code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
from scipy import signal
import scipy
import scipy.io as sio
import copy
import pylab as pl
import time
from IPython import display
# ## Chirp parameters
start_freq = 770000
band_freq = 80000
duration = 0.0004
samples_one_second = 10000000
rate = samples_one_second / start_freq
sample = start_freq * rate
npnts = int(sample * duration)
print("Smpale per cicle", rate, "Sample for one second", samples_one_second, "Total semples", npnts)
# ## Create the chirp
# +
timevec = np.linspace(0, duration, npnts)
adding_freq = np.linspace(0, band_freq, npnts)
chirp = np.sin(2*np.pi * (start_freq + adding_freq * 0.5) * timevec)
# chirp = signal.chirp(timevec,f0=start_freq,t1=duration,f1=start_freq + band_freq)
ciclestoshow = int(rate * 30)
plt.figure(figsize=(16,4))
plt.subplot(211)
plt.plot(timevec[:ciclestoshow], chirp[:ciclestoshow])
plt.title("Time domain Low")
plt.subplot(212)
plt.plot(timevec[-ciclestoshow:], chirp[-ciclestoshow:])
plt.title("Time domain High")
plt.tight_layout()
plt.show()
# +
hz = np.linspace(0, sample / 2, int(np.floor(npnts / 2) + 1))
spectrum = 2*np.abs(scipy.fftpack.fft(chirp)) / npnts
plt.figure(figsize=(16,4))
plt.stem(hz,spectrum[0:len(hz)])
plt.xlim([start_freq - band_freq,start_freq + band_freq*3])
plt.title('Power spectrum')
plt.show()
# -
# # Distance of an object in km
kilometer = 16.8
# # RX TX Chirp Mix
# +
light_speed_km = 300000
print('Theoretical max distance of a chirp of', light_speed_km * duration / 2, 'km')
smallest_measure_distance = light_speed_km * (1 / samples_one_second) / 2
print('smallest measure distance',smallest_measure_distance, 'km')
shift = int((1 / smallest_measure_distance) * kilometer)
print('shift', shift, 'out of', npnts, 'sample points, for a distance of', np.round(shift * smallest_measure_distance, 3), 'km')
distance_per_herz = (light_speed_km * duration / 2) / band_freq
print('Friquncy domain per distance', distance_per_herz, 'km per herz')
print()
chirp_time = np.linspace(0, duration, npnts)
chirp_freq = np.linspace(0, band_freq, npnts)
plt.plot(chirp_time, chirp_freq, label='TX chirp')
plt.plot(chirp_time[shift:], chirp_freq[:-shift], label='RX chirp')
plt.plot([chirp_time[shift],chirp_time[shift]], [0,chirp_freq[shift]], 'g-' , label='St Frequency {} hz'.format(np.round(chirp_freq[shift])))
plt.plot(chirp_time[shift],0, 'gv')
plt.plot(chirp_time[shift],chirp_freq[shift], 'g^')
plt.plot([chirp_time[shift],chirp_time[-1]],[chirp_freq[shift],chirp_freq[shift]], 'c:', label='IF signal')
plt.plot([chirp_time[-1]],[chirp_freq[shift]], 'c>')
plt.ylabel('Frequency')
plt.xlabel('Time')
plt.title('RX Reflection of corresponding object')
plt.legend()
plt.tight_layout()
plt.show()
# -
# ## Object detection
# +
# local synthesizer
tx = chirp[shift:]
# Object reflection signal
rx = chirp[:-shift]
# mixing all frequencies
mix = tx * rx
plt.figure(figsize=(16,4))
plt.plot(timevec[:-shift], mix)
plt.title("Time domain mix shift chirp")
plt.xlabel('Time')
plt.show()
accuracy = int(npnts * 5)
hz = np.linspace(0, sample / 2, int(np.floor(accuracy / 2) + 1))
# Get IF frequencies spectrum
fftmix = scipy.fftpack.fft(mix, n=accuracy)
ifSpectrum = np.abs(fftmix) / accuracy
ifSpectrum[1:] = ifSpectrum[1:] * 2
# Find local high as detection
hz_band_freq = hz[hz <= band_freq]
testIifSpectrum = ifSpectrum[:len(hz_band_freq)]
localMax = np.squeeze(np.where( np.diff(np.sign(np.diff(testIifSpectrum))) < 0)[0]) + 1
# Adjust trigger level
meanMax = testIifSpectrum[localMax].mean()
maxSpectrum = testIifSpectrum[localMax].max()
trigger = maxSpectrum * .8
# Frequency detection
valid_local_indexs = localMax[testIifSpectrum[localMax] > trigger]
colors = ['r','g','c','m','y']
plt.figure(figsize=(16,4))
plt.plot(hz_band_freq, testIifSpectrum,'b-o', label='spectrum')
# Convert chirp shift to distance
dist = smallest_measure_distance * shift
# Convert distance to frequency
scale = dist / distance_per_herz
plt.plot([scale,scale], [maxSpectrum, 0],'g--', label='closest distance {}'.format(np.round(dist,3)))
plt.plot([hz_band_freq[0],hz_band_freq[-1]],[trigger,trigger],'--',label='trigger level {}'.format(np.round(trigger,3)))
for i in range(len(valid_local_indexs)):
pos = valid_local_indexs[i]
freq = hz_band_freq[pos]
spect_val = testIifSpectrum[pos]
plt.plot(freq, spect_val,colors[i] + 'o', label='detection frq {} distance {}'.format(freq, np.round(freq * distance_per_herz, 2)))
plt.xlim([0,hz[valid_local_indexs[-1]] * 2])
plt.title("Friquncy domain IF signal")
plt.xlabel('Frequency')
plt.legend()
plt.show()
# +
# Low pass filter mixed IF
lowCut = band_freq * 1.2
nyquist = sample/2
transw = .1
order = npnts
# To avoid edge effect
longmix = np.concatenate((mix[::-1],mix,mix[::-1]))
# order must be odd
if order%2==0:
order += 1
shape = [ 1, 1, 0, 0 ]
frex = [ 0, lowCut-lowCut*transw, lowCut, nyquist ]
# define filter shape
# filter kernel
filtkern = signal.firls(order,frex,shape,fs=sample)
filtkern = filtkern * np.hanning(order)
nConv = len(filtkern) + len(longmix) - 1
lenfilter = len(filtkern)
half_filt_len = int(np.floor(lenfilter / 2))
filtkernFft = scipy.fftpack.fft(filtkern,n=nConv)
rowFft = scipy.fftpack.fft(longmix,n=nConv)
ifSignal = np.real(scipy.fftpack.ifft(rowFft * filtkernFft))
ifSignal = ifSignal[half_filt_len:-half_filt_len]
ifSignal = ifSignal[len(mix):-len(mix)]
siglen = len(ifSignal)
plt.figure(figsize=(16,4))
plt.plot(timevec[:siglen] * duration, ifSignal)
plt.title("Time domain IF signal (Mix low pass filter)")
plt.xlabel('Time')
plt.show()
# -
|
Notebook/1-Radar-rx-tx-mix.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Enconding Natural Data
#
# One of the primary challeges in natural language processing is properly encoding the text so that mathematical models such as neural networks can extract meaning from the text, and process it into some kind of result. The field has evolved quite a lot over time, and we're going to briefly present a few of the once-common strategies for language encoding then spend some time talking about the current favorite method: embeddings.
#
# ## Rules Based Systems
#
# At first there were rules based systems. Like many other systems at the time, these were "expert systems" designed with extensive help from linguists and grammatician. These systems would look at a sentnce and try to identify critical aspects of a sentence (the subject, object, key verb...), label the words into types (adjective, noun, verb adverb), and map their function (which adjectives describe which nouns?).
#
# These expert systems could produce some data structure representing the sentence or document, and then these structures could be used for some custom built purpose (grammar check, chat bot...)
#
# Unlike ML systems, these systems didn't have any real purpose for statistical based learning, and as a result the data structures didn't need to be strictly numerically based, nor did they have to explicitly encode individual words or longer sentences into pure numeric formats.
#
# ## Bag of Words
#
# Once statistical learning methods began advancing, NLP researchers needed pure numeric encodings that could meaningfully represent, words, sentences, or other strings of text. Some simple strategies like the classic one-hot-encoding were fleetingly popular — but the need for a vector the size of the entire vocabulary just to represent a single word was prohibitively costly for all but the simplest use cases.
#
# The "Bag of Words" encoding came next. Instead of representing single words at a time, this strategy reduces an entire string of text into a vector. Again the vector is the lenght of the vocabulary, but instead of one-hot the individual values are the number of times that word appears in a the text.
#
# For example, say our vocabulary is only 5 words:
#
# Hello, live, work, to, friend
#
# Each of these words is represented by a position in a vector, and we loop through the text to create the vector for our text.
#
# The sentence, "Hello friend" becomes the vector `[1, 0, 0, 0, 1]`
#
# "Live to work" and "work to live" both become the vector `[0, 1, 1, 1, 0]`
#
# For some simple tasks such an encoding can work reasonably well especially if the input texts are always short. For example, mapping Tweets to a binary positive/negative sentiment system. But such a simple system clearly discards much of the semantic meaning of the sentence by completely ignoring the order of the words. It also struggles with words like "live" that have multiple possible meanings ("The new feature is going live tomorrow." vs "She is going to live!")
#
# Such an encoding can indeed be used with a standard ANN. That said, none of the state of the art research is proceeding down this path.
#
#
# ## TF-IDF
#
# TF-IDF, or Term Frequencey Inverse Document Frequency is a way to turn an individual word into a numeric value rather than process a series of words the way Bag of Words would. The TF-IDF value is a representation of how common a word is within a particular document as well as how common the word is accross all documents in an entire corpus or dataset. We won't be using it, but you can view the [mathematical details on Wikipedia](https://en.wikipedia.org/wiki/Tf%E2%80%93idf) if you're curious
#
# While this is useful in some information retrieval contexts for many natural language tasks such as machine translation and sentiment analysis, information about a words commonality isn't especially helpful. Researchers needed a way to create a numeric representation of individual words that could somehow capture the semantic meaning of those words...
#
# ## Word Embeddings
#
# In 2013, a team of researchers at Google published two papers describing Word2Vec, a neural network that transformed individual words into vectors that could represent the word's semantic meaning. Word2Vec formed a strong foundation for reseach, and ultimately led to the creation of a now-standard tool: Word Embedding layers.
#
# Word embedding layers provide a generic interface for the first layer of any neural network that wants to process one word at a time as input. Their function is simple: Embedding layers act as a lookup table mapping a word from our vocabulary into a dense vector of a chosen length representing that word.
#
# The values associated with each position in the word-vector are learned during back-propagation just as the weights in a Dense layer would be, but don't require an entire matrix multiply as each word maps to a single vector of the matrix representing the entire vocabulary. This difference saves computational time, but learns through backpropagation very similarly to a Dense layer.
#
# Sometimes we may use a pre-trained network (such as Word2Vec) to create the word embeddings. Although it is now quite common to train an embedding layer as part of the network, which allows the word embeddings to learn patterns that are specific to the task at hand.
#
# Regardless, the result is a lookup that maps words to vectors and (if it works as expected) words that are closely related in the dataset result in vectors that are near each other in vector space. When it REALLY works, these embedding vectors can be thought of as a rich set of features extracted from the word.
#
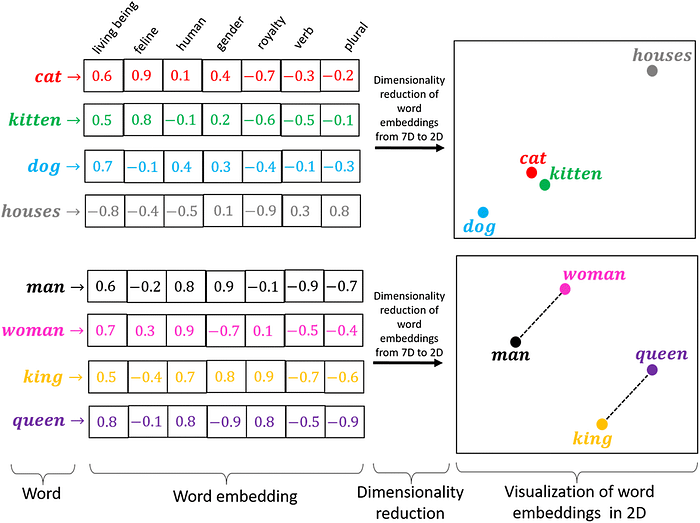
# Analysis of the embeddings themselves can be done. For example computing the cosine similarity between two words in the resulting vector space frequently reveals that semantically related words like cat and kitty are close to each other in the resulting vector space, as explained here [https://medium.com/@hari4om/word-embedding-d816f643140](https://medium.com/@hari4om/word-embedding-d816f643140)
#
# 
#
# > image from linked reading
|
08-recurrent-neural-networks/01-encoding-textual-data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Getting started
# ### Ranges
# A range is a rectangular group of cells. We define ranges from the top left to the bottom right. For example A1:B2.
# ### Formulas
# In B2, add 29 to 121. You should use the + operator. **It's really important here to put the = in front of the cell**.
# =29+121
# ### Exponents and parentheses
# = (7 + 2)^2 - 16
# ### Percentage
# = 4*21%
#
# ### Comparison operators
# = 0.4 < 1 will show a value of FALSE
# ### Data types: text and numbers
# ="Eggs"+1 will give an error. Hover over the cell to see the details.
#
# Fill in '= 0.40 * 21% in F2, **mind the ' in front**. Spaces are important here. By using the single quote, you ensure the formula will not be calculated.
#
# plain text, will be used if there's no other type recognized. You can force this with ', e.g.: 'text, or '2. Aligned to the left by default.
# ### Data types: currency and date
# Google Sheets tries to interpret the correct data type by default. For example, if you start a cell with $, it will be interpreted as a currency. Sometimes, it might be necessary to manually change a data type. You do this by selecting the cells you want to change and click on Format > Number in the menu bar.
#
# Select the values in D2:D5. These currently have type date. Using the format menu, change the format to use / instead of - to separate the date parts. Use the 'More date and time formats...' option in 'Format > Number > More formats'.
#
# Select the values in E2:E5. They currently have type currency, using the dollar $ sign. Change the format of these values to currency (rounded) You can use the format menu, or any other shortcuts you'd like.
# ### Data types: logic
# When you enter true or false in a cell, it's also recognized as a logical. Logicals are case insensitive, but Google Sheets will replace the value you entered by the capitalized logical: TRUE or FALSE.
#
# =true <> false, fill a cell with this will give a TRUE. <> is != in C++.
# ## References
# * Reference and absolute reference
# * Auto filling
# * Reactivity
# ### Cell references
# For example, you could make a reference to cell A1 in another cell: = A1. The cell will then always be the same as what's in A1. As cell references are case insensitive, = a1 would be fine as well.
#
# In D2 cell, write = C2
# In E2 cell, write = D2
# If C2 is changed to 20, then D2 and E2 will be changed to 20 automatically.
# ### Circular references
# But what happens when the referencing cell and the referenced cell are the same? In other words, what happens if a cell references itself? It will have problems.
# ### Copying references
# Referencing becomes especially useful if you start copying the references around. When you copy formulas containing references to neighbouring cells, the references will shift along.
#
# Assume you have = A1 in the cell with address B1. If you copy this cell to B2, the reference will change to = A2.
#
# To copy a cell to neighbouring cells, select the cell and move your mouse to the lower right corner. **The cursor should change to a "+"-sign. Drag and drop to where you want to copy.**
#
# ### Copying horizontally
# Just like you can copy cells vertically, you can just as well copy cells horizontally. Once again, when copying references, they will shift along. This time they will change columns.
#
# ### Copying columns
# Copy columns of references all at once!
#
# Start by filling in a reference to B2 in D2. Make sure to start the formula with a =.
#
# Copy D2 down to D11 vertically by dragging it downwards.
#
# Make sure D2:D11 is selected, copy this column one to the right to E2:E11, also by dragging.
# ### Mathematical operators and references
# Do calculations with references.
#
# Fill in the land area of China in miles in D2. To convert square kilometers to square miles, divide the values by 2.59. Use a reference to C2. That is: = C2/2.59
# Use the copying technique you learned in the previous exercises to fill down the square miles in D2:D11.
# ### Percentages and references
# First, fill in = B2 * 1.12% in D2. Then find the lower right corner to copy the values until D11 by dragging.
# Fill in = B2 / C2 in E2, then use your copy skills to fill out the columns until E11.
# ### Comparison operators and references
# In F2:F11, fill in a column which is TRUE each time the density is bigger than the world average and FALSE otherwise. The world average is 51. That is, fill with: =E2 > 51
#
# In G2:G11, fill in a column which is TRUE each time the continent is equal to "Asia" and FALSE otherwise. That is fill with: =D2="Asia"
# ### Absolute references
# If fill one cell with: = B2/B12 * 100, and they copy by dragging down, and then we will see B3/B13 * 100, B4/B14 * 100,... However, B13, B14, ... are not existed. So there will be error.
#
# We actually want B12 not changed in all the cells. So adding a `$` before B makes column B absolute (not changing when copying). Then adding a `$` before 12 makes row 12 absolute. That is, we fill the first cell with `=B2/$B$12 * 100`. When dragging down, we will have `B2/B12*100, B3/B12*100, B4/B12*100,...`
#
# ### Absolute references: row
#
# Fill D2 with `=C2/C$12 * 100.` The `$` only makes the row absolute.
# ### Absolute references: column
#
# Fill D2 with `=C2/$`C12` * 100`. The `$` only makes the row absolute.
|
dataManipulation/spreadsheets/Part I -- Spreadsheets basics/.ipynb_checkpoints/spreadsheet basics-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Load data
# +
import pickle
train_filename = "C:/Users/behl/Desktop/lung disease/train_data_sample_gray.p"
(train_labels, train_data, train_tensors) = pickle.load(open(train_filename, mode='rb'))
valid_filename = "C:/Users/behl/Desktop/lung disease/valid_data_sample_gray.p"
(valid_labels, valid_data, valid_tensors) = pickle.load(open(valid_filename, mode='rb'))
test_filename = "C:/Users/behl/Desktop/lung disease/test_data_sample_gray.p"
(test_labels, test_data, test_tensors) = pickle.load(open(test_filename, mode='rb'))
# +
def onhotLabels(label):
from sklearn.preprocessing import OneHotEncoder
enc = OneHotEncoder()
enc.fit(label)
return enc.transform(label).toarray()
train_labels = onhotLabels(train_labels)
valid_labels = onhotLabels(valid_labels)
test_labels = onhotLabels(test_labels)
# -
# # CapsNet model
# +
import os
import argparse
from keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras import callbacks
import numpy as np
from keras import layers, models, optimizers
from keras import backend as K
import matplotlib.pyplot as plt
from PIL import Image
from capsulelayers import CapsuleLayer, PrimaryCap, Length, Mask
def CapsNet(input_shape, n_class, routings):
"""
A Capsule Network on MNIST.
:param input_shape: data shape, 3d, [width, height, channels]
:param n_class: number of classes
:param routings: number of routing iterations
:return: Two Keras Models, the first one used for training, and the second one for evaluation.
`eval_model` can also be used for training.
"""
x = layers.Input(shape=input_shape)
# Layer 1: Just a conventional Conv2D layer
conv1 = layers.Conv2D(filters=256, kernel_size=9, strides=1, padding='valid', activation='relu', name='conv1')(x)
# Layer 2: Conv2D layer with `squash` activation, then reshape to [None, num_capsule, dim_capsule]
primarycaps = PrimaryCap(conv1, dim_capsule=8, n_channels=32, kernel_size=9, strides=2, padding='valid')
# Layer 3: Capsule layer. Routing algorithm works here.
digitcaps = CapsuleLayer(num_capsule=n_class, dim_capsule=16, routings=routings,
name='digitcaps')(primarycaps)
# Layer 4: This is an auxiliary layer to replace each capsule with its length. Just to match the true label's shape.
# If using tensorflow, this will not be necessary. :)
out_caps = Length(name='capsnet')(digitcaps)
# Decoder network.
y = layers.Input(shape=(n_class,))
masked_by_y = Mask()([digitcaps, y]) # The true label is used to mask the output of capsule layer. For training
masked = Mask()(digitcaps) # Mask using the capsule with maximal length. For prediction
# Shared Decoder model in training and prediction
decoder = models.Sequential(name='decoder')
decoder.add(layers.Dense(512, activation='relu', input_dim=16*n_class))
decoder.add(layers.Dense(1024, activation='relu'))
decoder.add(layers.Dense(np.prod(input_shape), activation='sigmoid'))
decoder.add(layers.Reshape(target_shape=input_shape, name='out_recon'))
# Models for training and evaluation (prediction)
train_model = models.Model([x, y], [out_caps, decoder(masked_by_y)])
eval_model = models.Model(x, [out_caps, decoder(masked)])
# manipulate model
noise = layers.Input(shape=(n_class, 16))
noised_digitcaps = layers.Add()([digitcaps, noise])
masked_noised_y = Mask()([noised_digitcaps, y])
manipulate_model = models.Model([x, y, noise], decoder(masked_noised_y))
return train_model, eval_model, manipulate_model
model, eval_model, manipulate_model = CapsNet(input_shape=train_tensors.shape[1:],
n_class=len(np.unique(train_labels)),
routings=4)
decoder.summary()
model.summary()
# +
from keras import backend as K
def binary_accuracy(y_true, y_pred):
return K.mean(K.equal(y_true, K.round(y_pred)))
def precision_threshold(threshold = 0.5):
def precision(y_true, y_pred):
threshold_value = threshold
y_pred = K.cast(K.greater(K.clip(y_pred, 0, 1), threshold_value), K.floatx())
true_positives = K.round(K.sum(K.clip(y_true * y_pred, 0, 1)))
predicted_positives = K.sum(y_pred)
precision_ratio = true_positives / (predicted_positives + K.epsilon())
return precision_ratio
return precision
def recall_threshold(threshold = 0.5):
def recall(y_true, y_pred):
threshold_value = threshold
y_pred = K.cast(K.greater(K.clip(y_pred, 0, 1), threshold_value), K.floatx())
true_positives = K.round(K.sum(K.clip(y_true * y_pred, 0, 1)))
possible_positives = K.sum(K.clip(y_true, 0, 1))
recall_ratio = true_positives / (possible_positives + K.epsilon())
return recall_ratio
return recall
def fbeta_score_threshold(beta = 1, threshold = 0.5):
def fbeta_score(y_true, y_pred):
threshold_value = threshold
beta_value = beta
p = precision_threshold(threshold_value)(y_true, y_pred)
r = recall_threshold(threshold_value)(y_true, y_pred)
bb = beta_value ** 2
fbeta_score = (1 + bb) * (p * r) / (bb * p + r + K.epsilon())
return fbeta_score
return fbeta_score
# +
def margin_loss(y_true, y_pred):
"""
Margin loss for Eq.(4). When y_true[i, :] contains not just one `1`, this loss should work too. Not test it.
:param y_true: [None, n_classes]
:param y_pred: [None, num_capsule]
:return: a scalar loss value.
"""
L = y_true * K.square(K.maximum(0., 0.9 - y_pred)) + \
0.5 * (1 - y_true) * K.square(K.maximum(0., y_pred - 0.1))
return K.mean(K.sum(L, 1))
def train(model, data, lr, lr_decay, lam_recon, batch_size, shift_fraction, epochs):
"""
Training a CapsuleNet
:param model: the CapsuleNet model
:param data: a tuple containing training and testing data, like `((x_train, y_train), (x_test, y_test))`
:param args: arguments
:return: The trained model
"""
# unpacking the data
(x_train, y_train), (x_test, y_test) = data
# callbacks
log = callbacks.CSVLogger('saved_models/CapsNet_log.csv')
tb = callbacks.TensorBoard(log_dir='saved_models/tensorboard-logs',
batch_size=batch_size, histogram_freq=0)
checkpoint = callbacks.ModelCheckpoint(filepath='saved_models/CapsNet.best.from_scratch.hdf5',
verbose=1, save_best_only=True)
cb_lr_decay = callbacks.LearningRateScheduler(schedule=lambda epoch: lr * (lr_decay ** epoch))
# compile the model
model.compile(optimizer='sgd', loss='binary_crossentropy',
metrics=[precision_threshold(threshold = 0.5),
recall_threshold(threshold = 0.5),
fbeta_score_threshold(beta=0.5, threshold = 0.5),
'accuracy'])
# Training without data augmentation:
# model.fit([x_train, y_train], [y_train, x_train], batch_size=batch_size, epochs=epochs,
# validation_data=[[x_test, y_test], [y_test, x_test]], callbacks=[log, tb, checkpoint, cb_lr_decay])
# Begin: Training with data augmentation ---------------------------------------------------------------------#
def train_generator(x, y, batch_size, shift_fraction=0.):
train_datagen = ImageDataGenerator(width_shift_range=shift_fraction,
height_shift_range=shift_fraction) # shift up to 2 pixel for MNIST
generator = train_datagen.flow(x, y, batch_size=batch_size)
while 1:
x_batch, y_batch = generator.next()
yield ([x_batch, y_batch], [y_batch, x_batch])
# Training with data augmentation. If shift_fraction=0., also no augmentation.
model.fit_generator(generator=train_generator(x_train, y_train, batch_size, shift_fraction),
steps_per_epoch=int(y_train.shape[0] / batch_size),
epochs=epochs,
validation_data=[[x_test, y_test], [y_test, x_test]],
callbacks=[log, tb, checkpoint, cb_lr_decay])
# End: Training with data augmentation -----------------------------------------------------------------------#
from utils import plot_log
plot_log('saved_models/CapsNet_log.csv', show=True)
return model
# -
train(model=model, data=((train_tensors, train_labels), (valid_tensors, valid_labels)),
lr=0.001, lr_decay=0.9, lam_recon=0.392, batch_size=32, shift_fraction=0.1, epochs=20)
# # Testing
model.load_weights('saved_models/CapsNet.best.from_scratch.hdf5')
prediction = eval_model.predict(test_tensors)
# +
threshold = 0.5
beta = 0.5
pre = K.eval(precision_threshold(threshold = threshold)(K.variable(value=test_labels),
K.variable(value=prediction[0])))
rec = K.eval(recall_threshold(threshold = threshold)(K.variable(value=test_labels),
K.variable(value=prediction[0])))
fsc = K.eval(fbeta_score_threshold(beta = beta, threshold = threshold)(K.variable(value=test_labels),
K.variable(value=prediction[0])))
print ("Precision: %f %%\nRecall: %f %%\nFscore: %f %%"% (pre, rec, fsc))
# -
K.eval(binary_accuracy(K.variable(value=test_labels),
K.variable(value=prediction[0])))
prediction[:30]
|
Capsule Network basic - SampleDataset.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import tensorflow as tf
import pandas as pd
import numpy as np
from glob import glob
import matplotlib.pyplot as plt
# + _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a" _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0"
dataset_path = "/kaggle/input/signlang/ArASL_Database_54K_Final/ArASL_Database_54K_Final"
label_path = "/kaggle/input/signlang/ArSL_Data_Labels.csv"
# -
image = glob(dataset_path + "/*/*.*")
len(image)
# example of pixel normalization
from numpy import asarray
from PIL import Image
# load image
image = Image.open(image[0])
pixels = asarray(image)
# confirm pixel range is 0-255
print('Data Type: %s' % pixels.dtype)
print('Min: %.3f, Max: %.3f' % (pixels.min(), pixels.max()))
# convert from integers to floats
pixels = pixels.astype('float32')
# normalize to the range 0-1
pixels /= 255.0
# confirm the normalization
print('Min: %.3f, Max: %.3f' % (pixels.min(), pixels.max()))
df = pd.read_csv(label_path)
df.head()
classes = df.Class.unique().tolist()
classes
# +
from tensorflow.keras.preprocessing.image import ImageDataGenerator
datagen = ImageDataGenerator(rescale=1.0/255, validation_split=0.2)
training_generator = datagen.flow_from_directory(
dataset_path,
target_size=(64, 64),
batch_size=32,
color_mode="grayscale",
classes = classes,
subset='training')
validation_generator = datagen.flow_from_directory(
dataset_path, # same directory as training data
target_size=(64, 64),
batch_size=32,
color_mode="grayscale",
classes = classes,
subset='validation') # set as validation data
# +
from tensorflow.keras import models
from tensorflow.keras import layers
import tensorflow as tf
with tf.device('/device:GPU:0'):
# build a 6-layer
model = models.Sequential()
model.add(layers.Conv2D(32, (3, 3), activation='relu', input_shape=(64, 64, 1)))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Conv2D(64, (3, 3), activation='relu'))
model.add(layers.MaxPooling2D((2, 2)))
model.add(layers.Conv2D(64, (3, 3), activation='relu'))
model.add(layers.Flatten())
model.add(layers.Dense(64, activation='relu'))
model.add(layers.Dense(32, activation='softmax'))
model.summary()
# +
from tensorflow.keras.optimizers import Adam
early_stopping = tf.keras.callbacks.EarlyStopping(patience=10, verbose=1)
checkpointer = tf.keras.callbacks.ModelCheckpoint('asl_char.h5',verbose=1,save_best_only=True)
optimizer = Adam(lr=0.001)
model.compile(optimizer=optimizer, loss='categorical_crossentropy', metrics=['accuracy'])
history = model.fit(
data_generator,
steps_per_epoch = data_generator.samples // 32,
validation_data = validation_generator,
validation_steps = validation_generator.samples // 32,
epochs=50, callbacks=[early_stopping, checkpointer])
# -
converter = tf.lite.TFLiteConverter.from_keras_model(model)
converter.experimental_new_converter = True
tflite_model = converter.convert()
open("asl_char.tflite", "wb").write(tflite_model)
|
train_model/train_char_model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="rX8mhOLljYeM"
# ##### Copyright 2019 The TensorFlow Authors.
# + cellView="form" colab_type="code" id="BZSlp3DAjdYf" colab={}
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="3wF5wszaj97Y"
# # Lướt nhanh cơ bản TensorFlow 2.0
# + [markdown] colab_type="text" id="DUNzJc4jTj6G"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/tutorials/quickstart/beginner"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />Xem trên TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/vi/tutorials/quickstart/beginner.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Chạy trên Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/vi/tutorials/quickstart/beginner.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />Xem mã nguồn trên GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/vi/tutorials/quickstart/beginner.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Tải notebook</a>
# </td>
# </table>
# + [markdown] id="BbU6UaFdS-WR" colab_type="text"
# Note: Cộng đồng TensorFlow Việt Nam đã dịch các tài liệu này từ nguyên bản tiếng Anh.
# Vì bản dịch này dựa trên sự cố gắng từ các tình nguyện viên, nên không thể đám bảo luôn bám sát
# [Tài liệu chính thức bằng tiếng Anh](https://www.tensorflow.org/?hl=en).
# Nếu bạn có đề xuất để cải thiện bản dịch này, vui lòng tạo PR đến repository trên GitHub của [tensorflow/docs](https://github.com/tensorflow/docs)
#
# Để đăng ký dịch hoặc duyệt lại nội dung bản dịch, các bạn hãy liên hệ
# [<EMAIL>](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs).
# + [markdown] colab_type="text" id="hiH7AC-NTniF"
# Đây là một tệp notebook [Google Colaboratory](https://colab.research.google.com/notebooks/welcome.ipynb). Các chương trình Python sẽ chạy trực tiếp trong trình duyệt, giúp bạn dễ dàng tìm hiểu và sử dụng TensorFlow. Để làm theo giáo trình này, chạy notebook trên Google Colab bằng cách nhấp vào nút ở đầu trang.
#
# 1. Trong Colab, kết nối đến Python runtime: Ở phía trên cùng bên phải của thanh menu, chọn *CONNECT*.
# 2. Chạy tất cả các ô chứa mã trong notebook: Chọn *Runtime* > *Run all*.
# + [markdown] colab_type="text" id="nnrWf3PCEzXL"
# Tải và cài đặt TensorFlow 2.0 RC. Import TensorFlow vào chương trình:
# + colab_type="code" id="0trJmd6DjqBZ" colab={}
from __future__ import absolute_import, division, print_function, unicode_literals
# Install TensorFlow
try:
# # %tensorflow_version only exists in Colab.
# %tensorflow_version 2.x
except Exception:
pass
import tensorflow as tf
# + [markdown] colab_type="text" id="7NAbSZiaoJ4z"
# Load và chuẩn bị [tập dữ liệu MNIST](http://yann.lecun.com/exdb/mnist/). Chuyển kiểu dữ liệu của các mẫu từ số nguyên sang số thực dấu phẩy động:
# + colab_type="code" id="7FP5258xjs-v" colab={}
mnist = tf.keras.datasets.mnist
(x_train, y_train), (x_test, y_test) = mnist.load_data()
x_train, x_test = x_train / 255.0, x_test / 255.0
# + [markdown] colab_type="text" id="BPZ68wASog_I"
# Xây dựng mô hình `tf.keras.Sequential` bằng cách xếp chồng các layers. Chọn trình tối ưu hoá (optimizer) và hàm thiệt hại (loss) để huấn luyện:
# + colab_type="code" id="h3IKyzTCDNGo" colab={}
model = tf.keras.models.Sequential([
tf.keras.layers.Flatten(input_shape=(28, 28)),
tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(10, activation='softmax')
])
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
# + [markdown] colab_type="text" id="ix4mEL65on-w"
# Huấn luyện và đánh giá mô hình:
# + colab_type="code" id="F7dTAzgHDUh7" colab={}
model.fit(x_train, y_train, epochs=5)
model.evaluate(x_test, y_test, verbose=2)
# + [markdown] colab_type="text" id="T4JfEh7kvx6m"
# Mô hình phân loại ảnh này, sau khi được huấn luyện bằng tập dữ liệu trên, đạt độ chính xác (accuracy) ~98%. Để tìm hiểu thêm, bạn có thể đọc [Giáo trình TensorFlow](https://www.tensorflow.org/tutorials/).
|
site/vi/tutorials/quickstart/beginner.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import numpy as np
import pandas as pd
from pandas import Series,DataFrame
# +
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style('whitegrid')
# %matplotlib inline
# -
from sklearn.datasets import load_boston
boston = load_boston()
print boston.DESCR
# +
plt.hist(boston.target,bins=50)
plt.xlabel('Prices in $1000s')
plt.ylabel('Number of houses')
# +
plt.scatter(boston.data[:,5],boston.target)
plt.ylabel('Price in $1000s')
plt.xlabel('Number of rooms')
# +
boston_df = DataFrame(boston.data)
boston_df.columns = boston.feature_names
boston_df.head()
# +
boston_df['Price'] = boston.target
boston_df.head()
# -
sns.lmplot('RM', 'Price',data=boston_df)
|
Linear_Regrsn_Price_Predict.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import time
import numpy as np
from keras.models import Sequential
from keras.layers import Dense, Activation
from sklearn.model_selection import StratifiedKFold
def gen_rand(n_size=1):
'''
This function return a n_size-dimensional random vector.
'''
return np.random.random(n_size)
class NN_DE(object):
def __init__(self, n_pop=10, n_neurons=5, F=0.4, Cr=0.9, p=1, change_scheme=True ,scheme='rand',
bounds=[-1, 1], max_sp_evals=np.int(1e5), sp_tol=1e-2):
#self.n_gens=n_gens
self.n_pop=n_pop
self.n_neurons=n_neurons
self.F=F*np.ones(self.n_pop)
self.Cr=Cr*np.ones(self.n_pop)
self.bounds=bounds
self.p=p
self.scheme=scheme
self.change_schame=change_scheme
self.max_sp_evals=max_sp_evals
self.sp_tol = sp_tol
self.sp_evals=0
self.interactions=0
# Build generic model
model = Sequential()
model.add(Dense(self.n_neurons, input_dim=100, activation='tanh'))
model.add(Dense(1, activation='tanh'))
model.compile( loss='mean_squared_error', optimizer = 'rmsprop', metrics = ['accuracy'] )
self.model=model
self.change_schame=False
self.n_dim=model.count_params()
#self.population=NN_DE.init_population(self, pop_size=self.n_pop,
# dim=self.n_dim, bounds=self.bounds)
#self.train_dataset= train_dataset
#self.test_dataset= test_dataset
def init_population(self, pop_size, dim, bounds=[-1,1]):
'''
This function initialize the population to be use in DE
Arguments:
pop_size - Number of individuals (there is no default value to this yet.).
dim - dimension of the search space (default is 1).
bounds - The inferior and superior limits respectively (default is [-1, 1]).
'''
return np.random.uniform(low=bounds[0], high=bounds[1], size=(pop_size, dim))
def keep_bounds(self, pop, bounds, idx):
'''
This function keep the population in the seach space
Arguments:
pop - Population;
bounds - The inferior and superior limits respectively
'''
#up_ = np.where(pop>bounds[1])
#down_ = np.where(pop<bounds[1])
#best_ = pop[idx]
#print(pop[pop<bounds[0]])
#print(down_)
#print(best_.shape)
pop[pop<bounds[0]] = bounds[0]; pop[pop>bounds[1]] = bounds[1]
#pop[pop<bounds[0]] = 0.5*(bounds[0]+best_[down_]); pop[pop>bounds[1]] = 0.5*(bounds[1]+best_[up_])
return pop
# Define the Fitness to be used in DE
def sp_fitness(self, target, score):
'''
Calculate the SP index and return the index of the best SP found
Arguments:
target: True labels
score: the predicted labels
'''
from sklearn.metrics import roc_curve
fpr, tpr, thresholds = roc_curve(target, score)
jpr = 1. - fpr
sp = np.sqrt( (tpr + jpr)*.5 * np.sqrt(jpr*tpr) )
idx = np.argmax(sp)
return sp[idx], tpr[idx], fpr[idx]#sp, idx, sp[idx], tpr[idx], fpr[idx]
def convert_vector_weights(self, pop, nn_model):
model = nn_model
generic_weights = model.get_weights()
hl_lim = generic_weights[0].shape[0]*generic_weights[0].shape[1]
w = []
hl = pop[:hl_lim]
ol = pop[hl_lim+generic_weights[1].shape[0]:hl_lim+generic_weights[1].shape[0]+generic_weights[1].shape[0]]
w.append(hl.reshape(generic_weights[0].shape))
w.append(pop[hl_lim:hl_lim+generic_weights[1].shape[0]])
w.append(ol.reshape(generic_weights[2].shape))
w.append(np.array(pop[-1]).reshape(generic_weights[-1].shape))
return w
def set_weights_to_keras_model_and_compute_fitness(self,pop, data, nn_model):
'''
This function will create a generic model and set the weights to this model and compute the fitness.
Arguments:
pop - The population of weights.
data - The samples to be used to test.
'''
fitness = np.zeros((pop.shape[0],3))
#test_fitness = np.zeros((pop.shape[0],3))
model=nn_model
if pop.shape[0]!= self.n_pop:
#print('Local seach ind...')
w = NN_DE.convert_vector_weights(self, pop=pop, nn_model=model)
model.set_weights(w)
y_score = model.predict(data[0])
fitness = NN_DE.sp_fitness(self, target=data[1], score=y_score)
# Compute the SP for test in the same calling to minimeze the evals
#test_y_score = model.predict(test_data[0])
#test_fitness = NN_DE.sp_fitness(self, target=test_data[1], score=test_y_score)
return fitness#, test_fitness
for ind in range(pop.shape[0]):
w = NN_DE.convert_vector_weights(self, pop=pop[ind], nn_model=model)
model.set_weights(w)
y_score = model.predict(data[0])
fitness[ind] = NN_DE.sp_fitness(self, target=data[1], score=y_score)
# Compute the SP for test in the same calling to minimeze the evals
#test_y_score = model.predict(test_data[0])
#test_fitness[ind] = NN_DE.sp_fitness(self, target=test_data[1], score=test_y_score)
#print('Population ind: {} - SP: {} - PD: {} - PF: {}'.format(ind, fitness[ind][0], fitness[ind][1], fitness[ind][2]))
return fitness#, test_fitness
def evolution(self, dataset):
self.population=NN_DE.init_population(self, pop_size=self.n_pop,
dim=self.n_dim, bounds=self.bounds)
r_NNDE = {}
fitness = NN_DE.set_weights_to_keras_model_and_compute_fitness(self, pop=self.population,
data=dataset,
nn_model=self.model)
best_idx = np.argmax(fitness[:,0])
# Create the vectors F and Cr to be adapted during the interactions
NF = np.zeros_like(self.F)
NCr = np.zeros_like(self.Cr)
# Create a log
r_NNDE['log'] = []
r_NNDE['log'].append((self.sp_evals, fitness[best_idx], np.mean(fitness, axis=0),
np.std(fitness, axis=0), np.min(fitness, axis=0), np.median(fitness, axis=0), self.F, self.Cr))
#r_NNDE['test_log'] = []
#r_NNDE['test_log'].append((self.sp_evals, test_fitness[best_idx], np.mean(test_fitness, axis=0),
#np.std(test_fitness, axis=0), np.min(test_fitness, axis=0), np.median(test_fitness, axis=0), self.F, self.Cr))
while self.sp_evals < self.max_sp_evals:
# ============ Mutation Step ===============
mutant = np.zeros_like(self.population)
for ind in range(self.population.shape[0]):
if gen_rand() < 0.1:
NF[ind] = 0.2 +0.2*gen_rand()
else:
NF[ind] = self.F[ind]
tmp_pop = np.delete(self.population, ind, axis=0)
choices = np.random.choice(tmp_pop.shape[0], 1+2*self.p, replace=False)
diffs = 0
for idiff in range(1, len(choices), 2):
diffs += NF[ind]*((tmp_pop[choices[idiff]]-tmp_pop[choices[idiff+1]]))
if self.scheme=='rand':
mutant[ind] = tmp_pop[choices[0]] + diffs
elif self.scheme=='best':
mutant[ind] = self.population[best_idx] + diffs
# keep the bounds
mutant = NN_DE.keep_bounds(self, mutant, bounds=[-1,1], idx=best_idx)
# ============ Crossover Step =============
trial_pop = np.copy(self.population)
K = np.random.choice(trial_pop.shape[1])
for ind in range(trial_pop.shape[0]):
if gen_rand() < 0.1:
NCr[ind] = 0.8 +0.2*gen_rand()
else:
NCr[ind] = self.Cr[ind]
for jnd in range(trial_pop.shape[1]):
if jnd == K or gen_rand()<NCr[ind]:
trial_pop[ind][jnd] = mutant[ind][jnd]
# keep the bounds
trial_pop = NN_DE.keep_bounds(self, trial_pop, bounds=[-1,1], idx=best_idx)
trial_fitness = NN_DE.set_weights_to_keras_model_and_compute_fitness(self, pop=trial_pop,
data=dataset,
nn_model=self.model)
self.sp_evals += self.population.shape[0]
# ============ Selection Step ==============
winners = np.where(trial_fitness[:,0]>fitness[:,0])
# Auto-adtaptation of F and Cr like NSSDE
self.F[winners] = NF[winners]
self.Cr[winners] = NCr[winners]
# Greedy Selection
fitness[winners] = trial_fitness[winners]
self.population[winners] = trial_pop[winners]
best_idx = np.argmax(fitness[:,0])
if self.interactions > 0.95*self.max_sp_evals/self.n_pop:
print('=====Interaction: {}====='.format(self.interactions+1))
print('Best NN found - SP: {} / PD: {} / FA: {}'.format(fitness[best_idx][0],
fitness[best_idx][1],
fitness[best_idx][2]))
#print('Test > Mean - SP: {} +- {}'.format(np.mean(test_fitness,axis=0)[0],
# np.std(test_fitness,axis=0)[0]))
# Local search like NSSDE
a_1 = gen_rand(); a_2 = gen_rand()
a_3 = 1.0 - a_1 - a_2
k, r1, r2 = np.random.choice(self.population.shape[0], size=3)
V = np.zeros_like(self.population[k])
for jdim in range(self.population.shape[1]):
V[jdim] = a_1*self.population[k][jdim] + a_2*self.population[best_idx][jdim] + a_3*(self.population[r1][jdim] - self.population[r2][jdim])
V = NN_DE.keep_bounds(self, V, bounds=self.bounds, idx=best_idx)
V_train_fitness = NN_DE.set_weights_to_keras_model_and_compute_fitness(self, pop=V,
data=dataset,
nn_model=self.model)
self.sp_evals += 1
if V_train_fitness[0] > fitness[k][0]:
#print('Found best model using local search...')
self.population[k] = V
if V_train_fitness[0] > fitness[best_idx][0]:
best_idx = k
# ======== Done interaction ===========
self.interactions += 1
r_NNDE['log'].append((self.sp_evals, fitness[best_idx], np.mean(fitness, axis=0),
np.std(fitness, axis=0), np.min(fitness, axis=0), np.median(fitness, axis=0), self.F, self.Cr))
#r_NNDE['test_log'].append((self.sp_evals, test_fitness[best_idx], np.mean(test_fitness, axis=0),
# np.std(test_fitness, axis=0), np.min(test_fitness, axis=0), np.median(test_fitness, axis=0), self.F, self.Cr))
#print('Fitness: ', fitness[:,0])
#print('Mean: ',np.mean(fitness[:,0]))
if np.mean(fitness[:,0]) > .9 and np.abs(np.mean(fitness[:,0])-fitness[best_idx][0])< self.sp_tol:
print('Stop by Mean Criteria...')
break
# Compute the test
#test_fitness = NN_DE.set_weights_to_keras_model_and_compute_fitness(self, pop=self.population,
# data=self.test_dataset, nn_model=self.model)
r_NNDE['champion weights'] = NN_DE.convert_vector_weights(self, self.population[best_idx], self.model)
r_NNDE['model'] = self.model
r_NNDE['best index'] = best_idx
r_NNDE['Best NN'] = fitness[best_idx]
r_NNDE['fitness'] = fitness
#r_NNDE['test_fitness'] = test_fitness
r_NNDE['population'] = self.population,
return r_NNDE
# +
data = np.load('/home/micael/MyWorkspace/RingerRepresentation/2channels/data17-18_13TeV.sgn_lhmedium_probes.EGAM2.bkg.vetolhvloose.EGAM7.samples.npz')
sgn = data['signalPatterns_etBin_2_etaBin_0']
bkg = data['backgroundPatterns_etBin_2_etaBin_0']
# Equilibrate the classes to make a controled tests
bkg = bkg[np.random.choice(bkg.shape[0], size=sgn.shape[0]),:]
#print(sgn.shape, bkg.shape)
sgn_trgt = np.ones(sgn.shape[0])
bkg_trgt = -1*np.ones(bkg.shape[0])
sgn_normalized = np.zeros_like(sgn)
for ind in range(sgn.shape[0]):
sgn_normalized[ind] = sgn[ind]/np.abs(np.sum(sgn[ind]))
bkg_normalized = np.zeros_like(bkg)
for ind in range(bkg.shape[0]):
bkg_normalized[ind] = bkg[ind]/np.abs(np.sum(bkg[ind]))
data_ = np.append(sgn_normalized, bkg_normalized, axis=0)
trgt = np.append(sgn_trgt, bkg_trgt)
# -
n_runs = 10
# +
# %%time
result_dict = {}
nn_de = NN_DE(n_pop=20, max_sp_evals=2e3, scheme='best', sp_tol=1e-3)
for irun in range(n_runs):
init_run_time = time.time()
print('Begin Run {}'.format(irun+1))
result_dict['Run {}'.format(irun+1)] = nn_de.evolution(dataset=(data_, trgt))
end_run_time = time.time()
print('Run {} - Time: {}'.format(irun+1, end_run_time - init_run_time))
# -
return_dict.keys()
r = []
m = []
for ifold in return_dict.keys():
print(ifold, '>', return_dict[ifold]['fitness'][return_dict[ifold]['best index']])
r.append(return_dict[ifold]['fitness'][return_dict[ifold]['best index']])
m.append(np.mean(return_dict[ifold]['fitness'], axis=0))
#print('population: {}+-{}'.format(np.around(np.mean(return_dict[ifold]['fitness'], axis=0),7),
# np.around(np.std(return_dict[ifold]['fitness'], axis=0),7)))
print(np.around(100*np.mean(r, axis=0),7), np.around(100*np.std(r, axis=0),7))
print('Pop: ', np.around(100*np.mean(m, axis=0),7), np.around(100*np.std(m, axis=0),7))
r_ = {}
for ifold in return_dict.keys():
print(len(return_dict[ifold]['log']))
checks = list(range(0,len(return_dict[ifold]['log'])))
r_[ifold]={}
r_[ifold]['train'] = []
#r_[ifold]['test'] = []
for icheck in checks:
#print(ifold, '>', return_dict[ifold]['log'][icheck][2])
r_[ifold]['train'].append(return_dict[ifold]['log'][icheck][2])
#r_[ifold]['test'].append(return_dict[ifold]['test_log'][icheck][2])
merits = {
'SP' : 'SP Index',
'PD' : 'PD',
'FA' : 'FA'
}
# +
import matplotlib.pyplot as plt
plt.style.use('_classic_test')
folds = ['Run 1', 'Run 2', 'Run 3', 'Run 4', 'Run 5', 'Run 6', 'Run 7', 'Run 8', 'Run 9', 'Run 10']
#x_axis = np.array([0., 20, 100, 1000,2000,3000, 4000, 5000, 6000, 7000, 8000, 9000, 10000])
for idx, imerit in enumerate(merits.keys()):
print('Plot: ', imerit)
#f, (ax1, ax2) = plt.subplots(1, 2, sharey=False, figsize=(15,5))
for ifold in folds:
plt.plot(np.array(r_[ifold]['train'])[:,idx], label=ifold)
plt.legend(fontsize='large', loc='best')
plt.title(merits[imerit]+' - Train', fontsize=15)
plt.xlabel('Interactions', fontsize=10)
plt.ylabel('Mean '+merits[imerit], fontsize=10)
plt.grid(True)
#ax2.plot(np.array(r_[ifold]['test'])[:,idx], label=ifold)
#ax2.legend(fontsize='large', loc='best')
#ax2.set_title(merits[imerit]+' - Test', fontsize=15)
#ax2.set_xlabel('Interactions', fontsize=10)
#ax2.set_ylabel('Mean '+merits[imerit], fontsize=10)
#ax2.grid(True)
#plt.savefig(merits[imerit]+'.rand1bin.2000evals.withLS.MeanStopCriteria.pdf',)
#plt.savefig(merits[imerit]+'.rand1bin.2000evals.withLS.MeanStopCriteria.png', dpi=150)
plt.show()
# -
print(type(return_dict))
return_dict = dict(return_dict)
print(type(return_dict))
return_dict.keys()
return_dict['CVO'] = CVO
return_dict['CVO']
# +
import pickle
with open('nnde.5neurons.rand1bin.2000evals.withLS.MeanStopCriteria.pickle', 'wb') as handle:
pickle.dump(return_dict, handle, protocol=pickle.HIGHEST_PROTOCOL)
# -
# # Here begin the Backpropagation
#
# Steps:
#
# 1. Get the champions and set the model.
# 2. Fit the model in each fold.
# 3. Get the results.
number_of_epoch = 100
# %%time
for i in range(number_of_epoch):
pass
return_dict['Fold 1'].keys()
return_dict['Fold 1']['champion weights']
# +
import time
inicio = time.time()
nn_de = NN_DE(n_pop=20, max_sp_evals=1e4, scheme='rand')
resultado = {}
for ifold, (train_index, test_index) in enumerate(CVO):
print("TRAIN:", train_index, "TEST:", test_index, "Fold: ", ifold)
resultado['Fold {}'.format(ifold+1)] = nn_de.evolution(train_dataset=(data_[train_index], trgt[train_index]),
test_dataset=(data_[test_index], trgt[test_index]))
fim=time.time()
print('Demorou - {} segundos'.format(fim-inicio))
# -
resultado['Fold 1']
resultado = {}
for train_index, test_index in skf.split(data_, trgt):
print("TRAIN:", train_index, "TEST:", test_index)
teste = NN_DE(n_pop=20,max_sp_evals=2e3, scheme='rand', sp_tol=0.1)
ev = teste.evolution(train_dataset=(data_, trgt), test_dataset=(data_, trgt))
ev.keys()
ev['log'][-1]
np.mean(ev['fitness'],axis=0), np.std(ev['fitness'], axis=0)
np.argmin(ev['fitness'][:,0]), ev['best index']
ev['fitness'][ev['best index']], ev['fitness'][np.argmin(ev['fitness'][:,0])]
|
dev_notebooks/classNN-DE.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# # Example curve fitting
# ## Error of the curve
#
# This script also shows the error of the curve fit, the error shows how far the actual data and the curve fit data are apart from eachother. Their are 4 types of errors shown:
#
# ### Max error
# The maximum error shows the highest difference between the actual data and the curve fit data at a certain point in the graph.
#
# ### Minimum error
# The minimum error shows the lowest difference between the actual data and the curve fit data at a certain point in the graph.
#
# ### Total error
# The total error shows the sum of all the differences between the actual data and the curve fit data.
#
# ### Average error
# The average error shows the average difference between the actual data and the curve fit data through the entire graph.
#
# ### Root mean squared error
# This is a indication of how accurate the simulated data is compared to the actual data. This rmse is the most important stat for our curve fitting model
# ## Import libraries
from numpy import arange
from numpy import sin
import numpy as np
from pandas import read_csv
from scipy.optimize import curve_fit
from matplotlib import pyplot
import math
from curveFitAlgorithm import *
# ## Dataset we are working with in the examples
#
# This dataset contains information about population vs employed
# +
# link of the tutorial https://machinelearningmastery.com/curve-fitting-with-python/
# plot "Population" vs "Employed"
# load the dataset
url = 'https://raw.githubusercontent.com/jbrownlee/Datasets/master/longley.csv'
dataframe = read_csv(url, header=None)
data = dataframe.values
# choose the input and output variables
x, y = data[:, 4], data[:, -1]
# plot input vs output
pyplot.scatter(x, y)
pyplot.show()
# -
# ## Polynomial regression curve fitting
#
# In statistics, polynomial regression is a form of regression analysis in which the relationship between the independent variable x and the dependent variable y is modelled as an nth degree polynomial in x. Polynomial regression fits a nonlinear relationship between the value of x and the corresponding conditional mean of y, denoted E(y |x). Although polynomial regression fits a nonlinear model to the data, as a statistical estimation problem it is linear, in the sense that the regression function E(y | x) is linear in the unknown parameters that are estimated from the data. For this reason, polynomial regression is considered to be a special case of multiple linear regression.
# +
#polynomial regression curve fitting
# choose the input and output variables
x, y = data[:, 4], data[:, -1]
curve_fit_algorithm = PolynomialRegressionFit(x, y)
y_line = curve_fit_algorithm.get_y_line()
x_line = curve_fit_algorithm.get_x_line()
# plot input vs output
pyplot.scatter(x, y)
# create a line plot for the mapping function
pyplot.plot(x_line, y_line, '-', color='red')
pyplot.show()
print('rmse: ', curve_fit_algorithm.get_rmse())
print('total error: ', curve_fit_algorithm.get_total_error())
print('max error: ', curve_fit_algorithm.get_max_error())
print('min error: ', curve_fit_algorithm.get_min_error())
print('average error: ', curve_fit_algorithm.get_average_error())
# -
# ## Sine wave curve fitting
#
# The sine-fit algorithm is a fitting algorithm based on parameter estimation. Sine function signal model is sampled at equal intervals. The least squares method is used to fit the sampling sequence to determine the amplitude, frequency, phase and DC component of the sine-wave, so as to obtain a sine function expression
#
# +
#Sine wave curve fitting
# choose the input and output variables
x, y = data[:, 4], data[:, -1]
curve_fit_algorithm = SineWaveFit(x, y)
y_line = curve_fit_algorithm.get_y_line()
x_line = curve_fit_algorithm.get_x_line()
# plot input vs output
pyplot.scatter(x, y)
# create a line plot for the mapping function
pyplot.plot(x_line, y_line, '-', color='red')
pyplot.show()
print('rmse: ', curve_fit_algorithm.get_rmse())
print('total error: ', curve_fit_algorithm.get_total_error())
print('max error: ', curve_fit_algorithm.get_max_error())
print('min error: ', curve_fit_algorithm.get_min_error())
print('average error: ', curve_fit_algorithm.get_average_error())
# -
# ## non-linear least squares curve fitting
#
# Non-linear least squares is the form of least squares analysis used to fit a set of m observations with a model that is non-linear in n unknown parameters (m ≥ n). It is used in some forms of nonlinear regression. The basis of the method is to approximate the model by a linear one and to refine the parameters by successive iterations.
#
# +
#non-linear least squares curve fitting
# choose the input and output variables
x, y = data[:, 4], data[:, -1]
curve_fit_algorithm = NonLinearLeastSquaresFit(x, y)
y_line = curve_fit_algorithm.get_y_line()
x_line = curve_fit_algorithm.get_x_line()
# plot input vs output
pyplot.scatter(x, y)
# create a line plot for the mapping function
pyplot.plot(x_line, y_line, '-', color='red')
pyplot.show()
print('rmse: ', curve_fit_algorithm.get_rmse())
print('total error: ', curve_fit_algorithm.get_total_error())
print('max error: ', curve_fit_algorithm.get_max_error())
print('min error: ', curve_fit_algorithm.get_min_error())
print('average error: ', curve_fit_algorithm.get_average_error())
# -
# ## Fifth degree polynomial
#
# Fifth degree polynomials are also known as quintic polynomials. Quintics have these characteristics:
#
# * One to five roots.
# * Zero to four extrema.
# * One to three inflection points.
# * No general symmetry.
# * It takes six points or six pieces of information to describe a quintic function.
# +
#Fifth degree polynomial
# choose the input and output variables
x, y = data[:, 4], data[:, -1]
curve_fit_algorithm = FifthDegreePolynomialFit(x, y)
y_line = curve_fit_algorithm.get_y_line()
x_line = curve_fit_algorithm.get_x_line()
# plot input vs output
pyplot.scatter(x, y)
# create a line plot for the mapping function
pyplot.plot(x_line, y_line, '-', color='red')
pyplot.show()
print('rmse: ', curve_fit_algorithm.get_rmse())
print('total error: ', curve_fit_algorithm.get_total_error())
print('max error: ', curve_fit_algorithm.get_max_error())
print('min error: ', curve_fit_algorithm.get_min_error())
print('average error: ', curve_fit_algorithm.get_average_error())
# -
# ## Linear curve fitting
#
#
# +
#linear curve fitting
# choose the input and output variables
x, y = data[:, 4], data[:, -1]
# plot input vs output
pyplot.scatter(x, y)
# For details about the algoritm read the curveFitAlgoritm.py file or the technical documentation
curve_fit_algorithm = LinearFit(x, y)
y_line = curve_fit_algorithm.get_y_line()
x_line = curve_fit_algorithm.get_x_line()
# create a line plot for the mapping function
pyplot.plot(x_line, y_line, '-', color='red')
pyplot.show()
print('rmse: ', curve_fit_algorithm.get_rmse())
print('total error: ', curve_fit_algorithm.get_total_error())
print('max error: ', curve_fit_algorithm.get_max_error())
print('min error: ', curve_fit_algorithm.get_min_error())
print('average error: ', curve_fit_algorithm.get_average_error())
# -
|
docs/curve fitting/example.ipynb
|
# ##### Copyright 2020 The OR-Tools Authors.
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#
# # least_diff
# <table align="left">
# <td>
# <a href="https://colab.research.google.com/github/google/or-tools/blob/master/examples/notebook/contrib/least_diff.ipynb"><img src="https://raw.githubusercontent.com/google/or-tools/master/tools/colab_32px.png"/>Run in Google Colab</a>
# </td>
# <td>
# <a href="https://github.com/google/or-tools/blob/master/examples/contrib/least_diff.py"><img src="https://raw.githubusercontent.com/google/or-tools/master/tools/github_32px.png"/>View source on GitHub</a>
# </td>
# </table>
# First, you must install [ortools](https://pypi.org/project/ortools/) package in this colab.
# !pip install ortools
# +
# Copyright 2010 <NAME> <EMAIL>
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""
Least diff problem in Google CP Solver.
This model solves the following problem:
What is the smallest difference between two numbers X - Y
if you must use all the digits (0..9) exactly once.
Compare with the following models:
* Choco : http://www.hakank.org/choco/LeastDiff2.java
* ECLiPSE : http://www.hakank.org/eclipse/least_diff2.ecl
* Comet : http://www.hakank.org/comet/least_diff.co
* Tailor/Essence': http://www.hakank.org/tailor/leastDiff.eprime
* Gecode : http://www.hakank.org/gecode/least_diff.cpp
* Gecode/R: http://www.hakank.org/gecode_r/least_diff.rb
* JaCoP : http://www.hakank.org/JaCoP/LeastDiff.java
* MiniZinc: http://www.hakank.org/minizinc/least_diff.mzn
* SICStus : http://www.hakank.org/sicstus/least_diff.pl
* Zinc : http://hakank.org/minizinc/least_diff.zinc
This model was created by <NAME> (<EMAIL>)
Also see my other Google CP Solver models:
http://www.hakank.org/google_cp_solver/
"""
from __future__ import print_function
from ortools.constraint_solver import pywrapcp
# Create the solver.
solver = pywrapcp.Solver("Least diff")
#
# declare variables
#
digits = list(range(0, 10))
a = solver.IntVar(digits, "a")
b = solver.IntVar(digits, "b")
c = solver.IntVar(digits, "c")
d = solver.IntVar(digits, "d")
e = solver.IntVar(digits, "e")
f = solver.IntVar(digits, "f")
g = solver.IntVar(digits, "g")
h = solver.IntVar(digits, "h")
i = solver.IntVar(digits, "i")
j = solver.IntVar(digits, "j")
letters = [a, b, c, d, e, f, g, h, i, j]
digit_vector = [10000, 1000, 100, 10, 1]
x = solver.ScalProd(letters[0:5], digit_vector)
y = solver.ScalProd(letters[5:], digit_vector)
diff = x - y
#
# constraints
#
solver.Add(diff > 0)
solver.Add(solver.AllDifferent(letters))
# objective
objective = solver.Minimize(diff, 1)
#
# solution
#
solution = solver.Assignment()
solution.Add(letters)
solution.Add(x)
solution.Add(y)
solution.Add(diff)
# last solution since it's a minimization problem
collector = solver.LastSolutionCollector(solution)
search_log = solver.SearchLog(100, diff)
# Note: I'm not sure what CHOOSE_PATH do, but it is fast:
# find the solution in just 4 steps
solver.Solve(
solver.Phase(letters, solver.CHOOSE_PATH, solver.ASSIGN_MIN_VALUE),
[objective, search_log, collector])
# get the first (and only) solution
xval = collector.Value(0, x)
yval = collector.Value(0, y)
diffval = collector.Value(0, diff)
print("x:", xval)
print("y:", yval)
print("diff:", diffval)
print(xval, "-", yval, "=", diffval)
print([("abcdefghij" [i], collector.Value(0, letters[i])) for i in range(10)])
print()
print("failures:", solver.Failures())
print("branches:", solver.Branches())
print("WallTime:", solver.WallTime())
print()
|
examples/notebook/contrib/least_diff.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + active=""
# Explore indicator names to identify indicators that are not documented each year and indicators named differently in consecutive years, to support renaming or exclusion decisions.
# -
import pandas as pd
import os
import re
years=['2014','2015','2016','2017']
dirpath='//Users/rony/Dropbox/2_projects/Health_Geo/tables/'
#check indicator names
indicators=[]
filenames=[]
indicator_names={}
for year in years:
input_path=dirpath+'{year}/prevalence/cleaned/'.format(year=year)
filenames=os.listdir(path=input_path)
filenames=[y for y in filenames if bool(re.search('.csv',y))==True] #clean none .csv files
indicators=[re.findall('(.*).csv',x)[0] for x in filenames]
if '.ipynb_checkpoints' in filenames:
filenames.remove('.ipynb_checkpoints')
year_int=int(year)
indicator_names[year]=indicators
#double iteration over the indicator list dictionary to identify names that differ for the same indicator
for year, indicators in indicator_names.items():
#print (year,indicators)
if int(year)<2017:
year_1=str(int(year)+1)
for year, indicators in indicator_names.items():
indicators_1=indicator_names[year_1]
if year!=year_1:
set1=set (indicators)
set2=set (indicators_1)
if set1 - set2 !={}:
print ('indicator names difference , {year} and {year1}:'.format (year=year, year1=year_1), set1 - set2)
if set2 - set1 !={}:
print ('indicator names difference, {year1} and {year}:'.format (year=year, year1=year_1), set2 - set1)
print ('-----------------')
#double iteration over the indicator list dictionary to identify names that differ for the same indicator
for year, indicators in indicator_names.items():
#print (year,indicators)
if int(year)<2017:
year_1=str(int(year)+1)
indicators_1=indicator_names[year_1]
set1=set (indicators)
set2=set (indicators_1)
print ('indicator names difference , {year} and {year1}:'.format (year=year, year1=year_1), set1 - set2)
print ('indicator names difference, {year1} and {year}:'.format (year=year, year1=year_1), set2 - set1)
print ()
print ('indicators:', indicators)
print ()
print ('indicators_1:', indicators_1)
print ()
print ('-----------------')
# + active=""
# indicator names difference , 2014 and 2015: {'Learning_Disabilities_(18+)', 'Cardiovascular_Disease_-_Primary_Prevention', 'Hypothyroidism'}
# indicator names difference, 2015 and 2014: {'Learning_Disabilities', 'Heart_Failure_Due_To_Left_Ventrical_Systolic_Dysfunction', 'Cardiovascular_Disease_-_Primary_Prevention_(30-74)', 'Obesity', 'Learning_Disabilities_All_Ages'}
# + active=""
# Summary of manual check and decisions: indicators_names_differences.xsls
# -
decisions=pd.read_csv('indicators_names_differences.csv')
decisions
|
preparation/data/prevalence/indicator_names.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import gzip
import json
import pandas as pd
path_data = '../dataset/reviews_Cell_Phones_and_Accessories_5.json.gz'
path_data_prepaired = '../dataset/dataset.json'
# +
def parse(path):
g = gzip.open(path, 'rb')
for l in g:
yield eval(l)
def getDF(path):
i = 0
df = {}
for d in parse(path):
df[i] = d
i += 1
return pd.DataFrame.from_dict(df, orient='index')
# %time df = getDF(path_data)
# -
# %time df.to_json(path_data_prepaired, orient='records')
df.shape
# # References (links):
#
# - 1. https://www.kaggle.com/varun08/sentiment-analysis-using-word2vec
#
# - 2. http://www.aclweb.org/anthology/C14-1008
#
# - 3. datasets are here http://jmcauley.ucsd.edu/data/amazon/
#
|
notebooks/preprocessing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Convolutional Neural Networks
#
# ## Project: Write an Algorithm for a Dog Identification App
#
# ---
#
# In this notebook, some template code has already been provided for you, and you will need to implement additional functionality to successfully complete this project. You will not need to modify the included code beyond what is requested. Sections that begin with **'(IMPLEMENTATION)'** in the header indicate that the following block of code will require additional functionality which you must provide. Instructions will be provided for each section, and the specifics of the implementation are marked in the code block with a 'TODO' statement. Please be sure to read the instructions carefully!
#
# > **Note**: Once you have completed all of the code implementations, you need to finalize your work by exporting the Jupyter Notebook as an HTML document. Before exporting the notebook to html, all of the code cells need to have been run so that reviewers can see the final implementation and output. You can then export the notebook by using the menu above and navigating to **File -> Download as -> HTML (.html)**. Include the finished document along with this notebook as your submission.
#
# In addition to implementing code, there will be questions that you must answer which relate to the project and your implementation. Each section where you will answer a question is preceded by a **'Question X'** header. Carefully read each question and provide thorough answers in the following text boxes that begin with **'Answer:'**. Your project submission will be evaluated based on your answers to each of the questions and the implementation you provide.
#
# >**Note:** Code and Markdown cells can be executed using the **Shift + Enter** keyboard shortcut. Markdown cells can be edited by double-clicking the cell to enter edit mode.
#
# The rubric contains _optional_ "Stand Out Suggestions" for enhancing the project beyond the minimum requirements. If you decide to pursue the "Stand Out Suggestions", you should include the code in this Jupyter notebook.
#
#
#
# ---
# ### Why We're Here
#
# In this notebook, you will make the first steps towards developing an algorithm that could be used as part of a mobile or web app. At the end of this project, your code will accept any user-supplied image as input. If a dog is detected in the image, it will provide an estimate of the dog's breed. If a human is detected, it will provide an estimate of the dog breed that is most resembling. The image below displays potential sample output of your finished project (... but we expect that each student's algorithm will behave differently!).
#
# 
#
# In this real-world setting, you will need to piece together a series of models to perform different tasks; for instance, the algorithm that detects humans in an image will be different from the CNN that infers dog breed. There are many points of possible failure, and no perfect algorithm exists. Your imperfect solution will nonetheless create a fun user experience!
#
# ### The Road Ahead
#
# We break the notebook into separate steps. Feel free to use the links below to navigate the notebook.
#
# * [Step 0](#step0): Import Datasets
# * [Step 1](#step1): Detect Humans
# * [Step 2](#step2): Detect Dogs
# * [Step 3](#step3): Create a CNN to Classify Dog Breeds (from Scratch)
# * [Step 4](#step4): Create a CNN to Classify Dog Breeds (using Transfer Learning)
# * [Step 5](#step5): Write your Algorithm
# * [Step 6](#step6): Test Your Algorithm
#
# ---
# <a id='step0'></a>
# ## Step 0: Import Datasets
#
# Make sure that you've downloaded the required human and dog datasets:
#
# **Note: if you are using the Udacity workspace, you *DO NOT* need to re-download these - they can be found in the `/data` folder as noted in the cell below.**
#
# * Download the [dog dataset](https://s3-us-west-1.amazonaws.com/udacity-aind/dog-project/dogImages.zip). Unzip the folder and place it in this project's home directory, at the location `/dog_images`.
#
# * Download the [human dataset](https://s3-us-west-1.amazonaws.com/udacity-aind/dog-project/lfw.zip). Unzip the folder and place it in the home directory, at location `/lfw`.
#
# *Note: If you are using a Windows machine, you are encouraged to use [7zip](http://www.7-zip.org/) to extract the folder.*
#
# In the code cell below, we save the file paths for both the human (LFW) dataset and dog dataset in the numpy arrays `human_files` and `dog_files`.
# +
import numpy as np
from glob import glob
# load filenames for human and dog images
human_files = np.array(glob("/data/lfw/*/*"))
dog_files = np.array(glob("/data/dog_images/*/*/*"))
# print number of images in each dataset
print('There are %d total human images.' % len(human_files))
print('There are %d total dog images.' % len(dog_files))
# -
# # Data Exploration and Exploratory Visualization
print(human_files)
print('The first element in dog_files: {}'.format(dog_files[0]))
print('There are {} human pictures.'.format(len(human_files)))
print('There are {} dog pictures.'.format(len(dog_files)))
import matplotlib.pyplot as plt
print('The first human file\'s shape is: {}'.format(plt.imread(human_files[0]).shape))
print('The second human file\'s shape is: {}'.format(plt.imread(human_files[1]).shape))
print('The third human file\'s shape is: {}'.format(plt.imread(human_files[2]).shape))
print('The first dig file\'s shape is: {}'.format(plt.imread(dog_files[0]).shape))
print('The second dog file\'s shape is: {}'.format(plt.imread(dog_files[1]).shape))
print('The third dog file\'s shape is: {}'.format(plt.imread(dog_files[2]).shape))
plt.imshow(plt.imread(human_files[0]))
plt.imshow(plt.imread(human_files[1]))
from tqdm import tqdm
human_shape_list = [(plt.imread(file).shape) for file in tqdm(human_files)]
dog_shape_list = [(plt.imread(file).shape) for file in tqdm(dog_files)]
import pandas as pd
human_shape_df, dog_shape_df = pd.DataFrame(), pd.DataFrame()
human_shape_df['human_shape'] = human_shape_list
dog_shape_df['dog_shape'] = dog_shape_list
display(human_shape_df.describe())
display(dog_shape_df.describe())
# <a id='step1'></a>
# ## Step 1: Detect Humans
#
# In this section, we use OpenCV's implementation of [Haar feature-based cascade classifiers](http://docs.opencv.org/trunk/d7/d8b/tutorial_py_face_detection.html) to detect human faces in images.
#
# OpenCV provides many pre-trained face detectors, stored as XML files on [github](https://github.com/opencv/opencv/tree/master/data/haarcascades). We have downloaded one of these detectors and stored it in the `haarcascades` directory. In the next code cell, we demonstrate how to use this detector to find human faces in a sample image.
# +
import cv2
import matplotlib.pyplot as plt
# %matplotlib inline
# extract pre-trained face detector
face_cascade = cv2.CascadeClassifier('haarcascades/haarcascade_frontalface_alt.xml')
# load color (BGR) image
img = cv2.imread(human_files[0])
# convert BGR image to grayscale
gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
# find faces in image
faces = face_cascade.detectMultiScale(gray)
# print number of faces detected in the image
print('Number of faces detected:', len(faces))
# get bounding box for each detected face
for (x,y,w,h) in faces:
# add bounding box to color image
cv2.rectangle(img,(x,y),(x+w,y+h),(255,0,0),2)
# convert BGR image to RGB for plotting
cv_rgb = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
# display the image, along with bounding box
plt.imshow(cv_rgb)
plt.show()
# -
# Before using any of the face detectors, it is standard procedure to convert the images to grayscale. The `detectMultiScale` function executes the classifier stored in `face_cascade` and takes the grayscale image as a parameter.
#
# In the above code, `faces` is a numpy array of detected faces, where each row corresponds to a detected face. Each detected face is a 1D array with four entries that specifies the bounding box of the detected face. The first two entries in the array (extracted in the above code as `x` and `y`) specify the horizontal and vertical positions of the top left corner of the bounding box. The last two entries in the array (extracted here as `w` and `h`) specify the width and height of the box.
#
# ### Write a Human Face Detector
#
# We can use this procedure to write a function that returns `True` if a human face is detected in an image and `False` otherwise. This function, aptly named `face_detector`, takes a string-valued file path to an image as input and appears in the code block below.
# returns "True" if face is detected in image stored at img_path
def face_detector(img_path):
img = cv2.imread(img_path)
gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
faces = face_cascade.detectMultiScale(gray)
return len(faces) > 0
# ### (IMPLEMENTATION) Assess the Human Face Detector
#
# __Question 1:__ Use the code cell below to test the performance of the `face_detector` function.
# - What percentage of the first 100 images in `human_files` have a detected human face?
# - What percentage of the first 100 images in `dog_files` have a detected human face?
#
# Ideally, we would like 100% of human images with a detected face and 0% of dog images with a detected face. You will see that our algorithm falls short of this goal, but still gives acceptable performance. We extract the file paths for the first 100 images from each of the datasets and store them in the numpy arrays `human_files_short` and `dog_files_short`.
# __Answer:__
# (You can print out your results and/or write your percentages in this cell)
# __98% human pictures are recognized as human face.__
# __17% dog pictures are recognized as human face.__
# +
from tqdm import tqdm
human_files_short = human_files[:100]
dog_files_short = dog_files[:100]
#-#-# Do NOT modify the code above this line. #-#-#
## TODO: Test the performance of the face_detector algorithm
## on the images in human_files_short and dog_files_short.
human = 0
dog = 0
for human_file in tqdm(human_files_short):
human += 1 * face_detector(human_file)
for dog_files in tqdm(dog_files_short):
dog += 1 * face_detector(dog_files)
print('{}% human pictures are recognized as human face.'.format(human))
print('{}% dog pictures are recognized as human face.'.format(dog))
# -
# We suggest the face detector from OpenCV as a potential way to detect human images in your algorithm, but you are free to explore other approaches, especially approaches that make use of deep learning :). Please use the code cell below to design and test your own face detection algorithm. If you decide to pursue this _optional_ task, report performance on `human_files_short` and `dog_files_short`.
# +
### (Optional)
### TODO: Test performance of anotherface detection algorithm.
### Feel free to use as many code cells as needed.
# -
# ---
# <a id='step2'></a>
# ## Step 2: Detect Dogs
#
# In this section, we use a [pre-trained model](http://pytorch.org/docs/master/torchvision/models.html) to detect dogs in images.
#
# ### Obtain Pre-trained VGG-16 Model
#
# The code cell below downloads the VGG-16 model, along with weights that have been trained on [ImageNet](http://www.image-net.org/), a very large, very popular dataset used for image classification and other vision tasks. ImageNet contains over 10 million URLs, each linking to an image containing an object from one of [1000 categories](https://gist.github.com/yrevar/942d3a0ac09ec9e5eb3a).
# +
import torch
import torchvision.models as models
# define VGG16 model
VGG16 = models.vgg16(pretrained=True)
# check if CUDA is available
use_cuda = torch.cuda.is_available()
# move model to GPU if CUDA is available
if use_cuda:
VGG16 = VGG16.cuda()
# -
# Given an image, this pre-trained VGG-16 model returns a prediction (derived from the 1000 possible categories in ImageNet) for the object that is contained in the image.
# ### (IMPLEMENTATION) Making Predictions with a Pre-trained Model
#
# In the next code cell, you will write a function that accepts a path to an image (such as `'dogImages/train/001.Affenpinscher/Affenpinscher_00001.jpg'`) as input and returns the index corresponding to the ImageNet class that is predicted by the pre-trained VGG-16 model. The output should always be an integer between 0 and 999, inclusive.
#
# Before writing the function, make sure that you take the time to learn how to appropriately pre-process tensors for pre-trained models in the [PyTorch documentation](http://pytorch.org/docs/stable/torchvision/models.html).
# +
from PIL import Image
import torchvision.transforms as transforms
def VGG16_predict(img_path):
'''
Use pre-trained VGG-16 model to obtain index corresponding to
predicted ImageNet class for image at specified path
Args:
img_path: path to an image
Returns:
Index corresponding to VGG-16 model's prediction
'''
## TODO: Complete the function.
## Load and pre-process an image from the given img_path
## Return the *index* of the predicted class for that image
transform = transforms.Compose([transforms.Resize(256),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize(mean=[0.485, 0.456, 0.406], std=[0.229, 0.224, 0.225])])
img = Image.open(img_path)
img = transform(img)
img = torch.unsqueeze(img, 0)
if torch.cuda.is_available():
img = img.cuda()
prediction = VGG16(img)
if torch.cuda.is_available():
prediction = prediction.cpu()
index = prediction.data.numpy().argmax()
return index # predicted class index
# -
# ### (IMPLEMENTATION) Write a Dog Detector
#
# While looking at the [dictionary](https://gist.github.com/yrevar/942d3a0ac09ec9e5eb3a), you will notice that the categories corresponding to dogs appear in an uninterrupted sequence and correspond to dictionary keys 151-268, inclusive, to include all categories from `'Chihuahua'` to `'Mexican hairless'`. Thus, in order to check to see if an image is predicted to contain a dog by the pre-trained VGG-16 model, we need only check if the pre-trained model predicts an index between 151 and 268 (inclusive).
#
# Use these ideas to complete the `dog_detector` function below, which returns `True` if a dog is detected in an image (and `False` if not).
### returns "True" if a dog is detected in the image stored at img_path
def dog_detector(img_path):
## TODO: Complete the function.
label = VGG16_predict(img_path)
logic = label >= 151 and label <= 268
return logic# true/false
# ### (IMPLEMENTATION) Assess the Dog Detector
#
# __Question 2:__ Use the code cell below to test the performance of your `dog_detector` function.
# - What percentage of the images in `human_files_short` have a detected dog?
# - What percentage of the images in `dog_files_short` have a detected dog?
# __Answer:__
# __0% and 100%__
# +
### TODO: Test the performance of the dog_detector function
### on the images in human_files_short and dog_files_short.
human = 0
dog = 0
for human_file in tqdm(human_files_short):
human += 1 * dog_detector(human_file)
for dog_files in tqdm(dog_files_short):
dog += 1 * dog_detector(dog_files)
print('{}% human pictures are recognized as dog.'.format(human))
print('{}% dog pictures are recognized as dog.'.format(dog))
# -
# We suggest VGG-16 as a potential network to detect dog images in your algorithm, but you are free to explore other pre-trained networks (such as [Inception-v3](http://pytorch.org/docs/master/torchvision/models.html#inception-v3), [ResNet-50](http://pytorch.org/docs/master/torchvision/models.html#id3), etc). Please use the code cell below to test other pre-trained PyTorch models. If you decide to pursue this _optional_ task, report performance on `human_files_short` and `dog_files_short`.
# +
### (Optional)
### TODO: Report the performance of another pre-trained network.
### Feel free to use as many code cells as needed.
# -
# ---
# <a id='step3'></a>
# ## Step 3: Create a CNN to Classify Dog Breeds (from Scratch)
#
# Now that we have functions for detecting humans and dogs in images, we need a way to predict breed from images. In this step, you will create a CNN that classifies dog breeds. You must create your CNN _from scratch_ (so, you can't use transfer learning _yet_!), and you must attain a test accuracy of at least 10%. In Step 4 of this notebook, you will have the opportunity to use transfer learning to create a CNN that attains greatly improved accuracy.
#
# We mention that the task of assigning breed to dogs from images is considered exceptionally challenging. To see why, consider that *even a human* would have trouble distinguishing between a Brittany and a Welsh Springer Spaniel.
#
# Brittany | Welsh Springer Spaniel
# - | -
# <img src="images/Brittany_02625.jpg" width="100"> | <img src="images/Welsh_springer_spaniel_08203.jpg" width="200">
#
# It is not difficult to find other dog breed pairs with minimal inter-class variation (for instance, Curly-Coated Retrievers and American Water Spaniels).
#
# Curly-Coated Retriever | American Water Spaniel
# - | -
# <img src="images/Curly-coated_retriever_03896.jpg" width="200"> | <img src="images/American_water_spaniel_00648.jpg" width="200">
#
#
# Likewise, recall that labradors come in yellow, chocolate, and black. Your vision-based algorithm will have to conquer this high intra-class variation to determine how to classify all of these different shades as the same breed.
#
# Yellow Labrador | Chocolate Labrador | Black Labrador
# - | -
# <img src="images/Labrador_retriever_06457.jpg" width="150"> | <img src="images/Labrador_retriever_06455.jpg" width="240"> | <img src="images/Labrador_retriever_06449.jpg" width="220">
#
# We also mention that random chance presents an exceptionally low bar: setting aside the fact that the classes are slightly imabalanced, a random guess will provide a correct answer roughly 1 in 133 times, which corresponds to an accuracy of less than 1%.
#
# Remember that the practice is far ahead of the theory in deep learning. Experiment with many different architectures, and trust your intuition. And, of course, have fun!
#
# ### (IMPLEMENTATION) Specify Data Loaders for the Dog Dataset
#
# Use the code cell below to write three separate [data loaders](http://pytorch.org/docs/stable/data.html#torch.utils.data.DataLoader) for the training, validation, and test datasets of dog images (located at `dog_images/train`, `dog_images/valid`, and `dog_images/test`, respectively). You may find [this documentation on custom datasets](http://pytorch.org/docs/stable/torchvision/datasets.html) to be a useful resource. If you are interested in augmenting your training and/or validation data, check out the wide variety of [transforms](http://pytorch.org/docs/stable/torchvision/transforms.html?highlight=transform)!
# +
import os
from torchvision import datasets
import torchvision
### TODO: Write data loaders for training, validation, and test sets
## Specify appropriate transforms, and batch_sizes
Dir = '/data/dog_images'
train_path = os.path.join(Dir, 'train')
val_path = os.path.join(Dir, 'valid')
test_path = os.path.join(Dir, 'test')
batch = 10
transform = transforms.Compose([transforms.Resize(224),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize(mean=[0.485, 0.456, 0.406], std=[0.229, 0.224, 0.225])])
train_data = torchvision.datasets.ImageFolder(root=train_path,transform=transform)
train_loader = torch.utils.data.DataLoader(train_data,
batch_size=batch,
shuffle=True,
)
val_data = torchvision.datasets.ImageFolder(root=val_path,transform=transform)
val_loader = torch.utils.data.DataLoader(val_data,
batch_size=batch,
shuffle=True,
)
test_data = torchvision.datasets.ImageFolder(root=test_path,transform=transform)
test_loader = torch.utils.data.DataLoader(test_data,
batch_size=batch,
shuffle=True,
)
loaders_scratch={'train':train_loader,'valid':val_loader,'test':test_loader}
# -
# **Question 3:** Describe your chosen procedure for preprocessing the data.
# - How does your code resize the images (by cropping, stretching, etc)? What size did you pick for the input tensor, and why?
# - Did you decide to augment the dataset? If so, how (through translations, flips, rotations, etc)? If not, why not?
#
# **Answer**:
# - I just resize the picture to 224 * 224 as in VGG model.
# - I do not augment the dataset because the plain preprocess reachs the required 10% accuracy.
# ### (IMPLEMENTATION) Model Architecture
#
# Create a CNN to classify dog breed. Use the template in the code cell below.
# +
import torch.nn as nn
import torch.nn.functional as F
# define the CNN architecture
class Net(nn.Module):
### TODO: choose an architecture, and complete the class
## Define layers of a CNN
def __init__(self):
super(Net, self).__init__()
self.conv1 = nn.Conv2d(3, 16, 3, padding=1)
self.batchnorm1 = nn.BatchNorm2d(16)
self.conv2 = nn.Conv2d(16, 32, 3, padding=1)
self.batchnorm2 = nn.BatchNorm2d(32)
self.conv3 = nn.Conv2d(32, 64, 3, padding=1)
self.batchnorm3 = nn.BatchNorm2d(64)
self.maxpool = nn.MaxPool2d(2, 2)
self.fc1 = nn.Linear(28 * 28 * 64, 512)
self.fc2 = nn.Linear(512, 133)
self.dropout = nn.Dropout(0.5)
def forward(self, x):
x = self.maxpool(F.relu(self.conv1(x)))
x = self.batchnorm1(x)
x = self.maxpool(F.relu(self.conv2(x)))
x = self.batchnorm2(x)
x = self.maxpool(F.relu(self.conv3(x)))
x = self.batchnorm3(x)
x = x.view(-1, 28 * 28 * 64)
x = self.dropout(x)
x = F.relu(self.fc1(x))
x = self.dropout(x)
x = self.fc2(x)
return x
#-#-# You so NOT have to modify the code below this line. #-#-#
# instantiate the CNN
model_scratch = Net()
# move tensors to GPU if CUDA is available
if use_cuda:
model_scratch.cuda()
# -
# __Question 4:__ Outline the steps you took to get to your final CNN architecture and your reasoning at each step.
# __Answer:__ It's natural to come up with a three Convolutional payer with two folly connected layer. Some tricks preventing overfitting are also involved. It's just the most naiive model.
# ### (IMPLEMENTATION) Specify Loss Function and Optimizer
#
# Use the next code cell to specify a [loss function](http://pytorch.org/docs/stable/nn.html#loss-functions) and [optimizer](http://pytorch.org/docs/stable/optim.html). Save the chosen loss function as `criterion_scratch`, and the optimizer as `optimizer_scratch` below.
# +
import torch.optim as optim
### TODO: select loss function
criterion_scratch = nn.CrossEntropyLoss()
### TODO: select optimizer
optimizer_scratch = optim.Adam(params = model_scratch.parameters(), lr = 0.01)
# -
# ### (IMPLEMENTATION) Train and Validate the Model
#
# Train and validate your model in the code cell below. [Save the final model parameters](http://pytorch.org/docs/master/notes/serialization.html) at filepath `'model_scratch.pt'`.
# +
from PIL import ImageFile
ImageFile.LOAD_TRUNCATED_IMAGES = True
def train(n_epochs, loaders, model, optimizer, criterion, use_cuda, save_path):
"""returns trained model"""
# initialize tracker for minimum validation loss
valid_loss_min = np.Inf
for epoch in tqdm(range(1, n_epochs+1)):
# initialize variables to monitor training and validation loss
train_loss = 0.0
valid_loss = 0.0
###################
# train the model #
###################
model.train()
for batch_idx, (data, target) in enumerate(loaders['train']):
# move to GPU
if use_cuda:
data, target = data.cuda(), target.cuda()
optimizer.zero_grad()
outputs=model(data)
loss=criterion(outputs,target)
loss.backward()
optimizer.step()
## record the average training loss, using something like
train_loss = train_loss + ((1 / (batch_idx + 1)) * (loss.data - train_loss))
######################
# validate the model #
######################
model.eval()
for batch_idx, (data, target) in enumerate(loaders['valid']):
# move to GPU
if use_cuda:
data, target = data.cuda(), target.cuda()
outputs = model(data)
loss = criterion(outputs, target)
valid_loss = valid_loss + ((1 / (batch_idx + 1)) * (loss.data - valid_loss))
# print training/validation statistics
print('Epoch: {} \tTraining Loss: {:.6f} \tValidation Loss: {:.6f}'.format(
epoch,
train_loss,
valid_loss
))
## TODO: save the model if validation loss has decreased
if valid_loss<valid_loss_min:
print('Validation loss descreased from {} to {}. save the model'.format(valid_loss_min,valid_loss))
torch.save(model.state_dict(), save_path) # save the model
valid_loss_min=valid_loss # update decreased validation loss
# return trained model
return model
# train the model
model_scratch = train(10, loaders_scratch, model_scratch, optimizer_scratch,
criterion_scratch, use_cuda, 'model_scratch.pt')
# load the model that got the best validation accuracy
model_scratch.load_state_dict(torch.load('model_scratch.pt'))
# -
# ### (IMPLEMENTATION) Test the Model
#
# Try out your model on the test dataset of dog images. Use the code cell below to calculate and print the test loss and accuracy. Ensure that your test accuracy is greater than 10%.
# +
def test(loaders, model, criterion, use_cuda):
# monitor test loss and accuracy
test_loss = 0.
correct = 0.
total = 0.
model.eval()
for batch_idx, (data, target) in enumerate(loaders['test']):
# move to GPU
if use_cuda:
data, target = data.cuda(), target.cuda()
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the loss
loss = criterion(output, target)
# update average test loss
test_loss = test_loss + ((1 / (batch_idx + 1)) * (loss.data - test_loss))
# convert output probabilities to predicted class
pred = output.data.max(1, keepdim=True)[1]
# compare predictions to true label
correct += np.sum(np.squeeze(pred.eq(target.data.view_as(pred))).cpu().numpy())
total += data.size(0)
print('Test Loss: {:.6f}\n'.format(test_loss))
print('\nTest Accuracy: %2d%% (%2d/%2d)' % (
100. * correct / total, correct, total))
# call test function
test(loaders_scratch, model_scratch, criterion_scratch, use_cuda)
# -
# ---
# <a id='step4'></a>
# ## Step 4: Create a CNN to Classify Dog Breeds (using Transfer Learning)
#
# You will now use transfer learning to create a CNN that can identify dog breed from images. Your CNN must attain at least 60% accuracy on the test set.
#
# ### (IMPLEMENTATION) Specify Data Loaders for the Dog Dataset
#
# Use the code cell below to write three separate [data loaders](http://pytorch.org/docs/master/data.html#torch.utils.data.DataLoader) for the training, validation, and test datasets of dog images (located at `dogImages/train`, `dogImages/valid`, and `dogImages/test`, respectively).
#
# If you like, **you are welcome to use the same data loaders from the previous step**, when you created a CNN from scratch.
## TODO: Specify data loaders
loaders_transfer={'train':train_loader,'valid':val_loader,'test':test_loader}
# ### (IMPLEMENTATION) Model Architecture
#
# Use transfer learning to create a CNN to classify dog breed. Use the code cell below, and save your initialized model as the variable `model_transfer`.
# +
import torchvision.models as models
import torch.nn as nn
## TODO: Specify model architecture
model_transfer = models.resnet152(pretrained=True)
for parameter in model_transfer.parameters():
parameter.requires_grad = False
num_in=model_transfer.fc.in_features
model_transfer.fc=nn.Linear(in_features=num_in,out_features=133)
if use_cuda:
model_transfer = model_transfer.cuda()
# -
# __Question 5:__ Outline the steps you took to get to your final CNN architecture and your reasoning at each step. Describe why you think the architecture is suitable for the current problem.
# __Answer:__ I think any pre-trained huge CNN would satisfy this dog project. So all I need to do is download one and put it into work.
#
# ### (IMPLEMENTATION) Specify Loss Function and Optimizer
#
# Use the next code cell to specify a [loss function](http://pytorch.org/docs/master/nn.html#loss-functions) and [optimizer](http://pytorch.org/docs/master/optim.html). Save the chosen loss function as `criterion_transfer`, and the optimizer as `optimizer_transfer` below.
criterion_transfer = nn.CrossEntropyLoss()
optimizer_transfer = optim.SGD(params=filter(lambda p: p.requires_grad, model_transfer.parameters()), lr=0.001)
# ### (IMPLEMENTATION) Train and Validate the Model
#
# Train and validate your model in the code cell below. [Save the final model parameters](http://pytorch.org/docs/master/notes/serialization.html) at filepath `'model_transfer.pt'`.
# +
# train the model
n_epochs = 10
model_transfer = train(n_epochs, loaders_transfer, model_transfer, optimizer_transfer, criterion_transfer, use_cuda, 'model_transfer.pt')
# load the model that got the best validation accuracy (uncomment the line below)
model_transfer.load_state_dict(torch.load('model_transfer.pt'))
# -
# ### (IMPLEMENTATION) Test the Model
#
# Try out your model on the test dataset of dog images. Use the code cell below to calculate and print the test loss and accuracy. Ensure that your test accuracy is greater than 60%.
test(loaders_transfer, model_transfer, criterion_transfer, use_cuda)
# ### (IMPLEMENTATION) Predict Dog Breed with the Model
#
# Write a function that takes an image path as input and returns the dog breed (`Affenpinscher`, `Afghan hound`, etc) that is predicted by your model.
# +
### TODO: Write a function that takes a path to an image as input
### and returns the dog breed that is predicted by the model.
# list of class names by index, i.e. a name can be accessed like class_names[0]
# class_names = [item[4:].replace("_", " ") for item in data_transfer['train'].classes]
class_names = [item[4:].replace("_", " ") for item in train_data.classes]
def predict_breed_transfer(img_path):
# load the image and return the predicted breed
transform = transforms.Compose([transforms.Resize(256),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize(mean=[0.485, 0.456, 0.406], std=[0.229, 0.224, 0.225])])
img = Image.open(img_path)
img = transform(img)
img = torch.unsqueeze(img, 0)
img = img.cuda()
output=model_transfer(img)
_, index = torch.max(output, 1)
return class_names[index]
# -
# ---
# <a id='step5'></a>
# ## Step 5: Write your Algorithm
#
# Write an algorithm that accepts a file path to an image and first determines whether the image contains a human, dog, or neither. Then,
# - if a __dog__ is detected in the image, return the predicted breed.
# - if a __human__ is detected in the image, return the resembling dog breed.
# - if __neither__ is detected in the image, provide output that indicates an error.
#
# You are welcome to write your own functions for detecting humans and dogs in images, but feel free to use the `face_detector` and `human_detector` functions developed above. You are __required__ to use your CNN from Step 4 to predict dog breed.
#
# Some sample output for our algorithm is provided below, but feel free to design your own user experience!
#
# 
#
#
# ### (IMPLEMENTATION) Write your Algorithm
# +
### TODO: Write your algorithm.
### Feel free to use as many code cells as needed.
def run_app(img_path):
## handle cases for a human face, dog, and neither
if face_detector(img_path):
print('Hello, human! ')
plt.imshow(plt.imread(img_path)); # display the image
plt.show()
print('You look like a...')
print(predict_breed_transfer(img_path))
elif dog_detector(img_path):
print('Hello, dog! ')
plt.imshow(plt.imread(img_path)); # display the image
plt.show()
print('Your predicted breed is...')
print(predict_breed_transfer(img_path))
else:
print('Error, the image is neither a human nor a dog!')
# -
# ---
# <a id='step6'></a>
# ## Step 6: Test Your Algorithm
#
# In this section, you will take your new algorithm for a spin! What kind of dog does the algorithm think that _you_ look like? If you have a dog, does it predict your dog's breed accurately? If you have a cat, does it mistakenly think that your cat is a dog?
#
# ### (IMPLEMENTATION) Test Your Algorithm on Sample Images!
#
# Test your algorithm at least six images on your computer. Feel free to use any images you like. Use at least two human and two dog images.
#
# __Question 6:__ Is the output better than you expected :) ? Or worse :( ? Provide at least three possible points of improvement for your algorithm.
# __Answer:__ (Three possible points for improvement)
#
# __The output cannot be better. It succesfully classified human and dog. As for breed classification, I am not an expert and cannot tell. But according to the test accuracy, it do a good job__
#
# - feed more data into training
# - when training use more epochs
# - choose the latest and strongest pre-trained model to transfer learning
# +
## TODO: Execute your algorithm from Step 6 on
## at least 6 images on your computer.
## Feel free to use as many code cells as needed.
## suggested code, below
for file in np.hstack((human_files[:3], dog_files[:3])):
run_app(file)
|
dog_app.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Comparing Urban and Rural Streams
# In this notebook we'll compare the hydrology of two streams in the Baltimore area:
# * [Grave Run](https://waterdata.usgs.gov/md/nwis/uv?site_no=01581830), a rural stream with only 0.3% impervious surfaces, and
# * [Dead Run](https://waterdata.usgs.gov/md/nwis/uv?site_no=01589330), an urban stream that is 39% impervious surfaces.
#
# These two watersheds are similar in size, topography, and geology. They also have very emo names.
# Start with the usual.
import hydrofunctions as hf
# %matplotlib inline
hf.__version__
# ```python
# # Use HydroCloud.org to find two appropriate sites. I've selected Grave Run and Dead Run. /n
# # You can find the size and percent imperviousness of the site in the Table View.
# hf.draw_map()
# ```
# 
# request data for our two sites for a three-year period.
sites = ['01581830', '01589330']
request = hf.NWIS(sites, start_date='2002-01-01', end_date='2005-01-01').get_data()
request.ok # Verify that the data request went fine.
# We'll store our data in a dataframe named 'Q'
Q = request.df()
Q.head() # Print the first five rows to verify.
# What are the names for our two sites?
request.siteName
# Modify our dataframe so that it only includes the columns with discharge in them.
Q = Q.iloc[:,[0,2]]
# What are the column names?
Q.columns
# Rename the columns to 'Urban' and 'Rural' so we remember which is which!
Q.rename(index=str, columns={"USGS:01581830:00060:00003": "Rural", "USGS:01589330:00060:00003": "Urban"}, inplace=True)
Q.head() # List the first five rows.
# Let's plot our data to create a hydrograph. plot() is a method that is built-in to dataframes.
Q.plot()
# Let's create a flow duration chart for our two sites!
# .flow_duration is a function included in Hydrofunctions. It accepts dataframes as input.
hf.flow_duration(Q)
# ## Description of the two sites
# If you look carefully at the **hydrograph** above, you can see that the orange urban site tends to have lower baseflow,
# but it also tends to have higher peaks during storms. Unfortunately, this obscures the hydrograph for the rural site a little!
#
# The second diagram is a **flow duration chart**. The default Y axis is logarithmic, with values ranging from less than 1 to about 400 cfs. The default X axis uses a logit scale to plot the chance of exceedance. Values range from greater than zero to less than one, with a value of 0.9 meaning that 90% of the flows are higher than this value. The logit scale stretches out the extreme high and extreme low values so that the distance from the center to one standard deviation is approximately the same as from two standard deviations out to three standard deviations. This approximates the probability scale that Flow Duration charts are often plotted on.
#
# Comparing the two sites, you can see that the orange urban site has lower baseflows than the blue rural site, but also has higher peak flows.
# let's compare stats for the two sites!
Q.describe()
|
Comparing_Urban_and_Rural_Streams.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.9 (''venv'': venv)'
# language: python
# name: python3
# ---
import nltk
# nltk.download()
text_file = open(r"Datasets\paragraph.txt","r")
paragraph = ''
for line in text_file:
paragraph+=line.replace('\n',' ')
print(paragraph)
# # Cleaning the Text
from nltk.corpus import stopwords
import re
from nltk.stem import PorterStemmer, WordNetLemmatizer
def getstem_lem(paragraph,method):
#sentence tokenization
sentences = nltk.sent_tokenize(paragraph)
#to store cleaned text
corpus=[]
for i in range(len(sentences)):
clean_text = re.sub('[^a-zA-Z]',' ',sentences[i])
clean_text = clean_text.lower()
clean_text = clean_text.split()
clean_text = [method(text) for text in clean_text if text not in set(stopwords.words('english'))]
clean_text = ' '.join(clean_text)
corpus.append(clean_text)
return corpus
# ### stemmer
ps = PorterStemmer()
stem_sentences = getstem_lem(paragraph,ps.stem)
print(stem_sentences)
# ### Lemmatizer
lemma = WordNetLemmatizer()
lemma_corpus = getstem_lem(paragraph,lemma.lemmatize)
print(lemma_corpus)
# Creating model for Bag of Words
from sklearn.feature_extraction.text import CountVectorizer
cv = CountVectorizer()
X = cv.fit_transform(lemma_corpus).toarray()
X
#
|
Codes/LearningCodes/3.Bag_of_Words.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Reconstruction Sandbox
# This notebook is a test-bed for regularization and reconstruction methods
# +
# %matplotlib notebook
# %load_ext autoreload
# %autoreload 2
# Load motiondeblur module and Dataset class
import libwallerlab.projects.motiondeblur as md
from libwallerlab.utilities.io import Dataset, isDataset
# Platform imports
import os, glob
from os.path import expanduser
# Debugging imports
import llops as yp
import matplotlib.pyplot as plt
import numpy as np
yp.config.setDefaultBackend('arrayfire')
yp.config.setDefaultDatatype('float32')
# -
# ## Define Output Path
output_path = os.path.expanduser('~/datasets/motiondeblur/res_target_snr_comparison')
if not os.path.exists(output_path):
os.mkdir(output_path)
# ## Load Data
# +
# dataset_full_path = os.path.expanduser('/Users/zfphil/datasets/motiondeblur/res_target_color_coded_raster_motiondeblur_2018_05_22_19_17_45/')
dataset_full_path = '/Users/zfphil/datasets/motiondeblur/02-06-19-MotionDeblur/res_line_400_coded_raster_100_motion_deblur_2019_02_06_09_05_38/'
# Create dataset object (loads metadata)
dataset = Dataset(dataset_full_path, use_median_filter=False, subtract_mean_dark_current=False, force_type='motion_deblur')#'res' not in dataset_label)
dataset.channel_mask = [0]
# -
# ## Perform Registration and Normalization
# +
force = False
# Perform registration
dataset.motiondeblur.register(force=force)
# Perform normalization
dataset.motiondeblur.normalize(force=force)
# Perform kernel shape
dataset.metadata.calibration['blur_vector'] = {'scale': {'axis': 1, 'factor': 1}}
# -
# ## Solve For Single Segment
# +
# dataset.motiondeblur.position_segment_indicies = [4]
dataset.frame_mask = [4,5,6]
# Create recon object
recon = md.recon.Reconstruction(dataset, alpha_blend_distance=1000, pad_mode='mean', estimate_background_poly=True)
# Perform reconstruction
recon.reconstruct(iteration_count=100, step_size=1, frame_number=4, mode='global', reg_types={})
# Save result
# recon.save(output_path, filename=recon.dataset.metadata.file_header + '_reprocess', formats=['png', 'npz'], save_raw=True, downsample=4)
recon.show()
# -
recon.save(output_path, filename=recon.dataset.metadata.file_header + '_reprocess', formats=['png', 'npz'], save_raw=True, downsample=4)
recon.show()
# ## Loop Over Segments
for segment_index in dataset.motiondeblur.position_segment_indicies_full:
# Set segment index
dataset.motiondeblur.position_segment_indicies = [segment_index]
# Create recon object
recon = md.recon.Reconstruction(dataset, alpha_blend_distance=1000, pad_mode='mean', )
# Perform reconstruction
# recon.reconstruct(iteration_count=-1, step_size=1, frame_number=4, mode='global', reg_types={'l2': 1e-4})
recon.reconstruct(iteration_count=100, step_size=1, frame_number=4, mode='global', reg_types={'l2': 1e-4})
# Save result
recon.save(output_path, filename=recon.dataset.metadata.file_header, formats=['png', 'npz'], save_raw=True, downsample=4)
# +
# Try with L1 Sparsity
|
notebooks/reconstructions/snr_comparison_recon_res.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # End-To-End Example: Tip Calculator
#
# The following code calculates the amount of tip you should leave based on the amount of the check and percentage you would like to tip.
