct recordings of 300 sec each (100 Hz sampling).
Imagining an EEG 'frame' of 61 x 61 (61 channels x 610 ms); each row can be thought of as a movie with ~492 frames. Each activity has a training set of 492 x 116 or 492 x 119 frames of data from a subset of subjects and sessions
```
# defining parameters for the model
batch_size = 32
img_width = 61
img_height = 61
num_channels = 61
print(f'Number of channels: {num_channels}')
# defining the number of samples
num_samples = 30000
print(f'Number of samples: {num_samples}')
# defining the number of frames
num_frames = num_samples/num_channels
print(f'image_size = 61 x 61')
print(f'Number of images per row: {num_frames}')
# defining the number of classes
num_classes = 5
print(f'Number of classes: {num_classes}')
# defining the number of epochs
num_training_epochs = num_frames * 116 * num_classes
print(f'num_training_epochs = {num_training_epochs.__round__()}')
# create labels for each dataframe with float16 precision
df_closed['label'] = 0
df_open['label'] = 1
df_math['label'] = 2
df_memory['label'] = 3
df_music['label'] = 4
df_closed_val['label'] = 0
df_open_val['label'] = 1
df_math_val['label'] = 2
df_memory_val['label'] = 3
df_music_val['label'] = 4
# force the labels to be float16 precision
df_closed['label'] = df_closed['label'].astype('float16')
df_open['label'] = df_open['label'].astype('float16')
df_math['label'] = df_math['label'].astype('float16')
df_memory['label'] = df_memory['label'].astype('float16')
df_music['label'] = df_music['label'].astype('float16')
df_closed_val['label'] = df_closed_val['label'].astype('float16')
df_open_val['label'] = df_open_val['label'].astype('float16')
df_math_val['label'] = df_math_val['label'].astype('float16')
df_memory_val['label'] = df_memory_val['label'].astype('float16')
df_music_val['label'] = df_music_val['label'].astype('float16')
# ensure that the dataframes are correctly labeled
df_music.head()
# Creating lists from each dataframe, each list contains one frame of data
list_df_closed = np.array_split(df_closed, 116)
print(list_df_closed[0].shape)
list_df_open = np.array_split(df_open, 116)
print(list_df_open[0].shape)
list_df_math = np.array_split(df_math, 119)
print(list_df_math[0].shape)
list_df_memory = np.array_split(df_memory, 119)
print(list_df_memory[0].shape)
list_df_music = np.array_split(df_music, 116)
print(list_df_music[0].shape)
list_df_closed_val = np.array_split(df_closed_val, 14)
print(list_df_closed_val[0].shape)
list_df_open_val = np.array_split(df_open_val, 14)
print(list_df_open_val[0].shape)
list_df_math_val = np.array_split(df_math_val, 14)
print(list_df_math_val[0].shape)
list_df_memory_val = np.array_split(df_memory_val, 14)
print(list_df_memory_val[0].shape)
list_df_music_val = np.array_split(df_music_val, 15)
print(list_df_music_val[0].shape)
# Create a training dataset with multiple sessions / subjects
training_examples = []
for i in range(116):
training_examples.append(list_df_closed[i])
training_examples.append(list_df_open[i])
training_examples.append(list_df_math[i])
training_examples.append(list_df_memory[i])
training_examples.append(list_df_music[i])
# Create a validation dataset with multiple sessions / subjects
validation_examples = []
for i in range(14):
validation_examples.append(list_df_closed_val[i])
validation_examples.append(list_df_open_val[i])
validation_examples.append(list_df_math_val[i])
validation_examples.append(list_df_memory_val[i])
validation_examples.append(list_df_music_val[i])
# defining parameters for the model
batch_size = 32
img_width = 61
img_height = 61
target_size = (img_width, img_height)
num_channels = 61
print(f'Number of channels: {num_channels}')
# defining the number of samples
num_samples = 30000
print(f'Number of samples: {num_samples}')
# defining the number of frames
num_frames = num_samples/num_channels
print(f'image_size = 61 x 61')
print(f'Number of images per row: {num_frames}')
# defining the number of classes
num_classes = 5
print(f'Number of classes: {num_classes}')
# defining the number of epochs
num_training_epochs = num_frames * 116 * num_classes
print(f'num_training_epochs = {num_training_epochs.__round__()}')
# creating a single training dataframe
training_examples = pd.concat(training_examples)
print(training_examples.shape)
# creating a single validation dataframe
validation_examples = pd.concat(validation_examples)
print(validation_examples.shape)
# ensuring uniform dtype
training_examples.dtypes
# create a separate target dataframe
target = training_examples.pop('label')
print(target.shape)
print(target.head())
dataset_ts = tf.convert_to_tensor(training_examples)
# dataset_ts_batches = dataset_ts.shuffle(buffer_size=10000).batch_size=batch_size #not working
# Create a model
model = tf.keras.Sequential([
tf.keras.layers.Flatten(input_shape=(30000, 1)),
tf.keras.layers.Dense(128, activation=tf.nn.relu),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(10, activation=tf.nn.softmax)
])
# Compile the model
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
# Train the model
model.fit(dataset_ts, target, epochs=3)
```
| github_jupyter |
# Wave Equation
#### John S Butler john.s.butler@tudublin.ie [Course Notes](https://johnsbutler.netlify.com/files/Teaching/Numerical_Analysis_for_Differential_Equations.pdf) [Github](https://github.com/john-s-butler-dit/Numerical-Analysis-Python)
## Overview
This notebook will implement the Forward Euler in time and Centered in space method to appoximate the solution of the wave equation.
## The Differential Equation
Condsider the one-dimensional hyperbolic Wave Equation:
$$ \frac{\partial u}{\partial t} +a\frac{\partial u}{\partial x}=0,$$
where $a=1$, with the initial conditions
$$ u(x,0)=1-\cos(x), \ \ 0 \leq x \leq 2\pi. $$
with wrap around boundary conditions.
```
# LIBRARY
# vector manipulation
import numpy as np
# math functions
import math
# THIS IS FOR PLOTTING
%matplotlib inline
import matplotlib.pyplot as plt # side-stepping mpl backend
import warnings
warnings.filterwarnings("ignore")
```
## Discete Grid
The region $\Omega$ is discretised into a uniform mesh $\Omega_h$. In the space $x$ direction into $N$ steps giving a stepsize of
$$ \Delta_x=\frac{2\pi-0}{N},$$
resulting in
$$x[j]=0+j\Delta_x, \ \ \ j=0,1,...,N,$$
and into $N_t$ steps in the time $t$ direction giving a stepsize of
$$ \Delta_t=\frac{1-0}{N_t}$$
resulting in
$$t[n]=0+n\Delta_t, \ \ \ n=0,...,K.$$
The Figure below shows the discrete grid points for $N=10$ and $Nt=100$, the known boundary conditions (green), initial conditions (blue) and the unknown values (red) of the Heat Equation.
```
N=20
Nt=100
h=2*np.pi/N
k=1/Nt
time_steps=100
time=np.arange(0,(time_steps+.5)*k,k)
x=np.arange(0,2*np.pi+h/2,h)
X, Y = np.meshgrid(x, time)
fig = plt.figure()
plt.plot(X,Y,'ro');
plt.plot(x,0*x,'bo',label='Initial Condition');
plt.xlim((-h,2*np.pi+h))
plt.ylim((-k,max(time)+k))
plt.xlabel('x')
plt.ylabel('time (ms)')
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.title(r'Discrete Grid $\Omega_h$ ',fontsize=24,y=1.08)
plt.show();
```
## Initial Conditions
The discrete initial conditions is,
$$ w[0,j]=1-\cos(x[j]), \ \ 0 \leq x[j] \leq \pi, $$
The figure below plots values of $w[j,0]$ for the inital (blue) conditions for $t[0]=0.$
```
w=np.zeros((time_steps+1,N+1))
b=np.zeros(N-1)
# Initial Condition
for j in range (0,N+1):
w[0,j]=1-np.cos(x[j])
fig = plt.figure(figsize=(12,4))
plt.plot(x,w[0,:],'o:',label='Initial Condition')
plt.xlim([-0.1,max(x)+h])
plt.title('Intitial Condition',fontsize=24)
plt.xlabel('x')
plt.ylabel('w')
plt.legend(loc='best')
plt.show()
```
## Boundary Conditions
To account for the wrap-around boundary conditions
$$w_{-1,n}=w_{N,n},$$
and
$$w_{N+1,n}=w_{0,n}.$$
```
xpos = np.zeros(N+1)
xneg = np.zeros(N+1)
for j in range(0,N+1):
xpos[j] = j+1
xneg[j] = j-1
xpos[N] = 0
xneg[0] = N
```
## The Explicit Forward Time Centered Space Difference Equation
The explicit Forward Time Centered Space difference equation of the Wave Equation is,
$$
\frac{w^{n+1}_{j}-w^{n}_{j}}{\Delta_t}+\big(\frac{w^n_{j+1}-w^n_{j-1}}{2\Delta_x}\big)=0.
$$
Rearranging the equation we get,
$$
w_{j}^{n+1}=w^{n}_{j}-\lambda a(w_{j+1}^{n}-w_{j-1}^{n}),
$$
for $j=0,...10$ where $\lambda=\frac{\Delta_t}{\Delta_x}$.
This gives the formula for the unknown term $w^{n+1}_{j}$ at the $(j,n+1)$ mesh points
in terms of $x[j]$ along the nth time row.
```
lamba=k/h
for n in range(0,time_steps):
for j in range (0,N+1):
w[n+1,j]=w[n,j]-lamba/2*(w[n,int(xpos[j])]-w[n,int(xneg[j])])
```
## Results
```
fig = plt.figure(figsize=(12,6))
plt.subplot(121)
for n in range (1,time_steps+1):
plt.plot(x,w[n,:],'o:')
plt.xlabel('x[j]')
plt.ylabel('w[j,n]')
plt.subplot(122)
X, T = np.meshgrid(x, time)
z_min, z_max = np.abs(w).min(), np.abs(w).max()
plt.pcolormesh( X,T, w, vmin=z_min, vmax=z_max)
#plt.xticks(np.arange(len(x[0:N:2])), x[0:N:2])
#plt.yticks(np.arange(len(time)), time)
plt.xlabel('x[j]')
plt.ylabel('time, t[n]')
clb=plt.colorbar()
clb.set_label('Temperature (w)')
#plt.colorbar()
plt.suptitle('Numerical Solution of the Wave Equation',fontsize=24,y=1.08)
fig.tight_layout()
plt.show()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/sevgiE/HU-BBY162-2022/blob/main/Python_101-2022.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
#Bölüm 00: Python'a Giriş
## Yazar Hakkında
SEVGİ EĞRİ
```
```
##Çalışma Defteri Hakkında
Bu çalışma defteri Google'ın Jupyter Notebook platformuna benzer özellikler taşıyan Google Colab üzerinde oluşturulmuştur. Google Colab, herhangi bir altyapı düzenlemesine ihtiyaç duymadan Web tabanlı olarak Python kodları yazmanıza ve çalıştırmanıza imkan veren ücretsiz bir platformdur. Platform ile ilgili detaylı bilgiye [https://colab.research.google.com/notebooks/intro.ipynb](https://colab.research.google.com/notebooks/intro.ipynb) adresinden ulaşabilirsiniz.
Python'a giriş seviyesinde 10 dersten oluşan bu çalışma defteri daha önce kodlama deneyimi olmayan öğrenenler için hazırlanmıştır. Etkileşimli yapısından dolayı hem konu anlatımlarının hem de çalıştırılabilir örneklerin bir arada olduğu bu yapı, sürekli olarak güncellenebilecek bir altyapıya sahiptir. Bu açıdan çalışma defterinin güncel sürümünü aşağıdaki adresten kontrol etmenizi tavsiye ederim.
Sürüm 1.0: [Python 101](https://github.com/orcunmadran/Python101/blob/main/Python_101.ipynb)
İyi çalışmalar ve başarılar :)
## Kullanım Şartları
Bu çalışma defteri aşağıda belirtilen şartlar altında, katkıda bulunanlara Atıf vermek ve aynı lisansla paylaşmak kaydıyla ticari amaç dahil olmak üzere her şekilde dağıtabilir, paylaşabilir, üzerinde değişiklik yapılarak yeniden kullanılabilir.
---

Bu çalışma defteri Jetbrains'in "Introduction to Python" dersi temel alınarak hazırlanmış ve Creative Commons [Atıf-AynıLisanslaPaylaş 4.0 Uluslararası Lisansı](http://creativecommons.org/licenses/by-sa/4.0/) ile lisanslanmıştır.
---
# Bölüm 01: Giriş
Bu bölümde:
* İlk bilgisayar programımız,
* Yorumlar yer almaktadır.
## İlk Bilgisayar Programımız
Geleneksel olarak herhangi bir programlama dilinde yazılan ilk program "Merhaba Dünya!"'dır.
**Örnek Uygulama:**
```
print("Merhaba Dünya!")
```
```
# Örnek uygulamayı çalıştır
print("Merhaba Dünya!")
```
**Görev:** Kendinizi dünyaya tanıtacak ilk bilgisayar programını yazın!
```
print("Merhaba ben Sinem!")
```
## Yorumlar
Python'daki yorumlar # "hash" karakteriyle başlar ve fiziksel çizginin sonuna kadar uzanır. Yorum yapmak için kullanılan # "hash" karakteri kod satırlarını geçici olarak devre dışı bırakmak amacıyla da kullanılabilir.
**Örnek Uygulama:**
```
# Bu ilk bilgisayar programım için ilk yorumum
print("# bu bir yorum değildir")
print("Merhaba!") # yorumlar kod satırının devamında da yapılabilir.
#print("Bu kod geçici olarak devre dışı bırakılmıştır.")
```
```
# Örnek uygulamayı çalıştır
# Bu ilk bilgisayar programım için ilk yorumum
print("# bu bir yorum değildir")
print("Merhaba!") # yorumlar kod satırının devamında da yapılabilir.
# print("Bu kod geçici olarak devre dışı bırakılmıştır.")
```
**Görev:** Python kodunuza yeni bir yorum ekleyin, mevcut satıra yorum ekleyin, yazılmış olan bir kod satırını geçici olarak devre dışı bırakın!
```
print("Bu satıbrın devamına bir yorum ekleyin")
print("Bu satırı devre dışı bırakın!")
print("bugün hava çok güzel")
print("#yağmurlu değil!")
```
# Bölüm 02: Değişkenler
Bu bölümde:
* Değişken nedir?,
* Değişken tanımlama,
* Değişken türleri,
* Değişken türü dönüştürme,
* Aritmetik operatörler,
* Artıtılmış atama operatörleri,
* Boolean operatörleri,
* Karşılaştırma operatörleri yer almaktadır.
## Değişken Nedir?
Değişkenler değerleri depolamak için kullanılır. Böylece daha sonra bu değişkenler program içinden çağırılarak atanan değer tekrar ve tekrar kullanılabilir. Değişkenlere metinler ve / veya sayılar atanabilir. Sayı atamaları direkt rakamların yazılması ile gerçekleştirilirken, metin atamalarında metin tek tırnak içinde ( 'abc' ) ya da çift tırnak ( "abc" ) içinde atanır.
Değişkenler etiketlere benzer ve atama operatörü olarak adlandırılan eşittir ( = ) operatörü ile bir değişkene bir değer atanabilir. Bir değer ataması zincirleme şeklinde gerçekleştirilebilir. Örneğin: a = b = 2
**Örnek Uygulama 1**
Aşağıda bir "zincir atama" örneği yer almaktadır. Değer olarak atanan 2 hem "a" değişkenine, hem de "b" değişkenine atanmaktadır.
```
a = b = 2
print("a = " + str(a))
print("b = " + str(b))
```
"a" ve "b" değişkenleri başka metinler ile birlikte ekrana yazdırılmak istendiğinde metin formatına çevrilmesi gerekmektedir. Bu bağlamda kullanılan "str(a)" ve "str(b)" ifadeleri eğitimin ilerleyen bölümlerinde anlatılacaktır.
```
# Örnek uygulamayı çalıştır
a = b = 2
print("a = " + str(a))
print("b = " + str(b))
```
**Örnek Uygulama 2**
```
adSoyad = "Orçun Madran"
print("Adı Soyadı: " + adSoyad)
```
```
# Örnek uygulamayı çalıştır
adSoyad = "Orçun Madran"
print("Adı Soyadı: " + adSoyad)
```
**Görev:** "eposta" adlı bir değişken oluşturun. Oluşturduğunuz bu değişkene bir e-posta adresi atayın. Daha sonra atadığınız bu değeri ekrana yazdırın. Örneğin: "E-posta: orcun[at]madran.net"
```
print ("eposta = sinemmandacı")
print("eposta")
```
## Değişken Tanımlama
Değişken isimlerinde uyulması gereken bir takım kurallar vardır:
* Rakam ile başlayamaz.
* Boşluk kullanılamaz.
* Alt tire ( _ ) haricinde bir noktalama işareti kullanılamaz.
* Python içinde yerleşik olarak tanımlanmış anahtar kelimeler kullanılamaz (ör: print).
* Python 3. sürümden itibaren latin dışı karakter desteği olan "Unicode" desteği gelmiştir. Türkçe karakterler değişken isimlerinde kullanılabilir.
**Dikkat:** Değişken isimleri büyük-küçük harfe duyarlıdır. Büyük harfle başlanan isimlendirmeler genelde *sınıflar* için kullanılır. Değişken isimlerinin daha anlaşılır olması için deve notasyonu (camelCase) ya da alt tire kullanımı tavsiye edilir.
**Örnek Uygulama:**
```
degisken = 1
kullaniciAdi = "orcunmadran"
kul_ad = "rafet"
```
Henüz tanımlanmamış bir değişken kullanıldığında derleyicinin döndürdüğü hatayı kodu çalıştırarak gözlemleyin!
```
degisken1 = "Veri"
print(degisken2)
```
**Görev:** Tanımladığınız değişkeni ekrana yazdırın!
```
degisken3 = 'Yeni veri' "merhaba ben sinem "
print(değişken4) #hatalı bölüm burada!
değişken5 = "merhaba bugün hava çok güzel"
print(değişken5)
```
## Değişken Türleri
Python'da iki ana sayı türü vardır; tam sayılar ve ondalık sayılar.
**Dikkat:** Ondalık sayıların yazımında Türkçe'de *virgül* (,) kullanılmasına rağmen, programlama dillerinin evrensel yazım kuralları içerisinde ondalık sayılar *nokta* (.) ile ifade edilir.
**Örnek Uygulama:**
```
tamSayi = 5
print(type(tamSayi)) # tamSayi değişkeninin türünü yazdırır
ondalikSayi = 7.4
print(type(ondalikSayi) # ondalikSayi değişkeninin türünü yazdırır
```
```
# Örnek uygulamayı çalıştır
tamSayi = 5
print(type(tamSayi))
ondalikSayi = 7.4
print(type(ondalikSayi)
```
**Görev:** "sayi" değişkeninin türünü belirleyerek ekrana yazdırın!
```
Sayi = 9.0
tamSayi = 9.0
print(float(tamSayi))
```
## Değişken Türü Dönüştürme
Bir veri türünü diğerine dönüştürmenize izin veren birkaç yerleşik fonksiyon (built-in function) vardır. Bu fonksiyonlar ("int()", "str()", "float()") uygulandıkları değişkeni dönüştürerek yeni bir nesne döndürürler.
**Örnek Uygulama**
```
sayi = 6.5
print(type(sayi)) # "sayi" değişkeninin türünü ondalık olarak yazdırır
print(sayi)
sayi = int(sayi) # Ondalık sayı olan "sayi" değişkenini tam sayıya dönüştürür
print(type(sayi))
print(sayi)
sayi = float(sayi) # Tam sayı olan "sayi" değişkenini ondalık sayıya dönüştürür
print(type(sayi))
print(sayi)
sayi = str(sayi) # "sayi" değişkeni artık düz metin halini almıştır
print(type(sayi))
print(sayi)
```
```
# Örnek uygulamayı çalıştır
sayi = 6.5
print(type(sayi))
print(sayi)
sayi = int(sayi)
print(type(sayi))
print(sayi)
sayi = float(sayi)
print(type(sayi))
print(sayi)
sayi = str(sayi)
print(type(sayi))
print(sayi)
```
**Görev:** Ondalık sayıyı tam sayıya dönüştürün ve ekrana değişken türünü ve değeri yazdırın!
```
sayi = 3.14
print(int(sayi))
```
## Aritmetik Operatörler
Diğer tüm programlama dillerinde olduğu gibi, toplama (+), çıkarma (-), çarpma (yıldız) ve bölme (/) operatörleri sayılarla kullanılabilir. Bunlarla birlikte Python'un üs (çift yıldız) ve mod (%) operatörleri vardır.
**Dikkat:** Matematik işlemlerinde geçerli olan aritmetik operatörlerin öncelik sıralamaları (çarpma, bölme, toplama, çıkarma) ve parantezlerin önceliği kuralları Python içindeki matematiksel işlemler için de geçerlidir.
**Örnek Uygulama:**
```
# Toplama işlemi
sayi = 7.0
sonuc = sayi + 3.5
print(sonuc)
# Çıkarma işlemi
sayi = 200
sonuc = sayi - 35
print(sonuc)
# Çarpma işlemi
sayi = 44
sonuc = sayi * 10
print(sonuc)
# Bölme işlemi
sayi = 30
sonuc = sayi / 3
print(sonuc)
# Üs alma işlemi
sayi = 30
sonuc = sayi ** 3
print(sonuc)
# Mod alma işlemi
sayi = 35
sonuc = sayi % 4
print(sonuc)
```
```
# Örnek uygulamayı çalıştır
# Toplama işlemi
sayi = 7.0
sonuc = sayi + 3.5
print(sonuc)
# Çıkarma işlemi
sayi = 200
sonuc = sayi - 35
print(sonuc)
# Çarpma işlemi
sayi = 44
sonuc = sayi * 10
print(sonuc)
# Bölme işlemi
sayi = 30
sonuc = sayi / 3
print(sonuc)
# Üs alma işlemi
sayi = 30
sonuc = sayi ** 3
print(sonuc)
# Mod alma işlemi
sayi = 35
sonuc = sayi % 4
print(sonuc)
```
**Görev:** Aşağıda değer atamaları tamamlanmış olan değişkenleri kullanarak ürünlerin peşin satın alınma bedelini TL olarak hesaplayınız ve ürün adı ile birlikte ekrana yazdırınız! İpucu: Ürün adını ve ürün bedelini tek bir satırda yazdırmak isterseniz ürün bedelini str() fonksiyonu ile düz metin değişken türüne çevirmeniz gerekir.
```
urunAdi = "Bisiklet"
urunBedeliAvro = 850
pariteAvroTL = 7
urunAdet = 3
pesinAdetIndirimTL = 500
butce =15000
hesapla = ((urunBedeliAvro + urunAdet) * pariteAvroTL) - pesinAdetIndirimTL
butceTamam = butce > hesapla
print(hesapla)
print("Alışveriş bütçeme uygun mu?" + str(butceTamam))
```
## Artırılmış Atama Operatörleri
Artırılmış atama, bir değişkenin mevcut değerine belirlenen değerin eklenerek ( += ) ya da çıkartılarak ( -= ) atanması işlemidir.
**Örnek Uygulama**
```
sayi = 8
sayi += 4 # Mevcut değer olan 8'e 4 daha ekler.
print(sayi)
sayi -= 6 # Mevcut değer olan 12'den 6 eksiltir.
print("Sayı = " + str(sayi))
```
```
# Örnek uygulama çalıştır
sayi = 8
sayi += 4
print(sayi)
sayi -= 6
print("Sayı = " + str(sayi))
```
**Görev:** Artıtılmış atama operatörleri kullanarak "sayi" değişkenine 20 ekleyip, 10 çıkartarak değişkenin güncel değerini ekrana yazdırın!
```
sayi = 55
sayi += 20
print(sayi)
sayi -= 10
print("Sayı = " + str(sayi))
```
## Boolean Operatörleri
Boolean, yalnızca **Doğru (True)** veya **Yanlış (False)** olabilen bir değer türüdür. Eşitlik (==) operatörleri karşılaştırılan iki değişkenin eşit olup olmadığını kontrol eder ve *True* ya da *False* değeri döndürür.
**Örnek Uygulama:**
```
deger1 = 10
deger2 = 10
esitMi = (deger1 == deger2) # Eşit olup olmadıkları kontrol ediliyor
print(esitMi) # Değişken "True" olarak dönüyor
deger1 = "Python"
deger2 = "Piton"
esitMi = (deger1 == deger2) # Eşit olup olmadıkları kontrol ediliyor
print(esitMi) # Değişken "False" olarak dönüyor
```
```
# Örnek uygulama çalıştır
deger1 = 10
deger2 = 10
esitMi = (deger1 == deger2)
print(esitMi)
deger1 = "Python"
deger2 = "Piton"
esitMi = (deger1 == deger2)
print(esitMi)
```
**Görev:** Atamaları yapılmış olan değişkenler arasındaki eşitliği kontrol edin ve sonucu ekrana yazıdırın!
```
sifre = "Python2020"
sifreTekrar = "Piton2020"
esitMi = (sifre == sifreTekrar)
print(esitMi)
```
## Karşılaştırma Operatörleri
Python'da, >=, <= , >, < vb. dahil olmak üzere birçok operatör bulunmaktadır. Python'daki tüm karşılaştırma operatörleri aynı önceliğe sahiptir. Karşılaştırma sonucunda boole değerleri (*True* ya da *False*) döner. Karşılaştırma operatörleri isteğe bağlı olarak arka arkaya da (zincirlenerek) kullanılabilir.
**Örnek Uygulama:**
```
deger1 = 5
deger2 = 7
deger3 = 9
print(deger1 < deger2 < deger3) # Sonuç "True" olarak dönecektir
```
```
# Örnek uygulama çalıştır
deger1 = 5
deger2 = 7
deger3 = 9
print(deger1 < deger2 < deger3)
```
**Görev:** Aşağıda değer atamaları tamamlanmış olan değişkenleri kullanarak ürünlerin peşin satın alınma bedelini TL olarak hesaplayın. Toplam satın alma bedeli ile bütçenizi karşılaştırın. Satın alma bedelini ve bütçenizi ekrana yazdırın. Ödeme bütçenizi aşıyorsa ekrana "False", aşmıyorsa "True" yazdırın.
```
urunAdi = "Bisiklet"
urunBedeliAvro = 850
kurAvro = 7
urunAdet = 3
pesinAdetIndirimTL = 500
butce = 15000
hesapla = ((urunBedeliAvro + urunAdet) * pariteAvroTL - pesinAdetIndirimTL)
butceTamam = butce > hesapla
print(hesapla)
print("Alışveriş bütçeme uygun mu?" + str(butceTamam))
```
# Bölüm 03: Metin Katarları
Bu bölümde:
* Birbirine bağlama,
* Metin katarı çarpımı,
* Metin katarı dizinleme,
* Metin katarı negatif dizinleme,
* Metin katarı dilimleme,
* In operatörü,
* Metin katarının uzunluğu,
* Özel karakterlerden kaçma,
* Basit metin katarı metodları,
* Metin katarı biçimlendirme yer almaktadır.
## Birbirine Bağlama
Birbirine bağlama artı (+) işlemini kullanarak iki metin katarının birleştirilmesi işlemine denir.
**Örnek Uygulama**
```
deger1 = "Merhaba"
deger2 = "Dünya"
selamlama = deger1 + " " + deger2
print(selamlama) # Çıktı: Merhaba Dünya
```
```
# Örnek uygulamayı çalışıtır
deger1 = "Merhaba"
deger2 = "Dünya"
selamlama = deger1 + " " + deger2
print(selamlama)
```
**Görev:** *ad*, *soyad* ve *hitap* değişkenlerini tek bir çıktıda birleştirecek kodu yazın!
```
hitap = "Öğr. Gör."
ad = "Orçun"
soyad = "Madran"
hitap = ad + " " + soyad
print(hitap)
# Çıktı: Öğr. Gör. Orçun Madran
```
## Metin Katarı Çarpımı
Python, metin katarlarının çarpım sayısı kadar tekrar ettirilmesini desteklemektedir.
**Örnek Uygulama**
```
metin = "Hadi! "
metniCarp = metin * 4
print(metniCarp) # Çıktı: Hadi! Hadi! Hadi! Hadi!
```
```
# Örnek uygulamayı çalıştır
metin = "Hadi!"
metniCarp = metin * 4
print(metniCarp)
```
**Görev:** Sizi sürekli bekleten arkadaşınızı uyarabilmek için istediğiniz sayıda "Hadi!" kelimesini ekrana yazdırın!
```
metin = "Hadi! "
metniCarp = metin * 15
print(metniCarp)
# Çıktı: Hadi! Hadi! Hadi! Hadi! ... Hadi!
metin = "hadi! \n"
metniCarp = metin * 15
print(metniCarp)
```
##Metin Katarı Dizinleme
Konumu biliniyorsa, bir metin katarındaki ilgili karaktere erişilebilir. Örneğin; str[index] metin katarındaki indeks numarasının karşılık geldiği karakteri geri döndürecektir. İndekslerin her zaman 0'dan başladığı unutulmamalıdır. İndeksler, sağdan saymaya başlamak için negatif sayılar da olabilir. -0, 0 ile aynı olduğundan, negatif indeksler -1 ile başlar.
**Örnek Uygulama**
```
metin = "Python Programlama Dili"
print("'h' harfini yakala: " + metin[3]) # Çıktı: 'h' harfini yakala: h"
```
```
# örnek uygulama çalıştır
metin = "Python Programlama Dili"
print("'h' harfini yakala: " + metin[3])
```
**Görev:** İndeks numarasını kullanarak metin katarındaki ikinci "P" harfini ekrana yazdırın!
```
metin = "Python Programlama Dili"
print("'P' harfini yakala:" + metin[7])
# Çıktı: P
```
## Metin Katarı Negatif Dizinleme
Metin katarının sonlarında yer alan bir karaktere daha rahat erişebilmek için indeks numarası negatif bir değer olarak belirlenebilir.
**Örnek Uygulama**
```
metin = "Python Programlama Dili"
dHarfi = metin[-4]
print(dHarfi) # Çıktı: D
```
```
# Örnek uygulama çalıştır
metin = "Python Programlama Dili"
dHarfi = metin[-4]
print(dHarfi)
```
**Görev:** Metin katarının sonunda yer alan "i" harfini ekrana yazdırın!
```
metin = "Python Programlama Dili"
iHarfi = metin[-1]
print(iHarfi)
#Çıktı: i
```
##Metin Katarı Dilimleme
Dilimleme, bir metin katarından birden çok karakter (bir alt katar oluşturmak) almak için kullanılır. Söz dizimi indeks numarası ile bir karaktere erişmeye benzer, ancak iki nokta üst üste işaretiyle ayrılmış iki indeks numarası kullanılır. Ör: str[ind1:ind2].
Noktalı virgülün solundaki indeks numarası belirlenmezse ilk karakterden itibaren (ilk karakter dahil) seçimin yapılacağı anlamına gelir. Ör: str[:ind2]
Noktalı virgülün sağındaki indeks numarası belirlenmezse son karaktere kadar (son karakter dahil) seçimin yapılacağı anlamına gelir. Ör: str[ind1:]
**Örnek Uygulama**
```
metin = "Python Programlama Dili"
dilimle = metin[:6]
print(dilimle) # Çıktı: Python
metin = "Python Programlama Dili"
print(metin[7:]) # Çıktı: Programlama Dili
```
```
# Örnek uygulama çalıştır
metin = "Python Programlama Dili"
dilimle = metin[:6]
print(dilimle)
metin = "Python Programlama Dili"
print(metin[7:])
```
**Görev:** Metin katarını dilemleyerek katarda yer alan üç kelimeyi de ayrı ayrı (alt alta) ekrana yazdırın!.
```
metin = "Python Programlama Dili"
dilimle = metin[:6]
print(dilimle)
metin = "Pyhton Proglamlama Dili"
dilimle2 = metin[6:18]
print(dilimle2)
metin = "Pyhton Proglamlama Dili"
dilimle3 = metin[18:23]
print(dilimle3)
# Çıktı:
# Python
# Programlama
# Dili
```
##In Operatörü
Bir metin katarının belirli bir harf ya da bir alt katar içerip içermediğini kontrol etmek için, in anahtar sözcüğü kullanılır.
**Örnek Uygulama**
```
metin = "Python Programlama Dili"
print("Programlama" in metin) # Çıktı: True
```
**Görev:** Metin katarında "Python" kelimesinin geçip geçmediğini kontrol ederek ekrana yazdırın!
```
metin = "Python Programlama Dili"
print("Python")
metin = "Python Proglamlama Dili"
print("lama")
```
##Metin Katarının Uzunluğu
Bir metin katarının kaç karakter içerdiğini saymak için len() yerleşik fonksiyonu kullanılır.
**Örnek Uygulama**
```
metin = "Python programlama dili"
print(len(metin)) # Çıktı: 23
```
```
# Örnek uygulamayı çalıştır
metin = "Python programlama dili"
print(len(metin))
```
**Görev:** Metin katarındaki cümlenin ilk yarısını ekrana yazdırın! Yazılan kod cümlenin uzunluğundan bağımsız olarak cümleyi ikiye bölmelidir.
```
metin = "Python programlama dili, dünyada eğitim amacıyla en çok kullanılan programlama dillerinin başında gelir."
# Çıktı: Python programlama dili, dünyada eğitim amacıyla en
print(len(metin))
```
## Özel Karakterlerden Kaçma
Metin katarları içerisinde tek ve çift tırnak kullanımı kimi zaman sorunlara yol açmaktadır. Bu karakterin metin katarları içerisinde kullanılabilmesi için "Ters Eğik Çizgi" ile birlikte kullanılırlar.
Örneğin: 'Önümüzdeki ay "Ankara'da Python Eğitimi" gerçekleştirilecek' cümlesindeki tek tırnak kullanımı soruna yol açacağından 'Önümüzdeki ay "Ankara\'da Python Eğitimi" gerçekleştirilecek' şeklinde kullanılmalıdır.
**İpucu:** Tek tırnaklı metin katarlarından kaçmak için çift tırnak ya da tam tersi kullanılabilir.
**Örnek Uygulama**
```
metin = 'Önümüzdeki ay "Ankara\'da Python Eğitimi" gerçekleştirilecektir.'
print(metin) #Çıktı: Önümüzdeki ay "Ankara'da Python Eğitimi" gerçekleştirilecektir.
metin = 'Önümüzdeki ay "Ankara'da Python Eğitimi" gerçekleştirilecektir.'
print(metin) # Çıktı: Geçersiz söz dizimi hatası dönecektir.
```
```
# Örnek uygulamayı çalıştır
metin = 'Önümüzdeki ay "Ankara\'da Python Eğitimi" gerçekleştirilecektir.'
print(metin)
# Örnek uygulamadaki hatayı gözlemle
metin = 'Önümüzdeki ay "Ankara'da Python Eğitimi" gerçekleştirilecektir.'
print(metin)
```
**Görev:** Metin katarındaki cümlede yer alan noktalama işaretlerinden uygun şekilde kaçarak cümleyi ekrana yazdırın!
```
metin = 'Bilimsel çalışmalarda "Python" kullanımı Türkiye\'de çok yaygınlaştı!'
print(metin)
```
##Basit Metin Katarı Metodları
Python içinde birçok yerleşik metin katarı fonksiyonu vardır. En çok kullanılan fonksiyonlardan bazıları olarak;
* tüm harfleri büyük harfe dönüştüren *upper()*,
* tüm harfleri küçük harfe dönüştüren *lower()*,
* sadece cümlenin ilk harfini büyük hale getiren *capitalize()* sayılabilir.
**İpucu:** Python'daki yerleşik fonksiyonların bir listesini görüntüleyebilmek için metin katarından sonra bir nokta (.) koyulur ve uygun olan fonksiyonlar arayüz tarafından otomatik olarak listelenir. Bu yardımcı işlevi tetiklemek için CTRL + Bolşuk tuş kombinasyonu da kullanılabilir.
**Örnek Uygulama**
```
metin = "Python Programlama Dili"
print(metin.lower()) # Çıktı: python programlama dili
print(metin.upper()) # Çıktı: PYTHON PROGRAMLAMA DILI
print(metin.capitalize()) # Çıktı: Python programlama dili
```
```
# Örnek uygulamayı çalıştır
metin = "Python Programlama Dili"
print(metin.lower())
print(metin.upper())
print(metin.capitalize())
```
**Görev:** *anahtarKelime* ve *arananKelime* değişkenlerinde yer alan metinler karşılaştırıldığında birbirlerine eşit (==) olmalarını sağlayın ve dönen değerin "True" olmasını sağlayın!
```
anahtarKelime = "Makine Öğrenmesi"
arananKelime = "makine öğrenmesi"
print(anahtarKelime.lower())
print(anahtarKelime == arananKelime)
# Çıktı: True
```
##Metin Katarı Biçimlendirme
Bir metin katarından sonraki % operatörü, bir metin katarını değişkenlerle birleştirmek için kullanılır. % operatörü, bir metin katarıdanki % s öğesini, arkasından gelen değişkenle değiştirir. % d sembolü ise, sayısal veya ondalık değerler için yer tutucu olarak kullanılır.
**Örnek Uygulama**
```
adsoyad = "Orçun Madran"
dogumTarihi = 1976
print("Merhaba, ben %s!" % adsoyad) # Çıktı: Merhaba, ben Orçun Madran!
print("Ben %d doğumluyum" % dogumTarihi) # Ben 1976 doğumluyum.
ad = "Orçun"
soyad = "Madran"
print("Merhaba, ben %s %s!" % (ad, soyad)) # Çıktı: Merhaba, ben Orçun Madran!
```
```
# Örnek uygulamayı çalıştır
adsoyad = "Orçun Madran"
dogumTarihi = 1976
print("Merhaba, ben %s!" % adsoyad)
print("Ben %d doğumluyum" % dogumTarihi)
# Örnek uygulamayı çalıştır
ad = "Orçun"
soyad = "Madran"
print("Merhaba, ben %s %s!" % (ad, soyad))
```
**Görev:** "Merhaba Orçun Madran, bu dönemki dersiniz 'Programlama Dilleri'. Başarılar!" cümlesini ekrana biçimlendirmeyi kullanarak (artı işaretini kullanmadan) yazdırın!
```
ad = "Orçun"
soyad = "Madran"
ders = "Programlama Dilleri"
print("Merhaba ben %s %s, bu dönemki dersiniz %s. Başarılar!" % (ad, soyad, ders))
# Çıktı: Merhaba Orçun Madran, bu dönemki dersiniz "Programlama Dilleri". Başarılar!
```
# Bölüm 04: Veri Yapılar
Bu bölümde:
* Listeler,
* Liste işlemleri,
* Liste öğeleri,
* Demetler (Tuples),
* Sözlükler,
* Sözlük değerleri ve anahtarları,
* In anahtar kelimesinin kullanımı yer almaktadır.
## Listeler
Liste, birden fazla değeri tek bir değişken adı altında saklamak için kullanabileceğiniz bir veri yapısıdır. Bir liste köşeli parantez arasında virgülle ayrılmış değerler dizisi olarak yazılır. Ör: liste = [deger1, deger2].
Listeler farklı türden öğeler içerebilir, ancak genellikle listedeki tüm öğeler aynı türdedir. Metin katarları gibi listeler de dizine eklenebilir ve dilimlenebilir. (Bkz. Bölüm 3).
**Örnek Uygulama**
```
acikListe = ["Açık Bilim", "Açık Erişim", "Açık Veri", "Açık Eğitim", "Açık Kaynak"] # acikListe adında yeni bir liste oluşturur
print(acikListe) # Çıktı: ['Açık Bilim', 'Açık Erişim', 'Açık Veri', 'Açık Eğitim', 'Açık Kaynak']
```
```
# Örnek uygulamayı çalıştır
acikListe = ["Açık Bilim", "Açık Erişim", "Açık Veri", "Açık Eğitim", "Açık Kaynak"]
print(acikListe)
```
**Görev 1:** acikListe içinde yer alan 3. liste öğesini ekrana yazıdırın!
```
acikListe = ["Açık Bilim", "Açık Erişim", "Açık Veri", "Açık Eğitim", "Açık Kaynak"]
print(acikListe[2])
```
**Görev 2:** acikListe içinde yer alan 4. ve 5. liste öğesini ekrana yazıdırın!
```
acikListe = ["Açık Bilim", "Açık Erişim", "Açık Veri", "Açık Eğitim", "Açık Kaynak"]
print(acikListe[3:5])
```
## Liste İşlemleri
append() fonksiyonunu kullanarak ya da artırılmış atama operatörü ( += ) yardımıyla listenin sonuna yeni öğeler (değerler) eklenebilir. Listelerin içindeki öğeler güncellenebilir, yani liste[indeksNo] = yeni_deger kullanarak içeriklerini değiştirmek mümkündür.
**Örnek Uygulama**
```
acikListe = ["Açık Bilim", "Açık Erişim", "Açık Veri", "Açık Eğitim", "Açık Kaynak"] # acikListe adında yeni bir liste oluşturur
print(acikListe) # Çıktı: ['Açık Bilim', 'Açık Erişim', 'Açık Veri', 'Açık Eğitim', 'Açık Kaynak']
acikListe += ["Açık Donanım", "Açık İnovasyon"] # listeye iki yeni öğe ekler
print(acikListe) # Çıktı: ['Açık Bilim', 'Açık Erişim', 'Açık Veri', 'Açık Eğitim', 'Açık Kaynak', 'Açık Donanım', 'Açık İnovasyon']
acikListe.append("Açık Veri Gazeteciliği") # listeye yeni bir öğe ekler
print(acikListe) # Çıktı: ['Açık Bilim', 'Açık Erişim', 'Açık Veri', 'Açık Eğitim', 'Açık Kaynak', 'Açık Donanım', 'Açık İnovasyon', 'Açık Veri Gazeteciliği']
acikListe[4] = "Açık Kaynak Kod" # listenin 5. öğesini değiştirir
print(acikListe) # Çıktı: ['Açık Bilim', 'Açık Erişim', 'Açık Veri', 'Açık Eğitim', 'Açık Kaynak Kod', 'Açık Donanım', 'Açık İnovasyon', 'Açık Veri Gazeteciliği']
```
```
# Örnek uygulamayı çalıştır
acikListe = ["Açık Bilim", "Açık Erişim", "Açık Veri", "Açık Eğitim", "Açık Kaynak"]
print(acikListe)
acikListe += ["Açık Donanım", "Açık İnovasyon"]
print(acikListe)
acikListe.append("Açık Veri Gazeteciliği")
print(acikListe)
acikListe[4] = "Açık Kaynak Kod"
print(acikListe)
```
**Görev:** bilgiBilim adlı bir liste oluşturun. Bu listeye bilgi bilim disiplini ile ilgili 3 adet anahtar kelime ya da kavram ekleyin. Bu listeyi ekrana yazdırın. Listeye istediğiniz bir yöntem ile (append(), +=) 2 yeni öğe ekleyin. Ekrana listenin son durumunu yazdırın. Listenizdeki son öğeyi değiştirin. Listenin son halini ekrana yazıdırn.
```
#bilgiBilim
bilgiBilim = ["bilgi", "veri", "bilim"]
print(bilgiBilim)
bilgiBilim.append("bby")
print(bilgiBilim)
```
## Liste Öğeleri
Liste öğelerini dilimleme (slice) yaparak da atamak mümkündür. Bu bir listenin boyutunu değiştirebilir veya listeyi tamamen temizleyebilir.
**Örnek Uygulama**
```
acikListe = ["Açık Bilim", "Açık Erişim", "Açık Veri", "Açık Eğitim", "Açık Kaynak"] # acikListe adında yeni bir liste oluşturur
print(acikListe) # Çıktı: ['Açık Bilim', 'Açık Erişim', 'Açık Veri', 'Açık Eğitim', 'Açık Kaynak']
acikListe[2:4] = ["Açık İnovasyon"] # "Açık Veri" ve "Açık Eğitim" öğelerinin yerine tek bir öğe ekler
print(acikListe) #Çıktı: ["Açık Bilim", "Açık Erişim", "Açık İnovasyon", "Açık Kaynak"]
acikListe[:2] = [] # listenin ilk iki öğesini siler
print(acikListe) #Çıktı: ["Açık İnovasyon", "Açık Kaynak"]
acikListe[:] = [] # listeyi temizler
print(acikListe) #Çıktı: []
```
```
# Örnek uygulamayı çalıştır
acikListe = ["Açık Bilim", "Açık Erişim", "Açık Veri", "Açık Eğitim", "Açık Kaynak"]
print(acikListe)
acikListe[2:4] = ["Açık İnovasyon"]
print(acikListe)
acikListe[:2] = []
print(acikListe)
acikListe[:] = []
print(acikListe)
```
**Görev:** Önceki görevde oluşturulan "bilgiBilim" adlı listenin istediğiniz öğesini silerek listenin güncel halini ekrana yazdırın. Listeyi tamamen temizleyerek listenin güncel halini ekrana yazdırın.
```
#bilgiBilim
bilgiBilim = ["bilgi","veri","bilim","bby"]
print(bilgiBilim)
bilgiBilim[2:3] = ["bilimsellik"]
print(bilgiBilim)
bilgiBilim[:] = []
print(bilgiBilim)
```
## Demetler (Tuples)
Demetler neredeyse listelerle aynı. Demetler ve listeler arasındaki tek önemli fark, demetlerin değiştirilememesidir. Demetlere öğe eklenmez, öğe değiştirilmez veya demetlerden öğe silinemez. Demetler, parantez içine alınmış bir virgül operatörü tarafından oluşturulur. Ör: demet = ("deger1", "deger2", "deger3"). Tek bir öğe demetinde ("d",) gibi bir virgül olmalıdır.
**Örnek Uygulama**
```
ulkeKodlari = ("TR", "US", "EN", "JP")
print(ulkeKodlari) # Çıktı: ('TR', 'US', 'EN', 'JP')
```
```
# Örnek uygulamayı çalıştır
ulkeKodlari = ("TR", "US", "EN", "JP")
print(ulkeKodlari)
```
**Görev:** Kongre Kütüphanesi konu başlıkları listesinin kodlarından oluşan bir demet oluşturun ve ekrana yazdırın! Oluşturulan demet içindeki tek bir öğeyi ekrana yazdırın!
```
#konuBasliklari
kongreKkonuB = ("felsefe", "din", "tarih", "siyaset", "hukuk", "eğitim")
print(kongreKkonuB)
print(kongreKkonuB[3])
```
## Sözlükler
Sözlük, listeye benzer, ancak sözlük içindeki değerlere indeks numarası yerine bir anahtara ile erişilebilir. Bir anahtar herhangi bir metin katarı veya rakam olabilir. Sözlükler ayraç içine alınır. Ör: sozluk = {'anahtar1': "değer1", 'anahtar2': "değer2"}.
**Örnek Uygulama**
```
adresDefteri = {"Hacettepe Üniversitesi": "hacettepe.edu.tr", "ODTÜ": "odtu.edu.tr", "Bilkent Üniversitesi": "bilkent.edu.tr"} # yeni bir sözlük oluşturur
print(adresDefteri) # Çıktı: {'Hacettepe Üniversitesi': 'hacettepe.edu.tr', 'ODTÜ': 'odtu.edu.tr', 'Bilkent Üniversitesi': 'bilkent.edu.tr'}
adresDefteri["Ankara Üniversitesi"] = "ankara.edu.tr" #sözlüğe yeni bir öğe ekler
print(adresDefteri) # Çıktı: {'Hacettepe Üniversitesi': 'hacettepe.edu.tr', 'ODTÜ': 'odtu.edu.tr', 'Bilkent Üniversitesi': 'bilkent.edu.tr', 'Ankara Üniversitesi': 'ankara.edu.tr'}
del adresDefteri ["Ankara Üniversitesi"] #sözlükten belirtilen öğeyi siler
print(adresDefteri) # Çıktı: {'Hacettepe Üniversitesi': 'hacettepe.edu.tr', 'ODTÜ': 'odtu.edu.tr', 'Bilkent Üniversitesi': 'bilkent.edu.tr'}
```
```
# Örnek uygulamayı çalıştır
adresDefteri = {"Hacettepe Üniversitesi": "hacettepe.edu.tr", "ODTÜ": "odtu.edu.tr", "Bilkent Üniversitesi": "bilkent.edu.tr"}
print(adresDefteri)
adresDefteri["Ankara Üniversitesi"] = "ankara.edu.tr"
print(adresDefteri)
del adresDefteri ["Ankara Üniversitesi"]
print(adresDefteri)
```
**Görev:** İstediğin herhangi bir konuda 5 öğeye sahip bir sözlük oluştur. Sözlüğü ekrana yazdır. Sözlükteki belirli bir öğeyi ekrana yazdır. Sözlükteki belirli bir öğeyi silerek sözlüğün güncel halini ekrana yazdır!
```
#sozluk
sozluk = {"one":"bir", "two":"iki", "three":"üç", "four":"dört", "five":"beş"}
print(sozluk)
del sozluk ["four"]
print(sozluk)
```
## Sözlük Değerleri ve Anahtarları
Sözlüklerde values() ve keys() gibi birçok yararlı fonksiyon vardır. Bir sozlük adı ve ardından noktadan sonra çıkan listeyi kullanarak geri kalan fonksiyolar incelenebilir.
**Örnek Uygulama**
```
adresDefteri = {"Hacettepe Üniversitesi": "hacettepe.edu.tr", "ODTÜ": "odtu.edu.tr", "Bilkent Üniversitesi": "bilkent.edu.tr"} # yeni bir sözlük oluşturur
print(adresDefteri) # Çıktı: {'Hacettepe Üniversitesi': 'hacettepe.edu.tr', 'ODTÜ': 'odtu.edu.tr', 'Bilkent Üniversitesi': 'bilkent.edu.tr'}
print(adresDefteri.values()) # Çıktı: dict_values(['hacettepe.edu.tr', 'odtu.edu.tr', 'bilkent.edu.tr'])
print(adresDefteri.keys()) # Çıktı: dict_keys(['Hacettepe Üniversitesi', 'ODTÜ', 'Bilkent Üniversitesi'])
```
```
# Örnek uygulamayı çalıştır
adresDefteri = {"Hacettepe Üniversitesi": "hacettepe.edu.tr", "ODTÜ": "odtu.edu.tr", "Bilkent Üniversitesi": "bilkent.edu.tr"}
print(adresDefteri)
print(adresDefteri.values())
print(adresDefteri.keys())
```
**Görev:** İstediğin bir konuda istediğin öğe saysına sahip bir sözlük oluştur. Sözlükler ile ilgili farklı fonksiyoları dene. Sonuçları ekrana yazdır!
```
#yeniSozluk
sozluk = {"yeni":"new", "küçük":"small", "büyük":"big"}
print(sozluk)
print(sozluk.values())
print(sozluk.keys())
```
##In Anahtar Kelimesi
"In" anahtar sözcüğü, bir listenin veya sözlüğün belirli bir öğe içerip içermediğini kontrol etmek için kullanılır. Daha önce metin katarlarındaki kullanıma benzer bir kullanımı vardır. "In" anahtar sözcüğü ile öğe kontrolü yapıldıktan sonra sonuç, öğe listede ya da sözlükte yer alıyorsa *True* yer almıyorsa *False* olarak geri döner.
**Dikkat**: Aranan öğe ile liste ya da sözlük içinde yer alan öğelerin karşılaştırılması sırasında büyük-küçük harf duyarlılığı bulunmaktadır. Ör: "Bilgi" ve "bilgi" iki farklı öğe olarak değerlendirilir.
**Örnek Uygulama**
```
bilgiKavramları = ["indeks", "erişim", "koleksiyon"] # yeni bir liste oluşturur
print("Erişim" in bilgiKavramları) # Çıktı: False
bilgiSozlugu = {"indeks": "index", "erişim": "access", "koleksiyon": "collection"} # yeni bir sozluk oluşturur
print("koleksiyon" in bilgiSozlugu.keys()) # çıktı: True
```
```
# Örnek uygulamayı çalıştır
bilgiKavramları = ["indeks", "erişim", "koleksiyon"]
print("Erişim" in bilgiKavramları)
bilgiSozlugu = {"indeks": "index", "erişim": "access", "koleksiyon": "collection"}
print("koleksiyon" in bilgiSozlugu.keys())
```
**Görev:** Bir liste ve bir sözlük oluşturun. Liste içinde istediğiniz kelimeyi aratın ve sonucunu ekrana yazdırın! Oluşturduğunuz sözlüğün içinde hem anahtar kelime (keys()) hem de değer (values()) kontrolü yaptırın ve sonucunu ekrana yazdırın!
```
#yeniListe
#yeniSozluk
liste = ["new","small","big"]
print("love" in liste)
sozluk = {"yeni":"new", "küçük":"small", "büyük":"big"}
print("küçük" in sozluk.keys())
```
#Bölüm 05: Koşullu İfadeler
Bu bölümde:
* Mantıksal operatörler,
* If cümleciği,
* Else ve elif kullanımı yer almatadır.
##Mantıksal Operatörler
Mantıksal operatörler ifadeleri karşılaştırır ve sonuçları *True* ya da *False* değerleriyle döndürür. Python'da üç tane mantıksal operatör bulunur:
1. "and" operatörü: Her iki yanındaki ifadeler doğru olduğunda *True* değerini döndürür.
2. "or" operatörü: Her iki tarafındaki ifadelerden en az bir ifade doğru olduğunda "True" değerini döndürür.
3. "not" operatörü: İfadenin tam tersi olarak değerlendirilmesini sağlar.
**Örnek Uygulama**
```
kullaniciAdi = "orcunmadran"
sifre = 123456
print(kullaniciAdi == "orcunmadran" and sifre == 123456) # Çıktı: True
kullaniciAdi = "orcunmadran"
sifre = 123456
print(kullaniciAdi == "orcunmadran" and not sifre == 123456) # Çıktı: False
cepTel = "05321234567"
ePosta = "orcunmadran@gmail.com"
print(cepTel == "" or ePosta == "orcunmadran@gmail.com" ) # Çıktı: True
```
```
# Örnek uygulamayı çalıştır
kullaniciAdi = "orcunmadran"
sifre = 123456
print(kullaniciAdi == "orcunmadran" and sifre == 123456)
kullaniciAdi = "orcunmadran"
sifre = 123456
print(kullaniciAdi == "orcunmadran" and not sifre == 123456)
cepTel = "05321234567"
ePosta = "orcunmadran@gmail.com"
print(cepTel == "" or ePosta == "orcunmadran@gmail.com" )
```
**Görev:** Klavyeden girilen kullanıcı adı ve şifrenin kayıtlı bulunan kullanıcı adı ve şifre ile uyuşup uyuşmadığını kontrol edin ve sonucu ekrana yazdırın!
```
#Sistemde yer alan bilgiler:
sisKulAdi = "yonetici"
sisKulSifre = "bby162"
#Klavyeden girilen bilgiler:
girKulAdi = input("Kullanıcı Adı: ")
girKulSifre = input("Şifre: ")
#kontrol
sonuc = sisKulAdi == girKulAdi and sisKulSifre == sisKulSifre
#sonuc
print(sonuc)
```
## If Cümleciği
"If" anahtar sözcüğü, verilen ifadenin doğru olup olmadığını kontrol ettikten sonra belirtilen kodu çalıştıran bir koşullu ifade oluşturmak için kullanılır. Python'da kod bloklarının tanımlanması için girinti kullanır.
**Örnek Uygulama**
```
acikKavramlar = ["bilim", "erişim", "veri", "eğitim"]
kavram = input("Bir açık kavramı yazın: ")
if kavram in acikKavramlar:
print(kavram + " açık kavramlar listesinde yer alıyor!")
```
```
# Örnek uygulamayı çalıştır
acikKavramlar = ["bilim", "erişim", "veri", "eğitim"]
kavram = input("Bir açık kavramı yazın: ")
if kavram in acikKavramlar:
print(kavram + " açık kavramlar listesinde yer alıyor!")
```
**Görev:** "acikSozluk" içinde yer alan anahtarları (keys) kullanarak eğer klavyeden girilen anahtar kelime sözlükte varsa açıklamasını ekrana yazdırın!
```
acikSozluk = {
"Açık Bilim" : "Bilimsel bilgi kamu malıdır. Bilimsel yayınlara ve verilere açık erişim bir haktır." ,
"Açık Erişim" : "Kamu kaynakları ile yapılan araştırmalar sonucunda üretilen yayınlara ücretsiz erişim" ,
"Açık Veri" : "Kamu kaynakları ile yapılan araştırma sonucunda üretilen verilere ücretsiz ve yeniden kullanılabilir biçimde erişim"
}
anahtar = input("Anahtar Kelime: ")
print(acikSozluk.keys())
#If
```
## Else ve Elif Kullanımı
"If" cümleciği içinde ikinci bir ifadenin doğruluğunun kontrolü için "Elif" ifadesi kullanılır. Doğruluğu sorgulanan ifadelerden hiçbiri *True* döndürmediği zaman çalışacak olan kod bloğu "Else" altında yer alan kod bloğudur.
**Örnek Uygulama**
```
gunler = ["Pazartesi", "Çarşamba", "Cuma"]
girilen = input("Gün giriniz: ")
if girilen == gunler[0]:
print("Programlama Dilleri")
elif girilen == gunler[1]:
print("Kataloglama")
elif girilen == gunler[2]:
print("Bilimsel İletişim")
else :
print("Kayıtlı bir gün bilgisi girmediniz!")
```
```
# Örnek uygulamayı çalıştır
gunler = ["Pazartesi", "Çarşamba", "Cuma"]
girilen = input("Gün giriniz: ")
if girilen == gunler[0]:
print("Programlama Dilleri")
elif girilen == gunler[1]:
print("Kataloglama")
elif girilen == gunler[2]:
print("Bilimsel İletişim")
else :
print("Kayıtlı bir gün bilgisi girmediniz!")
```
**Görev:** Klavyeden girilen yaş bilgisini kullanarak ekrana aşağıdaki mesajları yazdır:
* 21 yaş altı ve 64 yaş üstü kişilere: "Sokağa çıkma yasağı bulunmaktadır!"
* Diğer tüm kişilere: "Sokağa çıkma yasağı yoktur!"
* Klavyeden yaş harici bir bilgi girişi yapıldığında: "Yaşınızı rakam olarak giriniz!"
```
yas = int(input("Yaşınızı giriniz: "))
```
# Bölüm 06: Döngüler
Bu bölümde:
* for döngüsü,
* Metin katarlarında for döngüsü kullanımı,
* while döngüsü,
* break anahtar kelimesi,
* continue anahtar kelimesi yer almaktadır.
## for Döngüsü
for döngüleri belirli komut satırını ya da satırlarını yinelemek (tekrar etmek) için kullanılır. Her yinelemede, for döngüsünde tanımlanan değişken listedeki bir sonraki değere otomatik olarak atanacaktır.
**Örnek Uygulama**
```
for i in range(5): # i değerine 0-4 arası indeks değerleri otomatik olarak atanır
print(i) # Çıktı: Bu komut satırı toplam 5 kere tekrarlanır ve her satırda yeni i değeri yazdırılır
konular = ["Açık Bilim", "Açık Erişim", "Açık Veri"] # yeni bir liste oluşturur
for konu in konular:
print(konu) #Çıktı: Her bir liste öğesi alt alta satırlara yazdırılır
```
```
# Örnek uygulmayı çalıştır
for i in range(5):
print(i)
# Örnek uygulmayı çalıştır
konular = ["Açık Bilim", "Açık Erişim", "Açık Veri"]
for konu in konular:
print(konu)
```
**Görev:** Bir liste oluşturun. Liste öğelerini "for" döngüsü kullanarak ekrana yazdırın!
```
#liste
çiçekListe = ["gül","papatya","lale","yasemin","menekşe","lavanta"]
for liste in çiçekListe:
print(liste)
```
## Metin Katarlarında for Döngüsü Kullanımı
Metin Katarları üzerinde gerçekleştirilebilecek işlemler Python'daki listelerle büyük benzerlik taşırlar. Metin Katarını oluşturan öğeler (harfler) liste elemanları gibi "for" döngüsü yardımıyla ekrana yazdırılabilir.
**Örnek Uygulama**
```
cumle = "Bisiklet hem zihni hem bedeni dinç tutar!"
for harf in cumle: # Cümledeki her bir harfi ekrana satır satır yazdırır
print(harf)
```
```
# Örnek uygulamayı çalıştır
cumle = "Bisiklet hem zihni, hem bedeni dinç tutar!"
for harf in cumle:
print(harf)
```
**Görev:** İçinde metin katarı bulunan bir değişken oluşturun. Bu değişkende yer alan her bir harfi bir satıra gelecek şekilde "for" döngüsü ile ekrana yazdırın!
```
#degisken
değişken2 = "merhaba , bugün hava karlı ve soğuk"
for harf in değişken2:
print(harf)
```
## while Döngüsü
"While" döngüsü "if" cümleciğinin ifade şekline benzer. Koşul doğruysa döngüye bağlı kod satırı ya da satırları yürütülür (çalıştırılır). Temel fark, koşul doğru (True) olduğu olduğu sürece bağlı kod satırı ya da satırları çalışmaya devam eder.
