code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## This code is a demo for calling the Restartr API.
# ## Importing the required libraries below.
import requests
import json
# ### Format your data below:
# +
job={
"kind" : "test"
"value": "10"
}
r = json.dumps(job)
# -
# ### Example to call the observation api for restartr below:
x = requests.post('https://reccap.cloudapps.unc.edu/api/observation',
headers={"Content-Type": "application/json", "X-API-Key": "<put-api-key-here>"},
data=r)
print("The result of observation api", x.text)
# ### Example to call the query api for restartr below:
y = requests.post('https://reccap.cloudapps.unc.edu/api/query',
headers={"Content-Type": "application/json", "X-API-Key": "<put-api-key-here>"},
data=r)
print("The result of query api", y.text)
# ### Example to query records by "_id" below:
# +
# To query records by "_id" substitute the id below <place-id-here> after calling the observation api
# which will return an id .
# Example: s = {"_id": "5f2c507e8b501271fb7b1d76"}.
# The id above in the example can also be uses to query a record that is already present in the database.
s = {"_id": "<put-id-here>"}
y = requests.post('https://reccap.cloudapps.unc.edu/api/query',
headers={"Content-Type": "application/json", "X-API-Key": "<put-api-key-here>"},
data=json.dumps(s))
print( y.text)
# -
# ### Example to query records by sub-field below:
# +
# To query by sub-field the data will have to be formated as it is below.
# To run a query on existing data in database just use the Example below.
# Using the EXAMPLE below will return all data that has a StudyId of 100160.
# To run a query on a recently sent observation modify job_1 accordingly.
"""EXAMPLE:
job = {
"results.StudyId" : 100160,
"byField": "results.StudyId",
} """
job_1 = {
"results.<put-sub-field-here>" : <put-sub-field-value>,
"byField": "results.<put-sub-field-here>",
}
i = json.dumps(job_1)
y = requests.post('https://reccap.cloudapps.unc.edu/api/query',
headers={"Content-Type": "application/json", "X-API-Key": "<put-api-key-here>"},
data=i)
print("The result of query api", y.text)
# -
|
clinical-demo-notebooks/RestartrApiDemo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python Docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load
import numpy as np
import pandas as pd
import os
import matplotlib
import seaborn as sns
import matplotlib.pyplot as plt
from tqdm import tqdm_notebook
# %matplotlib inline
import cv2 as cv
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
from keras.models import Sequential
from keras.layers import Conv2D,MaxPool2D,DepthwiseConv2D,Flatten,Dense,Dropout,AveragePooling2D,BatchNormalization
from keras.preprocessing.image import ImageDataGenerator
from keras.optimizers import Adam
from keras.utils import to_categorical
from tensorflow.keras import regularizers
# -
from tensorflow.keras.preprocessing.image import ImageDataGenerator
img_height=512
img_width=512
batch_size=128
directory="../input//dl-hack-track-1-cv/train/"
# +
datagen=ImageDataGenerator(
rescale=1.0/255,
# featurewise_center=True,
# featurewise_std_normalization=True,
# rotation_range=20,
# width_shift_range=0.2,
# height_shift_range=0.2,
horizontal_flip=False,
data_format="channels_last",
validation_split=0.2,
dtype=tf.float32
)
# +
train_data=datagen.flow_from_directory(
"../input//dl-hack-track-1-cv/train/",
target_size=(512,512),
batch_size=batch_size,
color_mode="rgb",
class_mode='categorical',
shuffle=True,
subset='training'
)
val_data=datagen.flow_from_directory(
"../input//dl-hack-track-1-cv/train/",
target_size=(512,512),
batch_size=batch_size,
color_mode="rgb",
class_mode='categorical',
shuffle=True,
subset='validation'
)
# +
# model= keras.Sequential()
# # input shape is shape of images
# model.add(keras.Input(shape=(512,512,3)))
# model.add(DepthwiseConv2D(3,(2,2)))
# model.add(BatchNormalization())
# model.add(Conv2D(32,(3,3),activation = 'relu',padding='same'))
# model.add(BatchNormalization())
# model.add(MaxPool2D((2, 2)))
# model.add(Conv2D(48, (3, 3), activation='relu', padding='same'))
# model.add(Conv2D(64, (3, 3), activation='relu', padding='same'))
# model.add(MaxPool2D((2, 2)))
# model.add(Conv2D(80, (3, 3), activation='relu', padding='same'))
# model.add(Conv2D(128, (3, 3), activation='relu', padding='same'))
# model.add(MaxPool2D((2, 2)))
# model.add(Flatten())
# model.add(Dense(64, activation='relu'))
# model.add(Dense(16,activation="relu"))
# model.add(Dense(2,activation="sigmoid"))
# model.compile(
# optimizer=tf.keras.optimizers.Adam(0.00005),
# loss='sparse_categorical_crossentropy',
# metrics=['accuracy']
# )
# model.summary()
# #===========================================================
# batches=1
# q=True
# for x_batch, y_batch in train_data:
# if q:
# model.fit(x_batch, y_batch)
# q=False
# else:
# model.evaluate(x_batch,y_batch)
# q=True
# batches += 1
# if batches >= 70:
# # we need to break the loop by hand because
# # the generator loops indefinitely
# break
# +
def inflow(i):
#x=Conv2D(3,(3,3),padding="same")(i)
x=DepthwiseConv2D((2,2),strides=(2,2))(i)
x=BatchNormalization()(x)
#x=Conv2D(3,(3,3),padding="same",activation="relu")(x)
return x
def outflow(x):
x=Flatten()(x)
x=Dense(64,activation='relu')(x)
x=Dense(16,activation='relu')(x)
x=Dense(2,activation="sigmoid")(x)
return x
vgg19 =keras.applications.VGG16(weights='imagenet',input_shape=(256,256,3), include_top=False)
# for layer in vgg19.layers:
# layer.trainable=False
def midflow(x):
for layer in vgg19.layers:
#layer.trainable = False
x=layer(x)
return x
inputs=keras.Input(shape=(512,512,3))
y=inflow(inputs)
y=midflow(y)
outputs=outflow(y)
Model= keras.Model(inputs,outputs)
Model.summary()
# -
Model.compile(
optimizer=tf.keras.optimizers.Adam(0.000003),
loss='categorical_crossentropy',
metrics=['accuracy']
)
#Model.load_weights("transfer.h5")
Model.fit(train_data,epochs=1,validation_data=val_data)
Model.save("transfer.h5")
"""
Model: "model"
_________________________________________________________________
Layer (type) Output Shape Param #
=================================================================
input_2 (InputLayer) [(None, 512, 512, 3)] 0
_________________________________________________________________
depthwise_conv2d (DepthwiseC (None, 256, 256, 3) 15
_________________________________________________________________
batch_normalization (BatchNo (None, 256, 256, 3) 12
_________________________________________________________________
Vgg-16
_________________________________________________________________
flatten (Flatten) (None, 32768) 0
_________________________________________________________________
dense (Dense) (None, 64) 2097216
_________________________________________________________________
dense_1 (Dense) (None, 16) 1040
_________________________________________________________________
dense_2 (Dense) (None, 2) 34
=================================================================
Total params: 16,813,005
Trainable params: 16,812,999
Non-trainable params: 6
Model.compile(
optimizer=tf.keras.optimizers.Adam(0.000003),
loss='categorical_crossentropy',
metrics=['accuracy']
)
#Model.load_weights("transfer.h5")
Model.fit(train_data,epochs=1,validation_data=val_data)
Model.save("transfer.h5")
"""
# +
def classify(Id):
x=predictwithId(Id)
if x>0.975:
return 0.975
if x<0.025:
return 0.025
return float(x)
def predictwithId(Id):
path='../input/dl-hack-track-1-cv/test/'+str(Id)+".png"
image = tf.keras.preprocessing.image.load_img(path,
grayscale=False,
color_mode="rgb",
target_size=None,
interpolation="nearest")
input_arr = keras.preprocessing.image.img_to_array(image)
input_arr = np.array(input_arr)/255 # Convert single image to a batch.
result=Model.predict(np.reshape(input_arr,(1,512,512,3)))[:,1]
return result
# -
path='../input/dl-hack-track-1-cv/train/fake/'+str(4)+".png"
image = tf.keras.preprocessing.image.load_img(path,
grayscale=False,
color_mode="rgb",
target_size=None,
interpolation="nearest")
input_arr = keras.preprocessing.image.img_to_array(image)
input_arr = np.array(input_arr)/255 # Convert single image to a batch.
result=Model.predict(np.reshape(input_arr,(1,512,512,3)))[:,1]
float(result)
submission=pd.read_csv("../input/dl-hack-track-1-cv/sample_submission.csv")
submission['p_real']=submission['id'].apply(classify)
print(len(submission))
submission.set_index('id').to_csv('sample_submission.csv')
Model.save("transfer.h5")
# +
path='../input/dl-hack-track-1-cv/test/'+str(0)+".png"
image = tf.keras.preprocessing.image.load_img(path,
grayscale=False,
color_mode="rgb",
target_size=None,
interpolation="nearest")
input_arr = keras.preprocessing.image.img_to_array(image)
input_arr = np.array(input_arr)/255 # Convert single image to a batch.
input_arr = np.reshape(input_arr,(1,512,512,3))/255
model2 = tf.keras.models.load_model("m2.h5")
blocks = [7]
outputs = [model2.layers[i].output for i in blocks]
model2 = keras.Model( inputs= model2.inputs, outputs = outputs)
feature_map = model2.predict(input_arr)
for i,fmap in zip(blocks,feature_map):
fig = plt.figure(figsize=(20,15))
#https://stackoverflow.com/a/12444777
fig.suptitle("BLOCK_{}".format(i) , fontsize=20)
for i in range(1,features.shape[3]+1):
plt.subplot(8,8,i)
plt.imshow(fmap[:,:,i-1] , cmap='gray')
#print(fmap[:,:,i].shape)
plt.show()
# +
def inflow(i):
x=Conv2D(3,3,padding="same")(i)
x=DepthwiseConv2D((2,2),strides=(2,2))(x)
return x
vgg_model=keras.applications.VGG16(weights='imagenet', include_top=False, input_shape=(256,256,3))
model_aug=Sequential()
model_aug.add(Conv2D(3,3,padding="same",input_shape=(512,512,3)))
model_aug.add(DepthwiseConv2D((2,2),strides=(2,2)))
model_aug.add(vgg_model)
top_model=Sequential()
top_model.add(Flatten(input_shape=(2, 2, 512)))
#model_aug.add(Dropout(0.3))
top_model.add(Dense(64, activation='relu'))
top_model.add(Dense(1, activation='sigmoid'))
model_aug.add(top_model)
for layer in model_aug.layers[3].layers[:17]:
layer.trainable=False
model
# -
|
dl-hack-track-1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import pandas as pd
import glob
# All files and directories ending with .txt and that don't begin with a dot:
import os
import seaborn as sns
import matplotlib.pyplot as plt
import re
# # !pip install tweet-preprocessor
import preprocessor as p
import spacy
import string
import re
import nltk
from nltk.tokenize import TweetTokenizer, word_tokenize
from nltk.corpus import stopwords
import warnings
from wordcloud import WordCloud, STOPWORDS, ImageColorGenerator
from nltk.sentiment.vader import SentimentIntensityAnalyzer
from collections import Counter
warnings.filterwarnings('ignore')
data2 = pd.read_csv("Clean_1st.csv")
# data2 = data2.sample(n=1000)
# +
#This function is used to tokenize the tweets and remove the stop words which may not be useful for analysis
from nltk.tokenize import TweetTokenizer
nlp=spacy.load('en_core_web_sm')
tokenizer = TweetTokenizer()
data3 = data2[data2["Post Type"] == 'Original']
# data3['clean_soundbite'] = data3['Sound Bite Text'] #***********to delete************
textData = []
for items in data3['clean_soundbite']:
textData.append(items)
token_list = []
tokenizedData = []
for data in textData:
cleaned = [word.lower() for word in tokenizer.tokenize(data.lower()) if word not in nlp.Defaults.stop_words and len(word) > 2]
tokenizedData.append(' '.join(cleaned))
token_list.extend(cleaned)
# -
data3['iphone x'] = 0
data3['iphone 8'] = 0
data3['galaxy'] = 0
data3["tokenizedData"] = tokenizedData
for i in data3['Sound Bite Text'].index:
if re.search(r'iphone[\s]*x', data3['tokenizedData'][i].lower()):
data3['iphone x'][i] = 1
if re.search(r'iphone[\s]*8', data3['tokenizedData'][i].lower()):
data3['iphone 8'][i] = 1
if re.search(r'galaxy[\s]*s8', data3['tokenizedData'][i].lower()):
data3['galaxy'][i] = 1
# +
x = 0 #iphone x only tweets
i8 = 0 #iphone 8 only tweets
g = 0 #galaxy only tweets
for i in data3['tokenizedData'].index:
if data3['iphone x'][i] == 1 and data3['iphone 8'][i] == 0 and data3['galaxy'][i] == 0:
x+=1
if data3['iphone x'][i] == 0 and data3['iphone 8'][i] == 1 and data3['galaxy'][i] == 0:
i8+=1
if data3['iphone x'][i] == 0 and data3['iphone 8'][i] == 0 and data3['galaxy'][i] == 1:
g+=1
print(x, i8, g)
# -
iphone_x = data3[(data3['iphone x'] == 1) & (data3['iphone 8'] == 0) & (data3['galaxy']==0)]
iphone_8 = data3[(data3['iphone x'] == 0) & (data3['iphone 8'] == 1) & (data3['galaxy']==0)]
galaxy = data3[(data3['iphone x'] == 0) & (data3['iphone 8'] == 0) & (data3['galaxy']==1)]
# +
del_words = ["latest", "free", "mobile","iphone", "galaxy", "samsung", "plus", "apple", "new", "phone", "device", "smartphone", "year", "model", "google", "note"]
text = " ".join(str(review) for review in token_list if not any(word in str(review) for word in del_words))
wordcloud = WordCloud(background_color="white", collocations=False, max_words=30).generate(text)
plt.imshow(wordcloud, interpolation='bilinear')
plt.axis("off")
plt.show()
# +
combined_adjs = []
combined_nouns = []
def pos_tag(tokenized):
adjs = []
nouns = []
for tweet in tokenized['tokenizedData']:
for i in nlp(tweet):
if i.pos_ == 'ADJ':
adjs.append(i)
if i.pos_ == 'NOUN':
nouns.append(i)
combined_adjs.append(adjs)
combined_nouns.append(nouns)
combined_Set = [iphone_x, iphone_8, galaxy]
for i in combined_Set:
pos_tag(i)
# -
for i in range(len(combined_adjs)):
text = " ".join(str(review) for review in combined_adjs[i] if not any(word in str(review) for word in del_words))
wordcloud = WordCloud(background_color="white", collocations=False, max_words=25).generate(text)
plt.imshow(wordcloud, interpolation='bilinear')
plt.axis("off")
plt.show()
# +
#preparing data for senti analysis
combined_Set = [iphone_x, iphone_8, galaxy]
combined_corpus = []
for i in combined_Set:
token_data = i['tokenizedData']
corpus = []
for filler in token_data:
corpus.append(nlp(filler))
combined_corpus.append(corpus)
iphone_x["corpus"] = combined_corpus[0]
iphone_8["corpus"] = combined_corpus[1]
galaxy["corpus"] = combined_corpus[2]
# -
iphone_x["corpus"]
# +
#lemmatiztion commented for now. As I think it is not needed in senti analysis
# def lemma(corpus):
# lemmatizedData = []
# for i in range(len(corpus)):
# value =[]
# for data in corpus[i]:
# value.append(("").join(data.lemma_))
# lemmatizedData.append(value)
# return lemmatizedData
# combined_lemma = []
# for i in combined_corpus:
# token_data = i
# combined_lemma.append(lemma(token_data))
# +
#performing sentiment analysis using vader
sentimentAnalyser = SentimentIntensityAnalyzer()
polarity_combined = []
for corpus in combined_corpus:
polarity = []
for values in corpus:
polarity.append(sentimentAnalyser.polarity_scores(values.text))
polarity_combined.append(polarity)
# -
# extracting polarity compund scores
combined_compound = []
for i in polarity_combined:
compoundValues = []
for data in i:
compoundValues.append(data.get("compound"))
combined_compound.append(compoundValues)
# +
for i in range(len(combined_compound)):
plt.figure()
plt.hist(combined_compound[i])
plt.show
# +
#finding frequent words for highly poistive sentiments
pos_sent_combined = []
most_common_combined = []
for i in range(len(combined_compound)):
pos_senti = []
for j in range(len(combined_compound[i])):
if combined_compound[i][j]>0.75:
review = str(combined_corpus[i][j])
# print(review)
pos_senti.append(review)
pos_sent_combined.append((" ".join(pos_senti).split()))
most_common_combined.append(Counter(pos_sent_combined[i]).most_common(10))
# -
# getting top 5 words for each phone
top = 5
top_words_combined = []
for i in range (len(most_common_combined)):
top_words = []
for j in range(top):
word, count = most_common_combined[i][j]
top_words.append(word)
top_words_combined.append(top_words)
# +
#creating dataframes for average polarity score calulation for the top positive words for each mobile
df_iphnx = pd.DataFrame(list(zip(combined_corpus[0], combined_compound[0])), columns =['Review', 'Polarity'])
df_iphn8 = pd.DataFrame(list(zip(combined_corpus[1], combined_compound[1])), columns =['Review', 'Polarity'])
df_galaxy = pd.DataFrame(list(zip(combined_corpus[2], combined_compound[2])), columns =['Review', 'Polarity'])
avgscoring_combined = [df_iphnx, df_iphn8, df_galaxy]
# -
#calculating avergae polarity scores for each of the top words
for num in range(len(avgscoring_combined)):
for i in top_words_combined[num]:
avgscoring_combined[num][i] = 0 #creating a blank column for each of the top words.
for i in avgscoring_combined[num].index: #looping over each datasets index to calucalte mean
for word in top_words_combined[num]:
if word in str(avgscoring_combined[num].loc[i]["Review"]):
avgscoring_combined[num][word].loc[i] = 1 #populating column "word" if word is fiund in record index "i"
# +
phone_order= ["iphone x", "iphone 8", "galaxy"]
for num in range(len(avgscoring_combined)):
for i in top_words_combined[num]:
print("For ", phone_order[num] ," for top word ", i, " the avergae senti score is: ", round(avgscoring_combined[num][avgscoring_combined[num][i]==1]["Polarity"].mean(), 2))
# -
combined_corpus_bk = combined_corpus.copy()
# +
#performing lemmatization for topic modelling
# iphone_x = iphone_x.sample(200)
# iphone_8 = iphone_8.sample(1000)
# galaxy = galaxy.sample(1000)
#took smaller samples for faster processing
#remove combined_corpus_new.
#For final analysis use combined_corpus
import random
combined_corpus_new = [None for _ in range(3)] # for testing to be deleted
combined_corpus_new[0] = random.sample(combined_corpus[0], 200)
combined_corpus_new[1] = random.sample(combined_corpus[1], 2000)
combined_corpus_new[2] = random.sample(combined_corpus[2], 2000)
def lemma(corpus):
lemmatizedData = []
for i in range(len(corpus)):
value =[]
for data in corpus[i]:
value.append(("").join(data.lemma_))
lemmatizedData.append(value)
return lemmatizedData
combined_lemma = []
for i in combined_corpus_new: #change to combined_corpus
token_data = i
combined_lemma.append(lemma(token_data))
# +
#performing topic modeling
import gensim
from gensim.utils import simple_preprocess
import gensim.corpora as corpora
#removing words which appear in topic modelling and are not useful
del_words = [".", "-" "\\n","...", "-", "come", "you", "be", "like", "user", "s8s", "galaxys8", "not"
, "iphone8", "iphone8plus", "samsung", "#", "latest", "free", "mobile","iphone", "galaxy"
, "samsung", "plus", "apple", "new", "phone", "device", "smartphone", "year", "model", "google", "note"]
#https://towardsdatascience.com/end-to-end-topic-modeling-in-python-latent-dirichlet-allocation-lda-35ce4ed6b3e0
#https://github.com/priya-dwivedi/Deep-Learning/blob/master/topic_modeling/LDA_Newsgroup.ipynb
from pprint import pprint
topics_combined = []
for i in range(len(combined_lemma)):
texts = combined_lemma[i]
texts_1=[]
for sent in texts:
comb =[]
text = [word for word in sent if word.lower() not in del_words]
texts_1.append(text)
dictionary = corpora.Dictionary(texts_1)
bow_corpus = [dictionary.doc2bow(text) for text in texts_1]
# number of topics
num_topics = 20
# Build LDA model
lda_model = gensim.models.LdaMulticore(corpus=bow_corpus,
id2word=dictionary,
num_topics=num_topics)
# Print the Keyword in the 10 topics
pprint(lda_model.print_topics())
topics_combined.append(lda_model.print_topics())
# doc_lda = lda_model[bow_corpus]
# -
|
SocialMedia_Analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Reflecting Bear
# ## Background
# Panda Bear is confused. He is trying to work out how things should look when reflected in a mirror, but is getting the wrong results.
# In Bear's coordinates, the mirror lies along the first axis.
# But, as is the way with bears, his coordinate system is not orthonormal: so what he thinks is the direction perpendicular to the mirror isn't actually the direction the mirror reflects in.
# Help Bear write a code that will do his matrix calculations properly!
#
# ## Instructions
# In this assignment you will write a Python function that will produce a transformation matrix for reflecting vectors in an arbitrarily angled mirror.
#
# Building on the last assingment, where you wrote a code to construct an orthonormal basis that spans a set of input vectors, here you will take a matrix which takes simple form in that basis, and transform it into our starting basis.
# Recall the from the last video,
#
# \\( T = E T_E E^{-1} \\)
#
# You will write a function that will construct this matrix.
# This assessment is not conceptually complicated, but will build and test your ability to express mathematical ideas in code.
# As such, your final code submission will be relatively short, but you will receive less structure on how to write it.
#
# ### Matrices in Python
# For this exercise, we shall make use of the @ operator again.
# Recall from the last exercise, we used this operator to take the dot product of vectors.
# In general the operator will combine vectors and/or matrices in the expected linear algebra way,
# i.e. it will be either the vector dot product, matrix multiplication, or matrix operation on a vector, depending on it's input.
# For example to calculate the following expressions,
#
# \\( a = \mathbf{s}\cdot\mathbf{t} \\)
#
# \\( \mathbf{s} = A\mathbf{t} \\)
#
# \\( M = A B \\),
#
# One would use the code,
# ```python
# a = s @ t
# s = A @ t
# M = A @ B
# ```
# (This is in contrast to the \\(*\\) operator, which performs element-wise multiplication, or multiplication by a scalar.)
#
# You may need to use some of the following functions:
# ```python
# inv(A)
# transpose(A)
# gsBasis(A)
# ```
# These, respectively, take the inverse of a matrix, give the transpose of a matrix, and produce a matrix of orthonormal column vectors given a general matrix of column vectors - i.e. perform the Gram-Schmidt process.
# This exercise will require you to combine some of these functions.
#
# ### How to submit
# Edit the code in the cells below to complete the assignment.
# Once you are finished and happy with it, press the *Submit Assignment* button at the top of this notebook.
#
# Please don't change any of the function names, as these will be checked by the grading script.
#
# If you have further questions about submissions or programming assignments, here is a [list](https://www.coursera.org/learn/linear-algebra-machine-learning/discussions/weeks/1/threads/jB4klkn5EeibtBIQyzFmQg) of Q&A. You can also raise an issue on the discussion forum. Good luck!
# PACKAGE
# Run this cell first once to load the dependancies.
import numpy as np
from numpy.linalg import norm, inv
from numpy import transpose
from readonly.bearNecessities import *
# +
# GRADED FUNCTION
# You should edit this cell.
# In this function, you will return the transformation matrix T,
# having built it out of an orthonormal basis set E that you create from Bear's Basis
# and a transformation matrix in the mirror's coordinates TE.
def build_reflection_matrix(bearBasis) : # The parameter bearBasis is a 2×2 matrix that is passed to the function.
# Use the gsBasis function on bearBasis to get the mirror's orthonormal basis.
E = gsBasis(bearBasis)
# Write a matrix in component form that performs the mirror's reflection in the mirror's basis.
# Recall, the mirror operates by negating the last component of a vector.
# Replace a,b,c,d with appropriate values
TE = np.array([[1, 0],
[0, -1]])
# Combine the matrices E and TE to produce your transformation matrix.
T = E @ transpose(TE) @ inv(E)
# Finally, we return the result. There is no need to change this line.
return T
# -
# ## Test your code before submission
# To test the code you've written above, run the cell (select the cell above, then press the play button [ ▶| ] or press shift-enter).
# You can then use the code below to test out your function.
# You don't need to submit this cell; you can edit and run it as much as you like.
#
# The code below will show a picture of Panda Bear.
# If you have correctly implemented the function above, you will also see Bear's reflection in his mirror.
# The orange axes are Bear's basis, and the pink axes are the mirror's orthonormal basis.
# +
# First load Pyplot, a graph plotting library.
# %matplotlib inline
import matplotlib.pyplot as plt
# This is the matrix of Bear's basis vectors.
# (When you've done the exercise once, see what happns when you change Bear's basis.)
bearBasis = np.array( [[1, -1], [1.5, 2]])
# This line uses your code to build a transformation matrix for us to use.
T = build_reflection_matrix(bearBasis)
# Bear is drawn as a set of polygons, the vertices of which are placed as a matrix list of column vectors.
# We have three of these non-square matrix lists: bear_white_fur, bear_black_fur, and bear_face.
# We'll make new lists of vertices by applying the T matrix you've calculated.
reflected_bear_white_fur = T @ bear_white_fur
reflected_bear_black_fur = T @ bear_black_fur
reflected_bear_face = T @ bear_face
# This next line runs a code to set up the graphics environment.
ax = draw_mirror(bearBasis)
# We'll first plot Bear, his white fur, his black fur, and his face.
ax.fill(bear_white_fur[0], bear_white_fur[1], color=bear_white, zorder=1)
ax.fill(bear_black_fur[0], bear_black_fur[1], color=bear_black, zorder=2)
ax.plot(bear_face[0], bear_face[1], color=bear_white, zorder=3)
# Next we'll plot Bear's reflection.
ax.fill(reflected_bear_white_fur[0], reflected_bear_white_fur[1], color=bear_white, zorder=1)
ax.fill(reflected_bear_black_fur[0], reflected_bear_black_fur[1], color=bear_black, zorder=2)
ax.plot(reflected_bear_face[0], reflected_bear_face[1], color=bear_white, zorder=3);
# -
|
assignment-3/reflecting-bear.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="https://cybersecurity-excellence-awards.com/wp-content/uploads/2017/06/366812.png">
# <h1><center>Darwin Supervised Classification Model Building </center></h1>
# # Prior to getting started:
# First,
# <br>if you have just received a new api key from support, you will need to register your key and create a new user (see Register user cell)
#
# Second, in the Environment Variables cell:
# 1. Set your username and password to ensure that you're able to log in successfully
# 2. Set the path to the location of your datasets if you are using your own data. The path is set for the examples.
# 3. Set the dataset names accordingly
#
# Here are a few things to be mindful of:
# 1. For every run, check the job status (i.e. requested, failed, running, completed) and wait for job to complete before proceeding.
# 2. If you're not satisfied with your model and think that Darwin can benefit from extra training, use the resume function.
# ## Set Darwin SDK
from amb_sdk.sdk import DarwinSdk
ds = DarwinSdk()
ds.set_url('https://darwin-api.sparkcognition.com/v1/')
# ## Register user (if needed, read above)
# +
# Use only if you have a new api-key and
# no registered users - fill in the appropriate fields then execute
#Enter your support provided api key and api key password below to register/create new users
api_key = ''
api_key_pw = ''
status, msg = ds.auth_login(api_key_pw, api_key)
if not status:
print(msg)
#Create a new user
status, msg = ds.auth_register_user('username', 'password','<EMAIL>')
if not status:
print(msg)
# -
# ## Environment Variables
# +
#Set your user id and password accordingly
USER="[your Darwin user id]"
PW="[<PASSWORD>]"
# Set path to datasets - The default below assumes Jupyter was started from amb-sdk/examples/Enterprise/
# Modify accordingly if you wish to use your own data
PATH_TO_DATASET='../../sets/'
TRAIN_DATASET='cancer_train.csv'
TEST_DATASET='cancer_test.csv'
# A timestamp is used to create a unique name in the event you execute the workflow multiple times or with
# different datasets. File names must be unique in Darwin.
import datetime
ts = '{:%Y%m%d%H%M%S}'.format(datetime.datetime.now())
# -
# ## Import necessary libraries
# Import necessary libraries
# %matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
from IPython.display import Image
from time import sleep
import os
import numpy as np
from sklearn.metrics import classification_report
# # User Login
status, msg = ds.auth_login_user(USER,PW)
if not status:
print(msg)
# # Data Upload and Clean
# **Read dataset and view a file snippet**
# Preview dataset
df = pd.read_csv(os.path.join(PATH_TO_DATASET, TRAIN_DATASET))
df.head()
# **Upload dataset to Darwin**
# Upload dataset
status, dataset = ds.upload_dataset(os.path.join(PATH_TO_DATASET, TRAIN_DATASET))
if not status:
print(dataset)
# **Clean dataset**
# +
# clean dataset
target = "Diagnosis"
status, job_id = ds.clean_data(TRAIN_DATASET, target = target)
if status:
ds.wait_for_job(job_id['job_name'])
else:
print(job_id)
# -
# # Create and Train Model
# We will now build a model that will learn the class labels in the target column.<br> In the default cancer dataset, the target column is "Diagnosis". <br> You will have to specify your own target name for your custom dataset. <br> You can also increase max_train_time for longer training.
#
model = target + "_model0" + ts
status, job_id = ds.create_model(dataset_names = TRAIN_DATASET, \
model_name = model, \
max_train_time = '00:02')
if status:
ds.wait_for_job(job_id['job_name'])
else:
print(job_id)
# # Extra Training (Optional)
# Run the following cell for extra training, no need to specify parameters
# +
# Train some more
status, job_id = ds.resume_training_model(dataset_names = TRAIN_DATASET,
model_name = model,
max_train_time = '00:05')
if status:
ds.wait_for_job(job_id['job_name'])
else:
print(job_id)
# -
# # Analyze Model
# Analyze model provides feature importance ranked by the model. <br> It indicates a general view of which features pose a bigger impact on the model
# Retrieve feature importance of built model
status, artifact = ds.analyze_model(model)
sleep(1)
if status:
ds.wait_for_job(artifact['job_name'])
else:
print(artifact)
status, feature_importance = ds.download_artifact(artifact['artifact_name'])
# Show the 10 most important features of the model.
feature_importance[:10]
# # Predictions
# **Perform model prediction on the the training dataset.**
status, artifact = ds.run_model(TRAIN_DATASET, model)
sleep(1)
ds.wait_for_job(artifact['job_name'])
# Download predictions from Darwin's server.
status, prediction = ds.download_artifact(artifact['artifact_name'])
prediction.head()
# Create plots comparing predictions with actual target
unq = prediction[target].unique()[::-1]
p = np.zeros((len(prediction),))
a = np.zeros((len(prediction),))
for i,q in enumerate(unq):
p += i*(prediction[target] == q).values
a += i*(df[target] == q).values
#Plot predictions vs actual
plt.plot(a)
plt.plot(p)
plt.legend(['Actual','Predicted'])
plt.yticks([i for i in range(len(unq))],[q for q in unq]);
print(classification_report(df[target], prediction[target]))
# **Perform model prediction on a test dataset that wasn't used in training.** <br>
# Upload test dataset
status, dataset = ds.upload_dataset(os.path.join(PATH_TO_DATASET, TEST_DATASET))
if not status:
print(dataset)
# Clean test dataset
# +
# clean test dataset
status, job_id = ds.clean_data(TEST_DATASET, target = target, model_name = model)
if status:
ds.wait_for_job(job_id['job_name'])
else:
print(job_id)
# -
# Run model on test dataset.
status, artifact = ds.run_model(TEST_DATASET, model)
sleep(1)
ds.wait_for_job(artifact['job_name'])
# Create plots comparing predictions with actual target
# Create plots comparing predictions with actual target
status, prediction = ds.download_artifact(artifact['artifact_name'])
df = pd.read_csv(os.path.join(PATH_TO_DATASET, TEST_DATASET))
unq = prediction[target].unique()[::-1]
p = np.zeros((len(prediction),))
a = np.zeros((len(prediction),))
for i,q in enumerate(unq):
p += i*(prediction[target] == q).values
a += i*(df[target] == q).values
#Plot predictions vs actual
plt.plot(a)
plt.plot(p)
plt.legend(['Actual','Predicted'])
plt.yticks([i for i in range(len(unq))],[q for q in unq]);
print(classification_report(df[target], prediction[target]))
# ## Find out which machine learning model did Darwin use:
status, model_type = ds.lookup_model_name(model)
print(model_type['description']['best_genome'])
|
examples/Enterprise/Darwin Supervised Classification Example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Linear Regression
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
df = pd.read_csv("../../data/USA_Housing.csv")
df.head()
df.info()
df.columns
df.describe()
sns.pairplot(df)
sns.distplot(df.Price)
sns.heatmap(df.corr(), annot=True)
df.columns
X = df[['Avg. Area Income', 'Avg. Area House Age', 'Avg. Area Number of Rooms',
'Avg. Area Number of Bedrooms', 'Area Population']]
y = df['Price']
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.4, random_state=101)
from sklearn.linear_model import LinearRegression
lr = LinearRegression()
lr.fit(X_train, y_train)
print(lr.intercept_)
lr.coef_
X.columns
cdf = pd.DataFrame(lr.coef_, X.columns, )
|
Machine Learning/Linear Regression/Linear Regression-1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %%writefile myfile.txt
hello
this is a text file
myfile = open('myfile.txt')
pwd
myfile = open('myfile.txt')
myfile.read()
myfile.read()
myfile.seek(0)
myfile.read()
myfile.seek(0)
myfile.readlines()
myfile.seek(0)
myfile.readline()
myfile.readline()
myfile.seek(0)
myfile.readlines()
myfile.close()
with open('myfile.txt') as my_newfile:
contents = my_newfile.read()
contents
my_newfile
myfile = open('myfile.txt')
myfile.close()
with open('myfile.txt') as my_newfile:
cont = my_newfile.read()
cont
with open("myfile.txt",mode = 'r') as myfile:
content = myfile.read()
content
with open("myfile.txt",mode = 'w') as myfile:
content = myfile.read()
content
# %%writefile myfile.txt
This IS first line
This IS second line
This is third line
with open('myfile.txt', mode = 'r') as f:
print(f.read())
with open('myfile.txt' , mode = 'a') as f:
f.write('\nthis is 4th line')
with open('myfile.txt', mode = 'r') as f:
print(f.read())
with open('whatever.txt',mode = 'w') as f:
f.write('i just created this file')
with open('whatever.txt',mode = 'r') as f:
print(f.read())
|
IO with basic Files in Pythons.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.3 64-bit
# name: python38364bit1cb406a54609408c902ea62b2dcae9a3
# ---
# Import required modules
# + tags=[]
from datetime import datetime
import helper as hp
import numpy as np
import math
import matplotlib.pyplot as plt
import matplotlib.dates as mdates
# -
# Set Required Base Path (*Note*: relative paths or $Home (~) directory not working)
# +
hp.FIRED_BASE_FOLDER = "/Users/voelkerb/dump/FIRED/"
startDate = "2020.06.13" # None to select dataset first day
stopDate = "2020.07.16" # None to select dataset last day (timepoint)
# Get recording range from files and args
start, end = hp.getRecordingRange(startDate, stopDate)
measure = "p" # p,q,s
smoothing = 3
# -
# Select some appliances you are interested in
# + tags=[]
includeAppliances = ["kettle", "espresso machine", "stove", "television", "office pc", "router #1", "hairdryer"]
# Get list of all appliances in dataset
appliances = hp.getApplianceList()
# If specific appliances should be selected, based on script arguments
if len(includeAppliances) > 0: appliances = [a for a in includeAppliances if a in appliances]
print(appliances)
# -
# Load 1 Hz data of these appliances
# + tags=[]
# Load power of these appliances from 1Hz file (should be sufficient for histogram)
powers = hp.getPowerForAppliances(1, startTs=start, stopTs=end, appliances=appliances)
# Remove power which could not be extracted
for p,a in zip(powers, appliances):
if p is None: print("Could not extract power of appliance: " + str(a))
powers = [p for p in powers if p is not None]
# -
# Power histogram (distributuon of amount of power drawn by device).
# The OFF state is removed from the data by defining a threshold. Bin size represents the number of bins used for
# the histogram plotted. The more bins, the smoother the histogram.
# + tags=[]
off_threshold = 5.0
binsize = 200
# Calculate X,Y subplot grid
subplotsX = min(3, int(len(powers)))
subplotsY = max(1, int(len(powers)/3) + 1)
if len(powers)%3 == 0: subplotsY -= 1
# Init figure
fig, axes = plt.subplots(subplotsY,subplotsX, tight_layout=True)
# Flatten axis list
if subplotsY > 1: axes = [a for ax in axes for a in ax]
# Add data to each figure
for axis, power in zip(axes, powers):
axis.set_title(hp.prettyfyApplianceName(power["name"]))
data = power["data"][measure]
if smoothing > 1:
data = np.median(data[0:int(len(data)/smoothing)*smoothing].reshape(-1, smoothing), axis=1)
data = data[data > off_threshold]
hist, bins = np.histogram(data, binsize)
# thres = 0.0005*np.sum(hist)
# hist[hist < thres] = 0
# width = 1.0 * (bins[1] - bins[0])
# maxV = max([v for v,h in zip(bins,hist) if h > 0])
# minV = min([v for v,h in zip(bins,hist) if h > 0]) - width
center = (bins[:-1] + bins[1:]) * 0.5
axis.fill_between(center, hist, step="pre")
# axis.set_xlim([minV, maxV])
#axis.step(center, hist, align="center", width=width)
#axis.hist(data, bins=binsize) #, histtype="stepfilled")
axis.set_yticks([])
# Delete unused grid axes
for ax in axes[len(powers):]: fig.delaxes(ax)
# -
# Distribution of usage over time.
# + tags=[]
# Init figure
fig, axes = plt.subplots(subplotsY, subplotsX, tight_layout=True)
# Flatten axis list
if subplotsY > 1: axes = [a for ax in axes for a in ax]
startDate = datetime.fromtimestamp(start)
startDay = startDate.replace(hour=0, minute=0, second=0, microsecond=0)
if startDate != startDay: startDay = startDay + timedelta(days=1)
stopDay = datetime.fromtimestamp(end).replace(hour=0, minute=0, second=0, microsecond=0)
sampleStart = int(startDay.timestamp()-start)
sampleStop = int(stopDay.timestamp()-start)
# Add data to each figure
for axis, power in zip(axes, powers):
axis.set_title(hp.prettyfyApplianceName(power["name"]))
data = np.abs(power["data"][measure])[sampleStart:sampleStop]
# For data with some missing samples or too much samples
# This can happen for non finished days
days = int(len(data)/(24*60*60))
# e = days*(24*60*60)
# if len(data) > e: data = data[:e]
# elif len(data) < e: data = npconcatenate(data, np.zeros(e-len(data)))
# Split data in each day
data = data.reshape((days, 24, 60*60))
# mean for each hour
data = np.mean(data, axis=2)
# mean for each hour a day
data = np.mean(data, axis=0)
axis.bar(np.arange(0, 24, 1), height=data, align='edge', width=1.0)#
axis.set_xlim(0, 24)
axis.set_xticks(np.arange(0, 25, 4))
# Delete unused grid axes
for ax in axes[len(powers):]: fig.delaxes(ax)
# -
|
notebooks/applianceUsage.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Load
# ### Install Packages, load data
# +
# Install Kaggle from PIP
# ! pip install kaggle
# Download the data via API
# ! kaggle competitions download -c forest-cover-type-prediction
# +
# Import Packages
import kaggle
import numpy as np
import pandas as pd
np.random.seed(0)
# +
# Import Train and Test data from Kaggle
train_kaggle = pd.read_csv('../../data/raw/forest-cover-type-prediction/train.csv')
test_kaggle = pd.read_csv('../../data/raw/forest-cover-type-prediction/test.csv')
# Shuffle the data
# shuffle = np.random.permutation(np.arange(train_kaggle.shape[0]))
train_kaggle = train_kaggle.sample(frac = 1)
# Separate in to train/dev sets
train_pct = .5 # .8 for 80/20 split
split = int(train_kaggle.shape[0] * train_pct)
train_data = train_kaggle.iloc[:split,:-1].set_index('Id')
train_labels = train_kaggle.iloc[:split,].loc[:, ['Id', 'Cover_Type']].set_index('Id')
dev_data = train_kaggle.iloc[split:,:-1].loc[:,].set_index('Id')
dev_labels = train_kaggle.iloc[split:,].loc[:, ['Id', 'Cover_Type']].set_index('Id')
# -
print(train_data.shape)
print(dev_data.shape)
print(train_labels.shape)
print(dev_labels.shape)
# Write data to dataframes
train_data.to_csv('../../data/processed/train_data.csv')
train_labels.to_csv('../../data/processed/train_labels.csv')
dev_data.to_csv('../../data/processed/dev_data.csv')
dev_labels.to_csv('../../data/processed/dev_labels.csv')
|
src/data/.ipynb_checkpoints/data_load-checkpoint.ipynb
|
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .cs
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: .NET (C#)
// language: C#
// name: .net-csharp
// ---
#r "nuget:MNCD,1.0.0"
// +
using MNCD.Core;
using MNCD.Writers;
using System.Net.Http;
using Newtonsoft.Json;
public void VisualizeCommunities(Network network, List<Community> communities)
{
var writer = new EdgeListWriter();
var edge_list = writer.ToString(network, true);
var communityWriter = new ActorCommunityListWriter();
var community_list = communityWriter.ToString(network.Actors, communities, true);
var client = new HttpClient();
var uri = "https://mncd-viz.azurewebsites.net/api/multi-layer/slices-communities";
var body = new
{
edge_list = edge_list,
community_list = community_list,
image_format = "svg"
};
var json = JsonConvert.SerializeObject(body);
var content = new StringContent(json);
var result = client.PostAsync(uri, content).Result;
var svg = result.Content.ReadAsStringAsync().Result;
display(HTML(svg));
}
// +
using MNCD.Core;
// L1 L2
// 0 4
// | \ L1-L2 / |
// | 2 ----- 3 |
// | / \ |
// 1 5
var a = new List<Actor>
{
new Actor("a0"),
new Actor("a1"),
new Actor("a2"),
new Actor("a3"),
new Actor("a4"),
new Actor("a5"),
};
var e0 = new List<Edge>
{
new Edge(a[0], a[1]),
new Edge(a[0], a[2]),
new Edge(a[1], a[2]),
};
var e1 = new List<Edge>
{
new Edge(a[3], a[4]),
new Edge(a[3], a[5]),
new Edge(a[4], a[5])
};
var l0 = new Layer(e0) { Name = "Layer_0" };
var l1 = new Layer(e1) { Name = "Layer_1" };
var l = new List<Layer> { l0, l1 };
var i = new List<InterLayerEdge>
{
new InterLayerEdge(a[2], l0, a[3], l1)
};
var network = new Network(l, a)
{
InterLayerEdges = i
};
// -
// # CLECC Community Detection
// +
using MNCD.CommunityDetection.MultiLayer;
var communities = new CLECCCommunityDetection().Apply(network, 1, 2);
display(communities);
VisualizeCommunities(network, communities);
// -
// # ABACUS
// +
using MNCD.CommunityDetection.MultiLayer;
using MNCD.CommunityDetection.SingleLayer;
var communities = new ABACUS().Apply(network, n => new Louvain().Apply(n), 2);
display(communities);
VisualizeCommunities(network, communities);
|
examples/multi-layer.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # TP3 Kernel Methods for Machine Learning
# setup
import numpy as np
from sklearn import linear_model as lm
import sys
print(sys.version)
import sklearn
sklearn.__version__
# ## Tasks
# 1. Implement (naive) solvers to Ridge Regression, Weighted Ridge Regression and Logistic Ridge Regression (using Iteratively Reweighted Least Squares). See notes for the mathematical derivation.
# 2. Simulate some toy data to check if our solvers give correct solutions as provided by R.
# ## Solutions
# **Ridge Regression (RR)**
#
# Given $X \in \mathbb{R}^{n \times p}$ and $y \in \mathbb{R}^n$, solve
# $$
# \min_{\beta \in \mathbb{R}^p} \frac{1}{n} \|y - X \beta\|^2 + \lambda \|\beta\|^2 \,.
# $$
# Ridge Regression (RR)
def solveRR(y, X, lam):
n,p = X.shape
assert (len(y) == n)
A = X.T @ X
# Adjust diagonal due to Ridge
A[np.diag_indices_from(A)] += lam*n
b = X.T.dot(y)
# Find solution to the linear system Ax = b
beta = np.linalg.solve(A,b)
return (beta)
# **Weighted Ridge Regression (WRR)**
#
# Given $X \in \mathbb{R}^{n \times p}$ and $y \in \mathbb{R}^n$, and weights $w \in \mathbb{R}^n_+$, solve
# $$
# \min_{\beta \in \mathbb{R}^p} \frac{1}{n} \sum_{i=1}^n w_i (y_i - \beta^\top x_i)^2 + \lambda \|\beta\|^2 \,.
# $$
# Weighted Ridge Regression (WRR)
def solveWRR(y, X, w, lam):
y1 = np.sqrt(w)*y
X1 = (np.sqrt(w)*X.T).T
beta = solveRR(y1, X1, lam)
return (beta)
# **Logistic Ridge Regression (LRR)**
#
# Given $X \in \mathbb{R}^{n \times p}$ and $y \in \{-1,+1\}^n$, solve
# $$
# \min_{\beta \in \mathbb{R}^p} \frac{1}{n} \sum_{i=1}^n \log (1+e^{-y_i \beta^\top x_i}) + \lambda \|\beta\|^2 \,.
# $$
# Logistic Ridge Regression (LRR)
def solveLRR(y, X, lam):
# Parameters
L = 100
eps = 1e-3
sigmoid = lambda a: 1/(1+np.exp(-a))
n,p = X.shape
# Initialize
beta = np.zeros(p)
# Update (equiv to IRLS)
for k in range(L):
beta_old = beta
f = X.dot(beta_old)
w = sigmoid(f) * sigmoid(-f)
z = f + y / sigmoid(y*f)
beta = solveWRR(z, X, w, 2*lam)
if np.sum((beta-beta_old)**2) < eps:
break
return (beta)
# **Toy experiments**
# +
# Toy data
np.random.seed(12345)
n = 100
p = 20
X = np.random.normal(0,1,(n,p))
X = sklearn.preprocessing.scale(X)
y = np.sign(np.random.normal(0,1,n))
lam = 0.01
# Our solver
beta1 = solveRR(y, X, lam) # RR
# beta1 = solveLRR(y, X, lam) # LRR
# print(beta1)
# Python solver
beta2 = lm.Ridge(alpha=lam*n,fit_intercept=False,normalize=False).fit(X, y).coef_ # RR
# beta2 = lm.RidgeClassifier(alpha=2*n*lam,fit_intercept=False,normalize=False).fit(X, y).coef_ # LRR gives different results?
# print(beta2)
# Check
np.sum((beta1-beta2)**2)
|
Kernel-Method-for-ML-Codes/.ipynb_checkpoints/TP3-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
import pandas as pd
tip = pd.read_csv('../../../data/interim/US_cities_only/tip_US.csv')
tip.head()
# +
# Cleaning 'date' column
tip['date'] = pd.to_datetime(tip['date'])
# -
tip.info()
tip.describe()
# +
# Writing clean 'tip' dataframe to csv
tip.to_csv('../../../data/interim/clean_US_cities/tip_clean.csv', encoding='utf-8', index=False)
|
notebooks/exploratory/1-clean/1.5-clean-tip-data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (herschelhelp_internal)
# language: python
# name: helpint
# ---
# # EGS master catalogue
# ## Preparation of Pan-STARRS1 - 3pi Steradian Survey (3SS) data
#
# This catalogue comes from `dmu0_PanSTARRS1-3SS`.
#
# In the catalogue, we keep:
#
# - The `uniquePspsSTid` as unique object identifier;
# - The r-band position which is given for all the sources;
# - The grizy `<band>FApMag` aperture magnitude (see below);
# - The grizy `<band>FKronMag` as total magnitude.
#
# The Pan-STARRS1-3SS catalogue provides for each band an aperture magnitude defined as “In PS1, an 'optimal' aperture radius is determined based on the local PSF. The wings of the same analytic PSF are then used to extrapolate the flux measured inside this aperture to a 'total' flux.”
#
# The observations used for the catalogue where done between 2010 and 2015 ([ref](https://confluence.stsci.edu/display/PANSTARRS/PS1+Image+data+products)).
#
# **TODO**: Check if the detection flag can be used to know in which bands an object was detected to construct the coverage maps.
#
# **TODO**: Check for stellarity.
from herschelhelp_internal import git_version
print("This notebook was run with herschelhelp_internal version: \n{}".format(git_version()))
import datetime
print("This notebook was executed on: \n{}".format(datetime.datetime.now()))
# +
# %matplotlib inline
# #%config InlineBackend.figure_format = 'svg'
import matplotlib.pyplot as plt
plt.rc('figure', figsize=(10, 6))
from collections import OrderedDict
import os
from astropy import units as u
from astropy.coordinates import SkyCoord
from astropy.table import Column, Table
import numpy as np
from herschelhelp_internal.flagging import gaia_flag_column
from herschelhelp_internal.masterlist import nb_astcor_diag_plot, remove_duplicates
from herschelhelp_internal.utils import astrometric_correction, mag_to_flux
# +
OUT_DIR = os.environ.get('TMP_DIR', "./data_tmp")
try:
os.makedirs(OUT_DIR)
except FileExistsError:
pass
RA_COL = "ps1_ra"
DEC_COL = "ps1_dec"
# -
# ## I - Column selection
# +
imported_columns = OrderedDict({
"objID": "ps1_id",
"raMean": "ps1_ra",
"decMean": "ps1_dec",
"gFApMag": "m_ap_gpc1_g",
"gFApMagErr": "merr_ap_gpc1_g",
"gFKronMag": "m_gpc1_g",
"gFKronMagErr": "merr_gpc1_g",
"rFApMag": "m_ap_gpc1_r",
"rFApMagErr": "merr_ap_gpc1_r",
"rFKronMag": "m_gpc1_r",
"rFKronMagErr": "merr_gpc1_r",
"iFApMag": "m_ap_gpc1_i",
"iFApMagErr": "merr_ap_gpc1_i",
"iFKronMag": "m_gpc1_i",
"iFKronMagErr": "merr_gpc1_i",
"zFApMag": "m_ap_gpc1_z",
"zFApMagErr": "merr_ap_gpc1_z",
"zFKronMag": "m_gpc1_z",
"zFKronMagErr": "merr_gpc1_z",
"yFApMag": "m_ap_gpc1_y",
"yFApMagErr": "merr_ap_gpc1_y",
"yFKronMag": "m_gpc1_y",
"yFKronMagErr": "merr_gpc1_y"
})
catalogue = Table.read("../../dmu0/dmu0_PanSTARRS1-3SS/data/PanSTARRS1-3SS_EGS_v2.fits")[list(imported_columns)]
for column in imported_columns:
catalogue[column].name = imported_columns[column]
epoch = 2012
# Clean table metadata
catalogue.meta = None
# +
# Adding flux and band-flag columns
for col in catalogue.colnames:
if col.startswith('m_'):
errcol = "merr{}".format(col[1:])
# -999 is used for missing values
catalogue[col][catalogue[col] < -900] = np.nan
catalogue[errcol][catalogue[errcol] < -900] = np.nan
flux, error = mag_to_flux(np.array(catalogue[col]), np.array(catalogue[errcol]))
# Fluxes are added in µJy
catalogue.add_column(Column(flux * 1.e6, name="f{}".format(col[1:])))
catalogue.add_column(Column(error * 1.e6, name="f{}".format(errcol[1:])))
# Band-flag column
if "ap" not in col:
catalogue.add_column(Column(np.zeros(len(catalogue), dtype=bool), name="flag{}".format(col[1:])))
# TODO: Set to True the flag columns for fluxes that should not be used for SED fitting.
# -
catalogue[:10].show_in_notebook()
# ## II - Removal of duplicated sources
# We remove duplicated objects from the input catalogues.
# +
SORT_COLS = ['merr_ap_gpc1_r', 'merr_ap_gpc1_g', 'merr_ap_gpc1_i', 'merr_ap_gpc1_z', 'merr_ap_gpc1_y']
FLAG_NAME = 'ps1_flag_cleaned'
nb_orig_sources = len(catalogue)
catalogue = remove_duplicates(catalogue, RA_COL, DEC_COL, sort_col=SORT_COLS, flag_name=FLAG_NAME)
nb_sources = len(catalogue)
print("The initial catalogue had {} sources.".format(nb_orig_sources))
print("The cleaned catalogue has {} sources ({} removed).".format(nb_sources, nb_orig_sources - nb_sources))
print("The cleaned catalogue has {} sources flagged as having been cleaned".format(np.sum(catalogue[FLAG_NAME])))
# -
# ## III - Astrometry correction
#
# We match the astrometry to the Gaia one. We limit the Gaia catalogue to sources with a g band flux between the 30th and the 70th percentile. Some quick tests show that this give the lower dispersion in the results.
gaia = Table.read("../../dmu0/dmu0_GAIA/data/GAIA_EGS.fits")
gaia_coords = SkyCoord(gaia['ra'], gaia['dec'])
catalogue[RA_COL].unit = u.deg
catalogue[DEC_COL].unit = u.deg
nb_astcor_diag_plot(catalogue[RA_COL], catalogue[DEC_COL],
gaia_coords.ra, gaia_coords.dec)
# +
delta_ra, delta_dec = astrometric_correction(
SkyCoord(catalogue[RA_COL], catalogue[DEC_COL]),
gaia_coords
)
print("RA correction: {}".format(delta_ra))
print("Dec correction: {}".format(delta_dec))
# -
catalogue[RA_COL] += delta_ra.to(u.deg)
catalogue[DEC_COL] += delta_dec.to(u.deg)
nb_astcor_diag_plot(catalogue[RA_COL], catalogue[DEC_COL],
gaia_coords.ra, gaia_coords.dec)
# ## IV - Flagging Gaia objects
catalogue.add_column(
gaia_flag_column(SkyCoord(catalogue[RA_COL], catalogue[DEC_COL]), epoch, gaia)
)
# +
GAIA_FLAG_NAME = "ps1_flag_gaia"
catalogue['flag_gaia'].name = GAIA_FLAG_NAME
print("{} sources flagged.".format(np.sum(catalogue[GAIA_FLAG_NAME] > 0)))
# -
# # V - Saving to disk
catalogue.write("{}/PS1.fits".format(OUT_DIR), overwrite=True)
|
dmu1/dmu1_ml_EGS/1.10_PanSTARRS1-3SS.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/pliniodester/project_euler/blob/main/001.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="EBQiBCBMPq0i"
# ### **Problem 1 - Multiples of $3$ or $5$**
# If we list all the natural numbers below $10$ that are multiples of $3$ or $5$, we get $3$, $5$, $6$ and $9$. The sum of these multiples is $23$.
#
# Find the sum of all the multiples of $3$ or $5$ below $1000$.
# + colab={"base_uri": "https://localhost:8080/"} id="MFmgLJTZy94S" outputId="3872f220-87ac-475e-9c62-973f93503f0e"
N = 1000
sum = 0
for k in range(N):
if (k%3==0) | (k%5==0): # verify if k is divisible by 3 or 5
sum+=k
print(sum)
|
001.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Titanic Survival Machine Learning Model
#
# This notebook contains a **quick** attempt at building a machine learning model for predicting the surival of a passenger on the ship [Titanic](https://en.wikipedia.org/wiki/RMS_Titanic), based on the dataset hosted at [Kaggle](https://www.kaggle.com). The ultimate aim of this work is to demonstrate an archetypal 'data science workflow' that includes some data exploration, feature engineering, model training and selection and to yield a model that can be used in other projects downstream. We have already downloaded the data from Kaggle, in CSV format, to the `data` directory in this project's root directory.
# **WorkSheet done in DataLore**
# Link:- https://datalore.jetbrains.com/notebook/PaaYOjq7apUBBrizLygpah/FOl7hCaUmyt6V8leyl6ArH/
#
# ## Package Imports and Environment Setup
# +
import warnings
from datetime import datetime
from typing import Any, Callable, Dict, Iterable
import joblib
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
from seaborn import boxplot, countplot
from pandas import DataFrame
from numpy import ndarray
from sklearn.base import BaseEstimator
from sklearn.compose import make_column_transformer
from sklearn.ensemble import (
GradientBoostingClassifier, RandomForestClassifier, VotingClassifier)
from sklearn.exceptions import NotFittedError
from sklearn.impute import SimpleImputer
from sklearn.metrics import accuracy_score, roc_auc_score, make_scorer
from sklearn.model_selection import (
BaseCrossValidator, GridSearchCV, StratifiedKFold, train_test_split)
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import FunctionTransformer
from sklearn.preprocessing import OneHotEncoder
from sklearn.preprocessing import StandardScaler
from sklearn.svm import SVC
warnings.filterwarnings('ignore')
sns.set()
# -
# ## Load Dataset
data = pd.read_csv('/Users/Asus/Documents/Atmel Studio/MLtrain.csv')
data.info()
# ## Summary Statistics for Numeric Data
(data
.drop(['PassengerId'], axis=1)
.describe())
# ## Data Description
#
# - `Survived` - Survival (0 = No, 1 = Yes);
# - `Pclass` - Ticket class (1 = 1st, 2 = 2nd, 3 = 3rd);
# - `Sex` - sex (male/female);
# - `SibSp` - # of siblings / spouses aboard the Titanic (positive integer);
# - `Parch` - # of parents / children aboard the Titanic (positive integer);
# - `Ticket` - Ticket number (positive integer);
# - `Fare` - Passenger fare (positive real);
# - `Cabin` - Cabin number (string); and,
# - `Embarked` - Port of Embarkation (C = Cherbourg, Q = Queenstown, S = Southampton).
#
# Note, that some children travelled only with a nanny, therefore `parch=0` for them.
data.head()
# ## Data Visualisation
#
# We introduce a simple class for automating the creation of basic visualisations that are suitable for a binary classification task and which treat continuous and non-continuous features differently.
class VizFeatureBinaryClass:
"""Automate binary classifier feature visualisation."""
def __init__(self, label_col: str, feature_cols: Iterable[str],
data: DataFrame) -> None:
plot_data = data[feature_cols + [label_col]]
self.plots = {}
for feature in feature_cols:
fig, ax = plt.subplots()
plt.close(fig)
if plot_data[feature].dtype == 'float64':
plot = boxplot(x=label_col, y=feature, data=plot_data, ax=ax)
else:
plot = countplot(x=feature, hue=label_col, data=plot_data,
ax=ax)
plot.set_title(feature)
self.plots[feature] = plot
def show(self, feature: str) -> None:
"""Display single plot."""
plot = self.plots[feature]
display(plot.figure)
return None
def show_all(self) -> None:
"""Display all plots."""
for plot in self.plots.values():
display(plot.figure)
return None
# We now use the visualiation class to take a cursory look at the impact of each feature on class label assignment.
# +
viz_features = VizFeatureBinaryClass(
'Survived', ['Sex', 'Age', 'SibSp', 'Parch', 'Fare', 'Embarked'], data)
viz_features.show_all()
# -
# ## Extract Class Labels
#
# The class labels for this classification task are in the `Survived` column of the training data. We split this column into seperate vector for model training.
labels = data.Survived.values
# Take a look at the number of observations assigned to each class (i.e. look for the extent of the class imbalance).
survived_fraction = labels.sum() / labels.shape[0]
print(f'{survived_fraction:.1%} of passengers are classified as surviving')
# ## Spit Dataset into Train and Test Subsets
#
# Use stratified sampling to ensure that classes are fairly represented in both train and test data.
train_data, test_data, y_train, y_test = train_test_split(
data,
labels,
random_state=42,
test_size=0.1,
stratify=labels)
# ## Feature Engineering
#
# We will **drop** the following columns from the final set of features:
#
# - `PassengerId` - no apparent information content;
# - `Survived` - i.e. the class labels we wish to predict;
# - `Name` - no apparent predictive information content;
# - `Ticket` - no apparent information content; and,
# - `Cabin` - too many missing values to work with and `Pclass` and `Fare` should be adequate descriptors of cabin.
#
# We will need to infer the missing values for the following columns (plus label each observation as having missing data):
#
# - `Age` - using `Sex` and whether or not child or adult as inferred from `Name` and `Parch`; and,
# - `Embarked` - assign to a new category labelled as `UNKNOWN`.
#
# We will one-hot-encode the following categorical variables:
#
# - `Pclass`;
# - `Sex`; and,
# - `Embarked`.
# ### Missing Data Imputers
age_imputer = SimpleImputer(strategy='mean')
embarked_imputer = SimpleImputer(strategy='constant', fill_value='UNKNOWN')
# ### Configure and Fit Feature Engineering Pipeline
# +
feature_pipeline = make_column_transformer(
(make_pipeline(age_imputer, StandardScaler()), ['Age']),
(make_pipeline(embarked_imputer, OneHotEncoder()), ['Embarked']),
(StandardScaler(), ['Fare']),
(OneHotEncoder(), ['Pclass', 'Sex']))
feature_pipeline.fit(train_data);
# -
# ### Transform Train and Test Data
# +
X_train = feature_pipeline.transform(train_data)
X_test = feature_pipeline.transform(test_data)
X_train
# -
# ## Model Selection
#
# We are fairly certain that the final decision boundary needs to be non-linear and that model training will benefit from models that can use weighted loss function. With this in mind, we will test the following model classes:
#
# - Support Vector Machines (SVM);
# - Random Forests (RF);
# - Gradient Boosting Machine (GBM); and,
# - Voting Classifier based on combining the above models.
# ### Experimental Setup
#
# We will use 5-fold cross validation for hyper-parameter tuning and then asses performance on the test data.
cross_validator = StratifiedKFold(n_splits=5, random_state=42, shuffle=True)
# ### Test Metrics
#
# We will use the following metrics to assess model performance:
#
# - Area Under ROC Curve (AUC); and,
# - accuracy.
metrics = {'auc': roc_auc_score,
'accuray': accuracy_score}
# ### Automation
#
# We define a simple class for automating grid-search, cross-validation and metric calculation across all model types.
# +
MetricScorer = Callable[[Iterable[int], Iterable[int]], float]
class MLExperimentRunner:
"""Model training and metric calculation automation."""
def __init__(self,
metrics: Dict[str, MetricScorer],
cross_validator: BaseCrossValidator,
model_selection_metric: str,
X_train: ndarray,
y_train: ndarray,
X_test: ndarray,
y_test: ndarray,) -> None:
self.metrics = metrics
self.cv = cross_validator
self.selection_metric = model_selection_metric
self.X_train = X_train
self.y_train = y_train
self.X_test = X_test
self.y_test = y_test
return None
def run(self, estimator: BaseEstimator,
param_grid: Dict[str, Any], n_results: int = 10) -> BaseEstimator:
"""Run an experiment."""
scorers = {name: make_scorer(metric)
for name, metric in self.metrics.items()}
experiment = GridSearchCV(
estimator, param_grid, scoring=scorers, cv=self.cv,
refit=self.selection_metric, n_jobs=2)
experiment.fit(self.X_train, self.y_train)
self._display_results(experiment.cv_results_, n_results)
self._display_test_metrics(experiment.best_estimator_)
return experiment.best_estimator_
def _display_test_metrics(self, estimator: BaseEstimator) -> None:
"""Print metrics for test data to stdout."""
predictions = estimator.predict(self.X_test)
results = {
metric_name: metric_func(predictions, self.y_test)
for metric_name, metric_func in self.metrics.items()}
print('----------------------------------------')
print('-- TEST DATA METRICS (best estimator) --')
print('---------------------------------------')
for metric_name, metric_value in results.items():
print(f'{metric_name}: {metric_value:.4f}')
print('----------------------------------------')
return None
def _display_results(self, cv_results: Dict[str, Any],
n_results: int) -> None:
"""Display cross validation results as DataFrame."""
cv_data = DataFrame(cv_results)
metric_col_names = [
e
for name in self.metrics.keys()
for e in [f'mean_test_{name}', f'std_test_{name}']]
selection_metric_col_name = f'mean_test_{self.selection_metric}'
cv_data.sort_values(
by=selection_metric_col_name, ascending=False, inplace=True)
cv_data.reset_index(inplace=True)
display(cv_data[['params'] + metric_col_names].head(n_results))
return None
# -
# And we then generate an instance of the experiment runner for the current task.
ml_experiment = MLExperimentRunner(
metrics, cross_validator, 'auc', X_train, y_train, X_test, y_test)
# ### Support Vector Machine
svm_model = ml_experiment.run(
SVC(class_weight='balanced', gamma='auto', probability=True),
{'C': [0.01, 0.1, 1, 10, 100],
'kernel': ['linear', 'rbf']})
# ### Random Forest
rf_model = ml_experiment.run(
RandomForestClassifier(class_weight='balanced', random_state=42),
{'n_estimators': [10, 100, 1000],
'min_samples_split': [2, 4, 6, 8, 10],
'max_depth': [2, 4, 6, 8, 10, None]})
# ### Gradient Boosting Machine
gbm_model = ml_experiment.run(
GradientBoostingClassifier(random_state=42),
{'n_estimators': [10, 100, 1000],
'learning_rate': [0.01, 0.1, 1, 10],
'min_samples_split': [2, 4, 6, 8, 10],
'max_depth': [2, 4, 8, 10, None]})
# ### Voting Classifier
voting_model = ml_experiment.run(
VotingClassifier([
('svm', svm_model), ('rf', rf_model), ('gbm', gbm_model)]),
{'voting': ['hard', 'soft']})
# ## Assemble Final Prediction Pipeline
#
# We will choose the random forest model to combine with the feature pipeline and then persist to disk.
prediction_pipeline = make_pipeline(feature_pipeline, rf_model)
# ### Re-Fit Model on Entire Dataset
prediction_pipeline.fit(data, data.Survived)
# ### Test Pipeline
prediction_pipeline.predict(test_data)
# ### Persist Model
# +
base_model_name = 'titanic-ml'
directory = 'models'
timestamp = datetime.now().isoformat(timespec='seconds')
#joblib.dump(prediction_pipeline, f'{directory}/{base_model_name}-{timestamp}.joblib')
joblib.dump(prediction_pipeline,'models/titanic-ml.joblib')
# -
|
Supervised Learning/Model Automation/Titanic-ml.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
"""
In theory this should work with reasonable outputs from PPMS system. One needs certain fields, like System, Year. This
script generates various useful statistics for ScopeM members.
Assumptions:
1. Paid project hours are:
* coming from project LM/EM or other EM projects at 0 / 40 / 200 CHF / h
* when using Hour's value metrics (default)
** if activity brings [0,40)CHF income it is assumed that its duration is equal to the number of hours recorded
When the income is suggesting more than 40CHF / hour duration is COMPUTED such as each 40CHF correspond to one hour
2. Unpaid project hours are:
* coming from Trials
* coming from personal R&D
Python:
* requires python-3.2
:bug:
None known.
:organization:
ETH
"""
__authors__="""<NAME>"""
__contact__="xxx"
__license__="Cecill-C"
__date__="17-12-05"
__version__="0.1"
__docformat__= "restructuredtext en"
# ---------------------------------------------------------- imports
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# ---------------------------------------------------------- const
cfg_sstoma_id = "<NAME>"
cfg_andrzej_id = "<NAME>"
cfg_sfn_id = "<NAME>"
cfg_ppms_csv_path = 'detailed-list.csv'
cfg_ppms_csv_path = 'detailed-list-2017.csv'
cfg_hour_baseline = 40 #CHF per h
# ---------------------------------------------------------- reading files & adding columns
# reading and listing headers (tricks and hacks applied to fck strange output format)
df = pd.read_csv(cfg_ppms_csv_path, encoding='cp1250', skiprows=[0])
list(df.columns.values)
# removing records without a project since they are useless
df = df[ pd.notnull(df.Project) ]
# adding month to stratify data
dates = pd.to_datetime(df.Date, format="%d/%m/%Y")
df['Month'] = dates.dt.month
df['Year'] = dates.dt.year
# adding time in hours (decimal)
df['Total Booked Hours'] = df['Length booked'] / 60.
# making sens of Amount* column (* is added when it was not paid yet...)
def convert_amount_starred( a ):
if a.find('*') == -1: return float(a)
else: return float(a[:-1])
df['Amount*'] = df['Amount*'].apply(convert_amount_starred)
# money can be stored everywhere
df['Total Charged [CHF]'] = df['Amount*'] + df['Prepaid']
# adding ratio of charged to all (under assumption 40 CHF / hr.)
df['Total Charged Hours'] = df['Total Booked Hours'] * (df['Total Charged [CHF]'] > 0)
# backcalculating:
# for 0 CHF the actual length will be used
# for >40 CHF the hours will be incrased
df["Hour's value"] = pd.DataFrame([df['Total Booked Hours'], df['Total Charged [CHF]'] / cfg_hour_baseline]).max()
# adding a field for classification
df["PPMS_category"] = "NONE"
# listing columns
list(df.columns.values)
# ---------------------------------------------------------- filtering
# prepare filters
# divide into project types
cat_paid_projects = (df.Project.str.match('[LE]M/.*') | df.Project.str.match('\d+_\w*') | df.Project.str.match('\d+ \w*') | df.Project.str.match('\d+'))
cat_unpaid_projects = (df.Project.str.match('_Trials') | df.Project.str.match('_LM/ScopeM/Trials')| df.Project.str.match('0060_Trials')) # currently (2017) some projects which should be Trials are in Paid_projects
cat_rd_personal_dev = df.Project.str.match('.*_PD') | df.Project.str.match('_LM/ScopeM/Scientific Collaboration')
cat_teaching = (df.Project.str.match('.*(MOOC).*') | df.Project.str.match('_Teach.*') | df.Project.str.match('_LM/ScopeM/Schools') | df.Project.str.match('_Microscopy_Program') )
cat_maintenance = (df.Project.str.match('_Main.*') | df.Project.str.match('_LM/ScopeM/Maintenance') | df.Project.str.match('EAWAG SP5') | df.Project.str.match('_LM/ScopeM/Visitors') )
cat_rd_other = df.Project.str.match('.*(CellStar|Crispr|Flu).*') | df.Project.str.match('_H2020*') | df.Project.str.match('ETHProj201*')
cat_sstoma = (df.System == cfg_sstoma_id)
cat_sfn = (df.System == cfg_sfn_id)
cat_andrzej = (df.System == cfg_andrzej_id)
cat_ida = (df.System == cfg_sstoma_id) | (df.System == cfg_sfn_id) | (df.System == cfg_andrzej_id)
# assign a PPMS tag based on filters
# careful, order matches since some things fall into two exp.
df.loc[cat_maintenance, "PPMS_category"] = "PPMS_Maintenance"
df.loc[cat_paid_projects, "PPMS_category"] = "PPMS_Projects"
df.loc[cat_unpaid_projects, "PPMS_category"] = "PPMS_Trials"
df.loc[cat_rd_personal_dev, "PPMS_category"] = "PPMS_RD_Personal"
df.loc[cat_rd_other, "PPMS_category"] = "PPMS_RD"
df.loc[cat_teaching, "PPMS_category"] = "PPMS_Teaching"
# define groups of users: so far only IDA
users_labels = df.System.unique()
users_filters = [df.System == u for u in users_labels]
users = list(zip(users_labels, users_filters))
users.append(("IDA", cat_ida))
# ---------------------------------------------------------- displaying stats
# if activity brings [0,40)CHF income it is assumed that its duration is equal to the number of hours recorded
# When the income is suggesting more than 40CHF / hour duration is COMPUTED such as each 40CHF correspond to one hour
metrics = "Hour's value"
#metrics = 'Total Booked Hours'
# do text outputs and compute useful stats
print('# Summary: PPMS_project AVG booked hours using metrics ' + metrics )
u_in_years = df.groupby("Year")["System"].nunique()
print(df[ df.PPMS_category == "PPMS_Projects" ].groupby(['Year'])[ metrics ].aggregate(np.sum) / u_in_years)
print('\n')
print('# Summary: Total income from PPMS_projects' )
print(df[ df.PPMS_category == "PPMS_Projects" ].groupby(['Year'])[ 'Total Charged [CHF]' ].aggregate(np.sum))
print('\n')
with pd.option_context('display.max_rows', None, 'display.max_columns', 3):
for cat in pd.unique(df.PPMS_category):
print('# Summary: '+ cat+' hours computed using metrics ' + metrics )
print(df[ df.PPMS_category == cat].groupby([ 'Year', 'System'])[ metrics ].sum().round(0))
print('\n')
print('# Summary: Total booked hours computed using metrics ' + metrics )
print(df.groupby([ 'Year', 'System'])[ metrics ].sum().round(0))
print('\n')
print('# Summary: Total booked hours computed using metrics ' + metrics )
print(df.groupby([ 'Year', 'System', 'PPMS_category'])[ metrics ].sum().round(0))
print('\n')
print('# Summary: Total booked hours computed using metrics ' + metrics )
print(df.groupby([ 'Year', 'System', 'PPMS_category', 'Project' ])[ metrics ].sum().round(0))
print('\n')
# do plots by PPMS_category
for cat in pd.unique(df.PPMS_category):
df[ df.PPMS_category == cat ].groupby(['Year', 'System'])[ metrics ].aggregate(np.sum).unstack().plot(kind = 'bar', title = u[ 0 ]+': by Year, '+cat).legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.ylabel(metrics)
plt.show()
# do plots foir each user
for u in users:
df[ u[1] ].groupby(['Year'])[ metrics ].aggregate(np.sum).plot(kind = 'bar', title = u[ 0 ]+': by Year, total PPMS_hours').legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.ylabel(metrics)
plt.show()
for u in users:
df[ u[1] ].groupby(['Year', 'PPMS_category'])[ metrics ].aggregate(np.sum).unstack().plot(kind = 'bar', title = u[ 0 ]+': by Year, PPMS_category').legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.ylabel(metrics)
plt.show()
|
code/ppms/17_11_28_ppms_stats.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: TensorFlow2.0
# language: python
# name: tensorflow2.0
# ---
# # TFLite Converter with TensorFlow 2.x
# 
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
from pathlib import Path
gpus = tf.config.experimental.list_physical_devices('GPU')
tf.config.experimental.set_visible_devices(gpus[0], 'GPU')
tf.config.experimental.set_memory_growth(gpus[0], True)
PATH_DIR = Path.cwd()
dataset_dir = PATH_DIR.joinpath('bin/tf_tutorial_2')
saved_model_dir = dataset_dir.joinpath('original_model')
saved_h5_dir = dataset_dir.joinpath('model_original.h5')
# 
# # 1.0 Train a simple CNN on MNIST
# import the datatset
(X_train, y_train), (X_test, y_test) = tf.keras.datasets.mnist.load_data(path='mnist.npz')
ds_train = tf.data.Dataset.from_tensor_slices((X_train, y_train))
ds_test = tf.data.Dataset.from_tensor_slices((X_test, y_test))
# normalize dataset
def normalize(x, y):
return x / 255, y
# prepare the data
ds_train = ds_train.map(normalize).cache().batch(32).prefetch(tf.data.experimental.AUTOTUNE)
ds_test = ds_test.map(normalize).batch(32)
# create a simple cnn model
model = tf.keras.models.Sequential([
tf.keras.layers.Reshape((28,28,1)),
tf.keras.layers.Conv2D(32, 3, activation='relu', padding='same'),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.Conv2D(32, 3, strides=2, padding='same', activation='relu'),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.Conv2D(64, 3, padding='same', activation='relu'),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.Conv2D(64, 3, strides=2, padding='same', activation='relu'),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.Conv2D(128, 3, padding='same', activation='relu'),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.Conv2D(128, 3, strides=2, padding='same', activation='relu'),
tf.keras.layers.GlobalAveragePooling2D(),
tf.keras.layers.Dense(10, activation='softmax' )
])
# compile
model.compile(
loss='sparse_categorical_crossentropy',
optimizer=tf.keras.optimizers.Adam(0.001),
metrics=['accuracy'])
# train
history = model.fit(ds_train, steps_per_epoch=len(X_train)/32, epochs=15)
# evaluate
model.evaluate((ds_test))
# save model
model.save(saved_model_dir)
model.save(saved_h5_dir)
# ## 2.0 TF-Lite simple conversion
# - Energy
# - Size
# - Latency
# - Costs
# from keras model
converter = tf.lite.TFLiteConverter.from_keras_model(model)
# or from tf saved model
converter = tf.lite.TFLiteConverter.from_saved_model(saved_model_dir)
# last from concrete functions
converter = tf.lite.TFLiteConverter.from_concrete_funcions(tf_path_concrete_functions)
converter = tf.lite.TFLiteConverter.from_saved_model(saved_model_dir.as_posix())
# start conversion
tflite_model = converter.convert()
# save model
tflite_model_file = dataset_dir.joinpath('model_fp32.tflite')
tflite_model_file.write_bytes(tflite_model)
# # 3.0 Float16 quantization
#
# - **Size reduction:** up to 50 %
# - **Latency reduction:** 2x
# - **Accuracy:** Insignificant loss accuracy
converter = tf.lite.TFLiteConverter.from_saved_model(saved_model_dir.as_posix())
converter.optimizations = [tf.lite.Optimize.DEFAULT]
converter.target_spec.supported_types = [tf.float16]
tflite_model = converter.convert()
# save model
tflite_model_file = dataset_dir.joinpath('model_fp16.tflite')
tflite_model_file.write_bytes(tflite_model)
# # 4.0 Dynamic int8 range quantization
#
# - **Size reduction:** up to 75 %
# - **Latency reduction:** 2x/3x
# - **Accuracy:** Accuracy loss
converter = tf.lite.TFLiteConverter.from_saved_model(saved_model_dir.as_posix())
converter.optimizations = [tf.lite.Optimize.DEFAULT]
tflite_model = converter.convert()
# save model
tflite_model_file = dataset_dir.joinpath('model_int8_dynamic.tflite')
tflite_model_file.write_bytes(tflite_model)
# # 5.0 Integer quantization with float fallback
#
# - **Size reduction:** up to 75 %
# - **Latency reduction:** 3x/4x
# - **Accuracy:** Smallest loss accuracy
# +
num_calibration_steps = 1 # at least 100
def representative_dataset_gen():
for i in range(num_calibration_steps):
# Remember to pre-process your dataset as your training
imgs = X_train[i:i+1]
imgs = imgs / 255
yield [imgs.astype('float32')]
# -
converter = tf.lite.TFLiteConverter.from_saved_model(saved_model_dir.as_posix())
converter.optimizations = [tf.lite.Optimize.DEFAULT]
converter.representative_dataset = representative_dataset_gen
tflite_model = converter.convert()
# save model
tflite_model_file = dataset_dir.joinpath('model_int8_fb.tflite')
tflite_model_file.write_bytes(tflite_model)
# # 6.0 Full integer quantization (integer only)
#
# - **Size reduction:** up to 75 %
# - **Latency reductiion:** 3x/4x
# - **Accuracy:** Smallest loss accuracy
# ## 6.1 With TF >= 2.3
# +
num_calibration_steps = 1 # at least 100
def representative_dataset_gen():
for i in range(num_calibration_steps):
# Remember to pre-process your dataset as your training
imgs = X_train[i:i+1]
imgs = imgs / 255
yield [imgs.astype('float32')]
# -
converter = tf.lite.TFLiteConverter.from_saved_model(saved_model_dir.as_posix())
converter.optimizations = [tf.lite.Optimize.DEFAULT]
converter.representative_dataset = representative_dataset_gen
converter.target_spec.supported_ops = [tf.lite.OpsSet.TFLITE_BUILTINS_INT8]
converter.inference_input_type = tf.int8 # or tf.uint8
converter.inference_output_type = tf.int8 # or tf.uint8
tflite_model = converter.convert()
# save model
tflite_model_file = dataset_dir.joinpath('model_int8.tflite')
tflite_model_file.write_bytes(tflite_model)
# ## 6.2 With TF < 2.3
# +
num_calibration_steps = 1 # at least 100
def representative_dataset_gen():
for i in range(num_calibration_steps):
# Remember to pre-process your dataset as your training
imgs = X_train[i:i+1]
imgs = imgs / 255
yield [imgs.astype('float32')]
# -
converter = tf.compat.v1.lite.TFLiteConverter.from_keras_model_file(dataset_dir.joinpath('model_original.h5'))
converter.optimizations = [tf.lite.Optimize.OPTIMIZE_FOR_SIZE]
converter.representative_dataset = representative_dataset_gen
converter.experimental_new_converter = True
converter.target_spec.supported_ops = [tf.lite.OpsSet.TFLITE_BUILTINS_INT8]
converter.inference_input_type = tf.uint8
converter.inference_output_type = tf.uint8
tflite_model = converter.convert()
# save model
tflite_model_file = dataset_dir.joinpath('model_int8.tflite')
tflite_model_file.write_bytes(tflite_model)
|
Notebooks/0 - TF2.X Tutorials/tf_2_tflite_conversions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: research
# language: python
# name: research
# ---
# # Beta Extraction
#
# Replicating LSA regression using scikit-learn just to make sure they're okay.
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
# +
import json
import sys
import os
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
if sys.platform == 'darwin':
cfg = os.path.join("/Users", "njchiang", "CloudStation", "Grad",
"Research", "montilab-ucla", "analogy", "config", "project.json")
plat = "osx"
elif sys.platform == "linux":
import platform
if platform.linux_distribution()[0] == "debian":
cfg = os.path.join("/home", "njchiang", "data", "CloudStation", "Grad",
"Research", "montilab-ucla", "analogy", "config", "project.json")
plat = "linux"
else:
cfg = os.path.join("/u", "project", "monti", "Analysis", "Analogy",
"code", "analogy", "config", "project.json")
plat = "hoff"
else:
cfg = os.path.join("D:\\", "CloudStation", "Grad",
"Research", "montilab-ucla", "analogy", "config", "project.json")
plat = "win"
with open(cfg, "r") as f:
projectSettings = json.load(f)
paths = projectSettings["filepaths"]["{}Paths".format(plat)]
sys.path.append(paths["github"])
sys.path.append(paths["code"])
# -
from fmri.beta_extract import create_lss_from_lsa, load_aggregated_data, calc_scores
from fmri.analogy_utils import analysisSettings, contrastSettings, order, \
pu, pa, pv, compile_models, rsa, save_rois, load_rois
from sklearn.linear_model import LinearRegression
import statsmodels.api as sm
from sklearn.externals.joblib import Parallel, delayed
x = np.random.rand(100, 10)
y = np.random.rand(100, 2)
model = sm.OLS(y[:, 0], x)
results = model.fit()
scores = Parallel(n_jobs=-2)(delayed(calc_scores)(y[:, v], x) for v in range(y.shape[1]))
# Run LSS extraction!
maskname = "grayMatter-bin_mask"
for sub in projectSettings["subjects"]:
mask = pu.load_img(
os.path.join(paths["root"], "derivatives", sub, "masks",
"{}.nii.gz".format(maskname)), logger=None)
fmri_data, design = load_aggregated_data(paths, sub, maskname)
betas = np.zeros([144, fmri_data.shape[1]])
tstats = np.zeros([144, fmri_data.shape[1]])
for i, wordpair in enumerate(order.ABTag[::2]):
print(wordpair)
regressors = create_lss_from_lsa(design, wordpair, tt="AB")
# scores = Parallel(n_jobs=-2)(delayed(calc_scores)(fmri_data[:, v], regressors) for v in range(fmri_data.shape[1]))
# scores = np.array(scores)
# betas[i] = scores[:, 0]
# tstats[i] = scores[:, 1]
betas[i] = LinearRegression().fit(regressors, fmri_data).coef_[:, 0]
pu.unmask_img(betas, mask).to_filename(
os.path.join(paths["root"], "derivatives", sub, "betas",
"{}_task-analogy_betas-cope-LSS-condensed.nii.gz".format(sub)))
# pu.unmask_img(tstats, mask).to_filename(
# os.path.join(paths["root"], "derivatives", sub, "betas",
# "{}_task-analogy_betas-tstat-LSS-condensed.nii.gz".format(sub)))
# ## Scratch
maskname = "grayMatter-bin_mask"
mask = pu.load_img(
os.path.join(paths["root"], "derivatives", "sub-01", "masks",
"{}.nii.gz".format(maskname)), logger=None)
pu.unmask_img(betas, mask).to_filename(
os.path.join(paths["root"], "derivatives", "sub-01", "betas",
"sub-01_task-analogy_betas-LSS-condensed.nii.gz"))
# sort by columns and take every 2...3?
for tag in test:
if ":" in tag:
trialtype = tag.split("_")[1]
if trialtype == "AB":
this_tag = tag.split("_")[0].split("::")[0]
elif trialtype == "CD":
this_tag = tag.split("_")[0].split("::")[1]
else:
this_tag = tag.split("_")[0]
# +
n_trials = 108
test = des.matrix
i = []
nuisance = test[:, n_trials:]
trial_regs = test[:, :n_trials]
if sum(i) > 1:
trial = trial_regs[:, i]
trial_nuisance = np.sum(np.delete(trial_regs, i, axis=1), axis=1, keepdims=True)
else:
trial = np.zeros([trial_regs.shape[0], 1])
trial_nuisance = np.sum(trial_regs, axis=1, keepdims=True)
this_design = np.hstack([trial, trial_nuisance, nuisance])
# -
this_design
labels[labels.ABTag == "Hello Bloop"].index
test[:, i:i+1]
# +
frametimes = np.arange(0, fmri_data.shape[0], tr)
cond_ids = [i for i, r in test_l.iterrows()]
onsets = labels["Onset"]
durations = labels["Duration"]
design_kwargs = {"drift_model": "blank", "add_regs": motion.values}
LSA_des = pa.make_designmat(frametimes, cond_ids, onsets, durations,
design_kwargs=design_kwargs, logger=logger)
# -
for sub, runs in projectSettings["subjects"].items():
[regress_betas(paths, sub, r, maskname="template-bin_mask", write=True) for r in runs]
masks_dict = {"ba10": "dfc-left-BA10-bin_mask",
"lofc": "aal-LOFC-bin_mask",
"ba46": "dfc-left-BA46-bin_mask",
"lpitg": "anat-left-pITG-bin_mask",
"laitg": "anat-left-aITG-bin_mask",
"lastg": "anat-left-aSTG-bin_mask",
"lifg": "anat-ba44-45_mask",
"lips": "anat-LIPS_mask",
"v1": "aal-V1-bin_mask",
"csf": "csf-bin_mask"}
save_rois(masks_dict, t="LSA")
save_rois(masks_dict, t="cope-LSS")
save_rois(masks_dict, t="pymvpa")
|
notebooks/BetaExtract.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="59741d6d-450a-485e-b7ab-233b54a0bbbe"
# ## Modeling - customer response will be predicted based on the profile and when the last customer purchases
# + id="178a6e8d-5b95-4d48-8779-1330075c014e"
import numpy as np # linear algebra
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
import warnings
warnings.filterwarnings("ignore")
# PreProcessing
from sklearn.compose import ColumnTransformer
from sklearn.preprocessing import LabelEncoder, OneHotEncoder,RobustScaler,MinMaxScaler,StandardScaler
import category_encoders as ce
from sklearn.pipeline import Pipeline
from sklearn.impute import SimpleImputer
# Splitting Data
from sklearn.model_selection import train_test_split, StratifiedKFold, cross_val_score
# Resampling
from imblearn.over_sampling import RandomOverSampler, SMOTE
from imblearn.under_sampling import RandomUnderSampler, NearMiss
from imblearn.combine import SMOTETomek
from imblearn.pipeline import Pipeline
# Modeling, Fitting and Evaluation
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import classification_report,confusion_matrix, accuracy_score,f1_score, precision_score, roc_auc_score, plot_roc_curve,recall_score
from sklearn.tree import DecisionTreeClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.model_selection import RandomizedSearchCV, GridSearchCV
from sklearn.ensemble import RandomForestClassifier
import category_encoders as ce
from sklearn import metrics
from datetime import datetime
import datetime
# Boosting
from sklearn.ensemble import AdaBoostClassifier, GradientBoostingClassifier
from xgboost.sklearn import XGBClassifier
#feature Selection
from sklearn.feature_selection import SelectPercentile, RFE
#saving
import pickle
# + id="bbf24c75-095c-477f-b427-00da6c3214b3"
#load data
data = pd.read_csv('df_copy.csv')
# + id="1c462584-2f69-4337-bfd5-1f67a6d3ace1" outputId="04204dc1-69ae-4b21-80f4-facc959a1e35"
data.columns
# + id="adcd5b07-4a9c-424b-b55a-1be15f271327" outputId="f2d9bb6f-ded1-474c-c161-1878e91d858f"
#most successful campaign
campaign = data.loc[:,['Response','AcceptedCmp1','AcceptedCmp2','AcceptedCmp3','AcceptedCmp4','AcceptedCmp5']]
campaign = campaign.melt()
campaign = pd.crosstab(campaign["variable"], campaign["value"]).sort_values(0)
cols = list(campaign.columns)
a, b = cols.index(0), cols.index(1)
cols[b], cols[a] = cols[a], cols[b]
campaign = campaign[cols]
campaign.columns = "Yes","No"
campaign.plot.bar(stacked=True)
plt.title('Acceptance of Marketing Campaigns')
plt.xlabel('Campaign')
plt.ylabel('Acceptance')
plt.legend(title='Response',loc='upper left',bbox_to_anchor=(1, 0.5))
plt.show()
# + [markdown] id="2644cdd0-b7f4-4eca-8183-e584adb65765"
# ## Preprocessing
# + id="0ed91488-0984-44e6-ac13-34535f25c3f2"
# we will scale the numerical columns and encode the categorical columns
scale = Pipeline([
('scaling', RobustScaler()),
])
transformer = ColumnTransformer([
('encoder',OneHotEncoder(handle_unknown='ignore'),['Education','Marital_Status']),
('binary',ce.BinaryEncoder(),['Country']),
('scale',RobustScaler(),['Ages','Recency'])
],remainder='passthrough')
# + id="344448cf-80b6-4705-9d32-a8c9b167a154"
# predicting the customers response on based on their profile and their last purchase.
data=data.drop(['MntWines','MntFruits','MntMeatProducts','MntFishProducts','MntSweetProducts','MntGoldProds','NumDealsPurchases','NumWebPurchases','NumCatalogPurchases','NumStorePurchases','NumWebVisitsMonth','Complain','Dt_Customer','ID','AcceptedCmp3','AcceptedCmp4','AcceptedCmp5','AcceptedCmp1','AcceptedCmp2','Age_Group', 'Join_year',
'Join_month', 'Join_weekday', 'Dependents', 'Total_Mnt', 'Total_num_purchase', 'Total_accept', 'AOV'],axis=1)
# + id="4fcbb5b8-d8d8-4fbd-b5b0-6cd749f60f1d" outputId="f80d5674-fa5a-429a-aeaf-1fe9f55b2005"
data.columns
# + id="d9572e62-8ba8-4f3b-9427-b1f7511a332e" outputId="826c8f0d-5c8d-42c0-a2a1-976d8365c96b"
data.head()
# + id="126341df-717f-4e28-954a-5e61862ebeb2"
# Split the dataset into X , y
X=data.drop(['Response'],axis=1)
y=data['Response']
# + id="a5b38572-d982-411d-be65-7573f96ef7c2" outputId="42df1c06-6d23-47d5-e1af-3a35b436b360"
print(X.shape, y.shape)
# + id="5f8570d6-cbec-4855-a806-022392d8fe13" outputId="a503b548-eff4-4450-dce1-f6f4cd3b88b4"
#check transform: scale the training data
transformer.fit_transform(data)
# + id="c553ed14-990c-4ab6-8187-64ed6ee7200f" outputId="e8bf70f2-889f-448a-8e25-2434ec1e6ede"
X.head()
# + id="07d3348b"
# Data Splitting: Fit model to training data (70% of dataset) and Evaluate predictions on test data (30% of dataset)
# + id="d93fc25d-ff6d-49b7-b9ff-1245b6d84ea1"
# Split dataset into training set (70% of dataset) and test set (30% of dataset)
X_train,X_test,y_train,y_test=train_test_split(X,y,stratify=y,test_size=0.3,random_state=2020)
# + id="99d59b34-cd0a-4d52-a4ee-dc53a0f886db" outputId="41a287d8-e275-44f8-a632-0c63be5374bc"
print(X_train.shape, X_test.shape)
# + [markdown] id="7f790af3"
# ## Model Prediction
# + id="d02aebc3-d332-4f1c-acc7-36e9cdecd184" outputId="fa82c444-5f04-4e38-c816-d8abcf8de922"
#Checking how balanced is our target Data
data['Response'].value_counts()/data.shape[0]*100
# + id="d7df8720" active=""
# # Our data is imbalanced, so we will check F1 score instead of accuracy.
# + id="d011d5e5" active=""
# # We are going to try different Machine learning algorithms to compare the best results. Generally KNN and RandomFprest algorithm performed better than other algorithms in imbalanced data sets. We will also try some resampling techniques of balancing and compare the results.
# Hence we are trying below algorithms
# LogisticRegression
# DecisionTreeClassifier
# KNeighborsClassifier
# RandomForestClassifier
# + id="02bd4951-58c6-43c9-9ae6-fdbc38470718"
# Model selection
lr = LogisticRegression()
tree = DecisionTreeClassifier(random_state = 2020)
knn = KNeighborsClassifier()
rf = RandomForestClassifier(random_state = 2020)
# + id="0b223425-42e5-41b7-b871-c82eeddf6d0d"
# creating pipeline
lr_pipe = Pipeline([
('transform',transformer),
('lr',lr)
])
tree_pipe= Pipeline([
('transform',transformer),
('tree',tree)
])
knn_pipe =Pipeline([
('transform',transformer),
('knn',knn)
])
rf_pipe = Pipeline([
('transform',transformer),
('rf',rf)
])
# + id="28535337-1163-445a-895a-4aca631171d3" outputId="ba415405-dbb7-4cf0-f8f0-f0343818686c"
def model_evaluation(model, metric):
skfold = StratifiedKFold(n_splits = 5)
model_cv = cross_val_score(model, X_train, y_train, cv = skfold, scoring = metric)
return model_cv
lr_pipe_cv = model_evaluation(lr_pipe, 'precision')
tree_pipe_cv = model_evaluation(tree_pipe, 'precision')
knn_pipe_cv = model_evaluation(knn_pipe, 'precision')
rf_pipe_cv = model_evaluation(rf_pipe, 'precision')
for model in [lr_pipe,tree_pipe, knn_pipe,rf_pipe]:
model.fit(X_train, y_train)
score_mean = [lr_pipe_cv.mean(),tree_pipe_cv.mean(),knn_pipe_cv.mean(),rf_pipe_cv.mean()]
score_std = [lr_pipe_cv.std(),tree_pipe_cv.std(),knn_pipe_cv.std(),rf_pipe_cv.std()]
score_precision_score = [precision_score(y_test, lr_pipe.predict(X_test)),
precision_score(y_test, tree_pipe.predict(X_test)),
precision_score(y_test, knn_pipe.predict(X_test)),
precision_score(y_test, rf_pipe.predict(X_test))]
score_f1_score = [f1_score(y_test, lr_pipe.predict(X_test)),f1_score(y_test, tree_pipe.predict(X_test)),
f1_score(y_test, knn_pipe.predict(X_test)),f1_score(y_test, rf_pipe.predict(X_test))]
method_name = ['Logistic Regression','Decision Tree Classifier','KNN Classifier', 'Random Forest Classifier']
cv_result = pd.DataFrame({
'method': method_name,
'mean score': score_mean,
'std score': score_std,
'precision score': score_precision_score,
'f1 score': score_f1_score,
})
cv_result
# + id="f623f51a" outputId="ca66beb5-f022-4c03-d36e-86136cc631b9"
# confusion matrix KNN
knn_pipe.fit(X_train, y_train)
ypred=knn_pipe.predict(X_test)
print(classification_report(y_test,ypred))
print(metrics.confusion_matrix(y_test,ypred))
# + id="36ecc839" active=""
# knn_pipe.fit(X_train, y_train)
# y_pred_estimator = knn_pipe.predict(X_test)
# precision_estimator = precision_score(y_test, knn_pipe.predict(X_test))
# f1_estimator = f1_score(y_test, y_pred_estimator, average='binary')
#
# train_pred = knn_pipe.predict(X_train) #prediction on trained data
# print(X_train.shape, y_test.shape, train_pred.shape)
# train_precision_estimator = precision_score(y_train, train_pred)
# train_f1_estimator = f1_score(y_train, train_pred)
#
# score_list = [precision_estimator, f1_estimator, train_precision_estimator,train_f1_estimator]
# method_name = ['KNN Classifier_Test', 'KNN Classifier_Train']
# best_summary = pd.DataFrame({
# 'method': method_name,
# 'score': score_list
# })
# best_summary
# + id="742433c8" outputId="d339435e-32ff-4851-b43b-a78dfbefb8dc"
# confusion matrix RANDOM forest classifier
rf_pipe.fit(X_train, y_train)
ypred=rf_pipe.predict(X_test)
print(classification_report(y_test,ypred))
print(metrics.confusion_matrix(y_test,ypred))
# + id="837dea83-5f12-4457-9279-557a20e03c44" active=""
# Handling Imbalance Dataset: Random Resampling Imbalanced Datasets
# + id="6829187d-4c9d-497e-a8b4-bd5881eae927" active=""
# Random Under Sampling: Random undersampling involves randomly selecting examples from the majority class and deleting them from the training dataset.
# + id="fb5a2be2-b627-4af4-9ef5-9485cdba2ab7"
rus = RandomUnderSampler(random_state = 2020)
X_under, y_under = rus.fit_resample(X_train, y_train)
# + id="52159902-c8a9-484a-91a6-c9f207fafdc3"
lr_pipe_under = Pipeline([
('transformer', transformer),
('rus', rus),
('lr', lr)
])
tree_pipe_under = Pipeline([
('transformer', transformer),
('rus', rus),
('tree', tree)
])
knn_pipe_under = Pipeline([
('transformer', transformer),
('rus', rus),
('knn', knn)
])
rf_pipe_under = Pipeline([
('transformer', transformer),
('rus', rus),
('rf', rf)
])
# + id="c2f7b309-8348-4562-b50e-c024c6895d8c" outputId="bb576cdc-71ec-40ee-abe7-3639ece719f2"
def model_evaluation(model, metric):
skfold = StratifiedKFold(n_splits = 5)
model_cv = cross_val_score(model, X_train, y_train, cv = skfold, scoring = metric)
return model_cv
lr_under_cv = model_evaluation(lr_pipe_under, 'precision')
tree_under_cv = model_evaluation(tree_pipe_under, 'precision')
knn_under_cv = model_evaluation(knn_pipe_under, 'precision')
rf_under_cv = model_evaluation(rf_pipe_under, 'precision')
for model in [lr_pipe_under, tree_pipe_under, knn_pipe_under, rf_pipe_under]:
model.fit(X_train, y_train)
score_mean = [lr_under_cv.mean(), tree_under_cv.mean(), knn_under_cv.mean(),
rf_under_cv.mean()]
score_std = [lr_under_cv.std(), tree_under_cv.std(), knn_under_cv.std(),
rf_under_cv.std()]
score_precision_score = [precision_score(y_test, lr_pipe_under.predict(X_test)),
precision_score(y_test, tree_pipe_under.predict(X_test)),
precision_score(y_test, knn_pipe_under.predict(X_test)),
precision_score(y_test, rf_pipe_under.predict(X_test))]
score_f1_score = [f1_score(y_test, lr_pipe_under.predict(X_test)),f1_score(y_test, tree_pipe_under.predict(X_test)),
f1_score(y_test, knn_pipe_under.predict(X_test)),f1_score(y_test, rf_pipe_under.predict(X_test))]
method_name = ['Logistic Regression UnderSampling', 'Decision Tree Classifier UnderSampling',
'KNN Classifier UnderSampling', 'Random Forest Classifier UnderSampling']
under_result = pd.DataFrame({
'method': method_name,
'mean score': score_mean,
'std score': score_std,
'precision score': score_precision_score,
'f1 score': score_f1_score,
})
under_result
# + id="289ed8b7-a77d-49bc-908b-2cdea13de791" active=""
# # Random Over Sampling: Random oversampling involves randomly selecting examples from the minority class, with replacement, and adding them to the training dataset.
# + id="f1fb00d4-f1dc-4904-8607-19d66907c78a"
ros = RandomOverSampler(random_state = 2020)
X_over, y_over = ros.fit_resample(X_train, y_train)
# + id="f0f47365-3132-48d6-8e91-9b36b92fbb2c"
lr_pipe_over = Pipeline([
('transformer', transformer),
('ros', ros),
('lr', lr)
])
tree_pipe_over = Pipeline([
('transformer', transformer),
('ros', ros),
('tree', tree)
])
knn_pipe_over = Pipeline([
('transformer', transformer),
('ros', ros),
('knn', knn)
])
rf_pipe_over = Pipeline([
('transformer', transformer),
('ros', ros),
('rf', rf)
])
# + id="b024d5e5-9756-422d-b39c-5130fadcbee4" outputId="67f7e154-170e-4e0e-d855-6e1863360442"
def model_evaluation(model, metric):
skfold = StratifiedKFold(n_splits = 5)
model_cv = cross_val_score(model, X_train, y_train, cv = skfold, scoring = metric)
return model_cv
lr_over_cv = model_evaluation(lr_pipe_over, 'precision')
tree_over_cv = model_evaluation(tree_pipe_over, 'precision')
knn_over_cv = model_evaluation(knn_pipe_over, 'precision')
rf_over_cv = model_evaluation(rf_pipe_over, 'precision')
for model in [lr_pipe_over, tree_pipe_over, knn_pipe_over, rf_pipe_over]:
model.fit(X_train, y_train)
score_mean = [lr_over_cv.mean(), tree_over_cv.mean(), knn_over_cv.mean(),
rf_over_cv.mean()]
score_std = [lr_over_cv.std(), tree_over_cv.std(), knn_over_cv.std(),
rf_over_cv.std()]
score_precision_score = [precision_score(y_test, lr_pipe_over.predict(X_test)),
precision_score(y_test, tree_pipe_over.predict(X_test)),
precision_score(y_test, knn_pipe_over.predict(X_test)),
precision_score(y_test, rf_pipe_over.predict(X_test))]
score_f1_score = [f1_score(y_test, lr_pipe_over.predict(X_test)),f1_score(y_test, tree_pipe_over.predict(X_test)),
f1_score(y_test, knn_pipe_over.predict(X_test)),f1_score(y_test, rf_pipe_over.predict(X_test))]
method_name = ['Logistic Regression OverSampling', 'Decision Tree Classifier OverSampling',
'KNN Classifier OverSampling', 'Random Forest Classifier OverSampling']
over_summary = pd.DataFrame({
'method': method_name,
'mean score': score_mean,
'std score': score_std,
'precision score': score_precision_score,
'f1 score': score_f1_score,
})
over_summary
# + id="d34ad047-0cfb-49d4-af64-cc455f769171" active=""
# #NearMiss: NearMiss Algorithm – Undersampling-
# NearMiss – Version 1 : It selects samples of the majority class for which average distances to the k closest instances of the minority class is smallest.
# + id="5b4c8cb6-c0b4-4adc-accd-8cd68e2097dc"
nm = NearMiss(version = 1)
# + id="e86872b7-a239-43d0-95e1-a050b3b837b5"
lr_pipe_nm = Pipeline([
('transformer', transformer),
('nm', nm),
('lr', lr)
])
tree_pipe_nm = Pipeline([
('transformer', transformer),
('nm', nm),
('tree', tree)
])
knn_pipe_nm = Pipeline([
('transformer', transformer),
('nm', nm),
('knn', knn)
])
rf_pipe_nm = Pipeline([
('transformer', transformer),
('nm', nm),
('rf', rf)
])
# + id="6ba81688-f943-46a7-82e3-03ab8fa89986" outputId="6b8f13a9-06db-4567-f9ab-89bc7eaab991"
def model_evaluation(model, metric):
skfold = StratifiedKFold(n_splits = 5)
model_cv = cross_val_score(model, X_train, y_train, cv = skfold, scoring = metric)
return model_cv
lr_nm_cv = model_evaluation(lr_pipe_nm, 'precision')
tree_nm_cv = model_evaluation(tree_pipe_nm, 'precision')
knn_nm_cv = model_evaluation(knn_pipe_nm, 'precision')
rf_nm_cv = model_evaluation(rf_pipe_nm, 'precision')
for model in [lr_pipe_nm, tree_pipe_nm, knn_pipe_nm, rf_pipe_nm]:
model.fit(X_train, y_train)
score_mean = [lr_nm_cv.mean(), tree_nm_cv.mean(), knn_nm_cv.mean(),
rf_nm_cv.mean()]
score_std = [lr_nm_cv.std(), tree_nm_cv.std(), knn_nm_cv.std(),
rf_nm_cv.std()]
score_precision_score = [precision_score(y_test, lr_pipe_nm.predict(X_test)),
precision_score(y_test, tree_pipe_nm.predict(X_test)),
precision_score(y_test, knn_pipe_nm.predict(X_test)),
precision_score(y_test, rf_pipe_nm.predict(X_test))]
score_f1_score = [f1_score(y_test, lr_pipe_nm.predict(X_test)),f1_score(y_test, tree_pipe_nm.predict(X_test)),
f1_score(y_test, knn_pipe_nm.predict(X_test)),f1_score(y_test, rf_pipe_nm.predict(X_test))]
method_name = ['Logistic Regression NearMiss', 'Decision Tree Classifier NearMiss',
'KNN Classifier NearMiss', 'Random Forest Classifier NearMiss']
nm_summary = pd.DataFrame({
'method': method_name,
'mean score': score_mean,
'std score': score_std,
'precision score': score_precision_score,
'f1 score': score_f1_score,
})
nm_summary
# + id="03ae35f7-aa77-4dee-949c-c124c6afeedc" outputId="b854117b-0601-45db-bc82-94acb3ddb1ce"
#Summary Balancing Dataset
resume_balancing = pd.concat([under_result,over_summary,nm_summary], axis=0)
resume_balancing
# + id="e63d5b03-084a-4ccd-9720-fdb037e4fc06" active=""
# After balancing the dataset the f1 score is decreased or similar, so better model will be without balancing the dataset. Based on the above summary stable model are KNN and Random Forest Classifier.
# + id="df1df102-3ece-42f7-804d-3040b0e7e3af" active=""
# BOOSTING MODEL: we will now try few boosting model and compare the results
# + id="93ed9d29-72c4-468d-813b-4ca80445676f"
adaboost = AdaBoostClassifier(
tree,
n_estimators = 50,
learning_rate = 0.1,
random_state = 2020)
pipe_ada = Pipeline([
('transformer', transformer),
('adaboost', adaboost)
])
gradboost = GradientBoostingClassifier(
n_estimators = 50,
learning_rate = 0.1,
max_depth = 3,
random_state = 2020)
pipe_grad = Pipeline([
('transformer', transformer),
('gradboost', gradboost)
])
XGBOOST = XGBClassifier(
n_estimators = 50,
learning_rate = 0.1,
max_depth = 3,
random_state = 2020)
pipe_XGB = Pipeline([
('transformer', transformer),
('XGBOOST', XGBOOST)
])
# + id="c0d88adc-ba7e-40ec-9dc8-4e365a45a67a" outputId="a64dfed7-affa-4b16-ba12-c9d23775b454"
def model_evaluation(model, metric):
skfold = StratifiedKFold(n_splits = 5)
model_cv = cross_val_score(model, X_train, y_train, cv = skfold, scoring = metric, n_jobs = -1)
return model_cv
pipe_ada_cv = model_evaluation(pipe_ada, 'precision')
pipe_grad_cv = model_evaluation(pipe_grad, 'precision')
pipe_XGB_cv = model_evaluation(pipe_XGB, 'precision')
for model in [pipe_ada, pipe_grad, pipe_XGB]:
model.fit(X_train, y_train)
score_mean = [pipe_ada_cv.mean(), pipe_grad_cv.mean(), pipe_XGB_cv.mean()]
score_std = [pipe_ada_cv.std(), pipe_grad_cv.std(), pipe_XGB_cv.std()]
score_precision_score = [precision_score(y_test, pipe_ada.predict(X_test)),
precision_score(y_test, pipe_grad.predict(X_test)),
precision_score(y_test, pipe_XGB.predict(X_test))]
score_f1_score = [f1_score(y_test, pipe_ada.predict(X_test)),f1_score(y_test, pipe_grad.predict(X_test)),
f1_score(y_test, pipe_grad.predict(X_test))]
method_name = ['Ada Boost Classifier', 'Gradient Boost Classifier',
'XGB Classifier']
boost_summary = pd.DataFrame({
'method': method_name,
'mean score': score_mean,
'std score': score_std,
'precision score': score_precision_score,
'f1 score': score_f1_score,
})
boost_summary
# + id="e1ace324-1333-496f-a7af-423edfa3a620" outputId="f9bbc732-7176-4122-c946-ead403143f95"
resume_model = pd.concat([cv_result,boost_summary], axis=0)
resume_model
# + id="1b295bb6" active=""
# Based on the above resume, we will choose KNN Classifier because it has the highest F1 score.
# + id="cac0a7ca-dcfc-4471-9fa4-57fe6aca7791" active=""
# Based on the above resume, we will choose KNN Classifier, because it has the highest precision score and precision score between class 1 and 0 is balance.
#
# + id="c35366f6"
# KNN Classifier score
# + id="cb432395-e043-400e-a6b2-1707a204de87" outputId="07dce7e5-65fa-49fe-8e0e-a9fe6e87fc2e"
knn_pipe.fit(X_train, y_train)
y_pred_estimator = knn_pipe.predict(X_test)
precision_estimator = precision_score(y_test, knn_pipe.predict(X_test))
f1_estimator = f1_score(y_test, y_pred_estimator)
method_name = ['KNN Classifier_Test']
best_summary = pd.DataFrame({
'method': method_name,
'Precision_score': precision_estimator,
'F1_score': f1_estimator
})
best_summary
# + id="a1198650" outputId="52e9e706-ea04-45b7-9ad5-dacf9ce39e5b"
train_pred = knn_pipe.predict(X_train) #prediction on trained data
train_precision_estimator = precision_score(y_train, train_pred)
train_f1_estimator = f1_score(y_train, train_pred)
method_name = ['KNN Classifier_Train']
train_best_summary = pd.DataFrame({
'method': method_name,
'Precision_score': train_precision_estimator,
'F1_score': train_f1_estimator
})
train_best_summary
# + id="385f2e7c" outputId="67ff546b-d5ad-4049-d8d7-709b0ab9218b"
rf_pipe.fit(X_train, y_train)
rf_y_pred_estimator = rf_pipe.predict(X_test)
precision_estimator = precision_score(y_test,rf_y_pred_estimator)
f1_estimator = f1_score(y_test, rf_y_pred_estimator, average='binary')
#score_list = [precision_estimator,f1_estimator, train_precision_estimator ,train_f1_estimator]
method_name = ['Random Forest Classifier_Test']
rf_best_summary = pd.DataFrame({
'method': method_name,
'precision_score': precision_estimator,
'F1_score': f1_estimator
})
rf_best_summary
# + id="15e8bd35" outputId="707d7d52-99fb-41c3-fec6-a55b608d5932"
rf_train_pred = knn_pipe.predict(X_train) #prediction on trained data
train_precision_estimator = precision_score(y_train,rf_train_pred)
train_f1_estimator = f1_score(y_train, rf_train_pred)
method_name = ['Random Forest Classifier_Train']
rf_train_best_summary = pd.DataFrame({
'method': method_name,
'precision_score': train_precision_estimator,
'F1_score': train_f1_estimator
})
rf_train_best_summary
# + id="6eb34f33" outputId="9b4894fe-eb5a-4c88-df5c-8270d5c796b3"
pipe_XGB.fit(X_train, y_train)
XGB_y_pred_estimator = pipe_XGB.predict(X_test)
precision_estimator = precision_score(y_test,XGB_y_pred_estimator)
f1_estimator = f1_score(y_test, XGB_y_pred_estimator, average='binary')
#score_list = [precision_estimator,f1_estimator, train_precision_estimator ,train_f1_estimator]
method_name = ['XGB Classifier_Test']
XGB_best_summary = pd.DataFrame({
'method': method_name,
'precision_score': precision_estimator,
'F1_score': f1_estimator
})
XGB_best_summary
# + id="a9061c8d" outputId="6600108b-aab4-400b-8c68-2d082037f8cf"
XGB_train_pred = pipe_XGB.predict(X_train) #prediction on trained data
train_precision_estimator = precision_score(y_train,XGB_train_pred)
train_f1_estimator = f1_score(y_train, XGB_train_pred)
method_name = ['XGB Classifier_Train']
XGB_train_best_summary = pd.DataFrame({
'method': method_name,
'precision_score': train_precision_estimator,
'F1_score': train_f1_estimator
})
XGB_train_best_summary
# + id="16b8f82d" outputId="6635d07f-3d54-4cd0-ff28-f2302cb731e2"
train_pred = knn_pipe.predict(X_train) #prediction on trained data
f1_score(y_train, train_pred)
# + [markdown] id="cd818db8"
# precision score without tunning is better than after tunning, so the model will be choose is Random Forest Classifier without tuning.
# + id="4c4e643c" outputId="c047615e-7c49-4668-b752-bb0c71cace24"
# classification report and plotting confusion matrix
knn_cm = confusion_matrix(y_test,y_pred_estimator)
knn_clr = classification_report(y_test,y_pred_estimator, target_names=["NEGATIVE", "POSITIVE"])
plt.figure(figsize=(6, 6))
sns.heatmap(knn_cm, annot=True, fmt='g', vmin=0, cmap='Blues', cbar=False)
plt.xticks(ticks=np.arange(2) + 0.5, labels=["NEGATIVE", "POSITIVE"])
plt.yticks(ticks=np.arange(2) + 0.5, labels=["NEGATIVE", "POSITIVE"])
plt.xlabel("Predicted")
plt.ylabel("Actual")
plt.title("KNN_Confusion Matrix")
plt.show()
print("KNN_Confusion Matrix:\n----------------------\n", knn_cm)
print()
print("KNN_Classification Report:\n----------------------\n", knn_clr)
# + id="86d39f7a"
# + id="57829afb" outputId="faf0d07d-66a5-47b1-e85a-f0403dc241af"
# classification report and plotting confusion matrix
rf_cm = confusion_matrix(y_test,rf_y_pred_estimator)
rf_clr = classification_report(y_test,rf_y_pred_estimator, target_names=["NEGATIVE", "POSITIVE"])
plt.figure(figsize=(6, 6))
sns.heatmap(rf_cm, annot=True, fmt='g', vmin=0, cmap='Blues', cbar=False)
plt.xticks(ticks=np.arange(2) + 0.5, labels=["NEGATIVE", "POSITIVE"])
plt.yticks(ticks=np.arange(2) + 0.5, labels=["NEGATIVE", "POSITIVE"])
plt.xlabel("Predicted")
plt.ylabel("Actual")
plt.title("RF_Confusion Matrix")
plt.show()
print("RF_Confusion Matrix:\n----------------------\n", rf_cm)
print()
print("RF_Classification Report:\n----------------------\n", rf_clr)
# + id="5bb2403c" outputId="60b6e338-cb06-4caa-ddf3-0e787968c657"
# classification report and plotting confusion matrix
XGB_cm = confusion_matrix(y_test,XGB_y_pred_estimator)
XGB_clr = classification_report(y_test,XGB_y_pred_estimator, target_names=["NEGATIVE", "POSITIVE"])
plt.figure(figsize=(6, 6))
sns.heatmap(XGB_cm, annot=True, fmt='g', vmin=0, cmap='Blues', cbar=False)
plt.xticks(ticks=np.arange(2) + 0.5, labels=["NEGATIVE", "POSITIVE"])
plt.yticks(ticks=np.arange(2) + 0.5, labels=["NEGATIVE", "POSITIVE"])
plt.xlabel("Predicted")
plt.ylabel("Actual")
plt.title("XGB_Confusion Matrix")
plt.show()
print("XGB_Confusion Matrix:\n----------------------\n", XGB_cm)
print()
print("XGB_Classification Report:\n----------------------\n", XGB_clr)
# + id="33aa5169" active=""
# TN: Customers who are predicted will not respond to the campaign, in fact it does not respond.
# TP: Customers are predicted to respond the campaign, actually it does respond.
# FP: Customers who are predicted to respond the campaign, actually do not respond.
# FN: Predicted Customers do not respond to the campaign, actually respond.
#
# Error that occurred:
#
# FN: Wrong prediction, the company only loses prospective customers, but not financial losses
# FP: the company loses more such as time, energy and financial, because it has prepared everything for the campaign to people, but those people are not response.
# So the most influential mistake for financial losses is FP
# + id="5cdd1d5f" outputId="cbfe9dd6-0fa1-4d8f-e3e1-592932f7e656"
TP = knn_cm[1,1] # true positive
TN = knn_cm[0,0] # true negatives
FP = knn_cm[0,1] # false positives
FN = knn_cm[1,0] # false negatives
FP
# + id="d3cb6f0c" active=""
# Sensitivity is the metric that evaluates a model's ability to predict true positives of each available category. Specificity is the metric that evaluates a model's ability to predict true negatives of each available category.
