code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import numpy as np
import pandas as pd
from os.path import join
import matplotlib.pyplot as plt
from glob import glob
from keras.models import load_model
from matplotlib.colors import LogNorm
from scipy.ndimage import gaussian_filter, maximum_filter, minimum_filter
from deepsky.gan import unnormalize_multivariate_data
from skimage.morphology import disk
import pickle
data_path = "/glade/work/dgagne/spatial_storm_results_20171220/"
#data_path = "/Users/dgagne/data/spatial_storm_results_20171220/"
scores = ["auc", "bss"]
models = ["conv_net", "logistic_mean", "logistic_pca"]
imp_scores = {}
for model in models:
imp_scores[model] = {}
for score in scores:
score_files = sorted(glob(data_path + "var_importance_{0}_{1}_*.csv".format(model, score)))
imp_score_list = []
for score_file in score_files:
print(score_file)
imp_data = pd.read_csv(score_file, index_col="Index")
imp_score_list.append(((imp_data.iloc[0,0] - imp_data.loc[1:])).mean(axis=0))
imp_scores[model][score] = pd.concat(imp_score_list, axis=1).T
imp_scores[model][score].columns = imp_scores[model][score].columns.str.rstrip("_prev"
).str.replace("_", " "
).str.replace("-component of", ""
).str.replace("dew point temperature", "dewpoint"
).str.capitalize()
fig, axes = plt.subplots(3, 2, figsize=(12, 12))
plt.subplots_adjust(wspace=0.6)
model_titles = ["Conv. Net", "Logistic Mean", "Logistic PCA"]
for m, model in enumerate(models):
for s, score in enumerate(scores):
rankings = imp_scores[model][score].mean(axis=0).sort_values().index
axes[m,s ].boxplot(imp_scores[model][score].loc[:, rankings].values, vert=False,
boxprops={"color":"k"}, whiskerprops={"color":"k"},
medianprops={"color":"k"}, flierprops={"marker":".", "markersize":3},whis=[2.5, 97.5])
axes[m, s].set_yticklabels(imp_scores[model][score].loc[:, rankings].columns.str.replace(" mb", " hPa"))
axes[m, s].set_title(model_titles[m] + " " + score.upper())
axes[m, s].grid()
#axes[m, s].set_xscale("log")
if m == len(model_titles) - 1:
axes[m, s].set_xlabel("Decrease in " + score.upper(), fontsize=12)
plt.savefig("var_imp_box.pdf", dpi=300, bbox_inches="tight")
input_cols = imp_scores[model][score].columns
log_pca_coefs = np.zeros((30, 75))
for i in range(30):
with open("/Users/dgagne/data/spatial_storm_results_20171220/" + "hail_logistic_pca_sample_{0:03d}.pkl".format(i), "rb") as pca_pkl:
log_pca_model = pickle.load(pca_pkl)
log_pca_coefs[i] = log_pca_model.model.coef_
log_gan_coefs = np.zeros((30, 64))
for i in range(30):
with open("/Users/dgagne/data/spatial_storm_results_20171220/" + "logistic_gan_{0:d}_logistic.pkl".format(i), "rb") as gan_pkl:
log_gan_model = pickle.load(gan_pkl)
log_gan_coefs[i] = log_gan_model.coef_.ravel()
plt.boxplot(np.abs(log_gan_coefs.T))
np.abs(log_pca_coefs).min()
plt.figure(figsize=(6, 10))
plt.pcolormesh(np.abs(log_pca_coefs).T, norm=LogNorm(0.0001, 1))
plt.yticks(np.arange(0, 75, 5), input_cols)
plt.barh(np.arange(15), np.abs(log_pca_coefs).mean(axis=0).reshape(15, 5).mean(axis=1))
plt.yticks(np.arange(15), input_cols)
mean_imp_matrix = pd.DataFrame(index=imp_scores["conv_net"]["bss"].columns, columns=models, dtype=float)
mean_imp_rank_matrix = pd.DataFrame(index=imp_scores["conv_net"]["bss"].columns, columns=models, dtype=int)
for model in models:
mean_imp_matrix.loc[:, model] = imp_scores[model]["bss"].values.mean(axis=0)
rank = np.argsort(imp_scores[model]["bss"].values.mean(axis=0))
for r in range(rank.size):
mean_imp_rank_matrix.loc[mean_imp_rank_matrix.index[rank[r]], model] = rank.size - r
mean_imp_matrix["conv_net"].values[np.argsort(mean_imp_matrix["conv_net"].values)]
mean_imp_rank_matrix
| notebooks/spatial_hail_feature_importance.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python
# language: python3
# name: python3
# ---
#
# <a id='matplotlib'></a>
# <div id="qe-notebook-header" align="right" style="text-align:right;">
# <a href="https://quantecon.org/" title="quantecon.org">
# <img style="width:250px;display:inline;" width="250px" src="https://assets.quantecon.org/img/qe-menubar-logo.svg" alt="QuantEcon">
# </a>
# </div>
# # Matplotlib
#
#
# <a id='index-1'></a>
# ## Contents
#
# - [Matplotlib](#Matplotlib)
# - [Overview](#Overview)
# - [The APIs](#The-APIs)
# - [More Features](#More-Features)
# - [Further Reading](#Further-Reading)
# - [Exercises](#Exercises)
# - [Solutions](#Solutions)
# ## Overview
#
# We’ve already generated quite a few figures in these lectures using [Matplotlib](http://matplotlib.org/).
#
# Matplotlib is an outstanding graphics library, designed for scientific computing, with
#
# - high-quality 2D and 3D plots
# - output in all the usual formats (PDF, PNG, etc.)
# - LaTeX integration
# - fine-grained control over all aspects of presentation
# - animation, etc.
# ### Matplotlib’s Split Personality
#
# Matplotlib is unusual in that it offers two different interfaces to plotting.
#
# One is a simple MATLAB-style API (Application Programming Interface) that was written to help MATLAB refugees find a ready home.
#
# The other is a more “Pythonic” object-oriented API.
#
# For reasons described below, we recommend that you use the second API.
#
# But first, let’s discuss the difference.
# ## The APIs
#
#
# <a id='index-2'></a>
# ### The MATLAB-style API
#
# Here’s the kind of easy example you might find in introductory treatments
# + hide-output=false
%matplotlib inline
import matplotlib.pyplot as plt
plt.rcParams["figure.figsize"] = (10, 6) #set default figure size
import numpy as np
x = np.linspace(0, 10, 200)
y = np.sin(x)
plt.plot(x, y, 'b-', linewidth=2)
plt.show()
# -
# This is simple and convenient, but also somewhat limited and un-Pythonic.
#
# For example, in the function calls, a lot of objects get created and passed around without making themselves known to the programmer.
#
# Python programmers tend to prefer a more explicit style of programming (run `import this` in a code block and look at the second line).
#
# This leads us to the alternative, object-oriented Matplotlib API.
# ### The Object-Oriented API
#
# Here’s the code corresponding to the preceding figure using the object-oriented API
# + hide-output=false
fig, ax = plt.subplots()
ax.plot(x, y, 'b-', linewidth=2)
plt.show()
# -
# Here the call `fig, ax = plt.subplots()` returns a pair, where
#
# - `fig` is a `Figure` instance—like a blank canvas.
# - `ax` is an `AxesSubplot` instance—think of a frame for plotting in.
#
#
# The `plot()` function is actually a method of `ax`.
#
# While there’s a bit more typing, the more explicit use of objects gives us better control.
#
# This will become more clear as we go along.
# ### Tweaks
#
# Here we’ve changed the line to red and added a legend
# + hide-output=false
fig, ax = plt.subplots()
ax.plot(x, y, 'r-', linewidth=2, label='sine function', alpha=0.6)
ax.legend()
plt.show()
# -
# We’ve also used `alpha` to make the line slightly transparent—which makes it look smoother.
#
# The location of the legend can be changed by replacing `ax.legend()` with `ax.legend(loc='upper center')`.
# + hide-output=false
fig, ax = plt.subplots()
ax.plot(x, y, 'r-', linewidth=2, label='sine function', alpha=0.6)
ax.legend(loc='upper center')
plt.show()
# -
# If everything is properly configured, then adding LaTeX is trivial
# + hide-output=false
fig, ax = plt.subplots()
ax.plot(x, y, 'r-', linewidth=2, label='$y=\sin(x)$', alpha=0.6)
ax.legend(loc='upper center')
plt.show()
# -
# Controlling the ticks, adding titles and so on is also straightforward
# + hide-output=false
fig, ax = plt.subplots()
ax.plot(x, y, 'r-', linewidth=2, label='$y=\sin(x)$', alpha=0.6)
ax.legend(loc='upper center')
ax.set_yticks([-1, 0, 1])
ax.set_title('Test plot')
plt.show()
# -
# ## More Features
#
# Matplotlib has a huge array of functions and features, which you can discover
# over time as you have need for them.
#
# We mention just a few.
# ### Multiple Plots on One Axis
#
#
# <a id='index-3'></a>
# It’s straightforward to generate multiple plots on the same axes.
#
# Here’s an example that randomly generates three normal densities and adds a label with their mean
# + hide-output=false
from scipy.stats import norm
from random import uniform
fig, ax = plt.subplots()
x = np.linspace(-4, 4, 150)
for i in range(3):
m, s = uniform(-1, 1), uniform(1, 2)
y = norm.pdf(x, loc=m, scale=s)
current_label = f'$\mu = {m:.2}$'
ax.plot(x, y, linewidth=2, alpha=0.6, label=current_label)
ax.legend()
plt.show()
# -
# ### Multiple Subplots
#
#
# <a id='index-4'></a>
# Sometimes we want multiple subplots in one figure.
#
# Here’s an example that generates 6 histograms
# + hide-output=false
num_rows, num_cols = 3, 2
fig, axes = plt.subplots(num_rows, num_cols, figsize=(10, 12))
for i in range(num_rows):
for j in range(num_cols):
m, s = uniform(-1, 1), uniform(1, 2)
x = norm.rvs(loc=m, scale=s, size=100)
axes[i, j].hist(x, alpha=0.6, bins=20)
t = f'$\mu = {m:.2}, \quad \sigma = {s:.2}$'
axes[i, j].set(title=t, xticks=[-4, 0, 4], yticks=[])
plt.show()
# -
# ### 3D Plots
#
#
# <a id='index-5'></a>
# Matplotlib does a nice job of 3D plots — here is one example
# + hide-output=false
from mpl_toolkits.mplot3d.axes3d import Axes3D
from matplotlib import cm
def f(x, y):
return np.cos(x**2 + y**2) / (1 + x**2 + y**2)
xgrid = np.linspace(-3, 3, 50)
ygrid = xgrid
x, y = np.meshgrid(xgrid, ygrid)
fig = plt.figure(figsize=(10, 6))
ax = fig.add_subplot(111, projection='3d')
ax.plot_surface(x,
y,
f(x, y),
rstride=2, cstride=2,
cmap=cm.jet,
alpha=0.7,
linewidth=0.25)
ax.set_zlim(-0.5, 1.0)
plt.show()
# -
# ### A Customizing Function
#
# Perhaps you will find a set of customizations that you regularly use.
#
# Suppose we usually prefer our axes to go through the origin, and to have a grid.
#
# Here’s a nice example from [<NAME>](https://github.com/xcthulhu) of how the object-oriented API can be used to build a custom `subplots` function that implements these changes.
#
# Read carefully through the code and see if you can follow what’s going on
# + hide-output=false
def subplots():
"Custom subplots with axes through the origin"
fig, ax = plt.subplots()
# Set the axes through the origin
for spine in ['left', 'bottom']:
ax.spines[spine].set_position('zero')
for spine in ['right', 'top']:
ax.spines[spine].set_color('none')
ax.grid()
return fig, ax
fig, ax = subplots() # Call the local version, not plt.subplots()
x = np.linspace(-2, 10, 200)
y = np.sin(x)
ax.plot(x, y, 'r-', linewidth=2, label='sine function', alpha=0.6)
ax.legend(loc='lower right')
plt.show()
# -
# The custom `subplots` function
#
# 1. calls the standard `plt.subplots` function internally to generate the `fig, ax` pair,
# 1. makes the desired customizations to `ax`, and
# 1. passes the `fig, ax` pair back to the calling code.
# ## Further Reading
#
# - The [Matplotlib gallery](http://matplotlib.org/gallery.html) provides many examples.
# - A nice [Matplotlib tutorial](http://scipy-lectures.org/intro/matplotlib/index.html) by <NAME>, <NAME> and <NAME>.
# - [mpltools](http://tonysyu.github.io/mpltools/index.html) allows easy
# switching between plot styles.
# - [Seaborn](https://github.com/mwaskom/seaborn) facilitates common statistics plots in Matplotlib.
# ## Exercises
# ### Exercise 1
#
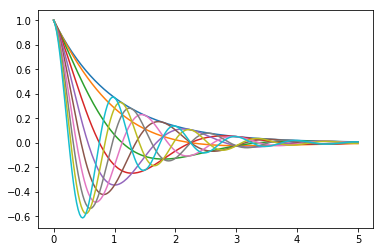
# Plot the function
#
# $$
# f(x) = \cos(\pi \theta x) \exp(-x)
# $$
#
# over the interval $ [0, 5] $ for each $ \theta $ in `np.linspace(0, 2, 10)`.
#
# Place all the curves in the same figure.
#
# The output should look like this
#
# 
# ## Solutions
# ### Exercise 1
#
# Here’s one solution
# + hide-output=false
def f(x, θ):
return np.cos(np.pi * θ * x ) * np.exp(- x)
θ_vals = np.linspace(0, 2, 10)
x = np.linspace(0, 5, 200)
fig, ax = plt.subplots()
for θ in θ_vals:
ax.plot(x, f(x, θ))
plt.show()
| tests/project/ipynb/matplotlib.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Introduction to Data Analysis
# **Data Analyst Nanodegree P2: Investigate a Dataset**
#
# **<NAME>**
#
# [List of Resources](#Resources)
#
# ## Introduction
#
# For the final project, you will conduct your own data analysis and create a file to share that documents your findings. You should start by taking a look at your dataset and brainstorming what questions you could answer using it. Then you should use Pandas and NumPy to answer the questions you are most interested in, and create a report sharing the answers. You will not be required to use statistics or machine learning to complete this project, but you should make it clear in your communications that your findings are tentative. This project is open-ended in that we are not looking for one right answer.
#
# ## Step One - Choose Your Data Set
#
# **Titanic Data** - Contains demographics and passenger information from 891 of the 2224 passengers and crew on board the Titanic. You can view a description of this dataset on the [Kaggle website](https://www.kaggle.com/c/titanic/data), where the data was obtained.
#
# From the Kaggle website:
#
# VARIABLE DESCRIPTIONS:
# survival Survival
# (0 = No; 1 = Yes)
# pclass Passenger Class
# (1 = 1st; 2 = 2nd; 3 = 3rd)
# name Name
# sex Sex
# age Age
# sibsp Number of Siblings/Spouses Aboard
# parch Number of Parents/Children Aboard
# ticket Ticket Number
# fare Passenger Fare
# cabin Cabin
# embarked Port of Embarkation
# (C = Cherbourg; Q = Queenstown; S = Southampton)
#
# SPECIAL NOTES:
# Pclass is a proxy for socio-economic status (SES)
# 1st ~ Upper; 2nd ~ Middle; 3rd ~ Lower
#
# Age is in Years; Fractional if Age less than One (1)
# If the Age is Estimated, it is in the form xx.5
#
# With respect to the family relation variables (i.e. sibsp and parch)some relations were ignored. The following are the definitions used for sibsp and parch.
#
# Sibling: Brother, Sister, Stepbrother, or Stepsister of Passenger Aboard Titanic
# Spouse: Husband or Wife of Passenger Aboard Titanic (Mistresses and Fiances Ignored)
# Parent: Mother or Father of Passenger Aboard Titanic
# Child: Son, Daughter, Stepson, or Stepdaughter of Passenger Aboard Titanic
#
# Other family relatives excluded from this study include cousins, nephews/nieces, aunts/uncles, and in-laws.
# Some children travelled only with a nanny, therefore parch=0 for them. As well, some travelled with very close friends or neighbors in a village, however, the definitions do not support such relations.
#
# ## Step Two - Get Organized
#
# Eventually you’ll want to submit your project (and share it with friends, family, and employers). Get organized before you begin. We recommend creating a single folder that will eventually contain:
#
# Using IPython notebook, containing both the code report of findings in the same document
# ## Step Three - Analyze Your Data
#
# Brainstorm some questions you could answer using the data set you chose, then start answering those questions. Here are some ideas to get you started:
#
# Titanic Data
# What factors made people more likely to survive?
# +
import pandas as pd
import numpy as np
from scipy import stats
import matplotlib.pyplot as plt
import matplotlib
# %pylab inline
matplotlib.style.use('ggplot')
# -
titanic_data = pd.read_csv('titanic_data.csv')
titanic_data.head()
# ## Step Four - Share Your Findings
#
# Once you have finished analyzing the data, create a report that shares the findings you found most interesting. You might wish to use IPython notebook to share your findings alongside the code you used to perform the analysis, but you can also use another tool if you wish.
#
# ## Step Five - Review
#
# Use the Project Rubric to review your project. If you are happy with your submission, then you're ready to submit your project. If you see room for improvement, keep working to improve your project.
# ## <a id='Resources'></a>List of Resources
#
# 1. Pandas documentation: http://pandas.pydata.org/pandas-docs/stable/index.html
# 2. Scipy ttest documentation: http://docs.scipy.org/doc/scipy-0.14.0/reference/generated/scipy.stats.ttest_rel.html
# 3. t-table: https://s3.amazonaws.com/udacity-hosted-downloads/t-table.jpg
# 4. Stroop effect Wikipedia page: https://en.wikipedia.org/wiki/Stroop_effect
| P2/p2_investigate_a_dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Controlled Abstention Networks (CAN) for regression tasks
# * authors: <NAME>, <NAME>
# * published in Barnes, <NAME>. and <NAME>: Controlled abstention neural networks for identifying skillful predictions for regression problems, submitted to JAMES, 04/2021.
# * code updated: April 22, 2021
# +
import os
import time
import sys
import pprint
import imp
import glob
from sklearn import preprocessing
import tensorflow as tf
import numpy as np
import data1d
import metrics
import abstentionloss
import network
import plots
import matplotlib.pyplot as plt
import matplotlib as mpl
mpl.rcParams['figure.facecolor'] = 'white'
mpl.rcParams['figure.dpi']= 150
np.warnings.filterwarnings('ignore', category=np.VisibleDeprecationWarning)
tf.print(f"sys.version = {sys.version}", output_stream=sys.stdout)
tf.print(f"tf.version.VERSION = {tf.version.VERSION}", output_stream=sys.stdout)
# -
# # Initialize Experiment
# +
checkpointDir = 'checkpoints/'
EXP_LIST = {
'data1d_constant':
{'exp_name': 'data1d_constant',
'loss': 'AbstentionLogLoss',
'updater': 'Constant', # set updater to Contant to run with fixed alpha
'nupd': np.nan,
'numClasses': 2,
'n_samples': [4000, 1000], #(number of noisy points, number of not noisy points)
'noise': [.5, .05],
'slope': [1., .7],
'yint': [-2., .6],
'x_sigma': [.25, .5],
'undersample': False,
'spinup': 0,
'hiddens': [5, 5],
'lr_init': .0001,
'batch_size': 32,
'np_seed': 99,
'act_fun': 'relu',
'kappa_setpoint': .1,
'fixed_alpha': .1, # fixed alpha (not using PID controller)
'n_spinup_epochs': 225,
'n_coarse_epochs': 0,
'n_epochs': 1000,
'patience': 200,
'boxcox': False,
'ridge_param': 0.,
},
}
EXPINFO = EXP_LIST['data1d_constant']
EXP_NAME = EXPINFO['exp_name']
pprint.pprint(EXPINFO, width=60)
# -
NP_SEED = EXPINFO['np_seed']
np.random.seed(NP_SEED)
tf.random.set_seed(NP_SEED)
# ## Internal functions
def get_long_name(exp_name, loss_str, setpoint, network_seed, np_seed):
# set experiment name
LONG_NAME = (
exp_name
+ '_' + loss_str
+ '_setpoint' + str(setpoint)
+ '_networkSeed' + str(network_seed)
+ '_npSeed' + str(np_seed)
)
return LONG_NAME
# +
def make_model(loss_str = 'RegressLogLoss', updater_str='Colorado', kappa=1.0e5, tau=1.0e5, spinup_epochs=0, coarse_epochs=0, setpoint=.5, nupd=10, network_seed=0):
# Define and train the model
tf.keras.backend.clear_session()
model = network.defineNN(hiddens=HIDDENS,
input_shape=X_train_std.shape[1],
output_shape=NUM_CLASSES,
ridge_penalty=RIDGE,
act_fun=ACT_FUN,
network_seed=network_seed)
if(loss_str=='AbstentionLogLoss'):
if(updater_str=='Constant'):
updater = getattr(abstentionloss, updater_str)(setpoint=setpoint,
alpha_init=FIXED_ALPHA,
)
loss_function = getattr(abstentionloss, loss_str)(kappa=kappa,
tau_fine=kappa,
updater=updater,
spinup_epochs=spinup_epochs,
coarse_epochs=coarse_epochs,
)
else:
updater = getattr(abstentionloss, updater_str)(setpoint=setpoint,
alpha_init=0.1,
length=nupd)
loss_function = getattr(abstentionloss, loss_str)(
kappa=kappa,
tau_fine=tau,
updater=updater,
spinup_epochs=spinup_epochs,
coarse_epochs=coarse_epochs,
)
model.compile(
optimizer=tf.keras.optimizers.SGD(lr=LR_INIT, momentum=0.9, nesterov=True),
loss = loss_function,
metrics=[
alpha_value,
metrics.AbstentionFraction(tau=tau),
metrics.MAE(),
metrics.MAECovered(tau=tau),
metrics.LikelihoodCovered(tau=tau),
metrics.LogLikelihoodCovered(tau=tau),
metrics.SigmaCovered(tau=tau),
]
)
else:
loss_function = getattr(abstentionloss, loss_str)()
model.compile(
optimizer=tf.keras.optimizers.SGD(lr=LR_INIT, momentum=0.9, nesterov=True),
loss = loss_function,
metrics=[
metrics.MAE(),
metrics.Likelihood(),
metrics.LogLikelihood(),
metrics.Sigma(),
]
)
# model.summary()
return model, loss_function
# +
def get_tau_vector(model,X):
y_pred = model.predict(X)
tau_dict = {}
for perc in np.around(np.arange(.1, 1.1, .1), 3):
tau_dict[perc] = np.percentile(y_pred[:, -1], 100-perc*100.)
return tau_dict
def alpha_value(y_true,y_pred):
return loss_function.updater.alpha
def scheduler(epoch, lr):
if epoch < LR_EPOCH_BOUND:
return lr
else:
return LR_INIT/2. # lr*tf.math.exp(-0.1)
class EarlyStoppingCAN(tf.keras.callbacks.Callback):
"""Stop training when the loss is at its min, i.e. the loss stops decreasing.
Arguments:
patience: Number of epochs to wait after min has been hit. After this
number of no improvement, training stops.
"""
def __init__(self, patience=0, updater_str='Colorado'):
super(EarlyStoppingCAN, self).__init__()
self.patience = patience
self.updater_str = updater_str
# best_weights to store the weights at which the minimum loss occurs.
self.best_weights = None
def on_train_begin(self, logs=None):
# The number of epoch it has waited when loss is no longer minimum.
self.wait = 0
# The epoch the training stops at.
self.stopped_epoch = 0
# Initialize the best to be the worse possible.
self.best = np.Inf
self.best_epoch = 0
# initialize best_weights to non-trained model
self.best_weights = self.model.get_weights()
def on_epoch_end(self, epoch, logs=None):
current = logs.get("val_loss")
if np.less(current, self.best):
if(self.updater_str=='Constant'):
abstention_error = 0.
else:
abstention_error = np.abs(logs.get("val_abstention_fraction") - setpoint)
if np.less(abstention_error, .1):
if (epoch >= EXPINFO['n_spinup_epochs']):
self.best = current
self.wait = 0
# Record the best weights if current results is better (greater).
self.best_weights = self.model.get_weights()
self.best_epoch = epoch
else:
self.wait += 1
if self.wait >= self.patience:
self.stopped_epoch = epoch
self.model.stop_training = True
print("Restoring model weights from the end of the best epoch.")
