code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Ch4.1 Simple Scatter Plots
# %matplotlib inline
import matplotlib.pyplot as plt
plt.style.use('seaborn-whitegrid')
import numpy as np
# ## Scatter Plots with ``plt.plot``
# +
x = np.linspace(0, 10, 30)
y = np.sin(x)
plt.plot(x, y, 'o', color='black');
# -
# specify options such as ``'-'``, ``'--'`` to control the line style
plt.plot(x, y, '-ok');
plt.scatter(x, y, color='black')
# Additional keyword arguments to ``plt.plot``
plt.plot(x, y, '-p', color='gray',
markersize=15, linewidth=4,
markerfacecolor='white',
markeredgecolor='gray',
markeredgewidth=2)
plt.ylim(-1.2, 1.2);
# ## Scatter Plots with ``plt.scatter``
plt.scatter(x, y, marker='o');
plt.scatter(x, y, marker='o', color='black');
# creating a random scatter plot with points of many colors and sizes
# +
rng = np.random.RandomState(0)
x = rng.randn(100)
y = rng.randn(100)
colors = rng.rand(100)
sizes = 1000 * rng.rand(100)
plt.scatter(x, y, c=colors, s=sizes, alpha=0.3,
cmap='viridis')
plt.colorbar(); # show color scale
|
III_DataEngineer_BDSE10/1905_Python/TeacherCode/datascience/Ch4.1_Simple_Scatter_Plots.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
import os
import phys
phys.Measurement.set_code_scale("m", 0.001)
import phys.newton
import phys.light
import numpy as np
import time
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
sim = phys.Simulation(bounds=np.array([1000, 1000, 1000]), cl_on=True, exit=lambda cond: len(cond.objects) == 0)
sim.add_objs(phys.light.generate_photons(1e5, bins=100, min=phys.light.E_from_wavelength(phys.Measurement(200e-9, "m**1")), max=phys.light.E_from_wavelength(phys.Measurement(700e-9, "m**1"))))
print("Done generating photons")
sim.add_step(0, phys.UpdateTimeStep(lambda s: phys.Measurement(0.00001, "s**1")))
sim.add_step(1, phys.newton.NewtonianKinematicsStep())
n = phys.Measurement(2.0e25, "m**-3")
A = phys.Measurement(5.1e-31, "m**2")
sim.add_step(2, phys.light.ScatterDeleteStep(n, A))
m1 = phys.light.ScatterMeasureStep(None, True, [phys.Measurement([1 / (n * A), np.nan, np.nan], "m**1"), phys.Measurement([9.80e4, np.nan, np.nan], "m**1")])
sim.add_step(3, m1)
sim.start()
while sim.running:
time.sleep(5)
print(sim.get_state())
# +
pos = 1 / (n * A)
pos2 = phys.Measurement(9.80e4, "m**1")
plt.plot(sim.ts, [x[1] for x in m1.data], label="n")
plt.ylabel("Photons")
plt.xlabel("Time (s)")
plt.title("Photon Count vs. Time (s)")
plt.show()
idx1 = int(((pos / sim.dt) / phys.light.c).__unscaled__())
idx2 = int((pos2 / sim.dt) / phys.light.c)
print(idx1, idx2)
print("Photons at plane 1: " + str(m1.data[idx1][2]))
print("Photons at plane 2: " + str(m1.data[idx2][2]))
expect = (1e5 / np.e)
print("Expected: " + str(expect))
print("Error: " + str((expect - m1.data[idx1][2]) / expect))
# -
|
examples/code_unit_scale_test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Quickstart
# ==========
#
# In this short tutorial we will guide you through a series of steps that
# will help you getting started using **SDV**.
#
# Model the dataset using SDV
# ---------------------------
#
# To model a multi table, relational dataset, we follow two steps. In the
# first step, we will load the data and configures the meta data. In the
# second step, we will use the SDV API to fit and save a hierarchical
# model. We will cover these two steps in this section using an example
# dataset.
#
# ### Load example data
#
# **SDV** comes with a toy dataset to play with, which can be loaded using
# the `sdv.load_demo` function:
# + attributes={"classes": [""], "id": ""}
from sdv import load_demo
metadata, tables = load_demo(metadata=True)
# -
# This will return two objects:
#
# 1. A `Metadata` object with all the information that **SDV** needs to
# know about the dataset.
# + attributes={"classes": [""], "id": ""}
metadata
# + attributes={"classes": [""], "id": ""}
metadata.visualize()
# -
# For more details about how to build the `Metadata` for your own dataset,
# please refer to the [Relational Metadata](relational_data/01_Relational_Metadata.ipynb)
# guide.
#
# 2. A dictionary containing three `pandas.DataFrames` with the tables
# described in the metadata object.
# + attributes={"classes": [""], "id": ""}
tables
# -
# ### Fit a model using the SDV API.
#
# First, we build a hierarchical statistical model of the data using
# **SDV**. For this we will create an instance of the `sdv.SDV` class and
# use its `fit` method.
#
# During this process, **SDV** will traverse across all the tables in your
# dataset following the primary key-foreign key relationships and learn
# the probability distributions of the values in the columns.
# + attributes={"classes": [""], "id": ""}
from sdv import SDV
sdv = SDV()
sdv.fit(metadata, tables)
# -
# Sample data from the fitted model
# ---------------------------------
#
# Once the modeling has finished you are ready to generate new synthetic
# data using the `sdv` instance that you have.
#
# For this, all you have to do is call the `sample_all` method from your
# instance passing the number of rows that you want to generate:
# + attributes={"classes": [""], "id": ""}
sampled = sdv.sample_all()
# -
# This will return a dictionary identical to the `tables` one that we
# passed to the SDV instance for learning, filled in with new synthetic
# data.
#
# <div class="alert alert-info">
#
# **Note**
#
# Only the parent tables of your dataset will have the specified number of
# rows, as the number of child rows that each row in the parent table has
# is also sampled following the original distribution of your dataset.
#
# </div>
# + attributes={"classes": [""], "id": ""}
sampled
# -
# Saving and Loading your model
# -----------------------------
#
# In some cases, you might want to save the fitted SDV instance to be able
# to generate synthetic data from it later or on a different system.
#
# In order to do so, you can save your fitted `SDV` instance for later
# usage using the `save` method of your instance.
# + attributes={"classes": [""], "id": ""}
sdv.save('sdv.pkl')
# -
# The generated `pkl` file will not include any of the original data in
# it, so it can be safely sent to where the synthetic data will be
# generated without any privacy concerns.
#
# Later on, in order to sample data from the fitted model, we will first
# need to load it from its `pkl` file.
# + attributes={"classes": [""], "id": ""}
sdv = SDV.load('sdv.pkl')
# -
# After loading the instance, we can sample synthetic data using its
# `sample_all` method like before.
# + attributes={"classes": [""], "id": ""}
sampled = sdv.sample_all(5)
sampled
|
tutorials/Quickstart.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Load data, train model
import numpy as np
import pandas as pd
from sklearn.metrics import log_loss
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
# +
# Load data
data = pd.read_csv('../data/brca_small.csv')
X = data.values[:, :-1]
Y = data.values[:, -1]
# Split data
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size=100, random_state=0)
X_train, X_val, Y_train, Y_val = train_test_split(X_train, Y_train, test_size=100, random_state=1)
# Normalize
mean = X_train.mean(axis=0)
std = X_train.std(axis=0)
X_train = (X_train - mean) / std
X_val = (X_val - mean) / std
X_test = (X_test - mean) / std
# -
def fit_model(x, y, x_val, y_val):
# Cross validate for C
C_list = np.arange(0.1, 1.0, 0.05)
best_loss = np.inf
best_C = None
for C in C_list:
# Fit model
model = LogisticRegression(C=C, penalty='l1', multi_class='multinomial',
solver='saga', max_iter=20000)
model.fit(x, y)
# Calculate loss
val_loss = log_loss(y_val, model.predict_proba(x_val))
# See if best
if val_loss < best_loss:
best_loss = val_loss
best_C = C
# Train model with all data
model = LogisticRegression(C=best_C, penalty='l1', multi_class='multinomial',
solver='saga', max_iter=10000)
model.fit(np.concatenate((x, x_val), axis=0),
np.concatenate((y, y_val), axis=0))
return model
# Train model
model = fit_model(X_train, Y_train, X_val, Y_val)
# # SHAP cooperative game
import matplotlib.pyplot as plt
from shapreg import removal, games, shapley, shapley_unbiased
# +
# Make model callable
model_lam = lambda x: model.predict_proba(x)
# Model extension
marginal_extension = removal.MarginalExtension(X_train, model_lam)
# -
# Set up game (single prediction)
instance = X_test[0]
game = games.PredictionGame(marginal_extension, instance)
# Calculate SHAP values
explanation = shapley.ShapleyRegression(game, batch_size=64)
# +
# Plot
fig, axarr = plt.subplots(2, 2, figsize=(12, 8))
label_names = ['Basal', 'Her2', 'LumA', 'LumB']
max_val = explanation.values.max()
min_val = explanation.values.min()
for i, name in enumerate(label_names):
# Select axis and SHAP values
ax = axarr[i // 2, i % 2]
plt.sca(ax)
values = explanation.values[:, i]
# Plot
plt.bar(np.arange(X.shape[1]), values)
plt.axhline(0, color='black', linewidth=0.5)
plt.title('{} Subtype'.format(name), fontsize=18)
plt.ylim(min_val * 1.2, max_val * 1.2)
plt.xlabel('Gene Index', fontsize=16)
plt.ylabel('SHAP Value', fontsize=16)
plt.tick_params(labelsize=14)
# Label most important genes
for i in range(len(values)):
if values[i] > 0.05:
plt.text(i - 5, values[i] + 0.02, data.columns[i], fontsize=14)
elif values[i] < (- 0.05):
plt.text(i - 5, values[i] - 0.06, data.columns[i], fontsize=14)
plt.tight_layout()
plt.show()
# -
|
notebooks/brca.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # UNCLASSIFIED
#
# Transcribed from FOIA Doc ID: 6689695
#
# https://archive.org/details/comp3321
# # (U) Regular Expressions (Regex)
# ## (U) Now You’ve Got Two Problems...
#
# > Some people, when confronted with a problem, think "I know, I'll use regular expressions." Now they have two
# problems.
# - <NAME>, 1997
#
# (U) A regular expression is a tool for finding and capturing patterns in text strings. It is very powerful and can be very complicated; the second problem referred to in the quote is a commentary on how regular expressions are essentially a separate programming language. As a rule of thumb, use the in operator or string methods like `find` or `startswith` if they are suitable for the task. When things get more complicated, use regular expressions, but try to use them sparingly, like a seasoning. At times it may be tempting to write one giant, powerful, super regular expression, but that is probably not the best thing to do.
#
# (U) The power of regular expressions is found in the special characters. Some, like `^` and `$`, are roughly equivalent to string methods `startswith` and `endswith`, while others are more flexible, especially `.` and `*`, which allow flexible matching.
# ## (U) Getting Stuff Done without Regex
"mike" in "so many mikes!"
"mike".startswith("mi")
"mike".endswith("ke")
"mike".find("k")
"mike".isalpha()
"mike".isdigit()
"mike".replace("k", "c")
# ## (U) Regular expressions in Python
#
# There are only a few common methods for using the **re** module, but they don't always do what you would first expect. Some functionality is exposed through _flags_, which are actually constants (i.e. `int` defined for the **re** module), which means that they can be combined by addition.
import re
re.match("c", "abcdef")
re.match("a", "abcdef")
re.search("c", "abcdef")
re.search("C", "abcdef")
re.search("C", "abcdef", re.I) # re.IGNORECASE
re.search("^c", "ab\ncdef")
re.search("^c", "ab\ncdef", re.M) # re.MULTILINE
re.search("^C", "ab\ncdef", re.M + re.I)
# (U) In both `match` and `search`, the _regular expression_ precedes the string to search. The difference between the two functions is that `match` works only at the beginning of the string, while `search` examines the whole string.
#
# (U) When repeatedly using the same regular expression, _compiling_ it can speed up processing. After a compiled regular expression is created, `find`, `search`, and other methods can be called on it, and given only the search string as a single argument.
c_re = re.compile("c")
c_re.search("abcde")
# ## Regex Operators
#
# ```
# . - matches any character but the newline character. Wildcard
# ^ - matches beginning of a string or newline
# $ - matches end of string
# * - 0 or more of something
# + - 1 or more of something
# ? - 0 or 1 of something
# *?, +?, ?? - don’t be greedy (see example below)
# {3} - match 3 of something
# {2,4} - match 2 to 4 of something
# \ - escape character
# [lrnLRN] - match any ONE of the letters l, r, n, L, R, N
# [a-m] - match any ONE of letters from a to m
# [a|m] - match letter a or m
# \w - match any letter, number, or underscore. Word characters
# \W - match any character that is NOT a letter, number, or underscore
# \s - match a space, tab, or newline character
# \S - match any character that is NOT a space, tab, or newline character
# \d - match a digit 0-9
# \D - match any character that is NOT a digit 0-9
# ```
re.search("\w*s$", "Mike likes cheese\nand Mike likes bees", re.M)
re.findall("\(\d{3}\)\d{3}-\d{4}", "Hello, I am a very bad terrorist. If you wanted to know, my phone number is (303)555-2345")
# greedy search will match everything between the 1st 'mi' and the last 'ke'
re.findall("mi.*ke", "i am looking for mike and not all this stuff in between mike, but micheal and ike is okay.")
# the '?' tells python we want a non-greedy search. It will only match from the first 'mi' to the first 'ke'
re.findall("mi.*?ke", "i am looking for mike and not all this stuff in between mike, but micheal and ike is okay.")
# ### Interlude
#
# How would we have recognized the bad terrorist's phone number without a regex? We could write a function that could recognize phone numbers. What would that function look like?
# +
def match_phone_numbers(text):
if len(text) != 13:
return False
if text[0] != '(' and text[4] != ')':
return False
if text[8] != '-':
return False
for i in range(1, 4):
if not text[i].isdecimal():
return False
for i in range(5, 8):
if not text[i].isdecimal():
return False
for i in range(9, len(text)):
if not text[i].isdecimal():
return False
return True
terror_message = "Hello, I am a very bad terrorist. If you wanted to know, my phone number is (303)555-2345"
for word in terror_message.split():
if match_phone_numbers(word):
print('Phone number found!')
print(word)
# -
# That function took up 16 lines and it can really only match phone numbers that are in the same format as our bad terrorist's number: (303)555-2345. What if there's a space between the area code and the main number? `.split()` will treat that as two separate words and we won't be able to match it. What if someone writes the area code separated from the rest of the number by a `-` instead of by parentheses `()`? The phone number is now going to be 12 characters long instead of 13 and our length check won't work anymore.
#
# There are other ways of breaking chunks of text up besides `.split()` that might help us, but regular expressions are ideal when you're looking for patterns instead of exact text because they provide the language for setting up our matches for us.
#
# Here's an example of a regular expression that will match multiple phone number formats:
phone_re = re.compile(r'(\(?\d{3}\)?-?\s?)?(\d{3}-\d{4})')
phone_re.search("Hello, I am a very bad terrorist. If you wanted to know, my phone number is 303-555-2345")
phone_re.search("Hello, I am a very bad terrorist. If you wanted to know, my phone number is (303) 555-2345")
phone_re.search("Hello, I am a very bad terrorist. If you wanted to know, my phone number is 555-2345")
phone_re.search("Hello, I am a very bad terrorist. If you wanted to know, my phone number is (303)555-2345")
# Let's look at our regular expression and break it down:
#
# `r'(\(?\d{3}\)?-?\s?)?(\d{3}-\d{4})'`
#
# Notice there is an `r` in front of our regular expression. It's very common to set up our regular expression patterns as raw strings since regular expression patterns usually have lots of escape characters in them. This can sometimes make things easier, especially if we need to match a literal backslash. It's probably a good idea to get in the habit of using raw strings for patterns even though it isn't always necessary.
#
# After the raw string starts we notice we have several parentheses in our pattern. Some of them are escaped with backslashes and others aren't. The escaped parenthesis characters are literal parenthesis and the non-escaped parenthesis are setting up an optional capture group. More about that in the next section, but basically most of our regular expression pattern is for matching different types of area codes and will let our pattern match a phone number that doesn't even have an area code.
#
# `(\(?\d{3}\)?-?\s?)?` is the part of our pattern that is specific to the area code. The outermost parenthesis set up the capture group for the area code and the trailing question mark makes the whole capture group optional.
#
# - `(` Start the capture group.
# - `\(?` Optionally match a literal opening parenthesis.
# - `\d{3}` Match three numeric characters.
# - `\)?` Optionally match a closing parenthesis.
# - `-?` Optionally match a hyphen.
# - `\s?` Optionally match a space.
# - `)?` Close the capture group and make the whole thing optional
#
# That takes care of the area code, but what about the remaining bits?
#
# - `(` Start a capture group for the main part of the phone number.
# - `\d{3}` Match three numbers.
# - `-` Match a hyphen.
# - `\d{4}` Match four numbers.
# - `)` Close our second capture group and keep it non-optional.
#
# The options in this section are not optional so something that looks like 555-1212 will always match whether there's an area code or not.
#
# This regular expression isn't perfect. It will match some weird but unlikely strings.
phone_re.findall('These are not phone numbers: (000)- 000-0000, 000)000-0000, (000- 000-0000')
# Even though it is catching some things we might want to exclude, it should catch all American style phone numbers. We could also improve it by adding a capture group to catch country codes. Sometimes you have to decide how much tolerance you have for false positives and whether it's worth the extra effort to craft a more precise regular expression.
# ### Adding some logic
#
# The previous problem with area codes matching can be solved by adding some or `|` operators to match more accurately.
#
# `(\d{3}-|\(\d{3}\)\s?)?` This capture group will capture either three digits without parens followed by a - or three digits surrounded with parenthesis and followed by an optional space.
#
# Let's try it on our bad phone numbers and some good phone numbers and see what it finds.
phone_re = re.compile(r'(\d{3}-|\(\d{3}\)\s?)(\d{3}-\d{4})')
# +
phone_nums = '(000)- 000-0000, 000)000-0000, (000- 000-0000, 555-1212, (801)555-1212, (801) 555-1212, 801-555-1212'
phone_re.findall(phone_nums)
# -
# Success? Sort of. It still matched the main parts of our invalid phone numbers because they still match but at least it excluded our invalid area codes. These days the area code isn't really optional anymore so maybe we can just drop the question mark from that capture group and require our matches to have valid area codes. As long as the area code portion is optional it's still going to match on the valid main number. There's more we could do to fine tune this to avoid matching invalid area codes but this seems like a good place to stop and move on. Let's learn more about capture groups.
# ## Capture Groups
#
# Put what you want to pull out of the strings in parentheses ()
my_string = "python is the best language for doing 'pro'gramming"
result = re.findall("'(\w+)", my_string)
print(result)
print(result[0])
# ## Matches and Groups
#
# (U) The return value from a successful call of `match` or `search` is a _match object_; an unsuccessful call returns `None`. First, this is suitable for use in `if` statements, such as `if c_re.search("abcde"): ...`. For complicated regular expressions, the match object has all the details about the substring that was matched, as well as any captured groups, i.e. regions surrounded by parentheses in the regular expression. These are available via the `group` and `groups` methods. Group 0 is always the whole matching string, after which remaining groups (which can be nested) are ordered according to the opening parenthesis.
m = re.match(r"(\w+) (\w+)", "<NAME>, physicist")
m.group()
m.group(1)
m.group(2)
m.groups()
# ## Other Methods
#
# (U) Other regular expression methods work through all matches in the string, although what is returned is not always straightforward, especially when captured groups are involved. We demonstrate out some basic uses without captured groups. When doing more complicated things, please remember: be careful, read the documentation, and do experiments to test!
re.findall("a.c", "abcadcaecafc") # returns list of strings
iterre = re.finditer("a.c","abcadcaecafc") # returns iterator of match objects
next(iterre)
re.split("a.", "abcadcaecafc") # returns list of strings.
# (U) The `sub` method returns a modified copy of the target string. The first argument is the regular expression to match, the second argument is what to replace it with -- which can be another string or a function, and the third argument is the string on which the substitutions are to be carried out. If the sub method is passed a function, the function should take a single match object as an argument and return a string. For some cases, if the substitution needs to reference captured groups from the regular expression, it can do so using the syntax `\g<number>`, which is the same as accessing the groups method within a function.
re.sub("a.*?c", "a--c", "abracadabra")
re.sub("a(.*?)c", "a\g<1>\g<1>c", "abracadabra")
def reverse_first_group(matchobj):
match = matchobj.group()
rev_group = matchobj.group(1)[::-1]
return match[:matchobj.start(1)] + rev_group + match[matchobj.end(1):]
re.sub("a(.*?)c", reverse_first_group, "abracadabra")
# (U) In the above, we used `start` and `end`, which are methods on a match object that take a single numeric argument -- the group number -- and return the starting and ending indices in the string of the captured group.
#
# (U) One final warning: if a group can be captured more than once, for instance when its definition is followed by a `+` or a `*`, then only the last occurrence of the group will be captured and stored.
# ## Resources:
#
# - Regular Expression Tester https://regex101.com/
# - Paste in some text to match against and see how different patterns will match against that text.
# - Python RegEx Module Documentation https://docs.python.org/3/library/re.html
# - Read the docs.
# - The book Automate the Boring Stuff with Python has a very good chapter about regular expressions. It's available in safari books.
# # UNCLASSIFIED
#
# Transcribed from FOIA Doc ID: 6689695
#
# https://archive.org/details/comp3321
|
Module - Regular Expressions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import nltk
import re
nltk.download('punkt')
nltk.download('stopwords')
nltk.download('wordnet')
train=pd.read_csv('./dataset/train.csv')
test=pd.read_csv('./dataset/test.csv')
print(train.shape, test.shape)
test=test.fillna(' ')
train=train.fillna(' ')
test['total']=test['title']+' '+test['author']+test['text']
train['total']=train['title']+' '+train['author']+train['text']
# +
from nltk.stem import WordNetLemmatizer
from nltk.corpus import stopwords
lemmatizer=WordNetLemmatizer()
for index,row in train.iterrows():
filter_sentence = ''
sentence = row['total']
sentence = re.sub(r'[^\w\s]','',sentence) #cleaning
words = nltk.word_tokenize(sentence) #tokenization
words = [w for w in words if not w in stopwords.words('english')] #stopwords removal
for word in words:
filter_sentence = filter_sentence + ' ' + str(lemmatizer.lemmatize(word)).lower()
train.loc[index,'total'] = filter_sentence
# -
train = train[['total','label']]
from sklearn.feature_extraction.text import TfidfTransformer
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfVectorizer
X_train = train['total']
Y_train = train['label']
count_vectorizer = CountVectorizer()
count_vectorizer.fit_transform(X_train)
freq_term_matrix = count_vectorizer.transform(X_train)
tfidf = TfidfTransformer(norm="l2")
tfidf.fit(freq_term_matrix)
tf_idf_matrix = tfidf.fit_transform(freq_term_matrix)
# +
test_counts = count_vectorizer.transform(test['total'].values)
test_tfidf = tfidf.transform(test_counts)
#split in samples
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(tf_idf_matrix, Y_train, random_state=0)
# +
from sklearn.metrics import confusion_matrix
from sklearn.linear_model import LogisticRegression
logreg = LogisticRegression(C=1e5)
logreg.fit(X_train, y_train)
pred = logreg.predict(X_test)
print('Accuracy of Lasso classifier on training set: {:.2f}'
.format(logreg.score(X_train, y_train)))
print('Accuracy of Lasso classifier on test set: {:.2f}'
.format(logreg.score(X_test, y_test)))
cm = confusion_matrix(y_test, pred)
cm
# +
from sklearn.naive_bayes import MultinomialNB
NB = MultinomialNB()
NB.fit(X_train, y_train)
pred = NB.predict(X_test)
print('Accuracy of NB classifier on training set: {:.2f}'
.format(NB.score(X_train, y_train)))
print('Accuracy of NB classifier on test set: {:.2f}'
.format(NB.score(X_test, y_test)))
cm = confusion_matrix(y_test, pred)
cm
# -
X_train = train['total']
Y_train = train['label']
from sklearn.pipeline import Pipeline
from sklearn.externals import joblib
from sklearn import linear_model
from sklearn.feature_extraction.text import TfidfTransformer
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfVectorizer
pipeline = Pipeline([
('vect', CountVectorizer()),
('tfidf', TfidfTransformer(norm='l2')),
('clf', LogisticRegression(C=1e5)),
])
pipeline.fit(X_train, Y_train)
filename = 'model.sav'
joblib.dump(pipeline, filename)
|
fakeNewsDetection.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1. Introduction to Python & Notebooks
# <datahub url>
# This is an interactive jupyter notebook document, accessible at: <http://datahub.berkeley.edu/user-redirect/interact?account=braddelong&repo=22-jupyter-ps01&branch=main&path=econ-135-s2022-ps01.ipynb>
#
# Page down through it, following the instructions…
#
# These computer programming problem set assignments are a required part of the course.
#
# Collaborating on the problem sets is more than okay—it is encouraged! Seek help from a classmate or an instructor or a roommate or a passerby when you get stuck! (Explaining things is beneficial, too—the best way to solidify your knowledge of a subject is to explain it.)
#
# But the work has to be your own: no cutting-&-pasting from others' problem sets, please! We want you to learn this stuff, and your fingers typing every keystroke is an important way of building muscle memory here.
#
# In fact, we strongly recommend that as you work through this notebook, whenever you come to a "code" cell—something intended not for you to read but also to direct the computer—the python interpreter—to do calculations, you (1) click on the code cell to bring it into your browser's focus; (2) click on the `+` button in the toolbar above to create a new code cell just below the one you were now in; and then (3) retype, line-by-line, the computer code in the cell (not the comment lines beginning with `#`s, but the code cells) while trying to figure out what the line of code is intended to tell the python interpreter to do. "Muscle"—in this case, fingertip—memory is an important but undervalued part of "active learning" here at Berkeley. In Germany, however, they have a term for it: _das Fingerspitzengefühl_; it's the kind of understanding-through-the-fingertips that a true expert has.
#
# In this problem set, you will learn how to:
#
# 1. navigate Jupyter notebooks (like this one);
# 2. write and evaluate some basic *expressions* in Python, the computer language of the course;
# 3. call *functions* to use code other people have written; and
# 4. break down python code into smaller parts to understand it.
# 5. Do some initial data explorations with Python in a notebook.
#
# With what looks to be a permanent and long-run partial moving-online of the university, the already important topic of “data science” seems likely to become even more foundational. Hence I am going to try to provide you with an introduction—to “data science”, and to the framework we will be using for problem sets that we hope will make things much easier for you and for us…
#
# When you are finished, satisfied, or stuck, print your notebook to pdf, & upload the pdf to the appropriate assignment bCourses page: <<https://bcourses.berkeley.edu/courses/1511359/assignments/8365146>>
#
# Also, please include with your submission all of your comments on and reactions to this assignment that you want us to know...
# + [markdown] deletable=false
# For reference, you might find it useful to read chapter 3 of the Data 8 textbook: <<http://www.inferentialthinking.com/chapters/03/programming-in-python.html>>. Chapters 1 <<https://www.inferentialthinking.com/chapters/01/what-is-data-science.html>> and 2 <<https://www.inferentialthinking.com/chapters/02/causality-and-experiments.html>> are worth skimming as well...
#
# ----
# + [markdown] deletable=false
# ## 1.1. Why Are We Doing This?
#
# First of all, we are doing this because our section leaders are overworked: teaching online takes more time and effort than teaching in person, and our section leaders were not overpaid before the 'rona arrived on these shores. Taking the bulk of the work of grading calculation assignments off of their backs is a plus—and it appears that the best way for us to do that is to distribute a number of the course assignements to you in this form: the form of a Python computer language "jupyter notebook"
#
# Second, we are doing this because learning jupyter notebooks and python may well turn out to be the intellectual equivalent for you of "eat your spinach": something that may seem unpleasant and unappetizing now, but that makes you stronger and more capable.
#
# A couple of years ago, I had coffee with a student who had graduated in 2007, and who attributed a large chunk of the interesting and successful career he has had in the past decade-and-a-half years—had in spite of the collapse of the job market in 2008-9—to the fact that he had been fluent in Microsoft Office when he left Berkeley. But now, he said—and I agree—it is not fluency in Word, Powerpoint, and (shudder) Excel that allows one to make oneself useful in a white-collar organization doing interesting work while one learns-by-doing, it is rather fluency in a language like Python and the ability to script hooks into various Big Data and other APIs.
#
# An analogy: Back in the medieval European university, people would learn the Trivium—the 'trivial' subjects of Grammar (how to write), Rhetoric (how to speak in public), and lLogic (how to think coherently)—then they would learn the Quadrivium of Arithmetic, Geometry, Music/Harmony, and Astronomy/Astrology; and last they would learn the advanced and professional subjects: Law or Medicine or Theology and Physics, Metaphysics, and Moral Philosophy.
#
# But a student would also learn two more things: how to learn by reading—how to take a book and get something useful out of it, without a requiring a direct hands-on face-to-face teacher; and (2) how to write a fine chancery hand so that they could prepare their own documents, for submission to secular courts or to religious bishops or even just put them in a form where they would be easily legible to any educated audience back in those days before screens-and-attachments, before screens-and-printers, before typewriters, before printing.
#
# In 1999 Python programming language creator <NAME> compared the ability to read, write, and use software you had built or modified yourself to search and analyze data and information collections. Guido predicted that mass programming, if it could be attained, would produce increases in societal power and changes in societal organization of roughly the same magnitude as mass literacy has had over the past several centuries. Guido may be right, and he may be wrong. But what is clear is that your lives may be richer, and you may have more options, if the data science and basic programming intellectual tools become a useful part of your intellectual panoplies.
#
# The Data Science tools may well turn out to be in the first half of the 2000s the equivalent of a _fine chancery hand_, just as a facility with the document formats and commands of the Microsoft office suite were the equivalent of a fine chancery hand at the end of the 1900s: practical, general skills that make you of immense value to most if not nearly all organizations. This—along with the ability to absorb useful knowledge without requiring hands-on person-to-person face-to-face training—will greatly boost your social power and your set of opportunities in your life.
#
# If we are right about its value.
#
# Why Jupyter and Python, rather than R-studio and R, or C++ and Matlab? Because Jupyter Project founder <NAME> has an office on the fourth floor of Evans. Because 40% of Berkeley undergraduates currently take Data 8 and so, taking account of other channels, more than half of Berkeley students are already going to graduate literonumerate in Python.
#
# Is fluency in something like Python in particular and data-science tools in general is becoming, for the 21st Century, the equivalent of what learning to write a fine chancery hand was if you went to Oxford or Cambridge University in the 14th Century? We do not know. We do think we should help you get ready.
#
# Third, why Jupyter and Python, rather than R-Studio and R, or C++ and MATLAB, or even (horrors!) Microsoft Ezcel? Because Jupyter Project founder <NAME> has an office on the fourth floor of Evans. Because half of Berkeley undergraduates currently take Data 8 and so, taking account of other channels, more than half of Berkeley students are already going to graduate literate in python.
#
# Let us get started!
#
# ----
# -
# ## 1.2. Jupyter Notebooks
#
# This browser window is now showing you a computer science object that is called a "Jupyter notebook". A notebook is a place to write programs and view their results, and also to write text. The text and programs are contained in little boxes called "cells". When the computer thinks that you are paying attention to a particular cell—and may click in it, and then edit it—it surrounds the cell with a blue box, and puts a blue line to the left of the cell.
#
# A notebook is an editable document object in which you can write computer programs; view their results; and comment, annotate, and explain what is going on. Project jupyter <https://en.wikipedia.org/wiki/Project_Jupyter> is headquartered here at Berkeley, where jupyter originator and ringmaster <NAME> <https://en.wikipedia.org/wiki/Fernando_Pérez_(software_developer)> works: its purpose is to build human-friendly frameworks for interactive computing. If you want to see what Fernando looks and sounds like, you can load and watch a 15-minute inspirational video by clicking on the line 'YouTubeVideo("Wd6a3JIFH0s")' in the cell immediately below this one, and then clicking on the '▶' symbol in the toolbar at the top of the sub-window this cell appears in:
# +
from IPython.display import YouTubeVideo
# The original URL is:
# https://www.youtube.com/watch?v=Wd6a3JIFH0s
YouTubeVideo("Wd6a3JIFH0s")
# -
# ### 1.2.1. Code Cells
#
# The _cell_ directly below this one contains one line of code in the Python 3 computer language. You _run_ this line as a program—the computer _executes_ the instructions the cell containss—when you click on the '▶' symbol in the toolbar at the top of this sub-window. (You could also have run the cell by, when the computer has focused on it by your clicking on it, pressing 'shift-return' or 'control-return' on your keyboard.) a code cell will execute all of the code it contains.
#
# The line of type in the cell is a _Python expression_.
#
#
# Now try running this cell:
print("Hello, World!")
# and this one:
print("\N{WAVING HAND SIGN}, \N{EARTH GLOBE ASIA-AUSTRALIA}!")
# Every 'print' expression commands the computer to print a line of symbols to the screen. Directions as to what is in the line of symbols that it prints are contained between the '(' and the ')' that follow the initial Python command 'print' on the expression line.
#
# The first of the two code cells above that you just ran is the standard first computer program to write, either if you are a new student or if you are testing out a new machine or testing out writing in a new programming language. it tells the computer to print out to the screen the symbols 'Hello, World!' What did the second code cell print to the screen, and why?
#
# Within a code cell, each 'print' statement's output is printed to the screen on a separate line. The lines are arranged in the order in which the computer, scanning from the top to the bottom of the code cell, encounters the lines.
#
# In the code cell immediately below, write and then execute a line of Python that prints out: "Friends don't let friends program in Excel!"
# +
# **answer**
# -
# Remember the <NAME> video? Let's look back on its code cell:
# This cell tells the computer to enlarge the set of Python commands it understands by finding the `YouTubeVideo` command in the `IPython.display` external library of commands, and then importing that command into its memory. Once it has done that, you can then ask the computer to execute that command—which then constructs a little YouTube video window and plays a YouTube video in it. You ask the computer to do so the in the fourth line of the code cell. Which YouTube video does the computer play? You tell it by writing, after `YouTubVideo` a `(`, then a `"`, then the reference ID of the video, then another `"`, and then, finally, a `'`.
#
# Between the first and the last lines of the code cell are two lines that are _comment lines_. Anything on a line that comes after a `#` symbol inside a code cell the computer takes to be a comment—something directed at humans, and not at computers, that it should ignore. The comment lines tell you—us—what the URL is for the video that the fourth line is going to command the computer to play.
#
# The cell also contains two blank lines. They are just there to give your eyes and brain visual clues to where to look for information, and thus make code cells easier to read.
#
# ----
# +
from IPython.display import YouTubeVideo
# The original URL is:
# https://www.youtube.com/watch?v=Wd6a3JIFH0s
YouTubeVideo("Wd6a3JIFH0s")
# -
# In the code cell immediately below, write and then execute a line of Python that plays your favorite Youtube video inside this notebook:
# +
# **answer**
# -
# ### 1.2.2. Text cells
# In a notebook, each rectangle containing text or code is called a *cell*.
#
# Text cells (like this one) can be edited by double-clicking on them. They're written in a simple format created by programmer-flaneur <NAME> called **markdown** <<http://daringfireball.net/projects/markdown/syntax>> to add formatting and section headings. You almost surely want to learn how to use markdown.
#
# After you edit a text cell, click the 'run cell' button at the top that looks like `▶` in the toolbar at the top of this window, or press 'shift-return' on your keyboard to confirm any changes to the text and formatting.
# **Question.** This paragraph is in its own text cell. Try editing it so that this sentence is the last sentence in the paragraph, and then click the 'run cell' `▶` button or 'shift-return'. This sentence, for example, should be deleted. So should this one.
# + [markdown] deletable=false
# **Answer.** This paragraph is in its own text cell. Try editing it so that this sentence is the last sentence in the paragraph, and then click the 'run cell' `▶` button or 'shift-return'.
# -
# ----
# ### 1.2.3. Writing notebooks
# You can use Jupyter notebooks for your own projects or documents. When you make your own notebook, you'll need to create your own cells for text and code.
#
# To add a cell, click the + button in the menu bar. It will start out as either a markdown text or a code cell. You can change it to the other type by clicking inside it so it's highlighted, clicking the drop-down box to the left of 'Python 3' in the menu bar, and choosing either 'Code' or 'Markdown'.
#
#
#
# **Question** Add a code cell below this one. Write code in it that prints out:
#
# A whole new cell! ♪🌏♪
#
# (That musical note symbol is like the Earth symbol. Its long-form name is
#
# `\N{EIGHTH NOTE}`
#
# .)
#
# Run your cell to verify that it works.
# +
# **answer**
# -
# The expression
#
# `"A whole new cell! \N{EIGHTH NOTE} \N{EARTH GLOBE ASIA-AUSTRALIA} \N{EIGHTH NOTE}"`
#
# is just a string of symbols. The quotation marks at the start and the end tell Python that it is a string of symbols. Similarly, the expression
#
# `"a"`
#
# is a (very short) string of symbols. What happens when we tell Python to print it?
print("a")
# What if we take out the quotation marks?
print(a)
# ----
# ### 1.2.4. "Errors"
#
# Python is a language, and like natural human languages, it has rules. It differs from natural language in two important ways:
#
# 1. The rules are *simple*. You can learn most of them in a few weeks and gain reasonable proficiency with the language in a semester.
# 2. The rules are *rigid*. If you're proficient in a natural language, you can understand a non-proficient speaker, glossing over small mistakes. A computer running Python code is not smart enough to do that.
#
# Whenever you write code, you'll make mistakes. When you run a code cell that has errors, Python will sometimes produce error messages to tell you what you did wrong.
#
# Errors are okay; even experienced programmers make many errors. When you make an error, you just have to find the source of the problem, fix it, and move on.
#
# When you run code, you have the option to, in the window menubar, click 'Run > Run All Cells', which will run all the code cells in the notebook in order. However, the notebook stops running code cells if it hits an error.
#
# We have made an error in the next code cell. Run it and see what happens. Then correct it:
# + deletable=false
print("This line is missing something."
# -
# When you ran the code cell above, you ought to have seen (minus the Data 8 staff's annotations) something like this:
#
# <img src="images/error.jpg" width="600" />
#
# The last line of the error output attempts to tell you what went wrong. The *syntax* of a language is its structure, and this 'SyntaxError' tells you that you have created an illegal structure. 'EOF' means 'end of file', so the message is saying Python expected you to write something more (in this case, a right parenthesis) before finishing the cell.
#
# There's a lot of terminology in programming languages, but you don't need to know it all in order to program effectively. If you see a cryptic message like this, you can often get by without deciphering it. (Of course, if you're frustrated, ask a neighbor or a staff member for help.) Googling the error message is often a very good strategy as well:
#
# <img src="images/googling-the-error-message-essential-book-orly_jpg_735×900_pixels.png" width="600" />
#
# Try to fix the code above so that you can run the cell and see the intended message instead of an error.
# + for_assignment_type="solution"
# **answer**
# -
# The first rule of computer programming: MAKING ERRORS IS NOT A PROBLEM. MAKING ERRORS IS AN OPPORTUNITY.
#
# ----
# ### 1.2.5. The Kernel
# The kernel is a program that executes the code inside your notebook and outputs the results. In the top right of your window, you can see a circle that indicates the status of your kernel. If the circle is empty (⚪), the kernel is idle and ready to execute code. If the circle is filled in (⚫), the kernel is busy running some code.
#
# Next to every code cell, you'll see some text that says 'In [...]'. Before you run the cell, you'll see 'In [ ]'. When the cell is running, you'll see 'In [*]'. If you see an asterisk (\*) next to a cell that doesn't go away, it's likely that the code inside the cell is taking too long to run, and it might be a good time to interrupt the kernel (discussed below). When a cell is finished running, you'll see a number inside the brackets, like so: 'In [1]'. The number corresponds to the order in which you run the cells; so, the first cell you run will show a 1 when it's finished running, the second will show a 2, and so on.
#
# You may run into problems where your kernel is stuck for an excessive amount of time, your notebook is very slow and unresponsive, or your kernel loses its connection. If this happens, try the following steps:
#
# 1. At the top of your screen, click **Kernel**, then **Interrupt**.
# 2. If that doesn't help, click **Kernel**, then **Restart**. If you do this, you will have to run your code cells from the start of your notebook up until where you paused your work.
# 3. If that doesn't help, restart your server. First, save your work by clicking **File** at the top left of your screen, then **Save and Checkpoint**. Next, click **Control Panel** at the top right. Choose **Stop My Server** to shut it down, then **Start My Server** to start it back up. Then, navigate back to the notebook you were working on. You'll still have to run your code cells again.
#
# ----
# ### 1.2.6. Libraries
# There are many add-ons and extensions to the core of python that are useful—indeed essential—to using it to get work done. They are contained in what are called libraries. The rest of this notebook needs three libraries. So let us tell the python interpreter to install them. Run the code cell below to do so:
# +
# install the numerical python, python data analysis, and mathematical
# plotting libraries for python
# !pip install numpy
# !pip install pandas
# !pip install matplotlib
import numpy as np
import pandas as pd
import matplotlib as mpl
# -
# ----
# # 2. Programming in Python
#
# ## 2.1. Python: Numbers & Variables
#
# ### 2.1.1. Variables
#
# Remember our line that went wrong: `print(a)`? The Python interpreter kernel burped up an error. It said "name is not defined". What does this mean? It means that, without the quotation marks telling Python the a is a string, it thought that the a was the name of something—a variable. But it wasn't. So it did not know what to do.
#
# Let's fix that:
a = "Hello, World!"
# This is an *assignment* statement. It assigns whatever is on the right of the equals sign as the *value* of what the name on the left is. In natural language, we have terminology that lets us quickly reference very complicated concepts. We don't say, "That's a large mammal with brown fur and sharp teeth!" Instead, we just say, "Bear!" In Python, we do this with *assignment statements*—something that has a name on the left side of an '=' sign, and an expression that can be evaluated on the right.
#
# Now what do you think will happen if we tell Python to
#
# `print(a)`
#
# ?
#
# Write a code cell and find out!
# +
# **answer**
# -
# ----
#
# Suppose that we assign something else to a?
# +
a = 3.25
print(a)
a = 6.02 * 10**23
print(a)
a = 1 * 2 * 3 * 4 * 5 * 6 * 7 * 8 * 9 * 10
print (a)
# -
# **Note**: If, on the last line of a cell, we leave off the `print`:
a
# When you run a notebook cell, if the last line has a _value_, then Jupyter helpfully prints out that value for you. However, it won't print out prior lines automatically:
print(2)
3
4
# Above, you should see that 4 is the value of the last expression, 2 is printed because it is within the scope of the print command, but 3 is lost forever because it was neither printed nor last.
#
# If we want Python to move to the next line as it prints, we use `\n` to tell it to do so:
print("stuff: now move to the next line \nmore stuff ")
# In the code cell below, write and execute a line of Python to print "stuff" on the first line, skip the second line, and then print "more stuff" on the third line:
# +
# **answer**
# -
# ----
# ### 2.1.2. Arithmetic
# The line in the next cell subtracts. Its value is what you'd expect. Run it:
3.25 - 1.5
# Many basic arithmetic operations are built into Python. The textbook section on [Expressions](http://www.inferentialthinking.com/chapters/03/1/expressions.html) describes all the arithmetic operators used in the course. The common operator that differs from typical math notation is '\*\*', which raises one number to the power of the other. **NOT '^'.** Thus '2**3' stands for $2^3$ and evaluates to 8.
#
# The order of operations and parentheses is the same as what you learned in elementary school. For example, compare:
3+6*5-6*3**2*2**3/4*7
# to:
4+(6*5-(6*3))**2*((2**3)/4*7)
# In standard math notation, the first expression is
#
# >$3 + 6 \times 5 - 6 \times 3^2 \times \frac{2^3}{4} \times 7$,
#
# while the second expression is
#
# >$3 + (6 \times 5 - (6 \times 3))^2 \times (\frac{(2^3)}{4} \times 7)$.
# There are some things to beware of in Python with respect to the standard arithmetic symbols:
print("2+2 =", 2+2)
print("'2'+'2' =", "2"+"2")
# If the designers of Python had been ruthlessly pedantic, they might have made us write
#
# define the name ten to hereafter have the value of 3 * 2 + 4
#
# instead. You will probably appreciate the brevity of `=`! But keep in mind that this is the real meaning. `=` does not assert or declare that what is on the left is equal to whqt is on the right. Rather, it takes what is on the right and stuffs it into the metaphorical box which has the name on the left.
#
# Suppose that you do want to use the equals sign in the normal way? You have to use `==`. Here is an example:
# +
a = 4
if (a == 4):
print("everything is ok")
else:
print("something is wrong!")
# +
a = 3
if (a == 4):
print("everything is ok")
else:
print("something is wrong!")
# +
a = 4
if (a = 4):
print("everything is ok")
else:
print("something is wrong!")
# -
# See? That last made the Python interpeter barf. By contrast, in the first of the three code cells, Python checked to make sure that variable a was in fact equal to four, and then reassured us. In the second of the thress code cells, Python checked, found out that a was not in fact equal to three, and called for help. But it just could not handle the third.
#
# ----
# A common pattern in Jupyter notebooks is to assign a value to a name and then immediately evaluate the name in the last line in the cell so that the value is displayed as output.
close_to_pi = 355/113
close_to_pi
# Another common pattern is that a series of lines in a single cell will build up a complex computation in stages, naming the intermediate results.
semimonthly_salary = 841.25
monthly_salary = 2 * semimonthly_salary
number_of_months_in_a_year = 12
yearly_salary = number_of_months_in_a_year * monthly_salary
yearly_salary
# Names in Python can have letters (upper- and lower-case letters are both okay, and count as different letters), underscores, and numbers. The first character can't be a number (otherwise a name might look like a number). And names can't contain spaces, since spaces are used to separate pieces of code from each other.
#
# Other than those rules, what you name something doesn't matter *to Python*. For example, this cell does the same thing as the above cell, except everything has a different name:
a = 841.25
b = 2 * a
c = 12
d = c * b
d
# **However**, *names are very important for making your code readable* to yourself and others. The cell above is shorter, but it's totally useless without an explanation of what it does. When you write any computer code, you must do so assuming that it will need to be read and used by the biggest idiot in the known world. Why? Because when you come back to your own code and need to use it, whether next year, next month, next week, tomorrow, or after dinner, you will soon discover that you are that idiot.
#
# ----
# ### 2.1.3. Lists
#
# Lists are ordered collections of objects that have an order. Lists allow us to store groups of variables under one name. The order then allows us to access the objects in the list for easy access and analysis. If you want an in-depth look at the capabilities of lists, take a look at <<https://www.tutorialspoint.com/python/python_lists.htm>>
#
# To initialize a list, you use brackets. Putting objects separated by commas in between the brackets will add them to the list. For example, we can create and name an empty list:
list_example = []
print(list_example)
# We can add an object to the end of a list:
list_example = list_example + [5]
print(list_example)
# Now we have a one-element list. And we can add another element:
list_example = list_example + [10]
print(list_example)
# to make a two-element list.
# In the code cell below, make and then print to screen a five-element list called "five_element_list", in which the elements are: 5, 4, 3, 2, "colorless green ideas":
# +
# **answer**
# -
# We can join—"concatenate"—two lists together:
#
list_example_two = list_example + [1, 3, 6, 'lists', 'are', 'fun', 4]
print(list_example_two)
# >**Digression**: It is, I think, a mistake for python to use '+' in this way. In arithmetic, '+' is simply addition. With lists, '+' smashes the two lists on either side together to make a bigger list. this 'overloading' of '+' can be a source of great confusion. For example:
# +
# overloading of the `+` operator considered harmful:
four = 4
print(" This '4' is a number:", four, "; so '+' is addition \n and so", four, "+", four, "=",
four + four)
four = [4]
print("\n This '4' is a list:", four, "; so '+' is list concatenation \n and so", four, "+", four, "=",
four + four)
four = '4'
print("\n Even worse is: this '4' is a string-of-symbols: \n", four, "; so '+' is symbol concatenation \n and so", four, "+", four, "=",
four + four)
# -
# To access not the list as a whole but an individual value in the list, simply count from the start of the list, and put the place of the object you want to access in brackets after the name of the list.
#
# **But you have to start counting from not one but zero.**
#
# Thus the initial object of a list has index 0, the second object of a list has index 1, and in the list above the eighth object has index 7:
selected_example = list_example_two[7]
print(selected_example)
# which is indeed the eighth item of 'list_example_two'.
#
# (This can become an enormous pain! I find it useful to keep my head straight to talk about "list item 7" and "the 8th item on the list"—which are, in fact, the same.)
#
# You may have noticed already that lists do not have to be made up of elements of the same kind.
#
# Indices do not have to be taken one at a time, either. Instead, we can take a 'slice;\' of indices, and return the elements at those indices as a separate list.
#
# Suppose we just want to select out items 4 through 6 from a list. We can do so:
selected_list = list_example_two[4:7]
print(selected_list)
# We can select out the largest and smallest items of a list via `min` and `max`:
# +
# A list containing six integers.
a_list = [1, 6, 4, 8, 13, 2]
# Another list containing six integers.
b_list = [4, 5, 2, 14, 9, 11]
print('Max of a_list:', max(a_list))
print('Min of b_list:', min(a_list))
# -
# ----
#
# A list is flexible: its elements can be anything. But often we want our lists to be things that we can do arithmetic on quickly and simply. to help us do this, we have the numerical python library: `numpy` or `np`, and the type pf lists we call _arrays_.
#
# Let us start by making an array that consists of the numbers from zero to nine:
# +
import numpy as np
example_array = np.array([0, 1, 2, 3, 4, 5, 6, 7, 8, 9])
print("Example Array:", example_array)
# -
# This could have been accomplished more quickly:
example_array_2 = np.arange(10)
print('Another Example Array: ', example_array_2)
# Multiplying a list and an array by a number produce different results:
# +
b_list = [4, 5, 2, 14, 9, 11]
print('Multiplying a list by 2: ', 2 * b_list)
b_array = np.array([4, 5, 2, 14, 9, 11])
print('Multiplying an array by 2: ', 2 * b_array)
# -
# ----
# ## 2.2. Looping
#
# Loops <<https://www.tutorialspoint.com/python/python_loops.htm>> are useful in manipulating, iterating over, or transforming large lists and arrays. The __for loop__ is useful in that it travels through a list, performing an action at each element. The following code cell moves through every element in example_array, adds it to the previous element in example_array, and copies this sum to a new array.
# +
# loop for calculating the labor force in the
# model over the next 100 years
Labor_Force = []
L = 10
n = .01
T = 100
for i in range(T + 1):
Labor_Force = Labor_Force + [L]
L = L * (1 + n)
print(Labor_Force)
# -
# The most important line in the above cell is the 'for i in...' line. This statement sets the structure of our loop, instructing the machine to stop at every number in `range(T+1)`, perform the indicated operations, and then move on. Once Python has stopped at every element in `range(T+1)`, the loop is completed and the final line, which outputs `Labor_Force`, is executed.
#
# In the code cell below, write a loop to calculate the labor force over the next hundred years if (a) its initial value is 50, and (b) if its growth rate is 2% per year:
# +
# **answer**
# -
# We have done our calculation. But the form in which the computer presents it to us is nearly unreadable.
#
# So now let us do a little magic, using the `pandas` or `pd` library::
# +
# calculating the labor force for the next 100 years
import numpy as np
import pandas as pd
Labor_Force = []
L = 100
n = .01
for i in range(101):
Labor_Force = Labor_Force + [L]
L = L * (1 + n)
pd.DataFrame(Labor_Force).plot(title = "Labor Force", xlabel = "Year",
ylabel = "Value")
# -
# Once we have told Python that we want it to think of the list `Labor Force` as a **pandas DataFrame**, simply adding `.plot()` at the end of our expression will create a graph. And we can put the labels and such for the graph inside the final `()`. We are not limited to simply plotting one series:
# +
# calculating the labor force and the efficiency-of-labor for the next 100 years
Year = []
Labor_Force = []
L = 10
n = 0.01
Efficiency_of_Labor = []
E = 10
g = 0.02
for i in range(101):
Labor_Force = Labor_Force + [L]
L = L * (1 + n)
Efficiency_of_Labor = Efficiency_of_Labor + [E]
E = E * (1 + g)
Year = Year + [i]
pd.DataFrame([Year, Labor_Force, Efficiency_of_Labor]
).transpose().plot()
# -
# And, once again, we can place legends and clean-up the graph by putting things inside the final parentheses:
pd.DataFrame([Year, Labor_Force, Efficiency_of_Labor]
).transpose().plot(
title = "Growth Model Variables", xlabel = "Year",
ylabel = "Value", legend = 0)
# ----
# ## 2.3. Reflection
#
# Do not let any of this scare you. It is about building tools so that you can think better. And, at the bottom, all we are doing is counting things—but doing so by standing on the shoulders of giants, who have figured out how to enable us to count things in incredibly compressed, incredibly powerful ways.
#
# Listen to physicist <NAME> explain how his QED model of light-matter interactions is simply counting things—adding up all the possible different ways a photon might interact with an electron—by analogy:
#
# >“An analogy.... The Maya Indians were interested in... [the War-Star] Venus.... To make calculations, the Maya had invented a system of bars and dots to represent numbers... and had rules by which to calculate and predict not only the risings and settings of Venus, but other celestial phenomena.... Only a few Maya priests could do such elaborate calculations.... Suppose we were to ask one of them how to do just one step in the process of predicting when Venus will next rise as a morning star—subtracting two numbers..... How would the priest explain?....
#
# >He could either teach us the... bars and dots and the rule... or he could tell us what he was really doing: ‘Suppose we want to subtract 236 from 584. First, count out 584 beans and put them in a pot. Then take out 236 beans and put them to one side. Finally, count the beans left in the pot. That number is the result....’ You might say, ‘My Quetzalcoatl! What tedium... what a job!’ To which the priest would reply, ‘That’s why we have the rules.... The rules are tricky, but they are a much more efficient way of getting the answer.... We can predict the appearance of Venus by counting beans (which is slow, but easy to understand) or by using the tricky rules (which is much faster, but you must spend years in school to learn them)’...
#
# Python is the equivalent of learning a bunch of such rules, and practicing applying them.
# ----
# ## 3. Growth Models
# ### 3.1. Behavioral Relationships
# Let us set out the basic Solow Growth Model:
#
# We assume that a particular economy’s labor force L's proportional rate $ g_L $ is a constant _for that particular economy_ n:
#
# > $ \frac{dL}{dt} = g_LL = nL $
#
# We also assume that a particular economy’s efficiency-of-labor E's proportional growth rate g is a constant _for that particular economy_ g:
#
# >$ \frac{dE}{dt} = gE $
#
# We assume the production function for output-per-worker y:
#
# > $ y = \kappa^\theta E $
#
# Total output is equal to the labor force times output per worker:
#
# > $ Y = yL $
#
# The capital stock is equal to capital-intensity times total output:
#
# > $ K = \kappa Y $
#
# Each year a fraction s of total gross output is invested and added to the capital stock, and a fraction $ \delta $ of the capital stock depreciates:
#
# > $ \frac{dK}{dt} = sY - \delta K
#
# The code cell below sets up this model, and prints out a graph of the calculated values for the variables:
# +
# basic solow growth model code cell:
# ----
# PARAMETERS AND INITIAL CONDITIONS BLOCK
T = 500 # number of years for model run
L_0 = 4 # initial labor force
n = 0.02 # labor-force growth rate
E_0 = 1 # initial labor effiency
g = 0.00 # labor efficiency growth rate
kappa_0 = 2 # initial capital-intensity
theta = 1 # salience of capital in the production function
alpha = theta/(1 + theta) # convenient transformation of theta
Y_0 = kappa_0**theta * E_0 * L_0 # initial total output
y_0 = kappa_0**theta * E_0 # initial productivity
K_0 = kappa_0 * Y_0 # initial capital stock
s = 0.2 # savings share of gross output
delta = 0.03 # capital depreciation rate
# END BLOCK
# ----
import numpy as np # import libraries
import pandas as pd
import matplotlib.pyplot as plt
L_series = [L_0] # initialize time-series lists
E_series = [E_0]
Y_series = [Y_0]
y_series = [y_0]
K_series = [K_0]
kappa_series = [kappa_0]
L = L_0 # initialize working variables
E = E_0
Y = Y_0
y = y_0
K = K_0
kappa = kappa_0
for i in range(T): # loop for calculating next-year values
L = (1 + n) * L
E = (1 + g) * E
K = (1 - delta) * K + s * Y
Y = K**alpha * (E * L)**(1 - alpha)
y = K**alpha * E
kappa = K/Y
L_series = L_series + [L] # update time series
E_series = E_series + [E]
K_series = K_series + [K]
Y_series = Y_series + [Y]
y_series = y_series + [y]
kappa_series = kappa_series + [kappa]
solow_df = pd.DataFrame() # create dataframe
solow_df['Labor Force'] = L_series
solow_df['Efficiency of Labor'] = E_series
solow_df['Capital Stock'] = K_series
solow_df['Total Output'] = Y_series
solow_df['Capital-Intensity'] = kappa_series
solow_df['Output per Worker'] = y_series
ax = solow_df.plot() # plot variables
ax.set_title("Model Run")
ax.set_xlabel("Date")
ax.set_ylabel("Value")
plt.show() # show plot
# -
# Suppose we just wanted to graph the labor force. Then—after running the code cell above—we would write this, and get the Python Interpreter to execute it:
# +
ax = solow_df['Labor Force'].plot() # plot the labor force
ax.set_title("Labor Force")
ax.set_xlabel("Date")
ax.set_ylabel("Value")
plt.show() # show plot
# -
# Now you write a code cell, plotting just capital-intensity:
# **answer**
# Now you write a code cell, plotting just output per worker:
# +
# **answer**
# -
# Suppose we started the economy in a different initial condition—with an initial capital-intensity of not 2 but 12. How much difference do we think it would make in the long run?
#
# Below is a copy of the Basic Solow Growth Model code cell. In it, in the `kappa_0 = 2` line, edit the line to change the 2 to 8, and then rerun the cell:
# +
# **answer**
# basic solow growth model code cell cell:
# ----
# PARAMETERS AND INITIAL CONDITIONS BLOCK
T = 500 # number of years for model run
L_0 = 4 # initial labor force
n = 0.02 # labor-force growth rate
E_0 = 1 # initial labor effiency
g = 0.00 # labor efficiency growth rate
kappa_0 = 12 # initial capital-intensity
theta = 1 # salience of capital in the production function
alpha = theta/(1 + theta) # convenient transformation of theta
Y_0 = kappa_0**theta * E_0 * L_0 # initial total output
y_0 = kappa_0**theta * E_0 # initial productivity
K_0 = kappa_0 * Y_0 # initial capital stock
s = 0.2 # savings share of gross output
delta = 0.03 # capital depreciation rate
# END BLOCK
# ----
import numpy as np # import libraries
import pandas as pd
import matplotlib.pyplot as plt
L_series = [L_0] # initialize time-series lists
E_series = [E_0]
Y_series = [Y_0]
y_series = [y_0]
K_series = [K_0]
kappa_series = [kappa_0]
L = L_0 # initialize working variables
E = E_0
Y = Y_0
y = y_0
K = K_0
kappa = kappa_0
for i in range(T): # loop for calculating next-year values
L = (1 + n) * L
E = (1 + g) * E
K = (1 - delta) * K + s * Y
Y = K**alpha * (E * L)**(1 - alpha)
y = K**alpha * E
kappa = K/Y
L_series = L_series + [L] # update time series
E_series = E_series + [E]
K_series = K_series + [K]
Y_series = Y_series + [Y]
y_series = y_series + [y]
kappa_series = kappa_series + [kappa]
solow_df = pd.DataFrame() # create dataframe
solow_df['Labor Force'] = L_series
solow_df['Efficiency of Labor'] = E_series
solow_df['Capital Stock'] = K_series
solow_df['Total Output'] = Y_series
solow_df['Capital-Intensity'] = kappa_series
solow_df['Output per Worker'] = y_series
ax = solow_df.plot() # plot variables
ax.set_title("Model Run")
ax.set_xlabel("Date")
ax.set_ylabel("Value")
plt.show() # show plot
# -
# Now plot capital-intensity for the new model run:
# +
# **answer**
# -
# Now, in the markdown cell below, write a paragraph explaining how different the model run is and what the differences between the model runs are for those two different initial conditions with respect to capital intensity. See iof upoi can give a clear and intuitive explanation for why the differences are what they are:
#
# **Answer**:
# You are finished!
#
# When you are finished, satisfied, or stuck, print your notebook to pdf, & upload the pdf to the appropriate assignment bCourses page: <https://bcourses.berkeley.edu/courses/1511359/assignments/8365146>
#
# Also, please include with your submission all of your comments on and reactions to this assignment that you want us to know...
|
econ-135-s2022-ps01.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from arcgis.gis import GIS
gis = GIS("https://www.arcgis.com", "username", "password")
sf_map = gis.map("San Francisco")
sf_map
# +
from IPython.display import display
items = gis.content.search('SF')
for item in items:
display(item)
# -
add_item = [bus_item for bus_item in items if bus_item.title == "SF_BusStops"]
sf_map.add_layer(add_item[0])
import pandas as pd
dataFrame = pd.read_csv("C:\PythonBook\ch12_ArcGIS_PythonAPI\Ch12_Data\SFPD_2016.csv", encoding = "ISO-8859-1")
dataFrame
dataFrame['PdDistrict'].describe()
import matplotlib.pyplot as py_plot
dataFrame['PdDistrict'].value_counts().plot(kind='bar')
# +
pd_data = r"C:\PythonBook\ch12_ArcGIS_PythonAPI\Ch12_Data\SFPD_2016.csv"
pd_data_properties = {'title': 'SFPD calls for 2016',
'description': 'All the SFPD calls for the year 2016',
'tags': 'SF PD, calls, csv' }
thumbnail_pic = r"C:\PythonBook\ch12_ArcGIS_PythonAPI\Ch12_Data\SF_PD.PNG"
sf_pd_item = gis.content.add(item_properties=pd_data_properties, data=pd_data,
thumbnail = thumbnail_pic)
sf_pd_item
# -
sf_pd_feature_layer = sf_pd_item.publish()
sf_pd_feature_layer
sf_crimemap = gis.map("Divisadero and Haight,San Francisco", zoomlevel=17)
sf_crimemap.add_layer(sf_pd_feature_layer)
sf_crimemap.height = '950px'
sf_crimemap
|
Chapter_12/Add Data to Map.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Comparison of Data Engineering Techniques
from Pipeline import Pipeline
from Compare import Compare
from StructuredData.LoadCSV import LoadCSV
from StructuredData.MissingValues import MissingValues
from StructuredData.Normalize import Normalize
from StructuredData.Factorize import Factorize
import warnings
warnings.filterwarnings('ignore')
csv_path = './DemoData/synthetic_classification.csv'
df = LoadCSV(csv_path)()
df.head(10)
# ### Comparison of pipelines
# In order to compare which feature engineering technique works better, we use pipelines on the model as shown below
# +
p1 = Pipeline([MissingValues(), Normalize(['1','2','3'])])
p2 = Pipeline([MissingValues()])
from sklearn.linear_model import LogisticRegression
model = LogisticRegression(random_state=0, solver='lbfgs', multi_class='ovr')
from sklearn.metrics import accuracy_score
p1_score, p2_score = Compare()(df,'0', model, p1, p2, accuracy_score)
output = "Performance of pipeline 1 is: {} \nPerformance of pipeline 2 is:{}"
print(output.format(p1_score,p2_score))
|
Comparison of pipelines.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
#Problem 1
#Create a generator that generates the squares
#of numbers up to some number N.
# -
def gensquares(N):
for num in range(N):
yield num**2
for x in gensquares(10):
print x
# +
#Problem 2
#Create a generator that yields "n" random
#numbers between a low and high number (that are inputs).
#Note: Use the random library. For example:
import random
random.randint(1,10)
# -
def rand_num(low,high,n):
for num in range(n):
yield random.randint(low,high)
for num in rand_num(1,10,12):
print num
# +
#Problem 3
#Use the iter() function to convert the string below
s='hello'
# -
s2=iter(s)
print next(s2)
# +
#Problem 4
#Explain a reason for using a generator instead of
#a normal function with a return statement.
# +
#When your output will take up a large amount of
#memory and you only intend to iterate through it
# -
# +
#Extra credit.
#Can you explain what gencomp is in the code below?
#Note: We never covered this in lecture, you will have
#to do some googling/stack overflowing
# +
my_list = [1,2,3,4,5]
gencomp = (item for item in my_list if item > 3)
for item in gencomp:
print item
# +
#This is a generator comprehension..instead of brackets [] like
#we use for list comprehensions, we use parenthesis () for
#generator comprehensions
|
Section9.3 Iterators and Generators Homework.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#
# Working through the example from
# https://github.com/kwmcbride/kipet_examples/blob/master/examples/example_7/Ex_7_concentration_input.py
#
# +
import sys
import kipet
import numpy as np
import matplotlib.pyplot as plt
# +
r1 = kipet.ReactionModel('reaction-1')
# Add the model parameters
k1 = r1.parameter('k1', value=2.0, bounds=(0.0, 5.0))
k2 = r1.parameter('k2', value=0.2, bounds=(0.0, 2.0))
# Declare the components and give the initial values
A = r1.component('A', value=0.001, known = False, bounds = (0.0, 3))
B = r1.component('B', value=0.0)
C = r1.component('C', value=0.0)
# -
filename = '/home/paperspace/learn_kipet/data_sets/Ex_1_C_data.txt'
data = kipet.read_data(filename)
data = data.iloc[::10,]
data.head()
r1.add_data(data = data, remove_negatives = True)
# +
# Define the reaction model
rate1 = k1 * A
rate2 = k2 * B
r1.add_ode('A', -rate1)
r1.add_ode('B', rate1 - rate2)
r1.add_ode('C', rate2)
# -
# Settings
r1.settings.collocation.nfe = 60
# +
#r1._create_pyomo_model()
# +
#r1._model.Z.pprint()
# -
# Run KIPET
r1.settings.solver.linear_solver = 'ma27'
r1.run_opt()
# Display the results
r1.results.show_parameters
fig, ax = plt.subplots()
for c in ['A','B','C']:
ax.plot(r1.results.Z.index, r1.results.Z[c])
ax.scatter(r1.results.Cm.index, r1.results.Cm[c])
r1.plot(jupyter=True)
r1.results
r1.results.Cm.index
r1.results.Z.index
tmp = np.isin(r1.results.Z.index, r1.results.Cm.index)
Zpred = r1.results.Z[tmp]
Zpred.shape, r1.results.Cm.shape
fig, ax = plt.subplots()
ax.scatter(Zpred['C'].values, r1.results.Cm['C'])
|
kipet_examples/.ipynb_checkpoints/Ex7_concentration_input_wk.py-checkpoint.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# ## Spatio-temporal functional data analysis for wireless sensor networks data 논문
load("hourly_data.RData")
ls() # 현재 존재하는 R 객체 확인
dim(full_data)
head(full_data)
library(mgcv)
library(ftsa)
library(MFPCA)
################################################################################
#### generate data : X ( time=24*60 days (2014/01/01~2014/3/1), grid=50 in Korea )
################################################################################
grid_pt <- unique(full_data$grid)[1:50] # grid 변수의 unique한 값 50개를 뽑음
X <- vector()
for (i in 1:length(grid_pt)){ # grid별로 각각 24*60개의 시계열 데이터를 뽑아서 rbind
X <- rbind(X, full_data[which(full_data$grid==grid_pt[i]),][1:(24*60),] )
}
colnames(X)[7] <- 'Rdate'
X$time_in_day <- rep(c(1:24), 60) # 시간 변수 추가(60일간의 시간별 데이터)
head(X)
na_index <- which(is.na(X$PM10)==T) # PM10이 NA인 값의 index 저장
na_index
# NA값들을 인접한 10개 데이터의 평균으로 보간
for (k in 1:length(na_index)){
X$PM10[ na_index[k] ] <- mean( X$PM10[ c( (na_index[k]-5):(na_index[k]+5) ) ], na.rm=T) ## interpolate NA terms
}
# +
## to see daily cyclical pattern. (Fig 2-(b)) - raw data version
# Example plot for grid k=1
k <- 1
grid1 <- X[which(X[,1] == grid_pt[k]), ]
i <- 1
tt <- c( (1 + 24*(i-1)) : (24*i) )
plot(grid1$Rdate[tt], grid1$PM10[tt], type='l', main=paste('grid ID =', grid_pt[k]), xlab='Time of day', ylab="PM10", ylim=range(grid1$PM10, na.rm=T) )
for (i in 2:60){
tt <- c( (1 + 24*(i-1)) : (24*i) )
lines(grid1$Rdate[1:24], grid1$PM10[tt], col=i)
}
# -
## Smoothing the data
# 비어있는 matrix 생성
smoothed_X <- matrix(nrow=length(unique(X$grid)), ncol=60*24) ## grid(50) * time(24*60)
dim(smoothed_X)
# +
for (k in 1:length(unique(X$grid))){
grid1 <- X[which(X[,1] == grid_pt[k]), ]
for (i in 1:60){
tt <- c( (1 + 24*(i-1)) : (24*i) )
b <- gam(PM10[tt]~s(time_in_day[tt], bs='cc', k=24), data=grid1, method="REML") # bs="cc": cyclic cubic regression splines
smoothed_X[k, tt] <- b$fitted.values
}
}
k <- 1
i <- 1
tt <- c( (1+24*(i-1) ) : (24*i) )
plot(grid1$Rdate[tt], smoothed_X[k,tt], type='l', main=paste('grid ID =', grid_pt[k]), xlab='Time of day', ylab="PM10", ylim=range(smoothed_X[k,], na.rm=T))
for (i in 2:60){
tt <- c( (1+24*(i-1) ) :(24*i) )
lines(grid1$Rdate[1:24], smoothed_X[k,tt], col=i)
}
# +
##### FPCA (Need to apply into two type of grid (ex. grassland vs forest))
grass_data <- matrix(apply(smoothed_X[1:25,], 2, mean), 24, 60) # grass에 해당하는 모든 grid를 각 시간마다 평균을 낸 데이터
forest_data <- matrix(apply(smoothed_X[26:50,], 2, mean), 24, 60)
fts_object_grass <- fts(x=c(1:24), grass_data) # fts(): functional 객체 생성
fit_grass <- ftsm(fts_object_grass, order=3, method='classical') # ftsm(): functional time series model - fpca, 24시간에 따라 pca(날짜가 아닌)
fts_object_forest <- fts(x=c(1:24), forest_data)
fit_forest <- ftsm(fts_object_forest, order=3, method='classical')
# -
dim(smoothed_X[1:25,])
length(apply(smoothed_X[1:25,], 2, mean))
names(fit_grass)
dim(grass_data)
dim(fit_grass$basis) # PC score & mean(or median)
fit_grass$varprop
par(mfrow=c(1,4))
plot(fit_grass$basis[,1], type='l', main='mean function', xlab='Time of day', ylab='', ylim=range(fit_grass$basis[,1], fit_forest$basis[,1]))
lines(fit_forest$basis[,1], col=2)
# legend('topright', c('grass', 'forest'), lty=1, col=c(1,2))
for (k in 2:4){
plot(fit_grass$basis[,k], type='l', main=paste(k ,'-th PC (phi)'), xlab='Time of day', ylab='', ylim=range(fit_grass$basis[,k], fit_forest$basis[,k]))
lines(fit_forest$basis[,k], col=2)
# legend('topright', c('grass', 'forest'), lty=1, col=c(1,2))
}
# +
#### Smoothed ANOVA
## 1. grass
k <- 2
z <- as.vector(fit_grass$coeff[,k])
z <- rep(z, times=25)
loc_effect <- as.numeric(unique(X$grid)[1:25])
loc_effect <- rep(loc_effect, each=60)
# days_effect= rep( unique(format((X$Rdate), format="%m-%d"))[-61] , 25)
days_effect <- rep(1:60, 25)
anova_data <- data.frame(z, loc_effect, days_effect)
b <- gamm(z~s(loc_effect)+s(days_effect)+te(loc_effect,days_effect), data=anova_data)
anova(b$gam)
|
Spatio-temporal FDA/code.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## Start the 3D Visualizer and the droidlet dashboard
# +
from droidlet.dashboard.o3dviz import O3DViz
# if False, opens a native window for the 3D visualization.
# If True, opens a web-based viewer for the 3D stuff that is available at port http://localhost:8889
web_streaming = False
o3dviz = O3DViz(web_streaming)
o3dviz.start()
from droidlet import dashboard
dashboard.start()
# this has to come after the `dashboard.start` function above
from droidlet.event import sio
# -
# ## Some useful imports
import math
import time
import cv2
from matplotlib import pyplot as plt
import open3d as o3d
import numpy as np
# ## Import and connect to the HelloRobotMover
from droidlet.lowlevel.hello_robot.hello_robot_mover import HelloRobotMover
mover = HelloRobotMover(ip="172.16.58.3") # ip of the robot
# ## Get the rgb, depth and globally-registered point-clouds
rgb_depth = mover.get_rgb_depth()
img = rgb_depth.rgb
plt.imshow(img)
plt.imshow(rgb_depth.depth)
def get_open3d_pcd(rgb_depth):
points, colors = rgb_depth.ptcloud.reshape(-1, 3), rgb_depth.rgb.reshape(-1, 3)
colors = colors / 255.
pcd = o3d.geometry.PointCloud()
pcd.points = o3d.utility.Vector3dVector(points)
pcd.colors = o3d.utility.Vector3dVector(colors)
return pcd
# +
point_cloud = get_open3d_pcd(rgb_depth)
o3dviz.put('pointcloud', point_cloud)
# -
# ## Get the SLAM obstacle map and base state in droidlet canonical co-ordinates
obstacle_map = mover.get_obstacles_in_canonical_coords()
base_state = mover.get_base_pos_in_canonical_coords()
# +
# Plot them
x, y = [m[0] for m in obstacle_map], [m[1] for m in obstacle_map]
plt.plot(y, x, 'o', markersize=1)
plt.plot(base_state[0], base_state[1], 'ro', markersize=12)
xorigin, yorigin = 0., 0.
newx = (base_state[0] - xorigin) * math.cos(base_state[2] * math.pi / 180)
newy = (base_state[1] - yorigin) * math.sin(base_state[2] * math.pi / 180)
plt.plot([base_state[0], newx], [base_state[1], newy], 'b')
# -
# you can also plot it into the dashboard
x, y, yaw = base_state
sio.emit("map",
{"x": x, "y": y, "yaw": yaw, "map": obstacle_map},
)
# ## Move the robot using it's navigation system
# +
# Move forward by 1 metre (x, y, yaw)
mover.move_relative([1.0, 0.0, 0.0], blocking=False)
# turn the robot
mover.turn(-math.radians(20))
# -
# ## Move the robot directly using it's base API
mover.bot.go_to_relative([1.0, 0.0, 0.0])
# ## Move it's camera
# Set the camera's tilt and pan
mover.bot.set_tilt(math.radians(-60) )
mover.bot.set_pan(math.radians(-30) )
# ## Stop the robot's actions immediately
mover.stop()
# ## Access mover's backend services directly via the RPC classes
# +
# mover.bot
# mover.slam
# mover.nav
# mover.cam
# -
|
examples_and_tutorials/notebooks/hello-robot-droidlet-intro.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# -*- coding: utf-8 -*-
from __future__ import unicode_literals
# text in Western (Windows 1252)
import numpy as np
# import StringIO
import math
import keras
from keras.models import Sequential
from keras.layers import Dense, Dropout, Merge
from keras.layers.merge import concatenate
from keras import regularizers
from keras.layers.convolutional import Conv1D
from keras.layers.convolutional import MaxPooling1D
from keras.constraints import maxnorm
from keras.layers import Flatten
from keras.optimizers import SGD
from keras.models import load_model
np.random.seed(7)
# +
# %run ../../../prepare_data.py
# import sys
# sys.path.insert(0, '../../')
# import preprare_data
# +
additional_data = [el[2] for el in content]
possible_variants = sorted(set(additional_data))
# -
np.array(X_other_features).shape
# +
X_other_features = create_X_features(content, feature_dictionary)
# -
feature_dictionary = create_feature_dictionary(content)
feature_dictionary
# X_train, X_other_features_train, y_train, X_validate, X_other_features_validate, y_validate = generate_full_matrix_inputs()
# save_inputs('../../internal_representations/inputs/shuffeled_matrix_train_inputs_other_features_output_11.h5', X_train, y_train, other_features = X_other_features_train)
# save_inputs('../../internal_representations/inputs/shuffeled_matrix_validate_inputs_other_features_output_11.h5', X_validate, y_validate, other_features = X_other_features_validate)
X_train, X_other_features_train, y_train = load_inputs('../../internal_representations/inputs/shuffeled_matrix_train_inputs_other_features_output_11.h5', other_features=True)
X_validate, X_other_features_validate, y_validate = load_inputs('../../internal_representations/inputs/shuffeled_matrix_validate_inputs_other_features_output_11.h5', other_features=True)
# X_other_features = create_X_features(content)
# print (X_other_features[178200])
decode_position = 30
print (decode_input(X_train[decode_position], dictionary))
# print (X_other_features_train[0])
print (decode_X_features(feature_dictionary, [X_other_features_train[decode_position]]))
# print (len(X_other_features_train[0]))
# +
# X = X[:100000]
# y = y[:100000]
# def unison_shuffled_copies(a, b):
# assert len(a) == len(b)
# p = np.random.permutation(len(a))
# return a[p], b[p]
# X, y = unison_shuffled_copies(X, y)
# print X.shape
# train_X = X.astype(np.float32)
# train_y = y.astype(np.int32)
num_examples = len(X_train) # training set size
# nn_input_dim = max_word * len(dictionary) # input layer dimensionality
# nn_output_dim = max_num_vowels * max_num_vowels # output layer dimensionality
nn_output_dim = 11
nn_hdim = 516
# Gradient descent parameters (I picked these by hand)
# epsilon = 1 # learning rate for gradient descent
# reg_lambda = 1 # regularization strength
# -
nn_output_dim
# +
word_processor = Sequential()
word_processor.add(Conv1D(43, (3), input_shape=(23, 43), padding='same', activation='relu'))
word_processor.add(Conv1D(43, (2), padding='same', activation='relu'))
word_processor.add(Conv1D(43, (2), padding='same', activation='relu'))
word_processor.add(MaxPooling1D(pool_size=2))
word_processor.add(Flatten())
word_processor.add(Dense(516, activation='relu', kernel_constraint=maxnorm(3)))
metadata_processor = Sequential()
metadata_processor.add(Dense(256, input_dim=167, activation='relu'))
model = Sequential()
model.add(Merge([word_processor, metadata_processor], mode='concat')) # Merge is your sensor fusion buddy
model.add(Dense(1024, input_dim=(516 + 256), activation='relu'))
model.add(Dropout(0.3))
model.add(Dense(1024, activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(nn_output_dim, activation='sigmoid'))
# np.random.seed(7)
# # create model
# model = Sequential()
# model.add(Conv1D(43, (3), input_shape=(23, 43), padding='same', activation='relu'))
# model.add(Dropout(0.2))
# model.add(Conv1D(43, (3), padding='same', activation='relu'))
# model.add(MaxPooling1D(pool_size=2))
# # model.add(Conv1D(43, (3), input_shape=(None, 43), padding='same', activation='relu', kernel_constraint=maxnorm(3)))
# model.add(Flatten())
# # model.add(Dense(1032, activation='relu', kernel_constraint=maxnorm(3)))
# # model.add(Dropout(0.2))
# model.add(Dense(516, activation='relu', kernel_constraint=maxnorm(3)))
# model.add(Dropout(0.2))
# model.add(Dense(nn_output_dim, activation='sigmoid'))
# +
epochs = 5
lrate = 0.1
decay = lrate/epochs
sgd = SGD(lr=lrate, momentum=0.9, decay=decay, nesterov=False)
model.compile(loss='mean_squared_error', optimizer='adam', metrics=['accuracy'])
# Compile model
# model.compile(loss='mean_squared_error', optimizer='adam', metrics=['accuracy'])
# -
# model.fit_generator((X, y), 536432/2, epochs=5)
# testX = X[:536432/16]
# print testX.shape
# testY = y[:536432/16]
# print testY.shape
model.fit([X_train, X_other_features_train], y_train, validation_data=([X_validate, X_other_features_validate], y_validate), epochs=5, batch_size=16)
# model.fit(X, y, epochs=5, validation_split=0.2, batch_size=10)
print(keras.__version__)
model.save('test.h5')
model = load_model('test.h5')
h5f.close()
# evaluate the model
scores = model.evaluate(X[429145:], y[429145:])
print("\n%s: %.2f%%" % (model.metrics_names[1], scores[1]*100))
# +
# calculate predictions
# test = generate_input_from_word('test', max_word, dictionary)
# print test.shape
# print test.T.shape
predictions = model.predict(np.array([generate_input_from_word('biotski', max_word, dictionary)]))
# round predictions
print decode_position(predictions[0], max_num_vowels)
b_pred = 0
ind = 0
i = 0
for el in predictions[0]:
if b_pred < el:
b_pred = el
ind = i
i += 1
print(ind)
# rounded = [round(x[0]) for x in predictions]
# print(rounded)
# -
predictions = model.predict(X[429145:])
decode_position(predictions[0], max_num_vowels)
# +
def test_accuracy(predictions, y):
dictionary, max_word, max_num_vowels, content, vowels, accetuated_vowels = create_dict()
num_of_pred = len(predictions)
num_of_correct_pred = 0
for i in range(predictions.shape[0]):
if decode_position(predictions[i], max_num_vowels) == decode_position(y[i], max_num_vowels):
num_of_correct_pred += 1
return (num_of_correct_pred/float(num_of_pred)) * 100
print(test_accuracy(predictions, y[429145:]))
# -
predictions.shape
print max_num_vowels
# +
dictionary, max_word, max_num_vowels, content, vowels, accetuated_vowels = create_dict()
feature_dictionary = create_feature_dictionary(content)
def generate_input_from_word(word, max_word, dictionary):
x = np.zeros((max_word, len(dictionary)))
j = 0
for c in list(word):
index = 0
for d in dictionary:
if c == d:
x[j, index] = 1
break
index += 1
j += 1
return x
# model = load_model()
# prediction = predict(model, generate_input_from_word('hidrija'))
# print decode_position(prediction[0])
# -
# %run ../../../prepare_data.py
generate_X_and_y(dictionary, max_word, max_num_vowels, content, vowels, accetuated_vowels, feature_dictionary)
|
cnn/word_accetuation/cnn_dictionary/v1_4/character_based_ffnn_keras.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import h5py
from astropy.table import Table
from astropy.constants import R_sun, R_earth
from glob import glob
X = []
labels = []
for lcpath, truthpath in zip(sorted(glob('../data/parallel_normed/*_simulated_transit_lcs.npy')),
sorted(glob('../data/parallel_normed/*_simulated_spots_occulted.npy'))):
# print(np.shape(np.load(lcpath).T))
X.append(np.load(lcpath).T)
labels.append(np.load(truthpath))
X = np.concatenate(X)[:, :, np.newaxis]# - np.array(X).mean()
X -= X.mean()
X /= X.ptp()
# X += 0.5
labels = np.hstack(labels)#[:, np.newaxis]
train_X = X
train_Y = labels
# print('Training data shape : ', train_X.shape, train_Y.shape)
from keras.utils import to_categorical
train_Y_one_hot = to_categorical(train_Y)
# test_Y_one_hot = to_categorical(test_Y)
from sklearn.model_selection import train_test_split
train_X, valid_X, train_label, valid_label = train_test_split(train_X, train_Y_one_hot,
test_size=0.2, random_state=13)
# +
from keras.models import load_model
import matplotlib.ticker as ticker
from sklearn.metrics import precision_recall_curve
paths = ['../data/model.hdf5'] + glob('data/model?.hdf5')
precisions = []
recalls = []
for path in paths:
model = load_model(path)
predict_y = model.predict(valid_X)
precision, recall, thresh = precision_recall_curve(valid_label[:, 0] == 1, predict_y[:, 0])
precisions.append(precision)
recalls.append(recall)
del model
# +
plt.figure(figsize=(3, 3))
labels = ['Deep', 'Shallow', 'Simplified', 'High Dropout', 'Small kernel', 'Sigmoid', 'SGD']
recall_thresh = 0.0
for i, recall, precision, label in zip(range(len(recalls)), recalls, precisions, labels):
# Skip shallow
if label != 'Shallow':
sort = np.argsort(recall)
recall = recall[sort]
precision = precision[sort]
auc = np.trapz(precision[recall > recall_thresh],
recall[recall > recall_thresh])
plt.plot(recall, precision, label=label+", AUC={0:.3f}".format(auc),
alpha=0.7 if i != 0 else 1, color=None if i != 0 else 'k',
zorder=0 if i != 0 else 10)
plt.xlim([0.3, 1.005])
plt.ylim([0.9, 1.00])
plt.xlabel('Recall')
plt.ylabel('Precision')
plt.legend(loc='lower left', fontsize=8, facecolor='none', frameon=False)
plt.grid(ls=':', color='silver')
plt.gca().xaxis.set_minor_locator(ticker.MultipleLocator(0.1))
plt.savefig('../plots/precision_recall.pdf', bbox_inches='tight')
# -
|
cnn/compare_models/compare.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 模型训练过程分析
#
# ## 引入第三方包
# +
import glob
import pickle
import numpy as np
import matplotlib.pyplot as plt
# -
# ## 加载训练过程记录
history_file = './pre-trained/history/optimizer/binary_ce/captcha_adam_binary_crossentropy_bs_100_epochs_100.history'
with open(history_file, 'rb') as f:
history = pickle.load(f)
# ## 训练过程可视化
# +
fig = plt.figure()
plt.subplot(2, 1, 1)
plt.plot(history['acc'])
plt.plot(history['val_acc'])
plt.title('Model Accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='lower right')
plt.subplot(2, 1, 2)
plt.plot(history['loss'])
plt.plot(history['val_loss'])
plt.title('Model Loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper right')
plt.tight_layout()
plt.show()
# -
# ## 定义过程可视化方法
def plot_training(history=None, metric='acc', title='Model Accuracy', loc='lower right'):
model_list = []
fig = plt.figure(figsize=(10, 8))
for key, val in history.items():
model_list.append(key.replace(HISTORY_DIR, '').rstrip('.history'))
plt.plot(val[metric])
plt.title(title)
plt.ylabel(metric)
plt.xlabel('epoch')
plt.legend(model_list, loc=loc)
plt.show()
# ## 加载预训练模型记录
HISTORY_DIR = './pre-trained/history/optimizer/binary_ce/'
history = {}
for filename in glob.glob(HISTORY_DIR + '*.history'):
print(filename)
with open(filename, 'rb') as f:
history[filename] = pickle.load(f)
for key, val in history.items():
print(key.replace(HISTORY_DIR, '').rstrip('.history'), val.keys())
# ## 准确率变化(训练集)
plot_training(history)
# ## 损失值变化(训练集)
plot_training(history, metric='loss', title='Model Loss', loc='upper right')
# ## 准确率变化(测试集)
plot_training(history, metric='val_acc', title='Model Accuracy (val)')
# ## 损失值变化(测试集)
plot_training(history, metric='val_loss', title='Model Loss (val)', loc='upper right')
|
notebook-examples/chapter-6/4_trainning_analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/chemicoPy/MACD-RSI-STOCHASTIC-strategy/blob/ccxt/MACD_RSI_STOCHASTIC_strategy_(ccxt).ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="qVUr_Q6v_926" outputId="69d39ae7-09fe-45b0-e393-30f25db286b3"
# !pip install ccxt
# !pip install pandas_ta
# !pip install schedule
# + id="3AQ-rbQiAcH4"
import ccxt
import pandas as pd
import numpy as np
from numpy import *
import pandas as pd
from pandas import DataFrame, Series
import matplotlib.pyplot as plt
from datetime import datetime, timedelta, date
from IPython.display import clear_output
import schedule
# + id="Hf1_spBMAcOR"
API_KEY = ""
API_SECRET = ""
# + id="XhNqM4i7AcYC"
# + colab={"base_uri": "https://localhost:8080/", "height": 72} id="WGXhfED0Acv6" outputId="908eabe1-675d-4086-86a7-695eb0899318"
""" Examples_of_coins_are = ["BNBUSDT", "ETHUSDT", "ADAUSDT", "DOTUSDT", "BTCUSDT", "FTMUSDT", "DOGEUSDT", "LINKUSDT", "SUSHIUSDT", "SOLUSDT",
"IOSTUSDT", "LTCUSDT", "XRPUSDT", "SXPUSDT", "MATICUSDT", "BCHUSDT"] """
coin = str(input("Enter coin name: ").upper())
interval = str(input("Enter time interval name e.g 5m for 5-minute: ").lower())
""" For example:
coin = 'BTCUSDT'
interval = '1m' """
# + colab={"base_uri": "https://localhost:8080/"} id="rR1vwjbLAc0s" outputId="c3f8e065-4ca7-472b-decf-3005510858ae"
""" Code line to see all columns of that data grabbed """
exchangepub = ccxt.binanceus() # You can choose/enter the ID you prefer here e.g binance, coinbase, coinmate, coinmetro e.t.c
exchange_id = 'binanceus'
exchange_class = getattr(ccxt, exchange_id)
exchange = exchange_class({
'apiKey': API_KEY,
'secret': API_SECRET,
'timeout': 3333,
'enableRateLimit': True,
})
index = coin.find('USDT')
symbol = coin[:index] + '/' + coin[index:]
instrument = exchange.fetchTickers(str(symbol))
print((instrument))
# + id="CZrGdDv3BRlH"
# + colab={"base_uri": "https://localhost:8080/"} id="Crc7ZdoXBRye" outputId="a1917565-3b90-4a97-9894-a3377a293b07"
import ccxt
exchangepub = ccxt.binanceus() # You can choose/enter the ID you prefer here e.g binanceus, coinbase, coinmate, coinmetro e.t.c
exchange_id = 'binanceus'
exchange_class = getattr(ccxt, exchange_id)
exchange = exchange_class({
'apiKey': API_KEY,
'secret': API_SECRET,
'timeout': 3333,
'enableRateLimit': True,
})
def orderSignal():
markets = exchange.fetch_ohlcv(coin, timeframe=interval, limit = 1000)
df = pd.DataFrame(markets, columns=["time", "open", "high", "low", "close", "volume"])
df.time = pd.to_datetime(df.time, unit='ms')
df.rename(columns = {'time':'dateTime'}, inplace = True)
df.set_index('dateTime', inplace=True)
df['open'] = pd.to_numeric(df['open'], errors='coerce')
df['high'] = pd.to_numeric(df['high'], errors='coerce')
df['low'] = pd.to_numeric(df['low'], errors='coerce')
df['close'] = pd.to_numeric(df['close'], errors='coerce')
df['volume'] = pd.to_numeric(df['volume'], errors='coerce')
return df
df = orderSignal()
print(df.tail())
print("")
print("Size of data:", len(df))
# + id="51mAp_4tBXwM"
# + [markdown] id="3g2XLN0_OQZg"
# # MACD - Moving Average Convergence Divergence
# + colab={"base_uri": "https://localhost:8080/", "height": 237} id="nl8xc4ygBX2J" outputId="11315d65-a306-41f2-acb9-3973ee906377"
def get_macd(price, slow, fast, smooth):
exp1 = price.ewm(span = fast, adjust = False).mean()
exp2 = price.ewm(span = slow, adjust = False).mean()
macd = pd.DataFrame(exp1 - exp2).rename(columns = {'close':'macd'})
signal = pd.DataFrame(macd.ewm(span = smooth, adjust = False).mean()).rename(columns = {'macd':'signal'})
hist = pd.DataFrame(macd['macd'] - signal['signal']).rename(columns = {0:'hist'})
frames = [macd, signal, hist]
df = pd.concat(frames, join = 'inner', axis = 1)
return df
macd = get_macd(df['close'], 26, 12, 9)
df = pd.concat([df, macd], join = 'inner', axis = 1)
df.tail()
# + id="st6ZwsLfLG_P" colab={"base_uri": "https://localhost:8080/", "height": 265} outputId="59f8aa39-4bbf-4078-edb5-bf3b75806eea"
def plot_macd(prices, macd, signal, hist):
ax1 = plt.subplot2grid((8,1), (0,0), rowspan = 5, colspan = 1)
ax2 = plt.subplot2grid((8,1), (5,0), rowspan = 3, colspan = 1)
ax1.plot(prices)
ax2.plot(macd, color = 'grey', linewidth = 1.5, label = 'MACD')
ax2.plot(signal, color = 'skyblue', linewidth = 1.5, label = 'SIGNAL')
for i in range(len(prices)):
if str(hist[i])[0] == '-':
ax2.bar(prices.index[i], hist[i], color = '#ef5350')
else:
ax2.bar(prices.index[i], hist[i], color = '#26a69a')
plt.legend(loc = 'lower right')
plot_macd(df['close'], df['macd'], df['signal'], df['hist'])
# + id="78_F8btoLHL2"
def implement_macd_strategy(prices, data):
buy_price = []
sell_price = []
macd_signal = []
signal = 0
for i in range(len(data)):
if data['macd'][i] > data['signal'][i]:
if signal != 1:
buy_price.append(prices[i])
sell_price.append(np.nan)
signal = 1
macd_signal.append(signal)
else:
buy_price.append(np.nan)
sell_price.append(np.nan)
macd_signal.append(0)
elif data['macd'][i] < data['signal'][i]:
if signal != -1:
buy_price.append(np.nan)
sell_price.append(prices[i])
signal = -1
macd_signal.append(signal)
else:
buy_price.append(np.nan)
sell_price.append(np.nan)
macd_signal.append(0)
else:
buy_price.append(np.nan)
sell_price.append(np.nan)
macd_signal.append(0)
return buy_price, sell_price, macd_signal
buy_price, sell_price, macd_signal = implement_macd_strategy(df['close'], df)
# + id="hFIC2gt6LHVv" colab={"base_uri": "https://localhost:8080/", "height": 281} outputId="ba13201c-b240-4937-b817-599ea6c4dc34"
ax1 = plt.subplot2grid((8,1), (0,0), rowspan = 5, colspan = 1)
ax2 = plt.subplot2grid((8,1), (5,0), rowspan = 3, colspan = 1)
ax1.plot(df['close'], color = 'skyblue', linewidth = 2, label = str(coin))
ax1.plot(df.index, buy_price, marker = '^', color = 'green', markersize = 10, label = 'BUY SIGNAL', linewidth = 0)
ax1.plot(df.index, sell_price, marker = 'v', color = 'r', markersize = 10, label = 'SELL SIGNAL', linewidth = 0)
ax1.legend()
ax1.set_title(str(coin+" MACD SIGNALS"))
ax2.plot(df['macd'], color = 'grey', linewidth = 1.5, label = 'MACD')
ax2.plot(df['signal'], color = 'skyblue', linewidth = 1.5, label = 'SIGNAL')
for i in range(len(df)):
if str(df['hist'][i])[0] == '-':
ax2.bar(df.index[i], df['hist'][i], color = '#ef5350')
else:
ax2.bar(df.index[i], df['hist'][i], color = '#26a69a')
plt.legend(loc = 'lower right')
plt.show()
# + id="Dwh-cZ-ndEJa"
macd_signal
# + id="0alTtxiZLHu2" colab={"base_uri": "https://localhost:8080/", "height": 455} outputId="87fdf400-1b86-4664-e94a-ba2b1aa6e793"
position = []
for i in range(len(macd_signal)):
if macd_signal[i] > 1:
position.append(0)
else:
position.append(1)
for i in range(len(df['close'])):
if macd_signal[i] == 1:
position[i] = 1
elif macd_signal[i] == -1:
position[i] = 0
else:
position[i] = position[i-1]
macd = df['macd']
signal = df['signal']
close_price = df['close']
macd_signal = pd.DataFrame(macd_signal).rename(columns = {0:'macd_signal'}).set_index(df.index)
position = pd.DataFrame(position).rename(columns = {0:'macd_position'}).set_index(df.index)
frames = [close_price, macd, signal, macd_signal, position]
strategy = pd.concat(frames, join = 'inner', axis = 1)
strategy
# + id="mpAma70eVadY"
# + [markdown] id="oo_3nGNyd6KA"
# # RSI - Relative Strength Index
# + id="ZxXDeqc_VakC"
def get_rsi(close, lookback):
ret = close.diff()
up = []
down = []
for i in range(len(ret)):
if ret[i] < 0:
up.append(0)
down.append(ret[i])
else:
up.append(ret[i])
down.append(0)
up_series = pd.Series(up)
down_series = pd.Series(down).abs()
up_ewm = up_series.ewm(com = lookback - 1, adjust = False).mean()
down_ewm = down_series.ewm(com = lookback - 1, adjust = False).mean()
rs = up_ewm/down_ewm
rsi = 100 - (100 / (1 + rs))
rsi_df = pd.DataFrame(rsi).rename(columns = {0:'rsi'}).set_index(close.index)
rsi_df = rsi_df.dropna()
return rsi_df[3:]
df['rsi_14'] = get_rsi(df['close'], 14)
df = df.dropna()
# + colab={"base_uri": "https://localhost:8080/", "height": 455} id="JegujVEreTom" outputId="fe54138b-048a-44ca-f555-2dc028f88b07"
df
# + id="wEuLm6oyeV4V"
# + colab={"base_uri": "https://localhost:8080/", "height": 259} id="am0tiX6xeaC9" outputId="90a73df1-f703-4897-dc1a-ec3e88b45336"
ax3 = plt.subplot2grid((10,1), (0,0), rowspan = 4, colspan = 1)
ax4 = plt.subplot2grid((10,1), (5,0), rowspan = 4, colspan = 1)
ax3.plot(df['close'], linewidth = 2.5)
ax3.set_title(str(coin+" CLOSE PRICE"))
ax4.plot(df['rsi_14'], color = 'orange', linewidth = 2.5)
ax4.axhline(30, linestyle = '--', linewidth = 1.5, color = 'grey')
ax4.axhline(70, linestyle = '--', linewidth = 1.5, color = 'grey')
ax4.set_title(str(coin+" RELATIVE STRENGTH INDEX"))
plt.show()
# + id="dm22GLBseTt9"
def implement_rsi_strategy(prices, rsi):
buy_price = []
sell_price = []
rsi_signal = []
signal = 0
for i in range(len(rsi)):
if rsi[i-1] > 30 and rsi[i] < 30:
if signal != 1:
buy_price.append(prices[i])
sell_price.append(np.nan)
signal = 1
rsi_signal.append(signal)
else:
buy_price.append(np.nan)
sell_price.append(np.nan)
rsi_signal.append(0)
elif rsi[i-1] < 70 and rsi[i] > 70:
if signal != -1:
buy_price.append(np.nan)
sell_price.append(prices[i])
signal = -1
rsi_signal.append(signal)
else:
buy_price.append(np.nan)
sell_price.append(np.nan)
rsi_signal.append(0)
else:
buy_price.append(np.nan)
sell_price.append(np.nan)
rsi_signal.append(0)
return buy_price, sell_price, rsi_signal
rsi_buy_price, rsi_sell_price, rsi_signal = implement_rsi_strategy(df['close'], df['rsi_14'])
# + colab={"base_uri": "https://localhost:8080/", "height": 455} id="RvVmjpCyiVwh" outputId="fbfabcf8-2ffe-4af7-ed8c-39bf79eb4bd3"
df
# + colab={"base_uri": "https://localhost:8080/"} id="ytyxBKJak5jN" outputId="4874c3e3-2f8e-4da0-bddb-95aa18a67ab6"
print(len(df.index))
# + colab={"base_uri": "https://localhost:8080/", "height": 259} id="fDmir9h8Vapz" outputId="efc2f78a-a0a8-4c3c-e8bd-6ebf3f6c11bc"
ax5 = plt.subplot2grid((10,1), (0,0), rowspan = 4, colspan = 1)
ax6 = plt.subplot2grid((10,1), (5,0), rowspan = 4, colspan = 1)
ax5.plot(df['close'], linewidth = 2.5, color = 'skyblue', label = 'RSI')
ax5.plot(df.index, rsi_buy_price, marker = '^', markersize = 10, color = 'green', label = 'BUY SIGNAL')
ax5.plot(df.index, rsi_sell_price, marker = 'v', markersize = 10, color = 'r', label = 'SELL SIGNAL')
ax5.set_title(str(coin+" RSI TRADE SIGNALS"))
ax6.plot(df['rsi_14'], color = 'orange', linewidth = 2.5)
ax6.axhline(30, linestyle = '--', linewidth = 1.5, color = 'grey')
ax6.axhline(70, linestyle = '--', linewidth = 1.5, color = 'grey')
plt.show()
# + id="v5OjDPlpmmwQ"
# + colab={"base_uri": "https://localhost:8080/", "height": 237} id="r-Q8XKTOg9--" outputId="bd7b3a92-0eea-4fed-93fb-f83542f5388b"
position = []
for i in range(len(rsi_signal)):
if rsi_signal[i] > 1:
position.append(0)
else:
position.append(1)
for i in range(len(df['close'])):
if rsi_signal[i] == 1:
position[i] = 1
elif rsi_signal[i] == -1:
position[i] = 0
else:
position[i] = position[i-1]
rsi = df['rsi_14']
close_price = df['close']
rsi_signal = pd.DataFrame(rsi_signal).rename(columns = {0:'rsi_signal'}).set_index(df.index)
position = pd.DataFrame(position).rename(columns = {0:'rsi_position'}).set_index(df.index)
frames = [close_price, rsi, rsi_signal, position]
strategy = pd.concat(frames, join = 'inner', axis = 1)
strategy.head()
# + id="fG_RMhhVg-HC"
# + [markdown] id="fk3CFuramqlW"
# # STOCHASTIC OSCILLATOR
# + colab={"base_uri": "https://localhost:8080/", "height": 281} id="bIXYspceg-NE" outputId="30368d8b-64c3-409a-e79b-e4c1f540ab43"
stoch_data = df.copy()
stoch_data['high'] = stoch_data['high'].rolling(14).max()
stoch_data['low'] = stoch_data['low'].rolling(14).min()
stoch_data['%k'] = (stoch_data["close"] - stoch_data['low'])*100/(stoch_data['high'] - stoch_data['low'])
stoch_data['%d'] = stoch_data['%k'].rolling(3).mean()
def plot_stoch(price, k, d):
ax7 = plt.subplot2grid((9, 1), (0,0), rowspan = 5, colspan = 1)
ax8 = plt.subplot2grid((9, 1), (6,0), rowspan = 3, colspan = 1)
ax7.plot(price)
ax7.set_title(f'{coin} STOCK PRICE')
ax8.plot(k, color = 'deepskyblue', linewidth = 1.5, label = '%K')
ax8.plot(d, color = 'orange', linewidth = 1.5, label = '%D')
ax8.axhline(80, color = 'black', linewidth = 1, linestyle = '--')
ax8.axhline(20, color = 'black', linewidth = 1, linestyle = '--')
ax8.set_title(f'{coin} STOCH')
ax8.legend()
plt.show()
plot_stoch(df['close'], stoch_data['%k'], stoch_data['%d'])
# + id="pvS43JWSmpTD"
def implement_stoch_strategy(prices, k, d):
buy_price = []
sell_price = []
stoch_signal = []
signal = 0
for i in range(len(prices)):
if k[i] < 20 and d[i] < 20 and k[i] < d[i]:
if signal != 1:
buy_price.append(prices[i])
sell_price.append(np.nan)
signal = 1
stoch_signal.append(signal)
else:
buy_price.append(np.nan)
sell_price.append(np.nan)
stoch_signal.append(0)
elif k[i] > 80 and d[i] > 80 and k[i] > d[i]:
if signal != -1:
buy_price.append(np.nan)
sell_price.append(prices[i])
signal = -1
stoch_signal.append(signal)
else:
buy_price.append(np.nan)
sell_price.append(np.nan)
stoch_signal.append(0)
else:
buy_price.append(np.nan)
sell_price.append(np.nan)
stoch_signal.append(0)
return buy_price, sell_price, stoch_signal
stoch_buy_price, stoch_sell_price, stoch_signal = implement_stoch_strategy(stoch_data['close'], stoch_data['%k'], stoch_data['%d'])
# + colab={"base_uri": "https://localhost:8080/", "height": 237} id="f0XMCaCFmpbs" outputId="f35eb71f-46d0-43ba-e68c-68872f547116"
position = []
for i in range(len(stoch_signal)):
if stoch_signal[i] > 1:
position.append(0)
else:
position.append(1)
for i in range(len(stoch_data['close'])):
if stoch_signal[i] == 1:
position[i] = 1
elif stoch_signal[i] == -1:
position[i] = 0
else:
position[i] = position[i-1]
k = stoch_data['%k']
d = stoch_data['%d']
close_price = stoch_data['close']
stoch_signal = pd.DataFrame(stoch_signal).rename(columns = {0:'stoch_signal'}).set_index(stoch_data.index)
position = pd.DataFrame(position).rename(columns = {0:'stoch_position'}).set_index(stoch_data.index)
frames = [close_price, k, d, stoch_signal, position]
strategy = pd.concat(frames, join = 'inner', axis = 1)
strategy.tail()
# + id="NioJS6I2mphX"
# + id="Jl0Aaisptg7u"
# + id="ZPr8gPEwtiOr"
# + id="SswtD3D6g-Tl"
|
MACD_RSI_STOCHASTIC_strategy_(ccxt).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] nbgrader={}
# # Integration Exercise 2
# + [markdown] nbgrader={}
# ## Imports
# + nbgrader={}
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
from scipy import integrate
# + [markdown] nbgrader={}
# ## Indefinite integrals
# + [markdown] nbgrader={}
# Here is a [table of definite integrals](http://en.wikipedia.org/wiki/List_of_definite_integrals). Many of these integrals has a number of parameters $a$, $b$, etc.
