text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
```
##====================================================
## このセルを最初に実行せよ---Run this cell initially.
##====================================================
import sys
if 'google.colab' in sys.modules:
!wget -P ./text https://www.eidos.ic.i.u-tokyo.ac.jp/~sato/assignments/project2/text/test_data.csv
!wget -P ./... | github_jupyter |
SOP038 - Install azure command line interface
=============================================
Steps
-----
### Common functions
Define helper functions used in this notebook.
```
# Define `run` function for transient fault handling, hyperlinked suggestions, and scrolling updates on Windows
import sys
import os
import ... | github_jupyter |
```
# cell used to import important library of the notebook
import numpy as np
import sys
from scipy import sparse
from scipy.spatial.distance import pdist, squareform
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
import pandas as pd
import networkx as nx
from sklearn.preprocessing import Stan... | github_jupyter |
# Preface
Learning a new language (programming or spoken) can be extremely difficult. Usually the first language you learn will probably be the hardest to learn. Trying to learn a second language will also be challenging, but hopefully you will be able to rely on your experience from the first language you learned to ... | github_jupyter |
```
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from sqlalchemy import create_engine
from hold import connection_string
engine = create_engine(f'{connection_string}', encoding='iso-8859-1', connect_args={'connect_timeout': 10})
gtdDF = pd.read_sql_table('global_terrorism', ... | github_jupyter |
# Mixture of Gaussians
```
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
```
Mixture of Gaussians (aka Expectation Maximation) is a clustering method. The idea of this model is simpel: for a given dataset, each point is generated by linearly combining multiple multivariate Gaussians.
## What... | github_jupyter |
# Tutorial 5:
## Random Forest Regression
Random Forests Regression is an ensemble learning method that combines multiple Decision Tree Regressions. The method uses a multitude of decision trees to train and predict values. Random Forests reduces the over-fitting in comparison to using a single Decision Tree model.
Fo... | github_jupyter |
```
from astropy.table import Table, Column
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib import colors
import os
import urllib.request
from tqdm import tqdm
import sys
os.chdir("/calvin1/benardorci/SimulationData")
os.getcwd()
Halos = np.load("/calvin1/benardorci/SimulationData... | github_jupyter |
```
import pandas as pd
from tqdm import tqdm
import spotipy
from spotipy.oauth2 import SpotifyClientCredentials #To access authorised Spotify data
client_id= "d8df752143c842238c774a4551d21546"
client_secret= "de7220a3104a43a0aed809594a5d92b1"
```
# Import data
```
pitchfork_df = pd.read_csv('pitchfork_reviews.tsv'... | github_jupyter |
<a href="https://colab.research.google.com/github/khbae/trading/blob/master/Petersen_Jupyter_Notebook.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Petersen - Estimating Standard Errors in Finance Panel Data Sets: Comparing Approaches (prepared ... | github_jupyter |
```
# import plaidml.keras
# plaidml.keras.install_backend()
# import os
# os.environ["KERAS_BACKEND"] = "plaidml.keras.backend"
# Importing useful libraries
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
from sklearn.preprocessing import MinMaxScaler
from keras.models import Sequential
from ker... | github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
```
# Derivadas aproximadas
Nem sempre é possível calcular a derivada de uma função.
Às vezes, a função em questão não é dada de forma explícita.
Por exemplo,
$$f(x) = \min_{|y| < x} \Big( \frac{\cos(2x^2 - 3y)}{20x - y} \Big).$$
Assim, teremos que _estimar_ a ... | github_jupyter |
# Introduction to Quantum Error Correction via the Repetition Code
## Introduction
Quantum computing requires us to encode information in qubits. Most quantum algorithms developed over the past few decades have assumed that these qubits are perfect: they can be prepared in any state we desire, and be manipulated with... | github_jupyter |
<a href="https://colab.research.google.com/github/rockerritesh/Artifical-Neural-Network/blob/master/NPL_and_SENTIMENT_CLASSIFICATION_using_simple_NEURAL_NETWORK.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
import nltk
from nltk.stem import P... | github_jupyter |
# Navigation
---
In this notebook, you will learn how to use the Unity ML-Agents environment for the first project of the [Deep Reinforcement Learning Nanodegree](https://www.udacity.com/course/deep-reinforcement-learning-nanodegree--nd893).