**Örnek Uygulama**
```
deger = 1
while deger <= 10:
print(deger) # Bu satır 10 kez tekrarlanacak
deger += 1 # Bu satır da 10 kez tekrarlanacak
print("Program bitti") # Bu satır sadece bir kez çalıştırılacak
```
```
# Örnek uygulamayı çalıştır
deger = 1
while deger <= 10:
print(deger)
deger += 1
print("Program bitti")
deger2 = 8
while deger2 <= 25:
print(deger2)
deger2 += 3
print("Tebrikler, tamamladınız!")
```
## break Anahtar Kelimesi
Asla bitmeyen döngüye sonsuz döngü adı verilir. Döngü koşulu daima doğru (True) olursa, böyle bir döngü sonsuz olur. "Break" anahtar kelimesi geçerli döngüden çıkmak için kullanılır.
**Örnek Uygulama**
```
sayi = 0
while True: # bu döngü sonsuz bir döngüdür
print(sayi)
sayi += 1
if sayi >= 5:
break # sayı değeri 5 olduğunda döngü otomatik olarak sonlanır
```
```
# Örnek Uygulamayı çalıştır
sayi = 0
while True:
print(sayi)
sayi += 1
if sayi >= 5:
break
rakam = 5
while True:
print(rakam)
rakam += 2
if rakam >= 15:
break
```
## continue Anahtar Kelimesi
"continue" anahtar kelimesi, o anda yürütülen döngü için döngü içindeki kodun geri kalanını atlamak ve "for" veya "while" deyimine geri dönmek için kullanılır.
```
for i in range(5):
if i == 3:
continue # i değeri 3 olduğu anda altta yer alan "print" komutu atlanıyor.
print(i)
```
```
# Örnek Uygulamayı çalıştır
for i in range(5):
if i == 3:
continue
print(i)
for i in range(9):
if i == 5:
continue
print(i)
```
**Görev: Tahmin Oyunu**
"while" döngüsü kullanarak bir tahmin oyunu tasarla. Bu tahmin oyununda, önceden belirlenmiş olan kelime ile klavyeden girilen kelime karşılaştırılmalı, tahmin doğru ise oyun "Bildiniz..!" mesajı ile sonlanmalı, yanlış ise tahmin hakkı bir daha verilmeli.
```
#Tahmin Oyunu
aylar = ["Ocak","Şubat","Mart","Nisan","Mayıs","Haziran","Temmuz","Ağustos","Eylül","Ekim","Kasım","Aralık"]
for ay in aylar:
print(ay)
if ay == "Nisan":
continue
print("Gratis'te indirim var!")
```
# Bölüm 07: Fonksiyonlar
## Fonksiyon Tanımlama (Definition)
Fonksiyonlar, yazılan kodu faydalı bloklara bölmenin, daha okunabilir hale getirmenin ve tekrar kullanmaya yardımcı olmanın kullanışlı bir yoludur. Fonksiyonlar "def" anahtar sözcüğü ve ardından fonksiyonun adı kullanılarak tanımlanır.
**Örnek Uygulama**
```
def merhaba_dunya(): # fonksiyon tanımlama, isimlendirme
print("Merhaba Dünya!") #fonksiyona dahil kod satırları
for i in range(5):
merhaba_dunya() # fonksiyon 5 kere çağırılacak
```
```
# Örnek uygulamayı çalıştır
def merhaba_dunya(): # fonksiyon tanımlama, isimlendirme
print("Merhaba Dünya!") #fonksiyona dahil kod satırları
for i in range(5):
merhaba_dunya() # fonksiyon 5 kere çağırılacak
```
##Fonksiyolarda Parametre Kullanımı
Fonksiyon parametreleri, fonksiyon adından sonra parantez () içinde tanımlanır. Parametre, iletilen bağımsız değişken için değişken adı görevi görür.
**Örnek Uygulama**
```
def foo(x): # x bir fonksiyon parametresidir
print("x = " + str(x))
foo(5) # 5 değeri fonksiyona iletilir ve değer olarak kullanılır.
```
```
# Örnek uygulamayı çalıştır
def foo(x):
print("x = " + str(x))
foo(5)
```
**Görev:** *karsila* fonksiyonunun tetiklenmesi için gerekli kod ve parametleri ekle!
```
#def karsila(kAd, kSoyad):
#print("Hoşgeldin, %s %s" % (kAd, kSoyad))
def karsila(ad,soyad):
print("***...***")
print("HOŞ GELDİNİZ! " + ad +" "+ soyad)
print("***...***")
karsila("Sinem","Mandacı")
```
##Return Değeri
Fonksiyonlar, "return" anahtar sözcüğünü kullanarak fonksiyon sonucunda bir değer döndürebilir. Döndürülen değer bir değişkene atanabilir veya sadece örneğin değeri yazdırmak için kullanılabilir.
**Örnek Uygulama**
```
def iki_sayi_topla(a, b):
return a + b # hesaplama işleminin sonucu değer olarak döndürülüyor
print(iki_sayi_topla(3, 12)) # ekrana işlem sonucu yazdırılacak
```
```
# Örnek uygulamayı çalıştır
def iki_sayi_topla(a, b):
return a + b
print(iki_sayi_topla(3, 12))
```
##Varsayılan Parametreler
Bazen bir veya daha fazla fonksiyon parametresi için varsayılan bir değer belirtmek yararlı olabilir. Bu, ihtiyaç duyulan parametrelerden daha az argümanla çağrılabilen bir fonksiyon oluşturur.
**Örnek Uygulama**
```
def iki_sayi_carp(a, b=2):
return a * b
print(iki_sayi_carp(3, 47)) # verilen iki degeri de kullanır
print(iki_sayi_carp(3)) # verilmeyen 2. değer yerine varsayılanı kullanır
```
```
# Örnek uygulamayı çalıştır
def iki_sayi_carp(a, b=2):
return a * b
print(iki_sayi_carp(3, 47))
print(iki_sayi_carp(3))
```
**Örnek Uygulama: Sayısal Loto**
Aşağıda temel yapısı aynı olan iki *sayısal loto* uygulaması bulunmaktadır: Fonksiyonsuz ve fonksiyonlu.
İlk sayısal loto uygulamasında herhangi bir fonksiyon kullanımı yoktur. Her satırda 1-49 arası 6 adet sayının yer aldığı 6 satır oluşturur.
İkinci sayısal loto uygulamsında ise *tahminEt* isimli bir fonksiyon yer almaktadır. Bu fonksiyon varsayılan parametrelere sahiptir ve bu parametreler fonksiyon çağırılırken değiştirilebilir. Böylece ilk uygulamadan çok daha geniş seçenekler sunabilir bir hale gelmiştir.
```
#Sayısal Loto örnek uygulama (fonksiyonsuz)
from random import randint
i = 0
secilenler = [0,0,0,0,0,0]
for rastgele in secilenler:
while i < len(secilenler):
secilen = randint(1, 49)
if secilen not in secilenler:
secilenler[i] = secilen
i+=1
print(sorted(secilenler))
i=0
#Sayısal Loto örnek uygulama (fonksiyonlu)
from random import randint
def tahminEt(rakam=6, satir=6, baslangic=1, bitis=49):
i = 0
secilenler = []
for liste in range(rakam):
secilenler.append(0)
for olustur in range(satir):
while i < len(secilenler):
secilen = randint(baslangic, bitis)
if secilen not in secilenler:
secilenler[i] = secilen
i+=1
print(sorted(secilenler))
i=0
tahminEt(10,6,1,60)
```
**Görev:** Bu görev genel olarak fonksiyon bölümünü kapsamaktadır.
Daha önce yapmış olduğunuz "Adam Asmaca" projesini (ya da aşağıda yer alan örneği) fonksiyonlar kullanarak oyun bittiğinde tekrar başlatmaya gerek duyulmadan yeniden oynanabilmesine imkan sağlayacak şekilde yeniden kurgulayın.
Oyunun farklı sekansları için farklı fonksiyonlar tanımlayarak oyunu daha optimize hale getirmeye çalışın.
Aşağıda bir adam asmaca oyununun temel özellikerine sahip bir örnek yer almaktadır.
```
#Fonksiyonsuz Adam Asmaca
from random import choice
adamCan = 3
kelimeler = ["bisiklet", "triatlon", "yüzme", "koşu"]
secilenKelime = choice(kelimeler)
print(secilenKelime)
dizilenKelime = []
for diz in secilenKelime:
dizilenKelime.append("_")
print(dizilenKelime)
while adamCan > 0:
girilenHarf = input("Bir harf giriniz: ")
canKontrol = girilenHarf in secilenKelime
if canKontrol == False:
adamCan-=1
i = 0
for kontrol in secilenKelime:
if secilenKelime[i] == girilenHarf:
dizilenKelime[i] = girilenHarf
i+=1
print(dizilenKelime)
print("Kalan can: "+ str(adamCan))
#fonksiyonlu adam asmaca
from random import choice
def oyunFon():
adamCan = 3
kelimeler = ["tez","yüksek lisans","doktora","makale","bilimsel kaynak"]
secilenKelime = []
dizilenKelime = []
butunKelimelerVar = False
sayac = 0
tekrarOyna = True
print("-------------------------\n\n\n\t\tADAM ASMACA \n\n\nLütfen zorluk derecesi giriniz (kolay, orta, zor): ")
x = input()
print (x)
if x == 'kolay':
for kelime in kelimeler:
if len(kelime) <= 5:
secilenKelime.append(kelime)
elif x == 'orta':
for kelime in kelimeler:
if len(kelime) > 5 and len(kelime)<= 7:
secilenKelime.append(kelime)
elif x == 'zor':
for kelime in kelimeler:
if len(kelime) > 7:
secilenKelime.append(kelime)
secim = choice(secilenKelime)
for diz in secim:
dizilenKelime.append("_")
print(dizilenKelime)
while adamCan >0 or tekrarOyna == True :
girilenHarf = input("Lütfen bir harf giriniz: ")
canKontrol = girilenHarf in secim
if canKontrol == False:
adamCan -=1
elif sayac+1 == len(secim):
print("\nTebrikler! Seçilen kelime: " + secim)
y = input("Tekrar oynamak ister misiniz?(evet, hayır):")
if y == 'hayır':
tekrarOyna == False
break
elif y == 'evet':
oyunFon()
else:
i = 0
for kontrol in secim:
if secim[i] == girilenHarf:
dizilenKelime[i] = girilenHarf
sayac +=1
i+=1
print(dizilenKelime)
print("\n")
print('Kalan can: ' + str(adamCan) + '\n')
if adamCan == 0:
print("Üzgünüm,Kaybettiniz")
oyunDurum = True
while oyunDurum == True:
oyunFon()
oyunDurum = False
```
# Bölüm 08: Sınıflar ve Nesneler
Bu bölümde:
* Sınıf ve nesne tanımlama,
* Değişkenlere erişim,
* self parametresi,
* init metodu yer almaktadır.
## Sınıf ve Nesne Tanımlama
Bir nesne değişkenleri ve fonksiyonları tek bir varlıkta birleştirir. Nesneler değişkenlerini ve fonksiyonlarını sınıflardan alır. Sınıflar bir anlamda nesnelerinizi oluşturmak için kullanılan şablonlardır. Bir nesneyi, fonksiyonların yanı sıra veri içeren tek bir veri yapısı olarak düşünebilirsiniz. Nesnelerin fonksiyonlarına yöntem (metod) denir.
**İpucu:** Sınıf isimlerinin baş harfi büyük yazılarak Python içindeki diğer öğelerden (değişken, fonksiyon vb.) daha rahat ayırt edilmeleri sağlanır.
**Örnek Uygulama**
```
class BenimSinifim: # yeni bir sınıfın tanımlanması
bsDegisken = 4 # sınıf içinde yer alan bir değişken
def bsFonksiyon(self): #sınıf içinde yer alan bir fonksiyon
print("Benim sınıfımın fonksiyonundan Merhaba!")
benimNesnem = BenimSinifim()
```
##Değişkenlere ve Fonksiyonlara Erişim
Sınıftan örneklenen bir nesnenin içindeki bir değişkene ya da fonksiyona erişmek için öncelikle nesnenin adı daha sonra ise değişkenin ya da fonkiyonun adı çağırılmalıdır (Ör: nesneAdi.degiskenAdi). Bir sınıfın farklı örnekleri (nesneleri) içinde tanımlanan değişkenlerin değerleri değiştirebilir.
**Örnek Uygulama 1**
```
class BenimSinifim: # yeni bir sınıf oluşturur
bsDegisken = 3 # sınıfın içinde bir değişken tanımlar
def bsFonksiyon(self): #sınıfın içinde bir fonksiyon tanımlar
print("Benim sınıfımın fonksiyonundan Merhaba!")
benimNesnem = BenimSinifim() #sınıftan yeni bir nesne oluşturur
for i in range(benimNesnem.bsDegisken): # oluşturulan nesne üzerinden değişkene ve fonksiyona ulaşılır
benimNesnem.bsFonksiyon()
benimNesnem.bsDegisken = 5 # sınıfın içinde tanımlanan değişkene yeni değer atanması
for i in range(benimNesnem.bsDegisken):
benimNesnem.bsFonksiyon()
```
```
# Örnek uygulama 1'i gözlemleyelim
class BenimSinifim:
bsDegisken = 3
def bsFonksiyon(self):
print("Benim sınıfımın fonksiyonundan Merhaba!")
benimNesnem = BenimSinifim()
for i in range(benimNesnem.bsDegisken):
benimNesnem.bsFonksiyon()
benimNesnem.bsDegisken = 5
for i in range(benimNesnem.bsDegisken):
benimNesnem.bsFonksiyon()
```
**Örnek Uygulama 2**
```
class Bisiklet:
renk = "Kırmızı"
vites = 1
def ozellikler(self):
ozellikDetay = "Bu bisiklet %s renkli ve %d viteslidir." % (self.renk, self.vites)
return ozellikDetay
bisiklet1 = Bisiklet()
bisiklet2 = Bisiklet()
print("Bisiklet 1: " + bisiklet1.ozellikler())
bisiklet2.renk = "Sarı"
bisiklet2.vites = 22
print("Bisiklet 2: " + bisiklet2.ozellikler())
```
```
# Örnek uygulama 2'i gözlemleyelim
class Bisiklet:
renk = "Kırmızı"
vites = 1
def ozellikler(self):
ozellikDetay = "Bu bisiklet %s renkli ve %d viteslidir." % (self.renk, self.vites)
return ozellikDetay
bisiklet1 = Bisiklet()
bisiklet2 = Bisiklet()
print("Bisiklet 1: " + bisiklet1.ozellikler())
bisiklet2.renk = "Sarı"
bisiklet2.vites = 22
print("Bisiklet 2: " + bisiklet2.ozellikler())
```
##self Parametresi
"self" parametresi bir Python kuralıdır. "self", herhangi bir sınıf yöntemine iletilen ilk parametredir. Python, oluşturulan nesneyi belirtmek için self parametresini kullanır.
**Örnek Uygulama**
Aşağıdaki örnek uygulamada **Bisiklet** sınıfının değişkenleri olan *renk* ve *bisiklet*, sınıf içindeki fonksiyonda **self** parametresi ile birlikte kullanılmaktadır. Bu kullanım şekli sınıftan oluşturulan nesnelerin tanımlanmış değişkenlere ulaşabilmeleri için gereklidir.
```
class Bisiklet:
renk = "Kırmızı"
vites = 1
def ozellikler(self):
ozellikDetay = "Bu bisiklet %s renkli ve %d viteslidir." % (self.renk, self.vites)
return ozellikDetay
```
```
# Örnek uygulamada "self" tanımlaması yapılmadığı zaman döndürülen hata kodunu inceleyin
class Bisiklet:
renk = "Kırmızı"
vites = 1
def ozellikler(self):
ozellikDetay = "Bu bisiklet %s renkli ve %d viteslidir." % (renk, vites) #tanımlama eksik
return ozellikDetay
bisiklet1 = Bisiklet()
bisiklet2 = Bisiklet()
print("Bisiklet 1: " + bisiklet1.ozellikler())
bisiklet2.renk = "Sarı"
bisiklet2.vites = 22
print("Bisiklet 2: " + bisiklet2.ozellikler())
```
##__init__ Metodu
__init__ fonksiyonu, oluşturduğu nesneleri başlatmak için kullanılır. init "başlat" ın kısaltmasıdır. __init__() her zaman yaratılan nesneye atıfta bulunan en az bir argüman alır: "self".
**Örnek Uygulama**
Aşağıdaki örnek uygulamada *sporDali* sınıfının içinde tanımlanan **init** fonksiyonu, sınıf oluşturulduğu anda çalışmaya başlamaktadır. Fonksiyonun ayrıca çağırılmasına gerek kalmamıştır.
```
class sporDali:
sporlar = ["Yüzme", "Bisiklet", "Koşu"]
def __init__(self):
for spor in self.sporlar:
print(spor + " bir triatlon branşıdır.")
triatlon = sporDali()
```
```
# Örnek uygulamayı çalıştır
class sporDali:
sporlar = ["Yüzme", "Bisiklet", "Koşu"]
def __init__(self):
for spor in self.sporlar:
print(spor + " bir triatlon branşıdır.")
triatlon = sporDali()
# Örnek uygulamayı > Duatlon
class sporDali:
sporlar = ["Yüzme", "Bisiklet", "Koşu"]
def __init__(self):
for spor in self.sporlar:
print(spor + " bir triatlon branşıdır.")
triatlon = sporDali()
```
#Bölüm 09: Modüller ve Paketler
##Modülün İçe Aktarılması
Python'daki modüller, Python tanımlarını (sınıflar, fonksiyonlar vb.) ve ifadelerini (değişkenler, listeler, sözlükler vb.) içeren .py uzantısına sahip Python dosyalarıdır.
Modüller, *import* anahtar sözcüğü ve uzantı olmadan dosya adı kullanılarak içe aktarılır. Bir modül, çalışan bir Python betiğine ilk kez yüklendiğinde, modüldeki kodun bir kez çalıştırılmasıyla başlatılır.
**Örnek Uygulama**
```
#bisiklet.py adlı modülün içeriği
"""
Bu modül içinde Bisiklet sınıfı yer almaktadır.
"""
class Bisiklet:
renk = "Kırmızı"
vites = 1
def ozellikler(self):
ozellikDetay = "Bu bisiklet %s renkli ve %d viteslidir." % (self.renk, self.vites)
return ozellikDetay
```
```
#bisikletler.py adlı Python dosyasının içeriği
import bisiklet
bisiklet1 = bisiklet.Bisiklet()
print("Bisiklet 1: " + bisiklet1.ozellikler())
```
**PyCharm Örneği**
bisiklet.py
---
bisikletler.py
##Colab'de Modülün İçe Aktarılması
Bir önceki bölümde (Modülün İçe Aktarılması) herhangi bir kişisel bilgisayarın sabit diski üzerinde çalışırken yerleşik olmayan (kendi yazdığımız) modülün içe aktarılması yer aldı.
Bu bölümde ise Colab üzerinde çalışırken yerleşik olmayan bir modülü nasıl içe aktarılacağı yer almakta.
**Örnek Uygulama**
Aşağıda içeriği görüntülenen *bisiklet.py* adlı Python dosyası Google Drive içerisinde "BBY162_Python_a_Giris.ipynb" dosyasının ile aynı klasör içinde bulunmaktadır.
```
#bisiklet.py adlı modülün içeriği
"""
Bu modül içinde Bisiklet sınıfı yer almaktadır.
"""
class Bisiklet:
renk = "Kırmızı"
vites = 1
def ozellikler(self):
ozellikDetay = "Bu bisiklet %s renkli ve %d viteslidir." % (self.renk, self.vites)
return ozellikDetay
```
```
# Google Drive'ın bir disk olarak görülmesi
from google.colab import drive
drive.mount('gdrive') # bağlanan diskin 'gdrive' adı ile tanımlanması.
import sys # bağlanan diskin fiziksel yolunun tespit edilmesi ve bağlantı yoluna eklenmesi
sys.path.append('/content/gdrive/My Drive/Colab Notebooks/BBY162 - Programlama ve Algoritmalar/')
import bisiklet # bisiklet.py içerisindeki 'bisiklet' modülünün içe aktarılması
bisiklet1 = bisiklet.Bisiklet()
print("Bisiklet 1: " + bisiklet1.ozellikler())
```
##Yerleşik Modüller (built-in)
Python aşağıdaki bağlantıda yer alan standart modüllerle birlikte gelir. Bu modüllerin *import* anahtar kelimesi ile çağrılması yeterlidir. Ayrıca bu modüllerin yüklenmesine gerek yoktur.
[Python Standart Modülleri](https://docs.python.org/3/library/)
**Örnek Uygulama**
```
import datetime
print(datetime.datetime.today())
```
```
# Örnek uygulamayı çalıştır
import datetime
print(datetime.datetime.today())
```
##from import Kullanımı
İçe aktarma ifadesinin bir başka kullanım şekli *from* anahtar kelimesinin kullanılmasıdır. *from* ifadesi ile modül adları paketin içinde alınarak direkt kullanıma hazır hale getirilir. Bu şekilde, içe aktarılan modül, modül_adı öneki olmadan doğrudan kullanılır.
**Örnek Uygulama**
```
#bisiklet.py adlı modülün içeriği
"""
Bu modül içinde Bisiklet sınıfı yer almaktadır.
"""
class Bisiklet:
renk = "Kırmızı"
vites = 1
def ozellikler(self):
ozellikDetay = "Bu bisiklet %s renkli ve %d viteslidir." % (self.renk, self.vites)
return ozellikDetay
```
```
# Google Drive'ın bir disk olarak görülmesi
from google.colab import drive
drive.mount('gdrive') # bağlanan diskin 'gdrive' adı ile tanımlanması.
import sys # bağlanan diskin fiziksel yolunun tespit edilmesi ve bağlantı yoluna eklenmesi
sys.path.append('/content/gdrive/My Drive/Colab Notebooks/BBY162 - Programlama ve Algoritmalar/')
from bisiklet import Bisiklet # bisiklet.py içerisindeki 'bisiklet' sınıfının içe aktarılması
bisiklet1 = Bisiklet() # bisiklet ön tanımlamasına gerek kalmadı
print("Bisiklet 1: " + bisiklet1.ozellikler())
```
#Bölüm 10: Dosya İşlemleri
##Dosya Okuma
Python, bilgisayarınızdaki bir dosyadan bilgi okumak ve yazmak için bir dizi yerleşik fonksiyona sahiptir. **open** fonksiyonu bir dosyayı açmak için kullanılır. Dosya, okuma modunda (ikinci argüman olarak "r" kullanılarak) veya yazma modunda (ikinci argüman olarak "w" kullanılarak) açılabilir. **open** fonksiyonu dosya nesnesini döndürür. Dosyanın saklanması için kapatılması gerekir.
**Örnek Uygulama**
```
#Google Drive Bağlantısı
from google.colab import drive
drive.mount('/gdrive')
dosya = "/gdrive/My Drive/Colab Notebooks/BBY162 - Programlama ve Algoritmalar/metin.txt"
f = open(dosya, "r")
for line in f.readlines():
print(line)
f.close()
```
Dosyanın sağlıklı şekilde okunabilmesi için Google Drive ile bağlantının kurulmuş olması ve okunacak dosyanın yolunun tam olarak belirtilmesi gerekmektedir.

```
#Google Drive Bağlantısı
from google.colab import drive
drive.mount('/gdrive')
dosya = "/gdrive/My Drive/Colab Notebooks/BBY162 - Programlama ve Algoritmalar/metin.txt"
f = open(dosya, "r")
for line in f.readlines():
print(line)
f.close()
```
##Dosya Yazma
Bir dosyayı ikinci argüman olarak "w" (yazma) kullanarak açarsanız, yeni bir boş dosya oluşturulur. Aynı ada sahip başka bir dosya varsa silineceğini unutmayın. Mevcut bir dosyaya içerik eklemek istiyorsanız "a" (ekleme) değiştiricisini kullanmalısınız.
**Örnek Uygulama**
Aşağıdaki örnekte dosya 'w' parametresi ile açıldığı için var olan dosyanın içindekiler silinir ve yeni veriler dosyaya yazılır. Dosyanın içindeki verilerin kalması ve yeni verilerin eklenmesi isteniyorsa dosya 'a' parametresi ile açılmalıdır.
```
#Google Drive Bağlantısı
from google.colab import drive
drive.mount('/gdrive')
dosya = "/gdrive/My Drive/Colab Notebooks/BBY162 - Programlama ve Algoritmalar/cikti.txt"
f = open(dosya, 'w') # Mevcut veriye ek veri yazılması için parametre: 'a'
f.write("test") # Her yeni verinin bir alt satıra yazdırılması "test\n"
f.close()
```
Kod çalıştırıldıktan sonra eğer *cikti.txt* adında bir dosya yoksa otomatik olarak oluşturulur ve istenilen içerik yazılır.
```
#Google Drive Bağlantısı
from google.colab import drive
drive.mount('/gdrive')
dosya = "/gdrive/My Drive/Colab Notebooks/BBY162 - Programlama ve Algoritmalar/cikti.txt"
f = open(dosya, 'w') # Mevcut veriye ek veri yazılması için parametre: 'a'
f.write("test") # Her yeni verinin bir alt satıra yazdırılması "test\n"
f.close()
```
| github_jupyter |
# Activity 12 - Breast Cancer Diagnosis Classification using Logistic Regression
In this activity we will be using the Breast Cancer dataset [https://archive.ics.uci.edu/ml/datasets/Breast+Cancer+Wisconsin+(Diagnostic)]( https://archive.ics.uci.edu/ml/datasets/Breast+Cancer+Wisconsin+(Diagnostic) ) available under the [UCI Machine Learning Repository] (https://archive.ics.uci.edu/ml/index.php). The dataset contains characteristics of the cell nuclei present in the digitized image of a fine needle aspirate (FNA) of a breast mass, with the labels _malignant_ and _benign_ for each cell nucleus. Throughout this activity we will use the measurements provided in the dataset to classify between malignant and benign cells.
## Import the Required Packages
For this exercise we will require the Pandas package for loading the data, the matplotlib package for plotting as well as scikit-learn for creating the Logistic Regression model, doing some feature selection as well as model selection. Import all of the required packages and relevant modules for these tasks.
## Load the Data
Load the Breast Cancer Diagnosis dataset using Pandas and examine the first 5 rows
Dissect the data into input (X) and output (y) variables
## Feature Engineering
We need to select the most appropriate features that will provide the most powerful classification model. We will use scikit-learn's select K best features sub-module under its feature selection module. Basically it examines the power of each feature against the target output based on a scoring function. You may read about the details here : https://scikit-learn.org/stable/modules/generated/sklearn.feature_selection.SelectKBest.html
Let us visualize how these 2 most important features correlate with the target (diagnosis) and how well do they separate the 2 classes of diagnosis.
## Constructing the Logistic Regression Model
Before we can construct the model we must first convert the dignosis values into labels that can be used within the model. Replace:
1. The diagnosis string *benign* with the value 0
2. The diagnosis string *malignant* with the value 1
Also, in order to impartially evaluate the model, we should split the training dataset into a training and a validation set.
Create the model using the *selected_features* and the assigned *diagnosis* labels
Compute the accuracy of the model against the validation set:
Construct another model using a random choice of *selected_features* and compare the performance:
Construct another model using all available information and compare the performance:
| github_jupyter |
# Multinomial Naive Bayes Classifier with MinMaxScaler and Quantile Transformer
This Code template is for Classification tasks using MultinomialNB based on the Naive Bayes algorithm for multinomially distributed data with feature rescaling technique MinMaxScaler and feature transformation technique Quantile Transformation in a pipeline.
## Required Packages
```
!pip install imblearn
import warnings
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as se
from imblearn.over_sampling import RandomOverSampler
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import MinMaxScaler
from sklearn.preprocessing import QuantileTransformer
from sklearn.naive_bayes import MultinomialNB
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import train_test_split
from sklearn.metrics import classification_report,plot_confusion_matrix
warnings.filterwarnings('ignore')
```
## Initialization
Filepath of CSV file
```
#filepath
file_path= ""
```
List of features which are required for model training .
```
#x_values
features=[]
```
Target feature for prediction.
```
#y_value
target=''
```
## Data Fetching
Pandas is an open-source, BSD-licensed library providing high-performance, easy-to-use data manipulation and data analysis tools.
We will use panda's library to read the CSV file using its storage path.And we use the head function to display the initial row or entry.
```
df=pd.read_csv(file_path)
df.head()
```
## Feature Selection
It is the process of reducing the number of input variables when developing a predictive model. Used to reduce the number of input variables to both reduce the computational cost of modelling and, in some cases, to improve the performance of the model.
We will assign all the required input features to X and target/outcome to Y.
```
X = df[features]
Y = df[target]
```
## Data Preprocessing
Since the majority of the machine learning models in the Sklearn library doesn't handle string category data and Null value, we have to explicitly remove or replace null values. The below snippet have functions, which removes the null value if any exists. And convert the string classes data in the datasets by encoding them to integer classes.
```
def NullClearner(df):
if(isinstance(df, pd.Series) and (df.dtype in ["float64","int64"])):
df.fillna(df.mean(),inplace=True)
return df
elif(isinstance(df, pd.Series)):
df.fillna(df.mode()[0],inplace=True)
return df
else:return df
def EncodeX(df):
return pd.get_dummies(df)
def EncodeY(df):
if len(df.unique())<=2:
return df
else:
un_EncodedT=np.sort(pd.unique(df), axis=-1, kind='mergesort')
df=LabelEncoder().fit_transform(df)
EncodedT=[xi for xi in range(len(un_EncodedT))]
print("Encoded Target: {} to {}".format(un_EncodedT,EncodedT))
return df
x=X.columns.to_list()
for i in x:
X[i]=NullClearner(X[i])
Y=NullClearner(Y)
Y=EncodeY(Y)
X=EncodeX(X)
X.head()
```
## Correlation Map
In order to check the correlation between the features, we will plot a correlation matrix. It is effective in summarizing a large amount of data where the goal is to see patterns.
```
f,ax = plt.subplots(figsize=(18, 18))
matrix = np.triu(X.corr())
se.heatmap(X.corr(), annot=True, linewidths=.5, fmt= '.1f',ax=ax, mask=matrix)
plt.show()
```
## Distribution Of Target Variable
```
plt.figure(figsize = (10,6))
se.countplot(Y)
```
## Data Splitting
The train-test split is a procedure for evaluating the performance of an algorithm. The procedure involves taking a dataset and dividing it into two subsets. The first subset is utilized to fit/train the model. The second subset is used for prediction. The main motive is to estimate the performance of the model on new data.
```
x_train,x_test,y_train,y_test=train_test_split(X,Y,test_size=0.2,random_state=123)
```
## Handling Target Imbalance
The challenge of working with imbalanced datasets is that most machine learning techniques will ignore, and in turn have poor performance on, the minority class, although typically it is performance on the minority class that is most important.
One approach to addressing imbalanced datasets is to oversample the minority class. The simplest approach involves duplicating examples in the minority class.We will perform overspampling using imblearn library.
```
x_train,y_train = RandomOverSampler(random_state=123).fit_resample(x_train, y_train)
```
## Model
With a multinomial event model, samples (feature vectors) represent the frequencies with which certain events have been generated by a multinomial probability that an event occurs.
The multinomial Naive Bayes classifier is suitable for classification with discrete features. The multinomial distribution normally requires integer feature counts. However, in practice, fractional counts such as tf-idf may also work.
Model Tuning Parameters
1. alpha : float, default=1.0
Additive (Laplace/Lidstone) smoothing parameter (0 for no smoothing).
2. fit_prior : bool, default=True
Whether to learn class prior probabilities or not. If false, a uniform prior will be used.
3. class_prior : array-like of shape (n_classes,), default=None
Prior probabilities of the classes. If specified the priors are not adjusted according to the data.
#### MinMax Scaler:
MinMax Scaler shrinks the data within the given range, usually of 0 to 1. It transforms data by scaling features to a given range. It scales the values to a specific value range without changing the shape of the original distribution.
#### Feature Transformation:
Quantile transforms are a technique for transforming numerical input or output variables to have a Gaussian or uniform probability distribution.
```
model = make_pipeline(QuantileTransformer(output_distribution='normal'),MinMaxScaler(),MultinomialNB())
model.fit(x_train, y_train)
```
## Model Accuracy
score() method return the mean accuracy on the given test data and labels.
In multi-label classification, this is the subset accuracy which is a harsh metric since you require for each sample that each label set be correctly predicted.
```
print("Accuracy score {:.2f} %\n".format(model.score(x_test,y_test)*100))
```
## Confusion Matrix
A confusion matrix is utilized to understand the performance of the classification model or algorithm in machine learning for a given test set where results are known.
```
plot_confusion_matrix(model,x_test,y_test,cmap=plt.cm.Blues)
```
## Classification Report
A Classification report is used to measure the quality of predictions from a classification algorithm. How many predictions are True, how many are False.
where:
- Precision:- Accuracy of positive predictions.
- Recall:- Fraction of positives that were correctly identified.
- f1-score:- percent of positive predictions were correct
- support:- Support is the number of actual occurrences of the class in the specified dataset.
```
print(classification_report(y_test,model.predict(x_test)))
```
#### Creator: Neel Pawar, Github: [Profile]( https://github.com/neel-ntp)
| github_jupyter |
```
from matplotlib import pyplot as plt
import numpy as np
import math
%matplotlib inline
```
# Numerical Solution of a Two-Dimensional Supersonic Flow: Prandtl-Meyer Expansion Wave
This notebook looks that what hppends when a nonviscous flow goes over an expansion corner at a supersonic velocity. We will be implmenting a space marching solution instead of a time marching solution. This example is taken from chapter 8 of John D. Anderson's book, *Computational Fluid Dynamics: The Basics with Applications*.
## Introduction: Physical Problem and Analytical Solution
A Prandtl-Meyer expansion wave is made up of an infinite number of infinitely weak shcokwaves that fan out around a corner. This is caused by the expansion of a compressible fluid at supersonic speeds. The leading edge of the fan makes an angle $\mu_1$ wtih respect to the upsttream flow and an agle $\mu_2$ with respect to the downstream flow. Analytically, these values are defined as
$\mu_1 = sin^{-1}\frac{1}{M_1}$
$\mu_2 = sin^{-1}\frac{1}{M_2}$
$M_1$ and $M_2$ are the upstream and downsteam mach numbers,respectively.
The flow through the fan is isentropic and we know that as a supersonic gas expands, the velocity increases as the temperature and density decrease. The flow before the fan is parallel to the wall boundary and after the expansion fan, the flow is parallel to the declined ramp. All of these flow characteristics hold true for the mjority of the flow except for the corner of the expansion fan. At this point, all of the arms of the fan branch out causing a disconinuous change in the flow direction. This will cause numerical problems if not hadled correctly.
For a calorically perfect gas, there is an analytical solution for that point based on the defelction angle of the ramp which we will call $\theta$. The simple relation is calculated as such.
$f_2 = f_1 + \theta$
The value of $f$ can be calculated with the Prandtl-Meyer function below
$f = \sqrt{\frac{\gamma + 1}{\gamma - 1}} tan^{-1}\sqrt{\frac{\gamma-1}{\gamma+1}(M^2 -1 )}-tan^{-1}\sqrt{M^2-1}$
This relation allows you to calculate the mach number if oyu know the upstream mach number and the angle of the downward slope $\theta$.
Once we have the mach number, we can use the isentropic flow relations to get the values for pressure and temperature downstream.
$p_2 = p_1 \Big \{ \frac{1+[(\gamma-1)/2]M_1^2}{1+[(\gamma-1)/2]M_2^2}\Big \} ^{\gamma / (\gamma -1)}$
$T_2 = T_1 \frac{1+[(\gamma-1)/2]M_1^2}{1+[(\gamma-1)/2]M_2^2}$
and we can calculate density using the following equation
$\rho_2 = \frac{p_2}{RT_2}$
## Numerical Solution
We will be using a space marching solution to solve this numerically. That means that instead of marching through steps of time, we will begin at the inflow boundary condition and progress down stream by increments of $\Delta x$. We will still be using MacCormack's method, but with space (in x-direction) instead of time.
### Governing Equations
For a steady state Euler flow in two dimensions,we will use the strong conservation form as displayed below.
$\frac{\partial F}{\partial x} = J - \frac{\partial G}{\partial y}$
The values of $F$ and $G$ are vectors that represent the flow variables as such.
$F_1 = \rho u$
$F_2 = \rho u^2 + p$
$F_3 = \rho u v$
$F_4 = \rho u \Big(e+\frac{V^2}{2}\Big)+pu$
$G_1 = \rho v$
$G_2 = \rho uv$
$G_3 = \rho v^2 +p$
$G_4 = \rho v \Big(e+\frac{V^2}{2}\Big)+pv$
We can simplify the energy equations for $F$ and $G$ with the following identity of a calorically perfect gas.
$e = c_v T = \frac{RT}{\gamma -1} = \frac{1}{\gamma - 1} = \frac{1}{\gamma -1}\frac{p}{\rho}$
$F_4 = \rho u \Big( \frac{1}{\gamma-1}\frac{p}{\rho} + \frac{u^2 + v^2}{2}\Big) + pu$
$ = \frac{1}{\gamma -1}pu + \rho u \frac{u^2 + v^2}{2} + pu$
Finally we have a form of the equation without e:
$F_4 = \frac{\gamma}{\gamma -1}pu + \rho u \frac{u^2 + v^2}{2}$
Doing the same thing for $G_4$, we get
$G_4 = \frac{\gamma}{\gamma -1}pv + \rho v \frac{u^2 + v^2}{2}$
Because we are not running this simulation with a time marching solution, we do not have a $U$ vector to ectract our primative variables. Instead we will be decoded from the flux variables $F$. We can use the following equations to do so
$\rho = \frac{-B + \sqrt{B^2 - 4AC}}{2A}$
$A = \frac{F^2_3}{2F_1} - F_4$
$B = \frac{\gamma}{\gamma - 1}F_1 F_2$
$C = -\frac{\gamma +1}{2(\gamma -1)}F_1^3$
$u = \frac{F_1}{\rho}$
$v = \frac{F_3}{F_1}$
$p = F_2 - F_1 u$
$T = \frac{p}{\rho R}$
As for the calculation of the $G$ vector values, we can get them in terms of $F$ instead of the primative variables.
$G_1 = \rho v = \rho \frac{F_3}{F_1}$
$G_2 = F_3$
$G_3 = \rho v^2 + p = \rho \Big(\frac{F_3}{F_1}\Big)^2 + p$
$p = F_2 - \rho u^2 = F_2 - \frac{F_1^2}{\rho}$
$G_3 = \rho \Big(\frac{F_3}{F_1}\Big)^2 + F_2 - \frac{F_1^2}{\rho}$
$G_4 =\frac{\gamma}{\gamma -1} pv + \rho v \frac{u^2 + v^2}{2} = \frac{\gamma}{\gamma -1} \Big(F_2 - \frac{F_1^2}{\rho}\Big)\frac{F_3}{F_1}+\frac{\rho}{2} \frac{F_3}{F_1}\Big[\Big(\frac{F_1}{\rho}\Big)^2 + \Big(\frac{F_3}{F_1}\Big)^2\Big]$
Before we begin setting upour simulation, we need to consider our geometry. Below is a plot of the geometry in the physical plane.
```
x = np.linspace(0,65,200)
h = np.zeros(200)
index = 0
for i in x:
if i <10:
h[index] = 40
else:
h[index] = 40+math.tan(0.09341002)*(x[index]-10) # make sure to convert to radians
index = index +1
#plot geometry
plt.figure(figsize=(7,7))
plt.plot(x,np.ones(200)*40,'black')
plt.plot(x,40-h,'black')
plt.ylim([-15,50])
plt.xlim([0,65])
plt.grid()
plt.title('Physical Plane Geometry: Ramp Angle = 5.352 deg')
plt.xlabel('x')
plt.ylabel('y')
```
This geometry does not work with a purely structured cartesian grid. In order to account for the declined ramp, we need to use a transformation that will show up in our governing equations.Here we will use a boundary fitted coordiate system with variables $\eta$ and $\zeta$. The symbol eta $\eta$ represents the spacing up and down in the physical plane and the symbol $\zeta$ represents the spacing left and right in the physical plane. The transformation from $(x,y)$ to $(\zeta,\eta)$ will be as follows
$\zeta = x$
$\eta = \frac{y-y_s (x)}{h(x)}$
Where $y_s (x)$ is the y location of the lower boundary surface, and $h(x)$ is the distance between the top and bottom boundaries. This transformation creates a nice cratesian mesh in the $(\zeta,\eta)$ plane where $\zeta$ is between 0 and L and $\eta$ varies from 0 to 1.0.
With this transformation, we also have adjust our governing equations. The following equations show the proper transformation.
$\frac{\partial}{\partial x} = \frac{\partial}{\partial \zeta} \Big( \frac{\partial \zeta}{\partial x}\Big) + \frac{\partial}{\partial \eta}\Big( \frac{\partial \eta}{\partial x}\Big)$
$\frac{\partial}{\partial y} = \frac{\partial}{\partial \zeta} \Big( \frac{\partial \zeta}{\partial y}\Big) + \frac{\partial}{\partial \eta}\Big( \frac{\partial \eta}{\partial y}\Big)$
Pluggin in our geometric transformation into the derivative transformation, we get the following metrics
$\frac{\partial \zeta}{\partial x} = 1$
$\frac{\partial \zeta}{\partial y} = 0$
$\frac{\partial \eta}{\partial x} = -\frac{1}{h} \frac{dy_s}{dx} - \frac{\eta}{h}\frac{\partial h}{\partial x}$
$\frac{\partial \eta}{\partial y} = \frac{1}{h}$
The one complicated term is $\partial \eta /\partial x$, so let's transform it into something simpler to deal with. For the x location of the corner of the expasion wave set to value $E$.
For $x \leq E$:
$y_s = 0$
$h = const$
For $x \geq E$:
$y_s = -(x-E)tan\theta$
$h = H +(x-E)tan\theta$
Differentiating these expressions, we get the following
For $x \leq E$:
$\frac{dy_s}{dx} = 0$
$\frac{dh}{dx} = 0$
For $x \geq E$:
$\frac{dy_s}{dx} = -tan\theta$
$\frac{dh}{dx} = tan\theta$
So the metric of $\partial \eta /\partial x$ can be described as:
$\frac{\partial \eta}{\partial x} = 0$ for $x \leq E$
$\frac{\partial \eta}{\partial x} = (1-\eta)\frac{tan\theta}{h}$ for $x \geq E$
Plugging these metrics back into the derivative transformation, we get that
$\frac{\partial}{\partial x} =\frac{\partial }{\partial \zeta} + \Big(\frac{\partial \eta}{\partial x}\Big)\frac{\partial}{\partial eta}$
$\frac{\partial}{\partial y} = \frac{1}{h}\frac{\partial }{\partial \eta}$
With $J=0$ we can write our govering equaiton.
$\frac{\partial F}{\partial x} = - \frac{\partial G}{\partial y}$
Now we we implement the transformation derivative to put our equation in the computational plane.
$\frac{\partial F}{\partial \zeta} + \Big(\frac{\partial \eta}{\partial x}\Big)\frac{\partial F}{\partial \eta} = -\frac{1}{h}\frac{\partial G}{\partial \eta}$
or rewritten as
$\frac{\partial F}{\partial \zeta} = - \Big[\Big(\frac{\partial \eta}{\partial x}\Big)\frac{\partial F}{\partial \eta}+\frac{1}{h}\frac{\partial G}{\partial \eta}\Big]$
To put this in terms of our flow equations, we have
Continuity: $\frac{\partial F_1}{\partial \zeta} = - \Big[\Big(\frac{\partial \eta}{\partial x}\Big)\frac{\partial F_1}{\partial \eta}+\frac{1}{h}\frac{\partial G_1}{\partial \eta}\Big]$
x momentum: $\frac{\partial F_2}{\partial \zeta} = - \Big[\Big(\frac{\partial \eta}{\partial x}\Big)\frac{\partial F_2}{\partial \eta}+\frac{1}{h}\frac{\partial G_2}{\partial \eta}\Big]$
y momentum: $\frac{\partial F_3}{\partial \zeta} = - \Big[\Big(\frac{\partial \eta}{\partial x}\Big)\frac{\partial F_3}{\partial \eta}+\frac{1}{h}\frac{\partial G_3}{\partial \eta}\Big]$
Energy: $\frac{\partial F_4}{\partial \zeta} = - \Big[\Big(\frac{\partial \eta}{\partial x}\Big)\frac{\partial F_4}{\partial \eta}+\frac{1}{h}\frac{\partial G_4}{\partial \eta}\Big]$
### The Setup
#### Artificial Viscosity
As we prepare to solve our problem, we need to address two big topics: artificial viscosity and boundary conditions. The artificial viscosity will be handled in the same fashion as the nozzle flow problem and will be represented as such.
$S'^t_i = \frac{C_y |(p')^{t'}_{i+1} - 2(p')^{t'}_i +(p')^{t'}_{i-1}|}{(p')^{t'}_{i+1} + 2(p')^{t'}_i +(p')^{t'}_{i+1}}(F^{t'}_{i+1}-2F^{t'}_{i}+F^{t'}_{i-1})$
This value will then be added to the calculation of the predictor and corrector step.
#### Boundary Conditions
As for the boundary condition, we want to enforce that at the wall there is no flow normal to the wall. We do this by uodating our primitive variables to ones that represent a flow that only has a tanget component of velocity to the wall. We do thsi by using Abbett's method. First first step is to calculate the angle the flow has with the wall and the mach number at the current node with the following equations
$\phi_1 = tan^{-1}\frac{v_1}{u_1}$
$(M_1)_{cal} = \frac{ \sqrt{(u_1)_{cal}^2+(v_1)_{cal}^2} }{(a_1)_{cal}}$
Then we use the Prandtl-Meyer function to get the value for $f_{cal}$
$f_{cal} = \sqrt{\frac{\gamma + 1}{\gamma - 1}} tan^{-1}\sqrt{\frac{\gamma-1}{\gamma+1}(M_{cal}^2 -1 )}-tan^{-1}\sqrt{M_{cal}^2-1}$
Then we correct the value of $f$ with $\phi$
$f_{act} = f_{cal}+\phi_1$
Then using the same equation used to et $f$, we backout the value for our new mach number.
Lastly, we use the following equations to get the rest of the primitive variables
$p_2 = p_1 \Big \{ \frac{1+[(\gamma-1)/2]M_1^2}{1+[(\gamma-1)/2]M_2^2}\Big \} ^{\gamma / (\gamma -1)}$
$T_2 = T_1 \frac{1+[(\gamma-1)/2]M_1^2}{1+[(\gamma-1)/2]M_2^2}$
$\rho_2 = \frac{p_2}{RT_2}$
There is one more thing to look out for when we are calculating the BC's and that is the change in slope of the ramp. After the horizontal part, the ramp changes to a decline. We account for this in the calculation of $f_{act}$
$f_{act} = f_{cal}+\phi_2$
$\psi = tan^{-1}\frac{|v_2|}{u_2}$
$\phi_2 = \theta - \psi$
As we prepare to run our simulation, let us define some functions that will help simplify our code later down the line. The first few funcitons will help with conversions. The following functions will help reduce repeated code and help prevent mistakes by reducing clutter. Additionally, by splitting up our code into small function we can test them individually and trouble shoot each part of the code with known input and output.
```
def primatives2F(u,v,rho,pres,gamma):
# Converting primitive variables to F vector
F1 = rho*u
F2 = rho*(u**2) + pres
F3 = rho * u * v
F4 = ( (gamma/(gamma-1)) * pres * u ) + ( rho * u * ((u**2 + v**2)/2) )
return F1,F2,F3,F4
def F2G(F1,F2,F3,F4):
# Calculate F values to G
gamma=1.4
A = ( (F3**2) / (2*F1) ) - F4
B = (gamma/ (gamma-1) ) * F1 * F2
C = -( (gamma+1) / ( 2 * (gamma-1) ) ) * F1**3
rho = (-B+( ( (B**2) - 4*A*C )**0.5 ) ) / (2*A)
G1 = rho*(F3/F1)
G2 = F3
G3 = (rho*(F3/F1)**2) + F2 - ((F1**2)/rho)
G4 = (gamma/(gamma-1)) * ( F2- ((F1**2)/rho) ) * (F3/F1) + ( (rho/2) * (F3/F1) * ( ((F1/rho)**2) + ((F3/F1)**2) ) )
return G1,G2,G3,G4
def F2Primatives(F1,F2,F3,F4,gamma):
# Extract primitive variables from F
A = ( (F3**2) / (2*F1) ) - F4
B = (gamma/ (gamma-1) ) * F1 * F2
C = -( (gamma+1) / ( 2 * (gamma-1) ) ) * F1**3
rho = (-B+( ( (B**2) - 4*A*C )**0.5 ) ) / (2*A)
u = F1/rho
v = F3/F1
pres = F2-F1*u
return u,v,rho,pres
```
The next set of function will help us calcualte our boundary conditions. There are a few steps to the BC calcualtion process. First we have to get the corrected mach value. To do this, we use the equations stated above to adjust the vlaue for $f$. However, we need to back out the coresponding mach number with the function: backOutMach. In this function, we iterate until we converge on a mach number that gets us close to our desired $f$. Once we have the corrected mach number, we extract the rest of the primatives with the mach2Primitives function.
```
def wallBC(u,v,rho,pres,x,theta,gamma,R):
# Calculate phi depending on x location
if x <= 10:
phi = math.atan(v/u)
else:
phi = theta - math.atan(abs(v)/u)
# Calculate corrected value
temp = pres/(rho*R)
a =((gamma*pres/rho)**0.5)
MCal = math.sqrt(u**2 + v**2)/a
fCal = math.sqrt((gamma+1)/(gamma-1)) * math.atan(math.sqrt( ( (gamma-1)/(gamma+1) ) * (MCal**2 -1) )) - math.atan(math.sqrt((MCal**2) -1))
fAct = fCal+phi
Mact = backOutMach(fAct,gamma)
pAct,TAct,rhoAact = mach2Primitives(MCal,Mact,pres,temp,R,gamma)
vNew = -u*math.tan(theta)# v is the tan(theta) component of u
uNew = u # u is kept the same
return uNew,vNew,rhoAact,pAct
def backOutMach(f,gamma):
# given some value of f, we need to backout M
# f input in deg converted to rad
guessM = 4
delta = 1
while abs(delta)>0.0000001:
fGuess = math.sqrt((gamma+1)/(gamma-1)) * math.atan(math.sqrt( ( (gamma-1)/(gamma+1) ) * (guessM**2 -1) )) - math.atan(math.sqrt(guessM**2 -1))
delta = fGuess - f
guessM = guessM - delta*0.1
return guessM
def mach2Primitives(Mcal,Mact,pcal,Tcal,R,gamma):
# Calculate primitives from mach numbers and new p & T
top = 1 + ((gamma-1)/2)*Mcal**2
bottom = 1 + ((gamma-1)/2)*Mact**2
pAct = pcal * (top/bottom) ** ( gamma/(gamma-1) )
TAct = Tcal * (top/bottom)
rhoAact = pAct/(R*TAct)
return pAct,TAct,rhoAact
```
### Iteration Scheme
```
def marchInX(u,v,rho,pres,gamma,dEtadx,deltaEta,deltaZeta,h,cVisc,x,theta,R):
"""
This function marches in the x-direction for
a space marching solution of a supersonic P-M
expansion fan problem
u - velocity in x
v - velocity in y
rho - density
pres - pressure
gamma - heat coefficient ratio (1.4 for air)
dEtadx - partial eta / partial x
deltaEta - step size of eta (analog to y but in computational plane)
deltaZeta - step size in zeta (analog to x but in computational plane)
cVisc - artificial viscosity coefficient
h - distance between top of gid and wall BC
x - x location of current node
theta - angle of ramp
R - gas constant
"""
numPts = 41
F1,F2,F3,F4 = primatives2F(u,v,rho,pres,gamma)
G1,G2,G3,G4 = F2G(F1,F2,F3,F4)
##########################
##### Predictor Step #####
##########################
# Initialize array for rate of change for predictor step
dF1dZeta = np.zeros(numPts)
dF2dZeta = np.zeros(numPts)
dF3dZeta = np.zeros(numPts)
dF4dZeta = np.zeros(numPts)
# Initialize array for first forward predictor step
F1Bar = np.zeros(numPts)
F2Bar = np.zeros(numPts)
F3Bar = np.zeros(numPts)
F4Bar = np.zeros(numPts)
G1Bar = np.zeros(numPts)
G2Bar = np.zeros(numPts)
G3Bar = np.zeros(numPts)
G4Bar = np.zeros(numPts)
# Initialize array for artificial viscosity
sf1 = np.zeros(numPts)
sf2 = np.zeros(numPts)
sf3 = np.zeros(numPts)
sf4 = np.zeros(numPts)
# Initialize array for parimitives forward step
uBar = np.zeros(numPts)
vBar = np.zeros(numPts)
rhoBar = np.zeros(numPts)
presBar = np.zeros(numPts)
for j in range(0,numPts-1):
dF1dZeta[j] = ( dEtadx[j] * ( (F1[j] - F1[j+1]) / (deltaEta) ) ) + (1/h) * ( (G1[j] - G1[j+1]) / (deltaEta) )
dF2dZeta[j] = ( dEtadx[j] * ( (F2[j] - F2[j+1]) / (deltaEta) ) ) + (1/h) * ( (G2[j] - G2[j+1]) / (deltaEta) )
dF3dZeta[j] = ( dEtadx[j] * ( (F3[j] - F3[j+1]) / (deltaEta) ) ) + (1/h) * ( (G3[j] - G3[j+1]) / (deltaEta) )
dF4dZeta[j] = ( dEtadx[j] * ( (F4[j] - F4[j+1]) / (deltaEta) ) ) + (1/h) * ( (G4[j] - G4[j+1]) / (deltaEta) )
sf1[j] = cVisc*(abs(pres[j+1]-2*pres[j]+pres[j-1])/(pres[j+1]+2*pres[j]+pres[j-1])) * (F1[j+1]-2*F1[j]+F1[j-1])
sf2[j] = cVisc*(abs(pres[j+1]-2*pres[j]+pres[j-1])/(pres[j+1]+2*pres[j]+pres[j-1])) * (F2[j+1]-2*F2[j]+F2[j-1])
sf3[j] = cVisc*(abs(pres[j+1]-2*pres[j]+pres[j-1])/(pres[j+1]+2*pres[j]+pres[j-1])) * (F3[j+1]-2*F3[j]+F3[j-1])
sf4[j] = cVisc*(abs(pres[j+1]-2*pres[j]+pres[j-1])/(pres[j+1]+2*pres[j]+pres[j-1])) * (F4[j+1]-2*F4[j]+F4[j-1])
if j==0:
F1Bar[j] = F1[j] + dF1dZeta[j]*deltaZeta + sf1[j]
F2Bar[j] = F2[j] + dF2dZeta[j]*deltaZeta + sf2[j]
F3Bar[j] = F3[j] + dF3dZeta[j]*deltaZeta + sf3[j]
F4Bar[j] = F4[j] + dF4dZeta[j]*deltaZeta + sf4[j]
else:
F1Bar[j] = F1[j] + dF1dZeta[j]*deltaZeta + sf1[j]
F2Bar[j] = F2[j] + dF2dZeta[j]*deltaZeta + sf2[j]
F3Bar[j] = F3[j] + dF3dZeta[j]*deltaZeta + sf3[j]
F4Bar[j] = F4[j] + dF4dZeta[j]*deltaZeta + sf4[j]
G1Bar[j],G2Bar[j],G3Bar[j],G4Bar[j] = F2G(F1Bar[j],F2Bar[j],F3Bar[j],F4Bar[j])
uBar[j],vBar[j],rhoBar[j],presBar[j] = F2Primatives(F1Bar[j],F2Bar[j],F3Bar[j],F4Bar[j],gamma)
"""
!!!CAUTION!!!
One thing to look out for here is that node 41 did not get any bar values
because there is no node to predict to, but we know from our BC's that
we will just set it to our initial condition values
"""
presBar[numPts-1]= pres[numPts-1]
F1Bar[numPts-1] = F1[numPts-1]
F2Bar[numPts-1] = F2[numPts-1]
F3Bar[numPts-1] = F3[numPts-1]
F4Bar[numPts-1] = F4[numPts-1]
##########################
##### Corrector Step #####
##########################
# Intitialize array for rate of change corrector step
dF1dZetaBar = np.zeros(numPts)
dF2dZetaBar = np.zeros(numPts)
dF3dZetaBar = np.zeros(numPts)
dF4dZetaBar = np.zeros(numPts)
# Averaged rate of change (corrector + predictor)/2
dF1dAvg = np.zeros(numPts)
dF2dAvg = np.zeros(numPts)
dF3dAvg = np.zeros(numPts)
dF4dAvg = np.zeros(numPts)
# Viscosity value for corrector step
sf1Bar = np.zeros(numPts)
sf2Bar = np.zeros(numPts)
sf3Bar = np.zeros(numPts)
sf4Bar = np.zeros(numPts)
for j in range(0,numPts-1):
if j ==0:# forward step for node 1 due to lack of node before it
dF1dZetaBar[j] = ( dEtadx[j] *( (F1Bar[j] - F1Bar[j+1]) / (deltaEta) ) ) + (1/h) * ( (G1Bar[j] - G1Bar[j+1]) / (deltaEta) )
dF2dZetaBar[j] = ( dEtadx[j] *( (F2Bar[j] - F2Bar[j+1]) / (deltaEta) ) ) + (1/h) * ( (G2Bar[j] - G2Bar[j+1]) / (deltaEta) )
dF3dZetaBar[j] = ( dEtadx[j] *( (F3Bar[j] - F3Bar[j+1]) / (deltaEta) ) ) + (1/h) * ( (G3Bar[j] - G3Bar[j+1]) / (deltaEta) )
dF4dZetaBar[j] = ( dEtadx[j] *( (F4Bar[j] - F4Bar[j+1]) / (deltaEta) ) ) + (1/h) * ( (G4Bar[j] - G4Bar[j+1]) / (deltaEta) )
else:# the rest of the nodes (excluding the last one get a rearwards step)
dF1dZetaBar[j] = ( dEtadx[j] *( (F1Bar[j-1] - F1Bar[j]) / (deltaEta) ) ) + (1/h) * ( (G1Bar[j-1] - G1Bar[j]) / (deltaEta) )
dF2dZetaBar[j] = ( dEtadx[j] *( (F2Bar[j-1] - F2Bar[j]) / (deltaEta) ) ) + (1/h) * ( (G2Bar[j-1] - G2Bar[j]) / (deltaEta) )
dF3dZetaBar[j] = ( dEtadx[j] *( (F3Bar[j-1] - F3Bar[j]) / (deltaEta) ) ) + (1/h) * ( (G3Bar[j-1] - G3Bar[j]) / (deltaEta) )
dF4dZetaBar[j] = ( dEtadx[j] *( (F4Bar[j-1] - F4Bar[j]) / (deltaEta) ) ) + (1/h) * ( (G4Bar[j-1] - G4Bar[j]) / (deltaEta) )
sf1Bar[j] = cVisc*(abs(presBar[j+1]-2*presBar[j]+presBar[j-1])/(presBar[j+1]+2*presBar[j]+presBar[j-1])) * (F1Bar[j+1]-2*F1Bar[j]+F1Bar[j-1])
sf2Bar[j] = cVisc*(abs(presBar[j+1]-2*presBar[j]+presBar[j-1])/(presBar[j+1]+2*presBar[j]+presBar[j-1])) * (F2Bar[j+1]-2*F2Bar[j]+F2Bar[j-1])
sf3Bar[j] = cVisc*(abs(presBar[j+1]-2*presBar[j]+presBar[j-1])/(presBar[j+1]+2*presBar[j]+presBar[j-1])) * (F3Bar[j+1]-2*F3Bar[j]+F3Bar[j-1])
sf4Bar[j] = cVisc*(abs(presBar[j+1]-2*presBar[j]+presBar[j-1])/(presBar[j+1]+2*presBar[j]+presBar[j-1])) * (F4Bar[j+1]-2*F4Bar[j]+F4Bar[j-1])
dF1dAvg[j] = 0.5*(dF1dZetaBar[j]+dF1dZeta[j])
dF2dAvg[j] = 0.5*(dF2dZetaBar[j]+dF2dZeta[j])
dF3dAvg[j] = 0.5*(dF3dZetaBar[j]+dF3dZeta[j])
dF4dAvg[j] = 0.5*(dF4dZetaBar[j]+dF4dZeta[j])
if j==0:
F1[j] = F1[j] + dF1dAvg[j]*deltaZeta + sf1Bar[j]
F2[j] = F2[j] + dF2dAvg[j]*deltaZeta + sf2Bar[j]
F3[j] = F3[j] + dF3dAvg[j]*deltaZeta + sf3Bar[j]
F4[j] = F4[j] + dF4dAvg[j]*deltaZeta + sf4Bar[j]
else:
F1[j] = F1[j] + dF1dAvg[j]*deltaZeta + sf1Bar[j]
F2[j] = F2[j] + dF2dAvg[j]*deltaZeta + sf2Bar[j]
F3[j] = F3[j] + dF3dAvg[j]*deltaZeta + sf3Bar[j]
F4[j] = F4[j] + dF4dAvg[j]*deltaZeta + sf4Bar[j]
# Extract primitives
u[j],v[j],rho[j],pres[j]= F2Primatives(F1[j],F2[j],F3[j],F4[j],gamma)
"""At this point, we have all of the updated
values for our flow except the boundary
conditions. The top boundary condition will
just be the free stream, but we will need to
do some calculations for the wall BC. Because
we never updated any value for the Nth node,
the BC is automatically enforced."""
u[0],v[0],rho[0],pres[0] = wallBC(u[0],v[0],rho[0],pres[0],x,theta,gamma,R)
return u,v,rho,pres
```
### Running the Solution and Results
```
def main(cCFL,cVisc):
# Flow variables
gamma = 1.4
R=287
# Set initial data line
u = np.ones([41,1])*678
v = np.zeros([41,1])
rho = np.ones([41,1])*1.23
pres = np.ones([41,1])*101000
mach = np.ones([41,1])*2
x=0
xValues = np.zeros([1,1])
xValues[0] = x
yValues = np.zeros([41,1])
yValues[:,0] = np.transpose(np.linspace(0,40,41))
# Initialize space for updated value arrays
uNew = np.zeros([41,1])
vNew = np.zeros([41,1])
rhoNew = np.zeros([41,1])
presNew = np.zeros([41,1])
thetaFreeStream = np.zeros(41)
mu = np.zeros(41)
bottom1 = np.zeros(41)
bottom2 = np.zeros(41)
i=0
while x<=65: # 65 is the x length of our grid
# Calculate h
if x<=10:
# Calculate h
h = 40
theta = 0
dEtadx = np.zeros(41)
else: # ramp starts at x=10
h = 40+math.tan(0.09341002)*(x-10)
theta = 0.09341002
eta = np.linspace(0,1,41)
dEtadx = (1-eta)*math.tan(theta)/h
#Calculate y values for post processing
y = np.zeros([41,1])
y[:,0] = np.transpose(np.linspace(40-h,40,41))
# Calculate deltaEta
deltaEta = 1/40
# Calculate deltaZeta
deltaY = h/40
for k in range(0,41):
thetaFreeStream[k] = math.atan(v[k,i]/u[k,i])
mu[k] = math.asin(1/mach[k,i])
bottom1[k] = abs(math.tan(thetaFreeStream[k]+mu[k]))
bottom2[k] = abs(math.tan(thetaFreeStream[k]-mu[k]))
deltaZeta1 = cCFL*(deltaY/(bottom1.max()))
deltaZeta2 = cCFL*(deltaY/(bottom2.max()))
deltaZeta = deltaZeta2
if deltaZeta1<deltaZeta2:
deltaZeta = deltaZeta1
# Solve for next step
uNew[:,0],vNew[:,0],rhoNew[:,0],presNew[:,0] = marchInX(u[:,i],v[:,i],rho[:,i],pres[:,i],gamma,dEtadx,deltaEta,deltaZeta,h,cVisc,x,theta,R)
# Append results of next step to result array
u = np.append(u,uNew,1)
v = np.append(v,vNew,1)
rho = np.append(rho,rhoNew,1)
pres = np.append(pres,presNew,1)
machNew =((uNew**2 + vNew**2)**0.5)/((gamma*presNew/rhoNew)**0.5)
mach = np.append(mach,machNew,1)
xValues = np.append(xValues,x)
yValues = np.append(yValues,y,1)
# Update x and i
x=x+deltaZeta
i=i+1
print('Done.')
return u,v,rho,pres,mach,xValues,yValues
u,v,rho,pres,mach,xValues,yValues = main(0.5,0.62)
```
With the simulation complete, we have the results for each point in our grid. For these conditions, the x axis was broken into 81 nodes so in total, this simulation had 3,321 grid points! Compared to most comercial solvers, this is very coarse, but for an beginning CFD researcher, this is leaps and bounds ahead of a 41 node 1-D simulation.