#
# Find five of these integrals and perform the following steps:
#
# 1. Typeset the integral using LateX in a Markdown cell.
# 1. Define an `integrand` function that computes the value of the integrand.
# 2. Define an `integral_approx` funciton that uses `scipy.integrate.quad` to peform the integral.
# 3. Define an `integral_exact` function that computes the exact value of the integral.
# 4. Call and print the return value of `integral_approx` and `integral_exact` for one set of parameters.
#
# Here is an example to show what your solutions should look like:
# + [markdown] nbgrader={}
# ### Example
# + [markdown] nbgrader={}
# Here is the integral I am performing:
#
# $$ I = \int_0^\infty \frac{dx}{x^2 + a^2} = \frac{\pi}{2a} $$
# + nbgrader={}
def integrand(x, a):
return 1.0/(x**2 + a**2)
def integral_approx(a):
# Use the args keyword argument to feed extra arguments to your integrand
I, e = integrate.quad(integrand, 0, np.inf, args=(a,))
return I
def integral_exact(a):
return 0.5*np.pi/a
print("Numerical: ", integral_approx(1.0))
print("Exact : ", integral_exact(1.0))
# + nbgrader={}
assert True # leave this cell to grade the above integral
# + [markdown] nbgrader={}
# ### Integral 1
# + [markdown] deletable=false nbgrader={"checksum": "e034fc7ac9c38bbb9c7c87db4b6c8e4e", "grade": true, "grade_id": "integrationex03a", "points": 1, "solution": true}
# $$ I_1 = \int_{0}^{\infty} \frac{\sin ^{2}px}{x^{2}}\ dx=\frac{\pi p}{2} $$
#
# + deletable=false nbgrader={"checksum": "6cff4e8e53b15273846c3aecaea84a3d", "solution": true}
def integrand1(x, p):
return np.sin(p*x)**2/(x**2)
def integral_approx1(p):
# Use the args keyword argument to feed extra arguments to your integrand
I, e = integrate.quad(integrand1, 0, np.inf, args=(p,))
return I
def integral_exact1(p):
return p*np.pi/2
print("Numerical: ", integral_approx1(1.0))
print("Exact : ", integral_exact1(1.0))
# + deletable=false nbgrader={"checksum": "b998cb1faa45ae86f0728d51dfa0e45c", "grade": true, "grade_id": "integrationex03b", "points": 1}
assert True # leave this cell to grade the above integral
# + [markdown] nbgrader={}
# ### Integral 2
# + [markdown] deletable=false nbgrader={"checksum": "c3191d99083f6d7cf804f95876e8a624", "grade": true, "grade_id": "integrationex03c", "points": 1, "solution": true}
# $$ I_2 = \int_0^\infty \frac {x}{e^{x}-1}\ dx= \frac {\pi^2}{6} $$
# + deletable=false nbgrader={"checksum": "6cff4e8e53b15273846c3aecaea84a3d", "solution": true}
def integrand2(x):
return x/(np.exp(x)-1)
def integral_approx2():
# Use the args keyword argument to feed extra arguments to your integrand
I, e = integrate.quad(integrand2, 0, np.inf)
return I
def integral_exact2():
return np.pi**2/6
print("Numerical: ", integral_approx2())
print("Exact : ", integral_exact2())
# + deletable=false nbgrader={"checksum": "4e20de120f0c45ba666f10ba9a6c82d8", "grade": true, "grade_id": "integrationex03d", "points": 1}
assert True # leave this cell to grade the above integral
# + [markdown] nbgrader={}
# ### Integral 3
# + [markdown] deletable=false nbgrader={"checksum": "c65f5242f7fa5525523b89899f6ca251", "grade": true, "grade_id": "integrationex03e", "points": 1, "solution": true}
# $$ I_3 = \int_0^a \frac{dx}{\sqrt{a^{2}-x^{2}}}=\frac{\pi }{2} $$
# + deletable=false nbgrader={"checksum": "6cff4e8e53b15273846c3aecaea84a3d", "solution": true}
def integrand3(x, a):
return 1.0/((a**2-x**2 )**(.5))
def integral_approx3(a):
# Use the args keyword argument to feed extra arguments to your integrand
I, e = integrate.quad(integrand3, 0, a, args=(a,))
return I
def integral_exact3(a):
return np.pi/2
print("Numerical: ", integral_approx3(17))
print("Exact : ", integral_exact3(17))
# + deletable=false nbgrader={"checksum": "8c60d256fe8559e423cf8946ae70ba8d", "grade": true, "grade_id": "integrationex03f", "points": 1}
assert True # leave this cell to grade the above integral
# + [markdown] nbgrader={}
# ### Integral 4
# + [markdown] deletable=false nbgrader={"checksum": "3a5d3b2070c78b64152c96681e8e6585", "grade": true, "grade_id": "integrationex03g", "points": 1, "solution": true}
# $$ I_4 =\int_0^\infty \frac{x \sin mx}{x^2+a^2}\ dx=\frac{\pi}{2}e^{-ma} $$
# + deletable=false nbgrader={"checksum": "6cff4e8e53b15273846c3aecaea84a3d", "solution": true}
def integrand4(x, m, a):
return (x*np.sin(m*x))/(x**2+a**2)
def integral_approx4(m, a):
# Use the args keyword argument to feed extra arguments to your integrand
I, e = integrate.quad(integrand4, 0, np.inf, args=(m,a,))
return I
def integral_exact4(m, a):
return (np.pi/2)*np.exp(-1*m*a)
print("Numerical: ", integral_approx4(.001,.001))
print("Exact : ", integral_exact4(.001,.001))
# + deletable=false nbgrader={"checksum": "88acfb75979c6551c8b3af758cd86acc", "grade": true, "grade_id": "integrationex03h", "points": 1}
assert True # leave this cell to grade the above integral
# + [markdown] nbgrader={}
# ### Integral 5
# + [markdown] deletable=false nbgrader={"checksum": "9dbb9f1159b3c089e60dd167d973cc59", "grade": true, "grade_id": "integrationex03i", "points": 1, "solution": true}
# $$ I_5 = \int_{-\infty}^\infty e^{-x^2}\,dx=\sqrt{\pi} $$
# + deletable=false nbgrader={"checksum": "6cff4e8e53b15273846c3aecaea84a3d", "solution": true}
def integrand5(x):
return (np.exp(-1*(x**2)))
def integral_approx5():
# Use the args keyword argument to feed extra arguments to your integrand
I, e = integrate.quad(integrand5, -1*np.inf, np.inf)
return I
def integral_exact5():
return np.pi**(1/2)
print("Numerical: ", integral_approx5())
print("Exact : ", integral_exact5())
# + deletable=false nbgrader={"checksum": "34f6cf778698f4b90fdadc09c2a0f120", "grade": true, "grade_id": "integrationex03j", "points": 1}
assert True # leave this cell to grade the above integral
|
assignments/assignment09/IntegrationEx02.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# formats: jl:light,ipynb
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.4.2
# language: julia
# name: julia-1.4
# ---
# +
"""
Tutorial: A reference implementation of configuration interactions singles.
"""
__authors__ = ["<NAME>", "<NAME>", "<NAME>"]
__credits__ = ["<NAME>", "<NAME>", "<NAME>", "<NAME>"]
__copyright_amp__ = "(c) 2014-2020, The Psi4Julia Developers"
__license__ = "BSD-3-Clause"
__date__ = "2020-08-03"
# -
# # Configuration Interaction Singles (CIS)
# ## I. Theoretical Overview
# In this tutorial, we will implement the configuration interaction singles method in the spin orbital notation. The groundwork for working in the spin orbital notation has been laid out in "Introduction to the Spin Orbital Formulation of Post-HF methods" [tutorial](../08_CEPA0_and_CCD/8a_Intro_to_spin_orbital_postHF.ipynb). It is highly recommended to work through that introduction before starting this tutorial.
# ### Configuration Interaction (CI)
#
# The configuration interaction wavefunction is constructed as a linear combination of the reference determinants and all singly, doubly, ... n-tuple excited determinants where n is the number of electrons in a given system:
#
# \begin{equation}
# \Psi_\mathrm{CI} = (1 + \hat{C_1} + \hat{C_2} + ...\hat{C_n)}\Phi
# \end{equation}
#
# Here, $\hat{C_n}$ is the n configuration excitation operator.
#
# In Full CI, all possible excitations are included in the wavefunction expansion. In truncated CI methods, only a subset of excitations are included.
# ## CIS
#
# In CIS, only single excitations from the occupied (indices i,j,k...) to the virtual (indices a,b,c...) orbitals are included. As a result, CIS gives transition energies to an excited state.
#
# Assuming we are using canonical Hartree-Fock spin orbitals($\{\mathrm{\psi_p}\}$) with orbital energies $\{\epsilon_p\}$, we can build a shifted CIS Hamiltonian matrix:
#
# \begin{equation}
# \tilde{\textbf{H}} = \textbf{H} - E_0 \textbf{I} = [\langle \Phi_P | \hat{H_e} - E_0|\Phi_Q \rangle],\,
# \Phi_P \in {\Phi_i^a}
# \end{equation}
#
# where $E_0$ is the ground state Hartree-Fock state energy given by $\langle \Phi | \hat{H_e}|\Phi \rangle$.
#
# The matrix elements of this shifted CIS Hamiltonian matrix can be evaluated using Slater's rules to give:
#
# \begin{equation}
# \langle \Phi_i^a | \hat{H_e} - E_0|\Phi_j^b \rangle = (\epsilon_a - \epsilon_i)\delta_{ij} \delta_{ab}
# + \langle aj || ib \rangle
# \end{equation}
#
# This then becomes a standard eigenvalue equation from which we can solve for the excitation energies and the wavefunction expansion coefficients:
#
# \begin{equation}
# \tilde{\textbf{H}} \textbf{c}_K = \Delta E_K\textbf{c}_K, \,\Delta E_K = E_K - E_0
# \end{equation}
#
#
# ## II. Implementation
# As with previous tutorials, let's begin by importing Psi4, NumPy, and TensorOperations and setting memory and output file options.
# +
# ==> Import Psi4, NumPy, & TensorOperations <==
using PyCall: pyimport
psi4 = pyimport("psi4")
np = pyimport("numpy") # used only to cast to Psi4 arrays
using TensorOperations: @tensor
# ==> Set Basic Psi4 Options <==
# Memory specifications
psi4.set_memory(Int(2e9))
numpy_memory = 2
# Output options
psi4.core.set_output_file("output.dat", false)
# -
# We now define the molecule and set Psi4 options:
# +
mol = psi4.geometry("""
0 1
O
H 1 1.1
H 1 1.1 2 104
symmetry c1
""")
psi4.set_options(Dict("basis" => "sto-3g",
"scf_type" => "pk",
"reference" => "rhf",
"mp2_type" => "conv",
"e_convergence" => 1e-8,
"d_convergence" => 1e-8))
# -
# We use Psi4 to compute the RHF energy and wavefunction and store them in variables `scf_e` and `scf_wfn`. We also check the memory requirements for computation:
# +
# Get the SCF wavefunction & energies
scf_e, scf_wfn = psi4.energy("scf", return_wfn=true)
# Check memory requirements
nmo = scf_wfn.nmo()
I_size = nmo^4 * 8e-9
println("\nSize of the ERI tensor will be $I_size GB.")
memory_footprint = I_size * 1.5
if I_size > numpy_memory
psi4.core.clean()
throw(OutOfMemoryError("Estimated memory utilization ($memory_footprint GB) exceeds " *
"allotted memory limit of $numpy_memory GB."))
end
# -
# We first obtain orbital information from our wavefunction. We also create an instance of MintsHelper to help build our molecular integrals:
# +
# Create instance of MintsHelper class
mints = psi4.core.MintsHelper(scf_wfn.basisset())
# Get basis and orbital information
nbf = mints.nbf() # Number of basis functions
nalpha = scf_wfn.nalpha() # Number of alpha electrons
nbeta = scf_wfn.nbeta() # Number of beta electrons
nocc = nalpha + nbeta # Total number of electrons
nso = 2nbf # Total number of spin orbitals
nvirt = nso - nocc # Number of virtual orbitals
# -
# We now build our 2-electron integral, a 4D tensor, in the spin orbital formulation. We also convert it into physicist's notation and antisymmetrize for easier manipulation of the tensor later on.
# +
"""
Spin blocks 2-electron integrals
Using np.kron, we project I and I tranpose into the space of the 2x2 ide
The result is our 2-electron integral tensor in spin orbital notation
"""
function spin_block_tei(I)
identity = [ 1.0 0.0; 0.0 1.0]
I = np.kron(identity, I)
np.kron(identity, permutedims(I, reverse(1:4)))
end
I = np.asarray(mints.ao_eri())
I_spinblock = spin_block_tei(I)
# Convert chemist's notation to physicist's notation, and antisymmetrize
# (pq | rs) ---> <pr | qs>
# <pr||qs> = <pr | qs> - <pr | sq>
gao = permutedims(I_spinblock, (1, 3, 2, 4)) - permutedims(I_spinblock, (1, 3, 4, 2));
# -
# We get the orbital energies from alpha and beta electrons and append them together. We spin-block the coefficients obtained from the reference wavefunction and convert them into NumPy arrays. There is a set corresponding to coefficients from alpha electrons and a set of coefficients from beta electrons. We then sort them according to the order of the orbital energies using argsort():
# +
# Get orbital energies, cast into NumPy array, and extend eigenvalues
eps_a = np.asarray(scf_wfn.epsilon_a())
eps_b = np.asarray(scf_wfn.epsilon_b())
eps = vcat(eps_a, eps_b)
# Get coefficients, block, and sort
Ca = np.asarray(scf_wfn.Ca())
Cb = np.asarray(scf_wfn.Cb())
C = [Ca zero(Ca); zero(Cb) Cb]; # direct sum
# Sort the columns of C according to the order of orbital energies
C = C[:, sortperm(eps)]
# Sort orbital energies
sort!(eps);
# -
# We now transform the 2-electron integral from the AO basis into the MO basis using the coefficients:
#
# Transform gao, which is the spin-blocked 4d array of physicist's notation,
# antisymmetric two-electron integrals, into the MO basis using MO coefficients
gmo = @tensor begin
gmo[P,Q,R,S] := gao[p,Q,R,S] * C[p,P]
gmo[p,Q,R,S] := gmo[p,q,R,S] * C[q,Q]
gmo[p,q,R,S] := gmo[p,q,r,S] * C[r,R]
gmo[p,q,r,S] := gmo[p,q,r,s] * C[s,S]
end
nothing
# Now that we have our integrals, coefficents, and orbital energies set up in with spin orbitals, we can start our CIS procedure. We first start by initializing the shifted Hamiltonion matrix $\tilde{\textbf{H}}$ (`HCIS`). Let's think about the size of $\tilde{\textbf{H}}$. We need all possible single excitations from the occupied to virtual orbitals. This is given by the number of occupied orbitals times the number of virtual orbitals (`nocc * nvirt`).
#
# The size of our matrix is thus `nocc * nvirt` by `nocc * nvirt`.
# Initialize CIS matrix.
# The dimensions are the number of possible single excitations
HCIS = zeros(nocc * nvirt, nocc * nvirt);
# Next, we want to build all possible excitations from occupied to virtual orbitals. We create two for-loops that will loop over the number of occupied orbitals and number of virtual orbitals, respectively, and store the combination of occupied and virtual indices as a tuple `(i, a)`. We put all tuples in a list called `excitations`.
# Build the possible excitations, collect indices into a list
excitations = []
for i in 1:nocc, a in nocc+1:nso
push!(excitations,(i, a))
end
# Now we can evaluate the matrix elements of the shifted CIS Hamiltonian matrix using the equation given above. For each element, there are several layers of indexing that we have to consider.
# First, there are the indices of the element itself, which gives the position of the element in the matrix. Indices `p` and `q` are used:
#
# `HCIS[p, q]`
#
# Second, there are two sets of excitations from occupied to virtual orbitals corresponding to the bra and ket of each matrix element. For these, we will take advantage of the `excitations` list that we build with the list of all possible excitations. We will use indices `i` and `a` to denote the excitation in the bra (`left_excitation`) and `j` and `b` to denote the excitation in the ket (`right_excitation`).
#
# To manage these indices, we will use the `enumerate` function.
#
# Note that a Kronecker delta $\delta_{pq}$ can be represented as `p == q`.
#
# Form matrix elements of shifted CIS Hamiltonian
for (p, left_excitation) in enumerate(excitations)
i, a = left_excitation
for (q, right_excitation) in enumerate(excitations)
j, b = right_excitation
HCIS[p, q] = (eps[a] - eps[i]) * (i == j) * (a == b) + gmo[a, j, i, b]
end
end
# We now use the composed function `eigen ∘ Hermitian` (for hermitian matrices) to diagonalize the shifted CIS Hamiltonian. This will give us the excitation energies (`ECIS`). These eigenvalues correspond to the CIS total energies for various excited states. The columns of matrix `CCIS` give us the coefficients which describe the relative contribution of each singly excited determinant to the excitation energy.
#
# +
# Diagonalize the shifted CIS Hamiltonian
using LinearAlgebra: eigen, Hermitian
ECIS, CCIS = (eigen ∘ Hermitian)(HCIS) ;
# -
# For a given excitation energy, each coefficent in the linear combination of excitations represents the amount that a particular excitation contributes to the overall excitation energy. The percentage contribution of each coefficient can be calculated by squaring the coefficent and multiplying by 100.
# Percentage contributions for each state vector
percent_contrib = @. round(CCIS^2 * 100);
# In addition to excitation energies, we want to print the excitations that contribute 10% or more to the overall energy, as well as their percent contribution.
#
# Note that `printfmt` allows us to print different sections to the same line without a line break.
# Print detailed information on significant excitations
using Formatting: printfmt
println("CIS:")
for state in eachindex(ECIS)
# Print state, energy
printfmt("State {1:3d} Energy (Eh) {2:10.7f} ", state, ECIS[state])
for (idx, excitation) in enumerate(excitations)
if percent_contrib[idx, state] > 10
i, a = excitation
# Print percentage contribution and the excitation
printfmt("{1:4d}% {2:2d} -> {3:2d} ", percent_contrib[idx, state], i, a)
end
end
printfmt("\n")
end
# ## References
# 1. Background paper:
# >"Toward a systematic molecular orbital theory for excited states"
# [[Foresman:1992:96](http://pubs.acs.org/doi/abs/10.1021/j100180a030)] <NAME>, <NAME>, <NAME>, <NAME>, *J. Phys. Chem.* **96**, 135 (1992).
#
#
# 2. Algorithms from:
# > [[CCQC:CIS](https://github.com/CCQC/summer-program/tree/master/7)] CCQC Summer Program, "CIS" accessed with https://github.com/CCQC/summer-program/tree/master/7.
#
|
Tutorials/09_Configuration_Interaction/9a_cis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:py3_spgh_dev]
# language: python
# name: conda-env-py3_spgh_dev-py
# ---
# ---------------
#
# **If any part of this notebook is used in your research, please cite with the reference found in** **[README.md](https://github.com/pysal/spaghetti#bibtex-citation).**
#
#
# ----------------
#
# ## Network-constrained spatial autocorrelation
# ### Performing and visualizing exploratory spatial data analysis
#
# **Author: <NAME>** **<<EMAIL>>**
#
# **This notebook is an advanced walk-through for:**
#
# 1. Demonstrating spatial autocorrelation with [pysal/esda](https://pysal.org/esda/)
# 2. Calculating [Moran's *I*](https://pysal.org/esda/generated/esda.Moran.html#esda.Moran) on a segmented network
# 3. Visualizing spatial autocorrelation with [pysal/splot](https://splot.readthedocs.io/en/latest/)
# %load_ext watermark
# %watermark
# +
import esda
import libpysal
import matplotlib
import matplotlib_scalebar
from matplotlib_scalebar.scalebar import ScaleBar
import numpy
import spaghetti
import splot
# %matplotlib inline
# %watermark -w
# %watermark -iv
# -
try:
from IPython.display import set_matplotlib_formats
set_matplotlib_formats("retina")
except ImportError:
pass
# ----------------
#
# ### Instantiating a `spaghetti.Network` object and a point pattern
# #### Instantiate the network from a `.shp` file
ntw = spaghetti.Network(in_data=libpysal.examples.get_path("streets.shp"))
ntw
# #### Extract network arcs as a `geopandas.GeoDataFrame`
_, arc_df = spaghetti.element_as_gdf(ntw, vertices=True, arcs=True)
arc_df.head()
# #### Associate the network with a point pattern
pp_name = "crimes"
pp_shp = libpysal.examples.get_path("%s.shp" % pp_name)
ntw.snapobservations(pp_shp, pp_name, attribute=True)
ntw.pointpatterns
# #### Extract the crimes point pattern as a `geopandas.GeoDataFrame`
pp_df = spaghetti.element_as_gdf(ntw, pp_name=pp_name)
pp_df.head()
# --------------------------
#
# ### 1. ESDA — Exploratory Spatial Data Analysis with [pysal/esda](https://esda.readthedocs.io/en/latest/)
#
# **The Moran's *I* test statistic allows for the inference of how clustered (or dispersed) a dataset is while considering both attribute values and spatial relationships. A value of closer to +1 indicates absolute clustering while a value of closer to -1 indicates absolute dispersion. Complete spatial randomness takes the value of 0. See the** [esda documentation](https://pysal.org/esda/) **for in-depth descriptions and tutorials.**
def calc_moran(net, pp_name, w):
"""Calculate a Moran's I statistic based on network arcs."""
# Compute the counts
pointpat = net.pointpatterns[pp_name]
counts = net.count_per_link(pointpat.obs_to_arc, graph=False)
# Build the y vector
arcs = w.neighbors.keys()
y = [counts[a] if a in counts.keys() else 0. for i, a in enumerate(arcs)]
# Moran's I
moran = esda.moran.Moran(y, w, permutations=99)
return moran, y
# #### Moran's *I* using the network representation's *W*
moran_ntwwn, yaxis_ntwwn = calc_moran(ntw, pp_name, ntw.w_network)
moran_ntwwn.I
# #### Moran's *I* using the graph representation's *W*
moran_ntwwg, yaxis_ntwwg = calc_moran(ntw, pp_name, ntw.w_graph)
moran_ntwwg.I
# **Interpretation:**
#
# * **Although both the network and graph representations (**``moran_ntwwn`` **and** ``moran_ntwwg``**, respectively) display minimal postive spatial autocorrelation, a slighly higher value is observed in the graph represention. This is likely due to more direct connectivity in the graph representation; a direct result of eliminating** [degree-2 vertices](https://en.wikipedia.org/wiki/Degree_(graph_theory))**. The Moran's *I* for both the network and graph representations suggest that network arcs/graph edges attributed with associated crime counts are nearly randomly distributed.**
#
# --------------------------------
#
# ### 2. Moran's *I* on a segmented network
# #### Moran's *I* on a network split into 200-meter segments
n200 = ntw.split_arcs(200.0)
n200
moran_n200, yaxis_n200 = calc_moran(n200, pp_name, n200.w_network)
moran_n200.I
# #### Moran's *I* on a network split into 50-meter segments
n50 = ntw.split_arcs(50.0)
n50
moran_n50, yaxis_n50 = calc_moran(n50, pp_name, n50.w_network)
moran_n50.I
# **Interpretation:**
#
# * **Contrary to above, both the 200-meter and 50-meter segmented networks (**``moran_n200`` **and** ``moran_n50``**, respectively) display minimal negative spatial autocorrelation, with slighly lower values being observed in the 200-meter representation. However, similar to above the Moran's *I* for both the these representations suggest that network arcs attributed with associated crime counts are nearly randomly distributed.**
#
# ---------------------------
#
# ### 3. Visualizing ESDA with `splot`
#
# **Here we are demonstrating** [spatial lag](https://pysal.org/esda/notebooks/spatialautocorrelation.html#Attribute-Similarity)**, which refers to attribute similarity. See the** [splot documentation](https://splot.readthedocs.io/en/latest/) **for in-depth descriptions and tutorials.**
from splot.esda import moran_scatterplot, lisa_cluster, plot_moran
# #### Moran scatterplot
#
# **Plotted with equal aspect**
moran_scatterplot(moran_ntwwn, aspect_equal=True);
# **Plotted without equal aspect**
moran_scatterplot(moran_ntwwn, aspect_equal=False);
# **This scatterplot demostrates the attribute values and associated attribute similarities in space (spatial lag) for the network representation's *W* (**``moran_ntwwn``**).**
#
# #### Reference distribution and Moran scatterplot
plot_moran(moran_ntwwn, zstandard=True, figsize=(10,4));
# **This figure incorporates the reference distribution of Moran's *I* values into the above scatterplot of the network representation's *W* (**``moran_ntwwn``**).**
#
# #### Local Moran's *l*
#
# **The demonstrations above considered the dataset as a whole, providing a global measure. The following demostrates the consideration of** [local spatial autocorrelation](https://pysal.org/esda/notebooks/spatialautocorrelation.html#Local-Autocorrelation:-Hot-Spots,-Cold-Spots,-and-Spatial-Outliers)**, providing a measure for each observation. This is best interpreted visually, here with another scatterplot colored to indicate** [relationship type](https://nbviewer.jupyter.org/github/pysal/splot/blob/master/notebooks/esda_morans_viz.ipynb#Visualizing-Local-Autocorrelation-with-splot---Hot-Spots,-Cold-Spots-and-Spatial-Outliers)**.**
#
# **Plotted with equal aspect**
p = 0.05
moran_loc_ntwwn = esda.moran.Moran_Local(yaxis_ntwwn, ntw.w_network)
fig, ax = moran_scatterplot(moran_loc_ntwwn, p=p, aspect_equal=True)
ax.set(xlabel="Crimes", ylabel="Spatial Lag of Crimes");
# **Plotted without equal aspect**
fig, ax = moran_scatterplot(moran_loc_ntwwn, aspect_equal=False, p=p)
ax.set(xlabel="Crimes", ylabel="Spatial Lag of Crimes");
# **Interpretation:**
#
# * **The majority of observations (network arcs) display no significant local spatial autocorrelation (shown in gray).**
#
# #### Plotting Local Indicators of Spatial Autocorrelation ([LISA](https://onlinelibrary.wiley.com/doi/epdf/10.1111/j.1538-4632.1995.tb00338.x))
f, ax = lisa_cluster(moran_loc_ntwwn, arc_df, p=p, figsize=(12,12), lw=5, zorder=0)
pp_df.plot(ax=ax, zorder=1, alpha=.25, color="g", markersize=30)
suptitle = "LISA for Crime-weighted Networks Arcs"
matplotlib.pyplot.suptitle(suptitle, fontsize=20, x=.51, y=.93)
subtitle = "Crimes ($n=%s$) are represented as semi-opaque green circles"
matplotlib.pyplot.title(subtitle % pp_df.shape[0], fontsize=15);
# -----------------
|
notebooks/network-spatial-autocorrelation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pickle as p
import numpy as np
import tensorflow as tf
import random
import lab1_utils as utils
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# # Intro to TensorFlow
#
# ## What is a Computation Graph?
#
# Everything in TensorFlow comes down to building a computation graph. What is a computation graph? Its just a series of math operations that occur in some order. Here is an example of a simple computation graph:
#
# <img src="files/computation-graph.png">
#
# This graph takes 2 inputs, (a, b) and computes an output (e). Each node in the graph is an operation that takes some input, does some computation, and passes its output to another node.
#
# We could make this computation graph in TensorFlow in the following way:
a = tf.placeholder(tf.float32)
b = tf.placeholder(tf.float32)
c = tf.add(a, b)
d = tf.sub(b, 1)
e = tf.mul(c, d)
# Tensorflow uses tf.placeholder to handle inputs to the model. This is like making a reservation at a restaurant. The restaurant reserves a spot for 5 people, but you are free to fill those seats with any set of friends you want to. tf.placeholder lets you specify that some input will be coming in, of some shape and some type. Only when you run the computation graph do you actually provide the values of this input data. You would run this simple computation graph like this:
with tf.Session() as session:
a_data, b_data = 3.0, 6.0
feed_dict = {a: a_data, b: b_data}
output = session.run([e], feed_dict=feed_dict)
print(output) # 45.0
# We use feed_dict to pass in the actual input data into the graph. We use session.run to get the output from the c operation in the graph. Since e is at the end of the graph, this ends up running the entire graph and returning the number 45 - cool!
# ## Neural Networks in Tensorflow
#
# We can define neural networks in TensorFlow using computation graphs. Here is an example, very simple neural network (just 1 perceptron):
#
# <img src="files/computation-graph-2.png">
#
# This graph takes an input, (x) and computes an output (out). It does it with what we learned in class, `out = sigmoid(W*x+b)`.
#
# We could make this computation graph in TensorFlow in the following way:
# +
n_input_nodes = 2
n_output_nodes = 1
x = tf.placeholder(tf.float32, (None, n_input_nodes))
W = tf.Variable(tf.ones((n_input_nodes, n_output_nodes)), dtype=tf.float32)
b = tf.Variable(tf.zeros(n_output_nodes), dtype=tf.float32)
'''TODO: Define the operation for z (use tf.matmul to multiply W and x).'''
z = #todo
'''TODO: Define the operation for out (use tf.sigmoid).'''
out = #todo
# -
# To run this graph, we again use session.run() and feed in our input via feed_dict.
test_input = [[0.5, 0.5]]
with tf.Session() as session:
tf.global_variables_initializer().run(session=session)
feed_dict = {x: test_input}
output = session.run([out], feed_dict=feed_dict)
print(output[0]) # This should output 0.73105. If not, double-check your code above
# We can also set the value of a tf.Variable when we make it. Below is an example where we set the value of tf.Variable ourselves. We've made a classification dataset for you to play around with, and see how the decision boundary changes with the model parameters (weights and bias). Try to get all the datapoints correct (green)!
# +
'''TODO: manually optimize weight_values and bias_value to classify points'''
# Modify weight_values, bias_value in the above code to adjust the decision boundary
# See if you can classify all the points correctly (all markers green)
weight_values = np.array([[-0.1], [0.2]]) # TODO change values and re-run
bias_value = np.array([[0.5]]) #TODO change values and re-run
# A pretty good boundary is made with:
# weight_values = np.array([[0.03], [0.12]])
# bias_value = np.array([[-0.5]])
x = tf.placeholder(tf.float32, (None, 2), name='x')
W = tf.Variable(weight_values, name='W', dtype=tf.float32)
b = tf.Variable(bias_value, name='b', dtype=tf.float32)
z = tf.matmul(x, W) + b
out = tf.sigmoid(z)
data = np.array([[2, 7], [1, 7], [3, 1], [3, 3], [4, 3], [4, 6], [6, 5], [7, 7], [7, 5], [2, 4], [2, 2]])
y = np.array([1, 1, 0, 0, 0, 1, 1, 1, 1, 0, 0])
with tf.Session() as session:
tf.global_variables_initializer().run(session=session)
utils.classify_and_plot(data, y, x, out, session)
# -
# ## Tweet Sentiment Analysis
# Let's move to a real-world task. We're going to be classifying tweets as positive, negative, or neutral. Check out the very negative tweet below:
# <img src="files/tweet-model.jpg" style="width: 500px;">
#
# ## Building the Model
#
# ### Building an MLP
#
# MLP or Multi-layer perceptron is a basic archetecture where where we multiply our representation with some matrix `W` and add some bias `b` and then apply some nonlineanity like `tanh` at each layer. Layers are fully connected to the next. As the network gets deeper, it's expressive power grows exponentially and so they can draw some pretty fancy decision boundaries. In this exercise, you'll build your own MLP, with 2 hidden layers (layers that aren't input or output).
#
# To make training more stable and efficient, we'll actually evaluate 128 tweets at a time, and take gradients with respect to the loss on the 128. We call this idea **training with mini-batches**.
#
# ### Step 1: Representing Words
#
# In this model, we’ll be representing tweets as [bag-of-words](https://en.wikipedia.org/wiki/Bag-of-words_model) (BOW) representations. BOW representations are vectors where each element index represents a different word and its value represents the number of times this word appears in our sentence. This means that each sentence will be represented by a vector that is vocab_size long. Our output labels will be represented as a vector of size n_classes (3). We get this data with some utility functions:
# +
# load data
X, y, index_to_word, sentences = utils.load_sentiment_data_bow()
X_train, y_train, X_test, y_test = utils.split_data(X, y)
vocab_size = X.shape[1]
n_classes = y.shape[1]
print("Tweet:", sentences[5])
print("Label:", y[5])
print("Bag of Words Representation:", X_train[5])
# -
# ### Step 2: Making Placeholders
# So we have our data loaded as numpy arrays. But remember, TensorFlow graphs begin with generic placeholder inputs, not actual data. We feed the actual data in later once the full graph has been defined. We define our placeholders like this:
# +
data_placeholder = tf.placeholder(tf.float32, shape=(None, vocab_size), name='data_placeholder')
'''TODO: Make the labels placeholder.''';
labels_placeholder = #todo
# -
# #### Why Do We Pass in None?
#
# A note about ‘None’ and fluid-sized dimensions:
#
# You may notice that the first dimension of shape of data_placeholder is ‘None’. data_placeholder should have shape (num_tweets, vocab_size). However, we don’t know how many tweets we are going to be passing in at a time, num_tweets is unknown. Its possible that we only want to pass in 1 tweet at a time, or 30, or 1,000. Thankfully, TensorFlow allows us to specify placeholders with fluid-sized dimensions. We can use None to specify some fluid dimension of our shape. When our data eventually gets passed in as a numpy array, TensorFlow can figure out what the value of the fluid-size dimension should be.
#
# ### Step 3: Define Network Parameters
# Let’s now define and initialize our network parameters.
#
# We'll our model parameters using tf.Variable. When you create a tf.Variable you pass a Tensor as its initial value to the Variable() constructor. A Tensor is a term for any N-dimensional matrix. There are a ton of different initial Tensor value functions you can use ([full list](https://www.tensorflow.org/api_docs/python/constant_op/)). All these functions take a list argument that determines their shape. Here we use tf.truncated_normal for our weights, and tf.zeros for our biases. Its important that the shape of these parameters are compatible. We’ll be matrix-multiplying the weights, so the last dimension of the previous weight matrix must equal the first dimension of the next weight matrix. Notice this pattern in the following tensor initialization code. Lastly, notice the size of the tensor for our last weights. We are predicting a vector of size n_classes so our network needs to end with n_classes nodes.
# +
# Define Network Parameters
# Here, we can define how many units will be in each hidden layer.
n_hidden_units_h0 = 512
n_hidden_units_h1 = 256
# Weights going from input to first hidden layer.
# The first value passed into tf.Variable is initialization of the variable.
# We initialize our weights to be sampled from a truncated normal, as this does something
# called symmetry breaking and keeps the neural network from getting stuck at the start.
# Since the weight matrix multiplies the previous layer to get inputs to the next layer,
# its size is prev_layer_size x next_layer_size.
h0_weights = tf.Variable(
tf.truncated_normal([vocab_size, n_hidden_units_h0]),
name='h0_weights')
h0_biases = tf.Variable(tf.zeros([n_hidden_units_h0]),
name='h0_biases')
'''TODO: Set up variables for the weights going into the second hidden layer.
You can check out the tf.Variable API here: https://www.tensorflow.org/api_docs/python/state_ops/variables.
''';
h1_weights = #todo
h1_biases = #todo
# Weights going into the output layer.
out_weights = tf.Variable(
tf.truncated_normal([n_hidden_units_h1, n_classes]),
name='out_weights')
out_biases = tf.Variable(tf.zeros([n_classes]),
name='out_biases')
# -
# ### Step 4: Build Computation Graph
#
# Now let’s define our computation graph.
#
# Our first operation in our graph is a tf.matmul of our data input and our first set of weights. tf.matmul performs a matrix multiplication of two tensors. This is why it is so important that the dimensions of data_placeholder and h0_weights align (dimension 1 of data_placeholder must equal dimension 0 of h0_weights). We then just add the h0_bias variable and perform a nonlinearity transformation, in this case we use tf.nn.relu (ReLU). We do this again for the next hidden layer, and the final output logits.
# +
# Define Computation Graphs
hidden0 = tf.nn.relu(tf.matmul(data_placeholder, h0_weights) + h0_biases)
'''TODO: write the computation to get the output of the second hidden layer.''';
hidden1 = #todo
logits = tf.matmul(hidden1, out_weights) + out_biases
# -
# ### Step 5: Define a Loss Functions
#
# We have defined our network, but we need a way to train it. Training a network comes down to optimizing our network to minimize a loss function, or a measure how good we're doing. Then, we can take the gradient with respect to that performance and move in the right direction.
#
# Since we are doing classification (pos vs neg), a common loss function to use is [cross entropy](https://colah.github.io/posts/2015-09-Visual-Information/):
#
# $$L( \Theta ) = - \sum_i y_i'\log{y_i} $$
#
# where $y$ is our predicted probability distribution and $y'$ is the true distribution. We can access this function in tensorflow with `tf.nn.softmax_cross_entropy_with_logits`.
#
# Note that this loss is 0 if the prediction is correct.
#
# +
'''TODO: Define the loss. Use tf.nn.softmax_cross_entropy_with_logits.'''
loss = #todo
learning_rate = 0.0002
# Define the optimizer operation. This is what will take the derivate of the loss
# with respect to each of our parameters and try to minimize it.
optimizer = tf.train.GradientDescentOptimizer(learning_rate).minimize(loss)
prediction = tf.nn.softmax(logits)
# Compute the accuracy
prediction_is_correct = tf.equal(tf.argmax(logits, 1), tf.argmax(labels_placeholder, 1))
accuracy = tf.reduce_mean(tf.cast(prediction_is_correct, tf.float32))
# -
# #### Quick Conceptual Note:
# Nearly everything we do in TensorFlow is an operation with inputs and outputs. Our loss variable is an operation, that takes the output of the last layer of the net, which takes the output of the 2nd-to-last layer of the net, and so on. Our loss can be traced back to the input as a single function. This is our full computation graph. We pass this to our optimizer which is able to compute the gradient for this computation graph and adjust all the weights in it to minimize the loss.
#
# ### Step 6: Training our Net
# We have our network, our loss function, and our optimizer ready, now we just need to pass in the data to train it. We pass in the data in chunks called mini-batches. We do backpropogation at the end of a batch based on the loss that results from all the examples in the batch. Using batches is just like Stochastic Gradient Descent, except instead of updating parameters after each example, we update them based on the gradient computed after *several* examples.
#
# To measure how well we're doing, we can't just look at how well our model performs on it's training data. It could be just memorizing the training data and perform terribly on data it hasn't seen before. To really measure how it performs in the wild, we need to present it with unseen data, and we can tune our hyper-parameters (like learning rate, num layers etc.) over this first unseen set, which we call our development (or validation) set. However, given that we optimized our hyper-parameters to the development set, to get a true fair assesment of the model, we test it with respect to a held-out test set at the end, and generally report those numbers.
#
# For now, we'll just use a training set and a testing set. We'll train with the training set and evaluate on the test set to see how well our model performs.
# +
num_steps = 3000
batch_size = 128
with tf.Session() as session:
# this operation initializes all the variables we made earlier.
tf.global_variables_initializer().run()
for step in range(num_steps):
# Generate a minibatch.
offset = (step * batch_size) % (X_train.shape[0] - batch_size)
batch_data = X_train[offset:(offset + batch_size), :]
batch_labels = y_train[offset:(offset + batch_size), :]
# Create a dictionary to pass in the batch data.
feed_dict_train = {data_placeholder: batch_data, labels_placeholder : batch_labels}
# Run the optimizer, the loss, the predictions.
# We can run multiple things at once and get their outputs.
_, loss_value_train, predictions_value_train, accuracy_value_train = session.run(
[optimizer, loss, prediction, accuracy], feed_dict=feed_dict_train)
# Print stuff every once in a while.
if (step % 100 == 0):
print("Minibatch train loss at step", step, ":", loss_value_train)
print("Minibatch train accuracy: %.3f%%" % (accuracy_value_train*100))
feed_dict_test = {data_placeholder: X_test, labels_placeholder: y_test}
loss_value_test, predictions_value_test, accuracy_value_test = session.run(
[loss, prediction, accuracy], feed_dict=feed_dict_test)
print("Test loss: %.3f" % loss_value_test)
print("Test accuracy: %.3f%%" % (accuracy_value_test*100))
# -
# Running this code, you’ll see the network train and output its performance as it learns. I was able to get it to 65.5% accuracy. This is just OK, considering random guessing gets you 33.3% accuracy. In the next tutorial, you'll learn some ways to improve upon this.
#
# ## Concluding Thoughts
# This was a brief introduction into TensorFlow. There is so, so much more to learn and explore, but hopefully this has given you some base knowledge to expand upon. As an additional exercise, you can see what you can do with this code to improve the performance. Ideas include: randomizing mini-batches, making the network deeper, using word embeddings (see below) rather than bag-of-words, trying different optimizers (like Adam), different weight initializations. We’ll explore some of these tomorrow.
#
# #### More on Word Embeddings
#
# In this lab we used Bag-of-Words to represent a tweet. Word Embeddings are a more meaningful representation. The basic idea is we represent a word with a vector $\phi$ by the context the word appears in. We do this by training a neural network to predict the context of words across a large training set. The weights of that neural network can then be thought of as a dense and useful representation that captures context. This is useful because now our representations of words captures actual semantic similarity.
#
# Word Embeddings capture all kinds of useful semantic relationships. For example, one cool emergent property is $ \phi(king) - \phi(queen) = \phi(man) - \phi(woman)$. To learn more about the magic behind word embeddings we recommend <NAME>'s [blog post](https://colah.github.io/posts/2014-07-NLP-RNNs-Representations/). A common tool for generating Word Embeddings is word2vec.
# # Solutions
# +
a = tf.placeholder(tf.float32)
b = tf.placeholder(tf.float32)
c = tf.add(a, b)
d = tf.sub(b, 1)
e = tf.mul(c, d)
with tf.Session() as session:
a_data, b_data = 3.0, 6.0
feed_dict = {a: a_data, b: b_data}
output = session.run([e], feed_dict=feed_dict)
print(output) # 45.0
n_input_nodes = 2
n_output_nodes = 1
x = tf.placeholder(tf.float32, (None, n_input_nodes))
W = tf.Variable(tf.ones((n_input_nodes, n_output_nodes)), dtype=tf.float32)
b = tf.Variable(tf.zeros(n_output_nodes), dtype=tf.float32)
'''TODO: Define the operation for z (use tf.matmul to multiply W and x).'''
z = tf.matmul(x, W) + b
'''TODO: Define the operation for out (use tf.sigmoid).'''
out = tf.sigmoid(z)
test_input = [[0.5, 0.5]]
with tf.Session() as session:
tf.global_variables_initializer().run(session=session)
feed_dict = {x: test_input}
output = session.run([out], feed_dict=feed_dict)
print(output[0]) # This should output 0.73105. If not, double-check your code above
'''TODO: manually optimize weight_values and bias_value to classify points'''
# Modify weight_values, bias_value in the above code to adjust the decision boundary
# See if you can classify all the points correctly (all markers green)
weight_values = np.array([[-0.1], [0.2]]) # TODO change values and re-run
bias_value = np.array([[0.5]]) #TODO change values and re-run
# A pretty good boundary is made with:
weight_values = np.array([[0.03], [0.12]])
bias_value = np.array([[-0.5]])
x = tf.placeholder(tf.float32, (None, 2), name='x')
W = tf.Variable(weight_values, name='W', dtype=tf.float32)
b = tf.Variable(bias_value, name='b', dtype=tf.float32)
z = tf.matmul(x, W) + b
out = tf.sigmoid(z)
data = np.array([[2, 7], [1, 7], [3, 1], [3, 3], [4, 3], [4, 6], [6, 5], [7, 7], [7, 5], [2, 4], [2, 2]])
y = np.array([1, 1, 0, 0, 0, 1, 1, 1, 1, 0, 0])
with tf.Session() as session:
tf.global_variables_initializer().run(session=session)
utils.classify_and_plot(data, y, x, out, session)
# load data
X, y, index_to_word, sentences = utils.load_sentiment_data_bow()
X_train, y_train, X_test, y_test = utils.split_data(X, y)
vocab_size = X.shape[1]
n_classes = y.shape[1]
print("Tweet:", sentences[5])
print("Label:", y[5])
print("Bag of Words Representation:", X_train[5])
data_placeholder = tf.placeholder(tf.float32, shape=(None, vocab_size), name='data_placeholder')
'''TODO: Make the labels placeholder.''';
labels_placeholder = tf.placeholder(tf.float32, shape=(None, n_classes), name='labels_placeholder')
# Define Network Parameters
# Here, we can define how many units will be in each hidden layer.
n_hidden_units_h0 = 512
n_hidden_units_h1 = 256
# Weights going from input to first hidden layer.
# The first value passed into tf.Variable is initialization of the variable.
# We initialize our weights to be sampled from a truncated normal, as this does something
# called symmetry breaking and keeps the neural network from getting stuck at the start.
# Since the weight matrix multiplies the previous layer to get inputs to the next layer,
# its size is prev_layer_size x next_layer_size.
h0_weights = tf.Variable(
tf.truncated_normal([vocab_size, n_hidden_units_h0]),
name='h0_weights')
h0_biases = tf.Variable(tf.zeros([n_hidden_units_h0]),
name='h0_biases')
'''TODO: Set up variables for the weights going into the second hidden layer.
You can check out the tf.Variable API here: https://www.tensorflow.org/api_docs/python/state_ops/variables.
''';
h1_weights = tf.Variable(
tf.truncated_normal([n_hidden_units_h0, n_hidden_units_h1]),
name='h1_weights')
h1_biases = tf.Variable(tf.zeros([n_hidden_units_h1]),
name='h1_biases')
# Weights going into the output layer.
out_weights = tf.Variable(
tf.truncated_normal([n_hidden_units_h1, n_classes]),
name='out_weights')
out_biases = tf.Variable(tf.zeros([n_classes]),
name='out_biases')
# Define Computation Graphs
hidden0 = tf.nn.relu(tf.matmul(data_placeholder, h0_weights) + h0_biases)
'''TODO: write the computation to get the output of the second hidden layer.''';
hidden1 = tf.nn.relu(tf.matmul(hidden0, h1_weights) + h1_biases)
logits = tf.matmul(hidden1, out_weights) + out_biases
'''TODO: Define the loss. Use tf.nn.softmax_cross_entropy_with_logits.'''
loss = tf.reduce_mean(
tf.nn.softmax_cross_entropy_with_logits(logits, labels_placeholder))
learning_rate = 0.0002
# Define the optimizer operation. This is what will take the derivate of the loss
# with respect to each of our parameters and try to minimize it.
optimizer = tf.train.GradientDescentOptimizer(learning_rate).minimize(loss)
prediction = tf.nn.softmax(logits)
# Compute the accuracy
prediction_is_correct = tf.equal(tf.argmax(logits, 1), tf.argmax(labels_placeholder, 1))
accuracy = tf.reduce_mean(tf.cast(prediction_is_correct, tf.float32))
num_steps = 3000
batch_size = 128
with tf.Session() as session:
# this operation initializes all the variables we made earlier.
tf.global_variables_initializer().run()
for step in range(num_steps):
# Generate a minibatch.
offset = (step * batch_size) % (X_train.shape[0] - batch_size)
batch_data = X_train[offset:(offset + batch_size), :]
batch_labels = y_train[offset:(offset + batch_size), :]
# Create a dictionary to pass in the batch data.
feed_dict_train = {data_placeholder: batch_data, labels_placeholder : batch_labels}
# Run the optimizer, the loss, the predictions.
# We can run multiple things at once and get their outputs.
_, loss_value_train, predictions_value_train, accuracy_value_train = session.run(
[optimizer, loss, prediction, accuracy], feed_dict=feed_dict_train)
# Print stuff every once in a while.
if (step % 100 == 0):
print("Minibatch train loss at step", step, ":", loss_value_train)
print("Minibatch train accuracy: %.3f%%" % (accuracy_value_train*100))
feed_dict_test = {data_placeholder: X_test, labels_placeholder: y_test}
loss_value_test, predictions_value_test, accuracy_value_test = session.run(
[loss, prediction, accuracy], feed_dict=feed_dict_test)
print("Test loss: %.3f" % loss_value_test)
print("Test accuracy: %.3f%%" % (accuracy_value_test*100))
# -
|
6.S191-Lab1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Import required modules
from urllib.request import urlopen
from bs4 import BeautifulSoup
import pandas as pd
# +
# Url of 2021 recieving stats page.
url = "https://www.pro-football-reference.com/years/2021/"
# Open URL and pass to beautiful soup
html = urlopen(url)
stats_page = BeautifulSoup(html)
# -
# Collect table headers
column_headers = stats_page.findAll('tr')[0]
column_headers = [i.getText() for i in column_headers.findAll('th')]
print(column_headers)
# +
# Collect table rows
rows = stats_page.findAll('tr')
# Get stats from each row
team_stats = []
for i in range(len(rows)):
team_stats.append([col.getText() for col in rows[i].findAll('th')])
team_stats.append([col.getText() for col in rows[i].findAll('td')])
# -
# Check first row of wr_stats
print(team_stats[0:5])
# +
# Create dataframe from the scraped headers and player stats
data = pd.DataFrame(team_stats, columns = column_headers[:])
data.head(20)
# -
# Adjust column names.
rec_data = data.rename(columns={"GS":"G's Started","Y/R":"Yds per Rec","1D":"1st Down Rec"})
rec_data.head()
# +
# Create data subset to change data types.
numerical_categories = ["Age","G's Started","Tgt","Rec","Ctch%","Yds","Yds per Rec","TD","1st Down Rec","Lng","Y/Tgt","R/G","Y/G","Fmb"]
# Create new data subset.
rec_data_subset = rec_data[["Player","Tm","Pos"] + numerical_categories]
rec_data_subset.head()
# -
# Check data types of all columns
rec_data_subset.dtypes
# Remove % sign from catch%.
rec_data_subset["Ctch%"] = rec_data_subset["Ctch%"].str.replace("%","")
rec_data_subset.head()
# Convert data to numerical data where necessary
for i in numerical_categories:
rec_data_subset[i] = pd.to_numeric(rec_data_subset[i])
rec_data_subset.head()
rec_data_subset.dtypes
# Import plotting libraries
import matplotlib
import hvplot.pandas
rec_data_subset = rec_data_subset.sort_values(by=['Yds'],ascending=False)
rec_data_subset.head()
|
NFL_team_stats.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/mdzafri/openvino-workshop/blob/main/LeNet/7_1_MNIST_with_LeNet.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="lsQJFOieRtEu"
# # MNIST with LeNet
# Understanding and Implementing LeNet-5 CNN Architecture
#
# Reference:
# 1. [<NAME>](https://towardsdatascience.com/understanding-and-implementing-lenet-5-cnn-architecture-deep-learning-a2d531ebc342)
# 2. [<NAME>](https://colab.research.google.com/github/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_06_2_cnn.ipynb)
# + [markdown] id="fFdjKP0kuL39"
# # 1. Dataset Preparation
#
#
# + id="D1uXvOTePrF3"
import numpy as np
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
# + id="LDYJf_n6oEH3"
# Load the dataset
num_classes = 10
input_shape = (28, 28, 1)
(x_train, y_train), (x_test, y_test) = keras.datasets.mnist.load_data()
# + id="MhFjvzUsqzX1"
# OPTIONAL: Display some dataset samples as an image
# %matplotlib inline
import matplotlib.pyplot as plt
import random
ROWS = 6
random_indices = random.sample(range(x_train.shape[0]), ROWS*ROWS)
sample_images = x_train[random_indices, :]
plt.clf()
fig, axes = plt.subplots(ROWS,ROWS,
figsize=(ROWS,ROWS),
sharex=True, sharey=True)
for i in range(ROWS*ROWS):
subplot_row = i//ROWS
subplot_col = i%ROWS
ax = axes[subplot_row, subplot_col]
plottable_image = np.reshape(sample_images[i,:], (28,28))
ax.imshow(plottable_image, cmap='gray_r')
ax.set_xbound([0,28])
plt.tight_layout()
plt.show()
# + id="-2JRMCQOq5i7" colab={"base_uri": "https://localhost:8080/"} outputId="cfed5053-4d13-48e2-df02-8302877e2f2e"
# Normalize images to the [0, 1] range
# This is to make the calculations more efficient
x_train = x_train.astype("float32") / 255
x_test = x_test.astype("float32") / 255
# Make sure images have shape (28, 28, 1)
x_train = np.expand_dims(x_train, -1)
x_test = np.expand_dims(x_test, -1)
print("x_train shape:", x_train.shape)
print(x_train.shape[0], "train samples")
print(x_test.shape[0], "test samples")
# convert class label vectors to binary class matrices (convert to 1-hot format)
y_train = keras.utils.to_categorical(y_train, num_classes)
y_test = keras.utils.to_categorical(y_test, num_classes)
# + [markdown] id="uAhXPgZuublL"
# # 2. Select/Design Model
#
# Choose one of these models to train. DO NOT RUN ALL CELLS HERE. Just choose one, then see the output.
# + id="LlWyId3LwNI5"
# 2-layer NN
model = keras.Sequential(
[
layers.Flatten(input_shape=(28, 28)), # Input layer
layers.Dense(100, activation='relu'), # Hidden layer(s)
layers.Dense(num_classes, activation='softmax') # Output layer
]
)
model.summary()
# + colab={"base_uri": "https://localhost:8080/", "height": 132} id="5OOcxMHIsB3f" outputId="75e2742a-589f-4df7-a882-404b9467d161"
# CNN LeNet model
model = keras.Sequential(
[
keras.Input(shape=input_shape)
layers.Conv2D(32, kernel_size=(3, 3), activation='relu'), #C1
layers.MaxPooling2D(pool_size=(2, 2)), #S2
layers.Conv2D(32, kernel_size=(3, 3), activation='relu'), #C3
layers.MaxPooling2D(pool_size=(2, 2)), #S4
layers.Flatten(), #Flatten
layers.Dense(64, activation='relu'), #C5
layers.Dense(num_classes, activation='softmax') #Output layer
]
)
model.summary()
# + id="m0cuVgLTQqp2" colab={"base_uri": "https://localhost:8080/"} outputId="0729ada8-0d7a-4a99-a2bf-135f64f451d7"
# LeNet-5 model
model = keras.Sequential(
[
layers.Conv2D(10, kernel_size=5, strides=1, activation='relu', padding='same', input_shape=x_train[0].shape), #C1
layers.AveragePooling2D(), #S2
layers.Conv2D(16, kernel_size=5, strides=1, activation='relu', padding='valid'), #C3
layers.AveragePooling2D(), #S4
layers.Flatten(), #Flatten
layers.Dense(120, activation='relu'), #C5
layers.Dense(84, activation='relu'), #F6
layers.Dense(num_classes, activation='softmax') #Output layer
]
)
model.summary()
# + [markdown] id="l8JAyLjWvLWH"
# # 3. Train the model
# + id="2Wm7qx7gP72Y" colab={"base_uri": "https://localhost:8080/"} outputId="bfaf329f-2166-484f-bc83-547ac820c108"
# set the loss, optimizer and metrics
model.compile(loss="categorical_crossentropy", optimizer=keras.optimizers.Adam(learning_rate=0.01), metrics=["accuracy"])
# train/fit the model
model.fit(x_train, y_train, batch_size=128, epochs=5, validation_split=0.1)
# + id="t-r9YVivP-L3" colab={"base_uri": "https://localhost:8080/"} outputId="9d23031f-f43f-4cd7-be8d-1517a3e251a4"
# Evaluate the trained model performance
score = model.evaluate(x_test, y_test)
print("Test loss:", score[0])
print("Test accuracy:", score[1])
# + [markdown] id="0supgTUEvNV5"
# # 4. Test the trained model
# + [markdown] id="ix_4txM0xWvz"
# ### Make a canvas for user to draw a digit
# + id="XpumNF-fuyV_"
# Make a canvas for user to draw a digit
# then save the drawing as a png file
# source: https://gist.github.com/korakot/8409b3feec20f159d8a50b0a811d3bca
from IPython.display import HTML, Image
from google.colab.output import eval_js
from base64 import b64decode
canvas_html = """
<canvas width=%d height=%d></canvas>
<button>Finish</button>
<script>
var canvas = document.querySelector('canvas')
var ctx = canvas.getContext('2d')
ctx.fillStyle = "white";
ctx.fillRect(0, 0, canvas.width, canvas.height);
ctx.lineWidth = %d
var button = document.querySelector('button')
var mouse = {x: 0, y: 0}
canvas.addEventListener('mousemove', function(e) {
mouse.x = e.pageX - this.offsetLeft
mouse.y = e.pageY - this.offsetTop
})
canvas.onmousedown = ()=>{
ctx.beginPath()
ctx.moveTo(mouse.x, mouse.y)
canvas.addEventListener('mousemove', onPaint)
}
canvas.onmouseup = ()=>{
canvas.removeEventListener('mousemove', onPaint)
}
var onPaint = ()=>{
ctx.lineTo(mouse.x, mouse.y)
ctx.stroke()
}
var data = new Promise(resolve=>{
button.onclick = ()=>{
resolve(canvas.toDataURL('image/png'))
}
})
</script>
"""
def draw(filename='drawing.png', w=150, h=150, line_width=10):
display(HTML(canvas_html % (w, h, line_width)))
data = eval_js("data")
binary = b64decode(data.split(',')[1])
with open(filename, 'wb') as f:
f.write(binary)
print("image saved as: ")
print(filename)
# return len(binary)
draw()
# + id="habVaeznymeh"
import matplotlib.pyplot as plt
from PIL import Image, ImageOps # import pillow image manipulation tool
# Load the image to be tested
user_image = Image.open('drawing.png')
user_image = ImageOps.grayscale(user_image)
user_image = ImageOps.invert(user_image)
# Resize to input_shape
user_image = user_image.resize((input_shape[0],input_shape[1]))
plt.imshow(user_image)
user_image = np.array(user_image).astype("float32") / 255
# user_image = np.expand_dims(user_image, axis=0)
user_image = user_image.reshape(-1, 28, 28, 1)
# print("user_image shape:", user_image.shape)
# Predict the class of the drawing
result = model.predict(user_image)
print(result)
result = np.argmax(result,axis=1)
print("The AI thinks this is the number:", result[0])
|
LeNet/7_1_MNIST_with_LeNet.ipynb
|
; -*- coding: utf-8 -*-
; ---
; jupyter:
; jupytext:
; text_representation:
; extension: .pro
; format_name: light
; format_version: '1.5'
; jupytext_version: 1.14.4
; kernelspec:
; display_name: IDL [conda env:gdl] *
; language: IDL
; name: conda-env-gdl-idl
; ---
; ## GDL demo notebook
; Demonstration of GDL [(gnudatalanguage)](https://github.com/gnudatalanguage/gdl)
;
; This notebook creates a Shepp-Logan phantom, projects it and then performs an FBP reconstruction.
; +
;; <NAME> and <NAME>, “The Fourier reconstruction of a head section,”
;; IEEE Trans. Nucl. Sci. 21(3), 21–43 (1974).
function shepplogan, size = size
if NOT keyword_set(size) then size = 256
phantom = fltarr(size,size)
tmp = (findgen(size)-((size-1)/2.0)) / (size/2.0)
xcor = rebin(tmp,size,size)
ycor = rebin(transpose(tmp),size,size)
tmp = fltarr(size,size)
aa={cx: 0.0, cy: 0.0, maj:0.69, min:0.92, theta: 0.0, val: 2.0 }
bb={cx: 0.0, cy:-0.0184, maj:0.6624, min:0.874, theta: 0.0, val:-0.98}
cc={cx: 0.22, cy: 0.0, maj:0.11, min:0.31, theta:-18.0, val:-0.02}
dd={cx:-0.22, cy: 0.0, maj:0.16, min:0.41, theta: 18.0, val:-0.02}
ee={cx: 0.0, cy: 0.35, maj:0.21, min:0.25, theta: 0.0, val:-0.01}
ff={cx: 0.0, cy: 0.1, maj:0.046, min:0.046, theta: 0.0, val:-0.01}
gg={cx: 0.0, cy:-0.1, maj:0.046, min:0.046, theta: 0.0, val:-0.01}
hh={cx:-0.08, cy:-0.605, maj:0.046, min:0.023, theta: 0.0, val:-0.01}
ii={cx: 0.0, cy:-0.605, maj:0.023, min:0.023, theta: 0.0, val:-0.01}
jj={cx: 0.06, cy:-0.605, maj:0.023, min:0.046, theta: 0.0, val:-0.01}
list = [aa,bb,cc,dd,ee,ff,gg,hh,ii,jj]
for n = 0, n_elements(list)-1 do begin
tmp = ((xcor-list[n].cx) / list[n].maj)^2 $
+ ((ycor-list[n].cy) / list[n].min)^2
if list[n].theta NE 0 then begin
nx = (size-1) * (list[n].cx + 1) / 2
ny = (size-1) * (list[n].cy + 1) / 2
tmp = rot(tmp, -list[n].theta, 1, nx, ny, /interp, /pivot)
endif
phantom[where(tmp LE 1.0)] += list[n].val
endfor
return, phantom < 1.1
end
; -
;; Enable inline plotting
!inline=1
phantom = shepplogan()
window, 0, xsize=256, ysize=256
tvscl, phantom > 0.95
; Now we define simple forward and backprojection functions:
; +
function proj, image, angle
sino_size_x = max(size(image,/dim))
sino_size_y = n_elements(angle)
sino = fltarr(sino_size_x, sino_size_y)
for aa=0, n_elements(angle)-1 do begin
sino[*,aa] = total(rot(image, angle[aa], /interp), 2)
endfor
return, sino
end
; +
function back, sino, angle
image_size = n_elements(sino[*,0])
image = fltarr(image_size,image_size)
for aa=0, n_elements(angle)-1 do begin
image += rot(rebin(sino[*,aa],[image_size,image_size]), -angle[aa], /interp)
endfor
return, image / n_elements(angle)
end
; -
; This allows us to create a sinogram:
angles = findgen(360)
sino = proj(phantom, angles)
window, 0, xsize=256, ysize=360
tvscl, sino
; On this we can apply a simple FBP reconstruction:
; +
;; <NAME>, “Revisit of the Ramp Filter,”
;; IEEE Trans. Nucl. Sci. 62(1), 131–136 (2015).
function fbp, sino, angles
ntot = n_elements(sino[*,0])
nang = n_elements(sino[0,*])
npos = ntot / 2 + 1 ; integer division needed !
nneg = ntot - npos
freq = findgen(ntot)
freq[npos:ntot-1] = REVERSE(freq[1:nneg])
freq[0] = 1
filter = -1 / (!pi * freq)^2
filter[where(freq mod 2 EQ 0)] *= -0.0
filter[0] = 0.25
filter = abs(fft(filter)) * ntot
filter[ntot/4:ntot/4+ntot/2-1] *= 0.0
filter = rebin(filter,ntot,nang)
;; apply filter to the sinogram
fsino = fft(sino, dim=1)
fsino *= filter
fsino = fft(fsino, dim=1, /overwrite, /inverse)
fsino = !pi * real_part(fsino)
;; backproject the filtered sinogram
return, back(fsino, angles)
end
; +
reconstruction = fbp(sino, angles)
window, 0, xsize=512, ysize=256
tvscl, [phantom, reconstruction] > 0.8
; -
; ### The end
|
tests/notebooks/ipynb_idl/demo_gdl_fbp.ipynb
|
# # 📝 Exercise 02
#
# The aim of this exercise is to find out whether a decision tree
# model is able to extrapolate.
#
# By extrapolation, we refer to values predicted by a model outside of the
# range of feature values seen during the training.
#
# We will first load the regression data.
# +
import pandas as pd
penguins = pd.read_csv("../datasets/penguins_regression.csv")
data_columns = ["Flipper Length (mm)"]
target_column = "Body Mass (g)"
data_train, target_train = penguins[data_columns], penguins[target_column]
# -
# <div class="admonition note alert alert-info">
# <p class="first admonition-title" style="font-weight: bold;">Note</p>
# <p class="last">If you want a deeper overview regarding this dataset, you can refer to the
# Appendix - Datasets description section at the end of this MOOC.</p>
# </div>
# First, create two models, a linear regression model and a decision tree
# regression model, and fit them on the training data. Limit the depth at
# 3 levels for the decision tree.
# +
# Write your code here.
# -
# Create a testing dataset, ranging from the minimum to the maximum of the
# flipper length of the training dataset. Get the predictions of each model
# using this test dataset.
# +
# Write your code here.
# -
# Create a scatter plot containing the training samples and superimpose the
# predictions of both model on the top.
# +
# Write your code here.
# -
# Now, we will check the extrapolation capabilities of each model. Create a
# dataset containing the value of your previous dataset. Besides, add values
# below and above the minimum and the maximum of the flipper length seen
# during training.
# +
# Write your code here.
# -
# Finally, make predictions with both model on this new testing set. Repeat
# the plotting of the previous exercise.
# +
# Write your code here.
|
notebooks/trees_ex_02.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Feature importance analysis
# for Tree Mortality Prediction based on Growth Patterns
# using Machine Learning Models
#
# __<NAME>__, Acton Boxborough Regional HS, <EMAIL>
# __<NAME>__, Department of Earth & Environment, Boston University, <EMAIL>
#
# ### Harvard Forest Ecology Symposium
# #### March 16-17, 2021
#
#
# data: [HF213](https://harvardforest1.fas.harvard.edu/exist/apps/datasets/showData.html?id=HF213)
#
# Use classification algorithms to predict A(live) or D(ead) labels in __mortality13__ and __mortality14__ columns using these features:
# - spp: USDA Plants database species code
# - dbh09: diameter at Breast Height (1.4m) in year 2009 (unit: centimeter / missing value: NA)
# - dbh11: diameter at Breast Height (1.4m) in year 2011 (unit: centimeter / missing value: NA)
# - dbh12: diameter at Breast Height (1.4m) in year 2012 (unit: centimeter / missing value: NA)
# - dbh13: diameter at Breast Height (1.4m) in year 2013 (unit: centimeter / missing value: NA)
#
#
#
#
# ##### Tree mortality prediction based on growth patterns using Machine Learning
# Feature importance analysis
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
# +
import sys, os, pathlib, shutil, platform
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.linear_model import LogisticRegression
from sklearn.neighbors import KNeighborsClassifier
from sklearn import svm
from sklearn.ensemble import RandomForestClassifier
from sklearn.feature_selection import RFE, RFECV
from sklearn.metrics import (
classification_report,
confusion_matrix,
accuracy_score,
roc_auc_score,
roc_curve,
auc,
)
from sklearn.model_selection import (
train_test_split,
StratifiedShuffleSplit,
StratifiedKFold,
)
from sklearn.preprocessing import OrdinalEncoder
from sklearn.compose import make_column_transformer
from sklearn.impute import SimpleImputer
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import OneHotEncoder
from sklearn.preprocessing import OrdinalEncoder
from sklearn.preprocessing import StandardScaler
# -
# # !pip install xgboost
import xgboost
print(xgboost.__version__)
import sklearn
print(sklearn.__version__)
MINIMUM_COUNT = 10
TRAIN_DATA = 0.6
# +
# # !/opt/conda/bin/conda install -c anaconda seaborn pandas scikit-learn -y
# -
# %matplotlib inline
# !pwd
# # !ls -la ./../data/hrvardf/HF213
dataFileName='hf213-01-hf-inventory.csv'
dataPathFull= pathlib.Path('./../data/harvardf/HF213') / dataFileName
myData = pd.read_csv(str(dataPathFull))
myData.shape
myData.head(2)
myData.tail(2)
myData.info()
myData.isnull().sum()
# myData.dropna()
# myData.isna().sum()
# basic descriptive statistics for numeric columns:
myData.describe()
corr=myData.corr()
plt.figure(figsize = (9, 7))
sns.heatmap(corr, cmap="RdBu",
xticklabels=corr.columns.values,
yticklabels=corr.columns.values)
plt.show()
# +
# myData.groupby('spp').size()
myCols = ['spp', 'mortality14', 'dmg14']
myData[myCols[0]].value_counts(dropna=False)
myData[myCols[1]].value_counts(dropna=False)
myData[myCols[2]].value_counts(dropna=False)
# myData.pivot_table(index = [myCols[0]]
# , columns = myCols[1]
# , values = myCols[2]
# , aggfunc=np.sum, fill_value=0)
# -
import seaborn as sns
sns.countplot(x= myData['spp'],label="spp Count")
plt.show()
# +
myData['spp'].value_counts(dropna=False)
removeSPP = myData['spp'].value_counts(dropna=False).loc[lambda x : x<MINIMUM_COUNT].index.tolist()
removeSPP
# filteredData = myData.replace(dict.fromkeys(removeSPP, 'TooFew'))
# filteredData['spp'].value_counts(dropna=False)
# +
featureColumn_01=['spp', 'dbh09', 'dbh11', 'dbh12']
# featureColumn_01=[ 'dbh09', 'dbh11', 'dbh12']
labelColumn_01 = 'mortality13'
featureColumn_02=['spp', 'dbh09', 'dbh11', 'dbh12', 'dbh13']
# featureColumn_02=['dbh09', 'dbh11', 'dbh12', 'dbh13']
labelColumn_02 = 'mortality14'
labelColumn = labelColumn_02
featureColumn = featureColumn_02
# +
sorted(set(featureColumn+[labelColumn]))
filteredDataML = myData[sorted(set(featureColumn+[labelColumn]))]
# filteredDataML.shape
# filteredDataML.head()
# filteredDataML[labelColumn].value_counts(dropna=False)
# -
sns.pairplot(filteredDataML)
from sklearn.impute import SimpleImputer
features_cols = [x for x in featureColumn if x != 'spp']
imputer = SimpleImputer(missing_values=np.nan, strategy='median')
imputer.fit(filteredDataML[features_cols])
imputed = imputer.transform(filteredDataML[features_cols])
filteredDataML[features_cols]=imputer.transform(filteredDataML[features_cols])
filteredDataML.columns
from itertools import combinations
list(combinations(sorted(features_cols), 2))
[print(col1, col2) for col1, col2 in combinations(sorted(features_cols), 2)]
# +
filteredDataML.columns
# col1= trainData.columns[0]
# col2= trainData.columns[1]
# trainData1 = trainData.join(pd.DataFrame(((trainData[col2]/(trainData[col1]) -1)*100).rename(f'{col1}to{col2}')))
filteredDataML = filteredDataML.join(
pd.concat([(((filteredDataML[col2]/(filteredDataML[col1]) -1)*100)
.rename(f'{col2}to{col1}'))
for col1, col2 in combinations(sorted(features_cols), 2)], 1))
# testData = testData.join(
# pd.concat([(((testData[col2]/(testData[col1]) -1)*100)
# .rename(f'{col2}to{col1}'))
# for col1, col2 in combinations(sorted(numCols), 2)], 1))
# validationData = validationData.join(
# pd.concat([(((validationData[col2]/(validationData[col1]) -1)*100)
# .rename(f'{col2}to{col1}'))
# for col1, col2 in combinations(sorted(numCols), 2)], 1))
# trainData.head(2)
# testData.head(2)
# validationData.head(2)
filteredDataML.columns
# +
mortality_13_14 = pd.crosstab(index=myData['mortality14']
,columns=myData['mortality13']
, margins=True)
mortality_13_14
mortality_spp = pd.crosstab(index=myData['spp']
,columns=myData['mortality14']
, margins=True)
mortality_spp
# -
corr=filteredDataML.corr()
corr
plt.figure(figsize = (9, 7))
sns.heatmap(corr, cmap="RdBu",
xticklabels=corr.columns.values,
yticklabels=corr.columns.values)
plt.show()
# +
filteredDataML[labelColumn].value_counts(dropna=False)
filteredDataML = filteredDataML[filteredDataML[labelColumn].isin(['D', 'A'])]
filteredDataML.shape
filteredDataML.head()
filteredDataML[labelColumn].value_counts(dropna=False)
# -
# oh_encoder = OneHotEncoder(handle_unknown='ignore')
# enc_lbl_df = pd.DataFrame((oh_encoder.fit_transform(filteredDataML[['spp']])).toarray())
# enc_lbl_df
# +
# https://scikit-learn.org/stable/auto_examples/ensemble/plot_stack_predictors.html#sphx-glr-auto-examples-ensemble-plot-stack-predictors-py
catCols = filteredDataML.columns[filteredDataML.dtypes == 'O']
numCols = filteredDataML.columns[filteredDataML.dtypes == 'float64']
catCols
numCols
# -
print(filteredDataML.loc[0:3])
# +
stratifySplit = StratifiedShuffleSplit(n_splits=1, train_size=TRAIN_DATA, random_state=1)
trainIdx, tstIdx = next(stratifySplit.split(filteredDataML, filteredDataML[labelColumn]))
# print("\n Train:", sorted(trainIdx))
len(trainIdx)
len(tstIdx)
filteredDataML.loc[filteredDataML.index.intersection(filteredDataML.index[trainIdx])].shape
filteredDataML[filteredDataML.index.isin(filteredDataML.index[trainIdx])].shape
aa=filteredDataML.loc[filteredDataML.index.intersection(filteredDataML.index[tstIdx])]
aa.shape
stratifySplit = StratifiedShuffleSplit(n_splits=1, train_size=TRAIN_DATA, test_size=1-TRAIN_DATA, random_state=1)
testIdx, validationIdx = next(stratifySplit.split(aa, aa[labelColumn]))
len(testIdx)
len(validationIdx)
filteredDataML.shape
# testIdx=tstIdx[testIdx]
# validationIdx=tstIdx[validationIdx]
# print("\n Test:", sorted(testIdx))
# print("\nValidation:", sorted(validationIdx))
# +
trainData=filteredDataML.loc[filteredDataML.index.intersection(filteredDataML.index[trainIdx]),:]
testData=aa.loc[aa.index.intersection(aa.index[testIdx]),:]
validationData = aa.loc[aa.index.intersection(aa.index[validationIdx]),:]
filteredDataML[labelColumn].value_counts(dropna=False)
trainData[labelColumn].value_counts(dropna=False)
testData[labelColumn].value_counts(dropna=False)
validationData[labelColumn].value_counts(dropna=False)
# +
ordinalEncoder = OrdinalEncoder()
ordinalEncoder.fit(filteredDataML[catCols])
ordinalEncoder.categories_
trainData[catCols] = ordinalEncoder.transform(trainData[catCols])
testData[catCols] = ordinalEncoder.transform(testData[catCols])
validationData[catCols] = ordinalEncoder.transform(validationData[catCols])
trainData.head()
# +
# from sklearn.impute import SimpleImputer
# imputer = SimpleImputer(missing_values=np.nan, strategy='median')
# imputer.fit(trainData[featureColumn])
# +
# trainData[featureColumn] = imputer.transform(trainData[featureColumn])
# testData[featureColumn] = imputer.transform(testData[featureColumn])
# validationData[featureColumn] = imputer.transform(validationData[featureColumn])
# +
# from itertools import combinations
# trainData.columns
# # col1= trainData.columns[0]
# # col2= trainData.columns[1]
# # trainData1 = trainData.join(pd.DataFrame(((trainData[col2]/(trainData[col1]) -1)*100).rename(f'{col1}to{col2}')))
# trainData = trainData.join(
# pd.concat([(((trainData[col2]/(trainData[col1]) -1)*100)
# .rename(f'{col2}to{col1}'))
# for col1, col2 in combinations(sorted(numCols), 2)], 1))
# testData = testData.join(
# pd.concat([(((testData[col2]/(testData[col1]) -1)*100)
# .rename(f'{col2}to{col1}'))
# for col1, col2 in combinations(sorted(numCols), 2)], 1))
# validationData = validationData.join(
# pd.concat([(((validationData[col2]/(validationData[col1]) -1)*100)
# .rename(f'{col2}to{col1}'))
# for col1, col2 in combinations(sorted(numCols), 2)], 1))
# trainData.head(2)
# testData.head(2)
# validationData.head(2)
# trainData.columns
# +
newNumCols = trainData.columns[trainData.dtypes == 'float64']
newNumCols = list(newNumCols)
newNumCols.remove(labelColumn)
featureColumn = newNumCols
newNumCols
# -
from sklearn.preprocessing import RobustScaler
robustScaler = RobustScaler(quantile_range = (0.1,0.9))
robustScaler = robustScaler.fit(trainData[newNumCols])
trainData[newNumCols] =robustScaler.transform(trainData[newNumCols])
testData[newNumCols] =robustScaler.transform(testData[newNumCols])
validationData[newNumCols] =robustScaler.transform(validationData[newNumCols])
trainData.describe()
testData.describe()
validationData.describe()
# +
# logistic regression for feature importance
# LogRegModel = LogisticRegression()
# LogRegModel.fit(trainData[featureColumn], trainData[labelColumn])
# # get importance
# importance = LogRegModel.coef_[0]
# # summarize feature importance
# print(featureColumn)
# for i,v in enumerate(importance):
# print('Feature: %0d, %s, Score: %.5f' % (i,featureColumn[i], v))
# # plot feature importance
# plt.bar([x for x in range(len(importance))], importance)
# plt.show()
# -
# ### Feature importance analysis
# for Tree Mortality Prediction based on Growth Patterns
# using Machine Learning Models
#
# +
# CART Feature Importance
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
from xgboost import XGBClassifier
DecTreeModel = DecisionTreeClassifier()
RFModel = RandomForestClassifier()
XGBModel = XGBClassifier()
def FI_Analysis(crt_model):
crt_model.fit(trainData[featureColumn], trainData[labelColumn])
# get importance
importance = crt_model.feature_importances_
# summarize feature importance
print(type(crt_model).__name__)
for i,v in enumerate(importance):
print('Feature: %0d, %s, Score: %.5f' % (i,featureColumn[i], v))
# plot feature importance
plt.bar([x for x in range(len(importance))], importance)
plt.show()
return importance
print(featureColumn)
importances = [FI_Analysis(crt_model) for crt_model in [DecTreeModel, RFModel, XGBModel]]
# +
model_names = [type(crt_model).__name__ for crt_model in [DecTreeModel, RFModel, XGBModel]]
imps = pd.DataFrame.from_dict(dict(zip(model_names, importances))).set_index([featureColumn])
imps
# plt.pcolor(imps)
# plt.yticks(np.arange(0.5, len(imps.index), 1), imps.index)
# plt.xticks(np.arange(0.5, len(imps.columns), 1), imps.columns)
# plt.show()
sns.heatmap(imps, annot=True)
# +
from sklearn.inspection import permutation_importance
from sklearn.metrics import cohen_kappa_score, roc_auc_score, make_scorer
roc_auc_scorer = make_scorer(roc_auc_score)
SVC_model = svm.SVC(kernel='rbf', random_state=0, gamma=.1, C=100, probability=True)
KNN_model = KNeighborsClassifier(n_neighbors=50)
def perm_FI(crt_model, crt_scorer=roc_auc_scorer):
crt_model.fit(trainData[featureColumn], trainData[labelColumn])
# perform permutation importance
results = permutation_importance(crt_model, trainData[featureColumn], trainData[labelColumn], scoring=crt_scorer)
# get importance
importance = results.importances_mean
print(type(crt_model).__name__)
# summarize feature importance
for i,v in enumerate(importance):
print('Feature: %0d, %s, Score: %.5f' % (i,featureColumn[i], v))
# plot feature importance
plt.bar([x for x in range(len(importance))], importance)
plt.show()
return importance
print(featureColumn)
model_list = [DecTreeModel, RFModel, XGBModel, SVC_model]
importances = [perm_FI(crt_model) for crt_model in model_list]
# +
model_names = [type(crt_model).__name__ for crt_model in model_list]
imps= pd.DataFrame.from_dict(dict(zip(model_names, importances))).set_index([featureColumn])
imps
sns.heatmap(imps, annot=True)
# +
from sklearn.metrics import cohen_kappa_score, roc_auc_score, make_scorer
cohen_kappa_scorer = make_scorer(cohen_kappa_score)
model_list = [DecTreeModel, RFModel, XGBModel, SVC_model]
importances = [perm_FI(crt_model, cohen_kappa_scorer) for crt_model in model_list]
# +
model_names = [type(crt_model).__name__ for crt_model in model_list]
imps= pd.DataFrame.from_dict(dict(zip(model_names, importances))).set_index([featureColumn])
imps
sns.heatmap(imps, annot=True)
# -
|
TreeMortalityPrediction_FeatImpAnalysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Import libraries
import pandas as pd
# ### Scrape 2020 Player Salaries
# +
# Loop through all pages for URL and scrape tables
# Set range for pages
pages = []
for i in range(2, 19):
pages.append(i)
# Scrape tables for 2020 page 1
url_2020 = 'http://www.espn.com/nba/salaries/_/seasontype/3'
tables_2020 = pd.read_html(url_2020)[0]
tables_2020['year'] = '2020'
# # Set variable for dataframe
tables_2020_pgs_df = pd.DataFrame()
# # Set URL variable for all other pages
for page in pages:
url_2020_pgs = f'http://www.espn.com/nba/salaries/_/page/{page}/seasontype/3'
tables_2020_pgs = pd.read_html(url_2020_pgs)[0]
tables_2020_pgs['year'] = '2020'
tables_2020_pgs_df = pd.concat(axis = 0, objs=[tables_2020_pgs, tables_2020_pgs_df])
# -
len(tables_2020)
len(tables_2020_pgs_df)
# ### Scrape all other years
# +
years = []
for i in range(2000, 2020):
years.append(i)
years
# +
# tables_all_others = []
tables_all_others_df = pd.DataFrame()
# Scrape tables for all other years - page 1
for year in years:
url_all_others = f'http://www.espn.com/nba/salaries/_/year/{year}/seasontype/3'
tables_all_others = pd.read_html(url_all_others)[0]
tables_all_others['year'] = year
tables_all_others_df = pd.concat(axis = 0, objs=[tables_all_others, tables_all_others_df])
# -
len(tables_all_others_df)
# +
years = []
for i in range(2000, 2020):
years.append(i)
tables_all_others_pgs_df = pd.DataFrame()
# Scrape tables for all other years - all other pages
for year in years:
for page in pages:
url_all_others_pgs = f'http://www.espn.com/nba/salaries/_/year/{year}/page/{page}/seasontype/3'
tables_all_others_pgs = pd.read_html(url_all_others_pgs)[0]
tables_all_others_pgs['year'] = year
tables_all_others_pgs_df = pd.concat(axis = 0, objs=[tables_all_others_pgs, tables_all_others_pgs_df])
# -
len(tables_all_others_pgs_df)
# Concat all dataframes
all_salaries_df = pd.concat(axis = 0, objs=[tables_2020, tables_2020_pgs_df, tables_all_others_df, tables_all_others_pgs_df])
len(all_salaries_df)
# +
# Clean dataframe
# Add column headers
all_salaries_df.columns = ['Rank', 'Player_Position', 'Team', 'Salary', 'Year']
# Drop na
all_salaries_df = all_salaries_df.dropna()
# Drop extra rows
drop_row = all_salaries_df[ (all_salaries_df['Rank'] == 'RK') & (all_salaries_df['Player_Position'] == 'NAME') & (all_salaries_df['Team'] == 'TEAM') & (all_salaries_df['Salary'] == 'SALARY')].index
all_salaries_df.drop(drop_row, inplace=True)
# Split Player_Position into 2 columns
all_salaries_df['Player'], all_salaries_df['Position'] = all_salaries_df['Player_Position'].str.split(',', 1).str
# Convert salary to interger
all_salaries_df['Salary'] = all_salaries_df['Salary'].replace('[\$,]','',regex=True).astype(int)
all_salaries_df['Rank'] = all_salaries_df['Rank'].astype(int)
all_salaries_df['Year'] = all_salaries_df['Year'].astype(int)
# -
len(all_salaries_df)
# Drop Player_Position column
all_salaries_df = all_salaries_df.drop('Player_Position', axis = 1)
all_salaries_df = all_salaries_df.sort_values(['Year', 'Rank'], ascending = (False, True))
# ## Adjust salary for inflation
# !pip install cpi
import cpi
adjusted_salaries_df = all_salaries_df.loc[(all_salaries_df['Year'] != 2020)&(all_salaries_df['Year'] != 2019)]
adjusted_salaries_df['Inflation'] = adjusted_salaries_df.apply(lambda x: cpi.inflate(x.Salary, x.Year), axis=1)
adjusted_salaries_df['Inflation'] = adjusted_salaries_df['Inflation'].astype(int)
adjusted_salaries_df.tail()
current_salaries_df = all_salaries_df.loc[(all_salaries_df['Year'] == 2020)|(all_salaries_df['Year'] == 2019)]
current_salaries_df['Inflation'] = current_salaries_df['Salary']
final_salaries_df = pd.concat([current_salaries_df, adjusted_salaries_df], axis=0).reset_index(drop=True)
final_salaries_df = final_salaries_df[['Rank', 'Team', 'Player', 'Position', 'Salary', 'Inflation']]
final_salaries_df
# ## Save to csv
final_salaries_df.to_csv('nba_salaries.csv', index=False)
|
group_files/jane/notebooks/salary_scrape_pandas.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="0rmnHFFLAbuK"
# <font size = "5"> **[MSE672: Introduction to TEM](https://gduscher.github.io/MSE672-Introduction-to-TEM/)** </font>
#
# <hr style="height:1px;border-top:4px solid #FF8200" />
#
# by
#
# <NAME>
#
# Materials Science & Engineering<br>
# Joint Institute of Advanced Materials<br>
# The University of Tennessee, Knoxville
#
# # Test Notebook
#
# [Download](https://raw.githubusercontent.com/gduscher/MSE672-Introduction-to-TEM//main/notebooks/TestNotebook.ipynb")
#
# [](
# https://colab.research.google.com/github/gduscher/MSE672-Introduction-to-TEM/blob/main/notebooks/TestNotebook.ipynb)
#
# + [markdown] id="zDbWs5s0AC1l"
# First we need to load the libraries we want to use. All of those are installed in Google colab and annaconda.
# + executionInfo={"elapsed": 1026, "status": "ok", "timestamp": 1609169925831, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02112921857027034506"}, "user_tz": 300} id="hldG1pG3ABo4"
import numpy as np
import matplotlib.pylab as plt
# + [markdown] id="L8BZWYPBAOCK"
# Now we run the next code cell to plot a graph.
# + colab={"base_uri": "https://localhost:8080/", "height": 267} executionInfo={"elapsed": 845, "status": "ok", "timestamp": 1609169942112, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02112921857027034506"}, "user_tz": 300} id="BJMl8k_9AGnh" outputId="3eec6ace-946a-47bc-f992-240f3e6226b2" pycharm={"name": "#%%\n"}
# Generate plotting values
t = np.linspace(0, 2*np.pi, 200)
x = 16 * np.sin(t)**3
y = 13 * np.cos(t) - 5 * np.cos(2*t) - 2 * np.cos(3*t) - np.cos(4*t)
plt.figure()
plt.plot(x,y, color='red', linewidth=2)
plt.text(-23, 0, 'I', ha='center', va='center', fontsize=206)
plt.text(20,0, 'MSE 672',va='center', fontsize=206)
plt.gca().set_aspect('equal')
# + [markdown] pycharm={"name": "#%% md\n"}
# Who would have thought that the formula of love is based on trigonometry?
# + pycharm={"name": "#%%\n"}
|
Introduction/TestNotebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 0.0 Importar pacotes
from matplotlib import gridspec
from datetime import datetime
from ipywidgets import fixed
from tabulate import tabulate
from matplotlib import pyplot as plt
import pandas as pd
import numpy as np
import plotly.express as px
import warnings
import tqdm
import ipywidgets as widgets
import seaborn as sns
import matplotlib.ticker as mtick
# ## 0.1 Funções Auxiliares
# +
pd.options.display.float_format = '{:.2f}'.format
plt.rcParams['figure.figsize'] = (11,7)
warnings.filterwarnings('ignore')
# -
# # 1.0 Dados
# + [markdown] heading_collapsed=true
# ## 1.1 Importando dataset
# + hidden=true
df = pd.read_csv('dataset/kc_house_data.csv')
# + [markdown] heading_collapsed=true
# ## 1.2 Dimensão dos dados
# + hidden=true
print('Número de linha:', df.shape[0])
print('Número de colunas:', df.shape[1])
# + [markdown] heading_collapsed=true
# ## 1.3 Tipos de variáveis
# + hidden=true
df.dtypes
# + [markdown] heading_collapsed=true
# ## 1.4 Alterando os tipos de variáveis
# + hidden=true
df['date'] = pd.to_datetime(df['date'])
# -
# ## 1.5 Limpeza de dados
# + [markdown] heading_collapsed=true
# ## 1.6 Criação de novas features
# + hidden=true
df['construcao'] = df['yr_built'].apply(lambda x: '> 1955' if x > 1955
else '< 1955')
df['porao'] = df['sqft_basement'].apply(lambda x: 'nao' if x == 0
else 'sim')
df['mes'] = df['date'].dt.month
df['ano'] = df['date'].dt.year
df['renovacao'] = df['yr_renovated'].apply(lambda x: 'sim' if x > 0 else
'nao' )
df['banheiro'] = df['bathrooms'].apply(lambda x: '0-3' if (x > 0 ) & (x < 3) else
'3-5' if (x > 3) & (x < 5) else
'5-8')
df['season'] = df['mes'].apply(lambda x: 'summer' if (x > 5) & (x < 8) else
'spring' if (x > 2) & (x < 5) else
'fall' if (x > 8) & (x < 12) else
'winter')
# + [markdown] heading_collapsed=true hidden=true
# ### 1.5.1 Duplicatas
# + hidden=true
df.sort_values('id').head(5)
# + hidden=true
df = df.drop_duplicates(subset = ['id'], keep = 'last')
df.sort_values('id').head(5)
# + [markdown] heading_collapsed=true hidden=true
# ### 1.5.1 Check NA
# + hidden=true
df.isna().sum()
# + [markdown] heading_collapsed=true hidden=true
# ### 1.5.2 Estatística descritiva
# + hidden=true
df = df.copy()
# + hidden=true
#Incluindo somente variáveis numéricas
atri_num = df.select_dtypes(include = ['int64', 'float64'])
#deletando a coluna 'ID'
atri_num = atri_num.iloc[:, 1: ]
# + hidden=true
#análise descritiva
#medidas de tendencia central
df_mean = pd.DataFrame(atri_num.apply(np.mean)).T
df_median = pd.DataFrame(atri_num.apply(np.median)).T
#medidas de dispersão
df_std = pd.DataFrame(atri_num.apply(np.std)).T
df_min = pd.DataFrame(atri_num.apply(np.min)).T
df_max = pd.DataFrame(atri_num.apply(np.max)).T
#concatenando
est = pd.concat( [df_mean, df_median, df_std, df_min, df_max ] ).T.reset_index()
#alterando o nome das colunas
est.columns = [ 'atributos','media', 'mediana', 'std', 'min', 'max']
print(est)
est.hist();
# + hidden=true
#substituindo o valor de 33 na coluna bedrooms por 3
linha = df[df['bedrooms'] == 33]
df = df.drop(15870)
df['bedrooms'].unique()
# -
# # 2.0 Exploração de dados
# + [markdown] heading_collapsed=true
# ## Correlação
# + hidden=true
matrix = np.triu(df.corr())
sns.heatmap(df.corr(), cmap = 'OrRd', square = True, annot = False, center = 0,
mask = matrix);
# -
# ## 2.1 Insights
df = df.copy()
df.dtypes
# ### **H1:** Imóveis com vista para a água são em média 30% mais caros
# +
h1 = df[['price', 'waterfront']].groupby('waterfront').mean().reset_index()
h1['waterfront'] = h1['waterfront'].astype(str)
sns.barplot(h1['waterfront'], h1['price']);
# -
# ### H2: Imóveis com data de construção menor que 1955 são em média 50% mais baratos.
#
# +
h2 = df[['construcao', 'price']].groupby('construcao').mean().reset_index()
sns.barplot(h2['construcao'], h2['price']);
# -
# ### H3: Imóveis sem porão possuem área total (sqrt_lot) são 40% maiores do que imóveis com porões.
#
# +
h3 = df[['porao', 'sqft_lot', 'price']].groupby('porao').sum().reset_index()
h3
sns.barplot(h3['porao'], h3['sqft_lot']);
# -
# ### H4: O crescimento do preço dos imóveis ano após ano (YoY) é de 10%
#
# +
h4 = df[['price', 'ano']].groupby('ano').sum().reset_index()
h4['ano'] = h4['ano'].astype(str)
sns.barplot(h4['ano'], h4['price']);
# -
# ### H4.1: O crescimento do preço dos imóveis MoM é de 10%
# +
h41 = df[['mes', 'price']].groupby('mes').sum().reset_index()
sns.set_theme(style="ticks")
sns.set_context('notebook')
sns.lineplot(data = h41, x = 'mes', y = 'price',
markers = True, style = (1), legend = False,
color = 'black');
# -
# ### H5: Imóveis com 3 banheiros tem um crescimento mês após mês (MoM) de 15%.
#
# +
h5 = df[(df['bathrooms'] == 3)]
h5 = h5[['mes', 'price']].groupby('mes').sum().reset_index()
sns.lineplot(data = h5, x = 'mes', y = 'price', markers = True, style = (1), legend = False, color = 'black');
# -
# ### H6: Imóveis que nunca foram reformadas (yr_renovated == 0) são em média 20% mais baratos.
#
# +
h6 = df[['price', 'renovacao']].groupby('renovacao').mean().reset_index()
sns.barplot(h6['renovacao'], h6['price']);
# -
# ### H7: Imóveis em más condições mas com boa vista (view) são 10% mais caros.
#
# +
h71 = df[df['condition'] == 1]
h7 = df[['price', 'view']].groupby('view').sum().reset_index()
sns.barplot(h7['view'], h7['price']);
# -
# ### H8: Imóveis antigos e não renovados são 40% mais baratos.
#
# +
h8 = df[df['renovacao'] == 1]
h8 = df[['construcao', 'price']].groupby('construcao').sum().reset_index()
sns.barplot(x ='construcao', y = 'price', data = h8 )
# -
# ### H9: Imóveis com mais banheiros são em média 5% mais caros.
#
# +
h9 = df[['banheiro', 'price', 'sqft_lot']].groupby('banheiro').mean().reset_index()
sns.barplot(x = 'banheiro', y = 'price', data = h9);
# -
# ### H10: Imóveis renovados recentemente são 35% mais caros.
# +
h10 = df[['construcao', 'price']].groupby('construcao').mean().reset_index()
sns.barplot(x = 'construcao', y = 'price', data = h10);
# -
# # 3.0 Questão de negócio
#
#
# ## 3.1 Quais são os imóveis que a House Rocket deveria comprar e por qual preço?
# + deletable=false editable=false run_control={"frozen": true}
# df = df.copy()
# -
# ### 3.1.1 Quais casas
# +
a = df[['zipcode', 'price']].groupby('zipcode').median().reset_index()
df2 = pd.merge(a, df, on='zipcode', how = 'inner')
df2 = df2.rename(columns = {'price_y' : 'price', 'price_x' : 'price_median'} ) #alterando nome das colunas
for i, row in df2.iterrows():
if (row['price_median'] >= row['price']) & (row['condition'] < 3):
df2.loc[i,'pay'] = 'sim'
else:
df2.loc[i, 'pay'] = 'nao'
# +
#Casas para comprar
compra = df2[df2['pay'] == 'sim']
compra['pay'].shape[0]
compra[['id', 'price', 'pay']]
# +
fil_zip = widgets.Dropdown(options = df2['pay'].unique().tolist(),
description = 'regiao',
disabled = False)
def update_map (df3, fil_zip):
house = df2[(df2['pay'] == fil_zip)][['id', 'lat', 'long', 'condition', 'pay', 'price']]
mapa = px.scatter_mapbox(house, lat = 'lat', lon = 'long',
size = 'price',
color = 'condition', opacity = 0.5, zoom = 10 )
mapa.update_layout( mapbox_style = 'open-street-map')
mapa.update_layout( height = 600, margin = {'b':0, 't':0, 'r':0, 'l': 0})
mapa.show()
widgets.interactive( update_map, df3 = fixed(df2), fil_zip = fil_zip)
# -
# ## 3.2 Uma vez comprado, qual é o melhor momento para vendê-lo e por qual preço?
# +
df3 = df2.copy()
df3 = df3[df3['pay'] == 'sim']
df4 = df3[['season', 'zipcode', 'price']].groupby(['zipcode', 'season']).median().reset_index()
df4 = df4.rename(columns = {'price' : 'price_medi_season', 'season': 'season_median'} )
df5 = pd.merge(df3, df4, on='zipcode', how = 'inner')
for i, row in df5.iterrows():
if (row['price_medi_season'] > row['price']):
df5.loc[i, 'sale'] = row['price'] * 1.1
else:
df5.loc[i, 'sale'] = row['price'] * 1.3
# -
lista2 = df5['season'].tolist()
df5[['season', 'price', 'id', 'sale']]
a = df5[['price_medi_season', 'price', 'sale', 'price_median', 'season', 'zipcode']]
a
sns.barplot(data = a, x = 'season', y = 'sale');
# ### 3.2.1 Resultado do negócio
df5 ['lucro'] = df5['sale'] - df5['price']
df5['lucro'].sum()
# +
values = [['Hipótese','Parecer','Dizeres',],
['H1','Verdadeira','Imóveis com vista para água são mais caros',],
['H2','Falsa','Imóveis com data de construção não parece afetar o preço'],
['H3',
'Verdadeira',
'Imóveis sem porão possuem maior área total e são mais caros'],
['H4',
'Verdadeira',
'Imóveis sem reforma são mais baratos'],
['H5',
'Falso',
'Imóveis com más condições e com vista ruim são mais caros'],
['H6',
'Verdadeira',
'Imóveis antigos que não foram renovados são mais baratos'],
['H7',
'Falso',
'Imóveis que possuem entre 3 – 5 banheiros são mais caros'],
['H8',
'Falso',
'O tempo de renovação não parece influenciar no preço'],
['H9',
'Falso',
'Os imóveis sofrem decaimento do preço em relação aos meses' ],
['H10',
'Falso',
'Imóveis com 3 banheiros sofrem decaimento do preço em relação aos meses']
]
print(tabulate (values, headers = 'firstrow'))
|
Projeto-Insight.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import time
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn import preprocessing as pp
from sklearn.model_selection import StratifiedKFold
from sklearn.metrics import accuracy_score
from sklearn import preprocessing
import xgboost as xgb
from sklearn.ensemble import BaggingClassifier
import lightgbm as lgb
from sklearn.naive_bayes import GaussianNB
from sklearn import preprocessing as pp
from sklearn.neighbors import KNeighborsClassifier
from sklearn import tree
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import cross_val_predict
from statistics import mode
from sklearn.model_selection import cross_val_score, cross_validate, train_test_split
from sklearn.ensemble import RandomForestClassifier
from sklearn.neighbors import KNeighborsClassifier
import xgboost as xgb
import lightgbm as lgb
#Todas las librerías para los distintos algoritmos
from sklearn.naive_bayes import GaussianNB
from sklearn.naive_bayes import ComplementNB
from sklearn.naive_bayes import BernoulliNB
from sklearn.naive_bayes import MultinomialNB
from sklearn.calibration import CalibratedClassifierCV
from sklearn.svm import LinearSVC
from sklearn.svm import OneClassSVM
from sklearn.svm import SVC
from sklearn.svm import NuSVC
from sklearn.ensemble import RandomForestClassifier
from sklearn.neighbors import KNeighborsClassifier
import matplotlib.pyplot as plt
import sklearn.metrics as metrics
from sklearn.neural_network import MLPClassifier
from sklearn.ensemble import BaggingClassifier
import statistics
from sklearn.preprocessing import LabelEncoder
from sklearn.metrics import confusion_matrix
from sklearn.metrics import classification_report
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
from sklearn.decomposition import PCA
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
import warnings
from mlxtend.classifier import StackingClassifier
from mlxtend.classifier import StackingCVClassifier
from sklearn.neural_network import MLPClassifier
from sklearn.ensemble import GradientBoostingClassifier
from pylab import rcParams
from collections import Counter
warnings.simplefilter('ignore')
# +
data_train= pd.read_csv("./datos/train.csv",na_values=["?"])