### 1. Start the Environment
We begin by importing some necessary packages... | github_jupyter |
## Introduction to Data Science
### Data Science Tasks: Recommender Systems
Based on [this](https://www.datacamp.com/community/tutorials/recommender-systems-python), [this](https://www.analyticsvidhya.com/blog/2016/06/quick-guide-build-recommendation-engine-python/) and [this](http://www.data-mania.com/blog/recommend... | github_jupyter |
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
```
## Object Creation
Creating a Series by passing a list of values, letting pandas create a default integer index (for the list)
```
s = pd.Series([1,3,5,np.nan,6,8])
s
```
Creating a DataFrame by passing numpy array, with datetime index a... | github_jupyter |
---
_You are currently looking at **version 1.1** of this notebook. To download notebooks and datafiles, as well as get help on Jupyter notebooks in the Coursera platform, visit the [Jupyter Notebook FAQ](https://www.coursera.org/learn/python-machine-learning/resources/bANLa) course resource._
---
## Assignment 4 - ... | github_jupyter |
# Adstock Campaign/Channel Attribution Model
This is a reference implementation of the adstock model that estimates the contribution of individual campaigns/channels to the total outcome (number of conversions, site traffic, etc.)
Assuming we observe activity data for several campaigns/channels as time series $x_{it}... | github_jupyter |
```
%matplotlib inline
import matplotlib.pyplot as plt
ax11 = plt.subplot(2, 2, 1)
ax12 = plt.subplot(2, 2, 2)
ax21 = plt.subplot(2, 2, 3)
ax22 = plt.subplot(2, 2, 4)
ax11.set_title("ax11")
ax12.set_title("ax12")
ax21.set_title("ax21")
ax22.set_title("ax22")
plt.tight_layout()
plt.show()
plt.savefig("images/subplots2.... | github_jupyter |
# Numerical Solution of the Wave Equation using the Finite Element Method
This notebook illustrates the numerical time-domain solution of the wave equation using the [Finite Element Method](https://en.wikipedia.org/wiki/Finite_element_method) (FEM). The method aims at an approximate solution by subdividing the area of... | github_jupyter |
# How to Use DeepBugs for Yourself
Follow along with this notebook to reproduce our replication of DeepBugs, tested on the switched-argument bug (i.e., the developer accidentally typed the arguments in reverse order.)
Or, feel free to just check out the pre-saved output - things can take a while to run.
You can also ... | github_jupyter |
# Learning to pivot, part 3
## Independence $\neq$ non significant
This example demonstrates that statistical independence of classifier predictions from the nuisance parameter does not imply that
classifier does not use nuisance parameter.
Main paper: https://arxiv.org/abs/1611.01046
```
try:
import mlhep2019
e... | github_jupyter |
```
from micromlgen import port
from sklearn.metrics import accuracy_score
from sklearn.svm import SVC
from sklearn.model_selection import train_test_split
import pandas as pd
import numpy as np
import random
# TYLKO ŻEBY OBLICZENIA BYŁY ZAWSZE TAKIE SAME
random.seed(123)
np.random.seed(123)
np.set_printoptions(pre... | github_jupyter |
# Distribution of insolation
**Note this should be updated to take advantage of the new xarray capabilities of the `daily_insolation` code.**
Here are some examples calculating daily average insolation at different locations and times.
These all use a function called `daily_insolation` in the module `insolation.py` ... | github_jupyter |
```
import csv
import os
import glob
import multiprocessing as mp
import pandas as pd
import numpy as np
pd.options.mode.chained_assignment = None
%matplotlib inline
import matplotlib.pyplot as plt
import matplotlib
matplotlib.style.use('ggplot')
from scipy.interpolate import interp1d
pd.set_option('display.float_forma... | github_jupyter |
# Completely optional
... but fun!