self.model.set_weights(self.best_weights)
def on_train_end(self, logs=None):
if self.stopped_epoch > 0:
print("Early stopping, setting to best_epoch = " + str(self.best_epoch + 1))
else:
self.best_epoch = np.nan
# -
# ## Make the data
X_train_std, onehot_train, X_val_std, onehot_val, X_test_std, onehot_test, xmean, xstd, tr_train, tr_val, tr_test = data1d.get_data(EXPINFO,to_plot=True)
# # Train the model
# +
#---------------------
LOSS = EXPINFO['loss']
UPDATER = EXPINFO['updater']
ACT_FUN = EXPINFO['act_fun']
NUPD = EXPINFO['nupd']
HIDDENS = EXPINFO['hiddens']
BATCH_SIZE = EXPINFO['batch_size']
LR_INIT = EXPINFO['lr_init']
NUM_CLASSES = EXPINFO['numClasses']
RIDGE = EXPINFO['ridge_param']
KAPPA_SEPOINT = EXPINFO['kappa_setpoint']
N_SPINUP_EPOCHS = EXPINFO['n_spinup_epochs']
N_COARSE_EPOCHS = EXPINFO['n_coarse_epochs']
PATIENCE = EXPINFO['patience']
FIXED_ALPHA = EXPINFO['fixed_alpha']
#---------------------
# Set parameters
LR_EPOCH_BOUND = 10000 # don't use the learning rate scheduler, but keep as an option
NETWORK_SEED = 0
SETPOINT_LIST = [-1., 0., .2,] # -1. = fit model for spinup period only with RegressionLogLoss
# 0. = fit model with RegressionLogLoss
# (0,1) = fit with AbstentionLogLoss for setpoint coverage,
# choose any value (0,1) if running with UPDATER = 'Constant'
for isetpoint, setpoint in enumerate(SETPOINT_LIST):
# set loss function to use ----
N_EPOCHS = EXPINFO['n_epochs']
if(setpoint==0.):
if(LOSS == 'AbstentionLogLoss'):
RUN_LOSS = 'RegressLogLoss'
else:
RUN_LOSS = LOSS
elif(setpoint==-1.):
if(LOSS == 'AbstentionLogLoss'):
RUN_LOSS = 'RegressLogLoss'
N_EPOCHS = EXPINFO['n_spinup_epochs']
else:
continue
else:
if(LOSS != 'AbstentionLogLoss' ):
continue
else:
RUN_LOSS = LOSS
#-------------------
LONG_NAME = get_long_name(EXP_NAME, RUN_LOSS, setpoint, NETWORK_SEED, NP_SEED)
model_name = 'saved_models/model_' + LONG_NAME
print(LONG_NAME)
#-------------------------------
# load the baseline spin-up model
if(setpoint>0):
spinup_file = 'saved_models/model_' + get_long_name(exp_name=EXP_NAME,
loss_str='RegressLogLoss',
setpoint=-1.,
network_seed=NETWORK_SEED,
np_seed=NP_SEED)
model_spinup, __ = make_model(loss_str = RUN_LOSS,
network_seed=NETWORK_SEED,
)
model_spinup.load_weights(spinup_file + '.h5')
tau_dict = get_tau_vector(model_spinup,X_val_std)
KAPPA = tau_dict[KAPPA_SEPOINT]
TAU = tau_dict[setpoint]
print(' kappa = ' + str(KAPPA) + ', tau = ' + str(TAU))
else:
TAU = np.nan
KAPPA = np.nan
#-------------------------------
# set random seed again
np.random.seed(NP_SEED)
# get the model
tf.keras.backend.clear_session()
# callbacks
lr_callback = tf.keras.callbacks.LearningRateScheduler(scheduler,verbose=0)
cp_callback = tf.keras.callbacks.ModelCheckpoint(
filepath = checkpointDir + 'model_' + LONG_NAME + '_epoch{epoch:03d}.h5',
verbose=0,
save_weights_only=True,
)
# define the model and loss function
if(RUN_LOSS=='AbstentionLogLoss'): # run with AbstentionLogLoss
es_can_callback = EarlyStoppingCAN(patience=PATIENCE,
updater_str=UPDATER,
)
model, loss_function = make_model(loss_str = RUN_LOSS,
updater_str=UPDATER,
kappa = KAPPA,
tau = TAU,
spinup_epochs=N_SPINUP_EPOCHS,
coarse_epochs=N_COARSE_EPOCHS,
setpoint=setpoint,
nupd=NUPD,
network_seed=NETWORK_SEED,
)
callbacks = [abstentionloss.AlphaUpdaterCallback(),
lr_callback,
cp_callback,
es_can_callback,
]
else: # run with RegressLogLoss
es_callback = tf.keras.callbacks.EarlyStopping(monitor='val_loss',
mode='min',
patience=PATIENCE,
restore_best_weights=True,
verbose=1)
model, loss_function = make_model(loss_str = RUN_LOSS,
network_seed=NETWORK_SEED,
)
callbacks = [lr_callback,
cp_callback,
es_callback,
]
#-------------------------------
# Train the model
start_time = time.time()
history = model.fit(
X_train_std,
onehot_train,
validation_data=(X_val_std, onehot_val),
batch_size=BATCH_SIZE,
epochs=N_EPOCHS,
shuffle=True,
verbose=0,
callbacks=callbacks
)
stop_time = time.time()
tf.print(f" elapsed time during fit = {stop_time - start_time:.2f} seconds\n")
model.save_weights(model_name + '.h5')
for f in glob.glob(checkpointDir + 'model_' + LONG_NAME + "_epoch*.h5"):
os.remove(f)
#-------------------------------
# Display the results
if(RUN_LOSS=='AbstentionLogLoss'):
best_epoch = es_can_callback.best_epoch
elif(setpoint==-1):
best_epoch = N_EPOCHS
else:
best_epoch = np.argmin(history.history['val_loss'])
exp_info=(RUN_LOSS, N_EPOCHS, setpoint, N_SPINUP_EPOCHS, HIDDENS, LR_INIT, LR_EPOCH_BOUND, BATCH_SIZE, NETWORK_SEED, best_epoch)
#---- plot nice predictions ----
y_pred = model.predict(X_test_std)
plots.plot_predictionscatter(X_test_std, y_pred, onehot_test[:,0], tr_test, LONG_NAME, showplot = True)
# -
| regression/main_1D.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/chuducthang77/coronavirus/blob/main/Recurring_mutation.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="I8UjHlnvKsB4"
# # Problem:
# We’ll identify mutations as re-occurred mutations if they happened earlier, then probably disappeared and again come back in some other virus sequences. By definition, recurring means something that happens over and over again at regular intervals. The timeline is 3 months. That means if a mutation 1st occurred in Jan 2020 and no mutations in Feb, Mar, and April and then appeared again in May and repeat the gap
# + colab={"base_uri": "https://localhost:8080/"} id="jAMOy7aTLWo9" outputId="aa3cb446-0081-4a19-cb57-f73c56dd13ef"
from google.colab import drive
drive.mount('/content/gdrive')
# + colab={"base_uri": "https://localhost:8080/"} id="kwPLYta6LXIZ" outputId="5f6ac725-0f5d-41db-a8af-d77c7f4cd659"
# %cd 'gdrive/MyDrive/Machine Learning/coronavirus/analysis'
# !ls
# + id="eP06VEZSLGYI"
import pandas as pd
import numpy as np
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="szLm6PP9LY2E" outputId="8401f46d-a918-4151-8256-66511c205e89"
#Read the csv
df = pd.read_csv('mutations_spike_msa_apr21.csv')
df
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="vy9NLzEdlWcI" outputId="8417ed54-7f34-4d99-b8d7-f5fa364cb262"
#Eliminate the empty row at the end of the file
df = df[df['Collection Date'].notna()]
df
# + colab={"base_uri": "https://localhost:8080/", "height": 453} id="y-CUpnwUMT1a" outputId="f565a981-b092-4843-e9e6-e4f5776b13f6"
#Create the column Month based on Collection Date
pd.options.mode.chained_assignment = None
dates = pd.to_datetime(df['Collection Date'], format='%Y-%m-%d')
dates = dates.dt.strftime('%m')
df['Month'] = dates
df['Month'] = df['Month'].astype(str).astype(int)
df
# + colab={"base_uri": "https://localhost:8080/", "height": 487} id="euN6wpsk8LFR" outputId="29b5e622-c356-4fa3-9c3d-b1da019cc57c"
#Convert the mutations columns to the list and expand for each item in the list to individual row
df = df.assign(names=df['Mutations'].str.split(',')).explode('names')
df = df.rename(columns={'names': 'Individual mutation'})
df
# + colab={"base_uri": "https://localhost:8080/"} id="HsRbfVVZ2sR6" outputId="75e956e7-f1e7-4fdf-9e71-8490e72f9da3"
#Check the mutation for the given interval
intervals = [{1,5,9},{2,6,10},{3,7,11},{4,8,12}]
result = []
for interval in intervals:
#Group the mutation by their individual mutation and create the set of month it occurs
temp_df = df.groupby('Individual mutation')['Month'].apply(set).reset_index()
#Keep only the set matching the given interval
temp_df = temp_df[temp_df['Month'] == interval]
#Keep only the unique individual mutation
result += list(temp_df['Individual mutation'].unique())
print(result)
# + id="jBpRGsc8WbMA"
#Save the output to the txt file
with open('output.txt', 'w') as output:
output.write(str(result))
| analysis/Recurring_mutation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import pandas as pd
import numpy as np
import heapq
import matplotlib.pyplot as plt
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfTransformer
from sklearn.pipeline import Pipeline
from sklearn.naive_bayes import MultinomialNB
from sklearn.model_selection import train_test_split
from sklearn import metrics
from time import time
from sklearn.datasets import fetch_20newsgroups
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.feature_extraction.text import HashingVectorizer
from sklearn.feature_selection import SelectKBest, chi2
from sklearn.linear_model import RidgeClassifier
from sklearn.pipeline import Pipeline
from sklearn.svm import LinearSVC
from sklearn.linear_model import SGDClassifier
from sklearn.linear_model import Perceptron
from sklearn.linear_model import PassiveAggressiveClassifier
from sklearn.naive_bayes import BernoulliNB, MultinomialNB
from sklearn.neighbors import KNeighborsClassifier
from sklearn.neighbors import NearestCentroid
from sklearn.ensemble import RandomForestClassifier
from sklearn.utils.extmath import density
from sklearn import metrics
def get_sample(ds, field, num=50):
ds_train = ds[ds[field]==1].sample(num)
ds_train = ds_train.append(ds[ds[field]==0].sample(num))
ds_train = ds_train.append(ds[ds[field]==-1].sample(num))
ds_train.shape
return ds_train
def build_model(X, y):
model = Pipeline([('vect', CountVectorizer())
,('tfidf', TfidfTransformer())
,('clf', MultinomialNB()),
])
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.5, random_state=42)
model = model.fit(X_train, y_train)
return model
def benchmark(clf,X_train, X_test, y_train, y_test):
#print('_' * 80)
#print("Training: ")
#print(clf)
t0 = time()
clf.fit(X_train, y_train)
train_time = time() - t0
#print("train time: %0.3fs" % train_time)
t0 = time()
pred = clf.predict(X_test)
test_time = time() - t0
#print("test time: %0.3fs" % test_time)
score = metrics.accuracy_score(y_test, pred)
#print("accuracy: %0.3f" % score)
if hasattr(clf, 'coef_'):
#print("dimensionality: %d" % clf.coef_.shape[1])
#print("density: %f" % density(clf.coef_))
if False and feature_names is not None:
print("top 10 keywords per class:")
for i, label in enumerate(target_names):
top10 = np.argsort(clf.coef_[i])[-10:]
print(trim("%s: %s" % (label, " ".join(feature_names[top10]))))
#print()
if False:
#print("classification report:")
print(metrics.classification_report(y_test, pred,
target_names=target_names))
if False:
#print("confusion matrix:")
print(metrics.confusion_matrix(y_test, pred))
#print()
clf_descr = str(clf).split('(')[0]
return clf, clf_descr, score, train_time, test_time, pred
def benchmark_models(X_train, X_test, y_train, y_test, vectorizer, path):
X_train = vectorizer.fit_transform(X_train)
X_test = vectorizer.transform(X_test)
results = []
for penalty in ["l2", "l1"]:
print('=' * 80)
print("%s penalty" % penalty.upper())
# Train Liblinear model
results.append(benchmark(LinearSVC(loss='l2', penalty=penalty,
dual=False, tol=1e-3),
X_train, X_test, y_train, y_test))
# Train SGD model
results.append(benchmark(SGDClassifier(alpha=.0001, n_iter=50,
penalty=penalty),
X_train, X_test, y_train, y_test))
# Train SGD with Elastic Net penalty
#print('=' * 80)
#print("Elastic-Net penalty")
results.append(benchmark(SGDClassifier(alpha=.0001, n_iter=50,
penalty="elasticnet"),
X_train, X_test, y_train, y_test))
# Train sparse Naive Bayes classifiers
#print('=' * 80)
#print("Naive Bayes")
results.append(benchmark(MultinomialNB(alpha=.01),
X_train, X_test, y_train, y_test))
results.append(benchmark(MultinomialNB(),
X_train, X_test, y_train, y_test))
results.append(benchmark(BernoulliNB(alpha=.01),
X_train, X_test, y_train, y_test))
#plot_scores(results,path)
return results
def plot_scores(results,path):
# make some plots
indices = np.arange(len(results))
results = [[x[i] for x in results] for i in range(6)]
clfs, clf_names, score, training_time, test_time, preds = results
training_time = np.array(training_time) / np.max(training_time)
test_time = np.array(test_time) / np.max(test_time)
plt.figure(figsize=(12, 8))
plt.title("Score")
plt.barh(indices, score, .2, label="score", color='navy')
plt.barh(indices + .3, training_time, .2, label="training time",
color='c')
plt.barh(indices + .6, test_time, .2, label="test time", color='darkorange')
plt.yticks(())
plt.legend(loc='best')
plt.subplots_adjust(left=.25)
plt.subplots_adjust(top=.95)
plt.subplots_adjust(bottom=.05)
for i, c in zip(indices, clf_names):
plt.text(-.3, i, c)
plt.savefig(path, format='eps')
def test_sample(model):
print('evaluating sample data')
docs_new = ['i agree with you', 'i disagree with you']
predicted = model.predict(docs_new)
for doc, stance in zip(docs_new, predicted):
print('%r => %s' % (doc, stance))
def calc_stats(clf_names, preds):
print('calculating f-scores')
ds_train['stance_pred'] = model.predict(ds_train.text)
types = ds_train.groupby(['type'])
for name, group in types:
#TODO: use only pos and neg inside groups
fscore=metrics.f1_score(group.stance, group.stance_pred, average='micro')
#f1_macro=metrics.f1_score(group.stance, group.stance_pred, labels=[-1,1], average='macro')
#print(name, fscore)
ds_train.loc[ds_train.type==name, 'fscore_nb'] = fscore
#ds_train.loc[ds_train.type==name, 'fscore_macro'] = f1_macro
f1_micro=metrics.f1_score(ds_train.stance, ds_train.stance_pred, average='micro')
f1_macro=metrics.f1_score(ds_train.stance, ds_train.stance_pred, average='macro')
ds_train['fscore_nb_micro'] = f1_micro
ds_train['fscore_nb_macro'] = f1_macro
fscores = ds_train.groupby('type').agg({'fscore_nb': 'mean'})
fscores = fscores.reset_index()
fscores.rename(columns={'fscore_nb': 'NB'}, inplace=True)
fscores
# fscores.loc[fscores.shape[0]] = ['F micro' , ds_train.fscore_nb_micro[0]]
# fscores.loc[fscores.shape[0]] = ['F macro' , ds_train.fscore_nb_macro[0]]
fscores['type'] = fscores['type'].str.replace('_', ' ')
#fscores.to_csv('../results/fscores.csv', index=False)
#print(fscores)
fmscores = ds_train[['fscore_nb_micro', 'fscore_nb_macro']].mean()
fmscores = fmscores.reset_index(name='NB')
f2 = ds_train[['fscore_nb_micro', 'fscore_nb_macro']].mean()
fmscores['index'] = fmscores['index'].str.replace('fscore_nb_' ,'F ')
fmscores['alg2'] = f2.values
fmscores.rename(columns={'index':'F score'}, inplace=True)
#fmscores.to_csv('../results/fmscores.csv', index=False)
#print(fmscores)
return fscores, fmscores
def benchmark_stats(X_test, y_test, results):
print('calculating f-scores')
ds_train = X_test.copy()
ds_train['y_test'] = y_test
print(y_test.shape)
results = heapq.nlargest(5, results, key=lambda x: x[2])
for r in results:
clf_name =r[1]
pred = r[5]
ds_train[clf_name] = pred
types = ds_train.groupby(['type'])
micro_stats = []
macro_stats = []
for r in results:
clf_name = r[1]
for name, group in types:
#TODO: use only pos and neg inside groups
#print(clf_name, group[clf_name].shape)
fscore=metrics.f1_score(group.y_test, group[clf_name], average='micro')
#f1_macro=metrics.f1_score(group.stance, group.stance_pred, labels=[-1,1], average='macro')
#print(name, fscore)
stat_name='fscore_'+clf_name
if not stat_name in micro_stats:
micro_stats.append(stat_name)
ds_train.loc[ds_train.type==name, stat_name] = fscore
#ds_train.loc[ds_train.type==name, 'fscore_macro'] = f1_macro
#print(len(ds_train.columns))
f1_micro=metrics.f1_score(ds_train.y_test, ds_train[clf_name], average='micro')
f1_macro=metrics.f1_score(ds_train.y_test, ds_train[clf_name], average='macro')
stat_name='fscore_micro'+clf_name
macro_stats.append(stat_name)
ds_train[stat_name] = f1_micro
stat_name='fscore_macro'+clf_name
macro_stats.append(stat_name)
ds_train[stat_name] = f1_macro
micro_stats.append('type')
fscores = ds_train[micro_stats].groupby('type').mean()
fscores = fscores.reset_index()
#fscores.rename(columns={'fscore_nb': 'NB'}, inplace=True)
fscores['type'] = fscores['type'].str.replace('_', ' ')
fscores['type'] = fscores['type'].str.replace('+', ' and ')
cols = [c.replace('fscore_', '') for c in fscores.columns]
fscores.columns = cols
#print(len(fscores.columns))
fmscores = ds_train[macro_stats].mean()
fmscores = fmscores.reset_index()
fmscores.columns = ['stat', 'value']
fmscores['stat'] = fmscores['stat'].str.replace('fscore_micro' ,'F micro ')
fmscores['stat'] = fmscores['stat'].str.replace('fscore_macro' ,'F macro ')
# fmscores['alg2'] = f2.values
# fmscores.rename(columns={'index':'F score'}, inplace=True)
#fmscores.to_csv('../results/fmscores.csv', index=False)
#print(fmscores)
return fscores, fmscores
# +
print('our english dataset...')
ds = pd.read_csv('../dataset/wiki/opinions_annotated.csv')
ds = ds[ds.lang=='en']
print('stance classification')
ds_train = get_sample(ds, 'stance')
X = ds_train[['text', 'type']]
y = ds_train.stance
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.5, random_state=42)
#vectorizer = TfidfVectorizer(sublinear_tf=True, max_df=0.5,stop_words='english')
vectorizer = TfidfVectorizer()
results = benchmark_models(X_train.text, X_test.text, y_train, y_test, vectorizer, '../results/opinions_stance_score_en.eps')
fscores, fmscores = benchmark_stats(X_test, y_test, results)
#choose the best model
#model = build_model(X,y)
model = heapq.nlargest(1, results, key=lambda x: x[2])[0][0]
print('best model: ' + str(model))
#test_sample(model)
X = vectorizer.transform(ds.text.values)
#X = ds.text
predicted = model.predict(X)
ds['stance_pred'] = predicted
fscores.to_csv('../results/opinions_fscores_stance_en.csv', index=False)
fmscores.to_csv('../results/opinions_fmscores_stance_en.csv', index=False)
print('sentiment classification')
ds_train = get_sample(ds, 'sentiment', 8)
X = ds_train[['text', 'type']]
y = ds_train.sentiment
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.5, random_state=42)
#vectorizer = TfidfVectorizer(sublinear_tf=True, max_df=0.5,stop_words='english')
vectorizer = TfidfVectorizer()
results = benchmark_models(X_train.text, X_test.text, y_train, y_test, vectorizer, '../results/opinions_sentiment_score_en.eps')
fscores, fmscores = benchmark_stats(X_test, y_test, results)
#choose the best model
#model = build_model(X,y)
model = heapq.nlargest(1, results, key=lambda x: x[2])[0][0]
print('best model: ' + str(model))
#test_sample(model)
X = vectorizer.transform(ds.text.values)
#X = ds.text
predicted = model.predict(X)
ds['sentiment_pred'] = predicted
fscores.to_csv('../results/opinions_fscores_sent_en.csv', index=False)
fmscores.to_csv('../results/opinions_fmscores_sent_en.csv', index=False)
ds.to_csv('../dataset/wiki/opinions_predicted_en.csv', index=False)
# +
print('our spanish dataset...')
ds = pd.read_csv('../dataset/wiki/opinions_annotated.csv')
ds = ds[ds.lang=='es']
print('stance classification')
ds_train = get_sample(ds, 'stance', 50)
X = ds_train[['text', 'type']]
y = ds_train.stance
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.5, random_state=42)
#vectorizer = TfidfVectorizer(sublinear_tf=True, max_df=0.5,stop_words='english')
vectorizer = TfidfVectorizer()
results = benchmark_models(X_train.text, X_test.text, y_train, y_test, vectorizer, '../results/opinions_stance_score_es.eps')
fscores, fmscores = benchmark_stats(X_test, y_test, results)
#choose the best model
#model = build_model(X,y)
model = heapq.nlargest(1, results, key=lambda x: x[2])[0][0]
print('best model: ' + str(model))
#test_sample(model)
X = vectorizer.transform(ds.text.values)
#X = ds.text
predicted = model.predict(X)
ds['stance_pred'] = predicted
fscores.to_csv('../results/opinions_fscores_stance_es.csv', index=False)
fmscores.to_csv('../results/opinions_fmscores_stance_es.csv', index=False)
print('sentiment classification')
ds_train = get_sample(ds, 'sentiment', 4)
X = ds_train[['text', 'type']]
y = ds_train.sentiment
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.5, random_state=42)
#vectorizer = TfidfVectorizer(sublinear_tf=True, max_df=0.5,stop_words='english')
vectorizer = TfidfVectorizer()
results = benchmark_models(X_train.text, X_test.text, y_train, y_test, vectorizer, '../results/opinions_sentiment_score_es.eps')
fscores, fmscores = benchmark_stats(X_test, y_test, results)
#choose the best model
#model = build_model(X,y)
model = heapq.nlargest(1, results, key=lambda x: x[2])[0][0]
print('best model: ' + str(model))
#test_sample(model)
X = vectorizer.transform(ds.text.values)
#X = ds.text
predicted = model.predict(X)
ds['sentiment_pred'] = predicted
fscores.to_csv('../results/opinions_fscores_sent_es.csv', index=False)
fmscores.to_csv('../results/opinions_fmscores_sent_es.csv', index=False)
ds.to_csv('../dataset/wiki/opinions_predicted_es.csv', index=False)
# +
print('aawd dataset...')
ds = pd.read_csv('../dataset/wiki/aawd_preprocessed.csv')
#ds = ds[ds.lang=='en']
ds_train = get_sample(ds, 'stance', 300)
X = ds_train[['text', 'type']]
print('stance classification')
y = ds_train.stance
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.5, random_state=42)
#vectorizer = TfidfVectorizer(sublinear_tf=True, max_df=0.5,stop_words='english')
vectorizer = TfidfVectorizer()
results = benchmark_models(X_train.text, X_test.text, y_train, y_test, vectorizer, '../results/awwd_stance_score.eps')
fscores, fmscores = benchmark_stats(X_test, y_test, results)
#choose the best model
model = build_model(X=ds_train.text,y = ds_train.stance)
#model = heapq.nlargest(1, results, key=lambda x: x[2])
test_sample(model)
#X = vectorizer.transform(ds.text.values)
X = ds.text
predicted = model.predict(X)
ds['stance_pred'] = predicted
ds.to_csv('../dataset/wiki/aawd_predicted.csv', index=False)
fscores.to_csv('../results/aawd_fscores_stance.csv', index=False)
fmscores.to_csv('../results/aawd_fmscores_stance.csv', index=False)
| books/3model .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PySpark
# language: python
# name: pyspark
# ---
spark.version
# + id="6hKq-n7G62-2"
from pyspark.sql import SparkSession
import matplotlib.pyplot as plt
# %matplotlib inline
import numpy as np
# + [markdown] id="V5_jvhfhtFCj"
# 1. Data loading
# -
spark = SparkSession.builder.appName("mvp-prediction").getOrCreate()
print('spark session created')
data_path = 'gs://6893-data/player_stats_2020_2021.csv'
data = spark.read.format("csv").option("header", "true").option("inferschema", "true").load(data_path)
data.show(3)
data.printSchema()
# + [markdown] id="CeRTQAUE6VfO"
# 2. Data preprocessing
# + id="3TKctNhO6bHG"
from pyspark.ml import Pipeline
from pyspark.ml.feature import OneHotEncoder, StringIndexer, VectorAssembler
# + id="_83QyptU_nDE"
#stages in our Pipeline
stages = []
# + id="BB4TOB6MBCJ3"
# Transform all features into a vector using VectorAssembler
numericCols = ["reb", "ast", "stl", "blk", "tov", "pts"]
assemblerInputs = numericCols
assembler = VectorAssembler(inputCols=assemblerInputs, outputCol="features")
stages += [assembler]
# + id="Ab0WDG00Bqc0"
pipeline = Pipeline(stages=stages)
pipelineModel = pipeline.fit(data)
preppedDataDF = pipelineModel.transform(data)
# + id="-x6nXJUiByOE"
preppedDataDF.take(3)
# + id="NPONX19OB2Tu"
# Keep relevant columns
cols = data.columns
selectedcols = ["features"] + cols
dataset = preppedDataDF.select(selectedcols)
display(dataset)
# + id="ZYB1oCw4CJuc"
### Randomly split data into training and test sets. set seed for reproducibility
#=====your code here==========
trainingData, testData = dataset.randomSplit([.70, .30], seed=100)
#===============================
print(trainingData.count())
print(testData.count())
# + [markdown] id="STxwMITSBLEH"
# 3. Modeling
# -
from pyspark.ml.classification import LogisticRegression, RandomForestClassifier
from pyspark.ml.evaluation import MulticlassClassificationEvaluator
# + id="2mej0dQPC22x"
# Fit model to prepped data
#LogisticRegression model, maxIter=10
#=====your code here==========
lrModel = LogisticRegression(featuresCol="features", labelCol="label", maxIter=10).fit(trainingData)
#===============================
# select example rows to display.
predictions = lrModel.transform(testData)
predictions.show()
# compute accuracy on the test set
evaluator = MulticlassClassificationEvaluator(labelCol="label", predictionCol="prediction", metricName="accuracy")
accuracy = evaluator.evaluate(predictions)
print("Test set accuracy = " + str(accuracy))
acc_hist = []
acc_hist.append(accuracy)
# + id="7AwIbeIwbpsY"
from pyspark.ml.classification import RandomForestClassifier
#Random Forest
#=====your code here==========
rfModel = RandomForestClassifier(featuresCol="features", labelCol="label").fit(trainingData)
#===============================
# select example rows to display.
predictions = rfModel.transform(testData)
predictions.show()
# compute accuracy on the test set
# evaluator = MulticlassClassificationEvaluator(labelCol="label", predictionCol="prediction", metricName="accuracy")
accuracy = evaluator.evaluate(predictions)
print("Test set accuracy = " + str(accuracy))
acc_hist.append(accuracy)
# + id="PHc1qAd6Skf1"
#NaiveBayes
#=====your code here==========
from pyspark.ml.classification import NaiveBayes
nbModel = NaiveBayes(featuresCol="features", labelCol="label").fit(trainingData)
#===============================
# select example rows to display.
predictions = nbModel.transform(testData)
predictions.show()
# compute accuracy on the test set
evaluator = MulticlassClassificationEvaluator(labelCol="label", predictionCol="prediction", metricName="accuracy")
accuracy = evaluator.evaluate(predictions)
print("Test set accuracy = " + str(accuracy))
acc_hist.append(accuracy)
# + id="PBbr8btnbyV3"
#Decision Tree
#=====your code here==========
from pyspark.ml.classification import DecisionTreeClassifier
dtModel = DecisionTreeClassifier(featuresCol="features", labelCol="label").fit(trainingData)
#===============================
# select example rows to display.
predictions = dtModel.transform(testData)
predictions.show()
# compute accuracy on the test set
evaluator = MulticlassClassificationEvaluator(labelCol="label", predictionCol="prediction", metricName="accuracy")
accuracy = evaluator.evaluate(predictions)
print("Test set accuracy = " + str(accuracy))
acc_hist.append(accuracy)
# + id="4nccBiy_b8KT"
#Gradient Boosting Trees
#=====your code here==========
from pyspark.ml.classification import GBTClassifier
gbtModel = GBTClassifier(featuresCol="features", labelCol="label").fit(trainingData)
#===============================
# select example rows to display.
predictions = gbtModel.transform(testData)
predictions.show()
# compute accuracy on the test set
evaluator = MulticlassClassificationEvaluator(labelCol="label", predictionCol="prediction", metricName="accuracy")
accuracy = evaluator.evaluate(predictions)
print("Test set accuracy = " + str(accuracy))
acc_hist.append(accuracy)
# + id="O9sNFLH0b_LH"
# Multi-layer Perceptron
#=====your code here==========
from pyspark.ml.classification import MultilayerPerceptronClassifier
mlpModel = MultilayerPerceptronClassifier(layers=[6, 5, 5, 2], seed=123, featuresCol="features", labelCol="label").fit(trainingData)
#===============================
# select example rows to display.
predictions = mlpModel.transform(testData)
predictions.show()
# compute accuracy on the test set
evaluator = MulticlassClassificationEvaluator(labelCol="label", predictionCol="prediction", metricName="accuracy")
accuracy = evaluator.evaluate(predictions)
print("Test set accuracy = " + str(accuracy))
acc_hist.append(accuracy)
# + id="AG_EmZcfcCIU"
# Linear Support Vector Machine
#=====your code here==========
from pyspark.ml.classification import LinearSVC
svmModel = LinearSVC(maxIter=10, regParam=0.1).fit(trainingData)
#===============================
# select example rows to display.
predictions = svmModel.transform(testData)
predictions.show()
# compute accuracy on the test set
evaluator = MulticlassClassificationEvaluator(labelCol="label", predictionCol="prediction", metricName="accuracy")
accuracy = evaluator.evaluate(predictions)
print("Test set accuracy = " + str(accuracy))
acc_hist.append(accuracy)
# + id="IvJc9VXrcGFU"
# One-vs-Rest
#=====your code here==========
from pyspark.ml.classification import LogisticRegression, OneVsRest
lr = LogisticRegression(maxIter=10, tol=1E-6, fitIntercept=True)
ovrModel = OneVsRest(classifier=lr).fit(trainingData)
#===============================
# select example rows to display.
predictions = ovrModel.transform(testData)
predictions.show()
# compute accuracy on the test set
evaluator = MulticlassClassificationEvaluator(labelCol="label", predictionCol="prediction", metricName="accuracy")
accuracy = evaluator.evaluate(predictions)
print("Test set accuracy = " + str(accuracy))
acc_hist.append(accuracy)
# -
print(len(acc_hist), acc_hist)
models = ['lr', 'rf', 'nb', 'dt', 'GBT', 'MLP', 'LSVM', 'ovr']
results = dict(zip(models, acc_hist))
results = sorted(results.items(), key=lambda x: x[1])
print(results)
accs = []
names = []
for i, t in enumerate(results):
accs.append(results[i][1])
names.append(results[i][0])
print(accs, names)
# + [markdown] id="_WIIE8pEDSR9"
# 4. Comparison and analysis
# + id="LpoclCFXD7tV"
# Rank models according to Test set accuracy
#=====your code here==========
fig = plt.figure()
ax = fig.add_axes([0,0,1,1])
ax.bar(range(len(results)), accs, align='center', tick_label=names, width=0.7)
#plt.xticks(range(len(results)), list(results.keys()))
plt.show()
#===============================
# + [markdown] id="HL3j030aa7M8"
# *your analysis*
# <br>
# The accuracy is sorted in an ascending order.
# <br>
# mlp < naive bayes < Linear SVM < Random Forest < Decision Tree < One-vs-Rest < Linear Regression < GBT
| algorithm/project_algor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
## OPERATION
# -
class Operation():
def __init__(self,input_nodes=[]):
self.input_nodes = input_nodes
self.output_nodes = []
for node in input_nodes:
node.output_nodes.append(self)
_default_graph.operations.append(self)
def compute(self):
pass
class add(Operation):
def __init__(self,x,y):
super().__init__([x,y])
def compute(self,x_var,y_var):
self.inputs = [x_var,y_var]
return x_var + y_var
class multiply(Operation):
def __init__(self,x,y):
super().__init__([x,y])
def compute(self,x_var,y_var):
self.inputs = [x_var,y_var]
return x_var * y_var
class matmul(Operation):
def __init__(self,x,y):
super().__init__([x,y])
def compute(self,x_var,y_var):
self.inputs = [x_var,y_var]
return x_var.dot(y_var)
class Placeholder():
def __init__(self):
self.output_nodes = []
_default_graph.placeholders.append(self)
class Variable():
def __init__(self,initial_value=None):
self.value = initial_value
self.output_nodes = []
self.output_node = []
_default_graph.variables.append(self)
class Graph():
def __init__(self):
self.operations = []
self.placeholders = []
self.variables = []
def set_as_default(self):
global _default_graph
_default_graph = self
# +
z = A*x + b
A = 10
b = 1
z = 10*x + 1
# -
g = Graph()
g.set_as_default()
A = Variable(10)
b = Variable(1)
x = Placeholder()
y = multiply(A,x)
z = add(y,b)
def traverse_postorder(operation):
"""
PostOrder Traversal of Nodes. Basically makes sure computations are done in
the correct order (Ax first , then Ax + b). Feel free to copy and paste this code.