data_test= pd.read_csv("./datos/test.csv",na_values=["?"])
data_trainCopia = data_train.copy()
data_testCopia = data_test.copy()
Nombre = LabelEncoder().fit(pd.read_csv("./datos/nombre.csv").Nombre)
Año = LabelEncoder().fit(pd.read_csv("./datos/ao.csv").Año)
Ciudad = LabelEncoder().fit(pd.read_csv("./datos/ciudad.csv").Ciudad)
Combustible = LabelEncoder().fit(pd.read_csv("./datos/combustible.csv").Combustible)
Consumo = LabelEncoder().fit(pd.read_csv("./datos/consumo.csv").Consumo)
Descuento = LabelEncoder().fit(pd.read_csv("./datos/descuento.csv").Descuento)
Kilometros = LabelEncoder().fit(pd.read_csv("./datos/kilometros.csv").Kilometros)
Mano = LabelEncoder().fit(pd.read_csv("./datos/mano.csv").Mano)
Potencia = LabelEncoder().fit(pd.read_csv("./datos/potencia.csv").Potencia)
Asientos = LabelEncoder().fit(pd.read_csv("./datos/asientos.csv").Asientos)
Motor_CC=LabelEncoder().fit(pd.read_csv("./datos/motor_cc.csv").Motor_CC)
Tipo_marchas=LabelEncoder().fit(pd.read_csv("./datos/Tipo_marchas.csv").Tipo_marchas)
data_trainCopia['Nombre']=data_trainCopia['Nombre'].fillna(mode(data_trainCopia['Nombre']))
data_trainCopia['Año']=data_trainCopia['Año'].fillna(mode(data_trainCopia['Año']))
data_trainCopia['Ciudad']=data_trainCopia['Ciudad'].fillna(mode(data_trainCopia['Ciudad']))
data_trainCopia['Kilometros']=data_trainCopia['Kilometros'].fillna(mode(data_trainCopia['Kilometros']))
data_trainCopia['Combustible']=data_trainCopia['Combustible'].fillna(mode(data_trainCopia['Combustible']))
data_trainCopia['Tipo_marchas']=data_trainCopia['Tipo_marchas'].fillna(mode(data_trainCopia['Tipo_marchas']))
data_trainCopia['Mano']=data_trainCopia['Mano'].fillna(mode(data_trainCopia['Mano']))
data_trainCopia['Consumo']=data_trainCopia['Consumo'].fillna(mode(data_trainCopia['Consumo']))
data_trainCopia['Motor_CC']=data_trainCopia['Motor_CC'].fillna(mode(data_trainCopia['Motor_CC']))
data_trainCopia['Potencia']=data_trainCopia['Potencia'].fillna(mode(data_trainCopia['Potencia']))
data_trainCopia['Asientos']=data_trainCopia['Asientos'].fillna(mode(data_trainCopia['Asientos']))
data_trainCopia['Descuento']=data_trainCopia['Descuento'].fillna(mode(data_trainCopia['Descuento']))
data_testCopia['Nombre']=data_testCopia['Nombre'].fillna(mode(data_testCopia['Nombre']))
data_testCopia['Año']=data_testCopia['Año'].fillna(mode(data_testCopia['Año']))
data_testCopia['Ciudad']=data_testCopia['Ciudad'].fillna(mode(data_testCopia['Ciudad']))
data_testCopia['Kilometros']=data_testCopia['Kilometros'].fillna(mode(data_testCopia['Kilometros']))
data_testCopia['Combustible']=data_testCopia['Combustible'].fillna(mode(data_testCopia['Combustible']))
data_testCopia['Tipo_marchas']=data_testCopia['Tipo_marchas'].fillna(mode(data_testCopia['Tipo_marchas']))
data_testCopia['Mano']=data_testCopia['Mano'].fillna(mode(data_testCopia['Mano']))
data_testCopia['Consumo']=data_testCopia['Consumo'].fillna(mode(data_testCopia['Consumo']))
data_testCopia['Motor_CC']=data_testCopia['Motor_CC'].fillna(mode(data_testCopia['Motor_CC']))
data_testCopia['Potencia']=data_testCopia['Potencia'].fillna(mode(data_testCopia['Potencia']))
data_testCopia['Asientos']=data_testCopia['Asientos'].fillna(mode(data_testCopia['Asientos']))
data_testCopia['Descuento']=data_testCopia['Descuento'].fillna(mode(data_testCopia['Descuento']))
#Eliminamos las columnas que no necesitamos
data_trainCopia=data_trainCopia.drop(['Descuento'], axis=1)
data_trainCopia=data_trainCopia.drop(['id'], axis=1)
data_testCopia=data_testCopia.drop(['Descuento'], axis=1)
data_testCopia=data_testCopia.drop(['id'], axis=1)
#Eliminamos los nan de los ids
data_trainCopia=data_trainCopia.dropna()
data_testCopia=data_testCopia.dropna()
#Codificación de las filas
data_trainCopia.Nombre = Nombre.transform(data_trainCopia.Nombre)
data_trainCopia.Año = Año.transform(data_trainCopia.Año)
data_trainCopia.Ciudad = Ciudad.transform(data_trainCopia.Ciudad)
data_trainCopia.Combustible = Combustible.transform(data_trainCopia.Combustible)
data_trainCopia.Potencia = Potencia.transform(data_trainCopia.Potencia)
data_trainCopia.Consumo = Consumo.transform(data_trainCopia.Consumo)
data_trainCopia.Kilometros = Kilometros.transform(data_trainCopia.Kilometros)
data_trainCopia.Mano = Mano.transform(data_trainCopia.Mano)
data_trainCopia.Motor_CC = Motor_CC.transform(data_trainCopia.Motor_CC)
data_trainCopia.Tipo_marchas = Tipo_marchas.transform(data_trainCopia.Tipo_marchas)
data_trainCopia.Asientos = Asientos.transform(data_trainCopia.Asientos)
#-------------------------------------------------------------------------------------------
data_testCopia.Nombre = Nombre.transform(data_testCopia.Nombre)
data_testCopia.Año = Año.transform(data_testCopia.Año)
data_testCopia.Ciudad = Ciudad.transform(data_testCopia.Ciudad)
data_testCopia.Combustible = Combustible.transform(data_testCopia.Combustible)
data_testCopia.Potencia = Potencia.transform(data_testCopia.Potencia)
data_testCopia.Consumo = Consumo.transform(data_testCopia.Consumo)
data_testCopia.Kilometros = Kilometros.transform(data_testCopia.Kilometros)
data_testCopia.Mano = Mano.transform(data_testCopia.Mano)
data_testCopia.Tipo_marchas = Tipo_marchas.transform(data_testCopia.Tipo_marchas)
data_testCopia.Asientos = Asientos.transform(data_testCopia.Asientos)
data_testCopia.Motor_CC = Motor_CC.transform(data_testCopia.Motor_CC)
target = pd.read_csv('./datos/precio_cat.csv')
target_train=data_trainCopia['Precio_cat']
data_trainCopia=data_trainCopia.drop(['Precio_cat'], axis=1)
data_trainCopia=data_trainCopia.astype(float)
data_testCopia=data_testCopia.astype(float)
data_testCopia_nor = (data_testCopia-data_trainCopia.mean(0))/data_trainCopia.std(0)
data_trainCopia_nor = (data_trainCopia-data_trainCopia.mean(0))/data_trainCopia.std(0)
atributos=data_trainCopia_nor[['Nombre','Ciudad', 'Año', 'Kilometros', 'Combustible','Tipo_marchas','Mano','Consumo','Motor_CC','Potencia', 'Asientos']]
from imblearn.over_sampling import SMOTE
Xo, yo = SMOTE().fit_resample(data_trainCopia_nor, target_train)
atributoYo=Xo[['Nombre','Ciudad', 'Año', 'Kilometros', 'Combustible','Tipo_marchas','Mano','Consumo','Motor_CC','Potencia', 'Asientos']]
# -
# +
lgbm1 = lgb.LGBMClassifier(learning_rate=0.055, objective='binary', n_estimators=740, n_jobs=2,
num_leaves=12, max_depth=-1,seed=46000)
scores = cross_val_score(lgbm1, Xo, yo, cv=5, scoring='accuracy')
print("Score Validacion Cruzada CON MODELO", np.mean(scores)*100)
lgbmEntrenado = lgbm1.fit(Xo, yo)
preLgbOVER = lgbmEntrenado.predict(data_testCopia_nor)
scores = cross_val_score(lgbmEntrenado, atributoYo, yo, cv=5, scoring='accuracy')
print("Score Validacion Cruzada CON MODELO", np.mean(scores)*100)
dfAux = pd.DataFrame({'id':data_test['id']})
dfAux.set_index('id', inplace=True)
dfFinal = pd.DataFrame({'id': data_test['id'], 'Precio_cat': preLgbOVER}, columns=['id', 'Precio_cat'])
dfFinal.set_index('id', inplace=True)
#dfFinal.to_csv("./soluciones/lgbmConDatosNormalizadosOverfiting9190.csv")
# -
# +
#SUBIDA DEL 29 QUE ME LLEGA A 0.79986
lgbm1 = lgb.LGBMClassifier(learning_rate=0.055, objective='binary', n_estimators=640, n_jobs=2,
num_leaves=20, max_depth=-1,seed=46000)
scores = cross_val_score(lgbm1, Xo, yo, cv=5, scoring='accuracy')
print("Score Validacion Cruzada CON MODELO", np.mean(scores)*100)
lgbmEntrenado = lgbm1.fit(Xo, yo)
preLgbOVER = lgbmEntrenado.predict(data_testCopia_nor)
scores = cross_val_score(lgbmEntrenado, atributoYo, yo, cv=5, scoring='accuracy')
print("Score Validacion Cruzada CON MODELO", np.mean(scores)*100)
dfAux = pd.DataFrame({'id':data_test['id']})
dfAux.set_index('id', inplace=True)
dfFinal = pd.DataFrame({'id': data_test['id'], 'Precio_cat': preLgbOVER}, columns=['id', 'Precio_cat'])
dfFinal.set_index('id', inplace=True)
dfFinal.to_csv("./soluciones/basura.csv")
# +
#CON ESTO SALE UN SCORE LOCAL DE 92.43
lgbm1 = lgb.LGBMClassifier(learning_rate=0.055, objective='binary', n_estimators=640, n_jobs=2,
num_leaves=22, max_depth=-1,seed=46000)
scores = cross_val_score(lgbm1, Xo, yo, cv=5, scoring='accuracy')
print("Score Validacion Cruzada CON MODELO", np.mean(scores)*100)
lgbmEntrenado = lgbm1.fit(Xo, yo)
preLgbOverSampling = lgbmEntrenado.predict(data_testCopia_nor)
scores = cross_val_score(lgbmEntrenado, atributoYo, yo, cv=5, scoring='accuracy')
print("Score Validacion Cruzada CON MODELO", np.mean(scores)*100)
dfAux = pd.DataFrame({'id':data_test['id']})
dfAux.set_index('id', inplace=True)
dfFinal = pd.DataFrame({'id': data_test['id'], 'Precio_cat': preLgbOverSampling}, columns=['id', 'Precio_cat'])
dfFinal.set_index('id', inplace=True)
#dfFinal.to_csv("./soluciones/LGBMOverSamplingDatosNormalizadosyscorelocal92_42.csv")
# +
#CON ESTO SALE UN SCORE LOCAL DE 92.42
lgbm1 = lgb.LGBMClassifier(learning_rate=0.055, objective='binary', n_estimators=620, n_jobs=2,
num_leaves=22, max_depth=-1,seed=46000)
scores = cross_val_score(lgbm1, Xo, yo, cv=5, scoring='accuracy')
print("Score Validacion Cruzada CON MODELO", np.mean(scores)*100)
lgbmEntrenado = lgbm1.fit(Xo, yo)
preLgbOVER = lgbmEntrenado.predict(data_testCopia_nor)
scores = cross_val_score(lgbmEntrenado, atributoYo, yo, cv=5, scoring='accuracy')
print("Score Validacion Cruzada CON MODELO", np.mean(scores)*100)
# +
#CON ESTO SALE UN SCORE LOCAL DE 92.56/
lgbm1 = lgb.LGBMClassifier(learning_rate=0.055, objective='binary', n_estimators=660, n_jobs=2,
num_leaves=20, max_depth=-1,seed=46000)
scores = cross_val_score(lgbm1, Xo, yo, cv=5, scoring='accuracy')
print("Score Validacion Cruzada CON MODELO", np.mean(scores)*100)
lgbmEntrenado = lgbm1.fit(Xo, yo)
preLgbOVER = lgbmEntrenado.predict(data_testCopia_nor)
scores = cross_val_score(lgbmEntrenado, atributoYo, yo, cv=5, scoring='accuracy')
print("Score Validacion Cruzada CON MODELO", np.mean(scores)*100)
# +
#CON ESTO SALE UN SCORE LOCAL DE 92.6369/
lgbm1 = lgb.LGBMClassifier(learning_rate=0.054, objective='binary', n_estimators=660, n_jobs=2,
num_leaves=20, max_depth=-1, seed=46000)
scores = cross_val_score(lgbm1, Xo, yo, cv=5, scoring='accuracy')
print("Score Validacion Cruzada CON MODELO", np.mean(scores)*100)
lgbmEntrenado = lgbm1.fit(Xo, yo)
preLgbOVER = lgbmEntrenado.predict(data_testCopia_nor)
scores = cross_val_score(lgbmEntrenado, atributoYo, yo, cv=5, scoring='accuracy')
print("Score Validacion Cruzada CON MODELO", np.mean(scores)*100)
# +
#CON ESTO SALE UN SCORE LOCAL DE 92.7105/
lgbm1 = lgb.LGBMClassifier(learning_rate=0.054, objective='binary', n_estimators=650, n_jobs=2,
num_leaves=20, max_depth=-1, seed=46000)
scores = cross_val_score(lgbm1, Xo, yo, cv=5, scoring='accuracy')
print("Score Validacion Cruzada CON MODELO", np.mean(scores)*100)
lgbmEntrenado = lgbm1.fit(Xo, yo)
preLgbOVER = lgbmEntrenado.predict(data_testCopia_nor)
scores = cross_val_score(lgbmEntrenado, atributoYo, yo, cv=5, scoring='accuracy')
print("Score Validacion Cruzada CON MODELO", np.mean(scores)*100)
# +
#CON ESTO SALE UN SCORE LOCAL DE 92.7197/
lgbm1 = lgb.LGBMClassifier(learning_rate=0.054, objective='binary', n_estimators=645, n_jobs=2,
num_leaves=20, max_depth=-1, seed=46000)
scores = cross_val_score(lgbm1, Xo, yo, cv=5, scoring='accuracy')
print("Score Validacion Cruzada CON MODELO", np.mean(scores)*100)
lgbmEntrenado = lgbm1.fit(Xo, yo)
preLgbOVER123 = lgbmEntrenado.predict(data_testCopia_nor)
scores = cross_val_score(lgbmEntrenado, atributoYo, yo, cv=5, scoring='accuracy')
print("Score Validacion Cruzada CON MODELO", np.mean(scores)*100)
dfAux = pd.DataFrame({'id':data_test['id']})
dfAux.set_index('id', inplace=True)
dfFinal = pd.DataFrame({'id': data_test['id'], 'Precio_cat': preLgbOVER123}, columns=['id', 'Precio_cat'])
dfFinal.set_index('id', inplace=True)
dfFinal.to_csv("./soluciones/LGBMOverSamplingDatosNormalizadosyscorelocal927197.csv")
# -
from imblearn.over_sampling import SMOTE
Xo, yo = SMOTE().fit_resample(data_trainCopia_nor, target_train)
atributoYo=Xo[['Nombre','Ciudad', 'Año', 'Kilometros', 'Combustible','Tipo_marchas','Mano','Consumo','Motor_CC','Potencia', 'Asientos']]
# +
#SUBIDA DEL 29 QUE ME LLEGA A 0.79986 con un score de 92,37 en local
lgbm1 = lgb.LGBMClassifier(learning_rate=0.055, objective='binary', n_estimators=640, n_jobs=2,
num_leaves=20, max_depth=-1,seed=46000)
scores = cross_val_score(lgbm1, Xo, yo, cv=5, scoring='accuracy')
print("Score Validacion Cruzada CON MODELO", np.mean(scores)*100)
lgbmEntrenado = lgbm1.fit(Xo, yo)
preLgbOVER = lgbmEntrenado.predict(data_testCopia_nor)
scores = cross_val_score(lgbmEntrenado, atributoYo, yo, cv=5, scoring='accuracy')
print("Score Validacion Cruzada CON MODELO", np.mean(scores)*100)
# +
#CON ESTO SALE UN SCORE LOCAL DE 92.7197/
lgbm1 = lgb.LGBMClassifier(learning_rate=0.054, objective='binary', n_estimators=645, n_jobs=2,
num_leaves=20, max_depth=-1, seed=46000)
scores = cross_val_score(lgbm1, Xo, yo, cv=5, scoring='accuracy')
print("Score Validacion Cruzada CON MODELO", np.mean(scores)*100)
lgbmEntrenado = lgbm1.fit(Xo, yo)
preLgbOVER123 = lgbmEntrenado.predict(data_testCopia_nor)
scores = cross_val_score(lgbmEntrenado, atributoYo, yo, cv=5, scoring='accuracy')
print("Score Validacion Cruzada CON MODELO", np.mean(scores)*100)
# +
#CON ESTO SALE UN SCORE LOCAL DE 92.74/
lgbm1 = lgb.LGBMClassifier(learning_rate=0.054, objective='binary', n_estimators=645, n_jobs=2,
num_leaves=20, max_depth=-1, seed=46000, reg_alpha=0.3)
scores = cross_val_score(lgbm1, Xo, yo, cv=5, scoring='accuracy')
print("Score Validacion Cruzada CON MODELO", np.mean(scores)*100)
lgbmEntrenado = lgbm1.fit(Xo, yo)
preLgbOVER123 = lgbmEntrenado.predict(data_testCopia_nor)
scores = cross_val_score(lgbmEntrenado, atributoYo, yo, cv=5, scoring='accuracy')
print("Score Validacion Cruzada CON MODELO", np.mean(scores)*100)
|
P3-Kaggle-Clasificacion/P3/.ipynb_checkpoints/prueba19Final-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# cd /Users/martin/Git/estates
import os
import boto3
import json
from decimal import Decimal
import pandas as pd
file = 'data/bronze/20210703-104535_1038932572.json'
with open(file) as f:
data = json.load(f)
columns = [
'estate_id',
'created_at',
'expires_at',
'estate_title',
'estate_description_short',
'estate_description_long',
'estate_locality_district',
'estate_disposition',
'estate_category_main_cb',
'estate_longitude',
'estate_latitude',
'estate_map_zoom',
'estate_images',
'seller_ico',
'seller_email',
'seller_numbers',
'seller_web',
'seller_address',
'seller_name',
'<NAME>',
'Užitná plocha',
'Podlaží',
'Stavba',
'Stav objektu',
'Poznámka k ceně',
'Energetická náročnost budovy',
'Vlastnictví',
'Vybavení',
'Výtah',
]
data
data = {
_id: {item: value for item, value in items.items() if item in columns}
for _id, items in data.items()
}
ddb_data = json.loads(json.dumps(data), parse_float=Decimal)
# +
dynamodb = boto3.resource('dynamodb')
table = dynamodb.Table('estates')
with table.batch_writer() as batch:
for _id, items in ddb_data.items():
batch.put_item(
Item={
'estate_id': _id,
'items': items
}
)
# -
|
notebooks/01_dynamodb.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Comparison star generator
# ## For information about your TESS target
#
# 1. Go to https://exofop.ipac.caltech.edu/tess/
# 2. Enter the TIC number in the TIC search box
# 2. Use the RA/Dec for the `ra` and `dec` variables in the cell below.
# 3. Use the "TESS mag" for the variable `Cmag` in the cell below.
# 4. Use the "depth" for the variable `depth` in the cell below.
# +
import functools
import re
from pathlib import Path
import requests
import numpy as np
from astropy import units as u
from astropy.table import Table
from astropy.coordinates import SkyCoord
from stellarphot.visualization.comparison_functions import (
read_file, set_up, match, mag_scale, in_field, wrap, make_markers,
viewer
)
from stellarphot.visualization.seeing_profile_functions import set_keybindings
# -
# ## Change the settings in this cell as needed
# +
directory_with_images = '.'
#This could be any image from roughly the middle of the sequence.
sample_image_for_finding_stars = 'TIC_470127886.01-S001-R001-C001-rp.fit'
# This name should be one of the "aliases" for the object if it is a TIC object
object_of_interest = 'TYC 4612-401-1'
# Get these variable from ExoFOP-TESS or Exoplanet ETD
ra_object = '22:04:28.27'
dec_object = '81:33:57.42'
Cmag = 10.76 # Use TESS magnitude for this
depth = 13280 / 1000 # Depth should be in parts per thousand, ExoFOP gives it in ppm.
aperture_output_file = 'aperture_locations.fits'
TESS_target = False # change to False (capitalized) if this is NOT a TESS object
if TESS_target:
server = "https://www.astro.louisville.edu"
gaia_aperture_url = server + f"/cgi-bin/gaia_to_aij/upload_request.cgi?ra={ra_object}&dec={dec_object}&mag={Cmag}&depth={depth}"
result = requests.get(gaia_aperture_url)
links = re.search('href="(.+)"', result.text.replace('\n', ''), )
target_file_contents = requests.get(server + links[1])
with open('gaia_stars.radec', 'wt') as f:
f.write(target_file_contents.text)
targets_from_file = read_file('gaia_stars.radec')
else:
targets_from_file = []
# -
coordinate = SkyCoord(ra=ra_object, dec=dec_object, unit=("hour", "degree"))
# +
ccd, vsx = \
set_up(sample_image_for_finding_stars,
directory_with_images=directory_with_images
)
apass, vsx_apass_angle, targets_apass_angle = match(ccd, targets_from_file, vsx)
apass_good_coord, good_stars = mag_scale(Cmag, apass, vsx_apass_angle,
targets_apass_angle,
brighter_dmag=3,
dimmer_dmag=2)
apass_comps = in_field(apass_good_coord, ccd, apass, good_stars)
box, iw = viewer()
make_markers(iw, ccd, targets_from_file, vsx, apass_comps,
name_or_coord=coordinate)
box
# -
# ### Get the table of marked stars.
all_table = iw.get_markers(marker_name='all')
elims = np.array([name.startswith('elim')
for name in all_table['marker name']])
elim_table = all_table[elims]
comp_table = all_table[~elims]
# ### Remove any that are marked for elimination as comparisons.
index, d2d, d3d = elim_table['coord'].match_to_catalog_sky(comp_table['coord'])
comp_table.remove_rows(index)
# ### Sort the table
# +
if targets_from_file:
target_coord = targets_from_file['coords'][0]
else:
target_coord = SkyCoord(ra=ra_object, dec=dec_object, unit=("hour", "degree"))
# Calculate how far each is from target
comp_table['separation'] = target_coord.separation(comp_table['coord'])
# Add dummy column for sorting in the order we want
comp_table['sort'] = np.zeros(len(comp_table))
# Set sort order
apass_mark = comp_table['marker name'] == 'APASS comparison'
vsx_mark = comp_table['marker name'] == 'VSX'
tess_mark = ((comp_table['marker name'] == 'TESS Targets') |
(comp_table['separation'] < 0.3 * u.arcsec))
comp_table['sort'][apass_mark] = 2
comp_table['sort'][vsx_mark] = 1
comp_table['sort'][tess_mark] = 0
# Ensure the target is always first
# Sort the table
comp_table.sort(['sort', 'separation'])
# Assign the IDs
comp_table['id'] = range(1, len(comp_table) + 1)
# -
# ### Display information on variables
new_vsx_mark = comp_table['marker name'] == 'VSX'
idx, _, _ = comp_table['coord'][new_vsx_mark].match_to_catalog_sky(vsx['coords'])
for our_name, vsx_name in zip(comp_table['id'][new_vsx_mark], vsx['Name'][idx]):
print(f'Our id number: {our_name}, VSX name: {vsx_name}')
# ### Label the stars with numbers
for star in comp_table:
star_id = star['id']
if star['marker name'] == 'TESS Targets':
iw._marker = functools.partial(iw.dc.Text, text=f'T{star_id}', fontsize=20, fontscale=False, color='green')
iw.add_markers(Table(data=[[star['x']+20], [star['y']-20]], names=['x', 'y']))
elif star['marker name'] == 'APASS comparison':
iw._marker = functools.partial(iw.dc.Text, text=f'C{star_id}', fontsize=20, fontscale=False, color='red')
iw.add_markers(Table(data=[[star['x']+20], [star['y']-20]], names=['x', 'y']))
elif star['marker name'] == 'VSX':
iw._marker = functools.partial(iw.dc.Text, text=f'V{star_id}', fontsize=20, fontscale=False, color='blue')
iw.add_markers(Table(data=[[star['x']+20], [star['y']-20]], names=['x', 'y']))
else:
print(f"Unrecognized marker name: {star['marker name']}")
comp_table.write(aperture_output_file, overwrite=True)
# ### Add 2.5 arcmin circle (do this after writing comparison table!)
#
# Otherwise the comparison table will contain an extra entry for this circle.
if targets_from_file:
target = targets_from_file[0]
iw.marker = {'color': 'yellow', 'radius': 268, 'type': 'circle'}
iw.add_markers(target, skycoord_colname='coords',
use_skycoord=True, marker_name='target')
|
stellarphot/notebooks/comp-stars-template.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Data Science)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:us-east-2:429704687514:image/datascience-1.0
# ---
# # Orchestrating Jobs, Model Registration, and Continuous Deployment with Amazon SageMaker
#
# Amazon SageMaker offers Machine Learning application developers and Machine Learning operations engineers the ability to orchestrate SageMaker jobs and author reproducible Machine Learning pipelines, deploy custom-build models for inference in real-time with low latency or offline inferences with Batch Transform, and track lineage of artifacts. You can institute sound operational practices in deploying and monitoring production workflows, deployment of model artifacts, and track artifact lineage through a simple interface, adhering to safety and best-practice paradigmsfor Machine Learning application development.
#
# The SageMaker Pipelines service supports a SageMaker Machine Learning Pipeline Domain Specific Language (DSL), which is a declarative Json specification. This DSL defines a Directed Acyclic Graph (DAG) of pipeline parameters and SageMaker job steps. The SageMaker Python Software Developer Kit (SDK) streamlines the generation of the pipeline DSL using constructs that are already familiar to engineers and scientists alike.
#
# The SageMaker Model Registry is where trained models are stored, versioned, and managed. Data Scientists and Machine Learning Engineers can compare model versions, approve models for deployment, and deploy models from different AWS accounts, all from a single Model Registry. SageMaker enables customers to follow the best practices with ML Ops and getting started right. Customers are able to standup a full ML Ops end-to-end system with a single API call.
# ## SageMaker Pipelines
#
# Amazon SageMaker Pipelines support the following activites:
#
# * Pipelines - A Directed Acyclic Graph of steps and conditions to orchestrate SageMaker jobs and resource creation.
# * Processing Job steps - A simplified, managed experience on SageMaker to run data processing workloads, such as feature engineering, data validation, model evaluation, and model interpretation.
# * Training Job steps - An iterative process that teaches a model to make predictions by presenting examples from a training dataset.
# * Conditional step execution - Provides conditional execution of branches in a pipeline.
# * Registering Models - Creates a model package resource in the Model Registry that can be used to create deployable models in Amazon SageMaker.
# * Creating Model steps - Create a model for use in transform steps or later publication as an endpoint.
# * Parameterized Pipeline executions - Allows pipeline executions to vary by supplied parameters.
# * Transform Job steps - A batch transform to preprocess datasets to remove noise or bias that interferes with training or inference from your dataset, get inferences from large datasets, and run inference when you don't need a persistent endpoint.
# ## Layout of the SageMaker ModelBuild Project Template
#
# The template provides a starting point for bringing your SageMaker Pipeline development to production.
#
# ```
# |-- codebuild-buildspec.yml
# |-- CONTRIBUTING.md
# |-- pipelines
# | |-- abalone
# | | |-- evaluate.py
# | | |-- __init__.py
# | | |-- pipeline.py
# | | `-- preprocess.py
# | |-- get_pipeline_definition.py
# | |-- __init__.py
# | |-- run_pipeline.py
# | |-- _utils.py
# | `-- __version__.py
# |-- README.md
# |-- sagemaker-pipelines-project.ipynb
# |-- setup.cfg
# |-- setup.py
# |-- tests
# | `-- test_pipelines.py
# `-- tox.ini
# ```
# A description of some of the artifacts is provided below:
# <br/><br/>
# Your codebuild execution instructions:
# ```
# |-- codebuild-buildspec.yml
# ```
# <br/><br/>
# Your pipeline artifacts, which includes a pipeline module defining the required `get_pipeline` method that returns an instance of a SageMaker pipeline, a preprocessing script that is used in feature engineering, and a model evaluation script to measure the Mean Squared Error of the model that's trained by the pipeline:
#
# ```
# |-- pipelines
# | |-- abalone
# | | |-- evaluate.py
# | | |-- __init__.py
# | | |-- pipeline.py
# | | `-- preprocess.py
#
# ```
# <br/><br/>
# Utility modules for getting pipeline definition jsons and running pipelines:
#
# ```
# |-- pipelines
# | |-- get_pipeline_definition.py
# | |-- __init__.py
# | |-- run_pipeline.py
# | |-- _utils.py
# | `-- __version__.py
# ```
# <br/><br/>
# Python package artifacts:
# ```
# |-- setup.cfg
# |-- setup.py
# ```
# <br/><br/>
# A stubbed testing module for testing your pipeline as you develop:
# ```
# |-- tests
# | `-- test_pipelines.py
# ```
# <br/><br/>
# The `tox` testing framework configuration:
# ```
# `-- tox.ini
# ```
# ### A SageMaker Pipeline
#
# The pipeline that we create follows a typical Machine Learning Application pattern of pre-processing, training, evaluation, and conditional model registration and publication, if the quality of the model is sufficient.
#
# 
#
# ### Getting some constants
#
# We get some constants from the local execution environment.
# +
import boto3
import sagemaker
region = boto3.Session().region_name
role = sagemaker.get_execution_role()
default_bucket = sagemaker.session.Session().default_bucket()
# Change these to reflect your project/business name or if you want to separate ModelPackageGroup/Pipeline from the rest of your team
model_package_group_name = f"AbaloneModelPackageGroup-Example"
pipeline_name = f"AbalonePipeline-Example"
# -
# ### Get the pipeline instance
#
# Here we get the pipeline instance from your pipeline module so that we can work with it.
# +
from pipelines.abalone.pipeline import get_pipeline
pipeline = get_pipeline(
region=region,
role=role,
default_bucket=default_bucket,
model_package_group_name=model_package_group_name,
pipeline_name=pipeline_name,
)
# -
# ### Submit the pipeline to SageMaker and start execution
#
# Let's submit our pipeline definition to the workflow service. The role passed in will be used by the workflow service to create all the jobs defined in the steps.
pipeline.upsert(role_arn=role)
# We'll start the pipeline, accepting all the default parameters.
#
# Values can also be passed into these pipeline parameters on starting of the pipeline, and will be covered later.
execution = pipeline.start()
# ### Pipeline Operations: examining and waiting for pipeline execution
#
# Now we describe execution instance and list the steps in the execution to find out more about the execution.
execution.describe()
# We can wait for the execution by invoking `wait()` on the execution:
execution.wait()
# We can list the execution steps to check out the status and artifacts:
execution.list_steps()
# ### Parameterized Executions
#
# We can run additional executions of the pipeline specifying different pipeline parameters. The parameters argument is a dictionary whose names are the parameter names, and whose values are the primitive values to use as overrides of the defaults.
#
# Of particular note, based on the performance of the model, we may want to kick off another pipeline execution, but this time on a compute-optimized instance type and set the model approval status automatically be "Approved". This means that the model package version generated by the `RegisterModel` step will automatically be ready for deployment through CI/CD pipelines, such as with SageMaker Projects.
execution = pipeline.start(
parameters=dict(
ProcessingInstanceType="ml.c5.xlarge",
ModelApprovalStatus="Approved",
)
)
execution.wait()
execution.list_steps()
|
mlops-template-gitlab/seedcode/mlops-gitlab-project-seedcode-model-build/sagemaker-pipelines-project.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Animations with Adampy
#
# By creating animations we can easily see at the evolution of an atmospheric event. As an example we can create an animation of the Saharan dust plume that reach Austria at the end of October 2018 by using the CAMS european PM10 model.
#
# With Adampy we can easily create animations that can be later displayed or embeded into a website or publication.
import geopandas as gpd
import numpy as np
from datetime import datetime, timedelta, date
import matplotlib.pyplot as plt
import adampy as adam
from IPython.display import Image
start_date = date(2018, 10, 15)
end_date = date(2018, 11, 15)
#collection = 'Z_CAMS_C_ECMF_TCNO2'
#collection = 'Z_CAMS_C_ECMF_TCSO2'
#collection = 'Z_CAMS_C_ECMF_GTCO3'
#collection = 'Z_CAMS_C_ECMF_PM10'
collection = 'EU_CAMS_SURFACE_PM10'
gif = adam.getAnimation('wcs.top-platform.eu', collection, start_date, end_date, frame_duration = 0.5, legend = False).get_data()
Image(url=gif)
|
notebooks/phiweek-2019/4. Animation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {}, "report_default": {"hidden": true}}}}
# # A dashboard to explore world population
#
# The Python API, along with the [Jupyter Dashboard](http://jupyter-dashboards-layout.readthedocs.io/) project enables Python developers to quickly build and prototype interactive web apps. This sample illustrates one such app which can be used to detect the changes in vegetation between the two dates. Increases in vegetation are shown in green, and decreases are shown in magenta.
#
# This sample uses the fast on-the-fly processing power of raster functions available in the `raster` module of the Python API.
#
# <blockquote>To run this sample you need `jupyter_dashboards` package in your conda environment. You can install it as shown below. For information on this, [refer to the install instructions](http://jupyter-dashboards-layout.readthedocs.io/en/latest/getting-started.html#installing-and-enabling)</blockquote>
#
# conda install jupyter_dashboards -c conda-forge
#
# This Dashboard uses the Geoenrichment API to alow users to pan around the world and explore the population of a selected AOI. Let's start by logging into ArcGIS Online and importing packages.
# -
# <img src="../../static/img/04_app_01.gif">
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {"hidden": true}}}}
from IPython.display import display, clear_output
import ipywidgets as widgets
from arcgis.features import SpatialDataFrame
from arcgis.geoenrichment import enrich
from arcgis.raster import ImageryLayer
from arcgis.geometry import Geometry
from arcgis.gis import GIS
gis = GIS(profile="your_online_profile")
img_svc ='https://landscape7.arcgis.com/arcgis/rest/services/World_Population_Density_Estimate_2016/ImageServer/'
img_lyr = ImageryLayer(img_svc, gis=gis)
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 8, "height": 4, "hidden": false, "row": 24, "width": 4}, "report_default": {"hidden": false}}}} language="html"
# <style>
# .intro {
# padding: 10px;
# color: #202020;
# font-family: 'Helvetica'
# }
# .map {
# border: solid;
# height: 450;
# }
# </style>
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"hidden": true}, "report_default": {"hidden": true}}}}
# run_btn on_click
def running(button_object):
viz_map.clear_graphics()
the_geom = Geometry(viz_map.extent)
enriched = SpatialDataFrame(enrich([the_geom], gis=gis))
if 'TOTPOP' not in list(enriched):
enriched_pop = -1
else:
enriched_pop = enriched.TOTPOP[0]
print('the Geom: {}'.format(the_geom))
print('Sample Count: {}'.format(img_lyr.properties.maxImageHeight))
samples = img_lyr.get_samples(the_geom)
sample_sum = sum([float(sample['value']) for sample in samples])
print('GeoEnrichment Population Results: {}'.format(enriched_pop))
title = 'GeoEnriched Cell'
for feature in enriched.to_featureset():
# Expand Attributes With New Lines
content = "{}".format(
'<br/>'.join(['%s: %s' % (key.upper(), value) for (key, value) in feature.attributes.items()])
)
# Add Cell to Map
viz_map.draw(feature.geometry, popup={'title': title, 'content': content})
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 24, "hidden": false, "row": 0, "width": null}, "report_default": {"hidden": false}}}}
# Create & Display Map
viz_map = gis.map('St Louis')
viz_map.add_layer(img_lyr)
display(viz_map)
# Create Submit Button & Set on_click
run_btn = widgets.Button(
description='Fetch Extent',
button_style='success',
tooltip='Collect GeoEnrichment for Extent',
layout=widgets.Layout(justify_content='center', margin='0px 0px 0px 10px')
)
run_btn.on_click(running)
# Handle Widget Layout
params = widgets.HBox(
[run_btn],
layout=widgets.Layout(justify_content='center', margin='10px')
)
display(params)
|
samples/02_power_users_developers/population_exploration_dashboard.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="WoAJAn5SAkY0" colab_type="text"
# ## Rare Labels
#
# - Values present for a small percentage
#
# - Usually present less than 5%
#
# - Concept of cardinality
#
# ## Rare label consequences
#
# - May add information in low cardinality
#
# - May add noise is high cardinality
#
#
# ### Engineering Rare Labels
#
# - Replacing by most frequent label
# - Grouping all rare labels together
#
# Categorical variables can have:
#
# - One predominant category
# - A small number of categories
# - High cardinality
#
#
# + id="42hbGwCeDd8-" colab_type="code" colab={}
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
# + id="Ds9gl_oFEATI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="57d6aed5-1dba-445a-e718-b00288fdf4a7"
from google.colab import drive
drive.mount('/content/gdrive')
data = pd.read_csv("gdrive/My Drive/Colab Notebooks/FeatureEngineering/trainh.csv")
# + id="3Jyw7xjcAkY6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 251} outputId="536e5b75-3bbe-47a2-b6f5-85fb891d96b3"
data.head()
# + id="9mQS3wfemry3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 329} outputId="5a7b27ea-ff70-46dd-8e67-8bd3e3c50def"
data.columns
# + id="3td36aFVAkZT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 763} outputId="0a32ffa2-e706-46e4-c984-11cd9ab62427"
# get number of categories in variables
categoricals = []
for col in data.columns:
if data[col].dtypes =='O':
print('{} categories : {} '.format(col, len(data[col].unique())))
categoricals.append(col)
# + id="8EdOnRtkp503" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 104} outputId="03013e91-68b8-4722-dda5-7ae6707b5ff8"
# Get variables with more than n categories
n = 8
cats = []
for col in data.columns:
if data[col].dtypes =='O':
if len(data[col].unique())>n:
print('{} categories : {} '.format(col, len(data[col].unique())))
cats.append(col)
# + id="6mL3xO1WqmQh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="5dc8277a-ac15-4bc7-a1ab-635fb4157989"
for col in cats:
if data[col].dtypes =='O': # if the variable is categorical
print(100*data.groupby(col)[col].count()/np.float(len(data)))
print()
# + id="nmQVTJM8AkY9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="6da6642e-be1e-4d82-b3e2-ba77b2592074"
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(data[cats], data.SalePrice,
test_size=0.2)
X_train.shape, X_test.shape
# + id="uyjA7GUWAkZE" colab_type="code" colab={}
def label_encoder(X_train, X_test, columns, na_flag = False):
import random
for col in columns:
mapper = {k:i for i, k in enumerate(X_train[col].unique(), 0)}
if na_flag:
mapper[np.nan] = np.nan
X_train.loc[:, col] = X_train.loc[:, col].map(mapper)
X_test.loc[:, col] = X_test.loc[:, col].map(mapper)
X_test[col] = X_test[col].fillna(random.choice(list(mapper.values())))
# + id="GwhzY2lyxBlQ" colab_type="code" colab={}
label_encoder(X_train, X_test, cats)
# + id="b4s6p7We4Iwt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 121} outputId="53045764-f344-475e-dc88-14933d1364fa"
X_train.isnull().sum()
# + id="1tEW_WpG4LWz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 121} outputId="92162fbb-f790-4da0-d588-a4653430de9d"
X_test.isnull().sum()
# + id="1VD9uGRxw7hs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="b2d17664-bbea-4941-ecb1-ccd9df6faf4c"
sns.set()
for i in cats:
plt.figure()
sns.distplot(X_train[i], kde=False)
# + id="8BuvhTmNxWzx" colab_type="code" colab={}
def new_label_imputation(Xtrain, Xtest, threshold, cats):
X_train, X_test = Xtrain.copy(), Xtest.copy()
for col in cats:
rows = len(X_train)
temp_df = pd.Series(100*X_train[col].value_counts() / rows)
nonrares = temp_df[temp_df>=threshold].index # non-rare labels
X_train[col] = np.where(Xtrain[col].isin(nonrares), Xtrain[col], 'rare')
X_test[col] = np.where(Xtest[col].isin(nonrares), Xtest[col], 'rare')
return X_train, X_test
# + id="cQ0nEWrAzbxZ" colab_type="code" colab={}
X_train_rare, X_test_rare = new_label_imputation(X_train, X_test, 10, cats)
# + id="XKkiQGztzKLK" colab_type="code" colab={}
label_encoder(X_train_rare, X_test_rare, cats)
# + id="2oyAs30YzCMJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="31dad17e-f63a-4c2f-a582-7741d4b4fe22"
sns.set()
for i in cats:
fig, ax = plt.subplots(1,2, figsize=(10,5))
sns.distplot(X_train[i], kde=False, ax=ax[0])
sns.distplot(X_train_rare[i], kde=False, ax=ax[1])
# + id="SMphyIVtyW-q" colab_type="code" colab={}
def frequent_imputation(Xtrain, Xtest, threshold, cats):
X_train, X_test = Xtrain.copy(), Xtest.copy()
for col in cats:
rows = len(X_train)
temp_df = pd.Series(100*X_train[col].value_counts() / rows)
nonrares = temp_df[temp_df>=threshold].index # non-rare labels
frequent_cat = X_train.groupby(col)[col].count().sort_values().tail(1).index.values[0]
X_train[col] = np.where(Xtrain[col].isin(nonrares), Xtrain[col], frequent_cat)
X_test[col] = np.where(Xtest[col].isin(nonrares), Xtest[col], frequent_cat)
return X_train, X_test
# + id="uHHwSQUl1xxq" colab_type="code" colab={}
X_train_freq, X_test_freq = frequent_imputation(X_train, X_test, 10, cats)
# + id="sL_u2ks318Pj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="97749e3a-c469-4485-902c-2050a193933c"
sns.set()
for i in cats:
fig, ax = plt.subplots(1,2, figsize=(10,5))
sns.distplot(X_train[i], kde=False, ax=ax[0])
sns.distplot(X_train_freq[i], kde=False, ax=ax[1])
# + id="LoMzlnkhAkZR" colab_type="code" colab={}
def regressor(X_train, y_train, X_test, y_test, cols, model):
from sklearn.metrics import mean_squared_error
model.fit(X_train[cols],y_train)
y_pred = model.predict(X_test[cols])
print(mean_squared_error(y_test, y_pred))
# + colab_type="code" id="Q6fHdmXlrcRo" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="e08e0fe9-7d3b-4c7a-926b-d0c42cbeab50"
from sklearn.linear_model import LinearRegression
model = LinearRegression()
regressor(X_train_rare, y_train, X_test_rare, y_test, cats, model)
regressor(X_train_freq, y_train, X_test_freq, y_test, cats, model)
# + colab_type="code" id="rf2qSlwzrcRs" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="e41f398f-1bb2-4c43-f014-2b5b40b0fcb9"
from sklearn.linear_model import RidgeCV
model = RidgeCV()
regressor(X_train, y_train, X_test, y_test, cats, model)
# + colab_type="code" id="1fl4DQLCrcRv" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="06544dc6-f8e0-48c1-f37e-11df03d0569c"
from sklearn.linear_model import Ridge
model = RidgeCV()
regressor(X_train, y_train, X_test, y_test, cats, model)
# + colab_type="code" id="2XSmYB49rcRy" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="252a8d4c-7f3a-4ae8-ad22-7bb126c9a3dc"
from sklearn.svm import SVR
model = SVR()
regressor(X_train, y_train, X_test, y_test, cats, model)
# + colab_type="code" id="nDWv80igrcR2" colab={"base_uri": "https://localhost:8080/", "height": 89} outputId="69ba2e4e-66da-4984-a5af-c903b3507214"
from sklearn.neural_network import MLPRegressor
model = MLPRegressor()
regressor(X_train, y_train, X_test, y_test, cats, model)
# + colab_type="code" id="ZtS9HIE8rcR5" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="ebe096b1-3fe6-4ade-f3c6-9d15f69c5375"
from sklearn.svm import LinearSVR
model = LinearSVR()
regressor(X_train, y_train, X_test, y_test, cats, model)
# + colab_type="code" id="nNYoHOujrcR7" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="4eb6cb61-ea1e-4c99-e943-b68ea8a14987"
from sklearn.ensemble import RandomForestRegressor
model = RandomForestRegressor()
regressor(X_train, y_train, X_test, y_test, cats, model)
# + colab_type="code" id="9Uc-0cjMrcR9" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="1017fa16-0c40-42e4-837f-a23a0c8487dd"
from sklearn.tree import DecisionTreeRegressor
model = DecisionTreeRegressor()
regressor(X_train, y_train, X_test, y_test, cats, model)
# + colab_type="code" id="_u07tym7rcR_" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="87def5e0-0c67-4bd6-b918-62a1e3729845"
from sklearn.linear_model import SGDRegressor
model = SGDRegressor()
regressor(X_train, y_train, X_test, y_test, cats, model)
|
FeatureEngineering_DataScience/Demo181_RareCategories_SomeCategories.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import numpy as np
import os
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
# %matplotlib inline
from astropy import table
from astropy.table import Table
from astropy.io import ascii
from tabulate import tabulate
import pyspherematch as sm
from astropy import units as u
from astropy.coordinates import SkyCoord, FK4, FK5
SAGA_DIR = os.environ['SAGA_DIR']
SAGA_DROPBOX= os.environ['SAGA_DROPBOX']
#import pandas as pd
from scipy.stats import spearmanr
# -
from palettable.colorbrewer.qualitative import Dark2_8
if 'plt' in locals() and hasattr(plt, 'rcParams'):
plt.rcParams['lines.linewidth'] = 2.5
plt.rcParams['font.size'] = 16.0
plt.rcParams['font.weight'] = 'medium'
plt.rcParams['font.family'] = 'serif'
plt.rcParams['axes.prop_cycle'] = plt.cycler(color=Dark2_8.mpl_colors)
plt.rcParams['legend.fontsize'] = 'medium'
plt.rcParams['legend.frameon'] = False
plt.rcParams['figure.dpi'] = 100
plt.rcParams['figure.figsize'] = 7, 6
plt.rcParams['xtick.major.size'] = 6
plt.rcParams['xtick.minor.size'] = 4
plt.rcParams['ytick.major.size'] = 6
plt.rcParams['ytick.minor.size'] = 4
# READ SUBMASTER LIST
master = Table.read(SAGA_DROPBOX+'hosts/submaster.ecsv.gz', format='ascii.ecsv')
# +
file = SAGA_DIR + '/data/sats.txt'
sats = ascii.read(file)
hid = np.in1d(master['NSAID'],sats['NSAID'])
hosts = master[hid]
for obj in hosts:
print obj['NSAID']
# +
nsa = Table.read (SAGA_DIR+'/cats/nsa_v0_1_3.fits')
nids = np.in1d(nsa['NSAID'],sats['NSAID'])
mass = nsa['MASS'][nids]
log_SM = np.log10(mass/0.7**2)
# +
fig = plt.subplots(figsize=(6,8), sharex=True)
plt.rcParams['font.size'] = 13.0
MWsats = 5
M31sats = 9
ns = sats['Ncompl']
np.append(ns,MWsats)
ylim = 11.5
yl = 10.25
# MK
ax = plt.subplot2grid((3,2), (0, 0))
ax.set_ylim(0, ylim)
ax.plot(hosts['M_K'],sats['Nsat'],'ko')
ax.plot(hosts['M_K'],sats['Ncompl'],'ko',markerfacecolor='none')
ax.plot([-24],[MWsats],'y*',color='#ff8c00',markersize=11)
ax.plot([-24.51-0.188],[M31sats],'*',color='#ba55d3',markersize=11) # hammer 2007
ax.xaxis.set_major_locator(ticker.MultipleLocator(0.5))
ax.set_ylabel('N$_{\mathrm{sats}}$')
ax.set_xlabel('$M_K$')
hp = hosts['M_K']
np.ma.append(hp,-24)
sr = spearmanr(hp,ns)
t = '$M_K$: ($r_s$={:04.2f}, p={:04.2f})'.format(sr[0],sr[1])
ax.text(-24.4,yl,t,fontsize=8)
print sr[1]
# Mr
ax = plt.subplot2grid((3,2), (0, 1))
ax.set_ylim(0, ylim)
ax.set_xlim(-22.15,-20.8)
ax.plot(hosts['M_r'],sats['Nsat'],'ko')
ax.plot(hosts['M_r'],sats['Ncompl'],'ko',markerfacecolor='none')
#ax.plot([-21. + 5.*np.log10(0.7)],[MWsats],'y*',color='#ff8c00',markersize=11)
ax.plot([-20.7 + 5.*np.log10(0.7)],[MWsats],'y*',color='#ff8c00',markersize=11)
ax.plot([-22.0],[M31sats],'*',color='#ba55d3',markersize=11) # hammer 2007
ax.xaxis.set_major_locator(ticker.MultipleLocator(0.5))
ax.set_xlabel('$M_r$')
ax.get_yaxis().set_ticklabels([])
hp = hosts['M_r']
np.ma.append(hp,-20.7+ 5.*np.log10(0.7))
sr = spearmanr(hp,ns)
#sr = spearmanr(hosts['M_r'],sats['Ncompl'])
t = '$M_r$: ($r_s$={:04.2f}, p={:04.2f})'.format(sr[0],sr[1])
ax.text(-21.85,yl,t,fontsize=8)
print sr[1]
# COLORS
# g-r
gr = hosts['M_g'] - hosts['M_r']
ax = plt.subplot2grid((3,2), (1, 0))
ax.plot(gr,sats['Nsat'],'ko')
ax.plot(gr,sats['Ncompl'],'ko',markerfacecolor='none')
ax.plot([0.68],[MWsats],'y*',color='#ff8c00',markersize=11)
ax.plot([0.708],[M31sats],'y*',color='#ba55d3',markersize=11)
ax.xaxis.set_major_locator(ticker.MultipleLocator(0.05))
ax.set_ylim(0, ylim)
ax.set_xlabel('$(g-r)_o$')
ax.set_ylabel('N$_{\mathrm{sats}}$')
hp = gr
np.ma.append(hp,0.68)
sr = spearmanr(hp,ns)
print sr
sr = spearmanr(gr,sats['Ncompl'])
print sr
print hp
print gr
t = '(g-r): ($r_s$={:04.2f}, p={:04.2f})'.format(sr[0],sr[1])
ax.text(0.705,yl,t,fontsize=8)
print sr[1]
# u-r
ur = hosts['M_u'] - hosts['M_r']
ax = plt.subplot2grid((3,2), (1, 1))
ax.plot(ur,sats['Nsat'],'ko')
ax.plot(ur,sats['Ncompl'],'ko',markerfacecolor='none')
ax.plot([2.043],[MWsats],'y*',color='#ff8c00',markersize=11)
ax.plot([2.32],[M31sats],'y*',color='#ba55d3',markersize=11)
ax.xaxis.set_major_locator(ticker.MultipleLocator(0.25))
ax.set_ylim(0, ylim)
ax.set_xlabel('$(u-r)_o$')
ax.get_yaxis().set_ticklabels([])
hp = ur
np.ma.append(hp,2.043)
sr = spearmanr(hp,ns)
#sr = spearmanr(ur,sats['Ncompl'])
t = '(u-r): ($r_s$={:04.2f}, p={:04.2f})'.format(sr[0],sr[1])
ax.text(2.1,yl,t,fontsize=8)
print sr[1]
# DERIVED PROPERTIES
# MSTAR
# 6.08 +/- 1.14 x 10^10
ax = plt.subplot2grid((3,2), (2, 0))
ax.plot(log_SM,sats['Nsat'],'ko',label='_nolabel_')
ax.plot(log_SM,sats['Ncompl'],'ko',markerfacecolor='none',label='_nolabel_')
ax.xaxis.set_major_locator(ticker.MultipleLocator(0.25))
ax.plot([np.log10(6.08e10)],[MWsats],'y*',color='#ff8c00',markersize=11,label='MW')
ax.plot([np.log10(10.5e10)],[M31sats],'y*',color='#ba55d3',markersize=11,label='M31')
ax.set_ylim(0, ylim)
ax.set_xlim(10.4, 11.1)
ax.set_xlabel('log[M$_{\mathrm{star}}$]')
ax.set_ylabel('N$_{\mathrm{sats}}$')
hp = log_SM
np.ma.append(hp,np.log10(6.08e10))
sr = spearmanr(hp,ns)
#sr = spearmanr(log_SM,sats['Ncompl'])
t = 'log[M$_{{star}}$] : ($r_s$={:04.2f}, p={:04.2f})'.format(sr[0],sr[1])
ax.text(10.55,yl,t,fontsize=8)
print sr[1]
ax.legend(fontsize=10,loc=4,numpoints = 1,frameon=True)
# SFR
# 1.65 +/- 0.19 msun/year
ax = plt.subplot2grid((3,2), (2, 1))
ax.plot(sats['logSFR'],sats['Nsat'],'ko')
ax.plot(sats['logSFR'],sats['Ncompl'],'ko',markerfacecolor='none')
ax.set_ylim(0, ylim)
ax.set_xlabel('log[SFR]')
ax.set_xlim(-1.28,1.6)
ax.get_yaxis().set_ticklabels([])
ax.plot([np.log10(1.65)],[MWsats],'y*',color='#ff8c00',markersize=11)
ax.plot([np.log10(0.7)],[M31sats],'y*',color='#ba55d3',markersize=11)
hp = sats['logSFR']
np.ma.append(hp,-1.28)
sr = spearmanr(hp,ns)
#sr = spearmanr(sats['logSFR'],sats['Ncompl'])
t = 'log[SFR] : ($r_s$={:04.2f}, p={:04.2f})'.format(sr[0],sr[1])
ax.text(-0.57,yl,t,fontsize=8)
print sr[1]
plt.tight_layout(w_pad=0)
plt.savefig('fig_nsats.pdf')
# -
# There are 9 M31 satellites.
# M33 -19.14
# M32 -16.73
# IC 10 -15.3
#
# NGC 205 -16.78
# NGC 147 -14.95
# NGC 185 -15.05
#
# And XXXII -12.55
# Andromeda II -12.87
# Andromeda VII -13.51
#
#
# Andromeda I -12.16
# And XXXI -12.0
# Andromeda VI -11.77
#
|
plot_Fig12_nsat.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Use numpy to convert to arrays
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
dataset = pd.read_csv('/Users/xinyuewang/Downloads/normalized_by_2000_dtpts.csv')
dataset
# Labels are the values we want to predict
labels = np.array(dataset['Normalized_Solar_lin'])
# Remove the labels from the features
# axis 1 refers to the columns
realtime_columns = ['DHI', 'DNI', 'GHI','Solar Zenith Angle', 'Surface Albedo','Temperature']
columns_all= ['DHI', 'DNI', 'GHI', 'Cloud Type', 'Dew Point',
'Solar Zenith Angle', 'Surface Albedo', 'Wind Speed',
'Wind Direction', 'Relative Humidity', 'Temperature', 'Pressure']
RFcolumns=['Relative Humidity','Solar Zenith Angle','GHI','Surface Albedo','Wind Direction']
multilinearColumns=['Surface Albedo','Temperature','Relative Humidity','Solar Zenith Angle','Pressure']
features= dataset[realtime_columns]
# Saving feature names for later use
feature_list = list(features.columns)
# Convert to numpy array
features = np.array(features)
features
print('The shape of our features is:', features.shape)
# Using Skicit-learn to split data into training and testing sets
from sklearn.model_selection import train_test_split
# Split the data into training and testing sets
train_features, test_features, train_labels, test_labels = train_test_split(features, labels, test_size = 0.25, random_state = 42)
print('Training Features Shape:', train_features.shape)
print('Training Labels Shape:', train_labels.shape)
print('Testing Features Shape:', test_features.shape)
print('Testing Labels Shape:', test_labels.shape)
# Import the model we are using
from sklearn.ensemble import RandomForestRegressor
# Instantiate model with 1000 decision trees
rf = RandomForestRegressor(n_estimators = 1000, random_state = 42)
# Train the model on training data
rf.fit(train_features, train_labels)
from sklearn import metrics
from sklearn.metrics import r2_score
# Use the forest's predict method on the test data
predictions = rf.predict(test_features)
# Calculate the absolute errors
errors = abs(predictions - test_labels)
# Print out the mean absolute error (mae)
print('Mean Absolute Error:', round(np.mean(errors), 2), 'watt.')
print('Mean Absolute Error:', metrics.mean_absolute_error(test_labels, predictions))
print('Mean squared Error::', metrics.mean_squared_error(test_labels, predictions))
print('R Squared Error:', r2_score(test_labels, predictions))
predictions
test_labels
plt.scatter(predictions,test_labels)
plt.plot([0,1],[0,1], 'r--')
# Calculate mean absolute percentage error (MAPE)
mape = 100 * (errors / test_labels)
# Calculate and display accuracy
accuracy = 100 - np.mean(mape)
print('Accuracy:', round(accuracy, 2), '%.')
# when choose all the 12 feature columns R2 = 0.78
# when choose top 5 feature from RF feature selection.
# Mean Absolute Error: 0.12 watt.
# Mean Absolute Error: 0.12456254980498581
# R Squared Error: 0.7111779101313127
# when choose top 5 feature from linear regression selection.
# Mean Absolute Error: 0.12 watt.
# Mean Absolute Error: 0.12005829970846314
# R Squared Error: 0.7256975802742123
residual = predictions - test_labels
i = 5000
plt.scatter(range(i), residual[:i])
dataset[columns_all]
from sklearn.model_selection import KFold
def kfold_CV(X_columns_all):
y=labels
kf = KFold(n_splits=10, shuffle=True)# Define the split - into 10 folds
kf.get_n_splits(X_columns_all, y)# returns the number of splitting iterations in the cross-validator
print(kf)
fig, ax = plt.subplots(figsize=(8,6))
mse = np.zeros((12, 10))
j = 0
#creat train and test vars
for train_index, test_index in kf.split(X_columns_all, y):
print("TRAIN:", train_index, "TEST:", test_index)
#print("x-train[X_train.columns[0:2]]: ", X_train[X_train.columns[0:2].values])
#print (X_train.shape, y_train.shape)
#print (X_test.shape, y_test.shape)
X_train, X_test = dataset[columns_all].iloc[train_index], dataset[columns_all].iloc[test_index]
y_train, y_test = y[train_index], y[test_index]
for i in range(1, 13):
rf = RandomForestRegressor(n_estimators = 100, random_state = 42)#change estimator
# Train the model on training data
reg = rf.fit(X_train[X_train.columns[0:i].values], y_train)#use first i features train
y_pred = rf.predict(X_test[X_test.columns[0:i].values])
mse[i-1, j] = metrics.mean_squared_error(y_test, y_pred)
ax.plot(np.linspace(1, 11, 12), mse[:, j], linewidth=4, color='b', alpha=0.09)
ax.set_ylabel('Mean Squared Error')
ax.set_xlabel('Degree of Polynomial')
j += 1
avg_mse = mse.mean(axis=1)
ax.plot(np.linspace(1, 11, 12), avg_mse, color='purple')
kfold_CV(dataset[columns_all])
|
examples/regression_methods/RandomForest_Regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: nama
# language: python
# name: nama
# ---
# %load_ext autoreload
# %autoreload 2
# +
import os
import torch
import numpy as np
from ray import tune
from ray.tune.schedulers import ASHAScheduler
from ray.tune.suggest.hyperopt import HyperOptSearch
from ray.tune.integration.wandb import WandbLoggerCallback
from src.data.utils import load_dataset, select_frequent_k
from src.eval import metrics
from src.models import swivel
# +
# Config
wandb_api_key_file = "" # "../.wandb-api-key"
given_surname = "given"
num_matches = 500
SAMPLE_SIZE = 30000
TRAIN_DATA_PATH = f"s3://familysearch-names/processed/tree-hr-{given_surname}-train-augmented.csv.gz"
# -
# NOTE: we're setting is_eval to False even though we use this dataset for evaluation
# it would be better if we re-loaded the dataset with is_eval=True and used that for evaluation
# but it may not matter much for hyperparameter optimization
input_names_train, weighted_actual_names_train, candidate_names_train = load_dataset(TRAIN_DATA_PATH, is_eval=False)
print(len(input_names_train))
print(len(candidate_names_train))
# sample the dataset
input_names_train_sample, weighted_actual_names_train_sample, candidate_names_train_sample = \
select_frequent_k(input_names_train,
weighted_actual_names_train,
candidate_names_train,
SAMPLE_SIZE)
print("sample input names", len(input_names_train_sample))
print("sample number of actuals", sum(len(wan) for wan in weighted_actual_names_train_sample))
print("sample candidate names", len(candidate_names_train_sample))
DEFAULT_VOCAB_SIZE = SAMPLE_SIZE
DEFAULT_EMBEDDING_DIM = 100
DEFAULT_CONFIDENCE_BASE = 0.18
DEFAULT_CONFIDENCE_SCALE = 0.5
DEFAULT_CONFIDENCE_EXPONENT = 0.3
DEFAULT_LEARNING_RATE = 0.14
DEFAULT_SUBMATRIX_SIZE = 2048 # Needs to be adjusted with full dataset
DEFAULT_NUM_EPOCHS = 30 # Needs to be adjusted with full dataset
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
print(device)
def compute_auc(model,
vocab,
input_names,
weighted_actual_names,
candidate_names):
best_matches = swivel.get_best_swivel_matches(model,
vocab,
input_names,
candidate_names,
k=num_matches,
batch_size=256,
add_context=True,
n_jobs=1,
progress_bar=False)
return metrics.get_auc(weighted_actual_names,
best_matches,
min_threshold=0.01,
max_threshold=2.0,
step=0.03,
distances=False)
def plot_pr_curve(model,
vocab,
input_names,
weighted_actual_names,
candidate_names):
best_matches = swivel.get_best_swivel_matches(model,
vocab,
input_names,
candidate_names,
k=num_matches,
batch_size=256,
add_context=True,
n_jobs=1,
progress_bar=False)
metrics.precision_weighted_recall_curve_at_threshold(weighted_actual_names,
best_matches,
min_threshold=0.01,
max_threshold=2.0,
step=0.05,
distances=False)
def train_model(param_config,
input_names,
weighted_actual_names,
candidate_names,
checkpoint_dir=None):
swivel_dataset = swivel.SwivelDataset(input_names,
weighted_actual_names,
param_config['vocab_size'])
swivel_vocab = swivel_dataset.get_vocab()
# Instantiate the model
model = swivel.SwivelModel(len(swivel_vocab),
param_config['embedding_dim'],
param_config['confidence_base'],
param_config['confidence_scale'],
param_config['confidence_exponent'])
# Init model biases
model.init_params(swivel_dataset.get_row_sums(), swivel_dataset.get_col_sums())
# Put model on device
model.to(device)
# Create optimizer
optimizer = torch.optim.Adagrad(model.parameters(),
lr=param_config['learning_rate'])
# Load checkpoint if exists
if checkpoint_dir:
model_state, optimizer_state = torch.load(
os.path.join(checkpoint_dir, "checkpoint"))
model.load_state_dict(model_state)
optimizer.load_state_dict(optimizer_state)
for epoch in range(param_config['num_epochs']):
loss_values = swivel.train_swivel(model,
swivel_dataset,
n_steps=0,
submatrix_size=param_config['submatrix_size'],
lr=param_config['learning_rate'],
device=device,
verbose=False,
optimizer=optimizer)
# Compute AUC on the train data
auc = compute_auc(model,
swivel_vocab,
input_names,
weighted_actual_names,
candidate_names)
# Checkpoint the model
with tune.checkpoint_dir(epoch) as checkpoint_dir:
path = os.path.join(checkpoint_dir, "checkpoint")
torch.save((model.state_dict(), optimizer.state_dict()), path)
# Report the metrics to Ray
tune.report(auc=auc, mean_loss=np.mean(loss_values))
# ### Search space for parameters
param_config = {
"vocab_size": DEFAULT_VOCAB_SIZE,
"embedding_dim": tune.grid_search([50, 100]),
"confidence_base": DEFAULT_CONFIDENCE_BASE, # tune.quniform(0.1, 0.2, 0.02),
"confidence_scale": DEFAULT_CONFIDENCE_SCALE, # tune.quniform(0.4, 0.5, 0.05),
"confidence_exponent": DEFAULT_CONFIDENCE_EXPONENT, # tune.quniform(0.2, .4, 0.05),
"learning_rate": DEFAULT_LEARNING_RATE, # tune.quniform(0.04, 0.3, 0.02),
"submatrix_size": DEFAULT_SUBMATRIX_SIZE, # Needs to be adjusted with full dataset
"num_epochs": DEFAULT_NUM_EPOCHS
}
# Will try to terminate bad trials early
# https://docs.ray.io/en/latest/tune/api_docs/schedulers.html
scheduler = ASHAScheduler(max_t=100,
grace_period=1,
reduction_factor=4)
# +
# Can provide multiple points
current_best_params = [
{
"vocab_size": DEFAULT_VOCAB_SIZE,
"embedding_dim": DEFAULT_EMBEDDING_DIM,
"confidence_base": DEFAULT_CONFIDENCE_BASE,
"confidence_scale": DEFAULT_CONFIDENCE_SCALE,
"confidence_exponent": DEFAULT_CONFIDENCE_EXPONENT,
"learning_rate": DEFAULT_LEARNING_RATE,
"submatrix_size": DEFAULT_SUBMATRIX_SIZE,
"num_epochs": DEFAULT_NUM_EPOCHS
}
]
# https://docs.ray.io/en/latest/tune/api_docs/suggestion.html#tune-hyperopt
search_alg = HyperOptSearch(points_to_evaluate=current_best_params)
# -
# ### Run HPO
# +
callbacks = []
if wandb_api_key_file:
callbacks.append(WandbLoggerCallback(
project="nama",
entity="nama",
group="60_swivel_tune_"+given_surname,
notes="",
config={
"SAMPLE_SIZE": SAMPLE_SIZE,
},
api_key_file=wandb_api_key_file
))
result = tune.run(tune.with_parameters(train_model,
input_names=input_names_train_sample,
weighted_actual_names=weighted_actual_names_train_sample,
candidate_names=candidate_names_train_sample),
resources_per_trial={'cpu': 0.5, 'gpu': 0.5},
config=param_config,
# scheduler=scheduler,
# search_alg=search_alg,
# num_samples=8,
# metric='auc',
# mode='max',
# checkpoint_score_attr='auc',
# time_budget_s=6*3600,
# keep_checkpoints_num=100,
progress_reporter=tune.JupyterNotebookReporter(
overwrite=False,
max_report_frequency=5*60
),
callbacks=callbacks
)
# -
# ### Get best model
# Get trial that has the highest AUC (can also do with mean_loss or any other metric)
best_trial_auc = result.get_best_trial(metric='auc', mode='max', scope='all')
# Parameters with the highest AUC
best_trial_auc.config
print(f"Best trial final train loss: {best_trial_auc.last_result['mean_loss']}")
print(f"Best trial final train auc: {best_trial_auc.last_result['auc']}")
# +
# Get checkpoint dir for best model
best_checkpoint_dir = best_trial_auc.checkpoint.value
# get vocab
swivel_dataset = swivel.SwivelDataset(input_names_train_sample,
weighted_actual_names_train_sample,
DEFAULT_VOCAB_SIZE)
swivel_vocab = swivel_dataset.get_vocab()
# Load best model
model_state, optimizer_state = torch.load(os.path.join(best_checkpoint_dir, 'checkpoint'))
best_trained_model = swivel.SwivelModel(len(swivel_vocab),
embedding_dim=best_trial_auc.config['embedding_dim'],
confidence_base=best_trial_auc.config['confidence_base'],
confidence_scale=best_trial_auc.config['confidence_scale'],
confidence_exponent=best_trial_auc.config['confidence_exponent'])
best_trained_model.load_state_dict(model_state)
# -
# ### Plot PR curve
# plot pr curve with best model
plot_pr_curve(best_trained_model,
swivel_vocab,
input_names_train_sample,
weighted_actual_names_train_sample,
candidate_names_train_sample)
# ### Demo
best_matches = swivel.get_best_swivel_matches(best_trained_model,
swivel_vocab,
input_names_train_sample,
candidate_names_train_sample,
k=num_matches,
batch_size=256,
add_context=True,
n_jobs=1,
progress_bar=False)
rndmx_name_idx = np.random.randint(len(input_names_train_sample))
print(f"Input name: {input_names_train_sample[rndmx_name_idx]}")
print("Nearest names:")
print(best_matches[rndmx_name_idx][:10])
print("Actual names:")
sorted(weighted_actual_names_train[rndmx_name_idx][:10], key=lambda k: k[1], reverse=True)
# ### Get all trials as DF
# All trials as pandas dataframe
df = result.results_df
df
df[(df["training_iteration"] == 30)].sort_values(by="auc")
|
notebooks/60_swivel_tune.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import requests
# ## Function builder
dictToSend = {'t_init':490,'t_end':550,'function':{
"function": "events",
"values": [
0.3,
{
"function": "transition",
"ftype": 0,
"initvalue": 0.3,
"endvalue": 1,
"concavity": 0,
"t_init":500,
"t_end":510
}
],
"days": [
[
0,
500
],
[
500,
540
]
],
"default": 0.3
}}
# + jupyter={"outputs_hidden": true}
res = requests.post('http://localhost:5003/function', json=dictToSend)
print('response from server:',res.text)
dictFromServer = res.json()
# + jupyter={"outputs_hidden": true}
dictFromServer['results']
# -
plt.plot(dictFromServer['results']['t'],dictFromServer['results']['function'])
import numpy as np
import matplotlib.pyplot as plt
np.linspace(0,10,101)
|
backend/backend_test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.3 64-bit
# name: python383jvsc74a57bd0aee8b7b246df8f9039afb4144a1f6fd8d2ca17a180786b69acc140d282b71a49
# ---
import pandas as pd
from functools import reduce
_MULTIINDEX_ = ["NOMBREDD", "NOMBREPV", "NOMBREDI"]
_PATH_ = "https://public.minsky.cc/maria/market_channel_factors_db"
df = pd.read_csv(_PATH_ + "/" + "databases/2019_03PreModelDatabaseOfWhitePotato.csv")
df_districtToLimaMarket = pd.read_csv(_PATH_ + "/" + 'databases/2019_04RoadTransportationDistrictToLimaMarket.csv')
df_districtToProvinceCapitalOfRegion = pd.read_csv(_PATH_ + "/" + 'databases/2019_04RoadTransportationDistrictToProvinceCapitalOfRegion.csv')
df_provinceCapitalToLimaMarket = pd.read_csv(_PATH_ + "/" + 'databases/2019_04RoadTransportationProvinceCapitalToLimaMarket.csv')
df = df.drop(columns=["Unnamed: 0"])
df.columns
df_districtToLimaMarket = df_districtToLimaMarket[["NOMBREDD", "NOMBREPV", "NOMBREDI", "districtTimeToLimaMarket"]]
df_districtToProvinceCapitalOfRegion = df_districtToProvinceCapitalOfRegion[["NOMBREDD", "NOMBREPV", "NOMBREDI", "districtTimeToProvinceCapitalOfRegion"]]
df_provinceCapitalToLimaMarket = df_provinceCapitalToLimaMarket[["NOMBREDD", "provinceCapitalToLimaMarket"]]
df_districtToLimaMarket.columns
df_districtToProvinceCapitalOfRegion.columns
df_provinceCapitalToLimaMarket.columns
def mergeDataFrames(dfArray, indices):
dfList = []
for df in dfArray:
dfList.append(df)
df = reduce(lambda x, y: pd.merge(x, y, how="left", on = indices, suffixes=("", "_y")), dfList)
filter = [col for col in df if col.endswith("y")]
#Eliminar
df_drop = df.drop(columns=filter)
#Seleccionar
df_keep = df[filter]
return df_drop
len(df_districtToLimaMarket), len(df_districtToProvinceCapitalOfRegion), len(df_provinceCapitalToLimaMarket)
df_transportation = mergeDataFrames([mergeDataFrames([df_districtToLimaMarket, df_districtToProvinceCapitalOfRegion], ["NOMBREDD", "NOMBREPV", "NOMBREDI"]), df_provinceCapitalToLimaMarket], ["NOMBREDD"])
df_transportation.columns
df_model = mergeDataFrames([df, df_transportation], _MULTIINDEX_)
df_model.columns
len(df_model)
df_model.head()
df_model.isnull().sum(axis=0)
# +
#df_model.to_csv("~/Desktop/DatosENA2019/2019_04ModelDatabaseOfWhitePotato.csv")
|
ENA2019/04BDatabaseModelWithTransportVariables.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 4.1. Evaluating the time taken by a command in IPython
n = 100000
# %timeit sum([1. / i**2 for i in range(1, n)])
# +
# %%timeit s = 0.
for i in range(1, n):
s += 1. / i**2
# +
import numpy as np
# %timeit np.sum(1. / np.arange(1., n) ** 2)
# -
|
chapter04_optimization/01_timeit.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Newton's Method for finding a root
#
#
# [Newton's method](https://en.wikipedia.org/wiki/Newton's_method) uses a clever insight to iteratively home in on the root of a function $f$. The central idea is to approximate $f$ by its tangent at some initial position $x_0$:
#
# $$
# y = f'(x_0) (x-x_0) + f(x_0)
# $$
#
# The $x$-intercept of this line is then closer to the root than the starting position $x_0$. That is, we need to solve the linear relation
#
# $$
# f'(x_n)(x_1-x_0) + f(x_0) = 0
# $$
#
# for the updated position $x_1 = x_0 - f(x_0)/f'(x_0)$. Repeating this sequence
#
# $$
# x_{n+1} = x_n - \frac{f(x_n)}{f'(x_n)}
# $$
#
# will yield a fixed point, which is the root of $f$ *if one exists in the vicinity of $x_0$*.
# + deletable=true editable=true
def newtons_method(f, df, x0, tol=1E-6):
x_n = x0
while abs(f(x_n)) > tol:
x_n = x_n - f(x_n)/df(x_n)
return x_n
# -
# ## Minimizing a function
#
# As the maximum and minimum of a function are defined by $f'(x) = 0$, we can use Newton's method to find extremal points by applying it to the first derivative. Let's try this with a simply function with known minimum:
# + deletable=true editable=true
# define a test function
def f(x):
return (x-3)**2 - 9
def df(x):
return 2*(x-3)
def df2(x):
return 2.
# + deletable=true editable=true
root = newtons_method(f, df, x0=0.1)
print ("root {0}, f(root) = {1}".format(root, f(root)))
# + deletable=true editable=true
minimum = newtons_method(df, df2, x0=0.1)
print ("minimum {0}, f'(minimum) = {1}".format(minimum, df(minimum)))
# -
# There is an important qualifier in the statement about fixed points: **a root needs to exist in the vicinity of $x_0$!** Let's see what happens if that's not the case:
def g(x):
return (x-3)**2 + 1
dg = df # same derivatives for f and g
newtons_method(g, dg, x0=0.1)
# As long as you don't interrupt the execution of this cell (Tip: click "Interrupt Kernel"), `newtons_method` will not terminate and come back with a result.
#
# With a little more defensive programming we can make sure that the function will terminate after a given number of iterations:
# + deletable=true editable=true
def newtons_method2(f, df, x0, tol=1E-6, maxiter=100000):
x_n = x0
for _ in range(maxiter):
x_n = x_n - f(x_n)/df(x_n)
if abs(f(x_n)) < tol:
return x_n
raise RuntimeError("Failed to find a minimum within {} iterations ".format(maxiter))
# -
newtons_method2(g, dg, x0=0.1)
# ## Using scipy.optimize
#
# scipy comes with a pretty feature-rich [optimization package](https://docs.scipy.org/doc/scipy/reference/optimize.html), for one- and multi-dimensional optimization. As so often, it's better (as in faster and more reliable) to leverage exisiting and battle-tested code than to try to implement it yourself.
#
# ### Exercise 1:
#
# Find the minimum of `f` with `scipy.optimize.minimize_scalar`. When done, visualize your result to confirm its correctness.
# To make this more interesting, we'll create a new multi-dimensional function that resembles `f`:
def h(x, p):
return np.sum((x-3)**p, axis=-1) - 9
# ### Exercise 2:
#
# In 2D, find the minimum of `h` for `p=2` with `scipy.optimimze.minimize`. Note that you have not been given a derivative of `h`. You can choose to compute it analytically, or see if `minimize` has options that allow you to work without.
#
# When done, visualize your result to confirm its correctness.
|
day4/Newton-Method.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Name: example_calibration_analysis.ipynb
# Authors: <NAME>
# Shows an example calibration analysis
# -
# General imports
import numpy as np
import matplotlib.pyplot as plt
import sys
import pandas as pd
from tqdm import tqdm
from scipy.interpolate import UnivariateSpline
# Adding path to module
sys.path.append("../")
# picture path
PICS = '../pics/'
# Module imports
from fourth_day import Fourth_Day, config
# Some example settings
config['scenario']['class'] = 'Calibration'
config['scenario']['light prop'] = {
"switch": True,
"x_pos": 5.,
"y_pos": 10.,
}
config['scenario']['detector'] = {
"switch": True,
"type": "PMTSpec_Func",
"response": True,
"acceptance": "Flat",
"mean detection prob": 1.
}
# The flasher position
config['calibration']['pos_arr'] = [2., 30.]
# The injected pulse
config['calibration']['light curve'] = {
396.: np.ones(100),
400.: np.ones(100),
506.: np.ones(100),
514.: np.ones(100),
545.: np.ones(100),
550.: np.ones(100),
}
samples = 50
sim_counts = []
for sample in tqdm(range(samples)):
# Varying the angle pointing by 1%
angle_offset_var = np.random.uniform(-0.01, 0.01, 12)
# Varying the opening angle by 1%
angle_opening_var = np.random.uniform(-0.01, 0.01, 12)
# Varying the quantum efficiency functions by 5%
qe_var = np.random.uniform(-0.01, 0.01, (12, 3))
# Varying the attenuation curve by 10 % at each point
atten_var = np.random.uniform(-0.15, 0.15, 23)
config['geometry']['detector properties']["PMTSpec_Func"] = {
"x_pos": 2.,
"y_pos": 5.,
"det num": 12, #12 pmts numbered by position
"x_offsets": np.array(
[0.1,0.,-0.1,0., 0.12,-0.12,-0.12,0.12, 0.2,-0.04,-0.2,0.04]
) / 2., #test radius 0.3 meter, real radius 0.15 meter
"y_offsets": np.array(
[0.,0.1,0.,-0.1, 0.12,0.12,-0.12,-0.12, 0.04,0.2,-0.04,-0.2]
) / 2.,
"angle offset": np.array([
90., 90., 90., 90., 90., 90.,
90., 90., 90., 90., 90., 90.]) * (1. + angle_offset_var), # In which direction the detector(s) points
"opening angle": np.array([
25., 25., 25., 25., 25., 25.,
25., 25., 25., 25., 25., 25.]) * (1. + angle_opening_var), # 25., # from dark box rotation test result: +-25 degrees
"quantum efficiency": "Func", # whether flat or function
"wavelength acceptance": np.array([ #position number,center wavelength,quantum efficiency (if flat)
[395., 405.],
[505., 515.],
[420., 430.],
[465., 475.],
[300., 600.],
[487., 497.],
[540., 560.],
[515., 535.],
[475., 485.],
[445., 455.],
[455., 465.],
[325., 375.],
]),
"quantum func": np.array([
[[395., 400., 405.], np.array([0.26, 0.26, 0.26]) * (1. + qe_var[0])],
[[505., 510., 515.], np.array([0.16, 0.16, 0.16]) * (1. + qe_var[1])],
[[420., 425., 430.], np.array([0.28, 0.28, 0.28]) * (1. + qe_var[2])],
[[465., 470., 475.], np.array([0.23, 0.23, 0.23]) * (1. + qe_var[3])],
[[300., 500., 600.], np.array([1., 1., 1.]) * (1. + qe_var[4])],
[[487., 490., 497.], np.array([0.1, 0.1, 0.1]) * (1. + qe_var[5])],
[[540., 550., 560.], np.array([0.1, 0.1, 0.1]) * (1. + qe_var[6])],
[[515., 525., 535.], np.array([0.13, 0.13, 0.13]) * (1. + qe_var[7])],
[[475., 480., 485.], np.array([0.2, 0.2, 0.2]) * (1. + qe_var[8])],
[[445., 450., 455.], np.array([0.2, 0.2, 0.2]) * (1. + qe_var[9])],
[[455., 460., 465.], np.array([0.23, 0.23, 0.23]) * (1. + qe_var[10])],
[[325., 350., 375.], np.array([0.3, 0.3, 0.3]) * (1. + qe_var[11])],
])
}
config['calibration']['attenuation curve'] = np.array([
[
299.,
329.14438502673795, 344.11764705882354, 362.2994652406417,
399.44415494181, 412.07970421102266, 425.75250006203635,
442.53703565845314, 457.1974490682151, 471.8380108687561,
484.3544504826423, 495.7939402962853, 509.29799746891985,
519.6903148961513, 530.0627807141617, 541.5022705278046,
553.9690811186382, 567.4929899004939, 580.9771954639073,
587.1609717362714, 593.3348222040249, 599.4391920395047,
602.4715253480235
],
np.array([
0.8,
0.6279453220864465,0.3145701363176568,
0.12591648888305143,0.026410321551339357, 0.023168667048510762,
0.020703255370450736, 0.019552708373076478,
0.019526153330089138, 0.020236306473695613,
0.02217620815962483, 0.025694647290888873,
0.031468126242251794, 0.03646434475343956,
0.04385011375530569, 0.05080729755501162,
0.061086337538657706, 0.07208875589035815, 0.09162216168767365,
0.11022281058708046, 0.1350811713674855, 0.18848851206491904,
0.23106528395398912
]) * (1. + atten_var)
])
# Creating a fourth_day object
fd = Fourth_Day()
# Launching solver
fd.sim()
sim_counts.append(fd.measured)
spl_atten = UnivariateSpline(
[
299.,
329.14438502673795, 344.11764705882354, 362.2994652406417,
399.44415494181, 412.07970421102266, 425.75250006203635,
442.53703565845314, 457.1974490682151, 471.8380108687561,
484.3544504826423, 495.7939402962853, 509.29799746891985,
519.6903148961513, 530.0627807141617, 541.5022705278046,
553.9690811186382, 567.4929899004939, 580.9771954639073,
587.1609717362714, 593.3348222040249, 599.4391920395047,
602.4715253480235
],
np.array([
0.8,
0.6279453220864465,0.3145701363176568,
0.12591648888305143,0.026410321551339357, 0.023168667048510762,
0.020703255370450736, 0.019552708373076478,
0.019526153330089138, 0.020236306473695613,
0.02217620815962483, 0.025694647290888873,
0.031468126242251794, 0.03646434475343956,
0.04385011375530569, 0.05080729755501162,
0.061086337538657706, 0.07208875589035815, 0.09162216168767365,
0.11022281058708046, 0.1350811713674855, 0.18848851206491904,
0.23106528395398912
]), k=1, s=0
)
# Distances
distances = np.array([
(config['calibration']['pos_arr'][0] - (fd._lucifer._det_geom['x_pos'] + fd._lucifer._det_geom["x_offsets"][i]))**2. +
(config['calibration']['pos_arr'][1] - (fd._lucifer._det_geom['y_pos'] + fd._lucifer._det_geom["y_offsets"][i]))**2.
for i in range(0, fd._lucifer._det_geom["det num"])])**(1./2.)