# + id="e33fafdd" outputId="a79db84b-ce00-487e-95fd-d4683c855008"
# Let's see the sensitivity of our Random Forest Classifier model
print('Sensivity :')
TP / float(TP+FN)
# + id="af7b1007" outputId="18befa76-c302-4f2b-e822-98fa15ba0d89"
# Let us calculate specificity
print('Specificity :')
TN / float(TN+FP)
# + id="74e2151d" active=""
# We can see that the value of Sensitivity and Specificity are close to each other
# + id="fa6d9125" outputId="55d6bf08-25ea-4345-9bc2-238c6c8020b2"
# Overall accuracy
ACC = float(TP+TN)/float(TP+FP+FN+TN)
ACC
# + id="4ea4e38a"
# + [markdown] id="5821c258-2398-4d7a-aacd-f9a91ef567be"
# Export the tested model to a pickle file
# + id="e15dda08-22c6-4da5-9f56-dd57495e102a"
# use pickle to store the model onto our disk.
filename = 'response_model.p'
#model_pickle = pickle.dump( grid_search, open( "loan_model.p", "wb" ) )
pickle.dump(knn_pipe, open(filename, 'wb'))
# + id="7eafd9e9-a01d-4438-b894-41c32686fd76" outputId="d411462c-af17-4430-cce5-87d92b28d715"
#When we load the pickle back:
loaded_model = pickle.load(open(filename, 'rb'))
result = loaded_model.score(X_test, y_test)
print(result)
# + id="1b0f26ea-68fd-464e-a11c-aaa66bb22502" outputId="74649c4b-053d-4bad-b857-2d096ca2298f"
# use our trained model to predict.
# calculate the predictions for each class
X_predict = knn_pipe.predict(X_test)
# create the solution dataframe
Result = pd.DataFrame(data=X_predict, columns=['Response'])
Result = Result['Response'].map({0:'N', 1:'Y'})
Result
# + id="2f2b6baa-44cd-441b-91c8-2554a8b4e767"
# save the solution as an csv file
Result.to_csv('Solution1.csv', header=True)
# + [markdown] id="6e597242-7c6b-4dcd-8c0d-d5fafe43dfc7"
# Export solution information to a file that will be useful later on in our prediction application
# + id="4b06ca34-39b2-438b-8371-e5d4500a6cb6"
import json
columns = {
'data_columns' : [col.lower() for col in X.columns]
}
with open("columns.json","w") as f:
f.write(json.dumps(columns))
# + id="e60ebd37-55e7-490a-b7fe-374790995170"
# + id="9e5c1cdd-f343-465c-8beb-17701db1ebd5"
df = pd.read_csv('df_copy.csv')
# + id="b72f6fc9-3f18-48fc-8893-82f90d97d684"
y_test_pred = knn_pipe.predict(df)
# + id="a9c0750e-27d9-4967-8235-8d8e0e18bd66"
df['Predict_Response'] = y_test_pred
df_final = df[['ID', 'Response', 'Predict_Response']]
# + id="4ca15c21-2f05-47b4-94bd-30b9b48733ec"
df_final['Response'] = df_final['Response'].map({0:'N', 1:'Y'})
# + id="6fdba4c0-6ce6-4e69-bafd-f08dfef935bb"
df_final['Predict_Response'] = df_final['Predict_Response'].map({0:'N', 1:'Y'})
# + id="afafe35f-f671-47bb-91b6-db9f61436bfc"
df_final.to_csv('Solution_2.csv', index=False)
# + id="0b215c3a-cbfb-46cc-ba43-2ac4ddd48f4a" outputId="10481560-fc17-46d1-bec4-d7bb781a2b16"
df_final.head()
# + id="d2d618b4-c6e1-4831-a2e9-5bb4be8fffe8" outputId="063650a1-b1c2-4f7f-c0d7-8583fb882139"
df_final.shape
# + id="a5a1d3bf-0dca-4054-9e05-c17b9fbd52da"
# + id="5dd794a5-aa2e-48f8-8fde-d7669be53806"
# + id="53b5ae45-2dd3-451a-a623-49542fc0bf9a"
# + id="6773cf96-de53-4dc5-adc4-97b75c241919"
|
New_model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 深度学习简介
#
# 你可能已经接触过编程,并开发过一两款程序。同时你可能读过关于深度学习或者机器学习的铺天盖地的报道,尽管很多时候它们被赋予了更广义的名字:人工智能。实际上,或者说幸运的是,大部分程序并不需要深度学习或者是更广义上的人工智能技术。例如,如果我们要为一台微波炉编写一个用户界面,只需要一点儿工夫我们便能设计出十几个按钮以及一系列能精确描述微波炉在各种情况下的表现的规则。再比如,假设我们要编写一个电子邮件客户端。这样的程序比微波炉要复杂一些,但我们还是可以沉下心来一步一步思考:客户端的用户界面将需要几个输入框来接受收件人、主题、邮件正文等,程序将监听键盘输入并写入一个缓冲区,然后将它们显示在相应的输入框中。当用户点击“发送”按钮时,我们需要检查收件人邮箱地址的格式是否正确,并检查邮件主题是否为空,或在主题为空时警告用户,而后用相应的协议传送邮件。
#
# 值得注意的是,在以上两个例子中,我们都不需要收集真实世界中的数据,也不需要系统地提取这些数据的特征。只要有充足的时间,我们的常识与编程技巧已经足够让我们完成任务。
#
# 与此同时,我们很容易就能找到一些连世界上最好的程序员也无法仅用编程技巧解决的简单问题。例如,假设我们想要编写一个判定一张图像中有没有猫的程序。这件事听起来好像很简单,对不对?程序只需要对每张输入图像输出“真”(表示有猫)或者“假”(表示无猫)即可。但令人惊讶的是,即使是世界上最优秀的计算机科学家和程序员也不懂如何编写这样的程序。
#
# 我们该从哪里入手呢?我们先进一步简化这个问题:若假设所有图像的高和宽都是同样的400像素大小,一个像素由红绿蓝三个值构成,那么一张图像就由近50万个数值表示。那么哪些数值隐藏着我们需要的信息呢?是所有数值的平均数,还是四个角的数值,抑或是图像中的某一个特别的点?事实上,要想解读图像中的内容,需要寻找仅仅在结合成千上万的数值时才会出现的特征,如边缘、质地、形状、眼睛、鼻子等,最终才能判断图像中是否有猫。
#
# 一种解决以上问题的思路是逆向思考。与其设计一个解决问题的程序,不如从最终的需求入手来寻找一个解决方案。事实上,这也是目前的机器学习和深度学习应用共同的核心思想:我们可以称其为“用数据编程”。与其枯坐在房间里思考怎么设计一个识别猫的程序,不如利用人类肉眼在图像中识别猫的能力。我们可以收集一些已知包含猫与不包含猫的真实图像,然后我们的目标就转化成如何从这些图像入手得到一个可以推断出图像中是否有猫的函数。这个函数的形式通常通过我们的知识来针对特定问题选定。例如,我们使用一个二次函数来判断图像中是否有猫,但是像二次函数系数值这样的函数参数的具体值则是通过数据来确定。
#
# 通俗来说,机器学习是一门讨论各式各样的适用于不同问题的函数形式,以及如何使用数据来有效地获取函数参数具体值的学科。深度学习是指机器学习中的一类函数,它们的形式通常为多层神经网络。近年来,仰仗着大数据集和强大的硬件,深度学习已逐渐成为处理图像、文本语料和声音信号等复杂高维度数据的主要方法。
#
# 我们现在正处于一个程序设计得到深度学习的帮助越来越多的时代。这可以说是计算机科学历史上的一个分水岭。举个例子,深度学习已经在你的手机里:拼写校正、语音识别、认出社交媒体照片里的好友们等。得益于优秀的算法、快速而廉价的算力、前所未有的大量数据以及强大的软件工具,如今大多数软件工程师都有能力建立复杂的模型来解决十年前连最优秀的科学家都觉得棘手的问题。
#
# 本书希望能帮助读者进入深度学习的浪潮中。我们希望结合数学、代码和样例让深度学习变得触手可及。本书不要求具有高深的数学或编程背景,我们将随着章节的发展逐一解释所需要的知识。更值得一提的是,本书的每一节都是一个可以独立运行的Jupyter记事本。读者可以从网上获得这些记事本,并且可以在个人电脑或云端服务器上执行它们。这样读者就可以随意改动书中的代码并得到及时反馈。我们希望本书能帮助和启发新一代的程序员、创业者、统计学家、生物学家,以及所有对深度学习感兴趣的人。
#
#
# ## 起源
#
# 虽然深度学习似乎是最近几年刚兴起的名词,但它所基于的神经网络模型和用数据编程的核心思想已经被研究了数百年。自古以来,人类就一直渴望能从数据中分析出预知未来的窍门。实际上,数据分析正是大部分自然科学的本质,我们希望从日常的观测中提取规则,并找寻不确定性。
#
# 早在17世纪,[雅各比·伯努利(1655--1705)](https://en.wikipedia.org/wiki/Jacob_Bernoulli)提出了描述只有两种结果的随机过程(如抛掷一枚硬币)的伯努利分布。大约一个世纪之后,[卡尔·弗里德里希·高斯(1777--1855)](https://en.wikipedia.org/wiki/Carl_Friedrich_Gauss)发明了今日仍广泛用在从保险计算到医学诊断等领域的最小二乘法。概率论、统计学和模式识别等工具帮助自然科学的实验学家们从数据回归到自然定律,从而发现了如欧姆定律(描述电阻两端电压和流经电阻电流关系的定律)这类可以用线性模型完美表达的一系列自然法则。
#
# 即使是在中世纪,数学家也热衷于利用统计学来做出估计。例如,在[雅各比·科贝尔(1460--1533)](https://www.maa.org/press/periodicals/convergence/mathematical-treasures-jacob-kobels-geometry)的几何书中记载了使用16名男子的平均脚长来估计男子的平均脚长。
#
# 
#
# 如图1.1所示,在这个研究中,16位成年男子被要求在离开教堂时站成一排并把脚贴在一起,而后他们脚的总长度除以16得到了一个估计:这个数字大约相当于今日的一英尺。这个算法之后又被改进,以应对特异形状的脚:最长和最短的脚不计入,只对剩余的脚长取平均值,即裁剪平均值的雏形。
#
# 现代统计学在20世纪的真正起飞要归功于数据的收集和发布。统计学巨匠之一[罗纳德·费雪(1890--1962)](https://en.wikipedia.org/wiki/Ronald_Fisher)对统计学理论和统计学在基因学中的应用功不可没。他发明的许多算法和公式,例如线性判别分析和费雪信息,仍经常被使用。即使是他在1936年发布的Iris数据集,仍然偶尔被用于演示机器学习算法。
#
# [克劳德·香农(1916--2001)](https://en.wikipedia.org/wiki/Claude_Shannon)的信息论以及[阿兰·图灵 (1912--1954)](https://en.wikipedia.org/wiki/Allan_Turing)的计算理论也对机器学习有深远影响。图灵在他著名的论文[《计算机器与智能》](https://www.jstor.org/stable/2251299)中提出了“机器可以思考吗?”这样一个问题 [1]。在他描述的“图灵测试”中,如果一个人在使用文本交互时不能区分他的对话对象到底是人类还是机器的话,那么即可认为这台机器是有智能的。时至今日,智能机器的发展可谓日新月异。
#
# 另一个对深度学习有重大影响的领域是神经科学与心理学。既然人类显然能够展现出智能,那么对于解释并逆向工程人类智能机理的探究也在情理之中。最早的算法之一是由[唐纳德·赫布(1904--1985)](https://en.wikipedia.org/wiki/Donald_O._Hebb)正式提出的。在他开创性的著作[《行为的组织》](http://s-f-walker.org.uk/pubsebooks/pdfs/The_Organization_of_Behavior-Donald_O._Hebb.pdf)中,他提出神经是通过正向强化来学习的,即赫布理论 [2]。赫布理论是感知机学习算法的原型,并成为支撑今日深度学习的随机梯度下降算法的基石:强化合意的行为、惩罚不合意的行为,最终获得优良的神经网络参数。
#
# 来源于生物学的灵感是神经网络名字的由来。这类研究者可以追溯到一个多世纪前的[亚历山大·贝恩(1818--1903)](https://en.wikipedia.org/wiki/Alexander_Bain)和[查尔斯·斯科特·谢灵顿(1857--1952)](https://en.wikipedia.org/wiki/Charles_Scott_Sherrington)。研究者们尝试组建模仿神经元互动的计算电路。随着时间发展,神经网络的生物学解释被稀释,但仍保留了这个名字。时至今日,绝大多数神经网络都包含以下的核心原则。
#
# * 交替使用线性处理单元与非线性处理单元,它们经常被称为“层”。
# * 使用链式法则(即反向传播)来更新网络的参数。
#
# 在最初的快速发展之后,自约1995年起至2005年,大部分机器学习研究者的视线从神经网络上移开了。这是由于多种原因。首先,训练神经网络需要极强的计算力。尽管20世纪末内存已经足够,计算力却不够充足。其次,当时使用的数据集也相对小得多。费雪在1936年发布的的Iris数据集仅有150个样本,并被广泛用于测试算法的性能。具有6万个样本的MNIST数据集在当时已经被认为是非常庞大了,尽管它如今已被认为是典型的简单数据集。由于数据和计算力的稀缺,从经验上来说,如核方法、决策树和概率图模型等统计工具更优。它们不像神经网络一样需要长时间的训练,并且在强大的理论保证下提供可以预测的结果。
#
# ## 发展
#
# 互联网的崛起、价廉物美的传感器和低价的存储器令我们越来越容易获取大量数据。加之便宜的计算力,尤其是原本为电脑游戏设计的GPU的出现,上文描述的情况改变了许多。一瞬间,原本被认为不可能的算法和模型变得触手可及。这样的发展趋势从如下表格中可见一斑。
#
# |年代|数据样本个数|内存|每秒浮点计算数|
# |:--|:-:|:-:|:-:|
# |1970|100(Iris)|1 KB|100 K(Intel 8080)|
# |1980|1 K(波士顿房价)|100 KB|1 M(Intel 80186)|
# |1990|10 K(手写字符识别)|10 MB|10 M(Intel 80486)|
# |2000|10 M(网页)|100 MB|1 G(Intel Core)|
# |2010|10 G(广告)|1 GB|1 T(NVIDIA C2050)|
# |2020|1 T(社交网络)|100 GB|1 P(NVIDIA DGX-2)|
#
# 很显然,存储容量没能跟上数据量增长的步伐。与此同时,计算力的增长又盖过了数据量的增长。这样的趋势使得统计模型可以在优化参数上投入更多的计算力,但同时需要提高存储的利用效率,例如使用非线性处理单元。这也相应导致了机器学习和统计学的最优选择从广义线性模型及核方法变化为深度多层神经网络。这样的变化正是诸如多层感知机、卷积神经网络、长短期记忆循环神经网络和Q学习等深度学习的支柱模型在过去10年从坐了数十年的冷板凳上站起来被“重新发现”的原因。
#
# 近年来在统计模型、应用和算法上的进展常被拿来与寒武纪大爆发(历史上物种数量大爆发的一个时期)做比较。但这些进展不仅仅是因为可用资源变多了而让我们得以用新瓶装旧酒。下面的列表仅仅涵盖了近十年来深度学习长足发展的部分原因。
#
# * 优秀的容量控制方法,如丢弃法,使大型网络的训练不再受制于过拟合(大型神经网络学会记忆大部分训练数据的行为) [3]。这是靠在整个网络中注入噪声而达到的,如训练时随机将权重替换为随机的数字 [4]。
#
# * 注意力机制解决了另一个困扰统计学超过一个世纪的问题:如何在不增加参数的情况下扩展一个系统的记忆容量和复杂度。注意力机制使用了一个可学习的指针结构来构建出一个精妙的解决方法 [5]。也就是说,与其在像机器翻译这样的任务中记忆整个句子,不如记忆指向翻译的中间状态的指针。由于生成译文前不需要再存储整句原文的信息,这样的结构使准确翻译长句变得可能。
#
# * 记忆网络 [6]和神经编码器—解释器 [7]这样的多阶设计使得针对推理过程的迭代建模方法变得可能。这些模型允许重复修改深度网络的内部状态,这样就能模拟出推理链条上的各个步骤,就好像处理器在计算过程中修改内存一样。
#
# * 另一个重大发展是生成对抗网络的发明 [8]。传统上,用在概率分布估计和生成模型上的统计方法更多地关注于找寻正确的概率分布,以及正确的采样算法。生成对抗网络的关键创新在于将采样部分替换成了任意的含有可微分参数的算法。这些参数将被训练到使辨别器不能再分辨真实的和生成的样本。生成对抗网络可使用任意算法来生成输出的这一特性为许多技巧打开了新的大门。例如生成奔跑的斑马 [9]和生成名流的照片 [10] 都是生成对抗网络发展的见证。
#
# * 许多情况下单个GPU已经不能满足在大型数据集上进行训练的需要。过去10年内我们构建分布式并行训练算法的能力已经有了极大的提升。设计可扩展算法的最大瓶颈在于深度学习优化算法的核心:随机梯度下降需要相对更小的批量。与此同时,更小的批量也会降低GPU的效率。如果使用1,024个GPU,每个GPU的批量大小为32个样本,那么单步训练的批量大小将是32,000个以上。近年来李沐 [11]、Yang You等人 [12]以及Xianyan Jia等人 [13]的工作将批量大小增至多达64,000个样例,并把在ImageNet数据集上训练ResNet-50模型的时间降到了7分钟。与之对比,最初的训练时间需要以天来计算。
#
# * 并行计算的能力也为至少在可以采用模拟情况下的强化学习的发展贡献了力量。并行计算帮助计算机在围棋、雅达利游戏、星际争霸和物理模拟上达到了超过人类的水准。
#
# * 深度学习框架也在传播深度学习思想的过程中扮演了重要角色。[Caffe](https://github.com/BVLC/caffe)、 [Torch](https://github.com/torch)和[Theano](https://github.com/Theano/Theano)这样的第一代框架使建模变得更简单。许多开创性的论文都用到了这些框架。如今它们已经被[TensorFlow](https://github.com/tensorflow/tensorflow)(经常是以高层API [Keras](https://github.com/keras-team/keras)的形式被使用)、[CNTK](https://github.com/Microsoft/CNTK)、 [Caffe 2](https://github.com/caffe2/caffe2) 和[Apache MXNet](https://github.com/apache/incubator-mxnet)所取代。第三代,即命令式深度学习框架,是由用类似NumPy的语法来定义模型的 [Chainer](https://github.com/chainer/chainer)所开创的。这样的思想后来被 [PyTorch](https://github.com/pytorch/pytorch)和MXNet的[Gluon API](https://github.com/apache/incubator-mxnet) 采用,后者也正是本书用来教学深度学习的工具。
#
# 系统研究者负责构建更好的工具,统计学家建立更好的模型。这样的分工使工作大大简化。举例来说,在2014年时,训练一个逻辑回归模型曾是卡内基梅隆大学布置给机器学习方向的新入学博士生的作业问题。时至今日,这个问题只需要少于10行的代码便可以完成,普通的程序员都可以做到。
#
# ## 成功案例
#
# 长期以来机器学习总能完成其他方法难以完成的目标。例如,自20世纪90年代起,邮件的分拣就开始使用光学字符识别。实际上这正是知名的MNIST和USPS手写数字数据集的来源。机器学习也是电子支付系统的支柱,可以用于读取银行支票、进行授信评分以及防止金融欺诈。机器学习算法在网络上被用来提供搜索结果、个性化推荐和网页排序。虽然长期处于公众视野之外,但是机器学习已经渗透到了我们工作和生活的方方面面。直到近年来,在此前认为无法被解决的问题以及直接关系到消费者的问题上取得突破性进展后,机器学习才逐渐变成公众的焦点。这些进展基本归功于深度学习。
#
# * 苹果公司的Siri、亚马逊的Alexa和谷歌助手一类的智能助手能以可观的准确率回答口头提出的问题,甚至包括从简单的开关灯具(对残疾群体帮助很大)到提供语音对话帮助。智能助手的出现或许可以作为人工智能开始影响我们生活的标志。
#
# * 智能助手的关键是需要能够精确识别语音,而这类系统在某些应用上的精确度已经渐渐增长到可以与人类比肩 [14]。
#
# * 物体识别也经历了漫长的发展过程。在2010年从图像中识别出物体的类别仍是一个相当有挑战性的任务。当年日本电气、伊利诺伊大学香槟分校和罗格斯大学团队在ImageNet基准测试上取得了28%的前五错误率 [15]。到2017年,这个数字降低到了2.25% [16]。研究人员在鸟类识别和皮肤癌诊断上,也取得了同样惊世骇俗的成绩。
#
# * 游戏曾被认为是人类智能最后的堡垒。自使用时间差分强化学习玩双陆棋的TD-Gammon开始,算法和算力的发展催生了一系列在游戏上使用的新算法。与双陆棋不同,国际象棋有更复杂的状态空间和更多的可选动作。“深蓝”用大量的并行、专用硬件和游戏树的高效搜索打败了加里·卡斯帕罗夫 [17]。围棋因其庞大的状态空间被认为是更难的游戏,AlphaGo在2016年用结合深度学习与蒙特卡罗树采样的方法达到了人类水准 [18]。对德州扑克游戏而言,除了巨大的状态空间之外,更大的挑战是游戏的信息并不完全可见,例如看不到对手的牌。而“冷扑大师”用高效的策略体系超越了人类玩家的表现 [19]。以上的例子都体现出了先进的算法是人工智能在游戏上的表现提升的重要原因。
#
# * 机器学习进步的另一个标志是自动驾驶汽车的发展。尽管距离完全的自主驾驶还有很长的路要走,但诸如[Tesla](http://www.tesla.com)、[NVIDIA](http://www.nvidia.com)、 [MobilEye](http://www.mobileye.com)和[Waymo](http://www.waymo.com)这样的公司发布的具有部分自主驾驶功能的产品展示出了这个领域巨大的进步。完全自主驾驶的难点在于它需要将感知、思考和规则整合在同一个系统中。目前,深度学习主要被应用在计算机视觉的部分,剩余的部分还是需要工程师们的大量调试。
#
# 以上列出的仅仅是近年来深度学习所取得的成果的冰山一角。机器人学、物流管理、计算生物学、粒子物理学和天文学近年来的发展也有一部分要归功于深度学习。可以看到,深度学习已经逐渐演变成一个工程师和科学家皆可使用的普适工具。
#
#
# ## 特点
#
# 在描述深度学习的特点之前,我们先回顾并概括一下机器学习和深度学习的关系。机器学习研究如何使计算机系统利用经验改善性能。它是人工智能领域的分支,也是实现人工智能的一种手段。在机器学习的众多研究方向中,表征学习关注如何自动找出表示数据的合适方式,以便更好地将输入变换为正确的输出,而本书要重点探讨的深度学习是具有多级表示的表征学习方法。在每一级(从原始数据开始),深度学习通过简单的函数将该级的表示变换为更高级的表示。因此,深度学习模型也可以看作是由许多简单函数复合而成的函数。当这些复合的函数足够多时,深度学习模型就可以表达非常复杂的变换。
#
# 深度学习可以逐级表示越来越抽象的概念或模式。以图像为例,它的输入是一堆原始像素值。深度学习模型中,图像可以逐级表示为特定位置和角度的边缘、由边缘组合得出的花纹、由多种花纹进一步汇合得到的特定部位的模式等。最终,模型能够较容易根据更高级的表示完成给定的任务,如识别图像中的物体。值得一提的是,作为表征学习的一种,深度学习将自动找出每一级表示数据的合适方式。
#
# 因此,深度学习的一个外在特点是端到端的训练。也就是说,并不是将单独调试的部分拼凑起来组成一个系统,而是将整个系统组建好之后一起训练。比如说,计算机视觉科学家之前曾一度将特征抽取与机器学习模型的构建分开处理,像是Canny边缘探测 [20] 和SIFT特征提取 [21] 曾占据统治性地位达10年以上,但这也就是人类能找到的最好方法了。当深度学习进入这个领域后,这些特征提取方法就被性能更强的自动优化的逐级过滤器替代了。
#
# 相似地,在自然语言处理领域,词袋模型多年来都被认为是不二之选 [22]。词袋模型是将一个句子映射到一个词频向量的模型,但这样的做法完全忽视了单词的排列顺序或者句中的标点符号。不幸的是,我们也没有能力来手工抽取更好的特征。但是自动化的算法反而可以从所有可能的特征中搜寻最好的那个,这也带来了极大的进步。例如,语义相关的词嵌入能够在向量空间中完成如下推理:“柏林 - 德国 + 中国 = 北京”。可以看出,这些都是端到端训练整个系统带来的效果。
#
# 除端到端的训练以外,我们也正在经历从含参数统计模型转向完全无参数的模型。当数据非常稀缺时,我们需要通过简化对现实的假设来得到实用的模型。当数据充足时,我们就可以用能更好地拟合现实的无参数模型来替代这些含参数模型。这也使我们可以得到更精确的模型,尽管需要牺牲一些可解释性。
#
# 相对其它经典的机器学习方法而言,深度学习的不同在于:对非最优解的包容、对非凸非线性优化的使用,以及勇于尝试没有被证明过的方法。这种在处理统计问题上的新经验主义吸引了大量人才的涌入,使得大量实际问题有了更好的解决方案。尽管大部分情况下需要为深度学习修改甚至重新发明已经存在数十年的工具,但是这绝对是一件非常有意义并令人兴奋的事。
#
# 最后,深度学习社区长期以来以在学术界和企业之间分享工具而自豪,并开源了许多优秀的软件库、统计模型和预训练网络。正是本着开放开源的精神,本书的内容和基于它的教学视频可以自由下载和随意分享。我们致力于为所有人降低学习深度学习的门槛,并希望大家从中获益。
#
#
# ## 小结
#
# * 机器学习研究如何使计算机系统利用经验改善性能。它是人工智能领域的分支,也是实现人工智能的一种手段。
# * 作为机器学习的一类,表征学习关注如何自动找出表示数据的合适方式。
# * 深度学习是具有多级表示的表征学习方法。它可以逐级表示越来越抽象的概念或模式。
# * 深度学习所基于的神经网络模型和用数据编程的核心思想实际上已经被研究了数百年。
# * 深度学习已经逐渐演变成一个工程师和科学家皆可使用的普适工具。
#
#
# ## 练习
#
# * 你现在正在编写的代码有没有可以被“学习”的部分,也就是说,是否有可以被机器学习改进的部分?
# * 你在生活中有没有这样的场景:虽有许多展示如何解决问题的样例,但缺少自动解决问题的算法?它们也许是深度学习的最好猎物。
# * 如果把人工智能的发展看作是新一次工业革命,那么深度学习和数据的关系是否像是蒸汽机与煤炭的关系呢?为什么?
# * 端到端的训练方法还可以用在哪里?物理学,工程学还是经济学?
# * 为什么应该让深度网络模仿人脑结构?为什么不该让深度网络模仿人脑结构?
#
#
#
# ## 参考文献
#
# [1] Machinery, C. (1950). Computing machinery and intelligence-AM Turing. Mind, 59(236), 433.
#
# [2] <NAME>. (1949). The organization of behavior; a neuropsycholocigal theory. A Wiley Book in Clinical Psychology., 62-78.
#
# [3] <NAME>., <NAME>., <NAME>., <NAME>., & <NAME>. (2014). Dropout: a simple way to prevent neural networks from overfitting. The Journal of Machine Learning Research, 15(1), 1929-1958.
#
# [4] <NAME>. (1995). Training with noise is equivalent to Tikhonov regularization. Neural computation, 7(1), 108-116.
#
# [5] <NAME>., <NAME>., & <NAME>. (2014). Neural machine translation by jointly learning to align and translate. arXiv preprint arXiv:1409.0473.
#
# [6] <NAME>., <NAME>., & <NAME>. (2015). End-to-end memory networks. In Advances in neural information processing systems (pp. 2440-2448).
#
# [7] <NAME>., & <NAME>. (2015). Neural programmer-interpreters. arXiv preprint arXiv:1511.06279.
#
# [8] <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., … & <NAME>. (2014). Generative adversarial nets. In Advances in neural information processing systems (pp. 2672-2680).
#
# [9] <NAME>., <NAME>., <NAME>., & <NAME>. (2017). Unpaired image-to-image translation using cycle-consistent adversarial networks. arXiv preprint.
#
# [10] <NAME>., <NAME>., <NAME>., & <NAME>. (2017). Progressive growing of gans for improved quality, stability, and variation. arXiv preprint arXiv:1710.10196.
#
# [11] <NAME>. (2017). Scaling Distributed Machine Learning with System and Algorithm Co-design (Doctoral dissertation, PhD thesis, Intel).
#
# [12] <NAME>., <NAME>., & <NAME>. Large batch training of convolutional networks. ArXiv e-prints.
#
# [13] <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., … & <NAME>. (2018). Highly Scalable Deep Learning Training System with Mixed-Precision: Training ImageNet in Four Minutes. arXiv preprint arXiv:1807.11205.
#
# [14] <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., … & <NAME>. (2017, March). The Microsoft 2016 conversational speech recognition system. In Acoustics, Speech and Signal Processing (ICASSP), 2017 IEEE International Conference on (pp. 5255-5259). IEEE.
#
# [15] <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., … & <NAME>. (2010). Imagenet classification: fast descriptor coding and large-scale svm training. Large scale visual recognition challenge.
#
# [16] <NAME>., <NAME>., & <NAME>. (2017). Squeeze-and-excitation networks. arXiv preprint arXiv:1709.01507, 7.
#
# [17] <NAME>., <NAME>., & <NAME>. (2002). Deep blue. Artificial intelligence, 134(1-2), 57-83.
#
# [18] <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., … & <NAME>. (2016). Mastering the game of Go with deep neural networks and tree search. nature, 529(7587), 484.
#
# [19] <NAME>., & <NAME>. (2017, August). Libratus: The superhuman ai for no-limit poker. In Proceedings of the Twenty-Sixth International Joint Conference on Artificial Intelligence.
#
# [20] <NAME>. (1986). A computational approach to edge detection. IEEE Transactions on pattern analysis and machine intelligence, (6), 679-698.
#
# [21] <NAME>. (2004). Distinctive image features from scale-invariant keypoints. International journal of computer vision, 60(2), 91-110.
#
# [22] <NAME>., & <NAME>. (1986). Introduction to modern information retrieval.
#
# ## 扫码直达[讨论区](https://discuss.gluon.ai/t/topic/746)
#
# 
|
1.chapter_introduction/deep-learning-intro.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/susanquiros/Big-Data-Challenge-/blob/main/main.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="-ryXKndegAwV"
# + colab={"base_uri": "https://localhost:8080/"} id="xvwFg_bZtHgw" outputId="8423d691-0ca5-4295-d1be-75acba3f81fc"
import os
# Find the latest version of spark 3.0 from http://www.apache.org/dist/spark/ and enter as the spark version
# For example:
# spark_version = 'spark-3.0.3'
spark_version = 'spark-3.1.2'
os.environ['SPARK_VERSION']=spark_version
# Install Spark and Java
# !apt-get update
# !apt-get install openjdk-8-jdk-headless -qq > /dev/null
# !wget -q http://www.apache.org/dist/spark/$SPARK_VERSION/$SPARK_VERSION-bin-hadoop2.7.tgz
# !tar xf $SPARK_VERSION-bin-hadoop2.7.tgz
# !pip install -q findspark
# Set Environment Variables
os.environ["JAVA_HOME"] = "/usr/lib/jvm/java-8-openjdk-amd64"
os.environ["SPARK_HOME"] = f"/content/{spark_version}-bin-hadoop2.7"
# Start a SparkSession
import findspark
findspark.init()
# + id="Tn78X89CtSqq"
from pyspark.sql import SparkSession
spark = SparkSession.builder.appName("CloudETL").config("spark.driver.extraClassPath","/content/postgresql-42.2.9.jar").getOrCreate()
# + colab={"base_uri": "https://localhost:8080/"} id="78jjwR5_zzBo" outputId="dc466bd1-4ca1-406f-e902-22a4fdb86603"
from pyspark import SparkFiles
url = 'https://s3.amazonaws.com/amazon-reviews-pds/tsv/sample_us.tsv'
spark.sparkContext.addFile(url)
df = spark.read.csv(SparkFiles.get('sample_us.tsv'), sep= '\t', header=True, inferSchema=True)
#showing the dataframe
df.show()
# + id="XfVDzGl33jvz"
#Schema for review_id_table
# review_id TEXT PRIMARY KEY NOT NULL,
# customer_id INTEGER,
# product_id TEXT,
# product_parent INTEGER,
# review_date DATE -- this should be in the formate yyyy-mm-dd
# + colab={"base_uri": "https://localhost:8080/"} id="Ddjenzny2tDR" outputId="d7f23ac7-4adf-4569-d165-5692d3e114be"
df.dtypes
# + id="y5fLFX2B3VkC"
#converting review_date to date format
|
main.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={"duration": 0.032637, "end_time": "2020-09-25T19:48:19.468299", "exception": false, "start_time": "2020-09-25T19:48:19.435662", "status": "completed"} tags=[] id="PB3EzyAeZje2"
# # **Sentiment Analysis with Deep Learning using BERT**
#
#
# ## **What is BERT?**
#
# BERT is a large-scale transformer-based Language Model that can be finetuned for a variety of tasks.
#
# For more information, the original paper can be found here (https://arxiv.org/abs/1810.04805).
#
# HuggingFace documentation (https://huggingface.co/transformers/model_doc/bert.html)
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" papermill={"duration": 0.049233, "end_time": "2020-09-25T19:48:19.549659", "exception": false, "start_time": "2020-09-25T19:48:19.500426", "status": "completed"} tags=[] id="U_f7Wnb-Zje6"
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python Docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the read-only "../input/" directory
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
# You can write up to 5GB to the current directory (/kaggle/working/) that gets preserved as output when you create a version using "Save & Run All"
# You can also write temporary files to /kaggle/temp/, but they won't be saved outside of the current session
# + [markdown] papermill={"duration": 0.04373, "end_time": "2020-09-25T19:48:19.638706", "exception": false, "start_time": "2020-09-25T19:48:19.594976", "status": "completed"} tags=[] id="2kNXLTeVZje9"
# ## 1: Exploratory Data Analysis and Preprocessing
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a" papermill={"duration": 1.602442, "end_time": "2020-09-25T19:48:21.286204", "exception": false, "start_time": "2020-09-25T19:48:19.683762", "status": "completed"} tags=[] id="l8CMcl8qZje9"
import torch
from tqdm.notebook import tqdm
# + papermill={"duration": 0.067018, "end_time": "2020-09-25T19:48:21.384464", "exception": false, "start_time": "2020-09-25T19:48:21.317446", "status": "completed"} tags=[] id="oJCcya0-Zje9"
df = pd.read_csv('corpus.csv',
names=['text', 'category'])
df = df[1:]
df.insert(0, 'id', range(1, 1 + len(df)))
df.set_index('id', inplace=True)
df = df[df.category != 'neutral']
# + colab={"base_uri": "https://localhost:8080/"} id="D8F9vJeSjMn1" outputId="aba49a68-c868-4d2b-dee1-9a71cdad7f8c"
# !pip install translate
# !pip install enchant
# + colab={"base_uri": "https://localhost:8080/"} id="xw1PG12YjN-s" outputId="6efe28a2-ff59-4c1e-8e4e-60e298a2bdd3"
import nltk
nltk.download('words')
from nltk.corpus import words
# + id="5JXpd_SgjouD"
from translate import Translator
translator = Translator(to_lang='English')
# + id="tEGohMYUo8F3"
vec = set()
# + id="Giz13fp3qEEK"
hindi_vocab = {}
# + colab={"base_uri": "https://localhost:8080/"} id="-YJF3Ro3pLqY" outputId="26b40245-764b-4388-88da-7030c651ade5"
en_words = words.words()
en_words = set(en_words)
len(en_words)
# + id="mBF8nyLSjPVb"
def trans(line):
arr = line.split(' ')
new_arr = []
for word in arr:
if word in en_words:
new_arr.append(word)
else:
vec.add(word)
# if word in hindi_vocab:
# new_arr.append(hindi_vocab[word])
# else:
# tr = translator.translate(word)
# hindi_vocab[word] = tr
# new_arr.append(tr)
return ' '.join(new_arr)
# + colab={"base_uri": "https://localhost:8080/", "height": 706, "referenced_widgets": ["0744076c47d94375abe5717db1684f0e", "eca7b50ffa5d4e54997f8563e29afacd", "6f9771c98c9c46ac85917d0bebc07862", "<KEY>", "<KEY>", "7108752e05d64e208657ee80c65dd73a", "0bd14d36d5bc44e3898b5feaf9ce08de", "<KEY>", "57fe1bc62e154d4a828ba56742079e54", "<KEY>", "<KEY>"]} id="XTBxR74RjQfJ" outputId="969e9cdd-0be2-4a37-e145-c6aa3b95023e"
new_text = []
for index, row in tqdm(df.iterrows()):
try:
new_text.append(trans(row.text))
except:
new_text.append('')
if index%250 == 0:
print(index)
# df['text'] = df['text'].apply(lambda line: trans(line))
# + colab={"base_uri": "https://localhost:8080/", "height": 501, "referenced_widgets": ["04662734f41748dd8d7a8f6bc3519dc2", "3e0d23bfed41494db5512e99b96dd487", "<KEY>", "fa29afe0db87496d897a87d6143c3042", "<KEY>", "e7fedd66f3ca4ebc89864aca2b7bdaea", "200311f0a5504e3a8ea15d05b968ba0f", "3c68ba28b8654b1ebbeb0a232a69fa5a", "16d184328a8747abb2e3104376b3ad67", "<KEY>", "1ff7a4c2169941ee91b9e4ae880d2123"]} id="wRn7rGZFpnir" outputId="f26cb088-9b46-47ac-84ff-2feced779aa1"
vec_list = list(vec)
for i,word in enumerate(tqdm(vec_list)):
tr = translator.translate(word)
hindi_vocab[word] = tr
# + papermill={"duration": 0.046982, "end_time": "2020-09-25T19:48:21.462996", "exception": false, "start_time": "2020-09-25T19:48:21.416014", "status": "completed"} tags=[] colab={"base_uri": "https://localhost:8080/", "height": 238} id="7bQgHecxZje-" outputId="be014734-01cb-494a-d02a-484608d80320"
df.head()
# + papermill={"duration": 0.041243, "end_time": "2020-09-25T19:48:21.535564", "exception": false, "start_time": "2020-09-25T19:48:21.494321", "status": "completed"} tags=[] colab={"base_uri": "https://localhost:8080/"} id="_b_AXYFFZje_" outputId="c8b79eac-5d95-485b-c277-ebc0003c84b5"
df.category.value_counts()
# + papermill={"duration": 0.04214, "end_time": "2020-09-25T19:48:21.684168", "exception": false, "start_time": "2020-09-25T19:48:21.642028", "status": "completed"} tags=[] colab={"base_uri": "https://localhost:8080/"} id="KgKKJghiZjfB" outputId="559a665a-9203-4f67-93ac-44e052bed983"
df.category.value_counts()
# + papermill={"duration": 0.039202, "end_time": "2020-09-25T19:48:21.755805", "exception": false, "start_time": "2020-09-25T19:48:21.716603", "status": "completed"} tags=[] id="ZARDZX2oZjfB"
possible_labels = df.category.unique()
# + papermill={"duration": 0.039653, "end_time": "2020-09-25T19:48:21.827872", "exception": false, "start_time": "2020-09-25T19:48:21.788219", "status": "completed"} tags=[] id="2n7gNOPJZjfB"
label_dict = {}
for index, possible_label in enumerate(possible_labels):
label_dict[possible_label] = index
# + papermill={"duration": 0.040137, "end_time": "2020-09-25T19:48:21.900939", "exception": false, "start_time": "2020-09-25T19:48:21.860802", "status": "completed"} tags=[] colab={"base_uri": "https://localhost:8080/"} id="qqsYs4pNZjfC" outputId="2a038a2d-9298-49c2-8f2b-8b561cd298fe"
label_dict
# + papermill={"duration": 0.044378, "end_time": "2020-09-25T19:48:21.979606", "exception": false, "start_time": "2020-09-25T19:48:21.935228", "status": "completed"} tags=[] id="i5QNZ3DEZjfC"
df.category = df['category'].map(label_dict)
# + papermill={"duration": 0.057347, "end_time": "2020-09-25T19:48:22.091890", "exception": false, "start_time": "2020-09-25T19:48:22.034543", "status": "completed"} tags=[] colab={"base_uri": "https://localhost:8080/", "height": 394} id="LpA_HUTxZjfD" outputId="5478b063-c55f-4f55-f45b-64867fe92a39"
df.head(10)
# + [markdown] papermill={"duration": 0.038215, "end_time": "2020-09-25T19:48:22.167600", "exception": false, "start_time": "2020-09-25T19:48:22.129385", "status": "completed"} tags=[] id="JqV2GTcpZjfD"
# Classes are imbalanced as visible
# + [markdown] papermill={"duration": 0.038175, "end_time": "2020-09-25T19:48:22.242791", "exception": false, "start_time": "2020-09-25T19:48:22.204616", "status": "completed"} tags=[] id="zbzBU3J5ZjfD"
# ## 2: Training/Validation Split
# + papermill={"duration": 1.053035, "end_time": "2020-09-25T19:48:23.332257", "exception": false, "start_time": "2020-09-25T19:48:22.279222", "status": "completed"} tags=[] id="D9cr5BIHZjfD"
from sklearn.model_selection import train_test_split
# + papermill={"duration": 0.072998, "end_time": "2020-09-25T19:48:23.459663", "exception": false, "start_time": "2020-09-25T19:48:23.386665", "status": "completed"} tags=[] id="ehpaRMK7ZjfD"
X_train, X_val, y_train, y_val = train_test_split(df.index.values,
df.category.values,
test_size=0.15,
random_state=42,
stratify=df.category.values)
# + papermill={"duration": 0.074634, "end_time": "2020-09-25T19:48:23.591300", "exception": false, "start_time": "2020-09-25T19:48:23.516666", "status": "completed"} tags=[] id="bCl2CePBZjfE"
df['data_type'] = ['not_set']*df.shape[0]
# + papermill={"duration": 0.064226, "end_time": "2020-09-25T19:48:23.712026", "exception": false, "start_time": "2020-09-25T19:48:23.647800", "status": "completed"} tags=[] colab={"base_uri": "https://localhost:8080/", "height": 238} id="kDFV1ZNvZjfE" outputId="4c4423f5-bc60-43ae-e5ec-c95d4789fbca"
df.head()
# + papermill={"duration": 0.063313, "end_time": "2020-09-25T19:48:23.821906", "exception": false, "start_time": "2020-09-25T19:48:23.758593", "status": "completed"} tags=[] id="W8hQRO_7ZjfE"
df.loc[X_train, 'data_type'] = 'train'
df.loc[X_val, 'data_type'] = 'val'
# + papermill={"duration": 0.064645, "end_time": "2020-09-25T19:48:23.923094", "exception": false, "start_time": "2020-09-25T19:48:23.858449", "status": "completed"} tags=[] colab={"base_uri": "https://localhost:8080/", "height": 206} id="WkOvtTxQZjfE" outputId="081542d8-4738-4b73-c9c0-2f06d311f126"
df.groupby(['category', 'data_type']).count()
# + [markdown] papermill={"duration": 0.049916, "end_time": "2020-09-25T19:48:24.020356", "exception": false, "start_time": "2020-09-25T19:48:23.970440", "status": "completed"} tags=[] id="sw9KYLScZjfF"
# # 3. Loading Tokenizer and Encoding our Data
# + colab={"base_uri": "https://localhost:8080/"} id="rzlbQxuwacL0" outputId="49402ba7-2543-448b-aa71-1170c99c5889"
# !pip install transformers
# + papermill={"duration": 6.608677, "end_time": "2020-09-25T19:48:30.679359", "exception": false, "start_time": "2020-09-25T19:48:24.070682", "status": "completed"} tags=[] id="JD8kJWXzZjfF"
from transformers import BertTokenizer
from transformers import AutoTokenizer, AutoModelForMaskedLM
from torch.utils.data import TensorDataset
# + papermill={"duration": 0.359537, "end_time": "2020-09-25T19:48:31.075982", "exception": false, "start_time": "2020-09-25T19:48:30.716445", "status": "completed"} tags=[] colab={"base_uri": "https://localhost:8080/", "height": 113, "referenced_widgets": ["5f8a230a099a4eef8cd097bac1ddedab", "0941b2a1ab7949e89dfce7455bcf997f", "546cb277e58f49a2a49afb73174fc966", "e24aeffa097344f2b882189e18f1ba9a", "163ad4170f5e4b4ea1ec51c93c826b77", "<KEY>", "<KEY>", "d968577ad7f44de393f3583693a12e5e", "<KEY>", "320034db051a40479644c2589eb532d0", "<KEY>", "1bcef91f7e224c0380fac00b6e164abc", "deac07c908d04d57b41b04eb50556766", "<KEY>", "5ebb9181d49044f88942e8ad196f6404", "d248e537a58b44e09c9486c663e37996", "<KEY>", "<KEY>", "<KEY>", "900ed4754e474325a49f992e5bd8ae2d", "<KEY>", "54e34a9ec02743919ee19bcaf6fb934d", "ad3bea9d87b94594b47bf32caf64b3a7", "aaed12551cc34765ba47fabe9ace8b9f", "2f7019091be4485fa404212404520eac", "599c780a19b447c8ace33f4ce50171aa", "<KEY>", "ce704783a44c4e9abf3685ed989d7374", "0f137359fe984fb7b10e4cb60da88984", "<KEY>", "1317ac4ca0b64a099fc0aa2598632401", "8f9ffaf3d45f45179fc5d2560ef5b5ec", "8d7b22a3fb374b19aca046f123fc02e1"]} id="aH3mf07xZjfF" outputId="2f855158-50af-4ed7-98fa-df6215e8098f"
tokenizer = AutoTokenizer.from_pretrained('xlm-roberta-base')
# + colab={"base_uri": "https://localhost:8080/", "height": 455} id="tQrL4kqya_BA" outputId="89373592-fbbf-4b9e-cfa6-2b9f8c621aa6"
df.dropna()
# + colab={"base_uri": "https://localhost:8080/"} id="eDyoFe0CbOwH" outputId="0702c094-6bcc-4ab2-bb02-d448c480758b"
df['text'] = df['text'].astype('str')
df[df.data_type=='train'].text.values
# + id="zDHlRFaYbd3F"
all_text_train = df[df.data_type=='train'].text.tolist()
# + papermill={"duration": 1.266904, "end_time": "2020-09-25T19:48:32.381780", "exception": false, "start_time": "2020-09-25T19:48:31.114876", "status": "completed"} tags=[] colab={"base_uri": "https://localhost:8080/"} id="RvCxZqElZjfF" outputId="29459241-40a0-463a-8cbb-8315b40d9219"
encoded_data_train = tokenizer.batch_encode_plus(
all_text_train,
add_special_tokens=True,
return_attention_mask=True,
pad_to_max_length=True,
max_length=256,
return_tensors='pt'
)
encoded_data_val = tokenizer.batch_encode_plus(
df[df.data_type=='val'].text.tolist(),
add_special_tokens=True,
return_attention_mask=True,
pad_to_max_length=True,
max_length=256,
return_tensors='pt'
)
input_ids_train = encoded_data_train['input_ids']
attention_masks_train = encoded_data_train['attention_mask']
labels_train = torch.tensor(df[df.data_type=='train'].category.values)
input_ids_val = encoded_data_val['input_ids']
attention_masks_val = encoded_data_val['attention_mask']
labels_val = torch.tensor(df[df.data_type=='val'].category.values)
# + colab={"base_uri": "https://localhost:8080/"} id="SNOtdwutbeTT" outputId="f0d419f0-bae9-40c8-9059-ac096f3ee6f7"
len(input_ids_train), len(attention_masks_train), len(labels_train)
# + papermill={"duration": 0.048886, "end_time": "2020-09-25T19:48:32.470406", "exception": false, "start_time": "2020-09-25T19:48:32.421520", "status": "completed"} tags=[] id="1b-ihvLJZjfG"
dataset_train = TensorDataset(input_ids_train,
attention_masks_train,
labels_train)
dataset_val = TensorDataset(input_ids_val,
attention_masks_val,
labels_val)
# + papermill={"duration": 0.046463, "end_time": "2020-09-25T19:48:32.554501", "exception": false, "start_time": "2020-09-25T19:48:32.508038", "status": "completed"} tags=[] colab={"base_uri": "https://localhost:8080/"} id="Ka76yNGBZjfG" outputId="83676048-1e87-4201-f613-d59d5d81165f"
len(dataset_train)
# + papermill={"duration": 0.067851, "end_time": "2020-09-25T19:48:32.660735", "exception": false, "start_time": "2020-09-25T19:48:32.592884", "status": "completed"} tags=[] colab={"base_uri": "https://localhost:8080/"} id="p_VGPdvlZjfG" outputId="b01d6b8b-9327-4796-e5bb-ee4528c10cc5"
dataset_val.tensors
# + [markdown] papermill={"duration": 0.038163, "end_time": "2020-09-25T19:48:32.739040", "exception": false, "start_time": "2020-09-25T19:48:32.700877", "status": "completed"} tags=[] id="S2PPOrl0ZjfG"
# # 4. Setting up BERT Pretrained Model
# + papermill={"duration": 0.045229, "end_time": "2020-09-25T19:48:32.823259", "exception": false, "start_time": "2020-09-25T19:48:32.778030", "status": "completed"} tags=[] id="ukyhArGpZjfG"
from transformers import AutoConfig, AutoModelForSequenceClassification
# + papermill={"duration": 35.879219, "end_time": "2020-09-25T19:49:08.742460", "exception": false, "start_time": "2020-09-25T19:48:32.863241", "status": "completed"} tags=[] colab={"base_uri": "https://localhost:8080/", "height": 156, "referenced_widgets": ["3cce6975fb6144d98e204b6c9a8afbb7", "c88bfe6a26ee4d6c8ba459a0a58c10aa", "820008a9708043d191b181d5098e4c0a", "2c16eb18e56b427787a5b5b4efccd6e7", "539361d9ae424a90b34f9a561874b798", "95950204b8d54881a095b702dea3b514", "ef1cbed97e10470c89fc549791e8fc7b", "344c8e0b5a5042e984e5fbc7a6fc6a3c", "56033219e26445539af37c59867515cf", "<KEY>", "17c5bdb5d17c48e0aa950af75f30606e"]} id="pXj2ldcDZjfH" outputId="0af42d03-ccb1-472c-a7f4-3c6bb9be9099"
model = AutoModelForSequenceClassification.from_pretrained(
'xlm-roberta-base',
num_labels = len(label_dict),
output_attentions = False,
output_hidden_states = False
)
# + [markdown] papermill={"duration": 0.041057, "end_time": "2020-09-25T19:49:08.825580", "exception": false, "start_time": "2020-09-25T19:49:08.784523", "status": "completed"} tags=[] id="YbVCrvo0ZjfH"
# # 5. Creating Data Loaders
# + papermill={"duration": 0.049908, "end_time": "2020-09-25T19:49:08.917274", "exception": false, "start_time": "2020-09-25T19:49:08.867366", "status": "completed"} tags=[] id="m_cZirsVZjfH"
from torch.utils.data import DataLoader, RandomSampler, SequentialSampler
# + papermill={"duration": 0.05037, "end_time": "2020-09-25T19:49:09.008401", "exception": false, "start_time": "2020-09-25T19:49:08.958031", "status": "completed"} tags=[] id="2ldIzMECZjfH"
batch_size = 4
dataloader_train = DataLoader(
dataset_train,
sampler=RandomSampler(dataset_train),
batch_size=batch_size
)
dataloader_val = DataLoader(
dataset_val,
sampler=RandomSampler(dataset_val),
batch_size=32
)
# + [markdown] papermill={"duration": 0.041482, "end_time": "2020-09-25T19:49:09.090888", "exception": false, "start_time": "2020-09-25T19:49:09.049406", "status": "completed"} tags=[] id="D87ySXdUZjfH"
# # 6. Setting Up Optimizer and Scheduler
# + papermill={"duration": 0.048269, "end_time": "2020-09-25T19:49:09.180451", "exception": false, "start_time": "2020-09-25T19:49:09.132182", "status": "completed"} tags=[] id="TwX6fkxiZjfH"
from transformers import AdamW, get_linear_schedule_with_warmup
# + papermill={"duration": 0.051332, "end_time": "2020-09-25T19:49:09.275138", "exception": false, "start_time": "2020-09-25T19:49:09.223806", "status": "completed"} tags=[] id="_-S-hXBvZjfI"
optimizer = AdamW(
model.parameters(),
lr = 1e-5,
eps = 1e-8
)
# + papermill={"duration": 0.04937, "end_time": "2020-09-25T19:49:09.366346", "exception": false, "start_time": "2020-09-25T19:49:09.316976", "status": "completed"} tags=[] id="7ufHbhbUZjfI"
epochs = 5
scheduler = get_linear_schedule_with_warmup(
optimizer,
num_warmup_steps=0,
num_training_steps = len(dataloader_train)*epochs
)
# + [markdown] papermill={"duration": 0.041344, "end_time": "2020-09-25T19:49:09.449933", "exception": false, "start_time": "2020-09-25T19:49:09.408589", "status": "completed"} tags=[] id="uRhVjKFWZjfI"
# # 7. Defining our Performance Metrics
# + papermill={"duration": 0.048777, "end_time": "2020-09-25T19:49:09.540839", "exception": false, "start_time": "2020-09-25T19:49:09.492062", "status": "completed"} tags=[] id="FzdDtGjVZjfI"
import numpy as np
from sklearn.metrics import f1_score
# + papermill={"duration": 0.050586, "end_time": "2020-09-25T19:49:09.634137", "exception": false, "start_time": "2020-09-25T19:49:09.583551", "status": "completed"} tags=[] id="ehPP23vfZjfI"
def f1_score_func(preds, labels):
preds_flat = np.argmax(preds, axis=1).flatten()
labels_flat = labels.flatten()
return f1_score(labels_flat, preds_flat, average = 'weighted')
# + papermill={"duration": 0.053645, "end_time": "2020-09-25T19:49:09.730645", "exception": false, "start_time": "2020-09-25T19:49:09.677000", "status": "completed"} tags=[] id="bh6xyhUCZjfI"
def accuracy_per_class(preds, labels):
label_dict_inverse = {v: k for k, v in label_dict.items()}
preds_flat = np.argmax(preds, axis=1).flatten()
labels_flat = labels.flatten()
for label in np.unique(labels_flat):
y_preds = preds_flat[labels_flat==label]
y_true = labels_flat[labels_flat==label]
print(f'Class: {label_dict_inverse[label]}')
print(f'Accuracy:{len(y_preds[y_preds==label])}/{len(y_true)}\n')
# + [markdown] papermill={"duration": 0.042252, "end_time": "2020-09-25T19:49:09.816391", "exception": false, "start_time": "2020-09-25T19:49:09.774139", "status": "completed"} tags=[] id="xApqMoJmZjfI"
# # 8. Creating our Training Loop
# + papermill={"duration": 0.060651, "end_time": "2020-09-25T19:49:09.919256", "exception": false, "start_time": "2020-09-25T19:49:09.858605", "status": "completed"} tags=[] id="Wj-k5_uDZjfJ"
import random
seed_val = 42
random.seed(seed_val)
np.random.seed(seed_val)
torch.manual_seed(seed_val)
torch.cuda.manual_seed_all(seed_val)
# + papermill={"duration": 7.770862, "end_time": "2020-09-25T19:49:17.733331", "exception": false, "start_time": "2020-09-25T19:49:09.962469", "status": "completed"} tags=[] colab={"base_uri": "https://localhost:8080/"} id="BzLmjw4aZjfJ" outputId="09572554-48ba-4f49-e87a-cfab663c20a5"
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
model.to(device)
print(device)
# + papermill={"duration": 0.055972, "end_time": "2020-09-25T19:49:17.832806", "exception": false, "start_time": "2020-09-25T19:49:17.776834", "status": "completed"} tags=[] id="bGMqbM8QZjfJ"
def evaluate(dataloader_val):
model.eval()
loss_val_total = 0
predictions, true_vals = [], []
for batch in tqdm(dataloader_val):
batch = tuple(b.to(device) for b in batch)
inputs = {'input_ids': batch[0],
'attention_mask': batch[1],
'labels': batch[2],
}
with torch.no_grad():
outputs = model(**inputs)
loss = outputs[0]
logits = outputs[1]
loss_val_total += loss.item()
logits = logits.detach().cpu().numpy()
label_ids = inputs['labels'].cpu().numpy()
predictions.append(logits)
true_vals.append(label_ids)
loss_val_avg = loss_val_total/len(dataloader_val)
predictions = np.concatenate(predictions, axis=0)
true_vals = np.concatenate(true_vals, axis=0)
return loss_val_avg, predictions, true_vals
# + papermill={"duration": 471.890013, "end_time": "2020-09-25T19:57:09.765836", "exception": false, "start_time": "2020-09-25T19:49:17.875823", "status": "completed"} tags=[] colab={"base_uri": "https://localhost:8080/", "height": 675, "referenced_widgets": ["1a346b5198fd4946a590a628a4e62f50", "<KEY>", "a7f044345c4646098c624724ab47672a", "3e571c1a95694ade9e4d50aeac8cf949", "<KEY>", "46eade9ea0dc45f9a827829aa013871d", "957a7f722dd342c8982e878ae5c59816", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "b431f376e1594367b0217e6f13c45e2b", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "603c258cfab14c4e9b8cc7e8f4aa2cef", "<KEY>", "fde0cf0181054d21b59483c246e628a6", "6d398ad115b145378a470371da57ebe9", "1867d4d3f5c34cbe9985fa7f7d387f45", "14cb49de11da44008d2aca3d7b4527fe", "e1e3d733ad4f460e9fffae0b0cd62228", "5dbf892ab1e841dab02c19069c02681b", "<KEY>", "<KEY>", "6ca0da51cc484456a8958603060a42da", "<KEY>", "44952405d09f4e85b14b4d9d63b6a634", "<KEY>", "75db72d8ee3845f2920500bbb61afa81", "5acd90298a2144059f03fb68b4173edb", "439c41a1027f43799859e8f606bec8e6", "<KEY>", "4a392c985e4d4633b59eadc82753cdeb", "<KEY>", "b25734e817fc4b629bd6f9b6e3408c2a", "e00f540c40844a35bbeb987c6f562cea", "<KEY>", "adebaeb8d9214e858159ec523c23d571", "<KEY>", "100e9961a52748d987d19e315df518ce", "<KEY>", "0be6d1aee1b5403f89a67b1f6b3675ef", "bbee4243a7e145ffb8c8e15969177da2", "<KEY>", "d392ac982e604ec9a429e5e0995663ec", "<KEY>", "<KEY>", "e1febc26c8154da184759ba8c31be6e3", "4f23a38ea686417b902559d147fe359c", "<KEY>", "<KEY>", "<KEY>", "58e0218300504a8d8b30f8517a13f879", "97beee9bb98e4876b056e4fd1d85eab7", "33c6c9b4366a4d51ac6ac60d1ed9f5dd", "<KEY>", "ec3d2729f2a04d57a70e8fe01579e40b", "<KEY>", "56196c1e88724d4e94914066dba813ba", "f34a7acc6a8c478c9be0cff66c570a8e", "adbaa0f43a454c27aefe37ed009cbaea", "<KEY>", "53e1258a54a848e99dd5e58d4361ab79", "<KEY>", "783966621c5543e3a8c97697365a66b0", "<KEY>", "<KEY>", "8c7342bec7ae473a946914f63ee07872", "9b8af1a671704398906f7ed241641587", "<KEY>", "e17f2f72a6de42b880f345757d3798e3", "db4f00fde62d420e8b7949eb07375217", "<KEY>", "0589ddaf50d54e5ca95c21527bd75f17", "<KEY>", "b58ac6e1ca3848cb9be9880d1b6ba42c", "96ad5bf7dd9d4b17a3212d110223e217", "0f3ddd8696e74f159858e7341050f141", "fe36a87a4ee74e2da289ffae6e8ee2b5", "<KEY>", "<KEY>", "<KEY>", "25027c027edc415b9b4ec4b625e66a3d", "4e39ff7a8c5244e096bedda59e56919e", "<KEY>", "<KEY>", "24de20ae22854f24a81df17af235281a", "<KEY>", "0fdbce334c5141508ae9dd50ec4692dd", "<KEY>", "<KEY>", "<KEY>", "b847dab73dfa466db3a66e9bbe242259", "d61593e4730841b4a9510c763dd1fafa", "<KEY>", "<KEY>", "f09d762467da4dca89b9dd88a544ca94", "95e7ca831ba0472ca415a02d54123d93", "<KEY>", "daa2cdf072874f4db8c0ce95c6c65505", "e4b4d25b218a4c40841e57a79e26ad66", "c0d2e24278f349e69537f32afed740f0", "<KEY>", "71b0ed9ea9c041efac66e5e60c160fbd", "<KEY>", "654b323c3e9a4b858a411e3a958a8d98", "effb986bfed1439cb98c60658eecadfb", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "4e0ff63b6dad4dfab21c872ef13bb553", "b01977360d144ad6bd7563cec1c79bef", "<KEY>", "bc6aaf33ef7643a2b917b8eac2f468a6", "101ce14917e343b3a0175a5641976594", "16bfaf053ff84118a7e2921e194f8032", "821e62397c5a450988071709fc5953d4", "3f3284c3010942aebe0fb1005ff99e81"]} id="M6Krjr6BZjfJ" outputId="9427b3eb-003c-40d3-cb57-004a9cd1bec9"
for epoch in tqdm(range(1, epochs+1)):
model.train()
loss_train_total = 0
progress_bar = tqdm(dataloader_train,
desc='Epoch {:1d}'.format(epoch),
leave=False,
disable=False)
for batch in progress_bar:
model.zero_grad()
batch = tuple(b.to(device) for b in batch)
inputs = {
'input_ids': batch[0],
'attention_mask': batch[1],
'labels': batch[2]
}
outputs = model(**inputs)
loss = outputs[0]
loss_train_total +=loss.item()
loss.backward()
torch.nn.utils.clip_grad_norm_(model.parameters(), 1.0)
optimizer.step()
scheduler.step()
progress_bar.set_postfix({'training_loss': '{:.3f}'.format(loss.item()/len(batch))})
#torch.save(model.state_dict(), f'Models/BERT_ft_Epoch{epoch}.model')
tqdm.write('\nEpoch {epoch}')
loss_train_avg = loss_train_total/len(dataloader_train)
tqdm.write(f'Training loss: {loss_train_avg}')
val_loss, predictions, true_vals = evaluate(dataloader_val)
val_f1 = f1_score_func(predictions, true_vals)
tqdm.write(f'Validation loss: {val_loss}')
tqdm.write(f'F1 Score (weighted): {val_f1}')
# + [markdown] papermill={"duration": 0.061737, "end_time": "2020-09-25T19:57:09.890836", "exception": false, "start_time": "2020-09-25T19:57:09.829099", "status": "completed"} tags=[] id="EQZjN_gcZjfJ"
# # 9. Evaluating our Model
# + papermill={"duration": 0.075568, "end_time": "2020-09-25T19:57:10.028406", "exception": false, "start_time": "2020-09-25T19:57:09.952838", "status": "completed"} tags=[] id="rCS9Iy6EZjfJ" colab={"base_uri": "https://localhost:8080/"} outputId="8e0efa24-a1f5-409c-87c8-75429b067334"
accuracy_per_class(predictions, true_vals)
# + id="7ovJ67f4eU8U"
|
BERT_with_MT.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lecture 60: Activity recognition using CNN-LSTM
# ## 60a: Train CNN for activity classification
#
# #### Note:
# 1. Run lecture59_preProc1.ipynb before running executing this notebook
# 2. Files lecture60a.ipynb, lecture60b.ipynb, lecture60c.ipynb are part of the same tutorial and are to be exeuted sequentially
# #### Dataset: [UCF101](https://www.crcv.ucf.edu/research/data-sets/ucf101/)
# +
# %matplotlib inline
import copy
import time
import torch
import numpy as np
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
import matplotlib.pyplot as plt
from torchvision import transforms,datasets, models
print(torch.__version__) # This code has been updated for PyTorch 1.0.0
# +
# Check availability of GPU
use_gpu = torch.cuda.is_available()
# use_gpu = False # Uncomment in case of GPU memory error
if use_gpu:
print('GPU is available!')
device = "cuda"
pinMem = True
else:
print('GPU is not available!')
device = "cpu"
pinMem = False
# +
# Loading data from folder using ImageFolder
trainDir = 'train_5class'
valDir = 'test_5class'
apply_transform = transforms.Compose([
transforms.Resize(256),
transforms.CenterCrop(224),
transforms.ToTensor()])
BatchSize = 128
# Training dataloader
train_dataset = datasets.ImageFolder(trainDir,transform=apply_transform)
trainLoader = torch.utils.data.DataLoader(train_dataset, batch_size=BatchSize, shuffle=True,num_workers=4, pin_memory=pinMem)
# Test dataloader
test_dataset = datasets.ImageFolder(valDir,transform=apply_transform)
testLoader = torch.utils.data.DataLoader(test_dataset, batch_size=BatchSize, shuffle=False,num_workers=4, pin_memory=pinMem)
# -
# Size of train and test datasets
print('No. of samples in train set: '+str(len(trainLoader.dataset)))
print('No. of samples in test set: '+str(len(testLoader.dataset)))
# ## Define network architecture
net = models.resnet18(pretrained=True)
print(net)
# Counting number of trainable parameters
totalParams = 0
for name,params in net.named_parameters():
print(name,'-->',params.size())
totalParams += np.sum(np.prod(params.size()))
print('Total number of parameters: '+str(totalParams))
# Modifying the last fully-connected layer for 5 classes
net.fc = nn.Linear(512,5)
net = net.to(device)
# ## Define loss function and optimizer
criterion = nn.NLLLoss() # Negative Log-likelihood
optimizer = optim.Adam(net.fc.parameters(), lr=1e-4) # Adam
# ## Train the network
# +
iterations = 10
trainLoss = []
trainAcc = []
testLoss = []
testAcc = []
start = time.time()
for epoch in range(iterations):
epochStart = time.time()
runningLoss = 0.0
avgTotalLoss = 0.0
running_correct = 0
net.train() # For training
batchNum = 1
for data in trainLoader:
inputs,labels = data
inputs, labels = inputs.to(device), labels.to(device)
outputs = net(inputs)
_, predicted = torch.max(outputs.data, 1)
running_correct += (predicted == labels.data).sum()
# Initialize gradients to zero
optimizer.zero_grad()
# Compute loss/error
loss = criterion(F.log_softmax(outputs,dim=1), labels)
# Backpropagate loss and compute gradients
loss.backward()
# Update the network parameters
optimizer.step()
# Accumulate loss per batch
runningLoss += loss.item()
batchNum += 1
avgTrainAcc = 100*float(running_correct)/float(len(trainLoader.dataset))
avgTrainLoss = runningLoss/(float(len(trainLoader.dataset))/BatchSize)
trainAcc.append(avgTrainAcc)
trainLoss.append(avgTrainLoss)
# Evaluating performance on test set for each epoch
net.eval() # For testing [Affects batch-norm and dropout layers (if any)]
running_correct = 0
with torch.no_grad():
for data in testLoader:
inputs,labels = data
inputs, labels = inputs.to(device), labels.to(device)
outputs = net(inputs)
_, predicted = torch.max(outputs.data, 1)
running_correct += (predicted == labels.data).sum()
loss = criterion(F.log_softmax(outputs,dim=1), labels)
runningLoss += loss.item()
avgTestLoss = runningLoss/(float(len(testLoader.dataset))/BatchSize)
avgTestAcc = 100*float(running_correct)/float(len(testLoader.dataset))
testAcc.append(avgTestAcc)
testLoss.append(avgTestLoss)
# Plotting training loss vs Epochs
fig1 = plt.figure(1)
plt.plot(range(epoch+1),trainLoss,'r-',label='train')
plt.plot(range(epoch+1),testLoss,'g-',label='test')
if epoch==0:
plt.legend(loc='upper left')
plt.xlabel('Epochs')
plt.ylabel('Loss')
# Plotting testing accuracy vs Epochs
fig2 = plt.figure(2)
plt.plot(range(epoch+1),trainAcc,'r-',label='train')
plt.plot(range(epoch+1),testAcc,'g-',label='test')
if epoch==0:
plt.legend(loc='upper left')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
epochEnd = time.time()-epochStart
print('Iteration: {:.0f} /{:.0f}; Training Loss: {:.6f} ; Training Acc: {:.3f}'\
.format(epoch + 1,iterations,avgTrainLoss,avgTrainAcc))
print('Iteration: {:.0f} /{:.0f}; Testing Loss: {:.6f} ; Testing Acc: {:.3f}'\
.format(epoch + 1,iterations,avgTestLoss,avgTestAcc))
print('Time consumed: {:.0f}m {:.0f}s'.format(epochEnd//60,epochEnd%60))
end = time.time()-start
print('Training completed in {:.0f}m {:.0f}s'.format(end//60,end%60))
# -
# ### Save trained model
torch.save(net.state_dict(), 'resnet18Pre_fcOnly5class_ucf101_10adam_1e-4_b128.pt')
|
lecture60a.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Zachary's Karate Club Community Detection Using the `NETWORK` Actionset in SAS Viya and Python
# In this example, we load the Zachary's Karate Club graph into CAS, and show how to detect communities using the network actionset.
#
# his example uses Zachary’s Karate Club data (<a href="https://go.documentation.sas.com/?docsetId=casmlnetwork&docsetTarget=casmlnetwork_network_references.htm&docsetVersion=8.5&locale=en&showBanner=walkup#casmlnetwork_networkzach_w77">Zachary 1977</a>), which describes social network friendships between 34 members of a karate club at a US university in the 1970s. This is one of the standard publicly available data tables for testing community detection algorithms. It contains 34 nodes and 78 links. The graph is shown below.
#
# ----------------
#
# The basic flow of this notebook is as follows:
# 1. Load the sample graph into a Pandas DataFrame as a set of links that represent the total graph.
# 2. Connect to our CAS server and load the actionsets we require.
# 3. Upload our sample graph to our CAS server.
# 4. Execute the community detection without fixed nodes using two resolutions (0.5 and 1.0).
# 5. Prepare and display the network plots showing the cliques.
#
# ----------------
# __Prepared by:__
# <NAME> (<i class="fa fa-github" aria-hidden="true"></i>: [dtherrick](www.github.com/dtherrick))
# ## Imports
#
# Our imports are broken out as follows:
#
# | Module | Method | Description |
# |:-----------------|:-----------------:|:----------------------------------------------------------------------------------:|
# | `os` | all | Allows access to environment variables. |
# | `sys` | all | Used to update our system path so Python can import our custom utility functions. |
# | `swat` | all | SAS Python module that orchestrates communicatoin with a CAS server. |
# | `pandas` | all | Data management module we use for preparation of local data. |
# | `networkx` | all | Used to manage graph data structures when plotting. |
# | `bokeh.io` | `output_notebook` | Utility function that allows rendering of Bokeh plots in Jupyter |
# | `bokeh.io` | `show` | Utility function that displays Bokeh plots |
# | `bokeh.layouts` | `gridplot` | Utility function that arranges Bokeh plots in a multi-plot grid |
# | `bokeh.palettes` | `Spectral8` | Eight-color palette used to differentiate node types. |
# | `bokehvis` | all | Custom module written to simplify plot rendering with Bokeh |
# +
import os
import sys
import swat
import pandas as pd
import networkx as nx
from bokeh.io import output_notebook, show
from bokeh.layouts import gridplot
from bokeh.palettes import Spectral8
sys.path.append(os.path.join(os.path.dirname(os.getcwd()),r"../../common/python"))
import bokehvis as vis
# tell our notebook we want to output with Bokeh
output_notebook()
# -
# ## Prepare the sample graph.
# * We pass a set of links, and a set of nodes. Nodes are passed this time because we define fix groups for later calculation on load.
colNames = ["from", "to"]
links = [
(0, 9), (0, 10), (0, 14), (0, 15), (0, 16), (0, 19), (0, 20), (0, 21), (0, 33),
(0, 23), (0, 24), (0, 27), (0, 28), (0, 29), (0, 30), (0, 31), (0, 32),
(2, 1),
(3, 1), (3, 2),
(4, 1), (4, 2), (4, 3),
(5, 1),
(6, 1),
(7, 1), (7, 5), (7, 6),
(8, 1), (8, 2), (8, 3), (8, 4),
(9, 1), (9, 3),
(10, 3),
(11, 1), (11, 5), (11, 6),
(12, 1),
(13, 1), (13, 4),
(14, 1), (14, 2), (14, 3), (14, 4),
(17, 6), (17, 7),
(18, 1), (18, 2),
(20, 1), (20, 2),
(22, 1), (22, 2),
(26, 24), (26, 25),
(28, 3), (28, 24), (28, 25),
(29, 3),
(30, 24), (30, 27),
(31, 2), (31, 9),
(32, 1), (32, 25), (32, 26), (32, 29),
(33, 3), (33, 9), (33, 15), (33, 16), (33, 19), (33, 21), (33, 23), (33, 24), (33, 30), (33, 31), (33, 32),
]
dfLinkSetIn = pd.DataFrame(links, columns=colNames)
# Let's start by looking at the basic network itself.
#
# We create a `networkx` graph and pass it to our `bokeh` helper function to create the initial plot.
# +
G_comm = nx.from_pandas_edgelist(dfLinkSetIn, 'from', 'to')
title = "Zachary's Karate Club"
hover = [('Node', '@index')]
nodeSize = 25
plot = vis.render_plot(graph=G_comm,
title=title,
hover_tooltips=hover,
node_size=nodeSize,
width=1200,
label_font_size="10px",
label_x_offset=-3)
show(plot)
# -
# ## Connect to CAS, load the actionsets we'll need, and upload our graph to the CAS server.
# +
host = os.environ['CAS_HOST_ORGRD']
port = int(os.environ['CAS_PORT'])
conn = swat.CAS(host, port)
conn.loadactionset("network")
# -
# ### Upload the local dataframe into CAS
conn.setsessopt(messageLevel="ERROR")
_ = conn.upload(dfLinkSetIn, casout='LinkSetIn')
conn.setsessopt(messageLevel="DEFAULT")
# ### Step 3: Calculate the communities (without fixed groups) in our graph using the `network` actionset.
# Since we've loaded our actionset, we can reference it using dot notation from our connection object.
#
# We use detection at two resolutions: 0.5 and 1.0
#
# Note that the Python code below is equivalent to this block of CASL:
# ```
# proc network
# links = mycas.LinkSetIn
# outNodes = mycas.NodeSetOut;
# community
# resolutionList = 1.0 0.5
# outLevel = mycas.CommLevelOut
# outCommunity = mycas.CommOut
# outOverlap = mycas.CommOverlapOut
# outCommLinks = mycas.CommLinksOut;
# run;
# ```
conn.network.community(links = {'name':'LinkSetIn'},
outnodes = {'name':'nodeSetOut', 'replace':True},
outLevel = {'name':'CommLevelOut', 'replace':True},
outCommunity = {'name':'CommOut', 'replace':True},
outOverlap = {'name':'CommOverlapOut', 'replace':True},
outCommLinks = {'name':'CommLinksOut', 'replace':True},
resolutionList = [0.5, 1]
)
# ### Step 4: Get the community results from CAS and prepare data for plotting
#
# ------
# In this step we fetch the node results from CAS, then add community assignments and node fill color as node attributes in our `networkx` graph.
#
# | Table | Description |
# |------------|-----------------------------------------------------------|
# | `NodeSetA` | Results and community labels for resolutions 0.5 and 1.0. |
#
# | Attribute Label | Description |
# |-------------------|--------------------------------------|
# | `community_0` | Community assignment, resolution 1.0 |
# | `community_1` | Community assignment, resolution 0.5 |
# +
# pull the node set locally so we can plot
comm_nodes_cas = conn.CASTable('NodeSetOut').to_dict(orient='index')
# make our mapping dictionaries that allow us to assign attributes
comm_nodes_0 = {v['node']:v['community_0'] for v in comm_nodes_cas.values()}
comm_nodes_1 = {v['node']:v['community_1'] for v in comm_nodes_cas.values()}
# set the attributes
nx.set_node_attributes(G_comm, comm_nodes_0, 'community_0')
nx.set_node_attributes(G_comm, comm_nodes_1, 'community_1')
# Assign the fill colors for the nodes.
for node in G_comm.nodes:
G_comm.nodes[node]['highlight_0'] = Spectral8[int(G_comm.nodes[node]['community_0'])]
G_comm.nodes[node]['highlight_1'] = Spectral8[int(G_comm.nodes[node]['community_1'])]
# -
# ### Create and display the plots
# +
title_0 = 'Community Detection Example 1: Resolution 1'
hover_0 = [('Node', '@index'), ('Community', '@community_0')]
title_1 = 'Community Detection Example 2: Resolution 0.5'
hover_1 = [('Node', '@index'), ('Community', '@community_1')]
# render the plots.
# reminder - we set nodeSize earlier in the notebook. Its value is 25.
plot_0 = vis.render_plot(graph=G_comm, title=title_0, hover_tooltips=hover_0, node_size=nodeSize, node_color='highlight_0', width=1200)
plot_1 = vis.render_plot(graph=G_comm, title=title_1, hover_tooltips=hover_1, node_size=nodeSize, node_color='highlight_1', width=1200)
# -
grid = gridplot([plot_0, plot_1], ncols=1)
show(grid)
# ## Clean up everything.
#
# Make sure we know what tables we created, drop them, and close our connection.
# (This is probably overkill, since everything in this session is ephemeral anyway, but good practice nonetheless.
# +
table_list = conn.tableinfo()["TableInfo"]["Name"].to_list()
for table in table_list:
conn.droptable(name=table, quiet=True)
conn.close()
|
examples/community/python/example_karateclub.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### validation or kfold is not added and also data augmentation is not added yet. Mainly the Ensemble is not performed. Due for tomorrow.
# %%capture
# !pip install ../input/segmentation-models-wheels/efficientnet_pytorch-0.6.3-py3-none-any.whl
# !pip install ../input/segmentation-models-wheels/pretrainedmodels-0.7.4-py3-none-any.whl
# !pip install ../input/segmentation-models-wheels/timm-0.3.2-py3-none-any.whl
# !pip install ../input/segmentation-models-wheels/segmentation_models_pytorch-0.1.3-py3-none-any.whl
# !pip install colorama -q
# +
# import wandb
# from wandb.keras import WandbCallback
# try:
# from kaggle_secrets import UserSecretsClient
# user_secrets = UserSecretsClient()
# api_key = user_secrets.get_secret("WANDB")
# wandb.login(key=api_key)
# anonymous = None
# except:
# anonymous = "must"
# print('To use your W&B account,\nGo to Add-ons -> Secrets and provide your W&B access token. Use the Label name as WANDB. \nGet your W&B access token from here: https://wandb.ai/authorize')
# +
# wandb.init(project="kaggle_sartorius_unet_ensemble", entity="somusan")
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
import os
import cv2
import pdb
import time
import warnings
import random
import numpy as np
import pandas as pd
from tqdm import tqdm_notebook as tqdm
from torch.optim.lr_scheduler import ReduceLROnPlateau
from sklearn.model_selection import KFold
import torch
import torch.nn as nn
from torch.nn import functional as F
import torch.optim as optim
import torch.backends.cudnn as cudnn
from torch.utils.data import DataLoader, Dataset, sampler
from matplotlib import pyplot as plt
import albumentations as A
from albumentations.pytorch import ToTensorV2
from PIL import Image
from sklearn.metrics import accuracy_score, f1_score
warnings.filterwarnings("ignore")
def fix_all_seeds(seed):
np.random.seed(seed)
random.seed(seed)
os.environ['PYTHONHASHSEED'] = str(seed)
torch.manual_seed(seed)
torch.cuda.manual_seed(seed)
torch.cuda.manual_seed_all(seed)
fix_all_seeds(2021)
# -
class config:
seed = 101
debug = False # set debug=False for Full Training
exp_name = 'Unet-effnetb2-512x512-aug2'
model_name = 'Unet'
backbone0 = 'efficientnet-b2'
backbone1 = 'efficientnet-b7'
backbone2 = 'efficientnet-b6'
backbone3 = 'efficientnet-b5'
batch_size = 8
train_bs = 24
valid_bs = 48
img_size = [512, 512]
epochs = 5
lr = 5e-3
scheduler = 'CosineAnnealingLR'
min_lr = 1e-6
T_max = int(100*6*1.8)
T_0 = 25
warmup_epochs = 0
wd = 1e-6
n_accumulate = 32//train_bs
# n_fold = 5
num_classes = 1
DEVICE = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
competition = 'sartorius'
_wandb_kernel = 'somusan'
# +
from colorama import init,Fore,Style
class PCOLOR:
init()
# COLORS
green = Fore.GREEN
red = Fore.RED
blue = Fore.BLUE
yellow = Fore.YELLOW
magneta = Fore.MAGENTA
cyan = Fore.CYAN
# BRIGHT COLORS
bgreen = Fore.GREEN + Style.BRIGHT
bred = Fore.RED + Style.BRIGHT
bblue = Fore.BLUE + Style.BRIGHT
byellow = Fore.YELLOW + Style.BRIGHT
bmagneta = Fore.MAGENTA + Style.BRIGHT
bcyan = Fore.CYAN + Style.BRIGHT
# +
SAMPLE_SUBMISSION = '../input/sartorius-cell-instance-segmentation/sample_submission.csv'
TRAIN_CSV = "../input/sartorius-cell-instance-segmentation/train.csv"
TRAIN_PATH = "../input/sartorius-cell-instance-segmentation/train"
TEST_PATH = "../input/sartorius-cell-instance-segmentation/test"
RESNET_MEAN = (0.485, 0.456, 0.406)
RESNET_STD = (0.229, 0.224, 0.225)
# (336, 336)
IMAGE_RESIZE = (512,512)
LEARNING_RATE = 5e-4
EPOCHS = 50
# -
df_train = pd.read_csv(TRAIN_CSV)
df_train.head()
# +
train_img_path = "../input/sartorius-cell-instance-segmentation/train/"
train_mask_path = "../input/sartorius-binary-mask-dataset"
train_img = [train_img_path +i for i in sorted(os.listdir(train_img_path))]
train_mask = [train_mask_path + '/' +i for i in sorted(os.listdir(train_mask_path))]
id_list = []
for i in train_img:
id_list.append(i.split("/")[-1].split(".")[0])
columns_tup = list(zip(id_list,train_img,train_mask))
df_prep = pd.DataFrame(columns_tup,columns=["id",'img_path','mask_path'])
print(df_prep["img_path"][0])
print(df_prep["mask_path"][0])
df_prep.head()
# plt.imshow(plt.imread(df_prep["img_path"][0]))
# plt.imshow(np.load(df_prep["mask_path"][0]))
# +
def rle_decode(mask_rle, shape, color=1):
'''
mask_rle: run-length as string formated (start length)
shape: (height,width) of array to return
Returns numpy array, 1 - mask, 0 - background
'''
s = mask_rle.split()
starts, lengths = [np.asarray(x, dtype=int) for x in (s[0:][::2], s[1:][::2])]
starts -= 1
ends = starts + lengths
img = np.zeros(shape[0] * shape[1], dtype=np.float32)
for lo, hi in zip(starts, ends):
img[lo : hi] = color
return img.reshape(shape)
def build_masks(df_train, image_id, input_shape):
height, width = input_shape
labels = df_train[df_train["id"] == image_id]["annotation"].tolist()
mask = np.zeros((height, width))
for label in labels:
mask += rle_decode(label, shape=(height, width))
mask = mask.clip(0, 1)
return mask
# +
class CellDataset(Dataset):
def __init__(self, df,transforms):
self.df = df
self.base_path = TRAIN_PATH
# self.transforms = Compose([Resize(IMAGE_RESIZE[0], IMAGE_RESIZE[1]),
# Normalize(mean=RESNET_MEAN, std=RESNET_STD, p=1),
# HorizontalFlip(p=0.5),
# VerticalFlip(p=0.5),
# ToTensorV2()])
self.transforms = transforms
self.gb = self.df.groupby('id')
self.image_ids = df.id.unique().tolist()
def __getitem__(self, idx):
image_id = self.image_ids[idx]
df = self.gb.get_group(image_id)
annotations = df['annotation'].tolist()
image_path = os.path.join(self.base_path, image_id + ".png")
# image = Image.open(image_path)
# print("image",image.size)
image = cv2.imread(image_path,cv2.IMREAD_COLOR)
# print("before reshape",image.shape)
# image = np.reshape(image,(3,512,512))
# image = image.resize((512,512))
# print("after reshape",image.shape)
mask = build_masks(df_train, image_id, input_shape=(520, 704))
mask = (mask >= 1).astype('float32')
augmented = self.transforms(image=image, mask=mask)
image = augmented['image'] #.type(torch.LongTensor)
# print("aug shape",image.shape)
mask = augmented['mask'] # .type(torch.LongTensor)
# print(mask.shape)
# print(image.resize((3,512,512)).shape)
image = np.reshape(image.numpy(),(3, 512, 512))
image = torch.from_numpy(image)
# print("img aug type-->",type(image))
# print("after aug -->", image.shape)
# print("mask after aug -->", mask.reshape((1, IMAGE_RESIZE[0], IMAGE_RESIZE[1])).shape)
return image, mask.reshape((1, IMAGE_RESIZE[0], IMAGE_RESIZE[1]))
def __len__(self):
return len(self.image_ids)
# +
train_transform = A.Compose([
A.Resize(*config.img_size),
A.CLAHE(p=0.35),
A.ColorJitter(p=0.5),
A.HorizontalFlip(p=0.5),
A.VerticalFlip(p=0.5),
A.ShiftScaleRotate(shift_limit=0.0625, scale_limit=0.1, rotate_limit=90, p=0.5),
A.OneOf([
A.GridDistortion(num_steps=5, distort_limit=0.05, p=1.0),
A.ElasticTransform(alpha=1, sigma=50, alpha_affine=50, p=1.0)
], p=0.25),
A.CoarseDropout(max_holes=8, max_height=config.img_size[0]//20, max_width=config.img_size[1]//20,
min_holes=5, fill_value=0, mask_fill_value=0, p=0.5),
ToTensorV2()], p=1.0)
valid_transform = A.Compose([
A.Resize(*config.img_size),
ToTensorV2()], p=1.0)
# +
# # ../input/sartorius-cell-instance-segmentation/train/0030fd0e6378.png
# # ../input/sartorius-binary-mask-dataset/0030fd0e6378.npy
# # img = cv2.imread("../input/sartorius-cell-instance-segmentation/train/0030fd0e6378.png")
# # # img = cv2.resize(img,(512,512,3))
# # # img = np.reshape(img,(512,512,3))
# # img = img.resize((IMAGE_RESIZE[0], IMAGE_RESIZE[1]))
# # print(img)
# from PIL import Image
# image = Image.open('./image_name.jpg')
# # print("image",image.size)
# new_image = image.resize((512,512))
# # new_image.size
# new_image
# +
# import requests
# image_url = "https://media.geeksforgeeks.org/wp-content/uploads/20190715202808/ybear3-300x224.jpg"
# img_data = requests.get(image_url).content
# with open('image_name.jpg', 'wb') as handler:
# handler.write(img_data)
# +
from sklearn.model_selection import train_test_split
train_data, valid_data = train_test_split(df_train,test_size = 0.2,random_state=42)
ds_train = CellDataset(train_data,train_transform)
ds_valid = CellDataset(valid_data,valid_transform)
image, mask = ds_valid[1]
# image.shape, mask.shape
# ds_train[0]
# +
# ds_train = ds_train.type(torch.LongTensor)
# +
print("Initializing Datasets and Dataloaders...")
# Creating our dataset
# train_dataset = SkinCancerDataset(csv_file=data_csv_file, root_dir=data_dir, transform=data_transforms['train'])
# print(len(train_dataset))
# train_dataset, val_dataset = torch.utils.data.random_split(train_dataset, [4000, 1000])
# Dataloader iterators, make sure to shuffle
train_dataloader = DataLoader(ds_train, batch_size=config.batch_size, shuffle=False,num_workers=4, pin_memory=True)
val_dataloader = DataLoader(ds_valid, batch_size=config.batch_size, shuffle=False,num_workers=4, pin_memory=True)
# Create training and validation dataloaders
dataloaders_dict = {'train': train_dataloader, 'val': val_dataloader}
# +
batch = next(iter(val_dataloader))
images, masks = batch
idx=1
plt.imshow(images[idx][0], cmap='bone')
plt.show()
plt.imshow(masks[idx][0], alpha=0.3)
plt.show()
plt.imshow(images[idx][0], cmap='bone')
plt.imshow(masks[idx][0], alpha=0.3)
plt.show()
# +
def dice_loss(input, target):
input = torch.sigmoid(input)
smooth = 1.0
iflat = input.view(-1)
tflat = target.view(-1)
intersection = (iflat * tflat).sum()
return ((2.0 * intersection + smooth) / (iflat.sum() + tflat.sum() + smooth))
class FocalLoss(nn.Module):
def __init__(self, gamma):
super().__init__()
self.gamma = gamma
def forward(self, input, target):
if not (target.size() == input.size()):
raise ValueError("Target size ({}) must be the same as input size ({})"
.format(target.size(), input.size()))
max_val = (-input).clamp(min=0)
loss = input - input * target + max_val + \
((-max_val).exp() + (-input - max_val).exp()).log()
invprobs = F.logsigmoid(-input * (target * 2.0 - 1.0))
loss = (invprobs * self.gamma).exp() * loss
return loss.mean()
class MixedLoss(nn.Module):
def __init__(self, alpha, gamma):
super().__init__()
self.alpha = alpha
self.focal = FocalLoss(gamma)
def forward(self, input, target):
loss = self.alpha*self.focal(input, target) - torch.log(dice_loss(input, target))
return loss.mean()
# +
# # !mkdir -p /root/.cache/torch/hub/checkpoints/
# # !cp ../input/pytorch-pretrained-image-models/resnet34.pth /root/.cache/torch/hub/checkpoints/resnet34-333f7ec4.pth
import torch
import collections.abc as container_abcs
torch._six.container_abcs = container_abcs
import segmentation_models_pytorch as smp
# -
model = smp.Unet(config.backbone0, encoder_weights="imagenet", classes=1,activation=None)
# model1 = smp.Unet(config.backbone1, encoder_weights="imagenet", classes=1,activation=None)
# model2 = smp.Unet(config.backbone2, encoder_weights="imagenet", classes=1,activation=None)
# model3 = smp.Unet(config.backbone3, encoder_weights="imagenet", classes=1,activation=None)
# +
# for i in val_dataloader:
# print(i.items())
# break
# +
# def valid_one_step(model,data):
# for k,v in data.items():
# data[k] = v.to(config.DEVICE)
# loss = model(**data)
# return loss
# def validation_one_epoch(model,data_loader):
# model.eval()
# total_loss = 0
# for batch_index, data in enumerate(data_loader):
# loss = validation_one_step(model,data)
# total_loss += loss
# return loss
# -
def eval_loop(model, criterion, eval_loader, device=config.DEVICE):
n_batches = len(val_dataloader)
print(n_batches)
running_loss = 0
model.eval()
with torch.no_grad():
accuracy, f1_scores = [], []
pbar = tqdm(eval_loader, desc='Iterating over evaluation data')
for imgs, masks in pbar:
# pass to device
imgs = imgs.to(device)
masks = masks.to(device)
# forward
out = model(imgs.float())
loss = criterion(out, masks)
running_loss += loss.item()#*imgs.shape[0]
# calculate predictions using output
predicted = (out > 0.5).float()
predicted = predicted.view(-1).cpu().numpy()
labels = masks.view(-1).cpu().numpy()
accuracy.append(accuracy_score(labels, predicted))
f1_scores.append(f1_score(labels, predicted))
acc = sum(accuracy)/len(accuracy)
f1 = sum(f1_scores)/len(f1_scores)
# running_loss /= len(eval_loader.sampler)
epoch_loss = running_loss / n_batches
return {
'accuracy':acc,
'f1_macro':f1,
'loss':running_loss}
# +
# torch.set_default_tensor_type("torch.cuda.FloatTensor")
n_batches = len(train_dataloader)
# model1.cuda()
model.to(config.DEVICE)
model.train()
criterion = MixedLoss(10.0, 2.0)
optimizer = optim.Adam(model.parameters(), lr=LEARNING_RATE)
for epoch in range(1, EPOCHS + 1):
print(f"Starting epoch: {epoch} / {EPOCHS}")
running_loss = 0.0
optimizer.zero_grad()
accuracy, f1_scores = [], []
for batch_idx, batch in enumerate(train_dataloader):
images, masks = batch
images, masks = images.to(config.DEVICE), masks.to(config.DEVICE)
outputs = model(images.float())
loss = criterion(outputs, masks)
loss.backward()
optimizer.step()
optimizer.zero_grad()
running_loss += loss.item()
predicted = (outputs > 0.5).float()
predicted = predicted.view(-1).cpu().numpy()
labels = masks.view(-1).cpu().numpy()
accuracy.append(accuracy_score(labels, predicted))
f1_scores.append(f1_score(labels, predicted))
val_eval = eval_loop(model,criterion,val_dataloader)
# print(type(val_eval))
# print(val_eval["loss"])
# print(val_eval["accuracy"])
# print(val_eval["f1_macro"])
val_loss = val_eval["loss"]
val_acc = val_eval["accuracy"]
val_f1 = val_eval["f1_macro"]
epoch_loss = running_loss / n_batches
epoch_acc = sum(accuracy)/len(accuracy)
epoch_f1 = sum(f1_scores)/len(f1_scores)
print(f"Epoch: {epoch} - Train Loss {epoch_loss:.4f} - Train acc {epoch_acc:.4f} - Train f1 {epoch_f1:.4f}")
print(f"Epoch: {epoch} - Validation Loss {val_loss} - Validation acc {val_acc} - Validation f1 {val_f1}")
# -
# class sartorius_dataset():
# def __init__(self,df,transforms):
# self.df = df
# self.img_path = df["img_path"].values()
# self.mask_path = df["mask_path"].values()
# self.transforms = transforms
# def __len__(self):
# return len(self.df)
# def __getitem__(self,idx):
# img_file = self.img_path[idx]
# img = cv2.imread(img_file,cv2.IMREAD_COLOR)
# mask_file = self.mask_path[idx]
# mask = np.load(mask_file)
# data = self.transforms(image = img,mask=mask)
# mask = data["mask"]
# mask = np.expand_dims(mask,axis=0)
# return data["image"], mask
class BuildDataset(torch.utils.data.Dataset):
def __init__(self, df, transforms=None):
self.df = df
# self.img_paths = df["img_path"].values()
self.img_paths = np.array(df['img_path'])
try: # if there is no mask then only send images --> test data
# self.mask_paths = df["mask_path"].values()
self.msk_paths = np.array(df['mask_path'])
except:
self.msk_paths = None
self.transforms = transforms
def __len__(self):
return len(self.df)
def __getitem__(self, index):
img_path = self.img_paths[index]
img = cv2.imread(img_path,cv2.IMREAD_COLOR)
if self.msk_paths is not None:
msk_path = self.msk_paths[index]
msk = np.load(msk_path)
if self.transforms:
data = self.transforms(image=img, mask=msk)
img = data['image']
msk = data['mask']
msk = np.expand_dims(msk, axis=0) # output_shape: (batch_size, 1, img_size, img_size)
return img, msk
else:
if self.transforms:
data = self.transforms(image=img)
img = data['image']
return img
# +
# class CellDataset(Dataset):
# def __init__(self, df):
# self.df = df
# self.base_path = TRAIN_PATH
# self.transforms = Compose([Resize(IMAGE_RESIZE[0], IMAGE_RESIZE[1]),
# Normalize(mean=RESNET_MEAN, std=RESNET_STD, p=1),
# HorizontalFlip(p=0.5),
# VerticalFlip(p=0.5),
# ToTensorV2()])
# self.gb = self.df.groupby('id')
# self.image_ids = df.id.
# +
unique().tolist()
# def __getitem__(self, idx):
# image_id = self.image_ids[idx]
# df = self.gb.get_group(image_id)
# annotations = df['annotation'].tolist()
# image_path = os.path.join(self.base_path, image_id + ".png")
# image = cv2.imread(image_path)
# mask = build_masks(df_train, image_id, input_shape=(520, 704))
# mask = (mask >= 1).astype('float32')
# augmented = self.transforms(image=image, mask=mask)
# image = augmented['image']
# mask = augmented['mask']
# return image, mask.reshape((1, IMAGE_RESIZE[0], IMAGE_RESIZE[1]))
# def __len__(self):
# return len(self.image_ids)
# +
train_transform = A.Compose([
A.Resize(*config.img_size),
A.CLAHE(p=0.35),
A.ColorJitter(p=0.5),
A.HorizontalFlip(p=0.5),
A.VerticalFlip(p=0.5),
A.ShiftScaleRotate(shift_limit=0.0625, scale_limit=0.1, rotate_limit=90, p=0.5),
A.OneOf([
A.GridDistortion(num_steps=5, distort_limit=0.05, p=1.0),
A.ElasticTransform(alpha=1, sigma=50, alpha_affine=50, p=1.0)
], p=0.25),
A.CoarseDropout(max_holes=8, max_height=config.img_size[0]//20, max_width=config.img_size[1]//20,
min_holes=5, fill_value=0, mask_fill_value=0, p=0.5),
ToTensorV2()], p=1.0)
valid_transform = A.Compose([
A.Resize(*config.img_size),
ToTensorV2()], p=1.0)
# +
from sklearn.model_selection import train_test_split
train_data, valid_data = train_test_split(df_prep,test_size = 0.2,random_state=42)
ds_train = BuildDataset(train_data,train_transform)
ds_valid = BuildDataset(valid_data,valid_transform)
image, mask = ds_valid[1]
image.shape, mask.shape
# ds_train[0]
# +
print("Initializing Datasets and Dataloaders...")
# Creating our dataset
# train_dataset = SkinCancerDataset(csv_file=data_csv_file, root_dir=data_dir, transform=data_transforms['train'])
# print(len(train_dataset))
# train_dataset, val_dataset = torch.utils.data.random_split(train_dataset, [4000, 1000])
# Dataloader iterators, make sure to shuffle
train_dataloader = DataLoader(ds_train, batch_size=config.batch_size, shuffle=False,num_workers=4, pin_memory=True)
val_dataloader = DataLoader(ds_valid, batch_size=config.batch_size, shuffle=False,num_workers=4, pin_memory=True)
# Create training and validation dataloaders
dataloaders_dict = {'train': train_dataloader, 'val': val_dataloader}
# -
dataloaders_dict
plt.imshow(image[0], cmap='bone')
plt.show()
plt.imshow(mask[0], alpha=0.3)
plt.show()
# +
# dl_train = DataLoader(ds_train, batch_size=64, num_workers=4, pin_memory=True, shuffle=False)
# -
len(val_dataloader)
batch = next(iter(val_dataloader))
images, masks = batch
idx=1
plt.imshow(images[idx][0], cmap='bone')
plt.show()
plt.imshow(masks[idx][0], alpha=0.3)
plt.show()
plt.imshow(images[idx][0], cmap='bone')
plt.imshow(masks[idx][0], alpha=0.3)
plt.show()
# +
def dice_loss(input, target):
input = torch.sigmoid(input)
smooth = 1.0
iflat = input.view(-1)
tflat = target.view(-1)
intersection = (iflat * tflat).sum()
return ((2.0 * intersection + smooth) / (iflat.sum() + tflat.sum() + smooth))
class FocalLoss(nn.Module):
def __init__(self, gamma):
super().__init__()
self.gamma = gamma
def forward(self, input, target):
if not (target.size() == input.size()):
raise ValueError("Target size ({}) must be the same as input size ({})"
.format(target.size(), input.size()))
max_val = (-input).clamp(min=0)
loss = input - input * target + max_val + \
((-max_val).exp() + (-input - max_val).exp()).log()
invprobs = F.logsigmoid(-input * (target * 2.0 - 1.0))
loss = (invprobs * self.gamma).exp() * loss
return loss.mean()
class MixedLoss(nn.Module):
def __init__(self, alpha, gamma):
super().__init__()
self.alpha = alpha
self.focal = FocalLoss(gamma)
def forward(self, input, target):
loss = self.alpha*self.focal(input, target) - torch.log(dice_loss(input, target))
return loss.mean()
# +
# # !mkdir -p /root/.cache/torch/hub/checkpoints/
# # !cp ../input/pytorch-pretrained-image-models/resnet34.pth /root/.cache/torch/hub/checkpoints/resnet34-333f7ec4.pth
import torch
import collections.abc as container_abcs
torch._six.container_abcs = container_abcs
import segmentation_models_pytorch as smp
# -
# # U-Net
model = smp.Unet(config.backbone0, encoder_weights="imagenet", classes=1,activation=None)
model1 = smp.Unet(config.backbone1, encoder_weights="imagenet", classes=1,activation=None)
model2 = smp.Unet(config.backbone2, encoder_weights="imagenet", classes=1,activation=None)
model3 = smp.Unet(config.backbone3, encoder_weights="imagenet", classes=1,activation=None)
# +
# Check model details
# model
# -
# ## Model0
# +
from torch.utils.tensorboard import SummaryWriter
def train_loop(model, optimizer, criterion, train_loader, device=config.DEVICE):
running_loss = 0
model.train()
pbar = tqdm(train_loader, desc='Iterating over train data')
for imgs, masks in pbar:
# pass to device
imgs = imgs.to(device)
masks = masks.to(device)
# forward
out = model(imgs)
loss = criterion(out, masks)
running_loss += loss.item()*imgs.shape[0] # += loss * current batch size
# optimize
optimizer.zero_grad()
loss.backward()
optimizer.step()
running_loss /= len(train_loader.sampler)
return running_loss
def eval_loop(model, criterion, eval_loader, device=config.DEVICE):
running_loss = 0
model.eval()
with torch.no_grad():
accuracy, f1_scores = [], []
pbar = tqdm(eval_loader, desc='Iterating over evaluation data')
for imgs, masks in pbar:
# pass to device
imgs = imgs.to(device)
masks = masks.to(device)
# forward
out = model(imgs)
loss = criterion(out, masks)
running_loss += loss.item()*imgs.shape[0]
# calculate predictions using output
predicted = (out > 0.5).float()
predicted = predicted.view(-1).cpu().numpy()
labels = masks.view(-1).cpu().numpy()
accuracy.append(accuracy_score(labels, predicted))
f1_scores.append(f1_score(labels, predicted))
acc = sum(accuracy)/len(accuracy)
f1 = sum(f1_scores)/len(f1_scores)
running_loss /= len(eval_loader.sampler)
return {
'accuracy':acc,
'f1_macro':f1,
'loss':running_loss}
def train(model, optimizer, criterion, train_loader, valid_loader,
device=config.DEVICE,
num_epochs=30,
valid_loss_min=np.inf,
logdir='logdir'):
tb_writer = SummaryWriter(log_dir=logdir)
for e in range(num_epochs):
# train for epoch
train_loss = train_loop(
model, optimizer, criterion, train_loader, device=device)
# evaluate on validation set
metrics = eval_loop(
model, criterion, valid_loader, device=device
)
# show progress
print_string = f'Epoch: {e+1} '
print_string+= f'TrainLoss: {train_loss:.5f} '
print_string+= f'ValidLoss: {metrics["loss"]:.5f} '
print_string+= f'ACC: {metrics["accuracy"]:.5f} '
print_string+= f'F1: {metrics["f1_macro"]:.3f}'
print(print_string)
# Tensorboards Logging
tb_writer.add_scalar('UNet/Train Loss', train_loss, e)
tb_writer.add_scalar('UNet/Valid Loss', metrics["loss"], e)
tb_writer.add_scalar('UNet/Accuracy', metrics["accuracy"], e)
tb_writer.add_scalar('UNet/F1 Macro', metrics["f1_macro"], e)
# save the model
if metrics["loss"] <= valid_loss_min:
torch.save(model.state_dict(), 'UNet.pt')
valid_loss_min = metrics["loss"]
# set_seed(21)
# model = UNet(3, 1).to(device)
model.to(device=config.DEVICE)
optimizer = optim.Adam(model.parameters(), lr=config.lr)
criterion = MixedLoss(10.0, 2.0)
train(model, optimizer, criterion, train_dataloader, val_dataloader)
# +
def save_checkpoint(state, filename="my_checkpoint.pth.tar"):
print("=> Saving checkpoint")
torch.save(state, filename)
def load_checkpoint(checkpoint, model):
print("=> Loading checkpoint")
model.load_state_dict(checkpoint["state_dict"])
def train_fn(loader, model, optimizer, loss_fn, scaler):
loop = tqdm(loader)
for batch_idx, (data, targets) in enumerate(loop):
data = data.to(device=DEVICE)
targets = targets.float().unsqueeze(1).to(device=DEVICE)
# forward
with torch.cuda.amp.autocast():
predictions = model(data)
loss = loss_fn(predictions, targets)
# backward
optimizer.zero_grad()
scaler.scale(loss).backward()
scaler.step(optimizer)
scaler.update()
# update tqdm loop
loop.set_postfix(loss=loss.item())
# model = UNET(in_channels=3, out_channels=1).to(config.DEVICE)
# loss_fn = nn.BCEWithLogitsLoss()
loss_fn = MixedLoss(10.0, 2.0)
optimizer = optim.Adam(model.parameters(), lr=config.lr)
# train_loader, val_loader = get_loaders(
# TRAIN_IMG_DIR,
# TRAIN_MASK_DIR,
# VAL_IMG_DIR,
# VAL_MASK_DIR,
# BATCH_SIZE,
# train_transform,
# val_transforms,
# NUM_WORKERS,
# PIN_MEMORY,
# )
# if LOAD_MODEL:
# load_checkpoint(torch.load("my_checkpoint.pth.tar"), model)
# check_accuracy(val_loader, model, device=DEVICE)
scaler = torch.cuda.amp.GradScaler()
for epoch in range(NUM_EPOCHS):
train_fn(train_loader, model, optimizer, loss_fn, scaler)
# save model
checkpoint = {
"state_dict": model.state_dict(),
"optimizer":optimizer.state_dict(),
}
save_checkpoint(checkpoint)
# check accuracy
check_accuracy(val_loader, model, device=DEVICE)
# print some examples to a folder
save_predictions_as_imgs(
val_loader, model, folder="saved_images/", device=DEVICE
)
# +
# from torch.multiprocessing import Pool, Process, set_start_method
# try:
# set_start_method('spawn', force=True)
# # except RuntimeError:
# pass
torch.set_default_tensor_type("torch.cuda.FloatTensor")
n_batches = len(train_dataloader)
# torch.multiprocessing.set_start_method('spawn')
model.to(device=DEVICE)
model.train()
criterion = MixedLoss(10.0, 2.0)
optimizer = optim.Adam(model.parameters(), lr=LEARNING_RATE)
for epoch in range(1, EPOCHS + 1):
print(f"Starting epoch: {epoch} / {EPOCHS}")
running_loss = 0.0
optimizer.zero_grad()
for batch_idx, batch in enumerate(train_dataloader):
# Predict
images, masks = batch
images, masks = images.cuda(), masks.cuda()
outputs = model(images)
loss = criterion(outputs, masks)
# Back prop
loss.backward()
optimizer.step()
optimizer.zero_grad()
running_loss += loss.item()
epoch_loss = running_loss / n_batches
print(f"Epoch: {epoch} - Train Loss {epoch_loss:.4f}")
# +
FILE = "./model0_50.pth"
torch.save(model.state_dict(), FILE)
wandb.save("./model0_50.pth")
# print(model.state_dict())
# loaded_model = Model(n_input_features=6)
# loaded_model.load_state_dict(torch.load(FILE)) # it takes the loaded dictionary, not the path file itself
# loaded_model.eval()
# -
# ### Model1
# +
torch.set_default_tensor_type("torch.cuda.FloatTensor")
n_batches = len(dl_train)
model1.cuda()
model1.train()
criterion = MixedLoss(10.0, 2.0)
optimizer = optim.Adam(model.parameters(), lr=LEARNING_RATE)
for epoch in range(1, EPOCHS + 1):
print(f"Starting epoch: {epoch} / {EPOCHS}")
running_loss = 0.0
optimizer.zero_grad()
for batch_idx, batch in enumerate(dl_train):
images, masks = batch
images, masks = images.cuda(), masks.cuda()
outputs = model(images)
loss = criterion(outputs, masks)
loss.backward()
optimizer.step()
optimizer.zero_grad()
running_loss += loss.item()
epoch_loss = running_loss / n_batches
print(f"Epoch: {epoch} - Train Loss {epoch_loss:.4f}")
# +
FILE1 = "./model1_50.pth"
torch.save(model1.state_dict(), FILE1)
wandb.save("./model1_50.pth")
# -
# ## Model2
# +
torch.set_default_tensor_type("torch.cuda.FloatTensor")
n_batches = len(dl_train)
model2.cuda()
model3.train()
criterion = MixedLoss(10.0, 2.0)
optimizer = optim.Adam(model.parameters(), lr=LEARNING_RATE)
for epoch in range(1, EPOCHS + 1):
print(f"Starting epoch: {epoch} / {EPOCHS}")
running_loss = 0.0
optimizer.zero_grad()
for batch_idx, batch in enumerate(dl_train):
images, masks = batch
images, masks = images.cuda(), masks.cuda()
outputs = model(images)
loss = criterion(outputs, masks)
loss.backward()
optimizer.step()
optimizer.zero_grad()
running_loss += loss.item()
epoch_loss = running_loss / n_batches
print(f"Epoch: {epoch} - Train Loss {epoch_loss:.4f}")
# +
FILE2 = "./model2_50.pth"
torch.save(model2.state_dict(), FILE2)
wandb.save("./model2_50.pth")
# -
# ## model3
# +
torch.set_default_tensor_type("torch.cuda.FloatTensor")
n_batches = len(dl_train)
model3.cuda()
model3.train()
criterion = MixedLoss(10.0, 2.0)
optimizer = optim.Adam(model.parameters(), lr=LEARNING_RATE)
for epoch in range(1, EPOCHS + 1):
print(f"Starting epoch: {epoch} / {EPOCHS}")
running_loss = 0.0
optimizer.zero_grad()
for batch_idx, batch in enumerate(dl_train):
# Predict
images, masks = batch
images, masks = images.cuda(), masks.cuda()
outputs = model(images)
loss = criterion(outputs, masks)
# Back prop
loss.backward()
optimizer.step()
optimizer.zero_grad()
running_loss += loss.item()
epoch_loss = running_loss / n_batches
print(f"Epoch: {epoch} - Train Loss {epoch_loss:.4f}")
# +
FILE3 = "./model3_50.pth"
torch.save(model3.state_dict(), FILE3)
wandb.save("./model3_50.pth")
# -
# # Predict
class TestCellDataset(Dataset):
def __init__(self):
self.test_path = TEST_PATH
self.image_ids = [f[:-4]for f in os.listdir(self.test_path)]
self.num_samples = len(self.image_ids)
self.transform = Compose([Resize(IMAGE_RESIZE[0], IMAGE_RESIZE[1]), Normalize(mean=RESNET_MEAN, std=RESNET_STD, p=1), ToTensorV2()])
def __getitem__(self, idx):
image_id = self.image_ids[idx]
path = os.path.join(self.test_path, image_id + ".png")
image = cv2.imread(path)
image = self.transform(image=image)['image']
return {'image': image, 'id': image_id}
def __len__(self):
return self.num_samples
del dl_train, ds_train, optimizer
ds_test = TestCellDataset()
dl_test = DataLoader(ds_test, batch_size=64, shuffle=False, num_workers=4, pin_memory=True)
# ### Utilities
# +
def post_process(probability, threshold=0.5, min_size=300):
mask = cv2.threshold(probability, threshold, 1, cv2.THRESH_BINARY)[1]
num_component, component = cv2.connectedComponents(mask.astype(np.uint8))
predictions = []
for c in range(1, num_component):
p = (component == c)
if p.sum() > min_size:
a_prediction = np.zeros((520, 704), np.float32)
a_prediction[p] = 1
predictions.append(a_prediction)
return predictions
def rle_encoding(x):
dots = np.where(x.flatten() == 1)[0]
run_lengths = []
prev = -2
for b in dots:
if (b>prev+1): run_lengths.extend((b + 1, 0))
run_lengths[-1] += 1
prev = b
return ' '.join(map(str, run_lengths))
# -
pd.read_csv(SAMPLE_SUBMISSION)
# +
def check_is_run_length(mask_rle):
if not mask_rle:
return True
s = mask_rle.split()
starts, lengths = [np.asarray(x, dtype=int) for x in (s[0:][::2], s[1:][::2])]
starts -= 1
ends = starts + lengths
start_prev = starts[0]
ok = True
for start in starts[1:]:
ok = ok and start > start_prev
start_prev = start
if not ok:
return False
return True
def create_empty_submission():
fs = os.listdir("../input/sartorius-cell-instance-segmentation/test")
df = pd.DataFrame([(f[:-4], "") for f in fs], columns=['id', 'predicted'])
df.to_csv("submission.csv", index=False)
# +
model.eval()
submission = []
for i, batch in enumerate(tqdm(dl_test)):
preds = torch.sigmoid(model(batch['image'].cuda()))
preds = preds.detach().cpu().numpy()[:, 0, :, :] # (batch_size, 1, size, size) -> (batch_size, size, size)
for image_id, probability_mask in zip(batch['id'], preds):
try:
#if probability_mask.shape != IMAGE_RESIZE:
# probability_mask = cv2.resize(probability_mask, dsize=IMAGE_RESIZE, interpolation=cv2.INTER_LINEAR)
probability_mask = cv2.resize(probability_mask, dsize=(704, 520), interpolation=cv2.INTER_LINEAR)
predictions = post_process(probability_mask)
for prediction in predictions:
#plt.imshow(prediction)
#plt.show()
try:
submission.append((image_id, rle_encoding(prediction)))
except:
print("Error in RL encoding")
except Exception as e:
print(f"Exception for img: {image_id}: {e}")
# Fill images with no predictions
image_ids = [image_id for image_id, preds in submission]
if image_id not in image_ids:
submission.append((image_id, ""))
df_submission = pd.DataFrame(submission, columns=['id', 'predicted'])
df_submission.to_csv('submission.csv', index=False)
if df_submission['predicted'].apply(check_is_run_length).mean() != 1:
print("Check run lenght failed")
create_empty_submission()
# -
# !rm -r ./wandb
|
Inference/unet-pretrained-ensemble-starter-draft-1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Process
# - A running instance of a computer program
#
# ## 1. Processes vs Threads
# - Process: Sidesteps GIL, Less need for synchronization, Can be paused and terminated, more resilient
# - Thread: Higher memory footprint, expensive content switches
#
# ## 2. Simple multiprocessing pattern
# +
import multiprocessing
import time
def do_some_work(val):
print("Doing some work in thread")
time.sleep(1)
print("echo: {}".format(val))
if __name__ == "__main__":
val = "text"
p = multiprocessing.Process(target=do_some_work, args=(val,))
p.start()
print("Start thread, process alive: {}".format(p.is_alive()))
# p.terminate() # Terminate the process
p.join()
print("End thread")
# -
# ## 3. Terms
# - Pickle: Process whereby a Python object hierarchy is converted into a byte stream. "Unpickling" is the inverse operation.
# - Deamon Process: A child process that does not prevent its parent process from exiting
#
# ## 4. Other operation
# - p.is_alive(): Check if process is alive
# - p.terminate(): Terminate a process
# - multiprocessing.cpu_count(): Check # of CPUs
#
# ## 5. Process pool
# - A process pool object which controls a pool of worker processes to which jobs can be submitted. It supports asynchronous results with timeouts and callbacks and has a parallel map implementation.
#
# ### 5.0. Pool example
# +
def do_work(data):
time.sleep(1)
return data**2
def start_process():
print("Start", multiprocessing.current_process().name)
if __name__ == "__main__":
pool_size = multiprocessing.cpu_count() * 2
pool = multiprocessing.Pool(processes=pool_size, initializer=start_process)
inputs = list(range(10))
# map(): block until it's ready
# map_async(): non-block and return a call back
# use .get() on call back object to get result
outputs = pool.map(do_work, inputs)
pool.close() # No more task accepted
pool.join() # Wait for the worker processes to exit
print("Outputs:", outputs)
# -
# ## 6. Inter-process Communication
# ### 6.0. Pipe
# - Communication between process
# +
import random
from multiprocessing import Pipe, Process
import time
def make_tuple(conn):
num = random.randint(1, 9)
conn.send(("Hi", num))
print(conn.recv())
def make_string(conn):
tup = conn.recv()
result = ""
substr, num = tup
for _ in range(num):
result += substr
print(result)
conn.send(result)
if __name__ == "__main__":
conn1, conn2 = Pipe(duplex=True)
p1 = Process(target=make_tuple, args=(conn1,))
p2 = Process(target=make_string, args=(conn2,))
p1.start()
p2.start()
p1.join()
p2.join()
print("Done")
# -
# ### 6.1. Queue
# - Pipe can only have two endpoints
# - Queue can have multiple producers and consumers
# +
""" Data flow
1. make_tuple -> ("Hi", num) -> make_string
2. sleep for 1 second
3. make_string -> result -> make_tuple
4. make_tuple print result by using queue.get()
"""
from multiprocessing import Queue
def make_tuple(queue):
num = random.randint(1, 9)
queue.put(("Hi", num))
time.sleep(1)
print(queue.get()) # Get from 'make_string'
def make_string(queue):
tup = queue.get()
result = ""
substr, num = tup
for _ in range(num):
result += substr
queue.put(result)
if __name__ == "__main__":
queue = Queue()
p1 = Process(target=make_tuple, args=(queue,))
p2 = Process(target=make_string, args=(queue,))
p1.start()
p2.start()
# -
# ## 7. Sharing State Between Processes
# ### 7.0. Value
# +
from multiprocessing import Value
import multiprocessing
import ctypes
counter = Value('i') # shared object of type int, defaults to 0
# shared object of type boolean, defaulting to False, unsynchronized
is_running = Value(ctypes.c_bool, False, lock=False)
my_lock = multiprocessing.Lock()
# Shared object of type long, with a lock specified
size_counter = Value('l', 0, lock=my_lock)
# -
# ### 7.1. Manager
# - Share variables between processes
# +
import multiprocessing
from multiprocessing import Process
def do_work(dictionary, item):
dictionary[item] = item ** 2
if __name__ == "__main__":
mgr = multiprocessing.Manager()
d = mgr.dict() # Shared dict
# Multiple processes work on same shared-dict
jobs = [
Process(target=do_work, args=(d, i)) for i in range(8)
]
for j in jobs:
j.start()
for j in jobs:
j.join()
print("Results:", d)
|
src/multiprocessing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
import numpy as np
# ## Concat
#
# Add rows or columns to an existing array.
matrix1 = np.arange(20).reshape(10,2)
arr1 = np.arange(10, 30, 2)
arr2 = np.array([99,1001])
matrix2 = np.array([96,97,98,99]).reshape(2,2)
for array1, array2 in zip(matrix1, arr1):
print(array1, array2)
# ### Add row(s) with axis=0
# add single row
np.concatenate((matrix1, arr2.reshape(1,-1)), axis=0)
# add multiple rows
np.concatenate((arr1, arr4), axis=0)
# ### Add column(s) with axis=1
# add column
np.concatenate((arr1, arr2.reshape(-1,1)), axis=1)
# ### Where is this useful?
#
# Let's create a scenario where data is generated on-the-fly and needs appending to our array. Think data matrix here.
# with concatenate
for i in range(10):
row = np.random.binomial(n=1, p=0.5, size=10).reshape(1,-1)
if i == 0:
array = np.array(row)
else:
array = np.concatenate((array, row), axis=0)
array
array.shape
# ### What about np.append()?
# with append
for i in range(10):
row = np.random.binomial(n=1, p=0.5, size=10).reshape(1,-1)
if i == 0:
array = np.array(row)
else:
array = np.append(array, row, axis=0)
array
array.shape
# It turns out that **np.append()** uses **np.concatenate()** under the hood. You may as well just use **np.concatenate()** since that's the case. It's faster for the same reason.
# ## Ravel
#
# Return a contiguous flattened 1D array.
matrix1
# C order is row-wise
ravel_C = np.ravel(matrix1, 'C')
ravel_C
ravel_C.shape
# F order is column-wise
ravel_F = np.ravel(matrix1, 'F')
ravel_F
ravel_F.shape
tensor3d = np.arange(27).reshape(3,3,3)
tensor3d
np.ravel(tensor3d, 'C')
np.ravel(tensor3d, 'F')
# ## Mesh-grid
#
# Returns a dense multi-dimensional “meshgrid”.
# first range sets number of rows and its values
# second range sets number of cols and its values
grid1 = np.mgrid[1:10:2,0:10:2]
grid1
grid1.shape
grid2 = np.mgrid[-100:100:5, range(3)]
grid2
grid2.shape
|
notebooks/Python/NumPy/Numpy_concat_append_ravel_mgrid.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Encapsulation in Python
# <ul><li>Encapsulation is the concept of bundling data and methods within a single unit. For example, when you create a class, it means you are implementing encapsulation. A class is an example of encapsulation as it binds all the data members (instance variables) and methods into a single unit.</li><li>
#
# Using encapsulation, we can hide an object’s internal representation from the outside. This is called information hiding.</li><li>
#
# Encapsulation allows us to restrict accessing variables and methods directly and prevent accidental data modification by creating private data members and methods within a class.</li><li>
#
# Encapsulation is a way to restrict access to methods and variables from outside of class. Whenever working with the class and dealing with sensitive data, providing access to all variables used within the class is not a good choice.</li></ul>
# +
# A program that displays employees information
class Employee:
# constructor
def __init__(self, name, salary, project):
# instance variables
self.name = name
self.salary = salary
self.project = project
# method to display employee's details
def show(self):
# accessing public instance variables
print("\nName: " , self.name, '\nSalary:', "N"+str(self.salary))
# method
def work(self):
print(self.name, 'is working on', self.project)
name = input("Enter your name: ")
salary = int(input("How much do you earn: "))
project = input("What project are you working on: ")
# creating object of a class
emp = Employee(name, salary, project)
# calling public method of the class
emp.show()
emp.work()
# -
# ## Access Modifiers in Python
# <ul><li>Encapsulation can be achieved by declaring the data members and methods of a class either as private or protected.</li><li> In Python, there is no direct access modifiers like public, private, and protected. </li><li>This can achieve by using single underscore and double underscores.</li></ul>
#
# Access modifiers limit access to the variables and methods of a class. Python provides three types of access modifiers private, public, and protected.
#
# <ul><li><b>Public Instance Variable:</b> Accessible anywhere from outside oclass.</li><li>
# <b>Private Instance Variable:</b> Accessible within the class.</li><li>
# <b>Protected Instance Variable:</b> Accessible within the class and its sub-classes.</li></ul>
# ### Public Instance Variable
# Public instance variables are accessible within and outside of a class. All member variables of the class are by default public.
# +
class Employee:
# constructor
def __init__(self, name, salary):
# public instance variables
self.name = name
self.salary = salary
# public instance methods
def show(self):
# accessing public instance variables
print("Name: ", self.name, 'Salary:', "N%s"%(self.salary))
# creating object of a class
emp = Employee('Abdurrahman', 500000)
# accessing public instance variables
print("Name: ", emp.name, 'Salary:', "N{}".format(emp.salary))
# calling public method of the class
emp.show()
# -
# ### Private Instance Variable
# <ul><li>Protect variables in the class by marking them private. To define a private variable add two underscores as a prefix at the start of a variable name.</li><li>
#
# Private instance variables are accessible only within the class, and we can’t access them directly from the class objects.</li></ul>
# +
class Employee:
# constructor
def __init__(self, name, salary):
# public instance variable
self.name = name
# private variable
self.__salary = salary
# creating object of a class
emp = Employee('<NAME>', 10000)
# accessing public instance variables
print("Name: ", emp.name)
# accessing private instance variable
print('Salary:', emp._Employee__salary)
# -
# To access private members from outside of a class using the following two approaches
#
# <ul><li>Create public method to access private members</li><li>
# Use name mangling</li></ul>
# #### Access Private member outside of a class using an instance method
# +
class Employee:
# constructor
def __init__(self, name, salary):
# public data member
self.name = name
# private member
self.__salary = salary
# public instance methods
def show(self):
# private members are accessible from a class
print("Name: ", self.name, '\nSalary:', self.__salary)
# creating object of a class
emp = Employee('<NAME>', 250000)
# calling public method of the class
emp.show()
# -
# #### Name Mangling to access private members
# <ul><li>Private and protected variables can be directly accessed from outside of a class through name mangling.</li><li> The name mangling is created on an identifier by adding two leading underscores and one trailing underscore, like this <b>_classname__dataMember</b>, where <b><i>classname</i></b> is the current class, and data member is the private variable name.</li></ul>
# +
class Employee:
# constructor
def __init__(self, name, salary):
# public instance variable
self.name = name
# private variable
self.__salary = salary
# creating object of a class
emp = Employee('<NAME>', 900000)
#direct access to public instance variable
print('Name:', emp.name)
# direct access to private instance variable using name mangling
print('Salary:', "N"+str(emp._Employee__salary))
# -
# ### Protected Instance Variable
# <ul><li>Protected instance variables are accessible within the class and also available to its sub-classes. </li><li>To define a protected variable, prefix the variable name with a single underscore <b>_</b>.</li><li>
#
# Protected instance variables are used when you implement inheritance and want to allow data members access to only child classes.</li></ul>
# +
# base class
class Company:
# base constructor
def __init__(self):
# Protected instance variable
self._project = "Blockchain Development"
# child class
class Employee(Company):
# child constructor
def __init__(self, name):
self.name = name
# invoke base constructor
Company.__init__(self)
def show(self):
print("Employee name :", self.name)
# Accessing protected instance variable in child class
print("Working on project :", self._project)
c = Employee("<NAME>")
c.show()
# Direct access to protected instance variable
print('Project:', c._project)
# -
# ## Getters and Setters in Python
# <ul><li>To implement proper encapsulation in Python, setters and getters can be used.</li><li> The primary purpose of using getters and setters in object-oriented programs is to ensure data encapsulation.</li><li> Use the getter method to access instance variables and the setter methods to modify the instance variables.</li></ul>
#
# In Python, private variables are not hidden fields like in other programming languages. The getters and setters methods are often used when:
#
# <ul><li>When we want to avoid direct access to private variables</li><li>
# To add validation logic for setting a value</li></ul>
# +
class Student:
def __init__(self, name, age):
# public instance variable
self.name = name
# private instance variable
self.__age = age
# getter method
def get_age(self):
return self.__age
# setter method
def set_age(self, age):
self.__age = age
stud = Student('<NAME>', 34)
# retrieving age using getter
print('Name:', stud.name, "\nAge:", stud.get_age())
# changing age using setter
stud.set_age(26)
# retrieving age using getter
print('\n\nName:', stud.name, "\nAge:", stud.get_age())
# +
class Student:
# contructor
def __init__(self, name, roll_no, age):
# public instance variable
self.name = name
# private instance variables to restrict access
# avoid direct data modification
self.__roll_no = roll_no
self.__age = age
def show(self):
print('Student Details:', self.name, self.__roll_no)
# getter methods
def get_roll_no(self):
return self.__roll_no
# setter method to modify instance varaible
# condition to allow data modification with rules
def set_roll_no(self, number):
if number > 50:
print('Invalid roll no. Please set correct roll number')
else:
self.__roll_no = number
# object instanc=tiation
info = Student('<NAME>', 10, 15)
# before Modify
info.show()
# changing roll number using setter
info.set_roll_no(120)
info.set_roll_no(25)
info.show()
# -
# ## Class Project I
# You have been contracted by the Registrar of Pan-Atlantic University (PAU) as an expert OOP developer to access the Student Information System (SIS) of PAU Student Council, inorder to classify the students grades, according to their age, into 3 categories; the pirates, the yankees and the bulls.
#
# Should you choose to accept this task, develop an OOP program that reads data from the sis.csv file as attached, following the instructions below:
#
# <b>Instructions:</b>
# <ul><li>If the student age is greater than 14 and less than 18, create a .csv file for that category called <b>The_Pirates</b> and display it.</li><li>
# If the student age is greater than 18 and less than 22, create a file for that category called <b>The_Yankees</b> and display it.</li><li>
# If the student age is greater than 22 create a file for that category called <b>The_Bulls</b> and display it.</li></ul>
#
import pandas as pd
df = pd.read_csv('sis.csv')
df
|
WEE11/Week 11 - Encapsulation Practice.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # CSE 110 Week 7
#
# ## 07 Prove: Assignment Milestone
#
#
# #### Project Description
# For this project, you will use this idea of looping, or iterating, through each pixel in an image to produce a green screen effect.
#
# #### Assignment
#
# 1. Downloading and saving image files
# 2. Installing and using a Python library
# 3. Opening an image file
# 4. Iterating through the pixels of an image
# 5. Using if statements to make decisions about how to handle each pixel
# 6. Producing a new image file and saving it to your computer
# +
from PIL import Image
# print("The Library is loaded correctly")
# Path to my directory where the images are stored
image_path = "/Users/vernwolfley/Documents/My-Education/CSE-110/assignment_files/cse110_images"
image_og = Image.open(image_path + "/beach.jpg")
# width, height = image_og.size
pixels_og = image_og.load()
# r, g, b = pixels_og[100, 200]
# pixels_og[100, 200] = (r, g, b)
# image_og.save("the_file_goes_here.jpg")
# print(image_og.size)
# print(pixels_og[100, 200])
# print(pixels_og[110, 210])
# print(pixels_og[120, 220])
# print(pixels_og[130, 230])
# Purple Square
# for y in range(100, 210):
# for x in range(100, 200):
# pixels_og[x, y] = (200, 8, 249) #C808F9
# change all pixel values add blue
for y in range(0, 600):
for x in range(0, 800):
r, g, b = pixels_og[x, y]
pixels_og[x, y] = (r, g, 200)
# pixels_og[100, 200] = (200, 8, 249) #C808F9
# pixels_og[101, 200] = (200, 8, 249) #C808F9
# pixels_og[102, 200] = (200, 8, 249) #C808F9
# pixels_og[103, 200] = (200, 8, 249) #C808F9
# pixels_og[104, 200] = (200, 8, 249) #C808F9
# pixels_og[105, 200] = (200, 8, 249) #C808F9
# pixels_og[100, 201] = (200, 8, 249) #C808F9
# pixels_og[101, 201] = (200, 8, 249) #C808F9
# pixels_og[102, 201] = (200, 8, 249) #C808F9
# pixels_og[103, 201] = (200, 8, 249) #C808F9
# pixels_og[104, 201] = (200, 8, 249) #C808F9
# pixels_og[105, 201] = (200, 8, 249) #C808F9
# pixels_og[100, 202] = (200, 8, 249) #C808F9
# pixels_og[101, 202] = (200, 8, 249) #C808F9
# pixels_og[102, 202] = (200, 8, 249) #C808F9
# pixels_og[103, 202] = (200, 8, 249) #C808F9
# pixels_og[104, 202] = (200, 8, 249) #C808F9
# pixels_og[105, 202] = (200, 8, 249) #C808F9
# pixels_og[100, 203] = (200, 8, 249) #C808F9
# pixels_og[101, 203] = (200, 8, 249) #C808F9
# pixels_og[102, 203] = (200, 8, 249) #C808F9
# pixels_og[103, 203] = (200, 8, 249) #C808F9
# pixels_og[104, 203] = (200, 8, 249) #C808F9
# pixels_og[105, 203] = (200, 8, 249) #C808F9
image_og.show()
file_loc = "../assignment_files/cse110_images/"
# image_og.save(file_loc + "purple_square.jpg")
# image_og.save("../assignment_files/cse110_images/blue_beach.jpg")
# -
|
jupyter-notebook-CSE-110/week07-prove-assignment.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Bayesian Statistics Seminar
#
# Copyright 2017 <NAME>
#
# MIT License: https://opensource.org/licenses/MIT
# +
from __future__ import print_function, division
# %matplotlib inline
import warnings
warnings.filterwarnings('ignore')
import numpy as np
import pandas as pd
import thinkbayes2
import thinkplot
# -
# ## Survival analysis
#
# Suppose that you are an auto insurance company interested in the time between collisions for a particular driver. If the probability of a collision is roughly constant over time, the time between collisions will follow an exponential distribution.