#
#
total = float(input("Enter Amount of Check: "))
tip = float(input("Enter the Tip Percentage: "))
tip_amount = total * tip
print ("You should leave this amount for a tip $%.2f" % (tip_amount))
# ## The Issues
#
# The issue with this program is that its not smart with the tip percentage. When I enter 15 is assumes 1500% not 15%.
#
# With our knowledge of strings and parsing we can make this program more intelligent:
#
# - When you enter `0.15` it uses `0.15`
# - When you enter `15` it assumes you meant `0.15` (divides by 100)
# - When you enter `15%` it assumes you meant `0.15` (removes the %, then divides by 100)
#
#
# Likewise we should do the same for currency input. Assuming the user might enter a $
# exploring how to parse percentages
x = "15 %"
y = float(x.replace('%',''))
if y >=1:
y = y/100
print(y)
# +
## Function: percentage - parses string input into a float as a percentage
## Arguments: text
## Returns float
def percentage(text):
number = float(text.replace('%',''))
if number >1:
number = number /100
return number
## Function: currency - parses string input into a float as currency
## Arguments: text
## Returns float
def currency(text):
number = float(text.replace('$',''))
return number
# -
total = currency(input("Enter Amount of Check: "))
tip = percentage(input("Enter the Tip Percentage: "))
tip_amount = total * tip
print ("You should leave this amount for a tip $%.2f" % (tip_amount))
|
content/lessons/06/End-To-End-Example/ETEE-Tip-Calculator.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.2 64-bit (''python_fundamentos_dsa'': conda)'
# name: python392jvsc74a57bd050c93bd6a88d5a56fafcaaf86f19d30d33d3068901c9b6ce1cc80821349e3a1a
# ---
# # <font color='blue'>Data Science Academy - Python Fundamentos - Capítulo 4</font>
#
# ## Download: http://github.com/dsacademybr
# Versão da Linguagem Python
from platform import python_version
print('Versão da Linguagem Python Usada Neste Jupyter Notebook:', python_version())
# # Expressões Lambda
# Definindo uma função - 3 linhas de código
def potencia(num):
result = pow(num, 2)
return result
potencia(5)
# Definindo uma função - 2 linhas de código
def potencia(num):
return pow(num, 2)
potencia(5)
# Definindo uma função - 1 linha de código
def potencia(num): return pow(num, 2)
potencia(5)
# Definindo uma expressão lambda
potencia = lambda num: num**2
potencia(5)
# Lembre: operadores de comparação retornam boolean, true or false
Par = lambda x: x%2==0
Par(3)
Par(4)
first = lambda s: s[0]
type(first)
first('Python')
reverso = lambda s: s[::-1]
type(reverso)
reverso('Python')
addNum = lambda x,y : x+y
addNum(2,3)
# # Fim
# ### Obrigado - Data Science Academy - <a href="http://facebook.com/dsacademybr">facebook.com/dsacademybr</a>
|
Cap03/Notebooks/DSA-Python-Cap03-07-Lambda.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import numpy as np
import time
import h5py
import keras
import pandas as pd
import math
import joblib
import matplotlib.pyplot as plt
from fuel.datasets.hdf5 import H5PYDataset
from sklearn.metrics import accuracy_score
from sklearn.model_selection import StratifiedShuffleSplit
from IPython.display import display
from keras.layers import Input, Dense, Lambda, Flatten, Reshape, BatchNormalization, Activation, Dropout
from keras.layers import Conv2D, Conv2DTranspose, MaxPooling2D, UpSampling2D
from keras.callbacks import EarlyStopping
from keras.optimizers import RMSprop, Adam, SGD
from keras.models import Model, Sequential
from keras.utils import np_utils
from keras import backend as K
from keras_tqdm import TQDMNotebookCallback
from keras.applications.inception_v3 import InceptionV3
from keras.preprocessing import image
from keras.models import Model
from keras.layers import Dense, GlobalAveragePooling2D
from keras import backend as K
# +
ftrain = H5PYDataset("../../data/cifar10/cifar10.hdf5", which_sets=('train',))
X_train, y_train = ftrain.get_data(ftrain.open(), slice(0, ftrain.num_examples))
X_train = np.moveaxis(X_train[:], 1, 3) / 255.
ftest = H5PYDataset("../../data/cifar10/cifar10.hdf5", which_sets=('test',))
X_test, y_test = ftest.get_data(ftest.open(), slice(0, ftest.num_examples))
X_test = np.moveaxis(X_test[:], 1, 3) / 255.
print(X_train.shape, y_train.shape)
print(X_test.shape, y_test.shape)
# +
# input image dimensions
img_rows, img_cols, img_chns = 32, 32, 3
# number of convolutional filters to use
num_classes = 10
batch_size = 100
original_img_size = (img_rows, img_cols, img_chns)
epochs = 1000
# +
def create_model():
x = Input(shape=(32, 32, 3))
up = UpSampling2D(size=7)(x)
base_model = InceptionV3(input_tensor=up, weights='imagenet', include_top=False)
avg_pool = GlobalAveragePooling2D()(base_model.output)
dense_1 = Dense(512, activation='relu')(avg_pool)
dropout_1 = Dropout(0.5)(dense_1)
y = Dense(10, activation='softmax')(dropout_1)
model = Model(inputs=x, outputs=y)
for layer in base_model.layers:
layer.trainable = False
opt = RMSprop(lr=0.0001, decay=1e-6)
model.compile(optimizer=opt, loss='categorical_crossentropy', metrics=['accuracy'])
return model
create_model().summary()
# -
results = []
for sample_size in [1000, 2000, 5000, 10000, 25000, 50000]:
start = time.time()
print('Fitting with sample_size: {}'.format(sample_size))
if sample_size < len(X_train):
sss = StratifiedShuffleSplit(n_splits=2, test_size=sample_size / len(X_train), random_state=0)
_, index = sss.split(X_train, y_train)
X, y = X_train[index[1]], y_train[index[1]]
else:
X, y = X_train, y_train
y = np_utils.to_categorical(y)
model = create_model()
model.fit(X, y, shuffle=True,
epochs=epochs,
batch_size=batch_size,
verbose=0,
callbacks=[TQDMNotebookCallback(),
EarlyStopping(monitor='loss', min_delta=0.01, patience=50)])
y_pred = np.argmax(model.predict(X_test), axis=-1)
score = accuracy_score(y_test, y_pred)
end = time.time()
elapsed = end - start
print(' * Accuracy: %.1f %%' % (100. * score))
print(' * Fit time elapsed: %.1fs' % elapsed)
results.append({'sample_size': sample_size, 'accuracy': score, 'time': elapsed})
df = pd.DataFrame(results)
display(df)
df.to_csv('inception_results.csv', index=False)
|
notebooks/vae-semi_supervised_learning/inception.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Classification Avancée
# # projet 4
# ## Séparateur à Vaste Marge et Méthode à Noyaux
#
# ## Machines à Vecteurs de Support (SVM)
#
# On commence par (essayer... d') installer un module qui permet de plus jolies visualisation que le matplotlib de base:
# !pip3 install seaborn --user
# ## Première partie : prise en main des SVM
# Cette partie est librement inspirée du travail de <NAME>, auteur du livre [Python Data Science Handbook](http://shop.oreilly.com/product/0636920034919.do). Son [GitHub](https://github.com/jakevdp/PythonDataScienceHandbook) (en anglais) regorge de fichiers utiles.
# Dans un premier temps, on va générer des données jouets, linéairement séparables :
# +
# %matplotlib inline
import matplotlib.pyplot as plt
#Un petit environment qui donne de meilleurs graphes
#import seaborn as sns; sns.set()
# fonction sklearn pour générer des données simples
from sklearn.datasets.samples_generator import make_blobs
X, y = make_blobs(n_samples=50, centers=2,
random_state=0, cluster_std=0.60)
# Affichage des données
plt.scatter(X[:, 0], X[:, 1], c=y, s=50, cmap='prism');
# -
# ### SVM linéaire (=perceptron à vaste marge)
# On va commencer par apprendre un SVM sans noyau (on dit "linéaire") à l'aide de scikit-learn :
#import de la classe - qui s'appelle SVC et pas SVM...
from sklearn.svm import SVC
#Définition du modèle
model = SVC(kernel='linear', C=1E10)
#Apprentissage sur les donnée
model.fit(X, y)
# On va utiliser une fonction d'affichage qui va bien, où tout ce qui est nécessaire est affiché. Il ne faut pas forcément la comprendre complètement de suite car on reviendra plus bas sur les points importants.
# +
import numpy as np
def affiche_fonction_de_decision(model, ax=None, plot_support=True):
"""Affiche le séparateur, les marges, et les vecteurs de support d'un SVM en 2D"""
if ax is None:
ax = plt.gca()
xlim = ax.get_xlim()
ylim = ax.get_ylim()
# création de la grille pour l'évaluation
x = np.linspace(xlim[0], xlim[1], 30)
y = np.linspace(ylim[0], ylim[1], 30)
Y, X = np.meshgrid(y, x)
xy = np.vstack([X.ravel(), Y.ravel()]).T
P = model.decision_function(xy).reshape(X.shape)
# affichage de l'hyperplan et des marges
ax.contour(X, Y, P, colors='k',
levels=[-1, 0, 1], alpha=0.5,
linestyles=['--', '-', '--'])
# Affichage des vecteurs de support
if plot_support:
ax.scatter(model.support_vectors_[:, 0],
model.support_vectors_[:, 1],
s=300, linewidth=1, facecolors='none', edgecolor='black');
ax.set_xlim(xlim)
ax.set_ylim(ylim)
# -
# Voyons ce que cela donne sur notre séparateur linéaire à vaste marge :
plt.scatter(X[:, 0], X[:, 1], c=y, s=50, cmap='prism')
affiche_fonction_de_decision(model);
# Sur ce graphe, on voit le séparateur (ligne pleine), les vecteurs de support (points entourés) et la marge (matérialisée par des lignes discontinues).
# On a ici le séparateur qui maximise la marge.
# Scikit-learn nous permet, après apprentissage, de récupérer les vecteurs de supports:
model.support_vectors_
# Seuls trois données sont utiles pour classer de nouvelles données. On peut s'en assurer en rajoutant des données sans changer le modèle :
#
# +
X2, y2 = make_blobs(n_samples=200, centers=2,
random_state=0, cluster_std=0.60)
model2 = SVC(kernel='linear', C=1E10)
model2.fit(X2, y2)
plt.scatter(X2[:, 0], X2[:, 1], c=y2, s=50, cmap='prism')
affiche_fonction_de_decision(model2);
# -
# ### SVM non linéaire
# Comme on l'a vu ce matin, la puissance des séparateurs linéaires est limitée (à des données linéairement séparables). Mais il est possible de contourner cette limitation par l'utilisation de noyaux.
#
# On va commencer par générer des données non-linéairement séparables :
# +
from sklearn.datasets.samples_generator import make_circles
X, y = make_circles(100, factor=.1, noise=.1)
clf = SVC(kernel='linear').fit(X, y)
plt.scatter(X[:, 0], X[:, 1], c=y, s=50, cmap='Dark2')
affiche_fonction_de_decision(clf, plot_support=False)
# -
# Clairement notre apprentissage de séparateur linéaire a échoué...
#
# On va manuellement ajouter une troisième dimension *z* :
z = np.exp(-(X ** 2).sum(1))
# On peut afficher les données augmentées et se rendre compte qu'elles sont linéairement séparables :
from mpl_toolkits.mplot3d import Axes3D
ax = plt.subplot(projection='3d')
ax.scatter3D(X[:, 0], X[:, 1], z, c=y, s=50, cmap='Dark2')
ax.view_init(elev=30, azim=30)
ax.set_xlabel('x')
ax.set_ylabel('y')
ax.set_zlabel('z')
# Le plan définit par z=0.7 (par exemple) sépare les 2 classes parfaitement.
#
# Bien entendu, la projection en plus grande dimension est capitale, et en choisissant un autre calcul pour *z* on aurait obtenu dans la plupart des cas des données non linéairement séparables.
#
# Et s'il fallait faire effectivement la projection, cela limiterait drastiquement la dimension de l'espace de plongement ainsi que le nombre de données traitables. C'est pourquoi l'utilisation de noyaux (kernels en anglais) est d'une grande efficacité.
#
# En Scikit-Learn, il suffit de modifier le paramètre *kernel* : jusqu'à présent, nous avons utilisé 'linear' comme valeur. On peut par exemple utiliser *rbf* pour 'radial basis function', le noyau gaussien, et il nous reste à trouver la bonne valeur du paramètre :
clf = SVC(kernel='rbf', gamma='auto', C=1E10)
clf.fit(X, y)
plt.scatter(X[:, 0], X[:, 1], c=y, s=50, cmap='Dark2')
affiche_fonction_de_decision(clf)
# On voit ici que le séparateur (et la marge associée) ne sont pas linéaire dans l'espace des données, mais qu'ils peuvent s'y représenter sans difficulté.
#
# Notons aussi que le nombre de vecteurs de support reste très petit.
# ### SVM à marge douce
#
# Il est aussi possible que le problème soit linéairement séparable (dans la dimension initiale des données ou dans un plongement) mais que le bruit (=la mauvaise qualité des données) empêche l'apprenant de trouver un séparateur.
#
# On utilise alors ce que l'on appelle un classifieur à marge douce : on autorise alors certains points à être dans la marge. C'est le role du paramètre *C* : pour des grosses valeurs, on est quasiment en marge dure, mais plus *C* prend des petites valeurs, plus les marges deviennent permissibles.
#
# On va prendre des données qui se chevauchent un peu :
X, y = make_blobs(n_samples=100, centers=2,
random_state=0, cluster_std=1.2)
plt.scatter(X[:, 0], X[:, 1], c=y, s=50, cmap='jet');
# On joue alors avec la valeur de *C*
# +
fig, ax = plt.subplots(1, 2, figsize=(16, 6))
fig.subplots_adjust(left=0.0625, right=0.95, wspace=0.1)
for axi, C in zip(ax, [10.0, 0.1]):
model = SVC(kernel='linear', C=C).fit(X, y)
axi.scatter(X[:, 0], X[:, 1], c=y, s=50, cmap='jet')
affiche_fonction_de_decision(model, axi)
axi.scatter(model.support_vectors_[:, 0],
model.support_vectors_[:, 1],
s=300, lw=1, facecolors='none');
axi.set_title('C = {0:.1f}'.format(C), size=14)
# -
# Bien entendu, il est possible de combiner l'utilisation d'un noyau et de marges douces.
# ### Tuner un SVM
# Tous les noyaux sont paramétrés : il est question ici d'étudier l'impact d'un (hyper)paramètre sur la qualité de l'apprentissage.
# On va générer des données qui ne sont pas linéairement séparables :
X, y = make_blobs(n_samples=200, centers=2,
random_state=0, cluster_std=1.3)
plt.scatter(X[:, 0], X[:, 1], c=y, s=50, cmap='prism')
# On va étudier 2 noyaux différents
# - le noyau polynomial (*kernel='poly'*) qui a 2 paramètres, *degree* qu'il faut faire varier entre 2 et 6 (au minimum), et *C* (lié à la 'douceur' de la marge)
# - le noyau gaussien (*kernel='rbf'*) qui a aussi 2 paramètres, *gamma*, qu'il faut faire varier de 1 à 0.01, et *C*
#
# A chaque fois, en plus de l'affichage des séparateurs, il serait intéressant de regarder combien de vecteurs de support le classifieur appris a besoin.
#
# A vous de jouer !
#
# +
# A vous
from sklearn.model_selection import train_test_split
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import classification_report
from sklearn.svm import SVC
# -
####
from sklearn.model_selection import train_test_split
# production de deux sous-échantillons
X_train, X_test, y_train, y_test = train_test_split(X,y,test_size=0.30, random_state=42)
# +
# trouver le bon parametre entre ces differents hyper parametres
tuned_parameters = [{'kernel': ['poly'], 'degree': [2,3,4,5,6],'C': [1, 10, 100, 1000]},
{'kernel': ['rbf'], 'gamma':[1,0.5,0.1,0.05,0.01],'C': [1, 10, 100, 1000]}]
scores = ['precision', 'recall']
for score in scores:
print("# Tuning hyper-parameters for %s" % score)
print()
clf = GridSearchCV(SVC(), tuned_parameters, cv=5, scoring='%s_macro' % score)
clf.fit(X_train, y_train)
print("Best parameters set found on development set:")
print()
print(clf.best_params_)
print()
print("Grid scores on development set:")
print()
means = clf.cv_results_['mean_test_score']
stds = clf.cv_results_['std_test_score']
for mean, std, params in zip(means, stds, clf.cv_results_['params']):
print("%0.3f (+/-%0.03f) for %r"
% (mean, std * 2, params))
print()
print("Detailed classification report:")
print()
print("The model is trained on the full development set.")
print("The scores are computed on the full evaluation set.")
print()
y_true, y_pred = y_test, clf.predict(X_test)
print(classification_report(y_true, y_pred))
print()
# +
#Best parameters set found on development set:
#{'C': 1, 'gamma': 0.1, 'kernel': 'rbf'}
# +
fig, ax = plt.subplots(1, 2, figsize=(16, 6))
fig.subplots_adjust(left=0.0625, right=0.95, wspace=0.1)
for axi, C in zip(ax, [10.0, 0.1]):
model = SVC(kernel='linear', C=C).fit(X, y)
axi.scatter(X[:, 0], X[:, 1], c=y, s=50, cmap='jet')
affiche_fonction_de_decision(model, axi)
axi.scatter(model.support_vectors_[:, 0],
model.support_vectors_[:, 1],
s=300, lw=1, facecolors='none');
axi.set_title('C = {0:.1f}'.format(C), size=14)
# -
###
#E10
#Exemple avec le noyau gaussien et des valeurs pour gamma et C
clf = SVC(kernel='rbf', gamma=0.1, C=1)
clf.fit(X, y)
plt.scatter(X[:, 0], X[:, 1], c=y, s=50, cmap='prism')
affiche_fonction_de_decision(clf)
print("Nombre de vecteurs de support (sur 200 données) :", len(clf.support_vectors_))
# +
#nombre de SVM 50 vecteurs
# -
# ## Deuxième partie : un traitement (presque) complet
# ### Préparation des données
# Nous allons utiliser un jeu de données réel - tiré de *Tsanas & Xifara : Accurate quantitative estimation of energy performance of residential buildings using statistical machine learning tools, Energy and Buildings, Vol. 49, pp. 560-567, 2012* - qu'il vous faut d'abord télécharger :
# !wget http://pageperso.lif.univ-mrs.fr/~remi.eyraud/data.csv
# Les 8 premières colonnes correspondent aux attributs descriptifs et les deux dernières, aux charges de chauffage et de climatisation (dans cet ordre).
# Pour les utiliser en Python, vous pourrez vous servir du code suivant :
data = np.loadtxt("./data.csv")
X = data[:,:-2]
Y = data[:,-2:]
Yheat = Y[:,0]
Ycool = Y[:,1]
# Le problème initial, tel que présenté ici, est un problème de régression. Nous allons d'abord le transformer en problème de classification. Par une méthode de clustering, on veut répartir les charges de chauffage et de climatisation en 3 classes : faibles, moyennes, élevées.
# Le seul trick : les Y sont des vecteurs et les classifieurs sklearn ont besoin d'array :
# il faut les reshaper : Yheat_vector = Yheat.reshape(-1,1)
Yheat_vector = Yheat.reshape(-1,1)
Ycool_vector = Ycool.reshape(-1,1)
# A vous de jouer :
# +
from sklearn.cluster import KMeans
# La suite ? il s'agit de définir un classifieur du k-means avec k=3
# et d'utiliser la méthode 'fit' sur les 2 ensembles de valeurs Y
kmeans = KMeans(n_clusters=3)
y_cool=kmeans.fit (Ycool_vector).labels_
y_heat=kmeans.fit (Yheat_vector).labels_
y_kmeans = kmeans.fit(Ycool_vector,Yheat_vector).labels_
#kmeans.labels_
# Après apprentissage du kmeans, les classes des données utilisées sont stockées dans mon_classifieur.labels_
# -
# ### Apprentissage
# Nous voulons comparer plusieurs méthodes d'apprentissage :
# 1. Les k-plus proches voisins (*KNeighborsClassifier* de la classe *sklearn.neighbors*, hyperparamètre à régler : *n_neighbors*)
# 2. Les arbres de décision (*DecisionTreeClassifier* de la classe *sklearn.tree*, hyperparamètre à régler : *max_depth*)
# 3. Perceptron linéaire (*Perceptron* de la classe *sklearn.linear_model*, hyperparamètre à régler : aucun/*max_iter* en sklearn 0.19 ou plus récent)
# 4. SVM à noyau gaussien (*SVC* avec *kernel='rbf'* de la classe *sklearn.svm*, hyperparamètre à régler : *gamma*)
# 5. SVM à noyau polynomial (*SVC* avec *kernel='poly'* de la classe *sklearn.svm*, hyperparamètre à régler : *degree*)
#
# Ecrivez le code permettant de :
# 1. Séparer les données en un échantillon d'apprentissage et un échantillon de test (80/20)
#
# 2. Sélectionner les meilleurs valeurs des hyperparamètres sur l'échantillon d'apprentissage par validation croisée en utilisant 10 folders
#
#
# separation des données 80/20
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X,y_kmeans,test_size=0.20, random_state=42)
# +
#Apprentissage trouver les bons params
from sklearn.model_selection import GridSearchCV
#1 Les k-plus proches voisins n_neighbors
from sklearn import neighbors as nn
import numpy as np
n = np.arange(1,126,1)
tuned_parameters = [{'n_neighbors': n}]
clf_knn = GridSearchCV(nn.KNeighborsClassifier(), tuned_parameters, cv=10)
clf_knn.fit(X_train, y_train)
print("Best parameters set found on development set knn:")
print(clf_knn.best_params_)
# +
#2Les arbres de décision *DecisionTreeClassifier* max_depth
from sklearn.tree import DecisionTreeClassifier
import numpy as np
m = np.arange(1,100,1)
tuned_parameters = [{'max_depth': m}]
clf_DT = GridSearchCV( DecisionTreeClassifier(), tuned_parameters, cv=10)
clf_DT.fit(X_train, y_train)
print("Best parameters set found on development set knn:")
print(clf_DT.best_params_)
# -
# 3 Perceptron linéaire max_iter
from sklearn.linear_model import Perceptron
import numpy as np
n = np.arange(1,1000,1)
tuned_parameters = [{'max_iter': n}]
clf_P = GridSearchCV(Perceptron(), tuned_parameters, cv=10)
clf_P.fit(X_train, y_train)
print("Best parameters set found on development set knn:")
print(clf_P.best_params_)
# +
#4 SVM à noyau gaussien
from sklearn.svm import SVC
import numpy as np
tuned_parameters = [ {'kernel': ['rbf'],
'gamma':[1,0.9,0.8,0.7,0.6,0.5,0.4,0.3,0.2,0.1,0.09,0.08,0.07,0.06,0.05,0.04,0.03,0.02,0.01]}]
clf_rbf = GridSearchCV(SVC(), tuned_parameters, cv=10)
clf_rbf.fit(X_train, y_train)
print("Best parameters set found on development set knn:")
print(clf_rbf.best_params_)
# +
#5. SVM à noyau polynomial (*SVC* avec *kernel='poly'* *degree*)
from sklearn.svm import SVC
import numpy as np
tuned_parameters = [ {'kernel': ['poly'],
'degree': [2,3,4,5]}]
clf_poly = GridSearchCV(SVC(), tuned_parameters, cv=10)
clf_poly.fit(X_train, y_train)
print("Best parameters set found on development set knn:")
print(clf_poly.best_params_)
# -
#Degree of the polynomial kernel function (‘poly’). Ignored by all other kernels.
# ### Analyse des résultats
# Afficher sur une courbe les scores de chacun des algorithmes avec la meilleure valeur d'hyperparamètre possible sur l'échantillon de test.
# +
# A vous
from sklearn.model_selection import cross_val_score
import matplotlib.pyplot as plt
# Puis : Commande uniquement Notebook : permet l'affichage des courbes
# %matplotlib inline
score = []
reussite_knn =[]
reussite_DT =[]
reussite_P =[]
reussite_rbf =[]
y = []
# Cross-validation pour chaque classifieur sur l échantillon test
#1 knn
clf_knn=nn.KNeighborsClassifier(n_neighbors=4)
clf_knn.fit(X_train, y_train)
s1 = clf_knn.score(X_test, y_test)
reussite_knn = cross_val_score(clf_knn, X_train, y_train, cv = 10)
reussite_knn_std = reussite_knn.std()
#2 DecisionTreeClassifier()
clf_DT = DecisionTreeClassifier(max_depth=15)
clf_DT.fit(X_train, y_train)
s2 = clf_DT.score(X_test, y_test)
reussite_DT = cross_val_score(clf_DT, X_train, y_train, cv = 10)
reussite_DT_std = reussite_DT.std()
#3
clf_P = Perceptron(max_iter=828)
clf_P.fit(X_train, y_train)
s3 = clf_P.score(X_test, y_test)
reussite_P = cross_val_score(clf_P, X_train, y_train, cv = 10)
reussite_P_std = reussite_P.std()
#4
clf_rbf = SVC(kernel='rbf',gamma=0.3)
clf_rbf.fit(X_train, y_train)
s4 = clf_rbf.score(X_test, y_test)
reussite_rbf = cross_val_score(clf_rbf, X_train, y_train, cv = 10)
reussite_rbf_std = reussite_rbf.std()
score =[s1 , s2 ,s3 ,s4]
y =[1,2,3,4]
plt.figure()
plt.plot(y,score)
plt.title("Evolution du score avec toutes les methodes")
plt.xlabel("Valeur random")
plt.ylabel("score")
plt.show()
# -
score
# +
# donc meilleur methode qui à le score s2 clf_DT = DecisionTreeClassifier(max_depth=15)
# -
# Pour chacune des méthodes, pour chaque meilleur hyperparamètre, calculer l'intervalle à 95% de confiance auquel le score doit appartenir en utilisant les résultats de la validation croisée. Si vous ne vous souvenez plus de comment on calcule un interval de confiance, vous pouvez consulter : https://fr.wikihow.com/calculer-un-intervalle-de-confiance
# +
# A vous
# Calculez l’écart-type de la population (σ) et la taille de l’échantillon (n). * le z score
# on a 95 % donc z score 1,96
# +
import math
n= len(X)
d =math.sqrt( n )
ref = 1.96
p = 0.95
zs = p * ref
m1 = (reussite_knn_std/d) * zs
m2 = (reussite_DT_std/d) * zs
m3 = (reussite_P_std/d) * zs
m4 = (reussite_rbf_std) * zs
# -
print(m1,m2,m3,m4)
# Quelle méthodes est la meilleure pour prédire la classe de frais de chauffage ? De frais de climatisation ?
# +
#on peut dire que la meilleur methode pour prédire les deux classe ces la 4 eme methode
#clf_rbf = SVC(kernel='rbf',gamma=0.3)
# puisque elle a un intervalle de confiance plus grand
# -
#
|
CA4.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python 201.2
# ## Nível intermediário em Python
#
# Os notebooks dessa segunda etapa, focam especificamente em features intermediárias/avançadas da linguagem.
#
# Tenha em mente que algumas questões apresentadas neste notebook, farão referência aos arquivos .py encontrados dentro do diretório src no mesmo nível.
# ### Iterators
#
# Iteradores em python estão por toda parte e são muito utilizados, durante este treinamento já utilizamos vários deles (como listas, tuplas e etc).
#
# *"Em programação de computadores, um iterador se refere tanto ao objeto que permite ao programador percorrer um container, (uma coleção de elementos) particularmente listas, quanto ao padrão de projetos Iterator, no qual um iterador é usado para percorrer um container e acessar seus elementos. O padrão Iterator desacopla os algoritmos dos recipientes, porém em alguns casos, os algoritmos são necessariamente específicos dos containers e, portanto, não podem ser desacoplados."* [wiki](https://pt.wikipedia.org/wiki/Iterador)
#
# O protocolo Iterator é definido na [PEP234](https://www.python.org/dev/peps/pep-0234/), que defini uma interface de iterações que objetos podem implementar.
#
# Uma das maneiras de se criar um objeto iterável é através do método built-in **iter()**.
alphabet = iter('abc')
while True:
try:
print('it: ', next(alphabet))
except StopIteration as e:
print('Exception : StopIteration')
break
# Entretanto, conforme a PEP 234 defini, podemos criar um objeto que seja iterável. Para isso é necessário criar uma classe e implementar dois métodos:
#
# - \_\_iter__: Método que deve retornar o próprio objeto a ser iterado (no caso **self**).
# - \_\_next__: Retorna o próximo valor da iteração. Caso tenha se esgotado os itens da iteração deve lançar a exceção **StopIteration**.
# +
# Implementação de Fibonacci baseada na Abordagem Iterativa
# https://pt.wikipedia.org/wiki/Sequ%C3%AAncia_de_Fibonacci#Abordagem_iterativa
class Fibonacci:
def __init__(self, number):
self.start(number)
def start(self, number):
self.number = number + 1
self.__index = 0
self.__j = 1
self.__i = 0
def __iter__(self):
return self
def __next__(self):
# Fibonacci
t = self.__index
# from 2 to number + 1
if 1 < self.__index <= self.number:
t = self.__i + self.__j
self.__i = self.__j
self.__j = t
# Iterator Exception
if self.__index == self.number:
raise StopIteration
self.__index += 1
return t
# 0, 1, 1, 2, 3, 5, 8, 13, 21, 34, 55, 89, 144,233, 377, 610, 987
# 1597, 2584, 4181, 6765, 10946
fib = Fibonacci(21)
nums = []
for i in fib:
nums.append(i)
print('Fibonacci Sequence: ')
print(nums)
print('-' * 20)
print('Restart and iterate over while and next:')
fib.start(8)
i = 0
while True:
try:
print(f'num {i}: ', next(fib))
i += 1
except StopIteration as si:
print('StopIteration : exception')
break
# -
# ### [Generators](https://docs.python.org/3/tutorial/classes.html#generators)
#
# Generators são ferramentas simples que nos permitem criar novos iteradores!
#
# Para utilizá-los devemos utilizar a palavras reservada **yield**, acoplando isso dentro de uma função.
#
# Os métodos **__iter__** e **__next__** são criados automaticamente quando utilizamos a criação de geradores.
#
# E um grande diferencial, é que eles mantém o estado de execução e consomem menos memória em relação a lista, já que nem todos os elementos são previamente gerados, mas sim são gerandos em tempo de execução.
# +
# gerado simples
def simple_gen(x):
for m in range(x):
yield m
m = simple_gen(10)
while True:
try:
print(next(m))
except StopIteration as si:
print('StopIteration : exception')
break
# +
# Exemplo de um gerador criado de objetos
class Cat:
def __init__(self, name):
self.name = name
def gerador_gatos(num):
for i in range(num):
yield Cat(f'Cat {i}')
gc = gerador_gatos(10)
print(next(gc).name)
print(next(gc).name)
# -
# O exemplo acima é bem verboso, podemos usar algo parecido com compreensão de listas, mas para geradores.
gc = (Cat(f'Cat {i}') for i in range(10))
print('Type:', type(gc))
print('First Iterable item:', next(gc).name)
# ### Coroutines
#
# Diferentemente dos geradores as corotinas são generalizações de subrotinas, elas podem consumir dados.
#
# Utilizando o operador **yield** podemos criar uma coroutine.
#
# Normalmente, são muito utilizadas para processamento em "paralelo" (na verdade concorrência) visto que ao chamar multíplas corotinas é possível alterando a execução devido a "parada" propiciada pelo operador yield.
#
# Nas novas versões da linguagem Python 3.5+, foi adicionado métodos assíncronos, usando async/await... na verdade as coroutines são o coração desse novo método de execução assíncrona na linguagem.
#
# Veremos async/await mais a frente no treinamento.
# +
import random
def producer(item):
print(f'Producing item {item}')
yield item
def consumer(name):
call = 0
while True:
item = list((yield))[0]
print(f'{name} n: {call} is doing work on item %s' % item)
call += 1
messengers = [consumer('Consumer 1'), consumer('Consumer 2')]
next(messengers[0])
next(messengers[1])
for i in range(8):
messenger = messengers[random.randint(0, 1)]
item = producer(f'item: {i}')
messenger.send(item)
# +
def coro():
hello = yield "Hello"
yield hello
c = coro()
print(next(c))
print(c.send("World"))
# -
# ### Programação Funcional
#
# Apesar do Python suportar algum tipo de programação funcional, originalmente a linguagem não foi construída para este tipo de estrutura, como eu caso de Haskell, Elm, Elixir.
#
# Entretanto, funções em Python são First-Class Citizen, existe Closure e algumas outras funções com aspecto funcional, entretanto, não existe necessariamente um paradigma funcional completo na linguagem.
#
# Mas vamos explorar melhor os aspectos funcionais da linguagem.
# #### map, reduce, filter e lambda
#
# Dentre as funções voltadas para o paradigma funcional temos map, reduce e filter.
#
# - **map**: Aplica uma função a cada elemento de uma sequência.
# - **reduce**: Aplica uma função a cada elemento de uma sequência e agrega a um total.
# - **filter**: Realiza um filtro de cada elemento dentro da sequência.
#
# Outro paradigma é a utilização de lambda expression, que é nada mais nada menos que funções anônimas de escopo restrito que aceitam argumentos e suportam apenas uma expressão.
# +
# Exemplo de map
lista = [1, 2, 3, 4, 5, 6, 7]
def sqrt(x):
return x*x
print('map : sqrt : ', list(map(sqrt, lista)))
# +
# Exemplo de reduce
from functools import reduce
def add(x, y):
return x + y
# (1 + 2) + (3 + 4) + (5 + 6) + 7 = 28
print('reduce = 28 : is ok? => ', reduce(add, lista) == 28)
# (1 + 2) + (3 + 4) = 10
print('reduce = 10 : is ok? => ', reduce(add, [1, 2, 3, 4]) == 10)
# +
# Exemplo de filter
lista = [None, 1, 2, 3, None, 4]
def gt(x):
"""greater than."""
if x:
return x > 2
return False
print('filter, remover None: ', list(filter(None, lista)))
print('filter, gt > 2 only:', list(filter(gt, lista)))
# -
# Claro, que ao invés de definirmos expressões tão curtas no formato de funções, podemos na verdade definir expressões lambdas para isso.
# +
print('sqrt: ', list(map(lambda x : x*x, [1, 2, 3, 4, 5])))
k = [[1, 2], [3, 4]]
# Colocando a função, gerada por lambda em uma variável!
sum_up = lambda x : x[0] + x[1]
r = list(map(sum_up, k))
print(f'Soma de {k}: ', r)
# -
# #### Special attributes
#
# - \_\_doc__
# - \_\_name__
# - \_\_defaults__
# - \_\_dict__
# - \_\_module__
# - more...
# +
def linear(a, x, b=0):
"""Simple linear funcion expression (a * x) + b."""
return (a * x) + b
k = linear
print(f"Function docs: {k.__doc__}")
print(f"Function name: {k.__name__}")
print(f"Function name: {k.__defaults__}")
# -
# #### Closure
#
# Closure refere-se a capacidade de se criar funções dentro do escopo de outras funções encapsulando assim determinado comportamento de modo interno apenas.
#
# Na prática Closure funciona quase como uma representação mais simples de um objeto. Em javascript nos meados da internet, essa definição de Closures foi muito utilizada (e ainda é muito ainda hoje, principalmente nos frameworks).
#
# Closure são bem importantes, pois através desse paradigma podemos definir comportamentos a nossas funções de modo padronizado, e closure é um primeiro passo para o entendimento de decoradores.
#
# +
def master(x):
y = 2
def slave():
return x + y
return slave
closure = master(5)
print('resultado: ', closure())
# -
# ### Collections
#
# Durante muito tempo, dentro da comunidade python existiu (e muitas vezes ainda é muito mencionado nos dias atuais) a menção de que python tem baterias inclusas.
#
# Dentre essas baterias inclusas existe o módulo de collections, o qual possui funcionalidades muitos interessantes e que podem facilitar muito a vida do desenvolvedor, para que o mesmo não precise reinvetar a roda!
#
# Vamos apresentar alguns dos principais, mas existem muitas outras classes dentro deste módulo. [Veja todos aqui](https://docs.python.org/3.7/library/collections.html).
#
# - **namedtuple**
# - **Counter**
# #### namedtuple
#
# As namedtuple são estruturas bem interessantes, elas fornecem o controle das tuplas, e ao mesmo tempo permitem uma estrutura de dados melhor e mais formatada.
#
# Muitas vezes elas são utilizadas para substituir pequenas classes que não possuem ações (métodos). Por esse mesmo motivo, em versões mais novas do Python foi implemetando os objetos com a assinatura de DataClass (que veremos mais a frente).
#
# Mesmo assim, namedtuples podem ser uma alternativas viável e interessante!
# +
from collections import namedtuple
Person = namedtuple('Person', ['name', 'age', 'gender'])
print('Type:', type(Person))
rodolfo = Person('Rodolfo', 36, 'Masculino')
print('Type:', type(rodolfo))
print(rodolfo.name)
# -
# #### Counter
#
# O objeto Counter permite realizar a contagem de hash.
#
# Parece uma tarefa estranha, mas caso você possua dicionários com valores, poderia ser meio verboso e chato criar uma rotina para realizar o merge e contagem de seus valores, por este motivo o Counter pode lhe ajudar muito!!!
# +
from collections import Counter
a = Counter({'a': 10, 'b': 5, 'c': 1})
b = Counter({'a': 1, 'b': 2, 'c': 0, 'd': 15})
print('Type:', type(a))
print(a + b)
# -
# ### DataClass ([PEP 557](https://www.python.org/dev/peps/pep-0557))
#
# Na nova versão da linguagem (3.7+) temos a implementação da **DataClass**, o qual cria uma **decorador** (que veremos mais a frente como utilizar), para anotar a classe e adicionar diversos métodos já implementados nela.
#
# Para efetivamente utilizarmos esta nova feature da linguagem, precisamos também utilizar a nova feature de type annotations (PEP), a qual adicionar tipos estáticos a linguagem python. Entretanto, as type annotations servem apenas como forma de documentação e inferência de tipos das funções para linters da linguagem python acoplados dentro de IDE's (como o PyCharm).
# +
from dataclasses import dataclass
# Old class to mimic DataClass
class OldPoint:
def __init__(self, x=0.0, y=0.0):
self.x = x
self.y = y
def __repr__(self):
return f'{self.__class__.__name__}(x={self.x}, y={self.y})'
def __eq__(self, other):
if other.__class__ is not self.__class__:
return NotImplemented
return other.x == self.x and other.y == self.y
def draw(self):
return (self.x, self.y)
# New Python 3.7 DataClass
@dataclass
class Point:
x: float = 0.0
y: float = 0.0
def draw(self):
return f'Drawing point at ({self.x}, {self.y})'
po0, po1 = OldPoint(), OldPoint()
pn0, pn1 = Point(), Point()
print(po0)
print(pn0)
print('Equals operation: ', po0 == po1)
print('Equals operation: ', pn0 == pn1)
print('Method call:', pn0.draw())
# -
# Portanto, uma DataClass, já implementa para nós diversas funcionalidades, que antigamente o desenvolvedor Python deveria manualmente implementar caso quisesse possuir uma classe bem estruturada.
#
# É possível ver mais exemplos de uso, e possíveis perigos no uso incorreto, no excelente [Blog Post](https://realpython.com/python-data-classes/) do site Real Python, assim como na [documentação oficial](https://docs.python.org/3/library/dataclasses.html) da linguagem.
# +
from dataclasses import dataclass
from typing import List
@dataclass
class Carta:
rank: str
naipe: str
@dataclass
class Baralho:
cartas: List[Carta]
baralho = Baralho([Carta('Q', 'Copa'), Carta('A', 'Espada')])
print(baralho)
# -
# ### Decorators
#
# Decorador é um design pattern em Python o qual permite que seja adicionada novas funcionalidades a um objeto existente sem modificar sua estrutura.
#
# Como funções são cidadãos de primeira classe, elas podem ser passadas como argumentos para serem executadas por outras funções (algo parecido que vimos em Closure).
#
# Neste sentido decoradores utilizam "closures" para trazer essa funcionalidade de uma forma simplificada.
#
# Vejamos um simples exemplo.
# +
def decorator(fn):
def wrapper(*args, **kwargs):
print('Before function...')
z = fn(*args, **kwargs)
print('After function...')
return z
return wrapper
def soma(x, y):
return x + y
# Without sintatic sugar
dec = decorator(soma)
print(dec(10, 2))
# -
# Na verdade, não precisamos criar toda essa estrutura complexa para nosso decorador, em Python temos uma estrutura sintática mais simples para definir um decorador em uma função.
#
# ```python
# @<nome_decorator>
# ```
#
# > P.S.: Note que usamos a sintaxe \*args e \*\*kwargs, dessa maneira seria possível passar qualquer quantidade de parâmetros para nossa função, fazendo dessa maneira com que nosso decorador fosse mais genérico!
# Examplo
@decorator
def subtracao(x, y):
return x - y
print(subtracao(10, 2))
# Para se criar uma melhor estrutura de um decorador, a melhor maneira é utilizar o pacote functools e envolver a função de wrapper com o decorador wraps. Isso permite que possamos manter a identidade de nossa função (caso ela tenha de ser inspecionada). Para isso precisamos adicionar duas novas linhas!
# +
# Este import
import functools
def dec(fn):
# Esta chamada ao decorador da functools
@functools.wraps(fn)
def wrapper(*args, **kwargs):
z = fn(*args, **kwargs)
return z
return wrapper
@dec
def mul(x, y):
return x * y
@decorator
def div(x, y):
return x / y
print('Example using functools:', mul.__name__)
print('Example NOT using functools:', div.__name__)
# -
# #### Múltiplos decoradores
#
# Podemos adicionar múltiplos decoradores, neste sentido eles serão executados na ordem em que foram apresentados. Onde um decorador chama o outro e por fim a função decorada.
#
# Por exemplo:
# +
def first(fn):
# Esta chamada ao decorador da functools
@functools.wraps(fn)
def wrapper(*args, **kwargs):
print('First decorator!')
return fn(*args, **kwargs)
return wrapper
def second(fn):
# Esta chamada ao decorador da functools
@functools.wraps(fn)
def wrapper(*args, **kwargs):
print('Second decorator!')
return fn(*args, **kwargs)
return wrapper
@second
@first
def func():
return('My function')
print(func())
# -
# #### Decoradores que recebem argumentos.
#
# Outra funcionalidade bem interessante é a de adicionar parâmetros a serem recebidos nos decoradores, isso é possível, entretanto temos de mudar nossa estrutura do decorador padrão genérico e adicionar uma nova layer a ela.
# +
def function(add):
def decorator(fn):
@functools.wraps(fn)
def wrapper(*args, **kwargs):
print(f'value of add param: {add}')
return fn(*args, **kwargs) + add
return wrapper
return decorator
@function(add=5)
def parametros(x, y):
return x + y
print('Total sum:', parametros(0, 0))
# -
# ### Referências
#
# - [The Ultimate Guide to Data Classes in Python 3.7](https://realpython.com/python-data-classes/)
# - [Primer on Python Decorators](https://realpython.com/primer-on-python-decorators/#decorating-classes)
# - [<NAME> - Practical decorators - PyCon 2019](https://youtu.be/MjHpMCIvwsY)
# - [A Curious Course on Coroutines and Concurrency](http://www.dabeaz.com/coroutines/Coroutines.pdf)
# - [Consider Coroutines to Run Many Functions Concurrently](https://effectivepython.com/2015/03/10/consider-coroutines-to-run-many-functions-concurrently/)
|
notebooks/Python201/Python201.2.ipynb
|
// -*- coding: utf-8 -*-
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .cpp
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: C++17
// language: C++17
// name: xcpp17
// ---
// # Integer logic and bit fiddling
// ## Preliminaries
//
// Let's load the mp++ runtime, include the ``integer.hpp`` header and import the user-defined literals:
// +
#pragma cling add_include_path("/srv/conda/envs/notebook/include")
#pragma cling add_library_path("/srv/conda/envs/notebook/lib")
#pragma cling load("mp++")
#include <mp++/integer.hpp>
using namespace mppp::literals;
// -
// Let's also include a few useful bits from the standard library:
#include <iostream>
// ## Logic and bit fiddling
// Multiprecision integers support the standard bitwise operators, such as the bitwise NOT operator:
~(0_z1)
~(123_z1)
// Negative integers are treated as-if they were represented using two’s complement:
~(-1_z1)
~(-124_z1)
// The bitwise OR, AND and XOR operators:
(0b01001010_z1 | 0b10010100_z1).to_string(2)
(0b11001010_z1 & 0b10010100_z1).to_string(2)
(0b11001010_z1 ^ 0b10010100_z1).to_string(2)
// Multiprecision integer arguments can be mixed with C++ integral arguments:
(0b01001010_z1 | 45).to_string(2)
(-123ll & 0b10010100_z1).to_string(2)
(255u ^ 0b10010100_z1).to_string(2)
// The in-place variants are supported as well:
{
auto n = 0b001_z1;
n |= 0b111_z1;
std::cout << n << '\n';
}
{
int n = -0b101010;
n &= 0b111_z1;
std::cout << n << '\n';
}
{
auto n = 0b001_z1;
n ^= 0b111_z1;
std::cout << n << '\n';
}
|
integer/04_bit_fiddling.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
# +
import matplotlib.pyplot as plt
# %matplotlib inline
from matplotlib import cm
import numpy as np
import shutil
# -
# ### Prepare data for TopicNet
from sklearn import datasets
from sklearn.datasets import fetch_20newsgroups
train_20 = fetch_20newsgroups(subset='train',
remove=('headers', 'footers', 'quotes'),)
test_20 = fetch_20newsgroups(subset='test',
remove=('headers', 'footers', 'quotes'),)
train_20.pop('DESCR')
labels = train_20.pop('target_names')
for k in train_20.keys():
print(len(train_20[k]), k)
test_20.pop('DESCR')
labels_test = test_20.pop('target_names')
for k in test_20.keys():
print(len(test_20[k]), k)
train_pd = pd.DataFrame(train_20).rename(columns = {'data':'raw_text'},)
#train_pd['raw_text'] = train_pd['raw_text'].apply(lambda x: x.decode('windows-1252'))
train_pd['id'] = train_pd.filenames.apply( lambda x: '.'.join(x.split('/')[-2:]).replace('.','_'))
test_pd = pd.DataFrame(test_20).rename(columns = {'data':'raw_text'})
#test_pd['raw_text'] = test_pd['raw_text'].apply(lambda x: x.decode('windows-1252'))
test_pd['id'] = test_pd.filenames.apply( lambda x: '.'.join(x.split('/')[-2:]))
# +
bad_names = [9976, 9977, 9978, 9979, 9980, 9981, 9982, 9983, 9984, 9985, 9986, 9987, 9988, 9990]
bad_names = [f"comp_os_ms-windows_misc_{i}" for i in bad_names]
bad_indices = train_pd.query("id in @bad_names").index
# -
# +
from nltk.corpus import wordnet
def nltk2wn_tag(nltk_tag):
if nltk_tag.startswith('J'):
return wordnet.ADJ
elif nltk_tag.startswith('V'):
return wordnet.VERB
elif nltk_tag.startswith('N'):
return wordnet.NOUN
elif nltk_tag.startswith('R'):
return wordnet.ADV
else:
return ''
# +
import nltk
import string
import pandas as pd
from glob import glob
from nltk.stem import WordNetLemmatizer
from nltk.corpus import stopwords
from nltk.collocations import BigramAssocMeasures, BigramCollocationFinder
from collections import Counter
import re
# +
pattern = re.compile('\S*@\S*\s?')
def vowpalize_sequence(sequence):
word_2_frequency = Counter(sequence)
del word_2_frequency['']
vw_string = ''
for word in word_2_frequency:
vw_string += word + ":" + str(word_2_frequency[word]) + ' '
return vw_string
def do_vw_for_me_please(dataframe):
bad_entries = []
tokenized_text = []
for indx, text in enumerate(dataframe['raw_text'].values):
try:
text = str(pattern.sub('', text))
except TypeError:
text=''
tokens = [tok for tok in nltk.wordpunct_tokenize(text.lower()) if len(tok) > 1]
tokenized_text.append(nltk.pos_tag(tokens))
dataframe['tokenized'] = tokenized_text
stop = set(stopwords.words('english'))
lemmatized_text = []
wnl = WordNetLemmatizer()
for text in dataframe['tokenized'].values:
lemmatized = [wnl.lemmatize(word, nltk2wn_tag(pos))
if nltk2wn_tag(pos) != ''
else wnl.lemmatize(word)
for word, pos in text ]
lemmatized = [word for word in lemmatized
if word not in stop and word.isalpha()]
lemmatized_text.append(lemmatized)
dataframe['lemmatized'] = lemmatized_text
bigram_measures = BigramAssocMeasures()
finder = BigramCollocationFinder.from_documents(dataframe['lemmatized'])
finder.apply_freq_filter(5)
set_dict = set(finder.nbest(bigram_measures.pmi,32100)[100:])
documents = dataframe['lemmatized']
bigrams = []
for doc in documents:
entry = ['_'.join([word_first, word_second])
for word_first, word_second in zip(doc[:-1],doc[1:])
if (word_first, word_second) in set_dict]
bigrams.append(entry)
dataframe['bigram'] = bigrams
vw_text = []
for index, data in dataframe.iterrows():
vw_string = ''
doc_id = data.id
lemmatized = '@lemmatized ' + vowpalize_sequence(data.lemmatized)
bigram = '@bigram ' + vowpalize_sequence(data.bigram)
vw_string = ' |'.join([doc_id, lemmatized, bigram])
vw_text.append(vw_string)
dataframe['vw_text'] = vw_text
print('num bad entries ', len(bad_entries))
print(bad_entries)
return dataframe
# -
train_pd = do_vw_for_me_please(train_pd)
display(train_pd.head())
test_pd = do_vw_for_me_please(test_pd)
display(test_pd.head())
# +
# ! mkdir 20_News_dataset
train_pd.drop(bad_indices).to_csv('20_News_dataset/train_preprocessed.csv')
test_pd.to_csv('20_News_dataset/test_preprocessed.csv')
|
topicnet/demos/20NG-PREPROCESSING.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MonaLIA Classification Analysis
#
# https://towardsdatascience.com/building-a-logistic-regression-in-python-step-by-step-becd4d56c9c8
#
# +
from __future__ import print_function
import torch
import os
import sys
import numpy as np
import pandas as pd
from sklearn import metrics
from sklearn.feature_selection import RFE
from sklearn import tree
from sklearn.linear_model import LogisticRegression
import statsmodels.api as sm
import matplotlib.pyplot as plt
import seaborn as sns
import pydotplus
from IPython.display import Image
# -
# ## Read the classification results from the IPython store of the previous script
# %store -r classified_df
print(classified_df.shape)
classified_df.info()
# #### Alternatively Read saved results from the flat file
classified_df = pd.read_csv('themes_262.tsv', sep='\t')
print(classified_df.shape)
classified_df.head()
# ## Output confusion matrix and statistics
# This code can be substituted with three lines of code but the output would be more cryptic
#
# confusion_matrix....
# classification_report...
# accuracy_score....
conf_matrix = pd.crosstab(index=classified_df.prediction, columns=classified_df.target, margins= True)
conf_matrix
conf_matrix_pct_target = pd.crosstab(index=classified_df.prediction, columns=classified_df.target)\
.div(conf_matrix.loc['All',:], axis=0) \
.dropna()\
.mul(100)\
.round(2)
conf_matrix_pct_target
# +
plt.figure(figsize = (10,7))
sns.heatmap(conf_matrix_pct_target, annot=True, cmap="YlGnBu")
# +
plt.figure(figsize = (10,7))
sns.clustermap(conf_matrix_pct_target, annot=True, cmap="YlGnBu", metric="correlation")
# +
prfs = metrics.precision_recall_fscore_support(y_true= classified_df.target,
y_pred= classified_df.prediction)
acc = metrics.accuracy_score(y_true= classified_df.target,
y_pred= classified_df.prediction)
output = pd.concat((pd.Series(conf_matrix.index[:-1]) ,
pd.Series(prfs[3].round(3)),
pd.Series(prfs[1].round(3)),
pd.Series(prfs[0].round(3)),
pd.Series(prfs[2].round(3)),
pd.Series(acc.round(3))),
axis=1 )
output.columns = ['class', 'support', 'recall', 'precision', 'f1 score', 'accuracy']
output
# -
classified_df['outcome'] = (classified_df.target == classified_df.prediction).astype(int)
classified_df.outcome.value_counts()
# ## Missclassifications
miss_classified_df = classified_df.loc[classified_df.outcome == False]
miss_classified_df.head()
miss_classified_df.groupby(miss_classified_df.target).count() \
.sort_values(by='prediction') \
.plot \
.bar(y='prediction' ,
title='Number of miss-classified images per class')
# +
#fig = plt.gcf()
#fig.set_size_inches(18.5, 5.5)
(miss_classified_df.pred_prob-miss_classified_df.target_prob).plot \
.hist(title = 'Histogram of differences between predicted and target probabilities for miss-classified images')
# -
# ## Outcome vs. continous variables
classified_df.info()
# +
fig, axes = plt.subplots(nrows=2, ncols=2)
fig.set_size_inches(18.5, 10)
classified_df.plot.scatter(y='outcome' , x='width', ax=axes[0,0])
classified_df.plot.scatter(y='outcome' , x='height', ax=axes[0,1])
classified_df['ar'] = classified_df.width/classified_df.height
classified_df.plot.scatter(y='outcome' , x='ar', ax=axes[1,0])
classified_df['inv_ar'] = classified_df.height/classified_df.width
ax = classified_df.plot.scatter(y='outcome' , x='inv_ar', ax=axes[1,1] )
ax.set_xlabel("1/ar")
# -
for col in ['width' , 'height', 'ar', 'inv_ar']:
X = np.where(classified_df[col].dtype == np.int64,
classified_df[col] // 100,
classified_df[col] * 10)
y=classified_df.outcome.values
logit_model=sm.Logit(y,X)
result=logit_model.fit()
print('=' * 78)
print(col + (' / 100' if classified_df[col].dtype == np.int64 else ' * 10'))
print('=' * 78)
print(result.summary())
print('exp(coef) = %.4f' % np.exp(result.params[0]))
print('=' * 78)
# #### Make up a continious variable that is a difference between predicted and target probabilities for missclassified images (negative) and a difference between target probability and the next class probability for correctly classified images (positive)
classified_df['prob_diff'] = np.where(classified_df.target==classified_df.prediction ,
classified_df.target_prob-classified_df.prob2,
classified_df.target_prob-classified_df.pred_prob)
# +
fig, axes = plt.subplots()
fig.set_size_inches(18.5, 5)
classified_df.prob_diff.sort_values() \
.plot(use_index=False)
axes.axhline(y=0 , color='k' , linewidth=0.5)
# +
fig, axes = plt.subplots()
fig.set_size_inches(18.5, 5)
sns.regplot(x='ar', y='prob_diff', data=classified_df)
plt.show()
X=classified_df.ar.values
y=classified_df.prob_diff.values
X1 = sm.add_constant(X)
lm_model=sm.OLS(y,X1)
result=lm_model.fit()
print(result.summary2())
#p = lm_model.fit().params
#print(p)
# -
classified_df = classified_df.drop(['ar', 'inv_ar'], axis=1)
# ## Work with Categorical variables
classified_df = classified_df.drop(['noticeArtForm', 'noticeDeno'], axis=1)
classified_df.info()
# #### Eliminate variables with missing data less than a threshold (TBD)
# +
kb_col = pd.Series(classified_df.columns)
kb_col = kb_col[kb_col.str.startswith('notice')]
kb_col = kb_col[4:]
kb_col_to_keep = []
miss_val_threshold = 0.33
for col in kb_col:
print(col)
print('Number of categories: %d' % classified_df[col].unique().size)
missing_count = classified_df[col].loc[(classified_df[col] == '')].count()
missing_count += classified_df[classified_df[col].isnull()].shape[0]
missing_values_ratio = missing_count / classified_df.shape[0]
if (missing_values_ratio < miss_val_threshold):
kb_col_to_keep.append(col)
print('Missing values: %d%%' % (missing_values_ratio * 100))
print()
kb_col_to_keep
# -
kb_col_to_keep.append('outcome')
categorical_data = classified_df[kb_col_to_keep].copy()
categorical_data.head()
categorical_data[categorical_data.isnull().any(axis=1)].head()
# #### Fill the missing values
categorical_data.replace("" , "z_missing", inplace=True)
categorical_data.fillna("z_missing", inplace=True)
categorical_data.head()
# #### Clean the technique columns a bit
categorical_data.noticeTechnique1 = categorical_data.noticeTechnique1.apply(lambda x: x.split('(')[0].strip())
#categorical_data.noticeTechnique2 = categorical_data.noticeTechnique2.apply(lambda x: x.split('(')[0].strip())
categorical_data.head()
# #### Reduce Number of categories
# +
n_rows = (categorical_data.shape[1]-1)//2 + (categorical_data.shape[1] - 1)%2
fig, axes = plt.subplots(nrows=n_rows, ncols=2)
fig.set_size_inches(18.5, 5 * n_rows)
for i , col in enumerate(kb_col_to_keep[:-1]):
subplot1 = pd.crosstab(categorical_data[col], categorical_data.outcome).plot(kind='bar', ax=axes[i//2 , int(i%2)] )
subplot1.set_title('Outcome by %s' % col)
subplot1.set_xlabel(col)
# +
categorical_data_reduced = categorical_data.copy()
cat_size_threshlod_dict = {'noticeRepresentationType': 40,
'noticePhotocredit': 15,
'noticeMuseum': 20,
'noticeTechnique1' : 30,
'noticeDenomination' : 25, }
cat_size_threshlod_dict
# +
n_rows = categorical_data.shape[1]-1
fig, axes = plt.subplots(nrows=n_rows, ncols=2)
fig.set_size_inches(18.5, 5 * n_rows)
for i , col in enumerate(kb_col_to_keep[:-1]):
pd.crosstab(categorical_data[col], categorical_data.outcome).plot(kind='bar', ax=axes[i , 0] )\
.set_title('Outcome by %s (before)' % col)
cat = categorical_data.groupby(by=col).size().sort_values(ascending=False)
categorical_data_reduced[col] = categorical_data[col].replace(cat[cat <= cat_size_threshlod_dict[col]].index.values , \
"z_other")
pd.crosstab(categorical_data_reduced[col], categorical_data_reduced.outcome).plot(kind='bar', ax=axes[i, 1] )\
.set_title('Outcome by %s (after)' % col)
# +
#for i , col in enumerate(categorical_data_reduced.columns[:-1]):
# categorical_data_reduced[col] = pd.Categorical(categorical_data_reduced[col])
#categorical_data_reduced.info()
# -
categorical_data_reduced.head()
pd.get_dummies(categorical_data_reduced, columns=['noticeTechnique1'], prefix = ['noticeTechnique1'.replace('notice', '' )]).iloc[:, offset:]
# +
offset = categorical_data_reduced.shape[1] - 1
for i , col in enumerate(categorical_data_reduced.columns[:-1]):
X= pd.get_dummies(categorical_data_reduced, columns=[col], prefix = [col.replace('notice', '' )]).iloc[:, offset:]
y=categorical_data_reduced.outcome
logit_model=sm.Logit(y,X)
result=logit_model.fit()
print('=' * 78)
print(col)
print('=' * 78)
print(result.summary())
print('exp(coef):')
print(np.exp(result.params))
print('=' * 78)
# -
# ## Feature selection
# #### Create Dummy Indicator variables
# +
categorical_data_copy = categorical_data_reduced.copy()
for i , col in enumerate(categorical_data_copy.columns[:-1]):
categorical_data_copy = pd.get_dummies(categorical_data_copy, columns=[col], prefix = [col.replace('notice', '' )])
categorical_data_copy.head()
# -
# #### Using scikit.learn Recursive Feature Elimination
# https://towardsdatascience.com/building-a-logistic-regression-in-python-step-by-step-becd4d56c9c8
# +
col_to_analize = pd.Series(categorical_data_copy.columns)[1:]
logreg = LogisticRegression()
rfe = RFE(logreg, 6)
rfe = rfe.fit(categorical_data_copy[col_to_analize], categorical_data_copy['outcome'] )
print(rfe.n_features_ )
print(rfe.support_)
print(rfe.ranking_)
print(col_to_analize[rfe.support_])
# -
# #### Build a decision tree for the selected features
# +
clf = tree.DecisionTreeClassifier(criterion = "gini", random_state = 100,
max_depth=3, min_samples_leaf=5)
X = categorical_data_copy[col_to_analize]
y = categorical_data_copy.outcome
clf.fit(X, y)
# -
# #### Decision Tree Visualization
#
# https://chrisalbon.com/machine_learning/trees_and_forests/visualize_a_decision_tree/
# +
# Create DOT data
dot_data = tree.export_graphviz(clf, out_file=None,
feature_names=categorical_data_copy.columns[1:],
filled = True,
rounded= True)
# Draw graph
graph = pydotplus.graph_from_dot_data(dot_data)
# Show graph
Image(graph.create_png())
# -
# #### Display Tree with Proprtions
# +
# Create DOT data
dot_data = tree.export_graphviz(clf, out_file=None,
feature_names=categorical_data_copy.columns[1:],
filled = True,
proportion = True,
rounded = True)
# Draw graph
graph = pydotplus.graph_from_dot_data(dot_data)
# Show graph
Image(graph.create_png())
# -
#
#
#
#
#
#
# ## Scrapbook
# #### R-style logistic regression
# +
import statsmodels.api as sm
model = sm.formula.glm("outcome ~ C(noticeTechnique2)",
family=sm.families.Binomial(), data=classified_df).fit()
print(model.summary())
# -
# #### Using Chi-square test to select variables one by one
# +
import scipy.stats as stats
p_values = []
for i, col in enumerate(categorical_data_copy.columns[1:]):
_, p , _, _ = stats.chi2_contingency(observed= pd.crosstab(categorical_data_copy[col], categorical_data_copy.outcome))
#_, p = stats.fisher_exact(pd.crosstab(classified_df_copy[col], classified_df_copy.outcome))
#print(col, round(p ,3))
p_values.append(round(p ,6))
p_s = pd.Series(p_values, index=categorical_data_copy.columns[1:])
p_s[p_s.values < 0.2].sort_values()
#plt.plot(p_values)
# -
print(metrics.classification_report(y_true= classified_df.target,
y_pred= classified_df.prediction,
target_names= test_set.classes))
miss_classified = (classified [(classified[:,0] - classified[:,1] != 0).nonzero(),:]).squeeze()
def to_class_label(idx):
return test_set.classes[idx]
# +
# JUst a reminder on what is what
what = 'chat'
tp = conf_matrix.loc[what,what]
print ('tp=%d' % tp)
fp = conf_matrix.loc[what,'All'] - tp
print ('fp=%d' % fp)
fn = conf_matrix.loc['All', what] - tp
print ('fp=%d' % fn)
recall = tp /(tp+fn)
print ('recall=%f' % recall)
prec = tp /(tp+fp)
print ('prec=%f' % prec)
fscore = 2 * prec * recall /(prec + recall)
print ('fscore=%f' % fscore)
# -
print(metrics.confusion_matrix(y_true= classified_df.target,
y_pred= classified_df.prediction))
# +
r = metrics.recall_score(y_true= classified_df.target,
y_pred= classified_df.prediction,
average= None)
p = metrics.precision_score(y_true= classified_df.target,
y_pred= classified_df.prediction,
average= None)
f = metrics.f1_score(y_true= classified_df.target,
y_pred= classified_df.prediction,
average= None)
a = metrics.accuracy_score(y_true= classified_df.target,
y_pred= classified_df.prediction)
output = pd.concat((pd.Series(test_set.classes) ,
pd.Series(r.round(3)),
pd.Series(p.round(3)),
pd.Series(f.round(3)),
pd.Series(a.round(3))),
axis=1 )
output.columns = ['class', 'recall', 'precision', 'f1 score', 'accuracy']
output
# +
prfs = metrics.precision_recall_fscore_support(y_true= classified_df.target,
y_pred= classified_df.prediction)
output = pd.concat((pd.Series(test_set.classes) ,
pd.Series(prfs[3].round(3)),
pd.Series(prfs[1].round(3)),
pd.Series(prfs[0].round(3)),
pd.Series(prfs[2].round(3)),
pd.Series(a.round(3))),
axis=1 )
output.columns = ['class', 'support', 'recall', 'precision', 'f1 score', 'accuracy']
output
# +
numberList = [1, 2, 3]
strList = ['one', 'two', 'three']
# No iterables are passed
result = zip()
#Converting itertor to list
resultList = list(result)
print(resultList)
# Two iterables are passed
result = zip(numberList, strList)
# Converting itertor to set
resultSet = set(result)
print(resultSet)
# +
np.random.seed(10)
# Sample data randomly at fixed probabilities
voter_race = np.random.choice(a= ["asian","black","hispanic","other","white"],
p = [0.05, 0.15 ,0.25, 0.05, 0.5],
size=1000)
# Sample data randomly at fixed probabilities
voter_party = np.random.choice(a= ["democrat","independent","republican"],
p = [0.4, 0.2, 0.4],
size=1000)
voters = pd.DataFrame({"race":voter_race,
"party":voter_party})
voters
voter_tab = pd.crosstab(voters.race, voters.party, margins = True)
voter_tab.columns = ["democrat","independent","republican","row_totals"]
voter_tab.index = ["asian","black","hispanic","other","white","col_totals"]
observed = voter_tab.iloc[0:5,0:3] # Get table without totals for later use
voter_tab
# -
60*186
# +
expected = np.outer(voter_tab["row_totals"][0:5],
voter_tab.loc["col_totals"][0:3]) / 1000
expected = pd.DataFrame(expected)
expected.columns = ["democrat","independent","republican"]
expected.index = ["asian","black","hispanic","other","white"]
expected
# -
pd.crosstab(classified_df_copy.artForm_peinture, classified_df_copy.outcome, margins= True)
# +
_, p , _, _ = stats.fisher_exact(observed= pd.crosstab(classified_df_copy.artForm_peinture, classified_df_copy.outcome))
p
# +
# stats.fisher_exact??
# -
classified_df.to_csv('image_classification_4000_without_birds.tsv', sep="\t")
classified_df = pd.read_csv('image_classification_4000.tsv', sep='\t')
print(classified_df.shape)
classified_df.head()
# +
#labels = classified_df.noticeArtForm.astype('category').cat.categories.tolist()
#replace_map_comp = {'noticeArtForm' : {k: v for k,v in zip(labels,list(range(1,len(labels)+1)))}}
#print(replace_map_comp['noticeArtForm'])
classified_df_copy = classified_df.copy()
#classified_df_copy.noticeArtForm.replace(replace_map_comp['noticeArtForm'], inplace=True)
#classified_df_copy.noticeArtForm = classified_df_copy.noticeArtForm.astype('category')
#classified_df_copy.noticeArtForm = classified_df_copy.noticeArtForm.cat.codes
classified_df_copy = pd.get_dummies(classified_df_copy, columns=['noticeArtForm'], prefix = ['artForm'])
classified_df_copy.head()
|
Notebooks 1.0/Pipeline/MonaLIA.STEP 5.Classification Analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import util
class SearchProblem:
"""
This class outlines the structure of a search problem, but doesn't implement
any of the methods (in object-oriented terminology: an abstract class).
You do not need to change anything in this class, ever.
"""
def getStartState(self):
"""
Returns the start state for the search problem.
"""
util.raiseNotDefined()
def isGoalState(self, state):
"""
state: Search state
Returns True if and only if the state is a valid goal state.
"""
util.raiseNotDefined()
def getSuccessors(self, state):
"""
state: Search state
For a given state, this should return a list of triples, (successor,
action, stepCost), where 'successor' is a successor to the current
state, 'action' is the action required to get there, and 'stepCost' is
the incremental cost of expanding to that successor.
"""
util.raiseNotDefined()
def getCostOfActions(self, actions):
"""
actions: A list of actions to take
This method returns the total cost of a particular sequence of actions.
The sequence must be composed of legal moves.
"""
util.raiseNotDefined()
# -
def tinyMazeSearch(problem):
"""
Returns a sequence of moves that solves tinyMaze. For any other maze, the
sequence of moves will be incorrect, so only use this for tinyMaze.
"""
from game import Directions
s = Directions.SOUTH
w = Directions.WEST
return [s, s, w, s, w, w, s, w]
def depthFirstSearch(problem):
"""
Search the deepest nodes in the search tree first.
Your search algorithm needs to return a list of actions that reaches the
goal. Make sure to implement a graph search algorithm.
To get started, you might want to try some of these simple commands to
understand the search problem that is being passed in:
print "Start:", problem.getStartState()
print "Is the start a goal?", problem.isGoalState(problem.getStartState())
print "Start's successors:", problem.getSuccessors(problem.getStartState())
"""
start_state = problem.getStartState()
start_path = []
start_cost = 0
# Taking fringe as a LIFO stack
fringe = util.Stack()
start_state_tuple = (start_state, start_path, start_cost)
fringe.push(start_state_tuple)
# Maintain a set of all the visited nodes
visited_states = set()
# Start a loop to find a possible path
while True:
# If we are out of fringes, no path exists
if fringe.isEmpty():
return []
# Removing that vertex from queue, whose neighbour will be
# visited now
next_state_tuple = fringe.pop()
next_state = next_state_tuple[0]
next_path = next_state_tuple[1]
next_cost = next_state_tuple[2]
# Check if the goal state is reached in the next node
if problem.isGoalState(next_state):
return next_path
# If the node has not been visited, add all its successors to the
# fringe stack
if not next_state in visited_states:
visited_states.add(next_state)
# Get all possible successor states
successor_state_tuples = problem.getSuccessors(next_state)
# Insert the states in the fringes
for idx in range(len(successor_state_tuples)):
successor_state_tuple = successor_state_tuples[idx]
successor_state = successor_state_tuple[0]
successor_path = successor_state_tuple[1]
successor_cost = successor_state_tuple[2]
if not successor_state in visited_states:
new_path = next_path + [successor_path]
new_tuple = (successor_state, new_path, successor_cost)
fringe.push(new_tuple)
def breadthFirstSearch(problem):
"""Search the shallowest nodes in the search tree first."""
start_state = problem.getStartState()
start_path = []
start_cost = 0
# Taking fringe as a FIFO queue
fringe = util.Queue()
start_state_tuple = (start_state, start_path, start_cost)
fringe.push(start_state_tuple)
# Maintain a set of all the visited nodes
visited_states = set()
# Start a loop to find a possible path
while True:
# If we are out of fringes, no path exists
if fringe.isEmpty():
return []
# Removing that vertex from queue, whose neighbour will be
# visited now
next_state_tuple = fringe.pop()
next_state = next_state_tuple[0]
next_path = next_state_tuple[1]
next_cost = next_state_tuple[2]
# Check if the goal state is reached in the next node
if problem.isGoalState(next_state):
return next_path
# If the node has not been visited, add all its successors to the
# fringe stack
if not next_state in visited_states:
visited_states.add(next_state)
# Get all possible successor states
successor_state_tuples = problem.getSuccessors(next_state)
# Insert the states in the fringes
for idx in range(len(successor_state_tuples)):
successor_state_tuple = successor_state_tuples[idx]
successor_state = successor_state_tuple[0]
successor_path = successor_state_tuple[1]
successor_cost = successor_state_tuple[2]
if not successor_state in visited_states:
new_path = next_path + [successor_path]
new_tuple = (successor_state, new_path, successor_cost)
fringe.push(new_tuple)
def uniformCostSearch(problem):
"""Search the node of least total cost first."""
start_state = problem.getStartState()
start_path = []
start_cost = 0
# Taking fringe as a priority queue
fringe = util.PriorityQueue()
start_state_tuple = (start_state, start_path, start_cost)
fringe.push(start_state_tuple, problem.getCostOfActions(start_path))
# Maintain a set of all the visited nodes
visited_states = set()
# Start a loop to find a possible path
while True:
# If we are out of fringes, no path exists
if fringe.isEmpty():
return []
# Removing that vertex from queue, whose neighbour will be
# visited now
next_state_tuple = fringe.pop()
next_state = next_state_tuple[0]
next_path = next_state_tuple[1]
next_cost = next_state_tuple[2]
# Check if the goal state is reached in the next node
if problem.isGoalState(next_state):
return next_path
# If the node has not been visited, add all its successors to the
# fringe stack
if not next_state in visited_states:
visited_states.add(next_state)
# Get all possible successor states
successor_state_tuples = problem.getSuccessors(next_state)
# Insert the states in the fringes
for idx in range(len(successor_state_tuples)):
successor_state_tuple = successor_state_tuples[idx]
successor_state = successor_state_tuple[0]
successor_path = successor_state_tuple[1]
successor_cost = successor_state_tuple[2]
if not successor_state in visited_states:
new_path = next_path + [successor_path]
new_cost = problem.getCostOfActions(new_path)
new_tuple = (successor_state, new_path, new_cost)
fringe.update(new_tuple, new_cost)
def nullHeuristic(state, problem=None):
"""
A heuristic function estimates the cost from the current state to the nearest
goal in the provided SearchProblem. This heuristic is trivial.
"""
return 0
def aStarSearch(problem, heuristic=nullHeuristic):
"""Search the node of least total cost first."""
start_state = problem.getStartState()
start_path = []
start_cost = 0
# Taking fringe as a priority queue
fringe = util.PriorityQueue()
start_state_tuple = (start_state, start_path, start_cost)
fringe.push(start_state_tuple, problem.getCostOfActions(start_path))
# Maintain a set of all the visited nodes
visited_states = set()
# Start a loop to find a possible path
while True:
# If we are out of fringes, no path exists
if fringe.isEmpty():
return []
# Removing that vertex from queue, whose neighbour will be
# visited now
next_state_tuple = fringe.pop()
next_state = next_state_tuple[0]
next_path = next_state_tuple[1]
next_cost = next_state_tuple[2]
# Check if the goal state is reached in the next node
if problem.isGoalState(next_state):
return next_path
# If the node has not been visited, add all its successors to the
# fringe stack
if not next_state in visited_states:
visited_states.add(next_state)
# Get all possible successor states
successor_state_tuples = problem.getSuccessors(next_state)
# Insert the states in the fringes
for idx in range(len(successor_state_tuples)):
successor_state_tuple = successor_state_tuples[idx]
successor_state = successor_state_tuple[0]
successor_path = successor_state_tuple[1]
successor_cost = successor_state_tuple[2]
if not successor_state in visited_states:
new_path = next_path + [successor_path]
new_cost = problem.getCostOfActions(new_path)
new_tuple = (successor_state, new_path, new_cost)
# Determine the heuristic cost
heuristic_cost = new_cost + heuristic(successor_state, problem)
fringe.update(new_tuple, heuristic_cost)
# Abbreviations
bfs = breadthFirstSearch
dfs = depthFirstSearch
astar = aStarSearch
ucs = uniformCostSearch
|
Untitled.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: kisaPy
# language: python
# name: kisapy
# ---
# # Paquete SMEPY
# ## Autor: <NAME>
# # Instalación
#
# Esta es la guía de uso del paquete 'smer'. Está alojado en "github.com/erikangulo/smepy"
#
# Este paquete se puede instalar fácilmente a través de la librería devtools, desde Github:
#
# También se puede instalar a través del código fuente usando la consola de R.
#
# Este paquete tiene dependencias únicamente para ciertas funciones concretas listadas a continuación. El paquete puede operar correctamente sin ellas excepto para las funciones mencionadas e indicará al usuario al ejecutar cada programa si necesita alguna dependencia.
#
# * jupyter notebook
# * pandas
# * numpy
# * seaborn
# * matplotlib
#
# Cargamos el paquete:
import smepy.dataset as ds
import smepy.modifications as md
import smepy.statistics as st
# # Uso
#
# El objetivo de este paquete es facilitar la aplicación de cálculos y estadísticos en datos a usuarios que dispongan de poca experiencia en el ámbito de la programación. También es adecuado para conseguir los resultados deseados automáticamente en pocas líneas de código sin necesidad de programar nada. Entre las funciones disponibles, se encuentran la lectura y escritura de datasets, cálculo de la varianza, curva ROC, discretización, normalización y estandarización de variables, cálculo de la correlación, cálculo de la entropía y distintos gráficos que representen los resultados.
#
# En este jupyter notebook presentaremos un tutorial de cómo usar el paquete
#
# # Gestión de Dataset
#
# En esta sección observaremos que son los Dataset y como podemos crearlos, cargarlos, guardarlos y visualizarlos.
#
# ## Creación de Dataset
#
# Los Dataset son unas clases que funcionan como tablas, donde cada fila corresponde a instancias y las columnas a las variables. Cada variable será de un tipo, ya sea numérica, lógica o de caracteres. Los dataset, además de la tabla, tendrán un nombre.
#
# Empecemos observando como se crea un Dataset.
#
# Los Dataset se deben crear con estructura de diccionario, donde las llaves son el nombre de las columnas y los valores son el contenido de las columnas. El único requisito que tiene es que ha de tener como mínimo dos filas y una columna (pues para elementos con una fila independientemente de las columnas ya podemos trabajar con un vector normal y corriente).
#
# A cada Dataset le podemos asignar un id o nombre, el cual se generará automáticamente con un número en caso de no especificarlo. Además, podemos decidir si factorizar alguna de sus columnas. De esta forma, aquellas que contengan menos valores distintos que el número indicado serán factorizadas.
# +
prueba1 = [3,3,5,8,9,5]
prueba2 = ["A", "B", "D", "R", "S", "P"]
dictPrueba = {"N" : prueba1, "L": prueba2}
dsPrueba = ds.Dataset(dictPrueba)
print(dsPrueba)
# -
# También podemos cambiarles el nombre a las columnas deseadas o directamente a todas ellas. Para cada columna deseada se indicará mediante un diccionario como llave los nombres antiguos y como valores los nombres nuevos. Para cambiar todas las columnas a la vez basta con crear un array con los nuevos nombres para las columnas
dsPrueba.nombres_columna({"N":"Num"}) #cambiar la columna deseada
print(dsPrueba)
dsPrueba.nombres_columna(["Num", "Let"]) #todas las columnas
print(dsPrueba)
# ## Lectura y guardado de Dataset
#
# Además de poder crear Dataset usando los comandos anteriores, también podemos crearlos desde un fichero. De la misma manera, podemos guardar un Dataset como fichero.
#
# El guardado de ficheros está limitado a formato csv, pero la lectura puede ser de formato csv o derivados como tsv. Para ello es necesario indicar que separador usa (por defecto ",") y el caracter usado para los números decimales (por defecto "."). De igual manera podemos asignar un nombre al dataset creado. Si el fichero dispone de encabezado, es decir, nombres asignados a cada columna como la primera instancia del fichero, se usarán como nombres del Dataset, en caso contrario, deberá indicarse con el parámetro 'encabezado' como False.
#
dsPuntos = ds.leer_datos("LecturaCSV_R.csv", nombre="Puntos", encabezado=True, sep=",", decimal='.')
print(dsPuntos)
# +
#ds.guardar_datos(dsPuntos, "Guardar.txt")
# -
# # Modificaciones del Dataset
#
# En esta sección veremos diferentes modificaciones que podemos realizar a nuestro Dataset. En estos casos no se modificarán el Dataset original y obtendremos una copia con dichas modificaciones aplicadas. La modificación que se realice aparecerá reflejada en el nombre del Dataset con las modificaciones.
# ## Estandarizar y normalizar
#
# La estandarización hace que los valores sigan una distribución normal con media 0 y desviación estandar 1, mientras que la normalización hace que los valores estén comprendidos en el rango entre 0 y 1, ambos incluídos.
#
# Se normalizarán y estandarizarán automáticamente las columnas numéricas
print(md.normalizar(dsPuntos))
print(md.estandarizar(dsPuntos))
# ## Filtrado
#
# Podemos obtener un subconjunto del Dataset con los filtros que se consideren oportunos. Para ello, deberemos indicar en base a los valores de qué columna queremos filtrar y la función de filtrado. El filtro deberá ir entre comillas simples, y si se compara con valores estos tendrán comillas dobles. Si el nombre de la columna tiene espacios deberá estar entre el caracter `
#comparamos dos columnas entre sí y obtenemos las filas que cumplan dicha condición
print(md.filtrar(dsPuntos, 'Intento < Puntuacion'))
#comparamos una columna con valor y obtenemos las filas que cumplan dicha condición
print(md.filtrar(dsPuntos, 'Participante == "Juan"'))
# ## Discretizar
#
# Mediante este proceso podemos factorizar las columnas numéricas en X tramos y sustituyendo cada elemento por el tramo al que pertenece. Esta factorización se denomina discretización.
#
# Para discretizar el Dataset, deberemos indicar una o varias columnas numéricas, cuantos puntos de corte se desean (los puntos de corte delimitan los tramos), y que algoritmo se usará para inferir los puntos de corte que nos delimiten los tramos para posteriormente discretizar. En caso de no indicar columnas, la discretización se aplicará automáticamente a todas las columnas numéricas del Dataset
#
# Uno de los algoritmos clásicos es equal width (igual anchura). Dado un vector de números reales y un número de intervalos, determina cuales son los puntos de corte para generar un vector categórico de tal manera que esos puntos están uniformemente distribuidos en el rango de los valores. Por ejemplo, si tenemos los valores (11.5, 10.2, 1.2, 0.5, 5.3, 20.5, 8.4) y queremos generar una variable categórica (su implementación en Python puede ser como un string) con cuatro posibles valores, tenemos que determinar tres puntos de corte, que serán valores que separen el intervalo (entre 0.5 y 20.5 en este caso) en 4 tramos de igual tamaño. Es decir, el primer tramo irá de 0.5 a 5.5, el segundo de 5.5 a 10.5, el tercero de 10.5 a 15.5 y el último de 15.5 a 20.5. Es decir, tendríamos 3 puntos de corte, 5.5, 10.5 y 15.5. Normalmente cuando se lleva a cabo esta tarea, ante un nuevo valor es necesario determinar a que intervalo pertence. Dado que cuando eso ocurre el valor puede estar fuera de los límites del vector original, el comienzo del primer tramos se suele considerar -infinito y el final del último como infinito. Es decir, una vez aplicado el algoritmo, el resultado sería un vector categórico de este estilo: '["I3", "I2", "I1", "I1", "I4", "I2"]', donde I1=(-infinito, 5.5], I2=(5.5, 10.5], I3=(10.5, 15.5], I4=(15.5, infinito). Otro de los algoritmos clásicos es equal frequency (igual frecuencia), donde el objetivo es buscar los puntos de corte que hagan que el número de valores del vector a discretizar que caen en cada uno de los intervalos sea el mismo (+- 1, según el número de intervalos y de puntos).
#
# Además, en caso de usar otro algoritmo de otro paquete de inferencia de puntos de corte, podemos discretizar también nuestro dataset usando los puntos de corte obtenidos.
#
# Veámos unos ejemplos.
# Discretizar en 3 tramos las columnas numéricas del Dataset a través del método 'Igual Frecuencia'
print(md.discretizar(dsPuntos))
# Discretizar en 4 tramos la segunda columna numéricas del Dataset a través del método 'Igual Anchura'
print(md.discretizar(dsPuntos, columnas=["Puntuacion"], metodo="anchura", puntos_corte=4))
# Discretizar la columna Puntuacion con los puntos de corte proporcionados
print(md.discretizar(dsPuntos, columnas=["Puntuacion"], metodo="manual", puntos_corte=[2,4,6]))
# # Cálculo de estadísticos en el Dataset
#
# En esta sección mostraremos diferentes estadísticos que podemos aplicar al Dataset y visualizaremos los resultados.
# ## Varianza y entropía
#
# Podemos usar el comando 'varianzas' para obtener la varianza correspondiente a cada columna que sea numérica.
#
st.varianzas(dsPuntos)
# Por defecto obtendremos la entropía normalizada, entre 0 y 1, el sistema se encarga en estos casos de detectar cuántos valores distintos hay. De todas formas, podemos obtenerla también sin normalizar si así quisiesemos. Por último, podemos representar las entropías a través de un gráfico también.
dfEst= {
'A':[1,2,3,4,5,6,7,8,9,10],
'B':[10, 20, 10, 10, 10, 30, 70, 80, 10, 90],
'C':['Verde', 'Amarillo', 'Azul', 'Amarillo', 'Verde', 'Rojo', 'Rojo', 'Rojo', 'Verde', 'Azul'],
'D':[0]*10
}
dsEst = ds.Dataset(dfEst)
st.entropias(dsEst, normalizar=True, plot=True)
# ## Boxplot
#
# Tener una idea de cómo son nuestros datos es interesante. Por ello, también se ofrece visualizar mediante un Boxplot cada columna del Dataset, simplemente usando la función 'graficoBoxplot'
st.graficoBoxplot(dsEst)
# ## Correlaciones e información mútua
#
# Es posible también observar cómo de relacionadas están las columnas del Dataset. Para las columnas numéricas usaremos 'correlaciones', mientras que para columnas discretas (factores y character) usaremos 'infmutuas'. Ambas funciones detectan automáticamente las columnas que son apropiadas. Podemos obtener también gráficos para analizar las relaciones.
dfEst2 = {'A': [33.33, 45.12, 84.32, 65.28, 14.67],
'B': [20.46, 78.36, 76.41, 16.81, 13.19],
'C': [8.91, 6.52, 13.35, 41.25, 30.40]
}
dsEst2 = ds.Dataset(dfEst2)
st.correlaciones(dsEst2, plot=True)
dfEst3= {
'A':["1","2","3","4","5","6","7","8","9","10"],
'B':["10", "20", "10", "10", "10", "30", "70", "80", "10", "90"],
'C':['Verde', 'Amarillo', 'Azul', 'Amarillo', 'Verde', 'Rojo', 'Rojo', 'Rojo', 'Verde', 'Azul']
}
dsEst3 = ds.Dataset(dfEst3)
st.infmutuas(dsEst3, plot=True)
|
docs/Tutorial_smpey.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Remote Backend Compiler - RBC
#
# In RBC, a function compilation process is split between the local host (client) and remote host (server).
# In client, the functions are compiled to LLVM IR string. The IR is sent to the server where it is compiled to machine code.
#
# When client calls the function, the arguments are sent to the server, the server executes the function call, and the results will be sent back to client as a response.
#
# To use RBC, import the `rbc` package and define remote JIT decorator. The remote JIT decorator has three use cases:
#
# 1. decorate Python functions that implementation will be used as a template for low-level functions
# 2. define signatures of the low-level functions
# 3. start/stop remote JIT server
import rbc
# ## Create Remote JIT decorator
rjit = rbc.RemoteJIT(host='localhost')
# One can start the server from a separate process as well as in background of the current process:
rjit.start_server(background=True)
# The server will be stopped at the end of this notebook, see below.
# ## Use `rjit` as decorator with signature specifications
#
# A function signature can be
# - in the form of a string, e.g. `"int64(int64, int64)"`,
# - or in the form of a numba function signature, e.g. `numba.int64(numba.int64, numba.int64)`,
# - or a `ctypes.CFUNCTYPE` instance, e.g. `ctypes.CFUNCTYPE(ctypes.c_int64, ctypes.c_int64, ctypes.c_int64)`.
#
# If a function uses annotations, these are also used for determining the signature of a function.
#
# For instance, the following example will define an `add` function for arguments with `int` or `float` type:
@rjit('f64(f64,f64)')
def add(a: int, b: int) -> int:
return a + b
[(device, target_info)] = rjit.targets.items()
print('\n'.join(map(str, add.get_signatures(target_info)))) # to view the currently defined signatures
# ## Try it out:
add(1, 2) # int inputs
add(1.5, 2.0) # float inputs
try: # complex inputs
add(1j, 2) # expect a failure
except Exception as msg:
print(msg)
# add support for complex inputs:
add.signature('complex128(complex128, complex128)')
add(1j, 2) # now it works
# ## Debugging
#
# For debugging, one can view the generated LLVM IR using the `add.describe` method or just printing the `add` object:
print(add)
# ## Stopping the RBC server
rjit.stop_server()
|
notebooks/rbc-simple.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (baobab)
# language: python
# name: baobab
# ---
# +
import numpy as np
import corner
import pandas as pd
import torch
from baobab.sim_utils import add_g1g2_columns
from baobab.data_augmentation.noise_lenstronomy import NoiseModelNumpy
import lenstronomy
print(lenstronomy.__path__)
import os
from baobab.data_augmentation.noise_lenstronomy import get_noise_sigma2_lenstronomy
import h0rton.tdlmc_utils as tdlmc_utils
from h0rton.configs import TrainValConfig, TestConfig
from h0rton.h0_inference import H0Posterior, plot_h0_histogram, h0_utils, plotting_utils
from h0rton.trainval_data import XYData
from torch.utils.data import DataLoader
import matplotlib.pyplot as plt
from astropy.cosmology import FlatLambdaCDM
from lenstronomy.LensModel.lens_model import LensModel
from lenstronomy.LensModel.lens_model_extensions import LensModelExtensions
from lenstronomy.LensModel.Solver.lens_equation_solver import LensEquationSolver
from lenstronomy.Cosmo.lens_cosmo import LensCosmo
from lenstronomy.Util import constants
from astropy.cosmology import FlatLambdaCDM
from lenstronomy.Plots import lens_plot
import lenstronomy.Util.util as util
import lenstronomy.Util.simulation_util as sim_util
import matplotlib.pyplot as plt
import numpy as np
from mpl_toolkits.axes_grid1 import make_axes_locatable
from lenstronomy.LensModel.lens_model_extensions import LensModelExtensions
from lenstronomy.Data.imaging_data import ImageData
from lenstronomy.Plots import plot_util
import scipy.ndimage as ndimage
from matplotlib.lines import Line2D
from matplotlib.patches import Patch
from baobab.configs import BaobabConfig
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
plt.rcParams.update(plt.rcParamsDefault)
plt.rc('font', family='STIXGeneral', size=15)
plt.rc('xtick', labelsize='medium')
plt.rc('ytick', labelsize='medium')
plt.rc('text', usetex=True)
plt.rc('axes', linewidth=2, titlesize='large', labelsize='medium')
# -
# # Curated lens examples gallery
#
# __Author:__ <NAME> (@jiwoncpark)
#
# __Created:__ 8/20/2020
#
# __Last run:__ 11/29/2020
#
# __Goals:__
# We compare the BNN-inferred, forward modeling, and precision ceiling H0 posteriors for four hand-picked lenses. The precision ceiling corresponds to the theoretical case of a perfectly known lens model. Any difference between the BNN-inferred posterior and the precision ceiling can be attributed to the lens model constraint.