# Attenuation function
def atten_func(distance, atten):
factor = np.exp(-distance * atten) / (4. * np.pi * distance**2.)
if factor > 1./2.:
factor = 1./2.
return factor
def dist_atten(distance):
return (4. * np.pi * distance**2.)
# Plotting standards
std_size = 6.
fontsize = 20.
lw=1.
h_length=0.2
export_dpi = 500
from matplotlib import rc
rc('font',**{'family':'sans-serif','sans-serif':['Helvetica']})
rc('text', usetex=False)
# Detector 0 evaluation
norm_fac = 1.
figure, (ax1) = plt.subplots(1, 1, figsize=(std_size, std_size * 6. / 8.), sharex=True)
det = 'Detector 0'
for sample in sim_counts:
atten_facs = np.array([np.mean(-np.log(sample[det].values * dist_atten(distances[0]) / 4. / 0.26) / distances[0])
for sample in sim_counts])
ax1.scatter(
range(len(sim_counts)), atten_facs / norm_fac, label=r'$\lambda = 398\;\mathrm{nm}$', color='b',
)
print(np.mean(atten_facs) / spl_atten(398.))
det = 'Detector 1'
for sample in sim_counts:
atten_facs = np.array([np.mean(-np.log(sample[det].values * dist_atten(distances[1]) / 8. / 0.16) / distances[1])
for sample in sim_counts])
ax1.scatter(
range(len(sim_counts)), atten_facs / norm_fac, label=r'$\lambda = 510\;\mathrm{nm}$', color='g',
)
print(np.mean(atten_facs) / spl_atten(510.))
det = 'Detector 6'
for sample in sim_counts:
atten_facs = np.array([np.mean(-np.log(sample[det].values * dist_atten(distances[2]) / 5. / 0.1) / distances[2])
for sample in sim_counts])
ax1.scatter(
range(len(sim_counts)), atten_facs / norm_fac, label=r'$\lambda = 547\;\mathrm{nm}$', color='r',
)
print(np.mean(atten_facs) / spl_atten(547.))
ax1.axhline(spl_atten(398.), color='b', lw=lw)
ax1.axhline(spl_atten(510.), color='g', lw=lw)
ax1.axhline(spl_atten(547.), color='r', lw=lw)
ax1.set_xscale('linear')
ax1.set_yscale('linear')
ax1.set_xlabel(r'$\mathrm{Run}$', fontsize=fontsize)
ax1.set_ylabel(r'$\mathrm{Attenuation\;Factor}$', fontsize=fontsize)
ax1.tick_params(axis = 'both', which = 'major', labelsize=fontsize, direction='in')
ax1.tick_params(axis = 'both', which = 'minor', labelsize=fontsize, direction='in')
# ax1.grid(True)
h, l = ax1.get_legend_handles_labels()
lgd1 = ax1.legend(h,l, loc=9, bbox_to_anchor=(0.5, +1.25),
ncol=6, fontsize=fontsize, handlelength=h_length,
fancybox=True, frameon=False)
# ax1.set_xlim(390., 400)
plt.tight_layout()
plt.show()
figure.savefig(PICS + "Calibration_Pop.png",
bbox_inches='tight', dpi=export_dpi)
|
examples/example_calibration_analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import cvxpy as cp
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.animation import FFMpegWriter
u = cp.Variable(1)
a = cp.Parameter(value=0.7)
b = cp.Parameter(value=0.1)
# x >= 0 + t = u
# x <= 1 + 0.3*t only valid till t = 1/0.7. But for some time horizon, it should be possible to keep the constraints
def xdot(x,u):
return u
def simulate_simple(x0,av,bv,c,movie_name=None):
t = 0
dt = 0.01
N = int(1.0/(1.0-c)*100)
x = x0
a.value = av
b.value = bv
xs = [x]
ts = [0]
if movie_name==None:
movie_name = 'temp.mp4'
fig = plt.figure(figsize=(2, 0.5))
ax = plt.axes(xlim=(-0.5,2),ylim=(-0.05,0.05))
plt.ion()
car1 = ax.scatter([t],[0],c='r',s=10)
car2 = ax.scatter([1 + c*t],[0],c='r',s=10)
vehicle = ax.scatter([x0],[0],c='g',s=10)
metadata = dict(title='Movie Test', artist='Matplotlib',comment='Movie support!')
writer = FFMpegWriter(fps=15, metadata=metadata)
ax.get_yaxis().set_visible(False)
ax.get_xaxis().set_visible(False)
# if 1:
with writer.saving(fig, movie_name, 100):
for i in range(N):
#h1
h1 = x - t # x>=t
h1_dot = xdot(x,u) - 1
#h2
h2 = 1 + c*t - x #x <= 1 + 0.3t
h2_dot = c - xdot(x,u)
if h1<0 or h2<0:
print("************* ERROR ****************")
#constraints
const = [h1_dot >= -a*h1]
const += [h2_dot >= -b*h2]
objective = cp.Maximize(u)
problem = cp.Problem(objective,const)
problem.solve(verbose=False)
if problem.status != 'optimal':
print(f"PROBLEM INFEASIBLE at :{i}, a:{a.value}, b:{b.value}")
return i, xs, ts
break
x = x + xdot(x,u.value)*dt
t += dt
xs.append(x)
ts.append(t)
fig.canvas.draw()
fig.canvas.flush_events()
car1.set_offsets([t,0])
car2.set_offsets([1+c*t,0])
vehicle.set_offsets([x,0])
writer.grab_frame()
if i==N-1:
print("Problem Successful",t)
return i+1, xs, ts
# -
#x=0.8, c = 0.7
x0 = 0.5
e0 = 0.5
C = 0.3
A = 2.5
B = 4.5
_, x1, t1 = simulate_simple(x0,A,B,C)
#x=0.8, c = 0.7
x0 = 0.5
e0 = 0.5
C = 0.3
A = 0.5
B = 1.5
_, x2, t2 = simulate_simple(x0,A,B,C)
#x=0.8, c = 0.7
x0 = 0.5
e0 = 0.5
C = 0.3
A = 1.0
B = 3.0
_, x3, t3 = simulate_simple(x0,A,B,C)
#x=0.8, c = 0.7
x0 = 0.5
e0 = 0.5
C = 0.3
A = 4.0
B = 1.0
_, x4, t4 = simulate_simple(x0,A,B,C)
#x=0.8, c = 0.7
x0 = 0.1
e0 = 0.5
C = 0.3
A = 20.4
B = 1.0
_, x5, t5 = simulate_simple(x0,A,B,C)
#x=0.8, c = 0.7
x0 = 0.1
e0 = 0.5
C = 0.3
A = 2.5
B = 4.5
_, x6, t6 = simulate_simple(x0,A,B,C)
# +
import matplotlib
font = {'family' : 'normal',
'weight' : 'normal',
'size' : 16}
matplotlib.rc('font', **font)
plt.plot(t1,x1,label='a=2.5,b=4.5')
plt.plot(t2,x2,label='a=0.5,b=1.5')
plt.plot(t3,x3,label='a=1.0,b=3.0')
plt.plot(t4,x4,label='a=3.0,b=1.0')
plt.plot(t5,x5,label='a=20.4,b=1.0')
plt.plot(t6,x6,label='a=2.5,b=4.5')
tmax = max(max(t1),max(t2))
t = np.linspace(0,tmax,30)
b1 = t
b2 = 1 + C*t
plt.axhline(y=0.0, color='k', linestyle='-')
plt.axvline(x=0.0, color='k', linestyle='-')
plt.plot(t,b1,'k--')
plt.plot(t,b2,'k--')
plt.xlabel('time')
plt.ylabel('x')
plt.legend()
plt.savefig("paths_example1.png")
plt.savefig("paths_example1.eps")
# -
#x=0.8, c = 0.7
x0 = 0.5
e0 = 0.5
C = 0.3
A = 2.5
B = 4.5
_, x_1, t_1 = simulate_simple(x0,A,B,C,movie_name='param1.mp4')
#x=0.8, c = 0.7
x0 = 0.5
e0 = 0.5
C = 0.3
A = 1.0
B = 3.0
_, x_2, t_2 = simulate_simple(x0,A,B,C,movie_name='param2.mp4')
#x=0.8, c = 0.7
x0 = 0.5
e0 = 0.5
C = 0.3
A = 3.0
B = 1.0
_, x_3, t_3 = simulate_simple(x0,A,B,C,movie_name='param3.mp4')
nsize = 20
x0 = 0.1
c = 0.7
A = np.linspace(0,5,nsize)
B = np.linspace(0,5,nsize)
horizons = np.zeros((nsize,nsize))
for i in range(nsize):
for j in range(nsize):
time_till_infeasibility , _, _ = simulate_simple(x0,A[i],B[j],c)
horizons[i,j] = time_till_infeasibility
print("DONE")
c = 0.7
fig2, ax2 = plt.subplots(1)
pos = ax2.imshow(horizons/(1.0/(1-c)),cmap='RdBu', interpolation='bilinear',extent=[0,5,5,0], vmin=0, vmax=100)
fig2.colorbar(pos, ax=ax2, label='Time Steps to infeasibility')
ax2.set_ylabel("a")
ax2.set_xlabel("b")
plt.show()
nsize = 5
x0 = 0.5
c = 0.7
A = np.linspace(0,5,nsize)
B = np.linspace(0,5,nsize)
horizons2 = np.zeros((nsize,nsize))
for i in range(nsize):
for j in range(nsize):
time_till_infeasibility , _, _ = simulate_simple(x0,A[i],B[j],c)
horizons2[i,j] = time_till_infeasibility
print("DONE")
c = 0.7
fig3, ax3 = plt.subplots(1)
pos3 = ax3.imshow(horizons2/(1.0/(1-c)),cmap='RdBu', interpolation='bilinear',extent=[0,5,5,0], vmin=0, vmax=100)
fig3.colorbar(pos3, ax=ax3, label='Time Steps to infeasibility')
ax3.set_ylabel("a")
ax3.set_xlabel("b")
plt.show()
|
car_example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # write errors to file (doesn't work in Jupyter, only outside of Jupyter)
# +
import datetime
import logging
LOG_FILENAME = 'errors_log.log'
logging.basicConfig(filename=LOG_FILENAME, level=logging.DEBUG)
logging.debug('Started run, time: ' + str(datetime.datetime.now()))
try:
print('run program')
variable = error_variable # force an error
logging.debug('Finished run, time: ' + str(datetime.datetime.now()))
except:
logging.exception('Got exception on main handler, time: ' + str(datetime.datetime.now()))
raise # use "raise" to exit program right away, without finishing
print('rest of program')
# -
# # write errors to Jupyter Notebook
# +
import datetime
import logging
logger = logging.getLogger()
logging.basicConfig(level=logging.DEBUG)
logging.debug('Started run, time: ' + str(datetime.datetime.now()))
try:
print('run program')
variable = error_variable # will error out
logging.debug('Finished run, time: ' + str(datetime.datetime.now()))
except:
logging.exception('Got exception on main handler, time: ' + str(datetime.datetime.now()))
# raise # use "raise" to exit program right away, without finishing
print('rest of program')
# -
|
code/.ipynb_checkpoints/6_error_handling-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="K4i5n883p_Xf"
# _Lambda School Data Science — Linear Models_
#
# # Doing Linear Regression
#
# ### Objectives
# - arrange data into X features matrix and y target vector
# - use scikit-learn for linear regression
# - use regression metric: MAE
# - do one-hot encoding
# - scale features
#
# ### Contents
# 1. Libraries
# 2. Pre-read
# 3. Process
# 4. Project
# + [markdown] colab_type="text" id="JHJ8JQjGTYro"
# # Libraries
# + [markdown] colab_type="text" id="LWZlqRnfl4sK"
# ### Install [category_encoders](http://contrib.scikit-learn.org/categorical-encoding/) (version 2+)
# - Local Anaconda: `conda install -c conda-forge category_encoders`
# - Google Colab: `pip install category_encoders`
# + colab={} colab_type="code" id="YS2w89kdl_2g"
# !pip install category_encoders
# + [markdown] colab_type="text" id="g74Pk4y0mA5b"
# ### Install [pandas-profiling](https://github.com/pandas-profiling/pandas-profiling) (version 2+)
# - `pip install -U pandas-profiling`
# + colab={} colab_type="code" id="5qg1pSAHl9Fg"
# # !pip install -U pandas-profiling
# + [markdown] colab_type="text" id="i6O0KEmmQ7OM"
# # Pre-reads
#
# #### [<NAME>, Python Data Science Handbook, Chapter 5.2, Introducing Scikit-Learn](https://jakevdp.github.io/PythonDataScienceHandbook/05.02-introducing-scikit-learn.html).
#
# Read up through “Supervised learning example: Simple linear regression”. You can stop when you get to “Supervised learning example: Iris classification.”
#
# + [markdown] colab_type="text" id="EZ7Oh030150T"
# # Process
#
# #### <NAME>, [Becoming a Data Scientist, PyData DC 2016 Talk](https://www.becomingadatascientist.com/2016/10/11/pydata-dc-2016-talk/)
#
# 
#
# + [markdown] colab_type="text" id="IEZu7RSd0O3w"
# ## Business Question --> Data Question --> Data Answer (for Supervised Learning)
#
# #### <NAME>, [Deep Learning with Python](https://github.com/fchollet/deep-learning-with-python-notebooks/blob/master/README.md), Chapter 4: Fundamentals of machine learning, "A universal workflow of machine learning"
#
# > **1. Define the problem at hand and the data on which you’ll train.** Collect this data, or annotate it with labels if need be.
#
# > **2. Choose how you’ll measure success on your problem.** Which metrics will you monitor on your validation data?
#
# > **3. Determine your evaluation protocol:** hold-out validation? K-fold validation? Which portion of the data should you use for validation?
#
# > **4. Develop a first model that does better than a basic baseline:** a model with statistical power.
#
# > **5. Develop a model that overfits.** The universal tension in machine learning is between optimization and generalization; the ideal model is one that stands right at the border between underfitting and overfitting; between undercapacity and overcapacity. To figure out where this border lies, first you must cross it.
#
# > **6. Regularize your model and tune its hyperparameters, based on performance on the validation data.** Repeatedly modify your model, train it, evaluate on your validation data (not the test data, at this point), modify it again, and repeat, until the model is as good as it can get.
#
# > **Iterate on feature engineering: add new features, or remove features that don’t seem to be informative.** Once you’ve developed a satisfactory model configuration, you can train your final production model on all the available data (training and validation) and evaluate it one last time on the test set.
#
# + [markdown] colab_type="text" id="kJhnVFGQRXS0"
# ## Define the data on which you'll train / Add new features or remove features
#
# #### <NAME>, [Python Data Science Handbook, Chapter 5.2, Introducing Scikit-Learn](https://jakevdp.github.io/PythonDataScienceHandbook/05.02-introducing-scikit-learn.html), Data Representation in Scikit-Learn
#
# > The best way to think about data within Scikit-Learn is in terms of tables of data.
#
# 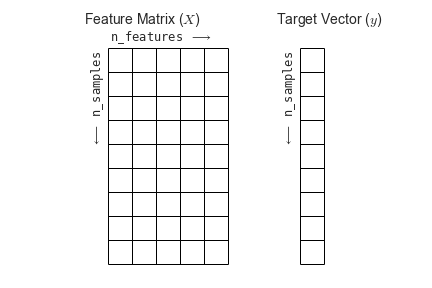
#
# > The samples (i.e., rows) always refer to the individual objects described by the dataset. For example, the sample might be a flower, a person, a document, an image, a sound file, a video, an astronomical object, or anything else you can describe with a set of quantitative measurements.
#
# > The features (i.e., columns) always refer to the distinct observations that describe each sample in a quantitative manner.
#
# > The information can be thought of as a two-dimensional numerical array or matrix, which we will call the _features matrix._ By convention, this features matrix is often stored in a variable named `X`. The features matrix is assumed to be two-dimensional, with shape `[n_samples, n_features]`, and is most often contained in a NumPy array or a Pandas `DataFrame`, though some Scikit-Learn models also accept SciPy sparse matrices.
#
# > In addition to the feature matrix `X`, we also generally work with a label or target array, which by convention we will usually call `y`. The target array is usually one dimensional, with length `n_samples`, and is generally contained in a NumPy array or Pandas `Series`.
#
# > Often one point of confusion is how the target array differs from the other features columns. The distinguishing feature of the target array is that it is usually the quantity we want to _predict from the data:_ in statistical terms, it is the dependent variable.
#
# #### Google Developers, [Machine Learning Glossary](https://developers.google.com/machine-learning/glossary/#l)
#
# > Each example in a labeled dataset consists of one or more features and a label.
#
# > For instance, in a housing dataset, the features might include the number of bedrooms, the number of bathrooms, and the age of the house, while the label might be the house's price.
#
# > In a spam detection dataset, the features might include the subject line, the sender, and the email message itself, while the label would probably be either "spam" or "not spam."
#
# #### Wikipedia, [Feature Engineering](https://en.wikipedia.org/wiki/Feature_engineering)
#
# > "Some machine learning projects succeed and some fail. What makes the difference? Easily the most important factor is the features used." — <NAME>, ["A Few Useful Things to Know about Machine Learning"](https://homes.cs.washington.edu/~pedrod/papers/cacm12.pdf)
#
# > "Coming up with features is difficult, time-consuming, requires expert knowledge. 'Applied machine learning' is basically feature engineering." — <NAME>, [Machine Learning and AI via Brain simulations](https://forum.stanford.edu/events/2011/2011slides/plenary/2011plenaryNg.pdf)
#
# > Feature engineering is the process of using domain knowledge of the data to create features that make machine learning algorithms work.
# + [markdown] colab_type="text" id="lSZBdwNg1Vvj"
# ## Determine evaluation protocol
#
# #### <NAME>, [Model Evaluation]( https://sebastianraschka.com/blog/2018/model-evaluation-selection-part4.html)
# > <img src="https://sebastianraschka.com/images/blog/2018/model-evaluation-selection-part4/model-eval-conclusions.jpg" width="600">
# + [markdown] colab_type="text" id="HFAHa5dD3BQt"
# ## Develop a first model that does better than a basic baseline
#
# ### Why begin with baselines?
#
# [My mentor](https://www.linkedin.com/in/jason-sanchez-62093847/) [taught me](https://youtu.be/0GrciaGYzV0?t=40s):
#
# >***Your first goal should always, always, always be getting a generalized prediction as fast as possible.*** You shouldn't spend a lot of time trying to tune your model, trying to add features, trying to engineer features, until you've actually gotten one prediction, at least.
#
# > The reason why that's a really good thing is because then ***you'll set a benchmark*** for yourself, and you'll be able to directly see how much effort you put in translates to a better prediction.
#
# > What you'll find by working on many models: some effort you put in, actually has very little effect on how well your final model does at predicting new observations. Whereas some very easy changes actually have a lot of effect. And so you get better at allocating your time more effectively.
#
# My mentor's advice is echoed and elaborated in several sources:
#
# [Always start with a stupid model, no exceptions](https://blog.insightdatascience.com/always-start-with-a-stupid-model-no-exceptions-3a22314b9aaa)
#
# > Why start with a baseline? A baseline will take you less than 1/10th of the time, and could provide up to 90% of the results. A baseline puts a more complex model into context. Baselines are easy to deploy.
#
# [Measure Once, Cut Twice: Moving Towards Iteration in Data Science](https://blog.datarobot.com/measure-once-cut-twice-moving-towards-iteration-in-data-science)
#
# > The iterative approach in data science starts with emphasizing the importance of getting to a first model quickly, rather than starting with the variables and features. Once the first model is built, the work then steadily focuses on continual improvement.
#
# [*Data Science for Business*](https://books.google.com/books?id=4ZctAAAAQBAJ&pg=PT276), Chapter 7.3: Evaluation, Baseline Performance, and Implications for Investments in Data
#
# > *Consider carefully what would be a reasonable baseline against which to compare model performance.* This is important for the data science team in order to understand whether they indeed are improving performance, and is equally important for demonstrating to stakeholders that mining the data has added value.
#
# ### What does baseline mean?
#
# Baseline is an overloaded term, as you can see in the links above. Baseline has multiple meanings:
#
# #### The score you'd get by guessing a single value
#
# > A baseline for classification can be the most common class in the training dataset.
#
# > A baseline for regression can be the mean of the training labels. —[<NAME>](https://twitter.com/koehrsen_will/status/1088863527778111488)
#
# #### The score you'd get by guessing in a more granular way
#
# > A baseline for time-series regressions can be the value from the previous timestep.
#
# #### Fast, first models that beat guessing
#
# What my mentor was talking about.
#
# #### Complete, tuned "simpler" model
#
# Can be simpler mathematically and computationally. For example, Logistic Regression versus Deep Learning.
#
# Or can be simpler for the data scientist, with less work. For example, a model with less feature engineering versus a model with more feature engineering.
#
# #### Minimum performance that "matters"
#
# To go to production and get business value.
#
# #### Human-level performance
#
# Your goal may to be match, or nearly match, human performance, but with better speed, cost, or consistency.
#
# Or your goal may to be exceed human performance.
# + [markdown] colab_type="text" id="aOW2hsiDBIk9"
# ## Use scikit-learn to fit a model
#
# #### <NAME>, [Python Data Science Handbook, Chapter 5.2, Introducing Scikit-Learn](https://jakevdp.github.io/PythonDataScienceHandbook/05.02-introducing-scikit-learn.html), Scikit-Learn's Estimator API
#
# > Most commonly, the steps in using the Scikit-Learn estimator API are as follows (we will step through a handful of detailed examples in the sections that follow).
#
# > 1. Choose a class of model by importing the appropriate estimator class from Scikit-Learn.
# > 2. Choose model hyperparameters by instantiating this class with desired values.
# > 3. Arrange data into a features matrix and target vector following the discussion above.
# > 4. Fit the model to your data by calling the `fit()` method of the model instance.
# > 5. Apply the Model to new data: For supervised learning, often we predict labels for unknown data using the `predict()` method.
# + [markdown] colab_type="text" id="T1hPspa5mkWT"
# # Project: Predict NYC apartment rent 🏠💸
#
# You'll use a real-world data with rent prices for a subset of apartments in New York City!
#
# + [markdown] colab_type="text" id="UbqeAzl9TK_y"
# ## Define the data on which you'll train
#
# - Get the data
# - What's the target?
# - Regression or classification?
# + colab={} colab_type="code" id="qEeVFTa0VWDE"
LOCAL = '../data/nyc/nyc-rent-2016.csv'
WEB = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Linear-Models/master/data/nyc/nyc-rent-2016.csv'
import pandas as pd
df = pd.read_csv(LOCAL)
assert df.shape == (48300, 34)
# + colab={} colab_type="code" id="MGzhihMGfp1p"
import pandas_profiling as pdp
# + [markdown] colab_type="text" id="nnJ6ioiesWsT"
# ## DO TRAIN/TEST SPLIT
#
# For this project, we'll split based on time.
#
# - Use data from April & May 2016 to train.
# - Use data from June 2016 to test.
#
# + colab={} colab_type="code" id="FYLE4Dzwskn-"
df['created'] = pd.to_datetime(df['created'], infer_datetime_format = True)
# + [markdown] colab_type="text" id="GMmPjCEWTXFn"
# ## Begin with baselines for regression
# + colab={} colab_type="code" id="Ap063j12tEWJ"
df['month'] = df['created'].dt.month
# -
train = df.query('month < 6') #df[df['month']<6]
test = df.query('month == 6') #df[df['month']==6]
train.shape, test.shape
train['price'].mean()
# +
import numpy as np
from sklearn.metrics import mean_absolute_error as mae
y_test = test['price']
y_pred = [train['price'].mean()]*len(y_test)
print(len(y_test), len(y_pred))
# -
print(mae(y_test,y_pred))
# + [markdown] colab_type="text" id="i3hDX7yUTbix"
# ## Use scikit-learn for linear regression, with 1 feature
#
#
#
# + [markdown] colab_type="text" id="HFWAop61CgCq"
# Follow the process from <NAME>, [Python Data Science Handbook, Chapter 5.2, Introducing Scikit-Learn](https://jakevdp.github.io/PythonDataScienceHandbook/05.02-introducing-scikit-learn.html), Scikit-Learn's Estimator API
#
# ### Choose a class of model by importing the appropriate estimator class from Scikit-Learn
#
# + colab={} colab_type="code" id="OMbOVWEDCfmO"
from sklearn.linear_model import LinearRegression
# + [markdown] colab_type="text" id="vATSdu5oD5NQ"
# ### Choose model hyperparameters by instantiating this class with desired values
#
# Refer to scikit-learn documentation to see what model hyperparameters you can choose. For example: [sklearn.linear_model.LinearRegression](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LinearRegression.html)
# + colab={} colab_type="code" id="CexmSzauEBnu"
linr = LinearRegression()
linr
# + [markdown] colab_type="text" id="oEsa7jtHC0L5"
# ### Arrange data into X features matrix and y target vector
# + colab={} colab_type="code" id="euIG2-5P_sdZ"
feature = ['bedrooms']
target = 'price'
X_train = train[feature]
y_train = train[target]
X_test = test[feature]
y_test = test[target]
# -
X_train.shape, y_train.shape
# + [markdown] colab_type="text" id="K8s3-WYWEKxN"
# ### Fit the model to your data by calling the `fit()` method of the model instance
# + colab={} colab_type="code" id="XTLnEzwUENb5"
linr.fit(X_train, y_train)
# + [markdown] colab_type="text" id="HynZZRL7ESvx"
# ### Apply the Model to new data: For supervised learning, often we predict labels for unknown data using the `predict()` method
# + colab={} colab_type="code" id="mo6h1SnMr9kP"
n_bedrm = 2
linr.predict([[n_bedrm]])
# -
#Slope, increase/decrease per feature
linr.coef_
#y intercept at 0
linr.intercept_
y_pred = linr.predict(X_test)
y_pred
# + [markdown] colab_type="text" id="6l-WbivHIwHh"
# ## Use regression metric: MAE
# + colab={} colab_type="code" id="r5G7hDzGG0B0"
mae(y_test,y_pred)
# +
import matplotlib.pyplot as plt
plt.scatter(X_train, y_train, alpha=0.01)
plt.plot(X_test, y_pred)
plt.xlabel('# of bedrooms')
plt.ylabel('Rent Price')
plt.title('LR with # of BR as a feature');
# + [markdown] colab_type="text" id="N_KGM3LOHyrW"
# ## Use scikit-learn for linear regression, with 2 features
# + [markdown] colab_type="text" id="Z_79qOeAH2ZU"
# Follow the process from <NAME>, [Python Data Science Handbook, Chapter 5.2, Introducing Scikit-Learn](https://jakevdp.github.io/PythonDataScienceHandbook/05.02-introducing-scikit-learn.html), Scikit-Learn's Estimator API
#
# ### Choose a class of model by importing the appropriate estimator class from Scikit-Learn
# + colab={} colab_type="code" id="E26qduGiH2_y"
from sklearn.linear_model import LinearRegression
# + [markdown] colab_type="text" id="--7julXYH3oC"
# ### Choose model hyperparameters by instantiating this class with desired values
# + colab={} colab_type="code" id="BkkoMxbsIXLR"
linr2 = LinearRegression()
# + [markdown] colab_type="text" id="pJNRFvK9IeWU"
# ### Arrange data into X features matrix and y target vector
# + colab={} colab_type="code" id="2290BJszIgrb"
features = ['bedrooms', 'bathrooms']
target2 = 'price'
X_train2 = train[features]
y_train2 = train[target]
X_test2 = test[features]
y_test2 = test[target]
# -
X_train2.shape, y_train.shape
# + [markdown] colab_type="text" id="BOa5Uj4jIjDR"
# ### Fit the model to your data by calling the `fit()` method of the model instance
# + colab={} colab_type="code" id="VZAUSsY0IjWa"
linr2.fit(X_train2, y_train2)
# + [markdown] colab_type="text" id="8GEyW2B3Imr2"
# ### Apply the Model to new data: For supervised learning, often we predict labels for unknown data using the `predict()` method
# + colab={} colab_type="code" id="3ubKZVRJInLV"
y_pred = linr2.predict(X_test2)
# + [markdown] colab_type="text" id="MDbG8jreI8Ip"
# ## Use regression metric: MAE
# + colab={} colab_type="code" id="xCs--47RI-He"
mae(y_test2, y_pred)
# -
linr2.coef_
linr2.intercept_
# + [markdown] colab_type="text" id="BM-KUzX8RqsS"
# ## Do one-hot encoding of categorical features
# + [markdown] colab_type="text" id="VqeBG7OoR72b"
# ### Which features are non-numeric?
# + colab={} colab_type="code" id="9ic36yUPRp2l"
train.describe(exclude='number')
# -
train['description'].head()
# + [markdown] colab_type="text" id="aN89gf1QSJG3"
# ### Check "cardinality" of non-numeric features
#
# [Cardinality](https://simple.wikipedia.org/wiki/Cardinality) means the number of unique values that a feature has:
# > In mathematics, the cardinality of a set means the number of its elements. For example, the set A = {2, 4, 6} contains 3 elements, and therefore A has a cardinality of 3.
#
# "One-hot encoding" adds a dimension for each unique value of each categorical feature. So, it may not be a good choice for "high cardinality" categoricals that have dozens, hundreds, or thousands of unique values.
# + colab={} colab_type="code" id="IkmTd6W3SNey"
import category_encoders as ce
# + [markdown] colab_type="text" id="ApYuyrb8SaDp"
# ### Explore `interest_level` feature
# + colab={} colab_type="code" id="ipS3vkvcRwub"
train['interest_level'].head()
# + [markdown] colab_type="text" id="QGM1e6ThSp5K"
# ### Encode `interest_level` feature
# + colab={} colab_type="code" id="28TRmEX_SuzM"
encoder = ce.OneHotEncoder(use_cat_names = True)
encoded = encoder.fit_transform(train['interest_level'])
# -
encoded
# + [markdown] colab_type="text" id="V15ZztGdccx5"
# ## Do one-hot encoding & Scale features,
# within a complete model fitting workflow.
#
# ### Why and how to scale features before fitting linear models
#
# Scikit-Learn User Guide, [Preprocessing data](https://scikit-learn.org/stable/modules/preprocessing.html)
# > Standardization of datasets is a common requirement for many machine learning estimators implemented in scikit-learn; they might behave badly if the individual features do not more or less look like standard normally distributed data: Gaussian with zero mean and unit variance.
#
# > The `preprocessing` module further provides a utility class `StandardScaler` that implements the `Transformer` API to compute the mean and standard deviation on a training set. The scaler instance can then be used on new data to transform it the same way it did on the training set.
#
# ### How to use encoders and scalers in scikit-learn
# - Use the **`fit_transform`** method on the **train** set
# - Use the **`transform`** method on the **validation** set
# + colab={} colab_type="code" id="Jf6r8sCDccDv"
from sklearn.preprocessing import StandardScaler as sc
feature3 = [
'bedrooms',
'bathrooms',
'latitude',
'longitude',
'interest_level',
'swimming_pool'
]
target3 = 'price'
X_train3 = train[feature3]
y_train3 = train[target3]
X_test3 = test[feature3]
y_test3 = test[target3]
# -
train[target3].shape
X_train3.shape, y_train3.shape
encoder2 = ce.OneHotEncoder(use_cat_names = True)
X_train3 = encoder2.fit_transform(X_train3)
X_test3 = encoder2.fit_transform(X_test3)
scaler = sc()
X_train3 = scaler.fit_transform(X_train3)
X_test3 = scaler.fit_transform(X_test3)
X_train3
X_train3.shape, y_train3.shape
linr3 = LinearRegression()
linr3.fit(X_train3, y_train3)
y_pred = linr3.predict(X_test3)
y_pred
mae(y_test3,y_pred)
# + [markdown] colab_type="text" id="70KA2iiTao3Z"
# # Assignment
# 1. Start a clean notebook. Follow the processes taught today.
# 2. Do train/test split. Use data from April & May 2016 to train. Use data from June 2016 to test.
# 3. Begin with baseline for regression.
# 4. Select two or more features.
# 5. Do one-hot encoding. (Remember it may not work with high cardinality categoricals.)
# 4. Use scikit-learn to fit a Linear Regression model on the train data.
# 5. Apply the model to predict rent prices for the test data.
# 6. Get the mean absolute error for the test data.
# 7. Get the model's coefficients and intercept.
# 8. Commit your notebook to your fork of the GitHub repo.
#
# _What's the best test MAE you can get? Share your score and features used with your cohort on Slack!_
#
# ### Stretch Goals
# - Try at least 3 different feature combinations.
# - Get regression metrics RMSE, MAE, and $R^2$, for both the train and test data.
# - [Engineer new features!](https://en.wikipedia.org/wiki/Feature_engineering)
#
# > "Some machine learning projects succeed and some fail. What makes the difference? Easily the most important factor is the features used." — <NAME>, ["A Few Useful Things to Know about Machine Learning"](https://homes.cs.washington.edu/~pedrod/papers/cacm12.pdf)
#
# > "Coming up with features is difficult, time-consuming, requires expert knowledge. 'Applied machine learning' is basically feature engineering." — <NAME>, [Machine Learning and AI via Brain simulations](https://forum.stanford.edu/events/2011/2011slides/plenary/2011plenaryNg.pdf)
#
# > Feature engineering is the process of using domain knowledge of the data to create features that make machine learning algorithms work.
#
# - Try different [scikit-learn scalers](https://scikit-learn.org/stable/modules/preprocessing.html)
# - Try [scikit-learn pipelines](https://scikit-learn.org/stable/modules/compose.html):
#
# > Pipeline can be used to chain multiple estimators into one. This is useful as there is often a fixed sequence of steps in processing the data, for example feature selection, normalization and classification. Pipeline serves multiple purposes here:
#
# > - **Convenience and encapsulation.** You only have to call fit and predict once on your data to fit a whole sequence of estimators.
# > - **Joint parameter selection.** You can grid search over parameters of all estimators in the pipeline at once.
# > - **Safety.** Pipelines help avoid leaking statistics from your test data into the trained model in cross-validation, by ensuring that the same samples are used to train the transformers and predictors.
|
module2-doing-linear-regression/doing_linear_regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Beginner's Python: Session Two - Politics and Social Sciences Questions
# +
#DO NOT change code in this cell but run it before going further
import matplotlib.pyplot as plt
def plot_demographics(continent_ls, continent_names, years):
fig, ax = plt.subplots(figsize = (10,8))
for i, continent in enumerate(continent_ls):
plt.plot(years[:12], continent[:12], '-o', label = continent_names[i])
if len(continent)>12:
plt.plot(years[11:], continent[11:], '-o', color = 'grey')
plt.title("World population by continent (in millions)")
plt.legend()
plt.show()
# -
# In the cell below there are seven lists defined, all of them together represent world population by continent and by year:
# - `years` is a list containing the years in which the population was recorded
# - `africa`, `asia`, ... , `oceania` are lists containing the number of population (in millions) of a given continent with values ordered respectively to the years in `years`
# </ul>
# example: the third value in <code>years</code> is 1700. The third value in <code>africa</code> is 106. This means that the population of Africa in 1700 was 106 milion people.
# +
years = ['1500', '1600', '1700', '1750', '1800', '1850', '1900', '1950', '1999', '2008', '2010', '2012']
africa = [86, 114, 106, 106, 107, 111, 133, 221, 783, 973, 1022, 1052]
asia = [282, 350, 411, 502, 635, 809, 947, 1402, 3700, 4054, 4164, 4250]
europe = [168, 170, 178, 190, 203, 276, 408, 547, 675, 732, 738, 740]
latin_america = [40, 20, 10, 16, 24, 38, 74, 167, 508, 577, 590, 603]
north_america = [6, 3, 2, 2, 7, 26, 82, 172, 312, 337, 345, 351]
oceania = [3, 3, 3, 2, 2, 2, 6, 13, 30, 34, 37, 38]
# -
# Using the `min()` and `max()` functions find what was the smallest and the largest number of population in Europe. Save these values in variables called `min_europe`, `max_europe`. Then print the following text: *Between year 1500 and 2012 the smallest recorded population of europe was: xxx milion people and the largest was: xxx milion people*
# Create two lists:
# - `continent_names` - a list containing the names of all continents (as strings)
# - `continent_ls` - a list contating the seven lists from the cell above in the same order as in `continent_names`
#
# **NOTE:** there is nothing wrong in creating list of lists. In fact, this is a very common procedure. If you can create a list of strings or list of integers then why you shouldn't be able to create a list of lists?
# If you did correctly your previous task then after running the cell below you should see a graph representing the world population by continents between 1500 and 2012. But there is something not quite right about this graph. Can you spot the mistake?
plot_demographics(continent_ls, continent_names, years)
# The problem is with the distances on the x axis. The distance between 1500 and 1600 and the distance between 2010 and 2012 are the same on this plot. This is a serious breach of rules of a good statistical data visualization. Do you know what might be the reason for that? Why Python does not interpret the values of years correctly?
# The answer is the **data types**. Note that the values in the `years` list are strings, this means that all of the years are treated as a text data rather than a numerical data. Which data type would do better?
#
# We would like to convert the `years` list of strings to a a new one containing the years in the same order but with a proper data type. This can be done by hand or using a shortcut - **list comprehensions**. We will learn more about them later on, but for now examine the example below.
# +
#example - conversion between list of doubles to list of integers
double_ls = [2.5, 2.8, 3.9, 10.1] #this is a list of doubles
int_ls = [int(x) for x in double_ls] #using this expression we can convert list of doubles to list of integers
# -
# Now try to do something similar with the `years` list. Do not create a new variable, instead overwrite the existing list.
# Now, the plot of world population should look a lot better. Execute the cell below to see if it works.
plot_demographics(continent_ls, continent_names, years)
# WHO has made some predictions for the number of population in 2150. These are presented in the table bellow:
# continent | population in 2150 (in millions) |
# --- | ---
# Africa | 2308
# Asia | 5561
# Europe| 517
# Latin America | 912
# North America | 938
# Oceania | 51
#
# Append the year 2150 to the `years` list and append the values of predicted population in 2150 to the corresponding lists defined previously.
# Now plot the extended results.
plot_demographics(continent_ls, continent_names, years)
|
session-two/subject_questions/session_two_politics_exercises.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] deletable=true editable=true
# # Filtering of Data in Insights Parsers and Rules
# In this tutorial we will investigate filters in insights-core, what they are, how they affect your components and how you can use them in your code. Documentation on filters can be found in the [insights-core documentation](https://insights-core.readthedocs.io/en/latest/api_index.html#module-insights.core.filters).
#
# The primary purposes of filters are:
#
# 1. to prevent the collection of sensitive information while enabling the collection of necessary information for analysis, and;
# 2. to reduce the amount of information collected.
#
# Filters are typically added in rule modules since the purpose of a rule is to analyze particular information and identify a problem, potential problem or fact about the system. A filter may also be added in a parse modules if it is required to enable parsing of the data. We will discuss this further when we look at the example. Filters added by rules and parsers are applied when the data is collected from a system. They are combined so that if they are added from multiple rules and parsers, each rule will receive all information that was collected by all filters for a given source. An example will help demonstrate this.
#
# Suppose you write some rules that needs information from `/var/log/messages`. This file could be very large and contain potentially sensitive information, so it is not desirable to collect the entire file. Let's say *rule_a* needs messages that indicate `my_special_process` has failed to start. And another rule, *rule_b* needs messages that indicate that `my_other_process` had the errors `MY_OTHER_PROCESS: process locked` or `MY_OTHER_PROCESS: memory exceeded`. Then the two rules could add the following filters to ensure that just the information they need is collected:
#
# *rule_a*:
# ```python
# add_filter(Specs.messages, 'my_special_process')
# ```
#
# *rule_b*:
# ```python
# add_filter(Specs.messages, ['MY_OTHER_PROCESS: process locked',
# 'MY_OTHER_PROCESS: memory exceeded'])
# ```
#
# The effect of this would be that when `/var/log/messages` is collected, the filters would be applied and only the lines containing the strings `'my_special_process'`, `'MY_OTHER_PROCESS: process locked'`, or `'MY_OTHER_PROCESS: memory exceeded'` would be collected. This significantly reduces the size of the data and the chance that sensitive information in `/var/log/messages` might be collected.
#
# While there are significant benefits to filtering, you must be aware that a datasource is being filtered or your rules could fail to identify a condition that may be present on a system. For instance suppose a rule *rule_c* also needs information from `/var/log/messages` about `process_xyz`. If *rule_c* runs with other rules like *rule_a* or *rule_b* then it would never see lines containing `"process_xyz"` appearing in `/var/log/messages` unless it adds a new filter. When any rule or parser adds a filter to a datasource, that data will be filtered for all components, not just the component adding the filter. Because of this it is important to understand when a datasource is being filtered so that your rule will function properly and include its own filters if needed.
# + [markdown] deletable=true editable=true
# ## Exploring Filters
# ### Unfiltered Data
# Suppose we want to write a rule that will evaluate the contents of the configuration file `death_star.ini` to determine if there are any vulnerabilities. Since this is a new data source that is not currently collected by insights-core we'll need to add three elements to collect, parse and evaluate the information.
# + deletable=true editable=true
""" Some imports used by all of the code in this tutorial """
import sys
sys.path.insert(0, "../..")
from __future__ import print_function
import os
from insights import run
from insights.specs import SpecSet
from insights.core import IniConfigFile
from insights.core.plugins import parser, rule, make_fail
from insights.core.spec_factory import simple_file
# + [markdown] deletable=true editable=true
# First we'll need to add a specification to collect the configuration file. Note that for purposes of this tutorial we are collecting from a directory where this notebook is located. Normally the file path would be an absolute path on your system or in an archive.
# + deletable=true editable=true
class Specs(SpecSet):
"""
Define a new spec to collect the file we need.
"""
death_star_config = simple_file(os.path.join(os.getcwd(), 'death_star.ini'), filterable=True)
# + [markdown] deletable=true editable=true
# Next we'll need to add a parser to parse the file being collected by the spec. Since this file is in INI format and insights-core provides the IniConfigFile parser, we can just use that to parse the file. See [the parser documentation](https://insights-core.readthedocs.io/en/latest/api_index.html#insights.core.IniConfigFile) to find out what methods that parser provides.
# + deletable=true editable=true
@parser(Specs.death_star_config)
class DeathStarCfg(IniConfigFile):
"""
Define a new parser to parse the spec. Since the spec is a standard INI format we
can use the existing IniConfigFile parser that is provided by insights-core.
See documentation here:
https://insights-core.readthedocs.io/en/latest/api_index.html#insights.core.IniConfigFile
"""
pass
# + [markdown] deletable=true editable=true
# Finally we can write the rule that will examine the contents of the parsed configuration file to determine if there are any vulnerabilities. In this INI file we can find the vulnerabilities by searching for keywords to find one that contains the string `vulnerability`. If any vulnerabilities are found the rule should return information in the form of a `response` that documents the vulnerabilities found, and tags them with the key `DS_IS_VULNERABLE`. If no vulnerabilities are found the rule should just drop out, effectively returning `None`.
# + deletable=true editable=true
@rule(DeathStarCfg)
def ds_vulnerable(ds_cfg):
"""
Define a new rule to look for vulnerable conditions that may be
included in the INI file. If found report them.
"""
vulnerabilities = []
for section in ds_cfg.sections():
print("Section: {}".format(section))
for item_key in ds_cfg.items(section):
print(" {}={}".format(item_key, ds_cfg.get(section, item_key)))
if 'vulnerability' in item_key:
vulnerabilities.append((item_key, ds_cfg.get(section, item_key)))
if vulnerabilities:
return make_fail('DS_IS_VULNERABLE', vulnerabilities=vulnerabilities)
# + [markdown] deletable=true editable=true
# Before we run the rule, lets look at the contents of the configuration file. It is in the format of a typical INI file and contains some interesting information. In particular we see that it does contain a keyword that should match the string we are looking for in the rule, *"major_vulnerability=ray-shielded particle exhaust vent"*. So we expect the rule to return results.
# + deletable=true editable=true
# !cat death_star.ini
# + [markdown] deletable=true editable=true
# Lets run our rule and find out. To run the rule we'll use the `insights.run()` function and as the argument pass in our rule object (note this is not a string but the actual object). The results returned will be an `insights.dr.broker` object that contains all sorts of information about the execution of the rule. You can explore more details of the `broker` in the [Insights Core Tutorial](https://github.com/RedHatInsights/insights-core/blob/master/docs/notebooks/Insights%20Core%20Tutorial.ipynb) notebook.
#
# The `print` statements in our rule provide output as it loops through the configuration file.
# + deletable=true editable=true
results = run(ds_vulnerable)
# + [markdown] deletable=true editable=true
# Now we are ready to look at the results. The results are stored in `results[ds_vulnerable]` where the rule object `ds_vulnerable` is the key into the dictionary of objects that your rule depended upon to execute, such as the parser `DeathStarCfg` and the spec `Spec.death_star_config`. You can see this by looking at those objects in results.
# + deletable=true editable=true
type(results[Specs.death_star_config])
# + deletable=true editable=true
type(results[DeathStarCfg])
# + deletable=true editable=true
type(results[ds_vulnerable])
# + [markdown] deletable=true editable=true
# Now lets look at the rule results to see if they match what we expected.
# + deletable=true editable=true
results[ds_vulnerable]
# + [markdown] deletable=true editable=true
# Success, it worked as we expected finding the vulnerability. Now lets look at how filtering can affect the rule results.
# + [markdown] deletable=true editable=true
# ### Filtering Data
# When we looked at the contents of the file you may have noticed some other interesting information such as this:
# ```
# # Keep this info secret
# [secret_stuff]
# username=dvader
# password=<PASSWORD>
# ```
# As a parser writer, if you know that a file could contain sensitive information, you may choose to filter it in the parser module to avoid collecting it. Usernames, passwords, hostnames, security keys, and other sensitive information should not be collected. In this case the `username` and `password` are in the configuration file, so we should add a filter to this parser to prevent them from being collected.
#
# How do we add a filter and avoid breaking the parser? Each parser is unique, so the parser writer must determine if a filter is necessary, and how to add a filter that will allow the parser to function with a minimal set of data. For instance a Yaml or XML parser might have a difficult time parsing a filtered Yaml or XML file.
#
# For our example, we are using an INI file parser. INI files are structured with sections which are identified as a section name in square brackets like `[section name]`, followed by items like `name` or `name=value`. One possible way to filter an INI file is to add the filter `"["` which will collect all lines with sections but no items. This can be successfully parsed by the INI parser, so that is how we'll filter out this sensitive information in our configuration file. We'll rewrite the parser adding the `add_filter(Specs.death_star_config, '[')` to filter all lines except those with a `'['` string.
# + deletable=true editable=true
from insights.core.filters import add_filter
add_filter(Specs.death_star_config, '[')
@parser(Specs.death_star_config)
class DeathStarCfg(IniConfigFile):
"""
Define a new parser to parse the spec. Since the spec is a standard INI format we
can use the existing IniConfigFile parser that is provided by insights-core.
See documentation here:
https://insights-core.readthedocs.io/en/latest/api_index.html#insights.core.IniConfigFile
"""
pass
# + [markdown] deletable=true editable=true
# Now lets run the rule again and see what happens. Do you expect the same results we got before?
# + deletable=true editable=true
results = run(ds_vulnerable)
results.get(ds_vulnerable, "No results") # Use .get method of dict so we can provide default other than None
# + [markdown] deletable=true editable=true
# Is that what you expected? Notice the output from the `print` statements in the rule, only the section names are printed. That is the result of adding the filter, only lines with `'['` (the sections) are collected and provided to the parser. This means that the lines we were looking for in the rule are no longer there, and that it appears our rule didn't find any vulnerabilities. Next we'll look at how to fix our rule to work with the filtered data.
# + [markdown] deletable=true editable=true
# ### Adding Filters to Rules
# We can add filters to a rule just like we added a filter to the parser, using the `add_filter()` method. The `add_filter` method requires a spec and a string or list/set of strings. In this case our rule is looking for the string `'vulnerability'` so we just need to add that to the filter.
# + deletable=true editable=true
add_filter(Specs.death_star_config, 'vulnerability')
@rule(DeathStarCfg)
def ds_vulnerable(ds_cfg):
"""
Define a new rule to look for vulnerable conditions that may be
included in the INI file. If found report them.
"""
vulnerabilities = []
for section in ds_cfg.sections():
print("Section: {}".format(section))
for item_key in ds_cfg.items(section):
print(" {}={}".format(item_key, ds_cfg.get(section, item_key)))
if 'vulnerability' in item_key:
vulnerabilities.append((item_key, ds_cfg.get(section, item_key)))
if vulnerabilities:
return make_fail('DS_IS_VULNERABLE', vulnerabilities=vulnerabilities)
# + [markdown] deletable=true editable=true
# Now lets run the rule again and see what happens.
# + deletable=true editable=true
results = run(ds_vulnerable)
results.get(ds_vulnerable, "No results") # Use .get method of dict so we can provide default other than None
# + [markdown] deletable=true editable=true
# Now look at the output from the `print` statements in the rule, the item that was missing is now included. By adding the string required by our rule to the spec filters we have successfully included the data needed by our rule to detect the problem. Also, by adding the filter to the parser we have eliminated the sensitive information from the input.
# + [markdown] deletable=true editable=true
# ## Determining if a Spec is Filtered
# When you are developing your rule, you may want to add some code, during development, to check if the spec you are using is filtered. This can be accomplished by looking at the spec in [insights/specs/__init__.py](https://github.com/RedHatInsights/insights-core/blob/master/insights/specs/__init__.py). Each spec is defined here as a `RegistryPoint()` type. If the spec is filtered it will have the parameter `filterable=True`, for example the following indicates that the messages log (`/var/log/messages`) will be filtered:
#
# ```
# messages = RegistryPoint(filterable=True)
# ```
#
# If you need to use a parser that relies on a filtered spec then you need to add your own filter to ensure that your rule will receive the data necessary to evaluate the rule conditions. If you forget to add a filter to your rule, if you include integration tests for your rule, `pytest` will indicate an exception like the following warning you that the `add_filter` is missing:
#
# ```
# telemetry/rules/tests/integration.py:7:
# _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _ _
#
# component = <function report at 0x7fa843094e60>, input_data = <InputData {name:test4-00000}>, expected = None
#
# def run_test(component, input_data, expected=None):
# if filters.ENABLED:
# mod = component.__module__
# sup_mod = '.'.join(mod.split('.')[:-1])
# rps = _get_registry_points(component)
# filterable = set(d for d in rps if dr.get_delegate(d).filterable)
# missing_filters = filterable - ADDED_FILTERS.get(mod, set()) - ADDED_FILTERS.get(sup_mod, set())
# if missing_filters:
# names = [dr.get_name(m) for m in missing_filters]
# msg = "%s must add filters to %s"
# > raise Exception(msg % (mod, ", ".join(names)))
# E Exception: telemetry.rules.plugins.kernel.overcommit must add filters to insights.specs.Specs.messages
#
# ../../insights/insights-core/insights/tests/__init__.py:114: Exception
#
# ```
#
# If you see this exception when you run tests then it means you need to include `add_filter` to your rule.
# + [markdown] deletable=true editable=true
# ## Turning Off Filtering Globally
# There are often times that you would want or need to turn off filtering in order to perform testing or to fully analyze some aspects of a system and diagnose problems. Also if you are running locally on a system you might want to collect all data unfiltered. You can to this by setting the environment variable `INSIGHTS_FILTERS_ENABLED=False` prior to running insights-core. This won't work inside this notebook unless you follow the directions below.
# + deletable=true editable=true
"""
This code will disable all filtering if it is run as the first cell when the notebook
is opened. After the notebook has been started you will need to click on the Kernel
menu and then the restart item, and then run this cell first before all others.
You would need to restart the kernel and then not run this cell to prevent disabling
filters.
"""
import os
os.environ['INSIGHTS_FILTERS_ENABLED'] = 'False'
# + deletable=true editable=true
results = run(ds_vulnerable)
results.get(ds_vulnerable, "No results") # Use .get method of dict so we can provide default other than None
# + [markdown] deletable=true editable=true
# ## Debugging Components
# If you are writing component code you may sometimes not see any results even though you expected them and no errors were displayed. That is because insights-core is catching the exceptions and saving them. In order to see the exceptions you can use the following method to display the results of a run and any errors that occurrerd.
# + deletable=true editable=true
def show_results(results, component):
"""
This function will show the results from run() where:
results = run(component)
run will catch all exceptions so if there are any this
function will print them out with a stack trace, making
it easier to develop component code.
"""
if component in results:
print(results[component])
else:
print("No results for: {}".format(component))
if results.exceptions:
for comp in results.exceptions:
print("Component Exception: {}".format(comp))
for exp in results.exceptions[comp]:
print(results.tracebacks[exp])
# + [markdown] deletable=true editable=true
# Here's an example of this function in use
# + deletable=true editable=true
@rule(DeathStarCfg)
def bad_rule(cfg):
# Force an error here
infinity = 1 / 0
# + deletable=true editable=true
results = run(bad_rule)
# + deletable=true editable=true
show_results(results, bad_rule)
# + deletable=true editable=true
|
docs/notebooks/Filters Tutorial.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ### Average Reward DQN
# \begin{eqnarray}
# \theta_{n+1} &=& \theta_n + a(n)\left(r(X_n,U_n) +\max_v Q(X_{n+1}, v; \theta_n) -f(Q;\theta_n) - Q(X_n, U_n;\theta_n)\right)\times \nonumber \\
# && \Big(\nabla_\theta Q(X_n, U_n;\theta_n) \Big)
# \end{eqnarray}
#
# for $n \geq 0$, where $v_n \in \mbox{Argmax} Q(X_{n+1}, \cdot ; \theta_n)$
# !python dqn_train.py --iter 1000 --expname Exp1
# ### Average Reward FGDQN
#
# \begin{eqnarray}
# \theta_{n+1} &=& \theta_n - a(n)\left(\overline{r(X_n,U_n) +\max_v Q(X_{n+1}, v; \theta_n) -f(Q;\theta_n) - Q(X_n, U_n;\theta_n)}\right)\times \nonumber \\
# && \Big(\nabla_\theta Q(X_{n+1}, v_n; \theta_n) - \nabla_\theta f(Q;\theta_n) - \nabla_\theta Q(X_n, U_n; \theta_n)\Big)
# \end{eqnarray}
#
# for $n \geq 0$, where $v_n \in \mbox{Argmax} Q(X_{n+1}, \cdot ; \theta_n)$
# !python fgdqn_train.py --iter 1000 --expname Exp1
# +
import pandas
import csv
import matplotlib.pyplot as plt
import numpy as np
colnames = ["Loss","State","Policy","Hamming_distance"]
data = pandas.read_csv('Data/FGDQN_1.csv', names=colnames)
loss = [float(x) for x in data.Loss.tolist()[1:-1]]
iters = np.arange(1,len(loss)+1).tolist()
plt.figure()
plt.plot(iters,loss)
plt.xlabel("Iteration", fontsize=12)
plt.ylabel("Loss", fontsize=12)
plt.show()
Hamming_distance = [float(x) for x in data.Hamming_distance.tolist()[1:-1]]
iters = np.arange(1,len(Hamming_distance)+1).tolist()
plt.figure()
plt.plot(iters,Hamming_distance)
plt.xlabel("Iteration", fontsize=12)
plt.ylabel("Hamming_distance", fontsize=12)
plt.show()
print("Final Policy", data.State.tolist()[-1])
# +
import pandas
import csv
import matplotlib.pyplot as plt
import numpy as np
colnames = ["Loss","State","Policy","Hamming_distance"]
data = pandas.read_csv('Data/DQN_1.csv', names=colnames)
loss = [float(x) for x in data.Loss.tolist()[1:-1]]
iters = np.arange(1,len(loss)+1).tolist()
plt.figure()
plt.plot(iters,loss)
plt.xlabel("Iteration", fontsize=12)
plt.ylabel("Loss", fontsize=12)
plt.show()
Hamming_distance = [float(x) for x in data.Hamming_distance.tolist()[1:-1]]
iters = np.arange(1,len(Hamming_distance)+1).tolist()
plt.figure()
plt.plot(iters,Hamming_distance)
plt.xlabel("Iteration", fontsize=12)
plt.ylabel("Hamming_distance", fontsize=12)
plt.show()
print("Final Policy", data.State.tolist()[-1])
|
src/forest/main.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/sartansartan/DS-Unit-1-Sprint-3-Statistical-Tests-and-Experiments/blob/master/module1-statistics-probability-and-inference/ASartan_LS_DS_132_Sampling_Confidence_Intervals_and_Hypothesis_Testing_Assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="11OzdxWTM7UR" colab_type="text"
# ## Assignment - Build a confidence interval
#
# A confidence interval refers to a neighborhood around some point estimate, the size of which is determined by the desired p-value. For instance, we might say that 52% of Americans prefer tacos to burritos, with a 95% confidence interval of +/- 5%.
#
# 52% (0.52) is the point estimate, and +/- 5% (the interval $[0.47, 0.57]$) is the confidence interval. "95% confidence" means a p-value $\leq 1 - 0.95 = 0.05$.
#
# In this case, the confidence interval includes $0.5$ - which is the natural null hypothesis (that half of Americans prefer tacos and half burritos, thus there is no clear favorite). So in this case, we could use the confidence interval to report that we've failed to reject the null hypothesis.
#
# But providing the full analysis with a confidence interval, including a graphical representation of it, can be a helpful and powerful way to tell your story. Done well, it is also more intuitive to a layperson than simply saying "fail to reject the null hypothesis" - it shows that in fact the data does *not* give a single clear result (the point estimate) but a whole range of possibilities.
#
# How is a confidence interval built, and how should it be interpreted? It does *not* mean that 95% of the data lies in that interval - instead, the frequentist interpretation is "if we were to repeat this experiment 100 times, we would expect the average result to lie in this interval ~95 times."
#
# For a 95% confidence interval and a normal(-ish) distribution, you can simply remember that +/-2 standard deviations contains 95% of the probability mass, and so the 95% confidence interval based on a given sample is centered at the mean (point estimate) and has a range of +/- 2 (or technically 1.96) standard deviations.
#
# Different distributions/assumptions (90% confidence, 99% confidence) will require different math, but the overall process and interpretation (with a frequentist approach) will be the same.
#
# Your assignment - using the data from the prior module ([congressional voting records](https://archive.ics.uci.edu/ml/datasets/Congressional+Voting+Records)):
#
#
# ### Confidence Intervals:
# 1. Generate and numerically represent a confidence interval
# 2. Graphically (with a plot) represent the confidence interval
# 3. Interpret the confidence interval - what does it tell you about the data and its distribution?
#
# ### Chi-squared tests:
# 4. Take a dataset that we have used in the past in class that has **categorical** variables. Pick two of those categorical variables and run a chi-squared tests on that data
# - By hand using Numpy
# - In a single line using Scipy
#
# + id="Ckcr4A4FM7cs" colab_type="code" colab={}
import pandas as pd
import numpy as np
# + id="ixRIHefLcQB8" colab_type="code" colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY> "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": "OK"}}, "base_uri": "https://localhost:8080/", "height": 70} outputId="300dd677-5360-4aeb-b953-15aeb85d0c3e"
from google.colab import files
uploaded = files.upload()
# + id="qBBOG4B3caD9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 278} outputId="b9005d86-1588-4aef-cdd1-b33a5dc995a6"
df = pd.read_csv("house-votes-84.data", header=None)
df.columns = ['Class Name', 'handicapped-infants', 'water-project-cost-sharing',
'adoption-of-the-budget-resolution', 'physician-fee-freeze', 'el-salvador-aid',
'religious-groups-in-schools', 'anti-satellite-test-ban', 'aid-to-nicaraguan-contras',
'mx-missile', 'immigration', 'synfuels-corporation-cutback', 'education-spending',
'superfund-right-to-sue', 'crime', 'duty-free-exports', 'export-administration-act-south-africa']
df = df.replace({"n":0, "y":int('1'), '?': np.NaN})
df.head()
# + id="du2CxmpWdDqy" colab_type="code" colab={}
from scipy import stats
# + id="u84zW75ndEgk" colab_type="code" colab={}
def confidence_interval(data, confidence=0.95):
data = np.array(data)
mean = np.mean(data)
n = len(data)
stderr = stats.sem(data)
t = stats.t.ppf((1 + confidence) / 2.0, n - 1)
interval = stderr * t
print (mean, mean - interval, mean + interval)
return mean, interval
# + id="tO_lJBJ4GHlg" colab_type="code" colab={}
columns = ['handicapped-infants', 'water-project-cost-sharing',
'adoption-of-the-budget-resolution', 'physician-fee-freeze', 'el-salvador-aid',
'religious-groups-in-schools', 'anti-satellite-test-ban', 'aid-to-nicaraguan-contras',
'mx-missile', 'immigration', 'synfuels-corporation-cutback', 'education-spending',
'superfund-right-to-sue', 'crime', 'duty-free-exports', 'export-administration-act-south-africa']
# + id="LloAfjwGHstY" colab_type="code" colab={}
df2 = df.dropna()
# + id="gSUleXLFGPew" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 302} outputId="9ae5e7ec-2c31-4d20-ac5f-7364ce770e20"
mean_list = []
interval_list = []
for col in columns:
mean, interval = confidence_interval(df2[col])
mean_list.append(mean)
interval_list.append(interval)
# + id="-R0bhkF0HULt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 302} outputId="3206fb6d-b6c8-476a-cf21-b2f63f8d84a9"
mean_list
# + id="E3EOXvrAJJrF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 302} outputId="6f27e586-4a7e-47bf-ba50-e68de99bd7d7"
interval_list
# + id="sXNdP0MxKuYl" colab_type="code" colab={}
mean_minus = mean_list - interval
mean_plus = mean_list + interval
# + id="hpgBL3gSJ0Kh" colab_type="code" colab={}
d = {'issue': columns, 'mean': mean_list, 'mean_minus': mean_minus, 'mean_plus': mean_plus}
df_final = pd.DataFrame(data=d)
df_final.set_index('issue', inplace=True)
# + id="-cyTPExPKCaX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 237} outputId="6dc2a6a1-f490-4901-b01a-a16eb9b9c733"
df_final.head()
# + id="PLfoO4q3LgZn" colab_type="code" colab={}
data2 = df_final
# + id="0i4HLLfQhCCY" colab_type="code" colab={}
import matplotlib.pyplot as plt
# + id="MEobemo2LW5p" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 269} outputId="2c043538-3d98-4773-ac7e-a5e9ad0ab38b"
fig2, ax2 = plt.subplots()
ax2.boxplot(data2);
# + id="TuybbtcFkp18" colab_type="code" colab={}
#Interpret the confidence interval - what does it tell you about the data and its distribution?
# + id="xQC31t1bkrsj" colab_type="code" colab={}
#The poll is belived to be accurate within ((mean_plus - mean_mins)/2), 19 times out of 20
# + id="-9X9eQh567C5" colab_type="code" colab={}
df_rep = df[df['Class Name'] == 'republican']
# + id="cWK2glgpMtl3" colab_type="code" colab={}
df_rep2 = df_rep.dropna()
# + id="sddofoY_7AHK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 302} outputId="5e9e87e5-c0a6-437e-adc1-0c55ff9b948d"
mean_list_rep = []
interval_list_rep = []
for col in columns:
mean, interval = confidence_interval(df_rep2[col])
mean_list_rep.append(mean)
interval_list_rep.append(interval)
# + id="3oVHbw0n7JNl" colab_type="code" colab={}
mean_minus_rep = mean_list_rep - interval
mean_plus = mean_list_rep + interval
d = {'issue': columns, 'mean': mean_list, 'mean_minus': mean_minus, 'mean_plus': mean_plus}
df_final_rep = pd.DataFrame(data=d)
df_final_rep.set_index('issue', inplace=True)
# + id="GFgzcKyDRtMF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 582} outputId="15772e6c-41f7-414e-9645-2f740b358825"
df_final_rep
# + id="9BzIsKZC7n8D" colab_type="code" colab={}
data_rep = df_final_rep
# + id="VuvtlR-JThF_" colab_type="code" colab={}
import matplotlib.style as style
# + id="Itv9X6Ha7kFb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 469} outputId="4601f6d4-ed85-4b44-b885-30b2f87575db"
fig3, ax3 = plt.subplots()
ax3.boxplot(data_rep);
style.use('fivethirtyeight')
fig3.size = (12,30)
fte_graph = ax3.boxplot(data_rep)
fte_graph.tick_params(axis = 'both', which = 'major', labelsize = 18)
# + id="reKlyxJ77Rys" colab_type="code" colab={}
#With 95% confidence between 0.12 and 0.24 republicans will support 'handicapped-infants' cause
# + id="PXt80XL4sCKA" colab_type="code" colab={}
#Take a dataset that we have used in the past in class that has categorical variables.
#Pick two of those categorical variables and run a chi-squared tests on that data
# By hand using Numpy
# In a single line using Scipy
# + id="3XVAFE_xsFVm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 348} outputId="ba1a9d01-f4c7-4c9a-f386-58281396e13e"
df = pd.read_csv('https://raw.githubusercontent.com/ryanleeallred/datasets/master/adult.csv', na_values=" ?")
df.head()
# + id="nQzU4VLVsKW9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 332} outputId="93d495b5-af8d-49a5-be85-f25849830c2f"
df = df.sort_values(by='marital-status')
# https://github.com/pandas-dev/pandas/issues/25278
contingency_table = pd.crosstab(df['marital-status'], df['salary'], margins=True)
contingency_table
# + id="ppnf6R5MtVXu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 53} outputId="493f4ce9-1ee5-41f6-bcab-b17b7e46ed6d"
row_sums = contingency_table.iloc[0:7, 2].values
col_sums = contingency_table.iloc[7, 0:2].values
print(row_sums)
print(col_sums)
# + id="U2yOfuSvt2nX" colab_type="code" colab={}
total = contingency_table.loc['All','All']
# + id="d2_1e-3wtrSv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 159} outputId="ebd28c0d-ea60-4c94-9a3f-942e57082d1b"
expected = []
for row_sum in row_sums:
expected_row = []
for column in col_sums:
expected_val = column*row_sum/total
expected_row.append(expected_val)
expected.append(expected_row)
expected = np.array(expected)
print(expected.shape)
print(expected)
# + id="RuLbyGJ0t8yj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 159} outputId="a3220303-ae85-4cc9-cf06-e251f0196d92"
observed = pd.crosstab(df['marital-status'], df['salary']).values
print(observed.shape)
observed
# + id="5xL_x3NmuPZ0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="b52a365f-2c81-46c2-f8bb-3490a62b3ecf"
chi_square = ((observed - expected)**2/(expected)).sum()
chi_square
# + id="vAWnAF_YuU6J" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 141} outputId="f8e0c854-17be-4743-abd2-b8a29d54de71"
chi_squared, p_value, dof, expected = stats.chi2_contingency(observed)
print(chi_squared, p_value, dof, expected)
# + id="8jeuZtQguxve" colab_type="code" colab={}
#Null Hypothesis: martial status is independent of salary.
#Due to a p-value of 0, we REJECT the null hypothesis that martial status and salary are independent,
#and conclude that there is an association between martial status and salary.
# + [markdown] id="4ohsJhQUmEuS" colab_type="text"
# ## Stretch goals:
#
# 1. Write a summary of your findings, mixing prose and math/code/results. *Note* - yes, this is by definition a political topic. It is challenging but important to keep your writing voice *neutral* and stick to the facts of the data. Data science often involves considering controversial issues, so it's important to be sensitive about them (especially if you want to publish).
# 2. Apply the techniques you learned today to your project data or other data of your choice, and write/discuss your findings here.
# 3. Refactor your code so it is elegant, readable, and can be easily run for all issues.
# + [markdown] id="nyJ3ySr7R2k9" colab_type="text"
# ## Resources
#
# - [Interactive visualize the Chi-Squared test](https://homepage.divms.uiowa.edu/~mbognar/applets/chisq.html)
# - [Calculation of Chi-Squared test statistic](https://en.wikipedia.org/wiki/Pearson%27s_chi-squared_test)
# - [Visualization of a confidence interval generated by R code](https://commons.wikimedia.org/wiki/File:Confidence-interval.svg)
# - [Expected value of a squared standard normal](https://math.stackexchange.com/questions/264061/expected-value-calculation-for-squared-normal-distribution) (it's 1 - which is why the expected value of a Chi-Squared with $n$ degrees of freedom is $n$, as it's the sum of $n$ squared standard normals)
|
module1-statistics-probability-and-inference/ASartan_LS_DS_132_Sampling_Confidence_Intervals_and_Hypothesis_Testing_Assignment.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
# -
from osu_util import read_osu_log, read_osu_log_multi, plot_osu
from collective_algo import bcast_algo_intelmpi, bcast_algo_openmpi3, bcast_algo_openmpi4
# ls osu_log/bcast/N8n288/
TOP_DIR = './osu_log/bcast/N8n288/run3/'
def plot_barh(df):
fig, axes = plt.subplots(2, 2, figsize=[12, 6])
for ax, message in zip(axes.flatten(), [1024, 8192, 65536, 1048576]):
df.loc[message].plot.barh(grid=True, ax=ax)
ax.set_xlim(0, df.loc[message]['default'] * 5)
ax.set_title('message size = {}'.format(message))
ax.set_xlabel('time (us)')
ax.vlines(df.loc[message]['default'], -0.5, df.shape[1], linestyle='--')
fig.tight_layout()
# ## OpenMPI3
# +
ompi3_dir = TOP_DIR + 'openmpi3/'
ompi3_file_list = [ompi3_dir + 'bcast_default.log'] + [ompi3_dir + 'bcast_algo{}.log'.format(i) for i in range(0, 7)]
ompi3_file_list
# -
df_ompi3 = read_osu_log_multi(ompi3_file_list, ['default'] + bcast_algo_openmpi3)
df_ompi3[::4]
plot_osu(df_ompi3)
plt.ylim(0, 8000)
plot_barh(df_ompi3)
# # OpenMPI4
# +
ompi4_dir = TOP_DIR + 'openmpi4/'
ompi4_file_list = [ompi4_dir + 'bcast_default.log'] + [ompi4_dir + 'bcast_algo{}.log'.format(i) for i in range(0, 10)]
ompi4_file_list
# -
df_ompi4 = read_osu_log_multi(ompi4_file_list, ['default'] + bcast_algo_openmpi4)
df_ompi4[::4]
plt.rcParams['font.size'] = 12
plot_barh(df_ompi4)
# ## MPICH
mpich_dir = TOP_DIR + 'mpich3/'
df_mpich = read_osu_log_multi([mpich_dir + 'bcast_default.log'], columns=['default'])
df_mpich[::4]
# ## Intel MPI
# +
impi_dir = TOP_DIR + '/intelmpi-tcp/'
impi_file_list = [impi_dir + 'bcast_default.log'] + [impi_dir + 'bcast_algo{}.log'.format(i) for i in range(1, 15)]
df_impi = read_osu_log_multi(impi_file_list, ['default'] + bcast_algo_intelmpi)
df_impi[::4]
# +
plt.rcParams['font.size'] = 10
plot_barh(df_impi)
# +
fig, axes = plt.subplots(1, 3, figsize=[14, 3])
plot_osu(df_impi.iloc[:, 0:5], ax=axes[0])
plot_osu(df_impi.iloc[:, 5:10], ax=axes[1])
plot_osu(df_impi.iloc[:, 10:], ax=axes[2])
# -
# ## Intel MPI EFA
# +
impi_efa_dir = TOP_DIR + '/intelmpi-efa/'
impi_efa_file_list = [impi_efa_dir + 'bcast_default.log'] + [impi_efa_dir + 'bcast_algo{}.log'.format(i) for i in range(1, 15)]
df_impi_efa = read_osu_log_multi(impi_efa_file_list, ['default'] + bcast_algo_intelmpi)
df_impi_efa[::4]
# -
plot_barh(df_impi_efa)
# # Put all together
# +
df_all = pd.concat(
[df_ompi3.add_suffix(' | OpenMPI 3'),
df_ompi4.add_suffix(' | OpenMPI 4'),
df_mpich.add_suffix(' | MPICH 3'),
df_impi.add_suffix(' | IntelMPI-TCP'),
df_impi_efa.add_suffix(' | IntelMPI-EFA')
], axis=1)
df_select = df_all.loc[65536]
df_select = df_select[df_select < df_select['default | OpenMPI 3']*1.2] # ignore cases that are too slower than default
df_select.sort_values(ascending=False).plot.barh(figsize=[10, 20], grid=True)
# -
# ## Only select default cases
df_default = df_all.filter(like='default', axis=1)
plot_osu(df_default)
df_default.loc[[1024, 8192, 65536]].T.plot.barh(grid=True)
# ## Only select important cases
df_important = df_all.filter(regex=r'(default|knomial|Knomial|split)')
df_important.index.name='message size (Bytes)'
# +
plt.rcParams['font.size'] = 14
df_important.loc[[16384, 65536]].T.plot.barh(grid=True, figsize=[8, 8])
plt.xlabel('time (us)')
plt.title('MPI_Bcast with 288 ranks on 8 nodes (c5n.18xlarge)')
plt.savefig('bcast_summary.png', dpi=144, bbox_inches='tight')
# -
df_important.loc[[262144, 1048576]].T.plot.barh(grid=True, figsize=[8, 8])
plt.xlabel('time (us)')
|
notebooks/bcast_summary.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Structural Reliability - Basics and Example
# This is a very condensed introduction to a simple structural reliability problem.
#
# ## Context:
# Structural reliability is defined as the complement of structural failure. Structural failure is defined as an event, where the load bearing capacity $R$ is smaller than the load effect $S$. Correspondingly, the failure probability is the defined as $P_F=\Pr(R\le S)=\Pr(R-S\le 0)$ and reliability as $\Pr(R-S>0)$.
# In general, $R$ and $S$ are not certainly known and are mathematically represented as random variables. If it is assumed that $R$ and $S$ can be represented as independent normal distributed random variables, a safety margin $M=R-S$ can be introduced, which is as well normal distributed with mean value $\mu_M=\mu_R-\mu_S$ and standard deviation $\sigma_M=\sqrt{\sigma_R^2+\sigma_S^2}$. The failure probability can then computed as
#
# $P_F=\Pr(R-S<0)=\Pr(M<0)=\Phi\left(\frac{0-\mu_M}{\sigma_M}\right)=\Phi\left(-\frac{\mu_R-\mu_S}{\sqrt{\sigma_R^2+\sigma_S^2}}\right)=\Phi\left(-\beta\right)$
#
# where $\beta=\frac{\mu_R-\mu_S}{\sqrt{\sigma_R^2+\sigma_S^2}}$ is referred to as the *reliability index*.
#
# ## Example:
# You have to design a beam that has to span $l=10$ m and has to carry a load $Q$ . The material that is available is glued laminated timber (Glulam) and the cross-section is specified to be rectangular with a width of $b =300$ mm and height $h$. The load is given in this project to be a uniform distributed load that is represented by its 50 years maximum value $Q$. The material property of interest in this case is the bending strength of the Glulam $F_{m}$ . The situation is illustrated in the following Figure.
#
# 
#
# The dominating effect ($S$) of the load $Q$ is the bending moment at mid-span, which is $Ql^{2}/8$. The elastic bending load bearing capacity of the rectangular cross-section $R$ is $F_{m}bh^{2}/6$.
# Structural failure is defined as the event when the load on a structure is larger than its load bearing capacity. In other words, failure is characterized by the difference between the load bearing capacity and the load being negative. The corresponding limit state is general referred to as *Ultimate Limit State (ULS)* and for this example expressed as:
#
# $g\left(R,S\right) = R-S = \left(\dfrac{bh^{2} }{6} \right)F_{m} -\left(\dfrac{l^{2} }{8} \right)Q\le 0$
#
# The limit state equation contains different variables, some of them are uncertain or random, as $Q,F_m$, and represented as Normal distributed with mean value and coefficient of variation, $\mu_Q=24.1$ N/mm, $V_Q=0.3$, and, $\mu_{F_m}=24.1$ MPa, $V_{F_m}=0.15$. For a chosen $h=800$ mm, the reliability index and the corresponding failure probability are computed as:
#
#
# +
## Packages and general settings
import numpy as np
import scipy as sp
import matplotlib.pyplot as plt
import scipy.stats
fontsizes=18
plt.rcParams.update({'font.size': fontsizes})
plt.rcParams.update({"font.family": "serif"})
plt.rcParams.update({"mathtext.fontset" : "cm"})
plt.rcParams.update({'font.serif': 'Times New Roman'})
plt.close('all')
# +
## Input
# Geometry
l = 10000 # [mm] span
b = 300 # [mm] width
#==============================================================================
# Material properties
mu_fm = 26.6 # [MPa] mean material resistance
cov_fm = 0.15 # coeff. of variation
std_fm = mu_fm*cov_fm # [MPa] standard deviation
mu_R = mu_fm*b/6
std_R = std_fm*b/6
#==============================================================================
# Load
mu_q = 24.1 # [N/mm] mean load
cov_q = 0.3 # coeff. of variation
std_q = mu_q*cov_q # [MPa] standard deviation
mu_S = (l**2/8)*mu_q
std_S = (l**2/8)*std_q
h1 = 800 # in [mm]
## Computation
# Reliability index :
beta = (mu_R*h1**2-mu_S)/(((std_R*h1**2)**2+(std_S)**2)**0.5)
# Probability of failure :
PF = sp.stats.norm.cdf(-beta)
# Results:
B1 = "Reliability Index Beta: {b:.2f} \n".format(b=beta)
P2 = "Probability of failure: {pf:.2e} \n".format(pf=PF)
print(B1)
print(P2)
# -
# We might be interested how the choice of $h$ affects the reliability index and the failure probability, say in the range of $h = [500,1500]$.
# +
# Reliability index as a function of the decision variable
BETA = lambda h: (mu_R*h**2-mu_S)/(((std_R*h**2)**2+(std_S)**2)**0.5)
h1 = np.linspace(500,1500, num=10000)
beta = BETA(h1)
PF = sp.stats.norm.cdf(-beta)
# Plot settings
plt.figure()
plt.subplot(121)
plt.plot(h1,beta, color='black',lw=2)
plt.xlabel('$h$ [m]',fontsize=fontsizes)
plt.ylabel(r'$\beta$',fontsize=fontsizes)
plt.xlim(500,1500)
plt.ylim(0,6)
plt.subplot(122)
plt.plot(h1,PF, color='black',lw=2)
plt.yscale('log')
plt.xlabel('$h$ [m]',fontsize=fontsizes)
plt.ylabel('$P_f$',fontsize=fontsizes)
plt.xlim(500,1500)
plt.ylim(1e-10,1e0)
plt.tight_layout()
plt.show()
# -
# Let's see how the relation between $\beta$ and the decision parameter $h$ is influenced by the coefficient of variation of the load:
# +
from IPython.display import display, Markdown, clear_output
import json
import ipywidgets as widgets
import scipy as sp
slider_covq = widgets.FloatSlider(
value=0.3,
min=0,
max=1.0,
step=0.05,
description='CoV load:',
orientation='horizontal',
readout=True,
readout_format='2.2f',)
button = widgets.Button(description='Refresh plot')
out = widgets.Output()
# Plot settings
def plot_beta(ax,beta):
PF = sp.stats.norm.cdf(-beta)
ax[0].plot(h1,beta, color='black',lw=2)
ax[0].set_xlabel('$h$ [m]')
ax[0].set_ylabel(r'$\beta$')
ax[0].set_xlim(500,1500)
ax[0].set_ylim(0,6)
ax[1].plot(h1,PF, color='black',lw=2)
ax[1].set_yscale('log')
ax[1].set_xlabel('$h$ [m]',fontsize=fontsizes)
ax[1].set_ylabel('$P_f$',fontsize=fontsizes)
ax[1].set_xlim(500,1500)
ax[1].set_ylim(1e-10,1e0)
plt.tight_layout()
plt.show()
def on_button_clicked(b):
with out:
clear_output()
std_q = mu_q*slider_covq.value # [MPa] standard deviation
std_S = (l**2/8)*std_q
BETA = lambda h: (mu_R*h**2-mu_S)/(((std_R*h**2)**2+(std_S)**2)**0.5)
h1 = np.linspace(500,1500, num=10000)
beta = BETA(h1)
fig, ax1 = plt.subplots(1,2)
plot_beta(ax1,beta)
button.on_click(on_button_clicked)
display(widgets.VBox([slider_covq,button,out]))
# +
from IPython.display import display, Markdown, clear_output
from matplotlib.figure import figaspect
import json
import ipywidgets as widgets
import scipy as sp
slider_h = widgets.FloatSlider(
value=1000,
min=500,
max=1500,
step=0.05,
description='$h$=',
orientation='horizontal',
readout=True,
readout_format='2.2f',)
slider_covq = widgets.FloatSlider(
value=0.3,
min=0,
max=1.0,
step=0.05,
description='CoV load:',
orientation='horizontal',
readout=True,
readout_format='2.2f',)
button = widgets.Button(description='Refresh plot')
out = widgets.Output()
# Plot settings
def plot_margin(ax,h,std_S,Pf):
S = np.linspace(mu_S-5*std_S,mu_S+5*std_S,100)
R = np.linspace(mu_R*h**2-5*std_R*h**2,mu_R*h**2+5*std_R*h**2,100)
std_M = np.sqrt(std_S**2+(std_R*h**2)**2)
mu_M = mu_R*h**2-mu_S
M1 = np.linspace(min(mu_M-5*std_M,-std_M),0,20)
M2 = np.linspace(0.1,max(mu_M+5*std_M,std_M),80)
M = np.concatenate((M1,M2))
y_S = sp.stats.norm.pdf(S,mu_S,std_S)
y_S /= np.trapz(y_S,S)
y_R = sp.stats.norm.pdf(R,mu_R*h**2,std_R*h**2)
y_R /=np.trapz(y_R,R)
y_M = sp.stats.norm.pdf(M,mu_M,std_M)
cte = np.trapz(y_M,M)
y_M /= cte
ax.plot(S,y_S, color='red',label = '$S$')
ax.plot(R,y_R, color='blue',label = '$R$')
ax.plot(M,y_M, color='black',label = '$M$')
ax.fill_between(M1, 0, sp.stats.norm.pdf(M1,mu_M,std_M)/cte,color='red',alpha = 0.4)
max_y = max(max(y_M),max(y_R),max(y_S))
ax.plot(np.array([0,0]),np.array([0,max_y*1.1]),'--k')
ax.set_xlabel('$S$,$R$,$M$')
ax.set_ylabel('PDF')
ax.set_xlim(min(-std_M,mu_M-3*std_M),max(mu_R*h**2+3*std_R*h**2,mu_S+3*std_S))
ax.set_ylim(0,max_y*1.1)
ax.set_xticks(np.array([0,mu_S,mu_M,mu_R*h**2]))
ax.set_xticklabels(['0','$\mu_S$','$\mu_M$','$\mu_R$'])
ax.set_yticks([])
ax.text(0,0,'$P_f$ = {p:.2e}'.format(p=Pf),horizontalalignment='right',verticalalignment='top')
ax.legend(loc = 'upper left', bbox_to_anchor=(1, 1))
plt.show()
def on_button_clicked(b):
with out:
clear_output()
h = slider_h.value # [MPa] standard deviation
std_q = mu_q*slider_covq.value
std_S = (l**2/8)*std_q
beta = (mu_R*h**2-mu_S)/(((std_R*h**2)**2+(std_S)**2)**0.5)
Pf = sp.stats.norm.cdf(-beta)
wp, hp = figaspect(1/3)
fig, ax2 = plt.subplots(figsize=(wp,hp))
plot_margin(ax2,h,std_S,Pf)
button.on_click(on_button_clicked)
display(widgets.VBox([slider_h,slider_covq,button,out]))
# -
|
SimpleReliability.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="dNPR9GYHcgU0" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1627552861376, "user_tz": -480, "elapsed": 6924, "user": {"displayName": "\u4f55\u51a0\u7def", "photoUrl": "", "userId": "03629691439050504910"}} outputId="ac899ffc-95f4-4d37-975e-4bfbd5527ee5"
# !pip install transformers
# + id="zIcm4lLIuAtc" executionInfo={"status": "ok", "timestamp": 1627552868940, "user_tz": -480, "elapsed": 7567, "user": {"displayName": "\u4f55\u51a0\u7def", "photoUrl": "", "userId": "03629691439050504910"}}
from transformers import BertTokenizer, BertForSequenceClassification, AdamW, BertConfig, get_linear_schedule_with_warmup
import torch
import pandas as pd
import numpy as np
import time
import datetime
import random
import matplotlib.pyplot as plt
import plotly.express as px
from sklearn.model_selection import train_test_split
from keras.preprocessing.sequence import pad_sequences
from torch.utils.data import TensorDataset, DataLoader, RandomSampler, SequentialSampler
# + colab={"base_uri": "https://localhost:8080/"} id="5z_cA5TPhj38" executionInfo={"status": "ok", "timestamp": 1627552868940, "user_tz": -480, "elapsed": 11, "user": {"displayName": "\u4f55\u51a0\u7def", "photoUrl": "", "userId": "03629691439050504910"}} outputId="6d8cab33-d8b6-4ea8-e35e-2fc8f0459918"
# 檢查GPU環境
if torch.cuda.is_available():
print ("yes")
else:
print("no")
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
# + id="wUfpN-2soxMc" executionInfo={"status": "ok", "timestamp": 1627552868940, "user_tz": -480, "elapsed": 8, "user": {"displayName": "\u4f55\u51a0\u7def", "photoUrl": "", "userId": "03629691439050504910"}}
seed_val = 666
random.seed(seed_val)
np.random.seed(seed_val)
torch.manual_seed(seed_val)
torch.cuda.manual_seed_all(seed_val)
# + [markdown] id="oHny-9M-gtKl"
# # Load Dataset
# + id="TcKBIThkuCkr" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1627552868941, "user_tz": -480, "elapsed": 9, "user": {"displayName": "\u4f55\u51a0\u7def", "photoUrl": "", "userId": "03629691439050504910"}} outputId="7d8680e9-d364-47f4-8efa-5c43103f18d6"
df = pd.read_excel("/content/testing_data(labeled).xlsx")
print(df.head())
print(df.sample(10))
# + id="bsvckEWouGbj" executionInfo={"status": "ok", "timestamp": 1627552868941, "user_tz": -480, "elapsed": 7, "user": {"displayName": "\u4f55\u51a0\u7def", "photoUrl": "", "userId": "03629691439050504910"}}
# Get the lists of sentences and their labels.
sentences = df.sentences.values
labels = df.labels.values
# + colab={"base_uri": "https://localhost:8080/"} id="ARw1VnjBg0j7" executionInfo={"status": "ok", "timestamp": 1627552868942, "user_tz": -480, "elapsed": 7, "user": {"displayName": "\u4f55\u51a0\u7def", "photoUrl": "", "userId": "03629691439050504910"}} outputId="bb54ff51-cd19-4b34-e146-c55cf91ef0ba"
# check class distribution
df.labels.value_counts(normalize = True)
# + [markdown] id="t3a22FLig6ZX"
# # 將sentence轉成token
# + id="pa2zB3TYuILj" colab={"base_uri": "https://localhost:8080/", "height": 246, "referenced_widgets": ["84ff67503cd94cf3afd5213caa751354", "5457ae4c7a4548678692cac42edb9c8b", "f6f46837f27340679acbaf50c4877107", "<KEY>", "<KEY>", "70e06116bff542239ce941c61ca8cecb", "4b7eec8a212a40b3ab23bca9c1c54420", "533886e3fd464ed888e2448ead55b69d", "<KEY>", "<KEY>", "<KEY>", "5ca4cab9ef8f424e9b2f139fac0fa336", "d23d9d9ab7334a85b9c262c47de50f68", "<KEY>", "06373e13269e4b859bad6269ed1d7298", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "9936028479cc414092a2e4c854d174e8", "0e4194990cc64076b82e64be4a9dd7ac", "<KEY>", "<KEY>", "343b781e0e5342c1b44e24b7a80a21b2", "<KEY>", "db0a1a4c93e146d29acd2835df6970a6", "<KEY>", "32f35e8380f54b7b8e33d8ede1f0e244", "<KEY>", "0dc40711e5c943c38ed6c9fb9deb156d", "d491c5e4e6be4e8f93249d099c0796d5", "45a39bff6ac244f49a9725f059797ec9"]} executionInfo={"status": "ok", "timestamp": 1627552869809, "user_tz": -480, "elapsed": 872, "user": {"displayName": "\u4f55\u51a0\u7def", "photoUrl": "", "userId": "03629691439050504910"}} outputId="35a72bea-e1ed-49ba-ff02-33e67fa5d91e"
## 範例的:載入BERT tokenizer
print('==== Loading BERT tokenizer ====\n')
tokenizer = BertTokenizer.from_pretrained('bert-base-uncased')
# + id="FiPwg-P1uKNq" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1627552870258, "user_tz": -480, "elapsed": 451, "user": {"displayName": "\u4f55\u51a0\u7def", "photoUrl": "", "userId": "03629691439050504910"}} outputId="e5c0f83a-cc45-43f0-a791-7d6b64478f30"
input_ids = []
for sent in sentences:
encoded_sent = tokenizer.encode(
sent,
add_special_tokens = True, # Add '[CLS]' and '[SEP]'
)
input_ids.append(encoded_sent)
print('==== Tokenized Sentences ====\n')
print('Original: ', sentences[0])
print('Token IDs:', input_ids[0])
# + id="gghLMd9UuOjL" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1627552870258, "user_tz": -480, "elapsed": 7, "user": {"displayName": "\u4f55\u51a0\u7def", "photoUrl": "", "userId": "03629691439050504910"}} outputId="03b98079-037a-4986-bb50-e59cd61438f0"
max_seq_len = max([len(sen) for sen in input_ids])
print('Max sentence length: ', max_seq_len)
# + id="z7PHkVe5uQRK" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1627552870259, "user_tz": -480, "elapsed": 7, "user": {"displayName": "\u4f55\u51a0\u7def", "photoUrl": "", "userId": "03629691439050504910"}} outputId="456bfb23-d7c4-46cf-ac9e-708c3efad48b"
MAX_LEN = max_seq_len
print('\nPadding/truncating all sentences to %d values...' % MAX_LEN)
print('\nPadding token: "{:}", ID: {:}'.format(tokenizer.pad_token, tokenizer.pad_token_id))
# Pad our input tokens with value 0.
# "post" indicates that we want to pad and truncate at the end of the sequence,
# as opposed to the beginning.
input_ids = pad_sequences(input_ids, maxlen=MAX_LEN, dtype="long",
value=0, truncating="post", padding="post")
print('\Done.')
# + id="M8qQM9VOuS3z" executionInfo={"status": "ok", "timestamp": 1627552870259, "user_tz": -480, "elapsed": 5, "user": {"displayName": "\u4f55\u51a0\u7def", "photoUrl": "", "userId": "03629691439050504910"}}
# attention masks
attention_masks = []
for sent in input_ids:
att_mask = [int(token_id > 0) for token_id in sent]
attention_masks.append(att_mask)
# + [markdown] id="Nh-7cyq0hV9D"
# # Split train dataset into train, validation and test sets
# + id="76nE_V8nuVBs" executionInfo={"status": "ok", "timestamp": 1627552870259, "user_tz": -480, "elapsed": 4, "user": {"displayName": "\u4f55\u51a0\u7def", "photoUrl": "", "userId": "03629691439050504910"}}
## 準備訓練集、驗證集
train_inputs, validation_inputs, train_labels, validation_labels = train_test_split(
input_ids, labels, random_state = 2021,
test_size=0.2,
)
# 同樣masks
train_masks, validation_masks, _, _ = train_test_split(
attention_masks, labels, random_state=2021,
test_size=0.2,
)
# + id="JKl82VjGuWiE" executionInfo={"status": "ok", "timestamp": 1627552881935, "user_tz": -480, "elapsed": 11680, "user": {"displayName": "\u4f55\u51a0\u7def", "photoUrl": "", "userId": "03629691439050504910"}}
# 轉換成torch tensors
train_inputs = torch.tensor(train_inputs).to(device)
validation_inputs = torch.tensor(validation_inputs).to(device)
train_labels = torch.tensor(train_labels).to(device)
validation_labels = torch.tensor(validation_labels).to(device)
train_masks = torch.tensor(train_masks).to(device)
validation_masks = torch.tensor(validation_masks).to(device)
# + id="w1Ja993kuYAM" executionInfo={"status": "ok", "timestamp": 1627552881936, "user_tz": -480, "elapsed": 4, "user": {"displayName": "\u4f55\u51a0\u7def", "photoUrl": "", "userId": "03629691439050504910"}}
# The DataLoader
batch_size = 16
# 建立訓練集的DataLoader
train_data = TensorDataset(train_inputs, train_masks, train_labels)
train_sampler = RandomSampler(train_data)
train_dataloader = DataLoader(train_data, sampler=train_sampler, batch_size=batch_size)
# 建立驗證集的DataLoader
validation_data = TensorDataset(validation_inputs, validation_masks, validation_labels)
validation_sampler = SequentialSampler(validation_data)
validation_dataloader = DataLoader(validation_data, sampler=validation_sampler, batch_size=batch_size)
# + [markdown] id="8KxmrAbyhwYW"
# # Import BERT-base pretrained model
# + id="1etS1v_muZxk" colab={"base_uri": "https://localhost:8080/", "height": 170, "referenced_widgets": ["d5591eb0f9d049acaf2c28a1cacb7733", "188162bb2a0c41cab78616cdddc1e5cc", "d651e43422094322b05f54f08e765178", "c8c233ec541a48fcb877d5a415616ae1", "c26af8bf369d458385dd897f730e6302", "65223d1bd6c14f4c940b05b7130af5c1", "ce7bc35e3bba467ca1318ae36940fa10", "3e96d744dc4c41bfbb8be689992a40fb"]} executionInfo={"status": "ok", "timestamp": 1627552893742, "user_tz": -480, "elapsed": 11809, "user": {"displayName": "\u4f55\u51a0\u7def", "photoUrl": "", "userId": "03629691439050504910"}} outputId="9423a79a-2be1-4629-8d8c-d78e256f70f4"
## Load BertForSequenceClassification, the pretrained BERT model with a single
model = BertForSequenceClassification.from_pretrained(
'bert-base-uncased',
num_labels = 3,
output_attentions = False,
output_hidden_states = False,
)
# + id="buFP--O_ubi0" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1627552893742, "user_tz": -480, "elapsed": 14, "user": {"displayName": "\u4f55\u51a0\u7def", "photoUrl": "", "userId": "03629691439050504910"}} outputId="284abe82-2c3f-4b06-a3a4-3da11cd10d03"
# Get all of the model's parameters as a list of tuples.
params = list(model.named_parameters())
print('The BERT model has {:} different named parameters.\n'.format(len(params)))
print('==== Embedding Layer ====\n')
for p in params[0:5]:
print("{:<55} {:>12}".format(p[0], str(tuple(p[1].size()))))
print('\n==== First Transformer ====\n')
for p in params[5:21]:
print("{:<55} {:>12}".format(p[0], str(tuple(p[1].size()))))
print('\n==== Output Layer ====\n')
for p in params[-4:]:
print("{:<55} {:>12}".format(p[0], str(tuple(p[1].size()))))
# + colab={"base_uri": "https://localhost:8080/"} id="LKiSbwJxh46N" executionInfo={"status": "ok", "timestamp": 1627552894155, "user_tz": -480, "elapsed": 423, "user": {"displayName": "\u4f55\u51a0\u7def", "photoUrl": "", "userId": "03629691439050504910"}} outputId="6c7e1e6d-e45a-4b7d-f8ed-72c5ba6afc1f"
# to cuda
model.to(device)
# + id="s1qIj2ALudtU" executionInfo={"status": "ok", "timestamp": 1627552894155, "user_tz": -480, "elapsed": 3, "user": {"displayName": "\u4f55\u51a0\u7def", "photoUrl": "", "userId": "03629691439050504910"}}
optimizer = AdamW(model.parameters(),
lr = 2e-5, # args.learning_rate - default is 5e-5, our notebook had 2e-5
eps = 1e-8 # args.adam_epsilon - default is 1e-8.