The most comprihensive way to plot our results is to display the 2D mesh with contour lines showing pressure and quivers showing the direction of the flow at each node. To do this, we will use the contourf and quiver functions from the matplotlib library.
```
x = np.linspace(0,65,200)
h = np.zeros(200)
index = 0
for i in x:
if i <10:
h[index] = 40
else:
h[index] = 40+math.tan(0.09341002)*(x[index]-10)
index = index +1
#plot geometry
plt.figure(figsize=(16,12))
plt.plot(x,np.ones(200)*40,'black')
plt.plot(x,40-h,'black')
plt.ylim([-15,50])
plt.xlim([0,65])
plt.grid()
plt.title('2D Supersonic Flow: Ramp Angle = 5.352 deg')
plt.xlabel('x')
plt.ylabel('y')
# Plot Results
xMatrix = np.array([xValues,]*41)
plt.contourf(xMatrix, yValues, pres, alpha=0.7)
plt.colorbar(label='Pressure N/m^2')
plt.quiver(xMatrix, yValues, u, v)
```
It is clear that our results match that of the physical and analytical solutions related to this problem. The Prandtl-Meyer expansion wave is shown clearly as the gas expands and speeds up past the expansion corner. In addition to pressure, we can look at the mach number of the flow. The plot below shows that the flow speeds up as it expands past the corner where the wall begins its decline.
```
#plot geometry
plt.figure(figsize=(16,12))
plt.plot(x,np.ones(200)*40,'black')
plt.plot(x,40-h,'black')
plt.ylim([-15,50])
plt.xlim([0,65])
plt.grid()
plt.title('2D Supersonic Flow: Ramp Angle = 5.352 deg')
plt.xlabel('x')
plt.ylabel('y')
# Plot Results
plt.contourf(xMatrix, yValues, mach,cmap=plt.cm.seismic, alpha=0.8)
plt.colorbar(label='Mach Number')
```
| github_jupyter |
## Training modules overview
```
from fastai.basic_train import *
from fastai.gen_doc.nbdoc import *
from fastai.vision import *
from fastai.callbacks import *
```
The fastai library is structured training around a [`Learner`](/basic_train.html#Learner) object that binds together a pytorch model, some data with an optimizer and a loss function, which then will allow us to launch training.
[`basic_train`](/basic_train.html#basic_train) contains the definition of this [`Learner`](/basic_train.html#Learner) class along with the wrapper around pytorch optimizer that the library uses. It defines the basic training loop that is used each time you call the [`fit`](/basic_train.html#fit) function in fastai (or one of its variants). This training loop is kept to the minimum number of instructions, and most of its customization happens in [`Callback`](/callback.html#Callback) objects.
[`callback`](/callback.html#callback) contains the definition of those, as well as the [`CallbackHandler`](/callback.html#CallbackHandler) that is responsible for the communication between the training loop and the [`Callback`](/callback.html#Callback) functions. It maintains a state dictionary to be able to provide to each [`Callback`](/callback.html#Callback) all the informations of the training loop, easily allowing any tweaks you could think of.
In [`callbacks`](/callbacks.html#callbacks), each [`Callback`](/callback.html#Callback) is then implemented in separate modules. Some deal with scheduling the hyperparameters, like [`callbacks.one_cycle`](/callbacks.one_cycle.html#callbacks.one_cycle), [`callbacks.lr_finder`](/callbacks.lr_finder.html#callbacks.lr_finder) or [`callback.general_sched`](/callbacks.general_sched.html#callbacks.general_sched). Others allow special kind of trainings like [`callbacks.fp16`](/callbacks.fp16.html#callbacks.fp16) (mixed precision) or [`callbacks.rnn`](/callbacks.rnn.html#callbacks.rnn). The [`Recorder`](/basic_train.html#Recorder) or [`callbacks.hooks`](/callbacks.hooks.html#callbacks.hooks) are useful to save some internal data.
[`train`](/train.html#train) then implements those callbacks with useful helper functions. Lastly [`metrics`](/metrics.html#metrics) contains all the functions you might want to call to evaluate your results.
## Walk-through of key functionality
We'll do a quick overview of the key pieces of fastai's training modules. See the separate module docs for details on each. We'll use the classic MNIST dataset for the training documentation, cut down to just 3's and 7's. To minimize the boilerplate in our docs we've defined a function to grab the data from <code>URLs.MNIST_SAMPLE</code> which will automatically download and unzip if not already done function, then we put it in an [`ImageDataBunch`](/vision.data.html#ImageDataBunch).
```
path = untar_data(URLs.MNIST_SAMPLE)
data = ImageDataBunch.from_folder(path)
```
### Basic training with [`Learner`](/basic_train.html#Learner)
We can create minimal simple CNNs using [`simple_cnn`](/layers.html#simple_cnn) (see [`models`](/vision.models.html#vision.models) for details on creating models):
```
model = simple_cnn((3,16,16,2))
```
The most important object for training models is [`Learner`](/basic_train.html#Learner), which needs to know, at minimum, what data to train with and what model to train.
```
learn = Learner(data, model)
```
That's enough to train a model, which is done using [`fit`](/basic_train.html#fit). If you have a CUDA-capable GPU it will be used automatically. You have to say how many epochs to train for.
```
learn.fit(1)
```
### Viewing metrics
To see how our training is going, we can request that it reports various [`metrics`](/metrics.html#metrics) after each epoch. You can pass it to the constructor, or set it later. Note that metrics are always calculated on the validation set.
```
learn.metrics=[accuracy]
learn.fit(1)
```
### Extending training with callbacks
You can use [`callback`](/callback.html#callback)s to modify training in almost any way you can imagine. For instance, we've provided a callback to implement Leslie Smith's 1cycle training method.
```
cb = OneCycleScheduler(learn, lr_max=0.01)
learn.fit(1, callbacks=cb)
```
The [`Recorder`](/basic_train.html#Recorder) callback is automatically added for you, and you can use it to see what happened in your training, e.g.:
```
learn.recorder.plot_lr(show_moms=True)
```
### Extending [`Learner`](/basic_train.html#Learner) with [`train`](/train.html#train)
Many of the callbacks can be used more easily by taking advantage of the [`Learner`](/basic_train.html#Learner) extensions in [`train`](/train.html#train). For instance, instead of creating OneCycleScheduler manually as above, you can simply call [`Learner.fit_one_cycle`](/train.html#fit_one_cycle):
```
learn.fit_one_cycle(1)
```
### Applications
Note that if you're training a model for one of our supported *applications*, there's a lot of help available to you in the application modules:
- [`vision`](/vision.html#vision)
- [`text`](/text.html#text)
- [`tabular`](/tabular.html#tabular)
- [`collab`](/collab.html#collab)
For instance, let's use [`create_cnn`](/vision.learner.html#create_cnn) (from [`vision`](/vision.html#vision)) to quickly fine-tune a pre-trained Imagenet model for MNIST (not a very practical approach, of course, since MNIST is handwriting and our model is pre-trained on photos!).
```
learn = create_cnn(data, models.resnet18, metrics=accuracy)
learn.fit_one_cycle(1)
```
| github_jupyter |
# Deploying Punctuation and Capitalization Model in JARVIS
[Transfer Learning Toolkit (TLT)](https://developer.nvidia.com/transfer-learning-toolkit) provides the capability to export your model in a format that can deployed using Nvidia [Jarvis](https://developer.nvidia.com/nvidia-jarvis), a highly performant application framework for multi-modal conversational AI services using GPUs.
This tutorial explores taking an .ejrvs model, the result of `tlt punctuation_and_capitalization export` command, and leveraging the Jarvis ServiceMaker framework to aggregate all the necessary artifacts for Jarvis deployment to a target environment. Once the model is deployed in Jarvis, you can issue inference requests to the server. We will demonstrate how quick and straightforward this whole process is.
## Learning Objectives
In this notebook, you will learn how to:
- Use Jarvis ServiceMaker to take a TLT exported .ejrvs and convert it to .jmir
- Deploy the model(s) locally on the Jarvis Server
- Send inference requests from a demo client using Jarvis API bindings.
## Prerequisites
Before going through the jupyter notebook, please make sure:
- You have access to NVIDIA NGC, and are able to download the Jarvis Quickstart [resources](https://ngc.nvidia.com/resources/ea-jarvis-stage:jarvis_quickstart/)
- Have an .ejrvs model file that you wish to deploy. You can obtain this from ``tlt <task> export`` (with ``export_format=JARVIS``).
<b>NOTE:</b> Please refer to the tutorial on *Punctuation And Capitalization using Transfer Learning Toolkit* for more details on training and exporting an .ejrvs model for punctuation and capitalization task.
## Jarvis ServiceMaker
Servicemaker is the set of tools that aggregates all the necessary artifacts (models, files, configurations, and user settings) for Jarvis deployment to a target environment. It has two main components as shown below:
### 1. Jarvis-build
This step helps build a Jarvis-ready version of the model. It’s only output is an intermediate format (called a JMIR) of an end to end pipeline for the supported services within Jarvis. We are taking a ASR QuartzNet Model in consideration<br>
`jarvis-build` is responsible for the combination of one or more exported models (.ejrvs files) into a single file containing an intermediate format called Jarvis Model Intermediate Representation (.jmir). This file contains a deployment-agnostic specification of the whole end-to-end pipeline along with all the assets required for the final deployment and inference. Please checkout the [documentation](http://docs.jarvis-ai.nvidia.com/release-1-0/service-nlp.html) to find out more.
```
# IMPORTANT: Set the following variables
# ServiceMaker Docker
JARVIS_SM_CONTAINER = "<Jarvis_Servicemaker_Image>"
# Directory where the .ejrvs model is stored $MODEL_LOC/*.ejrvs
MODEL_LOC = "<path_to_model_directory>"
# Name of the .erjvs file
MODEL_NAME = "<add model name>"
# Use the same key that .ejrvs model is encrypted with
KEY = "<add encryption key used for trained model>"
# Pull the ServiceMaker Image
!docker pull $JARVIS_SM_CONTAINER
# Syntax: jarvis-build <task-name> output-dir-for-jmir/model.jmir:key dir-for-ejrvs/model.ejrvs:key
!docker run --rm --gpus all -v $MODEL_LOC:/data $JARVIS_SM_CONTAINER -- \
jarvis-build punctuation -f /data/punct-capit.jmir:$KEY /data/$MODEL_NAME:$KEY
```
`NOTE:` Above, punct-capit-model.ejrvs is the punctuation and capitalization model obtained from `tlt punctuation_and_capitalization export`
### 2. Jarvis-deploy
The deployment tool takes as input one or more Jarvis Model Intermediate Representation (JMIR) files and a target model repository directory. It creates an ensemble configuration specifying the pipeline for the execution and finally writes all those assets to the output model repository directory.
```
# Syntax: jarvis-deploy -f dir-for-jmir/model.jmir:key output-dir-for-repository
!docker run --rm --gpus all -v $MODEL_LOC:/data $JARVIS_SM_CONTAINER -- \
jarvis-deploy -f /data/punct-capit.jmir:$KEY /data/models
```
## Start Jarvis Server
Once the model repository is generated, we are ready to start the Jarvis server. From this step onwards you need to download the Jarvis QuickStart Resource from NGC.
```
### Set the path to Jarvis directory
JARVIS_DIR = <path_to_jarvis_quickstart>
```
Next, we modify ``config.sh`` to enable relevant Jarvis services (nlp for Punctuation & Capitalization model), provide the encryption key, and path to the model repository (``jarvis_model_loc``) generated in the previous step among other configurations.
Pretrained versions of models specified in models_asr/nlp/tts are fetched from NGC. Since we are using our custom model, we can comment it in models_nlp (and any others that are not relevant to our use case).
### config.sh snippet
```
# Enable or Disable Jarvis Services
service_enabled_asr=false ## MAKE CHANGES HERE
service_enabled_nlp=true ## MAKE CHANGES HERE
service_enabled_tts=false ## MAKE CHANGES HERE
# Specify one or more GPUs to use
# specifying more than one GPU is currently an experimental feature, and may result in undefined behaviours.
gpus_to_use="device=0"
# Specify the encryption key to use to deploy models
MODEL_DEPLOY_KEY="tlt_encode" ## MAKE CHANGES HERE
# Locations to use for storing models artifacts
#
# If an absolute path is specified, the data will be written to that location
# Otherwise, a docker volume will be used (default).
#
# jarvis_init.sh will create a `jmir` and `models` directory in the volume or
# path specified.
#
# JMIR ($jarvis_model_loc/jmir)
# Jarvis uses an intermediate representation (JMIR) for models
# that are ready to deploy but not yet fully optimized for deployment. Pretrained
# versions can be obtained from NGC (by specifying NGC models below) and will be
# downloaded to $jarvis_model_loc/jmir by `jarvis_init.sh`
#
# Custom models produced by NeMo or TLT and prepared using jarvis-build
# may also be copied manually to this location $(jarvis_model_loc/jmir).
#
# Models ($jarvis_model_loc/models)
# During the jarvis_init process, the JMIR files in $jarvis_model_loc/jmir
# are inspected and optimized for deployment. The optimized versions are
# stored in $jarvis_model_loc/models. The jarvis server exclusively uses these
# optimized versions.
jarvis_model_loc="<add path>" ## MAKE CHANGES HERE (Replace with MODEL_LOC)
```
```
# Ensure you have permission to execute these scripts.
!cd $JARVIS_DIR && chmod +x ./jarvis_init.sh && chmod +x ./jarvis_start.sh
# Run Jarvis Init. This will fetch the containers/models
# YOU CAN SKIP THIS STEP IF YOU DID JARVIS DEPLOY
!cd $JARVIS_DIR && ./jarvis_init.sh config.sh
# Run Jarvis Start. This will deploy your model(s).
!cd $JARVIS_DIR && ./jarvis_start.sh config.sh
```
## Run Inference
Once the Jarvis server is up and running with your models, you can send inference requests querying the server.
To send GRPC requests, you can install Jarvis Python API bindings for client. This is available as a pip .whl with the QuickStart.
```
# IMPORTANT: Set the name of the whl file
JARVIS_API_WHL = "<add jarvis api .whl file name>"
# Install client API bindings
!cd $JARVIS_DIR && pip install $JARVIS_API_WHL
```
Run the following sample code from within the client docker container:
```
import grpc
import argparse
import os
import jarvis_api.jarvis_nlp_core_pb2 as jcnlp
import jarvis_api.jarvis_nlp_core_pb2_grpc as jcnlp_srv
import jarvis_api.jarvis_nlp_pb2 as jnlp
import jarvis_api.jarvis_nlp_pb2_grpc as jnlp_srv
class BertPunctuatorClient(object):
def __init__(self, grpc_server, model_name="jarvis_punctuation"):
# generate the correct model based on precision and whether or not ensemble is used
print("Using model: {}".format(model_name))
self.model_name = model_name
self.channel = grpc.insecure_channel(grpc_server)
self.jarvis_nlp = jcnlp_srv.JarvisCoreNLPStub(self.channel)
self.has_bos = True
self.has_eos = False
def run(self, input_strings):
if isinstance(input_strings, str):
# user probably passed a single string instead of a list/iterable
input_strings = [input_strings]
request = jcnlp.TextTransformRequest()
request.model.model_name = self.model_name
for q in input_strings:
request.text.append(q)
response = self.jarvis_nlp.TransformText(request)
return response.text[0]
def run_punct_capit(server,model,query):
print("Client app to test punctuation and capitalization on Jarvis")
client = BertPunctuatorClient(server, model_name=model)
result = client.run(query)
print(result)
run_punct_capit(server="localhost:50051",
model="jarvis_punctuation",
query="how are you doing")
```
You can stop all docker container before shutting down the jupyter kernel.
```
!docker stop $(docker ps -a -q)
```
| github_jupyter |
# Artificial Intelligence Nanodegree
## Machine Translation Project
In this notebook, sections that end with **'(IMPLEMENTATION)'** in the header indicate that the following blocks of code will require additional functionality which you must provide. Please be sure to read the instructions carefully!
## Introduction
In this notebook, you will build a deep neural network that functions as part of an end-to-end machine translation pipeline. Your completed pipeline will accept English text as input and return the French translation.
- **Preprocess** - You'll convert text to sequence of integers.
- **Models** Create models which accepts a sequence of integers as input and returns a probability distribution over possible translations. After learning about the basic types of neural networks that are often used for machine translation, you will engage in your own investigations, to design your own model!
- **Prediction** Run the model on English text.
```
%load_ext autoreload
%aimport helper, tests
%autoreload 1
import collections
import helper
import numpy as np
import project_tests as tests
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
from keras.models import Model
from keras.layers import CuDNNGRU, Input, Dense, TimeDistributed, Activation, RepeatVector, Bidirectional
from keras.layers.embeddings import Embedding
from keras.optimizers import Adam
from keras.losses import sparse_categorical_crossentropy
```
### Verify access to the GPU
The following test applies only if you expect to be using a GPU, e.g., while running in a Udacity Workspace or using an AWS instance with GPU support. Run the next cell, and verify that the device_type is "GPU".
- If the device is not GPU & you are running from a Udacity Workspace, then save your workspace with the icon at the top, then click "enable" at the bottom of the workspace.
- If the device is not GPU & you are running from an AWS instance, then refer to the cloud computing instructions in the classroom to verify your setup steps.
```
from tensorflow.python.client import device_lib
print(device_lib.list_local_devices())
```
## Dataset
We begin by investigating the dataset that will be used to train and evaluate your pipeline. The most common datasets used for machine translation are from [WMT](http://www.statmt.org/). However, that will take a long time to train a neural network on. We'll be using a dataset we created for this project that contains a small vocabulary. You'll be able to train your model in a reasonable time with this dataset.
### Load Data
The data is located in `data/small_vocab_en` and `data/small_vocab_fr`. The `small_vocab_en` file contains English sentences with their French translations in the `small_vocab_fr` file. Load the English and French data from these files from running the cell below.
```
# Load English data
english_sentences = helper.load_data('data/small_vocab_en')
# Load French data
french_sentences = helper.load_data('data/small_vocab_fr')
print('Dataset Loaded')
```
### Files
Each line in `small_vocab_en` contains an English sentence with the respective translation in each line of `small_vocab_fr`. View the first two lines from each file.
```
for sample_i in range(2):
print('small_vocab_en Line {}: {}'.format(sample_i + 1, english_sentences[sample_i]))
print('small_vocab_fr Line {}: {}'.format(sample_i + 1, french_sentences[sample_i]))
```
From looking at the sentences, you can see they have been preprocessed already. The puncuations have been delimited using spaces. All the text have been converted to lowercase. This should save you some time, but the text requires more preprocessing.
### Vocabulary
The complexity of the problem is determined by the complexity of the vocabulary. A more complex vocabulary is a more complex problem. Let's look at the complexity of the dataset we'll be working with.
```
english_words_counter = collections.Counter([word for sentence in english_sentences for word in sentence.split()])
french_words_counter = collections.Counter([word for sentence in french_sentences for word in sentence.split()])
print('{} English words.'.format(len([word for sentence in english_sentences for word in sentence.split()])))
print('{} unique English words.'.format(len(english_words_counter)))
print('10 Most common words in the English dataset:')
print('"' + '" "'.join(list(zip(*english_words_counter.most_common(10)))[0]) + '"')
print()
print('{} French words.'.format(len([word for sentence in french_sentences for word in sentence.split()])))
print('{} unique French words.'.format(len(french_words_counter)))
print('10 Most common words in the French dataset:')
print('"' + '" "'.join(list(zip(*french_words_counter.most_common(10)))[0]) + '"')
```
For comparison, _Alice's Adventures in Wonderland_ contains 2,766 unique words of a total of 15,500 words.
## Preprocess
For this project, you won't use text data as input to your model. Instead, you'll convert the text into sequences of integers using the following preprocess methods:
1. Tokenize the words into ids
2. Add padding to make all the sequences the same length.
Time to start preprocessing the data...
### Tokenize (IMPLEMENTATION)
For a neural network to predict on text data, it first has to be turned into data it can understand. Text data like "dog" is a sequence of ASCII character encodings. Since a neural network is a series of multiplication and addition operations, the input data needs to be number(s).
We can turn each character into a number or each word into a number. These are called character and word ids, respectively. Character ids are used for character level models that generate text predictions for each character. A word level model uses word ids that generate text predictions for each word. Word level models tend to learn better, since they are lower in complexity, so we'll use those.
Turn each sentence into a sequence of words ids using Keras's [`Tokenizer`](https://keras.io/preprocessing/text/#tokenizer) function. Use this function to tokenize `english_sentences` and `french_sentences` in the cell below.
Running the cell will run `tokenize` on sample data and show output for debugging.
```
def tokenize(x):
"""
Tokenize x
:param x: List of sentences/strings to be tokenized
:return: Tuple of (tokenized x data, tokenizer used to tokenize x)
"""
# TODO: Implement
tokenizer = Tokenizer()
tokenizer.fit_on_texts(x)
return tokenizer.texts_to_sequences(x), tokenizer
tests.test_tokenize(tokenize)
# Tokenize Example output
text_sentences = [
'The quick brown fox jumps over the lazy dog .',
'By Jove , my quick study of lexicography won a prize .',
'This is a short sentence .']
text_tokenized, text_tokenizer = tokenize(text_sentences)
print(text_tokenizer.word_index)
print()
for sample_i, (sent, token_sent) in enumerate(zip(text_sentences, text_tokenized)):
print('Sequence {} in x'.format(sample_i + 1))
print(' Input: {}'.format(sent))
print(' Output: {}'.format(token_sent))
```
### Padding (IMPLEMENTATION)
When batching the sequence of word ids together, each sequence needs to be the same length. Since sentences are dynamic in length, we can add padding to the end of the sequences to make them the same length.
Make sure all the English sequences have the same length and all the French sequences have the same length by adding padding to the **end** of each sequence using Keras's [`pad_sequences`](https://keras.io/preprocessing/sequence/#pad_sequences) function.
```
def pad(x, length=None):
"""
Pad x
:param x: List of sequences.
:param length: Length to pad the sequence to. If None, use length of longest sequence in x.
:return: Padded numpy array of sequences
"""
# TODO: Implement
return pad_sequences(x, maxlen=length, padding='post')
tests.test_pad(pad)
# Pad Tokenized output
test_pad = pad(text_tokenized)
for sample_i, (token_sent, pad_sent) in enumerate(zip(text_tokenized, test_pad)):
print('Sequence {} in x'.format(sample_i + 1))
print(' Input: {}'.format(np.array(token_sent)))
print(' Output: {}'.format(pad_sent))
```
### Preprocess Pipeline
Your focus for this project is to build neural network architecture, so we won't ask you to create a preprocess pipeline. Instead, we've provided you with the implementation of the `preprocess` function.
```
def preprocess(x, y):
"""
Preprocess x and y
:param x: Feature List of sentences
:param y: Label List of sentences
:return: Tuple of (Preprocessed x, Preprocessed y, x tokenizer, y tokenizer)
"""
preprocess_x, x_tk = tokenize(x)
preprocess_y, y_tk = tokenize(y)
preprocess_x = pad(preprocess_x)
preprocess_y = pad(preprocess_y)
# Keras's sparse_categorical_crossentropy function requires the labels to be in 3 dimensions
preprocess_y = preprocess_y.reshape(*preprocess_y.shape, 1)
return preprocess_x, preprocess_y, x_tk, y_tk
preproc_english_sentences, preproc_french_sentences, english_tokenizer, french_tokenizer =\
preprocess(english_sentences, french_sentences)
max_english_sequence_length = preproc_english_sentences.shape[1]
max_french_sequence_length = preproc_french_sentences.shape[1]
english_vocab_size = len(english_tokenizer.word_index)
french_vocab_size = len(french_tokenizer.word_index)
print('Data Preprocessed')
print("Max English sentence length:", max_english_sequence_length)
print("Max French sentence length:", max_french_sequence_length)
print("English vocabulary size:", english_vocab_size)
print("French vocabulary size:", french_vocab_size)
```
## Models
In this section, you will experiment with various neural network architectures.
You will begin by training four relatively simple architectures.
- Model 1 is a simple RNN
- Model 2 is a RNN with Embedding
- Model 3 is a Bidirectional RNN
- Model 4 is an optional Encoder-Decoder RNN
After experimenting with the four simple architectures, you will construct a deeper architecture that is designed to outperform all four models.
### Ids Back to Text
The neural network will be translating the input to words ids, which isn't the final form we want. We want the French translation. The function `logits_to_text` will bridge the gab between the logits from the neural network to the French translation. You'll be using this function to better understand the output of the neural network.
```
def logits_to_text(logits, tokenizer):
"""
Turn logits from a neural network into text using the tokenizer
:param logits: Logits from a neural network
:param tokenizer: Keras Tokenizer fit on the labels
:return: String that represents the text of the logits
"""
index_to_words = {id: word for word, id in tokenizer.word_index.items()}
index_to_words[0] = '<PAD>'
return ' '.join([index_to_words[prediction] for prediction in np.argmax(logits, 1)])
print('`logits_to_text` function loaded.')
```
### Model 1: RNN (IMPLEMENTATION)

A basic RNN model is a good baseline for sequence data. In this model, you'll build a RNN that translates English to French.
```
def simple_model(input_shape, output_sequence_length, english_vocab_size, french_vocab_size):
"""
Build and train a basic RNN on x and y
:param input_shape: Tuple of input shape
:param output_sequence_length: Length of output sequence
:param english_vocab_size: Number of unique English words in the dataset
:param french_vocab_size: Number of unique French words in the dataset
:return: Keras model built, but not trained
"""
# TODO: Build the layers
inputs = Input(input_shape[1:])
x = CuDNNGRU(128, return_sequences=True)(inputs)
predictions = TimeDistributed(Dense(french_vocab_size, activation='softmax'))(x)
model = Model(inputs=inputs, outputs=predictions)
model.compile(loss=sparse_categorical_crossentropy,
optimizer=Adam(),
metrics=['accuracy'])
return model
tests.test_simple_model(simple_model)
# Reshaping the input to work with a basic RNN
tmp_x = pad(preproc_english_sentences, max_french_sequence_length)
tmp_x = tmp_x.reshape((-1, preproc_french_sentences.shape[-2], 1))
# Train the neural network
simple_rnn_model = simple_model(
tmp_x.shape,
max_french_sequence_length,
english_vocab_size,
french_vocab_size)
simple_rnn_model.fit(tmp_x, preproc_french_sentences, batch_size=1024, epochs=10, validation_split=0.2)
# Print prediction(s)
print(logits_to_text(simple_rnn_model.predict(tmp_x[:1])[0], french_tokenizer))
```
### Model 2: Embedding (IMPLEMENTATION)

You've turned the words into ids, but there's a better representation of a word. This is called word embeddings. An embedding is a vector representation of the word that is close to similar words in n-dimensional space, where the n represents the size of the embedding vectors.
In this model, you'll create a RNN model using embedding.
```
def embed_model(input_shape, output_sequence_length, english_vocab_size, french_vocab_size):
"""
Build and train a RNN model using word embedding on x and y
:param input_shape: Tuple of input shape
:param output_sequence_length: Length of output sequence
:param english_vocab_size: Number of unique English words in the dataset
:param french_vocab_size: Number of unique French words in the dataset
:return: Keras model built, but not trained
"""
# TODO: Implement
inputs = Input(input_shape[1:])
x = Embedding(english_vocab_size, 128, input_length=input_shape[1])(inputs)
x = CuDNNGRU(128, return_sequences=True)(x)
predictions = TimeDistributed(Dense(french_vocab_size, activation='softmax'))(x)
model = Model(inputs=inputs, outputs=predictions)
model.compile(loss=sparse_categorical_crossentropy,
optimizer=Adam(),
metrics=['accuracy'])
return model
tests.test_embed_model(embed_model)
# TODO: Reshape the input
tmp_x = pad(preproc_english_sentences, max_french_sequence_length)
# TODO: Train the neural network
embed_rnn_model = embed_model(
tmp_x.shape,
max_french_sequence_length,
english_vocab_size,
french_vocab_size)
embed_rnn_model.fit(tmp_x, preproc_french_sentences, batch_size=1024, epochs=10, validation_split=0.2)
# TODO: Print prediction(s)
print(logits_to_text(embed_rnn_model.predict(tmp_x[:1])[0], french_tokenizer))
```
### Model 3: Bidirectional RNNs (IMPLEMENTATION)

One restriction of a RNN is that it can't see the future input, only the past. This is where bidirectional recurrent neural networks come in. They are able to see the future data.
```
def bd_model(input_shape, output_sequence_length, english_vocab_size, french_vocab_size):
"""
Build and train a bidirectional RNN model on x and y
:param input_shape: Tuple of input shape
:param output_sequence_length: Length of output sequence
:param english_vocab_size: Number of unique English words in the dataset
:param french_vocab_size: Number of unique French words in the dataset
:return: Keras model built, but not trained
"""
# TODO: Implement
inputs = Input(input_shape[1:])
x = Bidirectional(CuDNNGRU(128, return_sequences=True))(inputs)
predictions = TimeDistributed(Dense(french_vocab_size, activation='softmax'))(x)
model = Model(inputs=inputs, outputs=predictions)
model.compile(loss=sparse_categorical_crossentropy,
optimizer=Adam(),
metrics=['accuracy'])
return model
tests.test_bd_model(bd_model)
# TODO: Train and Print prediction(s)
tmp_x = pad(preproc_english_sentences, max_french_sequence_length)
tmp_x = tmp_x.reshape((-1, preproc_french_sentences.shape[-2], 1))
# Train the neural network
bd_rnn_model = bd_model(
tmp_x.shape,
max_french_sequence_length,
english_vocab_size,
french_vocab_size)
bd_rnn_model.fit(tmp_x, preproc_french_sentences, batch_size=1024, epochs=10, validation_split=0.2)
# Print prediction(s)
print(logits_to_text(bd_rnn_model.predict(tmp_x[:1])[0], french_tokenizer))
```
### Model 4: Encoder-Decoder (OPTIONAL)
Time to look at encoder-decoder models. This model is made up of an encoder and decoder. The encoder creates a matrix representation of the sentence. The decoder takes this matrix as input and predicts the translation as output.
Create an encoder-decoder model in the cell below.
```
def encdec_model(input_shape, output_sequence_length, english_vocab_size, french_vocab_size):
"""
Build and train an encoder-decoder model on x and y
:param input_shape: Tuple of input shape
:param output_sequence_length: Length of output sequence
:param english_vocab_size: Number of unique English words in the dataset
:param french_vocab_size: Number of unique French words in the dataset
:return: Keras model built, but not trained
"""
# OPTIONAL: Implement
# encoder input model
inputs = Input(input_shape[1:])
x = CuDNNGRU(128)(inputs)
x = RepeatVector(output_sequence_length)(x)
# decoder output model
x = CuDNNGRU(128, return_sequences=True)(x)
predictions = TimeDistributed(Dense(french_vocab_size, activation='softmax'))(x)
model = Model(inputs=inputs, outputs=predictions)
model.compile(loss=sparse_categorical_crossentropy,
optimizer=Adam(),
metrics=['accuracy'])
return model
tests.test_encdec_model(encdec_model)
# OPTIONAL: Train and Print prediction(s)
tmp_x = np.expand_dims(preproc_english_sentences,-1)
# Train the neural network
encdec_rnn_model = encdec_model(
tmp_x.shape,
max_french_sequence_length,
english_vocab_size,
french_vocab_size)
encdec_rnn_model.fit(tmp_x, preproc_french_sentences, batch_size=1024, epochs=10, validation_split=0.2)
# Print prediction(s)
print(logits_to_text(encdec_rnn_model.predict(tmp_x[:1])[0], french_tokenizer))
```
### Model 5: Custom (IMPLEMENTATION)
Use everything you learned from the previous models to create a model that incorporates embedding and a bidirectional rnn into one model.
```
def model_final(input_shape, output_sequence_length, english_vocab_size, french_vocab_size):
"""
Build and train a model that incorporates embedding, encoder-decoder, and bidirectional RNN on x and y
:param input_shape: Tuple of input shape
:param output_sequence_length: Length of output sequence
:param english_vocab_size: Number of unique English words in the dataset
:param french_vocab_size: Number of unique French words in the dataset
:return: Keras model built, but not trained
"""
# TODO: Implement
# encoder input model
inputs = Input(input_shape[1:])
x = Embedding(english_vocab_size, 128, input_length=input_shape[1])(inputs)
x = Bidirectional(CuDNNGRU(128))(x)
x = RepeatVector(output_sequence_length)(x)
# decoder output model
x = CuDNNGRU(128, return_sequences=True)(x)
predictions = TimeDistributed(Dense(french_vocab_size, activation='softmax'))(x)
model = Model(inputs=inputs, outputs=predictions)
model.compile(loss=sparse_categorical_crossentropy,
optimizer=Adam(),
metrics=['accuracy'])
return model
tests.test_model_final(model_final)
print('Final Model Loaded')
# TODO: Train the final model
tmp_x = preproc_english_sentences.copy()
# Train the neural network
model_rnn_final = model_final(
tmp_x.shape,
max_french_sequence_length,
english_vocab_size,
french_vocab_size)
model_rnn_final.fit(tmp_x, preproc_french_sentences, batch_size=1024, epochs=10, validation_split=0.2)
# Print prediction(s)
print(logits_to_text(model_rnn_final.predict(tmp_x[:1])[0], french_tokenizer))
```
## Prediction (IMPLEMENTATION)
```
def final_predictions(x, y, x_tk, y_tk):
"""
Gets predictions using the final model
:param x: Preprocessed English data
:param y: Preprocessed French data
:param x_tk: English tokenizer
:param y_tk: French tokenizer
"""
# TODO: Train neural network using model_final
model = model_final(
x.shape,
y.shape[1],
len(x_tk.word_index),
len(y_tk.word_index))
model.fit(x, y, batch_size=1024, epochs=100, validation_split=0.2, verbose=2)
## DON'T EDIT ANYTHING BELOW THIS LINE
y_id_to_word = {value: key for key, value in y_tk.word_index.items()}
y_id_to_word[0] = '<PAD>'
sentence = 'he saw a old yellow truck'
sentence = [x_tk.word_index[word] for word in sentence.split()]
sentence = pad_sequences([sentence], maxlen=x.shape[-1], padding='post')
sentences = np.array([sentence[0], x[0]])
predictions = model.predict(sentences, len(sentences))
print('Sample 1:')
print(' '.join([y_id_to_word[np.argmax(x)] for x in predictions[0]]))
print('Il a vu un vieux camion jaune')
print('Sample 2:')
print(' '.join([y_id_to_word[np.argmax(x)] for x in predictions[1]]))
print(' '.join([y_id_to_word[np.max(x)] for x in y[0]]))
final_predictions(preproc_english_sentences, preproc_french_sentences, english_tokenizer, french_tokenizer)
```
## Submission
When you're ready to submit, complete the following steps:
1. Review the [rubric](https://review.udacity.com/#!/rubrics/1004/view) to ensure your submission meets all requirements to pass
2. Generate an HTML version of this notebook
- Run the next cell to attempt automatic generation (this is the recommended method in Workspaces)
- Navigate to **FILE -> Download as -> HTML (.html)**
- Manually generate a copy using `nbconvert` from your shell terminal
```
$ pip install nbconvert
$ python -m nbconvert machine_translation.ipynb
```
3. Submit the project
- If you are in a Workspace, simply click the "Submit Project" button (bottom towards the right)
- Otherwise, add the following files into a zip archive and submit them
- `helper.py`
- `machine_translation.ipynb`
- `machine_translation.html`
- You can export the notebook by navigating to **File -> Download as -> HTML (.html)**.
```
!!python -m nbconvert *.ipynb
```
## Optional Enhancements
This project focuses on learning various network architectures for machine translation, but we don't evaluate the models according to best practices by splitting the data into separate test & training sets -- so the model accuracy is overstated. Use the [`sklearn.model_selection.train_test_split()`](http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.train_test_split.html) function to create separate training & test datasets, then retrain each of the models using only the training set and evaluate the prediction accuracy using the hold out test set. Does the "best" model change?
| github_jupyter |
# Compare Country Trajectories - Total Cases
> Comparing how countries trajectories of total cases are similar with Italy, South Korea and Japan
- comments: true
- author: Pratap Vardhan
- categories: [growth, compare, interactive]
- image: images/covid-compare-country-trajectories.png
- permalink: /compare-country-trajectories/
```
#hide
import pandas as pd
import altair as alt
from IPython.display import HTML
#hide
url = ('https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/'
'csse_covid_19_time_series/time_series_19-covid-Confirmed.csv')
df = pd.read_csv(url)
# rename countries
df['Country/Region'] = df['Country/Region'].replace({'Korea, South': 'South Korea'})
dt_cols = df.columns[~df.columns.isin(['Province/State', 'Country/Region', 'Lat', 'Long'])]
#hide
dff = (df.groupby('Country/Region')[dt_cols].sum()
.stack().reset_index(name='Confirmed Cases')
.rename(columns={'level_1': 'Date', 'Country/Region': 'Country'}))
dff['Date'] = pd.to_datetime(dff['Date'], format='%m/%d/%y')
#hide
MIN_CASES = 700
LAST_DATE = dt_cols[-1]
# sometimes last column may be empty, then go backwards
for c in dt_cols[::-1]:
if not df[c].fillna(0).eq(0).all():
LAST_DATE = c
break
countries = dff[dff['Date'].eq(LAST_DATE) & dff['Confirmed Cases'].ge(MIN_CASES) &
dff['Country'].ne('China')
].sort_values(by='Confirmed Cases', ascending=False)
countries = countries['Country'].values
#hide
SINCE_CASES_NUM = 100
dff2 = dff[dff['Country'].isin(countries)].copy()
days_since = (dff2.assign(F=dff2['Confirmed Cases'].ge(SINCE_CASES_NUM))
.set_index('Date')
.groupby('Country')['F'].transform('idxmax'))
dff2['Days since 100 cases'] = (dff2['Date'] - days_since.values).dt.days.values
dff2 = dff2[dff2['Days since 100 cases'].ge(0)]
#hide
def get_country_colors(x):
mapping = {
'Italy': 'black',
'Iran': '#A1BA59',
'South Korea': '#E45756',
'Spain': '#F58518',
'Germany': '#9D755D',
'France': '#F58518',
'US': '#2495D3',
'Switzerland': '#9D755D',
'Norway': '#C1B7AD',
'United Kingdom': '#2495D3',
'Netherlands': '#C1B7AD',
'Sweden': '#C1B7AD',
'Belgium': '#C1B7AD',
'Denmark': '#C1B7AD',
'Austria': '#C1B7AD',
'Japan': '#9467bd'}
return mapping.get(x, '#C1B7AD')
#hide_input
baseline_countries = ['Italy', 'South Korea', 'Japan']
max_date = dff2['Date'].max()
color_domain = list(dff2['Country'].unique())
color_range = list(map(get_country_colors, color_domain))
def make_since_chart(highlight_countries=[], baseline_countries=baseline_countries):
selection = alt.selection_multi(fields=['Country'], bind='legend',
init=[{'Country': x} for x in highlight_countries + baseline_countries])
base = alt.Chart(dff2, width=550).encode(
x='Days since 100 cases:Q',
y=alt.Y('Confirmed Cases:Q', scale=alt.Scale(type='log')),
color=alt.Color('Country:N', scale=alt.Scale(domain=color_domain, range=color_range)),
tooltip=list(dff2),
opacity=alt.condition(selection, alt.value(1), alt.value(0.05))
)
max_day = dff2['Days since 100 cases'].max()
ref = pd.DataFrame([[x, 100*1.33**x] for x in range(max_day+1)], columns=['Days since 100 cases', 'Confirmed Cases'])
base_ref = alt.Chart(ref).encode(x='Days since 100 cases:Q', y='Confirmed Cases:Q')
return (
base_ref.mark_line(color='black', opacity=.5, strokeDash=[3,3]) +
base_ref.transform_filter(
alt.datum['Days since 100 cases'] >= max_day
).mark_text(dy=-6, align='right', fontSize=10, text='33% Daily Growth') +
base.mark_line(point=True).add_selection(selection) +
base.transform_filter(
alt.datum['Date'] >= int(max_date.timestamp() * 1000)
).mark_text(dy=-8, align='right', fontWeight='bold').encode(text='Country:N')
).properties(
title=f"Compare {', '.join(highlight_countries)} trajectory with {', '.join(baseline_countries)}"
)
```
## Learning from Italy, South Korea & Japan
Italy, South Korea & Japan are three countries which show different growth rates and how it evolved over time.
**South Korea** flattened it's growth after 2 weeks since 100 cases. **Italy** continue to grew after 3rd week.
Where does your Country stand today?
<small>Click (Shift+ for multiple) on Countries legend to filter the visualization.</small>
```
#hide_input
HTML(f'<small class="float-right">Last Updated on {pd.to_datetime(LAST_DATE).strftime("%B, %d %Y")}</small>')
#hide_input
chart = make_since_chart()
chart
#hide_input
chart2 = make_since_chart(['Spain', 'Germany'])
chart2
```
{{LAST_DATE}}
```
#hide_input
chart3 = make_since_chart(['US', 'France'])
chart3
#hide_input
chart4 = make_since_chart(['Germany', 'United Kingdom'])
chart4
```
Select a country from the drop down list below to toggle the visualization.
```
#hide_input
base = alt.Chart(dff2, width=600).encode(
x='Days since 100 cases:Q',
y=alt.Y('Confirmed Cases:Q', scale=alt.Scale(type='log')),
color=alt.Color('Country:N', scale=alt.Scale(domain=color_domain, range=color_range), legend=None),
tooltip=['Country', 'Date', 'Confirmed Cases', 'Days since 100 cases']
)
country_selection = alt.selection_single(
name='Select', fields=['Country'],
bind=alt.binding_select(options=list(sorted(set(countries) - set(baseline_countries)))),
init={'Country': 'US'})
date_filter = alt.datum['Date'] >= int(max_date.timestamp() * 1000)
base2 = base.transform_filter(alt.FieldOneOfPredicate('Country', baseline_countries))
base3 = base.transform_filter(country_selection)
base4 = base3.transform_filter(date_filter)
max_day = dff2['Days since 100 cases'].max()
ref = pd.DataFrame([[x, 100*1.33**x] for x in range(max_day+1)], columns=['Days since 100 cases', 'Confirmed Cases'])
base_ref = alt.Chart(ref).encode(x='Days since 100 cases:Q', y='Confirmed Cases:Q')
base_ref_f = base_ref.transform_filter(alt.datum['Days since 100 cases'] >= max_day)
chart5 = (
base_ref.mark_line(color='black', opacity=.5, strokeDash=[3,3]) +
base_ref_f.mark_text(dy=-6, align='right', fontSize=10, text='33% Daily Growth') +
base2.mark_line(point=True, tooltip=True) +
base3.mark_line(point={'size':50}, tooltip=True) +
base2.transform_filter(date_filter).mark_text(dy=-8, align='right').encode(text='Country:N') +
base4.mark_text(dx=8, align='left', fontWeight='bold').encode(text='Country:N') +
base4.mark_text(dx=8, dy=12, align='left', fontWeight='bold').encode(text='Confirmed Cases:Q')
).add_selection(country_selection).properties(
title=f"Country's Trajectory compared to {', '.join(baseline_countries)}"
)
chart5
```
Interactive by [Pratap Vardhan](https://twitter.com/PratapVardhan)[^1]
[^1]: Source: ["2019 Novel Coronavirus COVID-19 (2019-nCoV) Data Repository by Johns Hopkins CSSE"](https://systems.jhu.edu/research/public-health/ncov/) [GitHub repository](https://github.com/CSSEGISandData/COVID-19). Link to [original notebook](https://github.com/pratapvardhan/notebooks/blob/master/covid19/covid19-compare-country-trajectories.ipynb).
| github_jupyter |
```
import numpy as np
import pandas as pd
import math
import sklearn
import sklearn.preprocessing
import datetime
import os
import matplotlib.pyplot as plt
import tensorflow as tf
import time
from sklearn.preprocessing import StandardScaler
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_squared_error, r2_score
# split data in 80%/10%/10% train/validation/test sets
valid_set_size_percentage = 10
test_set_size_percentage = 10
import warnings
warnings.filterwarnings('ignore')
print(tf.__version__)
```
### Model Training
```
num_data_df = pd.read_csv('num_data.csv')
num_data_df.head()
standardScaler=StandardScaler()
num_data_scaled_df = standardScaler.fit_transform(num_data_df)
seq_len = 24
# function to create train, validation, test data given stock data and sequence length
# the training sets are the sequences (20)
# this is the methods of time series prediction
def load_data(data_raw, seq_len):
# data_raw = stock.as_matrix() # convert to numpy array
data = []
# create all possible sequences of length seq_len
for index in range(len(data_raw) - seq_len):
data.append(data_raw[index: index + seq_len])
data = np.array(data);
valid_set_size = int(np.round(valid_set_size_percentage/100*data.shape[0]));
test_set_size = int(np.round(test_set_size_percentage/100*data.shape[0]));
train_set_size = data.shape[0] - (valid_set_size + test_set_size);
x_train = data[:train_set_size,:-1,:]
y_train = data[:train_set_size,-1,:]
x_valid = data[train_set_size:train_set_size+valid_set_size,:-1,:]
y_valid = data[train_set_size:train_set_size+valid_set_size,-1,:]
x_test = data[train_set_size+valid_set_size:,:-1,:]
y_test = data[train_set_size+valid_set_size:,-1,:]
return [x_train, y_train, x_valid, y_valid, x_test, y_test]
x_train, y_train, x_valid, y_valid, x_test, y_test = load_data(num_data_scaled_df, seq_len)
x_train
## Basic Cell RNN in tensorflow
index_in_epoch = 0;
perm_array = np.arange(x_train.shape[0])
np.random.shuffle(perm_array)
# function to get the next batch
def get_next_batch(batch_size):
global index_in_epoch, x_train, perm_array
start = index_in_epoch
index_in_epoch += batch_size
if index_in_epoch > x_train.shape[0]:
np.random.shuffle(perm_array) # shuffle permutation array
start = 0 # start next epoch
index_in_epoch = batch_size
end = index_in_epoch
return x_train[perm_array[start:end]], y_train[perm_array[start:end]]
# parameters
# We can divide the dataset of 2000 examples into batches of 500
# then it will take 4 iterations to complete 1 epoch
n_steps = seq_len-1
n_inputs = 16
n_neurons = 2
n_outputs = 16
n_layers = 2
learning_rate = 0.001
batch_size = 50
n_epochs = 50
train_set_size = x_train.shape[0]
test_set_size = x_test.shape[0]
tf.reset_default_graph()
# feed data into the graph through these placeholders.
X = tf.placeholder(tf.float32, [None, n_steps, n_inputs])
y = tf.placeholder(tf.float32, [None, n_outputs])
# use Basic RNN Cell
rnn_layer = tf.contrib.rnn.BasicRNNCell(num_units=n_neurons, activation=tf.nn.elu)
# use Basic LSTM Cell
lstm_layer = tf.contrib.rnn.BasicLSTMCell(num_units=n_neurons, activation=tf.nn.elu)
# use Basic GRU cell
gru_layer = tf.contrib.rnn.GRUCell(num_units=n_neurons, activation=tf.nn.leaky_relu)
# use LSTM Cell with peephole connections
#layers = [tf.contrib.rnn.LSTMCell(num_units=n_neurons,
# activation=tf.nn.leaky_relu, use_peepholes = True)
# for layer in range(n_layers)]
layers = [tf.contrib.rnn.BasicRNNCell(num_units=n_neurons, activation=tf.nn.elu)
for layer in range(n_layers)]
GRU = tf.contrib.rnn.MultiRNNCell(cells=[gru_layer])
LSTM = tf.contrib.rnn.MultiRNNCell(cells=[lstm_layer])
GRU_GRU = tf.contrib.rnn.MultiRNNCell(cells=[gru_layer,lstm_layer])
GRU_LSTM = tf.contrib.rnn.MultiRNNCell(cells=[gru_layer, lstm_layer])
LSTM_GRU = tf.contrib.rnn.MultiRNNCell(cells=[lstm_layer,gru_layer])
LSTM_LSTM = tf.contrib.rnn.MultiRNNCell(cells=[lstm_layer,lstm_layer])
multi_layer_cell = LSTM_GRU
#Creates a recurrent neural network specified by RNNCell cell
rnn_outputs, states = tf.nn.dynamic_rnn(multi_layer_cell, X, dtype=tf.float32)
stacked_rnn_outputs = tf.reshape(rnn_outputs, [-1, n_neurons])
stacked_outputs = tf.layers.dense(stacked_rnn_outputs, n_outputs) # Functional interface for the densely-connected layer
outputs = tf.reshape(stacked_outputs, [-1, n_steps, n_outputs])
outputs = outputs[:,n_steps-1,:] # keep only last output of sequence
loss = tf.reduce_mean(tf.square(outputs - y)) # loss function = mean squared error
# Instead of adapting the parameter learning rates based on the average first moment (the mean) as in
# RMSProp, Adam also makes use of the average of the second moments of the gradients (the uncentered variance).
optimizer = tf.train.AdamOptimizer(learning_rate=learning_rate)
training_op = optimizer.minimize(loss)
# run graph
start = time.process_time()
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
# Total number of training examples present in a single batch.
for iteration in range(int(n_epochs*train_set_size/batch_size)):
x_batch, y_batch = get_next_batch(batch_size) # fetch the next training batch
sess.run(training_op, feed_dict={X: x_batch, y: y_batch})
if iteration % int(5*train_set_size/batch_size) == 0:
mse_train = loss.eval(feed_dict={X: x_train, y: y_train})
mse_valid = loss.eval(feed_dict={X: x_valid, y: y_valid})
print('%.2f epochs: MSE train/valid = %.6f/%.6f'%(
iteration*batch_size/train_set_size, mse_train, mse_valid))
y_train_pred = sess.run(outputs, feed_dict={X: x_train})
y_valid_pred = sess.run(outputs, feed_dict={X: x_valid})
y_test_pred = sess.run(outputs, feed_dict={X: x_test})
print('time taken for model traning: {} for epoch: {}, n_neurons: {}, batch_size: {}, learning_rate: {}, n_steps: {}'
.format(time.process_time() - start, n_epochs, n_neurons, batch_size, learning_rate, n_steps ))
ft_list = []
for j in range(15):
ft_list.append([j, num_data_df.columns[j]])
print(ft_list)
ft = 4 #4 PM2.5
## show predictions
plt.figure(figsize=(35, 5));
plt.subplot(1,2,1);
plt.plot(np.arange(y_train.shape[0]), y_train[:,ft], color='blue', label='train target')
plt.plot(np.arange(y_train.shape[0], y_train.shape[0]+y_valid.shape[0]), y_valid[:,ft],
color='gray', label='valid target')
plt.plot(np.arange(y_train.shape[0]+y_valid.shape[0],
y_train.shape[0]+y_test.shape[0]+y_test.shape[0]),
y_test[:,ft], color='black', label='test target')
plt.plot(np.arange(y_train_pred.shape[0]),y_train_pred[:,ft], color='red',
label='test prediction')
plt.plot(np.arange(y_train_pred.shape[0], y_train_pred.shape[0]+y_valid_pred.shape[0]),
y_valid_pred[:,ft], color='orange', label='valid prediction')
plt.plot(np.arange(y_train_pred.shape[0]+y_valid_pred.shape[0],
y_train_pred.shape[0]+y_valid_pred.shape[0]+y_test_pred.shape[0]),
y_test_pred[:,ft], color='green', label='test prediction')
plt.title('past and future PM2.5 Level')
plt.xlabel('time [days]')
plt.ylabel('normalized PM2.5 Level')
plt.legend(loc='best');
plt.figure(figsize=(30, 15));
plt.subplot(1,1,1);
plt.plot(np.arange(y_train.shape[0], y_train.shape[0]+y_test.shape[0]),
y_test[:,ft], color='black', label='test target')
plt.plot(np.arange(y_train_pred.shape[0], y_train_pred.shape[0]+y_test_pred.shape[0]),
y_test_pred[:,ft], color='green', label='test prediction')
plt.title('future PM2.5 ')
plt.xlabel('time [days]')
plt.ylabel('normalized PM2.5')
plt.legend(loc='best');
plt.figure(figsize=(30, 15));
plt.subplot(1,1,1);
plt.plot(np.arange(y_train.shape[0], y_train.shape[0]+y_test.shape[0]),
y_test[:,ft], color='black', label='test target')
plt.plot(np.arange(y_train_pred.shape[0], y_train_pred.shape[0]+y_test_pred.shape[0]),
y_test_pred[:,ft], color='green', label='test prediction')
plt.title('future PM2.5 ')
plt.xlabel('time [days]')
plt.ylabel('normalized PM2.5')
plt.legend(loc='best');
from sklearn.metrics import mean_squared_error
from math import sqrt
def RMSE(y_actual, y_predicted):
return sqrt(mean_squared_error(y_actual, y_predicted))
print('mean_squared_error',mean_squared_error(y_train[:,4], y_train_pred[:,4]))
print('r2_score', r2_score(y_train[:,4], y_train_pred[:,4]))
print('RMSE', RMSE(y_train[:,4], y_train_pred[:,4]))
print()
print('mean_squared_error',mean_squared_error(y_valid[:,4], y_valid_pred[:,4]))
print('r2_score', r2_score(y_valid[:,4], y_valid_pred[:,4]))
print('RMSE', RMSE(y_valid[:,4], y_valid_pred[:,4]))
print()
print('mean_squared_error',mean_squared_error(y_test[:,4], y_test_pred[:,4]))
print('r2_score', r2_score(y_test[:,4], y_test_pred[:,4]))
print('RMSE', RMSE(y_test[:,4], y_test_pred[:,4]))
print()
y_test_pred[:,4]
for i in range(len(y_test)):
print(y_test[i, 4], y_test_pred[i,4])
```
### Linear Regression
```
linearRegression=LinearRegression()
linearRegression.fit(X_train,y_train)
y_pred=linearRegression.predict(X_test)
linearRegression.score(X_test, y_test)
n_results=100
fig, ax=plt.subplots(2,1,figsize=(12,8))
ax[0].plot(y_test.values[:n_results], color="red")
ax[1].plot(y_pred[:n_results], color="green")
print('mean_squared_error',mean_squared_error(y_test, y_pred))
print('r2_score', r2_score(y_test, y_pred))
```
| github_jupyter |
### Introduction
After learning about several clustering methods, we are tasked with applying these techniques to the [Boston Marathon dataset](https://github.com/llimllib/bostonmarathon). In this case, I have chosen data from the year 2013. Specifically, we are asked to determine which clustering technique is best for the marathon results and include a writeup on what we have discovered.
Something I didn't know about the Boston Marathon is that it includes wheelchair racers and hand cyclists. This dataset only includes those who ran on their feet and wheelchair racers, but it would be interesting to see if any clustering techniques are able to distinguish between runners, wheelchair racers, and gender, giving a total of 4 clusters.
### 1 - Importing the Data
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
from sklearn.cluster import KMeans, MeanShift, estimate_bandwidth, SpectralClustering
from sklearn.preprocessing import normalize
from sklearn.decomposition import PCA
from sklearn import metrics
df = pd.read_csv('boston_marathon_results.csv')
df.head()
df.info(memory_usage='deep')
df.describe()
```
#### 1.1 - Missing Data
There are 3 features with missing categories: ctz, state, and city. The feature 'ctz' is the citizenship the runner holds. Since 95% of the values are missing, I will remove this feature for this exercise.
```
df.isnull().sum()
```
In the states column, there are missing value because those races were run in a country outside of the United States. In this case, I will impute 'other'.
```
df[df['state'].isnull()]['country'].unique()
df['state'].fillna('Other', inplace = True)
```
Lastly, there is the matter of the missing city. This particular race was ran in Argentina. Most of the races in Argentina were run in Buenos Aires, so I will fill the missing city with Buenos Aires. I did try to search for the city, but failed to find the information.
```
df[df['city'].isnull()]['country']
df[df['country']=='ARG']['city'].value_counts()
df['city'].fillna('Buenos Aires', inplace = True)
```
### 2 - Data Cleaning
#### 2.1 - Creating Ground Truths
I know that when clustering, we don't always have ground truths. However, I can create one for easy comparison between clusters and models. In this case, I will have 4 categories: M, F, WM, and WF, which stands for male, female, wheelchair male, and wheelchair female respectively.
```
def get_ground_truth(gender, bib):
if bib[0] == 'W':
return 'W' + gender
else:
return gender
df['racertype'] = df.apply(lambda x: get_ground_truth(x['gender'], x['bib']), axis = 1)
```
The dataset is pretty unbalanced now, with only ~50 racers in wheelchairs finishing the race vs ~16k runners on foot. This could make it difficult to distinguish clusters. If anything, the official times (and splits) are a big separator between those in wheelchairs and non wheelchair racers. Below in the left chart, we see that the mean official times between wheelchair and non wheelchair racers differ by about 80 minutes, with wheelchair racers averaging ~120 minutes and non wheelchair racers somewhere between 200-220 minutes, depending on gender. On the right chart is a plot of official times vs age. Here we can see that there is a giant cluster for male runners, and inside is a subset of clusters for female runners. The wheelchair racers are more spread out but are clearly separate from the runners. They run into a similar problem of the genders blending together though.