#
# __Before_running:__
# 1. Train the BNN, e.g.
# ```bash
# python h0rton/train.py experiments/v2/train_val_cfg.json
# ```
#
# 2. Get inference results for the trained model and the precision ceiling, e.g.
# ```bash
# python h0rton/infer_h0_mcmc_default.py experiments/v2/mcmc_default.json
# python h0rton/infer_h0_simple_mc_truth.py experiments/v0/simple_mc_default.json
# ```
#
# 3. Summarize the inference results, e.g.
# ```bash
# python h0rton/summarize.py 2 mcmc_default
# python h0rton/summarize.py 0 mcmc_default
# ```
# Let's first read in some inference configs and truth metadata.
# +
n_test = 200
default_version_id = 3 # 1 HST orbit
truth_version_id = 0 # precision ceiling
default_version_dir = '/home/jwp/stage/sl/h0rton/experiments/v{:d}'.format(default_version_id)
truth_version_dir = '/home/jwp/stage/sl/h0rton/experiments/v{:d}'.format(truth_version_id)
default_summary = pd.read_csv(os.path.join(default_version_dir, 'summary.csv'), index_col=False).iloc[:n_test]
truth_summary = pd.read_csv(os.path.join(truth_version_dir, 'summary.csv'), index_col=False).iloc[:n_test]
true_H0 = 70.0
true_Om0 = 0.3
true_cosmo = FlatLambdaCDM(H0=true_H0, Om0=true_Om0)
# Join with metadata to get n_img
test_cfg_path = os.path.join(default_version_dir, 'mcmc_default.json')
test_cfg = TestConfig.from_file(test_cfg_path)
baobab_cfg = BaobabConfig.from_file(test_cfg.data.test_baobab_cfg_path)
test_dir = baobab_cfg.out_dir
metadata_path = os.path.join(test_dir, 'metadata.csv')
meta = pd.read_csv(metadata_path, index_col=None)
meta = add_g1g2_columns(meta)
meta['id'] = meta.index
default_summary = default_summary.merge(meta, on='id', how='inner', suffixes=['', '_y'])
truth_summary = truth_summary.merge(meta, on='id', how='inner', suffixes=['', '_y'])
# For getting noise kwargs
train_val_cfg = TrainValConfig.from_file(test_cfg.train_val_config_file_path)
# -
# We want to cover the whole range of H0 uncertainties, so select a lens from each quartile of H0 uncertainty.
pd.set_option('display.max_rows', None)
std_quantiles = np.quantile(default_summary['H0_std'].values, [0.25, 0.5, 0.75, 1])
#default_summary.sort_values('D_dt_sigma')
#default_summary.iloc[43]
#print(std_quantiles)
#np.argmin(default_summary['H0_std'].values)
example_lens_i = np.array([63, 37, 86, 43]) # IDs of four hand-picked lenses
#np.digitize(default_summary.loc[example_lens_i, 'H0_std'].values, std_quantiles, right=True)
def gaussian(x, mean, standard_deviation, amplitude):
"""Gaussian PDF"""
return amplitude * np.exp( - ((x - mean) / standard_deviation) ** 2)
# The below cell populates dictionaries associated with the BNN, forward modeling, and precision ceiling results.
# +
# Read in D_dt_samples from D_dt_dict files
from scipy import stats
n_test = 200 # number of lenses to visualize
version_id = 3 # ID of the version folder in experiments
prec_version_id = 0 # ID of the version folder corresponding to precision ceiling
true_H0 = 70.0
true_Om0 = 0.3
version_dir = '/home/jwp/stage/sl/h0rton/experiments/v{:d}'.format(version_id)
test_cfg_path = os.path.join(version_dir, 'mcmc_default.json')
test_cfg = TestConfig.from_file(test_cfg_path)
baobab_cfg = BaobabConfig.from_file(test_cfg.data.test_baobab_cfg_path)
train_val_cfg = TrainValConfig.from_file(test_cfg.train_val_config_file_path)
# Read in truth metadata
metadata = pd.read_csv(os.path.join(baobab_cfg.out_dir, 'metadata.csv'), index_col=None, nrows=n_test)
# Read in summary
summary = pd.read_csv(os.path.join(version_dir, 'summary.csv'), index_col=None, nrows=n_test)
ceiling_dir = os.path.join('/home/jwp/stage/sl/h0rton/experiments/v{:d}'.format(0), 'simple_mc_default')
samples_dir = os.path.join(version_dir, 'mcmc_default')
D_dt_dict_fnames = ['D_dt_dict_{0:04d}.npy'.format(lens_i) for lens_i in example_lens_i]
oversampling = 20
ceiling_samples_dict = {} # time delay precision ceiling
ceiling_weights_dict = {} # time delay precision ceiling
H0_samples_dict = {} # BNN-inferred H0 posterior
fm_samples_dict = {} # forward modeling H0 posterior
for i, lens_i in enumerate(example_lens_i):
truth_i = summary.iloc[lens_i]
# Populate ceiling dict
f_name_ceiling = 'h0_dict_{0:04d}.npy'.format(lens_i)
ceiling_dict = np.load(os.path.join(ceiling_dir, f_name_ceiling), allow_pickle=True).item()
ceiling_samples_dict[lens_i] = ceiling_dict['h0_samples']
ceiling_weights_dict[lens_i] = ceiling_dict['h0_weights']
# Populate BNN dict
f_name = 'D_dt_dict_{0:04d}.npy'.format(lens_i)
uncorrected_D_dt_samples = np.load(os.path.join(samples_dir, f_name), allow_pickle=True).item()['D_dt_samples'] # [old_n_samples,]
# Correct D_dt samples using k_ext
uncorrected_D_dt_samples = h0_utils.remove_outliers_from_lognormal(uncorrected_D_dt_samples, 3).reshape(-1, 1) # [n_samples, 1]
k_ext_rv = getattr(stats, test_cfg.kappa_ext_prior.dist)(**test_cfg.kappa_ext_prior.kwargs)
k_ext = k_ext_rv.rvs(size=[len(uncorrected_D_dt_samples), oversampling]) # [n_samples, oversampling]
if test_cfg.kappa_ext_prior.transformed:
D_dt_samples = (uncorrected_D_dt_samples*k_ext).flatten()
else:
D_dt_samples = (uncorrected_D_dt_samples/(1.0 - k_ext)).flatten() # [n_samples,]
# Convert D_dt into H0
cosmo_converter = h0_utils.CosmoConverter(truth_i['z_lens'], truth_i['z_src'], H0=true_H0, Om0=true_Om0)
H0_samples = cosmo_converter.get_H0(D_dt_samples)
H0_samples_dict[lens_i] = H0_samples
# Populate forward modeling dict
version_id = 2
fm_version_dir = '/home/jwp/stage/sl/h0rton/experiments/v{:d}'.format(version_id)
fm_samples_path = os.path.join(fm_version_dir, 'forward_modeling_{:d}'.format(lens_i), 'D_dt_dict_{0:04d}.npy'.format(lens_i))
fm_D_dt_samples = np.load(fm_samples_path, allow_pickle=True).item()['D_dt_samples']
fm_D_dt_samples = fm_D_dt_samples[int(fm_D_dt_samples.shape[0]*0.95):]
fm_D_dt_samples *= np.random.normal(1, 0.025, fm_D_dt_samples.shape)
fm_H0_samples = cosmo_converter.get_H0(fm_D_dt_samples)
fm_samples_dict[lens_i] = fm_H0_samples
# -
# Note that the below cells take a while (~10 min each) to run because of the caustics computation. We enable two plots: one that includes the precision ceiling (the paper version) and the other that doesn't (talk version, to minimize confusion). The paper version first:
# +
plt.close('all')
n_rows = 2
n_cols = 4
fig, axes = plt.subplots(n_rows, n_cols, figsize=(20, 8))
# H0 histograms
for col_i, lens_i in enumerate(example_lens_i):
axes[0, col_i].axvline(x=true_H0, linestyle='--', color='k', label='Truth = 70 km Mpc$^{-1}$ s$^{-1}$')
# Plot precision floor
truth_lens_info = truth_summary[truth_summary['id'] == lens_i].squeeze()
truth_H0_mean = truth_lens_info['H0_mean']
truth_H0_std = truth_lens_info['H0_std']
amp = 1.0/truth_H0_std/np.sqrt(2*np.pi)
popt = [truth_H0_mean, truth_H0_std, amp]
#truth_samples = np.random.normal(truth_H0_mean, truth_H0_std, 10000)
x_interval_for_fit = np.linspace(40, 100, 1000)
default_lens_info = default_summary[default_summary['id'] == lens_i].squeeze()
default_H0_mean = default_lens_info['H0_mean']
default_H0_std = default_lens_info['H0_std']
default_samples = np.random.normal(default_H0_mean, default_H0_std, 10000)
# Get max count
counts, bins = np.histogram(H0_samples_dict[lens_i], bins=40, range=[40, 100])
weight = np.ones_like(H0_samples_dict[lens_i])*amp/np.max(counts)
# Plot histogram of BNN samples
bins = np.linspace(40, 100, 30)
n, bins, _ = axes[0, col_i].hist(ceiling_samples_dict[lens_i],
weights=ceiling_weights_dict[lens_i],
bins=bins, alpha=1, range=[40, 100.0], edgecolor='tab:gray',
histtype='step', density=True, linewidth=2,
label='Time delay precision ceiling')
fm_counts, fm_bins = np.histogram(fm_samples_dict[lens_i], bins=bins, range=[40, 100])
fm_weight = np.max(n)/np.max(fm_counts)
_ = axes[0, col_i].hist(fm_samples_dict[lens_i],
#weights=np.ones_like(fm_samples_dict[lens_i])*fm_weight,
bins=bins, alpha=0.8, density=True, color='#8ca252', range=[40, 100.0],
edgecolor='#637939', histtype='stepfilled', linewidth=1.0,
label='Forward modeling posterior')
bnn_counts, bnn_bins = np.histogram(H0_samples_dict[lens_i], bins=bins, range=[40, 100])
bnn_weight = np.max(n)/np.max(bnn_counts)
_ = axes[0, col_i].hist(H0_samples_dict[lens_i],
#weights=np.ones_like(H0_samples_dict[lens_i])*bnn_weight,
bins=bins, alpha=0.8, density=True, color='#d6616b', range=[40, 100.0],
edgecolor='#843c39', histtype='stepfilled', linewidth=1.2,
label='BNN-inferred posterior')
# Plot forward modeling
fm_stats = h0_utils.get_normal_stats(fm_samples_dict[lens_i])
# Plot histogram of time delay precision ceiling
#axes[0, col_i].plot(x_interval_for_fit, gaussian(x_interval_for_fit, *popt), color='tab:gray', label='Time delay precision ceiling', lw=4)
axes[0, col_i].set_xticks(np.arange(40, 100 + 5, 10))
axes[0, col_i].set_xticks(np.arange(40, 100 + 1, 1), minor=True)
axes[0, col_i].set_yticks([])
axes[0, col_i].set_xlabel('$H_0$ (km Mpc$^{-1}$ s$^{-1}$)', fontsize=20)
subplot_legend_elements = [
Patch(facecolor='#d6616b', edgecolor='#843c39', alpha=0.8, label='{:0.1f} $\pm$ {:0.1f}'.format(default_H0_mean, default_H0_std)),
Patch(facecolor='#8ca252', edgecolor='#637939', alpha=0.8, label='{:0.1f} $\pm$ {:0.1f}'.format(fm_stats['mean'], fm_stats['std'])),
Patch(facecolor='white', edgecolor='tab:gray', linewidth=2, alpha=0.75, label='{:0.1f} $\pm$ {:0.1f}'.format(truth_H0_mean, truth_H0_std)),]
subplot_legend = axes[0, col_i].legend(handles=subplot_legend_elements, loc=[0.68, 0.68], framealpha=1.0, fontsize=20)
axes[0, col_i].add_artist(subplot_legend)
global_legend = axes[0, 0].legend(bbox_to_anchor=(0.03, 1.23, n_cols + 1.15, 0.102), loc='upper center', ncol=4, mode="expand", borderaxespad=-0.5, fontsize=20, frameon=False)
axes[0, 0].add_artist(global_legend)
axes[0, 0].set_ylabel('Density', fontsize=25)
bp = baobab_cfg.survey_info.bandpass_list[0]
exposure_time_factor = np.ones([1, 1, 1])
survey_object = baobab_cfg.survey_object_dict[bp]
# Dictionary of SingleBand kwargs
noise_kwargs = survey_object.kwargs_single_band()
# Factor of effective exptime relative to exptime of the noiseless images
exposure_time_factor[0, :, :] = train_val_cfg.data.eff_exposure_time[bp]/noise_kwargs['exposure_time']
noise_kwargs.update(exposure_time=train_val_cfg.data.eff_exposure_time[bp])
# Dictionary of noise models
noise_model = NoiseModelNumpy(**noise_kwargs)
# Noised images
for col_i, lens_i in enumerate(example_lens_i):
lens_info = default_summary[default_summary['id'] == lens_i].squeeze()
img = np.load(os.path.join(test_dir, 'X_{0:07d}.npy'.format(int(lens_i))))
# Add noise
img *= exposure_time_factor
#noise_map = noise_model.get_noise_map(img)
#img += noise_map
img = np.squeeze(img)
# Transform
img = np.log1p(img)
# Overlay caustic, critical curves
lens_model = LensModel(lens_model_list=['PEMD', 'SHEAR'], cosmo=true_cosmo, z_lens=lens_info['z_lens'], z_source=lens_info['z_src'])
kwargs_lens = [{'theta_E': lens_info['lens_mass_theta_E'], 'gamma': lens_info['lens_mass_gamma'], 'center_x': lens_info['lens_mass_center_x'], 'center_y': lens_info['lens_mass_center_y'], 'e1': lens_info['lens_mass_e1'], 'e2': lens_info['lens_mass_e2']}, {'gamma1': lens_info['external_shear_gamma1'], 'gamma2': lens_info['external_shear_gamma2']}]
x_source = lens_info['src_light_center_x']
y_source = lens_info['src_light_center_y']
plotting_utils.lens_model_plot_custom(img, axes[1, col_i], lensModel=lens_model, kwargs_lens=kwargs_lens, sourcePos_x=x_source, sourcePos_y=y_source, point_source=True, with_caustics=True, deltaPix=0.08, numPix=64)
axes[1, col_i].axis('off')
plt.subplots_adjust(wspace=0.4, hspace=0.3)
#fig.savefig('../curated_gallery.png', bbox_inches='tight', pad_inches=0)
plt.show()
# -
fig.savefig('../plots/curated_gallery.png', bbox_inches='tight', pad_inches=0)
# And now the talk version:
# +
plt.close('all')
n_rows = 2
n_cols = 4
fig, axes = plt.subplots(n_rows, n_cols, figsize=(20, 8))
# H0 histograms
for col_i, lens_i in enumerate(example_lens_i):
axes[0, col_i].axvline(x=true_H0, linestyle='--', color='k', label='Truth = 70 km Mpc$^{-1}$ s$^{-1}$')
# Plot precision floor
truth_lens_info = truth_summary[truth_summary['id'] == lens_i].squeeze()
truth_H0_mean = truth_lens_info['H0_mean']
truth_H0_std = truth_lens_info['H0_std']
amp = 1.0/truth_H0_std/np.sqrt(2*np.pi)
popt = [truth_H0_mean, truth_H0_std, amp]
#truth_samples = np.random.normal(truth_H0_mean, truth_H0_std, 10000)
x_interval_for_fit = np.linspace(40, 100, 1000)
default_lens_info = default_summary[default_summary['id'] == lens_i].squeeze()
default_H0_mean = default_lens_info['H0_mean']
default_H0_std = default_lens_info['H0_std']
default_samples = np.random.normal(default_H0_mean, default_H0_std, 10000)
# Get max count
counts, bins = np.histogram(H0_samples_dict[lens_i], bins=40, range=[40, 100])
weight = np.ones_like(H0_samples_dict[lens_i])*amp/np.max(counts)
# Plot histogram of BNN samples
bins = np.linspace(40, 100, 30)
#n, bins, _ = axes[0, col_i].hist(ceiling_samples_dict[lens_i],
# weights=ceiling_weights_dict[lens_i],
# bins=bins, alpha=1, range=[40, 100.0], edgecolor='tab:gray',
# histtype='step', density=True, linewidth=2,
# label='Time delay precision ceiling')
#fm_counts, fm_bins = np.histogram(fm_samples_dict[lens_i], bins=bins, range=[40, 100])
#fm_weight = np.max(n)/np.max(fm_counts)
_ = axes[0, col_i].hist(fm_samples_dict[lens_i],
#weights=np.ones_like(fm_samples_dict[lens_i])*fm_weight,
bins=bins, alpha=0.8, density=True, color='#8ca252', range=[40, 100.0],
edgecolor='#637939', histtype='stepfilled', linewidth=1.0,
label='Forward modeling posterior')
#bnn_counts, bnn_bins = np.histogram(H0_samples_dict[lens_i], bins=bins, range=[40, 100])
#bnn_weight = np.max(n)/np.max(bnn_counts)
_ = axes[0, col_i].hist(H0_samples_dict[lens_i],
#weights=np.ones_like(H0_samples_dict[lens_i])*bnn_weight,
bins=bins, alpha=0.8, density=True, color='#d6616b', range=[40, 100.0],
edgecolor='#843c39', histtype='stepfilled', linewidth=1.2,
label='BNN-inferred posterior')
# Plot forward modeling
fm_stats = h0_utils.get_normal_stats(fm_samples_dict[lens_i])
# Plot histogram of time delay precision ceiling
#axes[0, col_i].plot(x_interval_for_fit, gaussian(x_interval_for_fit, *popt), color='tab:gray', label='Time delay precision ceiling', lw=4)
axes[0, col_i].set_xticks(np.arange(40, 100 + 5, 10))
axes[0, col_i].set_xticks(np.arange(40, 100 + 1, 1), minor=True)
axes[0, col_i].set_yticks([])
axes[0, col_i].set_xlabel('$H_0$ (km Mpc$^{-1}$ s$^{-1}$)', fontsize=20)
subplot_legend_elements = [
Patch(facecolor='#d6616b', edgecolor='#843c39', alpha=0.8, label='{:0.1f} $\pm$ {:0.1f}'.format(default_H0_mean, default_H0_std)),
Patch(facecolor='#8ca252', edgecolor='#637939', alpha=0.8, label='{:0.1f} $\pm$ {:0.1f}'.format(fm_stats['mean'], fm_stats['std'])),]
#Patch(facecolor='white', edgecolor='tab:gray', linewidth=2, alpha=0.75, label='{:0.1f} $\pm$ {:0.1f}'.format(truth_H0_mean, truth_H0_std)),]
subplot_legend = axes[0, col_i].legend(handles=subplot_legend_elements, loc=[0.68, 0.68], framealpha=1.0, fontsize=20)
axes[0, col_i].add_artist(subplot_legend)
global_legend = axes[0, 0].legend(bbox_to_anchor=(0.03, 1.23, n_cols + 1.15, 0.102), loc='upper center', ncol=4, mode="expand", borderaxespad=-0.5, fontsize=20, frameon=False)
axes[0, 0].add_artist(global_legend)
axes[0, 0].set_ylabel('Density', fontsize=25)
bp = baobab_cfg.survey_info.bandpass_list[0]
exposure_time_factor = np.ones([1, 1, 1])
survey_object = baobab_cfg.survey_object_dict[bp]
# Dictionary of SingleBand kwargs
noise_kwargs = survey_object.kwargs_single_band()
# Factor of effective exptime relative to exptime of the noiseless images
exposure_time_factor[0, :, :] = train_val_cfg.data.eff_exposure_time[bp]/noise_kwargs['exposure_time']
noise_kwargs.update(exposure_time=train_val_cfg.data.eff_exposure_time[bp])
# Dictionary of noise models
noise_model = NoiseModelNumpy(**noise_kwargs)
# Noised images
for col_i, lens_i in enumerate(example_lens_i):
lens_info = default_summary[default_summary['id'] == lens_i].squeeze()
img = np.load(os.path.join(test_dir, 'X_{0:07d}.npy'.format(int(lens_i))))
# Add noise
img *= exposure_time_factor
#noise_map = noise_model.get_noise_map(img)
#img += noise_map
img = np.squeeze(img)
# Transform
img = np.log1p(img)
# Overlay caustic, critical curves
lens_model = LensModel(lens_model_list=['PEMD', 'SHEAR'], cosmo=true_cosmo, z_lens=lens_info['z_lens'], z_source=lens_info['z_src'])
kwargs_lens = [{'theta_E': lens_info['lens_mass_theta_E'], 'gamma': lens_info['lens_mass_gamma'], 'center_x': lens_info['lens_mass_center_x'], 'center_y': lens_info['lens_mass_center_y'], 'e1': lens_info['lens_mass_e1'], 'e2': lens_info['lens_mass_e2']}, {'gamma1': lens_info['external_shear_gamma1'], 'gamma2': lens_info['external_shear_gamma2']}]
x_source = lens_info['src_light_center_x']
y_source = lens_info['src_light_center_y']
#plotting_utils.lens_model_plot_custom(img, axes[1, col_i], lensModel=lens_model, kwargs_lens=kwargs_lens, sourcePos_x=x_source, sourcePos_y=y_source, point_source=True, with_caustics=True, deltaPix=0.08, numPix=64)
axes[1, col_i].axis('off')
plt.subplots_adjust(wspace=0.4, hspace=0.3)
#fig.savefig('../curated_cwp.png', dpi=100)
plt.show()
|
demo/[Paper]_Make_Curated_Lens_Examples_Gallery.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import requests, urlparse, os, re, urllib
# import sys
# # !{sys.executable} -m pip install asyncio
def get_file_name(url):
a = urlparse.urlparse(url)
return os.path.basename(a.path)
def normalize_url(url):
if not (url.startswith("http://") or url.startswith("https://")):
return 'http://' + url
return url
normalize_url("www.digimouth.com/news/media/2011/09/google-logo.jpg")
get_file_name("http://www.digimouth.com/news/media/2011/09/google-logo.jpg")
get_file_name("www.digimouth.com/news/media/2011/09/google-logo.jpg")
def validate_url(url):
regex = re.compile(
r'^(?:http|ftp)s?://' # http:// or https://
r'(?:(?:[A-Z0-9](?:[A-Z0-9-]{0,61}[A-Z0-9])?\.)+(?:[A-Z]{2,6}\.?|[A-Z0-9-]{2,}\.?)|' # domain...
r'localhost|' # localhost...
r'\d{1,3}\.\d{1,3}\.\d{1,3}\.\d{1,3}|' # ...or ipv4
r'\[?[A-F0-9]*:[A-F0-9:]+\]?)' # ...or ipv6
r'(?::\d+)?' # optional port
r'(?:/?|[/?]\S+)$', re.IGNORECASE)
return re.match(regex, url) is not None
validate_url("http://www.digimouth.com/news/media/2011/09/google-logo.jpg")
validate_url("www.digimouth.com/news/media/2011/09/google-logo.jpg")
normalize_url(url)
def download_image(url):
if not validate_url(normalize_url(url)):
print("Url '{0}' is not valid. Skipping...".format(url))
return
file_name = get_file_name(url)
print "Downloading: {0}...".format(file_name),
urllib.urlretrieve(url, file_name)
print("Done.")
download_image("http://site.meishij.net/r/58/25/3568808/a3568808_142682562777944.jpg")
download_image("https://steptosea.com/wp-content/uploads/2018/03/realty/1749/432d4a9ae31b30e5fab92c84db17fce1.JPG")
|
Downloading/Download_images.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # STOCK PRICE ANALYZER with TWITTER REACTION
#
# ### 2019-01-11 by <NAME>
#
# > Python library를 이용해 주식을 예측해봅니다
#
# - 필요한 module들을 다운받습니다
# - numpy, sklearn, matplotlib
# - Support Vector Machine을 이용해 주가를 예측해봅시다
#
# > 모델은 3가지를 사용합니다
#
# - 선형모델
# - 다항모델
# - RBF모델(Radial Bias Fuction)
#
# #### RBF의 특징은 다음과 같습니다
# - 은닉층이 1개이다
# - 유클리디안 거리를 사용한다
# - 역전파 알고리즘
# - 안정성 판별이 가능하다
# +
# %matplotlib inline
import csv
import numpy as np
from sklearn.svm import SVR
import matplotlib.pyplot as plt
dates=[]
prices=[]
# 데이터를 csv로부터 받아와 저장합니다
# dates에는 날짜가, prices에는 open 가격을 넣어줍니다
def get_data(filename):
with open(filename, 'r') as csvfile:
csvFileReader = csv.reader(csvfile)
next(csvFileReader)
for row in csvFileReader:
dates.append(int(row[0].split('-')[0]))
prices.append(float(row[1]))
return
# 주식을 예상해봅니다
def predict_prices(dates, prices, x):
dates = np.reshape(dates, (len(dates), 1))
# 선형
svr_lin = SVR(kernel= 'linear', C=1e3)
# 다항
svr_poly = SVR(kernel = 'poly', C=1e3, degree = 2)
# RBF
svr_rbf = SVR(kernel='rbf', C=1e3, gamma='auto')
# 데이터 훈련
svr_lin.fit(dates, prices)
svr_poly.fit(dates, prices)
svr_rbf.fit(dates, prices)
# 데이터 표현
plt.scatter(dates, prices, color='black', label='Data')
plt.plot(dates, svr_rbf.predict(dates), color='red', label='RBF model')
plt.plot(dates, svr_lin.predict(dates), color='green', label='Linear model')
plt.plot(dates, svr_poly.predict(dates), color='blue', label='Polynomial model')
# 그래프 레이블링
plt.xlabel('Date')
plt.ylabel('Price')
plt.title('Support Vector Regression')
plt.legend()
plt.show()
#훈련된 데이터 반환
return svr_rbf.predict(x)[0], svr_lin.predict(x)[0], svr_poly.predict(x)[0]
# CSV 파일 지정
get_data('snap.csv')
# 데이터 훈련
predicted_price = predict_prices(dates, prices, 21)
# 데이터 표시
print(predicted_price)
# +
import pandas as pd
#pandas의 read_csv를 사용하면
#다음과 같이 data를 불러올 수 있습니다
snap_data = pd.read_csv('snap.csv')
snap_data
# -
# # 더 쉽게 데이터를 받아오는 방법
# # quandl 모듈을 사용해 봅니다
#
# - start,end로 날짜를 지정하고
# - quandl.get으로 주식 데이터를 불러옵니다
# - 맨먼저 애플의 정보를 불러와봅니다
# - AAPL은 2018년 4월부터 주식정보가 없는데 이유가 뭘까요 .. ?
#
# ### 하루에 50번 넘게 불러오면 차단됩니다 ! 유의하세요
# - api를 등록하면 무제한으로 받아올 수 있습니다
# +
import pandas as pd
import quandl
import datetime
#시작 날짜와 마지막 날짜를 지정해줍니다
#마지막 날짜는 현재 컴퓨터 시간입니다
start = datetime.datetime(2018,3,1)
end = datetime.date.today()
s = "AAPL"
apple = quandl.get("WIKI/" + s, start_date=start, end_date=end)
# +
# apple 데이터의 타입을 확인해봅시다
type(apple)
#데이터를 읽고 정제하기 쉽게 DataFrame으로 불러오는것을 알수있습니다
#맨앞 몇개의 데이터만 살펴봅시다
apple.head()
# -
# ## 데이터에서 중요한 지표들
#
# - Open : 시장이 열리고 난 후 가격
# - High : 최고가
# - Low : 최저가
# - Close : 장이 마감한 후 가격
# - Volume : 주식 거래 량
# - Adj가 붙은 것은 조정된 가격을 나타냅니다
# +
#plot 차트를 한번 그려봅시다
# %pylab inline
pylab.rcParams['figure.figsize'] = (10,5)
apple['Open'].plot(grid=True)
# -
# # 나스닥 상위 20개 기업들의 트윗을 분석해 봅시다
#
# > https://www.nasdaq.com/screening/companies-by-industry.aspx?sortname=marketcap&sorttype=1&exchange=NASDAQ
#
# #### 상위 20개 기업들의 트윗을 감정분석
#
# - TextBlob을 이용해 polarity와 subjectivity를 분석해봅니다
# - polarity와 subjectivity가 어제,오늘 주식에 영향을 주는지 알아봅니다
# +
from textblob import TextBlob
import nltk
import tweepy
import config
from textblob import TextBlob
nltk.download('punkt')
nltk.download('averaged_perceptron_tagger')
# 커스터머키
consumer_key = config.consumer_key
consumer_secret = config.consumer_secret
# 엑세스토큰
access_token = config.access_token
access_token_secret = config.access_token_secret
auth = tweepy.OAuthHandler(consumer_key, consumer_secret)
auth.set_access_token(access_token, access_token_secret)
api = tweepy.API(auth)
# -
# # 긍정, 부정, 주관적 요소들을 분석해봅니다
#
# - polarity > 0일경우 긍정적, 0보다 작으면 부정적 반응
# - polarity=0인 경우는 기사일 확률이 높습니다
# - 트윗 반응이 긍정적인 기업과 부정적인 기업으로 나눠서 분석해봅시다
# - 주관적 반응도 함께 알아봅니다
# +
#나스닥 상위 20개 기업
nasdaq_20 = ['Amazon', 'Microsoft', 'Alphabet', 'Apple', 'Facebook', 'Intel', 'Cisco',
'Comcast', 'Pepsico', 'Netflix', 'Amgen', 'Adobe', 'PayPal', 'Broadcom',
'Costco', 'Texas Instruments', 'Twenty-First Century', 'Gilead Sciences',
'NVIDIA', 'GILD']
#기업들의 트윗분석을 넣어줄 리스트
nasdaq_20_polarity_positive = []
nasdaq_20_polarity_negative = []
nasdaq_20_subjectivity = []
#20개 나스닥의 긍정, 부정, 주관을 알아봅니다
for company in nasdaq_20:
company_tweet = api.search(company, len='en', count=100)
polarity_score = 0
subjectivity_score = 0
for tweet in company_tweet:
company = TextBlob(tweet.text)
polarity_score += company.sentiment.polarity
subjectivity_score += company.sentiment.subjectivity
#긍정적일 경우 nasdaq_20_polarity_positive에 추가해주고
#부정적일 경우 nasdaq_20_polarity_negative에 추가해줍니다
if polarity_score > 0:
nasdaq_20_polarity_positive.append(1)
nasdaq_20_polarity_negative.append(0)
else:
nasdaq_20_polarity_positive.append(0)
nasdaq_20_polarity_negative.append(1)
#주관을 나타내는 subjectivity는 그냥 모두 더해서 넣어줍니다
nasdaq_20_subjectivity.append(subjectivity_score)
# -
#DataFrame을 만들어 기업이름, 긍정&부정, 주관 데이터를 저장합니다
df_stock = pd.DataFrame({'company' : nasdaq_20,
'company_polarity_positive' : nasdaq_20_polarity_positive,
'company_polarity_negative' : nasdaq_20_polarity_negative,
'company_subjectivity' : nasdaq_20_subjectivity})
df_stock
# # 어제 주식과 오늘 주식을 비교해봅니다
#
# - 등락 여부를 나타내는 함수를 만듭니다
# - 어제의 Open, High, Low, Close를 모두 더해
# - 오늘과 비교해 봅시다
#
# > 오늘 주식 또한 Open, High, Low, Close를 모두 더합니다
#
#
# +
trend_data = []
#등락여부를 나타내는 함수
def stock_change(stock_data):
prev_price = 0
today_price = 0
#어제의 Open부터 Close까지 더합니다
for i in range(4):
prev_price += stock_data.iloc[0,i]
#오늘의 Open부터 Close까지 더합니다
for i in range(4):
today_price += stock_data.iloc[1,i]
#어제보다 올랐으면 1, 내렸으면 0을 반환합니다
if today_price > prev_price :
trend_data.append(1)
else:
trend_data.append(0)
# +
start_date = datetime.datetime(2018,1,10)
end_date = datetime.datetime(2018,1,11)
#주식을 불러오기 위해선 코드명이 필요합니다
#코드명을 넣어줍니다
#나중에는 크롤링을 이용해서 불러올수 있게 해보겠습니다
nasdaq_20_code = ['AMZN', 'MSFT', 'GOOGL', 'GOOG', 'AAPL', 'FB', 'INTC', 'CSCO', 'CMCSA', 'PEP', 'NFLX',
'AMGN', 'ADBE', 'PYPL', 'AVGO', 'TXN', 'COST', 'FOX', 'NVDA', 'GILD']
for i in range(12,20):
code = nasdaq_20_code[i]
stock_data = quandl.get("WIKI/" + code, start_date=start_date, end_date=end_date)
stock_change(stock_data)
# +
#DataFrame에 trend column을 만들고 분석한 trend를 넣어줍니다
df_stock.insert(4, 'trend', trend_data)
#trend 데이터를 살펴봅니다
#전날보다 증가한 기업이 16개, 전날보다 감소한 기업이 4개 있습니다
df_stock['trend'].value_counts()
# -
# # 트위터가 주가에 영향을 주는지 분석해봅니다
#
# - 히트맵을 그려서 상관관계를 알아봅시다
# - seaborn의 heatmap을 사용합니다
#
# > 긍정적,부정적영향과 subjectivity도 알아봅시다
# +
import seaborn as sns
heatmap_data = df_stock[['company_polarity_positive', 'company_polarity_negative', 'company_subjectivity','trend']]
colormap = plt.cm.RdBu
plt.figure(figsize=(18,10))
plt.title('Correlation Between features', size=15, y=1.1)
sns.heatmap(heatmap_data.astype(float).corr(), linewidths=0.1, vmax=1.0,
square=True, cmap=colormap, linecolor='white', annot=True, annot_kws={"size": 16})
# -
# # 히트맵을 살펴봅시다
#
# - trend와 company_polarity_positive는 상관관계가 -0.17로
# - 트윗반응이 긍정적일수록 주가가 내려갑니다 !?
# <br><br>
# - trend와 company_polarity_negative는 상관관계가 0.17로
# - 트윗반응이 부정적일수록 호재입니다
# <br>
#
# >주관적인 반응은 주가에 영향을 거의 미치지 않는것을 알 수 있습니다
|
StockPriceAnalyzer_SiHyung.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from sklearn.pipeline import Pipeline
import pandas as pd
import numpy as np
import seaborn as sns
import os
from collections import Counter
from matplotlib import pyplot as plt
import numpy as np
from discover_feature_relationships import discover
from common import *
# -
# watermark is optional - it shows the versions of installed libraries
# so it is useful to confirm your library versions when you submit bug reports to projects
# install watermark using
# # %install_ext https://raw.githubusercontent.com/rasbt/watermark/master/watermark.py
# %load_ext watermark
# show a watermark for this environment
# %watermark -d -m -v -p numpy,matplotlib,sklearn -g
train = pd.read_csv('aps_failure_training_set.csv',skiprows=20)
data = pre_processing(train,np.nan)
# +
from sklearn.neural_network import BernoulliRBM
from sklearn.pipeline import Pipeline
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import StandardScaler, MinMaxScaler, RobustScaler
from sklearn.feature_selection import SelectKBest
from sklearn.feature_selection import f_regression
from sklearn import svm
from sklearn.model_selection import cross_val_score, cross_validate
from sklearn.metrics import precision_score, recall_score
from sklearn.metrics import make_scorer
from sklearn import linear_model
from imblearn.over_sampling import SMOTE # or: import RandomOverSampler
from imblearn.pipeline import Pipeline as imbPipeline
# -
RBM_2 = BernoulliRBM(n_components=2)
RBM_15 = BernoulliRBM(n_components=15)
imp = SimpleImputer(missing_values=np.nan, strategy='median')
anova_filter = SelectKBest(f_regression, k=15)
scaler = StandardScaler()
clf = svm.SVC(kernel='linear')
logistic = linear_model.LogisticRegression(solver='lbfgs', max_iter=10000)
smote = SMOTE(random_state=444)
# # Pipe Lines
#
# To check the use of RBM, let's create 3 pipelines with similar steps:
# - Imputer (median value)
# - Scaler (Standard Scaler)
# - SMOTE
# - RBM or Feature selection using ANOVA
# - Logistic regression or SVM
# +
rbm_features_15_svm = imbPipeline(steps=[("imputer",imp),("scale",scaler),("smote",smote),('rbm', RBM_15),("svm",clf) ])
rbm_features_15_logit = imbPipeline(steps=[("imputer",imp),("scale",scaler),("smote",smote),('rbm', RBM_15),("logit",logistic) ])
anova_features_15 = Pipeline(steps=[("imputer",imp),("scale",scaler),('anova', anova_filter),("svm",clf) ])
anova_features_smote_15 = imbPipeline(steps=[("imputer",imp),("scale",scaler),("smote",smote),('anova', anova_filter),("svm",clf) ])
# -
X = data.drop("class",axis=1).values
y = data["class"].values
scoring = {'precison': make_scorer(precision_score),'recall': make_scorer(recall_score)}
scores = {}
scores["SVM"]=cross_validate(rbm_features_15_svm,X, y, cv=3, scoring=scoring)
scores["Logit"]=cross_validate(rbm_features_15_logit,X, y, cv=3, scoring=scoring)
scores["Anova_SVM"]=cross_validate(anova_features_15,X, y, cv=3, scoring=scoring)
scores["SMOTE_Anova_SVM"]= cross_validate(anova_features_smote_15,X, y, cv=3, scoring=scoring)
for k in scores.keys():
print(f"{k} :")
print("%15s => %02d%% +/- %02d%%"%("precision",np.mean(scores[k]['test_precison'])*100,np.std(scores[k]['test_precison'])*100*2))
print("%15s => %02d%% +/- %02d%%"%("recall",np.mean(scores[k]['test_recall'])*100,np.std(scores[k]['test_recall'])*100*2))
# For those using SMOTE allows a significant improvement in precision/recall (it does not actually converge for the pipeline using RBM without SMOTE).
#
# It will be interesting to look at the output of the RBM.
# # RBM ouput:
# +
rbm_output_smote = imbPipeline(steps=[("imputer",imp),("scale",scaler),("smote",smote),('rbm', RBM_15)])
rbm_output_no_smote = Pipeline(steps=[("imputer",imp),("scale",scaler),('rbm', RBM_15)])
# -
rbm_features = rbm_output_no_smote.fit_transform(X)
rbm_features_df = pd.DataFrame(rbm_features)
print(rbm_features_df.drop_duplicates().shape)
rbm_features_df.sample(10)
# # Understanding the output
#
# There seems to be a lot of duplicates, let's count the instance.
#
# For that we will calculate a unique column, as if each row was an encoding of a number in base 2.
categories=rbm_features_df.apply(lambda r:sum( x*(2**i) for i,x in enumerate(r)),axis=1)
categories_df = categories.to_frame().rename(columns={0:"categories"})
categories_df["Class"] = data["class"]
# It turns out that not all feature are 1 or 0, in rare case there are some float values (see below).
#
# To understand how the different values are distributed, we will just consider rounding then to make the output easier to read.
categories_df[categories_df.categories < np.round(categories_df.categories)]
pd.crosstab(np.round(categories_df.categories),categories_df.Class)
|
scania/20190208 RBM feature generation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import torch
from torch.autograd import Variable
from torch.optim import Adam
from torch import nn
import torch.nn.functional as F
from torchvision.datasets import mnist
from torchvision import transforms
from torch.utils.data import DataLoader
# +
input = torch.Tensor([[[[0.1, 0.15, 0.2, 0.25],
[0.3, 0.35, 0.4, 0.45],
[0.5, 0.55, 0.6, 0.65],
[0.7, 0.75, 0.8, 0.85]]]])
data = input[0][0]
weights_tensor = torch.Tensor([[[[0.5, 0.5, 0.5],
[0.5, 0.5, 0.5],
[0.5, 0.5, 0.5]]]])
weights = weights_tensor[0][0]
# -
conv = nn.Conv2d(in_channels=1, out_channels=1, kernel_size=3, bias=False)
conv._parameters['weight'] = weights_tensor
conv(input)
# manually calculate the convolution
# iterate over matrix (x, y direction), take submatrices with dimensions kernel_size
# => this gives us 4 submatrices
# matrix muliplicate each submatrix with the convolution weights and sum up the result
torch.Tensor([[sum((data[0:3, 0:3] * weights).flatten()), sum((data[0:3, 1:4] * weights).flatten())],
[sum((data[1:4, 0:3] * weights).flatten()), sum((data[1:4, 1:4] * weights).flatten())]])
# two output channels generate two sets of weights
conv = nn.Conv2d(in_channels=1, out_channels=2, kernel_size=3, bias=False)
conv(input)
weights = list(conv.parameters())[0]
torch.Tensor([
[[sum((data[0:3, 0:3] * weights[0]).flatten()), sum((data[0:3, 1:4] * weights[0]).flatten())],
[sum((data[1:4, 0:3] * weights[0]).flatten()), sum((data[1:4, 1:4] * weights[0]).flatten())]],
[[sum((data[0:3, 0:3] * weights[1]).flatten()), sum((data[0:3, 1:4] * weights[1]).flatten())],
[sum((data[1:4, 0:3] * weights[1]).flatten()), sum((data[1:4, 1:4] * weights[1]).flatten())]]
])
|
notebooks/convolution.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Mitigating Disparities in Ranking from Binary Data
# _**An example based on the Law School Admissions Council's National Longitudinal Bar Passage Study**_
#
#
# ## Contents
#
# 1. [What is Covered](#What-is-Covered)
# 1. [Overview](#Overview)
# 1. [Data](#Data)
# 1. [Unmitigated Predictor](#Unmitigated-Predictor)
# 1. [Mitigating Demographic Disparity with Grid Search](#Mitigating-Demographic-Disparity-with-Grid-Search)
# 1. [Comparing Probabilistic Predictors using the Dashboard](#Comparing-Probabilistic-Predictors-using-the-Dashboard)
# 1. [Obtaining Low-Disparity Classifiers](#Obtaining-Low-Disparity-Classifiers)
# 1. [Postprocessing](#Postprocessing)
# 1. [Exponentiated Gradient](#Exponentiated-Gradient)
# 1. [Comparing Classifiers using the Dashboard](#Comparing-Classifiers-using-the-Dashboard)
#
#
# ## What is Covered
#
# * **Domain:**
# * Education (law-school admissions). Please review the usage notes at the end of [Overview](#Overview).
#
# * **ML tasks:**
# * Prediction of the probability of success in a bar-passage exam based on binary classification data.
# * Binary classification.
# * Ranking based on probabilistic predictions.
#
# * **Fairness tasks:**
# * Assessment of unfairness using Fairlearn metrics and Fairlearn dashboard.
# * Mitigation of unfairness using Fairlearn mitigation algorithms.
#
# * **Performance metrics:**
# * Area under ROC curve.
# * Worst-case area under ROC curve.
# * Balanced accuracy.
#
# * **Fairness metrics:**
# * Demographic parity difference (for both binary and continuous predictions).
#
# * **Mitigation algorithms:**
# * `fairlearn.reductions.ExponentiatedGradient`
# * `fairlearn.reductions.GridSearch`
# * `fairlearn.postprocessing.ThresholdOptimizer`
#
# ## Overview
#
# We consider the task of ranking students for admission to law school using the data collected in [Law School Admissions Council's (LSAC) National Longitudinal Bar Passage Study](https://eric.ed.gov/?id=ED469370); specifically, the version downloaded from [Project SEAPHE](http://www.seaphe.org/databases.php). We highlight some of the fairness considerations that come up not only in school admissions, but also in other ranking scenarios. Necessarily, our example is simplified and ignores many real-world considerations specific to school admissions.
#
# The data set contains information about law students collected by LSAC between 1991 and 1997. Some of the information is available at the admission time (such as the undergraduate GPA and LSAT score), and some describes the performance of the students once admitted. We also have access to their self-identified race. To simplify this example, we will limit the attention to those self-identified as **black** and **white** (two largest groups) and restrict our attention to two features (undergraduate GPA and LSAT score).
#
# To help with ranking law school applicants, we train a model that uses the information that is available about a student at the admission time to predict the probability that they will pass their bar exam. The predictions of our model are intended to be used (among other factors) by admission officers to select the applicants. After training the initial model, we examine differences in the predictions it induces across two the two groups. We then mitigate these differences using three Fairlearn algorithms: `GridSearch`, `ThresholdOptimizer` and `ExponentiatedGradient`.
#
# **Usage notes:** This notebook is intended as an example of Fairlearn functionality and not a fully realistic case study of an admission scenario. In real world, one should think carefully about whether it is appropriate to rank or score individuals. Also, additional features beyond the two considered here (GPA and LSAT scores) should be considered in practice, as recommended by the authors of the [LSAC study](https://eric.ed.gov/?id=ED469370). Finally, in real-world settings, it would be inappropriate to restrict attention to only two of the subgroups without evaluating the impacts on other individuals.
# ## Data
#
# We download the data using the `tempeh` package, which already filters the set of students to black and white and splits them into training and test subsets. The training and test data sets are loaded in three parts:
#
# * **X_train**, **X_test**: features describing the training and test data; `tempeh` provides two features: `ugpa` (undegraduate GPA) and `lsat` (LSAT score)
#
# * **y_train**, **y_test**: labels of the training and test data; the labels are 0 or 1, indicating whether a student passed the bar exam by the 2nd attempt
#
# * **A_train**, **A_test**: self-identified race of each student (black or white)
# +
import numpy as np
import pandas as pd
from IPython.display import display, HTML
# Load the data using the tempeh package
from tempeh.configurations import datasets
dataset = datasets['lawschool_passbar']()
X_train, X_test = dataset.get_X(format=pd.DataFrame)
y_train, y_test = dataset.get_y(format=pd.Series)
A_train, A_test = dataset.get_sensitive_features(name='race', format=pd.Series)
# Combine all training data into a single data frame and glance at a few rows
all_train = pd.concat([X_train, y_train, A_train], axis=1)
display(all_train)
# -
# Now, let us examine the data more closely. We look at the distributions of `lsat` and `ugpa` by race (summarized via quartiles), and compare them with the bar passage rates.
# +
all_train_grouped = all_train.groupby('race')
counts_by_race = all_train_grouped[['lsat']].count().rename(
columns={'lsat': 'count'})
quartiles_by_race = all_train_grouped[['lsat','ugpa']].quantile([.25, .50, .75]).rename(
index={0.25: "25%", 0.5: "50%", 0.75: "75%"}, level=1).unstack()
rates_by_race = all_train_grouped[['pass_bar']].mean().rename(
columns={'pass_bar': 'pass_bar_rate'})
summary_by_race = pd.concat([counts_by_race, quartiles_by_race, rates_by_race], axis=1)
display(summary_by_race)
# -
# The majority of the students in the study are white. There is a notable gap between white and black students in their incoming academic credentials: the 75th percentile of the LSAT scores of black students is lower than the 25th percentile of the LSAT scores among white students. There is a less severe, but still substantial gap in UGPA. The achievement gap is greatly diminished in terms of the bar passage rate (78% for black students and 97% for white students). The authors of the [LSAC study](https://eric.ed.gov/?id=ED469370) conclude that this justifies admission practices that look beyond LSAT and UGPA. However, in this simplified example, we build predictors of bar passage from these two variables alone.
# ## Unmitigated Predictor
#
# We first train a standard logistic regression predictor that does not seek to incorporate any notion of fairness.
# +
from sklearn.linear_model import LogisticRegression
unmitigated_predictor = LogisticRegression(solver='liblinear', fit_intercept=True)
unmitigated_predictor.fit(X_train, y_train)
# -
# We view the probabilistic predictions produced by the logistic model as scores and evaluate the quality of the ranking they produce in terms of the [area under the ROC curve](https://en.wikipedia.org/wiki/Receiver_operating_characteristic#Area_under_the_curve) (AUC). AUC is equal to the probability that a randomly chosen positive example (i.e., a student who passes the bar) is scored above a randomly chosen negative example (i.e., a student who does not pass the bar). An AUC of 0.5 means that the scores are no better than a random coin flip, whereas AUC of 1.0 means that the scores perfectly separate positives from negatives. The AUC metric has two desirable properties: (1) it is preserved by monotone transformations of the score, and (2) it is not sensitive to the imbalance between positives and negatives, which is quite severe in our example, with the overall bar passage rate above 94%.
#
# Note that the logistic regression estimator above does not seek to optimize AUC directly, but only seeks to optimize the logistic loss. However, a good logistic loss is also expected to yield a good AUC.
#
# To obtain the AUC values for the overall student population as well as black and white subpopulations, we use the **group metric** variant of the `sklearn` metric [`roc_auc_score`](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.roc_auc_score.html).
# +
from fairlearn.metrics import roc_auc_score_group_summary
# a convenience function that transforms the result of a group metric call into a data frame
def summary_as_df(name, summary):
a = pd.Series(summary.by_group)
a['overall'] = summary.overall
return pd.DataFrame({name: a})
scores_unmitigated = pd.Series(unmitigated_predictor.predict_proba(X_test)[:,1], name="score_unmitigated")
auc_unmitigated = summary_as_df(
"auc_unmitigated", roc_auc_score_group_summary(y_test, scores_unmitigated, sensitive_features=A_test))
display(HTML('<span id="auc_unmitigated">'),
auc_unmitigated,
HTML('</span>'))
# -
# We next examine how the unmitigated predictor affects applicants of different race when it is used to score them. We plot the CDFs of the scores it generates for each group. We then consider all possible thresholds on the value of the score, and for each threshold check the fraction of black vs white students above the threshold. The largest observed difference across all possible thresholds is referred to as the **demographic parity difference** or **demographic disparity** (see [Agarwal et al. 2018](http://proceedings.mlr.press/v97/agarwal19d.html), where it is reffered to as SP disparity). Pictorially, this corresponds to the largest vertical difference between the two CDFs. Note that this disparity metric is preserved under monotone transformations of the scores.
# +
# %matplotlib inline
import matplotlib.pyplot as plt
from scipy.stats import cumfreq
def compare_cdfs(data, A, num_bins=100):
cdfs = {}
assert len(np.unique(A)) == 2
limits = ( min(data), max(data) )
s = 0.5 * (limits[1] - limits[0]) / (num_bins - 1)
limits = ( limits[0]-s, limits[1] + s)
for a in np.unique(A):
subset = data[A==a]
cdfs[a] = cumfreq(subset, numbins=num_bins, defaultreallimits=limits)
lower_limits = [v.lowerlimit for _, v in cdfs.items()]
bin_sizes = [v.binsize for _,v in cdfs.items()]
actual_num_bins = [v.cumcount.size for _,v in cdfs.items()]
assert len(np.unique(lower_limits)) == 1
assert len(np.unique(bin_sizes)) == 1
assert np.all([num_bins==v.cumcount.size for _,v in cdfs.items()])
xs = lower_limits[0] + np.linspace(0, bin_sizes[0]*num_bins, num_bins)
disparities = np.zeros(num_bins)
for i in range(num_bins):
cdf_values = np.clip([v.cumcount[i]/len(data[A==k]) for k,v in cdfs.items()],0,1)
disparities[i] = max(cdf_values)-min(cdf_values)
return xs, cdfs, disparities
def plot_and_compare_cdfs(data, A, num_bins=100, loc='best'):
xs, cdfs, disparities = compare_cdfs(data, A, num_bins)
for k, v in cdfs.items():
plt.plot(xs, v.cumcount/len(data[A==k]), label=k)
assert disparities.argmax().size == 1
d_idx = disparities.argmax()
xs_line = [xs[d_idx],xs[d_idx]]
counts = [v.cumcount[d_idx]/len(data[A==k]) for k, v in cdfs.items()]
ys_line = [min(counts), max(counts)]
plt.plot(xs_line, ys_line, 'o--')
disparity_label = "max disparity = {0:.3f}\nat {1:0.3f}".format(disparities[d_idx], xs[d_idx])
plt.text(xs[d_idx], 1, disparity_label, ha="right", va="top")
plt.xlabel(data.name)
plt.ylabel("cumulative frequency")
plt.legend(loc=loc)
plt.show()
display(HTML('<span id="disparity_unmitigated">'))
plot_and_compare_cdfs(scores_unmitigated, A_test)
display(HTML('</span>'))
# -
# We see that the largest disparity of about 0.6 occurs at the threshold value 0.94: only 23% of black students, but 83% of white students are above this threshold.
# ## Mitigating Demographic Disparity with Grid Search
#
# We next show how to mitigate the demographic disparity using the `GridSearch` algorithm of Fairlearn. We will use this algorithm to obtain several models that achieve various trade-offs between accuracy (measured by AUC) and demographic disparity.
#
# The `GridSearch` variant that we will use was developed for *classification* under demographic parity, but the experiments of
# [Agarwal et al. (2018)](http://proceedings.mlr.press/v97/agarwal19d.html) show that it also performs well for *logistic regression* (viewed as the probability prediction) under demographic parity. While the resulting logistic models mitigate the demographic disparity, they might not be well calibrated (unlike unmitigated logistic models), so we use Platt's scaling for [calibration](https://scikit-learn.org/stable/modules/calibration.html). Note that Platt's scaling is a monotone transformation, and so it has no effect on the AUC values or the demographic disparity of the resulting model. However, it makes the predicted scores interpretable as probabilities.
#
# `GridSearch` generates models corresponding to various Lagrange multiplier vectors of the underlying constraint optimization problem. We will compute 41 models on a grid of Lagrange multiplier vectors whose L1-norm is bounded by 10. For details on how the search works, refer to Section 3.4 of [Agarwal et. al (2018)](http://proceedings.mlr.press/v80/agarwal18a.html). The following cell may take a couple of minutes to run:
# +
from fairlearn.reductions import GridSearch, DemographicParity
from sklearn.calibration import CalibratedClassifierCV
sweep = GridSearch(LogisticRegression(solver='liblinear', fit_intercept=True),
constraints=DemographicParity(),
grid_size=41,
grid_limit=10)
sweep.fit(X_train, y_train, sensitive_features=A_train)
calibrated_predictors = []
for predictor in sweep.predictors_:
calibrated = CalibratedClassifierCV(base_estimator=predictor, cv='prefit', method='sigmoid')
calibrated.fit(X_train, y_train)
calibrated_predictors.append(calibrated)
# -
# We next assess the accuracy and disparity of the obtained predictors in a scatter plot, with *x* axis showing the worst-case AUC among the two subpopulations (of black and white students) and *y* axis showing the demographic disparity. Ideal models would be in the bottom right.
# +
from fairlearn.metrics import roc_auc_score_group_min
def auc_disparity_sweep_plot(predictors, names, marker='o', scale_size=1, zorder=-1):
roc_auc = np.zeros(len(predictors))
disparity = np.zeros(len(predictors))
for i in range(len(predictors)):
preds = predictors[i].predict_proba(X_test)[:,1]
roc_auc[i] = roc_auc_score_group_min(y_test, preds, sensitive_features=A_test)
_, _, dis = compare_cdfs(preds, A_test)
disparity[i] = dis.max()
plt.scatter(roc_auc, disparity,
s=scale_size * plt.rcParams['lines.markersize'] ** 2, marker=marker, zorder=zorder)
for i in range(len(roc_auc)):
plt.annotate(names[i], (roc_auc[i], disparity[i]), xytext=(3,2), textcoords="offset points", zorder=zorder+1)
plt.xlabel("worst-case AUC")
plt.ylabel("demographic disparity")
auc_disparity_sweep_plot(calibrated_predictors, names=range(len(calibrated_predictors)))
auc_disparity_sweep_plot([unmitigated_predictor], names=[''], marker='*', zorder=1, scale_size=5)
plt.show()
# -
# Model 33 has the lowest disparity, but its worst-case AUC is essentially the same as that of a coin flip. The unmitigated model, marked as a star, has a good worst-case AUC, but large disparity. We examine models 35 and 36: their AUC values are well above 0.6 and they substantially reduce the demographic disparity compared with the unmitigated model:
# +
scores_model35 = pd.Series(calibrated_predictors[35].predict_proba(X_test)[:,1], name="score_model35")
scores_model36 = pd.Series(calibrated_predictors[36].predict_proba(X_test)[:,1], name="score_model36")
auc_model35 = summary_as_df(
"auc_model35", roc_auc_score_group_summary(y_test, scores_model35, sensitive_features=A_test))
auc_model36 = summary_as_df(
"auc_model36", roc_auc_score_group_summary(y_test, scores_model36, sensitive_features=A_test))
display(HTML('<span id="grid_search_comparison">'),
pd.concat([auc_model35, auc_model36, auc_unmitigated], axis=1),
HTML('</span>'))
plot_and_compare_cdfs(scores_model35, A_test)
plot_and_compare_cdfs(scores_model36, A_test)
plot_and_compare_cdfs(scores_unmitigated, A_test)
# -
# ### Comparing Probabilistic Predictors using the Dashboard
#
# Next, we compare the three predictors above (unmitigated, model 35 and model 36) using `FairlearnDashboard`. The dashboard currently does not evaluate the demographic disparity of probabilistic scores, but instead evaluates the disparity in mean predictions—in this case, this amounts to the difference between mean predictions for the white and black subpopulations. However, thanks to the calibration step, the disparity in mean predictions qualitatively agrees with the demographic disparity (albeit this is not always the case and in general they could go in opposite directions).
from fairlearn.widget import FairlearnDashboard
FairlearnDashboard(sensitive_features=A_test, sensitive_feature_names=['Race'],
y_true=y_test,
y_pred={"unmitigated": scores_unmitigated, "model35": scores_model35, "model36": scores_model36})
# ## Obtaining Low-Disparity Classifiers
#
# In this section, we shift attention from the task of scoring and ranking students to the task of automatically classifying students, for example, in order to screen them for an interview or a deeper review of their application materials. Our goal is to obtain a _classifier_ that maximizes AUC while respecting demographic parity.
#
# The outputs of a classifier are either 0 or 1, so it is possible to re-interpret the AUC of a classifier as the *balanced accuracy*, meaning the accuracy under the distribution re-weighted to have the same mass of positive and negative examples. Demographic disparity can also be interpreted as the difference between the rates at which the students of either race are classified as 1; we refer to this rate as the _selection rate_.
#
# ### Postprocessing
#
# We first show how to obtain low-disparity classifiers by thresholding scores—such as the scores produced by unmitigated logistic regression—using the postprocessing algorithm of [Hardt et al. (2016)](https://arxiv.org/abs/1610.02413), implemented in the class `ThresholdOptimizer`. This algorithm finds thresholds that optimize accuracy subject to the constraint that there be no demographic disparity on the training data. Since our goal here is to optimize _balanced_ accuracy rather than accuracy, we first re-balance the data by randomly subsampling positive examples, so they are equal in number to negative examples. We then pass this re-balanced data set to `ThresholdOptimizer`. Since the accuracy of a classifier on the re-balanced data set is in expectation equal to the AUC on the original data, `ThresholdOptimizer` now seeks to optimize our desired accuracy metric.
# +
from sklearn.base import BaseEstimator, ClassifierMixin
from fairlearn.postprocessing import ThresholdOptimizer
# We want to apply ThresholdOptimizer to the probabilities returned
# by the unmitigated logistic regression predictor. Since ThresholdOptimizer
# applies thresholding to the output of predict(), but LogisticRegression
# returns probabilities (of both classes) in predict_proba(), we need to
# use the following wrapper for LogisticRegression.
class LogisticRegressionAsRegression(BaseEstimator, ClassifierMixin):
def __init__(self, logistic_regression_estimator):
self.logistic_regression_estimator = logistic_regression_estimator
def fit(self, X, y):
self.logistic_regression_estimator.fit(X, y)
return self
def predict(self, X):
# use predict_proba to get real values instead of 0/1, select only prob for 1
scores = self.logistic_regression_estimator.predict_proba(X)[:,1]
return scores
balanced_index_pass0 = y_train[y_train==0].index
balanced_index_pass1 = y_train[y_train==1].sample(n=balanced_index_pass0.size, random_state=0).index
balanced_index = balanced_index_pass0.union(balanced_index_pass1)
pp_estimator = ThresholdOptimizer(
estimator=LogisticRegressionAsRegression(unmitigated_predictor),
constraints="demographic_parity",
prefit=True)
pp_estimator.fit(X_train.iloc[balanced_index,:], y_train.iloc[balanced_index],
sensitive_features=A_train.iloc[balanced_index])
# -
# We next evaluate AUC (balanced accuracy) and demographic disparity (disparity in selection rates) of the black and white students on the test data; note that we use the actual test data (not a re-balanced version, which we only used for training purposes).
# +
from fairlearn.metrics import mean_prediction_group_summary
scores_pp = pd.Series(pp_estimator.predict(X_test, sensitive_features=A_test), name="scores_post")
auc_pp = summary_as_df(
"auc_post",
roc_auc_score_group_summary(y_test, scores_pp, sensitive_features=A_test))
sel_pp = summary_as_df(
"selection_post",
mean_prediction_group_summary(y_test, scores_pp, sensitive_features=A_test))
pp_summary = pd.concat([auc_pp, sel_pp], axis=1)
pp_summary.loc['disparity']=(pp_summary.loc['white']-pp_summary.loc['black']).abs()
pp_summary.loc['disparity', pp_summary.columns.str.startswith('auc')]='-'
display(pp_summary)
# -
# The original unmitigated scores have the demographic disparity around 0.6 (see [here](#disparity_unmitigated)). We see that `ThresholdOptimizer` dramatically reduces the disparity to around 0.1. At the same time, the AUC in each subpopulation is at or above 0.65, a moderate drop from the unmitigated values of 0.72 and 0.74 (see [here](#auc_unmitigated)). This is a more favorable trade-off than the one achieved by model 35 above, with the disparity of 0.4 and the worst-case AUC of around 0.62 (see [here](#grid_search_comparison)). However, note that `ThresholdOptimizer` is a classifier, and so it can only work as a crude ranker. Additionally, `ThresholdOptimizer` uses the sensitive feature (in this instance race) at the prediction time, by applying a different threshold to unmitigated scores depending on race. In some use cases, these two properties might be undesirable. We next show how to obtain a classifier that also seeks to achieve low demographic disparity, but without requiring access to the sensitive feature at the evaluation time.
#
# *Note*: `ThresholdOptimizer` produces randomized predictions, so the AUC and selection rate of postprocessing will vary if you re-run the cell above. Also, while `ThresholdOptimizer` is guaranteed to achieve zero demographic disparity on its training data, this does not mean it will achieve zero demographic disparity on the test data for several reasons: (1) the training data is balanced whereas test data is not, so test data comes from a different distribution than training data; (2) even if training and test data were coming from the same distribution, there would be some differences due to finite sample sizes.
# ### Exponentiated Gradient
#
# `ExponentiatedGradient` also seeks to find a classifier that optimizes accuracy while placing a constraint on the demographic disparity. However, it operates as a *reduction* to standard classification, taking any estimator as a black box. During its run it repeatedly re-fits the estimator on variously reweighted training data and eventually produces a randomized classifier of the same type as the provided black-box estimator. This means that if the black box does not have access to the sensitive feature, neither will the predictor fitted by `ExponentiatedGradient`.
#
# We next train two classifiers via `ExponentiatedGradient`. Both use `LogisticRegression` as a black box. However, one has only access to the original features (**X_train** and **X_test**), whereas the other one also has access to the sensitive features, which we include in the extended feature set (**XA_train** and **XA_test**). Both classifiers optimize AUC subject to the constraint that demographic disparity on training data is at most 0.01. We also set the convergence parameter `nu` to `1e-6` to optimize to numerical precision (the default is to optimize to statistical precision, which we override here).
# +
from fairlearn.reductions import ExponentiatedGradient
XA_train = pd.concat([X_train, A_train=='black'], axis=1).astype(float)
XA_test = pd.concat([X_test, A_test=='black'], axis=1).astype(float)
expgrad_X = ExponentiatedGradient(
LogisticRegression(solver='liblinear', fit_intercept=True),
constraints=DemographicParity(),
eps=0.01,
nu=1e-6)
expgrad_XA = ExponentiatedGradient(
LogisticRegression(solver='liblinear', fit_intercept=True),
constraints=DemographicParity(),
eps=0.01,
nu=1e-6)
expgrad_X.fit(
X_train.iloc[balanced_index,:],
y_train.iloc[balanced_index],
sensitive_features=A_train.iloc[balanced_index])
expgrad_XA.fit(
XA_train.iloc[balanced_index,:],
y_train.iloc[balanced_index],
sensitive_features=A_train.iloc[balanced_index])
# +
scores_expgrad_X = pd.Series(expgrad_X.predict(X_test), name="scores_expgrad_X")
scores_expgrad_XA = pd.Series(expgrad_XA.predict(XA_test), name="scores_expgrad_XA")
auc_expgrad_X = summary_as_df(
"auc_expgrad_X",
roc_auc_score_group_summary(y_test, scores_expgrad_X, sensitive_features=A_test))
sel_expgrad_X = summary_as_df(
"selection_expgrad_X",
mean_prediction_group_summary(y_test, scores_expgrad_X, sensitive_features=A_test))
auc_expgrad_XA = summary_as_df(
"auc_expgrad_XA",
roc_auc_score_group_summary(y_test, scores_expgrad_XA, sensitive_features=A_test))
sel_expgrad_XA = summary_as_df(
"selection_expgrad_XA",
mean_prediction_group_summary(y_test, scores_expgrad_XA, sensitive_features=A_test))
classifier_summary = pd.concat([auc_pp, sel_pp, auc_expgrad_X, sel_expgrad_X, auc_expgrad_XA, sel_expgrad_XA], axis=1)
classifier_summary.loc['disparity']=(classifier_summary.loc['white']-classifier_summary.loc['black']).abs()
classifier_summary.loc['disparity', classifier_summary.columns.str.startswith('auc')]='-'
display(classifier_summary)
# -
# We see that exponentiated gradient variants generally achieve lower disparity on this data than `ThresholdOptimizer`. Without access to the sensitive feature at the test time, this comes at the cost of bringing the AUC essentially to that of a random coin toss (AUC of **expgrad_X** is close to 0.5). With access to the sensitive feature, the overall AUC is comparable to that achieved by `ThresholdOptimizer`, but `ThresholdOptimizer` achieves a better worst-case AUC across the two population.
#
# *Note*: `ExponentiatedGradient` produces randomized predictions (similarly to `ThresholdOptimizer`), so the AUC and selection rate will vary if you re-run the cell above. Also, because of a mismatch between the training and test distributions and because of finite samples, we do not expect `ExponentiatedGradient` to achieve test disparity equal to 0.01.
#
# We next show that if we are willing to tolerate a larger demographic disparity, it is possible to achieve non-trivial AUC values even without access to the sensitive feature. We run `ExponentiatedGradient` with the bound on the training disparity equal to 0.3:
# +
expgrad_X_alt = ExponentiatedGradient(
LogisticRegression(solver='liblinear', fit_intercept=True),
constraints=DemographicParity(),
eps=0.3, # This has changed from 0.01 in the above examples
nu=1e-6)
expgrad_X_alt.fit(
X_train.iloc[balanced_index,:],
y_train.iloc[balanced_index],
sensitive_features=A_train.iloc[balanced_index])
scores_expgrad_X_alt = pd.Series(
expgrad_X_alt.predict(X_test), name="scores_expgrad_X_alt")
auc_expgrad_X_alt = summary_as_df(
"auc_expgrad_X_alt",
roc_auc_score_group_summary(y_test, scores_expgrad_X_alt, sensitive_features=A_test))
sel_expgrad_X_alt = summary_as_df(
"selection_expgrad_X_alt",
mean_prediction_group_summary(y_test, scores_expgrad_X_alt, sensitive_features=A_test))
auc_expgrad_X_alt.loc['disparity'] = '-'
sel_expgrad_X_alt.loc['disparity'] = (sel_expgrad_X_alt.loc['white'] - sel_expgrad_X_alt.loc['black']).abs()
display(pd.concat([auc_expgrad_X_alt, sel_expgrad_X_alt], axis=1))
# -
# ### Comparing Classifiers using the Dashboard
#
# We finish this section by comparing the four predictors above using `FairlearnDashboard`:
FairlearnDashboard(sensitive_features=A_test, sensitive_feature_names=['Race'],
y_true=y_test,
y_pred={"postprocessing": scores_pp,
"expgrad_X": scores_expgrad_X,
"expgrad_X_alt": scores_expgrad_X_alt,
"expgrad_XA": scores_expgrad_XA})
|
notebooks/Mitigating Disparities in Ranking from Binary Data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Retrieve published versions of articles on arXiv with `pyzotero`
# ## Load entire Zotero library
from pyzotero import zotero
zot = zotero.Zotero(library_id, 'user', api_key)
items = zot.everything(zot.top())
# ## Finds arXiv preprints in library
# + code_folding=[]
preprints = []
for item in items:
if 'publicationTitle' in item['data']:
if (
any('arxiv' in str for str in [item['data']['publicationTitle'].lower(), item['data']['url'].lower()]) and
(' ' not in item['data']['url'])
):
preprints.append(item)
# -
# ## Scrape URLs for DOIs
# +
import requests
import re
import textwrap
regex = 'name="citation_doi" content=(.*?)/>'
pattern = re.compile(regex)
new = {}
keys, titles, DOIs = [], [], []
for item in preprints:
r = requests.get(item['data']['url'])
match = re.search(pattern, r.text)
if match:
DOI = match[1].replace('"', '').strip()
new[item['data']['key']] = DOI
keys.append(item['data']['key'])
titles.append(item['data']['title'])
DOIs.append(DOI)
# -
print(f'{len(new)} DOIs have been found among {len(preprints)} preprints.')
print('\n')
print('\n'.join(f'{key:<9} {textwrap.shorten(title, width=60):<60} {DOI:<30}' for key, title, DOI in zip(keys, titles, DOIs)))
# ## Update items in library
success = []
for key, DOI in new.items():
to_update = zot.item(key)
to_update['data']['DOI'] = DOI
success.append(zot.update_item(to_update))
if all(success):
print('All preprints have been updated with new DOI.')
else:
pass
|
pyzotero/pyzotero-doi-retriever.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# -
#export
from exp.nb_02 import *
path= Path('../PCImages')
path2fm= '../FeatureMatsMerged/TheGreatCollection.txt'
path2colnames= 'FeatureMatIndex.txt'
fm= fm_from_txt(path2fm, path2colnames)
fm.shape
cols= ['id','date','time','label1','label2','label3','ind','wallcrash','temp','LP','HL',
't_stretch','t_relax','framecut','fps','medium','passage', 'ms_ch1','mf_ch1',
'mf_real_ch1','cv_nn_ch1','cvn_ch1','ms_ch2','mf_ch2','mf_real_ch2','cv_nn_ch2','cvn_ch2']
cols_filtered= [c for c in fm.columns if c not in cols]
cols_filtered= [c for c in cols_filtered if '2s' not in c]
len(cols_filtered)
#export
def get_filtered_cols(df, cols2keep=None):
if cols2keep is None:
cols= ['id','date','time','label1','label2','label3','ind','wallcrash','temp','LP','HL',
't_stretch','t_relax','framecut','fps','medium','passage', 'ms_ch1','mf_ch1',
'mf_real_ch1','cv_nn_ch1','cvn_ch1','ms_ch2','mf_ch2','mf_real_ch2','cv_nn_ch2','cvn_ch2']
else: cols= cols2keep
cols= [c for c in df.columns if c not in cols]
return [c for c in cols if '2s' not in c]
cols_filtered= get_filtered_cols(fm)
len(cols_filtered)
#export
class CellDataSet(Dataset):
def __init__(self, path, fm, cols2keep, label = 'label1', cell_phase = None, trfms=None,
random_sample=None):
self.fm = fm
self.cols2keep= cols2keep
self.y = self.fm[label]
self.id2label= id2label(self.fm.id, self.y)
if cell_phase is None:
self.data_files = get_filtered_files(path, img_ids=self.fm.id, labels=self.y)
else:
self.data_files= [
f for f in get_filtered_files(path, img_ids=self.fm.id, labels=self.y) if cell_phase in str(f)
]
if random_sample is not None: self.data_files= random.sample(self.data_files, random_sample)
self.trfms = get_trfms(trfms)
def __getindex__(self, idx):
return load_file(self.data_files[idx])
def __len__(self):
return len(self.data_files)
def get_row(self, img_id):
x_np= np.array(self.fm.loc[self.fm['id'].isin([img_id])][self.cols2keep]).flatten()
return torch.cuda.FloatTensor(x_np)
def __getitem__(self, idx):
if torch.is_tensor(idx):
idx = idx.tolist()
img_name = self.data_files[idx]
img_id, phase = split_fn(img_name)
label= self.id2label[img_id]
image = PIL.Image.open(img_name)
fm_row= self.get_row(img_id)
if self.trfms:
image = self.trfms(image)
return image, fm_row, label
#export
import torch.nn.functional as F
# # Concatenate models:
#export
class Learner():
def __init__(self, dataset, model):
self.ds= dataset
self.img_size, self.c= self.ds[0][0].shape[0], len(np.unique(self.ds.y))
self.model= model #get_model(model.cuda(), image_size=self.img_size, c=self.c)
self.loss= nn.CrossEntropyLoss()
def fit(self, epochs=1, bs=32, lr = 1e-5):
opt= torch.optim.Adam(self.model.parameters(), lr=lr)
train_dl, valid_dl= ds2dls(self.ds, bs=bs)
for epoch in range(epochs):
self.model.train()
for img_xb, data_xb, yb in train_dl:
img_xb= img_xb.to(device)
data_xb= data_xb.to(device)
yb= yb.to(device)
loss = self.loss(self.model(img_xb, data_xb), yb)
loss.backward()
opt.step()
opt.zero_grad()
self.model.eval()
with torch.no_grad():
tot_loss,tot_acc = 0.,0.
for img_xb, data_xb, yb in valid_dl:
img_xb= img_xb.to(device)
data_xb= data_xb.to(device)
yb= yb.to(device)
pred = self.model(img_xb, data_xb)
pred= pred.to(device)
tot_loss += self.loss(pred, yb)
tot_acc += accuracy(pred,yb)
nv = len(valid_dl)
print(epoch, tot_loss/nv, tot_acc/nv)
#return tot_loss/nv, tot_acc/nv
# ## Model for the feature matrix:
class FeatureMatrixModel(nn.Module):
def __init__(self, n_in, n_out):
super().__init__()
self.n_in = n_in
self.lin1 = nn.Linear(self.n_in, 200)
self.lin2 = nn.Linear(200, 70)
self.lin3 = nn.Linear(70, n_out)
self.bn1 = nn.BatchNorm1d(self.n_in)
self.bn2 = nn.BatchNorm1d(200)
self.bn3 = nn.BatchNorm1d(70)
self.drops = nn.Dropout(0.3)
def forward(self, x_cont):
x = self.bn1(x_cont)
x = F.relu(self.lin1(x))
x = self.drops(x)
x = self.bn2(x)
x = F.relu(self.lin2(x))
x = self.drops(x)
x = self.bn3(x)
x = self.lin3(x)
return x
class CombinedModel(nn.Module):
def __init__(self, n_out_cnn=50, n_out_data=50):
super(CombinedModel, self).__init__()
self.n_out_cnn, self.n_out_data= n_out_cnn, n_out_data
self.n_out= self.n_out_cnn + self.n_out_data
self.n_classes= 7
self.fc0= Resize(350).cuda()
self.cnn = models.resnet34(pretrained=True).cuda()
self.cnn.conv1= nn.Conv2d(1, 64, kernel_size=(7, 7), stride=(2, 2), padding=(3, 3), bias=False).cuda()
self.cnn.fc= nn.Linear(in_features=512, out_features=self.n_out_cnn).cuda()
self.fm_net= FeatureMatrixModel(len(cols_filtered), self.n_out_data).cuda()
self.fc1 = nn.Linear(self.n_out, self.n_classes).cuda()
def forward(self, image, data):
x1 = self.cnn(self.fc0(image))
x2 = self.fm_net(data)
x = torch.cat((x1, x2), dim=1)
x = F.relu(x)
x = self.fc1(x)
return x
# ### Ratio of model outputs:
ds= CellDataSet(path, fm, cols_filtered, label = 'label1', cell_phase = None,
trfms= [CenterCrop(size=350),
transforms.Grayscale(num_output_channels=1),
ToFloatTensor()],
random_sample=1000)
learn= Learner(ds, CombinedModel(50,10))
learn.fit(1, bs=8, lr=1e-5)
ds= CellDataSet(path, fm, cols_filtered, label = 'label1', cell_phase = None,
trfms= [CenterCrop(size=350),
transforms.Grayscale(num_output_channels=1),
ToFloatTensor()],
random_sample=1000)
learn= Learner(ds, CombinedModel(100,10))
learn.fit(1, bs=8, lr=1e-5)
ds= CellDataSet(path, fm, cols_filtered, label = 'label1', cell_phase = None,
trfms= [CenterCrop(size=350),
transforms.Grayscale(num_output_channels=1),
ToFloatTensor()],
random_sample=1000)
learn= Learner(ds, CombinedModel(100,100))
learn.fit(1, bs=8, lr=1e-5)
ds= CellDataSet(path, fm, cols_filtered, label = 'label1', cell_phase = None,
trfms= [CenterCrop(size=350),
transforms.Grayscale(num_output_channels=1),
ToFloatTensor()],
random_sample=1000)
learn= Learner(ds, CombinedModel(20,10))
learn.fit(1, bs=8, lr=1e-5)
ds= CellDataSet(path, fm, cols_filtered, label = 'label1', cell_phase = None,
trfms= [CenterCrop(size=350),
transforms.Grayscale(num_output_channels=1),
ToFloatTensor()],
random_sample=1000)
learn= Learner(ds, CombinedModel(50,40))
learn.fit(1, bs=8, lr=1e-5)
ds= CellDataSet(path, fm, cols_filtered, label = 'label1', cell_phase = None,
trfms= [CenterCrop(size=350),
transforms.Grayscale(num_output_channels=1),
ToFloatTensor()],
random_sample=1000)
learn= Learner(ds, CombinedModel(50,20))
learn.fit(1, bs=8, lr=1e-5)
# ### Dropout:
#export
class FeatureMatrixModel(nn.Module):
def __init__(self, n_in, n_out, p):
super().__init__()
self.n_in = n_in
self.lin1 = nn.Linear(self.n_in, 200)
self.lin2 = nn.Linear(200, 70)
self.lin3 = nn.Linear(70, n_out)
self.bn1 = nn.BatchNorm1d(self.n_in)
self.bn2 = nn.BatchNorm1d(200)
self.bn3 = nn.BatchNorm1d(70)
self.drops = nn.Dropout(p)
def forward(self, x_cont):
x = self.bn1(x_cont)
x = F.relu(self.lin1(x))
x = self.drops(x)
x = self.bn2(x)
x = F.relu(self.lin2(x))
x = self.drops(x)
x = self.bn3(x)
x = self.lin3(x)
return x
#export
class CombinedModel(nn.Module):
def __init__(self, n_out_cnn=50, n_out_data=50, p=0.5, n_features=44):
super(CombinedModel, self).__init__()
self.n_features= n_features
self.n_out_cnn, self.n_out_data= n_out_cnn, n_out_data
self.n_out= self.n_out_cnn + self.n_out_data
self.n_classes= 7
self.fc0= Resize(350).cuda()
self.cnn = models.resnet34(pretrained=True).cuda()
self.cnn.conv1= nn.Conv2d(1, 64, kernel_size=(7, 7), stride=(2, 2), padding=(3, 3), bias=False).cuda()
self.cnn.fc= nn.Linear(in_features=512, out_features=self.n_out_cnn).cuda()
self.fm_net= FeatureMatrixModel(n_features, self.n_out_data, p).cuda()
self.fc1 = nn.Linear(self.n_out, self.n_classes).cuda()
def forward(self, image, data):
x1 = self.cnn(self.fc0(image))
x2 = self.fm_net(data)
x = torch.cat((x1, x2), dim=1)
x = F.relu(x)
x = self.fc1(x)
return x
ds= CellDataSet(path, fm, cols_filtered, label = 'label1', cell_phase = None,
trfms= [CenterCrop(size=350),
transforms.Grayscale(num_output_channels=1),
ToFloatTensor()],
random_sample=1000)
learn= Learner(ds, CombinedModel(50,10, p=0.5))
learn.fit(1, bs=8, lr=1e-5)
ds= CellDataSet(path, fm, cols_filtered, label = 'label1', cell_phase = None,
trfms= [CenterCrop(size=350),
transforms.Grayscale(num_output_channels=1),
ToFloatTensor()],
random_sample=1000)
learn= Learner(ds, CombinedModel(50,10, p=0.7))
learn.fit(1, bs=8, lr=1e-5)
ds= CellDataSet(path, fm, cols_filtered, label = 'label1', cell_phase = None,
trfms= [CenterCrop(size=350),
transforms.Grayscale(num_output_channels=1),
ToFloatTensor()],
random_sample=1000)
learn= Learner(ds, CombinedModel(50,10, p=0.8))
learn.fit(1, bs=8, lr=1e-5)
ds= CellDataSet(path, fm, cols_filtered, label = 'label1', cell_phase = None,
trfms= [CenterCrop(size=350),
transforms.Grayscale(num_output_channels=1),
ToFloatTensor()],
random_sample=1000)
learn= Learner(ds, CombinedModel(50,10, p=0.8))
learn.fit(8, bs=8, lr=1e-5)
ds= CellDataSet(path, fm, cols_filtered, label = 'label1', cell_phase = None,
trfms= [CenterCrop(size=350),
transforms.Grayscale(num_output_channels=1),
ToFloatTensor()],
random_sample=1000)
learn= Learner(ds, CombinedModel(50,10, p=0.5))
learn.fit(8, bs=8, lr=1e-5)
ds= CellDataSet(path, fm, cols_filtered, label = 'label1', cell_phase = None,
trfms= [CenterCrop(size=350),
transforms.Grayscale(num_output_channels=1),
ToFloatTensor()],
random_sample=None)
learn= Learner(ds, CombinedModel(50,10, p=0.5))
learn.fit(1, bs=8, lr=1e-5)
# !python notebook2script.py 03_combined_model.ipynb
|
03_combined_model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Example: Compute Cost Matrix
# +
import os
import sys
import random
import numpy as np
import pandas as pd
from osrmcpy import OSRM, Coordinate
# -
# Matrix size (square matrix). Be aware of your hardware resources ;-)
N_LOCATIONS = 1000
MATRIX_SIZE = N_LOCATIONS**2
print(f'Matrix size is {MATRIX_SIZE:,}')
DATA_DIR = '/usr/local/app/data'
OSRM_DATASET = os.path.join(DATA_DIR, 'test_ireland', 'ireland-and-northern-ireland-latest.osrm')
osrm = OSRM(OSRM_DATASET.encode('utf-8'), contraction=True)
bottom_left = Coordinate(id=None, longitude=-6.57013, latitude=53.23382)
top_right = Coordinate(id=None, longitude=-6.23988, latitude=53.39977)
random.seed(19)
random_coordinate = lambda n: Coordinate(id=n, longitude=random.uniform(bottom_left.longitude, top_right.longitude),
latitude=random.uniform(bottom_left.latitude, top_right.latitude))
# %time coordinates = [random_coordinate(i) for i in range(N_LOCATIONS)]
# %time table = osrm.table(coordinates)
# %time df_table = pd.DataFrame(np.array(table).reshape(-1, 4), columns=['from', 'to', 'duration', 'distance'])
df_table.shape
df_table.head()
# compute costs only for specified indexes as sources
# %time table2 = osrm.table(coordinates, sources=[1, 2, 3])
# %time df_table2 = pd.DataFrame(np.array(table2).reshape(-1, 4), columns=['from', 'to', 'duration', 'distance'])
df_table2.shape
df_table2.head()
|
notebooks/osrmcpy_compute_matrix.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# These class functions are the adversarial attack systems for NER; if entities == True an entity attack is performed, if entities == False an entity context attack is performed. It has options for performing a Random Attack (default is set to False).
# +
import argparse
import glob
import logging
import os
import random
import criteria
# import tensorflow.compat.v1 as tf
# tf.disable_v2_behavior()
import tensorflow_hub as hub
import numpy as np
import torch
from seqeval.metrics import precision_score, recall_score, f1_score
# from tensorboardX import SummaryWriter
from torch.nn import CrossEntropyLoss
from torch.utils.data import DataLoader, RandomSampler, SequentialSampler, TensorDataset
from torch.utils.data.distributed import DistributedSampler
from tqdm import tqdm, trange
from utils_ner import convert_examples_to_features, get_labels, read_examples_from_file
from transformers import AdamW, WarmupLinearSchedule
from transformers import WEIGHTS_NAME, BertConfig, BertForTokenClassification, BertTokenizer
# from transformers import RobertaConfig, RobertaForTokenClassification, RobertaTokenizer
import tensorflow as tf
# +
import pickle
import pandas as pd
import re
def save_obj(obj, name):
with open(name + '.pkl', 'wb') as f:
pickle.dump(obj, f, pickle.HIGHEST_PROTOCOL)
def load_obj(name):
with open(name + '.pkl', 'rb') as f:
return pickle.load(f, encoding='latin1')
# +
# import os
# os.environ["CUDA_DEVICE_ORDER"] = "PCI_BUS_ID"
# os.environ["CUDA_VISIBLE_DEVICES"] = "5"
# -
class InputExample(object):
"""A single training/test example for token classification."""
def __init__(self, guid, words, labels):
"""Constructs a InputExample.
Args:
guid: Unique id for the example.
words: list. The words of the sequence.
labels: (Optional) list. The labels for each word of the sequence. This should be
specified for train and dev examples, but not for test examples.
"""
self.guid = guid
self.words = words
self.labels = labels
# +
class AdversarialBERT():
def __init__(self):
##can uncomment to use a cache version of the USE
# nw_cache_path = '/data/dirksonar/NER_data/42480c3c7f42bf87d36d4c58fc4374b08dae2909/'
self.embed = hub.load("https://tfhub.dev/google/universal-sentence-encoder/4")
self.stop_words_set = self.get_stopwords()
self.set_seed(1, 0)
def initialize_essential(self, entities):
if entities == True:
pass
if entities == False:
## get cos sim matrix
path = '/data/dirksonar/TextFooler/TextFooler-master/TextFooler-master/cos_sim_counter_fitting.npy'
self.cos_sim = np.load (path)
self.idx2word = {}
self.word2idx = {}
pathcf = '/data/dirksonar/TextFooler/TextFooler-master/TextFooler-master/counter-fitted-vectors.txt'
print("Building vocab...")
with open(pathcf, 'r') as ifile:
for line in ifile:
word = line.split()[0]
if word not in self.idx2word:
self.idx2word[len(self.idx2word)] = word
self.word2idx[word] = len(self.idx2word) - 1
def load_obj(self, name):
with open(name + '.pkl', 'rb') as f:
return pickle.load(f, encoding='latin1')
def save_obj(self, obj, name):
with open(name + '.pkl', 'wb') as f:
pickle.dump(obj, f, pickle.HIGHEST_PROTOCOL)
def initialize_model(self,modelpath, labelfile):
# labellst2 = [i for j in labellst for i in j]
# self.labels = list(set(labellst2))
# print(self.labels)
self.labels = get_labels(labelfile)
# self.labels = ['I-location', 'I-group', 'O', 'B-creative-work', 'I-product', 'B-corporation', 'I-corporation', 'B-product', 'I-creative-work', 'B-location', 'B-group', 'I-person', 'B-person']
self.num_labels = len(self.labels)
# Use cross entropy ignore index as padding label id so that only real label ids contribute to the loss later
self.pad_token_label_id = CrossEntropyLoss().ignore_index
self.max_seq_length = 128
self.tokenizer = BertTokenizer.from_pretrained(modelpath, do_lower_case = False)
self.model = BertForTokenClassification.from_pretrained(modelpath)
def set_seed(self,num, n_gpu):
# random.seed(num)
np.random.seed(num)
torch.manual_seed(num)
if n_gpu > 0:
torch.cuda.manual_seed_all(num)
def cosine_similarity(self, v1, v2):
mag1 = np.linalg.norm(v1)
mag2 = np.linalg.norm(v2)
if (not mag1) or (not mag2):
return 0
return np.dot(v1, v2) / (mag1 * mag2)
def test_similarity(self,text1, text2):
vecs = self.embed([text1, text2])['outputs']
v1 = vecs[0]
v2 = vecs[1]
return self.cosine_similarity(v1, v2)
def get_stopwords(self):
'''
:return: a set of 266 stop words from nltk. eg. {'someone', 'anyhow', 'almost', 'none', 'mostly', 'around', 'being', 'fifteen', 'moreover', 'whoever', 'further', 'not', 'side', 'keep', 'does', 'regarding', 'until', 'across', 'during', 'nothing', 'of', 'we', 'eleven', 'say', 'between', 'upon', 'whole', 'in', 'nowhere', 'show', 'forty', 'hers', 'may', 'who', 'onto', 'amount', 'you', 'yours', 'his', 'than', 'it', 'last', 'up', 'ca', 'should', 'hereafter', 'others', 'would', 'an', 'all', 'if', 'otherwise', 'somehow', 'due', 'my', 'as', 'since', 'they', 'therein', 'together', 'hereupon', 'go', 'throughout', 'well', 'first', 'thence', 'yet', 'were', 'neither', 'too', 'whether', 'call', 'a', 'without', 'anyway', 'me', 'made', 'the', 'whom', 'but', 'and', 'nor', 'although', 'nine', 'whose', 'becomes', 'everywhere', 'front', 'thereby', 'both', 'will', 'move', 'every', 'whence', 'used', 'therefore', 'anyone', 'into', 'meanwhile', 'perhaps', 'became', 'same', 'something', 'very', 'where', 'besides', 'own', 'whereby', 'whither', 'quite', 'wherever', 'why', 'latter', 'down', 'she', 'sometimes', 'about', 'sometime', 'eight', 'ever', 'towards', 'however', 'noone', 'three', 'top', 'can', 'or', 'did', 'seemed', 'that', 'because', 'please', 'whereafter', 'mine', 'one', 'us', 'within', 'themselves', 'only', 'must', 'whereas', 'namely', 'really', 'yourselves', 'against', 'thus', 'thru', 'over', 'some', 'four', 'her', 'just', 'two', 'whenever', 'seeming', 'five', 'him', 'using', 'while', 'already', 'alone', 'been', 'done', 'is', 'our', 'rather', 'afterwards', 'for', 'back', 'third', 'himself', 'put', 'there', 'under', 'hereby', 'among', 'anywhere', 'at', 'twelve', 'was', 'more', 'doing', 'become', 'name', 'see', 'cannot', 'once', 'thereafter', 'ours', 'part', 'below', 'various', 'next', 'herein', 'also', 'above', 'beside', 'another', 'had', 'has', 'to', 'could', 'least', 'though', 'your', 'ten', 'many', 'other', 'from', 'get', 'which', 'with', 'latterly', 'now', 'never', 'most', 'so', 'yourself', 'amongst', 'whatever', 'whereupon', 'their', 'serious', 'make', 'seem', 'often', 'on', 'seems', 'any', 'hence', 'herself', 'myself', 'be', 'either', 'somewhere', 'before', 'twenty', 'here', 'beyond', 'this', 'else', 'nevertheless', 'its', 'he', 'except', 'when', 'again', 'thereupon', 'after', 'through', 'ourselves', 'along', 'former', 'give', 'enough', 'them', 'behind', 'itself', 'wherein', 'always', 'such', 'several', 'these', 'everyone', 'toward', 'have', 'nobody', 'elsewhere', 'empty', 'few', 'six', 'formerly', 'do', 'no', 'then', 'unless', 'what', 'how', 'even', 'i', 'indeed', 'still', 'might', 'off', 'those', 'via', 'fifty', 'each', 'out', 'less', 're', 'take', 'by', 'hundred', 'much', 'anything', 'becoming', 'am', 'everything', 'per', 'full', 'sixty', 'are', 'bottom', 'beforehand'}
'''
stop_words = ['a', 'about', 'above', 'across', 'after', 'afterwards', 'again', 'against', 'ain', 'all', 'almost', 'alone', 'along', 'already', 'also', 'although', 'am', 'among', 'amongst', 'an', 'and', 'another', 'any', 'anyhow', 'anyone', 'anything', 'anyway', 'anywhere', 'are', 'aren', "aren't", 'around', 'as', 'at', 'back', 'been', 'before', 'beforehand', 'behind', 'being', 'below', 'beside', 'besides', 'between', 'beyond', 'both', 'but', 'by', 'can', 'cannot', 'could', 'couldn', "couldn't", 'd', 'didn', "didn't", 'doesn', "doesn't", 'don', "don't", 'down', 'due', 'during', 'either', 'else', 'elsewhere', 'empty', 'enough', 'even', 'ever', 'everyone', 'everything', 'everywhere', 'except', 'first', 'for', 'former', 'formerly', 'from', 'hadn', "hadn't", 'hasn', "hasn't", 'haven', "haven't", 'he', 'hence', 'her', 'here', 'hereafter', 'hereby', 'herein', 'hereupon', 'hers', 'herself', 'him', 'himself', 'his', 'how', 'however', 'hundred', 'i', 'if', 'in', 'indeed', 'into', 'is', 'isn', "isn't", 'it', "it's", 'its', 'itself', 'just', 'latter', 'latterly', 'least', 'll', 'may', 'me', 'meanwhile', 'mightn', "mightn't", 'mine', 'more', 'moreover', 'most', 'mostly', 'must', 'mustn', "mustn't", 'my', 'myself', 'namely', 'needn', "needn't", 'neither', 'never', 'nevertheless', 'next', 'no', 'nobody', 'none', 'noone', 'nor', 'not', 'nothing', 'now', 'nowhere', 'o', 'of', 'off', 'on', 'once', 'one', 'only', 'onto', 'or', 'other', 'others', 'otherwise', 'our', 'ours', 'ourselves', 'out', 'over', 'per', 'please','s', 'same', 'shan', "shan't", 'she', "she's", "should've", 'shouldn', "shouldn't", 'somehow', 'something', 'sometime', 'somewhere', 'such', 't', 'than', 'that', "that'll", 'the', 'their', 'theirs', 'them', 'themselves', 'then', 'thence', 'there', 'thereafter', 'thereby', 'therefore', 'therein', 'thereupon', 'these', 'they','this', 'those', 'through', 'throughout', 'thru', 'thus', 'to', 'too','toward', 'towards', 'under', 'unless', 'until', 'up', 'upon', 'used', 've', 'was', 'wasn', "wasn't", 'we', 'were', 'weren', "weren't", 'what', 'whatever', 'when', 'whence', 'whenever', 'where', 'whereafter', 'whereas', 'whereby', 'wherein', 'whereupon', 'wherever', 'whether', 'which', 'while', 'whither', 'who', 'whoever', 'whole', 'whom', 'whose', 'why', 'with', 'within', 'without', 'won', "won't", 'would', 'wouldn', "wouldn't", 'y', 'yet', 'you', "you'd", "you'll", "you're", "you've", 'your', 'yours', 'yourself', 'yourselves']
stop_words = set(stop_words)
return stop_words
def pos_filter(self, ori_pos, new_pos_list):
same = [True if ori_pos == new_pos
else False
for new_pos in new_pos_list]
return same
def pick_most_similar_words_batch(self, src_words, sim_mat, idx2word, ret_count=10, threshold=0.5):
"""
embeddings is a matrix with (d, vocab_size)
"""
sim_order = np.argsort(-sim_mat[src_words, :])[:, 1:1 + ret_count]
sim_words, sim_values = [], []
for idx, src_word in enumerate(src_words):
sim_value = sim_mat[src_word][sim_order[idx]]
mask = sim_value >= threshold
sim_word, sim_value = sim_order[idx][mask], sim_value[mask]
sim_word = [idx2word[id] for id in sim_word]
sim_words.append(sim_word)
sim_values.append(sim_value)
return sim_words, sim_values
def prepare_data_for_eval(self, examples):
features = convert_examples_to_features(examples, self.labels, self.max_seq_length, self.tokenizer,
# xlnet has a cls token at the end
cls_token=self.tokenizer.cls_token,
cls_token_segment_id=0,
sep_token=self.tokenizer.sep_token,
pad_token=self.tokenizer.convert_tokens_to_ids([self.tokenizer.pad_token])[0],
pad_token_segment_id= 0,
pad_token_label_id=self.pad_token_label_id
)
# Convert to Tensors and build dataset
all_input_ids = torch.tensor([f.input_ids for f in features], dtype=torch.long)
all_input_mask = torch.tensor([f.input_mask for f in features], dtype=torch.long)
all_segment_ids = torch.tensor([f.segment_ids for f in features], dtype=torch.long)
all_label_ids = torch.tensor([f.label_ids for f in features], dtype=torch.long)
dataset = TensorDataset(all_input_ids, all_input_mask, all_segment_ids, all_label_ids)
return dataset
def read_examples_from_list(self, data, mode): ## this does not incorporate labels but just adds 'O's
guid_index = 0
examples = []
for s in data: #data is a list
words = []
labels = []
guid_index += 1
for num, w in enumerate(s):
words.append(w)
labels.append("O")
# print(len(words))
# print(words)
examples.append(InputExample(guid="%s-%d".format(mode, guid_index),
words=words,
labels=labels))
return examples
def read_examples_from_lists_wlables(self, data, labels, mode): ## this does not incorporate labels but just adds 'O's
guid_index = 0
examples = []
for s, l in zip(data, labels): #data is a list
words = []
labels = []
guid_index += 1
for num, w in enumerate(s):
words.append(w)
labels.append(l[num])
examples.append(InputExample(guid="%s-%d".format(mode, guid_index),
words=words,
labels=labels))
return examples
def evaluate(self, eval_dataset, model, tokenizer, labels, pad_token_label_id, mode = 'test', prefix=""):
eval_sampler = SequentialSampler(eval_dataset)
eval_dataloader = DataLoader(eval_dataset, sampler=eval_sampler, batch_size=32)
# Eval!
eval_loss = 0.0
nb_eval_steps = 0
preds = None
out_label_ids = None
model.eval()
for batch in tqdm(eval_dataloader, desc="Evaluating"):
# batch = tuple(t.to(args.device) for t in batch)
with torch.no_grad():
inputs = {"input_ids": batch[0],
"attention_mask": batch[1],
"token_type_ids": batch[2],
# XLM and RoBERTa don"t use segment_ids
"labels": batch[3]}
outputs = model(**inputs)
tmp_eval_loss, logits = outputs[:2]
# if args.n_gpu > 1:
# tmp_eval_loss = tmp_eval_loss.mean() # mean() to average on multi-gpu parallel evaluating
eval_loss += tmp_eval_loss.item()
nb_eval_steps += 1
if preds is None:
preds = logits.detach().cpu().numpy()
out_label_ids = inputs["labels"].detach().cpu().numpy()
else:
preds = np.append(preds, logits.detach().cpu().numpy(), axis=0)
out_label_ids = np.append(out_label_ids, inputs["labels"].detach().cpu().numpy(), axis=0)
eval_loss = eval_loss / nb_eval_steps
probs = preds
preds = np.argmax(preds, axis=2)
label_map = {i: label for i, label in enumerate(labels)}
out_label_list = [[] for _ in range(out_label_ids.shape[0])]
preds_list = [[] for _ in range(out_label_ids.shape[0])]
old_preds_list = preds_list
for i in range(out_label_ids.shape[0]):
for j in range(out_label_ids.shape[1]):
if out_label_ids[i, j] != pad_token_label_id:
out_label_list[i].append(label_map[out_label_ids[i][j]])
preds_list[i].append(label_map[preds[i][j]])
results = {
"loss": eval_loss,
"precision": precision_score(out_label_list, preds_list),
"recall": recall_score(out_label_list, preds_list),
"f1": f1_score(out_label_list, preds_list)
}
# logger.info("***** Eval results %s *****", prefix)
# for key in sorted(results.keys()):
# logger.info(" %s = %s", key, str(results[key]))
return results, preds_list, probs, preds, old_preds_list, out_label_list, label_map
def get_entities (self, pred_tags):
ent_ids = []
ent_tags = []
for num, i in enumerate (pred_tags):
temp = []
temp2 = []
if i.startswith('B-'):
temp.append(num)
temp2.append(i)
for x in range(1, 20):
try:
if pred_tags[num+x].startswith ('I-'):
temp.append(num+x)
temp2.append(pred_tags[num+x])
else:
break
except IndexError:
break
ent_ids.append(temp)
ent_tags.append(temp2)
else:
pass
return ent_ids, ent_tags
def calcwordscore(self, orig_label, orig_prob, leave_1_probs, nw_labels, ix2):
w = (orig_prob.max() - leave_1_probs[ix2][orig_prob.argmax()] + (int(nw_labels[ix2] != orig_label) * (
leave_1_probs[ix2].max() - orig_prob[leave_1_probs[ix2].argmax()])))
# if nw_labels[ix2] != orig_label:
# print(nw_labels[ix2])
return w
def calcwordscore_Itag(self, orig_label, orig_prob, leave_1_probs, nw_labels, ix2):
w = (orig_prob.max() - leave_1_probs[ix2][orig_prob.argmax()] + (int(nw_labels[ix2][2:] != orig_label[2:]) * (
leave_1_probs[ix2].max() - orig_prob[leave_1_probs[ix2].argmax()])))
return w
def calculate_importance_scores (self, ent, preds_listnw, probsnw, leave_1_texts, sent_allowed, allowed_ids):
mean_import_scores = []
all_import_scores = []
for ix in ent: ##for each word in entity we calculate importance of words. ix is hte index of hte word in the sentence
orig_label = preds_listnw[0][ix]
orig_prob = probsnw[0][ix+1]
leave_1_probs= []
nw_labels =[]
for i in range(1, len(leave_1_texts)):
[leave_1_probs.append(probsnw[i][ix+1])]##new probabilities of this word ##need to add 1 to compensate for padded token
try:
[nw_labels.append (preds_listnw[i][ix])] ##new lables of this word
except IndexError:
return 0
import_scores = []
nw_leave_1_texts = leave_1_texts[1:]
for ix2, word in enumerate(nw_leave_1_texts):
if orig_label.startswith('I'):
wordscore = self.calcwordscore_Itag(orig_label, orig_prob, leave_1_probs, nw_labels, ix2)
else:
wordscore = self.calcwordscore(orig_label, orig_prob, leave_1_probs, nw_labels, ix2)
## the last part says what the probability of the new label (if there is one) without hte word minus that same prob WITH the word
import_scores.append(wordscore)
all_import_scores.append(import_scores) ##list of lists
sum_import_scores = np.sum(all_import_scores, axis =0)
import_score_threshold=-1
words_perturb = []
for idx, score in sorted(enumerate(sum_import_scores), key=lambda x: x[1], reverse=True):
if score > import_score_threshold and sent_allowed[idx] not in self.stop_words_set:
words_perturb.append((allowed_ids[idx], sent_allowed[idx]))
# print(words_perturb)
return words_perturb
def random_ranking(self, ent, leave_1_texts, sent_allowed, allowed_ids):
words_perturb = []
nw_leave_1_texts = leave_1_texts[1:]
for idx, word in enumerate(nw_leave_1_texts):
if sent_allowed[idx] not in self.stop_words_set:
words_perturb.append((allowed_ids[idx], sent_allowed[idx]))
random.seed(1)
random.shuffle(words_perturb)
# print(words_perturb)
return words_perturb
def get_adversarial_examples_per_sent(self, sent, origlbl, predlbl, ent_tags, ent_ids, random_attack = False, sim_synonyms=0.5, sim_score_threshold=0.8, import_score_threshold = -1, sim_predictor = None, synonym_num=50,batch_size=32):
if sim_predictor == None:
sim_predictor == self.embed
out_texts = []
# ent_ids, ent_tags = get_entities(origlbl)
taboo_ids = [i for j in ent_ids for i in j]
len_text = len(sent)
sent_allowed = [i for num, i in enumerate(sent) if num not in taboo_ids]
allowed_ids = [num for num, i in enumerate(sent) if num not in taboo_ids]
# print(sent_allowed)
pos_ls = criteria.get_pos(sent)
for entnum, ent in enumerate(ent_ids): ##for each entity
success= 0
# print(ent)
# print(sent)
# print(origlbl)
leave_1_texts = []
leave_1_texts.append(sent)
for idword, word in enumerate(sent):
if idword not in taboo_ids:
# print(text_ls)
a_text = sent[:idword] + ['<oov>'] + sent[min(idword+1, len_text):] #until the end of sentence or id + 1
leave_1_texts.append(a_text)
##need to prepare for eval
examples = self.read_examples_from_list(leave_1_texts, 'test')
dataset = self.prepare_data_for_eval(examples)
resultsnw, preds_listnw, probsnw, predsnw, old_preds_listnw, true_labelsnw, label_map2= self.evaluate(dataset, self.model, self.tokenizer, self.labels, self.pad_token_label_id)
#this gives predictions for all words but we need to pick out hte relevant ones
# print(preds_listnw[0])
# print(preds_listnw[0] == predlbl)
mean_import_scores = []
all_import_scores = []
if random_attack == False:
words_perturb = self.calculate_importance_scores (ent, preds_listnw, probsnw, leave_1_texts, sent_allowed, allowed_ids)
else:
words_perturb = self.random_ranking(ent, leave_1_texts, sent_allowed, allowed_ids)
# print(words_perturb)
if words_perturb == 0: ##there are none
out = 'This sentence was a problem'
out_texts.append(out)
else:
# find synonyms
words_perturb_idx = [self.word2idx[word] for idx, word in words_perturb if word in self.word2idx]
synonym_words, _ = self.pick_most_similar_words_batch(words_perturb_idx, self.cos_sim, self.idx2word, synonym_num, sim_synonyms)
synonyms_all = []
for idx, word in words_perturb:
if word in self.word2idx:
synonyms = synonym_words.pop(0)
if synonyms:
synonyms_all.append((idx, synonyms))
# start replacing and attacking until label changes
text_prime = sent[:]
text_cache = text_prime[:]
num_changed = 0
unchanged_ent = ent
# print(len(synonyms_all))
if len(synonyms_all) == 0:
out2 = 'No synonyms'
out_texts.append(out2)
break
for idx, synonyms in synonyms_all:
# print(len(synonyms))
# finalsim = self.test_similarity(' '.join(text_cache),' '.join(text_prime))
# print(finalsim)
new_texts = [text_prime[:idx] + [synonym] + text_prime[min(idx + 1, len_text):] for synonym in synonyms]
# print(len(new_texts))
new_texts.append(sent)
examples = self.read_examples_from_list(new_texts, 'test')#
# print(new_texts)
dataset = self.prepare_data_for_eval(examples)
results2, preds_list2, probs2, preds2, old_preds_list2, true_labels2, label_map2nw= self.evaluate(dataset,self.model, self.tokenizer, self.labels, self.pad_token_label_id)
# compute semantic similarity
semantic_sims = []
for i in new_texts:
# print(i)
# print(text_cache)
sim = self.test_similarity(' '.join(text_cache),' '.join(i))
# print(sim)
semantic_sims.append(sim)
# print(semantic_sims)
if len(ent) == 1: # it is a single word entity
# print('Branch of the single word')
rel_probs=[]
new_probs_mask=[]
ix = ent[0]
correct = probs2[-1][ix+1].argmax()
for i in range(0, (len(new_texts)-1)):
[rel_probs.append(probs2[i][ix+1])]
new_texts2 = new_texts[:-1]
for ix2, t in enumerate(new_texts2):
r = rel_probs[ix2]
new_probs_mask.append(correct != r.argmax())
new_probs_mask2 = np.array(new_probs_mask)
semsims = semantic_sims[:-1]
semantic_sims2 = np.array(semsims)
synonyms_pos_ls = [criteria.get_pos(new_text[max(idx - 4, 0):idx + 5])[min(4, idx)]
if len(new_text) > 10 else criteria.get_pos(new_text)[idx] for new_text in new_texts2]
pos_ls = criteria.get_pos(sent)
new_probs_mask2 *= (semantic_sims2 >= sim_score_threshold)
# print(new_probs_mask2)
pos_mask = np.array(self.pos_filter(pos_ls[idx], synonyms_pos_ls))
new_probs_mask2 *= pos_mask
if np.sum(new_probs_mask2) > 0: ##there is an instance that changes the label
z = (new_probs_mask2 * semantic_sims2).argmax()
if semantic_sims2[z] < sim_score_threshold:
# print('This is below the threshold')
break
text_prime[idx] = synonyms[z]
num_changed += 1
# print(text_prime)
# print('We will stop')
success = 1
success_p = success/1
break
else: ##no label change at all - But if not, then we select the word with the least confidence score of label y as the best replacement word for wi
nw_rel_probs = [i[correct] for i in rel_probs]
nw_rel_probs2 = nw_rel_probs + (semantic_sims2 < sim_score_threshold) + (1 - pos_mask)
new_label_prob_min = nw_rel_probs2.min()
new_label_prob_argmin = nw_rel_probs2.argmin()
if semantic_sims2[new_label_prob_argmin] < sim_score_threshold:
# print('This is below the threshold')
break
orig_prob = probs2[-1][ix+1].max()
if new_label_prob_min < orig_prob:
text_prime[idx] = synonyms[new_label_prob_argmin]
# print(text_prime)
# print(synonyms[new_label_prob_argmin])
num_changed += 1
# print('We will go on')
# else:
# print('Probs were not high enough')
else: #it is a multiword entity -- unchanged ent is the part of hte entity that has not been changed.