)
epochs = 4 # authors recommend between 2 and 4
total_steps = len(train_dataloader) * epochs
# Create the learning rate scheduler.
scheduler = get_linear_schedule_with_warmup(
optimizer,
num_warmup_steps = 0, # Default value
num_training_steps = total_steps
)
# + id="aHEtxGnVufVc" executionInfo={"status": "ok", "timestamp": 1627552894156, "user_tz": -480, "elapsed": 4, "user": {"displayName": "\u4f55\u51a0\u7def", "photoUrl": "", "userId": "03629691439050504910"}}
def flat_accuracy(preds, labels):
pred_flat = np.argmax(preds, axis=1).flatten()
labels_flat = labels.flatten()
return np.sum(pred_flat == labels_flat) / len(labels_flat)
# + id="Qps3l1q2ug08" executionInfo={"status": "ok", "timestamp": 1627552894156, "user_tz": -480, "elapsed": 4, "user": {"displayName": "\u4f55\u51a0\u7def", "photoUrl": "", "userId": "03629691439050504910"}}
def format_time(elapsed):
'''
Takes a time in seconds and returns a string hh:mm:ss
'''
# Round to the nearest second.
elapsed_rounded = int(round((elapsed)))
# Format as hh:mm:ss
return str(datetime.timedelta(seconds=elapsed_rounded))
# + id="3pVrAyNuuiN0" executionInfo={"status": "ok", "timestamp": 1627552894156, "user_tz": -480, "elapsed": 3, "user": {"displayName": "\u4f55\u51a0\u7def", "photoUrl": "", "userId": "03629691439050504910"}}
loss_values = []
val_loss_value = []
# + [markdown] id="cXOqXYcguj4U"
# # Train Process
# + id="xXva0XnEukVM" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1627552896620, "user_tz": -480, "elapsed": 2467, "user": {"displayName": "\u4f55\u51a0\u7def", "photoUrl": "", "userId": "03629691439050504910"}} outputId="f4e7b3bc-e200-4a94-91da-5cef906b6338"
# 統計整個訓練時長
total_t0 = time.time()
# For each Training epoch
for epoch_i in range(0, epochs):
# ========================================
# Training
# ========================================
print('======== Epoch {:} / {:} ========'.format(epoch_i + 1, epochs))
print('======== Training ========')
t0 = time.time()# Measure how long the training epoch takes.
total_loss = 0 # Reset the total loss for this epoch.
model.train()
for step, batch in enumerate(train_dataloader):
# 每經過N次迭代,就輸出進度資訊
if step % 5 == 0 and not step == 0:
elapsed = format_time(time.time() - t0)
print(' Batch {:>5,} of {:>5,}. Elapsed: {:}.'.format(step, len(train_dataloader), elapsed))
b_input_ids = batch[0].to(device)
b_input_mask = batch[1].to(device)
b_labels = batch[2].to(device)
model.zero_grad()
outputs = model(b_input_ids,
token_type_ids=None,
attention_mask=b_input_mask,
labels=b_labels)
loss = outputs[0]
total_loss += loss.item()
loss.backward()
torch.nn.utils.clip_grad_norm_(model.parameters(), 1.0)
optimizer.step()
scheduler.step()
avg_train_loss = total_loss / len(train_dataloader)
loss_values.append(avg_train_loss)
#print("")
print(" Average training loss: {0:.2f}".format(avg_train_loss))
print(" Training epcoh took: {:}".format(format_time(time.time() - t0)))
# ========================================
# Validation
# ========================================
print("")
print("======== Running Validation ========")
t0 = time.time()
model.eval()
eval_loss, eval_accuracy = 0, 0
nb_eval_steps, nb_eval_examples = 0, 0
for batch in validation_dataloader:
batch = tuple(t.to(device) for t in batch)
b_input_ids, b_input_mask, b_labels = batch
with torch.no_grad():
outputs = model(b_input_ids,
token_type_ids = None,
attention_mask = b_input_mask)
logits = outputs[0]
# 累加 loss
eval_loss += loss.item()
logits = logits.detach().cpu().numpy()
label_ids = b_labels.to('cpu').numpy()
tmp_eval_accuracy = flat_accuracy(logits, label_ids)
eval_accuracy += tmp_eval_accuracy
nb_eval_steps += 1
avg_val_loss = eval_loss / nb_eval_steps # len(validation_dataloader)
avg_val_accuracy = eval_accuracy/nb_eval_steps
validation_time = format_time(time.time() - t0)
val_loss_value.append(avg_val_loss)
#print("")
print(" Accuracy: {0:.2f}".format(avg_val_accuracy))
print(" Loss: {0:.2f}".format(avg_val_loss))
print(" Validation took: {:}".format(validation_time))
print("")
print("Training complete!")
print("Total training took {:} (h:mm:ss)".format(format_time(time.time()-total_t0)))
# + id="kp82-8zZupYd" colab={"base_uri": "https://localhost:8080/", "height": 542} executionInfo={"status": "ok", "timestamp": 1627552897949, "user_tz": -480, "elapsed": 1331, "user": {"displayName": "\u4f55\u51a0\u7def", "photoUrl": "", "userId": "03629691439050504910"}} outputId="87d7a134-11d8-4b7d-8486-7ba18e37d746"
f = pd.DataFrame(loss_values)
f.columns=['Loss']
fig = px.line(f, x=f.index, y=f.Loss)
fig.update_layout(title='Training loss of the Model',
xaxis_title='Epoch',
yaxis_title='Loss')
fig.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="8eSRmKCQmEgy" executionInfo={"status": "ok", "timestamp": 1627552897950, "user_tz": -480, "elapsed": 5, "user": {"displayName": "\u4f55\u51a0\u7def", "photoUrl": "", "userId": "03629691439050504910"}} outputId="8e11eabb-a324-47ab-dd5a-d35e90c0996a"
def show_train_history(train, validation):
plt.plot(train)
plt.plot(validation)
plt.title('Training History')
plt.ylabel("Loss")
plt.xlabel("Epoch")
plt.legend(['train','validation'],loc='upper right')
plt.show()
show_train_history(loss_values, val_loss_value)
# + id="aHneKXrpjuIu" executionInfo={"status": "ok", "timestamp": 1627552899314, "user_tz": -480, "elapsed": 1367, "user": {"displayName": "\u4f55\u51a0\u7def", "photoUrl": "", "userId": "03629691439050504910"}}
torch.save(model, 'model.pth')
# + id="EcP0hG3yjxS8" executionInfo={"status": "ok", "timestamp": 1627552900995, "user_tz": -480, "elapsed": 1683, "user": {"displayName": "\u4f55\u51a0\u7def", "photoUrl": "", "userId": "03629691439050504910"}}
# 保存訓練好的權重
torch.save(model.state_dict(), 'model_state_dict.pt')
|
Question_Scoring/BERTforSequenceClassification.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:genpen]
# language: python
# name: conda-env-genpen-py
# ---
# + heading_collapsed="false" tags=[]
import itertools
import numpy as np
import os
import seaborn as sns
from tqdm import tqdm
from dataclasses import asdict, dataclass, field
import vsketch
import shapely.geometry as sg
from shapely.geometry import box, MultiLineString, Point, MultiPoint, Polygon, MultiPolygon, LineString
import shapely.affinity as sa
import shapely.ops as so
import matplotlib.pyplot as plt
import pandas as pd
import vpype_cli
from typing import List, Generic
from genpen import genpen as gp, utils as utils
from scipy import stats as ss
import geopandas
from shapely.errors import TopologicalError
import functools
# %load_ext autoreload
# %autoreload 2
# + heading_collapsed="false"
paper_size = '19x24 inches'
border:float=30
paper = utils.Paper(paper_size)
drawbox = paper.get_drawbox(border)
buffer_style = 2
# + heading_collapsed="false"
poly = drawbox.buffer(-20)
pg = gp.PerlinGrid(poly, xstep=6, ystep=6, lod=4, falloff=None, noise_scale=0.0063, noiseSeed=5)
# + heading_collapsed="false"
f,ax = plt.subplots(figsize=(6,6))
ax.quiver(np.cos(pg.a), np.sin(pg.a), scale=50)
# + heading_collapsed="false"
(xcs, ycs), _ = gp.overlay_grid(pg.p, xstep=60, ystep=60)
particles = []
for x,y in itertools.product(xcs,ycs):
pos = (x+np.random.randn()*1.6, y+np.random.randn()*1.6)
p = gp.Particle(pos=pos,
grid=pg, stepsize=1)
if pg.p.contains(p.pos):
particles.append(p)
# + heading_collapsed="false"
for p in tqdm(particles):
for i in range(np.random.randint(1,290)):
p.step()
lss = [LineString(p.pts) for p in particles if len(p.pts) > 1]
# -
angle_gen = ss.uniform(loc=0, scale=100).rvs
lbs = [l.buffer(ss.uniform(loc=2, scale=12).rvs(), cap_style=1, join_style=1) for l in lss]
lbs = [sa.rotate(l, angle=angle_gen()) for l in lbs]
polymerge = gp.merge_Polygons(lbs).buffer(0.1, cap_style=2, join_style=2)
# + heading_collapsed="false"
stp = gp.ScaleTransPrms(d_buffer=-0.8,angles=45,d_translate_factor=0.7)
stp.d_buffers += np.random.uniform(-0.04, 0.04, size=stp.d_buffers.shape)
# + heading_collapsed="false"
fills = []
for p in polymerge:
P = gp.Poly(p)
P.fill_scale_trans(**stp.prms)
fills.append(P.fill)
# + heading_collapsed="false"
layer1 = gp.merge_LineStrings(fills)
# -
layer1
# %%time
bd = 0.001
l1b = layer1.buffer(bd, cap_style=1, join_style=1, resolution=16).buffer(-bd, cap_style=1, join_style=1).boundary
l1b = [l for l in l1b if l.length > 0.1]
l1b = gp.merge_LineStrings(l1b)
all_polys = polymerge
# + heading_collapsed="false"
(xcs, ycs), _ = gp.overlay_grid(pg.p, xstep=20, ystep=30)
particles = []
for x,y in itertools.product(xcs,ycs):
pos = (x+np.random.randn()*1.6, y+np.random.randn()*1.6)
p = gp.Particle(pos=pos,
grid=pg, stepsize=1)
if pg.p.contains(p.pos):
particles.append(p)
# + heading_collapsed="false"
for p in tqdm(particles):
for i in range(np.random.randint(1,190)):
p.step()
lss = [LineString(p.pts) for p in particles if len(p.pts) > 1]
# -
angle_gen = ss.uniform(loc=0, scale=130).rvs
lbs = [l.buffer(ss.uniform(loc=2, scale=19).rvs(), cap_style=1, join_style=1) for l in lss]
lbs = [sa.rotate(l, angle=angle_gen()) for l in lbs]
polymerge = gp.merge_Polygons(lbs).buffer(0.1, cap_style=2, join_style=2)
polymerge = gp.robust_difference(polymerge, all_polys)
# + heading_collapsed="false"
stp = gp.ScaleTransPrms(d_buffer=-0.8,angles=-45,d_translate_factor=0.7)
stp.d_buffers += np.random.uniform(-0.09, 0.09, size=stp.d_buffers.shape)
# + heading_collapsed="false"
fills = []
for p in polymerge:
P = gp.Poly(p)
P.fill_scale_trans(**stp.prms)
fills.append(P.fill)
# + heading_collapsed="false"
layer2 = gp.merge_LineStrings(fills)
# -
layer2
bd = 0.00001
l2b = layer2.buffer(bd, cap_style=1, join_style=1, resolution=8).buffer(-bd, cap_style=1, join_style=1).boundary
l2b
sk = vsketch.Vsketch()
sk.size(paper.page_format_mm)
sk.scale('1mm')
sk.penWidth('0.8mm')
sk.stroke(1)
sk.geometry(l1b)
# sk.stroke(2)
# sk.geometry(l2b)
sk.display(color_mode='none', mode='ipython')
# +
savepath = '/Users/naka/code/side/plotter_images/oned_outputs/0310_morse_flow_beams.svg'
sk.save(savepath)
# -
from tqdm import tqdm
for tolerance in [0.2, 0.4, 0.8, 1, 2, 4, 8, 16, 32,]:
sk.vpype(f' splitall linemerge --tolerance {tolerance}mm')
sk.vpype('linesort')
sk.display(color_mode='none', mode='ipython')
# +
savepath = '/Users/naka/code/side/plotter_images/oned_outputs/0310_morse_flow_beams_merged_sorted3.svg'
sk.save(savepath)
# + heading_collapsed="false"
sk vpype vsketch.Vsketch()
sk.size(paper.page_format_mm)
sk.scale('1mm')
sk.penWidth('0.3mm')
sk.stroke(1)
sk.geometry(layer1)
sk.stroke(2)
sk.geometry(layer2)
sk.penWidth('0.3')
# sk.vpype('splitall')
# for tolerance in [0.2,]:
# sk.vpype(f'linemerge --tolerance {tolerance}mm linesimplify --tolerance 0.1')
# sk.vpype('linesimplify --tolerance 0.1 linesort')
sk.display(color_mode='none', mode='ipython')
# -
lss = l1b
lbs = [l.buffer(ss.uniform(loc=0.5, scale=6).rvs(), cap_style=2, join_style=2) for l in lss]
polymerge = gp.merge_Polygons(lbs).buffer(0.1, cap_style=2, join_style=2)
# + heading_collapsed="false"
stp = gp.ScaleTransPrms(d_buffer=-0.9,angles=45,d_translate_factor=0.7)
stp.d_buffers += np.random.uniform(-0.04, 0.04, size=stp.d_buffers.shape)
# + heading_collapsed="false"
fills = []
for p in polymerge:
P = gp.Poly(p)
P.fill_scale_trans(**stp.prms)
fills.append(P.fill)
# -
layer2 = gp.merge_LineStrings(fills)
# + heading_collapsed="false"
sk vpype vsketch.Vsketch()
sk.size(paper.page_format_mm)
sk.scale('1mm')
sk.penWidth('0.3mm')
sk.stroke(1)
sk.geometry(layer1)
sk.stroke(2)
sk.geometry(layer2)
sk.penWidth('0.3')
# sk.vpype('splitall')
# for tolerance in [0.2,]:
# sk.vpype(f'linemerge --tolerance {tolerance}mm linesimplify --tolerance 0.1')
# sk.vpype('linesimplify --tolerance 0.1 linesort')
sk.display(color_mode='none', mode='ipython')
# + heading_collapsed="false"
sk vpype vsketch.Vsketch()
sk.size(paper.page_format_mm)
sk.scale('1mm')
sk.penWidth('0.3mm')
sk.stroke(1)
sk.geometry(layer1)
sk.stroke(2)
sk.geometry(layer2)
sk.penWidth('0.3')
# sk.vpype('splitall')
# for tolerance in [0.2,]:
# sk.vpype(f'linemerge --tolerance {tolerance}mm linesimplify --tolerance 0.1')
# sk.vpype('linesimplify --tolerance 0.1 linesort')
sk.display(color_mode='none', mode='ipython')
# -
# ## Try2
# + heading_collapsed="false"
paper_size = '19x24 inches'
border:float=30
paper = utils.Paper(paper_size)
drawbox = paper.get_drawbox(border)
buffer_style = 2
# + heading_collapsed="false"
poly = drawbox.buffer(-20)
pg = gp.PerlinGrid(poly, xstep=6, ystep=6, lod=4, falloff=None, noise_scale=0.0063, noiseSeed=7)
# + heading_collapsed="false"
(xcs, ycs), _ = gp.overlay_grid(pg.p, xstep=40, ystep=40)
particles = []
for x,y in itertools.product(xcs,ycs):
pos = (x+np.random.randn()*8.6, y+np.random.randn()*8.6)
p = gp.Particle(pos=pos,
grid=pg, stepsize=1)
if pg.p.contains(p.pos):
particles.append(p)
# + heading_collapsed="false"
for p in tqdm(particles):
for i in range(np.random.randint(1,90)):
p.step()
lss = [LineString(p.pts) for p in particles if len(p.pts) > 1]
# -
angle_gen = ss.uniform(loc=0, scale=70).rvs
lbs = [l.buffer(ss.uniform(loc=2, scale=9).rvs(), cap_style=1, join_style=1) for l in lss]
lbs = [sa.rotate(l, angle=angle_gen()) for l in lbs]
polymerge = gp.merge_Polygons(lbs).buffer(0.1, cap_style=2, join_style=2)
# + heading_collapsed="false"
stp = gp.ScaleTransPrms(d_buffer=-0.9,angles=45,d_translate_factor=0.7)
stp.d_buffers += np.random.uniform(-0.04, 0.04, size=stp.d_buffers.shape)
# + heading_collapsed="false"
fills = []
for p in polymerge:
P = gp.Poly(p)
P.fill_scale_trans(**stp.prms)
fills.append(P.fill)
# -
bd = 0.0001
l1b = layer1.buffer(bd, cap_style=1, join_style=1, resolution=8).buffer(-bd, cap_style=1, join_style=1).boundary
all_polys = polymerge
# + heading_collapsed="false"
(xcs, ycs), _ = gp.overlay_grid(pg.p, xstep=20, ystep=30)
particles = []
for x,y in itertools.product(xcs,ycs):
pos = (x+np.random.randn()*1.6, y+np.random.randn()*1.6)
p = gp.Particle(pos=pos,
grid=pg, stepsize=1)
if pg.p.contains(p.pos):
particles.append(p)
# + heading_collapsed="false"
for p in tqdm(particles):
for i in range(np.random.randint(1,190)):
p.step()
lss = [LineString(p.pts) for p in particles if len(p.pts) > 1]
# -
angle_gen = ss.uniform(loc=0, scale=130).rvs
lbs = [l.buffer(ss.uniform(loc=2, scale=19).rvs(), cap_style=1, join_style=1) for l in lss]
lbs = [sa.rotate(l, angle=angle_gen()) for l in lbs]
polymerge = gp.merge_Polygons(lbs).buffer(0.1, cap_style=2, join_style=2)
polymerge = gp.robust_difference(polymerge, all_polys)
# + heading_collapsed="false"
stp = gp.ScaleTransPrms(d_buffer=-0.8,angles=-45,d_translate_factor=0.7)
stp.d_buffers += np.random.uniform(-0.09, 0.09, size=stp.d_buffers.shape)
# + heading_collapsed="false"
fills = []
for p in polymerge:
P = gp.Poly(p)
P.fill_scale_trans(**stp.prms)
fills.append(P.fill)
# + heading_collapsed="false"
layer2 = gp.merge_LineStrings(fills)
# -
layer2
bd = 0.00001
l2b = layer2.buffer(bd, cap_style=1, join_style=1, resolution=8).buffer(-bd, cap_style=1, join_style=1).boundary
l2b
_l1b = gp.merge_LineStrings([l for l in l1b if l.length > 0.2])
sns.displot([np.log10(l.length) for l in _l1b])
from tqdm.autonotebook import tqdm
# +
sk = vsketch.Vsketch()
sk.size(paper.page_format_mm)
sk.scale('1mm')
sk.penWidth('0.3mm')
sk.stroke(1)
sk.geometry(_l1b)
for tolerance in [0.2, 0.4, 0.8, 1, 2, 4, 8, 16, 32, 64]:
sk.vpype(f'linemerge --tolerance {tolerance}mm')
sk.stroke(2)
sk.geometry(_l1b)
sk.vpype('linesort')
sk.display(color_mode='layer', mode='ipython')
# +
savepath = '/Users/naka/code/side/plotter_images/oned_outputs/0307_morse_flow_beams.svg'
sk.save(savepath)
# -
lbs = [l.buffer(ss.uniform(loc=1, scale=8).rvs(), cap_style=1, join_style=1) for l in _l1b]
polymerge = gp.merge_Polygons(lbs).buffer(0.1, cap_style=2, join_style=2)
polymerge = gp.robust_difference(polymerge, all_polys)
# + heading_collapsed="false"
stp = gp.ScaleTransPrms(d_buffer=-0.8,angles=-45,d_translate_factor=0.7)
stp.d_buffers += np.random.uniform(-0.09, 0.09, size=stp.d_buffers.shape)
# + heading_collapsed="false"
fills = []
for p in polymerge:
P = gp.Poly(p)
P.fill_scale_trans(**stp.prms)
fills.append(P.fill)
# + heading_collapsed="false"
layer2 = gp.merge_LineStrings(fills)
# -
# # try 4
# + heading_collapsed="false"
paper_size = '11x14 inches'
border:float=20
paper = utils.Paper(paper_size)
drawbox = paper.get_drawbox(border)
buffer_style = 2
# + heading_collapsed="false"
poly = drawbox.buffer(-20)
pg = gp.PerlinGrid(poly, xstep=6, ystep=6, lod=4, falloff=None, noise_scale=0.0063, noiseSeed=5)
# -
p = drawbox.centroid.buffer(70)
stp = gp.ScaleTransPrms(d_buffer=-0.3,angles=-45,d_translate_factor=0.7, n_iters=1000)
stp.d_buffers += np.random.uniform(-0.09, 0.09, size=stp.d_buffers.shape)
P = gp.Poly(p)
P.fill_scale_trans(**stp.prms)
fill = P.fill
fill
bd = 0.01
ml = fill.buffer(bd, cap_style=1, join_style=1, resolution=13).buffer(-bd, cap_style=1, join_style=1).boundary
ml = gp.merge_LineStrings([l for l in ml if l.length>0.2])
ml.buff
# +
sk = vsketch.Vsketch()
sk.size(paper.page_format_mm)
sk.scale('1mm')
sk.penWidth('0.3mm')
sk.stroke(1)
sk.geometry(ml)
for tolerance in [0.1, 0.2, 0.4, 0.8,]:
sk.vpype(f'linemerge --tolerance {tolerance}mm')
# sk.stroke(2)
# sk.geometry(_l1b)
sk.vpype('linesort')
sk.display(color_mode='layer', mode='ipython')
# +
savepath = '/Users/naka/code/side/plotter_images/oned_outputs/0311_morse_flow_circle.svg'
sk.save(savepath)
# -
|
scratch/033_flow_beams.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Mini Project: Sorting and Evaluating Math Expressions
# ## Week 3
# **Q1.** *Mergesort:* Modify your `mergesort(array)` function that you did in your cohort session to take one additional argument called `byfunc`, i.e. `mergesort(array, byfunc)`. If the caller does not specify the value of `byfunc`, its default value is `None`. When this argument is `None`, the function `mergesort` behaves similar to your cohort session by sorting the array according to its values. However, when the value of this argument is not `None` but rather some other function, your `mergesort` function should sort the array according to the value returned by this function.
#
# For example, instead of sorting an array of integers, we want to sort an array of tupple.
# ```python
# array = [(1, 2), (3, 2), (2, -1), (4, 7), (-1, -2)]
# ```
# We can define a function say `select()` as follows:
# ```python
# def select(item):
# return item[0]
# ```
#
# You can then should be able to call your `mergesort()` function in the following:
# ```python
# mergesort(array, select)
# ```
# which will sort the list of tuples according to the value of its *first* element (recall `item[0]` in `select()`). This means that if you want to sort based on the *second* element of the tuple, you can redefine select as:
# ```python
# def select(item):
# return item[1]
# ```
#
# You can also apply this to a list of objects, say `User` class objects.
# ```python
# array = [<User 1>, <User 2>, <User 3>, ..., <User 101>]
# ```
# You can define the following `select()` function to sort according to its `username` attribute.
# ```python
# def select(item):
# return item.username
# ```
#
# You can then call the `mergesort()` function as follows:
# ```python
# mergesort(array, select)
# ```
#
# Python allows you to write [lambda functions](https://realpython.com/python-lambda/) to replace your `select()` function definition. You can simply call merge sort with the following without defining `select()`.
# ```python
# mergesort(array, lambda item: item.username)
# ```
def mergesort(array, byfunc=None):
pass
array = [(1, 2), (3, 2), (2, -1), (4, 7), (-1, -2)]
mergesort(array, lambda item: item[0])
assert array == [(-1, -2), (1, 2), (2, -1), (3, 2), (4, 7)]
mergesort(array, lambda item: item[1])
assert array == [(-1, -2), (2, -1), (1, 2), (3, 2), (4, 7)]
# **Q2.** Create a class called `EvaluateExpression` to evaluate mathematical expressions for Integers. The class has the following property:
# - `expression`: which is a property with a get and set method. The set method of this property should check if the string contains any invalid characters. If there is any invalid character, it should set the internal property `expr` to an empty String. Otherwise, it should set the string as it is. Valid characters are: `0123456789+-*/()` and an empty space.
# - `expr`: which is a property that stores only valid expression. It is used internally to store the expression.
#
# During object instantiation, a string can be passed on to `__init__()`.
# - `__init__(expr)`: where expr is the mathematical expression to initialize the property `expr`. If nothing is provided it should initialize to an empty String. If the string contains other characters besides those in the valid characters list above, the property `expr` should be initialized to an empty string.
#
#
#
# +
class EvaluateExpression:
valid_char = '0123456789+-*/() '
def __init__(self, string=""):
pass
@property
def expression(self):
pass
@expression.setter
def expression(self, new_expr):
pass
# -
expr1 = EvaluateExpression()
assert expr1.expression == ""
expr2 = EvaluateExpression("1 + 2")
assert expr2.expression == "1 + 2"
expr2.expression = "3 * 4"
assert expr2.expression == "3 * 4"
expr2.expression = "3 & 4"
assert expr2.expression == ""
# **Q3.** The class `EvaluateExpression` also has the following method:
# - `insert_space()`: which is used to insert one empty space before an operator and another empty space after the operator in the `expression` property. The function should return a new String. Note that this means that if there are two operators side by side, there will be two empty space between them.
#
#
class EvaluateExpression:
# copy the other definitions
# from the previous parts
def insert_space(self):
pass
expr1 = EvaluateExpression("(1+2)")
assert expr1.insert_space() == " ( 1 + 2 ) "
expr1.expression = "((1+2)*3/(4-5))"
assert expr1.insert_space() == " ( ( 1 + 2 ) * 3 / ( 4 - 5 ) ) "
# ## Week 4
# **Q4.** The class `EvaluateExpression` also has the following methods:
# - `process_operator(operand_stack, operator_stack)`: which process one operator. This method should modify the Stacks provided in the arguments. Note that the division operator `/` should be considered as an integer division for this exercise. This means that you need to use `//` in Python.
class Stack:
pass
# +
class EvaluateExpression:
# copy the other definitions
# from the previous parts
def process_operator(self, operand_stack, operator_stack):
pass
# -
expr1 = EvaluateExpression()
operand_stack = Stack()
operator_stack = Stack()
operand_stack.push(3)
operand_stack.push(4)
operator_stack.push("+")
expr1.process_operator(operand_stack, operator_stack)
assert operand_stack.peek() == 7
operand_stack.push(5)
operator_stack.push("*")
expr1.process_operator(operand_stack, operator_stack)
assert operand_stack.peek() == 35
operand_stack.push(30)
operator_stack.push("-")
expr1.process_operator(operand_stack, operator_stack)
assert operand_stack.peek() == 5
operand_stack.push(2)
operator_stack.push("/")
expr1.process_operator(operand_stack, operator_stack)
assert operand_stack.peek() == 2
# **Q5.** The class `EvaluateExpression` also has the following methods:
# - `evaluate()`: which evaluate the mathematical expression contained in the property `expression`. The method should return an Integer. This method contains two processes:
# - Phase 1: In this phase, the code scans the expression from left to right to extract operands, operators, and the parentheses.
# 1. If the extracted character is an operand, push it to `operand_stack`.
# 1. If the extracted character is + or - operator, process all the operators at the top of the `operator_stack` and push the extracted operator to `operator_stack`. You should process all the operators as long as the `operator_stack` is not empty and the top of the `operator_stack` is not `(` or `)` symbols.
# 1. If the extracted character is a `*` or `/` operator, process all the `*` or `/` operators at the top of the `operator_stack` and push the extracted operator to `operator_stack`.
# 1. If the extracted character is a `(` symbol, push it to `operator_stack`.
# 1. If the extracted character is a `)` symbol, repeatedly process the operators from the top of `operator_stack` until seeing the `(` symbol on the stack.
# - Phase 2: Repeatedly process the operators from the top of `operator_stack` until `operator_stack` is empty.
#
class EvaluateExpression:
# copy the other definitions
# from the previous parts
def evaluate(self):
operand_stack = Stack()
operator_stack = Stack()
expression = self.insert_space()
tokens = expression.split()
pass
expr1 = EvaluateExpression("(1+2)*3")
assert expr1.evaluate() == 9
expr1.expression = "(1 + 2) * 4 - 3"
assert expr1.evaluate() == 9
expr2 = EvaluateExpression("(1+2 *4- 3)* (7/5 * 6)")
assert expr2.evaluate() == 36
|
mp_calc/mp2_exercises.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %pylab inline
from sympy import symbols
import numpy as np
f = lambda x, y: x * y
def find_gradient_2d(f):
x, y = symbols('x y')
f = f(x, y)
df_dx = f.diff(x)
df_dy = f.diff(y)
return np.array([df_dx, df_dy])
def eval_gradient_2d(grad, p):
x = grad[0].subs('x', p[0]).subs('y', p[1])
y = grad[1].subs('x', p[0]).subs('y', p[1])
return np.array([x.evalf(), y.evalf()])
def find_directional_deriv(grad, direction):
x = grad[0] * direction[0]
y = grad[1] * direction[1]
return np.array([x, y])
def eval_directional_deriv(dir_deriv, p):
x = dir_deriv[0].subs('x', p[0]).subs('y', p[1])
y = dir_deriv[1].subs('x', p[0]).subs('y', p[1])
return np.array([x.evalf(), y.evalf()])
grad = find_gradient_2d(f)
direction = np.array([1,0])
dir_deriv = find_directional_deriv(grad, direction)
dir_deriv
eval_gradient_2d(grad, [1,1])
# +
X, Y = np.meshgrid(np.arange(1, 10, 0.25), np.arange(1, 10, 0.25))
G = [eval_gradient_2d(grad, [x, y]) for x in X for y in Y]
print(G)
U = [g[0] for g in G]
V = [g[1] for g in G]
print(U,V)
fig, ax = plt.subplots(figsize=(12,12))
q = ax.quiver(X, Y, U, V)
ax.quiverkey(q, X=0.3, Y=1.1, U=10, label='Quiver key, length = 10', labelpos='E')
# -
|
Applied Math/Y2S2/.ipynb_checkpoints/Gradients & Directional Derivatives-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.datasets import make_blobs
datos= make_blobs(n_samples=200, n_features=2, centers=4)#200 ejemplos caracteristicas2 grupos4
datos
plt.scatter(datos[0][:,0], datos[0][:,1])
plt.show()
from sklearn.cluster import KMeans
modelo = KMeans(n_clusters=4)
modelo.fit(datos[0])
modelo.cluster_centers_
modelo.labels_
datos[1]
# +
fig, (ax1,ax2)= plt.subplots(1,2, figsize=(12,4))
ax1.scatter(datos[0][:,0], datos[0][:,1], c= modelo.labels_)
ax1.set_title('Algoritmo de K-medias')
ax2.scatter(datos[0][:,0], datos[0][:,1], c= datos[1])
ax2.set_title('Datos originales')
# -
flores = sns.load_dataset('iris')
x=flores.drop('species', axis=1)
modelo2 = KMeans(n_clusters=3)
modelo2.fit(x)
modelo2.cluster_centers_
modelo2.labels_
plt.scatter(modelo2.labels_,modelo2.cluster_centers_)
plt.show()
|
.ipynb_checkpoints/K-medias-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import math
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
# palette_colors = sns.color_palette('tab10')
# palette_dict = {continent: color for continent, color in zip(set(data.type), palette_colors)}
# import matplotlib.pyplot as plt
# import seaborn as sns
# -
data = pd.read_csv(r"C:\Users\pavan\Desktop\Mahesh_Visualization_Assignment\Sacramentorealestatetransactions.csv")
print("Total Zip Codes: ",len(data.zip.unique()))
print('\nSample:\n',(set(data.zip)))
data[data.zip==95648]
data.zip.value_counts()
zip_codes=set(data.zip)
# * Q1.Create a `grid of scatter plots` with each one representing the `sq_ft distribution` in a `single zipcode`, please also include ticks, labels and legend in your plot
# +
# grid_col_count = math.floor(len(zip_codes)**0.5)
# cnt =-1
# zip_codes = list(set(data.zip))
# fig,axes = plt.subplots(grid_col_count+1,grid_col_count,figsize=(15,15))
# for i in range(grid_col_count):
# for j in range(grid_col_count):
# cnt+=1
# axes[i,j].hist(data.loc[data['zip']==zip_codes[cnt],'sq__ft'], alpha=0.3)
# axes[i,j].axes.set_title(f"ZIP:{zip_codes[cnt]}",fontsize=10)
# for index,value in enumerate(zip_codes[cnt+1:]):
# axes[i+1,index].hist(data.loc[data['zip']==value,'sq__ft'], alpha=0.3)
# for i in range(4,8):
# fig.delaxes(axes[8,i])
# plt.subplots_adjust(wspace=0.5, hspace=0.8)
# # print(zip_codes[cnt],i,j)
# # print(zip_codes[cnt+1:])
# +
# fig = plt.figure() #create an empty plot windown
# for i in range(1,69):
# ax = fig.add_subplot(9,9,i)
# ax.scatter(np.arange(30), np.arange(30) + 3 * np.random.randn(30))
# -
n=math.floor(len(zip_codes)**0.5)+1
fig = plt.figure(figsize=(20,20))
for i,zip_code in enumerate(zip_codes):
ax = fig.add_subplot(n,n,i+1)
ax.hist(data.loc[data['zip']==zip_code,'sq__ft'].values, alpha=0.3)
ax.set_title(f"ZIP:{zip_code}",fontsize=10)
plt.subplots_adjust(wspace=0.5, hspace=0.8)
# * Q2.Create a grid of scatter plots with each one representing the `price distribution in a single zipcode`, annotate the `highest` and `lowest` price ones for each category of real estate: condo, residential and multi-family, please also include ticks, labels and legend in your plot
n=math.floor(len(zip_codes)**0.5)+1
fig = plt.figure(figsize=(20,20))
for i,zip_code in enumerate(zip_codes):
ax = fig.add_subplot(n,n,i+1)
ax.hist(data.loc[data['zip']==zip_code,'price'].values, alpha=0.3)
ax.set_title(f"ZIP:{zip_code}",fontsize=10)
# ax.text("max,min")
plt.subplots_adjust(wspace=0.5, hspace=0.8)
# * Q3.Create a grid of bar plots with each one representing a single zipcode and in that zipcode the sq_ft distribution is grouped by the category of condo, residential and multi-family, please also include ticks, labels and legend in your plot
# +
n=math.floor(len(zip_codes)**0.5)+1
ax_objs=[]
fig = plt.figure(figsize=(20,20))
for i,zip_code in enumerate(zip_codes):
ax = fig.add_subplot(n,n,i+1)
ax_objs.append(ax)
filter_data=data[data['zip']==zip_code]
sns.histplot(data=filter_data, x="sq__ft",hue='type')
ax.legend([],[], frameon=False)
ax.set_title(f"ZIP:{zip_code}",fontsize=10)
plt.subplots_adjust(wspace=0.5, hspace=0.8)
# ax.text("max,min")
# labels_handles = {
# label: handle for ax in fig.axes for handle, label in zip(*ax.get_legend_handles_labels())
# }
# fig.legend(
# labels_handles.values(),
# labels_handles.keys(),
# loc="upper center",
# bbox_to_anchor=(0.5, 0),
# bbox_transform=plt.gcf().transFigure,
# )
# plt.subplots_adjust(wspace=0.5, hspace=0.8)
# lines_labels = [ax.get_legend_handles_labels() for ax in fig.axes]
# lines, labels = [sum(lol, []) for lol in zip(*lines_labels)]
# # fig.legend(lines, labels)
# plt.show()
# fig.legend(ax_objs,labels=set(data.type),loc="upper right")
# -
# * Q4.Create a grid of bar plots with each one representing a single zipcode and in that zipcode the price distribution is grouped by the category of condo, residential and multi-family, please also include ticks, labels and legend in your plot
n=math.floor(len(zip_codes)**0.5)+1
ax_objs=[]
fig = plt.figure(figsize=(20,20))
for i,zip_code in enumerate(zip_codes):
ax = fig.add_subplot(n,n,i+1)
ax_objs.append(ax)
filter_data=data[data['zip']==zip_code]
sns.histplot(data=filter_data, x="price",hue='type')
ax.legend([],[], frameon=False)
ax.set(ylabel=None)
# ax.set(xticklabels=[])
ax.set(yticklabels=[])
ax.set_title(f"ZIP:{zip_code}",fontsize=10)
plt.subplots_adjust(wspace=0.5, hspace=1.0)
# * Plot the average price distribution based on zipcode for each category of real estate: condo, residential and multi-family, please also include ticks, labels and legend in your plot
z=data.groupby(['zip','type']).agg({'price': 'mean'}).reset_index()
plt.figure(figsize=(20,8))
# ax = sns.barplot(data=df, x='reads', estimator=len,y='interest', hue='interest' )
ax = sns.barplot(x='zip', y='price', data=z,hue='type',dodge=False,alpha=1)
h, l = ax.get_legend_handles_labels()
# ax.legend(h, labels, title="Interested in politics")
plt.xticks(size=15,rotation=90)
plt.yticks(size=15,rotation=0)
plt.xlabel('Zip Code', fontsize=18)
plt.ylabel('Price', fontsize=18)
plt.title("Average Price Distribution of ZIP's for all types",fontsize=19)
plt.legend(fontsize='x-large')
plt.show()
# * Plot the average price distribution based on city for each category of real estate: condo, residential and multi-family, please also include ticks, labels and legend in your plot
# +
z=data.groupby(['city','type']).agg({'price': 'mean'}).reset_index()
# z['city'] = z['city'].astype("int64")
plt.figure(figsize=(20,8))
# ax = sns.barplot(data=df, x='reads', estimator=len,y='interest', hue='interest' )
ax = sns.barplot(x='city', y='price', data=z,hue='type',dodge=False,alpha=1)
h, l = ax.get_legend_handles_labels()
# ax.legend(h, labels, title="Interested in politics")
plt.xticks(size=15,rotation=90)
plt.yticks(size=15,rotation=0)
plt.xlabel('City', fontsize=16)
plt.ylabel('Price', fontsize=16)
plt.legend(fontsize='large')
plt.title("Average Price Distribution of Cities for all types",fontsize=19)
plt.show()
# -
# ### SP500.csv file
# * Plot daily gain/loss for January of 2018, annotate the highest daily gain and its date, the highest daily loss and its date in January 2018
data = pd.read_csv(r"C:\Users\pavan\Desktop\Mahesh_Visualization_Assignment\SP500.csv")
data.head()
data["Date_"] = pd.to_datetime(data.Date,format='%Y-%m-%d')
data['Gain'] = data.Close - data.Open
# +
# data[['Gain']].sort_values(['Gain'],ascending=0)
# -
data = data.loc[data["Date_"].dt.year==2018,]
print("Total records in 2018: ",data.shape[0])
# +
import matplotlib.dates as mdates
fig=plt.figure(figsize=(15,8))
ax = fig.add_subplot(111)
ax = sns.barplot(x='Date_', y='Gain', data=data,color='blue')
plt.xticks(size=8,rotation=90)
plt.show()
ymax = max(data.Gain)
xpos = data[['Gain']].idxmax(axis = 0, skipna = True)[0]
xmax = data[['Date']].iloc[data.index==17134].values[0][0]#x[xpos]
ax.xaxis.set_major_formatter(mdates.DateFormatter('%Y-%m'))
# ax.annotate('local max', xy=('2018-02-06', 80), xytext=('2018-02-04', 25),
# arrowprops=dict(facecolor='black'),
# )
# plt.show()
# plt.savefig("test"+'.png')
# -
# * Make pair plot matrix of January 2018 SP500 data on high, low, adj close and volumn
plt.figure(figsize=(15,8))
sns.pairplot(data[['High','Low','Adj Close','Volume']])
plt.show()
|
Mahesh/Data Visualization/Visualization.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Training AU visualization model
# You will first need to gather the datasets for training. In this tutorial we use the datasets EmotioNet, DISFA Plus, and BP4d. After you download each model you should extract the labels and landmarks from each dataset. Detailed code on how to do that is described at the bottom of this tutorial. Once you have the labels and landmark files for each dataset you can train the AU visualization model with the following.
# +
# %matplotlib inline
import pandas as pd, numpy as np, matplotlib.pyplot as plt
from sklearn.cross_decomposition import PLSRegression
from sklearn.model_selection import KFold
from feat.plotting import predict, plot_face
from feat.utils import registration, neutral
from natsort import natsorted
import os, glob
import pandas as pd, numpy as np
import seaborn as sns
sns.set_style("white")
au_cols = [1, 2, 4, 5, 6, 7, 9, 10, 12, 14, 15, 17, 18, 20, 23, 24, 25, 26, 28, 43]
au_cols = [f"AU{au}" for au in au_cols]
base_dir = "/Storage/Projects/feat_benchmark/scripts/jcheong/openface_train"
labels_emotionet = pd.read_csv(os.path.join(base_dir, "emotionet_labels.csv"))
landmarks_emotionet = pd.read_csv(os.path.join(base_dir, "emotionet_landmarks.csv"))
print("EmotioNet: ", len(labels_emotionet))
labels_disfaplus = pd.read_csv(os.path.join(base_dir, "disfaplus_labels.csv"))
landmarks_disfaplus = pd.read_csv(os.path.join(base_dir, "disfaplus_landmarks.csv"))
# Disfa is rescaled to 0 - 1
disfaplus_aus = [col for col in labels_disfaplus.columns if "AU" in col]
labels_disfaplus[disfaplus_aus] = labels_disfaplus[disfaplus_aus].astype('float')/5
print("DISFA Plus: ", len(labels_disfaplus))
labels_bp4d = pd.read_csv(os.path.join(base_dir, "bp4d_labels.csv"))
landmarks_bp4d = pd.read_csv(os.path.join(base_dir, "bp4d_landmarks.csv"))
bp4d_pruned_idx = labels_bp4d.replace({9: np.nan})[au_cols].dropna(axis=1).index
print("BP4D: ", len(labels_bp4d))
# -
# We aggregate the datasets and specify the AUs we want to train.
# +
labels = pd.concat([
labels_emotionet.replace({999: np.nan}),
labels_disfaplus,
labels_bp4d.replace({9: np.nan}).iloc[bp4d_pruned_idx,:]
]).reset_index(drop=True)
landmarks = pd.concat([
landmarks_emotionet,
landmarks_disfaplus,
landmarks_bp4d.iloc[bp4d_pruned_idx,:]
]).reset_index(drop=True)
landmarks = landmarks.iloc[labels.index]
labels = labels[au_cols].fillna(0)
# -
# We train our model using PLSRegression with a minimum of 500 samples for each AU activation. We evaluate the model in a 3-fold split and retrain the model with all the data which is distributed with the package.
# +
min_pos_sample = 500
print('Pseudo balancing samples')
balY = pd.DataFrame()
balX = pd.DataFrame()
for AU in labels[au_cols].columns:
if np.sum(labels[AU]==1) > min_pos_sample:
replace = False
else:
replace = True
newSample = labels[labels[AU]>.5].sample(min_pos_sample, replace=replace, random_state=0)
balX = pd.concat([balX, newSample])
balY = pd.concat([balY, landmarks.loc[newSample.index]])
X = balX[au_cols].values
y = registration(balY.values, neutral)
# Model Accuracy in KFold CV
print("Evaluating model with KFold CV")
n_components=len(au_cols)
kf = KFold(n_splits=3)
scores = []
for train_index, test_index in kf.split(X):
X_train,X_test = X[train_index],X[test_index]
y_train,y_test = y[train_index],y[test_index]
clf = PLSRegression(n_components=n_components, max_iter=2000)
clf.fit(X_train,y_train)
scores.append(clf.score(X_test,y_test))
print('3-fold accuracy mean', np.round(np.mean(scores),2))
# Train real model
clf = PLSRegression(n_components=n_components, max_iter=2000)
clf.fit(X,y)
print('N_comp:',n_components,'Rsquare', np.round(clf.score(X,y),2))
# -
X.shape
# We visualize the results of our model. The regression was trained on labels 0-1 so we do not recommend exceeding 1 for the intensity. Setting the intensity to 2 will exaggerate the face and anything beyond that might give you strange faces.
# Plot results for each action unit
f,axes = plt.subplots(5,4,figsize=(12,18))
axes = axes.flatten()
# Exaggerate the intensity of the expression for clearer visualization.
# We do not recommend exceeding 2.
intensity = 2
for aui, auname in enumerate(axes):
try:
auname=au_cols[aui]
au = np.zeros(clf.n_components)
au[aui] = intensity
predicted = clf.predict([au]).reshape(2,68)
plot_face(au=au, model=clf,
vectorfield={"reference": neutral.T, 'target': predicted,
'color':'r','alpha':.6},
ax = axes[aui])
axes[aui].set(title=auname)
except:
pass
finally:
ax = axes[aui]
ax.axes.get_xaxis().set_visible(False)
ax.axes.get_yaxis().set_visible(False)
# Here is how we would export our model into an h5 format which can be loaded using our load_h5 function.
# +
# save out trained model
# import h5py
# hf = h5py.File('../feat/resources/pyfeat_aus_to_landmarks.h5', 'w')
# hf.create_dataset('coef', data=clf.coef_)
# hf.create_dataset('x_mean', data=clf._x_mean)
# hf.create_dataset('x_std', data=clf._x_std)
# hf.create_dataset('y_mean', data=clf._y_mean)
# hf.close()
# -
# Load h5 model
from feat.utils import load_h5
clf = load_h5('../../feat/resources/pyfeat_aus_to_landmarks.h5')
# ## Preprocessing datasets
# Here we provide sample code for how you might preprocess the datasets to be used in this tutorial.
#
#
# +
from PIL import Image, ImageOps
import math, cv2, csv
from scipy.spatial import ConvexHull
from skimage.morphology.convex_hull import grid_points_in_poly
from feat import Detector
import os, glob, pandas as pd, numpy as np
import matplotlib.pyplot as plt
from skimage import data, exposure
from skimage.feature import hog
from tqdm import tqdm
def padding(img, expected_size):
desired_size = expected_size
delta_width = desired_size - img.size[0]
delta_height = desired_size - img.size[1]
pad_width = delta_width // 2
pad_height = delta_height // 2
padding = (pad_width, pad_height, delta_width - pad_width, delta_height - pad_height)
return ImageOps.expand(img, padding)
def resize_with_padding(img, expected_size):
img.thumbnail((expected_size[0], expected_size[1]))
delta_width = expected_size[0] - img.size[0]
delta_height = expected_size[1] - img.size[1]
pad_width = delta_width // 2
pad_height = delta_height // 2
padding = (pad_width, pad_height, delta_width - pad_width, delta_height - pad_height)
return ImageOps.expand(img, padding)
def align_face_68pts(img, img_land, box_enlarge, img_size=112):
"""
img: image
img_land: landmarks 68
box_enlarge: relative size of face
img_size = 112
"""
leftEye0 = (img_land[2 * 36] + img_land[2 * 37] + img_land[2 * 38] + img_land[2 * 39] + img_land[2 * 40] +
img_land[2 * 41]) / 6.0
leftEye1 = (img_land[2 * 36 + 1] + img_land[2 * 37 + 1] + img_land[2 * 38 + 1] + img_land[2 * 39 + 1] +
img_land[2 * 40 + 1] + img_land[2 * 41 + 1]) / 6.0
rightEye0 = (img_land[2 * 42] + img_land[2 * 43] + img_land[2 * 44] + img_land[2 * 45] + img_land[2 * 46] +
img_land[2 * 47]) / 6.0
rightEye1 = (img_land[2 * 42 + 1] + img_land[2 * 43 + 1] + img_land[2 * 44 + 1] + img_land[2 * 45 + 1] +
img_land[2 * 46 + 1] + img_land[2 * 47 + 1]) / 6.0
deltaX = (rightEye0 - leftEye0)
deltaY = (rightEye1 - leftEye1)
l = math.sqrt(deltaX * deltaX + deltaY * deltaY)
sinVal = deltaY / l
cosVal = deltaX / l
mat1 = np.mat([[cosVal, sinVal, 0], [-sinVal, cosVal, 0], [0, 0, 1]])
mat2 = np.mat([[leftEye0, leftEye1, 1], [rightEye0, rightEye1, 1], [img_land[2 * 30], img_land[2 * 30 + 1], 1],
[img_land[2 * 48], img_land[2 * 48 + 1], 1], [img_land[2 * 54], img_land[2 * 54 + 1], 1]])
mat2 = (mat1 * mat2.T).T
cx = float((max(mat2[:, 0]) + min(mat2[:, 0]))) * 0.5
cy = float((max(mat2[:, 1]) + min(mat2[:, 1]))) * 0.5
if (float(max(mat2[:, 0]) - min(mat2[:, 0])) > float(max(mat2[:, 1]) - min(mat2[:, 1]))):
halfSize = 0.5 * box_enlarge * float((max(mat2[:, 0]) - min(mat2[:, 0])))
else:
halfSize = 0.5 * box_enlarge * float((max(mat2[:, 1]) - min(mat2[:, 1])))
scale = (img_size - 1) / 2.0 / halfSize
mat3 = np.mat([[scale, 0, scale * (halfSize - cx)], [0, scale, scale * (halfSize - cy)], [0, 0, 1]])
mat = mat3 * mat1
aligned_img = cv2.warpAffine(img, mat[0:2, :], (img_size, img_size), cv2.INTER_LINEAR, borderValue=(128, 128, 128))
land_3d = np.ones((int(len(img_land)/2), 3))
land_3d[:, 0:2] = np.reshape(np.array(img_land), (int(len(img_land)/2), 2))
mat_land_3d = np.mat(land_3d)
new_land = np.array((mat * mat_land_3d.T).T)
new_land = np.array(list(zip(new_land[:,0], new_land[:,1]))).astype(int)
return aligned_img, new_land
def extract_hog(image, detector):
im = cv2.imread(image)
detected_faces = np.array(detector.detect_faces(im)[0])
if np.any(detected_faces<0):
orig_size = np.array(im).shape
if np.where(detected_faces<0)[0][0]==1:
new_size = (orig_size[0], int(orig_size[1] + 2*abs(detected_faces[detected_faces<0][0])))
else:
new_size = (int(orig_size[0] + 2*abs(detected_faces[detected_faces<0][0])), orig_size[1])
im = resize_with_padding(Image.fromarray(im), new_size)
im = np.asarray(im)
detected_faces = np.array(detector.detect_faces(np.array(im))[0])
detected_faces = detected_faces.astype(int)
points = detector.detect_landmarks(np.array(im), [detected_faces])[0].astype(int)
aligned_img, points = align_face_68pts(im, points.flatten(), 2.5)
hull = ConvexHull(points)
mask = grid_points_in_poly(shape=np.array(aligned_img).shape,
verts= list(zip(points[hull.vertices][:,1], points[hull.vertices][:,0])) # for some reason verts need to be flipped
)
mask[0:np.min([points[0][1], points[16][1]]), points[0][0]:points[16][0]] = True
aligned_img[~mask] = 0
resized_face_np = aligned_img
fd, hog_image = hog(resized_face_np, orientations=8, pixels_per_cell=(8, 8),
cells_per_block=(2, 2), visualize=True, multichannel=True)
return fd, hog_image, points
# -
# Replace the paths so that it points to your local dataset directory.
# +
detector = Detector(face_model = "retinaface", landmark_model="mobilenet")
# Correct path to your downloaded dataset.
EmotioNet_images = np.sort(glob.glob("/Storage/Data/EmotioNet/imgs/*.jpg"))
labels = pd.read_csv("/Storage/Data/EmotioNet/labels/EmotioNet_FACS_aws_2020_24600.csv")
labels = labels.dropna(axis=0)
for col in labels.columns:
if "AU" in col:
kwargs = {col.replace("'", '').replace('"', '').replace(" ",""): labels[[col]]}
labels = labels.assign(**kwargs)
labels = labels.drop(columns = col)
labels = labels.assign(URL = labels.URL.apply(lambda x: x.split("/")[-1].replace("'", "")))
labels = labels.set_index('URL')
labels = labels.drop(columns = ["URL orig"])
aus_to_train = ['AU1','AU2','AU4','AU5', "AU6", "AU9","AU10", "AU12", "AU15","AU17",
"AU18","AU20", "AU24", "AU25", "AU26", "AU28", "AU43"]
with open('emotionet_labels.csv', "w", newline='') as csvfile:
writer = csv.writer(csvfile, delimiter=',')
writer.writerow(["URL"] + aus_to_train)
landmark_cols = [f"x_{i}" for i in range(68)] + [f"y_{i}" for i in range(68)]
with open('emotionet_landmarks.csv', "w", newline='') as csvfile:
writer = csv.writer(csvfile, delimiter=',')
writer.writerow(landmark_cols)
for ix, image in enumerate(tqdm(EmotioNet_images)):
try:
imageURL = os.path.split(image)[-1]
label = labels.loc[imageURL][aus_to_train]
fd, _, points = extract_hog(image, detector=detector)
with open('emotionet_labels.csv', "a+", newline='') as csvfile:
writer = csv.writer(csvfile, delimiter=',')
writer.writerow([imageURL]+list(label.values))
with open('emotionet_landmarks.csv', "a+", newline='') as csvfile:
writer = csv.writer(csvfile, delimiter=',')
writer.writerow(points.T.flatten())
except:
print(f"failed {image}")
|
notebooks/_build/html/_sources/content/dev_trainAUvisModel.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_mxnet_p36
# language: python
# name: conda_mxnet_p36
# ---
# # Amazon SageMaker を使用した $K$-means クラスタリング
#
# - 次の AWS ブログの内容を元にしたノートブックです [[blog](https://aws.amazon.com/jp/blogs/news/k-means-clustering-with-amazon-sagemaker/)]
# - SageMaker ビルトインである $k$-means クラスタリングを実演
# - 実装のベースは [[Scully'10](https://www.eecs.tufts.edu/~dsculley/papers/fastkmeans.pdf), [Mayerson'01](http://web.cs.ucla.edu/~awm/papers/ofl.pdf), [Guha et al.'03](https://papers.nips.cc/paper/4362-fast-and-accurate-k-means-for-large-datasets.pdf)]
import boto3
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from IPython.display import display
import io
import time
import copy
import json
import sys
import sagemaker.amazon.common as smac
import os
import mxnet as mx
from scipy.spatial.distance import cdist
import numpy as np
from numpy import array
import urllib.request
import gzip
import pickle
import sklearn.cluster
import sklearn
import re
import sagemaker
# +
# S3 バケットとプレフィックス
sess = sagemaker.Session()
bucket = sess.default_bucket()
prefix = 'sagemaker/DEMO-kmeans'
role = sagemaker.get_execution_role()
def get_gdelt(filename):
s3 = boto3.resource('s3')
s3.Bucket('gdelt-open-data').download_file('events/' + filename, '.gdelt.csv')
df = pd.read_csv('.gdelt.csv', sep='\t')
header = pd.read_csv('https://www.gdeltproject.org/data/lookups/CSV.header.historical.txt', sep='\t')
df.columns = header.columns
return df
data = get_gdelt('1979.csv')
data
# +
data = data[['EventCode', 'NumArticles', 'AvgTone', 'Actor1Geo_Lat', 'Actor1Geo_Long', 'Actor2Geo_Lat', 'Actor2Geo_Long']]
data['EventCode'] = data['EventCode'].astype(object)
events = pd.crosstab(index=data['EventCode'], columns='count').sort_values(by='count', ascending=False).index[:20]
#トレーニングデータを Sagemaker K-means に必要な protobuf 形式に変換するルーチン
def write_to_s3(bucket, prefix, channel, file_prefix, X):
buf = io.BytesIO()
smac.write_numpy_to_dense_tensor(buf, X.astype('float32'))
buf.seek(0)
boto3.Session().resource('s3').Bucket(bucket).Object(os.path.join(prefix, channel, file_prefix + '.data')).upload_fileobj(buf)
#上記のアクター場所とイベントに基づいて、データをフィルタリング
def transform_gdelt(df, events=None):
df = df[['AvgTone', 'EventCode', 'NumArticles', 'Actor1Geo_Lat', 'Actor1Geo_Long', 'Actor2Geo_Lat', 'Actor2Geo_Long']]
df['EventCode'] = df['EventCode'].astype(object)
if events is not None:
df = df[np.in1d(df['EventCode'], events)]
return pd.get_dummies(df[((df['Actor1Geo_Lat'] == 0) & (df['Actor1Geo_Long'] == 0) != True) &
((df['Actor2Geo_Lat'] == 0) & (df['Actor2Geo_Long'] == 0) != True)])
#トレーニングを準備し、S3 に保存
def prepare_gdelt(bucket, prefix, file_prefix, events=None, random_state=1729, save_to_s3=True):
df = get_gdelt(file_prefix + '.csv')
model_data = transform_gdelt(df, events)
train_data = model_data.sample(frac=1, random_state=random_state).as_matrix()
if save_to_s3:
write_to_s3(bucket, prefix, 'train', file_prefix, train_data)
return train_data
# 1979 年用のデータセットを使用。
train_79 = prepare_gdelt(bucket, prefix, '1979', events, save_to_s3=False)
# +
# 1979 年のデータセットから最初の 10000 データポイントを可視化するために TSNE を使用
from sklearn import manifold
tsne = manifold.TSNE(n_components=2, init='pca', random_state=1200)
X_tsne = tsne.fit_transform(train_79[:10000])
plt.figure(figsize=(6, 5))
X_tsne_1000 = X_tsne[:1000]
plt.scatter(X_tsne_1000[:, 0], X_tsne_1000[:, 1])
plt.show()
# +
BEGIN_YEAR = 1979
END_YEAR = 1980
for year in range(BEGIN_YEAR, END_YEAR):
train_data = prepare_gdelt(bucket, prefix, str(year), events)
# SageMaker k-means ECR image ARN
images = {'us-west-2': '174872318107.dkr.ecr.us-west-2.amazonaws.com/kmeans:latest',
'us-east-1': '382416733822.dkr.ecr.us-east-1.amazonaws.com/kmeans:latest',
'us-east-2': '404615174143.dkr.ecr.us-east-2.amazonaws.com/kmeans:latest',
'eu-west-1': '438346466558.dkr.ecr.eu-west-1.amazonaws.com/kmeans:latest'}
image = images[boto3.Session().region_name]
# +
from time import gmtime, strftime
output_time = strftime("%Y-%m-%d-%H-%M-%S", gmtime())
output_folder = 'kmeans-lowlevel-' + output_time
K = range(2, 12) # k が使用する範囲を変更
INSTANCE_COUNT = 1
run_parallel_jobs = True #一度に 1 つのジョブを実行するには、これを false にします。
#特に多数の EC2 インスタンスを 1 度に作成し、上限に達するのを避けたい場合
job_names = []
sagemaker_client = boto3.client('sagemaker')
# すべての k でジョブを起動する
for k in K:
print('starting train job:'+ str(k))
output_location = 's3://{}/kmeans_example/output/'.format(bucket) + output_folder
print('training artifacts will be uploaded to: {}'.format(output_location))
job_name = output_folder + str(k)
create_training_params = \
{
"AlgorithmSpecification": {
"TrainingImage": image,
"TrainingInputMode": "File"
},
"RoleArn": role,
"OutputDataConfig": {
"S3OutputPath": output_location
},
"ResourceConfig": {
"InstanceCount": INSTANCE_COUNT,
"InstanceType": "ml.c5.18xlarge",
"VolumeSizeInGB": 50
},
"TrainingJobName": job_name,
"HyperParameters": {
"k": str(k),
"feature_dim": "26",
"mini_batch_size": "1000"
},
"StoppingCondition": {
"MaxRuntimeInSeconds": 60 * 60
},
"InputDataConfig": [
{
"ChannelName": "train",
"DataSource": {
"S3DataSource": {
"S3DataType": "S3Prefix",
"S3Uri": "s3://{}/{}/train/".format(bucket, prefix),
"S3DataDistributionType": "FullyReplicated"
}
},
"CompressionType": "None",
"RecordWrapperType": "None"
}
]
}
sagemaker_client.create_training_job(**create_training_params)
status = sagemaker_client.describe_training_job(TrainingJobName=job_name)['TrainingJobStatus']
print(status)
if not run_parallel_jobs:
try:
sagemaker_client.get_waiter('training_job_completed_or_stopped').wait(TrainingJobName=job_name)
finally:
status = sagemaker_client.describe_training_job(TrainingJobName=job_name)['TrainingJobStatus']
print("Training job ended with status: " + status)
if status == 'Failed':
message = sagemaker_client.describe_training_job(TrainingJobName=job_name)['FailureReason']
print('Training failed with the following error: {}'.format(message))
raise Exception('Training job failed')
job_names.append(job_name)
# -
while len(job_names):
try:
sagemaker_client.get_waiter('training_job_completed_or_stopped').wait(TrainingJobName=job_names[0])
finally:
status = sagemaker_client.describe_training_job(TrainingJobName=job_name)['TrainingJobStatus']
print("Training job ended with status: " + status)
if status == 'Failed':
message = sagemaker_client.describe_training_job(TrainingJobName=job_name)['FailureReason']
print('Training failed with the following error: {}'.format(message))
raise Exception('Training job failed')
print(job_name)
info = sagemaker_client.describe_training_job(TrainingJobName=job_name)
job_names.pop(0)
colors = ['b', 'g', 'r']
markers = ['o', 'v', 's']
models = {}
distortions = []
for k in K:
s3_client = boto3.client('s3')
key = 'kmeans_example/output/' + output_folder +'/' + output_folder + str(k) + '/output/model.tar.gz'
s3_client.download_file(bucket, key, 'model.tar.gz')
print("Model for k={} ({})".format(k, key))
# !tar -xvf model.tar.gz
kmeans_model=mx.ndarray.load('model_algo-1')
kmeans_numpy = kmeans_model[0].asnumpy()
distortions.append(sum(np.min(cdist(train_data, kmeans_numpy, 'euclidean'), axis=1)) / train_data.shape[0])
models[k] = kmeans_numpy
# エルボーをプロット
plt.plot(K, distortions, 'bx-')
plt.xlabel('k')
plt.ylabel('distortion')
plt.title('Elbow graph')
plt.show()
# ## Set up hosting for the model
# 以下では [[notebook](https://github.com/hariby/amazon-sagemaker-examples/blob/master/sagemaker-python-sdk/1P_kmeans_lowlevel/kmeans_mnist_lowlevel.ipynb)] を参考にして、推論用のエンドポイントを構築します。
#
# In order to set up hosting, we have to import the model from training to hosting. A common question would be, why wouldn't we automatically go from training to hosting? And, in fact, the [k-means high-level example](/notebooks/sagemaker-python-sdk/1P_kmeans_highlevel/kmeans_mnist.ipynb) shows the functionality to do that. For this low-level example though it makes sense to show each step in the process to provide a better understanding of the flexibility available.
#
# ### Import model into hosting
# Next, you register the model with hosting. This allows you the flexibility of importing models trained elsewhere, as well as the choice of not importing models if the target of model creation is AWS Lambda, AWS Greengrass, Amazon Redshift, Amazon Athena, or other deployment target.
# +
primary_container = {
'Image': image,
'ModelDataUrl': info['ModelArtifacts']['S3ModelArtifacts']
}
create_model_response = sagemaker_client.create_model(
ModelName = job_name,
ExecutionRoleArn = role,
PrimaryContainer = primary_container
)
print(create_model_response['ModelArn'])
# -
# ### Create endpoint configuration
# Now, we'll create an endpoint configuration which provides the instance type and count for model deployment.
# +
endpoint_config_name = 'KMeansEndpointConfig-' + strftime("%Y-%m-%d-%H-%M-%S", gmtime())
print(endpoint_config_name)
create_endpoint_config_response = sagemaker_client.create_endpoint_config(
EndpointConfigName = endpoint_config_name,
ProductionVariants=[{
'InstanceType':'ml.m4.xlarge',
'InitialInstanceCount':1,
'ModelName':job_name,
'VariantName':'AllTraffic'}])
print("Endpoint Config Arn: " + create_endpoint_config_response['EndpointConfigArn'])
# -
# ### Create endpoint
# Lastly, the customer creates the endpoint that serves up the model, through specifying the name and configuration defined above. The end result is an endpoint that can be validated and incorporated into production applications. This takes 9-11 minutes to complete.
# +
# %%time
endpoint_name = 'KMeansEndpoint-' + strftime("%Y-%m-%d-%H-%M-%S", gmtime())
print(endpoint_name)
create_endpoint_response = sagemaker_client.create_endpoint(
EndpointName=endpoint_name,
EndpointConfigName=endpoint_config_name)
print(create_endpoint_response['EndpointArn'])
resp = sagemaker_client.describe_endpoint(EndpointName=endpoint_name)
status = resp['EndpointStatus']
print("Status: " + status)
try:
sagemaker_client.get_waiter('endpoint_in_service').wait(EndpointName=endpoint_name)
finally:
resp = sagemaker_client.describe_endpoint(EndpointName=endpoint_name)
status = resp['EndpointStatus']
print("Arn: " + resp['EndpointArn'])
print("Create endpoint ended with status: " + status)
if status != 'InService':
message = sagemaker_client.describe_endpoint(EndpointName=endpoint_name)['FailureReason']
print('Training failed with the following error: {}'.format(message))
raise Exception('Endpoint creation did not succeed')
# -
# ## Validate the model for use
# Finally, we'll validate the model for use. Let's generate a classification for a single observation from the trained model using the endpoint we just created.
# Simple function to create a csv from our numpy array
def np2csv(arr):
csv = io.BytesIO()
np.savetxt(csv, arr, delimiter=',', fmt='%g')
return csv.getvalue().decode().rstrip()
runtime = boto3.Session().client('runtime.sagemaker')
# +
payload = np2csv(train_79[0:10000])
response = runtime.invoke_endpoint(EndpointName=endpoint_name,
ContentType='text/csv',
Body=payload)
result = json.loads(response['Body'].read().decode())
cluster_ids = np.array([int(result['predictions'][i]['closest_cluster']) for i in range(len(result['predictions']))])
# -
plt.figure(figsize=(6, 5))
X_tsne_1000 = X_tsne[:1000]
plt.scatter(X_tsne_1000[:, 0], X_tsne_1000[:, 1], c=cluster_ids[:1000])
plt.show()
|
introduction_to_amazon_algorithms/kmeans_news_clustering/kmeans_news_clustering_lowlevel.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Google Colab Setup
# ---
#
# Make sure to select GPU in Runtime > Change runtime type > Hardware accelerator
#@title << Setup Google Colab by running this cell {display-mode: "form"}
import sys
if 'google.colab' in sys.modules:
# Clone GitHub repository
# !git clone https://github.com/pacm/rl-workshop.git
# Copy files required to run the code
# !cp -r "rl-workshop/agents" "rl-workshop/env" "rl-workshop/helpers" "rl-workshop/videos" .
# Install packages via pip
# !pip install -r "rl-workshop/colab-requirements.txt"
# Restart Runtime
import os
os.kill(os.getpid(), 9)
# Imports
# ---
# %run env/env.py
# %run helpers/rl_helpers.py
# %run agents/dqn.py
# %run agents/qlearning.py
# %run agents/random.py
# You might want to import other libraries
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
import numpy as np
from PIL import Image
# Intro to Q-Learning (compass Q-table)
# ---
#
# You can find a Q-learning implementation in `agents/`
#
# ```
# agents/
# ├── curiosity.py
# ├── dqn.py
# ├── logging.py
# ├── qlearning.py <-- Q-learning agent
# └── random.py
# ```
# +
# Environment without Skyscrapers + discharge
env = CompassQTable(DeliveryDrones())
env.env_params.update({'n_drones': 3, 'skyscrapers_factor': 0, 'stations_factor': 0, 'discharge': 0})
states = env.reset()
print('Observation space:', env.observation_space)
print('Initial state:', {drone_index: env.format_state(state) for drone_index, state in states.items()})
Image.fromarray(env.render(mode='rgb_array'))
# -
# Create the agents
agents = {drone.index: RandomAgent(env) for drone in env.drones}
agents[0] = QLearningAgent(
env,
gamma=0.95, # Discount factor
alpha=0.1, # Learning rate
# Exploration rate
epsilon_start=1, epsilon_decay=0.99, epsilon_end=0.01
)
agents
# Train agents
trainer = MultiAgentTrainer(env, agents, reset_agents=True, seed=0)
trainer.train(5000)
plot_rolling_rewards(trainer.rewards_log, drones_labels={0: 'Q-learning'})
agents[0].get_qtable()
plt.plot(agents[0].gamma**np.arange(100))
plt.title('Discount factor: {}'.format(agents[0].gamma))
plt.xlabel('Number of steps')
plt.ylabel('Discount')
plt.show()
rewards_log = test_agents(env, agents, n_steps=1000, seed=0)
plot_cumulative_rewards(rewards_log, drones_labels={0: 'Q-learning'})
path = os.path.join('videos', 'ql-compass.mp4')
render_video(env, agents, video_path=path, n_steps=120, fps=1, seed=0)
ColabVideo(path)
# Scaling Q-learning (compass + lidar Q-table)
# ---
#
# Let's see how Q-learning scales to larger observation spaces
# +
# Environment with skyscrapers but without discharge
env = LidarCompassQTable(DeliveryDrones())
env.env_params.update({'n_drones': 3, 'skyscrapers_factor': 3, 'stations_factor': 0, 'discharge': 0})
states = env.reset()
print('Observation space:', env.observation_space)
print('Sample state:', {drone_index: env.format_state(state) for drone_index, state in states.items()})
Image.fromarray(env.render(mode='rgb_array'))
# -
# Create the agents
agents = {drone.index: RandomAgent(env) for drone in env.drones}
agents[0] = QLearningAgent(
env,
gamma=0.95, # Discount factor
alpha=0.1, # Learning rate
# Exploration rate
epsilon_start=1, epsilon_decay=0.99, epsilon_end=0.01
)
agents
# Train agents
trainer = MultiAgentTrainer(env, agents, reset_agents=True, seed=0)
trainer.train(5000)
plot_rolling_rewards(trainer.rewards_log, drones_labels={0: 'Q-learning'})
rewards_log = test_agents(env, agents, n_steps=1000, seed=0)
plot_cumulative_rewards(rewards_log, drones_labels={0: 'Q-learning'})
path = os.path.join('videos', 'ql-compass-lidar-1st-try.mp4')
render_video(env, agents, video_path=path, n_steps=120, fps=1, seed=0)
ColabVideo(path)
# Issues with Q-learning
# ---
#
# Two issues here
#
# * Sparse "reward signal": pickup rate is around 1%
# * No generalization: need to explore entire space!
q_table = agents[0].get_qtable()
print('Q-table:', q_table.shape)
q_table.sample(10)
plt.plot(agents[0].epsilons)
plt.xlabel('Number of episodes')
plt.ylabel('Exploration rate (epsilon)')
plt.show()
# Possible solutions
# ---
# +
# (1/2) Sparse rewards: Create an intermediate "pickup" reward
env.env_params.update({
'n_drones': 3, 'pickup_reward': 0.99, 'delivery_reward': 1,
'skyscrapers_factor': 3, 'stations_factor': 0, 'discharge': 0})
states = env.reset()
# (2/2) Train longer ..
agents[0].epsilon = 1
agents[0].epsilon_decay = 0.999
set_seed(env, seed=0) # Make things deterministic
trainer.train(30000)
plot_rolling_rewards(
trainer.rewards_log,
events={'pickup': [0.99], 'delivery': [1], 'crash': [-1]},
drones_labels={0: 'Q-learning'})
# -
plt.plot(agents[0].epsilons)
plt.xlabel('Number of episodes')
plt.ylabel('Exploration rate (epsilon)')
plt.show()
rewards_log = test_agents(env, agents, n_steps=1000, seed=0)
plot_cumulative_rewards(
rewards_log,
events={'pickup': [0.99], 'delivery': [1], 'crash': [-1]},
drones_labels={0: 'Q-learning'}
)
# Overfitting issues: try with different seeds
# ---
rewards_log = test_agents(env, agents, n_steps=1000, seed=1)
plot_cumulative_rewards(
rewards_log,
events={'pickup': [0.99], 'delivery': [1], 'crash': [-1]},
drones_labels={0: 'Q-learning'}
)
# Pick a good seed for your video ;)
path = os.path.join('videos', 'ql-compass-lidar-2nd-try.mp4')
render_video(env, agents, video_path=path, n_steps=120, fps=1, seed=1)
ColabVideo(path)
# Q-learning limitations: discrete Q-table!
# ---
#
# Let's try Q-learning with the full environment: skyscrapers + charge
# +
env = LidarCompassChargeQTable(DeliveryDrones())
env.env_params.update({
'n_drones': 3, 'pickup_reward': 0.99, 'delivery_reward': 1,
'discharge': 10, 'charge': 20, 'charge_reward': -0.1 # (default values)
})
states = env.reset()
print('Observation space:', env.observation_space)
print('Sample state:', env.format_state(states[0]))
Image.fromarray(env.render(mode='rgb_array'))
# +
# Create the agents
agents = {drone.index: RandomAgent(env) for drone in env.drones}
agents[0] = QLearningAgent(
env, gamma=0.95, alpha=0.1,
epsilon_start=1, epsilon_decay=0.999, epsilon_end=0.01
)
trainer = MultiAgentTrainer(env, agents, reset_agents=True, seed=0)
trainer.train(35000)
plot_rolling_rewards(trainer.rewards_log, events={'pickup': [0.99], 'delivery': [1], 'crash': [-1], 'charging': [-0.1]})
# -
q_table = agents[0].get_qtable()
print('Q-table:', q_table.shape)
q_table.sample(10)
# Don't forget to test with different seeds
rewards_log = test_agents(env, agents, n_steps=1000, seed=0)
plot_cumulative_rewards(
rewards_log,
events={'pickup': [0.99], 'delivery': [1], 'crash': [-1], 'charging': [-0.1]},
drones_labels={0: 'Q-learning'}
)
path = os.path.join('videos', 'ql-compass-lidar-charge.mp4')
render_video(env, agents, video_path=path, n_steps=120, fps=1, seed=0)
ColabVideo(path)
# First tests with deep Q-learning (DQN)
# ---
#
# You can find a DQN implementation in `agents/`
#
# ```
# agents/
# ├── curiosity.py
# ├── dqn.py <-- DQN agent
# ├── logging.py
# ├── qlearning.py <-- Q-learning agent
# └── random.py
# ```
# +
# Create environment
env = LidarCompassChargeQTable(DeliveryDrones())
env.env_params.update({
'n_drones': 3, 'pickup_reward': 0.99, 'delivery_reward': 1
})
states = env.reset()
# Create the agents
agents = {drone.index: RandomAgent(env) for drone in env.drones}
agents[0] = DQNAgent(
env, DenseQNetworkFactory(env, hidden_layers=[256, 256]),
gamma=0.95, epsilon_start=1, epsilon_decay=0.999, epsilon_end=0.01,
memory_size=10000, batch_size=64, target_update_interval=5
)
trainer = MultiAgentTrainer(env, agents, reset_agents=True, seed=0)
agents[0].qnetwork
# -
# Train the agent
trainer.train(25000)
plot_rolling_rewards(
trainer.rewards_log, drones_labels={0: 'DQN'},
events={'pickup': [0.99], 'delivery': [1], 'crash': [-1], 'charging': [-0.1]})
plt.plot(agents[0].epsilons)
plt.xlabel('Number of episodes')
plt.ylabel('Exploration rate (epsilon)')
plt.show()
# Don't forget to test with different seeds!
rewards_log = test_agents(env, agents, n_steps=1000, seed=0)
plot_cumulative_rewards(
rewards_log, drones_labels={0: 'DQN'},
events={'pickup': [0.99], 'delivery': [1], 'crash': [-1], 'charging': [-0.1]})
# Inspect replay memory buffer
agents[0].inspect_memory(top_n=10, max_col=80)
# Take a moment to play with the different parameters: `memory_size`, `batch_size`, `target_update_interval` and the others
path = os.path.join('videos', 'dqn-compass-lidar-charge.mp4')
render_video(env, agents, video_path=path, n_steps=120, fps=1, seed=0)
ColabVideo(path)
# DQN and WindowedGrid (official environment)
# ---
# +
# Create environment
env = WindowedGridView(DeliveryDrones(), radius=3)
env.env_params.update({
'n_drones': 3, 'pickup_reward': 0.99, 'delivery_reward': 1
})
states = env.reset()
# Create the agents
agents = {drone.index: RandomAgent(env) for drone in env.drones}
agents[0] = my_agent = DQNAgent(
env, ConvQNetworkFactory(env, conv_layers=[
{'out_channels': 32, 'kernel_size': 3, 'stride': 1, 'padding': 1},
{'out_channels': 32, 'kernel_size': 3, 'stride': 1, 'padding': 1},
{'out_channels': 32, 'kernel_size': 3, 'stride': 1, 'padding': 1},
{'out_channels': 64, 'kernel_size': 3, 'stride': 1, 'padding': 1},
{'out_channels': 64, 'kernel_size': 3, 'stride': 1, 'padding': 1},
{'out_channels': 64, 'kernel_size': 3, 'stride': 1, 'padding': 1},
], dense_layers=[1024, 256]),
gamma=0.95, epsilon_start=1, epsilon_decay=0.99, epsilon_end=0.01,
memory_size=10000, batch_size=64, target_update_interval=5
)
trainer = MultiAgentTrainer(env, agents, reset_agents=True, seed=0)
agents[0].qnetwork
# -
# Train the agent
for run in range(10):
trainer.train(2500)
plot_rolling_rewards(
trainer.rewards_log, drones_labels={0: 'DQN'},
events={'pickup': [0.99], 'delivery': [1], 'crash': [-1], 'charging': [-0.1]})
path = os.path.join('videos', 'dqn-windowed.mp4')
render_video(env, agents, video_path=path, n_steps=120, fps=1, seed=0)
ColabVideo(path)
# Share your agent q-network
# ---
path = os.path.join('agents', 'dqn-agent.pt')
agents[0].save(path)
# agents[0].load(path) # Later, load the qnetwork!
|
02 Intro to Q-learning and DQN.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Deep Learning with Python
#
# ## 4.5 The universal workflow of machine learning
#
# > 机器学习的通用工作流程
#
# 1. 定义问题,收集数据集:定义问题与要训练的数据。收集这些数据,有需要的话用标签来标注数据。
#
# 2. 选择衡量成功的指标:选择衡量问题成功的指标。你要在验证数据上监控哪些指标?
#
# 3. 确定评估方法:留出验证? K 折验证?你应该将哪一部分数据用于验证?
#
# 4. 准备数据:预处理啦,特征工程啦。。
#
# 6. 开发比基准(比如随机预测)更好的模型,即一个具有统计功效的模型。
#
# 最后一层和损失的选择:
# 
#
# 7. 扩大规模,开发出过拟合的模型:加层、加单元、加轮次
#
# 8. 调节超参数,模型正则化:基于模型在验证数据上的性能来进行模型正则化与调节超参数。
#
#
|
ch4/4.5 The universal workflow of machine learning.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## This notebook handles mapping srprec to census tract, and then adding on census data
# imports
import pandas as pd
import numpy as np
import requests
import pickle as pkl
import matplotlib.pyplot as plt
plt.style.use('ggplot')
# +
# pd.set_option('display.max_columns', None)
# pd.set_option('display.max_rows', None)
# -
# ### Converter file
convert18 = pd.read_pickle('./data/df_convert18.pkl')
convert18.head(3)
convert18.shape
# ### Merge the full dataframes
# #### Steps:
# 1. Attach census data on 'tract', apply weights by 'tract', aggregate to 'srprec'.
# 2. Attach vote data on 'srprec'
# #### Step 1: Attach census data on 'tract', apply weights by 'tract', aggregate to 'srprec'.
# #### Census data set
census18_DP03 = pd.read_pickle('./census_data/DP03_subset.pkl')
census18_DP03.shape
census18_DP03.info()
census18_DP03.drop(['Geographic Area Name', 'GEO_ID'], axis=1, inplace=True)
# +
census18_DP03.head(3)
# -
census18_DP03.shape
# #### Merge census data into convert18 df on 'tract' to create combo df
# pd.merge(left=c, right=b, how='outer', left_on=['tract'], right_on=['tract'], indicator=True, suffixes=('_c', '_b'))
combo = pd.merge(left=convert18, right=census18_DP03, how='outer', left_on=['tract'], right_on=['tract'], indicator=True)
combo.head(10)
combo.shape
# #### Inspect resulting dataframe: 'srprec' with multiple tracts, nulls
#stackoverflow: https://stackoverflow.com/questions/14247586/how-to-select-rows-with-one-or-more-nulls-from-a-pandas-dataframe-without-listin
combo[pd.isnull(combo).any(axis=1)]
combo.loc[combo['srprec_orig'] == 14082]
combo.loc[combo['_merge'] == 'right_only']
combo.drop(index=2011, inplace=True)
combo['pctsrprec_tract'] = round(combo['pctsrprec_tract'], 2)
combo.head(10)
# #### Calculate weighted values for each tract
combo['hh_med_inc_wgt'] = round((combo['pctsrprec_tract']/100 *
combo['Estimate INCOME AND BENEFITS (IN 2018 INFLATION-ADJUSTED DOLLARS) Total households Median household income (dollars)'])
, 2)
combo.head(10)
combo.loc[combo['srprec_orig'] == 75116]
# #### Aggregate results to 'srprec'
srprec_agg = combo.groupby('srprec_orig').sum()
srprec_agg
srprec_agg.reset_index(inplace=True)
srprec_agg.head()
# #### Inspect
srprec_agg.loc[srprec_agg['srprec_orig'] == 75116]
# #### **DROP TRACT DETAIL**
# cols that don't make sense in aggregate:
# subindex
# tract
# test
srprec_census = srprec_agg[['srprec_orig', 'subindex', 'pctsrprec_tract', 'hh_med_inc_wgt']]
srprec_census
srprec_census.info()
# ### Merge in Vote data on 'srprec'
# +
#rename traceability column in advance of second merge
#should be dropped already by .groupby() aggregation step, but a reminder just in case:
# df.rename(columns={'_merge':'prec_merge'}, inplace=True)
# -
# #### Vote dataset
vote18 = pd.read_pickle('./data/trend18.pkl')
vote18.shape
vote18['srprec'] = vote18['srprec'].astype('int64')
vote18.info()
vote18.head(3)
# #### Merge vote data into srprec_census df on 'srprec'
combo = pd.merge(left=srprec_census, right=vote18, how='outer', left_on=['srprec_orig'], right_on=['srprec'], indicator=True)
combo.head()
combo.shape
# #### compare conversion fields
data18[['srprec', 'cddist']].info()
convert18[['srprec', 'tract', 'block']].info()
dp03_18sub[['tract']].info()
|
.ipynb_checkpoints/27_combine_census_data-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # Isotonic Regression
#
#
# An illustration of the isotonic regression on generated data. The
# isotonic regression finds a non-decreasing approximation of a function
# while minimizing the mean squared error on the training data. The benefit
# of such a model is that it does not assume any form for the target
# function such as linearity. For comparison a linear regression is also
# presented.
#
#
#
# +
print(__doc__)
# Author: <NAME> <<EMAIL>>
# <NAME> <<EMAIL>>
# License: BSD
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.collections import LineCollection
from sklearn.linear_model import LinearRegression
from sklearn.isotonic import IsotonicRegression
from sklearn.utils import check_random_state
n = 100
x = np.arange(n)
rs = check_random_state(0)
y = rs.randint(-50, 50, size=(n,)) + 50. * np.log1p(np.arange(n))
# #############################################################################
# Fit IsotonicRegression and LinearRegression models
ir = IsotonicRegression()
y_ = ir.fit_transform(x, y)
lr = LinearRegression()
lr.fit(x[:, np.newaxis], y) # x needs to be 2d for LinearRegression
# #############################################################################
# Plot result
segments = [[[i, y[i]], [i, y_[i]]] for i in range(n)]
lc = LineCollection(segments, zorder=0)
lc.set_array(np.ones(len(y)))
lc.set_linewidths(np.full(n, 0.5))
fig = plt.figure()
plt.plot(x, y, 'r.', markersize=12)
plt.plot(x, y_, 'g.-', markersize=12)
plt.plot(x, lr.predict(x[:, np.newaxis]), 'b-')
plt.gca().add_collection(lc)
plt.legend(('Data', 'Isotonic Fit', 'Linear Fit'), loc='lower right')
plt.title('Isotonic regression')
plt.show()
|
scikit-learn/plot_isotonic_regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Variational Autoencoder in TensorFlow
# The main motivation for this post was that I wanted to get more experience with both [Variational Autoencoders](http://arxiv.org/abs/1312.6114) (VAEs) using [Tensorflow](http://www.tensorflow.org/).
#
# Let us first do the necessary imports, load the data (MNIST), and define some helper functions
# +
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
# %matplotlib inline
np.random.seed(0)
tf.set_random_seed(0)
# +
# Download and read MNIST data.
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import gzip
import os
import tempfile
import numpy
from six.moves import urllib
from six.moves import xrange # pylint: disable=redefined-builtin
import tensorflow as tf
from tensorflow.contrib.learn.python.learn.datasets.mnist import read_data_sets
# -
# Load MNIST data in a format suited for tensorflow.
# The script input_data is available under this URL:
# https://raw.githubusercontent.com/tensorflow/tensorflow/master/tensorflow/examples/tutorials/mnist/input_data.py
# import input_data
mnist = read_data_sets('MNIST_data', one_hot=True)
n_samples = mnist.train.num_examples
We need to initialize the network wi
def xavier_init(fan_in, fan_out, constant=1):
""" Xavier initialization of network weights"""
# https://stackoverflow.com/questions/33640581/how-to-do-xavier-initialization-on-tensorflow
low = -constant*np.sqrt(6.0/(fan_in + fan_out))
high = constant*np.sqrt(6.0/(fan_in + fan_out))
return tf.random_uniform((fan_in, fan_out),
minval=low, maxval=high,
dtype=tf.float32)
# Based on this, we define now a class "VariationalAutoencoder" with a [sklearn](http://scikit-learn.org)-like interface that can be trained incrementally with mini-batches using partial_fit. The trained model can be used to reconstruct unseen input, to generate new samples, and to map inputs to the latent space.
class VariationalAutoencoder(object):
""" Variation Autoencoder (VAE) with an sklearn-like interface implemented using TensorFlow.
This implementation uses probabilistic encoders and decoders using Gaussian
distributions and realized by multi-layer perceptrons. The VAE can be learned
end-to-end.
See "Auto-Encoding Variational Bayes" by <NAME> Welling for more details.
"""
def __init__(self, network_architecture, transfer_fct=tf.nn.softplus,
learning_rate=0.001, batch_size=100):
self.network_architecture = network_architecture
self.transfer_fct = transfer_fct
self.learning_rate = learning_rate
self.batch_size = batch_size
# tf Graph input
self.x = tf.placeholder(tf.float32, [None, network_architecture["n_input"]])
# Create autoencoder network
self._create_network()
# Define loss function based variational upper-bound and
# corresponding optimizer
self._create_loss_optimizer()
# Initializing the tensor flow variables
init = tf.global_variables_initializer()
# Launch the session
self.sess = tf.InteractiveSession()
self.sess.run(init)
def _create_network(self):
# Initialize autoencode network weights and biases
network_weights = self._initialize_weights(**self.network_architecture)
# Use recognition network to determine mean and
# (log) variance of Gaussian distribution in latent
# space
self.z_mean, self.z_log_sigma_sq = \
self._recognition_network(network_weights["weights_recog"],
network_weights["biases_recog"])
# Draw one sample z from Gaussian distribution
n_z = self.network_architecture["n_z"]
eps = tf.random_normal((self.batch_size, n_z), 0, 1,
dtype=tf.float32)
# z = mu + sigma*epsilon
self.z = tf.add(self.z_mean,
tf.multiply(tf.sqrt(tf.exp(self.z_log_sigma_sq)), eps))
# Use generator to determine mean of
# Bernoulli distribution of reconstructed input
self.x_reconstr_mean = \
self._generator_network(network_weights["weights_gener"],
network_weights["biases_gener"])
def _initialize_weights(self, n_hidden_recog_1, n_hidden_recog_2,
n_hidden_gener_1, n_hidden_gener_2,
n_input, n_z):
all_weights = dict()
all_weights['weights_recog'] = {
'h1': tf.Variable(xavier_init(n_input, n_hidden_recog_1)),
'h2': tf.Variable(xavier_init(n_hidden_recog_1, n_hidden_recog_2)),
'out_mean': tf.Variable(xavier_init(n_hidden_recog_2, n_z)),
'out_log_sigma': tf.Variable(xavier_init(n_hidden_recog_2, n_z))}
all_weights['biases_recog'] = {
'b1': tf.Variable(tf.zeros([n_hidden_recog_1], dtype=tf.float32)),
'b2': tf.Variable(tf.zeros([n_hidden_recog_2], dtype=tf.float32)),
'out_mean': tf.Variable(tf.zeros([n_z], dtype=tf.float32)),
'out_log_sigma': tf.Variable(tf.zeros([n_z], dtype=tf.float32))}
all_weights['weights_gener'] = {
'h1': tf.Variable(xavier_init(n_z, n_hidden_gener_1)),
'h2': tf.Variable(xavier_init(n_hidden_gener_1, n_hidden_gener_2)),
'out_mean': tf.Variable(xavier_init(n_hidden_gener_2, n_input)),
'out_log_sigma': tf.Variable(xavier_init(n_hidden_gener_2, n_input))}
all_weights['biases_gener'] = {
'b1': tf.Variable(tf.zeros([n_hidden_gener_1], dtype=tf.float32)),
'b2': tf.Variable(tf.zeros([n_hidden_gener_2], dtype=tf.float32)),
'out_mean': tf.Variable(tf.zeros([n_input], dtype=tf.float32)),
'out_log_sigma': tf.Variable(tf.zeros([n_input], dtype=tf.float32))}
return all_weights
def _recognition_network(self, weights, biases):
# Generate probabilistic encoder (recognition network), which
# maps inputs onto a normal distribution in latent space.
# The transformation is parametrized and can be learned.
layer_1 = self.transfer_fct(tf.add(tf.matmul(self.x, weights['h1']),
biases['b1']))
layer_2 = self.transfer_fct(tf.add(tf.matmul(layer_1, weights['h2']),
biases['b2']))
z_mean = tf.add(tf.matmul(layer_2, weights['out_mean']),
biases['out_mean'])
z_log_sigma_sq = \
tf.add(tf.matmul(layer_2, weights['out_log_sigma']),
biases['out_log_sigma'])
return (z_mean, z_log_sigma_sq)
def _generator_network(self, weights, biases):
# Generate probabilistic decoder (decoder network), which
# maps points in latent space onto a Bernoulli distribution in data space.
# The transformation is parametrized and can be learned.
layer_1 = self.transfer_fct(tf.add(tf.matmul(self.z, weights['h1']),
biases['b1']))
layer_2 = self.transfer_fct(tf.add(tf.matmul(layer_1, weights['h2']),
biases['b2']))
x_reconstr_mean = \
tf.nn.sigmoid(tf.add(tf.matmul(layer_2, weights['out_mean']),
biases['out_mean']))
return x_reconstr_mean
def _create_loss_optimizer(self):
# The loss is composed of two terms:
# 1.) The reconstruction loss (the negative log probability
# of the input under the reconstructed Bernoulli distribution
# induced by the decoder in the data space).
# This can be interpreted as the number of "nats" required
# for reconstructing the input when the activation in latent
# is given.
# Adding 1e-10 to avoid evaluation of log(0.0)
reconstr_loss = \
-tf.reduce_sum(self.x * tf.log(1e-10 + self.x_reconstr_mean)
+ (1-self.x) * tf.log(1e-10 + 1 - self.x_reconstr_mean),
1)
# 2.) The latent loss, which is defined as the Kullback Leibler divergence
## between the distribution in latent space induced by the encoder on
# the data and some prior. This acts as a kind of regularizer.
# This can be interpreted as the number of "nats" required
# for transmitting the the latent space distribution given
# the prior.
latent_loss = -0.5 * tf.reduce_sum(1 + self.z_log_sigma_sq
- tf.square(self.z_mean)
- tf.exp(self.z_log_sigma_sq), 1)
self.cost = tf.reduce_mean(reconstr_loss + latent_loss) # average over batch
# Use ADAM optimizer
self.optimizer = \
tf.train.AdamOptimizer(learning_rate=self.learning_rate).minimize(self.cost)
def partial_fit(self, X):
"""Train model based on mini-batch of input data.
Return cost of mini-batch.
"""
opt, cost = self.sess.run((self.optimizer, self.cost),
feed_dict={self.x: X})
return cost
def transform(self, X):
"""Transform data by mapping it into the latent space."""
# Note: This maps to mean of distribution, we could alternatively
# sample from Gaussian distribution
return self.sess.run(self.z_mean, feed_dict={self.x: X})
def generate(self, z_mu=None):
""" Generate data by sampling from latent space.
If z_mu is not None, data for this point in latent space is
generated. Otherwise, z_mu is drawn from prior in latent
space.
"""
if z_mu is None:
z_mu = np.random.normal(size=self.network_architecture["n_z"])
# Note: This maps to mean of distribution, we could alternatively
# sample from Gaussian distribution
return self.sess.run(self.x_reconstr_mean,
feed_dict={self.z: z_mu})
def reconstruct(self, X):
""" Use VAE to reconstruct given data. """
return self.sess.run(self.x_reconstr_mean,
feed_dict={self.x: X})
# In general, implementing a VAE in tensorflow is relatively straightforward (in particular since we don not need to code the gradient computation). A bit confusing is potentially that all the logic happens at initialization of the class (where the graph is generated), while the actual sklearn interface methods are very simple one-liners.