It is not super important for understanding the basic fundamentals of deep learning.
"""
nodes_postorder = []
def recurse(node):
if isinstance(node, Operation):
for input_node in node.input_nodes:
recurse(input_node)
nodes_postorder.append(node)
recurse(operation)
return nodes_postorder
# +
class Session:
def run(self, operation, feed_dict = {}):
"""
operation: The operation to compute
feed_dict: Dictionary mapping placeholders to input values (the data)
"""
# Puts nodes in correct order
nodes_postorder = traverse_postorder(operation)
for node in nodes_postorder:
if type(node) == Placeholder:
node.output = feed_dict[node]
elif type(node) == Variable:
node.output = node.value
else: # Operation
node.inputs = [input_node.output for input_node in node.input_nodes]
node.output = node.compute(*node.inputs)
# Convert lists to numpy arrays
if type(node.output) == list:
node.output = np.array(node.output)
# Return the requested node value
return operation.output
# -
sess = Session()
result = sess.run(operation=z,feed_dict={x:10})
result
| FBTA/ANN/Perceptron.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from keras.datasets import mnist
import matplotlib.pyplot as plt
(X_train, y_train), (X_test, y_test) = mnist.load_data()
X_test.shape
y_test.shape
plt.imshow(X_test[90], cmap="gray")
y_test[90]
X = X_test.reshape(-1, 28*28)
y = y_test
X.shape
# +
# Step 1 - Preprocessing
# -
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
X_ = sc.fit_transform(X)
X_.shape
plt.imshow(X_[90].reshape(28,28) , cmap="gray")
# ## Sklearn PCA
from sklearn.decomposition import PCA
pca = PCA(n_components=2)
Z_pca = pca.fit_transform(X_)
Z_pca.shape
Z_pca
pca.explained_variance_
# ## Custom PCA
import numpy as np
# +
# Step 2 - Computer Covar matrix
# -
covar = np.dot(X_.T, X_)
covar.shape
# +
# Step - 3 Computer eigen vectors using SVD
# -
from numpy.linalg import svd
U, S, V = svd(covar)
U.shape
Ured = U[:, :2]
Ured.shape
# +
# Step 4 - Project of Data on New axis(Components)
# -
Z = np.dot(X_, Ured)
Z.shape
Z
# ## Visualize Dataset
import pandas as pd
new_dataset = np.hstack((Z, y.reshape(-1,1)))
dataframe = pd.DataFrame(new_dataset , columns=["PC1", "PC2", "label"])
dataframe.head()
import seaborn as sns
plt.figure(figsize=(15,15))
fg = sns.FacetGrid(dataframe, hue="label", height=10)
fg.map(plt.scatter, "PC1", "PC2")
fg.add_legend()
plt.show()
# # PCA with 784
pca = PCA()
Z_pca= pca.fit_transform(X_)
Z_pca.shape
cum_var_exaplined = np.cumsum(pca.explained_variance_ratio_)
cum_var_exaplined
plt.figure(figsize=(8,6))
plt.plot(cum_var_exaplined)
plt.grid()
plt.xlabel("n_components")
plt.ylabel("Cummulative explained variance")
plt.show()
| ml_repo/PCA/PCA.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia O3 1.6.0
# language: julia
# name: julia-o3-1.6
# ---
using Rocket
using ReactiveMP
using GraphPPL
using Distributions
using Plots
@model function kalman_filter()
x_t_min_mean = datavar(Float64)
x_t_min_var = datavar(Float64)
x_t_min ~ NormalMeanVariance(x_t_min_mean, x_t_min_var)
x_t ~ NormalMeanVariance(x_t_min, 1.0) + 1.0
γ_shape = datavar(Float64)
γ_rate = datavar(Float64)
γ ~ GammaShapeRate(γ_shape, γ_rate)
y = datavar(Float64)
y ~ NormalMeanPrecision(x_t, γ) where { q = MeanField() }
return x_t_min_mean, x_t_min_var, x_t_min, x_t, γ_shape, γ_rate, γ, y
end
function start_inference(data_stream)
model, (x_t_min_mean, x_t_min_var, x_t_min, x_t, γ_shape, γ_rate, γ, y) = kalman_filter()
x_t_min_prior = NormalMeanVariance(0.0, 1e7)
γ_prior = GammaShapeRate(0.001, 0.001)
x_t_stream = Subject(Marginal)
γ_stream = Subject(Marginal)
x_t_subscribtion = subscribe!(getmarginal(x_t, IncludeAll()), (x_t_posterior) -> begin
next!(x_t_stream, x_t_posterior)
update!(x_t_min_mean, mean(x_t_posterior))
update!(x_t_min_var, var(x_t_posterior))
end)
γ_subscription = subscribe!(getmarginal(γ, IncludeAll()), (γ_posterior) -> begin
next!(γ_stream, γ_posterior)
update!(γ_shape, shape(γ_posterior))
update!(γ_rate, rate(γ_posterior))
end)
setmarginal!(x_t, x_t_min_prior)
setmarginal!(γ, γ_prior)
data_subscription = subscribe!(data_stream, (d) -> update!(y, d))
return x_t_stream, γ_stream, () -> begin
unsubscribe!(x_t_subscribtion)
unsubscribe!(γ_subscription)
unsubscribe!(data_subscription)
end
end
# +
mutable struct DataGenerationProcess
previous :: Float64
process_noise :: Float64
observation_noise :: Float64
history :: Vector{Float64}
observations :: Vector{Float64}
end
function getnext!(process::DataGenerationProcess)
next = process.previous
process.previous = rand(Normal(process.previous, process.process_noise)) + 1.0
observation = next + rand(Normal(0.0, process.observation_noise))
push!(process.history, next)
push!(process.observations, observation)
return observation
end
function gethistory(process::DataGenerationProcess)
return process.history
end
function getobservations(process::DataGenerationProcess)
return process.observations
end
# +
n = 100
process = DataGenerationProcess(0.0, 1.0, 1.0, Float64[], Float64[])
stream = timer(100, 100) |> map_to(process) |> map(Float64, getnext!) |> take(n)
x_t_stream, γ_stream, stop_cb = start_inference(stream);
plot_callback = (posteriors) -> begin
IJulia.clear_output(true)
p = plot(mean.(posteriors), ribbon = std.(posteriors), label = "Estimation")
p = plot!(gethistory(process), label = "Real states")
p = scatter!(getobservations(process), ms = 2, label = "Observations")
p = plot(p, size = (1000, 400), legend = :bottomright)
display(p)
end
sub = subscribe!(x_t_stream |> scan(Vector{Marginal}, vcat, Marginal[]) |> map(Vector{Marginal}, reverse), lambda(
on_next = plot_callback,
on_error = (e) -> println(e)
))
# -
stop_cb()
IJulia.clear_output(false);
| demo/Infinite Data Stream.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="H3ZPVmgmMGHg" colab_type="code" colab={}
# ライブラリのインポート
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
from sklearn.tree import DecisionTreeRegressor
from sklearn.datasets import load_boston
# + id="lrY8TMglUGtE" colab_type="code" outputId="fdd9257c-2772-4ff0-950a-949d9367fe2e" executionInfo={"status": "ok", "timestamp": 1566453518531, "user_tz": -540, "elapsed": 1133, "user": {"displayName": "\u6bdb\u5229\u62d3\u4e5f", "photoUrl": "", "userId": "17854120745961292401"}} colab={"base_uri": "https://localhost:8080/", "height": 102}
# 住宅価格データセットのダウンロード
boston = load_boston()
# 特徴量に低所得者の割合(LSTAT)を選択し100行に絞り込み
X = boston.data[:100,[12]]
# 正解に住宅価格(MDEV)を設定し100行に絞り込み
y = boston.target[:100]
# 決定木回帰のモデルを作成
model = DecisionTreeRegressor(criterion='mse', max_depth=3, random_state=0)
# モデルの訓練
model.fit(X, y)
# + id="BTOo3xzfaD0C" colab_type="code" outputId="0f7389d8-7bcb-462e-d237-91fce0c1f577" executionInfo={"status": "ok", "timestamp": 1566453520064, "user_tz": -540, "elapsed": 894, "user": {"displayName": "\u6bdb\u5229\u62d3\u4e5f", "photoUrl": "", "userId": "17854120745961292401"}} colab={"base_uri": "https://localhost:8080/", "height": 295}
plt.figure(figsize=(8,4)) #プロットのサイズ指定
# 訓練データの最小値から最大値まで0.01刻みのX_pltを作成し、住宅価格を予測
X_plt = np.arange(X.min(), X.max(), 0.01)[:, np.newaxis]
y_pred = model.predict(X_plt)
# 訓練データ(低所得者の割合と住宅価格)の散布図と決定木回帰のプロット
plt.scatter(X, y, color='blue', label='data')
plt.plot(X_plt, y_pred, color='red',label='Decision tree')
plt.ylabel('Price in $1000s [MEDV]')
plt.xlabel('lower status of the population [LSTAT]')
plt.title('Boston house-prices')
plt.legend(loc='upper right')
plt.show()
# + id="MQOcQC5Y9U37" colab_type="code" colab={}
| chapter3/section3_7_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## part 1 ##
testdata = '''2199943210
3987894921
9856789892
8767896789
9899965678'''.split('\n')
testdata
with open('day9.txt') as fp:
puzzledata = fp.read().split('\n')[:-1]
puzzledata[-1]
def heightmap(data):
hmap = {}
nrows = len(data)
ncols = len(data[0])
for i, row in enumerate(data):
for j,c in enumerate(row):
height = int(c)
hmap[(i,j)] = height
return nrows, ncols, hmap
def isnbor(pos, nrows, ncols):
r, c = pos
return (0 <= r < nrows) and (0 <= c < ncols)
# +
def islow(pos, hmap, nrows, ncols):
h = hmap[pos]
row, col = pos
nbors = [(r,c) for (r,c) in [(row-1, col), (row+1, col), (row, col-1), (row, col+1)]
if isnbor((r,c), nrows, ncols)]
return all(hmap[nb] > h for nb in nbors)
# -
def lowpts(hmap, nrows, ncols):
return [pos for pos in hmap if islow(pos, hmap, nrows, ncols)]
def totrisk(pts, hmap):
return sum(hmap[pt]+1 for pt in pts)
testrows, testcols, testhmap = heightmap(testdata)
totrisk(lowpts(testhmap, testrows, testcols), testhmap)
puzzlerows, puzzlecols, puzzlehmap = heightmap(puzzledata)
totrisk(lowpts(puzzlehmap, puzzlerows, puzzlecols), puzzlehmap)
# ## part 2 ##
def walk(pos, hmap, basin):
r, c = pos
h = hmap[pos]
nbors = [(r-1, c), (r+1, c), (r, c-1), (r, c+1)]
for nb in nbors:
if (nb not in hmap) or (nb in basin):
continue
nbh = hmap[nb]
if nbh == 9:
continue
if (h < nbh):
basin.append(nb)
walk(nb, hmap, basin)
return basin
def getbasin(pt, hmap):
return walk(pt, hmap, [pt])
import math
def solve(hmap, nrows, ncols):
basin_sizes = []
for pt in lowpts(hmap, nrows, ncols):
basin = getbasin(pt, hmap)
basin_sizes.append(len(basin))
return math.prod(sorted(basin_sizes)[-3:])
solve(testhmap, testrows, testcols)
solve(puzzlehmap, puzzlerows, puzzlecols)
| day9.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## Libs
# +
import os
import shap
import pickle
import warnings
import numpy as np
import pandas as pd
import xgboost as xgb
import lightgbm as lgbm
from tqdm.notebook import tqdm
import matplotlib.pyplot as plt
from utils.general import viz_performance
from sklearn.tree import DecisionTreeClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import classification_report, confusion_matrix
from sklearn.ensemble import RandomForestClassifier, AdaBoostClassifier, ExtraTreesClassifier
from sklearn.model_selection import cross_validate, cross_val_score, KFold, train_test_split
warnings.filterwarnings('ignore')
# -
# ## Constants
SEED = 1
# ## Data Ingestion
DATA_RAW_PATH = os.path.join('..','data','raw')
MODEL_PATH = os.path.join('..', 'models')
DATA_RAW_NAME = ['abono.csv', 'aposentados.csv']
DATA_IMG_PATH = os.path.join('..', 'figures')
DATA_INTER_PATH = os.path.join('..','data','interim')
DATA_PROCESSED_PATH = os.path.join('..','data','processed')
DATA_INTER_NAME = 'interim.csv'
DATA_PROCESSED_NAME = 'processed.csv'
MODEL_NAME = 'model.pkl'
df = pd.read_csv(os.path.join(DATA_PROCESSED_PATH, DATA_PROCESSED_NAME), sep='\t')
df.head(3)
df.duplicated().sum()
# ## Modeling
df.RENDIMENTO_TOTAL.value_counts(normalize=True)
train = df.drop('RENDIMENTO_TOTAL', axis=1)
X_train, X_test, y_train, y_test = train_test_split(train, df.RENDIMENTO_TOTAL,
test_size=.3,
random_state=SEED,
stratify=df.RENDIMENTO_TOTAL)
# ### Baseline
reglog = LogisticRegression(
class_weight='balanced',
solver='saga',
random_state=SEED
)
viz_performance(X_train, X_test, y_train, y_test, reglog, ['0', '1'], figsize=(14,14))
plt.savefig(os.path.join(DATA_IMG_PATH,'baseline-reglog-metrics.png'), format='png')
# ### Models
models = [
('DecisionTree', DecisionTreeClassifier(random_state=SEED)),
('RandomForest', RandomForestClassifier(random_state=SEED)),
('ExtraTree', ExtraTreesClassifier(random_state=SEED)),
('Adaboost', AdaBoostClassifier(random_state=SEED)),
('XGBoost', xgb.XGBClassifier(random_state=SEED, verbosity=0)),
('LightGBM', lgbm.LGBMClassifier(random_state=SEED))
]
# +
original = pd.DataFrame()
for name, model in tqdm(models):
kfold = KFold(n_splits=5, random_state=SEED, shuffle=True)
score = cross_validate(model, X_train, y_train, cv=kfold, scoring=['precision_weighted','recall_weighted','f1_weighted'], return_train_score=True)
additional = pd.DataFrame({
'precision_train':np.mean(score['train_precision_weighted']),
'precision_test':np.mean(score['test_precision_weighted']),
'recall_train':np.mean(score['train_recall_weighted']),
'recall_test':np.mean(score['test_recall_weighted']),
'f1_train':np.mean(score['train_f1_weighted']),
'f1_test':np.mean(score['test_f1_weighted']),
}, index=[name])
new = pd.concat([original, additional], axis=0)
original = new
original
# +
results = []
names = []
for name, model in tqdm(models):
cv_results = cross_val_score(model, X_train, y_train, cv=kfold, scoring=('f1_weighted'))
results.append(cv_results)
names.append(name)
fig = plt.figure(figsize=(12,6))
fig.suptitle('Comparação entre algoritmos - F1-Score')
ax = fig.add_subplot(111)
plt.boxplot(results)
ax.set_xticklabels(names)
plt.ylabel('f1 score')
plt.xticks(rotation=45)
plt.grid(b=False)
plt.show()
plt.savefig(os.path.join(DATA_IMG_PATH,'models-score.png'), format='png')
# -
# ### Selected Model
# +
clf = lgbm.LGBMClassifier(random_state=SEED, max_depth=5)
clf.fit(X_train,y_train)
y_pred = clf.predict(X_test)
print(confusion_matrix(y_test, y_pred))
print(classification_report(y_test, y_pred))
# -
viz_performance(X_train, X_test, y_train, y_test, clf, ['0', '1'], figsize=(14,14))
plt.savefig(os.path.join(DATA_IMG_PATH,'final-lgbm-metrics.png'), format='png')
# +
explainer = shap.TreeExplainer(clf)
shap_values = explainer.shap_values(X_test)
shap.summary_plot(shap_values[1], X_test)
plt.savefig(os.path.join(DATA_IMG_PATH,'variable-performance.png'), format='png')
# -
# ### Export Model
pickle.dump(clf, open(os.path.join(MODEL_PATH,MODEL_NAME), 'wb'))
# ## Conclusions
#
# 1. Base foi dividida em 30% para teste e 70% para treino do modelo;
# 2. Variável alvo estava balanceada com proporções de ~51.5% e 48.4%;
# 3. Baseline de uma Regressao Logista que mostrou baixa separação entre os *target*, tendo um modelo que pouco generalizava;
# 4. Selecionado alguns modelos para avaliar performaces;
# 5. Utilzado técnicas de *cross validate* com *k-fold* = 5;
# 6. Como métrica a ser avaliada foi escolhida o *f1-score*, porém, baseado no problema que queremos atacar poderiamos escolher uma outra métrica;
# 7. O modelo baseado em árvore foi selecionado (LigthGBM) por ter um f1-score mais alto no conjunto de teste;
# 8. Com um modelo baseado em árvore conseguimos bater o nosso *baseline* tendo uma melhor separação das variáveis de acordo com *KS Statistic*, um AUC de 85%. Poderiamos variar o *threshold* para otimizar *recall* ou *precision*;
# 9. As variáveis que possuem um maior poder de predição são: NV_ESCOLARIDADE, PADRAO e CARGO;
# 10. A variável CLASSE apesar de ter um poder de predição positivo, também dificultou na predição, chegando a ter um SHAP value de -2;
#
# Obs.: Não houve tunagem de hiperparâmetros.
| notebooks/05 - MODELING.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.11 64-bit (''lightweight'': conda)'
# language: python
# name: python3
# ---
# # Optuna Fashion MNIST CNN Model Usage
# +
import os
import optuna
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
import torch.utils.data
from torchvision import datasets
from torchvision import transforms
# -
# ### 1. Set HyperParameter
# +
BATCHSIZE = 128
CLASSES = 10
EPOCHS = 10
DIR = os.getcwd()
DEVICE = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
print(DEVICE)
N_TRAIN_EXAMPLES = BATCHSIZE * 469 # Fashion MNIST Full Dataset size
N_VALID_EXAMPLES = BATCHSIZE * 79
# -
# ### 2. Get FashionMNIST to dataloader
def get_mnist():
# Load FashionMNIST dataset.
train_loader = torch.utils.data.DataLoader(
datasets.FashionMNIST(DIR, train=True, download=True, transform=transforms.ToTensor()),
batch_size=BATCHSIZE,
shuffle=True,
)
valid_loader = torch.utils.data.DataLoader(
datasets.FashionMNIST(DIR, train=False, transform=transforms.ToTensor()),
batch_size=BATCHSIZE,
shuffle=True,
)
return train_loader, valid_loader
# ### 3. Trial Model
class ConvNets(nn.Module):
def __init__(self, trial):
super().__init__()
self.out_layer = None
self.layer = self.define_model(trial)
self.in_features = None
self.in_channel = None
def define_model(self, trial):
n_layers = trial.suggest_int("n_layers", 1, 2)
layers = []
self.in_features = 28
self.in_channel = 1
for i in range(n_layers):
out_channel = trial.suggest_int(f"conv_c{i}", 32, 512),
kernel_size = trial.suggest_int(f"conv_k{i}", 3, 5)
stride = trial.suggest_int(f"conv_s{i}", 1, 2)
padding = trial.suggest_int(f"conv_p{i}", 0, 2)
layers.append(nn.Conv2d(self.in_channel, out_channel[0], kernel_size, stride, padding))
layers.append(nn.BatchNorm2d(out_channel[0]))
layers.append(nn.ReLU())
layers.append(nn.MaxPool2d(kernel_size=2,stride=2))
out_features = (self.in_features - kernel_size + 2*padding) // (stride) + 1
out_features = out_features // 2
print(f"conv_layer_{i}:", self.in_channel, out_channel, self.in_features, out_features, kernel_size, stride, padding)
self.in_features = out_features
self.in_channel = out_channel[0]
print('pow(self.in_features,2)* self.in_channel:',pow(self.in_features,2)* self.in_channel)
p = trial.suggest_float("dropout_l{}".format(i), 0.0, 0.5)
layers.append(nn.Flatten())
layers.append(nn.Linear(pow(self.in_features,2)* self.in_channel, 1000))
layers.append(nn.Dropout(p))
layers.append(nn.Linear(1000, 10))
layers.append(nn.LogSoftmax(dim=1))
return nn.Sequential(*layers)
def forward(self, x):
x = self.layer(x)
def forward(self, x):
x = self.layer(x)
return x
# ### 4. objective funtion for optuna
def objective(trial):
model = ConvNets(trial).to(DEVICE)
optimizer_name = trial.suggest_categorical("optimizer", ["Adam"])# "RMSprop"])
lr = trial.suggest_float("lr", 1e-5, 1e-3, log=True)
optimizer = getattr(optim, optimizer_name)(model.parameters(), lr=lr)
train_loader, valid_loader = get_mnist()
#epochs = trial.suggest_int("epochs", 1, EPOCHS)
#with tqdm(range(EPOCHS), total=EPOCHS, unit='epoch') as bar:
for epoch in range(EPOCHS):
#bar.set_description(f"Epoch {epoch}")
model.train()
for batch_idx, (data, target) in enumerate(train_loader):
if batch_idx * BATCHSIZE >= N_TRAIN_EXAMPLES:
break
data, target = data.to(DEVICE), target.to(DEVICE)
optimizer.zero_grad()
output = model(data)
loss = F.nll_loss(output, target)
loss.backward()
optimizer.step()
model.eval()
correct = 0
with torch.no_grad():
for batch_idx, (data, target) in enumerate(valid_loader):
if batch_idx * BATCHSIZE >= N_VALID_EXAMPLES:
break
data, target = data.to(DEVICE), target.to(DEVICE)
output = model(data)
pred = output.argmax(dim=1, keepdim=True)
correct += pred.eq(target.view_as(pred)).sum().item()
accuracy = correct / min(len(valid_loader.dataset), N_VALID_EXAMPLES)
trial.report(accuracy, epoch)
if trial.should_prune():
raise optuna.exceptions.TrialPruned()
return accuracy
# ### 5. Run optuna study
if __name__ == "__main__":
study = optuna.create_study(directions=["maximize"])
study.optimize(objective, n_trials=100)
| utils/03_optuna_mnist_cnn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Exercise 04 : Building the GAN network
# Import the required library functions
import tensorflow as tf
import numpy as np
from numpy.random import randn
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense
from matplotlib import pyplot
# Function to generate real samples
def realData(loc,batch):
# loc is the random location or mean around which samples are centred
# Generate numbers to right of the random point
xr = np.arange(loc,loc+(0.1*batch/2),0.1)
xr = xr[0:int(batch/2)]
# Generate numbers to left of the random point
xl = np.arange(loc-(0.1*batch/2),loc,0.1)
# Concatenating both these series
X1 = np.concatenate((xl,xr))
# Second dependent variable
X2 = np.sin(X1)
# Reshaping both the variables and then concatenating them to an array of independent variables
X1 = X1.reshape(batch,1)
X2 = X2.reshape(batch,1)
X = np.concatenate((X1,X2),axis = 1)
# Generating the labels for real data set which is 'ones'
y = np.ones((batch,1))
return X,y
# +
# Function to generate inputs for generator function
def fakeInputs(batch,infeats):
# Sample data points equal to (batch x input feature size) from a random distribution
genInput = randn(infeats * batch)
# Reshape the input
X = genInput.reshape(batch ,infeats)
return X
# -
# Function for the generator model
def genModel(infeats,outfeats):
# Defining the Generator model
Genmodel = Sequential()
Genmodel.add(Dense(32,activation = 'linear',kernel_initializer='he_uniform',input_dim=infeats))
Genmodel.add(Dense(32,activation = 'relu',kernel_initializer='he_uniform'))
Genmodel.add(Dense(64,activation = 'elu',kernel_initializer='he_uniform'))
Genmodel.add(Dense(32,activation = 'elu',kernel_initializer='he_uniform'))
Genmodel.add(Dense(32,activation = 'selu',kernel_initializer='he_uniform'))
Genmodel.add(Dense(outfeats,activation = 'selu'))
return Genmodel
# Function to create fake samples using the generator model
def fakedataGenerator(Genmodel,batch,infeats):
# first generate the inputs to the model
genInputs = fakeInputs(batch,infeats)
# use these inputs inside the generator model to generate fake distribution
X_fake = Genmodel.predict(genInputs)
# Generate the labels of fake data set
y_fake = np.zeros((batch,1))
return X_fake,y_fake
# Define the arguments like batch size,input feature size and output feature size
batch = 128
infeats = 10
outfeats = 2
# Next we develop the discriminator model which is a network having 4 layers
# Descriminator model as a function
def discModel(outfeats):
Discmodel = Sequential()
Discmodel.add(Dense(16, activation='relu',kernel_initializer = 'he_uniform',input_dim=outfeats))
Discmodel.add(Dense(16,activation='relu' ,kernel_initializer = 'he_uniform'))
Discmodel.add(Dense(16,activation='relu' ,kernel_initializer = 'he_uniform'))
Discmodel.add(Dense(1,activation='sigmoid'))
# Compiling the model
Discmodel.compile(loss='binary_crossentropy',optimizer='adam',metrics=['accuracy'])
return Discmodel
# Print the summary of the discriminator model
Discmodel = discModel(outfeats)
Discmodel.summary()
# Calling the Generator model function
Genmodel = genModel(infeats,outfeats)
Genmodel.summary()
# Let us visualize the initial fake data
x_fake, _ = fakedataGenerator(Genmodel,batch,infeats)
# Plotting the fake data using pyplot
pyplot.scatter(x_fake[:, 0], x_fake[:, 1], color='blue')
# Adding x and y labels
pyplot.xlabel('Feature 1 of the distribution')
pyplot.ylabel('Feature 2 of the distribution')
pyplot.show()
# Define the combined generator and discriminator model, for updating the generator
def ganModel(Genmodel,Discmodel):
# First define that discriminator model cannot be trained
Discmodel.trainable = False
Ganmodel = Sequential()
# First adding the generator model
Ganmodel.add(Genmodel)
# Next adding the discriminator model without training the parameters
Ganmodel.add(Discmodel)
# Compile the model for loss to optimise the Generator model
Ganmodel.compile(loss='binary_crossentropy',optimizer = 'adam')
return Ganmodel
# Initialise the gan model
gan_model = ganModel(Genmodel,Discmodel)
# Print summary of the GAN model
gan_model.summary()
# Defining the number of epochs
nEpochs = 20000
# Train the GAN network
for i in range(nEpochs):
# Generate the random number for generating real samples
loc = np.random.normal(3,1,1)
# Generate samples equal to the bath size from the real distribution
x_real, y_real = realData(loc,batch)
# Generate fake samples using the fake data generator function
x_fake, y_fake = fakedataGenerator(Genmodel,batch,infeats)
# train the discriminator on the real samples
Discmodel.train_on_batch(x_real, y_real)
# train the discriminator on the fake samples
Discmodel.train_on_batch(x_fake, y_fake)
# Generate new fake inputs for training the GAN network
x_gan = fakeInputs(batch,infeats)
# Create labels of the fake examples as 1 to fool the discriminator
y_gan = np.ones((batch, 1))
# Update the generator model through the discriminator model
gan_model.train_on_batch(x_gan, y_gan)
# Print the accuracy measures on the real and fake data for every 2000 epochs
if (i) % 2000 == 0:
# Generate samples equal to the bath size from the real distribution
x_real, y_real = realData(loc,batch)
# Evaluate Real distribution accuracy
_, realAccuracy = Discmodel.evaluate(x_real, y_real, verbose=0)
# Generate fake samples using the fake data generator function
x_fake,y_fake = fakedataGenerator(Genmodel,batch,infeats)
# Evaluate fake distribution accuracy levels
_, fakeAccuracy = Discmodel.evaluate(x_fake, y_fake, verbose=0)
print('Real accuracy:{R},Fake accuracy:{F}'.format(R=realAccuracy,F=fakeAccuracy))
# scatter plot real and fake data points
pyplot.scatter(x_real[:, 0], x_real[:, 1], color='red')
pyplot.scatter(x_fake[:, 0], x_fake[:, 1], color='blue')
pyplot.xlabel('Feature 1 of the distribution')
pyplot.ylabel('Feature 2 of the distribution')
# save plot to file
filename = 'GAN_Training_Plot%03d.png' % (i)
pyplot.savefig(filename)
pyplot.close()
| Exercise04/Exercise04.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Deep Learning with Python
#
# ## 3.5 Classifying newswires: a multiclass classification example
#
# > 新闻分类: 多分类问题
#
# [原文链接](https://livebook.manning.com/book/deep-learning-with-python/chapter-3/192)
#
# 3.4 节里我们不是把向量输入分成两类嘛,这节我们要把东西分成多类,即做“多分类(multi-class classification)”。
#
# 我们要把来自路透社的新闻分到 46 个话题种类里,
# 这里要求一条新闻只能属于一个类,所以具体来说,我们要做的是一个“单标签多分类(single-label, multiclass classification)”问题。
#
# ### 路透社数据集
#
# the Reuters dataset,路透社在 1986 年(比我老多了😂)发布的数据集,里面有 46 类新闻,训练集里每类至少 10 条数据。
#
# 这个玩具数据集和 IMDB、MNIST 一样,也在 Keras 里内置了:
# +
from tensorflow.keras.datasets import reuters
(train_data, train_labels), (test_data, test_labels) = reuters.load_data(
num_words=10000)
# -
# 这个数据集里面的数据和之前的 IMDB 一样,把单词翻译成了数字,然后我们只截取出现频率最高的10000个词。
#
# 咱们这个训练集里有 8K+ 条数据,测试集 2K+:
print(len(train_data), len(test_data))
# 咱们还是像搞 IMDB 时那样,把数据还原会文本看看:
# +
def decode_news(data):
reverse_word_index = {v: k for k, v in reuters.get_word_index().items()}
return ' '.join([reverse_word_index.get(i - 3, '?') for i in data])
# i - 3 是因为 0、1、2 为保留词 “padding”(填充)、“start of sequence”(序列开始)、“unknown”(未知词)
text = decode_news(train_data[0])
print(text)
# -
# 标签是 0~45 的数字:
train_labels[0]
# ### 数据准备
#
# 首先,还是把数据位向量化,直接套用我们搞 IMDB 时写的代码:
# +
import numpy as np
def vectorize_sequences(sequences, dimension=10000):
results = np.zeros((len(sequences), dimension))
for i, sequence in enumerate(sequences):
results[i, sequence] = 1.