```
df['racertype'].value_counts()
fig, (axis1, axis2) = plt.subplots(1, 2, figsize = (15, 4))
sns.pointplot(x = 'racertype', y = 'official', data = df, linestyles = '', ax = axis1)
sns.scatterplot(x = 'age', y = 'official', hue = 'racertype', data = df, ax = axis2)
axis1.set_title('Mean Official Times by Racer Type')
axis1.set_ylabel('Official Time (min)')
axis1.set_xlabel('Racer Type')
axis2.set_title('Relationship Between Age and Official Times')
axis2.set_ylabel('Official Time (min)')
axis2.set_xlabel('Age')
df.drop('gender', axis = 1, inplace = True)
```
#### 2.2 - Fixing Hidden Missing Values
I was wondering why the splits (i.e. 25k, 10k, etc) are objects instead of floats. Turns out if there are missing values, they are labeled as '-'. These values will be imputed by assuming the runner will have a steady pace over the whole marathon.
```
float_lst = ['25k', '10k', 'half', '30k', '5k', '20k', '35k', '40k']
marathon_distance = 42.19506847 # in km
splits_dict = {'25k': 25/marathon_distance, '10k': 10/marathon_distance, 'half': 0.5,
'30k': 30/marathon_distance, '5k': 5/marathon_distance, '20k': 20/marathon_distance,
'35k': 35/marathon_distance, '40k': 40/marathon_distance}
def get_distance(cols):
for i in range(len(float_lst)):
if cols[i]=='-':
return splits_dict[float_lst[i]] * cols[8]
for col in float_lst:
condition = df[col]=='-'
df.loc[condition, [col]] = df.loc[condition].apply(lambda x: get_distance(x[float_lst + ['official']].values), axis = 1)
df[col] = pd.to_numeric(df[col], downcast = 'float')
```
### 3 - Applying Cluster Techniques
Here, I would like to use 2 clustering techniques: kmeans and mean shift. I would have liked to apply spectral clustering to, but my computer cannot handle it.
#### 3.1 - Prepping the Data
Before applying the clustering techniques, I need to prep the data. There are two things I need to do to prep the data: 1) remove extraneous features, and 2) normalize the data. The features I think are unneeded are names and locations (ctz, country, state, city). I will also remove genderdiv because I'm afraid of target leakage.
```
df.drop(['name', 'genderdiv', 'ctz', 'country', 'state', 'city'], axis = 1, inplace = True)
```
I suspect bib numbers are important, since most of the racers in wheelchairs have numbers going up to the 100, maybe 150 at most. I am concerned however that there is a W in the bib number. There are two choices to deal with the W, either remove it or encode it. For now, I will try removing it to preserve the actual number.
```
def remove_bib_letters(col):
if col[0].isnumeric():
return col
else:
return col[1:]
df['bib'] = df['bib'].apply(lambda x: remove_bib_letters(x))
```
Lastly, I would like to normalize the data.
```
X = df.drop('racertype', axis = 1)
y = df['racertype']
X_norm = normalize(X)
```
In addition, I'd like to apply PCA to the data. For a first try, I'd like to reduce the data down to 4 features.
```
X_pca = PCA(4).fit_transform(X_norm)
```
#### 3.2 - K Means Clustering
Based on the crosstabs, it seems that the pca transformed data performs better than the non-pca data, but the main difference is identifying more female runners. Male runners and those in wheelchairs do not change much. In fact, it seems like both normalized and pca transformed data group the wheelchair racers as one group.
```
y_pred_kmeans = KMeans(n_clusters = 4, random_state=42).fit_predict(X_norm)
print(pd.crosstab(y_pred_kmeans, y))
y_pred_PCAkmeans = KMeans(n_clusters = 4, random_state=42).fit_predict(X_pca)
print(pd.crosstab(y_pred_PCAkmeans, y))
```
The adjusted rand index (ARI) compares how the pairs of data points are related to the ground truth in the new solution. Values range from 0 to 1, where 1 means the pairs are in complete agreement and 0 indicates perfect randomness. For both the normalized and pca transformed data, the ARI are pretty close to randomness, at around 0.018.
```
print('ARI for normalized data: {}'.format(metrics.adjusted_rand_score(y, y_pred_kmeans)))
print('ARI for pca transformed data: {}'.format(metrics.adjusted_rand_score(y, y_pred_PCAkmeans)))
```
A possible explanation is that I'm using the wrong number of clusters. Below, I've plotted the ARI against the number of clusters for the normalized and pca transformed data. It seems that the best ARI comes from 6 clusters for both transformed sets of data.
```
kmeans_norm_ari_lst = []
kmeans_pca_ari_lst = []
for k in range(2, 11):
tmp_pca = KMeans(n_clusters = k, random_state=42).fit_predict(X_pca)
kmeans_pca_ari_lst.append(metrics.adjusted_rand_score(y, tmp_pca))
tmp_norm = KMeans(n_clusters = k, random_state=42).fit_predict(X_norm)
kmeans_norm_ari_lst.append(metrics.adjusted_rand_score(y, tmp_norm))
plt.plot(range(2,11), kmeans_norm_ari_lst)
plt.plot(range(2,11), kmeans_pca_ari_lst)
plt.title('ARI for PCA transformed Data and Various Clusters - KMeans')
plt.xlabel('Number of Clusters (K)')
plt.ylabel('ARI')
plt.legend(['norm', 'pca'])
```
#### 3.3 - Mean Shift
With mean shift clustering, the algorithm predicts the number of clusters. With some adjusting to get the best ARI score, the predicted number of clusters for both normalized and pca transformed data was 4, and ARI ~0.026, which is not as good as k-means. On the positive side, mean shift seems to be able to distinguish between the genders in the wheelchair racers.
```
bandwidth_norm = estimate_bandwidth(X_norm, quantile=0.35, n_samples=1000)
ms_norm = MeanShift(bandwidth=bandwidth_norm, bin_seeding=True)
ms_norm.fit(X_norm)
labels_norm = ms_norm.labels_
cluster_centers_norm = ms_norm.cluster_centers_
n_clusters_norm = len(np.unique(labels_norm))
print("Number of estimated clusters: {}".format(n_clusters_norm))
print(pd.crosstab(labels_norm, y))
bandwidth_pca = estimate_bandwidth(X_pca, quantile=0.35, n_samples=1000)
ms_pca = MeanShift(bandwidth=bandwidth_pca, bin_seeding=True)
ms_pca.fit(X_pca)
labels_pca = ms_pca.labels_
cluster_centers_pca = ms_pca.cluster_centers_
n_clusters_pca = len(np.unique(labels_pca))
print("Number of estimated clusters: {}".format(n_clusters_pca))
print(pd.crosstab(labels_pca, y))
print('ARI for normalized data: {}'.format(metrics.adjusted_rand_score(y, labels_norm)))
print('ARI for pca transformed data: {}'.format(metrics.adjusted_rand_score(y, labels_pca)))
```
### 4 - Conclusions
In this notebook, I tried to use clustering techniques to separate racers into four categories: male runners, female runners, male wheelchair racers, and female wheelchair racers. Using k-means clustering, 6 was the optimal number of clusters for both normalized and pca transformed data, with an ARI of ~0.034. With mean shift clustering, 4 was the optimal number of clusters, with an ARI of ~0.026. From the crosstab information, it appears that both k-means and mean shift had a very difficult time distinguishing between male and female racers. However, mean shift clustering was able to distinguish between genders for the wheelchair racers.
| github_jupyter |
# 1A.soft - Tests unitaires, setup et ingéniérie logicielle
On vérifie toujours qu'un code fonctionne quand on l'écrit mais cela ne veut pas dire qu'il continuera à fonctionner à l'avenir. La robustesse d'un code vient de tout ce qu'on fait autour pour s'assurer qu'il continue d'exécuter correctement.
```
from jyquickhelper import add_notebook_menu
add_notebook_menu()
from pyensae.graphhelper import draw_diagram
```
## Petite histoire
Supposons que vous ayez implémenté trois fonctions qui dépendent les unes des autres. la fonction ``f3`` utilise les fonctions ``f1`` et ``f2``.
```
draw_diagram("blockdiag { f0 -> f1 -> f3; f2 -> f3;}")
```
Six mois plus tard, vous créez une fonction ``f5`` qui appelle une fonction ``f4`` et la fonction ``f2``.
```
draw_diagram('blockdiag { f0 -> f1 -> f3; f2 -> f3; f2 -> f5 [color="red"]; f4 -> f5 [color="red"]; }')
```
Ah au fait, ce faisant, vous modifiez la fonction ``f2`` et vous avez un peu oublié ce que faisait la fonction ``f3``... Bref, vous ne savez pas si la fonction ``f3`` sera impactée par la modification introduite dans la fonction ``f2`` ? C'est ce type de problème qu'on rencontre tous les jours quand on écrit un logiciel à plusieurs et sur une longue durée. Ce notebook présente les briques classiques pour s'assurer de la robustesse d'un logiciel.
* les tests unitaires
* un logiciel de suivi de source
* calcul de couverture
* l'intégration continue
* écrire un setup
* écrire la documentation
* publier sur [PyPi](https://pypi.python.org/pypi)
## Ecrire une fonction
N'importe quel fonction qui fait un calcul, par exemple une fonction qui résoud une équation du second degré.
```
def solve_polynom(a, b, c):
# ....
return None
```
## Ecrire un test unitaire
Un [test unitaire](https://fr.wikipedia.org/wiki/Test_unitaire) est une fonction qui s'assure qu'une autre fonction retourne bien le résultat souhaité. Le plus simple est d'utiliser le module standard [unittest](https://docs.python.org/3/library/unittest.html) et de quitter les notebooks pour utiliser des fichiers. Parmi les autres alternatives : [pytest](https://docs.pytest.org/en/latest/) et [nose](http://nose.readthedocs.io/en/latest/).
## Couverture ou coverage
La [couverture de code](https://fr.wikipedia.org/wiki/Couverture_de_code) est l'ensemble des lignes exécutées par les tests unitaires. Cela ne signifie pas toujours qu'elles soient correctes mais seulement qu'elles ont été exécutées une ou plusieurs sans provoquer d'erreur. Le module le plus simple est [coverage](https://coverage.readthedocs.io/en/coverage-4.4.1/). Il produit des rapports de ce type : [mlstatpy/coverage](https://codecov.io/github/sdpython/mlstatpy?branch=master).
## Créer un compte GitHub
[GitHub](https://github.com/) est un site qui contient la majorité des codes des projets open-source. Il faut créer un compte si vous n'en avez pas, c'est gratuit pour les projets open souce, puis créer un projet et enfin y insérer votre projet. Votre ordinateur a besoin de :
* [git](https://git-scm.com/)
* [GitHub destkop](https://desktop.github.com/)
Vous pouvez lire [GitHub Pour les Nuls : Pas de Panique, Lancez-Vous ! (Première Partie)](https://www.christopheducamp.com/2013/12/15/github-pour-nuls-partie-1/) et bien sûr faire plein de recherches internet.
**Note**
Tout ce que vous mettez sur GitHub pour un projet open-source est en accès libre. Veillez à ne rien mettre de personnel. Un compte GitHub fait aussi partie des choses qu'un recruteur ira regarder en premier.
## Intégration continue
L'[intégration continue](https://fr.wikipedia.org/wiki/Int%C3%A9gration_continue) a pour objectif de réduire le temps entre une modification et sa mise en production. Typiquement, un développeur fait une modification, une machine exécute tous les tests unitaires. On en déduit que le logiciel fonctionne sous tous les angles, on peut sans crainte le mettre à disposition des utilisateurs. Si je résume, l'intégration continue consiste à lancer une batterie de tests dès qu'une modification est détectée. Si tout fonctionne, le logiciel est construit et prêt à être partagé ou déployé si c'est un site web.
Là encore pour des projets open-source, il est possible de trouver des sites qui offre ce service gratuitement :
* [travis](https://travis-ci.com/) - [Linux](https://fr.wikipedia.org/wiki/Linux)
* [appveyor](https://www.appveyor.com/) - [Windows](https://fr.wikipedia.org/wiki/Microsoft_Windows) - 1 job à la fois, pas plus d'une heure.
* [circle-ci](https://circleci.com/) - [Linux](https://fr.wikipedia.org/wiki/Linux) et [Mac OSX](https://fr.wikipedia.org/wiki/MacOS) (payant)
* [GitLab-ci](https://about.gitlab.com/features/gitlab-ci-cd/)
A part [GitLab-ci](https://about.gitlab.com/features/gitlab-ci-cd/), ces trois services font tourner les tests unitaires sur des machines hébergés par chacun des sociétés. Il faut s'enregistrer sur le site, définir un fichier [.travis.yml](https://docs.travis-ci.com/user/customizing-the-build), [.appveyor.yml](https://www.appveyor.com/docs/appveyor-yml/) ou [circle.yml](https://circleci.com/docs/1.0/config-sample/) puis activer le projet sur le site correspondant. Quelques exemples sont disponibles à [pyquickhelper](https://github.com/sdpython/pyquickhelper) ou [scikit-learn](https://github.com/scikit-learn/scikit-learn). Le fichier doit être ajouté au projet sur *GitHub* et activé sur le site d'intégration continue choisi. La moindre modification déclenchera un nouveau *build*.permet
La plupart des sites permettent l'insertion de [badge](https://docs.travis-ci.com/user/status-images/) de façon à signifier que le *build* fonctionne.
```
from IPython.display import SVG
SVG("https://travis-ci.com/sdpython/ensae_teaching_cs.svg?branch=master")
SVG("https://codecov.io/github/sdpython/ensae_teaching_cs/coverage.svg?branch=master")
```
Il y a des badges un peu pour tout.
## Ecrire un setup
Le fichier ``setup.py`` détermin la façon dont le module python doit être installé pour un utilisateur qui ne l'a pas développé. Comment construire un setup : [setup](https://docs.python.org/3.6/distutils/setupscript.html).
## Ecrire la documentation
L'outil est le plus utilisé est [sphinx](https://www.sphinx-doc.org/en/master/). Saurez-vous l'utiliser ?
## Dernière étape : PyPi
[PyPi](https://pypi.python.org/pypi) est un serveur qui permet de mettre un module à la disposition de tout le monde. Il suffit d'uploader le module... [Packaging and Distributing Projects](https://packaging.python.org/tutorials/distributing-packages/) ou [How to submit a package to PyPI](http://peterdowns.com/posts/first-time-with-pypi.html). PyPi permet aussi l'insertion de badge.
```
SVG("https://badge.fury.io/py/ensae_teaching_cs.svg")
```
| github_jupyter |
# Get Started with Notebooks in Azure Machine Learning
Azure Machine Learning is a cloud-based service for creating and managing machine learning solutions. It's designed to help data scientists and machine learning engineers leverage their existing data processing and model development skills and frameworks, and scale their workloads to the cloud.
A lot of data science and machine learning work is accomplished in notebooks like this one. Notebooks consist of *cells*, some of which (like the one containing this text) are used for notes, graphics, and other content usually written using *markdown*; while others (like the cell below this one) contain code that you can run interactively within the notebook.
## The Azure Machine Learning Python SDK
You can run pretty much any Python code in a notebook, provided the required Python packages are installed in the environment where you're running it. In this case, you're running the notebook in a *Conda* environment on an Azure Machine Learning compute instance. This environment is installed in the compute instance by default, and contains common Python packages that data scientists typically work with. It also includes the Azure Machine Learning Python SDK, which is a Python package that enables you to write code that uses resources in your Azure Machine Learning workspace.
Run the cell below to import the **azureml-core** package and checking the version of the SDK that is installed.
```
import azureml.core
print("Ready to use Azure ML", azureml.core.VERSION)
```
## Connect to your workspace
All experiments and associated resources are managed within your Azure Machine Learning workspace. You can connect to an existing workspace, or create a new one using the Azure Machine Learning SDK.
In most cases, you should store workspace connection information in a JSON configuration file. This makes it easier to connect without needing to remember details like your Azure subscription ID. You can download the JSON configuration file from the blade for your workspace in the Azure portal or from the workspace details pane in Azure Machine Learning studio, but if you're using a compute instance within your workspace, the configuration file has already been downloaded to the root folder.
The code below uses the configuration file to connect to your workspace.
> **Note**: The first time you connect to your workspace in a notebook session, you may be prompted to sign into Azure by clicking the `https://microsoft.com/devicelogin` link, entering an automatically generated code, and signing into Azure. After you have successfully signed in, you can close the browser tab that was opened and return to this notebook.
```
from azureml.core import Workspace
ws = Workspace.from_config()
print(ws.name, "loaded")
```
## View Azure Machine Learning resources in the workspace
Now that you have a connection to your workspace, you can work with the resources. For example, you can use the following code to enumerate the compute resources in your workspace.
```
print("Compute Resources:")
for compute_name in ws.compute_targets:
compute = ws.compute_targets[compute_name]
print("\t", compute.name, ':', compute.type)
```
When you've finished exploring this notebook, you can save any changes you have made and close it.
| github_jupyter |
## Skin Deep Learning Training Notebook
```
import keras as K
from keras.layers import Dense, Dropout, Flatten, Conv2D, MaxPool2D
import tensorflow as TF
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import imageio
%matplotlib inline
import os
from glob import glob
import seaborn as sns
from PIL import Image
from sklearn.preprocessing import label_binarize
from sklearn.metrics import confusion_matrix
import itertools
from sklearn.model_selection import train_test_split
#Mounting Google Drive
from google.colab import drive
drive.mount('/content/gdrive')
```
### Loading and Preprocessing data
```
#Reading the metadata_csv to see what the current DataFrame looks like.
metadata_path = 'gdrive/My Drive/Google Colab Data/Skin/HAM10000_metadata.csv'
metadata = pd.read_csv(metadata_path)
metadata.head(5)
# This dictionary is useful for displaying more human-friendly labels later on
lesion_type_dict = {
'nv': 'Melanocytic nevi',
'mel': 'Melanoma',
'bkl': 'Benign keratosis-like lesions ',
'bcc': 'Basal cell carcinoma',
'akiec': 'Actinic keratoses',
'vasc': 'Vascular lesions',
'df': 'Dermatofibroma'
}
#Creating New Columns for better readability
newpath = 'gdrive/My Drive/Google Colab Data/Skin/HAM10000_images_part_1/'
metadata['path'] = metadata['image_id'].map(lambda x: newpath+x+".jpg")
print(list(metadata['path'])[6])
#Writes cell_type & cell_type_index features to the csv
metadata['cell_type'] = metadata['dx'].map(lesion_type_dict.get)
metadata['cell_type_idx'] = pd.Categorical(metadata['cell_type']).codes
# metadata.head(5)
#Resizing images to a 100x75x3 matrix and storing them as a new feature
#for the DF
metadata['image'] = metadata['path'].map(lambda x: np.asarray(Image\
.open(x).resize((100,75))))
# metadata.head(5)
#Plotting one image to confirm that the previous step was successful
plt.figure()
plt.imshow(metadata['image'][4])
```
### Cleaning and Preparing Data for Training
```
#Setting X & y values, in this case image is X and cell_type_idx is y
X = metadata['image'].values
y = metadata['cell_type_idx'].values
"""nx, ny represent the image resolution of the training dataset.
When we use this model for prediction later,
These values will be used to resize any images uploaded"""
nx = X[1].shape[1]
ny = X[1].shape[0]
#nc by convention; Referencing the number of channels used.
nc = X[1].shape[2]
m = X.shape[0]
#reshape X to a nicer shape and print dimentions
X = np.concatenate(X).reshape(m,ny,nx,nc)
X.shape
#np.save('temp.npy', [X, y, m, ny, nx, nc])
```
| github_jupyter |
## Beam Block
Testing json radar file being used in beam block calculation.
```
import matplotlib.pyplot as plt
import json
import pandas as pd
import pyart
from IPython.display import Image, display
get_ipython().magic(
'install_ext https://raw.github.com/cpcloud/ipython-\
autotime/master/autotime.py')
get_ipython().magic('load_ext autotime')
"""
pyart.retrieve.beam_block
=======================================
Calculates partial beam block(PBB) and cumulative beam block(CBB)
by using wradlib's beamblock and geotiff functions. PBB and CBB
are then used to created flags when a certain beam block fraction
is passed. Empty radar object is created using Py-ART and then
is filled with beam block data.
.. autosummary::
:toctreeL generated/
:template: dev_template.rst
beam_block_json
beam_block_json_flag
"""
import json
import numpy as np
import wradlib as wrl
def beam_block_json(json_file, tif_file,
beam_width=1.0):
"""
Beam Block Calculation
Parameters
----------
json_file : string
Name of json file containing radar data.
tif_name : string
Name of geotiff file to use for the
calculation
Other Parameters
----------------
beam_width : float
Radar's beam width for calculation.
Default value is 1.0.
Returns
-------
pbb_all : array
Array of partial beam block fractions for each
gate in each ray.
cbb_all : array
Array of cumulative beam block fractions for
each gate in each ray.
References
----------
Bech, J., B. Codina, J. Lorente, and D. Bebbington,
2003: The sensitivity of single polarization weather
radar beam blockage correction to variability in the
vertical refractivity gradient. J. Atmos. Oceanic
Technol., 20, 845–855
Heistermann, M., Jacobi, S., and Pfaff, T., 2013:
Technical Note: An open source library for processing
weather radar data (wradlib), Hydrol. Earth Syst.
Sci., 17, 863-871, doi:10.5194/hess-17-863-2013
Helmus, J.J. & Collis, S.M., (2016). The Python ARM
Radar Toolkit (Py-ART), a Library for Working with
Weather Radar Data in the Python Programming Language.
Journal of Open Research Software. 4(1), p.e25.
DOI: http://doi.org/10.5334/jors.119
"""
with open(json_file) as data:
json_data = json.load(data)
rasterfile = tif_file
data_raster = wrl.io.open_raster(rasterfile)
proj_raster = wrl.georef.wkt_to_osr(data_raster.GetProjection())
rastercoords, rastervalues = wrl.io.read_raster_data(rasterfile)
sitecoords = (np.float(json_data['variables']['longitude']['data']),
np.float(json_data['variables']['latitude']['data']),
np.float(json_data['variables']['altitude']['data']))
pbb_arrays = []
cbb_arrays = []
_range = np.array(json.loads(json_data['variables']['range']['data']))
beamradius = wrl.util.half_power_radius(_range, beam_width)
for i in range(
len(np.array(
json.loads(json_data['variables']['sweep_start_ray_index']['data'])))):
index_start = np.array(
json.loads(json_data['variables']['sweep_start_ray_index']['data']))[i]
index_end = np.array(
json.loads(json_data['variables']['sweep_end_ray_index']['data']))[i]
elevs = np.array(
json.loads(
json_data['variables']['elevation']['data']))[index_start:index_end + 1]
azimuths = np.array(
json.loads(
json_data['variables']['azimuth']['data']))[index_start:index_end + 1]
rg, azg = np.meshgrid(_range, azimuths)
rg, eleg = np.meshgrid(_range, elevs)
lon, lat, alt = wrl.georef.polar2lonlatalt_n(
rg, azg, eleg, sitecoords)
x_pol, y_pol = wrl.georef.reproject(
lon, lat, projection_target=proj_raster)
polcoords = np.dstack((x_pol, y_pol))
rlimits = (x_pol.min(), y_pol.min(), x_pol.max(), y_pol.max())
# Clip the region inside our bounding box
ind = wrl.util.find_bbox_indices(rastercoords, rlimits)
rastercoords = rastercoords[ind[1]:ind[3], ind[0]:ind[2], ...]
rastervalues = rastervalues[ind[1]:ind[3], ind[0]:ind[2]]
# Map rastervalues to polar grid points
polarvalues = wrl.ipol.cart2irregular_spline(
rastercoords, rastervalues, polcoords)
pbb = wrl.qual.beam_block_frac(polarvalues, alt, beamradius)
pbb = np.ma.masked_invalid(pbb)
pbb_arrays.append(pbb)
maxindex = np.nanargmax(pbb, axis=1)
cbb = np.copy(pbb)
# Iterate over all beams
for ii, index in enumerate(maxindex):
premax = 0.
for jj in range(index):
# Only iterate to max index to make this faster
if pbb[ii, jj] > premax:
cbb[ii, jj] = pbb[ii, jj]
premax = cbb[ii, jj]
else:
cbb[ii, jj] = premax
# beyond max index, everything is max anyway
cbb[ii, index:] = pbb[ii, index]
cbb_arrays.append(cbb)
pbb_all = np.ma.concatenate(pbb_arrays)
cbb_all = np.ma.concatenate(cbb_arrays)
return pbb_all, cbb_all
def beam_block_flag(pbb_all, cbb_all, pbb_threshold,
cbb_threshold):
""" Takes PBB and CBB arrays created from the
beam_block function and user chosen thresholds
to create and array of 1s and 0s, 1 is a flagged gate
where the fraction value is past the threshold. """
pbb_flags = np.empty_like(pbb_all)
pbb_flags[pbb_all > 0.95] = 3
pbb_flags[(pbb_all > 0.05) & (pbb_all < 0.95)] = 2
pbb_flags[pbb_all < 0.05] = 0
cbb_flags = np.empty_like(cbb_all)
cbb_flags[cbb_all > 0.95] = 3
cbb_flags[(cbb_all > 0.05) & (cbb_all < 0.95)] = 2
cbb_flags[cbb_all < 0.05] = 0
return pbb_flags, cbb_flags
def arrays_to_dict(pbb_all, cbb_all):
""" Function that takes the PBB and CBB arrays
and turns them into dictionaries to be used and added
to the pyart radar object. """
pbb_dict = {}
pbb_dict['coordinates'] = 'elevation, azimuth, range'
pbb_dict['units'] = 'unitless'
pbb_dict['data'] = pbb_all
pbb_dict['standard_name'] = 'partial_beam_block'
pbb_dict['long_name'] = 'Partial Beam Block Fraction'
pbb_dict['comment'] = 'Partial beam block fraction due to terrain'
cbb_dict = {}
cbb_dict['coordinates'] = 'elevation, azimuth, range'
cbb_dict['units'] = 'unitless'
cbb_dict['data'] = cbb_all
cbb_dict['standard_name'] = 'cumulative_beam_block'
cbb_dict['long_name'] = 'Cumulative Beam Block Fraction'
cbb_dict['comment'] = 'Cumulative beam block fraction due to terrain'
return pbb_dict, cbb_dict
def flags_to_dict(pbb_flags, cbb_flags):
""" Function that takes the PBB_flag and CBB_flag
arrays and turns them into dictionaries to be used
and added to the pyart radar object. """
pbb_flag_dict = {}
pbb_flag_dict['units'] = 'unitless'
pbb_flag_dict['data'] = pbb_flags
pbb_flag_dict['standard_name'] = 'partial_beam_block_flag'
pbb_flag_dict['long_name'] = 'Partial Beam Block Flag'
pbb_flag_dict['comment'] = 'Partial beam block fraction flag, ' \
'1 for flagged values, 0 for non-flagged.'
cbb_flag_dict = {}
cbb_flag_dict['units'] = 'unitless'
cbb_flag_dict['data'] = cbb_flags
cbb_flag_dict['standard_name'] = 'cumulative_beam_block_flag'
cbb_flag_dict['long_name'] = 'Cumulative Beam Block Flag'
cbb_flag_dict['comment'] = 'Cumulative beam block fraction flag, ' \
'1 for flagged values, 0 for non-flagged.'
return pbb_flag_dict, cbb_flag_dict
def empty_radar_beam_block_ppi(ngates, rays_per_sweep, nsweeps,
lon, lat, alt, range_start,
gate_space, elevations):
""" Creates a radar object with no fields based on
user inputed dimensions. The empty radar is to then
be used to add PBB, CBB and the flags for both. """
radar = pyart.testing.make_empty_ppi_radar(
ngates, rays_per_sweep, nsweeps)
radar.elevation['data'] = np.array([elevations] * rays_per_sweep)
radar.longitude['data'] = np.array([lon])
radar.latitude['data'] = np.array([lat])
radar.altitude['data'] = np.array([alt])
radar.azimuth['data'] = np.linspace(0, 360, rays_per_sweep)
radar.range['data'] = np.linspace(
range_start, (ngates - 1)*gate_space + range_start, ngates)
radar.fixed_angle['data'] = elevations
radar.metadata['instrument_name'] = 'beam_block_radar_object'
return radar
def empty_radar_beam_block_rhi(ngates, rays_per_sweep, nsweeps,
lon, lat, alt, range_start,
gate_space, azimuth, elev_start,
elev_end):
""" Creates a radar object with no fields based on
user inputed dimensions. The empty radar is to then
be used to add PBB, CBB and the flags for both. """
radar = pyart.testing.make_empty_rhi_radar(
ngates, rays_per_sweep, nsweeps)
nrays = rays_per_sweep * nsweeps
radar.longitude['data'] = np.array([lon])
radar.latitude['data'] = np.array([lat])
radar.altitude['data'] = np.array([alt])
# radar.azimuth['data'] = np.linspace(0, 360, rays_per_sweep)
radar.range['data'] = np.linspace(
range_start, (ngates - 1)*gate_space + range_start, ngates)
radar.elevation['data'] = np.linspace(elev_start, elev_end, nrays)
radar.azimuth['data'] = np.array([azimuth] * nrays)
radar.fixed_angle['data'] = np.array([azimuth])
radar.metadata['instrument_name'] = 'beam_block_radar_object'
return radar
# sitecoords = (-28.0257, 39.0916, 40.0)
elevations = np.array([0.5, 1.0, 2.0, 3.0, 4.0, 10.0, 11.0, 15.0,
20.0, 30.0, 40.0])
radar_ppi = empty_radar_beam_block_ppi(700, 430, 1, -28.0257,
39.0916, 40.0, 0, 100,
elevations=2.0)
tif_name = '/home/zsherman/beam_block/data/dtm_gra.tif'
azimuth = 147
radar_rhi = empty_radar_beam_block_rhi(1073, 990, 1, -28.0257,
39.0916, 40.0, 0, 100,
azimuth, 0.8, 90)
_range = radar_ppi.range['data']
sweep_start = radar_ppi.sweep_start_ray_index['data']
sweep_end = radar_ppi.sweep_end_ray_index['data']
az = radar_ppi.azimuth['data']
elev = radar_ppi.elevation['data']
fix_angle = radar_ppi.fixed_angle['data']
time = radar_ppi.time['data']
my_object = {
'variables' : {
'time' : {
'calendar' : 'gregorian',
'data' : json.dumps(time.tolist()),
'long_name' : 'time_in_seconds_since_volume_start',
'standard_name' : 'time',
'units' : 'seconds since 1989-01-01T00:00:01Z'
},
'altitude' : {
'data' : 40.0,
'units' : 'meters',
'positive' : 'up',
'long_name' : 'Altitude',
'standard_name' : 'Altitude'
},
'azimuth' : {
'data' : json.dumps(az.tolist()),
'comment' : 'Azimuth of antenna relative to true north',
'long_name' : 'azimuth_angle_from_true_north',
'units' : 'degrees',
'standard_name' : 'beam_azimuth_angle',
'axis' : 'radial_azimuth_coordinate'
},
'elevation' : {
'data' : json.dumps(elev.tolist()),
'comment' : 'Elevation of antenna relative to the horizontal plane',
'long_name' : 'elevation_angle_from_horizontal_plane',
'units' : 'degrees',
'standard_name' : 'beam_elevation_angle',
'axis' : 'radial_elevation_coordinate'
},
'fields' : {
},
'fixed_angle' : {
'data' : json.dumps(fix_angle),
'units' : 'degrees',
'long_name' : 'Target angle for sweep',
'standard_name' : 'target_fixed_angle'
},
'latitude' : {
'data' : 39.0916,
'units' : 'degrees_north',
'long_name' : 'Latitude',
'standard_name' : 'Latitude'
},
'longitude' : {
'data' : -28.0257,
'units' : 'degrees_east',
'long_name' : 'Longitude',
'standard_name' : 'Longitude'
},
'range' : {
'data' : json.dumps(_range.tolist())
},
'scan_type' : 'ppi',
'sweep_start_ray_index' : {
'data' : json.dumps(sweep_start.tolist()),
'units' : 'unitless',
'long_name' : 'Index of first ray in sweep, 0-based'
},
'sweep_end_ray_index' : {
'data' : json.dumps(sweep_end.tolist()),
'units' : 'unitless',
'long_name' : 'Index of last ray in sweep, 0-based'
}
}
}
json_file = '/home/zsherman/practice_json.json'
with open(json_file) as data:
json_data = json.load(data)
_range = np.array(
json.loads(json_data['variables']['range']['data']))
elevs = np.array(
json.loads(json_data['variables']['elevation']['data']))
azimuths = np.array(
json.loads(json_data['variables']['azimuth']['data']))
fixed_angle = np.array(
np.float(json_data['variables']['fixed_angle']['data']))
index_start = np.array(
json.loads(
json_data['variables']['sweep_start_ray_index']['data']))
index_end = np.array(
json.loads(
json_data['variables']['sweep_end_ray_index']['data']))
ngates = len(_range)
nrays = len(azimuths)
nsweeps = len(index_start)
time = np.array(
json.loads(json_data['variables']['time']['data']))
lon = np.array(
np.float(json_data['variables']['longitude']['data']))
lat = np.array(
np.float(json_data['variables']['latitude']['data']))
alt = np.array(
np.float(json_data['variables']['altitude']['data']))
radar = pyart.testing.make_empty_ppi_radar(ngates, 1, nsweeps)
radar.metadata = {'instrument_name': 'beam block'}
radar.nrays = nrays
radar.time['data'] = time
radar.range['data'] = _range.astype('float32')
radar.latitude['data'] = lat.astype('float64')
radar.longitude['data'] = lon.astype('float64')
radar.altitude['data'] = alt.astype('float64')
radar.sweep_number['data'] = np.arange(nsweeps, dtype='int32')
radar.sweep_start_ray_index['data'] = index_start.astype('int32')
radar.sweep_end_ray_index['data'] = index_end.astype('int32')
radar.fixed_angle['data'] = fixed_angle.astype('float32')
radar.azimuth['data'] = azimuths.astype('float32')
radar.elevation['data'] = elevs.astype('float32')
practice_radar = json.dumps(my_object)
practice_json = json.loads(practice_radar)
with open('/home/zsherman/practice_json.json', 'w') as outfile:
json.dump(my_object, outfile, sort_keys=True, indent=4,
ensure_ascii=False)
with open('/home/zsherman/practice_json.json') as data_file:
data = json.load(data_file)
var = practice_json['variables']
_range = np.array(json.loads(var['range']['data']))
var['azimuth']['data']
pd_json = pd.read_json(
'/home/zsherman/practice_json.json')
pd_json
json_file = '/home/zsherman/practice_json.json'
pbb_all, cbb_all = beam_block_json(
json_file, tif_name, 1.0)
pbb_json = {'paritial_beam_block' : json.dumps(pbb_all.tolist())}
data['variables']['partial_beam_block'] = {
'data' : json.dumps(pbb_all.tolist()),
'long_name' : 'Partial Beam Block',
'standard_name' : 'partial_beam_block'}
#from pprint import pprint
#pprint(data)
pbb_array = np.array(json.loads(data['variables']['partial_beam_block']['data']), dtype=np.float)
masked_array = np.ma.masked_invalid(pbb_array)
masked_array.max()
pbb_flags, cbb_flags = beam_block_flag(
pbb_all, cbb_all, 0.2, 0.2)
pbb_dict, cbb_dict = arrays_to_dict(
pbb_all, cbb_all)
pbb_flag_dict, cbb_flag_dict = flags_to_dict(
pbb_flags, cbb_flags)
pbb = masked_array
cbb = cbb_all
r = np.array(json.loads(practice_json['variables']['range']['data']))
az = np.array(json.loads(practice_json['variables']['azimuth']['data']))
el = np.array(json.loads(practice_json['variables']['elevation']['data']))
fig = plt.figure(figsize=(10, 7))
ax, dem = wrl.vis.plot_ppi(pbb, r=r,
az=az,
cmap=plt.cm.PuRd)
ax.set_xlim(-8000, 10000)
ax.set_ylim(-12000, 5000)
ax.plot(0, 0, 'ro')
ax.grid(True)
ax.annotate(' ARM ENA Site', (0, 0))
ticks = (ax.get_xticks()/1000).astype(np.int)
ax.set_xticklabels(ticks)
ticks = (ax.get_yticks()/1000).astype(np.int)
ax.set_yticklabels(ticks)
ax.set_title('Partial Beam Block 2.0 Degrees')
ax.set_xlabel("Kilometers")
ax.set_ylabel("Kilometers")
ax.set_axis_bgcolor('#E0E0E0')
plt.colorbar(dem, ax=ax)
plt.show()
```
| github_jupyter |
# Ray Tune - Search Algorithms and Schedulers
© 2019-2021, Anyscale. All Rights Reserved

This notebook introduces the concepts of search algorithms and schedulers which help optimize HPO. We'll see an example that combines the use of one search algorithm and one schedulers.
The full set of search algorithms provided by Tune is documented [here](https://docs.ray.io/en/latest/tune/api_docs/suggestion.html), along with information about implementing your own. The full set of schedulers provided is documented [here](https://docs.ray.io/en/latest/tune/api_docs/schedulers.html).
We need to install a few libraries. We'll explain what they are below.
```
!pip install hpbandster ConfigSpace
!python --version
```
> **NOTE:** If you are see **Python 3.6** in the output from the previous cell, run remove the `#` in the following cell and run it. This will fix a dependency bug needed for this notebook.
>
> Afterwards, **restart the kernel for this notebook**, using the circular error in the tool bar. After that, proceed with the rest of the notebook.
>
> If you have **Python 3.7** or later, skip these steps.
```
#!pip install statsmodels -U --pre
```
## About Search Algorithms
Tune integrates many [open source optimization libraries](https://docs.ray.io/en/latest/tune/api_docs/suggestion.html), each of which defines the parameter search space in its own way. Hence, you should read the corresponding documentation for an algorithm to understand the particular details of using it.
Some of the search algorithms supported include the following:
* [Bayesian Optimization](https://github.com/fmfn/BayesianOptimization): This constrained global optimization process builds upon bayesian inference and gaussian processes. It attempts to find the maximum value of an unknown function in as few iterations as possible. This is a good technique for optimization of high cost functions.
* [BOHB (Bayesian Optimization HyperBand](https://github.com/automl/HpBandSter): An algorithm that both terminates bad trials and also uses Bayesian Optimization to improve the hyperparameter search. It is backed by the [HpBandSter](https://docs.ray.io/en/latest/tune/api_docs/schedulers.html#tune-scheduler-bohb) library. BOHB is intended to be paired with a specific scheduler class: [HyperBandForBOHB](https://docs.ray.io/en/latest/tune/api_docs/schedulers.html#tune-scheduler-bohb).
* [HyperOpt](http://hyperopt.github.io/hyperopt): A Python library for serial and parallel optimization over awkward search spaces, which may include real-valued, discrete, and conditional dimensions.
* [Nevergrad](https://github.com/facebookresearch/nevergrad): HPO without computing gradients.
These and other algorithms are described in the [documentation](https://docs.ray.io/en/latest/tune/api_docs/suggestion.html).
A limitation of search algorithms used by themselves is they can't affect or stop training processes, for example early stopping of trials that are performing poorly. The schedulers can do this, so it's common to use a compatible search algorithm with a scheduler, as we'll show in the first example.
## About Schedulers
Tune includes distributed implementations of several early-stopping algorithms, including the following:
* [Median Stopping Rule](https://research.google.com/pubs/pub46180.html): It applies the simple rule that a trial is aborted if the results are trending below the median of the previous trials.
* [HyperBand](https://arxiv.org/abs/1603.06560): It structures search as an _infinite-armed, stochastic, exploration-only, multi-armed bandit_. See the [Multi-Armed Bandits lessons](../ray-rllib/multi-armed-bandits/00-Multi-Armed-Bandits-Overview.ipynb) for information on these concepts. The infinite arms correspond to the tunable parameters. Trying values stochastically ensures quick exploration of the parameter space. Exploration-only is desirable because for HPO, we aren't interested in _exploiting_ parameter combinations we've already tried (the usual case when using MABs where rewards are the goal). Intead, we need to explore as many new parameter combinations as possible.
* [ASHA](https://openreview.net/forum?id=S1Y7OOlRZ). This is an aynchronous version of HyperBand that improves on the latter. Hence it is recommended over the original HyperBand implementation.
Tune also includes a distributed implementation of [Population Based Training (PBT)](https://deepmind.com/blog/population-based-training-neural-networks). When the PBT scheduler is enabled, each trial variant is treated as a member of the _population_. Periodically, top-performing trials are checkpointed, which means your [`tune.Trainable`](https://docs.ray.io/en/latest/tune/api_docs/trainable.html#tune-trainable) object (e.g., the `TrainMNist` class we used in the previous exercise) has to support save and restore.
Low-performing trials clone the checkpoints of top performers and perturb the configurations in the hope of discovering an even better variation. PBT trains a group of models (or RLlib agents) in parallel. So, unlike other hyperparameter search algorithms, PBT mutates hyperparameters during training time. This enables very fast hyperparameter discovery and also automatically discovers good [annealing](https://en.wikipedia.org/wiki/Simulated_annealing) schedules.
See the [Tune schedulers](https://docs.ray.io/en/latest/tune/api_docs/schedulers.html) for a complete list and descriptions.
## Examples
Let's initialize Ray as before:
```
import ray
from ray import tune
ray.init(ignore_reinit_error=True)
```
### BOHB
BOHB (Bayesian Optimization HyperBand) is an algorithm that both terminates bad trials and also uses Bayesian Optimization to improve the hyperparameter search. The [Tune implementation](https://docs.ray.io/en/latest/tune/api_docs/suggestion.html#bohb-tune-suggest-bohb-tunebohb) is backed by the [HpBandSter library](https://github.com/automl/HpBandSter), which we must install, along with [ConfigSpace](https://automl.github.io/HpBandSter/build/html/quickstart.html#searchspace), which is used to define the search space specification:
We use BOHB with the scheduler [HyperBandForBOHB](https://docs.ray.io/en/latest/tune/api_docs/schedulers.html#bohb-tune-schedulers-hyperbandforbohb).
Let's try it. We'll use the same MNIST example from the previous lesson, but this time, we'll import the code from a file in this directory, `mnist.py`. Note that the implementation of `TrainMNIST` in the file has enhancements not present in the previous lesson, such as methods to support saving and restoring checkpoints, which are required to be used here. See the code comments for details.
```
from mnist import ConvNet, TrainMNIST, EPOCH_SIZE, TEST_SIZE, DATA_ROOT
```
Import and configure the `ConfigSpace` object we need for the search algorithm.
```
import ConfigSpace as CS
from ray.tune.schedulers.hb_bohb import HyperBandForBOHB
from ray.tune.suggest.bohb import TuneBOHB
config_space = CS.ConfigurationSpace()
# There are also UniformIntegerHyperparameter and UniformFloatHyperparameter
# objects for defining integer and float ranges, respectively. For example:
# config_space.add_hyperparameter(
# CS.UniformIntegerHyperparameter('foo', lower=0, upper=100))
config_space.add_hyperparameter(
CS.CategoricalHyperparameter('lr', choices=[0.001, 0.01, 0.1]))
config_space.add_hyperparameter(
CS.CategoricalHyperparameter('momentum', choices=[0.001, 0.01, 0.1, 0.9]))
config_space
experiment_metrics = dict(metric="mean_accuracy", mode="max")
search_algorithm = TuneBOHB(config_space, max_concurrent=4, **experiment_metrics)
scheduler = HyperBandForBOHB(
time_attr='training_iteration',
reduction_factor=4,
max_t=200,
**experiment_metrics)
```
Through experimentation, we determined that `max_t=200` is necessary to get good results. For the smallest learning rate and momentum values, it takes longer for training to converge.
```
analysis = tune.run(TrainMNIST,
scheduler=scheduler,
search_alg=search_algorithm,
num_samples=12, # Force it try all 12 combinations
verbose=1
)
stats = analysis.stats()
secs = stats["timestamp"] - stats["start_time"]
print(f'{secs:7.2f} seconds, {secs/60.0:7.2f} minutes')
print("Best config: ", analysis.get_best_config(metric="mean_accuracy"))
analysis.dataframe().sort_values('mean_accuracy', ascending=False).head()
analysis.dataframe()[['mean_accuracy', 'config/lr', 'config/momentum']].sort_values('mean_accuracy', ascending=False)
```
The runs in the previous lesson, for the class-based and the function-based Tune APIs, took between 12 and 20 seconds (on my machine), but we only trained for 20 iterations, where as here we went for 100 iterations. That also accounts for the different results, notably that a much smaller momentum value `0.01` and `0.1` perform best here, while for the the previous lesson `0.9` performed best. This is because a smaller momentum value will result in longer training times required, but more fine-tuned iterating to the optimal result, so more training iterations will favor a smaller momentum value. Still, the mean accuracies among the top three or four combinations are quite close.
## Exercise - Population Base Training
Read the [documentation]() on _population based training_ to understand what it is doing. The next cell configures a PBT scheduler and defines other things you'll need.
See also the discussion for the results in the [solutions](solutions/03-Search-Algos-and-Schedulers-Solutions.ipynb).
> **NOTE:** For a more complete example using MNIST and PyTorch, see [this example code](https://github.com/ray-project/ray/blob/master/python/ray/tune/examples/mnist_pytorch_lightning.py).
```
from ray.tune.schedulers import PopulationBasedTraining
pbt_scheduler = PopulationBasedTraining(
time_attr='training_iteration',
perturbation_interval=10, # Every N time_attr units, "perturb" the parameters.
hyperparam_mutations={
"lr": [0.001, 0.01, 0.1],
"momentum": [0.001, 0.01, 0.1, 0.9]
},
**experiment_metrics)
# Note: This appears to be needed to avoid a "key error", but in fact these values won't change
# in the analysis.dataframe() object, even though they will be tuned by the PBT scheduler.
# So when you look at the analysis.dataframe(), look at the `experiment_tag` to see the actual values!
config = {
"lr": 0.001, # Use the lowest values from the previous definition
"momentum": 0.001
}
```
Now run the the following cell, modified from above, which makes these changes:
1. Uses the new scheduler.
2. Removes the search_alg argument.
3. Adds the `config` argument.
4. Don't allow it to keep going past `0.97` accuracy for `600` iterations.
5. Use `1` for the `verbose` argument to reduce the "noise".
Then run it.
> **WARNING:** This will run for a few minutes.
```
analysis = tune.run(TrainMNIST,
scheduler=pbt_scheduler,
config=config,
stop={"mean_accuracy": 0.97, "training_iteration": 600},
num_samples=8,
verbose=1
)
stats = analysis.stats()
secs = stats["timestamp"] - stats["start_time"]
print(f'{secs:7.2f} seconds, {secs/60.0:7.2f} minutes')
```
Look at the `analysis` data of interest, as done previously. (You might want to focus on other columns in the dataframe.) How well does PBT work?
The final lesson in this tutorial discusses the new Ray SGD library.
```
ray.shutdown() # "Undo ray.init()".
```
| github_jupyter |
## RANDOM FOREST
*Random forest builds multiple decision trees and merges them together to get a more accurate and stable prediction.*
One big advantage of random forest is that it can be used for both **classification** and **regression problems**, which form the majority of current machine learning systems.
Random forest adds additional randomness to the model, while growing the trees. Instead of searching for the most important feature while splitting a node, it searches for the best feature among a random subset of features. This results in a wide diversity that generally results in a better model.
Therefore, in random forest, only a random subset of the features is taken into consideration by the algorithm for splitting a node. You can even make trees more random by additionally using random thresholds for each feature rather than searching for the best possible thresholds (like a normal decision tree does).
**WHY RANDOM FOREST ?**
- The same random forest algorithm or the random forest classifier can use for both classification and the regression task.
- Random forest classifier will handle the missing values.
- When we have more trees in the forest, random forest classifier won’t overfit the model.
- Can model the random forest classifier for categorical values also.
### OBJECTIVE : IMPLEMENT RANDOM FOREST FROM SCRATCH
### PREREQUISITE :
DECISION TREE ALGORITHM
- Refer the Code present in the [Decision_tree_functions]() to know the implementation of Decision tree.
- Our focus in this Notebook is to implement Random forest using the knowledge of Decision tree.
### DATA
We will be using a [Red Wine quality](https://www.kaggle.com/uciml/red-wine-quality-cortez-et-al-2009) from the Kaggle platform which has a very good collection of datasets.
The data is presented in CSV format.
Features:
- fixed acidity
- volatile acidity
- citric acid
- residual sugar
- chlorides
- free sulfur dioxide
- total sulfur dioxide
- densitythe density
- pHdescribes
- sulphatesa wine
- alcohol
- quality (OUTPUT VARIABLE)
Note that prices have been adjusted for dividends and splits.
### IMPORTED LIBRARIES
```
"""Import all the libraries required to complete the task"""
"""import the below functions from decision_tree_functions.py file
- decision_tree_algorithm
- decision_tree_predictions"""
```
## Load and Prepare Data
*Format of the data*
- the last column of the data frame must contain the label and it must also be called "label"
- there should be no missing values in the data frame"""
```
""" Loading the CSV file using pandas that contains the data and consider **quality** as the output variable"""
"""Replace the space in feature names with the '_' underscore"""
""" display the top 5 data instances"""
""" normalise the label and sort according to the index and plot """
""" Write a function to covert the label into binary classes
value below 5 is bad and above is good """
""" normalise the label and plot bar graph"""
```
Seed function is used to save the state of random function,
so that it can generate some random numbers on multiple execution of the code on the same machine or
on different machines (for a specific seed value). Seed value is the previous value number generated by the generator.
For the first time when there is no previous value, it uses current system time.
```
"""initiaise a random function using seedfunction"""
""" write a function to split the Data """
# 1. Train-Test-Split
```
## Random Forest
```
""" Write a function to create a bootstrap dataset with parameters as dataset and size of bootstrap dataset required"""
```
**Random Forest pseudocode:**
- Randomly select “k” features from total “m” features.
Where k << m
- Among the “k” features, calculate the node “d” using the best split point.
- Split the node into daughter nodes using the best split.
- Repeat 1 to 3 steps until “l” number of nodes has been reached.
- Build forest by repeating steps 1 to 4 for “n” number times to create “n” number of trees.
- The beginning of random forest algorithm starts with randomly selecting “k” features out of total “m” features. In the image, you can observe that we are randomly taking features and observations.
- In the next stage, we are using the randomly selected “k” features to find the root node by using the best split approach.
- The next stage, We will be calculating the daughter nodes using the same best split approach. Will the first 3 stages until we form the tree with a root node and having the target as the leaf node.
- Finally, we repeat 1 to 4 stages to create “n” randomly created trees. This randomly created trees forms the random forest.
```
""" Write a function to create a Random forest using decision tree function importes"""
```
Random forest prediction pseudocode:
- To perform prediction using the trained random forest algorithm uses the below pseudocode.
- Takes the test features and use the rules of each randomly created decision tree to predict the oucome and stores the predicted outcome (target)
- Calculate the votes for each predicted target.
- Consider the high voted predicted target as the final prediction from the random forest algorithm.
- To perform the prediction using the trained random forest algorithm we need to pass the test features through the rules of each randomly created trees. Suppose let’s say we formed 100 random decision trees to from the random forest.
- Each random forest will predict different target (outcome) for the same test feature. Then by considering each predicted target votes will be calculated. Suppose the 100 random decision trees are prediction some 3 unique targets x, y, z then the votes of x is nothing but out of 100 random decision tree how many trees prediction is x.
- Likewise for other 2 targets (y, z). If x is getting high votes. Let’s say out of 100 random decision tree 60 trees are predicting the target will be x. Then the final random forest returns the x as the predicted target.
- This concept of voting is known as **majority voting**.
```
"""Write the function to predict using random forest trees"""
"""Write a function to calculate accuracy where accuracy is mean of correct predictions"""
""" call the random forest function """
""" call the prediction function"""
""" calculate the accuracy"""
```
| github_jupyter |
```
# General
from os import path
from random import randrange
from sklearn.model_selection import train_test_split, GridSearchCV #cross validation
from sklearn.metrics import confusion_matrix, plot_confusion_matrix, make_scorer
from sklearn.metrics import accuracy_score, roc_auc_score, balanced_accuracy_score
from sklearn.preprocessing import LabelEncoder, StandardScaler
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import xgboost as xgb
from sklearn.ensemble import RandomForestClassifier
import pickle
import joblib
```
## TRAIN SET
```
trainDataFull = pd.read_csv("trainData.csv")
trainDataFull.head(3)
trainDataFull.info()
trainDataFull.describe()
trainData = trainDataFull.loc[:,'v1':'v99']
trainData.head(3)
trainLabels = trainDataFull.loc[:,'target']
trainLabels.unique()
# encode string class values as integers
label_encoder = LabelEncoder()
label_encoder = label_encoder.fit(trainLabels)
label_encoded_y = label_encoder.transform(trainLabels)
label_encoded_y
```
## Normalize
```
scaler = StandardScaler()
scaler.fit(trainData.values)
scaler.mean_
normalized_standart = scaler.transform(trainData)
X_train, X_test, y_train, y_test = train_test_split(normalized_standart,
label_encoded_y,
test_size = 0.3,
random_state = 33,
shuffle = True,
stratify = label_encoded_y)
```
## MODEL-2 (Random Forest Classifier)
```
RFC_model = RandomForestClassifier(n_estimators=1000,
verbose=2,
random_state=0,
criterion='gini')
RFC_model
RFC_model.fit(X_train, y_train)
# make predictions for test data
y_pred = RFC_model.predict(X_test)
y_pred
predictions = [round(value) for value in y_pred]
# evaluate predictions
accuracy = accuracy_score(y_test, predictions)
print("Accuracy: %.2f%%" % (accuracy * 100.0))
#fig = plt.figure(figsize=(10,10))
plot_confusion_matrix(RFC_model,
X_test,
y_test,
values_format='d')
```
## Save Valid Score
```
y_score = RFC_model.predict_proba(X_test)
y_score[0]
valid_score = pd.DataFrame(y_score, columns=['c1','c2','c3','c4','c5','c6','c7','c8','c9'])
valid_score
valid_score.to_csv('./results/valid-submission-RFC-standart-norm-3.csv', index = False)
```
## Save & Load Model
## joblib
```
# Save the model as a pickle in a file
joblib.dump(RFC_model, './model/model_RFC-standart-norm-3.pkl')
# Load the model from the file
RFC_model_from_joblib = joblib.load('./model/model_RFC-standart-norm-3.pkl')
# Use the loaded model to make predictions
RFC_model_predictions = RFC_model_from_joblib.predict(X_test)
# evaluate predictions
accuracy = accuracy_score(y_test, RFC_model_predictions)
print("Accuracy: %.2f%%" % (accuracy * 100.0))
```
## GridSearchCV
```
clf = GridSearchCV(RFC_model_model,
{'max_depth': [4, 6],
'n_estimators': [100, 200]},
verbose=1,
cv=2)
clf.fit(X_train,
y_train,
early_stopping_rounds=10,
eval_metric='mlogloss',
eval_set=[(X_train, y_train), (X_test, y_test)],
verbose=True)
print(clf.best_score_)
print(clf.best_params_)
# Save the model as a pickle in a file
joblib.dump(clf.best_estimator_, './model/clf.pkl')
# Load the model from the file
clf_from_joblib = joblib.load('./model/clf.pkl')
# Use the loaded model to make predictions
clf_predictions = clf_from_joblib.predict(X_test)
# evaluate predictions
accuracy = accuracy_score(y_test, clf_predictions)
print("Accuracy: %.2f%%" % (accuracy * 100.0))
```
# TEST
```
testData = pd.read_csv("testData.csv")
testData
test_normalized_standart = scaler.transform(testData.values)
test_normalized_standart
# Use the loaded model to make predictions
# test_predictions = RFC_model.predict(test_normalized_standart)
# test_predictions
# Use the loaded model to make predictions probability
test_predictions = RFC_model.predict_proba(test_normalized_standart)
test_predictions
result = pd.DataFrame(test_predictions, columns=['c1','c2','c3','c4','c5','c6','c7','c8','c9'])
result
result.to_csv('./results/test-submission-RFC-standart-norm-3.csv', index = False)
```
## REFERENCES
1- https://xgboost.readthedocs.io/en/latest/python/python_api.html#module-xgboost.sklearn
2- https://github.com/dmlc/xgboost/blob/master/demo/guide-python/sklearn_examples.py
3- https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.GridSearchCV.html
4- https://www.datacamp.com/community/tutorials/xgboost-in-python
5- https://scikit-learn.org/stable/modules/ensemble.html#voting-classifier
6- https://www.datacamp.com/community/tutorials/random-forests-classifier-python?utm_source=adwords_ppc&utm_campaignid=1455363063&utm_adgroupid=65083631748&utm_device=c&utm_keyword=&utm_matchtype=b&utm_network=g&utm_adpostion=&utm_creative=332602034364&utm_targetid=aud-392016246653:dsa-429603003980&utm_loc_interest_ms=&utm_loc_physical_ms=1012782&gclid=EAIaIQobChMI49HTjNO06wIVB-ztCh23nwMLEAAYASAAEgKKEvD_BwE
| github_jupyter |
# Session 5: Generative Networks
## Assignment: Generative Adversarial Networks, Variational Autoencoders, and Recurrent Neural Networks
<p class="lead">
<a href="https://www.kadenze.com/courses/creative-applications-of-deep-learning-with-tensorflow/info">Creative Applications of Deep Learning with Google's Tensorflow</a><br />
<a href="http://pkmital.com">Parag K. Mital</a><br />
<a href="https://www.kadenze.com">Kadenze, Inc.</a>
</p>
Continued from [session-5-part-1.ipynb](session-5-part-1.ipynb)...
# Table of Contents
<!-- MarkdownTOC autolink="true" autoanchor="true" bracket="round" -->
- [Overview](session-5-part-1.ipynb#overview)
- [Learning Goals](session-5-part-1.ipynb#learning-goals)
- [Part 1 - Generative Adversarial Networks \(GAN\) / Deep Convolutional GAN \(DCGAN\)](#part-1---generative-adversarial-networks-gan--deep-convolutional-gan-dcgan)
- [Introduction](session-5-part-1.ipynb#introduction)
- [Building the Encoder](session-5-part-1.ipynb#building-the-encoder)
- [Building the Discriminator for the Training Samples](session-5-part-1.ipynb#building-the-discriminator-for-the-training-samples)
- [Building the Decoder](session-5-part-1.ipynb#building-the-decoder)
- [Building the Generator](session-5-part-1.ipynb#building-the-generator)
- [Building the Discriminator for the Generated Samples](session-5-part-1.ipynb#building-the-discriminator-for-the-generated-samples)
- [GAN Loss Functions](session-5-part-1.ipynb#gan-loss-functions)
- [Building the Optimizers w/ Regularization](session-5-part-1.ipynb#building-the-optimizers-w-regularization)
- [Loading a Dataset](session-5-part-1.ipynb#loading-a-dataset)
- [Training](session-5-part-1.ipynb#training)
- [Equilibrium](session-5-part-1.ipynb#equilibrium)
- [Part 2 - Variational Auto-Encoding Generative Adversarial Network \(VAEGAN\)](#part-2---variational-auto-encoding-generative-adversarial-network-vaegan)
- [Batch Normalization](session-5-part-1.ipynb#batch-normalization)
- [Building the Encoder](session-5-part-1.ipynb#building-the-encoder-1)
- [Building the Variational Layer](session-5-part-1.ipynb#building-the-variational-layer)
- [Building the Decoder](session-5-part-1.ipynb#building-the-decoder-1)
- [Building VAE/GAN Loss Functions](session-5-part-1.ipynb#building-vaegan-loss-functions)
- [Creating the Optimizers](session-5-part-1.ipynb#creating-the-optimizers)
- [Loading the Dataset](session-5-part-1.ipynb#loading-the-dataset)
- [Training](session-5-part-1.ipynb#training-1)
- [Part 3 - Latent-Space Arithmetic](session-5-part-1.ipynb#part-3---latent-space-arithmetic)
- [Loading the Pre-Trained Model](session-5-part-1.ipynb#loading-the-pre-trained-model)
- [Exploring the Celeb Net Attributes](session-5-part-1.ipynb#exploring-the-celeb-net-attributes)
- [Find the Latent Encoding for an Attribute](session-5-part-1.ipynb#find-the-latent-encoding-for-an-attribute)
- [Latent Feature Arithmetic](session-5-part-1.ipynb#latent-feature-arithmetic)
- [Extensions](session-5-part-1.ipynb#extensions)
- [Part 4 - Character-Level Language Model](session-5-part-2.ipynb#part-4---character-level-language-model)
- [Part 5 - Pretrained Char-RNN of Donald Trump](session-5-part-2.ipynb#part-5---pretrained-char-rnn-of-donald-trump)
- [Getting the Trump Data](session-5-part-2.ipynb#getting-the-trump-data)
- [Basic Text Analysis](session-5-part-2.ipynb#basic-text-analysis)
- [Loading the Pre-trained Trump Model](session-5-part-2.ipynb#loading-the-pre-trained-trump-model)
- [Inference: Keeping Track of the State](session-5-part-2.ipynb#inference-keeping-track-of-the-state)
- [Probabilistic Sampling](session-5-part-2.ipynb#probabilistic-sampling)
- [Inference: Temperature](session-5-part-2.ipynb#inference-temperature)
- [Inference: Priming](session-5-part-2.ipynb#inference-priming)
- [Assignment Submission](session-5-part-2.ipynb#assignment-submission)
<!-- /MarkdownTOC -->
# First check the Python version
import sys
if sys.version_info < (3,4):
print('You are running an older version of Python!\n\n',
'You should consider updating to Python 3.4.0 or',
'higher as the libraries built for this course',
'have only been tested in Python 3.4 and higher.\n')
print('Try installing the Python 3.5 version of anaconda'
'and then restart `jupyter notebook`:\n',
'https://www.continuum.io/downloads\n\n')
# Now get necessary libraries
try:
import os
import numpy as np
import matplotlib.pyplot as plt
from skimage.transform import resize
from skimage import data
from scipy.misc import imresize
from scipy.ndimage.filters import gaussian_filter
import IPython.display as ipyd
import tensorflow as tf
from libs import utils, gif, datasets, dataset_utils, nb_utils
except ImportError as e:
print("Make sure you have started notebook in the same directory",
"as the provided zip file which includes the 'libs' folder",
"and the file 'utils.py' inside of it. You will NOT be able",
"to complete this assignment unless you restart jupyter",
"notebook inside the directory created by extracting",
"the zip file or cloning the github repo.")
print(e)
# We'll tell matplotlib to inline any drawn figures like so:
%matplotlib inline
plt.style.use('ggplot')
```
# Bit of formatting because I don't like the default inline code style:
from IPython.core.display import HTML
HTML("""<style> .rendered_html code {
padding: 2px 4px;
color: #c7254e;
background-color: #f9f2f4;
border-radius: 4px;
} </style>""")
```
<style> .rendered_html code {
padding: 2px 4px;
color: #c7254e;
background-color: #f9f2f4;
border-radius: 4px;
} </style>
<a name="part-4---character-level-language-model"></a>
# Part 4 - Character-Level Language Model
We'll now continue onto the second half of the homework and explore recurrent neural networks. We saw one potential application of a recurrent neural network which learns letter by letter the content of a text file. We were then able to synthesize from the model to produce new phrases. Let's try to build one. Replace the code below with something that loads your own text file or one from the internet. Be creative with this!