#### Geek-out about Pandas Expanding Rolling Windows follows (a.k.a. `"Let's measure the Earth!!"`)
Rolling windows are cool, especially because they forget the far past, and keep only the recent data "in mind" when performing operations. There are [many types of rolling window](htt... | github_jupyter |
# TOOLS_ELA
*tools_ela.py* is a command line tool that extracts information from original transcribed text. The provided information is:
1. Indexes of Words, Lemmas, Types
2. Frequencies of Lemmas, Types
3. Concordances
4. TTR
5. Collocations (for both words and lemmas)
6. N-grams
7. Min, Max and Mean lengths of Type... | github_jupyter |
```
import sys
sys.path.append('../../nucleon_elastic_FF/scripts/area51_files')
sys.path.append('../../nucleon_elastic_FF/scripts')
import sources
import utils
from lattedb.wavefunction.models import Hadron as wavefunction_Hadron
from lattedb.wavefunction.models import Hadron4D as wavefunction_Hadron4D
from lattedb.fer... | github_jupyter |
# MadMiner particle physics tutorial
# Part 3b: Training a score estimator
Johann Brehmer, Felix Kling, Irina Espejo, and Kyle Cranmer 2018-2019
In part 3a of this tutorial we will finally train a neural network to estimate likelihood ratios. We assume that you have run part 1 and 2a of this tutorial. If, instead of... | github_jupyter |
```
%load_ext watermark
%watermark -d -u -a 'Andreas Mueller, Kyle Kastner, Sebastian Raschka'
```
The use of watermark (above) is optional, and we use it to keep track of the changes while developing the tutorial material. (You can install this IPython extension via "pip install watermark". For more information, ple... | github_jupyter |
# Neural networks with PyTorch
Deep learning networks tend to be massive with dozens or hundreds of layers, that's where the term "deep" comes from. You can build one of these deep networks using only weight matrices as we did in the previous notebook, but in general it's very cumbersome and difficult to implement. Py... | github_jupyter |
## Step 1. 讀入檔案
```
# 下載 Gossiping 版 2005 至 2020 年,每五年的詞向量
!gdown --id "1gEL4v3wGgvqJnpWspISZvLeIL3GQZLB1" -O "Gossiping_2005.model" # 2005 年 Gossiping 板
!gdown --id "1yB9WPVDJVmmLLxbEHZroZP_cYMP0JUpC" -O "Gossiping_2010.model" # 2010 年 Gossiping 板
!gdown --id "1Vh8meq6hdte02nQ2-djclgpEKxFUC0YU" -O "Gossiping_2015.mod... | github_jupyter |
### Markov decision process
This week's methods are all built to solve __M__arkov __D__ecision __P__rocesses. In the broadest sense, an MDP is defined by how it changes states and how rewards are computed.
State transition is defined by $P(s' |s,a)$ - how likely are you to end at state $s'$ if you take action $a$ fro... | github_jupyter |
# Turbine isentropic efficiency
A steam turbine performs with an isentropic efficiency of $\eta_t = 0.84$. The inlet conditions are 4 MPa and 650°C, with a mass flow rate of 100 kg/s, and the exit pressure is 10 kPa. Assume the turbine is adiabatic.

**Problem:**
- Determine the ... | github_jupyter |
**Note**: Click on "*Kernel*" > "*Restart Kernel and Clear All Outputs*" in [JupyterLab](https://jupyterlab.readthedocs.io/en/stable/) *before* reading this notebook to reset its output. If you cannot run this file on your machine, you may want to open it [in the cloud <img height="12" style="display: inline-block" src... | github_jupyter |
# 1) Intro to Groupby Module
+ Groupby is like a container where each bundle has a common theme
```
import pandas as pd
fortune = pd.read_csv('Data/fortune1000.csv', index_col=['Rank'])
fortune.head(3)
```
## Groupby TIPS
- When we consider Groupby conditions, we want to group by using a column with smallest number o... | github_jupyter |
# Lab 3 Accuracy of Quantum Phase Estimation
Prerequisite
- [Ch.3.5 Quantum Fourier Transform](https://qiskit.org/textbook/ch-algorithms/quantum-fourier-transform.html)
- [Ch.3.6 Quantum Phase Estimation](https://qiskit.org/textbook/ch-algorithms/quantum-phase-estimation.html)
Other relevant materials
- [QCQI] Michae... | github_jupyter |
# Plot Performance Analysis
```
%load_ext autoreload
%autoreload 2
%matplotlib inline
import json
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
plt.rcParams['font.family'] = 'sans-serif'
plt.rcParams['font.sans-serif'] = 'Roboto Condensed'
sizes = [200, 500, 1000, 2000, 5000]
repeats = 10
```... | github_jupyter |
# 18 - Support Vector Machines
by [Alejandro Correa Bahnsen](albahnsen.com/)
version 0.1, Apr 2016
## Part of the class [Practical Machine Learning](https://github.com/albahnsen/PracticalMachineLearningClass)
This notebook is licensed under a [Creative Commons Attribution-ShareAlike 3.0 Unported License](http://c... | github_jupyter |
<h1 align="center">PROGRAMACIÓN DE COMPUTADORES </h1>
<h2 align="center">UNIVERSIDAD EAFIT</h2>
<h3 align="center">MEDELLÍN - COLOMBIA </h3>
<h2 align="center">Sesión 14 - Diccionarios</h2>
## Docente:
> <strong> *Carlos Alberto Álvarez Henao, I.C. Ph.D.* </strong>
### Diccionarios
>Un Diccionario es una estructu... | github_jupyter |
<div align="center">
<h1><img width="30" src="https://madewithml.com/static/images/rounded_logo.png"> <a href="https://madewithml.com/">Made With ML</a></h1>
Applied ML · MLOps · Production
<br>
Join 30K+ developers in learning how to responsibly <a href="https://madewithml.com/about/">deliver value</a> with ML.