return results
x_train = vectorize_sequences(train_data)
x_test = vectorize_sequences(test_data)
# -
# 然后就是这种效果:
x_train
# 然后要处理标签。我们可以把标签处理成整数张量,也可以用 `One-hot` 编码
# 对于分类这种问题,我们常用 one-hot 编码(也叫*分类编码*,categorical encoding)。
#
# 对于我们当前的问题,使用 one-hot 编码,即用除了标签索引位置为 1 其余位置全为 0 的向量:
# +
def to_one_hot(labels, dimension=46):
results = np.zeros((len(labels), dimension))
for i, label in enumerate(labels):
results[i, label] = 1.
return results
one_hot_train_labels = to_one_hot(train_labels)
one_hot_test_labels = to_one_hot(test_labels)
# -
# 其实,,,Keras 里自带了一个可以干这个事情的函数:
# +
from tensorflow.keras.utils import to_categorical
# 书上是 from keras.utils.np_utils import to_categorical 但,,,时代变了,而且咱这用的是 tensorflow.keras,所以稍微有点区别
one_hot_train_labels = to_categorical(train_labels)
one_hot_test_labels = to_categorical(test_labels)
# -
one_hot_train_labels
# ### 构建网络
#
# 这个问题和之前的电影评论分类问题还是差不多的,只是最后的解的可能从 2 -> 46,解空间大了太多了。
#
# 对于我们用的 Dense 层堆叠,每层都是接收上一层输出的信息作为输入。
# 所以,如果某一层丢失了一些信息,那么这些信息就再也不能被后面的层找回来了。
# 如果丢失的信息对分类没用,那这种丢失是好的、我们期望发生的;
# 但如果这些丢失的信息是对最后分类起作用的,那这种丢失就制约网络的结果了。
# 也就是说,这可能造成一种“信息瓶颈”。这种瓶颈在每一层都可能发生。
#
# 之前的电影评论分类最后只要 2 个结果,所以我们把层里的单元是用了 16 个,
# 即让机器在一个 16 维空间里学习,以及足够大了,不太会有“信息瓶颈“。
#
# 而我们现在的问题,解空间是 46 维的。
# 直接照搬之前的代码,让它在 16 维空间里学习,肯定有瓶颈!
#
# 解决瓶颈的办法很简单,直接增加层里的单元就好。这里我们是 16 -> 64:
# +
from tensorflow.keras import models
from tensorflow.keras import layers
model = models.Sequential()
model.add(layers.Dense(64, activation='relu', input_shape=(10000,)))
model.add(layers.Dense(64, activation='relu'))
model.add(layers.Dense(46, activation='softmax'))
# -
# 在最后一层,我们的输出是 46 维的,对应 46 种分类,
# 而这一层的激活函数是 softmax,和我们在训练 MNIST 时用的一样。
#
# 用 softmax 可以让网络输出在 46 种分类上的概率分布,即一个 46 维的向量,
# 其中第 i 个元素代表输入属于第 i 种分类的可能性,
# 并且这 46 个元素的总和为 `1`。
#
# ### 编译模型
#
# 编译模型,又要确定损失函数、优化器和优化的目标了。
#
# - 损失函数,分类问题嘛,还是用“分类交叉熵” categorical_crossentropy。
# - 优化器,其实对很多问题我们都是用 rmsprop
# - 目标还是一个,预测的精度 accuracy
model.compile(optimizer='rmsprop',
loss='categorical_crossentropy',
metrics=['accuracy'])
# ### 验证效果
#
# 我们还是要搞一个验证集来在训练过程中评估模型的。从训练集里分个 1K 条数据出来就好:
# +
x_val = x_train[:1000]
partial_x_train = x_train[1000:]
y_val = one_hot_train_labels[:1000]
partial_y_train = one_hot_train_labels[1000:]
# -
# ### 训练模型
#
# 好了,准备工作完成,又可以看到最迷人的训练过程了!
history = model.fit(partial_x_train,
partial_y_train,
epochs=20,
batch_size=512,
validation_data=(x_val, y_val))
# 🆗挺快的,照例,还是画图看看训练过程。
#
# 1. 训练过程中的损失
# +
import matplotlib.pyplot as plt
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(1, len(loss) + 1)
plt.plot(epochs, loss, 'bo-', label='Training loss')
plt.plot(epochs, val_loss, 'rs-', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
# -
# 2. 训练过程中的精度
# +
plt.clf()
acc = history.history['accuracy']
val_acc = history.history['val_accuracy']
plt.plot(epochs, acc, 'bo-', label='Training acc')
plt.plot(epochs, val_acc, 'rs-', label='Validation acc')
plt.title('Training and validation accuracy')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
# -
# Emmmm,说,第9轮 epochs 的时候开始过拟合了(你看validation的曲线抖在第9轮了一下)。
# 所以只要跑 9 轮就够了。
# +
model = models.Sequential()
model.add(layers.Dense(64, activation='relu', input_shape=(10000,)))
model.add(layers.Dense(64, activation='relu'))
model.add(layers.Dense(46, activation='softmax'))
model.compile(optimizer='rmsprop',
loss='categorical_crossentropy',
metrics=['accuracy'])
model.fit(partial_x_train,
partial_y_train,
epochs=9,
batch_size=512,
validation_data=(x_val, y_val))
# -
# 然后,用测试集测试一下:
results = model.evaluate(x_test, one_hot_test_labels, verbose=2)
print(results)
# 精度差不多 80%,其实还是不错的了,比随机瞎划线去分好多了。
#
# 如果随机划线去分类的话,对二元分类问题精度是 50 %,而对这 46 元的分类精度只要不到 19% 了:
# +
import copy
test_labels_copy = copy.copy(test_labels)
np.random.shuffle(test_labels_copy)
hits_array = np.array(test_labels) == np.array(test_labels_copy)
float(np.sum(hits_array)) / len(test_labels)
# -
# 调用 model 实例的 predict 方法,可以得到对输入在 46 个分类上的概率分布:
predictions = model.predict(x_test)
predictions
# 分别代表 46 个分类的可能
predictions[0].shape
# 总和为 1
np.sum(predictions[0])
# 最大的,即我们认为这条新闻属于这个分类
np.argmax(predictions[0])
# ### 处理标签和损失的另一种方法
#
# 前面提到了标签可以使用 one-hot 编码,或者直接把标签处理成整数张量:
y_train = np.array(train_labels)
y_test = np.array(test_labels)
# 用这种的话,损失函数也要跟着改,改成 sparse_categorical_crossentropy,
# 这个和 categorical_crossentropy 在数学上是一样的,只是接口不同:
model.compile(optimizer='rmsprop',
loss='sparse_categorical_crossentropy',
metrics=['acc'])
# ### 中间层维度足够大的重要性
#
# 之前讨论了关于“信息瓶颈”的事,然后我们就说对这个 46 维结果的网络,中间层的维度要足够大!
#
# 现在咱试试如果不够大(导致信息瓶颈)会怎么样,咱搞夸张一点,从 64 减到 4:
# +
model = models.Sequential()
model.add(layers.Dense(64, activation='relu', input_shape=(10000,)))
model.add(layers.Dense(4, activation='relu'))
model.add(layers.Dense(46, activation='softmax'))
model.compile(optimizer='rmsprop',
loss='categorical_crossentropy',
metrics=['accuracy'])
model.fit(partial_x_train,
partial_y_train,
epochs=20,
batch_size=128,
validation_data=(x_val, y_val))
# -
# 看看这,这训练出来比之前 64 维的差的不是一点半点哈,差距相当明显了。
#
# 发生这种效果的下降就是因为你给他学习的空间维度太低了,他把好多对分类有用的信息抛弃了。
#
# 那是不是越大越好?我们再试试把中间层加大一些:
# +
model = models.Sequential()
model.add(layers.Dense(64, activation='relu', input_shape=(10000,)))
model.add(layers.Dense(4096, activation='relu'))
model.add(layers.Dense(46, activation='softmax'))
model.compile(optimizer='rmsprop',
loss='categorical_crossentropy',
metrics=['accuracy'])
model.fit(partial_x_train,
partial_y_train,
epochs=20,
batch_size=128,
validation_data=(x_val, y_val))
# -
# 可以看到训练用的时间长了一点,电脑更暖手了一点,但效果却没有多大的提升。
# 这是由于第一层输入到中间层的只有 64 维嘛,中间层再大,也被第一层的瓶颈制约了。
#
# 在试试把第一层也加大!
# +
model = models.Sequential()
model.add(layers.Dense(512, activation='relu', input_shape=(10000,)))
model.add(layers.Dense(512, activation='relu'))
model.add(layers.Dense(46, activation='softmax'))
model.compile(optimizer='rmsprop',
loss='categorical_crossentropy',
metrics=['accuracy'])
model.fit(partial_x_train,
partial_y_train,
epochs=20,
batch_size=128,
validation_data=(x_val, y_val))
# -
# (稍微小一点,本来是用 4096 的,但太大了,咱乞丐版 mbp 跑的贼慢,跑完要20多分钟,我懒得等)
#
# 这个多浪费了好多时间,而且他很快就~~过泥河~~过拟合了,过得还过得很严重,画个图看一下:
# +
import matplotlib.pyplot as plt
loss = _.history['loss']
val_loss = _.history['val_loss']
epochs = range(1, len(loss) + 1)
plt.plot(epochs, loss, 'bo-', label='Training loss')
plt.plot(epochs, val_loss, 'rs-', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
# -
# 所以,太大了也不好。还是要有个度!
# ### 尝试使用更少/更多的层
#
# 1. 更少的层
# +
model = models.Sequential()
model.add(layers.Dense(46, activation='softmax', input_shape=(10000,)))
model.compile(optimizer='rmsprop',
loss='categorical_crossentropy',
metrics=['accuracy'])
history = model.fit(partial_x_train,
partial_y_train,
epochs=20,
batch_size=128,
validation_data=(x_val, y_val))
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(1, len(loss) + 1)
plt.plot(epochs, loss, 'bo-', label='Training loss')
plt.plot(epochs, val_loss, 'rs-', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
# -
# 快呀!结果稍微差了一点点。
# 2. 更多的层
# +
model = models.Sequential()
model.add(layers.Dense(64, activation='relu', input_shape=(10000,)))
model.add(layers.Dense(64, activation='relu'))
model.add(layers.Dense(64, activation='relu'))
model.add(layers.Dense(64, activation='relu'))
model.add(layers.Dense(46, activation='softmax'))
model.compile(optimizer='rmsprop',
loss='categorical_crossentropy',
metrics=['accuracy'])
history = model.fit(partial_x_train,
partial_y_train,
epochs=20,
batch_size=128,
validation_data=(x_val, y_val))
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(1, len(loss) + 1)
plt.plot(epochs, loss, 'bo-', label='Training loss')
plt.plot(epochs, val_loss, 'rs-', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
# -
# 所以,这个也不是越多越好呀!
| ch3/3.5 Classifying newswires.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.5
# language: python
# name: python3
# ---
# ## Segmenting and Clustering Neighborhoods in Toronto
import pandas as pd
# First task is to parse data from Wikipedia:
from IPython.display import IFrame
url = "https://en.wikipedia.org/wiki/List_of_postal_codes_of_Canada:_M"
IFrame(url, width=800, height=350)
# Data can be parsed using `BeautifulSoap`, but it's more straightforward just to use `Pandas` and its function `read_html`:
data, = pd.read_html(url, match="Postcode", skiprows=1)
data.columns = ["PostalCode", "Borough", "Neighborhood"]
data.head()
# Only process the cells that have an assigned borough. Ignore cells with a borough that is "Not assigned".
data = data[data["Borough"] != "Not assigned"]
data.head()
# More than one neighborhood can exist in one postal code area. For example, in the table on the Wikipedia page, you will notice that M5A is listed twice and has two neighborhoods: Harbourfront and Regent Park. These two rows will be combined into one row with the neighborhoods separated with a comma.
#
# Solution is to group data by `PostalCode` and aggregate columns. For borough, it's sufficient to pick first item from the resulting series and for neighbourhood, items are joined together using `", ".join(s)`:
borough_func = lambda s: s.iloc[0]
neighborhood_func = lambda s: ", ".join(s)
agg_funcs = {"Borough": borough_func, "Neighborhood": neighborhood_func}
data_temp = data.groupby(by="PostalCode").aggregate(agg_funcs)
data_temp.head()
# Some postprocessing is needed; reset the index and add columns back to right order:
data = data_temp.reset_index()[data.columns]
data.head()
# If a cell has a borough but a "Not assigned" neighborhood, then the neighborhood will be the same as the borough. So for the 9th cell in the table on the Wikipedia page, the value of the Borough and the Neighborhood columns will be Queen's Park.
data[data["Neighborhood"] == "Not assigned"]
# We can, for example, iterate through table and replace the values:
for (j, row) in data.iterrows():
if row["Neighborhood"] == "Not assigned":
borough = row["Borough"]
print("Replace \"Not assigned\" => %s in row %i" % (borough, j))
row["Neighborhood"] = borough
# To check data, examine row 85, which should be the only changed one:
data.iloc[83:88]
# The size of the data is now:
data.shape
# ## Determining coordinates for each neigbourhood
import sys
# !conda install -c conda-forge geopy --yes --prefix {sys.prefix}
# !conda install -c conda-forge folium=0.5.0 --yes --prefix {sys.prefix}
# !conda install -c conda-forge geocoder --yes --prefix {sys.prefix}
# Using geocoder with google service results `OVER_QUERY_LIMIT`: Keyless access to Google Maps Platform is deprecated. Please use an API key with all your API calls to avoid service interruption. For further details please refer to http://g.co/dev/maps-no-account. It seems to be quite hard to fetch location data from internet without api keys, so instead use the csv file approach this time:
locations = pd.read_csv("https://cocl.us/Geospatial_data")
locations.head()
locations.columns = ["PostalCode", "Latitude", "Longitude"]
data2 = pd.merge(data, locations, on='PostalCode')
data2.head()
# To check that merging was succesfull, find the first postal code `M5G` which should be `(43.657952, -79.387383)`:
data2[data2["PostalCode"] == "M5G"]
data2[data2["PostalCode"] == "M5A"]
# The ordering of the dataframe in assignment is unknown but clearly we have correct latitude and longitude now attached for each postal code.
# ## Explore data
# Let's filter only rows where Borough contains word Toronto and explore and cluster that.
subset = data2[data2['Borough'].str.contains("Toronto")]
subset.head()
subset.shape
# (copypaste all stuff from lab example notebook and so on ...)
| Segmenting and Clustering Neighborhoods in Toronto - final notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # Using simple NumPy operations for manipulating images
#
# This script illustrates how to use basic NumPy operations, such as slicing,
# masking and fancy indexing, in order to modify the pixel values of an image.
#
# +
import numpy as np
from skimage import data
import matplotlib.pyplot as plt
camera = data.camera()
camera[:10] = 0
mask = camera < 87
camera[mask] = 255
inds_x = np.arange(len(camera))
inds_y = (4 * inds_x) % len(camera)
camera[inds_x, inds_y] = 0
l_x, l_y = camera.shape[0], camera.shape[1]
X, Y = np.ogrid[:l_x, :l_y]
outer_disk_mask = (X - l_x / 2)**2 + (Y - l_y / 2)**2 > (l_x / 2)**2
camera[outer_disk_mask] = 0
plt.figure(figsize=(4, 4))
plt.imshow(camera, cmap='gray')
plt.axis('off')
plt.show()
| digital-image-processing/notebooks/numpy_operations/plot_camera_numpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Matrix factorization of an explicit feedback (ratings) matrix
#
# The goal of this exercise is to generate recommendations for movies using Matrix Factorization (MF) and Neural Network MF.
#
# See also:
# * https://arxiv.org/abs/1708.05031
# * http://hameddaily.blogspot.fr/2016/12/simple-matrix-factorization-with.html
# * https://nipunbatra.github.io/blog/2017/recommend-keras.html
#
# !pip install -q tensorflow==2.0.0-beta0
# !pip install -q matplotlib
# !pip install -q pandas
# !pip install -q numpy
# +
import tensorflow as tf
import numpy
import pandas as pd
import matplotlib
# %matplotlib inline
print("TensorFlow version: {}".format(tf.__version__))
print("Eager execution: {}".format(tf.executing_eagerly()))
# -
# ### Load the data
# The small MovieLens dataset: https://grouplens.org/datasets/movielens/100k/
# +
from io import BytesIO
from zipfile import ZipFile
from urllib.request import urlopen
resp = urlopen("http://files.grouplens.org/datasets/movielens/ml-100k.zip")
zipfile = ZipFile(BytesIO(resp.read()))
file = 'ml-100k/u.data'
df = pd.read_csv(zipfile.open(file), low_memory=False, skiprows=[0], sep='\t', names=['user', 'item', 'rate', 'time'])
df.head()
# -
# We split the dataset into training and test subset. We remove the mean rating.
#
#
# +
numpy.random.seed(42)
# split data into train and test set
msk = numpy.random.rand(len(df)) < 0.7
df_train = df[msk]
df_test = df[~msk]
user_index = [x-1 for x in df_train.user.values]
item_index = [x-1 for x in df_train.item.values]
user_index_test = [x-1 for x in df_test.user.values]
item_index_test = [x-1 for x in df_test.item.values]
rates = df_train.rate.values
rates_test = df_test.rate.values
num_ratings = len(rates)
num_ratings_test = len(rates_test)
mean_rating = numpy.mean(rates)
mean_rating_test = numpy.mean(rates_test)
rates = rates - mean_rating
rates_test = rates_test - mean_rating
print ("Mean (train) rating = " + str(mean_rating))
print ("Number of ratings (train/val/total) = " + str(num_ratings) + "/" + str(num_ratings_test) + "/" + str(num_ratings + num_ratings_test))
# -
# # Matrix Factorization
#
# In matrix factorization user rating r is formulated as an inner product of two latent vectors $u$ and $v$ which are two latent vectors in same space to represent the user interest and movie feature respectively.
#
# $r=u^Tv$
#
from IPython.display import Image
from IPython.core.display import HTML
Image(url= "https://4.bp.blogspot.com/-95QD5t9Lha4/Wd7uWnBZBeI/AAAAAAAADg4/xB4VnnxM0UgUp15lNmB3aHCXYGejpm4OACLcBGAs/s1600/matrix_factorization.png")
# ### Latent space
#
# We define these two latent variables in TensorFlow as follows: $U$ shows latent representation of user interest and $P$ that represents the latent values for items.
#
# The dimension of the latent space 'feature_len' is a parameter of the method.
#
# +
# variables
feature_len = 10
num_users = len(numpy.unique(df.user.values))
num_items = len(numpy.unique(df.item.values))
print("Number of users is {}".format(num_users))
print("Number of movies is {}".format(num_items))
print("The latent space has dimension {}".format(feature_len))
# -
# ### Model
# We define the user and item matrices and use their product to compute ratings R.
#
# +
# product embedding
item_input = tf.keras.layers.Input(shape=[1],name='Item')
item_embedding = tf.keras.layers.Embedding(num_items, feature_len, name='Item-Embedding')(item_input)
item_vec = tf.keras.layers.Flatten(name='FlattenItems')(item_embedding)
# user embedding
user_input = tf.keras.layers.Input(shape=[1],name='User')
user_embedding = tf.keras.layers.Embedding(num_users, feature_len, name='User-Embedding')(user_input)
user_vec = tf.keras.layers.Flatten(name='FlattenUsers')(user_embedding)
# rating
#user_vec_transp = tf.transpose(user_vec)
result = tf.keras.layers.dot([item_vec, user_vec], axes=1, name='DotProduct')
# initialize Keras model
model = tf.keras.Model([user_input, item_input], result)
model.summary()
# -
# ### Loss and optimizer
#
# To learn model parameters, we optimize the model with respect to mean squared error loss. As the optimization algorithm we use stochastic gradient descent (SGD) or Adam optimizer.
#
# +
loss = tf.keras.losses.MeanSquaredError()
# learning rate
lr = 0.001
learning_rate = tf.optimizers.schedules.ExponentialDecay(lr, decay_steps=100000,
decay_rate=0.96, staircase=True)
optimizer = tf.optimizers.Adam(learning_rate=learning_rate)
#optimizer = tf.optimizers.SGD(learning_rate=lr)
# -
# ### Metrics
# We define the accuracy metric.
# +
# Define accuracy
threshold = 1.0
def accuracy(desired_rates, predicted_rates):
diff_op = tf.subtract(predicted_rates, desired_rates, name='trainig_diff')
# Just measure the absolute difference against the threshold
good = tf.less(tf.abs(diff_op), threshold)
return tf.reduce_mean(tf.cast(good, tf.float32))
# -
# ### Compile the Keras model
# compile the model with the optimizerm, loss and the tracking metrics
model.compile(optimizer=optimizer, loss=loss, metrics=[accuracy])
# ### Training
#
# During training, we evaluate the accuracy on a validation set (validation_split=0.1).
#
# Question: Choose the number of epochs such that the model does not overfit the training set.
#
numberEpochs = 10
history = model.fit([user_index, item_index], rates, epochs=numberEpochs, verbose=1, validation_split=0.1)
# ### Visualize the training history
# We can visualize the training and validation loss.
# +
import matplotlib.pyplot as plt
def plot_history(history):
pd.Series(history.history['loss']).plot(logy=True, label='Training loss')
pd.Series(history.history['val_loss']).plot(logy=True, label='Validation loss')
pd.Series(history.history['accuracy']).plot(logy=True, label='Training accuracy')
pd.Series(history.history['val_accuracy']).plot(logy=True, label='Validation accuracy')
plt.legend()
plt.xlabel("Epoch")
plt.show()
plot_history(history)
# -
# ### Evaluation on the test dataset
# Let's look at the predictions for some users from the testset. We also compute MAE on the testset.
#
#
#
# +
# example
nr_sampled_users = 10
for index in numpy.random.choice(range(len(user_index_test)), size=nr_sampled_users):
u = [user_index_test[index]]
p = [item_index_test[index]]
r = rates_test[index] + mean_rating
rhat = model.predict([u, p]) + mean_rating_test
print ("rating for user " + str(u) + " for item " + str(p) + " is " + str(r) + " and our prediction is: " + str(rhat[0]))
# +
from sklearn.metrics import mean_absolute_error
def compute_error(model, user_index_test, item_index_test):
predicted_rates_test = model.predict([user_index_test, item_index_test])
return mean_absolute_error(rates_test, predicted_rates_test)
err_test = compute_error(model, user_index_test, item_index_test)
print("Mean absolute error on the test set: {}".format(err_test))
# -
# ### Generate recommendations from the trained model for a list of users
#
# Noe that we have trained a model to predict ratings, we generate recommendations for every user by ranking the movies by the predicted rating.
#
def create_recommendations(model, df, listOfUsers, nrRecommendations=20):
item_index = numpy.array(numpy.unique(df.item.values))-1
recommendations_set = {}
for user in listOfUsers:
user_index = numpy.ones(len(item_index)) * user
predicted_rates = model.predict([user_index, item_index]) #+ mean_rating
ranked_items_idx = numpy.argsort(predicted_rates, axis=0)[::-1].squeeze()
ranked_items = item_index[ranked_items_idx]
recommendations_set[user] = ranked_items[:nrRecommendations]
return recommendations_set
# ### Create validation set for every user
# We first create a validation set for every user which consists of all the products that the user rated higher than 3.5 (the value of the mean rate).
#
def create_validation_set(df, minRate=3.5, k=5):
validation_set = {}
for user in numpy.unique(df['user'].values) - 1:
rated_items = df[df['user'] == user]['item'].values -1
rates = df[df['user'] == user]['rate'].values
best_ranked_items = rated_items[numpy.where(rates > minRate)[0]]
if len(best_ranked_items) >= k:
validation_set[user] = best_ranked_items
return validation_set
# ### Compute precision@k
#
# In the context of recommendation systems we are most likely interested in recommending top-N items to the user. So it makes more sense to compute precision and recall metrics in the first N items instead of all the items. Thus the notion of precision at k where k is a user definable integer that is set by the user to match the top-N recommendations objective.
#
# Precision at k is the proportion of recommended items in the top-k set that are relevant.
#
#
def precisionAtK(validations_set, recommendations_set, k=5):
res = []
for user in validations_set.keys():
v = validations_set[user]
r = recommendations_set[user][:k]
ans = len(numpy.intersect1d(v, r)) / k
res.append(ans)
return numpy.mean(res)
# +
def compute_precisionAtK_from_recommendations(model, df_test, validation_set=None, nrRecommendations=20, k=10):
"""
Compute precisionAtK from recommendations and validation set. Generate recommendations applying \
'model' to dataset 'df_test'.
"""
if validation_set is None:
validation_set = create_validation_set(df_test, minRate=3.5, k=k)
recommendations_set = create_recommendations(model, df_test, \
validation_set.keys(), nrRecommendations=nrRecommendations)
precision = precisionAtK(validation_set, recommendations_set, k=k)
return precision
# +
k = 5
validation_set = create_validation_set(df_test, minRate=3.5, k=k)
precision = compute_precisionAtK_from_recommendations(model, df_test, validation_set=validation_set, \
nrRecommendations=20, k=k)
print("MF: Precision@{} is {}".format(k, precision))
# -
# ## Exercise
# Experiment with the parameter choice of the MF model and evaluate the setting in MAE and Precision@5. Compare various settings in the ResultsTable.
# +
class ResultsTable():
def __init__(self):
self.columns = ["name", "MAE", "Precision@5"]
self.df = pd.DataFrame(columns=self.columns)
def add(self, name="experimentName", mae=None, precision=None, overwrite=False):
data ={"name":name, "MAE":mae, "Precision@5": precision}
res = pd.Series(data, self.columns, name=name)
if len(self.df[self.df.name == name]) > 0:
if not overwrite:
raise Exception("Error: experiments name already exists. Change the name or set overwrite to True.")
else:
self.df = self.df.drop(self.df[self.df.name == name].index[0])
self.df = self.df.append(res, ignore_index=True)
else:
self.df = self.df.append(res, ignore_index=True)
def show(self):
display(self.df)
table = ResultsTable()
# -
# Compute the MAE and Precision@5.
# +
err_test = compute_error(model, user_index_test, item_index_test)
precision = compute_precisionAtK_from_recommendations(model, df_test, validation_set=validation_set, nrRecommendations=20, k=5)
print("MF: Precision@{} is {}".format(5, precision))
print("MF: MAE is {}".format(err_test))
table.add(name="MF", mae=err_test, precision=precision, overwrite=True)
table.show()
# -
## example:
experiment_name = "MF_" + "regUserItemL2"
# ### Question:
# - choose the regularization
# https://keras.io/regularizers/
# - loss
# - optimizer
# - learning rate
# - number of epochs
#
# Benchmark several experiments with different hyperparameters
#
#
# +
# regularizers
no_regularizer = None
regularizer_l2 = tf.keras.regularizers.l2(0.0001)
regularizer_l1 = tf.keras.regularizers.l1(0.0001)
regularizer_l1l2 = tf.keras.regularizers.l1_l2(0.0001)
regularizer_user = regularizer_l2
regularizer_product = regularizer_l2
# +
# product embedding
item_input = tf.keras.layers.Input(shape=[1],name='Item')
################ EMBEDDING AND REGULARIZER ##########################################################################
item_embedding = tf.keras.layers.Embedding(num_items, feature_len, name='Item-Embedding', \
embeddings_regularizer=regularizer_product)(item_input)
#################################################################################################
item_vec = tf.keras.layers.Flatten(name='FlattenItems')(item_embedding)
# user embedding
user_input = tf.keras.layers.Input(shape=[1],name='User')
################ EMBEDDING AND REGULARIZER ##########################################################################
user_embedding = tf.keras.layers.Embedding(num_users, feature_len,name='User-Embedding', \
embeddings_regularizer=regularizer_user)(user_input)
#################################################################################################
user_vec = tf.keras.layers.Flatten(name='FlattenUsers')(user_embedding)
# rating
result = tf.keras.layers.dot([item_vec, user_vec], axes=1, name='DotProduct')
# initialize Keras model
model = tf.keras.Model([user_input, item_input], result)
#model.summary()
# +
# choose the loss
# see https://keras.io/losses/
loss = tf.keras.losses.MeanSquaredError()
# learning rate
lr = 0.001
learning_rate = tf.optimizers.schedules.ExponentialDecay(lr, decay_steps=100000,
decay_rate=0.96, staircase=True)
# choose the optimizer
optimizer = tf.optimizers.Adam(learning_rate=learning_rate)
#optimizer = tf.optimizers.SGD(learning_rate=lr)
# compile the model with the optimizerm, loss and the tracking metrics
model.compile(optimizer=optimizer, loss=loss, metrics=[accuracy])
#model.summary()
numberEpochs = 10
history = model.fit([user_index, item_index], rates, epochs=numberEpochs, verbose=1, validation_split=0.1)
plot_history(history)
# -
# MAE
# +
err_test = compute_error(model, user_index_test, item_index_test)
precision = compute_precisionAtK_from_recommendations(model, df_test, validation_set=validation_set, \
nrRecommendations=20, k=5)
print("MF: Precision@{} is {}".format(5, precision))
print("MF: MAE is {}".format(err_test))
# -
table.add(name=experiment_name, mae=err_test, precision=precision, overwrite=True)
table.show()
#table.add(name="experiment_l1regularization", mae=err_test, precision=precision)
table.show()
# # Neural networks for recommendation
# Create a simple neural network for recommendation, or for estimating rating! This model is very similar to the earlier matrix factorisation models, but differs in the following ways:
#
# - Instead of taking a dot product of the user and the item embedding, we concatenate them and use them as features for our neural network. Thus, we are not constrained to the dot product way of combining the embeddings, and can learn complex non-linear relationships.