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
import tensorflow as tf
from six.moves import urllib
script = 'http://www.awesomefilm.com/script/biglebowski.txt'
txts = []
f, _ = urllib.request.urlretrieve(script, script.split('/')[-1])
with open(f, 'r') as fp:
txt = fp.read()
```
Let's take a look at the first part of this:
```
txt[:100]
```
We'll just clean up the text a little. This isn't necessary, but can help the training along a little. In the example text I provided, there is a lot of white space (those \t's are tabs). I'll remove them. There are also repetitions of \n, new lines, which are not necessary. The code below will remove the tabs, ending whitespace, and any repeating newlines. Replace this with any preprocessing that makes sense for your dataset. Try to boil it down to just the possible letters for what you want to learn/synthesize while retaining any meaningful patterns:
```
txt = "\n".join([txt_i.strip()
for txt_i in txt.replace('\t', '').split('\n')
if len(txt_i)])
```
Now we can see how much text we have:
```
len(txt)
```
In general, we'll want as much text as possible. But I'm including this just as a minimal example so you can explore your own. Try making a text file and seeing the size of it. You'll want about 1 MB at least.
Let's now take a look at the different characters we have in our file:
```
vocab = list(set(txt))
vocab.sort()
len(vocab)
print(vocab)
```
And then create a mapping which can take us from the letter to an integer look up table of that letter (and vice-versa). To do this, we'll use an `OrderedDict` from the `collections` library. In Python 3.6, this is the default behavior of dict, but in earlier versions of Python, we'll need to be explicit by using OrderedDict.
```
from collections import OrderedDict
encoder = OrderedDict(zip(vocab, range(len(vocab))))
decoder = OrderedDict(zip(range(len(vocab)), vocab))
encoder
```
We'll store a few variables that will determine the size of our network. First, `batch_size` determines how many sequences at a time we'll train on. The `seqence_length` parameter defines the maximum length to unroll our recurrent network for. This is effectively the depth of our network during training to help guide gradients along. Within each layer, we'll have `n_cell` LSTM units, and `n_layers` layers worth of LSTM units. Finally, we'll store the total number of possible characters in our data, which will determine the size of our one hot encoding (like we had for MNIST in Session 3).
```
# Number of sequences in a mini batch
batch_size = 100
# Number of characters in a sequence
sequence_length = 50
# Number of cells in our LSTM layer
n_cells = 128
# Number of LSTM layers
n_layers = 3
# Total number of characters in the one-hot encoding
n_chars = len(vocab)
```
Let's now create the input and output to our network. We'll use placeholders and feed these in later. The size of these need to be [`batch_size`, `sequence_length`]. We'll then see how to build the network in between.
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
X = tf.placeholder(tf.int32, shape=..., name='X')
# We'll have a placeholder for our true outputs
Y = tf.placeholder(tf.int32, shape=..., name='Y')
```
The first thing we need to do is convert each of our `sequence_length` vectors in our batch to `n_cells` LSTM cells. We use a lookup table to find the value in `X` and use this as the input to `n_cells` LSTM cells. Our lookup table has `n_chars` possible elements and connects each character to `n_cells` cells. We create our lookup table using `tf.get_variable` and then the function `tf.nn.embedding_lookup` to connect our `X` placeholder to `n_cells` number of neurons.
```
# we first create a variable to take us from our one-hot representation to our LSTM cells
embedding = tf.get_variable("embedding", [n_chars, n_cells])
# And then use tensorflow's embedding lookup to look up the ids in X
Xs = tf.nn.embedding_lookup(embedding, X)
# The resulting lookups are concatenated into a dense tensor
print(Xs.get_shape().as_list())
```
Now recall from the lecture that recurrent neural networks share their weights across timesteps. So we don't want to have one large matrix with every timestep, but instead separate them. We'll use `tf.split` to split our `[batch_size, sequence_length, n_cells]` array in `Xs` into a list of `sequence_length` elements each composed of `[batch_size, n_cells]` arrays. This gives us `sequence_length` number of arrays of `[batch_size, 1, n_cells]`. We then use `tf.squeeze` to remove the 1st index corresponding to the singleton `sequence_length` index, resulting in simply `[batch_size, n_cells]`.
```
with tf.name_scope('reslice'):
Xs = [tf.squeeze(seq, [1])
for seq in tf.split(1, sequence_length, Xs)]
```
With each of our timesteps split up, we can now connect them to a set of LSTM recurrent cells. We tell the `tf.nn.rnn_cell.BasicLSTMCell` method how many cells we want, i.e. how many neurons there are, and we also specify that our state will be stored as a tuple. This state defines the internal state of the cells as well as the connection from the previous timestep. We can also pass a value for the `forget_bias`. Be sure to experiment with this parameter as it can significantly effect performance (e.g. Gers, Felix A, Schmidhuber, Jurgen, and Cummins, Fred. Learning to forget: Continual prediction with lstm. Neural computation, 12(10):2451–2471, 2000).
```
cells = tf.nn.rnn_cell.BasicLSTMCell(num_units=n_cells, state_is_tuple=True, forget_bias=1.0)
```
Let's take a look at the cell's state size:
```
cells.state_size
```
`c` defines the internal memory and `h` the output. We'll have as part of our `cells`, both an `initial_state` and a `final_state`. These will become important during inference and we'll see how these work more then. For now, we'll set the `initial_state` to all zeros using the convenience function provided inside our `cells` object, `zero_state`:
```
initial_state = cells.zero_state(tf.shape(X)[0], tf.float32)
```
Looking at what this does, we can see that it creates a `tf.Tensor` of zeros for our `c` and `h` states for each of our `n_cells` and stores this as a tuple inside the `LSTMStateTuple` object:
```
initial_state
```
So far, we have created a single layer of LSTM cells composed of `n_cells` number of cells. If we want another layer, we can use the `tf.nn.rnn_cell.MultiRNNCell` method, giving it our current cells, and a bit of pythonery to multiply our cells by the number of layers we want. We'll then update our `initial_state` variable to include the additional cells:
```
cells = tf.nn.rnn_cell.MultiRNNCell(
[cells] * n_layers, state_is_tuple=True)
initial_state = cells.zero_state(tf.shape(X)[0], tf.float32)
```
Now if we take a look at our `initial_state`, we should see one `LSTMStateTuple` for each of our layers:
```
initial_state
```
So far, we haven't connected our recurrent cells to anything. Let's do this now using the `tf.nn.rnn` method. We also pass it our `initial_state` variables. It gives us the `outputs` of the rnn, as well as their states after having been computed. Contrast that with the `initial_state`, which set the LSTM cells to zeros. After having computed something, the cells will all have a different value somehow reflecting the temporal dynamics and expectations of the next input. These will be stored in the `state` tensors for each of our LSTM layers inside a `LSTMStateTuple` just like the `initial_state` variable.
```python
help(tf.nn.rnn)
Help on function rnn in module tensorflow.python.ops.rnn:
rnn(cell, inputs, initial_state=None, dtype=None, sequence_length=None, scope=None)
Creates a recurrent neural network specified by RNNCell `cell`.
The simplest form of RNN network generated is:
state = cell.zero_state(...)
outputs = []
for input_ in inputs:
output, state = cell(input_, state)
outputs.append(output)
return (outputs, state)
However, a few other options are available:
An initial state can be provided.
If the sequence_length vector is provided, dynamic calculation is performed.
This method of calculation does not compute the RNN steps past the maximum
sequence length of the minibatch (thus saving computational time),
and properly propagates the state at an example's sequence length
to the final state output.
The dynamic calculation performed is, at time t for batch row b,
(output, state)(b, t) =
(t >= sequence_length(b))
? (zeros(cell.output_size), states(b, sequence_length(b) - 1))
: cell(input(b, t), state(b, t - 1))
Args:
cell: An instance of RNNCell.
inputs: A length T list of inputs, each a `Tensor` of shape
`[batch_size, input_size]`, or a nested tuple of such elements.
initial_state: (optional) An initial state for the RNN.
If `cell.state_size` is an integer, this must be
a `Tensor` of appropriate type and shape `[batch_size, cell.state_size]`.
If `cell.state_size` is a tuple, this should be a tuple of
tensors having shapes `[batch_size, s] for s in cell.state_size`.
dtype: (optional) The data type for the initial state and expected output.
Required if initial_state is not provided or RNN state has a heterogeneous
dtype.
sequence_length: Specifies the length of each sequence in inputs.
An int32 or int64 vector (tensor) size `[batch_size]`, values in `[0, T)`.
scope: VariableScope for the created subgraph; defaults to "RNN".
Returns:
A pair (outputs, state) where:
- outputs is a length T list of outputs (one for each input), or a nested
tuple of such elements.
- state is the final state
Raises:
TypeError: If `cell` is not an instance of RNNCell.
ValueError: If `inputs` is `None` or an empty list, or if the input depth
(column size) cannot be inferred from inputs via shape inference.
```
Use the help on the functino `tf.nn.rnn` to create the `outputs` and `states` variable as below. We've already created each of the variable you need to use:
<h3><font color='red'>TODO! COMPLETE THIS SECTION!</font></h3>
```
outputs, state = tf.nn.rnn(cell=..., input=..., initial_state=...)
```
Let's take a look at the state now:
```
state
```
Our outputs are returned as a list for each of our timesteps:
```
outputs
```
We'll now stack all our outputs for every timestep. We can treat every observation at each timestep and for each batch using the same weight matrices going forward, since these should all have shared weights. Each timstep for each batch is its own observation. So we'll stack these in a 2d matrix so that we can create our softmax layer:
```
outputs_flat = tf.reshape(tf.concat(1, outputs), [-1, n_cells])
```
Our outputs are now concatenated so that we have [`batch_size * timesteps`, `n_cells`]
```
outputs_flat
```
We now create a softmax layer just like we did in Session 3 and in Session 3's homework. We multiply our final LSTM layer's `n_cells` outputs by a weight matrix to give us `n_chars` outputs. We then scale this output using a `tf.nn.softmax` layer so that they become a probability by exponentially scaling its value and dividing by its sum. We store the softmax probabilities in `probs` as well as keep track of the maximum index in `Y_pred`:
```
with tf.variable_scope('prediction'):
W = tf.get_variable(
"W",
shape=[n_cells, n_chars],
initializer=tf.random_normal_initializer(stddev=0.1))
b = tf.get_variable(
"b",
shape=[n_chars],
initializer=tf.random_normal_initializer(stddev=0.1))
# Find the output prediction of every single character in our minibatch
# we denote the pre-activation prediction, logits.
logits = tf.matmul(outputs_flat, W) + b
# We get the probabilistic version by calculating the softmax of this
probs = tf.nn.softmax(logits)
# And then we can find the index of maximum probability
Y_pred = tf.argmax(probs, 1)
```
To train the network, we'll measure the loss between our predicted outputs and true outputs. We could use the `probs` variable, but we can also make use of `tf.nn.softmax_cross_entropy_with_logits` which will compute the softmax for us. We therefore need to pass in the variable just before the softmax layer, denoted as `logits` (unscaled values). This takes our variable `logits`, the unscaled predicted outputs, as well as our true outputs, `Y`. Before we give it `Y`, we'll need to reshape our true outputs in the same way, [`batch_size` x `timesteps`, `n_chars`]. Luckily, tensorflow provides a convenience for doing this, the `tf.nn.sparse_softmax_cross_entropy_with_logits` function:
```python
help(tf.nn.sparse_softmax_cross_entropy_with_logits)
Help on function sparse_softmax_cross_entropy_with_logits in module tensorflow.python.ops.nn_ops:
sparse_softmax_cross_entropy_with_logits(logits, labels, name=None)
Computes sparse softmax cross entropy between `logits` and `labels`.
Measures the probability error in discrete classification tasks in which the
classes are mutually exclusive (each entry is in exactly one class). For
example, each CIFAR-10 image is labeled with one and only one label: an image
can be a dog or a truck, but not both.
**NOTE:** For this operation, the probability of a given label is considered
exclusive. That is, soft classes are not allowed, and the `labels` vector
must provide a single specific index for the true class for each row of
`logits` (each minibatch entry). For soft softmax classification with
a probability distribution for each entry, see
`softmax_cross_entropy_with_logits`.
**WARNING:** This op expects unscaled logits, since it performs a softmax
on `logits` internally for efficiency. Do not call this op with the
output of `softmax`, as it will produce incorrect results.
A common use case is to have logits of shape `[batch_size, num_classes]` and
labels of shape `[batch_size]`. But higher dimensions are supported.
Args:
logits: Unscaled log probabilities of rank `r` and shape
`[d_0, d_1, ..., d_{r-2}, num_classes]` and dtype `float32` or `float64`.
labels: `Tensor` of shape `[d_0, d_1, ..., d_{r-2}]` and dtype `int32` or
`int64`. Each entry in `labels` must be an index in `[0, num_classes)`.
Other values will result in a loss of 0, but incorrect gradient
computations.
name: A name for the operation (optional).
Returns:
A `Tensor` of the same shape as `labels` and of the same type as `logits`
with the softmax cross entropy loss.
Raises:
ValueError: If logits are scalars (need to have rank >= 1) or if the rank
of the labels is not equal to the rank of the labels minus one.
```
```
with tf.variable_scope('loss'):
# Compute mean cross entropy loss for each output.
Y_true_flat = tf.reshape(tf.concat(1, Y), [-1])
# logits are [batch_size x timesteps, n_chars] and
# Y_true_flat are [batch_size x timesteps]
loss = tf.nn.sparse_softmax_cross_entropy_with_logits(logits, Y_true_flat)
# Compute the mean over our `batch_size` x `timesteps` number of observations
mean_loss = tf.reduce_mean(loss)
```
Finally, we can create an optimizer in much the same way as we've done with every other network. Except, we will also "clip" the gradients of every trainable parameter. This is a hacky way to ensure that the gradients do not grow too large (the literature calls this the "exploding gradient problem"). However, note that the LSTM is built to help ensure this does not happen by allowing the gradient to be "gated". To learn more about this, please consider reading the following material:
http://www.felixgers.de/papers/phd.pdf
https://colah.github.io/posts/2015-08-Understanding-LSTMs/
```
with tf.name_scope('optimizer'):
optimizer = tf.train.AdamOptimizer(learning_rate=0.001)
gradients = []
clip = tf.constant(5.0, name="clip")
for grad, var in optimizer.compute_gradients(mean_loss):
gradients.append((tf.clip_by_value(grad, -clip, clip), var))
updates = optimizer.apply_gradients(gradients)
```
Let's take a look at the graph:
```
nb_utils.show_graph(tf.get_default_graph().as_graph_def())
```
Below is the rest of code we'll need to train the network. I do not recommend running this inside Jupyter Notebook for the entire length of the training because the network can take 1-2 days at least to train, and your browser may very likely complain. Instead, you should write a python script containing the necessary bits of code and run it using the Terminal. We didn't go over how to do this, so I'll leave it for you as an exercise. The next part of this notebook will have you load a pre-trained network.
```
with tf.Session() as sess:
init = tf.initialize_all_variables()
sess.run(init)
cursor = 0
it_i = 0
while it_i < 500:
Xs, Ys = [], []
for batch_i in range(batch_size):
if (cursor + sequence_length) >= len(txt) - sequence_length - 1:
cursor = 0
Xs.append([encoder[ch]
for ch in txt[cursor:cursor + sequence_length]])
Ys.append([encoder[ch]
for ch in txt[cursor + 1: cursor + sequence_length + 1]])
cursor = (cursor + sequence_length)
Xs = np.array(Xs).astype(np.int32)
Ys = np.array(Ys).astype(np.int32)
loss_val, _ = sess.run([mean_loss, updates],
feed_dict={X: Xs, Y: Ys})
if it_i % 100 == 0:
print(it_i, loss_val)
if it_i % 500 == 0:
p = sess.run(probs, feed_dict={X: np.array(Xs[-1])[np.newaxis]})
ps = [np.random.choice(range(n_chars), p=p_i.ravel())
for p_i in p]
p = [np.argmax(p_i) for p_i in p]
if isinstance(txt[0], str):
print('original:', "".join(
[decoder[ch] for ch in Xs[-1]]))
print('synth(samp):', "".join(
[decoder[ch] for ch in ps]))
print('synth(amax):', "".join(
[decoder[ch] for ch in p]))
else:
print([decoder[ch] for ch in ps])
it_i += 1
```
<a name="part-5---pretrained-char-rnn-of-donald-trump"></a>
# Part 5 - Pretrained Char-RNN of Donald Trump
Rather than stick around to let a model train, let's now explore one I've trained for you Donald Trump. If you've trained your own model on your own text corpus then great! You should be able to use that in place of the one I've provided and still continue with the rest of the notebook.
For the Donald Trump corpus, there are a lot of video transcripts that you can find online. I've searched for a few of these, put them in a giant text file, made everything lowercase, and removed any extraneous letters/symbols to help reduce the vocabulary (not that it's not very large to begin with, ha).
I used the code exactly as above to train on the text I gathered and left it to train for about 2 days. The only modification is that I also used "dropout" which you can see in the libs/charrnn.py file. Let's explore it now and we'll see how we can play with "sampling" the model to generate new phrases, and how to "prime" the model (a psychological term referring to when someone is exposed to something shortly before another event).
First, let's clean up any existing graph:
```
tf.reset_default_graph()
```
<a name="getting-the-trump-data"></a>
## Getting the Trump Data
Now let's load the text. This is included in the repo or can be downloaded from:
```
with open('trump.txt', 'r') as fp:
txt = fp.read()
```
Let's take a look at what's going on in here:
```
txt[:100]
```
<a name="basic-text-analysis"></a>
## Basic Text Analysis
We can do some basic data analysis to get a sense of what kind of vocabulary we're working with. It's really important to look at your data in as many ways as possible. This helps ensure there isn't anything unexpected going on. Let's find every unique word he uses:
```
words = set(txt.split(' '))
words
```
Now let's count their occurrences:
```
counts = {word_i: 0 for word_i in words}
for word_i in txt.split(' '):
counts[word_i] += 1
counts
```
We can sort this like so:
```
[(word_i, counts[word_i]) for word_i in sorted(counts, key=counts.get, reverse=True)]
```
As we should expect, "the" is the most common word, as it is in the English language: https://en.wikipedia.org/wiki/Most_common_words_in_English
<a name="loading-the-pre-trained-trump-model"></a>
## Loading the Pre-trained Trump Model
Let's load the pretrained model. Rather than provide a tfmodel export, I've provided the checkpoint so you can also experiment with training it more if you wish. We'll rebuild the graph using the `charrnn` module in the `libs` directory:
```
from libs import charrnn
```
Let's get the checkpoint and build the model then restore the variables from the checkpoint. The only parameters of consequence are `n_layers` and `n_cells` which define the total size and layout of the model. The rest are flexible. We'll set the `batch_size` and `sequence_length` to 1, meaning we can feed in a single character at a time only, and get back 1 character denoting the very next character's prediction.
```
ckpt_name = 'trump.ckpt'
g = tf.Graph()
n_layers = 3
n_cells = 512
with tf.Session(graph=g) as sess:
model = charrnn.build_model(txt=txt,
batch_size=1,
sequence_length=1,
n_layers=n_layers,
n_cells=n_cells,
gradient_clip=10.0)
saver = tf.train.Saver()
if os.path.exists(ckpt_name):
saver.restore(sess, ckpt_name)
print("Model restored.")
```
Let's now take a look at the model:
```
nb_utils.show_graph(g.as_graph_def())
n_iterations = 100
```
<a name="inference-keeping-track-of-the-state"></a>
## Inference: Keeping Track of the State
Now recall from Part 4 when we created our LSTM network, we had an `initial_state` variable which would set the LSTM's `c` and `h` state vectors, as well as the final output state which was the output of the `c` and `h` state vectors after having passed through the network. When we input to the network some letter, say 'n', we can set the `initial_state` to zeros, but then after having input the letter `n`, we'll have as output a new state vector for `c` and `h`. On the next letter, we'll then want to set the `initial_state` to this new state, and set the input to the previous letter's output. That is how we ensure the network keeps track of time and knows what has happened in the past, and let it continually generate.
```
curr_states = None
g = tf.Graph()
with tf.Session(graph=g) as sess:
model = charrnn.build_model(txt=txt,
batch_size=1,
sequence_length=1,
n_layers=n_layers,
n_cells=n_cells,
gradient_clip=10.0)
saver = tf.train.Saver()
if os.path.exists(ckpt_name):
saver.restore(sess, ckpt_name)
print("Model restored.")
# Get every tf.Tensor for the initial state
init_states = []
for s_i in model['initial_state']:
init_states.append(s_i.c)
init_states.append(s_i.h)
# Similarly, for every state after inference
final_states = []
for s_i in model['final_state']:
final_states.append(s_i.c)
final_states.append(s_i.h)
# Let's start with the letter 't' and see what comes out:
synth = [[encoder[' ']]]
for i in range(n_iterations):
# We'll create a feed_dict parameter which includes what to
# input to the network, model['X'], as well as setting
# dropout to 1.0, meaning no dropout.
feed_dict = {model['X']: [synth[-1]],
model['keep_prob']: 1.0}
# Now we'll check if we currently have a state as a result
# of a previous inference, and if so, add to our feed_dict
# parameter the mapping of the init_state to the previous
# output state stored in "curr_states".
if curr_states:
feed_dict.update(
{init_state_i: curr_state_i
for (init_state_i, curr_state_i) in
zip(init_states, curr_states)})
# Now we can infer and see what letter we get
p = sess.run(model['probs'], feed_dict=feed_dict)[0]
# And make sure we also keep track of the new state
curr_states = sess.run(final_states, feed_dict=feed_dict)
# Find the most likely character
p = np.argmax(p)
# Append to string
synth.append([p])
# Print out the decoded letter
print(model['decoder'][p], end='')
sys.stdout.flush()
```
<a name="probabilistic-sampling"></a>
## Probabilistic Sampling
Run the above cell a couple times. What you should find is that it is deterministic. We always pick *the* most likely character. But we can do something else which will make things less deterministic and a bit more interesting: we can sample from our probabilistic measure from our softmax layer. This means if we have the letter 'a' as 0.4, and the letter 'o' as 0.2, we'll have a 40% chance of picking the letter 'a', and 20% chance of picking the letter 'o', rather than simply always picking the letter 'a' since it is the most probable.
```
curr_states = None
g = tf.Graph()
with tf.Session(graph=g) as sess:
model = charrnn.build_model(txt=txt,
batch_size=1,
sequence_length=1,
n_layers=n_layers,
n_cells=n_cells,
gradient_clip=10.0)
saver = tf.train.Saver()
if os.path.exists(ckpt_name):
saver.restore(sess, ckpt_name)
print("Model restored.")
# Get every tf.Tensor for the initial state
init_states = []
for s_i in model['initial_state']:
init_states.append(s_i.c)
init_states.append(s_i.h)
# Similarly, for every state after inference
final_states = []
for s_i in model['final_state']:
final_states.append(s_i.c)
final_states.append(s_i.h)
# Let's start with the letter 't' and see what comes out:
synth = [[encoder[' ']]]
for i in range(n_iterations):
# We'll create a feed_dict parameter which includes what to
# input to the network, model['X'], as well as setting
# dropout to 1.0, meaning no dropout.
feed_dict = {model['X']: [synth[-1]],
model['keep_prob']: 1.0}
# Now we'll check if we currently have a state as a result
# of a previous inference, and if so, add to our feed_dict
# parameter the mapping of the init_state to the previous
# output state stored in "curr_states".
if curr_states:
feed_dict.update(
{init_state_i: curr_state_i
for (init_state_i, curr_state_i) in
zip(init_states, curr_states)})
# Now we can infer and see what letter we get
p = sess.run(model['probs'], feed_dict=feed_dict)[0]
# And make sure we also keep track of the new state
curr_states = sess.run(final_states, feed_dict=feed_dict)
# Now instead of finding the most likely character,
# we'll sample with the probabilities of each letter
p = p.astype(np.float64)
p = np.random.multinomial(1, p.ravel() / p.sum())
p = np.argmax(p)
# Append to string
synth.append([p])
# Print out the decoded letter
print(model['decoder'][p], end='')
sys.stdout.flush()
```
<a name="inference-temperature"></a>
## Inference: Temperature
When performing probabilistic sampling, we can also use a parameter known as temperature which comes from simulated annealing. The basic idea is that as the temperature is high and very hot, we have a lot more free energy to use to jump around more, and as we cool down, we have less energy and then become more deterministic. We can use temperature by scaling our log probabilities like so:
```
temperature = 0.5
curr_states = None
g = tf.Graph()
with tf.Session(graph=g) as sess:
model = charrnn.build_model(txt=txt,
batch_size=1,
sequence_length=1,
n_layers=n_layers,
n_cells=n_cells,
gradient_clip=10.0)
saver = tf.train.Saver()
if os.path.exists(ckpt_name):
saver.restore(sess, ckpt_name)
print("Model restored.")
# Get every tf.Tensor for the initial state
init_states = []
for s_i in model['initial_state']:
init_states.append(s_i.c)
init_states.append(s_i.h)
# Similarly, for every state after inference
final_states = []
for s_i in model['final_state']:
final_states.append(s_i.c)
final_states.append(s_i.h)
# Let's start with the letter 't' and see what comes out:
synth = [[encoder[' ']]]
for i in range(n_iterations):
# We'll create a feed_dict parameter which includes what to
# input to the network, model['X'], as well as setting
# dropout to 1.0, meaning no dropout.
feed_dict = {model['X']: [synth[-1]],
model['keep_prob']: 1.0}
# Now we'll check if we currently have a state as a result
# of a previous inference, and if so, add to our feed_dict
# parameter the mapping of the init_state to the previous
# output state stored in "curr_states".
if curr_states:
feed_dict.update(
{init_state_i: curr_state_i
for (init_state_i, curr_state_i) in
zip(init_states, curr_states)})
# Now we can infer and see what letter we get
p = sess.run(model['probs'], feed_dict=feed_dict)[0]
# And make sure we also keep track of the new state
curr_states = sess.run(final_states, feed_dict=feed_dict)
# Now instead of finding the most likely character,
# we'll sample with the probabilities of each letter
p = p.astype(np.float64)
p = np.log(p) / temperature
p = np.exp(p) / np.sum(np.exp(p))
p = np.random.multinomial(1, p.ravel() / p.sum())
p = np.argmax(p)
# Append to string
synth.append([p])
# Print out the decoded letter
print(model['decoder'][p], end='')
sys.stdout.flush()
```
<a name="inference-priming"></a>
## Inference: Priming
Let's now work on "priming" the model with some text, and see what kind of state it is in and leave it to synthesize from there. We'll do more or less what we did before, but feed in our own text instead of the last letter of the synthesis from the model.
```
prime = "obama"
temperature = 1.0
curr_states = None
n_iterations = 500
g = tf.Graph()
with tf.Session(graph=g) as sess:
model = charrnn.build_model(txt=txt,
batch_size=1,
sequence_length=1,
n_layers=n_layers,
n_cells=n_cells,
gradient_clip=10.0)
saver = tf.train.Saver()
if os.path.exists(ckpt_name):
saver.restore(sess, ckpt_name)
print("Model restored.")
# Get every tf.Tensor for the initial state
init_states = []
for s_i in model['initial_state']:
init_states.append(s_i.c)
init_states.append(s_i.h)
# Similarly, for every state after inference
final_states = []
for s_i in model['final_state']:
final_states.append(s_i.c)
final_states.append(s_i.h)
# Now we'll keep track of the state as we feed it one
# letter at a time.
curr_states = None
for ch in prime:
feed_dict = {model['X']: [[model['encoder'][ch]]],
model['keep_prob']: 1.0}
if curr_states:
feed_dict.update(
{init_state_i: curr_state_i
for (init_state_i, curr_state_i) in
zip(init_states, curr_states)})
# Now we can infer and see what letter we get
p = sess.run(model['probs'], feed_dict=feed_dict)[0]
p = p.astype(np.float64)
p = np.log(p) / temperature
p = np.exp(p) / np.sum(np.exp(p))
p = np.random.multinomial(1, p.ravel() / p.sum())
p = np.argmax(p)
# And make sure we also keep track of the new state
curr_states = sess.run(final_states, feed_dict=feed_dict)
# Now we're ready to do what we were doing before but with the
# last predicted output stored in `p`, and the current state of
# the model.
synth = [[p]]
print(prime + model['decoder'][p], end='')
for i in range(n_iterations):
# Input to the network
feed_dict = {model['X']: [synth[-1]],
model['keep_prob']: 1.0}
# Also feed our current state
feed_dict.update(
{init_state_i: curr_state_i
for (init_state_i, curr_state_i) in
zip(init_states, curr_states)})
# Inference
p = sess.run(model['probs'], feed_dict=feed_dict)[0]
# Keep track of the new state
curr_states = sess.run(final_states, feed_dict=feed_dict)
# Sample
p = p.astype(np.float64)
p = np.log(p) / temperature
p = np.exp(p) / np.sum(np.exp(p))
p = np.random.multinomial(1, p.ravel() / p.sum())
p = np.argmax(p)
# Append to string
synth.append([p])
# Print out the decoded letter
print(model['decoder'][p], end='')
sys.stdout.flush()
```
<a name="assignment-submission"></a>
# Assignment Submission
After you've completed both notebooks, create a zip file of the current directory using the code below. This code will make sure you have included this completed ipython notebook and the following files named exactly as:
session-5/
session-5-part-1.ipynb
session-5-part-2.ipynb
vaegan.gif
You'll then submit this zip file for your third assignment on Kadenze for "Assignment 5: Generative Adversarial Networks and Recurrent Neural Networks"! If you have any questions, remember to reach out on the forums and connect with your peers or with me.
To get assessed, you'll need to be a premium student! This will allow you to build an online portfolio of all of your work and receive grades. If you aren't already enrolled as a student, register now at http://www.kadenze.com/ and join the #CADL community to see what your peers are doing! https://www.kadenze.com/courses/creative-applications-of-deep-learning-with-tensorflow/info
Also, if you share any of the GIFs on Facebook/Twitter/Instagram/etc..., be sure to use the #CADL hashtag so that other students can find your work!
```
utils.build_submission('session-5.zip',
('vaegan.gif',
'session-5-part-1.ipynb',
'session-5-part-2.ipynb'))
```
| github_jupyter |
```
import scipy.sparse.linalg as spsplin
import numpy as np
import matplotlib.pyplot as plt
# Convergence of CG for different spectrum
n = 1000
A = np.random.randn(n, n)
# A = A.T @ A
Q, _ = np.linalg.qr(A)
A = Q @ np.diag([1, 10] + [100 + i for i in range(n-2)]) @ Q.T
eigvals = np.linalg.eigvalsh(A)
print(min(eigvals), max(eigvals))
plt.scatter(eigvals, np.ones(n))
q = np.linalg.cond(A)
print(q)
print((q - 1) / (q + 1))
print((np.sqrt(q) - 1) / (np.sqrt(q) + 1))
```
- Richardson iteration convergence
$$ \|x_{k+1} - x^* \|_2 \leq C \left(\frac{\kappa - 1}{\kappa+1}\right)^k $$
- Chebyshev iteration convergence
$$ \|x_{k+1} - x^* \|_2 \leq C \left(\frac{\sqrt{\kappa} - 1}{\sqrt{\kappa}+1}\right)^k $$
```
rhs = np.random.randn(n)
x0 = np.random.randn(n)
conv_cg = []
def cl_cg(x):
conv_cg.append(x.copy())
x_cg, info = spsplin.cg(A, rhs, callback=cl_cg, x0=x0, tol=1e-6)
np.linalg.norm(A @ x_cg - rhs)
def richardson(A, b, tau, x0, tol, max_iter):
x = x0.copy()
conv = []
for i in range(max_iter):
x = x - tau * (A @ x - b)
conv.append(x)
if np.linalg.norm(A @ x - b) < tol:
break
return x, conv
x_rich, conv_rich = richardson(A, rhs, 2 / (min(eigvals) + max(eigvals)), x0,
1e-6, 1000)
print(np.linalg.norm(A @ x_rich - rhs))
plt.semilogy([np.linalg.norm(A @ x - rhs) for x in conv_cg], label="CG")
plt.semilogy([np.linalg.norm(A @ x - rhs) for x in conv_rich], label="Richardson")
plt.legend(fontsize=20)
plt.grid(True)
plt.xlabel("Number of iterations", fontsize=24)
plt.ylabel("$||Ax_k - b||_2$", fontsize=24)
niters = 256
roots = [np.cos((np.pi * (2 * i + 1)) / (2 * niters)) for i in range(niters)]
lam_max = max(eigvals)
lam_min = min(eigvals)
taus = [(lam_max + lam_min - (lam_min - lam_max) * r) / 2 for r in roots]
x = np.zeros(n)
r = A.dot(x) - rhs
res_cheb = [np.linalg.norm(r)]
# Implementation may be non-optimal if number of iterations is not power of two
def good_shuffle(idx):
if len(idx) == 1:
return idx
else:
new_len = int(np.ceil((len(idx) / 2)))
new_idx = good_shuffle(idx[:new_len])
res_perm = []
perm_count = 0
for i in new_idx:
res_perm.append(i)
perm_count += 1
if perm_count == len(idx):
break
res_perm.append(len(idx) + 1 - i)
perm_count += 1
if perm_count == len(idx):
break
return res_perm
good_perm = good_shuffle([i for i in range(1, niters+1)])
# good_perm = [i for i in range(niters, 0, -1)]
# good_perm = [i for i in range(niters)]
# good_perm = np.random.permutation([i for i in range(1, niters+1)])
for i in range(niters):
x = x - 1.0/taus[good_perm[i] - 1] * r
r = A.dot(x) - rhs
res_cheb.append(np.linalg.norm(r))
plt.semilogy([np.linalg.norm(A @ x - rhs) for x in conv_cg], label="CG")
plt.semilogy([np.linalg.norm(A @ x - rhs) for x in conv_rich], label="Richardson")
plt.semilogy(res_cheb, label="Chebyshev")
plt.legend(fontsize=20)
plt.grid(True)
plt.xlabel("Number of iterations", fontsize=24)
plt.ylabel("$||Ax_k - b||_2$", fontsize=24)
%timeit np.linalg.solve(A, rhs)
%timeit spsplin.cg(A, rhs)
```
| github_jupyter |
# Multiple Comparisons
_This setup code is required to run in an IPython notebook_
```
%matplotlib inline
import matplotlib.pyplot as plt
import seaborn
seaborn.set_style("darkgrid")
plt.rc("figure", figsize=(16, 6))
plt.rc("savefig", dpi=90)
plt.rc("font", family="sans-serif")
plt.rc("font", size=14)
# Reproducability
import numpy as np
gen = np.random.default_rng(23456)
# Common seed used throughout
seed = gen.integers(0, 2 ** 31 - 1)
```
The multiple comparison procedures all allow for examining aspects of superior predictive ability. There are three available:
* `SPA` - The test of Superior Predictive Ability, also known as the Reality Check (and accessible as `RealityCheck`) or the bootstrap data snooper, examines whether any model in a set of models can outperform a benchmark.
* `StepM` - The stepwise multiple testing procedure uses sequential testing to determine which models are superior to a benchmark.
* `MCS` - The model confidence set which computes the set of models which with performance indistinguishable from others in the set.
All procedures take **losses** as inputs. That is, smaller values are preferred to larger values. This is common when evaluating forecasting models where the loss function is usually defined as a positive function of the forecast error that is increasing in the absolute error. Leading examples are Mean Square Error (MSE) and Mean Absolute Deviation (MAD).
## The test of Superior Predictive Ability (SPA)
This procedure requires a $t$-element array of benchmark losses and a $t$ by $k$-element array of model losses. The null hypothesis is that no model is better than the benchmark, or
$$ H_0: \max_i E[L_i] \geq E[L_{bm}] $$
where $L_i$ is the loss from model $i$ and $L_{bm}$ is the loss from the benchmark model.
This procedure is normally used when there are many competing forecasting models such as in the study of technical trading rules. The example below will make use of a set of models which are all equivalently good to a benchmark model and will serve as a *size study*.
#### Study Design
The study will make use of a measurement error in predictors to produce a large set of correlated variables that all have equal expected MSE. The benchmark will have identical measurement error and so all models have the same expected loss, although will have different forecasts.
The first block computed the series to be forecast.
```
import statsmodels.api as sm
from numpy.random import randn
t = 1000
factors = randn(t, 3)
beta = np.array([1, 0.5, 0.1])
e = randn(t)
y = factors.dot(beta)
```
The next block computes the benchmark factors and the model factors by contaminating the original factors with noise. The models are estimated on the first 500 observations and predictions are made for the second 500. Finally, losses are constructed from these predictions.
```
# Measurement noise
bm_factors = factors + randn(t, 3)
# Fit using first half, predict second half
bm_beta = sm.OLS(y[:500], bm_factors[:500]).fit().params
# MSE loss
bm_losses = (y[500:] - bm_factors[500:].dot(bm_beta)) ** 2.0
# Number of models
k = 500
model_factors = np.zeros((k, t, 3))
model_losses = np.zeros((500, k))
for i in range(k):
# Add measurement noise
model_factors[i] = factors + randn(1000, 3)
# Compute regression parameters
model_beta = sm.OLS(y[:500], model_factors[i, :500]).fit().params
# Prediction and losses
model_losses[:, i] = (y[500:] - model_factors[i, 500:].dot(model_beta)) ** 2.0
```
Finally the SPA can be used. The SPA requires the **losses** from the benchmark and the models as inputs. Other inputs allow the bootstrap sued to be changed or for various options regarding studentization of the losses. `compute` does the real work, and then `pvalues` contains the probability that the null is true given the realizations.
In this case, one would not reject. The three p-values correspond to different re-centerings of the losses. In general, the `consistent` p-value should be used. It should always be the case that
$$ lower \leq consistent \leq upper .$$
See the original papers for more details.
```
from arch.bootstrap import SPA
spa = SPA(bm_losses, model_losses, seed=seed)
spa.compute()
spa.pvalues
```
The same blocks can be repeated to perform a simulation study. Here I only use 100 replications since this should complete in a reasonable amount of time. Also I set `reps=250` to limit the number of bootstrap replications in each application of the SPA (the default is a more reasonable 1000).
```
# Save the pvalues
pvalues = []
b = 100
seeds = gen.integers(0, 2 ** 31 - 1, b)
# Repeat 100 times
for j in range(b):
if j % 10 == 0:
print(j)
factors = randn(t, 3)
beta = np.array([1, 0.5, 0.1])
e = randn(t)
y = factors.dot(beta)
# Measurement noise
bm_factors = factors + randn(t, 3)
# Fit using first half, predict second half
bm_beta = sm.OLS(y[:500], bm_factors[:500]).fit().params
# MSE loss
bm_losses = (y[500:] - bm_factors[500:].dot(bm_beta)) ** 2.0
# Number of models
k = 500
model_factors = np.zeros((k, t, 3))
model_losses = np.zeros((500, k))
for i in range(k):
model_factors[i] = factors + randn(1000, 3)
model_beta = sm.OLS(y[:500], model_factors[i, :500]).fit().params
# MSE loss
model_losses[:, i] = (y[500:] - model_factors[i, 500:].dot(model_beta)) ** 2.0
# Lower the bootstrap replications to 250
spa = SPA(bm_losses, model_losses, reps=250, seed=seeds[j])
spa.compute()
pvalues.append(spa.pvalues)
```
Finally the pvalues can be plotted. Ideally they should form a $45^o$ line indicating the size is correct. Both the consistent and upper perform well. The lower has too many small p-values.
```
import pandas as pd
pvalues = pd.DataFrame(pvalues)
for col in pvalues:
values = pvalues[col].values
values.sort()
pvalues[col] = values
# Change the index so that the x-values are between 0 and 1
pvalues.index = np.linspace(0.005, 0.995, 100)
fig = pvalues.plot()
```
#### Power
The SPA also has power to reject then the null is violated. The simulation will be modified so that the amount of measurement error differs across models, and so that some models are actually better than the benchmark. The p-values should be small indicating rejection of the null.
```
# Number of models
k = 500
model_factors = np.zeros((k, t, 3))
model_losses = np.zeros((500, k))
for i in range(k):
scale = (2500.0 - i) / 2500.0
model_factors[i] = factors + scale * randn(1000, 3)
model_beta = sm.OLS(y[:500], model_factors[i, :500]).fit().params
# MSE loss
model_losses[:, i] = (y[500:] - model_factors[i, 500:].dot(model_beta)) ** 2.0
spa = SPA(bm_losses, model_losses, seed=seed)
spa.compute()
spa.pvalues
```
Here the average losses are plotted. The higher index models are clearly better than the lower index models -- and the benchmark model (which is identical to model.0).
```
model_losses = pd.DataFrame(model_losses, columns=["model." + str(i) for i in range(k)])
avg_model_losses = pd.DataFrame(model_losses.mean(0), columns=["Average loss"])
fig = avg_model_losses.plot(style=["o"])
```
## Stepwise Multiple Testing (StepM)
Stepwise Multiple Testing is similar to the SPA and has the same null. The primary difference is that it identifies the set of models which are better than the benchmark, rather than just asking the basic question if any model is better.
```
from arch.bootstrap import StepM
stepm = StepM(bm_losses, model_losses)
stepm.compute()
print("Model indices:")
print([model.split(".")[1] for model in stepm.superior_models])
better_models = pd.concat([model_losses.mean(0), model_losses.mean(0)], 1)
better_models.columns = ["Same or worse", "Better"]
better = better_models.index.isin(stepm.superior_models)
worse = np.logical_not(better)
better_models.loc[better, "Same or worse"] = np.nan
better_models.loc[worse, "Better"] = np.nan
fig = better_models.plot(style=["o", "s"], rot=270)
```
## The Model Confidence Set
The model confidence set takes a set of **losses** as its input and finds the set which are not statistically different from each other while controlling the familywise error rate. The primary output is a set of p-values, where models with a pvalue above the size are in the MCS. Small p-values indicate that the model is easily rejected from the set that includes the best.
```
from arch.bootstrap import MCS
# Limit the size of the set
losses = model_losses.iloc[:, ::20]
mcs = MCS(losses, size=0.10)
mcs.compute()
print("MCS P-values")
print(mcs.pvalues)
print("Included")
included = mcs.included
print([model.split(".")[1] for model in included])
print("Excluded")
excluded = mcs.excluded
print([model.split(".")[1] for model in excluded])
status = pd.DataFrame(
[losses.mean(0), losses.mean(0)], index=["Excluded", "Included"]
).T
status.loc[status.index.isin(included), "Excluded"] = np.nan
status.loc[status.index.isin(excluded), "Included"] = np.nan
fig = status.plot(style=["o", "s"])
```
| github_jupyter |
# Lazada Product Review Data In Brief
----
## Description
First step of data exploration. Understanding data dan explore macro-characteristic of the data that we have to deal with
## Author
+ [Christian Wibisono](https://www.kaggle.com/christianwbsn)
```
import pandas as pd
import numpy as np
raw = pd.read_csv( "../data/raw/lazada_reviews.csv")
raw.head()
```
As we can see the data contain 2 columns *rating* and *review*
```
raw.info()
```
1. rating: numerical (1-5)
2. review: text
----
We have 280803 data points but there are 60570 missing values in review column.
Since our objective is to do **sentiment analysis** on text data, those missing value will be dropped later.
Those missing values couldn't tell us anything about the data
## Characteristic Of Marketplace Product Review in Bahasa Indonesia
```
raw['review'][:10]
```
## 1.Informal words
```
print(raw['review'][2])
print("---------------")
print(raw['review'][7])
```
Data contain words in slang words such as:
+ *ga* --> tidak
+ *mantab* --> mantap
+ *nyampe* --> sampai
## 2. Abbreviation
```
print(raw['review'][10])
print("---------------")
print(raw['review'][11])
```
Data contain abbreviation i.e:
+ *knp* --> kenapa
+ *yg* --> yang
+ *tp* --> tapi
## 3. Foreign Word
```
print(raw['review'][30])
print("---------------")
print(raw['review'][19])
```
Data contain foreign languange i.e:
+ *seller* --> penjual
+ *packing* --> pengemasan
+ *recommended* --> direkomendasikan
+ *happy birthday* --> selamat ulang tahun
## 4. Domain Spesific Word
```
print(raw['review'][10])
print("---------------")
print(raw['review'][14])
```
Our domain: Marketplace Product Review
Data contain domain spesific word i.e:
+ *flash sale*
+ *xiaomi*
+ *lazada*
+ *redmi*
+ *powerbank*
## 5. Emoji
```
print(raw['review'][12])
print("---------------")
print(raw['review'][31])
```
# 😃
This is the example of emoji found in our dataset
## 6. Emoticon
```
print(raw['review'][82])
print("---------------")
print(raw['review'][280323])
```
## :( sad or happy :)
## 7. Interjection
```
print(raw['review'][12983])
print("---------------")
print(raw['review'][222677])
```
Data contain interjection such as:
+ **wkwkwk**
+ **Hahahaha**
## 8. Typographical Error
```
print(raw['review'][15])
print("---------------")
print(raw['review'][8])
```
There are so **many** typographical error in our dataset. Single review can contain more than one typos
+ kadang**h** --> kadang(addition)
+ can**s**el --> cancel(replace)
+ pen**it**iman --> pengiriman(deletion,transposition,replace)
+ melalui**n** --> melalui(addition)
## 9. Irrelevant Text
```
print(raw['review'][1639])
print("---------------")
print(raw['review'][224544])
print("---------------")
print(raw['review'][80])
```
## 10. Completely Not in Bahasa
```
print(raw['review'][75])
print("---------------")
print(raw['review'][551])
```
## 11. Creative Isn't it?
```
print(raw['review'][38])
print("---------------")
print(raw['review'][21677])
```
## 12. Local *'Slang'* Words
```
print(raw['review'][17719])
print("---------------")
print(raw['review'][17025])
```
## 13. Unstandardized Words
```
print(raw['review'][0])
print("---------------")
print(raw['review'][5])
```
+ Sangatttttt --> sangat
+ lamaaaaaa --> lama
+ leleeeeeetttt --> lelet
+ ituuuuuuuu --> itu
## 14. Meaningless Words
```
print(raw['review'][224432])
print("---------------")
print(raw['review'][279396])
print("---------------")
print(raw['review'][279404])
print("---------------")
print(raw['review'][279403])
```
Know the meaning of those word? email me: [christian.wibisono7@gmail.com](mailto:christian.wibisono7@gmail.com)
## 15. Inconsistent Rating and Review
```
print("Rating " + str(raw['rating'][6970]) + ":" + raw['review'][6970])
print("---------------")
print("Rating " + str(raw['rating'][15908]) + ":" + raw['review'][15908])
```
Both condition appears in our dataset:
+ Positive review with negative rating
+ Negative review with positive rating
This condition can led to incorrectly labelled data
## 16. Reversed Words
```
print(raw['review'][228587])
print("---------------")
```
+ Kuy --> Yuk (reversed letter by letter)
| github_jupyter |
## Lecture 12:
* We are going to do RNN in this session.
* Very powerful concept. Hidden state of one cell is used as an input for another cell in addition to the original input.
For `RNN` the arguments that are passed are:
* `input_size`
* `hidden_size`
```
# Importing stock libraries
import numpy as np
import pandas as pd
import torch
from torch.utils.data import DataLoader, Dataset
```
### Experiment 1:
* Feeding `H E L L O` to RNN
* We will use 1 hot vector encoding for this
* $h = [1,0,0,0]$
* $e = [0,1,0,0]$
* $l = [0,0,1,0]$
* $o = [0,0,0,1]$
This is feeded to the network/cell one by one. Hence,
* `input size = 4`
* `hidden size = 2`
```
h = [1,0,0,0]
e = [0,1,0,0]
l = [0,0,1,0]
o = [0,0,0,1]
# Create an RNN cell with the desited input size and hidden size
cell = torch.nn.RNN(input_size=4, hidden_size=2,batch_first=True)
# Creating the one letter input
h = [1,0,0,0]
inputs = torch.Tensor([[h]])
print('Input size', inputs.size())
# initialize the hidden state
# (num_layers*num_directios, batch, hidden_size)
hidden = torch.Tensor(torch.rand(1,1,2))
# Feed one element at a time
# after each step, hidden contains the hidden state
out, hidden = cell(inputs, hidden)
print('Out', out.data)
print('Type', out.dtype)
print('Size',out.size())
print('Hidden', hidden.data)
# Create an RNN cell with the desited input size and hidden size, this time we are entering more than 1 charecter
cell = torch.nn.RNN(input_size=4, hidden_size=2,batch_first=True)
# hidden_size =2
# batch_size=1
# sequence_length=5
inputs = torch.Tensor([[h,e,l,l,o]])
print('Input size', inputs.size())
# initialize the hidden state
# (num_layers*num_directios, batch, hidden_size)
hidden = torch.Tensor(torch.rand(1,1,2))
# Feed one element at a time
# after each step, hidden contains the hidden state
out, hidden = cell(inputs, hidden)
print('Out', out.data)
print('Type', out.dtype)
print('Size',out.size())
print('Hidden', hidden.data)
# Create an RNN cell with the desited input size and hidden size, this time we are entering more than 1 word.
# hidden_size =2
# batch_size=2
# sequence_length=5
cell = torch.nn.RNN(input_size=4, hidden_size=2,batch_first=True)
# Creating the one letter input
inputs = torch.Tensor(
[
[h,e,l,l,o],
[e,l,l,o,l],
[l,l,e,e,l]
]
)
print('Input size', inputs.size())
# initialize the hidden state
# (num_layers*num_directios, batch, hidden_size)
hidden = torch.Tensor(torch.rand(1,3,2))
# Feed one element at a time
# after each step, hidden contains the hidden state
out, hidden = cell(inputs, hidden)
print('Out', out.data)
print('Type', out.dtype)
print('Size',out.size())
print('Hidden', hidden.data)
```
### Experiment 2:
* We will be feeding the string `hihell` to the network such that it gives us the output `ihello` basically predicting the next charter
* This is a sequence classification.
```
# Creating a project to convert hihell -> ihello
# Data prepration
idx2char = ['h', 'i', 'e', 'l', 'o']
x_data = [0,1,0,2,3,3] #hihell
x_data = [[0, 1, 0, 2, 3, 3]] # hihell
x_one_hot = [[[1, 0, 0, 0, 0], # h 0
[0, 1, 0, 0, 0], # i 1
[1, 0, 0, 0, 0], # h 0
[0, 0, 1, 0, 0], # e 2
[0, 0, 0, 1, 0], # l 3
[0, 0, 0, 1, 0]]] # l 3
y_data = [1, 0, 2, 3, 3, 4] # ihello
inputs = torch.Tensor(x_one_hot)
labels = torch.LongTensor(y_data)
# Parameters
num_classes =5
input_size =5 # One-hot size
hidden_size = 5 # output from the cell
batch_size=1 # one sentence
sequence_length=6
num_layers=1 # one layer run
class Model(torch.nn.Module):
def __init__(self, num_classes, input_size, hidden_size, num_layers, sequence_length):
super(Model, self).__init__()
self.num_classes = num_classes
self.input_size = input_size
self.hidden_size = hidden_size
self.num_layers = num_layers
self.sequence_length = sequence_length
self.rnn = torch.nn.RNN(input_size=5, hidden_size=5, batch_first=True)
def forward(self,x):
h_0 = torch.zeros(self.num_layers, x.size(0), self.hidden_size)
x = x.view(x.size(0), self.sequence_length, self.input_size)
outputs, hidden = self.rnn(x,h_0)
return outputs.view(-1, num_classes)
model = Model(num_classes, input_size, hidden_size, num_layers ,sequence_length)
print(model)
# Set loss and optimizer
criterion = torch.nn.CrossEntropyLoss()
optimus = torch.optim.SGD(model.parameters(), lr=0.1)
for epoch in range(100):
outputs = model(inputs)
optimus.zero_grad()
loss = criterion(outputs, labels)
loss.backward()
optimus.step()
_, idx = outputs.max(1)
idx = idx.data.numpy()
result_str = [idx2char[c] for c in idx.squeeze()]
if epoch%10 == 0:
print("epoch: %d, loss: %1.3f" % (epoch + 1, loss.item()))
print("Predicted string: ", ''.join(result_str))
print("Learning finished!")
```
### Experiment 3:
* We will be doing the same experiment as above but rather than using one hot embedding we will be using `embedding layer`.
```
# Creating a project to convert hihell -> ihello
# Data prepration
x_data = [[0,1,0,2,3,3]] #hihell
y_data = [1,0,2,3,3,4] #ihello
inputs = torch.LongTensor(x_data)
labels = torch.LongTensor(y_data)
labels.size(0)
# Parameters
embedding_size = 10
num_classes =5
input_size =5 # One-hot size
hidden_size = 5 # output from the cell
batch_size=1 # one sentence
sequence_length=6
num_layers=1 # one layer run
class Model(torch.nn.Module):
def __init__(self):
super(Model, self).__init__()
# self.num_classes = num_classes
# self.input_size = input_size
# self.hidden_size = hidden_size
# self.num_layers = num_layers
# self.sequence_length = sequence_length
self.embedding = torch.nn.Embedding(input_size, embedding_size)
self.rnn = torch.nn.RNN(input_size=embedding_size, hidden_size=5, batch_first=True)
self.fc = torch.nn.Linear(hidden_size, num_classes)
def forward(self,x):
h_0 = torch.zeros(num_layers, x.size(0),hidden_size)
emb = self.embedding(x)
emb = emb.view(batch_size, sequence_length, -1)
outputs, hidden = self.rnn(emb,h_0)
return self.fc(outputs.view(-1, num_classes))
model = Model()
print(model)
# Set loss and optimizer
criterion = torch.nn.CrossEntropyLoss()
optimus = torch.optim.Adam(model.parameters(), lr=0.1)
for epoch in range(100):
outputs = model(inputs)
optimus.zero_grad()
loss = criterion(outputs, labels)
loss.backward()
optimus.step()
_, idx = outputs.max(1)
idx = idx.data.numpy()
result_str = [idx2char[c] for c in idx.squeeze()]
if epoch%5 == 0:
print("epoch: %d, loss: %1.3f" % (epoch + 1, loss.item()))
print("Predicted string: ", ''.join(result_str))
print("Learning finished!")
```
| github_jupyter |
<a href="https://colab.research.google.com/github/SoIllEconomist/ds4b/blob/master/python_ds4b/python_basics/course_restart/Conditionals%20and%20Recursion.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Conditionals and Recursion
- `if` statement
- executes code depending on the state of the program
## Floor division and modulus
The **floor division** operator, //, divides two numbers and rounds down to an integer.
Suppose you have a movie with a runtime of 105 minutes. You might want to know how long that is in hours.
Conventional division returns a floating-point
```
minutes = 105
minutes / 60
```
But we don’t normally write hours with decimal points. Floor division returns the integer number of hours, rounding down:
```
hours = minutes // 60
hours
```
**Modulus operator**, %, which divides two numbers and returns the remainder.
```
remainder = minutes % 60
remainder
def movie_time(minutes):
"""Converts movie time from minutes
to hours minutes.
Output:
=========
X hrs Y mins
"""
hrs = minutes // 60
mins = minutes % 60
print(str(hrs) + ' hrs ' + str(mins) +' mins')
movie_time(105)
```
## Boolean expressions
A boolean expression is an expression that is either true or false. The following examples use the operator `==`, which compares two operands and produces `True` if they are equal and False otherwise:
```
5 == 5
5 == 6
type(True)
type(False)
```
The `==` operator is one of the relational operators; the others are:
| Relational Operator | Description |
|---------------------|---------------------------------|
| x != y | x is not equal to y |
| x > y | x is greater than y |
| x < y | x is less than y |
| x >= y | x is greater than or equal to y |
| x <= y | x is less than or equal to y |
## Logical operators
There are three **logical operators**:
- and, or, and not.
The meaning of these operators is similar to their meaning in English.
For example, `x > 0` **and** `x < 10` is true only if x is greater than 0 and less than 10.
```
5 > 0 and 5 < 10
20 > 0 and 20 < 10
```
`n%2 == 0` **or** `n%3 == 0` is true if either or both of the conditions is true, that is, if the number is divisible by 2 or 3
```
4 % 2 == 0 or 4 % 3 == 0
9 % 2 == 0 or 9 % 3 == 0
9 % 2 == 0 or 10 % 3 == 0
```
the not operator negates a boolean expression, so **not** `(x > y)` is true if x > y is false, that is, if x is less than or equal to y.
```
not (10 > 5)
(10 > 5)
```
## Conditional execution
In order to write useful programs, we almost always need the ability to check conditions and change the behavior of the program accordingly.
**Conditional statements** give us this ability. The simplest form is the if statement:
```
x = 10
if x > 0:
print('x is positive')
```
The boolean expression after if is called the **condition**. If it is true, the indented statement runs. If not, nothing happens.
`if` statements have the same structure as function definitions: a header followed by an indented body.
Statements like this are called **compound statements**.
## Alternative execution
A second form of the if statement is “alternative execution”, in which there are two possibilities and the condition determines which one runs.
The alternatives are called **branches**, because they are branches in the flow of execution.
```
x = 10
if x % 2 ==0:
print('x is even')
else:
print('x is odd')
x = 9
if x % 2 ==0:
print('x is even')
else:
print('x is odd')
```
## Chained conditionals
Sometimes there are more than two possibilities and we need more than two branches.
`elif` is an abbreviation of “else if”.
Exactly one branch will run.
There is no limit on the number of elif statements.
If there is an else clause, it has to be at the end, but there doesn’t have to be one.
One way to express a computation like that is a **chained conditional**:
```
x = 10
y = 11
if x < y:
print('x is less than y')
elif x > y:
print('x is greater than y')
else:
print('x and y are equal')
x = 12
y = 11
if x < y:
print('x is less than y')
elif x > y:
print('x is greater than y')
else:
print('x and y are equal')
x = 12
y = 12
if x < y:
print('x is less than y')
elif x > y:
print('x is greater than y')
else:
print('x and y are equal')
```
## Nested conditionals
- One conditional can also be nested within another.
- The outer conditional contains two branches.
- The first branch contains a simple statement.
- The second branch contains another if statement, which has two branches of its own.
- Those two branches are both simple statements, although they could have been conditional statements as well.
```
x = 12
y = 12
if x == y:
print('x and y are equal')
else:
if x < y:
print('x is less than y')
else:
print('x is greater than y')
x = 12
y = 11
if x == y:
print('x and y are equal')
else:
if x < y:
print('x is less than y')
else:
print('x is greater than y')
x = 10
y = 11
if x == y:
print('x and y are equal')
else:
if x < y:
print('x is less than y')
else:
print('x is greater than y')
```
Although the indentation of the statements makes the structure apparent, nested conditionals become difficult to read very quickly.
**It is a good idea to avoid them when you can.**
## Recursion
It is legal for one function to call another; it is also legal for a function to call itself.
It may not be obvious why that is a good thing, but it turns out to be one of the most magical things a program can do.
### What is recursion in Python?
Recursion is the process of defining something in terms of itself.
A physical world example would be to place two parallel mirrors facing each other. Any object in between them would be reflected recursively.
```
def countdown(n):
if n <= 0:
print('Blastoff!')
else:
print(n)
countdown(n-1)
```
- If n is 0 or negative, it outputs the word, “Blastoff!”