#
# Here's an example with parameter $\lambda = 0.5$.
# +
from thinkbayes2 import MakeExponentialPmf
pmf = MakeExponentialPmf(lam=0.5, high=30)
thinkplot.Pdf(pmf)
thinkplot.Config(xlabel='Lifetime', ylabel='PMF')
# -
# For the exponential distribution, the mean and standard deviation are $1/\lambda$.
#
# In this case they are only approximate because we truncated the distribution.
pmf.Mean(), pmf.Std()
# From the PMF, we can compute the CDF.
cdf = pmf.MakeCdf()
thinkplot.Cdf(cdf)
thinkplot.Config(xlabel='Lifetime', ylabel='CDF')
# And from the CDF, we can compute the survival function, which is the complement of the CDF.
#
# $SF(x) = Prob\{X > x\} = 1 - Prob\{X \le x\} = 1 - CDF(x)$
# +
from survival import MakeSurvivalFromCdf
sf = MakeSurvivalFromCdf(cdf)
thinkplot.Plot(sf)
thinkplot.Config(xlabel='Lifetime', ylabel='Survival function')
# -
# From the survival function we can get the hazard function, which is the probability of a collision at $x$, given no collision prior to $x$.
hf = sf.MakeHazardFunction()
thinkplot.Plot(hf)
thinkplot.Config(xlabel='Lifetime', ylabel='Hazard function')
# If the distribution is truly exponential, the hazard function is constant for all $x$.
#
# In this case it goes to 1 at the end, again because we truncated the distribution.
# **Exercise:** Go back and increase the value of `high`, and confirm that the hazard function is a constant until we approach the point where we cut off the distribution.
# Given the survival function, we can compute the distribution of remaining lifetime, conditioned on current age. The following function computes the mean remaining lifetime for a range of ages.
def RemainingLifetime(sf):
"""Computes remaining lifetime as a function of age.
sf: survival function
returns: Series that maps from age to remaining lifetime
"""
pmf = sf.MakePmf()
d = {}
for t in sorted(pmf.Values()):
pmf[t] = 0
if pmf.Total():
pmf.Normalize()
d[t] = pmf.Mean() - t
return pd.Series(d)
# And here's what it looks like for the exponential survival function.
mean_rem_life = RemainingLifetime(sf)
thinkplot.Plot(mean_rem_life)
thinkplot.Config(xlabel='Lifetime', ylabel='Survival function')
# The mean time until a collision is pretty much constant, until we approach the point where we truncate the distribution.
# ## The Weibull distribution
#
# The Weibull distribution is a generalization of the exponential distribution that takes an additional "shape" parameter, `k`.
#
# When `k=1`, the Weibull is an exponential distribution. Other values of `k` yield survival curves with different shapes, and hazard functions that increase, decrease, or both. So the Weibull family can capture a wide range of survival patterns.
# +
from thinkbayes2 import MakeWeibullPmf
pmf = MakeWeibullPmf(lam=2.0, k=1.5, high=30)
thinkplot.Pdf(pmf)
thinkplot.Config(xlabel='Lifetime', ylabel='PMF')
# -
# **Exercise**: In the previous section, replace the exponential distribution with a Weibull distribituion and run the analysis again. What can you infer about the values of the parameters and the behavior of the hazard function and remaining lifetime?
# ## Bayesian survival analysis
# Suppose you are the manager of a large building with many light fixtures. To figure out how often you will need to replace lightbulbs, you install 10 bulbs and measure the time until they fail.
#
# To generate some fake data, I'll choose a Weibull distribution and generate a random sample (let's suppose it's in years):
# +
def SampleWeibull(lam, k, n=1):
return np.random.weibull(k, size=n) * lam
data = SampleWeibull(lam=2, k=1.5, n=10)
data
# -
# **Exercise:** Write a class called `LightBulb` that inherits from `Suite` and provides a `Likelihood` function that takes an observed lifespan as data and a tuple, `(lam, k)`, as a hypothesis. It should return a likelihood proportional to the probability of the observed lifespan in a Weibull distribution with the given parameters.
#
# Test your method by creating a `LightBulb` object with an appropriate prior and update it with the data above.
#
# Plot the posterior distributions of `lam` and `k`. As the sample size increases, does the posterior distribution converge on the values of `lam` and `k` used to generate the sample?
# +
# Hint
from thinkbayes2 import Suite, Joint, EvalWeibullPdf
class LightBulb(Suite, Joint):
def Likelihood(self, data, hypo):
lam, k = hypo
x = data
like = 1
return like
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# -
# **Exercise:** Go back and run this analysis again with `n=20` and see if the posterior distributions seem to be converging on the actual parameters.
# ## Censored data
# **Exercise:** Now suppose that instead of observing a complete lifespan, you observe a lightbulb that has operated for 1 year and is still working. Write another version of `LightBulb` that takes data in this form and performs an update.
# +
# Hint
from thinkbayes2 import EvalWeibullCdf
class LightBulb2(Suite, Joint):
def Likelihood(self, data, hypo):
lam, k = hypo
x = data
like = 1
return like
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# -
# Note: based on this data alone, we can rule out some small values of `lam` and `k`, but we can't rule out large values. Without more data or a more informative prior, the results are not useful.
#
# To see why, try increasing the upper bounds in the prior distribition.
# **Exercise:** Suppose you install a light bulb and then you don't check on it for a year, but when you come back, you find that it has burned out. Extend `LightBulb` to handle this kind of data, too.
# +
# Hint
class LightBulb3(Suite, Joint):
def Likelihood(self, data, hypo):
lam, k = hypo
x = data
like = 1
return like
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# -
# This example has some of the same problems as the previous one. Based on this data alone, we can't pin down the parameters much.
# ## Pulling it together
# **Exercise:** Suppose you have 15 lightbulbs installed at different times over a 10 year period. When you observe them, some have died and some are still working. Write a version of `LightBulb` that takes data in the form of a `(flag, x)` tuple, where:
#
# 1. If `flag` is `eq`, it means that `x` is the actual lifespan of a bulb that has died.
# 2. If `flag` is `gt`, it means that `x` is the current age of a bulb that is still working, so it is a lower bound on the lifespan.
# 3. If `flag` is `lt`, it means that `x` is the elapsed time between installation and the first time the bulb is seen broken, so it is an upper bound on the lifespan.
# To help you test, I will generate some fake data.
#
# First, I'll generate a Pandas DataFrame with random start times and lifespans. The columns are:
#
# * `start`: time when the bulb was installed
#
# * `lifespan`: lifespan of the bulb in years
#
# * `end`: time when bulb died or will die
#
# * `age_t`: age of the bulb at t=10
# +
import pandas as pd
lam = 2
k = 1.5
n = 15
t_end = 10
starts = np.random.uniform(0, t_end, n)
lifespans = SampleWeibull(lam, k, n)
df = pd.DataFrame({'start': starts, 'lifespan': lifespans})
df['end'] = df.start + df.lifespan
df['age_t'] = t_end - df.start
df.head()
# -
# Now I'll process the DataFrame to generate data in the form we want for the update.
# +
data = []
for i, row in df.iterrows():
if row.end < t_end:
data.append(('eq', row.lifespan))
else:
data.append(('gt', row.age_t))
for pair in data:
print(pair)
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# -
# ## Prediction
#
# Suppose we know that, for a particular kind of lightbulb in a particular location, the distribution of lifespans is well modeled by a Weibull distribution with `lam=2` and `k=1.5`. If we install `n=100` lightbulbs and come back one year later, what is the distribution of `c`, the number of lightbulbs that have burned out?
# The probability that any given bulb has burned out comes from the CDF of the distribution.
lam = 2
k = 1.5
p = EvalWeibullCdf(1, lam, k)
p
# The number of bulbs that have burned out is distributed Binom(n, p).
#
# +
from thinkbayes2 import MakeBinomialPmf
n = 100
pmf_c = MakeBinomialPmf(n, p)
thinkplot.Pdf(pmf_c)
# -
# Or we can approximate the distribution with a random sample.
n = 100
sample = np.random.binomial(n, p, 1000)
pdf_c = thinkbayes2.EstimatedPdf(sample)
thinkplot.Pdf(pdf_c)
np.mean(sample), np.std(sample)
# **Exercise:** Now suppose that `lam` and `k` are not known precisely, but we have a `LightBulb` object that represents the joint posterior distribution of the parameters after seeing some data. Compute the posterior predictive distribution for `c`, the number of bulbs burned out after one year.
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# -
|
seminar04.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from numpy import *
from numpy.random import *
import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.cm as cm
import matplotlib.gridspec as gridspec
from matplotlib.colors import ListedColormap
from matplotlib import colors
import matplotlib.ticker as mticker
from mpl_toolkits.axes_grid1 import make_axes_locatable
import matplotlib.cm as cm
from scipy.stats import norm
from PlotFuncs import col_alpha,CurvedText
from PlotFuncs import BlackHoleSpins, AxionNeutron, MySaveFig
# %matplotlib inline
# Force range for lambda in [m] and m_a in [eV]
def mLambda(m_a):
return 0.1973*1e-6/m_a
# Scalar-nucleon coupling from Yukawa alpha
def g_scalar_nucleon(alph):
return sqrt(alph/1.37e37)
def MakeJoinedLimit_ScalarNucleon(files,fname,header,nvals=1000):
dirc = 'limit_data/ScalarNucleon/'
n = len(files)
m_min = 100000.0
m_max = 1e-10
dats = zeros(shape=(n,nvals))
for file in files:
dat = loadtxt(dirc+file+'.txt')
dat[:,0] = mLambda(dat[:,0])
m_min = min(m_min,amin(dat[:,0]))
m_max = max(m_max,amax(dat[:,0]))
m = logspace(log10(m_min),log10(m_max),nvals)
for i in range(0,n):
dat = flipud(loadtxt(dirc+files[i]+'.txt'))
x = mLambda(dat[:,0])
y = g_scalar_nucleon(dat[:,1])
ynew = interp(m,x,y)
ynew[m<amin(x)] = 1e0
ynew[m>amax(x)] = 1e0
dats[i,:] = ynew
dat = amin(dats,0)
DAT = column_stack((m,dat))
savetxt(dirc+fname,DAT,header=header)
return DAT
# Making the Union of the ISL and WEP tests:
dirc = 'limit_data/ScalarNucleon/'
ISL = ['IUPUI','Stanford','EotWash2006','EotWash2020','HUST2012','HUST2020','Irvine','Wuhan']
EP = ['EotWash_EP_1999','MICROSCOPE','EotWash_EP_2007_left','EotWash_EP_2007_right']
AllLims = ISL+EP
header_ISL = 'Union of inverse square law tests \n m [eV] \t g_s_nucleon [dimensionless]'
header_EP = 'Union of equivalence principle tests \n m [eV] \t g_s_nucleon [dimensionless]'
header_All = 'Union of all tests \n m [eV] \t g_s_nucleon [dimensionless]'
fname_ISL = 'Union_InverseSquareLaw.txt'
fname_EP = 'Union_EquivalencePrinciple.txt'
DAT_ISL = MakeJoinedLimit_ScalarNucleon(ISL,fname_ISL,header_ISL)
DAT_EP = MakeJoinedLimit_ScalarNucleon(EP,fname_EP,header_EP)
DAT_EP = MakeJoinedLimit_ScalarNucleon(AllLims,'Union.txt',header_All)
# -
def FigSetup(xlab=r'$m_a$ [eV]',ylab='$|g_{an}|$',\
g_min = 1.0e-26,g_max = 1.0e-10,\
m_min = 1.0e-16,m_max = mLambda(1.0e-10),\
lw=2.5,lfs=40,tfs=25,tickdir='out',\
Grid=False,Shape='Rectangular',mathpazo=False,
TopAndRightTicks=False,LambdaAxis=True,UnitAxis=True):
plt.rcParams['axes.linewidth'] = lw
plt.rc('text', usetex=True)
plt.rc('font', family='serif',size=tfs)
if mathpazo:
mpl.rcParams['text.latex.preamble'] = [r'\usepackage{mathpazo}']
if Shape=='Wide':
fig = plt.figure(figsize=(16.5,5))
elif Shape=='Rectangular':
fig = plt.figure(figsize=(16.5,11))
ax = fig.add_subplot(111)
ax.set_xlabel(xlab,fontsize=lfs)
ax.set_ylabel(ylab,fontsize=lfs)
ax.tick_params(which='major',direction=tickdir,width=2.5,length=13,right=TopAndRightTicks,top=TopAndRightTicks,pad=7)
ax.tick_params(which='minor',direction=tickdir,width=1,length=10,right=TopAndRightTicks,top=TopAndRightTicks)
ax.set_yscale('log')
ax.set_xscale('log')
ax.set_xlim([m_min,m_max])
ax.set_ylim([g_min,g_max])
locmaj = mpl.ticker.LogLocator(base=10.0, subs=(1.0, ), numticks=50)
locmin = mpl.ticker.LogLocator(base=10.0, subs=arange(2, 10)*.1,numticks=100)
ax.xaxis.set_major_locator(locmaj)
ax.xaxis.set_minor_locator(locmin)
ax.xaxis.set_minor_formatter(mpl.ticker.NullFormatter())
locmaj = mpl.ticker.LogLocator(base=10.0, subs=(1.0, ), numticks=100)
locmin = mpl.ticker.LogLocator(base=10.0, subs=arange(2, 10)*.1,numticks=100)
ax.yaxis.set_major_locator(locmaj)
ax.yaxis.set_minor_locator(locmin)
ax.yaxis.set_minor_formatter(mpl.ticker.NullFormatter())
if Shape=='Rectangular':
plt.xticks(rotation=20)
if Grid:
ax.grid(zorder=0)
if LambdaAxis:
ax2 = ax.twiny()
ax2.set_xlim([mLambda(m_min),mLambda(m_max)])
ax2.set_xscale('log')
if UnitAxis:
ax2.set_xlabel(r"$\lambda$",fontsize=lfs)
xticks = [696340e3,6371e3,1e3,1,0.01,1e-6,1e-9,1e-10]
xticklabels = [r'$R_\odot$',r'$R_\oplus$','km','m','cm',r'\textmu m','nm',r'\AA']
ax2.set_xticks(xticks)
ax2.set_xticklabels(xticklabels)
else:
ax2.set_xlabel(r"$\lambda$ [m]",fontsize=lfs)
locmaj = mpl.ticker.LogLocator(base=10.0, subs=(1.0, ), numticks=50)
locmin = mpl.ticker.LogLocator(base=10.0, subs=arange(2, 10)*.1,numticks=100)
ax2.xaxis.set_major_locator(locmaj)
ax2.xaxis.set_minor_locator(locmin)
ax2.xaxis.set_minor_formatter(mpl.ticker.NullFormatter())
plt.xticks(rotation=20)
ax2.tick_params(which='major',direction=tickdir,width=2.5,length=13,pad=7)
ax2.tick_params(which='minor',direction=tickdir,width=1,length=10)
plt.sca(ax)
return fig,ax
# # Scalar nucleon coupling
# +
dirc = "limit_data/ScalarNucleon/"
g_scale=1
lw=3
RG_col='mediumseagreen'
ISL_col='crimson'
ISL_text_col='darkred'
ISL_line_col='firebrick'
EP_col='rebeccapurple'
EP_text_col='indigo'
EP_line_col='purple'
small_fs=20
med_fs=30
large_fs=35
lfs=45
fig,ax = FigSetup(ylab=r'Scalar nucleon coupling, $g_s^N$',\
Shape='Rectangular',mathpazo=True)
# Stellar cooling:
dat = loadtxt(dirc+'RedGiant.txt')
dat[:,1] = sqrt(dat[:,1])*1.83
plt.plot(dat[:,0],dat[:,1],'k-',zorder=10,lw=lw)
plt.fill_between(dat[:,0],dat[:,1],y2=1e0,color=RG_col,alpha=1,zorder=10)
plt.text(1e3,0.3e-11,r'{\bf Red Giants}',color='k',zorder=10,fontsize=large_fs,ha='right')
# Fill in equivalence principle and inverse square law
zo = 0.01
DAT = loadtxt(dirc+'Union_EquivalencePrinciple.txt')
plt.fill_between(DAT[:,0],g_scale*DAT[:,1],y2=1e0,color=EP_col,alpha=0.4,zorder=zo)
DAT = loadtxt(dirc+'Union_InverseSquareLaw.txt')
plt.fill_between(DAT[:,0],g_scale*DAT[:,1],y2=1e0,color=ISL_col,alpha=0.4,zorder=zo)
zo = 0.01
text_col = ISL_text_col
line_col = ISL_line_col
# IUPUI differential force measurement https://arxiv.org/pdf/1410.7267.pdf
dat = loadtxt(dirc+'IUPUI.txt')
dat[:,0] = mLambda(dat[:,0])
dat[:,1] = g_scalar_nucleon(dat[:,1])*g_scale
plt.plot(dat[:,0],dat[:,1],'k-',zorder=zo,color=line_col,lw=lw)
txt = CurvedText(flipud(dat[:-30,0])*1.5,flipud(dat[:-30,1])/4,color=line_col,text=r'IUPUI',va = 'bottom',axes = ax,fontsize=small_fs)
# Eot-Wash
dat1 = loadtxt(dirc+'EotWash2006.txt') # EotWash 2006 https://arxiv.org/pdf/hep-ph/0611184.pdf
dat1[:,0] = mLambda(dat1[:,0])
dat1[:,1] = g_scalar_nucleon(dat1[:,1])*g_scale
dat2 = loadtxt(dirc+'EotWash2020.txt') # EotWash 2020 https://arxiv.org/pdf/2002.11761.pdf
dat2[:,0] = mLambda(dat2[:,0])
dat2[:,1] = g_scalar_nucleon(dat2[:,1])*g_scale
dat1 = dat1[dat1[:,0]<1.6e-3]
dat2 = dat2[dat2[:,0]>1.6e-3]
dat = vstack((dat2,dat1))
plt.plot(dat[:,0],dat[:,1],'k-',zorder=zo,color=line_col,lw=lw)
txt = CurvedText(flipud(dat[:-70,0])*1.7,flipud(dat[:-70,1])/4,color=line_col,text=r'Eöt-Wash',va = 'bottom',axes = ax,fontsize=small_fs)
# Stanford https://arxiv.org/pdf/0802.2350.pdf
dat = loadtxt(dirc+'Stanford.txt')
dat[:,0] = mLambda(dat[:,0])
dat[:,1] = g_scalar_nucleon(dat[:,1])*g_scale
plt.plot(dat[:,0],dat[:,1],'k-',zorder=zo,color=line_col,lw=lw)
plt.text(2.7e-2,0.6e-15,r'Stanford',rotation=77,rotation_mode='anchor',ha='center',color=line_col,fontsize=small_fs)
# HUST
dat1 = loadtxt(dirc+'HUST2012.txt') # HUST 2012 https://journals.aps.org/prl/abstract/10.1103/PhysRevLett.108.081101
dat1[:,0] = mLambda(dat1[:,0])
dat1[:,1] = g_scalar_nucleon(dat1[:,1])*g_scale
dat2 = loadtxt(dirc+'HUST2020.txt') # HUST 2020 https://journals.aps.org/prl/abstract/10.1103/PhysRevLett.124.051301
dat2[:,0] = mLambda(dat2[:,0])
dat2[:,1] = g_scalar_nucleon(dat2[:,1])*g_scale
dat1 = dat1[dat1[:,0]<3.7e-4]
dat2 = dat2[dat2[:,0]>3.7e-4]
dat = vstack((dat2,dat1))
plt.plot(dat[:,0],dat[:,1],'k-',zorder=zo,color=line_col,lw=lw)
txt = CurvedText(flipud(dat[:,0])*1.25,flipud(dat[:,1])/4,color=line_col,text=r'HUST',va = 'bottom',axes = ax,fontsize=small_fs)
# Irvine 10.1103/PhysRevD.32.3084
# dat = loadtxt(dirc+'Irvine.txt')
# dat[:,0] = mLambda(dat[:,0])
# dat[:,1] = g_scalar_nucleon(dat[:,1])*g_scale
# plt.plot(dat[:,0],dat[:,1],'k--',zorder=zo,color=line_col,lw=lw)
# plt.text(0.9e-7,0.7e-19,r'Irvine',rotation=-25,rotation_mode='anchor',ha='center',color=line_col,fontsize=small_fs)
# Wuhan AAF https://journals.aps.org/prl/pdf/10.1103/PhysRevLett.126.211101
dat = loadtxt(dirc+'Wuhan.txt')
dat[:,0] = mLambda(dat[:,0])
dat[:,1] = g_scalar_nucleon(dat[:,1])*g_scale
plt.plot(dat[:,0],dat[:,1],'-',lw=lw,color=line_col,zorder=100)
plt.text(4e-7,0.3e-20,r'Wuhan',rotation=-27,rotation_mode='anchor',ha='center',color=line_col,fontsize=small_fs)
text_col = EP_text_col
line_col = EP_line_col
# EotWash EP 10.1103/PhysRevD.61.022001
dat = loadtxt(dirc+'EotWash_EP_1999.txt')
dat[:,0] = mLambda(dat[:,0])
dat[:,1] = g_scalar_nucleon(dat[:,1])*g_scale
plt.plot(dat[:,0],dat[:,1],'k-',zorder=zo,color=line_col,lw=lw)
# EotWash EP test 2007 10.1103/PhysRevD.61.022001
dat1 = flipud(loadtxt(dirc+'EotWash_EP_2007_right.txt'))
dat1[:,0] = mLambda(dat1[:,0])
dat1[:,1] = g_scalar_nucleon(dat1[:,1])*g_scale
dat2 = flipud(loadtxt(dirc+'EotWash_EP_2007_left.txt'))
dat2[:,0] = mLambda(dat2[:,0])
dat2[:,1] = g_scalar_nucleon(dat2[:,1])*g_scale
dat_join = vstack((dat1[-1,:],dat2[0,:]))
dat_joined = vstack((dat1,dat2))
plt.plot(dat1[:,0],dat1[:,1],'k-',zorder=zo,color=line_col,lw=lw)
plt.plot(dat2[:,0],dat2[:,1],'k-',zorder=zo,color=line_col,lw=lw)
plt.plot(dat_join[:,0],dat_join[:,1],'k--',zorder=zo,color=line_col,lw=lw)
plt.text(9.5e-9,2e-23,r'E\"ot-Wash',rotation=18,rotation_mode='anchor',\
color=line_col,va = 'bottom',fontsize=small_fs+5)
# MICROSCOPE satellite https://arxiv.org/pdf/1712.00483.pdf
dat = loadtxt(dirc+'MICROSCOPE.txt')
dat[:,0] = mLambda(dat[:,0])
dat[:,1] = g_scalar_nucleon(dat[:,1])*g_scale
plt.plot(dat[:,0],dat[:,1],'k-',zorder=zo,color=line_col,lw=lw)
plt.text(1.5e-16,2.5e-25,'MICROSCOPE',color=line_col,fontsize=small_fs)
# Labels
plt.text(7e-3,0.7e-13,r'{\bf Inverse square}',color=ISL_text_col,ha='center',fontsize=large_fs)
plt.text(2e-3,0.7e-13/9,r'{\bf law tests}',color=ISL_text_col,ha='center',fontsize=large_fs)
plt.text(4e-12,1e-15,r'{\bf Equivalence}',color=EP_text_col,ha='center',fontsize=large_fs)
plt.text(4e-12,1e-15/9,r'{\bf principle tests}',color=EP_text_col,ha='center',fontsize=large_fs)
plt.gcf().text(0.89,0.15,r'{\bf Monopole-monopole}',fontsize=med_fs+10,ha='right')
ax3 = ax.twinx()
g_min = ax.get_ylim()[0]
g_max = ax.get_ylim()[1]
ax3.set_ylim([1.37e37*g_min**2,1.37e37*g_max**2])
ax3.set_yscale('log')
ax3.set_ylabel(r"$|\alpha|$",fontsize=lfs,rotation=0,labelpad=20)
ax3.tick_params(which='major',direction='out',width=2.5,length=13,pad=7)
ax3.tick_params(which='minor',direction='out',width=1,length=10)
plt.sca(ax)
#BlackHoleSpins(ax,label_position=[1e-12,3e-26],rotation=0,fs=20,PlotLine=False)
# QCD Axion band:
m_n = 0.93957
m_vals = array([1e-16,1e4])
g_QCD_lower = 1e-29*(m_vals/5.7e-3)
g_QCD_upper = 2e-21*(m_vals/5.7e-3)
for i in logspace(0,3,30):
ax.fill_between(m_vals,g_QCD_lower*i,y2=g_QCD_upper/i,color='orange',\
alpha=0.01,zorder=-100,lw=3)
ax.plot(m_vals,g_QCD_lower,'-',color='orange',alpha=1,lw=2)
ax.plot(m_vals,g_QCD_upper,'-',color='orange',alpha=1,lw=2)
ax.text(4e-2,0.89e-23,r'{\bf QCD axion}',fontsize=35,color='darkorange',rotation=38)
ax.text(3e-1,(0.89e-23)/2,r'(Yukawa)',fontsize=30,color='darkorange',rotation=38)
MySaveFig(fig,'ScalarNucleon')
#==============================================================================#
# +
fig,ax = FigSetup(ylab=r'Electron-nucleon coupling, $g_s^N g_p^e$',Shape='Rectangular',mathpazo=True,\
g_min=1e-38,g_max=5e-24)
lw = 3
small_fs = 20
med_fs = 25
#==============================================================================#
col = 'mediumseagreen'
line_col = 'darkgreen'
zo = 0
g_p_e_lim = loadtxt('limit_data/AxionElectron/RedGiants.txt')[0,1]
g_s_N_lim = sqrt(loadtxt('limit_data/ScalarNucleon/RedGiant.txt')[0,-1]*4*pi)
DAT = loadtxt('limit_data/ScalarNucleon/Union.txt')
plt.fill_between(DAT[:,0],g_p_e_lim*DAT[:,1],y2=1e0,color=col,alpha=0.1,zorder=zo)
plt.plot(DAT[:,0],g_p_e_lim*DAT[:,1],'--',color=line_col,lw=3)
plt.text(1e-7,0.4e-35,r'${\bf(Lab)}_s^N\times {\bf(Astro)}_p^e$',\
rotation=15,rotation_mode='anchor',ha='center',fontsize=30,color=line_col)
glim = g_p_e_lim*g_s_N_lim
plt.fill_between([1e-20,1e4],[glim,glim],y2=1e0,color=col,alpha=1,zorder=100)
plt.plot([1e-20,1e4],[glim,glim],'-',color=line_col,lw=3,zorder=100)
plt.text(1e3,0.3e-24,r'{\bf Red giants} $\mathbf{g}_\mathbf{s}^\mathbf{N} \times \mathbf{g}_\mathbf{p}^\mathbf{e}$',\
fontsize=35,color='k',zorder=101,ha='right')
#==============================================================================#
#==============================================================================#
dirc = 'limit_data/MonopoleDipole/ElectronNucleon/'
text_col = 'navy'
line_col = 'navy'
col = 'royalblue'
zo = 1
# QUAX limit https://arxiv.org/pdf/1705.06044.pdf
dat = loadtxt(dirc+'QUAX.txt')
dat[:,0] = mLambda(dat[:,0])
plt.plot(dat[:,0],dat[:,1],'-',lw=lw,color=line_col,zorder=zo)
plt.fill_between(dat[:,0],dat[:,1],y2=1,lw=lw,color=col,alpha=0.5,zorder=zo)
plt.text(2e-16,0.6e-29,'QUAX-$g_p g_s$',color=line_col,fontsize=med_fs)
# QUAX projection https://arxiv.org/pdf/1606.04751.pdf
dat = flipud(loadtxt(dirc+'QUAX_RLC.txt'))
dat[:,0] = mLambda(dat[:,0])
plt.plot(dat[:,0],dat[:,1],':',lw=lw,color=line_col)
plt.text(0.06e-4,1e-32,'QUAX-$g_p g_s$ (Proj.)',color=line_col,fontsize=small_fs,rotation=80)
# Torsio/Magnetometer
text_col = 'purple'
line_col = 'purple'
col = 'rebeccapurple'
zo = 1
# Washington (Hoedl) 10.1103/PhysRevLett.106.041801
dat = loadtxt(dirc+'Washington_Hoedl.txt')
plt.plot(dat[:,0],dat[:,1],'-',lw=lw,color=line_col,zorder=zo)
# Washington (Terrano) 10.1103/PhysRevLett.115.201801
dat = loadtxt(dirc+'Washington_Terrano.txt')
plt.plot(dat[:,0],dat[:,1],'-',lw=lw,color=line_col,zorder=zo)
plt.text(1.8e-4,0.5e-27,'Washington',color=line_col,fontsize=small_fs,rotation=71)
# # Amherst 10.1103/PhysRevLett.77.2170
# dat = loadtxt(dirc+'Amherst.txt')
# plt.plot(dat[:,0],dat[:,1],'-',lw=lw,color=line_col,zorder=zo)
# plt.fill_between(dat[:,0],dat[:,1],y2=1,lw=lw,color=col,alpha=0.5,zorder=zo)
# plt.text(0.8e-11,0.15e-28,'Amherst',color=line_col,fontsize=med_fs)
# EotWash https://arxiv.org/pdf/0808.2673.pdf
dat1 = flipud(loadtxt(dirc+'EotWash_left.txt'))
dat1[:,0] = mLambda(dat1[:,0])
plt.plot(dat1[:,0],dat1[:,1],'-',lw=lw,color=line_col,zorder=zo)
dat2 = flipud(loadtxt(dirc+'EotWash_right.txt'))
dat2[:,0] = mLambda(dat2[:,0])
plt.plot(dat2[:,0],dat2[:,1],'-',lw=lw,color=line_col,zorder=zo)
plt.plot([dat1[0,0],dat2[-1,0]],[dat1[0,1],dat2[-1,1]],'k--',lw=lw,color=line_col)
plt.text(6e-9,8e-33,r'E\"ot-Wash',color=line_col,\
fontsize=small_fs,rotation=32,rotation_mode='anchor',ha='center')
# NIST (Wineland) 10.1103/PhysRevLett.67.1735
dat = loadtxt(dirc+'NIST.txt')
dat[:,0] = mLambda(dat[:,0])
plt.plot(dat[:,0],dat[:,1],'-',lw=lw,color=line_col,zorder=zo)
plt.text(1e-12,1.5e-32,'NIST',color=line_col,fontsize=small_fs,rotation=43)
# SMILE https://arxiv.org/pdf/1801.02757.pdf
dat = loadtxt(dirc+'SMILE.txt')
dat[:,0] = mLambda(dat[:,0])
plt.plot(dat[:,0],dat[:,1],'-',lw=lw,color=line_col,zorder=zo)
plt.text(1e-9,0.5e-30,'SMILE',color=line_col,fontsize=small_fs,rotation=0)
# XENON1T S2
Xenon = loadtxt('limit_data/AxionElectron/XENON1T_DM_S2.txt')
plt.fill_between(Xenon[:,0],Xenon[:,1]*g_s_N_lim,y2=1e1,lw=lw,color='firebrick',zorder=-99)
plt.plot(Xenon[:,0],Xenon[:,1]*g_s_N_lim,'k-',zorder=-99)
plt.text(2e2,0.09e-25,r'Xenon1T',color='firebrick',fontsize=small_fs+2,rotation=0,ha='center')
plt.text(2e2/2,0.09e-25/5,r'${\rm(Lab)}_s^N$$\times$${\rm(DM)}_p^e$',color='firebrick',fontsize=small_fs-3,rotation=0,ha='center')
# Magnon
DAT = loadtxt('limit_data/ScalarNucleon/Union.txt')
Casper = loadtxt('limit_data/AxionElectron/Projections/Magnon.txt')
m = Casper[:,0]
Casper = interp(DAT[:,0],Casper[:,0],Casper[:,1])
Casper[DAT[:,0]>amax(m)] = nan
Casper[DAT[:,0]<amin(m)] = nan
y1 = Casper*DAT[:,1]
plt.plot(DAT[:,0],y1,':',lw=lw,color='crimson',zorder=zo)
plt.text(0.13e-1,5e-27,'Magnons',color='crimson',fontsize=small_fs,rotation=65,ha='center',rotation_mode='anchor')
plt.text(0.13e-1*4.5,5e-27/2,r'${\rm(Lab)}_s^N$$\times$${\rm(DM)}_p^e$',color='crimson',fontsize=small_fs-3,rotation=65,ha='center',rotation_mode='anchor')
# Union
DAT = loadtxt(dirc+'Union_TorsionMagnetometer.txt')
plt.fill_between(DAT[:,0],DAT[:,1],y2=1,lw=lw,color=col,alpha=0.3,zorder=zo)
plt.gcf().text(0.89,0.15,r'{\bf Monopole-dipole}',fontsize=med_fs+10,ha='right')
# QCD axion
m_e = 511.0/1e6
m_vals = array([1e-16,1e4])
g_QCD_lower = 1e-29*1e-9*0.024*m_e*(m_vals/5.7e-3)**2
g_QCD_upper = 2e-21*1e-9*1/3*m_e*(m_vals/5.7e-3)**2
for i in logspace(0,3,30):
ax.fill_between(m_vals,g_QCD_lower*i,y2=g_QCD_upper/i,color='orange',alpha=0.01,zorder=-100,lw=3)
ax.plot(m_vals,g_QCD_lower,color='orange',lw=2)
ax.plot(m_vals,g_QCD_upper,color='orange',lw=2)
ax.text(4e-1,0.9e-34,r'{\bf QCD axion}',fontsize=30,color='darkorange',rotation=58)
#BlackHoleSpins(ax,label_position=[1e-12,3e-38],rotation=0,fs=20,PlotLine=False)
MySaveFig(fig,'MonopoleDipole_ElectronNucleon')
# +
dirc = 'limit_data/MonopoleDipole/ElectronNucleon/'
# Make Union of nucleon electron limits
#==============================================================================#
dat1 = loadtxt(dirc+'Washington_Hoedl.txt')
dat2 = loadtxt(dirc+'Washington_Terrano.txt')
dat3 = flipud(loadtxt(dirc+'EotWash_left.txt'))
dat3[:,0] = mLambda(dat3[:,0])
dat4 = flipud(loadtxt(dirc+'EotWash_right.txt'))
dat4[:,0] = mLambda(dat4[:,0])
dat5 = flipud(loadtxt(dirc+'NIST.txt'))
dat5[:,0] = mLambda(dat5[:,0])
dat6 = flipud(loadtxt(dirc+'SMILE.txt'))
dat6[:,0] = mLambda(dat6[:,0])
m_min = amin(array([dat1[0,0],dat2[0,0],dat3[0,0],dat4[0,0],dat5[0,0],dat6[0,0]]))
m_max = amax(array([dat1[-1,0],dat2[-1,0],dat3[-1,0],dat4[-1,0],dat5[-1,0],dat6[-1,0]]))
nvals = 1000
files = [dat1,dat2,dat3,dat4,dat5,dat6]
m = logspace(log10(m_min),log10(m_max),nvals)
dats = zeros(shape=(6,nvals))
for i in range(0,6):
dat = files[i]
x = dat[:,0]
y = dat[:,1]
ynew = interp(m,x,y)
ynew[m<amin(x)] = 1e0
ynew[m>amax(x)] = 1e0
dats[i,:] = ynew
dat = amin(dats,0)
DAT = column_stack((m,dat))
dirc = 'limit_data/MonopoleDipole/ElectronNucleon/'
header = 'Union of torsion and magnetometer limits \n m [eV] \t g_s*g_p [dimensionless]'
fname = 'Union_TorsionMagnetometer.txt'
savetxt(dirc+fname,DAT,header=header)
#==============================================================================#
#==============================================================================#
DAT = loadtxt('limit_data/ScalarNucleon/Union.txt')
g_p_e_lim = loadtxt('limit_data/AxionElectron/RedGiants.txt')[0,1]
savetxt(dirc+'UnionAstroLab.txt',column_stack((DAT[:,0],g_p_e_lim*DAT[:,1])),\
header='Union of astro (g_p^e) x lab (g_s^N) limits \n m [eV] \t g^N_s*g^e_p [dimensionless]')
#==============================================================================#
# +
fig,ax = FigSetup(ylab=r'Nucleon-nucleon coupling, $g_s^N g_p^N$',Shape='Rectangular',mathpazo=True,\
g_min=1e-39,g_max=1e-20)
#==============================================================================#
lw = 3
small_fs = 20
med_fs = 25
col = 'mediumseagreen'
line_col = 'darkgreen'
zo = 0
g_p_N_lim = sqrt(7.7e-20) # Hot neutron star
g_s_N_lim = sqrt(loadtxt('limit_data/ScalarNucleon/RedGiant.txt')[0,-1]*4*pi)
DAT = loadtxt('limit_data/ScalarNucleon/Union.txt')
plt.fill_between(DAT[:,0],g_p_N_lim*DAT[:,1],y2=1e0,color=col,alpha=0.1,zorder=zo)
plt.plot(DAT[:,0],g_p_N_lim*DAT[:,1],'--',color=line_col,lw=3)
plt.text(1e-6,0.7e-32,r'${\bf(Lab)}_s^N\times {\bf(Astro)}_p^N$',rotation=15,rotation_mode='anchor',ha='center',fontsize=30,color=line_col)
glim = g_p_N_lim*g_s_N_lim
plt.fill_between([1e-20,1e4],[glim,glim],y2=1e0,color=col,alpha=1,zorder=100)
plt.plot([1e-20,1e4],[glim,glim],'-',color='k',lw=3,zorder=100)
plt.text(1e3,0.42e-21,r'{\bf Red giants $\mathbf{g}^\mathbf{N}_\mathbf{s}\times$ Neutron stars $\mathbf{g}_\mathbf{p}^\mathbf{N}$}',fontsize=32,color='k',ha='right',zorder=101)
#==============================================================================#
#==============================================================================#
# ARIADNE
dirc = 'limit_data/MonopoleDipole/NucleonNucleon/'
text_col = 'navy'
line_col = 'navy'
col = 'royalblue'
zo = 1
# ARIADNE https://arxiv.org/pdf/1403.1290.pdf
dat = loadtxt(dirc+'ARIADNE_projection1.txt')
dat[:,0] = mLambda(dat[:,0])
plt.plot(dat[:,0],dat[:,1],':',lw=lw,color=line_col,zorder=zo)
dat = loadtxt(dirc+'ARIADNE_projection2.txt')
dat[:,0] = mLambda(dat[:,0])
plt.plot(dat[:,0],dat[:,1],':',lw=lw,color=line_col,zorder=zo)
plt.text(20e-7,0.2e-34,'ARIADNE',color=line_col,fontsize=small_fs,ha='center')
plt.text(20e-7,0.2e-34/5,'(Projected)',color=line_col,fontsize=small_fs-2,ha='center')
plt.text(7.5e-7,0.15e-38,'ARIADNE (Upgrade)',color=line_col,fontsize=small_fs,rotation=48)
#==============================================================================#
#==============================================================================#
# Expts.
text_col = 'purple'
line_col = 'purple'
col = 'rebeccapurple'
zo = 1
# # Grenoble https://arxiv.org/pdf/1009.3434.pdf
# dat = loadtxt(dirc+'Grenoble.txt')
# dat[:,0] = mLambda(dat[:,0])
# plt.plot(dat[:,0],dat[:,1],'-',lw=lw,color=line_col,zorder=zo)
# plt.text(0.4e-4,0.3e-22,r'Grenoble',fontsize=small_fs,color=line_col,rotation=0)
# Amherst 10.1103/PhysRevLett.77.2170
# dat = loadtxt(dirc+'Amherst.txt')
# dat[:,1] *= 2 # to get 95%
# dat[:,0] = mLambda(dat[:,0])
# plt.plot(dat[:,0],dat[:,1],'-',lw=lw,color=line_col,zorder=zo)
# plt.text(0.3e-5,0.6e-25,r'Amherst',fontsize=small_fs,color=line_col,rotation=80)
# SMILE https://arxiv.org/pdf/1801.02757.pdf
dat = loadtxt(dirc+'SMILE.txt')
dat[:,0] = mLambda(dat[:,0])
plt.plot(dat[:,0],dat[:,1],'-',lw=lw,color=line_col,zorder=zo)
plt.text(3e-6,1e-27,'SMILE',color=line_col,fontsize=small_fs,rotation=78)
# Mainz 10.1103/PhysRevLett.111.100801
dat = loadtxt(dirc+'Mainz.txt')
dat[:,0] = mLambda(dat[:,0])
plt.plot(dat[:,0],dat[:,1],'-',lw=lw,color=line_col,zorder=zo)
plt.text(0.7e-4,0.7e-25,'Mainz',color=line_col,fontsize=small_fs,rotation=73)
# Washington 10.1103/PhysRevLett.68.135
dat = loadtxt(dirc+'Washington.txt')
dat[:,0] = mLambda(dat[:,0])
plt.plot(dat[:,0],dat[:,1],'-',lw=lw,color=line_col,zorder=zo)
plt.text(0.7e-9,0.5e-29,'Washington',color=line_col,fontsize=small_fs,rotation=38)
# union
DAT = loadtxt(dirc+'Union.txt')
plt.fill_between(DAT[:,0],DAT[:,1],y2=1,lw=lw,color=col,alpha=0.3,zorder=zo)
# CASPER
DAT = loadtxt('limit_data/ScalarNucleon/Union.txt')
Casper = loadtxt('limit_data/AxionNeutron/Projections/CASPEr_wind.txt')
Casper[:,1] *= 2*m_n
m = Casper[:,0]
Casper = interp(DAT[:,0],Casper[:,0],Casper[:,1])
Casper[DAT[:,0]>amax(m)] = nan
Casper[DAT[:,0]<amin(m)] = nan
plt.plot(DAT[:,0],Casper*DAT[:,1],':',lw=lw,color='crimson',zorder=zo)
plt.text(0.1e-9,0.3e-36,r'CASPEr-wind',color='crimson',fontsize=small_fs+2,rotation=28)
plt.text(0.1e-9*2,0.3e-36/6,r'${\rm(Lab)}_s^N$$\times$${\rm(DM)}_p^N$',color='crimson',fontsize=small_fs-2,rotation=28)
# Future comag
FC = loadtxt('limit_data/AxionNeutron/Projections/FutureComagnetometers.txt')
FC[:,1] *= 2*m_n
m = FC[:,0]
FC = interp(DAT[:,0],FC[:,0],FC[:,1])
FC[DAT[:,0]>amax(m)] = nan
FC[DAT[:,0]<amin(m)] = nan
imax = 395
plt.plot(DAT[0:imax,0],FC[0:imax]*DAT[0:imax,1],':',lw=lw,color='rebeccapurple',zorder=zo)
plt.text(1.5e-16,2e-36,'DM comag.',color='rebeccapurple',fontsize=small_fs,multialignment='center')
plt.text(1.5e-16,2e-36/15,r'${\rm(Lab)}_s^N$$\times$${\rm(DM)}_p^N$',color='rebeccapurple',fontsize=small_fs-2,rotation=0)
# QCD axion
m_n = 0.93957
m_vals = array([1e-16,1e4])
g_QCD_lower = 1e-29*1e-9*0.16*m_n*(m_vals/5.7e-3)**2
g_QCD_upper = 2e-21*1e-9*0.6*m_n*(m_vals/5.7e-3)**2
for i in logspace(0,3,30):
ax.fill_between(m_vals,g_QCD_lower*i,y2=g_QCD_upper/i,color='orange',alpha=0.01,zorder=-100,lw=3)
ax.plot(m_vals,g_QCD_lower,color='orange',lw=2)
ax.plot(m_vals,g_QCD_upper,color='orange',lw=2)
ax.text(4e-2,1e-32,r'{\bf QCD axion}',fontsize=30,color='darkorange',rotation=52)
plt.gcf().text(0.89,0.15,r'{\bf Monopole-dipole}',fontsize=med_fs+10,ha='right')
#BlackHoleSpins(ax,label_position=[1e-12,3e-39],rotation=0,fs=20,PlotLine=False)
MySaveFig(fig,'MonopoleDipole_NucleonNucleon')
# +
# Make union of nucleon-nucleon limits
#==============================================================================#
nvals = 1000
dat1 = flipud(loadtxt(dirc+'Washington.txt'))
dat1[:,0] = mLambda(dat1[:,0])
dat2 = flipud(loadtxt(dirc+'Amherst.txt'))
dat2[:,1] *= 2 # to get 95%
dat2[:,0] = mLambda(dat2[:,0])
dat3 = flipud(loadtxt(dirc+'SMILE.txt'))
dat3[:,0] = mLambda(dat3[:,0])
dat4 = flipud(loadtxt(dirc+'Mainz.txt'))
dat4[:,0] = mLambda(dat4[:,0])
m_min = amin(array([dat1[0,0],dat2[0,0],dat3[0,0],dat4[0,0]]))
m_max = amax(array([dat1[-1,0],dat2[-1,0],dat3[-1,0],dat4[-1,0]]))
files = [dat1,dat2,dat3,dat4]
nf = len(files)
m = logspace(log10(m_min),log10(m_max),nvals)
dats = zeros(shape=(nf,nvals))
for i in range(0,nf):
dat = files[i]
x = dat[:,0]
y = dat[:,1]
ynew = interp(m,x,y)
ynew[m<amin(x)] = 1e0
ynew[m>amax(x)] = 1e0
dats[i,:] = ynew
dat = amin(dats,0)
DAT = column_stack((m,dat))
dirc = 'limit_data/MonopoleDipole/NucleonNucleon/'
header = 'Union of nucleon-nucleon limits \n m [eV] \t g_s*g_p [dimensionless]'
fname = 'Union.txt'
savetxt(dirc+fname,DAT,header=header)
#==============================================================================#
#==============================================================================#
DAT = loadtxt('limit_data/ScalarNucleon/Union.txt')
g_p_N_lim = sqrt(7.7e-20) # Hot neutron star
savetxt(dirc+'UnionAstroLab.txt',column_stack((DAT[:,0],g_p_N_lim*DAT[:,1])),\
header='Union of astro (g_p^N) x lab (g_s^N) limits \n m [eV] \t g^N_s*g^N_p [dimensionless]')
#==============================================================================#
# +
#==============================================================================#
# Improvement plot
from PlotFuncs import MySquarePlot
fig,ax1 = MySquarePlot('$m_a$ [eV]','Improvement since 2012',size_y=8,lfs=35)
nm = 10000
m_min = 1.1e-15
m_max = 1e3
m = logspace(log10(m_min),log10(m_max),nm)
line_col = 'teal'
alph = 0.3
# Scalar
g_S_N_2012 = 0.5e-10
g_S_N_2020 = 1.1e-12
g_QCD_upper = 2e-21*(m/5.7e-3)
Raff = flipud(loadtxt('limit_data/ScalarNucleon/Raffelt.txt'))
m1 = mLambda(Raff[:,0])
y1 = interp(m,m1,Raff[:,1])
y1[y1==0] = g_S_N_2012
y1[y1>g_S_N_2012] =g_S_N_2012
My = loadtxt('limit_data/ScalarNucleon/Union.txt')
y2 = interp(m,My[:,0],My[:,1])
y2[y2==0] = g_S_N_2020
y2[y2>g_S_N_2020] =g_S_N_2020
mask = y1<y2
y1[mask] = y2[mask]
max1 = amax(y1/y2)
ax1.plot(m,y1/y2,'r-',lw=3,color=line_col,label=r'$g_s^N$')
ax1.plot(m,y1/g_QCD_upper,lw=3,color='orange')
ax1.fill_between(m,y1/g_QCD_upper,y2=1e10,color='orange',alpha=alph)
# Electron
g_p_e_2012 = 3e-13
g_p_e_2020 = 1.6e-13
g_QCD_upper = 2e-21*1e-9*1/3*m_e*(m/5.7e-3)**2
y1 = interp(m,m1,Raff[:,1]*g_p_e_2012)
y1[y1==0] = g_S_N_2012*g_p_e_2012
y1[y1>g_S_N_2012*g_p_e_2012] =g_S_N_2012*g_p_e_2012
y2 = interp(m,My[:,0],My[:,1]*g_p_e_2020)
y2[y2==0] = g_S_N_2020*g_p_e_2020
y2[y2>g_S_N_2020*g_p_e_2020] =g_S_N_2020*g_p_e_2020
mask = y1<y2
y1[mask] = y2[mask]
max2 = amax(y1/y2)
ax1.plot(m,y1/y2,'r--',lw=3,color=line_col,label=r'$g_s^N g_p^e$')
ax1.plot(m,y1/g_QCD_upper,'--',lw=3,color='orange')
ax1.fill_between(m,y1/g_QCD_upper,y2=1e10,color='orange',alpha=alph)
# Electron
g_p_N_2012 = 9.65e-10
g_p_N_2020 = 2.8e-10
g_QCD_upper = 2e-21*1e-9*0.26*m_n*(m/5.7e-3)**2
y1 = interp(m,m1,Raff[:,1]*g_p_N_2012)
y1[y1==0] = g_S_N_2012*g_p_N_2012
y1[y1>g_S_N_2012*g_p_N_2012] =g_S_N_2012*g_p_N_2012
y2 = interp(m,My[:,0],My[:,1]*g_p_N_2020)
y2[y2==0] = g_S_N_2020*g_p_N_2020
y2[y2>g_S_N_2020*g_p_N_2020] =g_S_N_2020*g_p_N_2020
mask = y1<y2
y1[mask] = y2[mask]
dat = flipud(loadtxt('limit_data/MonopoleDipole/NucleonNucleon/ARIADNE_projection2.txt'))
dat[:,0] = mLambda(dat[:,0])
y3 = interp(m,dat[:,0],dat[:,1])
y3[m>dat[-1,0]]=0
y3[m<dat[0,0]]=0
ax1.plot(m,y1/y3,'-.',color='navy',lw=lw)
dat = flipud(loadtxt('limit_data/MonopoleDipole/NucleonNucleon/ARIADNE_projection1.txt'))
dat[:,0] = mLambda(dat[:,0])
y3 = interp(m,dat[:,0],dat[:,1])
y3[m>dat[-1,0]]=0
y3[m<dat[0,0]]=0
ax1.plot(m,y1/y3,'-.',color='navy',lw=lw)
max3 = amax(y1/y2)
ax1.plot(m,y1/y2,'r-.',lw=3,color=line_col,label=r'$g_s^N g_p^N$')
ax1.plot(m,y1/g_QCD_upper,'-.',lw=3,color='orange')
ax1.fill_between(m,y1/g_QCD_upper,y2=1e10,color='orange',alpha=alph,label='QCD axion')
#plt.grid()
ax1.set_xscale('log')
ax1.set_yscale('log')
ax1.set_xlim([m_min,m_max])
ax1.set_ylim([1,1e6])
leg = plt.legend(fontsize=30,frameon=False,loc='upper left',edgecolor='k',labelspacing=0.1)
ax1.tick_params(pad=10)
plt.text(1e-3,1.3e4,r'ARIADNE',color='navy',fontsize=20)
#leg.get_frame().set_linewidth(2.5)
print(max1)
print(max2)
print(max3)
MySaveFig(fig,'Improvement')
# -
# QCD band of g_s^n
m_n = 0.93957
m_u = 2.3
m_d = 4.8
sig = 38
print('g_S^N (upper) = ',1.2e-10*sig*m_u*m_d/(m_n*1000*(m_u+m_d)**2)*(m_n/1e9))
print('g_S^N (lower) = ',1e-18*sig*m_u*m_d/(m_n*1000*(m_u+m_d)**2)*(m_n/1e9))
sig*m_u*m_d/(m_n*1000*(m_u+m_d)**2)*(m_n/1e9)
|
AxionCPV.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [](https://colab.research.google.com/github/MarcCote/TextWorld/blob/msr_summit_2021/notebooks/Handcrafting%20a%20game.ipynb)
# # Handcrafting a text-based game with TextWorld
# This tutorial shows how to make a custom text-based game using the TextWorld's API.
# ## Prerequisite
# Install TextWorld as described in the [README.md](https://github.com/microsoft/TextWorld#readme). Most of the time, a simple `pip install` should work.
#
# <span style="color:red">_*Notice here that TextWorld is installed with the `[vis]` extras to enable visual rendering of the world._</span>
# !pip install textworld[vis]
# ## `GameMaker`
# Let's make a game consisting of two rooms and where the goal is to place an object from the player's inventory inside a container located in the other room.
# +
import textworld
from textworld import GameMaker
# Make the generation process reproducible.
from textworld import g_rng # Global random generator.
g_rng.set_seed(20180916)
# GameMaker object for handcrafting text-based games.
M = GameMaker()
# -
# Documentation about the `GameMaker` API can be found at
# https://textworld.readthedocs.io/en/stable/textworld.generator.maker.html.
# #### First, define the layout of the world.
roomA = M.new_room("Room A")
roomB = M.new_room("Room B")
corridor = M.connect(roomA.east, roomB.west)
M.render()
# #### Place the player somewhere.
M.set_player(roomA)
M.render()
# #### Add two objects to Room A.
# +
note = M.new(type='o', name="note") # Provide the type and the name of the object.
note.infos.desc = "Bravo! You can read." # Text to display when issuing command "examine note".
supporter = M.new(type='s') # When not provided, names are automatically generated.
roomA.add(supporter) # Supporters are fixed in place.
roomA.add(note) # When added directly to a room, portable objects are put on the floor.
M.render()
# -
# ##### Supported types of objects
#
# type | name | description
# --------|-----|------------
# 'r' | room | can contain objects, can be connected with other rooms
# 'd' | door | (un)lockable, openable/closable
# 'c' | container | can hold objects, (un)lockable, openable/closable
# 's' | supporter | can hold objects
# 'o' | portable object | can be carried by the player
# 'k' | key | subtype of portable object, match a door or container's lock
# 'f' | food | subtype of portable object, can be eaten if edible
# 'oven' | oven | subtype of container, provide a heat source to cook food item
# 'stove' | stove | subtype of supporter, provide a heat source to cook food item
# #### Let's add a locked door between the two rooms.
door = M.new_door(corridor, name="door") # Door and room have their own "new" method, no need to provide the type.
M.add_fact("locked", door) # Add a fact about the door, i.e. it is locked.
M.render()
# ##### Supported types of facts
#
# fact/predicate | description
# ----------------|------------
# __match__(key, container/door) | The `key` matches the `container/door`'s lock.
# __open__(container/door) | The `container/door` is open.
# __closed__(container/door) | The `container/door` is closed.
# __locked__(container/door) | The `container/door` is locked.
# __edible__(food) | The `food` is edible, otherwise needs to be cooked first.
# #### Create a key object that be used to unlock the door and place on the supporter object.
# +
key = M.new(type="k", name="old key") # Create a 'k' (i.e. key) object.
M.add_fact("match", key, door) # Tell the game 'old key' is matching the 'door''s lock
supporter.add(key) # Add the 'old key' on the supporter.
M.render()
# -
# #### Create the object carried by the player and the container (initially closed).
# +
obj = M.new(type='o') # New portable object with a randomly generated name.
M.inventory.add(obj) # Add the object to the player's inventory.
container = M.new(type='c') # Creating a container with a randomly generated name
M.add_fact("closed", container) # Set the state of the container to 'closed'.
roomB.add(container) # Place the container in Room B
M.render()
# -
# #### Finally, record the quest.
# Type all commands you would like to be included in the game's quest. To end recording, interrupt the kernel (the stop button above).
#
# *Hint: take old key from plate, unlock door with old key, open it, go east, open refrigerator, insert coffee cup into refrigerator, quit.*
quest = M.record_quest()
# ### Check recorded quest
# Now, let's see what the generated quest description looks like.
print(" > ".join(quest.commands))
print("\n" + quest.desc)
# ### Trying out the game
# At any point, we can easily test the game we are currently building (if it can compile).
# Test the game by issuing commands from the generated walkthrough.
# You can set `walkthrough=False` if you want to play the game yourself.
M.test(walkthrough=True)
# ### Randomly generate quests for your game
import copy
M2 = copy.deepcopy(M) # Work on a copy.
M2.generate_distractors(10) # Add 10 objects for the agent to interact with.
M2.render()
M2.generate_random_quests(5, length=2) # Generate 5 random quests of length 2.
game = M2.build()
print(game.objective)
# Test the game by issuing commands from the walkthrough.
M2.test(walkthrough=True)
# ## Handcrafting your game
# +
import textworld
from textworld import GameMaker
# Make the generation process reproducible.
from textworld import g_rng # Global random generator.
g_rng.set_seed(20180329)
# GameMaker object for handcrafting text-based games.
M = GameMaker()
# ---> Build your game here using TextWorld GameMaker's API. <---
# Uncomment the following lines to record a quest.
# quest = M.record_quest()
# print(" > ".join(quest.commands))
# print("\n" + quest.desc)
M.test() # Try your game.
# -
# ## Common Errors
# - `FailedConstraintsError`: One the constraints has failed. You need to resolve the issue before being able to compile the game. The most common issues:
# - Two objects cannot have the exact same name. Names can be composed of multiple words.
# - A same objects was added twice to a container, supporter, room or inventory.
# - `ValueError: Could not determine status '' of container/door ...`: You have to provide the status (a fact) of the door/container, i.e. 'open', 'closed' or 'locked'.
|
notebooks/Handcrafting a game.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Instruction: make sure your code pass the assertion statements
# Given a positive integer N. The task is to write a Python program to check if the number is prime or not.
def is_prime(n):
# Implement your code below here.
# check input - n
if n <= 0 and isinstance(n, int):
return False
# Algo
for
while
# early return
return False
# DO NOT ALTER BELOW.
# assert is_prime(2)
# assert not is_prime(15)
# assert is_prime(7907)
# assert not is_prime(-1)
# assert not is_prime(0)
# int - positive / 0 / negative
# type must be int
#print(isinstance(13, int))
#print(isinstance(35.0, int))
(1)
13 / 1, 13
[2,3,4...12] ->
if 整除 -> False
else -> True
14/2 False
13/3 ?
13/4 ?
...
13/12 ?
True
(2)
14
< n prime number --> [2,3,5,7,11,13]
1M << 1M ~= 10%
(1/10)
(3)
# +
# Write a function rotate(ar[], d) that rotates arr[] of size n by d elements.
# Input ar = [1,2,3,4,5,6,7], d = 2
# Output [3,4,5,6,7,1,2]
# understand question and test case
# check input
# algorithm
# check return
def rotate(ar, d):
# #? ar is None or Empty?
# d >= 0
return []
# DO NOT ALTER BELOW.
assert rotate([1,2,3,4,5,6,7], 2) == [3,4,5,6,7,1,2]
[12|34567] + [12|34567]
|3456712|
7234561
76543 21
34567 12
3456712
--> in-place 消耗时间,不消耗空间
1234567
rotate_once
2345671
list[0] -> delete,
list[-1] append list[0]
234567 <- 1
3456712
4567123
100 times
assert rotate([1,2,3], 4) == [2,3,1]
1234
2341
3412
4123
100 % 4 = 0
len(ar)
# +
# Selection sort - implement a workable selection sort algorithm
# https://www.runoob.com/w3cnote/selection-sort.html 作为参考
# Input students would be a list of [student #, score], sort by score ascending order.
def selection_sort(students):
return [[]]
# DO NOT ALTER BELOW.
assert selection_sort([] == [])
assert selection_sort([[1, 100], [2, 70], [3, 95], [4, 66], [5, 98]]) == [[4, 66], [2, 70], [3, 95], [5, 98], [1, 100]]
# +
# Convert a list of Tuples into Dictionary
def convert(tup, di):
# Do NOT RETURN di, EDIT IN-PLACE
# DO NOT ALTER BELOW.
expected_dict = {}
convert((), expected_dict)
assert expected_dict == {}
convert(('key1', 'val1', 'key2', 'val2'), expected_dict)
assert expected_dict == {'key1': 'val1', 'key2': 'val2'}
# +
# 研究为什么 Python dict 可以做到常数级别的查找效率,将答案写在 Assignment1-answer.ipynb
|
w2-python/homework/Assignment1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="kLSLxAdBGHSd"
# # Notebook for building and evaluating LSN
#
# * **Data shapes**
# - X_MR: samples x 2 x ROIs (baseline and follow-up)
# - X_aux: samples x 4 (gentic, age, clinical_score_bl, clinical_score_followup) #Can be user defined
# - y: samples x n_classes (needs to be one-hot i.e. [0,1] )
#
#
# * **network architecture names**
# - MR_shape: number of MR ROIs
# - n_layers: number of layers for MR Siamese net
# - l1,l2, etc: number of nodes in each layer
# - MR_output: number of output nodes (embedding dimension) for each MR branch (baseline, follow-up)
# - use_aux: LSN can be built just with MR or MR + aux_features
# - aux_shape: number of aux features excluding genetic (i.e. apoe4 status)
# - aux_output: number nodes for aux output (single layer ANN)
# - output: final output of LSN == number of classes
# - reg: weight regulalization for the network
# - Example: net_arch = {'MR_shape':78,'n_layers':4,'l1':50,'l2':50,'l3':50,'l4':25,'MR_output':20,'use_aux':True,'aux_shape':3,'aux_output':2,'output':2,'reg':0.01}
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="WYpfpG7UGLnX"
# Basic imports
# %matplotlib inline
import tensorflow as tf
import numpy as np
import random
import time
import math
import pandas as pd
from datetime import datetime
import sys
import os
import pickle
sys.path.append(os.path.relpath("../lib/"))
from lsn import *
# -
# # Real CT + aux data example
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 51} colab_type="code" executionInfo={"elapsed": 376, "status": "ok", "timestamp": 1528988676971, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-rmY3GxNQ0TY/AAAAAAAAAAI/AAAAAAAAEdE/SePqq1Clrc4/s50-c-k-no/photo.jpg", "userId": "104945077116245412065"}, "user_tz": 240} id="CoG7X_s41pch" outputId="4cc20727-22d6-437f-d5f9-bbb89efc12a2"
kfold_dir = '/data/chamal/projects/nikhil/data/Traj_prediction/ADNI/exp_setup/KFolds/'
exp_name = 'Exp_502_ALL_ADNI_traj_MMSE_tp_var_tp_sKF.pkl'
exp_setup = pd.read_pickle(kfold_dir+exp_name)
print('exp_name: {}'.format(exp_name))
df = exp_setup['df']
ct_cols_bl = list(df.columns[pd.Series(df.columns).str.contains('CT_bl')])
ct_cols_followup =list(df.columns[pd.Series(df.columns).str.contains('CT_var_tp')])
# Just a single split (no k-fold) as an example
train_split = int(0.9*len(df))
subject_idx_test = df['PTID'][train_split:]
# Train-test splits
X_MR_train = np.stack((df[ct_cols_bl].values,df[ct_cols_followup].values),axis=1)[:train_split]
X_aux_train = df[['APOE4','AGE','MMSE_bl','MMSE_var_tp']].values[:train_split]
y_train = df['traj'].values[:train_split]
y_train = np.squeeze(np.vstack((1-y_train,y_train)).T)
X_MR_test = np.stack((df[ct_cols_bl].values,df[ct_cols_followup].values),axis=1)[train_split:]
X_aux_test = df[['APOE4','AGE','MMSE_bl','MMSE_var_tp']].values[train_split:]
y_test = df['traj'].values[train_split:]
y_test = np.squeeze(np.vstack((1-y_test,y_test)).T)
print('shapes of X_MR_train:{}, X_aux_train:{}, y_train:{}, \n\t X_MR_test:{}, X_aux_test:{}, y_test:{}'.format(X_MR_train.shape,X_aux_train.shape,y_train.shape,
X_MR_test.shape,X_aux_test.shape,y_test.shape))
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 85} colab_type="code" executionInfo={"elapsed": 243, "status": "ok", "timestamp": 1528988735549, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-rmY3GxNQ0TY/AAAAAAAAAAI/AAAAAAAAEdE/SePqq1Clrc4/s50-c-k-no/photo.jpg", "userId": "104945077116245412065"}, "user_tz": 240} id="BoTQj0-C5oD2" outputId="8bb452a7-b82d-45fb-f0c5-2438bcfa5783"
# Check differences in aux_feature distributions
print('train aux distributions')
class_0 = np.mean(X_aux_train[y_train[:,0]==0],axis=0)
class_1 = np.mean(X_aux_train[y_train[:,0]==1],axis=0)
print(class_0,class_1)
print('test aux distributions')
class_0 = np.mean(X_aux_test[y_test[:,0]==0],axis=0)
class_1 = np.mean(X_aux_test[y_test[:,0]==1],axis=0)
print(class_0,class_1)
# -
# # Train and test a sample LSN
#
# **Note there is no hyper param tuning in this example **
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 510} colab_type="code" executionInfo={"elapsed": 2258, "status": "ok", "timestamp": 1528990262472, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-rmY3GxNQ0TY/AAAAAAAAAAI/AAAAAAAAEdE/SePqq1Clrc4/s50-c-k-no/photo.jpg", "userId": "104945077116245412065"}, "user_tz": 240} id="tvUx-7fivyzv" outputId="17b66e25-2448-47a4-9224-022ba21e34e5"
# training params
lr = 0.001
n_epochs = 100
validate_after = 10
batch_size = 100
dropout = 0.75 #keep_prob
verbose = False # Do you want to print perf after every epoch??
save_model = False # If you want to save the model to the disk (not needed usually)
save_model_path = '/data/chamal/projects/nikhil/data/Traj_prediction/ADNI/exp_setup/TF_trained_models/'
net_arch = {'MR_shape':78,'n_layers':3,'l1':25,'l2':25,'l3':25,'l4':25,'l5':25,'MR_output':10,
'use_aux':True,'aux_shape':4,'aux_output':10,'output':2,'reg':0.01}
# minimal perf df --> append CV related attributes downstream.
perf_df = pd.DataFrame(columns=['subject_id','label','pred_prob','pred_label'])
tf.reset_default_graph()
with tf.Session(config=tf.ConfigProto(log_device_placement=True)) as sess:
# ----------------------Train model -------------------------------
data = {'X_MR':X_MR_train,'X_aux':X_aux_train,'y':y_train}
if check_data_shapes(data,net_arch):
print('train data <-> net_arch check passed')
lsn = siamese_net(net_arch)
optimizer = tf.train.AdamOptimizer(learning_rate = lr).minimize(lsn.loss)
tf.global_variables_initializer().run()
saver = tf.train.Saver()
cur_time = datetime.time(datetime.now())
print('\nStart training time: {}'.format(cur_time))
lsn, train_metrics = train_lsn(sess, lsn, data, optimizer, n_epochs, batch_size, dropout,validate_after,verbose)
# Save trained model
# See at the end of this notebook for the code for using pretrained model
if save_model:
print('saving model at {}'.format(save_model_path + 'lsn_example'))
saver.save(sess, save_model_path + 'lsn_example')
cur_time = datetime.time(datetime.now())
print('End training time: {}\n'.format(cur_time))
else:
print('train data <-> net_arch check failed')
# Test model (within same session)
data = {'X_MR':X_MR_test,'X_aux':X_aux_test,'y':y_test}
if check_data_shapes(data,net_arch):
print('test data <-> net_arch check passed')
_,test_metrics = test_lsn(sess,lsn,data)
# populate perf dataframe
perf_df['subject_id'] = subject_idx_test
perf_df['label'] = np.argmax(y_test,1)
perf_df['pred_prob'] = list(test_metrics['test_preds'])
perf_df['pred_label'] = np.argmax(test_metrics['test_preds'],1)
else:
print('test data <-> net_arch check failed')
# -
# # Plots
# +
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.manifold import TSNE
plt.figure(figsize=(16,9))
plt.style.use('seaborn-white')
sns.set(font_scale=2)
train_loss = train_metrics['train_loss']
valid_loss = train_metrics['valid_loss']
train_acc = train_metrics['train_acc']
valid_acc = train_metrics['valid_acc']
plt.subplot(2,2,1)
plt.plot(train_loss,label='train');
plt.plot(valid_loss,label='valid');
plt.title('Loss (innerloop)')
plt.xlabel('number of epoch x{}'.format(validate_after))
plt.legend()
plt.subplot(2,2,2)
plt.plot(train_acc,label='train');
plt.plot(valid_acc,label='valid');
plt.plot(np.tile(test_metrics['test_acc'],len(train_acc)),'--',label='test')
plt.title('Acc')
plt.xlabel('number of epoch x{}'.format(validate_after))
plt.legend()
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 729} colab_type="code" executionInfo={"elapsed": 10585, "status": "ok", "timestamp": 1528990279108, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-rmY3GxNQ0TY/AAAAAAAAAAI/AAAAAAAAEdE/SePqq1Clrc4/s50-c-k-no/photo.jpg", "userId": "104945077116245412065"}, "user_tz": 240} id="uwCrzzgCQJJP" outputId="34495065-a74d-41cb-fdee-3f5f847e6956"
#Plot TSNE
tsne_sampx = 500 #Too slow for large number of samples
train_embed_L = train_metrics['train_feature_L']
train_embed_R = train_metrics['train_feature_R']
test_embed_L = test_metrics['test_feature_L']
test_embed_R = test_metrics['test_feature_R']
train_tsne_embed_L = TSNE(n_components=2,init='pca').fit_transform(train_embed_L[:tsne_sampx])
test_tsne_embed_L = TSNE(n_components=2,init='pca').fit_transform(test_embed_L[:tsne_sampx])
train_tsne_embed_R = TSNE(n_components=2,init='pca').fit_transform(train_embed_R[:tsne_sampx])
test_tsne_embed_R = TSNE(n_components=2,init='pca').fit_transform(test_embed_R[:tsne_sampx])
train_df_L = pd.DataFrame(columns=['x','y','labels','subset'])
test_df_L = pd.DataFrame(columns=['x','y','labels','subset'])
train_df_R = pd.DataFrame(columns=['x','y','labels','subset'])
test_df_R = pd.DataFrame(columns=['x','y','labels','subset'])
train_df_L['x'] = train_tsne_embed_L[:,0]
train_df_L['y'] = train_tsne_embed_L[:,1]
train_df_L['labels'] = y_train[:tsne_sampx]
train_df_L['subset'] = np.tile('train_L',len(y_train[:tsne_sampx]))
train_df_R['x'] = train_tsne_embed_R[:,0]
train_df_R['y'] = train_tsne_embed_R[:,1]
train_df_R['labels'] = y_train[:tsne_sampx]
train_df_R['subset'] = np.tile('train_R',len(y_train[:tsne_sampx]))
test_df_L['x'] = test_tsne_embed_L[:,0]
test_df_L['y'] = test_tsne_embed_L[:,1]
test_df_L['labels'] = y_test[:tsne_sampx]
test_df_L['subset'] = np.tile('test_L',len(y_test[:tsne_sampx]))
test_df_R['x'] = test_tsne_embed_R[:,0]
test_df_R['y'] = test_tsne_embed_R[:,1]
test_df_R['labels'] = y_test[:tsne_sampx]
test_df_R['subset'] = np.tile('test_R',len(y_test[:tsne_sampx]))
plot_df = train_df_L.append(train_df_R).append(test_df_L).append(test_df_R)
sns.lmplot(x='x',y='y',hue='labels',col='subset',col_wrap=2, fit_reg=False, markers='.',data=plot_df,size=4);
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 369} colab_type="code" executionInfo={"elapsed": 804, "status": "ok", "timestamp": 1528989626700, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-rmY3GxNQ0TY/AAAAAAAAAAI/AAAAAAAAEdE/SePqq1Clrc4/s50-c-k-no/photo.jpg", "userId": "104945077116245412065"}, "user_tz": 240} id="2IA_YZkNHaVy" outputId="871ac4c9-2385-4a70-fcab-c2aa52e13aae"
# Plot preds
train_features = train_metrics['train_preds']
test_features = test_metrics['test_preds']
train_df = pd.DataFrame(columns=['x','y','labels','subset'])
test_df = pd.DataFrame(columns=['x','y','labels','subset'])
train_df['x'] = train_features[:,0]
train_df['y'] = train_features[:,1]
train_df['labels'] = y_train
train_df['subset'] = np.tile('train',len(y_train))
test_df['x'] = test_features[:,0]
test_df['y'] = test_features[:,1]
test_df['labels'] = y_test
test_df['subset'] = np.tile('test',len(y_test))
plot_df = train_df.append(test_df)
sns.lmplot(x='x',y='y',hue='labels',col='subset',fit_reg=False, markers='.',data=plot_df);
# -
# # Loading previously trained model
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="N6VZGx8n8Y1O"
# with tf.Session() as sess:
# new_saver = tf.train.import_meta_graph(save_model_path + 'lsn_example.meta')
# new_saver.restore(sess, tf.train.latest_checkpoint(save_model_path))
# _,test_metrics = test_lsn(sess,lsn,data)
|
notebooks/run_lsn.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Qiskit v0.35.0 (ipykernel)
# language: python
# name: python3
# ---
# # Variational Quantum Eigensolvers
#
# # ANALYSING TRADE-OFFS IN SYMMETRY-PRESERVING ANSATZ CIRCUITS FOR THE SCALABLE VARIATIONAL QUANTUM EIGENSOLVER ALGORITHM.
#
# VQE - hybrid classical-quantum algorithm that can find the ground state energy of various quantum systems.
#
# Has applications in quantum chemistry, where resources to classically simulate molecular wavefunctions increase exponentially with size of molecule.
#
# True speedups in computational chemistry are still far off and dependant on development of large fault-tolerant quantum computers, VQEs can already be used to determine ground state energy of smaller molecules to a high degree of accuracy
# $$\newcommand{\ket}[1]{\left|{#1}\right\rangle}$$
# $$\newcommand{\bra}[1]{\left\langle{#1}\right|}$$
# $$\newcommand{\braket}[2]{\left\langle{#1}\middle|{#2}\right\rangle}$$
# +
from typing import Optional, Union, Callable, cast
import matplotlib.pyplot as plt
import numpy as np
from qiskit import IBMQ, BasicAer, Aer
from qiskit.providers.aer import StatevectorSimulator
from qiskit.utils import QuantumInstance
from qiskit_nature.results import ElectronicStructureResult
from qiskit_nature.drivers import UnitsType, Molecule, QMolecule
from qiskit_nature.drivers.second_quantization import PySCFDriver, ElectronicStructureDriverType, ElectronicStructureMoleculeDriver
from qiskit_nature.problems.second_quantization import ElectronicStructureProblem
from qiskit_nature.converters.second_quantization import QubitConverter
from qiskit_nature.mappers.second_quantization import ParityMapper, JordanWignerMapper
from qiskit_nature.circuit.library import HartreeFock, UCC, UCCSD
from qiskit.opflow.primitive_ops import Z2Symmetries
from qiskit_nature.algorithms import GroundStateEigensolver
from qiskit_nature.algorithms import VQEUCCFactory
from qiskit.algorithms import NumPyMinimumEigensolver, VQE
from qiskit.algorithms.optimizers import SLSQP, COBYLA
# -
# ### Overview
#
# System is represented by a complex-valued wavefunction, and energy is associated with Hamiltonian $\mathbb{H}$, a matrix with eigenvalues that represent possible energies of the system.
#
# System's groudn state is eigenstate of the Hamiltonian with lowest eigenvalue.
#
# $\mathcal{H}\ket{\psi_g} = E_g\ket{\psi_g}$
#
# $E_g = \frac{ \bra{\psi_g}|\mathcal{H}|\ket{\psi_g} }{ \braket{\psi_g}{\psi_g} }$
#
# $E_g \leq \frac{ \bra{\psi}|\mathcal{H}|\ket{\psi} }{ \braket{\psi}{\psi} }$
#
# VQE leverages this inequality by parameterizing quantum state used to evaluate energy expectation, and varationally updating these parameters on a classical computer to find tight upper bound to the ground state.
# ### Algorithm
#
# * Parameterized quantum circuit prepares trialground state using some initial parameters values
# * A quantum measurement circuit estimates energy expectation in this trial state.
# * Classical optimization algorithm updates parameters to idealy decrease energy expectation of the next trail state.
# * Algorithm is iterates until energy expectation converges to some value, which is taken as the approx ground state energy.
# ### Challenges
#
# * Number of parameters required t prepare arbitrary quantum states increases problematically with system size, challenging the classical optimizer.
# ### Solutions
#
# * reduce number of parameters, hence reduce algorithmic search space.
# ### Classical Optimizers
#
# Responsible for determining parameter updates, thus for generating the new variational trial states.
#
# * Gradient Descent - identifying energy expectation as const function to be minimized, but has tendency of being stuch in local minima and has relatively high number of required quantum circuit evaluations. (poor optimizer for VQE).
#
# Popular optimizers for noise-free models i.e statevector simulations on classical computers. .e.g:
# * Nelder-Mead method
# * Sequential Least Squares Programming (SLSQP)
# * Constrained Optimization by Linear Optimization (COBYLA) - being more efficient at the expense of fewer(just one) energy expectation evaluations per iteration.
#
# VQE on physical quantum coputers incorporating noise models often involves more nuanced optimization schemes.
# ### Hamiltonian mapping
#
# How wavefunction of a molecule can be mapped onto qubit states of a quantum computer.
#
# General Hamiltonian for a quantum molecular system: (very intuitive)
#
# $$\mathcal{\hat{H}} = - \sum_i \frac{\nabla^2_i}{2} - \sum_{i, I} \frac{Z_I}{|r_i - R_I|} + \sum_{i\neq j} \frac{1}{2|r_i - r_j|}$$
#
# where: H = kinetic energy - electron-nucleus interaction + electron-electron interaction
#
# Where:
#
# $i$ and $j$ denote electrons,
#
# $I$ denotes nuclei,
#
# $Z$ is the atomic number of nuclei $I$,
#
# $r$ is position of the electron $i$, $j$,
#
# $R$ is position of each nucleus, which is fixed in space in accordance with the Bohr-Oppenheimer approximation.
#
# The $\textbf{second quantization}$ Hamiltonian in which electrons are treated as excitations in an electron field, will ultimately prove more useful in our effort to encode probelm onto quantum copmuter:
#
# $\begin{align}
# \mathcal{\hat{H}} = \sum_{p,q} h_{pq}a^\dagger_p a_q + \frac{1}{2}\sum_{p,q,r,s}h_{pqrs}a^\dagger_p a^\dagger_q a_r a_s
# \end{align}$
#
# The $a^\dagger$ operators (creation operator) in each term "excite" electron to orbitals $p$ and $(p, q)$ respectively. The $a$ operator (annihilation operator) in each term $de-excites$ electron from orbitals $q$ and $(r, s)$ respectively.
#
# The $h$ terms are coefficients known as the one an two-electron integrals and can be calculated relatively eficciently in terms of a small set of orbital basis states on a classical computer.
#
# We will restrict ourselves to single- and double-electron excitations to obtain a workable approximation of the more complicated true physical system.
#
# There are 3 common mappings that can be used to produce a Hamiltonian for distinguishable fermions i.e. qubits from a Hmailtonian for indistinguishable fermions i.e. electrons:
# * Jordan-Wigner mapping
# * Parity mapping
# * Bravyi-Kitaev mapping
#
# All the mappings produce a Hamitonian of the form:
#
# $\begin{equation}
# \mathcal{\hat{H}} = \sum_j \alpha_j (\prod_i \hat{\sigma^j_i})
# \end{equation}$
#
# This is to say the Hamiltonian will be a linear combination of products of Pauli matrices (with $i$ denoting qubit being acted upon) which can be executed on a quantum computer.
#
# Given a trial state, the energy expectation may be evaluated by measuring this superposition of Pauli operators, as below:
#
# $E(\vec{\theta}) = \sum^N_j \alpha_j \bra{\psi(\vec{\theta})} \prod_i \sigma_i^j \ket{\psi(\vec{\theta})}$
# ### Parameterized Wavefunction Ansatz
#
# Among most common wavefunction ansatz is the Unitary Coupled-CLuster Single and Double excitation (UCCSD) ansatz, used in foundational VQE paper by Perruzo et al.
#
# UCCSD is constructed by applying an exponentiated single and double-electron excitation operator to an initial state, commonly chossen to be the Hartree-Fock mean-field wavefunction (an unentangled state that decently approximates the ground state):
#
# $
# \begin{align}
# \ket{\psi(\theta)} = e^({\vec{T} - \vec{T}^\dagger}) \ket{\phi}\\\\
# \hat{T} = \sum_{i\in virt, j\in occ} t^j_i \hat{a}_i^\dagger + \sum_{i, j \in virt, k, l \in occ} t^{kl}_{ij} \hat{a}_i^\dagger \hat{a}_j^\dagger \hat{a}_k \hat{a}_l
# \end{align}
# $
#
# Where:
#
# "virt" denotes unoccupied orbitals,
#
# "occ" denotes occupied orbitals,
#
# $a\dagger$ creation operators excite electrons and $a$ annihilation operators de-excite electrons to/from orbiatls,
#
# t coefficients are the tunable parameters that are fed into classical optimizer.
#
# The $T$ operator is then converted via one of the 3 mappings into as effective Hamiltoniann operator on qubits, which may be subsequently executed on a quantum copmuter.
# ### Chemistry-Inspired Ansatz
#
# * UCCSD - UCC wit Singles and Doubles
# * UpCCD - Unitary Pair UCCD
# * OO-UCC - Orbital Optimized UCC
# * DUCC - Double UCC
#
#
# ### Hardware Efficient Ansatz
#
# * Symmetry Preserving State Preparation
# * Qubit Coupled Cluster Method
# ### Energy
#
# The hartree (symbol: Eh or Ha), also known as the Hartree energy, is the unit of energy in the Hartree atomic units system, named after the British physicist <NAME>. Its CODATA recommended value is Eh = 4.3597447222071(85)×10−18 J[1] = 27.211386245988(53) eV.[2]
#
# The hartree energy is approximately the electric potential energy of the hydrogen atom in its ground state and, by the virial theorem, approximately twice its ionization energy; the relationships are not exact because of the finite mass of the nucleus of the hydrogen atom and relativistic corrections.
#
# The hartree is usually used as a unit of energy in atomic physics and computational chemistry: for experimental measurements at the atomic scale, the electronvolt (eV) or the reciprocal centimetre (cm−1) are much more widely used.
#
# (Courtesy: Wikipedia: https://en.wikipedia.org/wiki/Hartree)
# ### Building a VQE in Qiskit
#
# VQE in Python using Qiskit, as open-source SDK for working with quantum computers, to find ground state energy and corresponding interatomic spearaton for several small molecules: diatomic hydrogen ($H_2$), lithium hydride ($LiH$), ionic helium hydride ($HeH+$) and hydroxide ($OH-$)
#
# ---
# * First the molecule structure was specified as a string in xyz coordinates, lithium hydroxide woudl specify as "Li 0.0 0.0 0.0; H 0.0 0.0 d" where d is the interatomic distance.
#
# For two-atom molecules, the z-coordinate of the second atom was varied to determine ground state energy as a function of interatomic distance.
#
# * A PySCF driver was initialized with this molecule specification, creating a data structure representing the molecule along with several useful calculated attributes, including the nuclear repulsion energy - a quantity later added to the VQE-determined electron energy to obtain the total molecular energy.
#
# The PySCF molecular data structure was then provided as input to the ElectronicStructureProblem class in Qiskit-Nature module. This class determines the Hamiltoninan of the molecule in terms of second-quantization operators, calculating the one- and two-electron integral coefficients discussed previously and returning the electronic operator in the form:
#
#
# Where:
#
# "+" is the excitation operator
#
# "-" is the de-excitation operator
#
# "I" is the identity operator
#
# "N" is the number operator (de-excitation followed by excitation)
#
# This operator was then converted into a qubit operator using the class qiskit_nature.converters.second_quantization,QubitConverter, with the mapping type set to the Jordan-Wigner transformation, yielding a summation of Pauli operator products:
#
#
#
# Note that the number of qubits needed to execute algorithm s equal to number of second quantization operators, which is defined as the number of molecular spin orbitals considered.
#
# * Next,the Hartree-Fock initial state preparation circuit and the UCCSD variational unitary transformation circuit. (with HF as its initial state) were retrieved from a library of optimized circuits in the qiskit_nature.circuits.library module, being sure to pass the Jordan-Wigner QubitConverter object created earlier as an argument to each circuit for consistency.
#
# Finally, these were all supplied to the VQE class in the module qiskit.algorithms, which simulated the UCCSD variational circuit repeatedly on a Qiskit Aer statevector backend to solve for the minimum eigenvalue of the qubit Hamiltonina, using the SLSQP classical optimizer to calculate parameter updates.
#
# This algorithm was repeated for intermolecular separations ranging between 0.2 and 2 Angstroms (~0.1 nanometres), storing the minimum energy as determined by the VQE at each iteration.
# +
state_sim = StatevectorSimulator()
from qiskit_nature.transformers.second_quantization.electronic import FreezeCoreTransformer
from qiskit_nature.algorithms import ExcitedStatesEigensolver, VQEUCCFactory
# vqe_energies = []
class MolecularVQE:
def __init__(self):
# H2
self.molecule_name = "H 0.0 0.0 0.0; H 0.0 0.0 "
self.backend = QuantumInstance(state_sim)
self.optimizer = SLSQP(maxiter=500)
self.vqe = VQE(
ansatz = None,
quantum_instance = self.backend,
optimizer = self.optimizer
)
def get_qubit_op(self, dist, mapper="parity"):
# Use PySCF, a classical computational chemistry software
# package, to compute the on-body and two-body integrals in
# electronic-orbital basis, necessary to form the Fermionic operator
driver = PySCFDriver(
atom = self.molecule_name + str(dist),
# unit = UnitsType.ANGSTROM,
# charge = 0,
# spin = 0,
# basis = "sto3g"
)
#molecule = Molecule(
# geometry=[
# ["H", [0.0, 0.0, 0.0]],
# ["H", [0.0, 0.0, 0.735]]
# ],
# charge=0,
# multiplicity=1
#)
#driver = ElectronicStructureMoleculeDriver(
# basis = "sto3g",
# driver_type=ElectronicStructureDriverType.PYSCF
#)
qmolecule = driver.run() # returns ElectronicStructureDriverResult
transformer = FreezeCoreTransformer()
qmolecule = transformer.transform(qmolecule)
es_problem = ElectronicStructureProblem(driver)
if mapper == "jw":
qubit_converter = QubitConverter(mapper=JordanWignerMapper())
elif mapper == "parity":
qubit_converter = QubitConverter(mapper=ParityMapper(), two_qubit_reduction=True)
return (es_problem, qubit_converter)
def run(self):
numpy_solver = NumPyMinimumEigensolver()
# distances = np.arange(0.3, 2.0, 0.05)
distances = np.arange(0.65, 1.0, 0.05)
exact_energies = []
vqe_energies = []
n = len(distances)
i = 1
for dist in distances:
print("Distance {}/{}".format(i, n))
i += 1
es_problem, qubit_converter = self.get_qubit_op(dist)
second_q_ops = es_problem.second_q_ops()
# Hamiltonian
main_op = second_q_ops[0]
#mine z2sym = Z2Symmetries.find_Z2_symmetries(second_q_ops)
qubit_op = qubit_converter.convert(
main_op,
num_particles = es_problem.num_particles,
#sector_locator = es_problem.symmetry_sector_locator(None, qubit_converter)
)
# aux_ops = qubit_converter.convert_match(
# second_q_ops[1:]
# )
# q_molecule_transformed = cast(QMolecule, es_problem.molecule_data_transformed)
# num_molecular_orbitals = q_molecule_transformed.num_molecular_orbitals
# num_particles = (q_molecule_transformed.num_alpha, q_molecule_transformed.num_beta)
# num_spin_orbitals = 2 * num_molecular_orbitals
num_particles = es_problem.num_particles
num_spin_orbitals = es_problem.num_spin_orbitals
# initial state is Hartree-Fock state
initial_state = HartreeFock(num_spin_orbitals, num_particles, qubit_converter)
# UCCSD ansatz for unitary update
ansatz = UCCSD()
ansatz.qubit_converter = qubit_converter
ansatz.num_particles = num_particles
ansatz.num_spin_orbitals = num_spin_orbitals
ansatz.initial_state = initial_state
self.vqe.ansatz = ansatz
solver = self.vqe
# solver = VQEUCCFactory(
# quantum_instance=self.backend,
# optimizer=self.optimizer,
# ansatz=ansatz,
# initial_state=initial_state
# )
uccsd_excited_states_calculation = ExcitedStatesEigensolver(qubit_converter, solver)
print(uccsd_excited_states_calculation)
# uccsd_ground_excited_states_properties = uccsd_excited_states_calculation.solve(es_problem)
print("Computing the minimum eigenvalue...")
# Approximate minimum eigensolver using VQE
vqe_result = solver.compute_minimum_eigenvalue(qubit_op)
print(vqe_result)
# print(np.real(vqe_result.eigenvalue) + nuclear_repulsion_energy)
# vqe_energies.append(np.real(vqe_result.eigenvalue) + nuclear_repulsion_energy)
vqe_energies.append(np.real(vqe_result.eigenvalue))
return (distances, vqe_energies, exact_energies)
# return self.vqe.ansatz
# -
# ### Results
#
# Demonstrate close correspondence between experimentally determined ground state energies and the ground state energies determined using VQEs. The ground state interatmoic distance was dtermined by finding the minimum on the plot of VQE ground state energy against interatomic distance.
#
# Find a theoretical curve of ground stte energy against interatomic distance - determined by diagonalizing the molecule Hamiltonian directly using NumpyMinimumEigensolver - check if it coincides with VQE curve.
# +
vqe = MolecularVQE()
res = vqe.run()
# res.draw("mpl")
# +
plt.plot(res[0], res[1], c="r", label="VQE")
# plt.rc("text", usetext=True)
plt.title(r"$H2$: Minimum energy vs. Interatomic distance")
plt.ylabel("Ground state energy (Hartree)")
plt.xlabel("Interatomic distance (A)")
plt.legend()
# plt.show()
idx = res[1].index(min(res[1]))
dist, min_energy = res[0][res[1].index(min(res[1]))], min(res[1])
print("Min Distance: {}\n\n".format(dist))
print("Min Energy: {}".format(min_energy))
# -
|
VQE/vqe_molecular_example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Basic training functionality
# + hide_input=true
from fastai.basic_train import *
from fastai.gen_doc.nbdoc import *
from fastai.vision import *
from fastai.distributed import *
# -
# [`basic_train`](/basic_train.html#basic_train) wraps together the data (in a [`DataBunch`](/basic_data.html#DataBunch) object) with a pytorch model to define a [`Learner`](/basic_train.html#Learner) object. This is where the basic training loop is defined for the [`fit`](/basic_train.html#fit) function. The [`Learner`](/basic_train.html#Learner) object is the entry point of most of the [`Callback`](/callback.html#Callback) functions that will customize this training loop in different ways (and made available through the [`train`](/train.html#train) module), notably:
#
# - [`Learner.lr_find`](/train.html#lr_find) will launch an LR range test that will help you select a good learning rate
# - [`Learner.fit_one_cycle`](/train.html#fit_one_cycle) will launch a training using the 1cycle policy, to help you train your model fast.
# - [`Learner.to_fp16`](/train.html#to_fp16) will convert your model in half precision and help you launch a training in mixed precision.
# + hide_input=true
show_doc(Learner, title_level=2)
# -
# The main purpose of [`Learner`](/basic_train.html#Learner) is to train `model` using [`Learner.fit`](/basic_train.html#Learner.fit). After every epoch, all *metrics* will be printed, and will also be available to callbacks.
#
# The default weight decay will be `wd`, which will be handled using the method from [Fixing Weight Decay Regularization in Adam](https://arxiv.org/abs/1711.05101) if `true_wd` is set (otherwise it's L2 regularization). If `bn_wd` is False then weight decay will be removed from batchnorm layers, as recommended in [Accurate, Large Minibatch SGD: Training ImageNet in 1 Hour](https://arxiv.org/abs/1706.02677). You can ensure that batchnorm layer learnable params are trained even for frozen layer groups, by enabling `train_bn`.
#
# To use [discriminative layer training](#Discriminative-layer-training) pass an [`nn.Module`](https://pytorch.org/docs/stable/nn.html#torch.nn.Module) for each layer group to be optimized with different settings.
#
# Any model files created will be saved in `path`/`model_dir`.
#
# You can pass a list of [`callbacks`](/callbacks.html#callbacks) that you have already created, or (more commonly) simply pass a list of callback functions to `callback_fns` and each function will be called (passing `self`) on object initialization, with the results stored as callback objects. For a walk-through, see the [training overview](/training.html) page. You may also want to use an `application` to fit your model, e.g. using the [`create_cnn`](/vision.learner.html#create_cnn) method:
# + hide_input=false
path = untar_data(URLs.MNIST_SAMPLE)
data = ImageDataBunch.from_folder(path)
learn = create_cnn(data, models.resnet18, metrics=accuracy)
learn.fit(1)
# -
# ### Model fitting methods
# + hide_input=true
show_doc(Learner.fit)
# -
# Uses [discriminative layer training](#Discriminative-layer-training) if multiple learning rates or weight decay values are passed. To control training behaviour, use the [`callback`](/callback.html#callback) system or one or more of the pre-defined [`callbacks`](/callbacks.html#callbacks).
# + hide_input=true
show_doc(Learner.fit_one_cycle)
# -
# Uses the [`OneCycleScheduler`](/callbacks.one_cycle.html#OneCycleScheduler) callback.
# + hide_input=true
show_doc(Learner.lr_find)
# -
# Runs the learning rate finder defined in [`LRFinder`](/callbacks.lr_finder.html#LRFinder), as discussed in [Cyclical Learning Rates for Training Neural Networks](https://arxiv.org/abs/1506.01186).
# ### See results
# + hide_input=true
show_doc(Learner.get_preds)
# + hide_input=true
show_doc(Learner.validate)
# + hide_input=true
show_doc(Learner.show_results)
# + hide_input=true
show_doc(Learner.predict)
# + hide_input=true
show_doc(Learner.pred_batch)
# + hide_input=true
show_doc(Learner.interpret, full_name='interpret')
# + hide_input=true
jekyll_note('This function only works in the vision application.')
# -
# ### Model summary
# + hide_input=true
show_doc(Learner.summary)
# -
# ### Test time augmentation
# + hide_input=true
show_doc(Learner.TTA, full_name = 'TTA')
# -
# Applies Test Time Augmentation to `learn` on the dataset `ds_type`. We take the average of our regular predictions (with a weight `beta`) with the average of predictions obtained through augmented versions of the training set (with a weight `1-beta`). The transforms decided for the training set are applied with a few changes `scale` controls the scale for zoom (which isn't random), the cropping isn't random but we make sure to get the four corners of the image. Flipping isn't random but applied once on each of those corner images (so that makes 8 augmented versions total).
# ### Gradient clipping
# + hide_input=true
show_doc(Learner.clip_grad)
# -
# ### Mixed precision training
# + hide_input=true
show_doc(Learner.to_fp16)
# -
# Uses the [`MixedPrecision`](/callbacks.fp16.html#MixedPrecision) callback to train in mixed precision (i.e. forward and backward passes using fp16, with weight updates using fp32), using all [NVIDIA recommendations](https://docs.nvidia.com/deeplearning/sdk/mixed-precision-training/index.html) for ensuring speed and accuracy.
# + hide_input=true
show_doc(Learner.to_fp32)
# -
# ### Distributed training
# + hide_input=true
show_doc(Learner.distributed, full_name='distributed')
# -
# ### Discriminative layer training
# When fitting a model you can pass a list of learning rates (and/or weight decay amounts), which will apply a different rate to each *layer group* (i.e. the parameters of each module in `self.layer_groups`). See the [Universal Language Model Fine-tuning for Text Classification](https://arxiv.org/abs/1801.06146) paper for details and experimental results in NLP (we also frequently use them successfully in computer vision, but have not published a paper on this topic yet). When working with a [`Learner`](/basic_train.html#Learner) on which you've called `split`, you can set hyperparameters in four ways:
#
# 1. `param = [val1, val2 ..., valn]` (n = number of layer groups)
# 2. `param = val`
# 3. `param = slice(start,end)`
# 4. `param = slice(end)`
#
# If we chose to set it in way 1, we must specify a number of values exactly equal to the number of layer groups. If we chose to set it in way 2, the chosen value will be repeated for all layer groups. See [`Learner.lr_range`](/basic_train.html#Learner.lr_range) for an explanation of the `slice` syntax).
#
# Here's an example of how to use discriminative learning rates (note that you don't actually need to manually call [`Learner.split`](/basic_train.html#Learner.split) in this case, since fastai uses this exact function as the default split for `resnet18`; this is just to show how to customize it):
# creates 3 layer groups
learn.split(lambda m: (m[0][6], m[1]))
# only randomly initialized head now trainable
learn.freeze()
learn.fit_one_cycle(1)
# all layers now trainable
learn.unfreeze()
# optionally, separate LR and WD for each group
learn.fit_one_cycle(1, max_lr=(1e-4, 1e-3, 1e-2), wd=(1e-4,1e-4,1e-1))
# + hide_input=true
show_doc(Learner.lr_range)
# -
# Rather than manually setting an LR for every group, it's often easier to use [`Learner.lr_range`](/basic_train.html#Learner.lr_range). This is a convenience method that returns one learning rate for each layer group. If you pass `slice(start,end)` then the first group's learning rate is `start`, the last is `end`, and the remaining are evenly geometrically spaced.
#
# If you pass just `slice(end)` then the last group's learning rate is `end`, and all the other groups are `end/10`. For instance (for our learner that has 3 layer groups):
learn.lr_range(slice(1e-5,1e-3)), learn.lr_range(slice(1e-3))
# + hide_input=true
show_doc(Learner.unfreeze)
# -
# Sets every layer group to *trainable* (i.e. `requires_grad=True`).
# + hide_input=true
show_doc(Learner.freeze)
# -
# Sets every layer group except the last to *untrainable* (i.e. `requires_grad=False`).
# + hide_input=true
show_doc(Learner.freeze_to)
# + hide_input=true
show_doc(Learner.split)
# -
# A convenience method that sets `layer_groups` based on the result of [`split_model`](/torch_core.html#split_model). If `split_on` is a function, it calls that function and passes the result to [`split_model`](/torch_core.html#split_model) (see above for example).
# ### Saving and loading models
# Simply call [`Learner.save`](/basic_train.html#Learner.save) and [`Learner.load`](/basic_train.html#Learner.load) to save and load models. Only the parameters are saved, not the actual architecture (so you'll need to create your model in the same way before loading weights back in). Models are saved to the `path`/`model_dir` directory.
# + hide_input=true
show_doc(Learner.load)
# + hide_input=true
show_doc(Learner.save)
# -
# ### Segmentation model
# + hide_input=true
show_doc(unet_learner)
# -
# Build a [`Learner`](/basic_train.html#Learner) for segmentation tasks from [`data`](/vision.data.html#vision.data), using `arch` that may be `pretrained` if that flag is `True`. `split_on` will overwrite the default way the layers are split for differential learning rates. The underlying model is a `DynamicUnet` with `blur`, `blur_final`, `self_attention`, `sigmoid` and `last_cross`. `norm_type` is passed to [`conv_layer`](/layers.html#conv_layer), the `kwargs` are passed to the [`Learner`](/basic_train.html#Learner) constructor.
# ### Other methods
# + hide_input=true
show_doc(Learner.init)
# -
# Initializes all weights (except batchnorm) using function `init`, which will often be from PyTorch's [`nn.init`](https://pytorch.org/docs/stable/nn.html#torch-nn-init) module.
# + hide_input=true
show_doc(Learner.mixup)
# -
# Uses [`MixUpCallback`](/callbacks.mixup.html#MixUpCallback).
# + hide_input=true
show_doc(Learner.backward)
# + hide_input=true
show_doc(Learner.create_opt)
# -
# You generally won't need to call this yourself - it's used to create the [`optim`](https://pytorch.org/docs/stable/optim.html#module-torch.optim) optimizer before fitting the model.
# + hide_input=true
show_doc(Learner.dl)
# + hide_input=true
show_doc(Recorder, title_level=2)
# -
# A [`Learner`](/basic_train.html#Learner) creates a [`Recorder`](/basic_train.html#Recorder) object automatically - you do not need to explicitly pass it to `callback_fns` - because other callbacks rely on it being available. It stores the smoothed loss, hyperparameter values, and metrics for each batch, and provides plotting methods for each. Note that [`Learner`](/basic_train.html#Learner) automatically sets an attribute with the snake-cased name of each callback, so you can access this through `Learner.recorder`, as shown below.
# + [markdown] hide_input=true
# ### Plotting methods
# + hide_input=true
show_doc(Recorder.plot)
# -
# This is mainly used with the learning rate finder, since it shows a scatterplot of loss vs learning rate.
learn = create_cnn(data, models.resnet18, metrics=accuracy)
learn.lr_find()
learn.recorder.plot()
# + hide_input=true
show_doc(Recorder.plot_losses)
# -
# Note that validation losses are only calculated once per epoch, whereas training losses are calculated after every batch.
learn.fit_one_cycle(2)
learn.recorder.plot_losses()
# + hide_input=true
show_doc(Recorder.plot_lr)
# -
learn.recorder.plot_lr(show_moms=True)
# + hide_input=true
show_doc(Recorder.plot_metrics)
# -
# Note that metrics are only collected at the end of each epoch, so you'll need to train at least two epochs to have anything to show here.
learn.recorder.plot_metrics()
# ### Callback methods
# You don't call these yourself - they're called by fastai's [`Callback`](/callback.html#Callback) system automatically to enable the class's functionality.
# + hide_input=true
show_doc(Recorder.on_backward_begin)
# + hide_input=true
show_doc(Recorder.on_batch_begin)
# + hide_input=true
show_doc(Recorder.on_epoch_end)
# + hide_input=true
show_doc(Recorder.on_train_begin)
# -
# ### Inner functions
# The following functions are used along the way by the [`Recorder`](/basic_train.html#Recorder) or can be called by other callbacks.
# + hide_input=true
show_doc(Recorder.add_metrics)
# + hide_input=true
show_doc(Recorder.add_metric_names)
# + hide_input=true
show_doc(Recorder.format_stats)
# -
# ## Module functions
# Generally you'll want to use a [`Learner`](/basic_train.html#Learner) to train your model, since they provide a lot of functionality and make things easier. However, for ultimate flexibility, you can call the same underlying functions that [`Learner`](/basic_train.html#Learner) calls behind the scenes:
# + hide_input=true
show_doc(fit)
# -
# Note that you have to create the `Optimizer` yourself if you call this function, whereas [`Learn.fit`](/basic_train.html#fit) creates it for you automatically.
# + hide_input=true
show_doc(train_epoch)
# -
# You won't generally need to call this yourself - it's what [`fit`](/basic_train.html#fit) calls for each epoch.
# + hide_input=true
show_doc(validate)
# -
# This is what [`fit`](/basic_train.html#fit) calls after each epoch. You can call it if you want to run inference on a [`DataLoader`](https://pytorch.org/docs/stable/data.html#torch.utils.data.DataLoader) manually.
# + hide_input=true
show_doc(get_preds)
# + hide_input=true
show_doc(loss_batch)
# -
# You won't generally need to call this yourself - it's what [`fit`](/basic_train.html#fit) and [`validate`](/basic_train.html#validate) call for each batch. It only does a backward pass if you set `opt`.
# ## Other classes
# + hide_input=true
show_doc(LearnerCallback, title_level=3)
# + hide_input=true
show_doc(RecordOnCPU, title_level=3)
# -
# ## Undocumented Methods - Methods moved below this line will intentionally be hidden
# + hide_input=true
show_doc(Learner.tta_only)
# + hide_input=true
show_doc(Learner.TTA)
# -
show_doc(RecordOnCPU.on_batch_begin)
# ## New Methods - Please document or move to the undocumented section
# + hide_input=false
show_doc(Learner.distributed)
# -
#
|
docs_src/basic_train.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.10.0 64-bit
# language: python
# name: python3
# ---
with open('1.txt') as f:
arr=[]
for lines in f:
arr.append(int(lines.strip()))
res=0
for i in range(1,len(arr)):
if arr[i]>arr[i-1]:
res+=1
print(res)
with open('1.txt') as f:
arr=[]
for lines in f:
arr.append(int(lines.strip()))
res=0
for i in range(2,len(arr)-1):
if sum(arr[i-1:i+2])>sum(arr[i-2:i+1]):
res+=1
print(res)
|
1/aoc2021_1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.7 64-bit (''venv'': venv)'
# metadata:
# interpreter:
# hash: ed2973e18a45e46195aea83f71ebc09fe8d3266afb265a31fa421a956b7d8fb0
# name: 'Python 3.7.7 64-bit (''venv'': venv)'
# ---
# # Query Wikidata for Belgian politicians
# +
from datetime import datetime as dt
from SPARQLWrapper import SPARQLWrapper, JSON
# -
# ## Fonctions
def get_rows():
"""Retrieve results from SPARQL"""
endpoint = "https://query.wikidata.org/bigdata/namespace/wdq/sparql"
sparql = SPARQLWrapper(endpoint)
statement = """
SELECT DISTINCT ?person ?personLabel ?dateBirth ?dateDeath WHERE {
?person wdt:P27 wd:Q31 .
?person wdt:P106 wd:Q82955 .
?person wdt:P569 ?dateBirth .
OPTIONAL {?person wdt:P570 ?dateDeath .}
SERVICE wikibase:label { bd:serviceParam wikibase:language "en" . }
}
ORDER BY ?personLabel
"""
sparql.setQuery(statement)
sparql.setReturnFormat(JSON)
results = sparql.query().convert()
rows = results['results']['bindings']
print(f"\n{len(rows)} Belgian politicians found\n")
return rows
def show(rows, name_filter=None, n=10):
"""Display n politicians (default=10)"""
date_format = "%Y-%m-%dT%H:%M:%SZ"
if name_filter:
rows = [row for row in rows if name_filter in row['personLabel']['value'].lower()]
print(f"Displaying the first {n}:\n")
for row in rows[:n]:
try:
birth_date = dt.strptime(row['dateBirth']['value'], date_format)
birth_year = birth_date.year
except ValueError:
birth_year = "????"
try:
death_date = dt.strptime(row['dateDeath']['value'], date_format)
death_year = death_date.year
except ValueError: # unknown death date
death_year = "????"
except KeyError: # still alive
death_year = ""
print(f"{row['personLabel']['value']} ({birth_year}-{death_year})")
# ## Obtenir une liste des politiciens belges depuis Wiki Data
# + tags=[]
rows = get_rows()
# -
rows[:10]
# ## Imprimer les 30 premiers politiciens
# + tags=[]
show(rows, n=30)
# -
# ## Filter pour afficher seulement les noms contenant "elio"
# + tags=[]
show(rows, name_filter="heyman")
# -
|
module1/s2_sparql.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # GLAM CSV Explorer
#
# [View code at GitHub](https://github.com/GLAM-Workbench/csv-explorer/blob/master/csv-explorer.ipynb) · [View details in GLAM Workbench](https://glam-workbench.net/csv-explorer/)
#
# Cultural institutions are making collection data available as machine readable downloads. But how can researchers explore the shape and meaning of this data? How do they know what types of questions they can ask?
#
# This notebook provides a quick overview of CSV-formatted data files, particularly those created by GLAM institutions (galleries, libraries, archives, and museums). The list of CSV files from Australian GLAM insitutions provided below [was harvested from state and national government data portals](https://glam-workbench.github.io/glam-data-portals/). You can select a file from the list or upload your own.
#
# The CSV Explorer looks at each column in the selected CSV file and tries to identify the type of data inside. It then attempts to tell you something useful about it. There's some more details about the process below.
#
# Given all the possible variations in recording and formatting data, there will be errors. But hopefully this will provide you with a useful starting point for further exploration.
# +
# This notebook is designed to run in Voila as an app (with the code hidden).
# To launch this notebook in Voila, just select 'View > Open with Voila in New Browser Tab'
# Your browser might ask for permission to open the new tab as a popup.
# -
# %%capture
import pandas as pd
from pandas.errors import ParserError
import statistics
import time
import os
import io
from urllib.parse import urlparse
from urllib.error import HTTPError
import ipywidgets as widgets
from IPython.display import display, HTML, clear_output
import altair as alt
from wordcloud import WordCloud
from slugify import slugify
# alt.renderers.enable('notebook')
#alt.data_transformers.enable('json', urlpath='files')
alt.data_transformers.enable('data_server')
#alt.data_transformers.enable('data_server_proxied', urlpath='.')
# +
#This is where the results go...
results = widgets.Output()
status = widgets.Output()
pd.set_option('display.max_columns', 10)
def read_csv(url, header=0, encoding=0):
'''
Loop through some encoding/parsing options to see if we can get the CSV to open properly.
'''
encodings = ['ISO-8859-1', 'latin-1']
headers = [None]
try:
if encoding > 0 and header > 0:
df = pd.read_csv(url, sep=None, engine='python', na_values=['-', ' '], encoding=encodings[encoding-1], header=headers[header-1])
elif encoding > 0:
df = pd.read_csv(url, sep=None, engine='python', na_values=['-', ' '], encoding=encodings[encoding-1])
elif header > 0:
df = pd.read_csv(url, sep=None, engine='python', na_values=['-', ' '], header=headers[header-1])
else:
df = pd.read_csv(url, sep=None, engine='python', na_values=['-', ' '])
except UnicodeDecodeError:
if encoding == len(encodings):
raise
else:
return read_csv(url=url, header=header, encoding=encoding+1)
except ParserError:
if header == len(headers):
raise
else:
return read_csv(url=url, header=header+1, encoding=encoding)
else:
return df
def analyse_csv(b):
'''
Try to open the CSV file, and start the analysis.
'''
results.clear_output()
status.clear_output()
error = ''
if tab.selected_index == 0:
row = csvs.loc[select_csv.value]
url = row['download_url']
title = row['file_title']
elif tab.selected_index == 1:
url = csv_url.value
parsed_url = urlparse(url)
title = os.path.basename(parsed_url.path)
elif tab.selected_index == 2:
# This will change in ipywidgets 8!
title, csv_content = list(csv_upload.value.items())[0]
url = io.BytesIO(csv_content['content'])
with results:
html = f'<hr><h2>{title}</h2>'
if tab.selected_index in [0, 1]:
html += f'<h4>Source</h4><p><a href="{url}">{url}</a> ({row["file_size"]})</p>'
display(HTML(html))
# display(status)
status.append_stdout('Downloading data...')
try:
df = read_csv(url)
except UnicodeDecodeError:
error = 'Unicode error: unable to read the CSV!'
except ParserError:
error = 'Parser error: unable to read the CSV!'
except HTTPError:
error = 'File not found!'
except:
error = 'Unable to read the CSV!'
status.clear_output()
if error:
display(HTML(f'<p class="alert alert-danger">{error}</p>'))
else:
rows, cols = df.shape
size = '<h4>Size</h4><ul>'
size += '<li>{} rows</li>'.format(rows)
size += '<li>{} columns</li></ul>'.format(cols)
cols = "<h4>Columns</h4><ol>"
for col in df.columns:
cols += '<li><a style="font-family: monospace" href="#{}" target="_self">{}</a></li>'.format(slugify(col), col)
cols += '</ol>'
display(HTML(size))
display(HTML(cols))
display(HTML('<h4>Sample</h4>'))
display(df.head())
analyse_columns(df)
# -
k, v = list({'foo': 'bar'}.items())[0]
# +
date_cutoff = 0.8
cutoff = 0.8
unique_cutoff = 0.1
category_count = 30
def display_dates(df, col):
# Better to group years first, so that the altair data isn't huge
# Get counts by year
counts = df[col].groupby([df[col].dt.year]).agg('count').to_frame()
# Get the full range of years
years = pd.Index([y for y in range(int(counts.index[0]), int(counts.index[-1]) + 1)])
# Set missing years to zero
counts = counts.reindex(years, fill_value=0)
counts = counts.reset_index()
counts.columns = [col, 'count']
chart = alt.Chart(counts).mark_area().encode(
x=alt.X(f'{col}:Q', axis=alt.Axis(format='c', title='Year', tickMinStep=1)),
y='count:Q',
tooltip=[alt.Tooltip('{}:O'.format(col), title='Year'), alt.Tooltip('count:Q', title='Count', format=',')],
color=alt.value('#5254a3')
).properties(
width=800
)
display(chart)
def display_categories(df, col):
counts = df[col].value_counts()
if counts.size > category_count:
counts = counts[:category_count].to_frame()
else:
counts = counts.to_frame()
counts = counts.reset_index()
counts.columns = [col, 'count']
chart = alt.Chart(counts).mark_bar().encode(
x='count:Q',
y=alt.Y('{}:N'.format(col), sort=alt.EncodingSortField(field='count', op='count', order='ascending')),
tooltip=[alt.Tooltip('{}:N'.format(col), title='Category'), alt.Tooltip('count:Q', title='Count', format=',')],
color=alt.value('#8ca252')
)
display(chart)
def display_wordcloud(df, col, collocates=True):
# Make a word cloud!
# The word cloud software splits the string into individual words and calculates their frquency
words = df[col].str.cat(sep=' ')
wordcloud = WordCloud(width=800, height=300, collocations=collocates).generate(words)
display(wordcloud.to_image())
def display_numbers(df, col, unique_count):
#display(df[col])
if unique_count <= 20:
# df[col].replace('0', np.NaN)
counts = df[col].value_counts().to_frame()
counts = counts.reset_index()
counts.columns = [col, 'count']
#display(counts)
chart = alt.Chart(counts).mark_bar().encode(
alt.X('{}:Q'.format(col)),
y='count',
tooltip=[alt.Tooltip('{}:Q'.format(col)), alt.Tooltip('count:Q', title='Count', format=',')],
color=alt.value('#ad494a')
)
else:
chart = alt.Chart(df).mark_bar().encode(
alt.X('{}:Q'.format(col), bin=alt.Bin(maxbins=10, nice=True)),
y='count()',
tooltip=[alt.Tooltip('{}:Q'.format(col), bin=alt.Bin(maxbins=10, nice=True), title='Range'), alt.Tooltip('count():Q', title='Count', format=',')],
color=alt.value('#ad494a')
)
display(chart)
def text_field(df, col, value_count, word_counts, details):
html = 'This looks like a text field.'
display(HTML(html))
median_word_count = statistics.median(word_counts)
collocates = True if median_word_count > 1 else False
details['Total number of words'] = word_counts.sum()
details['Highest number of words'] = word_counts.max()
details['Median number of words'] = median_word_count
details['Number of empty records'] = df[col].shape[0] - value_count
display_details(details)
wordcloud = display_wordcloud(df, col, collocates)
display(wordcloud.to_image())
#image_file = 'images/{}_cloud_{}.png'.format(slugify(col), int(time.time()))
#try:
# image.save(image_file)
#except FileNotFoundError:
# os.makedirs('images')
#display(HTML('<a href="{0}"><img src="{0}"></a>'.format(image_file)))
def textplus_field(df, col, value_count, unique_count, unique_ratio, word_counts, has_year, details, html):
median_word_count = statistics.median(word_counts)
collocates = True if median_word_count > 1 else False
mixed = False
details['Total number of words'] = word_counts.sum()
details['Highest number of words'] = word_counts.max()
details['Median number of words'] = median_word_count
details['Number of empty records'] = df[col].shape[0] - value_count
display_details(details)
has_mixed = df[col].str.contains(r'(?=\S*[a-zA-Z\/])(?=\S*[0-9])', regex=True)
if has_mixed.sum() / value_count > cutoff and median_word_count <= 2:
mixed = True
html = '<p>This columns contains a small number of words that combine letters and numbers. They\'re probably collection identifiers. Here\'s some examples:</p><ul>'
samples = df.loc[df[col].notna()][col].sample(5).to_list()
for sample in samples:
html += '<li>{}</li>'.format(sample)
html += '</ul>'
display(HTML(html))
elif unique_count <= category_count:
display(HTML(f'<p>This look like it contains categories. Let\'s look at the {category_count} most common.</p>'))
display_categories(df, col)
else:
try:
display(HTML('<p>This look like it contains text.</p>'))
wordcloud = display_wordcloud(df, col, collocates)
except ValueError:
pass
if unique_ratio < unique_cutoff:
display(HTML(f'<p>Less than {unique_cutoff:.2%} of the values are unique, let\'s look at the {category_count} most common.</p>'))
display_categories(df, col)
has_number = df[col].str.contains(r'\b\d+\b', regex=True)
# Check for dates
if has_year.sum() / value_count > cutoff and mixed is False:
html = '<p>Most of the values in this column include a number that looks like a year. It might be useful to convert them to dates.</p>'
df['{}_years_extracted'.format(col)] = df[col].str.extract(r'\b(1[7-9]{1}\d{2}|20[0-1]{1}\d{1})\b')
if df['{}_years_extracted'.format(col)].nunique(dropna=True) > 1:
df['{}_date_converted'.format(col)] = pd.to_datetime(df['{}_years_extracted'.format(col)], format='%Y', utc=True)
html += '<p>{:,} of {:,} values in this column were successfully parsed as dates.</p>'.format(df['{}_date_converted'.format(col)].dropna().size, value_count)
details = {}
details['Earliest date'] = df['{}_date_converted'.format(col)].min().strftime('%Y-%m-%d')
details['Latest date'] = df['{}_date_converted'.format(col)].max().strftime('%Y-%m-%d')
display(HTML(html))
display_details(details)
display_dates(df, '{}_date_converted'.format(col))
# Check for numbers
elif has_number.sum() / value_count > cutoff and mixed is False:
html = '<p>Most of the values in this column include a number. It might be useful to extract the values.</p>'
df['{}_numbers_extracted'.format(col)] = df[col].str.extract(r'\b(\d+)\b')
if df['{}_numbers_extracted'.format(col)].nunique(dropna=True) > 2:
df['{}_numbers_extracted'.format(col)] = pd.to_numeric(df['{}_numbers_extracted'.format(col)], errors='coerce', downcast='integer')
details = {}
details['Highest value'] = df['{}_numbers_extracted'.format(col)].max()
details['Lowest value'] = df['{}_numbers_extracted'.format(col)].dropna().min()
display(HTML(html))
display_details(details)
display_numbers(df, '{}_numbers_extracted'.format(col), unique_count)
def date_field(df, col, value_count, year_count, details, html):
default_dates = pd.to_datetime(df[col], infer_datetime_format=True, errors='coerce', utc=True)
default_dates_count = default_dates.dropna().size
dayfirst_dates = pd.to_datetime(df[col], infer_datetime_format=True, errors='coerce', dayfirst=True, yearfirst=True, utc=True)
dayfirst_dates_count = dayfirst_dates.dropna().size
if (default_dates_count / value_count > date_cutoff) and (default_dates_count >= dayfirst_dates_count):
df['{}_date_converted'.format(col)] = default_dates
elif (dayfirst_dates_count / value_count > date_cutoff) and (dayfirst_dates_count >= default_dates_count):
df['{}_date_converted'.format(col)] = dayfirst_dates
else:
# It's not a known date format, so let's just get the years
df['{}_years_extracted'.format(col)] = df[col].str.extract(r'\b(1[7-9]{1}\d{2}|20[0-1]{1}\d{1})\b')
df['{}_date_converted'.format(col)] = pd.to_datetime(df['{}_years_extracted'.format(col)], format='%Y', utc=True)
html += '<p>This looks like it contains dates.</p>'
html += '<p>{:,} of {:,} values in this column were successfully parsed as dates.</p>'.format(df['{}_date_converted'.format(col)].dropna().size, value_count)
details['Earliest date'] = df['{}_date_converted'.format(col)].min().strftime('%Y-%m-%d')
details['Latest date'] = df['{}_date_converted'.format(col)].max().strftime('%Y-%m-%d')
display(HTML(html))
display_details(details)
display_dates(df, '{}_date_converted'.format(col))
def url_field(df, col, details, html):
display_details(details)
html += '<p>It looks like this column contains urls. Here are some examples:</p><ul>'
samples = df.loc[df[col].notna()][col].sample(5).to_list()
for sample in samples:
html += '<li><a href="{0}">{0}</a></li>'.format(sample)
html += '</ul>'
display(HTML(html))
def unique_field(df, col, details, html):
display_details(details)
html += '<p>This column only contains one value:</p>'
html += '<blockquote>{}</blockquote>'.format(df[col].loc[df[col].first_valid_index()])
display(HTML(html))
def number_field(df, col, value_count, unique_count, unique_ratio, details, html):
has_year = df.loc[(df[col] >= 1700) & (df[col] <= 2019)]
if (has_year.size / value_count) > date_cutoff:
df['{}_date_converted'.format(col)] = pd.to_datetime(df[col], format='%Y', utc=True, errors='coerce')
html += '<p>This looks like it contains dates.</p>'
html += '<p>{:,} of {:,} values in this column were successfully parsed as dates.</p>'.format(df['{}_date_converted'.format(col)].dropna().size, value_count)
details['Earliest date'] = df['{}_date_converted'.format(col)].dropna().min().strftime('%Y-%m-%d')
details['Latest date'] = df['{}_date_converted'.format(col)].dropna().max().strftime('%Y-%m-%d')
display(HTML(html))
display_details(details)
display_dates(df, '{}_date_converted'.format(col))
else:
details['Highest value'] = df[col].max()
details['Lowest value'] = df[col].dropna().min()
display_details(details)
if unique_ratio > cutoff:
html = '{:.2%} of the values in this column are unique, so it\'s probably some sort of identifier.'.format(unique_ratio)
display(HTML(html))
if unique_count <= 20:
display_categories(df, col)
else:
display_numbers(df, col, unique_count)
#Check for geocoordinates?
def display_details(details):
details_df = pd.DataFrame.from_dict(details, orient='index', columns=[' '])
details_df.rename_axis('Summary', axis='columns', inplace=True)
details_df = details_df.style.set_table_styles([ dict(selector='th', props=[('text-align', 'left')] ) ])
display(details_df)
def analyse_columns(df):
enriched_df = df.copy()
#out = widgets.Output()
outputs = {}
for index, col in enumerate(enriched_df.columns):
display(HTML('<hr><h3 id="{}">{}. <code>{}</code></h3>'.format(slugify(col), index+1, col)))
details = {}
html = ''
# Are there any values in this column
value_count = enriched_df[col].dropna().size
details['Number of (non empty) values'] = '{:,} ({:.2%} of rows)'.format(value_count, (value_count / enriched_df[col].size))
if value_count:
# How many unique values are there in this column?
unique_count = enriched_df[col].nunique(dropna=True)
# What proportion of the values are unique?
unique_ratio = unique_count / value_count
details['Number of unique values'] = '{:,} ({:.2%} of non-empty values)'.format(unique_count, unique_ratio)
if unique_ratio == 1:
html += '<p>All the values in this column are unique, perhaps it''s some form of identifier.</p>'
if unique_count == 1:
unique_field(enriched_df, col, details, html)
# Check it's a string field
elif enriched_df[col].dtype == 'object':
word_counts = enriched_df[col].dropna().str.split().str.len().fillna(0)
median_word_count = statistics.median(word_counts)
# Check for the presence of years
# year_count = enriched_df[col].str.count(r'\b1[7-9]{1}\d{2}\b|\b20[0-1]{1}\d{1}\b').sum()
if enriched_df[col].str.startswith('http', na=False).sum() > 1:
url_field(enriched_df, col, details, html)
#elif median_word_count <= 4:
# How many have words that combine letters and numbers?
else:
# How many start with words (and no numbers in the first two words)?
starts_with_words = enriched_df[col].str.contains(r'^[a-zA-Z]+$|^(?:\b[a-zA-Z]{2,}\b\W*){2}', regex=True)
# How many have patterns that look like years?
has_year = enriched_df[col].str.contains(r'\b1[7-9]{1}\d{2}|20[0-1]{1}\d{1}\b', regex=True)
# If most don't start with words...
# This filters out titles or names that might include dates.
if (value_count - starts_with_words.sum()) / value_count > date_cutoff:
# If most contain years...
if (has_year.sum() / value_count) > date_cutoff:
date_field(enriched_df, col, value_count, has_year.sum(), details, html)
else:
textplus_field(enriched_df, col, value_count, unique_count, unique_ratio, word_counts, has_year, details, html)
else:
textplus_field(enriched_df, col, value_count, unique_count, unique_ratio, word_counts, has_year, details, html)
elif enriched_df[col].dtype in ['int64', 'float64']:
number_field(enriched_df, col, value_count, unique_count, unique_ratio, details, html)
else:
html = 'This column is empty.'
display(HTML(html))
# +
csvs = pd.read_csv('https://raw.githubusercontent.com/GLAM-Workbench/ozglam-data/master/glam-datasets-from-gov-portals-csvs.csv', parse_dates=['file_created'])
orgs = ['All'] + sorted(csvs['publisher'].unique().tolist())
datasets = ['All'] + sorted(csvs['dataset_title'].unique().tolist())
csvs.sort_values(by=['file_title', 'file_created'], inplace=True)
files = []
trigger = None
for row in csvs.itertuples():
files.append((f'{row.file_title} ({row.publisher}, {row.file_created.year})', row.Index))
def filter_files(field, value):
filtered_csvs = csvs.loc[csvs[field] == value]
filtered_files = []
for row in filtered_csvs.itertuples():
filtered_files.append((f'{row.file_title} ({row.publisher}, {row.file_created.year})', row.Index))
select_csv.options = filtered_files
def reset_options():
select_org.options = orgs
select_dataset.options = datasets
select_csv.options = files
select_org.value = orgs[0]
select_dataset.value = datasets[0]
#select_csv.value = files[0][1]
def filter_by_org(*args):
'''
Update the list of files in the selection dropdown based on the selected organisation.