# - We can now have a different dimension of user and item embeddings. This can be useful if one dimension is larger than the other.
experiment_name = "NNMF"
# +
n_latent_factors_user = 50
n_latent_factors_item = 100
item_input = tf.keras.layers.Input(shape=[1],name='Item')
item_embedding = tf.keras.layers.Embedding(num_items, n_latent_factors_item, name='Item-Embedding')(item_input)
item_vec = tf.keras.layers.Flatten(name='FlattenItems')(item_embedding)
item_vec = tf.keras.layers.Dropout(0.2)(item_vec)
user_input = tf.keras.layers.Input(shape=[1],name='User')
user_embedding = tf.keras.layers.Embedding(num_users, n_latent_factors_user,name='User-Embedding')(user_input)
user_vec = tf.keras.layers.Flatten(name='FlattenUsers')(user_embedding)
user_vec = tf.keras.layers.Dropout(0.2)(user_vec)
concat = tf.keras.layers.concatenate([item_vec, user_vec], name='Concat')
concat = tf.keras.layers.Dropout(0.2)(concat)
full1 = tf.keras.layers.Dense(200,name='FullyConnected', activation='relu')(concat)
full1 = tf.keras.layers.Dropout(0.2,name='Dropout')(full1)
# full2 = tf.keras.layers.Dense(100,name='FullyConnected-1', activation='relu')(full1)
# full2 = tf.keras.layers.Dropout(0.2,name='Dropout')(full2)
# full3 = tf.keras.layers.Dense(50,name='FullyConnected-2', activation='relu')(full2)
# full3 = tf.keras.layers.Dropout(0.2,name='Dropout')(full3)
# full4 = tf.keras.layers.Dense(20,name='FullyConnected-3', activation='relu')(full3)
result = tf.keras.layers.Dense(1, name='Activation')(full1)
lr = 0.0001
learning_rate = tf.optimizers.schedules.ExponentialDecay(lr, decay_steps=100000,
decay_rate=0.96, staircase=True)
adam = tf.optimizers.Adam(learning_rate=learning_rate)
model_NN = tf.keras.Model([user_input, item_input], result)
model_NN.compile(optimizer=adam, loss= 'mean_squared_error', metrics=[accuracy])
model_NN.summary()
# +
numberEpochs = 10
print_log = 1
history_NN = model_NN.fit([user_index, item_index], rates, epochs=numberEpochs, \
verbose=print_log, validation_split=0.1)
# -
plot_history(history_NN)
# +
experiment_name = "NNMF"
err_test = compute_error(model_NN, user_index_test, item_index_test)
precision = compute_precisionAtK_from_recommendations(model_NN, df_test, validation_set=validation_set, \
nrRecommendations=20, k=5)
print("MF: Precision@{} is {}".format(5, precision))
print("MF: MAE is {}".format(err_test))
table.add(name=experiment_name, mae=err_test, precision=precision, overwrite=True)
table.show()
# -
# ### Question:
# Change the model parameters, for example:
# - Change the number of layers of the NN
# - Remove/add dropout
#
#
#
# ### Design a simple baseline: predict average rate per item and recommend items with the highest rating
# +
gbo = df_test[["item", "rate"]].groupby("item").mean().reset_index()
df_gbo = pd.merge(df_test, gbo, on="item", suffixes=('', '_gbo'))
display(df_gbo[df_gbo.item == 1].head())
predicted_rates_gbo_test = df_gbo.rate_gbo.values
err_gbo_test = mean_absolute_error(rates_test + mean_rating_test, predicted_rates_gbo_test)
print("Mean absolute error on the test set: {}".format(err_gbo_test))
# -
# ### Generate recommendations
def recommend_highest_rates(df, userList, nrRecommendations=10):
gbo_set = {}
# compute mean rating per item
all_items = numpy.unique(df['item'].values)
gbos = df.groupby("item").mean().rate.reset_index().sort_values(by="rate", ascending=False).item.values[:nrRecommendations]
for user in userList:
gbo_set[user] = gbos
return gbo_set
# +
highest_rates_baseline = recommend_highest_rates(df, validation_set.keys(), nrRecommendations=20)
precision_baseline = precisionAtK(validation_set, highest_rates_baseline, k=5)
print("Baseline: Precision@{} is {}".format(5, precision_baseline))
table.add(name="gbo", mae=err_gbo_test, precision=precision_baseline, overwrite=True)
# -
# ### Final scores
display(table.df.sort_values(by="MAE"))
display(table.df.sort_values(by="Precision@5", ascending=False))
| application/part_1/ExerciseMF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="9JSOqsx3RtcH" colab_type="text"
# # Code-lab on Reinforcement Learning
# ## A Code-lab session by <NAME> ([@Krupagaliya](https://twitter.com/Krupagaliya)) at #DevFestAhm18
# + [markdown] id="J3xYM0hC4ivO" colab_type="text"
# 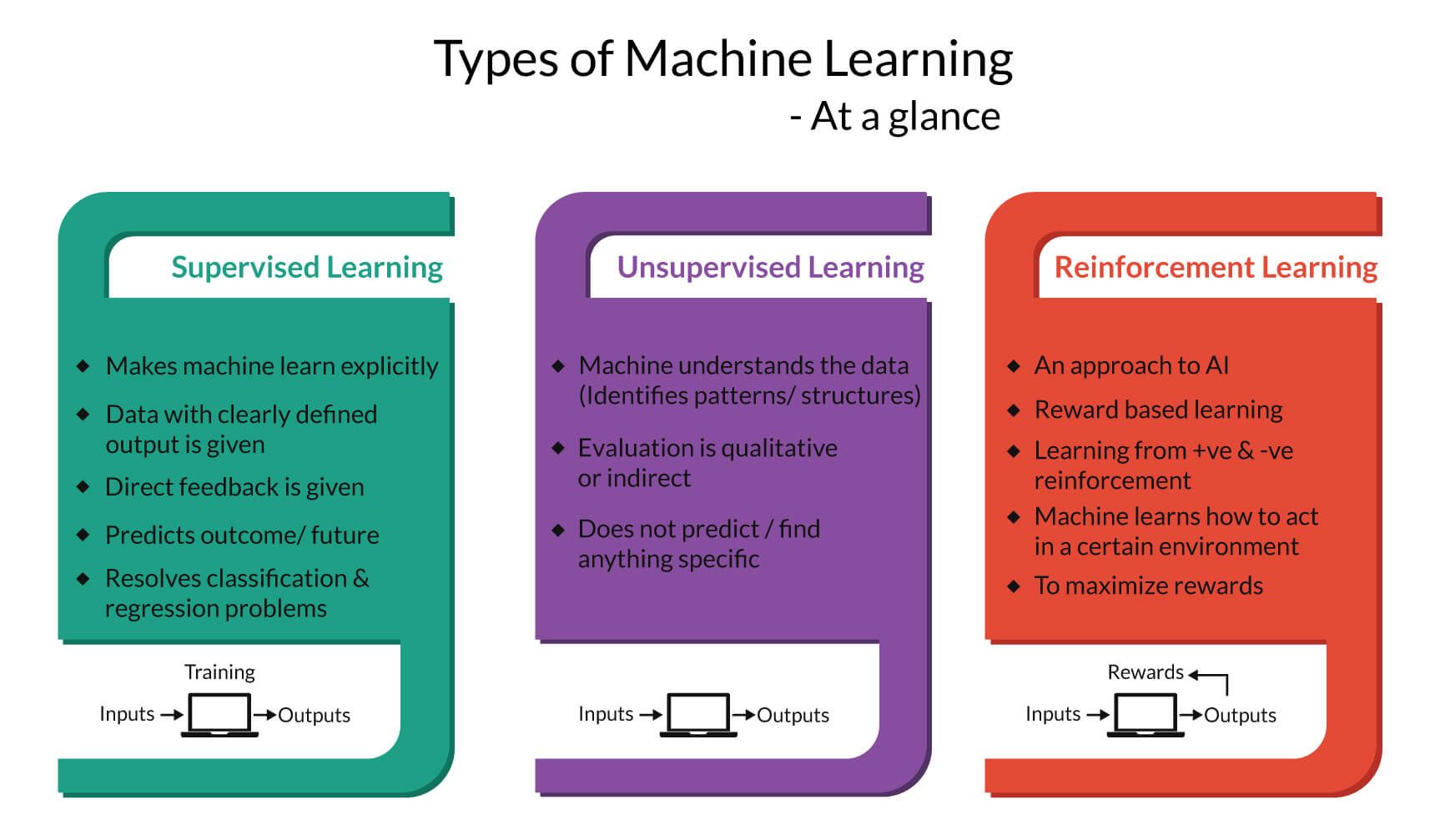
# + [markdown] id="vM6kEluL6sax" colab_type="text"
# ## Reinforcement Learning
# + [markdown] id="fysVo3bV66N_" colab_type="text"
#
#
# > A way of programming agents by rewards and punishments without needing to specify how the task to be achieved.
#
#
# > An agent that is decision-maker interacts with the environment and learns through trial-and-error
# 
#
#
#
# ---
#
#
#
# > **Cool, But how it learn ??**
#
#
#
#
# 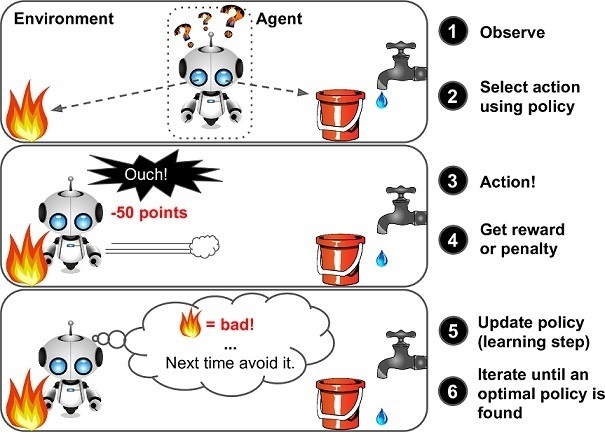
#
#
#
#
#
#
#
#
# + [markdown] id="hFJRZMAG-bh_" colab_type="text"
# # Applications
#
# 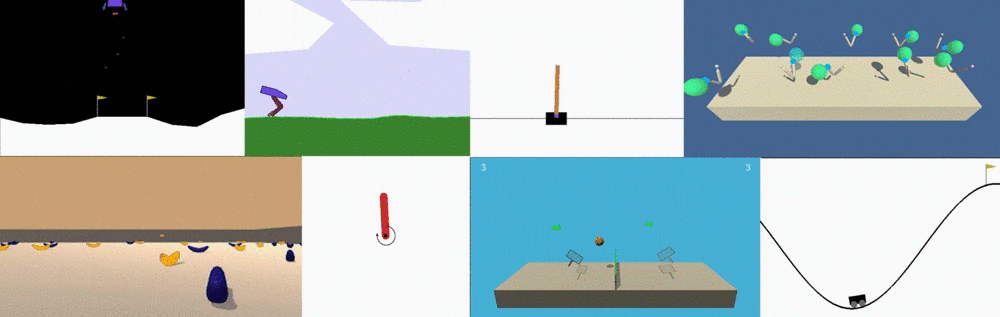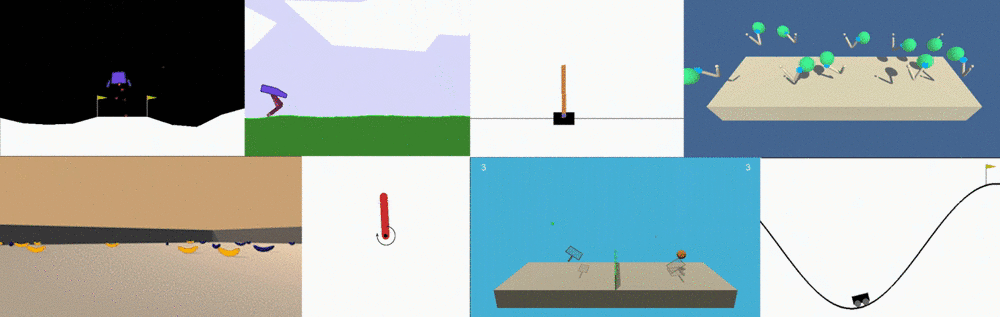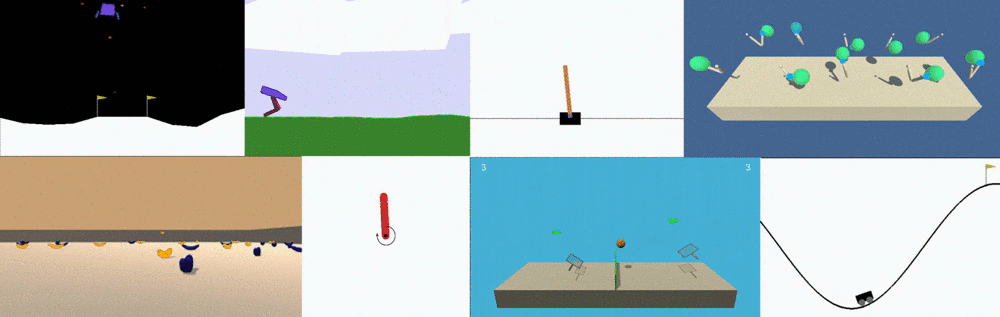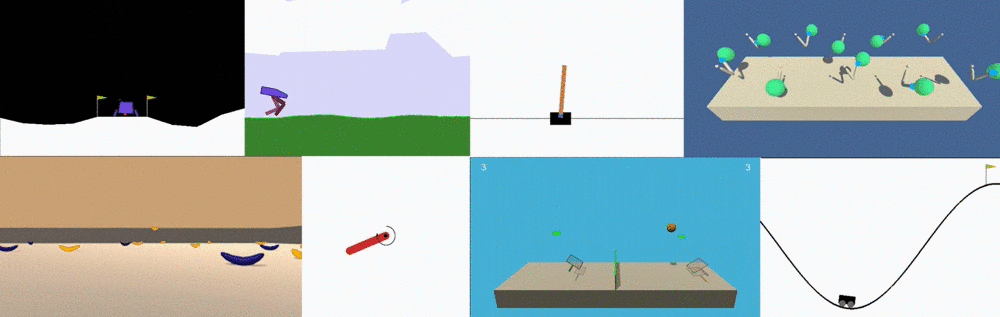
#
# ---
#  
#  
#
#
#
#
#
# + [markdown] id="4lYH6fsZOZNE" colab_type="text"
# # Q - Learning
# + [markdown] id="XlOSaEIGOhSy" colab_type="text"
# In Reinforcment Learning we want to obtain a function Q(s,a) that predicts best action a in states s in order to have maximum reward
# This function can be estimated using **Q-learning** , which iteratively updates Q(s,a) using the **Bellman Equation**
#
# *Where S - > state
# a - > Action*
#
#
# ```
# It is called Q-function, because it represents the “quality” of a certain action in a given state.
# Q values stored in table called Q- table (Quality table) which having 's' number of rows and 'a' number of columns
# ```
#
# 
#
# ---
#
#
# ** Q-learning Algorithm in form of flow chart**
#
# 
#
#
#
# *Reinforcement Learning libraries overview *
#
#
# 
# + [markdown] id="kiYRX0t7V41r" colab_type="text"
# # OpenAI Gym
# + [markdown] id="CzHymxxYZElP" colab_type="text"
# **[OpenAI Gym](https://gym.openai.com/) is a toolkit for developing and comparing reinforcement learning algorithms. This is the gym open-source library, which gives you access to a standardized set of environments.**
# + id="_3wtAyPl2tub" colab_type="code" outputId="e5230bb9-a305-4116-aec7-a38d1f258807" colab={"base_uri": "https://localhost:8080/", "height": 419}
#Installation of gym
# !pip install gym
# + id="aNz_1mM7Zyc3" colab_type="code" outputId="5a13b9fa-0470-4274-8873-e833d9b96f3a" colab={"base_uri": "https://localhost:8080/", "height": 14508}
#list of environments of gym
from gym import envs
envids = [spec.id for spec in envs.registry.all()]
for envid in sorted(envids):
print(envid)
# + id="tzXkRXluaRyV" colab_type="code" colab={}
import gym
# + id="M5jvhpOIciQr" colab_type="code" outputId="a4f2c0e4-36cf-4ebe-f8ac-27187568e782" colab={"base_uri": "https://localhost:8080/", "height": 219}
env = gym.make("Taxi-v2").env
env.render()
# + [markdown] id="vxwOn7aYc0dK" colab_type="text"
# **Explanation of Environment of Taxi-2**
#
#
# 
#
#
# * four different locations (R, G, Y, B)
# * Taxi environment has 5(row)×5(col)×5( 4+1= five passenger location)×4(destionation)=500 total possible states.
# * six possible actions: 0 = down, 1 = up, 2 = right, 3 = left, 4 = pickup, 5 = dropoff
#
#
#
#
#
#
# + id="tb3wJzmucr21" colab_type="code" outputId="8e0cbd71-429b-4a87-e563-84d91e61f327" colab={"base_uri": "https://localhost:8080/", "height": 54}
print(env.observation_space.n)
print(env.action_space.n)
# + id="qQH05Ywtgs7j" colab_type="code" outputId="c1258e15-432f-40ba-e6b4-a35d3570e87e" colab={"base_uri": "https://localhost:8080/", "height": 35}
env.reset()
env.step(2)
#Step the environment by one timestep. Returns
#observation: Observations of the environment
#reward: If your action was beneficial or not
#done: Indicates if we have successfully picked up and dropped off a passenger, also called one episode(one complete cycle)
#info: Additional info such as performance and latency for debugging purposes
# + id="hsNne9ljiNje" colab_type="code" outputId="f0979f57-25c9-4aa4-bf72-a1b92d5ff805" colab={"base_uri": "https://localhost:8080/", "height": 163}
env.render()
# + id="SJK4T89-iL8S" colab_type="code" outputId="0f5f431e-a95b-4d7b-a7a6-b0fd3f0a8a22" colab={"base_uri": "https://localhost:8080/", "height": 181}
#trying to reach at location same as given image
env.reset()
state = env.encode(3, 1, 2, 0) # (taxi row, taxi column, starting index(Blue color), destination index(Purple color))
print("State:", state)
env.s = state
env.render()
# + [markdown] id="arJZ48Tmjg-u" colab_type="text"
# # Self Drawing Cab with RL
# + [markdown] id="TqBZGL42juzF" colab_type="text"
# **Problem:** There are 4 locations (labeled by different letters), and our job is to pick up the passenger at one location and drop him off at another.
#
#
# * +20 points for successful drop-off
# * -1 point for every time-step it takes
# * -10 point penalty for illegal pick-up and drop-off actions
#
# **Conditions**
#
#
# * Save time of passenger
# * Take care of passenger
# * Drop him/her at right location
#
# Note: If car color "Yellow" then No passenger inside if "Green" then it having passenger
#
#
# + id="80GdBSPqhrlJ" colab_type="code" outputId="31dd8e57-d216-4e7c-ab00-862a230a8a64" colab={"base_uri": "https://localhost:8080/", "height": 35}
import numpy as np
q_table = np.zeros([env.observation_space.n, env.action_space.n])
q_table.shape
# + [markdown] id="01vibaNFlj_T" colab_type="text"
#
# * $\Large \alpha$ (alpha) is the learning rate (0<$\Large \alpha$<=1)
# * $\Large \gamma$ (gamma) is the discount factor ($0 \leq \gamma \leq 1$) - determines how much importance we want to give to future rewards. A high value for the discount factor (close to 1) captures the long-term effective award, whereas, a discount factor of 0 makes our agent consider only immediate reward, hence making it greedy.
#
#
#
# 
# + id="P4_nTvB4lPvY" colab_type="code" outputId="1b755940-0429-426d-99a7-cbde68dae275" colab={"base_uri": "https://localhost:8080/", "height": 54}
import random
from IPython.display import clear_output
# Hyperparameters
alpha = 0.1
gamma = 0.9
epsilon = 0.1
for i in range(1, 100001):
state = env.reset()
epochs, penalties, reward, = 0, 0, 0
done = False
while not done:
if random.uniform(0, 1) < epsilon:
action = env.action_space.sample() # Explore action space
else:
action = np.argmax(q_table[state]) # Exploit learned values
next_state, reward, done, info = env.step(action)
old_value = q_table[state, action]
next_max = np.max(q_table[next_state])
new_value = (1 - alpha) * old_value + alpha * (reward + gamma * next_max)
q_table[state, action] = new_value
if reward == -10:
penalties += 1
state = next_state
epochs += 1
if i % 100 == 0:
clear_output(wait=True)
print("Episode: ",i)
print("Training finished")
# + id="U5KZchhbleJ4" colab_type="code" outputId="04306a61-028a-43c3-b015-a721854fd122" colab={"base_uri": "https://localhost:8080/", "height": 54}
q_table[247] #check at value of q_table at perticular location #consider high
# + [markdown] id="4dqqXR0amTNJ" colab_type="text"
#
#
# > * Evaluate agent's performance after Q-learning
#
#
# + id="8Efg6L-umE39" colab_type="code" outputId="ffd77477-45dd-4f94-9607-210be2a53edf" colab={"base_uri": "https://localhost:8080/", "height": 72}
total_epochs, total_penalties = 0, 0
episodes = 100
for _ in range(episodes):
state = env.reset()
epochs, penalties, reward = 0, 0, 0
done = False
while not done:
action = np.argmax(q_table[state]) #mapping state to action
state, reward, done, info = env.step(action)
if reward == -10:
penalties += 1
epochs += 1
#print("Reward is:",reward)
total_penalties += penalties
total_epochs += epochs
print("Results after ",episodes," episodes:")
print("Average timesteps per episode:", (total_epochs / episodes))
print("Average penalties per episode:", (total_penalties / episodes))
# + [markdown] id="RJarz4U707W1" colab_type="text"
#
#
# > * As we can see from the evaluation, the agent's performance improved significantly and it incurred no penalties, which means it performed the correct pickup/dropoff actions with 100 different passengers.
#
#
#
#
# + id="F4r2wQ52maJY" colab_type="code" outputId="e01f855e-ff2e-40cb-a7f2-379c83c51b90" colab={"base_uri": "https://localhost:8080/", "height": 2054}
state = env.reset()
done = None
while done != True:
#take the action with the highest Q Value
action = np.argmax(q_table[state])
state, reward, done, info = env.step(action)
env.render()
# + [markdown] id="IsRT8-vamnPK" colab_type="text"
# # Deep Q Network
#
# Find the values of Q by Neural Netowork Approach
#
# 
#
# Left: Naive formulation of deep Q-network. Right: More optimized architecture of deep Q-network, used in DeepMind paper.
# + [markdown] id="SqT7ORCQ1x5g" colab_type="text"
# # Reference
#
#
#
# * [Reinforcement Learning:
# An Introduction](http://incompleteideas.net/book/bookdraft2018jan1.pdf)
# * [Material of ML/DL](https://goo.gl/3L9cGu)
#
#
# + [markdown] id="g8BTB-niSrPl" colab_type="text"
# # Thank You!
# Let's connect
# - [LinkedIn](https://www.linkedin.com/in/krupagaliya/)
# - [Twitter](https://twitter.com/Krupagaliya)
| cruft/devfestahm_rl.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sqlite3
from sqlite3 import Error
# +
def create_connection(db_file):
'''
create a database connection to the SQLite database
specified by the db_file
return: Connection object or None
'''
try:
conn = sqlite3.connect(db_file)
print("Sucessfully connected.")
return conn
except Error as e:
print(e)
return None
def select_all_records(conn):
'''
Query all rows in the people table
:param conn: the Connection object
:return: Nothing
'''
cur = conn.cursor()
cur.execute("SELECT * FROM people")
rows = cur.fetchall()
for row in rows:
print(row)
def show_table_names(conn):
'''
Prints list of all tables in databse
param: conn: the Connection object
return: Nothing
'''
cursor = conn.cursor()
cursor.execute("SELECT name FROM sqlite_master WHERE type='table';")
table_list = cursor.fetchall()
print(table_list)
def main():
database_name = "sample_db.db" #in present working dir
# create a database connection
conn = create_connection(database_name)
with conn:
print("1. Table names")
show_table_names(conn)
print("1. Get all records")
select_all_records(conn)
# -
if __name__ == '__main__':
main()
| test/tests/sql_api.ipynb |
# Transformers installation
# ! pip install transformers
# To install from source instead of the last release, comment the command above and uncomment the following one.
# # ! pip install git+https://github.com/huggingface/transformers.git
# # Training and fine-tuning
# Model classes in 🤗 Transformers are designed to be compatible with native PyTorch and TensorFlow 2 and can be used
# seamlessly with either. In this quickstart, we will show how to fine-tune (or train from scratch) a model using the
# standard training tools available in either framework. We will also show how to use our included
# `Trainer` class which handles much of the complexity of training for you.
#
# This guide assume that you are already familiar with loading and use our models for inference; otherwise, see the
# [task summary](https://huggingface.co/transformers/task_summary.html). We also assume that you are familiar with training deep neural networks in either
# PyTorch or TF2, and focus specifically on the nuances and tools for training models in 🤗 Transformers.
#
# Sections:
#
# - [pytorch](#pytorch)
# - [tensorflow](#tensorflow)
# - [trainer](#trainer)
# - [additional-resources](#additional-resources)
# <a id='pytorch'></a>
# ## Fine-tuning in native PyTorch
# Model classes in 🤗 Transformers that don't begin with `TF` are [PyTorch Modules](https://pytorch.org/docs/master/generated/torch.nn.Module.html), meaning that you can use them just as you would any
# model in PyTorch for both inference and optimization.
#
# Let's consider the common task of fine-tuning a masked language model like BERT on a sequence classification dataset.
# When we instantiate a model with `PreTrainedModel.from_pretrained`, the model configuration and
# pre-trained weights of the specified model are used to initialize the model. The library also includes a number of
# task-specific final layers or 'heads' whose weights are instantiated randomly when not present in the specified
# pre-trained model. For example, instantiating a model with
# `BertForSequenceClassification.from_pretrained('bert-base-uncased', num_labels=2)` will create a BERT model instance
# with encoder weights copied from the `bert-base-uncased` model and a randomly initialized sequence classification
# head on top of the encoder with an output size of 2. Models are initialized in `eval` mode by default. We can call
# `model.train()` to put it in train mode.
from transformers import BertForSequenceClassification
model = BertForSequenceClassification.from_pretrained('bert-base-uncased')
model.train()
# This is useful because it allows us to make use of the pre-trained BERT encoder and easily train it on whatever
# sequence classification dataset we choose. We can use any PyTorch optimizer, but our library also provides the
# `AdamW` optimizer which implements gradient bias correction as well as weight decay.
from transformers import AdamW
optimizer = AdamW(model.parameters(), lr=1e-5)
# The optimizer allows us to apply different hyperpameters for specific parameter groups. For example, we can apply
# weight decay to all parameters other than bias and layer normalization terms:
no_decay = ['bias', 'LayerNorm.weight']
optimizer_grouped_parameters = [
{'params': [p for n, p in model.named_parameters() if not any(nd in n for nd in no_decay)], 'weight_decay': 0.01},
{'params': [p for n, p in model.named_parameters() if any(nd in n for nd in no_decay)], 'weight_decay': 0.0}
]
optimizer = AdamW(optimizer_grouped_parameters, lr=1e-5)
# Now we can set up a simple dummy training batch using `PreTrainedTokenizer.__call__`. This returns
# a `BatchEncoding` instance which prepares everything we might need to pass to the model.
from transformers import BertTokenizer
tokenizer = BertTokenizer.from_pretrained('bert-base-uncased')
text_batch = ["I love Pixar.", "I don't care for Pixar."]