... | github_jupyter |
```
import numpy as np
import pandas as pd
import subprocess
import argparse
```
# Preprocessing RecSys 2017
For the RecSys 2017 dataset we first need to artificially create sessions out of the user internactions
```
def make_sessions(data,
session_th=30 * 60,
is_ordered=False, ... | github_jupyter |
# Feature Engineering - Business Attributes
```
import pandas as pd
import numpy as np
from sklearn.preprocessing import StandardScaler
rev_busi_Pho= pd.read_csv('../data/filtered_reviews_in_Phonex.csv', parse_dates=["date"])
busi = pd.read_csv('../data/business_data_subset.csv')
busi.head(1)
```
### Change dict attr... | github_jupyter |
This notebook assumes kafka (and zookeeper) have been started and are available at localhost:9092.
https://medium.com/better-programming/your-local-event-driven-environment-using-dockerised-kafka-cluster-6e84af09cd95
```
$ docker-compose up -d
```
You can explicitly create Kafka topics with appropriate replication a... | github_jupyter |
# Charts for **REINFORCE** or Monte-Carlo policy gradient
My notes on REINFORCE Algorithm.
## Symbol Lookup Table
| Symbol | Definition |
|-------------------- |----------------------------------------------------------... | github_jupyter |
```
from google.colab import drive
drive.mount('/content/drive')
cd /content/drive/My Drive/FYP/Sentiment Analysis/Implementation/Sentiment Analysis/Capsule/Dynamic _rounting_enhancement_Capsule_network
!pip install tensorflow==1.14.0
!pip install keras==2.1.5
```
# Dependencies
```
from __future__ import absolute... | github_jupyter |
# Week 3 Assessment: Orthogonal Projections
## Learning Objectives
In this week, we will write functions which perform orthogonal projections.
By the end of this week, you should be able to
1. Write code that projects data onto lower-dimensional subspaces.
2. Understand the real world applications of projections.
... | github_jupyter |
# APTOS 2019 Blindness Detection
New Idea Ref : https://www.kaggle.com/ratthachat/aptos-simple-preprocessing-decoloring-cropping
## Circle Crop
```
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import collections
import os
from pathlib import Path
from IPython.display i... | github_jupyter |
<h1>Quiz 1 : Pemahaman</h1>
1. Sebutkan apa saja kira2 preprocessing Data?
2. Jelaskan beberapa cara imputing missing value?
3. Kapan kita perlu melakukan feature centering dan scaling?
4. Bagaimana Data Science Workflow?
1. Berikut preprocessing data:
- Binarization
- Mean Removal
- Scaling
- Normalization
- Label e... | github_jupyter |
# Introduction to Random Forest
## Introduction
Random forests (also called random decision forests) construct multiple decision trees at training time. The output of a random forest is often the mode class of individual trees when it is a classification problem, or an average of prediction of individual trees when it... | github_jupyter |
# Midterm Answer Script
**Name**: Ferdous Zeaul Islam
**ID**: 173 1136 042
**Course**: CSE445 (Machine Learning)
**Faculty**: Dr. Sifat Momen (Sfm1)
**Section**: 01
**Semester**: Spring 2021
### N.B- please put the diabetes.csv dataset on the same directory as the ipynb file.
```
# only need th... | github_jupyter |
```
%matplotlib inline
%config InlineBackend.figure_format = "retina"
from matplotlib import rcParams
rcParams["savefig.dpi"] = 100
rcParams["figure.dpi"] = 100
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
session = tf.InteractiveSession()
from exoplanet import transit
T = tf.float64
... | github_jupyter |
## Making decisions with pandas
### quantile analysis: random data
Quintile analysis is a common framework for evaluating the efficacy of security factors