# print('Branch of the multiword')
correctlbl = []
correctlblname = []
# print('the unchanged ents are now:')
# print(unchanged_ent)
for ix in unchanged_ent:
correctlbl.append(probs2[-1][ix+1].argmax())
correctlblname.append(preds_list2[-1][ix])
# print(correctlblname)
new_probs_mask_temp=[]
all_rel_probs = []
for number, ix in enumerate(unchanged_ent):
rel_probs=[]
new_probs_mask=[]
for i in range(0, (len(new_texts)-1)):
[rel_probs.append(probs2[i][ix+1])]
all_rel_probs.append(rel_probs)
correct = probs2[-1][ix+1].argmax()
new_texts2 = new_texts[:-1]
c = correctlblname[number]
if c.startswith('I'):
# print(c)
for ix2, t in enumerate(new_texts2):
pl = preds_list2[ix2][ix] ##predicted label
if pl[2:] == c[2:]: ##if the predicted label is either the B or I version of the originla label = no label change
new_probs_mask.append(False)
else:
new_probs_mask.append(True)
else:
for ix2, t in enumerate(new_texts2):
r = rel_probs[ix2]
new_probs_mask.append(correct != r.argmax()) ## true is label has been changed.
new_probs_mask_temp.append(np.array(new_probs_mask))
m = np.matrix(new_probs_mask_temp)
unchanged_lbls = [] ## contains hte indexes of hte words changed in the entity- not the ix in the sentence
# print(len(new_texts))
if len(new_texts) > 0: ## there are some synonyms
# print(len(new_texts))
# print(m)
for i in range(0, (len(new_texts)-1)):
# print(i)
try:
z = np.argwhere(m[:,i] == 0)[:,0]
unchanged_lbls.append(z.flatten().tolist())
except IndexError:
print(m)
z = []
unchanged_lbls.append(z)
# print('the unchanged labels for each synonym are:')
# print(unchanged_lbls)
new_probs_mask2 = np.sum(new_probs_mask_temp,axis = 0) ##true if any of the entity words have changed label with higher number for more label changes.
semsims = semantic_sims[:-1]
semantic_sims2 = np.array(semsims)
synonyms_pos_ls = [criteria.get_pos(new_text[max(idx - 4, 0):idx + 5])[min(4, idx)]
if len(new_text) > 10 else criteria.get_pos(new_text)[idx] for new_text in new_texts2]
pos_ls = criteria.get_pos(sent)
pos_mask = np.array(self.pos_filter(pos_ls[idx], synonyms_pos_ls))
new_probs_mask2 *= (semantic_sims2 >= sim_score_threshold)
new_probs_mask2 *= pos_mask
## are there any that change all labels? MAKE A MASK
entmax = len(unchanged_ent)
# w = np.argwhere(new_probs_mask2 == entmax)
# lstw = w.flatten().tolist()
# print('entmax: ')
# print(entmax)
new_probs_mask_all = [1 if x == entmax else 0 for x in new_probs_mask2]
# print(new_probs_mask_all)
# print(new_probs_mask2)
# print(np.sum(new_probs_mask2))
if np.sum(new_probs_mask_all) > 0: ## there are synonyms that can change all the labels! BEST SCENARIO
z= (new_probs_mask_all * semantic_sims2).argmax()
if semantic_sims2[z] < sim_score_threshold:
# print('This is below the threshold')
break
text_prime[idx] = synonyms[z]
num_changed += 1
# print(text_prime)
# print('We will stop')
success = 1
success_p = success/1
break
elif np.sum(new_probs_mask2) > 0: ##there is an instance that changes the label
winners = np.argwhere(new_probs_mask2 == np.amax(new_probs_mask2))
# print('some have changed')
lstwinners = winners.flatten().tolist() ##the ix of ones that make the highest number of words change but also conform to filters
winner_probs = []
for a in lstwinners:
u = unchanged_lbls[a]
##retrieve correct prob of unchanged labels to compare
rp = [i for num, i in enumerate(all_rel_probs) if num in u] ##ones for right entities
rp2 = [i[a] for i in rp] #get the ones for this synonym
cor = [i for num, i in enumerate(correctlbl) if num in u] ##ix of correct labels for relevant entities
rp_out = []
for a,b in zip(cor, rp2):
rp_out.append(np.array(b[a]))
rp_out2 = np.array(rp_out)
rp_sum = np.sum(rp_out2,axis =0) #summed confidence of all the relevant words in the entity
winner_probs.append(rp_sum)
##choose the lowest winner prob - lowest for other entities
winner_probs2 = np.array(winner_probs)
new_label_prob_min = winner_probs2.min()
new_label_prob_argmin = winner_probs2.argmin()
winning_ix = lstwinners[new_label_prob_argmin]
##change the unchanged ent for next iteration
u = unchanged_lbls[winning_ix]
nw_unchanged_ent = [i for num,i in enumerate(unchanged_ent) if num in u] ##actual indexes in sentences
unchanged_ent = nw_unchanged_ent
# print(unchanged_ent)
if semantic_sims2[winning_ix] < sim_score_threshold:
print('This is below the threshold')
break
# for ix in unchanged_ent:
# o = [probs2[-1][ix+1].max()]
# o2 = np.sum(np.array(o))
# if new_label_prob_min < o2:
text_prime[idx] = synonyms[winning_ix]
# print(text_prime)
num_changed += 1
# print('We have changed some but will continue')
success += 1
success_p = success/len(ent)
print(success_p)
text_cache2 = text_prime
if success_p == 1:
break
else: ##no label change at all - But if not, then we select the word with the least confidence score of label y as the best replacement word for wi
# print('no label change')
rp_out = []
for a,b in zip(correctlbl, all_rel_probs): #correct is the correct labels and all rel probs is the probabilities for hte wrods in the entities
rp = [x[a] for x in b]
rp_out.append(np.array(rp))
rp_sum = np.sum(rp_out,axis =0) #summed confidence of all the words in the entity
nw_rel_probs2 = rp_sum + (semantic_sims2 < sim_score_threshold) + (1 - pos_mask)
new_label_prob_min = nw_rel_probs2.min()
new_label_prob_argmin = nw_rel_probs2.argmin()
# print(semantic_sims2[new_label_prob_argmin])
# print(semantic_sims2.max())
if semantic_sims2[new_label_prob_argmin] < sim_score_threshold:
# print('This is below the threshold')
break
##get orig prob
for ix in unchanged_ent:
o = [probs2[-1][ix+1].max()]
o2 = np.sum(np.array(o))
if new_label_prob_min < o2:
text_prime[idx] = synonyms[new_label_prob_argmin]
num_changed += 1
##calculate semantic sim
try:
if 0<success_p<1:
# print('We are reverting!')
# print(text_prime)
text_prime= text_cache2
# print(text_cache2)
except UnboundLocalError:
pass
finalsim = self.test_similarity(' '.join(text_cache),' '.join(text_prime))
max_possible = len(synonyms_all)
if success == 0:
success_p = 0
out_texts.append(tuple([text_prime, num_changed, max_possible, success_p, finalsim]))
return out_texts
def retrieve_entities(self,devdata, traindata,testdata):
##first make a list of entities possible
alldata = pd.concat([devdata, traindata,testdata])
words = list(alldata['words'])
tags = list(alldata['ner'])
flattags = [i for j in tags for i in j]
flattags2 = [i[2:] for i in flattags]
tagset = set(flattags2)
taglist = [i for i in list(tagset) if i != '' and i != 'MISC']
print(taglist)
allent = []
ent_ids = []
ent_tags = []
for i in tags:
e1, e2 = self.get_entities(i)
ent_ids.append(e1)
ent_tags.append(e2)
for i in taglist: ##for each entity
temp = []
# print(i)
for a,b,c,d in zip(tags, words, ent_ids, ent_tags):
for num, t in enumerate(d):
if t[0][2:] == i: ##is it hte correct entity
l = []
ix = c[num] ##get the ids
[l.append(b[j]) for j in ix]
temp.append(l)
allent.append(temp)
return taglist, allent
# def test_get_adversarial_sent_entities (self, sent, origlbl, predlbl, ent_tags, ent_ids, taglst, allent, num_sample=50, sim_score_threshold=0.8, sim_predictor=None, batch_size=32):
# if sim_predictor == None:
# sim_predictor == self.embed
# out_texts = []
# # ent_idsold, ent_tagsold = get_entities(origlbl)
# # random.seed(1)
# len_text= len(sent)
# # print(ent_ids)
# for entnum, ent in enumerate(ent_ids):
# # print(ent)
# e = ent_tags[entnum]
# e1 = e[0][2:]
# # print(e1)
# try:
# corix = taglst.index(e1)
# correct_entlst = allent[corix]
# bcor = 'B-' + e1
# icor = 'I-' + e1
# # print(bcor)
# start = ent[0]
# end = ent[-1]
# idx = start
# sam = random.sample(correct_entlst, num_sample)
# print(sam)
# except ValueError:
# pass
def get_adversarial_sent_entities (self, sent, origlbl, predlbl, ent_tags, ent_ids, taglst, allent, num_sample=50, sim_score_threshold=0.8, sim_predictor=None, batch_size=32):
if sim_predictor == None:
sim_predictor == self.embed
out_texts = []
# ent_idsold, ent_tagsold = get_entities(origlbl)
# random.seed(1)
len_text= len(sent)
# print(ent_ids)
for entnum, ent in enumerate(ent_ids):
# print(ent)
e = ent_tags[entnum]
e1 = e[0][2:]
# print(e1)
try:
corix = taglst.index(e1)
correct_entlst = allent[corix]
bcor = 'B-' + e1
icor = 'I-' + e1
# print(bcor)
start = ent[0]
end = ent[-1]
idx = start
sam = random.sample(correct_entlst, num_sample)
comparewith = []
entidx = []
for syn in sam:
cw = [bcor]
if len(syn) > 1:
[cw.append(icor) for i in range(1, len(syn))]
comparewith.append(cw)
ei = [ent[0]]
if len(syn) > 1:
[ei.append(ent[0] + i) for i in range(1, len(syn))]
entidx.append(ei)
# print(sam)
text_prime = sent[:]
text_cache = text_prime[:]
# print(sam)
new_texts = [text_prime[:start] + syn + text_prime[end+1:] for syn in sam]
# print(new_texts)
examples = self.read_examples_from_list(new_texts, 'test')#
# print(new_texts)
dataset = self.prepare_data_for_eval(examples)
results2, preds_list2, probs2, preds2, old_preds_list2, true_labels2, label_map2nw= self.evaluate(dataset,self.model, self.tokenizer, self.labels, self.pad_token_label_id)
# compute semantic similarity
semantic_sims = []
for i in new_texts:
sim = self.test_similarity(' '.join(text_cache),' '.join(i))
semantic_sims.append(sim)
new_probs_mask = []
##make a mask of if anyone managed
for a, b, c in zip(comparewith, entidx, preds_list2):
temp = []
try:
z = [c[i] for i in b] # the prediction
except IndexError:
print(comparewith)
print(entidx)
print(preds_list2)
break
for num, j in enumerate(z): ## j is hte predicted label and a is the GOOD label
if j == a[num]: ##if the label did not change
temp.append(0)
if a[num] == icor and j == bcor: ##if the old lable was I and the new one is B
temp.append(0)
else:
temp.append(1) ## this word did change
t = np.sum(temp)/len(temp)
new_probs_mask.append(t) ##relative amount of entity that was changed.
new_probs_mask2 = np.array(new_probs_mask)
semantic_sims2 = np.array(semantic_sims)
#mask if similarity is too low
new_probs_mask2 *= (semantic_sims2 >= sim_score_threshold)
##did anyone manage? if yes output this.
if np.sum(new_probs_mask2) > 0: ##there is an instance that changes at least a part of a label
success = np.amax(new_probs_mask2)
new_probs_mask3 = [1 if x == success else 0 for x in new_probs_mask2]
chosen = sam[(new_probs_mask3 * semantic_sims2).argmax()]
# print(chosen)
text_done = text_prime[:start] + chosen + text_prime[end+1:]
success = 1
# print(text_done)
# print(text_cache)
finalsim = self.test_similarity(' '.join(text_cache),' '.join(text_done))
out_texts.append(tuple([text_done, success, finalsim]))
else:
success = 0 ##if not, output this.
chosen = random.choice(sam)
text_done = text_prime[:start] + chosen + text_prime[end+1:]
finalsim = self.test_similarity(' '.join(text_cache),' '.join(text_done))
out_texts.append(tuple([text_done, success, finalsim]))
except ValueError:#happens when a MISC value occurs
# print('the issue is this')
# out_texts.append('There is a ValueError here')
pass
return out_texts
def first_prediction(self, sents, true_labels):
examples = self.read_examples_from_lists_wlables(sents, true_labels, 'test')
dataset = self.prepare_data_for_eval(examples)
results, preds_list, probs, preds, old_preds_list, true_labelsx, label_map= self.evaluate(dataset, self.model, self.tokenizer, self.labels, self.pad_token_label_id)
return results, preds_list, probs, preds, old_preds_list, true_labelsx, label_map
def identify_unfit_sents (self, true_labels, sents):
only_ent = []
no_ent = []
pat = '[A-Za-z]+'
for num, origlbl in enumerate(true_labels):
ent_ids, ent_tags = self.get_entities(origlbl)
taboo_ids = [i for j in ent_ids for i in j]
sent = sents[num]
len_text = len(sent)
nw_txt = [i for num,i in enumerate(sent) if num not in taboo_ids]
nw_txt2 = [i.lower() for i in nw_txt]
filt_txt= [i for i in nw_txt2 if i not in self.stop_words_set]
filt_txt2 = [i for i in filt_txt if re.match(pat, i)]
if len(taboo_ids) == 0:
no_ent.append(1)
only_ent.append(0)
else:
no_ent.append(0)
if len(taboo_ids) == len_text:
only_ent.append(1)
elif len(filt_txt2) == 0:
only_ent.append(1)
else:
only_ent.append(0)
return only_ent, no_ent
def filter_sentences_correctness (self, true_labels, pred_labels):
correct_ent_ids = []
correct_ent_tags = []
only_wrong = []
for a,b in zip(true_labels, pred_labels):
a2ids, a2tags = self.get_entities(a) ##correct entities
c = 0 ##counter for how many correct
nw_entids= []
nw_enttags= []
for ent, enttag in zip(a2ids, a2tags): ##per correct entity check
c2 = 0
start = ent[0]
end = ent[-1]
if end == start:
b2 = b[start:(start+1)]
else:
b2 = b[start:(end+1)]
if b2 == enttag: ##definitely correct if the same
c =+1
c2 =+ 1
else:
btag = enttag[0]
s = set(b2)
if btag in s: #if the B-tag is correct then it is not completely wrong / missing
c =+ 1
c2 =+ 1
else:
pass
if c2 > 0:
nw_entids.append(ent)
nw_enttags.append(enttag)
if c == 0: ##no correct at all
only_wrong.append(1) ##there is no correct entity
# correct_ent_ids.append(None)
else:
only_wrong.append(0) ##there is at least one correct entity
correct_ent_ids.append(nw_entids)
correct_ent_tags.append(nw_enttags)
return only_wrong, correct_ent_ids, correct_ent_tags
def filter_unfit(self,true_labels, sents, ids, only_ent, no_ent):
remove = []
##remove unfit
for a,b in zip(only_ent, no_ent):
if a==1 or b==1:
remove.append(1)
else:
remove.append(0)
nw_sents = [i for num, i in enumerate(sents) if remove[num] == 0]
nw_true_labels = [i for num, i in enumerate(true_labels) if remove[num] == 0]
nw_ids = [i for num, i in enumerate(ids) if remove[num] == 0]
return nw_sents, nw_true_labels, nw_ids
def filter_unfit_r2 (self, true_labels, sents, ids, correct_ent_ids, correct_ent_tags, preds_list, only_wrong):
nw_sents = [i for num, i in enumerate(sents) if only_wrong[num] == 0]
nw_true_labels = [i for num, i in enumerate(true_labels) if only_wrong[num] == 0]
nw_ids = [i for num, i in enumerate(ids) if only_wrong[num] == 0]
nw_cor_ent_ids = [i for num, i in enumerate(correct_ent_ids) if only_wrong[num] == 0]
nw_cor_ent_tags = [i for num, i in enumerate(correct_ent_tags) if only_wrong[num] == 0 ]
nw_preds_list = [i for num, i in enumerate(preds_list) if only_wrong[num] == 0 ]
return nw_true_labels, nw_sents, nw_ids, nw_cor_ent_ids, nw_cor_ent_tags, nw_preds_list
def main(self, data, modelpath, savepath, savepath2, labelfile='labels.txt', devdata= None, traindata= None, alltestdata=None, random_attack = False, make_first_prediction= False, entities = False, sim_synonyms=0.5, sim_score_threshold= 0.8):
# print(sim_score_threshold)
self.initialize_essential(entities)
already_fully_wrong = [] #no need to break
not_fit_for_adversarial = [] # no words besides entities and stopwords
self.initialize_model(modelpath, labelfile)
##get sents and labels -- input data needs to be a aggregated df (grouped by id)
sents = list(data['words'])
true_labels = list(data['ner'])
try:
ids = list(data['id'])
except KeyError:
ids = list(data.index.values)
##first filter for if sentences either miss entities or miss changeable words - there is nothing to change
if entities == False:
only_ent, no_ent = self.identify_unfit_sents (true_labels, sents)
only_ent_ids = [i for num, i in enumerate(ids) if only_ent[num] == 1]
no_ent_ids = [i for num, i in enumerate(ids) if no_ent[num] == 1]
nw_sents, nw_true_labels, nw_ids = self.filter_unfit (true_labels, sents, ids, only_ent, no_ent)
# print(nw_sents[0])
else:
nw_sents = sents
nw_true_labels = true_labels
nw_ids = ids
only_ent_ids =[]
no_ent_ids = []
#make or cache first_prediction
if make_first_prediction== True:
print(self.labels)
results, preds_list, probs, preds, old_preds_list, true_labelsx, label_map = self.first_prediction(nw_sents, nw_true_labels)
self.save_obj(preds_list,'first_preds_list')
self.save_obj(probs,'first_probs')
else:
preds_list = self.load_obj('first_preds_list')
probs = self.load_obj('first_probs')
if entities == False:
sp1 = savepath2 + 'first_results_context'
save_obj(results, sp1)
sp2 = savepath2 + 'first_predictions_context'
save_obj(preds_list, sp2)
sp3 = savepath2 + 'first_probs_context'
save_obj(probs, sp3)
sp4 = savepath2 + 'only_ents'
save_obj(only_ent_ids, sp4)
if entities == True:
sp1 = savepath2 + 'first_results_entities'
save_obj(results, sp1)
sp2 = savepath2 + 'first_predictions_entities'
save_obj(preds_list, sp2)
sp3 = savepath2 + 'first_probs_entities'
save_obj(probs, sp3)
##second filter -do not need to change if they are already wrong (if some of entities are wrong only those are changed)
only_wrong, correct_ent_ids, correct_ent_tags = self.filter_sentences_correctness (nw_true_labels, preds_list)
only_wrong_ids = [i for num, i in enumerate(nw_ids) if only_wrong[num] == 1]
nwer_true_labels, nwer_sents, nwer_ids, correct_ent_ids, correct_ent_tags, nw_preds_list = self.filter_unfit_r2 (nw_true_labels, nw_sents, nw_ids, correct_ent_ids, correct_ent_tags, preds_list, only_wrong)
# print(len(nw_preds_list))
if entities == True:
sp1 = savepath2 + 'only_wrong_ids'
save_obj(only_wrong_ids, sp1)
out_texts = []
if entities == False:
if random_attack == False:
for num, sent in enumerate(nwer_sents):
print(sent)
##initialize
print(num +1)
print(len(nwer_sents))
predlbl = nw_preds_list[num]
origlbl = nwer_true_labels[num]
ent_tags = correct_ent_tags[num]
ent_ids = correct_ent_ids[num]
##run
out_text = self.get_adversarial_examples_per_sent(sent, origlbl, predlbl, ent_tags, ent_ids, sim_synonyms = 0.5, sim_score_threshold = 0.8)
# print(out_text)
out_texts.append(out_text)
save_obj(out_texts,savepath)
else:
for num, sent in enumerate(nwer_sents):
##initialize
print(num + 1)
print(len(nwer_sents))
predlbl = nw_preds_list[num]
origlbl = nwer_true_labels[num]
ent_tags = correct_ent_tags[num]
ent_ids = correct_ent_ids[num]
print(ent_tags)
##run
out_text = self.get_adversarial_examples_per_sent(sent, origlbl, predlbl, ent_tags, ent_ids, random_attack = True, sim_synonyms = sim_synonyms, sim_score_threshold = sim_score_threshold)
out_texts.append(out_text)
save_obj(out_texts,savepath)
if entities == True:
taglst, allent = self.retrieve_entities(devdata, traindata, alltestdata)
for num, sent in enumerate(nwer_sents):
##initialize
print(num + 1)
# if num > 10:
# break
print(sent)
predlbl = nw_preds_list[num]
origlbl = nwer_true_labels[num]
ent_tags = correct_ent_tags[num]
ent_ids = correct_ent_ids[num]
##run
out_text = self.get_adversarial_sent_entities (sent, origlbl, predlbl, ent_tags, ent_ids, taglst, allent, sim_score_threshold=sim_score_threshold)
out_texts.append(out_text)
save_obj(out_texts,savepath)
if entities == None:
# for num, sent in enumerate(nwer_sents):
# out_texts = []
return only_ent_ids, no_ent_ids, only_wrong_ids, nwer_sents, results
return only_ent_ids, no_ent_ids, only_wrong_ids, out_texts
# -
# # EXAMPLE USAGE
# +
##loading data
path = '/data/WNUT_AdversarialSample.tsv'
test = pd.read_csv(path, sep = '\t')
devdata= pd.read_csv('/data/WNUT_devdata.tsv', sep = '\t')
traindata = pd.read_csv('/data/WNUT_traindata.tsv', sep = '\t')
testdata = pd.read_csv('/data/WNUT_testdata.tsv', sep = '\t')
# +
# EXAMPLE OF ENTITY ATTACK -- entities argument set to True
modelpath = '/NER_data/WNUT/bert_e4_lr5/'
##The first save path is for the adversarial sentences
savepath = '/NER_data/WNUT/BERT1/output_bert1_entity'
##The second save path is other output
savepath2 = '/NER_data/WNUT/BERT1/'
only_ent_ids, no_ent_ids, only_wrong_ids, out_texts = AdversarialBERT().main(test, modelpath,savepath, savepath2, 'labels.txt', devdata2, traindata2, testdata2, random_attack =False, make_first_prediction= True, entities = True)
##only ent ids are those sentences with only entities and no other words -- not relevant for entity attack
## no ent ids are those sentences without any entities - EXCLUDED FROM ENTITY ATTACK
## only wrong ids are those sentences with only wrong predictions of entities - EXCLUDED FROM ENTITY ATTACK
## out texts are the provided adversarial sentences.
# +
# EXAMPLE CONTEXT ATTACK -- entities argument set to False
modelpath = '/NER_data/WNUT/bert_e4_lr5/'
##The first save path is for the adversarial sentences
savepath = '/NER_data/WNUT/BERT1/output_bert1_context'
##The second save path is other output
savepath2 = '/NER_data/WNUT/BERT1/'
only_ent_ids, no_ent_ids, only_wrong_ids, out_texts = AdversarialBERT().main(test, modelpath,savepath, savepath2, 'labels.txt', devdata, traindata, testdata, random_attack =False, make_first_prediction= True, entities = False)
##only ent ids are those sentences with only entities and no other words -- EXCLUDED FROM CONTEXT ATTACK
## no ent ids are those sentences without any entities - EXCLUDED FROM CONTEXT ATTACK
## only wrong ids are those sentences with only wrong predictions of entities - EXCLUDED FROM CONTEXT ATTACK
## out texts are the adversarial sentences.
|
src/AdversarialAttackNER.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ---
# title: coal crawling
# date: 2020-07-18
# category: coal
# tags:
# - web crawling
# - Selenium
# - coal
# - IoT
# toc: true
# ---
# ### 목표 데이터 정의
# ---
# 
# ### Selenium을 이용한 스크래핑
# ---
# +
#크롬의 해당 이미지 url 접속 코드
Search_term = 'Bituminous coal'
url = https://www.google.co.in/search?q=+search_term+"&tbm=isch“
browser = webdriver.Chrome('chromedriver.exe')
browser.get(url)
#스크롤 코드
browser.execute_script('window.scrollBy(0,10000)')
#해당 클래스 이름을 가진 요소 찾는 코드
browser.find_elements_by_class_name("rg_i")
#이미지로 저장하는 코드
el.screenshot(str(idx) + ".png")
# -
# ### 스크래핑 결과
# ---
# 
# ### Dataset
# ---
#
# 연갈탄과 유연탄 사진
#
# Trainset – 연갈탄 20개, 유연탄 120개
#
# Testset – 연갈탄 6개, 유연탄 14개
#
# validationset – 연갈탄 3개, 유연탄 7개
#
# ### 마치며
# ---
#
# 석탄 분류기 제조를 위해 연갈탄 및 유연탄 데이터를 수집 해 봤습니다.
#
# 읽어주셔서 감사합니다.
|
_ipynbs/.ipynb_checkpoints/coal_crawling-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
#Monthly, flex, walk-up, and staff annual passes used for round trips
monthlyRound = []
flexRound = []
walkRound = []
staffRound = []
#Monthly, flex, walk-up, and staff annual passes used for one-way trips
monthlyOne = []
flexOne = []
walkOne = []
staffOne = []
# -
# %matplotlib inline
df = pd.read_csv('metro-bike-share-trip-data.csv')
#Replaces spaces with _ and get rid of parenthesis
df.columns = df.columns.str.replace(' ', '_').str.replace('(', '').str.replace(')', '')
df["Trip_ID"].astype(int)
df["Duration"].astype(int)
df["Start_Time"].astype(str)
df["End_Time"].astype(str)
df["Starting_Station_ID"].astype(float)
df["Starting_Station_Latitude"].astype(float)
df["Starting_Station_Longitude"].astype(float)
df["Ending_Station_ID"].astype(float)
df["Ending_Station_Latitude"].astype(float)
df["Ending_Station_Longitude"].astype(float)
df["Bike_ID"].astype(float)
df["Trip_Route_Category"].astype(str)
df["Passholder_Type"].astype(str)
df["Starting_Lat-Long"].astype(str)
df["Ending_Lat-Long"].astype(str)
#Figuring out different values for Trip_Routes in df
df.Trip_Route_Category.unique()
#Figuring out different values for Passholder_Types in df
df.Passholder_Type.unique()
#For loop that iterates through rows, checking what pass is used for Round Trip or One-Way Trip
for index, row in df.iterrows():
passUsed = row["Passholder_Type"]
tripCategory = row["Trip_Route_Category"]
if passUsed == "Monthly Pass" and tripCategory == "Round Trip":
monthlyRound.append(passUsed)
elif passUsed == "Flex Pass" and tripCategory == "Round Trip":
flexRound.append(passUsed)
elif passUsed == "Walk-Up" and tripCategory == "Round Trip":
walkRound.append(passUsed)
elif passUsed == "Staff Annual" and tripCategory == "Round Trip":
staffRound.append(passUsed)
elif passUsed == "Monthly Pass" and tripCategory == "One Way":
monthlyOne.append(passUsed)
elif passUsed == "Flex Pass" and tripCategory == "One Way":
flexOne.append(passUsed)
elif passUsed == "Walk-Up" and tripCategory == "One Way":
walkOne.append(passUsed)
elif passUsed == "Staff Annual" and tripCategory == "One Way":
staffOne.append(passUsed)
# +
#Different types of Passholder types for Round Trip
print ("Round Trips Counts: ")
print ("Monthly passes count - {}".format(len(monthlyRound)))
print ("Flex passes count - {}".format(len(flexRound)))
print ("Walk-up passes count - {}".format(len(walkRound)))
print ("Staff Annual passes count - {}\n".format(len(staffRound)))
#Different types of Passholder types for One Way Trip
print ("One Way Trips Counts:")
print ("Monthly passes count - {}".format(len(monthlyOne)))
print ("Flex passes count - {}".format(len(flexOne)))
print ("Walk-up passes count - {}".format(len(walkOne)))
print ("Staff Annual passes count - {}\n".format(len(staffOne)))
|
data/Trip Route Category - PassHolder Type Combo Breakdown.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#Import libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import plotly.express as px
import seaborn as sns
from sklearn.preprocessing import PolynomialFeatures
from sklearn.pipeline import make_pipeline
from sklearn.linear_model import LinearRegression
from sklearn.linear_model import Ridge
from sklearn import preprocessing
from sklearn.model_selection import StratifiedKFold
from keras.models import Sequential
from keras.layers import Dense
pd.set_option('display.max_rows', None)
pd.set_option('display.max_columns', None)
# %matplotlib inline
# +
#Import global confirmed cases
data = pd.read_csv('COVIDGlobalConfirmed.txt')
data = pd.concat([data.iloc[:, 1:2], data.iloc[:, 4:]], axis = 1)
data = data.set_index('Country/Region')
#Calculate new cases from total cases
data2 = data.diff(axis = 1).iloc[:, 1:]
#Keep only regions with more than 500 total cases
data3 = data2[data2.sum(axis = 1) > 500]
#Remove cruise ship assuming not representative of normal spread
data3 = data3[data3.index != 'Diamond Princess']
#Reshape data for plotting
data4 = data3.T.reset_index()
data4['index'] = pd.to_datetime(data4['index'])
# +
#Create template for cleaning data
#Pull first country's data
c1 = pd.concat([data4['index'], data4.iloc[:, 1:2]], axis = 1)
#Add number to country name in case there are duplicates
c1['country'] = c1.columns[1] + str(0)
#Rename columns
c1.columns = ['date', 'new', 'country']
#Drop leading zeros up until first case
c1 = c1[c1['date'] >= c1[c1['new'] > 0]['date'].min()]
#Calculate the total previous to data point
c1['prevTtl'] = c1['new'].cumsum() - c1['new']
#Append new cases from day before
c1['prevNew'] = c1.new.shift(1)
#Calculate days from peak
c1['rel2Peak'] = c1['date'] - c1[c1['new'] == c1['new'].max()]['date'].min()
r2p = []
for i in range(len(c1['rel2Peak'])):
r = c1['rel2Peak'].iloc[i].days
r2p.append(r)
c1['rel2Peak'] = r2p
c1['peak'] = c1[c1['new'] == c1['new'].max()]['new'].values[0]
#Calculate peak prior to data point date
prevPeak = []
for i in range(len(c1)):
pP = c1[c1['date'] < c1.iloc[i:i+1, :]['date'][c1.iloc[i:i+1, :]['date'].index[0]]]['new'].max()
prevPeak.append(pP)
c1['prevPeak'] = prevPeak
#Reset and drop index
c1 = c1.reset_index(drop=True)
#Calculate days from prevPeak
r2P = [0]
for j in range(len(c1) - 1):
d = c1[c1['new'] == c1.iloc[j+1:j+2, :]['prevPeak'][j+1]]['date'].min()
d2 = c1.iloc[j+1:j+2, :]['date'] - d
r2P.append(d2[j+1].days)
c1['R2P'] = r2P
#Rolling average new cases
c1['avgNew3'] = c1.new.rolling(3).mean()
#Rolling sum new cases
c1['sumNew3'] = c1.new.rolling(3).sum()
#Rolling avg change in new
c1['avgChng3'] = c1['new'].diff().rolling(3).mean()
#Rolling average sum change in new
c1['sumChng3'] = c1['new'].diff().rolling(3).sum()
#Replace NaN values with zeros
c1 = c1.fillna(0)
#Clean and reshape data for plotting
forPlot = c1
for i in range(data4.shape[1] - 3):
c = pd.concat([data4['index'], data4.iloc[:, i+2:i+3]], axis = 1)
c['country'] = c.columns[1] + str(i+1)
c.columns = ['date', 'new', 'country']
c = c[c['date'] >= c[c['new'] > 0]['date'].min()]
c['prevTtl'] = c['new'].cumsum() - c['new']
c['prevNew'] = c.new.shift(1)
c['rel2Peak'] = c['date'] - c[c['new'] == c['new'].max()]['date'].min()
r2p = []
for k in range(len(c['rel2Peak'])):
r = c['rel2Peak'].iloc[k].days
r2p.append(r)
c['rel2Peak'] = r2p
c['peak'] = c[c['new'] == c['new'].max()]['new'].values[0]
prevPeak = []
for i in range(len(c)):
pP = c[c['date'] < c.iloc[i:i+1, :]['date'][c.iloc[i:i+1, :]['date'].index[0]]]['new'].max()
prevPeak.append(pP)
c['prevPeak'] = prevPeak
c = c.reset_index(drop=True)
#Calculate days from prevPeak
r2P = [0]
for j in range(len(c) - 1):
d = c[c['new'] == c.iloc[j+1:j+2, :]['prevPeak'][j+1]]['date'].min()
d2 = c.iloc[j+1:j+2, :]['date'] - d
r2P.append(d2[j+1].days)
c['R2P'] = r2P
c['avgNew3'] = c.new.rolling(3).mean()
c['sumNew3'] = c.new.rolling(3).sum()
c['avgChng3'] = c['new'].diff().rolling(3).mean()
c['sumChng3'] = c['new'].diff().rolling(3).sum()
c = c.fillna(0)
forPlot = pd.concat([forPlot, c])
forPlot = forPlot.reset_index()
forPlot.columns = ['day', 'date', 'new', 'country', 'prevTtl', 'prevNew', 'rel2Peak', 'peak', 'prevPeak', 'R2P', 'avgNew3', 'sumNew3', 'avgChng3', 'sumChng3']
# +
#Doing a visual check just to see what the genreal curve looks like
c = 'Algeria1'
#View a plot of the data
plt.figure(figsize=(20,10))
plt.scatter(forPlot[forPlot['country'] == c]['day'], forPlot[forPlot['country'] == c]['new'], color='navy', s=30, marker='o', label="training points")
X = pd.concat([forPlot[forPlot['country'] == c]['day'], forPlot[forPlot['country'] == c].iloc[:, 4:]], axis = 1)
y = forPlot[forPlot['country'] == c]['new']
interaction = PolynomialFeatures(degree=3, include_bias=False, interaction_only=True)
#X = interaction.fit_transform(X)
model = make_pipeline(PolynomialFeatures(5), Ridge())
model.fit(X['day'].values.reshape(-1, 1), np.array(y).reshape(-1, 1))
y_plot = model.predict(X['day'].values.reshape(-1, 1))
plt.plot(forPlot[forPlot['country'] == c]['day'], y_plot)
plt.legend(loc='lower left')
print(c)
plt.show()
# +
#Label data based on curve metrics
flatPrior = []
increasing = []
top = []
pastPeak = []
for fP in forPlot['country'].unique():
md = forPlot[forPlot['new'] > 1].groupby('country').min()['date']
f = len(forPlot[(forPlot['date'] < md[md.index == fP][0]) & (forPlot['country'] == fP)])
flatPrior.append(f)
p2P = forPlot[(forPlot.index < forPlot[(forPlot['country'] == fP) & (forPlot['rel2Peak'] == -1)].index[0]) & (forPlot['country'] == fP)]
i = len(p2P[p2P['date'] >= p2P[p2P['new'] > 1]['date'].min()])
increasing.append(i)
t = len(forPlot[(forPlot['country'] == fP) & (forPlot['rel2Peak'] >= -1) & (forPlot['rel2Peak'] <= 1)])
top.append(t)
p = len(forPlot[(forPlot['country'] == fP) & (forPlot['rel2Peak'] > 1)])
pastPeak.append(p)
targets = []
for n in range(105):
for r in range(flatPrior[n]):
targets.append([0, 0])
for up in range(increasing[n]):
targets.append([1, 1])
for s in range(top[n]):
targets.append([0, 1])
for p in range(pastPeak[n]):
targets.append([1, 0])
forPlot['targets'] = targets
#Add a single digit target for plotting purposes
targets2 = []
for n in range(105):
for r in range(flatPrior[n]):
targets2.append(0)
for up in range(increasing[n]):
targets2.append(1)
for s in range(top[n]):
targets2.append(2)
for p in range(pastPeak[n]):
targets2.append(3)
forPlot['targets2'] = targets2
# +
#Visual check to make sure labels were applied properly
c = 'Afghanistan0'
#View a plot of the data
fig = px.scatter(forPlot[forPlot['country'] == c], x='day', y='new', color = 'targets2', trendline="lowess")
fig.show()
# +
#new = forPlot['targets'].str.split(" ", n = 1, expand = True)
#forPlot["incline"]= new[0]
#forPlot["height"]= new[1]
#Shuffle and split x and y, test and train and scale
forPlot2 = forPlot.sample(frac=1)
n_train = int(len(forPlot)*0.8)
X = pd.concat([forPlot2['day'], forPlot2.iloc[:, 4:6], forPlot2.iloc[:, 8:13]], axis = 1)
y = forPlot2['targets'].reset_index(drop=True)
scalerX = preprocessing.StandardScaler()
X2 = scalerX.fit_transform(X)
y2 = []
for i in range(len(y)):
y2.append(np.array(y[i]).reshape(-1, 1))
y3 = np.array(y2).reshape(5743, 2)
X_train = X2[:n_train]
X_test = X2[n_train:]
y_train = np.array(y2[:n_train]).reshape(4594, 2)
y_test = np.array(y2[n_train:]).reshape(1149, 2)
# +
#Build model
model = Sequential()
model.add(Dense(12, input_dim=8, activation='relu'))
model.add(Dense(8, activation='relu'))
model.add(Dense(2, activation='sigmoid'))
#Compile
model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
#Fit model
model.fit(X_train, y_train, epochs=150, batch_size=10)
# +
#Generate predictions for spot tests
preds = model.predict(X_train)
#Evaluate model on unseen data
scores = model.evaluate(X_test, y_test, verbose=0)
print("%s: %.2f%%" % (model.metrics_names[1], scores[1]*100))
# -
|
Classifying COVID Status.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="UrjQGgr5nUHC"
# <h1> Imports and Set-Up
# + id="eGl9mcc0nOMP"
# !pip3 install higher
import matplotlib.pyplot as plt
import numpy as np
import torch.nn as nn
import torch
from torch.autograd import Variable
import random
from higher import innerloop_ctx
import warnings
#The code includes extensive warnings when run so have used this to ignore them
warnings.filterwarnings("ignore")
#Set random seeds for reproducibility of results
torch.manual_seed(0)
random.seed(0)
np.random.seed(0)
# set GPU or CPU depending on available hardware
# help from: https://stackoverflow.com/questions/46704352/porting-pytorch-code-from-cpu-to-gpu
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
print(f"Available device: {device}")
if device == "cuda:0":
# set default so all tensors are on GPU, if available
# help from: https://stackoverflow.com/questions/46704352/porting-pytorch-code-from-cpu-to-gpu
torch.set_default_tensor_type('torch.cuda.FloatTensor')
domain_type = "multidim_sine"
# + [markdown] id="T3KVOwFXFOY0"
# <h1> Data Loading and Generation
# + [markdown] id="nMUUm70ufKHH"
# This Sine function generator is based on the repostory: https://github.com/AdrienLE/ANIML/blob/master/ANIML.ipynb
# + id="a3X51uGHDvSV"
class SineWaveTask_multi:
def __init__(self,dimensions=20):
self.dimensions = dimensions
self.a = []
self.b = []
for dim in range(self.dimensions):
self.a.append(np.random.uniform(0.1, 5.0))
self.b.append(np.random.uniform(0, 2*np.pi))
self.train_x = None
def f(self, x,a,b):
return a * np.sin(x + b)
def training_set(self, size=10, force_new=False):
if self.train_x is None and not force_new:
self.train_x = np.random.uniform(-5, 5, size)
x = self.train_x
elif not force_new:
x = self.train_x
else:
x = np.random.uniform(-5, 5, size)
y = self.f(x,self.a[0],self.b[0])[:,None]
for dim in range(self.dimensions-1):
y = np.concatenate((y,self.f(x,self.a[dim+1],self.b[dim+1])[:,None]),axis=-1)
return torch.Tensor(x[:,None]), torch.Tensor(y)
def test_set(self, size=50):
x = np.linspace(-5, 5, size)
y = self.f(x,self.a[0],self.b[0])[:,None]
for dim in range(self.dimensions-1):
y = np.concatenate((y,self.f(x,self.a[dim+1],self.b[dim+1])[:,None]),axis=-1)
return torch.Tensor(x[:,None]), torch.Tensor(y)
TRAIN_SIZE = 20000
TEST_SIZE = 1000
SINE_TRAIN = [SineWaveTask_multi() for _ in range(TRAIN_SIZE)]
SINE_TEST = [SineWaveTask_multi() for _ in range(TEST_SIZE)]
# + id="<KEY>"
x, y_true = SINE_TRAIN[0].training_set()
# + colab={"base_uri": "https://localhost:8080/"} id="jInHQnwIKKxH" outputId="8760f64a-0585-432f-fa26-e9189ee9a6dd"
y_true.shape
# + [markdown] id="cu4urLF7Q88A"
# <h1> Neural Network Model
# + id="R1B0YTz6ytyN"
# Define network
class Neural_Network_multi(nn.Module):
def __init__(self, input_size=1, hidden_size=40, output_size=20):
super(Neural_Network_multi, self).__init__()
# network layers
self.hidden1 = nn.Linear(input_size,hidden_size)
self.hidden2 = nn.Linear(hidden_size,hidden_size)
self.output_layer = nn.Linear(hidden_size,output_size)
#Activation functions
self.relu = nn.ReLU()
def forward(self, x):
x = self.hidden1(x)
x = self.relu(x)
x = self.hidden2(x)
x = self.relu(x)
x = self.output_layer(x)
y = x
return y
# + [markdown] id="G-ExWACxQ3mt"
# <h1> Helper functions
# + id="1zyNHFXdOnug"
# The Minimum Square Error is used to evaluate the difference between prediction and ground truth
criterion = nn.MSELoss()
def copy_existing_model(model):
# Function to copy an existing model
# We initialize a new model
new_model = Neural_Network_multi()
# Copy the previous model's parameters into the new model
new_model.load_state_dict(model.state_dict())
return new_model
def get_samples_in_good_format(wave, num_samples=10, force_new=False):
#This function is used to sample data from a wave
x, y_true = wave.training_set(size=num_samples, force_new=force_new)
# We add [:,None] to get the right dimensions to pass to the model: we want K x 1 (we have scalars inputs hence the x 1)
# Note that we convert everything torch tensors
x = torch.tensor(x)
y_true = torch.tensor(y_true)
return x.to(device),y_true.to(device)
def initialization_to_store_meta_losses():
# This function creates lists to store the meta losses
global store_train_loss_meta; store_train_loss_meta = []
global store_test_loss_meta; store_test_loss_meta = []
def test_set_validation(model,new_model,wave,lr_inner,k,store_test_loss_meta):
# This functions does not actually affect the main algorithm, it is just used to evaluate the new model
new_model = training(model, wave, lr_inner, k)
# Obtain the loss
loss = evaluation(new_model, wave)
# Store loss
store_test_loss_meta.append(loss)
def train_set_evaluation(new_model,wave,store_train_loss_meta):
loss = evaluation(new_model, wave)
store_train_loss_meta.append(loss)
def print_losses(epoch,store_train_loss_meta,store_test_loss_meta,printing_step=1000):
if epoch % printing_step == 0:
print(f'Epochh : {epoch}, Average Train Meta Loss : {np.mean(store_train_loss_meta)}, Average Test Meta Loss : {np.mean(store_test_loss_meta)}')
#This is based on the paper update rule, we calculate the difference between parameters and then this is used by the optimizer, rather than doing the update by hand
def reptile_parameter_update(model,new_model):
# Zip models for the loop
zip_models = zip(model.parameters(), new_model.parameters())
for parameter, new_parameter in zip_models:
if parameter.grad is None:
parameter.grad = torch.tensor(torch.zeros_like(parameter))
# Here we are adding the gradient that will later be used by the optimizer
parameter.grad.data.add_(parameter.data - new_parameter.data)
# Define commands in order needed for the metaupdate
# Note that if we change the order it doesn't behave the same
def metaoptimizer_update(metaoptimizer):
# Take step
metaoptimizer.step()
# Reset gradients
metaoptimizer.zero_grad()
def metaupdate(model,new_model,metaoptimizer):
# Combine the two previous functions into a single metaupdate function
# First we calculate the gradients
reptile_parameter_update(model,new_model)
# Use those gradients in the optimizer
metaoptimizer_update(metaoptimizer)
def evaluation(new_model, wave, num_samples=10, force_new=False, item = False):
# Get data
x, label = get_samples_in_good_format(wave,num_samples=num_samples, force_new=force_new)
# Make model prediction
prediction = new_model(x)
# Get loss
if item == True: #Depending on whether we need to return the loss value for storing or for backprop
loss = criterion(prediction,label).item()
else:
loss = criterion(prediction,label)
return loss
def training(model, wave, lr_k, k):
# Create new model which we will train on
new_model = copy_existing_model(model)
# Define new optimizer
koptimizer = torch.optim.SGD(new_model.parameters(), lr=lr_k)
# Update the model multiple times, note that k>1 (do not confuse k with K)
for i in range(k):
# Reset optimizer
koptimizer.zero_grad()
# Evaluate the model
loss = evaluation(new_model, wave, item = False)
# Backpropagate
loss.backward()
koptimizer.step()
return new_model
# for MAML -- see MAML cell for additional citations around structure inspiration
def task_specific_train_and_eval(model, T_i, inner_loop_optimizer, N=1):
#Description of the loop formulation from https://higher.readthedocs.io/en/latest/toplevel.html
with innerloop_ctx(model, inner_loop_optimizer, copy_initial_weights = False) as (fmodel,diffopt):
#get our input data and our label
x, label = get_samples_in_good_format(T_i,num_samples=num_samples, force_new= True)
per_step_loss = []
for _ in range(N):
#Get the task specific loss for our model
task_specifc_loss = criterion(fmodel(x), label)
#Step through the inner gradient
diffopt.step(task_specifc_loss)
per_step_loss.append(task_specifc_loss.item())
held_out_task_specific_loss = evaluation(fmodel, T_i, num_samples=num_samples, force_new=True)
return held_out_task_specific_loss, per_step_loss, fmodel
# + [markdown] id="-4Ps8P2IRCmF"
# <h1> Reptile
# + id="8ogpg_DHizlC"
#Define important variables
epochs = int(1e5) # number of epochs
lr_meta=0.001 # Learning rate for meta model (outer loop)
printing_step=1000 # how many epochs should we wait to print the loss
lr_k=0.01 # Internal learning rate
k=5 # Number of internal updates for each task
# Initializations
initialization_to_store_meta_losses()
model = Neural_Network_multi()
metaoptimizer = torch.optim.Adam(model.parameters(), lr=lr_meta)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="-4-zQWWKFt3s" outputId="747b2ee4-fc6a-487e-98a4-a35d0cf2f7b2"
# Training loop
for epoch in range(epochs):
# Sample a sine wave (Task from training data)
wave = random.sample(SINE_TRAIN, 1)
# Update model predefined number of times based on k
new_model = training(model, wave[0], lr_k, k)
# Evalaute the loss for the training data
train_set_evaluation(new_model,wave[0],store_train_loss_meta)
#Meta-update --> Get gradient for meta loop and update
metaupdate(model,new_model,metaoptimizer)
# Evalaute the loss for the test data
# Note that we need to sample the wave from the test data
wave = random.sample(SINE_TEST, 1)
test_set_validation(model,new_model,wave[0],lr_k,k,store_test_loss_meta)
# Print losses every 'printing_step' epochs
print_losses(epoch,store_train_loss_meta,store_test_loss_meta,printing_step)
# + [markdown] id="bQjoz6FYctJM"
# <h1> Few Shot learning with new meta-model
# + [markdown] id="m-SPUG5Bfpe9"
# The model performs good few shot learning
# + colab={"base_uri": "https://localhost:8080/", "height": 351} id="GY84TNs8JXVH" outputId="1f8f1545-9c7f-4c55-bd7a-520642859f4d"
wave = SineWaveTask_multi();
k_shot_updates = 4
initialization_to_store_meta_losses()
for shots in range(k_shot_updates):
new_model = training(model, wave, lr_k, shots)
train_set_evaluation(new_model,wave,store_train_loss_meta)
plt.plot(store_train_loss_meta,label = 'Loss')
plt.legend()
plt.xlabel('k shots')
# + [markdown] id="5lL1NN2OPBSD"
# ## Second-Order MAML
# +
'''
Handling computation graphs and second-order backprop help and partial inspiration from:
- https://discuss.pytorch.org/t/how-to-save-computation-graph-of-a-gradient/128286/2
- https://discuss.pytorch.org/t/when-do-i-use-create-graph-in-autograd-grad/32853/3
- https://lucainiaoge.github.io/download/PyTorch-create_graph-is-true_Tutorial_and_Example.pdf
- https://www.youtube.com/watch?v=IkDw22a8BDE
- https://discuss.pytorch.org/t/how-to-manually-update-network-parameters-while-keeping-track-of-its-computational-graph/131642/2
- https://discuss.pytorch.org/t/how-to-calculate-2nd-derivative-of-a-likelihood-function/15085/3
- https://pytorch.org/tutorials/recipes/recipes/zeroing_out_gradients.html
- https://higher.readthedocs.io/en/latest/toplevel.html
Neural network configuration and helper class functions copied directly from
-https://github.com/AdrienLE/ANIML/blob/master/ANIML.ipynb
Note, different ways to refer to the task-specific vs. meta/aggregate updates to the parameters
Sometimes called "inner" and "outer" loop, respectively
Here, refered to as "task_specific" and "agg"/meta" (the latter, for consistency w/ ocariz code)
'''
#Instantiate the model network
model = Neural_Network_multi()
# move to the current device (GPU or CPU)
# help from: https://stackoverflow.com/questions/46704352/porting-pytorch-code-from-cpu-to-gpu
model.to(device)
T = 25 # num tasks
N = 1 # number of inner loop steps (notation from: https://www.bayeswatch.com/2018/11/30/HTYM/)
num_samples = 10 # number of samples to draw from the task
lr_task_specific = 0.01 # task specific learning rate
lr_meta = 0.001 # meta-update learning rate
num_epochs = 10000#70001 #Number of iterations for outer loop
printing_step = 5000 # show log of loss every x epochs
#Used to store the validation losses
metaLosses = []
metaValLosses = []
#Meta-optimizer for the outer loop
meta_optimizer = torch.optim.Adam(model.parameters(), lr = lr_meta)
cosScheduler = torch.optim.lr_scheduler.CosineAnnealingLR(optimizer=meta_optimizer, T_max=num_epochs,
eta_min=0, verbose = False)
#Inner optimizer, we were doing this by hand previously
inner_loop_optimizer = torch.optim.SGD(model.parameters(), lr = lr_task_specific)
for epoch in range(num_epochs):
cosScheduler.step(epoch=epoch)
# store loss over all tasks to then do a large meta-level update of initial params
# idea/help from video: https://www.youtube.com/watch?v=IkDw22a8BDE
meta_loss = None
#Sample a new wave each time
waves = [SineWaveTask_multi() for _ in range(T)]
#Loop through all of the tasks
for i, T_i in enumerate(waves):
held_out_task_specific_loss, _, _ = task_specific_train_and_eval(model, T_i, inner_loop_optimizer, N)
if meta_loss is None:
meta_loss = held_out_task_specific_loss
else:
meta_loss += held_out_task_specific_loss
meta_optimizer.zero_grad()
meta_loss /= T
meta_loss.backward()
meta_optimizer.step()
metaLosses.append(meta_loss.item())
# validation
val_wave = SineWaveTask_multi() # our own addition -- can vary
val_loss, _, _ = task_specific_train_and_eval(model, val_wave, inner_loop_optimizer, N)
metaValLosses.append(val_loss.item())
if epoch % printing_step == 0:
print("Iter = ", epoch, " Current Loss", np.mean(metaLosses), " Val Loss: ", np.mean(metaValLosses))
# saving model help from:
# https://pytorch.org/tutorials/beginner/saving_loading_models.html
torch.save(model.state_dict(), f"{domain_type}_maml_model.pt")
# -
# <h1> Few Shot learning with new meta-model (MAML)
# +
# run k-shot to check how rapidly we are able to adapt to unseen tasks
# starting w/ a single unseen task
test_wave = SineWaveTask_multi()
num_k_shots = 10
# use model returned from earlier optimization
inner_loop_optimizer = torch.optim.SGD(model.parameters(), lr = lr_task_specific)
held_out_task_specific_loss, metaTrainLosses, _ = task_specific_train_and_eval(model, test_wave, inner_loop_optimizer, num_k_shots)
plt.plot(metaTrainLosses)
plt.xlim([0,num_k_shots])
# +
all_losses = []
num_eval = 100
num_k_shots = 10
for test_eval in range(num_eval):
test_wave = SineWaveTask_multi()
# use model returned from earlier optimization
inner_loop_optimizer = torch.optim.SGD(model.parameters(), lr = lr_task_specific)
held_out_task_specific_loss, metaTrainLosses, _ = task_specific_train_and_eval(model, test_wave, inner_loop_optimizer, num_k_shots)
all_losses.append(np.array(metaTrainLosses))
all_losses = np.array(all_losses)
np.save(f"maml_ca_multi_sine_{num_k_shots}.npy", all_losses)
fig, ax = plt.subplots(figsize=(8,4))
mean_loss = np.mean(all_losses, axis=0)
# confidence interval plotting help from: https://stackoverflow.com/questions/59747313/how-to-plot-confidence-interval-in-python
y = mean_loss
x = list(range(num_k_shots))
ci = 1.96 * np.std(all_losses, axis=0)**2/np.sqrt(len(y))
ax_size=16
title_size=18
ax.plot(x, y, linewidth=3, label=f"Mean Loss")
ax.fill_between(x, (y-ci), (y+ci), alpha=.5,label=f"95% CI")
ax.set_xlabel("Gradient Steps",fontsize=ax_size)
ax.set_ylabel("Mean Squared Error (MSE)",fontsize=ax_size)
ax.set_title("Sine Wave Regression: k-Shot Evaluation",fontsize=title_size)
ax.legend()#loc="upper right")
plt.savefig("sine_ca_wave_multidim_reg_kshot.png")
# -
analysis_steps = [0, 1, num_k_shots-1]
for analysis_step in analysis_steps:
print(f"Step: {analysis_step}, Error: {mean_loss[analysis_step]}, Var: {ci[analysis_step]}")
# +
## Second-Order MAML
# +
'''
Handling computation graphs and second-order backprop help and partial inspiration from:
- https://discuss.pytorch.org/t/how-to-save-computation-graph-of-a-gradient/128286/2
- https://discuss.pytorch.org/t/when-do-i-use-create-graph-in-autograd-grad/32853/3
- https://lucainiaoge.github.io/download/PyTorch-create_graph-is-true_Tutorial_and_Example.pdf
- https://www.youtube.com/watch?v=IkDw22a8BDE
- https://discuss.pytorch.org/t/how-to-manually-update-network-parameters-while-keeping-track-of-its-computational-graph/131642/2
- https://discuss.pytorch.org/t/how-to-calculate-2nd-derivative-of-a-likelihood-function/15085/3
- https://pytorch.org/tutorials/recipes/recipes/zeroing_out_gradients.html
- https://higher.readthedocs.io/en/latest/toplevel.html
Neural network configuration and helper class functions copied directly from
-https://github.com/AdrienLE/ANIML/blob/master/ANIML.ipynb
Note, different ways to refer to the task-specific vs. meta/aggregate updates to the parameters
Sometimes called "inner" and "outer" loop, respectively
Here, refered to as "task_specific" and "agg"/meta" (the latter, for consistency w/ ocariz code)
'''
#Instantiate the model network
model = Neural_Network_multi()
# move to the current device (GPU or CPU)
# help from: https://stackoverflow.com/questions/46704352/porting-pytorch-code-from-cpu-to-gpu
model.to(device)
T = 25 # num tasks
N = 1 # number of inner loop steps (notation from: https://www.bayeswatch.com/2018/11/30/HTYM/)
num_samples = 10 # number of samples to draw from the task
lr_task_specific = 0.01 # task specific learning rate
lr_meta = 0.001 # meta-update learning rate
num_epochs = 10000#70001 #Number of iterations for outer loop
printing_step = 5000 # show log of loss every x epochs
#Used to store the validation losses
metaLosses = []
metaValLosses = []
#Meta-optimizer for the outer loop
meta_optimizer = torch.optim.Adam(model.parameters(), lr = lr_meta)
#Inner optimizer, we were doing this by hand previously
inner_loop_optimizer = torch.optim.SGD(model.parameters(), lr = lr_task_specific)
for epoch in range(num_epochs):
# store loss over all tasks to then do a large meta-level update of initial params
# idea/help from video: https://www.youtube.com/watch?v=IkDw22a8BDE
meta_loss = None
#Sample a new wave each time
waves = [SineWaveTask_multi() for _ in range(T)]
#Loop through all of the tasks
for i, T_i in enumerate(waves):
held_out_task_specific_loss, _, _ = task_specific_train_and_eval(model, T_i, inner_loop_optimizer, N)
if meta_loss is None:
meta_loss = held_out_task_specific_loss
else:
meta_loss += held_out_task_specific_loss
meta_optimizer.zero_grad()
meta_loss /= T
meta_loss.backward()
meta_optimizer.step()
metaLosses.append(meta_loss.item())
# validation
val_wave = SineWaveTask_multi() # our own addition -- can vary
val_loss, _, _ = task_specific_train_and_eval(model, val_wave, inner_loop_optimizer, N)
metaValLosses.append(val_loss.item())
if epoch % printing_step == 0:
print("Iter = ", epoch, " Current Loss", np.mean(metaLosses), " Val Loss: ", np.mean(metaValLosses))
# saving model help from:
# https://pytorch.org/tutorials/beginner/saving_loading_models.html
torch.save(model.state_dict(), f"{domain_type}_maml_model.pt")
|
archive/Alg_Comparison/Domain_CA_multidimensional_sinewave_plot_explore.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from scipy import stats
import numpy as np
import pandas as pd
import os
def spearman(df_weekrank, df_monthrank, indicator):
"""
@param: indicator could only be "Sharp-Omega", "ASKSR-Value", "Riskiness"
@return: a rank correlation between the indicator in weekly order and monthly order
"""
week_rank = list(df_weekrank[indicator])
month_rank = list(df_monthrank[indicator])
#check whether the rank is invalid
# this part is only for riskiness R
indexes = []
for i in range(len(week_rank)):
if(week_rank[i] == '-' or month_rank[i] == '-'):
indexes.append(i)
for pos in sorted(indexes, reverse=True):
del week_rank[pos]
del month_rank[pos]
for i in range(len(month_rank)):
if isinstance(month_rank[i], str):
month_rank[i] = float(month_rank[i])
if isinstance(week_rank[i], str):
week_rank[i] = float(week_rank[i])
res = stats.spearmanr(week_rank, month_rank)
return res[0]
def kendalltau(df_weekrank, df_monthrank, indicator):
"""
@param: indicator could only be "Sharp-Omega", "ASKSR-Value", "Riskiness"
@return: a rank correlation between the indicator in weekly order and monthly order
"""
week_rank = list(df_weekrank[indicator])
month_rank = list(df_monthrank[indicator])
#check whether the rank is invalid
# this part is only for riskiness R
indexes = []
for i in range(len(week_rank)):
if(week_rank[i] == '-' or month_rank[i] == '-'):
indexes.append(i)
for pos in sorted(indexes, reverse=True):
del week_rank[pos]
del month_rank[pos]
for i in range(len(month_rank)):
if isinstance(month_rank[i], str):
month_rank[i] = float(month_rank[i])
if isinstance(week_rank[i], str):
week_rank[i] = float(week_rank[i])
res = stats.kendalltau(week_rank, month_rank)
return res[0]
if __name__ == '__main__':
work_dir = os.getcwd()
df_weekrank = pd.read_csv(os.path.join(work_dir, "ranking", "weekly_ranking.csv"), index_col = 0)
df_monthrank = pd.read_csv(os.path.join(work_dir, "ranking", "monthly_ranking.csv"), index_col = 0)
res1 = spearman(df_weekrank, df_monthrank, "Sharp-Omega")
res2 = kendalltau(df_weekrank, df_monthrank, "Sharp-Omega")
omega_res = [res1, res2]
omega_res
res1 = spearman(df_weekrank, df_monthrank, "ASKSR-Value")
res2 = kendalltau(df_weekrank, df_monthrank, "ASKSR-Value")
asksr_res = [res1, res2]
asksr_res
res1 = spearman(df_weekrank, df_monthrank, "Riskiness")
res2 = kendalltau(df_weekrank, df_monthrank, "Riskiness")
riskness_res = [res1, res2]
riskness_res
df_weekrank
# # Correlation among indicators
def spearman_cf(df):
"""
@param: df is either monthly data or weekly data
@return: a rank correlation between the indicators
"""
S_rank = list(df["S_Rank"])
A_rank = list(df["A_Rank"])
R_rank = list(df["R_Rank"])
#check whether the rank is invalid
# this part is only for riskiness R
indexes = []
for i in range(len(R_rank)):
if(R_rank[i] == '-'):
indexes.append(i)
for pos in sorted(indexes, reverse=True):
del S_rank[pos]
del A_rank[pos]
del R_rank[pos]
for i in range(len(S_rank)):
if isinstance(S_rank[i], str):
S_rank[i] = float(S_rank[i])
if isinstance(A_rank[i], str):
A_rank[i] = float(A_rank[i])
if isinstance(R_rank[i], str):
R_rank[i] = float(R_rank[i])
res1 = stats.spearmanr(S_rank, A_rank)
res2 = stats.spearmanr(S_rank, R_rank)
res3 = stats.spearmanr(R_rank, A_rank)
print("S/A", res1[0])
print()
print("S/R", res2[0])
print()
print("A/R", res3[0])
return res1[0], res2[0], res3[0]
def kendalltau_cf(df):
"""
@param: df is either monthly data or weekly data
@return: a rank correlation between the indicators
"""
S_rank = list(df["S_Rank"])
A_rank = list(df["A_Rank"])
R_rank = list(df["R_Rank"])
#check whether the rank is invalid
# this part is only for riskiness R
indexes = []
for i in range(len(R_rank)):
if(R_rank[i] == '-'):
indexes.append(i)
for pos in sorted(indexes, reverse=True):
del S_rank[pos]
del A_rank[pos]
del R_rank[pos]
for i in range(len(S_rank)):
if isinstance(S_rank[i], str):
S_rank[i] = float(S_rank[i])
if isinstance(A_rank[i], str):
A_rank[i] = float(A_rank[i])
if isinstance(R_rank[i], str):
R_rank[i] = float(R_rank[i])
res1 = stats.kendalltau(S_rank, A_rank)
res2 = stats.kendalltau(S_rank, R_rank)
res3 = stats.kendalltau(R_rank, A_rank)
print("S/A", res1[0])
print()
print("S/R", res2[0])
print()
print("A/R", res3[0])
return res1[0], res2[0], res3[0]
spearman_cf(df_weekrank)
kendalltau_cf(df_weekrank)
spearman_cf(df_monthrank)
kendalltau_cf(df_monthrank)
|
Rank_correlation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/standroidbeta/DS-Unit-1-Sprint-3-Data-Storytelling/blob/master/Unit%201%20Sprint%20Challenge%203%20/DS_Unit_1_Sprint_Challenge_3_Data_Storytelling.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="i-n_5en3ER1o"
# # Data Science Unit 1 Sprint Challenge 3
#
# # Data Storytelling
#
# In this sprint challenge you'll work with a dataset from **FiveThirtyEight's article, [Every Guest Jon Stewart Ever Had On ‘The Daily Show’](https://fivethirtyeight.com/features/every-guest-jon-stewart-ever-had-on-the-daily-show/)**!
# + [markdown] colab_type="text" id="Thm2n5FF2Fnp"
# # Part 0 — Run this starter code
#
# You don't need to add or change anything here. Just run this cell and it loads the data for you, into a dataframe named `df`.
#
# (You can explore the data if you want, but it's not required to pass the Sprint Challenge.)
# + colab_type="code" id="0rTHgzJIuRS7" colab={}
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
url = 'https://raw.githubusercontent.com/fivethirtyeight/data/master/daily-show-guests/daily_show_guests.csv'
df = pd.read_csv(url).rename(columns={'YEAR': 'Year', 'Raw_Guest_List': 'Guest'})
def get_occupation(group):
if group in ['Acting', 'Comedy', 'Musician']:
return 'Acting, Comedy & Music'
elif group in ['Media', 'media']:
return 'Media'
elif group in ['Government', 'Politician', 'Political Aide']:
return 'Government and Politics'
else:
return 'Other'
df['Occupation'] = df['Group'].apply(get_occupation)
# + [markdown] colab_type="text" id="OS0nW1vz1itX"
# # Part 1 — What's the breakdown of guests’ occupations per year?
#
# For example, in 1999, what percentage of guests were actors, comedians, or musicians? What percentage were in the media? What percentage were in politics? What percentage were from another occupation?
#
# Then, what about in 2000? In 2001? And so on, up through 2015.
#
# So, **for each year of _The Daily Show_, calculate the percentage of guests from each occupation:**
# - Acting, Comedy & Music
# - Government and Politics
# - Media
# - Other
#
# #### Hints:
# You can make a crosstab. (See pandas documentation for examples, explanation, and parameters.)
#
# You'll know you've calculated correctly when the percentage of "Acting, Comedy & Music" guests is 90.36% in 1999, and 45% in 2015.
#
# **Optional Bonus Challenge:** Do additional insightful data exploration.
# + colab_type="code" id="sRMc0H_5z6ff" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="e1db855f-1989-45a0-c22f-462aeffcd998"
df.info()
# + id="q0eXbt0ZCyIz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 607} outputId="f5c21fbc-b4f0-43c8-ca03-d7c1f3b4dc46"
ct = pd.crosstab(df['Year'], df['Occupation'])
ct
# + id="RUFYjUi1JWDd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 607} outputId="bf537eb0-d573-49e2-f668-4f9f223dd9e0"
percents = ct.div(ct.sum(axis=1), axis=0)
percents
# + id="AfZrdpHE_DnZ" colab_type="code" colab={}
percents = percents.drop(columns=['Other'])
# + id="a4lPEpEzLFWn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 418} outputId="77f36ad7-e50a-4652-84b3-ff2100809247"
formated = percents.style.format("{:.2%}")
formated
# + id="7-fvFEfebeKE" colab_type="code" colab={}
# + [markdown] colab_type="text" id="Nqf9oJJDDu-d"
# # Part 2 — Recreate this explanatory visualization:
# + colab_type="code" id="scozkHQc0_eD" outputId="a8fb111c-8b29-49e9-e8a0-aaa56653c2bd" colab={"base_uri": "https://localhost:8080/", "height": 406}
from IPython.display import display, Image
png = 'https://fivethirtyeight.com/wp-content/uploads/2015/08/hickey-datalab-dailyshow.png'
example = Image(png, width=500)
display(example)
# + [markdown] colab_type="text" id="W7lw3JzAE6BJ"
# **Hints:**
# - You can choose any Python visualization library you want. I've verified the plot can be reproduced with matplotlib, pandas plot, or seaborn. I assume other libraries like altair or plotly would work too.
# - If you choose to use seaborn, you may want to upgrade the version to 0.9.0.
#
# **Expectations:** Your plot should include:
# - 3 lines visualizing "occupation of guests, by year." The shapes of the lines should look roughly identical to 538's example. Each line should be a different color. (But you don't need to use the _same_ colors as 538.)
# - Legend or labels for the lines. (But you don't need each label positioned next to its line or colored like 538.)
# - Title in the upper left: _"Who Got To Be On 'The Daily Show'?"_ with more visual emphasis than the subtitle. (Bolder and/or larger font.)
# - Subtitle underneath the title: _"Occupation of guests, by year"_
#
# **Optional Bonus Challenge:**
# - Give your plot polished aesthetics, with improved resemblance to the 538 example.
# - Any visual element not specifically mentioned in the expectations is an optional bonus.
# + colab_type="code" id="E8XBAr8rz_Na" colab={"base_uri": "https://localhost:8080/", "height": 342} outputId="f516776b-ee9b-4d8e-e732-089e410f3d34"
plt.style.use('fivethirtyeight')
guests_graph = percents.plot();
plt.title("Who Got To Be On 'The Daily Show'?",
x=-.1, y=1.08, fontsize=18, fontweight='bold', loc='left')
# guests_graph.set(xticks=range(0, 4, 2015)
# guests_graph.text(x = 0, y = 3, s = "Who Got To Be On 'The Daily Show'?",
# fontsize = 18, weight = 'bold')
plt.suptitle('Occupation of Guests, by year', x=0, y=.95, fontsize =16, horizontalalignment='left');
# + [markdown] colab_type="text" id="LuacMjSf2ses"
# # Part 3 — Who were the top 10 guests on _The Daily Show_?
#
# **Make a plot** that shows their names and number of appearances.
#
# **Add a title** of your choice.
#
# **Expectations:** It's ok to make a simple, quick plot: exploratory, instead of explanatory.
#
# **Optional Bonus Challenge:** You can change aesthetics and add more annotation. For example, in a relevant location, could you add the text "19" to show that <NAME> appeared 19 times on _The Daily Show_? (And so on, for each of the top 10 guests.)
# + colab_type="code" id="tbwfBN3HsFlh" colab={}
appearances = df.drop(columns=['GoogleKnowlege_Occupation', 'Show', 'Group','Occupation'])
appearances.head(10)
# + id="wg3nEZwrqo17" colab_type="code" colab={}
num_appearances = appearances['Guest'].value_counts()
top_ten = num_appearances.head(10).sort_values(ascending=False)
top_ten
# + id="GCTaqnVzsOBt" colab_type="code" colab={}
top_ten.plot.bar()
plt.title("Who were the top 10 guests on 'The Daily Show'?", fontweight='bold');
|
Unit 1 Sprint Challenge 3 /DS_Unit_1_Sprint_Challenge_3_Data_Storytelling.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Activity 5: Assembling a Deep Learning System
# In this activity, we will train the first version of our LSTM model using Bitcoin daily closing prices. These prices will be organized using the weeks of both 2016 and 2017. We do that because we are interested in predicting the prices of a week's worth of trading.
#
# Let's go ahead and import our data.
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
from tensorflow.keras.models import load_model
plt.style.use('seaborn-white')
# ### Shaping Data
# Neural networks typically work with vectors and tensors, both mathematical objects that organize data in a number of dimensions.
train = pd.read_csv('data/train_dataset.csv')
train.head()
# LSTM networks require vectors with three dimensions. These dimensions are:
#
# * **Period length**: The period length, i.e. how many observations is there on a period.
# * **Number of periods**: How many periods are available in the dataset.
# * **Number of features**: Number of features available in the dataset.
#
# We will create weekly groups then rearrange the resulting array to match those dimensions.
def create_groups(data, group_size=7):
"""
Creates distinct groups from a given continuous series.
Parameters
----------
data: np.array
Series of continious observations.
group_size: int, default 7
Determines how large the groups are. That is,
how many observations each group contains.
Returns
-------
A Numpy array object.
"""
samples = list()
for i in range(0, len(data), group_size):
sample = list(data[i:i + group_size])
if len(sample) == group_size:
samples.append(np.array(sample).reshape(1, group_size).tolist())
return np.array(samples)
data = create_groups(train['close_point_relative_normalization'].values, 7)
len(data)
X_train = data[:-1,:].reshape(1, 186, 7)
Y_validation = data[-1].reshape(1, 7)
# ### Load Our Model
# Let's start by loading our previously trained model.
model = load_model('bitcoin_lstm_v0.h5')
# ### Make Predictions
#
# %%time
history = model.fit(
x=X_train, y=Y_validation,
epochs=100)
model.save('bitcoin_lstm_v0_trained.h5')
pd.Series(history.history['loss']).plot(linewidth=2, figsize=(14, 4), color='#d35400')
def denormalize(series, last_value):
result = last_value * (series + 1)
return result
predictions = model.predict(x=X_train)[0]
last_weeks_value = train[train['date'] == train['date'].max()]['close'].values[0]
denormalized_prediction = denormalize(predictions, last_weeks_value)
pd.DataFrame(denormalized_prediction).plot(linewidth=2, figsize=(6, 4), color='#d35400', grid=True)
full_series = list(train['close'].values) + list(denormalized_prediction)
pd.DataFrame(full_series[-7*8:]).plot(linewidth=2, figsize=(14, 4), color='#d35400', grid=True)
plt.axvline(len(full_series[-7*8:]) - 7)
# ### Summary
# In this activity, we have assembled a complete deep learning system: from data to prediction. The model created in this activity need a number of improvements before it can be considered useful. However, it serves as a great starting point from which we will continuously improve.
|
old/Chapter02/activity_5/Activity_5_Assembling_a_Deep_Learning_System.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib.pyplot as plt
import numpy as np
import pickle as pkl
get_ipython().magic('matplotlib inline')
# -
import torch
from torchvision import datasets
from torchvision import transforms
# +
transform = transforms.Compose([
transforms.Resize(32),
transforms.ToTensor(),
])
dataset = datasets.ImageFolder('pokemon',transform=transform)
batch_size = 128
num_workers = 4
dataloader = torch.utils.data.DataLoader(dataset,batch_size=batch_size,num_workers=num_workers,shuffle=True)
# -
dataiter = iter(dataloader)
images, labels = dataiter.next()
fig = plt.figure(figsize=(25, 4))
plot_size=20
for idx in np.arange(plot_size):
ax = fig.add_subplot(2, plot_size/2, idx+1, xticks=[], yticks=[])
ax.imshow(np.transpose(images[idx], (1, 2, 0)))
ax.set_title(str(labels[idx].item()))
# +
img = images[0]
print('Min: ', img.min())
print('Max: ', img.max())
print(img.shape)
# -
def scale(x, feature_range=(-1, 1)):
''' Scale takes in an image x and returns that image, scaled
with a feature_range of pixel values from -1 to 1.
This function assumes that the input x is already scaled from 0-1.'''
minq,maxq = feature_range
x = x * (maxq-minq) + minq
return x
# +
scaled_img = scale(img)
print('Scaled min: ', scaled_img.min())
print('Scaled max: ', scaled_img.max())
# +
import torch.nn as nn
import torch.nn.functional as F
# helper conv function
def conv(in_channels, out_channels, kernel_size, stride=2, padding=1, batch_norm=True):
"""Creates a convolutional layer, with optional batch normalization.
"""
layers = []
conv_layer = nn.Conv2d(in_channels, out_channels,
kernel_size, stride, padding, bias=False)
# append conv layer
layers.append(conv_layer)
if batch_norm:
# append batchnorm layer
layers.append(nn.BatchNorm2d(out_channels))
# using Sequential container
return nn.Sequential(*layers)
# -
class Discriminator(nn.Module):
def __init__(self, conv_dim=32):
super(Discriminator, self).__init__()
self.conv_dim = conv_dim
self.layer_1 = conv(3,conv_dim,4,batch_norm=False)
self.layer_2 = conv(conv_dim,conv_dim*2,4,batch_norm=True)
self.layer_3 = conv(conv_dim*2,conv_dim*4,4,batch_norm=True)
self.fc = nn.Linear(4*4*conv_dim*4,1)
def forward(self, x):
x = F.leaky_relu(self.layer_1(x),0.2)
x = F.leaky_relu(self.layer_2(x),0.2)
x = F.leaky_relu(self.layer_3(x),0.2)
x = x.view(-1,4*4*self.conv_dim*4)
x = self.fc(x)
return x
def deconv(in_channels, out_channels, kernel_size, stride=2, padding=1, batch_norm=True):
"""Creates a transposed-convolutional layer, with optional batch normalization.
"""
layers = []
layers.append(nn.ConvTranspose2d(in_channels,out_channels,kernel_size,stride,padding,bias = False))
if(batch_norm):
layers.append(nn.BatchNorm2d(out_channels))
return nn.Sequential(*layers)
class Generator(nn.Module):
def __init__(self, z_size, conv_dim=32):
super(Generator, self).__init__()
self.conv_dim = conv_dim
self.fc = nn.Linear(z_size,conv_dim*4*4*4)
self.t_conv1 = deconv(conv_dim*4,conv_dim*2,4)
self.t_conv2 = deconv(conv_dim*2,conv_dim,4)
self.t_conv3 = deconv(conv_dim,3,4,batch_norm=False)
def forward(self, x):
x = self.fc(x)
x = x.view(-1,4*self.conv_dim,4,4)
x = F.leaky_relu(self.t_conv1(x))
x = F.leaky_relu(self.t_conv2(x))
x = self.t_conv3(x)
x = F.tanh(x)
return x
# +
conv_dim = 32
z_size = 100
D = Discriminator(conv_dim)
G = Generator(z_size=z_size, conv_dim=conv_dim)
print(D)
print()
print(G)
# -
train_on_gpu = torch.cuda.is_available()
if train_on_gpu:
# move models to GPU
G.cuda()
D.cuda()
print('GPU available for training. Models moved to GPU')
else:
print('Training on CPU.')
# +
def real_loss(D_out, smooth=False):
batch_size = D_out.size(0)
# label smoothing
if smooth:
# smooth, real labels = 0.9
labels = torch.ones(batch_size)*0.9
else:
labels = torch.ones(batch_size) # real labels = 1
# move labels to GPU if available
if train_on_gpu:
labels = labels.cuda()
# binary cross entropy with logits loss
criterion = nn.BCEWithLogitsLoss()
# calculate loss
loss = criterion(D_out.squeeze(), labels)
return loss
def fake_loss(D_out):
batch_size = D_out.size(0)
labels = torch.zeros(batch_size) # fake labels = 0
if train_on_gpu:
labels = labels.cuda()
criterion = nn.BCEWithLogitsLoss()
# calculate loss
loss = criterion(D_out.squeeze(), labels)
return loss
# +
import torch.optim as optim
# params
lr = 0.0002
beta1=0.5
beta2=0.999
# Create optimizers for the discriminator and generator
d_optimizer = optim.Adam(D.parameters(), lr, [beta1, beta2])
g_optimizer = optim.Adam(G.parameters(), lr, [beta1, beta2])
# +
import pickle as pkl
# training hyperparams
num_epochs = 500
# keep track of loss and generated, "fake" samples
samples = []
losses = []
print_every = 300
# Get some fixed data for sampling. These are images that are held
# constant throughout training, and allow us to inspect the model's performance
sample_size=16
fixed_z = np.random.uniform(-1, 1, size=(sample_size, z_size))
fixed_z = torch.from_numpy(fixed_z).float()
# train the network
for epoch in range(num_epochs):
for batch_i, (real_images, _) in enumerate(dataloader):
batch_size = real_images.size(0)
# important rescaling step
real_images = scale(real_images)
# ============================================
# TRAIN THE DISCRIMINATOR
# ============================================
d_optimizer.zero_grad()
# 1. Train with real images
# Compute the discriminator losses on real images
if train_on_gpu:
real_images = real_images.cuda()
D_real = D(real_images)
d_real_loss = real_loss(D_real)
# 2. Train with fake images
# Generate fake images
z = np.random.uniform(-1, 1, size=(batch_size, z_size))
z = torch.from_numpy(z).float()
# move x to GPU, if available
if train_on_gpu:
z = z.cuda()
fake_images = G(z)
# Compute the discriminator losses on fake images
D_fake = D(fake_images)
d_fake_loss = fake_loss(D_fake)
# add up loss and perform backprop
d_loss = d_real_loss + d_fake_loss
d_loss.backward()
d_optimizer.step()
# =========================================
# TRAIN THE GENERATOR
# =========================================
g_optimizer.zero_grad()
# 1. Train with fake images and flipped labels
# Generate fake images
z = np.random.uniform(-1, 1, size=(batch_size, z_size))
z = torch.from_numpy(z).float()
if train_on_gpu:
z = z.cuda()
fake_images = G(z)
# Compute the discriminator losses on fake images
# using flipped labels!
D_fake = D(fake_images)
g_loss = real_loss(D_fake) # use real loss to flip labels
# perform backprop
g_loss.backward()
g_optimizer.step()
# Print some loss stats
if batch_i % print_every == 0:
# append discriminator loss and generator loss
losses.append((d_loss.item(), g_loss.item()))
# print discriminator and generator loss
print('Epoch [{:5d}/{:5d}] | d_loss: {:6.4f} | g_loss: {:6.4f}'.format(
epoch+1, num_epochs, d_loss.item(), g_loss.item()))
## AFTER EACH EPOCH##
# generate and save sample, fake images
G.eval() # for generating samples
if train_on_gpu:
fixed_z = fixed_z.cuda()
samples_z = G(fixed_z)
samples.append(samples_z)
G.train() # back to training mode
# Save training generator samples
with open('train_samples.pkl', 'wb') as f:
pkl.dump(samples, f)
# -
fig, ax = plt.subplots()
losses = np.array(losses)
plt.plot(losses.T[0], label='Discriminator', alpha=0.5)
plt.plot(losses.T[1], label='Generator', alpha=0.5)
plt.title("Training Losses")
plt.legend()
def view_samples(epoch, samples):
fig, axes = plt.subplots(figsize=(16,4), nrows=2, ncols=8, sharey=True, sharex=True)
for ax, img in zip(axes.flatten(), samples[epoch]):
img = img.detach().cpu().numpy()
img = np.transpose(img, (1, 2, 0))
img = ((img +1)*255 / (2)).astype(np.uint8) # rescale to pixel range (0-255)
ax.xaxis.set_visible(False)
ax.yaxis.set_visible(False)
im = ax.imshow(img.reshape((32,32,3)))
_ = view_samples(-1, samples)
|
dcgan_pokemon.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 6.4: データの関係の可視化
# +
# リスト 6.4.1 tips データセットの読み込み
import seaborn as sns
tips = sns.load_dataset("tips")
tips.head()
# +
# リスト 6.4.2 relplot() 関数で描画した散布図
sns.relplot(data=tips, x="total_bill", y="tip", kind="scatter")
# +
# リスト 6.4.3 time 列で色分けした散布図
sns.relplot(data=tips, x="total_bill", y="tip", hue="time")
# +
# リスト 6.4.4 day 列でマーカーを分けた散布図
sns.relplot(data=tips, x="total_bill", y="tip", hue="time", style="day")
# +
# リスト 6.4.5 要素の値を色で表現した散布図
sns.relplot(data=tips, x="total_bill", y="tip", hue="size")
# +
# リスト 6.4.6 要素の値を大きさで表現した散布図
sns.relplot(data=tips, x="total_bill", y="tip", size="size")
# +
# リスト 6.4.7 fmri データセットの読み込み
fmri = sns.load_dataset("fmri")
fmri.head()
# +
# リスト 6.4.8 折れ線グラフの描画
import matplotlib.pyplot as plt
sorted_fmri = fmri.sort_values("timepoint")
fig, ax = plt.subplots()
ax.plot(sorted_fmri["timepoint"], sorted_fmri["signal"])
# +
# リスト 6.4.9 relplot 関数を用いて折れ線グラフを描画
sns.relplot(data=fmri, x="timepoint", y="signal", kind="line")
# +
# リスト 6.4.10 塗りつぶし色を標準偏差で描画
sns.relplot(data=fmri, x="timepoint", y="signal", kind="line", ci="sd")
# +
# リスト 6.4.11 データの集計をしない折れ線グラフ
sns.relplot(data=fmri, x="timepoint", y="signal", kind="line", estimator=None)
# +
# リスト 6.4.12 event 列のデータの種類
fmri["event"].unique()
# +
# リスト 6.4.13 region 列のデータの種類
fmri["region"].unique()
# +
# リスト 6.4.14 色と線種で分類した複数の折れ線グラフ
sns.relplot(
data=fmri, x="timepoint", y="signal", kind="line", hue="event", style="region"
)
# +
# リスト 6.4.15 ファセットを利用して散布図を描画
sns.relplot(data=tips, x="total_bill", y="tip", col="time", row="smoker")
# +
# リスト 6.4.16 ファセットを利用して折れ線グラフを描画
sns.relplot(
data=fmri, x="timepoint", y="signal", kind="line", row="event", col="region"
)
|
notebooks/6-04.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### get_all_tweets will get all the tweets related to the parameter you passed typed in there while calling the method.
#
# for example in our main we called method by passing "BLOOMBERG QUINT"
# +
import tweepy
import csv
consumer_key = "WG6WMvY5QnoKgb52MG9MZFAmt"
consumer_secret = "<KEY>"
access_key = "<KEY>"
access_secret = "<KEY>"
def get_all_tweets(screen_name):
auth = tweepy.OAuthHandler(consumer_key, consumer_secret)
auth.set_access_token(access_key, access_secret)
api = tweepy.API(auth)
alltweets = []
new_tweets = api.user_timeline(screen_name = screen_name,count=200)
alltweets.extend(new_tweets)
oldest = alltweets[-1].id - 1
while len(new_tweets) > 0:
print( "getting tweets before", oldest)
new_tweets = api.user_timeline(screen_name = screen_name,count=200,max_id=oldest)
alltweets.extend(new_tweets)
oldest = alltweets[-1].id - 1
print (len(alltweets)," downloaded")
outtweets = [[tweet.id_str, tweet.retweet_count, tweet.favorite_count, tweet.created_at, tweet.text.encode("utf-8")] for tweet in alltweets]
with open('%s_tweets.csv' % screen_name, 'w',newline='') as f:
writer = csv.writer(f)
writer.writerow(["id","number_of_retweets","number_of_favorites","created_at","text"])
writer.writerows(outtweets)
pass
if __name__ == '__main__':
get_all_tweets("BloombergQuint")
|
Scraping/TwitterScraping/AccessingBySearch/ScrapingTwitterBySearchName.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
sys.path.append('../scripts/')
from mcl import *
from kf import *
# %matplotlib widget
class EstimatedLandmark(Landmark):
def __init__(self):
super().__init__(0,0)
self.cov = None
def draw(self, ax, elems):
if self.cov is None:
return
##推定位置に青い星を描く##
c = ax.scatter(self.pos[0], self.pos[1], s=100, marker="*", label="landmarks", color="blue")
elems.append(c)
elems.append(ax.text(self.pos[0], self.pos[1], "id:" + str(self.id), fontsize=10))
##誤差楕円を描く##
e = sigma_ellipse(self.pos, self.cov, 3)
elems.append(ax.add_patch(e))
class MapParticle(Particle):
def __init__(self, init_pose, weight, landmark_num):
super().__init__(init_pose, weight)
self.map = Map()
for i in range(landmark_num):
self.map.append_landmark(EstimatedLandmark())
def init_landmark_estimation(self, landmark, z, distance_dev_rate, direction_dev):
landmark.pos = z[0]*np.array([np.cos(self.pose[2] + z[1]), np.sin(self.pose[2] + z[1])]).T + self.pose[0:2]
H = matH(self.pose, landmark.pos)[0:2,0:2] #カルマンフィルタのHの右上2x2を取り出す
Q = matQ(distance_dev_rate*z[0], direction_dev)
landmark.cov = np.linalg.inv(H.T.dot( np.linalg.inv(Q) ).dot(H))
def observation_update_landmark(self, landmark, z, distance_dev_rate, direction_dev): ###fastslam4landestm
estm_z = IdealCamera.observation_function(self.pose, landmark.pos) #ランドマークの推定位置から予想される計測値
if estm_z[0] < 0.01: # 推定位置が近すぎると計算がおかしくなるので回避
return
H = - matH(self.pose, landmark.pos)[0:2,0:2] #ここは符号の整合性が必要
Q = matQ(distance_dev_rate*estm_z[0], direction_dev)
K = landmark.cov.dot(H.T).dot( np.linalg.inv(Q + H.dot(landmark.cov).dot(H.T)) )
landmark.pos = K.dot(z - estm_z) + landmark.pos
landmark.cov = (np.eye(2) - K.dot(H)).dot(landmark.cov)
def observation_update(self, observation, distance_dev_rate, direction_dev): ###fastslam4obsupdate
for d in observation:
z = d[0]
landmark = self.map.landmarks[d[1]]
if landmark.cov is None:
self.init_landmark_estimation(landmark, z, distance_dev_rate, direction_dev)
else: #追加
self.observation_update_landmark(landmark, z, distance_dev_rate, direction_dev)
class FastSlam(Mcl):
def __init__(self, init_pose, particle_num, landmark_num, motion_noise_stds={"nn":0.19, "no":0.001, "on":0.13, "oo":0.2},\
distance_dev_rate=0.14, direction_dev=0.05):
super().__init__(None, init_pose, particle_num, motion_noise_stds, distance_dev_rate, direction_dev)
self.particles = [MapParticle(init_pose, 1.0/particle_num, landmark_num) for i in range(particle_num)]
self.ml = self.particles[0]
def observation_update(self, observation):
for p in self.particles:
p.observation_update(observation, self.distance_dev_rate, self.direction_dev) #self.mapを削除
self.set_ml()
self.resampling()
def draw(self, ax, elems):
super().draw(ax, elems)
self.ml.map.draw(ax, elems)
# +
def trial():
time_interval = 0.1
world = World(30, time_interval, debug=False)
###真の地図を作成###
m = Map()
for ln in [(-4,2), (2,-3), (3,3)]: m.append_landmark(Landmark(*ln))
world.append(m)
### ロボットを作る ###
init_pose = np.array([0,0,0]).T
pf = FastSlam(init_pose,100, len(m.landmarks))
a = EstimationAgent(time_interval, 0.2, 10.0/180*math.pi, pf)
r = Robot(init_pose, sensor=Camera(m), agent=a, color="red")
world.append(r)
world.draw()
trial()
# +
#a.estimator.particles[10].map.landmarks[2].cov
# +
#math.sqrt(0.0025)
# -
|
section_fastslam/fastslam4.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Taller evaluable sobre la extracción, transformación y visualización de datos usando IPython
# **<NAME>**
# <EMAIL>
# Universidad Nacional de Colombia, Sede Medellín
# Facultad de Minas
# Medellín, Colombia
# # Instrucciones
# En la carpeta 'Taller' del repositorio 'ETVL-IPython' se encuentran los archivos 'Precio_Bolsa_Nacional_($kwh)_'*'.xls' en formato de Microsoft Excel, los cuales contienen los precios históricos horarios de la electricidad para el mercado eléctrico Colombiano entre los años 1995 y 2017 en COL-PESOS/kWh. A partir de la información suministrada resuelva los siguientes puntos usando el lenguaje de programación Python.
# # Preguntas
#
# **1.--** Lea los archivos y cree una tabla única concatenando la información para cada uno de los años. Imprima el encabezamiento de la tabla usando `head()`.
import os
import pandas as pd
# +
x=[]
for n in range(1995,2018):
if n<2000:
skip=3
else:
skip=2
filename='Precio_Bolsa_Nacional_($kwh)_'+str(n)
if n>= 2016:
filename+='.xls'
else:
filename+='.xlsx'
y=pd.read_excel(filename,skiprows=skip,parse_cols=24)
x.append(y)
z=pd.concat(x)
len(z)
index=list(range(0,len(z)))
z.index = index
# -
# **2.--** Compute e imprima el número de registros con datos faltantes.
r=len(z)-len(z.dropna())
r
# **3.--** Compute e imprima el número de registros duplicados.
m=z[z.duplicated()]
len(m)
# **4.--** Elimine los registros con datos duplicados o datos faltantes, e imprima la cantidad de registros que quedan (registros completos).
#print(len(z))
m2=z.dropna()
#print(len(m2))
m3=m2.drop_duplicates()
print(len(m3))
# **5.--** Compute y grafique el precio primedio diario.
# +
#Precio promedio diario
import matplotlib.pyplot as plt
import matplotlib
# %matplotlib inline
m4=m3
m4['mean']=m3.mean(axis=1)
st=pd.to_datetime(m3['Fecha'],infer_datetime_format=True)
m4['Fecha']=st
m4['dia']=st.dt.dayofweek
m4['mes']=st.dt.month
m4['año']=st.dt.year
plt.plot(m4['Fecha'],m4['mean'],)
plt.ylabel('$')
plt.xlabel('año')
plt.title('Precio Promedio Diario del kWh 1995-2017')
plt.show()
# -
# **6.--** Compute y grafique el precio máximo por mes.
# +
w = []
m5=m4
for n in range(len(m3['Fecha'])):
w.append(str(m3.iloc[n,0])[0:7])
m5['key']=w
# +
#Precio Máximo del mes
# %matplotlib inline
y=list(m4.axes[1])
m5['max']=m4[y[1:25]].apply(max,axis=1)
b=m5.groupby('key').max()['max']
b.plot()
plt.ylabel('$')
plt.xlabel('Mes')
plt.title('Precio Máximo Mes de kWh')
# -
# **7.--** Compute y grafique el precio mínimo mensual.
# +
#Precio minimo mensual
# %matplotlib inline
m6=m5
y=list(m4.axes[1])
m6['min']=m4[y[1:25]].apply(min,axis=1)
b3=m6.groupby('key').min()['min']
b3.plot()
plt.ylabel('$')
plt.xlabel('Mes')
plt.title('Precio Mínimo Mes de kWh')
# -
# **8.--** Haga un gráfico para comparar el precio máximo del mes (para cada mes) y el precio promedio mensual.
# +
#Precio máximo del mes y precio promedio mensual
b4=m6.groupby('key').mean()['mean']
plt.figure()
b4.plot(legend='mean')
b.plot(legend='max')
plt.ylabel('$')
plt.xlabel('Mes')
plt.title('Comparativo Precio Máximo y promedio Mes de kWh')
# -
# **9.--** Haga un histograma que muestre a que horas se produce el máximo precio diario para los días laborales.
# +
#Histograma de horas con precio máximo días laborales
# %matplotlib inline
import numpy as np
from datetime import datetime, date, time, timedelta
import calendar
fecha=[]
fecha=m6['Fecha']
m6['Fecha']=pd.to_datetime(m6['Fecha'], format="%Y-%m-%d")
m6['Dia']=m6['Fecha'].dt.weekday_name
Lab = m6['Dia'].isin(['Monday','Tuesday','Wednesday','Thursday','Friday'])
Lab = m6[Lab]
indicador = ['{}'.format(n) for n in range(len(Lab))]
Lab.index = indicador
t=[]
for n in range(len(Lab)):
x = pd.Series(Lab.loc[str(n)]).values[1:25]
t.append ([i for i, e in enumerate(x) if e == max(x)])
a=[]
for n in range(len(t)):
for i in range (len(t[n])):
a.append(t[n][i])
rep=[]
for n in range (24):
rep.append(a.count(n))
plt.xlabel("Horas")
plt.ylabel("$/kWh")
plt.bar(range(24),rep,color='r',width = 0.9)
plt.show()
# -
# **10.--** Haga un histograma que muestre a que horas se produce el máximo precio diario para los días sabado.
# +
#Histograma de horas con precio máximo día Sabado
Sab = m6['Dia'].isin(['Saturday'])
Sab = m6[Sab]
indicador = ['{}'.format(n) for n in range(len(Sab))]
Sab.index = indicador
s=[]
for n in range(len(Sab)):
x = pd.Series(Sab.loc[str(n)]).values[1:25]
s.append ([i for i, e in enumerate(x) if e == max(x)])
a=[]
for n in range(len(s)):
for i in range (len(s[n])):
a.append(s[n][i])
rep=[]
for n in range (24):
rep.append(a.count(n))
plt.xlabel("Hora")
plt.ylabel("Frecuencia")
plt.title('Sabado')
plt.bar(range(24),rep,color='blue',width = 0.9)
plt.show()
# -
# **11.--** Haga un histograma que muestre a que horas se produce el máximo precio diario para los días domingo.
# +
#Histograma de horas con precio máximo día Domingo
Sun = m6['Dia'].isin(['Sunday'])
Sun = m6[Sun]
indicador = ['{}'.format(n) for n in range(len(Sun))]
Sun.index = indicador
s=[]
for n in range(len(Sun)):
x = pd.Series(Sun.loc[str(n)]).values[1:25]
s.append ([i for i, e in enumerate(x) if e == max(x)])
a=[] # Este Fragmento hace una matriz de la matriz u que contenia horas en que el maximo se repetia.
for n in range(len(s)):
for i in range (len(s[n])):
a.append(s[n][i])
rep=[]
for n in range (24):
rep.append(a.count(n))
plt.bar(range(24),rep,color='g',width = 0.9)
plt.show()
# -
# **12.--** Imprima una tabla con la fecha y el valor más bajo por año del precio de bolsa.
#Matriz con valores mínimos anuales - Se eliminaron los datos de Cero ya que este precio no es lógico
matrizSinCero = m6[m6>0].dropna()
Agrupac=matrizSinCero.groupby('año')['min'].idxmin()
ValorMinAnio=matrizSinCero.loc[Agrupac]
ValorMinAnio.filter(['año','Fecha','min'], axis=1)
# **13.--** Haga una gráfica en que se muestre el precio promedio diario y el precio promedio mensual.
# +
# Precio promedio mensual
b4=m6.groupby('key').mean()['mean']
plt.figure()
b4.plot(legend='promMes')
plt.show
plt.ylabel('$')
plt.xlabel('Mes')
plt.title('Promedio Mes')
plt.show()
# +
# precio promedio diario
plt.plot(m4['Fecha'],m4['mean'],)
plt.ylabel('$')
plt.xlabel('año')
plt.title('Precio Promedio Diario del kWh 1995-2017')
plt.show()
# -
# ---
|
Taller ETVL CarlosCeballos.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # PyDMD
# ## Tutorial 7: Dynamic mode decomposition with control
# In this tutorial we will show how to extend the dynamic mode decomposition to incorporate the effect of control (this technique has been introduced in the paper [Dynamic mode decomposition with control](https://arxiv.org/abs/1409.6358)).
# First of all we import the `DMDc` class from the pydmd package, we set matplotlib for the notebook and we import numpy and scipy.
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import scipy
from pydmd import DMDc
# -
# Now, we create our dataset: since we want to add the control, the evolution of the complex system can be formally summarized as:
# $$
# \mathbf{x}_{k+1} = \mathbf{A}\mathbf{x}_k + \mathbf{B}\mathbf{u}_k,
# $$where the operators $\mathbf{A}$ and $\mathbf{B}$ are the ones we will approximate using DMD. So, for a demostrative purpose, we create the original snapshots by using two random operators.
def create_system(n, m):
A = scipy.linalg.helmert(n, True)
B = np.random.rand(n, n)-.5
x0 = np.array([0.25]*n)
u = np.random.rand(n, m-1)-.5
snapshots = [x0]
for i in range(m-1):
snapshots.append(A.dot(snapshots[i])+B.dot(u[:, i]))
snapshots = np.array(snapshots).T
return {'snapshots': snapshots, 'u': u, 'B': B, 'A': A}
# We got 25 snapshots of the evolving system.
s = create_system(25, 10)
print(s['snapshots'].shape)
# Now, we can compute as usually the DMD algorithm on the data: the `fit` method in this version take as arguments the snapshots and the control input (the $\mathbf{B}$ operator can be also passed). In this case, we do not perform any truncation.
dmdc = DMDc(svd_rank=-1)
dmdc.fit(s['snapshots'], s['u'])
# Let us visualize the original system and the reconstructed one: also because without truncation, the plots are the same!
# +
plt.figure(figsize=(16,6))
plt.subplot(121)
plt.title('Original system')
plt.pcolor(s['snapshots'].real)
plt.colorbar()
plt.subplot(122)
plt.title('Reconstructed system')
plt.pcolor(dmdc.reconstructed_data().real)
plt.colorbar()
plt.show()
# -
# Well, we built the approximation for $\mathbf{A}$ and for $\mathbf{B}$; we can now test the system with a different control input: differently by the other versions, we can pass as argument of the `reconstructed_data` method the control input we want to test.
# +
new_u = np.exp(s['u'])
plt.figure(figsize=(8,6))
plt.pcolor(dmdc.reconstructed_data(new_u).real)
plt.colorbar()
plt.show()
# -
# Finally, we can also use a different timestep for the reconstruction, obviously passing the proper control input (the number of inputs we pass has to be the number of reconstructed snapshots we try to compute, except for the first snapshots). We continue halving the timestep and create a new random input.
# +
dmdc.dmd_time['dt'] = .5
new_u = np.random.rand(s['u'].shape[0], dmdc.dynamics.shape[1]-1)
plt.figure(figsize=(8,6))
plt.pcolor(dmdc.reconstructed_data(new_u).real)
plt.colorbar()
plt.show()
|
tutorials/tutorial-7-dmdc.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Ref :
# https://beckernick.github.io/oversampling-modeling/
# https://machinelearningmastery.com/standard-machine-learning-datasets-for-imbalanced-classification/
# https://towardsdatascience.com/methods-for-dealing-with-imbalanced-data-5b761be45a18
# +
import numpy as np
import pandas as pd
from sklearn.ensemble import RandomForestClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.model_selection import train_test_split
from sklearn.metrics import recall_score
from sklearn.metrics import f1_score
from imblearn.over_sampling import SMOTE
random_seed = 100
# -
# ### Read Data
df = pd.read_csv('creditcard.csv', header=None)
df.columns = ['col_' + str(col) for col in df.columns]
df.sample(10)
# ## check imblanace of target class
df_features = df.drop(['col_30'], axis=1)
df_target = df['col_30']
df_target.value_counts()
# ## Keep 10% of data aside to validate which technique would really work ?
x, x_real_world, y, y_real_world = train_test_split(df_features, df_target, test_size = .1, random_state=random_seed)
# # SMOTE Before Split
# +
sm = SMOTE(random_state=random_seed)
x_res, y_res = sm.fit_resample(x, y)
print("\nCounts before oversampling : \n", y.value_counts())
print("\nCounts after oversampling : \n", y_res.value_counts())
df_oversampled = pd.concat([x_res,y_res], axis=1)
# -
# ### Note : Not all rows are duplicated
print(df_oversampled.duplicated().sum())
df_oversampled.to_csv('resample_before_split.csv', index=False)
# ### Best Model ?
# +
x_train_res, x_test_res, y_train_res, y_test_res = train_test_split(x_res, y_res, test_size = .2, random_state=random_seed)
clf_dt = DecisionTreeClassifier(random_state=random_seed)
clf_dt.fit(x_train_res, y_train_res)
print("Train set f1_score : ", f1_score(y_train_res, clf_dt.predict(x_train_res)))
print("Test set f1_score : ", f1_score(y_test_res, clf_dt.predict(x_test_res)))
# -
# # What happend in Production : resampling before split ?
print("Real world data Accuracy : ", clf_dt.score(x_real_world, y_real_world)) # Big red, because of this you never identify your mistakes
print("Real world data Recall : ", recall_score(y_real_world, clf_dt.predict(x_real_world)))
print("Real World f1_score : ", f1_score(y_real_world, clf_dt.predict(x_real_world)))
# # SMOTE, Oversampling after split
# +
# Train, test split the original data
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size = .2, random_state=random_seed)
# Oversample using SMOTE
sm = SMOTE(random_state=random_seed)
x_train_res, y_train_res = sm.fit_resample(x_train, y_train)
print("\nCounts before oversampling : \n", y_train.value_counts())
print("\nCounts after oversampling : \n", y_train_res.value_counts())
# -
clf_dt = DecisionTreeClassifier(random_state=random_seed)
clf_dt.fit(x_train_res, y_train_res)
print("Train set f1_score : ", f1_score(y_train_res, clf_dt.predict(x_train_res)))
print("Test set f1_score : ", f1_score(y_test, clf_dt.predict(x_test)))
print("Real world data Accuracy : ", clf_dt.score(x_real_world, y_real_world)) # Big red, because of this you never identify your mistakes
print("Real world data Recall : ", recall_score(y_real_world, clf_dt.predict(x_real_world)))
print("Real World f1_score : ", f1_score(y_real_world, clf_dt.predict(x_real_world)))
|
Correct approach to oversampling.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Click "Edit App" to see the code
# # Improve figures layout
#
# In this tutorial we'll learn how to make good figures, which would be suitable for your reports or for a publication or for an Honours thesis.
# # The Jupyter Notebook
#
# Let's start by loading the Python packages, note that this time there is one extra package, which will be used to modify the ticks on the figure's axes.
# +
# python packages
import pandas as pd # Dataframes and reading CSV files
import numpy as np # Numerical libraries
import matplotlib.pyplot as plt # Plotting library
from lmfit import Model # Least squares fitting library
from matplotlib.ticker import (MultipleLocator,
FormatStrFormatter,
AutoMinorLocator)
# -
# First of all we want to define the figure's size and the font size that we want to use in the various parts of the figure. We use a variable _fontsize_ for the largest font in the image and then define the font size on the axes and the legend as a proportion of its value. This is useful because if we change the figure's size, we'll have to change the font size too.
# Define the figure's parameters
fontsize=28
figureParameters = {'figure.figsize' : (12,8),
'legend.fontsize': fontsize*0.7,
'axes.labelsize' : fontsize,
'axes.titlesize' : fontsize,
'xtick.labelsize': fontsize*0.8,
'ytick.labelsize': fontsize*0.8,
'xtick.direction': "in", # tick marks inside the frame
'ytick.direction': "out", # tick marks outside the frame
'axes.linewidth' : 3,
'axes.titlepad' : 25}
# We then define a function that we can call to customise the axes of a figure.
def prettyTicks(ax):
# Add tick marks on all sides of the figure
ax.xaxis.set_ticks_position('both')
ax.yaxis.set_ticks_position('both')
# Define the spacing of the major tick marks
# It is useful to comment this lines the first time
# we run the code to avoid errors because of an excessive
# number of tick marks
ax.xaxis.set_major_locator(MultipleLocator(2))
ax.yaxis.set_major_locator(MultipleLocator(0.01))
ax.yaxis.set_minor_locator(AutoMinorLocator())
ax.xaxis.set_minor_locator(AutoMinorLocator())
ax.tick_params(which='minor', length=6, width=3, color='black')
ax.tick_params(which='major', length=12, width=3, color='black')
# As a working example we can use a dataset for the Lambert-Beer law.
# We load the data in a DataFrame and fit them with a line, similarly to what we did in the previous tutorial.
# +
# Import the data
data = pd.read_csv("../miscData/LB.csv")
data.columns = ("X","Y")
nval = len(data.index)
# fit the data with a line
def lline(x,m,q):
return m*x + q
fitModel = Model(lline)
initialParameters = fitModel.make_params(m=1,q=1)
result = fitModel.fit(data["Y"], initialParameters, x=data["X"])
# Extract the best fitting parameters
mValue = result.best_values["m"]
qValue = result.best_values["q"]
# Create an array with the best fittig line
xl = np.arange(0,11)
plotLine = lline(xl,mValue,qValue)
# -
# For illustrative purposes, let's just make a graph without any customisations/
# +
# Create the figure with the data points and their fit
fig = plt.figure()
ax = fig.gca()
ax.scatter(data["X"], data["Y"])
ax.plot(xl,plotLine)
# Make the figure
plt.show()
# -
# Let's now make a better figure. These are the steps that we are going to follow:
# 1. Set figure parameters that we have defined earlier
# 2. Create new figure and axes objects
# 3. Add data and the fit line to the axes
# 4. Set the axes' labels
# 5. Set the limits of the axes
# 6. Make pretty tick marks by calling the function we defined before
# 7. Add legend to the plot in the upper left corner
# 8. Display the figure
#
# These commands have to be put in one box otherwise there can be problems with the figure representation.
# +
plt.rcParams.update(figureParameters)
fig2 = plt.figure()
ax = fig2.gca()
ax.scatter(data["X"], data["Y"], label="Data", s=75)
ax.plot(xl,plotLine, color='red', lw=3, label="Average")
ax.set(xlabel="Concentration (mmol/L)")
ax.set(ylabel="Absorbance")
plt.xlim([0,10])
plt.ylim([0,0.04])
prettyTicks(ax)
ax.legend(loc="upper left")
plt.show()
# -
# Although the plot is not perfect (tick marks should all be pointing inward or outward), it is a much better figure than the default one, and it can be used in a scientific document.