encoding = tokenizer(text_batch, return_tensors='pt', padding=True, truncation=True)
input_ids = encoding['input_ids']
attention_mask = encoding['attention_mask']
# When we call a classification model with the `labels` argument, the first returned element is the Cross Entropy loss
# between the predictions and the passed labels. Having already set up our optimizer, we can then do a backwards pass and
# update the weights:
labels = torch.tensor([1,0]).unsqueeze(0)
outputs = model(input_ids, attention_mask=attention_mask, labels=labels)
loss = outputs.loss
loss.backward()
optimizer.step()
# Alternatively, you can just get the logits and calculate the loss yourself. The following is equivalent to the previous
# example:
from torch.nn import functional as F
labels = torch.tensor([1,0])
outputs = model(input_ids, attention_mask=attention_mask)
loss = F.cross_entropy(outputs.logits, labels)
loss.backward()
optimizer.step()
# Of course, you can train on GPU by calling `to('cuda')` on the model and inputs as usual.
#
# We also provide a few learning rate scheduling tools. With the following, we can set up a scheduler which warms up for
# `num_warmup_steps` and then linearly decays to 0 by the end of training.
from transformers import get_linear_schedule_with_warmup
scheduler = get_linear_schedule_with_warmup(optimizer, num_warmup_steps, num_train_steps)
# Then all we have to do is call `scheduler.step()` after `optimizer.step()`.
loss.backward()
optimizer.step()
scheduler.step()
# We highly recommend using `Trainer`, discussed below, which conveniently handles the moving parts
# of training 🤗 Transformers models with features like mixed precision and easy tensorboard logging.
# ### Freezing the encoder
# In some cases, you might be interested in keeping the weights of the pre-trained encoder frozen and optimizing only the
# weights of the head layers. To do so, simply set the `requires_grad` attribute to `False` on the encoder
# parameters, which can be accessed with the `base_model` submodule on any task-specific model in the library:
for param in model.base_model.parameters():
param.requires_grad = False
# <a id='tensorflow'></a>
# ## Fine-tuning in native TensorFlow 2
# Models can also be trained natively in TensorFlow 2. Just as with PyTorch, TensorFlow models can be instantiated with
# `PreTrainedModel.from_pretrained` to load the weights of the encoder from a pretrained model.
from transformers import TFBertForSequenceClassification
model = TFBertForSequenceClassification.from_pretrained('bert-base-uncased')
# Let's use `tensorflow_datasets` to load in the [MRPC dataset](https://www.tensorflow.org/datasets/catalog/glue#gluemrpc) from GLUE. We can then use our built-in
# `glue_convert_examples_to_features` to tokenize MRPC and convert it to a
# TensorFlow `Dataset` object. Note that tokenizers are framework-agnostic, so there is no need to prepend `TF` to
# the pretrained tokenizer name.
from transformers import BertTokenizer, glue_convert_examples_to_features
import tensorflow as tf
import tensorflow_datasets as tfds
tokenizer = BertTokenizer.from_pretrained('bert-base-uncased')
data = tfds.load('glue/mrpc')
train_dataset = glue_convert_examples_to_features(data['train'], tokenizer, max_length=128, task='mrpc')
train_dataset = train_dataset.shuffle(100).batch(32).repeat(2)
# The model can then be compiled and trained as any Keras model:
optimizer = tf.keras.optimizers.Adam(learning_rate=3e-5)
loss = tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True)
model.compile(optimizer=optimizer, loss=loss)
model.fit(train_dataset, epochs=2, steps_per_epoch=115)
# With the tight interoperability between TensorFlow and PyTorch models, you can even save the model and then reload it
# as a PyTorch model (or vice-versa):
from transformers import BertForSequenceClassification
model.save_pretrained('./my_mrpc_model/')
pytorch_model = BertForSequenceClassification.from_pretrained('./my_mrpc_model/', from_tf=True)
# <a id='trainer'></a>
# ## Trainer
# We also provide a simple but feature-complete training and evaluation interface through `Trainer`
# and `TFTrainer`. You can train, fine-tune, and evaluate any 🤗 Transformers model with a wide range
# of training options and with built-in features like logging, gradient accumulation, and mixed precision.
# +
## PYTORCH CODE
from transformers import BertForSequenceClassification, Trainer, TrainingArguments
model = BertForSequenceClassification.from_pretrained("bert-large-uncased")
training_args = TrainingArguments(
output_dir='./results', # output directory
num_train_epochs=3, # total # of training epochs
per_device_train_batch_size=16, # batch size per device during training
per_device_eval_batch_size=64, # batch size for evaluation
warmup_steps=500, # number of warmup steps for learning rate scheduler
weight_decay=0.01, # strength of weight decay
logging_dir='./logs', # directory for storing logs
)
trainer = Trainer(
model=model, # the instantiated 🤗 Transformers model to be trained
args=training_args, # training arguments, defined above
train_dataset=train_dataset, # training dataset
eval_dataset=test_dataset # evaluation dataset
)
# +
## TENSORFLOW CODE
from transformers import TFBertForSequenceClassification, TFTrainer, TFTrainingArguments
model = TFBertForSequenceClassification.from_pretrained("bert-large-uncased")
training_args = TFTrainingArguments(
output_dir='./results', # output directory
num_train_epochs=3, # total # of training epochs
per_device_train_batch_size=16, # batch size per device during training
per_device_eval_batch_size=64, # batch size for evaluation
warmup_steps=500, # number of warmup steps for learning rate scheduler
weight_decay=0.01, # strength of weight decay
logging_dir='./logs', # directory for storing logs
)
trainer = TFTrainer(
model=model, # the instantiated 🤗 Transformers model to be trained
args=training_args, # training arguments, defined above
train_dataset=tfds_train_dataset, # tensorflow_datasets training dataset
eval_dataset=tfds_test_dataset # tensorflow_datasets evaluation dataset
)
# -
# Now simply call `trainer.train()` to train and `trainer.evaluate()` to evaluate. You can use your own module as
# well, but the first argument returned from `forward` must be the loss which you wish to optimize.
#
# `Trainer` uses a built-in default function to collate batches and prepare them to be fed into the
# model. If needed, you can also use the `data_collator` argument to pass your own collator function which takes in the
# data in the format provided by your dataset and returns a batch ready to be fed into the model. Note that
# `TFTrainer` expects the passed datasets to be dataset objects from `tensorflow_datasets`.
#
# To calculate additional metrics in addition to the loss, you can also define your own `compute_metrics` function and
# pass it to the trainer.
# +
from sklearn.metrics import accuracy_score, precision_recall_fscore_support
def compute_metrics(pred):
labels = pred.label_ids
preds = pred.predictions.argmax(-1)
precision, recall, f1, _ = precision_recall_fscore_support(labels, preds, average='binary')
acc = accuracy_score(labels, preds)
return {
'accuracy': acc,
'f1': f1,
'precision': precision,
'recall': recall
}
# -
# Finally, you can view the results, including any calculated metrics, by launching tensorboard in your specified
# `logging_dir` directory.
# <a id='additional-resources'></a>
# ## Additional resources
# - [A lightweight colab demo](https://colab.research.google.com/drive/1-JIJlao4dI-Ilww_NnTc0rxtp-ymgDgM?usp=sharing)
# which uses `Trainer` for IMDb sentiment classification.
#
# - [🤗 Transformers Examples](https://github.com/huggingface/transformers/tree/master/examples) including scripts for
# training and fine-tuning on GLUE, SQuAD, and several other tasks.
#
# - [How to train a language model](https://colab.research.google.com/github/huggingface/blog/blob/master/notebooks/01_how_to_train.ipynb), a detailed
# colab notebook which uses `Trainer` to train a masked language model from scratch on Esperanto.
#
# - [🤗 Transformers Notebooks](https://huggingface.co/transformers/notebooks.html) which contain dozens of example notebooks from the community for
# training and using 🤗 Transformers on a variety of tasks.
| transformers_doc/training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import h2o
from h2o.estimators.gbm import H2OGradientBoostingEstimator
from h2o.grid.grid_search import H2OGridSearch
h2o.init()
DATAFILE = "/mnt/fs-h2o/Telco-Customer-Churn.csv"
data = h2o.import_file(DATAFILE, destination_frame= "gbm_data")
data.head()
data.columns
# mark as categorical
data['SeniorCitizen'] = data['SeniorCitizen'].asfactor()
# build the list of features I want to use (excluding customeriD from list)
features = ['gender',
'SeniorCitizen',
'Partner',
'Dependents',
'tenure',
'PhoneService',
'MultipleLines',
'InternetService',
'OnlineSecurity',
'OnlineBackup',
'DeviceProtection',
'TechSupport',
'StreamingTV',
'StreamingMovies',
'Contract',
'PaperlessBilling',
'PaymentMethod',
'MonthlyCharges',
'TotalCharges']
train, valid = data.split_frame([0.85], seed=1234, destination_frames= ["gbm_train",
"gbm_valid"])
# +
# grid search for hyper-parameter optimization
# +
# this is one possible set of params, chosen after some initial trials. We can add more with longer exec time
ntrees_opt = [10, 20, 25, 30, 40, 50]
max_depth_opt = [3,4,5,6,7]
learn_rate_opt = [0.01, 0.05, 0.1, 0.2, 0.3, 0.5]
hyper_parameters = {"ntrees": ntrees_opt, "max_depth": max_depth_opt, "learn_rate": learn_rate_opt}
# -
gs = H2OGridSearch(H2OGradientBoostingEstimator, hyper_params=hyper_parameters)
# important to give an ID to the model, to make easier to find it in H2O Flow
# %time gs.train(x = features, y = 'Churn', training_frame = train, validation_frame = valid, model_id = "gbm_churn")
# sort the grid in order of decreasing recall (top is higher recall)
grid = gs.get_grid(sort_by="recall", decreasing=True)
grid
# this way we get the Top Model !
best_gbm = grid.models[0]
# let's see model's characteristics
best_gbm
# ### Train the model using parameters identified.
| H2O-GBM-Churn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import numpy as np
from scripts import m2
data = pd.read_csv("/Users/Mason/cosc301/course-project-solo_6023/data/processed/all_alpha_20.csv")
# -
fig=plt.figure(figsize=(14,8), dpi= 100, facecolor='w', edgecolor='k')
countdata = sns.countplot(data=data, x="Type", palette = "crest", order= data['Type'].value_counts().index)
countdata.set(title="Number of 2020 Vehicles by vehicle type", xlabel="Type of Vehicle", ylabel= "Count of Vehciels")
fig=plt.figure(figsize=(14,8), dpi= 100, facecolor='w', edgecolor='k')
countdata = sns.countplot(data=data, x="Fuel", palette = "mako")
countdata.set(title="Number of 2020 Vehicles by fuel type", xlabel="Fuel Type", ylabel= "Number of Models")
fig=plt.figure(figsize=(14,8), dpi= 100, facecolor='w', edgecolor='k')
countdata = sns.regplot(data=data, x="Displacement", y="Combined MPG", logx=True, scatter_kws={"color": "mediumturquoise"}, line_kws={"color": "darkcyan"})
countdata.set(title="Relation between MPG and engine size", xlabel= "Displacement (in Liters)")
fig=plt.figure(figsize=(14,8), dpi= 100, facecolor='w', edgecolor='k')
countdata = sns.stripplot(data=data, x="Combined MPG", y="Type", orient="h", palette="crest", order = data['Type'].value_counts().index)
countdata.set(title="Fuel economy by vehicle type", xlabel= "Combined MPG")
| analysis/Graphs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# I was asked to revisit the difference between function declaration and calling, class declaration and instantiation and method declaration and calling.
#
# We use `def` to declare a function. What follows is what we call the _function signature_, then a `:` and then the _function body_ which is indented by four spaces and often ends with a `return` statement:
def square(x):
return x**2
# Using the function signature we can also call the function. The arguments declared in the signature are then used to process and we get back whatever we evaluate after the `return`:
square(2)
# The `class` keyword introduces a similar syntax with very different meaning. Like for a function we give a name to the class. By convention this name is capitalized.
#
# Anything following in parentheses is not a parameter, but a so called _superclass_ (or _parent class_). In fact, all objects in Python inherit from `object`. The part in parentheses in `class Classname(object)` is usually omitted.
#
# The parameters we call upon _instantiation_ of a class are found in the special method `__init__`. Special methods are methods with leading and trailing double underscores. Python uses those for internal purposes, they can be defined but should not be called directly.
#
# As `__init__` is a method, it has access to a `self` parameter. If we call the class name (_instantiation_) we create a **new** object of this class. The `__init__` method forges this new object into the shape of the class. The class is the blueprint for any new instance of it.
# +
class Powerizer(object):
def __init__(self, to):
self.to = to
def power(self, x):
return x**self.to
def square(self, x):
return x**2
p = Powerizer(to=3)
# -
# In the example class we create objects which are _powerizers_ and carry a state in the attribute `to`. Any two instances of this class can carry different state in `to`.
p.to
# A method is very similar to a function. The key differences are, that methods live inside classes and their first argument is always `self`. This first argument is handed to the method by Python itself. We only ever need to call methods with the parameters following this `self`.
#
# Using `self`, the method has access to the individual state an object carries. For instance, `p` has its `to` attribute set to $3$. The `power` method raises its argument to the power of `to` (in the case of `p` this is the power of $3$).
#
# Notice also the `p.square` method. It is similar to the `square` function in what it does. It would have access to the state of the `Powerizer` instance it is called on, we just chose to ignore this `self`.
p.power(3)
# # The Zen of Python
# As we have seen throughout this semester, there are often many paths to the same result in Python. However, they are rarely equal. An order of elegance and _pythonism_ separates them.
#
# In times of distress we can turn to the _Zen of Python_ for guidance among these paths. You can summon it with
#
# ~~~python
# import this
# ~~~
import this
# We've already seen namespaces to great extent. Here is the summary:
#
# + To **look** into a namespace use the `dir(namespace)` function
# + To **move** into a namespace use the dot: `.`
# + Namespaces are available on all Python objects, including modules
# + There is a global namespace which you can discover by calling `dir()` without arguments
import numpy as np
np.square([2,3,4,5])
# ## Identity and equality
# Reconsider the last part of the Pre-Exercise 11. In certain cases there certainly could be two distinct fungi at the same position. The description disregards such cases on purpose, of course, as to not complicate the matter. But what if we wanted to know if a fungus merely grows at the same spot or is actually the very same fungus as another?
#
# In other words: How do we compare objects for (in-)equality and identity?
class A:
pass
a1 = A()
id(a1)
# As we can see, even an object of a completely empty class `A` has an identity. This identity can be queried using the `id` function.
a2 = A()
id(a2)
# Another object of the same class does not have the same `id`. So surely they are not the same:
a2 is a1
# But what about objects that can be equal while they are not the same?
#
# Consider the class `Circle` which has a radius. Two circles are equal if the have the same radius.
# +
class Circle:
def __init__(self, radius):
self.radius = radius
c1 = Circle(1)
c2 = Circle(1)
# -
# iIf we want to compare for equality, rather than identity, we already have learnt to use `==`.
#
# The following should yield `True` given our reasoning:
c1 == c2, c1 is c2
# What has happened here?
#
# Behind the scenes Python uses identity as equality if we don't tell it otherwise. Here is how we can define our own equality.
#
# It works very similar to how we defined our `distance` method above:
# +
class Circle:
def __init__(self, radius):
self.radius = radius
def __eq__(self, other):
return self.radius == other.radius
c1 = Circle(1)
c2 = Circle(1)
c3 = c2
# -
# Now we can distinguish if two circles really **are** the same, or if they merely satisfy our conditions of equality.
#
# This can be especially useful if we want to treat the two circles differently later on.
c1 == c3, \
c1 is c3, \
c2 is c3, \
c2 == c3
# ## Mutability and its pitfalls
# The issue of identity is especially pressing when mutating objects. Lists e.g. are mutable:
# +
l1 = list()
l2 = l1 # assign l1 to l2
l1.append('a')
# appending on l2 also appends on l1 -> They are the same object!
print(l1)
print(l2)
l1 is l2 # This is the reason they seem to both change
# -
# It may seem intuitive, that if we say `l1 = l2` we get this result. But this does not work if we re-assign rather than mutate. For example with integers:
i = 1
j = i
j += 1
i,j
# or with tuples:
t = (1,2)
u = t
u += (3,)
t,u
# This is why lists being mutable and tuples being immutable is such an important thing to know in Python.
#
# In more obscure use cases like the following, the mutability of lists (and object attributes in Python in general) can catch us off guard:
# +
li = [list()] * 3 # create a list of three empty lists
print(li) # check the result is correct
li[0].append('a') # append `1` to the first list in the super-list
print(li) # what happened?
# -
# Now we can also understand better how list comprehensions work:
# +
li = [list() for _ in range(3)]
print(li)
li[0].append('a') # append `1` to the first list in the super-list
print(li) # what happened?
# -
# ## Multiple inheritance
# We now know that a class `B` can inherit methods from a class `A` by defining it as `B(A)`. But what if another class `C` inherits from `B`? Will it also inherit from `A`?
# +
class Animal:
lifepoints = 63
class Rabbit(Animal):
pass
class Wolf(Animal):
pass
wolf1 = Wolf()
wolf1.lifepoints # it does!
# -
# This was foreseeable. Let's have a look at a more interesting case: *Multiple inheritance*
#
# Let's say we have two separate classes:
# + `Predator` defines an attribute `lifepoints` as `63`
# + `Prey` defined an attribute, also named `lifepoints`, as `42`
#
# The class `Fox` inherits from both `Predator` and `Prey`, just as `Hog` but in reverse order:
# +
class Predator:
lifepoints = 63
class Prey:
lifepoints = 42
class Fox(Predator, Prey):
pass
class Hog(Prey, Predator):
pass
fox1 = Fox()
hog1 = Hog()
fox1.lifepoints, hog1.lifepoints
# -
# We can observe, that the order of inheritance is in fact important!
#
# If we want to have a look at the order at which attribute and method names are looked up, we can use the `__mro__` attribute of a class:
Fox.__mro__
Hog.__mro__
# Just above I mentioned a method or attribute can be *overridden*. Let's see how that works.
#
# Say in the last example we also want to add a fusion power plant. It is like a normal power plant, but we need to know how much hydrogen it uses:
# +
class PowerPlant:
def __init__(self, f, out):
self.f = f
self.out = out
self.inoperation = 'no'
self.shape = '.'
def emissions(self):
return self.f * self.out
def turn_on(self):
self.inoperation = 'yes'
def draw(self):
plt.scatter(rd.randint(0,5), rd.randint(0,5)
, marker=self.shape
, s = self.out / 10
, color = 'red' if self.inoperation == 'no' else 'green'
)
class FusionPlant(PowerPlant):
def __init__(self, hydrogen_demand):
self.hydrogen_demand = hydrogen_demand
fp = FusionPlant(300)
fp.hydrogen_demand
# -
# That seems to work, but the mindful reader may have noticed: We do not initialize `PowerPlant` proper!
#
# We can observe this easily:
fp.shape, fp.out. fp.f
# So how would we do this? We call `super` !
# +
class FusionPlant(PowerPlant):
def __init__(self, f, out, hydrogen_demand):
super(FusionPlant, self).__init__(f, out)
self.hydrogen_demand = hydrogen_demand
fp = FusionPlant(0.1, 10_000, 300)
fp.hydrogen_demand, fp.out, fp.shape
| jupyter/Exercises 13.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# variables are defined with =
x = 4 # running this code has no output, but the value is stored.
print(x)
y = x**2
print(y)
x = x**2
print(x)
print(y) # doesn't change value of y
# for loops are fun.
for i in range(5):
print(i)
# if statement
if 3 > 0:
print("Yes!")
# +
# inside/outside loop is determined by indentation
counter = 0
for i in range(10,20): #i starts at 10 , ends at 19. Think half-open interval [10, 20)
if i % 2 == 0: # checks if i is even. Think i mod 2 or remainder when i is divided by 2.
counter += 1 # shorthand for counter = counter + 1
print("this is i: {}, this is counter: {}".format(i,counter)) # prints variable values inside string
# print(f"this is i: {i}, this is counter: {counter}) # f-string works in python 3.6+
print(counter)
# -
#What will this code output? Why?
for i in range(3,8):
counter = 0
for j in range(6):
if i == j:
counter +=1
print(counter)
# +
# fix the above code so that the counter actually counts how many times i equals j
# +
# write a loop that prints the squares of 3,4,5,6
# +
# write a triple loop that counts how often i > j > k where 0 <= i <= 6, 4 <= j <= 8, 2 <= k <= 5. Answer should be 7.
# don't forget colons at the end of for/if statements.
# if statement condition can be written as:
# i > j > k
# or
# i > j and j > k
# -
# write a loop that prints the next 10 numbers of the fibonacci sequence given the two starting values. Harder.
x=1
y=1
print(x)
print(y)
| basics2_loops.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# ### Sigmoid Layer
from IPython.display import Image
Image("sigmoid.png")
# ### Artifical Neural Networks
Image("nn.PNG")
# ### Back Propogation
Image("backprop.png")
# conda update scikit-learn
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
from sklearn.neural_network import MLPClassifier
from sklearn.model_selection import train_test_split
# %matplotlib inline
data=pd.read_csv('mnist.csv')
df_x=data.iloc[:,1:]
df_y=data.iloc[:,0]
x_train, x_test, y_train, y_test = train_test_split(df_x, df_y, test_size=0.2, random_state=4)
nn=MLPClassifier(activation='logistic',solver='sgd',hidden_layer_sizes=(10,15),random_state=1)
nn.fit(x_train,y_train)
pred=nn.predict(x_test)
#activation logistic with hidden layer sizes-> 45,90 Gave 92 % accuracy
#activation relu with hidden layer sizes-> 45,90 gave 89 % accuracy
#Test with different combinatations of learning rate, activation and other hyper parameters and mesure the accuracy
a=y_test.values
a
count=0
for i in range(len(pred)):
if pred[i]==a[i]:
count=count+1
count
len(pred)
6824/8400.0
| Data_Science/Neural Networks and BackPropogation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Comparing CLMM to NumCosmo
# This notebook compares `CLMM` to `NumCosmo`.
import numpy as np
import cluster_toolkit as ct
from astropy import constants, cosmology, units
import math
# For NumCosmo
import gi
gi.require_version('NumCosmo', '1.0')
gi.require_version('NumCosmoMath', '1.0')
from gi.repository import NumCosmo as Nc
from gi.repository import NumCosmoMath as Ncm
# +
import sys
import os
os.environ['CLMM_MODELING_BACKEND'] = 'ccl'
from clmm import theory as pp
# -
# ## Configuration
# ### Cosmology
# +
# If CCL is installed, define a CCL cosmology object
# import pyccl as ccl
# cosmo_ccl = ccl.Cosmology(Omega_c=0.27, Omega_b=0.045, h=0.67, A_s=2.1e-9, n_s=0.96)
# If CCL is not installed, use astropy instead with the following lines
#from astropy.cosmology import FlatLambdaCDM
#astropy_cosmology_object = FlatLambdaCDM(H0=70, Om0=0.27, Ob0=0.045)
#cosmo_ccl = pp.cclify_astropy_cosmo(astropy_cosmology_object)
cosmo_ccl = pp.Cosmology (H0 = 70.0, Omega_dm0 = 0.27 - 0.045, Omega_b0 = 0.045)
# -
# ### Cluster/source config
# Select density profile and profile parametrization options
density_profile_parametrization = 'nfw'
mass_Delta = 200 # Mass defintion: mean matter density
cluster_mass = 1.0e15 # Msun
cluster_concentration = 4.0
z_max = 3.0
z_cluster = 1.0
z_source = 2.0
# ### Vectors for inputs
z_a = np.linspace (0.01, 3.0, 1000)
a_a = 1.0 / (1.0 + z_a)
r3d = np.logspace (-2, 2, 100)
# ### NumCosmo
# Pass configs to `NumCosmo`
# +
#########################
# Initializing NumCosmo #
#########################
Ncm.cfg_init ()
##################
# Set Cosmology ##
##################
cosmo = Nc.HICosmo.new_from_name (Nc.HICosmo, "NcHICosmoDEXcdm")
cosmo.omega_x2omega_k ()
cosmo.props.w = -1.0
cosmo.props.H0 = cosmo_ccl['H0']
cosmo.param_set_by_name ("H0", cosmo_ccl['H0'])
cosmo.param_set_by_name ("Omegab", cosmo_ccl['Omega_b0'])
cosmo.param_set_by_name ("Omegac", cosmo_ccl['Omega_dm0'])
cosmo.param_set_by_name ("w", -1.0)
cosmo.param_set_by_name ("Omegak", 0.0)
cosmo.param_set_by_name ("Tgamma0", 0.0)
##########################
# Config Cluster/Source ##
##########################
# NumCosmo Density profile
dp = Nc.HaloDensityProfileNFW.new (Nc.HaloDensityProfileMassDef.MEAN, mass_Delta)
dp.props.log10MDelta = math.log10 (cluster_mass)
dp.props.cDelta = cluster_concentration
dist = Nc.Distance.new (z_max)
smd = Nc.WLSurfaceMassDensity.new (dist)
smd.prepare (cosmo)
# -
# ## Comparisons
# ### Function to make automatic comparisons
import pylab as plt
def compare(x, y_clmm, y_nc, x_name='x', y_name='func', subplots_pars={'figsize':(6, 4)}):
clmm_name, nc_name = '%s_{clmm}'%y_name, '%s_{nc}'%y_name
diff = y_nc/y_clmm - 1.
print('rel diff min: %g\nrel diff max: %g'%(min(abs(diff)), max(abs(diff))))
fig, axs = plt.subplots(2, sharex=True, **subplots_pars)
fig.subplots_adjust(hspace=0)
axs[0].plot(x, y_clmm, label='clmm', lw=3)
axs[0].plot(x, y_nc, label='nc')
axs[1].plot(x, diff, c='r')
axs[1].axhline(0, ls=':', c='0', lw=.5)
axs[0].legend()
axs[0].set_ylabel('$%s$'%y_name)
axs[1].set_xlabel('$%s$'%x_name)
axs[1].set_ylabel('$%s/%s-1$'%(nc_name, clmm_name))
return fig, axs
# ### Quick test of all theory function
# +
Da_a = cosmo_ccl.eval_da_a1a2(a_a)
nc_Da_a = np.array ([dist.angular_diameter (cosmo, z) * cosmo.RH_Mpc () for z in z_a])
fig, axs = compare (z_a, Da_a, nc_Da_a, x_name='redshift', y_name='D_a(z)')
axs[0].set_yscale ('log')
# +
rho = pp.compute_3d_density (r3d, mdelta = cluster_mass, cdelta = cluster_concentration,
z_cl = z_cluster, cosmo = cosmo_ccl, delta_mdef = mass_Delta)
nc_rho = np.array ([dp.eval_density (cosmo, r3d_i, z_cluster) for r3d_i in r3d])
fig, axs = compare(r3d, rho, nc_rho, x_name='radius [Mpc]', y_name='\\rho')
axs[0].set_xscale('log')
# +
Sigma = pp.compute_surface_density (r3d, cluster_mass, cluster_concentration, z_cl=z_cluster,
cosmo=cosmo_ccl, delta_mdef=mass_Delta,
halo_profile_model='nfw')
nc_Sigma = np.array ([smd.sigma (dp, cosmo, r3d_i, z_cluster) for r3d_i in r3d])
fig, axs = compare (r3d, Sigma, nc_Sigma, x_name='radius [Mpc]', y_name='\\Sigma')
axs[0].set_xscale ('log')
# +
DeltaSigma = pp.compute_excess_surface_density(r3d, cluster_mass, cluster_concentration,
z_cl=z_cluster, cosmo=cosmo_ccl, delta_mdef=mass_Delta,
halo_profile_model='nfw')
nc_SigmaMean = np.array ([smd.sigma_mean (dp, cosmo, r3d_i, z_cluster) for r3d_i in r3d])
nc_DeltaSigma = nc_SigmaMean - nc_Sigma
fig, axs = compare(r3d, DeltaSigma, nc_DeltaSigma, x_name='radius [Mpc]', y_name=r'\Delta\Sigma')
axs[0].set_xscale('log')
# +
Sigmac = pp.compute_critical_surface_density (cosmo_ccl, z_cluster = z_cluster, z_source = z_source)
nc_Sigmac = smd.sigma_critical (cosmo, z_source, z_cluster, z_cluster)
print ("% 22.15g % 22.15g %e" % (Sigmac, nc_Sigmac, Sigmac / nc_Sigmac - 1.0))
# +
gammat = pp.compute_tangential_shear(r3d, mdelta=cluster_mass, cdelta=cluster_concentration, z_cluster=z_cluster,
z_source=z_source, cosmo=cosmo_ccl, delta_mdef=mass_Delta,
halo_profile_model='nfw', z_src_model='single_plane')
nc_gammat = np.array ([smd.shear (dp, cosmo, r3d_i, z_source, z_cluster, z_cluster) for r3d_i in r3d])
fig, axs = compare(r3d, gammat, nc_gammat, x_name='radius [mpc]', y_name=r'{\gamma_t}')
axs[0].set_xscale('log')
# +
kappa = pp.compute_convergence(r3d, mdelta=cluster_mass, cdelta=cluster_concentration,
z_cluster=z_cluster, z_source=z_source,
cosmo=cosmo_ccl, delta_mdef=mass_Delta,
halo_profile_model='nfw',
z_src_model='single_plane')
nc_kappa = np.array ([smd.convergence (dp, cosmo, r3d_i, z_source, z_cluster, z_cluster) for r3d_i in r3d])
fig, axs = compare(r3d, kappa, nc_kappa, x_name='radius [mpc]', y_name=r'\kappa')
axs[0].set_xscale('log')
# +
gt = pp.compute_reduced_tangential_shear(r3d, mdelta=cluster_mass, cdelta=cluster_concentration,
z_cluster=z_cluster, z_source=z_source, cosmo=cosmo_ccl, delta_mdef=mass_Delta,
halo_profile_model='nfw', z_src_model='single_plane')
nc_gt = np.array ([smd.reduced_shear (dp, cosmo, r3d_i, z_source, z_cluster, z_cluster) for r3d_i in r3d])
fig, axs = compare(r3d, gt, nc_gt, x_name='radius [mpc]', y_name=r'{g_t}')
axs[0].set_xscale('log')
axs[0].set_yscale('symlog')
# -
nc_mu = np.array ([smd.magnification (dp, cosmo, r3d_i, z_source, z_cluster, z_cluster) for r3d_i in r3d])
# ### Save the numcosmo outputs
np.savetxt('radial_profiles.txt',
np.array([r3d, nc_rho, nc_Sigma, nc_DeltaSigma, nc_gammat, nc_kappa, nc_gt, nc_mu]).T,
header='r3d rho Sigma DeltaSigma gammat kappa gt mu')
np.savetxt('angular_diameter_distance.txt',
np.array([a_a, z_a, nc_Da_a]).T,
header='a_a z_a Da_a')
# +
config_dict = {'dl': dist.angular_diameter (cosmo, z_cluster) * cosmo.RH_Mpc (),
'z_source': z_source,
'cluster_mass': cluster_mass,
'G[m3/km.s2]': Ncm.C.G(),
'aexp_cluster': 1./(1.+z_cluster),
'density_profile_parametrization': density_profile_parametrization,
'ds': dist.angular_diameter (cosmo, z_source) * cosmo.RH_Mpc (),
'cosmo_Ob0': cosmo_ccl['Omega_b0'],
'aexp_source': 1./(1.+z_source),
'dsl': dist.angular_diameter_z1_z2 (cosmo, z_cluster, z_source) * cosmo.RH_Mpc (),
'z_cluster': z_cluster,
'mass_Delta': mass_Delta,
'lightspeed[km/s]': Ncm.C.c()/1000.,
'cosmo_H0': cosmo_ccl['H0'],
'cluster_concentration': cluster_concentration,
'cosmo_Om0': cosmo_ccl['Omega_m0'],
'nc_Sigmac': nc_Sigmac,
'Msun[kg]': Ncm.C.mass_solar(),
'pc_to_m': Ncm.C.pc()}
import json
with open('config.json', 'w') as fp:
json.dump(config_dict, fp)
# -
| examples/NumCosmo/modeling_cmp_numcosmo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7 (DL)
# language: python
# name: dl
# ---
# # Chapter 12 - Principal Components Analysis with scikit-learn
#
# This notebook contains code accompanying Chapter 12 Principal Components Analysis with scikit-learn in *Practical Discrete Mathematics* by <NAME> and <NAME>.