'''
if select_org.value == 'All':
select_dataset.options = datasets
select_dataset.value = datasets[0]
select_csv.options = files
else:
filter_files('publisher', select_org.value)
if select_dataset.value != 'All':
selected_org = sorted(csvs.loc[csvs['dataset_title'] == select_dataset.value]['publisher'].unique().tolist())[0]
if selected_org != select_org.value:
filtered_datasets = ['All'] + sorted(csvs.loc[csvs['publisher'] == select_org.value]['dataset_title'].unique().tolist())
select_dataset.value = 'All'
select_dataset.options = filtered_datasets
else:
filtered_datasets = ['All'] + sorted(csvs.loc[csvs['publisher'] == select_org.value]['dataset_title'].unique().tolist())
select_dataset.value = 'All'
select_dataset.options = filtered_datasets
def filter_by_dataset(*args):
'''
Update the list of files in the selection dropdown based on the selected organisation.
'''
if select_dataset.value == 'All':
if select_org.value != 'All':
filter_files('publisher', select_org.value)
else:
filter_files('dataset_title', select_dataset.value)
selected_org = sorted(csvs.loc[csvs['dataset_title'] == select_dataset.value]['publisher'].unique().tolist())[0]
#select_org.options = filtered_orgs
if selected_org != select_org.value:
select_org.value = selected_org
def clear_all(b):
reset_options()
csv_url.value = ''
results.clear_output()
select_org = widgets.Dropdown(
options=orgs,
description='',
disabled=False,
layout=widgets.Layout(width='100%')
)
select_dataset = widgets.Dropdown(
options=datasets,
description='',
disabled=False,
layout=widgets.Layout(width='100%')
)
select_csv = widgets.Dropdown(
options=files,
description='',
disabled=False,
layout=widgets.Layout(width='100%')
)
csv_url = widgets.Text(
placeholder='Enter the url of a CSV file',
description='Url:',
disabled=False,
layout=widgets.Layout(width='100%')
)
csv_upload = widgets.FileUpload(
accept='.csv',
multiple=False
)
clear_button = widgets.Button(
description='Clear',
disabled=False,
button_style='', # 'success', 'info', 'warning', 'danger' or ''
tooltip='Clear current data',
icon=''
)
analyse_button = widgets.Button(
description='Analyse CSV',
disabled=False,
button_style='primary', # 'success', 'info', 'warning', 'danger' or ''
tooltip='Analyse CSV',
icon=''
)
# Update the file list when you select an org
select_org.observe(filter_by_org)
# Update the file list when you select an org
select_dataset.observe(filter_by_dataset)
clear_button.on_click(clear_all)
analyse_button.on_click(analyse_csv)
select_org_note = widgets.HTML('Filter by organisation:')
select_dataset_note = widgets.HTML('Filter by dataset:')
select_note = widgets.HTML('Select a CSV file:')
select_tab = widgets.VBox([select_note, select_csv, select_org_note, select_org, select_dataset_note, select_dataset])
tab = widgets.Tab(children=[select_tab, csv_url, csv_upload])
tab.set_title(0, 'Select CSV')
tab.set_title(1, 'Enter CSV url')
tab.set_title(2, 'Upload CSV')
display(widgets.VBox([tab, widgets.HBox([analyse_button, clear_button]), results, status]))
# -
# ----
#
# ## More information
#
# The GLAM CSV Explorer is a [Jupyter](http://jupyter.org/) notebook, combining live Python code with text and widgets in a form that's easy to hack and build upon. The app makes heavy use of [Pandas](https://pandas.pydata.org/), the all-purpose toolkit for working with tabular data. Pandas is quick and powerful, but has so many options it can be difficult to know where to start. You might like to poke around in the code for ideas.
#
# To analyse a CSV, the explorer looks at things like the datatype of a column, and the number of unique values it holds. It also applies a variety of regular expressions to look for dates and numbers. Depending on what it finds, it extracts some summary information, and tries to visualise the results using [WordCloud](https://github.com/amueller/word_cloud) and [Altair](https://altair-viz.github.io/index.html).
# ----
#
# Created by [<NAME>](https://timsherratt.org/) for the [GLAM Workbench](https://glam-workbench.net/). Support me by becoming a [GitHub sponsor](https://github.com/sponsors/wragge)!
#
# Work on this notebook was supported by the [Humanities, Arts and Social Sciences (HASS) Data Enhanced Virtual Lab](https://tinker.edu.au/).
|
csv-explorer.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="4GDelxAZefoW" outputId="3de89a52-846a-45e3-8c6b-b1242636e753"
x=int(input("Enter your salary"))
if x>250000 :
print("you have to pay tax ")
if x<250000 :
print("you do not have to pay the tax")
# + id="fT0okQc8fJXV"
|
Python_basics.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Variant effect prediction
# Variant effect prediction offers a simple way to predict effects of SNVs using any model that uses DNA sequence as an input. Many different scoring methods can be chosen, but the principle relies on in-silico mutagenesis. The default input is a VCF and the default output again is a VCF annotated with predictions of variant effects.
#
# For details please take a look at the documentation in Postprocessing/Variant effect prediction. This iPython notebook goes through the basic programmatic steps that are needed to preform variant effect prediction. First a variant-centered approach will be taken and secondly overlap-based variant effect prediction will be presented. For details in how this is done programmatically, please refer to the documentation.
# ## Variant centered effect prediction
# Models that accept input `.bed` files can make use of variant-centered effect prediction. This procedure starts out from the query VCF and generates genomic regions of the length of the model input, centered on the individual variant in the VCF.The model dataloader is then used to produce the model input samples for those regions, which are then mutated according to the alleles in the VCF:
#
# 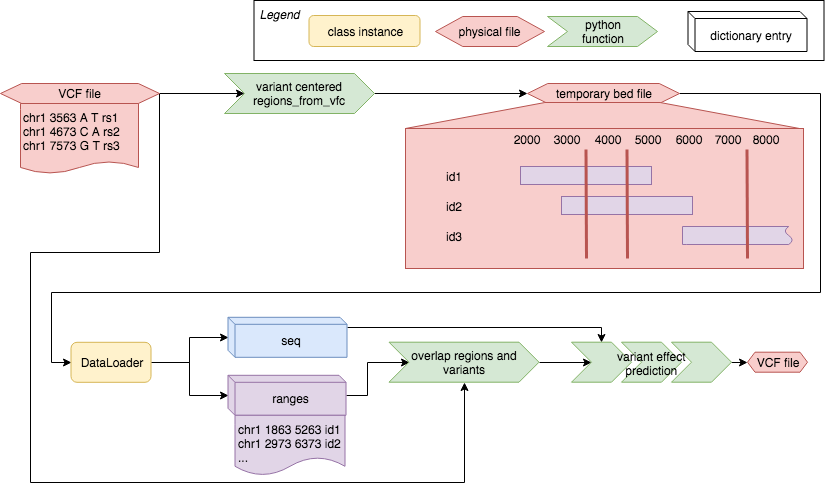
#
# First an instance of `SnvCenteredRg` generates a temporary bed file with regions matching the input sequence length defined in the model.yaml input schema. Then the model dataloader is used to preduce the model input in batches. These chunks of data are then modified by the effect prediction algorithm, the model batch prediction function is triggered for all mutated sequence sets and finally the scoring method is applied.
#
# The selected scoring methods compare model predicitons for sequences carrying the reference or alternative allele. Those scoring methods can be `Diff` for simple subtraction of prediction, `Logit` for substraction of logit-transformed model predictions, or `DeepSEA_effect` which is a combination of `Diff` and `Logit`, which was published in the Troyanskaya et al. (2015) publication.
#
# This ipython notebook assumes that it is executed in an environment in which all dependencies for the following models are installed: `DeepSEA/vaariantEffects`, `HAL`, `labranchor`, `MaxEntScan`, and `rbp` are installed, as well as the `--vep` flag has to be used during installing the dependencies. Now let's start out by loading the DeepSEA model and dataloader factory:
import kipoi
model_name = "DeepSEA/variantEffects"
# get the model
model = kipoi.get_model(model_name)
# get the dataloader factory
Dataloader = model.default_dataloader
# Next we will have to define the variants we want to look at, let's look at a sample VCF in chromosome 22:
# !head -n 40 example_data/clinvar_donor_acceptor_chr22.vcf
# Now we will define path variable for vcf input and output paths and instantiate a VcfWriter, which will write out the annotated VCF:
import kipoi_veff
from kipoi_veff import VcfWriter
# The input vcf path
vcf_path = "example_data/clinvar_donor_acceptor_chr22.vcf"
# The output vcf path, based on the input file name
out_vcf_fpath = vcf_path[:-4] + "%s.vcf"%model_name.replace("/", "_")
# The writer object that will output the annotated VCF
writer = VcfWriter(model, vcf_path, out_vcf_fpath)
# Then we need to instantiate an object that can generate variant-centered regions (`SnvCenteredRg` objects). This class needs information on the model input sequence length which is extracted automatically within `ModelInfoExtractor` objects:
# Information extraction from dataloader and model
model_info = kipoi_veff.ModelInfoExtractor(model, Dataloader)
# vcf_to_region will generate a variant-centered regions when presented a VCF record.
vcf_to_region = kipoi_veff.SnvCenteredRg(model_info)
# Now we can define the required dataloader arguments, omitting the `intervals_file` as this will be replaced by the automatically generated bed file:
dataloader_arguments = {"fasta_file": "example_data/hg19_chr22.fa"}
# This is the moment to run the variant effect prediction:
import kipoi_veff.snv_predict as sp
from kipoi_veff.scores import Diff, DeepSEA_effect
sp.predict_snvs(model,
Dataloader,
vcf_path,
batch_size = 32,
dataloader_args=dataloader_arguments,
vcf_to_region=vcf_to_region,
evaluation_function_kwargs={'diff_types': {'diff': Diff("mean"), 'deepsea_effect': DeepSEA_effect("mean")}},
sync_pred_writer=writer)
# In the example above we have used the variant scoring method `Diff` and `DeepSEA_effect` from `kipoi_veff` plug-in. As mentioned above variant scoring methods calculate the difference between predictions for reference and alternative, but there is another dimension to this: Models that have the `use_rc: true` flag set in their model.yaml file (DeepSEA/variantEffects does) will not only be queried with the reference and alternative carrying input sequences, but also with the reverse complement of the the sequences. In order to know of to combine predictions for forward and reverse sequences there is a initialisation flag (here set to: `"mean"`) for the variant scoring methods. `"mean"` in this case means that after calculating the effect (e.g.: Difference) the average over the difference between the prediction for the forward and for the reverse sequence should be returned. Setting `"mean"` complies with what was used in the Troyanskaya et al. publication.
#
# Now let's look at the output:
# Let's print out the first 40 lines of the annotated VCF (up to 80 characters per line maximum)
with open("example_data/clinvar_donor_acceptor_chr22DeepSEA_variantEffects.vcf") as ifh:
for i,l in enumerate(ifh):
long_line = ""
if len(l)>80:
long_line = "..."
print(l[:80].rstrip() +long_line)
if i >=40:
break
# This shows that variants have been annotated with variant effect scores. For every different scoring method a different INFO tag was created and the score of every model output is concantenated with the `|` separator symbol. A legend is given in the header section of the VCF. The name tag indicates with model was used, wich version of it and it displays the scoring function label (`DIFF`) which is derived from the scoring function label defined in the `evaluation_function_kwargs` dictionary (`'diff'`).
# The most comprehensive representation of effect preditions is in the annotated VCF. Kipoi offers a VCF parser class that enables simple parsing of annotated VCFs:
# +
from kipoi_veff.parsers import KipoiVCFParser
vcf_reader = KipoiVCFParser("example_data/clinvar_donor_acceptor_chr22DeepSEA_variantEffects.vcf")
#We can have a look at the different labels which were created in the VCF
print(list(vcf_reader.kipoi_parsed_colnames.values()))
# -
# We can see that two scores have been saved - `'DEEPSEA_EFFECT'` and `'DIFF'`. Additionally there is `'rID'` which is the region ID - that is the ID given by the dataloader for a genomic region which was overlapped with the variant to get the prediction that is listed in the effect score columns mentioned before. Let's take a look at the VCF entries:
import pandas as pd
entries = [el for el in vcf_reader]
print(pd.DataFrame(entries).head().iloc[:,:7])
# Another way to access effect predicitons programmatically is to keep all the results in memory and receive them as a dictionary of pandas dataframes:
#
effects = sp.predict_snvs(model,
Dataloader,
vcf_path,
batch_size = 32,
dataloader_args=dataloader_arguments,
vcf_to_region=vcf_to_region,
evaluation_function_kwargs={'diff_types': {'diff': Diff("mean"), 'deepsea_effect': DeepSEA_effect("mean")}},
return_predictions=True)
# For every key in the `evaluation_function_kwargs` dictionary there is a key in `effects` and (the equivalent of an additional INFO tag in the VCF). Now let's take a look at the results:
for k in effects:
print(k)
print(effects[k].head().iloc[:,:4])
print("-"*80)
# We see that for `diff` and `deepsea_effect` there is a dataframe with variant identifiers as rows and model output labels as columns. The DeepSEA model predicts 919 tasks simultaneously hence there are 919 columns in the dataframe.
# ## Overlap based prediction
# Models that cannot predict on every region of the genome might not accept a `.bed` file as dataloader input. An example of such a model is a splicing model. Those models only work in certain regions of the genome. Here variant effect prediction can be executed based on overlaps between the regions generated by the dataloader and the variants defined in the VCF:
#
# 
# The procedure is similar to the variant centered effect prediction explained above, but in this case no temporary bed file is generated and the effect prediction is based on all the regions generated by the dataloader which overlap any variant in the VCF. If a region is overlapped by two variants the effect of the two variants is predicted independently.
#
# Here the VCF has to be tabixed so that a regional lookup can be performed efficiently, this can be done by using the `ensure_tabixed` function, the rest remains the same as before:
# +
import kipoi
from kipoi_veff import VcfWriter
from kipoi_veff import ensure_tabixed_vcf
# Use a splicing model
model_name = "HAL"
# get the model
model = kipoi.get_model(model_name)
# get the dataloader factory
Dataloader = kipoi.get_dataloader_factory(model_name)
# The input vcf path
vcf_path = "example_data/clinvar_donor_acceptor_chr22.vcf"
# Make sure that the vcf is bgzipped and tabixed, if not then generate the compressed vcf in the same place
vcf_path_tbx = ensure_tabixed_vcf(vcf_path)
# The output vcf path, based on the input file name
out_vcf_fpath = vcf_path[:-4] + "%s.vcf"%model_name.replace("/", "_")
# The writer object that will output the annotated VCF
writer = VcfWriter(model, vcf_path, out_vcf_fpath)
# -
# This time we don't need an object that generates regions, hence we can directly define the dataloader arguments and run the prediction:
# +
from kipoi_veff import predict_snvs
from kipoi_veff.scores import Diff
dataloader_arguments = {"gtf_file":"example_data/Homo_sapiens.GRCh37.75.filtered_chr22.gtf",
"fasta_file": "example_data/hg19_chr22.fa"}
effects = predict_snvs(model,
Dataloader,
vcf_path_tbx,
batch_size = 32,
dataloader_args=dataloader_arguments,
evaluation_function_kwargs={'diff_types': {'diff': Diff("mean")}},
sync_pred_writer=writer,
return_predictions=True)
# -
# Let's have a look at the VCF:
# A slightly convoluted way of printing out the first 40 lines and up to 80 characters per line maximum
with open("example_data/clinvar_donor_acceptor_chr22HAL.vcf") as ifh:
for i,l in enumerate(ifh):
long_line = ""
if len(l)>80:
long_line = "..."
print(l[:80].rstrip() +long_line)
if i >=40:
break
# And the prediction output this time is less helpful because it's the ids that the dataloader created which are displayed as index. In general it is advisable to use the output VCF for more detailed information on which variant was overlapped with which region fo produce a prediction.
for k in effects:
print(k)
print(effects[k].head())
print("-"*80)
# ## Command-line based effect prediction
# The above command can also conveniently be executed using the command line:
#
import json
import os
model_name = "DeepSEA/variantEffects"
dl_args = json.dumps({"fasta_file": "example_data/hg19_chr22.fa"})
out_vcf_fpath = vcf_path[:-4] + "%s.vcf"%model_name.replace("/", "_")
scorings = "diff deepsea_effect"
command = ("kipoi veff score_variants {model} "
"--dataloader_args='{dl_args}' "
"-i {input_vcf} "
"-o {output_vcf} "
"-s {scorings}").format(model=model_name,
dl_args=dl_args,
input_vcf=vcf_path,
output_vcf=out_vcf_fpath,
scorings=scorings)
# Print out the command:
print(command)
# ! $command
# ## Batch prediction
# Since the syntax basically doesn't change for different kinds of models a simple for-loop can be written to do what we just did on many models:
import kipoi
# Run effect predicton
models_df = kipoi.list_models()
models_substr = ["HAL", "MaxEntScan", "labranchor", "rbp"]
models_df_subsets = {ms: models_df.loc[models_df["model"].str.contains(ms)] for ms in models_substr}
# +
# Run variant effect prediction using a basic Diff
import kipoi
from kipoi_veff import ensure_tabixed_vcf
import kipoi_veff.snv_predict as sp
from kipoi_veff import VcfWriter
from kipoi_veff.scores import Diff
splicing_dl_args = {"gtf_file":"example_data/Homo_sapiens.GRCh37.75.filtered_chr22.gtf",
"fasta_file": "example_data/hg19_chr22.fa"}
dataloader_args_dict = {"HAL": splicing_dl_args,
"labranchor": splicing_dl_args,
"MaxEntScan":splicing_dl_args,
"rbp": {"fasta_file": "example_data/hg19_chr22.fa",
"gtf_file":"example_data/Homo_sapiens.GRCh37.75_chr22.gtf"}
}
for ms in models_substr:
model_name = models_df_subsets[ms]["model"].iloc[0]
#kipoi.pipeline.install_model_requirements(model_name)
model = kipoi.get_model(model_name)
vcf_path = "example_data/clinvar_donor_acceptor_chr22.vcf"
vcf_path_tbx = ensure_tabixed_vcf(vcf_path)
out_vcf_fpath = vcf_path[:-4] + "%s.vcf"%model_name.replace("/", "_")
writer = VcfWriter(model, vcf_path, out_vcf_fpath)
print(model_name)
Dataloader = model.default_dataloader
dataloader_arguments = dataloader_args_dict[ms]
model_info = kipoi_veff.ModelInfoExtractor(model, Dataloader)
vcf_to_region = None
if ms == "rbp":
vcf_to_region = kipoi_veff.SnvCenteredRg(model_info)
sp.predict_snvs(model,
Dataloader,
vcf_path_tbx,
batch_size = 32,
dataloader_args=dataloader_arguments,
vcf_to_region=vcf_to_region,
evaluation_function_kwargs={'diff_types': {'diff': Diff("mean")}},
sync_pred_writer=writer)
writer.close()
# -
# let's validate that things have worked:
# ! wc -l example_data/clinvar_donor_acceptor_chr22*.vcf
|
notebooks/variant_effect_prediction.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ___
#
# <a href='http://www.pieriandata.com'> <img src='../Pierian_Data_Logo.png' /></a>
# ___
# # Random Forest Project
#
# For this project we will be exploring publicly available data from [LendingClub.com](www.lendingclub.com). Lending Club connects people who need money (borrowers) with people who have money (investors). Hopefully, as an investor you would want to invest in people who showed a profile of having a high probability of paying you back. We will try to create a model that will help predict this.
#
# Lending club had a [very interesting year in 2016](https://en.wikipedia.org/wiki/Lending_Club#2016), so let's check out some of their data and keep the context in mind. This data is from before they even went public.
#
# We will use lending data from 2007-2010 and be trying to classify and predict whether or not the borrower paid back their loan in full. You can download the data from [here](https://www.lendingclub.com/info/download-data.action) or just use the csv already provided. It's recommended you use the csv provided as it has been cleaned of NA values.
#
# Here are what the columns represent:
# * credit.policy: 1 if the customer meets the credit underwriting criteria of LendingClub.com, and 0 otherwise.
# * purpose: The purpose of the loan (takes values "credit_card", "debt_consolidation", "educational", "major_purchase", "small_business", and "all_other").
# * int.rate: The interest rate of the loan, as a proportion (a rate of 11% would be stored as 0.11). Borrowers judged by LendingClub.com to be more risky are assigned higher interest rates.
# * installment: The monthly installments owed by the borrower if the loan is funded.
# * log.annual.inc: The natural log of the self-reported annual income of the borrower.
# * dti: The debt-to-income ratio of the borrower (amount of debt divided by annual income).
# * fico: The FICO credit score of the borrower.
# * days.with.cr.line: The number of days the borrower has had a credit line.
# * revol.bal: The borrower's revolving balance (amount unpaid at the end of the credit card billing cycle).
# * revol.util: The borrower's revolving line utilization rate (the amount of the credit line used relative to total credit available).
# * inq.last.6mths: The borrower's number of inquiries by creditors in the last 6 months.
# * delinq.2yrs: The number of times the borrower had been 30+ days past due on a payment in the past 2 years.
# * pub.rec: The borrower's number of derogatory public records (bankruptcy filings, tax liens, or judgments).
# # Import Libraries
#
# **Import the usual libraries for pandas and plotting. You can import sklearn later on.**
# ## Get the Data
#
# ** Use pandas to read loan_data.csv as a dataframe called loans.**
# ** Check out the info(), head(), and describe() methods on loans.**
# # Exploratory Data Analysis
#
# Let's do some data visualization! We'll use seaborn and pandas built-in plotting capabilities, but feel free to use whatever library you want. Don't worry about the colors matching, just worry about getting the main idea of the plot.
#
# ** Create a histogram of two FICO distributions on top of each other, one for each credit.policy outcome.**
#
# *Note: This is pretty tricky, feel free to reference the solutions. You'll probably need one line of code for each histogram, I also recommend just using pandas built in .hist()*
# ** Create a similar figure, except this time select by the not.fully.paid column.**
# ** Create a countplot using seaborn showing the counts of loans by purpose, with the color hue defined by not.fully.paid. **
# ** Let's see the trend between FICO score and interest rate. Recreate the following jointplot.**
# ** Create the following lmplots to see if the trend differed between not.fully.paid and credit.policy. Check the documentation for lmplot() if you can't figure out how to separate it into columns.**
# # Setting up the Data
#
# Let's get ready to set up our data for our Random Forest Classification Model!
#
# **Check loans.info() again.**
# ## Categorical Features
#
# Notice that the **purpose** column as categorical
#
# That means we need to transform them using dummy variables so sklearn will be able to understand them. Let's do this in one clean step using pd.get_dummies.
#
# Let's show you a way of dealing with these columns that can be expanded to multiple categorical features if necessary.
#
# **Create a list of 1 element containing the string 'purpose'. Call this list cat_feats.**
# **Now use pd.get_dummies(loans,columns=cat_feats,drop_first=True) to create a fixed larger dataframe that has new feature columns with dummy variables. Set this dataframe as final_data.**
# ## Train Test Split
#
# Now its time to split our data into a training set and a testing set!
#
# ** Use sklearn to split your data into a training set and a testing set as we've done in the past.**
# ## Training a Decision Tree Model
#
# Let's start by training a single decision tree first!
#
# ** Import DecisionTreeClassifier**
from sklearn.tree import DecisionTreeClassifier
# **Create an instance of DecisionTreeClassifier() called dtree and fit it to the training data.**
# ## Predictions and Evaluation of Decision Tree
# **Create predictions from the test set and create a classification report and a confusion matrix.**
# ## Training the Random Forest model
#
# Now its time to train our model!
#
# **Create an instance of the RandomForestClassifier class and fit it to our training data from the previous step.**
# ## Predictions and Evaluation
#
# Let's predict off the y_test values and evaluate our model.
#
# ** Predict the class of not.fully.paid for the X_test data.**
# **Now create a classification report from the results. Do you get anything strange or some sort of warning?**
# **Show the Confusion Matrix for the predictions.**
# **What performed better the random forest or the decision tree?**
# # Great Job!
|
Udemy/Refactored_Py_DS_ML_Bootcamp-master/15-Decision-Trees-and-Random-Forests/02-Decision Trees and Random Forest Project.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Sweeps - Capacitance matrix
# ### Prerequisite
# You need to have a working local installation of Ansys
# ## 1. Perform the necessary imports and create a QDesign in Metal first.
# %load_ext autoreload
# %autoreload 2
import qiskit_metal as metal
from qiskit_metal import designs, draw
from qiskit_metal import MetalGUI, Dict, Headings
from qiskit_metal.analyses.quantization import LOManalysis
from qiskit_metal.renderers.renderer_ansys.ansys_renderer import QAnsysRenderer
# +
design = designs.DesignPlanar()
gui = MetalGUI(design)
from qiskit_metal.qlibrary.qubits.transmon_pocket import TransmonPocket
# -
design.variables['cpw_width'] = '15 um'
design.variables['cpw_gap'] = '9 um'
# ### In this example, the design consists of 4 qubits and 4 CPWs.
# +
# Allow running the same cell here multiple times to overwrite changes
design.overwrite_enabled = True
## Custom options for all the transmons
options = dict(
# Some options we want to modify from the defaults
# (see below for defaults)
pad_width = '425 um',
pocket_height = '650um',
# Adding 4 connectors (see below for defaults)
connection_pads=dict(
readout = dict(loc_W=+1,loc_H=-1, pad_width='200um'),
bus1 = dict(loc_W=-1,loc_H=+1, pad_height='30um'),
bus2 = dict(loc_W=-1,loc_H=-1, pad_height='50um')
)
)
## Create 4 transmons
q1 = TransmonPocket(design, 'Q1', options = dict(
pos_x='+2.42251mm', pos_y='+0.0mm', **options))
q2 = TransmonPocket(design, 'Q2', options = dict(
pos_x='+0.0mm', pos_y='-0.95mm', orientation = '270', **options))
q3 = TransmonPocket(design, 'Q3', options = dict(
pos_x='-2.42251mm', pos_y='+0.0mm', orientation = '180', **options))
q4 = TransmonPocket(design, 'Q4', options = dict(
pos_x='+0.0mm', pos_y='+0.95mm', orientation = '90', **options))
from qiskit_metal.qlibrary.tlines.meandered import RouteMeander
RouteMeander.get_template_options(design)
options = Dict(
lead=Dict(
start_straight='0.2mm',
end_straight='0.2mm'),
trace_gap='9um',
trace_width='15um')
def connect(component_name: str, component1: str, pin1: str, component2: str, pin2: str,
length: str, asymmetry='0 um', flip=False, fillet='90um'):
"""Connect two pins with a CPW."""
myoptions = Dict(
fillet=fillet,
hfss_wire_bonds = True,
pin_inputs=Dict(
start_pin=Dict(
component=component1,
pin=pin1),
end_pin=Dict(
component=component2,
pin=pin2)),
total_length=length)
myoptions.update(options)
myoptions.meander.asymmetry = asymmetry
myoptions.meander.lead_direction_inverted = 'true' if flip else 'false'
return RouteMeander(design, component_name, myoptions)
asym = 140
cpw1 = connect('cpw1', 'Q1', 'bus2', 'Q2', 'bus1', '5.6 mm', f'+{asym}um')
cpw2 = connect('cpw2', 'Q3', 'bus1', 'Q2', 'bus2', '5.7 mm', f'-{asym}um', flip=True)
cpw3 = connect('cpw3', 'Q3', 'bus2', 'Q4', 'bus1', '5.6 mm', f'+{asym}um')
cpw4 = connect('cpw4', 'Q1', 'bus1', 'Q4', 'bus2', '5.7 mm', f'-{asym}um', flip=True)
gui.rebuild()
gui.autoscale()
# -
# ## 2 Metal passes information to the simulator "q3d" to extract the capacitance matrix.
#
c1 = LOManalysis(design, "q3d")
# Prepare data to pass as arguments for method run_sweep().
# +
render_design_argument_qcomps = ['Q1']
render_design_argument_endcaps = [('Q1', 'readout'), ('Q1', 'bus1'),('Q1', 'bus2')]
# -
# To identify the agruments that you can change.
# They will change based on the simulation software used.
c1.sim.setup
# +
# For the simulation software, if you don't want to use the default values,
# you can update them as seen below.
# If a setup named "sweeper_q3d_setup" exists in the project, it will be deleted,
# and a new setup will be added.
c1.sim.setup.name = "sweeper_q3d_setup"
c1.sim.setup.freq_ghz = 5.6
c1.sim.setup.max_passes = 9
c1.sim.setup.min_passes = 2
c1.sim.setup.percent_error = 0.45
# -
#
#
# We will look at modifying the pad_gap of qubit 1, to see how it impacts the anharmonicity of the qubit.
#
# The steps will be;
# - Connect to Ansys Q3D.
# - Rebuild QComponents in Metal.
# - Render QComponents within Q3D and setup the simulation.
# - Delete/Clear the Q3D between each simulation.
# - Using the capacitance matrices, LOM for each value in option_sweep is found.
#
# #### Returns a dict and return code. If the return code is zero, there were no errors detected.
# #### The dict has: key = each value used to sweep, value = capacitance matrix
#
# #### This could take minutes or hours based on the complexity of the design.
#
#
sweep_data, return_code = c1.run_sweep(q1.name,
'pad_gap',
['20um', '30um', '40um'],
render_design_argument_qcomps,
render_design_argument_endcaps,
design_name="GetCapacitance",
box_plus_buffer=True)
# +
from pandas import DataFrame
ec_val = []
for opt_val in sweep_data.keys():
ec_val.append([opt_val,sweep_data[opt_val]['variables']['lumped_oscillator']['EC']])
df=DataFrame(ec_val,columns = ['Sweep Value', 'Ec'])
df
# -
# We can grab specific values from the results as seen below;
sweep_data.keys()
# For each value of option, there is a set of data.
sweep_data['20um'].keys()
sweep_data['20um']['variables'].keys()
sweep_data['20um']['sim_variables'].keys()
# +
if return_code ==0:
print(f'{sweep_data.keys()} \n')
for key in sweep_data.keys():
print(f'\nkey={key}')
option_name = sweep_data[key]['option_name']
print(f'option_name[\'{key}\'][\'option_name\']={option_name}')
variables = sweep_data[key]['variables']
sim_variables = sweep_data[key]['sim_variables']
print(f'variables={variables}')
print(f'sim_variables={sim_variables}')
# -
# # READ THIS BEFORE running the cell.
# This cell is to demonstrate that if you have already executed c1.sim.run(), you don't have to set up
# the environment again for c1.run_sweep(). In another words, if you don't pass updated arguments to
# c1.run_sweep(), then c1.run_sweep() looks for the previous desgin arguments.
#
# If you pass anything more than these three arguments: qcomp_name, option_name, option_sweep .....
# Then NOTHING will be used from previous run.
# ```
# c1.sim.solution_order = 'Medium'
# c1.sim.auto_increase_solution_order = 'False'
#
#
# c1.sim.run(components=render_design_argument_qcomps,
# open_terminations=render_design_argument_endcaps)
# ```
#
# Because c1.sim.setup.run has the information from last run, this is OK.
#
# ```
# sweep_data, return_code = c1.run_sweep(q1.name,
# 'pad_gap',
# ['20um', '30um', '40um'])
# ```
c1.sim.close()
# Uncomment next line if you would like to close the gui
gui.main_window.close()
|
tutorials/4 Analysis/C. Parametric sweeps/4.21 Capacitance matrix.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # EPS VGEO X-ray 18MV water phantom simulation
#
# `Geant4 v10.7.3`
#
# - AMD EPYC 7643 (2.3GHz / 3.6GHz) 48 cores x 2 SMT : AlmaLinux 8.5
# - AMD Ryzen Threadripper PRO 3995WX (2.7GHz / 4.2GHz) 64 cores x 1 SMT : Ubuntu 20.04LTS
# - Intel Xeon Gold 6240 (2.6GHz / 3.9GHz) 18 cores x 2 HT : Ubuntu 20.04LTS
# - Intel Core i9-12900K (3.2GHz / 5.2GHz + 2.4GHz / 3.9GHz) 8 cores(P) HT + 8 cores (E) : Ubuntu 20.04LTS
# - Apple M1 4 cores (P) + 4 cores (E) (ARMv8) : macOS Monterey 12.1
# +
import pandas
import json
import os
data_dir = "./data/10.7.3/"
cpu_list = os.listdir(data_dir)
cpu_list
# -
ecal_df = []
for cpu in cpu_list:
ecal_df.append(pandas.read_json(data_dir + cpu + '/vgeo_x18.json'))
ecal_df[-1] = ecal_df[-1].sort_values('thread')
import matplotlib.pyplot as plt
import numpy
plt.figure(figsize=(16,8))
plt.title("EPS (Ecal e1000)")
plt.xlabel("#threads")
plt.ylabel("EPS (events/msec)")
plt.xlim([0,200])
plt.ylim([0,400000])
plt.grid(which="both", color="#cccccc")
plot = plt.plot(ecal_df[1]['thread'], ecal_df[1]['eps']*1000, 'o-', markersize=8, label='AMD EPYC 7643 (48cx2 SMT)')
plot = plt.plot(ecal_df[3]['thread'], ecal_df[3]['eps']*1000, 'o-', markersize=8, label='AMD Ryzen Threadripper Pro 3995WX (64c SMT)')
plot = plt.plot(ecal_df[4]['thread'], ecal_df[4]['eps']*1000, 'o-', markersize=8, label='Intel Xeon Gold 6240 (18cx2 HT)')
plot = plt.plot(ecal_df[2]['thread'], ecal_df[2]['eps']*1000, 'o-', markersize=8, label='Intel Core-i9 12900K (8H/HP+4E)')
plot = plt.plot(ecal_df[0]['thread'], ecal_df[0]['eps']*1000, 'o-', markersize=8, label='Apple M1 (4H+4E)')
plt.legend()
|
bench/plots/multi-thread/eps-vgeo-x18.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: master_thesis
# language: python
# name: master_thesis
# ---
# # Stablecoin Billionaires<br> Descriptive Analysis of the Ethereum-based Stablecoin ecosystem
# ## by <NAME>, 01.07.2020
# # Analytics Part of the thesis
import numpy as np
import pandas as pd
import matplotlib
import matplotlib.pyplot as plt
from datetime import datetime
from collections import Counter
from matplotlib import rc
# +
#plots
exchanges = 'plots/exchanges.csv'
dai_tx_over_date = 'plots/dai/dai_txs_over_date.csv'
dai_balances = 'plots/dai/dai_balances.csv'
dai_positive_cumulated_balances = 'plots/dai/dai_positive_cumulated_balances.csv'
dai_circulating_supply = 'plots/dai/dai_circulating_supply.csv'
tether_tx_over_date = 'plots/tether/tether_txs_over_date.csv'
tether_balances = 'plots/tether/tether_balances.csv'
tether_positive_cumulated_balances = 'plots/tether/tether_positive_cumulated_balances.csv'
tether_circulating_supply = 'plots/tether/tether_circulating_supply.csv'
tether_chunk5 = 'data/tether/transfer/5_tether_transfer_10176691-10370273.csv'
usdc_tx_over_date = 'plots/usdc/usdc_txs_over_date.csv'
usdc_balances = 'plots/usdc/usdc_balances.csv'
usdc_positive_cumulated_balances = 'plots/usdc/usdc_positive_cumulated_balances.csv'
usdc_circulating_supply = 'plots/usdc/usdc_circulating_supply.csv'
paxos_tx_over_date = 'plots/paxos/paxos_txs_over_date.csv'
paxos_balances = 'plots/paxos/paxos_balances.csv'
paxos_positive_cumulated_balances = 'plots/paxos/paxos_positive_cumulated_balances.csv'
paxos_circulating_supply = 'plots/paxos/paxos_circulating_supply.csv'
paxos_tx_count_to = 'plots/paxos/paxos_tx_count_to.csv'
paxos_tx_count_from = 'plots/paxos/paxos_tx_count_from.csv'
trueusd_tx_over_date = 'plots/trueusd/trueusd_txs_over_date.csv'
trueusd_balances = 'plots/trueusd/trueusd_balances.csv'
trueusd_positive_cumulated_balances = 'plots/trueusd/trueusd_positive_cumulated_balances.csv'
trueusd_circulating_supply = 'plots/trueusd/trueusd_circulating_supply.csv'
binanceusd_tx_over_date = 'plots/binanceusd/binanceusd_txs_over_date.csv'
binanceusd_balances = 'plots/binanceusd/binanceusd_balances.csv'
binanceusd_positive_cumulated_balances = 'plots/binanceusd/binanceusd_positive_cumulated_balances.csv'
binanceusd_circulating_supply = 'plots/binanceusd/binanceusd_circulating_supply.csv'
husd_tx_over_date = 'plots/husd/husd_txs_over_date.csv'
husd_balances = 'plots/husd/husd_balances.csv'
husd_positive_cumulated_balances = 'plots/husd/husd_positive_cumulated_balances.csv'
husd_circulating_supply = 'plots/husd/husd_circulating_supply.csv'
#data
#tether
tether_transfer = 'data/tether/transfer/tether_transfers.csv'
tether_mint = 'data/tether/issue/tether_issue.csv'
tether_burn = 'data/tether/destroyedblackfunds/tether_destroyedblackfunds.csv'
#usdc
usdc_transfer = 'data/usdc/transfer/0_usdc_transfer_6082465-10370273.csv'
usdc_mint = 'data/usdc/mint/usdc_mint.csv'
usdc_burn = 'data/usdc/burn/usdc_burn.csv'
#paxos
paxos_transfer = 'data/paxos/transfer/0_paxos_transfer_6294931-10370273.csv'
paxos_mint = 'data/paxos/supplyincreased/paxos_supplyincreased.csv'
paxos_burn = 'data/paxos/supplydecreased/paxos_supplydecreased.csv'
#dai
dai_transfer = 'data/dai/transfer/0_dai_transfer_8928158-10370273.csv'
dai_mint = 'data/dai/mint/dai_mint.csv'
dai_burn = 'data/dai/burn/dai_burn.csv'
#trueusd
trueusd_transfer = 'data/trueusd/transfer/0_trueUSD_transfer_5198636-10370273.csv'
trueusd_mint = 'data/trueusd/mint/trueusd_mint.csv'
trueusd_burn = 'data/trueusd/burn/trueusd_burn.csv'
#binance usd
binanceusd_transfer = 'data/binanceusd/transfer/0_binanceusd_transfer_8493105-10370273.csv'
binanceusd_mint = 'data/binanceusd/supplyincreased/binanceusd_supplyincreased.csv'
binanceusd_burn = 'data/binanceusd/supplydecreased/binanceusd_supplydecreased.csv'
#husd
husd_transfer = 'data/husd/transfer/0_husd_transfer_8174400-10370273.csv'
husd_mint = 'data/husd/issue/husd_issue.csv'
husd_burn = 'data/husd/redeem/husd_redeem.csv'
_all_transfer = [tether_tx_over_date, usdc_tx_over_date, paxos_tx_over_date, dai_tx_over_date, trueusd_tx_over_date, binanceusd_tx_over_date, husd_tx_over_date]
_all = [usdc_transfer,usdc_mint,
usdc_burn,paxos_transfer,paxos_mint,paxos_burn,dai_transfer,tether_transfer,tether_mint,tether_burn,
dai_mint,dai_burn,trueusd_transfer,trueusd_mint,trueusd_burn,
binanceusd_transfer,binanceusd_mint,binanceusd_burn,husd_transfer,husd_mint,husd_burn]
_all_tx = [usdc_transfer,paxos_transfer,dai_transfer,tether_transfer,
trueusd_transfer,binanceusd_transfer,husd_transfer]
_all_mint = [usdc_mint,paxos_mint,dai_mint,tether_mint,
trueusd_mint,binanceusd_mint,husd_mint]
_all_burn = [usdc_burn,paxos_burn,dai_burn,tether_burn,
trueusd_burn,binanceusd_burn,husd_burn]
_all_non_transfer = [tether_mint,tether_burn,usdc_mint,usdc_burn,paxos_mint,
paxos_burn,dai_mint,dai_burn,trueusd_mint,trueusd_burn,
binanceusd_mint,binanceusd_burn,husd_mint,husd_burn]
_all_positive_cumulated_balances = [dai_positive_cumulated_balances, tether_positive_cumulated_balances,
usdc_positive_cumulated_balances, paxos_positive_cumulated_balances,
trueusd_positive_cumulated_balances,
binanceusd_positive_cumulated_balances, husd_positive_cumulated_balances]
_all_balances = [dai_balances, tether_balances, usdc_balances, paxos_balances,
trueusd_balances, binanceusd_balances, husd_balances]
# -
# <center></center>
# <center></center>
# <center></center>
# # Data
rows = 0
for i in _all:
rows += pd.read_csv(i).describe().loc['count','timestamp']
print('The dataset consisists of {:.0f} rows'.format(rows))
# Non Transfer Events
rows_non_transfer = 0
for i in _all_non_transfer:
rows_non_transfer += pd.read_csv(i).describe().loc['count','timestamp']
print('The dataset consisists of {:.0f} rows that are not Transfer Events'.format(rows_non_transfer))
print('In total, {:,.0f} Transfer Events'.format(48983417-352065))
print('In total, {:.2f}% of all Events are Transfer Events'.format((48983417-352065)/48983417*100))
print('{:,.0f} observations in total'.format(48631352*9+352065*8-74))
# <center></center>
# ## Unique Transactions (!Transfers)'
# +
rows_transactions = 0
for i in _all_tx:
rows_transactions += len(pd.read_csv(i)['txhash'].unique())
print('The Transfer Events were included in {:,.0f} transactions'.format(rows_transactions))
# -
# <center></center>
# <center></center>
# <center></center>
# # 4.1.1 Transfers
# ## Transfers per Transaction
print('Transfers/Transaction: {:,.2f}'.format(48631352/46204011))
# ## TetherTransfers/Total Transfers
print('Tether Transfers/Total Transfers: {:,.2f}'.format(39501960/48631352))
# ## Tether Transfer Rise 2020
print('Transfers from January 1st until July 1st increased by {:,.1f} %'.format(pd.read_csv(tether_tx_over_date, index_col=0).loc[945,]['txs']/pd.read_csv(tether_tx_over_date, index_col=0).loc[764,]['txs']*100))
# ## Others
idx = pd.date_range('01/01/2020', '30/06/2020')
_all = {}
for i in _all_transfer:
token = i.split('/')[1]
df = pd.read_csv(i, index_col='dates').drop('Unnamed: 0', axis=1)
df.index = pd.DatetimeIndex(df.index)
_all[token] = df.reindex(idx, fill_value=0).loc['01/01/2020':,:]
others = _all['usdc']+_all['paxos']+_all['dai']+_all['trueusd']+_all['binanceusd']+_all['husd']
print('Transfers from January 1st until July 1st increased by {:,.1f} % from {:,.0f} to {:,.0f}'.format(float(others.loc['2020-06-30',]/others.loc['2020-01-01',])*100, float(others.loc['2020-01-01',]), float(others.loc['2020-06-30',])))
# ## Paxos Scammer
fr = pd.read_csv(paxos_tx_count_from, index_col='Unnamed: 0').sort_values('txs', ascending = False)
to = pd.read_csv(paxos_tx_count_to, index_col='Unnamed: 0').sort_values('txs', ascending = False)
fr = pd.DataFrame(fr.loc[:fr.index[2],'txs'])
fr['tag'] = ['MMM BSC', 'Scammer', 'Kyber: Contract']
to = pd.DataFrame(to.loc[:to.index[2],'txs'])
to['tag'] = ['MMM BSC', '-', 'Scammer']
to.append(fr).groupby([to.append(fr).index, to.append(fr)['tag']]).sum().sort_values('txs')
len(pd.read_csv(paxos_transfer))
print('Share of MMMBSC Scammer: {:.2f} %'.format(650256/2580722*100))
print('Share of both Scammers: {:.2f} %'.format((650254+135484)/2580722*100))
# <center></center>
# <center></center>
# # DAI ATH Analysis
df = pd.read_csv(dai_transfer)
# ## Most active DAI addresses
pd.DataFrame(dict(Counter(df['txto'])), index=[0]).T[0].sort_values().iloc[-5:]
# ## Avg. Transfers from/to DSR
# +
ts = 1573603200
addr = '0x0000000000000000000000000000000000000000'
txs_from = []
txs_to = []
while ts < 1593561601:
df2 = df[(df['timestamp'] > ts) & (df['timestamp'] < ts+86400)]
txs_from.append(len(df2[df2['txfrom'] == addr].index))
txs_to.append(len(df2[df2['txto'] == addr].index))
ts += 86400
txs = (sum(txs_to)+sum(txs_from))/len(txs_to)
print('On average, {:,.1f} transfers to the address\nOn average {:,.1f} transfers from the address'.format(sum(txs_to)/len(txs_to),sum(txs_from)/len(txs_from)))
ts = 1583971200
_df = df[(df['timestamp'] >= ts+86400*0) & (df['timestamp'] < ts+86400*1)]
_to_12 = len(_df[_df['txto'] == addr].index)
_df = df[(df['timestamp'] >= ts+86400*1) & (df['timestamp'] < ts+86400*2)]
_to_13 = len(_df[_df['txto'] == addr].index)
_df = df[(df['timestamp'] >= ts+86400*0) & (df['timestamp'] < ts+86400*1)]
_fr_12 = len(_df[_df['txfrom'] == addr].index)
_df = df[(df['timestamp'] >= ts+86400*1) & (df['timestamp'] < ts+86400*2)]
_fr_13 = len(_df[_df['txfrom'] == addr].index)
df_12 = _to_12 + _fr_12
df_13 = _to_13 + _fr_13
print('\nOn 12 & 13 March:')
print('{:,.1f} and {:,.1f} transfers to and from the address\n'.format(df_12 ,df_13))
print('Increase by {:,.2f} % on 12 March'.format(df_12/txs*100))
print('Withdrawals increased by {:,.2f} % on 12 March'.format(_fr_12/(sum(txs_from)/len(txs_from))*100))
print('Deposits increased by {:,.2f} % on 12 March\n'.format(_to_12/(sum(txs_to)/len(txs_to))*100))
print('Increase by {:,.2f} % on 13 March'.format(df_13/txs*100))
print('Withdrawals increased by {:,.2f} % on 13 March'.format(_fr_13/(sum(txs_from)/len(txs_from))*100))
print('Deposits increased by {:,.2f} % on 13 March\n'.format(_to_13/(sum(txs_to)/len(txs_to))*100))
print('Increase by {:,.2f} % on both days'.format((df_12+df_13)/(txs*2)*100))
# -
# ## Avg. Transfers from/to OasisDEX
# +
ts = 1573603200
addr = '0x794e6e91555438afc3ccf1c5076a74f42133d08d'
txs_from = []
txs_to = []
while ts < 1593561601:
df2 = df[(df['timestamp'] > ts) & (df['timestamp'] < ts+86400)]
txs_from.append(len(df2[df2['txfrom'] == addr].index))
txs_to.append(len(df2[df2['txto'] == addr].index))
ts += 86400
txs = (sum(txs_to)+sum(txs_from))/len(txs_to)
print('On average, {:,.1f} transfers to the address\nOn average {:,.1f} transfers from the address'.format(sum(txs_to)/len(txs_to),sum(txs_from)/len(txs_from)))
ts = 1583971200
_df = df[(df['timestamp'] >= ts+86400*0) & (df['timestamp'] < ts+86400*1)]
_to_12 = len(_df[_df['txto'] == addr].index)
_df = df[(df['timestamp'] >= ts+86400*1) & (df['timestamp'] < ts+86400*2)]
_to_13 = len(_df[_df['txto'] == addr].index)
_df = df[(df['timestamp'] >= ts+86400*0) & (df['timestamp'] < ts+86400*1)]
_fr_12 = len(_df[_df['txfrom'] == addr].index)
_df = df[(df['timestamp'] >= ts+86400*1) & (df['timestamp'] < ts+86400*2)]
_fr_13 = len(_df[_df['txfrom'] == addr].index)
df_12 = _to_12 + _fr_12
df_13 = _to_13 + _fr_13
print('\nOn 12 & 13 March:')
print('{:,.1f} and {:,.1f} transfers to and from the address\n'.format(df_12 ,df_13))
print('Increase by {:,.2f} % on 12 March'.format(df_12/txs*100))
print('Increase by {:,.2f} % on 13 March'.format(df_13/txs*100))
print('Increase by {:,.2f} % on both days'.format((df_12+df_13)/(txs*2)*100))
# -
# ## Avg. Transfers from/to Uniswap: DAI
# +
ts = 1573603200
addr = '0x2a1530c4c41db0b0b2bb646cb5eb1a67b7158667'
txs_from = []
txs_to = []
while ts < 1593561601:
df2 = df[(df['timestamp'] > ts) & (df['timestamp'] < ts+86400)]
txs_from.append(len(df2[df2['txfrom'] == addr].index))
txs_to.append(len(df2[df2['txto'] == addr].index))
ts += 86400
txs = (sum(txs_to)+sum(txs_from))/len(txs_to)
print('On average, {:,.1f} transfers to the address\nOn average {:,.1f} transfers from the address'.format(sum(txs_to)/len(txs_to),sum(txs_from)/len(txs_from)))
ts = 1583971200
_df = df[(df['timestamp'] >= ts+86400*0) & (df['timestamp'] < ts+86400*1)]
_to_12 = len(_df[_df['txto'] == addr].index)
_df = df[(df['timestamp'] >= ts+86400*1) & (df['timestamp'] < ts+86400*2)]
_to_13 = len(_df[_df['txto'] == addr].index)
_df = df[(df['timestamp'] >= ts+86400*0) & (df['timestamp'] < ts+86400*1)]
_fr_12 = len(_df[_df['txfrom'] == addr].index)
_df = df[(df['timestamp'] >= ts+86400*1) & (df['timestamp'] < ts+86400*2)]
_fr_13 = len(_df[_df['txfrom'] == addr].index)
df_12 = _to_12 + _fr_12
df_13 = _to_13 + _fr_13
print('\nOn 12 & 13 March:')
print('{:,.1f} and {:,.1f} transfers to and from the address\n'.format(df_12 ,df_13))
print('Increase by {:,.2f} % on 12 March'.format(df_12/txs*100))
print('Increase by {:,.2f} % on 13 March'.format(df_13/txs*100))
print('Increase by {:,.2f} % on both days'.format((df_12+df_13)/(txs*2)*100))
# -
# ## Avg. Transfers from/to Kyber
# +
ts = 1573603200
addr = '0x65bf64ff5f51272f729bdcd7acfb00677ced86cd'
txs_from = []
txs_to = []
while ts < 1593561601:
df2 = df[(df['timestamp'] > ts) & (df['timestamp'] < ts+86400)]
txs_from.append(len(df2[df2['txfrom'] == addr].index))
txs_to.append(len(df2[df2['txto'] == addr].index))
ts += 86400
txs = (sum(txs_to)+sum(txs_from))/len(txs_to)
print('On average, {:,.1f} transfers to the address\nOn average {:,.1f} transfers from the address'.format(sum(txs_to)/len(txs_to),sum(txs_from)/len(txs_from)))
ts = 1583971200
_df = df[(df['timestamp'] >= ts+86400*0) & (df['timestamp'] < ts+86400*1)]
_to_12 = len(_df[_df['txto'] == addr].index)
_df = df[(df['timestamp'] >= ts+86400*1) & (df['timestamp'] < ts+86400*2)]
_to_13 = len(_df[_df['txto'] == addr].index)
_df = df[(df['timestamp'] >= ts+86400*0) & (df['timestamp'] < ts+86400*1)]
_fr_12 = len(_df[_df['txfrom'] == addr].index)
_df = df[(df['timestamp'] >= ts+86400*1) & (df['timestamp'] < ts+86400*2)]
_fr_13 = len(_df[_df['txfrom'] == addr].index)
df_12 = _to_12 + _fr_12
df_13 = _to_13 + _fr_13
print('\nOn 12 & 13 March:')
print('{:,.1f} and {:,.1f} transfers to and from the address\n'.format(df_12 ,df_13))
print('Increase by {:,.2f} % on 12 March'.format(df_12/txs*100))
print('Increase by {:,.2f} % on 13 March'.format(df_13/txs*100))
print('Increase by {:,.2f} % on both days'.format((df_12+df_13)/(txs*2)*100))
# -
# ## Avg. Transfers from/to Compound Dai
# +
ts = 1573603200
addr = '0x5d3a536e4d6dbd6114cc1ead35777bab948e3643'
txs_from = []
txs_to = []
while ts < 1593561601:
df2 = df[(df['timestamp'] > ts) & (df['timestamp'] < ts+86400)]
txs_from.append(len(df2[df2['txfrom'] == addr].index))
txs_to.append(len(df2[df2['txto'] == addr].index))
ts += 86400
txs = (sum(txs_to)+sum(txs_from))/len(txs_to)
print('On average, {:,.1f} transfers to the address\nOn average {:,.1f} transfers from the address'.format(sum(txs_to)/len(txs_to),sum(txs_from)/len(txs_from)))
ts = 1583971200
_df = df[(df['timestamp'] >= ts+86400*0) & (df['timestamp'] < ts+86400*1)]
_to_12 = len(_df[_df['txto'] == addr].index)
_df = df[(df['timestamp'] >= ts+86400*1) & (df['timestamp'] < ts+86400*2)]
_to_13 = len(_df[_df['txto'] == addr].index)
_df = df[(df['timestamp'] >= ts+86400*0) & (df['timestamp'] < ts+86400*1)]
_fr_12 = len(_df[_df['txfrom'] == addr].index)
_df = df[(df['timestamp'] >= ts+86400*1) & (df['timestamp'] < ts+86400*2)]
_fr_13 = len(_df[_df['txfrom'] == addr].index)
df_12 = _to_12 + _fr_12
df_13 = _to_13 + _fr_13
print('\nOn 12 & 13 March:')
print('{:,.1f} and {:,.1f} transfers to and from the address\n'.format(df_12 ,df_13))
print('Increase by {:,.2f} % on 12 March'.format(df_12/txs*100))
print('Increase by {:,.2f} % on 13 March'.format(df_13/txs*100))
# -
# <center></center>
# <center></center>
# # Gas Cost Comparison
# ## Average Gas Fees All Time
gas_teth = 27244892832.678*57439.360/10**18
gas_dai_ = 17711503439.366*472827.895/10**18
gas_usdc = 20520686635.884*385927.692/10**18
gas_paxo = 15768757613.356*152639.472/10**18
gas_tusd = 20112704549.078*90049.336/10**18
gas_bina = 33194878037.906*260947.284/10**18
gas_husd = 47252501697.708*44784.038/10**18
pd.DataFrame({'gascosts': [gas_teth,gas_dai_,gas_usdc,gas_paxo,gas_tusd,gas_bina,gas_husd]},
index=['gas_teth','gas_dai_','gas_usdc','gas_paxo','gas_tusd','gas_bina','gas_husd'])
# ## Average Gas Fees 2020
ga = {}
for i in _all_tx:
tk = i.split('/')[1]
_tf = pd.read_csv(i, usecols = ['timestamp','gas_price','gas_used'])
_tf = _tf[_tf['timestamp'] >= 1577836800] #01.01.2020
_tf['costs'] = _tf['gas_used']*_tf['gas_price']/10**18
ga[tk] = sum(_tf['costs'])/len(_tf['costs'])
ga
print('DAI/USDT: {}\nUSDC/USDT: {}\nBUSD/USDT: {}'.format(0.007909282581493824/0.0018932924837845818,
0.009161195861560514/0.0018932924837845818,
0.007751093391282079/0.0018932924837845818))
# # DeFi Impact on DAI's Gas Fee 2020
# +
dai_ = pd.read_csv(dai_transfer)
dai = dai_[dai_['timestamp'] >= 1577836800]
fr_da = dai[dai['txfrom'].isin(['0x0000000000000000000000000000000000000000', # DSR
'0x794e6e91555438afc3ccf1c5076a74f42133d08d', # OasisDEX
'0x2a1530c4c41db0b0b2bb646cb5eb1a67b7158667', # Uniswap: DAI
'0x65bf64ff5f51272f729bdcd7acfb00677ced86cd', # Kyber: Contract
'0x39755357759ce0d7f32dc8dc45414cca409ae24e', # Eth2Dai: Old Contract
'0x5d3a536e4d6dbd6114cc1ead35777bab948e3643', # Compound DAI
'0x11111254369792b2ca5d084ab5eea397ca8fa48b', # 1inch.exchange
'0xd152f549545093347a162dce210e7293f1452150', # Disperse.app
'0x31e085afd48a1d6e51cc193153d625e8f0514c7f'])] # Kyber: Reserve ...
to_da = dai[dai['txto'].isin(['0x0000000000000000000000000000000000000000', # DSR
'0x794e6e91555438afc3ccf1c5076a74f42133d08d', # OasisDEX
'0x2a1530c4c41db0b0b2bb646cb5eb1a67b7158667', # Uniswap: DAI
'0x65bf64ff5f51272f729bdcd7acfb00677ced86cd', # Kyber: Contract
'0x39755357759ce0d7f32dc8dc45414cca409ae24e', # Eth2Dai: Old Contract
'0x5d3a536e4d6dbd6114cc1ead35777bab948e3643', # Compound DAI
'0x11111254369792b2ca5d084ab5eea397ca8fa48b', # 1inch.exchange
'0xd152f549545093347a162dce210e7293f1452150', # Disperse.app
'0x31e085afd48a1d6e51cc193153d625e8f0514c7f'])] # Kyber: Reserve ...
perc_defi = (len(fr_da)+len(to_da))/len(dai)
df = fr_da.loc[:,['gas_price', 'gas_used']].append(to_da.loc[:,['gas_price', 'gas_used']])
df['costs'] = df['gas_price']*df['gas_used']/10**18
avg_defi_cost = df['costs'].mean()
print('DeFi/USDT: {}\nAvg. DeFi Transfer costs: {}\nDeFi share of total: {}'.format(avg_defi_cost/0.001893292,
avg_defi_cost,
perc_defi))
# -
# # DeFi Impact DAI's Gas Fee on since May '20
# +
dai_ = pd.read_csv(dai_transfer)
dai = dai_[dai_['timestamp'] >= 1588291200]
fr_da = dai[dai['txfrom'].isin(['0x0000000000000000000000000000000000000000', # DSR
'0x794e6e91555438afc3ccf1c5076a74f42133d08d', # OasisDEX
'0x2a1530c4c41db0b0b2bb646cb5eb1a67b7158667', # Uniswap: DAI
'0x65bf64ff5f51272f729bdcd7acfb00677ced86cd', # Kyber: Contract
'0x39755357759ce0d7f32dc8dc45414cca409ae24e', # Eth2Dai: Old Contract
'0x5d3a536e4d6dbd6114cc1ead35777bab948e3643', # Compound DAI
'0x11111254369792b2ca5d084ab5eea397ca8fa48b', # 1inch.exchange
'0xd152f549545093347a162dce210e7293f1452150', # Disperse.app
'0x31e085afd48a1d6e51cc193153d625e8f0514c7f'])] # Kyber: Reserve ...
to_da = dai[dai['txto'].isin(['0x0000000000000000000000000000000000000000', # DSR
'0x794e6e91555438afc3ccf1c5076a74f42133d08d', # OasisDEX
'0x2a1530c4c41db0b0b2bb646cb5eb1a67b7158667', # Uniswap: DAI
'0x65bf64ff5f51272f729bdcd7acfb00677ced86cd', # Kyber: Contract
'0x39755357759ce0d7f32dc8dc45414cca409ae24e', # Eth2Dai: Old Contract
'0x5d3a536e4d6dbd6114cc1ead35777bab948e3643', # Compound DAI
'0x11111254369792b2ca5d084ab5eea397ca8fa48b', # 1inch.exchange
'0xd152f549545093347a162dce210e7293f1452150', # Disperse.app
'0x31e085afd48a1d6e51cc193153d625e8f0514c7f'])] # Kyber: Reserve ...
perc_defi = (len(fr_da)+len(to_da))/len(dai)
df = fr_da.loc[:,['gas_price', 'gas_used']].append(to_da.loc[:,['gas_price', 'gas_used']])
df['costs'] = df['gas_price']*df['gas_used']/10**18
dai['costs'] = (dai['gas_price']*dai['gas_used']/10**18)
avg_defi_cost = df['costs'].mean()
print('DeFi/USDT: {}\nAvg. DeFi Transfer costs: {}\nDeFi share of total: {}'.format(avg_defi_cost/0.001893292,
avg_defi_cost,
perc_defi,
))
print('Avg. since May against tether: {}'.format((sum(dai['costs'])/len(dai)/0.001893292)))
# -
# ## DSR costs
dsr = dai_[(dai_['txfrom'] == '0x0000000000000000000000000000000000000000') | (dai_['txto'] == '0x0000000000000000000000000000000000000000') ]
dsr['costs'] = dsr['gas_price']*dsr['gas_used']/10**18
print('Avg. DSR Transfer costs: {}\nDSR/USDT: {}'.format(sum(dsr['costs'])/len(dsr['costs']),
sum(dsr['costs'])/len(dsr['costs'])/0.0018932924837845818))
# # DeFi Impact on USDC's Gas Fee 2020
# +
usdc = pd.read_csv(usdc_transfer)
usdc = usdc[usdc['timestamp'] >= 1577836800]
fr_da = usdc[usdc['txfrom'].isin(['0xd152f549545093347a162dce210e7293f1452150', # Disperse.app
'0x97dec872013f6b5fb443861090ad931542878126', # Uniswap: USDC
'0x65bf64ff5f51272f729bdcd7acfb00677ced86cd', # Kyber: Contract
'0x39aa39c021dfbae8fac545936693ac917d5e7563', # Compound USD Coin
'0x11111254369792b2ca5d084ab5eea397ca8fa48b', # 1inch.exchange
'0x9ae49c0d7f8f9ef4b864e004fe86ac8294e20950', # Kyber: Old Contract
'0x04046027549f739edfd5b2a78efdbaf0f0bf4514', # BlockFi
'0x65b0bf8ee4947edd2a500d74e50a3d757dc79de0', # Nexo: Wallet
'0x32a3256a4b15badd4a6e072a03d23404d925a5cf', # Celsius Network: Contract
'0xaf38668f4719ecf9452dc0300be3f6c83cbf3721'])] # Nuo Network: Kernel Escrow
to_da = usdc[usdc['txto'].isin(['0xd152f549545093347a162dce210e7293f1452150', # Disperse.app
'0x97dec872013f6b5fb443861090ad931542878126', # Uniswap: USDC
'0x65bf64ff5f51272f729bdcd7acfb00677ced86cd', # Kyber: Contract
'0x39aa39c021dfbae8fac545936693ac917d5e7563', # Compound USD Coin
'0x11111254369792b2ca5d084ab5eea397ca8fa48b', # 1inch.exchange
'0x9ae49c0d7f8f9ef4b864e004fe86ac8294e20950', # Kyber: Old Contract
'0x04046027549f739edfd5b2a78efdbaf0f0bf4514', # BlockFi
'0x65b0bf8ee4947edd2a500d74e50a3d757dc79de0', # Nexo: Wallet
'0x32a3256a4b15badd4a6e072a03d23404d925a5cf', # Celsius Network: Contract
'0xaf38668f4719ecf9452dc0300be3f6c83cbf3721'])] # Nuo Network: Kernel Escrow
perc_defi = (len(fr_da)+len(to_da))/len(usdc)
df = fr_da.loc[:,['gas_price', 'gas_used']].append(to_da.loc[:,['gas_price', 'gas_used']])
df['costs'] = df['gas_price']*df['gas_used']/10**18
avg_defi_cost = df['costs'].mean()
print('DeFi/USDT Costs: {}\nAvg. DeFi Transfer costs: {}\nDeFi share of total: {}'.format(avg_defi_cost/0.001893292,
avg_defi_cost,
perc_defi))
# -
# # DeFi Impact on USDC's Gas Fee since May '20
# +
usdc = pd.read_csv(usdc_transfer)
usdc = usdc[usdc['timestamp'] >= 1588291200]
fr_da = usdc[usdc['txfrom'].isin(['0xd152f549545093347a162dce210e7293f1452150', # Disperse.app
'0x97dec872013f6b5fb443861090ad931542878126', # Uniswap: USDC
'0x65bf64ff5f51272f729bdcd7acfb00677ced86cd', # Kyber: Contract
'0x39aa39c021dfbae8fac545936693ac917d5e7563', # Compound USD Coin
'0x11111254369792b2ca5d084ab5eea397ca8fa48b', # 1inch.exchange
'0x9ae49c0d7f8f9ef4b864e004fe86ac8294e20950', # Kyber: Old Contract
'0x04046027549f739edfd5b2a78efdbaf0f0bf4514', # BlockFi
'0x65b0bf8ee4947edd2a500d74e50a3d757dc79de0', # Nexo: Wallet
'0x32a3256a4b15badd4a6e072a03d23404d925a5cf', # Celsius Network: Contract
'0xaf38668f4719ecf9452dc0300be3f6c83cbf3721'])] # Nuo Network: Kernel Escrow
to_da = usdc[usdc['txto'].isin(['0xd152f549545093347a162dce210e7293f1452150', # Disperse.app
'0x97dec872013f6b5fb443861090ad931542878126', # Uniswap: USDC
'0x65bf64ff5f51272f729bdcd7acfb00677ced86cd', # Kyber: Contract
'0x39aa39c021dfbae8fac545936693ac917d5e7563', # Compound USD Coin
'0x11111254369792b2ca5d084ab5eea397ca8fa48b', # 1inch.exchange
'0x9ae49c0d7f8f9ef4b864e004fe86ac8294e20950', # Kyber: Old Contract
'0x04046027549f739edfd5b2a78efdbaf0f0bf4514', # BlockFi
'0x65b0bf8ee4947edd2a500d74e50a3d757dc79de0', # Nexo: Wallet
'0x32a3256a4b15badd4a6e072a03d23404d925a5cf', # Celsius Network: Contract
'0xaf38668f4719ecf9452dc0300be3f6c83cbf3721'])] # Nuo Network: Kernel Escrow
perc_defi = (len(fr_da)+len(to_da))/len(usdc)
df = fr_da.loc[:,['gas_price', 'gas_used']].append(to_da.loc[:,['gas_price', 'gas_used']])
df['costs'] = df['gas_price']*df['gas_used']/10**18
usdc['costs'] = (usdc['gas_price']*usdc['gas_used']/10**18)
avg_defi_cost = df['costs'].mean()
print('DeFi/USDT: {}\nAvg. DeFi Transfer costs: {}\nDeFi share of total: {}'.format(avg_defi_cost/0.001893292,
avg_defi_cost,
perc_defi))
print('Avg. since May against tether: {}'.format((sum(usdc['costs'])/len(usdc)/0.001893292)))
# -
# <center></center>
# <center></center>
# ## Most expensive transfer
aa = pd.read_csv(tether_chunk5, usecols=['txhash','gas_price', 'gas_used'])
aa["costs"] = (aa["gas_price"]/10**18).multiply(aa["gas_used"])
aa = aa[aa['costs'] == max(aa['costs'])]
print(aa['txhash'].tolist())
aa
# <center></center>
# <center></center>
# # Circulating supply Analytics
su = 0
for i, j in zip(_all_mint,_all_burn):
tk = i.split('/')[1]
tk2 = j.split('/')[1]
assert(tk==tk2)
if tk in ['dai', 'trueusd', 'paxos', 'binanceusd']:
dec=18
elif tk in ['husd']:
dec=8
else:
dec=6
print(tk)
df = pd.read_csv(i)
df2 = pd.read_csv(j)
su += sum(df['txvalue'].astype(float)/10**dec) - sum(df2['txvalue'].astype(float)/10**dec)
print('In total, {:,.0f} Stablecoins'.format(su))
# ## USDT supply
sum(pd.read_csv(tether_mint)['txvalue']/10**6)-sum(pd.read_csv(tether_burn)['txvalue']/10**6)
# ## USDC supply
sum(pd.read_csv(usdc_mint)['txvalue']/10**6)-sum(pd.read_csv(usdc_burn)['txvalue']/10**6)
# ## Others supply
7800726550-6037847550.677356
# ## USDT supply/Others
6037847550.677356/1762878999.3226442
# ## USDT supply/total Stablecoin supply
6037847550.677356/7800726550
# ## Mint Events
# +
tks = {}
avg = {}
rows = 0
for i in _all_mint:
tk = i.split('/')[1]
if tk in ['dai', 'trueusd', 'paxos', 'binanceusd']:
dec=18
elif tk in ['husd']:
dec=8
else:
dec=6
df = pd.read_csv(i)
rws = len(df)
me = round((df['txvalue'].astype(float)/10**dec).mean())
rows += rws
tks[tk] = rws
avg[tk] = me
print('In total, {:,.0f} Mint Events'.format(rows))
tks, avg
# -
# ## HUSD mint + burn events / total events
'{:.2f} %'.format((3598+6992)/(3598+6992 + 36821)*100)
# ## DAI mint + burn events / total events
'{:.2f} %'.format((150131+167956)/(150131+167956 + 2647517)*100)
# ## Burn Events
# +
tks = {}
avg = {}
rows = 0
for i in _all_burn:
tk = i.split('/')[1]
if tk in ['dai', 'trueusd', 'paxos', 'binanceusd']:
dec=18
elif tk in ['husd']:
dec=8
else:
dec=6
df = pd.read_csv(i)
rws = len(df)
me = round((df['txvalue'].astype(float)/10**dec).mean())
rows += rws
tks[tk] = rws
avg[tk] = me
print('In total, {:,.0f} Burn Events'.format(rows))
tks, avg
# -
# ## Number of events influencing the total supply
170386+181679
# ## DAI/Total
(150131+167956)/(170386+181679)
# ## Dai Max Mint
# +
df = pd.read_csv(dai_mint, index_col=0)
mean_all = []
ts = 1573603200
dailymax = 0
while ts < 1593561600:
dailysum = sum(df[(df['timestamp'] >= ts) & (df['timestamp'] < ts+86400) ]['txvalue'].astype(float)/10**18)
if dailysum > dailymax:
dailymax = dailysum
day = str(datetime.utcfromtimestamp(ts))[0:10]
tos = len(df[(df['timestamp'] >= ts) & (df['timestamp'] < ts+86400) ]['txto'].unique())
lar = max(df[(df['timestamp'] >= ts) & (df['timestamp'] < ts+86400) ]['txvalue'])
mean_all.append(dailysum)
ts += 86400
dailymax, day, tos, lar, dailymax/(sum(mean_all)/len(mean_all))
# -
# ## Dai Max Burn
# +
df = pd.read_csv(dai_burn, index_col=0)
mean_all = []
ts = 1573603200
dailymax = 0
while ts < 1593561600:
dailysum = sum(df[(df['timestamp'] >= ts) & (df['timestamp'] < ts+86400) ]['txvalue'].astype(float)/10**18)
if dailysum > dailymax:
dailymax = dailysum
day = str(datetime.utcfromtimestamp(ts))[0:10]
tos = len(df[(df['timestamp'] >= ts) & (df['timestamp'] < ts+86400) ]['txto'].unique())
lar = max(df[(df['timestamp'] >= ts) & (df['timestamp'] < ts+86400) ]['txvalue'])
mean_all.append(dailysum)
ts += 86400
dailymax, day, tos, lar, dailymax/(sum(mean_all)/len(mean_all))
# -
df = pd.read_csv(usdc_burn)
df
# ## Describe Mint and burns
mb = pd.DataFrame()
for i, j in zip(_all_mint,_all_burn):
tk = i.split('/')[1]
tk2 = j.split('/')[1]
assert(tk==tk2)
if tk in ['dai', 'trueusd', 'paxos', 'binanceusd']:
dec=18
elif tk in ['husd']:
dec=8
else:
dec=6
df = pd.read_csv(i)
df2 = pd.read_csv(j)
mn = pd.DataFrame({tk+'_mint':df['txvalue'].astype(float)/10**dec})
mn2 = pd.DataFrame({tk+'_burn':df2['txvalue'].astype(float)/10**dec})
mb = pd.concat([mb, mn], axis=1)
mb = pd.concat([mb, mn2], axis=1)
pd.set_option('display.float_format', lambda x: format(x, ',.0f'))
mb.describe()
# ## Tether Mints in 2020
df = pd.read_csv(tether_mint)
to_mi = sum(df['txvalue'])
ts = 1577836800 # 01.01.2020
sum(df[df['timestamp'] >= ts]['txvalue'])/10**6,sum(df[df['timestamp'] >= ts]['txvalue'])/to_mi
# ## DAI mints to unique addresses
len(pd.read_csv(dai_mint, index_col=0)['txto'].unique())
# ## DAI burns from unique addresses
len(pd.read_csv(dai_burn, index_col=0)['address'].unique())
# ## Tether burns from unique addresses
len(pd.read_csv(tether_burn, index_col=0)['address'].unique())
# ## USDC mints to unique addresses
df = pd.read_csv(usdc_mint, index_col=0)
len(df['address'].unique()),df['address'].unique()
df = pd.read_csv(usdc_mint, index_col=0)
sum(df[df['address'] == '0x55fe002aeff02f77364de339a1292923a15844b8']['txvalue']/10**6)
# ## USDC burns from unique addresses
df = pd.read_csv(usdc_burn, index_col=0)
len(df['address'].unique()),df['address'].unique()
df = pd.read_csv(usdc_burn, index_col=0)
sum(df[df['address'] == '0xd4c1315948125cd20c11c5e9565a3632c1710055']['txvalue']/10**6)
# ## TrueUSD mints to unique addresses
len(pd.read_csv(trueusd_mint, index_col=0)['address'].unique())
# ## TrueUSD burns from unique addresses
len(pd.read_csv(trueusd_burn, index_col=0)['address'].unique())
# ## Paxos, BinanceUSD and HUSD mints to unique addresses
len(pd.read_csv(paxos_mint, index_col=0)['address'].unique()),len(pd.read_csv(binanceusd_mint, index_col=0)['address'].unique()),len(pd.read_csv(husd_mint, index_col=0)['address'].unique())
# ## Paxos, BinanceUSD and HUSD burns from unique addresses
len(pd.read_csv(paxos_burn, index_col=0)['address'].unique()),len(pd.read_csv(binanceusd_burn, index_col=0)['address'].unique()),len(pd.read_csv(husd_burn, index_col=0)['address'].unique())
# ## DAI total Mint
sum(pd.read_csv(dai_mint, index_col=0)['txvalue'].astype(float)/10**18)
# ## DAI total Burn
sum(pd.read_csv(dai_burn, index_col=0)['txvalue'].astype(float)/10**18)
# ## Tether total Burn
sum(pd.read_csv(tether_burn, index_col=0)['txvalue'].astype(float)/10**6)
# ## USDC total Mint
sum(pd.read_csv(usdc_mint, index_col=0)['txvalue'].astype(float)/10**6)
# ## USDC total Burn
sum(pd.read_csv(usdc_burn, index_col=0)['txvalue'].astype(float)/10**6)
# ## TrueUSD total Mint
sum(pd.read_csv(trueusd_mint, index_col=0)['txvalue'].astype(float)/10**18)
# ## TrueUSD total Burn
sum(pd.read_csv(trueusd_burn, index_col=0)['txvalue'].astype(float)/10**18)
# ## PAX, BUSD and HUSD total Mint
sum(pd.read_csv(paxos_mint, index_col=0)['txvalue'].astype(float)/10**18),sum(pd.read_csv(binanceusd_mint, index_col=0)['txvalue'].astype(float)/10**18),sum(pd.read_csv(husd_mint, index_col=0)['txvalue'].astype(float)/10**8)
# ## PAX, BUSD and HUSD total Burn
sum(pd.read_csv(paxos_burn, index_col=0)['txvalue'].astype(float)/10**18),sum(pd.read_csv(binanceusd_burn, index_col=0)['txvalue'].astype(float)/10**18),sum(pd.read_csv(husd_burn, index_col=0)['txvalue'].astype(float)/10**8)
df = pd.read_csv(paxos_burn, index_col=0)#
df[df['address'] != '0x5195427ca88df768c298721da791b93ad11eca65']
# <center></center>
# <center></center>
# <center></center>
# <center></center>
# <center></center>
# <center></center>
# # Address Analytics
# ## Total Addresses
# +
addr = []
toke = {}
for i in _all_balances:
tk = i.split('/')[1]
df = pd.read_csv(i)
df = df['address'].tolist()
addr = addr + df
toke[tk] = '{:,.0f}'.format(len(df))
print('Total unique addresses: {:,.0f}'.format(len(set(addr))))
toke
# -
# ## Tethers share of addresses
6301972/8372918
# ## Total positive Addresses
fr = pd.read_csv('plots/summary/from.csv')
to = pd.read_csv('plots/summary/to.csv')
df = fr.append(to).groupby('Unnamed: 0').sum()
tether_bal = pd.read_csv(tether_positive_cumulated_balances, index_col='Unnamed: 0').drop('cum', axis=1).set_index('address')
usdc_bal = pd.read_csv(usdc_positive_cumulated_balances, index_col='Unnamed: 0').drop('cum', axis=1).set_index('address')
dai_bal = pd.read_csv(dai_positive_cumulated_balances, index_col='Unnamed: 0').drop('cum', axis=1).set_index('address')
paxos_bal = pd.read_csv(paxos_positive_cumulated_balances, index_col='Unnamed: 0').drop('cum', axis=1).set_index('address')
binanceusd_bal = pd.read_csv(binanceusd_positive_cumulated_balances, index_col='Unnamed: 0').drop('cum', axis=1).set_index('address')
trueusd_bal = pd.read_csv(trueusd_positive_cumulated_balances, index_col='Unnamed: 0').drop('cum', axis=1).set_index('address')
husd_bal = pd.read_csv(husd_positive_cumulated_balances, index_col='Unnamed: 0').drop('cum', axis=1).set_index('address')
df['bal_tether'] = tether_bal
df['bal_usdc'] = usdc_bal
df['bal_dai'] = dai_bal
df['bal_paxos'] = paxos_bal
df['bal_binanceusd'] = binanceusd_bal
df['bal_trueusd'] = trueusd_bal
df['bal_husd'] = husd_bal
df = df.fillna(0)
df['sum'] = df['bal_tether']+df['bal_usdc']+df['bal_dai']+df['bal_paxos']+df['bal_binanceusd']+df['bal_trueusd'] +df['bal_husd']
df['normsum'] = df['sum']/sum(df['sum'])
df = df.sort_values('sum')
# ## Describe Dataset
df.describe()
# ## Addresses with a positive balance
print('{} addresses with {} USD'.format(len(df[df['sum'] > 0]), sum(df[df['sum'] > 0 ]['sum'])))
# ## Addresses with less than 1 dollar in Stablecoins
print('{} addresses with {} USD'.format(len(df[(df['sum'] <= 1) & (df['sum'] > 0)]), sum(df[(df['sum'] <= 1) & (df['sum'] > 0)]['sum']) ))
# ## Addresses with less than 0.01 dollar in Stablecoins
print('{} addresses with {} USD'.format(len(df[(df['sum'] < 0.01) & (df['sum'] > 0)]), sum(df[(df['sum'] < 0.01) & (df['sum'] > 0)]['sum']) ))
# ## Addresses withmore than 1 mio dollar in Stablecoins
print('{} addresses with {} USD'.format(len(df[df['sum'] > 1e6]), sum(df[df['sum'] > 1e6]['sum'])))
# +
df_ = df[df['sum']>0]
dfsu = sum(df_['sum'])
a = df_.iloc[-round(len(df_)/10):,:] # 10 %
b = df_.iloc[-round(len(df_)/20):,:] # 5 %
x = df_.iloc[-round(len(df_)/100):,:] # 1 %
y = df_.iloc[-round(len(df_)/1000):,:] # 0.1 %
z = df_.iloc[-round(len(df_)/10000):,:] # 0.01 %
al = len(a)
bl = len(b)
xl = len(x)
yl = len(y)
zl = len(z)
asu = sum(a['sum'])
bsu = sum(b['sum'])
xsu = sum(x['sum'])
ysu = sum(y['sum'])
zsu = sum(z['sum'])
print('{:,.2f} addresses ({:,.2f}%) own {:,.2f}; {:,.2f}% of total'.format(al, 10, asu, asu/dfsu*100))
print('{:,.2f} addresses ({:,.2f}%) own {:,.2f}; {:,.2f}% of total'.format(bl, 5, bsu, bsu/dfsu*100))
print('{:,.2f} addresses ({:,.2f}%) own {:,.2f}; {:,.2f}% of total'.format(xl, 1, xsu, xsu/dfsu*100))
print('{:,.2f} addresses ({:,.2f}%) own {:,.2f}; {:,.2f}% of total'.format(yl, 0.1, ysu, ysu/dfsu*100))
print('{:,.2f} addresses ({:,.2f}%) own {:,.2f}; {:,.2f}% of total'.format(zl, 0.1, zsu, zsu/dfsu*100))
# -
# ## Desribe positive addresses
ds = df[(df.loc[:,'bal_tether':'bal_husd']>0).any(axis=1)].replace(0, np.NaN).describe().style.format("{:,.2f}")
ds
# ## Addresses with a balance larger than 10 USD
print('Tether Addresses with a balance larger than 10 USD: {:,.0f}'.format(len(df[(df['bal_tether']>10)])))
print('USDC Addresses with a balance larger than 10 USD: {:,.0f}'.format(len(df[(df['bal_usdc']>10)])))
print('Paxos Addresses with a balance larger than 10 USD: {:,.0f}'.format(len(df[(df['bal_paxos']>10)])))
print('TrueUSD Addresses with a balance larger than 10 USD: {:,.0f}'.format(len(df[(df['bal_trueusd']>10)])))
print('BinanceUSD Addresses with a balance larger than 10 USD: {:,.0f}'.format(len(df[(df['bal_binanceusd']>10)])))
print('HUSD Addresses with a balance larger than 10 USD: {:,.0f}'.format(len(df[(df['bal_husd']>10)])))
print('DAI Addresses with a balance larger than 10 USD: {:,.0f}'.format(len(df[(df['bal_dai']>10)])))
# ## Positive Addresses with Tether/Positive Addresses
'Positive Addresses with Tether/Positive Addresses: {:.2f}%'.format(
len(df[(df['bal_tether']>0)]) # positive tether addresses
/
len(df[(df.loc[:,'bal_tether':'bal_husd']>0).any(axis=1)])
*100
)
# ## Summed balance of the 20 richest addresses for each token
te,uc,da,pa,bu,tu,hu = [0]*7
for i,e in df.tail(20).iloc[::-1].iterrows():
a,b,c,d,e,f,g = e.tolist()[1:-2]
te += a
uc += b
da += c
pa += d
bu += e
tu += f
hu += g
te,uc,da,pa,bu,tu,hu
df.tail(20)
# # Rich List (Latex)
te,uc,da,pa,bu,tu,hu= [0]*7
for i,e in df.tail(20).iloc[::-1].iterrows():
su = sum(e.tolist()[1:-2])
_a,_b,_c,_d,_e,_f,_g = e.tolist()[1:-2]
te += _a/1e6
uc += _b/1e6
da += _c/1e6
pa += _d/1e6
bu += _e/1e6
tu += _f/1e6
hu += _g/1e6
ba = [j.replace('0.00', '-') if float(j) == 0 else j for j in ['{:,.2f}'.format(i/1e6) for i in e.tolist()[1:-2]]]
print(i[0:5] +' & & ' + ' & '.join(ba) + ' & {:,.2f} \\\[-0.6ex] \n\hline'.format(su/1e6))
to=te+uc+da+pa+bu+tu+hu
print('$\sum$&&{:,.2f}&{:,.2f}&{:,.2f}&{:,.2f}&{:,.2f}&{:,.2f}&{:,.2f}&{:,.0f}\\\[-0.6ex]'.format(te,uc,da,pa,bu,tu,hu,to))
# ## Balance of richest address
df.reset_index().loc[df.loc[:,'bal_tether'].argmax(),:]['bal_tether']
# ## Balance of 2nd richest address
sum(df.loc['0xbe0eb53f46cd790cd13851d5eff43d12404d33e8','bal_tether':'bal_husd'])
# ## ... and its tokens
df.loc['<KEY>','bal_tether':'bal_husd'].apply(lambda x: format(x, '6f'))
# ## Balance of 3rt richest address
sum(df.loc['0x39aa39c021dfbae8fac545936693ac917d5e7563','bal_tether':'bal_husd'])
# ## Balance of richest HUSD address
sum(df.loc['0x1062a747393198f70f71ec65a582423dba7e5ab3','bal_tether':'bal_husd'])
# ## Balance of richest DAI address
sum(df.loc['0x1e0447b19bb6ecfdae1e4ae1694b0c3659614e4e','bal_tether':'bal_husd'])
df.loc['0x1e0447b19bb6ecfdae1e4ae1694b0c3659614e4e','bal_tether':'bal_husd']
df.tail(83)
# ## Are there addresses that have more than one Stablecoin?
df_dai = set(pd.read_csv(dai_positive_cumulated_balances)['address'])
df_tet = set(pd.read_csv(tether_positive_cumulated_balances)['address'])
df_usd = set(pd.read_csv(usdc_positive_cumulated_balances)['address'])
df_pax = set(pd.read_csv(paxos_positive_cumulated_balances)['address'])
df_hus = set(pd.read_csv(husd_positive_cumulated_balances)['address'])
df_bin = set(pd.read_csv(binanceusd_positive_cumulated_balances)['address'])
df_tus = set(pd.read_csv(trueusd_positive_cumulated_balances)['address'])
# +
#Tether and DAI Stablecoin
def get_similar_addr_1(df, df2, df3=None, df4=None, df5=None, df6=None, df7=None):
if df3:
df2 = set(list(df2)+list(df3))
if df4:
df2 = set(list(df2)+list(df4))
if df5:
df2 = set(list(df2)+list(df5))
if df6:
df2 = set(list(df2)+list(df6))
if df7:
df2 = set(list(df2)+list(df7))
print()
return [i for i in df if i in df2]
def get_similar_addr_2(df, df2, df3=None, df4=None, df5=None, df6=None, df7=None):
if df7:
return [i for i in df if i in df2 and i in df3 and i in df4 and i in df5 and i in df6 and i in df7]
print()
return [i for i in df if i in df2]
# -
# ## Addresses that have USDT and at least one other Stablecoin
addrs = get_similar_addr_1(list(df_tet),df_dai,df_usd,df_pax,df_hus,df_bin,df_tus)
to_ba = len(df_tet)
'Addresses with USDT and at least one other Stablecoin: {:,.0f}({:,.2f}%)'.format(len(addrs),len(addrs)/to_ba*100)
# ## Addresses that have DAI and at least one other Stablecoin
addrs = get_similar_addr_1(list(df_dai),df_tet,df_usd,df_pax,df_hus,df_bin,df_tus)
to_ba = len(df_dai)
'Addresses with DAI and at least one other Stablecoin: {:,.0f}({:,.2f}%)'.format(len(addrs),len(addrs)/to_ba*100)
# ## Addresses that have USDC and at least one other Stablecoin
addrs = get_similar_addr_1(list(df_usd),df_tet,df_dai,df_pax,df_hus,df_bin,df_tus)
to_ba = len(df_usd)
'Addresses with USDC and at least one other Stablecoin: {:,.0f}({:,.2f}%)'.format(len(addrs),len(addrs)/to_ba*100)
# ## Addresses that have PAX and at least one other Stablecoin
addrs = get_similar_addr_1(list(df_pax),df_tet,df_usd,df_dai,df_hus,df_bin,df_tus)
to_ba = len(df_pax)
'Addresses with PAX and at least one other Stablecoin: {:,.0f}({:,.2f}%)'.format(len(addrs),len(addrs)/to_ba*100)
# ## Addresses that have TUSD and at least one other Stablecoin
addrs = get_similar_addr_1(list(df_tus),df_tet,df_usd,df_pax,df_hus,df_bin,df_dai)
to_ba = len(df_tus)
'Addresses with TUSD and at least one other Stablecoin: {:,.0f}({:,.2f}%)'.format(len(addrs),len(addrs)/to_ba*100)
# ## Addresses that have BUSD and at least one other Stablecoin
addrs = get_similar_addr_1(list(df_bin),df_tet,df_usd,df_pax,df_hus,df_dai,df_tus)
to_ba = len(df_bin)
'Addresses with BUSD and at least one other Stablecoin: {:,.0f}({:,.2f}%)'.format(len(addrs),len(addrs)/to_ba*100)
# ## Addresses that have HUSD and at least one other Stablecoin
addrs = get_similar_addr_1(list(df_hus),df_tet,df_usd,df_pax,df_dai,df_bin,df_tus)
to_ba = len(df_hus)
'Addresses with HUSD and at least one other Stablecoin: {:,.0f}({:,.2f}%)'.format(len(addrs),len(addrs)/to_ba*100)
# ## Addresses owing all Stablecoins analyzed (latex formated)
# +
_all = get_similar_addr_2(df_tet, df_dai,df_pax, df_usd, df_tus, df_hus, df_bin)
for i in _all:
print(i, '& \\\ [0ex]')
# -
# # Billionaires
# ## Huobi
ex = pd.read_csv(exchanges, header=None)
df = ex.loc[0:73,:]
bal = 0
for i in _all_positive_cumulated_balances:
balance = pd.read_csv(i)
for i in df[0]:
val = balance['balance'][balance['address'] == i]
if not val.empty:
bal += balance['balance'][balance['address'] == i].values[0]
else:
pass
print('Huobi Total Balance: {:,.0f} on Ethereum\nAdditional 124,680,800* and 64,389,947** on Bitcoin'.format(bal))
print('and 627,942,845*** on Tron')
print('------------------------\nIn Total: {:,.0f} USD without Tron\n------------------------\n'.format(bal+124680800+64389947))
print('*35hK24tcLEWcgNA4JxpvbkNkoAcDGqQPsP, **1HckjUpRGcrrRAtFaaCAUaGjsPx9oYmLaZ', '***TNaRAoLUyYEV2uF7GUrzSjRQTU8v5ZJ5VR')
# ## Huobi share of total Stablecoin supply
897573359/7800726550
# ## Binance
# +
ex = pd.read_csv(exchanges, header=None)
df = ex.loc[74:88,:]
bal = 0
for i in _all_positive_cumulated_balances:
balance = pd.read_csv(i)
for i in df[0]:
val = balance['balance'][balance['address'] == i]
if not val.empty:
bal += balance['balance'][balance['address'] == i].values[0]
else:
pass
print('Binance Total Balance: {:,.0f} on Ethereum\nAdditional 112,413,357* on Bitcoin'.format(bal))
print('and 257,387,652** on Tron')
print('------------------------\nIn Total: {:,.0f} USD\n------------------------\n'.format(bal+112413357+257387652))
print('*1FoWyxwPXuj4C6abqwhjDWdz6D4PZgYRjA, **TWd4WrZ9wn84f5x1hZhL4DHvk738ns5jwb')
# -
# ## Binance share of total Stablecoin supply
642624504/7800726550
# ## Latex code for Appendix with all identified addresses
ex = pd.read_csv(exchanges).iloc[:,:]
for i, e in ex.iterrows():
print('\\footnotesize{',e[0],'}', ' & ', '\\footnotesize{', e[1],'}', ' \\\ [-2.3ex]')
|
data_analytics.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Module 3 Discussion Grading - Peer Evaluation Grade Calculation
# <NAME>
#
# 2021-10-31
# ## Objective
#
# The objective of this file is to append all submitted peer-evaluation forms.
# ## Data Description
#
# Excel files downloaded from Canvas were used. Some students submtited Excel files, but changed the format of the table. It's adjusted mannually.
#
# Submissions with wrong group numbers (such as Group number = 10) are excluded.
#
# Submissions with wrong names where the errors cannot be parsed by me were excluded.
#
# cd C:\Users\jillb\OneDrive - UBC\CONS 302 TA 2021Fall\grading\module 3\new
# +
import os
import glob
os.chdir(r'C:\Users\jillb\OneDrive - UBC\CONS 302 TA 2021Fall\grading\module 3\new')
FileList = glob.glob('*.xlsx')
# -
import pandas as pd
# Loop through all Excel files, append all rows together (Columns should be the same)
#
# Issue: "File is not a recognized excel file".
#
# Debug process: Checked Pandas documentation; Tried different attributes (concat, append, etc); Deleted "," and "+" from Excel names; Renamed all Excel files as numbers. Did not work. Then suspected some files were corrupted. Re-downloaded all submissions from Canvas. Did not work. Moved all Excel files to a new directory, renamed the directory so that it does not have the same name with the deleted compressed folder. Did not work. Then ordered all students' Excel files based on the file size (learned this method from Jason Sutherland). One student's file is extradionarily smaller than everyone else's. Tried to open it mannually, but could not open because "the file extension or file format is not valid". Deleted that file.
#
# Then it worked.
#
#
# +
finalexcelsheet = pd.DataFrame()
for File in FileList:
# combining multiple excel worksheets
# into single data frames
df = pd.concat(pd.read_excel(File, sheet_name=None),
ignore_index=True, sort=False)
# Appending excel files one by one
finalexcelsheet = finalexcelsheet.append(
df, ignore_index=True)
# -
# cd C:\Users\jillb\OneDrive - UBC\CONS 302 TA 2021Fall\grading\module 3
finalexcelsheet.to_excel(r'Final.xlsx', index=False)
# Note:
#
# Many repetitive columns occurred because some students changed the column names. Told them not to but still did. Maybe some students' Excel automatically change that? I rememeber I had similar issues with Excel automatically change the date format, resulting in errors in REDCap data updating. Probably I should just make Column names' first letter upper case for everyone.
#
# Some new column occurred because students added notes to empty columns. Considering students would like to give reasons to why their teammates deserve the score, I should probably make next time's sample Excel file consist of a column named "notes".
#
#
#
#
|
CONS 302 101 2021W1 Module 3 Discussion Grading - Peer Evaluation Grade Calculation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6 (Develer Science)
# language: python
# name: develer-science
# ---
# ##### (exceprt from Python Machine Learning Essentials, Supplementary Materials)
# ### Sections
#
# - [Implementing a perceptron learning algorithm in Python](#Implementing-a-perceptron-learning-algorithm-in-Python)
# - [Training a perceptron model on the Iris dataset](#Training-a-perceptron-model-on-the-Iris-dataset)
# - [Adaptive linear neurons and the convergence of learning](#Adaptive-linear-neurons-and-the-convergence-of-learning)
# - [Implementing an adaptive linear neuron in Python](#Implementing-an-adaptive-linear-neuron-in-Python)
# +
# Display plots in notebook
# %matplotlib inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# -
# ## Implementing a perceptron learning algorithm in Python
# [[back to top](#Sections)]
from ann import Perceptron
# +
# Perceptron?
# -
# <br>
# <br>
# ### Training a perceptron model on the Iris dataset
# [[back to top](#Sections)]
# #### Reading-in the Iris data
# +
from sklearn.datasets import load_iris
iris = load_iris()
X = iris.data
y = iris.target
data = np.hstack((X, y[:, np.newaxis]))
labels = iris.target_names
features = iris.feature_names
df = pd.DataFrame(data, columns=iris.feature_names+['label'])
df.label = df.label.map({k:v for k,v in enumerate(labels)})
df.tail()
# -
# <br>
# <br>
# #### Plotting the Iris data
# +
# select setosa and versicolor
y = df.iloc[0:100, 4].values
y = np.where(y == 'setosa', -1, 1)
# extract sepal length and petal length
X = df.iloc[0:100, [0, 2]].values
# plot data
plt.scatter(X[:50, 0], X[:50, 1],
color='red', marker='o', label='setosa')
plt.scatter(X[50:100, 0], X[50:100, 1],
color='blue', marker='x', label='versicolor')
plt.xlabel('petal length [cm]')
plt.ylabel('sepal length [cm]')
plt.legend(loc='upper left')
plt.show()
# -
# <br>
# <br>
# #### Training the perceptron model
# +
ppn = Perceptron(eta=0.1, n_iter=10)
ppn.fit(X, y)
plt.plot(range(1, len(ppn.errors_) + 1), ppn.errors_, marker='o')
plt.xlabel('Epochs')
plt.ylabel('Number of misclassifications')
plt.tight_layout()
plt.show()
# -
# <br>
# <br>
# #### A function for plotting decision regions
# +
from matplotlib.colors import ListedColormap
def plot_decision_regions(X, y, classifier, resolution=0.02):
# setup marker generator and color map
markers = ('s', 'x', 'o', '^', 'v')
colors = ('red', 'blue', 'lightgreen', 'gray', 'cyan')
cmap = ListedColormap(colors[:len(np.unique(y))])
# plot the decision surface
x1_min, x1_max = X[:, 0].min() - 1, X[:, 0].max() + 1
x2_min, x2_max = X[:, 1].min() - 1, X[:, 1].max() + 1
xx1, xx2 = np.meshgrid(np.arange(x1_min, x1_max, resolution),
np.arange(x2_min, x2_max, resolution))
Z = classifier.predict(np.array([xx1.ravel(), xx2.ravel()]).T)
Z = Z.reshape(xx1.shape)
plt.contourf(xx1, xx2, Z, alpha=0.4, cmap=cmap)
plt.xlim(xx1.min(), xx1.max())
plt.ylim(xx2.min(), xx2.max())
# plot class samples
for idx, cl in enumerate(np.unique(y)):
plt.scatter(x=X[y == cl, 0], y=X[y == cl, 1],
alpha=0.8, c=[cmap(idx)],
marker=markers[idx], label=cl)
# +
plot_decision_regions(X, y, classifier=ppn)
plt.xlabel('sepal length [cm]')
plt.ylabel('petal length [cm]')
plt.legend(loc='upper left')
plt.tight_layout()
plt.show()
# -
# <br>
# <br>
# ## Adaptive linear neurons and the convergence of learning
# [[back to top](#Sections)]
# ### Implementing an adaptive linear neuron in Python
from ann import AdalineGD
# +
# AdalineGD?
# +
fig, ax = plt.subplots(nrows=1, ncols=2, figsize=(8, 4))
ada1 = AdalineGD(n_iter=10, eta=0.01).fit(X, y)
ax[0].plot(range(1, len(ada1.cost_) + 1), np.log10(ada1.cost_), marker='o')
ax[0].set_xlabel('Epochs')
ax[0].set_ylabel('log(Sum-squared-error)')
ax[0].set_title('Adaline - Learning rate 0.01')
ada2 = AdalineGD(n_iter=10, eta=0.0001).fit(X, y)
ax[1].plot(range(1, len(ada2.cost_) + 1), ada2.cost_, marker='o')
ax[1].set_xlabel('Epochs')
ax[1].set_ylabel('Sum-squared-error')
ax[1].set_title('Adaline - Learning rate 0.0001')
plt.tight_layout()
plt.show()
# -
# <br>
# <br>
# #### Standardizing features and re-training adaline
# standardize features
X_std = np.copy(X)
X_std[:,0] = (X[:,0] - X[:,0].mean()) / X[:,0].std()
X_std[:,1] = (X[:,1] - X[:,1].mean()) / X[:,1].std()
# +
ada = AdalineGD(n_iter=15, eta=0.01)
ada.fit(X_std, y)
plot_decision_regions(X_std, y, classifier=ada)
plt.title('Adaline - Gradient Descent')
plt.xlabel('sepal length [standardized]')
plt.ylabel('petal length [standardized]')
plt.legend(loc='upper left')
plt.tight_layout()
plt.show()
plt.plot(range(1, len(ada.cost_) + 1), ada.cost_, marker='o')
plt.xlabel('Epochs')
plt.ylabel('Sum-squared-error')
plt.tight_layout()
plt.show()
# -
# <br>
# <br>
# ### Large scale machine learning and stochastic gradient descent
# [[back to top](#Sections)]
from ann import AdalineSGD
# +
# AdalineSGD?
# +
ada = AdalineSGD(n_iter=15, eta=0.01, random_state=1)
ada.fit(X_std, y)
plot_decision_regions(X_std, y, classifier=ada)
plt.title('Adaline - Stochastic Gradient Descent')
plt.xlabel('sepal length [standardized]')
plt.ylabel('petal length [standardized]')
plt.legend(loc='upper left')
plt.tight_layout()
plt.show()
plt.plot(range(1, len(ada.cost_) + 1), ada.cost_, marker='o')
plt.xlabel('Epochs')
plt.ylabel('Average Cost')
plt.tight_layout()
plt.show()
# -
ada.partial_fit(X_std[0, :], y[0])
|
4_archmage/1.1.1. Perceptron and Adaline.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data analyses in Python
#
# Imagine you have a `.csv` file (could also be `.txt`, `.log`, `.dat`, `.sav`, etc.) with some data in it and want to analyze that data. Your `worklfow` to do so will most likely consist of different steps: `loading the data`, `"cleaning" the data`, `exploring the data`, `analyzing the data` and `visualization of the data/results`. In the following, we'll briefly go through all of these steps.
#
# As you might be aware, the first step is always to load necessary `packages` and/or `functions`. Most of the time, it is not clear what exactly is needed along the `worklfow`. Hence, starting with respective `packages/functions` we're sure about is a good idea. This is most likely based on the data you want to analyze. As we want to have a look at some data in a `.csv` file, `numpy` is a good starting point.
# However, to already provide you with the full list of `packages` we are going to explore, here you::
#
# - [numpy](https://numpy.org/)
# - [pandas](https://pandas.pydata.org/)
# - [scipy's stats module](https://docs.scipy.org/doc/scipy/reference/stats.html)
# - [statsmodels](http://www.statsmodels.org/stable/index.html)
# - [seaborn](seaborn.pydata.org)
#
# Ok, nuff said, let's go to work:
import numpy as np
# Using `np.` + `tab` provides us with a nice overview of `numpy`'s functionality:
np.
# In case, we are not sure about how a certain function works or simply want to know more about it, we can use the `help` function:
help(np.array)
# Based on our goal, the `genfromtxt` function looks useful, as we initially need to load the data from the `.csv` file:
my_data_numpy = np.genfromtxt('data/brain_size.csv', delimiter=';')
# With that, we generated a variable called `my_data_numpy`. Now, let's check its `type`.
type(my_data_numpy)
# It is a `numpy.ndarray` and within it, the data is stored:
my_data_numpy
# As we saw in the `help` function above, `objects` and `data types` come with certain `functionality`:
my_data_numpy.
# We can, for example, check the `shape`, that is the `dimensionality`, of the data:
my_data_numpy.shape
# This returns not a `numpy.ndarray`, but a `tuple`:
type(my_data_numpy.shape)
# It is also possible to `concatenate functionality`. E.g., we could `transpose` our `numpy.ndarray` and check its resulting `shape` within one command:
my_data_numpy.transpose().shape
# Is it possible to `view only certain parts` of the data, i.e. the second row? Yes, using `slicing`.
my_data_numpy[1]
# The output is a `numpy.ndarray` again:
type(my_data_numpy[1])
# If we want to be more specific, it is also possible to only view one value, i.e. the fourth value of the second row:
my_data_numpy[1, 3]
# Now, the `data type` has changed to `numpy.float64`:
type(my_data_numpy[1, 3])
# However, getting more than one value, i.e. multiple values of the second row, results in a `numpy.ndarray` again:
my_data_numpy[1, 3:6]
# Let's look at our data again:
my_data_numpy
# Even though it's a small dataset, there's already a lot going on: different `data types`, different `columns`, etc. and apparently not everything is `"numpy"` compatible. `Numpy` is a great tool and very powerful, building the foundation for a lot of `python libraries`, but in a lot of cases you might want to prefer `packages` that build upon `numpy` and are intended for specific purposes. A good example for that is the amazing `pandas` library that should be your first address for everything `data wrangling`. In particular, this refers to `tabular data`, that is `multiple observations` or `samples` described by a set of different `attributes` or `features`. The data can than be seen as a `2D table`, or `matrix`, with `columns` giving the different `attributes` of the data, and rows the `observations`. Let's try `pandas`, but at first, we have to import it:
import pandas as pd
# Now we can check its functionality:
pd.
# `read_csv` looks helpful regarding loading the data:
my_data_pandas = pd.read_csv('data/brain_size.csv', delimiter=',')
# What do we have? A `pandas dataframe`:
type(my_data_pandas)
# Before, our data was in `np.ndarray` format:
type(my_data_numpy)
# How does our data look now?
my_data_pandas
# Even though we already have more information as in the `numpy array`, e.g., `headers`, `strings` and `indexes`, it still looks off. What's the problem? Well, we see that our data has a `;` as a `delimiter`, but we indicated `,` as delimiter when loading our data. Therefore, it is important to carefully check your data and beware of its specifics. Let's reload our data with the fitting `delimiter`:
my_data_pandas = pd.read_csv('data/brain_size.csv', delimiter=';')
# Investigating our `dataframe`, we see that it worked as expected this time:
my_data_pandas
# Thinking about our `numpy.ndarray` version, we see a difference in the `shape` of the data, which is related to the `header`:
my_data_pandas.shape
# What can we do with our `dataframe`:
my_data_pandas.
# For example we can and should rename `columns` with uninformative names:
my_data_pandas.rename(columns={'Unnamed: 0': 'sub-id'})
# That looks a bit more informative, doesn't it? Let's have a look at our columns again
my_data_pandas.columns
# Wait a minute, it's not `renamed`. Did we do something wrong? Let's check the respective functionality:
help(my_data_pandas.rename)
# Checking the functionality more in depth, a `dataframe` with the new `column names` is returned, but the old one `not automatically changed`. Hence, we have to do it again, this overwriting the original `dataframe`:
my_data_pandas=my_data_pandas.rename(columns={'Unnamed: 0': 'sub-id'})
my_data_pandas.columns
# Pandas also allows the easy and fast `exploration` of our data:
my_data_pandas.describe()
# Unfortunately, not all `columns` are there. But why is that? We need to investigate the `columns` more closely, beginning with one that was included:
type(my_data_pandas['sub-id'])
# The data in the `columns` is a `pandas series`, not a `dataframe` or `numpy.ndarray`, again with its own functionality. Nevertheless, it was included in our `numerical summary`. Let's check the first missing `column`:
type(my_data_pandas['Hair'])
# Well, that's not very informative on its own, as it's also a `pandas series`, but was not included. Maybe the `data type` is the problem? Luckily, the `pandas dataframe` object comes with a helpful functionality:
my_data_pandas.dtypes
# And a bit more closely using `indexing`:
type(my_data_pandas['Hair'][0])
# The data in `my_data_pandas['Hair']` has the `type str` and as you might have already guessed: it's rather hard to compute `summary statistics` from a `str`. We could re-code it, but given there are only two values, this might not be very useful for our current aim:
my_data_pandas['Hair'].unique()
# What about the other `missing columns`, e.g., `height`?
type(my_data_pandas['Height'][0])
# The `data type` is yet again `str`, but how many values do we have?
my_data_pandas['Height'].unique()
# Hm, we can see that `height` contains `numerical values`. However, the `data type` is `str`. Here it can be useful to change the `data type`, using `pandas dataframe` object functionality:
my_data_pandas['Height'].astype(float)
# And we're getting another `error`. This time, it's related to a `missing data point`, which needs to be addressed before the `conversion` is possible. We can simply use the `replace` functionality to `replace` the `missing data point` to `NaN`, which should as allow to do the `conversion`:
my_data_pandas['Height'] = my_data_pandas['Height'].replace('.', np.nan)
my_data_pandas['Height'] = my_data_pandas['Height'].astype(float)
# Let's check if the `column` is now included in the `summary`:
my_data_pandas.describe()
# Now, we can do the same for the `Weight` column, `concatenating` all necessary functions in one line:
my_data_pandas['Weight'] = my_data_pandas['Weight'].replace('.', np.nan).astype(float)
# Is `Weight` now included?
my_data_pandas.describe()
# We can also compute one statistical value for one column, for example the `mean` using `numpy`:
np.mean(my_data_pandas['Weight'])
# But the same is also possible using inbuilt `pandas data frame` functionality:
my_data_pandas['Weight'].mean()
# We can do the same for the standard deviation:
np.std(my_data_pandas['Weight'])
my_data_pandas['Weight'].std()
# Here we can see, the same `functionality` can lead to different `results`, potentially based on `different implementations`. Thus, always make sure to check every part of your code and re-run it to see if you get the same outputs. As you can see here, using a `jupyter notebook` for your analyses, this is comparably straightforward. Additionally, you can document each step of your workflow, from data loading, inspection, changes, etc. . While you should of course always use `version control` on your data, the format we've explored nicely allows to redo your analyses (excluding the `computational reproducibility` and `numerical instability` aspect). On top of that, you can document the executed steps so that your future self and everyone else knows what's going on. Enough chit-chat, now that we've loaded and inspected our data, as well as fixed some errors it's time to do some statistics. To show you a few nice `packages` that are out there, we will run different `analyses` with different `packages`. We will explore `pingouin`, `scipy`, `statsmodels` and `seaborn`.
# <img src="https://github.com/raphaelvallat/pingouin/blob/master/docs/pictures/logo_pingouin.png?raw=true" height="300" width="700"/>
#
#
#
# ### _Pingouin is an open-source statistical package written in Python 3 and based mostly on Pandas and NumPy._
#
#
# - ANOVAs: one- and two-ways, repeated measures, mixed, ancova
# - Post-hocs tests and pairwise comparisons
# - Robust correlations
# - Partial correlation, repeated measures correlation and intraclass correlation
# - Linear/logistic regression and mediation analysis
# - Bayesian T-test and Pearson correlation
# - Tests for sphericity, normality and homoscedasticity
# - Effect sizes and power analysis
# - Parametric/bootstrapped confidence intervals around an effect size or a correlation coefficient
# - Circular statistics
# - Plotting: Bland-Altman plot, Q-Q plot, etc...
#
# **Pingouin is designed for users who want simple yet exhaustive statistical functions.**
#
#
# ##### **material scavenged from [10 minutes to Pingouin](https://pingouin-stats.org/index.html) and [the pingouin docs](https://pingouin-stats.org/api.html)
#
# Let's import the `package`:
import pingouin as pg
# ### Correlations
#
# "In the broadest sense correlation is any statistical association, though in common usage it most often refers to how close two variables are to having a linear relationship with each other" - [Wikipedia](https://en.wikipedia.org/wiki/Pearson_correlation_coefficient)
#
# `Pingouin` supports a variety of [measures of correlation](https://pingouin-stats.org/generated/pingouin.corr.html#pingouin.corr). When talking about `correlation`, we commonly mean the `Pearson correlation coefficient`, also referred to as `Pearson's r`:
#
# <img src="https://wikimedia.org/api/rest_v1/media/math/render/svg/93185aed3047ef42fa0f1b6e389a4e89a5654afa"/>
#
# Computing `Pearson's r` using `pingouin` is as easy as:
pearson_correlation = pg.corr(my_data_pandas['FSIQ'], my_data_pandas['VIQ'])
display(pearson_correlation)
cor_coeeficient = pearson_correlation['r']
# The output we got, is the `test summary`:
#
# - 'n' : Sample size (after NaN removal)
# - 'outliers' : number of outliers (only for 'shepherd' or 'skipped')
# - 'r' : Correlation coefficient
# - 'CI95' : [95% parametric confidence intervals](https://en.wikipedia.org/wiki/Confidence_interval)
# - 'r2' : [R-squared](https://en.wikipedia.org/wiki/Coefficient_of_determination)
# - 'adj_r2' : [Adjusted R-squared](https://en.wikipedia.org/wiki/Coefficient_of_determination#Adjusted_R2)
# - 'p-val' : one or two tailed p-value
# - 'BF10' : Bayes Factor of the alternative hypothesis (Pearson only)
# - 'power' : achieved power of the test (= 1 - type II error)
# What if we want to compute `pairwise correlations` between `columns` of a `dataframe`? With `pingouin` that's one line of code and we can even sort the results based on a `test statistic` of interest, e.g. `r2`:
pg.pairwise_corr(my_data_pandas, columns=['FSIQ', 'VIQ', 'Weight']).sort_values(by=['r2'], ascending=False)
# ### Before we calculate: `Testing statistical premises`
#
# Statistical procedures can be classfied into either [`parametric`](https://en.wikipedia.org/wiki/Parametric_statistics) or `non parametric` prcedures, which require different necessary preconditions to be met in order to show consistent/robust results.
# Generally people assume that their data follows a gaussian distribution, which allows for parametric tests to be run.
# Nevertheless it is essential to first test the distribution of your data to decide if the assumption of normally distributed data holds, if this is not the case we would have to switch to non parametric tests.
# ### [<NAME> normality test](https://pingouin-stats.org/generated/pingouin.normality.html#pingouin.normality)
#
# Standard procedure for testing `normal distribution` tests if the `distribution` of your data `deviates significantly` from a `normal distribution`.
# The function we're using returns the following information:
#
# - W : Test statistic
#
# - p : float
# P-value
#
# - normal : boolean
# True if data comes from a normal distribution.
pg.normality(my_data_pandas['Height'], alpha=.05)
# ### [Henze-Zirkler multivariate normality test](https://pingouin-stats.org/generated/pingouin.multivariate_normality.html#pingouin.multivariate_normality)
#
# The same procedure, but for [multivariate normal distributions](https://en.wikipedia.org/wiki/Multivariate_normal_distribution).
pg.multivariate_normality(my_data_pandas[['Height', 'Weight','VIQ']], alpha=.05)
# ### [Testing for homoscedasticity](https://pingouin-stats.org/generated/pingouin.homoscedasticity.html?highlight=homoscedasticity#pingouin.homoscedasticity)
#
#
# "In statistics, a sequence or a vector of random variables is homoscedastic /ˌhoʊmoʊskəˈdæstɪk/ if all random variables in the sequence or vector have the same finite variance." - Wikipedia
#
# returns:
#
# equal_var : boolean True if data have equal variance.
#
# p : float P-value.
#
# Note: This function first tests if the data are normally distributed using the Shapiro-Wilk test. If yes, then the homogeneity of variances is measured using the Bartlett test. If the data are not normally distributed, the Levene test, which is less sensitive to departure from normality, is used.
pg.homoscedasticity(my_data_pandas[['VIQ', 'FSIQ']], alpha=.05)
# ## Parametric tests
# ## Student's t-test: the simplest statistical test
#
# ### 1-sample t-test: testing the value of a population mean
#
# tests if the population mean of data is likely to be equal to a given value (technically if observations are drawn from a Gaussian distributions of given population mean).
#
#
# `pingouin.ttest` returns the T_statistic, the p-value, the [degrees of freedom](https://en.wikipedia.org/wiki/Degrees_of_freedom_(statistics), the [Cohen d effect size](https://en.wikiversity.org/wiki/Cohen%27s_d), the achieved [power](https://en.wikipedia.org/wiki/Power_(statistics%29) of the test ( = 1 - type II error (beta) = [P(Reject H0|H1 is true)](https://deliveroo.engineering/2018/12/07/monte-carlo-power-analysis.html)), and the [Bayes Factor](https://en.wikipedia.org/wiki/Bayes_factor) of the alternative hypothesis
#
#
#
#
pg.ttest(my_data_pandas['VIQ'],0)
# ### 2-sample t-test: testing for difference across populations
#
# We have seen above that the mean VIQ in the dark hair and light hair populations
# were different. To test if this is significant, we do a 2-sample t-test:
light_viq = my_data_pandas[my_data_pandas['Hair'] == 'light']['VIQ']
dark_viq = my_data_pandas[my_data_pandas['Hair'] == 'dark']['VIQ']
pg.ttest(light_viq, dark_viq)
# ### Plot achieved power of a paired T-test
#
# Plot the curve of achieved power given the effect size (Cohen d) and the sample size of a paired T-test.
# +
import matplotlib.pyplot as plt
import seaborn as sns
sns.set(style='ticks', context='notebook', font_scale=1.2)
d = 0.5 # Fixed effect size
n = np.arange(5, 80, 5) # Incrementing sample size
# Compute the achieved power
pwr = pg.power_ttest(d=d, n=n, contrast='paired', tail='two-sided')
# Start the plot
plt.plot(n, pwr, 'ko-.')
plt.axhline(0.8, color='r', ls=':')
plt.xlabel('Sample size')
plt.ylabel('Power (1 - type II error)')
plt.title('Achieved power of a paired T-test')
sns.despine()
# -
# ### Non parametric tests:
#
#
# Unlike the parametric test these do not require the assumption of normal distributions.
#
# "`Mann-Whitney U Test` (= Wilcoxon rank-sum test). It is the non-parametric version of the independent T-test.
# Mwu tests the hypothesis that data in x and y are samples from continuous distributions with equal medians. The test assumes that x and y are independent. This test corrects for ties and by default uses a continuity correction." - [mwu-function](https://pingouin-stats.org/generated/pingouin.mwu.html#pingouin.mwu)
#
# Test summary
#
# - 'W-val' : W-value
# - 'p-val' : p-value
# - 'RBC' : matched pairs rank-biserial correlation (effect size)
# - 'CLES' : common language effect size
pg.mwu(light_viq, dark_viq)
# "`Wilcoxon signed-rank test` is the non-parametric version of the paired T-test.
#
# The Wilcoxon signed-rank test tests the null hypothesis that two related paired samples come from the same distribution. A continuity correction is applied by default." - [wilcoxon - func](https://pingouin-stats.org/generated/pingouin.wilcoxon.html#pingouin.wilcoxon)
#
pg.wilcoxon(light_viq, dark_viq, tail='two-sided')
# ### `scipy.stats` - Hypothesis testing: comparing two groups
#
# For simple [statistical tests](https://en.wikipedia.org/wiki/Statistical_hypothesis_testing), it is also possible to use the `scipy.stats` sub-module of [`scipy`](http://docs.scipy.org/doc/).
from scipy import stats
# ### 1-sample t-test: testing the value of a population mean
#
# `scipy.stats.ttest_1samp` tests if the population mean of data is likely to be equal to a given value (technically if observations are drawn from a Gaussian distributions of given population mean). It returns the [T statistic](https://en.wikipedia.org/wiki/Student%27s_t-test), and the [p-value](https://en.wikipedia.org/wiki/P-value) (see the function's help):
stats.ttest_1samp(my_data_pandas['VIQ'], 100)
# With a p-value of 10^-28 we can claim that the population mean for the IQ (VIQ measure) is not 0.
# ### 2-sample t-test: testing for difference across populations
#
# We have seen above that the mean VIQ in the dark hair and light hair populations
# were different. To test if this is significant, we do a 2-sample t-test
# with `scipy.stats.ttest_ind`:
light_viq = my_data_pandas[my_data_pandas['Hair'] == 'light']['VIQ']
dark_viq = my_data_pandas[my_data_pandas['Hair'] == 'dark']['VIQ']
stats.ttest_ind(light_viq, dark_viq)
# ## Paired tests: repeated measurements on the same indivuals
#
# PIQ, VIQ, and FSIQ give 3 measures of IQ. Let us test if FISQ and PIQ are significantly different. We can use a 2 sample test:
stats.ttest_ind(my_data_pandas['FSIQ'], my_data_pandas['PIQ'])
# The problem with this approach is that it forgets that there are links
# between observations: FSIQ and PIQ are measured on the same individuals.
#
# Thus the variance due to inter-subject variability is confounding, and
# can be removed, using a "paired test", or ["repeated measures test"](https://en.wikipedia.org/wiki/Repeated_measures_design):
stats.ttest_rel(my_data_pandas['FSIQ'], my_data_pandas['PIQ'])
# This is equivalent to a 1-sample test on the difference::
stats.ttest_1samp(my_data_pandas['FSIQ'] - my_data_pandas['PIQ'], 0)
# T-tests assume Gaussian errors. We can use a [Wilcoxon signed-rank test](https://en.wikipedia.org/wiki/Wilcoxon_signed-rank_test), that relaxes this assumption:
stats.wilcoxon(my_data_pandas['FSIQ'], my_data_pandas['PIQ'])
# **Note:** The corresponding test in the non paired case is the [Mann–Whitney U test](https://en.wikipedia.org/wiki/Mann%E2%80%93Whitney_U), `scipy.stats.mannwhitneyu`.
# + [markdown] solution2="hidden" solution2_first=true
# ### Exercise 2
#
# * Test the difference between weights in people with dark and light hair.
# * Use non-parametric statistics to test the difference between VIQ in people with dark and light hair.
# + solution2="hidden"
light_weight = my_data_pandas[my_data_pandas['Hair'] == 'light']['Weight']
dark_weight = my_data_pandas[my_data_pandas['Hair'] == 'dark']['Weight']
stats.ttest_ind(light_weight, dark_weight, nan_policy='omit')
# + solution2="hidden"
stats.mannwhitneyu(light_viq, dark_viq)
# + [markdown] solution2="hidden"
# **Conclusion**: we find that the data does not support the hypothesis that people with dark and light hair have different VIQ.
# +
# Create solution here
# -
# # `statsmodels` - use "formulas" to specify statistical models in Python
#
# Use `statsmodels` to perform linear models, multiple factors or analysis of variance.
#
#
# ## A simple linear regression
#
# Given two set of observations, `x` and `y`, we want to test the hypothesis that `y` is a linear function of `x`.
#
# In other terms:
#
# $y = x * coef + intercept + e$
#
# where $e$ is observation noise. We will use the [statsmodels](http://statsmodels.sourceforge.net) module to:
#
# 1. Fit a linear model. We will use the simplest strategy, [ordinary least squares](https://en.wikipedia.org/wiki/Ordinary_least_squares) (OLS).
# 2. Test that $coef$ is non zero.
#
# First, we generate simulated data according to the model. Then we specify an OLS model and fit it:
from statsmodels.formula.api import ols
model = ols("FSIQ ~ VIQ", my_data_pandas).fit()
# **Note:** For more about "formulas" for statistics in Python, see the [statsmodels documentation](http://statsmodels.sourceforge.net/stable/example_formulas.html).
# We can inspect the various statistics derived from the fit:
print(model.summary())
# ### Terminology
#
# Statsmodels uses a statistical terminology: the `y` variable in statsmodels is called *endogenous* while the `x` variable is called *exogenous*. This is discussed in more detail [here](http://statsmodels.sourceforge.net/devel/endog_exog.html). To simplify, `y` (endogenous) is the value you are trying to predict, while `x` (exogenous) represents the features you are using to make the prediction.
# + [markdown] solution2="hidden" solution2_first=true
# ### Exercise 3
#
# Retrieve the estimated parameters from the model above.
# **Hint**: use tab-completion to find the relevant attribute.
# + solution2="hidden"
model.params
# +
# Create solution here
# -
# ## Categorical variables: comparing groups or multiple categories
model = ols("VIQ ~ Hair + 1", my_data_pandas).fit()
print(model.summary())
# ### Tips on specifying model
#
# ***Forcing categorical*** - the 'Hair' is automatically detected as a categorical variable, and thus each of its different values is treated as different entities.