# Equally good figures can probably be done with Excel, but, in my opinion, this approach has the advantage that once a script is prepared all figures of a paper/thesis can be readily produced with a consistent style and little extra effort.
#
# The figure could be saved to a file by adding this command to a Python cell.
# ```python
# fig2.savefig("figure.png")
# ```
|
codeSnippets/8_prettyFigure.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext sppl.magics
import matplotlib.pyplot as plt
import numpy as np
# %%sppl model
from sppl.transforms import Sqrt
X ~= norm(loc=0, scale=2)
if (X < 1):
Z ~= -X**3+X**2+6*X
else:
Z~= -5*Sqrt(X)+11
# %sppl_to_graph model
# n = %sppl_get_namespace model
model = n.model
# Conditioning model on Event concerning the transformed variable Z
# changes the structure of the SPE.
modelc = model.condition((0 < n.Z) < 2)
# %sppl_to_graph modelc
|
examples/piecewise-transformation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + language="javascript"
# <!-- Ignore this block -->
# IPython.OutputArea.prototype._should_scroll = function(lines) {
# return false;
# }
# -
import warnings
warnings.filterwarnings("ignore", category= FutureWarning)
# +
import pandas as pd
import numpy as np
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.svm import SVC
from sklearn.pipeline import Pipeline
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import classification_report
from sklearn.ensemble import AdaBoostClassifier
import matplotlib
"""Set RC Params for matplotlib global"""
font = {'size' : 16}
matplotlib.rc('font', **font)
import matplotlib.pyplot as plt
# -
"""Splits the provided pandas dataframe into training and test dataset"""
def splitDataSet(inputDataframe, trainSetSize):
trainSet = inputDataframe.sample(frac = trainSetSize)
testSet = inputDataframe.drop(trainSet.index)
trainSet.reset_index(inplace = True)
testSet.reset_index(inplace = True)
return trainSet, testSet
# ## Load Data
inputData = pd.read_csv("data/SMSSpamCollection", sep="\t", header=None)
inputData.columns = ['target', 'text']
inputData.head()
# ## Encoding the target variables
# +
encodingDictionary = {"spam":1,
"ham":0}
inputData['target'] = inputData.target.apply(lambda x: encodingDictionary[x])
# -
# ## Split into train set and test set
trainSet, testSet = splitDataSet(inputData, 0.7)
# +
yTrain = trainSet.target.values
xTrain = trainSet.text.values
yTest = testSet.target.values
xTest = testSet.text.values
# -
# # Train SVM model
# <img src="https://upload.wikimedia.org/wikipedia/commons/thumb/2/2a/Svm_max_sep_hyperplane_with_margin.png/220px-Svm_max_sep_hyperplane_with_margin.png">
# +
"""Load Stop words"""
with open("data/stopwords") as file:
stopwordlist = file.readlines()
stopwordlist = [x.strip() for x in stopwordlist]
stopwords = set(stopwordlist)
# +
"""Define classification Pipeline"""
classificationPipeline = Pipeline([
("vectorizer", TfidfVectorizer(stop_words = stopwords, lowercase = True, ngram_range=(1, 3))),
("classifier", SVC())
])
# +
"""Define grid for performing grid search"""
parameterGrid = {
"classifier__C":np.arange(0.5, 10, 0.5),
"classifier__kernel":["rbf", "linear", "poly"],
"classifier__verbose":[True]
}
# +
"""Perform grid search"""
model = GridSearchCV(classificationPipeline, cv = 5, n_jobs = 3, param_grid = parameterGrid,
verbose = 1, refit = True)
model.fit(xTrain, yTrain)
# -
# ## Grid search results
scores = pd.DataFrame(model.cv_results_)
scores.head(57)
# +
plt.figure(figsize = (20, 15))
plt.title("Mean score for SVM")
plt.plot(model.cv_results_["mean_test_score"])
plt.plot(model.cv_results_["mean_test_score"], "bo", label = "model with different hyper parameters")
plt.xlabel('Models', fontsize = 15)
plt.ylabel('Mean test score', fontsize = 15)
plt.xticks(range(0, len(model.cv_results_['params'])), model.cv_results_['params'], rotation = 90)
plt.legend()
plt.show()
# -
# ## Classification Report
prediction = model.predict(xTest)
print(classification_report(yTest, prediction, target_names=["ham", "spam"]))
# # Train model for AdaBoost classifier
classificationPipeline = Pipeline([
("vectorizer", TfidfVectorizer(stop_words = stopwords, lowercase = True, ngram_range=(1, 3))),
("classifier", AdaBoostClassifier())
])
parameterGrid = {
"classifier__n_estimators":range(50,100),
"classifier__algorithm":["SAMME", "SAMME.R"]
}
# +
modelBoosted = GridSearchCV(classificationPipeline, cv = 5, n_jobs = 3, param_grid = parameterGrid,
verbose = 1, refit = True)
modelBoosted.fit(xTrain, yTrain)
# +
plt.figure(figsize = (30, 25))
plt.title("Mean score for AdaBoost")
plt.plot(modelBoosted.cv_results_["mean_test_score"])
plt.plot(modelBoosted.cv_results_["mean_test_score"], "bo", label = "model with different hyper parameters")
plt.xticks(range(0, len(modelBoosted.cv_results_['params'])), modelBoosted.cv_results_['params'], rotation = 90)
plt.xlabel('Models', fontsize = 15)
plt.ylabel('Mean test score', fontsize = 15)
plt.legend()
plt.show()
# -
# ## Classification Report
prediction = modelBoosted.predict(xTest)
print(classification_report(yTest, prediction, target_names=["ham", "spam"]))
# Both SVM and AdaBoostClassifier seems to be performing good. For SVM C parameter value gives constant performanace after some value.<br>
# The F1 score for both SVM and Adaboostclassifier is same. There is better recall for SVM for spam class as compared to Adaboost classifier. However Adaboost classifier is providing better recall for spam.
|
sklearn/svm/spamFilter/SpamFilterSKLearn.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Random Forests
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.datasets import load_breast_cancer
cancer = load_breast_cancer()
from sklearn.ensemble import RandomForestClassifier
X_train, X_test, y_train, y_test = train_test_split(
cancer.data, cancer.target, stratify=cancer.target, random_state=1)
rf = RandomForestClassifier(n_estimators=100).fit(X_train, y_train)
rf.feature_importances_
pd.Series(rf.feature_importances_,
index=cancer.feature_names).plot(kind="barh")
# # Exercise
# Use a random forest classifier or random forest regressor on a dataset of your choice.
# Try different values of n_estimators and max_depth and see how they impact performance and runtime.
# Tune ``max_features`` with GridSearchCV.
|
Bootcamp-materials/Day1-Machine-Learning/notebooks/07.2 - Random Forests.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import csv
import json
import requests
OPENFOODFACTS_API = 'https://world.openfoodfacts.org/api/v0'
REDMART_SEARCH_API = 'https://api.redmart.com/v1.6.0/catalog/search'
# -
# ## Redmart: API
# #### Category: Redmart Label
params = {
'q': 'redmart',
'category': 'redmart-label',
'pageSize': 1,
'sort': 1024
}
response = requests.get(REDMART_SEARCH_API, params=params)
response_json = response.json()
print(response_json['products'])
|
redmart/Redmart API.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (herschelhelp_internal)
# language: python
# name: helpint
# ---
# This notebook prepare the catalogues that will be analysed by CIGALE for SED fitting and physical parameter estimation.
# +
import numpy as np
import os
os.environ['LOG_LEVEL'] = 'INFO'
from astropy.table import Table
from herschelhelp.filters import correct_galactic_extinction
from herschelhelp.external import convert_table_for_cigale
# -
SUFFIX = '20180219'
master_catalogue = Table.read("../../dmu32/dmu32_NGP/data/NGP_{}_cigale.fits".format(SUFFIX))
len(master_catalogue)
# # Best sources
#
# Define a good far-IR measurement as:
# - an existing flux in the band;
# - the flag from XID+ must not be set;
# - the signal to noise ratio must be over 2.
# +
# good = {}
# for band in ['pacs_green', 'pacs_red', 'spire_250', 'spire_350', 'spire_500']:
# good[band] = (~np.isnan(master_catalogue['f_{}'.format(band)]) &
# ~master_catalogue['flag_{}'.format(band)])
# good[band][good[band]] &= (master_catalogue[good[band]]['f_{}'.format(band)] /
# master_catalogue[good[band]]['ferr_{}'.format(band)] >= 2)
# -
# We will keep only sources with at leat 2 good far-IR measurements (we may actually use less sources are not all may have a redshift).
# +
# combined_good = np.sum(list(good.values()), axis=0) >= 2
# +
# print("Number of good sources: {}".format(np.sum(combined_good)))
# -
# Only sources with at least two optical and at least two near infrared detections
optnir = ((master_catalogue['flag_optnir_det'] == 3)
| (master_catalogue['flag_optnir_det'] == 7))
# # Main catalogue for CIGALE
#best_catalogue = master_catalogue[combined_good].copy()
best_catalogue = master_catalogue[optnir].copy()
# Correction for galactic extinction
best_catalogue = correct_galactic_extinction(best_catalogue, inplace=True)
# Convertion to CIGALE format
best_catalogue = convert_table_for_cigale(best_catalogue, inplace=True, remove_zerofluxes=True)
# ## Band selection
#
# We want to use only one filter for similar bands. We define an order of preference and set to NaN the flux in the lower prefered bands when a prefered band is available. Some band may have a 0 flux, we set there values to NaN.
# +
u_bands = [ ]
g_bands = ["90prime_g", "gpc1_g"]
r_bands = ["90prime_r", "gpc1_r"]
i_bands = [ "gpc1_i"]
z_bands = ["mosaic_z", "decam_z", "gpc1_z"]
y_bands = [ "gpc1_y"]
def remove_unneeded_fluxes(list_of_bands):
for band_idx, band in enumerate(list_of_bands[:-1]):
mask = ~np.isnan(best_catalogue[band])
for lower_band in list_of_bands[band_idx+1:]:
best_catalogue[lower_band][mask] = np.nan
best_catalogue["{}_err".format(lower_band)][mask] = np.nan
# -
remove_unneeded_fluxes(g_bands)
remove_unneeded_fluxes(u_bands)
remove_unneeded_fluxes(r_bands)
remove_unneeded_fluxes(i_bands)
remove_unneeded_fluxes(z_bands)
remove_unneeded_fluxes(y_bands)
best_catalogue.write("data_tmp/NGP_cigale_optnir_extcor_20180129.fits")
# # Catalogue using spectroscopic redshift
best_catalogue = master_catalogue[optnir].copy()
best_catalogue.remove_column("redshift")
best_catalogue["zspec"].name = "redshift"
best_catalogue = best_catalogue[~np.isnan(best_catalogue["redshift"])]
print("Number of sources with z-spec: {}".format(len(best_catalogue)))
# Correction for galactic extinction
best_catalogue = correct_galactic_extinction(best_catalogue, inplace=True)
# Convertion to CIGALE format
os.environ['LOG_LEVEL'] = 'INFO'
best_catalogue = convert_table_for_cigale(best_catalogue, inplace=True, remove_zerofluxes=True)
remove_unneeded_fluxes(g_bands)
remove_unneeded_fluxes(u_bands)
remove_unneeded_fluxes(r_bands)
remove_unneeded_fluxes(i_bands)
remove_unneeded_fluxes(z_bands)
remove_unneeded_fluxes(y_bands)
best_catalogue.write("data_tmp/NGP_cigale_optnir_extcor_zspec_{}.fits".format(SUFFIX), overwrite=True)
|
dmu28/dmu28_NGP/CIGALE_catalogue_preparation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import requests
from bs4 import BeautifulSoup
soup=BeautifulSoup(requests.get('https://www.exchangerates.org.uk/Dollars-to-Egyptian-Pounds-currency-conversion-page.html').text,'html.parser')
soup
data=soup.find("span",attrs={"id":"shd2b;"}).get_text()
data
print(int(input("Enter your USD Amount: "))*float(data),'EGP')
|
assignment2.2 Task2 (from USD To EGP only).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# cd C:\Users\wanzheng\Desktop\1.3---实训\《Python数据分析与应用》-配套资料\01-数据和代码\第3章\01-任务程序\data
import numpy as np
import matplotlib.pyplot as plt
plt.rcParams['font.sans-serif'] = 'SimHei' # 设置中文显示
plt.rcParams['axes.unicode_minus'] = False
data = np.load('国民经济核算季度数据.npz')
data
name = data['columns'] # 提取其中的columns数组,视为数据的标签
values = data['values'] # 提取其中的values数组,数据的存在位置
name
values
# ### 散点图
plt.figure(figsize=(8, 7)) # 设置画布
plt.scatter(vbalues[:, 0], values[:, 2], marker='o') # 绘制散点图
plt.xlabel('年份') # 添加横轴标签
plt.ylabel('生产总值(亿元)') # 添加y轴名称
plt.xticks(range(0, 70, 4), values[range(0, 70, 4), 1], rotation=45)
plt.title('2000-2017年季度生产总值散点图') # 添加图表标题
# plt.savefig('../tmp/2000-2017年季度生产总值散点图.png')
plt.show()
plt.figure(figsize=(8, 7)) ## 设置画布
## 绘制散点1
plt.scatter(values[:, 0], values[:, 3], marker='o', c='red')
## 绘制散点2
plt.scatter(values[:, 0], values[:, 4], marker='D', c='blue')
## 绘制散点3
plt.scatter(values[:, 0], values[:, 5], marker='v', c='yellow')
plt.xlabel('年份') ## 添加横轴标签
plt.ylabel('生产总值(亿元)') ## 添加纵轴标签
plt.xticks(range(0, 70, 4), values[range(0, 70, 4), 1], rotation=45)
plt.title('2000-2017年各产业季度生产总值散点图') ## 添加图表标题
plt.legend(['第一产业', '第二产业', '第三产业']) ## 添加图例
# plt.savefig('../tmp/2000-2017年各产业季度生产总值散点图.png')
plt.show()
# ### 折线图
plt.figure(figsize=(8, 7)) ## 设置画布
## 绘制折线图
plt.plot(values[:, 0], values[:, 2], color='m', linestyle='--')
plt.xlabel('年份') ## 添加横轴标签
plt.ylabel('生产总值(亿元)') ## 添加y轴名称
plt.xticks(range(0, 70, 4), values[range(0, 70, 4), 1], rotation=45)
plt.title('2000-2017年季度生产总值折线图') ## 添加图表标题
# plt.savefig('../tmp/2000-2017年季度生产总值折线图.png')
plt.show()
# 代码 3-8
plt.figure(figsize=(8, 7)) ## 设置画布
plt.plot(values[:, 0], values[:, 2], color='r', linestyle='--',
marker='o') ## 绘制折线图
plt.xlabel('年份') ## 添加横轴标签
plt.ylabel('生产总值(亿元)') ## 添加y轴名称
plt.xticks(range(0, 70, 4), values[range(0, 70, 4), 1], rotation=45)
plt.title('2000-2017年季度生产总值点线图', rotation=45) ## 添加图表标题
# plt.savefig('../tmp/2000-2017年季度生产总值点线图.png')
plt.show()
# 代码 3-9
plt.figure(figsize=(8, 7)) ## 设置画布
plt.plot(values[:, 0], values[:, 3], 'bs-',
values[:, 0], values[:, 4], 'ro-.',
values[:, 0], values[:, 5], 'gH--') ## 绘制折线图
plt.xlabel('年份') ## 添加横轴标签
plt.ylabel('生产总值(亿元)') ## 添加y轴名称
plt.xticks(range(0, 70, 4), values[range(0, 70, 4), 1], rotation=-45)
plt.title('2000-2017年各产业季度生产总值折线图') ## 添加图表标题
plt.legend(['第一产业', '第二产业', '第三产业'])
plt.savefig('../tmp/2000-2017年季度各产业生产总值折线图.png')
plt.show()
plt.rcParams['font.sans-serif'] = 'SimHei' ## 设置中文显示
plt.rcParams['axes.unicode_minus'] = False
data = np.load('国民经济核算季度数据.npz')
name = data['columns'] ## 提取其中的columns数组,视为数据的标签
values = data['values'] ## 提取其中的values数组,数据的存在位置
p = plt.figure(figsize=(12, 12)) ##设置画布
## 子图1
ax1 = p.add_subplot(2, 1, 1)
plt.scatter(values[:, 0], values[:, 3], marker='o', c='r') ## 绘制散点
plt.scatter(values[:, 0], values[:, 4], marker='D', c='b') ## 绘制散点
plt.scatter(values[:, 0], values[:, 5], marker='v', c='y') ## 绘制散点
plt.ylabel('生产总值(亿元)') ## 添加纵轴标签
plt.title('2000-2017年各产业季度生产总值散点图') ## 添加图表标题
plt.legend(['第一产业', '第二产业', '第三产业']) ## 添加图例
## 子图2
ax2 = p.add_subplot(2, 1, 2)
plt.scatter(values[:, 0], values[:, 6], marker='o', c='r') ## 绘制散点
plt.scatter(values[:, 0], values[:, 7], marker='D', c='b') ## 绘制散点
plt.scatter(values[:, 0], values[:, 8], marker='v', c='y') ## 绘制散点
plt.scatter(values[:, 0], values[:, 9], marker='8', c='g') ## 绘制散点
plt.scatter(values[:, 0], values[:, 10], marker='p', c='c') ## 绘制散点
plt.scatter(values[:, 0], values[:, 11], marker='+', c='m') ## 绘制散点
plt.scatter(values[:, 0], values[:, 12], marker='s', c='k') ## 绘制散点
## 绘制散点
plt.scatter(values[:, 0], values[:, 13], marker='*', c='purple')
## 绘制散点
plt.scatter(values[:, 0], values[:, 14], marker='d', c='brown')
plt.legend(['农业', '工业', '建筑', '批发', '交通',
'餐饮', '金融', '房地产', '其他'])
plt.xlabel('年份') ## 添加横轴标签
plt.ylabel('生产总值(亿元)') ## 添加纵轴标签
plt.xticks(range(0, 70, 4), values[range(0, 70, 4), 1], rotation=45)
# plt.savefig('../tmp/2000-2017年季度各行业生产总值散点子图.png')
plt.show()
# 代码 3-11
p1 = plt.figure(figsize=(8, 7)) ## 设置画布
## 子图1
ax3 = p1.add_subplot(2, 1, 1)
plt.plot(values[:, 0], values[:, 3], 'b-',
values[:, 0], values[:, 4], 'r-.',
values[:, 0], values[:, 5], 'g--') ## 绘制折线图
plt.ylabel('生产总值(亿元)') ## 添加纵轴标签
plt.title('2000-2017年各产业季度生产总值折线图') ## 添加图表标题
plt.legend(['第一产业', '第二产业', '第三产业']) ## 添加图例
## 子图2
ax4 = p1.add_subplot(2, 1, 2)
plt.plot(values[:, 0], values[:, 6], 'r-', ## 绘制折线图
values[:, 0], values[:, 7], 'b-.', ## 绘制折线图
values[:, 0], values[:, 8], 'y--', ## 绘制折线图
values[:, 0], values[:, 9], 'g:', ## 绘制折线图
values[:, 0], values[:, 10], 'c-', ## 绘制折线图
values[:, 0], values[:, 11], 'm-.', ## 绘制折线图
values[:, 0], values[:, 12], 'k--', ## 绘制折线图
values[:, 0], values[:, 13], 'r:', ## 绘制折线图
values[:, 0], values[:, 14], 'b-') ## 绘制折线图
plt.legend(['农业', '工业', '建筑', '批发', '交通',
'餐饮', '金融', '房地产', '其他'])
plt.xlabel('年份') ## 添加横轴标签
plt.ylabel('生产总值(亿元)') ## 添加纵轴标签
plt.xticks(range(0, 70, 4), values[range(0, 70, 4), 1], rotation=45)
# plt.savefig('../tmp/2000-2017年季度各行业生产总值折线子图.png')
plt.show()
# ### 直方图
values[-1, 3:6]
label = ['第一产业', '第二产业', '第三产业'] ## 刻度标签
plt.figure(figsize=(6, 5)) ## 设置画布
plt.bar(range(3), values[-1, 3:6], width=0.5) ## 绘制散点图
plt.xlabel('产业') ## 添加横轴标签
plt.ylabel('生产总值(亿元)') ## 添加y轴名称
plt.xticks(range(3), label)
plt.title('2017年第一季度各产业国民生产总值直方图') ## 添加图表标题
plt.show()
# ### 饼图
# 代码 3-13
plt.figure(figsize=(6, 6)) ## 将画布设定为正方形,则绘制的饼图是正圆
label = ['第一产业', '第二产业', '第三产业'] ## 定义饼状图的标签,标签是列表
explode = [0.2, 0.01, 0.01] ## 设定各项离心n个半径
plt.pie(
values[-1, 3:6], explode=explode, labels=label, autopct='%1.1f%%', radius=1, rotatelabels=False) ## 绘制饼图
plt.title('2017年第一季度各产业国民生产总值饼图')
plt.savefig('../tmp/2017年第一季度各产业生产总值占比饼图')
plt.show()
# ### 箱线图
# 代码 3-14
label = ['第一产业', '第二产业', '第三产业'] ## 定义标签
gdp = (list(values[:, 3]), list(values[:, 4]), list(values[:, 5]))
plt.figure(figsize=(6, 4))
plt.boxplot(gdp, notch=True, vert=True, labels=label, meanline=True)
plt.title('2000-2017各产业国民生产总值箱线图')
plt.show()
# ### 练习
# #### 1
data = np.load('国民经济核算季度数据.npz')
name = data['columns'] ## 提取其中的columns数组,视为数据的标签
values = data['values']## 提取其中的values数组,数据的存在位置
plt.rcParams['font.sans-serif'] = 'SimHei' ## 设置中文显示
plt.rcParams['axes.unicode_minus'] = False
label1 = ['第一产业','第二产业','第三产业']## 刻度标签1
label2 = ['农业','工业','建筑','批发','交通',
'餐饮','金融','房地产','其他']## 刻度标签2
# +
p = plt.figure(figsize=(12, 12))
## 子图1
ax1 = p.add_subplot(2, 2, 1)
plt.bar(range(3), values[0, 3:6], width=0.5)
plt.xlabel('产业') ## 添加横轴标签
plt.ylabel('生产总值(亿元)') ## 添加y轴名称
plt.xticks(range(3), label1)
plt.title('2000年第一季度国民生产总值产业构成分布直方图')
## 子图2
ax2 = p.add_subplot(2, 2, 2)
plt.bar(range(3), values[-1, 3:6], width=0.5)
plt.xlabel('产业') ## 添加横轴标签
plt.ylabel('生产总值(亿元)') ## 添加y轴名称
plt.xticks(range(3), label1)
plt.title('2017年第一季度国民生产总值产业构成分布直方图')
## 子图3
ax3 = p.add_subplot(2, 2, 3)
plt.bar(range(9), values[0, 6:], width=0.5)
plt.xlabel('行业') ## 添加横轴标签
plt.ylabel('生产总值(亿元)') ## 添加y轴名称
plt.xticks(range(9), label2)
plt.title('2000年第一季度国民生产总值行业构成分布直方图') ## 添加图表标题
## 子图4
ax4 = p.add_subplot(2, 2, 4)
plt.bar(range(9), values[-1, 6:], width=0.5) ## 绘制散点图
plt.xlabel('行业') ## 添加横轴标签
plt.ylabel('生产总值(亿元)') ## 添加y轴名称
plt.xticks(range(9), label2)
plt.title('2017年第一季度国民生产总值行业构成分布直方图') ## 添加图表标题
## 保存并显示图形
plt.show()
# -
# #### 2
# +
# 代码 3-16
label1 = ['第一产业', '第二产业', '第三产业'] ## 标签1
label2 = ['农业', '工业', '建筑', '批发', '交通', '餐饮', '金融', '房地产', '其他'] ## 标签2
explode1 = [0.01, 0.01, 0.01]
explode2 = [0.01, 0.01, 0.01, 0.01, 0.01, 0.01, 0.01, 0.01, 0.01]
p = plt.figure(figsize=(12, 12))
## 子图1
ax1 = p.add_subplot(2, 2, 1)
plt.pie(
values[0, 3:6], explode=explode1, labels=label1,
autopct='%1.1f%%') ## 绘制散点图
plt.title('2000年第一季度国民生产总值产业构成分布饼图')
## 子图2
ax2 = p.add_subplot(2, 2, 2)
plt.pie(
values[-1, 3:6], explode=explode1, labels=label1,
autopct='%1.1f%%') ## 绘制散点图
plt.title('2017年第一季度国民生产总值产业构成分布饼图')
## 子图3
ax3 = p.add_subplot(2, 2, 3)
plt.pie(
values[0, 6:], explode=explode2, labels=label2,
autopct='%1.1f%%') ## 绘制散点图
plt.title('2000年第一季度国民生产总值行业构成分布饼图') ## 添加图表标题
## 子图4
ax4 = p.add_subplot(2, 2, 4)
plt.pie(
values[-1, 6:], explode=explode2, labels=label2,
autopct='%1.1f%%') ## 绘制散点图
plt.title('2017年第一季度国民生产总值行业构成分布饼图') ## 添加图表标题
## 保存并显示图形
# plt.savefig('../tmp/国民生产总值构成分布饼图.png')
plt.show()
# -
# #### 3
# +
# 代码 3-17
label1 = ['第一产业', '第二产业', '第三产业'] ## 标签1
label2 = ['农业', '工业', '建筑', '批发', '交通', '餐饮', '金融', '房地产', '其他'] ## 标签2
gdp1 = (list(values[:, 3]), list(values[:, 4]), list(values[:, 5]))
gdp2 = ([list(values[:, i]) for i in range(6, 15)])
p = plt.figure(figsize=(8, 8))
## 子图1
ax1 = p.add_subplot(2, 1, 1)
## 绘制散点图
plt.boxplot(gdp, notch=True, labels=label1, meanline=True)
plt.title('2000-2017各产业国民生产总值箱线图')
plt.ylabel('生产总值(亿元)') ## 添加y轴名称
## 子图2
ax2 = p.add_subplot(2, 1, 2)
## 绘制散点图
plt.boxplot(gdp2, notch=True, labels=label2, meanline=True)
plt.title('2000-2017各行业国民生产总值箱线图')
plt.xlabel('行业') ## 添加横轴标签
plt.ylabel('生产总值(亿元)') ## 添加y轴名称
plt.show()
# -
import numpy as np
import matplotlib.pyplot as plt
data = np.random.normal(size =100 , loc = 0 , scale = 1)
plt.boxplot(data , sym='o' , whis=0.05)
# print(data)
plt.show()
# cd C:\Users\wanzheng\PycharmProjects\Days\01-数据和代码\第3章\02-习题程序\data
# +
iris = np.load('iris.npz')['data'][:,:-1]
name = np.load('iris.npz')['features_name']
plt.rcParams['font.sans-serif'] = 'SimHei'
p = plt.figure(figsize=(16,16)) ##设置画布
plt.title('iris散点图矩阵')
for i in range(4):
for j in range(4):
p.add_subplot(4,4,(i*4)+(j+1))
plt.scatter(iris[:,i],iris[:,j])## 绘制散点图
plt.xlabel(name[i])
plt.ylabel(name[j])
plt.show()
|
matplotlib2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Create a QComponent - Advanced
from qiskit_metal import draw, Dict
from qiskit_metal.toolbox_metal import math_and_overrides
from qiskit_metal.qlibrary.core import QComponent
import qiskit_metal as metal
design = metal.designs.DesignPlanar()
# ## Qubits and Junctions
#
# The vast majority of junction management is actually under the QRenderers. The only information that a component designer needs to provide, is a linestring and width which indicates the location and orientation of a given junction. We can see this from a couple extracted lines of code from `TransmonPocket`
#
# `...`
#
# `rect_jj = draw.LineString([(0, -pad_gap / 2), (0, +pad_gap / 2)])`
#
# `...`
#
# `self.add_qgeometry('junction', dict(rect_jj=rect_jj), width=p.inductor_width)`
#
#
# In this case, the linestring is drawn between the two charge islands of the `TransmonPocket`. Much more of the junctions options are from renderer options added when the QRenderers are initiated. These are covered more in the renderer tutorials and sessions.
#
# It should be noted, currently multiple junctions in a component will receive the same renderer options. This is fine if, say, making a symmetric SQUID, though if trying to have asymmetry, (or, say fluxonium), a manner to handled multiple junction renderer options in a component is required.
from qiskit_metal.qlibrary.qubits.transmon_pocket import TransmonPocket
# ?TransmonPocket
# ## Exteriors, Interiors, and MultiPolygons
#
# As was shown in 3.1, there is a great amount of flexibility already present in Metal for what a component can be, though as it is still in development, there are some limitations with respect to if renderers can accurately render a given shape, say, a multi-faceted polygon where some facets are composed of splines. What capabilities are currently missing and would be beneficial to be added are all part of the development process.
#
# Currently, a poly can be generated with interior cut outs, such as the smiley face previously,
# +
face = draw.shapely.geometry.Point(0, 0).buffer(1)
eye = draw.shapely.geometry.Point(0, 0).buffer(0.2)
eye_l = draw.translate(eye, -0.4, 0.4)
eye_r = draw.translate(eye, 0.4, 0.4)
smile = draw.shapely.geometry.Point(0, 0).buffer(0.8)
cut_sq = draw.shapely.geometry.box(-1, -0.3, 1, 1)
smile = draw.subtract(smile, cut_sq)
face = draw.subtract(face, smile)
face = draw.subtract(face, eye_r)
face = draw.subtract(face, eye_l)
face
# -
# This differs from qgeometries which have `subtract=True`, as that specifically sets that geometry to be "etched" from the ground plane. The polygon face is composed of an exterior;
face.exterior
# and interiors, such as;
face.interiors[0]
# A renderer must recognize the difference between these shapes, as the current QRenderers do. This allows for the component designer to generate complex shapes, without having to worry about how to add the qgeometries in any particular manner. This is also true with MultiPolygons.
big_square = draw.rectangle(10,10,0,0)
cut_rectangle = draw.rectangle(12,1,0,0)
multi_poly = draw.subtract(big_square, cut_rectangle)
multi_poly
type(multi_poly)
# The MultiPolygon can still just be passed to add_qgeometry as one would with a regular polygon. It is broken up behind the scenes so two separate rectangles (with the appropriate coordinates) are added to the poly qgeometry table. This is handled by the add_qgeometry method of QGeometryTables.
# ?metal.qgeometries.QGeometryTables.add_qgeometry
# This method also handles rounding of coordinates to try and avoid any numerical errors. It is called by `metal.qlibrary.core.QComponent.add_qgeometry` and should not be called directly.
# ## QComponent Inheritance
#
# As is the case with python classes, one can extend a given component by creating a qcomponent which inherits said class, making it a parent/child relationship. While python does support multiple inheritances, Metal may run into some bugs, so it is best to keep inheritances as single paths of heritage.
#
# A good example is `TransmonPocketCL`, which adds a "charge line" the a "standard" `TransmonPocket`. As can be seen in the below code, none of the charge islands or other connection pads are present, but will still be generated via the `super().make()` line in the `make()` method.
# +
import numpy as np
from qiskit_metal import draw, Dict
from qiskit_metal.qlibrary.qubits.transmon_pocket import TransmonPocket
class TransmonPocketCL(TransmonPocket): # pylint: disable=invalid-name
"""
The base `TransmonPocketCL` class
Inherits `TransmonPocket` class
Description:
Create a standard pocket transmon qubit for a ground plane,
with two pads connected by a junction (see drawing below).
Connector lines can be added using the `connection_pads`
dictionary. Each connector line has a name and a list of default
properties.
This is a child of TransmonPocket, see TransmonPocket for the variables and
description of that class.
::
_________________
| |
|_______________| ^
________x________ | N
| | |
|_______________|
.. image::
Component_Qubit_Transmon_Pocket_CL.png
Charge Line:
* make_CL (bool): If a chargeline should be included.
* cl_gap (string): The cpw dielectric gap of the charge line.
* cl_width (string): The cpw width of the charge line.
* cl_length (string): The length of the charge line 'arm' coupling the the qubit pocket.
Measured from the base of the 90 degree bend.
* cl_ground_gap (string): How much ground is present between the charge line and the
qubit pocket.
* cl_pocket_edge (string): What side of the pocket the charge line is.
-180 to +180 from the 'west edge', will round to the nearest 90.
* cl_off_center (string): Distance from the center axis the qubit pocket is referenced to
"""
component_metadata = Dict(short_name='Q', _qgeometry_table_poly='True')
"""Component metadata"""
default_options = Dict(
make_CL=True,
cl_gap='6um', # the cpw dielectric gap of the charge line
cl_width='10um', # the cpw trace width of the charge line
# the length of the charge line 'arm' coupling the the qubit pocket.
cl_length='20um',
# Measured from the base of the 90 degree bend
cl_ground_gap=
'6um', # how much ground between the charge line and the qubit pocket
# -180 to +180 from the 'left edge', will round to the nearest 90.
cl_pocket_edge='0',
cl_off_center=
'100um', # distance from the center axis the qubit pocket is built on
)
"""Default drawing options"""
def make(self):
"""Define the way the options are turned into QGeometry."""
super().make()
if self.options.make_CL == True:
self.make_charge_line()
#####################################################################
def make_charge_line(self):
"""Creates the charge line if the user has charge line option to TRUE
"""
# Grab option values
name = 'Charge_Line'
p = self.p
cl_arm = draw.box(0, 0, -p.cl_width, p.cl_length)
cl_cpw = draw.box(0, 0, -8 * p.cl_width, p.cl_width)
cl_metal = draw.unary_union([cl_arm, cl_cpw])
cl_etcher = draw.buffer(cl_metal, p.cl_gap)
port_line = draw.LineString([(-8 * p.cl_width, 0),
(-8 * p.cl_width, p.cl_width)])
polys = [cl_metal, cl_etcher, port_line]
# Move the charge line to the side user requested
cl_rotate = 0
if (abs(p.cl_pocket_edge) > 135) or (abs(p.cl_pocket_edge) < 45):
polys = draw.translate(
polys, -(p.pocket_width / 2 + p.cl_ground_gap + p.cl_gap),
-(p.pad_gap + p.pad_height) / 2)
if (abs(p.cl_pocket_edge) > 135):
p.cl_rotate = 180
else:
polys = draw.translate(
polys, -(p.pocket_height / 2 + p.cl_groundGap + p.cl_gap),
-(p.pad_width) / 2)
cl_rotate = 90
if (p.cl_pocket_edge < 0):
cl_rotate = -90
# Rotate it to the pockets orientation
polys = draw.rotate(polys, p.orientation + cl_rotate, origin=(0, 0))
# Move to the final position
polys = draw.translate(polys, p.pos_x, p.pos_y)
[cl_metal, cl_etcher, port_line] = polys
# Generating pins
points = list(draw.shapely.geometry.shape(port_line).coords)
self.add_pin(name, points, p.cl_width) # TODO: chip
# Adding to element table
self.add_qgeometry('poly', dict(cl_metal=cl_metal))
self.add_qgeometry('poly', dict(cl_etcher=cl_etcher), subtract=True)
# -
# We can see this is the case by generating a TransmonPocketCL in the GUI.
gui = metal.MetalGUI(design)
# + tags=["nbsphinx-thumbnail"]
my_transmon_cl = TransmonPocketCL(design,'my_transmon_cl',options=dict(connection_pads=dict(a=dict(),b=dict(loc_W=-1))))
gui.rebuild()
gui.autoscale()
gui.screenshot()
# -
my_transmon_cl.options
# We can see that `my_transmon_cl` inherited the appropriate options from `TransmonPocket`, and even got the junction renderer options since its parent class does declare `_qgeometry_table_junction='True'`
gui.main_window.close()
|
docs/tut/2-From-components-to-chip/2.32-Create-a-QComponent-Advanced.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: lang101
# language: python
# name: lang101
# ---
# __Libraries__
# +
# system tools
import os
import sys
sys.path.append(os.path.join(".."))
# pandas, numpy, gensim
import pandas as pd
import numpy as np
import gensim.downloader
# import my classifier utility functions - see the Github repo!
import utils.classifier_utils as clf
# Import plotting function and embedding matrix function
#import ass6_utils as utils
# Machine learning stuff
from sklearn.model_selection import train_test_split
from sklearn.feature_extraction.text import CountVectorizer, TfidfVectorizer
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import ShuffleSplit # cross-validation
from sklearn import metrics
from sklearn.preprocessing import MinMaxScaler
from sklearn.preprocessing import LabelBinarizer
# tools from tensorflow
import tensorflow as tf
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import (Dense, Embedding,
Flatten, GlobalMaxPool1D, Conv1D)
from tensorflow.keras.optimizers import SGD, Adam # optimization algorithms
from tensorflow.keras.utils import plot_model
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
from tensorflow.keras.regularizers import L2 # regularization
# matplotlib
import matplotlib.pyplot as plt
# +
def plot_history(H, epochs):
"""
Utility function for plotting model history using matplotlib
H: model history
epochs: number of epochs for which the model was trained
"""
plt.style.use("fivethirtyeight")
plt.figure()
plt.plot(np.arange(0, epochs), H.history["loss"], label="train_loss")
plt.plot(np.arange(0, epochs), H.history["val_loss"], label="val_loss")
plt.plot(np.arange(0, epochs), H.history["accuracy"], label="train_acc")
plt.plot(np.arange(0, epochs), H.history["val_accuracy"], label="val_acc")
plt.title("Training Loss and Accuracy")
plt.xlabel("Epoch #")
plt.ylabel("Loss/Accuracy")
plt.legend()
plt.tight_layout()
plt.show()
def create_embedding_matrix(filepath, word_index, embedding_dim):
"""
A helper function to read in saved GloVe embeddings and create an embedding matrix
filepath: path to GloVe embedding
word_index: indices from keras Tokenizer
embedding_dim: dimensions of keras embedding layer
"""
vocab_size = len(word_index) + 1 # Adding again 1 because of reserved 0 index
embedding_matrix = np.zeros((vocab_size, embedding_dim))
with open(filepath) as f:
for line in f:
word, *vector = line.split()
if word in word_index:
idx = word_index[word]
embedding_matrix[idx] = np.array(
vector, dtype=np.float32)[:embedding_dim]
return embedding_matrix
# -
# ## LOGISTIC CLASSIFIER AS BENCHMARK
# __Load data__
# +
filename = os.path.join("..", "assignments", "data", "Game_of_Thrones_Script.csv")
data = pd.read_csv(filename, lineterminator = "\n")
data = data.loc[:, ("Season", "Sentence")]
# -
# Create training and test split using sklearn
X_train, X_test, y_train, y_test = train_test_split(chunks_sentences,
labels,
test_size=0.25,
random_state=42)
# __Create training data__
# +
sentences = data['Sentence'].values
labels = data['Season'].values
X_train, X_test, y_train, y_test = train_test_split(sentences,
labels,
test_size=0.25,
random_state=42)
# -
# __Vectorize__
# +
vectorizer = CountVectorizer()
X_train_feats = vectorizer.fit_transform(X_train)
X_test_feats = vectorizer.transform(X_test)
# -
# __Create logistic regression classifier__
classifier = LogisticRegression(random_state=42, max_iter = 1000).fit(X_train_feats, y_train)
# __Evaluate__
# +
y_pred = classifier.predict(X_test_feats)
classifier_metrics = metrics.classification_report(y_test, y_pred)
print(classifier_metrics)
# -
clf.plot_cm(y_test, y_pred, normalized=True)
# __Cross-validation___
# +
X_vect = vectorizer.fit_transform(sentences)
title = "Learning Curves (Logistic Regression)"
cv = ShuffleSplit(n_splits=100, test_size=0.2, random_state=0)
model = LogisticRegression(random_state=42)
clf.plot_learning_curve(model, title, X_vect, labels, cv=cv, n_jobs=4)
# -
# ## DEEP NEURAL NETWORK
# ### Model 1
# __Binarize labels__
lb = LabelBinarizer()
y_train = lb.fit_transform(y_train)
y_test = lb.fit_transform(y_test)
# __Create Word Embeddings__
# +
tokenizer = Tokenizer(num_words=5000)
tokenizer.fit_on_texts(X_train)
X_train_toks = tokenizer.texts_to_sequences(X_train)
X_test_toks = tokenizer.texts_to_sequences(X_test)
vocab_size = len(tokenizer.word_index) + 1
# -
# __Padding__
# +
maxlen = 100
X_train_pad = pad_sequences(X_train_toks,
padding='post',
maxlen=maxlen)
X_test_pad = pad_sequences(X_test_toks,
padding='post',
maxlen=maxlen)
# -
# __Define model architecture__
# +
embedding_dim = 100
l2 = L2(0.0001)
model = Sequential()
model.add(Embedding(input_dim=vocab_size,
output_dim=embedding_dim,
input_length=maxlen))
model.add(Conv1D(128, 5,
activation='relu',
kernel_regularizer=l2))
model.add(GlobalMaxPool1D())
model.add(Dense(10, activation='relu',
kernel_regularizer=l2))
model.add(Dense(8,
activation='softmax'))
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
model.summary()
# -
# __Train and evaluate__
history = model.fit(X_train_pad, y_train,
epochs=20,
verbose=False,
validation_data=(X_test_pad, y_test),
batch_size=10)
loss, accuracy = model.evaluate(X_train_pad, y_train, verbose=False)
print("Training Accuracy: {:.4f}".format(accuracy))
loss, accuracy = model.evaluate(X_test_pad, y_test, verbose=False)
print("Testing Accuracy: {:.4f}".format(accuracy))
plot_history(history, epochs=20)
# ### Model 4
# __Use Pretrained Word Embeddings__
tf.keras.backend.clear_session()
# +
# #!wget http://nlp.stanford.edu/data/glove.6B.zip
# #!unzip -q glove.6B.zip
# -
embedding_matrix = create_embedding_matrix('../glove/glove.6B.100d.txt',
tokenizer.word_index,
embedding_dim)
# +
embedding_dim = 100
l2 = L2(0.0001)
model = Sequential()
model.add(Embedding(vocab_size,
embedding_dim,
weights=[embedding_matrix],
input_length=maxlen,
trainable=False))
model.add(Conv1D(128, 5,
activation='relu',
kernel_regularizer=l2))
model.add(GlobalMaxPool1D())
model.add(Dense(10,
activation='relu',
kernel_regularizer=l2))
model.add(Dense(8,
activation='softmax'))
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
model.summary()
# +
history = model.fit(X_train_pad, y_train,
epochs=50,
verbose=False,
validation_data=(X_test_pad, y_test),
batch_size=10)
loss, accuracy = model.evaluate(X_train_pad, y_train, verbose=False)
print("Training Accuracy: {:.4f}".format(accuracy))
loss, accuracy = model.evaluate(X_test_pad, y_test, verbose=False)
print("Testing Accuracy: {:.4f}".format(accuracy))
plot_history(history, epochs = 50)
# -
# ### Model 5
# __Retraining the GloVe embeddings__
# +
embedding_dim = 100
l2 = L2(0.0001)
model = Sequential()
model.add(Embedding(vocab_size,
embedding_dim,
weights=[embedding_matrix],
input_length=maxlen,
trainable=True))
model.add(Conv1D(128, 5,
activation='relu',
kernel_regularizer=l2))
model.add(GlobalMaxPool1D())
model.add(Dense(10,
activation='relu',
kernel_regularizer=l2))
model.add(Dense(8, activation='softmax'))
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
# +
history = model.fit(X_train_pad, y_train,
epochs=50,
verbose=False,
validation_data=(X_test_pad, y_test),
batch_size=10)
# evaluate
loss, accuracy = model.evaluate(X_train_pad, y_train, verbose=False)
print("Training Accuracy: {:.4f}".format(accuracy))
loss, accuracy = model.evaluate(X_test_pad, y_test, verbose=False)
print("Testing Accuracy: {:.4f}".format(accuracy))
# plot
plot_history(history, epochs = 50)
|
notebooks/assignment6_notebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="./Img/Label_02.png">
# 5 февраля 2021 года <br>
# Семинар <br>
# ПИ19-3, ПИ19-4 - 3 подгруппа<br>
# ПИ19-4, ПИ19-5 - 4 подгруппа
#
# 6 февраля 2021 года <br>
# Семинар <br>
# ПИ19-2, ПИ19-3, ПИ19-4 - 2 подгруппа
# # Тема 1. SQLAlchemy и язык выражений SQL
# SQL Expression Language
# # Введение
# ## SQLAlchemy и язык выражений SQL
#
# SQLAlchemy - библиотека Пайтон, которая устраняет разрыв между реляционными базами данных и традиционным программированием. Хотя SQLAlchemy позволяет «опуститься» до необработанного SQL для выполнения запросов, она поощряет мышление более высокого уровня за счет более «питонического» и дружественного подхода к запросам и обновлению базы данных. SQLAlchemy используется для взаимодействия с широким спектром баз данных. Она позволяет создавать модели данных и запросы в манере, напоминающей обычные классы и операторы Python.
#
# Язык выражений SQL (SQL Expression Language), называемый также Кор (Core, ядро) - это инструмент SQLAlchemy для представления общих операторов и выражений SQL в стиде Пайтон. Он ориентирован на фактическую схему базы данных и стандартизирован таким образом, что обеспечивает единообразный язык для большого числа серверных баз данных.
#
# SQLAlchemy Core имеет представление, ориентированное на схему, таблицы, ключи и индексы, как и традиционный SQL. SQLAlchemy Core эффективен в отчетах, анализе и других применениях, где полезно иметь возможность жестко контролировать запрос или работать с немоделированными данными. Надежный пул соединений с базой данных и оптимизация набора результатов идеально подходят для работы с большими объемами данных.
# ## Кодирование
#
# Работать с SQLAlchemy удобно в интерактивной среде "чтение-оценка-вывод" (read-evaluate-print-loop REPL), в такой, как интерактивный ноутбук ipython http://ipython.org/
#
# Установка Юпитер Ноутбук
# https://jupyter.readthedocs.io/en/latest/install/notebook-classic.html
#
# Для освоения и работы с ноутбуком iPython, рекомендуется пакет программ Анаконда (Anaconda).
#
# https://www.anaconda.com/products/individual
#
# Анаконда содержит Пайтон, Юпитер ноутбук и другие часто используемые приложения для научных вычислений и обработки данных.
#
# Порядок установки и запуска интерктивной среды Юпитер ноутбук
#
# 1. Загрузить Анаконда
# 2. Установить Анаконда
# 3. Запустить Юпитер ноутбук. Для этого использовать команду меню, либо консольную команду
#
# => jupyter notebook
# ## Установка SQLAlchemy
# +
# pip3 install sqlalchemy
# -
# ## Установка драйверов баз данных
#
# По **_умолчанию_** SQLAlchemy поддерживает SQLite3 без дополнительных драйверов. Для подключения к другим базам данных необходимы дополнительные драйверы баз данных.
#
# - PostgreSQL. <br>Установка драйвера Psycopg2: http://initd.org/psycopg/
# - MySQL (требуется версия MySQL 4.1 и выше)
# - Другие
#
# SQLAlchemy также можно использовать вместе с Drizzle, Firebird, Oracle, Sybase и Microsoft SQL Server. Сообщество также предоставило внешние диалекты для многих других баз данных, таких как IBM DB2, Informix, Amazon Redshift, EXASolution, SAP SQL Anywhere, Monet и многих других. Создание диалектов поддерживается SQLAlchemy.
# +
# PostgreSQL
# # ! pip install psycopg2
# MySQL
# # ! pip install pymysql
# -
# ## Соединение с базой данных
#
# Чтобы подключиться к базе данных, нужно создать механизм (движок) SQLAlchemy. Механизм SQLAlchemy создает общий интерфейс с базой данных для выполнения операторов SQL.
#
# SQLAlchemy предоставляет функцию для создания механизма с учетом *строки подключения* и, возможно, некоторых дополнительных именованных (keywords) аргументов. Строка подключения может содержать:
#
# - Тип базы данных (Postgres, MySQL, etc.);
# - Диалект, если он отличается от установленного по умолчанию для конкретного типа базы данных (Psycopg2, PyMySQL и т. д.);
# - Дополнительные данные аутентификации (имя пользователя и пароль);
# - Расположение базы данных (файл или имя хоста сервера базы данных);
# - Дополнительный порт сервера базы данных;
# - Необязательное имя базы данных.
#
# Строка подключения позволяют нам использовать конкретный файл или место хранения. В примере 1 определяется файл базы данных SQLite с именем <TT>listings.db</tt>:
# - хранящийся в текущем каталоге;
# - в памяти;
# - с указанием полного пути к файлу (Unix и Windows).
#
# В Windows строка подключения будет иметь вид engine4; \\\ требуются для экранирования символа "слэш".
#
# Функция <TT>create_engine</tt> возвращает экземпляр механизма SQLAlchemy.
# +
# 0-1. Создание механизма SQLAlchemy со строкой подключения SQLite
import sqlalchemy
from sqlalchemy import create_engine
engine = create_engine('sqlite:///listings.db')
# engine2 = create_engine('sqlite:///:memory:')
# engine3 = create_engine('sqlite:////home/Airbnb/listings.db')
# engine4 = create_engine('sqlite:///c:\\Users\\Airbnb\\listings.db')
# -
# PostgreSQL<br>Пример 2. Cоздание механизма для локальной базы данных PostgreSQL с именем mydb
#
# `
# from sqlalchemy import create_engine
# engine=create_engine('postgresql+psycopg2://username:password@localhost:5432/mydb')
# `
# MySQL. <br> Пример 3. Создание механизма для удаленной БД MySQL
#
# `
# from sqlalchemy import create_engine
# engine = create_engine('mysql+pymysql://username:password''@mysql01.mikhail.internal/listings', pool_recycle=3600)
# `
# Теперь, когда создан экземпляр механизма соединения с базой данных, мы можем начать использовать SQLAlchemy Core чтобы связать наше приложение с сервисами базы данных.
# # 1.1. Схема и типы данных
# В процессе выполнения кода примеров этой темы нам понадобятся библиотеки
import pandas as pd
import numpy as np
from datetime import datetime
from sqlalchemy import (MetaData, Table, Column, Integer, Numeric, String, DateTime,
Boolean, ForeignKey, create_engine, PrimaryKeyConstraint,
UniqueConstraint, CheckConstraint, Index, insert, BigInteger)
# В SQLAlchemy имеется четыре категории типов данных:
# - Универсальный
# - Стандартный SQL
# - Зависящий от поставщика
# - Определяется пользователем
#
# Универсальная категория типов данных предназначена для сопоставления типов данных в Python и SQL.
pd.DataFrame(['BigInteger,int,BIGINT'.split(','),
'Boolean,bool,BOOLEAN or SMALLINT'.split(','),
'Date,datetime.date,DATE (SQLite: STRING)'.split(','),
'DateTime,datetime.datetime,DATETIME (SQLite: STRING)'.split(','),
'Enum,str,ENUM or VARCHAR'.split(','),
'Float,float or Decimal,FLOAT or REAL'.split(','),
'Integer,int,INTEGER'.split(','),
'Interval,datetime.timedelta,INTERVAL or DATE from epoch'.split(','),
'LargeBinary,byte,BLOB or BYTEA'.split(','),
'Numeric,decimal.Decimal,NUMERIC or DECIMAL'.split(','),
'Unicode,unicode,UNICODE or VARCHAR'.split(','),
'Text,str,CLOB or TEXT'.split(','),
'Time,datetime.time,DATETIME'.split(',')],
columns='SQLAlchemy,Python,SQL'.split(','))
# Стандартные типы (например CHAR и NVARCHAR) используются в случаях, когда универсальные типы не отвечают требованиям из-за конкретной структуры данных.
#
# Типы, зависящие от поставщика. Пример: поле JSON в PostgreSQL.
#
# <TT>fromsqlalchemy.dialects.postgresqlimport JSON</tt>
# ## Метаданные
#
# Метаданные используются для связывания структуры базы данных. Метаданные полезно рассматривать как каталог объектов таблиц с дополнительной информацией о механизме и соединении. Метаданные необходимо импортировать и инициализировать. Инициализируем экземпляр объектов MetaData:
# +
# 1-1
from sqlalchemy import MetaData
metadata = MetaData()
# -
# ## Таблицы
#
# Объекты таблиц инициализируются в SQLAlchemy Core путем вызова конструктора <TT>Table</tt> с именем таблицы и метаданными, аргументы считаются объектами столбцов. Столбцы создаются путем вызова Column с именем, типом и затем аргументами, которые представляют дополнительные конструкции и ограничения SQL. В примере 1-2 создадим таблицу, которая может использоваться для перечня объектов размещения гостиничного бизнеса airbnb: http://insideairbnb.com/get-the-data.html
#
# +
# 1-2
listing=Table('listing',metadata,
Column('listing_id',Integer(),primary_key=True),
Column('listing_name',String(50),index=True),
Column('listing_url',String(255)),
Column('host_id',Integer()),
Column('neighbourhood_id',Integer()),
Column('amenities',String(300)),
Column('property_type_id',Integer()),
Column('room_type_id',Integer()),
Column('bedrooms',Integer()),
Column('beds',Integer()),
Column('price',Numeric(7,2)),
CheckConstraint('price >= 0.00', name='listing_price_positive'),
extend_existing=True
)
# -
pd.DataFrame({'En':['listing_id','listing_name','listing_url','host_id','neighbourhood_id',
'amenities','property_type_id','room_type_id','bedrooms','beds','price'],
'Ru':['идентификатор объекта размещения','имя объекта размещения',
'адрес веб-страницы','идентификатор владельца',
'идентификатор местоположения','оборудование, удобства',
'тип собственности','тип помещения','число спален','число кроватей','цена']})
# ### Дополнительные аргументы
#
# Рассмотрим использование дополнительных аргументов <TT>nullable, unique, onupdate</tt>
# +
# 1-3
user=Table('user',metadata,
Column('user_id',Integer(),primary_key=True),
Column('user_name',String(15),nullable=False,unique=True),
Column('email_address',String(255),nullable=False),
Column('phone',String(20),nullable=False),
Column('password',String(25),nullable=False),
Column('created_on',DateTime(),default=datetime.now),
Column('updated_on',DateTime(),default=datetime.now,onupdate=datetime.now)
)
# -
# ### Ключи и ограничения
#
# Ключи и ограничения задают с помощью объектов <TT>PrimaryKeyConstraint, UniqueConstraint, CheckConstraint</tt>
# #### Первичный ключ
# В примерах 1-2 и 1-3 столбцы `listing_id` и `user_id` объявлялись первичными ключами с помощью ключевого слова <TT>primary_key</tt>. Также, можно определить составной первичный ключ, присвоив параметру <TT>primary_key</tt> значение <TT>True</tt> для нескольких столбцов. Таким образом, ключ рассматривается как кортеж, в котором столбцы, помеченные как ключ, присутствуют в порядке, в котором они были определены в таблице. Первичные ключи также могут быть определены после столбцов в конструкторе таблицы, как показано в следующем фрагменте.
#
# `user=Table('user',metadata,
# Column('user_name',String(15),nullable=False,unique=True),
# Column('email_address',String(255),nullable=False),
# Column('phone',String(20),nullable=False),
# Column('password',String(25),nullable=False),
# Column('created_on',DateTime(),default=datetime.now),
# Column('updated_on',DateTime(),default=datetime.now,onupdate=datetime.now),
# PrimaryKeyConstraint('user_id', name='user_pk'),
# extend_existing=True
# )`
# #### Уникальность
# Другое распространенное ограничение - ограничение уникальности, которое используется, чтобы гарантировать, что в данном поле значения не повторяются.
#
# <TT>UniqueConstraint('user_name', name='uix_username')</tt>
# #### Проверка значения
# Этот тип ограничения используется, чтобы гарантировать, что данные, предоставленные для столбца, соответствуют набору критериев, определенных пользователем. В следующем фрагменте кода мы гарантируем, что <TT>price</tt> не может быть меньше 0,00:
#
# <TT>CheckConstraint('price >= 0.00', name='listing_price_positive')</tt>
# ### Индексы
#
# В примере 1-2 создан индекс для столбца listing_name. Когда индексы создаются, как показано в этом примере, они получают имена <TT>ix_listings_listing_name</tt>. Мы также можем определить индекс, используя явный тип конструкции. Можно обозначить несколько столбцов, разделив их запятой. Можно добавить аргумент ключевого слова <TT>unique = True</tt>, чтобы индекс был уникальным. При явном создании индексов они передаются в конструктор таблиц после столбцов. Чтобы имитировать указанный индекс явным способом, в конструктор <TT>Table</tt> требуется добавить <TT>Index('ix_listings_listing_name', 'listing_name')</tt>
# Мы также можем создавать функциональные индексы для ситуаций, когда часто требуется запрос на основе нескольких полей БД. Например, если мы хотим искать по параметрам оборудования ("удобства") и цены в качестве объединенного элемента, можно определить функциональный индекс для оптимизации поиска:
Index('ix_am_price', listing.c.amenities, listing.c.price)
# ### Связи и внешние ключи
# Тепрь, когда имеются пользователи и объекты размещения, необходимо обеспечить связи, позволяющие пользователям бронировать те или иные объекты. Рассмотрим схему данных.
# <img src=./Img/listing_schema.png width="560">
# Создадим таблицы для заказов `order`, `line_item`, таблицу местоположений `neighbourhood`, таблицы типов собственности и комнат `property_type`, `room_type`.
# +
# 1-4
order = Table('order', metadata,
Column('order_id', Integer(),primary_key=True),
Column('user_id', ForeignKey('user.user_id')),
Column('confirmed', Boolean(),default=False),
Column('order_price', Integer()),
extend_existing=True
)
# +
# 1-5
line_item = Table('line_item', metadata,
Column('line_item_id', Integer(), primary_key=True),
Column('order_id', ForeignKey('order.order_id')),
Column('listing_id', ForeignKey('listing.listing_id')),
Column('item_start_date', DateTime()),
Column('item_end_date', DateTime()),
Column('item_price', Integer()),
extend_existing=True
)
# +
# 1-6
neighbourhood=Table('neighbourhood',metadata,
Column('neighbourhood_id',Integer(),primary_key=True),
Column('neighbourhood_name',String(30)),
)
# +
# 1-7
property_type=Table('property_type',metadata,
Column('property_type_id',Integer(),primary_key=True),
Column('property_type_name',String(30))
)
# +
# 1-8
room_type=Table('room_type',metadata,
Column('room_type_id',Integer(),primary_key=True),
Column('property_type_name',String(30))
)
# -
# В примерах 1-4 и 1-5 внешние ключи задаются с помощью строки: <TT>'order.order_id'</tt>. Также существует явный способ задания ограничений по внешнему ключу: <TT>ForeignKeyConstraint(['order_id'], ['order.order_id'])</tt>
# ## Сохранение таблиц
#
# Все таблицы и определения связаны с экземпляром метаданных. Сохранение схемы в базе данных осуществляется посредством вызова метода <TT>create_all()</tt> в экземпляре метаданных с движком, в котором он должен создавать эти таблицы. По умолчанию create_all не будет пытаться воссоздать таблицы, которые уже существуют в базе данных, и его можно запускать несколько раз. Движок (механизм) SQLAlchemy определен нами ранее в примере 0-1, экземпляр метаданных создан ранее в примере 1-1. Теперь осуществим вызов метода <TT>create_all()</tt>
metadata.create_all(engine)
# ## Дополнительно: DB Browser for SQLite
# https://sqlitebrowser.org/
# ### Контрольная работа
#
# 1. Используя библиотеку SQLAlchemy и Юпитер Ноутбук создать реляционную базу данных (до 5-6 таблиц), отражающих финансово-экономческую деятельность отдела предприятия или решающих какую-либо задачу в рамках работы отдела или предпрития.
# 2. Создать схему (нарисовать).
# 3. Написать аннтоцию, кратко о смысле деятельности, отраженной в структуре данных.
# 4. Схему в pdf, png, jpeg и т. д. приложить к письму с результатом.
#
# Результат в формате Юпит<NAME> прислать на почту <EMAIL>.
#
# В теме письма указать подгруппу, Фимилию, ИО. В отдельной ячейке ноутбука указать подгруппу, Фимилию, ИО.
# <img src="./Img/Label_02.png">
# 19 февраля 2021 года <br>
# Семинар <br>
# ПИ19-3, ПИ19-4 - 3 подгруппа<br>
# ПИ19-4, ПИ19-5 - 4 подгруппа
#
# 20 февраля 2021 года <br>
# Семинар <br>
# ПИ19-2, ПИ19-3, ПИ19-4 - 2 подгруппа
# # 1.2. Работа с данными
# Теперь, когда в нашей базе данных есть таблицы, приступим к работе с данными внутри этих таблиц. Мы рассмотрим, как вставлять, извлекать и удалять данные, а затем научимся сортировать, группировать и использовать связи в наших данных. Мы будем использовать язык SQLExpression Language (SEL), предоставляемый SQLAlchemy Core. Начнем с изучения того, как вставлять данные.
# ## Вставка данных
#
# Вставим первую строку `ListingsAmsterdam.csv`.
am=pd.read_csv('ListingsAmsterdam.csv',sep=';')
am.head(5)
am['neighbourhood_cleansed'].unique() #Уникальные значения
values_count = am['neighbourhood_cleansed'].value_counts().sort_index() #Сколько раз встречается
print(values_count)
r = pd.DataFrame(values_count.reset_index())
print(r)
print(type(r))
am_neigh.index=range(1,len(am_neigh)+1)
am_neigh
# +
print("PREVIOUS")
print(am)
result = pd.DataFrame(am['neighbourhood_cleansed'].value_counts())
print("RESULT")
print(result)
am_neigh=pd.DataFrame(am['neighbourhood_cleansed'].value_counts()).sort_index().reset_index()
am_neigh.index=range(1,len(am_neigh)+1)
am_neigh
# -
am_prop=pd.DataFrame(am['property_type'].value_counts()).sort_index().reset_index()
am_prop.index=range(1,len(am_prop)+1)
am_prop
am_room=pd.DataFrame(am['room_type'].value_counts()).sort_index().reset_index()
am_room.index=range(1,len(am_room)+1)
am_room
am.loc[0]["host_id"]
check = am.loc[0,'amenities'][:300]
print(check)
# Создадим оператор вставки, чтобы поместить объект размещения в таблицу. Для этого мы можем вызвать метод `insert()` для таблицы `listing`, а затем использовать оператор `values()` с аргументами для каждого столбца.
# +
# 2-1 Одиночная вставка как метод
ins=listing.insert().values(
listing_id=20168,
listing_name='Studio with private bathroom in the centre 1',
listing_url='https://www.airbnb.com/rooms/20168',
host_id=59484,
neighbourhood_id=4,
property_type_id=35,
room_type_id=3,
amenities=am.loc[0,'amenities'][:300],
bedrooms=1,
beds=1,
price=236
)
print(str(ins))
# -
# `print(str(ins))` показывает нам фактический оператор SQL, который будет выполнен. Наши значения были заменены на: `column_name` в этом операторе SQL, именно так SQLAlchemy представляет параметры, отображаемые с помощью функции `str()`. Параметры используются, чтобы гарантировать, что наши данные были правильно экранированы, что снижает проблемы безопасности, такие как атаки с использованием SQL-инъекций. По-прежнему можно получить параметры, посмотрев на скомпилированную версию оператора вставки, потому что каждая внутренняя часть базы данных может обрабатывать параметры по-разному (это контролируется диалектом).
#
# Метод `compile()` для объекта `ins` возвращает объект `SQLCompiler`, который дает нам доступ к фактическим параметрам, которые будут отправлены с запросом через атрибут params:
ins.compile().params
# Теперь, когда у нас есть полное представление об операторе вставки и мы понимаем, что будет вставлено в таблицу, мы можем использовать метод `execute()`, чтобы отправить оператор в базу данных, которая вставит запись в таблицу.
connection=engine.connect()
result=connection.execute(ins)
# Мы также можем получить идентификатор вставленной записи, обратившись к атрибуту `inserted_primary_key`.
result.inserted_primary_key
# Обратите внимание, что значения `beds` и `bedrooms` в таблице `am` веществнного типа, в то время, как в базе данных эти поля целочисленного типа. В данном случае все работает корректно, так как параметрам `bedrooms` и`beds` значения присваиваются в явном виде, но если их получать из столбцов `bedrooms` и `beds` таблицы `df`, то в таблице `df` потребуется преобразование типов, его можно выполнить (с предварительной очисткой данных) так:
#
# `
# df['bedrooms']=df['bedrooms'].fillna(0)
# df['bedrooms']=pd.to_numeric(df['bedrooms'], downcast='integer')
# df['beds']=df['beds'].fillna(0)
# df['beds']=pd.to_numeric(df['beds'], downcast='integer')`
# Метод `execute()` использует оператор `insert` и другие параметры для компиляции SQL-выражения с помощью компилятора соответствующего диалекта базы данных. Этот компилятор строит нормальное параметризованное выражение и возвращается в метод `execute`, который отправляет оператор SQL в базу данных через соответствующее соединение. Затем сервер базы данных выполняет оператор и возвращает результат операции.
# <img src=./Img/Execute_method.png alt="Execute method" width="350">
# В дополнение к вставке в качестве метода экземпляра объекта `Table`, `insert` также доступен как функция верхнего уровня для случаев, когда таблица изначально неизвестна. Например, компания airbnb дополнительно к объектам в Амстердаме может иметь еще одну таблицу для Лиона (Франция): `listing_lyon`. Использование функции вставки позволяет использовать один оператор и подменять таблицы.
#
# `ins=insert(listing).values(
# listing_id=20168,
# listing_name='Studio with private bathroom in the centre 1',
# listing_url='https://www.airbnb.com/rooms/20168',
# host_id=59484,
# neighbourhood_id=1,
# property_type_id=1,
# room_type_id=1,
# amenities=df.loc[0,'amenities'],
# bedrooms=1,
# beds=1,
# price=236
# )`
lyon=pd.read_csv('ListingsLyon.csv',sep=';')
lyon.head(2)
lyon.info()
lyon.loc[0]
lyon_neigh=pd.DataFrame(lyon['neighbourhood_cleansed'].value_counts()).sort_index().reset_index()
lyon_neigh.index=range(1,lyon_neigh.shape[0]+1)
lyon_neigh
lyon_prop=pd.DataFrame(lyon['property_type'].value_counts()).sort_index().reset_index()
lyon_prop.index=range(1,len(lyon_prop)+1)
lyon_prop
lyon_room=pd.DataFrame(lyon['room_type'].value_counts()).sort_index().reset_index()
lyon_room.index=range(1,len(lyon_room)+1)
lyon_room
listing_lyon=Table('listing_lyon',metadata,
Column('listing_id',Integer(),primary_key=True),
Column('listing_name',String(50),index=True),
Column('listing_url',String(255)),
Column('host_id',Integer()),
Column('neighbourhood_id',Integer()),
Column('amenities',String(300)),
Column('property_type_id',Integer()),
Column('room_type_id',Integer()),
Column('bedrooms',Integer()),
Column('beds',Integer()),
Column('price',Numeric(7,2)),
CheckConstraint('price >= 0.00', name='lyon_price_positive'),
extend_existing=True
)
metadata.create_all(engine)
# +
#2-3. Функция insert
ins=insert(listing_lyon).values(
listing_id=int(lyon.loc[0,'id']),
listing_name=lyon.loc[0,'name'],
listing_url=lyon.loc[0,'listing_url'],
host_id=int(lyon.loc[0,'host_id']),
neighbourhood_id=5,
amenities=lyon.loc[0,'amenities'][:300],
property_type_id=1,
room_type_id=1,
bedrooms=int(lyon.loc[0,'bedrooms']),
beds=int(lyon.loc[0,'beds']),
price=lyon.loc[0,'price']
)
print(str(ins))
print(ins.compile().params)
# -
result=connection.execute(ins)
result.inserted_primary_key
# Метод `execute` объекта `connection` может принимать значения в качестве именованных аргументов, которые передаются после выражения. Когда выражение компилируется, он добавляет названия именованных аргументов в список столбцов, а каждое из их значений в часть ЗНАЧЕНИЯ оператора SQL.
# района, тип собственности, тип комнаты для вставляемой записи
print(am.loc[1,'id'])
print(am.loc[1,'neighbourhood_cleansed'])
print(am.loc[1,'property_type'])
print(am.loc[1,'room_type'])
# +
# 2-4
ins = listing.insert()
result = connection.execute(
ins,
listing_id=int(am.loc[1,'id']),
listing_name=am.loc[1,'name'],
listing_url=am.loc[1,'listing_url'],
host_id=int(am.loc[1,'host_id']),
neighbourhood_id=5,
amenities=am.loc[1,'amenities'][:300],
property_type_id=30,
room_type_id=3,
bedrooms=int(am.loc[1,'bedrooms']),
beds=int(am.loc[1,'beds']),
price=am.loc[1,'price']
)
result.inserted_primary_key
# -
# Хотя такой способ не часто используется на практике для одиночных вставок, он дает иллюстрацию компиляции и сборки оператора перед отправкой на сервер базы данных. Мы можем вставить сразу несколько записей, используя список словарей с данными. Воспользуемся этими знаниями, чтобы вставить еще две записи в таблицу listing (Пример 2-5).
for rec in range(2,4):
for column in ['id','neighbourhood_cleansed','property_type','room_type']:
print(am.loc[rec,column])
print()
listing_list=[
{
'listing_id' : int(am.loc[2,'id']),
'listing_name' : am.loc[2,'name'],
'listing_url' : am.loc[2,'listing_url'],
'host_id' : int(am.loc[2,'host_id']),
'neighbourhood_id' : 4,
'amenities' : am.loc[2,'amenities'][:300],
'property_type_am' : 19,
'room_type_id' : 3,
'bedrooms' : int(am.loc[2,'bedrooms']),
'beds' : int(am.loc[2,'beds']),
'price' : am.loc[2,'price']
},
{
'listing_id' : int(am.loc[3,'id']),
'listing_name' : am.loc[3,'name'],
'listing_url' : am.loc[3,'listing_url'],
'host_id' : int(am.loc[3,'host_id']),
'neighbourhood_id' : 4,
'amenities' : am.loc[3,'amenities'][:300],
'property_type_am' : 19,
'room_type_id' : 3,
'bedrooms' : int(am.loc[3,'bedrooms']),
'beds' : int(am.loc[3,'beds']),
'price' : am.loc[3,'price']
}
]
result=connection.execute(ins,listing_list)
# Словари в списке должны иметь одинаковые ключи. SQLAlchemy компилирует выражение с первым словарем в списке и завершится ошибкой, если последующие словари будут другими.
# ### Запросы к данным
#
# Чтобы начать построение запроса, мы начнем с использования функции выбора, которая аналогична стандартному оператору SQL SELECT. Для начала выберем все записи в нашей таблице файлов listing.
from sqlalchemy.sql import select
s = select([listing])
rp = connection.execute(s)
results = rp.fetchall()
results
|
Course II/Python_SQL/pract/pract3/T01_Core/T01_Core.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Get data via SQLAlchemy
import sqlalchemy, pickle
from sqlalchemy.ext.declarative import declarative_base
from sqlalchemy import Column, Integer, String
from sqlalchemy.orm import sessionmaker
from sqlalchemy import and_, or_
import MySQLdb, pickle
import pandas as pd
# ### Access to MySQL Server
pw = pickle.load(open("../crawler/mysql_pw.pickle", "rb"))
engine = sqlalchemy.create_engine("mysql+mysqldb://root:" + pw + "@192.168.3.11/project_rookie")
# ### ORM
Base = declarative_base()
def disp(datas):
for data in datas:
print(data)
def orm_to_df(orm):
df = pd.DataFrame(columns=['artist', 'album','freq_billboard'])
for i in orm:
data = {
'artist' : i.__dict__['artist'],
'album' : i.__dict__['album'],
'freq_billboard' : i.__dict__['freq_billboard'],
}
df.loc[len(df)] = data
return df
# ### ORM - Debut Album
class Debut(Base):
__tablename__ = 'debut_album'
index = Column(Integer, primary_key=True)
artist = Column(String)
album = Column(String)
genre = Column(String)
single_count = Column(Integer)
def __init__(self, index, artist, album, genre, single_count):
self.index = index
self.artist = artist
self.album = album
self.genre = genre
self.single_count = single_count
def __repr__(self):
return "<index {}, {}, {}, {}, {}>".format(self.index, self.artist, self.album, self.genre, self.single_count)
Base.metadata.create_all(engine)
Session = sessionmaker(bind=engine)
session = Session()
results = session.query(Debut).all()
results[0].__dict__
hiphop = session.query(Debut).filter(and_(~Debut.genre.contains(['K-pop']), Debut.genre.contains(["hop"])))
hiphop_df = orm_to_df(hiphop)
print(hiphop_df.shape)
hiphop_df.head()
rap = session.query(Debut).filter(and_(~Debut.genre.contains(['K-pop']), Debut.genre.contains(["Rap"])))
rap_df = orm_to_df(rap)
print(rap_df.shape)
rap_df.head()
trap = session.query(Debut).filter(and_(~Debut.genre.contains(['K-pop']), Debut.genre.contains(["trap"])))
trap_df = orm_to_df(trap)
print(trap_df.shape)
trap_df.head()
hiphop_df = pd.concat([hiphop_df, rap_df, trap_df])
hiphop_df.drop_duplicates(['artist', 'album'], inplace=True)
hiphop_df['genre'] = "hiphop"
hiphop_df
rnb = session.query(Debut).filter(and_(~Debut.genre.contains(['K-pop']), Debut.genre.contains(["R&B"])))
rnb_df = orm_to_df(rnb)
rnb_df['genre'] = 'rnb'
print(rnb_df.shape)
rnb_df.head()
soul = session.query(Debut).filter(and_(~Debut.genre.contains(['K-pop']), Debut.genre.contains(["soul"])))
soul_df = orm_to_df(soul)
soul_df['genre'] = 'soul'
print(soul_df.shape)
soul_df.head()
funk = session.query(Debut).filter(and_(~Debut.genre.contains(['K-pop']), Debut.genre.contains(["funk"])))
funk_df = orm_to_df(funk)
funk_df['genre'] = 'funk'
print(funk_df.shape)
funk_df.head()
pop = session.query(Debut).filter(and_(~Debut.genre.contains(['K-pop']), Debut.genre.contains(["pop"])))
pop_df = orm_to_df(pop)
pop_df['genre'] = 'pop'
print(pop_df.shape)
pop_df.head()
# ### Concat all genres and drop duplicates rows (overlapping genres)
df_list = pd.concat([hiphop_df, rap_df, trap_df, rnb_df, soul_df, funk_df, pop_df])
len(df_list)
df_list = df_list.drop_duplicates(['artist', 'album'])
print(df_list.shape)
df_list.head()
# ### Save to csv file
df_list.to_csv('../data/final_list_debut_album.csv', index=False)
|
merging_data/target_value_labeling.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # gtburst
#
# This tutorial provides a step-by-step guide to using the **gtburst** GUI for GRB and solar flare analysis of GBM and LAT data.
# ## What it is used for?
#
# Gtburst can be used to do the following:
# * GBM data:
# * download data from the GBM Trigger catalog, select data analysis interval, and interactively fit the background
# * write out pha and rsp files for spectral analysis in either XSPEC or RMFIT
#
# * LLE data:
# * download data from the Fermi LAT Low-Energy Events Catalog, select data analysis interval, and interactively fit the background
# * write out pha and rsp files for spectral analysis in either XSPEC or RMFIT
#
# * LAT data:
# * download photon events and spacecraft data from the LAT Data server
# * produce navigation plots to allow user to select optimal time intervals and zenith cuts
# * do photon selections based on energy, Region Of Interest (ROI), time, zenith, event class
# * produce counts maps
# * do likelihood analysis given a simple spectral model and background models
# * localization using **gtfindsrc** or a TS map
# * writes out pha and rsp files for spectral analysis in either XSPEC or RMFIT
# ## Prerequisites:
#
# * Internet connection
# * Recent version of **gtburst**
# * Latest installation of the Fermitools
# # Updating gtburst
#
# Before starting this analysis thread, make sure you have the latest version of **gtburst**. **gtburst** adopts a "release early, release often" model, thus it is a good habit to check often for updates. **gtburst** can be easily updated via the Update functionality. `Menu: Update -> Update` to the latest version.
#
# <img src='https://fermi.gsfc.nasa.gov/ssc/data/analysis/scitools/images/gtburst/image01.png'>
#
# This syncs with the [github repository](https://github.com/giacomov/gtburst). You may need to sync twice initially to get to the latest version. After each update, you'll need to restart gtburst, which is automatically prompted.
# # Immediate help from the interface
#
# * At any step in any analysis task, you can read in the lower left corner a description on what you are supposed to do and some hints about things to keep in mind.
#
# * If you don't remember what a given parameter in a command means, you can always click on the question mark "?" on the right of the parameter to get a short description of its meaning.
# # Analysis of a Gamma-Ray Burst
#
# For this thread, we will analyze GRB080916C, one of the brightest LAT GRBs on record. All outputs from GUI window are in `gtburst.log`.
#
# Steps:
#
# 1. To start using **gtburst** for any analysis
# * Open a terminal, and cd into a directory where you wish the gtburst output files to go. You may wish to make a sub directory specific to the object being analyzed to keep files organized (e.g. GRB080916C).
# * Type **gtburst** at command line.
#
#
# 2. Downloading data
#
# **gtburst** can download GBM, LLE, and LAT data from the FSSC, can be run on datasets previously downloaded, or can be loaded as a custom dataset.
# !gtburst
# <img src='https://fermi.gsfc.nasa.gov/ssc/data/analysis/scitools/images/gtburst/image02.png'>
#
# Download datasets ...
#
# <img src='https://fermi.gsfc.nasa.gov/ssc/data/analysis/scitools/images/gtburst/image03.png'>
# The trigger details can either be entered manually, or retrieved from the trigger catalog which includes both Swift-BAT and Fermi-GBM triggers.
#
# Click `Browse triggers`
#
# <img src='https://fermi.gsfc.nasa.gov/ssc/data/analysis/scitools/images/gtburst/image04.png'>
# Triggers can be sorted by any of the columns by clicking on the header, or filtered by trigger type.
#
# Note: GBM triggers may not appear immediately in this database, as they depend on the GBM team populating the table, which is done within 48 hours of the trigger.
#
# Select the trigger and click `Done` (bn080916009 for GRB 080916C in this example).
#
# A box will pop up to confirm data selection. Time Tagged Events (TTE) versus binned CTIME data. TTE data files have finer time resolution and are therefore larger, and can be binned as needed.
#
# <img src='https://fermi.gsfc.nasa.gov/ssc/data/analysis/scitools/images/gtburst/image05.png'>
# 3. GBM & LLE Data Analysis
#
# * A box will pop up to select the datasets for the following analysis. The number of degrees in parentheses is the angle from the individual detector boresight to the GRB. By default, the 3 or 4 smallest angle NaI's, the 1 smallest angle BGO, and LAT & LLE data will be selected. The backgrounds will be fit and spectral files created for only the selected detectors. The user may select additional detectors.
#
# <img src='https://fermi.gsfc.nasa.gov/ssc/data/analysis/scitools/images/gtburst/image06.png'>
# * Make spectra for XSPEC - will walk user through background and source selection, and output rsp & pha files.
#
# <img src='https://fermi.gsfc.nasa.gov/ssc/data/analysis/scitools/images/gtburst/image08.png'>
# * By default, the user will click to select the time intervals for source and background selection. The source interval will be chosen only once for the first detector used, which is the smallest angle NaI by default (n3 in this example).
#
# Click `Run` to select interactively.
#
# <img src='https://fermi.gsfc.nasa.gov/ssc/data/analysis/scitools/images/gtburst/image09.png'>
# * Zoom into the GRB emission using the <img src='https://fermi.gsfc.nasa.gov/ssc/data/analysis/scitools/images/gtburst/image10.png'> button, and drawing a box around the GRB emission. Click the button again to exit the zoom feature.
#
# At any time, click <img src='https://fermi.gsfc.nasa.gov/ssc/data/analysis/scitools/images/gtburst/image11.png'> to zoom back out. You can repeatedly refine the box smaller.
#
# Once zoomed sufficiently, click once at the beginning of the interval, and again at the end. Once you are happy with the selection, click `Done`.
#
# <img src='https://fermi.gsfc.nasa.gov/ssc/data/analysis/scitools/images/gtburst/image12.png'>
# * Background fitting - click next, and then `Run` to interactively fit the background. Now zoom in to a region around the burst emission to fit the background. A few hundred seconds or so before and after the burst is usually sufficient. Select one interval prior to the burst (by clicking at the beginning and end) and another interval after the burst, where the background is approximately flat. Then click `Done`.
#
# <img src='https://fermi.gsfc.nasa.gov/ssc/data/analysis/scitools/images/gtburst/image13.png'>
# * An automated fit to the background, which fits various polynomials is performed, to find the best fit. A plot showing the background-subtracted light curve is produced. If you are happy with the result, click `OK`, otherwise `Retry` to select new background intervals.
#
# <img src='https://fermi.gsfc.nasa.gov/ssc/data/analysis/scitools/images/gtburst/image14.png'>
# * You will now need to produce a background fit for each subsequent detector. The same source interval is assumed. Redo e-f for each detector including BGO & LLE.
#
# * When all detectors are completed, click Next, and then Run to produce the output files, and Finish to go back to the starting menu. The directory where you're running gtburst should now contain the following files, which can be read into XSPEC to conduct a joint spectral fit as described in http://fermi.gsfc.nasa.gov/ssc/data/analysis/scitools/combined_LAT_GBM.html
# ```
# bn080916009_n3_srcspectra.pha
# bn080916009_n3_weightedrsp.rsp
# bn080916009_n3_bkgspectra.bak
# bn080916009_n4_srcspectra.pha
# bn080916009_n4_weightedrsp.rsp
# bn080916009_n4_bkgspectra.bak
# bn080916009_n6_srcspectra.pha
# bn080916009_n6_weightedrsp.rsp
# bn080916009_n6_bkgspectra.bak
# bn080916009_b0_srcspectra.pha
# bn080916009_b0_weightedrsp.rsp
# bn080916009_b0_bkgspectra.bak
# bn080916009_LAT-LLE_srcspectra.pha
# bn080916009_LAT-LLE_weightedrsp.rsp
# bn080916009_LAT-LLE_bkgspectra.bak
# ```
# 4. LAT Data Analysis
# * After downloading a dataset or loading data from a directory, it's best to start with the navigation plot.
#
# Click Tools->Make navigation plots
#
# <img src='https://fermi.gsfc.nasa.gov/ssc/data/analysis/scitools/images/gtburst/image15.png'>
#
# The navigation plots will pop up in a separate window. The upper plot is the angle between the GRB RA/Dec and the Fermi zenith angle. This indicates when the GRB is and is not occulted by the Earth. A zenith angle cut of 100 is fairly standard for a GRB, but can be adjusted slightly higher if the source is very bright at that time, or lower if the source is fainter and the user is concerned about Earth limb contamination.
#
# The lower panel of the navigation plot is the angle between the GRB RA/Dec and the boresight of the LAT. This indicates when the source is within the LAT FoV. The size of the LAT FoV is dependent on energy and event class.
#
# The navigation plots are in reference to the GRB localization in the GBM GRB catalog, which may be the best available GBM position (~few deg), an announced LAT position (~0.1-1 deg), or a much more accurate position from follow-up (~arcsec). The user can manually adjust the position in the GUI window at this time, or later based upon the counts map. If the user changes the R.A. and Dec. in the initial window, making the navigation plots again will update the plots using the new position.
#
# <img src='https://fermi.gsfc.nasa.gov/ssc/data/analysis/scitools/images/gtburst/image16.png'>
# ***
# * Likelihood Analysis
#
# Click `Tasks->Make likelihood analysis`
#
# <img src='https://fermi.gsfc.nasa.gov/ssc/data/analysis/scitools/images/gtburst/image17.png'>
# <img src='https://fermi.gsfc.nasa.gov/ssc/data/analysis/scitools/images/gtburst/image18.png'>
#
# The first step is filtering for the counts map, which can be repeated and optimized. The parameters:
# rad - radius of interest (degrees). Customary values corresponds to the 95% containment of the PSF at the emin energy. If you use `emin=100` MeV, rad should be 12 deg for any Transient class and 10 deg for Source or cleaner classes.
#
# irf - the event class - in GRB analysis transient class is usually sufficient for short (<100 s) timescales and spectral analysis, and source class is better for longer intervals and localizations. Event Class recommendations for different analyses are discussed at: http://fermi.gsfc.nasa.gov/ssc/data/analysis/documentation/Cicerone/Cicerone_Data/LAT_DP.html
#
# zmax - Zenith angle cut. If the parameter `strategy=time` (default), any time interval where any part of the ROI is at a Zenith angle larger than zmax is excluded. This is the normal choice for GRB and SF analysis. If `strategy=events`, all events with a Zenith angle larger than zmax will be excluded. The user is strongly advised against using `strategy=events`, unless he/she understands exactly its implication, since such choice can introduce systematic uncertainties in the analysis difficult to estimate.
#
# Tstart - time to start analysis relative to GRB trigger. This can be specified either as a time from the trigger time, or as a Mission Elapsed Time (MET). The interface will automatically understand, since MET numbers are very big.
#
# Tstop - time to stop analysis relative to GRB trigger. This can be specified either as a time from the trigger time, or as a Mission Elapsed Time (MET). The interface will automatically understand, since MET numbers are very big.
#
# Emin - minimum energy for analysis in MeV. Normal value is 100 MeV, as going below that requires special attention.
#
# Emax - maximum energy for analysis in MeV
#
# Skybinsize - binning for map
#
# Thetamax - This is an additional cut which will exclude from the analysis time intervals in which the position of the source is at more than Thetamax degrees from the center of the LAT field of view. Since the PSF of the LAT becomes worse and more uncertain at high off-axis angles, this can be used when analyzing bright bursts to reduce the errors on the localization. It is usually not necessary to change this value.
#
# Strategy - method of zenith angle cut
# Using `strategy=time` (the standard value) will exclude from the analysis all time intervals in which any part of the ROI is at Zenith angles larger than the zmax value.
#
# Do not change this value unless you know what you are doing! Using `strategy=events` will exclude from the analysis all events with a Zenith angle larger than zmax, which can introduce subtle systematic errors in the analysis difficult to judge.
#
# Click `Run`:
#
# <img src='https://fermi.gsfc.nasa.gov/ssc/data/analysis/scitools/images/gtburst/image19.png'>
# You will then see the resulting counts map and photon energy as a function of time.
#
# You can click on photons on the right plot, which will be highlighted on the left plot with a small white circle. This is helpful for determining if a particular high energy photon is clustered near others. A small text box will also appear with the ID of the run in which the event was detected, the event ID, the Zenith and the off-axis (theta) angle of that event.
#
# You can also zoom in the left plot, and only the photons within your zoomed area will remain in the right plot. This is useful for example to figure out which photons are close to the source position. Note that if you zoom in the right plot, the left plot will NOT change since this would require a new run of the command.
#
# Click `Next`:
# <img src='https://fermi.gsfc.nasa.gov/ssc/data/analysis/scitools/images/gtburst/image20.png'>
# Next we choose the components of our likelihood model. This command will produce a XML file containing your likelihood model, as described here ([link_to_likelihood_tutorial]). For source class, a particle model of isotr template is appropriate. The other defaults are sufficient. Click `Run`; once that finishes, click `Next`.
#
# Gtburst will automatically add nearby bright catalog sources to your XML file. Once the dialog box finishes, click `Run`.
#
# A window summarizing the fit parameters of the model will pop up. You can modify the parameters (e.g. fixing index to some value), or simply click `Done` to leave everything as it is.
#
# If you make changes, be sure to click `Save`, and click `Done` when finished. Then, once the window with the list of parameters is closed, click `Next`.
#
# <img src='https://fermi.gsfc.nasa.gov/ssc/data/analysis/scitools/images/gtburst/image21.png'>
# Now you will be given options on the outputs of the likelihood analysis.
#
# `Optimizeposition=yes` will call **gtfindsrc** at the end of the likelihood analysis and attempt to improve the GRB localization.
#
# `Showmodelimage=yes` will create a model map and display it. This does not have any impact on the actual analysis, but allow you to see a representation of your final model.
#
# `Spectralfiles=yes` will create the pha and rsp files necessary to do spectral analysis in XSPEC or rmfit.
#
# Each of these steps will make the analysis take longer. Click `Run`.
#
# <img src='https://fermi.gsfc.nasa.gov/ssc/data/analysis/scitools/images/gtburst/image22.png'>
# Results:
#
# Likelihood fit result parameters of GRB, relevant nearby sources, and background models. You must close this window to proceed.
#
# <img src='https://fermi.gsfc.nasa.gov/ssc/data/analysis/scitools/images/gtburst/image23.png'>
# Resulting count map and likelihood model image. GRB 080916C is well detected!
#
# <img src='https://fermi.gsfc.nasa.gov/ssc/data/analysis/scitools/images/gtburst/image24.png'>
#
# If a substantially improved position is available, enter this position in the start window, which can be reached by clicking Finish, then repeat likelihood analysis from that position.
# ***
# * TS map - If photon clustering isn't entirely obvious or to potentially improve the localization further, one can create a TS map, which can take a while to run, especially on a long dataset. For very bright GRBs, note that the binning may be more of a limiting factor then the photon statistics on localization determination.
#
# `Tasks->Find source` in TS Map
#
# Follow similar steps as the likelihood analysis, ending up with a map and localization. For GRB 080916C, the localization is not improved by the TS map because the statistical error is smaller than the TS map binsize.
# <img src='https://fermi.gsfc.nasa.gov/ssc/data/analysis/scitools/images/gtburst/image25.png'>
# * Choose new center for likelihood analysis, useful for localizing a GBM burst, based upon visual determination of the position of the cluster of photons in the counts map.