#
# We can now define a simple fuction which trains the VAE using mini-batches:
def train(network_architecture, learning_rate=0.001,
batch_size=100, training_epochs=10, display_step=5):
vae = VariationalAutoencoder(network_architecture,
learning_rate=learning_rate,
batch_size=batch_size)
# Training cycle
for epoch in range(training_epochs):
avg_cost = 0.
total_batch = int(n_samples / batch_size)
# Loop over all batches
for i in range(total_batch):
batch_xs, _ = mnist.train.next_batch(batch_size)
# Fit training using batch data
cost = vae.partial_fit(batch_xs)
# Compute average loss
avg_cost += cost / n_samples * batch_size
# Display logs per epoch step
if epoch % display_step == 0:
print("Epoch:", '%04d' % (epoch+1),
"cost=", "{:.9f}".format(avg_cost))
return vae
# ## Illustrating reconstruction quality
# We can now train a VAE on MNIST by just specifying the network topology. We start with training a VAE with a 20-dimensional latent space.
# +
network_architecture = \
dict(n_hidden_recog_1=500, # 1st layer encoder neurons
n_hidden_recog_2=500, # 2nd layer encoder neurons
n_hidden_gener_1=500, # 1st layer decoder neurons
n_hidden_gener_2=500, # 2nd layer decoder neurons
n_input=784, # MNIST data input (img shape: 28*28)
n_z=20) # dimensionality of latent space
vae = train(network_architecture, training_epochs=75)
# -
# Based on this we can sample some test inputs and visualize how well the VAE can reconstruct those. In general the VAE does really well.
# +
x_sample = mnist.test.next_batch(100)[0]
x_reconstruct = vae.reconstruct(x_sample)
plt.figure(figsize=(8, 12))
for i in range(5):
plt.subplot(5, 2, 2*i + 1)
plt.imshow(x_sample[i].reshape(28, 28), vmin=0, vmax=1, cmap="gray")
plt.title("Test input")
plt.colorbar()
plt.subplot(5, 2, 2*i + 2)
plt.imshow(x_reconstruct[i].reshape(28, 28), vmin=0, vmax=1, cmap="gray")
plt.title("Reconstruction")
plt.colorbar()
plt.tight_layout()
# -
# ## Illustrating latent space
# Next, we train a VAE with 2d latent space and illustrates how the encoder (the recognition network) encodes some of the labeled inputs (collapsing the Gaussian distribution in latent space to its mean). This gives us some insights into the structure of the learned manifold (latent space)
# +
network_architecture = \
dict(n_hidden_recog_1=500, # 1st layer encoder neurons
n_hidden_recog_2=500, # 2nd layer encoder neurons
n_hidden_gener_1=500, # 1st layer decoder neurons
n_hidden_gener_2=500, # 2nd layer decoder neurons
n_input=784, # MNIST data input (img shape: 28*28)
n_z=2) # dimensionality of latent space
vae_2d = train(network_architecture, training_epochs=75)
# -
x_sample, y_sample = mnist.test.next_batch(5000)
z_mu = vae_2d.transform(x_sample)
plt.figure(figsize=(8, 6))
plt.scatter(z_mu[:, 0], z_mu[:, 1], c=np.argmax(y_sample, 1))
plt.colorbar()
plt.grid()
# An other way of getting insights into the latent space is to use the generator network to plot reconstructions at the positions in the latent space for which they have been generated:
# +
nx = ny = 20
x_values = np.linspace(-3, 3, nx)
y_values = np.linspace(-3, 3, ny)
canvas = np.empty((28*ny, 28*nx))
for i, yi in enumerate(x_values):
for j, xi in enumerate(y_values):
z_mu = np.array([[xi, yi]]*vae.batch_size)
x_mean = vae_2d.generate(z_mu)
canvas[(nx-i-1)*28:(nx-i)*28, j*28:(j+1)*28] = x_mean[0].reshape(28, 28)
plt.figure(figsize=(8, 10))
Xi, Yi = np.meshgrid(x_values, y_values)
plt.imshow(canvas, origin="upper", cmap="gray")
plt.tight_layout()
# -
# ## Summary
# In summary, tensorflow is well suited to rapidly implement a prototype of machine learning models like VAE. The resulting code could be easily executed on GPUs as well (requiring just that tensorflow with GPU support was installed). VAE allows learning probabilistic encoders and decoders of data in an end-to-end fashion.
# %load_ext watermark
# %watermark -a "RKP" -d -v -m -p numpy,sklearn,tensorflow
|
.ipynb_checkpoints/vae-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Modeling and Simulation in Python
#
# Case study.
#
# Copyright 2017 <NAME>
#
# License: [Creative Commons Attribution 4.0 International](https://creativecommons.org/licenses/by/4.0)
#
# +
# Configure Jupyter so figures appear in the notebook
# %matplotlib inline
# Configure Jupyter to display the assigned value after an assignment
# %config InteractiveShell.ast_node_interactivity='last_expr_or_assign'
# import functions from the modsim.py module
from modsim import *
# -
# ### Unrolling
#
# Let's simulate a kitten unrolling toilet paper. As reference material, see [this video](http://modsimpy.com/kitten).
#
# The interactions of the kitten and the paper roll are complex. To keep things simple, let's assume that the kitten pulls down on the free end of the roll with constant force. Also, we will neglect the friction between the roll and the axle.
#
# 
#
# This figure shows the paper roll with $r$, $F$, and $\tau$. As a vector quantity, the direction of $\tau$ is into the page, but we only care about its magnitude for now.
# We'll start by loading the units we need.
radian = UNITS.radian
m = UNITS.meter
s = UNITS.second
kg = UNITS.kilogram
N = UNITS.newton
# And a few more parameters in the `Params` object.
params = Params(Rmin = 0.02 * m,
Rmax = 0.055 * m,
Mcore = 15e-3 * kg,
Mroll = 215e-3 * kg,
L = 47 * m,
tension = 2e-4 * N,
t_end = 120 * s)
# `make_system` computes `rho_h`, which we'll need to compute moment of inertia, and `k`, which we'll use to compute `r`.
def make_system(params):
"""Make a system object.
params: Params with Rmin, Rmax, Mcore, Mroll,
L, tension, and t_end
returns: System with init, k, rho_h, Rmin, Rmax,
Mcore, Mroll, ts
"""
unpack(params)
init = State(theta = 0 * radian,
omega = 0 * radian/s,
y = L)
area = pi * (Rmax**2 - Rmin**2)
rho_h = Mroll / area
k = (Rmax**2 - Rmin**2) / 2 / L / radian
return System(init=init, k=k, rho_h=rho_h,
Rmin=Rmin, Rmax=Rmax,
Mcore=Mcore, Mroll=Mroll,
t_end=t_end)
# Testing `make_system`
system = make_system(params)
system.init
# Here's how we compute `I` as a function of `r`:
def moment_of_inertia(r, system):
"""Moment of inertia for a roll of toilet paper.
r: current radius of roll in meters
system: System object with Mcore, rho, Rmin, Rmax
returns: moment of inertia in kg m**2
"""
unpack(system)
Icore = Mcore * Rmin**2
Iroll = pi * rho_h / 2 * (r**4 - Rmin**4)
return Icore + Iroll
# When `r` is `Rmin`, `I` is small.
moment_of_inertia(system.Rmin, system)
# As `r` increases, so does `I`.
moment_of_inertia(system.Rmax, system)
# ## Exercises
#
# Write a slope function we can use to simulate this system. Here are some suggestions and hints:
#
# * `r` is no longer part of the `State` object. Instead, we compute `r` at each time step, based on the current value of `y`, using
#
# $y = \frac{1}{2k} (r^2 - R_{min}^2)$
#
# * Angular velocity, `omega`, is no longer constant. Instead, we compute torque, `tau`, and angular acceleration, `alpha`, at each time step.
#
# * I changed the definition of `theta` so positive values correspond to clockwise rotation, so `dydt = -r * omega`; that is, positive values of `omega` yield decreasing values of `y`, the amount of paper still on the roll.
#
# * Your slope function should return `omega`, `alpha`, and `dydt`, which are the derivatives of `theta`, `omega`, and `y`, respectively.
#
# * Because `r` changes over time, we have to compute moment of inertia, `I`, at each time step.
#
# That last point might be more of a problem than I have made it seem. In the same way that $F = m a$ only applies when $m$ is constant, $\tau = I \alpha$ only applies when $I$ is constant. When $I$ varies, we usually have to use a more general version of Newton's law. However, I believe that in this example, mass and moment of inertia vary together in a way that makes the simple approach work out. Not all of my collegues are convinced.
# +
# Solution goes here
# -
# Test `slope_func` with the initial conditions.
# +
# Solution goes here
# -
# Run the simulation.
# +
# Solution goes here
# -
# And look at the results.
results.tail()
# Check the results to see if they seem plausible:
#
# * The final value of `theta` should be about 220 radians.
#
# * The final value of `omega` should be near 4 radians/second, which is less one revolution per second, so that seems plausible.
#
# * The final value of `y` should be about 35 meters of paper left on the roll, which means the kitten pulls off 12 meters in two minutes. That doesn't seem impossible, although it is based on a level of consistency and focus that is unlikely in a kitten.
#
# * Angular velocity, `omega`, should increase almost linearly at first, as constant force yields almost constant torque. Then, as the radius decreases, the lever arm decreases, yielding lower torque, but moment of inertia decreases even more, yielding higher angular acceleration.
# Plot `theta`
# +
def plot_theta(results):
plot(results.theta, color='C0', label='theta')
decorate(xlabel='Time (s)',
ylabel='Angle (rad)')
plot_theta(results)
# -
# Plot `omega`
# +
def plot_omega(results):
plot(results.omega, color='C2', label='omega')
decorate(xlabel='Time (s)',
ylabel='Angular velocity (rad/s)')
plot_omega(results)
# -
# Plot `y`
# +
def plot_y(results):
plot(results.y, color='C1', label='y')
decorate(xlabel='Time (s)',
ylabel='Length (m)')
plot_y(results)
|
code/kitten.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# version 1.0.1
# # + 
# # **Web Server Log Analysis with Apache Spark**
#
# ####This lab will demonstrate how easy it is to perform web server log analysis with Apache Spark.
#
# ####Server log analysis is an ideal use case for Spark. It's a very large, common data source and contains a rich set of information. Spark allows you to store your logs in files on disk cheaply, while still providing a quick and simple way to perform data analysis on them. This homework will show you how to use Apache Spark on real-world text-based production logs and fully harness the power of that data. Log data comes from many sources, such as web, file, and compute servers, application logs, user-generated content, and can be used for monitoring servers, improving business and customer intelligence, building recommendation systems, fraud detection, and much more.
# ### How to complete this assignment
#
# ####This assignment is broken up into sections with bite-sized examples for demonstrating Spark functionality for log processing. For each problem, you should start by thinking about the algorithm that you will use to *efficiently* process the log in a parallel, distributed manner. This means using the various [RDD](http://spark.apache.org/docs/latest/api/python/pyspark.html#pyspark.RDD) operations along with [`lambda` functions](https://docs.python.org/2/tutorial/controlflow.html#lambda-expressions) that are applied at each worker.
#
# ####This assignment consists of 4 parts:
# #### *Part 1*: Apache Web Server Log file format
# #### *Part 2*: Sample Analyses on the Web Server Log File
# #### *Part 3*: Analyzing Web Server Log File
# #### *Part 4*: Exploring 404 Response Codes
# ### **Part 1: Apache Web Server Log file format**
# ####The log files that we use for this assignment are in the [Apache Common Log Format (CLF)](http://httpd.apache.org/docs/1.3/logs.html#common). The log file entries produced in CLF will look something like this:
# `127.0.0.1 - - [01/Aug/1995:00:00:01 -0400] "GET /images/launch-logo.gif HTTP/1.0" 200 1839`
#
# ####Each part of this log entry is described below.
# * `127.0.0.1`
# ####This is the IP address (or host name, if available) of the client (remote host) which made the request to the server.
#
# * `-`
# ####The "hyphen" in the output indicates that the requested piece of information (user identity from remote machine) is not available.
#
# * `-`
# ####The "hyphen" in the output indicates that the requested piece of information (user identity from local logon) is not available.
#
# * `[01/Aug/1995:00:00:01 -0400]`
# ####The time that the server finished processing the request. The format is:
# `[day/month/year:hour:minute:second timezone]`
# * ####day = 2 digits
# * ####month = 3 letters
# * ####year = 4 digits
# * ####hour = 2 digits
# * ####minute = 2 digits
# * ####second = 2 digits
# * ####zone = (\+ | \-) 4 digits
#
# * `"GET /images/launch-logo.gif HTTP/1.0"`
# ####This is the first line of the request string from the client. It consists of a three components: the request method (e.g., `GET`, `POST`, etc.), the endpoint (a [Uniform Resource Identifier](http://en.wikipedia.org/wiki/Uniform_resource_identifier)), and the client protocol version.
#
# * `200`
# ####This is the status code that the server sends back to the client. This information is very valuable, because it reveals whether the request resulted in a successful response (codes beginning in 2), a redirection (codes beginning in 3), an error caused by the client (codes beginning in 4), or an error in the server (codes beginning in 5). The full list of possible status codes can be found in the HTTP specification ([RFC 2616](https://www.ietf.org/rfc/rfc2616.txt) section 10).
#
# * `1839`
# ####The last entry indicates the size of the object returned to the client, not including the response headers. If no content was returned to the client, this value will be "-" (or sometimes 0).
#
# ####Note that log files contain information supplied directly by the client, without escaping. Therefore, it is possible for malicious clients to insert control-characters in the log files, *so care must be taken in dealing with raw logs.*
#
# ### NASA-HTTP Web Server Log
# ####For this assignment, we will use a data set from NASA Kennedy Space Center WWW server in Florida. The full data set is freely available (http://ita.ee.lbl.gov/html/contrib/NASA-HTTP.html) and contains two month's of all HTTP requests. We are using a subset that only contains several days worth of requests.
# ### **(1a) Parsing Each Log Line**
# ####Using the CLF as defined above, we create a regular expression pattern to extract the nine fields of the log line using the Python regular expression [`search` function](https://docs.python.org/2/library/re.html#regular-expression-objects). The function returns a pair consisting of a Row object and 1. If the log line fails to match the regular expression, the function returns a pair consisting of the log line string and 0. A '-' value in the content size field is cleaned up by substituting it with 0. The function converts the log line's date string into a Python `datetime` object using the given `parse_apache_time` function.
# +
import re
import datetime
from pyspark.sql import Row
month_map = {'Jan': 1, 'Feb': 2, 'Mar':3, 'Apr':4, 'May':5, 'Jun':6, 'Jul':7,
'Aug':8, 'Sep': 9, 'Oct':10, 'Nov': 11, 'Dec': 12}
def parse_apache_time(s):
""" Convert Apache time format into a Python datetime object
Args:
s (str): date and time in Apache time format
Returns:
datetime: datetime object (ignore timezone for now)
"""
return datetime.datetime(int(s[7:11]),
month_map[s[3:6]],
int(s[0:2]),
int(s[12:14]),
int(s[15:17]),
int(s[18:20]))
def parseApacheLogLine(logline):
""" Parse a line in the Apache Common Log format
Args:
logline (str): a line of text in the Apache Common Log format
Returns:
tuple: either a dictionary containing the parts of the Apache Access Log and 1,
or the original invalid log line and 0
"""
match = re.search(APACHE_ACCESS_LOG_PATTERN, logline)
if match is None:
return (logline, 0)
size_field = match.group(9)
if size_field == '-':
size = long(0)
else:
size = long(match.group(9))
return (Row(
host = match.group(1),
client_identd = match.group(2),
user_id = match.group(3),
date_time = parse_apache_time(match.group(4)),
method = match.group(5),
endpoint = match.group(6),
protocol = match.group(7),
response_code = int(match.group(8)),
content_size = size
), 1)
# -
# A regular expression pattern to extract fields from the log line
APACHE_ACCESS_LOG_PATTERN = '^(\S+) (\S+) (\S+) \[([\w:/]+\s[+\-]\d{4})\] "(\S+) (\S+)\s*(\S*)\s*" (\d{3}) (\S+)'
# ### **(1b) Configuration and Initial RDD Creation**
# ####We are ready to specify the input log file and create an RDD containing the parsed log file data. The log file has already been downloaded for you.
#
# ####To create the primary RDD that we'll use in the rest of this assignment, we first load the text file using [`sc.textfile(logFile)`](http://spark.apache.org/docs/latest/api/python/pyspark.html#pyspark.SparkContext.textFile) to convert each line of the file into an element in an RDD.
# ####Next, we use [`map(parseApacheLogLine)`](http://spark.apache.org/docs/latest/api/python/pyspark.html#pyspark.RDD.map) to apply the parse function to each element (that is, a line from the log file) in the RDD and turn each line into a pair [`Row` object](http://spark.apache.org/docs/latest/api/python/pyspark.sql.html#pyspark.sql.Row).
# ####Finally, we cache the RDD in memory since we'll use it throughout this notebook.
# +
import sys
import os
from test_helper import Test
baseDir = os.path.join('data')
inputPath = os.path.join('cs100', 'lab2', 'apache.access.log.PROJECT')
logFile = os.path.join(baseDir, inputPath)
def parseLogs():
""" Read and parse log file """
parsed_logs = (sc
.textFile(logFile)
.map(parseApacheLogLine)
.cache())
access_logs = (parsed_logs
.filter(lambda s: s[1] == 1)
.map(lambda s: s[0])
.cache())
failed_logs = (parsed_logs
.filter(lambda s: s[1] == 0)
.map(lambda s: s[0]))
failed_logs_count = failed_logs.count()
if failed_logs_count > 0:
print 'Number of invalid logline: %d' % failed_logs.count()
for line in failed_logs.take(20):
print 'Invalid logline: %s' % line
print 'Read %d lines, successfully parsed %d lines, failed to parse %d lines' % (parsed_logs.count(), access_logs.count(), failed_logs.count())
return parsed_logs, access_logs, failed_logs
parsed_logs, access_logs, failed_logs = parseLogs()
# -
# ### **(1c) Data Cleaning**
# #### Notice that there are a large number of log lines that failed to parse. Examine the sample of invalid lines and compare them to the correctly parsed line, an example is included below. Based on your observations, alter the `APACHE_ACCESS_LOG_PATTERN` regular expression below so that the failed lines will correctly parse, and press `Shift-Enter` to rerun `parseLogs()`.
#
# `127.0.0.1 - - [01/Aug/1995:00:00:01 -0400] "GET /images/launch-logo.gif HTTP/1.0" 200 1839`
#
# #### If you not familar with Python regular expression [`search` function](https://docs.python.org/2/library/re.html#regular-expression-objects), now would be a good time to check up on the [documentation](https://developers.google.com/edu/python/regular-expressions). One tip that might be useful is to use an online tester like http://pythex.org or http://www.pythonregex.com. To use it, copy and paste the regular expression string below (located between the single quotes ') and test it against one of the 'Invalid logline' above.
# +
# TODO: Replace <FILL IN> with appropriate code
# This was originally '^(\S+) (\S+) (\S+) \[([\w:/]+\s[+\-]\d{4})\] "(\S+) (\S+)\s*(\S*)" (\d{3}) (\S+)'
APACHE_ACCESS_LOG_PATTERN = '^(\S+) (\S+) (\S+) \[([\w:/]+\s[+\-]\d{4})\] "(\S+) (\S+)\s*(\S*)\s*" (\d{3}) (\S+)'
parsed_logs, access_logs, failed_logs = parseLogs()
# -
# TEST Data cleaning (1c)
Test.assertEquals(failed_logs.count(), 0, 'incorrect failed_logs.count()')
Test.assertEquals(parsed_logs.count(), 1043177 , 'incorrect parsed_logs.count()')
Test.assertEquals(access_logs.count(), parsed_logs.count(), 'incorrect access_logs.count()')
# ### **Part 2: Sample Analyses on the Web Server Log File**
#
# ####Now that we have an RDD containing the log file as a set of Row objects, we can perform various analyses.
#
# #### **(2a) Example: Content Size Statistics**
#
# ####Let's compute some statistics about the sizes of content being returned by the web server. In particular, we'd like to know what are the average, minimum, and maximum content sizes.
#
# ####We can compute the statistics by applying a `map` to the `access_logs` RDD. The `lambda` function we want for the map is to extract the `content_size` field from the RDD. The map produces a new RDD containing only the `content_sizes` (one element for each Row object in the `access_logs` RDD). To compute the minimum and maximum statistics, we can use [`min()`](http://spark.apache.org/docs/latest/api/python/pyspark.html#pyspark.RDD.min) and [`max()`](http://spark.apache.org/docs/latest/api/python/pyspark.html#pyspark.RDD.max) functions on the new RDD. We can compute the average statistic by using the [`reduce`](http://spark.apache.org/docs/latest/api/python/pyspark.html#pyspark.RDD.reduce) function with a `lambda` function that sums the two inputs, which represent two elements from the new RDD that are being reduced together. The result of the `reduce()` is the total content size from the log and it is to be divided by the number of requests as determined using the [`count()`](http://spark.apache.org/docs/latest/api/python/pyspark.html#pyspark.RDD.count) function on the new RDD.
# Calculate statistics based on the content size.
content_sizes = access_logs.map(lambda log: log.content_size).cache()
print 'Content Size Avg: %i, Min: %i, Max: %s' % (
content_sizes.reduce(lambda a, b : a + b) / content_sizes.count(),
content_sizes.min(),
content_sizes.max())
# #### **(2b) Example: Response Code Analysis**
# ####Next, lets look at the response codes that appear in the log. As with the content size analysis, first we create a new RDD by using a `lambda` function to extract the `response_code` field from the `access_logs` RDD. The difference here is that we will use a [pair tuple](https://docs.python.org/2/tutorial/datastructures.html?highlight=tuple#tuples-and-sequences) instead of just the field itself. Using a pair tuple consisting of the response code and 1 will let us count how many records have a particular response code. Using the new RDD, we perform a [`reduceByKey`](http://spark.apache.org/docs/latest/api/python/pyspark.html#pyspark.RDD.reduceByKey) function. `reduceByKey` performs a reduce on a per-key basis by applying the `lambda` function to each element, pairwise with the same key. We use the simple `lambda` function of adding the two values. Then, we cache the resulting RDD and create a list by using the [`take`](http://spark.apache.org/docs/latest/api/python/pyspark.html#pyspark.RDD.take) function.
# Response Code to Count
responseCodeToCount = (access_logs
.map(lambda log: (log.response_code, 1))
.reduceByKey(lambda a, b : a + b)
.cache())
responseCodeToCountList = responseCodeToCount.take(100)
print 'Found %d response codes' % len(responseCodeToCountList)
print 'Response Code Counts: %s' % responseCodeToCountList
assert len(responseCodeToCountList) == 7
assert sorted(responseCodeToCountList) == [(200, 940847), (302, 16244), (304, 79824), (403, 58), (404, 6185), (500, 2), (501, 17)]
# #### **(2c) Example: Response Code Graphing with `matplotlib`**
# ####Now, lets visualize the results from the last example. We can visualize the results from the last example using [`matplotlib`](http://matplotlib.org/). First we need to extract the labels and fractions for the graph. We do this with two separate `map` functions with a `lambda` functions. The first `map` function extracts a list of of the response code values, and the second `map` function extracts a list of the per response code counts divided by the total size of the access logs. Next, we create a figure with `figure()` constructor and use the `pie()` method to create the pie plot.
labels = responseCodeToCount.map(lambda (x, y): x).collect()
print labels
count = access_logs.count()
fracs = responseCodeToCount.map(lambda (x, y): (float(y) / count)).collect()
print fracs
# +
import matplotlib.pyplot as plt
def pie_pct_format(value):
""" Determine the appropriate format string for the pie chart percentage label
Args:
value: value of the pie slice
Returns:
str: formated string label; if the slice is too small to fit, returns an empty string for label
"""
return '' if value < 7 else '%.0f%%' % value
fig = plt.figure(figsize=(4.5, 4.5), facecolor='white', edgecolor='white')
colors = ['yellowgreen', 'lightskyblue', 'gold', 'purple', 'lightcoral', 'yellow', 'black']
explode = (0.05, 0.05, 0.1, 0, 0, 0, 0)
patches, texts, autotexts = plt.pie(fracs, labels=labels, colors=colors,
explode=explode, autopct=pie_pct_format,
shadow=False, startangle=125)
for text, autotext in zip(texts, autotexts):
if autotext.get_text() == '':
text.set_text('') # If the slice is small to fit, don't show a text label
plt.legend(labels, loc=(0.80, -0.1), shadow=True)
pass
# -
# #### **(2d) Example: Frequent Hosts**
# ####Let's look at hosts that have accessed the server multiple times (e.g., more than ten times). As with the response code analysis in (2b), first we create a new RDD by using a `lambda` function to extract the `host` field from the `access_logs` RDD using a pair tuple consisting of the host and 1 which will let us count how many records were created by a particular host's request. Using the new RDD, we perform a `reduceByKey` function with a `lambda` function that adds the two values. We then filter the result based on the count of accesses by each host (the second element of each pair) being greater than ten. Next, we extract the host name by performing a `map` with a `lambda` function that returns the first element of each pair. Finally, we extract 20 elements from the resulting RDD - *note that the choice of which elements are returned is not guaranteed to be deterministic.*
# +
# Any hosts that has accessed the server more than 10 times.
hostCountPairTuple = access_logs.map(lambda log: (log.host, 1))
hostSum = hostCountPairTuple.reduceByKey(lambda a, b : a + b)
hostMoreThan10 = hostSum.filter(lambda s: s[1] > 10)
hostsPick20 = (hostMoreThan10
.map(lambda s: s[0])
.take(20))
print 'Any 20 hosts that have accessed more then 10 times: %s' % hostsPick20
# An example: [u'192.168.3.11', u'172.16.31.10', u'slip1-32.acs.ohio-state.edu', u'lapdog-14.baylor.edu', u'172.16.58.3', u'gs1.cs.ttu.edu', u'haskell.limbex.com', u'alfred.uib.no', u'172.16.58.3', u'manaus.bologna.maraut.it', u'dialup98-110.swipnet.se', u'slip-ppp02.feldspar.com', u'ad03-053.compuserve.com', u'srawlin.opsys.nwa.com', u'172.16.31.10', u'ix-den7-23.ix.netcom.com', u'192.168.127.12', u'w20-575-104.mit.edu', u'192.168.3.11', u'ns.rmc.com']
# -
# #### **(2e) Example: Visualizing Endpoints**
# ####Now, lets visualize the number of hits to endpoints (URIs) in the log. To perform this task, we first create a new RDD by using a `lambda` function to extract the `endpoint` field from the `access_logs` RDD using a pair tuple consisting of the endpoint and 1 which will let us count how many records were created by a particular host's request. Using the new RDD, we perform a `reduceByKey` function with a `lambda` function that adds the two values. We then cache the results.
#
# ####Next we visualize the results using `matplotlib`. We previously imported the `matplotlib.pyplot` library, so we do not need to import it again. We perform two separate `map` functions with `lambda` functions. The first `map` function extracts a list of endpoint values, and the second `map` function extracts a list of the visits per endpoint values. Next, we create a figure with `figure()` constructor, set various features of the plot (axis limits, grid lines, and labels), and use the `plot()` method to create the line plot.
# +
endpoints = (access_logs
.map(lambda log: (log.endpoint, 1))
.reduceByKey(lambda a, b : a + b)
.cache())
ends = endpoints.map(lambda (x, y): x).collect()
counts = endpoints.map(lambda (x, y): y).collect()
fig = plt.figure(figsize=(8,4.2), facecolor='white', edgecolor='white')
plt.axis([0, len(ends), 0, max(counts)])
plt.grid(b=True, which='major', axis='y')
plt.xlabel('Endpoints')
plt.ylabel('Number of Hits')
plt.plot(counts)
pass
# -
# #### **(2f) Example: Top Endpoints**
# ####For the final example, we'll look at the top endpoints (URIs) in the log. To determine them, we first create a new RDD by using a `lambda` function to extract the `endpoint` field from the `access_logs` RDD using a pair tuple consisting of the endpoint and 1 which will let us count how many records were created by a particular host's request. Using the new RDD, we perform a `reduceByKey` function with a `lambda` function that adds the two values. We then extract the top ten endpoints by performing a [`takeOrdered`](http://spark.apache.org/docs/latest/api/python/pyspark.html#pyspark.RDD.takeOrdered) with a value of 10 and a `lambda` function that multiplies the count (the second element of each pair) by -1 to create a sorted list with the top endpoints at the bottom.
# +
# Top Endpoints
endpointCounts = (access_logs
.map(lambda log: (log.endpoint, 1))
.reduceByKey(lambda a, b : a + b))
topEndpoints = endpointCounts.takeOrdered(10, lambda s: -1 * s[1])
print 'Top Ten Endpoints: %s' % topEndpoints
assert topEndpoints == [(u'/images/NASA-logosmall.gif', 59737), (u'/images/KSC-logosmall.gif', 50452), (u'/images/MOSAIC-logosmall.gif', 43890), (u'/images/USA-logosmall.gif', 43664), (u'/images/WORLD-logosmall.gif', 43277), (u'/images/ksclogo-medium.gif', 41336), (u'/ksc.html', 28582), (u'/history/apollo/images/apollo-logo1.gif', 26778), (u'/images/launch-logo.gif', 24755), (u'/', 20292)], 'incorrect Top Ten Endpoints'
# -
# ### **Part 3: Analyzing Web Server Log File**
#
# ####Now it is your turn to perform analyses on web server log files.
# #### **(3a) Exercise: Top Ten Error Endpoints**
# ####What are the top ten endpoints which did not have return code 200? Create a sorted list containing top ten endpoints and the number of times that they were accessed with non-200 return code.
#
# ####Think about the steps that you need to perform to determine which endpoints did not have a 200 return code, how you will uniquely count those endpoints, and sort the list.
#
# ####You might want to refer back to the previous Lab (Lab 1 Word Count) for insights.
# +
# TODO: Replace <FILL IN> with appropriate code
# HINT: Each of these <FILL IN> below could be completed with a single transformation or action.
# You are welcome to structure your solution in a different way, so long as
# you ensure the variables used in the next Test section are defined (ie. endpointSum, topTenErrURLs).
not200 = access_logs.filter(lambda log: log.response_code != 200)
endpointCountPairTuple = not200.map(lambda log: (log.endpoint, 1))
endpointSum = endpointCountPairTuple.reduceByKey(lambda a, b : a + b)
topTenErrURLs = endpointSum.takeOrdered(10, lambda s: -1 * s[1])
print 'Top Ten failed URLs: %s' % topTenErrURLs
# -
# TEST Top ten error endpoints (3a)
Test.assertEquals(endpointSum.count(), 7689, 'incorrect count for endpointSum')
Test.assertEquals(topTenErrURLs, [(u'/images/NASA-logosmall.gif', 8761), (u'/images/KSC-logosmall.gif', 7236), (u'/images/MOSAIC-logosmall.gif', 5197), (u'/images/USA-logosmall.gif', 5157), (u'/images/WORLD-logosmall.gif', 5020), (u'/images/ksclogo-medium.gif', 4728), (u'/history/apollo/images/apollo-logo1.gif', 2907), (u'/images/launch-logo.gif', 2811), (u'/', 2199), (u'/images/ksclogosmall.gif', 1622)], 'incorrect Top Ten failed URLs (topTenErrURLs)')
# #### **(3b) Exercise: Number of Unique Hosts**
# ####How many unique hosts are there in the entire log?
#
# ####Think about the steps that you need to perform to count the number of different hosts in the log.
# +
# TODO: Replace <FILL IN> with appropriate code
# HINT: Do you recall the tips from (3a)? Each of these <FILL IN> could be an transformation or action.
hosts = access_logs.map(lambda log: (log.host, 1))
uniqueHosts = hosts.reduceByKey(lambda a, b : a + b)
uniqueHostCount = uniqueHosts.count()
print 'Unique hosts: %d' % uniqueHostCount
# -
# TEST Number of unique hosts (3b)
Test.assertEquals(uniqueHostCount, 54507, 'incorrect uniqueHostCount')
# #### **(3c) Exercise: Number of Unique Daily Hosts**
# ####For an advanced exercise, let's determine the number of unique hosts in the entire log on a day-by-day basis. This computation will give us counts of the number of unique daily hosts. We'd like a list sorted by increasing day of the month which includes the day of the month and the associated number of unique hosts for that day. Make sure you cache the resulting RDD `dailyHosts` so that we can reuse it in the next exercise.
#
# ####Think about the steps that you need to perform to count the number of different hosts that make requests *each* day.
# ####*Since the log only covers a single month, you can ignore the month.*
# +
# TODO: Replace <FILL IN> with appropriate code
dayToHostPairTuple = (access_logs.map(lambda log : ((log.date_time.day, log.host), 1)))
dayGroupedHosts = dayToHostPairTuple.reduceByKey(lambda v1,v2: v1+v2).map(lambda (k,v) : k)
dayHostCount = dayGroupedHosts.map(lambda (k, v) : (k,1)).reduceByKey(lambda v1, v2: v1+v2)
dailyHosts = (dayHostCount
.sortByKey()
.cache())
dailyHostsList = dailyHosts.take(30)
print 'Unique hosts per day: %s' % dailyHostsList
# -
# TEST Number of unique daily hosts (3c)
Test.assertEquals(dailyHosts.count(), 21, 'incorrect dailyHosts.count()')
Test.assertEquals(dailyHostsList, [(1, 2582), (3, 3222), (4, 4190), (5, 2502), (6, 2537), (7, 4106), (8, 4406), (9, 4317), (10, 4523), (11, 4346), (12, 2864), (13, 2650), (14, 4454), (15, 4214), (16, 4340), (17, 4385), (18, 4168), (19, 2550), (20, 2560), (21, 4134), (22, 4456)], 'incorrect dailyHostsList')
Test.assertTrue(dailyHosts.is_cached, 'incorrect dailyHosts.is_cached')
# #### **(3d) Exercise: Visualizing the Number of Unique Daily Hosts**
# ####Using the results from the previous exercise, use `matplotlib` to plot a "Line" graph of the unique hosts requests by day.
# #### `daysWithHosts` should be a list of days and `hosts` should be a list of number of unique hosts for each corresponding day.
# #### * How could you convert a RDD into a list? See the [`collect()` method](http://spark.apache.org/docs/latest/api/python/pyspark.html?highlight=collect#pyspark.RDD.collect)*
# +
# TODO: Replace <FILL IN> with appropriate code
daysWithHosts = dailyHosts.map(lambda (k,v): k).collect()
hosts = dailyHosts.map(lambda (k,v): v).collect()
# -
# TEST Visualizing unique daily hosts (3d)
test_days = range(1, 23)
test_days.remove(2)
Test.assertEquals(daysWithHosts, test_days, 'incorrect days')
Test.assertEquals(hosts, [2582, 3222, 4190, 2502, 2537, 4106, 4406, 4317, 4523, 4346, 2864, 2650, 4454, 4214, 4340, 4385, 4168, 2550, 2560, 4134, 4456], 'incorrect hosts')
fig = plt.figure(figsize=(8,4.5), facecolor='white', edgecolor='white')
plt.axis([min(daysWithHosts), max(daysWithHosts), 0, max(hosts)+500])
plt.grid(b=True, which='major', axis='y')
plt.xlabel('Day')
plt.ylabel('Hosts')
plt.plot(daysWithHosts, hosts)
pass
# #### **(3e) Exercise: Average Number of Daily Requests per Hosts**
# ####Next, let's determine the average number of requests on a day-by-day basis. We'd like a list by increasing day of the month and the associated average number of requests per host for that day. Make sure you cache the resulting RDD `avgDailyReqPerHost` so that we can reuse it in the next exercise.
# ####To compute the average number of requests per host, get the total number of request across all hosts and divide that by the number of unique hosts.
# ####*Since the log only covers a single month, you can skip checking for the month.*
# ####*Also to keep it simple, when calculating the approximate average use the integer value - you do not need to upcast to float*
# +
# TODO: Replace <FILL IN> with appropriate code
dayAndHostTuple = access_logs.map(lambda log: ((log.date_time.day, log.host),1)).reduceByKey(lambda v1, v2 : v1+v2)
groupedByDay = dayAndHostTuple.map(lambda ((k,v),cnt): (k,(1,cnt))).reduceByKey(lambda v1, v2 : (v1[0]+v2[0], v1[1]+v2[1]))
sortedByDay = groupedByDay.sortByKey()
avgDailyReqPerHost = (sortedByDay
.map(lambda (k,v): (k, v[1]/v[0]))
.cache())
avgDailyReqPerHostList = avgDailyReqPerHost.take(30)
print 'Average number of daily requests per Hosts is %s' % avgDailyReqPerHostList
# -
# TEST Average number of daily requests per hosts (3e)
Test.assertEquals(avgDailyReqPerHostList, [(1, 13), (3, 12), (4, 14), (5, 12), (6, 12), (7, 13), (8, 13), (9, 14), (10, 13), (11, 14), (12, 13), (13, 13), (14, 13), (15, 13), (16, 13), (17, 13), (18, 13), (19, 12), (20, 12), (21, 13), (22, 12)], 'incorrect avgDailyReqPerHostList')
Test.assertTrue(avgDailyReqPerHost.is_cached, 'incorrect avgDailyReqPerHost.is_cache')
# #### **(3f) Exercise: Visualizing the Average Daily Requests per Unique Host**
# ####Using the result `avgDailyReqPerHost` from the previous exercise, use `matplotlib` to plot a "Line" graph of the average daily requests per unique host by day.
# #### `daysWithAvg` should be a list of days and `avgs` should be a list of average daily requests per unique hosts for each corresponding day.
# +
# TODO: Replace <FILL IN> with appropriate code
daysWithAvg = avgDailyReqPerHost.map(lambda (k,v): k).take(30)
avgs = avgDailyReqPerHost.map(lambda(k,v): v).take(30)
# -
# TEST Average Daily Requests per Unique Host (3f)
Test.assertEquals(daysWithAvg, [1, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22], 'incorrect days')
Test.assertEquals(avgs, [13, 12, 14, 12, 12, 13, 13, 14, 13, 14, 13, 13, 13, 13, 13, 13, 13, 12, 12, 13, 12], 'incorrect avgs')
fig = plt.figure(figsize=(8,4.2), facecolor='white', edgecolor='white')
plt.axis([0, max(daysWithAvg), 0, max(avgs)+2])
plt.grid(b=True, which='major', axis='y')
plt.xlabel('Day')
plt.ylabel('Average')
plt.plot(daysWithAvg, avgs)
pass
# ### **Part 4: Exploring 404 Response Codes**
#
# ####Let's drill down and explore the error 404 response code records. 404 errors are returned when an endpoint is not found by the server (i.e., a missing page or object).
# #### **(4a) Exercise: Counting 404 Response Codes**
# #### Create a RDD containing only log records with a 404 response code. Make sure you `cache()` the RDD `badRecords` as we will use it in the rest of this exercise.
#
# #### How many 404 records are in the log?
# +
# TODO: Replace <FILL IN> with appropriate code
badRecords = (access_logs
.filter(lambda log : log.response_code==404)
.cache()
)
print 'Found %d 404 URLs' % badRecords.count()
# -
# TEST Counting 404 (4a)
Test.assertEquals(badRecords.count(), 6185, 'incorrect badRecords.count()')
Test.assertTrue(badRecords.is_cached, 'incorrect badRecords.is_cached')
# #### **(4b) Exercise: Listing 404 Response Code Records**
# ####Using the RDD containing only log records with a 404 response code that you cached in part (4a), print out a list up to 40 **distinct** endpoints that generate 404 errors - *no endpoint should appear more than once in your list.*
# +
# TODO: Replace <FILL IN> with appropriate code
badEndpoints = badRecords.map(lambda log: (log.endpoint,1))
badUniqueEndpoints = badEndpoints.reduceByKey(lambda v1,v2 :1).map(lambda (k, v): k)
badUniqueEndpointsPick40 = badUniqueEndpoints.take(40)
print '404 URLS: %s' % badUniqueEndpointsPick40
# +
# TEST Listing 404 records (4b)
badUniqueEndpointsSet40 = set(badUniqueEndpointsPick40)
Test.assertEquals(len(badUniqueEndpointsSet40), 40, 'badUniqueEndpointsPick40 not distinct')
# -
# #### **(4c) Exercise: Listing the Top Twenty 404 Response Code Endpoints**
# ####Using the RDD containing only log records with a 404 response code that you cached in part (4a), print out a list of the top twenty endpoints that generate the most 404 errors.
# ####*Remember, top endpoints should be in sorted order*
# +
# TODO: Replace <FILL IN> with appropriate code
badEndpointsCountPairTuple = badRecords.map(lambda log: (log.endpoint, 1))
badEndpointsSum = badEndpointsCountPairTuple.reduceByKey(lambda v1, v2: v1+v2)
badEndpointsTop20 = badEndpointsSum.takeOrdered(20, lambda (k,v): -1*v)
print 'Top Twenty 404 URLs: %s' % badEndpointsTop20
# -
# TEST Top twenty 404 URLs (4c)
Test.assertEquals(badEndpointsTop20, [(u'/pub/winvn/readme.txt', 633), (u'/pub/winvn/release.txt', 494), (u'/shuttle/missions/STS-69/mission-STS-69.html', 431), (u'/images/nasa-logo.gif', 319), (u'/elv/DELTA/uncons.htm', 178), (u'/shuttle/missions/sts-68/ksc-upclose.gif', 156), (u'/history/apollo/sa-1/sa-1-patch-small.gif', 146), (u'/images/crawlerway-logo.gif', 120), (u'/://spacelink.msfc.nasa.gov', 117), (u'/history/apollo/pad-abort-test-1/pad-abort-test-1-patch-small.gif', 100), (u'/history/apollo/a-001/a-001-patch-small.gif', 97), (u'/images/Nasa-logo.gif', 85), (u'/shuttle/resources/orbiters/atlantis.gif', 64), (u'/history/apollo/images/little-joe.jpg', 62), (u'/images/lf-logo.gif', 59), (u'/shuttle/resources/orbiters/discovery.gif', 56), (u'/shuttle/resources/orbiters/challenger.gif', 54), (u'/robots.txt', 53), (u'/elv/new01.gif>', 43), (u'/history/apollo/pad-abort-test-2/pad-abort-test-2-patch-small.gif', 38)], 'incorrect badEndpointsTop20')
# #### **(4d) Exercise: Listing the Top Twenty-five 404 Response Code Hosts**
# ####Instead of looking at the endpoints that generated 404 errors, let's look at the hosts that encountered 404 errors. Using the RDD containing only log records with a 404 response code that you cached in part (4a), print out a list of the top twenty-five hosts that generate the most 404 errors.
# +
# TODO: Replace <FILL IN> with appropriate code
errHostsCountPairTuple = badRecords.map(lambda log: (log.host, 1))
errHostsSum = errHostsCountPairTuple.reduceByKey(lambda v1, v2: v1+v2)
errHostsTop25 = errHostsSum.takeOrdered(25, lambda (k,v): -1*v)
print 'Top 25 hosts that generated errors: %s' % errHostsTop25
# +
# TEST Top twenty-five 404 response code hosts (4d)
Test.assertEquals(len(errHostsTop25), 25, 'length of errHostsTop25 is not 25')
Test.assertEquals(len(set(errHostsTop25) - set([(u'maz3.maz.net', 39), (u'piweba3y.prodigy.com', 39), (u'gate.barr.com', 38), (u'm38-370-9.mit.edu', 37), (u'ts8-1.westwood.ts.ucla.edu', 37), (u'nexus.mlckew.edu.au', 37), (u'172.16.17.32', 33), (u'172.16.31.10', 27), (u'spica.sci.isas.ac.jp', 27), (u'www-d4.proxy.aol.com', 26), (u'www-c4.proxy.aol.com', 25), (u'192.168.127.12', 25), (u'192.168.127.12', 25), (u'internet-gw.watson.ibm.com', 24), (u'scooter.pa-x.dec.com', 23), (u'crl5.crl.com', 23), (u'piweba5y.prodigy.com', 23), (u'onramp2-9.onr.com', 22), (u'slip145-189.ut.nl.ibm.net', 22), (u'172.16.58.3.sap2.artic.edu', 21), (u'gn2.getnet.<EMAIL>', 20), (u'msp1-16.nas.mr.net', 20), (u'isou24.vilspa.esa.es', 19), (u'dial055.mbnet.mb.ca', 19), (u'tigger.nashscene.com', 19)])), 0, 'incorrect errHostsTop25')
# -
# #### **(4e) Exercise: Listing 404 Response Codes per Day**
# ####Let's explore the 404 records temporally. Break down the 404 requests by day (`cache()` the RDD `errDateSorted`) and get the daily counts sorted by day as a list.
# ####*Since the log only covers a single month, you can ignore the month in your checks.*
# +
# TODO: Replace <FILL IN> with appropriate code
errDateCountPairTuple = badRecords.map(lambda log: (log.date_time.day, 1))
errDateSum = errDateCountPairTuple.reduceByKey(lambda v1, v2: v1+v2)
errDateSorted = (errDateSum
.sortByKey()
.cache())
errByDate = errDateSorted.take(30)
print '404 Errors by day: %s' % errByDate
# -
# TEST 404 response codes per day (4e)
Test.assertEquals(errByDate, [(1, 243), (3, 303), (4, 346), (5, 234), (6, 372), (7, 532), (8, 381), (9, 279), (10, 314), (11, 263), (12, 195), (13, 216), (14, 287), (15, 326), (16, 258), (17, 269), (18, 255), (19, 207), (20, 312), (21, 305), (22, 288)], 'incorrect errByDate')
Test.assertTrue(errDateSorted.is_cached, 'incorrect errDateSorted.is_cached')
# #### **(4f) Exercise: Visualizing the 404 Response Codes by Day**
# ####Using the results from the previous exercise, use `matplotlib` to plot a "Line" or "Bar" graph of the 404 response codes by day.
# +
# TODO: Replace <FILL IN> with appropriate code
daysWithErrors404 = errDateSorted.map(lambda (k,v): k).take(30)
errors404ByDay = errDateSorted.map(lambda (k,v):v).take(30)
# -
# TEST Visualizing the 404 Response Codes by Day (4f)
Test.assertEquals(daysWithErrors404, [1, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22], 'incorrect daysWithErrors404')
Test.assertEquals(errors404ByDay, [243, 303, 346, 234, 372, 532, 381, 279, 314, 263, 195, 216, 287, 326, 258, 269, 255, 207, 312, 305, 288], 'incorrect errors404ByDay')
fig = plt.figure(figsize=(8,4.2), facecolor='white', edgecolor='white')
plt.axis([0, max(daysWithErrors404), 0, max(errors404ByDay)])
plt.grid(b=True, which='major', axis='y')
plt.xlabel('Day')
plt.ylabel('404 Errors')
plt.plot(daysWithErrors404, errors404ByDay)
pass
# #### **(4g) Exercise: Top Five Days for 404 Response Codes **
# ####Using the RDD `errDateSorted` you cached in the part (4e), what are the top five days for 404 response codes and the corresponding counts of 404 response codes?
# +
# TODO: Replace <FILL IN> with appropriate code
topErrDate = errDateSorted.takeOrdered(5, lambda (k,v):-1*v)
print 'Top Five dates for 404 requests: %s' % topErrDate
# -
# TEST Five dates for 404 requests (4g)
Test.assertEquals(topErrDate, [(7, 532), (8, 381), (6, 372), (4, 346), (15, 326)], 'incorrect topErrDate')
# #### **(4h) Exercise: Hourly 404 Response Codes**
# ####Using the RDD `badRecords` you cached in the part (4a) and by hour of the day and in increasing order, create an RDD containing how many requests had a 404 return code for each hour of the day (midnight starts at 0). Cache the resulting RDD hourRecordsSorted and print that as a list.
# +
# TODO: Replace <FILL IN> with appropriate code
hourCountPairTuple = badRecords.map(lambda log: (log.date_time.hour, 1))
hourRecordsSum = hourCountPairTuple.reduceByKey(lambda v1, v2: v1+v2)
hourRecordsSorted = (hourRecordsSum
.sortByKey()
.cache())
errHourList = hourRecordsSorted.take(24)
print 'Top hours for 404 requests: %s' % errHourList
# -
# TEST Hourly 404 response codes (4h)
Test.assertEquals(errHourList, [(0, 175), (1, 171), (2, 422), (3, 272), (4, 102), (5, 95), (6, 93), (7, 122), (8, 199), (9, 185), (10, 329), (11, 263), (12, 438), (13, 397), (14, 318), (15, 347), (16, 373), (17, 330), (18, 268), (19, 269), (20, 270), (21, 241), (22, 234), (23, 272)], 'incorrect errHourList')
Test.assertTrue(hourRecordsSorted.is_cached, 'incorrect hourRecordsSorted.is_cached')
# #### **(4i) Exercise: Visualizing the 404 Response Codes by Hour**
# ####Using the results from the previous exercise, use `matplotlib` to plot a "Line" or "Bar" graph of the 404 response codes by hour.
# +
# TODO: Replace <FILL IN> with appropriate code
hoursWithErrors404 = hourRecordsSorted.map(lambda (k,v): k).take(24)
errors404ByHours = hourRecordsSorted.map(lambda (k,v): v).take(24)
# -
# TEST Visualizing the 404 Response Codes by Hour (4i)
Test.assertEquals(hoursWithErrors404, [0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23], 'incorrect hoursWithErrors404')
Test.assertEquals(errors404ByHours, [175, 171, 422, 272, 102, 95, 93, 122, 199, 185, 329, 263, 438, 397, 318, 347, 373, 330, 268, 269, 270, 241, 234, 272], 'incorrect errors404ByHours')
fig = plt.figure(figsize=(8,4.2), facecolor='white', edgecolor='white')
plt.axis([0, max(hoursWithErrors404), 0, max(errors404ByHours)])
plt.grid(b=True, which='major', axis='y')
plt.xlabel('Hour')
plt.ylabel('404 Errors')
plt.plot(hoursWithErrors404, errors404ByHours)
pass
|
cs1001x_lab2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import json
from pathlib import Path
NEMO_PATH = Path('/src/NeMo/')
# +
import json
from pathlib import Path
decoder = json.JSONDecoder()
def read_json(file):
res = []
with open(file, 'r') as f:
line = f.readline()
while line:
res.append(decoder.raw_decode(line)[0])
line = f.readline()
return res
# +
train_all = []
val_all = []
roots = [
'/opt/storage/datasets/audio/japanese/CSJ',
'/opt/storage/datasets/audio/japanese/jvs_ver1',
'/opt/storage/datasets/audio/japanese/JNAS'
]
# 入れたくないコーパスは適宜コメントアウト
for r in roots:
train = Path(r) / 'mix_train_manifest.json'
val = Path(r) / 'mix_val_manifest.json'
train_all.extend(read_json(train))
val_all.extend(read_json(val))
# +
with open(NEMO_PATH / 'examples/asr/conf/mix_train_manifest-test.json', 'w') as f:
for metadata in train_all:
json.dump(metadata, f, ensure_ascii=False)
f.write('\n')
with open(NEMO_PATH / 'examples/asr/conf/mix_val_manifest-test.json', 'w') as f:
for metadata in val_all:
json.dump(metadata, f, ensure_ascii=False)
f.write('\n')
# -
|
prepare_conf/gather.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="BJsrTjueHRPH"
# # Introduction to Descriptive and Predictive Models
#
# This module introduces the core ideas of *machine learning*, which is subdivided into **unsupervised** and **supervised** machine learning.
#
# We break it into three segments:
#
# 1. Representing data for building machine learning models.
# 2. Unsupervised machine learning.
# 3. Supervised machine learning.
# + [markdown] colab_type="text" id="Wp7qRQdWL5tZ"
# # 3. Supervised Machine Learning
#
# Within supervised machine learning, we often use techniques based on regression -- including artificial neural networks.
#
# Let's start with the technique of **gradient descent**, useful for doing regression. We'll soon use it for training neural networks, but we'll start simpler.
# + [markdown] colab_type="text" id="829riMBRIoXm"
# ## 3.4 Gradient Descent
#
# ### 3.4.1 Gradient Descent with a Linear Function
#
# To look at gradient descent, let's first consider a simple example, with a linear value for our prediction. (We'll relax this in a moment to consider the logistic function.)
#
# We can define the cost function to be Mean Squared Error as follows:
#
# $MSE = {1 \over n}\Sigma^n_{i=1}(\hat{y}^{(i)} - y^{(i)})$
#
# where
#
# $\hat{y}^{(i)} = \sigma(w^T \cdot x^{(i)})$.
#
# For this part we'll generate a simple artificial dataset with two "blob" clusters.
#
# + colab_type="code" id="FQO-gsoQRk-3" colab={}
import numpy as np
from sklearn.datasets import make_blobs
(X, y) = make_blobs(n_samples=1000, n_features=2, centers=2, \
cluster_std=1.10, random_state=42)
# + colab_type="code" id="12PLfC4HSaIQ" outputId="54977b7c-8a96-4cd3-f2ba-d9d7d147824e" colab={"base_uri": "https://localhost:8080/", "height": 282}
import matplotlib.pyplot as plt
plt.scatter(x=X[:, 0], y=X[:, 1], c=y)
# + colab_type="code" id="d6UGS25F9OfK" outputId="228025f3-fe46-42ad-a459-01a04d903db6" colab={"base_uri": "https://localhost:8080/", "height": 554}
# Reload our data
from sklearn.model_selection import train_test_split
import sklearn.metrics
X_train, X_test, y_train, y_test = train_test_split(X, \
y, \
test_size=0.30)
y_train
# + colab_type="code" id="IZEoq_-_Buif" outputId="adab4c1b-0af6-4006-8f5c-fcab59aa0ebd" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Here's our cost function, as mean squared error
def cost_fn(X, y, p, fn):
if len(X.shape) == 1:
# Single row, return the squared error
return np.square(y - fn(X[p]))
else:
# Matrix, return the mean of the squared errors
return np.average([cost_fn(X[i], y[i], p, fn) for i in range (0, X.shape[0])])
# We'll excerpt one instance from the training set
x_sample = X_train[0]
y_sample = y_train[0]
x_sample
# + colab_type="code" id="krwhAxsAJzb6" outputId="506bebf6-deea-4807-eb87-0ea44ecadecd" colab={"base_uri": "https://localhost:8080/", "height": 268}
import matplotlib.pyplot as plt
x_coord = range(-10,10)
plt.figure(figsize=(12, 4))
# Now let's plot the error function for a *single instance*, for each feature,
# for a range of weights and the *linear* function x*w
for feature in range(0, 2):
ax = plt.subplot(1, 2, feature+1)
ax.plot(x_coord, [cost_fn(x_sample, y_sample, feature, lambda x: x*w) for w in x_coord])
ax.set_ylabel('Feature ' + str(feature))
# + [markdown] colab_type="text" id="9wzzFO_XxXps"
# If you look above, each feature has a parabola, with a different scale and a different minimum.
# + [markdown] colab_type="text" id="wO7Z9-sAxror"
# Now we'll look at *all* of the instances, for one feature.
# + colab_type="code" id="kYbiurxNJnEM" outputId="34df872b-5c9c-41d1-af87-1dc97723ff78" colab={"base_uri": "https://localhost:8080/", "height": 265}
import matplotlib.pyplot as plt
x_coord = range(-10,10)
plt.figure(figsize=(12, 4))
# Now let's plot the error function for each feature,
# for a range of weights and the *linear* function x*w
for feature in range(0, 2):
ax = plt.subplot(1, 2, feature+1)
ax.plot(x_coord, [cost_fn(X_train, y_train, feature, lambda x: x*w) for w in x_coord])
ax.set_ylabel('Feature ' + str(feature))
# + [markdown] colab_type="text" id="lfsbc2cM6yP8"
# ## Gradient Descent for Training Logistic Regression
#
# Let's try this with logistic regression, using the sigmoid function...
# + colab_type="code" id="MZPIMFNr60Gq" outputId="8bf2b90b-c49a-4fe4-8462-dd34701ce6b2" colab={"base_uri": "https://localhost:8080/", "height": 353}
import numpy as np
# Here is our sigmoid function for making
# predictions with logistic regression
# or with perceptron-style neural nets
def prediction(x):
return 1.0 / (1 + np.exp(-x))
def gradient_descent(epochs, eta, X, w, y):
"""
The gradient descent iterates for *epochs* rounds, making a step
of size eta. It will be adjusting w, based on the prediction for each
instance vs y and the overall error.
"""
# We'll use this list to accumulate
# the error
overall_error = []
# Iterate over each epoch
for i in range(epochs):
# This is y-hat, the predictions for each
# class label before they are thresholded
# based on comparison with 0.5
predictions = prediction(X.dot(w))
# The overall error, as a vector
error = (predictions - y)
# Record the MSE so we can plot it
mean_sq_error = np.sum(error ** 2) / X.shape[0]
overall_error.append(mean_sq_error)
# Now we update the weights.
# The gradient is based on the partial derivative
# of the MSE with respect to w.
gradient = 2 / X.shape[0] * X.T.dot(error)
w = w - eta * gradient
return w, overall_error
# Training rounds or epochs
epochs = 500
eta = 0.01
w = np.random.randn(2)
weights, evect = gradient_descent(epochs, eta, X, w, y)
print (weights)
print (evect)
# Plot the mean-squared error
plt.plot(range(0,epochs), evect)
plt.xlabel('Gradient Descent Epoch #')
plt.ylabel('Mean-squared error (loss)')
# + colab_type="code" id="r7CAqODetbXZ" outputId="6961f362-1f06-4abf-dbe3-a1583c5e7238" colab={"base_uri": "https://localhost:8080/", "height": 279}
x_coord = range(-10,10)
plt.figure(figsize=(12, 4))
# Now let's plot the error function for each feature,
# for a range of weights and the output of the sigmoid function over x*w
for feature in range(0, 2):
ax = plt.subplot(1, 2, feature+1)
ax.plot(x_coord, [cost_fn(X_train, y_train, feature, lambda x: prediction(x*w)) for w in x_coord],
marker='x')
ax.axvline(x=weights[feature], color='red')
ax.set_ylabel('Cost vs weight on feature ' + str(feature))
ax.set_xlabel('Weight w' + str(feature))
# + [markdown] colab_type="text" id="89LKGhu4837u"
# We've trained the weights -- let's make predictions!
# + colab_type="code" id="4ypZeyVe0A__" outputId="50f4af1c-715a-46bb-fe4a-f2656201a619" colab={"base_uri": "https://localhost:8080/", "height": 1000}
for item in range(len(X_test)):
predicted_label = 0 if prediction(X_test[item].dot(weights)) < 0.5 else 1
print('Prediction {} vs {}'.format(predicted_label, y_test[item]))
# + colab_type="code" id="9S9e_B9M9MA0" outputId="79ed210f-7e36-42e2-bb98-6aca99dc155b" colab={"base_uri": "https://localhost:8080/", "height": 252}
from sklearn.linear_model import Perceptron
clf = Perceptron(random_state=42)
clf.fit(X_train, y_train)
clf.predict(X_test)
# + [markdown] colab_type="text" id="kc9AKcW6fGS3"
# # Multilayer Perceptrons
#
# Let's load some data suitable for training a simple multilayer perceptron. The MNIST dataset has handwritten letters and we can train a classifier to predict the written digit.
# + colab_type="code" id="JzAWrlyAcUPo" colab={}
from sklearn.datasets import fetch_openml
# Load data from https://www.openml.org/d/554
X, y_str = fetch_openml('mnist_784', version=1, return_X_y=True)
y = np.array([int(x) for x in y_str])
# + colab_type="code" id="JBPW8R69fElU" outputId="3142f3fe-aadf-4164-a067-997a3ae7422c" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Each row has 784 pixels
X[1].shape
# + colab_type="code" id="uFQaPvP_fVod" colab={}
# Which really represents a 28x28 grid...
img = X[1].reshape(28,28)
# + colab_type="code" id="Tn5vBy1EfgpV" outputId="ee507820-c9c3-470a-f172-0d55811643fe" colab={"base_uri": "https://localhost:8080/", "height": 282}
import matplotlib.pyplot as plt
plt.imshow(img, cmap='gray')
# + colab_type="code" id="ac6kF9R1fo-w" outputId="48fbf040-841d-417d-cf7d-30f003e2e2d9" colab={"base_uri": "https://localhost:8080/", "height": 703}
plt.figure(figsize=(12, 12))
for img in range(100):
ax = plt.subplot(10, 10, img+1)
plt.imshow(X[img].reshape(28,28), cmap='gray')
# + colab_type="code" id="gLN9sFJohF_B" colab={}
X_train, X_test, y_train, y_test = train_test_split(X, \
y, \
test_size=0.30)
# + colab_type="code" id="59rKlgZMgb6B" outputId="62b9a7dc-840a-474c-a736-08d4c38a729e" colab={"base_uri": "https://localhost:8080/", "height": 87}
from sklearn.neural_network import MLPClassifier
mlp = MLPClassifier(hidden_layer_sizes=(10,10))
mlp.fit(X_train, y_train)
mlp.predict(X_test)
# + [markdown] colab_type="text" id="COWEM5zJnTwX"
# ## Implementing the MLP
#
# Let's see how this would look under the covers!
# + colab_type="code" id="7nnDAwr5imBs" outputId="acf515d2-b04a-4102-8a70-d39e3bffd5bf" colab={"base_uri": "https://localhost:8080/", "height": 134}
from scipy.special import expit
class FFClassifier:
layer_weights = []
costs = []
epochs = 100
eta = 0.001
decrease_eta = 0.000001
# Initialize weights for each layer
def __init__(self, features, hidden_layers, classes, epochs=100, eta=0.001):
# Initial weights = 1 for each feature, plus 1 for bias
first = np.random.uniform(-1.0, 1.0, size=hidden_layers[0] * (features+1))\
.reshape(hidden_layers[0], features+1)
self.layer_weights.append(first)
self.epochs = epochs
self.eta = eta
print ('Initial hidden layer:', first.shape)
# "Middle" hidden layers
for layer in range(0, len(hidden_layers) - 1):
hidden = np.random.uniform(-1.0, 1.0, size=(hidden_layers[layer]+1) * hidden_layers[layer+1])\
.reshape(hidden_layers[layer+1], hidden_layers[layer]+1)
self.layer_weights.append(hidden)
print ('Hidden layer:', hidden.shape)
# Last
last_layer = len(hidden_layers) - 1
hidden = np.random.uniform(-1.0, 1.0, size=(hidden_layers[last_layer]+1) * classes)\
.reshape(classes, hidden_layers[last_layer]+1)
self.layer_weights.append(hidden)
print ('Last layer:', hidden.shape)
return
# expit is a version of sigmoid
def activation(self, sigma):
return expit(sigma)
def feed_forward(self, X):
sums = []
layer_outputs = []
layer = 0
X_with_bias = np.ones((X.shape[1] + 1, X.shape[0]))
X_with_bias[1:, 0:] = X.T
# Dot products for weights from inputs to next
# layer
result = self.layer_weights[0].dot(X_with_bias)
sums.append(result)
# Run them through sigmoid
result = self.activation(result)
layer_outputs.append(result)
print (result.shape)
for layer in range(1, len(self.layer_weights)):
layer_with_bias = np.ones((result.shape[0] + 1, result.shape[1]))
layer_with_bias[1:, :] = result
result = self.layer_weights[layer].dot(layer_with_bias)
sums.append(result)
result = self.activation(result)
layer_outputs.append(result)
print (result.shape)
return sums, layer_outputs, result
@staticmethod
def onehot_encode(y):
classes = np.unique(y).shape[0]
ret = np.zeros((classes, y.shape[0]))
for i, v in enumerate(y):
ret[v, i] = 1.0
return ret.T
def cost(self, y_onehot, output):
term1 = -y_enc * (np.log(output + 1e-5))
term2 = (1. - y_enc) * np.log(1. - output + 1e-5)
return np.sum(term1 - term2)
def fit(self, X, y_onehot):
X_tmp = X.copy()
y_tmp = y_onehot.copy()
eta = self.eta
for i in range(self.epochs):
eta /= (1 + self.decrease_eta)
shuffle = np.random.permutation(y_tmp.shape[0])
X_tmp, y_tmp = X_tmp[shuffle], y_tmp[:, shuffle]
sums, layer_outputs, result = self.feed_forward(X_tmp)
cost = self.cost(y_tmp, result)
self.costs.append(cost)
def predict(self, X):
sums, layer_outputs, result = self.feed_forward(X)
return np.argmax(result, axis=0)
clf = FFClassifier(X.shape[1], [15, 15], 10)
y_oh = FFClassifier.onehot_encode(y_train)
clf.predict(X_train)
# + colab_type="code" id="w0ocPcAWn877" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="016829f3-4f5c-4472-fa0b-4ff4d798922a"
y
# + colab_type="code" id="8B5FoCeGn3NT" colab={}
|
opends4all-resources/opends4all-machine-learning/SUPERVISED-ML-NN-neural-nets-intermediate.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 06 Machine Learning
# Logistic regression and randon forest models are used to predict the divorce of male and female actors.
#
# I analyed seperated data from male and female found that the sex does no have a significant contribution to the divorce. Therefore, I combine all actor and actress data and perform ML model.
#
# +
# Data manipulation
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import ast
# Import sklearn.preprocessing.StandardScaler
from sklearn.preprocessing import MinMaxScaler, StandardScaler
from sklearn import svm, tree, linear_model, neighbors, naive_bayes, ensemble, discriminant_analysis, gaussian_process
from sklearn.model_selection import train_test_split
from sklearn.metrics import r2_score, mean_squared_error
from sklearn.ensemble import RandomForestClassifier
#from xgboost import XGBClassifier
#Common Model Helpers
from sklearn.preprocessing import OneHotEncoder, LabelEncoder
from sklearn import feature_selection
from sklearn import model_selection
from sklearn import metrics
import warnings
warnings.filterwarnings('ignore')
# %matplotlib inline
# +
#import pandas as pd
df_X = pd.read_csv('export_df_X_forML.csv')
df_y = pd.read_csv('export_df_y_forML.csv')
# male data
df_X_M = pd.read_csv('export_df_X_M_forML.csv')
df_y_M = pd.read_csv('export_df_y_M_forML.csv')
# female data
df_X_F = pd.read_csv('export_df_X_F_forML.csv')
df_y_F = pd.read_csv('export_df_y_F_forML.csv')
# remove data
df_X = df_X.drop(["Unnamed: 0"], axis = 1)
df_y = df_y.drop(["Unnamed: 0"], axis = 1)
df_X_M = df_X_M.drop(["Unnamed: 0"], axis = 1)
df_y_M = df_y_M.drop(["Unnamed: 0"], axis = 1)
df_X_F = df_X_F.drop(["Unnamed: 0"], axis = 1)
df_y_F = df_y_F.drop(["Unnamed: 0"], axis = 1)
# -
df_X.columns
# rename the feature columns
df_X.rename(columns = {'age_diff':'age difference', 'zodiac_sp':'zodiac (spouse)',
'num_of_child_cl':'num of child', 'num_of_child_sp_cl':'num of child (spouse)',
'num_of_role':'num of role', 'num_of_role_sp': 'num of role (spouse)',
'geo_distance': 'geo distance', 'age_m_1':'age at 1st marriage',
'age_m_sp_1': 'age at 1st marriage (spouse)'}, inplace=True)
# ## Standardization
# Use Standardization to linear model to make algorithm less senstive to outlier
# +
# Initialize a scaler, then apply it to the features
scaler = StandardScaler()
df_X_ss = scaler.fit_transform(df_X)
df_X_M_ss = scaler.fit_transform(df_X_M)
df_X_F_ss = scaler.fit_transform(df_X_F)
# -
# Split data
# +
# Import train_test_split
from sklearn.model_selection import train_test_split
# Split the 'features' and 'income' data into training and testing sets
X_train, X_test, y_train, y_test = train_test_split(df_X_ss, df_y, test_size = 0.3, random_state = 12)
# seperated male and female dataset
X_M_train, X_M_test, y_M_train, y_M_test = train_test_split(df_X_M_ss, df_y_M, test_size = 0.3, random_state = 32)
X_F_train, X_F_test, y_F_train, y_F_test = train_test_split(df_X_F_ss, df_y_F, test_size = 0.3, random_state = 32)
# Show the results of the split
print("Training set for total has {} samples.".format(X_train.shape[0]))
print("Testing set for total has {} samples.".format(X_test.shape[0]))
# -
# ## Logistic regression
# Use grid search to find the optimized hyperparameter
# +
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import GridSearchCV
# Create regularization penalty space
penalty = ['l1', 'l2']
# Create regularization hyperparameter space
C = np.logspace(-3, 10, 20)
solver = ['liblinear', 'saga']
#multi_class = ['ovr']
# Create hyperparameter options
hyperparameters = dict(C=C, penalty=penalty, solver = solver)
# Create logistic regression
lr_model = LogisticRegression(random_state=32) # Instantiate
# Create grid search using 5-fold cross validation
clf = GridSearchCV(lr_model, hyperparameters, cv=5, verbose=0)
# +
# Fit grid search
best_lr_model = clf.fit(X_train, y_train) #Fit
y_test_preds_lr = best_lr_model.predict(X_test)
print('Accuracy of logistic regression classifier on test set: {:.4f}'.format(clf.best_score_))
# View best hyperparameters
print('Best Penalty:', best_lr_model.best_estimator_.get_params()['penalty'])
print('Best C:', best_lr_model.best_estimator_.get_params()['C'])
print('Best Model:', best_lr_model.best_estimator_)
# -
from sklearn.metrics import confusion_matrix, classification_report
confusion_matrix = confusion_matrix(y_test, y_test_preds_lr)
print(confusion_matrix)
from sklearn.metrics import classification_report
print(classification_report(y_test, y_test_preds_lr))
print(classification_report(y_test, y_test_preds_lr))
# +
from sklearn.metrics import roc_auc_score
from sklearn.metrics import roc_curve
logit_roc_auc= roc_auc_score(y_test, best_lr_model.predict(X_test) )
fpr, tpr, thresholds = roc_curve(y_test, best_lr_model.predict(X_test))
plt.figure()
plt.plot(fpr, tpr, label='Logistic Regression (area = %0.2f)' % logit_roc_auc)
plt.plot([0, 1], [0, 1],'r--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic')
plt.legend(loc="lower right")
plt.savefig('Log_ROC')
plt.show()
# -
# ## Random forest
# Use grid search anc cross validation to find the optimized hyperparameter
# +
from sklearn.ensemble import RandomForestRegressor
from sklearn.model_selection import RandomizedSearchCV
# Number of trees in random forest
n_estimators = [int(x) for x in np.linspace(start = 200, stop = 2000, num = 10)]
# Number of features to consider at every split
max_features = ['auto', 'sqrt']
# Maximum number of levels in tree
max_depth = [int(x) for x in np.linspace(2, 50, num = 10)]
max_depth.append(None)
# Minimum number of samples required to split a node
min_samples_split = [2, 5, 10, 20]
# Minimum number of samples required at each leaf node
min_samples_leaf = [2, 4, 8, 10]
# Method of selecting samples for training each tree
bootstrap = [True, False]
# Create the random grid
random_grid = {'n_estimators': n_estimators,
'max_features': max_features,
#'criterion': criterion,
'max_depth': max_depth,
'min_samples_split': min_samples_split,
'min_samples_leaf': min_samples_leaf,
'bootstrap': bootstrap}
print(random_grid)
# -
# ### Random search training
# Instantiate the random search and fit.
#
# **1.** Use the random grid to search for best hyperparameters. Search across 100 different combinations, and use all available cores
# First create the base model to tune
# Random search of parameters, using 3 fold cross validation,
rf_model = RandomizedSearchCV(estimator = rf_model, param_distributions = random_grid,
n_iter = 100, cv = 3, verbose=2, random_state=42, n_jobs = -1)
rf_model.fit(X_train, y_train)
print('Best Parameters:', rf_model.best_params_)
print('Best Model:', rf_model.best_estimator_)
# +
from sklearn.metrics import accuracy_score
best_rf_model= rf_model.best_estimator_
y_train_pred_rf= best_rf_model.predict(X_train).round()
y_test_pred_rf= best_rf_model.predict(X_test).round()
accuracy = accuracy_score(y_test, y_test_pred_rf)
print('Accuracy of random forest classifier on train set: {:.2f}'.format(accuracy_score(y_train, y_train_pred_rf)))
print('Accuracy of random forest classifier on test set: {:.2f}'.format(accuracy_score(y_test, y_test_pred_rf)))
# -
from sklearn.metrics import confusion_matrix, classification_report
confusion_matrix = confusion_matrix(y_test, y_test_pred_rf)
print(confusion_matrix)
# +
# feature importance
importance = best_rf_model.feature_importances_
# Sort feature importances in descending order
indices = np.argsort(importance)[::-1]
# Rearrange feature names so they match the sorted feature importances
names = [df_X.columns[i] for i in indices]
std = np.std([tree.feature_importances_ for tree in best_rf_model.estimators_], axis=0)
# Create plot
fig, ax = plt.subplots()
#plt.figure()
# Create plot title
plt.title("Feature Importance")
# Add bars
ax.barh(range(df_X.shape[1]), importance[indices], xerr=std[indices], align='center', color='lightblue', ecolor='black')
# Add feature names as y-axis labels
plt.yticks(range(df_X.shape[1]), names, rotation=0)
# Show plot
plt.show()
# -
# Learning:
# - Age, age of the first marrage are the most important feautres
# - Sex is acturally not a very important contrintor to determine the devorce
# - Number of child and number of role are not very important features
# - Geo distance between spouses are least important feature
# Next, we look at the data of male and female actors seperately
#
# **1.** Male actor data
# +
rf_model = RandomForestRegressor()
# Random search of parameters, using 3 fold cross validation,
# search across 100 different combinations, and use all available cores
rf_random_M = RandomizedSearchCV(estimator = rf_model, param_distributions = random_grid,
n_iter = 100, cv = 3, verbose=2, random_state=42, n_jobs = -1)
rf_random_M.fit(X_M_train, y_M_train)
# +
from sklearn.metrics import accuracy_score
best_rf_model_M = rf_random_M.best_estimator_
y_train_pred_rf_M = best_rf_model_M.predict(X_M_train).round()
y_test_pred_rf_M = best_rf_model_M.predict(X_M_test).round()
print('Accuracy of random forest classifier on train set (Male): {:.2f}'.format(accuracy_score(y_M_train, y_train_pred_rf_M)))
print('Accuracy of random forest classifier on test set (Male): {:.2f}'.format(accuracy_score(y_M_test, y_test_pred_rf_M)))
# +
importanceM = best_rf_model_M.feature_importances_
# Sort feature importances in descending order
indices = np.argsort(importanceM)[::-1]
# Rearrange feature names so they match the sorted feature importances
names_M = [df_X_M.columns[i] for i in indices]
std = np.std([tree.feature_importances_ for tree in best_rf_model_M.estimators_], axis=0)
# Create plot
fig, ax = plt.subplots()
#plt.figure()
# Create plot title
plt.title("Feature Importance")
# Add bars
ax.barh(range(df_X_M.shape[1]), importanceM[indices], xerr=std[indices], align='center', color='lightblue', ecolor='black')
# Add feature names as y-axis labels
plt.yticks(range(df_X_M.shape[1]), names_M, rotation=0)
# Show plot
plt.show()
# -
# **2.** Female actor data
# +
from sklearn.ensemble import RandomForestClassifier
#rfF= RandomForestClassifier(n_estimators= 200, min_samples_split= 5, min_samples_leaf= 4,
# max_features= 'auto', max_depth= 10, bootstrap= True)
rf_random_F = RandomizedSearchCV(estimator = rf_model, param_distributions = random_grid,
n_iter = 100, cv = 3, verbose=2, random_state=42, n_jobs = -1)
rf_random_F.fit(X_F_train, y_F_train)
# -
print('Best Parameters:', rf_random_F.best_params_)
print('Best Model:', rf_random_F.best_estimator_)
# +
from sklearn.metrics import accuracy_score
best_rf_model_F = rf_random_F.best_estimator_
y_train_pred_rf_F = best_rf_model_F.predict(X_F_train).round()
y_test_pred_rf_F = best_rf_model_F.predict(X_F_test).round()
print('Accuracy of random forest classifier on train set (Male): {:.2f}'.format(accuracy_score(y_F_train, y_train_pred_rf_F)))
print('Accuracy of random forest classifier on test set (Male): {:.2f}'.format(accuracy_score(y_F_test, y_test_pred_rf_F)))
# +
importanceF = best_rf_model_F.feature_importances_
# Sort feature importances in descending order
indices = np.argsort(importanceF)[::-1]
# Rearrange feature names so they match the sorted feature importances
names_F = [df_X_F.columns[i] for i in indices]
std = np.std([tree.feature_importances_ for tree in best_rf_model_F.estimators_], axis=0)
# Create plot
fig, ax = plt.subplots()
#plt.figure()
# Create plot title
plt.title("Feature Importance")
# Add bars
ax.barh(range(df_X_F.shape[1]), importanceF[indices], xerr=std[indices], align='center', color='lightblue', ecolor='black')
# Add feature names as y-axis labels
plt.yticks(range(df_X_F.shape[1]), names_F, rotation=0)
# Show plot
plt.show()
# -
|
notebooks/06_ML_model_building_tuning.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h2> Test macierzy numpy 2D <h2>
import numpy as np
a = [[11,12,13] , [21,22,23] , [31,32,33] ]
# Casting a to NP Array -> A
A = np.array(a)
print(type(A))
print(A.ndim)
print(A.shape)
# +
a=np.array([0,1])
b=np.array([1,0])
np.dot(a,b)
# -
|
numpy_2d_test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction to Hadoop MapReduce on Palmetto
# Run the following commands to check that your `.jhubrc` file contains the correct modules
# !cat ~/.jhubrc
# !cat ~/.bashrc
# If the modules are not there
# ! echo "module load openjdk/1.8.0_222-b10-gcc/8.3.1 hadoop/3.2.1-gcc/8.3.1" >> ~/.jhubrc
# ! echo "module load openjdk/1.8.0_222-b10-gcc/8.3.1 hadoop/3.2.1-gcc/8.3.1" >> ~/.bashrc
# Check again
# !cat ~/.jhubrc
# !cat ~/.bashrc
# ### Restart the JupyterLab server
# ## Launch the cluster
# !./init_hadoop.sh
# ## Test the cluster
# !./test_hadoop.sh
# ## Stop the cluster
# !./stop_hadoop.sh
|
intro-to-hadoop-00.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Filtrado de grupos
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
data =pd.read_csv("../../datasets/customer-churn-model/Customer Churn Model.txt")
data.head()
data.describe()
# ### Forma 1: Dividir utilizando la distribución normal
random_array = np.random.normal(0,1,3333)
plt.hist(random_array)
check = random_array < 0.8
check
check[0:10]
training = data[check]
training.describe()
test = data[~check]
test.describe()
# # Con la librería sklearn
from sklearn.model_selection import train_test_split
(train, test) = train_test_split(data, test_size = 0.2)
train.describe()
test.describe()
# ### Con shuffle
import sklearn
# +
shuffled_data = sklearn.utils.shuffle(data)
# -
shuffled_data.head()
cutid = int( 0.8 * len(data));
print (cutid)
train2 = shuffled_data[0:cutid]
test2= shuffled_data[cutid+1:len(data)]
train2.describe()
test2.describe()
|
notebooks/mios/12. Dividir el dataset en train y test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: otrepipe
# language: python
# name: otrepipe
# ---
# ## Searching for bouts for a day of alsa recording
# +
# %matplotlib inline
import os
import glob
import socket
import logging
import numpy as np
import pandas as pd
from scipy.io import wavfile
from scipy import signal
from matplotlib import pyplot as plt
from importlib import reload
logger = logging.getLogger()
handler = logging.StreamHandler()
formatter = logging.Formatter(
'%(asctime)s %(name)-12s %(levelname)-8s %(message)s')
handler.setFormatter(formatter)
logger.addHandler(handler)
logger.setLevel(logging.INFO)
logger.info('Running on {}'.format(socket.gethostname()))
# -
from ceciestunepipe.file import bcistructure as et
# ### Get the file locations for a session (day) of recordings
# +
reload(et)
sess_par = {'bird': 's_b1253_21',
'sess': '2021-06-29'}
exp_struct = et.get_exp_struct(sess_par['bird'], sess_par['sess'], ephys_software='alsa')
raw_folder = exp_struct['folders']['alsa']
# -
exp_struct['folders']
# #### search parameters
# Spectrograms are computed using librosa
#
# Additional parameters are for bout search criteria and functions to read the data
# +
# function for getting one channel out of a wave file
def read_wav_chan(wav_path: str, chan_id: int=0) -> tuple:
s_f, x = wavfile.read(wav_path, mmap=True)
return s_f, x[:, chan_id]
def sess_file_id(f_path):
n = int(os.path.split(f_path)[1].split('-')[-1].split('.wav')[0])
return n
hparams = {
# spectrogram
'num_freq':1024, #1024# how many channels to use in a spectrogram #
'preemphasis':0.97,
'frame_shift_ms':5, # step size for fft
'frame_length_ms':10, #128 # frame length for fft FRAME SAMPLES < NUM_FREQ!!!
'min_level_db':-55, # minimum threshold db for computing spe
'ref_level_db':110, # reference db for computing spec
'sample_rate':None, # sample rate of your data
# spectrograms
'mel_filter': False, # should a mel filter be used?
'num_mels':1024, # how many channels to use in the mel-spectrogram
'fmin': 500, # low frequency cutoff for mel filter
'fmax': 12000, # high frequency cutoff for mel filter
# spectrogram inversion
'max_iters':200,
'griffin_lim_iters':20,
'power':1.5,
# Added for the searching
'read_wav_fun': read_wav_chan, # function for loading the wav_like_stream (has to returns fs, ndarray)
'file_order_fun': sess_file_id, # function for extracting the file id within the session
'min_segment': 30, # Minimum length of supra_threshold to consider a 'syllable' (ms)
'min_silence': 2000, # Minmum distance between groups of syllables to consider separate bouts (ms)
'min_bout': 5000, # min bout duration (ms)
'peak_thresh_rms': 0.55, # threshold (rms) for peak acceptance,
'thresh_rms': 0.25, # threshold for detection of syllables
'mean_syl_rms_thresh': 0.3, #threshold for acceptance of mean rms across the syllable (relative to rms of the file)
'max_bout': 120000, #exclude bouts too long
'l_p_r_thresh': 100, # threshold for n of len_ms/peaks (typycally about 2-3 syllable spans
'waveform_edges': 1000, #get number of ms before and after the edges of the bout for the waveform sample
'bout_auto_file': 'bout_auto.pickle', # extension for saving the auto found files
'bout_curated_file': 'bout_checked.pickle', #extension for manually curated files (coming soon)
}
# -
# #### Get one wave file
# +
one_wav_path = os.path.join(exp_struct['folders']['alsa'], '09-00-01-02.wav')
s_f, x = read_wav_chan(one_wav_path)
hparams['sample_rate'] = s_f
# -
x.shape
plt.plot(x[:50000])
# ### try the function to search for bouts in the file
from ceciestunepipe.util.sound import boutsearch as bs
reload(bs)
# +
#bpd, x, p = bs.get_bouts_in_file(one_wav_path, hparams)
bouts_file_path = '/mnt/sphere/speech_bci/derived_data/s_b1253_21/2021-06-29/alsa/bouts_ceciestunepipe/bout_auto.pickle'
# -
bpd = pd.read_pickle(bouts_file_path)
plt.plot(bpd['waveform'].values[0])
bpd
# ### inspect the bouts and curate them
# #### visualize one bout
bpd['spectrogram'] = bpd['waveform'].apply(lambda x: bs.gimmepower(x, hparams)[2])
bpd.iloc[0]
# +
def viz_bout(df: pd.Series, ax_arr=None, sub_sample=10):
# get the power and the spectrogram
sxx = df['spectrogram'][:, ::sub_sample]
x = df['waveform'][::sub_sample]
if ax_arr is None:
fig, ax_arr = plt.subplots(nrows=2, figsize=(20, 4))
ax_arr[0].imshow(np.sqrt(sxx[::-1]), aspect='auto', cmap='inferno')
ax_arr[1].plot(x)
fig, ax = plt.subplots(nrows=2, figsize=(20, 3))
viz_bout(bpd.iloc[2], ax)
# -
# #### use it in a widget
#
from IPython.display import display, clear_output
import ipywidgets as widgets
from traitlets import CInt, link
class Counter(widgets.DOMWidget):
value = CInt(0)
value.tag(sync=True)
# + jupyter={"source_hidden": true}
class VizBout():
def __init__(self, hparams, bouts_pd):
self.bout = None
self.bouts_pd = bouts_pd
self.bout_series = None
self.is_bout = None
self.bout_counter = None
self.bout_id = None
self.buttons = {}
self.m_pick = None
self.fig = None
self.fig_ax = {}
self.fig_width = 2
self.sub_sample = 10
self.x = None
self.sxx = None
self.init_widget()
self.init_fig()
def init_fig(self):
self.fig = plt.figure()
self.fig_ax['waveform'] = self.fig.add_axes([0,0,self.fig_width,0.5])
self.fig_ax['spectrogram'] = self.fig.add_axes([0,.5,self.fig_width,0.5])
for ax in self.fig_ax.values():
ax.xaxis.set_ticks([])
ax.yaxis.set_ticks([])
self.show()
def init_widget(self):
self.bout_counter = Counter()
self.is_bout = widgets.Checkbox(description='is bout')
self.buttons['Next'] = widgets.Button(description="Next", button_style='info',
icon='plus')
self.buttons['Prev'] = widgets.Button(description="Prev", button_style='warning',
icon='minus')
self.buttons['Check'] = widgets.Button(description="Check", button_style='success',
icon='check')
self.buttons['Uncheck'] = widgets.Button(description="Uncheck", button_style='danger',
icon='wrong')
[b.on_click(self.button_click) for b in self.buttons.values()]
left_box = widgets.VBox([self.buttons['Prev'], self.buttons['Uncheck']])
right_box = widgets.VBox([self.buttons['Next'], self.buttons['Check']])
button_box = widgets.HBox([left_box, right_box])
self.m_pick = widgets.IntSlider(value=0, min=0, max=self.bouts_pd.index.size-1,step=1,
description="MotCandidate")
link((self.m_pick, 'value'), (self.bout_counter, 'value'))
self.update_bout()
display(button_box)
display(self.m_pick)
display(self.is_bout)
def button_click(self, button):
self.bout_id = self.bout_counter.value
curr_bout = self.bout_counter
if button.description == 'Next':
curr_bout.value += 1
elif button.description == 'Prev':
curr_bout.value -= 1
elif button.description == 'Check':
self.bouts_pd.set_value(self.bout_id, 'bout_check', True)
curr_bout.value += 1
elif button.description == 'Uncheck':
self.bouts_pd.set_value(self.bout_id, 'bout_check', False)
curr_bout.value += 1
def slider_change(self, change):
logger.info('slider changed')
#self.bout_counter = change.new
#clear_output(True)
self.update_bout()
self.show()
def bout_checked(self, bc):
# print "bout checked"
# print bc['new']
# print self.motiff
self.bouts_pd.set_value(self.bout_id, 'bout_check', bc['new'])
def update_bout(self):
self.bout_id = self.bout_counter.value
self.bout_series = self.bouts_pd.iloc[self.bout_id]
self.x = self.bout_series['waveform'][::self.sub_sample]
self.sxx = self.bout_series['spectrogram'][::self.sub_sample]
def show(self):
#self.fig.clf()
#self.init_fig()
# update
# self.update_bout()
#plot
logger.info('showing')
ax = self.fig_ax['spectrogram']
ax.imshow(self.sxx[::-1], cmap='inferno', aspect='auto')
ax.grid(False)
ax = self.fig_ax['waveform']
ax.plot(self.x)
self.fig.canvas.draw()
#display(self.fig)
#ax.canvas.draw()
self.is_bout.value = bool(self.bout_series['bout_check'])
self.is_bout.observe(self.bout_checked, names='value')
self.m_pick.observe(self.slider_change, names='value')
#display(self.fig)
plt.show()
ms = VizBout(hparams, bpd);
# +
fig, ax_arr = plt.subplots(nrows=2, figsize=(20, 3))
sxx = bpd.iloc[0]['spectrogram'][:, ::10]
ax_arr[0].imshow(sxx[::-1], aspect='auto', cmap='inferno')
|
notebooks/searchbout_s_b1253_21-Copy1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="WLAHnf5g4aLw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="2a8be7cc-e732-4390-b6bd-8099015a8b12" executionInfo={"status": "ok", "timestamp": 1581535914321, "user_tz": -60, "elapsed": 28523, "user": {"displayName": "<NAME>\u0144", "photoUrl": "", "userId": "12996647961383840972"}}
# !pip install datadotworld
# !pip install datadotworld[pandas]
# + id="5jta1-wV4-CH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 55} outputId="f0a6bda6-3fbd-4b24-a56d-e6b7b39b55c6" executionInfo={"status": "ok", "timestamp": 1581535996506, "user_tz": -60, "elapsed": 7388, "user": {"displayName": "<NAME>\u0144", "photoUrl": "", "userId": "12996647961383840972"}}
# !dw configure
# + id="V_uVqsVR3ZCC" colab_type="code" colab={}
from google.colab import drive
import pandas as pd
import numpy as np
import datadotworld as dw
# + id="gfrjl1lD5IRD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 126} outputId="f34fd07e-837d-4e14-f7f5-8afdf32de5bc" executionInfo={"status": "ok", "timestamp": 1581536082027, "user_tz": -60, "elapsed": 8654, "user": {"displayName": "<NAME>\u0144", "photoUrl": "", "userId": "12996647961383840972"}}
drive.mount("/content/drive")
# + id="GN5ue6xj5MsT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="d362541c-f129-46c5-9a95-f70fa0920178" executionInfo={"status": "ok", "timestamp": 1581536137342, "user_tz": -60, "elapsed": 574, "user": {"displayName": "<NAME>\u0144", "photoUrl": "", "userId": "12996647961383840972"}}
# cd "drive/My Drive/Colab Notebooks/dw_matrix"
# + id="Tdu0IwyM5bF9" colab_type="code" colab={}
# !echo 'data' > .gitignore
# + id="A_GbwzU15l9k" colab_type="code" colab={}
# !git add .gitignore
# + id="LziClGMR6Fw7" colab_type="code" colab={}
data = dw.load_dataset('datafiniti/mens-shoe-prices')
# + id="ZHQ0IsfV6V6r" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 126} outputId="f0ea966a-299a-4fa5-cf98-5a44de1a7ec6" executionInfo={"status": "ok", "timestamp": 1581536389387, "user_tz": -60, "elapsed": 1825, "user": {"displayName": "<NAME>\u0144", "photoUrl": "", "userId": "12996647961383840972"}}
df = data.dataframes['7004_1']
df.shape
# + id="ZDpKsPYF6Z-6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 653} outputId="77610742-8ff3-4ebf-8e08-f086720afd8b" executionInfo={"status": "ok", "timestamp": 1581536401800, "user_tz": -60, "elapsed": 604, "user": {"displayName": "<NAME>\u0144", "photoUrl": "", "userId": "12996647961383840972"}}
df.sample(5)
# + id="4wf6cHSX6la1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 230} outputId="74082af2-9057-4aa4-fe18-bdaff668aa70" executionInfo={"status": "ok", "timestamp": 1581536423628, "user_tz": -60, "elapsed": 496, "user": {"displayName": "<NAME>\u0144", "photoUrl": "", "userId": "12996647961383840972"}}
df.columns
# + id="eJG_9RM56qyP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 106} outputId="e4b56143-58ad-46a4-a74f-01942edc7f8a" executionInfo={"status": "ok", "timestamp": 1581536453545, "user_tz": -60, "elapsed": 520, "user": {"displayName": "<NAME>\u0144", "photoUrl": "", "userId": "12996647961383840972"}}
df.prices_currency.unique()
# + id="iRCPqR0Z6yFO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 266} outputId="fc9fe74f-c332-4472-c9ee-231d86ff6701" executionInfo={"status": "ok", "timestamp": 1581536510918, "user_tz": -60, "elapsed": 584, "user": {"displayName": "<NAME>\u0144", "photoUrl": "", "userId": "12996647961383840972"}}
df.prices_currency.value_counts(normalize=True)
# + id="cFbHNFBI68qN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="31fa9619-40c2-48d9-e0a1-26e6135ffd01" executionInfo={"status": "ok", "timestamp": 1581536627676, "user_tz": -60, "elapsed": 544, "user": {"displayName": "<NAME>\u0144", "photoUrl": "", "userId": "12996647961383840972"}}
df_usd = df[ df.prices_currency == 'USD' ].copy()
df_usd.shape
# + id="umBEh-kA7clZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 283} outputId="b2727556-817c-4c06-9d9c-2d1342a5d32a" executionInfo={"status": "ok", "timestamp": 1581536867460, "user_tz": -60, "elapsed": 875, "user": {"displayName": "<NAME>\u0144", "photoUrl": "", "userId": "12996647961383840972"}}
df_usd['prices_amountmin'] = df_usd.prices_amountmin.astype(np.float)
df_usd['prices_amountmin'].hist()
# + id="VYpDhlIL7nFN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="4c4b1e6c-4f54-482b-d5fe-fd8d595e6658" executionInfo={"status": "ok", "timestamp": 1581536988569, "user_tz": -60, "elapsed": 602, "user": {"displayName": "<NAME>\u0144", "photoUrl": "", "userId": "12996647961383840972"}}
filter_max = np.percentile(df_usd['prices_amountmin'], 99)
filter_max
# + id="7CH2lLxa8ib_" colab_type="code" colab={}
df_usd_filter = df_usd[ df_usd['prices_amountmin'] < filter_max ]
# + id="l6YmUu3e81Gc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 283} outputId="aac20ed3-0618-4af7-bae1-4023896c1d06" executionInfo={"status": "ok", "timestamp": 1581537106586, "user_tz": -60, "elapsed": 935, "user": {"displayName": "<NAME>\u0144", "photoUrl": "", "userId": "12996647961383840972"}}
df_usd_filter.prices_amountmin.hist(bins=100)
# + id="gHVnvlt_9NWs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="04548f03-7e47-4a89-c480-931cf6586aa3" executionInfo={"status": "ok", "timestamp": 1581537250207, "user_tz": -60, "elapsed": 1747, "user": {"displayName": "<NAME>\u0144", "photoUrl": "", "userId": "12996647961383840972"}}
!