#
# An integer column can be forced to be treated as categorical using:
#
# ```python
# model = ols('VIQ ~ C(Hair)', my_data_pandas).fit()
# ```
#
# ***Intercept***: We can remove the intercept using `- 1` in the formula, or force the use of an intercept using `+ 1`.
# ### Link to t-tests between different FSIQ and PIQ
#
# To compare different types of IQ, we need to create a "long-form" table, listing IQs, where the type of IQ is indicated by a categorical variable:
data_fisq = pd.DataFrame({'iq': my_data_pandas['FSIQ'], 'type': 'fsiq'})
data_piq = pd.DataFrame({'iq': my_data_pandas['PIQ'], 'type': 'piq'})
data_long = pd.concat((data_fisq, data_piq))
print(data_long[::8])
model = ols("iq ~ type", data_long).fit()
print(model.summary())
# We can see that we retrieve the same values for t-test and corresponding p-values for the effect of the type of IQ than the previous t-test:
stats.ttest_ind(my_data_pandas['FSIQ'], my_data_pandas['PIQ'])
# ## Multiple Regression: including multiple factors
#
# Consider a linear model explaining a variable `z` (the dependent
# variable) with 2 variables `x` and `y`:
#
# $z = x \, c_1 + y \, c_2 + i + e$
#
# Such a model can be seen in 3D as fitting a plane to a cloud of (`x`,
# `y`, `z`) points (see the following figure).
# +
from mpl_toolkits.mplot3d import Axes3D
x = np.linspace(-5, 5, 21)
# We generate a 2D grid
X, Y = np.meshgrid(x, x)
# To get reproducable values, provide a seed value
np.random.seed(1)
# Z is the elevation of this 2D grid
Z = -5 + 3*X - 0.5*Y + 8 * np.random.normal(size=X.shape)
# Plot the data
fig = plt.figure()
ax = fig.gca(projection='3d')
surf = ax.plot_trisurf(my_data_pandas['VIQ'].to_numpy(), my_data_pandas['PIQ'].to_numpy(),
my_data_pandas['FSIQ'].to_numpy(), cmap=plt.cm.plasma)
ax.view_init(20, -120)
ax.set_xlabel('X')
ax.set_ylabel('Y')
ax.set_zlabel('Z')
# -
model = ols('FSIQ ~ VIQ + PIQ', my_data_pandas).fit()
print(model.summary())
# ## Post-hoc hypothesis testing: analysis of variance (ANOVA)
#
# In the above iris example, we wish to test if the petal length is different between versicolor and virginica, after removing the effect of sepal width. This can be formulated as testing the difference between the coefficient associated to versicolor and virginica in the linear model estimated above (it is an Analysis of Variance, [ANOVA](https://en.wikipedia.org/wiki/Analysis_of_variance). For this, we write a **vector of 'contrast'** on the parameters estimated with an [F-test](https://en.wikipedia.org/wiki/F-test):
print(model.f_test([0, 1, -1]))
# Is this difference significant?
# + [markdown] solution2="hidden" solution2_first=true
# ### Exercise 4
#
# Going back to the brain size + IQ data, test if the VIQ of people with dark and light hair are different after removing the effect of brain size, height, and weight.
# + solution2="hidden"
data = pd.read_csv('data/brain_size.csv', sep=';', na_values=".")
model = ols("VIQ ~ Hair + Height + Weight + MRI_Count", data).fit()
print(model.summary())
# +
# Create solution here
# -
# ### Throwback to pingouin and pandas
#
# Remember `pingouin`? As briefly outlined, it can also compute `ANOVA`s and other types of models fairly easy. For example, let's compare `VIQ` between `light` and `dark` `hair`ed participants.
pg.anova(dv='VIQ', between='Hair', data=my_data_pandas,
detailed=True)
# It gets even better, `pandas` actually support some `pingouin` functions directly as an in-built `dataframe method`:
my_data_pandas.anova(dv='VIQ', between='Hair', detailed=True)
# # `seaborn` - use visualization for statistical exploration
#
# [Seaborn](http://stanford.edu/~mwaskom/software/seaborn/) combines simple statistical fits with plotting on `pandas dataframes`, `numpy arrays`, etc. .
# ## Pairplot: scatter matrices
#
# We can easily have an intuition on the interactions between continuous variables using `seaborn.pairplot` to display a scatter matrix:
import seaborn
seaborn.set()
seaborn.pairplot(my_data_pandas, vars=['FSIQ', 'PIQ', 'VIQ'], kind='reg')
# Categorical variables can be plotted as the hue:
seaborn.pairplot(my_data_pandas, vars=['FSIQ', 'VIQ', 'PIQ'], kind='reg', hue='Hair')
# ## lmplot: plotting a univariate regression
#
# A regression capturing the relation between one variable and another, e.g. `FSIQ` and `VIQ`, can be plotted using `seaborn.lmplot`:
seaborn.lmplot(y='FSIQ', x='VIQ', data=my_data_pandas)
# ### Robust regression
# Given that, in the above plot, there seems to be a couple of data points that are outside of the main cloud to the right, they might be outliers, not representative of the population, but driving the regression.
#
# To compute a regression that is less sensitive to outliers, one must use a [robust model](https://en.wikipedia.org/wiki/Robust_statistics). This is done in seaborn using ``robust=True`` in the plotting functions, or in `statsmodels` by replacing the use of the OLS by a "Robust Linear Model", `statsmodels.formula.api.rlm`.
# # Testing for interactions
#
# Does `FSIQ` increase more with `PIQ` for people with dark hair than with light hair?
seaborn.lmplot(y='FSIQ', x='PIQ', hue='Hair', data=my_data_pandas)
# The plot above is made of two different fits. We need to formulate a single model that tests for a variance of slope across the population. This is done via an ["interaction"](http://statsmodels.sourceforge.net/devel/example_formulas.html#multiplicative-interactions).
from statsmodels.formula.api import ols
result = ols(formula='FSIQ ~ PIQ + Hair + PIQ * Hair', data=my_data_pandas).fit()
print(result.summary())
# # Take home messages
#
# * Hypothesis testing and p-value give you the **significance** of an effect / difference
#
# * **Formulas** (with categorical variables) enable you to express rich links in your data
#
# * **Visualizing** your data and simple model fits matters!
#
# * **Conditioning** (adding factors that can explain all or part of the variation) is an important modeling aspect that changes the interpretation.
|
notebooks/python_statistics.ipynb
|
% -*- coding: utf-8 -*-
% ---
% jupyter:
% jupytext:
% text_representation:
% extension: .m
% format_name: light
% format_version: '1.5'
% jupytext_version: 1.14.4
% kernelspec:
% display_name: Octave
% language: octave
% name: octave
% ---
% # <center>Alkalmazott Analízis II.</center>
% ## <center>7. Feladatsor</center>
%
% <img src="coding.gif" width="700">
%
% <br>
% ## Lineáris többlépéses módszerek
%
% ### Konzisztencia
%
% 1.feladat. Taylor-sorfejtés útján határozzuk meg az $y_n-\frac{4}{3}y_{n-1}+\frac{1}{3}y_{n-2}=\frac{2}{3}hf_n$ kétlépéses módszer konzisztencia rendjét!
%
% 2.feladat. Mennyi a konzisztencia rendje az alábbi kétlépéses módszereknek?
%
% (a) $y_n-\frac{4}{3}y_{n-1}+\frac{1}{3}y_{n-2}=\frac{2}{3}hf_n$<br>
% (b) $y_n-y_{n-1}=h\Big(\frac{3}{2}f_{n-1}-\frac{1}{2}f_{n-2}\Big)$<br>
% (c) $y_n-y_{n-2}=2hf_{n-1}$
%
% 3.feladat. Határozzuk meg az $y_n+a_1y_{n-1}+a_2y_{n-2}=h(b_1f_{n-1}+b_2f_{n-2})$ explicit kétlépéses módszer együtthatóit úgy, hogy a konzisztencia rendje minél magasabb legyen.
% <br>
%
A=[1 1 0 0; 1 2 1 1; 1 4 2 4; 1 8 3 12];
b=[-1 0 0 0]';
A\b
% 4.feladat. Oldjuk meg az $y_n-4y_{n-1}+3y_{n-2}=-2hf_{n-2}$ módszerrel az
%
% \begin{cases}
% \dot y(t)= -y(t),\ \ \ t\in[0,1]&\\
% y(0)=1 &
% \end{cases}
%
% egyenletet $h=1/10$ választással. Nézzük meg minden egyes lépés után, hogy a hiba hogyan változik. Konvergens-e a módszer?
t=linspace(0,1,11);h=1/10;
y(1)=1; y(2)=exp(-h);
for i=3:11
y(i)=4*y(i-1)-(3-2*h)*y(i-2);
end
y;
exp(-t);
abs(y-exp(-t))
% ### Erős stabilitás
%
% 5.feladat. Az alábbi módszerek közül melyek lesznek erõsen stabilak?
%
% (a) $y_n+4y_{n-1}-5y_{n-2}=h(4f_{n-1}+2f_{n-2})$<br>
% (b) $y_n-y_{n-2}=\frac{h}{2}(f_n+4f_{n-1}+f_{n-2})$<br>
% (c) $y_n-\frac{4}{3}y_{n-1}+\frac{1}{3}y_{n-2}=\frac{2}{3}hf_n$
%
% 6.feladat. Mutassuk meg, hogy az Adams módszerek erõsen stabilak!
%
%5.feladat
%(a)
p1 = [1 4 -5];
r1 = roots(p1)
%(b)
p2 = [1 0 -1];
r2 = roots(p2)
%(c)
p3 = [1 -4/3 1/3];
r3 = roots(p3)
|
7.feladatsor.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import cv2
import numpy as np
import matplotlib.pyplot as plt
from PIL import Image
import torch
import tensorflow as tf
from torchvision import transforms
n = 16 #299
img = np.ones((120, 120), dtype=np.float32)
cv2.circle(img, center=(60, 60), radius=30, color=(0, 0, 0), thickness=1)
plt.imshow(img, cmap='gray')
img = torch.from_numpy(img)
# ## PIL
pil_img = transforms.functional.to_pil_image(img).resize((n, n), Image.BILINEAR)
plt.imshow(pil_img, cmap='gray')
# ## OpenCV
cv2_img = cv2.resize(img.numpy(), (n, n), interpolation=cv2.INTER_LINEAR)
plt.imshow(cv2_img, cmap='gray')
# ## TensorFlow
tensor_img = torch.unsqueeze(img, dim=2)
tensor_img = tf.image.resize(tensor_img, [n, n], antialias=True)
tensor_img = tf.squeeze(tensor_img).numpy()
plt.imshow(tensor_img, cmap='gray')
# ## PyTorch
torch_img = transforms.functional.to_pil_image(img)
torch_img = torch.unsqueeze(img, dim=2)
transform = transforms.Resize(n)
torch_img = transform(img)
torch_img = torch_img.numpy().reshape(n, n)
plt.imshow(torch_img, cmap='gray')
|
GLCIC/demo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="MTv5tY1iw9w6"
import numpy as np
import pandas as pd
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
from tensorflow.keras.layers.experimental.preprocessing import TextVectorization
from keras.optimizers import Adam
# + id="qYDRgFrjX0XX" outputId="2881d9c7-5154-4818-d995-2335f6077da3"
# Loading the dataset from the 'train' directory
batch_size = 512
seed = 1337 # Keep the seed same for both 'train' & 'validation' to avoid overlap
train_ds = keras.preprocessing.text_dataset_from_directory(
"../input/positionalembedding/hw2-ycbs-273-intro-to-prac-ml (1)/train",
batch_size=batch_size,
label_mode='int',
validation_split=0.2,
subset='training',
seed=seed)
val_ds = keras.preprocessing.text_dataset_from_directory(
"../input/positionalembedding/hw2-ycbs-273-intro-to-prac-ml (1)/train",
batch_size=batch_size,
label_mode='int',
validation_split=0.2,
subset='validation',
seed=seed)
text_only_train_ds = train_ds.map(lambda x, y: x)
# + id="-Hc61aLYvste"
from tensorflow.keras import layers
# from tensorflow.keras.layers.experimental.preprocessing import TextVectorization
max_length = 600
max_tokens = 20000
text_vectorization = layers.experimental.preprocessing.TextVectorization(
max_tokens=max_tokens,
output_mode="int",
output_sequence_length=max_length,
)
text_vectorization.adapt(text_only_train_ds)
int_train_ds = train_ds.map(lambda x, y: (text_vectorization(x), y))
int_val_ds = val_ds.map(lambda x, y: (text_vectorization(x), y))
# int_test_ds = test_ds.map(lambda x, y: (text_vectorization(x), y))
# + id="p_2Qjvw_vsqv"
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
class TransformerEncoder(layers.Layer):
def __init__(self, embed_dim, dense_dim, num_heads, **kwargs):
super().__init__(**kwargs)
self.embed_dim = embed_dim
self.dense_dim = dense_dim
self.num_heads = num_heads
self.attention = layers.MultiHeadAttention(
num_heads=num_heads, key_dim=embed_dim)
self.dense_proj = keras.Sequential(
[layers.Dense(dense_dim, activation="relu"),
layers.Dense(embed_dim),]
)
self.layernorm_1 = layers.LayerNormalization()
self.layernorm_2 = layers.LayerNormalization()
def call(self, inputs, mask=None):
if mask is not None:
mask = mask[:, tf.newaxis, :]
attention_output = self.attention(
inputs, inputs, attention_mask=mask)
proj_input = self.layernorm_1(inputs + attention_output)
proj_output = self.dense_proj(proj_input)
return self.layernorm_2(proj_input + proj_output)
def get_config(self):
config = super().get_config()
config.update({
"embed_dim": self.embed_dim,
"num_heads": self.num_heads,
"dense_dim": self.dense_dim,
})
return config
# + id="e7-gPn0WnQ1s"
class PositionalEmbedding(layers.Layer):
def __init__(self, sequence_length, input_dim, output_dim, **kwargs):
super().__init__(**kwargs)
self.token_embeddings = layers.Embedding(
input_dim=input_dim, output_dim=output_dim)
self.position_embeddings = layers.Embedding(
input_dim=sequence_length, output_dim=output_dim)
self.sequence_length = sequence_length
self.input_dim = input_dim
self.output_dim = output_dim
def call(self, inputs):
length = tf.shape(inputs)[-1]
positions = tf.range(start=0, limit=length, delta=1)
embedded_tokens = self.token_embeddings(inputs)
embedded_positions = self.position_embeddings(positions)
return embedded_tokens + embedded_positions
def compute_mask(self, inputs, mask=None):
return tf.math.not_equal(inputs, 0)
def get_config(self):
config = super().get_config()
config.update({
"output_dim": self.output_dim,
"sequence_length": self.sequence_length,
"input_dim": self.input_dim,
})
return config
# +
vocab_size = 20000
sequence_length = 600
embed_dim = 256
num_heads = 2
dense_dim = 32
inputs = keras.Input(shape=(None,), dtype="int64")
x = PositionalEmbedding(sequence_length, vocab_size, embed_dim)(inputs)
x = TransformerEncoder(embed_dim, dense_dim, num_heads)(x)
x = layers.GlobalMaxPooling1D()(x)
x = layers.Dropout(0.5)(x)
outputs = layers.Dense(4, activation="softmax")(x)
model = keras.Model(inputs, outputs)
# -
#setting compile and learning rate
adam = Adam(lr = 3e-4)
model.compile(optimizer="adam",
loss="sparse_categorical_crossentropy",
metrics=["accuracy"])
model.summary()
callbacks_list = [
keras.callbacks.EarlyStopping(
monitor="val_accuracy",
patience = 5,
),
keras.callbacks.ModelCheckpoint(
filepath="full_transformer_encoder.keras",
monitor="val_accuracy",
save_best_only=True,
)
]
model_history = model.fit(int_train_ds, validation_data=int_val_ds, epochs=10, callbacks=callbacks_list)
model = keras.models.load_model(
"full_transformer_encoder.keras",
custom_objects={"TransformerEncoder": TransformerEncoder,
"PositionalEmbedding": PositionalEmbedding})
print(f"Test acc: {model.evaluate(int_val_ds)[1]:.3f}")
import matplotlib.pyplot as plt
#draw plot for visualization
history_dict = model_history.history
loss_values = history_dict["loss"]
val_loss_values = history_dict["val_loss"]
epochs = range(1, len(loss_values) + 1)
plt.plot(epochs, loss_values, "bo", label="Training loss")
plt.plot(epochs, val_loss_values, "b", label="Validation loss")
plt.title("Training and validation loss")
plt.xlabel("Epochs")
plt.ylabel("Loss")
plt.legend()
plt.show()
plt.show()
# +
# Using the trained model to make prediction on unseen (test) data
# Here we use the 'adapted' text_vectorization layer and include it as part of a prediction_model
prediction_model = tf.keras.Sequential(
[text_vectorization, model])
prediction_model.compile(
loss=keras.losses.SparseCategoricalCrossentropy(),
optimizer='adam',
metrics=['accuracy'])
# Test it with `val_ds`, which yields raw strings
loss, accuracy = prediction_model.evaluate(val_ds)
print("Accuracy: {:2.2%}".format(accuracy))
# + id="hiDpXMt-vsGp"
# Read the test data in the form of a dataframe
df_test_data = pd.read_csv('../input/positionalembedding/data_test_df.csv')
inputs = df_test_data['data']
# + id="7S4Xsv4lvr5V"
# Make sure you use the 'prediction_model' and not the trained 'model' alone
# If you use the 'model' object, you will run int error as the data is still in the 'text' format and needs vectorization
predicted_scores = prediction_model.predict(inputs)
predicted_scores[0:5]
# + id="Ssq2FB1Lvr2y"
# populating the dataframe to make a submission on Kaggle
df_predictions = pd.DataFrame(predicted_scores, columns=['solution_' + str(i+1) for i in range(4)])
df_predictions.index.rename('Id', inplace=True)
df_predictions.head(30)
# + id="MpCGJ70Cvrr9"
# If using colab, then download this and submit on Kaggle
df_predictions.to_csv('df_predictions_6.csv')
# + id="DfU0toA4wwrK"
|
positional-embedding.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="wKY1p5iyzY0f"
# # Final project - Applied Mathematics
#
# + [markdown] id="VBCjvry_65ug"
# Members
# - <NAME> A00834191
# - <NAME> A01197044
# + [markdown] id="XZmEvzPT6SDj"
# In this python notebook we implemented several machine learning and statistical models to predict the inflation in Mexico, using the bi-weekly data recorded by INEGI.
# + [markdown] id="2vVDsmYRzYYm"
# ### INEGI data
# - INPC alogn with its components, extracted from [INEGI](https://www.inegi.org.mx/app/tabulados/default.aspx?nc=ca56_2018)
# - INPC per city, extracted from [INEGI](https://www.inegi.org.mx/app/tabulados/default.aspx?nc=ca62_2018)
# - INPC classifiying by object, extracted from [INEGI](https://www.inegi.org.mx/app/tabulados/default.aspx?nc=ca58_2018)
#
# [Inflation calculator](https://www.inegi.org.mx/app/indicesdeprecios/calculadorainflacion.aspx)
#
# [Price index](https://www.inegi.org.mx/app/indicesdeprecios/Estructura.aspx?idEstructura=112001300030&T=%C3%8Dndices%20de%20Precios%20al%20Consumidor&ST=Inflaci%C3%B3n%20Mensual)
#
# [INEGI main page (check graphics)](https://www.inegi.org.mx/temas/inpc/#Informacion_general)
# + [markdown] id="30kqU_3-4B1G"
# ## Process data
# + [markdown] id="A8M1Bb2-335n"
# ### Libraries
# + colab={"base_uri": "https://localhost:8080/"} id="mXgZHMlzCDBl" outputId="7a26ed80-ee12-4859-def7-c0f207d145ce"
# !pip install pystan==2.19.1.1 && pip install prophet
# + id="1Jo740twzDlq" colab={"base_uri": "https://localhost:8080/"} outputId="17d0a445-4ce5-4745-fe5c-209d71d002f3"
import datetime as dt
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import sklearn as sk
import tensorflow as tf
import warnings
from pandas.plotting import autocorrelation_plot
from prophet import Prophet
from prophet.plot import plot_plotly, plot_components_plotly
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import train_test_split
from statsmodels.tsa.arima_model import ARIMA
# + [markdown] id="4dPUqW5D9Nej"
# ### INEGI Dataframes
# + id="T7mGBOzY4IJR"
# INEGI Data
inpc_components_link = 'https://docs.google.com/spreadsheets/d/e/2PACX-1vSpdpTVzL6d_p4qhhkuVHxMMXIYKnITeyFtd98_e575z4MPiBtWdb8WKqmzXAlWYg/pub?gid=1239599080&single=true&output=csv'
inpc_per_city_link = 'https://docs.google.com/spreadsheets/d/e/2PACX-1vTJ_JokBZWk1rFvOWK-frzbLo9cOw_IzyLkXyFbGejKytzyBkuoaUrz3ydCL5PH3A/pub?gid=988073853&single=true&output=csv'
inpc_per_objects_link = 'https://docs.google.com/spreadsheets/d/e/2PACX-1vSTBQ9lwW-BX20fU8_wR0Ux2IzPTVe8yf6px5vFED9EzaijnzBKsjKn4jHRi2GEEQ/pub?gid=1466962329&single=true&output=csv'
# DataFrames
df_components = pd.read_csv(inpc_components_link)
df_city = pd.read_csv(inpc_per_city_link)
df_objects = pd.read_csv(inpc_per_objects_link)
# Parse dates
months = ['Ene', 'Feb', 'Mar', 'Abr',
'May', 'Jun', 'Jul', 'Ago',
'Sep', 'Oct', 'Nov', 'Dic']
def change_format_date(old_date):
date_splitted = old_date.split(' ')
day = '1' if date_splitted[0] == '1Q' else '15'
month = str(months.index(date_splitted[1]) + 1)
year = date_splitted[2]
parsed_date = '-'.join([year, month, day])
return parsed_date
df_components['Fecha'] = df_components['Fecha'].apply(lambda date: change_format_date(date))
df_city['Fecha'] = df_city['Fecha'].apply(lambda date: change_format_date(date))
df_objects['Fecha'] = df_objects['Fecha'].apply(lambda date: change_format_date(date))
# + [markdown] id="HDK4FsCC9hgG"
# ## Statistical models
# + [markdown] id="80vReg3Y9p-M"
# ### Linear Regression
# + id="BVNhevP79cqs"
def linear_regression(timeSerie, test_size=0.2):
# Given a time serie, train a model that uses the position of time and
# previous value to predict the next values.
# f(t, x_t) -> x_{t+1}
X = timeSerie.copy()
y = X.copy()
X.pop(0)
y.pop()
X = [[idx, x] for idx, x in enumerate(X)]
X, y = np.array(X), np.array(y)
# Train-test split
X_train, X_test, y_train, y_test = train_test_split(X, y,
test_size=test_size,
shuffle=False)
# Train model
model = LinearRegression().fit(X_train, y_train)
# Predict
y_predict = model.predict(X_test)
y_predict = []
last_known_t, last_known_x = X_train[-1]
for _ in range(len(X_test)):
y_hat = model.predict(np.array([[last_known_t, last_known_x]], dtype=object))
y_predict.append(y_hat)
last_known_t += 1
last_known_x = y_hat
return y_train, y_test, y_predict
# + [markdown] id="KmaKgaGL9s50"
# ### ARIMA
# + id="XVPDEI3ZHF30"
def arima(timeSerie, test_size=0.2, order=(5, 1, 0)):
# Given a time serie, train an ARIMA model to predict next values.
X = timeSerie.copy()
train_size_X = int(len(X) * (1 - test_size))
# Train-test split
X_train, X_test = X[:train_size_X], X[train_size_X:]
# Train model, and predict
y_predict = []
history = X_train.copy()
for _ in range(len(X_test)):
model = ARIMA(np.array(history, dtype=object), order=order)
model_fit = model.fit()
y_hat = model_fit.forecast()[0]
y_predict.append(y_hat)
history.append(y_hat)
return X_train, X_test, y_predict
# + [markdown] id="6RczwrweLbT4"
# ### Prophet
# + id="8IUp3s1vJfur"
def prophet(timeSerie, dates, test_size=0.2, periods=365):
X = timeSerie.copy()
train_size_X = int(len(X) * (1 - test_size))
# Train-test split
X_train, X_test = X[:train_size_X], X[train_size_X:]
dates_train, dates_test = dates[:train_size_X], dates[train_size_X:]
# Train model
df = pd.DataFrame({'ds': dates_train, 'y':X_train})
model = Prophet()
model.fit(df)
# Predict
future = model.make_future_dataframe(periods=len(X_test))
forecast = model.predict(future)
y_predict = forecast['yhat'].to_numpy(dtype=float)[-len(X_test):]
y_predict_upper = forecast['yhat_upper'].to_numpy(dtype=float)[-len(X_test):]
y_predict_lower = forecast['yhat_lower'].to_numpy(dtype=float)[-len(X_test):]
"""
# Plotting prophet
fig1 = model.plot(forecast)
fig1.show()
fig2 = model.plot_components(forecast)
fig2.show()
plot_plotly(model, forecast)
plot_components_plotly(model, forecast)
"""
return X_train, X_test, y_predict, y_predict_lower, y_predict_upper
# + [markdown] id="150Xb1T__0yb"
# ## Machine Learning models
#
# + [markdown] id="0UCfhwSO_WSM"
# ### Multi-Layer Perceptron
#
# + id="1ltWTMnD_n3Y"
def multi_layer_perceptron(timeSerie, look_back=10, test_size=0.2, epochs=100, verbose=False):
# Given a time serie, train a model that uses the last 'look_back' values
# to predict the next value.
# f(x_{t-4}, x_{t-3}, x_{t-2}, x_{t-1}, x_{t}) -> x_{t+1}
X, y = [], []
for idx in range(len(timeSerie) - look_back):
X.append(timeSerie[idx : idx + look_back])
y.append(timeSerie[idx + look_back])
X, y = np.array(X), np.array(y)
# Train-test split
X_train, X_test, y_train, y_test = train_test_split(X, y,
test_size=test_size,
shuffle=False)
# Architecture of model
model = tf.keras.Sequential([
tf.keras.layers.Dense(10, activation='relu', input_shape=(look_back,)),
tf.keras.layers.Dense(8, activation='relu'),
tf.keras.layers.Dense(1)
])
model.compile(loss=tf.keras.losses.mean_squared_error,
optimizer=tf.keras.optimizers.Adam(),
metrics=['mse', 'mae'])
# Train model
model.fit(X_train, y_train,
epochs=epochs,
verbose=verbose)
# Predict
y_predict = []
last_known_xs = X_train[-1]
for _ in range(len(X_test)):
y_hat = model.predict(np.array([last_known_xs]))
y_predict.append(y_hat[0])
last_known_xs = np.append(last_known_xs, y_hat[0])
last_known_xs = np.delete(last_known_xs, 0)
return y_train, y_test, y_predict
# + [markdown] id="BqwwTucXFWeU"
# ### Long Short Term-Memory
# + id="ZFb5mYC-FVYd"
def long_short_term_memory(timeSerie, look_back=10, test_size=0.2, batch_size=8, epochs=350, verbose=False):
# Given a time serie, train a model that uses the last 'look_back' values
# to predict the next value.
# f(x_{t-4}, x_{t-3}, x_{t-2}, x_{t-1}, x_{t}) -> x_{t+1}
X, y = [], []
for idx in range(len(timeSerie) - look_back):
x = timeSerie[idx : idx + look_back]
X.append([[t] for t in x])
y.append(timeSerie[idx + look_back])
X, y = np.array(X), np.array(y)
# Train-test split
X_train, X_test, y_train, y_test = train_test_split(X, y,
test_size=test_size,
shuffle=False)
# Architecture of model
model = tf.keras.Sequential([
tf.keras.layers.Input(shape=(look_back, 1)),
tf.keras.layers.LSTM(5, activation='tanh'),
tf.keras.layers.Dense(1)
])
model.compile(loss=tf.keras.losses.mean_squared_error,
optimizer=tf.keras.optimizers.Adam(),
metrics=['mse', 'mae'])
# Train model
model.fit(X_train, y_train,
epochs=epochs,
batch_size=batch_size,
verbose=verbose)
# Predict
y_predict = []
last_known_xs = X_train[-1]
for _ in range(len(X_test)):
y_hat = model.predict(np.array([last_known_xs]))
y_predict.append(y_hat[0])
last_known_xs = np.append(last_known_xs, y_hat)
last_known_xs = np.delete(last_known_xs, 0)
last_known_xs = [[x] for x in last_known_xs]
return y_train, y_test, y_predict
# + [markdown] id="MW1iZzikQmoG"
# ## Bemnchark
# + [markdown] id="lrytHXDFtbZ_"
# ### Plotting functions
# + id="3PDuV9e_gNsU"
def particular_plot(dates_train, dates_test, y_train, y_test, y_predict=None, model_name='', ticks=10, suffix='', y_predict_lower=None, y_predict_upper=None):
fig, ax = plt.subplots()
# Plotting
plt.ion()
plt.plot(dates_train, y_train, color='red', label='Train')
plt.plot(dates_test, y_test, color='blue', label='Test')
plt.plot(dates_test, y_predict, color='green', label='Prediction')
if y_predict_lower is not None:
plt.plot(dates_test, y_predict_lower, color='yellowgreen', label='Lower limit')
if y_predict_upper is not None:
plt.plot(dates_test, y_predict_upper, color='darkgreen', label='Upper limit')
# Configuration
plt.xlabel('Time')
plt.ylabel('INPC')
plt.title(model_name)
inv_ticks = (len(dates_train) + len(dates_test) - 1)//ticks + 1
ax.set_xticks(ax.get_xticks()[::inv_ticks])
ax.tick_params(axis="x", labelrotation=-60)
ax.legend()
# Show
plt.ioff()
plt.savefig(f'{model_name}{suffix}.png', dpi=333, transparent=True)
fig.show()
def show_plots(dates, y_train, y_test, y_predict=None, model_name='', percentage_closeup=0.95, ticks_normal=12, ticks_closeup=10, y_predict_lower=None, y_predict_upper=None):
dates_train = dates[:len(y_train)+1]
dates_test = dates[len(y_train) : len(y_train) + len(y_test)]
y_train_ = list(y_train)
y_train_.append(y_test[0])
particular_plot(dates_train, dates_test,
y_train_, y_test, y_predict,
model_name, ticks_normal,
y_predict_lower=y_predict_lower,
y_predict_upper=y_predict_upper)
closer_point = int(len(dates_train) * percentage_closeup)
dates_train_closeup = dates_train[closer_point:]
y_train_closeup = y_train_[closer_point:]
particular_plot(dates_train_closeup, dates_test,
y_train_closeup, y_test, y_predict,
model_name, ticks_closeup, suffix='_closeup',
y_predict_lower=y_predict_lower,
y_predict_upper=y_predict_upper)
# + [markdown] id="r4FUZdTCtgtI"
# ### Plotting each model
# + id="281ZEEINEM6w"
def get_series(days=None, biweeks=None):
if biweeks is None:
biweeks = days // 15 + 1 # Approximation of bi-weeks
dates = df_components['Fecha'].to_numpy()[-biweeks:]
timeSerie = list(df_components['INPC'].to_numpy())[-biweeks:]
return timeSerie, dates
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="IXEMDPXGQ76j" outputId="fc46f81a-0641-4f90-d54c-a0024224c517"
timeSerie, dates = get_series(biweeks=len(df_components['Fecha'].to_numpy()))
## Linear regression
y_train_lr, y_test_lr, y_predict_lr = linear_regression(timeSerie)
show_plots(dates, y_train_lr, y_test_lr, y_predict_lr, 'Linear Regression', 0.85)
## ARIMA
y_train_ar, y_test_ar, y_predict_ar = arima(timeSerie)
show_plots(dates, y_train_ar, y_test_ar, y_predict_ar, 'ARIMA', 0.85)
## Prophet
y_train_fb, y_test_fb, y_predict_fb, y_predict_lower_fb, y_predict_upper_fb = prophet(timeSerie, dates)
show_plots(dates, y_train_fb, y_test_fb, y_predict_fb, 'Prophet', 0.85, y_predict_lower=y_predict_lower_fb, y_predict_upper=y_predict_upper_fb)
## MLP
y_train_mlp, y_test_mlp, y_predict_mlp = multi_layer_perceptron(timeSerie, epochs=200)
show_plots(dates, y_train_mlp, y_test_mlp, y_predict_mlp, 'Multi-Layer Perceptron', 0.85)
## LSTM
y_train_lstm, y_test_lstm, y_predict_lstm = long_short_term_memory(timeSerie, epochs=200)
show_plots(dates, y_train_lstm, y_test_lstm, y_predict_lstm, 'Long Short Term-Memory', 0.85)
# + colab={"base_uri": "https://localhost:8080/", "height": 593} id="hSij8l4NKimL" outputId="399f4446-c219-4c16-f29d-9dabf1fc534d"
fig, ax = plt.subplots()
# Plotting
plt.ion()
dates_train_lr = dates[:len(y_train_lr)]
dates_test_lr = dates[len(y_train_lr) : len(y_train_lr) + len(y_test_lr)]
plt.plot(dates_train_lr, y_train_lr, color='red', label='Train')
plt.plot(dates_test_lr, y_test_lr, color='blue', label='Test')
models_data = [
[y_train_lr, y_test_lr, y_predict_lr, 'Linear Regression'],
[y_train_ar, y_test_ar, y_predict_ar, 'ARIMA'],
[y_train_fb, y_test_fb, y_predict_fb, 'Prophet'],
[y_train_mlp, y_test_mlp, y_predict_mlp, 'MLP'],
[y_train_lstm, y_test_lstm, y_predict_lstm, 'LSTM']
]
for y_train_model, y_test_model, y_predict_model, model_name in models_data:
plt.plot(dates[len(y_train_model) : len(y_train_model) + len(y_test_model)], y_predict_model, label=model_name)
# Configuration
plt.xlabel('Time')
plt.ylabel('INPC')
plt.title('Benchmark models')
ticks = 10
inv_ticks = (len(dates_train) + len(dates_test) - 1)//ticks + 1
ax.set_xticks(ax.get_xticks()[::inv_ticks])
ax.tick_params(axis="x", labelrotation=-60)
ax.legend()
# Show
plt.ioff()
plt.savefig('benchmark_models.png', dpi=333, transparent=True)
fig.show()
# + id="Vv5XnDXXMyWz" colab={"base_uri": "https://localhost:8080/", "height": 593} outputId="940f21ed-4a5d-4443-e7f2-5fb755b3abf4"
fig, ax = plt.subplots()
# Plotting
plt.ion()
percentage_closeup=0.85
closer_point = int(len(y_train_lr) * percentage_closeup)
dates_train_lr = dates[closer_point:len(y_train_lr)]
dates_test_lr = dates[len(y_train_lr) : len(y_train_lr) + len(y_test_lr)]
y_train_ = list(y_train)
y_train_.append(y_test[0])
plt.plot(dates_train_lr, y_train_lr[closer_point:], color='red', label='Train')
plt.plot(dates_test_lr, y_test_lr, color='blue', label='Test')
models_data = [
[y_train_lr, y_test_lr, y_predict_lr, 'Linear Regression'],
[y_train_ar, y_test_ar, y_predict_ar, 'ARIMA'],
[y_train_fb, y_test_fb, y_predict_fb, 'Prophet'],
[y_train_mlp, y_test_mlp, y_predict_mlp, 'MLP'],
[y_train_lstm, y_test_lstm, y_predict_lstm, 'LSTM']
]
for y_train_model, y_test_model, y_predict_model, model_name in models_data:
plt.plot(dates[len(y_train_model) : len(y_train_model) + len(y_test_model)], y_predict_model, label=model_name)
# Configuration
plt.xlabel('Time')
plt.ylabel('INPC')
plt.title('Benchmark models')
ticks = 10
inv_ticks = (len(dates_train) + len(dates_test) - 1)//ticks + 1
ax.set_xticks(ax.get_xticks()[::inv_ticks])
ax.tick_params(axis="x", labelrotation=-60)
ax.legend()
# Show
plt.ioff()
plt.savefig('benchmark_models_closeup.png', dpi=333, transparent=True)
fig.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="hpNzqBj3VZPB" outputId="32aa0fb0-02be-469b-c592-fbf5d0e73c75"
dates_train_lr[-1]
# + colab={"base_uri": "https://localhost:8080/"} id="LeDQrgmpZ0aa" outputId="74aff983-ee3e-4e3a-b6bc-aa73b12ac571"
y_predict_lstm
# + id="mOCDBw1iEoc8"
timeSerie, dates = get_series(biweeks=54)
plot_models(timeSerie, dates)
# + colab={"base_uri": "https://localhost:8080/"} id="1emDvdRXVmPO" outputId="85512c8d-e819-48d0-f65e-459f640e7292"
from scipy.stats import pearsonr, spearmanr
def calculate_errors(y_predict, y_test):
if isinstance(y_predict[0], np.ndarray):
y_predict = [ x[0] for x in y_predict ]
covariance = np.cov(y_predict, y_test)
corr, _ = pearsonr(y_predict, y_test)
corr_2, _ = spearmanr(y_predict, y_test)
return mean_squared_error(y_test, y_predict),covariance[0][1], corr, corr_2
print("""
\\begin{table}[H]
\\centering
\\begin{tabular}{|l|r|r|r|r|}
\\hline
\\multicolumn{1}{|c|}{\\textbf{Models}} & \\multicolumn{1}{c|}{\\textbf{Mean Square Error}} & \\multicolumn{1}{c|}{\\textbf{Covariance}} & \\multicolumn{1}{c|}{\\textbf{\\begin{tabular}[c]{@{}c@{}}Pearson\\\\ correlation\\end{tabular}}} & \\multicolumn{1}{c|}{\\textbf{\\begin{tabular}[c]{@{}c@{}}Spearman\\\\ correlation\\end{tabular}}} \\\\ \hline
""")
for _, y_test_model, y_predict_model, model_name in models_data:
mse, cov, pearson, spearman_c = calculate_errors(y_predict_model, y_test_model)
print("{} & {:.4f} & {:.4f} & {:.4f} & {:.4f}".format(model_name, mse, cov, pearson, spearman_c), end='\\\\ \\hline\n')
print("""
\\end{tabular}
\\caption{Benchmark results}
\\label{table:benchmark}
\\end{table}
""" )
|
AppliedMathematics_FinalProject.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exponential Distribution
#
# The Exponential distribution arises out of processes involving either temporal or spatial distances between events. The canonical example is that of time *between* arrivals of customers at a bank or coffee shop. The general name for such a generating process is a Poisson process and a number of probability distributions arise out of such processes. There are a number of key assumptions surrounding Poisson processes including:
#
# * continuous
# * constant average rate
# * independent events
#
# Therefore, if the *average* rate of phone calls is constant, if phone calls occur continuously (or approximately so), and if the phone calls are independent, the time between phone calls will follow an Exponential distribution. As with all such distributions, they are *models* and we can often use the models even if various assumptions are not met. We indicate a random variable $X$ distributed according to the Exponential distribution as $X \sim exp(\lambda)$ although it is often seen as $X \sim exp(\beta)$ where $\lambda = \frac{1}{\beta}$ so some care is required.
#
# For now, we want to tell a different generating story for the Exponential distribution. The Exponential distribution is characterized by a single parameter, $\lambda$, called the rate parameter. This is the rate of death per unit of time or rate of phone calls per unit of time. For now, we're going to take this as some kind of general failure rate.
#
# Let us assume we have a system of a single component that fails if that component fails. The component lasts anywhere from 1 to 365 days which we can model with a (discrete) Uniform distribution, which we saw in a previous section.
#
# We could simulate this process as we've done before and generate the following data (printing out the first 10 to inspect).
from numpy.random import randint, seed
import numpy as np
seed(1504120447)
data = randint(1, 366, 10000)
data[0:10]
# So what does that look like as a histogram of days to failure:
# %matplotlib inline
# +
import seaborn as sns
import matplotlib.pyplot as plt
sns.set(style="whitegrid")
# +
figure = plt.figure(figsize=(10,6))
axes = figure.add_subplot(1, 1, 1)
values, base = np.histogram( data, bins=90, density=True)
axes.plot( base[:-1], values, color="DimGray")
axes.set_title( "Distribution of Failures, 1 to 365 days")
axes.set_ylabel( "Probability Density")
axes.set_xlabel( "X")
axes.set_ylim(0, 0.005)
plt.show()
plt.close()
# -
# which is about as we'd expect. We have 10,000 processes whose behavior can be characterized by a Uniform distribution and the sum of those Uniform distributions is itself Uniform.
#
# Now we're going to change it up a bit. We assume that we have 10,000 systems as before except that each system has 3 components. Each component fails in 1 to 365 days but the entire system fails as soon as *any* of the 3 components fails. The parameters for `randint` are: min, max, number of samples. We want 3 in this case.
np.min( randint(1, 366, 3))
data = np.array([np.min(randint(1, 366, 3)) for _ in range(10000)])
data[0:10]
# And we'll plot this again:
# +
figure = plt.figure(figsize=(10,6))
axes = figure.add_subplot(1, 1, 1)
values, base = np.histogram( data, bins=91, density=True)
axes.plot( base[:-1], values, color="DimGray")
axes.set_title( "Distribution of Failures, 1 to 365 days")
axes.set_ylabel( "Probability Density")
axes.set_xlabel( "Days to Fail")
plt.show()
plt.close()
# -
# This is interesting. As the system has gotten more complicated, under our assumptions, it has gotten more fragile. Our systems have a higher probability of failing within 1 to 50 days than in 300 to 350 days which is very different than before. More systems fail earlier. What if we up the number of components to 10?
data = np.array([np.min(randint(1, 366, 10)) for _ in range(10000)])
data[0:10]
# +
figure = plt.figure(figsize=(10,6))
axes = figure.add_subplot(1, 1, 1)
values, base = np.histogram( data, bins=90, density=True)
axes.plot( base[:-1], values, color="DimGray")
axes.set_title( "Distribution of Failures, 1 to 365 days")
axes.set_ylabel( "Probability Density")
axes.set_xlabel( "Days to Fail")
plt.show()
plt.close()
# -
# At this point, we are almost exactly what the corresponding Exponential distribution would be. If we take the average of days to failure:
days_to_failure = np.mean( data) # beta
days_to_failure
# we get a parameter often called $\beta$. It turns out that $\lambda = \frac{1}{\beta}$ so `failures_per_day` here is the number of system failures we see per day.
failures_per_day = 1/days_to_failure # lambda
failures_per_day
# If we start with or have the rate, of course, we can always invert *that* to get the time we have to wait (on average) to see a failure or `days_to_failure`. Scipy's `expon` function is parameterized in terms of $\beta$ rather than $\lambda$. Pay attention to your library's documentation. If you have one, you can always get to the other.
from scipy.stats import expon
def drange(start, stop, step):
r = start
while r < stop:
yield r
r += step
# +
figure = plt.figure(figsize=(10,6))
axes = figure.add_subplot(1, 1, 1)
axes.set_title( "Exponential Distribution with $\lambda=%.2f$" % days_to_failure)
axes.set_ylabel( "Probability Density")
axes.set_xlabel( "Days to Fail")
xs = [expon.pdf( k, 0, days_to_failure) for k in drange( 1, 365, 1)]
axes.plot( xs, color="DimGray")
plt.show()
plt.close()
# -
# One think to keep an eye on are the different representations and presentations of probability distributions in different contexts.
#
# Mathematically, the Exponential distribution is parameterized in terms of $\lambda$ or wait time (for example, "days until a failture), the *code* from the Scipy library actually takes the rate, $\beta$, (for example, "failures per day"). You should make sure you read the documentation of the libraries you use as this sort of thing happens quite a lot when dealing with mathematical distributions.
#
# It's worth mentioning that we just derived the Exponential distribution using the rather famous *Central Limit Theorem*. But wait, what? Doesn't the Central Limit Theorem prove that a combination of random variables leads to a *Normal* distribution? Actually, no. That's just the most famous example. What the Central Limit Theorem actually says is that combinations of random variables tend towards specific *limiting distributions*. What limiting distribution depends on how the random variables are combined or interact. Here the variables interact in such a way (using `min`) that we get the Exponential distribution. Later, we will see what's required for the the Central Limit Theorem to result in a Normal distribution as well as a Log-Normal distribution.
# ## Estimation
#
# We've basically seen the main components of the Exponential distribution. Let's take the example of a call center.
#
# If your call center receives 2 calls per hour ($\beta$) *on average* then the time between calls ($\lambda$) is 0.5 hours on average. In order to estimate the Exponential distribution from data, you need to have data expressed in terms of events per unit just like calls per hour.
#
# The Method of Moments estimate of $\beta$ is:
#
# $\hat{\beta} = m_1$
#
# and as we've already mentioned, $\lambda = \frac{1}{\beta}$, so:
#
# $\hat{\lambda} = \frac{1}{\hat{beta}}$
#
# $mean = \beta = \frac{1}{\lambda}$
#
# but
#
# $variance = \beta^2 = \frac{1}{\lambda^2}$
#
# this is another one of those cases where *if* you are using data that is distributed according to the Exponential distribution, the data's variance is not what a descriptive statistics function is going to spit out.
# ## Memoryless
#
# One of the key characteristics of the Exponential distribution is that it is memoryless. If you wait for an event, stop, and then continue to wait, the distribution of time between events still follows the same Exponential distribution. For example, if you wait for a phone call, the Exponential distribution may say that from the last phone, it may take 5 minutes. 3 minutes in you may stop waiting and then resume waiting 10 minutes later. The next phone call is still most likely to take 5 minutes to occur on average.
#
# Because of this, the Exponential distribution is not a good model for situations that *require* memory. For example, the Exponential distribution is not a good model for the death rate of humans, largely because our components don't have binary states of failure.
# ## Shifted Exponential Distribution
#
# It is sometimes the case that there is a *minimum* amount of time that occurs between events, $L$, instead of 0. A good example of this is something like compile times or run times. Given a CPU and memory and absolutely no other processes running, it may take 10 minutes for a codebase to compile. If you recompile it, other processes may be running and so the compile time will be longer. In general, the compile times will have an Exponential distribution but the lowest value will be 10 minutes or $L$.
#
# In such a case, we need to model our process with a *Shifted* Exponential Distribution, $X \sim Exp(\lambda, L)$. The general approach to such modeling is to estimate $\hat{L}$ using the mininmum value of your data set, subtract $\hat{L}$ from your data set, estimate $\lambda$ and use those two parameters for your Exponential distribution.
#
# This illustrates up a very important point. Models like this are only good for *stationary* processes. As the codebase increases, we naturally expect $L$ to increase which means that overtime, we need to re-estimate our model. We'll talk more about this later.
# ## Gamma Distribution
#
# The Exponential distribution is a special case of the Gamma distribution. The Gamma distribution arises most generally in waiting times between Poisson distributed events (we'll talk about the Poisson distribution shortly...Poisson distributions are counts).
#
# In one of the parameterizations of the Gamma distribution (there are several), the Gamma distribution is described by a shape parameter $k$ and scale parameter $\theta$. If $X \sim Gamma(1, \frac{1}{\lambda})$ then $X ~ \sim Exp(\lambda)$. $k$ is the number of occurences of the event and $\theta$ is the mean number events per (time/space) unit. It is what we've called $\beta$ before (sorry, notation for these is not consistent--I do try, though).
#
# So what does this mean in terms of the Exponential distribution? Our Poisson distributed event is a single failure ("occurence" or "success") of any of the $n$ components. The Gamma distribution is useful in cases where $k > 1$.
#
# The Method of Moments estimators are:
#
# $\hat{k} = \frac{m_1^2}{m_2}$
#
# $\hat{\theta} = \frac{m_2}{m_1}$
#
# Note that libraries parameterize the Gamma distribution in a variety of ways. You may actually see $b$ instead of $k$ which is just $\frac{1}{k}$ (as is $scale = \beta = \theta = \frac{1}{\lambda}$. You may need to "play" with the library to see what it's doing if the documentation isn't clear.
#
# The mean and variance are:
#
# $mean = k\theta$
#
# $variance = k\theta^2$
#
# Again, there are other parameterizations and many uses for the Gamma distribution [Wikipedia](https://en.wikipedia.org/wiki/Gamma_distribution).
# ## Erlang Distribution
#
# The Erlang distribution describes waiting times in queueing systems. It was developed by <NAME> and is another special case of the Gamma distribution where $k$ is an integer. The Erlang distribution is the *sum* of $k$ independent variables $x_i \sim exp(\lambda)$. [Wikipedia](https://en.wikipedia.org/wiki/Erlang_distribution)
# ## Weibull Distribution
#
# Finally, the Weibull distribution is a variation on the Exponential distribution that is *not* memoryless. If you find that the process you wish to model *does* depend on time then you may want to consider a Weibull distribution.
#
# As with the Gamma distribution, the Weibull distribution has two parameters $k$, for "shape", and $\lambda$, for the "rate" (sometimes parameterized with the inverse or "scale"). The key advantage of the Weibull distribution over the Exponential is in the $k$ which has the following interpretations:
#
# * $k < 1$ - the failure rate is decreasing over time. This represents a "fail fast" process.
# * $k = 1$ - the failure rate is constant over time (the Exponential distribution).
# * $k > 1$ - the failure rate is increasing over time. This represents an "aging" process.
#
# There are entire books on the Weibull distribution and its uses. [Wikipedia](https://en.wikipedia.org/wiki/Weibull_distribution)
#
# The mean and variance as well as the parameters are a bit complicated because they are functions of $k$ (and because they depend on time, ultimately $t$).
#
# $mean = \lambda \Gamma(1 + \frac{1}{k})$
#
# $variance = \lambda^2 [\Gamma(1+\frac{2}{k}) - (\Gamma(1 + \frac{1}{k}))^2]$
#
# where $\Gamma()$ is the gamma *function*. If you decide that a Weibull distribution would be a good model for your data, you should consult a good reference.
|
fundamentals_2018.9/distributions/bestiary/exponential.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction to Regular Expressions in Python
#
#
# The term "regular expression" is a formal, linguistic term you might be interested to read about (https://en.wikipedia.org/wiki/Regular_language). For our purposes, regular expressions (AKA "regexes" or a "regex") is a way to formally describe some string of characters that we want to find. Regexes are an entirely separate DSL (domain-specific language) that we use inside Python, just like in the previous chapter we use SQL statements to communite with SQLite. While it's a bit of a drag to have to learn yet another language, the bonus is that you can use regular expressions in many places besides Python including with command line tools like `grep` and `awk` as well as within other languages like Perl and Rust.
#
# We can `import re` to use the Python regular expression module and use it to search text. For instance, in the tic-tac-toe exercise, we needed to see if the `--player` argument was exactly one character that was either an 'X' or an 'O'. Here's code that can do that:
for player in ['X', 'A', 'O', '5']:
if len(player) == 1 and (player == 'X' or player == 'O'):
print('{} OK'.format(player))
else:
print('{} bad'.format(player))
# A shorter way to write this could be:
for player in ['X', 'A', 'B', '5']:
if len(player) == 1 and player in 'XO':
print('{} OK'.format(player))
else:
print('{} bad'.format(player))
# It's not too onerous, but it quickly gets worse as we get more complicated requirements. In that same exercise, we needed to check if `--state` was exactly 9 characters composed entirely of ".", "X", "O":
for state in ['XXX...OOO', 'XXX...OOA']:
#print([(x, x in 'XO.') for x in state])
print(state, 'OK' if len(state) == 9 and
all(map(lambda x: x in 'XO.', state)) else 'No')
# Can we make this simpler? Well, when we were starting out with the Unix command line, one exercise had us using `grep` to look for lines that start with vowels. One solution was:
#
# ````
# $ grep -io '^[aeiou]' scarlet.txt | sort | uniq -c
# 59 A
# 10 E
# 91 I
# 20 O
# 6 U
# 651 a
# 199 e
# 356 i
# 358 o
# 106 u
# ````
#
# We used square brackets `[]` to enumerate all the vowels `[aeiou]` and used the `-i` flag to `grep` to indicate it should match case **insensitively**. Additionally, the `^` indicated that the match should occur at the start of the string. Those were regular expressions we were using.
#
# The regex allows us to **describe** what we want rather than **implement** the code to find what we want. We can create a class of allowed characters with `[XO]` and additionally constraint it to be exactly one character wide with `{1}` after the class. (Note that `{}` for match length can be in the format `{exactly}`, `{min,max}`, `{min,}`, or `{,max}`.)
#
# To use regular expressions:
import re
# Now let's describe our pattern using a character class `[XO]` and the length `{1}`:
for player in ['X', 'O', 'A']:
print(player, re.match('[XO]{1}', player))
# We can extend this to our state problem:
state = 'XXX...OOO'
print(state, re.match('[XO.]{9}', state))
state = 'XXX...OOA'
print(state, re.match('[XO.]{9}', state))
# ## Building regular expressions
#
# How do we match a number?
print(re.match('1', '1'))
# But that only works for just "1"
print(re.match('2', '1'))
# How do we match all the numbers from 0 to 9? We can create a character class that contains that range:
print(re.match('[0-9]', '1'))
# There is a short-hand for the character class `[0-9]` that is `\d` (digit)
re.match('\d', '1')
# But this only matches the first number we see:
re.match('\d', '123')
# We can use `{}` to indicate `{min,max}`, `{min,}`, `{,max}`, or `{exactly}`:
print(re.match('\d{1,4}', '8005551212'))
print(re.match('\d{1,}', '8005551212'))
print(re.match('\d{,5}', '8005551212'))
print(re.match('\d{8}', '8005551212'))
# ## match vs search
#
# Note that we are using `re.match` which requires the regex to match **at the beginning of the string**:
print(re.match('\d{10}', 'That number to call is 8005551212!'))
# If you want to match anywhere in the string, use `re.search`:
for s in ['123', 'abc456', '789def']:
print(s, re.search('\d{3}', s))
# To anchor your match to the beginning of the string, use the `^`:
for s in ['123', 'abc456', '789def']:
print(s, re.search('^\d{3}', s))
# Use `$` for the end of the string:
for s in ['123', 'abc456', '789def']:
print(s, re.search('\d{3}$', s))
# And use both to say that the entire string from beginning to end must match:
for s in ['123', 'abc456', '789def']:
print(s, re.search('^\d{3}$', s))
# Returning to our previous problem of trying to see if we got *exactly* one "X" or "O" for our tic-tac-toe player:
for player in ['X', 'O', 'XX', 'OO']:
print(player, re.match('[XO]{1}', player))
# The problem is that there is a match of `[XO]{1}` in the strings "XX" and "OO" -- there *is* exactly one X or O at the beginning of those strings. Since `re.match` already anchors the match to the beginning of the string, we could just add `$` to the end of our pattern:
for player in ['X', 'O', 'XX', 'OO']:
print(player, re.match('[XO]{1}$', player))
# Or use `re.search` with `^$` to indicate a match over the entire string:
for player in ['X', 'O', 'XX', 'OO']:
print(player, re.search('^[XO]{1}$', player))
# ## Matching SSNs and Dates
#
# What if we wanted to recognize a US SSN (social security number)?
# We will use `re.compile` to create the regex and use it in a `for` loop:
ssn_re = re.compile('\d{3}-\d{2}-\d{4}')
for s in ['123456789', '123-456-789', '123-45-6789']:
print('{}: {}'.format(s, ssn_re.match(s)))
# SSNs always use a dash (`-`) as a number separator, but dates do not.
date_re = re.compile('\d{4}-\d{2}-\d{2}')
dates = ['1999-01-02', '1999/01/02']
for d in dates:
print('{}: {}'.format(d, date_re.match(d)))
# Just as we created a character class with `[0-9]` to represent all the numbers from 0 to 9, we can create a class to represent the separators "/" and "-" with `[/-]`. As regular expressions get longer, it makes sense to break each unit onto a different line and use Python's literal string expression to join them into a single string. As a bonus, we can comment on each unit of the regex.
# +
date_re = re.compile('\d{4}' # year
'[/-]' # separator
'\d{2}' # month
'[/-]' # separator
'\d{2}') # day
dates = ['1999-01-02', '1999/01/02']
for d in dates:
print('{}: {}'.format(d, date_re.match(d)))
# -
# You may notice that certain elements are repeated. If we followed DRY (Don't Repeat Yourself), we might want to make variables to hold each piece, but then we could not use the literal string joining trick above. In that case, just go back to using `+` to join strings:
# +
sep = '[/-]'
four_digits = '\d{4}'
two_digits = '\d{2}'
date_re = re.compile(four_digits + # year
sep + # separator
two_digits + # month
sep + # separator
two_digits) # day
dates = ['1999-01-02', '1999/01/02']
for d in dates:
print('{}: {}'.format(d, date_re.match(d)))
# -
# Dates are not always written YYYY-MM-DD where the month/day are zero-padded left, e.g., "01" instead of "1". How could we handle that? Change our `two_digits` from `\d{2}` (exactly two) to `\d{1,2}` (one or two):
# +
sep = '[/-]'
four_digits = '\d{4}'
two_digits = '\d{1,2}'
date_re = re.compile(four_digits + # year
sep + # separator
two_digits + # month
sep + # separator
two_digits) # day
dates = ['1999-01-01', '1999/01/02', '1999/1/2']
for d in dates:
print('{}: {}'.format(d, date_re.match(d)))
# -
# If we wanted to extract each part of the date (year, month, day), we can use parentheses `()` around the parts we want to capture into `groups`. The group "0" is the whole string that was match, and they are numbered sequentially after that for each group.
#
# Can you change the regex to match all three strings?
# +
date_re = re.compile('(\d{4})' # capture year (group 1)
'[/-]' # separator
'(\d{1,2})' # capture month (group 2)
'[/-]' # separator
'(\d{1,2})') # capture day (group 3)
dates = ['1999-01-02', '1999/1/2', '1999.01.01']
for d in dates:
match = date_re.match(d)
print('{}: {}'.format(d, 'match' if match else 'miss'))
if match:
print(match.groups())
print('year:', match.group(1))
print()
# -
# As we add more groups, it can be confusing to remember them by their positions, so we can name them with `?P<name>` just inside the opening paren.
# +
date_re = re.compile('(?P<year>\d{4})'
'[/-]'
'(?P<month>\d{1,2})'
'[/-]'
'(?P<day>\d{1,2})')
dates = ['1999-1-2', '1999/01/02', '1999.01.01']
for d in dates:
match = date_re.match(d)
print('{}: {}'.format(d, 'match' if match else 'miss'))
if match:
print('{} = year "{}" month "{}" day "{}"'.format(d,
match.group('year'),
match.group('month'),
match.group('day')))
print()
# -
# ## Matching US Phone Numbers
#
# What if we wanted to match a US phone number?
phone_re = re.compile('(\d{3})' # area code
' ' # a space
'\d{3}' # prefix
'-' # dash
'\d{4}') # line number
print(phone_re.match('(800) 555-1212'))
# Why didn't that work?
#
# What do those parentheses do again? They group!
#
# So we need to indicate that the parens are literal things to match by using backslashes `\` to escape them.
phone_re = re.compile('\(' # left paren
'\d{3}' # area code
'\)' # right paren
' ' # space
'\d{3}' # prefix
'-' # dash
'\d{4}') # line number
print(phone_re.match('(800) 555-1212'))
# We could also use character classes to make this more readable:
# +
phone_re = re.compile('[(]' # left paren
'\d{3}' # area code
'[)]' # right paren
' ' # space
'\d{3}' # prefix
'-' # dash
'\d{4}') # line number
print(phone_re.match('(800) 555-1212'))
# -
# There is not always a space after the area code, and it may sometimes it may be more than one space (or a tab?). We can use the `\s` to indicate any type of whitespace and `*` to indicate zero or more:
phone_re = re.compile('[(]' # left paren
'\d{3}' # area code
'[)]' # right paren
'\s*' # zero or more spaces
'\d{3}' # prefix
'-' # dash
'\d{4}') # line number
phones = ['(800)555-1212', '(800) 555-1212', '(800) 555-1212']
for phone in phones:
print('{}\t{}'.format(phone, phone_re.match(phone)))
# When the parens around the area code are optional, usually there is a dash to separate the area code:
# +
phone_re = re.compile('[(]?' # optional left paren
'\d{3}' # area code
'[)]?' # optional right paren
'[-]?' # optional dash
'\s*' # zero or more whitespace
'\d{3}' # prefix
'-' # dash
'\d{4}') # line number
phones = ['(800)555-1212', '(800) 555-1212', '800-555-1212']
for phone in phones:
print('{}\t{}'.format(phone, phone_re.match(phone)))
# -
# This has the affect of matching a dash after parens which is generally not a valid format:
# +
phone_re = re.compile('[(]?' # optional left paren
'\d{3}' # three digits
'[)]?' # optional right paren
'[-]?' # optional dash
'\s*' # zero or more spaces
'\d{3}' # three digits
'-' # dash
'\d{4}') # four digits
phone_re.match('(800)-555-1212')
# -
# We really have to create two regexes to handle these cases:
# +
phone_re1 = re.compile('[(]'
'\d{3}'
'[)]'
'\s*'
'\d{3}'
'-'
'\d{4}')
phone_re2 = re.compile('\d{3}'
'-'
'\d{3}'
'-'
'\d{4}')
phones = ['(800)555-1212', '(800) 555-1212', '800-555-1212', '(800)-555-1212']
for phone in phones:
match1 = phone_re1.match(phone)
match2 = phone_re2.match(phone)
print('{}\t{}'.format(phone, 'match' if match1 or match2 else 'miss'))
# -
# I worked with a graphic artist who always insisted on using dots as the number separator, and sometimes there are no separators at all. The combination of these two regexes find the valid formats and skip the invalid one.
# +
phone_re1 = re.compile('[(]'
'\d{3}'
'[)]'
'\s*'
'\d{3}'
'[.-]'
'\d{4}')
phone_re2 = re.compile('\d{3}'
'[.-]?'
'\d{3}'
'[.-]?'
'\d{4}')
phones = ['8005551212', '(800)555-1212', '(800) 555-1212',
'800-555-1212', '(800)-555-1212', '800.555.1212']
for phone in phones:
match = phone_re1.match(phone) or phone_re2.match(phone)
print('{}\t{}'.format(phone, 'match' if match else 'miss'))
# -
# OK, now let's normalize the numbers by using parens to capture the area code, prefix, and line number and then create a standard representation.
# +
phone_re1 = re.compile('[(]'
'(\d{3})' # group 1
'[)]'
'\s*'
'(\d{3})' # group 2
'[.-]'
'(\d{4})') # group 3
phone_re2 = re.compile('(\d{3})' # group 1
'[.-]?'
'(\d{3})' # group 2
'[.-]?'
'(\d{4})') # group 3
phones = ['8005551212', '(800)555-1212', '(800) 555-1212',
'800-555-1212', '(800)-555-1212', '800.555.1212']
for phone in phones:
match = phone_re1.match(phone) or phone_re2.match(phone)
standard = '{}-{}-{}'.format(match.group(1),
match.group(2),
match.group(3)) if match else 'miss'
print('{}\t{}'.format(phone, standard))
# -
# And if we add named capture groups...
# +
phone_re1 = re.compile('[(]'
'(?P<area_code>\d{3})'
'[)]'
'\s*'
'(?P<prefix>\d{3})'
'[.-]'
'(?P<line_num>\d{4})')
phone_re2 = re.compile('(?P<area_code>\d{3})'
'[.-]?'
'(?P<prefix>\d{3})'
'[.-]?'
'(?P<line_num>\d{4})')
phones = ['8005551212', '(800)555-1212', '(800) 555-1212',
'800-555-1212', '(800)-555-1212', '800.555.1212']
for phone in phones:
match = phone_re1.match(phone) or phone_re2.match(phone)
standard = '{}-{}-{}'.format(match.group('area_code'),
match.group('prefix'),
match.group('line_num')) if match else 'miss'
print('{}\t{}'.format(phone, standard))
# -
# And if we add named capture groups and named groups in `format`:
# +
phone_re1 = re.compile('[(]'
'(?P<area_code>\d{3})'
'[)]'
'\s*(?P<prefix>\d{3})'
'[.-]'
'(?P<line_num>\d{4})')
phone_re2 = re.compile('(?P<area_code>\d{3})'
'[.-]?'
'(?P<prefix>\d{3})'
'[.-]?'
'(?P<line_num>\d{4})')
phones = ['8005551212', '(800)555-1212', '(800) 555-1212',
'800-555-1212', '(800)-555-1212', '800.555.1212']
for phone in phones:
match = phone_re1.match(phone) or phone_re2.match(phone)
tmpl = '{area_code}-{prefix}-{line_num}'
standard = tmpl.format(prefix=match.group('prefix'),
area_code=match.group('area_code'),
line_num=match.group('line_num')) if match else 'miss'
print('{}\t{}'.format(phone, standard))
# -
# ## ENA Metadata
#
# Let's examine the ENA metadata from the XML parsing example. We see there are many ways that latitude/longitude have been represented:
#
# ````
# $ ./xml_ena.py *.xml | grep lat_lon
# attr.lat_lon : 27.83387,-65.4906
# attr.lat_lon : 29.3 N 122.08 E
# attr.lat_lon : 28.56_-88.70377
# attr.lat_lon : 39.283N 76.611 W
# attr.lat_lon : 78 N 5 E
# attr.lat_lon : missing
# attr.lat_lon : 0.00 N, 170.00 W
# attr.lat_lon : 11.46'45.7" 93.01'22.3"
# ````
# How can we go about parsing all the various ways this data has been encoded? Regular expressions provide us a way to describe in very specific way what we want.
#
# Let's start just with the idea of matching a number (where "number" is a string that could be parsed into a number) like "27.83387":
print(re.search('\d', '27.83387'))
# The `\d` pattern means "any number" which is the same as `[0-9]` where the `[]` creates a class of characters and `0-9` expands to all the numbers from zero to nine. The problem is that it only matches one number, `2`. Change it to `\d+` to indicate "one or more numbers":
re.search('\d+', '27.83387')
# Now let's capture the decimal point:
re.search('\d+.', '27.83387')
# You might think that's perfect, but the `.` has a special meaning in regex. It means "one of anything", so it matches this, too:
re.search('\d+.', '27x83387')
# To indicate we want a literal `.` we have to make it `\.` (backslash-escape):
print(re.search('\d+\.', '27.83387'))
print(re.search('\d+\.', '27x83387'))
# Notice that the second try returns nothing.
#
# To capture the bit after the `.`, add more numbers:
re.search('\d+\.\d+', '27.83387')
# But we won't always see floats. Can we make this regex match integers, too? We can indicate that part of a pattern is optional by putting a `?` after it. Since we need more than one thing to be optional, we need to wrap it in parens:
print(re.search('\d+\.\d+', '27'))
print(re.search('\d+(\.\d+)?', '27'))
print(re.search('\d+(\.\d+)?', '27.83387'))
# What if there is a negative symbol in front? Add `-?` (an optional dash) at the beginning:
print(re.search('-?\d+(\.\d+)?', '-27.83387'))
print(re.search('-?\d+(\.\d+)?', '27.83387'))
print(re.search('-?\d+(\.\d+)?', '-27'))
print(re.search('-?\d+(\.\d+)?', '27'))
# Sometimes we actually find a `+` at the beginning, so we can make an optional character class `[+-]?`:
print(re.search('[+-]?\d+(\.\d+)?', '-27.83387'))
print(re.search('[+-]?\d+(\.\d+)?', '+27.83387'))
print(re.search('[+-]?\d+(\.\d+)?', '27.83387'))
# Now we can match things that basically look like a floating point number or an integer, both positive and negative.
#
# Usually the data we want to find it part of a larger string, however, and the above fails to capture more than one thing, e.g.:
print(re.search('[+-]?\d+(\.\d+)?', 'Lat is "-27.83387" and lon is "+132.43."'))
# We really need to match more than once using our pattern matching to extract data. We saw earlier that we can use parens to group optional patterns, but the parens also end up creating a **capture group** that we can refer to by position:
re.findall('([+-]?\d+(\.\d+)?)','Lat is "-27.83387" and lon is "+132.43."')
# OK, it was a bit unexpected that we have matches for both the whole float and the decimal part. This is because of the dual nature of the parens, and in the case of using them to group the optional part we are also creating another capture. If we change `()` to `(?:)`, we make this a non-capturing group:
re.findall('([+-]?\d+(?:\.\d+)?)', 'Lat is "-27.83387" and lon is "+132.43."')
# There are many resources you can use to thoroughly learn regular expressions, so I won't try to cover them completely here. I will mostly try to introduce the general idea and show you some useful regexes you could steal.
#
# Here is an example of how you can embed regexes in your Python code. This version can parse all the versions of latitude/longitude shown above. This code uses parens to create capture groups which it then uses `match.group(n)` to extract:
#
# ````
# $ cat -n parse_lat_lon.py
# 1 #!/usr/bin/env python3
# 2
# 3 import os
# 4 import re
# 5 import sys
# 6
# 7 args = sys.argv[1:]
# 8
# 9 if len(args) != 1:
# 10 print('Usage: {} FILE'.format(os.path.basename(sys.argv[0])))
# 11 sys.exit(1)
# 12
# 13 file = args[0]
# 14
# 15 float_ = r'[+-]?\d+\.*\d*'
# 16 ll1 = re.compile('(' + float_ + ')\s*[,_]\s*(' + float_ + ')')
# 17 ll2 = re.compile('(' + float_ + ')(?:\s*([NS]))?(?:\s*,)?\s+(' + float_ +
# 18 ')(?:\s*([EW])?)')
# 19 loc_hms = r"""
# 20 \d+\.\d+'\d+\.\d+"
# 21 """.strip()
# 22 ll3 = re.compile('(' + loc_hms + ')\s+(' + loc_hms + ')')
# 23
# 24 for line in open(file):
# 25 line = line.rstrip()
# 26 ll_match1 = ll1.search(line)
# 27 ll_match2 = ll2.search(line)
# 28 ll_match3 = ll3.search(line)
# 29
# 30 if ll_match1:
# 31 lat, lon = ll_match1.group(1), ll_match1.group(2)
# 32 lat = float(lat)
# 33 lon = float(lon)
# 34 print('lat = {}, lon = {}'.format(lat, lon))
# 35 elif ll_match2:
# 36 lat, lat_dir, lon, lon_dir = ll_match2.group(
# 37 1), ll_match2.group(2), ll_match2.group(
# 38 3), ll_match2.group(4)
# 39 lat = float(lat)
# 40 lon = float(lon)
# 41
# 42 if lat_dir == 'S':
# 43 lat *= -1
# 44
# 45 if lon_dir == 'W':
# 46 lon *= -1
# 47 print('lat = {}, lon = {}'.format(lat, lon))
# 48 elif ll_match3:
# 49 lat, lon = ll_match3.group(1), ll_match3.group(2)
# 50 print('lat = {}, lon = {}'.format(lat, lon))
# 51 else:
# 52 print('No match: "{}"'.format(line))
# $ cat lat_lon.txt
# attr.lat_lon : 27.83387,-65.4906
# attr.lat_lon : 29.3 N 122.08 E
# attr.lat_lon : 28.56_-88.70377
# This line will not be included
# attr.lat_lon : 39.283N 76.611 W
# attr.lat_lon : 78 N 5 E
# attr.lat_lon : missing
# attr.lat_lon : 0.00 N, 170.00 W
# attr.lat_lon : 11.46'45.7" 93.01'22.3"
# $ ./parse_lat_lon.py lat_lon.txt
# lat = 27.83387, lon = -65.4906
# lat = 29.3, lon = 122.08
# lat = 28.56, lon = -88.70377
# No match: "This line will not be included"
# lat = 39.283, lon = -76.611
# lat = 78.0, lon = 5.0
# No match: "attr.lat_lon : missing"
# lat = 0.0, lon = -170.0
# lat = 11.46'45.7", lon = 93.01'22.3"
# ````
#
# We see a similar problem with "collection_date":
#
# ````
# $ ./xml_ena.py *.xml | grep collection
# attr.collection_date : March 24, 2014
# attr.collection_date : 2013-08-15/2013-08-28
# attr.collection_date : 20100910
# attr.collection_date : 02-May-2012
# attr.collection_date : Jul-2009
# attr.collection_date : missing
# attr.collection_date : 2013-12-23
# attr.collection_date : 5/04/2012
# ````
#
# Imagine how you might go about parsing all these various representations of dates. Be aware that parsing date/time formats is so problematic and ubiquitous that many people have already written modules to assist you!
#
# To run the code below, you will need to install the `dateparser` module:
#
# ````
# $ python3 -m pip install dateparser
# ````
import dateparser
for date in ['March 24, 2014',
'2013-08-15',
'20100910',
'02-May-2012',
'Jul-2009',
'5/04/2012']:
print('{:15}\t{}'.format(date, dateparser.parse(date)))
# You can see it's not perfect, e.g., "20100910" should be "2010-09-10" and "Jul-2009" should not resolve to the 26th of July, but, honestly, what should it be? (Is the 1st any better?!) Still, this saves you writing a lot of code. And, trust me, **THIS IS REAL DATA**! While trying to parse latitude, longitude, collection date, and depth for 35K marine metagenomes from the ENA, I wrote a hundreds of lines of code and dozens of regular expressions!
# ## Exercises
#
# Write the regular expressions to parse the year, month, and day from the following date formats found in SRA metadata. When no day is present, e.g., "2/14," use "01" for the day.
import re
d1 = "2012-03-09T08:59"
print(d1, re.match('', d1))
d2 = "2012-03-09T08:59:03"
d3 = "2017-06-16Z"
d4 = "2015-01"
d5 = "2015-01/2015-02"
d6 = "2015-01-03/2015-02-14"
d7 = "20100910"
d8 = "12/06"
d9 = "2/14"
d10 = "2/14-12/15"
d11 = "2017-06-16Z"
# "Excel" format! What is that?! Look it up.
d12 = "34210"
d13 = "Dec-2015"
d14 = "March-2017"
d15 = "May, 2017"
d16 = "March-April 2017"
d17 = "July of 2011"
d18 = "2008 August"
# Now combine all your code from the previous cell to normalize all the dates into the same format.
# +
dates = ["2012-03-09T08:59", "2012-03-09T08:59:03", "2017-06-16Z",
"2015-01", "2015-01/2015-02", "2015-01-03/2015-02-14",
"20100910", "12/06", "2/14", "2/14-12/15", "2017-06-16Z",
"34210", "Dec-2015", "March-2017", "May, 2017",
"March-April 2017", "July of 2011", "2008 August"]
for date in dates:
year = '1999'
month = '01'
day = '01'
print('{}-{}-{}\t{}'.format(year, month, day, date))
# -
|
lectures/12-regular-expressions/re.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import selenium
from selenium import webdriver
from selenium.webdriver.common.keys import Keys
wd =webdriver.Chrome()
wd.get("http://fb.com")
username = wd.find_element_by_id("email")
password = wd.find_element_by_id("pass")
username.send_keys("your username")
password.send_keys("<PASSWORD>")
sub=wd.find_element_by_id("loginbutton")
sub.submit()
|
fb_login_using_selenium.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.5.4
# language: julia
# name: julia-1.5
# ---
# # Markov model using Soss.jl
# <NAME> (@sdwfrost), 2021-03-08
#
# ## Introduction
#
# The Markov model approach taken here is:
#
# - Stochastic
# - Discrete in time
# - Discrete in state
#
# This tutorial uses the `Soss.jl` package to simulate the Markov model, and is based on one of the demos, the original version of which can be found [here](https://github.com/cscherrer/Soss.jl/blob/master/demos/sir.jl).
#
# ## Libraries
using Random
using Soss
using DataFrames
using StatsPlots
using BenchmarkTools
# ## Utility functions
@inline function rate_to_proportion(r::Float64,t::Float64)
1-exp(-r*t)
end;
# ## Transitions
#
# A `@model` in `Soss.jl` needs to have a variable `state` to store the internal state. The following model describes a single step of the Markov process. In addition to the state variables, `S`, `I`, and `R`, we also keep track of the time, `t`.
sir_markov = @model state,p begin
# Unpack parameters
β = p.β
c = p.c
γ = p.γ
δt = p.δt
# Unpack starting counts
t0 = state.t
S0 = state.S
I0 = state.I
R0 = state.R
N = S0 + I0 + R0
# Transitions between states
S_I ~ Binomial(S0, rate_to_proportion(β*c*I0/N,δt))
I_R ~ Binomial(I0, rate_to_proportion(γ,δt))
# Updated state
t = t0 + δt
S = S0 - S_I
I = I0 + S_I - I_R
R = R0 + I_R
next = (p=p, state=(t=t,S=S,I=I,R=R))
end;
# The following `@model` takes the transitions defined above and plugs them into a Markov chain.
sir_model = @model u0,p begin
x ~ MarkovChain(p, sir_markov(state=u0,p=p))
end;
# ## Time domain
δt = 0.1
nsteps = 400
tmax = nsteps*δt;
# ## Initial conditions
u0 = (t=0.0, S=990, I=10, R=0); # t,S,I,R
# ## Parameter values
p = (β=0.05, c=10.0, γ=0.25, δt=δt);
# ## Random number seed
Random.seed!(1234);
# ## Running the model
#
# To run the model, we first instantiate a random number generator for our model with our initial conditions and parameters, and then make repeated calls until we exceed the number of iterations. An array is used to store the state variables.
r = rand(sir_model(u0=u0,p=p));
data = [u0]
for (n,s) in enumerate(r.x)
n>nsteps && break
push!(data,s)
end;
# ## Post-processing
#
# We can convert the output to a dataframe for convenience.
df_markov = DataFrame(data);
# ## Plotting
#
# We can now plot the results.
@df df_markov plot(:t,
[:S :I :R],
label=["S" "I" "R"],
xlabel="Time",
ylabel="Number")
# ## Benchmarking
@benchmark begin
r = rand(sir_model(u0=u0,p=p));
data = [u0]
for (n,s) in enumerate(r.x)
n>nsteps && break
push!(data,s)
end
end
|
notebook/markov_soss/markov_soss.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Image recognition android application project
# - Flower image classification with Baseline CNN (20200823)
# - 4685 training set with 5 class, 100 iterations, 16 batch
#
# ### Reference
# - [Advanced Computer Vision with TensorFlow], https://stephan-osterburg.gitbook.io/coding/coding/ml-dl/tensorfow
# ## 1. Import packages
# +
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import os
import sys
import tarfile
import glob
from six.moves import urllib
from glob import glob
import random
import shutil
import keras
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten
from keras.layers.convolutional import Conv2D, MaxPooling2D
from keras.models import load_model
import tensorflow.compat.v1 as tf
tf.disable_v2_behavior()
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
# -
# ## 2. Load and Explore the Flower Dataset
def load_data_files(base_dir):
folder_name = "dataset/flower_photos"
RAW_DATASET = os.path.join(base_dir, folder_name)
abs_dir = os.path.join(os.getcwd(), folder_name)
sub_dir = os.listdir(abs_dir)
data_dic = {}
for class_name in sub_dir:
imgs = glob(os.path.join(RAW_DATASET,class_name,"*.jpg"))
data_dic[class_name] = imgs
print("Class: {}".format(class_name))
print("Number of images: {} \n".format(len(imgs)))
return data_dic
BASE_DIR = os.getcwd()
data_dic = load_data_files(BASE_DIR)
# ## 3. Split train and validation dataset
# Create new directory and copy files to it
def copy_files_to_directory(files, directory):
if not os.path.exists(directory):
os.makedirs(directory)
print("Created directory: {}".format(directory))
for f in files:
shutil.copy(f, directory)
print("Copied {} files.\n".format(len(files)))
def train_validation_split(base_dir, data_dic, split_ratio=0.2):
FLOWER_DATASET = os.path.join(base_dir,"flower_dataset")
if not os.path.exists(FLOWER_DATASET):
os.makedirs(FLOWER_DATASET)
for class_name, imgs in data_dic.items():
idx_split = int(len(imgs) * split_ratio)
random.shuffle(imgs)
validation = imgs[:idx_split]
train = imgs[idx_split:]
copy_files_to_directory(train, os.path.join(FLOWER_DATASET,"train",class_name))
copy_files_to_directory(validation, os.path.join(FLOWER_DATASET,"validation",class_name))
# +
# BASE_DIR = os.getcwd()
# train_validation_split(BASE_DIR, data_dic, split_ratio=0.2)
# -
# ## 4. Image preprocessing
batch_size = 16
num_classes = 5
epochs = 100
# +
preprocessing_image = tf.keras.preprocessing.image
train_datagen = preprocessing_image.ImageDataGenerator(
rescale=1./255,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True)
test_datagen = preprocessing_image.ImageDataGenerator(rescale=1./255)
# +
BASE_DIR = os.getcwd()
train_generator = train_datagen.flow_from_directory(
os.path.join(BASE_DIR, "flower_dataset/train"),
target_size=(32, 32),
batch_size=batch_size,
class_mode='categorical')
validation_generator = test_datagen.flow_from_directory(
os.path.join(BASE_DIR, "flower_dataset/validation"),
target_size=(32, 32),
batch_size=batch_size,
class_mode='categorical')
# -
# ## 5. Baseline CNN model
model = Sequential()
model.add(Conv2D(32, kernel_size=(5, 5), strides=(1, 1), padding='same',
activation='relu',
input_shape=(32, 32, 3)))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(64, (5, 5), activation='relu', padding='same'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(1024, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(num_classes, activation='softmax'))
model.summary()
model.compile(loss='categorical_crossentropy', optimizer='sgd', metrics=['accuracy'])
# model.compile(loss='categorical_crossentropy', optimizer=adam(lr=0.0005), metrics=['accuracy'])
# ## 6. Training
history = model.fit_generator(
train_generator,
steps_per_epoch = 4685//batch_size,
epochs=epochs,
validation_data = validation_generator,
validation_steps=20)
# ## 7. Accuracy Analysis and Visualization
def plot_accuracy_and_loss(history):
plt.figure(1, figsize= (15, 10))
# plot train and test accuracy
plt.subplot(221)
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title('Baseline CNN Accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
# plot train and test loss
plt.subplot(222)
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('Baseline CNN loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper right')
plt.show()
plot_accuracy_and_loss(history)
# +
print("-- Evaluate --")
scores_train = model.evaluate_generator(
train_generator,
steps = 5)
scores_val = model.evaluate_generator(
validation_generator,
steps = 5)
print("%s: %.2f%%" %(model.metrics_names[1], scores_train[1]*100))
print("%s: %.2f%%" %(model.metrics_names[1], scores_val[1]*100))
print("-- Predict --")
output_train = model.predict_generator(train_generator, steps=5)
output_val = model.predict_generator(validation_generator, steps=5)
np.set_printoptions(formatter={'float': lambda x: "{0:0.3f}".format(x)})
print(train_generator.class_indices)
print(output_train)
print(validation_generator.class_indices)
print(output_val)
# -
# ## 8. Save and Load model
# +
# save model architecture
model_json = model.to_json()
open('3_model.json', 'w').write(model_json)
# save model's learned weights
model.save_weights('3_weights.h5', overwrite=True)
# +
# Load trained model
from keras.models import model_from_json
json_file = open("3_model.json", "r")
loaded_model_json = json_file.read()
json_file.close()
loaded_model = model_from_json(loaded_model_json)
# model weight load
loaded_model.load_weights("3_weights.h5")
print("Loaded model from disk")
|
Image_recognition_DeepLearning_Models/20200823_Model_Selection_with_more_data/3_Baseline_CNN_with_flower_dataset.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
from dotenv import load_dotenv, find_dotenv
from os.path import join, dirname, basename, exists, isdir
### Load environmental variables from the project root directory ###
# find .env automagically by walking up directories until it's found
dotenv_path = find_dotenv()
# load up the entries as environment variables
load_dotenv(dotenv_path)
# now you can get the variables using their names
# Check whether a network drive has been specified
DATABASE = os.environ.get("NETWORK_URL")
if DATABASE == 'None':
pass
else:
pass
#mount network drive here
# set up directory pathsa
CURRENT_DIR = os.getcwd()
PROJ = dirname(dotenv_path) # project root directory
DATA = join(PROJ, 'data') #data directory
RAW_EXTERNAL = join(DATA, 'raw_external') # external data raw directory
RAW_INTERNAL = join(DATA, 'raw_internal') # internal data raw directory
INTERMEDIATE = join(DATA, 'intermediate') # intermediate data directory
FINAL = join(DATA, 'final') # final data directory
RESULTS = join(PROJ, 'results') # output directory
FIGURES = join(RESULTS, 'figures') # figure output directory
PICTURES = join(RESULTS, 'pictures') # picture output directory
# make folders specific for certain data
folder_name = ''
if folder_name != '':
#make folders if they don't exist
if not exists(join(RAW_EXTERNAL, folder_name)):
os.makedirs(join(RAW_EXTERNAL, folder_name))
if not exists(join(INTERMEDIATE, folder_name)):
os.makedirs(join(INTERMEDIATE, folder_name))
if not exists(join(FINAL, folder_name)):
os.makedirs(join(FINAL, folder_name))
print('Standard variables loaded, you are good to go!')
# +
import pandas as pd
import re
# import data
data = pd.read_csv(f"{INTERMEDIATE}/proteomics/acetate_usages.csv", index_col=0)
data.head()
# get cell volumes
cell_volumes = pd.read_csv(f"{RAW_INTERNAL}/proteomics/growth_conditions.csv", index_col=0)
cell_volumes = cell_volumes["Single cell volume [fl]1"]
# remove the first two rows of LB
cell_volumes = cell_volumes.loc[~cell_volumes.index.duplicated(keep='first')]
# rename the number 3 in there
cell_volumes = cell_volumes.rename({'Osmotic-stress glucose3':'Osmotic-stress glucose_uncertainty'}, axis='index')
rename_dict = {i:re.sub(r'\W+', '', i).lower() for i in cell_volumes.index}
cell_volumes = cell_volumes.rename(rename_dict, axis='index')
# -
data.head()
# +
# Finally, convert to mmol/gDW:
water_content = 0.3
cell_density = 1.105e-12
# Iterate through the dataset and multiply by the corresponding cell volume, to get mmol/fL:
for (col_name, d) in data.iteritems():
chemo_name = "acetate"
try:
data[col_name] = data[col_name] * cell_volumes.loc[chemo_name]
except:
print(chemo_name)
data = data * cell_density * water_content
# convert into counts
data = data * 6.022e+23 / 1000
# -
data_for_UP2genes = pd.read_csv(f"{RAW_EXTERNAL}/raw_proteomics_all.csv")
data_for_UP2genes = data_for_UP2genes.drop([0])
UP2gene = dict(zip(data_for_UP2genes["UP"], data_for_UP2genes["Frist_Gene"]))
# translate index uniprot names to gene names and remove enzymes not included in the data
data = data.loc[list(set(data.index).intersection(UP2gene.keys()))]
data.index = [UP2gene[i] for i in list(set(data.index).intersection(UP2gene.keys()))]
#data["pyr_1_original"] =
# +
data_for_UP2genes.index = data_for_UP2genes['Frist_Gene']
data_for_UP2genes = data_for_UP2genes.loc[~data_for_UP2genes.index.duplicated(keep='first')]
data = data.loc[~data.index.duplicated(keep='first')]
data['ac_original'] = data_for_UP2genes.loc[data.index, ['acetate']]
data['ac_original_1'] = data_for_UP2genes.loc[data.index, ['acetate.1']]
data['ac_original_2'] = data_for_UP2genes.loc[data.index, ['acetate.2']]
data
# -
data.to_csv(f"{INTERMEDIATE}/proteomics/acetate_usages_counts.csv")
|
data_science/code/modeling/chemostat_0.12/conc2count_acetate.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="https://image.aladin.co.kr/product/16902/16/cover500/k202534274_1.jpg" width="200" height="200"><br>
# # Chapter 9 문자열 처리하기
#
# 판다스에서 다루는 데이터의 대부분은 문자열로 저장되어 있다. 어떤 경우에는 정수, 실수와 같은 숫자 데이터도 문자열로 저장된다.이번에는 판다스가 아니라 파이썬으로 문자열을 처리하는 방법에 대해 알아보자. 목차는 다음과 같다.
#
# - 9-1: 문자열 다루기
# - 9-2: 문자열 메서드
# - 9-3: 문자열 포메팅
# - 9-4: 정규식으로 문자열 처리에 날개 달기
#
# ## 9-2 문자열 다루기
#
# ### 파이썬과 문자열
# 문자열은 작은따옴표나 큰따옴표로 감싸서 만든다. 다음은 작은따옴표로 grail, a acratch라는 문자열 데이터를 만들어 변수 word, sent에 저장한 것이다.
word = 'grail'
sent = 'a scratch'
# ### 인덱스로 문자열 추출하기
# 데이터프레임에서 인덱스를 이용하여 원하는 데이터를 추출했던 것을 기억하자. 문자열도 인덱스를 사용하여 추출할 수 있다. 문자는 길이가 1인 문자열로 생각하면 된다.
#
# ### 문자열 추출하기
#
# #### 1.
#
# 각 문자열의 첫 번째 문자는 오른쪽과 같은 방법으로 추출 한다.
# +
word = '<PASSWORD>'
sent = 'a scratch'
print(word[0])
print(sent[0])
# -
# #### 2.
# 인덱스 슬라이싱을 사용하면 여러 개의 문자를 한 번에 추출할 수 있습니다. 오른쪽은 인덱스 슬라이싱을 사용하여 0~2번째 인덱스의 문자를 추출한 것인다. 이때 대괄호에 지정한 왼쪽 범위는 포함하고 오른쪽 범위는 포함하지 않는다. 즉, 인덱스가 0,1,2인 문자를 추출한다.
print(word[0:3])
# #### 3.
# 음수를 사용해도 문자열을 추출할 수 있다. 만약 인덱스를 -1로 지정하여 문자열을 추출하면 마지막 문자가 추출이 된다. 다음은 다양한 방법으로 음수 범위의 인덱스 슬라이싱을 한 것이다.
# +
print(sent[-1])
print(sent[-9:-8])
print(sent[0:-8])
# -
# ### 전체 문자열 추출하기
# 왼쪽 범위가 비어 있으면 문자열의 첫 번째 위치부터 문자열을 추출한다. 반대로 오른쪽 범위가 비어 있으면 문자열의 마지막 위치까지 문자열을 추출한다. 즉, word[0:3]과 word[:3]은 같은 범위의 문자열을 추출한다. 다음을 보자.
#
#
# ### 왼쪽이나 오른쪽 범위를 지정하지 않고 문자열 추출하기
#
# #### 1.
# 왼쪽 범위를 비우고 문자열을 추출해보자.
print(word[0:3])
print(word[:3])
# #### 2.
# 만약 오른쪽 범위를 비우면 문자열의 마지막 위치까지 문자열을 추출하게 된다.
# +
print(sent[2:len(sent)])
print(sent[2:])
# -
# #### 3.
# 양쪽을 모두 비우면 전체 문자열을 추출한다.
print(sent[ : ])
# #### 4.
# 만약 문자열을 일정한 간격으로 건너뛰며 추출해야 한다면 콜론을 하나 더 추가 하여 추출 간격을 지정하면 된다. 오른쪽은 전체 문자열을 추출하되 다음 추출 문자는 자신을 포함하여 거리가 2인 인덱스의 문자를 추출한 것이다.
print(sent[::2])
# ## 9-2 문자열 메서드
#
# 지금까지 인덱스 슬라이싱으로 문자열을 추출하는 방법을 알아보았다. 그런데 문자열이 너무 길어서 내가 원하는 문자가 몇 번째 인덱스에 잇는지 파악하기 어렵거나 문자열에 포함된 소문자를 모두 대문자로 바꾸고 싶다면 어떻게 해야 할까? 이런 경우에는 문자열 메서드를 사용하면 된다. 자주 사용하는 문자열 메서드와 간단한 실습 코드는 다음 표를 확인해보자.
#
# #### 문자열 메서드
#
# |메서드|설명|
# |:-:|:-:|
# |capitalize|첫 문자를 대문자로 변환|
# |count|문자열의 개수를 반환|
# |startswith|문자열이 특정 문자로 시작하면 참|
# |endswith|문자열이 특정 문자로 끝나면 참|
# |find|찾을 문자열의 첫 번째 인덱스를 반환. 실패시 -1 반환|
# |index|find 메서드와 같은 역할을 수행하지만 실패시 ValueError를 반환|
# |isalpha|모든 문자가 알파벳이면 참|
# |isdecimal|모든 문자가 숫자면 참|
# |isalnum|모든 문자가 알파벳이거나 숫자면 참|
# |lower|모든 문자를 소문자로 변환|
# |upper|모든 문자를 대문자로 변환|
# |replace|문자열의 문자를 다른 문자로 교체 |
# |strip|문자열의 맨 앞과 맨 뒤에 잇는 빈 칸을 제거|
# |split|구분자를 지정하여 문자열을 나누고, 나눈 값들의 리스트를 반환|
# |partition|split 메서드와 비슷한 역할을 수행하지만 구분자도 반환|
# |center|지정한 너비로 문자열을 늘이고 문자열을 가운데 정렬|
# |zfill|문자열의 빈 칸을 '0'으로 채운다.|
# #### 파이썬 문자열 메서드 - 실습 코드
print("black Knight".capitalize())
"It's just a flesh wound!".count('u')
"Halt! Who goes there!".startswith('Halt')
"coconut".endswith("nut")
"It's just a flesh wound!".find('u')
"It's just a flesh wound!".index('scratch')
"old woman".isalpha()
"37".isdecimal()
"I'm 37".isalnum()
"Black Knight".lower()
"Black Knight".upper()
"flesh wound!".replace('flesh wound', 'scratch')
"I'm not dead.".strip()
"NI! NI! NI! ni!".split(sep = " ")
"3,4".partition(",")
"nine".center(10)
"9".zfill(5)
# ### join, splitlines, replace 메서드 실습하기
#
# #### 1. join 메서드
#
# join 메서드는 문자열을 연결하여 새로운 문자열을 반환하는 메서드이다. 다음은 분리된 좌표 데이터를 합친 것이다. 이때 join 메서드 앞에 문자를 지정하면 해당 문자를 단어 사이에 넣어 연결해 주면 된다.
# +
d1 = '40'
m1 = "46'"
s1 = '52.837"'
u1 = 'N'
d2 = '73'
m2 = "58'"
s2 = '26.302"'
u2 = 'W'
coords = ' '.join([d1, m1, s1, u1, d2, m2, s2, u2])
coords
# -
# #### 2. splitlines 메서드
#
# 이번에는 splitlines 메서드에 대해 알아보자. splitlines 메서드는 여러 행을 가진 문자열을 분리한 다음 리스트로 반환 한다. multi_str에 저장된 문자열을 splitlines 메서드로 분리하면 다음과 같이 변환된다.
# +
multi_str = """Guard: What? Ridden on a horse?
King arthur: Yes!
Guard: You're using coconuts!
King Arthur: What?
Guard: You've got ... coconut[s] and you're bangin' ''em together/
"""
print(multi_str)
multi_str_split = multi_str.splitlines()
print(multi_str_split)
# -
# #### 3.
# 인덱스 슬라이싱을 응용하면 특정 문자열만 가져올 수도 있다. 다음은 Guard의 대사만 가져온 것
guard = multi_str_split[::2]
print(guard)
# #### 4. replace 메서드
#
# 만약 Guard의 대사에서 'Guard:'라는 문자열을 빼고 싶다면 replace 메서드를 사용하자.
guard = multi_str.replace("Guard: ", "").splitlines()[::2]
print(guard)
# ## 9-3 문자열 포메팅
#
# ### 문자열 포메팅 하기
#
# 문자열 포메팅은 문자열을 편리하게 출력할 수 있게 해주는 기능이다. 예를 들어 I can swim, I can fly, I can run과 같은 문자열은 I can이라는 문자열에 swim, run, fly와 같은 단어만 바꿔 넣어 출력 하는 것이 더 편리 하다. 즉, 문자열 포메팅이란 출력할 문자열의 형식을 지정하거나 변수를 조합하여 출력하는 방법이다.
#
# #### 1.
# 다음과 같이 단어를 삽입할 위치를 {}로 지정하고 format 메서드에 원하는 단어를 전달하면 {}의 위치에 전달한 단어를 삽입해 출력한다. 이때 {}를 플레이스 홀더라고 부른다.
# +
var = 'flesh wound'
s = "It's just a {}!"
print(s.format(var))
print(s.format('scratch'))
# -
# #### 2.
# 플레이스 홀더는 여러 번 사용해도 된다. 하지만 여러 단어를 전달할 때는 어떻게 해야 할까? format 메서드에 여러 단어를 전달하려면 인덱스 개념을 응용하면 된다. 다음은 1개의 단어만 사용한다. 따라서 인덱스를 0으로 지정한 플레이스 홀더를 사용한다.
# +
s = """Black Kinight: 'Tis but a {0}.
King Arthur: A {0}? Your arm's off!
"""
print(s.format('scratch'))
# -
# #### 3.
#
# 플레이스 홀더에는 변수를 지정해도 된다. 단, format 메서드에 전달하는 문자열도 변수에 담아 전달해야 한다.
s = 'Hayden Planentarium Coordinates: {lat}, {lon}'
print(s.format(lat = '40.7815 N', lon = '73.9733 W'))
# ### 숫자 데이터 포메팅하기
#
# 이번에는 숫자 데이터 포매팅에 대해 알아보자. 숫자 데이터 포매팅ㄹ을 사용하면 실수의 소수점을 어디까지 출력할지 등을 설정할 수 있다.
#
# #### 1.
# 숫자 데이터도 플레이스 홀더를 사용할 수 있다.
print('Some digits of pi: {}'.format(3.14159265359))
# #### 2.
# 플레이스 홀더에:, 를 넣으면 쉼표를 넣어 숫자를 표현할 수도 있다.
print("In 2005, Lu Chao of china recited {:,} digits of pi".format(67890))
# #### 3.
# 소수는 좀 더 다양한 방법으로 포매팅 가능하다. 다음은 플레이스 홀더에 0:.4, 0:.4%를 넣어 7/67890의 결과값을 포매팅 한 것이다. {0:.4}와 {0:.4% }에서 0은 format 메서드에서 전달받을 값(7/67890)의 인덱스를 의미하고 .4는 소수점 이하의 숫자를 4개까지 술력하겠다는 것을 의미한다. 이때 %를 사용하면 7/67890의 결과값을 백분율로 환산하여 출력한다.
print("I remember {0:.4} or {0:.4%} of what LuChao recited".format(7/67890))
# #### 4.
# 만약 사용자의 아이디가 5자리로 숫자로 표현되어야 한다면 42와 같은 2자리의 값은 00042로 출력해야 한다. 다음은 format의 0번째 값을 5자리의 수로 표현하되 빈 칸을 0으로 채워 출력한 것이다.
# ### % 연산자로 포매팅하기
#
# 지금까지는 플레이스 홀더를 이용하여 여러 가지 포매팅을 해보았다. 이번에는 % 연산자로 포매팅을 실습해 보자,
#
# #### 1.
# 만약 삽입할 값이 10진수라면 값을 삽입할 위치에 %d라고 입력해야 한다. 그런 다음 % 연산자를 이용하여 삽입할 값(7)을 지정하여 출력한다.
s = 'I only know %d digits of pi' % 7
print(s)
# #### 2.
# 만약 삽입할 값이 문자열이라면 값을 삽입할 위치에 %s라고 입력해야 한다. 이때 %와 s사이에 소괄호를 사용하여 변수를 지정한다.
print('Some digits of %(cont)s: %(value).2f' % {'cont': 'e', 'value': 2.718})
# ### f-strings 포매팅 사용
# f-strings 포매팅은 파이썬 3.6버전에서 새로 도입한 포매팅이다. f-strings 포매팅의 사용방법은 format 메서드의 사용 방법과 동일하다. 하지만 문자열 앞에 f를 붙인다는 점이다. f-strings포매팅은 읽기 쉬울 뿐만 아니라 문자열을 빠르게 처리한다는 장점이 있다. 다음을 보자.
# +
var = 'flesh wound'
s = f"It's just a {var}!"
print(s)
lat = '40.7815 N'
lon = '73.9733 W'
s = f'Hayen Planetarium Coordinates: {lat}, {lon}'
print(s)
# -
# ## 9-4 정규식으로 문자열 처리에 날개 달기
#
# ### 정규식이란?
#
# 수만 개의 문자열 중에서 내가 원하는 패턴의 문자열만 추출하는 방법을 알아보자. find 메서드를 사용해도 좋지만 정규표현식을 사용하면 더 편리하다.
#
# ### 정규식 표현 - 문법, 특수문자
#
# 정규식 표현에서 사용되는 문법과 특수문자를 다음과 같이 표로 정리 해두 었다. 정규식을 사용하려면 다음 표를 참고하여 찾고자 하는 문자열의 정규식 패턴들 만들어야 한다.
#
# ### 정규식 표현 - 메서드
# 다음은 정규식 메서드이다 정규식 메서드는 정규식 패턴을 사용하지 않아도 메서드를 호출하는 방법으로 원하는 패턴의 문자열을 찾을 수 있도록 해준다.
#
# - search: 첫 번째로 찾은 패턴의 양 끝 인덱스를 반환
# - match: 문자열의 처음붕터 검색하여 찾아낸 패턴의 양 끝 인덱스를 반환
# - fullmatch: 전체 문자열이 일치하는지 검사
# - split: 지정한 패턴으로 잘라낸 문자여을 리스트로 반환
# - findall: 지정한 패턴을 찾아 리스트로 반환
# - finditer: findall 메서드와 기능은 동일하지만 iterator를 반환
# - sub: 첫 번째로 전달한 패턴을 두 번째 인자로 전달한 값으로 교체
#
# ### 정규식으로 전화번호 패턴 찾기
#
# 전화번호와 같은 단순한 데이터도 복잡하고 다양한 정규식이 필요하다. 다음을 보자.
#
# #### 1.
# 다음과 같이 re 모듈과 테스트용 문자열을 준비 한다.
# +
import re
tele_num = '1234567890'
# -
# #### 2.
# match 메서드를 사용하여 길이가 10인 숫자를 확인해 보자. pattern 인자에는 10개의 숫자를 의미하는 10개의 \d를, string에는 테스트용 문자열인 tele_num을 전달했다. 만약 패턴을 찾으면 Match 오브젝트를 반환한다. Match 오브젝트를 출력하면 span에는 찾은 패턴의 인덱스가, match에는 찾은 패턴의 문자열이 있는 것을 확인할 수 있다.
# +
m = re.match(pattern='\d\d\d\d\d\d\d\d\d\d', string = tele_num)
print(type(m))
print(m)
# -
# #### 3.
# 이때 bool 메서드에 m을 전달하면 True나 False를 얻을 수 있다. 즉, match 메서드가 반환한 Match 오브젝트는 bool 메서드로, True, False를 판단할 수 있다.
# +
print(bool(m))
if m:
print('match')
else:
print('no match')
# -
# #### 4.
# Match 오브젝트에는 찾아낸 패턴의 정보를 확인할 수 있는 다양한 메서드가 있다. start와 end 메서드는 첫 번째와 마지막 인덱스를 반환한다. span 메서드는 찾은 패턴의 첫 번째와 마지막 인덱스를 한 번에 반환한다. group 메서드는 찾아낸 패턴을 반환 한다.
# +
print(m.start())
print(m.end())
print(m.span())
print(m.group())
# -
# #### 5.
# 그런데 전화번호를 입력하는 방법은 1234567890이 아니라 123-456-7890같은 형식을 사용하게 된다. 다음은 이와 같은 패튼을 검사 해보자.
# +
tele_num_spaces = '123 456 7890'
m = re.match(pattern = '\d{10}', string = tele_num_spaces)
print(m)
if m:
print('match')
else:
print('no match')
# -
# #### 6.
# 위의 문제를 해결하려면 정규식을 다시 작성해야 한다. 다음과 같이 빈 칸을 의미하는 정규식 \s?를 넣어 패턴을 다시 만들자.
p = '\d{3}\s?\d{3}\s?\d{4}'
m = re.match(pattern=p, string=tele_num_spaces)
print(m)
# #### 7.
# 지역 코드는 소괄호로 감싸고 나머지 번호는 반각 기호로 구분한 전화번호의 정규식은 다음과 같이 작성한다.
# +
tele_num_space_paren_dash = '(123) 456-7890'
p = '\(?\d{3}\)?\s?\d{3}\s?-?\d{4}'
m = re.match(pattern = p, string = tele_num_space_paren_dash)
print(m)
# -
# #### 8.
# 국가 코드까지 있는 전화번호의 정규식은 다음과 같이 작성 한다.
# +
cnty_tele_num_space_paren_dash = '+1 (123) 456-7890'
p = '\+1\s?\(?\d{3}\)?\s?\d{3}\s?-?\d{4}'
m = re.match(pattern=p, string=cnty_tele_num_space_paren_dash)
print(m)
# -
# ### compile 메서드로 정규식 메서드 사용
#
# 패턴을 반복해서 사용하려면 compile 메서드로 패턴을 컴파일한 다음 변수에 저장하여 사용하면 된다. 다음은 앞에서 실습한 내용 중 하나를 compile 메서드로 처리한 것이다. 패턴을 컴파일한 다음 변수에 저장했기 때문에 정규식 메서드를 반복해서 사용할 수 있어 매우 편리하다.
p = re.compile('\d{10}')
s = '1234567890'
m = p.match(s)
print(m)
|
Chapter_9.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sh
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Bash
# language: bash
# name: bash
# ---
ls -lah ../data
ls -lah ../ref
# +
kallisto="$HOME/bin/kallisto_linux-v0.43.1/kallisto"
#$kallisto index -i "../ref/cele_tx.idx" "../ref/Caenorhabditis_elegans.WBcel235.cdna.all.fa.gz"
# -
ls -lah ../data/ig01/trim
# +
# Run kallisto quant for samples
base_dir="../data"
for sample in $(ls $base_dir | grep ^ig)
do
echo "Running kallisto quant for sample "$sample
output="../results/kallisto/$sample"
if [ ! -d "$output" ]; then
mkdir -p $output
fi
read1="$base_dir/$sample/trim/R1_pe.fastq.gz"
read2="$base_dir/$sample/trim/R2_pe.fastq.gz"
$kallisto quant \
-i "../ref/cele_tx.idx" \
-t 30 \
-o $output -b 100 <(zcat $read1) <(zcat $read2)
done
# -
ls -lah ../results/kallisto/ig05
head ../results/kallisto/ig05/abundance.tsv
head ../results/kallisto/ig05/abundance.tsv
|
nlabseq/templates/kallisto_quantification.ipynb
|
# ---
# jupyter:
# jupytext:
# formats: ipynb,py:percent
# text_representation:
# extension: .py
# format_name: percent
# format_version: '1.3'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %% [markdown]
#
# <p align="center">
# <img width="100%" src="../../../multimedia/mindstorms_51515_logo.png">
# </p>
#
# # `aat_ms5`
# Python program to control the AAT MS5 robot on patrol mode.
# The tank will move in circles, while the droid in the blaster will be looking for any sympathisants of the Republic.
# When the distance sensor detects a target, the tank will stop and the droid will center the
# blasters to fire!
#
# You can find a video of the robot functioning [here](https://www.youtube.com/watch?v=Ma7CmThktUg&feature=youtu.be&ab_channel=ArturoMoncada-Torres).
#
# # Required robot
# * AAT MS5 (you can find the [instructions for building it here](https://arturomoncadatorres.com/aat-ms5/))
#
# <img src="../multimedia/aat_ms5.png" width="50%" align="center">
#
# # Source code
# You can find the code in the accompanying [`.py` file](https://github.com/arturomoncadatorres/lego-mindstorms/blob/main/mocs/aat_ms5/programs/aat_ms5.py). To get it running, simply copy and paste it in a new Mindstorms project.
#
# # Imports
# %%
from mindstorms import MSHub, Motor, MotorPair, ColorSensor, DistanceSensor, App
from mindstorms.control import wait_for_seconds, wait_until, Timer
from mindstorms.operator import greater_than, greater_than_or_equal_to, less_than, less_than_or_equal_to, equal_to, not_equal_to
import math
import hub
# %%
print("-"*15 + " Execution started " + "-"*15 + "\n")
# %% [markdown]
# # Initialize hub
# Notice we won't be using the standard `MSHub`, but rather the "raw" `hub`.
# It is a little lower level, but it allows us making more things.
# Fore more information, see [Maarten Pennings brilliant explanation and unofficial documentation about it](https://github.com/maarten-pennings/Lego-Mindstorms/blob/main/ms4/faq.md#why-are-there-so-many-ways-to-do--in-python).
# %%
# hub = MSHub()
# %%
# hub.status_light.on('black')
hub.led(0, 0, 0)
# %% [markdown]
# # Initialize motors
# %%
print("Configuring motors...")
motor_steer = Motor('A') # Front wheels (for steering)
motor_power = Motor('C') # Back wheels (for moving)
motor_turret = Motor('B') # Turrent spinning
# %%
print("Setting motors to position 0...")
motor_steer.run_to_position(45, speed=100)
motor_steer.run_to_position(0, speed=100)
motor_turret.run_to_position(0, speed=75)
print("DONE!")
# %% [markdown]
# # Initialize distance sensor
# %%
print("Initializing distance sensor...")
distance_sensor = DistanceSensor('D')
print("DONE!")
# %% [markdown]
# # Put the AAT MS5 in motion
#
# The tank will move until the distance sensor detects an obstacle.
#
# The steering is given by `POSITION`.
# * A value between `0` and `90` will steer the tank to the left.
# - A value closer to `0` will make the tank turn wider.
# - A value closer to `90` will make the tank turn tighter.
#
# * A value between `270` and `360` will steer the tank to the right.
# - A value closer to `270` will make the tank turn tighter.
# - A value closer to `360` will make the tank turn wider.
# %%
POSITION = 270
print("Steering...")
motor_steer.run_to_position(POSITION, speed=35)
print("DONE!")
# %% [markdown]
# The tank speed is given by `SPEED`. It should have a value between `-100` and `100`.
# * A negative value will move the tank forward.
# * A positive value will move the tank backwards.
#
# Recommended value is `-50`
# %%
SPEED = -50
print("Moving...")
motor_power.start(SPEED)
print("DONE!")
# %% [markdown]
# # Configure the patrolling
# We will move the turret constantly. It will go from left to right and from
# right to left. When an obstacle is detected, the turret will go back to the
# initial position and "fire".
#
#
#
# ## Define distance function
# As part of the program, we need to continuously check if the
# measured distance is less than 10 cm.
# %%
OBSTACLE_DISTANCE = 10 # [cm]
# %% [markdown]
# However, if the sensor reads no measure, it will return a `None`, which
# in turn will generate an error (since we cannot do a comparision
# between a `None` and something else).
#
# To solve this, we will define our own cuestom distance function.
# This way, when the sensor has no reading, we will just return
# a (simulated) very long distance (instead of returning a `None`).
# This will allow us to safely do the comparision.
# %%
def my_get_distance_cm():
"""
Parameters
----------
None
Returns
-------
dist:
Distance value (in cm).
If the sensor returns a None, it returns a very large value (1000).
"""
distance = distance_sensor.get_distance_cm()
if distance == None:
distance = 10000
return distance
# %% [markdown]
# ## Patrolling
# Now, in order to be able to stop the turret at any moment
# (and not until the motor has completed a whole sweep),
# we will use co-routines.
#
# > This is a simplified version [David Lechner's trick](https://community.legoeducation.com/discuss/viewtopic/66/110), which I've used before in [Charlie's `drum_solo`](https://nbviewer.jupyter.org/github/arturomoncadatorres/lego-mindstorms/blob/main/base/charlie/programs/drum_solo.ipynb?flush_cache=True).
# In this case, we are only controlling one motor (the turret) and we don't depend on time
# (but rather on the motor position). Thus, we don't need a timer.
#
# We need to define a function for moving the turret.
# Pay attention to the comments, since they explain how using
# co-routines work. It isn't very hard, but it isn't trivial either.
# %%
def move_turret():
"""
Moves the AAT MS5 turrent.
Parameters
----------
None
Returns
-------
None
"""
# First, we need to define the coroutine.
# In this case, we only need one (corresponding to the turret motor).
# Notice how the definition is very similar to that of a function.
# Coroutines also have input parameters.
# However, they have no "output" (i.e., return), but actually a yield.
def background_turret(angle):
"""
Parameters
----------
angle:
The angle at which the turret turns.
In practice, this value is twice the original angle, since
it moves completely from one side to the other (and not from
the center to one side). That is why it is passed to this
function multiplied by two.
In degrees.
"""
# We want to make sure counted degrees are initialized at 0.
motor_turret.set_degrees_counted(0)
# Notice that we use the absolute value of the counted degrees.
# This is to ensure that it works on the way back (when the measured
# degrees would be negative).
curr_turret_position = math.fabs(motor_turret.get_degrees_counted())
# Here, we check if the motor has reached the desired angle.
while curr_turret_position < angle:
# If you wish to see the current turret position and the target angle,
# uncomment the following line.
# print(str(curr_turret_position) + " - " + str(angle))
# We update the turret current position.
curr_turret_position = math.fabs(motor_turret.get_degrees_counted())
# If the turret hasn't reached the desired angle, we reach this yield.
# yield lets the rest of the program run until we come back
# here again later to check if the condition was met.
yield
def turret_patrol():
while True:
# This is how we receive a parameter.
# In this case, it corresponds to the angle the motor should move.
angle_action = yield
# We make sure we only execute code if the received
# value was transmitted correctly.
if not angle_action == None:
# We will start to move the turret...
motor_turret.start(10)
# ...and check if its angle exceeded the maximum allowed.
# First we move the turret from left to right...
yield from background_turret(angle_action*2)
hub.sound.beep(150, 200, hub.sound.SOUND_SIN) # Play simple tone
# ...and from right to left (exactly same process, but inverted speed).
motor_turret.start(-10)
yield from background_turret(angle_action*2)
# hub.sound.play("/extra_files/Ping")
hub.sound.beep(150, 200, hub.sound.SOUND_SIN) # Play simple tone
# We assume that the movement is immediate and takes no time.
# This isn't completely true, but for now it works.
# Since turret_patrol() is a coroutine and uses yield
# (i.e., it isn't a function and thus has no return), it will NOT
# run here when we call it. Instead, it will just be created as a generator object.
# This generator will be used to run the functions one yield (i.e., step) at a time.
turret_generator = turret_patrol()
# Now we will actually start the task.
# The task (turret patrolling) will be run as long as the distance sensor
# doesn't detect an obstacle.
while my_get_distance_cm() > OBSTACLE_DISTANCE:
next(turret_generator)
turret_generator.send(TURRET_ANGLE)
wait_for_seconds(0.01) # Small pause between steps.
return None
# %% [markdown]
# After we have defined the turret movement, we can now make the AAT MS5 patrol until it finds those pesky Republic supporters!
# %%
TURRET_ANGLE = 40
print("Initializing turret with angle " + str(TURRET_ANGLE) + "...")
motor_turret.set_default_speed(10)
motor_turret.run_for_degrees(-TURRET_ANGLE)
print("DONE!")
print("Starting patrolling...")
move_turret()
print("DONE!")
# %% [markdown]
# Once it finds an enemy (i.e, it detects an obstacle), it will stop and center the turret.
# %%
print("Enemy detected! Attack!")
motor_power.stop() # Stop the movement
motor_turret.run_to_position(0, speed=75) # Center the turret
# %% [markdown]
# Then, it will fire three blasters. Each blaster will come with a sound and an
# animation of the blaster moving in the hub.
#
# First, lets define the frames of the animation.
# %%
print("Defining animation frames...")
frames = ['00000:00000:00000:00000:00000',
'00900:00000:00000:00000:00000',
'00700:00900:00000:00000:00000',
'00500:00700:00900:00000:00000',
'00000:00500:00700:00900:00000',
'00000:00000:00500:00700:00900',
'00000:00000:00000:00500:00700',
'00000:00000:00000:00000:00500',
'00000:00000:00000:00000:00000']
n_frames = len(frames)
t_pause = 0.05 # Pause between frames (in seconds)
print("DONE!")
# %% [markdown]
# Then, let's proceed with the actual fire!
# %%
print("Firing blasters...")
n_blasters = 3
for ii in range(0, n_blasters):
# Play blaster sound.
hub.sound.play("/extra_files/Laser")
# Display blaster animation.
for ii in range(0, n_frames):
img = hub.Image(frames[ii])
hub.display.show(img)
wait_for_seconds(t_pause)
wait_for_seconds(0.5)
print("DONE!")
# %%
print("Target eliminated.")
# %%
print("-"*15 + " Execution ended " + "-"*15 + "\n")
|
mocs/aat_ms5/programs/aat_ms5.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/mkirby1995/DS-Unit-2-Sprint-3-Classification-Validation/blob/master/LS_DS_231_Logistic_Regression.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="N7SXF6jEBd5_"
# # Lambda School Data Science - Logistic Regression
#
# Logistic regression is the baseline for classification models, as well as a handy way to predict probabilities (since those too live in the unit interval). While relatively simple, it is also the foundation for more sophisticated classification techniques such as neural networks (many of which can effectively be thought of as networks of logistic models).
# + [markdown] colab_type="text" id="E7-AOngjadRN"
# ## Lecture - Where Linear goes Wrong
# ### Return of the Titanic 🚢
#
# You've likely already explored the rich dataset that is the Titanic - let's use regression and try to predict survival with it. The data is [available from Kaggle](https://www.kaggle.com/c/titanic/data), so we'll also play a bit with [the Kaggle API](https://github.com/Kaggle/kaggle-api).
# + [markdown] id="QZAPkIhP6w9K" colab_type="text"
# ### Get data, option 1: Kaggle API
#
# #### Sign up for Kaggle and get an API token
# 1. [Sign up for a Kaggle account](https://www.kaggle.com/), if you don’t already have one.
# 2. [Follow these instructions](https://github.com/Kaggle/kaggle-api#api-credentials) to create a Kaggle “API Token” and download your `kaggle.json` file. If you are using Anaconda, put the file in the directory specified in the instructions.
#
# _This will enable you to download data directly from Kaggle. If you run into problems, don’t worry — I’ll give you an easy alternative way to download today’s data, so you can still follow along with the lecture hands-on. And then we’ll help you through the Kaggle process after the lecture._
# + [markdown] id="rdXsWPp26w9S" colab_type="text"
# #### Put `kaggle.json` in the correct location
#
# - ***If you're using Anaconda,*** put the file in the directory specified in the [instructions](https://github.com/Kaggle/kaggle-api#api-credentials).
#
# - ***If you're using Google Colab,*** upload the file to your Google Drive, and run this cell:
# + id="yZXyy7826w9U" colab_type="code" outputId="c8d2c2fe-61de-4400-d105-4f220b26ff4c" colab={"base_uri": "https://localhost:8080/", "height": 555}
from google.colab import drive
drive.mount('/content/drive')
# %env KAGGLE_CONFIG_DIR=/content/drive/My Drive/
# + [markdown] id="14U2QB0L6w9g" colab_type="text"
# #### Install the Kaggle API package and use it to get the data
#
# You also have to join the Titanic competition to have access to the data
# + colab_type="code" id="MnHLWPYDcyIe" colab={}
# !pip install kaggle
# + id="YE1R2LRp6w93" colab_type="code" colab={}
# !kaggle competitions download -c titanic
# + [markdown] id="Fa_9ckjS6w9-" colab_type="text"
# ### Get data, option 2: Download from the competition page
# 1. [Sign up for a Kaggle account](https://www.kaggle.com/), if you don’t already have one.
# 2. [Go to the Titanic competition page](https://www.kaggle.com/c/titanic) to download the [data](https://www.kaggle.com/c/titanic/data).
# + [markdown] id="o1wVSs6Z6w-A" colab_type="text"
# ### Get data, option 3: Use Seaborn
#
# ```
# import seaborn as sns
# train = sns.load_dataset('titanic')
# ```
#
# But Seaborn's version of the Titanic dataset is not identical to Kaggle's version, as we'll see during this lesson!
# + [markdown] id="_Nn11BA46w-B" colab_type="text"
# ### Read data
# + id="aHdT77fr_itb" colab_type="code" colab={}
import pandas as pd
train = pd.read_csv('train.csv')
test = pd.read_csv('test.csv')
train.shape, test.shape
# + id="tfHLV-ktA8lc" colab_type="code" colab={}
train.head()
# + id="uO4u-JeWBhU9" colab_type="code" colab={}
test.head()
# + id="0Q1TZ_R7BiiV" colab_type="code" colab={}
train.isnull().sum()
# + id="SjSy99mZBlFG" colab_type="code" colab={}
test.isnull().sum()
# + id="eCBk14pDBmac" colab_type="code" colab={}
# + [markdown] id="w7ZY2xeX6w-I" colab_type="text"
# ### How would we try to do this with linear regression?
# + [markdown] id="jxweEr-b6w-J" colab_type="text"
# https://scikit-learn.org/stable/modules/impute.html
# + colab_type="code" id="fcxfpsjdFJwM" colab={}
from sklearn.impute import SimpleImputer
from sklearn.linear_model import LinearRegression
features = ['Pclass', 'Age', 'Fare']
target = 'Survived'
X_train = train[features]
Y_train = train[target]
X_test = test[features]
imputer = SimpleImputer()
X_train_imputed = imputer.fit_transform(X_train)
X_test_imputed = imputer.transform(X_test)
lin_reg = LinearRegression()
lin_reg.fit(X_train_imputed, Y_train)
# + id="5HXN0QQpE-o5" colab_type="code" colab={}
import numpy as np
test_case = np.array([[1, 5, 500]]) #Rich 5 yr old in first class
lin_reg.predict(test_case)
# + [markdown] id="lT6UW4QOFk9i" colab_type="text"
# super good survival chance 119%
# + [markdown] id="OJ7nMbxH6w-N" colab_type="text"
# ### How would we do this with Logistic Regression?
# + colab_type="code" id="dpUm8Dl-u2aB" colab={}
from sklearn.linear_model import LogisticRegression
log_reg = LogisticRegression(solver='lbfgs')
log_reg.fit(X_train_imputed, Y_train)
print(log_reg.predict(test_case))
print(log_reg.predict_proba(test_case))
# + id="5Ww052bkKH0E" colab_type="code" colab={}
log_reg.predict(X_test_imputed)
# + [markdown] id="1gxkIuca6w-w" colab_type="text"
# ### How accurate is the Logistic Regression?
# + id="6J5DmvZQ6w-y" colab_type="code" colab={}
# + [markdown] id="Hi4en0Cn6w-3" colab_type="text"
# ### What's the math for the Logistic Regression?
#
# https://en.wikipedia.org/wiki/Logistic_function
#
# https://en.wikipedia.org/wiki/Logistic_regression#Probability_of_passing_an_exam_versus_hours_of_study
# + colab_type="code" id="9Bq-54noR1uE" colab={}
# + [markdown] id="9-nsrVAP6w-9" colab_type="text"
# ## Feature Engineering
#
# Get the [Category Encoder](http://contrib.scikit-learn.org/categorical-encoding/) library
#
# If you're running on Google Colab:
#
# ```
# # # !pip install category_encoders
# ```
#
# If you're running locally with Anaconda:
#
# ```
# # # !conda install -c conda-forge category_encoders
# ```
# + id="b5whcSZZ6w--" colab_type="code" colab={}
# + [markdown] colab_type="text" id="iblW74C8afuR"
# ## Assignment: real-world classification
#
# We're going to check out a larger dataset - the [FMA Free Music Archive data](https://github.com/mdeff/fma). It has a selection of CSVs with metadata and calculated audio features that you can load and try to use to classify genre of tracks. To get you started:
# + [markdown] id="WBPaXK3Y6w_M" colab_type="text"
# ### Get and unzip the data
# + [markdown] id="eRc-9a4D6w_N" colab_type="text"
# #### Google Colab
# + colab_type="code" id="SsySnuKaKtQf" outputId="046ceb24-5ddd-4214-e5ce-6dc8dffb25eb" colab={"base_uri": "https://localhost:8080/", "height": 437}
# !wget https://os.unil.cloud.switch.ch/fma/fma_metadata.zip
# !unzip fma_metadata.zip
# + [markdown] id="Dz8rFhmp6w_Q" colab_type="text"
# #### Windows
# - Download the [zip file](https://os.unil.cloud.switch.ch/fma/fma_metadata.zip)
# - You may need to use [7zip](https://www.7-zip.org/download.html) to unzip it
#
#
# #### Mac
# - Download the [zip file](https://os.unil.cloud.switch.ch/fma/fma_metadata.zip)
# - You may need to use [p7zip](https://superuser.com/a/626731) to unzip it
# + [markdown] id="IgT3j3n56w_R" colab_type="text"
# ### Look at first 3 lines of raw file
# + id="Rp6lckeM6w_S" colab_type="code" outputId="e935bf41-0aca-4bf0-90bd-fbaa24b73af1" colab={"base_uri": "https://localhost:8080/", "height": 174}
# !head -n 4 fma_metadata/tracks.csv
# + [markdown] id="4AlDvAka6w_V" colab_type="text"
# ### Read with pandas
# https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.read_csv.html
# + id="1tvkmh3u6w_W" colab_type="code" colab={}
import pandas as pd
from sklearn.impute import SimpleImputer
import matplotlib.pyplot as plt
import numpy as np
from sklearn.linear_model import LinearRegression
from sklearn.linear_model import LogisticRegression
# + id="PcNAXsVLoyXu" colab_type="code" colab={}
tracks = pd.read_csv('fma_metadata/tracks.csv', header=[0,1], index_col=0)
# + id="nvEWmKuNrcOX" colab_type="code" outputId="14f7980a-ac53-4ec8-f044-eb8cf2b5837a" colab={"base_uri": "https://localhost:8080/", "height": 590}
tracks.head()
# + id="A6UAJUcRrfv1" colab_type="code" outputId="ef2d561c-9219-4b70-a866-8ce66359ca70" colab={"base_uri": "https://localhost:8080/", "height": 34}
tracks.shape
# + [markdown] id="cALwu774cRvB" colab_type="text"
#
#
# ---
#
#
# + [markdown] id="2vrz_qipbXd0" colab_type="text"
# ####Create Df for Artist Album and Track
# + id="s3r6nG5filaJ" colab_type="code" colab={}
tracks['album', 'track_genre'] = tracks['track', 'genre_top']
tracks['artist', 'track_genre'] = tracks['track', 'genre_top']
# + id="ZisLmEwGbU_-" colab_type="code" colab={}
album_df = tracks['album']
artist_df = tracks['artist']
track_df = tracks['track']
# + [markdown] id="5lVKCu9TcJkZ" colab_type="text"
#
#
# ---
#
#
# + [markdown] id="rDD16fzt6w_Z" colab_type="text"
# ### Fit Logistic Regression!
# + [markdown] id="bJMlUZfHd7O_" colab_type="text"
# ####Clean and Concatenate dataframes into one large dataframe
# + [markdown] id="dfC6kQcNflNk" colab_type="text"
# #####Clean album_df
# + id="7nK5Y33ieKfy" colab_type="code" colab={}
album_df = album_df.drop(columns=['information',
'engineer',
'producer',
'tags',
'type',
'date_created',
'date_released',
'title'])
# + id="8pq_1nsWeE83" colab_type="code" colab={}
album_df = album_df.rename(columns={'comments': 'album_comments',
'favorites': 'album_favorites',
'id': 'album_id',
'listens': 'album_listens',
'tracks': 'album_track_number'})
# + [markdown] id="xmWHDLRPiaXy" colab_type="text"
#
#
# ---
#
#
# + [markdown] id="Q9mDQvGhfnyj" colab_type="text"
# #####Clean artist_df
# + id="OM93woXQf-y9" colab_type="code" colab={}
art_col_drop = []
for _ in range(len(artist_df.dtypes)):
#print(artist_df.dtypes[_])
if artist_df.dtypes[_] == object:
art_col_drop.append(artist_df.columns[_])
# + id="AV_ErcpNfqEn" colab_type="code" colab={}
artist_df = artist_df.drop(columns=art_col_drop)
artist_df = artist_df.drop(columns=['latitude', 'longitude'])
# + id="mtK8tIAgfqUv" colab_type="code" colab={}
artist_df = artist_df.rename(columns={'comments': 'artist_comments',
'favorites': 'artist_favorites',
'id': 'artist_id'})
# + [markdown] id="EI3P_pncibEz" colab_type="text"
#
#
# ---
#
#
# + [markdown] id="A_wzsBLCg3vA" colab_type="text"
# #####clean track_df
# + id="GTQSnIBIhLFQ" colab_type="code" colab={}
track_df = track_df.drop(columns=['composer',
'date_created',
'date_recorded',
'genres',
'genres_all',
'information',
'language_code',
'license',
'lyricist',
'publisher',
'tags',
'title'])
# + id="6ZL__mlag5W7" colab_type="code" colab={}
track_df = track_df.rename(columns={'bit_rate':'track_bit_rate',
'comments':'track_comments',
'duration':'track_duration',
'favorites':'track_favorites',
'genre_top':'track_genre',
'interest':'track_interest',
'listens':'track_listens',
'number':'track_number',})
# + [markdown] id="XDVbUi46ib3T" colab_type="text"
#
#
# ---
#
#
# + [markdown] id="pdoz_VJlicQo" colab_type="text"
# #####Concatenate Dataframes
# + id="-54k_f7sifzk" colab_type="code" colab={}
df = pd.concat([album_df, artist_df, track_df], axis=0, sort=False)
# + id="NHwD0HWrjyBF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 337} outputId="921b6200-1b0d-43e7-fd5a-cd53d80c1e4a"
df.describe()
# + id="GtKnqtkOkVpz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 173} outputId="db0daa06-1ae0-49d8-afc0-1fe123188b41"
df.describe(exclude='number')
# + [markdown] id="OcoP8rRgd7k7" colab_type="text"
#
#
# ---
#
#
# + [markdown] id="7JdDX8an3Ruy" colab_type="text"
# #####Encode `genre_top`
# + id="V9W1MxtUz4iw" colab_type="code" colab={}
subset = df.dropna(subset=['track_genre'])
# + id="7KvZAIfJ1uYy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 102} outputId="a010a57d-0a8a-41b2-a387-1f38c5852549"
subset['track_genre'].replace({'Rock':1,
'Experimental':2,
'Electronic':2,
'Hip-Hop':1,
'Folk':1,
'Pop':2,
'Instrumental':3,
'International':3,
'Classical':3,
'Jazz':1,
'Old-Time / Historic':3,
'Spoken':4,
'Country':1,
'Soul-RnB':1,
'Blues':1,
'Easy Listening':4},
inplace=True)
# + [markdown] id="PfFwwWUT3adl" colab_type="text"
#
#
# ---
#
#
# + [markdown] colab_type="text" id="kQUVlUKQMPPW"
# This dataset is bigger than many you've worked with so far, and while it should fit in Colab, it can take awhile to run. That's part of the challenge!