- Otherwise, it outputs n and then calls a function named countdown—itself—passing n-1 as an argument.
```
countdown(5)
```
## Infinite recursion
- **Infinite recursion** is when a recursion never reaches a base case, it goes on making recursive calls forever, and the program never terminates.
In most programming environments, a program with infinite recursion does not really run forever. Python reports an error message when the maximum recursion depth is reached:
```
def recursion():
recursion()
recursion()
```
## Keyboard Input
- Python provides a built-in function called `input` that stops the program and waits for the user to type something.
- When the user presses Return or Enter, the program resumes and input returns what the user typed as a string.
```
text = input()
text
number = input('Pick a number between 1 and 3:\n')
number
int(number)
```
What if they typed out the digits instead?
```
number = input('Pick a number between 1 and 3:\n')
int(number)
```
| github_jupyter |
### Distribution of Named Entities in Regular and Non-Regular Articles
In this notebook, we will look at the distribution of different named entities in regular and non-regular articles. We would primarily be looking at the number of mentions of that entity by the document length and draw a distribution for each entity for both the regular and non-regular articles. Below are the common named entities that are referenced:
**Different Types of Named Entities**
`PERSON`: People, including fictional.
`NORP`: Nationalities or religious or political groups.
`ORG`: Companies, agencies, institutions, etc.
`LOCATION`: mountain ranges, bodies of water, counter, cities, states, buildings, airports, highways, bridges, etc.
`PRODUCT`: Objects, vehicles, foods, etc. (Not services.)
`EVENT`: Named hurricanes, battles, wars, sports events, etc.
`WORK_OF_ART`: Titles of books, songs, etc.
`LAW`: Named documents made into laws.
`LANGUAGE`: Any named language.
`DATE`: Absolute or relative dates or periods.
`TIME`: Times smaller than a day.
`PERCENT`: Percentage, including ”%“.
`MONEY`: Monetary values, including unit.
`QUANTITY`: Measurements, as of weight or distance.
`ORDINAL`: “first”, “second”, etc.
`CARDINAL`: Numerals that do not fall under another type.
```
#Importing Libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import spacy
from tqdm import tqdm
#Reading the data
df = pd.read_csv('combined_data.csv')
df['label'] = df['label'].apply(lambda x: 'regular' if x == 'regular' or x == '0' else 'other')
df.head()
#Creating Sample
df_other = df[df.label=='other'].reset_index(drop=True)
df_regular = df[df.label=='regular'].sample(n=len(df[df.label=='other']), replace=False, random_state=0).reset_index(drop=True)
df_sample = df_other.append(df_regular).reset_index(drop=True)
df_sample = df_sample.sample(frac=1)
#Collecting Entities
regular_entities_all = {'PERSON': [], 'NORP': [], 'FAC': [], 'ORG': [], 'GPE': [], 'LOC': [],
'PRODUCT': [], 'EVENT': [], 'WORK_OF_ART': [], 'LAW': [], 'LANGUAGE': [],
'DATE': [], 'LANGUAGE': [], 'DATE': [], 'TIME': [], 'PERCENT': [], 'MONEY':[],
'QUANTITY': [], 'ORDINAL': [], 'CARDINAL': []}
op_entities_all = {'PERSON': [], 'NORP': [], 'FAC': [], 'ORG': [], 'GPE': [], 'LOC': [],
'PRODUCT': [], 'EVENT': [], 'WORK_OF_ART': [], 'LAW': [], 'LANGUAGE': [],
'DATE': [], 'LANGUAGE': [], 'DATE': [], 'TIME': [], 'PERCENT': [], 'MONEY':[],
'QUANTITY': [], 'ORDINAL': [], 'CARDINAL': []}
for i, row in tqdm(df_sample.iterrows()):
nlp = spacy.load('en_core_web_sm')
article = row['article_text']
try:
doc = nlp(article)
regular_entities = {}
op_entities = {}
for entity in doc.ents:
if row['label'] == 'regular':
if entity.label_ not in regular_entities.keys():
regular_entities[entity.label_] = 1
else:
regular_entities[entity.label_] +=1
else:
if entity.label_ not in op_entities.keys():
op_entities[entity.label_] = 1
else:
op_entities[entity.label_] +=1
for i in op_entities.keys():
op_entities_all[i].append(op_entities[i]/len(doc))
for i in regular_entities.keys():
regular_entities_all[i].append(regular_entities[i]/len(doc))
except:
print(i)
continue
#Preparing DataFrame from the constructed dictionary
op_ent_df = pd.DataFrame(dict([ (k,pd.Series(v)) for k,v in op_entities_all.items() ]))
op_ent_df['LOCATION'] = op_ent_df['FAC'] + op_ent_df['GPE'] + op_ent_df['LOC']
op_ent_df = op_ent_df.drop(columns=['FAC', 'GPE', 'LOC'])
reg_ent_df = pd.DataFrame(dict([ (k,pd.Series(v)) for k,v in regular_entities_all.items() ]))
reg_ent_df['LOCATION'] = reg_ent_df['FAC'] + reg_ent_df['GPE'] + reg_ent_df['LOC']
reg_ent_df = reg_ent_df.drop(columns=['FAC', 'GPE', 'LOC'])
#Preparing the charts
fig, ax = plt.subplots(4,4, figsize=(18,12))
fig.suptitle('Distribution of Named Entities', fontweight='bold')
plt.subplots_adjust(hspace=0.5)
plt.style.use('ggplot')
entities = reg_ent_df.columns
for i in range(4):
for j in range(4):
bins = np.histogram(np.hstack((op_ent_df[entities[0]],
reg_ent_df[entities[0]]))[~np.isnan(np.hstack((op_ent_df[entities[0]],
reg_ent_df[entities[0]])))],
bins=20)[1]
_ = ax[i, j].hist(op_ent_df[entities[0]], alpha=0.5, bins = bins, label='Non-Regular')
_ = ax[i, j].hist(reg_ent_df[entities[0]], alpha=0.5, bins = bins, label='Regular')
_ = ax[i, j].set_title(entities[0], fontweight='bold')
_ = ax[i, j].legend()
entities = entities[1:]
_ = ax[3, 0].set_xlabel('Instances by article length', fontweight='bold')
_ = ax[3, 1].set_xlabel('Instances by article length', fontweight='bold')
_ = ax[3, 2].set_xlabel('Instances by article length', fontweight='bold')
_ = ax[3, 3].set_xlabel('Instances by article length', fontweight='bold')
_ = ax[0, 0].set_ylabel('Articles', fontweight='bold')
_ = ax[1, 0].set_ylabel('Articles', fontweight='bold')
_ = ax[2, 0].set_ylabel('Articles', fontweight='bold')
_ = ax[3, 0].set_ylabel('Articles', fontweight='bold')
```
**Observations**
As we can observe from the above charts, there is some striking differences in the distribution of named entities in the two classes:
* **Non-Regular Articles** have many more mentions of person names, languages and nationalities as compared to regular articles.
* **Regular Articles** on the other hand have much more mentions of locations, quantity, time and date.
* The distribution is heavily right-skewed for most of the entity types
| github_jupyter |
## **Waste classification**
The project concerns waste classification to determine if it may be recycle or not. The dataset contains waste images recyclable and organic ones. We will try to train a model that will be able to recognise them all.
It is a binary classification problem and we will train the model in TensorFlow.
**Data**
The dataset contains images divided into two classes: organic waste and recycle waste and it is split into train data (85%) and test data (15%).
Training dataset contains 22564 images while test dataset 2513 images.
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import PIL
import keras
import tensorflow as tf
from tensorflow.keras import layers
from tensorflow.keras.models import Sequential
from keras.layers import Conv2D, MaxPool2D, Activation
from keras.layers import Dropout, Dense, Flatten, BatchNormalization
cd '/content/drive/My Drive/Kaggle'
```
### **Data preparation**
```
train_dir = '/content/drive/My Drive/Kaggle/DATASET/TRAIN'
test_dir = '/content/drive/My Drive/Kaggle/DATASET/TEST'
```
We load images with **image_dataset_from_directory** - this method returns an object **tf.data.Dataset**.
We define some of parameters:
```
batch_size = 32
img_height = 64
img_width = 64
```
Create a dataset:
```
train_ds = tf.keras.preprocessing.image_dataset_from_directory(
train_dir,
validation_split=0.2,
subset='training',
seed=123,
image_size=(img_height, img_width),
batch_size=batch_size)
test_ds = tf.keras.preprocessing.image_dataset_from_directory(
test_dir,
validation_split=0.2,
subset="validation",
seed=123,
image_size=(img_height, img_width),
batch_size=batch_size)
```
The class names can be found in the attribute class_names. They correspond to directory names in alphabetical order.
```
class_names = train_ds.class_names
print(class_names)
```
**Data visualization**
Here there are first 9 images from the training dataset.
```
plt.figure(figsize= (10,10))
for images, labels in train_ds.take(1):
for i in range(9):
ax = plt.subplot(3, 3, i + 1)
plt.imshow(images[i].numpy().astype('uint8'))
plt.title(class_names[labels[i]])
plt.axis('off')
```
Shape of data:
```
for image_batch, labels_batch in train_ds:
print(image_batch.shape)
print(labels_batch.shape)
break
```
**Data standardization**
The RGB channel values fall into the [0, 255] range. We have to standardize values to be in the [0, 1] range. We use a Rescaling layer for it.
```
normalization_layer = layers.experimental.preprocessing.Rescaling(1./255)
normalized_ds = train_ds.map(lambda x, y: (normalization_layer(x), y))
X_train, y_train = next(iter(normalized_ds))
first_image = X_train[0]
print(np.min(first_image), np.max(first_image))
normalized_test = test_ds.map(lambda x, y: (normalization_layer(x), y))
X_test, y_test = next(iter(normalized_test))
first_img = X_test[0]
print(np.min(first_img), np.max(first_img))
```
### **Creating the model**
We build model with Convolutional Neural Network (CNN) and we will test a few models.
The model consists of three convolution blocks with a max pool layer in each of them. There's a fully connected layer with three Dense layers. First two layers are activated by a relu activation function and the last is activated by a sigmoid activation function.
Compile the model:
We choose the Adam optimizer and the loss function: binary_crossentropy. To view training and validation accuracy for each training epoch, we use metrics "accuracy".
```
model = Sequential([
tf.keras.layers.Conv2D(32, (3,3), activation='relu', padding='same', input_shape=(64, 64, 3)),
tf.keras.layers.MaxPool2D((2,2)),
tf.keras.layers.Conv2D(32, (3,3), activation='relu', padding='same'),
tf.keras.layers.MaxPool2D((2,2)),
tf.keras.layers.Conv2D(64, (3,3), activation='relu', padding='same'),
tf.keras.layers.MaxPool2D((2,2)),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(units=128, activation="relu"),
tf.keras.layers.Dense(units=64, activation="relu"),
tf.keras.layers.Dense(units=1, activation="sigmoid")
])
model.compile(optimizer = 'adam', loss = 'binary_crossentropy',
metrics = ['accuracy'])
model.summary()
```
**Train the model**
We train the model for 30 epoch.
```
history = model.fit(
X_train, y_train,
epochs = 30,
validation_data=(X_test, y_test),
verbose=1
)
loss, accuracy = model.evaluate(X_train,y_train, verbose=False)
print("Training Accuracy: {:.4f}".format(accuracy))
loss, accuracy = model.evaluate(X_test,y_test, verbose=False)
print("Testing Accuracy: {:.4f}".format(accuracy))
```
**Visualization training results**
The plots of loss and accuracy on the training and validation sets.
```
def plot_graphs(history, string):
plt.plot(history.history[string])
plt.plot(history.history['val_'+string])
plt.xlabel("Epochs")
plt.ylabel(string)
plt.legend([string, 'val_'+string])
plt.show()
plot_graphs(history, "accuracy")
plot_graphs(history, "loss")
```
From the above plots one can see that difference in accuracy between training and validation set is noticeable. We are dealing with overfitting. The training accuracy is increasing over time, whereas validation accuracy achieved 78% in the training process.
In this case from this dataset is difficult to achieve good accuracy with a CNN created from scratch.
**Model 2**
In model two we add additional convolution layers and we reduce the complexity of architecture in fully connected layer to have a better accuracy. We also are adding Dropout layer to fight with overfitting in the training process.
```
model2 = Sequential([
tf.keras.layers.Conv2D(32, (3,3), input_shape=(64, 64, 3), activation='relu', padding='same'),
tf.keras.layers.MaxPool2D((2,2)),
tf.keras.layers.Conv2D(32, (3,3), activation='relu', padding='same'),
tf.keras.layers.MaxPool2D((2,2)),
tf.keras.layers.Conv2D(64, (3,3), activation='relu', padding='same'),
tf.keras.layers.MaxPool2D((2,2)),
tf.keras.layers.Conv2D(128, (3,3), activation='relu', padding='same'),
tf.keras.layers.MaxPool2D((2,2)),
tf.keras.layers.Flatten(),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(64, activation="relu"),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(32, activation="relu"),
tf.keras.layers.Dense(1, activation="sigmoid")
])
model2.compile(optimizer = 'adam', loss = 'binary_crossentropy',
metrics = ['accuracy'])
model2.summary()
history_2 = model2.fit(
X_train, y_train,
epochs = 30,
validation_data=(X_test, y_test),
verbose=1
)
loss, accuracy = model2.evaluate(X_train,y_train, verbose=False)
print("Training Accuracy: {:.4f}".format(accuracy))
loss, accuracy = model2.evaluate(X_test,y_test, verbose=False)
print("Testing Accuracy: {:.4f}".format(accuracy))
plot_graphs(history_2, "accuracy")
plot_graphs(history_2, "loss")
```
The validation accuracy is achieved 81% in the training process.
The training accuracy is increasing over time, whereas validation accuracy stalls around 80% in the training process. We also see that the overfitting slightly decreased. The validation loss reaches a minimum value after 5 epochs, and then stabilizes, with the training loss decreasing linearly until the values are close to 0.
**Model 3**
We also may add Batch Normalization to reduce overfitting.
```
model3 = Sequential([
tf.keras.layers.Conv2D(32, (3,3), input_shape=(64, 64, 3), activation='relu', padding='same'),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.MaxPool2D((2,2)),
tf.keras.layers.Conv2D(32, (3,3), activation='relu', padding='same'),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.MaxPool2D((2,2)),
tf.keras.layers.Conv2D(64, (3,3), activation='relu', padding='same'),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.MaxPool2D((2,2)),
tf.keras.layers.Conv2D(128, (3,3), activation='relu', padding='same'),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.MaxPool2D((2,2)),
tf.keras.layers.Flatten(),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(64, activation="relu"),
tf.keras.layers.Dense(32, activation="relu"),
tf.keras.layers.Dense(1, activation="sigmoid")
])
model3.compile(optimizer='adam', loss = 'binary_crossentropy',
metrics = ['accuracy'])
model3.summary()
```
In this case we train the model for 40 epoch.
```
history_3 = model3.fit(
X_train, y_train,
epochs = 40,
validation_data=(X_test, y_test),
verbose=1
)
loss, accuracy = model3.evaluate(X_test,y_test, verbose=False)
print("Testing Accuracy: {:.4f}".format(accuracy))
plot_graphs(history_3, "accuracy")
plot_graphs(history_3, "loss")
```
In this case we achaived the worst validation accuracy equal to 65% in the training process. We see that the training accuracy is increasing over time, whereas validation accuracy stalls around 65% in the training process. Unfortunately the overfitting has grown significantly.
```
```
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/Image/image_stats_by_band.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/Image/image_stats_by_band.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://mybinder.org/v2/gh/giswqs/earthengine-py-notebooks/master?filepath=Image/image_stats_by_band.ipynb"><img width=58px src="https://mybinder.org/static/images/logo_social.png" />Run in binder</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/Image/image_stats_by_band.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API and geemap
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geemap](https://github.com/giswqs/geemap). The **geemap** Python package is built upon the [ipyleaflet](https://github.com/jupyter-widgets/ipyleaflet) and [folium](https://github.com/python-visualization/folium) packages and implements several methods for interacting with Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, and `Map.centerObject()`.
The following script checks if the geemap package has been installed. If not, it will install geemap, which automatically installs its [dependencies](https://github.com/giswqs/geemap#dependencies), including earthengine-api, folium, and ipyleaflet.
**Important note**: A key difference between folium and ipyleaflet is that ipyleaflet is built upon ipywidgets and allows bidirectional communication between the front-end and the backend enabling the use of the map to capture user input, while folium is meant for displaying static data only ([source](https://blog.jupyter.org/interactive-gis-in-jupyter-with-ipyleaflet-52f9657fa7a)). Note that [Google Colab](https://colab.research.google.com/) currently does not support ipyleaflet ([source](https://github.com/googlecolab/colabtools/issues/60#issuecomment-596225619)). Therefore, if you are using geemap with Google Colab, you should use [`import geemap.eefolium`](https://github.com/giswqs/geemap/blob/master/geemap/eefolium.py). If you are using geemap with [binder](https://mybinder.org/) or a local Jupyter notebook server, you can use [`import geemap`](https://github.com/giswqs/geemap/blob/master/geemap/geemap.py), which provides more functionalities for capturing user input (e.g., mouse-clicking and moving).
```
# Installs geemap package
import subprocess
try:
import geemap
except ImportError:
print('geemap package not installed. Installing ...')
subprocess.check_call(["python", '-m', 'pip', 'install', 'geemap'])
# Checks whether this notebook is running on Google Colab
try:
import google.colab
import geemap.eefolium as emap
except:
import geemap as emap
# Authenticates and initializes Earth Engine
import ee
try:
ee.Initialize()
except Exception as e:
ee.Authenticate()
ee.Initialize()
```
## Create an interactive map
The default basemap is `Google Satellite`. [Additional basemaps](https://github.com/giswqs/geemap/blob/master/geemap/geemap.py#L13) can be added using the `Map.add_basemap()` function.
```
Map = emap.Map(center=[40,-100], zoom=4)
Map.add_basemap('ROADMAP') # Add Google Map
Map
```
## Add Earth Engine Python script
```
# Add Earth Engine dataset
image = ee.Image('USDA/NAIP/DOQQ/m_3712213_sw_10_1_20140613')
Map.setCenter(-122.466123, 37.769833, 17)
Map.addLayer(image, {'bands': ['N', 'R','G']}, 'NAIP')
geometry = image.geometry()
means = image.reduceRegions(geometry, ee.Reducer.mean().forEachBand(image), 10)
print(means.getInfo())
```
## Display Earth Engine data layers
```
Map.addLayerControl() # This line is not needed for ipyleaflet-based Map.
Map
```
| github_jupyter |
```
import keras
keras.__version__
from keras import backend as K
K.clear_session()
```
# Generating images
This notebook contains the second code sample found in Chapter 8, Section 4 of [Deep Learning with Python](https://www.manning.com/books/deep-learning-with-python?a_aid=keras&a_bid=76564dff). Note that the original text features far more content, in particular further explanations and figures: in this notebook, you will only find source code and related comments.
---
## Variational autoencoders
Variational autoencoders, simultaneously discovered by Kingma & Welling in December 2013, and Rezende, Mohamed & Wierstra in January 2014,
are a kind of generative model that is especially appropriate for the task of image editing via concept vectors. They are a modern take on
autoencoders -- a type of network that aims to "encode" an input to a low-dimensional latent space then "decode" it back -- that mixes ideas
from deep learning with Bayesian inference.
A classical image autoencoder takes an image, maps it to a latent vector space via an "encoder" module, then decode it back to an output
with the same dimensions as the original image, via a "decoder" module. It is then trained by using as target data the _same images_ as the
input images, meaning that the autoencoder learns to reconstruct the original inputs. By imposing various constraints on the "code", i.e.
the output of the encoder, one can get the autoencoder to learn more or less interesting latent representations of the data. Most
commonly, one would constraint the code to be very low-dimensional and sparse (i.e. mostly zeros), in which case the encoder acts as a way
to compress the input data into fewer bits of information.
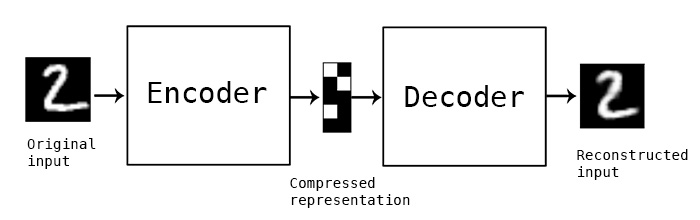
In practice, such classical autoencoders don't lead to particularly useful or well-structured latent spaces. They're not particularly good
at compression, either. For these reasons, they have largely fallen out of fashion over the past years. Variational autoencoders, however,
augment autoencoders with a little bit of statistical magic that forces them to learn continuous, highly structured latent spaces. They
have turned out to be a very powerful tool for image generation.
A VAE, instead of compressing its input image into a fixed "code" in the latent space, turns the image into the parameters of a statistical
distribution: a mean and a variance. Essentially, this means that we are assuming that the input image has been generated by a statistical
process, and that the randomness of this process should be taken into accounting during encoding and decoding. The VAE then uses the mean
and variance parameters to randomly sample one element of the distribution, and decodes that element back to the original input. The
stochasticity of this process improves robustness and forces the latent space to encode meaningful representations everywhere, i.e. every
point sampled in the latent will be decoded to a valid output.
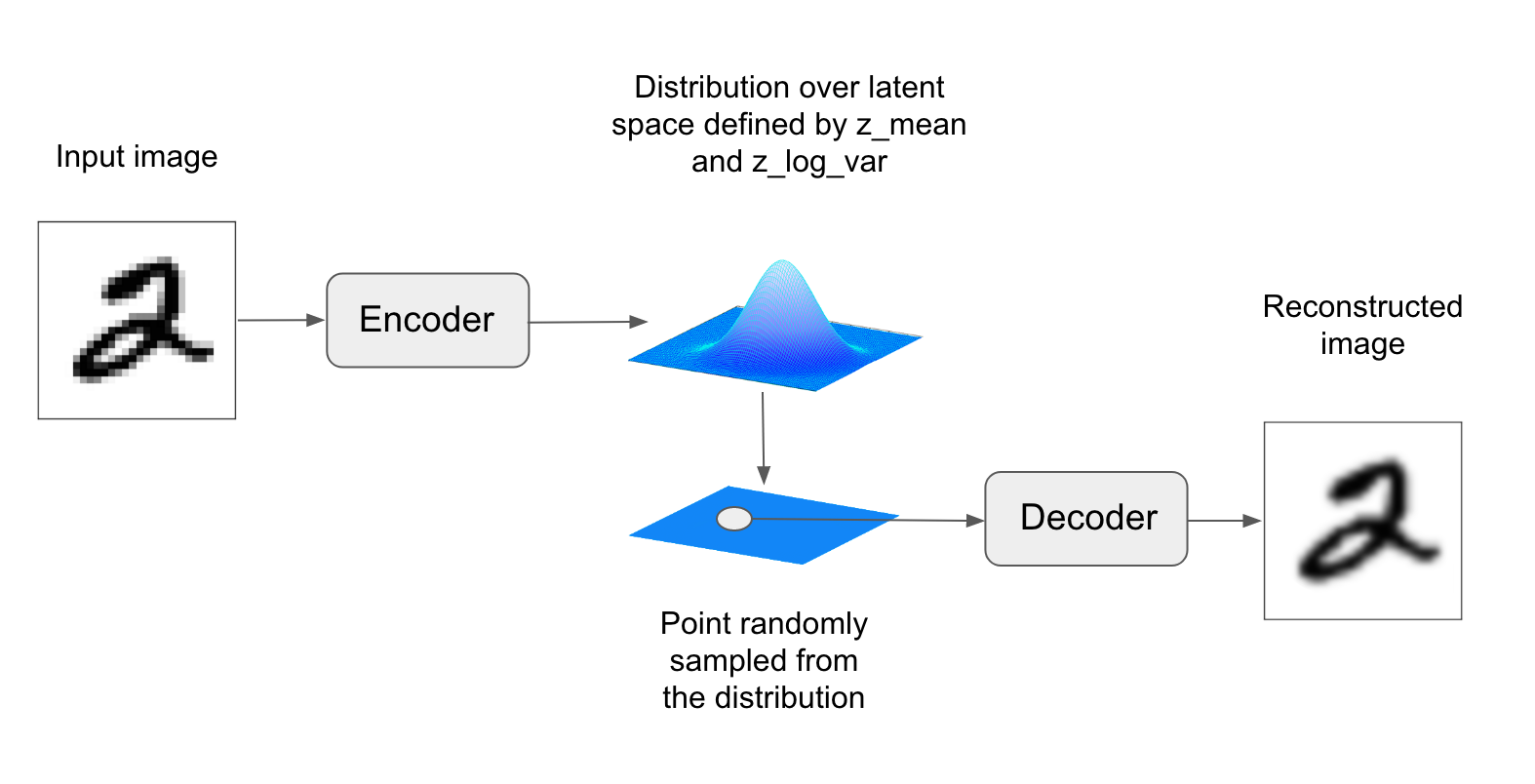
In technical terms, here is how a variational autoencoder works. First, an encoder module turns the input samples `input_img` into two
parameters in a latent space of representations, which we will note `z_mean` and `z_log_variance`. Then, we randomly sample a point `z`
from the latent normal distribution that is assumed to generate the input image, via `z = z_mean + exp(z_log_variance) * epsilon`, where
epsilon is a random tensor of small values. Finally, a decoder module will map this point in the latent space back to the original input
image. Because `epsilon` is random, the process ensures that every point that is close to the latent location where we encoded `input_img`
(`z-mean`) can be decoded to something similar to `input_img`, thus forcing the latent space to be continuously meaningful. Any two close
points in the latent space will decode to highly similar images. Continuity, combined with the low dimensionality of the latent space,
forces every direction in the latent space to encode a meaningful axis of variation of the data, making the latent space very structured
and thus highly suitable to manipulation via concept vectors.
The parameters of a VAE are trained via two loss functions: first, a reconstruction loss that forces the decoded samples to match the
initial inputs, and a regularization loss, which helps in learning well-formed latent spaces and reducing overfitting to the training data.
Let's quickly go over a Keras implementation of a VAE. Schematically, it looks like this:
```
# Encode the input into a mean and variance parameter
z_mean, z_log_variance = encoder(input_img)
# Draw a latent point using a small random epsilon
z = z_mean + exp(z_log_variance) * epsilon
# Then decode z back to an image
reconstructed_img = decoder(z)
# Instantiate a model
model = Model(input_img, reconstructed_img)
# Then train the model using 2 losses:
# a reconstruction loss and a regularization loss
```
Here is the encoder network we will use: a very simple convnet which maps the input image `x` to two vectors, `z_mean` and `z_log_variance`.
```
import keras
from keras import layers
from keras import backend as K
from keras.models import Model
import numpy as np
img_shape = (28, 28, 1)
batch_size = 16
latent_dim = 2 # Dimensionality of the latent space: a plane
input_img = keras.Input(shape=img_shape)
x = layers.Conv2D(32, 3,
padding='same', activation='relu')(input_img)
x = layers.Conv2D(64, 3,
padding='same', activation='relu',
strides=(2, 2))(x)
x = layers.Conv2D(64, 3,
padding='same', activation='relu')(x)
x = layers.Conv2D(64, 3,
padding='same', activation='relu')(x)
shape_before_flattening = K.int_shape(x)
x = layers.Flatten()(x)
x = layers.Dense(32, activation='relu')(x)
z_mean = layers.Dense(latent_dim)(x)
z_log_var = layers.Dense(latent_dim)(x)
```
Here is the code for using `z_mean` and `z_log_var`, the parameters of the statistical distribution assumed to have produced `img_img`, to
generate a latent space point `z`. Here, we wrap some arbitrary code (built on top of Keras backend primitives) into a `Lambda` layer. In
Keras, everything needs to be a layer, so code that isn't part of a built-in layer should be wrapped in a `Lambda` (or else, in a custom
layer).
```
def sampling(args):
z_mean, z_log_var = args
epsilon = K.random_normal(shape=(K.shape(z_mean)[0], latent_dim),
mean=0., stddev=1.)
return z_mean + K.exp(z_log_var) * epsilon
z = layers.Lambda(sampling)([z_mean, z_log_var])
```
This is the decoder implementation: we reshape the vector `z` to the dimensions of an image, then we use a few convolution layers to obtain a final
image output that has the same dimensions as the original `input_img`.
```
# This is the input where we will feed `z`.
decoder_input = layers.Input(K.int_shape(z)[1:])
# Upsample to the correct number of units
x = layers.Dense(np.prod(shape_before_flattening[1:]),
activation='relu')(decoder_input)
# Reshape into an image of the same shape as before our last `Flatten` layer
x = layers.Reshape(shape_before_flattening[1:])(x)
# We then apply then reverse operation to the initial
# stack of convolution layers: a `Conv2DTranspose` layers
# with corresponding parameters.
x = layers.Conv2DTranspose(32, 3,
padding='same', activation='relu',
strides=(2, 2))(x)
x = layers.Conv2D(1, 3,
padding='same', activation='sigmoid')(x)
# We end up with a feature map of the same size as the original input.
# This is our decoder model.
decoder = Model(decoder_input, x)
# We then apply it to `z` to recover the decoded `z`.
z_decoded = decoder(z)
```
The dual loss of a VAE doesn't fit the traditional expectation of a sample-wise function of the form `loss(input, target)`. Thus, we set up
the loss by writing a custom layer with internally leverages the built-in `add_loss` layer method to create an arbitrary loss.
```
class CustomVariationalLayer(keras.layers.Layer):
def vae_loss(self, x, z_decoded):
x = K.flatten(x)
z_decoded = K.flatten(z_decoded)
xent_loss = keras.metrics.binary_crossentropy(x, z_decoded)
kl_loss = -5e-4 * K.mean(
1 + z_log_var - K.square(z_mean) - K.exp(z_log_var), axis=-1)
return K.mean(xent_loss + kl_loss)
def call(self, inputs):
x = inputs[0]
z_decoded = inputs[1]
loss = self.vae_loss(x, z_decoded)
self.add_loss(loss, inputs=inputs)
# We don't use this output.
return x
# We call our custom layer on the input and the decoded output,
# to obtain the final model output.
y = CustomVariationalLayer()([input_img, z_decoded])
```
Finally, we instantiate and train the model. Since the loss has been taken care of in our custom layer, we don't specify an external loss
at compile time (`loss=None`), which in turns means that we won't pass target data during training (as you can see we only pass `x_train`
to the model in `fit`).
```
from keras.datasets import mnist
vae = Model(input_img, y)
vae.compile(optimizer='rmsprop', loss=None)
vae.summary()
# Train the VAE on MNIST digits
(x_train, _), (x_test, y_test) = mnist.load_data()
x_train = x_train.astype('float32') / 255.
x_train = x_train.reshape(x_train.shape + (1,))
x_test = x_test.astype('float32') / 255.
x_test = x_test.reshape(x_test.shape + (1,))
vae.fit(x=x_train, y=None,
shuffle=True,
epochs=10,
batch_size=batch_size,
validation_data=(x_test, None))
```
Once such a model is trained -- e.g. on MNIST, in our case -- we can use the `decoder` network to turn arbitrary latent space vectors into
images:
```
import matplotlib.pyplot as plt
from scipy.stats import norm
# Display a 2D manifold of the digits
n = 15 # figure with 15x15 digits
digit_size = 28
figure = np.zeros((digit_size * n, digit_size * n))
# Linearly spaced coordinates on the unit square were transformed
# through the inverse CDF (ppf) of the Gaussian
# to produce values of the latent variables z,
# since the prior of the latent space is Gaussian
grid_x = norm.ppf(np.linspace(0.05, 0.95, n))
grid_y = norm.ppf(np.linspace(0.05, 0.95, n))
for i, yi in enumerate(grid_x):
for j, xi in enumerate(grid_y):
z_sample = np.array([[xi, yi]])
z_sample = np.tile(z_sample, batch_size).reshape(batch_size, 2)
x_decoded = decoder.predict(z_sample, batch_size=batch_size)
digit = x_decoded[0].reshape(digit_size, digit_size)
figure[i * digit_size: (i + 1) * digit_size,
j * digit_size: (j + 1) * digit_size] = digit
plt.figure(figsize=(10, 10))
plt.imshow(figure, cmap='Greys_r')
plt.show()
```
The grid of sampled digits shows a completely continuous distribution of the different digit classes, with one digit morphing into another
as you follow a path through latent space. Specific directions in this space have a meaning, e.g. there is a direction for "four-ness",
"one-ness", etc.
| github_jupyter |
Create a TS+biology tracers initial file for one bathymetry based on a restart file from a different bathymetry
```
import netCDF4 as nc
import numpy as np
from salishsea_tools import nc_tools
%matplotlib inline
```
# New Bathymetry (via its mesh mask)
```
mesh = nc.Dataset('/home/sallen/MEOPAR/grid/mesh_mask201702.nc')
mbathy = mesh.variables['mbathy'][0,:,:]
#used to calculate number of vertical ocean grid cells at each (i,j) (1=land point)
gdepw = mesh.variables['gdepw_0'][0,:,:,:]
surface_tmask = mesh.variables['tmask'][0,0,:,:]
surface_tmask = np.abs(surface_tmask-1)
tmask = mesh.variables['tmask'][0,:,:,:]
tmask = np.abs(tmask-1)
lats = mesh.variables['nav_lat'][:]
lons = mesh.variables['nav_lon'][:]
mesh.close()
# calculate bathymetry based on meshmask
NEMO_bathy = np.zeros(mbathy.shape)
for i in range(NEMO_bathy.shape[1]):
for j in range(NEMO_bathy.shape[0]):
level = mbathy[j,i]
NEMO_bathy[j,i] = gdepw[level,j,i]
NEMO_bathy = np.ma.masked_array(NEMO_bathy, mask = surface_tmask)
```
# Old Bathymetry (based on its mesh mask)
```
oldmesh = nc.Dataset('/home/sallen/MEOPAR/NEMO-forcing/grid/mesh_mask201702.nc')
oldmbathy =oldmesh.variables['mbathy'][0,:,:]
#used to calculate number of vertical ocean grid cells at each (i,j) (1=land point)
oldgdepw = oldmesh.variables['gdepw_0'][0,:,:,:]
oldsurface_tmask = oldmesh.variables['tmask'][0,0,:,:]
oldsurface_tmask = np.abs(oldsurface_tmask-1)
oldtmask = oldmesh.variables['tmask'][0,:,:,:]
oldtmask = np.abs(oldtmask-1)
oldmesh.close()
```
# Restart Files to Get Tracers (TS and Biology)
```
dataphys = nc.Dataset('/results/SalishSea/nowcast-green/06mar17/SalishSea_02002320_restart.nc')
databio = nc.Dataset('/results/SalishSea/nowcast-green/06mar17/SalishSea_02002320_restart_trc.nc')
physical = ['tn', 'sn']
biological = ['TRNDON', 'TRNMICZ','TRNMYRI','TRNNH4','TRNNO3','TRNTRA',
'TRNPHY','TRNDIAT','TRNPON','TRNSi','TRNbSi']
varas = {}
for vb in physical:
varas[vb] = dataphys.variables[vb][0, :]
for vb in biological:
print (vb)
varas[vb] = databio.variables[vb][0, :]
dataphys.close()
databio.close()
varall = physical + biological
```
# Fill in any Missing Data Points
```
def find_mean(varas, varall, i, j, k, dd, oldtmask):
for vb in varall:
imin = max(i-dd, 0)
imax = min(i+dd, 897)
jmin = max(j-dd, 0)
jmax = min(j+dd, 397)
temporary = np.sum(varas[vb][k, imin:imax+1, jmin:jmax+1]*(1-oldtmask[k, imin:imax+1, jmin:jmax+1]))
count = np.sum(1-oldtmask[k, imin:imax+1, jmin:jmax+1])
if count == 0:
varas[vb][k, i, j] = 0
else:
varas[vb][k, i, j] = temporary/count
return varas
def fillit(kmax, oldtmask, varas, varall):
dd = 1
bad = 1
while bad > 0:
dd += 1
good = 1
while good > 0:
good = 0; bad = 0; already = 0
for k in range(kmax+1):
for i in range(1, 898):
for j in range(1, 398):
if tmask[k,i,j] < oldtmask[k,i,j]:
if varas['sn'][k, i, j] > 0:
already = already + 1
else:
varas = find_mean(varas, varall, i, j, k, dd, oldtmask)
if varas['sn'][k, i, j] > 0:
good = good + 1
else:
bad = bad + 1
print ('dd', dd, 'good', good)
print ('already', already, 'bad', bad)
```
This can take a very long time if the bathymetries are very different, aka add a new long river. If you want you can do it in pieces by starting with the first argument at say 5 and then slowly increasing it. You do need to go to 39 finally. For new bathy201702 versus old bathy201702 its fast.
```
fillit(39, oldtmask, varas, varall)
```
# Write your Initial File
```
# build nc file
new_initialfile = nc.Dataset('Bathy201702_06mar17.nc', 'w')
nc_tools.init_dataset_attrs(
new_initialfile,
title='All tracers for Bathymetry 201702 from nowcast-green 06mar17',
notebook_name='Tools/I_ForcingFiles/Initial/Initial_from_Restart_Bathy201702',
nc_filepath='sea_initial/v201702b/Bathy201702_06mar17.nc',
# nc_filepath='tracers/initial/Bathy201702_18aug17.nc',
comment='All Tracers, physical and biological')
new_initialfile.createDimension('y', 898)
new_initialfile.createDimension('x', 398)
new_initialfile.createDimension('deptht', 40)
new_initialfile.createDimension('time_counter', None)
thevara = {}
for vb in varall:
thevara[vb] = new_initialfile.createVariable(
vb, 'float32', ('time_counter', 'deptht', 'y', 'x'), zlib=True,
least_significant_digit=1e-5, fill_value=-99)
thevara[vb][0] = varas[vb]
print (np.max(thevara[vb]))
new_initialfile
new_initialfile.close()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/m-rafiul-islam/driver-behavior-model/blob/main/Finding_SSE.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
#!/usr/bin/env python3
# -*- coding: utf-8 -*-
"""
Created on Mon Jan 24 15:17:39 2022
@author: rafiul
"""
# import scipy.integrate as integrate
# from scipy.integrate import odeint
import sys
import os
# from geneticalgorithm import geneticalgorithm as ga
#from geneticalgorithm_pronto import geneticalgorithm as ga
# from ga import ga
import numpy as np
import scipy.integrate as integrate
from scipy import special
from scipy.interpolate import interp1d
import pandas as pd
# f = interp1d(nth_car_data['time'],nth_car_data['speed'])
###
version='007'
output_folder = 'results/version%s/' % version
isExist = os.path.exists(output_folder)
if not isExist:
os.makedirs(output_folder)
# python parameter_est_68.py $SLURM_ARRAY_TASK_ID 2 10 500
try:
filename = sys.argv[0]
SLURM_ID = int(sys.argv[1])
except:
filename = ''
SLURM_ID = 1
nth_car = SLURM_ID+1
if len(sys.argv)>2:
ga_iteration = int(sys.argv[2])
else:
ga_iteration = 3#5
def RK4(func, X0, ts):
"""
Runge Kutta 4 solver.
"""
dt = ts[1] - ts[0]
nt = len(ts)
X = np.zeros((nt, X0.shape[0]),dtype=np.float64)
X[0] = X0
for i in range(nt-1):
k1 = func(X[i], ts[i])
k2 = func(X[i] + dt/2. * k1, ts[i] + dt/2.)
k3 = func(X[i] + dt/2. * k2, ts[i] + dt/2.)
k4 = func(X[i] + dt * k3, ts[i] + dt)
X[i+1] = X[i] + dt / 6. * (k1 + 2. * k2 + 2. * k3 + k4)
return X
# see this link for model and paramterts https://en.wikipedia.org/wiki/Intelligent_driver_model
# DOI: 10.1098/rsta.2010.0084
# @jit(nopython=True)
def idm_model(x,t):
X,V = x[0],x[1]
dX,dV = np.zeros(1,dtype=np.float64), np.zeros(1,dtype=np.float64)
dX = V # Differtial Equation 1
###
s = position_LV(t) - X - 5 # 5 = length of the car
deltaV = V - speed_LV(t)
sstar = s0+V*T + (V*deltaV)/(2*np.sqrt(a*b))
# ###
dV = a*(1-(V/V_0)**delta - (sstar/s)**2) # Differtial Equation 2
return np.array([dX,dV],dtype=np.float64)
# @jit(nopython=True)
def speed_LV(t):
return interp1d(nth_car_data['time'],nth_car_data['speed'],bounds_error=False)(t)
def position_LV(t):
return interp1d(nth_car_data['time'],postion_of_the_LV,bounds_error=False)(t)
def fractional_idm_model_1d(V,t,X):
# index = round(t) #convert into integer number
current_position_of_follower = X
###
s = position_LV(t) - current_position_of_follower - 5 # 5 = length of the car
deltaV = V - speed_LV(t)
sstar = s0+V*T + (V*deltaV)/(2*np.sqrt(a*b))
# ###
dV = a*(1-(V/V_0)**delta - (sstar/s)**2) # Differtial Equation 2
return dV
def speed_error(sol,nth_car_speed):
return np.sum((sol[1,:-1]-nth_car_speed[1:])**2)
def gap_error(sol,postion_of_the_LV):
return np.sum((sol[0,:]-postion_of_the_LV)**2)
def caputoEuler_1d(a, f, y0, tspan, x0_f):
"""Use one-step Adams-Bashforth (Euler) method to integrate Caputo equation
D^a y(t) = f(y,t)
Args:
a: fractional exponent in the range (0,1)
f: callable(y,t) returning a numpy array of shape (d,)
Vector-valued function to define the right hand side of the system
y0: array of shape (d,) giving the initial state vector y(t==0)
tspan (array): The sequence of time points for which to solve for y.
These must be equally spaced, e.g. np.arange(0,10,0.005)
tspan[0] is the intial time corresponding to the initial state y0.
Returns:
y: array, with shape (len(tspan), len(y0))
With the initial value y0 in the first row
Raises:
FODEValueError
See also:
K. Diethelm et al. (2004) Detailed error analysis for a fractional Adams
method
C. Li and F. Zeng (2012) Finite Difference Methods for Fractional
Differential Equations
"""
#(d, a, f, y0, tspan) = _check_args(a, f, y0, tspan)
N = len(tspan)
h = (tspan[N-1] - tspan[0])/(N - 1)
c = special.rgamma(a) * np.power(h, a) / a
w = c * np.diff(np.power(np.arange(N), a))
fhistory = np.zeros(N - 1, dtype=np.float64)
y = np.zeros(N, dtype=np.float64)
x = np.zeros(N, dtype=np.float64)
y[0] = y0;
x[0] = x0_f;
for n in range(0, N - 1):
tn = tspan[n]
yn = y[n]
fhistory[n] = f(yn, tn, x[n])
y[n+1] = y0 + np.dot(w[0:n+1], fhistory[n::-1])
x[n+1] = x[n] + y[n+1] * h
return np.array([x,y])
def error_func_idm(variable_X):
# varbound=np.array([[a*lf,a*uf],[lf*delta,uf*delta],[lf*beta,uf*beta]])
a = variable_X[0]
delta = variable_X[1]
beta = variable_X[2]
x0 = np.array([initial_position,initial_velocity],dtype=np.float64) #initial position and velocity
# Classical ODE
# sol = integrate.odeint(idm_model, x0, time_span)
sol = RK4(idm_model, x0, time_span)
sol = sol.transpose(1,0)
# print(np.sum((sol[1,:-1]-nth_car_speed[1:])**2))
return np.sum((sol[1,1:]-nth_car_speed[:-1])**2)
def error_func_fidm(variable_X):
# varbound=np.array([[a*lf,a*uf],[lf*delta,uf*delta],[lf*beta,uf*beta]])
a = variable_X[0]
delta = variable_X[1]
beta = variable_X[2]
alpha = variable_X[3]
if alpha > .99999:
alpha = .99999
sol = caputoEuler_1d(alpha,fractional_idm_model_1d, initial_velocity, time_span, initial_position) #, args=(number_groups,beta_P,beta_C,beta_A,v,w,mu_E,mu_A,mu_P,mu_C,p,q,contact_by_group))
return np.sum((sol[1,1:]-nth_car_speed[:-1])**2)
######################################
# Global variables
# see this link for model and paramterts https://en.wikipedia.org/wiki/Intelligent_driver_model
V_0 = 20 # desired speed m/s
s0 = 30
T = 1.5
nth_car = 2
# a=1.5
# b = 1.67
# delta = 4.0
# beta = 2
# find best values for our model
# a_alpha = 1.2
# ######################################
# Actual data
# df = pd.read_csv('RAllCarDataTime350.csv')
git_raw_url = 'https://raw.githubusercontent.com/m-rafiul-islam/driver-behavior-model/main/RAllCarDataTime350.csv'
df = pd.read_csv(git_raw_url)
nth_car_data = df.loc[df['nthcar'] == nth_car, :]
nth_car_speed = np.array(df.loc[df['nthcar'] == nth_car,'speed'])
# leader vehicle profile
# 7 m/s - 25.2 km/h 11 m/s - 39.6 km/h 18 m/s - 64.8 km/h 22 m/s - 79.2 km/h
# 25 km/h -- 6.95 m/s 40 km/h -- 11.11 m/s 60 km/h -- 16.67 m/s
# dt=1 #time step -- 1 sec
time_span = np.array(nth_car_data['time'])
dt = time_span[1]-time_span[0]
# speed_of_the_LV = 15*np.ones(600+1) # we will need data
# speed_of_the_LV = np.concatenate((np.linspace(0,7,60),7*np.ones(120),np.linspace(7,11,60), 11*np.ones(120), np.linspace(11,0,60) ))# we will need data
speed_of_the_LV = nth_car_speed
num_points = len(speed_of_the_LV)
postion_of_the_LV = np.zeros(num_points)
initla_position_of_the_LV = 18.45 # 113
postion_of_the_LV[0] = initla_position_of_the_LV
for i in range(1,num_points):
postion_of_the_LV[i] = postion_of_the_LV[i-1] + dt*(speed_of_the_LV[i]+speed_of_the_LV[i-1])/2
# plt.figure()
# plt.subplot(211)
# plt.plot(speed_of_the_LV)
# plt.xlabel('time')
# plt.ylabel('speed of the leader vehicle')
# plt.subplot(212)
# plt.plot(postion_of_the_LV)
# plt.xlabel('time')
# plt.ylabel('postion of the leader vehicle')
# simulation_time = 35
# time_span = np.linspace(0, simulation_time, int(simulation_time /dt)+1)
#
# alpha_list = [.95,.9,.8]
initial_position = 0.
initial_velocity = 6.72
x0 = np.array([initial_position,initial_velocity],dtype=np.float64) #initial position and velocity
# f,ax=plt.subplots(2,1,figsize=(10,10))
# ax[0].plot(time_span,postion_of_the_LV,label = 'position of the LV')
# ax[1].plot(time_span,speed_of_the_LV,label = 'speed of the LV')
# # Classical ODE
# sol = integrate.odeint(idm_model, x0, time_span)
# ax[0].plot(time_span,sol[:,0], label='position of the FV using alpha = 1.00')
# ax[1].plot(time_span,sol[:,1], label='speed of the FV using alpha = 1.00')
# print(speed_error(sol,nth_car_speed))
# # Fractional ODE
# for alpha in alpha_list:
# #sol = fintegrate_mod.fodeint_mod(alpha,fractional_idm_model_mod, x0, ts) #, args=(number_groups,beta_P,beta_C,beta_A,v,w,mu_E,mu_A,mu_P,mu_C,p,q,contact_by_group))
# sol = caputoEuler_1d(alpha,fractional_idm_model_1d, initial_velocity, time_span, initial_position) #, args=(number_groups,beta_P,beta_C,beta_A,v,w,mu_E,mu_A,mu_P,mu_C,p,q,contact_by_group))
# ax[0].plot(time_span,sol[0], label='position of the FV using alpha = %.2f' %alpha)
# ax[1].plot(time_span,sol[1], label='speed of the FV using alpha = %.2f' %alpha)
# print(speed_error(sol,nth_car_speed))
# ax[0].set_xlabel('time (sec)')
# ax[0].set_ylabel('position (m)')
# ax[0].legend()
# ax[1].set_xlabel('time (sec)')
# ax[1].set_ylabel('speed (m/s)')
# ax[1].legend()
# plt.savefig('simulation.pdf',dpi=300)
# https://pypi.org/project/geneticalgorithm/
# try:
# from geneticalgorithm import geneticalgorithm as ga
# except:
# !pip install geneticalgorithm
# coff a, delta
alpha =1
a=1.5
b = 1.67
delta = 4.0
beta = 2
variable_X = [1,1,1]
error_func_idm(variable_X)
```
| github_jupyter |
# Branching and Merging II: Rewriting History
## Rewriting History
1. Amending a Commit
- You can change the most recent commit
- Change the commit message
- Change the project files
- This creates a new SHA-1 (rewrites history)
- `git commit --amend -m "add fileC.txt"`
- Optionally use the `--no edit` option to reuse the previous commit message
2. Interactive Rebase
- Interactive rebase lets you edit commit using commands
- The commits can belong to any branch
- **The commit history is changed - do not use for shared commit**
- `git rebase -i <after-this-commit>`
- Commits in the current branch after `<after-this-commit>` are listed in an editor and can be modified
- <img src="./images/git_47.png" width="400">
- Interactive Rebase Option provides the following:
- <img src="./images/git_48.png" width="200">
- <img src="./images/git_49.png" width="200">
- Delete a Commit
- The commit's work is not used!
- Squash a Commit
- Applies a new (squashed) commit to an older commit
- Combines the commit messages
- Removes the newer commit
- *Note: A fixup is like a squash, but the squashed commit's message is discarded*
- Squash VS. Delete
- Squash: Combines this commit with the older commit, creating a single commit
- The work of both commits in included
- Delete: No changes from this commit are applied
- The diff thrown out
- The work of this commit is lost
- Greater change of a merge conflict
3. Squash Merges
- Merges the tip of the feature branch (D) onto the top of the base branch (C)
- There is a chance of a merge conflict
- Places the results in the **staging area**
- The results can then be commited (E)
- <img src="./images/git_50.png" width="200">
- Feature Commits?
- After the **featureX** label is deleted, commits B and D are no longer part of any named branch
- A squash merge *rewrites the commit history*
- <img src="./images/git_51.png" width="200">
- Squash Merge Commands:
- `git checkout master`
- `git merge --squash featureX`
- `git commit`
- accept or modify the squash message
- `git branch -D featureX`
- Squash Merge with Fast-Forward
- `git checkout master`
- `git merge --squash featureX`
- `git commit`
- accept or modify the squash message
- `git branch -D featureX`
- Review
- You can amend the most recent commit's message and/or commited files
- It creates a new SHA-1
- Interactive rebase allows you to rewrite the history of a branch
- A squash reduces multiple commits into a single commit
## Review: Rewriting History
- You can amend the most recent commit's message and/or commited files
- It creates a new SHA-1
- Interactive rebase allows you to rewrite the history of a branch
- A squash reduces multiple commits into a single commit
| github_jupyter |
# Checking Location Usage Policy of a Container
for this example we will be using `rdflib` which can be installed using pip
`pip install rdflib`
Since the usage policy contains a named graph, we will use `rdflib.ConjunctiveGraph` class instead of `rdflib.Graph`.
```
from rdflib import ConjunctiveGraph, Namespace, RDF, RDFS, OWL, URIRef
```
In order to access Usage Policy of a semantic container, we access via API on `/api/meta/usage` which will return a usage policy in RDF graph (using `trig` serialization format)
For convenience in querying an RDF Graph, we define some Namespace which we will bind it to some prefixes in the graph later on.
```
#SEMCON_URL = "https://vownyourdata.zamg.ac.at:9600/api/meta/"
SEMCON_URL = "https://vownyourdata.zamg.ac.at:9505/api/meta/"
SC_FILE = Namespace(SEMCON_URL)
SCO = Namespace("http://w3id.org/semcon/ns/ontology#")
SPL = Namespace("http://www.specialprivacy.eu/langs/usage-policy#")
SPL_LOC = Namespace("http://www.specialprivacy.eu/vocabs/locations#")
g = ConjunctiveGraph()
#namespace binding, useful for printout and querying
g.bind('owl', OWL)
g.bind('', SC_FILE)
g.bind('sc', SCO)
g.bind('spl', SPL)
g.bind('loc', SPL_LOC)
#load the actual data
g.parse(SEMCON_URL+"usage", format="trig")
#print the number of RDF triples
print(len(g))
```
First thing first, we will have a look on the policy
```
# print some human-readable serialization
print(g.serialize(format="trig").decode("utf-8"))
```
The policy is defined using OWL vocabulary as a set of restriction on each aspect : Data policy, Processing policy, purpose, recipient, and storage (which contain location and duration).
instead of 'pretty' format above, we can also see how it look as list of triples of subject, predicate, and object
```
# print list of triples
print(g.serialize(format="nt").decode("utf-8"))
```
In order to extract information from the graph, we will use SPARQL query on the graph.
```
qloc = """
SELECT ?q
WHERE {
?policy a owl:Class; owl:equivalentClass [
owl:intersectionOf ?i
].
?i rdf:rest*/rdf:first [
owl:onProperty spl:hasStorage;
owl:someValuesFrom/owl:intersectionOf ?s
].
?s rdf:rest*/rdf:first [
owl:onProperty spl:hasLocation;
owl:someValuesFrom ?q
].
}"""
q = g.query(qloc)
for s in q:
print(s)
qunion = """
SELECT ?q
WHERE {
?policy a owl:Class; owl:equivalentClass [
owl:intersectionOf ?i
].
?i rdf:rest*/rdf:first [
owl:onProperty spl:hasStorage;
owl:someValuesFrom/owl:intersectionOf ?s
].
?s rdf:rest*/rdf:first [
owl:onProperty spl:hasLocation;
owl:someValuesFrom/owl:unionOf ?l
].
?l rdf:rest*/rdf:first ?q.
}
"""
r = list(q)
#check if the returned object is a union or not
if isinstance(r[0][0], URIRef):
print(r[0][0])
else:
loc = g.query(qunion)
for l in loc:
print(l[0])
```
| github_jupyter |
# Maximum Likelihood Estimation
In this notebook, we will see what Maximum Likelihood Estimation is and how it can be used.
**Contents**:
* **[Introduction](#Introduction)**
* **[Mathematical Formulation](#Mathematical-Formulation)**
* **[Difference between Probability and Likelihood](#Difference-between-Probability-and-Likelihood)**
* **[Probability Distributions, Estimations and Codes](#Probability-Distributions,-Estimations-and-Codes)**
* **[Bernoulli Distribution](#Bernoulli-Distribution)**
* **[Binomial Distribution](#Binomial-Distribution)**
* **[Poisson Distribution](#Poisson-Distribution)**
* **[Exponential Distribution](#Exponential-Distribution)**
* **[Normal Distribution](#Normal-Distribution)**
* **[Summary](#Summary)**
* **[References](#References)**
---
# <a id="Introduction">Introduction</a>
We often need to study populations for various purposes such as understanding its characteristics, for finding patterns and to devise better solutions using these information. However, for large populations, it is neither possible nor feasible to examine each individual in a population to understand these properties and characteristics. Hence, the statisticians make use of random sampling. The measurements are, therefore, made on much smaller samples for analysis and drawing conclusions about the population.
Analysing the data becomes a lot more easier if we are able to identify distribution of the population, using the sample. Any probability distribution is uniquely defined by some parameters. Therefore, if we somehow find/identify these parameters, we can easily study the population. This process of finding parameters for the population by analysing the samples from that population is called **estimation**.
We can say that whenever our estimation is good, on random sampling again, we are **likelier** to obtain a sample which would be very similar to our original sample. In other words, more the chances of getting a similar sample with our estimates of the parameters, more is the **likelihood** that our estimation is satisfactorily correct.
**Maximum likelihood estimation** is basically the process of estimating these parameters based on our sample, such that the likelihood of the population to be defined with those parameters is maximised.
---
# <a id="Mathematical-Formulation">Mathematical Formulation</a>
Now that we have a gist of the problem we are trying to solve here, let's define it more formally.
Suppose we have obtained a random sample from our population, given by $x_1, \ x_2, \ x_3, \ ..., \ x_n$. <br>
Now, using this sample, we want to assume some distribution for our population. Let that distribution be defined as $D(\theta)$, <br>
where $\theta = [\theta_1 \ \theta_2 \ \theta_3 \ ... \ \theta_k]^T$ is the vector defining the parameters for our assumed distribution $D$.
Now, using our random sample, we want to determine the values of $\theta$ which will make our distribution $D$ most likely to define our population. In order to quantify how likely it is for our assumed distribution $D$ with its estimated parameters $\theta$ to define the population, we make use of **likelihood function**.
We define the likelihood function,
$L : \{\theta, \ x_1, \ x_2, \ x_3, \ ..., \ x_n\} \longrightarrow \mathbb{R}_0^+$ as follows: <br>
$$
\begin{aligned}
L(\theta \ | \ x_1, \ x_2, \ x_3, \ ..., \ x_n)
& = P(X_1 = x_1,X_2 = x_2,X_3 = x_3, \ ..., \ X_n = x_n) \\
& = f(x_1 | \theta)\ .\ f(x_2 | \theta)\ .\ f(x_3 | \theta)\ .\ ...\ .\ f(x_n | \theta) \\
& = \prod_{i = 1}^n f(x_i | \theta)
\end{aligned}
$$
Here, $f$ is the probability mass/density function for our distribution $D$.
**Note**: Since the sample is selected randomly from a large population, we can assume that selection of a datum into the sample is **independent** from selection of any other datum. This assumption of independence allows us to multiply the probability mass/density functions for each datum to calculate the overall likelihood.
The **goal** of **Maximum Likelihood Estimation** is to find $\theta$ such that the likelihood function value gets maximised. For any likelihood function which is **concave**, or **negatively convexed**, we can find its maximum likelihood estimate by equating the first partial derivatives w.r.t. each parameter $\theta_j, \forall j \in \{1, 2, 3, \ ..., \ k\}$ to $0$. As we will see later, for commonly occuring probability distributions, the likelihood functions are indeed concave.
$$
\begin{aligned}
\therefore \ \theta
& = \text{arg} \ \max_{\theta \ \in \ \Theta} \ L(\theta \ | \ x_1, x_2, x_3, \ ..., \ x_n) \\
\Longrightarrow \ \theta_j
& = \text{arg} \left( \frac{\partial L}{\partial \theta_j} = 0 \right)
\ \forall \ j \in \{1, \ 2, \ 3, \ ..., \ k\} \ | \ \text{L is concave}
\end{aligned}
$$
Equating partial derivatives to $0$ would be difficult here because product of probability mass/density function for all the elements of the sample would require extended application of chain rule.
Hence, we make use of **logarithm function**. Logarithm function is monotonically strictly increasing function. Therefore, wherever $L$ will maximise, at the same parameter values, $\log(L)$ will also maximize. Therefore, instead of maximising the likelihood estimation, we maximise the **log likelihood estimation**.
$$
\begin{aligned}
L(\theta \ | \ x_1, x_2, x_3, \ ..., \ x_n) & = \prod_{i = 1}^n f(x_i \ | \ \theta) \\
\Longrightarrow \log\left(L(\theta \ | \ x_1, x_2, x_3, \ ..., \ x_n)\right)
& = \log\left(\prod_{i = 1}^n f(x_i \ | \ \theta)\right) \\
& = \sum_{i = 1}^n \log\left(f(x_i \ | \ \theta)\right) \\
\Longrightarrow \frac{\partial}{\partial \theta_j}
\log\left(L(\theta \ | \ x_1, x_2, x_3, \ ..., \ x_n)\right)
& = \sum_{i = 1}^n \frac{\partial}{\partial \theta_j} \log\left(f(x_i \ | \ \theta)\right),
\ \forall \ j \in \{1, 2, 3, \ ..., \ k\} \\
& = \sum_{i = 1}^n \frac{1}{f(x_i \ | \ \theta)} \frac{\partial}{\partial \theta_j} f(x_i \ | \ \theta),
\ \forall \ j \in \{1, 2, 3, \ ..., \ k\} \\
\therefore \ \text{for concave likelihood functions}, \ \theta_j
& = \text{arg} \left(\sum_{i = 1}^n \frac{1}{f(x_i \ | \ \theta)} \frac{\partial}{\partial \theta_j}
f(x_i \ | \ \theta) = 0\right), \ \forall \ j \in \{1, 2, 3, \ ..., \ k\} \\
\end{aligned}
$$
Now that we have seen the way to maximise the likelihood of estimation, let's do this for some probability distributions in the next section.
---
# <a id="Difference-between-Probability-and-Likelihood">Difference between Probability and Likelihood</a>
In the beginning, one may think probability and likelihood to be the same thing with only thier names different. This is primarily because the mathematical formula for both of these entities are quite similar. So let's see how likelihood is different from probability.
Roughly speaking, **probability** tells you the **possibility of observing specific results** or outcomes given a **particular distribution**. When we compute probability, we are already aware about the distribution parameters and hence the **distribution is fixed**. Therefore, with the help of probability, we are able to determine how probable it is to obtain the results in consideration for an event.