#### What is a factor ?
A factor is a method for scoring/ranking sets of securities. For a particular point in time and for a
particular set of securities, a fac... | github_jupyter |
## 3.4 编辑段落
### 3.4.1 段落首行缩进调整
许多出版社要求文章段落必须首行缩进,若想调整段落首行缩进的距离,可以使用`\setlength{\parindent}{长度}`命令,在`{长度}`处填写需要设置的距离即可。
【**例1**】使用`\setlength{\parindent}{长度}`命令调整段落首行缩进为两字符。
```tex
\documentclass[12pt]{article}
\setlength{\parindent}{2em}
\begin{document}
In \LaTeX, We can use the setlength command to adjust the i... | github_jupyter |
```
from NADINEmainloop import NADINEmain, NADINEmainId
from NADINEbasic import NADINE
from utilsNADINE import dataLoader, plotPerformance
import random
import torch
import numpy as np
# random seed control
np.random.seed(0)
torch.manual_seed(0)
random.seed(0)
# load data
dataStreams = dataLoader('../dataset/pmnist2.ma... | github_jupyter |
# scikit-learn中的逻辑回归
```
import numpy as np
import matplotlib.pyplot as plt
np.random.seed(666)
X = np.random.normal(0, 1, size=(200, 2))
y = np.array((X[:,0]**2+X[:,1])<1.5, dtype='int')
for _ in range(20):
y[np.random.randint(200)] = 1
y
plt.scatter(X[y==0,0], X[y==0,1])
plt.scatter(X[y==1,0], X[y==1,1])
from s... | github_jupyter |
```
!date
```
# Preprocessing FASTQ files into a sample by gene matrix
### Download files and software
```
!git clone https://github.com/pachterlab/BLCSBGLKP_2020.git
!mkdir temporary
!chmod +x BLCSBGLKP_2020/data/kb/parseSS.py
!BLCSBGLKP_2020/data/kb/parseSS.py < BLCSBGLKP_2020/data/kb/SampleSheet.csv > temporary/m... | github_jupyter |
```
import pandas as pd
import numpy as np
#Visualization tools
from matplotlib import pyplot as plt
import seaborn as sns; sns.set()
import itertools
#ML steps structure
from sklearn.pipeline import FeatureUnion, Pipeline
#Preprocessing
from sklearn.preprocessing import FunctionTransformer, MinMaxScaler
from sklear... | github_jupyter |
# Import
```
import gc
import os
import random
import numpy as np
import pandas as pd
```
# Random seed initialize
```
def random_seed_initialize(seed=42):
random.seed(seed)
os.environ['PYTHONHASHSEED'] = str(seed)
np.random.seed(seed)
random_seed_initialize()
```
# Reduce memory Function
```
def redu... | github_jupyter |
```
import os
import numpy
from numpy import arange, sin, exp, pi, diff, floor, asarray
from scipy.io import savemat
from utils.simulation_utils import add_noise, quantize
from utils.gps_l1ca_utils import generate_GPS_L1CA_code
from utils.acquisition_utils import coarse_acquire
from utils.utils import PSKSignal, sample... | github_jupyter |
# SimSwap for videos
Reference: [my changes to the official notebook](https://gist.github.com/woctezuma/78a98b73cbba8cba478d99c8c50bc359)
## Prepare code
```
%cd /content
!git clone https://github.com/woctezuma/SimSwap
%cd /content/SimSwap
!git checkout no-logo
!pip install insightface==0.2.1 onnxruntime moviepy > ... | github_jupyter |
# City street network orientations
Compare the spatial orientations of city street networks with OSMnx.
- [Overview of OSMnx](http://geoffboeing.com/2016/11/osmnx-python-street-networks/)
- [GitHub repo](https://github.com/gboeing/osmnx)
- [Examples, demos, tutorials](https://github.com/gboeing/osmnx-examples)
... | github_jupyter |
# Object Detection Demo
Welcome to the object detection inference walkthrough! This notebook will walk you step by step through the process of using a pre-trained model to detect objects in an image. Make sure to follow the [installation instructions](https://github.com/tensorflow/models/blob/master/research/object_de... | github_jupyter |
# Predicting Boston Housing Prices
## Using XGBoost in SageMaker (Batch Transform)
_Deep Learning Nanodegree Program | Deployment_
---
As an introduction to using SageMaker's High Level Python API we will look at a relatively simple problem. Namely, we will use the [Boston Housing Dataset](https://www.cs.toronto.ed... | github_jupyter |
```
import numpy as np
import pandas as pd
import scipy
import matplotlib.pyplot as plt
from pandas.api.types import CategoricalDtype
from plotnine import *
from scipy.stats import *
import scikit_posthocs as sp
data = pd.read_csv("./NewCols.csv")
dataControl = pd.read_csv("./control.csv")
dataLps= pd.read_csv("./lps... | github_jupyter |
# Desafio 3
Neste desafio, iremos praticar nossos conhecimentos sobre distribuições de probabilidade. Para isso,
dividiremos este desafio em duas partes:
1. A primeira parte contará com 3 questões sobre um *data set* artificial com dados de uma amostra normal e
uma binomial.