#
# ## Eigenvalues and eigenvectors, orthogonal bases
#
# ### Example: Pizza nutrition
import pandas as pd
dataset = pd.read_csv('pizza.csv')
dataset.head()
# ### Example: Computing eigenvalues and eigenvectors
import numpy as np
A = np.array([[3,1], [1,3]])
l, v = np.linalg.eig(A)
print("The eigenvalues are:\n ",l)
print("The eigenvectors are:\n ", v)
# ## The scikit-learn implementation of PCA
#
# We will start by importing the dataset and then dropping the brand column from it. This is done to make sure that all our feature variables are numbers and hence can be scaled/normalized. We will then create another variable called target which will contain the names of the brands of pizzas.
# +
import pandas as pd
dataset = pd.read_csv('pizza.csv')
#Dropping the brand name column before standardizing the data
df_num = dataset.drop(["brand"], axis=1)
# Setting the brand name column as the target variable
target = dataset['brand']
# -
# Now that we have the dataset in order, we will then normalize the columns of the dataset to make sure that the mean for a variable is 0 and the variance is 1 and then we will run PCA on the dataset.
# +
#Scaling the data
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
scaler.fit(df_num)
scaled_data = scaler.transform(df_num)
#Applying PCA to the scaled data
from sklearn.decomposition import PCA
#Reducing the dimesions to 2 components so that we can have a 2D visualization
pca = PCA(n_components = 2)
pca.fit(scaled_data)
#Applying to our scaled dataset
scaled_data_pca = pca.transform(scaled_data)
#Check the shape of the original dataset and the new dataset
print("The dimensions of the original dataset is: ", scaled_data.shape)
print("The dimensions of the dataset after performing PCA is: ", scaled_data_pca.shape)
# -
# Now we have reduced our 7-dimensional dataset to its 2 principal components as can be seen from the dimensions shown above. We will move forward with plotting the principal components to check whether 2 principal components were enough to capture the variability in the dataset – the different nutritional content of pizzas produced by different companies.
# +
#Plotting the principal components
import matplotlib.pyplot as plt
import seaborn as sns
sns.scatterplot(scaled_data_pca[:,0], scaled_data_pca[:,1], target)
plt.legend(loc="best")
plt.gca().set_aspect("equal")
plt.xlabel("Principal Component 1")
plt.ylabel("Principal Component 2")
plt.show()
# -
# Now, we will move on to perform PCA in a way where we do not choose the number of desired principal components, rather we choose the number of principal components that add up to a certain desired variance. The Python implementation of this is very similar to the previous way with very slight changes to the code as shown below.
# +
import pandas as pd
dataset = pd.read_csv('pizza.csv')
#Dropping the brand name column before standardizing the data
df_num = dataset.drop(["brand"], axis=1)
# Setting the brand name column as the target variable
target = dataset['brand']
#Scaling the data (Step 1)
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
scaler.fit(df_num)
scaled_data = scaler.transform(df_num)
#Applying PCA to the scaled data
from sklearn.decomposition import PCA
#Setting the variance to 0.95
pca = PCA(n_components = 0.95)
pca.fit(scaled_data)
#Applying to our scaled dataset
scaled_data_pca = pca.transform(scaled_data)
#Check the shape of the original dataset and the new dataset
print("The dimensions of the original dataset are: ", scaled_data.shape)
print("The dimensions of the dataset after performing PCA is: ", scaled_data_pca.shape)
# -
# As we can see from the above output, 3 principal components are required to capture 95% of the variance in the dataset. This means that by choosing 2 principal directions previously, we were capturing < 95% of the variance in the dataset. Despite capturing < 95% of the variance, we were able to visualize the fact that the pizzas produced by different companies have different nutritional contents.
# ## An application to real-world data
#
# The first step is to import the data as shown below. It is going to take some time since it is a big dataset, hence hang tight. The dataset contains images of 70000 digits (0-9) where each image has 784 features.
# +
#Importing the dataset
from sklearn.datasets import fetch_openml
mnist_data = fetch_openml('mnist_784', version = 1)
# Choosing the independent (X) and dependent variables (y)
X,y = mnist_data["data"], mnist_data["target"]
# -
# Now that we have the dataset imported, we will move on to visualize the image of a digit to get familiar with the dataset. For visualization, we will use the `matplotlib` library. We will visualize the 50000th digit image. Feel free to check out other digit images of your choice – make sure to use an index between 0 and 69999. We will set colormap to "binary" to output a grayscale image.
# +
#Plotting one of the digits
import matplotlib.pyplot as plt
plt.figure(1)
#Plotting the 50000th digit
digit = X[50000]
#Reshaping the 784 features into a 28x28 matrix
digit_image = digit.reshape(28,28)
plt.imshow(digit_image, cmap='binary')
plt.show()
# -
# Next, we will apply PCA to this dataset to reduce its dimension from $28*28=784$ to a lower number. We will plot the proportion of the variation that is reflected by PCA-reduced dimensional data of different dimensions.
# +
#Scaling the data
from sklearn.preprocessing import StandardScaler
scaled_mnist_data = StandardScaler().fit_transform(X)
print(scaled_mnist_data.shape)
#Applying PCA to ur dataset
from sklearn.decomposition import PCA
pca = PCA(n_components=784)
mnist_data_pca = pca.fit_transform(scaled_mnist_data)
#Calculating cumulative variance captured by PCs
import numpy as np
variance_percentage = pca.explained_variance_/np.sum(pca.explained_variance_)
#Calculating cumulative variance
cumulative_variance = np.cumsum(variance_percentage)
#Plotting cumalative variance
import matplotlib.pyplot as plt
plt.figure(2)
plt.plot(cumulative_variance)
plt.xlabel('Number of principal components')
plt.ylabel('Cumulative variance explained by PCs')
plt.grid()
plt.show()
| Chapter 12 - Principal Components Analysis with scikit-learn/Chapter12.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="../images/QISKit-c.gif" alt="Note: In order for images to show up in this jupyter notebook you need to select File => Trusted Notebook" width="250 px" align="left">
# ## _*Winning The Game of Magic Square with Quantum Pseudo-Telepathy *_
#
# The latest version of this notebook is available on https://github.com/QISKit/qiskit-tutorial.
# ***
# ### Contributors
# <NAME>
# ## Introduction
#
# We have seen that quantum entanglement enables phenomena that we often see in science fiction, such as, (quantum) teleportation. Now, we will see that it can give rise to a kind of telepathy between two separated parties. Well, strictly speaking, quantum entanglement does not allow communication. However, it can be used for [Quantum Pseudo-Telepathy](https://en.wikipedia.org/wiki/Quantum_pseudo-telepathy), that is, parties sharing entangled states can be seen as if having some kind of communication to outside observers.
#
# Here, we consider [the Magic Square Game](https://en.wikipedia.org/wiki/Quantum_pseudo-telepathy), which is also known as *The Mermin-Peres Magic Square Game*. The magic square is a $3\times 3$ matrix whose entries are either $0$ or $1$ such that the sum of each row is even, and the sum of each column is odd. Notice that such *magic* square is impossible: because there are odd number of entries, the sum of rows implies that the sum of $1$s must be even, but the sum of columns implies that it must be odd. A contradiction.
#
# The magic square game is played by a referee against two distant parties, say, Alice and Bob. In the game, the referee sends an integer $a \in \{1,2,3\}$ to Alice who must answer with the $a$-th row of the magic square, and an integer $b \in \{1,2,3\}$ to Bob, who must return the $b$-th column of the magic square. Alice and Bob win the game if the sum of entries of Alice's answer is even, the sum of Bob's answer is odd, and their intersecting answer is the same. Otherwise, the referee wins. Prior to the start of the game, Alice and Bob can meet to discuss their strategy and/or share random bits and entangled states, but they are not allowed to communicate during the game.
#
# For example, a simple strategy for Alice and Bob is to answer to the referee according to the following $3x3$ Boolean matrix:
#
# $$
# \begin{pmatrix}
# 1 & 1 & 0\\
# 0 & 1 & 1\\
# 0 & 1 & ?
# \end{pmatrix}.
# $$
#
# That is, for $a = 1$ and $b = 2$, Alice's answer is $110$, while Bob's is $111$ and they win. However, they lose when the referee sends them $a = 3$ and $b=3$, because Alice's and Bob's answers do not satisfy the requirement. It can be shown that in the classical setting the winning probability of Alice and Bob is at most $8/9$ (in the above example, there are eight out of nine combinations of $a$ and $b$ that result in Alice and Bob winning the game).
#
#
# ## Quantum Winning Strategy
#
# However, with shared quantum states Alice and Bob can always win the game regardless of the values of $a$ and $b$. We show the winning strategy following [Brassard, Broadbent, and Tapp, 2004](https://arxiv.org/abs/quant-ph/0407221).
#
# ### Preparing the environment
# First, as usual we prepare the environment.
# +
# Checking the version of PYTHON; we only support > 3.5
import sys
if sys.version_info < (3,5):
raise Exception('Please use Python version 3.5 or greater.')
# useful additional packages
import matplotlib.pyplot as plt
# %matplotlib inline
import numpy as np
# useful math functions
from math import pi, cos, acos, sqrt
import random
# importing the QISKit
from qiskit import QuantumProgram
import Qconfig
# import basic plot tools
from qiskit.tools.visualization import plot_histogram
# -
# ### Sharing entangled quantum states
#
# Prior to the start of the game, Alice and Bob share the following quantum state. The first two qubits go to Alice, and the rest to Bob.
#
# $$
# |\psi\rangle = \frac{1}{2}|0011\rangle - \frac{1}{2}|0110\rangle - \frac{1}{2}|1001\rangle + \frac{1}{2}|1100\rangle
# $$
#
# To generate such quantum state, we first prepare $4$ qubits and the corresponding $4$ classical bits to record the measurement later. Below is a quantum circuit to create the above entangled state.
# +
Q_program = QuantumProgram()
Q_program.set_api(Qconfig.APItoken, Qconfig.config["url"]) # set the APIToken and API url
N = 4
# Creating registers
qr = Q_program.create_quantum_register("qr", N)
# for recording the measurement on qr
cr = Q_program.create_classical_register("cr", N)
circuitName = "sharedEntangled"
sharedEntangled = Q_program.create_circuit(circuitName, [qr], [cr])
#Create uniform superposition of all strings of length 2
for i in range(2):
sharedEntangled.h(qr[i])
#The amplitude is minus if there are odd number of 1s
for i in range(2):
sharedEntangled.z(qr[i])
#Copy the content of the fist two qubits to the last two qubits
for i in range(2):
sharedEntangled.cx(qr[i], qr[i+2])
#Flip the last two qubits
for i in range(2,4):
sharedEntangled.x(qr[i])
# -
# ### Alice's and Bob's operations
#
# Receiving $a \in \{1,2,3\}$, Alice applies the unitary matrix $A_a$ on her qubits, where $A_a$ is one of the followings:
#
# $$
# A_1 = \frac{1}{\sqrt{2}}
# \begin{pmatrix}
# i & 0 & 0 & 1\\
# 0 & -i& 1 & 0\\
# 0 & i & 1 & 0\\
# 1 & 0 & 0 & i
# \end{pmatrix},~
# A_2 = \frac{1}{2}
# \begin{pmatrix}
# i & 1 & 1 & i\\
# -i& 1 &-1 & i\\
# i & 1 &-1 &-i\\
# -i& 1 & 1 &-i
# \end{pmatrix},~
# A_3 = \frac{1}{2}
# \begin{pmatrix}
# -1&-1&-1& 1\\
# 1 & 1&-1& 1\\
# 1 &-1& 1& 1\\
# 1 &-1&-1&-1
# \end{pmatrix}
# $$
#
# Meanwhile, receiving $b \in \{1,2,3\}$, Bob applies the unitary matrix $B_b$ on his qubits, where $B_b$ is one of the followings:
#
# $$
# B_1 = \frac{1}{2}
# \begin{pmatrix}
# i&-i& 1& 1\\
# -i&-i& 1&-1\\
# 1& 1&-i& i\\
# -i& i& 1& 1
# \end{pmatrix},~
# B_2 = \frac{1}{2}
# \begin{pmatrix}
# -1& i& 1& i\\
# 1& i& 1&-i\\
# 1&-i& 1& i\\
# -1&-i& 1&-i
# \end{pmatrix},~
# B_3 = \frac{1}{\sqrt{2}}
# \begin{pmatrix}
# 1& 0 & 0 & 1\\
# -1& 0 & 0 & 1\\
# 0 & 1 & 1 & 0\\
# 0 & 1 &-1 & 0
# \end{pmatrix}
# $$
#
# After applying their unitary operators, Alice and Bob independently measure their qubits in the computational basis and use the outcome as the answer to their first two bits, while their third bits can be inferred from the parity with each of their first two bits: even for Alice and odd for Bob.
#
# Below are the circuits of Alice's and Bob's operations.
# +
#we first define controlled-u gates required to assign phases
from math import pi
def ch(qProg, a, b):
""" Controlled-Hadamard gate """
qProg.h(b)
qProg.sdg(b)
qProg.cx(a, b)
qProg.h(b)
qProg.t(b)
qProg.cx(a, b)
qProg.t(b)
qProg.h(b)
qProg.s(b)
qProg.x(b)
qProg.s(a)
return qProg
def cu1pi2(qProg, c, t):
""" Controlled-u1(phi/2) gate """
qProg.u1(pi/4.0, c)
qProg.cx(c, t)
qProg.u1(-pi/4.0, t)
qProg.cx(c, t)
qProg.u1(pi/4.0, t)
return qProg
def cu3pi2(qProg, c, t):
""" Controlled-u3(pi/2, -pi/2, pi/2) gate """
qProg.u1(pi/2.0, t)
qProg.cx(c, t)
qProg.u3(-pi/4.0, 0, 0, t)
qProg.cx(c, t)
qProg.u3(pi/4.0, -pi/2.0, 0, t)
return qProg
# -
# The last two gates will be used to assign amplitudes with $i$ phase and realize the following unitaries:
#
# $$
# U_\mbox{cu1pi2} =
# \begin{pmatrix}
# 1 & 0 & 0 & 0\\
# 0 & 1 & 0 & 0\\
# 0 & 0 & 1 & 0\\
# 0 & 0 & 0 & i
# \end{pmatrix},~
# U_\mbox{cu3pi2} = \frac{1}{\sqrt{2}}
# \begin{pmatrix}
# \sqrt{2} & 0 & 0 & 0\\
# 0 & \sqrt{2} & 0 & 0\\
# 0 & 0 & 1 & -i\\
# 0 & 0 & -i & 1
# \end{pmatrix}
# $$
#
# Next, we define circuits used by Alice and Bob for each of their inputs.
# +
# dictionary for Alice's operations/circuits
aliceCircuits = {}
# Quantum circuits for Alice when receiving idx in 1, 2, 3
for idx in range(1, 4):
circuitName = "Alice"+str(idx)
aliceCircuits[circuitName] = Q_program.create_circuit(circuitName, [qr], [cr])
theCircuit = aliceCircuits[circuitName]
if idx == 1:
#the circuit of A_1
theCircuit.x(qr[1])
theCircuit.cx(qr[1], qr[0])
theCircuit = cu1pi2(theCircuit, qr[1], qr[0])
theCircuit.x(qr[0])
theCircuit.x(qr[1])
theCircuit = cu1pi2(theCircuit, qr[0], qr[1])
theCircuit.x(qr[0])
theCircuit = cu1pi2(theCircuit, qr[0], qr[1])
theCircuit = cu3pi2(theCircuit, qr[0], qr[1])
theCircuit.x(qr[0])
theCircuit = ch(theCircuit, qr[0], qr[1])
theCircuit.x(qr[0])
theCircuit.x(qr[1])
theCircuit.cx(qr[1], qr[0])
theCircuit.x(qr[1])
elif idx == 2:
theCircuit.x(qr[0])
theCircuit.x(qr[1])
theCircuit = cu1pi2(theCircuit, qr[0], qr[1])
theCircuit.x(qr[0])
theCircuit.x(qr[1])
theCircuit = cu1pi2(theCircuit, qr[0], qr[1])
theCircuit.x(qr[0])
theCircuit.h(qr[0])
theCircuit.h(qr[1])
elif idx == 3:
theCircuit.cz(qr[0], qr[1])
theCircuit.swap(qr[0], qr[1])
theCircuit.h(qr[0])
theCircuit.h(qr[1])
theCircuit.x(qr[0])
theCircuit.x(qr[1])
theCircuit.cz(qr[0], qr[1])
theCircuit.x(qr[0])
theCircuit.x(qr[1])
#measure the first two qubits in the computational basis
theCircuit.measure(qr[0], cr[0])
theCircuit.measure(qr[1], cr[1])
# dictionary for Bob's operations/circuits
bobCircuits = {}
# Quantum circuits for Bob when receiving idx in 1, 2, 3
for idx in range(1,4):
circuitName = "Bob"+str(idx)
bobCircuits[circuitName] = Q_program.create_circuit(circuitName, [qr], [cr])
theCircuit = bobCircuits[circuitName]
if idx == 1:
theCircuit.x(qr[2])
theCircuit.x(qr[3])
theCircuit.cz(qr[2], qr[3])
theCircuit.x(qr[3])
theCircuit.u1(pi/2.0, qr[2])
theCircuit.x(qr[2])
theCircuit.z(qr[2])
theCircuit.cx(qr[2], qr[3])
theCircuit.cx(qr[3], qr[2])
theCircuit.h(qr[2])
theCircuit.h(qr[3])
theCircuit.x(qr[3])
theCircuit = cu1pi2(theCircuit, qr[2], qr[3])
theCircuit.x(qr[2])
theCircuit.cz(qr[2], qr[3])
theCircuit.x(qr[2])
theCircuit.x(qr[3])
elif idx == 2:
theCircuit.x(qr[2])
theCircuit.x(qr[3])
theCircuit.cz(qr[2], qr[3])
theCircuit.x(qr[3])
theCircuit.u1(pi/2.0, qr[3])
theCircuit.cx(qr[2], qr[3])
theCircuit.h(qr[2])
theCircuit.h(qr[3])
elif idx == 3:
theCircuit.cx(qr[3], qr[2])
theCircuit.x(qr[3])
theCircuit.h(qr[3])
#measure the third and fourth qubits in the computational basis
theCircuit.measure(qr[2], cr[2])
theCircuit.measure(qr[3], cr[3])
# -
# ### A quantum program for one round of the game
#
# Prior to the start of the game, Alice and Bob share the entangled quantum states as described before. Notice that this is performed before their receiving inputs. After sharing entanglement, they are not allowed to communicate. Next, an integer $a$ is given to Alice, and $b$ to Bob. Alice and Bob then independently perform operations with one the circuits defined previously based on their inputs. They generate their answers (three bits each) based on their measurement outcomes so that the parity of Alice's answer is even, and Bob's answer is odd. Here is a program for one round of the game using the circuits previously defined.
# +
a, b = random.randint(1,3), random.randint(1,3) #generate random integers
print("The values of a and b are, resp.,", a,b)
aliceCircuit = aliceCircuits["Alice"+str(a)]
bobCircuit = bobCircuits["Bob"+str(b)]
circuitName = "Alice"+str(a)+"Bob"+str(b)
Q_program.add_circuit(circuitName, sharedEntangled+aliceCircuit+bobCircuit)
backend = "local_qasm_simulator"
##backend = "ibmqx2"
shots = 1 # We perform a one-shot experiment
results = Q_program.execute([circuitName], backend=backend, shots=shots)
answer = results.get_counts(circuitName)
print(answer)
for key in answer.keys():
aliceAnswer = [int(key[-1]), int(key[-2])]
bobAnswer = [int(key[-3]), int(key[-4])]
if sum(aliceAnswer) % 2 == 0:#the sume of Alice answer must be even
aliceAnswer.append(0)
else:
aliceAnswer.append(1)
if sum(bobAnswer) % 2 == 1:#the sum of Bob answer must be odd
bobAnswer.append(0)
else:
bobAnswer.append(1)
break
print("Alice answer for a = ", a, "is", aliceAnswer)
print("Bob answer for b = ", b, "is", bobAnswer)
if(aliceAnswer[b-1] != bobAnswer[a-1]): #check if the intersection of their answers is the same
print("Alice and Bob lost")
else:
print("Alice and Bob won")
# -
# ### Checking Alice's and Bob's answers for all combinations of their inputs
#
# Finally, we can try every combination of $a$ and $b$ to see that Alice and Bob can always win surely. One can also try to run the code below with `ibmqx2` backend and check that the winning probability can be higher than $8/9$ on real devices (unfortunately, perfect probability cannot be achieved due to noise and gate errors).
# +
backend = "local_qasm_simulator"
#backend = "ibmqx2"
shots = 10 # We perform 10 shots of experiments for each round
nWins = 0
nLost = 0
for a in range(1,4):
for b in range(1,4):
print("Asking Alice and Bob with a and b are, resp.,", a,b)
rWins = 0
rLost = 0
aliceCircuit = aliceCircuits["Alice"+str(a)]
bobCircuit = bobCircuits["Bob"+str(b)]
circuitName = "Alice"+str(a)+"Bob"+str(b)
Q_program.add_circuit(circuitName, sharedEntangled+aliceCircuit+bobCircuit)
if backend == "ibmqx2":
ibmqx2_backend = Q_program.get_backend_configuration('ibmqx2')
ibmqx2_coupling = ibmqx2_backend['coupling_map']
results = Q_program.execute([circuitName], backend=backend, shots=shots, coupling_map=ibmqx2_coupling, max_credits=3, wait=10, timeout=240)
else:
results = Q_program.execute([circuitName], backend=backend, shots=shots)
answer = results.get_counts(circuitName)
for key in answer.keys():
kfreq = answer[key] #frequencies of keys obtained from measurements
aliceAnswer = [int(key[-1]), int(key[-2])]
bobAnswer = [int(key[-3]), int(key[-4])]
if sum(aliceAnswer) % 2 == 0:
aliceAnswer.append(0)
else:
aliceAnswer.append(1)
if sum(bobAnswer) % 2 == 1:
bobAnswer.append(0)
else:
bobAnswer.append(1)
#print("Alice answer for a = ", a, "is", aliceAnswer)
#print("Bob answer for b = ", b, "is", bobAnswer)
if(aliceAnswer[b-1] != bobAnswer[a-1]):
#print(a, b, "Alice and Bob lost")
nLost += kfreq
rLost += kfreq
else:
#print(a, b, "Alice and Bob won")
nWins += kfreq
rWins += kfreq
print("\t#wins = ", rWins, "out of ", shots, "shots")
print("Number of Games = ", nWins+nLost)
print("Number of Wins = ", nWins)
print("Winning probabilities = ", (nWins*100.0)/(nWins+nLost))
# -
# ## About Quantum Pseudo-Telepathy for the Magic Square Game
# The winning strategy described in this note is from [Brassard et al. 2004](https://arxiv.org/abs/quant-ph/0407221), where there listed many other interesting games that can be accomplished with shared entanglement. The Magic Square game was first proposed by [Aravind, 2002](https://arxiv.org/abs/quant-ph/0206070) based on the work of [Mermin, 1990](http://blog.umd.edu/phil858/files/2015/10/Mermin1990-2jvtdbh.pdf) and [Peres, 1990](http://blog.umd.edu/phil858/files/2015/10/Peres1990-2g8uzi6.pdf).
#
# [Gawron et al., 2008](https://arxiv.org/pdf/0801.4848v1.pdf) showed that the winning probabilities of Magic Square Games is related to the noise of the circuits. [Ozaydin, 2016](https://arxiv.org/abs/1609.03881) provided theoretical analysis of the winning probabilities of the thermal entangled state of the spin system for the Magic Square game: the higher the temperature, the lower the winning probability. Interestingly, [Pawela et al. 2013](http://journals.plos.org/plosone/article?id=10.1371/journal.pone.0064694) showed that it is possible to achieve higher winning probability under noisy circuits by employing Semidefinite Programming for noise mitigation.
# %run "../version.ipynb"
| 5_games/quantum_magic_square.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Web scrapping using python
#
# #### References
# 1. [Practical Introduction to Web Scraping in Python](https://realpython.com/python-web-scraping-practical-introduction/)
# 2. [Web Scraping using Python](https://www.datacamp.com/community/tutorials/web-scraping-using-python)
# +
# $ python3 -m venv venv
# $ . ./venv/bin/activate
# +
# #!pip install --upgrade pip
# -
#Better
# #!pip install requests BeautifulSoup4 fire
# +
from requests import get
from requests.exceptions import RequestException
from contextlib import closing
from bs4 import BeautifulSoup
import pandas as pd
import os, sys
import fire
# +
# #%%writefile ../pyscrap_url.py
def simple_get(url):
"""
Attempts to get the content at `url` by making an HTTP GET request.
If the content-type of response is some kind of HTML/XML, return the
text content, otherwise return None.
"""
try:
with closing(get(url, stream=True)) as resp:
if is_good_response(resp):
return resp.content #.encode(BeautifulSoup.original_encoding)
else:
return None
except RequestException as e:
log_error('Error during requests to {0} : {1}'.format(url, str(e)))
return None
def is_good_response(resp):
"""
Returns True if the response seems to be HTML, False otherwise.
"""
content_type = resp.headers['Content-Type'].lower()
return (resp.status_code == 200
and content_type is not None
and content_type.find('html') > -1)
def log_error(e):
"""
It is always a good idea to log errors.
This function just prints them, but you can
make it do anything.