#
# Your tasks:
# - Clean up the variable names in the dataframe
# - Use logistic regression to fit a model predicting (primary/top) genre
# - Inspect, iterate, and improve your model
# - Answer the following questions (written, ~paragraph each):
# - What are the best predictors of genre?
# - What information isn't very useful for predicting genre?
# - What surprised you the most about your results?
#
# *Important caveats*:
# - This is going to be difficult data to work with - don't let the perfect be the enemy of the good!
# - Be creative in cleaning it up - if the best way you know how to do it is download it locally and edit as a spreadsheet, that's OK!
# - If the data size becomes problematic, consider sampling/subsetting, or [downcasting numeric datatypes](https://www.dataquest.io/blog/pandas-big-data/).
# - You do not need perfect or complete results - just something plausible that runs, and that supports the reasoning in your written answers
#
# If you find that fitting a model to classify *all* genres isn't very good, it's totally OK to limit to the most frequent genres, or perhaps trying to combine or cluster genres as a preprocessing step. Even then, there will be limits to how good a model can be with just this metadata - if you really want to train an effective genre classifier, you'll have to involve the other data (see stretch goals).
#
# This is real data - there is no "one correct answer", so you can take this in a variety of directions. Just make sure to support your findings, and feel free to share them as well! This is meant to be practice for dealing with other "messy" data, a common task in data science.
# + [markdown] id="dpK78pV9uPyS" colab_type="text"
# #### Model
# + id="dejNVoP9uZEO" colab_type="code" colab={}
target = 'track_genre'
features = subset.drop(columns='track_genre').columns.tolist()
# + id="dBYbUFzB0z3l" colab_type="code" colab={}
features = subset[features]
target = subset[target]
imputer = SimpleImputer()
features_imputed = imputer.fit_transform(features)
# + id="Twf2wyAauMz-" colab_type="code" outputId="d5f30965-f570-4b19-e8c1-d359ad28ea6c" colab={"base_uri": "https://localhost:8080/", "height": 139}
log_reg = LogisticRegression(solver='lbfgs', multi_class ='auto', max_iter=1000)
log_reg.fit(features_imputed, target)
# + id="L6hKgyuC5lL6" colab_type="code" outputId="c791db6a-9580-4448-c656-9522e6b7d46b" colab={"base_uri": "https://localhost:8080/", "height": 34}
log_reg.score(features_imputed, target)
# + [markdown] colab_type="text" id="wlI5OXfSag9C"
# ## Resources and stretch goals
#
# - Check out the other .csv files from the FMA dataset, and see if you can join them or otherwise fit interesting models with them
# - [Logistic regression from scratch in numpy](https://blog.goodaudience.com/logistic-regression-from-scratch-in-numpy-5841c09e425f) - if you want to dig in a bit more to both the code and math (also takes a gradient descent approach, introducing the logistic loss function)
# - Create a visualization to show predictions of your model - ideally show a confidence interval based on error!
# - Check out and compare classification models from scikit-learn, such as [SVM](https://scikit-learn.org/stable/modules/svm.html#classification), [decision trees](https://scikit-learn.org/stable/modules/tree.html#classification), and [naive Bayes](https://scikit-learn.org/stable/modules/naive_bayes.html). The underlying math will vary significantly, but the API (how you write the code) and interpretation will actually be fairly similar.
# - Sign up for [Kaggle](https://kaggle.com), and find a competition to try logistic regression with
# - (Not logistic regression related) If you enjoyed the assignment, you may want to read up on [music informatics](https://en.wikipedia.org/wiki/Music_informatics), which is how those audio features were actually calculated. The FMA includes the actual raw audio, so (while this is more of a longterm project than a stretch goal, and won't fit in Colab) if you'd like you can check those out and see what sort of deeper analysis you can do.
# + [markdown] id="lGW152MB7Iis" colab_type="text"
# ###Visualization
# + id="EKRZza59JG3z" colab_type="code" colab={}
feature_list = [features['album_comments'],
features['album_favorites'],
features['album_id'],
features['album_listens'],
features['album_track_number'],
features['artist_comments'],
features['artist_favorites'],
features['artist_id'],
features['track_bit_rate'],
features['track_comments'],
features['track_duration'],
features['track_favorites'],
features['track_interest'],
features['track_listens'],
features['track_number']]
# + id="58ny9e51I7DW" colab_type="code" colab={}
features['total'] = sum(feature_list)
# + id="q8CoYxxIG2Yw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a5f8c559-bc12-466c-8ca6-b8dadf0e1fd0"
log_reg.predict(features_imputed)
# + id="Njqba6gTH_qi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 269} outputId="ed2e0024-79db-4ce6-f0eb-1d8af5aa81ab"
plt.scatter(features['total'], log_reg.predict(features_imputed));
# + id="sFbktFl9G1Cz" colab_type="code" colab={}
|
LS_DS_231_Logistic_Regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
import gensim
from os import path
from glob import glob
import numpy as np
with open("/Users/Belal/Projects/jobs/i2x_job/keyword_xtract/script.txt", "r") as f:
data = f.read()
model.save_word2vec_format("shizer", fvocab="shizer_vocab.txt", binary=True)
# +
# ====== KEYWORD EXTRACTION ======
# ================================
stopword = "/Users/Belal/Projects/jobs/i2x_job/keyword_xtract/stop_words/sklearn_stopwords.txt"
inputt = "/Users/Belal/Projects/jobs/i2x_job/keyword_xtract/script.txt"
# Initialize RAKE object,
rake_object = Rake(stop_words_path=stopword, min_char_length=4,
max_words_length=4, min_keyword_frequency=3)
# 2. run on RAKE on a given text
sample_file = open(inputt, 'r')
text = sample_file.read()
keywords = rake_object.run(text)
# +
# ======= KEYWORD RANKING ========
# ================================
model = "/Users/Belal/Projects/jobs/i2x_job/keyword_xtract/w2v_models/GoogleNews-vectors-negative300.bin.gz"
print("loading Word2Vec model...")
model = gensim.models.KeyedVectors.load_word2vec_format(model, limit=150000, binary=True)
print("loaded model!")
# +
test = "evaluation/"
test_dirs = glob(path.join(test, "*txt"))
test_docs = [doc.read() for doc in [open(test_file, "r") for test_file in test_dirs]]
test_vecs = [get_avg_feature_vecs([doc],
model=model,
num_features=model.vector_size)
for doc in test_docs]
# +
from itertools import compress
index2word_set = set(model.index2word)
bool_split = [word[0] in index2word_set for word in keywords]
keyword_in_model = list(compress(keywords, bool_split))
# +
#sort keywords + choose how many to use
sorted_keyword = sorted(keyword_in_model, key=lambda x: x[1], reverse=True)
sorted_keyword = sorted(keyword_in_model, key=lambda x: x[1], reverse=True)
n_keywords = int(0.25*len(sorted_keyword))
keyword_list = sorted_keyword[0:n_keywords]
# -
keyword_vecs = [(model.word_vec(word[0])) for word in sorted_keyword]
# generating candidate words from test docs (optional)
# test_words = generate_candidate_keywords(split_sentences(test_docs[0]), stopword_pattern=stopword,
# min_char_length=2, max_words_length=2)
# +
test = "../evaluation/"
test_dirs = glob(path.join(test, "*txt"))
test_docs = [doc.read() for doc in [open(test_file, "r") for test_file in test_dirs]]
test_vecs = [get_avg_feature_vecs([doc],
model=model,
num_features=model.vector_size)
for doc in test_docs]
# -
#Ranking
from sklearn.metrics import pairwise
x=[]
for vec in test_vecs:
for key_word in keyword_vecs:
x.append(pairwise.cosine_similarity(X=key_word.reshape(1,-1), Y = vec.reshape(1,-1)))
x=[]
for vec in test_vecs:
x.append([pairwise.cosine_similarity(X=key_word.reshape(1,-1), Y = vec.reshape(1,-1)) for key_word in keyword_vecs])
z=np.zeros_like(x[0])
for doc in x:
sum_keyword = z + doc
len(keyword_list)
names_key = [k[0] for k in sorted_keyword]
# adding cosine similarities to get a single 'rank' for each keyword
z=np.zeros_like(x[0])
for y in x:
z=z+y
newlist = z/3
final = list(zip(names_key, newlist))
fff=[]
for i in range(len(names_key)):
fff.append((names_key[i], newlist[i][0][0]))
ranked = sorted(fff, key=lambda x: x[1], reverse=True)
ranked
out = "bla.txt"
file.
# +
# ========= SAVE OUTPUT ==========
# ================================
print("saving results")
with open("bla.txt","w") as f:
for line in ranked:
strs=" score: ".join(str(x) for x in line)
f.write(strs+"\n")
# -
for line in ranked:
strs=" score: ".join(str(x) for x in line)
print(strs)
# ###
# +
# #!/usr/bin/env python
# -*- coding: utf-8 -*-
# Implementation of RAKE - Rapid Automatic Keyword Extraction algorithm
# as described in:
# <NAME>., <NAME>, <NAME>, and <NAME> (2010).
# Automatic keyword extraction from individual documents.
# In <NAME> and <NAME> (Eds.), Text Mining: Applications and Theory.unknown: John Wiley and Sons, Ltd.
#
# NOTE: The original implementation (available at - https://github.com/zelandiya/RAKE-tutorial)
# has been extended and updated to work with Python 3 and to include more specific functionality
import re
import operator
import six
from six.moves import range
# Required functions for RAKE
def is_number(s):
try:
float(s) if '.' in s else int(s)
return True
except ValueError:
return False
def load_stop_words(stop_word_file):
"""
Utility function to load stop words from a file and return as a list of words
@param stop_word_file Path and file name of a file containing stop words.
@return list A list of stop words.
"""
stop_words = []
for line in open(stop_word_file):
if line.strip()[0:1] != "#":
for word in line.split(): # in case more than one per line
stop_words.append(word)
return stop_words
def separate_words(text, min_word_return_size):
"""
Utility function to return a list of all words that are have a length greater than a specified number of characters.
@param text The text that must be split in to words.
@param min_word_return_size The minimum no of characters a word must have to be included.
"""
splitter = re.compile('[^a-zA-Z0-9_\\+\\-/]')
words = []
for single_word in splitter.split(text):
current_word = single_word.strip().lower()
#leave numbers in phrase, but don't count as words, since they tend to invalidate scores of their phrases
if len(current_word) > min_word_return_size and current_word != '' and not is_number(current_word):
words.append(current_word)
return words
def split_sentences(text):
"""
Utility function to return a list of sentences.
@param text The text that must be split in to sentences.
"""
sentence_delimiters = re.compile(u'[\\[\\]\n.!?,;:\t\\-\\"\\(\\)\\\'\u2019\u2013]')
sentences = sentence_delimiters.split(text)
return sentences
def build_stop_word_regex(stop_word_file_path):
stop_word_list = load_stop_words(stop_word_file_path)
stop_word_regex_list = []
for word in stop_word_list:
word_regex = '\\b' + word + '\\b'
stop_word_regex_list.append(word_regex)
stop_word_pattern = re.compile('|'.join(stop_word_regex_list), re.IGNORECASE)
return stop_word_pattern
def generate_candidate_keywords(sentence_list, stopword_pattern, min_char_length=1, max_words_length=5):
phrase_list = []
for s in sentence_list:
tmp = re.sub(stopword_pattern, '|', s.strip())
phrases = tmp.split("|")
for phrase in phrases:
phrase = phrase.strip().lower()
if phrase != "" and is_acceptable(phrase, min_char_length, max_words_length):
phrase_list.append(phrase)
return phrase_list
def is_acceptable(phrase, min_char_length, max_words_length):
# a phrase must have a min length in characters
if len(phrase) < min_char_length:
return 0
# a phrase must have a max number of words
words = phrase.split()
if len(words) > max_words_length:
return 0
digits = 0
alpha = 0
for i in range(0, len(phrase)):
if phrase[i].isdigit():
digits += 1
elif phrase[i].isalpha():
alpha += 1
# a phrase must have at least one alpha character
if alpha == 0:
return 0
# a phrase must have more alpha than digits characters
if digits > alpha:
return 0
return 1
def calculate_word_scores(phraseList):
word_frequency = {}
word_degree = {}
for phrase in phraseList:
word_list = separate_words(phrase, 0)
word_list_length = len(word_list)
word_list_degree = word_list_length - 1
# if word_list_degree > 3: word_list_degree = 3 #exp.
for word in word_list:
word_frequency.setdefault(word, 0)
word_frequency[word] += 1
word_degree.setdefault(word, 0)
word_degree[word] += word_list_degree # orig.
# word_degree[word] += 1/(word_list_length*1.0) #exp.
for item in word_frequency:
word_degree[item] = word_degree[item] + word_frequency[item]
# Calculate Word scores = deg(w)/freq(w)
word_score = {}
for item in word_frequency:
word_score.setdefault(item, 0)
word_score[item] = word_degree[item] / (word_frequency[item] * 1.0) #orig.
# word_score[item] = word_frequency[item]/(word_degree[item] * 1.0) #exp.
return word_score
def generate_candidate_keyword_scores(phrase_list, word_score, min_keyword_frequency=1):
keyword_candidates = {}
for phrase in phrase_list:
if min_keyword_frequency > 1:
if phrase_list.count(phrase) < min_keyword_frequency:
continue
keyword_candidates.setdefault(phrase, 0)
word_list = separate_words(phrase, 0)
candidate_score = 0
for word in word_list:
candidate_score += word_score[word]
keyword_candidates[phrase] = candidate_score
return keyword_candidates
class Rake(object):
def __init__(self, stop_words_path, min_char_length=1, max_words_length=5, min_keyword_frequency=1):
self.__stop_words_path = stop_words_path
self.__stop_words_pattern = build_stop_word_regex(stop_words_path)
self.__min_char_length = min_char_length
self.__max_words_length = max_words_length
self.__min_keyword_frequency = min_keyword_frequency
def run(self, text):
sentence_list = split_sentences(text)
phrase_list = generate_candidate_keywords(sentence_list, self.__stop_words_pattern,
self.__min_char_length, self.__max_words_length)
word_scores = calculate_word_scores(phrase_list)
keyword_candidates = generate_candidate_keyword_scores(phrase_list, word_scores, self.__min_keyword_frequency)
sorted_keywords = sorted(six.iteritems(keyword_candidates), key=operator.itemgetter(1), reverse=True)
return sorted_keywords
test=None
# Testing + debugging RAKE on pre-defined text block
if test:
text = "Compatibility of systems of linear constraints over the set of natural numbers. " \
"Criteria of compatibility of a system of linear Diophantine equations, strict inequations," \
" and nonstrict inequations are considered. Upper bounds for components of a minimal set of " \
"solutions and algorithms of construction of minimal generating sets of solutions for all types" \
" of systems are given. These criteria and the corresponding algorithms for constructing a minimal" \
" supporting set of solutions can be used in solving all the considered" \
" types of systems and systems of mixed types."
# Split text into sentences
sentenceList = split_sentences(text)
stoppath = "stop_words/sklearn_stopwords.txt"
stopwordpattern = build_stop_word_regex(stoppath)
# generate candidate keywords
phraseList = generate_candidate_keywords(sentenceList, stopwordpattern)
# calculate individual word scores
wordscores = calculate_word_scores(phraseList)
# generate candidate keyword scores
keywordcandidates = generate_candidate_keyword_scores(phraseList, wordscores)
if debug:
print(keywordcandidates)
sortedKeywords = sorted(six.iteritems(keywordcandidates), key=operator.itemgetter(1), reverse=True)
if debug:
print(sortedKeywords)
totalKeywords = len(sortedKeywords)
if debug:
print(totalKeywords)
print(sortedKeywords[0:(totalKeywords // 3)])
rake = Rake("stop_words/sklearn_stopwords.txt")
keywords = rake.run(text)
print(keywords)
# +
def make_feature_vec(words, model, num_features):
"""
Function to average all of the word vectors in a given paragraph
:param words:
:param model:
:param num_features:
:return:
"""
# Pre-initialize an empty numpy array (for speed)
feature_vec = np.zeros((num_features,), dtype="float32")
n_words = 0
# Index2word is a list that contains the names of the words in
# the model's vocabulary. Convert it to a set, for speed
index2word_set = set(model.index2word)
# Loop over each word in the review and, if it is in the model's
# vocabulary, add its feature vector to the total
for word in words:
if word in index2word_set:
n_words += 1
feature_vec = np.add(feature_vec, model[word])
# Divide the result by the number of words to get the average
feature_vec = np.divide(feature_vec, n_words)
return feature_vec
def get_avg_feature_vecs(reviews, model, num_features):
# Given a set of reviews (each one a list of words), calculate
# the average feature vector for each one and return a 2D numpy array
#
# Initialize a counter
counter = 0
#
# Pre-allocate a 2D numpy array, for speed
review_feature_vecs = np.zeros((len(reviews), num_features), dtype="float32")
#
# Loop through the reviews
for review in reviews:
# Print a status message
# if counter % 1000 == 0:
print("Review %d of %d" % (counter, len(reviews)))
# Call the function (defined above) that makes average feature vectors
review_feature_vecs[counter] = make_feature_vec(review, model, num_features)
# Increment the counter
counter += 1
return review_feature_vecs
# -
|
notebooks/RAKE.ipynb
|
# ---
# jupyter:
# jupytext:
# split_at_heading: true
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#hide
#skip
! [ -e /content ] && pip install -Uqq fastai # upgrade fastai on colab
#export
from fastai.torch_basics import *
from fastai.data.all import *
from fastai.text.core import *
#hide
from nbdev.showdoc import *
# +
#default_exp text.data
#default_cls_lvl 3
# -
# # Text data
#
# > Functions and transforms to help gather text data in a `Datasets`
# ## Backwards
#
# Reversing the text can provide higher accuracy with an ensemble with a forward model. All that is needed is a `type_tfm` that will reverse the text as it is brought in:
#export
def reverse_text(x): return x.flip(0)
t = tensor([0,1,2])
r = reverse_text(t)
test_eq(r, tensor([2,1,0]))
# ## Numericalizing
# Numericalization is the step in which we convert tokens to integers. The first step is to build a correspondence token to index that is called a vocab.
#export
def make_vocab(count, min_freq=3, max_vocab=60000, special_toks=None):
"Create a vocab of `max_vocab` size from `Counter` `count` with items present more than `min_freq`"
vocab = [o for o,c in count.most_common(max_vocab) if c >= min_freq]
special_toks = ifnone(special_toks, defaults.text_spec_tok)
for o in reversed(special_toks): #Make sure all special tokens are in the vocab
if o in vocab: vocab.remove(o)
vocab.insert(0, o)
vocab = vocab[:max_vocab]
return vocab + [f'xxfake' for i in range(0, 8-len(vocab)%8)]
# If there are more than `max_vocab` tokens, the ones kept are the most frequent.
#
# > Note: For performance when using mixed precision, the vocabulary is always made of size a multiple of 8, potentially by adding `xxfake` tokens.
count = Counter(['a', 'a', 'a', 'a', 'b', 'b', 'c', 'c', 'd'])
test_eq(set([x for x in make_vocab(count) if not x.startswith('xxfake')]),
set(defaults.text_spec_tok + 'a'.split()))
test_eq(len(make_vocab(count))%8, 0)
test_eq(set([x for x in make_vocab(count, min_freq=1) if not x.startswith('xxfake')]),
set(defaults.text_spec_tok + 'a b c d'.split()))
test_eq(set([x for x in make_vocab(count,max_vocab=12, min_freq=1) if not x.startswith('xxfake')]),
set(defaults.text_spec_tok + 'a b c'.split()))
# +
#export
class TensorText(TensorBase): pass
class LMTensorText(TensorText): pass
TensorText.__doc__ = "Semantic type for a tensor representing text"
LMTensorText.__doc__ = "Semantic type for a tensor representing text in language modeling"
# -
#export
class Numericalize(Transform):
"Reversible transform of tokenized texts to numericalized ids"
def __init__(self, vocab=None, min_freq=3, max_vocab=60000, special_toks=None):
store_attr('vocab,min_freq,max_vocab,special_toks')
self.o2i = None if vocab is None else defaultdict(int, {v:k for k,v in enumerate(vocab)})
def setups(self, dsets):
if dsets is None: return
if self.vocab is None:
count = dsets.counter if getattr(dsets, 'counter', None) is not None else Counter(p for o in dsets for p in o)
if self.special_toks is None and hasattr(dsets, 'special_toks'):
self.special_toks = dsets.special_toks
self.vocab = make_vocab(count, min_freq=self.min_freq, max_vocab=self.max_vocab, special_toks=self.special_toks)
self.o2i = defaultdict(int, {v:k for k,v in enumerate(self.vocab) if v != 'xxfake'})
def encodes(self, o): return TensorText(tensor([self.o2i [o_] for o_ in o]))
def decodes(self, o): return L(self.vocab[o_] for o_ in o)
num = Numericalize(min_freq=2)
num.setup(L('This is an example of text'.split(), 'this is another text'.split()))
start = 'This is an example of text '
# If no `vocab` is passed, one is created at setup from the data, using `make_vocab` with `min_freq` and `max_vocab`.
# +
start = 'This is an example of text'
num = Numericalize(min_freq=1)
num.setup(L(start.split(), 'this is another text'.split()))
test_eq(set([x for x in num.vocab if not x.startswith('xxfake')]),
set(defaults.text_spec_tok + 'This is an example of text this another'.split()))
test_eq(len(num.vocab)%8, 0)
t = num(start.split())
test_eq(t, tensor([11, 9, 12, 13, 14, 10]))
test_eq(num.decode(t), start.split())
# -
num = Numericalize(min_freq=2)
num.setup(L('This is an example of text'.split(), 'this is another text'.split()))
test_eq(set([x for x in num.vocab if not x.startswith('xxfake')]),
set(defaults.text_spec_tok + 'is text'.split()))
test_eq(len(num.vocab)%8, 0)
t = num(start.split())
test_eq(t, tensor([0, 9, 0, 0, 0, 10]))
test_eq(num.decode(t), f'{UNK} is {UNK} {UNK} {UNK} text'.split())
#hide
df = pd.DataFrame({'texts': ['This is an example of text', 'this is another text']})
tl = TfmdLists(df, [attrgetter('text'), Tokenizer.from_df('texts'), Numericalize(min_freq=2)])
test_eq(tl, [tensor([2, 8, 9, 10, 0, 0, 0, 11]), tensor([2, 9, 10, 0, 11])])
# ## LM_DataLoader -
#export
def _maybe_first(o): return o[0] if isinstance(o, tuple) else o
#export
def _get_tokenizer(ds):
tok = getattr(ds, 'tokenizer', None)
if isinstance(tok, Tokenizer): return tok
if isinstance(tok, (list,L)):
for t in tok:
if isinstance(t, Tokenizer): return t
#export
def _get_lengths(ds):
tok = _get_tokenizer(ds)
if tok is None: return
return tok.get_lengths(ds.items)
#export
#TODO: add backward
@delegates()
class LMDataLoader(TfmdDL):
"A `DataLoader` suitable for language modeling"
def __init__(self, dataset, lens=None, cache=2, bs=64, seq_len=72, num_workers=0, **kwargs):
self.items = ReindexCollection(dataset, cache=cache, tfm=_maybe_first)
self.seq_len = seq_len
if lens is None: lens = _get_lengths(dataset)
if lens is None: lens = [len(o) for o in self.items]
self.lens = ReindexCollection(lens, idxs=self.items.idxs)
# The "-1" is to allow for final label, we throw away the end that's less than bs
corpus = round_multiple(sum(lens)-1, bs, round_down=True)
self.bl = corpus//bs #bl stands for batch length
self.n_batches = self.bl//(seq_len) + int(self.bl%seq_len!=0)
self.last_len = self.bl - (self.n_batches-1)*seq_len
self.make_chunks()
super().__init__(dataset=dataset, bs=bs, num_workers=num_workers, **kwargs)
self.n = self.n_batches*bs
def make_chunks(self): self.chunks = Chunks(self.items, self.lens)
def shuffle_fn(self,idxs):
self.items.shuffle()
self.make_chunks()
return idxs
def create_item(self, seq):
if seq>=self.n: raise IndexError
sl = self.last_len if seq//self.bs==self.n_batches-1 else self.seq_len
st = (seq%self.bs)*self.bl + (seq//self.bs)*self.seq_len
txt = self.chunks[st : st+sl+1]
return LMTensorText(txt[:-1]),txt[1:]
@delegates(TfmdDL.new)
def new(self, dataset=None, seq_len=None, **kwargs):
lens = self.lens.coll if dataset is None else None
seq_len = self.seq_len if seq_len is None else seq_len
return super().new(dataset=dataset, lens=lens, seq_len=seq_len, **kwargs)
show_doc(LMDataLoader, title_level=2)
# `dataset` should be a collection of numericalized texts for this to work. `lens` can be passed for optimizing the creation, otherwise, the `LMDataLoader` will do a full pass of the `dataset` to compute them. `cache` is used to avoid reloading items unnecessarily.
#
# The `LMDataLoader` will concatenate all texts (maybe `shuffle`d) in one big stream, split it in `bs` contiguous sentences, then go through those `seq_len` at a time.
#hide
bs,sl = 4,3
ints = L([0,1,2,3,4],[5,6,7,8,9,10],[11,12,13,14,15,16,17,18],[19,20],[21,22]).map(tensor)
dl = LMDataLoader(ints, bs=bs, seq_len=sl)
list(dl)
test_eq(list(dl),
[[tensor([[0, 1, 2], [5, 6, 7], [10, 11, 12], [15, 16, 17]]),
tensor([[1, 2, 3], [6, 7, 8], [11, 12, 13], [16, 17, 18]])],
[tensor([[3, 4], [8, 9], [13, 14], [18, 19]]),
tensor([[4, 5], [9, 10], [14, 15], [19, 20]])]])
bs,sl = 4,3
ints = L([0,1,2,3,4],[5,6,7,8,9,10],[11,12,13,14,15,16,17,18],[19,20],[21,22,23],[24]).map(tensor)
dl = LMDataLoader(ints, bs=bs, seq_len=sl)
test_eq(list(dl),
[[tensor([[0, 1, 2], [6, 7, 8], [12, 13, 14], [18, 19, 20]]),
tensor([[1, 2, 3], [7, 8, 9], [13, 14, 15], [19, 20, 21]])],
[tensor([[3, 4, 5], [ 9, 10, 11], [15, 16, 17], [21, 22, 23]]),
tensor([[4, 5, 6], [10, 11, 12], [16, 17, 18], [22, 23, 24]])]])
#hide
#Check lens work
dl = LMDataLoader(ints, lens=ints.map(len), bs=bs, seq_len=sl)
test_eq(list(dl),
[[tensor([[0, 1, 2], [6, 7, 8], [12, 13, 14], [18, 19, 20]]),
tensor([[1, 2, 3], [7, 8, 9], [13, 14, 15], [19, 20, 21]])],
[tensor([[3, 4, 5], [ 9, 10, 11], [15, 16, 17], [21, 22, 23]]),
tensor([[4, 5, 6], [10, 11, 12], [16, 17, 18], [22, 23, 24]])]])
dl = LMDataLoader(ints, bs=bs, seq_len=sl, shuffle=True)
for x,y in dl: test_eq(x[:,1:], y[:,:-1])
((x0,y0), (x1,y1)) = tuple(dl)
#Second batch begins where first batch ended
test_eq(y0[:,-1], x1[:,0])
test_eq(type(x0), LMTensorText)
#hide
#test new works
dl = LMDataLoader(ints, bs=bs, seq_len=sl, shuffle=True)
dl1 = dl.new()
test_eq(dl1.seq_len, sl)
dl2 = dl.new(seq_len=2)
test_eq(dl2.seq_len, 2)
# ### Showing -
#export
@typedispatch
def show_batch(x: TensorText, y, samples, ctxs=None, max_n=10, trunc_at=150, **kwargs):
if ctxs is None: ctxs = get_empty_df(min(len(samples), max_n))
if trunc_at is not None: samples = L((s[0].truncate(trunc_at),*s[1:]) for s in samples)
ctxs = show_batch[object](x, y, samples, max_n=max_n, ctxs=ctxs, **kwargs)
display_df(pd.DataFrame(ctxs))
return ctxs
#export
@typedispatch
def show_batch(x: LMTensorText, y, samples, ctxs=None, max_n=10, trunc_at=150, **kwargs):
samples = L((s[0].truncate(trunc_at), s[1].truncate(trunc_at)) for s in samples)
return show_batch[TensorText](x, None, samples, ctxs=ctxs, max_n=max_n, trunc_at=None, **kwargs)
# ## Classification
# For classification, we deal with the fact that texts don't all have the same length by using padding.
# export
class Pad_Input(ItemTransform):
def encodes(self,samples, pad_idx=1, pad_fields=0, pad_first=False, backwards=False):
"Function that collect `samples` and adds padding"
self.pad_idx = pad_idx
pad_fields = L(pad_fields)
max_len_l = pad_fields.map(lambda f: max([len(s[f]) for s in samples]))
if backwards: pad_first = not pad_first
def _f(field_idx, x):
if field_idx not in pad_fields: return x
idx = pad_fields.items.index(field_idx) #TODO: remove items if L.index is fixed
sl = slice(-len(x), sys.maxsize) if pad_first else slice(0, len(x))
pad = x.new_zeros(max_len_l[idx]-x.shape[0])+pad_idx
x1 = torch.cat([pad, x] if pad_first else [x, pad])
if backwards: x1 = x1.flip(0)
return retain_type(x1, x)
return [tuple(map(lambda idxx: _f(*idxx), enumerate(s))) for s in samples]
def decodes(self, o:TensorText):
pad_idx = self.pad_idx if hasattr(self,'pad_idx') else 1
return o[o != pad_idx]
pad_input=Pad_Input()
# `pad_idx` is used for the padding, and the padding is applied to the `pad_fields` of the samples. The padding is applied at the beginning if `pad_first` is `True`, and if `backwards` is added, the tensors are flipped.
test_eq(pad_input([(tensor([1,2,3]),1), (tensor([4,5]), 2), (tensor([6]), 3)], pad_idx=0),
[(tensor([1,2,3]),1), (tensor([4,5,0]),2), (tensor([6,0,0]), 3)])
test_eq(pad_input([(tensor([1,2,3]), (tensor([6]))), (tensor([4,5]), tensor([4,5])), (tensor([6]), (tensor([1,2,3])))], pad_idx=0, pad_fields=1),
[(tensor([1,2,3]),(tensor([6,0,0]))), (tensor([4,5]),tensor([4,5,0])), ((tensor([6]),tensor([1, 2, 3])))])
test_eq(pad_input([(tensor([1,2,3]),1), (tensor([4,5]), 2), (tensor([6]), 3)], pad_idx=0, pad_first=True),
[(tensor([1,2,3]),1), (tensor([0,4,5]),2), (tensor([0,0,6]), 3)])
test_eq(pad_input([(tensor([1,2,3]),1), (tensor([4,5]), 2), (tensor([6]), 3)], pad_idx=0, backwards=True),
[(tensor([3,2,1]),1), (tensor([5,4,0]),2), (tensor([6,0,0]), 3)])
x = pad_input([(TensorText([1,2,3]),1), (TensorText([4,5]), 2), (TensorText([6]), 3)], pad_idx=0)
test_eq(x, [(tensor([1,2,3]),1), (tensor([4,5,0]), 2), (tensor([6,0,0]), 3)])
test_eq(pad_input.decode(x[1][0]), tensor([4,5]))
#hide
#Check retain type
x = [(TensorText([1,2,3]),1), (TensorText([4,5]), 2), (TensorText([6]), 3)]
y = pad_input(x, pad_idx=0)
for s in y: test_eq(type(s[0]), TensorText)
# Pads `x` with `pad_idx` to length `pad_len`. If `pad_first` is false, all padding is appended to `x`, until `x` is len `pad_len`. Otherwise ff `pad_first` is true, then chunks of size `seq_len` are prepended to `x`, the remainder of the padding is appended to `x`.
#export
def pad_chunk(x,pad_idx=1, pad_first=True, seq_len=72, pad_len=10):
"Pad `x` by adding padding by chunks of size `seq_len`"
l = pad_len - x.shape[0]
pad_chunk = x.new_zeros((l//seq_len) * seq_len) + pad_idx
pad_res = x.new_zeros(l % seq_len) + pad_idx
x1 = torch.cat([pad_chunk, x, pad_res]) if pad_first else torch.cat([x, pad_chunk, pad_res])
return retain_type(x1, x)
print('pad_first: ',pad_chunk(torch.tensor([1,2,3]),seq_len=3,pad_idx=0,pad_len=8))
print('pad_last: ',pad_chunk(torch.tensor([1,2,3]),seq_len=3,pad_idx=0,pad_len=8,pad_first=False))
# `pad_input_chunk` is the version of `pad_chunk` that works over a list of lists.
#export
@delegates(pad_chunk)
def pad_input_chunk(samples, n_inp=1,**kwargs):
"Pad `samples` by adding padding by chunks of size `seq_len`"
max_len = max([len(s[n]) for s in samples for n in range(n_inp)])
padeds = [[pad_chunk(s[n],pad_len=max_len,**kwargs) for n in range(n_inp) ] for s in samples]
return [(*p, *s[n_inp:]) for p,s in zip(padeds,samples)]
# The difference with the base `pad_input` is that most of the padding is applied first (if `pad_first=True`) or at the end (if `pad_first=False`) but only by a round multiple of `seq_len`. The rest of the padding is applied to the end (or the beginning if `pad_first=False`). This is to work with `SequenceEncoder` with recurrent models.
pad_input_chunk([(TensorText([1,2,3,4,5,6]),TensorText([1,2]),1)], pad_idx=0, seq_len=3,n_inp=2)
# +
test_eq(pad_input_chunk([(tensor([1,2,3,4,5,6]),1), (tensor([1,2,3]), 2), (tensor([1,2]), 3)], pad_idx=0, seq_len=2),
[(tensor([1,2,3,4,5,6]),1), (tensor([0,0,1,2,3,0]),2), (tensor([0,0,0,0,1,2]), 3)])
test_eq(pad_input_chunk([(tensor([1,2,3,4,5,6]),), (tensor([1,2,3]),), (tensor([1,2]),)], pad_idx=0, seq_len=2),
[(tensor([1,2,3,4,5,6]),), (tensor([0,0,1,2,3,0]),), (tensor([0,0,0,0,1,2]),)])
test_eq(pad_input_chunk([(tensor([1,2,3,4,5,6]),), (tensor([1,2,3]),), (tensor([1,2]),)], pad_idx=0, seq_len=2, pad_first=False),
[(tensor([1,2,3,4,5,6]),), (tensor([1,2,3,0,0,0]),), (tensor([1,2,0,0,0,0]),)])
test_eq(pad_input_chunk([(TensorText([1,2,3,4,5,6]),TensorText([1,2]),1)], pad_idx=0, seq_len=2,n_inp=2),
[(TensorText([1,2,3,4,5,6]),TensorText([0,0,0,0,1,2]),1)])
# -
# `Transform` version of `pad_input_chunk`. This version supports types, decoding, and the other functionality of `Transform`
#export
class Pad_Chunk(DisplayedTransform):
"Pad `samples` by adding padding by chunks of size `seq_len`"
def __init__(self, pad_idx=1, pad_first=True, seq_len=72,decode=True,**kwargs):
store_attr('pad_idx, pad_first, seq_len,seq_len')
super().__init__(**kwargs)
def before_call(self, b):
"Set `self.max_len` before encodes"
self.max_len = max([x.shape[0] for xs in b for x in xs if isinstance(x,TensorText)])
def __call__(self, b, **kwargs):
self.before_call(b)
return super().__call__(tuple(b), **kwargs)
def encodes(self, x:TensorText):
return pad_chunk(x,pad_idx=self.pad_idx, pad_first=self.pad_first, seq_len=self.seq_len, pad_len=self.max_len)
def decodes(self, o:TensorText):
return o[o != self.pad_idx] if self.decode else o
# Here is an example of `Pad_Chunk`
pc=Pad_Chunk(pad_idx=0,seq_len=3)
out=pc([(TensorText([1,2,3,4,5,6]),TensorText([1,2]),1)])
print('Inputs: ',*[(TensorText([1,2,3,4,5,6]),TensorText([1,2]),1)])
print('Encoded: ',*out)
print('Decoded: ',*pc.decode(out))
# +
pc=Pad_Chunk(pad_idx=0, seq_len=2)
test_eq(pc([(TensorText([1,2,3,4,5,6]),1), (TensorText([1,2,3]), 2), (TensorText([1,2]), 3)]),
[(tensor([1,2,3,4,5,6]),1), (tensor([0,0,1,2,3,0]),2), (tensor([0,0,0,0,1,2]), 3)])
pc=Pad_Chunk(pad_idx=0, seq_len=2)
test_eq(pc([(TensorText([1,2,3,4,5,6]),), (TensorText([1,2,3]),), (TensorText([1,2]),)]),
[(tensor([1,2,3,4,5,6]),), (tensor([0,0,1,2,3,0]),), (tensor([0,0,0,0,1,2]),)])
pc=Pad_Chunk(pad_idx=0, seq_len=2, pad_first=False)
test_eq(pc([(TensorText([1,2,3,4,5,6]),), (TensorText([1,2,3]),), (TensorText([1,2]),)]),
[(tensor([1,2,3,4,5,6]),), (tensor([1,2,3,0,0,0]),), (tensor([1,2,0,0,0,0]),)])
pc=Pad_Chunk(pad_idx=0, seq_len=2)
test_eq(pc([(TensorText([1,2,3,4,5,6]),TensorText([1,2]),1)]),
[(TensorText([1,2,3,4,5,6]),TensorText([0,0,0,0,1,2]),1)])
# +
#export
def _default_sort(x): return len(x[0])
@delegates(TfmdDL)
class SortedDL(TfmdDL):
"A `DataLoader` that goes throught the item in the order given by `sort_func`"
def __init__(self, dataset, sort_func=None, res=None, **kwargs):
super().__init__(dataset, **kwargs)
self.sort_func = _default_sort if sort_func is None else sort_func
if res is None and self.sort_func == _default_sort: res = _get_lengths(dataset)
self.res = [self.sort_func(self.do_item(i)) for i in range_of(self.dataset)] if res is None else res
if len(self.res) > 0: self.idx_max = np.argmax(self.res)
def get_idxs(self):
idxs = super().get_idxs()
if self.shuffle: return idxs
return sorted(idxs, key=lambda i: self.res[i], reverse=True)
def shuffle_fn(self,idxs):
idxs = np.random.permutation(len(self.dataset))
idx_max = np.where(idxs==self.idx_max)[0][0]
idxs[0],idxs[idx_max] = idxs[idx_max],idxs[0]
sz = self.bs*50
chunks = [idxs[i:i+sz] for i in range(0, len(idxs), sz)]
chunks = [sorted(s, key=lambda i: self.res[i], reverse=True) for s in chunks]
sort_idx = np.concatenate(chunks)
sz = self.bs
batches = [sort_idx[i:i+sz] for i in range(0, len(sort_idx), sz)]
sort_idx = np.concatenate(np.random.permutation(batches[1:-1])) if len(batches) > 2 else np.array([],dtype=np.int)
sort_idx = np.concatenate((batches[0], sort_idx) if len(batches)==1 else (batches[0], sort_idx, batches[-1]))
return iter(sort_idx)
@delegates(TfmdDL.new)
def new(self, dataset=None, **kwargs):
if 'val_res' in kwargs and kwargs['val_res'] is not None: res = kwargs['val_res']
else: res = self.res if dataset is None else None
return super().new(dataset=dataset, res=res, **kwargs)
# -
# `res` is the result of `sort_func` applied on all elements of the `dataset`. You can pass it if available to make the init much faster by avoiding an initial pass over the whole dataset. For example if sorting by text length (as in the default `sort_func`, called `_default_sort`) you should pass a list with the length of each element in `dataset` to `res` to take advantage of this speed-up.
#
# To get the same init speed-up for the validation set, `val_res` (a list of text lengths for your validation set) can be passed to the `kwargs` argument of `SortedDL`. Below is an example to reduce the init time by passing a list of text lengths for both the training set and the validation set:
#
# ```
# # Pass the training dataset text lengths to SortedDL
# srtd_dl=partial(SortedDL, res = train_text_lens)
#
# # Pass the validation dataset text lengths
# dl_kwargs = [{},{'val_res': val_text_lens}]
#
# # init our Datasets
# dsets = Datasets(...)
#
# # init our Dataloaders
# dls = dsets.dataloaders(...,dl_type = srtd_dl, dl_kwargs = dl_kwargs)
# ```
#
# If `shuffle` is `True`, this will shuffle a bit the results of the sort to have items of roughly the same size in batches, but not in the exact sorted order.
ds = [(tensor([1,2]),1), (tensor([3,4,5,6]),2), (tensor([7]),3), (tensor([8,9,10]),4)]
dl = SortedDL(ds, bs=2, before_batch=partial(pad_input, pad_idx=0))
test_eq(list(dl), [(tensor([[ 3, 4, 5, 6], [ 8, 9, 10, 0]]), tensor([2, 4])),
(tensor([[1, 2], [7, 0]]), tensor([1, 3]))])
ds = [(tensor(range(random.randint(1,10))),i) for i in range(101)]
dl = SortedDL(ds, bs=2, create_batch=partial(pad_input, pad_idx=-1), shuffle=True, num_workers=0)
batches = list(dl)
max_len = len(batches[0][0])
for b in batches:
assert(len(b[0])) <= max_len
test_ne(b[0][-1], -1)
# ## TransformBlock for text
# To use the data block API, you will need this build block for texts.
#export
class TextBlock(TransformBlock):
"A `TransformBlock` for texts"
@delegates(Numericalize.__init__)
def __init__(self, tok_tfm, vocab=None, is_lm=False, seq_len=72, backwards=False, **kwargs):
type_tfms = [tok_tfm, Numericalize(vocab, **kwargs)]
if backwards: type_tfms += [reverse_text]
return super().__init__(type_tfms=type_tfms,
dl_type=LMDataLoader if is_lm else SortedDL,
dls_kwargs={'seq_len': seq_len} if is_lm else {'before_batch': Pad_Chunk(seq_len=seq_len)})
@classmethod
@delegates(Tokenizer.from_df, keep=True)
def from_df(cls, text_cols, vocab=None, is_lm=False, seq_len=72, backwards=False, min_freq=3, max_vocab=60000, **kwargs):
"Build a `TextBlock` from a dataframe using `text_cols`"
return cls(Tokenizer.from_df(text_cols, **kwargs), vocab=vocab, is_lm=is_lm, seq_len=seq_len,
backwards=backwards, min_freq=min_freq, max_vocab=max_vocab)
@classmethod
@delegates(Tokenizer.from_folder, keep=True)
def from_folder(cls, path, vocab=None, is_lm=False, seq_len=72, backwards=False, min_freq=3, max_vocab=60000, **kwargs):
"Build a `TextBlock` from a `path`"
return cls(Tokenizer.from_folder(path, **kwargs), vocab=vocab, is_lm=is_lm, seq_len=seq_len,
backwards=backwards, min_freq=min_freq, max_vocab=max_vocab)
# For efficient tokenization, you probably want to use one of the factory methods. Otherwise, you can pass your custom `tok_tfm` that will deal with tokenization (if your texts are already tokenized, you can pass `noop`), a `vocab`, or leave it to be inferred on the texts using `min_freq` and `max_vocab`.
#
# `is_lm` indicates if we want to use texts for language modeling or another task, `seq_len` is only necessary to tune if `is_lm=False`, and is passed along to `pad_input_chunk`.
show_doc(TextBlock.from_df)
# Here is an example using a sample of IMDB stored as a CSV file:
# +
path = untar_data(URLs.IMDB_SAMPLE)
df = pd.read_csv(path/'texts.csv')
imdb_clas = DataBlock(
blocks=(TextBlock.from_df('text', seq_len=72), CategoryBlock),
get_x=ColReader('text'), get_y=ColReader('label'), splitter=ColSplitter())
dls = imdb_clas.dataloaders(df, bs=64)
dls.show_batch(max_n=2)
# -
# `vocab`, `is_lm`, `seq_len`, `min_freq` and `max_vocab` are passed to the main init, the other argument to `Tokenizer.from_df`.
show_doc(TextBlock.from_folder)
# `vocab`, `is_lm`, `seq_len`, `min_freq` and `max_vocab` are passed to the main init, the other argument to `Tokenizer.from_folder`.
# ## TextDataLoaders -
# +
#export
class TextDataLoaders(DataLoaders):
"Basic wrapper around several `DataLoader`s with factory methods for NLP problems"
@classmethod
@delegates(DataLoaders.from_dblock)
def from_folder(cls, path, train='train', valid='valid', valid_pct=None, seed=None, vocab=None, text_vocab=None, is_lm=False,
tok_tfm=None, seq_len=72, backwards=False, **kwargs):
"Create from imagenet style dataset in `path` with `train` and `valid` subfolders (or provide `valid_pct`)"
splitter = GrandparentSplitter(train_name=train, valid_name=valid) if valid_pct is None else RandomSplitter(valid_pct, seed=seed)
blocks = [TextBlock.from_folder(path, text_vocab, is_lm, seq_len, backwards) if tok_tfm is None else TextBlock(tok_tfm, text_vocab, is_lm, seq_len, backwards)]
if not is_lm: blocks.append(CategoryBlock(vocab=vocab))
get_items = partial(get_text_files, folders=[train,valid]) if valid_pct is None else get_text_files
dblock = DataBlock(blocks=blocks,
get_items=get_items,
splitter=splitter,
get_y=None if is_lm else parent_label)
return cls.from_dblock(dblock, path, path=path, seq_len=seq_len, **kwargs)
@classmethod
@delegates(DataLoaders.from_dblock)
def from_df(cls, df, path='.', valid_pct=0.2, seed=None, text_col=0, label_col=1, label_delim=None, y_block=None,
text_vocab=None, is_lm=False, valid_col=None, tok_tfm=None, seq_len=72, backwards=False, **kwargs):
"Create from `df` in `path` with `valid_pct`"
blocks = [TextBlock.from_df(text_col, text_vocab, is_lm, seq_len, backwards) if tok_tfm is None else TextBlock(tok_tfm, text_vocab, is_lm, seq_len, backwards)]
if y_block is None and not is_lm:
blocks.append(MultiCategoryBlock if is_listy(label_col) and len(label_col) > 1 else CategoryBlock)
if y_block is not None and not is_lm: blocks += (y_block if is_listy(y_block) else [y_block])
splitter = RandomSplitter(valid_pct, seed=seed) if valid_col is None else ColSplitter(valid_col)
dblock = DataBlock(blocks=blocks,
get_x=ColReader("text"),
get_y=None if is_lm else ColReader(label_col, label_delim=label_delim),
splitter=splitter)
return cls.from_dblock(dblock, df, path=path, seq_len=seq_len, **kwargs)
@classmethod
def from_csv(cls, path, csv_fname='labels.csv', header='infer', delimiter=None, **kwargs):
"Create from `csv` file in `path/csv_fname`"
df = pd.read_csv(Path(path)/csv_fname, header=header, delimiter=delimiter)
return cls.from_df(df, path=path, **kwargs)
TextDataLoaders.from_csv = delegates(to=TextDataLoaders.from_df)(TextDataLoaders.from_csv)
# -
show_doc(TextDataLoaders, title_level=2)
# You should not use the init directly but one of the following factory methods. All those factory methods accept as arguments:
#
# - `text_vocab`: the vocabulary used for numericalizing texts (if not passed, it's inferred from the data)
# - `tok_tfm`: if passed, uses this `tok_tfm` instead of the default
# - `seq_len`: the sequence length used for batch
# - `bs`: the batch size
# - `val_bs`: the batch size for the validation `DataLoader` (defaults to `bs`)
# - `shuffle_train`: if we shuffle the training `DataLoader` or not
# - `device`: the PyTorch device to use (defaults to `default_device()`)
show_doc(TextDataLoaders.from_folder)
# If `valid_pct` is provided, a random split is performed (with an optional `seed`) by setting aside that percentage of the data for the validation set (instead of looking at the grandparents folder). If a `vocab` is passed, only the folders with names in `vocab` are kept.
#
# Here is an example on a sample of the IMDB movie review dataset:
#slow
path = untar_data(URLs.IMDB)
dls = TextDataLoaders.from_folder(path)
dls.show_batch(max_n=3)
show_doc(TextDataLoaders.from_df)
# `seed` can optionally be passed for reproducibility. `text_col`, `label_col` and optionally `valid_col` are indices or names of columns for texts/labels and the validation flag. `label_delim` can be passed for a multi-label problem if your labels are in one column, separated by a particular char. `y_block` should be passed to indicate your type of targets, in case the library did no infer it properly.
#
# Here are examples on subsets of IMDB:
path = untar_data(URLs.IMDB_SAMPLE)
dls = TextDataLoaders.from_df(df, path=path, text_col='text', label_col='label', valid_col='is_valid')
dls.show_batch(max_n=3)
dls = TextDataLoaders.from_df(df, path=path, text_col='text', is_lm=True, valid_col='is_valid')
dls.show_batch(max_n=3)
show_doc(TextDataLoaders.from_csv)
# Opens the csv file with `header` and `delimiter`, then pass all the other arguments to `TextDataLoaders.from_df`.
dls = TextDataLoaders.from_csv(path=path, csv_fname='texts.csv', text_col='text', label_col='label', valid_col='is_valid')
dls.show_batch(max_n=3)
# ## Export -
#hide
from nbdev.export import notebook2script
notebook2script()
|
nbs/31_text.data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import json
import random
from deap import creator, base, tools, algorithms
import numpy as np
# +
creator.create("FitnessMax", base.Fitness, weights=(1.0,))
creator.create("Individual", np.ndarray, fitness=creator.FitnessMax)
toolbox = base.Toolbox()
toolbox.register("attr_bool", random.randint, 0, 1)
toolbox.register("individual", tools.initRepeat, creator.Individual, toolbox.attr_bool, n=100)
toolbox.register("population", tools.initRepeat, np.ndarray, toolbox.individual)
def evalOneMax(individual):
return sum(individual),
toolbox.register("evaluate", evalOneMax)
toolbox.register("mate", tools.cxTwoPoint)
toolbox.register("mutate", tools.mutFlipBit, indpb=0.05)
toolbox.register("select", tools.selTournament, tournsize=3)
population = toolbox.population(n=300)
NGEN=40
for gen in range(NGEN):
offspring = algorithms.varAnd(population, toolbox, cxpb=0.5, mutpb=0.1)
fits = toolbox.map(toolbox.evaluate, offspring)
for fit, ind in zip(fits, offspring):
ind.fitness.values = fit
population = toolbox.select(offspring, k=len(population))
top10 = tools.selBest(population, k=10)
# -
print(top10[0])
creator.create("FitnessMax", base.Fitness, weights=(1.0,))
creator.create("Individual", np.ndarray, fitness=creator.FitnessMax)
def generate_individual():
BT = np.array([
[ 0., 0., 0., 0., 0.],
[ 1/2., 1/2., 0., 0., 0.],
[ 1/2., 0., 1/2., 0., 0.],
[ 1., 0., 0., 1., 0.],
[ 0., 1/6., 1/3., 1/3., 1/6.]
], dtype=np.float32)*random.random()
return BT
toolbox = base.Toolbox()
toolbox.register("")
np.flo
creator.create("indi", lis, fitness=C)
|
notebooks/DEAP Playground.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_mxnet_p36
# language: python
# name: conda_mxnet_p36
# ---
# # Sentiment Classification for Movie Review Dataset (Korean)
#
# 본 핸즈온에서는 네이버 영화 리뷰에 대한 감정(0: 부정, 1: 긍정)을 요약한 네이버 영화 리뷰 데이터셋으로 AutoGluon 훈련을 수행합니다.
# +
# GPU 인스턴스를 사용하시면 아래 주석을 해제하고 본 코드 셀을 실행해 주세요.
# # !pip install --upgrade mxnet-cu100
# +
import os
import mxnet as mx
num_gpus = mx.context.num_gpus()
if num_gpus == 0:
os.environ['AUTOGLUON_TEXT_TRAIN_WITHOUT_GPU'] = '1'
# -
import numpy as np
import warnings
import matplotlib.pyplot as plt
warnings.filterwarnings('ignore')
np.random.seed(123)
# <br>
#
# ## 1. Data preparation and Training
#
# https://github.com/e9t/nsmc/ 에 공개된 네이버 영화 리뷰 데이터셋을 다운로드합니다.
# 훈련 데이터는 총 15만건이며, 테스트 데이터는 총 5만건입니다.
save_path = 'ag-02-sentiment-classifcation-kor'
# !rm -rf $save_path input
# !wget -nc https://raw.githubusercontent.com/e9t/nsmc/master/ratings_train.txt -P ./input/
# !wget -nc https://raw.githubusercontent.com/e9t/nsmc/master/ratings_test.txt -P ./input/
# +
import pandas as pd
import numpy as np
train_df = pd.read_csv('./input/ratings_train.txt', header=0, delimiter='\t')
test_df = pd.read_csv('./input/ratings_test.txt', header=0, delimiter='\t')
train_df = train_df[['document', 'label']]
test_df = test_df[['document', 'label']]
# +
from autogluon.tabular import TabularDataset, TabularPredictor
train_data = TabularDataset(train_df)
test_data = TabularDataset(test_df)
subsample_size = 1000 # subsample data for faster demo, try setting this to larger values
train_data = train_data.sample(n=subsample_size, random_state=0)
test_data = test_data.sample(n=subsample_size, random_state=0)
train_data.head(5)
# -
# 간단한 전처리를 수행합니다. 특수 문자와 한글 외 문자들을 제거하고 중복값 및 결측값을 제외합니다.
def basic_preprocess(data):
data.drop_duplicates(subset = ['document'], inplace=True)
data['document'] = data['document'].str.replace("[^ㄱ-ㅎㅏ-ㅣ가-힣 ]","")
#data['document'] = data['document'].str.replace("[\,\(\)\{\}\[\]\`\'\!\?\:\;\-\=]", " ") # 특수문자 제거만 사용
data = data.dropna(how='any')
return data
train_data = basic_preprocess(train_data)
test_data = basic_preprocess(test_data)
# 훈련 지정 시 좀 더 세부적인 하이퍼파라메터 설정이 필요하다면, 사전 정의된 preset을 사용하시면 편리합니다. TextPredictor는 사전 훈련된 BERT, RoBERT, ELECTRA가 내장되어 있으며, 한국어를 비롯한 다국어에 대한 훈련이 필요하면 `multi_cased_bert_base_fuse_late` preset을 사용하시면 됩니다.
from autogluon.text import ag_text_presets, list_presets
list_presets()
# +
from autogluon.text import TextPredictor
predictor = TextPredictor(label='label', eval_metric='acc', path=save_path)
predictor.set_verbosity(2)
predictor.fit(train_data, presets='multi_cased_bert_base_fuse_late', time_limit=120)
# -
# <br>
#
# ## 2. Evaluation and Prediction
# ### Evaluation
#
# `predictor.evaluation()`를 사용하여 평가를 쉽게 수행할 수 있으며, F1 score 등의 추가 metric도 지정 가능합니다.
if num_gpus > 0:
test_score = predictor.evaluate(test_data, metrics=['acc', 'f1'])
print(test_score)
# ### Prediction
# `predictor.predict()`으로 예측을 수행할 수 있습니다.
sentence1 = "이 영화 너무너무 재미있어요. 인생 최고의 영화입니다. 최고!"
sentence2 = "평점 1점도 아깝습니다..비추"
predictions = predictor.predict({'document': [sentence1, sentence2]})
print('"Sentence":', sentence1, '"Predicted Sentiment":', predictions[0])
print('"Sentence":', sentence2, '"Predicted Sentiment":', predictions[1])
# ### Extract Embeddings
# 훈련된 predictor를 사용하여 임베딩 벡터에 매핑하는 임베딩을 추출할 수도 있습니다.
embeddings = predictor.extract_embedding(test_data)
print(embeddings)
# TSNE를 사용하여 추출된 임베딩을 시각화합니다. 성능 향상을 위해서는 다국어 모델보다는 한국어 전용 모델로 훈련하는 것이 더 유리합니다.
if num_gpus > 0:
from sklearn.manifold import TSNE
embeddings = predictor.extract_embedding(test_data)
print(embeddings)
X_embedded = TSNE(n_components=2, random_state=123).fit_transform(embeddings)
for val, color in [(0, 'red'), (1, 'blue')]:
idx = (test_data['label'].to_numpy() == val).nonzero()
plt.scatter(X_embedded[idx, 0], X_embedded[idx, 1], c=color, label=f'label={val}')
plt.legend(loc='best')
|
2.nlp/02_sentiment_classification_navermovie_kor.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.10.0 64-bit
# name: python3
# ---
# # Accounts Merge
# Given a list of accounts where each element accounts[i] is a list of strings, where the first element accounts[i][0] is a name, and the rest of the elements are emails representing emails of the account.
# Now, we would like to merge these accounts. Two accounts definitely belong to the same person if there is some common email to both accounts. Note that even if two accounts have the same name, they may belong to different people as people could have the same name. A person can have any number of accounts initially, but all of their accounts definitely have the same name. After merging the accounts, return the accounts in the following format: the first element of each account is the name, and the rest of the elements are emails in sorted order. The accounts themselves can be returned in any order.
#
# ### Example 1:
# - Input: accounts = [["John","<EMAIL>","<EMAIL>"],["John","<EMAIL>","<EMAIL>"],["Mary","<EMAIL>"],["John","<EMAIL>"]]
# - Output: [["John","<EMAIL>","<EMAIL>","<EMAIL>"],["Mary","<EMAIL>"],["John","<EMAIL>"]]
# - Explanation: The first and second John's are the same person as they have the common email "<EMAIL>".
# The third John and Mary are different people as none of their email addresses are used by other accounts.
# We could return these lists in any order, for example the answer [['Mary', '<EMAIL>'], ['John', '<EMAIL>'], ['John', '<EMAIL>', '<EMAIL>', '<EMAIL>']] would still be accepted.
# +
def acc_merge(accounts: list) -> list:
res = {}
for a in accounts:
if not a[0] in res:
res[a[0]] = [set(a[1:])]
else:
curr = 0
lst = res[a[0]]
tmp = set(a[1:])
while curr < len(lst):
if lst[curr].intersection(tmp):
tmp = lst[curr].union(tmp)
lst.pop(curr)
else:
curr += 1
lst.append(tmp)
return [[k] + sorted(list(y)) for k,v in res.items() for y in v]
acc_merge([
["John","<EMAIL>","<EMAIL>"],
["John","<EMAIL>","<EMAIL>"],
["Mary","<EMAIL>"],
["John","<EMAIL>"]])
|
python-data-structures/interview-fb/accounts-merge.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + dc={"key": "4"} deletable=false editable=false run_control={"frozen": true} tags=["context"] id="vPaFUDv-ZFkm" colab_type="text"
# ## 1. Google Play Store apps and reviews
# <p>Mobile apps are everywhere. They are easy to create and can be lucrative. Because of these two factors, more and more apps are being developed. In this notebook, we will do a comprehensive analysis of the Android app market by comparing over ten thousand apps in Google Play across different categories. We'll look for insights in the data to devise strategies to drive growth and retention.</p>
# <p><img src="https://play.google.com/intl/en_us/badges/static/images/badges/en_badge_web_generic.png" alt="Google Play logo"></p>
# <p>Let's take a look at the data, which consists of two files:</p>
# <ul>
# <li><code>apps.csv</code>: contains all the details of the applications on Google Play. There are 13 features that describe a given app.</li>
# <li><code>user_reviews.csv</code>: contains 100 reviews for each app, <a href="https://www.androidpolice.com/2019/01/21/google-play-stores-redesigned-ratings-and-reviews-section-lets-you-easily-filter-by-star-rating/">most helpful first</a>. The text in each review has been pre-processed and attributed with three new features: Sentiment (Positive, Negative or Neutral), Sentiment Polarity and Sentiment Subjectivity.</li>
# </ul>
# + id="i3YfE801eaNi" colab_type="code" outputId="93202fa7-ffc8-47be-ff15-fd4881e8cb3b" colab={"base_uri": "https://localhost:8080/", "height": 217}
# !pip install chart_studio
# !pip install plotly==4.7.1
# + id="o4fJ1453caOt" colab_type="code" colab={}
import chart_studio.plotly as py
import numpy as np
# + dc={"key": "4"} tags=["sample_code"] id="moAzvTZFZFkn" colab_type="code" outputId="cfc0db2b-570b-44a0-8680-0600de0ad7c5" colab={"base_uri": "https://localhost:8080/", "height": 330}
# Read in dataset
import pandas as pd
apps_with_duplicates = pd.read_csv("https://raw.githubusercontent.com/Sankalp679/SHALA/Assignments/DataCamp_Projects/The-Android-App-Market-on-Google-Play/Datasets/apps.csv")
# # Drop duplicates
apps =apps_with_duplicates. drop_duplicates()
# Print the total number of apps
print('Total number of apps in the dataset = ', apps.shape[0])
# # Have a look at a random sample of 5 rows
n = 5
apps.sample(n)
# + dc={"key": "11"} deletable=false editable=false run_control={"frozen": true} tags=["context"] id="blbe7dSRZFks" colab_type="text"
# ## 2. Data cleaning
# <p>The three features that we will be working with most frequently henceforth are <code>Installs</code>, <code>Size</code>, and <code>Price</code>. A careful glance of the dataset reveals that some of these columns mandate data cleaning in order to be consumed by code we'll write later. Specifically, the presence of special characters (<code>, $ +</code>) and letters (<code>M k</code>) in the <code>Installs</code>, <code>Size</code>, and <code>Price</code> columns make their conversion to a numerical data type difficult. Let's clean by removing these and converting each column to a numeric type.</p>
# + dc={"key": "11"} tags=["sample_code"] id="54T87HvyZFks" colab_type="code" colab={}
# List of characters to remove
chars_to_remove = [',','$','+','M','K']
# List of column names to clean
cols_to_clean = ['Installs','Size', 'Price']
# Loop for each column
for col in cols_to_clean:
# Replace each character with an empty string
for char in chars_to_remove:
apps[col] = apps[col].str.replace(char, '')
# Convert col to numeric
apps[col] = pd.to_numeric(apps[col])
# + dc={"key": "18"} deletable=false editable=false run_control={"frozen": true} tags=["context"] id="LMfHoVy8ZFkv" colab_type="text"
# ## 3. Exploring app categories
# <p>With more than 1 billion active users in 190 countries around the world, Google Play continues to be an important distribution platform to build a global audience. For businesses to get their apps in front of users, it's important to make them more quickly and easily discoverable on Google Play. To improve the overall search experience, Google has introduced the concept of grouping apps into categories.</p>
# <p>This brings us to the following questions:</p>
# <ul>
# <li>Which category has the highest share of (active) apps in the market? </li>
# <li>Is any specific category dominating the market?</li>
# <li>Which categories have the fewest number of apps?</li>
# </ul>
# <p>We will see that there are <code>33</code> unique app categories present in our dataset. <em>Family</em> and <em>Game</em> apps have the highest market prevalence. Interestingly, <em>Tools</em>, <em>Business</em> and <em>Medical</em> apps are also at the top.</p>
# + dc={"key": "18"} tags=["sample_code"] id="xtPlq3-eZFkv" colab_type="code" outputId="c801fd9a-42a0-4005-f28d-08a64cc17329" colab={"base_uri": "https://localhost:8080/", "height": 558}
import plotly.graph_objects as go
# Print the total number of unique categories
num_categories = len(apps['Category'].unique())
print('Number of categories = ', num_categories)
# Count the number of apps in each 'Category' and sort them in descending order
num_apps_in_category = apps['Category'].value_counts().sort_values(ascending = False)
fig = go.Figure(
data = [go.Bar(
x = num_apps_in_category.index, # index = category name
y = num_apps_in_category.values, # value = count
)]
)
fig.show()
# + dc={"key": "25"} deletable=false editable=false run_control={"frozen": true} tags=["context"] id="VvvnOiT3ZFky" colab_type="text"
# ## 4. Distribution of app ratings
# <p>After having witnessed the market share for each category of apps, let's see how all these apps perform on an average. App ratings (on a scale of 1 to 5) impact the discoverability, conversion of apps as well as the company's overall brand image. Ratings are a key performance indicator of an app.</p>
# <p>From our research, we found that the average volume of ratings across all app categories is <code>4.17</code>. The histogram plot is skewed to the right indicating that the majority of the apps are highly rated with only a few exceptions in the low-rated apps.</p>
# + dc={"key": "25"} tags=["sample_code"] id="9zy_wxhWZFkz" colab_type="code" outputId="41c0533b-dc06-4735-a87e-b6c9099afe42" colab={"base_uri": "https://localhost:8080/", "height": 558}
# Average rating of apps
avg_app_rating = apps['Rating'].mean()
print('Average app rating = ', avg_app_rating)
fig = go.Figure(
# Distribution of apps according to their ratings
data = [go.Histogram(
x = apps['Rating'])],
# Vertical dashed line to indicate the average app rating
layout = {'shapes': [{
'type' :'line',
'x0': avg_app_rating,
'y0': 0,
'x1': avg_app_rating,
'y1': 1000,
'line': { 'dash': 'dashdot'}}]})
fig.show()
# + dc={"key": "32"} deletable=false editable=false run_control={"frozen": true} tags=["context"] id="1H_HjtI6ZFk1" colab_type="text"
# ## 5. Size and price of an app
# <p>Let's now examine app size and app price. For size, if the mobile app is too large, it may be difficult and/or expensive for users to download. Lengthy download times could turn users off before they even experience your mobile app. Plus, each user's device has a finite amount of disk space. For price, some users expect their apps to be free or inexpensive. These problems compound if the developing world is part of your target market; especially due to internet speeds, earning power and exchange rates.</p>
# <p>How can we effectively come up with strategies to size and price our app?</p>
# <ul>
# <li>Does the size of an app affect its rating? </li>
# <li>Do users really care about system-heavy apps or do they prefer light-weighted apps? </li>
# <li>Does the price of an app affect its rating? </li>
# <li>Do users always prefer free apps over paid apps?</li>
# </ul>
# <p>We find that the majority of top rated apps (rating over 4) range from 2 MB to 20 MB. We also find that the vast majority of apps price themselves under \$10.</p>
# + dc={"key": "32"} tags=["sample_code"] id="3dgaIH0DZFk2" colab_type="code" outputId="56175c98-fd41-454a-d065-1d3cc52dbe44" colab={"base_uri": "https://localhost:8080/", "height": 865}
# %matplotlib inline
import seaborn as sns
sns.set_style("darkgrid")
import warnings
warnings.filterwarnings("ignore")
# Plot size vs. rating
plt1 = sns.jointplot(x = apps['Size'], y = apps['Rating'], kind = 'hex')
# Subset out apps whose type is 'Paid'
paid_apps = apps[apps['Type'] == 'Paid']
# Plot price vs. rating
plt2 = sns.jointplot(x = paid_apps['Price'], y = paid_apps['Rating'])
# + dc={"key": "39"} deletable=false editable=false run_control={"frozen": true} tags=["context"] id="6GtWn4fQZFk4" colab_type="text"
# ## 6. Relation between app category and app price
# <p>So now comes the hard part. How are companies and developers supposed to make ends meet? What monetization strategies can companies use to maximize profit? The costs of apps are largely based on features, complexity, and platform.</p>
# <p>There are many factors to consider when selecting the right pricing strategy for your mobile app. It is important to consider the willingness of your customer to pay for your app. A wrong price could break the deal before the download even happens. Potential customers could be turned off by what they perceive to be a shocking cost, or they might delete an app they’ve downloaded after receiving too many ads or simply not getting their money's worth.</p>
# <p>Different categories demand different price ranges. Some apps that are simple and used daily, like the calculator app, should probably be kept free. However, it would make sense to charge for a highly-specialized medical app that diagnoses diabetic patients. Below, we see that <em>Medical and Family</em> apps are the most expensive. Some medical apps extend even up to $80! All game apps are reasonably priced below \$20.</p>
# + dc={"key": "39"} tags=["sample_code"] id="ux7zb6kqZFk5" colab_type="code" outputId="0b470248-151b-4698-966c-9e55ae7a8be7" colab={"base_uri": "https://localhost:8080/", "height": 1000}
import matplotlib.pyplot as plt
fig, ax = plt.subplots()
fig.set_size_inches(15, 8)
# Select a few popular app categories
popular_app_cats = apps[apps.Category.isin(['GAME', 'FAMILY', 'PHOTOGRAPHY',
'MEDICAL', 'TOOLS', 'FINANCE',
'LIFESTYLE','BUSINESS'])]
# # Examine the price trend by plotting Price vs Category
ax = sns.stripplot(x = popular_app_cats['Price'], y = popular_app_cats['Category'], jitter=True, linewidth=1)
ax.set_title('App pricing trend across categories')
# # Apps whose Price is greater than 200
apps_above_200 = popular_app_cats[['Category', 'App', 'Price']][popular_app_cats['Price'] > 200]
apps_above_200
# + dc={"key": "46"} deletable=false editable=false run_control={"frozen": true} tags=["context"] id="DfJlkvW3ZFk8" colab_type="text"
# ## 7. Filter out "junk" apps
# <p>It looks like a bunch of the really expensive apps are "junk" apps. That is, apps that don't really have a purpose. Some app developer may create an app called <em>I Am Rich Premium</em> or <em>most expensive app (H)</em> just for a joke or to test their app development skills. Some developers even do this with malicious intent and try to make money by hoping people accidentally click purchase on their app in the store.
# <br>
# Let's filter out these junk apps and re-do our visualization. The distribution of apps under $100 becomes clearer.</p>
# + dc={"key": "46"} tags=["sample_code"] id="sqYi1Ke0ZFk8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 513} outputId="d175a58f-3354-4bfe-ef9f-94385a546f1d"
# Select apps priced below $100
apps_under_100 = popular_app_cats[popular_app_cats['Price'] < 100]
fig, ax = plt.subplots()
fig.set_size_inches(15, 8)
# Examine price vs category with the authentic app
ax = sns.stripplot(x=apps_under_100['Price'], y=apps_under_100['Category'],
jitter=True, linewidth=1)
ax.set_title('App pricing trend across categories after filtering for junk apps')
plt.show()
# + dc={"key": "53"} deletable=false editable=false run_control={"frozen": true} tags=["context"] id="ik_Q_9dRZFk_" colab_type="text"
# ## 8. Popularity of paid apps vs free apps
# <p>For apps in the Play Store today, there are five types of pricing strategies: free, freemium, paid, paymium, and subscription. Let's focus on free and paid apps only. Some characteristics of free apps are:</p>
# <ul>
# <li>Free to download.</li>
# <li>Main source of income often comes from advertisements.</li>
# <li>Often created by companies that have other products and the app serves as an extension of those products.</li>
# <li>Can serve as a tool for customer retention, communication, and customer service.</li>
# </ul>
# <p>Some characteristics of paid apps are:</p>
# <ul>
# <li>Users are asked to pay once for the app to download and use it.</li>
# <li>The user can't really get a feel for the app before buying it.</li>
# </ul>
# <p>Are paid apps installed as much as free apps? It turns out that paid apps have a relatively lower number of installs than free apps, though the difference is not as stark as I would have expected!</p>
# + dc={"key": "53"} tags=["sample_code"] id="SDddJQdcZFk_" colab_type="code" outputId="d18f67c2-0878-4473-8764-323dd293c242" colab={"base_uri": "https://localhost:8080/", "height": 542}
fig = go.Figure()
trace0 = fig.add_trace(go.Box(
# Data for paid apps
y=apps[apps['Type'] == 'Paid']['Installs'],
name = 'Paid'
))
trace1 = fig.add_trace(go.Box(
# Data for free apps
y=apps[apps['Type'] == 'Free']['Installs'],
name = 'Free'
))
fig.update_layout(
title = "Number of downloads of paid apps vs. free apps",
yaxis = dict(
type = 'log',
autorange = True))
# Add trace0 and trace1 to a list for plotting
fig.show()
# + dc={"key": "60"} deletable=false editable=false run_control={"frozen": true} tags=["context"] id="7fZMWU0rZFlC" colab_type="text"
# ## 9. Sentiment analysis of user reviews
# <p>Mining user review data to determine how people feel about your product, brand, or service can be done using a technique called sentiment analysis. User reviews for apps can be analyzed to identify if the mood is positive, negative or neutral about that app. For example, positive words in an app review might include words such as 'amazing', 'friendly', 'good', 'great', and 'love'. Negative words might be words like 'malware', 'hate', 'problem', 'refund', and 'incompetent'.</p>
# <p>By plotting sentiment polarity scores of user reviews for paid and free apps, we observe that free apps receive a lot of harsh comments, as indicated by the outliers on the negative y-axis. Reviews for paid apps appear never to be extremely negative. This may indicate something about app quality, i.e., paid apps being of higher quality than free apps on average. The median polarity score for paid apps is a little higher than free apps, thereby syncing with our previous observation.</p>
# <p>In this notebook, we analyzed over ten thousand apps from the Google Play Store. We can use our findings to inform our decisions should we ever wish to create an app ourselves.</p>
# + dc={"key": "60"} tags=["sample_code"] id="7CylGiR3ZFlC" colab_type="code" outputId="e4ee64f9-d2dd-458f-8b7f-5582d9c4db7a" colab={"base_uri": "https://localhost:8080/", "height": 513}
# Load user_reviews.csv
import seaborn as sns
import matplotlib.pyplot as plt
reviews_df = pd.read_csv('https://raw.githubusercontent.com/Sankalp679/SHALA/Assignments/DataCamp_Projects/The-Android-App-Market-on-Google-Play/Datasets/user_reviews.csv')
# # Join and merge the two dataframe
merged_df = pd.merge(apps,reviews_df, on ='App', how = "inner")
merged_df
# Drop NA values from Sentiment and Translated_Review columns
merged_df = merged_df.dropna(subset=['Sentiment', 'Translated_Review'])
sns.set_style('ticks')
fig, ax = plt.subplots()
fig.set_size_inches(11, 8)
# # User review sentiment polarity for paid vs. free apps
ax = sns.boxplot(x ='Type', y ='Sentiment_Polarity', data = merged_df)
ax.set_title('Sentiment Polarity Distribution')
plt.show()
|
DataCamp_Projects/The-Android-App-Market-on-Google-Play/notebook (2).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import os
# # Dataset
SimilarPairs = pd.read_csv('./archive/matchpairsDevTrain.csv')
temp_folders = pd.DataFrame(SimilarPairs.iloc[:,0])
useful_folders = pd.DataFrame(temp_folders.iloc[:,0].unique()).copy()
complete_folder_list = pd.DataFrame(os.listdir('./dataset/'))
useless_folders = complete_folder_list.copy()
cond = useless_folders[0].isin(useful_folders[0])
useless_folders.drop(useless_folders[cond].index, inplace = True)
# <b>Triplet Generation Idea:</b> <br>
# Idea is to choose Anchor and positive from useful_folders and negative from any useless_folder <br>
#
# - Total Unique IDs (With more than 1 image) = 788 <br>
# - num of images = 4441
# - Total Unique IDs (with only 1 image) = 4961 <br>
# - num of images = 8792
#
# <i>Will Always use Offline Triplet Generation Scheme after 20 epochs</i>
#
# <b>Possible Number of Triplets</b><br>
# <i>n_IDs * possible_anchor * possible_positives * possible_negatives = 112,394,314,349,408 </i> <br>
# 788 * 4441 * (4441-788) * 8792 = 112394314349408 (112 Trillion+ triplets) <BR>
#
# <b>Practicle Generation of Triplets Scheme</b> <br>
# 788 * 2 * 1* 250 = 394000 (394k Triplets) <br>
#
# - Will Always Choose 1st Image as Anchor and Second Image as Positive
# - Will Choose a random image from the pool as negative Image
788 * 2 * 1 * 250
# ## Finding all possible Triplets
# +
num_of_images_in_useful_folders = 0
for i,row in useful_folders.iterrows():
path = "./dataset/{0}/".format(row[0])
num_of_images_in_useful_folders += len(os.listdir(path))
num_of_images_in_useless_folders = 0
for i,row in useless_folders.iterrows():
path = "./dataset/{0}/".format(row[0])
num_of_images_in_useless_folders += len(os.listdir(path))
# -
num_of_images_in_useless_folders
import glob
import cv2
import numpy as np
import torch
from torch.utils.data import Dataset, DataLoader
# +
imgs_path = "dataset/"
file_list = glob.glob(imgs_path + "*")
data = {}
class_map = {}
classes = []
i = 0
for class_path in file_list:
#print(class_path)
class_name = class_path.split("\\")[-1]
#classes.append(class_name)
#class_map[class_name] = i
#i += 1
#print(class_name)
if (len(glob.glob(class_path + "\\*.jpg")) > 1 ):
classes.append(class_name)
images = []
for img_path in glob.glob(class_path + "\\*.jpg"):
images.append(img_path)
data[class_name] = images
print(data)#classes)
# -
df = pd.DataFrame(data)
df2 = pd.DataFrame(df.iloc[:,1].unique())
df
class LFW_Dataset(Dataset):
def __init__(self):
# For crop but might remove if not essential
# but controls the cropping of image
# self.img_dim = (100, 100)
# This is a dictionary with key as class name and values as a list of images path
self.data = {}
# class list
self.classes = []
# home Directory of dataset
self.imgs_path = "dataset/"
# class mapping might remove if not essential
# i = 0
# self.class_map = {}
# gets all folders name inside the dataset directory
names_list = glob.glob(self.imgs_path + "*")
#considering each folder as a unique class
for class_path in names_list:
#Spliting Class name from Path
class_name = class_path.split("\\")[-1]
# Mapping Class name to number
#self.class_map[class_name] = i
#i += 1
# Only Select those classes with more than 1 image
# if class have more than one Image
if (len(glob.glob(class_path + "\\*.jpg")) > 1 ):
#generating list of classes
self.classes.append(class_name)
images = []
for img_path in glob.glob(class_path + "\\*.jpg"):
images.append(img_path)
self.data[class_name] = images
# return total length of data
def __len__(self):
return len(self.data)
# This function must return a single datapoint when asked
# datapoint is a triplet, so it must return a triplet
# And i am returning a triplet
def __getitem__(self, idx):
# randomly pick two random classes
postive_class, negative_class = np.random.choice(classes, size=2, replace=False)
# randomly pick two images from positive_class
anchor_image , positive_image = np.random.choice(class_imgs[positive_class], size=2, replace=False)
# randomly pick one image from negative class
negative_image = np.random.choice(class_imgs[negative_class])
# create a triplet
triplet = (anchor_image, positive_image, negative_image)
#returns immutable tuple
return triplet
#img_path, class_name = self.data[idx]
#img = cv2.imread(img_path)
#img = cv2.resize(img, self.img_dim)
#img_tensor = torch.from_numpy(img)
#img_tensor = img_tensor.permute(2, 0, 1)
#class_id = torch.tensor([class_id])
#return img_tensor, class_id
# # Deep Neural Network
|
lfw-ds.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + run_control={"frozen": false, "read_only": false}
# %%typecheck
import re
import datetime
from dateutil._common import weekday
import dateutil.relativedelta as rd
import numpy as np
from pyqstrat.holiday_calendars import Calendar, get_date_from_weekday
from typing import Tuple, Mapping
FUTURE_CODES_INT = {'F': 1, 'G': 2, 'H': 3, 'J': 4, 'K': 5, 'M': 6, 'N': 7, 'Q': 8, 'U': 9, 'V': 10, 'X': 11, 'Z': 12}
FUTURES_CODES_INVERTED: Mapping[int, str] = {v: k for k, v in FUTURE_CODES_INT.items()}
FUTURE_CODES_STR = {'F': 'jan', 'G': 'feb', 'H': 'mar', 'J': 'apr', 'K': 'may', 'M': 'jun',
'N': 'jul', 'Q': 'aug', 'U': 'sep', 'V': 'oct', 'X': 'nov', 'Z': 'dec'}
def future_code_to_month(future_code: str) -> str:
'''
Given a future code such as "X", return the month abbreviation, such as "nov"
Args:
future_code (str): the one letter future code
>>> future_code_to_month('X')
'nov'
'''
assert len(future_code) == 1, f'Future code must be a single character: {future_code}'
if future_code not in FUTURE_CODES_STR: raise Exception(f'unknown future code: {future_code}')
return FUTURE_CODES_STR[future_code]
if future_code not in FUTURE_CODES_INT: raise Exception(f'unknown future code: {future_code}')
return FUTURE_CODES_INT[future_code]
def future_code_to_month_number(future_code: str) -> int:
'''
Given a future code such as "X", return the month number (from 1 - 12)
Args:
future_code (str): the one letter future code
>>> future_code_to_month_number('X')
11
'''
assert len(future_code) == 1, f'Future code must be a single character: {future_code}'
if future_code not in FUTURE_CODES_INT: raise Exception(f'unknown future code: {future_code}')
return FUTURE_CODES_INT[future_code]
def get_future_code(month: int) -> str:
'''
Given a month number such as 3 for March, return the future code for it, e.g. H
>>> get_future_code(3)
'H'
'''
return FUTURES_CODES_INVERTED[month]
class EminiFuture:
calendar = Calendar.get_calendar(Calendar.NYSE)
@staticmethod
def get_current_symbol(curr_date: datetime.date) -> str:
'''
>>> assert(EminiFuture.get_current_symbol(datetime.date(2019, 3, 14)) == 'ESH9')
>>> assert(EminiFuture.get_current_symbol(datetime.date(2019, 3, 15)) == 'ESM9')
>>> assert(EminiFuture.get_current_symbol(datetime.date(2020, 3, 14)) == 'ESH0')
'''
year = curr_date.year
month = curr_date.month
day = curr_date.day
third_friday = EminiFuture.calendar.third_friday_of_month(month, year).astype(datetime.date)
if month < 3 or (month == 3 and day < third_friday.day): month_str = 'H'
elif month < 6 or (month == 6 and day < third_friday.day): month_str = 'M'
elif month < 9 or (month == 9 and day < third_friday.day): month_str = 'U'
elif month < 12 or (month == 12 and day < third_friday.day): month_str = 'Z'
else:
month_str = 'H'
year += 1
base = 2010 if year < 2020 else 2020
fut_symbol = 'ES' + month_str + str(year - base)
return fut_symbol
@staticmethod
def get_previous_symbol(curr_future_symbol: str) -> str:
'''
>>> assert(EminiFuture.get_previous_symbol('ESH9') == 'ESZ8')
'''
month = curr_future_symbol[2]
year = int(curr_future_symbol[3])
prev_month = {'H': 'Z', 'M': 'H', 'U': 'M', 'Z': 'U'}[month]
prev_year = year if prev_month != 'Z' else year - 1
if prev_year == -1: prev_year == 9
return f'ES{prev_month}{prev_year}'
@staticmethod
def get_next_symbol(curr_future_symbol: str) -> str:
'''
>>> assert(EminiFuture.get_next_symbol('ESZ8') == 'ESH9')
'''
month = curr_future_symbol[2]
year = int(curr_future_symbol[3])
next_month = {'Z': 'H', 'H': 'M', 'M': 'U', 'U': 'Z'}[month]
next_year = year if next_month != 'H' else year + 1
if next_year == 10: next_year == 0
return f'ES{next_month}{next_year}'
@staticmethod
def get_expiry(fut_symbol: str) -> np.datetime64:
'''
>>> assert(EminiFuture.get_expiry('ESH8') == np.datetime64('2018-03-16T08:30'))
'''
month_str = fut_symbol[-2: -1]
year_str = fut_symbol[-1:]
month = future_code_to_month_number(month_str)
assert(isinstance(month, int))
year = int(year_str)
year_base = 2020 if year < 5 else 2010
year = year_base + int(year_str)
expiry_date = EminiFuture.calendar.third_friday_of_month(month, year).astype(datetime.date)
return np.datetime64(expiry_date) + np.timedelta64(8 * 60 + 30, 'm')
class EminiOption:
calendar = Calendar.get_calendar(Calendar.NYSE)
@staticmethod
def decode_symbol(name: str) -> Tuple[weekday, int, int, int]:
'''
>>> EminiOption.decode_symbol('E1AF8')
(MO, 2018, 1, 1)
'''
if re.match('EW[1-4].[0-9]', name): # Friday
year = int('201' + name[-1:])
if year in [2010, 2011]: year += 10
week = int(name[2:3])
month = future_code_to_month_number(name[3:4])
return rd.FR, year, month, week
if re.match('E[1-5]A.[0-9]', name): # Monday
year = int('201' + name[-1:])
if year in [2010, 2011]: year += 10
week = int(name[1:2])
month = future_code_to_month_number(name[3:4])
return rd.MO, year, month, week
if re.match('E[1-5]C.[0-9]', name): # Wednesday
year = int('201' + name[-1:])
if year in [2010, 2011]: year += 10
week = int(name[1:2])
month = future_code_to_month_number(name[3:4])
return rd.WE, year, month, week
if re.match('EW[A-Z][0-9]', name): # End of month
year = int('201' + name[-1:])
if year in [2010, 2011]: year += 10
week = -1
month = future_code_to_month_number(name[2:3])
return rd.WE, year, month, week
else:
raise Exception(f'could not decode: {name}')
@staticmethod
def get_expiry(symbol: str) -> np.datetime64:
'''
>>> EminiOption.get_expiry('EW2Z5')
numpy.datetime64('2015-12-11T15:00')
>>> EminiOption.get_expiry('E3AF7')
numpy.datetime64('2017-01-17T15:00')
>>> EminiOption.get_expiry('EWF0')
numpy.datetime64('2020-01-31T15:00')
'''
assert ':' not in symbol, f'{symbol} contains: pass in option root instead'
weekday, year, month, week = EminiOption.decode_symbol(symbol)
expiry = get_date_from_weekday(weekday.weekday, year, month, week)
if weekday in [rd.WE, rd.FR]:
expiry = EminiOption.calendar.add_trading_days(expiry, num_days=0, roll='backward')
else:
expiry = EminiOption.calendar.add_trading_days(expiry, num_days=0, roll='forward')
# Option expirations changed on 9/20/2015 from 3:15 to 3 pm -
# See https://www.cmegroup.com/market-regulation/files/15-384.pdf
expiry += np.where(expiry < np.datetime64('2015-09-20'), np.timedelta64(15 * 60 + 15, 'm'), np.timedelta64(15, 'h'))
return expiry
if __name__ == "__main__":
import doctest
doctest.testmod(optionflags=doctest.NORMALIZE_WHITESPACE)
|
pyqstrat/src_nb/markets.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.7 64-bit (''venv'': venv)'
# name: python3
# ---
# # Tutorial: How to use weirwood pyfinance API
#
# This python API allows you to retrieve financial statements (filings) and daily prices from more than 9000 companies.
#
# In this tutorial you will learn how to use pyfinance python API for:
#
# - Download financial statements from US companies
# - Download daily stock prices
# # Prerequisites: Install pyfinance
#
# The following code block will download pyfinance package in your environment. The only requirement is to have python > 3.X
# Once the download finishes remember to reload the kernel
pip install weirwood_pyfinance
# # Import FinTen
#
# In order to use pyfinance API you will need to connect to Finten Backend. The following line will show you how to instantiate Finten without login required
from weirwood_pyfinance import FinTen
finten = FinTen()
# # Get filings
#
# With _get_filings('COMPANY_TICKER')_ you can download any of the available public companies financial statements as we show in the following chunk.
tesla_filings = finten.get_filings('TSLA')
tesla_filings.head()
# The returning object is a pandas dataframe which columns contain numerical information about the company and metadata, such as the release date of the filing, the type of filing, the industry which the company belongs to... For the numerical values it returns variables such as Assets, Liabilities, Equity.. and other relevant financial information about the compamny requested
tesla_filings.info()
# # Available companies
#
# Using _.get_tickers()_ we can get a list of available companies
all_available_tickers = finten.get_tickers()
all_available_tickers
#
# # Get prices
# Finally, we can also download daily prices using _.get_prices('COMPANY_TICKER')_ public method. The return type is also a pandas dataframe with the daily stock market price of the company.
tesla_prices = finten.get_prices('TSLA')
tesla_prices.tail()
|
pyfinance example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from skimage import data, color
import numpy as np
import matplotlib.pyplot as plt
def show_image(image, title='Image', cmap_type='gray'):
plt.imshow(image, cmap=cmap_type)
plt.title(title)
plt.axis('off')
plt.show()
# +
# Import the modules from skimage
# Load the rocket image
rocket = data.rocket()
# Convert the image to grayscale
gray_scaled_rocket = color.rgb2gray(rocket)
# Show the original image
show_image(rocket, 'Original RGB image')
# Show the grayscale image
show_image(gray_scaled_rocket, 'Grayscale image')
# -
type(rocket)
red = rocket[:, :, 0]
plt.hist(red.reshape(-1),bins=256)
# +
from skimage.filters import threshold_otsu,threshold_local
thers = threshold_otsu(gray_scaled_rocket)
img = gray_scaled_rocket > thers
show_image(img)
# +
block_size = 97
local_thresh = threshold_local(gray_scaled_rocket, block_size)
img = gray_scaled_rocket > local_thresh
show_image(img)
show_image(gray_scaled_rocket)
# -
|
courses file/image processing in python/capter1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="T8IkEZ1mF3MP"
# # Time Series Forecasting System
# Time Series is a big component of our everyday lives. They are in fact used in medicine (EEG analysis), finance (Stock Prices) and electronics (Sensor Data Analysis). Many Machine Learning models have been created in order to tackle these types of tasks, two examples are ARIMA (AutoRegressive Integrated Moving Average) models and RNNs (Recurrent Neural Networks).
#
# + [markdown] colab_type="text" id="0a1a3jB3GYI8"
# # Data Source
#
# For Time series analysis, we are going to deal with Stock market Analysis. This dataset is based US-based stocks daily price and volume data.
# Dataset taken for analysis is IBM stock market data from 2006-01-01 to 2018-01-01.
#
# Below are the key fields in the dataset:
#
# __`Date, Open, High, Low, Close, Volume, Name`__
# + [markdown] colab_type="text" id="QJiCYDCPH0JX"
# # Import Libraries
# + colab={} colab_type="code" id="I1SfYI6nebcN"
import warnings
warnings.filterwarnings('ignore')
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
# + [markdown] colab_type="text" id="7ftJJhAPH9ZJ"
# # Load Data
# + colab={"base_uri": "https://localhost:8080/", "height": 195} colab_type="code" id="zLpZyyGucahj" outputId="c3cef8f3-bc13-42e6-f496-4096a13e4bd3"
df = pd.read_csv("IBM_2006-01-01_to_2018-01-01.csv")
df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" id="HqoaX9Zy8wvW" outputId="03cf2223-ec08-45fa-8803-714664d9fe6e"
print(df.shape)
print(df.columns)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="QL9CyyYg83ow" outputId="ffcc3cf0-1be9-4b2d-81d6-4023e849aeaf"
# Cleaning up the data
df.isnull().values.any()
df = df.dropna()
df.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 225} colab_type="code" id="FXk5auRRh2O6" outputId="38ecc2d6-fb6f-405c-e6aa-72f2a59f9791"
df.index = pd.to_datetime(df['Date'])
df.head()
# + [markdown] colab_type="text" id="HDxYwSHoJmDP"
# # Note
# This dataset is composed of different features.We will just examine the “Open” stock prices feature. This same analysis can be repeated for most of the other features.
# + [markdown] colab_type="text" id="piiRMy-_IIZv"
# # Visualization
# -
# ### Visualizing the High and Low prices of IBM
# + colab={"base_uri": "https://localhost:8080/", "height": 285} colab_type="code" id="c7B6d-PTfKuC" outputId="190fa06f-a8f0-413f-b94b-efab802c758d"
dr = df[['High', 'Low']]
dr.plot()
plt.title('IBM Returns');
# -
# ### Q1: Visualize the Open and Close prices of IBM
# + colab={"base_uri": "https://localhost:8080/", "height": 285} colab_type="code" id="vWGWKmhKgJOr" outputId="c59eae5b-c44a-4333-9057-8fc405d01986"
# -
# ### Q2: Visualize the Open and Close Cumulative Prices of IBM
# + colab={"base_uri": "https://localhost:8080/", "height": 285} colab_type="code" id="8Cc0Zseff-4k" outputId="621dc52c-8621-45ef-a7f0-2d8c76fb7e73"
# + [markdown] colab_type="text" id="J0nwpAKqKDfH"
# #### Before we start working on Time Series forecasting, Let's analyse the autocorrelation plot of the “Open” feature with respect to a few lag values
# -
# ### Auto-correlation plot with Lag 1
# + colab={"base_uri": "https://localhost:8080/", "height": 350} colab_type="code" id="ivjXx85Bf_h2" outputId="dd5ce4e7-8dc9-428e-a958-8e5b9ce42f80"
# START_CODE_HERE - plot the Autocorrelation plot for feature 'Open'
from pandas.plotting import lag_plot
plt.figure(figsize=(5,5))
lag_plot(df['Open'], lag=1)
plt.title('IBM Autocorrelation plot - Lag 1');
# END_CODE_HERE
# -
# ### Q3: Visualize the Auto-Correlation plot for IBM Open prices with Lag 5
# + colab={"base_uri": "https://localhost:8080/", "height": 350} colab_type="code" id="1BWxkN2ZhDGq" outputId="85296efa-0345-459e-a91e-adc3459c6def"
# -
# We see a definite linear trend in the auto-correlation plot telling us there is some correlation in prices with respect to prices from previous 1 / 5 days of lag which sets up the stage of forecasting future prices based on past price data
# + [markdown] colab_type="text" id="1n4_JM9Lhb5K"
# ## Build Train-Test Datasets
# + [markdown] colab_type="text" id="KLTiykRpKh46"
# #### Now, Let's divide the data into a training and test set. Once done so, we can plot both on the same figure in order to get a feeling of how does our Time Series looks like
# + colab={} colab_type="code" id="dlEMHunIgLjE"
train_data, test_data = df.iloc[0:int(len(df)*0.8), :], df.iloc[int(len(df)*0.8):, :]
# + colab={"base_uri": "https://localhost:8080/", "height": 458} colab_type="code" id="nXQdYUp0isTi" outputId="8d99b465-1ca5-42ad-a9ff-9fa9c80ff3ff"
plt.figure(figsize=(12,7))
plt.title('IBM Prices')
plt.xlabel('Dates')
plt.ylabel('Prices')
plt.plot(train_data['Open'], 'blue', label='Training Data')
plt.plot(test_data['Open'], 'green', label='Testing Data')
plt.legend();
# + [markdown] colab_type="text" id="wJhAHw6PKrQh"
# # ARIMA (AutoRegressive Integrated Moving Average)
#
# The acronym of ARIMA stands for:
#
# AutoRegressive(AR) = the model takes advantage of the connection between a predefined number of lagged observations and the current one.
#
# Integrated(I) = differencing between raw observations (eg. subtracting observations at different time steps).
#
# Moving Average(MA) = the model takes advantage of the relationship between the residual error and the observations.
#
# The ARIMA model makes use of three main parameters (p,d,q). These are:
#
# p = number of lag observations.
#
# d = the degree of differencing.
#
# q = the size of the moving average window.
#
#
# + [markdown] colab_type="text" id="Ftf6yg95L7CR"
# ## Understaning the ARIMA Model
#
# ### the ARIMA parameters - used to help model the major aspects of a times series: seasonality, trend, and noise. These parameters are labeled p,d,and q. You have already learnt a fair bit of this in the curriculum but following is a brief refresher.
#
# __p:__ is the parameter associated with the auto-regressive aspect of the model, which incorporates past values. For example, forecasting that if it rained a lot over the past few days, you state its likely that it will rain tomorrow as well.
#
# __d:__ is the parameter associated with the integrated part of the model, which effects the amount of differencing to apply to a time series. You can imagine an example of this as forecasting that the amount of rain tomorrow will be similar to the amount of rain today, if the daily amounts of rain have been similar over the past few days.
#
# __q:__ is the parameter associated with the moving average part of the model.
#
# ### Approach to determine the parameters
# There are many ways to choose these values statistically, such as looking at auto-correlation plots, correlation plots, domain experience, etc.
#
# Another approach is to perform a grid search over multiple values of p,d,q using some sort of performance criteria. The Akaike information criterion (AIC) is an estimator of the relative quality of statistical models for a given set of data. Given a collection of models for the data, AIC estimates the quality of each model, relative to each of the other models.
#
#
# In this exercise, we will look into the statistical method of getting these values from auto-correlation and correlation plots.
# + [markdown] colab_type="text" id="iP5Qup5I7l0x"
# ### Stationarity of the data - Determine the d value
#
# Stationarity typically indicates various statistical measures of the time series do not change over time. Thus, a time series is stationary when its mean, variance and auto-correlation, etc., are constant over time.
#
# Most time-series forecasting models typically perform well when the series is stationary and hence it is important to find out if your time-series dataset is stationary.
#
# ARIMAs that include differencing (i.e., d > 0) assume that the data becomes stationary after differencing. This is called difference-stationary.
#
# Auto-correlation plots are an easy way to determine whether your time series is sufficiently stationary for modeling.
#
# If the plot does not appear relatively stationary, your model will likely need a differencing term.
#
# The Augmented Dickey-Fuller test is an important statistical test which we will use to prove if the series is stationary or not and take necessary steps in case it is not stationary.
# + colab={"base_uri": "https://localhost:8080/", "height": 390} colab_type="code" id="0TfnfVdMleJY" outputId="069bc3f8-aa6b-44f5-8c05-b5e31913d179"
window = 7
train_series = train_data['Open']
#Determing rolling statistics
rolmean = train_series.rolling(window).mean()
rolstd = train_series.rolling(window).std()
#Plot rolling statistics:
fig = plt.figure(figsize=(10, 6))
orig = plt.plot(train_series, color='blue',label='Original')
mean = plt.plot(rolmean, color='red', label='Rolling Mean')
std = plt.plot(rolstd, color='black', label = 'Rolling Std')
plt.legend(loc='best')
plt.title('Rolling Mean & Standard Deviation');
# + colab={"base_uri": "https://localhost:8080/", "height": 151} colab_type="code" id="RM44jwLammeY" outputId="509bda4d-22da-43e7-cde6-bedbb0f24dce"
from statsmodels.tsa.stattools import adfuller
dftest = adfuller(train_series, autolag='AIC')
dfoutput = pd.Series(dftest[0:4], index=['Test Statistic','p-value','#Lags Used','Number of Observations Used'])
for key,value in dftest[4].items():
dfoutput['Critical Value (%s)'%key] = value
dfoutput
# + [markdown] colab_type="text" id="R1YJhVNzZD87"
# If the p-value is small beyond a specific significance level threshold, let's consider that to be a standard value of 0.05, then we can say the series is stationary. F
#
# rom the above statistics, we can observe that the p-value is 0.539 which proves that our series is not stationary.
#
# To get stationary data, there are many techniques. We can use log, differencing and so on. Let's use a first order differencing here.
# -
# ### Q4: Apply a first order differencing on the training data
#
# Hint: Check out the __`diff()`__ function in pandas and try using it on the __`train_series`__ dataset
# + colab={} colab_type="code" id="EomzoS5494th"
train_diff = <YOUR CODE HERE>
train_diff = train_diff.dropna(inplace = False)
# -
# ### Q5: Visualize Rolling statistics for differenced train data
# + colab={"base_uri": "https://localhost:8080/", "height": 390} colab_type="code" id="kStK1sqjn_df" outputId="883dce6e-9e28-441d-d14b-aca927d7c283"
#Determing rolling statistics
#Plot rolling statistics:
# -
# ### Q6: Compute AD-Fuller Stats for differenced train data
# + colab={"base_uri": "https://localhost:8080/", "height": 151} colab_type="code" id="OgNoMUJ1oE7V" outputId="01ee99ed-aa58-41eb-a54e-ad3a2a3c5f28"
# + [markdown] colab_type="text" id="-T_wDkhRZdqm"
# After differencing, the p-value is extremely small. Thus this series is very likely to be stationary.
# + [markdown] colab_type="text" id="QhY85n0qa85g"
# ### ACF Plots (Auto Correlation Function):
# ACF is an auto-correlation function which gives us correlation of any series with its lagged values(previous timestep values).
#
# ACF plot describes the correlation of the current value with the previous lagged values(specified by *lags*).
#
# For example, how the dependency chain is followed as direct dependency .... $S_{t-2} --> S_{t-1} --> S_t$*
#
# Also, ACF finds correlation between $S_{t-2} --> S_t$ (indirect dependency).
#
# * --> = represents dependency
#
# #### Limitation:
# ACF is not very accurate as indirect dependency is affected by direct dependency and so the plots are always above the confidence band(as shown below).
# + [markdown] colab_type="text" id="fXEjp5BubI-L"
# ### PACF Plots: Pearson Auto Correlation Function:
# PACF plots models the indirect dependencies and is not affected by the direct dependencies.
#
# $S_{t-2} --> S_t$
#
# From the below example we can see how today's value is affected by the last 10 days.
#
# The points that lie inside the blue confidence band do not correlate with or affect today's value. In ACF, we saw that all values are above the confidence band(as $S_{t-2} --> S{t}$ is affected by $S_{t-1} --> S_t$), which is not a good representation of the correlation.
#
#
# In PACF, indirect dependencies are modelled well.
# + [markdown] colab_type="text" id="9nG-g8DDZhL5"
# ## ACF and PACF - AR and MA Intuition
# The partial autocorrelation at lag k is the correlation that results after removing the effect of any correlations due to the terms at shorter lags.
#
# ### Autoregression Intuition
# Consider a time series that was generated by an autoregression (AR) process with a lag of k.
#
# We know that the ACF describes the autocorrelation between an observation and another observation at a prior time step that includes direct and indirect dependence information.
#
# This means we would expect the ACF for the AR(k) time series to be strong to a lag of k and the inertia of that relationship would carry on to subsequent lag values, trailing off at some point as the effect was weakened.
#
# We know that the PACF only describes the direct relationship between an observation and its lag. This would suggest that there would be no correlation for lag values beyond k.
#
# This is exactly the expectation of the ACF and PACF plots for an AR(k) process.
#
# ### Moving Average Intuition
# Consider a time series that was generated by a moving average (MA) process with a lag of k.
#
# Remember that the moving average process is an autoregression model of the time series of residual errors from prior predictions. Another way to think about the moving average model is that it corrects future forecasts based on errors made on recent forecasts.
#
# We would expect the ACF for the MA(k) process to show a strong correlation with recent values up to the lag of k, then a sharp decline to low or no correlation. By definition, this is how the process was generated.
#
# For the PACF, we would expect the plot to show a strong relationship to the lag and a trailing off of correlation from the lag onwards.
#
# Again, this is exactly the expectation of the ACF and PACF plots for an MA(k) process.
# -
# ### Plot ACF and PACF on the original train series
# + colab={"base_uri": "https://localhost:8080/", "height": 499} colab_type="code" id="TKCS364Bq2BP" outputId="6c65562b-c492-4984-b2dd-edf97a2e682e"
from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
fig, ax = plt.subplots(2, 1, figsize=(12,8))
plot_acf(train_series, ax=ax[0]); #
plot_pacf(train_series, ax=ax[1]);
# -
# ### Q7: Plot ACF and PACF on the differenced train series
# + colab={"base_uri": "https://localhost:8080/", "height": 499} colab_type="code" id="FPxG3Hkwpl6u" outputId="74e69c20-903d-45f4-a111-ddfcc15af99a"
# + [markdown] colab_type="text" id="z4veOUVxSP8j"
# # How to determine p, d, q
#
# It's easy to determine d. In our case, we see the first order differencing make the ts stationary. Hence d = 1
#
# AR model might be investigated first with lag length selected from the PACF or via empirical investigation. In our case, it's clearly that within 5 lags the AR is significant. Which means, we can use AR = 5 i.e, p = 5
#
# To avoid the potential for incorrectly specifying the MA order to be too high we set MA = 0 i.e q = 0 by taking a look at the ACF plot though we do have a value of 5 which is significant considering the interval but we start off with the first lag value i.e q = 0.
#
# Hence:
#
# - p=5
# - d=1
# - q=0
# + [markdown] colab_type="text" id="dCEcnGXsdLkl"
# # Evaluation of ARIMA Model
#
# In order to evaluate the ARIMA model,we can use two different error functions:
#
# - Mean Squared Error (MSE)
# - Symmetric Mean Absolute Percentage Error (SMAPE)
#
# SMAPE is commonly used as an accuracy measure based on relative errors
# + [markdown] colab_type="text" id="iClH4YhNdSCB"
# ### SMAPE
#
# 
#
# SMAPE is not currently supported in Scikit-learn as a loss function, therefore we first create this function.
# + colab={} colab_type="code" id="v36vtXA1dZPo"
def smape_kun(y_true, y_pred):
# START_CODE_HERE
return np.mean((np.abs(y_pred - y_true) * 200 / (np.abs(y_pred) + np.abs(y_true))))
# END_CODE_HERE
# -
# ### Q8: Difference the Test Series
# + colab={} colab_type="code" id="_YRY19919xXD"
test_series = test_data['Open']
test_diff = <YOUR CODE HERE>
test_diff = test_diff.dropna(inplace = False)
# -
# ### Q9: Train and Forecast using ARIMA Model by filling in the necessary blocks
#
# Note: Here we will use a rolling point-based prediction for the ARIMA model where we tried to predict every day's (t) stock price in the test data by using both the training data as well as the previous (n - t) days of test data also to fit the model.
# Of course this is not the only way for forecasting and you can do it in multiple ways e.g just use train data to forecast, use a window of days to forecast including test data and so on.
# + colab={} colab_type="code" id="9_0v9P7OtoVu"
from statsmodels.tsa.arima_model import ARIMA
from sklearn.metrics import mean_squared_error
# + colab={"base_uri": "https://localhost:8080/", "height": 202} colab_type="code" id="IUvMaWIRg4VR" outputId="6dca97a1-65c1-4cc3-9adf-905995a54abe"
# %%time
history = [x for x in train_diff]
predictions = list()
for t in range(len(test_diff)):
# START_CODE_HERE - call the ARIMA Method with history and params
model = <YOUR CODE HERE> # initialize the model with history and right order of parameters
model_fit = <YOUR CODE HERE> # fit the model
# END_CODE_HERE
output = <YOUR CODE HERE> # use forecast on the fitted model
yhat = output[0][0]
predictions.append(yhat)
obs = test_diff[t]
history.append(obs)
if t % 100 == 0:
print('Test Series Point: {}\tPredicted={}, Expected={}'.format(t, yhat, obs))
# -
# ### Reverse Transform the forecasted values
#
# This is very important. Since we used differencing of the first order in the series before training, we need to reverse transform the values to get meaningful price forecasts.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="unArWBuS7o5z" outputId="cee241e1-e170-4063-e676-752fc5dbb70d"
reverse_test_diff = np.r_[test_series.iloc[0], test_diff].cumsum()
reverse_predictions = np.r_[test_series.iloc[0], predictions].cumsum()
reverse_test_diff.shape, reverse_predictions.shape
# -
# ### Evaluate model performance
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" id="NGDHu85n2clu" outputId="af8b3b73-5757-4c07-ba29-ed6a37ba0616"
error = mean_squared_error(reverse_test_diff, reverse_predictions)
print('Testing Mean Squared Error: %.3f' % error)
error2 = smape_kun(reverse_test_diff, reverse_predictions)
print('Symmetric Mean absolute percentage error: %.3f' % error2)
# + [markdown] colab_type="text" id="kVShn5zgMPcX"
# The loss results for this model are available above. According to the MSE, the model loss is quite low but for SMAPE is instead consistently higher. One of the main reason for this discrepancy is because SMAPE is commonly used loss a loss function for Time Series problems and can, therefore, provide a more reliable analysis. That showed there is still room for improvement of our model.
# + [markdown] colab_type="text" id="g5XoIrqDMe7c"
# ## Let's Visualize the forecast results
# + colab={} colab_type="code" id="V3S7iMXv8nKg"
reverse_test_diff_series = pd.Series(reverse_test_diff)
reverse_test_diff_series.index = test_series.index
reverse_predictions_series = pd.Series(reverse_test_diff)
reverse_predictions_series.index = test_series.index
# -
# ### Visualizing train, test and forecast prices
# + colab={"base_uri": "https://localhost:8080/", "height": 458} colab_type="code" id="LEBu1A6P8aJp" outputId="38f7847a-465b-4fa0-a14d-f5d78b70c5c5"
plt.figure(figsize=(12,7))
plt.title('IBM Prices')
plt.xlabel('Dates')
plt.ylabel('Prices')
plt.plot(train_series, color='blue', label='Training Prices')
plt.plot(reverse_test_diff_series, color='green', marker='.', label='Testing Prices - Reverse Diff Transform')
plt.plot(reverse_test_diff_series, color='red', linestyle='--', label='Forecasted Prices - Reverse Diff Transform')
plt.legend();
# -
# ### Q10: Visualize only test and forecast prices
# + colab={"base_uri": "https://localhost:8080/", "height": 458} colab_type="code" id="fOKhtSM79hM_" outputId="694f2659-1a0e-4665-d453-13<PASSWORD>6cda28"
# + [markdown] colab_type="text" id="8k-cTIUyMzd_"
# This analysis using ARIMA has performed pretty well in forecasting prices
# + [markdown] colab_type="text" id="fcM7rwQFCqHL"
# # Time Series Forecasting with Deep Learning
#
# The approach uses sequential models, to be more specific - LSTMs, to build a deep learning model that predicts the 'Open' Stock prices of IBM over a period of two years by using data from the previous 10 years.
# + [markdown] colab_type="text" id="Ege_yD9YDmSN"
# ### LSTM: A brief overview
#
# What are LSTMs? : https://medium.com/deep-math-machine-learning-ai/chapter-10-1-deepnlp-lstm-long-short-term-memory-networks-with-math-21477f8e4235
#
# Long short-term memory (LSTM) units (or blocks) are a building unit for layers of a recurrent neural network (RNN). A RNN composed of LSTM units is often called an LSTM network. A common LSTM unit is composed of a cell, an input gate, an output gate and a forget gate. The cell is responsible for "remembering" values over arbitrary time intervals; hence the word "memory" in LSTM. Each of the three gates can be thought of as a "conventional" artificial neuron, as in a multi-layer (or feedforward) neural network: that is, they compute an activation (using an activation function) of a weighted sum. Intuitively, they can be thought as regulators of the flow of values that goes through the connections of the LSTM; hence the denotation "gate". There are connections between these gates and the cell.
#
# The expression long short-term refers to the fact that LSTM is a model for the short-term memory which can last for a long period of time. An LSTM is well-suited to classify, process and predict time series given time lags of unknown size and duration between important events. LSTMs were developed to deal with the exploding and vanishing gradient problem when training traditional RNNs.
#
# Source: [Wikipedia](https://en.wikipedia.org/wiki/Long_short-term_memory)
# + [markdown] colab_type="text" id="c8hK0f8wb79O"
# ### Headers
# + colab={} colab_type="code" id="Y4Fo2a_Rg5_w"
# Let's load the libraries and dependencies for the deep learning model
from sklearn.preprocessing import MinMaxScaler
# %tensorflow_version 1.x
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, LSTM, Dropout, GRU, Bidirectional
from tensorflow.keras.optimizers import SGD
# + [markdown] colab_type="text" id="VuF5ziiYcDD-"
# ### Load Data
# + colab={"base_uri": "https://localhost:8080/", "height": 225} colab_type="code" id="loos_QCqjZl8" outputId="8db32178-6d24-41a5-aca4-c8886d2ad4d0"
df = pd.read_csv("IBM_2006-01-01_to_2018-01-01.csv")
df.isnull().values.any()
df = df.dropna()
df.index = pd.to_datetime(df['Date'])
df.head()
# + [markdown] colab_type="text" id="EeR4wBTNSAUX"
# # Note
# This dataset is composed of different features.we will just examine the "Open" stock prices feature. This same analysis can be repeated for most of the other features.
# + [markdown] colab_type="text" id="Nvliw7fqMCcF"
# ## Build Train-Test Datasets
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="e51pJz3Ajs0u" outputId="9d7fa2bd-044f-4f6f-cc4d-8930eb8b61f5"
# Splitting the train and test set considering 'Open' feature from the dataset
train_data, test_data = df.iloc[0:int(len(df)*0.8), :], df.iloc[int(len(df)*0.8):, :]
train_series = train_data['Open']
test_series = test_data['Open']
train_series.shape, test_series.shape
# + [markdown] colab_type="text" id="vSp2L8YIMfKN"
# ### Q11: Visualize train and test price data
# + colab={"base_uri": "https://localhost:8080/", "height": 458} colab_type="code" id="gBnSZGfSj6Cs" outputId="6e071f16-4d78-413f-b175-37e8cb603955"
# + [markdown] colab_type="text" id="k8f0sPbUMvg7"
# ### Scaling
# As stock prices can vary across a wide range, we scale the data to have zero mean and unit variance.
#
# This is done to ensure that the gradient descent is sooner when learning a deep learning model
# -
# ### Q12: Use the initialized min-max scaler to scale the prices in train_series
# + colab={} colab_type="code" id="-7D3Hvo_kC2F"
sc = MinMaxScaler(feature_range=(0,1))
# START_CODE_HERE
training_set_scaled = <YOUR CODE HERE>
# END_CODE_HERE
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="Rms7Nr3LIeuV" outputId="5a92caf3-f015-44c1-fbe5-e7451ba445eb"
training_set_scaled.shape
# + [markdown] colab_type="text" id="aEaPD_p2Nu87"
# ### Train Data Preparation
#
# Train data uses the previous 60 days (two months) data to predict the stock price of the next day.
# The data is prepared just like a sliding window approach, where
# *window_size = 60*
#
# Sample image for sliding window:
# 
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="3LHfiLnhkGpN" outputId="278d904e-00b5-44fd-9ca9-cc6bc37c7249"
#1 output and 60 values inputs
# So for each element of training set (output), we have 60 previous training set elements (input)
X_train = []
y_train = []
for i in range(60, len(training_set_scaled)):
X_train.append(training_set_scaled[i-60:i,0])
y_train.append(training_set_scaled[i,0])
X_train, y_train = np.array(X_train), np.array(y_train)
X_train.shape, y_train.shape
# + [markdown] colab_type="text" id="YdzzWChQROAg"
# #### Reshape X_train
#
# Now we reshape X_train in the format like:
#
# (batch_size, timesteps, input_dim) => (m, features, $x_{i1}$)
#
# The X_train should be now: (2709, 60, 1)
#
# 60 features = 60 day sliding window
#
# $x_{i1}$ = 1 data point for each feature and i represents the feature
#
#
# + colab={} colab_type="code" id="KQRlDlXMkKCV"
# Reshaping X_train for efficient modeling
X_train = np.reshape(X_train, (X_train.shape[0],X_train.shape[1], 1))
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="I_FLbtZkT-RF" outputId="70bd8ffd-f73b-4091-ef7f-d594ec30873b"
X_train.shape
# + [markdown] colab_type="text" id="PIrlm79SVn5i"
# ### LSTM Regression model
#
# We use [LSTM](https://www.tensorflow.org/api_docs/python/tf/keras/layers/LSTM):
# * units - output dimensions
# * return_sequences is set to True to get all the hidden state vectors information
#
# The model uses 2 LSTM layers followed by a Dense Layer with a single neuron to output regression prediction.
# + [markdown] colab_type="text" id="W1gSynqKGGPJ"
# #### Similar Model Architecture (dimensions not exact)
#
# 
#
# -
# ### Q13: Build the LSTM based forecasting DL Model architecture
#
# Hints:
#
# - Fill the second LSTM layer using an LSTM cell with 64 units, remember NOT to set return_sequences to True as we are only concerned about passing the last sequence output to the next layer
# - Fill the Output layer with 1 unit
# - Compile the model with mentioned optimizer and loss values
# + colab={"base_uri": "https://localhost:8080/", "height": 319} colab_type="code" id="Vhe8p4iqkPfB" outputId="682d878c-bdd6-4d28-e72a-1b8cb3c3f24b"
regressor = Sequential()
# First LSTM layer with Dropout regularisation
regressor.add(LSTM(units=64, return_sequences=True, input_shape=(X_train.shape[1],1)))
regressor.add(Dropout(0.2))
# Second LSTM layer
<YOUR CODE HERE>
# The output layer
<YOUR CODE HERE>
# Compiling the RNN - optimizer(rmsprop)and loss(mean squared error)
<YOUR CODE HERE>
regressor.summary()
# -
# ### Train the model
# + colab={"base_uri": "https://localhost:8080/", "height": 554} colab_type="code" id="MBRin4eMAsT_" outputId="91760fb0-6085-42c1-f507-8adf135eadb6"
regressor.fit(X_train,y_train, epochs=15, batch_size=64, validation_split=0.1, shuffle=False)
# + [markdown] colab_type="text" id="tXyIJG9vauFj"
# ### Test Data Forecasting
#
# #### Data Preparation:
# Lets prepare the test data just like we did with the train data.
#
# Remember to start forecasting on the first day of the test data, we need the last 60 days of train data.
#
# Thus, the following steps have been performed so first 60 entires of test set have 60 previous values from the train dataset
# -
# ### Q14: Get the last 60 records from train_series
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="4mLalxGGCxzf" outputId="1df0414a-9ab3-4443-e527-ca56226c674e"
train_last60 = <YOUR CODE HERE>
print(train_last60.shape)
assert train_last60.shape == (60,), ("Oops! There is a data dimension mismatch error. Hint: Slice the last 60 records from train_series")
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="vhfLNVCnCrNE" outputId="7c6324e3-572a-491a-e8e8-ab7b9ec99e91"
test_series.shape
# -
# ### Q15: Combine both train_last60 and test_series together
#
# Hint: Check pandas __`concat()`__
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="_XrJWDeZC_nG" outputId="2ceb51ec-08f9-440b-a6be-9cfc26883ab5"
new_test_series = <YOUR CODE HERE>
print(new_test_series.shape)
assert new_test_series.shape == (664,), ("Oops! There is a data dimension mismatch error. Hint: Use pandas concat with the right axis parameter")
# -
# ### Q16: Scale the test dataset (new_test_series) using the trained MinMaxScaler transformer - sc
#
# Hint: Don't fit the scaler again here since it has already been trained
# + colab={} colab_type="code" id="E6EEBJmFkRsB"
test_set_scaled = <YOUR CODE HERE>
# -
# ### Prepare Test dataset Windows of 60 days each
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="ja7H-jR0kWvX" outputId="ccc648fb-249a-4fd0-b4d7-a6f9ae827a0b"
# Preparing X_test and predicting the prices
X_test = []
for i in range(60,len(test_set_scaled)):
X_test.append(test_set_scaled[i-60:i,0])
X_test = np.array(X_test)
X_test = np.reshape(X_test, (X_test.shape[0],X_test.shape[1],1))
X_test.shape
# -
# ### Model Prediction and Reverse Transform of Prices
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="BBQ-3C53DZ5W" outputId="8beb0870-626b-4e77-bcae-3d8b05107a0d"
predicted_stock_price = regressor.predict(X_test)
predicted_stock_price_revtrans = sc.inverse_transform(predicted_stock_price).ravel()
predicted_stock_price_revtrans_series = pd.Series(predicted_stock_price_revtrans)
predicted_stock_price_revtrans_series.index = test_series.index
predicted_stock_price_revtrans_series.shape, test_series.shape
# + [markdown] colab_type="text" id="Dhjde2l_Dx1y"
# ## Model Evaluation
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" id="iLDpDXl5kcnH" outputId="16f9238f-2d13-4c19-bed3-b3281f0388b7"
# Evaluating our model
error = mean_squared_error(test_series, predicted_stock_price_revtrans_series)
print('Testing Mean Squared Error: %.3f' % error)
error2 = smape_kun(test_series, predicted_stock_price_revtrans_series)
print('Symmetric Mean absolute percentage error: %.3f' % error2)
# + [markdown] colab_type="text" id="tR06LgzkD58c"
# ## Visualizing the results from model predictions
# -
# ### Visualize train, test and forecasted prices
# + colab={"base_uri": "https://localhost:8080/", "height": 458} colab_type="code" id="g32mqimHkYja" outputId="c998166f-1e23-4aac-d1bf-98ee4bfe82ee"
plt.figure(figsize=(12,7))
plt.title('IBM Prices')
plt.xlabel('Dates')
plt.ylabel('Prices')
plt.plot(train_series, color='blue', label='Training Prices')
plt.plot(test_series, color='green', label='Testing Prices')
plt.plot(predicted_stock_price_revtrans_series, color='red', linestyle='--', label='Forecasted Prices - Reverse Transform')
plt.legend();
# -
# ### Q17: Visualize only test and forecast prices
# + colab={"base_uri": "https://localhost:8080/", "height": 458} colab_type="code" id="UH7f5oS8Fqxy" outputId="25bb2da0-b464-4a96-f357-41fead67b1ba"
# + [markdown] colab_type="text" id="c1xuGfJuQTOp"
# # Conclusion
#
# Remember we did a rolling point-based prediction for the ARIMA model where we tried to predict every day's (t) stock price in the test data by using both the training data as well as the previous (n - t) days of test data also to fit the model which gave it such good results vs. the LSTM model where we used 2 months of rolling window price data to predict the next day's price.
|
notebook/mec-18.5.1-time-series-analysis-mini-project/Mini_Project_Time_Series_Forecasting.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Regression with BIWI head pose dataset
# This is a more advanced example to show how to create custom datasets and do regression with images. Our task is to find the center of the head in each image. The data comes from the [BIWI head pose dataset](https://data.vision.ee.ethz.ch/cvl/gfanelli/head_pose/head_forest.html#db), thanks to <NAME> al. We have converted the images to jpeg format, so you should download the converted dataset from [this link](https://s3.amazonaws.com/fast-ai-imagelocal/biwi_head_pose.tgz).