2. A segunda parte será sobre a an... | github_jupyter |
# Image classification transfer learning demo
1. [Introduction](#Introduction)
2. [Prerequisites and Preprocessing](#Prequisites-and-Preprocessing)
3. [Fine-tuning the Image classification model](#Fine-tuning-the-Image-classification-model)
4. [Set up hosting for the model](#Set-up-hosting-for-the-model)
1. [Import ... | github_jupyter |
# Concise Implementation of Softmax Regression
:label:`sec_softmax_concise`
Just as high-level APIs of deep learning frameworks
made it much easier
to implement linear regression in :numref:`sec_linear_concise`,
we will find it similarly (or possibly more)
convenient for implementing classification models. Let us stic... | github_jupyter |
# Yelp dataset: CAVI vs noise
Experiments that explore the performance of CAVI at different levels of noise.
```
import collabclass
import json
import matplotlib.pyplot as plt
import numpy as np
```
## Loading & preparing the data
```
classes = (
"AZ",
"NV",
"ON",
"OH",
"NC",
"PA",
"... | github_jupyter |
```
# Source/Reference: https://www.tensorflow.org/tutorials/structured_data/time_series
# Reasoning for explaining TF guides/tutorials:
# You will become comfortable with reading other tutorials/guides on TF2.0/Keras
# Pre-req:
# - LSTMs, RNNs, GRUs chapter
# - Previous code-walkthrough sessions
```
## Dataset
`... | github_jupyter |
```
import geemap
import ee
import os
import geopandas as gpd
import json
import requests
from geemap import geojson_to_ee, ee_to_geojson
from ipyleaflet import GeoJSON
Map = geemap.Map()
Map
# Get ESA World cover
world_cover_collection = ee.ImageCollection("ESA/WorldCover/v100")
world_cover_image = world_cover_collec... | github_jupyter |
```
###########################
# -*- coding: utf-8 -*- #
# PHM_data_challenge_2019 #
# Author: Huet Zhu #
# Date:2019.5 #
# All Rights Reserved #
###########################
# # 基于机器学习的飞控系统故障诊断方案设计
from __future__ import division
import numpy as np
import thundergbm
import pickle
import pand... | github_jupyter |
```
# Copyright 2020 Fagner Cunha
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writ... | github_jupyter |
# Time series forecasting using ARIMA
### Import necessary libraries
```
%matplotlib notebook
import numpy
import pandas
import datetime
import sys
import time
import matplotlib.pyplot as ma
import statsmodels.tsa.seasonal as st
import statsmodels.tsa.arima_model as arima
import statsmodels.tsa.stattools as tools
```... | github_jupyter |
# A Cantera Simulation Using RMG-Py
```
from IPython.display import display, Image
from rmgpy.chemkin import load_chemkin_file
from rmgpy.tools.canteramodel import Cantera, get_rmg_species_from_user_species
from rmgpy.species import Species
```
Load the species and reaction from the RMG-generated chemkin file `chem_... | github_jupyter |
*Poonam Ligade*
*1st Feb 2017*
----------
This notebook is like note to self.
I am trying to understand various components of Artificial Neural Networks aka Deep Learning.
Hope it might be useful for someone else here.
I am designing neural net on MNIST handwritten digits images to identify their correct label ... | github_jupyter |
# Numpy and Matplotlib #
These are two of the most fundamental parts of the scientific python "ecosystem". Most everything else is built on top of them. This is an introduction to python written by [Ryan Abernathy at Columbia University](https://ocean-transport.github.io/index.html) for his module on [Research Computi... | github_jupyter |
# Clustering CIML
Clustering experiment on CIML.
**Motivation:** During CIML supervised learning on multiple classification experiments, where the classes are cloud operators providing the VMs to run CI jobs, the classes predicted with the best metrics were those with the higher amount of samples in the dataset.