# + id="nsjCR8Tz90R7" colab_type="code" colab={}
|
matrix_one/day3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import nltk
nltk.download('punkt')
nltk.download('wordnet')
from nltk.stem import WordNetLemmatizer
lemmatizer = WordNetLemmatizer()
import json
import pickle
import numpy as np
from keras.models import Sequential
from keras.layers import Dense, Activation, Dropout
from keras.optimizers import SGD
import random
words=[]
classes = []
documents = []
ignore_words = ['?', '!']
data_file = open('IntentFile').read()
intents = json.loads(data_file)
data_file
for intent in intents['intents']:
for pattern in intent['patterns']:
# take each word and tokenize it
w = nltk.word_tokenize(pattern)
words.extend(w)
# adding documents
documents.append((w, intent['tag']))
# adding classes to our class list
if intent['tag'] not in classes:
classes.append(intent['tag'])
words = [lemmatizer.lemmatize(w.lower()) for w in words if w not in ignore_words]
words = sorted(list(set(words)))
classes = sorted(list(set(classes)))
print (len(documents), "documents")
print (len(classes), "classes", classes)
print (len(words), "unique lemmatized words", words)
pickle.dump(words,open('words.pkl','wb'))
pickle.dump(classes,open('classes.pkl','wb'))
# initializing training data
training = []
output_empty = [0] * len(classes)
for doc in documents:
# initializing bag of words
bag = []
# list of tokenized words for the pattern
pattern_words = doc[0]
# lemmatize each word - create base word, in attempt to represent related words
pattern_words = [lemmatizer.lemmatize(word.lower()) for word in pattern_words]
# create our bag of words array with 1, if word match found in current pattern
for w in words:
bag.append(1) if w in pattern_words else bag.append(0)
# output is a '0' for each tag and '1' for current tag (for each pattern)
output_row = list(output_empty)
output_row[classes.index(doc[1])] = 1
training.append([bag, output_row])
# shuffle our features and turn into np.array
random.shuffle(training)
training = np.array(training)
# create train and test lists. X - patterns, Y - intents
train_x = list(training[:,0])
train_y = list(training[:,1])
print("Training data created")
# Create model - 3 layers. First layer 128 neurons, second layer 64 neurons and 3rd output layer contains number of neurons
# equal to number of intents to predict output intent with softmax
model = Sequential()
model.add(Dense(128, input_shape=(len(train_x[0]),), activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(64, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(len(train_y[0]), activation='softmax'))
# Compile model. Stochastic gradient descent with Nesterov accelerated gradient gives good results for this model
sgd = SGD(lr=0.01, decay=1e-6, momentum=0.9, nesterov=True)
model.compile(loss='categorical_crossentropy', optimizer=sgd, metrics=['accuracy'])
#fitting and saving the model
hist = model.fit(np.array(train_x), np.array(train_y), epochs=200, batch_size=5, verbose=1)
model.save('chatbot_model.h5', hist)
print("model created")
|
Training_Chatbot.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/carlaolivei/data_visualization/blob/main/visualizacao_de_dados_covid_vacina.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="g-ZOF8fkOVgw"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import missingno as msgo
import seaborn as sns
#import geopandas as gpd
import plotly.express as px
# + [markdown] id="cyjdSNaFmcw-"
# <h1> <h3>Sobre o Dataset</h3>
#
# O conjunto de dados contém dados sobre o total da vacinação mundial e novas mortes por covid em 2021.
#
# [Fonte do Dataset](http://www.un.org/en/development/desa/population/migration/data/empirical2/migrationflows.shtml).
# + colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 108} id="Y6WZRbPfOxGk" outputId="ccde0e3a-3002-4d1f-9a83-47f95aabfada"
from google.colab import files
uploaded = files.upload()
for fn in uploaded.keys():
print('User uploaded file "{name}" with length {length} bytes'.format(
name=fn, length=len(uploaded[fn])))
print("Upload concluído!")
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="4eGiUZ5QPEto" outputId="8b906142-2d5b-4c0b-c4db-9e9056cedfb0"
df = pd.read_csv('covid-vaccination-vs-death_ratio.csv')
df.head()
# + colab={"base_uri": "https://localhost:8080/"} id="dayGNVnUPaXt" outputId="03dc0dc8-45a1-4170-f857-f31d1a8beec2"
df.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 300} id="x7K-n3qWPdFg" outputId="687fc920-4eb8-4d47-9d2d-08a62ec55543"
df.describe()
# + [markdown] id="scVntlMiF7Gv"
# <h3>Total de países do dataset</h3>
# + colab={"base_uri": "https://localhost:8080/"} id="wY-aoTwlQITJ" outputId="44ab6439-5abd-41d2-89b1-bf5658cf7ccf"
df['country'].unique().shape
# + [markdown] id="KxaItIk1Fqm2"
# <h3>Total da população</h3>
# + colab={"base_uri": "https://localhost:8080/"} id="XdEM60LyQPB3" outputId="bb84213e-ca5a-4112-c5fb-580e8b8cf07c"
#total da população
total_populacao = df['population'].unique().sum()
total_populacao
# + [markdown] id="bRRqTtLvF3bS"
# <h3>Total da população vacinada completamente contra o covid em 2021</h3>
# + colab={"base_uri": "https://localhost:8080/"} id="BQuop8u-QX42" outputId="6fd014ca-1607-46f6-cf44-401d1c8b26a2"
#total de pessoas completamente vacinadas contra a covid
df_pessoas_vacinadas = df.groupby('country').agg({'people_fully_vaccinated':max})
sum_vac_completa = df_pessoas_vacinadas['people_fully_vaccinated'].sum()
sum_vac_completa
# + [markdown] id="uujRjwEeoGkU"
# <h3> Porcentagem das pessoas completamente vacinadas contra a covid em 2021</h3>
# + colab={"base_uri": "https://localhost:8080/", "height": 361} id="5r6fLDo0n7Pw" outputId="241f8c6d-776d-4c86-a0b2-5a8e07ef5fa3"
plt.figure(figsize=(8, 4),dpi=100) #tamanho do gráfico)
plt.pie(
[total_populacao,sum_vac_completa],
autopct='%1.1f%%', #mostra em percentagem
startangle=90, #ângulo inicial 90°
colors=['slategray','darkseagreen'],
labels=['Total da Populução','Pessoas completamente vacinadas'])
plt.title('Porcentagem das pessoas completamente vacinadas contra a covid em 2021')
plt.show()
# + [markdown] id="A6I0rw55s9KE"
# <h3> Porcentagem das pessoas que tomaram pelo menos uma dose da vacina contra covid em 2021</h3>
# + id="obWd0gBvQn1h"
df_vac_uma_dose = df.groupby('country').agg({'people_vaccinated':max}).reset_index()
sum_vac_uma_dose = df_vac_uma_dose['people_vaccinated'].sum()
# + colab={"base_uri": "https://localhost:8080/", "height": 361} id="JL6ZslGVvLKz" outputId="693dd72f-43d2-419d-b94f-dc337afba547"
plt.figure(figsize=(8, 4),dpi=100) #tamanho do gráfico)
plt.pie(
[total_populacao,sum_vac_uma_dose],
autopct='%1.1f%%', #mostra em percentagem
startangle=90, #ângulo inicial 90°
colors=['slategray','darkseagreen'],
labels=['Total da Populução','Pessoas que tomaram pelo menos uma dose'])
plt.title('Porcentagem das pessoas que tomaram pelo menos uma dose da vacina contra a covid em 2021')
plt.show()
# + [markdown] id="zobZLIM3wLW3"
# <h3> Total de pessoas vacinadas contra covid vs novas mortes em 2021</h3>
# + id="BcHjQQV_Q4mj"
data = df.groupby('date').agg('sum').reset_index()
# + colab={"base_uri": "https://localhost:8080/", "height": 394} id="U5hGMRwcRNr5" outputId="8c14b23d-603e-44f2-8843-332c5257df0d"
x = data['date']
y1 = data['total_vaccinations']
y2 = data['New_deaths']
#linha 1
fig, ax1 = plt.subplots(1,1,figsize=(11,5), dpi= 80)
ax1.plot(x,y1,color='tab:green')
#linha 2
ax2 = ax1.twinx()
ax2.plot(x,y2,color='tab:red')
#configuração da linha 1
ax1.set_xlabel('date',fontsize=15)
ax1.tick_params(axis='x',rotation=0,labelsize=12)
ax1.set_ylabel('Total de vacina',color='tab:green',fontsize=15)
ax1.tick_params(axis='y',rotation=0,labelcolor='tab:green')
ax1.grid(alpha=0.4)
#configuração da linha 2
ax2.set_ylabel('Novas mortes',fontsize=15,color='tab:red')
ax2.tick_params(axis='y',rotation=0,labelcolor='tab:red')
ax2.set_xticks(np.arange(0,len(x),60))#
ax2.set_title('Total de pessoas vacinadas contra covid vs novas mortes em 2021')
plt.show()
# + [markdown] id="YoiyxqKC3EM2"
# <h3>10 Países com vacinação completa contra a covid em 2021</h3>
# + id="EESEcdLRVTsF"
df_paises_vac_completa = df.groupby('country').agg({'people_fully_vaccinated':max})
df_paises_vac_completa.sort_values(['people_fully_vaccinated'], ascending=False, axis=0, inplace=True)
df_paises_vac_completa_top10 = df_paises_vac_completa.head(10)
# + colab={"base_uri": "https://localhost:8080/", "height": 428} id="iUkDkriSEzXS" outputId="a5648ba8-1564-484c-8be3-14c55b334ad7"
ax = df_paises_vac_completa_top10['people_fully_vaccinated'].plot(kind='bar', figsize=(15, 5), color = 'cadetblue')
ax.set_xticklabels(ax.get_xticklabels(), rotation=40, ha="right")
plt.xlabel('Países') # add to x-label to the plot
plt.ylabel('Total de vacinas completas') # add y-label to the plot
plt.title('10 Paíse com maior número de vacinas completas contra covid em 2021') # add title to the plot
plt.show()
# + [markdown] id="oRHMn_t82Y3i"
# <h3>10 Países com maior número de novas mortes por covid em 2021</h3>
# + id="DEjCwicarGj4"
df_paises_mortes = df.groupby('country').sum('New_deaths')
df_paises_mortes.sort_values(['New_deaths'], ascending=False, axis=0, inplace=True)
df_paises_mortes_top10 = df_paises_mortes.head(10)
# + colab={"base_uri": "https://localhost:8080/", "height": 428} id="5-rp_YKanyOo" outputId="55af0e3c-54f5-4749-dea5-6d136ef308ae"
ax = df_paises_mortes_top10['New_deaths'].plot(kind='bar', figsize=(15, 5), color = 'tomato')
ax.set_xticklabels(ax.get_xticklabels(), rotation=40, ha="right")
plt.xlabel('Países') # add to x-label to the plot
plt.ylabel('Novas mortes') # add y-label to the plot
plt.title('10 Paíse com maior número de novas mortes por covid em 2021') # add title to the plot
plt.show()
# + [markdown] id="x6sg85d5k2ZB"
# <h1>Analisando o Brasil</h1>
# + id="e-AEfFd0j1kJ"
df_brasil = df[df['country']=='Brazil']
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="BZphTLl_U5D7" outputId="33f2416d-430c-4810-83c7-5898c8250f1e"
df_brasil.head()
# + colab={"base_uri": "https://localhost:8080/"} id="a7X-OJ2FVsal" outputId="dc981615-52e0-4232-e62b-123ae5c10e6a"
df_brasil.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 300} id="9hihC9VrV6RK" outputId="1155c39d-0278-44cb-9c77-3c58c75fc13e"
df_brasil.describe()
# + [markdown] id="sTM1_eTGGYfC"
# <h3>Total da população do Brasil</h3>
# + colab={"base_uri": "https://localhost:8080/"} id="HTj11tVoWJtk" outputId="bd1d6df2-b07e-4a70-9f56-8fc8705482e3"
total_pop_brasil = df_brasil['population'].unique().sum()
total_pop_brasil
# + [markdown] id="m02-sUQ8Gbp6"
# <h3>Total da população completamente vacinada contra o covid no Brasil em 2021</h3>
# + colab={"base_uri": "https://localhost:8080/"} id="rBn2Sa7gWi3h" outputId="0ddd8c5c-6b99-4ca6-92d2-25375683483c"
df_pais_brasil = df_brasil.agg({'people_fully_vaccinated':max})
sum_vac_completa_brasil = df_pais_brasil['people_fully_vaccinated'].sum()
sum_vac_completa_brasil
# + [markdown] id="_8wvEPVlHciw"
# <h3> Porcentagem das pessoas completamente vacinadas contra a covid no Brasil em 2021</h3>
# + colab={"base_uri": "https://localhost:8080/", "height": 361} id="lTnMUlJbHBxD" outputId="8b475349-8975-4901-c5e0-236985fc6f9d"
plt.figure(figsize=(8, 4),dpi=100) #tamanho do gráfico)
plt.pie(
[total_pop_brasil,sum_vac_completa_brasil],
autopct='%1.1f%%', #mostra em percentagem
startangle=90, #ângulo inicial 90°
colors=['yellowgreen','yellow'],
labels=['Total da Populução Brasil','Pessoas completamente vacinadas'])
plt.title('Porcentagem das pessoas completamente vacinadas contra a covid no Brasil em 2021')
plt.show()
# + [markdown] id="m77P8XXmIVRt"
# <h3> Porcentagem das pessoas que tomaram pelo menos uma dose da vacina contra covid no Brasil em 2021</h3>
# + colab={"base_uri": "https://localhost:8080/"} id="bff3iJe8aNC-" outputId="39c0dfc2-54a6-400c-f1ec-5c0b5ee864e7"
df_pais_brasil_vac = df_brasil.agg({'people_vaccinated':max})
sum_vac_brasil = df_pais_brasil_vac['people_vaccinated'].sum()
sum_vac_brasil
# + colab={"base_uri": "https://localhost:8080/", "height": 361} id="zu3eAAMIIuHM" outputId="e36c67aa-2de6-4600-da28-4c24ee3dc571"
plt.figure(figsize=(8, 4),dpi=100) #tamanho do gráfico)
plt.pie(
[total_pop_brasil,sum_vac_brasil],
autopct='%1.1f%%', #mostra em percentagem
startangle=90, #ângulo inicial 90°
colors=['yellowgreen','yellow'],
labels=['Total da Populução Brasil','Pessoas com pelo menos uma dose da vacina'])
plt.title('Porcentagem das pessoas com pelo menos uma dose da vacina contra covid no Brasil em 2021')
plt.show()
# + [markdown] id="KMhpF-UUJRBF"
# <h3> Total de pessoas vacinadas contra covid vs novas mortes no Brasil em 2021</h3>
# + id="0jeTVZ7LlUKj"
data_brasil = df_brasil.groupby('date').agg('sum').reset_index()
# + colab={"base_uri": "https://localhost:8080/", "height": 411} id="bUQySdmslxNt" outputId="820cd529-d1a4-4c2d-fb8c-2a164b0d3a28"
#preparing data
x = data_brasil['date']
y1 = data_brasil['total_vaccinations']
y2 = data_brasil['New_deaths']
#linha 1
fig, ax1 = plt.subplots(1,1,figsize=(9,5), dpi= 80)
ax1.plot(x,y1,color='tab:green')
#linha 2
ax2 = ax1.twinx()
ax2.plot(x,y2,color='tab:red')
#configuração da linha 1
ax1.set_xlabel('Data',fontsize=15)
ax1.tick_params(axis='x',rotation=0,labelsize=12)
ax1.set_ylabel('Total de Pessoas Vacinadas',color='tab:green',fontsize=15)
ax1.tick_params(axis='y',rotation=0,labelcolor='tab:green')
ax1.grid(alpha=0.4)
#configuração da linha 2
ax2.set_ylabel('Novas Mortes',fontsize=15,color='tab:red')
ax2.tick_params(axis='y',rotation=0,labelcolor='tab:red')
ax2.set_xticks(np.arange(0,len(x),60))#
ax2.set_title('Total de Pessoas Vacinadas vs Novas Mortes no Brasil em 2021')
|
visualizacao_de_dados_covid_vacina.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import dataframe_image as dfi
from matplotlib import rcParams
from matplotlib import pyplot as plt
rcParams['figure.figsize'] = [15, 5]
rcParams['font.size'] = 15
rcParams['axes.spines.top'] = False
rcParams['axes.spines.right'] = False
rcParams['figure.autolayout'] = True
# +
import os, requests
fname = []
for j in range(3):
fname.append('steinmetz_part%d.npz'%j)
url = ['https://osf.io/agvxh/download']
url.append('https://osf.io/uv3mw/download')
url.append('https://osf.io/ehmw2/download')
for i in range(len(url)):
if not os.path.isfile(fname[i]):
try:
r = requests.get(url[i])
except requests.ConnectionError:
print("Data could not download!")
else:
if r.status_code != requests.codes.ok:
print("Data could not download!")
else:
with open(fname[i], "wb") as fid:
fid.write(r.content)
steinmetz_data = np.array([])
for i in range(len(fname)):
steinmetz_data = np.hstack((steinmetz_data, np.load('steinmetz_part%d.npz'%i, allow_pickle=True)['dat']))
|
old_notebooks/spikes_and_behaviour_analyses.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import torch
import torch.utils.data as data
import torchvision.transforms as transforms
import numpy as np
from pytorch_nsynth_lib.nsynth import NSynth
from IPython.display import Audio
import librosa
import librosa.display
import phase_operation
from tqdm import tqdm
import h5py
# -
import spec_ops as spec_ops
import phase_operation as phase_op
import spectrograms_helper as spec_helper
train_data = h5py.File('../data/Nsynth_melspec_IF_pitch.hdf5', 'w')
# audio samples are loaded as an int16 numpy array
# rescale intensity range as float [-1, 1]
toFloat = transforms.Lambda(lambda x: x / np.iinfo(np.int16).max)
# use instrument_family and instrument_source as classification targets
dataset = NSynth(
"../data/nsynth/nsynth-train",
transform=toFloat,
blacklist_pattern=[ "string"], # blacklist string instrument
categorical_field_list=["instrument_family","pitch"])
loader = data.DataLoader(dataset, batch_size=1, shuffle=True)
def expand(mat):
expand_vec = np.expand_dims(mat[:,125],axis=1)
expanded = np.hstack((mat,expand_vec,expand_vec))
return expanded
# +
spec_list=[]
pitch_list=[]
IF_list =[]
mel_spec_list=[]
mel_IF_list=[]
pitch_set =set()
count=0
for samples, instrument_family, pitch, targets in loader:
pitch = targets['pitch'].data.numpy()[0]
if pitch < 24 or pitch > 84:
# print("pitch",pitch)
continue
sample = samples.data.numpy().squeeze()
spec = librosa.stft(sample, n_fft=2048, hop_length = 512)
magnitude = np.log(np.abs(spec)+ 1.0e-6)[:1024]
# print("magnitude Max",magnitude.max(),"magnitude Min",magnitude.min())
angle =np.angle(spec)
# print("angle Max",angle.max(),"angle Min",angle.min())
IF = phase_operation.instantaneous_frequency(angle,time_axis=1)[:1024]
magnitude = expand(magnitude)
IF = expand(IF)
logmelmag2, mel_p = spec_helper.specgrams_to_melspecgrams(magnitude, IF)
# pitch = targets['pitch'].data.numpy()[0]
assert magnitude.shape ==(1024, 128)
assert IF.shape ==(1024, 128)
# spec_list.append(magnitude)
# IF_list.append(IF)
pitch_list.append(pitch)
mel_spec_list.append(logmelmag2)
mel_IF_list.append(mel_p)
pitch_set.add(pitch)
count+=1
if count%10000==0:
print(count)
# -
# train_data.create_dataset("Spec", data=spec_list)
# train_data.create_dataset("IF", data=IF_list)
train_data.create_dataset("pitch", data=pitch_list)
train_data.create_dataset("mel_Spec", data=mel_spec_list)
train_data.create_dataset("mel_IF", data=mel_IF_list)
|
Make Training Data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import logging
import importlib
importlib.reload(logging) # see https://stackoverflow.com/a/21475297/1469195
log = logging.getLogger()
log.setLevel('INFO')
import sys
logging.basicConfig(format='%(asctime)s %(levelname)s : %(message)s',
level=logging.INFO, stream=sys.stdout)
# +
# %%capture
import os
import site
os.sys.path.insert(0, '/home/schirrmr/code/reversible/')
os.sys.path.insert(0, '/home/schirrmr/braindecode/code/braindecode/')
os.sys.path.insert(0, '/home/schirrmr/code/explaining/reversible//')
# %load_ext autoreload
# %autoreload 2
import numpy as np
import logging
log = logging.getLogger()
log.setLevel('INFO')
import sys
logging.basicConfig(format='%(asctime)s %(levelname)s : %(message)s',
level=logging.INFO, stream=sys.stdout)
import matplotlib
from matplotlib import pyplot as plt
from matplotlib import cm
# %matplotlib inline
# %config InlineBackend.figure_format = 'png'
matplotlib.rcParams['figure.figsize'] = (12.0, 1.0)
matplotlib.rcParams['font.size'] = 14
import seaborn
seaborn.set_style('darkgrid')
from reversible2.sliced import sliced_from_samples
from numpy.random import RandomState
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.autograd import Variable
import numpy as np
import copy
import math
import itertools
import torch as th
from braindecode.torch_ext.util import np_to_var, var_to_np
from reversible2.splitter import SubsampleSplitter
from reversible2.view_as import ViewAs
from reversible2.invert import invert
from reversible2.affine import AdditiveBlock
from reversible2.plot import display_text, display_close
from reversible2.bhno import load_file, create_inputs
# -
import sklearn.datasets
X,y = sklearn.datasets.make_moons(200, shuffle=False, noise=1e-4)
plt.figure(figsize=(4,4))
plt.scatter(X[:100,0], X[:100,1])
plt.scatter(X[100:,0], X[100:,1])
train_inputs_a = np_to_var(X[0:100:2], dtype=np.float32)
train_inputs_b = np_to_var(X[1:100:2], dtype=np.float32)
cuda = False
# ### let's try less data
import sklearn.datasets
X,y = sklearn.datasets.make_moons(20, shuffle=False, noise=1e-4)
plt.figure(figsize=(4,4))
plt.scatter(X[:10,0], X[:10,1])
plt.scatter(X[10:,0], X[10:,1])
train_inputs_a = np_to_var(X[0:10:2], dtype=np.float32)
train_inputs_b = np_to_var(X[1:10:2], dtype=np.float32)
cuda = False
plt.figure(figsize=(4,4))
plt.scatter(var_to_np(train_inputs_a)[:,0], var_to_np(train_inputs_a)[:,1])
plt.scatter(var_to_np(train_inputs_b)[:,0], var_to_np(train_inputs_b)[:,1])
# +
from reversible2.distribution import TwoClassDist
from reversible2.blocks import dense_add_block, conv_add_block_3x3
from reversible2.rfft import RFFT, Interleave
from reversible2.util import set_random_seeds
from torch.nn import ConstantPad2d
import torch as th
from reversible2.splitter import SubsampleSplitter
set_random_seeds(2019011641, cuda)
feature_model_a = nn.Sequential(
dense_add_block(2,200),
dense_add_block(2,200),
dense_add_block(2,200),
dense_add_block(2,200),
)
feature_model_b = nn.Sequential(
dense_add_block(2,200),
dense_add_block(2,200),
dense_add_block(2,200),
dense_add_block(2,200),
)
if cuda:
feature_model_a.cuda()
if cuda:
feature_model_b.cuda()
from reversible2.ot_exact import ot_euclidean_loss_for_samples
class_dist_a = TwoClassDist(2,0, [0,1])
if cuda:
class_dist_a.cuda()
class_dist_b = TwoClassDist(2,0, [0,1])
if cuda:
class_dist_b.cuda()
optim_model_a = th.optim.Adam(feature_model_a.parameters())
optim_model_b = th.optim.Adam(feature_model_b.parameters())
optim_dist_a = th.optim.Adam(class_dist_a.parameters(), lr=1e-2)
optim_dist_b = th.optim.Adam(class_dist_b.parameters(), lr=1e-2)
# +
mix_log_stds_a = th.zeros(len(train_inputs_a), 2, requires_grad=True)
mix_log_stds_b = th.zeros(len(train_inputs_b), 2, requires_grad=True)
optim_log_stds_a = th.optim.Adam([mix_log_stds_a], lr=1e-2)
optim_log_stds_b = th.optim.Adam([mix_log_stds_b], lr=1e-2)
from reversible2.gaussian import get_gaussian_log_probs
def get_mix_gauss_samples(n_samples, mix_means, mix_stds):
samples = th.randn(100, *mix_means.shape)
samples = ((samples * mix_stds.unsqueeze(0)) + mix_means.unsqueeze(0))
return samples
# +
n_epochs = 1000
n_samples = 200
for i_epoch in range(n_epochs):
for model, dist, mix_log_stds, optim_stds, optim_model, optim_dist, other_model, other_dist, train_inputs in ((
feature_model_a, class_dist_a, mix_log_stds_a, optim_log_stds_a,
optim_model_a, optim_dist_a, feature_model_b, class_dist_b, train_inputs_a),
(
feature_model_b, class_dist_b, mix_log_stds_b, optim_log_stds_b,
optim_model_b, optim_dist_b, feature_model_a, class_dist_a, train_inputs_b)):
with th.no_grad():
outs = model(train_inputs).detach()
mixed = get_mix_gauss_samples(n_samples, outs, th.exp(mix_log_stds))
mixed = mixed.view(-1, mixed.shape[-1])
translated = other_model(invert(model, mixed))
log_probs = other_dist.get_total_log_prob(0, translated)
nll_loss = -th.mean(log_probs)
optim_stds.zero_grad()
nll_loss.backward()
optim_stds.step()
outs = model(train_inputs)
mixed = get_mix_gauss_samples(n_samples, outs, th.exp(mix_log_stds))
own_log_prob = dist.get_total_log_prob(0, mixed)
nll_loss = -th.mean(own_log_prob)
optim_model.zero_grad()
optim_dist.zero_grad()
#optim_stds.zero_grad()
nll_loss.backward()
optim_model.step()
optim_dist.step()
#optim_stds.step()
if i_epoch % (n_epochs // 20) == 0:
fig, axes = plt.subplots(1,3, figsize=(16,5))
radians = np.linspace(0,2*np.pi,24)
circle_points = np.stack([np.cos(radians), np.sin(radians)], axis=-1)
circle_th = np_to_var(circle_points, device=train_inputs.device, dtype=np.float32)
for i_model, (model, dist, inputs, mix_log_stds, other_model) in enumerate((
(feature_model_a, class_dist_a, train_inputs_a, mix_log_stds_a, feature_model_b),
(feature_model_b, class_dist_b, train_inputs_b, mix_log_stds_b, feature_model_a),)):
outs = model(train_inputs)
for ax in axes[1:]:
ax.scatter(var_to_np(outs)[:,0], var_to_np(outs)[:,1], s=30, label="Real data {:d}".format(i_model),
c=[seaborn.color_palette()[i_model * 2]])
stds = th.exp(mix_log_stds)
circles_per_point = outs.unsqueeze(1) + (circle_th.unsqueeze(0) * stds.unsqueeze(1))
for c in var_to_np(circles_per_point):
axes[1].plot(c[:,0], c[:,1],color=seaborn.color_palette()[i_model * 2],
alpha=0.5, lw=0.5)
translated_circles = other_model(invert(model, circles_per_point.view(-1, circles_per_point.shape[-1]))).view(
*circles_per_point.shape)
for c in var_to_np(translated_circles):
axes[2].plot(c[:,0], c[:,1],color=seaborn.color_palette()[i_model * 2],
alpha=0.5, lw=0.5)
in_circles = invert(model, circles_per_point.view(-1, circles_per_point.shape[-1])).view(
*circles_per_point.shape)
for c in var_to_np(in_circles):
axes[0].plot(c[:,0], c[:,1],color=seaborn.color_palette()[i_model * 2],
alpha=0.5, lw=0.5)
out_circle = (circle_th * dist.get_mean_std(0)[1]) + dist.get_mean_std(0)[0]
axes[1].plot(var_to_np(out_circle)[:,0], var_to_np(out_circle)[:,1],color=seaborn.color_palette()[i_model * 2],
alpha=0.5, lw=0.5)
in_circle = invert(model, out_circle)
axes[0].plot(var_to_np(in_circle)[:,0], var_to_np(in_circle)[:,1],color=seaborn.color_palette()[i_model * 2],
alpha=1, lw=3,)
axes[0].scatter(var_to_np(inputs)[:,0], var_to_np(inputs)[:,1], s=30, label="Real data {:d}".format(i_model),
c=[seaborn.color_palette()[i_model * 2]])
axes[0].set_title("Input space")
axes[0].axis('equal')
axes[1].set_title("Output space")
axes[1].axis('equal')
axes[2].set_title("Output space swapped")
axes[2].axis('equal')
display_close(fig)
for model, dist, train_inputs in [(feature_model_a, class_dist_a, train_inputs_a),
(feature_model_b, class_dist_b, train_inputs_b)]:
fig,axes = plt.subplots(1,2, figsize=(10,4))
rng = RandomState(201904114)
outs = model(train_inputs)
other_X = sklearn.datasets.make_moons(200, shuffle=False, noise=1e-4)[0][:100]
other_ins = np_to_var(other_X, dtype=np.float32)
other_outs = model(other_ins)
axes[0].plot(var_to_np(other_outs[:,0]), var_to_np(other_outs[:,1]), label="All Outputs",
color=seaborn.color_palette()[1])
axes[0].scatter(var_to_np(outs[:,0]), var_to_np(outs[:,1]), s=30, c=[seaborn.color_palette()[0]],
label="Actual data outputs")
axes[0].axis('equal')
axes[0].set_title("Output space")
plt.axis('equal')
samples = dist.get_samples(0, 100)
inverted = invert(model, samples)
axes[1].scatter(var_to_np(inverted)[:,0], var_to_np(inverted)[:,1], s=30, label="Fake/Unknown Samples",
c=[seaborn.color_palette()[1]])
axes[1].scatter(var_to_np(train_inputs)[:,0], var_to_np(train_inputs)[:,1], s=30, label="Real data",
c=[seaborn.color_palette()[0]])
axes[1].legend(bbox_to_anchor=(1,1,0,0))
axes[1].set_title("Input space")
axes[1].axis('equal')
display_close(fig)
# -
out_circle.shape
fig, axes = plt.subplots(1,3, figsize=(16,5))
radians = np.linspace(0,2*np.pi,24)
circle_points = np.stack([np.cos(radians), np.sin(radians)], axis=-1)
circle_th = np_to_var(circle_points, device=train_inputs.device, dtype=np.float32)
for i_model, (model, dist, inputs, mix_log_stds, other_model) in enumerate((
(feature_model_a, class_dist_a, train_inputs_a, mix_log_stds_a, feature_model_b),
(feature_model_b, class_dist_b, train_inputs_b, mix_log_stds_b, feature_model_a),)):
outs = model(train_inputs)
for ax in axes[1:]:
ax.scatter(var_to_np(outs)[:,0], var_to_np(outs)[:,1], s=30, label="Real data {:d}".format(i_model),
c=[seaborn.color_palette()[i_model * 2]])
stds = th.exp(mix_log_stds)
circles_per_point = outs.unsqueeze(1) + (circle_th.unsqueeze(0) * stds.unsqueeze(1))
for c in var_to_np(circles_per_point):
axes[1].plot(c[:,0], c[:,1],color=seaborn.color_palette()[i_model * 2],
alpha=0.5, lw=0.5)
translated_circles = other_model(invert(model, circles_per_point.view(-1, circles_per_point.shape[-1]))).view(
*circles_per_point.shape)
for c in var_to_np(translated_circles):
axes[2].plot(c[:,0], c[:,1],color=seaborn.color_palette()[i_model * 2],
alpha=0.5, lw=0.5)
in_circles = invert(model, circles_per_point.view(-1, circles_per_point.shape[-1])).view(
*circles_per_point.shape)
for c in var_to_np(in_circles):
axes[0].plot(c[:,0], c[:,1],color=seaborn.color_palette()[i_model * 2],
alpha=0.5, lw=0.5)
out_circle = (circle_th * dist.get_mean_std(0)[1]) + dist.get_mean_std(0)[0]
axes[1].plot(var_to_np(out_circle)[:,0], var_to_np(out_circle)[:,1],color=seaborn.color_palette()[i_model * 2],
alpha=0.5, lw=0.5)
in_circle = invert(model, out_circle)
axes[0].plot(var_to_np(in_circle)[:,0], var_to_np(in_circle)[:,1],color=seaborn.color_palette()[i_model * 2],
alpha=0.5, lw=1,)
axes[0].scatter(var_to_np(inputs)[:,0], var_to_np(inputs)[:,1], s=30, label="Real data {:d}".format(i_model),
c=[seaborn.color_palette()[i_model * 2]])
axes[0].set_title("Input space")
axes[0].axis('equal')
axes[1].set_title("Output space")
axes[1].axis('equal')
axes[2].set_title("Output space swapped")
axes[2].axis('equal')
# ### old more data
# +
from reversible2.distribution import TwoClassDist
from reversible2.blocks import dense_add_block, conv_add_block_3x3
from reversible2.rfft import RFFT, Interleave
from reversible2.util import set_random_seeds
from torch.nn import ConstantPad2d
import torch as th
from reversible2.splitter import SubsampleSplitter
set_random_seeds(2019011641, cuda)
feature_model_a = nn.Sequential(
dense_add_block(2,200),
dense_add_block(2,200),
dense_add_block(2,200),
dense_add_block(2,200),
)
feature_model_b = nn.Sequential(
dense_add_block(2,200),
dense_add_block(2,200),
dense_add_block(2,200),
dense_add_block(2,200),
)
if cuda:
feature_model_a.cuda()
if cuda:
feature_model_b.cuda()
from reversible2.ot_exact import ot_euclidean_loss_for_samples
class_dist_a = TwoClassDist(2,0, [0,1])
if cuda:
class_dist_a.cuda()
class_dist_b = TwoClassDist(2,0, [0,1])
if cuda:
class_dist_b.cuda()
optim_model_a = th.optim.Adam(feature_model_a.parameters())
optim_model_b = th.optim.Adam(feature_model_b.parameters())
optim_dist_a = th.optim.Adam(class_dist_a.parameters(), lr=1e-2)
optim_dist_b = th.optim.Adam(class_dist_b.parameters(), lr=1e-2)
# +
mix_log_stds_a = th.zeros(len(train_inputs_a), 2, requires_grad=True)
mix_log_stds_b = th.zeros(len(train_inputs_b), 2, requires_grad=True)
optim_log_stds_a = th.optim.Adam([mix_log_stds_a], lr=1e-2)
optim_log_stds_b = th.optim.Adam([mix_log_stds_b], lr=1e-2)
from reversible2.gaussian import get_gaussian_log_probs
def get_mix_gauss_samples(n_samples, mix_means, mix_stds):
samples = th.randn(100, *mix_means.shape)
samples = ((samples * mix_stds.unsqueeze(0)) + mix_means.unsqueeze(0))
return samples
# +
n_epochs = 1000
n_samples = 200
for i_epoch in range(n_epochs):
for model, dist, mix_log_stds, optim_stds, optim_model, optim_dist, other_model, other_dist, train_inputs in ((
feature_model_a, class_dist_a, mix_log_stds_a, optim_log_stds_a,
optim_model_a, optim_dist_a, feature_model_b, class_dist_b, train_inputs_a),
(
feature_model_b, class_dist_b, mix_log_stds_b, optim_log_stds_b,
optim_model_b, optim_dist_b, feature_model_a, class_dist_a, train_inputs_b)):
with th.no_grad():
outs = model(train_inputs).detach()
mixed = get_mix_gauss_samples(n_samples, outs, th.exp(mix_log_stds))
mixed = mixed.view(-1, mixed.shape[-1])
translated = other_model(invert(model, mixed))
log_probs = other_dist.get_total_log_prob(0, translated)
nll_loss = -th.mean(log_probs)
optim_stds.zero_grad()
nll_loss.backward()
optim_stds.step()
outs = model(train_inputs)
mixed = get_mix_gauss_samples(n_samples, outs, th.exp(mix_log_stds))
own_log_prob = dist.get_total_log_prob(0, mixed)
nll_loss = -th.mean(own_log_prob)
optim_model.zero_grad()
optim_dist.zero_grad()
#optim_stds.zero_grad()
nll_loss.backward()
optim_model.step()
optim_dist.step()
#optim_stds.step()
if i_epoch % (n_epochs // 20) == 0:
fig, axes = plt.subplots(1,3, figsize=(16,5))
radians = np.linspace(0,2*np.pi,24)
circle_points = np.stack([np.cos(radians), np.sin(radians)], axis=-1)
circle_th = np_to_var(circle_points, device=train_inputs.device, dtype=np.float32)
for i_model, (model, dist, inputs, mix_log_stds, other_model) in enumerate((
(feature_model_a, class_dist_a, train_inputs_a, mix_log_stds_a, feature_model_b),
(feature_model_b, class_dist_b, train_inputs_b, mix_log_stds_b, feature_model_a),)):
outs = model(train_inputs)
for ax in axes[1:]:
ax.scatter(var_to_np(outs)[:,0], var_to_np(outs)[:,1], s=30, label="Real data {:d}".format(i_model),
c=[seaborn.color_palette()[i_model * 2]])
stds = th.exp(mix_log_stds)
circles_per_point = outs.unsqueeze(1) + (circle_th.unsqueeze(0) * stds.unsqueeze(1))
for c in var_to_np(circles_per_point):
axes[1].plot(c[:,0], c[:,1],color=seaborn.color_palette()[i_model * 2],
alpha=0.5, lw=0.5)
translated_circles = other_model(invert(model, circles_per_point.view(-1, circles_per_point.shape[-1]))).view(
*circles_per_point.shape)
for c in var_to_np(translated_circles):
axes[2].plot(c[:,0], c[:,1],color=seaborn.color_palette()[i_model * 2],
alpha=0.5, lw=0.5)
in_circles = invert(model, circles_per_point.view(-1, circles_per_point.shape[-1])).view(
*circles_per_point.shape)
for c in var_to_np(in_circles):
axes[0].plot(c[:,0], c[:,1],color=seaborn.color_palette()[i_model * 2],
alpha=0.5, lw=0.5)
axes[0].scatter(var_to_np(inputs)[:,0], var_to_np(inputs)[:,1], s=30, label="Real data {:d}".format(i_model),
c=[seaborn.color_palette()[i_model * 2]])
samples = dist.get_samples(0, 100)
inverted = invert(model, samples)
axes[0].scatter(var_to_np(inverted)[:,0], var_to_np(inverted)[:,1], s=30, label="Fake data {:d}".format(i_model),
c=[seaborn.color_palette()[i_model * 2 + 1]])
for ax in axes[1:]:
ax.scatter(var_to_np(samples)[:,0], var_to_np(samples)[:,1], s=30, label="Fake data {:d}".format(i_model),
c=[seaborn.color_palette()[i_model * 2 + 1]])
axes[0].set_title("Input space")
axes[0].axis('equal')
axes[1].set_title("Output space")
axes[1].axis('equal')
axes[2].set_title("Output space swapped")
axes[2].axis('equal')
display_close(fig)
for model, dist, train_inputs in [(feature_model_a, class_dist_a, train_inputs_a),
(feature_model_b, class_dist_b, train_inputs_b)]:
fig,axes = plt.subplots(1,2, figsize=(10,4))
rng = RandomState(201904114)
outs = model(train_inputs)
other_X = sklearn.datasets.make_moons(200, shuffle=False, noise=1e-4)[0][:100]
other_ins = np_to_var(other_X, dtype=np.float32)
other_outs = model(other_ins)
axes[0].plot(var_to_np(other_outs[:,0]), var_to_np(other_outs[:,1]), label="All Outputs",
color=seaborn.color_palette()[1])
axes[0].scatter(var_to_np(outs[:,0]), var_to_np(outs[:,1]), s=30, c=[seaborn.color_palette()[0]],
label="Actual data outputs")
axes[0].axis('equal')
axes[0].set_title("Output space")
plt.axis('equal')
samples = dist.get_samples(0, 100)
inverted = invert(model, samples)
axes[1].scatter(var_to_np(inverted)[:,0], var_to_np(inverted)[:,1], s=30, label="Fake/Unknown Samples",
c=[seaborn.color_palette()[1]])
axes[1].scatter(var_to_np(train_inputs)[:,0], var_to_np(train_inputs)[:,1], s=30, label="Real data",
c=[seaborn.color_palette()[0]])
axes[1].legend(bbox_to_anchor=(1,1,0,0))
axes[1].set_title("Input space")
axes[1].axis('equal')
display_close(fig)
# +
n_epochs = 2001
n_samples = 200
for i_epoch in range(n_epochs):
for model, dist, mix_log_stds, optim_stds, optim_model, optim_dist, other_model, other_dist, train_inputs in ((
feature_model_a, class_dist_a, mix_log_stds_a, optim_log_stds_a,
optim_model_a, optim_dist_a, feature_model_b, class_dist_b, train_inputs_a),
(
feature_model_b, class_dist_b, mix_log_stds_b, optim_log_stds_b,
optim_model_b, optim_dist_b, feature_model_a, class_dist_a, train_inputs_b)):
with th.no_grad():
outs = model(train_inputs).detach()
mixed = get_mix_gauss_samples(n_samples, outs, th.exp(mix_log_stds))
mixed = mixed.view(-1, mixed.shape[-1])
translated = other_model(invert(model, mixed))
log_probs = other_dist.get_total_log_prob(0, translated)
nll_loss = -th.mean(log_probs)
optim_stds.zero_grad()
nll_loss.backward()
optim_stds.step()
outs = model(train_inputs)
mixed = get_mix_gauss_samples(n_samples, outs, th.exp(mix_log_stds))
own_log_prob = dist.get_total_log_prob(0, mixed)
nll_loss = -th.mean(own_log_prob)
optim_model.zero_grad()
optim_dist.zero_grad()
#optim_stds.zero_grad()
nll_loss.backward()
optim_model.step()
optim_dist.step()
#optim_stds.step()
if i_epoch % (n_epochs // 20) == 0:
display_text("Epoch {:d} of {:d}".format(i_epoch, n_epochs))
display_text("Std 0: {:.1E} {:.1E}".format(*class_dist_a.get_mean_std(0)[1].detach().numpy()))
display_text("Std 1: {:.1E} {:.1E}".format(*class_dist_b.get_mean_std(0)[1].detach().numpy()))
fig, axes = plt.subplots(1,3, figsize=(16,5))
radians = np.linspace(0,2*np.pi,24)
circle_points = np.stack([np.cos(radians), np.sin(radians)], axis=-1)
circle_th = np_to_var(circle_points, device=train_inputs.device, dtype=np.float32)
for i_model, (model, dist, inputs, mix_log_stds, other_model) in enumerate((
(feature_model_a, class_dist_a, train_inputs_a, mix_log_stds_a, feature_model_b),
(feature_model_b, class_dist_b, train_inputs_b, mix_log_stds_b, feature_model_a),)):
outs = model(train_inputs)
for ax in axes[1:]:
ax.scatter(var_to_np(outs)[:,0], var_to_np(outs)[:,1], s=30, label="Real data {:d}".format(i_model),
c=[seaborn.color_palette()[i_model * 2]])
stds = th.exp(mix_log_stds)
circles_per_point = outs.unsqueeze(1) + (circle_th.unsqueeze(0) * stds.unsqueeze(1))
for c in var_to_np(circles_per_point):
axes[1].plot(c[:,0], c[:,1],color=seaborn.color_palette()[i_model * 2],
alpha=0.5, lw=0.5)
translated_circles = other_model(invert(model, circles_per_point.view(-1, circles_per_point.shape[-1]))).view(
*circles_per_point.shape)
for c in var_to_np(translated_circles):
axes[2].plot(c[:,0], c[:,1],color=seaborn.color_palette()[i_model * 2],
alpha=0.5, lw=0.5)
in_circles = invert(model, circles_per_point.view(-1, circles_per_point.shape[-1])).view(
*circles_per_point.shape)
for c in var_to_np(in_circles):
axes[0].plot(c[:,0], c[:,1],color=seaborn.color_palette()[i_model * 2],
alpha=0.5, lw=0.5)
axes[0].scatter(var_to_np(inputs)[:,0], var_to_np(inputs)[:,1], s=30, label="Real data {:d}".format(i_model),
c=[seaborn.color_palette()[i_model * 2]])
samples = dist.get_samples(0, 100)
inverted = invert(model, samples)
axes[0].scatter(var_to_np(inverted)[:,0], var_to_np(inverted)[:,1], s=30, label="Fake data {:d}".format(i_model),
c=[seaborn.color_palette()[i_model * 2 + 1]])
for ax in axes[1:]:
ax.scatter(var_to_np(samples)[:,0], var_to_np(samples)[:,1], s=30, label="Fake data {:d}".format(i_model),
c=[seaborn.color_palette()[i_model * 2 + 1]])
axes[0].set_title("Input space")
axes[0].axis('equal')
axes[1].set_title("Output space")
axes[1].axis('equal')
axes[2].set_title("Output space swapped")
axes[2].axis('equal')
display_close(fig)
for model, dist, train_inputs in [(feature_model_a, class_dist_a, train_inputs_a),
(feature_model_b, class_dist_b, train_inputs_b)]:
fig,axes = plt.subplots(1,2, figsize=(10,4))
rng = RandomState(201904114)
outs = model(train_inputs)
other_X = sklearn.datasets.make_moons(200, shuffle=False, noise=1e-4)[0][:100]
other_ins = np_to_var(other_X, dtype=np.float32)
other_outs = model(other_ins)
axes[0].plot(var_to_np(other_outs[:,0]), var_to_np(other_outs[:,1]), label="All Outputs",
color=seaborn.color_palette()[1])
axes[0].scatter(var_to_np(outs[:,0]), var_to_np(outs[:,1]), s=30, c=[seaborn.color_palette()[0]],
label="Actual data outputs")
axes[0].axis('equal')
axes[0].set_title("Output space")
plt.axis('equal')
samples = dist.get_samples(0, 100)
inverted = invert(model, samples)
axes[1].scatter(var_to_np(inverted)[:,0], var_to_np(inverted)[:,1], s=30, label="Fake/Unknown Samples",
c=[seaborn.color_palette()[1]])
axes[1].scatter(var_to_np(train_inputs)[:,0], var_to_np(train_inputs)[:,1], s=30, label="Real data",
c=[seaborn.color_palette()[0]])
axes[1].legend(bbox_to_anchor=(1,1,0,0))
axes[1].set_title("Input space")
axes[1].axis('equal')
display_close(fig)
# +
display(class_dist_b.get_mean_std(0)[1].detach().numpy())
# -
class_dist_b.get_mean_std(0)
# +
display_text("Epoch {:d} of {:d}".format(i_epoch, n_epochs))
display_text("Std 0: {:.1E} {:.1E}".format(*class_dist_a.get_mean_std(0)[1].detach().numpy()))
display_text("Std 1: {:.1E} {:.1E}".format(*class_dist_b.get_mean_std(0)[1].detach().numpy()))
fig, axes = plt.subplots(1,3, figsize=(16,5))
radians = np.linspace(0,2*np.pi,24)
circle_points = np.stack([np.cos(radians), np.sin(radians)], axis=-1)
circle_th = np_to_var(circle_points, device=train_inputs.device, dtype=np.float32)
for i_model, (model, dist, inputs, mix_log_stds, other_model) in enumerate((
(feature_model_a, class_dist_a, train_inputs_a, mix_log_stds_a, feature_model_b),
(feature_model_b, class_dist_b, train_inputs_b, mix_log_stds_b, feature_model_a),)):
outs = model(train_inputs)
for ax in axes[1:]:
ax.scatter(var_to_np(outs)[:,0], var_to_np(outs)[:,1], s=30, label="Real data {:d}".format(i_model),
c=[seaborn.color_palette()[i_model * 2]])
stds = th.exp(mix_log_stds)
circles_per_point = outs.unsqueeze(1) + (circle_th.unsqueeze(0) * stds.unsqueeze(1))
for c in var_to_np(circles_per_point):
axes[1].plot(c[:,0], c[:,1],color=seaborn.color_palette()[i_model * 2],
alpha=0.5, lw=0.5)
translated_circles = other_model(invert(model, circles_per_point.view(-1, circles_per_point.shape[-1]))).view(
*circles_per_point.shape)
for c in var_to_np(translated_circles):
axes[2].plot(c[:,0], c[:,1],color=seaborn.color_palette()[i_model * 2],
alpha=0.5, lw=0.5)
in_circles = invert(model, circles_per_point.view(-1, circles_per_point.shape[-1])).view(
*circles_per_point.shape)
for c in var_to_np(in_circles):
axes[0].plot(c[:,0], c[:,1],color=seaborn.color_palette()[i_model * 2],
alpha=0.5, lw=0.5)
axes[0].scatter(var_to_np(inputs)[:,0], var_to_np(inputs)[:,1], s=30, label="Real data {:d}".format(i_model),
c=[seaborn.color_palette()[i_model * 2]])
samples = dist.get_samples(0, 100)
inverted = invert(model, samples)
axes[0].scatter(var_to_np(inverted)[:,0], var_to_np(inverted)[:,1], s=30, label="Fake data {:d}".format(i_model),
c=[seaborn.color_palette()[i_model * 2 + 1]])
axes[0].set_title("Input space")
axes[0].axis('equal')
axes[1].set_title("Output space")
axes[1].axis('equal')
axes[2].set_title("Output space swapped")
axes[2].axis('equal')
display_close(fig)
for model, dist, train_inputs in [(feature_model_a, class_dist_a, train_inputs_a),
(feature_model_b, class_dist_b, train_inputs_b)]:
fig,axes = plt.subplots(1,2, figsize=(10,4))
rng = RandomState(201904114)
outs = model(train_inputs)
other_X = sklearn.datasets.make_moons(200, shuffle=False, noise=1e-4)[0][:100]
other_ins = np_to_var(other_X, dtype=np.float32)
other_outs = model(other_ins)
axes[0].plot(var_to_np(other_outs[:,0]), var_to_np(other_outs[:,1]), label="All Outputs",
color=seaborn.color_palette()[1])
axes[0].scatter(var_to_np(outs[:,0]), var_to_np(outs[:,1]), s=30, c=[seaborn.color_palette()[0]],
label="Actual data outputs")
axes[0].axis('equal')
axes[0].set_title("Output space")
plt.axis('equal')
samples = dist.get_samples(0, 100)
inverted = invert(model, samples)
axes[1].scatter(var_to_np(inverted)[:,0], var_to_np(inverted)[:,1], s=30, label="Fake/Unknown Samples",
c=[seaborn.color_palette()[1]])
axes[1].scatter(var_to_np(train_inputs)[:,0], var_to_np(train_inputs)[:,1], s=30, label="Real data",
c=[seaborn.color_palette()[0]])
axes[1].legend(bbox_to_anchor=(1,1,0,0))
axes[1].set_title("Input space")
axes[1].axis('equal')
display_close(fig)
|
notebooks/toy-1d-2d-examples/MoonTwoInvNetsAgain.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.5 64-bit (''base'': conda)'
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import string
data = pd.read_csv("Datasets/Metadata_of_Kaggle dataset/data.csv")
data.head()
# # First View Of Data
data.info()
# # Feature Engineering
# ## Checking if ther are any Null Values in Numerical Features
num_categories = [feature for feature in data.columns if data[feature].dtypes != "O"] # here "O" basically means object which represents string values
data[num_categories].isnull().sum()
# ## Checking if ther are any Null Values in Categorical Features
cat_categories = [feature for feature in data.columns if data[feature].dtypes == "O"]
data[cat_categories].isnull().sum()
# ### Arround 50% of the Description Data is missing but in my knowledge we dont Need Description Column so i will completely remove it.
data = data.drop(columns="description")
# ### Slicing the **ct** Column and making 3 new Columns ["year", "month", "time"] so that we can use this Later in EDA Part.
data[['year','month','time']] = data['ct'].str.split('-',expand=True)
data.head()
# # EDA for The Dataset
# +
medal_distribution = data["DataSetMedals"].value_counts()[1:].reset_index()
plt.figure(figsize=(10,7))
sns.barplot(x="index" ,y="DataSetMedals" ,data=medal_distribution)
plt.title("Medals Distribusion Bar Plot")
plt.xlabel("Medals")
plt.ylabel("Medals Distribution")
#plt.xticks(rotation=90)
plt.show()
# +
year_plot = data["year"].value_counts().reset_index()
plt.figure(figsize=(10,7))
sns.barplot(x="index" ,y="year" ,data=year_plot)
plt.title("Medals Distributed Among The Kagglers Bar Plot")
plt.xlabel("Year")
plt.ylabel("Medals Distributed")
#plt.xticks(rotation=90)
plt.show()
# -
top10_titles = data["title"].value_counts().head(10).reset_index()
plt.figure(figsize=(10,7))
sns.barplot(x="index" ,y="title" ,data=top10_titles)
plt.title("Top 10 Tag Distributed Among The Kagglers Bar Plot")
plt.xlabel("Tag Names")
plt.ylabel("Tag Count")
#plt.xticks(rotation=90)
plt.show()
# +
top10_category = data["key"].value_counts()[1:].head(10).reset_index()
top10_category["index"] = top10_category["index"].str.replace("[", "", regex=True).replace("]", "", regex=True).replace("'", "", regex=True)
plt.figure(figsize=(20,7))
sns.barplot(x="index" ,y="key" ,data=top10_category)
plt.title("Top 10 Category Distribution Among The Kagglers Bar Plot")
plt.xlabel("Category Name")
plt.ylabel("Category Count")
plt.xticks(rotation=80)
plt.show()
# -
#
# # Results Of **EDA**
#
# #### 1) According to the Medal Distribution bar plot Most of the users Got Bronze Medal than Silver and Gold.
# #### 2) According to the Data Year 2020 was the time when The users were wost active and so the Medal Created/Distributed in large numbers and second place comes 2021.
# #### 3) According to the Data the most trending Title Tags are **Dataset** and **Titanic**.
# #### 4) According to the Data the most Common Categories in which Most of the Medals were Distributed Are Business, Earth and Temperature, Art and Entertainment.
# ### So in order to get the Maximum Numbers of Medals you Should use Tags like **Dataset** and **Titanic** wherever possible. And you can try to Create more Kernal, Datasets or take part in Competition which falls in the Category of **Business**, **Earth and Temperature**, **Art and Entertainment** Because these Are *Hot Topics*.
#
# ## Author
#
# - [@everydaycodings(Kaggle)](https://www.kaggle.com/everydaycodings)
# - [@everydaycodings(Github)](https://github.com/everydaycodings)
#
# #### If You Lived this Kernal, Please Don't Forget To Give a UP Vote To It, This keeps me motivated to Make more Such Kernels.
|
Metadata_of_Kaggle_dataset.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="dKHHWgRDH9O6"
# !pip install pyyaml==5.1
import torch
TORCH_VERSION = ".".join(torch.__version__.split(".")[:2])
CUDA_VERSION = torch.__version__.split("+")[-1]
print("torch: ", TORCH_VERSION, "; cuda: ", CUDA_VERSION)
# Install detectron2 that matches the above pytorch version
# See https://detectron2.readthedocs.io/tutorials/install.html for instructions
# !pip install detectron2 -f https://dl.fbaipublicfiles.com/detectron2/wheels/$CUDA_VERSION/torch$TORCH_VERSION/index.html
# If there is not yet a detectron2 release that matches the given torch + CUDA version, you need to install a different pytorch.
# exit(0) # After installation, you may need to "restart runtime" in Colab. This line can also restart runtime
# + id="3yFIRqlhIIen"
# !nvidia-smi
# !nvcc --version
# + id="u3g2phu2IK1u"
from google.colab import drive
drive.mount('/content/gdrive')
project_path = '/content/gdrive/MyDrive/madeira'
images_path = f'{project_path}'
# + id="EHqekWE4ItfF" executionInfo={"status": "ok", "timestamp": 1648162484937, "user_tz": 180, "elapsed": 1083, "user": {"displayName": "oi oi", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "03679624378915265090"}}
# Some basic setup:
# Setup detectron2 logger
import detectron2
from detectron2.utils.logger import setup_logger
setup_logger()
# import some common libraries
import numpy as np
import os, json, cv2, random
import datetime
from google.colab.patches import cv2_imshow
# import some common detectron2 utilities
from detectron2 import model_zoo
from detectron2.engine import DefaultPredictor
from detectron2.config import get_cfg
from detectron2.utils.visualizer import Visualizer
from detectron2.data import MetadataCatalog, DatasetCatalog
from detectron2.engine import DefaultTrainer
from detectron2.config import get_cfg
from detectron2.checkpoint import DetectionCheckpointer
# + id="1KQ_cA0nIwVt"
from detectron2.data.datasets import register_coco_instances
register_coco_instances("train_coco", {}, f"{images_path}/train_coco/annotations.json", f'{images_path}/train_coco')
train_dataset = DatasetCatalog.get("train_coco")
train_metadata = MetadataCatalog.get("train_coco")
register_coco_instances("test_coco", {}, f"{images_path}/test_coco/annotations.json", f'{images_path}/test_coco')
test_dataset = DatasetCatalog.get("test_coco")
test_metadata = MetadataCatalog.get("test_coco")
# + id="erWgy6KTIyJ9" executionInfo={"status": "ok", "timestamp": 1648162490669, "user_tz": 180, "elapsed": 274, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "03679624378915265090"}}
from detectron2.data import DatasetCatalog, MetadataCatalog, build_detection_test_loader, build_detection_train_loader
from detectron2.data import detection_utils as utils
import detectron2.data.transforms as T
import copy
def custom_mapper(dataset_dict):
dataset_dict = copy.deepcopy(dataset_dict)
image = utils.read_image(dataset_dict["file_name"], format="BGR")
transform_list = [
T.RandomBrightness(0.9, 1.1),
T.RandomContrast(0.9, 1.1),
T.RandomSaturation(0.9, 1.1),
T.RandomFlip(prob=0.5, horizontal=False, vertical=True),
T.RandomFlip(prob=0.5, horizontal=True, vertical=False),
T.RandomCrop("relative", (0.4, 0.4))
]
image, transforms = T.apply_transform_gens(transform_list, image)
dataset_dict["image"] = torch.as_tensor(image.transpose(2, 0, 1).astype("float32"))
annos = [
utils.transform_instance_annotations(obj, transforms, image.shape[:2])
for obj in dataset_dict.pop("annotations")
if obj.get("iscrowd", 0) == 0
]
instances = utils.annotations_to_instances(annos, image.shape[:2])
dataset_dict["instances"] = utils.filter_empty_instances(instances)
return dataset_dict
class AugTrainer(DefaultTrainer):
@classmethod
def build_train_loader(cls, cfg):
return build_detection_train_loader(cfg, mapper=custom_mapper)
# + id="Ijk6Ao-SI-lw"
# If first training
cfg = get_cfg()
cfg.OUTPUT_DIR = f'{project_path}/model/best'
os.makedirs(cfg.OUTPUT_DIR, exist_ok=True)
cfg.merge_from_file(model_zoo.get_config_file('COCO-InstanceSegmentation/mask_rcnn_R_50_FPN_3x.yaml'))
cfg.MODEL.WEIGHTS = model_zoo.get_checkpoint_url("COCO-InstanceSegmentation/mask_rcnn_R_50_FPN_3x.yaml")
cfg.DATASETS.TRAIN = ("train_coco",)
cfg.DATASETS.TEST = ()
cfg.DATALOADER.NUM_WORKERS = 2
cfg.SOLVER.IMS_PER_BATCH = 6
cfg.SOLVER.BASE_LR = 0.002
cfg.SOLVER.MAX_ITER = (300)
cfg.MODEL.SEM_SEG_HEAD.NUM_CLASSES = 2
cfg.MODEL.ROI_HEADS.NUM_CLASSES = 2
cfg.MODEL.ROI_HEADS.BATCH_SIZE_PER_IMAGE = (128)
trainer = AugTrainer(cfg)
checkpointer = DetectionCheckpointer(trainer.model, save_dir=cfg.OUTPUT_DIR)
# + id="eKPRhodhJXyf"
# Train
import os
cfg.SOLVER.MAX_ITER = (300)
trainer.resume_or_load(resume=False)
trainer.resume_or_load()
trainer.train()
# + colab={"base_uri": "https://localhost:8080/", "height": 604} id="8R54iVK-JtXO" executionInfo={"status": "ok", "timestamp": 1648163737496, "user_tz": 180, "elapsed": 3083, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "03679624378915265090"}} outputId="408e730e-bec4-4cf2-c9a5-8b46647e06de"
from detectron2.utils.visualizer import ColorMode
cfg.MODEL.WEIGHTS = os.path.join(cfg.OUTPUT_DIR, "model_final.pth")
cfg.MODEL.ROI_HEADS.SCORE_THRESH_TEST = 0.5
predictor = DefaultPredictor(cfg)
#im = cv2.imread(f'{images_path}/de frente.jpeg')
im = cv2.imread(f'{images_path}/test_coco/JPEGImages/1_3.jpg')
print(test_metadata.thing_classes)
outputs = predictor(im) # format is documented at https://detectron2.readthedocs.io/tutorials/models.html#model-output-format
v = Visualizer(im[:, :, ::-1],
metadata=train_metadata,
scale=0.5,
instance_mode=ColorMode.IMAGE_BW # remove the colors of unsegmented pixels. This option is only available for segmentation models
)
out = v.draw_instance_predictions(outputs["instances"].to("cpu"))
cv2_imshow(out.get_image()[:, :, ::-1])
|
Segmentacao de instancia - toras.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # What are `TargetPixelFile` objects?
# Target Pixel Files (TPFs) are a file common to Kepler/K2 and the TESS mission. They contain movies of the pixel data centered on a single target star.
#
# TPFs can be thought of as stacks of images, with one image for every timestamp the telescope took data. Each timestamp is referred to as a **cadence**. These images are cut out 'postage stamps' of the full observation to make them easier to work with.
#
# TPFs are given in FITS files, which you can read more about [here](https://fits.gsfc.nasa.gov/fits_primer.html). *Lightkurve* includes tools for you to work directly with these files easily and intuitively.
#
# In this tutorial we'll cover the basics of working with TPFs. In *lightkurve* there are classes to work with each mission. For example `KeplerTargetPixelFile` deals with data from the Kepler (and K2) mission. `TessTargetPixelFile` deals with data from the TESS mission. We'll use a Kepler TPF as an example.
#
# To load a `KeplerTargetPixelFile` from a local path or remote url, simply call Lightkurve's `read` function using the location of the file as the parameter:
import lightkurve as lk
tpf = lk.read("https://archive.stsci.edu/pub/kepler/target_pixel_files/0069/006922244/kplr006922244-2010078095331_lpd-targ.fits.gz")
# You can also search for the url automatically using the `search_targetpixelfile()` function. This will search for the right file in the [MAST data archive](https://archive.stsci.edu/kepler/) which holds all of the Kepler and K2 data.
# In this case we want the Target Pixel File with Kepler ID 6922244 for Quarter 4 (Kepler's observations were split into quarters of a year):
from lightkurve import search_targetpixelfile
tpf = search_targetpixelfile('KIC 6922244', quarter=4).download()
# You can also pass the name of the target or its astronomical coordinates as a parameter to `search_targetpixelfile()`.
#
# The above code has created a variable named `tpf` which is a Python object of type `KeplerTargetPixelFile`:
tpf
# We can access lots of meta data using this object in a simple way. For example, we can find the mission name, and the quarter that the data was taken in by typing the following:
tpf.mission
tpf.quarter
# You can find the full list of properties in the [API documentation](https://docs.lightkurve.org/api/lightkurve.targetpixelfile.KeplerTargetPixelFile.html#lightkurve.targetpixelfile.KeplerTargetPixelFile) on this object.
# The most interesting data in a `KeplerTargetPixelFile` object are the `flux` and `time` values which give access to the brightness of the observed target over time. You can access the timestamps of the observations using the `time` property:
tpf.time
# By default, `time` is in the Kepler-specific *Barycentric Kepler Julian Day* format (BKJD). You can easily convert this into [AstroPy Time objects](http://docs.astropy.org/en/stable/time/) using the `astropy_time` property:
tpf.astropy_time
# In turn, this gives you access to human-readable ISO timestamps using the `astropy_time.iso` property:
tpf.astropy_time.iso
# **Beware:** these timestamps are in the Solar System Barycentric frame (TDB) and do not include corrections for light travel time or leap seconds. To use a different time scale, such as the Earth-centered UTC system, you can use [AstroPy's time scale conversion features](http://docs.astropy.org/en/stable/time/#time-scale). For example:
tpf.astropy_time.utc.iso
# Next, let's look at the actual image data, which is available via the `flux` property:
tpf.flux.shape
# The `flux` data is a 4116x5x5 array in units electrons/second. The first axis is the time axis, and the images themselves are 5 pixels by 5 pixels. You can use the `plot` method on the `KeplerTargetPixelFile` object to view the data. (By default, this will show just one cadence of the data. But you can pass the cadence you want to look at to the `frame` keyword if you would like to check a particular flux point for thruster firings, cosmic rays or asteroids.)
# %matplotlib inline
tpf.plot(frame=0);
# The values shown in this image are also directly accessible as an array:
tpf.flux[0]
# You can use normal `numpy` methods on these to find the shape, mean etc!
# We can now turn this Target Pixel File into a light curve, with a single flux value for every time value. Each of the pixels are 4 arcseconds across. The point spread function (PSF) of the telescope causes the light from the star fall onto several different pixels, which can be seen in the image above. Because of this spreading, we have to sum up many pixels to collect all the light from the source. To do this we sum up all the pixels in an **aperture**. An aperture is a pixel mask, where we take only the pixels related to the target.
# The *Kepler* pipeline adds an aperture mask to each target pixel file. This aperture determines which pixels are summed to create a 1-D light curve of the target. There are some science cases where you might want to create a different aperture. For example, there may be a nearby contaminant or you may want to measure the background.
#
# The standard pipeline aperture is easily accessed in a `KeplerTargetPixelFile` object using `tpf.pipeline_mask`, which is a boolean array:
tpf.pipeline_mask
# We can also plot this aperture over the target pixel file above to see if the flux of the star is all contained within the aperture.
tpf.plot(aperture_mask=tpf.pipeline_mask);
# Now that we have the aperture we can create a Simple Aperture Photometry light curve in the next tutorial.
# Finally, note that you can inspect all the raw metadata of the target by taking a look at the 'header' of the FITS file, which contains information about the data set. Let's just print the first 10 lines:
tpf.get_header()[:10]
# We can look at the values in the second extension of the fits file by accessing the AstroPy FITS `HDUList` object. For example, to look at all the column titles:
tpf.hdu[1].header['TTYPE*']
|
docs/source/tutorials/01-target-pixel-files.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# %matplotlib inline
import seaborn as sns
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.ensemble import IsolationForest
from sklearn.ensemble import RandomForestClassifier
from Utils.utils import conf_matrix, read_mat
# default plot settings
plt.rcParams['figure.figsize'] = [12, 10]
plt.rcParams['axes.facecolor'] = 'lightskyblue'
plt.rcParams['axes.titlepad'] = 35.0
rng = np.random.RandomState(42)
# -
customPalette = ['#630C3A', '#39C8C6', '#D3500C', '#FFB139']
sns.set_palette(customPalette)
sns.palplot(customPalette)
def plot_data(dframe,x='x', y='y',col='label'):
sns.lmplot(data=dframe, x=x, y=y, col=col,
fit_reg=False, legend=True, legend_out=True)
def model_fitting(df, xidx, ylabel,test_size, random_state, model1, model2):
Data = df.iloc[:,xidx].values
Target = df[ylabel].values
X_train, X_test, y_train, y_test = train_test_split(Data, Target, test_size=test_size, random_state=random_state, stratify=Target)
print('Training Features Shape:', X_train.shape)
print('Training Labels Shape:', y_train.shape)
print('Testing Features Shape:', X_test.shape)
print('Testing Labels Shape:', y_test.shape)
model1.fit(X_train)
model2.fit(X_train,y_train)
return ((X_test, y_test), model1, model2)
def plot_prep(data_splits, model, actual_inlier, model_inlier, model_name="model"):
X_test, y_test = data_splits
y_pred = model.predict(X_test)
y_test_new = []
for yt in y_test.tolist():
if yt==actual_inlier:
y_test_new.append('Good')
else:
y_test_new.append('Bad')
y_pred_new = []
for yt in y_pred.tolist():
if yt==model_inlier:
y_pred_new.append('Good')
else:
y_pred_new.append('Bad')
data = np.stack((X_test[:, 2], X_test[:,1], y_test_new, y_pred_new)).T
new_df = pd.DataFrame(data, columns=['x', 'y', '{}.actual'.format(model_name), '{}.predicted'.format(model_name)])
new_df['x'] = new_df['x'].astype('float')
new_df['y'] = new_df['y'].astype('float')
return (new_df, 'x', 'y', ('{}.actual'.format(model_name), '{}.predicted'.format(model_name)))
def conf_prep(df, model_name):
y_act = df['{}.actual'.format(model_name)].values.tolist()
y_pred = df['{}.predicted'.format(model_name)].values.tolist()
y_act = [0 if x=='Good' else 1 for x in y_act]
y_pred = [0 if x=='Good' else 1 for x in y_pred]
return (y_act, y_pred, [0,1])
# **Case - Dataset #1**<br/>
# The original Statlog (Landsat Satellite) dataset from UCI machine learning repository is a multi-class classification dataset. Here, the training and test data are combined. The smallest three classes, i.e. 2, 4, 5 are combined to form the outliers class, while all the other classes are combined to form an inlier class.
#
# *Description: X = Multi-dimensional point data, y = labels (1 = outliers, 0 = inliers)*
filepath = 'resources/anomaly_datasets/satellite.mat'
df_satellite = read_mat(filepath)
df_satellite.head()
# +
'''Initialize Models Here'''
clf1 = IsolationForest(behaviour='new', max_samples=100,
random_state=rng, contamination=0.3)
rfc1 = RandomForestClassifier(n_estimators=1000, random_state=rng, max_features=1.,
class_weight='balanced', n_jobs=-1)
features = range(0,df_satellite.shape[1]-1)
data_splits, clf1_trained, rfc1_trained = model_fitting(df_satellite, features, 'label', 0.20, rng, clf1, rfc1)
# -
df1_iso, plot_x, plot_y, cols = plot_prep(data_splits,clf1_trained, 0, 1, "iforest")
plot_data(df1_iso, x=plot_x, y=plot_y, col=cols[0])
plot_data(df1_iso, x=plot_x, y=plot_y, col=cols[1])
df1_rf, plot_x, plot_y, cols = plot_prep(data_splits,rfc1_trained, 0, 0, "rforest")
plot_data(df1_rf, x=plot_x, y=plot_y, col=cols[0])
plot_data(df1_rf, x=plot_x, y=plot_y, col=cols[1])
# +
test1_iso, pred1_iso, labels1_iso = conf_prep(df1_iso, "iforest")
test1_rf, pred1_rf, labels1_ref = conf_prep(df1_rf, "rforest")
fig = plt.figure(figsize=(16,12))
conf_matrix(test1_iso, pred1_iso, fig, labels=labels1_iso, z=1 )
conf_matrix(test1_rf, pred1_rf, fig, labels= labels1_ref, model_name="Random Forest", z=2)
fig.subplots_adjust(wspace=0.8)
plt.show()
# -
# **Case - Dataset #2**<br/>
# A very imbalanced dataset for ICU discharge of patients. A bad discharge is when the patient had to be readmitted within 90 days of discharge from the ICU. A very few cases of bad discharge are present in the dataset which indicates that such cases can simply be anomalies.
#
# *Description: X = Multi-dimensional point data, y = labels (1 = outliers, 0 = inliers)*
'''Isolation and Random Forest on really bad dataset'''
path = 'resources/pacemed/'
df_icu = pd.read_csv(path+'cleaned.csv',index_col=None, sep=',')
df_icu['bad_discharge'] = df_icu['bad_discharge'].astype('int')
df_icu.head()
# +
'''Initialize Models Here'''
clf2 = IsolationForest(behaviour='new', max_samples=100,
random_state=rng, contamination=0.1)
rfc2 = RandomForestClassifier(n_estimators=1000, random_state=rng, max_features=1.,
class_weight='balanced', n_jobs=-1)
features = range(1,df_icu.shape[1]-1)
data_splits, clf2_trained, rfc2_trained = model_fitting(df_icu, features, 'bad_discharge', 0.20, rng, clf2, rfc2)
# -
df2_iso, plot_x, plot_y, cols = plot_prep(data_splits,clf2_trained, 0, 1, "iforest")
plot_data(df2_iso, x=plot_x, y=plot_y, col=cols[0])
plot_data(df2_iso, x=plot_x, y=plot_y, col=cols[1])
df2_rf, plot_x, plot_y, cols = plot_prep(data_splits,rfc2_trained, 0, 0, "rforest")
plot_data(df2_rf, x=plot_x, y=plot_y, col=cols[0])
plot_data(df2_rf, x=plot_x, y=plot_y, col=cols[1])
# +
test2_iso, pred2_iso, labels2_iso = conf_prep(df2_iso, "iforest")
test2_rf, pred2_rf, labels2_ref = conf_prep(df2_rf, "rforest")
fig = plt.figure(figsize=(16,12))
conf_matrix(test2_iso, pred2_iso, fig, labels=labels2_iso, z=1 )
conf_matrix(test2_rf, pred2_rf, fig, labels= labels2_ref, model_name="Random Forest", z=2)
fig.subplots_adjust(wspace=0.8)
plt.show()
# -
# **Case - Dataset #3**<br/>
# The original ForestCover/Covertype dataset from UCI machine learning repository is a multiclass classification dataset. It is used in predicting forest cover type from cartographic variables only (no remotely sensed data). This study area includes four wilderness areas located in the Roosevelt National Forest of northern Colorado. These areas represent forests with minimal human-caused disturbances, so that existing forest cover types are more a result of ecological processes rather than forest management practices. This dataset has 54 attributes (10 quantitative variables, 4 binary wilderness areas and 40 binary soil type variables). Here, outlier detection dataset is created using only 10 quantitative attributes. Instances from class 2 are considered as normal points and instances from class 4 are anomalies. The anomalies ratio is 0.9%. Instances from the other classes are omitted.
#
# *Description: X = Multi-dimensional point data, y = labels (1 = outliers, 0 = inliers)*
#
#
#
filepath = 'resources/anomaly_datasets/cover.mat'
df_cover = read_mat(filepath)
df_cover.head()
# +
'''Initialize Models Here'''
clf3 = IsolationForest(behaviour='new', max_samples=100,
random_state=rng, contamination=0.009)
rfc3 = RandomForestClassifier(n_estimators=1000, random_state=rng, max_features=1.,
class_weight='balanced', n_jobs=-1)
features = range(0,df_cover.shape[1]-1)
data_splits, clf3_trained, rfc3_trained = model_fitting(df_cover, features, 'label', 0.20, rng, clf3, rfc3)
# -
df3_iso, plot_x, plot_y, cols = plot_prep(data_splits,clf3_trained, 0, 1, "iforest")
plot_data(df3_iso, x=plot_x, y=plot_y, col=cols[0])
plot_data(df3_iso, x=plot_x, y=plot_y, col=cols[1])
df3_rf, plot_x, plot_y, cols = plot_prep(data_splits,rfc3_trained, 0, 0, "rforest")
plot_data(df3_rf, x=plot_x, y=plot_y, col=cols[0])
plot_data(df3_rf, x=plot_x, y=plot_y, col=cols[1])
# +
test3_iso, pred3_iso, labels3_iso = conf_prep(df3_iso, "iforest")
test3_rf, pred3_rf, labels3_ref = conf_prep(df3_rf, "rforest")
fig = plt.figure(figsize=(16,12))
conf_matrix(test3_iso, pred3_iso, fig, labels=labels3_iso, z=1 )
conf_matrix(test3_rf, pred3_rf, fig, labels= labels3_ref, model_name="Random Forest", z=2)
fig.subplots_adjust(wspace=0.8)
plt.show()
# -
|
tutorials/ML_tutorials/Ensemble _Trees_Comparison.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import cv2
import os
import matplotlib.pyplot as plt
import numpy as np
from IPython.display import clear_output
# %matplotlib inline
# +
# cap = cv2.VideoCapture(0)
# success,prevFrame = cap.read()
# sizeOfFrame = prevFrame.size
# print(prevFrame.shape)
# counter = 0
# while(success):
# success, curFrame = cap.read()
# dist_euclidean = 0
# if cv2.waitKey(33) == ord('q'):
# break
# cv2.imshow('frame',prevFrame)
# euclideanDist = sum(sum(np.sqrt(sum((curFrame-prevFrame)^2))))
# euclideanDist /= sizeOfFrame
# print((sum((curFrame-prevFrame)^2)).shape)
# print(euclideanDist)
# if(euclideanDist>0.05):
# print('IamReady')
# prevFrame = curFrame
# cap.release()
# cv2.destroyAllWindows()
# +
# cap = cv2.VideoCapture('./data/inputVideo/jump_input.mp4')
# success,prevFrame = cap.read()
# sizeOfFrame = prevFrame.size
# print(prevFrame.shape)
# frame_array = []
# height, width, layers = prevFrame.shape
# size = (width, height)
# counter = 0
# while(success):
# success, curFrame = cap.read()
# dist_euclidean = 0
# if(not success):
# break
# euclideanDist = sum(sum(np.sqrt(sum((curFrame-prevFrame)^2))))
# euclideanDist /= sizeOfFrame
# frame_array.append(prevFrame)
# if(euclideanDist<0.095):
# avgFrame = prevFrame/2+curFrame/2
# frame_array.append(avgFrame)
# print('IamReady')
# if(count>500):
# break
# counter += 1
# prevFrame = curFrame
# plt.imshow(prevFrame)
# out = cv2.VideoWriter('./data/outputVideo/4.avi', cv2.VideoWriter_fourcc(*'MJPG'),48, size)
# # out = cv2.VideoWriter('./data/outputVideo/4.avi', cv2.VideoWriter_fourcc(*'DIVX'),48, size)
# # 'M','J','P','G'
# l = len(frame_array)
# for i in range(l):
# out.write(np.uint8(frame_array[i]))
# out.release()
# print(count)
# -
frame_array
# +
cap = cv2.VideoCapture('./inputVideo/jump_input.mp4')
success,prevFrame = cap.read()
sizeOfFrame = prevFrame.size
print(prevFrame.shape)
frame_array = []
height, width, layers = prevFrame.shape
size = (width, height)
out = cv2.VideoWriter('./data/outputVideo/8.avi', cv2.VideoWriter_fourcc(*'DIVX'),48,size)
counter = 0
while(success):
success, curFrame = cap.read()
dist_euclidean = 0
if(not success):
break
euclideanDist = sum(sum(np.sqrt(sum((curFrame-prevFrame)^2))))
euclideanDist /= sizeOfFrame
out.write(np.uint8(prevFrame))
if(euclideanDist<0.095):
avgFrame = prevFrame/2+curFrame/2
out.write(np.uint8(avgFrame))
print("I am inserted")
else:
print("I am not inserted")
prevFrame = curFrame
plt.imshow(prevFrame)
out.release()
# -
|
.ipynb_checkpoints/FrameInsert_eculidean-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <div align="right"><i><NAME><br>12 August 2019</i></div>
#
# # Tracking Trump: Electoral Votes Edition
#
# Sites such as [RealClearPolitics](https://www.realclearpolitics.com/epolls/other/president_trump_job_approval-6179.html), [538](https://projects.fivethirtyeight.com/trump-approval-ratings/), and [Mourning Consult](https://morningconsult.com/tracking-trump/) track presidential approval ratings (currently about 43% approval and 53% disapproval for a net -10%). Do approval ratings predict election results? There are three big caveats:
#
# 1. Today is not election day 2020.
#
# 2. Approval polls are not votes.
#
# 3. Popular votes are not electoral votes.
#
# We can't be conclusive about the first two points, but this notebook can use state-by-state approval polls to
# compute expected electoral votes, under the assumption that Trump wins the electoral votes of states he has positive net approval (and for the purposes of computation we'll count half the electoral votes for states where approval exactly equals disapproval).
#
#
# # TL;DR for policy wonks
#
# As of August 2019, Trump would expect **172 electoral votes** under these assumptions (you need **270** to win). If you list states in order of his approval, the key turning-point state is Pennsylvania; he'd need to win that and every state in which he is more popular. He currently is **7% behind in Pennsylvania**; we call that the *margin*.
#
#
# # The details for data science nerds
#
# We don't know who else will be on the ballot and what their approval levels will be, we don't know if there is systematic bias in the polling data, we don't know how many people will vote for a candidate they disapprove of or against a candidate they approve of, and we don't know who will decline to vote.
# I have five ways of understanding the fluidity of the situation:
#
# - **Undecided**: If many voters are undecided, the net approval could change a lot. So I track the number of states for which at least 5% of voters are undecided. At the inauguration in 2017, all 51 states (including DC) had at least 5% undecided; now there is only one such state (Alaska). Overall 4% of voters are undecided. Most people have made up their mind. In [one poll](https://www.pbs.org/newshour/politics/57-percent-of-voters-say-they-wont-support-trump-in-2020) 57% said they would definitely not vote for Trump in 2020; other polls have this in the 50% to 55% range.
#
# - **Variance**: How much are voters changing their minds from month to month in each state? I track the standard deviation, 𝝈, of the net approval for each state over the last 12 months.
#
# - **Movement**: What's the most a state's net approval could be expected to move, due to random fluctuations (that is, assuming there is no big event that changes people's minds)? I define the maximum expected **movement** of a state as 1/5 of the undecided voters (i.e. assume the undecided voters broke 60/40 one way or the other) plus 2 standard deviations in the net approval over the last 12 months.
#
# - **Swing state**: I define a swing state as one whose maximum expected movement is greater than the absolute value of the net approval. There are 13 such states now; if Trump won them all, he would still lose the election with only 237 electoral votes.
#
# - **Margin**: Suppose a future event swings voters in one direction, across the board in all the key states. How much of a swing would be necessary to change the election outcome? We call that the **margin**. Today **Trump's margin is 7%:** if he got 7% more votes in 8 key states he would be over 270 electoral votes. (This could come, for example, by convincing 3% of undecided voters to break for him at a 2 to 1 ratio, and then convincing 3% of disapproving voters to switch to approving.)
#
# # Data and Code
#
# First fetch the state-by-state, month-by-month approval data from the **[Tracking Trump](https://morningconsult.com/tracking-trump/)** web page at *Morning Consult*
# and cache it locally:
# ! curl -o evs.html https://morningconsult.com/tracking-trump-2/
# Now some imports:
# %matplotlib inline
import matplotlib.pyplot as plt
import re
import ast
from collections import namedtuple
from IPython.display import display, Markdown
from statistics import stdev
# Additional data: the variable `data` contains the [electoral votes by state](https://www.britannica.com/topic/United-States-Electoral-College-Votes-by-State-1787124) and the [partisan lean by state](https://github.com/fivethirtyeight/data/tree/master/partisan-lean) (how much more Republican (plus) or Democratic (minus) leaning the state is compared to the country as a whole, across recent elections). The variable `net_usa` has the [country-wide net presidential approval](https://projects.fivethirtyeight.com/trump-approval-ratings/) by month.
# +
data = { # From https://github.com/fivethirtyeight/data/tree/master/partisan+lean
# a dict of {"state name": (electoral_votes, partisan_lean)}
"Alabama": (9, +27), "Alaska": (3, +15), "Arizona": (11, +9),
"Arkansas": (6, +24), "California": (55, -24), "Colorado": (9, -1),
"Connecticut": (7, -11), "Delaware": (3, -14), "District of Columbia": (3, -43),
"Florida": (29, +5), "Georgia": (16, +12), "Hawaii": (4, -36),
"Idaho": (4, +35), "Illinois": (20, -13), "Indiana": (11, +18),
"Iowa": (6, +6), "Kansas": (6, +23), "Kentucky": (8, +23),
"Louisiana": (8, +17), "Maine": (4, -5), "Maryland": (10, -23),
"Massachusetts": (11, -29), "Michigan": (16, -1), "Minnesota": (10, -2),
"Mississippi": (6, +15), "Missouri": (10, +19), "Montana": (3, +18),
"Nebraska": (5, +24), "Nevada": (6, +1), "New Hampshire": (4, +2),
"New Jersey": (14, -13), "New Mexico": (5, -7), "New York": (29, -22),
"North Carolina": (15, +5), "North Dakota": (3, +33), "Ohio": (18, +7),
"Oklahoma": (7, +34), "Oregon": (7, -9), "Pennsylvania": (20, +1),
"Rhode Island": (4, -26), "South Carolina": (9, +17), "South Dakota": (3, +31),
"Tennessee": (11, +28), "Texas": (38, +17), "Utah": (6, +31),
"Vermont": (3, -24), "Virginia": (13, 0), "Washington": (12, -12),
"West Virginia": (5, +30), "Wisconsin": (10, +1), "Wyoming": (3, +47)}
net_usa = { # From https://projects.fivethirtyeight.com/trump-approval-ratings/
'1-Jan-17': +10, # a dict of {date: country-wide-net-approval}
'1-Feb-17': 0, '1-Mar-17': -6, '1-Apr-17': -13, '1-May-17': -11,
'1-Jun-17': -16, '1-Jul-17': -15, '1-Aug-17': -19, '1-Sep-17': -20,
'1-Oct-17': -17, '1-Nov-17': -19, '1-Dec-17': -18, '1-Jan-18': -18,
'1-Feb-18': -15, '1-Mar-18': -14, '1-Apr-18': -13, '1-May-18': -12,
'1-Jun-18': -11, '1-Jul-18': -10, '1-Aug-18': -12, '1-Sep-18': -14,
'1-Oct-18': -11, '1-Nov-18': -11, '1-Dec-18': -10, '1-Jan-19': -12,
'1-Feb-19': -16, '1-Mar-19': -11, '1-Apr-19': -11, '1-May-19': -12,
'1-Jun-19': -12, '1-Jul-19': -11}
# -
# Now the code to parse and manipulate the data:
# +
class State(namedtuple('_', 'name, ev, lean, approvals, disapprovals')):
'''A State has a name, the number of electoral votes, the partisan lean,
and two dicts of {date: percent}: approvals and disapprovals'''
def parse_page(filename='evs.html', data=data):
"Read data from the file and return (list of dates, list of `State`s, last date)."
# File format: Date headers, then [state, approval, disapproval ...]
# [["Demographic","1-Jan-17","","1-Feb-17","", ... "1-Apr-19",""],
# ["Alabama","62","26","65","29", ... "61","35"], ... ] =>
# State("Alabama", 9, approvals={"1-Jan-17": 62, ...}, disapprovals={"1-Jan-17": 26, ...}), ...
text = re.findall(r'\[\[.*?\]\]', open(filename).read())[0]
header, *table = ast.literal_eval(text)
dates = header[1::2] # Every other header entry is a date
states = [State(name, *data[name],
approvals=dict(zip(dates, map(int, numbers[0::2]))),
disapprovals=dict(zip(dates, map(int, numbers[1::2]))))
for (name, *numbers) in table]
return dates, states, dates[-1]
dates, states, now = parse_page()
assert len(states) == 51 and sum(s.ev for s in states) == 538
def EV(states, date=now, swing=0) -> int:
"Total electoral votes with net positive approval (plus half the votes for net zero)."
return sum(s.ev * (1/2 if net(s, date) + swing == 0 else int(net(s, date) + swing > 0))
for s in states)
def margin(states, date=now) -> int:
"What's the least swing that would lead to a majority?"
return next(swing for swing in range(-50, 50) if EV(states, date, swing) >= 270)
def net(state, date=now) -> int: return state.approvals[date] - state.disapprovals[date]
def undecided(state, date=now) -> int: return 100 - state.approvals[date] - state.disapprovals[date]
def movement(state, date=now) -> float: return undecided(state, date) / 5 + 2 * 𝝈(state)
def 𝝈(state, recent=dates[-12:]) -> float: return stdev(net(state, d) for d in recent)
def is_swing(state) -> bool: return abs(net(state)) < movement(state)
# -
# # Current expected electoral votes, with various swings
# How many Electoral votes would Trump expect to get today?
EV(states)
# What across-the-board increase in approval would he need to win?
margin(states)
# How many votes does he get with various swings?
{s: EV(states, swing=s)
for s in range(11)}
# We see that:
# - Trump is currently leading in states with only **172** electoral votes;
# - The margin is **7%** (if he got 7% more popular in key states, his expected total would be 270.5).
# - Swings from 0 to 10% produce electoral vote totals from 172 to 286.
# # Electoral votes by month
#
# The following plot shows, for each month in office, the expected number of electoral votes with error bars indicating a 3% swing in either direction (Why 3%? That was the [average error](https://fivethirtyeight.com/features/the-polls-are-all-right/) in national presidential polls in 2016: Clinton was predicted by polls to win the popular vote by 6% but actually only won by 3%.) Trump hasn't been above 270 since 4 months into his term, and even with the 3% swing, since 6 months in.
# +
def labels(xlab, ylab): plt.xlabel(xlab); plt.ylabel(ylab); plt.grid(True); plt.legend()
plt.rcParams["figure.figsize"] = [12, 10]
plt.style.use('fivethirtyeight')
def plot1(states, dates, swing=3):
N = len(dates)
err = [[EV(states, date) - EV(states, date, -swing) for date in dates],
[EV(states, date, swing) - EV(states, date) for date in dates]]
plt.plot(range(N), [270] * N, color='darkorange', label="270 EVs")
plt.errorbar(range(N), [EV(states, date) for date in dates], fmt='D-',
yerr=err, ecolor='grey', capsize=7, label='Trump EVs ±3% swing')
labels('Months into term', 'Electoral Votes')
plot1(states, dates)
# -
# # Margin and country-wide net approval by month
#
# The next plot gives the swing margin needed to reach 270 for each month, along with the country-wide net approval. Trump has been in negative territory on all metrics since his fourth month in office. He's been net -10% or worse every month since his third in office. His necessary margin has been 4% or worse every month since his seventh. We see that the state-by-state margin roughly correlates with the country-wide net approval, but not exactly.
# +
def plot2(states, dates):
N = len(dates)
plt.plot(range(N), [0] * N, label='Net zero', color='darkorange')
plt.plot(range(N), [-margin(states, date) for date in dates], 'D-', label='Margin to 270')
plt.plot(range(N), [net_usa[date] for date in dates], 'go-', label='Country-wide Net')
labels('Months into term', 'Net popularity')
plot2(states, dates)
# -
# # Month-by-month summary table
#
# For each month, we show the expected electoral vote total (**EVs**), the swing margin needed to get to 270 (**Margin**), the overall (popular vote) net approval across the whole country (**Country**), and then the total percentage of undecided voters and in parentheses the number of states with at least 5% undecided.
# Note that the country-wide vote is not all that correlated with the state-by-state margin: recently the state-by-state margin has held at 7% while the country-wide net approval has ranged from -10% to -16%, and when the state-by-state margin jumped to 11%, the country-wide measure stayed right in the middle at 12%.
# +
def header(head) -> str: return head + '\n' + '-'.join('|' * head.count('|'))
def markdown(fn) -> callable: return lambda *args: display(Markdown('\n'.join(fn(*args))))
@markdown
def by_month(states, dates=reversed(dates)):
yield header('|Month|EVs|Margin|Country|Undecided|')
for date in dates:
month = date.replace('1-', '').replace('-', ' 20')
yield (f'|{month}|{int(EV(states, date))}|{margin(states, date)}%|{net_usa[date]}%'
f'|{sum(s.ev * undecided(s, date) for s in states) / 538:.0f}% '
f'({sum(undecided(s, date) > 5 for s in states)})|')
by_month(states)
# -
# # State-by-state summary table
#
# Below is each state sorted by net approval, with the state's maximum expected movement, and electoral vote allotment, followed by the cumulative running total of electoral votes and the percentages of approval, disapprovals, and undecided in the state, and finally the standard deviation of the net approval over the last 12 months. By going down the **Total** column, you can see what it takes to win.
#
# The **CAPITALIZED bold state names** are the **swing states**, which I define as states in which the absolute value of net approval is less than two standard deviations of the net approval over time, plus a fifth of the undecided voters. The idea is that if we are just dealing with random sampling variation, you could expect future approval to be within two standard deviations 95% of the time, and if the undecideds split 60/40, then a candidate could get a net fifth of them. So it would be very unusual for the non-bold states to flip, unless some events change perception of the candidates.
#
# This analysis says that to win, Trump would need to take *all* the swing states, plus Ohio, Arizona, and Pennsylvannia, which are traditionally considered swing states, but are not under my model because Trump currently trails by a lot (6 0r 7% in each state), and movement there is low.
#
# +
@markdown
def by_state(states, d=now):
total = 0
yield header('|State|Net|Move|EV|Total|+|-|?|𝝈|')
for s in sorted(states, key=net, reverse=True):
total += s.ev
b = '**' * is_swing(s)
yield (f'|{swing_name(s)}|{b}{net(s):+d}%{b}|{b}{movement(s):.0f}%{b}|{s.ev}|{total}'
f'|{s.approvals[d]}%|{s.disapprovals[d]}%|{undecided(s, now)}%|{𝝈(s):3.1f}%|')
def swing_name(s) -> str: return ('**' + s.name.upper() + '**') if is_swing(s) else s.name
by_state(states)
# -
# # Popularity Above Replacement President (PARP) table
#
# Fivethirtyeight is a combination sports/politics site, and it has a lot of statistics about sports players and how much better they are than the average replacement player. Given that, they [decided](https://fivethirtyeight.com/features/the-states-where-trump-is-more-and-less-popular-than-he-should-be/) to rate the president's approval versus each state's overall approval of his party (in recent elections), which is a way of rating the president's performance versus an average replacement candidate from the same party. I'll duplicate that work and keep it up to date.
#
# There are only five states where Trump is exceeding a replacement Republican (i.e., has a positive PARP): one deep-red southern state, Mississippi, and three deep-blue coastal states, Hawaii, Delaware, and Rhode Island. Again, the swing states are **BOLD CAPITALIZED**.
# +
def parp(state) -> int: return net(state) - state.lean
@markdown
def by_parp(states, d=now):
yield header('|State|PARP|Net|Lean|EV|')
for s in sorted(states, key=parp, reverse=True):
yield (f'|{swing_name(s)}|{parp(s):+d}|{net(s):+d}|{s.lean:+d}|{s.ev}|')
by_parp(states)
|
ipynb/Electoral Votes.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import json
import pandas as pd
schema = json.loads(open("data/taxi-cab-classification/schema.json", 'r').read())
columns = [x['name'] for x in schema]
','.join(columns)
data = pd.read_csv("data/taxi-cab-classification/train.csv")
data
with open("data/taxi-cab-classification/train.csv") as f:
print(f.__next__())
import matplotlib
plt = data['tips'].hist(bins=50)
matplotlib.pyplot.show()
data.isna().any()[data.isna().any() == True].index
|
examples/taxi-cab-classification/read_data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Leak Location model
# ## Introduction
# This code is part of Fuel leak detection and location based on NPW.
# When a leak occurs, the fluid is forced to change direction suddenly while in motion. Because of this change, two pressure waves are generated and propagated through the fluid, one upstream and the other downstream. This model aims to find a good estimation for the velocity of sound in the fuel.
# 
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as seabornInstance
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
from sklearn import metrics
# %matplotlib inline
from sklearn.model_selection import cross_val_score
from sklearn import datasets, linear_model
import statsmodels.api as sm
from scipy import stats
# -
dataset = pd.read_csv('Data10.csv')
dataset.shape
dataset['TimeDiff'] = dataset['TimeDiff']/1000 #convert ms to sec
#dataset['Distance'] = dataset['Distance']*1000 #convert km to m
dataset['Press1'] = dataset['Press1']*100/4095 #convert to bar
dataset['Press2'] = dataset['Press2']*100/4095 #convert to bar
dataset.head()
dataset.describe()
# +
df=dataset.loc[:,('Distance','TimeDiff','Press1','Press2','Winter','Summer')]
dataset2=dataset.loc[:,('TimeDiff','Press1','Press2','Winter','Summer')]
#df=dataset.loc[:,('Distance','TimeDiff','V','Press1','Press2','Season_num','Winter','Spring','Summer')]
#dataset2=dataset.loc[:,('TimeDiff','Press1','Press2','Winter','Spring','Summer')]
# -
df
df.corr()
import seaborn as sn
corrMatrix = df.corr()
sn.heatmap(corrMatrix, annot=True)
plt.show()
# +
import scipy.stats as stats
stats.f_oneway(dataset['V'][dataset['Season'] == 'Summer'],
dataset['V'][dataset['Season'] == 'Winter'],
dataset['V'][dataset['Season'] == 'In between'])
# -
plt.figure(figsize=(15,10))
plt.tight_layout()
seabornInstance.distplot(dataset['V'])
#
X = dataset2
y = dataset['Distance']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=0)
def powerset(seq):
"""
Returns all the subsets of this set. This is a generator.