# %matplotlib inline
from fastai2.basics import *
from fastai2.callback.all import *
from fastai2.vision.all import *
from fastai2.notebook.showdoc import *
# ## Getting and converting the data
path = untar_data(URLs.BIWI_HEAD_POSE)
cal = np.genfromtxt(path/'01'/'rgb.cal', skip_footer=6); cal
fname = '09/frame_00667_rgb.jpg'
def img2txt_name(f): return path/f'{str(f)[:-7]}pose.txt'
img = PILImage.create(path/fname)
img.show();
ctr = np.genfromtxt(img2txt_name(fname), skip_header=3); ctr
# +
def convert_biwi(coords):
c1 = coords[0] * cal[0][0]/coords[2] + cal[0][2]
c2 = coords[1] * cal[1][1]/coords[2] + cal[1][2]
return tensor([c1,c2])
def get_ctr(f):
ctr = np.genfromtxt(img2txt_name(f), skip_header=3)
return convert_biwi(ctr)
def get_ip(img,pts): return TensorPoint.create(pts, sz=img.size)
# -
get_ctr(fname)
ctr = get_ctr(fname)
ax = img.show(figsize=(6, 6))
get_ip(img, ctr).show(ctx=ax);
# ## Creating a dataset
dblock = DataBlock(blocks=(ImageBlock, PointBlock),
get_items=get_image_files,
splitter=FuncSplitter(lambda o: o.parent.name=='13'),
get_y=get_ctr)
dbunch = dblock.databunch(path, path=path, bs=64, batch_tfms=[*aug_transforms(size=(120,160)), Normalize(*imagenet_stats)])
dbunch.show_batch(max_n=9, figsize=(9,6))
# ## Train model
#TODO: look in after_item for c
dbunch.c = dbunch.train_dl.after_item.c
learn = cnn_learner(dbunch, resnet34)
learn.lr_find()
lr = 2e-2
learn.fit_one_cycle(5, slice(lr))
learn.save('stage-1')
learn.load('stage-1');
learn.show_results(max_n=6)
# ## Data augmentation
def repeat_one_file(path):
items = get_image_files(path)
return [items[0]] * 500
dblock = DataBlock(blocks=(ImageBlock, PointBlock),
get_items=repeat_one_file,
splitter=RandomSplitter(),
get_y=get_ctr)
tfms = aug_transforms(max_rotate=20, max_zoom=1.5, max_lighting=0.5, max_warp=0.4, p_affine=1., p_lighting=1., size=(120,160))
dbunch = dblock.databunch(path, path=path, bs=64, batch_tfms=[*tfms, Normalize(*imagenet_stats)])
dbunch.show_batch(max_n=9, figsize=(8,6))
|
dev/course/lesson3-head-pose.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Example Without Render
import gym
import gym_gvgai
[env.id for env in gym.envs.registry.all() if env.id.startswith('gvgai')]
env = gym.make('gvgai-aliens-lvl4-v0')
env.reset()
sum_score = 0
for i in range(1000):
action_id = env.action_space.sample()
state, reward, isOver, debug = env.step(action_id)
sum_score += reward
print('Action: {} Reward: {} SumScore: {} Done: {}'.format(action_id, reward, sum_score, isOver))
if isOver:
print('Game over at game tick {}'.format(i+1))
break
# ## Example With Render
# +
import gym
import gym_gvgai
import matplotlib.pyplot as plt
from IPython import display
def show_state(env, step=0, name="", info=""):
plt.figure(3)
plt.clf()
plt.imshow(env.render(mode='rgb_array'))
plt.title("%s | Step: %d %s" % (name, step, info))
plt.axis('off')
display.clear_output(wait=True)
display.display(plt.gcf())
# -
env = gym.make('gvgai-zelda-lvl0-v0')
# +
# %matplotlib inline
env.reset()
score = 0
for i in range(10):
show_state(env, i, "Aliens", str(score))
#env.render()
state, reward, isOver, debug = env.step(env.action_space.sample())
score += reward
if(isOver):
break
# -
[method_name for method_name in dir(env)]
[method_name for method_name in dir(env.unwrapped)]
state.shape
|
sketch.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Output models
# Class models for different types of output files.
# - OutputRun (files of type output_r00001.csv)
# - OutputMV (files of type output_mv00001.csv)
# - OutputSum (files of type output_sum.csv)
# - OutputSsoln (files of type output_ssoln.csv)
# - OutputSolMat (files of type solutionsmatrix.csv)
#
# I have not created any models for the types *output_sen.dat* or *output_log.dat* because I don't think we need to ingest these data files for any calculation into the database so far.
from pydantic import BaseModel, ValidationError, validator, Field
from pydantic.generics import GenericModel
from typing import Generic, TypeVar, Generic, Optional, Dict, Type
import pandas as pd
# %run marxan_utils.ipynb
MARXAN_FOLDER = '/home/jovyan/work/datasets/raw/Marxan_BLM/BLM_0.001'
InputFile = DatFile(f'{MARXAN_FOLDER}/input.dat')
InputFile.read()
userInputFile = inputDatFile.from_dat(InputFile.data)
# +
class OutputRun(BaseModel):
"""
Class of files _r00001.csv and _best.csv
A file is produced for each repeat run containing a list of all the planning units selected in the solution for that run
"""
# General Parameters
PUID: int = Field(title='Planning Unit id',
description='List of planning units')
SOLUTION: int = Field(title='Solution',
description='Planning unit included in this solution if 1, not included if 0',
ge =0, le =1)
class OutputMV(BaseModel):
"""
Class of files _mv00001.csv or _mvbest.csv
This file contains information about the representation of conservation features in the solution for each run.\
The file contains a total of nine columns which basically report on how the solution performed relative to the targets.\
Some of these are simply a summary of the information provided in the Conservation Feature File
"""
# General Parameters
Conservation_Feature: int = Field(title = 'Conservation feature id',
description ='The unique ID number of the conservation feature')
Feature_Name: str = Field(title = 'Feature Name',
description = 'The optional alphabetic name of the conservation feature.\
If no name has been specified then nothing will appear in this column.')
Target: float = Field(title= 'target',
description = 'The target level of representation (if any) for that conservation feature')
Amount_Held: float = Field(title = 'Amount held',
description = 'The amount of that conservation feature captured in the reserve system')
Occurrence_Target: float = Field(title = 'Occurrence target',
description ='The target number of occurrences in the reserve system for that conservation feature')
Occurrences_Held: float = Field(title= 'Ocurrence s held',
description= 'The number of occurrences of the conservation feature captured in the solution.\
Again, only occurrences in valid clumps are included')
Separation_Target: float = Field (title= 'Separation target',
description = 'The number of mutually and adequately separated occurrences of that\
conservation feature required in the reserve system')
Separation_Achieved: float = Field(title = 'Separation Achieved',
description= 'The number reported here will be the lowest of either: the number of \
separate occurrences that are actually achieved in the reserve system ; or the target \
number of separate occurrences. The separation count (see Appendix B-1.3.1) never exceeds \
the separation target for that feature. This is a convention which speeds up the execution \
of the software but it means that no information is given about how far this target is exceeded.' )
Target_Met: str = Field(title= 'Target met',
description = 'An alphabetic variable that returns ‘yes’ if all the targets set for that feature are met,\
otherwise it returns ‘no’')
MPM: float = Field (title= 'Minimum Proportion Met',
description= 'Propotion of target met, min in 0 max i 1',
ge =0, le =1)
class OutputSum(BaseModel):
"""
Class of files _sum.csv
This file contains the summary information for each repeat run.\
It contains nine columns, which basically report on how the solution performed relative to the targets
"""
Run_Number: int = Field(title='Run number',
description='Which of the repeat runs (or solutions) the output pertains to')
Score: float = Field(title= 'Score',
description ='This is the overall objective function value for the solution from that run.\
This includes not only the cost of the planning units and the boundary length but also the penalties\
for failing to adequately represent all conservation features or exceeding the cost threshold.\
It is useful to know this value because it is how Marxan chooses the ‘best’ solution out of your repeat runs.')
Cost: float = Field(title='Cost',
description = 'This is the total cost of the reserve system as determined solely by the costs given to each planning unit.')
Planning_Units: int = Field(title= 'Planning Units',
description = 'The number of planning units contained in the solution for that run')
Connectivity: float = Field (title ='Connectivity',
description= 'The total boundary length of the reserve system.\
If boundary length is not being considered in the analyses (i.e. no Boundary Length File is provided),\
then this value will read ‘0.0’.')
Connectivity_Total: float = Field (title ='Connectivity Total',
description= 'Total boundary of planning units in study area.')
Connectivity_In:float = Field (title ='Connectivity In',
description= 'Sum of shared boundary between selected planning units.')
Connectivity_Edge:float = Field (title ='Connectivity Edge',
description= 'Same as Connectivity')
Connectivity_Out:float = Field (title ='Connectivity Out',
description= 'Sum of the outer boundaries of unselected planning units.')
Connectivity_In_Fraction: float = Field (title ='Connectivity In Fraction',
description= 'Connectivity_In/Connectivity_Total - the larger this fraction,\
the more spatially compact the solution.')
Penalty: float = Field(title = 'Penalty',
description = 'The penalty that was added to the objective function because the reserve system\
failed to meet the representation targets for all features. If all features are adequately represented\
then the penalty value will be either 0.0 or “-0.0”. (Because of round-off error it is not likely to be\
exactly equal to 0, but with only one decimal place presented the round-off error will probably be hidden).\
The penalty is useful to know because it can give you an idea of the cost required to meet the remaining targets,\
this is something that is not captured simply by looking at the shortfall. It is also another way to rank\
the success of runs, looking only at those solutions that have a low penalty.')
Shortfall: float = Field(title ='Shortfall',
description = 'The amount by which the targets for conservation features have not been met\
in the solution for that run. The shortfall reported here is the total shortfall summed across\
all conservation features. The shortfall is a good indication of whether missing conservation\
features are very close or very far from their targets. If there are a number of conservation\
features which have missed their targets but the combined shortfall is very sma ll then a planner\
might not be too concerned.')
Missing_Values: int = Field (title ='Missing Values',
description= 'The number of features that did not achieve their targets in the final solution for that run.\
This is screened according to the ‘misslevel’, which has been set in the Input Parameter File.\
If the miss level is set to 1 then every conservation feature which falls below its target level\
is counted as missing. If the miss level is set lower than 1 (e.g. 0.98), Marxan may not report a\
feature as missing even if the reserve system contains slightly less than the target amount.')
MPM: float = Field(title ='Minimum Proportion Met',
description= 'The Minimum Proportion Met for the worst performing feature.\
That is, this value corresponds to the lowest MPM value in the missing value file.')
class OutputSsoln(BaseModel):
"""
Class of files _ssoln.csv
Summed solution provides the selection frequency of each planning unit across all runs.\
Each line has the ID number of a planning unit and the number of times\
that planning unit was selected in the final solution across all repeat runs
"""
planning_unit: int = Field(title='Planning Unit',
description= 'ID number of a planning unit')
number: int = Field(title= 'Number',
description= 'Number of times a planning unit was selected in the final solution across all repeat runs')
# OutputSolMat
### Depends on the number of Planning Units:
### Create a class dynamically depending on the number of planning units of the file
### https://stackoverflow.com/questions/62267544/pydantic-generate-model-from-dict
#OutputSolMat = createDynamicModel(filename= filename, name= 'OutputSolMat', dict_def= d)
class OutputSolutionsMatrix(BaseModel):
SolutionsMatrix: str = Field(..., title='Solution number',
description='Solution number')
PU: Dict[str, float] = Field(..., title='Planning unit selection',
description='Pllaning unites selectes in each solution. The dictionary parameters represent:\
key = Planning unit number (P1, P2, P3...), value= selection in this solution (0= False, 1 = True)')
# @validator('SolutionsMatrix')
# # Check that the number of output solutions is the same as in the input.dat file
# def SolutionsMatrix_is_valid(cls, method:str) -> str:
# InputFile = DatFile(f'{MARXAN_FOLDER}/input.dat')
# InputFile.read()
# userInputFile = inputDatFile.from_dat(InputFile.data)
# solNum = userInputFile.NUMREPS
# if len('SolutionsMatrix') != solNum:
# raise ValueError(f"Solutions in input file is {solNum} but got {len(SolutionsMatrix)}")
# return method
# @validator('PU')
# # Check that the number of output planning units is the same as in the pu.dat file
# def PU_is_valid(cls, method:dict) -> dict:
# InputFile = DatFile(f'{MARXAN_FOLDER}/input.dat')
# InputFile.read()
# userInputFile = inputDatFile.from_dat(InputFile.data)
# filename = f'{MARXAN_FOLDER}/{userInputFile.INPUTDIR}/{userInputFile.PUNAME}'
# userPlanningUnits = CreateListModelFromFile(f'{MARXAN_FOLDER}/{userInputFile.INPUTDIR}/{userInputFile.PUNAME}', planningUnits)
# puNum =len(userPlanningUnits)
# if len(PU) != puNum:
# raise ValueError(f"PU in input file is {puNum} but got {len(PU)}")
# return method
# -
# Test OutputRun
filename = f'{MARXAN_FOLDER}/{userInputFile.OUTPUTDIR}/{userInputFile.SCENNAME}_r00001.csv'
CreateListModelFromFile(filename, OutputRun)[0:2]
# Test OutputMV
filename = f'{MARXAN_FOLDER}/{userInputFile.OUTPUTDIR}/{userInputFile.SCENNAME}_mv00001.csv'
CreateListModelFromFile(filename, OutputMV)[0:2]
# Test OutputSum
filename = f'{MARXAN_FOLDER}/{userInputFile.OUTPUTDIR}/{userInputFile.SCENNAME}_sum.csv'
CreateListModelFromFile(filename, OutputSum)[0:2]
# Test OutputSsoln
filename = f'{MARXAN_FOLDER}/{userInputFile.OUTPUTDIR}/{userInputFile.SCENNAME}_ssoln.csv'
CreateListModelFromFile(filename, OutputSsoln)[0:2]
# Test OutputSolMat
filename = f'{MARXAN_FOLDER}/{userInputFile.OUTPUTDIR}/{userInputFile.SCENNAME}_solutionsmatrix.csv'
CreateListModelFromFile(filename, OutputSolutionsMatrix)[0:1]
|
data/notebooks/Lab/marxan_outputs.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Offline Plotting Tutorial
#
# The new dataset comes with a tool for offline (i.e. not live as the data are coming in) plotting. This notebook explains how to use it and what it is capable of plotting.
#
# The tool in question is the function `plot_by_id`.
# +
# %matplotlib notebook
import numpy as np
import qcodes as qc
from typing import List, Dict, Tuple, Any
import matplotlib.pyplot as plt
import qcodes as qc
from qcodes import ParamSpec, new_data_set, new_experiment
from qcodes.dataset.plotting import plot_by_id
from qcodes.dataset.database import initialise_database
# -
# First we make an experimental run, so that we have something to plot.
# if you just deleted your database file, you'll need to init a new one
initialise_database()
new_experiment('test_plot_by_id', 'nosample')
# +
# Make a handful of parameters to be used in the examples
x = ParamSpec('x', 'numeric', label='Voltage', unit='V')
t = ParamSpec('t', 'numeric', label='Time', unit='s')
y = ParamSpec('y', 'numeric', label='Voltage', unit='V', depends_on=[x])
y2 = ParamSpec('y2', 'numeric', label='Current', unit='A', depends_on=[x])
z = ParamSpec('z', 'numeric', label='Majorana number', unit='Anyon', depends_on=[x, t])
# -
# ## A single, simple 1D sweep
# +
data_set = new_data_set('1D-sweep')
data_set.add_parameter(x)
data_set.add_parameter(y)
# +
# %%time
xvals = np.linspace(-3.4, 4.2, 250)
for xnum in xvals:
noise = np.random.randn()*0.1 # multiplicative noise yeah yeah
data_set.add_result({'x': xnum, 'y': 2*(xnum+noise)**3 - 5*(xnum+noise)**2})
data_set.mark_complete()
# -
# Now let us plot that run. The function `plot_by_id` takes the `run_id` of the run to plot as a positional argument. Furthermore, the user may specify the matplotlib axis object (or list of axis objects) to plot on.
# If no axes are specified, the function creates new axis object(s). The function returns a tuple of a list of the axes and a list of the colorbar axes (just `None`s if there are no colorbars).
axes, cbaxes = plot_by_id(data_set.run_id)
# Using the returned axis, we can e.g. change the plot linewidth and color. We refer to the matplotlib documentation for details on matplotlib plot customization.
my_ax = axes[0]
line = my_ax.lines[0]
line.set_color('#223344')
line.set_linewidth(3)
# ## Two interleaved 1D sweeps
#
# Now we make a run where two parameters are measured as a function of the same parameter.
data_set = new_data_set('interleaved-1Ds')
data_set.add_parameter(x)
data_set.add_parameter(y)
data_set.add_parameter(y2)
# +
xvals = np.linspace(-5, 5, 250)
for xnum in xvals:
data_set.add_result({'x': xnum, 'y': xnum**2})
data_set.add_result({'x': xnum, 'y2': -xnum**2})
data_set.mark_complete()
# -
# In such a situation, `plot_by_id` by default creates a new axis for **each** dependent parameter. Sometimes this is not desirable; we'd rather have both plots on the same axis. In such a case, we might pass the same axis twice to `plot_by_id`.
axes, cbaxes = plot_by_id(data_set.run_id)
# Let's do that now
fig, ax = plt.subplots(1)
axes, cbaxes = plot_by_id(data_set.run_id, axes=[ax, ax])
# ## Regular 2D rectangular sweep scan
#
# For 2D plots, a colorbar is usually present. As mentioned above, `plot_by_id` returns this.
data_set = new_data_set('regular-2D-scan')
data_set.add_parameter(x)
data_set.add_parameter(t)
data_set.add_parameter(z)
# +
xvals = np.linspace(-4, 5, 50)
tvals = np.linspace(-500, 1500, 25)
for xv in xvals:
for tv in tvals:
# just some arbitrary semi good looking function
zv = np.sin(2*np.pi*xv)*np.cos(2*np.pi*0.001*tv) + 0.001*tv
data_set.add_result({'x': xv, 't': tv, 'z': zv})
data_set.mark_complete()
# -
axes, colorbars = plot_by_id(data_set.run_id)
# A somewhat normal situation is that the colorbar was somehow mislabelled. Using the returned colorbar, the label can be overwritten.
colorbar = colorbars[0]
colorbar.set_label('Correct science label')
# ## Warped 2D rectangular sweep scan
#
# A nice feature of the new dataset is that the grid may be warped; it makes no difference.
# Here we warp the x axis of the previous scan to increase the resolution in the right half plane.
data_set = new_data_set('warped-2D-scan')
data_set.add_parameter(x)
data_set.add_parameter(t)
data_set.add_parameter(z)
# +
xvals = np.linspace(-4, 5, 50) + np.cos(2/9*np.pi*xvals+np.pi/4)
tvals = np.linspace(-500, 1500, 25)
for xv in xvals:
for tv in tvals:
# just some arbitrary semi good looking function
zv = np.sin(2*np.pi*xv)*np.cos(2*np.pi*0.001*tv) + 0.001*tv
data_set.add_result({'x': xv, 't': tv, 'z': zv})
data_set.mark_complete()
# -
axes, cbaxes = plot_by_id(data_set.run_id)
# ## Interrupted 2D scans (a hole in the cheese)
#
# In case a sweep in interrupted, the entire grid will not be filled out. This is also supported,
# in fact, any single rectangular hole is allowed
data_set = new_data_set('warped-with-hole-2D-scan')
data_set.add_parameter(x)
data_set.add_parameter(t)
data_set.add_parameter(z)
# +
xvals = np.linspace(-4, 5, 50) + np.cos(2/9*np.pi*xvals+np.pi/4)
tvals = np.linspace(-500, 1500, 25)
# define two small forbidden range functions
def no_x(xv):
if xv > 0 and xv < 3:
return True
else:
return False
def no_t(tv):
if tv > 0 and tv < 450:
return True
else:
return False
for xv in xvals:
for tv in tvals:
if no_x(xv) and no_t(tv):
continue
else:
# just some arbitrary semi good looking function
zv = np.sin(2*np.pi*xv)*np.cos(2*np.pi*0.001*tv) + 0.001*tv
data_set.add_result({'x': xv, 't': tv, 'z': zv})
data_set.mark_complete()
# -
axes, colorbars = plot_by_id(data_set.run_id)
# ## Fancy plotting
# As a final example, let us combine several plots in one window.
#
# We first make a little grid of axes.
fig, figaxes = plt.subplots(2, 2)
# Next, we make some runs (shamelessly copy-pasting from above).
# +
# First run
data_set = new_data_set('1D-sweep')
data_set.add_parameter(x)
data_set.add_parameter(y)
xvals = np.linspace(-3.4, 4.2, 250)
for xnum in xvals:
noise = np.random.randn()*0.1 # multiplicative noise yeah yeah
data_set.add_result({'x': xnum, 'y': 2*(xnum+noise)**3 - 5*(xnum+noise)**2})
data_set.mark_complete()
rid1 = data_set.run_id
# Second run
data_set = new_data_set('2D-sweep')
data_set.add_parameter(x)
data_set.add_parameter(t)
data_set.add_parameter(z)
xvals = np.linspace(-4, 5, 50)
tvals = np.linspace(-500, 1500, 25)
for xv in xvals:
for tv in tvals:
# just some arbitrary semi good looking function
zv = np.sin(2*np.pi*xv)*np.cos(2*np.pi*0.001*tv) + 0.001*tv
data_set.add_result({'x': xv, 't': tv, 'z': zv})
data_set.mark_complete()
rid2 = data_set.run_id
# -
# And then we put them just where we please.
axes, colorbars = plot_by_id(rid1, figaxes[0, 0])
axes, colorbars = plot_by_id(rid2, figaxes[1, 1], colorbars)
# Note that if we want to replot on an axis with a colorbar we probably also want to reuse the colorbar
axes, colorbars = plot_by_id(rid2, figaxes[1, 1], colorbars)
fig.tight_layout()
|
docs/examples/DataSet/Offline Plotting Tutorial.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=true editable=true
# # Import required packages
# + deletable=true editable=true
import os
import math
import glob
import cv2
from collections import deque
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
from moviepy.editor import VideoFileClip
# %matplotlib inline
# + [markdown] deletable=true editable=true
# # Create a utility class for camera calibration
#
# * This is used for calibrating camera and undistorting the images
# + deletable=true editable=true
class cam_util():
"""
util class for camera operations
"""
ret = None
mtx = None
dist = None
rvecs = None
tvecs = None
# Arrays to store object points and image points from all the images.
objpoints = [] # 3d points in real world space
imgpoints = [] # 2d points in image plane.
def gen_camera_points(self):
"""
generate objpoints and impoints from calibration images
"""
objp = np.zeros((6*9,3), np.float32)
objp[:,:2] = np.mgrid[0:9,0:6].T.reshape(-1,2)
# Make a list of calibration images
images = glob.glob('camera_cal/calibration*.jpg')
# Step through the list and search for chessboard corners
for fname in images:
img = cv2.imread(fname)
gray = cv2.cvtColor(img,cv2.COLOR_BGR2GRAY)
# Find the chessboard corners
ret, corners = cv2.findChessboardCorners(gray, (9,6),None)
# If found, add object points, image points
if ret == True:
self.objpoints.append(objp)
self.imgpoints.append(corners)
def undistort(self, img):
"""
undistort an image with camera matrix
"""
if self.mtx is None:
self.ret, self.mtx, self.dist, self.rvecs, self.tvecs = cv2.calibrateCamera(self.objpoints, self.imgpoints,
img.shape[:2],None,None)
h, w = img.shape[:2]
newcameramtx, roi=cv2.getOptimalNewCameraMatrix(self.mtx, self.dist, (w,h), 1, (w,h))
dst = cv2.undistort(img, self.mtx, self.dist, None, newcameramtx)
x,y,w,h = roi
return dst[y:y+h, x:x+w]
def clean_mat(self):
"""
Reset camera calibration
"""
self.ret = None
self.mtx = None
self.dist = None
self.rvecs = None
self.tvecs = None
# + [markdown] deletable=true editable=true
# # Create a class to keep track of lane detections
#
# * Here we use the average of last maxSamples to identify the lane
# + deletable=true editable=true
class Line():
"""
class to store detected lane stats
"""
def __init__(self, maxSamples=15):
self.maxSamples = maxSamples
# x values of the last n fits of the line
self.recent_xfitted = deque(maxlen=self.maxSamples)
#polynomial coefficients for the most recent fit
self.current_fit = [np.array([False])]
#polynomial coefficients averaged over the last n iterations
self.best_fit = None
#difference in fit coefficients between last and new fits
self.diffs = np.array([0,0,0], dtype='float')
#average x values of the fitted line over the last n iterations
self.bestx = None
# was the line detected in the last iteration?
self.detected = False
#radius of curvature of the line in some units
self.radius_of_curvature = None
#distance in meters of vehicle center from the line
self.line_base_pos = None
def update_lane(self, ally, allx):
"""
Function to update the stats
"""
# get the mean as the best x
self.bestx = np.mean(allx, axis=0)
# fit a 2 order polynomial
new_fit = np.polyfit(ally, allx, 2)
# calculate the difference between last fit and new fit
self.diffs = np.subtract(self.current_fit, new_fit)
# update current fit
self.current_fit = new_fit
# add the new fit to the queue
self.recent_xfitted.append(self.current_fit)
# Use the queue mean as the best fit
self.best_fit = np.mean(self.recent_xfitted, axis=0)
# meters per pixel in y dimension
ym_per_pix = 30/720
# meters per pixel in x dimension
xm_per_pix = 3.7/700
# Calculate radius of curvature
fit_cr = np.polyfit(ally*ym_per_pix, allx*xm_per_pix, 2)
y_eval = np.max(ally)
self.radius_of_curvature = ((1 + (2*fit_cr[0]*y_eval*ym_per_pix + fit_cr[1])**2)**1.5) / np.absolute(2*fit_cr[0])
# + deletable=true editable=true
# Utility Functions
def get_roi(img, vertices):
"""
Apply mask and get region of interest within the mask
"""
mask = np.zeros_like(img)
if len(img.shape) > 2:
channel_count = img.shape[2]
ignore_mask_color = (255,) * channel_count
else:
ignore_mask_color = 255
cv2.fillPoly(mask, vertices, ignore_mask_color)
masked_image = cv2.bitwise_and(img, mask)
return masked_image
def hide_roi(img, vertices):
"""
Apply mask and get region of interest outside the mask
"""
mask = np.zeros_like(img)
mask=mask+255
if len(img.shape) > 2:
channel_count = img.shape[2]
ignore_mask_color = (0,) * channel_count
else:
ignore_mask_color = 0
cv2.fillPoly(mask, vertices, ignore_mask_color)
masked_image = cv2.bitwise_and(img, mask)
return masked_image
def drow_on_images(img, vertices):
"""
Draw ploygon on image
"""
cv2.polylines(img, [vertices], True, (255,255,255), 2)
plot_img(img, 'img drawing', True)
def plot_img(img, step, show_stages=False):
"""
plot image
"""
if show_stages:
print('######################## '+step+' ########################')
plt.imshow(img, cmap='gray')
plt.show()
def plot_hist(histogram, show_stages=False):
"""
plot histogram
"""
if show_stages:
print('######################## histogram ########################')
plt.plot(histogram)
plt.show()
# + [markdown] deletable=true editable=true
# # Use the lane pixals identified to fit a ploygon and draw it back on the original image
# + deletable=true editable=true
def write_stats(img):
"""
Write lane stats on image
"""
font = cv2.FONT_HERSHEY_SIMPLEX
size = 1
weight = 2
color = (255,70,0)
cv2.putText(img,'Left Curve : '+ '{0:.2f}'.format(left_line.radius_of_curvature)+' m',(10,30), font, size, color, weight)
cv2.putText(img,'Right Curve : '+ '{0:.2f}'.format(right_line.radius_of_curvature)+' m',(10,60), font, size, color, weight)
cv2.putText(img,'Left Lane Pos: '+ '{0:.2f}'.format(left_line.bestx),(10,100), font, size, color, weight)
cv2.putText(img,'Right Lane Pos: '+ '{0:.2f}'.format(right_line.bestx),(10,130), font, size, color, weight)
cv2.putText(img,'Distance from center: '+ "{0:.2f}".format(left_line.line_base_pos)+' m',(10,180), font, size, color, weight)
def draw_lane(undist, img, Minv):
"""
Draw the detected lane bak on the image
"""
# Generate x and y values for plotting
ploty = np.linspace(300, 700)
# Create an image to draw the lines on
warp_zero = np.zeros_like(img).astype(np.uint8)
color_warp = np.dstack((warp_zero, warp_zero, warp_zero))
left_fit = left_line.best_fit
right_fit = right_line.best_fit
if left_fit is not None and right_fit is not None:
left_fitx = left_fit[0]*ploty**2 + left_fit[1]*ploty + left_fit[2]
# Recast the x and y points into usable format for cv2.fillPoly()
pts_left = np.array([np.transpose(np.vstack([left_fitx, ploty]))])
right_fitx = right_fit[0]*ploty**2 + right_fit[1]*ploty + right_fit[2]
pts_right = np.array([np.flipud(np.transpose(np.vstack([right_fitx, ploty])))])
pts = np.hstack((pts_left, pts_right))
# Draw the lane onto the warped blank image
cv2.fillPoly(color_warp, np.int_([pts]), (20,120, 80))
# Warp the blank back to original image space using inverse perspective matrix (Minv)
newwarp = cv2.warpPerspective(color_warp, Minv, (img.shape[1], img.shape[0]))
# Combine the result with the original image
result = cv2.addWeighted(undist, 1, newwarp, 0.3, 0)
write_stats(result)
return result
return undist
# + [markdown] deletable=true editable=true
# # Here we validate the detected lines and add them to the lane class
#
# ## A valid detection satisfies below rules
#
# * Minmum number of pixals must be greater than 2000
# * Left lane mean should be more than a minimum
# * Right lane mean should be less then a minimum
# * Lane width whoud be atlest 300 and atmost 800
# * New detections must be within 100px of the average of last n detections
# + deletable=true editable=true
def validate_Update_lane(img, nonzero, nonzerox, nonzeroy, left_lane_inds, right_lane_inds, show_stages=False):
"""
Validate the detected lane ids and update the lane stats if valid.
"""
# Extract left and right line pixel positions
left_line_allx = nonzerox[left_lane_inds]
left_line_ally = nonzeroy[left_lane_inds]
right_line_allx = nonzerox[right_lane_inds]
right_line_ally = nonzeroy[right_lane_inds]
# Discard the detections if any of the detected lane is less than 2000 pixals.
# This is done because for very small size the poly fit function gives unpredictable results.
# A better approch would be to use the largest lane curvature to extend the other one
if len(left_line_allx) <= 2000 or len(right_line_allx) <= 2000:
left_line.detected = False
right_line.detected = False
return
left_x_mean = np.mean(left_line_allx, axis=0)
right_x_mean = np.mean(right_line_allx, axis=0)
lane_width = np.subtract(right_x_mean, left_x_mean)
# Discard the detections if the lane with is too large or too small
if left_x_mean > 450 or right_x_mean < 850:
left_line.detected = False
right_line.detected = False
return
if lane_width < 300 or lane_width > 800:
left_line.detected = False
right_line.detected = False
return
# Update the lane stats if the current detection is the first one or
# the detection is within 100 pixals of the last n detection mean
if left_line.bestx is None or np.abs(np.subtract(left_line.bestx, np.mean(left_line_allx, axis=0))) < 100:
left_line.update_lane(left_line_ally, left_line_allx)
left_line.detected = True
else:
left_line.detected = False
if right_line.bestx is None or np.abs(np.subtract(right_line.bestx, np.mean(right_line_allx, axis=0))) < 100:
right_line.update_lane(right_line_ally, right_line_allx)
right_line.detected = True
else:
right_line.detected = False
# Calculate the distance of car from center of lane
lane_center = right_line.bestx - left_line.bestx
left_line.line_base_pos = ((img.shape[1]*0.5 - lane_center)*3.7)/700
right_line.line_base_pos = left_line.line_base_pos
# + [markdown] deletable=true editable=true
# # Find the lane using sliding window technique
#
# * Use the minimum of bottom 1/4 of the histogram to find the initial left and right base
# * Use the base points to find more points within a margin and min number of pixals
# * Using
# * windows size = 9
# * margin = 80
# * min pixals = 30
# + deletable=true editable=true
def window_search(img, nonzero, nonzerox, nonzeroy, show_stages=False):
"""
Perform a sliding window search to detect lane pixals.
"""
# Temp image to draw detections on
out_img = np.dstack((img, img, img))*255
# Calculate histogram
histogram = np.sum(img[img.shape[0]*.75:,:], axis=0)
plot_hist(histogram, show_stages)
# Take the midpoint and use the max on each side as starting point
midpoint = np.int(histogram.shape[0]/2)
leftx_base = np.argmax(histogram[0:midpoint])
rightx_base = np.argmax(histogram[midpoint:histogram.shape[0]]) + midpoint
# Choose the number of sliding windows
nwindows = 9
# Set height of windows
window_height = np.int(img.shape[0]/nwindows)
# Current positions to be updated for each window
leftx_current = leftx_base
rightx_current = rightx_base
# Set the width of the windows +/- margin
margin = 80
# Set minimum number of pixels found to recenter window
minpix = 30
# Create empty lists to receive left and right lane pixel indices
left_lane_inds = []
right_lane_inds = []
# Step through the windows one by one
for window in range(nwindows):
# Identify window boundaries in x and y (and right and left)
win_y_low = img.shape[0] - (window+1)*window_height
win_y_high = img.shape[0] - window*window_height
win_xleft_low = leftx_current - margin
win_xleft_high = leftx_current + margin
win_xright_low = rightx_current - margin
win_xright_high = rightx_current + margin
# Draw the windows on the visualization image
cv2.rectangle(out_img,(win_xleft_low,win_y_low),(win_xleft_high,win_y_high),(0,255,0), 2)
cv2.rectangle(out_img,(win_xright_low,win_y_low),(win_xright_high,win_y_high),(0,255,0), 2)
# Identify the nonzero pixels in x and y within the window
good_left_inds = ((nonzeroy >= win_y_low)
& (nonzeroy < win_y_high)
& (nonzerox >= win_xleft_low)
& (nonzerox < win_xleft_high)).nonzero()[0]
good_right_inds = ((nonzeroy >= win_y_low)
& (nonzeroy < win_y_high)
& (nonzerox >= win_xright_low)
& (nonzerox < win_xright_high)).nonzero()[0]
left_lane_inds.append(good_left_inds)
right_lane_inds.append(good_right_inds)
# If found > minpix pixels, recenter next window on their mean position
if len(good_left_inds) > minpix:
leftx_current = np.int(np.mean(nonzerox[good_left_inds]))
if len(good_right_inds) > minpix:
rightx_current = np.int(np.mean(nonzerox[good_right_inds]))
# Concatenate the arrays of indices
left_lane_inds = np.concatenate(left_lane_inds)
right_lane_inds = np.concatenate(right_lane_inds)
plot_img(out_img, 'sliding window marked', show_stages)
return left_lane_inds, right_lane_inds
# + [markdown] deletable=true editable=true
# # Find Lanes Wrapper
#
# * If left or right lane found in the last iteration. Get the pixals in a margin of 30 and validate
#
# * If the validation fails or this is the first iteration use the sliding window technique to find lanes and then validate.
# + deletable=true editable=true
def find_lanes(img, show_stages=False):
"""
Lane finding wrapper function
"""
# Get the foreground pixals
nonzero = img.nonzero()
nonzeroy = np.array(nonzero[0])
nonzerox = np.array(nonzero[1])
# If the last detection was successful take the non zero pixals within the 30 pixal margin as the new detections
if left_line.detected and right_line.detected:
margin = 30
left_lane_inds = ((nonzerox > (left_line.current_fit[0]*(nonzeroy**2) + left_line.current_fit[1]*nonzeroy + left_line.current_fit[2] - margin))
& (nonzerox < (left_line.current_fit[0]*(nonzeroy**2) + left_line.current_fit[1]*nonzeroy + left_line.current_fit[2] + margin)))
right_lane_inds = ((nonzerox > (right_line.current_fit[0]*(nonzeroy**2) + right_line.current_fit[1]*nonzeroy + right_line.current_fit[2] - margin))
& (nonzerox < (right_line.current_fit[0]*(nonzeroy**2) + right_line.current_fit[1]*nonzeroy + right_line.current_fit[2] + margin)))
# Update the lane detections
validate_Update_lane(img, nonzero, nonzerox, nonzeroy, left_lane_inds, right_lane_inds)
# If first detection or the last detection was unsuccessful perform a sliding window search
else:
#print('doing window search')
left_lane_inds, right_lane_inds = window_search(img, nonzero, nonzerox, nonzeroy, show_stages)
# Update the lane detections
validate_Update_lane(img, nonzero, nonzerox, nonzeroy, left_lane_inds, right_lane_inds)
# + [markdown] deletable=true editable=true
# # Warp the image to get birds' eye view
#
# * Use source points
# * bounding_top_right = [img_shape[1]*0.5 + 90,img_shape[0]*0.70]
# * bounding_btm_right = [img_shape[1]*0.5 + 450,img_shape[0]]
# * bounding_btm_left = [img_shape[1]*0.5 - 400,img_shape[0]]
# * bounding_top_left = [img_shape[1]*0.5 - 60,img_shape[0]*0.70]
#
# * Destinations points
# * bounding_top_right = [img_shape[1]*0.5 + 250,img_shape[0]*0.60]
# * bounding_btm_right = [img_shape[1]*0.5 + 390,img_shape[0]]
# * bounding_btm_left = [img_shape[1]*0.5 - 345,img_shape[0]]
# * bounding_top_left = [img_shape[1]*0.5 - 205,img_shape[0]*0.60]
#
# * Get perpective transform
# * Get inverse perpective transform
# * warp the image using perspective transform
#
# + deletable=true editable=true
def warp(img):
"""
Warp the image to get birds eye view.
"""
img_shape = img.shape
bounding_top_right = [img_shape[1]*0.5 + 90,img_shape[0]*0.70]
bounding_btm_right = [img_shape[1]*0.5 + 450,img_shape[0]]
bounding_btm_left = [img_shape[1]*0.5 - 400,img_shape[0]]
bounding_top_left = [img_shape[1]*0.5 - 60,img_shape[0]*0.70]
# Select source points
pts1 = np.float32([bounding_top_right,bounding_btm_right,bounding_btm_left,bounding_top_left])
# Select destination points
pts2 = np.float32([[img_shape[1]*0.5 + 250,img_shape[0]*0.60],
[img_shape[1]*0.5 + 390,img_shape[0]],
[img_shape[1]*0.5 - 345,img_shape[0]],
[img_shape[1]*0.5 - 205,img_shape[0]*0.60]])
# Get Perspective Transform
M = cv2.getPerspectiveTransform(pts1, pts2)
# Get inverse Perspective Transform
Minv = cv2.getPerspectiveTransform(pts2, pts1)
# Apply warp transform on source image
dst = cv2.warpPerspective(img, M, (img.shape[1], img.shape[0]), flags=cv2.INTER_LINEAR)
return dst, Minv
# + [markdown] deletable=true editable=true
# # Threshold
#
# * Use color threshold
# * The number of lane pixals must be considerably less than the background pixals and have a minimum value.
# * We use this to recursively increase or decrease the minimum threshold value to find the optimal value.
# * Use Sobel operator to find gradients
# * Combine the two to get the result
# + deletable=true editable=true
def rec_threshold(img, roi, t_min=140, t_max=255):
"""
Funtion to apply recursive threshold with increasing/decreasing boundries
based on the area of lane within a region of interest.
"""
binary = np.zeros_like(img)
binary[(img >= t_min) & (img <= t_max)] = 1
# retrun last val if the threshold levels reach minimum or maximum.
if t_min <= 40 or t_min >= 220:
return binary
binary_1 = get_roi(binary, roi)
#print(np.sum(binary_1.nonzero()))
if np.sum(binary_1.nonzero()) > 9800000:
binary = rec_threshold(img, roi, t_min+10)
elif np.sum(binary_1.nonzero()) < 100000:
binary = rec_threshold(img, roi, t_min-10)
return binary
def threshold(img, roi, show_stages=False):
"""
Apply threshold
"""
# Convert image the HSV
hsv = cv2.cvtColor(img, cv2.COLOR_RGB2HSV)
# Take v channel
v_channel = hsv[:,:,2]
plot_img(v_channel, 'v channel', show_stages)
# Apply threshold to find lane
v_binary = rec_threshold(v_channel, roi)
plot_img(v_binary, 'color threshold', show_stages)
# Convert image to grayscale
gray = cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
# Take the derivative in x
sobelx = cv2.Sobel(gray, cv2.CV_32F, 1, 0)
#sobelx = cv2.Sobel(sobelx, cv2.CV_32F, 0, 1) # Take the derivative
#plot_img(sobelx, show_stages)
# Absolute x derivative to
abs_sobelx = np.absolute(sobelx)
#accentuate lines away from horizontal
scaled_sobel = np.uint8(255*abs_sobelx/np.max(abs_sobelx))
#plot_img(sobelx, show_stages)
sxbinary = np.zeros_like(scaled_sobel)
# perform threshold
sxbinary[(scaled_sobel >= 100) & (scaled_sobel <= 255)] = 1
plot_img(sobelx, 'sobel', show_stages)
color_binary = np.dstack(( np.zeros_like(sxbinary), sxbinary, v_binary))
combined_binary = np.zeros_like(sxbinary)
# conbine color and sobel threshold
combined_binary[(v_binary == 1) | (sxbinary == 1)] = 1
plot_img(combined_binary, 'combined threshold', show_stages)
return combined_binary
# + [markdown] deletable=true editable=true
# # Apply all the steps
#
# * Undistort the image
# * Apply perspective transform
# * Apply threshold
# * Find lanes
# * Draw the result back on image
# + deletable=true editable=true
def process_image(image, show_stages=False):
"""
Wrapper function for all image processing
"""
# Undistort the image
undistorted = cam.undistort(image)
plot_img(undistorted, 'undistorted', show_stages)
# Apply perpective transform
img, Minv = warp(undistorted)
plot_img(img, 'warped', show_stages)
# Get points for region of interst
vertices = np.array([[(image.shape[1]*0.1,image.shape[0]-50),
(image.shape[1]*0.5-100,image.shape[0]*0.60),
(image.shape[1]*0.5+100,image.shape[0]*0.60),
(image.shape[1]*0.95,image.shape[0]-50)]],
dtype=np.int32)
# Apply threshold
img = threshold(img, vertices, show_stages)
vertices = np.array([[(200,img.shape[0]),
(200,0),
(1050,0),
(1050,img.shape[0])]], dtype=np.int32)
# Get roi
img = get_roi(img, vertices)
# Find Lanes
find_lanes(img, show_stages)
# Draw lanes on image
res = draw_lane(undistorted, img, Minv);
#plot_img(res, show_stages)
return res
# + [markdown] deletable=true editable=true
# # Generate obj points and img points
# + deletable=true editable=true
# init camera
cam = cam_util()
cam.gen_camera_points()
# + [markdown] deletable=true editable=true
# # Calibrate camera and undistort the chessbaord images
# + deletable=true editable=true
# Undistort a sample calibration image
cal_dir = "camera_cal/"
cal_images = glob.glob(cal_dir+'*.jpg')
for cal_image in cal_images:
cimg = mpimg.imread(cal_image)
cimg_undistort = cam.undistort(cimg)
cv2.imwrite('output_images/undistort_'+cal_image.split('/')[1],cimg_undistort)
print('calibration done')
# + deletable=true editable=true
# Clean camera matrix
cam.clean_mat()
# + [markdown] deletable=true editable=true
# # Test on images
# + deletable=true editable=true
# Test on images
test_dir = "test_images/"
test_images = glob.glob(test_dir+'test*.jpg')
#test_images = glob.glob(test_dir+'straight_lines*.jpg')
#test_images = glob.glob(test_dir+'*.jpg')
for test_image in test_images:
left_line = Line()
right_line = Line()
image = mpimg.imread(test_image)
res = process_image(image, False)
#plot_img(res, True)
print('######################## Sample Stages ########################')
print()
# display stages for a sample image
left_line = Line()
right_line = Line()
image = mpimg.imread('test_images/test3.jpg')
plot_img(image, 'Initial', True)
res = process_image(image, True)
plot_img(res, 'Final', True)
# + [markdown] deletable=true editable=true
# # Test on videos
# + deletable=true editable=true
# Test on Videos
# Clean data for video
#"""
left_line = Line()
right_line = Line()
cam.clean_mat()
project_video_res = 'project_video_res.mp4'
clip1 = VideoFileClip("project_video.mp4")
project_video_clip = clip1.fl_image(process_image)
project_video_clip.write_videofile(project_video_res, audio=False)
#"""
# Clean data for video
#"""
left_line = Line()
right_line = Line()
cam.clean_mat()
challenge_video_res = 'challenge_video_res.mp4'
clip2 = VideoFileClip('challenge_video.mp4')
challenge_video_clip = clip2.fl_image(process_image)
challenge_video_clip.write_videofile(challenge_video_res, audio=False)
#"""
# Clean data for video
#"""
left_line = Line()
right_line = Line()
cam.clean_mat()
harder_challenge_video_res = 'harder_challenge_video_res.mp4'
clip2 = VideoFileClip('harder_challenge_video.mp4')
harder_challenge_video_clip = clip2.fl_image(process_image)
harder_challenge_video_clip.write_videofile(harder_challenge_video_res, audio=False)
#"""
# + deletable=true editable=true
|
.ipynb_checkpoints/car-lane-detection-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # December 2017: Advent of Code Solutions
#
# <NAME>
#
# I'm doing the [Advent of Code](https://adventofcode.com) puzzles, just like [last year](https://github.com/norvig/pytudes/blob/master/ipynb/Advent%20of%20Code.ipynb). My terms of engagement are:
#
# * You'll need to follow the links in the section headers (e.g. **[Day 1](https://adventofcode.com/2017/day/1)**) to understand what each puzzle is asking; I won't repeat the puzzle description.
# * What you see is mostly the algorithm I came up with first, although sometimes I go back and refactor for clarity.
# * I'll clean up the code a bit: adding docstrings, making variable names longer and more descriptive, adding `assert` statements.
# * I will discuss any errors I made along the way; usually I won't show the erroneous code, just a description of what I did wrong.
# * The way Advent of Code works is that you read the puzzle descriotion for Part One, but only when you correctly solve it do you get to see Part Two. This is typical in software development: you deploy some code, and then some new requirements arise. So it makes sense to program by creating small functions and data types that form a *vocabulary* for the domain at hand, and can be recombined to solve new problems in the domain.
# * Each day's code should run in a few seconds; certainly less than a minute. (As it turns out, the total run time for all my solutions was just under a minute.)
# * There is a contest to see who can solve each day's puzzle fastest; I do not expect to be competitive.
#
#
#
# # Day 0: Imports and Utility Functions
#
# I might need these:
# +
# Python 3.x Utility Functions
# %matplotlib inline
import matplotlib.pyplot as plt
import os
import urllib.request
import re
import numpy as np
import math
import random
import time
from collections import Counter, defaultdict, namedtuple, deque, abc, OrderedDict
from functools import lru_cache
from statistics import mean, median, mode, stdev, variance
from itertools import (permutations, combinations, chain, cycle, product, islice,
takewhile, zip_longest, count as count_from)
from heapq import heappop, heappush
from numba import jit
letters = '<KEY>'
cache = lru_cache(None)
cat = ''.join
Ø = frozenset() # Empty set
inf = float('inf')
BIG = 10 ** 999
################ Functions for Input, Parsing
def Input(day, year=2017):
"Open this day's input file."
directory = 'advent{}/'.format(year)
filename = directory+'input{}.txt'.format(day)
try:
return open(filename)
except FileNotFoundError:
if not os.path.exists(directory):
os.makedirs(directory)
urllib.request.urlretrieve("https://raw.githubusercontent.com/norvig/pytudes/master/data/" + filename, filename)
return Input(day)
def Inputstr(day, year=2017):
"The contents of this day's input file as a str."
return Input(day, year).read().rstrip('\n')
def Array(lines):
"Parse an iterable of str lines into a 2-D array. If `lines` is a str, splitlines."
if isinstance(lines, str): lines = lines.splitlines()
return mapt(Vector, lines)
def Vector(line):
"Parse a str into a tuple of atoms (numbers or str tokens)."
return mapt(Atom, line.replace(',', ' ').split())
def Integers(text):
"Return a tuple of all integers in a string."
return mapt(int, re.findall(r'-?\b\d+\b', text))
def Atom(token):
"Parse a str token into a number, or leave it as a str."
try:
return int(token)
except ValueError:
try:
return float(token)
except ValueError:
return token
def error(err=RuntimeError, *args): raise err(*args)
################ Functions on Iterables
def first(iterable, default=None):
"The first item in an iterable, or default if it is empty."
return next(iter(iterable), default)
def first_true(iterable, pred=None, default=None):
"""Returns the first true value in the iterable.
If no true value is found, returns *default*
If *pred* is not None, returns the first item
for which pred(item) is true."""
# first_true([a,b,c], default=x) --> a or b or c or x
# first_true([a,b], fn, x) --> a if fn(a) else b if fn(b) else x
return next(filter(pred, iterable), default)
def nth(iterable, n, default=None):
"Returns the nth item of iterable, or a default value"
return next(islice(iterable, n, None), default)
def upto(iterable, maxval):
"From a monotonically increasing iterable, generate all the values <= maxval."
# Why <= maxval rather than < maxval? In part because that's how Ruby's upto does it.
return takewhile(lambda x: x <= maxval, iterable)
identity = lambda x: x
def groupby(iterable, key=identity):
"Return a dict of {key(item): [items...]} grouping all items in iterable by keys."
groups = defaultdict(list)
for item in iterable:
groups[key(item)].append(item)
return groups
def grouper(iterable, n, fillvalue=None):
"""Collect data into fixed-length chunks:
grouper('ABCDEFG', 3, 'x') --> ABC DEF Gxx"""
args = [iter(iterable)] * n
return zip_longest(*args, fillvalue=fillvalue)
def overlapping(iterable, n):
"""Generate all (overlapping) n-element subsequences of iterable.
overlapping('ABCDEFG', 3) --> ABC BCD CDE DEF EFG"""
if isinstance(iterable, abc.Sequence):
yield from (iterable[i:i+n] for i in range(len(iterable) + 1 - n))
else:
result = deque(maxlen=n)
for x in iterable:
result.append(x)
if len(result) == n:
yield tuple(result)
def pairwise(iterable):
"s -> (s0,s1), (s1,s2), (s2, s3), ..."
return overlapping(iterable, 2)
def sequence(iterable, type=tuple):
"Coerce iterable to sequence: leave alone if already a sequence, else make it `type`."
return iterable if isinstance(iterable, abc.Sequence) else type(iterable)
def join(iterable, sep=''):
"Join the items in iterable, converting each to a string first."
return sep.join(map(str, iterable))
def powerset(iterable):
"Yield all subsets of items."
items = list(iterable)
for r in range(len(items)+1):
for c in combinations(items, r):
yield c
def quantify(iterable, pred=bool):
"Count how many times the predicate is true."
return sum(map(pred, iterable))
def length(iterable):
"Same as len(list(iterable)), but without consuming memory."
return sum(1 for _ in iterable)
def shuffled(iterable):
"Create a new list out of iterable, and shuffle it."
new = list(iterable)
random.shuffle(new)
return new
flatten = chain.from_iterable
################ Functional programming
def mapt(fn, *args):
"Do a map, and make the results into a tuple."
return tuple(map(fn, *args))
def map2d(fn, grid):
"Apply fn to every element in a 2-dimensional grid."
return tuple(mapt(fn, row) for row in grid)
def repeat(n, fn, arg, *args, **kwds):
"Repeat arg = fn(arg) n times, return arg."
return nth(repeatedly(fn, arg, *args, **kwds), n)
def repeatedly(fn, arg, *args, **kwds):
"Yield arg, fn(arg), fn(fn(arg)), ..."
yield arg
while True:
arg = fn(arg, *args, **kwds)
yield arg
def compose(f, g):
"The function that computes f(g(x))."
return lambda x: f(g(x))
################ Making immutable objects
class Set(frozenset):
"A frozenset, but with a prettier printer."
def __repr__(self): return '{' + join(sorted(self), ', ') + '}'
def canon(items, typ=None):
"Canonicalize these order-independent items into a hashable canonical form."
typ = typ or (cat if isinstance(items, str) else tuple)
return typ(sorted(items))
################ Math Functions
def transpose(matrix): return tuple(zip(*matrix))
def isqrt(n):
"Integer square root (rounds down)."
return int(n ** 0.5)
def ints(start, end, step=1):
"The integers from start to end, inclusive: range(start, end+1)"
return range(start, end + 1, step)
def floats(start, end, step=1.0):
"Yield floats from start to end (inclusive), by increments of step."
m = (1.0 if step >= 0 else -1.0)
while start * m <= end * m:
yield start
start += step
def multiply(numbers):
"Multiply all the numbers together."
result = 1
for n in numbers:
result *= n
return result
import operator as op
operations = {'>': op.gt, '>=': op.ge, '==': op.eq,
'<': op.lt, '<=': op.le, '!=': op.ne,
'+': op.add, '-': op.sub, '*': op.mul,
'/': op.truediv, '**': op.pow}
################ 2-D points implemented using (x, y) tuples
def X(point): return point[0]
def Y(point): return point[1]
origin = (0, 0)
HEADINGS = UP, LEFT, DOWN, RIGHT = (0, -1), (-1, 0), (0, 1), (1, 0)
def turn_right(heading): return HEADINGS[HEADINGS.index(heading) - 1]
def turn_around(heading):return HEADINGS[HEADINGS.index(heading) - 2]
def turn_left(heading): return HEADINGS[HEADINGS.index(heading) - 3]
def add(A, B):
"Element-wise addition of two n-dimensional vectors."
return mapt(sum, zip(A, B))
def neighbors4(point):
"The four neighboring squares."
x, y = point
return ( (x, y-1),
(x-1, y), (x+1, y),
(x, y+1))
def neighbors8(point):
"The eight neighboring squares."
x, y = point
return ((x-1, y-1), (x, y-1), (x+1, y-1),
(x-1, y), (x+1, y),
(x-1, y+1), (x, y+1), (x+1, y+1))
def cityblock_distance(P, Q=origin):
"Manhatten distance between two points."
return sum(abs(p - q) for p, q in zip(P, Q))
def distance(P, Q=origin):
"Straight-line (hypotenuse) distance between two points."
return sum((p - q) ** 2 for p, q in zip(P, Q)) ** 0.5
def king_distance(P, Q=origin):
"Number of chess King moves between two points."
return max(abs(p - q) for p, q in zip(P, Q))
################ Debugging
def trace1(f):
"Print a trace of the input and output of a function on one line."
def traced_f(*args):
result = f(*args)
print('{}({}) = {}'.format(f.__name__, ', '.join(map(str, args)), result))
return result
return traced_f
def grep(pattern, iterable):
"Print lines from iterable that match pattern."
for line in iterable:
if re.search(pattern, line):
print(line)
class Struct:
"A structure that can have any fields defined."
def __init__(self, **entries): self.__dict__.update(entries)
def __repr__(self):
fields = ['{}={}'.format(f, self.__dict__[f])
for f in sorted(self.__dict__)]
return 'Struct({})'.format(', '.join(fields))
################ A* and Breadth-First Search (tracking states, not actions)
def always(value): return (lambda *args: value)
def Astar(start, moves_func, h_func, cost_func=always(1)):
"Find a shortest sequence of states from start to a goal state (where h_func(s) == 0)."
frontier = [(h_func(start), start)] # A priority queue, ordered by path length, f = g + h
previous = {start: None} # start state has no previous state; other states will
path_cost = {start: 0} # The cost of the best path to a state.
Path = lambda s: ([] if (s is None) else Path(previous[s]) + [s])
while frontier:
(f, s) = heappop(frontier)
if h_func(s) == 0:
return Path(s)
for s2 in moves_func(s):
g = path_cost[s] + cost_func(s, s2)
if s2 not in path_cost or g < path_cost[s2]:
heappush(frontier, (g + h_func(s2), s2))
path_cost[s2] = g
previous[s2] = s
def bfs(start, moves_func, goals):
"Breadth-first search"
goal_func = (goals if callable(goals) else lambda s: s in goals)
return Astar(start, moves_func, lambda s: (0 if goal_func(s) else 1))
# +
def tests():
"Tests for my utility functions."
# Functions for Input, Parsing
assert Array('''1 2 3
4 5 6''') == ((1, 2, 3),
(4, 5, 6))
assert Vector('testing 1 2 3.') == ('testing', 1, 2, 3.0)
assert Integers('test1 (2, -3), #4') == (2, -3, 4)
assert Atom('123.4') == 123.4 and Atom('x') == 'x'
# Functions on Iterables
assert first('abc') == first(['a', 'b', 'c']) == 'a'
assert first_true([0, None, False, {}, 42, 43]) == 42
assert nth('abc', 1) == nth(iter('abc'), 1) == 'b'
assert cat(upto('abcdef', 'd')) == 'abcd'
assert cat(['do', 'g']) == 'dog'
assert groupby([-3, -2, -1, 1, 2], abs) == {1: [-1, 1], 2: [-2, 2], 3: [-3]}
assert list(grouper(range(8), 3)) == [(0, 1, 2), (3, 4, 5), (6, 7, None)]
assert list(overlapping((0, 1, 2, 3, 4), 3)) == [(0, 1, 2), (1, 2, 3), (2, 3, 4)]
assert list(overlapping('abcdefg', 4)) == ['abcd', 'bcde', 'cdef', 'defg']
assert list(pairwise((0, 1, 2, 3, 4))) == [(0, 1), (1, 2), (2, 3), (3, 4)]
assert sequence('seq') == 'seq'
assert sequence((i**2 for i in range(5))) == (0, 1, 4, 9, 16)
assert join(range(5)) == '01234'
assert join(range(5), ', ') == '0, 1, 2, 3, 4'
assert transpose(((1, 2, 3), (4, 5, 6))) == ((1, 4), (2, 5), (3, 6))
assert isqrt(9) == 3 == isqrt(10)
assert ints(1, 100) == range(1, 101)
assert identity('anything') == 'anything'
assert set(powerset({1, 2, 3})) == {
(), (1,), (1, 2), (1, 2, 3), (1, 3), (2,), (2, 3), (3,)}
assert quantify(['testing', 1, 2, 3, int, len], callable) == 2 # int and len are callable
assert quantify([0, False, None, '', [], (), {}, 42]) == 1 # Only 42 is truish
assert set(shuffled('abc')) == set('abc')
# Functional programming
assert mapt(math.sqrt, [1, 9, 4]) == (1, 3, 2)
assert map2d(abs, ((1, -2, -3), (-4, -5, 6))) == ((1, 2, 3), (4, 5, 6))
assert repeat(3, isqrt, 256) == 2
assert compose(isqrt, abs)(-9) == 3
# Making immutable objects
assert Set([1, 2, 3, 3]) == {1, 2, 3}
assert canon('abecedarian') == 'aaabcdeeinr'
assert canon([9, 1, 4]) == canon({1, 4, 9}) == (1, 4, 9)
# Math
assert transpose([(1, 2, 3), (4, 5, 6)]) == ((1, 4), (2, 5), (3, 6))
assert isqrt(10) == isqrt(9) == 3
assert ints(1, 5) == range(1, 6)
assert list(floats(1, 5)) == [1., 2., 3., 4., 5.]
assert multiply(ints(1, 10)) == math.factorial(10) == 3628800
# 2-D points
P = (3, 4)
assert X(P) == 3 and Y(P) == 4
assert cityblock_distance(P) == cityblock_distance(P, origin) == 7
assert distance(P) == distance(P, origin) == 5
assert turn_right(UP) == turn_left(DOWN) == turn_around(LEFT) == RIGHT
# Search
assert Astar((4, 4), neighbors8, distance) == [(4, 4), (3, 3), (2, 2), (1, 1), (0, 0)]
assert bfs((4, 4), neighbors8, {origin}) == [(4, 4), (3, 3), (2, 2), (1, 1), (0, 0)]
forty2 = always(42)
assert forty2() == forty2('?') == forty2(4, 2) == 42
return 'pass'
tests()
# -
# # [Day 1](https://adventofcode.com/2017/day/1): Inverse Captcha
#
# This was easier than I remember last year's puzzles being:
#
# +
digits = mapt(int, Inputstr(1))
N = len(digits)
N, digits[:10]
# -
sum(digits[i]
for i in range(N)
if digits[i] == digits[i - 1])
# ## Part Two
sum(digits[i]
for i in range(N)
if digits[i] == digits[i - N // 2])
# # [Day 2](https://adventofcode.com/2017/day/2): Corruption Checksum
#
rows2 = Array('''790 99 345 1080 32 143 1085 984 553 98 123 97 197 886 125 947
302 463 59 58 55 87 508 54 472 63 469 419 424 331 337 72
899 962 77 1127 62 530 78 880 129 1014 93 148 239 288 357 424
2417 2755 254 3886 5336 3655 5798 3273 5016 178 270 6511 223 5391 1342 2377
68 3002 3307 166 275 1989 1611 364 157 144 3771 1267 3188 3149 156 3454
1088 1261 21 1063 1173 278 1164 207 237 1230 1185 431 232 660 195 1246
49 1100 136 1491 647 1486 112 1278 53 1564 1147 1068 809 1638 138 117
158 3216 1972 2646 3181 785 2937 365 611 1977 1199 2972 201 2432 186 160
244 86 61 38 58 71 243 52 245 264 209 265 308 80 126 129
1317 792 74 111 1721 252 1082 1881 1349 94 891 1458 331 1691 89 1724
3798 202 3140 3468 1486 2073 3872 3190 3481 3760 2876 182 2772 226 3753 188
2272 6876 6759 218 272 4095 4712 6244 4889 2037 234 223 6858 3499 2358 439
792 230 886 824 762 895 99 799 94 110 747 635 91 406 89 157
2074 237 1668 1961 170 2292 2079 1371 1909 221 2039 1022 193 2195 1395 2123
8447 203 1806 6777 278 2850 1232 6369 398 235 212 992 7520 7304 7852 520
3928 107 3406 123 2111 2749 223 125 134 146 3875 1357 508 1534 4002 4417''')
sum(abs(max(row) - min(row)) for row in rows2)
# ## Part Two
# +
def evendiv(row):
return first(a // b for a in row for b in row if a > b and a // b == a / b)
sum(map(evendiv, rows2))
# -
# This day was also very easy. It was nice that my pre-defined `array` function did the whole job of parsing the input. In Part One, I was slowed down by a typo: I had `"="` instead of `"-"` in `"max(row) - min(row)"`. I was confused by Python's misleading error message, which said `"SyntaxError: keyword can't be an expression"`. Later on, <NAME> explained to me that the message meant that in `abs(max(row)=...)` it thought that `max(row)` was a keyword argument to `abs`, as in `abs(x=-1)`.
#
# In Part Two, note that to check that `a/b` is an exact integer, I used `a // b == a / b`, which I think is more clear than the marginally-faster expression one would typically use here, `a % b == 0`, which requires you to think about two things: division and the modulus operator (is it `a % b` or `b % a`?).
# # [Day 3](https://adventofcode.com/2017/day/3): Spiral Memory
#
# For today the data is just one number:
M = 277678
# This puzzle takes some thinking, not just fast typing. I decided to break the problem into three parts:
# - Generate a spiral (by writing a new function called `spiral`).
# - Find the Nth square on the spiral (with my function `nth`).
# - Find the distance from that square to the center (with my function `cityblock_distance`).
#
# I suspect many people will do all three of these in one function. That's probably the best way to get the answer really quickly, but I'd rather be clear than quick (and I'm anticipating that `spiral` will come in handy in Part Two), so I'll factor out each part, obeying the *single responsibility principle*.
#
# Now I need to make `spiral()` generate the coordinates of squares on an infinite spiral, in order, going out from the center square, `(0, 0)`. After the center square, the spiral goes 1 square right, then 1 square up, then 2 square left, then 2 square down, thus completing one revolution; then it does subsequent revolutions. In general if the previous revolution ended with *s* squares down, then the next revolution consists of *s*+1 squares right, *s*+1 squares up, *s*+2 squares left and *s*+2 down. A small test confirms that this matches the example diagram in the puzzle description (although I had a bug on my first try because I only incremented `s` once per revolution, not twice):
# +
def spiral():
"Yield successive (x, y) coordinates of squares on a spiral."
x = y = s = 0 # (x, y) is the position; s is the side length.
yield (x, y)
while True:
for (dx, dy) in (RIGHT, UP, LEFT, DOWN):
if dy: s += 1 # Increment side length before RIGHT and LEFT
for _ in range(s):
x += dx; y += dy
yield (x, y)
list(islice(spiral(), 10))
# -
# Now we can find the `N`th square. As this is Python, indexes start at 0, whereas the puzzle description starts counting at 1, so I have to subtract 1. Then I can find the distance to the origin:
nth(spiral(), M - 1)
cityblock_distance(_)
# ## Part Two
#
# I can re-use my `spiral` generator, yay! Here's a function to sum the neighboring squares (I can use my `neighbors8` function, yay!):
def spiralsums():
"Yield the values of a spiral where each square has the sum of the 8 neighbors."
value = defaultdict(int)
for p in spiral():
value[p] = sum(value[q] for q in neighbors8(p)) or 1
yield value[p]
list(islice(spiralsums(), 12))
# Looks good, so let's get the answer:
first(x for x in spiralsums() if x > M)
# # [Day 4](https://adventofcode.com/2017/day/4): High-Entropy Passphrases
#
# This is the first time I will have to store an input file and read it with the function `Input`. It should be straightforward, though:
# +
def is_valid(line): return is_unique(line.split())
def is_unique(items): return len(items) == len(set(items))
quantify(Input(4), is_valid)
# -
# ## Part Two
# +
def is_valid2(line): return is_unique(mapt(canon, line.split()))
quantify(Input(4), is_valid2)
# -
# That was easy, and I started on time, but the leaders were still three times faster than me!
# # [Day 5](https://adventofcode.com/2017/day/5): A Maze of Twisty Trampolines, All Alike
#
# Let's first make sure we can read the data/program okay:
# +
program = mapt(int, Input(5))
program[:10]
# -
# Now I'll make a little interpreter, `run`, which takes a program, loads it into memory,
# and executes the instruction, maintaining a program counter, `pc`, and doing the incrementing/branching as described in the puzzle,
# until the program counter no longer points to a location in memory:
# +
def run(program):
memory = list(program)
pc = steps = 0
M = len(memory)
while 0 <= pc < M:
steps += 1
oldpc = pc
pc += memory[pc]
memory[oldpc] += 1
return steps
run(program)
# -
# ## Part Two
#
# Part Two seems tricky, so I'll include an optional argument, `verbose`, and check if the printout it produces matches the example in the puzzle description:
# +
@jit
def run2(program, verbose=False):
memory = list(program)
pc = steps = 0
M = len(memory)
while 0 <= pc < M:
steps += 1
oldpc = pc
pc += memory[pc]
memory[oldpc] += (-1 if memory[oldpc] >= 3 else 1)
if verbose: print(steps, pc, memory)
return steps
run2([0, 3, 0, 1, -3], True)
# -
# That looks right, so I can solve the puzzle:
run2(program)
# Thanks to [<NAME>](https://github.com/ClementSreeves) for the suggestion of making a distinction between the `program` and the `memory`. In my first version, `run` would mutate the argument, which was OK for a short exercise, but not best practice for a reliable API. And thanks to [<NAME>](https://github.com/maxalbert) for speeding up the loop by pulling the `len(memory)` out of the loop.
# # [Day 6](https://adventofcode.com/2017/day/6): Memory Reallocation
# I had to read the puzzle description carefully, but then it is pretty clear what to do. I'll keep a set of previously seen configurations, which will all be tuples. But in the function `spread`, I want to mutate the configuration of banks, so I will convert to a list at the start, then convert back to a tuple at the end.
# +
banks = Vector('4 10 4 1 8 4 9 14 5 1 14 15 0 15 3 5')
def realloc(banks):
"How many cycles until we reach a configuration we've seen before?"
seen = {banks}
for cycles in count_from(1):
banks = spread(banks)
if banks in seen:
return cycles
seen.add(banks)
def spread(banks):
"Find the area with the most blocks, and spread them evenly to following areas."
banks = list(banks)
maxi = max(range(len(banks)), key=lambda i: banks[i])
blocks = banks[maxi]
banks[maxi] = 0
for i in range(maxi + 1, maxi + 1 + blocks):
banks[i % len(banks)] += 1
return tuple(banks)
# -
spread((0, 2, 7, 0))
realloc((0, 2, 7, 0))
# These tests look good; let's solve the problem:
realloc(banks)
# ## Part Two
#
# Here I will just replace the `set` of `seen` banks with a `dict` of `{bank: cycle_number}`; everything else is the same, and the final result is the current cycle number minus the cycle number of the previously-seen tuple of banks.
# +
def realloc2(banks):
"When we hit a cycle, what is the length of the cycle?"
seen = {banks: 0}
for cycles in count_from(1):
banks = spread(banks)
if banks in seen:
return cycles - seen[banks]
seen[banks] = cycles
realloc2((0, 2, 7, 0))
# -
realloc2(banks)
# # [Day 7](https://adventofcode.com/2017/day/7): Recursive Circus
# First I'll read the data into two dicts as follows: the input line
#
# tcmdaji (40) -> wjbdxln, amtqhf
#
# creates the two entries:
#
# weight['tcmdaji'] = 40
# above['tcmdaji'] = ['wjbdxln', 'amtqhf']
# +
def towers(lines):
"Return (weight, above) dicts."
weight = {}
above = {}
for line in lines:
name, w, *rest = re.findall(r'\w+', line)
weight[name] = int(w)
above[name] = set(rest)
return weight, above
weight, above = towers(Input(7))
programs = set(above)
# -
# Now the root progam is the one that is not above anything:
programs - set(flatten(above.values()))
# ## Part Two
#
# A program is *wrong* if it is the bottom of a tower that is a different weight from all its sibling towers:
def wrong(p): return tower_weight(p) not in map(tower_weight, siblings(p))
# Here we define `tower_weight`, `siblings`, and the `below` dict:
# +
def tower_weight(p):
"Total weight for the tower whose root (bottom) is p."
return weight[p] + sum(map(tower_weight, above[p]))
def siblings(p):
"The other programs at the same level as this one."
if p not in below:
return Ø # the root has no siblings
else:
return above[below[p]] - {p}
below = {a: b for b in programs for a in above[b]}
# -
set(filter(wrong, programs))
# So these four programs are wrong. Which one should we correct? The one that is wrong, and has no wrong program above it:
# +
def wrongest(programs):
return first(p for p in programs
if wrong(p)
and not any(wrong(p2) for p2 in above[p]))
wrongest(programs)
# -
# Now what should we correct it to? To the weight that makes it the same weight as the sibling towers:
# +
def correct(p):
"Return the weight that would make p's tower's weight the same as its sibling towers."
delta = tower_weight(first(siblings(p))) - tower_weight(p)
return weight[p] + delta
correct(wrongest(programs))
# -
# # [Day 8](https://adventofcode.com/2017/day/8): Memory Reallocation
#
# This one looks easy: a simple interpreter for straight-line code where each instruction has 7 tokens. It is nice that my `Array` function parses the whole program.
# +
program8 = Array(Input(8))
def run8(program):
"Run the program and return final value of registers."
registers = defaultdict(int)
for (r, inc, delta, _if, r2, cmp, amount) in program:
if operations[cmp](registers[r2], amount):
registers[r] += delta * (+1 if inc == 'inc' else -1)
return registers
max(run8(program8).values())
# -
# ## Part Two
#
# Here I modify the interpreter to keep track of the highest value of any register at any time.
# +
def run8_2(program):
registers = defaultdict(int)
highest = 0
for (r, inc, delta, _if, r2, cmp, amount) in program:
if operations[cmp](registers[r2], amount):
registers[r] += delta * (+1 if inc == 'inc' else -1)
highest = max(highest, registers[r])
return highest
run8_2(program8)
# -
# # [Day 9](https://adventofcode.com/2017/day/9): Stream Processing
#
# For this problem I could have defined a single parser that handles all five magic characters, `'{<!>}'`, but I think it is easier to first clean up the garbage, using regular expressions:
# +
text1 = re.sub(r'!.', '', Inputstr(9)) # Delete canceled characters
text2 = re.sub(r'<.*?>', '', text1) # Delete garbage
text2[:70]
# -
# Now I can deal with the nested braces (which can't be handled with regular expressions). The puzzle says "*Each group is assigned a score which is one more than the score of the group that immediately contains it,*" which is the same as saying that a group's score is its nesting level, a quantity that increases with each open-brace character, and decreases with each close-brace:
# +
def total_score(text):
"Total of group scores; each group scores one more than the group it is nested in."
total = 0
level = 0 # Level of nesting
for c in text:
if c == '{':
level += 1
total += level
elif c == '}':
level -= 1
return total
total_score(text2)
# -
# ## Part Two
# At first I thought that the amount of garbage is just the difference in lengths of `text2` and `text3`:
len(text1) - len(text2)
# But this turned out to be wrong; it counts the angle brackets themselves s being deleted, whereas the puzzle is actually asking how many character between the angle brackets are deleted. So that would be:
# +
text3 = re.sub(r'<.*?>', '<>', text1) # Delete garbage inside brackets, but not brackets
len(text1) - len(text3)
# -
# # [Day 10](https://adventofcode.com/2017/day/10): Stream Processing
# I have to do a bunch of reversals of substrings of `stream`. It looks complicated so I will include a `verbose` argument to `knothash` and confirm it works on the example puzzle. I break out the reversal into a separate function, `rev`. The way I handle reversal interacting with wraparound is that I first move all the items before the reversal position to the end of the list, then I do the reversal, then I move them back.
# +
stream = (63,144,180,149,1,255,167,84,125,65,188,0,2,254,229,24)
def knothash(lengths, N=256, verbose=False):
"Do a reversal for each of the numbers in `lengths`."
nums = list(range(N))
pos = skip = 0
for L in lengths:
nums = rev(nums, pos, L)
if verbose: print(nums)
pos = (pos + L + skip) % N
skip += 1
return nums[0] * nums[1]
def rev(nums, pos, L):
"Reverse nums[pos:pos+L], handling wrap-around."
# Move first pos elements to end, reverse first L, move pos elements back
nums = nums[pos:] + nums[:pos]
nums[:L] = reversed(nums[:L])
nums = nums[-pos:] + nums[:-pos]
return nums
# -
# Reverse [0, 1, 2]:
assert rev(list(range(5)), 0, 3) == [2, 1, 0, 3, 4]
# Reverse [4, 0, 1], wrapping around:
assert rev(list(range(5)), 4, 3) == [0, 4, 2, 3, 1]
# Duplicate the example output
assert knothash((3, 4, 1, 5), N=5, verbose=True) == 12
# That's correct, but the first time through I got it wrong because I forgot the `"% N"` on the update of `pos`.
knothash(stream)
# ## Part Two
#
# Now it gets *really* complicated: string processing, the suffix, hex string output, and dense hashing. But just take them one at a time:
# +
stream2 = '63,144,180,149,1,255,167,84,125,65,188,0,2,254,229,24'
def knothash2(lengthstr, N=256, rounds=64, suffix=(17, 31, 73, 47, 23),
verbose=False):
"Do a reversal for each length; repeat `rounds` times."
nums = list(range(N))
lengths = mapt(ord, lengthstr) + suffix
pos = skip = 0
for round in range(rounds):
for L in lengths:
nums = rev(nums, pos, L)
if verbose: print(nums)
pos = (pos + L + skip) % N
skip += 1
return hexstr(dense_hash(nums))
def hexstr(nums):
"Convert a sequence of (0 to 255) ints into a hex str."
return cat(map('{:02x}'.format, nums))
def dense_hash(nums, blocksize=16):
"XOR each block of nums, return the list of them."
return [XOR(block) for block in grouper(nums, blocksize)]
def XOR(nums):
"Exclusive-or all the numbers together."
result = 0
for n in nums:
result ^= n
return result
assert XOR([65, 27, 9, 1, 4, 3, 40, 50, 91, 7, 6, 0, 2, 5, 68, 22]) == 64
assert hexstr([255, 0, 17]) == 'ff0011'
assert knothash2('') == 'a2582a3a0e66e6e86e3812dcb672a272'
knothash2(stream2)
# -
# I had a bug: originally I used `'{:x}'` as the format instead of `'{:02x}'`; the later correctly formats `0` as `'00'`, not `'0'`.
# # [Day 11](https://adventofcode.com/2017/day/11): Hex Ed
#
# The first thing I did was search [`[hex coordinates]`](https://www.google.com/search?source=hp&ei=Ft4xWoOqKcy4jAOs76a4CQ&q=hex+coordinates), and the #1 result (as I expected) was <NAME>'s "[Hexagonal Grids](https://www.redblobgames.com/grids/hexagons/)" page. I chose his "odd-q vertical layout" to define the six headings as (dx, dy) deltas:
headings6 = dict(n=(0, -1), ne=(1, 0), se=(1, 1), s=(0, 1), sw=(-1, 0), nw=(-1, -1))
# Now I can read the path, follow it, and see where it ends up. From there, we have to compute how far we are from the origin: I can use my `king_distance` function for that—the number of moves a Chess King would take.
# +
path = Vector(Inputstr(11))
def follow(path):
"Follow each step of the path; return final distance to origin."
pos = origin
for dir in path:
pos = add(pos, headings6[dir])
return king_distance(pos)
follow(path)
# -
# This one seemed so easy that I didn't bother testing it on the simple examples in the puzzle; all I did was confirm that the answer for my puzzle input was correct.
#
# ## Part Two
#
# This looks pretty easy; repeat Part One, but keep track of the maximum number of steps we get from the origin at any point in the path:
# +
def follow2(path):
"Follow each step of the path; return the farthest away we ever got."
pos = origin
maxsteps = 0
for dir in path:
pos = add(pos, headings6[dir])
maxsteps = max(maxsteps, king_distance(pos))
return maxsteps
follow2(path)
# -
# Again, no tests, just the final answer.
#
# # [Day 12](https://adventofcode.com/2017/day/12): Digital Plumber
#
# First I'll parse the data, creating a dict of `{program: direct_group_of_programs}`:
# +
def groups(lines):
"Dict of {i: {directly_connected_to_i}"
return {lhs: {lhs} | set(rhs)
for (lhs, _, *rhs) in Array(lines)}
assert groups(Input(12))[0] == {0, 659, 737}
# -
# That looks good. I recognize this as a [Union-Find](https://en.wikipedia.org/wiki/Disjoint-set_data_structure) problem, for which there are efficient algorithms. But for this small example, I don't need efficiency, I need clarity and simplicity. So I'll write `merge` to take a dict and merge together the sets that are connected:
# +
def merge(G):
"Merge all indirectly connected groups together."
for i in G:
for j in list(G[i]):
if G[i] != G[j]:
G[i].update(G[j])
G[j] = G[i]
return G
G = merge(groups(Input(12)))
# -
len(G[0])
# That's the answer for Part One.
#
# ## Part Two
#
# I did almost all the work; I just need to count the number of distinct groups. That's a set of sets, and regular `set`s are not hashable, so I use my `Set` class:
len({Set(G[i]) for i in G})
# # [Day 13](https://adventofcode.com/2017/day/13): Packet Scanners
#
# First thing: The puzzle says the data is *depth: range*, but `range` has a meaning in Python, so I'll use the term *width* instead.
#
# Second thing: I misread the puzzle description and mistakenly thought the scanners were going in a circular route,
# so that they'd be at the top at any time that is 0 mod *width*. That gave the wrong answer and I realized the scanners are actually going back-and-forth, so with a width of size *n*, it takes *n* - 1 steps to get to the bottom, and *n* - 1 steps to get back to the top, so the scanner will be
# at the top at times that are multiples of 2(*n* - 1). For example, with width 3, that would be times 0, 4, 8, ...
# +
def trip_severity(scanners):
"The sum of severities for each time the packet is caught."
return sum((d * w if caught(d, w) else 0)
for (d, w) in scanners)
def caught(depth, width):
"Does the scanner at this depth/width catch the packet?"
return depth % (2 * (width - 1)) == 0
example = ((0, 3), (1, 2), (4, 4), (6, 4))
assert trip_severity(example) == 24
# -
scanners = mapt(Integers, Input(13))
scanners[:5]
trip_severity(scanners)
# ## Part Two
#
# A packet is safe if no scanner catches it. We now have the possibility of a delay, so I update `caught` to allow for an optional delay, and define `safe_delay`:
# +
def caught(depth, width, delay=0):
"Does the scanner at this depth/width catch the packet with this delay?"
return (depth + delay) % (2 * (width - 1)) == 0
def safe_delay(scanners):
"Find the first delay such that no scanner catches the packet."
safe = lambda delay: not any(caught(d, w, delay) for (d, w) in scanners)
return first(filter(safe, count_from(0)))
safe_delay(example)
# -
safe_delay(scanners)
# # [Day 14](https://adventofcode.com/2017/day/14): Disk Defragmentation
#
# I found this puzzle description confusing: are they talking about what I call `knothash`, or is it `knothash2`? I decided for the latter, which turned out to be right:
key = '<KEY>'
# +
def bits(key, i):
"The bits in the hash of this key with this row number."
hash = knothash2(key + '-' + str(i))
return format(int(hash, base=16), '0128b')
sum(bits(key, i).count('1') for i in range(128))
# -
# ## Part Two
#
# So as not to worry about running off the edge of the grid, I'll surround the grid with `'0'` bits:
# +
def Grid(key, N=128+2):
"Make a grid, with a border around it."
return border('0', (list(bits(key, i)) for i in range(128)))
def border(fill, grid):
"Surround a grid with a border of fill cells."
rows = [[fill] + list(row) + [fill]
for row in grid]
empty = [fill] * len(rows[0])
return [empty] + rows + [empty]
# -
# To find a region, start at some `(x, y)` position and [flood fill](https://en.wikipedia.org/wiki/Flood_fill) to neighbors that have the same value (a `'1'` bit).
def flood(grid, x, y, val, R):
"For all cells with value val connected to grid[x][y], give them region number R."
if grid[y][x] == val:
grid[y][x] = R
for x2, y2 in neighbors4((x, y)):
flood(grid, x2, y2, val, R)
def flood_all(grid, val='1'):
"Label all regions with consecutive ints starting at 1."
R = 0 # R is the region number
for y in range(1, len(grid) - 1):
for x in range(1, len(grid) - 1):
if grid[y][x] == val:
R += 1
flood(grid, x, y, val, R)
return R
flood_all(Grid(key))
# # [Day 15](https://adventofcode.com/2017/day/15): Dueling Generators
#
# My personalized inputs for this puzzle are `516` and `190`; the other numbers are shared by all puzzle-solvers. I decided to make infinite generators of numbers, using `gen`:
# +
@jit
def gen(prev, factor, m=2147483647):
"Generate a sequence of numbers according to the rules; stop at 0."
while prev:
prev = (prev * factor) % m
yield prev
def judge(A, B, N, mask=2**16-1):
"How many of the first N numbers from A and B agree in the masked bits (default last 16)?"
return quantify(a & mask == b & mask
for (a, b, _) in zip(A, B, range(N)))
def A(): return gen(516, 16807)
def B(): return gen(190, 48271)
# %time judge(A(), B(), 40*10**6)
# -
# Notice I also decided to use `@jit` (i.e. `numba.jit`) to speed things up, since this is the slowest-running day yet.
#
# ## Part Two
#
# A small change: only consider numbers that match the **criteria** of being divisible by 4 or 8, respectively;
# +
def criteria(m, iterable):
"Elements of iterable that are divisible by m"
return (n for n in iterable if n % m == 0)
# %time judge(criteria(4, A()), criteria(8, B()), 5*10**6)
# -
# When I got this solution on Day 15, I was happy to end there. But looking back, after Day 25, I noticed this day's run time was the slowest of all, so I wondered if I could speed things up, using `@jit`. Unfortunately, `@jit` doesn't work with generators, so I'll have to rewrite the code:
# +
@jit
def duelgen(prev1=516, factor1=16807, prev2=190, factor2=48271,
m=2147483647, mask=2**16-1, N=40*10**6):
matches = 0
for _ in range(N):
prev1 = (prev1 * factor1) % m
prev2 = (prev2 * factor2) % m
matches += (prev1 & mask == prev2 & mask)
return matches
# %time duelgen()
# -
# That was an excellent speedup (and the same answer); I'll leave optimizing Part Two as an exercise for the reader.
# # [Day 16](https://adventofcode.com/2017/day/16): Permutation Promenade
#
# Let's read the input and check that it looks reasonable:
dance = Vector(Inputstr(16))
dance[:10]
len(dance)
# I'll define `perform` to perform the dance:
# +
dancers = 'abcdefghijklmnop'
def perform(dance, dancers=dancers):
D = deque(dancers)
def swap(i, j): D[i], D[j] = D[j], D[i]
for move in dance:
op, arg = move[0], move[1:]
if op == 's': D.rotate(int(arg))
elif op == 'x': swap(*Integers(arg))
elif op == 'p': swap(D.index(arg[0]), D.index(arg[2]))
return cat(D)
perform(dance)
# -
# That's the right answer.
#
# ## Part Two
#
# My first thought was to define a dance as a permutation: a list of numbers `[11, 1, 9, ...]` which says that the net effect of the dance is that the first dancer (`a`) ends up in position, the second (`b`) stays in position 1, and so on. Applying that permutation once is a lot faster than interpreting all 10,000 moves of the dance, and it is feasible to apply the permutation a billion times. I tried that (code not shown here), but that was a mistake: it took 15 minutes to run, and it got the wrong answer. The problem is that a dance is *not* just a permutation, because a dance can reference dancer *names*, not just positions.
#
# It would take about 10,000 times 20 minutes to perform a billion repetitions of the dance, so that's out. But even though the dance is not a permutation, it might repeat after a short period. Let's check:
seen = {dancers: 0}
d = dancers
for i in range(1, 1000):
d = perform(dance, d)
if d in seen:
print(d, 'is seen in iterations', (seen[d], i))
break
seen[d] = i
# So we get back to the start position after 56 repetitions of the dance. What happens after a billion repetitions?
1000000000 % 56
# The end position after a billion repetitions is the same as after 48:
# +
def whole(N, dance, dancers=dancers):
"Repeat `perform(dance)` N times."
for i in range(N):
dancers = perform(dance, dancers)
return dancers
whole(48, dance)
# -
#
# # [Day 17](https://adventofcode.com/2017/day/17): Spinlock
#
# This one looks pretty easy:
# +
step = 314
def spinlock(step=step, N=2017):
"Make N inserts into the buffer, skipping ahead by `step` each time."
buf = [0]
pos = 0
for i in ints(1, N):
pos = (pos + step) % i + 1
buf[pos:pos] = [i]
return buf
buf = spinlock()
buf[buf.index(2017)+1]
# -
# That's the right answer.
#
# ## Part Two
#
# But Part Two is not so easy, if we care about the run time. Insertion into a `list` has to move all the elements after the insertion down, so insertion is O(N) and `spinlock` is O(N<sup>2</sup>). That's no problem when N = 2017, but when N is 50 million? We're gonna need a bigger boat, where by "boat" I mean algorithm or data structure. My first thought is a (circular) linked list, because insertion is O(1). I can implement the three key methods: `skip` to move ahead, `insert` to add a new node after the current one, and `find` to find a piece of data (with a linear search):
class Node:
"A Node in a singly-linked list"
__slots__ = ('data', 'next') # Declaring slots makes it more efficient
def __init__(self, data, next): self.data, self.next = data, next
def skip(self, n):
"Skip ahead n nodes, and return that node."
node = self
for i in range(n):
node = node.next
return node
def insert(self, value):
"Insert a new node with the given value after this node."
self.next = Node(value, self.next)
return self.next
def find(self, value):
"Find the node with the given data value."
node = self
while node.data != value:
node = node.next
return node
# Now I can rewrite `spinlock` to use this class:
def spinlock2(step=step, N=2017):
node = Node(0, None)
node.next = node # Make node be a circular linked list
for i in ints(1, N):
node = node.skip(step).insert(i)
return node
# Let's replicate the Part One results:
spinlock2().find(2017).next.data
# Good news! We get the same answer. But how fast/slow is it?
# %time spinlock2(N=100000)
# Bad news! More than a second for just 100,000 insertions, which projects to over 10 minutes for 50 million insertions. I did in fact try
#
# spinlock2(N=50000000).find(0).next.data
#
# and it eventually gave the right answer, but while it was running I had plenty of time to think.
# I realized that, if we go back to the original `spinlock` version, the value `0` will always be in `buf[0]`, and the value we are looking for will always be in `buf[1]`. So I can create a version of `spinlock` that only keeps track of `buf[0:2]`. That should run in a few seconds, not minutes:
# +
def spinlock3(step=step, N=2017):
"Make N inserts into a simulated buffer, but ignore all except buf[0:2]."
pos = 0
buf = [0, 0]
for i in ints(1, N):
pos = (pos + step) % i + 1
if pos <= 1:
buf[pos] = i
return buf
# %time spinlock3(N=50000000)[1]
# -
# The moral of the story is *keep your eyes on the prize*. I got distracted because I asked the wrong question. I asked myself "how can I make my solution in `spinlock` faster?" and answered myself "insertion is O(N<sup>2</sup>) and it should be O(N)." I knew how to do that, with a linked list, but that was the right answer to the wrong question. I should have asked myself "how do I solve Part Two quickly," concentrating on solving the actual problem. Once I did that, I realized I didn't need all those insertions: not doing them at all is a better idea than doing them faster.
# # [Day 18](https://adventofcode.com/2017/day/17): Duet
#
# First, read the input, and take a peak at it:
program18 = Array(Input(18))
program18[:10]
# Now write an interpreter for the assembly language:
# +
def run18(program):
"Interpret the assembly language program; return recovered `snd`."
regs = defaultdict(int)
pc = snd = 0
while True:
instr = program[pc]
pc += 1
op, x, y = instr[0], instr[1], instr[-1]
vy = value(regs, y)
if op == 'snd': snd = regs[x]
elif op == 'set': regs[x] = vy
elif op == 'add': regs[x] += vy
elif op == 'mul': regs[x] *= vy
elif op == 'mod': regs[x] %= vy
elif op == 'jgz' and regs[x] > 0: pc += vy - 1
elif op == 'rcv' and regs[x] != 0: return snd
def value(regs, y): return (y if isinstance(y, int) else regs[y])
run18(program18)
# -
# That was easy. (One tricky bit: the `pc` is incremented by 1 every time through the loop, regardless of the instruction. Therefore, the `'jgz'` jump instruction increments by "`vy - 1`" so that the net increment is "`vy`".)
#
# ## Part Two
#
# In Part Two we have to run two copies of the program, and send messages between them. I'll break up the loop in `run18` into
# two functions. First, `run18_2`, creates (in `ps`) two structures to hold the state variables necessary to run a program:
# - `id`: The id number (0 or 1) of this copy of the program.
# - `pc`: The program counter.
# - `sends`: A count of the number of `snd` instructions executed.
# - `regs`: A dict of the program registers (`a` to `z`).
# - `status`: A program has a status which can be:
# * `'run'` when it is ready to execute an instruction,
# * `'wait'` when it is waiting for a value to arrive in its input queue, or
# * `'end'` when the `pc` has run off the end of the program and it has terminated.
#
# `run18_2` repeatedly calls the second function, `step18(program, p)` to execute one instruction of `program` with the state variables in `p`. I choose randomly which of the two programs to step on each iteration. The function exits when neither copy of the program can run, according to their status.
def run18_2(program):
"Run two copies of program, with different state variables. Return final states."
Qs = [deque(), deque()]
ps = [Struct(id=id, pc=0, sends=0, regs=defaultdict(int, p=id), status='run')
for id in (0, 1)]
while any(p.status == 'run' for p in ps):
step18(program, Qs, random.choice(ps))
return ps
# `step18` has most of the guts of thee previous `run18` function, but with a few changes:
# - State variables are accessed indirectly: `p.pc` instead of just `pc`.
# - If the `pc` is out of bounds, the program terminates; the status is set to `'end'`.
# - The `snd` instruction sends a value to the other program's queue.
# - The `rcv` instruction pops a value off the queue if there is one, otherwise the status is set to `'wait'`.
# - The "`X`" in "`jgz X Y`" might be an integer, not a register name, so use `vx = value(p.regs, x)`. I was stuck for a *long* time before I realized this. Finally I tried the strategy of *look carefully at the input*. I noticed the instruction `"jgz 1 3"`, and it was a simple change to make the program work.
# +
def step18(program, Qs, p):
"Execute one instruction in program, using state variables in p."
if p.pc < 0 or p.pc > len(program):
p.status = 'end'
else:
instr = program[p.pc]
op, x, y = instr[0], instr[1], instr[-1]
vx, vy = value(p.regs, x), value(p.regs, y)
if op == 'snd': Qs[1-p.id].append(vy); p.sends += 1
elif op == 'set': p.regs[x] = vy
elif op == 'add': p.regs[x] += vy
elif op == 'mul': p.regs[x] *= vy
elif op == 'mod': p.regs[x] %= vy
elif op == 'jgz' and vx > 0: p.pc += vy - 1
elif op == 'rcv':
if not Qs[p.id]:
p.status = 'wait'
return # don't update pc; try again next time
else:
p.regs[x] = Qs[p.id].popleft()
p.status = 'run'
p.pc += 1
run18_2(program18)[1].sends
# -
# # [Day 19](https://adventofcode.com/2017/day/19): A Series of Tubes
# At first I was confused; I thought this was a maze-following problem where I had to make a choice of directions at every turn. Actually, the direction is always determined: keep going in the current direction as long as possible, but when we hit a `'+'` character, find the new direction to go in (there will only be one possibility). Leave breadcrumbs (the `'.'` character) so that we don't back up along a previously-followed path. As in Day 14, the grid is surrounded by a border of space characters so that we don't have to worry about `(x, y)` going off the edge.
# +
diagram = Inputstr(19)
def follow_tubes(diagram):
"Follow [-+|] lines, yielding characters along the path."
grid = border(' ', diagram.splitlines())
x, y = grid[1].index('|'), 1
dx, dy = 0, 1
while grid[y][x] != ' ':
yield grid[y][x]
if grid[y][x] == '+':
dx, dy = new_direction(grid, x, y)
grid[y][x] = '.' # Leave a breadcrumb
x += dx; y += dy
def new_direction(grid, x, y):
"Find a direction that continues the path."
for (dx, dy) in (UP, DOWN, RIGHT, LEFT):
if grid[y+dy][x+dx] not in (' ', '.'):
return dx, dy
# -
cat(filter(str.isalpha, follow_tubes(diagram)))
# That's the right answer.
# ## Part Two
#
# This is a surprisingly easy Part Two; I already generated the characters in the path; all I have to do is count them:
length(follow_tubes(diagram))
# # [Day 20](https://adventofcode.com/2017/day/20): Particle Swarm
#
# I'll create structures for particles, each will have fields for particle's number (`id`), position (`p`), velocity(`v`), and acceleration (`a`). I have `particles` as a function that creartes a collection, and not a collection in its own right, because I anticipate that I will want to mutate particles, so I'll need a fresh copy every time I want to do something with them.
# +
def particles(lines=tuple(Input(20))):
"Parse the input file into a list of particles."
return [Particle(id, *grouper(Integers(line), 3))
for id, line in enumerate(lines)]
def Particle(id, p, v, a): return Struct(id=id, p=p, v=v, a=a)
particles()[:5]
# -
# I'm not quite sure how to determine what "in the long run" means, so I'll just interpret it as meaning "after 1000 updates."
# +
def update(particles):
"Update velocity and position of all particles."
for r in particles:
r.v = add(r.v, r.a)
r.p = add(r.p, r.v)
return particles
def closest(particles):
"Find the particle closest to origin."
return min(particles, key=lambda r: sum(map(abs, r.p)))
# Answer: the id of the particle closest to origin after 1000 updates
closest(repeat(1000, update, particles())).id
# -
# ## Part Two
#
# I'll add the function `remove_collisions`, and now the thing we repeatedly do is the composition of `remove_collisions` and `update`. Also, instead of finding the `id` of the `closest` particle, now we just need to count the number of surviving particles:
# +
def remove_collisions(particles):
"Eliminate particles that are in the same place as another."
num_particles_at = Counter(r.p for r in particles)
return [r for r in particles if num_particles_at[r.p] == 1]
# Answer: number of particles remaining after collisions removed
len(repeat(1000, compose(remove_collisions, update), particles()))
# -
# I got the right answer both times, so my assumption that "in the long run" means "1000 updates" turned out to work for my input data, but I feel bad that it is not guaranteed to work for all input data.
# # [Day 21](https://adventofcode.com/2017/day/21): Fractal Art
#
# Today looks like a complex one, so I'll break the code up into more chunks and have more test assertions than usual. I can identify the following important data types:
#
# - `Enhancements`: a `dict` of `{grid: larger_grid}` rewrite rules.
# - `grid`: a square of 0-or-1 pixels, such as `((0, 1), (0, 1))`. The function `Pixels` translates text into this form.
#
# I define the functions `rotate` and `flip`; the puzzle descriptions says "When searching for a rule to use, rotate and flip the pattern as necessary," but I'm going to be doing many searches, and only one initialization of the rule set, so it will be more efficient to do the rotating and flipping just once:
# +
def Enhancements(lines):
"Create a dict of {grid: enhanced_grid}; include all rotations/flips."
enhancements = {}
for line in lines:
lhs, rhs = map(Pixels, line.split('=>'))
for rot in range(4):
enhancements[lhs] = enhancements[flip(lhs)] = rhs
lhs = rotate(lhs)
return enhancements
def Pixels(text):
"Translate the str '.#/.#' to the grid ((0, 1), (0, 1))"
bits = {'#': 1, '.': 0}
return tuple(tuple(bits[p] for p in row.strip())
for row in text.split('/'))
def rotate(subgrid):
"Rotate a subgrid 90 degrees clockwise."
return tuple(zip(*reversed(subgrid)))
def flip(subgrid):
"Reverse every row of the subgrid."
return tuple(tuple(reversed(row)) for row in subgrid)
# -
# Let's test some assertions, and then look at all the 2x2 enhancement rulesfrom my input file; with rotations and flips there should be 2<sup>4</sup> = 16 entries:
# +
assert Pixels('../##') == ((0, 0), (1, 1))
assert rotate(((0, 0), (1, 1))) == ((1, 0), (1, 0))
assert flip(((0, 0, 1), (1, 1, 0))) == ((1, 0, 0), (0, 1, 1))
Enhancements('''
../.. => .../.#./.#.
#./.. => .../#../#..
##/.. => #.#/.#./.#.
.#/#. => ##./##./...
##/#. => .##/###/#..
##/## => .##/#../##.
'''.strip().splitlines())
# -
assert len(_) == 2 ** 4
# Looks good; let's create the complete `enhancements` for my data. There should be 2<sup>4</sup> + 2<sup>9</sup> = 528 entries:
# +
enhancements = Enhancements(Input(21))
assert len(enhancements) == 2 ** 4 + 2 ** 9
# -
# Now on each iteration we `enhance` the grid by first dividing it into pieces with `divide_grid`, then using my utility function `map2d` to apply `enhancements` to each piece, and then call `stitch_grid` to put all the pieces back together into a bigger grid:
# +
def enhance(grid):
"Divide the drid into pieces, enhance each piece, and stitch them together."
return stitch_grid(map2d(enhancements.get, divide_grid(grid)))
def divide_grid(grid):
"Divide the grid into d x d pieces and enhance each piece."
N = len(grid[0])
d = (2 if N % 2 == 0 else 3 if N % 3 == 0 else error())
return [[tuple(row[c:c+d] for row in grid[r:r+d])
for c in range(0, N, d)]
for r in range(0, N, d)]
def stitch_grid(pieces):
"Stitch the pieces back into one big grid."
N = sum(map(len, pieces[0]))
return tuple(tuple(getpixel(pieces, r, c)
for c in range(N))
for r in range(N))
def getpixel(pieces, r, c):
"The pixel at location (r, c), from a matrix of d x d pieces."
# Use `//` to find the right piece, and `%` to find the pixel within the piece
d = len(pieces[0][0])
piece = pieces[r // d][c // d]
return piece[r % d][c % d]
# +
# Some tests
corners = Pixels('#..#/..../..../#..#')
pieces = [[((1, 0),
(0, 0)), ((0, 1),
(0, 0))],
[((0, 0),
(1, 0)), ((0, 0),
(0, 1))]]
assert divide_grid(corners) == pieces
assert stitch_grid(pieces) == corners
# -
# An extended test
grid = Pixels('.#./..#/###')
grid
divide_grid(_)
map2d(enhancements.get, _)
stitch_grid(_)
divide_grid(_)
map2d(enhancements.get, _)
stitch_grid(_)
sum(flatten(_))
# That looks right; Let's try to solve the whole puzzle:
sum(flatten(repeat(5, enhance, grid)))
# That's correct!
#
# ## Part Two
#
# Huh — It looks like I don't need to change any code for Part Two, just do `18` repetitions instead of `5`.
#
# Well, almost. Doing that gave an answer (in a few seconds); but the answer was wrong. I carefully looked over all my code, and realized there was a place where I had swapped the order of `r` and `c`. Once I fixed that (the fix is already incorporated above), I got the right answer:
# %time sum(flatten(repeat(18, enhance, grid)))
# # [Day 22](https://adventofcode.com/2017/day/22): Sporifica Virus
#
# This one looks to be of medium difficulty. One important choice: since we are dealing with "a seemingly-infinite two-dimensional grid of compute nodes," and I think it will be sparse, I'll represent the grid with a `set` of the positions of infected nodes, rather than with a 2-dimensional array. I'll define a `namedtuple` to hold the state of the network: the current position of the virus, its heading, the number of infections caused so far, and the set of infected nodes:
# +
Net = namedtuple('Net', 'current, heading, caused, infected')
def parse_net(lines):
"Read the initial state of the network."
lines = list(lines)
center = (len(lines) // 2, len(lines[0].strip()) // 2)
return Net(center, UP, 0,
{(x, y)
for (y, row) in enumerate(lines)
for (x, node) in enumerate(row)
if node == '#'})
# +
test = '''
..#
#..
...
'''.strip().splitlines()
parse_net(test)
# -
# Now the logic for one step of the simulation, called a *burst*:
def burst(net):
"Simulate the virus through one step and return the new state of the network."
(current, heading, caused, infected) = net
heading = (turn_right if current in infected else turn_left)(heading)
if current in infected:
infected.remove(current)
else:
caused += 1
infected.add(current)
return Net(add(current, heading), heading, caused, infected)
# We're supposed to get 5 infections caused in the first 7 steps:
repeat(7, burst, parse_net(test))
# And 41 out of 70:
repeat(70, burst, parse_net(test))
# This testing revealed a problem: I had (yet again) messed up the order of (x, y). (I find it confusing that there are two traditional orders: (x, y) and (row, col), and this is yet another reminder that I have to pay extra attention to keep them straight.) After fixing that, I was
# ready to solve the problem:
repeat(10000, burst, parse_net(Input(22))).caused
# ## Part Two
#
# It looks like I can't re-use any of my code from Part One (except by copy-and-paste). I have the following concerns:
# - I want to replace the `set` of `infected` nodes with a `dict`, `status[node]`, which can be `I`, `F`, `C`, or `W` (default `C` for clean).
# - I need to run 10,000,000 steps, so I want it to be efficient.
# - I have some confidence from doing Part One successfully, so I'm comfortable stressing efficiency over simplicity.
# I'll use variables inside a function, `bursts`, that does `N` repetitions; I'll avoid creating a new `Net` object each iteration.
def bursts(N, net):
"Run N steps of bursts on the network depicted by `lines`."
(current, heading, caused, infected) = net
status = defaultdict(lambda: 'C', {pos: 'I' for pos in infected})
for _ in range(N):
S = status[current]
if S == 'C':
heading = turn_left(heading)
status[current] = 'W'
elif S == 'W':
# heading unchanged
status[current] = 'I'
caused += 1
elif S == 'I':
heading = turn_right(heading)
status[current] = 'F'
elif S == 'F':
heading = turn_around(heading)
status[current] = 'C'
current = add(current, heading)
return caused
# Of the first 100 bursts of the test network, 26 will result in infection
assert bursts(100, parse_net(test)) == 26
# I had another bug here that gave me the wrong answer the first time: I had put the "`caused += 1`" line under the condition where the status *was* `'I'`, whereas it actually belongs under the condition where the status *becomes* `'I'`. With that fix, I get the right answer:
# %time bursts(10000000, parse_net(Input(22)))
#
#
# # [Day 23](https://adventofcode.com/2017/day/23): Coprocessor Conflagration
#
# Part One looks straightforward. I won't make the "register X might be an integer" mistake again:
#
#
# +
def run23(program):
regs = {L: 0 for L in 'abcdefgh'}
pc = 0
mulcount = 0
while 0 <= pc < len(program):
op, X, Y = program[pc]
pc += 1
if op == 'set': regs[X] = value(regs, Y)
elif op == 'sub': regs[X] -= value(regs, Y)
elif op == 'mul': regs[X] *= value(regs, Y); mulcount += 1
elif op == 'jnz' and value(regs, X): pc += value(regs, Y) - 1
return mulcount
run23(Array(Input(23)))
# -
# ## Part Two
#
# The hint of "You'll need to **optimize the program**" reminded me of a puzzle from 2016 where I had to understand what the program was doing and make it more efficient. It wasn't obvious what Day 23's program was doing, but I began the process of re-writing it as a Python program, converting the `jnz` instructions to `if` and `while` statements. Eventually I realized that the inner loop was doing "`b % d`", and my program became the following:
# +
@jit
def run23_2():
a = 1
d = e = f = g = h = 0
b = 99
c = b
if a:
b *= 100
b -= -100000
c = b
c -= -17000
while True:
f = 1
d = 2
e = 2
while True:
if b % d == 0:
f = 0
d -= -1
g = d - b
if g == 0:
if f == 0:
h -= -1
g = b - c
if g == 0:
return h
b -= -17
break
# %time run23_2()
# -
# The `numba.jit` decorator really helps here, speeding up execution from 13 seconds to 1 second.
#
# # [Day 24](https://adventofcode.com/2017/day/24): Electromagnetic Moat
#
# First I will read the data and store it as a table of `{port_number: [components_with_that_port]}`. I also define two simple utility functions:
# +
def component_table(pairs):
"Make a table of {port: {components_with_that_port}"
ctable = defaultdict(set)
for pair in pairs:
ctable[pair[0]].add(pair)
ctable[pair[1]].add(pair)
return ctable
ctable = component_table(map(Integers, Input(24)))
def other_port(component, port):
"The other port in a two-port component."
return (component[1] if component[0] == port else component[0])
def strength(chain): return sum(flatten(chain))
# -
# We are dealing with an optimization problem involving paths in a graph (called *chains* in this problem), and we're looking for the chain that maximizes `strength`. I'll represent a chain as a tuple of components. I could have defined a single function that traverses the graph and also keeeps track of the maximum, but I think it is cleaner to keep the two aspects of the problem separate. First a function to generate all possible chains:
def chains(chain=(), port=0, ctable=ctable):
"Given a partial chain ending in `port`, yield all chains that extend it."
yield chain
for c in ctable[port]:
if c not in chain:
# Extend with components, c, that match port but are not already in chain
yield from chains(chain + (c,), other_port(c, port), ctable)
# And then asking for the strength of the strongest chain:
# %time strength(max(chains(), key=strength))
# I was worried it was going to be slow, so I measured the `%time`, but it turned out not too bad.
#
# ## Part Two
#
# Now we want to find the strength of the longest chain, but if there is a tie, pick the strongest one:
# +
def length_and_strength(c): return len(c), strength(c)
# %time strength(max(chains(), key=length_and_strength))
# -
# I think I made the right choice in defining things the way I did. My code is simple, and gets the right answers in a few seconds. But I realize there are some inefficiencies:
#
# - Calculating the strength of a chain is O(N), but since we always form new chains by extending an old chain (for which we know the strength) with one new component, calculating the strength of the new chain could be O(1).
# - A chain is a `tuple`, so checking "`c not in chain`" is O(N). If the chain were a `set`, it would be O(1).
# - A new chain is created by *copying* the previous chain and appending a new component. A more efficient approach is to *mutate* the chain by adding a component, and then removing the component when it is time to consider other possibilities. This is called *backtracking*.
#
# Here is a backtracking implementation. It keeps track of a single `chain`, `port`, and `strength`. A call to `recurse(best_strength)` returns the best strength, either the one passed in, or one found by adding components to the current chain. When `recurse` returns, `chain`, `port`, and `strength` are reset to their original values, and the best strength found is returned as the value of the call to `recurse`. This is indeed faster (and gives the same answer):
# +
def strongest_chain(ctable=ctable):
"""Return the strength of the strongest chain, using backtracking."""
chain = set()
port = 0
strength = 0
def recurse(best_strength):
nonlocal chain, port, strength
for c in ctable[port] - chain:
# Update chain, port, strength
# then recurse and possibly update best_strength
# then backtrack and restore chain, port, strength
chain.add(c)
port = other_port(c, port)
strength += sum(c)
best_strength = max(strength, recurse(best_strength))
chain.remove(c)
port = other_port(c, port)
strength -= sum(c)
return best_strength
return recurse(0)
# %time strongest_chain()
# -
# You can decide whether the saving in time is worth the complication in code.
# # [Day 25](https://adventofcode.com/2017/day/25): The Halting Problem
#
# I won't write a parser for my input; instead I'll translate it into a `dict` by hand:
def machine():
"machine()[state][value] == (new_value, move, new_state)}"
L, R = -1, +1
A, B, C, D, E, F = 'ABCDEF'
return {A: [(1, R, B), (0, L, C)],
B: [(1, L, A), (1, R, D)],
C: [(0, L, B), (0, L, E)],
D: [(1, R, A), (0, R, B)],
E: [(1, L, F), (1, L, C)],
F: [(1, R, D), (1, R, A)]}
# Now a simple interpreter for machines like this:
# +
def turing(machine, state, steps):
"Run the Turing machine for given number of steps, then return tape."
tape = defaultdict(int)
cursor = 0
for step in range(steps):
tape[cursor], move, state = machine[state][tape[cursor]]
cursor += move
return tape
sum(turing(machine(), 'A', 12667664).values())
# -
# There is no **Part Two** today; we're done!
#
# # A Note on Reuse
#
# One interesting question: for what days did my Part Two code reuse the Part One code? How so?
# Here are my answers:
#
#
# * **Total Reuse (11 days)**: The major function defined in Part One is called again in Part Two:
# <br>Days 3 (`spiral`), 6 (`spread`, but `realloc2` is copy-edit), 9, 12, 14 (`bits`),
# 15 (`A, B, gen, judge`), 16 (`perform`), 19 (`follow_tubes`), 20 (`update, particles`), 21 (`enhance`),
# 24 (`chains`, `strength`)
#
# * **Generalization (1 day)**: A major function from Part One is generalized in Part Two (e.g. by adding an optional parameter):
# <br>Days 13 (`caught`)
#
# * **Copy-edit (7 days)**: The major function from Part One is copied and edited for Part Two:
# <br>Days 5 (`run2`), 8 (`run8_2`), 10 (`knothash2`), 11 (`follow2`), 17 (`spinlock2`), 18 (`run18_2`), 22 (`parse_net2`, `burst2`)
#
# * **All new (5 days)**: All the code for Part Two (except possibly reading and parsing the input) is brand new:
# <br>Days 1, 2, 4, 7, 23
#
# I think I did a reasonably good job of facilitating reuse. It seems like using generators and higher-order functions like `repeat` helps.
#
#
# # Verification and Run Times
#
# A little test harness and a report on all the run times that are over 5 seconds per day:
# +
# %%time
def run_tests(tests, short=5.0):
"Run daily test assertions; report times > `short` seconds."
for day in sorted(tests):
t0 = time.time()
assert tests[day]()
dt = time.time() - t0
if dt > short:
print('Day {:2d}: {:4.1f} sec'.format(day, dt))
run_tests({
1: lambda: sum(digits[i] for i in range(N) if digits[i] == digits[i - 1]) == 1158 and
sum(digits[i] for i in range(N) if digits[i] == digits[i - N // 2]) == 1132,
2: lambda: sum(abs(max(row) - min(row)) for row in rows2) == 46402 and
sum(map(evendiv, rows2)) == 265,
3: lambda: cityblock_distance(nth(spiral(), M - 1)) == 475 and
first(x for x in spiralsums() if x > M) == 279138,
4: lambda: quantify(Input(4), is_valid) == 337 and
quantify(Input(4), is_valid2) == 231,
5: lambda: run(program) == 364539 and
run2(program) == 27477714,
6: lambda: realloc(banks) == 12841 and
realloc2(banks) == 8038,
7: lambda: first(programs - set(flatten(above.values()))) == 'wiapj' and
correct(wrongest(programs)) == 1072,
8: lambda: max(run8(program8).values()) == 6828 and
run8_2(program8) == 7234,
9: lambda: total_score(text2) == 9662 and
len(text1) - len(text3) == 4903,
10: lambda: knothash(stream) == 4480 and
knothash2(stream2) == 'c500ffe015c83b60fad2e4b7d59dabc4',
11: lambda: follow(path) == 705 and
follow2(path) == 1469,
12: lambda: len(G[0]) == 115 and
len({Set(G[i]) for i in G}) == 221,
13: lambda: trip_severity(scanners) == 1504 and
safe_delay(scanners) == 3823370,
14: lambda: sum(bits(key, i).count('1') for i in range(128)) == 8316 and
flood_all(Grid(key)) == 1074,
15: lambda: duelgen() == 597 and
judge(criteria(4, A()), criteria(8, B()), 5*10**6) == 303,
16: lambda: perform(dance) == 'lbdiomkhgcjanefp' and
whole(48, dance) == 'ejkflpgnamhdcboi',
17: lambda: spinlock2().find(2017).next.data == 355 and
spinlock3(N=50*10**6)[1] == 6154117,
18: lambda: run18(program18) == 7071 and
run18_2(program18)[1].sends == 8001,
19: lambda: cat(filter(str.isalpha, follow_tubes(diagram))) == 'VEBTPXCHLI' and
quantify(follow_tubes(diagram)) == 18702,
20: lambda: closest(repeat(1000, update, particles())).id == 243 and
len(repeat(1000, compose(remove_collisions, update), particles())) == 648,
21: lambda: sum(flatten(repeat(5, enhance, grid))) == 147 and
sum(flatten(repeat(18, enhance, grid))) == 1936582,
22: lambda: repeat(10000, burst, parse_net(Input(22))).caused == 5460 and
bursts(10000000, parse_net(Input(22))) == 2511702,
23: lambda: run23(Array(Input(23))) == 9409 and
run23_2() == 913,
24: lambda: strongest_chain() == 1695 and
strength(max(chains(), key=length_and_strength)) == 1673,
25: lambda: sum(turing(machine(), 'A', 12667664).values()) == 4769
})
# -
# All the days together run in a but less than a minute; only 4 days take more than 5 seconds each; and only 2 take more than 10 seconds.
#
# # Development Time
#
# Here is a plot of the time it took to program solutions each day, for me, the first person to finish, and the hundredth person. My mean time to solve is a little slower than the 100th solver, and five times slower than the first solver.
# +
def plot_times(times):
plt.style.use('seaborn-whitegrid')
X = ints(1, len(times[0]) - 2)
for (mark, label, *Y) in times:
label = '{} (μ={:.0f} min)'.format(label, mean(Y))
plt.plot(X, Y, mark, label=label)
plt.xlabel('Day Number');
plt.ylabel('Minutes to Solve Both Parts')
plt.legend(loc='upper left')
plot_times([
('o--', 'Me',
4, 6, 20, 5, 12, 30, 33, 10, 21, 40, 13, 12, 30, 41, 13, 64, 54, 74, 50, 18, 40, 25, 50, 10, 10),
('v:', '100th',
6, 6, 23, 4, 5, 9, 25, 8, 12, 25, 12, 9, 22, 25, 10, 27, 16, 41, 18, 21, 45, 20, 54, 21, 11),
('^:', '1st',
1, 1, 4, 1, 2, 3, 10, 3, 4, 6, 3, 2, 6, 5, 2, 5, 5, 10, 5, 7, 10, 6, 19, 6, 2)])
# -
# I asked [<NAME>](https://github.com/kevmo314), last year's overall time leader and my colleague at Google, how he manages to go so fast. His answers:
#
# - "My code tends to be eccentrically terse."
# - "I try to minimize the amount of code I write: each line of code is just another chance for a typo."
# - "I save the most time by just observing that a problem is an adaptation of a common problem" (such as a topological sort, or union-find, or A* search, or the Chinese Remainder Theorem).