On the other hand, in likelihood, we deal with the reverse scenario. Likelihood tells you the possibility that something that you have observed is specifically distributed. This means, for the case of **likelihood** calculation, we already have our **results fixed**. In this case, the **the distribution parameters** play the role of **variables**. Therefore, using our results, we try to find how well some particular distribution fits on our results.
Mathematically, <br>
- Probability is described by a function $P : \{x_1, \ x_2, \ x_3, \ ..., \ x_n, \theta\} \longrightarrow [0, 1]$, computed as $P(x_1, \ x_2, \ x_3, \ ..., \ x_n \ | \ \theta)$
- Whereas, likelihood can be written as a function $L : \{x_1, \ x_2, \ x_3, \ ..., \ x_n, \theta\} \longrightarrow \mathbb{R}_0^+$, computed as $L(\theta \ | \ x_1, \ x_2, \ x_3, \ ..., \ x_n)$
where, $x_1, \ x_2, \ x_3, \ ..., \ x_n$ are the results/observations, and $\theta$ is the vector of distribution parameters
Let's consider an example to understand this better. <br>
Suppose we are given an exponential distribution with its rate, i.e., $\lambda$ equal to $2$. This is how the probability density function for this distribution will look:
```
# make matplotlib plots plotted inline with the notebook
%matplotlib inline
# import packages
import matplotlib.pyplot as plt
import numpy as np
import scipy.stats as ss
# find 10000 exponentially distributed values between 0 and 3
rate = 2 # lambda
x = np.linspace(0, 3, 10000) # 10000 uniformly spaced values between 0 and 3
y_pdf = ss.expon.pdf(x, scale=(1 / rate))
# scale is inverse of lambda, pdf function returns pdf values
# plot the values to visualise the distribution
plt.plot(x, y_pdf) # plot x againtst y values
plt.title('Plot of exponential distribution with λ = 2') # give title
plt.ylabel('Probability Density Function') # give y axis label
plt.xlabel('Random variable') # give x axis label
plt.show() # show the plot
```
Now suppose we want to know how likely is it that we'll find a waiting time, $x$, between two successive events as $x \geq 0.75 \wedge x \leq 1.5$ with this distribution. For that, we calculate the probability as $P(0.75 \leq x \leq 1.5 \ | \ \lambda = 2)$. This will be same as finding area under the curve.
```
# define the bounds
lower_bound = 0.75
upper_bound = 1.5
# get indices in x with values within these bounds
mask = (x >= lower_bound) & (x <= upper_bound)
x_area = x[mask] # get values of x which are between the bounds
y_area = y_pdf[mask] # get corresponding values for the PDF function
plt.plot(x, y_pdf); # plot the PDF
plt.fill_between(x_area, y_area, 0, alpha=0.5) # plot area under the curve
plt.title('Probability that x is between 0.75 and 1.5, when λ = 2')
plt.ylabel('Probability Density Function')
plt.xlabel('Random variable')
plt.show()
```
Now, this area, when calculated, will be equal to equal to $\text{CDF}(1.5) - \text{CDF}(0.75)$, which is equal to 0.2492 (approx.).
```
# code to find the probability value:
ss.expon.cdf(upper_bound) - ss.expon.cdf(lower_bound)
```
In this case, if you would see, we had the value of the distribution parameter, that is $\lambda$ already fixed to $2$. And using those fixed values, we calculated how probable it would be to find observations between $0.75$ and $1.5$.
Now, suppose we already have some results - let's say $x = 0.75$. Now using this, in case of likelihood, we want to evaluate how likely is it that this datum was obtained from a specific distribution. So let's try to first see the likelihood when $\lambda = 2$. For that, we calculate the likelihood as: $L(\lambda = 2 \ | \ x = 0.75) = \text{pdf}(x = 0.75 \ | \ \lambda = 2)$.
```
# define the observation and the rate values
observation = 0.75
rate = 2
# compute the likelihood value
likelihood = ss.expon.pdf(observation, scale=(1 / rate))
plt.plot(x, y_pdf) # plot the PDF
plt.plot([0, observation], [likelihood, likelihood], '-r') # plot the abscissa
# plot the oordinate
plt.plot([observation, observation], [0, likelihood], '-r')
plt.plot([observation], [likelihood], 'or') # plot the point
plt.annotate(f'({observation}, {round(likelihood, 4)})', xy=(observation, likelihood)) # annotate the point
plt.title('Likelihood that λ = 2 when x = 0.75')
plt.ylabel('Probability Density Function')
plt.xlabel('Random variable')
plt.xlim(0, 3) # define range on x axis
plt.ylim(0, 2) # define range on y axis
plt.show()
```
In this case, we can see that likelihood came out to be a single point on the PDF function curve. Hence, the likelihood that $\lambda = 2$ for exponential distribution when $x = 0.75$ is $0.4463$.
Similarly, for $\lambda = 0.5$, the likelihood will be: $0.3436$. <br>
This means it is likelier that this observation has been drawn from an exponential distribution with rate $= 2$ rather than rate $= 0.5$.
```
# define the observation and the rate values
observation = 0.75
rate = 0.5
# compute the likelihood value
likelihood = ss.expon.pdf(observation, scale=(1 / rate))
y_pdf = ss.expon.pdf(x, scale=(1 / rate)) # pdf values with scale changed
plt.plot(x, y_pdf); # plot the PDF
plt.plot([0, observation], [likelihood, likelihood], '-r'); # plot the abscissa
# plot the oordinate
plt.plot([observation, observation], [0, likelihood], '-r');
plt.plot([observation], [likelihood], 'or'); # plot the point
plt.annotate(f'({observation}, {round(likelihood, 4)})', xy=(observation, likelihood)); # annotate the point
plt.title('Likelihood that λ = 0.5 when x = 0.75')
plt.ylabel('Probability Density Function')
plt.xlabel('Random variable')
plt.xlim(0, 3)
plt.ylim(0, 2)
plt.show()
```
---
# <a id="Probability-Distributions,-Estimations-and-Codes">Probability Distributions, Estimations and Codes</a>
Now that we know what maximum likelihood estimation is and how we are supposed to do it, let's use it to estimate distributions for samples taken from some commonly occurring probability distributions.
## <a id="Bernoulli-Distribution">Bernoulli Distribution</a>
Bernoulli Distribution is a **discrete** distribution which can be used to model a **single yes-no event**. An example of such an event is a coint toss. A coin can either land with heads up, or tails up. Since only two outcomes are possible in a Bernoulli trial, therefore, it has only **one parameter**, i.e., the **probability of obtaining one of the outcomes**. For the other outcome, because of the two outcomes being complementary, the probability for it to occurr is determined automatically.
Mathematically, <br>
A random variable $X$ is said to have a Bernoulli Distribution if, for $x \in X$:
$$
f(x) = \left\{
\begin{matrix}
p & \text{if } x = 1 \\
1 - p & \text{if } x = 0 \\
0 & \text{otherwise}
\end{matrix}
\right.
$$
Here, $f$ is the probability mass function, $p$ is the probability for one of the two possible outcomes to happen, $x = 1$ implies that the first outcome has successfully occurred. Therefore, when $x = 0$, the first outcome has failed to occur which implies that the second outcome has occurred. Hence, the probability mass function can be condensed to the following representation:
$$
f(x) = p^x(1 - p)^{1 - x}, \ x \in \{0, 1\}
$$
Now let's say we have a sample, $x_1, \ x_2, \ x_3, \ ..., \ x_n$ which we have obtained supposedly from $n$ independent and identical Bernoulli trials. So, we will now need to estimate what parameter $p$, for our assumed Bernoulli Distribution, will best describe our sample. For this, we proceed onto calculating the likelihood.
$$
\begin{aligned}
L(p \ | \ x_1, x_2, x_3, \ ..., \ x_n) & = \prod_{i = 1}^n f(x_i \ | \ p) \\
& = \prod_{i = 1}^n p^{x_i}(1 - p)^{1 - x_i}
\end{aligned}
$$
Therefore, the log likelihood function would be:
$$
\begin{aligned}
\log L(p \ | \ x_1, x_2, x_3, \ ..., \ x_n)
& = \log\left(\prod_{i = 1}^n p^{x_i}(1 - p)^{1 - x_i}\right) \\
& = \sum_{i = 1}^n \log\left(p^{x_i}(1 - p)^{1 - x_i}\right) \\
& = \sum_{i = 1}^n \log\left(p^{x_i}\right) + \sum_{i = 1}^n \log\left((1 - p)^{1 - x_i}\right) \\
& = \log(p) \sum_{i = 1}^n x_i + \log(1 - p) \sum_{i = 1}^n (1 - x_i) \\
\text{Let's consider} \ \frac{1}{n}\sum_{i = 1}^n x_i
& = c, \ \text{where} \ c \ \text{is a constant for a given sample, }
0 \leq c \leq 1 \text{, independent of } p. \\
\therefore \log L(p \ | \ x_1, x_2, x_3, \ ..., \ x_n)
& = n[c\log(p) + (1 - c)\log(1 - p)] \\
\end{aligned}
$$
Let's try to visualise this function. Since $n$ will only scale the function, we can ignore it for now. Also, the surface plot did not give a representation intuitive enough that the function is concave, so we have made these plots for $c \in \{0.1, 0.5, 0.9\}$
```
# include packages
import numpy as np
import matplotlib.pyplot as plt
# make arrays for p and c
p = np.linspace(0.001, 0.999, 1000)
c_s = np.linspace(0.1, 0.9, 3)
# define the figure
fig = plt.figure()
ax = plt.axes()
# plot the log-likelihood function for each value of c
for c in c_s:
# compute log-likelihood function values
log_likelihood = c * np.log(p) + (1 - c) * np.log(1 - p)
ax.plot(p, log_likelihood, label=f'c = {c}') # plot log-likelihood function
plt.legend() # put legend on plot
plt.xlabel('Probability(x = 1), p')
plt.ylabel('Log-likelihood')
plt.title('Log-likelihood function for Bernoulli Distribution')
plt.show()
```
Here we observe that log-likelihood function is concave and we can, therefore, perform maximum likelihood estimation by equating partial derivates of this log-likelihood function w.r.t. $p$ to be $0$.
Partially differentiating log-likelihood function w.r.t. $p$, we get:
$$
\begin{aligned}
\frac{\partial}{\partial p}\log L(p \ | \ x_1, x_2, x_3, \ ..., \ x_n)
& = n\left[c \frac{\partial \log(p)}{\partial p}
+ (1 - c) \frac{\partial \log(1 - p)}{\partial p}\right] \\
& = n\left[c \frac{1}{p} - (1 - c) \frac{1}{1 - p}\right] \\
\end{aligned}
$$
Equating the partial derivative with 0, to find $p$ which gives maximum likelihood, we get:
$$
\begin{aligned}
\frac{\partial}{\partial p}\log L(p \ | \ x_1, x_2, x_3, \ ..., \ x_n) & = 0 \\
\Longrightarrow \frac{c}{p} & = \frac{1 - c}{1 - p} \\
\Longrightarrow p & = c \\
\therefore p & = \frac{1}{n} \sum_{i = 1}^n x_i = \bar{x}
\end{aligned}
$$
Hence, we find that for **MLE of Bernoulli Distribution**, the parameter $p$ should be kept as the **mean of the sample**. Now let's demonstrate this on an example.
```
# Let actual distribution parameter, p, be uniformly picked between 0 and 1
p = np.random.rand()
N = [1, 10, 100, 1000] # perform MLE with these many points in samples
sample = {} # to store the obtained samples
for n in N:
# Simulate n identical and independent Bernoulli trials
sample_n = []
for i in range(n):
# perform single Bernoulli trial and add the result to sample
sample_n.append(np.random.binomial(n=1, p=p))
sample[n] = sample_n # add sample to dictionary
print(sample[100]) # representation of a sample obtained
# Now, let's try to perform maximum likelihood estimation using these samples.
# We saw that we estimate p to be equal to the sample mean
for n in N:
p_hat = np.mean(sample[n])
print(f'For n = %4d, we estimate the parameter p to be {p_hat}' % n)
# Now, let's see the actual value of p ;)
print(f'The actual value of p is {p}')
```
From this example, we see that with the help of maximum likelihood estimation, for large samples, we are able to estimate the values of the distribution parameter reasonably well.
---
## <a id="Binomial-Distribution">Binomial Distribution</a>
Binomial Distribution is a discrete probability distribution which models the **number of successes** in an experiment in which $n$ **Bernoulli trials** were conducted successively and independently, such that probability for success in any of these trials is $p$. For example, number of heads obtained in $10$ coin tosses will be binomially distributed.
Formally,
For a random variable $X \sim \text{Bin}(n, p)$ if for $x \in X$, we have:
$$
f(x) = \left\{
\begin{matrix}
\frac{n!}{(n - x)! x!} p^x (1 - p)^{n - x}
& \text{if } x \in \{z \ : \ z \in \mathbb{Z} \ \wedge \ 0 \leq z \leq n\} \\
0 & \text{otherwise }
\end{matrix}
\right.
$$
where, $f$ is the probability mass function and $x_i$ is the number of successes in $i^{th}$ Bernoulli trial.
Now suppose we have a sample $x = [x_1 \ x_2 \ x_3 \ ... \ x_m]^T$, obtained after performing identical experiments $m$ times, and we assume it to come from a binomial distribution. Therefore, its likelihood function and log-likelihood function will be defined as:
$$
\begin{aligned}
L(p \ | \ n, x) & = \prod_{i = 1}^m f(x_i \ | \ n, p) \\
& = \prod_{i = 1}^m \frac{n!}{(n - x_i)! x_i!} p^{x_i} (1 - p)^{n - x_i} \\
\therefore \log L(p \ | \ n, x)
& = \log\left(\prod_{i = 1}^m \frac{n!}{(n - x_i)! x_i!} p^{x_i} (1 - p)^{n - x_i}\right) \\
& = \sum_{i = 1}^m \log\left(\frac{n!}{(n - x_i)! x_i!} p^{x_i} (1 - p)^{n - x_i}\right) \\
& = \sum_{i = 1}^m \log\left(\frac{n!}{(n - x_i)! x_i!}\right)
+ \sum_{i = 1}^m \log\left(p^{x_i} (1 - p)^{n - x_i}\right) \\
& = \sum_{i = 1}^m \log\left(\frac{n!}{(n - x_i)! x_i!}\right)
+ \log(p) \sum_{i = 1}^m x_i + \log(1 - p) \sum_{i = 1}^m (n - x_i) \\
\text{Let's consider} \ \frac{1}{m}\sum_{i = 1}^m x_i
& = c, \ \text{where} \ c \ \text{is a constant for a given sample, }
0 \leq c \leq n \text{, independent of } p. \\
\therefore \log L(p \ | \ n, x)
& = \sum_{i = 1}^m \log\left(\frac{n!}{(n - x_i)! x_i!}\right)
+ m[c\log(p) + (n - c)\log(1 - p)] \\
\end{aligned}
$$
Since $\sum_{i = 1}^m \log\left(\frac{n!}{(n - x_i)! x_i!}\right)$ is constant for any given sample, and $m$ would only scale the rest of the terms, we can drop these two things while log-likelihood maximisation.
Let $\log L'(p \ | \ n, x) = c\log(p) + (n - c)\log(1 - p)$.
Therefore, we have:
$$
\begin{aligned}
\frac{\partial}{\partial p} \log L'(p \ | \ n, x)
& = c \frac{\partial}{\partial p} \log(p) + (n - c) \frac{\partial}{\partial p} \log(1 - p) \\
& = \frac{c}{p} - \frac{(n - c)}{1 - p} \\
\Longrightarrow \frac{\partial^2}{\partial p^2} \log L'(p \ | \ n, x)
& = - \frac{c}{p^2} - \frac{(n - c)}{{(1 - p)}^2} < 0 \
\forall \ p \ \in \ (0, 1), \text{ implying concavity} \\
\therefore \text{maxima of } L' \text{ will be when }
\frac{\partial}{\partial p} \log L'(p \ | \ n, x) & = 0 \\
\Longrightarrow \frac{c}{p} & = \frac{(n - c)}{1 - p} \\
\Longrightarrow p & = \frac{c}{n} \\
\Longrightarrow p & = \frac{1}{mn} \sum_{i = 1}^m x_i \\
& = \frac{1}{n} \bar{x}
\end{aligned}
$$
Therefore, for **MLE of Binomial Distribution**, the parameter $p$ should be kept as the **mean of proportion of successes across all Binomial trials**. Let's see an example for the same.
```
# import packages
import numpy as np
import matplotlib.pyplot as plt
import scipy
# Let actual distribution parameter, p, be uniformly picked between 0 and 1
p = np.random.rand()
# Now, n has to be fixed, but it can be anything. So let n be 50
n = 50
# Now, suppose we conduct the Binomial trials m times, Let M = m_i for ith trial
M = [1, 10, 100, 1000] # perform MLE with these many points in samples
sample = {} # to store the obtained samples
for m in M:
# Simulate m identical and independent Binomial trials
sample_m = []
for i in range(m):
# perform n Bernoulli trials and add the result to sample
sample_m.append(np.random.binomial(n=n, p=p))
sample[m] = sample_m # add sample to dictionary
print(sample[100]) # a typical sample
# Now, let's try to perform maximum likelihood estimation using these samples.
# We saw that we estimate p to be equal to the mean of total number of successes
for m in M:
p_hat = np.mean(sample[m]) / n
print(f'For m = %4d, we estimate the parameter p to be {p_hat}' % m)
# Plot histogram for sample with 1000 Binomial experiments
# plot the histogram
plt.hist(sample[1000], bins=n, range=(0, n), label='Histogram for sample')
# Plot the distribution
# find nPx for all values of x
x = np.linspace(0, n, num=(n + 1))
n_s = n * np.ones(n + 1)
perms = scipy.special.comb(n_s, x) # calculate nPx
# calculate probabilities - p^x * (1 - p)^(n - x)
p_hat = np.mean(sample[1000]) / n
probs = (p_hat ** x) * ((1 - p_hat) ** (n - x))
y = np.multiply(perms, probs) # multiply perms with probs
# scale y in range of sample counts
y = y / np.max(y) * scipy.stats.mode(sample[1000]).count[0]
# distribution shifted by 0.5 in plot to match with histogram bars
plt.plot(x + 0.5, y, label='Estimated Binomial Distribution (scaled)')
plt.title('Histogram to show distribution of sample')
plt.xlabel('Value of x observed')
plt.ylabel('Counts')
plt.legend(bbox_to_anchor=(1.05, 1), loc='best') # put legend outside the plot
plt.show()
```
Hence, we see that the binomial distribution with the value of its parameter that we calculated can estimate the sample well.
```
# Now, let's see the actual value of p
print(f'The actual value of p is {p}')
```
Contrary to the example in Bernoulli distribution, we see that Binomial distribution parameter, $p$, can be estimated with decent accuracy even if we do not have a lot of points in the sample, given by $m$, provided our sample has been taken from results of large number of Bernoulli trials, $n$.
---
## <a id="Poisson-Distribution">Poisson Distribution</a>
Poisson distribution is used to model the **number of occurrences** of any event which has a **constant mean rate** and whose occurrence is **independent of the last occurrence**, in a **fixed span** of quantifiable entity such as time, distance, area, volume etc. For example, the number of structural defects found per volume of an engineering material, or the number of calls received in an hour such that the average number of calls received in a day is usually the same.
The probability mass function for Poisson distribution is given as:
$$
f(k \ | \ \lambda) =
\left\{
\begin{matrix}
\frac{\lambda^k e^{-\lambda}}{k!} & k \in \mathbb{Z}^+_0 \\
0 & \text{otherwise} \\
\end{matrix}
\right.
$$
where $k$ determines the number of occurrences of an event in a fixed span which has a parameter, expected number of occurrences in that span, given by $\lambda$.
Therefore, if we have a sample $x = [x_1 \ x_2 \ x_3 \ ... \ x_m]^T$ in which $x_i$ denotes the number of occurrences of the event in consideration in $i^{th}$ Poisson experiment, then its likelihood and log-likelihood estimation for the parameter $\lambda$ will be given as:
$$
\begin{aligned}
L(\lambda \ | \ x) & = \prod_{i = 1}^m f(x_i \ | \ \lambda) \\
& = \prod_{i = 1}^m \frac{\lambda^{x_i} e^{-\lambda}}{{x_i}!} \\
\therefore \log L(\lambda \ | \ x)
& = \log \left(\prod_{i = 1}^m \frac{\lambda^{x_i} e^{-\lambda}}{{x_i}!}\right) \\
& = \sum_{i = 1}^m \log \left(\frac{\lambda^{x_i} e^{-\lambda}}{{x_i}!}\right) \\
& = \log \lambda \sum_{i = 1}^m x_i + \sum_{i = 1}^m (-\lambda)
- \sum_{i = 1}^m \log({x_i}!) \\
\text{Let } \sum_{i = 1}^m x_i & = a \text{, where } a \text{ is a constant for a given sample.} \\
\therefore \log L(\lambda \ | \ x)
& = a \log \lambda -\lambda m - \sum_{i = 1}^m \log({x_i}!) \\
\end{aligned}
$$
Now, for a given sample, $\sum_{i = 1}^m \log({x_i}!)$ will be a constant independent of $\lambda$, and therefore can be dropped from consideration for maximisation of likelihood. Let $\log L'(\lambda \ | \ x) = a \log \lambda -\lambda m$. Therefore, differentiating w.r.t. to $\lambda$, we get:
$$
\begin{aligned}
\frac{\partial}{\partial \lambda} \log L'(\lambda \ | \ x)
& = a \frac{\partial}{\partial \lambda} \log \lambda - m \frac{\partial}{\partial \lambda} \lambda \\
& = \frac{a}{\lambda} - m \\
\frac{\partial^2}{\partial \lambda^2} \log L'(\lambda \ | \ x)
& = - \frac{a}{\lambda^2} < 0 \ \forall \ \lambda \ \in \ \mathbb{R}^+ \text{, implying concavity} \\
\therefore \text { for maximum log-likelihood estimation,} \\
\frac{\partial}{\partial \lambda} \log L'(\lambda \ | \ x) & = 0 \\
\Longrightarrow \frac{a}{\lambda} - m & = 0 \\
\Longrightarrow \lambda & = \frac{a}{m} \\
& = \frac{1}{m} \sum_{i = 1}^m x_i \\
& = \bar{x}
\end{aligned}
$$
This means for **Poisson Distribution**, our parameter, $\lambda$, should be equal to the **sample arithmetic mean** for maximum-likelihood estimation. Next, let's see an example on a practical dataset.
We will be using the **Prussian Horse Kick Data** (Andrews & Herzberg, 1985). This is a publicly available dataset containing the number of soldier deaths due to horse-kicks in Prussian Cavalry between 1875 and 1894, on an annual basis across 14 different cavalry corps. So let's get started.
```
# import packages
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import scipy
# load the data from the text file stored in 'data' directory
horse_kick_data = pd.read_csv('../data/HorseKicks.txt', sep='\t', index_col='Year')
horse_kick_data.head()
# let us flatten our dataframe into an array,
# such that each element of the array represents the number of deaths
# due to horse-kicks per corp per year
sample = horse_kick_data.to_numpy().flatten()
sample
# Let us now estimate the distribution parameter, 𝜆
lambda_hat = np.mean(sample)
lambda_hat
# Let us now plot the histogram as well as the estimated distribution
# plot the histogram
plt.hist(sample, bins=(sample.max() + 1), range=(0, sample.max() + 1),
label='Histogram for sample')
# plot the distribution
# find the pmf values
x_s = np.array(range(sample.max() + 1))
y_s = (lambda_hat ** x_s) * np.exp(-lambda_hat) / scipy.special.factorial(x_s)
# scale the pmf values
y_s = y_s / y_s.max() * scipy.stats.mode(sample).count[0]
# distribution shifted by 0.5 in the plot
plt.plot(x_s + 0.5, y_s, label='Estimated Poisson Distribution (Scaled)')
# customize plot
plt.title('Sample Histogram and Estimated Distribution')
plt.xlabel('Number of deaths per corp per year')
plt.ylabel('Counts')
plt.legend()
plt.show()
# Table to show Expected counts with the estimated distribution
# vs the actual counts observed from the sample
counts = pd.DataFrame() # define an empty dataframe
expected_counts = y_s
actual_counts = []
# find actual counts for all values in the sample
for x in x_s:
actual_counts.append((sample == x).sum())
# assign column-wise values to the dataframe
counts['Number of deaths per corp per year'] = x_s
counts['Actual Counts'] = actual_counts
counts['Expected Counts'] = expected_counts
# change index column
counts = counts.set_index('Number of deaths per corp per year')
# see the resultant dataframe
counts
```
Therefore, we see that our estimated distribution fits well to the sample and can be further used to determine other relevant information.
---
## <a id="Exponential-Distribution">Exponential Distribution</a>
**Exponential Distribution** models the **span of entities** such as time or distance **between consecutive occurrences** of an event following **Poisson Distribution**. Unlike the previously seen distributions, this is a **continuous distribution** since span can take continuous values. Like Poisson process, this distribution also has a property of **memorylessness**, which means that the previous occurrences and the time between them do not affect the occurrences happening later.
Probability density function for Exponential Distribution is given by:
$$
f(x \ | \ \lambda) = \left\{
\begin{matrix}
\lambda e^{-\lambda x} & x \geq 0 \\
0 & \text{otherwise} \\
\end{matrix}
\right.
$$
where $\lambda$ is the mean occurrence rate of the event under consideration, and $x$ determines the span of entity observed between consecutive occurrences.
For a sample $x = [x_1 \ x_2 \ x_3 \ ... \ x_m]^T$, such that $x_i$ is the span of entitiy between the occurrence of $i^{th}$ and $(i + 1)^{th}$ events, the likelihood and log-likelihood functions and the condition for maximum likelihood estimation would be as follows:
$$
\begin{aligned}
L(\lambda \ | \ x) & = \prod_{i = 1}^m f(x_i \ | \ \lambda) \\
& = \prod_{i = 1}^m \lambda e^{-\lambda x_i} \\
\therefore \log L(\lambda \ | \ x)
& = \log \left(\prod_{i = 1}^m \lambda e^{-\lambda x_i}\right) \\
& = \sum_{i = 1}^m \log \left(\lambda e^{-\lambda x_i}\right) \\
& = m \log \lambda - \lambda \sum_{i = 1}^m x_i \\
\text{Let } \sum_{i = 1}^m x_i & = a \text{, where } a \text{ is a constant for a given sample.} \\
\therefore \log L(\lambda \ | \ x)
& = m \log \lambda -\lambda a \\
\therefore \frac{\partial}{\partial \lambda} \log L(\lambda \ | \ x)
& = m \frac{\partial}{\partial \lambda} \log \lambda - a \frac{\partial}{\partial \lambda} \lambda \\
& = \frac{m}{\lambda} - a \\
\Longrightarrow \frac{\partial^2}{\partial \lambda^2} \log L(\lambda \ | \ x)
& = - \frac{m}{\lambda^2} < 0 \ \forall \ \lambda \ \in \ \mathbb{R}^+ \text{, implying concavity} \\
\Longrightarrow \text{For maximum log-likelihood estimation,} \\
\frac{\partial}{\partial \lambda} \log L(\lambda \ | \ x) & = 0 \\
\Longrightarrow \frac{m}{\lambda} - a & = 0 \\
\Longrightarrow \lambda & = \frac{m}{a} \\
& = \frac{m}{\sum_{i = 1}^m x_i} \\
& = \frac{1}{\bar{x}}
\end{aligned}
$$
Therefore, we see that for maximum likelihood estimation of **Exponential Distribution**, the parameter $\lambda$ should be equal to the **reciprocal of sample arithmetic mean**. Let's see an example. In this example, we shall refer the **Mileage** (Modarres M et al., 2017) standard dataset from the reliability package (needed to be installed additionally), which contains the information of distances after which failures occurred in vehicles.
```
# pip install reliability
# run the above stated command first to install reliability package
# import packages
import reliability
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# import the dataset and see its help
mileage = reliability.Datasets.mileage
print(help(mileage))
# get list of distances at which failures occurred
failures = mileage().failures
# sort the list so that distance between consecutive failures can be calculated
failures.sort()
# see the list
print(failures)
# let's populate the list of time taken between consecutive failures
time_between_failures = []
for i in range(len(failures) - 1):
time_between_failures.append(failures[i + 1] - failures[i])
# let's see the final data obtained
print(time_between_failures)
```
Let's now try to fit exponential distribution on it.
```
# estimate lambda
lambda_hat = 1 / np.mean(time_between_failures)
lambda_hat # these many average failures per unit distance covered
# Let's plot the time between failures and the estimated distribution
# plot the histogram and store bin counts
bin_counts = plt.hist(time_between_failures, bins=20,
range=(0, np.max(time_between_failures)),
label='Histogram')[0]
# plot the estimated distribution
# find pdf values
x_s = np.linspace(0, np.max(time_between_failures), 20)
y_s = lambda_hat * np.exp(-lambda_hat * x_s)
# scale pdf values
y_s = y_s / np.max(y_s) * np.max(bin_counts)
# plot x_s vs y_s
plt.plot(x_s, y_s, label='Estimated Distribution (scaled)')
# customize the plot
plt.title('Sample Histogram and Estimated Distribution')
plt.xlabel('Distance between failures')
plt.ylabel('Counts')
plt.legend()
plt.show()
```
We see that our estimated exponential distribution fits the data reasonably well.
---
## <a id="Normal-Distribution">Normal Distribution</a>
**Normal distribution** is one of the commonly occurring distributions found in the nature. It can model **real-valued values** whose distribution are not known. It has a central values, called **mean**, and the distribution is even at both sides of the mean, whose spread is determined by another parameter, that is **variance**. More formally, a random variable $X \sim N(\mu, \sigma^2)$, where $\mu$ is the mean, and $\sigma^2$ is the variance, will follow the following **probability density function**:
$$
f(x \ | \ \mu, \sigma^2) = \frac{1}{\sqrt{2 \pi \sigma^2}} e^{- \frac{(x - \mu)^2}{2 \sigma^2}}
$$
For a sample $x = [x_1 \ x_2 \ x_3 \ ... \ x_m]^T$, the likelihood and log-likelihood functions would be defined as follows:
$$
\begin{aligned}
L(\mu, \sigma^2 \ | \ x) & = \prod_{i = 1}^m f(x_i \ | \ \mu, \sigma^2) \\
& = \prod_{i = 1}^m \left( \frac{1}{\sqrt{2 \pi \sigma^2}}
e^{- \frac{(x_i - \mu)^2}{2 \sigma^2}} \right) \\
\therefore \log L(\mu, \sigma^2 \ | \ x)
& = \log \left( \prod_{i = 1}^m \frac{1}{\sqrt{2 \pi \sigma^2}}
e^{- \frac{(x_i - \mu)^2}{2 \sigma^2}} \right) \\
& = \sum_{i = 1}^m \log \left( \frac{1}{\sqrt{2 \pi \sigma^2}}
e^{- \frac{(x_i - \mu)^2}{2 \sigma^2}} \right) \\
& = \sum_{i = 1}^m \log \left( \frac{1}{\sqrt{2 \pi \sigma^2}} \right)
- \sum_{i = 1}^m \left( \frac{(x_i - \mu)^2}{2 \sigma^2} \right) \\
\end{aligned}
$$
Now, in case of normal distribution, we have two independent parameters and hence, we will need to maximise the likelihood w.r.t. both of them. We can do this by calculating the partial derivatives w.r.t. both of them and then equating both of them to $0$, provided that the log-likelihood function w.r.t. both of them is concave.
Differentiating w.r.t. $\mu$, we get:
$$
\begin{aligned}
\frac{\partial}{\partial \mu} \log L(\mu, \sigma^2 \ | \ x)
& = \sum_{i = 1}^m \frac{\partial}{\partial \mu} \log \left( \frac{1}{\sqrt{2 \pi \sigma^2}} \right)
- \sum_{i = 1}^m \frac{\partial}{\partial \mu} \left( \frac{(x_i - \mu)^2}{2 \sigma^2} \right) \\
& = 0 - \sum_{i = 1}^m \left(\frac{2 (x_i - \mu)}{2 \sigma^2} \right)
\frac{\partial}{\partial \mu} (x_i - \mu) \\
& = \frac{1}{\sigma^2} \sum_{i = 1}^m (x_i - \mu) \\
& = \frac{1}{\sigma^2} \left( \sum_{i = 1}^m x_i - m \mu \right) \\
\therefore \frac{\partial^2}{\partial \mu^2} \log L(\mu, \sigma^2 \ | \ x)
& = - \frac{1}{\sigma^2} m < 0 \ \forall \ \mu \in \mathbb{R}, \sigma^2 \in \mathbb{R}^+
\text{, implying concavity} \\
\Longrightarrow \text{For maximum log-likelihood estimation,} \\
\frac{\partial}{\partial \mu} \log L(\mu, \sigma^2 \ | \ x) & = 0 \\
\Longrightarrow \frac{1}{\sigma^2} \left( \sum_{i = 1}^m x_i - m \mu \right) & = 0 \\
\Longrightarrow m \mu & = \sum_{i = 1}^m x_i \\
\Longrightarrow \mu & = \frac{1}{m} \sum_{i = 1}^m x_i \\
& = \bar{x}
\end{aligned}
$$
Therefore, for maximum likelihood estimation, **mean**, $\mu$, of the estimated normal distribution should be equal to the **sample mean**.
Differentiating w.r.t. $\sigma$, we get:
$$
\begin{aligned}
\frac{\partial}{\partial \sigma} \log L(\mu, \sigma^2 \ | \ x)
& = \sum_{i = 1}^m \frac{\partial}{\partial \sigma}
\log \left( \frac{1}{\sqrt{2 \pi \sigma^2}} \right)
- \sum_{i = 1}^m \frac{\partial}{\partial \sigma} \left( \frac{(x_i - \mu)^2}{2 \sigma^2} \right) \\
& = \sum_{i = 1}^m \left[
\frac{\partial}{\partial \sigma} \log \left( \frac{1}{\sqrt{2 \pi}} \right)
- \frac{\partial}{\partial \sigma} \log (\sigma) \right]
+ \sum_{i = 1}^m \left( \frac{(x_i - \mu)^2}{\sigma^3} \right) \\
& = - \frac{1}{\sigma} \sum_{i = 1}^m 1
+ \frac{1}{\sigma^3} \sum_{i = 1}^m (x_i - \mu)^2 \\
& = - \frac{m}{\sigma} + \frac{1}{\sigma^3} \sum_{i = 1}^m (x_i - \mu)^2 \\
\therefore \frac{\partial^2}{\partial \sigma^2} \log L(\mu, \sigma^2 \ | \ x)
& = \frac{m}{\sigma^2} - \frac{3}{\sigma^4} \sum_{i = 1}^m (x_i - \mu)^2 \\
& = \frac{1}{\sigma^2} \left[m - \frac{3}{\sigma^2} \sum_{i = 1}^m (x_i - \mu)^2 \right]
\end{aligned}
$$
Since the second partial derivative of our log-likelihood function w.r.t. $\sigma$ is continuous, $\forall \ \sigma \in \mathbb{R}^+$, it means that wherever the log-likelihood function reaches an extremum, the first partial derivative w.r.t. $\sigma$ should be $0$.
$$
\begin{aligned}
\frac{\partial}{\partial \sigma} \log L(\mu, \sigma^2 \ | \ x) & = 0 \\
\Longrightarrow - \frac{m}{\sigma} + \frac{1}{\sigma^3} \sum_{i = 1}^m (x_i - \mu)^2 & = 0 \\
\Longrightarrow \frac{1}{\sigma} \left[ - m + \frac{1}{\sigma^2} \sum_{i = 1}^m (x_i - \mu)^2 \right]
& = 0 \\
\Longrightarrow \sigma & \rightarrow +\infty \\
\text{or} \\
\sigma^2 & = \frac{1}{m} \sum_{i = 1}^m (x_i - \mu)^2
\end{aligned}
$$
Now,
$$
\begin{aligned}
\lim_{\sigma \to +\infty} \left(m - \frac{3}{\sigma^2}
\sum_{i = 1}^m (x_i - \mu)^2 \right) & > 0 \ \forall \ m \in \mathbb{Z}^+ \\
\Longrightarrow \lim_{\sigma \to +\infty}
\frac{1}{\sigma^2} \left(m - \frac{3}{\sigma^2} \sum_{i = 1}^m (x_i - \mu)^2 \right)
& \to 0^+ \ \forall \ m \in \mathbb{Z}^+ \\
\end{aligned}
$$
This means that when $\sigma \to +\infty$, log-likelihood function attains minimum value.
When
$$
\begin{aligned}
\sigma^2 & = \frac{1}{m} \sum_{i = 1}^m (x_i - \mu)^2, \\
\left(m - \frac{3}{\sigma^2} \sum_{i = 1}^m (x_i - \mu)^2 \right)
& = - 2 m \\
\Longrightarrow
\frac{1}{\sigma^2} \left(m - \frac{3}{\sigma^2} \sum_{i = 1}^m (x_i - \mu)^2 \right)
& < 0 \ \forall \ m \in \mathbb{Z}^+ \\
\end{aligned}
$$
This means that this is a point of maximum for the log-likelihood function.
Therefore, even though log-likelihood function in this case is not concave w.r.t. $\sigma$, we still see that the function gets maximised when $\frac{\partial}{\partial \sigma} \log L(\mu, \sigma^2 \ | \ x) = 0$, with $\sigma$ being finite.
Therefore, for **maximisation of likelihood** for **normal distribution**, **mean parameter**, $\mu$, should be equal to **sample mean** and the **variance**, $\sigma^2$, should be equal to **sample variance (biased)**.
Let's see an example of maximum likelihood estimation for normally distributed data. For this example, we use **Housefly Wing Lengths** (Sokal & Rohlf, 1971) data, which as the name suggests, contains information for wing lengths of 100 houseflies.
```
# import packages
import numpy as np
import matplotlib.pyplot as plt
import scipy.stats
# read and store data
wing_lengths = [] # create an empty list
# open the file containing the data
text_file = open('../data/Housefly_wing_lengths.txt')
# store data
for line in text_file:
wing_lengths.append(int(line)) # append datum into list
# close the file
text_file.close()
# convert list to numpy array
wing_lengths = np.array(wing_lengths)
print(wing_lengths)
# estimate parameters
# estimate mean
mu = np.mean(wing_lengths) # calculate sample mean
# estimate variance
sigma_square = np.mean((wing_lengths - mu) ** 2)
print(f'Estimated mean = {mu}, variance = {sigma_square}')
# Plot the histogram as well as the estimated distribution
# plot the histogram
plt.hist(wing_lengths, bins=(wing_lengths.max() - wing_lengths.min() + 1),
range=(wing_lengths.min(), wing_lengths.max() + 1),
label='Histogram for the data')
# plot the distribution
# find the pdf values
x_s = np.array(range(wing_lengths.min(), wing_lengths.max() + 1))
y_s = np.exp(-((x_s - mu) ** 2) / (2 * sigma_square)) / np.sqrt(2 * np.pi * sigma_square)
# scale the pmf values
y_s = y_s / y_s.max() * scipy.stats.mode(wing_lengths).count[0]
# distribution shifted by 0.5 so that values can be matched with histogram bars' centre
plt.plot(x_s + 0.5, y_s, label='Estimated Normal Distribution (Scaled)')
# customize plot
plt.title('Sample Histogram and Estimated Distribution')
plt.xlabel('Wing length (mm)')
plt.ylabel('Counts')
plt.legend(bbox_to_anchor=(1.05, 1), loc='best')
plt.show()
```
We observe that our estimated distribution fits very well to the data.
---
# <a id="Summary">Summary</a>
- **Maximum likelihood estimation** is the process of **determining parameters** of an **assumed distribution** for the **given sample**, such that the **likelihood** of the distribution to be defining the sample gets **maximised**.
- **Likelihood function**, $L:[\theta, x] \longrightarrow \mathbb{R}^+_0$ is defined as $L(\theta \ | \ x) = \prod_{i = 1}^n f(x_i | \theta)$, where $f$ is the **probability density/mass function**, $\theta$ is the vector of $k$ **distribution parameters**, $x$ is the vector of $n$ **sample points**.
- We usually **maximise** the **log-likelihood functions** since calculating derivatives is easier for them. Since logarithm function is a monotonically strictly increasing function, the parameter values where log-likelihood function is maximised are same as that for maximisation of likelihood functions.
- For MLE in case of **concave likelihood functions** (w.r.t. $\theta_j$), $\theta_j = \text{arg} \left(\sum_{i = 1}^n \frac{1}{f(x_i \ | \ \theta)} \frac{\partial}{\partial \theta_j} f(x_i \ | \ \theta) = 0\right), \ \forall \ j \in \{1, 2, 3, \ ..., \ k\}$
- **Likelihood** is **different** from **probability**. In probabiltiy, parameters are given and we determine possibility of getting certain outcomes. In **likelihood** calculation, **outcomes are given** by default and we determine the likeliness of the sample to have come from a probability distribution with some specific parameters (**find which parameters will make the distribution fit best on the sample**).
- MLE for **Bernoulli Distribution**, Bernoulli$(p)$, $\Longrightarrow p = \bar{x}$. That is, **probability of success**, $p$ equals **sample mean**, $\bar{x}$.
- MLE for **Binomial Distribution**, Bin$(n, p)$, $\Longrightarrow p = \frac{\bar{x}}{n}$, where, $\bar{x} = \frac{1}{m} \sum_{i = 1}^m x_i$, $m$ is the number of sample points. Here, $n$ is the number of Bernoulli trials conducted for each sample point, $p$ is the probability of success of a Bernoulli trial, and $x_i$ is the number of successes in $i^{th}$ sample point.
- MLE for **Poisson Distribution**, Poisson$(\lambda)$, $\Longrightarrow \lambda = \bar{x}$. This means that **average rate**, $\lambda$, should be equal to the **sample mean**, $\bar{x}$.
- MLE for **Exponential Distribution**, Exp$(\lambda)$, $\Longrightarrow \lambda = \frac{1}{\bar{x}}$. This means that **mean rate**, $\lambda$, should be equal to the **reciprocal** of the **sample mean**, $\bar{x}$.
- MLE for **Normal Distribution**, $N(\mu, \sigma^2)$, $\Longrightarrow \mu = \bar{x}$ and $\sigma^2 = \frac{1}{m} \sum_{i = 1}^m (x_i - \bar{x})^2$. This means that the **distribution mean**, $\mu$, should be equal to the **sample mean**, $\bar{x}$, and the **distribution variance**, $\sigma^2$, should be equal to the **sample variance (biased)**.
---
# <a id="References">References</a>
1. Andrews D. F., & Herzberg A. M. (1985). *Prussian Horse-Kick Data* \[Data set\]. A Collection of Problems from Many Fields for the Student and Research Worker. http://www.randomservices.org/random/data/HorseKicks.html
2. Brooks-Bartlett J. (2018). *Probability concepts explained: Maximum likelihood estimation*. Towards Data Science. https://towardsdatascience.com/probability-concepts-explained-maximum-likelihood-estimation-c7b4342fdbb1
3. Maximum Likelihood Estimation. (2020). In _Wikipedia_. https://en.wikipedia.org/wiki/Maximum_likelihood_estimation
4. Modarres M et al. (2017). *Standard Datasets: Mileage* \[Data set\]. Reliability Engineering and Risk Analysis: A Practical Guide (3rd ed.). https://reliability.readthedocs.io/en/latest/Datasets.html
5. Piech C. (2018). *Maximum Likelihood Estimation*. CS109: Probability for Computer Scientists. Stanford University. http://web.stanford.edu/class/archive/cs/cs109/cs109.1192/lectureNotes/21%20-%20MLE.pdf
6. Sokal R. R., & Rohlf F. J. (1971). Housefly Wing Lengths \[Data set\]. Biometry, The Principles and Practice of Statistics in Biological Research. *International Review of Hydrobiology 52*(2), 328-329. https://doi.org/10.1002/iroh.19710560218 https://seattlecentral.edu/qelp/sets/057/057.html
7. StatQuest with Josh Starmer. (2018). *Maximum Likelihood for the Binomial Distribution, Clearly Explained!!!* \[Video\]. YouTube. https://youtu.be/4KKV9yZCoM4
8. StatQuest with Josh Starmer. (2018). *Maximum Likelihood for the Exponential Distribution, Clearly Explained! V2.0* \[Video\]. YouTube. https://youtu.be/p3T-_LMrvBc
9. StatQuest with Josh Starmer. (2018). *Maximum Likelihood For the Normal Distribution, step-by-step!* \[Video\]. YouTube. https://youtu.be/Dn6b9fCIUpM
10. StatQuest with Josh Starmer. (2017). *StatQuest: Maximum Likelihood, clearly explained!!!* \[Video\]. YouTube. https://youtu.be/XepXtl9YKwc
11. StatQuest with Josh Starmer. (2018). *StatQuest: Probability vs Likelihood* \[Video\]. YouTube. https://youtu.be/pYxNSUDSFH4
12. The Pennsylvania State University. Eberly College of Science. (2020). *1.2 - Maximum Likelihood Estimation*. STAT 414 | Introduction to Mathematical Statistics. https://online.stat.psu.edu/stat415/lesson/1/1.2
13. The Pennsylvania State University. Eberly College of Science. (2020). *1.5 - Maximum-likelihood (ML) Estimation*. STAT 504 | Analysis of Discrete Data. https://online.stat.psu.edu/stat504/node/28/
---
| github_jupyter |
```
import arviz as az
import matplotlib.pyplot as plt
import numpy as np
import pymc3 as pm
%load_ext watermark
az.style.use('arviz-darkgrid')
```
# Model comparison
To demonstrate the use of model comparison criteria in PyMC3, we implement the **8 schools** example from Section 5.5 of Gelman et al (2003), which attempts to infer the effects of coaching on SAT scores of students from 8 schools. Below, we fit a **pooled model**, which assumes a single fixed effect across all schools, and a **hierarchical model** that allows for a random effect that partially pools the data.
The data include the observed treatment effects and associated standard deviations in the 8 schools.
```
J = 8
y = np.array([28, 8, -3, 7, -1, 1, 18, 12])
sigma = np.array([15, 10, 16, 11, 9, 11, 10, 18])
```
### Pooled model
```
with pm.Model() as pooled:
mu = pm.Normal('mu', 0, sigma=1e6)
obs = pm.Normal('obs', mu, sigma=sigma, observed=y)
trace_p = pm.sample(2000, return_inferencedata=True)
az.plot_trace(trace_p);
```
### Hierarchical model
```
with pm.Model() as hierarchical:
eta = pm.Normal('eta', 0, 1, shape=J)
mu = pm.Normal('mu', 0, sigma=10)
tau = pm.HalfNormal('tau', 10)
theta = pm.Deterministic('theta', mu + tau*eta)
obs = pm.Normal('obs', theta, sigma=sigma, observed=y)
trace_h = pm.sample(2000, target_accept=0.9, return_inferencedata=True)
az.plot_trace(trace_h, var_names='mu');
az.plot_forest(trace_h, var_names='theta');
```
### Leave-one-out Cross-validation (LOO)
LOO cross-validation is an estimate of the out-of-sample predictive fit. In cross-validation, the data are repeatedly partitioned into training and holdout sets, iteratively fitting the model with the former and evaluating the fit with the holdout data. Vehtari et al. (2016) introduced an efficient computation of LOO from MCMC samples (without the need for re-fitting the data). This approximation is based on importance sampling. The importance weights are stabilized using a method known as Pareto-smoothed importance sampling (PSIS).
### Widely-applicable Information Criterion (WAIC)
WAIC (Watanabe 2010) is a fully Bayesian criterion for estimating out-of-sample expectation, using the computed log pointwise posterior predictive density (LPPD) and correcting for the effective number of parameters to adjust for overfitting.
By default ArviZ uses LOO, but WAIC is also available.
```
pooled_loo = az.loo(trace_p, pooled)
pooled_loo.loo
hierarchical_loo = az.loo(trace_h, hierarchical)
hierarchical_loo.loo
```
ArviZ includes two convenience functions to help compare LOO for different models. The first of these functions is `compare`, which computes LOO (or WAIC) from a set of traces and models and returns a DataFrame.
```
df_comp_loo = az.compare({"hierarchical": trace_h, "pooled": trace_p})
df_comp_loo
```
We have many columns, so let's check out their meaning one by one:
0. The index is the names of the models taken from the keys of the dictionary passed to `compare(.)`.
1. **rank**, the ranking of the models starting from 0 (best model) to the number of models.
2. **loo**, the values of LOO (or WAIC). The DataFrame is always sorted from best LOO/WAIC to worst.
3. **p_loo**, the value of the penalization term. We can roughly think of this value as the estimated effective number of parameters (but do not take that too seriously).
4. **d_loo**, the relative difference between the value of LOO/WAIC for the top-ranked model and the value of LOO/WAIC for each model. For this reason we will always get a value of 0 for the first model.
5. **weight**, the weights assigned to each model. These weights can be loosely interpreted as the probability of each model being true (among the compared models) given the data.
6. **se**, the standard error for the LOO/WAIC computations. The standard error can be useful to assess the uncertainty of the LOO/WAIC estimates. By default these errors are computed using stacking.
7. **dse**, the standard errors of the difference between two values of LOO/WAIC. The same way that we can compute the standard error for each value of LOO/WAIC, we can compute the standard error of the differences between two values of LOO/WAIC. Notice that both quantities are not necessarily the same, the reason is that the uncertainty about LOO/WAIC is correlated between models. This quantity is always 0 for the top-ranked model.
8. **warning**, If `True` the computation of LOO/WAIC may not be reliable.
9. **loo_scale**, the scale of the reported values. The default is the log scale as previously mentioned. Other options are deviance -- this is the log-score multiplied by -2 (this reverts the order: a lower LOO/WAIC will be better) -- and negative-log -- this is the log-score multiplied by -1 (as with the deviance scale, a lower value is better).
The second convenience function takes the output of `compare` and produces a summary plot in the style of the one used in the book [Statistical Rethinking](http://xcelab.net/rm/statistical-rethinking/) by Richard McElreath (check also [this port](https://github.com/aloctavodia/Statistical-Rethinking-with-Python-and-PyMC3) of the examples in the book to PyMC3).
```
az.plot_compare(df_comp_loo, insample_dev=False);
```
The empty circle represents the values of LOO and the black error bars associated with them are the values of the standard deviation of LOO.
The value of the highest LOO, i.e the best estimated model, is also indicated with a vertical dashed grey line to ease comparison with other LOO values.
For all models except the top-ranked one we also get a triangle indicating the value of the difference of WAIC between that model and the top model and a grey errobar indicating the standard error of the differences between the top-ranked WAIC and WAIC for each model.
### Interpretation
Though we might expect the hierarchical model to outperform a complete pooling model, there is little to choose between the models in this case, given that both models gives very similar values of the information criteria. This is more clearly appreciated when we take into account the uncertainty (in terms of standard errors) of LOO and WAIC.
## Reference
[Gelman, A., Hwang, J., & Vehtari, A. (2014). Understanding predictive information criteria for Bayesian models. Statistics and Computing, 24(6), 997–1016.](https://doi.org/10.1007/s11222-013-9416-2)
[Vehtari, A, Gelman, A, Gabry, J. (2016). Practical Bayesian model evaluation using leave-one-out cross-validation and WAIC. Statistics and Computing](http://link.springer.com/article/10.1007/s11222-016-9696-4)
```
%watermark -n -u -v -iv -w
```
| github_jupyter |
```
import pandas as pd
import numpy as np
from functions import *
import random
import csv
from sklearn.linear_model import LogisticRegression
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.tree import ExtraTreeClassifier
from sklearn.neural_network import MLPClassifier
from sklearn.neighbors import RadiusNeighborsClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.linear_model import SGDClassifier
from sklearn.gaussian_process import GaussianProcessClassifier
from sklearn.ensemble import AdaBoostClassifier
from sklearn.ensemble import BaggingClassifier
from sklearn.naive_bayes import BernoulliNB
from sklearn.calibration import CalibratedClassifierCV
from sklearn.svm import LinearSVC
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.linear_model import LogisticRegressionCV
from sklearn.discriminant_analysis import QuadraticDiscriminantAnalysis
#from sklearn.mixture import DPGMM
remove_fight_island = False
#Turn off warnings
import warnings
warnings.filterwarnings("ignore")
#Load models
#REMINDER: We are going to need to use 'eval' to get the models usable
with open('../data/models.csv', newline='') as f:
reader = csv.reader(f)
models = list(reader)
print(len(models))
###SELECT MODEL TO OPTIMIZE
#model_num = 19
models
df = pd.read_csv("../data/kaggle_data/ufc-master.csv")
#Let's put all the labels in a dataframe
df['label'] = ''
#If the winner is not Red or Blue we can remove it.
mask = df['Winner'] == 'Red'
df['label'][mask] = 0
mask = df['Winner'] == 'Blue'
df['label'][mask] = 1
#df["Winner"] = df["Winner"].astype('category')
df = df[(df['Winner'] == 'Blue') | (df['Winner'] == 'Red')]
#Make sure lable is numeric
df['label'] = pd.to_numeric(df['label'], errors='coerce')
#Let's fix the date
df['date'] = pd.to_datetime(df['date'])
#Create a label df:
label_df = df['label']
#Let's create an odds df too:
odds_df = df[['R_odds', 'B_odds']]
#Split the test set. We are always(?) going to use the last 200 matches as the test set, so we don't want those around
#as we pick models
df_train = df[250:]
odds_train = odds_df[250:]
label_train = label_df[250:]
df_test = df[:250]
odds_test = odds_df[:250]
label_test = label_df[:250]
print(len(df_test))
print(len(odds_test))
print(len(label_test))
print(len(df_train))
print(len(odds_train))
print(len(label_train))
if remove_fight_island:
##Let's remove the Fight island contests and see how that affects score
df_test_no_fight_island = df_test[(df_test['location'] != 'Abu Dhabi, Abu Dhabi, United Arab Emirates')]
df_train_no_fight_island = df_train[(df_train['location'] != 'Abu Dhabi, Abu Dhabi, United Arab Emirates')]
df_test = df_test_no_fight_island
df_train = df_train_no_fight_island
print(len(df_test))
print(len(df_train))
display(df_train)
display(df_test)
#Set a value for the nulls in the ranks
weightclass_list = ['B_match_weightclass_rank', 'R_match_weightclass_rank', "R_Women's Flyweight_rank", "R_Women's Featherweight_rank", "R_Women's Strawweight_rank", "R_Women's Bantamweight_rank", 'R_Heavyweight_rank', 'R_Light Heavyweight_rank', 'R_Middleweight_rank', 'R_Welterweight_rank', 'R_Lightweight_rank', 'R_Featherweight_rank', 'R_Bantamweight_rank', 'R_Flyweight_rank', 'R_Pound-for-Pound_rank', "B_Women's Flyweight_rank", "B_Women's Featherweight_rank", "B_Women's Strawweight_rank", "B_Women's Bantamweight_rank", 'B_Heavyweight_rank', 'B_Light Heavyweight_rank', 'B_Middleweight_rank', 'B_Welterweight_rank', 'B_Lightweight_rank', 'B_Featherweight_rank', 'B_Bantamweight_rank', 'B_Flyweight_rank', 'B_Pound-for-Pound_rank']
df_train[weightclass_list] = df_train[weightclass_list].fillna(17)
df_test[weightclass_list] = df_test[weightclass_list].fillna(17)
df_test
#df_test.to_csv('test.csv')
#1. Set features
#2. Set Hyperparameters
#3. Set EV
#4. Remove Features
models[1][model_num]
test_model_name = models[0][model_num]
test_model = eval(models[1][model_num])
test_model_features = eval(models[2][model_num])
test_model_ev = eval(models[3][model_num])
old_test_model = test_model
old_test_model_features = test_model_features
old_test_model_ev = test_model_ev
print(df_test.columns.tolist()) #Keeps us from truncating
#1. set features
my_pos_features = ['R_odds', 'B_odds', 'R_ev', 'B_ev',
'location', 'country', 'title_bout', 'weight_class', 'gender',
'no_of_rounds', 'B_current_lose_streak', 'B_current_win_streak',
'B_draw', 'B_avg_SIG_STR_landed', 'B_avg_SIG_STR_pct', 'B_avg_SUB_ATT',
'B_avg_TD_landed', 'B_avg_TD_pct', 'B_longest_win_streak', 'B_losses',
'B_total_rounds_fought', 'B_total_title_bouts',
'B_win_by_Decision_Majority', 'B_win_by_Decision_Split',
'B_win_by_Decision_Unanimous', 'B_win_by_KO/TKO', 'B_win_by_Submission',
'B_win_by_TKO_Doctor_Stoppage', 'B_wins', 'B_Stance', 'B_Height_cms',
'B_Reach_cms', 'B_Weight_lbs', 'R_current_lose_streak',
'R_current_win_streak', 'R_draw', 'R_avg_SIG_STR_landed',
'R_avg_SIG_STR_pct', 'R_avg_SUB_ATT', 'R_avg_TD_landed', 'R_avg_TD_pct',
'R_longest_win_streak', 'R_losses', 'R_total_rounds_fought',
'R_total_title_bouts', 'R_win_by_Decision_Majority',
'R_win_by_Decision_Split', 'R_win_by_Decision_Unanimous',
'R_win_by_KO/TKO', 'R_win_by_Submission',
'R_win_by_TKO_Doctor_Stoppage', 'R_wins', 'R_Stance', 'R_Height_cms',
'R_Reach_cms', 'R_Weight_lbs', 'R_age', 'B_age', 'lose_streak_dif',
'win_streak_dif', 'longest_win_streak_dif', 'win_dif', 'loss_dif',
'total_round_dif', 'total_title_bout_dif', 'ko_dif', 'sub_dif',
'height_dif', 'reach_dif', 'age_dif', 'sig_str_dif', 'avg_sub_att_dif',
'avg_td_dif', 'empty_arena', 'B_match_weightclass_rank', 'R_match_weightclass_rank',
"R_Women's Flyweight_rank", "R_Women's Featherweight_rank", "R_Women's Strawweight_rank",
"R_Women's Bantamweight_rank", 'R_Heavyweight_rank', 'R_Light Heavyweight_rank',
'R_Middleweight_rank', 'R_Welterweight_rank', 'R_Lightweight_rank', 'R_Featherweight_rank',
'R_Bantamweight_rank', 'R_Flyweight_rank', 'R_Pound-for-Pound_rank', "B_Women's Flyweight_rank",
"B_Women's Featherweight_rank", "B_Women's Strawweight_rank", "B_Women's Bantamweight_rank",
'B_Heavyweight_rank', 'B_Light Heavyweight_rank', 'B_Middleweight_rank', 'B_Welterweight_rank',
'B_Lightweight_rank', 'B_Featherweight_rank', 'B_Bantamweight_rank', 'B_Flyweight_rank',
'B_Pound-for-Pound_rank', 'better_rank']
print(test_model_name)
print(test_model)
print(test_model_features)
print(test_model_ev)
def save_model():
score = evaluate_model(test_model, test_model_features, test_model_ev, df_train, label_train, odds_train, df_test, label_test,
odds_test, verbose = True)
models[0][model_num] = test_model_name
models[1][model_num] = test_model
models[2][model_num] = test_model_features
models[3][model_num] = test_model_ev
models[4][model_num] = score
with open('../data/models.csv', 'w', newline='') as outfile:
writer = csv.writer(outfile)
for row in models:
print("HI")
writer.writerow(row)
outfile.close()
def print_model():
print()
print(test_model_name)
print(test_model)
print(test_model_features)
print(test_model_ev)
print()
keep_going = True
#keep_going = False
while(keep_going):
#1. Set Features
#get_best_features(pos_features, m, df, cur_features, labels, odds, scale=False)
test_model_features = (get_best_features(my_pos_features, test_model, df_train, test_model_features, label_train, odds_train,
min_ev=test_model_ev))
print_model()
save_model()
#2 Set hyperparameters
#def tune_hyperparameters(input_model, input_features, input_df, input_labels, odds_input):
test_model = tune_hyperparameters(test_model, test_model_features, df_train, label_train, odds_train,
min_ev=test_model_ev)
print_model()
save_model()
#3. Set EV
#def tune_ev(input_model, input_features, input_df, input_labels, odds_input, verbose=False):
test_model_ev = tune_ev(test_model, test_model_features, df_train, label_train, odds_train, verbose=False)
old_test_model_features = test_model_features #This prevents
#an uneccesary loop
print_model()
save_model()
#4. Remove Features
#def remove_to_improve(cur_features, m, df, labels, odds, scale=False, min_ev = 0):
test_model_features = remove_to_improve(test_model_features, test_model, df_train, label_train, odds_train, min_ev = test_model_ev)
keep_going = False
if old_test_model != test_model:
print("The hyperparameters are different")
print("OLD:")
print(old_test_model)
print("NEW:")
print(test_model)
keep_going = True
old_test_model = test_model
if old_test_model_features != test_model_features:
print("The features are different")
print("OLD:")
print(old_test_model_features)
print("NEW:")
print(test_model_features)
keep_going = True
old_test_model_features = test_model_features
if old_test_model_ev != test_model_ev:
print("The EV is different")
print("OLD:")
print(old_test_model_ev)
print("NEW:")
print(test_model_ev)
keep_going = True
old_test_model_ev = test_model_ev
print_model()
save_model()
#Get new score:
#def evaluate_model(input_model, input_features, input_ev, train_df, train_labels, train_odds, test_df, test_labels, test_odds, verbose=True):
score = evaluate_model(test_model, test_model_features, test_model_ev, df_train, label_train, odds_train, df_test, label_test,
odds_test, verbose = True)
```
print(test_model_name)
print(models[0][model_num])
print()
print(test_model)
print(eval(models[1][model_num]))
print()
print(test_model_features)
print(eval(models[2][model_num]))
print()
print(test_model_ev)
print(eval(models[3][model_num]))
```
models[0][model_num] = test_model_name
models[1][model_num] = test_model
models[2][model_num] = test_model_features
models[3][model_num] = test_model_ev
models[4][model_num] = score
with open('../data/models.csv', 'w', newline='') as outfile:
writer = csv.writer(outfile)
for row in models:
print("HI")
writer.writerow(row)
outfile.close()
with open('../data/models.csv', newline='') as f:
reader = csv.reader(f)
data = list(reader)
print(len(data))
data
```
| github_jupyter |
## Table of Contents
* [Introduction](#Introduction)
* [Common Model Inputs](#Common-Model-Inputs)
* [Model-Independent Inputs](#Model-Independent-Inputs)
* [Storage Object](#Storage-Object)
* [Forward Curve](#Forward-Curve)
* [Interest Rates Curve](#Interest-Rates-Curve)
* [Settlement Rule](#Settlement-Rule)
* [Valuation Date, Inventory and Discount Deltas](#Valuation-Date,-Inventory-and-Discount-Deltas)
* [Least Square Monte Carlo Common Inputs](#Least-Square-Monte-Carlo-Common-Inputs)
* [Three-Factor Seasonal Model](#Three-Factor-Seasonal-Model)
* [Three-Factor Seasonal Model Specific Inputs](#Three-Factor-Seasonal-Model-Specific-Inputs)
* [Three-Factor Price Dynamics Parameters](#Three-Factor-Price-Dynamics-Parameters)
* [Three-Factor Basis Functions](#Three-Factor-Basis-Functions)
* [Calling Three Factor Model](#Calling-Three-Factor-Seasonal-Model)
* [General Multi-Factor Model](#General-Multi-Factor-Model)
* [Multi-Factor Model Specific Inputs](#Multi-Factor-Model-Specific-Inputs)
* [Multi-Factor Price Dynamics Parameters](#Multi-Factor-Price-Dynamics-Parameters)
* [Factor Volatility Curves and Mean Reversions](#Factor-Volatility-Curves-and-Mean-Reversions)
* [Factor Correlations](#Factor-Correlations)
* [Multi-Factor Basis Functions](#Multi-Factor-Basis-Functions)
* [Calling the Multi Factor Model](#Calling-the-Multi-Factor-Model)
* [Valuation Results](#Valuation-Results)
* [NPV Properties](#NPV-Properties)
* [Deltas](#Deltas)
* [Intrinsic Profile](#Intrinsic-Profile)
* [Trigger Prices](#Trigger-Prices)
* [Simulated Values](#Simulated-Values)
* [Expected Profile](#Expected-Profile)
* [Instrumentation](#Instrumentation)
* [Progress Callback](#Progress-Callback)
* [Logging](#Logging)
* [References](#References)
***
## Introduction
This notebook describes how to use the LSMC (Least Squares Monte Carlo) energy storage valuation models in the Python package [cmdty-storage](https://pypi.org/project/cmdty-storage/). As a prerequisite study the notebook [creating_storage_instances](creating_storage_instances.ipynb) which describes how to create instances of the CmdtyStorage class used to represent the characteristics of the storage facility.