We ... | github_jupyter |
<a href="https://colab.research.google.com/github/finerbrighterlighter/myanmar_covid19/blob/master/exponential_growth.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
#Libraries
```
import statsmodels.api as sm
import pandas as pd
import numpy as np... | github_jupyter |
### I. Discover API Documentation
First, let's take a look at the [API Docs](https://www.weatherapi.com/docs/) for the free WeatherAPI.
When performing discovery on an API, look for answers to the following questions:
1. What endpoints are available and what data sources do they offer?
2. How is Authentication perfor... | github_jupyter |
# Using LightGBM as designed (not through sklearn API)
## Automatically Encode Categorical Columns
I've been encoding the geo_level columns as numeric this whole time. Can it perform better by using categorical columns?
LGBM can handle categorical features directly. No need to OHE them. But they must be ints.
1. L... | github_jupyter |
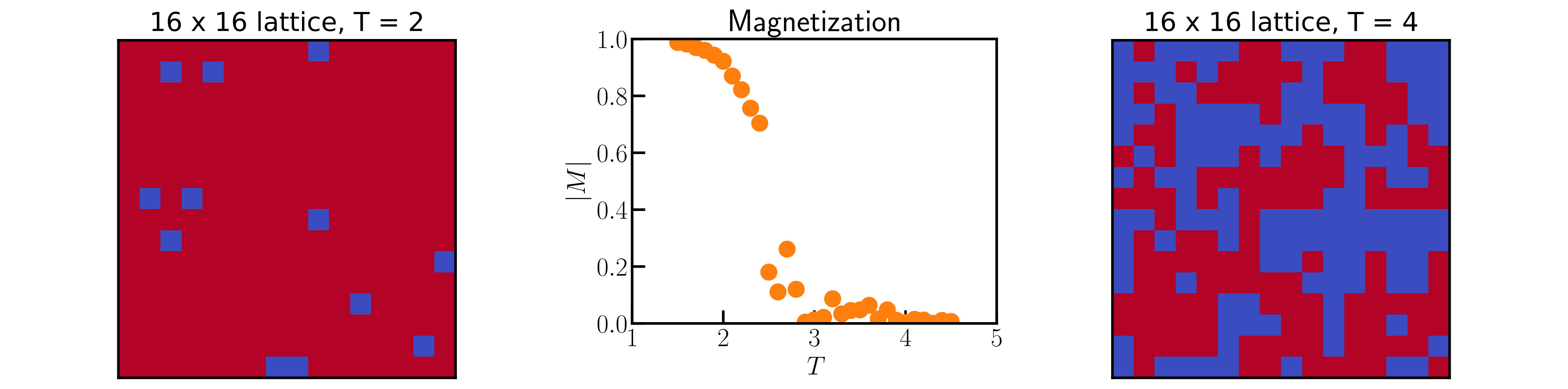
# Monte Carlo simulations of the 2D Ising model
*Authors: Enze Chen (University of California, Berkeley)*
This notebook guides you through the steps of setting up a Monte Carlo (MC) simulation of the 2D ferrom... | github_jupyter |
# Deep Learning & Art: Neural Style Transfer
Welcome to the second assignment of this week. In this assignment, you will learn about Neural Style Transfer. This algorithm was created by Gatys et al. (2015) (https://arxiv.org/abs/1508.06576).
**In this assignment, you will:**
- Implement the neural style transfer alg... | github_jupyter |
# Git
Git is the version control software used in the workshop.
## Installation
- Go to the following link and choose the correct version for your operating system: https://git-scm.com/downloads.
- Following the download, run the installer as per usual on your machine.
- **Windows**: You may leave all selection widg... | github_jupyter |
# Cross Validation - Gabbar
```
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
import warnings
warnings.filterwarnings("ignore")
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns; sns.set_style('ticks')
from sklearn.model_selection import cross_val_score
p... | github_jupyter |
# Modelo lindo de infecção progressiva estocastica
## Introdução
A alguns dias atrás, vi no facebok a seguinte imagem:

<div style="margin-top: 9px; background-color: #efefef; padding-top:10px; padding-bottom:10px;margin-bottom: 9px;box-shadow: 5px 5px 5px 0px rgba(87, 87, 87, 0.2);">
<center>
<h2>Table of Contents</h2>
</center>
<ol>
<li><a href... | github_jupyter |
# Example InfluxDB Jupyter notebook - stream data
This example demonstrates how to query data from InfluxDB 2.0 using Flux and display results in real time.