#
# Tasks -> Interactively recenter ROI
#
# Make photon selections, click `Run`. Once it finishes, click `Next`, and `Run`.
#
# Click on a new center position on the left counts map. Click `Run`, and finish. Then repeat the likelihood analysis step at this new location.
|
GRBAnalysis/3.GRBAnalysisGTBurst/3.GRBAnalysisGTBurst.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ### Copyright (C) 2022 <NAME> and s.o who's unknown :)
# ### Date: 2/20/2022
# + id="mBMoPLmGbrIn"
from amalearn.reward import RewardBase
from amalearn.agent import AgentBase
from amalearn.environment import EnvironmentBase
from matplotlib.patches import Rectangle
import matplotlib.pyplot as plt
import numpy as np
import gym
# -
# + id="pH6sNHxPbrIs"
class Environment(EnvironmentBase):
obstacles = [(1, 7), (1, 8), (2, 7), (2, 8), (3, 7), (3, 8), (4, 7), (4, 8),
(12, 6), (12, 7), (13, 6), (13, 7), (14, 6), (14, 7), (15, 6), (15, 7),
(8, 13), (8, 14), (8, 15), (9, 13), (9, 14), (9, 15)]
def __init__(self, actionPrice, goalReward, punish, obstacles=obstacles,
i_limit=15, j_limit=15, p=0.8, goal=(1, 1), start=(15, 15), container=None):
""" initialize your variables """
state_space = gym.spaces.MultiDiscrete([i_limit, j_limit])
action_space = gym.spaces.Discrete(9)
super(Environment, self).__init__(action_space, state_space, container)
self.state_space = state_space
self.obstacles = obstacles
self.actionPrice = actionPrice
self.goalReward = goalReward
self.punish = punish
self.i_limit = i_limit
self.j_limit = j_limit
self.p = p
self.goal = goal
self.start = start
self.state = start
self.state_p = None
# -------------------------------------------------------------------------------------------------------------
def isStatePossible(self, state):
"""if given state is possible (not out of the grid and not obstacle) return ture"""
i_in = range(1, self.i_limit + 1)
j_in = range(1, self.j_limit + 1)
return False if state in self.obstacles or state[0] not in i_in or state[1] not in j_in else True
# -------------------------------------------------------------------------------------------------------------
def isAccessible(self, state, state_p):
"""if given state is Accesible (we can reach state_p by doing an action from state) return true"""
if self.isStatePossible(state) and self.isStatePossible(state_p):
if (np.abs(np.subtract(state, state_p)) <= (1, 1)).all():
return True
return False
# -------------------------------------------------------------------------------------------------------------
def getTransitionStatesAndProbs(self, state, action, state_p):
"""return probability of transition or T(sp,a,s)"""
_, actions = self.available_actions(state)
if action in actions:
available_states = self.available_states(actions, state)
if self.next_state(action, state) == state_p:
return self.p
elif state_p in available_states:
return (1 - self.p) / (len(available_states) - 1)
else: return 0
else: return 0
# -------------------------------------------------------------------------------------------------------------
def getReward(self, state, action, state_p):
"""return reward of transition"""
# The Goal Achieved
if state_p == self.goal:
return self.goalReward
elif self.isAccessible(state, state_p):
return self.actionPrice
# Hit the obstacles
else:
return self.punish
# -------------------------------------------------------------------------------------------------------------
def calculate_reward(self, action):
return
# -------------------------------------------------------------------------------------------------------------
def terminated(self, state):
return state == self.goal
# -------------------------------------------------------------------------------------------------------------
def available_actions(self, state):
actions = []
numbers = []
num = 0
for i in range(-1, 2):
for j in range(-1, 2):
state_p = tuple(np.add(state, (j, i)))
if self.isAccessible(state, state_p):
actions.append((j, i))
numbers.append(num)
num += 1
return numbers, actions
# -------------------------------------------------------------------------------------------------------------
def action_num2dim(self, num):
if num < 3:
return (np.mod(num, 3) -1, -1)
elif num < 6:
return (np.mod(num, 3) -1, 0)
else:
return (np.mod(num, 3) -1, 1)
# -------------------------------------------------------------------------------------------------------------
def get_states(self):
states = []
for i in range(1, self.i_limit + 1):
for j in range(1, self.j_limit + 1):
if self.isStatePossible((i, j)):
states.append((i, j))
return states
# -------------------------------------------------------------------------------------------------------------
def available_states(self, actions, state):
states_p = []
for action in actions:
states_p.append(self.next_state(action, state))
return states_p
# -------------------------------------------------------------------------------------------------------------
def next_state(self, action, state):
return tuple(np.add(state, action))
# -------------------------------------------------------------------------------------------------------------
def reset(self):
self.state = self.start
# -------------------------------------------------------------------------------------------------------------
def observe(self):
return self.state
# -------------------------------------------------------------------------------------------------------------
def render(self):
return
# -------------------------------------------------------------------------------------------------------------
def close(self):
return
# -
# + id="898Jlhsycyes"
class Agent(AgentBase):
def __init__(self, environment, theta=0.1, discount=0.9, free_stay=False):
#initialize a random policy and V(s) = 0 for each state
self.environment = environment
self.width = self.environment.i_limit
self.height = self.environment.j_limit
#init V
self.V = [[0] * (self.width + 1) for _ in range(self.height + 1)]
#init policy
self.policy = np.random.randint(0, 9, (self.width + 1, self.height + 1))
super(Agent, self).__init__(id, environment)
self.discount = discount
self.theta = theta
self.free_stay = free_stay
# -------------------------------------------------------------------------------------------------------------
def policy_evaluation(self):
while True:
delta = 0
pre_delta = delta
for state in self.environment.get_states():
v = self.V[state[0]][state[1]]
action = self.policy[state[0]][state[1]]
numbers, actions = self.environment.available_actions(state)
value = 0
for act in actions:
state_p = self.environment.next_state(act, state)
if self.free_stay:
reward = 0 if act == (0, 0) else self.environment.getReward(state, environment.action_num2dim(action), state_p)
else:
reward = self.environment.getReward(state, environment.action_num2dim(action), state_p)
probability = self.environment.getTransitionStatesAndProbs(state, environment.action_num2dim(action), state_p)
value += probability * (reward + self.discount * self.V[state_p[0]][state_p[1]])
self.V[state[0]][state[1]] = value
pre_delta = delta
delta = max([delta, np.abs(v - self.V[state[0]][state[1]])])
if delta < self.theta or delta == pre_delta:
break
return self.V
# -------------------------------------------------------------------------------------------------------------
def policy_improvement(self):
unchanged = True
for state in self.environment.get_states():
pre_action = self.policy[state[0]][state[1]]
acts = []
numbers, actions = self.environment.available_actions(state)
for _, act1 in zip(numbers, actions):
value = 0
for _, act2 in zip(numbers, actions):
state_p = self.environment.next_state(act2, state)
reward = self.environment.getReward(state, act1, state_p)
probability = self.environment.getTransitionStatesAndProbs(state, act1, state_p)
value += probability * (reward + self.discount * self.V[state_p[0]][state_p[1]])
acts.append(value)
best_act = np.argmax(acts)
self.policy[state[0]][state[1]] = numbers[best_act]
if numbers[best_act] != pre_action:
unchanged = False
return unchanged
# -------------------------------------------------------------------------------------------------------------
def value_opt_func(self):
while True:
delta = 0.1
pre_delta = delta
for state in self.environment.get_states():
acts = []
numbers, actions = self.environment.available_actions(state)
for _, act1 in zip(numbers, actions):
value = 0
for _, act2 in zip(numbers, actions):
state_p = self.environment.next_state(act2, state)
reward = self.environment.getReward(state, act1, state_p)
probability = self.environment.getTransitionStatesAndProbs(state, act1, state_p)
value += probability * (reward + self.discount * self.V[state_p[0]][state_p[1]])
acts.append(value)
best_act = np.max(acts)
pre_delta = delta
delta = max([delta, np.abs(best_act - self.V[state[0]][state[1]])])
self.V[state[0]][state[1]] = best_act
if delta < self.theta or delta == pre_delta:
break
return self.V
# -------------------------------------------------------------------------------------------------------------
def value_extraction(self):
for state in self.environment.get_states():
pre_action = self.policy[state[0]][state[1]]
acts = []
numbers, actions = self.environment.available_actions(state)
for _, act1 in zip(numbers, actions):
value = 0
for _, act2 in zip(numbers, actions):
state_p = self.environment.next_state(act2, state)
reward = self.environment.getReward(state, act1, state_p)
probability = self.environment.getTransitionStatesAndProbs(state, act1, state_p)
value += probability * (reward + self.discount * self.V[state_p[0]][state_p[1]])
acts.append(value)
best_act = np.argmax(acts)
self.policy[state[0]][state[1]] = numbers[best_act]
# -------------------------------------------------------------------------------------------------------------
def policy_iteration(self):
unchanged = False
while not unchanged:
self.V = self.policy_evaluation()
unchanged = self.policy_improvement()
# -------------------------------------------------------------------------------------------------------------
def value_iteration(self):
self.V = self.value_opt_func()
self.value_extraction()
# -------------------------------------------------------------------------------------------------------------
def take_action(self, mode='policy') -> (object, float, bool, object):
if mode == 'policy':
self.policy_iteration()
elif mode == 'value':
self.value_iteration()
# -------------------------------------------------------------------------------------------------------------
def visualize_policy(self):
plt.gcf().set_size_inches(5, 5)
ax = plt.gca()
ax.set_xticks(range(1, environment.i_limit + 1))
ax.set_yticks(range(environment.j_limit, 0, -1))
ax.axes.xaxis.set_ticklabels([])
ax.axes.yaxis.set_ticklabels([])
for i in range(environment.i_limit, 0, -1):
temp = (environment.i_limit + 1) - i
for j in range(environment.j_limit, 0, -1):
num = agent.policy[i, j]
plt.gca().text(j - 0.5, temp - 0.5, str(num), va='center', ha='center')
plt.gca().add_patch(Rectangle((j, temp), -1, -1, fill=True, color='white', alpha=0.5))
plt.grid(True)
plt.show()
# -------------------------------------------------------------------------------------------------------------
def visualize_values(self):
value = self.V
plt.gcf().set_size_inches(7, 7)
ax = plt.gca()
ax.set_xticks(range(1, environment.i_limit + 1))
ax.set_yticks(range(environment.j_limit, 0, -1))
ax.axes.xaxis.set_ticklabels([])
ax.axes.yaxis.set_ticklabels([])
for i in range(environment.i_limit, 0, -1):
temp = (environment.i_limit + 1) - i
for j in range(environment.j_limit, 0, -1):
c = round(value[i][j])
plt.gca().text(j - 0.5, temp - 0.5, str(c), va='center', ha='center')
plt.gca().add_patch(Rectangle((j, temp), -1, -1, fill=True, color='white', alpha=0.5))
plt.grid(True)
plt.show()
# -------------------------------------------------------------------------------------------------------------
def get_path(self):
dead = self.width * self.height
achived = False
path = []
states = []
start = self.environment.start
curr = start
count = 0
while count < dead:
num = self.policy[curr[0]][curr[1]]
path.append(num)
states.append(curr)
direction = self.environment.action_num2dim(num)
curr = tuple(np.add(curr, direction))
count += 1
if self.environment.terminated(curr):
achived = True
break
return path, states[1:], achived
# -
def plotter(environment, agent):
plt.gcf().set_size_inches(10, 10)
ax = plt.gca()
ax.set_xticks(range(1, environment.i_limit + 1))
ax.set_yticks(range(environment.j_limit, 0, -1))
ax.axes.xaxis.set_ticklabels([])
ax.axes.yaxis.set_ticklabels([])
path, states, achived = agent.get_path()
no = 1
for i in range(environment.i_limit, 0, -1):
temp = (environment.i_limit + 1) - i
for j in range(environment.j_limit, 0, -1):
if (i, j) == environment.start:
plt.gca().text(j - 0.5, temp - 0.5, str('S'), va='center', ha='center')
plt.gca().add_patch(Rectangle((j, temp), -1, -1, fill=True, color='purple', alpha=0.5))
if (i, j) in environment.obstacles:
plt.gca().add_patch(Rectangle((j, temp), -1, -1, fill=True, color='black', alpha=0.5))
elif (i, j) in states:
num = path[states.index((i, j))]
direction = environment.action_num2dim(num)
plt.gca().text(j - 0.5, temp - 0.5, str(no), va='center', ha='center')
plt.gca().add_patch(Rectangle((j, temp), -1, -1, fill=True, color='blue', alpha=0.5))
no += 1
elif (i, j) == environment.goal:
plt.gca().text(j - 0.5, temp - 0.5, str('F'), va='center', ha='center')
plt.gca().add_patch(Rectangle((j, temp), -1, -1, fill=True, color='green', alpha=0.5))
plt.grid(True)
plt.show()
# ### Q1.
environment = Environment(actionPrice=-0.01, goalReward=1000, punish=-1)
agent = Agent(environment)
agent.take_action()
plotter(environment, agent)
agent.visualize_policy()
# ### Q2.
environment = Environment(actionPrice=0, goalReward=1000, punish=-0.01)
agent = Agent(environment)
agent.take_action()
plotter(environment, agent)
agent.visualize_policy()
# ### Q3.
environment = Environment(actionPrice=-1, goalReward=100, punish=-10)
agent = Agent(environment, free_stay=True)
agent.take_action()
plotter(environment, agent)
agent.visualize_policy()
# ### Q4.
# +
discount_factors = [0.1, 0.01, 0.001, 0]
for discount in discount_factors:
print('|_ Discount Factor: {}'.format(discount))
environment = Environment(actionPrice=-0.01, goalReward=1000, punish=-1)
agent = Agent(environment, discount=discount)
agent.take_action()
plotter(environment, agent)
agent.visualize_policy()
# -
# ### Q5.
environment = Environment(actionPrice=-0.01, goalReward=1000, punish=-1)
agent = Agent(environment)
agent.take_action(mode='value')
plotter(environment, agent)
agent.visualize_policy()
# ### Extra.
# +
obstacles = [(1, 7), (1, 8), (2, 7), (2, 8), (3, 7), (3, 8), (4, 7), (4, 8),
(6, 4), (6, 5), (6, 6), (6, 7), (6, 8), (6, 9), (6, 10),
(7, 4), (7, 5), (7, 6), (7, 7), (7, 8), (7, 9), (7, 10),
(8, 4), (8, 5), (9, 4), (9, 5),
(12, 6), (12, 7), (13, 6), (13, 7), (14, 6), (14, 7), (15, 6), (15, 7),
(8, 13), (8, 14), (8, 15), (9, 13), (9, 14), (9, 15)]
environment = Environment(obstacles=obstacles, actionPrice=-0.01, goalReward=1000, punish=-1)
agent = Agent(environment)
agent.take_action()
plotter(environment, agent)
agent.visualize_policy()
# -
# ### Finito
|
FinalProj/ML_HW04.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # 9AR_omim_enrichment_dotplot
#
library(tidyverse)
omim_df = read.csv('omim_enrichment//mono_poly_dz_df_filt.csv',stringsAsFactor=F)
dim(omim_df)
omim_df = omim_df%>%
mutate(pval_adj = p.adjust(hypergeom_pval,'fdr'))%>%
filter( pval_adj<0.10)%>%
mutate(neglogpval = pmin(5,-log10(pval_adj)))
colnames(omim_df)
dim(omim_df)
length(unique(omim_df$mono_dz))
omim_df%>%group_by(poly_dz_abbr)%>%tally()
omim_df%>%select(-poly_geneset)
# +
ggplot(omim_df%>%filter(poly_dz_abbr!=''), aes(x=poly_dz_abbr, y=mono_dz, label= overlap))+
# geom_point()+
geom_text()+
# theme(axis.text.x=element_text(angle = -45, hjust = 0))+
# theme_classic()+
xlab('disease')+ylab('Rare/Mendelian diseases')+
ggtitle('Gene overlap between monogenic diseases and daSNP eGenes')# between monogenic neurological diseases and neuropsychiatric eGenes')
ggsave('omim_enrichment/mono_poly_dotplot.pdf',width=10, height=7)
# -
|
9AR_omim_enrichment_dotplot.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/jfaure3/.github.io/blob/master/Bases_de_Python.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="xclU-bKWADiU" colab_type="text"
# #Divers astuces et fonctions intéressantes
# + [markdown] id="R_BqShcQ-Hlt" colab_type="text"
# Ne pas aller à la ligne avec un print en spécifiant le caractère de fin.
#
# Ici end=' ' c'est une espace.
# + id="H4DDKPkq-Hl5" colab_type="code" colab={}
def affiche(mot='rien'):
"""pas de retour à la ligne, mais une espace"""
for I in range(3):
print(I,mot,end=' ')
affiche()
# + [markdown] id="Ofj1t2Gm-HmS" colab_type="text"
# Associer un caractère avec son point de code de façon biunivoque.
# + id="XRt-5kSh-HmW" colab_type="code" colab={}
def code_cesar(mot='Bonjour tout le monde', decalage=3):
"""César avec un décallage de 1"""
return "".join([chr(ord(c)+decalage) for c in mot])
code_cesar()
# + [markdown] id="wEjvDfkl-Hmo" colab_type="text"
# Ecrire des if en ligne.
# + id="Jft0W4rf-Hmt" colab_type="code" colab={}
def autre_ecriture_si(masse):
"""écrite de if en ligne"""
return 3*masse if masse<=20 else (2*masse if masse<50 else masse)
autre_ecriture_si(20)
# + [markdown] id="TsdKBGkOEToP" colab_type="text"
# Liste en compréhension avec des entrées.
# + id="Sc3A-jtD-HnJ" colab_type="code" colab={}
def si_en_ligne():
"""liste en compréhention"""
return sum([float(input('jour '+str(I)+' : ')) for I in range(7)])/7
si_en_ligne()
# + [markdown] id="P_1spp_5-HnU" colab_type="text"
# Liste de nombres pseudo-aléatoires, mais périodiques... (principe des tiroirs de Dirichlet)
# + id="sdb3B8pX-HnZ" colab_type="code" colab={}
import matplotlib.pyplot as plt
def pseudo_alea(n=100):
"""liste de nombres pseudo-aléatoires"""
liste=[13]
for I in range(n):
liste.append((16805*liste[I] + 1) % 32768)
plt.subplot(1,2,1)
plt.plot(liste,'.')
plt.title('nombres')
pileFace=[bin(nb)[-1] for nb in liste]
plt.subplot(1,2,2 )
plt.plot(pileFace,'.')
plt.title('bits de poids faibles')
plt.show()
pseudo_alea()
# + [markdown] id="hu8qnz-Y-Hnt" colab_type="text"
# ##Récursivité
# + id="olL0jfzF-Hnz" colab_type="code" colab={}
def puissance_de_2(n=10):
"""retourne la puissance n-ième de 2"""
return 1 if n==0 else 2*puissance_de_2(n-1)
puissance_de_2()
# + id="5MGkDRT7-HoB" colab_type="code" colab={}
def PGCD(a,b):
"""retourne le PGCD à partir de l'algorithme d'Euclide"""
return b if a%b==0 else PGCD(b,a%b)
PGCD(30,42)
# + id="8nwNDESB-HoW" colab_type="code" colab={}
def base_10_vers_n_recursif(nb=6007,n=5):
"""retourne l'écriture de nb en base n"""
return str(nb) if nb//n==0 else base_10_vers_n_recursif(nb//n,n)+str(nb%n)
base_10_vers_n_recursif(1020,2)
# + [markdown] id="rhI17JHr-Hos" colab_type="text"
# avec les listes différence entre le tri en place a.sort() et sorted(a) qui produit une autre liste
# + id="_z2_IGFT-Ho3" colab_type="code" colab={}
def deux_tris(liste=[2,1,8,6,5,7,5,6]):
"""illusrte le tri en place et le tri dans une autre liste"""
print(sorted(liste))#renvoie une autre liste
print(liste)
liste.sort()#en place
print(liste)
deux_tris()
# + [markdown] id="CDfKtLL2-HpJ" colab_type="text"
# L'encodage des caractères
# + id="5s-RJcDC-HpO" colab_type="code" colab={}
print("\N{GREEK CAPITAL LETTER DELTA}")
print("\u0394")
# + [markdown] id="u_1qmFag-Hpg" colab_type="text"
# Pour tester une égalité, l'astuce est de voir si les deux nombres sont \og vraiment \fg{} proches.
# + id="WtV5bkxe-Hpk" colab_type="code" colab={}
from math import *
def egalite(a=0.3,b=0.1+0.1+0.1):
"""retourne le test d'égalité stricte et le test de proximité"""
return (a==b, isclose(a,b))
egalite()
# + [markdown] id="f7cOhPVQ_rNA" colab_type="text"
# #Les graphiques avec matplotlib
# + [markdown] id="hV4qJ48s-Hpx" colab_type="text"
# Affichage de courbes avec matplotlib.pyplot
# + id="ToSYTXC--Hp1" colab_type="code" colab={}
import numpy as np
import matplotlib.pyplot as plt
def graphiques(f = lambda x : x**2):
"""tracés des courbes"""
X=np.linspace(-2,1.5,100)#100 valeurs de -2 à 1.5 dans un ndarray de numpy
Y=f(X)
plt.plot(X,Y,'r-')#- -- ^ s -. : b g r c m y k w , linewidth=0.8, marker="+", label="Trajet 2"
plt.ylabel('$C_f$')
plt.title("f en fonction de x")
plt.show()
graphiques()
# + [markdown] id="QU0fv4HK-HqJ" colab_type="text"
# Graphiques statistiques avec matplotlib
# + id="PjZ_qYWO-HqM" colab_type="code" colab={}
import numpy as np
import matplotlib.pyplot as plt
def graphiques():
"""graphique statistiques"""
x = [1,2,3,4,5]
donnees=[20,5,60,10,20]
noms=['A','B','C','D','E']
explode=(0, 0, 0.15, 0,0)#sépart la part de 'B'
plt.subplot(1,2,1)
plt.pie(donnees, explode=explode, labels=noms, autopct='%1.2f%%', startangle=90, shadow=True)
plt.axis('equal')# un camembert bien rond
plt.subplot(1,2,2)
plt.bar(x,donnees, width=0.5, color=(0.1,0.2,0.3, 1.0) )
plt.xticks(x,noms)
plt.grid()
plt.savefig('circulaire et barres.png')
plt.show()
graphiques()
# + [markdown] id="qX__TWMP-Hqk" colab_type="text"
# histogramme et diagramme en boite
# + id="mogRdL5c-Hqo" colab_type="code" colab={}
from random import *
import matplotlib.pyplot as plt
def graphiques(n=1000):
"""simule n fois la somme de 5 dés"""
X1=[randint(1,6)+randint(1,6)+randint(1,6)+randint(1,6)+randint(1,6) for I in range(n)]
X2=[randint(1,6)+randint(1,6)+randint(1,6)+randint(1,6)+randint(1,6)for I in range(n)]
plt.subplot(1,2,1)
plt.boxplot([X1,X2])
plt.subplot(1,2,2)
plt.hist(X1,bins=[0,10,15,18,20,22,25,31],color=(1,0,0,1))
plt.hist(X2,bins=[0,10,15,18,20,22,25,31],color=(0,0,1,0.5))
plt.grid()
plt.savefig('boite et histogramme.png')
plt.show()
graphiques()
|
Bases_de_Python.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import codecs, json
# -
with open('/Users/calmaleh/Desktop/school/project_course/jeppesen/ac_poor_2.bsad') as json_file:
json_data = json.load(json_file)
# +
frames = []
for j in range(len(json_data['tables'])):
df = pd.DataFrame(np.array(json_data['tables'][j]['table'])[:,:],
columns = json_data['tables'][j]['header']['variables'][:])
df['state'] = json_data['tables'][j]['header']['flightphase']
if df['state'][0] == 'cruise':
frames.append(df)
df = pd.concat(frames,ignore_index=True)
df = df[['DISA','ALTITUDE','MASS','TAS','FUELFLOW']]
# -
## treat the poor and rich data the same; methods shouldn't change (TAS being equivalent to MACH)
## normalize data beforehand might be an important factor? due to the different units (MACH -> %, TAS -> knots)
|
.ipynb_checkpoints/poor_and_limits_parse_test-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MSDS 430 Milestone 1
# ## California Wildfires
# Source: https://www.kaggle.com/rtatman/188-million-us-wildfires/download#FPA_FOD_20170508.sqlite
#
# Dates: 1992 - 2015
# Step 2 (5 pts.): Read in your data file and use pandas to inspect the first 5 lines and the last 5 lines.
import pandas
import numpy as np
import matplotlib.pyplot as plt
import sqlite3
import seaborn as sns
import pandas_profiling
import stats
conn = sqlite3.connect("../data/FPA_FOD_20170508.sqlite")
cur = conn.cursor()
# ## Getting to understand the data
# While Kaggle's documentation goes into detail about what's inside the SQLite file, I like to see for myself what tables are inside the instance I have in my memory, just to confirm they match. They do.
cur.execute("SELECT name FROM sqlite_master WHERE type='table';")
output = cur.fetchall()
tables = [table[0] for table in output]
tables.sort() # sorting alphabetically for readability
i = 0
while i < len(tables) - 1:
if len(tables[i]) > 10 & len(tables[i]) <= 15:
tabs = "\t\t\t\t"
elif len(tables[i]) > 15:
tabs = "\t\t\t\t\t"
# elif len(tables[i]) < 10:
# tabs = "\t\t"
# else:
# tabs = "\t\t\t\t"
print(tables[i] + tabs + tables[i + 1])
i += 2
#for table in tables:
# print(table)
# ## Getting column information
query = cur.execute("SELECT * FROM fires;") # this will point cur.description to the table fires
names = [description[0] for description in cur.description]
num_cols_fires = len(names) # this I will not sort because the order should coordinate with data
print(names, "\n\n--> column count sqlite file : ", num_cols_fires)
# ## SQLite Rowcount
# Before converting into pandas Dataframe, want to know the number of rows in SQLite file to assert conversion was successful.
cur.execute("SELECT COUNT(*) FROM fires;")
num_rows_fires = cur.fetchone()[0]
print("row count sqlite file : ", num_rows_fires)
sqlite_dim = (num_rows_fires, num_cols_fires) # storing sqlite dimension information for convenience
# ## Creating the dataframe from SQLite file
# According to Kaggle's documentation, FOD_ID is the global unique identifier. This will become the pandas index column.
df = pandas.read_sql("SELECT * FROM fires;", conn, index_col="FOD_ID")
# # First 5 elements
df.head(5) # first 5 elements
# # Last 5 elements
df.tail(5) # last 5 elements
# Step 3 (5 pts.): Use numpy to perform a few calculations relevant to your goals for the final project. Explain each calculation by including comments within your code.
# ## Relevant Calculations
# 1. Extracting California information from US-wide Dataset
# 2. Basic descriptive statistics
# 3. Fire frequency by year
# +
# Extracting CA info
CA = df[df['STATE']=='CA']
CA.head()
# +
# descriptive statistics
CA.describe()
# +
# missings
CA.isnull().sum()
# +
# missings as a proportion to population
proportions = 100 * (round(CA.isnull().sum() / len(CA), 4))
print(proportions)
# +
# dropping useless columns
CA.drop(columns = ["ICS_209_INCIDENT_NUMBER", "ICS_209_NAME",
"MTBS_ID", "MTBS_FIRE_NAME", "COMPLEX_NAME",
"COUNTY", "FIPS_CODE", "FIPS_NAME", "LOCAL_FIRE_REPORT_ID",
"OWNER_CODE", "FIRE_CODE"], inplace = True)
# -
"ICS_209_INCIDENT_NUMBER" in CA.columns # checking the drop worked correctly
# Step 4 (5 pts.): Use matplotlib to create a visual display relevant to your goals for the final project. Explain your graph by including comments within your code.
# ## Using matplotlib to visualize California wildfire frequencies over time
# +
# This gives a frequency distribution of all the years
years = CA["FIRE_YEAR"]
plt.hist(years, bins=len(CA["FIRE_YEAR"].unique()))
plt.title("Fire Year Frequencies")
plt.xlabel("Year")
plt.ylabel("Frequency")
plt.show()
# +
# curious about discovery date, how or if it differs
plt.hist(CA["DISCOVERY_DATE"], bins=len(CA["FIRE_YEAR"].unique()))
plt.title("Discovery Date Frequencies")
plt.xlabel("Year")
plt.ylabel("Frequency")
plt.show()
# +
# seeing whether the dates are the datetype
CA.dtypes
# +
# reviewing the missings after dropping useless columns
updated_proportions = 100 * (round(CA.isnull().sum() / len(CA), 4))
print(updated_proportions)
# +
# dropping missings for now; will find another strategy since this is removing about half the data
CA_without_missings = CA.dropna()
# +
# all proportions should now have 0% of their data containing missings
should_be_0 = 100 * (round(CA_without_missings.isnull().sum() / len(CA_without_missings), 4))
print(should_be_0)
# +
# filtering for only numeric datatypes in prep for Pearson coefficiant
numerics = ['int16', 'int32', 'int64', 'float16', 'float32', 'float64']
df = CA_without_missings.select_dtypes(include=numerics)
# +
# Seaborn for practical scatter plots including the Pearson coefficient.
mean = np.zeros(3)
cov = np.random.uniform(.2, .4, (3, 3))
cov += cov.T
cov[np.diag_indices(3)] = 1
data = np.random.multivariate_normal(mean, cov, 100)
def corrfunc(x, y, **kws):
r = stats.corr(x, y)
ax = plt.gca()
ax.annotate("r = {:.2f}".format(r),
xy=(.1, .9), xycoords=ax.transAxes)
g = sns.PairGrid(df, palette=["red"])
g.map_upper(plt.scatter, s=10)
g.map_diag(sns.distplot, kde=False)
g.map_lower(sns.kdeplot, cmap="Blues_d")
g.map_lower(corrfunc)
# -
CA_without_missings.columns
|
Fall2019/MSDS430/FinalProject/milestones/Draw_CA.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import datetime
anyday=datetime.datetime(2017,05,16
).strftime("%w")
anyday
from collections import defaultdict
dd = defaultdict(list)
dd['20']=3
dd['20']
a=['a','b']
dd=dict.fromkeys(a)
# +
dd
# -
a=['s','b','d','s']
set(a)
b=list(set(a))
b
b.sort()
b
anyday=datetime.datetime(2017,12,31
).strftime("%W")
anyday
#weekflag格式为'2017-20'(即2014年第35周)
def getfirstday(weekflag):
yearnum = weekflag[0:4] #取到年份
weeknum = weekflag[5:7] #取到周
stryearstart = yearnum +'0101' #当年第一天
yearstart = datetime.datetime.strptime(stryearstart,'%Y%m%d') #格式化为日期格式
yearstartcalendarmsg = yearstart.isocalendar() #当年第一天的周信息
yearstartweek = yearstartcalendarmsg[1]
yearstartweekday = yearstartcalendarmsg[2]
yearstartyear = yearstartcalendarmsg[0]
if yearstartyear < int (yearnum):
daydelat = (8-int(yearstartweekday))+(int(weeknum)-1)*7
else :
daydelat = (8-int(yearstartweekday))+(int(weeknum)-2)*7
first_day = (yearstart+datetime.timedelta(days=daydelat)).date()
last_day =first_day+datetime.timedelta(days=6)
return (first_day,last_day)
getfirstday('2017-20')
import re
# +
emailorphone="<EMAIL>"
p3=re.compile('^0\d{2,3}\d{7,8}$|^1[358]\d{9}$|^147\d{8}|[^\._-][\w\.-]+@(?:[A-Za-z0-9]+\.)+[A-Za-z]+)
emailorphonematch=p3.match(emailorphone)
if emailorphone:
print emailorphonematch.group()
else:
print "phone or email error..."
# -
p3=re.compile('^0\d{2,3}\d{7,8}$|^1[358]\d{9}$|^147\d{8}|[^\._-][\w\.-]+@(?:[A-Za-z0-9]+\.)+[A-Za-z]+’)
emailorphone="2014-34sdf2017-34"
p3=re.compile('^\d{4}-\d{2}').match(emailorphone)
p3
match.group()
a=[1200, 300, 200, 900, 300]
sum_list=sum(a)
def change(param,tempSum=0):
'''
累加器
'''
result=[]
for i in param:
tempSum=i+tempSum
result.append(tempSum)
return result
y_change=change(a)
y_change_data=[abs(i-sum_list) for i in y_change]
y_change
y_change_data
a = range(1,11)
b = range(1,10)
c = sum([item for item in a if item in b])
print c
|
Untitled.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] tags=[]
# # Example: CanvasXpress line Chart No. 3
#
# This example page demonstrates how to, using the Python package, create a chart that matches the CanvasXpress online example located at:
#
# https://www.canvasxpress.org/examples/line-3.html
#
# This example is generated using the reproducible JSON obtained from the above page and the `canvasxpress.util.generator.generate_canvasxpress_code_from_json_file()` function.
#
# Everything required for the chart to render is included in the code below. Simply run the code block.
# +
from canvasxpress.canvas import CanvasXpress
from canvasxpress.js.collection import CXEvents
from canvasxpress.render.jupyter import CXNoteBook
cx = CanvasXpress(
render_to="line3",
data={
"z": {
"Annt1": [
"Desc:1",
"Desc:2",
"Desc:3",
"Desc:4"
],
"Annt2": [
"Desc:A",
"Desc:B",
"Desc:A",
"Desc:B"
],
"Annt3": [
"Desc:X",
"Desc:X",
"Desc:Y",
"Desc:Y"
],
"Annt4": [
5,
10,
15,
20
],
"Annt5": [
8,
16,
24,
32
],
"Annt6": [
10,
20,
30,
40
]
},
"x": {
"Factor1": [
"Lev:1",
"Lev:2",
"Lev:3",
"Lev:1",
"Lev:2",
"Lev:3"
],
"Factor2": [
"Lev:A",
"Lev:B",
"Lev:A",
"Lev:B",
"Lev:A",
"Lev:B"
],
"Factor3": [
"Lev:X",
"Lev:X",
"Lev:Y",
"Lev:Y",
"Lev:Z",
"Lev:Z"
],
"Factor4": [
5,
10,
15,
20,
25,
30
],
"Factor5": [
8,
16,
24,
32,
40,
48
],
"Factor6": [
10,
20,
30,
40,
50,
60
]
},
"y": {
"vars": [
"V1",
"V2",
"V3",
"V4"
],
"smps": [
"S1",
"S2",
"S3",
"S4",
"S5",
"S6"
],
"data": [
[
5,
10,
25,
40,
45,
50
],
[
95,
80,
75,
70,
55,
40
],
[
25,
30,
45,
60,
65,
70
],
[
55,
40,
35,
30,
15,
1
]
]
}
},
config={
"graphOrientation": "vertical",
"graphType": "Line",
"legendPosition": "right",
"lineDecoration": "pattern",
"smpLabelRotate": 90,
"smpTitle": "Collection of Samples",
"smpTitleFontStyle": "italic",
"theme": "blackAndWhite",
"title": "Random Data"
},
width=613,
height=613,
events=CXEvents(),
after_render=[],
other_init_params={
"version": 35,
"events": False,
"info": False,
"afterRenderInit": False,
"noValidate": True
}
)
display = CXNoteBook(cx)
display.render(output_file="line_3.html")
|
tutorials/notebook/cx_site_chart_examples/line_3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/ryanleeallred/DS-Unit-1-Sprint-3-Statistical-Tests-and-Experiments/blob/master/module2-sampling-confidence-intervals-and-hypothesis-testing/LS_DS_132_Sampling_Confidence_Intervals_and_Hypothesis_Testing_Assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="11OzdxWTM7UR"
# ## Assignment - Build a confidence interval
#
# A confidence interval refers to a neighborhood around some point estimate, the size of which is determined by the desired p-value. For instance, we might say that 52% of Americans prefer tacos to burritos, with a 95% confidence interval of +/- 5%.
#
# 52% (0.52) is the point estimate, and +/- 5% (the interval $[0.47, 0.57]$) is the confidence interval. "95% confidence" means a p-value $\leq 1 - 0.95 = 0.05$.
#
# In this case, the confidence interval includes $0.5$ - which is the natural null hypothesis (that half of Americans prefer tacos and half burritos, thus there is no clear favorite). So in this case, we could use the confidence interval to report that we've failed to reject the null hypothesis.
#
# But providing the full analysis with a confidence interval, including a graphical representation of it, can be a helpful and powerful way to tell your story. Done well, it is also more intuitive to a layperson than simply saying "fail to reject the null hypothesis" - it shows that in fact the data does *not* give a single clear result (the point estimate) but a whole range of possibilities.
#
# How is a confidence interval built, and how should it be interpreted? It does *not* mean that 95% of the data lies in that interval - instead, the frequentist interpretation is "if we were to repeat this experiment 100 times, we would expect the average result to lie in this interval ~95 times."
#
# For a 95% confidence interval and a normal(-ish) distribution, you can simply remember that +/-2 standard deviations contains 95% of the probability mass, and so the 95% confidence interval based on a given sample is centered at the mean (point estimate) and has a range of +/- 2 (or technically 1.96) standard deviations.
#
# Different distributions/assumptions (90% confidence, 99% confidence) will require different math, but the overall process and interpretation (with a frequentist approach) will be the same.
#
# Your assignment - using the data from the prior module ([congressional voting records](https://archive.ics.uci.edu/ml/datasets/Congressional+Voting+Records)):
#
#
# ### Confidence Intervals:
# 1. Generate and numerically represent a confidence interval
# 2. Graphically (with a plot) represent the confidence interval
# 3. Interpret the confidence interval - what does it tell you about the data and its distribution?
#
# ### Chi-squared tests:
# 4. Take a dataset that we have used in the past in class that has **categorical** variables. Pick two of those categorical variables and run a chi-squared tests on that data
# - By hand using Numpy
# - In a single line using Scipy
#
# + colab={} colab_type="code" id="Ckcr4A4FM7cs"
# TODO - your code!
# -
# Start by importing the necessary libraries
import pandas as pd
import scipy.stats as sps
import numpy as np
# +
df= pd.read_csv(r'C:\Users\Administrator\Downloads\house-votes-84.data', header=None)
# Fix column names to match the issue voted on
df = df.rename(columns={0:'party', 1:'handicapped-infants', 2:'water-project',
3:'budget', 4:'phys-fee-freeze', 5:'elsalvador-aid', 6:'religious-groups-in-schools',
7:'anti-satellite', 8:'nicaraguan-aid', 9:'mx-missile', 10:'immigration',
11:'synfuels', 12:'education', 13:'superfund', 14:'crime', 15:'duty-free-exp',
16:'export-adm-sa'})
# -
df.head()
# Change votes to numeric format with 1 representing 'y', and 0 represeting 'n'. NaN fills for '?'
df= df.replace({'y':1, 'n':0, '?':np.NaN})
dem = df[df['party']== 'democrat']
rep = df[df['party']== 'republican']
# +
def statsGet(self, sample=True, confidence=0.95, get='none', h0='none'):
'''This section of the function will place the passed data into a numpy array with the variable name data.
Secondly defines N, mean, and variance.'''
data= np.array(self)
N= len(data)
mean= sum(data)/N
'''Define the function for finding variance'''
def variance(data, sample=True):
if sample!= True:
diff= [x- mean for x in data]
variance= sum([i**2 for i in diff])/N
return variance
else:
diff= [x- mean for x in data]
variance= sum([i**2 for i in diff])/(N-1)
return variance
'''Define the function for finding the sample deviation'''
def deviation(data, sample=True):
if sample!= True:
return variance(data, sample=False)**.5
else:
return variance(data)**.5
'''Define the function for finding the standard error'''
def stderr(data, sample=True):
if sample!= True:
return deviation(data, sample=False)/(N**.5)
else:
return deviation(data)/(N**.5)
'''Define Interval'''
def interval(data, sample=True):
if sample!= True:
return stderr(data, sample=False)* sps.t.ppf((1 + confidence) / 2, N)
else:
return stderr(data)* sps.t.ppf((1 + confidence) / 2, N - 1)
def format2(value):
return '{:.2f}'.format(value)
if sample!= True:
'''Values for statistical analysis for population data.'''
if get == 'none':
raise ValueError('No analysis requested')
if get == 'ci':
return [mean-interval(data, sample=False), mean+interval(data, sample=False)]
if get == 'ttest':
if h0 == 'none':
raise ValueError('Null Hypothesis not indicated.')
else:
return sps.ttest_1samp(data, h0, nan_policy='omit')
if get == 'dev':
return deviation(data, sample=False)
'''Values for statistical analysis for sample data.'''
else:
if get == 'none':
raise ValueError('No analysis requested')
if get == 'ci':
return [mean-interval(data), mean+interval(data)]
if get == 'ttest':
if h0 == 'none':
raise ValueError('Null Hypothesis not indicated.')
else:
return sps.ttest_1samp(data, h0, nan_policy='omit')
if get == 'dev':
return deviation(data)
# -
statsGet(dem['budget'].dropna(), get='ci')
# +
'''This is a most extremely simple graph that I made in ~15 seconds.'''
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
dem_budget= dem['budget'].dropna()
dem_water= dem['water-project'].dropna()
fig = plt.figure(figsize=(10, 8))
plt.subplots()
dem_budget_mean= dem['budget'].dropna().mean()
plt.bar(1, dem_budget_mean)
plt.vlines(1, statsGet(dem_budget, get='ci')[0], statsGet(dem_budget, get='ci')[1])
dem_waterp_mean= dem_water.mean()
plt.bar(2, dem_waterp_mean)
plt.vlines(2, statsGet(dem_water, get='ci')[0], statsGet(dem_water, get='ci')[1])
# + [markdown] colab_type="text" id="4ohsJhQUmEuS"
# ## Stretch goals:
#
# 1. Write a summary of your findings, mixing prose and math/code/results. *Note* - yes, this is by definition a political topic. It is challenging but important to keep your writing voice *neutral* and stick to the facts of the data. Data science often involves considering controversial issues, so it's important to be sensitive about them (especially if you want to publish).
# 2. Apply the techniques you learned today to your project data or other data of your choice, and write/discuss your findings here.
# 3. Refactor your code so it is elegant, readable, and can be easily run for all issues.
# + [markdown] colab_type="text" id="nyJ3ySr7R2k9"
# ## Resources
#
# - [Interactive visualize the Chi-Squared test](https://homepage.divms.uiowa.edu/~mbognar/applets/chisq.html)
# - [Calculation of Chi-Squared test statistic](https://en.wikipedia.org/wiki/Pearson%27s_chi-squared_test)
# - [Visualization of a confidence interval generated by R code](https://commons.wikimedia.org/wiki/File:Confidence-interval.svg)
# - [Expected value of a squared standard normal](https://math.stackexchange.com/questions/264061/expected-value-calculation-for-squared-normal-distribution) (it's 1 - which is why the expected value of a Chi-Squared with $n$ degrees of freedom is $n$, as it's the sum of $n$ squared standard normals)
|
module2-sampling-confidence-intervals-and-hypothesis-testing/LS_DS_132_Sampling_Confidence_Intervals_and_Hypothesis_Testing_Assignment.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # References
# * [Brief introduction to Python and Jupyter for Machine Learning (ML)](http://github.com/machine-learning-helpers/induction-python), with:
# + [this Altair section on that introduction](http://github.com/machine-learning-helpers/induction-python/tree/master/tutorials/altair)
# * [Altair - Basic Statistical Visualization](https://altair-viz.github.io/getting_started/starting.html#starting)
# # [The Data](https://altair-viz.github.io/getting_started/starting.html#the-data)
import pandas as pd
data = pd.DataFrame({'a': list('CCCDDDEEE'),
'b': [2, 7, 4, 1, 2, 6, 8, 4, 7]})
# # [The Chart Object](https://altair-viz.github.io/getting_started/starting.html#the-chart-object)
import altair as alt
chart = alt.Chart(data)
# # [Encodings and Marks](https://altair-viz.github.io/getting_started/starting.html#encodings-and-marks)
alt.Chart(data).mark_point()
alt.Chart(data).mark_point().encode(
x='a',
)
alt.Chart(data).mark_point().encode(
x='a',
y='b'
)
# # [Data Transformation: Aggregation](https://altair-viz.github.io/getting_started/starting.html#data-transformation-aggregation)
alt.Chart(data).mark_point().encode(
x='a',
y='average(b)'
)
alt.Chart(data).mark_bar().encode(
x='a',
y='average(b)'
)
alt.Chart(data).mark_bar().encode(
y='a',
x='average(b)'
)
# # [Aside: Examining the JSON Output](https://altair-viz.github.io/getting_started/starting.html#aside-examining-the-json-output)
chart = alt.Chart(data).mark_bar().encode(
x='a',
y='average(b)',
)
print(chart.to_json())
y = alt.Y('average(b):Q')
print(y.to_json())
y = alt.Y(field='b', type='quantitative', aggregate='average')
print(y.to_json())
alt.Chart(data).mark_bar().encode(
alt.Y('a', type='nominal'),
alt.X('b', type='quantitative', aggregate='average')
)
# # [Customizing your Visualization](https://altair-viz.github.io/getting_started/starting.html#customizing-your-visualization)
alt.Chart(data).mark_bar(color='firebrick').encode(
alt.Y('a', axis=alt.Axis(title='category')),
alt.X('average(b)', axis=alt.Axis(title='avg(b) by category'))
)
# # [Publishing your Visualization](https://altair-viz.github.io/getting_started/starting.html#publishing-your-visualization)
chart = alt.Chart(data).mark_bar().encode(
x='a',
y='average(b)',
)
chart.save('chart.html')
# + language="sh"
#
# cat chart.html
# -
|
tutorials/altair/altair-viz-jupyter-02.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="cT5cdSLPX0ui"
# # Intro to Object Detection Colab
#
# Welcome to the object detection colab! This demo will take you through the steps of running an "out-of-the-box" detection model in SavedModel format on a collection of images.
#
#
# + [markdown] colab_type="text" id="vPs64QA1Zdov"
# Imports
# + colab={} colab_type="code" id="OBzb04bdNGM8"
# !pip install -U --pre tensorflow=="2.2.0"
# + colab={} colab_type="code" id="NgSXyvKSNHIl"
import os
import pathlib
# Clone the tensorflow models repository if it doesn't already exist
if "models" in pathlib.Path.cwd().parts:
while "models" in pathlib.Path.cwd().parts:
os.chdir('..')
elif not pathlib.Path('models').exists():
# !git clone --depth 1 https://github.com/tensorflow/models
# + colab={} colab_type="code" id="rhpPgW7TNLs6"
# Install the Object Detection API
# %%bash
# cd models/research/
protoc object_detection/protos/*.proto --python_out=.
# cp object_detection/packages/tf2/setup.py .
python -m pip install .
# + colab={} colab_type="code" id="yn5_uV1HLvaz"
import io
import os
import scipy.misc
import numpy as np
import six
import time
from six import BytesIO
import matplotlib
import matplotlib.pyplot as plt
from PIL import Image, ImageDraw, ImageFont
import tensorflow as tf
from object_detection.utils import visualization_utils as viz_utils
# %matplotlib inline
# + colab={} colab_type="code" id="-y9R0Xllefec"
def load_image_into_numpy_array(path):
"""Load an image from file into a numpy array.
Puts image into numpy array to feed into tensorflow graph.
Note that by convention we put it into a numpy array with shape
(height, width, channels), where channels=3 for RGB.
Args:
path: a file path (this can be local or on colossus)
Returns:
uint8 numpy array with shape (img_height, img_width, 3)
"""
img_data = tf.io.gfile.GFile(path, 'rb').read()
image = Image.open(BytesIO(img_data))
(im_width, im_height) = image.size
return np.array(image.getdata()).reshape(
(im_height, im_width, 3)).astype(np.uint8)
# Load the COCO Label Map
category_index = {
1: {'id': 1, 'name': 'Faizan'},
2: {'id': 2, 'name': 'Ayan'},
3: {'id': 3, 'name': 'Rehan'},
4: {'id': 4, 'name': 'Seema'},
5: {'id': 5, 'name': 'Suffyan'}
}
# + colab={} colab_type="code" id="QwcBC2TlPSwg"
# Download the saved model and put it into models/research/object_detection/test_data/
# !wget http://download.tensorflow.org/models/object_detection/tf2/20200711/efficientdet_d0_coco17_tpu-32.tar.gz
# !tar -xf efficientdet_d0_coco17_tpu-32.tar.gz
# !mv efficientdet_d0_coco17_tpu-32/ models/research/object_detection/test_data/
# + colab={} colab_type="code" id="Z2p-PmKLYCVU"
start_time = time.time()
tf.keras.backend.clear_session()
detect_fn = tf.saved_model.load('models/research/object_detection/inference_graph/saved_model')
end_time = time.time()
elapsed_time = end_time - start_time
print('Elapsed time: ' + str(elapsed_time) + 's')
# + colab={} colab_type="code" id="vukkhd5-9NSL"
import time
image_dir = 'path/to/image/dir'
image = '*.jpg'
elapsed = []
for i in range(2):
image_path = os.path.join(image_dir, image)
image_np = load_image_into_numpy_array(image_path)
input_tensor = np.expand_dims(image_np, 0)
start_time = time.time()
detections = detect_fn(input_tensor)
end_time = time.time()
elapsed.append(end_time - start_time)
plt.rcParams['figure.figsize'] = [42, 21]
label_id_offset = 1
image_np_with_detections = image_np.copy()
viz_utils.visualize_boxes_and_labels_on_image_array(
image_np_with_detections,
detections['detection_boxes'][0].numpy(),
detections['detection_classes'][0].numpy().astype(np.int32),
detections['detection_scores'][0].numpy(),
category_index,
use_normalized_coordinates=True,
max_boxes_to_draw=200,
min_score_thresh=.40,
agnostic_mode=False)
plt.subplot(2, 1, i+1)
plt.imshow(image_np_with_detections)
mean_elapsed = sum(elapsed) / float(len(elapsed))
print('Elapsed time: ' + str(mean_elapsed) + ' second per image')
# +
import io
import os
import scipy.misc
import numpy as np
import six
import time
import glob
from IPython.display import display
from six import BytesIO
import matplotlib
import matplotlib.pyplot as plt
from PIL import Image, ImageDraw, ImageFont
import tensorflow as tf
from object_detection.utils import ops as utils_ops
from object_detection.utils import label_map_util
from object_detection.utils import visualization_utils as vis_util
# %matplotlib inline
# -
def load_image_into_numpy_array(path):
"""Load an image from file into a numpy array.
Puts image into numpy array to feed into tensorflow graph.
Note that by convention we put it into a numpy array with shape
(height, width, channels), where channels=3 for RGB.
Args:
path: a file path (this can be local or on colossus)
Returns:
uint8 numpy array with shape (img_height, img_width, 3)
"""
img_data = tf.io.gfile.GFile(path, 'rb').read()
image = Image.open(BytesIO(img_data))
(im_width, im_height) = image.size
return np.array(image.getdata()).reshape(
(im_height, im_width, 3)).astype(np.uint8)
labelmap_path = 'models/research/object_detection/training/label_map.pbtxt'
category_index = label_map_util.create_category_index_from_labelmap(labelmap_path, use_display_name=True)
category_index
# + jupyter={"outputs_hidden": true}
tf.keras.backend.clear_session()
model = tf.saved_model.load('models/research/object_detection/inference_graph/saved_model')
# -
def run_inference_for_single_image(model, image):
image = np.asarray(image)
# The input needs to be a tensor, convert it using `tf.convert_to_tensor`.
input_tensor = tf.convert_to_tensor(image)
# The model expects a batch of images, so add an axis with `tf.newaxis`.
input_tensor = input_tensor[tf.newaxis,...]
# Run inference
model_fn = model.signatures['serving_default']
output_dict = model_fn(input_tensor)
# All outputs are batches tensors.
# Convert to numpy arrays, and take index [0] to remove the batch dimension.
# We're only interested in the first num_detections.
num_detections = int(output_dict.pop('num_detections'))
output_dict = {key:value[0, :num_detections].numpy()
for key,value in output_dict.items()}
output_dict['num_detections'] = num_detections
# detection_classes should be ints.
output_dict['detection_classes'] = output_dict['detection_classes'].astype(np.int64)
# Handle models with masks:
if 'detection_masks' in output_dict:
# Reframe the the bbox mask to the image size.
detection_masks_reframed = utils_ops.reframe_box_masks_to_image_masks(
output_dict['detection_masks'], output_dict['detection_boxes'],
image.shape[0], image.shape[1])
detection_masks_reframed = tf.cast(detection_masks_reframed > 0.5,
tf.uint8)
output_dict['detection_masks_reframed'] = detection_masks_reframed.numpy()
return output_dict
for image_path in glob.glob('images/test/*.jpg'):
image_np = load_image_into_numpy_array(image_path)
output_dict = run_inference_for_single_image(model, image_np)
vis_util.visualize_boxes_and_labels_on_image_array(
image_np,
output_dict['detection_boxes'],
output_dict['detection_classes'],
output_dict['detection_scores'],
category_index,
instance_masks=output_dict.get('detection_masks_reframed', None),
use_normalized_coordinates=True,
line_thickness=8)
Image._show(Image.fromarray(image_np))
|
research/object_detection/colab_tutorials/inference_from_saved_model_tf2_colab.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Hierarchical clustering demo
#
# ## 1. Algorithm implementation
# It should be a tree, so we create a Node class first.
class BiClusterNode:
def __init__(self,vec,left=None,right=None,distance=0.0,id=None):
self.left=left
self.right=right
self.vec=vec
self.id=id
self.distance=distance
# We still need all different distance metrics
# +
from math import sqrt
def manhattan(v1,v2):
res=0
dimensions=min(len(v1),len(v2))
for i in range(dimensions):
res+=abs(v1[i]-v2[i])
return res
def euclidean(v1,v2):
res=0
dimensions=min(len(v1),len(v2))
for i in range(dimensions):
res+=pow(abs(v1[i]-v2[i]),2)
return sqrt(float(res))
def cosine(v1,v2):
dotproduct=0
dimensions=min(len(v1),len(v2))
for i in range(dimensions):
dotproduct+=v1[i]*v2[i]
v1len=0
v2len=0
for i in range (dimensions):
v1len+=v1[i]*v1[i]
v2len+=v2[i]*v2[i]
v1len=sqrt(v1len)
v2len=sqrt(v2len)
# we need distance here -
# we convert cosine similarity into distance
return 1.0-(float(dotproduct)/(v1len*v2len))
def pearson(v1,v2):
# Simple sums
sum1=sum(v1)
sum2=sum(v2)
# Sums of the squares
sum1Sq=sum([pow(v,2) for v in v1])
sum2Sq=sum([pow(v,2) for v in v2])
# Sum of the products
pSum=sum([v1[i]*v2[i] for i in range(min(len(v1),len(v2)))])
# Calculate r (Pearson score)
numerator=pSum-(sum1*sum2/len(v1))
denominator=sqrt((sum1Sq-pow(sum1,2)/len(v1))*(sum2Sq-pow(sum2,2)/len(v1)))
if denominator==0: return 1.0
# we need distance here -
# we convert pearson correlation into distance
return 1.0-numerator/denominator
def tanimoto(v1,v2):
c1,c2,shared=0,0,0
for i in range(len(v1)):
if v1[i]!=0 or v2[i]!= 0:
if v1[i]!=0: c1+=1 # in v1
if v2[i]!=0: c2+=1 # in v2
if v1[i]!=0 and v2[i]!=0: shared+=1 # in both
# we need distance here -
# we convert tanimoto overlap into distance
return 1.0-(float(shared)/(c1+c2-shared))
# -
# Hierarchical clustering algorithm.
def hcluster(rows,distance=euclidean):
distances={}
currentclustid=-1
# Clusters are initially just the rows
clust=[BiClusterNode(rows[i],id=i) for i in range(len(rows))]
while len(clust)>1:
lowestpair=(0,1)
closest=distance(clust[0].vec,clust[1].vec)
# loop through every pair looking for the smallest distance
for i in range(len(clust)):
for j in range(i+1,len(clust)):
# distances is the cache of distance calculations
if (clust[i].id,clust[j].id) not in distances:
distances[(clust[i].id,clust[j].id)]=distance(clust[i].vec,clust[j].vec)
d=distances[(clust[i].id,clust[j].id)]
if d<closest:
closest=d
lowestpair=(i,j)
# calculate the average of the two clusters
mergevec=[
(clust[lowestpair[0]].vec[i]+clust[lowestpair[1]].vec[i])/2.0
for i in range(len(clust[0].vec))]
# create the new cluster
newcluster=BiClusterNode(mergevec,left=clust[lowestpair[0]],
right=clust[lowestpair[1]],
distance=closest,id=currentclustid)
# cluster ids that weren't in the original set are negative
currentclustid-=1
del clust[lowestpair[1]]
del clust[lowestpair[0]]
clust.append(newcluster)
return clust[0]
# Function for printing resulting dendrogram.
def printhclust(clust,labels=None,n=0):
# indent to make a hierarchy layout
for i in range(n):
print (' ', end="")
if clust.id<0:
# negative id means that this is branch
print ('-')
else:
# positive id means that this is an endpoint
if labels==None: print (clust.id)
else: print (labels[clust.id])
# now print the right and left branches
if clust.left!=None: printhclust(clust.left,labels=labels,n=n+1)
if clust.right!=None: printhclust(clust.right,labels=labels,n=n+1)
# And to draw the dendrogram.
# +
# draw hierarchical clusters
from PIL import Image,ImageDraw
def getheight(clust):
# Is this an endpoint? Then the height is just 1
if clust.left==None and clust.right==None: return 1
# Otherwise the height is the same of the heights of
# each branch
return getheight(clust.left)+getheight(clust.right)
def getdepth(clust):
# The distance of an endpoint is 0.0
if clust.left==None and clust.right==None: return 0
# The distance of a branch is the greater of its two sides
# plus its own distance
return max(getdepth(clust.left),getdepth(clust.right))+clust.distance
def drawdendrogram(clust,labels,jpeg='clusters.jpg'):
# height and width
h=getheight(clust)*20
w=1200
depth=getdepth(clust)
# width is fixed, so scale distances accordingly
scaling=float(w-150)/depth
# Create a new image with a white background
img=Image.new('RGB',(w,h),(255,255,255))
draw=ImageDraw.Draw(img)
draw.line((0,h/2,10,h/2),fill=(255,0,0))
# Draw the first node
drawnode(draw,clust,10,(h/2),scaling,labels)
img.save(jpeg,'JPEG')
def drawnode(draw,clust,x,y,scaling,labels):
if clust.id<0:
h1=getheight(clust.left)*20
h2=getheight(clust.right)*20
top=y-(h1+h2)/2
bottom=y+(h1+h2)/2
# Line length
ll=clust.distance*scaling
# Vertical line from this cluster to children
draw.line((x,top+h1/2,x,bottom-h2/2),fill=(255,0,0))
# Horizontal line to left item
draw.line((x,top+h1/2,x+ll,top+h1/2),fill=(255,0,0))
# Horizontal line to right item
draw.line((x,bottom-h2/2,x+ll,bottom-h2/2),fill=(255,0,0))
# Call the function to draw the left and right nodes
drawnode(draw,clust.left,x+ll,top+h1/2,scaling,labels)
drawnode(draw,clust.right,x+ll,bottom-h2/2,scaling,labels)
else:
# If this is an endpoint, draw the item label
draw.text((x+5,y-7),labels[clust.id],(0,0,0))
# -
# ## 2. Same toy dataset: clustering papers by title
# The input is a vector of words for each document, created from the paper titles in file [titles.txt](titles.txt), and stored in file [titles_vectors.txt](titles_vectors.txt).
# The same function to read vectors file
def read_vector_file(file_name):
f = open(file_name)
lines=[line for line in f]
# First line is the column headers
colnames=lines[0].strip().split('\t')[:]
print(colnames)
rownames=[]
data=[]
for line in lines[1:]:
p=line.strip().split('\t')
# First column in each row is the rowname
if len(p)>1:
rownames.append(p[0])
# The data for this row is the remainder of the row
data.append([float(x) for x in p[1:]])
return rownames,colnames,data
# The same function to rotate the matrix.
def rotatematrix(data):
newdata=[]
for i in range(len(data[0])):
newrow=[data[j][i] for j in range(len(data))]
newdata.append(newrow)
return newdata
# Build hierarchy of clusters and print it.
# +
file_name = "titles_vectors.txt"
docs,words,data=read_vector_file(file_name)
clust=hcluster(data,distance=euclidean)
print ('clusters of documents')
printhclust(clust,labels=docs)
# -
drawdendrogram(clust,docs,jpeg='docsclust_image.jpg')
# +
from PIL import Image # to load images
from IPython.display import display # to display images
pil_im = Image.open('docsclust_image.jpg')
display(pil_im)
# -
# And these are our original documents:
# - `['d1', {'human': 1, 'interface': 1, 'computer': 1}]`
# - `['d2', {'survey': 1, 'user': 1, 'computer': 1, 'system': 1, 'response': 1, 'time': 1}]`
# - `['d3', {'eps': 1, 'user': 1, 'interface': 1, 'system': 1}]`
# - `['d4', {'system': 2, 'human': 1, 'eps': 1}]`
# - `['d5', {'user': 1, 'response': 1, 'time': 1}]`
# - `['d6', {'trees': 1}]`
# - `['d7', {'graph': 1, 'trees': 1}]`
# - `['d8', {'graph': 1, 'minors': 1, 'trees': 1}]`
# - `['d9', {'graph': 1, 'minors': 1, 'survey': 1}]`
# ## 3. Clustering words by their co-occurrence in documents
# If we want to cluster words by their occurrences in the documents, all we need to do is to transpose the matrix.
rdata=rotatematrix(data)
clust=hcluster(rdata,distance=euclidean)
print ('hierarchical clusters of words')
printhclust(clust,labels=words)
drawdendrogram(clust,words,jpeg='wordsclust_image.jpg')
pil_im = Image.open('wordsclust_image.jpg')
display(pil_im)
# Copyright © 2020 <NAME>. All rights reserved.
|
hierarchical_clustering_demo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: "\u201Ctorch1.1\u201D"
# language: python
# name: python3
# ---
import random
import os
import sys
import torch.backends.cudnn as cudnn
import torch.optim as optim
import torch.utils.data
import numpy as np
from seriesDataset import seriesDataset
from torchvision import datasets
from torchvision import transforms
# +
# source_root = '/home/iot/jupyter/root_dir/zkx2/letterSim'
source_root = '/home/iot/jupyter/root_dir/zkx2/shapeletX'
labelMap = {"a":0, "b":1, "c":2, "d":3, "e":4}
source_dataset = seriesDataset( source_root, labelMap , False)
dataloader_source = torch.utils.data.DataLoader(
dataset=source_dataset,
batch_size=10,
shuffle=True,
num_workers=8
)
test_root = '/home/iot/jupyter/root_dir/zkx2/UserData/user1/letter'
test_dataset = seriesDataset(test_root , labelMap , False)
dataloader_test = torch.utils.data.DataLoader(
dataset= test_dataset,
batch_size=20,
shuffle=True,
num_workers=8
)
# +
import torch.nn as nn
class KaiNet(nn.Module):
def __init__(self):
super(KaiNet, self).__init__()
self.feature = nn.Sequential()
self.feature.add_module('f_conv1', nn.Conv1d(1,64,kernel_size=3))
self.feature.add_module('f_bn', nn.BatchNorm1d(64))
self.feature.add_module('f_relu', nn.ReLU(True))
self.feature.add_module('f_conv2', nn.Conv1d(64,128,kernel_size=3))
self.feature.add_module('f_bn2', nn.BatchNorm1d(128))
self.feature.add_module('f_relu2', nn.ReLU(True))
self.feature.add_module('f_conv3', nn.Conv1d(128,128,kernel_size=3))
self.feature.add_module('f_bn3', nn.BatchNorm1d(128))
self.feature.add_module('f_relu3', nn.ReLU(True))
self.class_classifier = nn.Sequential()
self.class_classifier.add_module('c_fc1', nn.Linear(250*128, 32))
self.class_classifier.add_module("c_bn1", nn.BatchNorm1d(32))
self.class_classifier.add_module('c_relu', nn.ReLU(True))
self.class_classifier.add_module('c_fc2', nn.Linear(32,5))
def forward(self, input_data ):
input_data = input_data.view(-1 , 1, 256)
x = self.feature(input_data.float())
x = x.view(-1 , 250*128)
class_output = self.class_classifier(x)
return class_output
# -
class MyTransformer(nn.Module):
def __init__(self):
super(MyTransformer , self).__init__()
self.encoder_layer = nn.TransformerEncoderLayer(256, nhead=8, dropout=0.1)
self.transformer_encoder = nn.TransformerEncoder(self.encoder_layer , num_layers=1)
self.classifier = nn.Sequential()
self.classifier.add_module( 'c_fc1' ,nn.Linear( 256 , 128))
self.classifier.add_module('c_relu' , nn.ReLU(True))
self.classifier.add_module( 'c_fc2' , nn.Linear(128 , 5))
def forward(self, input_data):
input_data = input_data.view(-1 , 1, 256)
x = input_data.float()
x = self.transformer_encoder(x)
x = x.view(-1 , 256 )
y = self.classifier(x)
return y
# +
# my_net = KaiNet()
my_net = MyTransformer()
cuda = True
lr = 1e-4
optimizer = optim.Adam(my_net.parameters(), lr=lr)
loss_class = torch.nn.CrossEntropyLoss()
if cuda:
my_net = my_net.cuda()
loss_class = loss_class.cuda()
# -
for epoch in range(50):
data_source_iter = iter(dataloader_source)
len_dataloader = len(dataloader_source)
my_net.train()
for i in range(len_dataloader):
s_img, s_label = data_source_iter.next()
my_net.zero_grad()
if cuda:
s_img = s_img.cuda()
s_label = s_label.cuda()
class_output = my_net(input_data = s_img)
s_label = s_label.view((s_label.shape[0]))
err_s_label = loss_class(class_output , s_label)
err_s_label.backward()
optimizer.step()
sys.stdout.write('\r epoch:%d step:%d err_s_label:%f'%(epoch, i , err_s_label))
sys.stdout.flush()
# test
my_net.eval()
test_iter = iter(dataloader_test)
len_test =len(dataloader_test)
n_total = 0
n_correct = 0
for j in range(len_test):
data_test = test_iter.next()
t_img, t_label = data_test
batch_size = len(t_label)
if cuda:
t_img = t_img.cuda()
t_label = t_label.cuda()
class_output = my_net(t_img)
pred = class_output.data.max(1 , keepdim=True)[1]
n_correct += pred.eq(t_label.data.view_as(pred)).cpu().sum()
n_total += batch_size
accu = n_correct.data.numpy() * 1.0 / n_total
print("accuracy:%f"%(accu))
test_root = '/home/iot/jupyter/root_dir/zkx2/UserData/user10/letter'
test_dataset = seriesDataset(test_root , labelMap , True)
dataloader_test = torch.utils.data.DataLoader(
dataset= test_dataset,
batch_size=20,
shuffle=True,
num_workers=8
)
test_iter = iter(dataloader_test)
len_test =len(dataloader_test)
n_total = 0
n_correct = 0
for j in range(len_test):
data_test = test_iter.next()
t_img, t_label = data_test
batch_size = len(t_label)
if cuda:
t_img = t_img.cuda()
t_label = t_label.cuda()
class_output = my_net(t_img)
pred = class_output.data.max(1 , keepdim=True)[1]
n_correct += pred.eq(t_label.data.view_as(pred)).cpu().sum()
n_total += batch_size
accu = n_correct.data.numpy() * 1.0 / n_total
print("accuracy:%f"%(accu))
|
DataProcess/net/DANN.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="rIh7FpkJ63RE" outputId="1bef3ebf-749d-42da-8621-7e7a428c2258"
# !unzip "/content/drive/MyDrive/Colab Notebooks/curso word2vec/cbow_s300.zip"
# !unzip "/content/drive/MyDrive/Colab Notebooks/curso word2vec/skip_s300.zip"
# + [markdown] id="cHxCW4e7SXTP"
# ## Libs Usadas
# + id="tRCwLVlkDuaa" colab={"base_uri": "https://localhost:8080/"} outputId="deb66911-4064-4ae7-e93a-e991759f7ec8"
import nltk
import string
import numpy as np
import pandas as pd
from gensim.models import KeyedVectors
from sklearn.dummy import DummyClassifier
from sklearn.metrics import classification_report
from sklearn.linear_model import LogisticRegression
from sklearn.feature_extraction.text import CountVectorizer
nltk.download('punkt')
# + [markdown] id="Nj5UrukxSZ3o"
# ## Leitura dos dados
# + id="tyIX5pogHAAI"
artigo_treino = pd.read_csv('/content/drive/MyDrive/Colab Notebooks/curso word2vec/treino.csv')
artigo_teste = pd.read_csv('/content/drive/MyDrive/Colab Notebooks/curso word2vec/teste.csv')
# + colab={"base_uri": "https://localhost:8080/", "height": 697} id="gZtIeGydHe8C" outputId="ee72ac8c-a678-477b-fd1f-9e740202752d"
display(artigo_treino.head())
display(artigo_treino.shape)
display(artigo_teste.head())
display(artigo_teste.shape)
# + [markdown] id="KfZZyN_ASJfi"
# ## Estudo de como funciona o Word2Vec
# + id="s2jIj4qULF3N"
modelo = KeyedVectors.load_word2vec_format('/content/cbow_s300.txt')
# + colab={"base_uri": "https://localhost:8080/"} id="za_Hxg36UTgH" outputId="3713a454-d986-40e5-cd8d-10c0a4d90f2b"
modelo.most_similar('carro')
# + colab={"base_uri": "https://localhost:8080/"} id="fsVBmk8NNM4N" outputId="f4fd26d2-3cd8-490c-d4df-bad315e3c3b6"
modelo.most_similar(positive=["brasil", "argentina", "uruguai"])
# + colab={"base_uri": "https://localhost:8080/"} id="o1COBdLhOrmN" outputId="b3e5ac83-2904-4c2d-edd6-346ef0e5db84"
'''
Possível maneira de descobrir o plural de palavras.
nuvens -> nuvem
estrelas -> estrelas
Como descobrir o plural da palavra "estrelas" com base nas outras informações?
nuvens + estrela - nuvem = estrelas
As operações vetoriais deveriam resultar assim.
'''
modelo.most_similar(positive=["nuvens", "estrela"], negative=["nuvem"])
# + colab={"base_uri": "https://localhost:8080/"} id="PnHEhb4oQCxI" outputId="14f5a3da-edee-4338-acd9-26f776c107dd"
'''
Nem tudo funciona perfeitamente.
Nesse caso abaixo podemos ver o claro vies humano.
Onde é associado a figura da mulher a enfermeira em vez de médica.
'''
modelo.most_similar(positive=["médico", "mulher"], negative=["homem"])
# + colab={"base_uri": "https://localhost:8080/"} id="blyTpZPIQoNC" outputId="dbadac02-d3e4-44fd-a7ba-6ac46da4a8dd"
'''
Para um caso em que não temos tanto um vies humano, a relação funciona
corretamente, pois nesse caso é mais comum associar a mulher o cargo
de professora.
Assim, é muito importante fazer um estudo dos dados que foram analisados.
'''
modelo.most_similar(positive=["professor", "mulher"], negative=["homem"])
# + [markdown] id="tP72VOu1SQ4a"
# ## Tokenização
# #### Vetorização de textos
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="csembmydRxKU" outputId="f00fb56a-72a4-4100-861b-7a8ae29dd9aa"
artigo_treino.title.loc[8]
# + id="lCIWCg1gmCjE"
'''
Função que tokeniza um texto separando pela pontuação.
Recebe uma frase como entrada.
'''
def tokenizador(texto):
lista_alfanumerica = list()
texto_min = texto.lower()
for token_valido in nltk.word_tokenize(texto_min):
if token_valido in string.punctuation: continue
lista_alfanumerica.append(token_valido)
return lista_alfanumerica
# + id="KzfxjOm3rajS"
'''
Função que cria e ajusta a frase de entrada para um vetor.
Recebe uma frase que já foi tokenizada como entrada.
'''
def combinacao_vetores_soma(palavras_numeros):
vetor_resultante = np.zeros(300)
for pn in palavras_numeros:
try:
vetor_resultante += modelo.get_vector(pn)
except KeyError:
if pn.isnumeric():
pn = '0'*len(pn)
vetor_resultante += modelo.get_vector(pn)
else:
vetor_resultante += modelo.get_vector('unknown')
return vetor_resultante
# + colab={"base_uri": "https://localhost:8080/"} id="bS_U_YiWsKoN" outputId="67a13f48-d62c-43bd-d64c-313f6f4e6cd5"
'''
Ainda existem erros nesse algoritmo.
Como o fato de palavras escritas erradas gerarem erro no Word2Vec.
Também temos o problema dos números, onde é preciso transformar
todos para zero.
Isso foi descrito no artigo científico do projeto do Word2Vec que
estamos usando.
As correções foram feitas na função acima com o try : except.
'''
palavras_numeros = tokenizador("texto alura")
vetor_texto = combinacao_vetores_soma(palavras_numeros)
print(len(vetor_texto))
print(vetor_texto)
# + id="_WTJdHVG0tyX"
'''
Função para transformar o conjunto de treino e teste em uma
matriz de vetores.
Recebe o conjunto de textos do DataFrame como entrada.
'''
def matriz_vetores(textos):
x = len(textos)
y = 300
matriz = np.zeros((x, y))
for i in range(x):
palavras_numeros = tokenizador(textos.iloc[i])
matriz[i] = combinacao_vetores_soma(palavras_numeros)
return matriz
# + id="fofmHobH20la"
matriz_vetores_treino = matriz_vetores(artigo_treino.title)
matriz_vetores_teste = matriz_vetores(artigo_teste.title)
# + [markdown] id="Kk-ra_5FW3oH"
# # Treinamento e Comparações
# + [markdown] id="R6WunipxXe7i"
# ## Treinamento com CBOW
# + colab={"base_uri": "https://localhost:8080/"} id="G-00ERmd3MRG" outputId="3feba821-e06e-4ca7-a29b-0b623e822157"
print(matriz_vetores_treino.shape)
print(matriz_vetores_teste.shape)
# + [markdown] id="CWmv5ZxeW8H-"
# ### Regressão Logística
# + id="TJA2mpzZD3YE" colab={"base_uri": "https://localhost:8080/"} outputId="b574f5ce-bf6a-4bb0-87ab-9800aa9cb008"
lr = LogisticRegression(max_iter=200)
lr.fit(matriz_vetores_treino, artigo_treino.category)
# + colab={"base_uri": "https://localhost:8080/"} id="uQ8ziEklSyId" outputId="033c507c-1306-451a-930f-3f74cdf1c4e2"
label_prevista = lr.predict(matriz_vetores_teste)
lr.score(matriz_vetores_teste, artigo_teste.category)
# + colab={"base_uri": "https://localhost:8080/"} id="H0ccNsPoUdlN" outputId="19750b6d-c2a6-4459-935d-1748fc099d47"
cr = classification_report(artigo_teste.category, label_prevista)
print(cr)
# + [markdown] id="F8sEnZHNW__c"
# ### Dummy Classifier
# + colab={"base_uri": "https://localhost:8080/"} id="vHWC0WTQWZTL" outputId="d0e01e58-997f-4110-cf9e-e1f3b4455aef"
dc = DummyClassifier()
dc.fit(matriz_vetores_treino, artigo_treino.category)
# + colab={"base_uri": "https://localhost:8080/"} id="P22gWmDaWg6s" outputId="9b8c1ed8-de53-441d-95bd-860ee1919bbe"
label_prevista_dc = dc.predict(matriz_vetores_teste)
dc.score(matriz_vetores_teste, artigo_teste.category)
# + colab={"base_uri": "https://localhost:8080/"} id="3mhDaDNbWrEV" outputId="0698d48a-f81a-4a7b-ceed-3e3061d40d8e"
cr_dc = classification_report(artigo_teste.category, label_prevista_dc)
print(cr_dc)
# + [markdown] id="0yMKoC3zXkqp"
# ## Treinamento com Skip-gram
# + id="K1qfzk2jXn3m"
modelo_skipgram = KeyedVectors.load_word2vec_format("skip_s300.txt")
# + id="yCuAojo94RIw"
def combinacao_vetores_soma_skipgram(palavras_numeros):
vetor_resultante = np.zeros(300)
for pn in palavras_numeros:
try:
vetor_resultante += modelo_skipgram.get_vector(pn)
except KeyError:
if pn.isnumeric():
pn = '0'*len(pn)
vetor_resultante += modelo_skipgram.get_vector(pn)
else:
vetor_resultante += modelo_skipgram.get_vector('unknown')
return vetor_resultante
def matriz_vetores_skipgram(textos):
x = len(textos)
y = 300
matriz = np.zeros((x, y))
for i in range(x):
palavras_numeros = tokenizador(textos.iloc[i])
matriz[i] = combinacao_vetores_soma_skipgram(palavras_numeros)
return matriz
# + id="ZMQpdhZC40ZC"
matriz_vetores_treino_skipgram = matriz_vetores_skipgram(artigo_treino.title)
matriz_vetores_teste_skipgram = matriz_vetores_skipgram(artigo_teste.title)
# + colab={"base_uri": "https://localhost:8080/"} id="0E4yVn7j49Vp" outputId="06c1b3ae-119a-4bcb-85c1-62d55e73c3bf"
lr_skipgram = LogisticRegression(max_iter=300)
lr_skipgram.fit(matriz_vetores_treino_skipgram, artigo_treino.category)
# + colab={"base_uri": "https://localhost:8080/"} id="KBhdYbX65Nag" outputId="e783c951-cc2e-4f98-f3a2-a16ad6550121"
label_prevista_skipgram = lr.predict(matriz_vetores_teste_skipgram)
lr_skipgram.score(matriz_vetores_teste_skipgram, artigo_teste.category)
# + colab={"base_uri": "https://localhost:8080/"} id="CfZKExKr52sB" outputId="e7d8797f-1962-4f33-8eee-4d67aecc8358"
cr_skipgram = classification_report(artigo_teste.category, label_prevista_skipgram)
print(cr_skipgram)
|
2 - Word Embedding/Word2Vec.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/NikuDubenco/DS-Unit-2-Regression-2/blob/master/Nicolae_Dubenco_DS_SC_8_Regression_2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="rPh-uurrzprt" colab_type="text"
# _Lambda School Data Science, Unit 2_
#
# # Regression 2 Sprint Challenge: Predict drugstore sales 🏥
#
# For your Sprint Challenge, you'll use real-world sales data from a German drugstore chain, from Jan 2, 2013 — July 31, 2015.
#
# You are given three dataframes:
#
# - `train`: historical sales data for 100 stores
# - `test`: historical sales data for 100 different stores
# - `store`: supplemental information about the stores
#
#
# The train and test set do _not_ have different date ranges. But they _do_ have different store ids. Your task is _not_ to forecast future sales from past sales. **Your task is to predict sales at unknown stores, from sales at known stores.**
# + id="jKOFk_6nzpru" colab_type="code" colab={}
import pandas as pd
train = pd.read_csv('https://drive.google.com/uc?export=download&id=1E9rgiGf1f_WL2S4-V6gD7ZhB8r8Yb_lE')
test = pd.read_csv('https://drive.google.com/uc?export=download&id=1vkaVptn4TTYC9-YPZvbvmfDNHVR8aUml')
store = pd.read_csv('https://drive.google.com/uc?export=download&id=1rZD-V1mWydeytptQfr-NL7dBqre6lZMo')
assert train.shape == (78400, 7)
assert test.shape == (78400, 7)
assert store.shape == (200, 10)
# + [markdown] id="PrvIj4olzprv" colab_type="text"
# The dataframes have a variety of columns:
#
# - **Store** - a unique Id for each store
# - **DayOfWeek** - integer, 1-6
# - **Date** - the date, from Jan 2, 2013 — July 31, 2015.
# - **Sales** - the units of inventory sold on a given date (this is the target you are predicting)
# - **Customers** - the number of customers on a given date
# - **Promo** - indicates whether a store is running a promo on that day
# - **SchoolHoliday** - indicates the closure of public schools
# - **StoreType** - differentiates between 4 different store models: a, b, c, d
# - **Assortment** - describes an assortment level: a = basic, b = extra, c = extended
# - **CompetitionDistance** - distance in meters to the nearest competitor store
# - **CompetitionOpenSince[Month/Year]** - gives the approximate year and month of the time the nearest competitor was opened
# - **Promo2** - Promo2 is a continuing and consecutive promotion for some stores: 0 = store is not participating, 1 = store is participating
# - **Promo2Since[Year/Week]** - describes the year and calendar week when the store started participating in Promo2
# - **PromoInterval** - describes the consecutive intervals Promo2 is started, naming the months the promotion is started anew. E.g. "Feb,May,Aug,Nov" means each round starts in February, May, August, November of any given year for that store
# + [markdown] id="Txb785Qdzprw" colab_type="text"
# This Sprint Challenge has three parts. To demonstrate mastery on each part, do all the required instructions. To earn a score of "3" for the part, also do the stretch goals.
# + id="Mb5TvhQwj8Bd" colab_type="code" outputId="e7146469-3944-48b4-a836-b99856b949e2" colab={"base_uri": "https://localhost:8080/", "height": 359}
train.head(10)
# + id="4PQlNNgylCFC" colab_type="code" outputId="5d1d3e00-2d78-4bf0-d5be-a0a42d486447" colab={"base_uri": "https://localhost:8080/", "height": 34}
train['Store'].unique().sum()
# + id="c6Tbvi_elO-p" colab_type="code" outputId="2adefbdd-e09e-436c-e792-770f3a682dd5" colab={"base_uri": "https://localhost:8080/", "height": 297}
train.describe()
# + id="dbYGoiA8mn3d" colab_type="code" outputId="3d573e8f-e7d9-43fd-f573-eb0ebc4c509d" colab={"base_uri": "https://localhost:8080/", "height": 221}
train.info()
# + id="DsPcGOZ0kE3E" colab_type="code" outputId="db9f82cb-559a-49ce-ce42-053d9e7d54ca" colab={"base_uri": "https://localhost:8080/", "height": 359}
test.head(10)
# + id="Yk-8nw9skxAZ" colab_type="code" outputId="b31da475-dbe8-4239-ea44-0e8a6d598906" colab={"base_uri": "https://localhost:8080/", "height": 34}
test['Store'].unique().sum()
# + id="CZhdLkb1lg7k" colab_type="code" outputId="1f74ba54-dad6-4d52-b862-28b3802066ec" colab={"base_uri": "https://localhost:8080/", "height": 297}
test.describe()
# + id="I6Ls791BmsJK" colab_type="code" outputId="d42b7fac-42d8-4895-860c-7ac1b95a0211" colab={"base_uri": "https://localhost:8080/", "height": 221}
test.info()
# + id="NM91GWV7kQys" colab_type="code" outputId="7a917906-5c0b-45ea-832d-986363345c22" colab={"base_uri": "https://localhost:8080/", "height": 379}
store.head(10)
# + id="lMezhyD1l27W" colab_type="code" outputId="bef35ebf-7d40-48ed-9ddb-a9a4897448ed" colab={"base_uri": "https://localhost:8080/", "height": 297}
store.describe()
# + id="x8WOzXOZmDJT" colab_type="code" outputId="ecc994cc-5c07-4942-a6fe-3dcfaca4183a" colab={"base_uri": "https://localhost:8080/", "height": 173}
store.describe(exclude='number')
# + id="sJWgZFYEmRkp" colab_type="code" outputId="a5fc34e1-edff-45b9-e863-ccc19e19ad7f" colab={"base_uri": "https://localhost:8080/", "height": 272}
store.info()
# + id="44XFb1YpmdPx" colab_type="code" outputId="80697468-194d-455a-92f4-d5d2e9322d65" colab={"base_uri": "https://localhost:8080/", "height": 204}
store.isna().sum()
# + [markdown] id="B9NV3COuzprw" colab_type="text"
# ## 1. Wrangle relational data, Log-transform the target
# - Merge the `store` dataframe with the `train` and `test` dataframes.
# - Arrange the X matrix and y vector for the train and test sets.
# - Log-transform the target for the train and test set.
# - Plot the target's distribution for the train set, before and after the transformation.
#
# #### Stretch goals
# - Engineer 3+ more features.
# + id="k8gdnxzPs2h2" colab_type="code" colab={}
# %matplotlib inline
import numpy as np
import seaborn as sns
# + id="qjvBbl78zprx" colab_type="code" outputId="43630225-f172-4d59-a9ee-084cea65b954" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Merge the store dataframe with the train and test dataframes
train = train.merge(store, left_on='Store', right_on='Store', how='left')
test = test.merge(store, left_on='Store', right_on='Store', how='left')
train.shape, test.shape
# + id="9T_UPczCrH2W" colab_type="code" colab={}
# Arrange the X matrix and y vector for the train and test sets
target = 'Sales'
X_train = train.drop(columns=target)
X_test = test.drop(columns=target)
y_train = train[target]
y_test = test[target]
# + id="6TD58Fcmts8l" colab_type="code" outputId="7305308b-5393-4778-c64b-de1bb01eb794" colab={"base_uri": "https://localhost:8080/", "height": 283}
# Plot the target's distribution for the train set, before the transformation
sns.distplot(y_train);
# + id="U4FQ3mdLrzpW" colab_type="code" colab={}
# Log-transform the target for the train and test set
y_train_log = np.log1p(y_train)
y_test_log = np.log1p(y_test)
# + id="ITuF4myAsE5u" colab_type="code" outputId="3a82c773-2b2b-49cc-fe80-b2fd5893045c" colab={"base_uri": "https://localhost:8080/", "height": 283}
# Plot the target's distribution for the train set, after the transformation
sns.distplot(y_train_log);
# + [markdown] id="xiljXNuKzprz" colab_type="text"
# ## 2. Fit and validate your model
# - **Use Gradient Boosting** or any type of regression model.
# - **Beat the baseline:** The estimated baseline Root Mean Squared Logarithmic Error is 0.90, if we guessed the mean sales for every prediction. Remember that RMSE with the log-transformed target is equivalent to RMSLE with the original target. Try to get your error below 0.20.
# - **To validate your model, choose any one of these options:**
# - Split the train dataframe into train and validation sets. Put all dates for a given store into the same set. Use xgboost `early_stopping_rounds` with the validation set.
# - Or, use scikit-learn `cross_val_score`. Put all dates for a given store into the same fold.
# - Or, use scikit-learn `RandomizedSearchCV` for hyperparameter optimization. Put all dates for a given store into the same fold.
# - **Get the Validation Error** (multiple times if you try multiple iterations) **and Test Error** (one time, at the end).
#
# #### Stretch goal
# - Optimize 3+ hyperparameters by searching 10+ "candidates" (possible combinations of hyperparameters).
# + id="uH60EmvJv54O" colab_type="code" outputId="34e39882-41d2-4f2c-c2a2-4e6fb4c3e3b3" colab={"base_uri": "https://localhost:8080/", "height": 292}
# !pip install category_encoders
# + id="9sBgGuPBvcbS" colab_type="code" outputId="77b7d377-2126-40b3-f2ca-fdc654b2cf39" colab={"base_uri": "https://localhost:8080/", "height": 51}
# %%time
import category_encoders as ce
from sklearn.ensemble import RandomForestRegressor
from xgboost import XGBRegressor
from sklearn.metrics import mean_squared_error
from sklearn.pipeline import make_pipeline
# + id="WWNccxI5zprz" colab_type="code" outputId="33c86bdc-bd8e-4ed6-dd3e-10e27c598ac0" colab={"base_uri": "https://localhost:8080/", "height": 85}
# Use Gradient Boosting in pipeline together with Ordinal Encoder
pipeline = make_pipeline(
ce.OrdinalEncoder(),
XGBRegressor(n_estimators=1000, n_jobs=-1))
pipeline.fit(X_train, y_train_log)
y_pred_log = pipeline.predict(X_test)
print('Validation Error =', np.sqrt(mean_squared_error(y_test_log, y_pred_log)))
# + id="-yrglG5ZyAFk" colab_type="code" outputId="2f199264-b87b-4367-b0e4-bd5685bf1641" colab={"base_uri": "https://localhost:8080/", "height": 289}
# Use scikit-learn cross_val_score to validate my model
from sklearn.model_selection import cross_val_score
scores = cross_val_score(pipeline, X_train, y_train_log, cv=5,
scoring='neg_mean_squared_error')
print('RMSLE for 5 folds:', np.sqrt(-scores))
# + id="1Dn9YeKFIwxm" colab_type="code" outputId="3eeab0d8-9810-4b4f-f4ce-1da2b8ede278" colab={"base_uri": "https://localhost:8080/", "height": 1836}
from scipy.stats import randint, uniform
from sklearn.model_selection import RandomizedSearchCV
param_distributions = { 'randomforestregressor__n_estimators': randint(50, 200),
'randomforestregressor__max_features': uniform(),
'randomforestregressor__min_samples_leaf': [1, 10, 50]}
search = RandomizedSearchCV(pipeline,
param_distributions=param_distributions,
n_iter=2,
cv=2,
scoring='neg_mean_squared_error',
verbose=10,
return_train_score=True,
n_jobs=-1)
search.fit(X_train, y_train_log)
print('Best hyperparameters', search.best_params_)
print('Cross-validation RMSLE', np.sqrt(-search.best_score_))
# + [markdown] id="JGqeEpRmzpr1" colab_type="text"
# ## 3. Plot model interpretation visualizations
# - Choose any one of these options:
# - Permutation Importances plot
# - Partial Dependency Plot, 1 feature isolation
# - Partial Dependency Plot, 2 feature interaction
#
# #### Stretch goals
# - Plot 2+ visualizations.
# - Use permutation importances for feature selection.
# + id="c7a0cB6A7j0e" colab_type="code" outputId="2dcb8360-5b3b-4f4d-83d0-1193f672c18d" colab={"base_uri": "https://localhost:8080/", "height": 581}
# !pip install eli5 pdpbox
# + id="Hw43UFQyDuxn" colab_type="code" colab={}
train = pd.read_csv('https://drive.google.com/uc?export=download&id=1E9rgiGf1f_WL2S4-V6gD7ZhB8r8Yb_lE')
test = pd.read_csv('https://drive.google.com/uc?export=download&id=1vkaVptn4TTYC9-YPZvbvmfDNHVR8aUml')
store = pd.read_csv('https://drive.google.com/uc?export=download&id=1rZD-V1mWydeytptQfr-NL7dBqre6lZMo')
assert train.shape == (78400, 7)
assert test.shape == (78400, 7)
assert store.shape == (200, 10)
# + id="8UfNrK5c8Och" colab_type="code" colab={}
# Engineer 3+ more features
def wrangle(X):
X = X.copy()
# Merge data
X = X.merge(store, left_on='Store', right_on='Store', how='left').fillna(0)
# Engineer date
X['Date'] = pd.to_datetime(X['Date'], infer_datetime_format=True)
X['DateYear'] = X['Date'].dt.year
X['DateMonth'] = X['Date'].dt.month
X = X.drop(columns='Date')
# Drop Sales as our goal is to predict its
X = X.drop(columns='Sales')
return X
# + id="AFM2UNwm_fwI" colab_type="code" colab={}
X_train = wrangle(train)
X_test = wrangle(test)
# + id="ZeOJFJJZzpr1" colab_type="code" outputId="2ebe6633-513d-4c2d-8832-4924cf11d98f" colab={"base_uri": "https://localhost:8080/", "height": 357}
# Permutation Importances plot
import eli5
from eli5.sklearn import PermutationImportance
encoder = ce.OrdinalEncoder()
X_train_encoded = encoder.fit_transform(X_train)
X_test_encoded = encoder.transform(X_test)
model = XGBRegressor(n_estimators=1000, n_jobs=-1)
model.fit(X_train_encoded, y_train_log)
permuter = PermutationImportance(model, scoring='neg_mean_squared_error',
cv='prefit', n_iter=2, random_state=42)
permuter.fit(X_test_encoded, y_test_log)
feature_names = X_test_encoded.columns.tolist()
eli5.show_weights(permuter, top=None, feature_names=feature_names)
# + id="9lG2svFSAysJ" colab_type="code" outputId="e65f0c9d-9c41-4876-bd52-885aa0b88045" colab={"base_uri": "https://localhost:8080/", "height": 501}
# Partial Dependency Plot, 1 feature isolation
from pdpbox.pdp import pdp_isolate, pdp_plot
feature = 'Customers'
isolated = pdp_isolate( model=model,
dataset=X_test_encoded,
model_features=X_test_encoded.columns,
feature=feature)
pdp_plot(isolated, feature_name=feature, figsize=(14,8));
# + id="nfL8Fw5LGFVC" colab_type="code" outputId="e4f6f716-8b6f-4db2-fa80-513f992e21da" colab={"base_uri": "https://localhost:8080/", "height": 585}
# Partial Dependency Plot, 2 feature interaction
from pdpbox.pdp import pdp_interact, pdp_interact_plot
features = ['Customers', 'Promo']
interaction = pdp_interact( model=model,
dataset=X_test_encoded,
model_features=X_test_encoded.columns,
features=features)
pdp_interact_plot(interaction, plot_type='grid', feature_names=features);
|
Nicolae_Dubenco_DS_SC_8_Regression_2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import struct, socket
import numpy as np
import linecache, bisect
import csv
import operator
import json
import os
try:
import ipywidgets as widgets # For jupyter/ipython >= 1.4
except ImportError:
from IPython.html import widgets
from IPython.display import display, Javascript, clear_output
with open('/etc/duxbay.conf') as conf:
for line in conf.readlines():
if "DBNAME=" in line: DBNAME = line.split("=")[1].strip('\n').replace("'","");
elif "IMPALA_DEM=" in line: IMPALA_DEM = line.split("=")[1].strip('\n').replace("'","");
spath = os.getcwd()
path = spath.split("/")
date = path[len(path)-1]
dpath = '/'.join(['data' if var == 'ipynb' else var for var in path]) + '/'
cpath = '/'.join(['context' if var == 'ipynb' else var for var in path][:len(path)-2]) + '/'
sconnect = dpath + 'flow_scores.csv'
threats_file = dpath + 'threats.csv'
iploc = cpath + 'iploc.csv'
nwloc = cpath + 'networkcontext_1.csv'
anchor_ip = ''
ir_f = ''
threat_name = ''
iplist = ''
top_inbound_b=''
top_results = 20
if os.path.isfile(iploc):
iplist = np.loadtxt(iploc,dtype=np.uint32,delimiter=',',usecols={0}, converters={0: lambda s: np.uint32(s.replace('"',''))})
else:
print "No iploc.csv file was found, Map View map won't be created"
# -
# **Functions Definition**
# +
def display_controls(ip_list):
container = widgets.HBox(width=550, height=150)
h_container = widgets.Box(width=550, height=200)
label = widgets.HTML(value='<h2>Expanded Search</h2>')
ip_select = widgets.Select(options=ip_list,height=min(len(ip_list)*18+18,150),width=200)
search_button = widgets.Button(description='Search')
container.children = [ip_select,search_button]
h_container.children = [label,container]
display(h_container)
def search_ip(b):
global anchor_ip
global top_inbound_b
anchor_ip = ip_select.value
if anchor_ip != "":
clear_output()
removeWidget(1)
print "Searching for ip: " + anchor_ip
global ir_f
ir_f = dpath + "ir-" + anchor_ip + ".tsv"
if not os.path.isfile(ir_f) or (os.path.isfile(ir_f) and file_is_empty(ir_f)):
imp_query = (" \"SELECT min(treceived) as firstSeen, max(treceived) as lastSeen, sip as srcIP, dip as dstIP, " +
"sport as SPort, dport AS Dport, count(sip) as conns, max(ipkt) as maxPkts, avg(ipkt) " +
"as avgPkts, max(ibyt) as maxBytes, avg(ibyt) as avgBytes FROM "+DBNAME+".flow WHERE " +
"y="+ date[0:4] +" AND m="+ date[4:6] +" AND d="+ date[6:] +" " +
" AND (sip =\'" + anchor_ip + "\' OR dip=\'" + anchor_ip + "\') GROUP BY sip, dip,sport,dport\" ")
# !impala-shell -i $IMPALA_DEM --print_header -B --output_delimiter='\t' -q $imp_query -o $ir_f
clear_output()
print "\n Looking for additional details..."
get_in_out_and_twoway_conns()
add_geospatial_info()
add_network_context()
print anchor_ip + ": connected to " + str(len(inbound.keys())) + " IPs"
top_inbound_b = get_top_bytes(inbound,top_results)
top_inbound_conns = get_top_conns(inbound,top_results)
top_inbound_b.update(top_inbound_conns) # merge the two dictionaries
display_threat_box(anchor_ip)
search_button.on_click(search_ip)
def display_threat_box(ip):
container_summary = widgets.HBox(width=550, height=150)
separator = widgets.Box(width=550, height=15)
main_container = widgets.Box(width=550, height=200)
threat_header = widgets.HTML(value='<h4>Threat summary for ' + anchor_ip +'</h4>')
threat_title_box = widgets.Text(value='',width=200, placeholder='Threat Title')
threat_summary_box = widgets.Textarea(value='',width=300, height=100)
save_button = widgets.Button(description='Save')
container_summary.children = [threat_summary_box,save_button]
main_container.children = [threat_header, threat_title_box, separator, container_summary]
display(main_container)
def save_threat_summary(b):
clear_output()
removeWidget(1)
global threat_name
global top_inbound_b
print "Creating Story Board elements ..."
generate_attack_map_file(anchor_ip, top_inbound_b, outbound, twoway)
generate_stats(anchor_ip, top_inbound_b, outbound, twoway, threat_name)
generate_dendro(anchor_ip, top_inbound_b, outbound, twoway, date)
details_inbound(anchor_ip,top_inbound_b)
add_threat(anchor_ip, threat_title_box.value, threat_summary_box.value.replace('\n', '\\n'))
print "Story board successfully created for {0}".format(anchor_ip)
save_button.on_click(save_threat_summary)
def details_inbound(ip, inbound):
if ip != "" and len(inbound) > 0:
if os.path.isfile(ir_f):
sbdet_f = dpath + "sbdet-" + ip + ".tsv"
if not os.path.isfile(sbdet_f) or (os.path.isfile(sbdet_f) and file_is_empty(sbdet_f)):
imp_query = "SELECT min(treceived) as tstart, max(treceived) as tend, sip as srcIP, "
+ "dip as dstIP, proto as Proto, sport as SPort, dport AS Dport,ipkt as "
+ "Pkts, ibyt as Bytes FROM "+DBNAME+".flow WHERE "
+ "y="+ date[0:4] +" AND m="+ date[4:6] +" AND d="+ date[6:]
+ " AND ((dip IN({0}) "
+ "AND sip ='{1}') OR "
+ "(sip IN({0}) "
+ "AND dip ='{1}')) GROUP BY sip, dip, proto, sport, dport, ipkt, ibyt SORT BY tstart"
ips = "'" + "','".join(inbound.keys()) + "'"
imp_query = imp_query.format(ips,ip)
# !impala-shell -i $IMPALA_DEM --print_header -B --output_delimiter='\t' -q $imp_query -o $sbdet_f
print "Timeline successfully created"
else:
print "Timeline couldn't be created"
def generate_dendro(ip, inbound, outbound, twoway, date):
dendro_fpath = dpath + 'threat-dendro-' + anchor_ip + ".json"
obj = {
'name':ip,
'children': [],
'time': date
}
#----- Add Inbound Connections-------#
obj["children"].append({'name': 'Inbound Only', 'children': [], 'impact': 0})
in_ctxs = {}
for ip in inbound:
if 'nwloc' in inbound[ip] and len(inbound[ip]['nwloc']) > 0:
ctx = inbound[ip]['nwloc'][2] # get the machine type Only for vast Data
if ctx not in in_ctxs:
in_ctxs[ctx] = 1
else:
in_ctxs[ctx] += 1
for ctx in in_ctxs:
obj["children"][0]['children'].append({
'name': ctx,
'impact': in_ctxs[ctx]
})
#------ Add Outbound ----------------#
obj["children"].append({'name': 'Outbound Only', 'children': [], 'impact': 0})
out_ctxs = {}
for ip in outbound:
if 'nwloc' in outbound[ip] and len(outbound[ip]['nwloc']) > 0:
ctx = outbound[ip]['nwloc'][2] # get the machine type Only for vast Data
if ctx not in out_ctxs:
out_ctxs[ctx] = 1
else:
out_ctxs[ctx] += 1
for ctx in out_ctxs:
obj["children"][1]['children'].append({
'name': ctx,
'impact': out_ctxs[ctx]
})
#------ Add Outbound ----------------#
obj["children"].append({'name': 'two way', 'children': [], 'impact': 0})
tw_ctxs = {}
for ip in twoway:
if 'nwloc' in twoway[ip] and len(twoway[ip]['nwloc']) > 0:
ctx = twoway[ip]['nwloc'][2] # get the machine type Only for vast Data
if ctx not in tw_ctxs:
tw_ctxs[ctx] = 1
else:
tw_ctxs[ctx] += 1
for ctx in tw_ctxs:
obj["children"][2]['children'].append({
'name': ctx,
'impact': tw_ctxs[ctx]
})
with open(dendro_fpath, 'w') as dendro_f:
dendro_f.write(json.dumps(obj))
print 'Incident progression successfully created'
def generate_stats(ip, inbound, outbound, twoway, threat_name):
stats_fpath = dpath + 'stats-' + anchor_ip + ".json"
obj = {
'name':threat_name,
'children': [],
'size': len(inbound) + len(outbound) + len(twoway)
}
#----- Add Inbound Connections-------#
obj["children"].append({'name': 'Inbound Only', 'children': [], 'size': len(inbound)})
in_ctxs = {}
for ip in inbound:
full_ctx = ''
if 'nwloc' in inbound[ip] and len(inbound[ip]['nwloc']) > 0:
full_ctx = inbound[ip]['nwloc'][2].split('.')[0]
ctx = get_ctx_name(full_ctx) # get the machine type Only for vast Data
if ctx not in in_ctxs:
in_ctxs[ctx] = 1
else:
in_ctxs[ctx] += 1
for ctx in in_ctxs:
obj["children"][0]['children'].append({
'name': ctx,
'size': in_ctxs[ctx]
})
#------ Add Outbound ----------------#
obj["children"].append({'name': 'Outbound Only', 'children': [], 'size': len(outbound)})
out_ctxs = {}
for ip in outbound:
full_ctx = ''
if 'nwloc' in outbound[ip] and len(outbound[ip]['nwloc']) > 0:
full_ctx = outbound[ip]['nwloc'][2].split('.')[0]
ctx = get_ctx_name(full_ctx) # get the machine type Only for vast Data
if ctx not in out_ctxs:
out_ctxs[ctx] = 1
else:
out_ctxs[ctx] += 1
for ctx in out_ctxs:
obj["children"][1]['children'].append({
'name': ctx,
'size': out_ctxs[ctx]
})
#------ Add Twoway ----------------#
obj["children"].append({'name': 'two way', 'children': [], 'size': len(twoway)})
tw_ctxs = {}
for ip in twoway:
full_ctx = ''
if 'nwloc' in twoway[ip] and len(twoway[ip]['nwloc']) > 0:
full_ctx = twoway[ip]['nwloc'][2].split('.')[0]
ctx = get_ctx_name(full_ctx) # get the machine type Only for vast Data
if ctx not in tw_ctxs:
tw_ctxs[ctx] = 1
else:
tw_ctxs[ctx] += 1
for ctx in tw_ctxs:
obj["children"][2]['children'].append({
'name': ctx,
'size': tw_ctxs[ctx]
})
json_str = json.dumps(obj)
with open(stats_fpath, 'w') as stats_f:
stats_f.write(json_str)
print 'Stats file successfully created'
def get_ctx_name(full_context):
ctx= 'DMZ'
if "VPN" in full_context:
ctx = "VPN"
elif "DMZ" in full_context:
ctx = "DMZ"
elif "Proxy" in full_context:
ctx = "Proxy"
elif "FW" in full_context:
ctx = "FW"
return ctx
def display_expanded_search():
external_ips = []
c_ips=[]
clear_output()
if os.path.isfile(threats_file) and (os.path.isfile(threats_file) and not file_is_empty(threats_file)):
with open(threats_file, 'r') as th:
t_read = csv.reader(th, delimiter=',')
t_read.next()
for row in t_read:
if row[0] != '' : c_ips.append(row[0])
with open(sconnect, 'r') as f:
reader = csv.reader(f, delimiter=',')
reader.next()
#Internal Netflows use case:
for row in reader:
#sev,tstart,srcIP,dstIP,sport,dport,proto,flag,ipkt,ibyt,lda_score,rank,srcIpInternal,destIpInternal,srcGeo,dstGeo,
# 0 , 1 , 2 , 3 , 4 , 5 , 6 , 7 , 8 , 9 , 10 , 11 , 12 , 13 , 14 , 15 ,
#srcDomain,dstDomain,srcIP_rep,dstIP_rep
# 16 , 17 , 18 , 19
if row[0] == '1':
srcIP = ''
dstIP = ''
if row[2] not in external_ips and row[2] not in c_ips:
external_ips.append(row[2])
if row[3] not in external_ips and row[3] not in c_ips:
external_ips.append(row[3])
if len(external_ips) > 0:
display_controls(external_ips)
else:
print "There are not high risk connections."
# calculate number of inbound only, two-way, and outbound only
# build dict of IP addresses
# firstSeen,lastSeen,srcIP, dstIP, sport,dport,conns, maxPkts, avgPkts,maxBytes, avgBytes
def get_in_out_and_twoway_conns():
global inbound
inbound = {}
global outbound
outbound = {}
global twoway
twoway = {}
srcdict = {}
dstdict = {}
conns_dict= {}
if os.path.isfile(ir_f):
with open(ir_f, 'r') as f:
reader = csv.reader(f,delimiter='\t')
reader.next() #skip headers
rowct = 0
for row in reader:
if row != []:
srcdict[row[2]] = {
'ip_int': struct.unpack("!L", socket.inet_aton(row[2]))[0],
'dst_ip': row[3],
'dst_ip_int': struct.unpack("!L", socket.inet_aton(row[3]))[0],
'conns': int(row[6]),
'maxbytes': int(row[9])
}
dstdict[row[3]] = {
'ip_int': struct.unpack("!L", socket.inet_aton(row[3]))[0],
'src_ip': row[2],
'src_ip_int': struct.unpack("!L", socket.inet_aton(row[2]))[0],
'conns': int(row[6]),
'maxbytes': int(row[9])
}
rowct +=1
if rowct > 0:
for result in srcdict:
if result in dstdict:
twoway[result] = srcdict[result]
else:
outbound[result] = srcdict[result]
for result in dstdict:
if result not in srcdict:
inbound[result] = dstdict[result]
print "Input, Output & Two way connections file detected."
else:
print "Couldn't find any matching connections."
#=========== Adds GEO IP information to the outbound, inbound and twoway connections==============================#
def add_geospatial_info():
# get geospatial info, only when iplocation file is available
if iplist != '':
for srcip in outbound:
reader = csv.reader([linecache.getline(iploc, bisect.bisect(iplist,outbound[srcip]['ip_int'])).replace('\n','')])
outbound[srcip]['geo'] = reader.next()
reader = csv.reader([linecache.getline(iploc, bisect.bisect(iplist,outbound[srcip]['dst_ip_int'])).replace('\n','')])
outbound[srcip]['geo_dst'] = reader.next()
for dstip in twoway:
reader = csv.reader([linecache.getline(iploc, bisect.bisect(iplist,twoway[dstip]['ip_int'])).replace('\n','')])
twoway[dstip]['geo'] = reader.next()
for srcip in inbound:
reader = csv.reader([linecache.getline(iploc, bisect.bisect(iplist,inbound[srcip]['ip_int'])).replace('\n','')])
inbound[srcip]['geo'] = reader.next()
reader = csv.reader([linecache.getline(iploc, bisect.bisect(iplist,inbound[srcip]['src_ip_int'])).replace('\n','')])
inbound[srcip]['geo_src'] = reader.next()
# need some way to combine timelines of outbound and two-way with big picture inbound only
# get network context - get start and end ranges
# need some way to combine timelines of outbound and two-way with big picture inbound only
# get network context - get start and end ranges
def add_network_context():
nwdict = {}
if os.path.isfile(nwloc) :
with open(nwloc, 'r') as f:
reader = csv.reader(f,delimiter=',')
reader.next()
#address range, description
for row in reader:
if '/' in row[0]:
#Range in subnet
iprange = row[0].split('/')
if len(iprange) < 2:
ipend = 0
else:
ipend = int(iprange[1])
nwdict[row[0]] = [struct.unpack("!L", socket.inet_aton(iprange[0]))[0],
struct.unpack("!L", socket.inet_aton(iprange[0]))[0]+2**(32-ipend)-1, row[1]]
elif '-' in row[0]:
#IP Range
iprange = row[0].split('-')
nwdict[row[0]] = [struct.unpack("!L", socket.inet_aton(iprange[0].replace(" ", "")))[0],
struct.unpack("!L", socket.inet_aton(iprange[1].replace(" ", "")))[0], row[1]]
else:
#Exact match
nwdict[row[0]] = [struct.unpack("!L", socket.inet_aton(row[0]))[0],
struct.unpack("!L", socket.inet_aton(row[0]))[0], row[1]]
for srcip in outbound:
temp_ip = struct.unpack("!L", socket.inet_aton(srcip))[0]
if srcip in nwdict:
inbound[srcip]['nwloc'] = nwdict[srcip]
else:
matchingVals = [x for x in nwdict if nwdict[x][1] >= temp_ip and nwdict[x][0] <= temp_ip]
outbound[srcip]['nwloc'] = nwdict[matchingVals[0]] if len(matchingVals) > 0 else ''
for dstip in twoway:
temp_ip = struct.unpack("!L", socket.inet_aton(dstip))[0]
if dstip in nwdict:
twoway[dstip]['nwloc'] = nwdict[dstip]
else:
matchingVals = [x for x in nwdict if nwdict[x][1] >= temp_ip and nwdict[x][0] <= temp_ip]
twoway[dstip]['nwloc'] = nwdict[matchingVals[0]] if len(matchingVals) > 0 else ''
for srcip in inbound:
temp_ip = struct.unpack("!L", socket.inet_aton(srcip))[0]
if srcip in nwdict:
inbound[srcip]['nwloc'] = nwdict[srcip]
else:
matchingVals = [x for x in nwdict if nwdict[x][1] >= temp_ip and nwdict[x][0] <= temp_ip]
inbound[srcip]['nwloc'] = nwdict[matchingVals[0]] if len(matchingVals) > 0 else ''
def generate_attack_map_file(ip, inbound, outbound, twoway):
if iplist != '':
globe_fpath = dpath + 'globe-' + anchor_ip + ".json"
globe_json = {}
globe_json['type'] = "FeatureCollection"
globe_json['sourceips'] = []
globe_json['destips'] = []
for srcip in twoway:
try:
row = twoway[srcip]['geo']
globe_json['destips'].append({
'type': 'Feature',
'properties': {
'location':row[8],
'ip':srcip,
'type':1
},
'geometry': {
'type': 'Point',
'coordinates': [float(row[7]), float(row[6])]
}
})
except ValueError:
pass
for dstip in outbound:
try:
row = outbound[dstip]['geo']
dst_geo = outbound[dstip]['geo_dst']
globe_json['sourceips'].append({
'type': 'Feature',
'properties': {
'location':row[8],
'ip':dstip,
'type':3
},
'geometry': {
'type': 'Point',
'coordinates': [float(row[7]), float(row[6])]
}
})
globe_json['destips'].append({
'type': 'Feature',
'properties': {
'location':row[8],
'ip':outbound[dstip]['dst_ip'],
'type':3
},
'geometry': {
'type': 'Point',
'coordinates': [float(dst_geo[7]), float(dst_geo[6])]
}
})
except ValueError:
pass
for dstip in inbound:
try:
row = inbound[dstip]['geo']
dst_geo = inbound[dstip]['geo_src']
globe_json['sourceips'].append({
'type': 'Feature',
'properties': {
'location':row[8],
'ip':dstip,
'type':2
},
'geometry': {
'type': 'Point',
'coordinates': [float(row[7]), float(row[6])]
}
})
globe_json['destips'].append({
'type': 'Feature',
'properties': {
'location':row[8],
'ip':inbound[dstip]['src_ip'],
'type':2
},
'geometry': {
'type': 'Point',
'coordinates': [float(dst_geo[7]), float(dst_geo[6])]
}
})
except ValueError:
pass
json_str = json.dumps(globe_json)
with open(globe_fpath, 'w') as globe_f:
globe_f.write(json_str)
print "Geolocation map successfully created"
else:
print "The map can't be created without an iploc file"
def add_threat(ip,threat_title, threat_comment):
content = ''
try:
threat_f = open(threats_file, 'r')
content = threat_f.read()
if '{0}|{1}|{2}\n'.format(ip,threat_title,threat_comment) not in content:
content += '{0}|{1}|{2}\n'.format(ip,threat_title,threat_comment)
threat_f.close()
except:
content = 'ip|title|summary\n'
content += '{0}|{1}|{2}\n'.format(ip,threat_title,threat_comment)
threat_fw = open(threats_file, 'w')
threat_fw.write(content)
threat_fw.close()
def get_top_bytes(conns_dict, top):
print "Now looking at the top "+ str(top) +" connections per bytes:"
topbytes = sorted(conns_dict.iteritems(), key=lambda (x,y): y['maxbytes'], reverse=True)
topbytes = topbytes[0:top]
for item in topbytes:
print item[0], "|", item[1]["maxbytes"]
return dict(topbytes)
def get_top_conns(conns_dict, top):
print "Now looking at the top "+str(top)+" connections per number of connections:"
topconns = sorted(conns_dict.iteritems(), key=lambda (x,y): y['conns'], reverse=True)
topconns = topconns[0:top]
for item in topconns:
print item[0], "|", item[1]["conns"]
return dict(topconns)
def file_is_empty(path):
return os.stat(path).st_size==0
def removeWidget(index):
js_command = "$('.widget-area > .widget-subarea > .widget-box:eq({0})').remove();".format(index)
display(Javascript(js_command))
# -
display_expanded_search()
|
spot-demo/assets/spot-oa/ipynb/flow/20160708/Threat_Investigation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: GPUEnv
# language: python
# name: venv
# ---
import tensorflow as tf
import pandas as pd
import numpy as np
import os
import matplotlib.pyplot as plt
import matplotlib.mlab as mlab
import matplotlib as mpl
import cv2
import time
import PIL
from tensorflow import keras
from tensorflow.keras import layers
from tensorflow.keras.applications.resnet50 import ResNet50
from tensorflow.keras.preprocessing import image
from tensorflow.keras.applications.resnet50 import preprocess_input, decode_predictions
import numpy as np
from keras.initializers import glorot_normal
physical_devices = tf.config.experimental.list_physical_devices('GPU')
if len(physical_devices) > 0:
tf.config.experimental.set_memory_growth(physical_devices[0], True)
from matplotlib import pyplot
from keras.layers import Conv2D, MaxPooling2D, Flatten, Dense, Dropout
# +
"""
Calculates dot product of x[0] and x[1] for mini_batch
Assuming both have same size and shape
@param
x -> [ (size_minibatch, total_pixels, size_filter), (size_minibatch, total_pixels, size_filter) ]
"""
def dot_product(x):
return keras.backend.batch_dot(x[0], x[1], axes=[1,1]) / x[0].get_shape().as_list()[1]
"""
Calculate signed square root
@param
x -> a tensor
"""
def signed_sqrt(x):
return keras.backend.sign(x) * keras.backend.sqrt(keras.backend.abs(x) + 1e-9)
"""
Calculate L2-norm
@param
x -> a tensor
"""
def L2_norm(x, axis=-1):
return keras.backend.l2_normalize(x, axis=axis)
# +
'''
Take outputs of last layer of VGG and load it into Lambda layer which calculates outer product.