# - "A lot of it is just finding patterns and not making mistakes."
# - "For AoC it's important to just read the input/output and skip all the instructions first. Especially for the first few days, you can guess what the problem is based on the sample input/output."
|
ipynb/Advent 2017.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %pylab inline
from astropy.io import fits
import astropy.units as u
import pickle as pickle
import os as os
import pandas as pd
from spectools_ir.utils import extract_hitran_data, spec_convol, make_rotation_diagram, get_molmass
from spectools_ir.utils import compute_thermal_velocity, sigma_to_fwhm, fwhm_to_sigma, wn_to_k, spec_convol_R
from spectools_ir.utils import get_miri_mrs_resolution, get_miri_mrs_wavelengths, make_miri_mrs_figure
from spectools_ir.flux_calculator import calc_fluxes, make_lineshape
from spectools_ir.slabspec import make_spec
from spectools_ir.slab_fitter import Config, LineData,Retrieval
from spectools_ir.slab_fitter import corner_plot, trace_plot, find_best_fit, compute_model_fluxes
from spectools_ir.slab_fitter import calc_solid_angle, calc_radius
from spectools_ir.slab_fitter import read_data_from_file, get_samples
import seaborn as sb
import dataframe_image as dfi
# -
import spectools_ir
spectools_ir.__file__
# # Compare Star Properties
star = pd.read_csv("/Users/erichegonzales/Desktop/eriche-thesis/star_properties.csv")
dfi.export(star, 'star_properties.png')
star
#
# # Compare Star Properties and Parameters
data = pd.read_csv("/Users/erichegonzales/Desktop/eriche-thesis/star_data.csv")
#print(data)
data
# +
data_t = data.loc[data['disk_type'] == 'Transitional']
data_c = data.loc[data['disk_type'] == 'Classical']
data_h = data.loc[data['disk_type'] == 'HAeBe']
distance_t = data['distance_au'][data['disk_type'] == 'Transitional']
sol_mass_t = data['solar_mass'][data['disk_type'] == 'Transitional']
sol_lum_t = data['solar_lum'][data['disk_type'] == 'Transitional']
sol_temp_t = data['solar_temp'][data['disk_type'] == 'Transitional']
ntot_t = data['log_ntot'][data['disk_type'] == 'Transitional']
disk_temp_t = data['temp'][data['disk_type'] == 'Transitional']
solid_angle_t = data['log_omega'][data['disk_type'] == 'Transitional']
col_density_t = data['col_density'][data['disk_type'] == 'Transitional']
disk_radius_t = data['disk_radius'][data['disk_type'] == 'Transitional']
disk_area_t = data['disk_area'][data['disk_type'] == 'Transitional']
distance_c = data['distance_au'][data['disk_type'] == 'Classical']
sol_mass_c = data['solar_mass'][data['disk_type'] == 'Classical']
sol_lum_c = data['solar_lum'][data['disk_type'] == 'Classical']
sol_temp_c = data['solar_temp'][data['disk_type'] == 'Classical']
ntot_c = data['log_ntot'][data['disk_type'] == 'Classical']
disk_temp_c = data['temp'][data['disk_type'] == 'Classical']
solid_angle_c = data['log_omega'][data['disk_type'] == 'Classical']
col_density_c = data['col_density'][data['disk_type'] == 'Classical']
disk_radius_c = data['disk_radius'][data['disk_type'] == 'Classical']
disk_area_c = data['disk_area'][data['disk_type'] == 'Classical']
distance_h = data['distance_au'][data['disk_type'] == 'HAeBe']
sol_mass_h = data['solar_mass'][data['disk_type'] == 'HAeBe']
sol_lum_h = data['solar_lum'][data['disk_type'] == 'HAeBe']
sol_temp_h = data['solar_temp'][data['disk_type'] == 'HAeBe']
ntot_h = data['log_ntot'][data['disk_type'] == 'HAeBe']
disk_temp_h = data['temp'][data['disk_type'] == 'HAeBe']
solid_angle_h = data['log_omega'][data['disk_type'] == 'HAeBe']
col_density_h = data['col_density'][data['disk_type'] == 'HAeBe']
disk_radius_h = data['disk_radius'][data['disk_type'] == 'HAeBe']
disk_area_h = data['disk_area'][data['disk_type'] == 'HAeBe']
#(data.loc[data['disk_type'] == 'Transitional'])['solar_mass']
# -
print(data['ntot_pos'].min())
print(data['ntot_pos'].max())
# ax1.set_ylim(ymin=-1.5e24, ymax=1.5e24)
#
# #neg = data['ntot_neg'].str[1:]
# neg = data['ntot_neg'].str[1:][data['disk_type'] == 'HAeBe']
# ntot_neg1 = neg.astype(float)
# ntot_neg2 = 10**ntot_neg1
#
# #pos = data['ntot_pos']
# pos = data['ntot_pos'][data['disk_type'] == 'HAeBe']
# ntot_pos1 = pos.astype(float)
# ntot_pos2 = 10**ntot_pos1
#
# error = ntot_pos2 + ntot_neg2
# #ax1.errorbar(data['solar_mass'], data['col_density'], yerr=error)
# ax1.errorbar(sol_mass_h, col_density_h, yerr=error,fmt ='o', color='darkgreen')
# +
#1
fig=plt.figure(figsize=(8, 5))
ax1=fig.add_subplot(111)
ax1.scatter(sol_mass_t, col_density_t, label='Transitional', marker ='x', s=100)
ax1.scatter(sol_mass_c, col_density_c, label='Classial', marker ='o', s=100)
ax1.scatter(sol_mass_h, col_density_h, label='Herbig Ae/Be', marker ='^', s=100)
ax1.set_xlabel('Stellar Mass [M_☉]', fontsize=14)
ax1.set_ylabel('CO Gas Column Density [cm^2]', fontsize=14)
ax1.set_title('Stellar Mass vs. CO Gas Column Density', fontsize=16)
ax1.set_ylim(ymin=-0.5e23, ymax=1.5e23)
ax1.legend(fontsize=12)
#ax1.legend(loc="upper center", bbox_to_anchor=(0.5, 1.2), ncol=2)
neg = data['ntot_neg']
disk_radius_neg = neg.astype(float)
pos = data['ntot_pos']
disk_radius_pos = pos.astype(float)
error = disk_radius_neg + disk_radius_pos
ax1.errorbar(data['solar_mass'], data['col_density'], yerr=error, xerror=None, fmt ='o', markersize=4, color='firebrick')
# -
# m, b = np.polyfit(sol_mass_t, col_density_t, 1)
# plt.plot(sol_mass_t, m*sol_mass_t + b)
#
# m1, b1 = np.polyfit(sol_mass_t, col_density_t, 1)
# plt.plot(sol_mass_t, m1*sol_mass_t + b1)
# ### Column density stays around constant regardless of stellar mass
# +
#2
fig=plt.figure(figsize=(8, 5))
ax1=fig.add_subplot(111)
ax1.scatter(sol_lum_t, col_density_t, label='Transitional', marker ='x', s=100)
ax1.scatter(sol_lum_c, col_density_c, label='Classial', marker ='o', s=100)
ax1.scatter(sol_lum_h, col_density_h, label='Herbig Ae/Be', marker ='^', s=100)
ax1.set_xlabel('Stellar Luminosity [L_☉]', fontsize=14)
ax1.set_ylabel('CO Gas Column Density [cm^2]', fontsize=14)
ax1.set_title('Stellar Luminosity vs. CO Gas Column Density', fontsize=16)
ax1.set_ylim(ymin=-0.5e23, ymax=1.5e23)
ax1.legend(fontsize=12)
neg = data['ntot_neg']
disk_radius_neg = neg.astype(float)
pos = data['ntot_pos']
disk_radius_pos = pos.astype(float)
error = disk_radius_neg + disk_radius_pos
ax1.errorbar(data['solar_lum'], data['col_density'], yerr=error, xerror=None, fmt ='o', markersize=4, color='firebrick')
# -
# ### Column density stays around constant regardless of stellar luminosity
# +
#3
fig=plt.figure(figsize=(8, 5))
ax1=fig.add_subplot(111)
ax1.scatter(sol_temp_t, col_density_t, label='Transitional', marker ='x', s=100)
ax1.scatter(sol_temp_c, col_density_c, label='Classial', marker ='o', s=100)
ax1.scatter(sol_temp_h, col_density_h, label='Herbig Ae/Be', marker ='^', s=100)
ax1.set_xlabel('Stellar Temperature [K]', fontsize=14)
ax1.set_ylabel('Gas Column Density [cm^2]', fontsize=14)
ax1.set_title('Stellar Temperature vs. Gas Column Density', fontsize=16)
ax1.set_ylim(ymin=-0.5e23, ymax=1.5e23)
ax1.legend(fontsize=12)
neg = data['ntot_neg']
disk_radius_neg = neg.astype(float)
pos = data['ntot_pos']
disk_radius_pos = pos.astype(float)
error = disk_radius_neg + disk_radius_pos
ax1.errorbar(data['solar_temp'], data['col_density'], yerr=error, xerror=None, fmt ='o', markersize=4, color='firebrick')
# -
# neg = data['disk_radius_neg'][data['disk_type'] == 'Classical']
# disk_radius_neg = neg.astype(float)
#
# pos = data['disk_radius_pos'][data['disk_type'] == 'Classical']
# disk_radius_pos = pos.astype(float)
#
# error = disk_radius_neg + disk_radius_pos
# ax1.errorbar(sol_mass_c, disk_radius_c, yerr=error, xerror=None, fmt ='o', color='darkgreen')
# ### Column density stays around constant regardless of stellar temperature
# +
#4
fig=plt.figure(figsize=(8, 5))
ax1=fig.add_subplot(111)
ax1.scatter(sol_mass_t, disk_radius_t, label='Transitional', marker ='x', s=100)
ax1.scatter(sol_mass_c, disk_radius_c, label='Classial', marker ='o', s=100)
ax1.scatter(sol_mass_h, disk_radius_h, label='Herbig Ae/Be', marker ='^', s=100)
ax1.set_xlabel('Stellar Mass [M_☉]', fontsize=14)
ax1.set_ylabel('CO Gas Radius [AU]', fontsize=14)
ax1.set_title('Stellar Mass vs. CO Gas Radius', fontsize=16)
ax1.legend(fontsize=12)
neg = data['disk_radius_neg']
disk_radius_neg = neg.astype(float)
pos = data['disk_radius_pos']
disk_radius_pos = pos.astype(float)
error = disk_radius_neg + disk_radius_pos
ax1.errorbar(data['solar_mass'], data['disk_radius'], yerr=error, xerror=None, fmt ='o', markersize=4, color='firebrick')
# -
# ### Positive correlation between stellar mass and disk radius
# Higher mass means a higher gravitational pull, so more gas is pulled in
# +
#5
fig=plt.figure(figsize=(8, 5))
ax1=fig.add_subplot(111)
ax1.scatter(sol_lum_t, disk_radius_t, label='Transitional', marker ='x', s=100)
ax1.scatter(sol_lum_c, disk_radius_c, label='Classial', marker ='o', s=100)
ax1.scatter(sol_lum_h, disk_radius_h, label='Herbig Ae/Be', marker ='^', s=100)
ax1.set_xlabel('Stellar Luminosity [L_☉]', fontsize=14)
ax1.set_ylabel('CO Gas Radius [AU]', fontsize=14)
ax1.set_title('Stellar Luminosity vs. CO Gas Radius', fontsize=16)
ax1.legend(fontsize=12)
neg = data['disk_radius_neg']
disk_radius_neg = neg.astype(float)
pos = data['disk_radius_pos']
disk_radius_pos = pos.astype(float)
error = disk_radius_neg + disk_radius_pos
ax1.errorbar(data['solar_lum'], data['disk_radius'], yerr=error, xerror=None, fmt ='o', markersize=4, color='firebrick')
# +
#6
fig=plt.figure(figsize=(8, 5))
ax1=fig.add_subplot(111)
ax1.scatter(sol_temp_t, disk_radius_t, label='Transitional', marker ='x', s=100)
ax1.scatter(sol_temp_c, disk_radius_c, label='Classial', marker ='o', s=100)
ax1.scatter(sol_temp_h, disk_radius_h, label='Herbig Ae/Be', marker ='^', s=100)
ax1.set_xlabel('Stellar Temperature [K]', fontsize=14)
ax1.set_ylabel('CO Gas Radius [AU]', fontsize=14)
ax1.set_title('Stellar Temperature vs. CO Gas Radius', fontsize=16)
ax1.legend(fontsize=12)
neg = data['disk_radius_neg']
disk_radius_neg = neg.astype(float)
pos = data['disk_radius_pos']
disk_radius_pos = pos.astype(float)
error = disk_radius_neg + disk_radius_pos
ax1.errorbar(data['solar_temp'], data['disk_radius'], yerr=error, xerror=None, fmt ='o', markersize=4, color='firebrick')
# +
#7
fig=plt.figure(figsize=(8, 5))
ax1=fig.add_subplot(111)
ax1.scatter(sol_mass_t, disk_temp_t, label='Transitional', marker ='x', s=100)
ax1.scatter(sol_mass_c, disk_temp_c, label='Classial', marker ='o', s=100)
ax1.scatter(sol_mass_h, disk_temp_h, label='Herbig Ae/Be', marker ='^', s=100)
ax1.set_xlabel('Stellar Mass [M_☉]', fontsize=14)
ax1.set_ylabel('CO Gas Temperature [K]', fontsize=14)
ax1.set_title('Stellar Mass vs. CO Gas Temperature', fontsize=16)
ax1.set_ylim(ymax = 2500)
ax1.legend(fontsize=12)
neg = data['abs_temp_neg']
disk_radius_neg = neg.astype(float)
pos = data['temp_pos']
disk_radius_pos = pos.astype(float)
error = disk_radius_neg + disk_radius_pos
ax1.errorbar(data['solar_mass'], data['temp'], yerr=error, xerror=None, fmt ='o', markersize=4, color='firebrick')
# +
#8
fig=plt.figure(figsize=(8, 5))
ax1=fig.add_subplot(111)
ax1.scatter(sol_lum_t, disk_temp_t, label='Transitional', marker ='x', s=100)
ax1.scatter(sol_lum_c, disk_temp_c, label='Classial', marker ='o', s=100)
ax1.scatter(sol_lum_h, disk_temp_h, label='Herbig Ae/Be', marker ='^', s=100)
ax1.set_xlabel('Stellar Luminosity [L_☉]', fontsize=14)
ax1.set_ylabel('CO Gas Temperature [K]', fontsize=14)
ax1.set_title('Stellar Luminosity vs. CO Gas Temperature', fontsize=16)
ax1.set_ylim(ymax = 2500)
ax1.legend(fontsize=12)
neg = data['abs_temp_neg']
disk_radius_neg = neg.astype(float)
pos = data['temp_pos']
disk_radius_pos = pos.astype(float)
error = disk_radius_neg + disk_radius_pos
ax1.errorbar(data['solar_lum'], data['temp'], yerr=error, xerror=None, fmt ='o', markersize=4, color='firebrick')
# +
#9
fig=plt.figure(figsize=(8, 5))
ax1=fig.add_subplot(111)
ax1.scatter(sol_temp_t, disk_temp_t, label='Transitional', marker ='x', s=100)
ax1.scatter(sol_temp_c, disk_temp_c, label='Classial', marker ='o', s=100)
ax1.scatter(sol_temp_h, disk_temp_h, label='Herbig Ae/Be', marker ='^', s=100)
ax1.set_xlabel('Stellar Temperature [K]', fontsize=14)
ax1.set_ylabel('CO Gas Temperature [K]', fontsize=14)
ax1.set_title('Stellar Temperature vs. CO Gas Temperature', fontsize=16)
ax1.set_ylim(ymax = 2500)
ax1.legend(fontsize=12)
neg = data['abs_temp_neg']
disk_radius_neg = neg.astype(float)
pos = data['temp_pos']
disk_radius_pos = pos.astype(float)
error = disk_radius_neg + disk_radius_pos
ax1.errorbar(data['solar_temp'], data['temp'], yerr=error, xerror=None, fmt ='o', markersize=4, color='firebrick')
# +
#10
fig=plt.figure(figsize=(8, 5))
ax1=fig.add_subplot(111)
ax1.scatter(col_density_t, disk_radius_t, label='Transitional', marker ='x', s=100)
ax1.scatter(col_density_c, disk_radius_c, label='Classial', marker ='o', s=100)
ax1.scatter(col_density_h, disk_radius_h, label='Herbig Ae/Be', marker ='^', s=100)
ax1.set_xlabel('CO Gas Column Density [cm^2]', fontsize=14)
ax1.set_ylabel('CO Gas Radius [AU]', fontsize=14)
ax1.set_title('CO Gas Column Density vs. CO Gas Radius', fontsize=16)
ax1.set_xlim(xmin = -0.5e23, xmax= 1.5e23)
ax1.legend(fontsize=12)
neg = data['disk_radius_neg']
disk_radius_neg = neg.astype(float)
pos = data['disk_radius_pos']
disk_radius_pos = pos.astype(float)
error = disk_radius_neg + disk_radius_pos
ax1.errorbar(data['col_density'], data['disk_radius'], yerr=error, xerror=None, fmt ='o', markersize=4, color='firebrick')
# +
#11
fig=plt.figure(figsize=(8, 5))
ax1=fig.add_subplot(111)
ax1.scatter(disk_radius_t, disk_temp_t, label='Transitional', marker ='x', s=100)
ax1.scatter(disk_radius_c, disk_temp_c, label='Classial', marker ='o', s=100)
ax1.scatter(disk_radius_h, disk_temp_h, label='Herbig Ae/Be', marker ='^', s=100)
ax1.set_xlabel('CO Gas Radius [AU]', fontsize=14)
ax1.set_ylabel('CO Gas Temperature [cm^2]', fontsize=14)
ax1.set_title('CO Gas Radius vs. CO Gas Temperature', fontsize=16)
ax1.set_ylim(ymax = 2500)
ax1.legend(fontsize=12)
neg = data['abs_temp_neg']
disk_radius_neg = neg.astype(float)
pos = data['temp_pos']
disk_radius_pos = pos.astype(float)
error = disk_radius_neg + disk_radius_pos
ax1.errorbar(data['disk_radius'], data['temp'], yerr=error, xerror=None, fmt ='o', markersize=4, color='firebrick')
# +
#12
fig=plt.figure(figsize=(8, 5))
ax1=fig.add_subplot(111)
ax1.scatter(disk_temp_t, col_density_t, label='Transitional', marker ='x', s=100)
ax1.scatter(disk_temp_c, col_density_c, label='Classial', marker ='o', s=100)
ax1.scatter(disk_temp_h, col_density_h, label='Herbig Ae/Be', marker ='^', s=100)
ax1.set_xlabel('CO Gas Temperature [AU]', fontsize=14)
ax1.set_ylabel('CO Gas Column Density [cm^2]', fontsize=14)
ax1.set_title('CO Gas Temperature vs. CO Gas Column Density', fontsize=16)
ax1.set_ylim(ymin=-0.5e23, ymax=1.5e23)
ax1.legend(fontsize=12)
neg = data['ntot_neg']
disk_radius_neg = neg.astype(float)
pos = data['ntot_pos']
disk_radius_pos = pos.astype(float)
error = disk_radius_neg + disk_radius_pos
ax1.errorbar(data['temp'], data['col_density'], yerr=error, xerror=None, fmt ='o', markersize=4, color='firebrick')
# -
|
data/compare_data/compare_properties.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 4.10 – Practice Problem 1
#
# ## Question
#
# A vessel contains a mixture of benzene, $C_6 H_6$, and toluene, $C_7 H_8$. At 373K the pressure in the vessel is 1.53 bar(abs). Using the data provioded, and any others necessary, find the mass fractions of benzene in the liquid and vapour. Assume the liquids and vapours behave ideally. Recall the formula for Antoines equation: $\log_{10} p^* = A - \frac{B}{T + C}$. Where T is in kelvin for this question.
#
# | Component | Are | Cool | Antoine Constants |
# |-----------|:-------:|---------:|-----------------------------------------------------------|
# | | A | B | C |
# | Benzene | 4.72583 | 1660.652 | -1.461 |
# | Toluene | 4.07827 | 1343.943 | -53.773 |
#
#
# ## Answer
#
# From antoines equation, we can find the vapour partial pressure of toluene and benze.
#
# $$ p^*_b = 10^{\space A - \frac{B}{T + C}} = 10^{\space 4.72583 - \frac{1660.652}{373 - 1.461}} = 1.8037 \space \text{bar}$$
#
# $$ p^*_t = 10^{\space A - \frac{B}{T + C}} = 10^{\space 4.07827 - \frac{1343.943}{373 - 53.773}} = 0.7384 \space \text{bar} $$
#
# Now we can solve for the liquid mole fractions of the two substances with two equations and two unknowns using Raoult's law and the fact that the mole fractions must add to one:
#
# 1. $x_b + x_t = 1$
#
# 2. $P = x_b \cdot p^*_b + x_t \cdot p^*_t$
#
# $$ x_t = 1 - x_b $$
#
# $$ P = x_b \cdot p^*_b + (1 - x_b)\cdot p^*_t $$
#
# $$ 1.53 \space \text{bar} = x_b \cdot (1.8037 \space \text{bar}) + (1 - x_b)\cdot (0.7384 \space \text{bar}) $$
#
# $$ x_b = 0.743 $$
#
# $$ \therefore \space x_t = 0.257 $$
#
# Next we can solve for the vapour mole fractions using Raoult's law:
#
# $$ y_b = \frac{x_b \cdot p^*_b}{P} = \frac{1.340}{1.53} = 0.876 $$
#
# $$ y_t = 1 - y_b = 0.124 $$
#
# Finally we can solve for the mass fractions using the molecular weight of the benzne and toluene and a basis of 1 mole:
#
# $$ MW_b = 78.11 \space \frac{g}{mol} $$
#
# $$ MW_t = 92.14 \space \frac{g}{mol} $$
#
# $$ m_b = x_b \times MW_b = 0.743 \space mol \times 78.11 \space \frac{g}{mol} = 58.03 \space g $$
#
# $$ m_t = x_t \times MW_t = 0.257 \space mol \times 92.14 \space \frac{g}{mol} = 23.68 \space g $$
#
# $$ \therefore \space w_b = \frac{m_b}{m_b + m_t} = \frac{58.03}{58.03 + 23.68} = 0.710 $$
#
# and we will do the same for the vapour mass fractions:
#
# $$ m_b = y_b \times MW_b = 0.876 \space mol \times 78.11 \space \frac{g}{mol} = 68.42 \space g $$
#
# $$ m_t = y_t \times MW_t = 0.124 \space mol \times 92.14 \space \frac{g}{mol} = 11.43 \space g $$
#
# $$ \therefore \space w_b = \frac{m_b}{m_b + m_t} = \frac{68.42}{68.42 + 11.43} = 0.857 $$
|
Modules/Module-4-Separation-II/4.10 - Practice Problem 1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import json
import scipy.interpolate
import matplotlib.pyplot as plt
import xml.etree.ElementTree as ET
from collections import OrderedDict
from pprint import pprint
camera="Xsens"
if camera=="Kinect":
form=".txt"
elif camera=="Xsens":
form=".txt"
file_name="./Données/%s/chris1/chris1_1_transformed%s"%(camera,form)
print(file_name)
with open(file_name) as f:
data = json.load(f, object_pairs_hook=OrderedDict)
Times=list(data['positions'].keys())
positions=data['positions']
fps=60
frames_count=len(list(positions.keys()))
# +
def body_positions(body_Part,Times,positions):
x_bPart_values={}
y_bPart_values={}
z_bPart_values={}
tronq_times=[]
for time in Times:
bParts=list(positions[time].keys())
if body_Part in bParts:
x_bPart_values[time]=positions[time][body_Part][1]
y_bPart_values[time]=positions[time][body_Part][2]
z_bPart_values[time]=positions[time][body_Part][0]
tronq_times.append(time)
tronq_times=np.array(tronq_times)
x_bPart_values_list=list(x_bPart_values.values())
x_bPart_values_array=np.array(x_bPart_values_list)
y_bPart_values_list=list(y_bPart_values.values())
y_bPart_values_array=np.array(y_bPart_values_list)
z=z_bPart_values_list=list(z_bPart_values.values())
z_bPart_values_array=np.array(z_bPart_values_list)
return(x_bPart_values_array,y_bPart_values_array,z_bPart_values_array,tronq_times)
def interpolation(x_bPart_values_array,y_bPart_values_array,z_bPart_values_array,Times_float,new_times_array):
tau = Times_float[-1] - Times_float[0]
#new_times_array = np.arange(0, tau, tau/len(y_bPart_values_array))
#new_times_array = np.arange(0, 1628/60, 1/30)
#new_times_array = np.arange(0, 2*1628/60, 1/30)
new_xbPart_values = np.zeros(new_times_array.shape)
new_ybPart_values = np.zeros(new_times_array.shape)
new_zbPart_values = np.zeros(new_times_array.shape)
y_gen = scipy.interpolate.interp1d(([t-Times_float[0] for t in Times_float]), y_bPart_values_array)
y_gen(new_times_array)
print(len(y_gen(new_times_array)))
for i in range(len(new_times_array)):
new_ybPart_values[i]=y_gen(new_times_array[i])
x_gen = scipy.interpolate.interp1d(([t-Times_float[0] for t in Times_float]), x_bPart_values_array)
x_gen(new_times_array)
for i in range(len(new_times_array)):
new_xbPart_values[i]=x_gen(new_times_array[i])
z_gen = scipy.interpolate.interp1d(([t-Times_float[0] for t in Times_float]), z_bPart_values_array)
z_gen(new_times_array)
for i in range(len(new_times_array)):
new_zbPart_values[i]=z_gen(new_times_array[i])
return(new_xbPart_values,new_ybPart_values,new_zbPart_values,list(new_times_array))
def new_body_positions(body_part,Times,positions,times_array):
x_bPart_values_array,y_bPart_values_array,z_bPart_values_array,tronq_times=body_positions(body_part,Times,positions)
Times_float=[]
for time in tronq_times:
Times_float.append(float(time))
new_xbPart_values,new_ybPart_values,new_zbPart_values,new_Times_float=interpolation(x_bPart_values_array,y_bPart_values_array,z_bPart_values_array,Times_float,new_times_array)
print("t ",len(new_Times_float),"y ",len(new_ybPart_values))
plt.plot(new_Times_float,new_ybPart_values,'red')
plt.title("y values after interpolation %s"%body_part)
plt.show()
plt.plot(new_Times_float,new_xbPart_values,'blue')
plt.title("x values after interpolation %s"%body_part)
plt.show()
new_bPart_Positions=np.stack((new_xbPart_values,new_ybPart_values,new_zbPart_values),axis=-1)
return(new_bPart_Positions,new_Times_float)
def stackPositions(body_Part,Times,positions):
x_bPart_values_array,y_bPart_values_array,z_bPart_values_array=body_positions(body_Part,Times,positions)
All_positions=np.stack((x_bPart_values_array,y_bPart_values_array,z_bPart_values_array),axis=-1)
return(All_positions)
# -
T=float(list(positions.keys())[-1])
# +
bParts=list(list(positions.values())[0].keys())
T=27.69323690681233
#new_body_pos,new_Times_float_mSpine=new_body_positions('mSpine',Times,positions)
new_positions={}
#fps=frames_count/T-1
fps=30
new_times_array = np.arange(0, T, 1/fps)
for time in new_times_array:
new_positions[str(time)]={}
for bpart in bParts:
#if bpart=='mSpine':
# for i in range(len(new_body_pos)):
# new_positions[str(new_Times_float_mSpine[i])][bpart]=list(new_mSpine_positions[i])
#else:
new_body_pos=new_body_positions(bpart,Times,positions,new_times_array)[0]
for i in range(len(new_body_pos)):
new_positions[str(new_times_array[i])][bpart]=list(new_body_pos[i])
# -
interpolated_data={}
interpolated_data['positions']=new_positions
with open("./Données/Xsens/chris1/chris1_1_interpolated.txt", 'w') as outfile:
json.dump(interpolated_data, outfile, sort_keys = True, indent = 4,
ensure_ascii = False)
Times
|
Algo/.ipynb_checkpoints/Interpolation_DataXSens-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.7 64-bit (''my_env'': conda)'
# language: python
# name: python3
# ---
# +
import os
from typing import List
import numpy as np
from PIL import Image
import argparse
import sys
from tqdm import tqdm
from numba import njit
FILENAME = '1B1B1K2-3p1N2-6k1-R7-5P2-4q3-7R-1B6.jpeg'
# -
orig_image = Image.open(f"../data/raw/train/{FILENAME}")
orig_image = np.array(orig_image)
@njit
def crop(image: np.ndarray) -> List[np.ndarray]:
parts = []
for r in range(0, image.shape[0], 50):
for c in range(0, image.shape[1], 50):
parts.append(image[r : r + 50, c : c + 50, :])
return parts
cropped_image = crop(orig_image)
lst = ['s', 's']
lst.extend((ord('3') - 48) * 'E')
lst
@njit
def transform_label(filename):
orig_label = filename.split(".")[0]
transformed_label = []
ranks = orig_label.split("-")
for rank in ranks:
for letter in rank:
if letter in "0123456789":
transformed_label.extend((ord(letter) - 48) * 'E')
else:
transformed_label.append(letter)
return transformed_label
transform_label(FILENAME)
|
notebooks/1.2-hcm-crop-images-faster.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Personal tweets sentiment analysis
#
# ## This paper presents user interaction with the developed model
#
# interaction interface:<br>
# 1: You will need to enter screenname in variable "user_screen_name"<br>
# 2: Enter the required number of current tweets to view in variable "tweets_count"<br>
# 3: Run first three jupyter cells(if you didn't get http: link on the output, run second and third cell again)<br>
# 4: Output of the third cell is a link for visualization in the browser. Open this link, later we will need it.<br>
# 5: Run fourth cell. In the output of this cell, you will see a dataframe. In the "raw_data" column there are actual tweets of the person you are interested in. in the "target" column there are corresponding tweet sentiment scores<br>
# 6: After full cell execution. follow the link obtained in step 4<br>
# ### visualization setup
# 1: In the left part of the interface, find the drop-down list "Color by" and select "Sentiment"<br>
# This will help to distinguish between positively and negatively used words in the context of the speech of person being analyzed<br>
# Also when choosing a word of interest, after it, in brackets is indicated an accurate assessment positivity of the word used in the contest of this person’s speech<br>
# 2: On the right side of the interface, locate the “Search” field. With the help of it you can find the words of interest, and analyze words in the original speech space
user_screen_name = "realDonaldTrump"
tweets_count = 200
tweets_to_print = 10
# %run thread.py
thread_start()
url = ! curl -s http://localhost:4040/api/tunnels | python3 -c \
"import sys, json; print(json.load(sys.stdin)['tunnels'][0]['public_url'])"
print(url[0]+"/#projector")
# %run main.py
main_func(user_screen_name, tweets_count, tweets_to_print)
|
Tweet_sentiment.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Date: 26/12/2020
# # Building a Deep Neural Network from scratch.
#
# This project focuses on building a Deep Neural Network from scratch. The aim of this project is to : -<br>
# 1) Understand the concepts and working of DNN.<br>
# 2) Using ReLU.<br>
# 3) Coding it.<br>
# ## Import Modules
import numpy as np
import h5py
import matplotlib.pyplot as plt
import pandas as pd
import scipy
# %matplotlib inline
# ## Import Data
raw_train_data = h5py.File('train_catvnoncat.h5')
train_keys = list(raw_train_data.keys())
# +
X_train_raw = np.array(raw_train_data[train_keys[1]])
y_train_raw = np.array(raw_train_data[train_keys[2]])
classes = np.array(raw_train_data[train_keys[0]])
# -
print(f'The shape of X_train_raw is:{X_train_raw.shape}')
print(f'The shape of y_train_raw is:{y_train_raw.shape}')
# +
y_train = y_train_raw.reshape((1, y_train_raw.shape[0]))
print(f'The shape of y_train is:{y_train.shape}')
# +
X_train = X_train_raw.reshape((X_train_raw.shape[0], -1)).T
print(f'The shape of X_train is:{X_train.shape}')
# -
# ## Normalization
X_train = X_train/255.0
# ## Model development
# +
'''
Defining number of units in each layers.
Inputs: -
------
units_in_layers - Number of units for each layer in the form of a list.
Output: -
------
Prints number of units in each layer.
'''
units_in_layers = [X_train.shape[0], 5, 1]
total_layers = len(units_in_layers)
print('Number of units in each layers are:')
for i in range(len(units_in_layers)):
print(str(i) + f' layer has {units_in_layers[i]} units')
print('---------------------------------------------------')
print(f'Total number of layers are {total_layers}')
# -
def initialization_func(units_in_layers, total_layers):
'''
This function initializes weights and bias for the layers.
Inputs: -
------
units_in_layers - Number of units for each layer in the form of a list.
total_layers - Total number of layers.
Returns: -
------
weights - dictionary containing weights and bias.
'''
np.random.seed(5)
weights = {}
for i in range(1, total_layers):
weights['W' + str(i)] = np.random.randn(units_in_layers[i],
units_in_layers[i - 1])*0.01
weights['b' + str(i)] = np.zeros((units_in_layers[i],
1))
return weights
def forward_activation(A_ref, W, b, activation):
'''
Implements the forward propagation and calculates the required Activation.
Inputs: -
------
A_ref - Activations from previous layer (Initially it is equal to the input data).
W - Weight associated with the corresponding layer.
b - Bias associated with the corresponding layer.
activation - Name of the activation function to be implemented.
Returns: -
------
a - Activation value for the next layer.
forward_cache - List containing values of A_ref, W, b, Z which is used during
backward propagation.
Example: - For initial layer the forward_cache will be
forward_cache = (A_ref 0, W1, b1 z1)
0 --> refers to the input layer or activation values of the previous layer.
1 --> refers to the values associated with the 1st layer.
'''
if activation == 'sigmoid':
z = np.matmul(W, A_ref) + b
a = 1/(1 + np.exp(-z))
forward_cache = (A_ref, W, b, z)
return a, forward_cache
elif activation == 'relu':
z = np.matmul(W, A_ref) + b
a = np.maximum(0, z)
forward_cache = (A_ref, W, b, z)
return a, forward_cache
# +
def backward_activation_sigmoid(z):
'''
Calculates the derivative of sigmoid activation.
Inputs: -
------
z - Forward propagation value. This value is collected from **forward_cache**
of **forward_activation** function.
Returns: -
------
Derivative of sigmoid function.
'''
sig = 1/(1 + np.exp(-z))
return sig*(1 - sig)
def backward_activation_relu(z):
'''
Calculates the derivative of ReLU activation.
Inputs: -
------
z - Forward propagation value. This value is collected from **forward_cache**
of **forward_activation** function.
Returns: -
------
Derivative of ReLU function.
'''
relu_grad = z.copy()
relu_grad[relu_grad <= 0] = 0
relu_grad[relu_grad > 0] = 1
return relu_grad
# -
def forward_prop_func(weights, X_train):
'''
Implements the forward propagation and calculates the Activation for every layer.
Inputs: -
------
weights - Initial weights calculated from **initialization_func** function.
X_train - Input data.
Returns: -
------
A_last - Activation value of the last layer.
cache - List of tuples containing the values of A_ref, W, b, Z of every layer.
'''
L = len(list(weights.keys()))//2
A_ref = X_train.copy()
cache = []
for i in range(L - 1):
A_next, forward_cache = forward_activation(A_ref, weights['W' + str(i + 1)],
weights['b' + str(i + 1)], activation = 'relu')
A_ref = A_next
cache.append(forward_cache)
A_last, forward_cache = forward_activation(A_ref, weights['W' + str(L)],
weights['b' + str(L)], activation = 'sigmoid')
cache.append(forward_cache)
return A_last, cache
def cost_func(A_last, y_train):
'''
Calculates the cost for a given value of weights.
Inputs: -
------
A_last - Activation value of the last layer calculated
from **forward_prop_func** function.
y_train - Label data.
Returns: -
------
train_cost - Training cost for a given value of weights.
dA_last - Derivative of Loss with respect to the last activation layer.
'''
epsilon = 1e-5
train_cost = (-(y_train*np.log(A_last + epsilon) +
(1 - y_train)*np.log(1 - A_last + epsilon))).mean()
dA_last = (-(np.divide(y_train, A_last + epsilon)
- np.divide((1 - y_train), (1 - A_last + epsilon))))
return train_cost, dA_last
def dZ_func(dA_last, Z, activation):
'''
Calculates derivative of Loss with respect to the forward propagation layer.
Inputs: -
------
dA_last - Derivative of Loss with respect to the last activation layer.
This value is collected from **cost_func** function.
Z - This value is collected from **forward_cache** of **forward_activation** function.
activation - Name of the activation function to be implemented.
Returns: -
------
dZ values.
'''
if activation == 'relu':
return dA_last*backward_activation_relu(Z)
elif activation == 'sigmoid':
return dA_last*backward_activation_sigmoid(Z)
def grad(cache, dA_last, activation):
'''
Calculates gradients.
Inputs: -
------
cache - List of tuples containing the values of A_ref, W, b, Z of every layer.
dA_last - Derivative of Loss with respect to the last activation layer.
This value is collected from **cost_func** function.
activation - Name of the activation function to be implemented.
Returns: -
------
dA_prev - Derivative of Loss with respect to the previous activation layer.
dw_ - Weight gradient.
db_ - Bias gradient.
'''
A_prev, W, b, Z = cache
m = A_prev.shape[1]
dz = dZ_func(dA_last, Z, activation)
dw_ = (1.0/m)*np.matmul(dz, A_prev.T)
db_ = (1.0/m)*(dz).sum(axis = 1, keepdims = True)
dA_prev = np.matmul(W.T, dz)
return dA_prev, dw_, db_
def back_prop_func(A_last, cache, y_train):
'''
Calculates gradients.
Inputs: -
------
A_last - Activation value of the last layer calculated
from **forward_prop_func** function.
cache - List of tuples containing the values of A_ref, W, b, Z of every layer.
y_train - Label data.
Returns: -
------
grads - Dictionary of gradients.
train_cost - Value of training cost.
'''
grads = {}
L = len(cache)
train_cost, dA_last = cost_func(A_last, y_train)
current_cache = cache[-1]
dA_temp, dw_temp, db_temp = grad(current_cache, dA_last, activation = 'sigmoid')
grads['dA' + str(L)] = dA_temp
grads['dW' + str(L)] = dw_temp
grads['dB' + str(L)] = db_temp
for t in reversed(range(L - 1)):
current_cache = cache[t]
grads['dA' + str(t + 1)], grads['dW' + str(t + 1)], grads['dB' + str(t + 1)] = grad(current_cache, grads['dA' + str(t + 2)], activation = 'relu')
return grads, train_cost
def update_parameters(weights, grads, learning_rate):
'''
Updates the weights.
Inputs: -
------
weights - Dictionary containing weights and bias.
grads - Dictionary of gradients. This value is collected from **back_prop_func**.
learning_rate - Value of learning rate.
Returns: -
------
Updated values of weights.
'''
lr = learning_rate
L = len(weights) // 2
for l in range(L):
weights["W" + str(l+1)] = weights["W" + str(l+1)] - lr* grads["dW" + str(l+1)]
weights["b" + str(l+1)] = weights["b" + str(l+1)] - lr * grads["dB" + str(l+1)]
return weights
# ## Final Model
def nn_model(units_in_layers, total_layers, X_train, y_train,
iterations, learning_rate):
'''
Final model.
Inputs: -
------
units_in_layers - Number of units for each layer in the form of a list.
total_layers - Total number of layers.
X_train - Input data.
y_train - Label data.
iterations - Iteration for optimization loop.
learning_rate - Value of learning rate.
Returns: -
------
weights - Learned weights.
training_cost - List of training cost.
'''
weights = initialization_func(units_in_layers, total_layers)
training_cost = []
for i in range(iterations):
A_last_calc, cache = forward_prop_func(weights, X_train, X_test)
grads, train_cost = back_prop_func(A_last_calc, cache, y_train)
weights = update_parameters(weights, grads, learning_rate)
if i%200 == 0:
training_cost.append(train_cost)
return weights, training_cost
# ## Model checking
# +
units_in_layers = [X_train.shape[0], 500, 5, 1]
total_layers = len(units_in_layers)
u, co = nn_model(units_in_layers, total_layers, X_train, y_train,
X_test, y_test, 3000, 0.01)
# -
final_weights = u
training_cost = co
# +
# Plotting the cost
plt.figure(figsize = (8, 5))
plt.plot(training_cost, label = 'Training cost')
plt.title('Variation of Cost', fontsize = 20)
plt.xlabel('Every 200th iteration')
plt.ylabel('Cost')
plt.legend()
plt.show()
# -
# # Conclusion
# The Deep Neural Network code is running correctly and it is capable of reducing the cost.
|
Machine Learning/Building DNN from scratch/Building DNN from scratch.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Make SMIRKS from clustered fragments
#
# This notebook will showcase how `ChemPer`'s `ClusterGraph` creates SMIRKS patterns from a group of user specified molecular fragments.
# For example, imagine we wanted to create a SMIRKS pattern for an angle type that appears in many molecules.
# `ClusterGraph` collects the SMIRKS decorators from every molecule and stores them in a highlyspecific SMIRKS pattern.
#
# The ultimate goal for chemper is to create a hierarchical list of SMIRKS patterns that retains fragment clustering.
# We could use this tool to generate SMIRKS patterns for the SMIRNOFF force field format allowing use to create data driven, direct chemical percpeption.
#
# For example, if your initial clusters had 4 types of carbon-carbon bonds (single, aromatic, double, and triple), you would expect the final SMIRKS patterns to reflect those four categories.
#
# The first step here is to store possible decorators for atoms and bonds in a given cluster. In this notebook we will use example SMIRKS patterns as a way of identifying groups of molecular fragments. Then we will use `ClusterGraph` to create highly specific SMIRKS for these same fragments.
# import statements
from chemper.mol_toolkits import mol_toolkit
from chemper.graphs.cluster_graph import ClusterGraph
from chemper.chemper_utils import create_tuples_for_clusters
# ### create_tuples_for_clusters
#
# This is a utility function inside ChemPer which extracts atom indices which match a specific SMIRKS pattern.
#
# Help on function create_tuples_for_clusters in module chemper.chemper_utils:
# For example, lets assume you wanted to find all of the
# atoms that match this SMIRKS list
# * "any", `'[*:1]~[*:2]'`
# * "single", `'[*:1]-[*:2]'`
#
# In this case, the "any" bond would match all bonds, but then
# the "single" would match all single bonds.
# If you were typing Ethene (C=C) then you expect the double bond
# between carbon atoms 0 and 1 to match "any" bond and all C-H bonds to match "single".
#
# The output in this case would be:
# ```python
# [ ('any', [[ (0, 1) ]] ),
# ('single', [[ (0, 2), (0, 3), (1,4), (1,5) ]] )
# ]
# ```
# ## Clustering from other SMIRKS
#
# This example attempts to show how `ClusterGraph` creates a SMIRKS for already clustered sub-graphs.
#
# Here, we will consider two types of angles around tetrahedral carbon atoms.
# In this hierarchical list `c1` would match ANY angle around a tetrahedral carbon (indicated with the connectivity `X4` on atom `:2`).
# Then `c2` would match angles where both outer atoms are hydrogens, just H-C-H angles, meaning those angles would be assigned `c2` and NOT `c1`.
#
# We will use the utility function `create_tuples_for_clusters` (described above) to identify atoms in each example moleucle that match each of these angle types.
smirks_list = [
("c1", "[*:1]~[#6X4:2]-[*:3]"),
("c2", "[#1:1]-[#6X4:2]-[#1:3]"),
]
for label, smirks in smirks_list:
print(label,'\t',smirks)
# ## Start with a single molecule
#
# For the first example, we will start with just one molecule (ethane) and extract the clusters of atoms matching each angle type.
#
# Ethane has a total of 12 sets of angles, all of which can be categorized by the two SMIRKS patterns `c1` or `c2`
# * 6 with the form H-C-C - type c1
# * 6 with the form H-C-H - type c2
#
# First we need to extract the atoms for each of these categories. We use tuples of atom indices to represent these two clusters which are identified using the `create_tuple_for_cluster` utilities function.
mol = mol_toolkit.MolFromSmiles('CC')
atom_index_list = create_tuples_for_clusters(smirks_list, [mol])
for label, mol_list in atom_index_list:
print(label)
for mol_idx, atom_list in enumerate(mol_list):
print('\tmolecule ', mol_idx)
for atoms in atom_list:
print('\t\t', atoms)
# Next, we will look at the `ClusterGraph` for the set of atoms matching the angle type `c1` (`[*:1]~[#6X4:2]-[*:3]`). `ClusterGraph` works by only storing the unique combination of atom decorators. That means that even though we are using six sets of atoms there is only one set of decorators for each atom in the SMIRKS patterns
c1_atoms = atom_index_list[0][1]
graph = ClusterGraph([mol], c1_atoms)
print(graph.as_smirks())
# ### Adding Layers
#
# Similar to the `ChemPerGraph`s described in the `single_mol_smirks` example. We can add atoms outside those indexed in `ClusterGraph`. This is done with the key word `layers`. The specified number of layers corresponds to the number of bonds away from an indexed atom should be included in the SMIRKS. As with `ChemPerGraph`s, you can also use the keyword `"all"` to include all atoms in a molecule in the SMIRKS pattern. For ethane, this would result in the same SMIRKS as specifying 1 layer:
print("layers = 0")
graph = ClusterGraph([mol], c1_atoms, layers=1)
print(graph.as_smirks())
print('-'*80)
print("layers='all'")
graph = ClusterGraph([mol], c1_atoms, layers='all')
print(graph.as_smirks())
# ## Multiple molecules
#
# Now that you have the general idea, lets consider a more complex case,
# Lets create a `ClusterGraph` for both labels in the `smirks_list` from above for the hydrocarbons listed below.
#
# First we need to create the molecules and use `create_tuple_for_cluster` to find group the angles by category.
smiles = ['CC', 'CCC', 'C1CC1', 'CCCC', 'CC(C)C', 'C1CCC1', 'CCCCC']
mols = [mol_toolkit.MolFromSmiles(s) for s in smiles]
atom_index_list = create_tuples_for_clusters(smirks_list, mols)
for label, mol_list in atom_index_list:
print(label)
for mol_idx, atom_list in enumerate(mol_list):
print('\tmolecule ', mol_idx)
for atoms in atom_list:
print('\t\t', atoms)
# Now lets make a `ClusterGraph` object for both `c1` and `c2`.
# In these patterns you will see lists of decorators on each atom. In the SMIRKS lanage `','` stands for 'OR'. So in the case of `"[#6AH1X4x0!r+0,#6AH2X4x0!r+0:1]"` both decorator sets (`"#6AH1X4x0!r+0"` or `"#6AH2X4x0!r+0"`) could match up with atom `:1`
c1_graph = ClusterGraph(mols, atom_index_list[0][1])
print('c1\n'+'-'*50)
print(c1_graph.as_smirks())
c2_graph = ClusterGraph(mols, atom_index_list[1][1])
print()
print('c2\n'+'-'*50)
print(c2_graph.as_smirks())
# ### Identifying common decorators
#
#
# You might notice that some SMIRKS decorators in each atom list are very similar. For example, all of our atoms are neutral so they all have the decorator `"+0"` to indicate a formal charge of zero.
#
# We can take advantage of these commonalities and group decorators together using the SMIRKS `";"` symbol for ANDing decorators. For example, in `"[#6,#7;+0:1]"` the atom is either carbon (`#6`) or (`,`) nitrogen (`#7`) and (`;`) it has a zero formal charge (`+0`).
#
# In the `ChemPer` graph language you can group like decorators using the keyword `compress`. In that case we get these SMIRKS patterns for `c1` and `c2` instead:
print('c1\n'+'-'*50)
print(c1_graph.as_smirks(compress=True))
print()
print('c2\n'+'-'*50)
print(c2_graph.as_smirks(compress=True))
# ### Adding layers
#
# As shown above we could also add `layers` to the `ClusterGraph`s with multiple molecules.
for l in [1,2,3]:
print('layers = ', l)
c1_graph = ClusterGraph(mols, atom_index_list[0][1], layers=l)
print('c1\n'+'-'*50)
print(c1_graph.as_smirks())
c2_graph = ClusterGraph(mols, atom_index_list[1][1], layers=l)
print()
print('c2\n'+'-'*50)
print(c2_graph.as_smirks())
print('\n', '='*80, '\n')
# ## Where do you go from here
#
# As you see above, the `ClusterGraph` SMIRKS are significantly more complicated and specific than the input SMIRKS.
# For example, the input SMIRKS for `c1` is `[*:1]~[#6X4:2]-[*:3]`,
# however `ClusterGraph` creates this monstrosity:
#
# ```
# [#6AH1X4x0!r+0,#6AH2X4x0!r+0,#6AH2X4x2r3+0,#6AH2X4x2r4+0,#6AH3X4x0!r+0:1]-[#6AH1X4x0!r+0,#6AH2X4x0!r+0,#6AH2X4x2r3+0,#6AH2X4x2r4+0,#6AH3X4x0!r+0:2]-[#1AH0X1x0!r+0,#6AH2X4x0!r+0,#6AH2X4x2r3+0,#6AH2X4x2r4+0,#6AH3X4x0!r+0:3]
# ```
#
# Although this pattern becomes a bit less complex with the compression:
# ```
# [*!rH1x0,*!rH2x0,*!rH3x0,*H2r3x2,*H2r4x2;#6;+0;A;X4:1]-[*!rH1x0,*!rH2x0,*!rH3x0,*H2r3x2,*H2r4x2;#6;+0;A;X4:2]-[#1!rH0X1x0,#6!rH2X4x0,#6!rH3X4x0,#6H2X4r3x2,#6H2X4r4x2;+0;A:3]
# ```
#
# Our goal is to generate a hierarchical list of SMIRKS would could recover the same chemistry in a different list of molecules. In order to do this we would want to generate the SMIRKS patterns for different clusters and then remove unnecessary decorators.
#
# To meet this purpose we created the `SMIRKSifier`. For details on this topic see the notebook `smirksifying_clusters` in this example folder.
|
examples/smirks_from_molecules.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/anicelysantos/tutoriais-dados-realpython/blob/main/python_csv.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="YBbMS-q0s2se"
# Esse tutorial pode ser encontrado aqui https://realpython.com/python-csv/
# + [markdown] id="BHpOBU9uii92"
# # Lendo e escrevendo arquivos CSV em Python
# + colab={"base_uri": "https://localhost:8080/"} id="4mY2jXJJia1t" outputId="5cab18db-546a-41bd-c6ef-e96e56ae91fc"
import csv
with open ('/content/drive/MyDrive/dados_pandas/Real Python/employee_birthday.csv') as csv_file:
csv_reader = csv.reader(csv_file, delimiter=',')
line_count = 0
for row in csv_reader:
if line_count == 0:
print(f'Column names are {",".join(row)}')
line_count += 1
else:
print(f'\t {row[0]} works in the {row[1]} department, and was born in {row[2]}.' )
line_count += 1
print(f'Processed {line_count} lines.')
# + [markdown] id="g2VHgFLVnxKm"
# **Fazendo a mesma leitura usando dicionários**
# + colab={"base_uri": "https://localhost:8080/"} id="2z_sGFr7n2Ct" outputId="0d7a8eb1-2ece-4d57-bfaf-6bc1146be9a3"
import csv
with open ('/content/drive/MyDrive/dados_pandas/Real Python/employee_birthday.csv', mode='r') as csv_file:
csv_reader = csv.DictReader(csv_file)
line_count = 0
for row in csv_reader:
if line_count == 0:
print(f'Column names are {",".join(row)}')
line_count += 1
print(f'\t{row["name"]} works in the {row["department"]} department, and was born in {row["birthday month"]}.')
line_count += 1
print(f'Processed {line_count} lines.')
# + [markdown] id="PVOI9q2fqZb9"
# **Gravando arquivos CSV com `csv`**
#
# + id="m0HG1eJpqWU2"
import csv
with open('employee_file.csv', mode='w') as employee_file:
employee_writer = csv.writer(employee_file, delimiter=',', quotechar='"', quoting = csv.QUOTE_MINIMAL)
employee_writer.writerow(['<NAME>', 'Accounting', 'November'])
employee_writer.writerow(['<NAME>','IT','March'])
# + [markdown] id="mFky97FCuNgt"
# **Usando dicionários para fazer a mesma coisa**
# + id="vM5HSNlbuSdE"
import csv
with open('employee_file2.csv', mode='w') as csv_file:
fieldnames = ['emp_name','dept','birth_month']
writer = csv.DictWriter(csv_file, fieldnames=fieldnames)
writer.writeheader()
writer.writerow({'emp_name':'<NAME>', 'dept':'Accounting','birth_month':'November'})
writer.writerow({'emp_name':'<NAME>', 'dept':'IT','birth_month':'March'})
# + [markdown] id="o5YdGZu_vd8V"
#
# + [markdown] id="o6gvXq-AvgSs"
# # Analisando arquivos CSV com a biblioteca Pandas
# + id="fpAVvhDoKMhM"
import pandas as pd
# + colab={"base_uri": "https://localhost:8080/"} id="n3SCqGrCKOy3" outputId="18027bce-071b-4270-cc7d-fa6c9a6693d9"
df = pd.read_csv('/content/drive/MyDrive/dados_pandas/Real Python/hrdata.csv')
print(df)
# + colab={"base_uri": "https://localhost:8080/"} id="u9qOzvPnLAbQ" outputId="bdccf644-9665-4e5b-98a0-126dfbe9642c"
print(type(df['Hire'][0]))
# + colab={"base_uri": "https://localhost:8080/"} id="2JgvO-5PLN8X" outputId="90a79ce9-35e8-4647-a056-136ba5c6bab2"
#Transformar uma coluna em índice
df = pd.read_csv('/content/drive/MyDrive/dados_pandas/Real Python/hrdata.csv', index_col='Name')
print(df)
# + colab={"base_uri": "https://localhost:8080/"} id="knA7JF_SLnNo" outputId="ba05563e-72a2-4578-b691-00927336be4f"
#Mudar o formato da coluna data para date
df = pd.read_csv('/content/drive/MyDrive/dados_pandas/Real Python/hrdata.csv', index_col='Name', parse_dates=['Hire Date'])
print(df)
# + colab={"base_uri": "https://localhost:8080/"} id="bhw2Fnb4L5vb" outputId="33237a73-4a79-4171-987c-a75abb5e6fed"
print(type(df['Hire Date'][0]))
# + colab={"base_uri": "https://localhost:8080/"} id="-_OidubqMzR4" outputId="d593f80c-96c7-4b15-c648-1a0a7ff314bd"
df = pd.read_csv('/content/drive/MyDrive/dados_pandas/Real Python/hrdata.csv', index_col='Employee', parse_dates=['Hired'], header=0, names=['Employee','Hired','Salary','Sick Days'])
print(df)
# + [markdown] id="G2o3CxIUN5sR"
# **Salvando um CSV**
# + id="xS-7WW5PNitc"
df = pd.read_csv('/content/drive/MyDrive/dados_pandas/Real Python/hrdata.csv', index_col='Employee',parse_dates=['Hired'],header=0, names=['Employee','Hired','Salary','Sick Days'])
# + id="zGBw-g3HOb5P"
df.to_csv('hrdata_modified.csv')
# + colab={"base_uri": "https://localhost:8080/", "height": 234} id="pD3nsnipOqa7" outputId="867c7ed8-24c3-466e-9e47-b13d8e266f9c"
df_m = pd.read_csv('/content/hrdata_modified.csv')
df_m
|
python_csv.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import multiprocessing as mp
import itertools as it
import functools as ft
import pickle
import sys
import numpy as np
import pandas as pd
import time
import sklearn
import sklearn.preprocessing as pre
import scipy.sparse as sp
(set([1, 2]), set([3, 4]))
temp = lambda x,y: frozenset(x | y)
ft.reduce(temp, list(map(lambda x: frozenset([x]), [1,2,3,4])))
{**{1:'a', 2:'b'}, **{2:'b', 3:'d'}}
# +
def get_cell_sets(row, oe_csr):
return oe_csr[row['lower']:row['upper']].sum(axis=0)
def first_candidates(cells, cell_sets, min_shared_cells):
count_filter = cell_sets.apply(np.sum) > min_shared_cells
return list(map(lambda x: frozenset([x]), ((cells[count_filter])))), {frozenset([x]):y for x,y in cell_sets[count_filter].to_dict().items()}
def intersector(tuple_of_candidates, cell_sets):
new_candidate_maker = lambda x,y: x | y
return new_candidate_maker(*tuple_of_candidates), np.logical_and(cell_sets[tuple_of_candidates[0]], cell_sets[tuple_of_candidates[1]])
def intersect_chunk(chunk_of_tuples, cell_sets, min_shared_cells, q):
new_cell_set = dict([intersector(x, cell_sets) for x in chunk_of_tuples])
new_cell_set = {x:y for x,y in new_cell_set.items() if np.sum(y) > min_shared_cells}
q.put(new_cell_set)
return
def pickle_cells(cells, cell_sets, k):
'''These files are gonna be decently big. Do not want to keep them in memory.'''
with open('cell_' + str(k) + '.pickle', 'wb') as f:
pickle.dump(cells, f, pickle.HIGHEST_PROTOCOL)
with open('cell_sets_' + str(k) + '.pickle', 'wb') as f:
pickle.dump(cell_sets, f, pickle.HIGHEST_PROTOCOL)
# -
def fast_gather_gene_sets(dat, min_shared_cells = 100, min_percent_cells = None, max_cluster_size = sys.maxsize):
st = time.time()
begin = st
cores = max(mp.cpu_count()-1, 1)
total_cells = dat['barcode'].nunique()
if(min_percent_cells is not None):
min_shared_cells = int(min_percent_cells * total_cells)
cell_id_dict = {y:x for x,y in enumerate(dat['symbol'].unique())}
dat['symbol'] = dat['symbol'].map(cell_id_dict)
cells = dat['symbol'].unique()
dat.sort_values(by='symbol',inplace=True)
slices = pd.DataFrame(dat.groupby('symbol').count().cumsum())
slices.columns = ['upper']
slices['lower'] = [0] + list(slices['upper'])[0:-1]
lab_enc = pre.LabelEncoder()
one_hot = pre.OneHotEncoder(categories='auto')
oe_data = one_hot.fit_transform((lab_enc.fit_transform(dat['barcode'].values)).reshape(-1,1))
get_cell_partial = ft.partial(get_cell_sets, oe_csr=oe_data)
cell_sets = slices.apply(get_cell_partial, axis=1)
en = time.time()
print('Formatted data in ' + str(en-st) + ' seconds')
cells, cell_sets = first_candidates(cells, cell_sets, min_shared_cells)
print(str(len(cells)) + ' genes made have > ' + str(min_shared_cells) + ' cells')
k = 2
n = len(cells)
pickle_cells(cells, cell_sets, k)
while(len(cells) > 0 and k < max_cluster_size):
st = time.time()
candidates_iter = filter(lambda x: len(set(x)) == k, it.combinations(cells, 2))
q = mp.JoinableQueue()
kwarg_dict = {'cell_sets':cell_sets, 'min_shared_cells':min_shared_cells, 'q':q}
for i in range(cores-1):
p = mp.Process(target=intersect_chunk, args=(it.islice(candidates_iter, n // cores),), kwargs=kwarg_dict)
p.start()
p = mp.Process(target=intersect_chunk, args=(candidates_iter,), kwargs=kwarg_dict)
p.start()
out = []
for i in range(cores):
out.append(q.get())
q.task_done()
print('Done with queue!')
q.join()
q.close()
cell_sets = ft.reduce(lambda x, y: {**x, **y}, out)
cells = list(cell_sets.keys())
k+= 1
n = len(cells)
en = time.time()
print('Found ' + str(n) + ' remaining genes with > ' + str(min_shared_cells) + ' of size: ' +str(k-1))
print('Iteration took: ' + str(en-st) + ' seconds')
if(n == 0):
print('Terminated! Total run time: ' + str(en - begin) + ' seconds')
else:
pickle_cells(cells, cell_sets, k-1)
dat = pd.read_csv('./cord_blood_kinases.csv', sep=',', header=0, index_col=0)
fast_gather_gene_sets(dat, min_percent_cells = 0.01)
|
demos/fast_gather_gene_sets.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Load dataset
from pandas_overview import DataFrameSummary, DataFrameOverview
import pandas as pd
df = pd.read_excel("../data/gap_58_dataset.xlsx")
df.head()
# # Create DataFrameSummary
dfs = DataFrameSummary(df)
dfs.columns_types
dfs.types
# # summary()
dfs.summary()
dfs.summary(columns=['WELL_LABEL', 'DP_PRES_DROP'])
# # type_summary()
dfs.type_summary('numeric')
dfs.type_summary('constant')
# # columns_of_type()
dfs[dfs.columns_of_type("constant")]
dfs[dfs.columns_of_type("unique")]
# # DataFrameOverview.overview()
DataFrameOverview.overview(dfs, first_level=1)
|
notebooks/01 - GAP network scan.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Reviews Classification with BERT Using Pytorch
# Bidirectional Encoder Representations from Transformers. BERT is a **text representation technique** which is a fusion of variety of state-of-the-art deep learning algorithms, such as bidirectional encoder LSTM and Transformers.
# + jupyter={"outputs_hidden": true}
import torch
import pandas as pd
from tqdm.notebook import tqdm
from sklearn.model_selection import train_test_split
# + jupyter={"outputs_hidden": true}
df = pd.read_csv('Data\cleaned_reviews.csv')
df.head()
# + jupyter={"outputs_hidden": true}
index = []
for i,j in enumerate(df['reviews']):
index.append(i)
df['index'] = index
# + jupyter={"outputs_hidden": true}
X_train, X_val, y_train, y_val = train_test_split(df[:1000].index.values,
df[:1000].label.values,
test_size = .25,
random_state = 14,
stratify = df[:1000].label.values)
# + jupyter={"outputs_hidden": true}
df['data_type'] = ['not_set']*df.shape[0]
# + jupyter={"outputs_hidden": true}
df.loc[X_train , 'data_type'] = 'train'
df.loc[X_val , 'data_type'] = 'val'
# + jupyter={"outputs_hidden": true}
from transformers import BertTokenizer
from torch.utils.data import TensorDataset
# + jupyter={"outputs_hidden": true}
tokenizer = BertTokenizer.from_pretrained('bert-base-uncased',
do_lower_case = True
)
# + jupyter={"outputs_hidden": true}
encoded_train_data = tokenizer.batch_encode_plus(
df[df.data_type == 'train'][:1000].reviews.values,
add_special_tokens=True,
return_attention_mask=True,
pad_to_max_length=True,
max_length=256,
return_tensors='pt'
)
encoded_val_data = tokenizer.batch_encode_plus(
df[df.data_type == 'val'][:1000].reviews.values,
add_special_tokens=True,
return_attention_mask=True,
pad_to_max_length=True,
max_length=256,
return_tensors='pt'
)
input_ids_train = encoded_train_data['input_ids']
attention_mask_train = encoded_train_data['attention_mask']
labels_train = torch.tensor(df[df.data_type == 'train'][:1000].label.values)
input_ids_val = encoded_val_data['input_ids']
attention_mask_val = encoded_val_data['attention_mask']
labels_val = torch.tensor(df[df.data_type == 'val'][:1000].label.values)
# + jupyter={"outputs_hidden": true}
train_dataset = TensorDataset(input_ids_train, attention_mask_train, labels_train)
val_dataset = TensorDataset(input_ids_val, attention_mask_val, labels_val)
# + jupyter={"outputs_hidden": true}
len(train_dataset), len(val_dataset)
# + jupyter={"outputs_hidden": true}
from transformers import BertForSequenceClassification
# + jupyter={"outputs_hidden": true}
model = BertForSequenceClassification.from_pretrained(
'bert-base-uncased',
num_labels = 2,
output_attentions = False,
output_hidden_states = False
)
# + jupyter={"outputs_hidden": true}
from torch.utils.data import DataLoader, RandomSampler, SequentialSampler
# + jupyter={"outputs_hidden": true}
data_loader_train = DataLoader(
train_dataset,
sampler = RandomSampler(train_dataset),
batch_size = 32
)
data_loader_val = DataLoader(
val_dataset,
sampler = RandomSampler(val_dataset),
batch_size = 32
)
# + jupyter={"outputs_hidden": true}
from transformers import AdamW, get_linear_schedule_with_warmup
# + jupyter={"outputs_hidden": true}
optimizer = AdamW(
model.parameters(),
eps=1e-8,
lr=1e-5
)
# + jupyter={"outputs_hidden": true}
epochs = 2
scheduler = get_linear_schedule_with_warmup(
optimizer,
num_warmup_steps=0,
num_training_steps=len(data_loader_train)*10 # 10: number of epochs....
)
# + jupyter={"outputs_hidden": true}
import numpy as np
# + jupyter={"outputs_hidden": true}
from sklearn.metrics import f1_score
# + jupyter={"outputs_hidden": true}
def f1_score_func(preds, labels):
preds_flat = np.argmax(preds, axis = 1).flatten()
labels_flat = labels.flatten()
return f1_score(labels_flat, preds_flat, average='weighted')
# + jupyter={"outputs_hidden": true}
def accuracy_per_class(preds, labels):
labels_inv = {v : k for k, v in labels.items()}
preds_flat = np.argmax(preds, axis = 1).flatten()
labels_flat = labels.flatten()
for label in np.unique(labels_flat):
y_preds = preds_flat[labels_flat==label]
y_true = labels_flat[labels_flat==label]
print(f'class: {label_inv[label]}')
print(f'accuracy: {len(y_pred[y_preds == label])}/{len(y_ture)}\n')
# + jupyter={"outputs_hidden": true}
import random
seed_val = 17
random.seed(seed_val)
np.random.seed(seed_val)
torch.manual_seed(seed_val)
torch.cuda.manual_seed_all(seed_val)
# + jupyter={"outputs_hidden": true}
def evaluate(dataloader_val):
model.eval()
loss_val_total = 0
predictions, true_vals = [], []
for batch in dataloader_val:
batch = tuple(b for b in batch)
inputs = {'input_ids': batch[0],
'attention_mask': batch[1],
'labels': batch[2],
}
with torch.no_grad():
outputs = model(**inputs)
loss = outputs[0]
logits = outputs[1]
loss_val_total += loss.item()
logits = logits.detach().cpu().numpy()
label_ids = inputs['labels'].cpu().numpy()
predictions.append(logits)
true_vals.append(label_ids)
loss_val_avg = loss_val_total/len(dataloader_val)
predictions = np.concatenate(predictions, axis=0)
true_vals = np.concatenate(true_vals, axis=0)
return loss_val_avg, predictions, true_vals
# + jupyter={"outputs_hidden": true}
# model training loop.. it takes a very long time .
# in this cell we re-train the BERT model on our dataset.
for epoch in tqdm(range(1, epochs+1)):
model.train()
loss_train_total = 0
progress_par = tqdm(data_loader_train,
desc = 'Epoch {:1d}'.format(epoch),
leave = False,
disable = False
)
for batch in progress_par:
model.zero_grad()
batch = tuple(b for b in batch)
inputs = {
'input_ids' :batch[0],
'attention_mask' :batch[1],
'labels' :batch[2]
}
outputs = model(**inputs)
loss = outputs[0]
loss_train_total += loss.item()
loss.backward()
torch.nn.utils.clip_grad_norm_(model.parameters(), 1.0)
optimizer.step()
scheduler.step()
progress_par.set_postfix({'training_loss' : '{:.3f}'.format(loss.item()/len(batch))})
torch.save(model.state_dict(), f'Models/BERT_ft_epoch{epoch}.model')
tqdm.write(f'epoch: {epoch}')
loss_train_avg = loss_train_total/len(data_loader_train)
val_loss, preds, true_vals = evaluate(data_loader_val)
val_f1 = f1_score_func(preds, true_vals)
tqdm.write(f'accuracy: {val_f1} - train_loss: {loss_train_avg} - val_loss: {val_loss}')
|
BERT with Pytorch.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# # Equation of State (EOS) for a Real Gas
# ## Chem 113, Spring 2021, <NAME>
# In General Chemistry, you have learned the ideal gas law:
#
# $$p V = n R T.$$
#
# If we define molar volume as
#
# $$ \bar{V}=\frac{V}{n},$$
#
# the ideal gas law reads
#
# $$ P \bar{V} = R T \;\;\;\textrm{or}\;\;\; P = \frac{RT}{\bar{V}}.$$
#
# In order to quantify the non-ideality of a real gas, we define *compressibility factor* as
#
# $$ Z=\frac{P \bar{V}}{RT}. $$
#
# load some libraries for numerical operations and plotting
import numpy as np
from scipy.optimize import curve_fit
import matplotlib.pyplot as plt
# %matplotlib inline
# Set the parameters first (*units are very important*)
# Parameters
R = 0.082058 # L atm / (mol K)
T = 300.0 # K
# Load experimental data at 300 K (Ref: [<NAME>, <NAME>, and <NAME>. J. Comput. Phys. 17 (1975) 401](https://www.sciencedirect.com/science/article/pii/002199917590042X))
# +
# Load the data
exp = np.loadtxt('Z_300.txt') # the exp data file has 2 columns
P_exp = exp[:,0] # the 1st column is the pressure in the unit of atm
Z_exp = exp[:,1] # the 2nd column is Z
# take a look at the data
plt.plot(P_exp, Z_exp, 'r-o', label="Exp")
plt.xlabel('Pressure (atm)')
plt.ylabel('Z')
plt.xlim((0,500))
plt.ylim((0.9,1.2))
plt.legend()
plt.show()
# -
# Compute $\bar{V}$ first from $Z$
#
# $$\bar{V} = \frac{ZRT}{P}$$
# compute Vbar from Z, Vbar is in the unit of L/mol
Vbar = np.divide(Z_exp * R * T, P_exp)
# In order to fit the $P$ vs. $\bar{V}$ according to the vdW equation, we have to define a function for vdw equation.
#
# $$ P = \frac{RT}{\bar{V}-b}-\frac{a}{\bar{V}^2} $$
# +
def vdw(x, a, b):
'''
x: Vbar, molar volume
a, b: vdW parameters
'''
return R*T/(x-b)-a/x/x
popt, pcov = curve_fit(vdw, Vbar, P_exp, [1.3, 0.03])
a = popt[0]
b = popt[1]
print("a value from fit (L^2 atm / mol^2):", a)
print("b value from fit (L / mol):", b)
# -
# compute predicted P and Z from vdw equation
P_vdw = vdw(Vbar,*popt)
Z_vdw = np.multiply(P_vdw, Vbar)/R/T
# Plot the results and compare them
plt.plot(Vbar, Z_exp, 'ro', label="Exp")
plt.plot(Vbar, Z_vdw, 'b-', label="vdW")
plt.xlabel('Molar volume (L/mol)')
plt.ylabel('Z')
plt.xlim((0,1.5))
plt.ylim((0.9,1.2))
plt.legend()
plt.show()
# Figure out the rough size of the molecule, using
#
# $$ b = \frac{2\pi \sigma^3}{3}N_A $$
# make sure the units are correct
# b: L/mol = dm^3 / mol
# NA = 6.02214076e23 /mol
# sigma will be in dm, needs to time 10^9 to Angstrom
NA = 6.02215076e23
d = (3.0*b/2/np.pi/NA)**(1.0/3.0)*1e9
print("The rough estimate of the diameter of the molecule in Angstrom:", d)
# This molecule is Argon, and its $\sigma$ is commonly taken to be 3.41 Angstrom (Table 16.7), and the parameter $b$ for Ar is 0.03183 L/mol (Table 16.3). Our simple estimates based on a single set of experimental data are reasonably good, slightly underestimating both $b$ and $\sigma$.
|
courses/Ch16_EOS/eos.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import scipy as sp
import scipy.special
import pandas as pd
import scipy.stats
import numpy as np
# # <font face="gotham" color="purple"> Point Estimator
# To estimate the population parameters (e.g. average income of man with a master degree), we compute corresponding characteristics of the sample, referred to as **sample statistic**.
#
# Any types of single value sample statistic is called **point estimator**, previously we have seen **Sample mean, standard deviation, variance**, which all are point estimators of populations. For instance, $\bar{x}$ is the point estimator of $\mu$ and $s^2$ the point estimator of $\sigma^2$.
# # <font face="gotham" color="purple"> Sampling Distribution
# The pointer estimators themselves are random variables. For instance we have 10 samples of household income, so each sample has a mean value, which are most probably different than each other, therefore the sample mean is essentially a random variable.|
#
# The distribution of point estimator is called the **sampling distribution**, i.e. the distribution of sample statistics. The knowledge of its properties will enable us to make statements about how close the point estimates (e.g. sample mean) to the population parameters.
# ## <font face="gotham" color="purple"> Sampling Distribution of $\bar{X}$
# If point estimators are unbiased, then **mean of sampling distribution** and **standard error of the mean** are:
# $$
# E(\bar{X})=\mu\\
# \sigma_{\bar{X}}=\frac{\sigma}{\sqrt{n}}\sqrt{\frac{N-n}{N-1}}
# $$
# The name of _standard error_ generally is used when referring to the standard deviation of a sampling distribution.
#
# To illustrate the sampling distribution of $\bar{X}$, we will generate a population of $100000$ people's height with standard deviation of $3$, mean of $170$cm.
mu = 170
sigma = 3
pop_size = 100000
pop_height = sp.stats.norm.rvs(mu, sigma, size = pop_size)
# Now pretend that we know nothing of the population, but we are able to draw as many samples as possible with zero cost (this will never be possible in real world, but perfect illustration to sampling distribution). After drawing each sample we calculate the sample mean then append onto the list ```sample_mean```, finally we plot the histogram.
#
# Here we plot six sampling distributions with increasing sample sizes, the larger the sample size $n$, the smaller the standard errors are. The red dashed line is a normal distribution $z\sim N(170, 3)$, in contrast, the histogram of sample mean is much concentrated.
# +
sample_sizes = [2**i for i in range(1, 7)]
number_of_sample = 1000
sample_means = []
norm_pdf_x = np.linspace(160, 180, 100)
norm_pdf_y = sp.stats.norm.pdf(norm_pdf_x, mu, sigma)
fig, axs = plt.subplots(nrows=3, ncols=2, figsize=(18, 18))
for indx, ax in enumerate(axs.flatten()):
for i in range(number_of_sample):
sample_means.append(np.mean(np.random.choice(pop_height, size = sample_sizes[indx])))
n, bins, patches = ax.hist(sample_means, bins = 100, label = 'Sampling Distribution of Mean of Height', density= True)
ax.axvline(x = mu, color = 'tomato', lw = 3)
ax.plot(norm_pdf_x, norm_pdf_y, 'r--', label="$z\sim N(170, 3)$")
ax.set_xlim(160, 180)
ax.annotate('number of sample: {}\nsample size: {}\nstandard error: {}'
.format(10000, sample_sizes[indx], round(np.std(sample_means, ddof=1), 2)), (160.5, np.max(n)*.90), size = 11)
fig.suptitle('Sampling Distribution Of Means With Various Sample Size', y = 0.9, size = 18)
plt.show()
# -
# From $\sigma_{\bar{x}}=\frac{\sigma}{\sqrt{n}}\sqrt{\frac{N-n}{N-1}}$, we can see as $n\rightarrow N$, then $\sigma_{\bar{x}}\rightarrow 0$.
#
# Though raising sample size can contract the variation of sampling distribution, but efficiency drops relatively fast. Take a look at graph of $\sqrt{n}$, the slope is steep in the beginning, then flattens as the sample size increases.
x = np.linspace(0, 10, 1000)
y = np.sqrt(x)
fig, ax = plt.subplots(figsize=(12,5))
ax.plot(x, y, color = 'tomato', lw = 3)
ax.set_ylim(0, 4)
ax.grid()
ax.set_title('$y=\sqrt{n}$', size = 18)
plt.show()
# There are also sampling distribution of sample variances.
# +
sample_sizes = [2**i for i in range(1, 18,3)]
number_of_sample = 1000
sample_vars = []
fig, axs = plt.subplots(nrows=3, ncols=2, figsize=(18, 18))
for indx, ax in enumerate(axs.flatten()):
for i in range(number_of_sample):
sample_vars.append(np.var(np.random.choice(pop_height, size = sample_sizes[indx]), ddof = 1))
n, bins, patches = ax.hist(sample_vars, bins = 100, label = 'Sampling Distribution of Variance of Height', density= True)
ax.axvline(x = np.mean(sample_vars), color = 'tomato', lw = 3)
ax.annotate('number of sample: {}\nsample size: {}\nstandard error: {:.2f}\nmean: {:.2f}'
.format(10000, sample_sizes[indx], np.std(sample_vars, ddof=1), np.mean(sample_vars)), (30, np.max(n)*.80), size = 11)
fig.suptitle('Sampling Distribution Of Variances With Various Sample Size', y = 0.9, size = 18)
plt.show()
# -
# # <font face="gotham" color="purple"> Central Limit Theorem
# We mentioned above that properties of sampling distribution can help us making meaningful inferences about population. The most important property is **Central Limit Theorem** which guarantees that with large sample size, we can safely assume the sample mean follows a normal distribution. With this information, we can make statistical inferences such as how far away the sample mean is from the population mean?
#
# Here is the classical definition:
#
# $\{\bar{X}_1,...\bar{X}_2\}$ is a random i.i.d. sequence drawn from an unknown distribution whose $E(X_i)=\mu$ and $\text{Var}(X_i)=\sigma^2$, as $n\rightarrow \infty$, we have
# $$
# \sqrt{n}(\bar{X}_n-\mu)\xrightarrow[]{d} N(0, \sigma^2)
# $$
#
# Simply speaking, sampling distribution of sample mean will converge to a normal distribution when sample size is large enough. The figure below shows three different distributions other than normal distribution, as the sample size increases their sampling distributions converge to normal distribution.
#
# The code is in module ```plot_material```.
from plot_material import central_limit_theorem_plot
central_limit_theorem_plot()
# # <font face="gotham" color="purple"> Sample Size and Estimation Accuracy
# Here is a concrete example that demonstrates how sample sizes influence the estimation accuracy.
#
# To estimate the average height of female in the city of Helsinki, we measure the height of $n = 30$ female adults, we have the sample mean but we also would like to know if the sample mean is within $\pm 1.5cm$ of population mean. And we need to have a key but unrealistic assumption: we know the population standard deviation of the height: $\sigma = 6 cm$.
#
# Step 1: Because we know the $\sigma$, we can calculate the standard error of the mean by using
#
# \begin{equation}
# \sigma_{\bar{x}}=\frac{\sigma}{\sqrt{n}}= \frac{6}{\sqrt{30}}
# \end{equation}
#
#
sigma = 6
std_err = sigma/np.sqrt(30)
std_err
# Step 2: Calculate of range in terms of standard errors.
z_right = 1.5/std_err # plus or minus zr/zl units of std errors
z_left = -1.5/std_err
p = sp.stats.norm.cdf(z_right)-sp.stats.norm.cdf(z_left)
print('The probability of sample mean (sample size = 30) being within 1.5cm of pop mean is {0:.2f}%.'.format(p*100))
# Now let's increase the sample size to 60 persons.
# +
sigma = 6
ss = 60
std_err = sigma/np.sqrt(ss)
zr = 1.5/std_err # plus or minus zr/zl units of std errors
zl = -1.5/std_err
p = sp.stats.norm.cdf(zr)-sp.stats.norm.cdf(zl)
print('The probability of sample mean (sample size = {0}) being within 1.5cm of pop mean is {1:.2f}%.'.format(ss, p*100))
# -
# And this figure can show how the sample size can influence the estimation accuracy. We will come back to similar topics in interval estimations.
sigma = 6
p = np.zeros(100)
for i in range(2, 100):
std_err = sigma/np.sqrt(i)
zr = 1.5/std_err # plus or minus zr/zl units of std errors
zl = -1.5/std_err
p[i] = sp.stats.norm.cdf(zr)-sp.stats.norm.cdf(zl)
fig, ax = plt.subplots(figsize = (12, 7))
ax.plot(np.arange(100), p, lw = 3, color = 'r')
ax.set_xlabel('Sample Size', size = 16)
ax.set_ylabel('Probability', size = 16)
ax.set_title('The probability of sample mean being within 1.5cm of pop mean',size = 18)
plt.show()
# # <font face="gotham" color="purple"> Interval Estimation With Known $\sigma$
# The example above is exactly an **interval estimation** of population mean with known $\sigma$. We will dive deeper here, once you understand the logic of interval estimation, you will have much easier time in later chapters.
#
# We have talked about _Central Limit Theorem_ which guarantees that $\bar{X}$ is normally distributed with sufficiently large samples. But we almost always convert $\bar{X}$ into a standard normal distribution before making any inferences.
# $$
# Z=\frac{\bar{x}-\mu}{\sigma/\sqrt{n}}
# $$
# Essentially, it is a $z$-score of $\bar{X}$ as we mentioned in chapter 1. To make any meaningful interval estimation, we need a notation $z_{\alpha/2}$, such as $z_{0.025}=1.96$. It means the area to the right of $1.96$ is $0.025$.
#
# To establish an interval
# $$
# P\bigg(-z_{\alpha/2} < \frac{\bar{x}-\mu}{\sigma/\sqrt{n}} < z_{\alpha/2}\bigg)=.95
# $$
# With a bit rearrangement, which means that with $1-\alpha$ probability the $X$ will fall in this range.
# $$
# P\bigg(\mu-z_{\alpha/2}\frac{\sigma}{\sqrt{n}} < \bar{x}< \mu+z_{\alpha/2}\frac{\sigma}{\sqrt{n}}\bigg)=1-\alpha
# $$
#
# The sample mean of women's height in Helsinki will fall in
# $$
# P\bigg(170-1.96\frac{6}{\sqrt{30}} < \bar{x}< 170+1.96\frac{6}{\sqrt{30}}\bigg)=.95
# $$
print(170-1.96*6/np.sqrt(30))
print(170+1.96*6/np.sqrt(30))
# However expression above is less common in practice, more commonly $\mu$ is set as the centre of the interval, a slight rearrangement gives us
# $$
# P\bigg(\bar{x}-Z_{\alpha/2}\frac{\sigma}{\sqrt{n}}<\mu<\bar{x}+Z_{\alpha/2}\frac{\sigma}{\sqrt{n}}\bigg)
# $$
#
# this is officially called **confidence interval estimator of population mean**. It states that there is $1-\alpha$ probability that $\bar{X}$ will equal to a value such that the interval will include the population mean.
# ## <font face="gotham" color="purple"> Example of Sleeping Hours
# Suppose we have a sample of ten old person (>70 years old) whose average sleeping time recorded as below
sleep_time = [8.3, 6.9, 4.3, 10.8, 7.9, 9.6, 6.8, 5.6, 7.7]
# The standard deviation of the population sleeping time is $2$, enough information to calculate the confidence interval of $95\%$.
LCL = np.mean(sleep_time)-1.96*2/np.sqrt(len(sleep_time))
UCL = np.mean(sleep_time)+1.96*2/np.sqrt(len(sleep_time))
print('Confidence interval of 95% is ({:.2f}, {:.2f}).'.format(LCL, UCL))
# Because the sample size is considerably small, the confidence interval doesn't provide much info either.
#
# Here we've created a class for population of old people's sleeping time with some methods for easy computation of interval estimators, i.e. ```conf_interval``` for _known $\sigma$_ and ```conf_interval_t``` for _unknown $\sigma$_.
class OldPeopleSleep:
def __init__(self, mean, std, pop_size):
self.std = std
self.population = sp.stats.norm.rvs(loc=mean, scale=std, size=pop_size)
def sampling(self, size):
self.sample = np.random.choice(self.population, size)
def conf_interval(self, confidence_level, two_tail = True):
'''For simplicity, we only show a two tail confidence interval.'''
if two_tail == True:
ppf = 1 - (1 - confidence_level)/2
z = round(sp.stats.norm.ppf(ppf), 2)
LCL = np.mean(self.sample)-z*self.std/np.sqrt(len(self.sample))
UCL = np.mean(self.sample)+z*self.std/np.sqrt(len(self.sample))
return LCL, UCL
def conf_interval_t(self, confidence_level, two_tail = True):
'''For simplicity, we only show a two tail confidence interval.'''
if two_tail == True:
ppf = 1 - (1 - confidence_level)/2
t = round(sp.stats.t.ppf(ppf, df = len(self.sample)-1), 2)
LCL_t = np.mean(self.sample)-t*np.std(self.sample, ddof = 1)/np.sqrt(len(self.sample))
UCL_t = np.mean(self.sample)+t*np.std(self.sample, ddof = 1)/np.sqrt(len(self.sample))
return LCL_t, UCL_t
# Generate a population and take a sample of 100 persons, calculate the confidence interval
population_mean = 7.5
old_ppl = OldPeopleSleep(population_mean, std=2, pop_size=100000)
old_ppl.sampling(100)
old_ppl.conf_interval(.95, two_tail=True)
# We will simulate repetitive sampling and show that $95\%$ of time the confidence interval can include the population mean. We draw sample ```simu_time``` times, each time calculate its confidence interval based on sampling distribution, if the population mean is included in confidence interval, we append a $1$ onto a list named ```conf_inclusion```, otherwise append a $0$.
#
# It shows that round $95\%$ of times the confidence interval will indeed include population mean.
conf_inclusion =[]
simu_time = 10000
for i in range(simu_time):
old_ppl.sampling(100)
conf_interval = old_ppl.conf_interval(.95, two_tail=True)
if conf_interval[0] <= population_mean and population_mean <= conf_interval[1]:
conf_inclusion.append(1)
else:
conf_inclusion.append(0)
print('Total number of simulation: {}'.format(simu_time))
print('Number of inclusion: {}'.format(np.sum(conf_inclusion)))
print('Inclusion rate: {}'.format(np.sum(conf_inclusion)/len(conf_inclusion)))
# # <font face="gotham" color="purple"> Interval Estimation With Unknown $\sigma$
# In practice, we barely have the knowledge of the $\sigma$, then $s$ is used instead. Because of unknown $\sigma$, the sampling distribution of mean will be based on $t$-distribution. A $t$-distribution with large degree of freedom is very close to a normal distribution, while smaller degree of freedom will cause fatter tails in $t$-distribution than normal distribution.
#
# The notation of interval estimator based on $t$-distribution is similar to the one based on the normal distribution
# $$
# P\bigg(\bar{x}-t_{\alpha/2}\frac{s}{\sqrt{n}}<\mu<\bar{x}+t_{\alpha/2}\frac{s}{\sqrt{n}}\bigg)
# $$
# where $s$ is the sample standard deviation
# $$
# s = \sqrt{\frac{\sum(x_i - \bar{x})^2}{n-1}}
# $$
# We can compare the confidence based on normal and $t$-distribution, very similar.
population_mean = 7.5
old_ppl = OldPeopleSleep(population_mean, std=2, pop_size=100000)
old_ppl.sampling(10)
print(old_ppl.conf_interval(.95, two_tail=True))
print(old_ppl.conf_interval_t(.95, two_tail=True))
conf_inclusion_t =[]
simu_time = 10000
for i in range(simu_time):
old_ppl.sampling(100)
conf_interval_t = old_ppl.conf_interval_t(.95, two_tail=True)
if conf_interval_t[0] <= population_mean and population_mean <= conf_interval_t[1]:
conf_inclusion_t.append(1)
else:
conf_inclusion_t.append(0)
print('Total number of simulation: {}'.format(simu_time))
print('Number of inclusion: {}'.format(np.sum(conf_inclusion)))
print('Inclusion rate: {}'.format(np.sum(conf_inclusion)/len(conf_inclusion)))
|
Chapter 3 - Point and Interval Estimation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:dspy3]
# language: python
# name: conda-env-dspy3-py
# ---
# # Scraping Google Images
#
# simple_image_download, imported here as "simp" will scrape the first N (no_images) images displayed for a search term.
# Here the search term is looped over from a list of house plants in a .csv
from simple_image_download import simple_image_download as simp
import pandas as pd
import os
# **Initial Setup**
# +
no_images = 2000 # number of images to download for each search
# load plants list
plants_df = pd.read_csv('../data/house_plants.csv',
names=['name', 'type'], skiprows=1,)
# initialize downloader
response = simp.simple_image_download
# -
# **Download all plants**
# download houseplants
for plant in plants_df.name.values:
response().download(f'{plant} houseplant', no_images)
# **Tidy directory**
# +
# create google images directory (this will overwrite)
os.makedirs('../google_images/', exist_ok=True)
# rename to remove spaces
for plant in plants_df.name.values:
os.system(
f'mv -r ../simple_images/{plant}_plant/ ../google_images/{plant}')
|
notebooks/01_scrape_google_images.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # LAB 3b: BigQuery ML Model Linear Feature Engineering/Transform.
#
# **Learning Objectives**
#
# 1. Create and evaluate linear model with BigQuery's ML.FEATURE_CROSS
# 1. Create and evaluate linear model with BigQuery's ML.FEATURE_CROSS and ML.BUCKETIZE
# 1. Create and evaluate linear model with ML.TRANSFORM
#
#
# ## Introduction
# In this notebook, we will create multiple linear models to predict the weight of a baby before it is born, using increasing levels of feature engineering using BigQuery ML. If you need a refresher, you can go back and look how we made a baseline model in the previous notebook [BQML Baseline Model](../solutions/3a_bqml_baseline_babyweight.ipynb).
#
# We will create and evaluate a linear model using BigQuery's ML.FEATURE_CROSS, create and evaluate a linear model using BigQuery's ML.FEATURE_CROSS and ML.BUCKETIZE, and create and evaluate a linear model using BigQuery's ML.TRANSFORM.
#
# Each learning objective will correspond to a __#TODO__ in this student lab notebook -- try to complete this notebook first and then review the [solution notebook](../solutions/3b_bqml_linear_transform_babyweight.ipynb).
# + [markdown] colab_type="text" id="hJ7ByvoXzpVI"
# ## Load necessary libraries
# + [markdown] colab_type="text" id="mC9K9Dpx1ztf"
# Check that the Google BigQuery library is installed and if not, install it.
# + colab={"base_uri": "https://localhost:8080/", "height": 609} colab_type="code" id="RZUQtASG10xO" outputId="5612d6b0-9730-476a-a28f-8fdc14f4ecde" language="bash"
# sudo pip freeze | grep google-cloud-bigquery==1.6.1 || \
# sudo pip install google-cloud-bigquery==1.6.1
# + [markdown] colab_type="text" id="clnaaqQsXkwC"
# ## Verify tables exist
#
# Run the following cells to verify that we previously created the dataset and data tables. If not, go back to lab [1b_prepare_data_babyweight](../solutions/1b_prepare_data_babyweight.ipynb) to create them.
# -
# %%bigquery
-- LIMIT 0 is a free query; this allows us to check that the table exists.
SELECT * FROM babyweight.babyweight_data_train
LIMIT 0
# %%bigquery
-- LIMIT 0 is a free query; this allows us to check that the table exists.
SELECT * FROM babyweight.babyweight_data_eval
LIMIT 0
# + [markdown] colab_type="text" id="FbSRbuJ-fYtK"
# ## Lab Task #1: Model 1: Apply the ML.FEATURE_CROSS clause to categorical features
#
# BigQuery ML now has ML.FEATURE_CROSS, a pre-processing clause that performs a feature cross with syntax ML.FEATURE_CROSS(STRUCT(features), degree) where features are comma-separated categorical columns and degree is highest degree of all combinations.
# -
# #### Create model with feature cross.
# + cellView="both" colab={} colab_type="code" id="Z3U2FxVklrlU"
# %%bigquery
CREATE OR REPLACE MODEL
babyweight.model_1
OPTIONS (
MODEL_TYPE="LINEAR_REG",
INPUT_LABEL_COLS=["weight_pounds"],
L2_REG=0.1,
DATA_SPLIT_METHOD="NO_SPLIT") AS
SELECT
# TODO: Add base features and label
ML.FEATURE_CROSS(
# TODO: Cross categorical features
) AS gender_plurality_cross
FROM
babyweight.babyweight_data_train
# + [markdown] colab_type="text" id="G6tpoYhcIgs4"
# #### Create two SQL statements to evaluate the model.
# + cellView="both" colab={} colab_type="code" id="t10fGqSfIgtA"
# %%bigquery
SELECT
*
FROM
ML.EVALUATE(MODEL babyweight.model_1,
(
SELECT
# TODO: Add same features and label as training
FROM
babyweight.babyweight_data_eval
))
# + cellView="both" colab={} colab_type="code" id="uC-gyvAmIgtE"
# %%bigquery
SELECT
# TODO: Select just the calculated RMSE
FROM
ML.EVALUATE(MODEL babyweight.model_1,
(
SELECT
# TODO: Add same features and label as training
FROM
babyweight.babyweight_data_eval
))
# -
# ## Lab Task #2: Model 2: Apply the BUCKETIZE Function
#
# Bucketize is a pre-processing function that creates "buckets" (e.g bins) - e.g. it bucketizes a continuous numerical feature into a string feature with bucket names as the value with syntax ML.BUCKETIZE(feature, split_points) with split_points being an array of numerical points to determine bucket bounds.
# #### Apply the BUCKETIZE function within FEATURE_CROSS.
# * Hint: Create a model_2.
# +
# %%bigquery
CREATE OR REPLACE MODEL
babyweight.model_2
OPTIONS (
MODEL_TYPE="LINEAR_REG",
INPUT_LABEL_COLS=["weight_pounds"],
L2_REG=0.1,
DATA_SPLIT_METHOD="NO_SPLIT") AS
SELECT
weight_pounds,
is_male,
mother_age,
plurality,
gestation_weeks,
ML.FEATURE_CROSS(
STRUCT(
is_male,
ML.BUCKETIZE(
# TODO: Bucketize mother_age
) AS bucketed_mothers_age,
plurality,
ML.BUCKETIZE(
# TODO: Bucketize gestation_weeks
) AS bucketed_gestation_weeks
)
) AS crossed
FROM
babyweight.babyweight_data_train
# + [markdown] colab_type="text" id="AVPXGKZ374v7"
# #### Create three SQL statements to EVALUATE the model.
#
# Let's now retrieve the training statistics and evaluate the model.
# -
# %%bigquery
SELECT * FROM ML.TRAINING_INFO(MODEL babyweight.model_2)
# We now evaluate our model on our eval dataset:
# %%bigquery
SELECT
*
FROM
ML.EVALUATE(MODEL babyweight.model_2,
(
SELECT
# TODO: Add same features and label as training
FROM
babyweight.babyweight_data_eval))
# Let's select the `mean_squared_error` from the evaluation table we just computed and square it to obtain the rmse.
# %%bigquery
SELECT
SQRT(mean_squared_error) AS rmse
FROM
ML.EVALUATE(MODEL babyweight.model_2,
(
SELECT
# TODO: Add same features and label as training
FROM
babyweight.babyweight_data_eval))
# ## Lab Task #3: Model 3: Apply the TRANSFORM clause
#
# Before we perform our prediction, we should encapsulate the entire feature set in a TRANSFORM clause. This way we can have the same transformations applied for training and prediction without modifying the queries.
# Let's apply the TRANSFORM clause to the model_3 and run the query.
# +
# %%bigquery
CREATE OR REPLACE MODEL
babyweight.model_3
TRANSFORM(
# TODO: Add base features and label as you would in select
# TODO: Add transformed features as you would in select
)
OPTIONS (
MODEL_TYPE="LINEAR_REG",
INPUT_LABEL_COLS=["weight_pounds"],
L2_REG=0.1,
DATA_SPLIT_METHOD="NO_SPLIT") AS
SELECT
*
FROM
babyweight.babyweight_data_train
# -
# Let's retrieve the training statistics:
# %%bigquery
SELECT * FROM ML.TRAINING_INFO(MODEL babyweight.model_3)
# We now evaluate our model on our eval dataset:
# %%bigquery
SELECT
*
FROM
ML.EVALUATE(MODEL babyweight.model_3,
(
SELECT
*
FROM
babyweight.babyweight_data_eval
))
# Let's select the `mean_squared_error` from the evaluation table we just computed and square it to obtain the rmse.
# %%bigquery
SELECT
SQRT(mean_squared_error) AS rmse
FROM
ML.EVALUATE(MODEL babyweight.model_3,
(
SELECT
*
FROM
babyweight.babyweight_data_eval
))
# ## Lab Summary:
# In this lab, we created and evaluated a linear model using BigQuery's ML.FEATURE_CROSS, created and evaluated a linear model using BigQuery's ML.FEATURE_CROSS and ML.BUCKETIZE, and created and evaluated a linear model using BigQuery's ML.TRANSFORM and L2 regularization.
# Copyright 2019 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License
|
courses/machine_learning/deepdive2/structured/labs/3b_bqml_linear_transform_babyweight.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="vnRzMA4KfJy5"
# + [markdown] id="PsOpdI2tLtUM"
# ## STATSMODEL
# + id="_3eDcPbxLtUM" colab={"base_uri": "https://localhost:8080/"} outputId="d9a91eb5-2c68-4fcf-80ee-f8f443d89769"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
import statsmodels.api as sm
# + id="MU9nqY8hLtUN" colab={"base_uri": "https://localhost:8080/", "height": 419} outputId="f60f2ffb-b29e-46d1-c2fc-e899d7bc6433"
df=sm.datasets.macrodata.load_pandas().data #in order to load data from statsmodels
df
# + id="0omPirRoLtUO" colab={"base_uri": "https://localhost:8080/"} outputId="d324e83a-8115-4651-a503-28c66c67e8a6"
#if you want information from statsmodels dataset then:
print(sm.datasets.macrodata.NOTE)
# + id="JrZSFh_ELtUO"
# + id="d5fNJtr8LtUO"
#we can convert years data to date by using pandas
#BUT statsmodels also has the capability to do that:
df.index = pd.Index(sm.tsa.datetools.dates_from_range('1959Q1','2009Q3'))
# + id="Sd1yJi2XLtUO" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="a4b6db94-0eda-46e1-8820-effd22838193"
df.head()
# + id="s44cTL77LtUO" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="ffcdde39-3d55-4fd6-82c6-5f76efe08465"
df['realgdp'].plot()
# + id="V0QDmclxLtUP"
#we are gonna use stats model to get the trend.
result=sm.tsa.filters.hpfilter(df['realgdp'])
# + id="MhJDHxmBLtUP" colab={"base_uri": "https://localhost:8080/"} outputId="7da0dc8c-8cef-445a-dd3a-b4f6925df8d1"
type(result)
# + id="cYFLmW6ILtUP"
#this hpfilter gives a tuple. 1- is the cycle and 2- is the trend
#in order to catch the trend line we can store that tuple into two diff vars
gdp_cycle,gdp_trend=sm.tsa.filters.hpfilter(df['realgdp'])
# + id="quRhoiJhLtUP"
#adding a trend coloum to the df
df['trend'] = gdp_trend
df['Cycle'] = gdp_cycle
# + id="WlSB8prCLtUQ" colab={"base_uri": "https://localhost:8080/", "height": 283} outputId="bfc2e9e2-da5b-4036-92b2-e3808f1a7a0d"
#ploting the realgdp with the trend line in order to show the trend of the realgdp.
df[['realgdp','trend']]['2000-03-31':].plot()
|
Recognising_Trend_in_Time_Series_Data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
print('''创建数组''')
arr1 = np.array([2,3,4])
arr2 = np.array([(1.3,9,2.0),(7,6,1)])
arr3 = np.zeros([2,3])
arr4 = np.identity(3)
arr5 = np.random.random(size = (2,3))
arr6 = np.arange(5,20,3)
arr7 = np.linspace(0,2,9)
print(arr1)
print(arr2)
print(arr3)
print(arr4)
print(arr5)
print(arr6)
print(arr7)
print(arr2.shape)
print(arr2.ndim)
print(arr2.size)
print(arr2.dtype.name)
print(type(arr2))
def f(x,y):
return 10*x+y
arr8 = np.fromfunction(f,(4,3),dtype = int)
print(arr8)
print(arr8[1,2])
print(arr8[0:2,:])
print(arr8[:,1])
print(arr8[-1])
for row in arr8:
print(row)
for element in arr8.flat:
print(element)
print('''数组的运算''')
arr9 = np.array([[2,1],[1,2]])
arr10 = np.array([[1,2],[3,4]])
print(arr9 - arr10)
print(arr9**2)
print(3*arr10)
print(arr9*arr10)
print(np.dot(arr9,arr10))
print(arr10.T)
print(np.linalg.inv(arr10))
print('''Numpy通用函数''')
print(np.exp(arr9))
print(np.sin(arr9))
print(np.sqrt(arr9))
print(np.add(arr9,arr10))
arr11 = np.vstack((arr9,arr10))
print(arr11)
arr12 = np.hstack((arr9,arr10))
print(arr12)
print(np.hsplit(arr12,2))
print(np.vsplit(arr11,2))
import pandas as pd
data = {
'id':['Jack','Sarah','Mike'],
'age':[18,35,20],
'cash':[10.53,500.7,13.6]
}
df = pd.DataFrame(data)
print(df)
df2 = pd.DataFrame(data, columns = ['id','age','cash'], index = ['one','two','three'])
print(df2)
print(df2['id'])
s = pd.Series({'a':4,'b':9,'c':16},name='number')
print(s)
print(s[0])
print(s[:3])
print(s['a'])
s['d']=25
print(s)
import numpy as np
print(np.sqrt(s))
print(s*s)
print (df['id'])
df['rich'] = df['cash']>200.0
print(df)
del df['rich']
print(df)
# +
import numpy as np
import matplotlib.pyplot as plt
x = np.linspace(0,10,100)
y = np.sin(x)
z = np.cos(x)
plt.figure(figsize=(8,4))
plt.plot(x,y,label="$sin(x)$",color='red',linewidth=2)
plt.plot(x,z,"b--",label="$cos(x^2)$")
plt.xlabel("Time(s)")
plt.ylabel("Volt")
plt.title("Pyplot First Example")
plt.ylim(-1.2,1.2)
plt.legend()
plt.show()
# -
import matplotlib.pylab as plt
import numpy as np
plt.subplot(2,1,1)
plt.subplot(2,2,3)
plt.subplot(2,2,4)
plt.show()
# +
import matplotlib.pylab as plt
import numpy as np
plt.subplot(2,1,1)
n = 12
X = np.arange(n)
Y1 = (1-X/float(n)) * np.random.uniform(0.5,1.0,n)
Y2 = (1-X/float(n)) * np.random.uniform(0.5,1.0,n)
plt.bar(X, +Y1, facecolor = '#9999ff', edgecolor='white')
plt.bar(X, -Y2, facecolor = '#ff9999', edgecolor='white')
for x,y in zip(X,Y1):
plt.text(x+0.4, y+0.05, '%.2f' % y, ha='center',va='bottom')
plt.ylim(-1.25, +1.25)
plt.subplot(2,2,3)
n = 20
z = np.random.uniform(0,1,n)
plt.pie(z)
plt.subplot(2,2,4)
x = np.linspace(-np.pi,np.pi,256,endpoint=True)
Y_C, Y_S = np.cos(x),np.sin(x)
plt.plot(x,Y_C,color='blue',linewidth=2.5,linestyle="-")
plt.plot(x,Y_S,color='red',linewidth=2.5,linestyle="-")
plt.xlim(x.min()*1.1,x.max()*1.1)
plt.xticks([-np.pi,-np.pi/2,0,np.pi/2,np.pi],
[r'$-\pi$',r'$-\pi/2$',r'$0$',r'$+\pi/2$',r'$+\pi$'])
plt.ylim(Y_C.min()*1.1,Y_C.max()*1.1)
plt.yticks([-1,0,+1],
[r'$-1$',r'$0$',r'$+1$'])
plt.show()
# -
from bokeh.plotting import figure, output_file,show
x = [1,2,3,4,5]
y = [6,7,2,4,5]
output_file("lines.html",title="line plot example")
p = figure(title = "simple line example", x_axis_label="x",y_axis_label="y")
p.line(x,y,legend="Line A.",line_width=2)
show(p)
# +
import numpy as np
import matplotlib.pyplot as plt
from sklearn.datasets import load_boston
from sklearn.linear_model import LinearRegression
boston = load_boston()
#print(boston.keys())
#print(boston.feature_names)
x = boston.data[:,np.newaxis, 5]
y = boston.target
lm = LinearRegression()
lm.fit(x,y)
print(u'方程的确定性系数(R^2): %.2f' % lm.score(x,y))
plt.scatter(x,y,color='green')
plt.plot(x,lm.predict(x),color='blue',linewidth=3)
plt.xlabel('Average Number of Rooms per Dwellings(RM)')
plt.ylabel('Hosing Price')
plt.title('2D Demo of Linear Regression')
plt.show()
# -
|
Practice.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=true editable=true
# This is an implementation of the *harder* option for Assignment 3 of coursera's [Applied Plotting, Charting & Data Representation in Python](https://www.coursera.org/learn/python-plotting/home/welcome).
#
#
# # Description
#
# A challenge that users face is that, for a given y-axis value (e.g.
# 42,000), it is difficult to know which x-axis values are most likely to
# be representative, because the confidence levels overlap and their
# distributions are different (the lengths of the confidence interval bars
# are unequal). One of the solutions the authors propose for this problem
# (Figure 2c) is to allow users to indicate the y-axis value of interest
# (e.g. 42,000) and then draw a horizontal line and color bars based on
# this value. So bars might be colored red if they are definitely above
# this value (given the confidence interval), blue if they are definitely
# below this value, or white if they contain this value.
#
# *Even Harder option:* Add interactivity to the above, which allows the
# user to click on the y axis to set the value of interest. The bar colors
# should change with respect to what value the user has selected.
#
# # Imports
#
# All the imports were created by third-parties (taken from pypi).
#
#
# + deletable=true editable=true
import matplotlib.pyplot as pyplot
import numpy
import pandas
import scipy.stats as stats
import seaborn
# + [markdown] deletable=true editable=true
#
# # Some Plotting Setup
#
#
# + deletable=true editable=true
# %matplotlib notebook
style = seaborn.axes_style("whitegrid")
style["axes.grid"] = False
seaborn.set_style("whitegrid", style)
# + [markdown] deletable=true editable=true
#
# # The Data
#
# There data set will be four normally-distributed, randomly generated data sets each representing a simulated data set for a given year.
#
#
# ## `numpy.random.normal`
#
# This is from the `numpy.random.normal` doc-string:
#
# `normal(loc=0.0, scale=1.0, size=None)`
#
# Draw random samples from a normal (Gaussian) distribution.
#
# The probability density function of the normal distribution, first
# derived by <NAME>re and 200 years later by both Gauss and Laplace
# independently [2]\_, is often called the bell curve because of
# its characteristic shape (see the example below).
#
# The normal distributions occurs often in nature. For example, it
# describes the commonly occurring distribution of samples influenced
# by a large number of tiny, random disturbances, each with its own
# unique distribution.
#
#
# ## Parameters
#
#
# ### `loc` : float or array\_like of floats
#
# Mean ("centre") of the distribution.
#
#
# ### `scale` : float or array\_like of floats
#
# Standard deviation (spread or "width") of the distribution.
#
#
# ### `size` : int or tuple of ints, optional
#
# Output shape. If the given shape is, e.g., `(m, n, k)`, then
# `m * n * k` samples are drawn. If size is `None` (default),
# a single value is returned if `loc` and `scale` are both scalars.
# Otherwise, `np.broadcast(loc, scale).size` samples are drawn.
#
#
# + deletable=true editable=true
numpy.random.seed(12345)
data = pandas.DataFrame([numpy.random.normal(33500,150000,3650),
numpy.random.normal(41000,90000,3650),
numpy.random.normal(41000,120000,3650),
numpy.random.normal(48000,55000,3650)],
index=[1992,1993,1994,1995])
# + [markdown] deletable=true editable=true
#
#
# + deletable=true editable=true
data.T.describe()
# + [markdown] deletable=true editable=true
# Comparing the sample to the values fed to the `normal` function it appears that even with 3,650 values, it's still not exactly what we asked for.
# + deletable=true editable=true
data.T.plot.kde()
seaborn.despine()
# + [markdown] deletable=true editable=true
# 1992, the plot with the largest spread looks kind of lumpy. Their means look surprisingly close, but that's probably because the large standaard deviation distorts the scale.
#
#
# + deletable=true editable=true
data.T.plot.box()
seaborn.despine()
# + [markdown] deletable=true editable=true
# The box-plot shows once again that there centers are relatively close. But 1992 and 1994 have considerably more spread than 1993 and especially more than 1995.
#
#
# # Interval Check
#
# This is the class that implements the plotting. It colors the bar-plots based on whether the value given is within a bar's confidence interval (white), below the confidence interval (blue) or above the confidence interval (red). It's set up to work with the easiest case so the `color_bars` method has to be overridden to make it work for this case.
#
#
# + deletable=true editable=true
class IntervalCheck(object):
"""colors plot based on whether a value is in range
Args:
data (DataFrame): frame with data of interest as columns
confidence_interval (float): probability we want to exceed
"""
def __init__(self, data, confidence_interval=0.95, title="Confidence Intervals"):
self.data = data
self.confidence_interval = confidence_interval
self.title = title
self._intervals = None
self._lows = None
self._highs = None
self._errors = None
self._means = None
self._errors = None
self._figure = None
self._axes = None
self._bars = None
self.horizontal_line = None
self.line_label = None
return
@property
def figure(self):
if self._figure is None:
"""A pyplot figure"""
self._figure = pyplot.figure()
return self._figure
@property
def axes(self):
if self._axes is None:
"""the current axes for self.figure"""
self._axes = self.figure.gca()
self._axes.set_title(self.title)
return self._axes
@property
def bars(self):
"""the bar-plot-objects"""
if self._bars is None:
self._bars = self.axes.bar(self.data.columns, self.means,
yerr=self.errors)
return self._bars
@property
def intervals(self):
"""list of high and low interval tuples"""
if self._intervals is None:
data = (self.data[column] for column in self.data)
self._intervals = [stats.norm.interval(alpha=self.confidence_interval,
loc=datum.mean(),
scale=datum.sem())
for datum in data]
return self._intervals
@property
def lows(self):
"""the low-ends for the confidence intervals
Returns:
numpy.array of low-end confidence interval values
"""
if self._lows is None:
self._lows = numpy.array([low for low, high in self.intervals])
return self._lows
@property
def highs(self):
"""high-ends for the confidence intervals
Returns:
numpy.array of high-end values for confidence intervals
"""
if self._highs is None:
self._highs = numpy.array([high for low, high in self.intervals])
return self._highs
@property
def means(self):
"""the means of the data-arrays"""
if self._means is None:
self._means = self.data.mean()
return self._means
@property
def errors(self):
"""The size of the errors, rather than the ci values"""
if self._errors is None:
self._errors = self.highs - self.means
return self._errors
def print_intervals(self):
"""print org-mode formatted table of the confidence intervals"""
intervals = pandas.DataFrame({column: self.intervals[index]
for index, column in enumerate(self.data.columns)},
index="low high".split())
try:
print(tabulate(intervals, tablefmt="orgtbl", headers="keys"))
except NameError:
# not supported
pass
return
def draw_value(self, value):
"""draws the horizontal line and value"""
if self.horizontal_line:
self.horizontal_line.set_ydata(value)
self.line_label.set_y(value)
self.line_label.set_text("{0:.2f}".format(value))
else:
self.horizontal_line = pyplot.axhline(value,
axes=self.axes,
color="darkorange")
self.line_label = pyplot.text(self.data.columns[0],
value,
"{0:.2f}".format(value),
axes=self.axes,
bbox={"facecolor": "white", "boxstyle": "round"})
return
def setup_bars(self, value):
"""sets up the horizontal line, value and bars
Args:
value (float): value to compare to distributions
"""
x_labels = [str(index) for index in self.data.columns]
for bar in self.bars:
bar.set_edgecolor("royalblue")
pyplot.xticks(self.data.columns, x_labels)
self.draw_value(value)
return
def color_bars(self, value):
"""colors the bars based on the value
this is the easiest case
Args:
value (float): value to compare to the distribution
"""
for index, bar in enumerate(self.bars):
if value < self.lows[index]:
bar.set_color('crimson')
elif self.lows[index] <= value <= self.highs[index]:
bar.set_color('w')
bar.set_edgecolor("royalblue")
else:
bar.set_color("royalblue")
return
def __call__(self, value):
"""plots the data and value
* blue bar if value above c.i.
* white bar if value in c.i.
* red bar if value is below c.i.
Args:
value (float): what to compare to the data
"""
self.setup_bars(value)
self.color_bars(value)
return
# + [markdown] deletable=true editable=true
#
# # Harder
#
# This is the class that implements the harder coloring scheme were a gradient is used instead of just three colors.
# + deletable=true editable=true
class Harder(IntervalCheck):
"""implements the harder problem
Uses a gradient instead of just 3 colors
"""
def __init__(self, *args, **kwargs):
super(Harder, self).__init__(*args, **kwargs)
self._colors = None
self._proportions = None
return
@property
def colors(self):
"""array of rgb color triples"""
if self._colors is None:
# could have been done with straight fractions
# but I find it easier to think in terms of
# 0..255
base = list(range(0, 255, 51))
full = [255] * 6
blue = numpy.array(base + full)
blue = blue/255
base.reverse()
red = numpy.array(full + base)
red = red/255
tail = base[:]
base.reverse()
green = numpy.array(base + [255] + tail)/255
self._colors = numpy.array([red, green, blue]).T
return self._colors
@property
def proportions(self):
"""array of upper limits for the value to find the matching color
"""
if self._proportions is None:
self._proportions = numpy.linspace(0.09, 1, 10)
return self._proportions
def color_bars(self, value):
"""colors the bars based on the value
this is the harder case
Args:
value (float): value to compare to the distribution
"""
mapped_values = [(value - low)/(high - low)
for low, high in self.intervals]
for index, mapped_value in enumerate(mapped_values):
if mapped_value < 0:
self.bars[index].set_color(self.colors[0])
continue
if mapped_value >= 1:
self.bars[index].set_color(self.colors[-1])
continue
for p_index, proportion in enumerate(self.proportions):
if mapped_value <= proportion:
color = self.colors[p_index]
self.bars[index].set_color(color)
self.bars[index].set_edgecolor("royalblue")
break
return
# + [markdown] deletable=true editable=true
# # Even Harder
#
# This is the class that adds interactivity to the Harder case.
# + deletable=true editable=true
class EvenHarder(Harder):
"""the interactive version of Harder"""
@property
def figure(self):
"""pyplot figure
As a side-effect registers on_click with the canvas
"""
if self._figure is None:
self._figure = pyplot.figure()
self._figure.canvas.mpl_connect("button_press_event",
self.on_click)
return self._figure
def on_click(self, event):
"""event-handler to update the plot"""
if event.ydata:
self.draw_value(event.ydata)
self.color_bars(event.ydata)
return
def __call__(self, value=0):
"""add a default value since this is interactive"""
super(EvenHarder, self).__call__(value)
return
# + [markdown] deletable=true editable=true
#
# # Examples
#
# First, I'll take a look at the values for the confidence intervals so that I can find values to plot. Here are the confidence intervals for the data I created.
#
#
# + deletable=true editable=true
plotter = EvenHarder(data=data.T)
plotter.print_intervals()
# + [markdown] deletable=true editable=true
# Here's a value that is below all the confidence intervals.
#
#
# + deletable=true editable=true
value = 42000
plotter(value)
# + [markdown] deletable=true editable=true
#
#
|
2_visualizing_data/week_3/assignment_3_harder.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Practice
# ### Write a program that goes over(iterates) numbers from 1 to 100(included)
# * For each number it prints Fizz if number divides by 5 even
# * For each number it prints Buzz if number divides by 7 evenly
# * If number divides by 5 AND 7 print FizzBuzz
# * print number if it does not divide by 5 nor 7
# * 1,2,3,4,Fizz,6,Buzz,8,9,Fizz,11,.......34,FizzBuzz,36
for n in range(20):
if n % 2 == 0:
print(n,"is even!")
else:
print(n, "is odd!")
for c in range(1, 100+1):
end = ", " # instead of default newline for print
if c%5 == 0 and c%7 == 0:
print("FizzBuzz", end=end)
elif c%5 == 0:
print("FiZZ", end=end)
elif c%7 == 0:
print("Buzz", end=end)
else:
print(c, end=end)
# +
txt = "" # this has to go outside otherwise we will keep rewriting the txt
for c in range(1,101):
# print("working on ", c, "txt is", txt)
if c%5 == 0:
txt += "Fizz"
if c%7 == 0:
txt += "Buzz"
if c%5 != 0 and c%7 !=0: # this means number
txt += str(c) # c is number we need to convert to string
if c < 100:
txt += ", "
print("My fizzbuzz")
print(txt)
# -
|
core/FizzBuzz.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import time
import numpy as np
import h5py
import matplotlib.pyplot as plt
import scipy
from PIL import Image
from scipy import ndimage
from dnn_app_utils_v2 import *
# %matplotlib inline
plt.rcParams['figure.figsize'] = (5.0, 4.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
# %load_ext autoreload
# %autoreload 2
np.random.seed(1)
# -
train_x_orig, train_y, test_x_orig, test_y, classes = load_data()
# Example of a picture
index = 10
plt.imshow(train_x_orig[index])
print ("y = " + str(train_y[0,index]) + ". It's a " + classes[train_y[0,index]].decode("utf-8") + " picture.")
# +
# Explore your dataset
m_train = train_x_orig.shape[0]
num_px = train_x_orig.shape[1]
m_test = test_x_orig.shape[0]
print ("Number of training examples: " + str(m_train))
print ("Number of testing examples: " + str(m_test))
print ("Each image is of size: (" + str(num_px) + ", " + str(num_px) + ", 3)")
print ("train_x_orig shape: " + str(train_x_orig.shape))
print ("train_y shape: " + str(train_y.shape))
print ("test_x_orig shape: " + str(test_x_orig.shape))
print ("test_y shape: " + str(test_y.shape))
# -
#
# <img src="https://zedwnutwhnkzdykkpsnyql.coursera-apps.org/notebooks/Week%204/Deep%20Neural%20Network%20Application:%20Image%20Classification/images/imvectorkiank.png" style="width:450px;height:300px;">
#
# <caption><center> <u>Figure 1</u>: Image to vector conversion. <br> </center></caption>
#
# +
# Reshape the training and test examples
train_x_flatten = train_x_orig.reshape(train_x_orig.shape[0], -1).T # The "-1" makes reshape flatten the remaining dimensions
test_x_flatten = test_x_orig.reshape(test_x_orig.shape[0], -1).T
# Standardize data to have feature values between 0 and 1.
train_x = train_x_flatten/255.
test_x = test_x_flatten/255.
print ("train_x's shape: " + str(train_x.shape))
print ("test_x's shape: " + str(test_x.shape))
# -
# ## 搭建两层神经网络
# 一个两层的神经网络模型图如下:
#
# <img src="https://zedwnutwhnkzdykkpsnyql.coursera-apps.org/notebooks/Week%204/Deep%20Neural%20Network%20Application:%20Image%20Classification/images/LlayerNN_kiank.png" style="width:650px;height:400px;">
# <caption><center> <u>Figure 3</u>: L-layer neural network. <br> 该模型可以概括为: ***[LINEAR -> RELU] $\times$ (L-1) -> LINEAR -> SIGMOID***</center></caption>
#
# 我们正式开始构建两层的神经网络:
#
def two_layer_model(X,Y,layers_dims,learning_rate=0.0075,num_iterations=3000,print_cost=False,isPlot=True):
"""
实现一个两层的神经网络,【LINEAR->RELU】 -> 【LINEAR->SIGMOID】
参数:
X - 输入的数据,维度为(n_x,例子数)
Y - 标签,向量,0为非猫,1为猫,维度为(1,数量)
layers_dims - 层数的向量,维度为(n_y,n_h,n_y)
learning_rate - 学习率
num_iterations - 迭代的次数
print_cost - 是否打印成本值,每100次打印一次
isPlot - 是否绘制出误差值的图谱
返回:
parameters - 一个包含W1,b1,W2,b2的字典变量
"""
np.random.seed(1)
grads = {}
costs = []
(n_x,n_h,n_y) = layers_dims
"""
初始化参数
"""
parameters = initialize_parameters(n_x, n_h, n_y)
W1 = parameters["W1"]
b1 = parameters["b1"]
W2 = parameters["W2"]
b2 = parameters["b2"]
"""
开始进行迭代
"""
for i in range(0,num_iterations):
#前向传播
A1, cache1 = linear_activation_forward(X, W1, b1, "relu")
A2, cache2 = linear_activation_forward(A1, W2, b2, "sigmoid")
#计算成本
cost = compute_cost(A2,Y)
#后向传播
##初始化后向传播
dA2 = - (np.divide(Y, A2) - np.divide(1 - Y, 1 - A2))
##向后传播,输入:“dA2,cache2,cache1”。 输出:“dA1,dW2,db2;还有dA0(未使用),dW1,db1”。
dA1, dW2, db2 = linear_activation_backward(dA2, cache2, "sigmoid")
dA0, dW1, db1 = linear_activation_backward(dA1, cache1, "relu")
##向后传播完成后的数据保存到grads
grads["dW1"] = dW1
grads["db1"] = db1
grads["dW2"] = dW2
grads["db2"] = db2
#更新参数
parameters = update_parameters(parameters,grads,learning_rate)
W1 = parameters["W1"]
b1 = parameters["b1"]
W2 = parameters["W2"]
b2 = parameters["b2"]
#打印成本值,如果print_cost=False则忽略
if i % 100 == 0:
#记录成本
costs.append(cost)
#是否打印成本值
if print_cost:
print("第", i ,"次迭代,成本值为:" ,np.squeeze(cost))
#迭代完成,根据条件绘制图
if isPlot:
plt.plot(np.squeeze(costs))
plt.ylabel('cost')
plt.xlabel('iterations (per tens)')
plt.title("Learning rate =" + str(learning_rate))
plt.show()
#返回parameters
return parameters
# 我们现在开始加载数据集
# +
import lr_utils
train_set_x_orig , train_set_y , test_set_x_orig , test_set_y , classes = lr_utils.load_dataset()
train_x_flatten = train_set_x_orig.reshape(train_set_x_orig.shape[0], -1).T
test_x_flatten = test_set_x_orig.reshape(test_set_x_orig.shape[0], -1).T
train_x = train_x_flatten / 255
train_y = train_set_y
test_x = test_x_flatten / 255
test_y = test_set_y
# -
# 数据集加载完成,开始正式训练:
# +
n_x = 12288
n_h = 7
n_y = 1
layers_dims = (n_x,n_h,n_y)
parameters = two_layer_model(train_x, train_set_y, layers_dims = (n_x, n_h, n_y), num_iterations = 2500, print_cost=True,isPlot=True)
# -
# 迭代完成之后我们就可以进行预测了,预测函数如下:
def predict(X, y, parameters):
"""
该函数用于预测L层神经网络的结果,当然也包含两层
参数:
X - 测试集
y - 标签
parameters - 训练模型的参数
返回:
p - 给定数据集X的预测
"""
m = X.shape[1]
n = len(parameters) // 2 # 神经网络的层数
p = np.zeros((1,m))
#根据参数前向传播
probas, caches = L_model_forward(X, parameters)
for i in range(0, probas.shape[1]):
if probas[0,i] > 0.5:
p[0,i] = 1
else:
p[0,i] = 0
print("准确度为: " + str(float(np.sum((p == y))/m)))
return p
# 预测函数构建好了我们就开始预测,查看训练集和测试集的准确性:
predictions_train = predict(train_x, train_y, parameters) #训练集
predictions_test = predict(test_x, test_y, parameters) #测试集
# ## 搭建多层神经网络
# 我们首先来看看多层的网络的结构吧~
# <img src="https://zedwnutwhnkzdykkpsnyql.coursera-apps.org/notebooks/Week%204/Deep%20Neural%20Network%20Application:%20Image%20Classification/images/LlayerNN_kiank.png" style="width:650px;height:400px;">
# <caption><center> <u>Figure 3</u>: L-layer neural network. <br> 该模型可以概括为: ***[LINEAR -> RELU] $\times$ (L-1) -> LINEAR -> SIGMOID***</center></caption>
#
# ```python
# def initialize_parameters_deep(layers_dims):
# ...
# return parameters
# def L_model_forward(X, parameters):
# ...
# return AL, caches
# def compute_cost(AL, Y):
# ...
# return cost
# def L_model_backward(AL, Y, caches):
# ...
# return grads
# def update_parameters(parameters, grads, learning_rate):
# ...
# return parameters
# ```
def L_layer_model(X, Y, layers_dims, learning_rate=0.0075, num_iterations=3000, print_cost=False,isPlot=True):
"""
实现一个L层神经网络:[LINEAR-> RELU] *(L-1) - > LINEAR-> SIGMOID。
参数:
X - 输入的数据,维度为(n_x,例子数)
Y - 标签,向量,0为非猫,1为猫,维度为(1,数量)
layers_dims - 层数的向量,维度为(n_y,n_h,···,n_h,n_y)
learning_rate - 学习率
num_iterations - 迭代的次数
print_cost - 是否打印成本值,每100次打印一次
isPlot - 是否绘制出误差值的图谱
返回:
parameters - 模型学习的参数。 然后他们可以用来预测。
"""
np.random.seed(1)
costs = []
parameters = initialize_parameters_deep(layers_dims)
for i in range(0,num_iterations):
AL , caches = L_model_forward(X,parameters)
cost = compute_cost(AL,Y)
grads = L_model_backward(AL,Y,caches)
parameters = update_parameters(parameters,grads,learning_rate)
#打印成本值,如果print_cost=False则忽略
if i % 100 == 0:
#记录成本
costs.append(cost)
#是否打印成本值
if print_cost:
print("第", i ,"次迭代,成本值为:" ,np.squeeze(cost))
#迭代完成,根据条件绘制图
if isPlot:
plt.plot(np.squeeze(costs))
plt.ylabel('cost')
plt.xlabel('iterations (per tens)')
plt.title("Learning rate =" + str(learning_rate))
plt.show()
return parameters
# +
train_set_x_orig , train_set_y , test_set_x_orig , test_set_y , classes = lr_utils.load_dataset()
train_x_flatten = train_set_x_orig.reshape(train_set_x_orig.shape[0], -1).T
test_x_flatten = test_set_x_orig.reshape(test_set_x_orig.shape[0], -1).T
train_x = train_x_flatten / 255
train_y = train_set_y
test_x = test_x_flatten / 255
test_y = test_set_y
# +
layers_dims = [12288, 20, 7, 5, 1] # 5-layer model
parameters = L_layer_model(train_x, train_y, layers_dims, num_iterations = 2500, print_cost = True,isPlot=True)
# +
# 训练完成,我们看一下预测:
pred_train = predict(train_x, train_y, parameters) #训练集
pred_test = predict(test_x, test_y, parameters) #测试集
# -
# ## 分析
# 我们可以看一看有哪些东西在L层模型中被错误地标记了,导致准确率没有提高。
# +
def print_mislabeled_images(classes, X, y, p):
"""
绘制预测和实际不同的图像。
X - 数据集
y - 实际的标签
p - 预测
"""
a = p + y
mislabeled_indices = np.asarray(np.where(a == 1))
plt.rcParams['figure.figsize'] = (40.0, 40.0) # set default size of plots
num_images = len(mislabeled_indices[0])
for i in range(num_images):
index = mislabeled_indices[1][i]
plt.subplot(2, num_images, i + 1)
plt.imshow(X[:,index].reshape(64,64,3), interpolation='nearest')
plt.axis('off')
plt.title("Prediction: " + classes[int(p[0,index])].decode("utf-8") + " \n Class: " + classes[y[0,index]].decode("utf-8"))
print_mislabeled_images(classes, test_x, test_y, pred_test)
# -
# 分析一下我们就可以得知原因了:
# 模型往往表现欠佳的几种类型的图像包括:
#
# * 猫身体在一个不同的位置
# * 猫出现在相似颜色的背景下
# * 不同的猫的颜色和品种
# * 相机角度
# * 图片的亮度
# * 比例变化(猫的图像非常大或很小)
# ## 【选做】
# 我们使用自己图片试试?
# 我们把一张图片放在一个特定位置,然后识别它。
# +
## START CODE HERE ##
my_image = "my_image.jpg" # change this to the name of your image file
my_label_y = [1] # the true class of your image (1 -> cat, 0 -> non-cat)
## END CODE HERE ##
fname = "images/" + my_image
image = np.array(ndimage.imread(fname, flatten=False))
my_image = scipy.misc.imresize(image, size=(num_px,num_px)).reshape((num_px*num_px*3,1))
my_predicted_image = predict(my_image, my_label_y, parameters)
plt.imshow(image)
print ("y = " + str(np.squeeze(my_predicted_image)) + ", your L-layer model predicts a \"" + classes[int(np.squeeze(my_predicted_image)),].decode("utf-8") + "\" picture.")
# -
|
01神经网络和深度学习/Code编程作业/deeplearning第1专题编程作业/deeplearning编程作业/week4/Deep Neural Network Application Image Classification/第四周-一步步搭建多层神经网络以及应用2-图像分类.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.10.1 64-bit
# language: python
# name: python3
# ---
# ## Instructions
#
# Write a program that works out whether if a given year is a leap year. A normal year has 365 days, leap years have 366, with an extra day in February.
#
# This is how you work out whether if a particular year is a leap year.
#
# on every year that is evenly divisible by 4
#
# **except** every year that is evenly divisible by 100
#
# **unless** the year is also evenly divisible by 400
# +
#input year
year = int(input("Which year do you want to check? "))
# +
#checks if it's a leap year
if year % 4 == 0:
if year % 100 == 0:
if year % 400 == 0:
print("Leap year.")
else:
print("Not leap year.")
else:
print("Leap year.")
else:
print("Not leap year.")
|
Day-3/3. Leap_Year.ipynb
|