"""
if len(seq) <= 0:
yield []
else:
for item in powerset(seq[1:]):
yield [seq[0]]+item
yield item
allEst = pd.DataFrame({'Model':[0],'Len':[0],'Subset': [' '],'rsquared_adj':[0.0],'AIC':[0],'BIC':[0],'mse_resid':[0],'CV':[0.0],
'fvalue':[0],'f_pvalue':[0]})
predictors = ['TimeDiff','Press1','Press2','Winter','Summer']
subsets = [set(s) for s in powerset(predictors)]
lm = LinearRegression()
m = 0
for sb in subsets:
if len(sb):
X7 = sm.add_constant(X_train.loc[:,sb])
#X7 = X_train.loc[:,sb]
est7 = sm.regression.linear_model.OLS(y_train, X7)
est7 = est7.fit()
allEst.at[m,'Model'] = m+1
allEst.at[m,'Len'] = len(sb)
substr = ', '.join(str(e) for e in sb)
allEst.at[m,'Subset'] = str(substr)
allEst.at[m,'rsquared_adj'] = est7.rsquared_adj
allEst.at[m,'AIC'] = est7.aic
allEst.at[m,'BIC'] = est7.bic
scores = cross_val_score(lm, X7, y_train, scoring='r2', cv=10)
allEst.at[m,'CV'] = scores.mean()
allEst.at[m,'fvalue'] = est7.fvalue
allEst.at[m,'f_pvalue'] = est7.f_pvalue
allEst.at[m,'mse_resid'] = est7.mse_resid
m = m+1
allEst
allEst.to_csv('all_est.csv', encoding='utf-8', index=False)
allEst = pd.DataFrame({'Model':[0],'Len':[0],'Subset': [' '],'rsquared_adj':[0.0],'AIC':[0],'BIC':[0],'mse_resid':[0],'CV':[0.0],
'fvalue':[0],'f_pvalue':[0]})
predictors = ['TimeDiff','Press1','Press2','Winter','Summer']
subsets = [set(s) for s in powerset(predictors)]
lm = LinearRegression()
m = 0
for sb in subsets:
if len(sb):
X7 = X_train.loc[:,sb]
#X7 = sm.add_constant(X_train.loc[:,sb])#
est7 = sm.regression.linear_model.OLS(y_train, X7)
est7 = est7.fit()
allEst.at[m,'Model'] = m+1
allEst.at[m,'Len'] = len(sb)
substr = ', '.join(str(e) for e in sb)
allEst.at[m,'Subset'] = str(substr)
allEst.at[m,'rsquared_adj'] = est7.rsquared_adj
allEst.at[m,'AIC'] = est7.aic
allEst.at[m,'BIC'] = est7.bic
scores = cross_val_score(lm, X7, y_train, scoring='r2', cv=10)
allEst.at[m,'CV'] = scores.mean()
allEst.at[m,'fvalue'] = est7.fvalue
allEst.at[m,'f_pvalue'] = est7.f_pvalue
allEst.at[m,'mse_resid'] = est7.mse_resid
m = m+1
a1 = allEst[allEst['Len'] == 1].sort_values(by='rsquared_adj', ascending=False)
a1
a2 = allEst[allEst['Len'] == 2].sort_values(by='rsquared_adj', ascending=False)
a2
a3 = allEst[allEst['Len'] == 3].sort_values(by='rsquared_adj', ascending=False)
a3
a4 = allEst[allEst['Len'] == 4].sort_values(by='rsquared_adj', ascending=False)
a4
a5 = allEst[allEst['Len'] == 5].sort_values(by='rsquared_adj', ascending=False)
a5
allEstRes = a5.append(a1.iloc[0,:]).append(a2.iloc[0,:]).append(a3.iloc[0,:]).append(a4.iloc[0,:])
allEstRes.sort_values(by='Len')
Subset1 = ['TimeDiff']
Subset2 = ['Press2', 'TimeDiff']
Subset3 = ['Press2', 'Winter','TimeDiff']
Subset4 = ['Press2', 'Winter', 'Summer', 'TimeDiff']
Subset5 = ['Press2', 'Press1', 'Summer', 'Winter', 'TimeDiff']
X1 = sm.add_constant(X_train[Subset1])
est1 = sm.regression.linear_model.OLS(y_train, X1)
est1 = est1.fit()
print(est1.summary())
X1_0 = X_train[Subset1]
est1_0 = sm.regression.linear_model.OLS(y_train, X1_0)
est1_0 = est1_0.fit()
print(est1_0.summary())
X2 = sm.add_constant(X_train[Subset2])
est2 = sm.regression.linear_model.OLS(y_train, X2)
est2 = est2.fit()
print(est2.summary())
X2_0 = X_train[Subset2]
est2_0 = sm.regression.linear_model.OLS(y_train, X2_0)
est2_0 = est2_0.fit()
print(est2_0.summary())
X3 = sm.add_constant(X_train[Subset3])
est3 = sm.regression.linear_model.OLS(y_train, X3)
est3 = est3.fit()
print(est3.summary())
X3_0 = X_train[Subset3]
est3_0 = sm.regression.linear_model.OLS(y_train, X3_0)
est3_0 = est3_0.fit()
print(est3_0.summary())
X4 = sm.add_constant(X_train[Subset4])
est4 = sm.regression.linear_model.OLS(y_train, X4)
est4 = est4.fit()
print(est4.summary())
X4_0 = X_train[Subset4]
est4_0 = sm.regression.linear_model.OLS(y_train, X4_0)
est4_0 = est4_0.fit()
print(est4_0.summary())
X5 = sm.add_constant(X_train[Subset5])
est5 = sm.regression.linear_model.OLS(y_train, X5)
est5 = est5.fit()
print(est5.summary())
X5_0 = X_train[Subset5]
est5_0 = sm.regression.linear_model.OLS(y_train, X5_0)
est5_0 = est5_0.fit()
print(est5_0.summary())
# +
from statsmodels.nonparametric.smoothers_lowess import lowess
residuals = est1.resid
fitted = est1.fittedvalues
smoothed = lowess(residuals,fitted)
top3 = abs(residuals).sort_values(ascending = False)[:3]
plt.rcParams.update({'font.size': 16})
plt.rcParams["figure.figsize"] = (8,7)
fig, ax = plt.subplots()
ax.scatter(fitted, residuals, edgecolors = 'k', facecolors = 'none')
ax.plot(smoothed[:,0],smoothed[:,1],color = 'r')
ax.set_ylabel('Residuals')
ax.set_xlabel('Fitted Values')
ax.set_title('Residuals vs. Fitted')
ax.plot([min(fitted),max(fitted)],[0,0],color = 'k',linestyle = ':', alpha = .3)
for i in top3.index:
ax.annotate(i,xy=(fitted[i],residuals[i]))
plt.show()
# +
sorted_student_residuals = pd.Series(est1.get_influence().resid_studentized_internal)
sorted_student_residuals.index = est1.resid.index
sorted_student_residuals = sorted_student_residuals.sort_values(ascending = True)
df = pd.DataFrame(sorted_student_residuals)
df.columns = ['sorted_student_residuals']
df['theoretical_quantiles'] = stats.probplot(df['sorted_student_residuals'], dist = 'norm', fit = False)[0]
rankings = abs(df['sorted_student_residuals']).sort_values(ascending = False)
top3 = rankings[:3]
fig, ax = plt.subplots()
x = df['theoretical_quantiles']
y = df['sorted_student_residuals']
ax.scatter(x,y, edgecolor = 'k',facecolor = 'none')
ax.set_title('Normal Q-Q')
ax.set_ylabel('Standardized Residuals')
ax.set_xlabel('Theoretical Quantiles')
ax.plot([np.min([x,y]),np.max([x,y])],[np.min([x,y]),np.max([x,y])], color = 'r', ls = '--')
for val in top3.index:
ax.annotate(val,xy=(df['theoretical_quantiles'].loc[val],df['sorted_student_residuals'].loc[val]))
plt.show()
# +
residuals = est2.resid
fitted = est2.fittedvalues
smoothed = lowess(residuals,fitted)
top3 = abs(residuals).sort_values(ascending = False)[:3]
plt.rcParams.update({'font.size': 16})
plt.rcParams["figure.figsize"] = (8,7)
fig, ax = plt.subplots()
ax.scatter(fitted, residuals, edgecolors = 'k', facecolors = 'none')
ax.plot(smoothed[:,0],smoothed[:,1],color = 'r')
ax.set_ylabel('Residuals')
ax.set_xlabel('Fitted Values')
ax.set_title('Residuals vs. Fitted')
ax.plot([min(fitted),max(fitted)],[0,0],color = 'k',linestyle = ':', alpha = .3)
for i in top3.index:
ax.annotate(i,xy=(fitted[i],residuals[i]))
plt.show()
# +
sorted_student_residuals = pd.Series(est2.get_influence().resid_studentized_internal)
sorted_student_residuals.index = est2.resid.index
sorted_student_residuals = sorted_student_residuals.sort_values(ascending = True)
df = pd.DataFrame(sorted_student_residuals)
df.columns = ['sorted_student_residuals']
df['theoretical_quantiles'] = stats.probplot(df['sorted_student_residuals'], dist = 'norm', fit = False)[0]
rankings = abs(df['sorted_student_residuals']).sort_values(ascending = False)
top3 = rankings[:3]
fig, ax = plt.subplots()
x = df['theoretical_quantiles']
y = df['sorted_student_residuals']
ax.scatter(x,y, edgecolor = 'k',facecolor = 'none')
ax.set_title('Normal Q-Q')
ax.set_ylabel('Standardized Residuals')
ax.set_xlabel('Theoretical Quantiles')
ax.plot([np.min([x,y]),np.max([x,y])],[np.min([x,y]),np.max([x,y])], color = 'r', ls = '--')
for val in top3.index:
ax.annotate(val,xy=(df['theoretical_quantiles'].loc[val],df['sorted_student_residuals'].loc[val]))
plt.show()
# -
allEst.plot(x='Len', y='mse_resid', style='o')
#plt.title('Press2 vs V')
plt.xlabel('Number of Predictors')
plt.ylabel('R^2')
plt.show()
best_subset = allEstRes
best_subset
best_subset.plot(x='Len', y='rsquared_adj', style='o')
#plt.title('Press2 vs V')
plt.xlabel('Number of Predictors')
plt.ylabel('R^2')
plt.show()
X1_test = sm.add_constant(X_test[Subset1])
X1_0_test = X_test[Subset1]
X2_test = sm.add_constant(X_test[Subset2])
X2_0_test = X_test[Subset2]
X3_test = sm.add_constant(X_test[Subset3])
X3_0_test = X_test[Subset3]
X4_test = sm.add_constant(X_test[Subset4])
X4_0_test = X_test[Subset4]
X5_test = sm.add_constant(X_test[Subset5])
X5_0_test = X_test[Subset5]
y_pred1 = est1.predict(X1_test)
y_pred1_0 = est1_0.predict(X1_0_test)
y_pred2 = est2.predict(X2_test)
y_pred2_0 = est2_0.predict(X2_0_test)
y_pred3 = est3.predict(X3_test)
y_pred3_0 = est3_0.predict(X3_0_test)
y_pred4 = est4.predict(X4_test)
y_pred4_0 = est4_0.predict(X4_0_test)
y_pred5 = est5.predict(X5_test)
y_pred5_0 = est5_0.predict(X5_0_test)
df = pd.DataFrame({'Actual': y_test, 'Predicted1': y_pred1, 'Predicted1_0': y_pred1_0,'Predicted2': y_pred2, 'Predicted2_0': y_pred2_0,'Predicted3': y_pred3, 'Predicted3_0': y_pred3_0
,'Predicted4': y_pred4, 'Predicted4_0': y_pred4_0,'Predicted5': y_pred5, 'Predicted5_0': y_pred5_0})
df
df.to_csv('prediction.csv', encoding='utf-8', index=False)
plt.scatter(X_test['TimeDiff'], df['Actual'], color='gray')
plt.plot(X_test['TimeDiff'], y_pred1, color='blue', linewidth=1)
plt.plot(X_test['TimeDiff'], y_pred2_0, color='red', linewidth=1)
plt.plot(X_test['TimeDiff'], y_pred3_0, color='green', linewidth=1)
plt.show()
print('Mean Absolute Error:', metrics.mean_absolute_error(y_test, y_pred1))
print('Mean Squared Error:', metrics.mean_squared_error(y_test, y_pred1))
print('Root Mean Squared Error:', np.sqrt(metrics.mean_squared_error(y_test, y_pred1)))
# +
# sum of square of residuals
ssr = np.sum((y_pred1 - y_test)**2)
# total sum of squares
sst = np.sum((y_test - np.mean(y_test))**2)
# R2 score
r2_score = 1 - (ssr/sst)
# -
r2_score
print('Mean Absolute Error:', metrics.mean_absolute_error(y_test, y_pred1_0))
print('Mean Squared Error:', metrics.mean_squared_error(y_test, y_pred1_0))
print('Root Mean Squared Error:', np.sqrt(metrics.mean_squared_error(y_test, y_pred1_0)))
print('Mean Absolute Error:', metrics.mean_absolute_error(y_test, y_pred2))
print('Mean Squared Error:', metrics.mean_squared_error(y_test, y_pred2))
print('Root Mean Squared Error:', np.sqrt(metrics.mean_squared_error(y_test, y_pred2)))
print('Mean Absolute Error:', metrics.mean_absolute_error(y_test, y_pred2_0))
print('Mean Squared Error:', metrics.mean_squared_error(y_test, y_pred2_0))
print('Root Mean Squared Error:', np.sqrt(metrics.mean_squared_error(y_test, y_pred2_0)))
print('Mean Absolute Error:', metrics.mean_absolute_error(y_test, y_pred3))
print('Mean Squared Error:', metrics.mean_squared_error(y_test, y_pred3))
print('Root Mean Squared Error:', np.sqrt(metrics.mean_squared_error(y_test, y_pred3)))
print('Mean Absolute Error:', metrics.mean_absolute_error(y_test, y_pred3_0))
print('Mean Squared Error:', metrics.mean_squared_error(y_test, y_pred3_0))
print('Root Mean Squared Error:', np.sqrt(metrics.mean_squared_error(y_test, y_pred3_0)))
print('Mean Absolute Error:', metrics.mean_absolute_error(y_test, y_pred4))
print('Mean Squared Error:', metrics.mean_squared_error(y_test, y_pred4))
print('Root Mean Squared Error:', np.sqrt(metrics.mean_squared_error(y_test, y_pred4)))
print('Mean Absolute Error:', metrics.mean_absolute_error(y_test, y_pred4_0))
print('Mean Squared Error:', metrics.mean_squared_error(y_test, y_pred4_0))
print('Root Mean Squared Error:', np.sqrt(metrics.mean_squared_error(y_test, y_pred4_0)))
print('Mean Absolute Error:', metrics.mean_absolute_error(y_test, y_pred5))
print('Mean Squared Error:', metrics.mean_squared_error(y_test, y_pred5))
print('Root Mean Squared Error:', np.sqrt(metrics.mean_squared_error(y_test, y_pred5)))
print('Mean Absolute Error:', metrics.mean_absolute_error(y_test, y_pred5_0))
print('Mean Squared Error:', metrics.mean_squared_error(y_test, y_pred5_0))
print('Root Mean Squared Error:', np.sqrt(metrics.mean_squared_error(y_test, y_pred5_0)))
# ### Under 25 km
df_25 = df[df['Actual'] < 25]
y_25 = df_25['Actual']
y_pred_25 = df_25['Predicted3_0']
print('Mean Absolute Error:', metrics.mean_absolute_error(y_25, y_pred_25))
print('Mean Squared Error:', metrics.mean_squared_error(y_25, y_pred_25))
print('Root Mean Squared Error:', np.sqrt(metrics.mean_squared_error(y_25, y_pred_25)))
# ## 25 - 50 km
df_50 = df[(df['Actual'] >= 25) & (df['Actual'] < 50)]
y_50 = df_50['Actual']
y_pred_50 = df_50['Predicted3_0']
print('Mean Absolute Error:', metrics.mean_absolute_error(y_50, y_pred_50))
print('Mean Squared Error:', metrics.mean_squared_error(y_50, y_pred_50))
print('Root Mean Squared Error:', np.sqrt(metrics.mean_squared_error(y_50, y_pred_50)))
|
LeakLocation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] hide_input=true
# # Risk analysis
# -
# ### Generating data
# +
# Importing and data
import theano.tensor as T
import theano
import sys, os
sys.path.append("../")
sys.path.append("../gempy")
# Importing GeMpy modules
import gempy as GeMpy
# Reloading (only for development purposes)
#import importlib
#importlib.reload(GeMpy)
# Usuful packages
import numpy as np
import pandas as pn
import matplotlib.pyplot as plt
# This was to choose the gpu
os.environ['CUDA_LAUNCH_BLOCKING'] = '1'
# Default options of printin
np.set_printoptions(precision = 6, linewidth= 130, suppress = True)
# %matplotlib qt5
# #%matplotlib notebook
# Importing the data from csv files and settign extent and resolution
geo_data = GeMpy.create_data([0,2000,0,2000,-2000,0],[ 50,50,50],
path_f = os.pardir+"/input_data/FabLessPoints_Foliations.csv",
path_i = os.pardir+"/input_data/FabLessPoints_Points.csv")
# -
# Assigning series to formations as well as their order (timewise)
GeMpy.set_data_series(geo_data, {"fault":geo_data.formations[4],
"Rest":np.delete(geo_data.formations, 4)},
order_series = ["fault",
"Rest",
], verbose=0)
geo_data.n_faults = 1
GeMpy.data_to_pickle(GeMpy.select_series(geo_data, ['Rest']), 'NoFault')
# + run_control={"marked": true}
GeMpy.data_to_pickle(geo_data, 'BasicFault')
# -
data_int = GeMpy.InterpolatorInput(geo_data, dtype='float64',
verbose = ['potential_field_at_interfaces'])
f = data_int.compile_th_fn()
i = data_int.get_input_data()
s = f(*i)
s.shape
np.save('pot_field', s[0,1,:])
a = (np.zeros_like(geo_data.grid.grid[:, 0]).astype(bool).
reshape(9,9,9))
b = np.indices((9,9,9))
b[2, 0, :, :]
b[1, 0, :, :]
b[1,0,2:4,2:4]
a[7,4,0] = True
geo_data.grid.grid[np.ravel(a)]
geo_data.grid.grid[0, 4, 7]
# +
# np.indices?
# -
GeMpy.plot_data(geo_data)
# +
from mpl_toolkits.mplot3d import Axes3D
import matplotlib.pyplot as plt
import numpy as np
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
xs = geo_data.grid.grid[:,0]
ys = geo_data.grid.grid[:,1]
zs = geo_data.grid.grid[:,2]
ax.scatter(xs, ys, zs, )
ax.set_xlabel('X Label')
ax.set_ylabel('Y Label')
ax.set_zlabel('Z Label')
plt.show()
# -
geo_data.interfaces
geo_data.extent
# + run_control={"marked": false}
import visualization_vtk as vv
vv.visualize(geo_data)
# +
# #!/usr/bin/env python
import vtk
# Create a superquadric
superquadricSource = vtk.vtkSuperquadricSource()
superquadricSource.SetPhiRoundness(3.1)
superquadricSource.SetThetaRoundness(1.0)
superquadricSource.Update() # needed to GetBounds later
renderer = vtk.vtkRenderer()
mapper = vtk.vtkPolyDataMapper()
mapper.SetInputConnection(superquadricSource.GetOutputPort())
superquadricActor = vtk.vtkActor()
superquadricActor.SetMapper(mapper)
cubeAxesActor = vtk.vtkCubeAxesActor()
cubeAxesActor.SetBounds(superquadricSource.GetOutput().GetBounds())
cubeAxesActor.SetCamera(renderer.GetActiveCamera())
cubeAxesActor.GetTitleTextProperty(0).SetColor(1.0, 0.0, 0.0)
cubeAxesActor.GetLabelTextProperty(0).SetColor(1.0, 0.0, 0.0)
cubeAxesActor.GetTitleTextProperty(1).SetColor(0.0, 1.0, 0.0)
cubeAxesActor.GetLabelTextProperty(1).SetColor(0.0, 1.0, 0.0)
cubeAxesActor.GetTitleTextProperty(2).SetColor(0.0, 0.0, 1.0)
cubeAxesActor.GetLabelTextProperty(2).SetColor(0.0, 0.0, 1.0)
cubeAxesActor.DrawXGridlinesOn()
cubeAxesActor.DrawYGridlinesOn()
cubeAxesActor.DrawZGridlinesOn()
if vtk.VTK_MAJOR_VERSION > 5:
cubeAxesActor.SetGridLineLocation(vtk.VTK_GRID_LINES_FURTHEST)
cubeAxesActor.XAxisMinorTickVisibilityOff()
cubeAxesActor.YAxisMinorTickVisibilityOff()
cubeAxesActor.ZAxisMinorTickVisibilityOff()
renderer.AddActor(cubeAxesActor)
renderer.AddActor(superquadricActor)
renderer.GetActiveCamera().Azimuth(30)
renderer.GetActiveCamera().Elevation(30)
renderer.ResetCamera()
renderWindow = vtk.vtkRenderWindow()
renderWindow.AddRenderer(renderer)
renderWindowInteractor = vtk.vtkRenderWindowInteractor()
renderWindowInteractor.SetRenderWindow(renderWindow)
renderWindow.Render()
renderWindowInteractor.Start()
del renderWindow, renderWindowInteractor
# -
GeMpy.data_to_pickle(geo_data)
geo_data.data_to_pickle()
geo_data.formations
# Assigning series to formations as well as their order (timewise)
GeMpy.set_data_series(geo_data, {"fault":geo_data.formations[4],
"Rest":np.delete(geo_data.formations, 4)},
order_series = ["fault",
"Rest",
], verbose=0)
geo_data_s = GeMpy.select_series(geo_data, ["fault"])
# Select series to interpolate (if you do not want to interpolate all)
data_interp = GeMpy.set_interpolator(geo_data,
verbose = ['potential_field_at_interfaces'],
dtype='float64')
# This are the shared parameters and the compilation of the function. This will be hidden as well at some point
input_data_T = data_interp.interpolator.tg.input_parameters_list()
data_interp.interpolator.tg.len_series_i.get_value()
# This prepares the user data to the theano function
input_data_P = data_interp.interpolator.data_prep(u_grade=[3])
# HERE when we compile we have to pass the number of formations that are faults!!
debugging = theano.function(input_data_T, data_interp.interpolator.tg.whole_block_model(0), on_unused_input='ignore',
allow_input_downcast=True, profile=True);
# Solution of theano
sol = debugging(input_data_P[0], input_data_P[1], input_data_P[2], input_data_P[3],input_data_P[4], input_data_P[5])
data_interp.rescaling_factor*data_interp.interpolator.tg.a_T.get_value()
# +
GeMpy.plot_potential_field(geo_data, sol[0, 1, :], 25, direction='x')
# -
GeMpy.plot_section(geo_data, 100, block=sol[0,0, :], direction='y', plot_data=True)
import numpy as np
import pandas as pn
geo_res = pn.read_csv('voxet_sub.vox')
geo_res = geo_res.iloc[9:]
#, geo_data.formations
ip_addresses = geo_res['nx 500'].unique()#geo_data.interfaces["formation"].unique()
ip_dict = dict(zip(ip_addresses, range(1, len(ip_addresses) + 1)))
#ip_dict['Murchison'] = 0
#ip_dict['out'] = 0
#ip_dict['SimpleMafic'] = 4
geo_res_num = geo_res['nx 500'].replace(ip_dict)
a = geo_res_num.as_matrix()
# + run_control={"marked": true}
#perth = np.tile(geo_res_num.as_matrix().reshape(500, 500, 8), (1,1,10))
perth = a.reshape(500, 500, 8, order='F')
#perth = np.tile(geo_res_num.as_matrix().reshape(500, 500, 8), (1,1,10))
perth = np.repeat(perth, 10, axis=2)
# -
perth.shape
ip_dict
perth.shape
# +
from evtk.hl import imageToVTK
import numpy as np
# Dimensions
nx, ny, nz = perth.shape
ncells = nx * ny * nz
npoints = (nx + 1) * (ny + 1) * (nz + 1)
# Variables
lith = perth
imageToVTK("./PerthBasin",
# cellData = {"lith_cell" : lith},
pointData = {"lith_point" : lith}
)
# +
from evtk.hl import imageToVTK
import numpy as np
# Dimensions
nx, ny, nz = 200,200,200
ncells = nx * ny * nz
npoints = (nx + 1) * (ny + 1) * (nz + 1)
# Variables
lith = sol[0,0, :].reshape( (nx, ny, nz), order = 'C')
imageToVTK("./FabianModel_point_good_res",
# cellData = {"lith_cell" : lith},
pointData = {"lith_point" : lith}
)
# -
sol
# +
from skimage import measure
vertices, simplices = measure.marching_cubes(sol[1,:].reshape(50,50,50), -0.559606,
spacing=(10.0, 10.0, 10.0),)
x,y,z = zip(*vertices)
# vertices, simplices = measure.marching_cubes(sol[1,:].reshape(50,50,50), 0.43724)
# x2,y2,z2 = zip(*vertices)
# vertices, simplices = measure.marching_cubes(sol[1,:].reshape(50,50,50), 0.414782)
# x3,y3,z3 = zip(*vertices)
# vertices, simplices = measure.marching_cubes(sol[1,:].reshape(50,50,50), 0.220775)
# x4,y4,z4 = zip(*vertices)
# -
vertices.max()
# + run_control={"marked": false}
import vtk
from vtk import *
#setup points and vertices
Points = vtk.vtkPoints()
Triangles = vtk.vtkCellArray()
Triangle = vtk.vtkTriangle()
for p in vertices*0.4:
Points.InsertNextPoint(p)
#Unfortunately in this simple example the following lines are ambiguous.
#The first 0 is the index of the triangle vertex which is ALWAYS 0-2.
#The second 0 is the index into the point (geometry) array, so this can range from 0-(NumPoints-1)
#i.e. a more general statement is triangle->GetPointIds()->SetId(0, PointId);
for i in simplices:
Triangle.GetPointIds().SetId(0, i[0])
Triangle.GetPointIds().SetId(1, i[1])
Triangle.GetPointIds().SetId(2, i[2])
Triangles.InsertNextCell(Triangle)
polydata = vtk.vtkPolyData()
polydata.SetPoints(Points)
polydata.SetPolys(Triangles)
polydata.Modified()
if vtk.VTK_MAJOR_VERSION <= 5:
polydata.Update()
writer = vtk.vtkXMLPolyDataWriter();
writer.SetFileName("Fabian_f.vtp");
if vtk.VTK_MAJOR_VERSION <= 5:
writer.SetInput(polydata)
else:
writer.SetInputData(polydata)
writer.Write()
# -
import numpy as np
vertices = np.load('vertices.npy')
simpleces = np.load('simplices.npy')
Triangle1.
Triangle1.GetPointIds().SetId
Vertices
Res = np.array([x,y,z]).T
Non_res_deep = np.array([x4,y4,z4]).T
seal = np.array([x3,y3,z3]).T
sec_res = np.array([x2,y2,z2]).T
#np.save('block_faults', sol[0,0,:])
np.save('vertices', vertices)
np.save('simplices', simplices)
# + run_control={"marked": false}
from mpl_toolkits.mplot3d import Axes3D
import matplotlib.pyplot as plt
import numpy as np
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
# For each set of style and range settings, plot n random points in the box
# defined by x in [23, 32], y in [0, 100], z in [zlow, zhigh].
m = "."
ax.scatter(x, y, z, marker=m)
ax.scatter(x2, y2, z2, c='r', marker=m)
ax.scatter(x3, y3, z3, c='g', marker=m)
ax.scatter(x4, y4, z4, c='black', marker=m)
ax.set_xlabel('X Label')
ax.set_ylabel('Y Label')
ax.set_zlabel('Z Label')
plt.show()
# -
np.min(z)
#
# ## PyMC3
# Input data is a list of arrays with the different values as follows
#
# - Dips position
# - Dip Angles
# - Azimuth
# - Polarity
# - Interfaces points:
# - Reference points
# - Rest points
data_interp.interfaces['formation'].unique()
select = data_interp.interpolator.pandas_rest_layer_points['formation'] == 'Reservoir'
# It is important to notice that in the array ref_layer_points, the values are tiled so it would be necessary to change all of them. At some point it would be nice to tile them in theano I guess.
#
# So the next step is to compile the theano graph that creates the GeMpy model, not as a function but as a theano operation. Basically this is like compress a bunch of theano operations in one
# +
# Everytime you want to compile the theano graph just reset all the theano shared with the next 3 lines of code. I am
# investigating to try to avoid it but in any case at some point it will be hidden to the user
data_interp = GeMpy.set_interpolator(geo_data)
# This are the shared parameters and the compilation of the function. This will be hidden as well at some point
input_data_T = data_interp.interpolator.tg.input_parameters_list()
# This prepares the user data to the theano function
input_data_P = data_interp.interpolator.data_prep(u_grade=[3,3])
geomodel = theano.OpFromGraph(input_data_T, [data_interp.interpolator.tg.whole_block_model(1)], on_unused_input='ignore')
# -
# Because now the GeMpy model is a theano operation and not a theano function, to call it we need to use theano variables (with theano functions we call them with python variables). This is very easy to modify, we just need to use theano shared to convert our python input data into theano variables.
#
# The pymc3 objects are already theano variables (pm.Normal and so on). Now the trick is that using the theano function T.set_subtensor, we can change one deterministic value of the input arrays(the ones printed in the cell above) by a stochastic pymc3 object. Then with the new arrays we just have to call the theano operation and pymc will do the rest
# + run_control={"marked": false}
# This is the creation of the model
import pymc3 as pm
theano.config.compute_test_value = 'off'
model = pm.Model()
with model:
# Stochastic value
reservoir = pm.Normal('reservoir', np.array([0], dtype='float64')
, sd=np.array([0.09], dtype='float64'), dtype='float64', shape=(1))
# We convert a python variable to theano.shared
ref = theano.shared(input_data_P[4])
rest = theano.shared(input_data_P[5])
# We add the stochastic value to the correspondant array
ref = pm.Deterministic('reference', T.set_subtensor(
ref[T.nonzero(T.cast(select.as_matrix(), "int8"))[0], 2],
ref[T.nonzero(T.cast(select.as_matrix(), "int8"))[0], 2]+reservoir))
rest = pm.Deterministic('rest', T.set_subtensor(
rest[T.nonzero(T.cast(select.as_matrix(), "int8"))[0], 2],
rest[T.nonzero(T.cast(select.as_matrix(), "int8"))[0], 2]+reservoir))#
geo_model = pm.Deterministic('GeMpy', geomodel(theano.shared(input_data_P[0]),
theano.shared(input_data_P[1]),
theano.shared(input_data_P[2]),
theano.shared(input_data_P[3]),
ref, rest))
# -
theano.config.compute_test_value = 'ignore'
# This is the sampling
# BEFORE RUN THIS FOR LONG CHECK IN THE MODULE THEANOGRAF THAT THE FLAG THEANO OPTIMIZER IS IN 'fast_run'!!
with model:
# backend = pm.backends.ndarray.NDArray('geomodels')
step = pm.NUTS()
trace = pm.sample(30, init=None, step=step, )
np.save('models',trace.get_values('GeMpy'))
for i in range(100):
GeMpy.plot_section(geo_data, 25, block=trace.get_values('GeMpy')[i][0, :],
direction='y', plot_data=False)
plt.show()
p = model.profile(geo_model)
p.summary()
def calculate_prob_lith(models, n_samples = 100):
import copy
v_lith = np.unique(models[0,0,:])
prob_lith = np.zeros(len(v_lith), dtype = object)
for i, pid in enumerate(v_lith):
prob_lith[i] = copy.deepcopy(models[i,0,:])
prob_lith[i] = np.zeros_like(models[i,0,:])
for lith in model[-1:-n_samples:-1]:
prob_lith[i]+= (lith == pid)/float(len(models[-1:-n_samples:-1, 0, :]))
calculate_prob_lith(trace.get_values('GeMpy')[:,0, :], n_samples=2000)
for i in trace.get_values('GeMpy'):
GeMpy.plot_section(geo_data, 20, direction = 'y', block = i, plot_data = False)
plt.show()
# +
# Cheaper 3D plot
# So far this is a simple 3D visualization. I have to adapt it into GeMpy
sol = trace['GeMpy'][5]
lith0 = sol == 0
lith1 = sol == 2
lith2 = sol == 3
lith3 = sol == 4
lith4 = sol == 5
import ipyvolume.pylab as p3
p3.figure(width=800)
blue = p3.scatter(geo_data.grid.grid[:,0][lith0],
geo_data.grid.grid[:,1][lith0],
geo_data.grid.grid[:,2][lith0], marker='box', color = 'blue', size = 0.1 )
p3.scatter(geo_data.grid.grid[:,0][lith1],
geo_data.grid.grid[:,1][lith1],
geo_data.grid.grid[:,2][lith1], marker='box', color = 'yellow', size = 2.2 )
p3.scatter(geo_data.grid.grid[:,0][lith2],
geo_data.grid.grid[:,1][lith2],
geo_data.grid.grid[:,2][lith2], marker='box', color = 'green', size = 2.2 )
p3.scatter(geo_data.grid.grid[:,0][lith3],
geo_data.grid.grid[:,1][lith3],
geo_data.grid.grid[:,2][lith3], marker='box', color = 'pink', size = 2.2 )
p3.scatter(geo_data.grid.grid[:,0][lith4],
geo_data.grid.grid[:,1][lith4],
geo_data.grid.grid[:,2][lith4], marker='box', color = 'red', size = 2.2 )
#p3.xlim(np.min(geo_data.grid.grid[:,0]),np.min(geo_data.grid.grid[:,0]))
#p3.ylim(np.min(geo_data.grid.grid[:,1]),np.max(geo_data.grid.grid[:,1]))
#p3.zlim(np.min(geo_data.grid.grid[:,2]),np.min(geo_data.grid.grid[:,2]))#np.max(geo_data.grid.grid[:,2]))
#p3.show()
from ipywidgets import FloatSlider, ColorPicker, VBox, jslink
size = FloatSlider(min=0, max=30, step=0.1)
size_selected = FloatSlider(min=0, max=30, step=0.1)
color = ColorPicker()
color_selected = ColorPicker()
jslink((blue, 'size'), (size, 'value'))
#jslink((blue, 'size_selected'), (size_selected, 'value'))
#jslink((quiver, 'color'), (color, 'value'))
#jslink((quiver, 'color_selected'), (color_selected, 'value'))
VBox([p3.gcc(), size])
# -
# The uncertainty in this case is minimum but you get the idea
(trace['GeMpy'][0]-trace['GeMpy'][2]).sum()
for i in trace.get_values('GeMpy'):
GeMpy.plot_section(geo_data, 13, block = i, plot_data = False)
plt.show()
|
Prototype Notebook/FabianThesis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Remove input cells at runtime (nbsphinx)
import IPython.core.display as d
d.display_html('<script>jQuery(function() {if (jQuery("body.notebook_app").length == 0) { jQuery(".input_area").toggle(); jQuery(".prompt").toggle();}});</script>', raw=True)
# # Energy reconstruction (MODEL)
# This notebook contains the same code as in `protopipe.scripts.model_diagnostic`.
# It should be used to test the performance of the trained model **before** use it to estimate the energy of DL2 events.
#
# In fact, what happens in a *protopipe* analysis is that part of the TRAINING sample is used for *testing* the models to get some preliminary diagnostics.
# This notebook shows this camera-wise preliminary diagnostics.
#
# Settings and setup of the plots are done using the same configuration file used for training the model.
#
# **Developers**
# Please, if you have any contribution regarding this part, do it here and not in the relevant sections of the main code, which are now discontinued.
# ## Table of contents
# * [Feature importance](#Feature-importance)
# * [Feature distributions](#Feature-distributions)
# * [Migration distribution](#Migration-distribution)
# * [Energy resolution and bias](#Energy-resolution-and-bias)
# + [markdown] nbsphinx="hidden"
# ## Imports
# [back to top](#Table-of-contents)
# +
import gzip
import glob
from os import path
import pickle
import joblib
import yaml
import numpy as np
import pandas as pd
from scipy.optimize import curve_fit
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
plt.rcParams.update({'figure.max_open_warning': 0})
# + [markdown] nbsphinx="hidden"
# ## Functions and classes
# [back to top](#Table-of-contents)
# -
def load_config(name):
"""Load YAML configuration file."""
try:
with open(name, "r") as stream:
cfg = yaml.load(stream, Loader=yaml.FullLoader)
except FileNotFoundError as e:
print(e)
raise
return cfg
def load_obj(name ):
"""Load object in binary"""
with gzip.open(name, 'rb') as f:
return pickle.load(f)
def plot_hist(ax, data, nbin, limit, norm=False, yerr=False, hist_kwargs={}, error_kw={}):
"""Utility function to plot histogram"""
bin_edges = np.linspace(limit[0], limit[-1], nbin + 1, True)
y, tmp = np.histogram(data, bins=bin_edges)
weights = np.ones_like(y)
if norm is True:
weights = weights / float(np.sum(y))
if yerr is True:
yerr = np.sqrt(y) * weights
else:
yerr = np.zeros(len(y))
centers = 0.5 * (bin_edges[1:] + bin_edges[:-1])
width = bin_edges[1:] - bin_edges[:-1]
ax.bar(centers, y * weights, width=width, yerr=yerr, error_kw=error_kw, **hist_kwargs)
return ax
def plot_distributions(feature_list,
data_list,
nbin=30,
hist_kwargs_list={},
error_kw_list={},
ncols=2):
"""Plot feature distributions for several data set. Returns list of axes."""
n_feature = len(feature_list)
nrows = int(n_feature / ncols) if n_feature % ncols == 0 else int((n_feature + 1) / ncols)
fig, axes = plt.subplots(nrows=nrows, ncols=ncols, figsize=(5 * ncols, 5 * nrows))
if nrows == 1 and ncols == 1:
axes = [axes]
else:
axes = axes.flatten()
for i, colname in enumerate(feature_list):
ax = axes[i]
# Range for binning
range_min = min([data[colname].min() for data in data_list])
range_max = max([data[colname].max() for data in data_list])
myrange = [range_min, range_max]
for j, data in enumerate(data_list):
ax = plot_hist(
ax=ax, data=data[colname], nbin=nbin, limit=myrange,
norm=True, yerr=True,
hist_kwargs=hist_kwargs_list[j],
error_kw=error_kw_list[j]
)
ax.set_xlabel(colname)
ax.set_ylabel('Arbitrary units')
ax.legend(loc='upper left')
ax.grid()
plt.tight_layout()
return fig, axes
def get_evt_subarray_model_output(data,
weight_name=None,
keep_cols=['reco_energy'],
model_output_name='score_img',
model_output_name_evt='score'):
"""
Returns DataStore with keepcols + score/target columns of model at the
level-subarray-event.
Parameters
----------
data: `~pandas.DataFrame`
Data frame
weight_name: `str`
Variable name in data frame to weight events with
keep_cols: `list`, optional
List of variables to keep in resulting data frame
model_output_name: `str`, optional
Name of model output (image level)
model_output_name: `str`, optional
Name of averaged model output (event level)
Returns
--------
data: `~pandas.DataFrame`
Data frame
"""
keep_cols += [model_output_name]
keep_cols += [weight_name]
new_data = data[keep_cols].copy(deep=True)
new_data[model_output_name_evt] = np.zeros(len(new_data))
new_data.set_index(["tel_id"], append=True, inplace=True)
new_data[model_output_name_evt] = new_data.groupby(["obs_id", "event_id"]).apply(
lambda g: np.average(g[model_output_name], weights=g[weight_name])
)
# Remove columns
new_data = new_data.drop(columns=[model_output_name])
# Remove duplicates
new_data = new_data[~new_data.index.duplicated(keep="first")]
return new_data
class ModelDiagnostic(object):
"""
Base class for model diagnostics.
Parameters
----------
model: `~sklearn.base.BaseEstimator`
Best model
feature_name_list: list
List of the features used to buil the model
target_name: str
Name of the target (e.g. score, gamaness, energy, etc.)
"""
def __init__(self, model, feature_name_list, target_name):
self.model = model
self.feature_name_list = feature_name_list
self.target_name = target_name
def plot_feature_importance(self, ax, **kwargs):
"""
Plot importance of features
Parameters
----------
ax: `~matplotlib.axes.Axes`
Axis
"""
if ax is None:
import matplotlib.pyplot as plt
ax = plt.gca()
importance = self.model.feature_importances_
importance, feature_labels = \
zip(*sorted(zip(importance, self.feature_name_list), reverse=True))
bin_edges = np.arange(0, len(importance)+1)
bin_width = bin_edges[1:] - bin_edges[:-1] - 0.1
ax.bar(bin_edges[:-1], importance, width=bin_width, **kwargs)
ax.set_xticks(np.arange(0, len(importance)))
ax.set_xticklabels(feature_labels, rotation=75)
return ax
def plot_features(self, data_list,
nbin=30,
hist_kwargs_list={},
error_kw_list={},
ncols=2):
"""
Plot model features for different data set (e.g. training and test samples).
Parameters
----------
data_list: list
List of data
nbin: int
Number of bin
hist_kwargs_list: dict
Dictionary with histogram options
error_kw_list: dict
Dictionary with error bar options
ncols: int
Number of columns
"""
return plot_distributions(
self.feature_name_list,
data_list,
nbin,
hist_kwargs_list,
error_kw_list, ncols
)
def add_image_model_output(self):
raise NotImplementedError("Please Implement this method")
class RegressorDiagnostic(ModelDiagnostic):
"""
Class to plot several diagnostic plots for regression.
Parameters
----------
model: sklearn.base.BaseEstimator
Scikit model
feature_name_list: str
List of features
target_name: str
Name of target (e.g. `mc_energy`)
data_train: `~pandas.DataFrame`
Data frame
data_test: `~pandas.DataFrame`
Data frame
"""
def __init__(self, model, feature_name_list, target_name, data_train, data_test, output_name):
super().__init__(model, feature_name_list, target_name)
self.data_train = data_train
self.data_test = data_test
self.target_estimation_name = self.target_name
self.output_name = output_name
self.output_name_img = output_name + '_img'
# Compute and add target estimation
self.data_train = self.add_image_model_output(
self.data_train,
col_name=self.output_name_img
)
self.data_test = self.add_image_model_output(
self.data_test,
col_name=self.output_name_img
)
@staticmethod
def plot_resolution_distribution(ax, y_true, y_reco, nbin=100, fit_range=[-3,3],
fit_kwargs={}, hist_kwargs={}):
"""
Compute bias and resolution with a gaussian fit
and return a plot with the fit results and the migration distribution.
"""
def gauss(x, ampl, mean, std):
return ampl * np.exp(-0.5 * ((x - mean) / std) ** 2)
if ax is None:
ax = plt.gca()
migration = (y_reco - y_true) / y_true
bin_edges = np.linspace(fit_range[0], fit_range[-1], nbin + 1, True)
y, tmp = np.histogram(migration, bins=bin_edges)
x = (bin_edges[:-1] + bin_edges[1:]) / 2
try:
param, cov = curve_fit(gauss, x, y)
except:
param = [-1, -1, -1]
cov = [[]]
#print('Not enough stat ? (#evts={})'.format(len(y_true)))
plot_hist(
ax=ax, data=migration, nbin=nbin,
yerr=False,
norm=False,
limit=fit_range,
hist_kwargs=hist_kwargs
)
ax.plot(x, gauss(x, param[0], param[1], param[2]), **fit_kwargs)
return ax, param, cov
def add_image_model_output(self, data, col_name):
data[col_name] = self.model.predict(data[self.feature_name_list])
return data
# + [markdown] nbsphinx="hidden"
# ## Load models
# [back to top](#Table-of-contents)
# -
# Please, if you modify this notebook through a pull request empty these variables before pushing
analysesDir = "" # Where all your analyses are stored
analysisName = "" # The name of this analysis
# +
configuration = f"{analysesDir}/{analysisName}/configs/regressor.yaml"
cfg = load_config(configuration)
model_type = cfg["General"]["model_type"]
method_name = cfg["Method"]["name"]
inDir = f"{analysesDir}/{analysisName}/estimators/energy_regressor"
cameras = [model.split('/')[-1].split('_')[2] for model in glob.glob(f"{inDir}/regressor*.pkl.gz")]
# +
data = {camera : dict.fromkeys(["model", "data_scikit", "data_train", "data_test"]) for camera in cameras}
for camera in cameras:
data[camera]["data_scikit"] = load_obj(
glob.glob(f"{inDir}/data_scikit_{model_type}_{method_name}_*_{camera}.pkl.gz")[0]
)
data[camera]["data_train"] = pd.read_pickle(
glob.glob(f"{inDir}/data_train_{model_type}_{method_name}_*_{camera}.pkl.gz")[0]
)
data[camera]["data_test"] = pd.read_pickle(
glob.glob(f"{inDir}/data_test_{model_type}_{method_name}_*_{camera}.pkl.gz")[0]
)
modelName = f"{model_type}_*_{camera}_{method_name}.pkl.gz"
data[camera]["model"] = joblib.load(glob.glob(f"{inDir}/{modelName}")[0])
# + [markdown] nbsphinx="hidden"
# ## Settings and setup
# [back to top](#Table-of-contents)
# +
# Energy (both true and reconstructed)
nbins = cfg["Diagnostic"]["energy"]["nbins"]
energy_edges = np.logspace(
np.log10(cfg["Diagnostic"]["energy"]["min"]),
np.log10(cfg["Diagnostic"]["energy"]["max"]),
nbins + 1,
True,
)
# -
diagnostic = dict.fromkeys(cameras)
for camera in cameras:
diagnostic[camera] = RegressorDiagnostic(
model=data[camera]["model"],
feature_name_list=cfg["FeatureList"],
target_name="true_energy",
data_train=data[camera]["data_train"],
data_test=data[camera]["data_test"],
output_name="reco_energy",
)
# ## Benchmarks
# [back to top](#Table-of-contents)
# ### Feature importance
# [back to top](#Table-of-contents)
for camera in cameras:
plt.figure(figsize=(5, 5))
ax = plt.gca()
ax = diagnostic[camera].plot_feature_importance(
ax,
**{"alpha": 0.7, "edgecolor": "black", "linewidth": 2, "color": "darkgreen"}
)
ax.set_ylabel("Feature importance")
ax.grid()
plt.title(camera)
plt.tight_layout()
# ### Feature distributions
# [back to top](#Table-of-contents)
for camera in cameras:
print(" ====================================================================================")
print(f" {camera} ")
print(" ====================================================================================")
fig, axes = diagnostic[camera].plot_features(
data_list=[data[camera]["data_train"], data[camera]["data_test"]],
nbin=30,
hist_kwargs_list=[
{
"edgecolor": "blue",
"color": "blue",
"label": "Gamma training",
"alpha": 0.2,
"fill": True,
"ls": "-",
"lw": 2,
},
{
"edgecolor": "blue",
"color": "blue",
"label": "Gamma test",
"alpha": 1,
"fill": False,
"ls": "--",
"lw": 2,
},
],
error_kw_list=[
dict(ecolor="blue", lw=2, capsize=2, capthick=2, alpha=0.2),
dict(ecolor="blue", lw=2, capsize=2, capthick=2, alpha=0.2),
],
ncols=3,
)
plt.title(camera)
fig.tight_layout()
# ### Migration distribution
# [back to top](#Table-of-contents)
for camera in cameras:
# Compute averaged energy
# print("Process test sample...")
print(" ====================================================================================")
print(f" {camera} ")
print(" ====================================================================================")
data_test_evt = get_evt_subarray_model_output(
data[camera]["data_test"],
weight_name="hillas_intensity_reco",
keep_cols=["tel_id", "true_energy"],
model_output_name="reco_energy_img",
model_output_name_evt="reco_energy",
)
ncols = 5
nrows = (
int(nbins / ncols) if nbins % ncols == 0 else int((nbins + 1) / ncols)
)
if nrows == 0:
nrows = 1
ncols = 1
fig, axes = plt.subplots(nrows=nrows, ncols=ncols, figsize=(5 * 5, 10))
try:
axes = axes.flatten()
except:
axes = [axes]
bias = []
resolution = []
energy_centres = []
for ibin in range(len(energy_edges) - 1):
ax = axes[ibin]
test_data = data_test_evt.query(
"true_energy >= {} and true_energy < {}".format(
energy_edges[ibin], energy_edges[ibin + 1]
)
)
# print("Estimate energy for {} evts".format(len(test_data)))
er = test_data["reco_energy"]
emc = test_data["true_energy"]
opt_hist = {
"edgecolor": "black",
"color": "darkgreen",
"label": "data",
"alpha": 0.7,
"fill": True,
}
opt_fit = {"c": "red", "lw": 2, "label": "Best fit"}
ax, fit_param, cov = diagnostic[camera].plot_resolution_distribution(
ax=ax,
y_true=emc,
y_reco=er,
nbin=50,
fit_range=[-2, 2],
hist_kwargs=opt_hist,
fit_kwargs=opt_fit,
)
if fit_param[2] < 0: # negative value are allowed for the fit
fit_param[2] *= -1
label = "[{:.2f},{:.2f}] TeV\n#Evts={}\nmean={:.2f}\nstd={:.2f}".format(
energy_edges[ibin],
energy_edges[ibin + 1],
len(er),
fit_param[1],
fit_param[2],
)
ax.set_ylabel("# Events")
ax.set_xlabel("(E_reco - E_true) / E_true")
ax.set_xlim([-2, 2])
ax.grid()
evt_patch = mpatches.Patch(color="white", label=label)
data_patch = mpatches.Patch(color="blue", label="data")
fit_patch = mpatches.Patch(color="red", label="best fit")
ax.legend(loc="best", handles=[evt_patch, data_patch, fit_patch])
plt.tight_layout()
#print(
# " Fit results: ({:.3f},{:.3f} TeV)".format(
# energy_edges[ibin], energy_edges[ibin + 1]
# )
#)
#try:
# print(" - A : {:.3f} +/- {:.3f}".format(fit_param[0], cov[0][0]))
# print(" - mean : {:.3f} +/- {:.3f}".format(fit_param[1], cov[1][1]))
# print(" - std : {:.3f} +/- {:.3f}".format(fit_param[2], cov[2][2]))
#except:
# print(" ==> Problem with fit, no covariance...".format())
# continue
bias.append(fit_param[1])
resolution.append(fit_param[2])
energy_centres.append(
(energy_edges[ibin] + energy_edges[ibin + 1]) / 2.0
)
plt.show()
# ### Energy resolution and bias
# [back to top](#Table-of-contents)
for camera in cameras:
plt.figure(figsize=(5, 5))
ax = plt.gca()
ax.plot(
energy_centres,
resolution,
marker="s",
color="darkorange",
label="Resolution",
)
ax.plot(energy_centres, bias, marker="s", color="darkgreen", label="Bias")
ax.set_xlabel("True energy [TeV]")
ax.set_ylabel("Energy resolution")
ax.set_xscale("log")
ax.grid()
ax.legend()
ax.set_ylim([-0.2, 1.2])
plt.title(camera)
plt.tight_layout()
|
docs/contribute/benchmarks/MODELS/benchmarks_MODELS_energy.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
withdraw=int(input())
money=float(input())
charges=0.50
if (withdraw%5==0 and withdraw<= money-charges):
print(money-withdraw-charges)
else:
print(money)
|
Python/ATM.ipynb
|
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .scala
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Scala
// language: scala
// name: scala
// ---
// ## Refining till success or error; <NAME>
//
// Here we illustrate running a sequence of tasks till either some criterion for success is met or we have an error (usually a timeout) or all tasks run with no success. Here we seek the statement of Modus Ponens, but with two types in the distribution.
import $cp.bin.`provingground-core-jvm-3d48753.fat.jar`
import provingground._ , interface._, HoTT._, learning._
repl.pprinter() = {
val p = repl.pprinter()
p.copy(
additionalHandlers = p.additionalHandlers.orElse {
translation.FansiShow.fansiHandler
}
)
}
val A = "A" :: Type
val B = "B" :: Type
val MPAB = A ->: (A ->: B) ->: B
def lp(n: Int) = LocalProver(TermState(FiniteDistribution.unif(A, B), FiniteDistribution.unif(A, B))).sharpen(math.pow(2, n))
def ns(n: Int) = lp(n).nextState.map{ns => n -> ns}
val bT = Utils.bestTask[(Int, TermState)]((1 to 30).map(ns), {case (_, s) => s.typs(MPAB) > 0} )
import monix.execution.Scheduler.Implicits.global
val bF = bT.runToFuture
bF.map(_.get._2.typs(MPAB))
val bT2 = Utils.bestTask[(Int, TermState)]((-5 to 25).map(ns), {case (_, s) => s.typs(MPAB) > 0} )
val bF2 = bT2.runToFuture
// ## Conclusions
//
// * Correctly ran until success, even when the success was not in the first step.
// * Huge speedup over running to the end
// * The desired form of Modus Ponens was found soon.
|
notes/2019-09-18-refine-till-success.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the "../input/" directory.
# For example, running this (by clicking run or pressing Shift+Enter) will list the files in the input directory
from subprocess import check_output
#print(check_output(["ls", "../input"]).decode("utf8"))
# Any results you write to the current directory are saved as output.
# Load in the Client Name data
# Make sure all names uppercase (there are some mixed instances)
pd.set_option('display.max_rows', 30)
path = '/home/zongyi/bimbo_data/'
# path = '/Users/zonemercy/jupyter_notebook/bimbo_data/'
vf = pd.read_csv(path+'cliente_tabla.csv',header=0)
vf['NombreCliente'] = vf['NombreCliente'].str.upper()
# -
vf['NombreCliente'].value_counts()[0:10]
# Let's also generate a list of individual word frequency across all names
def tfidf_score_list(vf2, list_len):
from sklearn.feature_extraction.text import TfidfVectorizer
v = TfidfVectorizer()
vf2['New'] = 'na'
a = " ".join(vf2['NombreCliente'])
vf2['New'][0] = a
tfidf = v.fit_transform(vf2['New'])
feature_names = v.get_feature_names()
freq = []
doc = 0
feature_index = tfidf[doc,:].nonzero()[1]
tfidf_scores = zip(feature_index, [tfidf[doc, x] for x in feature_index])
for w, s in [(feature_names[i], s) for (i, s) in tfidf_scores]:
freq.append((w.encode('utf-8'),s))
del vf2['New']
import numpy as np
names = ['word','score']
formats = ['S50','f8']
dtype = dict(names = names, formats=formats)
array = np.array(freq, dtype=dtype)
b = np.sort(array, order='score')
if list_len > len(b)+1:
list_len = len(b)+1
for i in range(1,list_len):
print(b[-i])
tfidf_score_list(vf, 200)
# print(vf[vf['NombreCliente'].str.contains('.*CAFE.*')])
print len(vf[vf['NombreCliente'].str.contains('NEZ|JOSE|NZO|TES')])
# +
# --- Begin Filtering for specific terms
# Note that the order of filtering is significant.
# For example:
# The regex of .*ERIA.* will assign "FRUITERIA" to 'Eatery' rather than 'Fresh Market'.
# In other words, the first filters to occur have a bigger priority.
def filter_specific(vf2):
# Known Large Company / Special Group Types
vf2['NombreCliente'] = vf2['NombreCliente'].str.replace('.*REMISION.*','Consignment')
vf2['NombreCliente'] = vf2['NombreCliente'].str.replace('.*DISTRIBUIDORA.*','Distribut')
vf2['NombreCliente'] = vf2['NombreCliente'].replace(['.*WAL MART.*','.*SAMS CLUB.*'],'Walmart', regex=True)
vf2['NombreCliente'] = vf2['NombreCliente'].str.replace('.*OXXO.*','Oxxo Store')
vf2['NombreCliente'] = vf2['NombreCliente'].str.replace('.*CONASUPO.*','Govt Store')
vf2['NombreCliente'] = vf2['NombreCliente'].str.replace('.*BIMBO.*','Bimbo Store')
# General term search for a random assortment of words I picked from looking at
# their frequency of appearance in the data and common spanish words for these categories
# vf2['NombreCliente'] = vf2['NombreCliente'].replace(['.*NEZ.*','.*JOSE.*','.*NZO.*','.*TES.*'],'Xicans', regex=True)
vf2['NombreCliente'] = vf2['NombreCliente'].replace(['.*COLEG.*','.*UNIV.*','.*ESCU.*','.*INSTI.*',\
'.*PREPAR.*'],'School', regex=True)
vf2['NombreCliente'] = vf2['NombreCliente'].str.replace('.*PUESTO.*','Post')
vf2['NombreCliente'] = vf2['NombreCliente'].replace(['.*FARMA.*','.*HOSPITAL.*','.*CLINI.*'],'Hospital', regex=True)
vf2['NombreCliente'] = vf2['NombreCliente'].replace(['.*CAFE.*','.*CREMERIA.*','.*DULCERIA.*',\
'.*REST.*','.*BURGER.*','.*TACO.*', '.*TORTA.*',\
'.*TAQUER.*','.*HOT DOG.*',\
'.*COMEDOR.*', '.*ERIA.*','.*BURGU.*'],'Eatery', regex=True)
vf2['NombreCliente'] = vf2['NombreCliente'].str.replace('.*SUPER.*','Supermarket')
vf2['NombreCliente'] = vf2['NombreCliente'].replace(['.*COMERCIAL.*','.*BODEGA.*','.*DEPOSITO.*',\
'.*ABARROTES.*','.*MERCADO.*','.*CAMBIO.*',\
'.*MARKET.*','.*MART .*','.*MINI .*',\
'.*PLAZA.*','.*MISC.*','.*ELEVEN.*','.*SEVEN.*','.*EXP.*',\
'.*SNACK.*', '.*PAPELERIA.*', '.*CARNICERIA.*',\
'.*LOCAL.*','.*COMODIN.*','.*PROVIDENCIA.*'
],'General Market'\
, regex=True)
vf2['NombreCliente'] = vf2['NombreCliente'].replace(['.*VERDU.*','.*FRUT.*'],'Fresh Market', regex=True)
vf2['NombreCliente'] = vf2['NombreCliente'].replace(['.*HOTEL.*','.*MOTEL.*'],'Hotel', regex=True)
vf2['NombreCliente'] = vf2['NombreCliente'].replace(['.*NEZ.*','.*JOSE.*','.*NZO.*','.*TES.*'],'Xicans', regex=True)
vf2['NombreCliente'] = vf2['NombreCliente'].str.replace('.*MODELOR.*','Modelor')
vf2['NombreCliente'] = vf2['NombreCliente'].str.replace('.*ARTELI.*','Arteli')
vf2['NombreCliente'] = vf2['NombreCliente'].str.replace('.*CALIMAX.*','Calimax')
# -
filter_specific(vf)
# --- Begin filtering for more general terms
# The idea here is to look for names with particles of speech that would
# not appear in a person's name.
# i.e. "Individuals" should not contain any participles or numbers in their names.
def filter_participle(vf2):
vf2['NombreCliente'] = vf2['NombreCliente'].replace([
'.*LA .*','.*EL .*','.*DE .*','.*LOS .*','.*DEL .*','.*Y .*', '.*SAN .*', '.*SANTA .*',\
'.*AG .*','.*LAS .*','.*MI .*','.*MA .*', '.*II.*', '.*[0-9]+.*'\
],'Small Franchise', regex=True)
filter_participle(vf)
# Any remaining entries should be "Individual" Named Clients, there are some outliers.
# More specific filters could be used in order to reduce the percentage of outliers in this final set.
def filter_remaining(vf2):
def function_word(data):
# Avoid the single-words created so far by checking for upper-case
if (data.isupper()) and (data != "NO IDENTIFICADO"):
return 'Individual'
else:
return data
vf2['NombreCliente'] = vf2['NombreCliente'].map(function_word)
filter_remaining(vf)
vf['NombreCliente'].value_counts()
# +
from sklearn import preprocessing
le = preprocessing.LabelEncoder()
le.fit(vf['NombreCliente'])
print le.classes_
vf['c_clt'] = le.transform(vf['NombreCliente'])
# -
vf.head()
len(vf) #935362
vf = vf[['Cliente_ID','c_clt']]
# vf = vf[vf['Cliente_ID'].unique()]
vf = vf.drop_duplicates(subset='Cliente_ID', keep='last')
vf
vf.to_csv(path+'clients.csv',index=False)
|
Bimbo/client_clf_tfidf.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import random
def sigmoid(z):
return (1/(1+np.exp(-z)))
def propagation(b,X_hat,y_hat):
N=X_hat.shape[0]
z=np.dot(X_hat,b)
s=sigmoid(z)
temp=-(y_hat*np.log(s+0.001)+(1-y_hat)*np.log(1-s+0.001))
L= np.mean(temp,axis=0,keepdims=True)
x1=sigmoid(z)-y_hat
dL= 1/N*np.dot(x1.T,X_hat)
propagate={"L":L,"dL":dL}
return propagate
# + In this section you need to update weight b by gradient descent based on the training data X_hat and y_hat.
# + In his pice of code, dl is the derivative of loss with respect to weight matrix b.
# + Fill None parts in the code.
#
def update(b,X_hat,y_hat,eta):
dl=propagation(b,X_hat,y_hat)["dL"].T
update=b-eta*dl
return update
def init(m,n,method):
if method=='zero':
b=np.zeros((m,n))
elif method=='random':
b=np.random.randn(m,n)
else:
raise Exception('Choose correct method: zero or random')
return b
def b_opt(X_hat,y_hat,eta,steps,initialization):
N=X_hat.shape[0]
p=X_hat.shape[1]
b=init(p,1,initialization)
for i in range(steps):
b1=update(b,X_hat,y_hat,eta)
b=update(b1,X_hat,y_hat,eta)
loss=propagation(b,X_hat,y_hat)["L"]
if i%10 == 0:
print ("Loss after iteration %i= %f" %(i, loss))
return b
# # 2
# + In this section based on the probability of the output in the binary classifier algorithm, complete the code by filling the None parts.
def predict(x,X_hat,y_hat,eta,steps=20):
b_optimal=b_opt(X_hat,y_hat,eta,steps,'random')
z=np.dot(x.T,b_optimal)
prob=sigmoid(z)
if None:
return 1
else:
return 0
def predict2(x,X_train,y_train,eta,steps,initialization):
N=X_train.shape[0]
one=np.ones((N,1))
X_hat=np.concatenate((one,X_train),axis=1)
b_optimal=b_opt(X_hat,y_train,eta,steps,initialization)
xnew=np.append(1,x).reshape(-1,1)
z=np.dot(xnew.T,b_optimal)
prob=sigmoid(z)
if None:
return 1
else:
return 0
# ## Making Data for Binary Classifier
def making_data(n_sample,shape_type):
ratio=2*np.pi/n_sample
t=np.arange(0,2*np.pi,ratio)
if shape_type=='circle':
noise=np.random.randn(t.shape[0])/8
x1=np.sin(t)
y1=np.cos(t)+noise
noise2=np.random.randn(t.shape[0])/8
x2=1.5*np.sin(t)
y2=1.5*np.cos(t)+noise2
elif shape_type=='wave':
noise=np.random.randn(t.shape[0])/2
x1=t
y1=np.cos(2*t)+noise
noise2=np.random.randn(t.shape[0])/2
x2=t
y2=1.5*np.cos(2*t)+2+noise2
elif shape_type=='linear':
noise=np.random.randn(t.shape[0])
x1=t
y1=t+noise
noise2=np.random.randn(t.shape[0])
x2=t
y2=t+3+noise2
elif shape_type=='cluster':
noise=np.random.randn(t.shape[0])/3
x1=noise*np.cos(t)
y1=noise*np.sin(t)+noise
noise2=np.random.randn(t.shape[0])/3
x2=noise2*np.cos(t)+1
y2=noise2*np.sin(t)+1.5+noise2
elif shape_type=='lorenz':
rot=(lambda theta:np.array([[np.cos(theta), -np.sin(theta)],[np.sin(theta), np.cos(theta)]]) )
noise=np.random.randn(t.shape[0])/4
x01=noise*np.cos(t)
y01=3*noise*np.sin(t)
xy0=np.array([[x01,y01]]).T
xy=np.dot(rot(np.pi/4),xy0)
x1=xy[0,:,:]
y1=xy[1,:,:]
noise2=np.random.randn(t.shape[0])/4
x02=noise2*np.cos(t)+1
y02=3*noise2*np.sin(t)+1.5
xy02=np.array([[x02,y02]]).T
xy2=np.dot(rot(-np.pi/4),xy02)
x2=xy2[0,:,:]
y2=xy2[1,:,:]
else:
raise Exception('Insert true shape name: circle or crescent or linear or wave or cluster or lorenz')
xy_red=np.column_stack((x1,y1))
l_red=np.ones((xy_red.shape[0],1)).astype(int)
red=np.column_stack((xy_red,l_red))
xy2_red=np.column_stack((x2,y2))
l_blue=np.zeros((xy2_red.shape[0],1)).astype(int)
blue=np.column_stack((xy2_red,l_blue))
total=np.concatenate([red,blue])
X=total[:,:2]
y=[item[0] for item in total[:,2:]]
y=np.array(y).astype(int)
return X, y
def plot_data(X,y):
df = pd.DataFrame(dict(x=X[:,0], y=X[:,1], label=y))
colors = {1:'red', 0:'blue'}
fig, ax = plt.subplots()
grouped = df.groupby('label')
for key, group in grouped:
group.plot(ax=ax, kind='scatter', x='x', y='y', label=key, color=colors[key])
return plt.show()
# # Split Data
def split(X,y,ratio):
total=np.column_stack((X,y))
np.random.shuffle(total)
M=int(X.shape[0]*ratio)
X=total[:,:2]
y=total[:,2]
y=y.reshape(-1,1)
train_X, test_X=X[:M], X[M:]
train_y, test_y=y[:M], y[M:]
return train_X, test_X, train_y, test_y
def plot_decision_boundary(X,y,ratio,filename):
train_X, test_X, train_y, test_y=split(X,y,ratio)
one=np.ones((train_X.shape[0],1))
xhat=np.column_stack([one, train_X])
yhat=train_y
yhat=yhat.reshape(-1,1)
w=b_opt(xhat,yhat,0.5,10,'random')
x1=np.arange(np.min(X[:,0]),np.max(X[:,0]),0.01)
x2=-w[1]/w[2]*x1-w[0]/w[2]
y.reshape(-1,)
df = pd.DataFrame(dict(x=X[:,0], y=X[:,1], label=y))
colors = {1:'red', 0:'blue'}
fig, ax = plt.subplots()
grouped = df.groupby('label')
for key, group in grouped:
group.plot(ax=ax, kind='scatter', x='x', y='y', label=key, color=colors[key])
plt.plot(x1,x2,'go-')
plt.savefig(filename,dpi=400)
return plt.show()
X1, y1=making_data(500,'linear')
X2, y2=making_data(500,'cluster')
X3, y3=making_data(500,'circle')
X4, y4=making_data(500,'wave')
X5, y5=making_data(500,'lorenz')
plot_data(X1,y1)
plot_decision_boundary(X1,y1,0.8,"HD1.jpg")
# # 3
# + Plot decision boundary for (X2,y2) with learning rate=0.01
# + Plot decision boundary for (X3,y3) with learning rate=0.1
# + Plot decision boundary for (X4,y4) with learning rate=0.5
# + Plot decision boundary for (X5,y5) with learning rate=0.0001
|
Deep Learning with Tensorflow/Logistic_Regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="RAoPUnFDSLD1"
import pandas as pd
# + id="kt2PI1ItDK2p"
from urllib import parse
from ast import literal_eval
import requests
# + id="IRFKzAXNDOLN"
def get_sise(code, start_time, end_time, time_from='day'):
get_param = { 'symbol':code, 'requestType':1, 'startTime':start_time, 'endTime':end_time, 'timeframe':time_from }
get_param = parse.urlencode(get_param)
url="https://api.finance.naver.com/siseJson.naver?%s"%(get_param)
response = requests.get(url)
return literal_eval(response.text.strip())
# + id="UZKxKEArDRKe"
data_sec = get_sise('005930', '20160101', '20220228', 'day')
# + id="__9Beh7aEEjD" outputId="904d85d7-390a-4a49-f2db-e7bbee6d0783" colab={"base_uri": "https://localhost:8080/"}
data_sec[0:3]
# + id="V4-b22Q7EQ1d"
data_df = pd.DataFrame(data_sec, columns=['Date', 'open', 'high', 'low', 'close', 'volume', 'foreigner'])
# + id="td8tU0x_LZWo" outputId="ab97fee2-9c4d-4907-9b2a-6d1d719995f7" colab={"base_uri": "https://localhost:8080/"}
data_df.dtypes
# + id="3Kb0znq2LpHk"
data_df.drop(data_df.index[0], inplace=True)
# + id="sZ6pVRHSMXpn" outputId="031a851a-49c7-4e0a-afaf-7f937c04b2bc" colab={"base_uri": "https://localhost:8080/", "height": 143}
data_df.head(3)
# + id="Dm-dV24ULwqw"
data_df.drop(data_df.columns[6], axis=1, inplace=True)
# + id="DCcHKwipL1_X"
data_df['Date'] = pd.to_datetime(data_df['Date'])
# + id="4oMRjKuvS92S"
data_df.sort_values(by=['Date'], inplace=True, ascending=False)
# + id="O4bo7kJUL45n" outputId="1ddcf070-9ce6-4981-bcb2-c35d2ef07254" colab={"base_uri": "https://localhost:8080/", "height": 143}
data_df.tail(3)
# + id="CMxn3JDzDUqQ"
data_df.set_index('Date', drop=True, inplace=True)
# + colab={"base_uri": "https://localhost:8080/"} id="x7i80B7kS5mT" outputId="622cce02-60b0-41f9-b27d-e8b0939e4769"
data_df.shape
# + id="OhjwLlwvNO7z" outputId="a982026e-e714-4674-d81f-8a4eb95c6d8c" colab={"base_uri": "https://localhost:8080/", "height": 175}
data_df.tail(3)
# + id="Gzqr0sfBIEOY" outputId="284a90d6-4bd9-44c7-aa62-da6f9887138d" colab={"base_uri": "https://localhost:8080/"}
data_df.shape
# + id="1X6MlGOwI_N5"
no_of_days = data_df.shape[0]
# + id="ewku1JonJFnR" outputId="f37bcf1c-567f-440d-b588-e8ee08597fba" colab={"base_uri": "https://localhost:8080/"}
no_of_days
# + id="vLV51RfYn6wy"
data_df['ma20'] = data_df['close'].rolling(window=20).mean() # 20일 이동평균
data_df['stddev'] = data_df['close'].rolling(window=20).std() # 20일 이동표준편차
data_df['upper'] = data_df['ma20'] + 2*data_df['stddev'] # 상단밴드
data_df['lower'] = data_df['ma20'] - 2*data_df['stddev'] # 하단밴드
data_df = data_df[19:] # 20일 이동평균을 구했기 때문에 20번째 행부터 값이 들어가 있음
# + colab={"base_uri": "https://localhost:8080/", "height": 455} id="LezYKd3hn5JK" outputId="620429b7-9a2a-4cf5-a363-845748a3ee88"
data_df
# + id="jE_MK_5HpIY5"
# + id="q5DDN2cNom4v"
import matplotlib.pyplot as plt
# + colab={"base_uri": "https://localhost:8080/", "height": 329} id="5OtLqrpzohxg" outputId="d01fb184-0fda-406f-f68d-ff05d97f0465"
plt.figure(figsize=(15, 5))
plt.plot(data_df.index, data_df['close'], label='close')
plt.plot(data_df.index, data_df['upper'], linestyle='dashed', label='Upper band')
plt.plot(data_df.index, data_df['ma20'], linestyle='dashed', label='Moving Average 20')
plt.plot(data_df.index, data_df['lower'], linestyle='dashed', label='Lower band')
plt.title(f'{"SEC"}(005930)BBands(20 Days, 2 std)')
plt.legend(loc='best');
# + id="k-Dht7VpSHkd"
|
Exercise/BBands_sec.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # DTM-based filtrations: demo
#
# <NAME>, https://raphaeltinarrage.github.io/
# <br/>
# Nov 2020
# This page describes the implementation and use of the method described in our paper *DTM-based Filtrations* https://arxiv.org/abs/1811.04757.
# ## Background
#
# The method described in this notebook has been designed to answer the following estimation problem: given a sample of a submanifold, potentially containing anomalous points, estimate the homology groups of the submanifold.
#
# The **DTM-filtration** is a filtration of the Euclidean space $\mathbb{R}^n$, based on a finite point cloud $X \subset \mathbb{R}^n$, and two parameters: $m \in [0,1)$ and $p \in [1, +\infty]$. The parameter $m$ corresponds to the DTM and the parameter $p$ corresponds to the radius functions.
# The DTM-filtration is a particular case of the weighted Čech filtrations.
#
# Let $\mu$ denote the empirical probability measure on $X$, and $\mathrm{d}_{\mu,m}\colon \mathbb{R}^n\rightarrow[0, +\infty)$ the DTM on $\mu$ with parameter $m$.
# The DTM-filtration $W[X,m,p]$ is defined as the collection of subsets $(W^t[X,m,p])_{t \geq 0}$ where
# $$W^t[X,m,p] = \bigcup_{x \in X} \overline{\mathcal{B}}\big(x,(t^p - \mathrm{d}_{\mu,m}^p(x))^\frac{1}{p}\big),$$
# and where $\overline{\mathcal{B}}(x,r)$ denotes the closed ball of center $x$ and radius $r$ if $r \geq 0$, or the emptyset if $r < 0$.
#
# The corresponding persistent module of $i^\text{th}$ homology is obtained by applying the $i^\text{th}$ homology functor to $V[X,m,p]$. Throughout this notebook, we will compute homology over the finite field $\mathbb{Z}/2\mathbb{Z}$.
#
# We shall implement the Vietoris-Rips version of the DTM-filtration, still refered here as the DTM-filtration.
# ## Datasets
#
# We present here two cases of application of the DTM-filtrations, and compare them with the usual Čech or Rips filtrations:
# - the circle $\mathbb{S}_1$ with anomalous points
# - the necklace (union of a circle and spheres)
# ## Package
#
# The functions are contained in the `Velour` package (https://pypi.org/project/velour/).
# <br/>
# It is based on the `Gudhi` library (https://gudhi.inria.fr/python/latest/).
import velour
import numpy as np
import matplotlib.pyplot as plt
# # First dataset: circle with anomalous points
# We start with a noisy sample $X$ of the unit circle in $\mathbb{R}^2$.
# We aim to recover the homology of the circle: Betti numbers $(\beta_0,\beta_1) = (1,1)$.
# +
N_observation = 150 #number of points sampled on the circle
N_anomalous = 50 #number of anomalous points
X = velour.SampleOnCircle(N_observation, N_anomalous) #sample with anomalous points
velour.PlotPointCloud(X) #displays the point cloud
# -
# We first build the usual Rips filtration on this dataset.
# +
filtration_max = 0.5 #maximal filtration value for Rips complex
st_Rips = velour.RipsComplex(X, filtration_max=filtration_max) #builds the Rips filtration
velour.PlotPersistenceDiagram(st_Rips) #displays the persistence diagram
# -
# On this diagram, $0$-homology points are represented in red, and $1$-homology in green.
# Note that the $1$-homology feature of the circle does not appear clearly here.
#
# We now illustrate a DTM-filtration on $X$.
# First, we compute the values of the DTM, with a given parameter $m \in [0,1)$.
# +
m = 0.1 #parameter for the DTM
DTM_values = velour.DTM(X,X,m) #computes the values of the DTM of parameter m
velour.PlotPointCloud(X, values = DTM_values) #draws X and the values of DTM
# -
# Notice that the DTM takes greater values on the anomalous points.
#
# Now we build a DTM-filtration, and plot its persistence diagram. Remind that filtration depends on a parameter $p \in [1, +\infty]$.
# +
p = 1 #parameter for the DTM-filtration
filtration_max = 1 #maximal filtration value for DTM-filtration
dimension_max = 2 #maximal dimension to expand the complex
st_DTM = velour.DTMFiltration(X, m, p, dimension_max = dimension_max, filtration_max=filtration_max)
#creates a DTM-filtration
velour.PlotPersistenceDiagram(st_DTM) #displays the persistence diagram
# -
# Here, the $1$-homology feature of the circle (green point) looks separated from the noise on the diagram. The DTM-filtration is able to recognize the underlying circle.
# We show below how greater values of $p$ tend to sparsify the persistence diagram.
# +
P = [1.5, 4.62, np.inf]
for p in P:
st_DTM = velour.DTMFiltration(X, m, p, dimension_max = dimension_max, filtration_max=filtration_max)
#creates a DTM-filtration
velour.PlotPersistenceDiagram(st_DTM) #displays the persistence diagram
plt.title('Parameter p = '+repr(p))
# -
# ## Second dataset: necklace
# We sample points on the union of a circle and three spheres in $\mathbb{R}^3$.
# Its Betti numbers are $(\beta_0,\beta_1, \beta_3) = (1,4,3)$.
# +
N_observation = 350 #number of points sampled on each component of the necklace
X = velour.SampleOnNecklace(N_observation) #samples points on necklace
velour.PlotPointCloud(X, plot_axis = True) #plots the point cloud
# -
# Instead of computing the Rips filtration on $X$, which contains `4*N_observation` vertices, we build the alpha-complex, which contains less simplices.
# +
st_alpha = velour.AlphaComplex(X) #creates an alpha-complex
velour.PlotPersistenceDiagram(st_alpha) #displays the persistence diagram
# -
# Blue points represent the $2$-homology.
# Between the values $t \in [0, 0.2]$ one can read the homology of the underlying object: Betti numbers $\beta_0 = 1$, $\beta_1 = 4$ and $\beta_2 = 3$.
# We can see this better on the barcodes. In order to ease the reading, only bars of length greater than `eps = 0.05` are plotted.
velour.PlotPersistenceBarcodes(st_alpha, eps = 0.05) #displays the persistence barcodes
# We now build a DTM-filtration on $X$.
# First, we illustrate how the value of the parameter $m$ can influence the DTM. When the underlying measure admits a density on a $d$-dimensional submanifold $\mathcal{M}$, there exists a constant $C>0$ such that the DTM on $\mathcal{M}$ is lower than $C m^\frac{1}{d}$.
# +
' DTM with parameter m = 0.05 '
m = 0.05
DTM_values = velour.DTM(X,X,m)
velour.PlotPointCloud(X, values = DTM_values)
plt.title('Parameter m = '+repr(m))
' DTM with parameter m = 0.3 '
m = 0.3
DTM_values = velour.DTM(X,X,m)
velour.PlotPointCloud(X, values = DTM_values)
plt.title('Parameter m = '+repr(m))
# -
# It appears that, for a small value of $m$, the DTM takes lower values on the circle than the spheres, while for a greater value of $m$, the DTM takes high values on the circle and outside the spheres.
# This observation is consistent with the previous remark: the DTM on the circle is lower than $C m$, while on the sphere it is lower than $C' m^\frac{1}{2}$.
# Since the number of points of $X$ is large, we use the function `velour.AlphaDTMFiltration()`.
# This construction only is heuristic, but reduces the number of simplices drastically.
# We start with the parameter $m = 0.05$.
# +
m = 0.05
p = 1
dimension_max = 3
st_DTM_alpha = velour.AlphaDTMFiltration(X, m, p, dimension_max) #builds the alpha-DTM filtration
velour.PlotPersistenceDiagram(st_DTM_alpha) #displays the persistence diagram
# -
# One can still read the homology of $X$ on this diagram, for the values of $t$ close to 1 (Betti numbers $(\beta_0,\beta_1,\beta_2) = (1,4,3)$). This can also be read on the barcodes. Only bars of length larger than $0.2$ are plotted.
velour.PlotPersistenceBarcodes(st_DTM_alpha, tmax = 2, eps = 0.2) #displays the persistence barcodes
# We now compute the DTM-filtration with the parameter `m = 0.3`.
# +
m = 0.3
p = 1
dimension_max = 3
st_DTM_alpha = velour.AlphaDTMFiltration(X, m, p, dimension_max) #builds the alpha-DTM filtration
velour.PlotPersistenceDiagram(st_DTM_alpha) #displays the persistence diagram
# -
# On this diagram, the homology of the spheres does not appear anymore, and one reads $(\beta_0,\beta_1,\beta_2) = (3,0,0)$ on the first part of the diagram, and $(\beta_0,\beta_1,\beta_2) = (1,1,0)$ on the second part.
# See also the barcodes below (only bars of length larger than $0.25$ are represented).
velour.PlotPersistenceBarcodes(st_DTM_alpha, tmax = 2, eps = 0.25) #displays the persistence barcodes
# Let us compare the DTM-filtrations for several values of $p$.
# +
m = 0.05
P = [1.2, 6.62, np.inf]
dimension_max = 3
for p in P:
st_DTM_alpha = velour.AlphaDTMFiltration(X, m, p, dimension_max) #builds the alpha-DTM filtration
velour.PlotPersistenceDiagram(st_DTM_alpha) #displays the persistence diagram
plt.title('Parameter p = '+repr(p))
# -
# Notice that, for large values of $p$, the $2$-homology (in blue) of the underlying spheres does not appear clearly on the diagram. Information have been lost.
# ## Take-home message
# Dependance of the DTM on its parameters:
# - higher values of $p$ tend to simplify the persistence diagram,
# - different values of $m$ may highlight various areas of the dataset.
#
# This simple implementation can be used as follows:
# ```
# # X is a Nxn np.array
# m = .1
# p = 1
# dimension_max = 2
#
# st = velour.DTMFiltration(X, m, p, dimension_max) #builds the DTM-filtration
# gudhi.plot_persistence_diagram(st.persistence()) #displays the persistence diagram
# ```
|
Demo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/john-s-butler-dit/Numerical-Analysis-Python/blob/master/Chapter%2010%20-%20Hyperbolic%20Equations/1001_Wave%20Equation.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="CDO6MV51DoQN"
# # Wave Equation
# #### <NAME> <EMAIL> [Course Notes](https://johnsbutler.netlify.com/files/Teaching/Numerical_Analysis_for_Differential_Equations.pdf) [Github](https://github.com/john-s-butler-dit/Numerical-Analysis-Python)
#
# ## Overview
# This notebook will implement the Forward Euler in time and Centered in space method to appoximate the solution of the wave equation.
#
# ## The Differential Equation
# Condsider the one-dimensional hyperbolic Wave Equation:
# \begin{equation} \frac{\partial u}{\partial t} +a\frac{\partial u}{\partial x}=0,\end{equation}
# where $a=1$, with the initial conditions
# \begin{equation} u(x,0)=1-\cos(x), \ \ 0 \leq x \leq 2\pi. \end{equation}
# with wrap around boundary conditions.
#
#
# + id="v8FbXNtIDoQQ"
# LIBRARY
# vector manipulation
import numpy as np
# math functions
import math
# THIS IS FOR PLOTTING
# %matplotlib inline
import matplotlib.pyplot as plt # side-stepping mpl backend
import warnings
warnings.filterwarnings("ignore")
# + [markdown] id="nvUNuPPPDoQR"
# ## Discete Grid
# The region $\Omega$ is discretised into a uniform mesh $\Omega_h$. In the space $x$ direction into $N$ steps giving a stepsize of
# \begin{equation} \Delta_x=\frac{2\pi-0}{N},\end{equation}
# resulting in
# \begin{equation}x[j]=0+j\Delta_x, \ \ \ j=0,1,...,N,\end{equation}
# and into $N_t$ steps in the time $t$ direction giving a stepsize of
# \begin{equation} \Delta_t=\frac{1-0}{N_t}\end{equation}
# resulting in
# \begin{equation}t[n]=0+n\Delta_t, \ \ \ n=0,...,K.\end{equation}
# The Figure below shows the discrete grid points for $N=10$ and $Nt=100$, the known boundary conditions (green), initial conditions (blue) and the unknown values (red) of the Heat Equation.
# + id="n5PyInBnDoQS" outputId="2b9405c2-1244-4918-b21f-357090e3000e" colab={"base_uri": "https://localhost:8080/", "height": 324}
N=20
Nt=100
h=2*np.pi/N
k=1/Nt
time_steps=100
time=np.arange(0,(time_steps+.5)*k,k)
x=np.arange(0,2*np.pi+h/2,h)
X, Y = np.meshgrid(x, time)
fig = plt.figure()
plt.plot(X,Y,'ro');
plt.plot(x,0*x,'bo',label='Initial Condition');
plt.xlim((-h,2*np.pi+h))
plt.ylim((-k,max(time)+k))
plt.xlabel('x')
plt.ylabel('time (ms)')
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.title(r'Discrete Grid $\Omega_h$ ',fontsize=24,y=1.08)
plt.show();
# + [markdown] id="OnIx7JtEDoQT"
# ## Initial Conditions
#
# The discrete initial conditions is,
# \begin{equation} w[0,j]=1-\cos(x[j]), \ \ 0 \leq x[j] \leq \pi, \end{equation}
# The figure below plots values of $w[j,0]$ for the inital (blue) conditions for $t[0]=0.$
# + id="39auVS78DoQT" outputId="856e4718-de10-4f22-dc9f-08d5bdf297bd" colab={"base_uri": "https://localhost:8080/", "height": 304}
w=np.zeros((time_steps+1,N+1))
b=np.zeros(N-1)
# Initial Condition
for j in range (0,N+1):
w[0,j]=1-np.cos(x[j])
fig = plt.figure(figsize=(12,4))
plt.plot(x,w[0,:],'o:',label='Initial Condition')
plt.xlim([-0.1,max(x)+h])
plt.title('Intitial Condition',fontsize=24)
plt.xlabel('x')
plt.ylabel('w')
plt.legend(loc='best')
plt.show()
# + [markdown] id="Mw_bC8CVDoQU"
# ## Boundary Conditions
# To account for the wrap-around boundary conditions
# \begin{equation}w_{-1,n}=w_{N,n},\end{equation}
# and
# \begin{equation}w_{N+1,n}=w_{0,n}.\end{equation}
# + id="NuHcK-e3DoQV"
xpos = np.zeros(N+1)
xneg = np.zeros(N+1)
for j in range(0,N+1):
xpos[j] = j+1
xneg[j] = j-1
xpos[N] = 0
xneg[0] = N
# + [markdown] id="O8lJ-AnYDoQW"
# ## The Explicit Forward Time Centered Space Difference Equation
# The explicit Forward Time Centered Space difference equation of the Wave Equation is,
# \begin{equation}
# \frac{w^{n+1}_{j}-w^{n}_{j}}{\Delta_t}+\big(\frac{w^n_{j+1}-w^n_{j-1}}{2\Delta_x}\big)=0.
# \end{equation}
# Rearranging the equation we get,
# \begin{equation}
# w_{j}^{n+1}=w^{n}_{j}-\lambda a(w_{j+1}^{n}-w_{j-1}^{n}),
# \end{equation}
# for $j=0,...10$ where $\lambda=\frac{\Delta_t}{\Delta_x}$.
#
# This gives the formula for the unknown term $w^{n+1}_{j}$ at the $(j,n+1)$ mesh points
# in terms of $x[j]$ along the nth time row.
# + id="RHwq9gEDDoQX"
lamba=k/h
for n in range(0,time_steps):
for j in range (0,N+1):
w[n+1,j]=w[n,j]-lamba/2*(w[n,int(xpos[j])]-w[n,int(xneg[j])])
# + [markdown] id="TsVfD7sEDoQY"
# ## Results
# + id="hOx5ScRMDoQY" outputId="b7e48e97-cb43-4b66-95f6-31ec49aafbd5" colab={"base_uri": "https://localhost:8080/", "height": 487}
fig = plt.figure(figsize=(12,6))
plt.subplot(121)
for n in range (1,time_steps+1):
plt.plot(x,w[n,:],'o:')
plt.xlabel('x[j]')
plt.ylabel('w[j,n]')
plt.subplot(122)
X, T = np.meshgrid(x, time)
z_min, z_max = np.abs(w).min(), np.abs(w).max()
plt.pcolormesh( X,T, w, vmin=z_min, vmax=z_max)
#plt.xticks(np.arange(len(x[0:N:2])), x[0:N:2])
#plt.yticks(np.arange(len(time)), time)
plt.xlabel('x[j]')
plt.ylabel('time, t[n]')
clb=plt.colorbar()
clb.set_label('Temperature (w)')
#plt.colorbar()
plt.suptitle('Numerical Solution of the Wave Equation',fontsize=24,y=1.08)
fig.tight_layout()
plt.show()
# + id="IIVsYob6DoQZ"
# + id="Kql3zdeNDoQZ"
|
Chapter 10 - Hyperbolic Equations/1001_Wave Equation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="m7I35Gzz78o1" colab_type="text"
# # Modelling Solar generation across Multiple Sites
#
# This example shows how `timeserio` helps building deep learning models for time series forecasting. Especially,
# we deal with the case of many related timeseries.
#
# We demonstrate some core functionality and concepts, without striving for model accuracy or seeking out additional features like historic weather forecasts.
#
# We will be using the dataset on solar (photo-voltaic, PV) generation potential across Europe, as collected by [SETIS](https://setis.ec.europa.eu/EMHIRES-datasets). The dataset presents solar generation, normalized to the solar capacity installed as of 2015.
# + [markdown] id="xmOm7b2v78o3" colab_type="text"
# ## Download the data
# + id="ZF43VGYC78o5" colab_type="code" outputId="80dd4565-2506-46e6-9c2d-a46380cf595d" colab={"base_uri": "https://localhost:8080/", "height": 238}
# !mkdir -p ~/tmp/datasets; cd ~/tmp/datasets; wget https://setis.ec.europa.eu/sites/default/files/EMHIRES_DATA/Solar/EMHIRESPV_country_level.zip; unzip -o EMHIRESPV_country_level.zip; rm EMHIRESPV_country_level.zip
# + [markdown] id="2wBXQhv978o-" colab_type="text"
# ## Download data and save in a more performant format
# + id="j-Uau75L78o_" colab_type="code" colab={}
import pandas as pd
import numpy as np
# + id="ZkBVOL2R78pD" colab_type="code" outputId="a07d3694-d82c-4527-db74-b0e4f023b8ec" colab={"base_uri": "https://localhost:8080/", "height": 51}
# %%time
df = pd.read_excel("~/tmp/datasets/EMHIRESPV_TSh_CF_Country_19862015.xlsx")
# + id="D02Lit7v78pH" colab_type="code" outputId="2c9ade8d-e6d9-47b0-ab5b-f785ce5d3a6e" colab={"base_uri": "https://localhost:8080/", "height": 142}
df.head(3)
# + [markdown] id="p84G29sj78pL" colab_type="text"
# ### Reshape data to tall format
# We add a `country` column to identify each table row.
# + id="tST-xAnJ78pM" colab_type="code" colab={}
id_vars = ['Time_step', 'Date', 'Year', 'Month', 'Day', 'Hour']
country_vars = list(set(df.columns) - set(id_vars))
df_tall = pd.melt(df, id_vars=id_vars, value_vars=country_vars, var_name="country", value_name="generation")
# + id="P9UUdSRo_dOz" colab_type="code" outputId="9d5742c6-0253-440c-b70d-ceb1ee6b9499" colab={"base_uri": "https://localhost:8080/", "height": 359}
df_tall.head(10)
# + [markdown] id="YFrPxsA578pi" colab_type="text"
# ## Data Exploratory
# + id="BR8RNvXO78pj" colab_type="code" colab={}
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# + id="egEOQNwo78pn" colab_type="code" outputId="c6ccd26c-584f-4873-bc2b-f387ef3b0262" colab={"base_uri": "https://localhost:8080/", "height": 51}
# %%time
df = df_tall
# + id="3x96zbR278pq" colab_type="code" outputId="f6e82559-ecc0-40dd-d83e-ba4bcc4b9f88" colab={"base_uri": "https://localhost:8080/", "height": 34}
print(' '.join(sorted(df['country'].unique())))
# + id="-hlmipwZ78pu" colab_type="code" colab={}
plot_countries = ['ES', 'UK', 'FI', ]
# + id="pHL7XfO078pz" colab_type="code" outputId="1cc5705e-d0ad-4b04-b0a1-6a4c13c35ee2" colab={"base_uri": "https://localhost:8080/", "height": 364}
plt.figure(figsize=(15, 3))
sns.lineplot(
data=df[(df['Time_step'] < 500) & (df['country'].isin(plot_countries))],
x='Date', y='generation', hue='country',
)
# + id="G88lEZ5u78p3" colab_type="code" colab={}
df_monthly = df.groupby(['Month', 'country']).mean().reset_index()
# + id="V3NH2xER78p6" colab_type="code" outputId="fccd87cc-987d-4171-9af0-377dc8b81e3d" colab={"base_uri": "https://localhost:8080/", "height": 242}
plt.figure(figsize=(15, 3))
sns.lineplot(
data=df_monthly[df_monthly['country'].isin(plot_countries)],
x='Month', y='generation', hue='country',
)
# + id="LgZqgorv78p-" colab_type="code" colab={}
df_daily = df.groupby(['Hour', 'country']).mean().reset_index()
# + id="IQsDAG8K78qB" colab_type="code" outputId="84d2221e-49fd-4bef-8090-3d66f4ae54c8" colab={"base_uri": "https://localhost:8080/", "height": 242}
plt.figure(figsize=(15, 3))
sns.lineplot(
data=df_daily[df_daily['country'].isin(plot_countries)],
x='Hour', y='generation', hue='country',
)
# + id="EY6WEObB78qE" colab_type="code" colab={}
df_yearly = df.groupby(['Year', 'country']).mean().reset_index()
# + id="KOi2qIRf78qI" colab_type="code" outputId="4a42e7dd-040f-4d18-e146-871bbabdb29a" colab={"base_uri": "https://localhost:8080/", "height": 242}
plt.figure(figsize=(15, 3))
sns.lineplot(
data=df_yearly[df_yearly['country'].isin(plot_countries)],
x='Year', y='generation', hue='country',
)
# + [markdown] id="w1l0YV4U78qL" colab_type="text"
# ### Preliminary observations
#
# The timeseries for different countries exhibit a lot of similarity - they will have similar daily and seaonal shapes. At the same time, the curves have different scaling (due to latitudes and weather), and different time offsets (due to longitude). We can build models to incorporate these as external features, or learn the relevant features from the available data only!
# + [markdown] id="2_xsYzthwUiV" colab_type="text"
# ### Store to parquet
# [Apache Parquet](https://parquet.apache.org/documentation/latest/) is a much preferred data format for columnar numerical data - it is much faster to read (see below), is fully compatible with tools like `pandas` and Spark, and allows easy partitioning of large datasets.
# + id="tpJHQuHjwUiW" colab_type="code" outputId="95a495fb-161d-4fdb-a2cf-4fb5847f82f0" colab={"base_uri": "https://localhost:8080/", "height": 51}
# %%time
df_tall.to_parquet("/content/tall.parquet")
# + id="UzQcIUAlCRGW" colab_type="code" outputId="d5d5b80e-6591-426c-f8b7-08f5ba0d24dc" colab={"base_uri": "https://localhost:8080/", "height": 68}
# !ls /content -l
# + id="bF9otCaAwUij" colab_type="code" colab={}
from google.colab import files
files.download("/content/tall.parquet")
# + [markdown] id="ARaXenQwwUia" colab_type="text"
# ### Store to partitioned parquet (For Distributed Environment)
#
#
# ```python
# # # %%time
# df_tall.to_parquet("~/tmp/datasets/EMHIRESPV_TSh_CF_Country_19862015_partitioned/", partition_cols=["country"])
#
# # # !tree -h --filelimit=10 ~/tmp/datasets
# ```
|
MultiModel/1_Solar_Generation_Data_Preparation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
x = np.array([["Germany","France"],["Berlin","Paris"]])
y = np.array([["Hungary","Austria"],["Budapest","Vienna"]])
x.shape
x
y.shape
y
# The default is row-wise concatenation for a 2D array
print('Joining two arrays along axis 0')
np.concatenate((x,y))
# Column-wise
print('Joining two arrays along axis 1')
np.concatenate((x,y), axis = 1)
a = np.array([1, 2, 3])
b = np.array([2, 3, 4])
# one array in the top if another
np.stack((a, b))
studentId = np.array([1,2,3,4])
name = np.array(["Alice","Beth","Cathy","Dorothy"])
scores = np.array([65,78,90,81])
np.stack((studentId, name, scores))
np.stack((studentId, name, scores)).shape
np.stack((studentId, name, scores), axis =1)
np.stack((studentId, name, scores), axis =1)
np.stack((studentId, name, scores), axis =1).shape
# vstack
# Stacks row wise
np.vstack((studentId, name, scores))
# hstack
# Stacks column wise
np.hstack((studentId, name, scores))
np.hstack((studentId, name, scores)).shape
|
VectorStacking_m02_demo07.ipynb
|