Prerequisites:
1. Start InfluxDB: `./scripts/influxdb-restart.sh`
2. Start Telegraf: `telegraf -config ./notebooks/telegraf.conf`
3. Install the following depen... | github_jupyter |
# Eel imports
Now let's take a look at a cut of data on eel product imports. The data come from [a foreign trade database maintained by NOAA](https://www.st.nmfs.noaa.gov/commercial-fisheries/foreign-trade/).
The CSV file lives here: `../data/eels.csv`.
We'll start by importing pandas and creating a data frame.
```... | github_jupyter |
# Documentation for the creation and usage of the heat pump library (hplib)
This documentation covers the database preperation, validation and usage for simulation. If you're only interested in using hplib for simulation purpose, you should have a look into chapter [4. How to simulate](#how-to-simulate).
1. [Definiti... | github_jupyter |
# Day 6 - Voronoi diagram
<figure style="float: right; max-width: 25em; margin: 1em">
<img src="https://upload.wikimedia.org/wikipedia/commons/6/6d/Manhattan_Voronoi_Diagram.svg"
alt="Manhattan Voronoi diagram illustration from Wikimedia"/>
<figcaption style="font-style: italic; font-size: smaller">
Manhattan Vo... | github_jupyter |
# 简单理解 SCF 中的 DIIS
> 创建时间:2019-10-23
这篇文档将会简单地叙述 GGA 为代表的 SCF DIIS。
DIIS 是一种 (几乎) 专门用于自洽场过程加速的算法。关于 DIIS 的算法与数学论述,这里并不作展开。这里推荐 C. David Sherrill 的笔记 [^Sherrill-note] 与 Psi4NumPy 的 Jupyter Notebook [^psi4numpy-note] 作为拓展阅读。
这篇笔记会借助 PySCF 的 DIIS 程序,对 Fock 矩阵进行外推。我们将描述在第 $t$ 步 DIIS 过程之后,如何更新第 $t+1$ 步的 Fock 矩阵。我们 **并不会... | github_jupyter |
```
import os
import torch
import torch.nn as nn
import torchvision.models as models
from torchvision import transforms
from PIL import Image
import numpy as np
from tqdm.notebook import tqdm
from sklearn.metrics import confusion_matrix
from IPython.display import display
from src.data_loader import read_dataset, get_... | github_jupyter |
# DO NOT RUN THIS FILE
This file extracts datas from multiple raw XML data files and create json file as needed for model
## Load packages
```
import warnings
warnings.filterwarnings('ignore')
import xml.etree.ElementTree as ET
import sys
import os
import pandas as pd
import json
import xmltodict
import nltk, re
```
... | github_jupyter |

<a href="https://hub.callysto.ca/jupyter/hub/user-redirect/git-pull?repo=https%3A%2F%2Fgithub.com%2Fcallysto%2Fcurriculum-notebooks&branch=master&subPath=Science/ReflectionsOfLightByPlaneAndSph... | github_jupyter |
# 1. Sequence Manager
```
%matplotlib inline
%load_ext autoreload
%autoreload 2
import sys
sys.path.append('../..')
from vimms.SequenceManager import *
data_dir = os.path.join(os.path.abspath(os.path.join(os.path.join(os.getcwd(),".."),"..")),'tests','fixtures')
dataset_file = os.path.join(data_dir, 'QCB_22May19_1.p')... | github_jupyter |
```
seed = 42
import pandas as pd
import numpy as np
import random
np.random.seed(seed)
random.seed(seed)
```
Load the data
```
data = pd.read_csv('data/EEG_data.csv')
data.columns
"""
load subtitle vectors
"""
sub_vec_path = 'data/subtitles/subtitle_vecs.npy'
sub_vecs = np.load(sub_vec_path)
sub_vec_dim = sub_vec... | github_jupyter |
# Neighbors of neighbors
In this notebook, we demonstrate how neighbor-based filters work in the contexts of measurements of cells in tissues. We also determine neighbor of neighbors and extend the radius of such filters.
```
import pyclesperanto_prototype as cle
import numpy as np
import matplotlib
from numpy.random... | github_jupyter |
## Linear Regression with eaget api (立即执行)
```
import matplotlib.pyplot as plt
import numpy as np
import tensorflow as tf
import tensorflow.contrib.eager as tfe
tfe.enable_eager_execution()
# Training Data
train_X = [3.3, 4.4, 5.5, 6.71, 6.93, 4.168, 9.779, 6.182, 7.59, 2.167,
7.042, 10.791, 5.313, 7.997, ... | github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.