Here both bi-linear branches have same shape.
z -> output shape tuple
x -> outpur og VGG tensor
y -> copy of x as we modify x, we use x, y for outer product.
'''
def build_model():
tensor_input = keras.layers.Input(shape=[150,150,3])
# load pre-trained model
tensor_input = keras.layers.Input(shape=[150,150,3])
model_detector = keras.applications.vgg16.VGG16(
input_tensor=tensor_input,
include_top=False,
weights='imagenet')
model_detector2 = keras.applications.vgg16.VGG16(
input_tensor=tensor_input,
include_top=False,
weights='imagenet')
model_detector2 = keras.models.Sequential(layers=model_detector2.layers)
for i, layer in enumerate(model_detector2.layers):
layer._name = layer.name +"_second"
model2 = keras.models.Model(inputs=[tensor_input], outputs = [model_detector2.layers[-1].output])
x = model_detector.layers[17].output
z = model_detector.layers[17].output_shape
y = model2.layers[17].output
print(model_detector.summary())
print(model2.summary())
# rehape to (batch_size, total_pixels, filter_size)
x = keras.layers.Reshape([z[1] * z[2] , z[-1]])(x)
y = keras.layers.Reshape([z[1] * z[2] , z[-1]])(y)
# outer products of x, y
x = keras.layers.Lambda(dot_product)([x, y])
# rehape to (batch_size, filter_size_vgg_last_layer*filter_vgg_last_layer)
x = keras.layers.Reshape([z[-1]*z[-1]])(x)
# signed_sqrt
x = keras.layers.Lambda(signed_sqrt)(x)
# L2_norm
x = keras.layers.Lambda(L2_norm)(x)
# FC-Layer
initializer = tf.keras.initializers.GlorotNormal()
x = keras.layers.Dense(units=258,
kernel_regularizer=keras.regularizers.l2(0.0),
kernel_initializer=initializer)(x)
tensor_prediction = keras.layers.Activation("softmax")(x)
model_bilinear = keras.models.Model(inputs=[tensor_input],
outputs=[tensor_prediction])
# Freeze VGG layers
for layer in model_detector.layers:
layer.trainable = False
sgd = keras.optimizers.SGD(lr=1.0,
decay=0.0,
momentum=0.9)
model_bilinear.compile(loss="categorical_crossentropy",
optimizer=sgd,
metrics=["categorical_accuracy"])
model_bilinear.summary()
return model_bilinear
# -
model = build_model()
# +
def train_model(epochs):
hist = model.fit_generator(
train_generator,
epochs=epochs,
validation_data=val_generator,
workers=3,
verbose=1
)
model.save_weights("./bilinear_weights/val_acc_" + hist.history['val_categorical_accuracy'][-1] +"_"+ str(epochs)+ ".h5")
return hist
# -
train_datagen = image.ImageDataGenerator(
rotation_range=40,
width_shift_range=0.2,
height_shift_range=0.2,
fill_mode='nearest',
rescale=1./255,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True)
test_datagen = image.ImageDataGenerator(rescale=1./255)
train_generator = train_datagen.flow_from_directory(
'../nut_snacks/dataset_split/train',
target_size=(150, 150),
color_mode="rgb",
batch_size=32,
subset='training',
class_mode='categorical')
val_generator = test_datagen.flow_from_directory(
'../nut_snacks/dataset_split/val',
target_size=(150, 150),
color_mode="rgb",
batch_size=32,
subset='training',
class_mode='categorical')
test_generator = test_datagen.flow_from_directory(
'../nut_snacks/dataset_split/test',
target_size=(150, 150),
color_mode="rgb",
shuffle = False,
class_mode=None,
batch_size=1)
hist =train_model(epochs=20)
hist =train_model(epochs=20)
# +
for layer in model.layers:
layer.trainable = True
sgd = keras.optimizers.SGD(lr=1e-3, decay=1e-9, momentum=0.9)
model.compile(loss="categorical_crossentropy", optimizer=sgd, metrics=["categorical_accuracy"])
# -
hist =train_model(epochs=30)
model.save('./model_bilin')
model2 = keras.models.load_model('./model_bilin')
preds = model2.predict_generator(test_generator, verbose=1)
preds_cls_idx = preds.argmax(axis=-1)
preds_cls_idx
idx_to_cls = {v: k for k, v in train_generator.class_indices.items()}
preds_cls = np.vectorize(idx_to_cls.get)(preds_cls_idx)
preds_cls
true_lables = []
true_lables_upc_idx_map = {}
true_lables_img = {}
upc_list = os.listdir('../nut_snacks/dataset_split/test/')
idx = 0
for upc in upc_list:
img_folder = '../nut_snacks/dataset_split/test/' + upc +'/'
img_list = os.listdir(img_folder)
for img in img_list:
true_lables.append(upc)
true_lables_upc_idx_map[idx] = upc
true_lables_img[idx] = img
idx += 1
len(true_lables)
wrong_predicted = []
count = 0
for idx in range(0, len(preds_cls)):
if preds_cls[idx] != true_lables[idx]:
wrong_predicted.append(idx)
else:
count += 1
count
len(wrong_predicted)
accuracy = count/len(preds_cls)
accuracy
wrong_pred_upc = set()
for label in wrong_predicted:
wrong_pred_upc.add(true_lables_upc_idx_map[label])
len(wrong_pred_upc), len(wrong_predicted)
images_pred_wrong = []
for label in wrong_predicted:
images_pred_wrong.append(true_lables_img[label])
len(images_pred_wrong)
d={}
for i in range(0 ,len(images_pred_wrong)):
if preds_cls[i] not in d.keys():
d[preds_cls[i]] = 1
else:
d[preds_cls[i]] += 1
d
for i in range(0 ,len(images_pred_wrong)):
img ='../nut_snacks/dataset_split/test/' + true_lables_upc_idx_map[wrong_predicted[i]] + '/' + images_pred_wrong[i]
print(img, preds_cls[i])
from sklearn.metrics import f1_score,precision_score,recall_score,accuracy_score
f1 = f1_score(true_lables, preds_cls, average='weighted')
precision = precision_score(true_lables, preds_cls, average='weighted')
recall = recall_score(true_lables, preds_cls, average='weighted')
accuracy = accuracy_score(true_lables, preds_cls)
print("f1 :", f1)
print("precision :", precision)
print("recall :", recall)
print("accuracy :", accuracy)
|
BILINEAR.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# ## Description
#
# UPDATE: In this version of the kernel we will try to test the idea of selecting features using LOFO. For more details about LOFO please see Ahmet Erdem's kernel available [at this link](https://www.kaggle.com/divrikwicky/instantgratification-lofo-feature-importance). The feature selection step is going to slow down the training process, so this new version will run longer than 1 minute. If you want to see the original kernel that runs less than a minute please refer to Version 1 of this kernel.
#
# The original kernel scores 0.99610 on the LB. Unfortunately, we won't be able to use this result as a baseline for comparison because we won't be able to submit our work to LB: in order for LOFO to work, an external package, `lofo-importance`, must be loaded but the usage of external packages is banned by the competion rules. However, it is possible to compute the cross-validation score for the QDA model without LOFO. As a matter of fact, I have already done it in a different kernel: [link](https://www.kaggle.com/graf10a/tuning-512-separate-qda-models) (see the "Repeat Using the Standard Parameters" section). The result was a CV score of 0.96629. Let's see if selecting features with LOFO can improve this baseline.
#
# SPOILER: Basically, the resutl is very inconclusive -- the combined AUC went up from 0.96629 to 0.96727, the fold-average AUC went down from 0.96628 to 0.96213, and the standard deviation increased from 9e-05 to 0.0097. It would be nice to submit it to the LB to see how well it performs.
# ## Setting things up
# ### Loading Libraries
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
import warnings
warnings.filterwarnings('ignore')
import numpy as np
import pandas as pd
from sklearn.pipeline import Pipeline
from sklearn.metrics import roc_auc_score
from sklearn.model_selection import StratifiedKFold
from sklearn.feature_selection import VarianceThreshold
from sklearn.preprocessing import StandardScaler
from sklearn.discriminant_analysis import QuadraticDiscriminantAnalysis
# -
# ### Loading Data
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
# %%time
train = pd.read_csv('../input/train.csv')
test = pd.read_csv('../input/test.csv')
train['wheezy-copper-turtle-magic'] = train['wheezy-copper-turtle-magic'].astype('category')
test['wheezy-copper-turtle-magic'] = test['wheezy-copper-turtle-magic'].astype('category')
# -
# ### Computing LOFO Importance
#
# Here is the adapted code from [Ahmet's notebook](https://www.kaggle.com/divrikwicky/instantgratification-lofo-feature-importance):
# +
from lofo import LOFOImportance, FLOFOImportance, plot_importance
from tqdm import tqdm_notebook
def get_model():
return Pipeline([('scaler', StandardScaler()),
('qda', QuadraticDiscriminantAnalysis(reg_param=0.111))
])
features = [c for c in train.columns if c not in ['id', 'target', 'wheezy-copper-turtle-magic']]
def get_lofo_importance(wctm_num):
sub_df = train[train['wheezy-copper-turtle-magic'] == wctm_num]
sub_features = [f for f in features if sub_df[f].std() > 1.5]
lofo_imp = LOFOImportance(sub_df, target="target",
features=sub_features,
cv=StratifiedKFold(n_splits=4, random_state=42, shuffle=True), scoring="roc_auc",
model=get_model(), n_jobs=4)
return lofo_imp.get_importance()
features_to_remove = []
potential_gain = []
n_models=512
for i in tqdm_notebook(range(n_models)):
imp = get_lofo_importance(i)
features_to_remove.append(imp["feature"].values[-1])
potential_gain.append(-imp["importance_mean"].values[-1])
print("Potential gain (AUC):", np.round(np.mean(potential_gain), 5))
# -
# ## Building the QDA Classifier with LOFO
# ### Preparing Things for Cross-Validation
# +
clf_name='QDA'
NFOLDS=25
RS=42
oof=np.zeros(len(train))
preds=np.zeros(len(test))
# -
# ### Training the Classifiers on All Data
# +
# %%time
print(f'Cross-validation for the {clf_name} classifier:')
default_cols = [c for c in train.columns if c not in ['id', 'target', 'wheezy-copper-turtle-magic']]
# BUILD 512 SEPARATE NON-LINEAR MODELS
for i in range(512):
# EXTRACT SUBSET OF DATASET WHERE WHEEZY-MAGIC EQUALS i
X = train[train['wheezy-copper-turtle-magic']==i].copy()
Y = X.pop('target').values
X_test = test[test['wheezy-copper-turtle-magic']==i].copy()
idx_train = X.index
idx_test = X_test.index
X.reset_index(drop=True,inplace=True)
#cols = [c for c in X.columns if c not in ['id', 'wheezy-copper-turtle-magic']]
cols = [c for c in default_cols if c != features_to_remove[i]]
X = X[cols].values # numpy.ndarray
X_test = X_test[cols].values # numpy.ndarray
# FEATURE SELECTION (USE APPROX 40 OF 255 FEATURES)
vt = VarianceThreshold(threshold=1.5).fit(X)
X = vt.transform(X) # numpy.ndarray
X_test = vt.transform(X_test) # numpy.ndarray
# STRATIFIED K FOLD
auc_all_folds=np.array([])
folds = StratifiedKFold(n_splits=NFOLDS, random_state=RS)
for fold_num, (train_index, val_index) in enumerate(folds.split(X, Y), 1):
X_train, Y_train = X[train_index, :], Y[train_index]
X_val, Y_val = X[val_index, :], Y[val_index]
pipe = Pipeline([('scaler', StandardScaler()),
(clf_name, QuadraticDiscriminantAnalysis(reg_param=0.111)),
])
pipe.fit(X_train, Y_train)
oof[idx_train[val_index]] = pipe.predict_proba(X_val)[:,1]
preds[idx_test] += pipe.predict_proba(X_test)[:,1]/NFOLDS
auc = roc_auc_score(Y_val, oof[idx_train[val_index]])
auc_all_folds = np.append(auc_all_folds, auc)
# PRINT CROSS-VALIDATION AUC FOR THE CLASSFIER
auc_combo = roc_auc_score(train['target'].values, oof)
auc_folds_average = np.mean(auc_all_folds)
std = np.std(auc_all_folds)/np.sqrt(NFOLDS)
print(f'The combined CV score is {round(auc_combo,5)}.')
print(f'The folds average CV score is {round(auc_folds_average,5)}.')
print(f'The standard deviation is {round(std, 5)}.')
# -
# ## Creating the Submission File
#
# All done! At this point we are ready to make our submission file! (We won't be able to submit it but let's make it anyway.)
sub = pd.read_csv('../input/sample_submission.csv')
sub['target'] = preds
sub.to_csv('submission.csv',index=False)
sub.shape
sub.head()
|
5 instant gratification/single-qda-lb-0-96610-time-1-min.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: feml
# language: python
# name: feml
# ---
# ## Target guided encodings
#
# In the previous lectures in this section, we learned how to convert a label into a number, by using one hot encoding, replacing by a digit or replacing by frequency or counts of observations. These methods are simple, make (almost) no assumptions and work generally well in different scenarios.
#
# There are however methods that allow us to capture information while pre-processing the labels of categorical variables. These methods include:
#
# - Ordering the labels according to the target
# - Replacing labels by the target mean (mean encoding / target encoding)
# - Replacing the labels by the probability ratio of the target being 1 or 0
# - Weight of evidence.
#
# All of the above methods have something in common:
#
# - the encoding is **guided by the target**, and
# - they create a **monotonic relationship** between the variable and the target.
#
#
# ### Monotonicity
#
# A monotonic relationship is a relationship that does one of the following:
#
# - (1) as the value of one variable increases, so does the value of the other variable; or
# - (2) as the value of one variable increases, the value of the other variable decreases.
#
# In this case, as the value of the independent variable (predictor) increases, so does the target, or conversely, as the value of the variable increases, the target value decreases.
#
#
#
# ### Advantages of target guided encodings
#
# - Capture information within the category, therefore creating more predictive features
# - Create a monotonic relationship between the variable and the target, therefore suitable for linear models
# - Do not expand the feature space
#
#
# ### Limitations
#
# - Prone to cause over-fitting
# - Difficult to cross-validate with current libraries
#
#
# ### Note
#
# The methods discussed in this and the coming 3 lectures can be also used on numerical variables, after discretisation. This creates a monotonic relationship between the numerical variable and the target, and therefore improves the performance of linear models. I will discuss this in more detail in the section "Discretisation".
#
# ===============================================================================
#
# ## Probability Ratio Encoding
#
# These encoding is suitable for classification problems only, where the target is binary.
#
# For each category, we calculate the mean of target=1, that is the probability of the target being 1 ( P(1) ), and the probability of the target=0 ( P(0) ). And then, we calculate the ratio P(1)/P(0), and replace the categories by that ratio.
#
#
# ## In this demo:
#
# We will see how to perform one hot encoding with:
# - pandas
# - Feature-Engine
#
# And the advantages and limitations of each implementation using the Titanic dataset.
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# to split the datasets
from sklearn.model_selection import train_test_split
# for encoding with feature-engine
from feature_engine.encoding import PRatioEncoder
# +
# load dataset
data = pd.read_csv(
'../titanic.csv',
usecols=['cabin', 'sex', 'embarked', 'survived'])
data.head()
# +
# let's remove obserrvations with na in embarked
data.dropna(subset=['embarked'], inplace=True)
data.shape
# +
# Now we extract the first letter of the cabin
# to create a simpler variable for the demo
data['cabin'] = data['cabin'].astype(str).str[0]
# +
# and we remove the observations where cabin = T
# because they are too few
data = data[data['cabin']!= 'T']
data.shape
# +
# let's have a look at how many labels each variable has
for col in data.columns:
print(col, ': ', len(data[col].unique()), ' labels')
# -
# let's explore the unique categories
data['cabin'].unique()
data['sex'].unique()
data['embarked'].unique()
# ### Encoding important
#
# We calculate the ratio P(1)/P(0) using the train set, and then use those mappings in the test set.
#
# Note that to implement this in pandas, we need to keep the target in the training set.
# +
# let's separate into training and testing set
X_train, X_test, y_train, y_test = train_test_split(
data[['cabin', 'sex', 'embarked', 'survived']], # this time we keep the target!!
data['survived'], # target
test_size=0.3, # percentage of obs in test set
random_state=0) # seed to ensure reproducibility
X_train.shape, X_test.shape
# -
# ### Explore original relationship between categorical variables and target
# +
# let's explore the relationship of the categories with the target
for var in ['cabin', 'sex', 'embarked']:
fig = plt.figure()
fig = X_train.groupby([var])['survived'].mean().plot()
fig.set_title('Relationship between {} and Survival'.format(var))
fig.set_ylabel('Mean Survival')
plt.show()
# -
# You can see that the relationship between the target and cabin and embarked goes up and down, depending on the category.
#
#
# ## Probability ratio encoding with pandas
#
#
# ### Advantages
#
# - quick
# - returns pandas dataframe
#
# ### Limitations of pandas:
#
# - it does not preserve information from train data to propagate to test data
# +
# let's calculate the probability of survived = 1 per category
prob_df = X_train.groupby(['cabin'])['survived'].mean()
# and capture it into a dataframe
prob_df = pd.DataFrame(prob_df)
prob_df
# +
# and now the probability of survived = 0
prob_df['died'] = 1 - prob_df['survived']
prob_df
# +
# and now the ratio
prob_df['ratio'] = prob_df['survived'] / prob_df['died']
prob_df
# +
# and now let's capture the ratio in a dictionary
ordered_labels = prob_df['ratio'].to_dict()
ordered_labels
# +
# now, we replace the labels with the ratios
X_train['cabin'] = X_train['cabin'].map(ordered_labels)
X_test['cabin'] = X_test['cabin'].map(ordered_labels)
# +
# let's explore the result
X_train['cabin'].head(10)
# +
# we can turn the previous commands into 2 functions
def find_category_mappings(df, variable, target):
tmp = pd.DataFrame(df.groupby([variable])[target].mean())
tmp['non-target'] = 1 - tmp[target]
tmp['ratio'] = tmp[target] / tmp['non-target']
return tmp['ratio'].to_dict()
def integer_encode(train, test, variable, ordinal_mapping):
train[variable] = train[variable].map(ordinal_mapping)
test[variable] = test[variable].map(ordinal_mapping)
# +
# and now we run a loop over the remaining categorical variables
for variable in ['sex', 'embarked']:
mappings = find_category_mappings(X_train, variable, 'survived')
integer_encode(X_train, X_test, variable, mappings)
# +
# let's see the result
X_train.head()
# +
# let's inspect the newly created monotonic relationship
# between the categorical variables and the target
for var in ['cabin', 'sex', 'embarked']:
fig = plt.figure()
fig = X_train.groupby([var])['survived'].mean().plot()
fig.set_title('Monotonic relationship between {} and Survival'.format(var))
fig.set_ylabel('Mean Survived')
plt.show()
# -
# Note the monotonic relationships between the mean target and the categories.
#
# ### Note
#
# Replacing categorical labels with this code and method will generate missing values for categories present in the test set that were not seen in the training set. Therefore it is extremely important to handle rare labels before-hand. I will explain how to do this, in a later notebook.
#
# **In addition, it will create NA or Inf if the probability of target = 0 is zero, as the division by zero is not defined.**
# ## Probability Ratio Encoding with Feature-Engine
#
# If using Feature-Engine, instead of pandas, we do not need to keep the target variable in the training dataset.
# +
# let's separate into training and testing set
X_train, X_test, y_train, y_test = train_test_split(
data[['cabin', 'sex', 'embarked']], # predictors
data['survived'], # target
test_size=0.3, # percentage of obs in test set
random_state=0) # seed to ensure reproducibility
X_train.shape, X_test.shape
# -
ratio_enc = PRatioEncoder(
encoding_method = 'ratio',
variables=['cabin', 'sex', 'embarked'])
# +
# when fitting the transformer, we need to pass the target as well
# just like with any Scikit-learn predictor class
ratio_enc.fit(X_train, y_train)
# +
# in the encoder dict we see the P(1)/P(0) for each
# category for each of the indicated variables
ratio_enc.encoder_dict_
# +
# this is the list of variables that the encoder will transform
ratio_enc.variables_
# +
X_train = ratio_enc.transform(X_train)
X_test = ratio_enc.transform(X_test)
# let's explore the result
X_train.head()
# -
# **Note**
#
# If the argument variables is left to None, then the encoder will automatically identify all categorical variables. Is that not sweet?
#
# The encoder will not encode numerical variables. So if some of your numerical variables are in fact categories, you will need to re-cast them as object before using the encoder.
#
# If there is a label in the test set that was not present in the train set, the encoder will through and error, to alert you of this behaviour.
#
# Finally, if the probability of target = 0 is zero for any category, the encoder will raise an error as the division by zero is not defined.
|
Section-06-Categorical-Encoding/06.07-Probability-Ratio-Encoding.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
# !wget https://huseinhouse-storage.s3-ap-southeast-1.amazonaws.com/bert-bahasa/dictionary-pos.json
# !wget https://huseinhouse-storage.s3-ap-southeast-1.amazonaws.com/bert-bahasa/session-pos.pkl
# + _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a" _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0"
import pickle
import json
import tensorflow as tf
import numpy as np
# -
with open('session-pos.pkl', 'rb') as fopen:
data = pickle.load(fopen)
data.keys()
train_X = data['train_X']
test_X = data['test_X']
train_Y = data['train_Y']
test_Y = data['test_Y']
with open('dictionary-pos.json') as fopen:
dictionary = json.load(fopen)
dictionary.keys()
word2idx = dictionary['word2idx']
idx2word = {int(k): v for k, v in dictionary['idx2word'].items()}
tag2idx = dictionary['tag2idx']
idx2tag = {int(k): v for k, v in dictionary['idx2tag'].items()}
char2idx = dictionary['char2idx']
list(zip([idx2word[d] for d in train_X[-1]], [idx2tag[d] for d in train_Y[-1]]))
def generate_char_seq(batch):
x = [[len(idx2word[i]) for i in k] for k in batch]
maxlen = max([j for i in x for j in i])
temp = np.zeros((batch.shape[0],batch.shape[1],maxlen),dtype=np.int32)
for i in range(batch.shape[0]):
for k in range(batch.shape[1]):
for no, c in enumerate(idx2word[batch[i,k]]):
temp[i,k,-1-no] = char2idx[c]
return temp
generate_char_seq(data['train_X'][:10]).shape
class Model:
def __init__(
self,
dim_word,
dim_char,
dropout,
learning_rate,
hidden_size_char,
hidden_size_word,
num_layers,
):
def cells(size, reuse = False):
return tf.contrib.rnn.DropoutWrapper(
tf.nn.rnn_cell.LSTMCell(
size,
initializer = tf.orthogonal_initializer(),
reuse = reuse,
),
output_keep_prob = dropout,
)
def luong(embedded, size):
attention_mechanism = tf.contrib.seq2seq.LuongAttention(
num_units = hidden_size_word, memory = embedded
)
return tf.contrib.seq2seq.AttentionWrapper(
cell = cells(hidden_size_word),
attention_mechanism = attention_mechanism,
attention_layer_size = hidden_size_word,
)
self.word_ids = tf.placeholder(tf.int32, shape = [None, None])
self.char_ids = tf.placeholder(tf.int32, shape = [None, None, None])
self.labels = tf.placeholder(tf.int32, shape = [None, None])
self.maxlen = tf.shape(self.word_ids)[1]
self.lengths = tf.count_nonzero(self.word_ids, 1)
self.word_embeddings = tf.Variable(
tf.truncated_normal(
[len(word2idx), dim_word], stddev = 1.0 / np.sqrt(dim_word)
)
)
self.char_embeddings = tf.Variable(
tf.truncated_normal(
[len(char2idx), dim_char], stddev = 1.0 / np.sqrt(dim_char)
)
)
word_embedded = tf.nn.embedding_lookup(
self.word_embeddings, self.word_ids
)
char_embedded = tf.nn.embedding_lookup(
self.char_embeddings, self.char_ids
)
s = tf.shape(char_embedded)
char_embedded = tf.reshape(
char_embedded, shape = [s[0] * s[1], s[-2], dim_char]
)
for n in range(num_layers):
(out_fw, out_bw), (
state_fw,
state_bw,
) = tf.nn.bidirectional_dynamic_rnn(
cell_fw = cells(hidden_size_char),
cell_bw = cells(hidden_size_char),
inputs = char_embedded,
dtype = tf.float32,
scope = 'bidirectional_rnn_char_%d' % (n),
)
char_embedded = tf.concat((out_fw, out_bw), 2)
output = tf.reshape(
char_embedded[:, -1], shape = [s[0], s[1], 2 * hidden_size_char]
)
word_embedded = tf.concat([word_embedded, output], axis = -1)
for n in range(num_layers):
(out_fw, out_bw), (
state_fw,
state_bw,
) = tf.nn.bidirectional_dynamic_rnn(
cell_fw = luong(word_embedded, hidden_size_word),
cell_bw = luong(word_embedded, hidden_size_word),
inputs = word_embedded,
dtype = tf.float32,
scope = 'bidirectional_rnn_word_%d' % (n),
)
word_embedded = tf.concat((out_fw, out_bw), 2)
logits = tf.layers.dense(word_embedded, len(idx2tag))
y_t = self.labels
log_likelihood, transition_params = tf.contrib.crf.crf_log_likelihood(
logits, y_t, self.lengths
)
self.cost = tf.reduce_mean(-log_likelihood)
self.optimizer = tf.train.AdamOptimizer(
learning_rate = learning_rate
).minimize(self.cost)
mask = tf.sequence_mask(self.lengths, maxlen = self.maxlen)
self.tags_seq, tags_score = tf.contrib.crf.crf_decode(
logits, transition_params, self.lengths
)
self.tags_seq = tf.identity(self.tags_seq, name = 'logits')
y_t = tf.cast(y_t, tf.int32)
self.prediction = tf.boolean_mask(self.tags_seq, mask)
mask_label = tf.boolean_mask(y_t, mask)
correct_pred = tf.equal(self.prediction, mask_label)
correct_index = tf.cast(correct_pred, tf.float32)
self.accuracy = tf.reduce_mean(tf.cast(correct_pred, tf.float32))
class Model:
def __init__(
self,
dim_word,
dim_char,
dropout,
learning_rate,
hidden_size_char,
hidden_size_word,
num_layers,
):
def cells(size, reuse = False):
return tf.contrib.rnn.DropoutWrapper(
tf.nn.rnn_cell.LSTMCell(
size,
initializer = tf.orthogonal_initializer(),
reuse = reuse,
),
output_keep_prob = dropout,
)
def luong(embedded, size):
attention_mechanism = tf.contrib.seq2seq.LuongAttention(
num_units = hidden_size_word, memory = embedded
)
return tf.contrib.seq2seq.AttentionWrapper(
cell = cells(hidden_size_word),
attention_mechanism = attention_mechanism,
attention_layer_size = hidden_size_word,
)
self.word_ids = tf.placeholder(tf.int32, shape = [None, None])
self.char_ids = tf.placeholder(tf.int32, shape = [None, None, None])
self.labels = tf.placeholder(tf.int32, shape = [None, None])
self.maxlen = tf.shape(self.word_ids)[1]
self.lengths = tf.count_nonzero(self.word_ids, 1)
self.word_embeddings = tf.Variable(
tf.truncated_normal(
[len(word2idx), dim_word], stddev = 1.0 / np.sqrt(dim_word)
)
)
self.char_embeddings = tf.Variable(
tf.truncated_normal(
[len(char2idx), dim_char], stddev = 1.0 / np.sqrt(dim_char)
)
)
word_embedded = tf.nn.embedding_lookup(
self.word_embeddings, self.word_ids
)
char_embedded = tf.nn.embedding_lookup(
self.char_embeddings, self.char_ids
)
s = tf.shape(char_embedded)
char_embedded = tf.reshape(
char_embedded, shape = [s[0] * s[1], s[-2], dim_char]
)
for n in range(num_layers):
(out_fw, out_bw), (
state_fw,
state_bw,
) = tf.nn.bidirectional_dynamic_rnn(
cell_fw = cells(hidden_size_char),
cell_bw = cells(hidden_size_char),
inputs = char_embedded,
dtype = tf.float32,
scope = 'bidirectional_rnn_char_%d' % (n),
)
char_embedded = tf.concat((out_fw, out_bw), 2)
output = tf.reshape(
char_embedded[:, -1], shape = [s[0], s[1], 2 * hidden_size_char]
)
word_embedded = tf.concat([word_embedded, output], axis = -1)
for n in range(num_layers):
(out_fw, out_bw), (
state_fw,
state_bw,
) = tf.nn.bidirectional_dynamic_rnn(
cell_fw = luong(word_embedded, hidden_size_word),
cell_bw = luong(word_embedded, hidden_size_word),
inputs = word_embedded,
dtype = tf.float32,
scope = 'bidirectional_rnn_word_%d' % (n),
)
word_embedded = tf.concat((out_fw, out_bw), 2)
logits = tf.layers.dense(word_embedded, len(idx2tag))
y_t = self.labels
log_likelihood, transition_params = tf.contrib.crf.crf_log_likelihood(
logits, y_t, self.lengths
)
self.cost = tf.reduce_mean(-log_likelihood)
self.optimizer = tf.train.AdamOptimizer(
learning_rate = learning_rate
).minimize(self.cost)
mask = tf.sequence_mask(self.lengths, maxlen = self.maxlen)
self.tags_seq, tags_score = tf.contrib.crf.crf_decode(
logits, transition_params, self.lengths
)
self.tags_seq = tf.identity(self.tags_seq, name = 'logits')
y_t = tf.cast(y_t, tf.int32)
self.prediction = tf.boolean_mask(self.tags_seq, mask)
mask_label = tf.boolean_mask(y_t, mask)
correct_pred = tf.equal(self.prediction, mask_label)
correct_index = tf.cast(correct_pred, tf.float32)
self.accuracy = tf.reduce_mean(tf.cast(correct_pred, tf.float32))
# +
tf.reset_default_graph()
sess = tf.InteractiveSession()
dim_word = 128
dim_char = 256
dropout = 0.8
learning_rate = 1e-3
hidden_size_char = 128
hidden_size_word = 128
num_layers = 2
batch_size = 64
model = Model(dim_word,dim_char,dropout,learning_rate,hidden_size_char,hidden_size_word,num_layers)
sess.run(tf.global_variables_initializer())
# +
string = 'KUALA LUMPUR: Sempena sambutan Aidilfitri minggu depan, Perdana Menteri Tun Dr <NAME> dan Menteri Pengangkutan <NAME> <NAME> menitipkan pesanan khas kepada orang ramai yang mahu pulang ke kampung halaman masing-masing. Dalam video pendek terbitan Jabatan Keselamatan Jalan Raya (JKJR) itu, Dr Mahathir menasihati mereka supaya berhenti berehat dan tidur sebentar sekiranya mengantuk ketika memandu.'
import re
def entities_textcleaning(string, lowering = False):
"""
use by entities recognition, pos recognition and dependency parsing
"""
string = re.sub('[^A-Za-z0-9\-\/() ]+', ' ', string)
string = re.sub(r'[ ]+', ' ', string).strip()
original_string = string.split()
if lowering:
string = string.lower()
string = [
(original_string[no], word.title() if word.isupper() else word)
for no, word in enumerate(string.split())
if len(word)
]
return [s[0] for s in string], [s[1] for s in string]
def char_str_idx(corpus, dic, UNK = 0):
maxlen = max([len(i) for i in corpus])
X = np.zeros((len(corpus), maxlen))
for i in range(len(corpus)):
for no, k in enumerate(corpus[i][:maxlen][::-1]):
val = dic[k] if k in dic else UNK
X[i, -1 - no] = val
return X
# +
from tqdm import tqdm
import time
EARLY_STOPPING, CURRENT_CHECKPOINT, CURRENT_ACC, EPOCH = 2, 0, 0, 0
while True:
lasttime = time.time()
if CURRENT_CHECKPOINT == EARLY_STOPPING:
print('break epoch:%d\n' % (EPOCH))
break
lasttime = time.time()
train_acc, train_loss, test_acc, test_loss = 0, 0, 0, 0
pbar = tqdm(
range(0, train_X.shape[0], batch_size), desc = 'train minibatch loop'
)
for i in pbar:
index = min(i + batch_size, train_X.shape[0])
batch_x = train_X[i : index]
batch_char = generate_char_seq(batch_x)
batch_y = train_Y[i : index]
acc, cost, _ = sess.run(
[model.accuracy, model.cost, model.optimizer],
feed_dict = {
model.word_ids: batch_x,
model.char_ids: batch_char,
model.labels: batch_y
},
)
assert not np.isnan(cost)
train_loss += cost
train_acc += acc
pbar.set_postfix(cost = cost, accuracy = acc)
pbar = tqdm(
range(0, test_X.shape[0], batch_size), desc = 'test minibatch loop'
)
for i in pbar:
index = min(i + batch_size, test_X.shape[0])
batch_x = test_X[i : index]
batch_char = generate_char_seq(batch_x)
batch_y = test_Y[i : index]
acc, cost = sess.run(
[model.accuracy, model.cost],
feed_dict = {
model.word_ids: batch_x,
model.char_ids: batch_char,
model.labels: batch_y
},
)
assert not np.isnan(cost)
test_loss += cost
test_acc += acc
pbar.set_postfix(cost = cost, accuracy = acc)
train_loss /= len(train_X) / batch_size
train_acc /= len(train_X) / batch_size
test_loss /= len(test_X) / batch_size
test_acc /= len(test_X) / batch_size
print('time taken:', time.time() - lasttime)
print(
'epoch: %d, training loss: %f, training acc: %f, valid loss: %f, valid acc: %f\n'
% (EPOCH, train_loss, train_acc, test_loss, test_acc)
)
sequence = entities_textcleaning(string)[1]
X_seq = char_str_idx([sequence], word2idx, 2)
X_char_seq = generate_char_seq(X_seq)
predicted = sess.run(model.tags_seq,
feed_dict = {
model.word_ids: X_seq,
model.char_ids: X_char_seq,
},
)[0]
for i in range(len(predicted)):
print(sequence[i],idx2tag[predicted[i]])
if test_acc > CURRENT_ACC:
print(
'epoch: %d, pass acc: %f, current acc: %f'
% (EPOCH, CURRENT_ACC, test_acc)
)
CURRENT_ACC = test_acc
CURRENT_CHECKPOINT = 0
else:
CURRENT_CHECKPOINT += 1
EPOCH += 1
# +
sequence = entities_textcleaning('<NAME> 19977')[1]
X_seq = char_str_idx([sequence], word2idx, 2)
X_char_seq = generate_char_seq(X_seq)
predicted = sess.run(model.tags_seq,
feed_dict = {
model.word_ids: X_seq,
model.char_ids: X_char_seq,
},
)[0]
for i in range(len(predicted)):
print(sequence[i],idx2tag[predicted[i]])
# -
def pred2label(pred):
out = []
for pred_i in pred:
out_i = []
for p in pred_i:
out_i.append(idx2tag[p])
out.append(out_i)
return out
# +
real_Y, predict_Y = [], []
pbar = tqdm(
range(0, len(test_X), batch_size), desc = 'validation minibatch loop'
)
for i in pbar:
batch_x = test_X[i : min(i + batch_size, test_X.shape[0])]
batch_char = generate_char_seq(batch_x)
batch_y = test_Y[i : min(i + batch_size, test_X.shape[0])]
predicted = pred2label(sess.run(model.tags_seq,
feed_dict = {
model.word_ids: batch_x,
model.char_ids: batch_char,
},
))
real = pred2label(batch_y)
predict_Y.extend(predicted)
real_Y.extend(real)
# -
from sklearn.metrics import classification_report
print(classification_report(np.array(real_Y).ravel(), np.array(predict_Y).ravel(),
digits = 6))
# +
saver = tf.train.Saver(tf.trainable_variables())
saver.save(sess, 'luong/model.ckpt')
strings = ','.join(
[
n.name
for n in tf.get_default_graph().as_graph_def().node
if ('Variable' in n.op
or 'Placeholder' in n.name
or 'logits' in n.name
or 'alphas' in n.name)
and 'Adam' not in n.name
and 'beta' not in n.name
and 'OptimizeLoss' not in n.name
and 'Global_Step' not in n.name
]
)
strings.split(',')
# +
def freeze_graph(model_dir, output_node_names):
if not tf.gfile.Exists(model_dir):
raise AssertionError(
"Export directory doesn't exists. Please specify an export "
'directory: %s' % model_dir
)
checkpoint = tf.train.get_checkpoint_state(model_dir)
input_checkpoint = checkpoint.model_checkpoint_path
absolute_model_dir = '/'.join(input_checkpoint.split('/')[:-1])
output_graph = absolute_model_dir + '/frozen_model.pb'
clear_devices = True
with tf.Session(graph = tf.Graph()) as sess:
saver = tf.train.import_meta_graph(
input_checkpoint + '.meta', clear_devices = clear_devices
)
saver.restore(sess, input_checkpoint)
output_graph_def = tf.graph_util.convert_variables_to_constants(
sess,
tf.get_default_graph().as_graph_def(),
output_node_names.split(','),
)
with tf.gfile.GFile(output_graph, 'wb') as f:
f.write(output_graph_def.SerializeToString())
print('%d ops in the final graph.' % len(output_graph_def.node))
def load_graph(frozen_graph_filename):
with tf.gfile.GFile(frozen_graph_filename, 'rb') as f:
graph_def = tf.GraphDef()
graph_def.ParseFromString(f.read())
with tf.Graph().as_default() as graph:
tf.import_graph_def(graph_def)
return graph
# -
freeze_graph('luong', strings)
# +
import boto3
bucketName = 'huseinhouse-storage'
Key = 'luong/frozen_model.pb'
outPutname = "v27/pos/luong-pos.pb"
s3 = boto3.client('s3',
aws_access_key_id='',
aws_secret_access_key='')
s3.upload_file(Key,bucketName,outPutname)
# -
|
session/pos/pos-luong.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:PIC16B] *
# language: python
# name: conda-env-PIC16B-py
# ---
# ### This notebook is a special explaination for data cleaning and analysis. Although we have defined a few functions in `training_module.py` and `training.py` with explanatory docstrings, we will not call them in this notebook instead we will reproduce those codes in order to explain in detail as much as possible.
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# ### The following is the process of data cleaning, please see the comments next to the code for details
# +
#data cleaning
#https://www.kaggle.com/paramaggarwal/fashion-product-images-small
def df_drop(styles, col, item):
"""
This function drops certain columns
input: styles, dataframe
col, the item we want to drop in this coloumn
item, which item we want to drop
"""
for i in item:
styles = styles.drop(styles[styles[col] == i].index)
return styles
def get_df():
"""
this function get and clean the data, return a dataframe
"""
styles = pd.read_csv("/Users/pingkefan/Desktop/archive/styles.csv", error_bad_lines=False)
styles = styles.drop(["productDisplayName"],axis = 1) #drop useless column, we do not need name to do recommendation
styles = styles.drop(["year"],axis = 1) #drop useless column, we do not need year to do recommendation
styles = styles[(styles.masterCategory=='Apparel')| (styles.masterCategory=='Footwear')] # drop useless rows, we are not recommend acessories
styles = styles.drop(styles[styles["subCategory"] == "Innerwear"].index) # drop useless row, we are not recommend innerwears, only outfits.
styles = styles.dropna() # drop NA
styles = df_drop(styles,"subCategory", ["Apparel Set", "Dress","Loungewear and Nightwear","Saree","Socks"]) # we only recommend outfits.
styles["subCategory"] = styles["subCategory"].transform(lambda x: "Footwear" if(x in ["Shoes","Flip Flops","Sandal"]) else x) # Group them into one category.
styles = styles.drop(labels=[6695,16194,32309,36381,40000], axis=0) # drop incomplete rows
return styles
styles = get_df()
styles
# -
len(styles.baseColour.unique()),styles.baseColour.unique()
# ### We can find that we actually have 43 colors, some of which are very close and difficult to distinguish. We originally had a model that specifically recognizes colors, but the accuracy was very low. So later we decided to extract colors directly from the picture instead of predicting.
# +
import random
from random import randint
import seaborn as sns
df = styles.copy()
df_colors = df.groupby(["baseColour"]).size().reset_index(name="counts").sort_values(by=["counts"], ascending=False)
colors = [f'#{random.randint(0, 0xFFFFFF):06x}' for n in range(df.baseColour.nunique())]
fig, axes = plt.subplots(1, 2, figsize=(15, 10))
sns.barplot(x="baseColour", y="counts", data=df_colors.head(10), palette="hls", ax=axes[0])
axes[1].pie(list(df_colors.counts.values), labels=list(df_colors.baseColour.values), autopct="%1.1f%%", shadow=True, startangle=90, pctdistance=0.85,colors=colors)
center_circle = plt.Circle((0, 0), 0.70, fc='black')
fig = plt.gcf()
fig.gca().add_artist(center_circle)
plt.tight_layout()
plt.show()
# -
plt.figure(figsize=(7,20))
styles.baseColour.value_counts().sort_values().plot(kind='barh')
# ### When we were using the color model, we also found that the model often incorrectly recognizes black color. We think this is due to the fact that there are more black clothes in the data set.
# change the original color column
styles.loc[(styles.baseColour=='Red')|
(styles.baseColour=='Brown')|
(styles.baseColour=='Coffee Brown')|
(styles.baseColour=='Maroon')|
(styles.baseColour=='Rust')|
(styles.baseColour=='Burgundy')|
(styles.baseColour=='Mushroom Brown'),"colorgroup"] = "Red"
styles.loc[(styles.baseColour=='Copper'),"colorgroup"] = "Dark Orange"
styles.loc[(styles.baseColour=='Orange')|
(styles.baseColour=='Bronze')|
(styles.baseColour=='Skin')|
(styles.baseColour=='Nude'),"colorgroup"] = "Orange"
styles.loc[(styles.baseColour=='Gold')|
(styles.baseColour=='Khaki')|
(styles.baseColour=='Beige')|
(styles.baseColour=='Mustard')|
(styles.baseColour=='Tan')|
(styles.baseColour=='Metallic'),"colorgroup"]= "Dark Yellow"
styles.loc[(styles.baseColour=='Yellow'),"colorgroup"] = "Yellow"
styles.loc[(styles.baseColour=='Lime Green'),"colorgroup"]= "Green"
styles.loc[(styles.baseColour=='Green')|
(styles.baseColour=='Sea Green')|
(styles.baseColour=='Fluorescent Green')|
(styles.baseColour=='Olive'),"colorgroup"] = "Dark Green"
styles.loc[(styles.baseColour=='Teal')|
(styles.baseColour=='Turquoise Blue'),"colorgroup"] = "Light Blue"
styles.loc[(styles.baseColour=='Blue'),"colorgroup"]= "Blue"
styles.loc[(styles.baseColour=='Navy Blue'),"colorgroup"] = "Dark Blue"
styles.loc[(styles.baseColour=='Purple')|
(styles.baseColour=='Lavender'),"colorgroup"] = "Purple"
styles.loc[(styles.baseColour=='Pink')|
(styles.baseColour=='Magenta')|
(styles.baseColour=='Peach')|
(styles.baseColour=='Rose')|
(styles.baseColour=='Mauve'),"colorgroup"] = "Pink"
styles.loc[(styles.baseColour=='Black')|
(styles.baseColour=='Charcoal'),"colorgroup"] = "Black"
styles.loc[(styles.baseColour=='White')|
(styles.baseColour=='Off White')|
(styles.baseColour=='Cream'),"colorgroup"] = "White"
styles.loc[(styles.baseColour=='Grey')|
(styles.baseColour=='Silver')|
(styles.baseColour=='Taupe')|
(styles.baseColour=='Grey Melange'),"colorgroup"] = "Grey"
styles.loc[(styles.baseColour=='Multi'),"colorgroup"] = "Multi"
plt.figure(figsize=(7,20))
styles.articleType.value_counts().sort_values().plot(kind='barh')
# ### Similarly, we also found that there are a lot of T-shirts in the data set, which is consistent with the fact that our model often misidentifies to T-shirts
# ### There is another bias here, which may offend users.
plt.figure(figsize=(7,7))
styles.gender.value_counts().sort_values().plot(kind='barh')
# ### We found that sometimes our model would incorrectly identify women clothes as men clothes. Moreover, we have never seen boys and girls and unisex in the recognition results.
|
A special explanation for data cleaning and analysis(bias).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
s = pd.Series(list('asdesdaesdaesdasaeda'))
s
s.unique()
s.value_counts()
dados = pd.read_csv('dados/aluguel.csv', sep = ';')
dados.Tipo.unique()
dados.Tipo.value_counts()
|
Estudos/Python_Data_Science/Pandas/Curso_Pandas/extras/Contadores.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="sTQX9UT37-JK" colab_type="text"
# # Taxon name information
#
# ## Input Name
#
# Enter the taxon name.
# + id="M4SzDNwTCMR_" colab_type="code" cellView="form" colab={}
#@title String fields
taxonNameFull = 'Solanum baretiae' #@param {type:"string"}
taxonName = taxonNameFull.split(" ")
# + [markdown] id="-TvgbfkB2LJm" colab_type="text"
# ## Initialisation
#
# ### Importing Libraries
# + id="3qsD568A7zBO" colab_type="code" outputId="db7b4a88-efa5-4400-8d47-d5e9bcbf0b93" executionInfo={"status": "ok", "timestamp": 1591702063265, "user_tz": -120, "elapsed": 9601, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg1lAOBAHfckOiCDYDPymyViRps7Tf4dAtzyvtEMw=s64", "userId": "15182443477784263859"}} colab={"base_uri": "https://localhost:8080/", "height": 69}
# !pip install -q SPARQLWrapper
# !pip install -q pykew
import requests
import json
import pandas as pd
from rdflib import *
from urllib.error import HTTPError
from google.colab import data_table
from IPython.display import display, Markdown, Latex, Image
import xml.etree.ElementTree as ET
from SPARQLWrapper import SPARQLWrapper, JSON
import warnings
warnings.filterwarnings('ignore') #suppress some warnings coming from RDFLib
import pykew.powo as powo
from pykew.powo_terms import Name
import pykew.ipni as ipni
import matplotlib.pyplot as plt
import numpy as np
import matplotlib.dates as mdates
from datetime import datetime
# + id="iMQQX3mXZu_t" colab_type="code" colab={}
columns = ['scientificName', 'acceptedScientificName', 'recordedBy', 'year', 'eventDate', 'genus', 'species']
#columns = ['scientificName', 'acceptedScientificName']
# + [markdown] id="mz1oO3Os8RMl" colab_type="text"
# ### Defining the APIs
# + id="SOCUPnyk8Uwq" colab_type="code" colab={}
gbif_base_url = 'https://api.gbif.org/v1/'
gbif_occurence = gbif_base_url + 'occurrence/search'
gbif_species = gbif_base_url + 'species/match' # to retrieve the taxonKey
plazi_base_url = 'http://tb.plazi.org/GgServer/rdf/'
# + [markdown] id="xTKbr6EZSPDg" colab_type="text"
# ### Defining functions
# + id="Dfy3O2ujSTR2" colab_type="code" colab={}
def get_types(key,type_status):
type_search = {'taxonKey' : key, 'typeStatus' : type_status, 'limit' : 20}
type_request = requests.get(gbif_occurence, params=type_search)
if type_request.status_code == 200:
types = json.loads(type_request.text)
endOfRecords = types['endOfRecords']
df = pd.DataFrame(types['results'])
iteration = 1
while not endOfRecords:
type_search = {'taxonKey' : key, 'typeStatus' : type_status, 'limit' : 20, 'offset' : iteration*20}
type_request = requests.get(gbif_occurence, params=type_search)
types = json.loads(type_request.text)
endOfRecords = types['endOfRecords']
dfe = pd.DataFrame(types['results'])
df = df.append(dfe, ignore_index=True)
iteration += 1
else:
print('Something went wrong')
return df
# + [markdown] id="qpaLQNw0V5wn" colab_type="text"
# ## Get synonyms from POWO
# + id="3tkrXbmFIhEo" colab_type="code" outputId="d7e310b5-97ae-4248-99c2-7fab7b9575c3" executionInfo={"status": "ok", "timestamp": 1591702065064, "user_tz": -120, "elapsed": 11370, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg1lAOBAHfckOiCDYDPymyViRps7Tf4dAtzyvtEMw=s64", "userId": "15182443477784263859"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
p_query = {Name.genus : taxonName[0], Name.species : taxonName[1]}
name = []
synonyms = []
results = ipni.search(p_query)
spec = []
for r in results:
if r['rank'] == 'spec.':
spec.append(r)
for s in spec:
powo_result = powo.lookup(s['fqId'])
try:
for synonym in powo_result['synonyms']:
synonyms.append(synonym)
for synonym in synonyms:
print(synonym['name'])
except KeyError:
print('No synonyms found')
# + [markdown] id="A7ii1duaL5oW" colab_type="text"
# ## Get type specimen from GBIF
#
# ### Get the taxon key
# + id="b92i4tS4L-VR" colab_type="code" colab={}
taxonKey_search = {'name' : taxonNameFull, 'strict' : True, 'verbose' : True}
request = requests.get(gbif_species, params=taxonKey_search)
if request.status_code == 200:
req_json = json.loads(request.text)
if req_json['matchType'] == 'NONE':
print('Species not found, try another one')
assert(False)
taxonKey = req_json['usageKey']
speciesKey = req_json['speciesKey']
genusKey = req_json['genusKey']
genus = req_json['genus']
species = req_json['species'].split(' ')[1]
else:
print('Something went wrong, try different Scientific Name')
assert(False)
# + [markdown] id="ukl_AsrRM83y" colab_type="text"
# ### Look for type specimen in GBIF
# + id="b6el3mf1NCM4" colab_type="code" outputId="1e5beba7-8693-4d8b-e6e7-89b3093a817d" executionInfo={"status": "ok", "timestamp": 1591703163385, "user_tz": -120, "elapsed": 7951, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg1lAOBAHfckOiCDYDPymyViRps7Tf4dAtzyvtEMw=s64", "userId": "15182443477784263859"}} colab={"base_uri": "https://localhost:8080/", "height": 1585}
types_list = ['Holotype', 'Isotype', 'Paratype', 'Syntype', 'Lectotype', 'Isolectotype', 'Neotype', 'Isoneotype', 'Epitype', 'Type']
for type_name in types_list:
print('List of ' + type_name)
print('--------------------')
types = get_types(taxonKey, type_name)
if len(types) > 0:
adjusted_columns = []
for item in columns:
if item in types.columns:
adjusted_columns.append(item)
types1 = types[adjusted_columns]
display(data_table.DataTable(types1, include_index=False, num_rows_per_page=20))
try:
if len(types) > 0:
#display(data_table.DataTable(types[columns], include_index=False, num_rows_per_page=20))
# create a timeline
# first drop the nan in eventdate
types = types[types['eventDate'].notna()]
names = types['scientificName'].tolist()
dates = [datetime.strptime(d, "%Y-%m-%dT%H:%M:%S") for d in types.eventDate]
# Choose some nice levels
levels = np.tile([-5, 5, -3, 3, -1, 1],
int(np.ceil(len(dates)/6)))[:len(dates)]
# Create figure and plot a stem plot with the date
fig, ax = plt.subplots(figsize=(8.8, 4), constrained_layout=True)
ax.set(title="Specimen eventDates")
markerline, stemline, baseline = ax.stem(dates, levels,
linefmt="C3-", basefmt="k-",
use_line_collection=True)
plt.setp(markerline, mec="k", mfc="w", zorder=3)
# Shift the markers to the baseline by replacing the y-data by zeros.
markerline.set_ydata(np.zeros(len(dates)))
# annotate lines
vert = np.array(['top', 'bottom'])[(levels > 0).astype(int)]
for d, l, r, va in zip(dates, levels, names, vert):
ax.annotate(r, xy=(d, l), xytext=(-3, np.sign(l)*3),
textcoords="offset points", va=va, ha="right")
# format xaxis with 62 month intervals
ax.get_xaxis().set_major_locator(mdates.MonthLocator(interval=63))
ax.get_xaxis().set_major_formatter(mdates.DateFormatter("%b %Y"))
plt.setp(ax.get_xticklabels(), rotation=30, ha="right")
# remove y axis and spines
ax.get_yaxis().set_visible(False)
for spine in ["left", "top", "right"]:
ax.spines[spine].set_visible(False)
ax.margins(y=0.1)
plt.show()
else:
print('No ' + type_name + ' found')
except KeyError:
print('KeyError')
pass
print('\n \n')
# + [markdown] id="CyyhG8XEXUWF" colab_type="text"
# ### Get taxonomic treatments
# + id="fo5otPzejLfQ" colab_type="code" outputId="89c5761d-d514-4300-a9bf-4be0b4b5a30a" executionInfo={"status": "ok", "timestamp": 1591703198308, "user_tz": -120, "elapsed": 2001, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg1lAOBAHfckOiCDYDPymyViRps7Tf4dAtzyvtEMw=s64", "userId": "15182443477784263859"}} colab={"base_uri": "https://localhost:8080/", "height": 35}
plazi_sparql = 'https://treatment.ld.plazi.org/sparql'
p_query = """
PREFIX rdf: <http://www.w3.org/1999/02/22-rdf-syntax-ns#>
PREFIX dwc: <http://rs.tdwg.org/dwc/terms/>
PREFIX treat: <http://plazi.org/vocab/treatment#>
PREFIX dc: <http://purl.org/dc/elements/1.1/>
SELECT * WHERE {{
?tc dwc:genus "{0}" .
?tc dwc:species "{1}" .
?tc a <http://filteredpush.org/ontologies/oa/dwcFP#TaxonConcept> .
OPTIONAL {{ ?tc treat:hasTaxonName ?tn . }}
OPTIONAL {{ ?augmentingTreatment treat:augmentsTaxonConcept ?tc .
?augmentingTreatment dc:creator ?augmentingTreatmentCreator .}}
OPTIONAL {{ ?definingTreatment treat:definesTaxonConcept ?tc .
?definingTreatment dc:creator ?definingTreatmentCreator .}}
}}
""".format(genus,species)
treatments = []
sparql = SPARQLWrapper(plazi_sparql)
sparql.setQuery(p_query)
sparql.setReturnFormat(JSON)
results = sparql.query().convert()
for result in results["results"]["bindings"]:
try:
treatments.append(result['definingTreatment']['value'])
except KeyError:
pass
try:
treatments.append(result['augmentingTreatment']['value'])
except KeyError:
pass
treatments = list(set(treatments))
print(treatments)
|
notebooks/Name_information_types.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# STAT 453: Deep Learning (Spring 2021)
# Instructor: <NAME> (<EMAIL>)
#
# Course website: http://pages.stat.wisc.edu/~sraschka/teaching/stat453-ss2021/
# GitHub repository: https://github.com/rasbt/stat453-deep-learning-ss21
#
# ---
# %load_ext watermark
# %watermark -a '<NAME>' -v -p torch
# # MLP with Dropout
# ## Imports
import torch
import numpy as np
import matplotlib.pyplot as plt
# From local helper files
from helper_evaluation import set_all_seeds, set_deterministic
from helper_train import train_model
from helper_plotting import plot_training_loss, plot_accuracy, show_examples
from helper_dataset import get_dataloaders_mnist
# ## Settings and Dataset
# +
##########################
### SETTINGS
##########################
RANDOM_SEED = 123
BATCH_SIZE = 256
NUM_HIDDEN_1 = 75
NUM_HIDDEN_2 = 45
NUM_EPOCHS = 50
DEVICE = torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')
# -
set_all_seeds(RANDOM_SEED)
set_deterministic()
# +
##########################
### MNIST DATASET
##########################
train_loader, valid_loader, test_loader = get_dataloaders_mnist(
batch_size=BATCH_SIZE,
validation_fraction=0.1)
# Checking the dataset
for images, labels in train_loader:
print('Image batch dimensions:', images.shape)
print('Image label dimensions:', labels.shape)
print('Class labels of 10 examples:', labels[:10])
break
# -
# ## Model
class MultilayerPerceptron(torch.nn.Module):
def __init__(self, num_features, num_classes, drop_proba,
num_hidden_1, num_hidden_2):
super().__init__()
self.my_network = torch.nn.Sequential(
# 1st hidden layer
torch.nn.Flatten(),
torch.nn.Linear(num_features, num_hidden_1, bias=False),
torch.nn.BatchNorm1d(num_hidden_1),
torch.nn.ReLU(),
# 2nd hidden layer
torch.nn.Linear(num_hidden_1, num_hidden_2, bias=False),
torch.nn.BatchNorm1d(num_hidden_2),
torch.nn.ReLU(),
# output layer
torch.nn.Linear(num_hidden_2, num_classes)
)
def forward(self, x):
logits = self.my_network(x)
return logits
# +
torch.manual_seed(RANDOM_SEED)
model = MultilayerPerceptron(num_features=28*28,
num_hidden_1=NUM_HIDDEN_1,
num_hidden_2=NUM_HIDDEN_2,
drop_proba=0.5,
num_classes=10)
model = model.to(DEVICE)
optimizer = torch.optim.SGD(model.parameters(), lr=0.1)
minibatch_loss_list, train_acc_list, valid_acc_list = train_model(
model=model,
num_epochs=NUM_EPOCHS,
train_loader=train_loader,
valid_loader=valid_loader,
test_loader=test_loader,
optimizer=optimizer,
device=DEVICE)
plot_training_loss(minibatch_loss_list=minibatch_loss_list,
num_epochs=NUM_EPOCHS,
iter_per_epoch=len(train_loader),
results_dir=None,
averaging_iterations=20)
plt.show()
plot_accuracy(train_acc_list=train_acc_list,
valid_acc_list=valid_acc_list,
results_dir=None)
plt.ylim([80, 100])
plt.show()
|
L11/code/batchnorm.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import seaborn as sns
import scipy.io as sio
import matplotlib.pyplot as plt
sns.set_style('whitegrid', {'xtick.color': '.15',
'xtick.direction': u'in',
'xtick.major.size': 10,
'xtick.minor.size': 5,
'ytick.color': '.15',
'ytick.direction': u'in',
'ytick.major.size': 10,
'ytick.minor.size': 5,
'grid.color': '.8',
'axes.edgecolor': '.4',
'grid.linestyle': '',
'axes.linewidth': 1.0,
'grid.linewidth': 0.5})
plt.rc('font', size=20)
paired = sns.color_palette("Paired", 10)
plt.rcParams['ytick.right'] = plt.rcParams['ytick.labelright'] = True
plt.rcParams['ytick.left'] = plt.rcParams['ytick.labelleft'] = False
# +
f = plt.figure()
f.set_size_inches(12,24)
plt.rc('axes', axisbelow=False)
plt.subplots_adjust(hspace=0)
plt.subplots_adjust(wspace=0)
dat = sio.loadmat('flexure_profiles.mat')
ax1 = plt.subplot2grid((5,1),(0,0),rowspan=1,colspan=1)
ax1.axhline(0,color=[0.5,0.5,0.5],linewidth=1)
ax1.plot(dat['lat_T0653_gt3L'],dat['z_wv_T0653_gt3L'],color=paired[4],linewidth=2) #Worldview Profile
ax1.plot(dat['lat_T0653_gt3L'],dat['z_profile_T0653_gt3L'],color='k',linewidth=2) #Flexure Model
ax1.plot(dat['lat_is2_0653_1l'],dat['z_is2_0653_1l'],color='#7B2281',linewidth=2) #IS2 Profile
ax1.xaxis.set_major_formatter(plt.NullFormatter())
ax1.set_ylim([-3,13])
ax1.set_xlim([-72.5,-72.1])
ax1.set_ylabel('$\Delta$ h (m)')
ax1.set_xlim([-72.475,-72.2])
ax1.yaxis.set_label_position("right")
ax1.set_xticks([-72.45,-72.4,-72.35,-72.3,-72.25,-72.2])
ax1 = plt.subplot2grid((5,1),(1,0),rowspan=1,colspan=1)
ax1.axhline(0,color=[0.5,0.5,0.5],linewidth=1)
ax1.plot(dat['lat_T0081_gt1L'],dat['z_wv_T0081_gt1L'],color=paired[4],linewidth=2)
ax1.plot(dat['lat_T0081_gt1L'],dat['z_profile_T0081_gt1L'],color='k',linewidth=2)
ax1.plot(dat['lat_T0081_gt1L'],dat['z_uplift_T0081_gt1L'],color='k',linewidth=2,linestyle='--')
ax1.plot(dat['lat_is2_0081_1l'],dat['z_is2_0081_1l'],color='#C96C29',linewidth=3)
ax1.xaxis.set_major_formatter(plt.NullFormatter())
ax1.set_ylim([-40,40])
ax1.set_yticks([-30,-15,0,15,30])
ax1.set_ylabel('$\Delta$ h (m)')
ax1.set_xlim([-72.475,-72.2])
ax1.yaxis.set_label_position("right")
ax1.set_xticks([-72.45,-72.4,-72.35,-72.3,-72.25,-72.2])
ax1 = plt.subplot2grid((5,1),(2,0),rowspan=1,colspan=1)
ax1.axhline(0,color=[0.5,0.5,0.5],linewidth=1)
ax1.plot(dat['lat_T1095_gt1L'],dat['z_wv_T1095_gt1L'],color=paired[4],linewidth=2)
ax1.plot(dat['lat_T1095_gt1L'],dat['z_profile_T1095_gt1L'],color='k',linewidth=2)
ax1.plot(dat['lat_T1095_gt1L'],dat['z_uplift_T1095_gt1L'],color='k',linewidth=2,linestyle='--')
ax1.plot(dat['lat_is2_1095_1l'],dat['z_is2_1095_1l'],color='#418869',linewidth=3)
ax1.xaxis.set_major_formatter(plt.NullFormatter())
ax1.set_ylim([-40,40])
ax1.set_yticks([-30,-15,0,15,30])
ax1.set_ylabel('$\Delta$ h (m)')
ax1.set_xlim([-72.475,-72.2])
ax1.yaxis.set_label_position("right")
ax1.set_xticks([-72.45,-72.4,-72.35,-72.3,-72.25,-72.2])
ax1 = plt.subplot2grid((5,1),(3,0),rowspan=1,colspan=1)
ax1.axhline(0,color=[0.5,0.5,0.5],linewidth=1)
ax1.plot(dat['lat_T1095_gt1R'],dat['z_wv_T1095_gt1R'],color=paired[4],linewidth=2)
ax1.plot(dat['lat_T1095_gt1R'],dat['z_profile_T1095_gt1R'],color='k',linewidth=2)
ax1.plot(dat['lat_T1095_gt1R'],dat['z_uplift_T1095_gt1R'],color='k',linewidth=2,linestyle='--')
ax1.plot(dat['lat_is2_1095_1r'],dat['z_is2_1095_1r'],color='#418869',linewidth=3)
ax1.xaxis.set_major_formatter(plt.NullFormatter())
ax1.set_ylim([-40,40])
ax1.set_yticks([-30,-15,0,15,30])
ax1.set_ylabel('$\Delta$ h (m)')
ax1.set_xlim([-72.475,-72.2])
ax1.yaxis.set_label_position("right")
ax1.set_xticks([-72.45,-72.4,-72.35,-72.3,-72.25,-72.2])
ax1 = plt.subplot2grid((5,1),(4,0),rowspan=1,colspan=1)
ax1.axhline(0,color=[0.5,0.5,0.5],linewidth=1)
ax1.plot(dat['lat_T1095_gt2L'],dat['z_wv_T1095_gt2L'],color=paired[4],linewidth=2)
ax1.plot(dat['lat_T1095_gt2L'],dat['z_profile_T1095_gt2L'],color='k',linewidth=2)
ax1.plot(dat['lat_is2_1095_2l'],dat['z_is2_1095_2l'],color='#418869',linewidth=3)
plt.xlabel('Latitude')
ax1.set_ylim([-5,25])
ax1.set_yticks([0,10,20])
ax1.set_ylabel('$\Delta$ h (m)')
ax1.set_xlim([-72.475,-72.2])
ax1.yaxis.set_label_position("right")
ax1.set_xticks([-72.45,-72.4,-72.35,-72.3,-72.25,-72.2])
plt.savefig('figure_2.pdf',bbox_inches='tight')
|
Fig2_height_change_profiles.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # CVE.ICU - 2021
# ---
# This information from this website is from a [jupyter notebook](https://jupyter.org/) that automatically pulls all [JSON Data](https://nvd.nist.gov/vuln/data-feeds#JSON_FEED) from the NVD and perform some fundamental data analysis and graphing. If you have any questions or suggestions, please join the [discussion here](https://github.com/jgamblin/cve.icu/discussions) and check out the [source code](https://github.com/jgamblin/cve.icu/). Questions? Reach out to [@jgamblin](https://www.twitter.com/jgamblin).
#
# **Yearly Data**
# - [All](https://cve.icu/)
# - [2022](2022.html)
# - [2021](2021.html)
# - [2020](2020.html)
# - [2019](2019.html)
#
# **Other Data**
# - [CVE Growth Predictions](prophet.html)
# - [CVE Calendar All Years](calendar.html)
# - [CNA Map](cnamap.html)
# ## CVE Data
# + tags=[]
from IPython.core.magic import register_cell_magic
from IPython.display import Markdown
import calplot
import datetime
from datetime import date
import glob
import json
import logging
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import warnings
@register_cell_magic
def markdown(line, cell):
return Markdown(cell.format(**globals()))
logging.getLogger('matplotlib.font_manager').disabled = True
warnings.filterwarnings("ignore")
pd.set_option('display.width', 500)
pd.set_option('display.max_rows', 50)
pd.set_option('display.max_columns', 10)
# + tags=[]
row_accumulator = []
for filename in glob.glob('nvdcve-1.1-*.json'):
with open(filename, 'r', encoding='utf-8') as f:
nvd_data = json.load(f)
for entry in nvd_data['CVE_Items']:
cve = entry['cve']['CVE_data_meta']['ID']
try:
assigner = entry['cve']['CVE_data_meta']['ASSIGNER']
except KeyError:
published_date = 'Missing_Data'
try:
published_date = entry['publishedDate']
except KeyError:
published_date = 'Missing_Data'
try:
attack_vector = entry['impact']['baseMetricV3']['cvssV3']['attackVector']
except KeyError:
attack_vector = 'Missing_Data'
try:
attack_complexity = entry['impact']['baseMetricV3']['cvssV3']['attackComplexity']
except KeyError:
attack_complexity = 'Missing_Data'
try:
privileges_required = entry['impact']['baseMetricV3']['cvssV3']['privilegesRequired']
except KeyError:
privileges_required = 'Missing_Data'
try:
user_interaction = entry['impact']['baseMetricV3']['cvssV3']['userInteraction']
except KeyError:
user_interaction = 'Missing_Data'
try:
scope = entry['impact']['baseMetricV3']['cvssV3']['scope']
except KeyError:
scope = 'Missing_Data'
try:
confidentiality_impact = entry['impact']['baseMetricV3']['cvssV3']['confidentialityImpact']
except KeyError:
confidentiality_impact = 'Missing_Data'
try:
integrity_impact = entry['impact']['baseMetricV3']['cvssV3']['integrityImpact']
except KeyError:
integrity_impact = 'Missing_Data'
try:
availability_impact = entry['impact']['baseMetricV3']['cvssV3']['availabilityImpact']
except KeyError:
availability_impact = 'Missing_Data'
try:
base_score = entry['impact']['baseMetricV3']['cvssV3']['baseScore']
except KeyError:
base_score = '0.0'
try:
base_severity = entry['impact']['baseMetricV3']['cvssV3']['baseSeverity']
except KeyError:
base_severity = 'Missing_Data'
try:
exploitability_score = entry['impact']['baseMetricV3']['exploitabilityScore']
except KeyError:
exploitability_score = 'Missing_Data'
try:
impact_score = entry['impact']['baseMetricV3']['impactScore']
except KeyError:
impact_score = 'Missing_Data'
try:
cwe = entry['cve']['problemtype']['problemtype_data'][0]['description'][0]['value']
except IndexError:
cwe = 'Missing_Data'
try:
description = entry['cve']['description']['description_data'][0]['value']
except IndexError:
description = ''
new_row = {
'CVE': cve,
'Published': published_date,
'AttackVector': attack_vector,
'AttackComplexity': attack_complexity,
'PrivilegesRequired': privileges_required,
'UserInteraction': user_interaction,
'Scope': scope,
'ConfidentialityImpact': confidentiality_impact,
'IntegrityImpact': integrity_impact,
'AvailabilityImpact': availability_impact,
'BaseScore': base_score,
'BaseSeverity': base_severity,
'ExploitabilityScore': exploitability_score,
'ImpactScore': impact_score,
'CWE': cwe,
'Description': description,
'Assigner' : assigner
}
if not description.startswith('** REJECT **'): # disputed, rejected and other non issues start with '**'
row_accumulator.append(new_row)
nvd = pd.DataFrame(row_accumulator)
nvd['Published'] = pd.to_datetime(nvd['Published'])
thisyear = ((nvd['Published'] > '2021-01-01') & (nvd['Published'] < '2021-12-31'))
nvd = nvd.loc[thisyear]
nvd = nvd.sort_values(by=['Published'])
nvd = nvd.reset_index(drop=True)
nvd['BaseScore'] = pd.to_numeric(nvd['BaseScore']);
nvd['BaseScore'] = pd.to_numeric(nvd['BaseScore']);
nvd['BaseScore'] = nvd['BaseScore'].replace(0, np.NaN);
nvdcount = nvd['Published'].count()
nvdunique = nvd['Published'].nunique()
startdate = date(2021, 1, 1)
enddate = date(2021, 12, 31)
numberofdays = enddate - startdate
per_day = nvdcount/numberofdays.days
# -
Markdown(f"Total Number of CVEs: **{nvd['CVE'].count()}**<br />Average CVEs Per Day: **{per_day.round(2)}**<br />Average CVSS Score: **{nvd['BaseScore'].mean().round(2)}**")
Markdown(f"## CVE Graphs")
Month_Graph = nvd['Published'].groupby(nvd.Published.dt.to_period("M")).agg('count')
Year_Graph = nvd['Published'].groupby(nvd.Published.dt.to_period("Y")).agg('count')
Week_Graph = nvd['Published'].groupby(nvd.Published.dt.to_period("W")).agg('count')
Day_Graph = nvd['Published'].groupby(nvd.Published.dt.to_period("D")).agg('count')
Markdown(f"### CVE Calendar")
dfs = nvd['Published'].apply(lambda x: pd.to_datetime(x, errors='coerce', format='%Y/%m/%d'))
df = dfs.value_counts()
df = df.to_frame()
df.index = df.index.strftime('%m/%d/%Y')
df.index = pd.to_datetime(df.index, format='%m/%d/%Y')
calplot.calplot(df.T.squeeze(), cmap='jet', dropzero=True, edgecolor="Grey", textcolor="White", textformat='{:.0f}', textfiller='', suptitle='CVEs Per Day', figsize=(25,3));
Markdown(f"### CVE Per Month Graph")
# +
cg = Month_Graph.plot.area(colormap='jet', figsize=(16, 8), title='CVEs Per Month')
plt.grid()
cg.set_ylabel("New CVEs");
cg.set_xlabel("Date");
# -
Markdown(f"### CVE Per Week Graph")
# +
cg = Week_Graph.plot.area(colormap='jet', figsize=(16, 8), title='CVEs Per Week')
plt.grid()
cg.set_ylabel("New CVEs");
cg.set_xlabel("Date");
# -
Markdown(f"### CVE Per Day Graph")
# +
cg = Day_Graph.plot.area(colormap='jet', figsize=(16, 8), title='CVEs Per Day')
plt.grid()
cg.set_ylabel("New CVEs");
cg.set_xlabel("Date");
# -
Markdown(f"## CVSS Data")
nvd['BaseScore'].plot(kind="hist", colormap='jet', figsize=(16, 8), title='CVSS Scores');
Markdown(f"## CNA Data")
Markdown(f"### CNA Assigner Graph")
nvd_frequency = nvd['Assigner'].value_counts()
nvd_frequency = nvd_frequency.reset_index()
nvd_frequency.columns = ['Assigner', 'counts']
nvd_frequency[nvd_frequency.counts > 100].head(50)
nvd_frequency_no_mitre = nvd_frequency[~nvd_frequency.Assigner.str.contains('<EMAIL>')]
nvd_frequency_no_mitre = nvd_frequency_no_mitre[nvd_frequency_no_mitre.counts > 100].head(20)
plt.figure(figsize=(10,10))
plt.barh("Assigner", "counts", data = nvd_frequency_no_mitre, color="#001d82")
plt.xlabel("CVEs");
plt.ylabel("") ;
plt.title("Top 20 CNAs");
Markdown(f"## CWE Data")
nvd_cwe = nvd['CWE'].value_counts()
nvd_cwe = nvd_cwe.reset_index()
nvd_cwe.columns = ['CWE', 'counts']
nvd_cwe = nvd_cwe[~nvd_cwe.CWE.str.contains('Missing_')]
nvd_cwe = nvd_cwe[~nvd_cwe.CWE.str.contains('NVD')]
nvd_cwe = nvd_cwe[nvd_cwe.counts > 100].head(25)
plt.figure(figsize=(10,10));
plt.barh("CWE", "counts", data = nvd_cwe, color="#001d82");
plt.xlabel("Count");
plt.ylabel("CWE") ;
plt.title("Most Common CWE in CVE Records");
Markdown(f"## More CVE Data")
Markdown(f"### Top CNA Assigner")
nvd_frequency_no_mitre.style.hide_index()
Markdown(f"### CVEs By Identifier")
# +
print("CVE-1999:\t%s" % len(nvd[nvd['CVE'].str.contains('CVE-1999-')]))
print("CVE-2000:\t%s" % len(nvd[nvd['CVE'].str.contains('CVE-2000-')]))
print("CVE-2001:\t%s" % len(nvd[nvd['CVE'].str.contains('CVE-2001-')]))
print("CVE-2002:\t%s" % len(nvd[nvd['CVE'].str.contains('CVE-2002-')]))
print("CVE-2003:\t%s" % len(nvd[nvd['CVE'].str.contains('CVE-2003-')]))
print("CVE-2004:\t%s" % len(nvd[nvd['CVE'].str.contains('CVE-2004-')]))
print("CVE-2005:\t%s" % len(nvd[nvd['CVE'].str.contains('CVE-2005-')]))
print("CVE-2006:\t%s" % len(nvd[nvd['CVE'].str.contains('CVE-2006-')]))
print("CVE-2007:\t%s" % len(nvd[nvd['CVE'].str.contains('CVE-2007-')]))
print("CVE-2008:\t%s" % len(nvd[nvd['CVE'].str.contains('CVE-2008-')]))
print("CVE-2009:\t%s" % len(nvd[nvd['CVE'].str.contains('CVE-2009-')]))
print("CVE-2010:\t%s" % len(nvd[nvd['CVE'].str.contains('CVE-2010-')]))
print("CVE-2011:\t%s" % len(nvd[nvd['CVE'].str.contains('CVE-2011-')]))
print("CVE-2012:\t%s" % len(nvd[nvd['CVE'].str.contains('CVE-2012-')]))
print("CVE-2013:\t%s" % len(nvd[nvd['CVE'].str.contains('CVE-2013-')]))
print("CVE-2014:\t%s" % len(nvd[nvd['CVE'].str.contains('CVE-2014-')]))
print("CVE-2015:\t%s" % len(nvd[nvd['CVE'].str.contains('CVE-2015-')]))
print("CVE-2016:\t%s" % len(nvd[nvd['CVE'].str.contains('CVE-2016-')]))
print("CVE-2017:\t%s" % len(nvd[nvd['CVE'].str.contains('CVE-2017-')]))
print("CVE-2018:\t%s" % len(nvd[nvd['CVE'].str.contains('CVE-2018-')]))
print("CVE-2019:\t%s" % len(nvd[nvd['CVE'].str.contains('CVE-2019-')]))
print("CVE-2020:\t%s" % len(nvd[nvd['CVE'].str.contains('CVE-2020-')]))
print("CVE-2021:\t%s" % len(nvd[nvd['CVE'].str.contains('CVE-2021-')]))
print("CVE-2022:\t%s" % len(nvd[nvd['CVE'].str.contains('CVE-2022-')]))
# -
Markdown(f"This report is updated automatically every day, last generated on: **{datetime.datetime.now()}**")
|
cveicu2021.nbconvert.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + [markdown] origin_pos=0
# # 深度卷积神经网络(AlexNet)
# :label:`sec_alexnet`
#
# 在LeNet提出后,卷积神经网络在计算机视觉和机器学习领域中很有名气。但卷积神经网络并没有主导这些领域。这是因为虽然LeNet在小数据集上取得了很好的效果,但是在更大、更真实的数据集上训练卷积神经网络的性能和可行性还有待研究。事实上,在上世纪90年代初到2012年之间的大部分时间里,神经网络往往被其他机器学习方法超越,如支持向量机(support vector machines)。
#
# 在计算机视觉中,直接将神经网络与其他机器学习方法进行比较也许不公平。这是因为,卷积神经网络的输入是由原始像素值或是经过简单预处理(例如居中、缩放)的像素值组成的。但在使用传统机器学习方法时,从业者永远不会将原始像素作为输入。在传统机器学习方法中,计算机视觉流水线是由经过人的手工精心设计的特征流水线组成的。对于这些传统方法,大部分的进展都来自于对特征有了更聪明的想法,并且学习到的算法往往归于事后的解释。
#
# 虽然上世纪90年代就有了一些神经网络加速卡,但仅靠它们还不足以开发出有大量参数的深层多通道多层卷积神经网络。此外,当时的数据集仍然相对较小。除了这些障碍,训练神经网络的一些关键技巧仍然缺失,包括启发式参数初始化、随机梯度下降的变体、非挤压激活函数和有效的正则化技术。
#
# 因此,与训练*端到端*(从像素到分类结果)系统不同,经典机器学习的流水线看起来更像下面这样:
#
# 1. 获取一个有趣的数据集。在早期,收集这些数据集需要昂贵的传感器(在当时最先进的图像也就100万像素)。
# 2. 根据光学、几何学、其他知识以及偶然的发现,手工对特征数据集进行预处理。
# 3. 通过标准的特征提取算法,如SIFT(尺度不变特征变换) :cite:`Lowe.2004`和SURF(加速鲁棒特征) :cite:`Bay.Tuytelaars.Van-Gool.2006`或其他手动调整的流水线来输入数据。
# 4. 将提取的特征送入最喜欢的分类器中(例如线性模型或其它核方法),以训练分类器。
#
# 如果你和机器学习研究人员交谈,你会发现他们相信机器学习既重要又美丽:优雅的理论去证明各种模型的性质。机器学习是一个正在蓬勃发展、严谨且非常有用的领域。然而,如果你和计算机视觉研究人员交谈,你会听到一个完全不同的故事。他们会告诉你图像识别的诡异事实————推动领域进步的是数据特征,而不是学习算法。计算机视觉研究人员相信,从对最终模型精度的影响来说,更大或更干净的数据集、或是稍微改进的特征提取,比任何学习算法带来的进步要大得多。
#
# ## 学习表征
#
# 另一种预测这个领域发展的方法————观察图像特征的提取方法。在2012年前,图像特征都是机械地计算出来的。事实上,设计一套新的特征函数、改进结果,并撰写论文是盛极一时的潮流。SIFT :cite:`Lowe.2004`、SURF :cite:`Bay.Tuytelaars.Van-Gool.2006`、HOG(定向梯度直方图) :cite:`Dalal.Triggs.2005`、[bags of visual words](https://en.wikipedia.org/wiki/Bag-of-words_model_in_computer_vision)和类似的特征提取方法占据了主导地位。
#
# 另一组研究人员,包括<NAME>、<NAME>、<NAME>、<NAME>、<NAME>和<NAME>,想法则与众不同:他们认为特征本身应该被学习。此外,他们还认为,在合理地复杂性前提下,特征应该由多个共同学习的神经网络层组成,每个层都有可学习的参数。在机器视觉中,最底层可能检测边缘、颜色和纹理。事实上,<NAME>、<NAME>和<NAME>提出了一种新的卷积神经网络变体*AlexNet*。在2012年ImageNet挑战赛中取得了轰动一时的成绩。AlexNet以<NAME>的名字命名,他是论文 :cite:`Krizhevsky.Sutskever.Hinton.2012`的第一作者。
#
# 有趣的是,在网络的最底层,模型学习到了一些类似于传统滤波器的特征抽取器。 :numref:`fig_filters`是从AlexNet论文 :cite:`Krizhevsky.Sutskever.Hinton.2012`复制的,描述了底层图像特征。
#
# 
# :width:`400px`
# :label:`fig_filters`
#
# AlexNet的更高层建立在这些底层表示的基础上,以表示更大的特征,如眼睛、鼻子、草叶等等。而更高的层可以检测整个物体,如人、飞机、狗或飞盘。最终的隐藏神经元可以学习图像的综合表示,从而使属于不同类别的数据易于区分。尽管一直有一群执着的研究者不断钻研,试图学习视觉数据的逐级表征,然而很长一段时间里这些尝试都未有突破。深度卷积神经网络的突破出现在2012年。突破可归因于两个关键因素。
#
# ### 缺少的成分:数据
#
# 包含许多特征的深度模型需要大量的有标签数据,才能显著优于基于凸优化的传统方法(如线性方法和核方法)。
# 然而,限于早期计算机有限的存储和90年代有限的研究预算,大部分研究只基于小的公开数据集。例如,不少研究论文基于加州大学欧文分校(UCI)提供的若干个公开数据集,其中许多数据集只有几百至几千张在非自然环境下以低分辨率拍摄的图像。这一状况在2010年前后兴起的大数据浪潮中得到改善。2009年,ImageNet数据集发布,并发起ImageNet挑战赛:要求研究人员从100万个样本中训练模型,以区分1000个不同类别的对象。ImageNet数据集由斯坦福教授李飞飞小组的研究人员开发,利用谷歌图像搜索(Google Image Search)对每一类图像进行预筛选,并利用亚马逊众包(Amazon Mechanical Turk)来标注每张图片的相关类别。这种规模是前所未有的。这项被称为ImageNet的挑战赛推动了计算机视觉和机器学习研究的发展,挑战研究人员确定哪些模型能够在更大的数据规模下表现最好。
#
# ### 缺少的成分:硬件
#
# 深度学习对计算资源要求很高,训练可能需要数百个迭代轮数,每次迭代都需要通过代价高昂的许多线性代数层传递数据。这也是为什么在20世纪90年代至21世纪初,优化凸目标的简单算法是研究人员的首选。然而,用GPU训练神经网络改变了这一格局。*图形处理器*(Graphics Processing Unit,GPU)早年用来加速图形处理,使电脑游戏玩家受益。GPU可优化高吞吐量的$4 \times 4$矩阵和向量乘法,从而服务于基本的图形任务。幸运的是,这些数学运算与卷积层的计算惊人地相似。由此,英伟达(NVIDIA)和ATI已经开始为通用计算操作优化gpu,甚至把它们作为*通用GPU*(general-purpose GPUs,GPGPU)来销售。
#
# 那么GPU比CPU强在哪里呢?
#
# 首先,我们深度理解一下中央处理器(Central Processing Unit,CPU)的*核心*。
# CPU的每个核心都拥有高时钟频率的运行能力,和高达数MB的三级缓存(L3Cache)。
# 它们非常适合执行各种指令,具有分支预测器、深层流水线和其他使CPU能够运行各种程序的功能。
# 然而,这种明显的优势也是它的致命弱点:通用核心的制造成本非常高。
# 它们需要大量的芯片面积、复杂的支持结构(内存接口、内核之间的缓存逻辑、高速互连等等),而且它们在任何单个任务上的性能都相对较差。
# 现代笔记本电脑最多有4核,即使是高端服务器也很少超过64核,因为它们的性价比不高。
#
# 相比于CPU,GPU由$100 \sim 1000$个小的处理单元组成(NVIDIA、ATI、ARM和其他芯片供应商之间的细节稍有不同),通常被分成更大的组(NVIDIA称之为warps)。
# 虽然每个GPU核心都相对较弱,有时甚至以低于1GHz的时钟频率运行,但庞大的核心数量使GPU比CPU快几个数量级。
# 例如,NVIDIA最近一代的Ampere GPU架构为每个芯片提供了高达312 TFlops的浮点性能,而CPU的浮点性能到目前为止还没有超过1 TFlops。
# 之所以有如此大的差距,原因其实很简单:首先,功耗往往会随时钟频率呈二次方增长。
# 对于一个CPU核心,假设它的运行速度比GPU快4倍,你可以使用16个GPU内核取代,那么GPU的综合性能就是CPU的$16 \times 1/4 = 4$倍。
# 其次,GPU内核要简单得多,这使得它们更节能。
# 此外,深度学习中的许多操作需要相对较高的内存带宽,而GPU拥有10倍于CPU的带宽。
#
# 回到2012年的重大突破,当<NAME>和<NAME>实现了可以在GPU硬件上运行的深度卷积神经网络时,一个重大突破出现了。他们意识到卷积神经网络中的计算瓶颈:卷积和矩阵乘法,都是可以在硬件上并行化的操作。
# 于是,他们使用两个显存为3GB的NVIDIA GTX580 GPU实现了快速卷积运算。他们的创新[cuda-convnet](https://code.google.com/archive/p/cuda-convnet/)几年来它一直是行业标准,并推动了深度学习热潮。
#
# ## AlexNet
#
# 2012年,AlexNet横空出世。它首次证明了学习到的特征可以超越手工设计的特征。它一举打破了计算机视觉研究的现状。
# AlexNet使用了8层卷积神经网络,并以很大的优势赢得了2012年ImageNet图像识别挑战赛。
#
# AlexNet和LeNet的架构非常相似,如 :numref:`fig_alexnet`所示。
# 注意,这里我们提供了一个稍微精简版本的AlexNet,去除了当年需要两个小型GPU同时运算的设计特点。
#
# 
# :label:`fig_alexnet`
#
# AlexNet和LeNet的设计理念非常相似,但也存在显著差异。
# 首先,AlexNet比相对较小的LeNet5要深得多。
# AlexNet由八层组成:五个卷积层、两个全连接隐藏层和一个全连接输出层。
# 其次,AlexNet使用ReLU而不是sigmoid作为其激活函数。
# 下面,让我们深入研究AlexNet的细节。
#
# ### 模型设计
#
# 在AlexNet的第一层,卷积窗口的形状是$11\times11$。
# 由于ImageNet中大多数图像的宽和高比MNIST图像的多10倍以上,因此,需要一个更大的卷积窗口来捕获目标。
# 第二层中的卷积窗口形状被缩减为$5\times5$,然后是$3\times3$。
# 此外,在第一层、第二层和第五层卷积层之后,加入窗口形状为$3\times3$、步幅为2的最大汇聚层。
# 而且,AlexNet的卷积通道数目是LeNet的10倍。
#
# 在最后一个卷积层后有两个全连接层,分别有4096个输出。
# 这两个巨大的全连接层拥有将近1GB的模型参数。
# 由于早期GPU显存有限,原版的AlexNet采用了双数据流设计,使得每个GPU只负责存储和计算模型的一半参数。
# 幸运的是,现在GPU显存相对充裕,所以我们现在很少需要跨GPU分解模型(因此,我们的AlexNet模型在这方面与原始论文稍有不同)。
#
# ### 激活函数
#
# 此外,AlexNet将sigmoid激活函数改为更简单的ReLU激活函数。
# 一方面,ReLU激活函数的计算更简单,它不需要如sigmoid激活函数那般复杂的求幂运算。
# 另一方面,当使用不同的参数初始化方法时,ReLU激活函数使训练模型更加容易。
# 当sigmoid激活函数的输出非常接近于0或1时,这些区域的梯度几乎为0,因此反向传播无法继续更新一些模型参数。
# 相反,ReLU激活函数在正区间的梯度总是1。
# 因此,如果模型参数没有正确初始化,sigmoid函数可能在正区间内得到几乎为0的梯度,从而使模型无法得到有效的训练。
#
# ### 容量控制和预处理
#
# AlexNet通过dropout( :numref:`sec_dropout`)控制全连接层的模型复杂度,而LeNet只使用了权重衰减。
# 为了进一步扩充数据,AlexNet在训练时增加了大量的图像增强数据,如翻转、裁切和变色。
# 这使得模型更健壮,更大的样本量有效地减少了过拟合。
# 我们将在 :numref:`sec_image_augmentation`中更详细地讨论数据扩充。
#
# + origin_pos=3 tab=["tensorflow"]
import tensorflow as tf
from d2l import tensorflow as d2l
def net():
return tf.keras.models.Sequential([
# 这里,我们使用一个11*11的更大窗口来捕捉对象。
# 同时,步幅为4,以减少输出的高度和宽度。
# 另外,输出通道的数目远大于LeNet
tf.keras.layers.Conv2D(filters=96, kernel_size=11, strides=4,
activation='relu'),
tf.keras.layers.MaxPool2D(pool_size=3, strides=2),
# 减小卷积窗口,使用填充为2来使得输入与输出的高和宽一致,且增大输出通道数
tf.keras.layers.Conv2D(filters=256, kernel_size=5, padding='same',
activation='relu'),
tf.keras.layers.MaxPool2D(pool_size=3, strides=2),
# 使用三个连续的卷积层和较小的卷积窗口。
# 除了最后的卷积层,输出通道的数量进一步增加。
# 在前两个卷积层之后,汇聚层不用于减少输入的高度和宽度
tf.keras.layers.Conv2D(filters=384, kernel_size=3, padding='same',
activation='relu'),
tf.keras.layers.Conv2D(filters=384, kernel_size=3, padding='same',
activation='relu'),
tf.keras.layers.Conv2D(filters=256, kernel_size=3, padding='same',
activation='relu'),
tf.keras.layers.MaxPool2D(pool_size=3, strides=2),
tf.keras.layers.Flatten(),
# 这里,全连接层的输出数量是LeNet中的好几倍。使用dropout层来减轻过拟合
tf.keras.layers.Dense(4096, activation='relu'),
tf.keras.layers.Dropout(0.5),
tf.keras.layers.Dense(4096, activation='relu'),
tf.keras.layers.Dropout(0.5),
# 最后是输出层。由于这里使用Fashion-MNIST,所以用类别数为10,而非论文中的1000
tf.keras.layers.Dense(10)
])
# + [markdown] origin_pos=4
# [**我们构造一个**]高度和宽度都为224的(**单通道数据,来观察每一层输出的形状**)。
# 它与 :numref:`fig_alexnet`中的AlexNet架构相匹配。
#
# + origin_pos=7 tab=["tensorflow"]
X = tf.random.uniform((1, 224, 224, 1))
for layer in net().layers:
X = layer(X)
print(layer.__class__.__name__, 'output shape:\t', X.shape)
# + [markdown] origin_pos=8
# ## 读取数据集
#
# 尽管本文中AlexNet是在ImageNet上进行训练的,但我们在这里使用的是Fashion-MNIST数据集。因为即使在现代GPU上,训练ImageNet模型,同时使其收敛可能需要数小时或数天的时间。
# 将AlexNet直接应用于Fashion-MNIST的一个问题是,[**Fashion-MNIST图像的分辨率**]($28 \times 28$像素)(**低于ImageNet图像。**)
# 为了解决这个问题,(**我们将它们增加到$224 \times 224$**)(通常来讲这不是一个明智的做法,但我们在这里这样做是为了有效使用AlexNet架构)。
# 我们使用`d2l.load_data_fashion_mnist`函数中的`resize`参数执行此调整。
#
# + origin_pos=9 tab=["tensorflow"]
batch_size = 128
train_iter, test_iter = d2l.load_data_fashion_mnist(batch_size, resize=224)
# + [markdown] origin_pos=10
# ## [**训练AlexNet**]
#
# 现在,我们可以开始训练AlexNet了。与 :numref:`sec_lenet`中的LeNet相比,这里的主要变化是使用更小的学习速率训练,这是因为网络更深更广、图像分辨率更高,训练卷积神经网络就更昂贵。
#
# + origin_pos=11 tab=["tensorflow"]
lr, num_epochs = 0.01, 10
d2l.train_ch6(net, train_iter, test_iter, num_epochs, lr, d2l.try_gpu())
# + [markdown] origin_pos=12
# ## 小结
#
# * AlexNet的架构与LeNet相似,但使用了更多的卷积层和更多的参数来拟合大规模的ImageNet数据集。
# * 今天,AlexNet已经被更有效的架构所超越,但它是从浅层网络到深层网络的关键一步。
# * 尽管AlexNet的代码只比LeNet多出几行,但学术界花了很多年才接受深度学习这一概念,并应用其出色的实验结果。这也是由于缺乏有效的计算工具。
# * Dropout、ReLU和预处理是提升计算机视觉任务性能的其他关键步骤。
#
# ## 练习
#
# 1. 试着增加迭代轮数。对比LeNet的结果有什么不同?为什么?
# 1. AlexNet对于Fashion-MNIST数据集来说可能太复杂了。
# 1. 尝试简化模型以加快训练速度,同时确保准确性不会显著下降。
# 1. 设计一个更好的模型,可以直接在$28 \times 28$图像上工作。
# 1. 修改批量大小,并观察模型精度和GPU显存变化。
# 1. 分析了AlexNet的计算性能。
# 1. 在AlexNet中主要是哪部分占用显存?
# 1. 在AlexNet中主要是哪部分需要更多的计算?
# 1. 计算结果时显存带宽如何?
# 1. 将dropout和ReLU应用于LeNet-5,效果有提升吗?再试试预处理会怎么样?
#
# + [markdown] origin_pos=15 tab=["tensorflow"]
# [Discussions](https://discuss.d2l.ai/t/1862)
#
|
d2l/tensorflow/chapter_convolutional-modern/alexnet.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
# +
#os.getcwd()
# +
#os.chdir(os.path.join(os.getcwd(),"src"))
# -
from src.algorithm import *
from PIL import Image
for i in range(475,478):
print(i)
image_name = str(i)
img = load_image("examples/{}.jpg".format(image_name))
edges = get_contours(img)
perspective = get_perspective_from_contours(edges[0])
print(perspective)
board =get_boards_from_perspective(img, perspective)
plt.imshow(board._board[-1].getImage())
plt.show()
image_folder = "../data/"
save_folder = image_folder + image_name
if not os.path.exists(save_folder):
os.makedirs(save_folder)
for i in range(len(board._board)): #len(board._board)
img = board._board[i].getImage()
plt.imshow(img)
im = Image.fromarray(img)
im.save("{}/{}.png".format(save_folder, str(i)+"s" + image_name))
#image = plt.savefig("{}/{}.png".format(save_folder, str(i)))
|
data_label.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import time
# - There is a data dir which contains two txt files
# - Use your GoogleFu to figure out how to use Python to open these files
# - Save "books_published "books_published_last_two_years.txt" in the variable "recent_books"
# - Save "all_coding_books.txt" in the variable "coding_books"
recent_books = open("data/books_published_last_two_years.txt","r")
coding_books = open("data/all_coding_books.txt","r")
# Print how many books each file has
recent_books_lines = recent_books.readlines()
coding_books_lines = coding_books.readlines()
len(recent_books_lines)-1
len(coding_books_lines)-1
# Problem: Using a loop, find which books "recent_books" and "coding_books" have in common (time how long it takes).
# +
# Starter code
start = time.time() # This allows you to time the code
# Your code here
common_books = []
for i in recent_books_lines:
if i in coding_books_lines:
common_books = common_books + [i]
print(f"Execution time: {time.time() - start}") # This prints how long it took to run your code
# -
len(common_books)
# Problem: Can you think of a way to make this code run faster? Anything YOU (emphaiss on you!!) can think of is fair game (time your code).