The cmdty-storage package contains two functions to value storage, each with different assumptions about the price dynamics:
* The function **multi_factor_value** values assuming a very general multi-factor model which can be called with an arbitrary number of factors..
* The functions **three_factor_seasonal_value** is a specific case of the more general multi-factor model which reduces the number of parameters to a minimum level which appropriately describe the seasonality of energy prices.
As there are a large number of complex inputs to both functions, this notebook first describes the inputs, starting with those common between both functions. Next details of how to call the functions is provided, followed by details of the results generated by both models. The final section describe intrumentation of the model usage in order to hook into calculation progress and tracing information.
```
# Common Imports
from cmdty_storage import CmdtyStorage, three_factor_seasonal_value, multi_factor_value, RatchetInterp
import pandas as pd
import numpy as np
import ipywidgets as ipw
from IPython.display import display
```
***
## Common Model Inputs
This section describes the inputs which are common between three factor seasonal valuation model and the multi-factor model.
### Model-Independent Inputs
The following inputs would always be necessary for any methodology for valuing storage, irrespective of what model is used.
#### Storage Object
For all examples the following storage with ratchets is used. See the notebook [creating_storage_instances](creating_storage_instances.ipynb) which describes the various other ways of creating CmdtyStorage instances depending on the characteristics of the storage facility being modelled.
```
storage = CmdtyStorage(
freq='D',
storage_start = '2021-04-01',
storage_end = '2022-04-01',
injection_cost = 0.1,
withdrawal_cost = 0.25,
ratchets = [
('2021-04-01', # For days after 2021-04-01 (inclusive) until 2022-10-01 (exclusive):
[
(0.0, -150.0, 250.0), # At min inventory of zero, max withdrawal of 150, max injection 250
(2000.0, -200.0, 175.0), # At inventory of 2000, max withdrawal of 200, max injection 175
(5000.0, -260.0, 155.0), # At inventory of 5000, max withdrawal of 260, max injection 155
(7000.0, -275.0, 132.0), # At max inventory of 7000, max withdrawal of 275, max injection 132
]),
('2022-10-01', # For days after 2022-10-01 (inclusive):
[
(0.0, -130.0, 260.0), # At min inventory of zero, max withdrawal of 130, max injection 260
(2000.0, -190.0, 190.0), # At inventory of 2000, max withdrawal of 190, max injection 190
(5000.0, -230.0, 165.0), # At inventory of 5000, max withdrawal of 230, max injection 165
(7000.0, -245.0, 148.0), # At max inventory of 7000, max withdrawal of 245, max injection 148
]),
],
ratchet_interp = RatchetInterp.LINEAR
)
```
#### Forward Curve
The forward curve used for valuation is specified as an instance of pandas.Series, with index of type PeriodIndex with granularity consistent with the frequency string **freq** used to create the CmdtyStorage instance.
The following natural gas forward curve is used in all examples. This is a monthly curve, converted to daily granularity with piecewise flat interpolation. To create smooth daily curves, with day-of-week seasonality see [Cmdty Curves](https://github.com/cmdty/curves).
```
monthly_index = pd.period_range(start='2021-04-25', periods=25, freq='M')
monthly_fwd_prices = [16.61, 15.68, 15.42, 15.31, 15.27, 15.13, 15.96, 17.22, 17.32, 17.66,
17.59, 16.81, 15.36, 14.49, 14.28, 14.25, 14.32, 14.33, 15.30, 16.58,
16.64, 16.79, 16.64, 15.90, 14.63]
fwd_curve = pd.Series(data=monthly_fwd_prices, index=monthly_index).resample('D').fillna('pad')
%matplotlib inline
fwd_curve.plot(title='Forward Curve')
```
#### Interest Rates Curve
Act/365 continuously compounded interest rates should be specified as a Pandas Series instance with daily granularity. In these examples, for simplicity a linearly interpolated rates curve is used.
```
rates = [0.005, 0.006, 0.0072, 0.0087, 0.0101, 0.0115, 0.0126]
rates_pillars = pd.PeriodIndex(freq='D', data=['2021-04-25', '2021-06-01', '2021-08-01', '2021-12-01', '2022-04-01',
'2022-12-01', '2023-12-01'])
ir_curve = pd.Series(data=rates, index=rates_pillars).resample('D').asfreq('D').interpolate(method='linear')
ir_curve.plot(title='Interest Rate Curve')
```
#### Settlement Rule
This is a mapping from delivery period to the date on which cash settlement occurs. In a real-world usage scenario this would depend on the settlement rules for the forward market used to trade around the storage. This is specified as a function mapping from a pandas.Period instance (which represents the commodity delivery period) to the date on which this commodity delivery would be cash settle. The return instance needs to be date-like.
In the example below this is expressed as a function which maps to the 20th day of the following month.
```
def settlement_rule(delivery_date):
return delivery_date.asfreq('M').asfreq('D', 'end') + 20
```
#### Valuation Date, Inventory and Discount Deltas
* **val_date** is the current time period as of when the valuation is performed. In this example as string is used, but instances of date, datetime or pandas.Period are also admissible.
* **inventory** is the current volume of commodity held in storage.
* **discount_deltas** is a boolean flag indicating whether the deltas should be either:
* True: discounted and as such maintain the definition of a sensitivity of the NPV to the forward price as a partial derivative.
* False: in the case where the forward market is OTC and not margined, the undiscounted deltas calculated should be used as the negative of the forward hedge to maintain delta neutrality. This is because a single unit of non-marginned forward has a delta of one discounted.
```
val_date = '2021-04-25'
inventory = 1500.0
discount_deltas = True
```
### Least Square Monte Carlo Common Inputs
The following are inputs to both the Three-Factor Seasonal model and the Multi-Factor model.
* **num_sims** is the number of Monte Carlo simulations (or paths) used for the valuation. The higher this integer is, the more accurate the valuation will be, but the longer it will take to calculate.
* **seed** is an optional integer argument which is used as the seed to the Mersenne Twister random number generator. With all other inputs being equal, repeated runs of the valuation with the same seed will create identical simulated spot price paths. If the seed is omitted, a random seed will be used, and hence different spot prices will be generated even if all of the other inputs are identical. Specifying a seed is advisable in production usage as you will to be able to reproduce a valuation at any other time.
* **fwd_sim_seed** is another optional integer argument used if you want to specify the seed for the second forward spot price simulation. If omitted, unlike for when the seed argument is omitted, the second spot price will not use a random seed, rather it will use a continuation of the stream of random numbers generated by the first spot simulations, used for regressions.
```
num_sims = 500
seed = 12
fwd_sim_seed = 25
```
Both model functions include two other optional arguments which have quite a technical usage. As such, in most cases these should be omitted from the function calls, as they are in the example below, so that the default values are used.
* **num_inventory_grid_points**: during the backward induction, at each time step the inventory space is discretized into a grid. At these inventory points the Bellman Equation is used to calculate the optimal decisions. This argument determines the maximum number of points in this grid. The higher the integer specified, the more accurate the valuation will be, but the slower the calculation will run.
* **numerical_tolerance** is used as a tolerance at various points in the calculation to determine if two floating point numbers are considered equal.
***
## Three Factor Seasonal Model
This section gives an example of using the storage valuation model where the commodity price dynamics are assumed to follow a three-factor seasonal model. The stochastic price model is described in [this paper](https://github.com/cmdty/core/blob/master/docs/three_factor_seasonal_model/three_factor_seasonal_model.pdf), which is a special case of the more general [multi-factor model](https://github.com/cmdty/core/blob/master/docs/multifactor_price_process/multifactor_price_process.pdf). The three factor model is particularly suited power and natural gas which show seaonality in the volatility and correlation. TODO reference Kyos paper.
### Three Factor Seasonal Model Specific Inputs
The following inputs are specific to the three-factor seasonal model.
#### Three Factor Price Dynamics Parameters
There are four numeric arguments which determine the spot price dynamics. See the [the three-factor paper](https://github.com/cmdty/core/blob/master/docs/three_factor_seasonal_model/three_factor_seasonal_model.pdf) for a description and details of calibration.
```
spot_mean_reversion = 91.0
spot_vol = 0.85
long_term_vol = 0.30
seasonal_vol = 0.19
```
#### Three Factor Basis Functions
The basis functions form an integral part of the Least Squares Monte Carlo valuation technique. Each basis function represents a linear component of the parameterisation of the expected continuation value given the inventory after an inject/withdraw decision. See the [original paper](https://people.math.ethz.ch/~hjfurrer/teaching/LongstaffSchwartzAmericanOptionsLeastSquareMonteCarlo.pdf) by Longstaff and Schwartz for further details of the significance of basis functions in the valuation of financial derivatives.
The **basis_funcs** argument should be assigned to a string representing the basis functions. The format of this string should be the individual basis function deliminted by the '+' character. Any whitespace does not change the semantics of the string. Each individual basis function can consist of the following components:
* **1** for the contstant term. This should always be included on it's own, without multiplication with any other factors.
* **s** for the spot price.
* **x_st** for the short-term factor, the Brownian motion $z_{spot}(t)$.
* **x_lt** for the long-term factor, the Brownian motion $z_{long}(t)$.
* **x_sw** for the seasonal Summer-Winter factor, the Brownian motion $z_{seas}(t)$.
These can be augmented with the following operators:
* **\*** to multiply components, e.g. 's * x_st' is the spot price multiplied by the short-term factor.
* **\*\*** to raise to the power of, e.g. 'x_sw ** 2' is the Summer-Winter factor raised to the power of 2.
The example below includes basis functions for the first two powers of all four stochastic components, the product of the spot price and the short-term factor, and the constant term.
```
three_factor_basis_functions = '1 + x_st + x_sw + x_lt + s + x_st**2 + x_sw**2 + x_lt**2 + s**2 + s * x_st'
```
#### Calling Three Factor Seasonal Model
The three factor seasonal model can be run using the function **three_factor_seasonal_value**. This returns an instance of **MultiFactorValuationResults**, which has a property for the NPV, as printed in the example below. See the section [Valuation Results](#Valuation-Results) for more details on other information included in the results.
```
three_factor_results = three_factor_seasonal_value(
cmdty_storage = storage,
val_date = val_date,
inventory = inventory,
fwd_curve = fwd_curve,
interest_rates = ir_curve,
settlement_rule = settlement_rule,
discount_deltas=discount_deltas,
num_sims = num_sims,
seed = seed,
fwd_sim_seed = fwd_sim_seed,
spot_mean_reversion = spot_mean_reversion,
spot_vol = spot_vol,
long_term_vol = long_term_vol,
seasonal_vol = seasonal_vol,
basis_funcs = three_factor_basis_functions
)
'{0:,.0f}'.format(three_factor_results.npv)
```
***
## General Multi-Factor Model
This section gives an example of using the storage valuation model where the commodity price dynamics are assumed to follow the price dynamics described in [this paper](https://github.com/cmdty/core/blob/master/docs/multifactor_price_process/multifactor_price_process.pdf). This model of price dynamics can be seen as a general framework which can be used to construct more specific commodity models based to the characteristics of the commodity.
### Multi-Factor Model Specific Inputs
The following inputs are specific to the multi-factor model.
#### Multi Factor Price Dynamics Parameters
Multi-factor price dynamics parameters are demonstrated below using an example of a two-factor model with one long-term non-mean reverting factor, and one short-term factor. Such a model could be suitable to model commodities where there is no seasonality in volatility and there is strong correlation between different points on the forward curve, e.g. crude oil.
##### Factor Volatility Curves and Mean Reversions
The factors argument should be assigned to an iterable of tuples. The number of tuples equals the number of factors. Tuple $i$ should have two elements:
1. The $i^{th}$ factor mean reversion, i.e. $\alpha_i$ from the document, as a float
2. The $i^{th}$ factor volatility function, i.e. $\sigma_i(T)$, as a Pandas Series, with index of type PeriodIndex with frequency the same as that assigned to **freq** when creating the storage object.
The example code below creates the factors with constant factor volatilities.
```
# Parameters for a constant volatility 2-factor model
long_term_vol = 0.14
long_term_mean_reversion = 0.0
short_term_vol = 0.64
short_term_mean_reversion = 16.2
# Construct the factors argument of multi_factor_value function
factor_index = pd.period_range(start=val_date, end=storage.end, freq=storage.freq)
long_term_vol_curve = pd.Series(index=factor_index, dtype='float64')
long_term_vol_curve[:] = long_term_vol
short_term_vol_curve = pd.Series(index=factor_index, dtype='float64')
short_term_vol_curve[:] = short_term_vol
factors = [(short_term_mean_reversion, short_term_vol_curve),
(long_term_mean_reversion, short_term_vol_curve)]
```
##### Factor Correlations
The correlation between the different driving Brownian motion factors is specified as a matrix where element $i, j$ corresponds to $\rho_{i, j}$, the correlation between factors $i$ and $j$. For an n-factor model this should be an n by n 2-dimensional numpy ndarray. In the example below `factor_corrs_matrix` is created for this purpose.
However, in certain cases the factor correlations can be of different types:
* For a one-factor model factors can be `None` as factor correlations are not relevant.
* For a two-factor model a scalar of float type can be used. In the example below `factor_corrs` could be used without having to create the numpy array.
```
factor_corrs = 0.64
factor_corrs_matrix = np.array([[1.0, factor_corrs],
[factor_corrs, 1.0]])
```
##### Multi-Factor Basis Functions
The basis function string will be very similar to that described in [Three Factor Basis Functions](#Three-Factor-Basis-Functions) section above. The only difference is that instead of **x_st**, **x_lt** and **x_sw**, strings of the form **xi** should be used, where $i$ is the factor index, starting at 0 for the first factor. For example use **x0** for the first factor, **x1** for the second factor and so on. Hence **xi** should only be used with $i$ values from 0 to $n-1$ for an $n$-factor model.
In the example below `multi_factor_basis_funcs` is created to use the first two powers for all factors and the spot price, as well as all cross-products between factors and the spot price. Finally the constant term is included.
```
multi_factor_basis_funcs = '1 + x0 + x1 + s + x0**2 + x1**2 + s**2 + x0*x1 + x1*s + x0*s'
```
#### Calling the Multi Factor Model
The multi-factor model can be run using the function **multi_factor_value**. This returns an instance of **MultiFactorValuationResults**, which has a property for the NPV, as printed in the example below. See the section [Valuation Results](#Valuation-Results) for more details on other information included in the results.
```
multi_factor_results = multi_factor_value(
cmdty_storage = storage,
val_date = val_date,
inventory = inventory,
fwd_curve = fwd_curve,
interest_rates = ir_curve,
settlement_rule = settlement_rule,
discount_deltas=discount_deltas,
num_sims = num_sims,
seed = seed,
fwd_sim_seed = fwd_sim_seed,
factors = factors,
factor_corrs = factor_corrs_matrix,
basis_funcs = multi_factor_basis_funcs
)
'{0:,.0f}'.format(multi_factor_results.npv)
```
***
## Valuation Results
Both functions **three_factor_seasonal_value** and **multi_factor_value** return instances of **MultiFactorValuationResults**. This class derives from NamedTuple and has many properties for the results of the optimisation, plus other calculation metadata. The subsections below describe these properties.
### NPV Properties
The the following attributes give information on the NPV (Net Present Value):
* **npv** is the full NPV including the option value.
* **intrinsic_npv** is the NPV calculated using a static optimisation assuming prices are deterministic, i.e. giving the storage no option value.
* **extrinsic_npv** is the value of just the optionality, i.e. npv minus intrinsic_npv.
```
print('Full NPV:\t{0:,.0f}'.format(three_factor_results.npv))
print('Intrinsic NPV: \t{0:,.0f}'.format(three_factor_results.intrinsic_npv))
print('Extrinsic NPV: \t{0:,.0f}'.format(three_factor_results.extrinsic_npv))
```
### Deltas
The **deltas** property is of type pandas.Series and contains the senstivity of the full NPV to the forward price, for each future delivery period. This is calculated using pathwise differentiation as part of the Monte Carlo simulation, hence if not many simulation paths are used, as in this example, it can look quite jagged. With a higher number of simulations this will be smoother.
```
three_factor_results.deltas.plot(title='Deltas')
```
### Intrinsic Profile
The **intrinsic_profile** property is of type pandas.DataFrame and contains information for each future time period on the optimisation performed for the intrinsic valuation.
```
three_factor_results.intrinsic_profile
```
### Trigger Prices
The **trigger_prices** property contains information on "trigger prices" which are approximate spot price levels at which the exercise decision changes.
* The withdraw trigger price is the spot price level, at time of nomination, above which the optimal decision will change to withdraw.
* The inject trigger price is the spot price level, at time of nomination, below which the optimal decision will change to inject.
The columns 'withdraw_volume' and 'inject_volume' contain the volumes withdrawn or injected at the trigger prices respectively. Note that these volumes could have lower absolute value to the storage maximum withdrawal and injection rates, because inventory constraints might mean that these decision volumes aren't achievable.
```
display(three_factor_results.trigger_prices[0:10])
ax = three_factor_results.trigger_prices['inject_trigger_price'].plot(
title='Trigger Prices Versus Forward Curve', legend=True)
three_factor_results.trigger_prices['withdraw_trigger_price'].plot(legend=True)
fwd_curve['2021-04-25' : '2022-04-01'].plot(legend=True)
ax.legend(['Inject Trigger Price', 'Withdraw Trigger', 'Forward Curve'])
```
### Simulated Values
**MultiFactorValuationResults** has a number of properties of type pandas.DataFrame which contain information at the individual simulation level from the internals of the Least Squares Monte Carlo valuation. Each of these DataFrame instances will have index (row label) of type PeriodIndex with frequency corresponding to the frequency string **freq** argument used to create the CmdtyStorage instance. These correspond to the periods for which the storage is active. The column labels will be integers corresponding to the Monte Carlo simulation number, hence will be in the range from 0 (inclusive) to num_sims (exclusive). The following is a list of such properties:
* **sim_spot_regress** is the first set of simulated spot prices are used to calculate the exercise continuation values via regression during a backward induction.
* **sim_spot_valuation** is the second set of simulated spot prices for which exercise (inject/withdraw) decisions are simulated going forward in time using the regression derived continuation values. The final NPV is calulated using the cash flow derived from these exercise decisions.
* **sim_inject_withdraw** contains the simulation optimal inject or withdraw volumes.
* **sim_inventory** contains the simulation inventory. For each simulation and time period this will be the cummulative sum of the starting inventory and the inject/withdraw volumes up until the time period.
* **sim_cmdty_consumed** contains the simulation commodity consumed upon injection/withdrawal as described in [this section](creating_storage_instances.ipynb#commodity_consumed) of the creating_storage_instances notebook.
* **sim_inventory_loss** contains the simulated inventory loss, as described [here](creating_storage_instances.ipynb#inventory_cost_loss).
* **sim_net_volume** contains the simulated net volume which will need to be in place at nomination time due to injection/withdrawal. This equals minus sim_inject_withdraw (because withdrawals are represented by negative numbers) minus sim_cmdty_consumed.
* **sim_pv** contains the simulated PV at each time period. This equals the discounted sum of:
* Cash flow generated by purchasing or selling commodity on the spot market to inject or withdraw respectively, which is minus the corresponding volume in sim_inject_withdraw multiplied by the simulated spot price from sim_spot_valuation.
* Minus the inject/withdraw cost.
* Minus the corresponding value in sim_cmdty_consumed multiplied by the simulated spot price from sim_spot_valuation. This is the cost of commodity consumed upon injection/withdrawal.
* Minus the inventory cost.
The following example shows usage of the sim_spot_valuation property to chart the mean, 10th and 90th percentile of simulated spot prices against the initial forward curve.
```
sim_spot_valuation_mean = three_factor_results.sim_spot_valuation.aggregate(func='mean', axis='columns', raw = True)
sim_spot_valuation_90th = three_factor_results.sim_spot_valuation.aggregate(func=np.percentile, q=90, axis='columns', raw = True)
sim_spot_valuation_10h = three_factor_results.sim_spot_valuation.aggregate(func=np.percentile, q=10, axis='columns', raw = True)
sim_spot_valuation_mean.plot(title = 'Statistics of Simulated Spot Prices', legend=True)
fwd_curve['2021-04-25' : '2022-04-01'].plot(legend=True)
sim_spot_valuation_10h.plot(legend=True)
ax = sim_spot_valuation_90th.plot(legend=True)
ax.legend(['Mean', 'Forward Curve', '10th Percentile', '90th Percentile'])
```
### Expected Profile
The **expected_profile** property is a Pandas DataFrame containing the expected value of several quantities simulated during the valuation.
```
three_factor_results.expected_profile
```
***
## Instrumentation
There are two ways to instrument the valuation code: an on-progress call-back and logging. This section describes both of these.
### Progress Callback
Both valuation functions have an optional argument **on_progress_update** which should be assigned to a callable (i.e. a function) which accepts one argument and has void return. When running the valuation the callable will be invoked with a float instance passed in to the single argument with a numeric approximation of the percentage progress of the total calculation. A typical usage scenario would be to wire this up to a progress bar. This is particularly important from a usability perspective, when a high number of Monte-Carlo simulations are used, hence the calculations can take a long time.
The example below demonstrates this with a IPython progress bar widget.
```
progress_wgt = ipw.FloatProgress(min=0.0, max=1.0)
def on_progress(progress):
progress_wgt.value = progress
def btn_value_progress_clicked(b):
on_progress_demo_results = three_factor_seasonal_value(
cmdty_storage = storage,
val_date = '2021-04-25',
inventory = 1500.0,
fwd_curve = fwd_curve,
interest_rates = ir_curve,
settlement_rule = lambda decision_date: decision_date.asfreq('M').asfreq('D', 'end') + 20,
spot_mean_reversion = 45.0,
spot_vol = 0.65,
long_term_vol = 0.21,
seasonal_vol = 0.18,
num_sims = 500,
basis_funcs = '1 + x_st + x_sw + x_lt + x_st**2 + x_sw**2 + x_lt**2 + x_st**3 + x_sw**3 + x_lt**3',
discount_deltas=True,
on_progress_update = on_progress
)
btn_calc_value_progress = ipw.Button(description='Calc Storage Value')
btn_calc_value_progress.on_click(btn_value_progress_clicked)
display(btn_calc_value_progress)
display(progress_wgt)
```
### Logging
The cmdty_storage package makes use of the [standard Python logging module](https://docs.python.org/3/library/logging.html) by exposing a logger object at the level of the cmdty_storage package, which is instantiated with the name 'cmdty.storage'. A log handler can be added to this in order to direct the logged information to the desired destination. See the [logging advanced tutorial](https://docs.python.org/3/howto/logging.html#logging-advanced-tutorial) for more information on using appenders.
The following uses slightly modified code from an example in the ipywideget [documentation](https://ipywidgets.readthedocs.io/en/latest/examples/Output%20Widget.html#Integrating-output-widgets-with-the-logging-module) to create a custom logging handler which prints to an [Output widget](https://ipywidgets.readthedocs.io/en/latest/examples/Output%20Widget.html).
```
import cmdty_storage
import logging
class OutputWidgetHandler(logging.Handler):
""" Custom logging handler sending logs to an output widget """
def __init__(self, *args, **kwargs):
super(OutputWidgetHandler, self).__init__(*args, **kwargs)
layout = {
'width': '100%',
'height': '160px',
'border': '1px solid black',
'overflow_y': 'auto',
}
self.out = ipw.Output(layout=layout)
def emit(self, record):
""" Overload of logging.Handler method """
formatted_record = self.format(record)
new_output = {
'name': 'stdout',
'output_type': 'stream',
'text': formatted_record+'\n'
}
self.out.outputs = self.out.outputs + (new_output, )
def show_logs(self):
""" Show the logs """
display(self.out)
def clear_logs(self):
""" Clear the current logs """
self.out.clear_output()
handler = OutputWidgetHandler()
handler.setFormatter(logging.Formatter('%(asctime)s - [%(levelname)s] %(message)s'))
cmdty_storage.logger.setLevel(logging.INFO)
def btn_value_logging_clicked(b):
cmdty_storage.logger.addHandler(handler)
logging_demo_results = three_factor_seasonal_value(
cmdty_storage = storage,
val_date = '2021-04-25',
inventory = 1500.0,
fwd_curve = fwd_curve,
interest_rates = ir_curve,
settlement_rule = lambda decision_date: decision_date.asfreq('M').asfreq('D', 'end') + 20,
spot_mean_reversion = 45.0,
spot_vol = 0.65,
long_term_vol = 0.21,
seasonal_vol = 0.18,
num_sims = 500,
basis_funcs = '1 + x_st + x_sw + x_lt + x_st**2 + x_sw**2 + x_lt**2 + x_st**3 + x_sw**3 + x_lt**3',
discount_deltas=True,
)
cmdty_storage.logger.removeHandler(handler)
def btn_on_clear_logs_clicked(b):
handler.clear_logs()
btn_calc_value_logging = ipw.Button(description='Calc Storage Value')
btn_calc_value_logging.on_click(btn_value_logging_clicked)
display(btn_calc_value_logging)
btn_clear_logs = ipw.Button(description='Clear Logs')
btn_clear_logs.on_click(btn_on_clear_logs_clicked)
display(btn_clear_logs)
handler.show_logs()
```
***
## References
* Boogert, A. and de Jong, C. (). Gas storage valuation using a multi-factor process. J.Energy Mark., 4:1–24, 12 2011. doi: 10.21314/JEM.2011.067.
* Fowler, J. C. (2020). *Multi-factor Commodity Price Process: Spot and Forward Price Simulation*. URL https://github.com/cmdty/core/blob/master/docs/multifactor_price_process/multifactor_price_process.pdf.
| github_jupyter |
# Interpolating Network Sets
Frequently a set of `Networks` is recorded while changing some other parameters; like temperature, voltage, current, etc. Once this set of data acquired, it is sometime usefull to estimate the behaviour of the network for parameter values that lie in between those that have been mesured. For that purpose, interpolating a network from a set of network is possible. This example demonstrates how to do this using [NetworkSets](../../tutorials/NetworkSet.ipynb).
```
import skrf as rf
import matplotlib.pyplot as plt
import numpy as np
rf.stylely()
```
## Narda 3752 phase shifter
In this example, we are characterizing a old [narda phase shifter 3752](https://nardamiteq.com/docs/119-PHASESHIFTERS.PDF) at 1.5 GHz.
 :
In order to deduce the phase shift that one can obtain at this specific frequency, we have measured scattering parameters in the 1-2 GHz band at 19 positions of the phase knob (from 0 to 180). These measurements are loaded into a [NetworkSets](../../tutorials/NetworkSet.ipynb) object:
```
# Array containing the 19 phase shift indicator values
indicators_mes = np.linspace(0, 180, num=19) # from 0 to 180 per 10
ntw_set = rf.NetworkSet.from_zip('phase_shifter_measurements/phase_shifter_measurements.zip')
print('ntw_set contains', len(ntw_set), 'networks')
```
We extract from the network set the phase shift and $S_{11}$ at the specific frequency of 1.5 GHz:
```
f = '1.5 GHz'
phases_mes = np.squeeze([ntw[f].s21.s_deg for ntw in ntw_set])
s11_mes = np.squeeze([ntw[f].s11.s_db for ntw in ntw_set])
```
We would like however to get the phase shift values for intermediate settings of this phase shifter. To that purpose, let's create a network from the interpolation of the measured networks.
```
# the indicator values for which we want to interpolate the network
indicators = np.linspace(0, 180, num=181) # every degrees for 0 to 180
phases_interp = [ntw_set.interpolate_from_network(indicators_mes, phi)['1.5GHz'].s21.s_deg for phi in indicators]
phases_interp = np.squeeze(phases_interp)
s11_interp = [ntw_set.interpolate_from_network(indicators_mes, phi, interp_kind='quadratic')['1.5GHz'].s11.s_db for phi in indicators]
s11_interp = np.squeeze(s11_interp)
print('We have interpolated the network for', len(phases_interp), 'points')
```
Now let's plot everything
```
fig, (ax1, ax2) = plt.subplots(2,1, figsize=(10,7), sharex=True)
ax1.set_title('Narda Phase Shifter vs Indicator @ 1.5 GHz', fontsize=10)
ax1.plot(indicators, s11_interp, label='Interpolated network')
ax1.plot(indicators_mes, s11_mes, '.', ms=10, label='Measurements')
ax1.legend()
ax1.set_ylabel(r'$S_{11}$ [dB]')
ax2.plot(indicators, phases_interp, label='Interpolated network')
ax2.plot(indicators_mes, phases_mes, '.', ms=10, label='Measurements')
ax2.set_xlabel('Phase shifter indicator')
ax2.set_ylabel('$S_{21}$ [deg]')
ax2.fill_between(indicators, phases_mes[0], phases_mes[-1], alpha=0.2, color='r')
ax2.text(40, 25, 'not available zone')
ax2.legend()
ax2.set_ylim(-200, 200)
fig.tight_layout()
```
| github_jupyter |
# The Mesh: Where do things live?
<img src="images/FiniteVolume.png" width=70% align="center">
<h4 align="center">Figure 3. Anatomy of a finite volume cell.</h4>
To bring our continuous equations into the computer, we need to discretize the earth and represent it using a finite(!) set of numbers. In this tutorial we will explain the discretization in 2D and generalize to 3D in the notebooks. A 2D (or 3D!) mesh is used to divide up space, and we can represent functions (fields, parameters, etc.) on this mesh at a few discrete places: the nodes, edges, faces, or cell centers. For consistency between 2D and 3D we refer to faces having area and cells having volume, regardless of their dimensionality. Nodes and cell centers naturally hold scalar quantities while edges and faces have implied directionality and therefore naturally describe vectors. The conductivity, $\sigma$, changes as a function of space, and is likely to have discontinuities (e.g. if we cross a geologic boundary). As such, we will represent the conductivity as a constant over each cell, and discretize it at the center of the cell. The electrical current density, $\vec{j}$, will be continuous across conductivity interfaces, and therefore, we will represent it on the faces of each cell. Remember that $\vec{j}$ is a vector; the direction of it is implied by the mesh definition (i.e. in $x$, $y$ or $z$), so we can store the array $\bf{j}$ as *scalars* that live on the face and inherit the face's normal. When $\vec{j}$ is defined on the faces of a cell the potential, $\vec{\phi}$, will be put on the cell centers (since $\vec{j}$ is related to $\phi$ through spatial derivatives, it allows us to approximate centered derivatives leading to a staggered, second-order discretization).
# Implementation
```
%matplotlib inline
import numpy as np
from SimPEG import Mesh, Utils
import matplotlib.pyplot as plt
plt.set_cmap(plt.get_cmap('viridis')) # use a nice colormap!
```
## Create a Mesh
A mesh is used to divide up space, here we will use [SimPEG's mesh class](http://docs.simpeg.xyz/content/api_core/api_Mesh.html) to define a simple tensor mesh. By "Tensor Mesh" we mean that the mesh can be completely defined by the tensor products of vectors in each dimension; for a 2D mesh, we require one vector describing the cell widths in the x-direction and another describing the cell widths in the y-direction.
Here, we define and plot a simple 2D mesh using SimPEG's mesh class. The cell centers boundaries are shown in blue, cell centers as red dots and cell faces as green arrows (pointing in the positive x, y - directions). Cell nodes are plotted as blue squares.
```
# Plot a simple tensor mesh
hx = np.r_[2., 1., 1., 2.] # cell widths in the x-direction
hy = np.r_[2., 1., 1., 1., 2.] # cell widths in the y-direction
mesh2D = Mesh.TensorMesh([hx,hy]) # construct a simple SimPEG mesh
mesh2D.plotGrid(nodes=True, faces=True, centers=True) # plot it!
# This can similarly be extended to 3D (this is a simple 2-cell mesh)
hx = np.r_[2., 2.] # cell widths in the x-direction
hy = np.r_[2.] # cell widths in the y-direction
hz = np.r_[1.] # cell widths in the z-direction
mesh3D = Mesh.TensorMesh([hx,hy,hz]) # construct a simple SimPEG mesh
mesh3D.plotGrid(nodes=True, faces=True, centers=True) # plot it!
```
## Counting things on the Mesh
Once we have defined the vectors necessary for construsting the mesh, it is there are a number of properties that are often useful, including keeping track of the
- number of cells: **`mesh.nC`**
- number of cells in each dimension: **`mesh.vnC`**
- number of faces: **`mesh.nF`**
- number of x-faces: **`mesh.nFx`** (and in each dimension **`mesh.vnFx`** ...)
and the list goes on. Check out [SimPEG's mesh documentation](http://docs.simpeg.xyz/content/api_core/api_FiniteVolume.html) for more.
```
# Construct a simple 2D, uniform mesh on a unit square
mesh = Mesh.TensorMesh([10, 8])
mesh.plotGrid()
"The mesh has {nC} cells and {nF} faces".format(nC=mesh.nC, nF=mesh.nF)
# Sometimes you need properties in each dimension
("In the x dimension we have {vnCx} cells. This is because our mesh is {vnCx} x {vnCy}.").format(
vnCx=mesh.vnC[0],
vnCy=mesh.vnC[1]
)
# Similarly, we need to keep track of the faces, we have face grids in both the x, and y
# directions.
("Faces are vectors so the number of faces pointing in the x direction is {nFx} = {vnFx0} x {vnFx1} "
"In the y direction we have {nFy} = {vnFy0} x {vnFy1} faces").format(
nFx=mesh.nFx,
vnFx0=mesh.vnFx[0],
vnFx1=mesh.vnFx[1],
nFy=mesh.nFy,
vnFy0=mesh.vnFy[0],
vnFy1=mesh.vnFy[1]
)
```
## Simple properties of the mesh
There are a few things that we will need to know about the mesh and each of it's cells, including the
- cell volume: **`mesh.vol`**,
- face area: **`mesh.area`**.
For consistency between 2D and 3D we refer to faces having area and cells having volume, regardless of their dimensionality.
```
# On a uniform mesh, not suprisingly, the cell volumes are all the same
plt.colorbar(mesh.plotImage(mesh.vol, grid=True)[0])
plt.title('Cell Volumes');
# All cell volumes are defined by the product of the cell widths
assert (np.all(mesh.vol == 1./mesh.vnC[0] * 1./mesh.vnC[1])) # all cells have the same volume on a uniform, unit cell mesh
print("The cell volume is the product of the cell widths in the x and y dimensions: "
"{hx} x {hy} = {vol} ".format(
hx = 1./mesh.vnC[0], # we are using a uniform, unit square mesh
hy = 1./mesh.vnC[1],
vol = mesh.vol[0]
)
)
# Similarly, all x-faces should have the same area, equal to that of the length in the y-direction
assert np.all(mesh.area[:mesh.nFx] == 1.0/mesh.nCy) # because our domain is a unit square
# and all y-faces have an "area" equal to the length in the x-dimension
assert np.all(mesh.area[mesh.nFx:] == 1.0/mesh.nCx)
print(
"The area of the x-faces is {xFaceArea} and the area of the y-faces is {yFaceArea}".format(
xFaceArea=mesh.area[0],
yFaceArea=mesh.area[mesh.nFx]
)
)
mesh.plotGrid(faces=True)
# On a non-uniform tensor mesh, the first mesh we defined, the cell volumes vary
# hx = np.r_[2., 1., 1., 2.] # cell widths in the x-direction
# hy = np.r_[2., 1., 1., 1., 2.] # cell widths in the y-direction
# mesh2D = Mesh.TensorMesh([hx,hy]) # construct a simple SimPEG mesh
plt.colorbar(mesh2D.plotImage(mesh2D.vol, grid=True)[0])
plt.title('Cell Volumes');
```
## Grids and Putting things on a mesh
When storing and working with features of the mesh such as cell volumes, face areas, in a linear algebra sense, it is useful to think of them as vectors... so the way we unwrap is super important.
Most importantly we want some compatibility with <a href="https://en.wikipedia.org/wiki/Vectorization_(mathematics)#Compatibility_with_Kronecker_products">**Kronecker products**</a> as we will see later! This actually leads to us thinking about unwrapping our vectors column first. This column major ordering is inspired by linear algebra conventions which are the standard in Matlab, Fortran, Julia, but sadly not Python. To make your life a bit easier, you can use our MakeVector `mkvc` function from Utils.
```
from SimPEG.Utils import mkvc
mesh = Mesh.TensorMesh([3,4])
vec = np.arange(mesh.nC)
row_major = vec.reshape(mesh.vnC, order='C')
print('Row major ordering (standard python)')
print(row_major)
col_major = vec.reshape(mesh.vnC, order='F')
print('\nColumn major ordering (what we want!)')
print(col_major)
# mkvc unwraps using column major ordering, so we expect
assert np.all(mkvc(col_major) == vec)
print('\nWe get back the expected vector using mkvc: {vec}'.format(vec=mkvc(col_major)))
```
### Grids on the Mesh
When defining where things are located, we need the spatial locations of where we are discretizing different aspects of the mesh. A SimPEG Mesh has several grids. In particular, here it is handy to look at the
- Cell centered grid: **`mesh.gridCC`**
- x-Face grid: **`mesh.gridFx`**
- y-Face grid: **`mesh.gridFy`**
```
# gridCC
"The cell centered grid is {gridCCshape0} x {gridCCshape1} since we have {nC} cells in the mesh and it is {dim} dimensions".format(
gridCCshape0=mesh.gridCC.shape[0],
gridCCshape1=mesh.gridCC.shape[1],
nC=mesh.nC,
dim=mesh.dim
)
# The first column is the x-locations, and the second the y-locations
mesh.plotGrid()
plt.plot(mesh.gridCC[:,0], mesh.gridCC[:,1],'ro')
# gridFx
"Similarly, the x-Face grid is {gridFxshape0} x {gridFxshape1} since we have {nFx} x-faces in the mesh and it is {dim} dimensions".format(
gridFxshape0=mesh.gridFx.shape[0],
gridFxshape1=mesh.gridFx.shape[1],
nFx=mesh.nFx,
dim=mesh.dim
)
mesh.plotGrid()
plt.plot(mesh.gridCC[:,0], mesh.gridCC[:,1],'ro')
plt.plot(mesh.gridFx[:,0], mesh.gridFx[:,1],'g>')
```
## Putting a Model on a Mesh
In [index.ipynb](index.ipynb), we constructed a model of a block in a whole-space, here we revisit it having defined the elements of the mesh we are using.
```
mesh = Mesh.TensorMesh([100, 80]) # setup a mesh on which to solve
# model parameters
sigma_background = 1. # Conductivity of the background, S/m
sigma_block = 10. # Conductivity of the block, S/m
# add a block to our model
x_block = np.r_[0.4, 0.6]
y_block = np.r_[0.4, 0.6]
# assign them on the mesh
sigma = sigma_background * np.ones(mesh.nC) # create a physical property model
block_indices = ((mesh.gridCC[:,0] >= x_block[0]) & # left boundary
(mesh.gridCC[:,0] <= x_block[1]) & # right boudary
(mesh.gridCC[:,1] >= y_block[0]) & # bottom boundary
(mesh.gridCC[:,1] <= y_block[1])) # top boundary
# add the block to the physical property model
sigma[block_indices] = sigma_block
# plot it!
plt.colorbar(mesh.plotImage(sigma)[0])
plt.title('electrical conductivity, $\sigma$')
```
## Next up ...
In the [next notebook](divergence.ipynb), we will work through defining the discrete divergence.
| github_jupyter |
ПИ19-4 Деменчук Георгий КР
### 1. Создайте отображение базы данных студентов *Students_2021.sqlite*
```
from sqlalchemy.ext.automap import automap_base
from sqlalchemy.orm import Session
from sqlalchemy import create_engine
from datetime import datetime
engine = create_engine('sqlite:///Students_2021.sqlite')
Base = automap_base()
Base.prepare(engine, reflect=True)
session = Session(engine)
```
### 2. Напишите запрос, позволяющий получить из таблицы *exam_marks* значения столбца *mark* (экзаменационная оценка) для всех студентов, исключив из списка повторение одинаковых строк.
```
city,exam_marks,student,university,subject,lecturer,subj_lect = Base.classes.values()
from sqlalchemy.orm import Session
session = Session(engine)
for item in session.query(exam_marks.mark).distinct().all():
print(item.mark)
```
### 3. Напишите запрос для получения списка студентов без определенного места жительства.
```
for item in session.query(student.name, student.surname).where(student.city == None).all():
print(item)
```
### 4. Напишите запрос для получения списка студентов, проживающих в Воронеже и не получающих стипендию.
```
from sqlalchemy import and_
for item in session.query(student.name, student.surname, student.city, student.stipend).where(
and_(
student.city == "Воронеж",
student.stipend == 0
)
).all():
print(item)
```
### 5. Напишите запрос для получения списка университетов, расположенных в Москве и имеющих рейтинг меньший,
чем у НГУ. Значение рейтинга НГУ получите с помощью отдельного запроса или подзапроса.
```
from sqlalchemy import and_
subquery = session.query(university.rating).where(university.univ_name=="НГУ").subquery()
query1 = session.query(university.univ_name).where(
and_(
university.city == "Москва",
university.rating < (subquery)
)
)
for item in query1.all():
print(item)
```
### 6. Напишите запрос, выполняющий вывод находящихся в таблице EXAM_MARKS
номеров предметов обучения, экзамены по которым сдавались между 1 и 21 марта 2020 г.
```
from datetime import datetime
first_date = datetime.strptime('2020-03-1', '%Y-%m-%d')
second_date = datetime.strptime('2020-03-21', '%Y-%m-%d')
for item in session.query(exam_marks.subj_id, exam_marks.mark, exam_marks.exam_date).where(
exam_marks.exam_date.between(first_date, second_date)
).all():
print(item)
```
### 7. Напишите запрос, который выполняет вывод названий предметов обучения, начинающихся на букву ‘И’.
```
for item in session.query(subject.subj_id, subject.subj_name, subject.hour).where(
subject.subj_name.like("И%")
).all():
print(item)
```
### 8. Напишите запрос, выбирающий сведения о студентах, у которых имена начинаются на букву ‘И’ или ‘С’.
```
from sqlalchemy import or_
for item in session.query(student.name, student.surname).where(
or_(
student.name.like("И%"),
student.name.like("C%")
)
).all():
print(item)
```
### 9. Напишите запрос для получения списка предметов обучения, названия которых состоят из более одного слова.
```
for item in session.query(subject.subj_id, subject.subj_name, subject.hour).where(
subject.subj_name.like("% %")
).all():
print(item)
```
### 10. Напишите запрос для получения списка студентов, фамилии которых состоят из трех букв.
```
from sqlalchemy.sql.expression import func
for item in session.query(student.name, student.surname).where(
func.length(student.surname) == 3
).all():
print(item)
```
### 11. Составьте запрос для таблицы STUDENT таким образом, чтобы получить результат в следующем виде. Распечатайте первые 9 записей результата.
```
И. Иванов 1982-12-03
П. Петров 1980-12-01
В. Сидоров 1979-06-07
...
```
```
from sqlalchemy.sql.expression import func
for item in session.query(
cast(func.substr(student.name,1,1), String(20))+"."+
cast(student.surname, String(20))+" "+
cast(func.strftime('%Y-%m-%d',student.birthday), String(20))
).all():
print(item)
```
### 12. Напишите запрос для получения списка студентов, фамилии которых начинаются на ‘Ков’ или на ‘Куз’.
```
from sqlalchemy import or_
for item in session.query(student.surname, student.name).where(
or_(
student.surname.like("Ков%"),
student.surname.like("Куз%")
)
).all():
print(item)
```
### 13. Напишите запрос для получения списка предметов, названия которых оканчиваются на ‘ия’.
```
for item in session.query(subject.subj_id, subject.subj_name, subject.hour).where(
subject.subj_name.like("%ия")
).all():
print(item)
```
### 14. Напишите запрос для выбора из таблицы EXAM_MARKS записей, для которых отсутствуют значения оценок (поле MARK).
```
for item in session.query(exam_marks.subj_id, exam_marks.mark).where(
exam_marks.mark == None
).all():
print(item)
```
### 15. Составьте запрос, выводящий фамилии, имена студентов и величину получаемых ими стипендий,
при этом значения стипендий должны быть увеличены в 100 раз.
```
from sqlalchemy import cast, Integer
for item in session.query(student.surname, student.name, cast((student.stipend * 100), Integer)).all():
print(item)
```
### 16. Составьте запрос для таблицы UNIVERSITY таким образом,
чтобы выходная таблица содержала всего один столбец в следующем виде:
Код-10; ВГУ-г.ВОРОНЕЖ; Рейтинг=296.
```
from sqlalchemy import cast, String
for item in session.query("Код-"+cast(university.univ_id, String(20))+"; "+
cast(university.univ_name, String(20))+"-г."+
cast(university.city, String(20)) +"; Рейтинг="+
cast(university.rating, String(20)) +".").all():
print(item)
```
### 17. Напишите запрос для подсчета количества студентов, сдававших экзамен по предмету обучения с идентификатором 10.
```
from sqlalchemy import cast, String
for item in session.query(
func.count(exam_marks.student_id)
).where(exam_marks.subj_id == 10).all():
print(item)
print("-------")
from sqlalchemy import cast, String
for item in session.query(exam_marks.student_id
).where(exam_marks.subj_id == 10).all():
print(item)
```
### 18. Напишите запрос, который позволяет подсчитать в таблице EXAM_MARKS количество различных предметов обучения.
```
for item in session.query(exam_marks.subj_id).distinct().all():
print(item)
print("-------")
session.query(exam_marks.subj_id).distinct().count()
```
### 19. Напишите запрос, который для каждого студента выполняет выборку его идентификатора и минимальной из полученных им оценок.
```
for item in session.query(exam_marks.student_id, func.min(exam_marks.mark)).group_by(
exam_marks.student_id
).all():
print(item)
```
### 20. Напишите запрос, который для каждого предмета обучения выводит наименование предмета и максимальное значение номера семестра, в котором этот предмет преподается.
```
for item in session.query(subject.subj_name,
func.max(subject.semester.label("max_semester"))).group_by(subject.subj_name).all():
print(item)
```
### 21. Напишите запрос, который для каждого конкретного дня сдачи экзамена выводит данные о количестве студентов, сдававших экзамен в этот день.
```
for item in session.query(
func.count(func.distinct(exam_marks.student_id)).label("Кол-во студентов"),
exam_marks.exam_date.label("Дата экзамена"),
exam_marks.exam_id.label("id экзамена")
).group_by(exam_marks.exam_date).all():
print(item)
```
### 22. Напишите запрос, выдающий средний балл для каждого студента.
```
for item in session.query(
exam_marks.student_id,
func.avg(exam_marks.mark)
).group_by(exam_marks.student_id).all():
print(item)
```
### 23. Напишите запрос, выдающий средний балл для каждого экзамена.
```
for item in session.query(
exam_marks.exam_id,
func.avg(exam_marks.mark)
).group_by(exam_marks.exam_id).all():
print(item)
```
| github_jupyter |
# Check LSLGA in DR8
#### John Moustakas
```
import os, pdb
import numpy as np
from glob import glob
import fitsio
from astropy.table import vstack, Table, hstack
import seaborn as sns
sns.set(context='talk', style='ticks', font_scale=1.6)
%matplotlib inline
dr8dir = '/global/project/projectdirs/cosmo/data/legacysurvey/dr8'
outdir = '/global/project/projectdirs/desi/users/ioannis/dr8-lslga'
lslgafile = '/global/project/projectdirs/cosmo/staging/largegalaxies/v2.0/LSLGA-v2.0.fits'
lslga = Table(fitsio.read(lslgafile))
#lslga
def build_lslga_dr8(northsouth, overwrite=False):
cols = ['RA', 'DEC', 'TYPE', 'BRICKNAME', 'REF_CAT',
'FRACDEV', 'SHAPEEXP_R', 'SHAPEEXP_E1', 'SHAPEEXP_E2',
'SHAPEDEV_R', 'SHAPEDEV_E1', 'SHAPEDEV_E2', 'REF_ID']
outfile = os.path.join(outdir, 'dr8-lslga-{}.fits'.format(northsouth))
if not os.path.isfile(outfile) or overwrite:
out = []
catfile = glob(os.path.join(dr8dir, northsouth, 'sweep', '8.0', 'sweep-*.fits'))
#catfile = glob(os.path.join(dr8dir, northsouth, 'tractor', '???', 'tractor*.fits'))
for ii, ff in enumerate(catfile):#[:3]):
if ii % 20 == 0:
print('{} / {}'.format(ii, len(catfile)))
cc = fitsio.read(ff, columns=cols, upper=True)
ckeep = np.where(cc['REF_CAT'] == b'L2')[0]
out.append(Table(cc[ckeep]))
out = vstack(out)
# find and remove duplicates
#uval, cnt = np.unique(out['REF_ID'], return_counts=True)
#dup_refid = uval[cnt > 1]
# now match against the LSLGA
match = np.hstack([np.where(refid == lslga['LSLGA_ID'])[0] for refid in out['REF_ID']])
lss = lslga[match]
lss.rename_column('RA', 'RA_LSLGA')
lss.rename_column('DEC', 'DEC_LSLGA')
lss.rename_column('TYPE', 'MORPHTYPE')
out = hstack((lss, out))
print('Writing {}'.format(outfile))
out.write(outfile, overwrite=True)
else:
print('Reading {}'.format(outfile))
out = Table(fitsio.read(outfile))
return out
%time north = build_lslga_dr8('north', overwrite=False)
%time south = build_lslga_dr8('south', overwrite=False)
def merge_north_south(overwrite=False):
mergedfile = os.path.join(outdir, 'dr8-lslga-northsouth.fits')
if not os.path.isfile(mergedfile) or overwrite:
out = []
for northsouth in ('north', 'south'):
catfile = os.path.join(outdir, 'dr8-lslga-{}.fits'.format(northsouth))
cat = Table(fitsio.read(catfile))
cat['REGION'] = northsouth
out.append(cat)
out = vstack(out)
print('Writing {}'.format(mergedfile))
out.write(mergedfile, overwrite=True)
else:
print('Reading {}'.format(mergedfile))
out = Table(fitsio.read(mergedfile))
return out
%time cat = merge_north_south(overwrite=True)
cat
urefid, uindx = np.unique(cat['REF_ID'], return_index=True)
dup = np.delete(np.arange(len(cat)), uindx)
cat[dup]
cat[cat['REF_ID'] == 462171]
ww = [tt.strip() == 'REX' for tt in cat['TYPE']]
cat[ww]
```
#### Test out the code for adding this catalog as a separate layer
```
from astrometry.util.fits import fits_table
T = fits_table(os.path.join(outdir, 'dr8-lslga-northsouth.fits'))
def get_ba_pa(e1, e2):
ee = np.hypot(e1, e2)
ba = (1 - ee) / (1 + ee)
#pa = -np.rad2deg(np.arctan2(e2, e1) / 2)
pa = 180 - (-np.rad2deg(np.arctan2(e2, e1) / 2))
return ba, pa
def lslga_model(T):
ba, pa = [], []
for Tone in T:
ttype = Tone.type.strip()
if ttype == 'DEV' or ttype == 'COMP':
e1, e2 = Tone.shapedev_e1, Tone.shapedev_e2
_ba, _pa = get_ba_pa(e1, e2)
elif ttype == 'EXP' or ttype == 'REX':
e1, e2 = Tone.shapeexp_e1, Tone.shapeexp_e2
_ba, _pa = get_ba_pa(e1, e2)
else: # PSF
_ba, _pa = np.nan, np.nan
ba.append(_ba)
pa.append(_pa)
T.ba_model = np.hstack(ba)
T.pa_model = np.hstack(pa)
T.radius_model_arcsec = T.fracdev * T.shapedev_r + (1 - T.fracdev) * T.shapeexp_r
T.radius_arcsec = T.d25 / 2. * 60.
return T
rr = lslga_model(T[:10])
rr.pa, rr.pa_model
rr.ba, rr.ba_model
rr.radius_arcsec, rr.radius_arcsec_model
```
| github_jupyter |
Uno de los problemas más comunes con que nos solemos encontrar al desarrollar cualquier programa informático, es el de procesamiento de texto. Esta tarea puede resultar bastante trivial para el cerebro humano, ya que nosotros podemos detectar con facilidad que es un número y que una letra, o cuales son palabras que cumplen con un determinado patrón y cuales no; pero estas mismas tareas no son tan fáciles para una computadora. Es por esto, que el procesamiento de texto siempre ha sido uno de los temas más relevantes en las ciencias de la computación. Luego de varias décadas de investigación se logró desarrollar un poderoso y versátil lenguaje que cualquier computadora puede utilizar para reconocer patrones de texto; este lenguale es lo que hoy en día se conoce con el nombre de expresiones regulares; las operaciones de validación, búsqueda, extracción y sustitución de texto ahora son tareas mucho más sencillas para las computadoras gracias a las expresiones regulares.
## ¿Qué son las Expresiones Regulares?
Las expresiones regulares, a menudo llamada también regex, son unas secuencias de caracteres que forma un patrón de búsqueda, las cuales son formalizadas por medio de una sintaxis específica. Los patrones se interpretan como un conjunto de instrucciones, que luego se ejecutan sobre un texto de entrada para producir un subconjunto o una versión modificada del texto original. Las expresiones regulares pueden incluir patrones de coincidencia literal, de repetición, de composición, de ramificación, y otras sofisticadas reglas de reconocimiento de texto . Las expresiones regulares deberían formar parte del arsenal de cualquier buen programador ya que un gran número de problemas de procesamiento de texto pueden ser fácilmente resueltos con ellas.
```
import re
pat = r"(Jaime)"
m = re.findall(pat, "Jaime Rodriguez irá con Jaime Perez al cine.")
```
Esta es una expresión regular que busca un patrón de correo electrónico:
`/[\w._%+-]+@[\w.-]+\.[a-zA-Z]{2,4}/`
Pero, no te preocupes ... no tienes que entenderlo ahora. La buena noticia es que una expresión regular compleja es simplemente la combinación de varias expresiones regulares muy simples. "¡Divide y vencerás!"
| abc… | Letters |
|-|-|
| 123… | Digits |
| \d | Any Digit |
| \D | Any Non-digit character |
| . | Any Character |
| \. | Period |
| [abc] | Only a, b, or c |
| [^abc] | Not a, b, nor c |
| [a-z] | Characters a to z |
| [0-9] | Numbers 0 to 9 |
| \w | Any Alphanumeric character |
| \W | Any Non-alphanumeric character |
| {m} | m Repetitions |
| {m,n} | m to n Repetitions |
| * | Zero or more repetitions |
| + | One or more repetitions |
| ? | Optional character |
| \s | Any Whitespace |
| \S | Any Non-whitespace character |
| ^…$ | Starts and ends |
| (…) | Capture Group |
| (a(bc)) | Capture Sub-group |
| (.*) | Capture all |
| (abc\|def) | Matches abc or def |
## Raw strings
Los *raw strings* de Python son strings usuales a los que les antecede una r. Por ejemplo ``r"SciData"`` es un raw string. Su caraterística principal es que no consideran el símbolo `\` como un escape.
```
saludo = "Hola\nMundo.\tYa casi terminamos"
print(saludo)
r_saludo = r"Hola\nMundo.\tYa casi terminamos"
print(r_saludo)
```
## Findall: Encontrar todos los patrones en un string y que devuelva todas las coincidencias.
```
pat = r"Python"
txt = "Programamos en Python con los módulos pytorch y numpy."
matchs = re.findall(pat, txt)# r is for raw strings
print(matchs)
pat2 = r"py"
re.findall(pat2, txt)
pat3 = r"[Pp]y"
re.findall(pat3, txt)
mensaje = "Mi número es 610-742-8645"
pat4 = "[0-9]"
re.findall(pat4, mensaje)
mensaje = "Mi número es 610-742-8645"
pat4 = "\d"
re.findall(pat4, mensaje)
mensaje = "Mi número es 610-742-8645"
pat4 = r"\d\d\d-\d\d\d-\d\d\d\d"
re.findall(pat4, mensaje)
mensaje = "Mi número es 610-742-865"
pat4 = r"\d{3}-\d{3}-\d{3}"
re.findall(pat4, mensaje)
txt2 = "Mis números son 610-742-845 y 663852642"
pat4 = r"\d{3}-?\d{3}-?\d{3}"
re.findall(pat4, txt2)
```
## Group: to get the matched string
```
txt2 = "Mis números son 610-742-845 y 663852642"
pat4 = r"(\d{3})-?(\d{3})-?(\d{3})"
m = re.findall(pat4, txt2)
m[0]
m[0][0]
txt = "casa, caja, cama"
result3 = re.findall("ca[sjm]a", txt)
print(result3)
```
## ? Character: a lo más una coincidencia
```
expr = re.compile(r'Pyt(ho)?n')
match = expr.search("Python! a great language.")
match.group()
match = expr.search("Pytn! a great language.")
match.group()
match = expr.search("Pythohon a great language")
print(match)
```
## * Character: cero o más coincidencias
## + Character: al menos una coincidencia
## {} : n coincidencias
## {m,n} entre m y n coincidencias (máximo de coincidencias)
## Match the exact number of characters
```
string = 'The date is 22/10/2019'
lst = re.findall('[0-9]{1,2}\/[0-9]{1,2}\/[0-9]{4}', string)
print(lst)
```
## ^ start and end $
```
regex = re.compile(r'[^aeiouAEIOU]')
string = "The language is PythOn"
regex.findall(string)
```
## Escaping Special Characters
```
str = 'Sentences have dots.How do we escape them?\n'
lst = re.findall('.', str)
print(lst)
lst1 = re.findall('\n', str)
print(lst1)
```
[Tutorial](https://regexone.com/lesson/introduction_abcs)
[Problems](https://regexone.com/problem/matching_decimal_numbers)
## Reference:
https://docs.python.org/3/library/re.html
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.