# +
start = time.time()
# Your code here
common_books = [(i in coding_books_lines) for i in recent_books_lines]
print(f"Execution time: {time.time() - start}")
# -
¯\_(ツ)_/¯
|
CodeChallenge .ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: mmdetection
# language: python
# name: mmdetection
# ---
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
import os
import numpy as np
from comparsion import read_coco_eval
coco_path = [
"../detection/cm_rcnn_10164_step",
"../detection/cm_rcnn_10164_step0",
"../detection/cm_rcnn_10164_exp",
"../detection/cm_rcnn_10164_poly",
"../detection/cm_rcnn_10164_inv",
"../detection/cm_rcnn_10164_cos",
"../detection/cm_rcnn_10164_m56",
"../detection/cm_rcnn_10132",
"../detection/cm_rcnn_101",
"../detection/cm_rcnn_50",
"../detection/cm_rcnn_dconv",
"../detection/mask_rcnn_r50",
"../detection/cascade_rcnn_r50",
"../detection/faster_rcnn_r50",
"../detection/retinanet_r50",
]
result = read_coco_eval(coco_path, save=None)
result
name = [
"cm_rcnn_10164_a3",
"cm_rcnn_10164_a4",
"cm_rcnn_10164_a5",
"cm_rcnn_10164_a10",
"cm_rcnn_10164_a34",
"cm_rcnn_10164_a3510",
"cm_rcnn_10164_s1",
"cm_rcnn_10164_s2",
"cm_rcnn_10164_s3",
]
read_coco_eval(name, save=None)
|
concrete/coco_eval_comparsion.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
from dataicer import DirectoryHandler, list_handlers, ZipHandler
from dataicer.plugins import get_numpy_handlers
import numpy as np
import pathlib
np_data = np.arange(10)
dict_data = dict(a=1, b="hello")
list_data = [1, 2, 3, "world"]
# Saves a new folder called `first_write.ice`
# +
wp = (pathlib.Path(".")).absolute()
with ZipHandler("first_write", get_numpy_handlers(), mode="w") as dh:
dh.ice(np_data=np_data, dict_data=dict_data, list_data=list_data)
print(dh.path)
# with ZipHandler("first_write", get_numpy_handlers(), mode="a") as dh:
# dh.ice(np_data=np_data)
# -
with ZipHandler("first_write", get_numpy_handlers(), mode="a") as dh:
print(dh.keys())
dh.ice(np_data=np_data, dict_data=dict_data, list_data=list_data)
dh = ZipHandler("first_write", get_numpy_handlers(), mode="w", working_path=".")
dh.open()
# +
with DirectoryHandler("first_write", get_numpy_handlers(), mode="w") as dh:
dh.ice(np_data=np_data, dict_data=dict_data, list_data=list_data)
with DirectoryHandler("first_write", get_numpy_handlers(), mode="a") as dh:
dh.ice(np_data=np_data)
# -
(pathlib.Path(".")/"dataicer/examples").absolute()
# Append or update will overwrite the previously saved file
# +
# more_data = np_data.copy()
with DirectoryHandler("first_write", get_numpy_handlers(), mode="a") as dh:
print(dh.keys())
print(dh.path)
b = dh._key_get_uuid("np_data")
# dh.ice(np_data=np_data)
# -
b
import jsonpickle as jp
jp.decode(b)
|
examples/example_icing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.2
# language: julia
# name: julia-1.6
# ---
# # Getting started with JuMP
# This tutorial is aimed at providing a quick introduction to writing JuMP code.
# ## What is JuMP?
# JuMP ("Julia for Mathematical Programming") is an open-source modeling
# language that is embedded in Julia. It allows users to formulate various
# classes of optimization problems (linear, mixed-integer, quadratic, conic
# quadratic, semidefinite, and nonlinear) with easy-to-read code. These problems
# can then be solved using state-of-the-art open-source and commercial solvers.
# JuMP also makes advanced optimization techniques easily accessible from a
# high-level language.
# ## Installation
# JuMP is a package for Julia. From Julia, JuMP is installed by using the
# built-in package manager.
# ```julia
# import Pkg
# Pkg.add("JuMP")
# ```
# You also need to include a Julia package which provides an appropriate solver.
# One such solver is `GLPK.Optimizer`, which is provided by the
# [GLPK.jl package](https://github.com/jump-dev/GLPK.jl).
# ```julia
# import Pkg
# Pkg.add("GLPK")
# ```
# See Installation Guide for a list of other solvers you can use.
# ## An example
# Let's try to solve the following linear programming problem by using JuMP and
# GLPK. We will first look at the complete code to solve the problem and then go
# through it step by step.
# $$
# \begin{aligned}
# & \min & 12x + 20y \\
# & \;\;\text{s.t.} & 6x + 8y \geq 100 \\
# & & 7x + 12y \geq 120 \\
# & & x \geq 0 \\
# & & y \in [0, 3] \\
# \end{aligned}
# $$
using JuMP
using GLPK
model = Model(GLPK.Optimizer)
@variable(model, x >= 0)
@variable(model, 0 <= y <= 3)
@objective(model, Min, 12x + 20y)
@constraint(model, c1, 6x + 8y >= 100)
@constraint(model, c2, 7x + 12y >= 120)
print(model)
optimize!(model)
@show termination_status(model)
@show primal_status(model)
@show dual_status(model)
@show objective_value(model)
@show value(x)
@show value(y)
@show shadow_price(c1)
@show shadow_price(c2)
# ## Step-by-step
# Once JuMP is installed, to use JuMP in your programs, we just need to write:
using JuMP
# We also need to include a Julia package which provides an appropriate solver.
# We want to use `GLPK.Optimizer` here which is provided by the `GLPK.jl`
# package.
using GLPK
# A model object is a container for variables, constraints, solver options, etc.
# Models are created with the `Model` function. The model can be created
# with an optimizer attached with default arguments by calling the constructor
# with the optimizer type, as follows:
model = Model(GLPK.Optimizer)
# Variables are modeled using `@variable`:
@variable(model, x >= 0)
# They can have lower and upper bounds.
@variable(model, 0 <= y <= 30)
# The objective is set using `@objective`:
@objective(model, Min, 12x + 20y)
# Constraints are modeled using `@constraint`. Here `c1` and `c2` are
# the names of our constraint.
@constraint(model, c1, 6x + 8y >= 100)
@constraint(model, c2, 7x + 12y >= 120)
print(model)
# To solve the optimization problem, call the `optimize!` function.
optimize!(model)
# **Info**
# The `!` after optimize is just part of the name. It's nothing special.
# Julia has a convention that functions which mutate their arguments should
# end in `!`. A common example is `push!`.
# Now let's see what information we can query about the solution.
# `termination_status` tells us why the solver stopped:
termination_status(model)
# In this case, the solver found an optimal solution. We should also check
# `primal_status` to see if the solver found a primal feasible point:
primal_status(model)
# and `dual_status` to see if the solver found a dual feasible point:
dual_status(model)
# Now we know that our solver found an optimal solution, and has a primal and a
# dual solution to query.
# Query the objective value using `objective_value`:
objective_value(model)
# The primal solution using `value`:
value(x)
value(y)
# and the dual solution using `shadow_price`:
shadow_price(c1)
shadow_price(c2)
# ## Variable basics
model = Model()
# ### Variable bounds
# All of the variables we have created till now have had a bound. We can also
# create a free variable.
@variable(model, free_x)
# While creating a variable, instead of using the <= and >= syntax, we can also
# use the `lower_bound` and `upper_bound` keyword arguments.
@variable(model, keyword_x, lower_bound = 1, upper_bound = 2)
# We can query whether a variable has a bound using the `has_lower_bound` and
# `has_upper_bound` functions. The values of the bound can be obtained using the
# `lower_bound` and `upper_bound` functions.
has_upper_bound(keyword_x)
upper_bound(keyword_x)
# Note querying the value of a bound that does not exist will result in an error.
lower_bound(free_x)
# JuMP also allows us to change the bounds on variable. We will learn this in
# the problem modification tutorial.
# ### Containers
# We have already seen how to add a single variable to a model using the
# `@variable` macro. Let's now look at more ways to add variables to a
# JuMP model.
# JuMP provides data structures for adding collections of variables to a model.
# These data structures are referred to as Containers and are of three types:
# `Arrays`, `DenseAxisArrays`, and `SparseAxisArrays`.
# #### Arrays
# JuMP arrays are created in a similar syntax to Julia arrays with the addition
# of specifying that the indices start with 1. If we do not tell JuMP that the
# indices start at 1, it will create a `DenseAxisArray` instead.
@variable(model, a[1:2, 1:2])
# An n-dimensional variable $x \in {R}^n$ having a bound $l \preceq x \preceq u$
# ($l, u \in {R}^n$) is added in the following manner.
# +
n = 10
l = [1; 2; 3; 4; 5; 6; 7; 8; 9; 10]
u = [10; 11; 12; 13; 14; 15; 16; 17; 18; 19]
@variable(model, l[i] <= x[i = 1:n] <= u[i])
# -
# Note that while working with Containers, we can also create variable bounds
# depending upon the indices:
@variable(model, y[i = 1:2, j = 1:2] >= 2i + j)
# #### DenseAxisArrays
# `DenseAxisArrays` are used when the required indices are not one-based integer
# ranges. The syntax is similar except with an arbitrary vector as an index as
# opposed to a one-based range.
# An example where the indices are integers but do not start with one.
@variable(model, z[i = 2:3, j = 1:2:3] >= 0)
# Another example where the indices are an arbitrary vector.
@variable(model, w[1:5, ["red", "blue"]] <= 1)
# #### SparseAxisArrays
# `SparseAxisArrays` are created when the indices do not form a rectangular set.
# For example, this applies when indices have a dependence upon previous indices
# (called triangular indexing).
@variable(model, u[i = 1:3, j = i:5])
# We can also conditionally create variables by adding a comparison check that
# depends upon the named indices and is separated from the indices by a
# semi-colon (;).
@variable(model, v[i = 1:9; mod(i, 3) == 0])
# ### Variable types
# The last argument to the `@variable` macro is usually the variable type. Here
# we'll look at how to specify the variable type.
# #### Integer variables
# Integer optimization variables are constrained to the set $x \in {Z}$
@variable(model, integer_x, Int)
# or
@variable(model, integer_z, integer = true)
# #### Binary variables
# Binary optimization variables are constrained to the set $x \in \{0, 1\}$.
@variable(model, binary_x, Bin)
# or
@variable(model, binary_z, binary = true)
# ## Constraint basics
model = Model()
@variable(model, x)
@variable(model, y)
@variable(model, z[1:10]);
# ### Constraint references
# While calling the `@constraint` macro, we can also set up a constraint
# reference. Such a reference is useful for obtaining additional information
# about the constraint, such as its dual solution.
@constraint(model, con, x <= 4)
# ### Containers
# Just as we had containers for variables, JuMP also provides `Arrays`,
# `DenseAxisArrays`, and `SparseAxisArrays` for storing collections of
# constraints. Examples for each container type are given below.
# #### Arrays
@constraint(model, [i = 1:3], i * x <= i + 1)
# #### DenseAxisArrays
@constraint(model, [i = 1:2, j = 2:3], i * x <= j + 1)
# #### SparseAxisArrays
@constraint(model, [i = 1:2, j = 1:2; i != j], i * x <= j + 1)
# ### Constraints in a loop
# We can add constraints using regular Julia loops
for i in 1:3
@constraint(model, 6x + 4y >= 5i)
end
# or use for each loops inside the `@constraint` macro.
@constraint(model, [i in 1:3], 6x + 4y >= 5i)
# We can also create constraints such as $\sum _{i = 1}^{10} z_i \leq 1$
@constraint(model, sum(z[i] for i in 1:10) <= 1)
# ## Objective functions
# While the recommended way to set the objective is with the `@objective`
# macro, the functions `set_objective_sense` and `set_objective_function`
# provide an equivalent lower-level interface.
# +
using GLPK
model = Model(GLPK.Optimizer)
@variable(model, x >= 0)
@variable(model, y >= 0)
set_objective_sense(model, MOI.MIN_SENSE)
set_objective_function(model, x + y)
optimize!(model)
# -
objective_value(model)
# To query the objective function from a model, we use the `objective_sense`,
# `objective_function`, and `objective_function_type` functions.
objective_sense(model)
objective_function(model)
objective_function_type(model)
# ## Vectorized syntax
# We can also add constraints and an objective to JuMP using vectorized linear
# algebra. We'll illustrate this by solving an LP in standard form i.e.
# $$
# \begin{aligned}
# & \min & c^T x \\
# & \;\;\text{s.t.} & A x = b \\
# & & x \succeq 0 \\
# & & x \in \mathbb{R}^n
# \end{aligned}
# $$
# +
vector_model = Model(GLPK.Optimizer)
A = [
1 1 9 5
3 5 0 8
2 0 6 13
]
b = [7; 3; 5]
c = [1; 3; 5; 2]
@variable(vector_model, x[1:4] >= 0)
@constraint(vector_model, A * x .== b)
@objective(vector_model, Min, c' * x)
optimize!(vector_model)
# -
objective_value(vector_model)
# ---
#
# *This notebook was generated using [Literate.jl](https://github.com/fredrikekre/Literate.jl).*
|
getting_started_with_JuMP.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data ETL - Function Testing
#
# This notebook implements the functions developed for Twitter data ETL. See notebooks 1.2 and 1.3, as well as documentation to understand the steps developed/implemented in these functions.
# +
import os
import sys
from os.path import join
import pandas as pd
import numpy as np
project_dir = join(os.getcwd(), os.pardir)
raw_dir = join(project_dir, 'data', 'raw')
raw_fname = 'data_pull_sample.json'
sys.path.append(project_dir)
from src.data import transform, load_es
# -
# Initialize Elastic Search and Kibana docker containers:
os.chdir(project_dir)
# !make database
os.chdir(join(project_dir, 'notebooks'))
df = transform(join(raw_dir, raw_fname))
load_es(df, verbose=True)
|
notebooks/1.4-jf-data-etl-testing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# Lecture 32: Convolutional Autoencoder for Representation Learning
# ===
# %matplotlib inline
import torch
from PIL import Image
import matplotlib.pyplot as plt
import numpy as np
import torchvision
import torchvision.transforms as transforms
from torch.autograd import Variable
from torch.autograd import Function
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
import copy
import time
# Load Data:
# ==
# +
transform = transforms.Compose([transforms.ToTensor()])
BatchSize = 2000
trainset = torchvision.datasets.CIFAR10(root='./CIFAR10', train=True,
download=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=BatchSize,
shuffle=True, num_workers=4) # Creating dataloader
testset = torchvision.datasets.CIFAR10(root='./CIFAR10', train=False,
download=True, transform=transform)
testloader = torch.utils.data.DataLoader(testset, batch_size=BatchSize,
shuffle=False, num_workers=4) # Creating dataloader
classes = ('plane', 'car', 'bird', 'cat',
'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
# -
# Check availability of GPU
use_gpu = torch.cuda.is_available()
if use_gpu:
print('GPU is available!')
# Convolutional Autoencoder:
# ==
class autoencoder(nn.Module):
def __init__(self):
super(autoencoder, self).__init__()
self.conv_encoder = nn.Sequential(
nn.Conv2d(in_channels=3, out_channels=64, kernel_size=3, stride=2, padding=1),
nn.LeakyReLU(0.1),
nn.Conv2d(in_channels=64, out_channels=128, kernel_size=3, stride=2, padding=1),
nn.LeakyReLU(0.1),
nn.Conv2d(in_channels=128, out_channels=128, kernel_size=3, stride=2, padding=1),
nn.LeakyReLU(0.1))
self.fc_encoder = nn.Sequential(
nn.Linear(128*4*4,1024),
nn.LeakyReLU(0.1))
self.fc_decoder = nn.Sequential(
nn.Linear(1024,128*4*4),
nn.LeakyReLU(0.1))
self.conv_decoder = nn.Sequential(
nn.Conv2d(in_channels=128, out_channels=128, kernel_size=3, stride=1, padding=1),
nn.LeakyReLU(0.1),
nn.Upsample(scale_factor=2, mode='bilinear'),
nn.Conv2d(in_channels=128, out_channels=64, kernel_size=3, stride=1, padding=1),
nn.LeakyReLU(0.1),
nn.Upsample(scale_factor=2, mode='bilinear'),
nn.Conv2d(in_channels=64, out_channels=64, kernel_size=3, stride=1, padding=1),
nn.LeakyReLU(0.1),
nn.Upsample(scale_factor=2, mode='bilinear'),
nn.Conv2d(in_channels=64, out_channels=3, kernel_size=3, stride=1, padding=1),
nn.ReLU())
def forward(self, x):
x = self.conv_encoder(x)
x = x.view(-1, 128*4*4)
x = self.fc_encoder(x)
x = self.fc_decoder(x)
x = x.view(-1, 128,4,4)
x = self.conv_decoder(x)
return x
net = autoencoder()
if use_gpu:
net = net.cuda()
print(net)
init_weights = copy.deepcopy(net.conv_encoder[0].weight.data)
# Train Autoencoder:
# ==
# +
iterations = 10
criterion = nn.MSELoss()
optimizer = optim.Adam(net.parameters(), lr=1e-3)
trainLoss = []
for epoch in range(iterations): # loop over the dataset multiple times
epochStart = time.time()
runningLoss = 0.0
for i, data in enumerate(trainloader, 0):
# get the inputs
inputs, labels = data
# wrap them in Variable
if use_gpu:
inputs = Variable(inputs).cuda()
else:
inputs = Variable(inputs)
optimizer.zero_grad() # zeroes the gradient buffers of all parameters
outputs = net(inputs) # forward
loss = criterion(outputs, inputs) # calculate loss
loss.backward() # backpropagate the loss
optimizer.step()
runningLoss += loss.data[0]
trainLoss.append((runningLoss/(60000/BatchSize)))
epochEnd = time.time()-epochStart
print('Iteration: {:.0f} /{:.0f} ; Training Loss: {:.6f} ; Time consumed: {:.0f}m {:.0f}s '\
.format(epoch + 1,iterations,runningLoss/(60000/BatchSize),epochEnd//60,epochEnd%60))
print('Finished Training')
fig = plt.figure()
plt.plot(range(epoch+1),trainLoss,'g-',label='Loss')
plt.legend(loc='best')
plt.xlabel('Epochs')
plt.ylabel('Training loss')
# -
# Weights Visualization:
# ====
# functions to show an image
def imshow(img, strlabel):
npimg = img.numpy()
npimg = np.abs(npimg)
fig_size = plt.rcParams["figure.figsize"]
fig_size[0] = 10
fig_size[1] = 10
plt.rcParams["figure.figsize"] = fig_size
plt.figure()
plt.title(strlabel)
plt.imshow(np.transpose(npimg, (1, 2, 0)))
# +
trained_weights = copy.deepcopy(net.conv_encoder[0].weight.data)
d_weights = init_weights - trained_weights
if use_gpu:
init_weights = init_weights.view(64,3,3,3).cpu()
trained_weights = trained_weights.view(64,3,3,3).cpu()
d_weights = d_weights.view(64,3,3,3).cpu()
else:
init_weights = init_weights.view(64,3,3,3)
trained_weights = trained_weights.view(64,3,3,3)
d_weights = d_weights.view(64,3,3,3)
imshow(torchvision.utils.make_grid(init_weights,nrow=8,normalize=True),'Initial Weights')
imshow(torchvision.utils.make_grid(trained_weights,nrow=8,normalize=True),'Trained Weights')
imshow(torchvision.utils.make_grid(d_weights,nrow=8,normalize=True), 'Weight update')
# -
# Modifying the autoencoder for classification:
# ==
# +
class Model(nn.Module):
def __init__(self):
super(Model, self).__init__()
self.conv = net.conv_encoder
self.fc1 = net.fc_encoder
self.fc2 = nn.Sequential(nn.Linear(1024, 10))
def forward(self, x):
x = self.conv(x)
x = x.view(-1, 128*4*4)
x = self.fc1(x)
x = self.fc2(x)
return x
net = Model()
print(net)
if use_gpu:
net = net.cuda()
else:
net = net
# Copying initial weights for visualization
cll_weights = copy.deepcopy(net.conv[0].weight.data)
# -
# Train Classifier:
# ==
# +
iterations = 10
optimizer = optim.Adam(net.parameters(), lr=1e-3)
criterion = nn.CrossEntropyLoss()
trainLoss = []
testacc = []
for epoch in range(iterations): # loop over the dataset multiple times
epochStart = time.time()
runningLoss = 0.0
for i, data in enumerate(trainloader, 0):
# get the inputs
inputs, labels = data
# wrap them in Variable
if use_gpu:
inputs, labels = Variable(inputs).cuda(), Variable(labels).cuda()
else:
inputs, labels = Variable(inputs), Variable(labels)
optimizer.zero_grad() # zeroes the gradient buffers of all parameters
outputs = net(inputs) # forward
loss = criterion(outputs, labels) # calculate loss
loss.backward() # backpropagate the loss
optimizer.step()
runningLoss += loss.data[0]
correct = 0
total = 0
for data in testloader:
inputs, labels = data
if use_gpu:
inputs, labels = Variable(inputs).cuda(), labels.cuda()
else:
inputs, labels = Variable(inputs), labels
outputs = net(inputs)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum()
trainLoss.append((runningLoss/(60000/BatchSize)))
testacc.append(100 * correct /float(total))
epochEnd = time.time()-epochStart
print('Iteration: {:.0f} /{:.0f} ; Training Loss: {:.6f} ; Testing Acc: {:.3f} ; Time consumed: {:.0f}m {:.0f}s '\
.format(epoch + 1,iterations,runningLoss/(60000/BatchSize),100 * correct /float(total),epochEnd//60,epochEnd%60))
print('Finished Training')
fig = plt.figure()
plt.plot(range(epoch+1),trainLoss,'g-',label='Train Loss')
plt.legend(loc='best')
plt.xlabel('Epochs')
plt.ylabel('Training loss')
fig = plt.figure()
plt.plot(range(epoch+1),testacc,'r-',label='Test Acc')
plt.legend(loc='best')
plt.xlabel('Epochs')
plt.ylabel('Test Accuracy')
# -
# Encoder Weights Visualization:
# ==
# +
cll_weights_ft = copy.deepcopy(net.conv[0].weight.data)
d_weights = cll_weights-cll_weights_ft
if use_gpu:
cll_weights = cll_weights.view(64,3,3,3).cpu()
cll_weights_ft = cll_weights_ft.view(64,3,3,3).cpu()
d_weights = d_weights.view(64,3,3,3).cpu()
else:
cll_weights = cll_weights.view((64,3,3,3))
cll_weights_ft = cll_weights_ft.view((64,3,3,3))
d_weights = d_weights.view((64,3,3,3))
imshow(torchvision.utils.make_grid(cll_weights,nrow=8,normalize=True),'Trained Weights')
imshow(torchvision.utils.make_grid(cll_weights_ft,nrow=8,normalize=True),'Finetuned Weights')
imshow(torchvision.utils.make_grid(d_weights,nrow=8,normalize=True), 'Weight update')
# -
# Performance of different Classes:
# ==
# +
class_correct = list(0. for i in range(10))
class_total = list(0. for i in range(10))
for data in testloader:
images, labels = data
if use_gpu:
outputs = net(Variable(images.cuda()))
_, predicted = torch.max(outputs.data.cpu(), 1)
else:
outputs = net(Variable(images))
_, predicted = torch.max(outputs.data, 1)
c = (predicted == labels).squeeze()
for i in range(BatchSize):
label = labels[i]
class_correct[label] += c[i]
class_total[label] += 1
for i in range(10):
print('Accuracy of %5s : %f %%' % (
classes[i], 100 * class_correct[i] / float(class_total[i])))
# -
|
lecture32.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1 align="center">Métodos Numéricos</h1>
# <h1 align="center">Capítulo 5: Diferenciación e integración numérica</h1>
# <h1 align="center">2021/02</h1>
# <h1 align="center">MEDELLÍN - COLOMBIA </h1>
# <table>
# <tr align=left><td><img align=left src="https://github.com/carlosalvarezh/Metodos_Numericos/blob/master/images/CC-BY.png?raw=true">
# <td>Text provided under a Creative Commons Attribution license, CC-BY. All code is made available under the FSF-approved MIT license.(c) <NAME></td>
# </table>
# ***
#
# ***Docente:*** <NAME>, I.C. D.Sc.
#
# ***e-mail:*** <EMAIL>
#
# ***skype:*** carlos.alberto.alvarez.henao
#
# ***Linkedin:*** https://www.linkedin.com/in/carlosalvarez5/
#
# ***github:*** https://github.com/carlosalvarezh/Metodos_Numericos
#
# ***Herramienta:*** [Jupyter](http://jupyter.org/)
#
# ***Kernel:*** Python 3.8
#
#
# ***
# <a id='TOC'></a>
# + [markdown] toc=true
# <h1>Tabla de Contenidos<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Diferenciación-Numérica" data-toc-modified-id="Diferenciación-Numérica-1"><span class="toc-item-num">1 </span>Diferenciación Numérica</a></span><ul class="toc-item"><li><span><a href="#Introducción" data-toc-modified-id="Introducción-1.1"><span class="toc-item-num">1.1 </span>Introducción</a></span></li><li><span><a href="#Series-de-Taylor" data-toc-modified-id="Series-de-Taylor-1.2"><span class="toc-item-num">1.2 </span>Series de Taylor</a></span></li><li><span><a href="#Esquemas-de-diferencias-finitas-para-la-primera-derivada" data-toc-modified-id="Esquemas-de-diferencias-finitas-para-la-primera-derivada-1.3"><span class="toc-item-num">1.3 </span>Esquemas de diferencias finitas para la primera derivada</a></span><ul class="toc-item"><li><span><a href="#Esquema-de-primer-orden-hacia-adelante-(forward)" data-toc-modified-id="Esquema-de-primer-orden-hacia-adelante-(forward)-1.3.1"><span class="toc-item-num">1.3.1 </span>Esquema de primer orden hacia adelante (forward)</a></span></li><li><span><a href="#Esquema-de-primer-orden-hacia-atrás-(backward)" data-toc-modified-id="Esquema-de-primer-orden-hacia-atrás-(backward)-1.3.2"><span class="toc-item-num">1.3.2 </span>Esquema de primer orden hacia atrás (backward)</a></span></li><li><span><a href="#Esquema-de-segundo-orden-(central)" data-toc-modified-id="Esquema-de-segundo-orden-(central)-1.3.3"><span class="toc-item-num">1.3.3 </span>Esquema de segundo orden (central)</a></span></li><li><span><a href="#Resumen-esquemas-diferencias-finitas-para-la-primera-derivada" data-toc-modified-id="Resumen-esquemas-diferencias-finitas-para-la-primera-derivada-1.3.4"><span class="toc-item-num">1.3.4 </span>Resumen esquemas diferencias finitas para la primera derivada</a></span></li></ul></li><li><span><a href="#Esquemas-de-diferencias-finitas-para-la-segunda-derivada" data-toc-modified-id="Esquemas-de-diferencias-finitas-para-la-segunda-derivada-1.4"><span class="toc-item-num">1.4 </span>Esquemas de diferencias finitas para la segunda derivada</a></span></li><li><span><a href="#Implementación-computacional-de-algunos-esquemas-de-diferencias-finitas" data-toc-modified-id="Implementación-computacional-de-algunos-esquemas-de-diferencias-finitas-1.5"><span class="toc-item-num">1.5 </span>Implementación computacional de algunos esquemas de diferencias finitas</a></span></li></ul></li><li><span><a href="#Integración-Numérica" data-toc-modified-id="Integración-Numérica-2"><span class="toc-item-num">2 </span>Integración Numérica</a></span><ul class="toc-item"><li><span><a href="#Introducción" data-toc-modified-id="Introducción-2.1"><span class="toc-item-num">2.1 </span>Introducción</a></span></li><li><span><a href="#Fórmulas-de-integración-de-Newton---Cotes" data-toc-modified-id="Fórmulas-de-integración-de-Newton---Cotes-2.2"><span class="toc-item-num">2.2 </span>Fórmulas de integración de <em>Newton - Cotes</em></a></span></li><li><span><a href="#Regla-trapezoidal" data-toc-modified-id="Regla-trapezoidal-2.3"><span class="toc-item-num">2.3 </span>Regla trapezoidal</a></span><ul class="toc-item"><li><span><a href="#Regla-trapezoidal-de-aplicación-simple" data-toc-modified-id="Regla-trapezoidal-de-aplicación-simple-2.3.1"><span class="toc-item-num">2.3.1 </span>Regla trapezoidal de aplicación simple</a></span></li><li><span><a href="#Regla-trapezoidal-de-aplicación-múltiple" data-toc-modified-id="Regla-trapezoidal-de-aplicación-múltiple-2.3.2"><span class="toc-item-num">2.3.2 </span>Regla trapezoidal de aplicación múltiple</a></span></li><li><span><a href="#Implementación-computacional" data-toc-modified-id="Implementación-computacional-2.3.3"><span class="toc-item-num">2.3.3 </span>Implementación computacional</a></span></li><li><span><a href="#Error-en-la-aplicación-de-la-regla-trapezoidal" data-toc-modified-id="Error-en-la-aplicación-de-la-regla-trapezoidal-2.3.4"><span class="toc-item-num">2.3.4 </span>Error en la aplicación de la regla trapezoidal</a></span></li></ul></li><li><span><a href="#Reglas-de-Simpson" data-toc-modified-id="Reglas-de-Simpson-2.4"><span class="toc-item-num">2.4 </span>Reglas de Simpson</a></span><ul class="toc-item"><li><span><a href="#Regla-de-Simpson1/3-de-aplicación-simple" data-toc-modified-id="Regla-de-Simpson1/3-de-aplicación-simple-2.4.1"><span class="toc-item-num">2.4.1 </span>Regla de Simpson1/3 de aplicación simple</a></span></li><li><span><a href="#Error-en-la-regla-de-Simpson-1/3-de-aplicación-simple" data-toc-modified-id="Error-en-la-regla-de-Simpson-1/3-de-aplicación-simple-2.4.2"><span class="toc-item-num">2.4.2 </span>Error en la regla de Simpson 1/3 de aplicación simple</a></span></li><li><span><a href="#Regla-de-simpson1/3-de-aplicación-múltiple" data-toc-modified-id="Regla-de-simpson1/3-de-aplicación-múltiple-2.4.3"><span class="toc-item-num">2.4.3 </span>Regla de simpson1/3 de aplicación múltiple</a></span></li><li><span><a href="#Implementación-computacional-regla-de-Simpson1/3-de-aplicación-múltiple" data-toc-modified-id="Implementación-computacional-regla-de-Simpson1/3-de-aplicación-múltiple-2.4.4"><span class="toc-item-num">2.4.4 </span>Implementación computacional regla de Simpson1/3 de aplicación múltiple</a></span></li><li><span><a href="#Regla-de-Simpson-3/8-de-aplicación-simple" data-toc-modified-id="Regla-de-Simpson-3/8-de-aplicación-simple-2.4.5"><span class="toc-item-num">2.4.5 </span>Regla de Simpson 3/8 de aplicación simple</a></span></li><li><span><a href="#Regla-de-Simpson3/8-de-aplicación-múltiple" data-toc-modified-id="Regla-de-Simpson3/8-de-aplicación-múltiple-2.4.6"><span class="toc-item-num">2.4.6 </span>Regla de Simpson3/8 de aplicación múltiple</a></span></li><li><span><a href="#Implementación-computacional-de-la-regla-de-Simpson3/8-de-aplicación-múltiple" data-toc-modified-id="Implementación-computacional-de-la-regla-de-Simpson3/8-de-aplicación-múltiple-2.4.7"><span class="toc-item-num">2.4.7 </span>Implementación computacional de la regla de Simpson3/8 de aplicación múltiple</a></span></li></ul></li><li><span><a href="#Cuadratura-de-Gauss" data-toc-modified-id="Cuadratura-de-Gauss-2.5"><span class="toc-item-num">2.5 </span>Cuadratura de Gauss</a></span><ul class="toc-item"><li><span><a href="#Introducción" data-toc-modified-id="Introducción-2.5.1"><span class="toc-item-num">2.5.1 </span>Introducción</a></span></li><li><span><a href="#Determinación-de-los-coeficientes" data-toc-modified-id="Determinación-de-los-coeficientes-2.5.2"><span class="toc-item-num">2.5.2 </span>Determinación de los coeficientes</a></span></li><li><span><a href="#Cambios-de-los-límites-de-integración" data-toc-modified-id="Cambios-de-los-límites-de-integración-2.5.3"><span class="toc-item-num">2.5.3 </span>Cambios de los límites de integración</a></span></li><li><span><a href="#Fórmulas-de-punto-superior" data-toc-modified-id="Fórmulas-de-punto-superior-2.5.4"><span class="toc-item-num">2.5.4 </span>Fórmulas de punto superior</a></span></li><li><span><a href="#Ejemplo-Cuadratura-de-Gauss" data-toc-modified-id="Ejemplo-Cuadratura-de-Gauss-2.5.5"><span class="toc-item-num">2.5.5 </span>Ejemplo Cuadratura de Gauss</a></span></li></ul></li></ul></li></ul></div>
# -
# ## Diferenciación Numérica
# ### Introducción
# La [diferenciación numérica](https://en.wikipedia.org/wiki/Numerical_differentiation) se emplea para determinar (estimar) el valor de la derivada de una función en un punto específico. No confundir con la derivada de una función, pues lo que se obtendrá es un valor puntual y no una función. En este capítulo nos centraremos únicamente en ecuiaciones unidimensionales.
# [Volver a la Tabla de Contenido](#TOC)
# ### Series de Taylor
# De la [serie de Taylor](https://en.wikipedia.org/wiki/Taylor_series)
#
# <a id='Ec5_1'></a>
# \begin{equation*}
# f(x_{i \pm 1}) = f(x_i) \pm f'(x_i)h + \frac{f''(x_i)h^2}{2!} \pm \frac{f'''(x_i)h^3}{3!} + \ldots
# \label{eq:Ec5_1} \tag{5.1}
# \end{equation*}
#
# con $h=\Delta x = x_{i+1}-x_i$ siendo el tamaño de paso.
#
# Dada que la serie contiene infinitos términos, partir de la ecuación ($5.1$) se pueden obtener infinitos esquemas numéricos para determinar cada una de las infinitas derivadas de dicho polinomio. En este curso usaremos la técnica de [Diferencias Finitas](https://en.wikipedia.org/wiki/Finite_difference) para desarrollarlas.
# [Volver a la Tabla de Contenido](#TOC)
# ### Esquemas de diferencias finitas para la primera derivada
# #### Esquema de primer orden hacia adelante (forward)
# De la ecuación [(5.1)](#Ec5_1) tomando los valores positivos, que involucran únicamente términos hacia adelante, se trunca la serie hasta la primera derivada y se realiza un despeje algebraico para llegar a:
#
# <a id='Ec5_2'></a>
# \begin{equation*}
# f'(x_i) = \frac{f(x_{i+1})-f(x_i)}{h} + \mathcal{O}(h)
# \label{eq:Ec5_2} \tag{5.2}
# \end{equation*}
#
# se puede observar que el término $\mathcal{O}(h)$ indica que el error es de orden lineal, es decir, si se reduce el tamaño de paso, $h$, a la mitad, el error se reducirá a la mitad. Si se reduc el tamaño de paso a una cuarta parte, el error se reducirá, linealmente, una cuarta parte.
# [Volver a la Tabla de Contenido](#TOC)
# #### Esquema de primer orden hacia atrás (backward)
# De la ecuación [(5.1)](#Ec5_1) tomando los valores negativos, que involucran únicamente términos hacia atrás (backward), se trunca la serie hasta la primera derivada y se realiza un despeje algebraico para llegar a:
#
# <a id='Ec5_1'></a>
# \begin{equation*}
# f'(x_i) = \frac{f(x_{i})-f(x_{i-1})}{h} + \mathcal{O}(h)
# \label{eq:Ec5_3} \tag{5.3}
# \end{equation*}
#
# se observa que se llega a una expresión similar a la de la ecuación [(5.2)](#Ec5_2), pero de esta vez, se tiene en cuenta es el valor anterior al punto $x_i$. También se observa que el error es de orden lineal, por lo que se mantiene un esquema de primer orden.
#
#
# [Volver a la Tabla de Contenido](#TOC)
# #### Esquema de segundo orden (central)
# Una forma de aumentar el orden de estos esquemas, es realizar el truncamiento de la *serie de Taylor* hasta la segunda derivada, hacia adelante y hacia atras, y realizar su resta aritmética.
#
# <a id='Ec5_4'></a>
# \begin{equation*}
# \begin{split}
# f(x_{i+1}) & = f(x_i) + f'(x_i)h + \frac{f''(x_i)h^2}{2!} \\
# - \\
# f(x_{i-1}) & = f(x_i) - f'(x_i)h + \frac{f''(x_i)h^2}{2!} \\
# \hline \\
# f(x_{i+1}) - f(x_{i-1}) & = 2 f'(x_i)h
# \end{split}
# \label{eq:Ec5_4} \tag{5.4}
# \end{equation*}
#
# de la anterior ecuación, despejando el término que corresponde a la primera derivada queda:
#
# <a id='Ec5_5'></a>
# \begin{equation*}
# \begin{split}
# f'(x_i) = \frac{f(x_{i+1}) - f(x_{i-1})}{2h} + \mathcal{O}(h^2)
# \end{split}
# \label{eq:Ec5_5} \tag{5.5}
# \end{equation*}
#
# se llega al esquema de diferencias finitas central para la primera derivada, que es de orden dos, es decir, si se disminuye el tamaño de paso, $h$, a la mitad, el error se disminuye una cuarta partes. En principio, esta es una mejor aproximación que los dos esquemas anteriores. La selección del esquema dependerá de la disponibilidad de puntos y del fenómeno físico a tratar.
# [Volver a la Tabla de Contenido](#TOC)
# #### Resumen esquemas diferencias finitas para la primera derivada
# Como la serie de Taylor es infinita, se podrían determinar infinitos esquemas de diferentes ordenes para la primera derivada. En la siguiente tabla se presentan algunos esquemas de diferencias finitas para la primera derivada de diferentes órdenes. Se deja al estudiante la consulta de otros esquemas.
#
# |***Esquema***|***Función***|***Error***|
# |:-----:|:-----:|:---:|
# |***Forward***|$$f´(x_0)=\frac{f(x_0+h)-f(x_0)}{h}$$|$$\mathcal{O}(h)$$|
# | |$$f´(x_0)=\frac{-3f(x_0)+4f(x_0+h)-f(x_0+2h)}{2h}$$|$$\mathcal{O}(h^2)$$|
# |***Central***|$$f´(x_0)=\frac{f(x_0+h)-f(x_0-h)}{2h}$$|$$\mathcal{O}(h^2)$$|
# | |$$f´(x_0)=\frac{f(x_0-2h)-8f(x_0-h)+8f(x_0+h)-f(x_0+2h)}{12h}$$|$$\mathcal{O}(h^4)$$|
# |***Backward***|$$f´(x_0)=\frac{f(x_0)-f(x_0-h)}{h}$$|$$\mathcal{O}(h)$$|
# | |$$f´(x_0)=\frac{f(x_0-2h)-4f(x_0-h)+3f(x_0)}{2h}$$|$$\mathcal{O}(h^2)$$|
#
# [Volver a la Tabla de Contenido](#TOC)
# ### Esquemas de diferencias finitas para la segunda derivada
# Siguiendo con la misma forma de abordar el problema para la primera derivada, si se amplian los términos en la serie de Taylor hasta la tercera derivada tanto hacia adelante como hacia atrás, y se suman, se llega a:
#
# \begin{equation*}
# \begin{split}
# f(x_{i+1}) & = f(x_i) + f'(x_i)h + \frac{f''(x_i)h^2}{2!} + \frac{f'''(x_i)h^3}{3!}\\
# + \\
# f(x_{i-1}) & = f(x_i) - f'(x_i)h + \frac{f''(x_i)h^2}{2!} - \frac{f'''(x_i)h^3}{3!}\\
# \hline \\
# f(x_{i+1}) + f(x_{i-1}) & = 2 f(x_i) + 2f''(x_i)\frac{h^2}{2!} + \mathcal{O}(h^3)
# \end{split}
# \label{eq:Ec5_6} \tag{5.6}
# \end{equation*}
#
# Despejando para el término de la segunda derivada, se llega a:
#
# <a id='Ec5_7'></a>
# \begin{equation*}
# \begin{split}
# f''(x_i) = \frac{f(x_{i+1}) - 2f(x_i) + f(x_{i-1})}{h^2} + \mathcal{O}(h^3)
# \end{split}
# \label{eq:Ec5_7} \tag{5.7}
# \end{equation*}
#
# Que corresponde a un esquema de diferencias finitas de segundo orden para la segunda derivada. A este esquema también se le llama "*molécula de tres puntos*"
#
# Igual que para la primera derivada, se pueden determinar infinitos esquemas de diferentes órdenes para la segunda derivada, y derivadas superiores. A continuación se muestra un cuadro resumen de algunos esquemas de diferencias finitas para la segunda derivada. Se deja al estudiante la revisión de esquemas de mayor orden para la segunda derivada y derivadas superiores.
#
# |***Esquema***|***Función***|***Error***|
# |:-----:|:-----:|:---:|
# |***Forward***|$$f''(x_0)=\frac{f(x_0)-2f(x_0+h)+f(x_0+2h)}{h^2}$$|$$\mathcal{O}(h)$$|
# | |$$f''(x_0)=\frac{2f(x_0)-5f(x_0+h)+4f(x_0+2h)-f(x_0+3h)}{h^2}$$|$$\mathcal{O}(h^2)$$|
# |***Central***|$$f''(x_0)=\frac{f(x_0-h)-2f(x_0)+f(x_0+h)}{h^2}$$|$$\mathcal{O}(h^2)$$|
# | |$$f''(x_0)=\frac{-f(x_0-2h)+16f(x_0-h)-30f(x_0)+16f(x_0+h)-f(x_0+2h)}{12h^2}$$|$$\mathcal{O}(h^4)$$|
# |***Backward***|$$f''(x_0)=\frac{f(x_0-2h)-2f(x_0-h)+f(x_0)}{h}$$|$$\mathcal{O}(h^2)$$|
# | |$$f''(x_0)=\frac{-f(x_0-3h)+4f(x_0-2h)-5f(x_0-h)+2f(x_0)}{h^2}$$|$$\mathcal{O}(h^2)$$|
#
# [Volver a la Tabla de Contenido](#TOC)
# ### Implementación computacional de algunos esquemas de diferencias finitas
# A manera de ejemplo, se implementarán algunos esquemas simples de diferencias finitas para la primera derivada. Se deja como actividad a los estudiantes la implementación de otros esquemas para las diferentes derivadas.
# +
import numpy as np
import matplotlib.pyplot as plt
import sympy as sym
sym.init_printing()
# +
#Esquemas de diferencias finitas para la primera derivada
def df1df(x0, h):
# Esquema de diferencias finitas para la primera derivada hacia adelante (forward)
return (f(x0 + h) - f(x0)) / h
def df1db(x0, h):
# Esquema de diferencias finitas para la primera derivada hacia atrás (backward)
return (f(x0) - f(x0 - h) ) / h
def df1dc(x0,h):
# Esquema de diferencias finitas para la primera derivada central (central)
return (f(x0 + h) - f(x0 - h) ) / (2 * h)
# -
#funcion a determinar el valor de la derivada
def f(x):
return 2*x**3 - 3*x**2 + 5*x+0.8
# +
#cálculo y evaluación de la primera derivada empleando cálculo simbólico
def df1de(x0):
x = sym.Symbol('x')
df = sym.diff(f(x), x)
#print(df)
df1 = df.evalf(subs={x:x0})
return df1
# +
h = 0.1
x0 = 0.8
print("1st derivative \t Value \t\t Error(%)")
print('---------------------------------------')
pde = df1de(x0)
pdf = df1df(x0, h)
epdf = abs((pde - pdf) / pde * 100)
print("forward \t {0:6.4f} \t {1:6.2f}".format(pdf,epdf))
pdb = df1db(x0, h)
epdb = abs((pde - pdb) / pde * 100)
print("backward \t {0:6.4f} \t {1:6.2f}".format(pdb,epdb))
pdc = df1dc(x0,h)
epdc = abs((pde - pdc) / pde * 100)
print("central \t {0:6.4f} \t {1:6.2f}".format(pdc, epdc))
print("exacta \t\t {0:6.4f} \t {1}".format(pde, ' -'))
# -
# [Volver a la Tabla de Contenido](#TOC)
# ## Integración Numérica
# ### Introducción
# La [integración numérica](https://en.wikipedia.org/wiki/Numerical_integration) aborda una amplia gama de algoritmos para determinar el valor numérico (aproximado) de una integral definida. En este curso nos centraremos principalmente en los métodos de cuadratura, tanto de interpolación como [gaussiana](https://en.wikipedia.org/wiki/Gaussian_quadrature), como dos ejemplos de dichos algoritmos.
#
# El problema a tratar en este capítulo es la solución aproximada de la función
#
# <a id='Ec5_8'></a>
# \begin{equation*}
# \begin{split}
# I = \int_a^b f(x) dx
# \end{split}
# \label{eq:Ec5_8} \tag{5.8}
# \end{equation*}
#
# [Volver a la Tabla de Contenido](#TOC)
# ### Fórmulas de integración de *Newton - Cotes*
# La idea básica en la integración numérica es cambiar una función difícil de integrar, $f(x)$, dada por la ecuación [(5.8)](#Ec5_8), por una función más simple, $p_n(x)$,
#
# <a id='Ec5_9'></a>
# \begin{equation*}
# \begin{split}
# \widetilde{I} \approx \int_{a=x_0}^{b=x_n} p_{n}(x) dx
# \end{split}
# \label{eq:Ec5_9} \tag{5.9}
# \end{equation*}
#
# Cabe resaltar que en integración numérica no se conocerá la función a integrar, solo se dispondrá de una serie de $n+1$ puntos $(x_i, y_i), i = 0, 1, 2, \ldots, n$, y a partir de ellos se construye un polinomio interpolante de grado $n$, $p_n$, entre los valores de los límites de integración $a = x_0$ y $b=x_n$. $p_n(x)$ es un polinomio de interpolación de la forma
#
# <a id='Ec5_10'></a>
# \begin{equation*}
# \begin{split}
# p_n(x)=a_0+a_1x+a_2x^2+\ldots+a_{n-1}x^{n-1}+a_nx^n
# \end{split}
# \label{eq:Ec5_10} \tag{5.10}
# \end{equation*}
#
# Las fórmulas de integración de [*Newton - Cotes*](https://en.wikipedia.org/wiki/Newton%E2%80%93Cotes_formulas), también llamadas de <a id='Quadrature'></a>[cuadratura](https://en.wikipedia.org/wiki/Quadrature_(mathematics)), son un grupo de fórmulas de integración numérica de tipo interpolación, evaluando la función en puntos equidistantes, para determinar un valor aproximado de la integral. Si no se tienen puntos espaciados, otros métodos deben ser usados, como por ejemplo cuadratura gaussiana, que se verá al final del capítulo.
#
# La forma general de las fórmulas de Newton - Cotes está dada por la función:
#
# <a id='Ec5_11'></a>
# \begin{equation*}
# \begin{split}
# p_n(x)=\sum \limits_{i=0}^n f(x_i)L_{in}(x)
# \end{split}
# \label{eq:Ec5_11} \tag{5.11}
# \end{equation*}
#
# donde
#
# <a id='Ec5_12'></a>
# \begin{equation*}
# \begin{split}
# L_{in}(x)=\frac{(x-x_0)\ldots(x-x_{i-1})(x-x_{i+1})\ldots(x-x_n)}{(x_i-x_0)\ldots(x_i-x_{i-1})(x_i-x_{i+1})\ldots(x_i-x_n)}
# \end{split}
# \label{eq:Ec5_12} \tag{5.12}
# \end{equation*}
#
# es el polinomio de Lagrange, de donde se deduce que:
#
# <a id='Ec5_13'></a>
# \begin{equation*}
# \begin{split}
# \int_a^b p(x)dx=(b-a)\sum \limits_{i=0}^n f(x_i) \frac{1}{(b-a)} \int_a^b L_{in}(x)dx
# \end{split}
# \label{eq:Ec5_13} \tag{5.13}
# \end{equation*}
#
# entonces,
#
# <a id='Ec5_14'></a>
# \begin{equation*}
# \begin{split}
# \int_a^b f(x)dx \approx \int_a^b p(x)dx=(b-a)\sum \limits_{i=0}^n w_if(x_i)
# \end{split}
# \label{eq:Ec5_14} \tag{5.14}
# \end{equation*}
#
# donde los pesos, $w_i$ de la función son representados por
#
# <a id='Ec5_15'></a>
# \begin{equation*}
# \begin{split}
# w_i=\frac{1}{(b-a)} \int_a^b L_{in}(x)dx
# \end{split}
# \label{eq:Ec5_15} \tag{5.15}
# \end{equation*}
#
# A partir de esta idea se obtienen los diferentes esquemas de integración numérica de *Newton - Cotes*
# [Volver a la Tabla de Contenido](#TOC)
# ### Regla trapezoidal
# #### Regla trapezoidal de aplicación simple
# La [regla trapezoidal](https://en.wikipedia.org/wiki/Trapezoidal_rule) emplea una aproximación de la función mediante una línea recta
#
# <p float="center">
# <img src="https://github.com/carlosalvarezh/Analisis_Numerico/blob/master/images/C05_Img03_TrapezoidalRule.PNG?raw=true" width="250" />
# </p>
#
# <div style="text-align: right"> Fuente: <a href="https://upload.wikimedia.org/wikipedia/commons/4/40/Trapezoidal_rule_illustration.svg">wikipedia.com</a> </div>
#
# y corresponde al caso en el que el polinomio en la ecuación [(5.11)](#Ec5_11) es de primer orden
#
#
# \begin{equation*}
# \begin{split}
# I=\int_{a}^{b}f(x)dx \approx \int_a^b \left[ f(a) + \frac{f(b)-f(a)}{b-a}(x-a)\right]dx
# = (b-a)\frac{f(a)+f(b)}{2}
# \end{split}
# \label{eq:Ec5_16} \tag{5.16}
# \end{equation*}
#
# Geométricamente, es equivalente a aproximar el área del trapezoide bajo la línea recta que conecta $f(a)$ y $f(b)$. La integral se representa como:
#
# $$I ≈ \text{ancho} \times \text{altura promedio}$$
#
# El error en la regla trapezoidal simple se puede determinar como:
#
# \begin{equation*}
# \begin{split}
# E_t=-\frac{1}{12}f''(\xi)(b-a)^3
# \end{split}
# \label{eq:Ec5_17} \tag{5.17}
# \end{equation*}
#
# [Volver a la Tabla de Contenido](#TOC)
# #### Regla trapezoidal de aplicación múltiple
# Una manera de mejorar la exactitud de la regla trapezoidal es dividir el intervalo de integración de $a$ a $b$ en un número $n$ de segmentos y aplicar el método a cada uno de ellos. Las ecuaciones resultantes son llamadas fórmulas de integración de múltiple aplicación o compuestas.
#
# <p float="center">
# <img src="https://github.com/carlosalvarezh/Analisis_Numerico/blob/master/images/C05_Img04_TrapezoidalRuleMultiple.gif?raw=true" width="350" />
# </p>
#
# <div style="text-align: right"> Fuente: <a href="https://en.wikipedia.org/wiki/Trapezoidal_rule#/media/File:Trapezium2.gif">wikipedia.com</a> </div>
#
# Hay $n+1$ puntos base igualmente espaciados $(x_0, x_1, x_2, \ldots, x_n)$. En consecuencia hay $n$ segmentos de igual anchura: $h = (b–a) / n$. Si $a$ y $b$ son designados como $x_0$ y $x_n$ respectivamente, la integral total se representará como:
#
# \begin{equation*}
# \begin{split}
# I=\int_{x_0}^{x_1}f(x)dx+\int_{x_1}^{x_2}f(x)dx+\int_{x_2}^{x_3}f(x)dx+\ldots+\int_{x_{n-2}}^{x_{n-1}}f(x)dx+\int_{x_{n-1}}^{x_n}f(x)dx
# \end{split}
# \label{eq:Ec5_18} \tag{5.18}
# \end{equation*}
#
# Al sustituir la regla trapezoidal simple en cada integrando, se tiene
#
# \begin{equation*}
# \begin{split}
# I\approx \left(f(x_0)+f(x_1)\right)\frac{h}{2}+\left(f(x_1)+f(x_2)\right)\frac{h}{2}+\left(f(x_2)+f(x_3)\right)\frac{h}{2}+\ldots\left(f(x_{n-2})+f(x_{n-1})\right)\frac{h}{2}+\left(f(x_{n-1})+f(x_n)\right)\frac{h}{2}
# \end{split}
# \label{eq:Ec5_19} \tag{5.19}
# \end{equation*}
#
# ahora, agrupando términos
#
# \begin{equation*}
# \begin{split}
# I\approx \frac{h}{2}\left[ f(x_0) + 2\sum_{i=1}^{n-1}f(x_i)+f(x_n) \right]
# \end{split}
# \label{eq:Ec5_20} \tag{5.20}
# \end{equation*}
#
# donde $h=(b-a)/n$
# [Volver a la Tabla de Contenido](#TOC)
# #### Implementación computacional
import numpy as np
import matplotlib.pyplot as plt
def trapezoidal(x):
n = len(x)
h = (x[-1] - x[0]) / n
suma = 0
for i in range(1, n-1):
suma += funcion(x[i])
return h * (funcion(x[0]) + 2 * suma + funcion(x[-1])) / 2
def funcion(x):
return 4 / (1 + x**2)
a = 0
b = 1
n = 2
x = np.linspace(a, b, n+1)
I = trapezoidal(x)
I
# [Volver a la Tabla de Contenido](#TOC)
# #### Error en la aplicación de la regla trapezoidal
# Recordando que estos esquemas provienen de la serie truncada de Taylor, el error se puede obtener determinando el primer término truncado en el esquema, que para la regla trapezoidal de aplicación simple corresponde a:
#
# \begin{equation*}
# \begin{split}
# E_t=-\frac{1}{12}f''(\xi)(b-a)^3
# \end{split}
# \label{eq:Ec5_21} \tag{5.21}
# \end{equation*}
#
# donde $f''(\xi)$ es la segunda derivada en el punto $\xi$ en el intervalo $[a,b]$, y $\xi$ es un valor que maximiza la evaluación de esta segunda derivada.
#
# Generalizando este concepto a la aplicación múltiple de la regla trapezoidal, se pueden sumar cada uno de los errores en cada segmento para dar:
#
# \begin{equation*}
# \begin{split}
# E_t=-\frac{(b-a)^3}{12n^3}\sum\limits_{i=1}^n f''(\xi_i)
# \end{split}
# \label{eq:Ec5_22} \tag{5.22}
# \end{equation*}
#
# el anterior resultado se puede simplificar estimando la media, o valor promedio, de la segunda derivada para todo el intervalo
#
# <a id='Ec5_23'></a>
# \begin{equation*}
# \begin{split}
# \bar{f''} \approx \frac{\sum \limits_{i=1}^n f''(\xi_i)}{n}
# \end{split}
# \label{eq:Ec5_23} \tag{5.23}
# \end{equation*}
#
# de esta ecuación se tiene que $\sum f''(\xi_i)\approx nf''$, y reemplazando en la ecuación [(5.23)](#Ec5_23)
#
# \begin{equation*}
# \begin{split}
# E_t \approx \frac{(b-a)^3}{12n^2}\bar{f''}
# \end{split}
# \label{eq:Ec5_24} \tag{5.24}
# \end{equation*}
#
# De este resultado se observa que si se duplica el número de segmentos, el error de truncamiento se disminuirá a una cuarta parte.
# [Volver a la Tabla de Contenido](#TOC)
# ### Reglas de Simpson
# Las [reglas de Simpson](https://en.wikipedia.org/wiki/Simpson%27s_rule) son esquemas de integración numérica en honor al matemático [*Thomas Simpson*](https://en.wikipedia.org/wiki/Thomas_Simpson), utilizado para obtener la aproximación de la integral empleando interpolación polinomial sustituyendo a $f(x)$.
#
# [Volver a la Tabla de Contenido](#TOC)
# #### Regla de Simpson1/3 de aplicación simple
# La primera regla corresponde a una interpolación polinomial de segundo orden sustituida en la ecuación [(5.8)](#Ec5_8)
#
# <p float="center">
# <img src="https://github.com/carlosalvarezh/Analisis_Numerico/blob/master/images/C05_Img05_SimpsonRule13.PNG?raw=true" width="350" />
# </p>
#
# <div style="text-align: right"> Fuente: <a href="https://upload.wikimedia.org/wikipedia/commons/c/ca/Simpsons_method_illustration.svg">wikipedia.com</a> </div>
#
#
# \begin{equation*}
# \begin{split}
# I=\int_a^b f(x)dx \approx \int_a^b p_2(x)dx
# \end{split}
# \label{eq:Ec5_25} \tag{5.25}
# \end{equation*}
#
# del esquema de interpolación de Lagrange para un polinomio de segundo grado, visto en el capitulo anterior, y remplazando en la integral arriba, se llega a
#
# \begin{equation*}
# \begin{split}
# I\approx\int_{x0}^{x2} \left[\frac{(x-x_1)(x-x_2)}{(x_0-x_1)(x_0-x_2)}f(x_0)+\frac{(x-x_0)(x-x_2)}{(x_1-x_0)(x_1-x_2)}f(x_1)+\frac{(x-x_0)(x-x_1)}{(x_2-x_0)(x_2-x_1)}f(x_2)\right]dx
# \end{split}
# \label{eq:Ec5_26} \tag{5.26}
# \end{equation*}
#
# realizando la integración de forma analítica y un manejo algebraico, resulta
#
# \begin{equation*}
# \begin{split}
# I\approx\frac{h}{3} \left[ f(x_0)+4f(x_1)+f(x_2)\right]
# \end{split}
# \label{eq:Ec5_27} \tag{5.27}
# \end{equation*}
#
# donde $h=(b-a)/2$ y los $x_{i+1} = x_i + h$
# A continuación, vamos a comparar graficamente las funciones "exacta" (con muchos puntos) y una aproximada empleando alguna técnica de interpolación para $n=3$ puntos (Polinomio interpolante de orden $2$).
# +
from scipy.interpolate import barycentric_interpolate
# usaremos uno de los tantos métodos de interpolación dispobibles en las bibliotecas de Python
n = 3 # puntos a interpolar para un polinomio de grado 2
xp = np.linspace(a,b,n) # generación de n puntos igualmente espaciados para la interpolación
fp = funcion(xp) # evaluación de la función en los n puntos generados
x = np.linspace(a, b, 100) # generación de 100 puntos igualmente espaciados
y = barycentric_interpolate(xp, fp, x) # interpolación numérica empleando el método del Baricentro
fig = plt.figure(figsize=(9, 6), dpi= 80, facecolor='w', edgecolor='k')
ax = fig.add_subplot(111)
l, = plt.plot(x, y)
plt.plot(x, funcion(x), '-', c='red')
plt.plot(xp, fp, 'o', c=l.get_color())
plt.annotate('Función "Real"', xy=(.63, 1.5), xytext=(0.8, 1.25),arrowprops=dict(facecolor='black', shrink=0.05),)
plt.annotate('Función interpolada', xy=(.72, 1.75), xytext=(0.4, 2),arrowprops=dict(facecolor='black', shrink=0.05),)
plt.grid(True) # muestra la malla de fondo
plt.show() # muestra la gráfica
# -
# Se observa que hay una gran diferencia entre las áreas que se estarían abarcando en la función llamada "*real*" (que se emplearon $100$ puntos para su generación) y la función *interpolada* (con únicamente $3$ puntos para su generación) que será la empleada en la integración numérica (aproximada) mediante la regla de *Simpson $1/3$*.
#
# Conscientes de esto, procederemos entonces a realizar el cálculo del área bajo la curva del $p_3(x)$ empleando el método de *Simpson $1/3$*
# Creemos un programa en *Python* para que nos sirva para cualquier función $f(x)$ que queramos integrar en cualquier intervalo $[a,b]$ empleando la regla de integración de *Simpson $1/3$*:
# se ingresan los valores del intervalo [a,b]
a = float(input('Ingrese el valor del límite inferior: '))
b = float(input('Ingrese el valor del límite superior: '))
# +
# cuerpo del programa por la regla de Simpson 1/3
h = (b-a)/2 # cálculo del valor de h
x0 = a # valor del primer punto para la fórmula de S1/3
x1 = x0 + h # Valor del punto intermedio en la fórmula de S1/3
x2 = b # valor del tercer punto para la fórmula de S1/3
fx0 = funcion(x0) # evaluación de la función en el punto x0
fx1 = funcion(x1) # evaluación de la función en el punto x1
fx2 = funcion(x2) # evaluación de la función en el punto x2
int_S13 = h / 3 * (fx0 + 4*fx1 + fx2)
#erel = np.abs(exacta - int_S13) / exacta * 100
print('el valor aproximado de la integral por la regla de Simpson1/3 es: ', int_S13, '\n')
#print('el error relativo entre el valor real y el calculado es: ', erel,'%')
# -
# [Volver a la Tabla de Contenido](#TOC)
# #### Error en la regla de Simpson 1/3 de aplicación simple
# El problema de calcular el error de esta forma es que realmente no conocemos el valor exacto. Para poder calcular el error al usar la regla de *Simpson 1/3*:
#
# \begin{equation*}
# \begin{split}
# -\frac{h^5}{90}f^{(4)}(\xi)
# \end{split}
# \label{eq:Ec5_28} \tag{5.28}
# \end{equation*}
#
# será necesario derivar cuatro veces la función original: $f(x)=e^{x^2}$. Para esto, vamos a usar nuevamente el cálculo simbólico (siempre deben verificar que la respuesta obtenida es la correcta!!!):
from sympy import *
x = symbols('x')
# Derivamos cuatro veces la función $f(x)$ con respecto a $x$:
deriv4 = diff(4 / (1 + x**2),x,4)
deriv4
# y evaluamos esta función de la cuarta derivada en un punto $0 \leq \xi \leq 1$. Como la función $f{^{(4)}}(x)$ es creciente en el intervalo $[0,1]$ (compruébelo gráficamente y/o por las técnicas vistas en cálculo diferencial), entonces, el valor que hace máxima la cuarta derivada en el intervalo dado es:
x0 = 1.0
evald4 = deriv4.evalf(subs={x: x0})
print('El valor de la cuarta derivada de f en x0={0:6.2f} es {1:6.4f}: '.format(x0, evald4))
# Calculamos el error en la regla de *Simpson$1/3$*
errorS13 = abs(h**5*evald4/90)
print('El error al usar la regla de Simpson 1/3 es: {0:6.6f}'.format(errorS13))
# Entonces, podemos expresar el valor de la integral de la función $f(x)=e^{x^2}$ en el intervalo $[0,1]$ usando la *Regla de Simpson $1/3$* como:
#
# <div class="alert alert-block alert-warning">
# $$\color{blue}{\int_0^1 \frac{4}{1 + x^2}dx} = \color{green}{3,133333} \color{red}{+ 0.004167}$$
# </div>
# Si lo fuéramos a hacer "a mano" $\ldots$ aplicando la fórmula directamente, con los siguientes datos:
#
# $h = \frac{(1.0 - 0.0)}{2.0} = 0.5$
#
# $x_0 = 0.0$
#
# $x_1 = 0.5$
#
# $x_2 = 1.0$
#
# $f(x) = \frac{4}{1 + x^2}$
#
# sustituyendo estos valores en la fórmula dada:
#
#
# $\int_0^1\frac{4}{1 + x^2}dx \approx \frac{0.5}{3} \left[f(0)+4f(0.5)+f(1)\right]$
#
# $\int_0^1\frac{4}{1 + x^2}dx \approx \frac{0.5}{3} \left[ \frac{4}{1 + 0^2} + 4\frac{4}{1 + 0.5^2} + \frac{4}{1 + 1^2} \right] \approx 3.133333$
# [Volver a la Tabla de Contenido](#TOC)
# #### Regla de simpson1/3 de aplicación múltiple
# Al igual que en la regla Trapezoidal, las reglas de Simpson también cuentan con un esquema de aplicación múltiple (llamada también compuesta). Supongamos que se divide el intervalo $[a,b]$ se divide en $n$ sub intervalos, con $n$ par, quedando la integral
#
# \begin{equation*}
# \begin{split}
# I=\int_{x_0}^{x_2}f(x)dx+\int_{x_2}^{x_4}f(x)dx+\ldots+\int_{x_{n-2}}^{x_n}f(x)dx
# \end{split}
# \label{eq:Ec5_29} \tag{5.29}
# \end{equation*}
#
# y sustituyendo en cada una de ellas la regla de Simpson1/3, se llega a
#
# \begin{equation*}
# \begin{split}
# I \approx 2h\frac{f(x_0)+4f(x_1)+f(x_2)}{6}+2h\frac{f(x_2)+4f(x_3)+f(x_4)}{6}+\ldots+2h\frac{f(x_{n-2})+4f(x_{n-1})+f(x_n)}{6}
# \end{split}
# \label{eq:Ec5_30} \tag{5.30}
# \end{equation*}
#
#
# entonces la regla de Simpson compuesta (o de aplicación múltiple) se escribe como:
#
# \begin{equation*}
# \begin{split}
# I=\int_a^bf(x)dx\approx \frac{h}{3}\left[f(x_0) + 2 \sum \limits_{j=1}^{n/2-1} f(x_{2j}) + 4 \sum \limits_{j=1}^{n/2} f(x_{2j-1})+f(x_n)\right]
# \end{split}
# \label{eq:Ec5_31} \tag{5.31}
# \end{equation*}
#
# donde $x_j=a+jh$ para $j=0,1,2, \ldots, n-1, n$ con $h=(b-a)/n$, $x_0=a$ y $x_n=b$.
# [Volver a la Tabla de Contenido](#TOC)
# #### Implementación computacional regla de Simpson1/3 de aplicación múltiple
# [Volver a la Tabla de Contenido](#TOC)
# #### Regla de Simpson 3/8 de aplicación simple
# Resulta cuando se sustituye la función $f(x)$ por una interpolación de tercer orden:
#
# \begin{equation*}
# \begin{split}
# I=\int_{a}^{b}f(x)dx = \frac{3h}{8}\left[ f(x_0)+3f(x_1)+3f(x_2)+f(x_3) \right]
# \end{split}
# \label{eq:Ec5_32} \tag{5.32}
# \end{equation*}
#
# Realizando un procedimiento similar al usado para la regla de *Simpson $1/3$*, pero esta vez empleando $n=4$ puntos:
# +
# usaremos uno de los tantos métodos de interpolación dispobibles en las bibliotecas de Python
n = 4 # puntos a interpolar para un polinomio de grado 2
xp = np.linspace(0,1,n) # generación de n puntos igualmente espaciados para la interpolación
fp = funcion(xp) # evaluación de la función en los n puntos generados
x = np.linspace(0, 1, 100) # generación de 100 puntos igualmente espaciados
y = barycentric_interpolate(xp, fp, x) # interpolación numérica empleando el método del Baricentro
fig = plt.figure(figsize=(9, 6), dpi= 80, facecolor='w', edgecolor='k')
ax = fig.add_subplot(111)
l, = plt.plot(x, y)
plt.plot(x, funcion(x), '-', c='red')
plt.plot(xp, fp, 'o', c=l.get_color())
plt.annotate('"Real"', xy=(.63, 1.5), xytext=(0.8, 1.25),arrowprops=dict(facecolor='black', shrink=0.05),)
plt.annotate('Interpolación', xy=(.72, 1.75), xytext=(0.4, 2),arrowprops=dict(facecolor='black', shrink=0.05),)
plt.grid(True) # muestra la malla de fondo
plt.show() # muestra la gráfica
# +
# cuerpo del programa por la regla de Simpson 3/8
h = (b - a) / 3 # cálculo del valor de h
int_S38 = 3 * h / 8 * (funcion(a) + 3*funcion(a + h) + 3*funcion(a + 2*h) + funcion(a + 3*h) + 3 * funcion(a+4*h)/8)
erel = np.abs(np.pi - int_S38) / np.pi * 100
print('el valor aproximado de la integral utilizando la regla de Simpson 3/8 es: ', int_S38, '\n')
print('el error relativo entre el valor real y el calculado es: ', erel,'%')
# -
# Para poder calcular el error al usar la regla de *Simpson 3/8*:
#
# <div class="alert alert-block alert-warning">
# $$\color{red}{-\frac{3h^5}{80}f^{(4)}(\xi)}$$
# </div>
#
# será necesario derivar cuatro veces la función original. Para esto, vamos a usar nuevamente el cálculo simbólico (siempre deben verificar que la respuesta obtenida es la correcta!!!):
errorS38 = 3*h**5*evald4/80
print('El error al usar la regla de Simpson 3/8 es: ',errorS38)
# Entonces, podemos expresar el valor de la integral de la función $f(x)=e^{x^2}$ en el intervalo $[0,1]$ usando la *Regla de Simpson $3/8$* como:
#
# <div class="alert alert-block alert-warning">
# $$\color{blue}{\int_0^1\frac{4}{1 + x^2}dx} = \color{green}{3.138462} \color{red}{- 0.001852}$$
# </div>
# Aplicando la fórmula directamente, con los siguientes datos:
#
# $h = \frac{(1.0 - 0.0)}{3.0} = 0.33$
#
# $x_0 = 0.0$, $x_1 = 0.33$, $x_2 = 0.66$, $x_3 = 1.00$
#
# $f(x) = \frac{4}{1 + x^2}$
#
# sustituyendo estos valores en la fórmula dada:
#
# $\int_0^1\frac{4}{1 + x^2}dx \approx \frac{3\times0.3333}{8} \left[ \frac{4}{1 + 0^2} + 3\frac{4}{1 + 0.3333^2} +3\frac{4}{1 + 0.6666^2} + \frac{4}{1 + 1^2} \right] \approx 3.138462$
#
#
# Esta sería la respuesta si solo nos conformamos con lo que podemos hacer usando word...
# [Volver a la Tabla de Contenido](#TOC)
# #### Regla de Simpson3/8 de aplicación múltiple
# Dividiendo el intervalo $[a,b]$ en $n$ sub intervalos de longitud $h=(b-a)/n$, con $n$ múltiplo de 3, quedando la integral
#
# \begin{equation*}
# \begin{split}
# I=\int_{x_0}^{x_3}f(x)dx+\int_{x_3}^{x_6}f(x)dx+\ldots+\int_{x_{n-3}}^{x_n}f(x)dx
# \end{split}
# \label{eq:Ec5_33} \tag{5.33}
# \end{equation*}
#
# sustituyendo en cada una de ellas la regla de Simpson3/8, se llega a
#
# \begin{equation*}
# \begin{split}
# I=\int_a^bf(x)dx\approx \frac{3h}{8}\left[f(x_0) + 3 \sum \limits_{i=0}^{n/3-1} f(x_{3i+1}) + 3 \sum \limits_{i=0}^{n/3-1}f(x_{3i+2})+2 \sum \limits_{i=0}^{n/3-2} f(x_{3i+3})+f(x_n)\right]
# \end{split}
# \label{eq:Ec5_34} \tag{5.34}
# \end{equation*}
#
# donde en cada sumatoria se deben tomar los valores de $i$ cumpliendo que $i=i+3$.
# [Volver a la Tabla de Contenido](#TOC)
# #### Implementación computacional de la regla de Simpson3/8 de aplicación múltiple
# +
#
# -
# [Volver a la Tabla de Contenido](#TOC)
# ### Cuadratura de Gauss
# #### Introducción
# Retomando la idea inicial de los esquemas de [cuadratura](#Quadrature), el valor de la integral definida se estima de la siguiente manera:
#
# <a id='Ec5_35'></a>
# \begin{equation*}
# \begin{split}
# I=\int_a^b f(x)dx \approx \sum \limits_{i=0}^n c_if(x_i)
# \end{split}
# \label{eq:Ec5_35} \tag{5.35}
# \end{equation*}
#
# Hasta ahora hemos visto los métodos de la regla trapezoidal y las reglas de Simpson más empleadas. En estos esquemas, la idea central es la distribución uniforme de los puntos que siguen la regla $x_i=x_0+ih$, con $i=0,1,2, \ldots, n$ y la evaluación de la función en estos puntos.
#
# Supongamos ahora que la restricción de la uniformidad en el espaciamiento de esos puntos fijos no es más considerada y se tiene la libertad de evaluar el área bajo una recta que conecte a dos puntos cualesquiera sobre la curva. Al ubicar estos puntos en forma “inteligente”, se puede definir una línea recta que equilibre los errores negativos y positivos
#
# <p float="center">
# <img src="https://github.com/carlosalvarezh/Analisis_Numerico/blob/master/images/C05_Img06_GQ01.PNG?raw=true" width="750" />
# </p>
#
# <div style="text-align: right"> Fuente: <a href="http://artemisa.unicauca.edu.co/~cardila/Chapra.pdf"><NAME>., <NAME>. Métodos Numéricos para ingenieros, 5a Ed. <NAME>. 2007</a> </div>
#
# De la figura de la derecha, se disponen de los puntos $x_0$ y $x_1$ para evaluar la función $f(x)$. Expresando la integral bajo la curva de forma aproximada dada en la la ecuación ([5.35](#Ec5_35)), y empleando los límites de integración en el intervalo $[-1,1]$ por simplicidad (después se generalizará el concepto a un intervalo $[a,b]$), se tiene
#
# <a id='Ec5_36'></a>
# \begin{equation*}
# \begin{split}
# I=\int_{-1}^1 f(x)dx \approx c_0f(x_0)+c_1f(x_1)
# \end{split}
# \label{eq:Ec5_36} \tag{5.36}
# \end{equation*}
#
# [Volver a la Tabla de Contenido](#TOC)
# #### Determinación de los coeficientes
# se tiene una ecuación con cuatro incógnitas ($c_0, c_1, x_0$ y $x_1$) que se deben determinar. Para ello, supongamos que disponemos de un polinomio de hasta grado 3, $f_3(x)$, de donde podemos construir cuatro ecuaciones con cuatro incógnitas de la siguiente manera:
#
# - $f_3(x)=1$:
#
# <a id='Ec5_37'></a>
# \begin{equation*}
# \begin{split}
# \int_{-1}^1 1dx = c_0 \times 1 + c_1 \times 1 = c_0 + c_1 = 2
# \end{split}
# \label{eq:Ec5_37} \tag{5.37}
# \end{equation*}
#
# - $f_3(x)=x$:
#
# <a id='Ec5_38'></a>
# \begin{equation*}
# \begin{split}
# \int_{-1}^1 xdx = c_0x_0 + c_1x_1 = 0
# \end{split}
# \label{eq:Ec5_38} \tag{5.38}
# \end{equation*}
#
# - $f_3(x)=x^2$:
#
# <a id='Ec5_39'></a>
# \begin{equation*}
# \begin{split}
# \int_{-1}^1 x^2dx = c_0x^2_0 + c_1x^2_1 = \frac{2}{3}
# \end{split}
# \label{eq:Ec5_39} \tag{5.39}
# \end{equation*}
#
# y por último
#
# - $f_3(x)=x^3$:
#
# <a id='Ec5_40'></a>
# \begin{equation*}
# \begin{split}
# \int_{-1}^1 x^3dx = c_0x^3_0 + c_1x^3_1 = 0
# \end{split}
# \label{eq:Ec5_40} \tag{5.40}
# \end{equation*}
#
# resolviendo simultáneamente las dos primeras ecuaciones para $c_0$ y $c_1$ en térm,inos de $x_0$ y $x_1$, se llega a
#
# <a id='Ec5_41'></a>
# \begin{equation*}
# \begin{split}
# c_0=\frac{2x_1}{x_1-x_0}, \quad c_1=-\frac{2x_0}{x_1-x_0}\end{split}
# \label{eq:Ec5_41} \tag{5.41}
# \end{equation*}
#
# reemplazamos estos dos valores en las siguientes dos ecuaciones
#
# <a id='Ec5_42'></a>
# \begin{equation*}
# \begin{split}
# \frac{2}{3}=\frac{2x_0^2x_1}{x_1-x_0}-\frac{2x_0x_1^2}{x_1-x_0}
# \end{split}
# \label{eq:Ec5_42} \tag{5.42}
# \end{equation*}
#
# <a id='Ec5_43'></a>
# \begin{equation*}
# \begin{split}
# 0=\frac{2x_0^3x_1}{x_1-x_0}-\frac{2x_0x_1^3}{x_1-x_0}
# \end{split}
# \label{eq:Ec5_43} \tag{5.43}
# \end{equation*}
#
# de la ecuación ([5.43](#Ec5_43)) se tiene
#
# <a id='Ec5_44'></a>
# \begin{equation*}
# \begin{split}
# x_0^3x_1&=x_0x_1^3 \\
# x_0^2 &= x_1^2
# \end{split}
# \label{eq:Ec5_44} \tag{5.44}
# \end{equation*}
#
# de aquí se tiene que $|x_0|=|x_1|$ (para considerar las raíces negativas recuerde que $\sqrt{a^2}= \pm a = |a|$), y como se asumió que $x_0<x_1$, entonces $x_0<0$ y $x_1>0$ (trabajando en el intervalo $[-1,1]$), llegándose finalmente a que $x_0=-x_1$. Reemplazando este resultado en la ecuación ([5.42](#Ec5_42))
#
# <a id='Ec5_45'></a>
# \begin{equation*}
# \begin{split}
# \frac{2}{3}=2\frac{x_1^3+x_1^3}{2x_1}
# \end{split}
# \label{eq:Ec5_45} \tag{5.45}
# \end{equation*}
#
# despejando, $x_1^2=1/3$, y por último se llega a que
#
# <a id='Ec5_46'></a>
# \begin{equation*}
# \begin{split}
# x_0=-\frac{\sqrt{3}}{3}, \quad x_1=\frac{\sqrt{3}}{3}
# \end{split}
# \label{eq:Ec5_46} \tag{5.46}
# \end{equation*}
#
# reemplazando estos resultados en la ecuación ([5.41](#Ec5_41)) y de la ecuación ([5.37](#Ec5_37)), se tiene que $c_0=c_1=1$. Reescribiendo la ecuación ([5.36](#Ec5_36)) con los valores encontrados se llega por último a:
#
# <a id='Ec5_47'></a>
# \begin{equation*}
# \begin{split}
# I=\int_{-1}^1 f(x)dx &\approx c_0f(x_0)+c_1f(x_1) \\
# &= f \left( \frac{-\sqrt{3}}{3}\right)+f \left( \frac{\sqrt{3}}{3}\right)
# \end{split}
# \label{eq:Ec5_47} \tag{5.47}
# \end{equation*}
#
#
# Esta aproximación realizada es "exacta" para polinomios de grado menor o igual a tres ($3$). La aproximación trapezoidal es exacta solo para polinomios de grado uno ($1$).
#
# ***Ejemplo:*** Calcule la integral de la función $f(x)=x^3+2x^2+1$ en el intervalo $[-1,1]$ empleando tanto las técnicas analíticas como la cuadratura de Gauss vista.
#
#
# - ***Solución analítica (exacta)***
#
# $$\int_{-1}^1 (x^3+2x^2+1)dx=\left.\frac{x^4}{4}+\frac{2x^3}{3}+x \right |_{-1}^1=\frac{10}{3}$$
#
#
# - ***Aproximación numérica por Cuadratura de Gauss***
#
# \begin{equation*}
# \begin{split}
# \int_{-1}^1 (x^3+2x^2+1)dx &\approx1f\left(-\frac{\sqrt{3}}{3} \right)+1f\left(\frac{\sqrt{3}}{3} \right) \\
# &=-\frac{3\sqrt{3}}{27}+\frac{2\times 3}{9}+1+\frac{3\sqrt{3}}{27}+\frac{2\times 3}{9}+1 \\
# &=2+\frac{4}{3} \\
# &= \frac{10}{3}
# \end{split}
# \end{equation*}
#
# [Volver a la Tabla de Contenido](#TOC)
# #### Cambios de los límites de integración
# Obsérvese que los límites de integración de la ecuación ([5.47](#Ec5_47)) son de $-1$ a $1$. Esto se hizo para simplificar las matemáticas y para hacer la formulación tan general como fuera posible. Asumamos ahora que se desea determinar el valor de la integral entre dos límites cualesquiera $a$ y $b$. Supongamos también, que una nueva variable $x_d$ se relaciona con la variable original $x$ de forma lineal,
#
# <a id='Ec5_48'></a>
# \begin{equation*}
# \begin{split}
# x=a_0+a_1x_d
# \end{split}
# \label{eq:Ec5_48} \tag{5.48}
# \end{equation*}
#
# si el límite inferior, $x=a$, corresponde a $x_d=-1$, estos valores podrán sustituirse en la ecuación ([5.48](#Ec5_48)) para obtener
#
# <a id='Ec5_49'></a>
# \begin{equation*}
# \begin{split}
# a=a_0+a_1(-1)
# \end{split}
# \label{eq:Ec5_49} \tag{5.49}
# \end{equation*}
#
# de manera similar, el límite superior, $x=b$, corresponde a $x_d=1$, para dar
#
# <a id='Ec5_50'></a>
# \begin{equation*}
# \begin{split}
# b=a_0+a_1(1)
# \end{split}
# \label{eq:Ec5_50} \tag{5.50}
# \end{equation*}
#
# resolviendo estas ecuaciones simultáneamente,
#
# <a id='Ec5_51'></a>
# \begin{equation*}
# \begin{split}
# a_0=(b+a)/2, \quad a_1=(b-a)/2
# \end{split}
# \label{eq:Ec5_51} \tag{5.51}
# \end{equation*}
#
# sustituyendo en la ecuación ([5.48](#Ec5_48))
#
# <a id='Ec5_52'></a>
# \begin{equation*}
# \begin{split}
# x=\frac{(b+a)+(b-a)x_d}{2}
# \end{split}
# \label{eq:Ec5_52} \tag{5.52}
# \end{equation*}
#
# derivando la ecuación ([5.52](#Ec5_52)),
#
# <a id='Ec5_53'></a>
# \begin{equation*}
# \begin{split}
# dx=\frac{b-a}{2}dx_d
# \end{split}
# \label{eq:Ec5_53} \tag{5.53}
# \end{equation*}
#
# Las ecuacio es ([5.51](#Ec5_51)) y ([5.52](#Ec5_52)) se pueden sustituir para $x$ y $dx$, respectivamente, en la evaluación de la integral. Estas sustituciones transforman el intervalo de integración sin cambiar el valor de la integral. En este caso
#
# <a id='Ec5_54'></a>
# \begin{equation*}
# \begin{split}
# \int_a^b f(x)dx = \frac{b-a}{2} \int_{-1}^1 f \left( \frac{(b+a)+(b-a)x_d}{2}\right)dx_d
# \end{split}
# \label{eq:Ec5_54} \tag{5.54}
# \end{equation*}
#
# Esta integral se puede aproximar como,
#
# <a id='Ec5_55'></a>
# \begin{equation*}
# \begin{split}
# \int_a^b f(x)dx \approx \frac{b-a}{2} \left[f\left( \frac{(b+a)+(b-a)x_0}{2}\right)+f\left( \frac{(b+a)+(b-a)x_1}{2}\right) \right]
# \end{split}
# \label{eq:Ec5_55} \tag{5.55}
# \end{equation*}
# [Volver a la Tabla de Contenido](#TOC)
# #### Fórmulas de punto superior
# La fórmula anterior para la cuadratura de Gauss era de dos puntos. Se pueden desarrollar versiones de punto superior en la forma general:
#
# <a id='Ec5_56'></a>
# \begin{equation*}
# \begin{split}
# I \approx c_0f(x_0) + c_1f(x_1) + c_2f(x_2) +\ldots+ c_{n-1}f(x_{n-1})
# \end{split}
# \label{eq:Ec5_56} \tag{5.56}
# \end{equation*}
#
# con $n$, el número de puntos.
#
# Debido a que la cuadratura de Gauss requiere evaluaciones de la función en puntos espaciados uniformemente dentro del intervalo de integración, no es apropiada para casos donde se desconoce la función. Si se conoce la función, su ventaja es decisiva.
#
# En la siguiente tabla se presentan los valores de los parámertros para $1, 2, 3, 4$ y $5$ puntos.
#
# |$$n$$ | $$c_i$$ | $$x_i$$ |
# |:----:|:----------:|:-------------:|
# |$$1$$ |$$2.000000$$| $$0.000000$$ |
# |$$2$$ |$$1.000000$$|$$\pm0.577350$$|
# |$$3$$ |$$0.555556$$|$$\pm0.774597$$|
# | |$$0.888889$$| $$0.000000$$ |
# |$$4$$ |$$0.347855$$|$$\pm0.861136$$|
# | |$$0.652145$$|$$\pm0.339981$$|
# |$$5$$ |$$0.236927$$|$$\pm0.906180$$|
# | |$$0.478629$$|$$\pm0.538469$$|
# | |$$0.568889$$| $$0.000000$$ |
#
import numpy as np
import pandas as pd
GaussTable = [[[0], [2]], [[-1/np.sqrt(3), 1/np.sqrt(3)], [1, 1]], [[-np.sqrt(3/5), 0, np.sqrt(3/5)], [5/9, 8/9, 5/9]], [[-0.861136, -0.339981, 0.339981, 0.861136], [0.347855, 0.652145, 0.652145, 0.347855]], [[-0.90618, -0.538469, 0, 0.538469, 0.90618], [0.236927, 0.478629, 0.568889, 0.478629, 0.236927]], [[-0.93247, -0.661209, -0.238619, 0.238619, 0.661209, 0.93247], [0.171324, 0.360762, 0.467914, 0.467914, 0.360762, 0.171324]]]
display(pd.DataFrame(GaussTable, columns=["Integration Points", "Corresponding Weights"]))
def IG(f, n):
n = int(n)
return sum([GaussTable[n - 1][1][i]*f(GaussTable[n - 1][0][i]) for i in range(n)])
def f(x): return x**9 + x**8
IG(f, 5.0)
# [Volver a la Tabla de Contenido](#TOC)
# #### Ejemplo Cuadratura de Gauss
# Determine el valor aproximado de:
#
# $$\int_0^1 \frac{4}{1+x^2}dx$$
#
# empleando cuadratura gaussiana de dos puntos.
#
# Reemplazando los parámetros requeridos en la ecuación ([5.55](#Ec5_55)), donde $a=0$, $b=1$, $x_0=-\sqrt{3}/3$ y $x_1=\sqrt{3}/3$
#
# \begin{equation*}
# \begin{split}
# \int_0^1 f(x)dx &\approx \frac{1-0}{2} \left[f\left( \frac{(1+0)+(1-0)\left(-\frac{\sqrt{3}}{3}\right)}{2}\right)+f\left( \frac{(1+0)+(1-0)\left(\frac{\sqrt{3}}{3}\right)}{2}\right) \right]\\
# &= \frac{1}{2} \left[f\left( \frac{1-\frac{\sqrt{3}}{3}}{2}\right)+f\left( \frac{1+\frac{\sqrt{3}}{3}}{2}\right) \right]\\
# &= \frac{1}{2} \left[ \frac{4}{1 + \left( \frac{1-\frac{\sqrt{3}}{3}}{2} \right)^2}+\frac{4}{1 + \left( \frac{1+\frac{\sqrt{3}}{3}}{2} \right)^2} \right]\\
# &=3.147541
# \end{split}
# \end{equation*}
#
# Ahora veamos una breve implementación computacional
import numpy as np
def fxG(a, b, x):
xG = ((b + a) + (b - a) * x) / 2
return funcion(xG)
def GQ2(a,b):
c0 = 1.0
c1 = 1.0
x0 = -1.0 / np.sqrt(3)
x1 = 1.0 / np.sqrt(3)
return (b - a) / 2 * (c0 * fxG(a,b,x0) + c1 * fxG(a,b,x1))
print(GQ2(0,1))
# [Volver a la Tabla de Contenido](#TOC)
from IPython.core.display import HTML
def css_styling():
styles = open('./nb_style.css', 'r').read()
return HTML(styles)
css_styling()
|
Cap05_IntegracionNumerica.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PaddlePaddle 2.1.0 (Python 3.5)
# language: python
# name: py35-paddle1.2.0
# ---
# The following additional libraries are needed to run this
# notebook. Note that running on Colab is experimental, please report a Github
# issue if you have any problem.
# !pip install git+https://github.com/d2l-ai/d2l-zh@release # installing d2l
# # 7.2. 使用块的网络(VGG)
# :label:`sec_vgg`
#
# 虽然 AlexNet 证明深层神经网络卓有成效,但它没有提供一个通用的模板来指导后续的研究人员设计新的网络。
# 在下面的几个章节中,我们将介绍一些常用于设计深层神经网络的启发式概念。
#
# 与芯片设计中工程师从放置晶体管到逻辑元件再到逻辑块的过程类似,神经网络结构的设计也逐渐变得更加抽象。研究人员开始从单个神经元的角度思考问题,发展到整个层次,现在又转向模块,重复各层的模式。
#
# 使用块的想法首先出现在牛津大学的 [视觉几何组(visualgeometry Group)](http://www.robots.ox.ac.uk/~vgg/) (VGG)的 *VGG网络* 中。通过使用循环和子程序,可以很容易地在任何现代深度学习框架的代码中实现这些重复的结构。
#
# ## (**7.2.1. VGG块**)
#
# 经典卷积神经网络的基本组成部分是下面的这个序列:
# 1. 带填充以保持分辨率的卷积层;
# 1. 非线性激活函数,如ReLU;
# 1. 池化层,如最大池化层。
#
# 而一个 VGG 块与之类似,由一系列卷积层组成,后面再加上用于空间下采样的最大池化层。在最初的 VGG 论文 :cite:`Simonyan.Zisserman.2014` 中,作者使用了带有 $3\times3$ 卷积核、填充为 1(保持高度和宽度)的卷积层,和带有 $2 \times 2$ 池化窗口、步幅为 2(每个块后的分辨率减半)的最大池化层。在下面的代码中,我们定义了一个名为 `vgg_block` 的函数来实现一个 VGG 块。
#
# 该函数有三个参数,分别对应于卷积层的数量 `num_convs`、输入通道的数量 `in_channels`
# 和输出通道的数量 `out_channels`.
#
# +
import sys
import time
import paddle
from paddle import nn, optimizer
def vgg_block(num_convs, in_channels, out_channels):
blk = []
for i in range(num_convs):
if i == 0:
blk.append(nn.Conv2D(in_channels, out_channels, kernel_size=3, padding=1))
else:
blk.append(nn.Conv2D(out_channels, out_channels, kernel_size=3, padding=1))
blk.append(nn.ReLU())
blk.append(nn.MaxPool2D(kernel_size=2, stride=2))
return nn.Sequential(*blk)
# -
# ## [**7.2.2. VGG网络**]
#
# 与 AlexNet、LeNet 一样,VGG 网络可以分为两部分:第一部分主要由卷积层和池化层组成,第二部分由全连接层组成。如 :numref:`fig_vgg` 中所示。
#
# 
# :width:`400px`
# :label:`fig_vgg`
#
#
# VGG神经网络连续连接 :numref:`fig_vgg` 的几个 VGG 块(在 `vgg_block` 函数中定义)。其中有超参数变量 `conv_arch` 。该变量指定了每个VGG块里卷积层个数和输出通道数。全连接模块则与AlexNet中的相同。
#
# 原始 VGG 网络有 5 个卷积块,其中前两个块各有一个卷积层,后三个块各包含两个卷积层。
# 第一个模块有 64 个输出通道,每个后续模块将输出通道数量翻倍,直到该数字达到 512。由于该网络使用 8 个卷积层和 3 个全连接层,因此它通常被称为 VGG-11。
#
conv_arch = ((1, 1, 64), (1, 64, 128), (2, 128, 256), (2, 256, 512), (2, 512, 512))
fc_features = 512 * 7 * 7 # 根据卷积层的输出算出来的
fc_hidden_units = 4096 # 任意
# 下面的代码实现了 VGG-11。可以通过在 `conv_arch` 上执行 for 循环来简单实现。
#
# +
class FlattenLayer(nn.Layer):
def __init__(self):
super(FlattenLayer, self).__init__()
def forward(self, x): # x shape: (batch, *, *, ...)
return x.reshape((x.shape[0], -1))
def vgg(conv_arch, fc_features, fc_hidden_units=4096):
net = nn.Sequential()
# 卷积层部分
for i, (num_convs, in_channels, out_channels) in enumerate(conv_arch):
net.add_sublayer("vgg_block_" + str(i+1), vgg_block(num_convs, in_channels, out_channels))
# 全连接层部分
net.add_sublayer("fc", nn.Sequential(
FlattenLayer(),
nn.Linear(fc_features, fc_hidden_units),
nn.ReLU(),
nn.Dropout(0.5),
nn.Linear(fc_hidden_units, fc_hidden_units),
nn.ReLU(),
nn.Dropout(0.5),
nn.Linear(fc_hidden_units, 10)
))
return net
# -
# 接下来,我们将构建一个高度和宽度为 224 的单通道数据样本,以[**观察每个层输出的形状**]。
#
# +
net = vgg(conv_arch, fc_features, fc_hidden_units)
X = paddle.rand((1, 1, 224, 224))
# named_children获取一级子模块及其名字(named_modules会返回所有子模块,包括子模块的子模块)
for name, blk in net.named_children():
X = blk(X)
print(name, 'output shape: ', X.shape)
# -
# 正如你所看到的,我们在每个块的高度和宽度减半,最终高度和宽度都为7。最后再展平表示,送入全连接层处理。
#
# ## 7.2.3. 训练模型
#
# [**由于VGG-11比AlexNet计算量更大,因此我们构建了一个通道数较少的网络**],足够用于训练Fashion-MNIST数据集。
#
ratio = 8
small_conv_arch = [(1, 1, 64//ratio), (1, 64//ratio, 128//ratio), (2, 128//ratio, 256//ratio),
(2, 256//ratio, 512//ratio), (2, 512//ratio, 512//ratio)]
net = vgg(small_conv_arch, fc_features // ratio, fc_hidden_units // ratio)
# 除了使用略高的学习率外,[**模型训练**]过程与 :numref:`sec_alexnet` 中的 AlexNet 类似。
#
# +
import paddle.vision.datasets as datasets
import paddle.vision.transforms as transforms
import sys
def load_data_fashion_mnist(batch_size, resize=None):
"""Download the fashion mnist dataset and then load into memory."""
trans = []
if resize:
trans.append(transforms.Resize(size=resize))
trans.append(transforms.ToTensor())
transform = transforms.Compose(trans)
mnist_train = datasets.FashionMNIST(mode='train', download=True, transform=transform)
mnist_test = datasets.FashionMNIST(mode='test', download=True, transform=transform)
if sys.platform.startswith('win'):
num_workers = 0 # 0表示不用额外的进程来加速读取数据
else:
num_workers = 0
train_iter = paddle.io.DataLoader(mnist_train, batch_size=batch_size, shuffle=True, num_workers=num_workers)
test_iter = paddle.io.DataLoader(mnist_test, batch_size=batch_size, shuffle=False, num_workers=num_workers)
return train_iter, test_iter
# -
def evaluate_accuracy(data_iter, net):
acc_sum, n = 0.0, 0
with paddle.no_grad():
for X, y in data_iter:
if isinstance(net, nn.Layer):
net.eval() # 评估模式, 这会关闭dropout
acc_sum += (net(X).argmax(axis=1) == y.flatten()).astype('float32').sum().numpy()[0]
net.train() # 改回训练模式
else: # 自定义的模型, 3.13节之后不会用到, 不考虑GPU
if('is_training' in net.__code__.co_varnames): # 如果有is_training这个参数
# 将is_training设置成False
acc_sum += (net(X, is_training=False).argmax(dim=1) == y).float().sum().item()
else:
acc_sum += (net(X).argmax(dim=1) == y).float().sum().item()
n += y.shape[0]
return acc_sum / n
def train_ch5(net, train_iter, test_iter, batch_size, optimi, num_epochs):
loss = nn.CrossEntropyLoss()
batch_count = 0
for epoch in range(num_epochs):
train_l_sum, train_acc_sum, n, start = 0.0, 0.0, 0, time.time()
for idx, (X, y) in enumerate(train_iter):
y_hat = net(X)
l = loss(y_hat, y)
optimi.clear_grad()
l.backward()
optimi.step()
train_l_sum += l.numpy()[0]
train_acc_sum += (y_hat.argmax(axis=1) == y.flatten()).astype('float32').sum().numpy()[0]
n += y.shape[0]
batch_count += 1
test_acc = evaluate_accuracy(test_iter, net)
print('epoch %d, loss %.4f, train acc %.3f, test acc %.3f, time %.1f sec'
% (epoch + 1, train_l_sum / batch_count, train_acc_sum / n, test_acc, time.time() - start))
# +
batch_size = 64
# 如出现“out of memory”的报错信息,可减小batch_size或resize
train_iter, test_iter = load_data_fashion_mnist(batch_size, resize=224)
lr, num_epochs = 0.001, 5
optimi = optimizer.Adam(parameters=net.parameters(), learning_rate=lr)
train_ch5(net, train_iter, test_iter, batch_size, optimi, num_epochs)
# -
# ## 7.2.4. 小结
#
# * VGG-11 使用可复用的卷积块构造网络。不同的 VGG 模型可通过每个块中卷积层数量和输出通道数量的差异来定义。
# * 块的使用导致网络定义的非常简洁。使用块可以有效地设计复杂的网络。
# * 在VGG论文中,Simonyan和Ziserman尝试了各种架构。特别是他们发现深层且窄的卷积(即$3 \times 3$)比较浅层且宽的卷积更有效。
#
# ## 7.2.5. 练习
#
# 1. 打印层的尺寸时,我们只看到 8 个结果,而不是 11 个结果。剩余的 3 层信息去哪了?
# 1. 与 AlexNet 相比,VGG 的计算要慢得多,而且它还需要更多的显存。分析出现这种情况的原因。
# 1. 尝试将Fashion-MNIST数据集图像的高度和宽度从 224 改为 96。这对实验有什么影响?
# 1. 请参考 VGG 论文 :cite:`Simonyan.Zisserman.2014` 中的表1构建其他常见模型,如 VGG-16 或 VGG-19。
#
# [Discussions](https://discuss.d2l.ai/t/1866)
#
|
Notebook/7/7.2.vgg.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "aae66268-03f9-445d-9716-d504b943ae61", "showTitle": false, "title": ""}
from pyspark.sql import SparkSession
spark = SparkSession.builder.appName('decisiontree').getOrCreate()
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "c6fac6b1-cc5f-4cb6-9370-9def59723142", "showTitle": false, "title": ""}
from pyspark.ml.pipeline import Pipeline
from pyspark.ml.classification import RandomForestClassifier,GBTClassifier,DecisionTreeClassifier
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "99b14779-bc38-4d92-a0a6-ab9d1a6ff4fb", "showTitle": false, "title": ""}
df = spark.read.format('libsvm').load('/FileStore/tables/sample_libsvm_data.txt')
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "c4825ac6-3aac-4c90-baad-489e6e96810a", "showTitle": false, "title": ""}
df.show()
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "5bc55525-a4df-462e-a7fb-b8258904f3e1", "showTitle": false, "title": ""}
train_df,test_df = df.randomSplit([0.7,0.3])
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "0b56fc8a-6650-4ffb-ba15-c479af57c6cd", "showTitle": false, "title": ""}
dtc = DecisionTreeClassifier()
rfc = RandomForestClassifier(numTrees=100)
gbt = GBTClassifier()
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "8f66b226-b39b-4262-9685-c03500358777", "showTitle": false, "title": ""}
dtc_model = dtc.fit(train_df)
rfc_model = rfc.fit(train_df)
gbt_model = gbt.fit(train_df)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "90857be6-20d8-445f-9856-4688d753659b", "showTitle": false, "title": ""}
dtc_preds = dtc_model.transform(test_df)
rfc_preds = rfc_model.transform(test_df)
gbt_preds = gbt_model.transform(test_df)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "b1f25beb-d7ab-4831-b60f-06d9009d8033", "showTitle": false, "title": ""}
dtc_preds.show()
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "3c29bc35-062e-4d98-82f7-bb7174ab1964", "showTitle": false, "title": ""}
from pyspark.ml.evaluation import MulticlassClassificationEvaluator
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "95c98b0b-1bbe-47a9-ab2f-52da334528ce", "showTitle": false, "title": ""}
acc_eval = MulticlassClassificationEvaluator(metricName='accuracy')
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "26542bcc-e25e-4938-b848-01e0b26a3cea", "showTitle": false, "title": ""}
print('DTC Accuracy:')
acc_eval.evaluate(dtc_preds)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "531ccce9-995a-4a06-aa7c-f855e238c05b", "showTitle": false, "title": ""}
print('RFC Accuracy:')
acc_eval.evaluate(rfc_preds)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "17b133d3-4fcc-4277-81fc-a66b2baf822e", "showTitle": false, "title": ""}
print('GBT Accuracy:')
acc_eval.evaluate(gbt_preds)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "a5c944c8-9845-46d4-a03c-89d371ab400b", "showTitle": false, "title": ""}
dtc_model.featureImportances
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "b1ccb7d3-7673-46ef-86db-3b3ec00c76ff", "showTitle": false, "title": ""}
rfc_model.featureImportances
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "397ec1ad-6746-4928-93b5-0f3d1241a9d2", "showTitle": false, "title": ""}
spark = SparkSession.builder.appName('collegedecisiontree').getOrCreate()
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "4cab1d76-232d-4fda-8768-a4268d8bff19", "showTitle": false, "title": ""}
df = spark.read.csv('/FileStore/tables/College.csv',inferSchema=True,header=True)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "97bdd98c-33e3-4794-a7d7-587804b58197", "showTitle": false, "title": ""}
df.printSchema()
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "9ced7672-565c-4780-80da-01c7bc300d5d", "showTitle": false, "title": ""}
df.head(1)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "1263ed05-6b47-4b7f-91f8-0a61f9cfe7bc", "showTitle": false, "title": ""}
from pyspark.ml.feature import VectorAssembler
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "59fc862d-01a2-483a-b6db-2bedd1df5b94", "showTitle": false, "title": ""}
df.columns
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "cbbfcead-fc80-47b3-83b7-8db7b6f2e5d5", "showTitle": false, "title": ""}
assembler = VectorAssembler(inputCols=[
'Apps',
'Accept',
'Enroll',
'Top10perc',
'Top25perc',
'F_Undergrad',
'P_Undergrad',
'Outstate',
'Room_Board',
'Books',
'Personal',
'PhD',
'Terminal',
'S_F_Ratio',
'perc_alumni',
'Expend',
'Grad_Rate'
],outputCol='features')
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "2f65e22d-b623-437c-9e2d-445f11249b21", "showTitle": false, "title": ""}
output = assembler.transform(df)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "215efee1-f6ae-481e-97ca-82b221d8b0b2", "showTitle": false, "title": ""}
from pyspark.ml.feature import StringIndexer
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "786633ef-2fcf-4bed-80ac-1407c3f0c09a", "showTitle": false, "title": ""}
indexer = StringIndexer(inputCol='Private',outputCol='Private_index')
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "b2f0934b-6ab1-44a8-8148-b34d2d5eabb2", "showTitle": false, "title": ""}
output_fixed = indexer.fit(output).transform(output)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "7425e498-907c-4f14-9f19-d3d1cad5de99", "showTitle": false, "title": ""}
output_fixed.printSchema()
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "c3815a6a-32ef-43eb-85dc-d8ee7dc7f583", "showTitle": false, "title": ""}
final_df = output_fixed.select('features','Private_index')
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "8d63872d-cc4c-48c3-ab8d-860824b13737", "showTitle": false, "title": ""}
train_df,test_df = final_df.randomSplit([0.7,0.3])
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "eadfeea0-eeed-4259-a615-8ab5b8213eac", "showTitle": false, "title": ""}
from pyspark.ml.classification import DecisionTreeClassifier,GBTClassifier,RandomForestClassifier
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "0d3807e0-7d32-4a8d-a8df-312ba9ce555e", "showTitle": false, "title": ""}
from pyspark.ml import Pipeline
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "0cf2a8b7-ef9b-4dff-b0b5-81a5690182c6", "showTitle": false, "title": ""}
dtc = DecisionTreeClassifier(labelCol='Private_index',featuresCol='features')
rfc = RandomForestClassifier(labelCol='Private_index',featuresCol='features')
gbt = GBTClassifier(labelCol='Private_index',featuresCol='features')
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "09bc08b9-8ced-447e-a956-88542dffc4c6", "showTitle": false, "title": ""}
dtc_model = dtc.fit(train_df)
rfc_model = rfc.fit(train_df)
gbt_model = gbt.fit(train_df)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "cf0564a4-2c2b-4443-89ad-c3800ad637f1", "showTitle": false, "title": ""}
dtc_preds = dtc_model.transform(test_df)
rfc_preds = rfc_model.transform(test_df)
gbt_preds = gbt_model.transform(test_df)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "ac2e2a9a-9ff7-4064-8cf8-89d2c8a5d180", "showTitle": false, "title": ""}
from pyspark.ml.evaluation import BinaryClassificationEvaluator
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "bfe0148d-3477-47bb-8b1a-e2ff23102220", "showTitle": false, "title": ""}
binary_eval = BinaryClassificationEvaluator(labelCol='Private_index')
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "6c2745c1-9872-438f-bcc0-62b0588bf351", "showTitle": false, "title": ""}
print('DTC Accuracy')
print(binary_eval.evaluate(dtc_preds))
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "7ed37be4-d970-4dc9-b143-a54dd9e6a6bb", "showTitle": false, "title": ""}
print('RFC Accuracy')
print(binary_eval.evaluate(rfc_preds))
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "a0021e1b-1f4e-4307-bf98-6b125f2c8447", "showTitle": false, "title": ""}
gbt_preds.printSchema()
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "958ffe5c-b282-499a-866c-ac843e0203a2", "showTitle": false, "title": ""}
rfc_preds.printSchema()
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "4c4fe5bf-91b0-4847-adff-9b706256a1e5", "showTitle": false, "title": ""}
binary_eval_gbt = BinaryClassificationEvaluator(labelCol='Private_index',rawPredictionCol='prediction')
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "30a7a5ce-f3b3-4115-8098-622dcf70e686", "showTitle": false, "title": ""}
print('GBT Correct Accuracy')
print(binary_eval_gbt.evaluate(gbt_preds))
print('GBT Incorrect Accuracy')
print(binary_eval.evaluate(gbt_preds))
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "e8410d74-6742-460b-b633-e3b98cdfbbb8", "showTitle": false, "title": ""}
rfc = RandomForestClassifier(numTrees=150,labelCol='Private_index',featuresCol='features')
rfc_model = rfc.fit(train_df)
rfc_preds = rfc_model.transform(test_df)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "14157877-b7c0-44de-9e09-8f9ad9ec3719", "showTitle": false, "title": ""}
print('RFC Accuracy')
print(binary_eval.evaluate(rfc_preds))
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "aacaf23b-c98d-4402-9511-14ce10fd7d4b", "showTitle": false, "title": ""}
from pyspark.ml.evaluation import MulticlassClassificationEvaluator
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "51a453c7-3fd6-4720-b377-7853ac852050", "showTitle": false, "title": ""}
acc_eval = MulticlassClassificationEvaluator(labelCol='Private_index',metricName='accuracy')
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "d184054e-124c-44a2-8f5d-15e267176f61", "showTitle": false, "title": ""}
rfc_acc = acc_eval.evaluate(rfc_preds)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "bb8a2148-6a30-47c3-8c6b-c5cb3880ca25", "showTitle": false, "title": ""}
rfc_acc
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "cfdbfab2-a2bf-42b8-9370-f1feb123269a", "showTitle": false, "title": ""}
|
projects/Spark and Python for Big Data with PySpark/Udemy_Section 13 Decision Trees and Random Forests.ipynb
|