"""
print(e)
def get_elements(url, tag='',search={}, fname=None):
"""
Downloads a page specified by the url parameter
and returns a list of strings, one per tag element
"""
if isinstance(url,str):
response = simple_get(url)
else:
#if already it is a loaded html page
response = url
if response is not None:
html = BeautifulSoup(response, 'html.parser')
res = []
if tag:
for li in html.select(tag):
for name in li.text.split('\n'):
if len(name) > 0:
res.append(name.strip())
if search:
soup = html
r = ''
if 'find' in search.keys():
print('findaing',search['find'])
soup = soup.find(**search['find'])
r = soup
if 'find_all' in search.keys():
print('findaing all of',search['find_all'])
r = soup.find_all(**search['find_all'])
if r:
for x in list(r):
if len(x) > 0:
res.extend(x)
return res
# Raise an exception if we failed to get any data from the url
raise Exception('Error retrieving contents at {}'.format(url))
if get_ipython().__class__.__name__ == '__main__':
fire(get_tag_elements)
# -
res = get_elements('https://africafreak.com/100-most-influential-twitter-users-in-africa', tag='h2')
# +
res
#print(res)
for i in res[100:]:
res.remove(i)
#print(res)
names_infl = []
handle_infl = []
for r in res:
split_data = r.split('.',maxsplit=1)[1].rsplit('(',maxsplit=1)
name = split_data[0].split(',')[0].strip()
handle = split_data[1].split(')',maxsplit=1)[0]
names_infl.append(name)
handle_infl.append(handle)
#print(f'{name}:{handle}'
#print(handle_infl)
#df_influencer_name = pd.DataFrame(names_infl)
#df_influencer_handle = pd.DataFrame(handle_infl)
#df_influencer_name.to_csv('C:/Users/HP/Desktop/CV, P.Statement and others/10 Academy/Topinfluencer_handle.csv', index = False)
#print(df_influencer_handle)
# -
url= 'https://www.atlanticcouncil.org/blogs/africasource/african-leaders-respond-to-coronavirus-on-twitter/#east-africa'
response = simple_get(url)
# +
url= 'https://www.atlanticcouncil.org/blogs/africasource/african-leaders-respond-to-coronavirus-on-twitter/#east-africa'
response = get(url).content
re_gov = get_elements(response, tag='blockquote')
names = []
handles = []
for r in re_gov:
split_data = r.split('— ',maxsplit=1)[1].rsplit('(',maxsplit=1)
#print(split_data)
name = split_data[0].split(',')[0].strip()
handle = split_data[1].rsplit(')',maxsplit=1)[0]
names.append(name)
handles.append(handle)
nam_handle = f'{name}:{handle}'
print(handles)
#print(df_gvrn)
#res[:2]
# -
df_gvrn = pd.DataFrame(handles, columns=["Twitter Handles"])
df_gvrn.to_csv (r'C:\Users\HP\Desktop\CV, P.Statement and others\10 Academy\gov_influencers_handles.csv', index = False, header=True)
#print(df_gvrn)
# +
import sys
import os
import json
import pandas as pd
import matplotlib.pyplot as plt
import re
import string
import matplotlib.dates as mdates
import seaborn as sns
#sns.set()
# to view all columns
#pd.set_option("display.max.columns", None)
# -
# #!pip install tweepy
import tweepy
from tweepy.streaming import StreamListener
from tweepy import OAuthHandler
from tweepy import Stream
consumer_key = "re8a7dLArl8y2Tic312dbRWxH"
consumer_secret = "<KEY>"
access_token = "<KEY>"
access_token_secret = "<KEY>"
auth = OAuthHandler(consumer_key, consumer_secret)
auth.set_access_token(access_token, access_token_secret)
api = tweepy.API(auth)
#getting tweets for 100 influencers
tweets = []
tweetCount=1
for i in handle_infl:
try:
results=api.user_timeline(id=i, count=tweetCount)
except tweepy.TweepError as e:
continue
for tweet in results:
tweets.append(tweet.text)
#print(tweets)
#to convert tweets to csv
influencer_tweets = pd.DataFrame(tweets, columns=["Influencer Tweets"] )
print(influencer_tweets)
# +
# getting followers for 100 influencers
# Calling the get_user function with our parameters
followers = []
for i in handle_infl:
try:
results = api.get_user(id=i)
except tweepy.TweepError as e:
continue
followers.append(results.followers_count)
#print(results.followers_count)
#print(followers)
# -
# followers dataframe
df_followers = pd.DataFrame(followers, columns = ["Number of followers"])
df_followers.to_csv('C:/Users/HP/Desktop/CV, P.Statement and others/10 Academy/infl_followers.csv', index = False)
print(df_followers.head())
# +
# getting no of likes for 100 influencers
likes = []
for i in handle_infl:
try:
results = api.get_user(id=i)
except tweepy.TweepError as e:
continue
likes.append(results.favourites_count)
#print(results.followers_count)
#print(likes)
df_likes = pd.DataFrame(likes, columns=["No of likes"])
df_likes.to_csv (r'C:\Users\HP\Desktop\CV, P.Statement and others\10 Academy\top_influencers_likes.csv', index = False, header=True)
#print(df_likes)
# +
# getting no of following for 100 influencers
following = []
for i in handle_infl:
try:
results = api.get_user(id=i)
except tweepy.TweepError as e:
continue
following.append(results.friends_count)
print(following)
# +
# followers dataframe
d_following = pd.DataFrame(following, columns = ["Number of following"])
d_following.to_csv('C:/Users/HP/Desktop/CV, P.Statement and others/10 Academy/following.csv', index = False)
print(d_following.head())
# +
#getting retweets
no_of_retweets = []
for x in handle_infl:
try:
tweets = tweepy.Cursor(api.user_timeline, id=x, lang="en").items()
for tweet in tweets:
no_of_retweets.append(tweet.retweet_count)
#print(tweet.retweet_count)
except tweepy.TweepError as e:
continue
print(no_of_retweets)
#retweets dataframe
df_retweets = pd.DataFrame(no_of_retweets, columns=["No of retweets"])
df_retweets.to_csv (r'C:\Users\HP\Desktop\CV, P.Statement and others\10 Academy\finalJ_retweets.csv', index = False, header=True)
print(df_retweets)
# +
#getting no of tweets shared
no_tweets_shared = []
for i in handle_infl:
try:
results = api.get_user(id=i)
except tweepy.TweepError as e:
continue
no_tweets_shared.append(results.statuses_count)
print(no_tweets_shared)
# -
# ### Popularity reach
# ### Reach Score
# ### Relevance Score
#popularity reach
#Reach Score = followers - following
reach = pd.concat([df_followers,d_following], axis=1)
print(reach.head())
reach['reach_score']= reach["Number of followers"] - reach["Number of following"]
print(reach.head())
# +
retweets=[]
for i in id_num:
try:
results = api.retweets(id=i,count=10)
except tweepy.TweepError as e:
continue
#print(results)
#retweets.append(results.text)
#print(retweets)
# +
#hash_tags
hash_tags = []
for i in handle_infl:
try:
results = api.get_user(id=i)
except tweepy.TweepError as e:
continue
if hasattr(status, "entities"):
entities = status.entities
# get hashtags
if "hashtags" in entities:
for ent in entities["hashtags"]:
if ent is not None:
if "text" in ent:
hashtag = ent["text"]
if hashtag is not None:
hashtags.append(hashtag)
for status in Cursor(auth_api.user_timeline, id=handle).items():
tweet_count+= 1
if hasattr(status, "entities"):
entities = status.entities
# get hashtags
if "hashtags" in entities:
for ent in entities["hashtags"]:
if ent is not None:
if "text" in ent:
hashtag = ent["text"]
if hashtag is not None:
hashtags.append(hashtag)
# +
#getting follower for gov
followers = []
for i in name:
results = api.get_user(id=i)
followers.append(results.followers_count)
#print(results.followers_count)
#print(followers)
# followers dataframe
#df_followers = pd.DataFrame(followers, columns = ["Number of followers"])
#df_followers.to_csv('C:/Users/HP/Desktop/CV, P.Statement and others/10 Academy/gov_followers.csv', index = False)
#print(df_followers.head())
# +
# The Twitter users who we want to get tweets from
name = handles
# Calling the get_user function with our parameters
#print(tweet.retweet_count)
#print(tweet.favorite_count)
#no of likes
likes = []
for i in name:
results = api.get_user(id=i)
likes.append(results.favourites_count)
#print(results.followers_count)
#print(likes)
df_likes = pd.DataFrame(likes, columns=["Twitter Handles"])
df_likes.to_csv (r'C:\Users\HP\Desktop\CV, P.Statement and others\10 Academy\gov_influencers_likes.csv', index = False, header=True)
#print(df_likes)
# +
#no of retweets
name = handles
retweets = []
for i in name:
results = api.user_timeline(id=i)
retweets.
#print(results)
if hasattr(results, 'retweet_count'):
retweets = results.retweet_count
#if retweets is not None:
# retweet_count.append(retweets)
print(retweets)
# +
name = handles
following = []
for i in name:
results = api.get_user(id=i)
following.append(results.friends_count)
print(following)
df_ffl = pd.DataFrame(following, columns=["Number of following"])
df_ffl.to_csv (r'C:\Users\HP\Desktop\CV, P.Statement and others\10 Academy\gov_influencers_following.csv', index = False, header=True)
#print(df_ffl)
# -
reach_score = pd.concat([df_gvrn,df_ffl], axis=1)
#print(reach_score)
#reach_score.sort_order()
len(api.retweets(197493438))
# +
# Creating the API object while passing in auth information
# The Twitter user who we want to get tweets from
name = handles
# Calling the get_user function with our parameters
followers = []
for i in name:
results = api.get_user(id=i)
# print(results)
#followers.append(results.followers_count)
#print(results.followers_count)
#print(followers)
# followers dataframe
#df_followers = pd.DataFrame(followers, columns = ["Number of followers"])
#df_followers.to_csv('C:/Users/HP/Desktop/CV, P.Statement and others/10 Academy/gov_followers.csv', index = False)
#print(df_followers.head())
# -
# ## Web scrapping using bash script
# If the web site has a quite simple HTML, you can easily use curl to perform the request and then extract the needed values using bash commands grep, cut , sed, ..
#
# This tutorial is adapted from [this](https://medium.com/@LiliSousa/web-scraping-with-bash-690e4ee7f98d) medium article
# + language="bash"
#
# # curl the page and save content to tmp_file
# #url = "https://www.atlanticcouncil.org/blogs/africasource/african-leaders-respond-to-coronavirus-on-twitter/#east-africa"
# #curl -X GET $url -o tmp_file
#
#
# #!/bin/bash
#
# # write headers to CSV file
# echo "Name, twitter_id" >> extractData.csv
# n="1"
# while [ $n -lt 2 ]
# do
#
# #get title
# title=$(cat tmp_file | grep "class=\"twitter-tweet\"" | cut -d ';' -f1 )
# echo $title
# #get author
# #twitter_id=$(cat tmp_file |grep -A1 "class=\"css-901oao css-16my406 r-1qd0xha r-ad9z0x r-bcqeeo r-qvutc0\"" | tail -1)
#
# #echo "$title, $twitter_id" >> extractData.csv
# #echo "$title, $twitter_id"
#
# n=$[$n+1]
#
# done
# -
| week1/notebook/scrapping_starter updated.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Wisawasi/basics/blob/master/24_example_deep_learning.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Yp6lb-pIsZj8"
# <img height="45px" src="https://avatars3.githubusercontent.com/u/20337240?s=460&v=4" align="left" hspace="10px" vspace="0px">
#
# <h1>รวมตัวอย่าง Machine learning และ Deep learning</h1>
#
# <hr>
#
# <a href='https://www.facebook.com/programmerthai'><font color='green'>สรุปและเรียบเรียงโดย โปรแกรมเมอร์ไทย thai programmer</font></a>
#
#
# + [markdown] id="zv5f6VJPSkOD"
# # วิธีใช้งาน colab
#
# Colab พัฒนาโดย google เป็นหน้าเว็บเอาไว้ใช้เขียนโค้ด python ได้ผ่านทางหน้าเว็บ และเชื่อมต่อกับ google drive ของเราก็ได้ด้วย ซึ่ง colab หน้านี้**ได้รวบรวมเนื้อหา Python ของม.ต้น** เอาไว้ แต่ก่อนอื่นอยากให้ดูวิธีการใช้งาน colab เสียก่อน
#
# <font color='red'>** ต้องมี Gmail และ google drive</font>
#
# <hr/>
#
# <h3>วิธี 1) กด "Open in playgrouhd"</h3>
# <div>
# <img height="200px" src="https://www.patanasongsivilai.com/blog/wp-content/uploads/2020/05/python_matayom_4.png" align="center" hspace="10px" vspace="0px">
#
#
# <hr/>
#
# <h3>วิธี 2) กด Save a copy in Drive ... เพื่อบันทึกไว้ google drive เครื่องเรา</h3>
#
# <img height="300px" src="https://www.patanasongsivilai.com/blog/wp-content/uploads/2019/11/python_matayom_1.png" align="center" hspace="10px" vspace="0px">
#
# <br/>
#
# <img height="200px" src="https://www.patanasongsivilai.com/blog/wp-content/uploads/2019/11/python_matayom_2.png" align="center" hspace="10px" vspace="0px">
#
# <hr/>
#
# <h3>จากนั้นถึงจะสามารถรันได้</h3>
# <img height="200px" src="https://www.patanasongsivilai.com/blog/wp-content/uploads/2019/11/python_matayom_3.png" align="center" hspace="10px" vspace="0px">
# + [markdown] id="zj-YJroM3594"
# # import โมดูลสำคัญๆ
# + id="4SAMRUNp6FJX" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="6fbea839-9ef1-4209-e860-389e10c2b24b"
import keras
from keras.datasets import mnist
from keras.layers import Dense, Dropout
from keras.models import Sequential
from matplotlib import pyplot as plt
from keras.layers.core import Dense, Activation, Dropout, Flatten
from keras.layers.recurrent import SimpleRNN, LSTM, GRU
from keras.models import Sequential
from keras.regularizers import l2
from keras import backend as K
from keras.layers import Conv2D, Conv1D, MaxPooling2D, MaxPooling1D
# + [markdown] id="1cEaN4FUX6g3"
# # Datasets
#
# ฟังก์ชั่นดาวนโหลด datasets ได้แก่ cifar10, cifar100, imdb, reuters, mnist, fashion_mnist และ boston_housing
# + id="XzZmRZn-X5lO"
def get_datasets(param='mnist'):
x_train = None; y_train = None; x_test = None; y_test = None
if param == 'cifar10':
from keras.datasets import cifar10
(x_train, y_train), (x_test, y_test) = cifar10.load_data()
elif param == 'cifar100':
from keras.datasets import cifar100
(x_train, y_train), (x_test, y_test) = cifar100.load_data(label_mode='fine')
elif param == 'imdb':
from keras.datasets import imdb
(x_train, y_train), (x_test, y_test) = imdb.load_data(path="imdb.npz",
num_words=None,
skip_top=0,
maxlen=None,
seed=113,
start_char=1,
oov_char=2,
index_from=3)
elif param == 'reuters':
from keras.datasets import reuters
(x_train, y_train), (x_test, y_test) = reuters.load_data(path="reuters.npz",
num_words=None,
skip_top=0,
maxlen=None,
test_split=0.2,
seed=113,
start_char=1,
oov_char=2,
index_from=3)
# Optional
word_index = reuters.get_word_index(path="reuters_word_index.json")
elif param == 'mnist':
from keras.datasets import mnist
(x_train, y_train), (x_test, y_test) = mnist.load_data()
elif param == 'fashion_mnist':
from keras.datasets import fashion_mnist
(x_train, y_train), (x_test, y_test) = fashion_mnist.load_data()
elif param == 'boston_housing':
from keras.datasets import boston_housing
(x_train, y_train), (x_test, y_test) = boston_housing.load_data()
return x_train, y_train, x_test, y_test
# + id="LkafliG0PHld" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="efa3c76a-1aa6-4f50-9a62-eb4b16eb2553"
"""
from sklearn.model_selection import train_test_split
def get_datasets():
digits = datasets.load_digits()
x = digits.data
y = digits.target
x_train, x_test, y_train, y_test = train_test_split(x, y,
test_size=0.33,
random_state=42)
return x_train, x_test, y_train, y_test
"""
# + [markdown] id="AR_fImt_HTRq"
# # Model
# + [markdown] id="-3gmT7ZTHGNW"
# ## Model 1: Nearest neighbors
#
# โมเดลแบบที่ 1
#
# ---
# + id="a5EKWIGmHAi6"
def train_nearest_neighbors(Xtrain, Ytrain, Xtest, Yexpected):
count = 0
numSample,_ = Xtest.shape
# compute distance with Manhattan formula
Ypredicted = np.zeros(numSample)
for index in range(0,numSample):
# minus with broadcasting in numpy
ditanceList = np.sum(np.abs(Xtrain - Xtest[index]), axis=1)
# min distance at first of list and get Index
minDistanceIndex = np.argsort(ditanceList)[0]
Ypredicted[index] = Ytrain[minDistanceIndex]
count = count+1
if count % 100 == 0:
print("progress %d %%" % (count/numSample*100))
# Calculate accuracy (True in python is 1, and False is 0
accuracy = np.sum(Yexpected == Ypredicted)/ len(Yexpected) * 100
print("Accuracy: %.4f" % accuracy)
print("Classification report")
print(metrics.classification_report(Yexpected, Ypredicted))
# + [markdown] id="uQ7F-abfGStQ"
# ## Model 2: Support vector
#
# โมเดลแบบที่ 2
#
#
# ---
#
# + id="aUg0Q6DPGHJr"
from sklearn import datasets, svm, metrics
def train_support_vector(Xtrain, Ytrain, Xtest, Yexpected):
# a support vector classifier
classifier = svm.SVC(gamma=0.001)
# learning
classifier.fit(Xtrain, Ytrain)
# predict
Ypredicted = classifier.predict(Xtest)
# Calculate accuracy (True in python is 1, and False is 0
accuracy = np.sum(Yexpected == Ypredicted)/ len(Yexpected) * 100
print("Accuracy: %.4f" % accuracy)
print("Classification report")
print(metrics.classification_report(Yexpected, Ypredicted))
# + [markdown] id="gfgSREBADOno"
# ## Model 3: Logistic regression (1 neural)
# โมเดลแบบที่ 3
#
#
# ---
#
# + id="OW-vkbjxDr9u"
def build_logistic_regression(features):
model = Sequential()
# L2 is weight regularization penalty, also known as weight decay, or Ridge
model.add(Dense(input_dim=features, output_dim=10, W_regularizer=l2(0.20)))
# now model.output_shape == (None, 10)
# note: `None` is the batch dimension.
#
model.add(Activation("softmax"))
# algorithim to train models use RMSprop
# compute loss with function: categorical crossentropy
model.compile(optimizer='rmsprop',
loss='categorical_crossentropy',
metrics=['accuracy'])
return model
# + [markdown] id="cIyGa_k2dZ04"
# ## Model 4: Neural network (Multilayer Perceptron: MLP)
#
# โมเดลแบบที่ 4
#
#
# ---
#
#
# + id="BC-scfWGbNrE"
def build_MLP(features):
model = Sequential()
model.add(Dense(input_dim=features, units=200, activation="relu"))
model.add(Dense(input_dim=features, units=200, activation="relu"))
model.add(Dropout(rate = 1-0.6)) # reduce overfitting
model.add(Dense(units=10, activation="softmax"))
# algorithim to train models use RMSprop
# compute loss with function: categorical crossentropy
model.compile(optimizer='rmsprop',
loss='categorical_crossentropy',
metrics=['accuracy'])
return model
# + [markdown] id="415xnN6NxJ78"
# ## Model 5: Convolutional Neural Networks (CNN) with Conv2D
#
# โมเดลแบบที่ 5
#
#
# ---
#
# + id="NelPeZ6_xnLw"
import math
def reshapeCNN2D_Input(X):
exampleNum, D = X.shape
W = int(math.sqrt(D))
#assert W == 8 # size of image == 8 x 8
# change shape of image data
if K.image_dim_ordering() == 'th':
# backend is Theano
# Image dimension = chanel x row x column (chanel = 1, if it is RGB: chanel = 3)
XImg = X.reshape(exampleNum, 1, W, W)
else:
# 'tf' backend is Tensorflow
# Image dimension = row x column x chanel (chanel = 1, if it is RGB: chanel = 3)
XImg = X.reshape(exampleNum, W, W, 1)
return XImg
# + id="UqvAkMq_xDg0"
def build_CNN_2D(image_shape):
model = Sequential()
model.add(Conv2D(filters=100, kernel_size=(3, 3), padding='same', input_shape=image_shape))
model.add(Conv2D(filters=100, kernel_size=(3, 3), padding='same'))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2,2)))
model.add(Dropout(0.5))
model.add(Flatten())
model.add(Dense(10))
model.add(Activation('softmax'))
# algorithim to train models use ADAdelta
# compute loss with function: categorical crossentropy
model.compile(optimizer='adadelta',
loss='categorical_crossentropy',
metrics=['accuracy'])
return model
# + [markdown] id="dtAkZVkryBVQ"
# ## Model 6: Convolutional Neural Networks (CNN) with Convolution1D
#
# โมเดลแบบที่ 6
#
#
# ---
#
# + id="E5GIXzoWJO92"
def reshapeCNN1D_Input(X):
exampleNum, D = X.shape
W = int(math.sqrt(D))
#assert W == 8 # size of image == 8 x 8
return X.reshape(exampleNum, W, W)
# + id="_4k0ryzEx-9l"
def build_CNN_1D(image_shape):
model = Sequential()
model.add(Conv1D(filters=100, kernel_size= 3, padding='same', input_shape=image_shape))
model.add(Conv1D(filters=100, kernel_size=3, padding='same'))
model.add(Activation('relu'))
model.add(MaxPooling1D(pool_size=2))
model.add(Dropout(0.5))
model.add(Flatten())
model.add(Dense(10))
model.add(Activation('softmax'))
# algorithim to train models use ADAdelta
# compute loss with function: categorical crossentropy
model.compile(optimizer='adadelta',
loss='categorical_crossentropy',
metrics=['accuracy'])
return model
# + [markdown] id="jCGJjFb-as_I"
# ## Model 7: Recurrent Neural Network (RNN)
#
# โมเดลแบบที่ 7
#
#
# ---
#
#
# + id="lwIrZdP9a3DD"
def getSequenceInput(X):
exampleNum, D = X.shape
W = int(math.sqrt(D))
#assert W == 8 # size of image == 8 x 8
# Dimension = row x colum (without chanel)
XImg = X.reshape(exampleNum, W, W)
return XImg
# + id="1WOKGTCbarC_"
def build_RNN(image_shape):
sequence, features = image_shape
model = Sequential()
model.add(SimpleRNN(input_shape=(sequence, features),
units=100, dropout=0.2, recurrent_dropout=0.2, return_sequences=True))
model.add(SimpleRNN(100, dropout=0.2, recurrent_dropout=0.2, return_sequences=False))
model.add(Dense(10))
model.add(Activation("softmax"))
# algorithim to train models use RMSProp
# compute loss with function: categorical crossentropy
model.compile(optimizer='rmsprop',
loss='categorical_crossentropy',
metrics=['accuracy'])
return model
# + [markdown] id="cVfpU73UYZC5"
# ## Model 8: Long short-term memory (LSTM)
#
# โมเดลแบบที่ 8
#
#
# ---
#
# + id="dou_owuMYXa6"
def build_LSTM(image_shape):
sequence, features = image_shape
model = Sequential()
model.add(LSTM( input_shape=(sequence, features),
units=100, dropout=0.2, recurrent_dropout=0.2, return_sequences=True))
model.add(LSTM(100, dropout=0.2, recurrent_dropout=0.2, return_sequences=False))
model.add(Dense(10))
model.add(Activation("softmax")) #outputs are independent
# algorithim to train models use RMSProp
# compute loss with function: categorical crossentropy
model.compile(optimizer='rmsprop',
loss='categorical_crossentropy',
metrics=['accuracy'])
return model
# + [markdown] id="xwo6uDcWYgzE"
# ## Model 9: Gated Recurrent Unit (GRU)
#
# โมเดลแบบที่ 9
#
#
# ---
#
#
#
# + id="zuTn1-rQYe_Z"
def build_GRU(image_shape):
sequence, features = image_shape
model = Sequential()
model.add(GRU( input_shape=(sequence, features),
units=100, dropout=0.2, recurrent_dropout=0.2, return_sequences=True))
model.add(GRU(100, dropout=0.2, recurrent_dropout=0.2, return_sequences=False))
model.add(Dense(10))
model.add(Activation("softmax")) #outputs are independent
# algorithim to train models use RMSProp
# compute loss with function: categorical crossentropy
model.compile(optimizer='rmsprop',
loss='categorical_crossentropy',
metrics=['accuracy'])
return model
# + [markdown] id="CYsNYxuuf5w4"
# # Visualize loss function and accuracy
#
# # # copy code from
#
# https://gist.github.com/stared/dfb4dfaf6d9a8501cd1cc8b8cb806d2e
# + id="0N6SwfYAf48Q"
import numpy as np
from matplotlib import pyplot as plt
from IPython.display import clear_output
class PlotLearning(keras.callbacks.Callback):
def on_train_begin(self, logs={}):
#self.i = 0
self.x = []
self.losses = []
self.val_losses = []
self.acc = []
self.val_acc = []
self.fig = plt.figure()
self.logs = []
def on_epoch_end(self, epoch, logs={}):
print("epoch: %d " % epoch)
self.logs.append(logs)
#self.x.append(self.i)
self.x.append(epoch)
self.losses.append(logs.get('loss'))
self.val_losses.append(logs.get('val_loss'))
self.acc.append(logs.get('acc'))
self.val_acc.append(logs.get('val_acc'))
#self.i += 1
if epoch%10 !=0 or epoch==0:
return # skip plot graph
f, (ax1, ax2) = plt.subplots(2, 1, sharex=True)
clear_output(wait=True)
ax1.set_yscale('log')
ax1.plot(self.x, self.losses, label="loss")
ax1.plot(self.x, self.val_losses, label="val_loss")
ax1.legend()
ax2.plot(self.x, self.acc, label="accuracy")
ax2.plot(self.x, self.val_acc, label="validation accuracy")
ax2.legend()
plt.show();
plot = PlotLearning()
# + [markdown] id="199FOZWfi9Xp"
# # Utility function
# + id="aX5jJGzyqUrV"
def restoreImg(X):
_, D = X.shape
W = int(math.sqrt(D))
assert D == W * W
X_image = X.reshape((-1, W, W))
return X_image
# + id="4JW8iz0wjMMg"
def decode(Ydigits):
# np.argmax: Returns the indices of the maximum values along an axis.
return np.array( [ np.argmax(list) for list in Ydigits] )
def get_prob(Y_predict):
Y_predict=np.array(Y_predict)
return np.array( [ np.max(list) for list in Y_predict] )
# + id="_Z7u7u7AlQjS"
def plotExampleImg(title, X_image, Ydigits, Y_predict=None):
fig, axarr = plt.subplots(2, 5)
axList = np.reshape(axarr, (2*5,))
plt.gcf().canvas.set_window_title(title)
fig.set_facecolor('#FFFFFF')
assert X_image.shape[0] == Ydigits.shape[0]
for num in range(0,10): # label 0 to 9
selectIndex = np.where(Ydigits == num)[0] # select all indexs followed the label number
digitsImg = X_image[selectIndex]
# random images
#Return random integers from 0 (inclusive) to high (exclusive).
if len(digitsImg)==0:
print("++picture of %s is not found+++" % num)
continue
randomIndex = np.random.randint(0, digitsImg.shape[0])
#axList[num].imshow(digitsImg[randomIndex], cmap=plt.cm.gray)
plt.gray()
axList[num].set_axis_off() # turn off axis x, y
axList[num].imshow(digitsImg[randomIndex])
if Y_predict is not None:
assert Ydigits.shape[0] == Y_predict.shape[0]
ySelect = Y_predict[selectIndex]
axList[num].set_title("%s=> %.2f%%" % (num, ySelect[randomIndex]*100))
else:
axList[num].set_title("Number %s" % num)
plt.tight_layout()
plt.show()
# + id="6yOmbxCak7Pm"
from sklearn import metrics
def testModel(model, X_image, Xtest, Yexpected, title_graph=""):
Ypredicted = model.predict(Xtest, verbose=0)
#Yexpected = decode(Yexpected) # convert binary to digits 0-9
Ypredicted_decode = decode(Ypredicted) # convert binary to digits 0-9
print("Classification report")
print(metrics.classification_report(Yexpected, Ypredicted_decode))
Y_max = np.array( [ np.max(list) for list in Ypredicted] )
plotExampleImg(title_graph, X_image, Ypredicted_decode, Y_max)
# + [markdown] id="HQkYMCNQbQZq"
# **ฟังก์ชั่นเตรียม datasets**
# + id="XYyjvWl3cOm3"
def preprocess_input(x_train, x_test):
# for input
x_train = x_train.reshape(60000,784)
x_test = x_test.reshape(10000,784)
return x_train, x_test
# + id="gaYPvtmROey6"
def preprocess_label(y_train, y_test):
# for label
y_train = keras.utils.to_categorical(y_train,10)
y_test = keras.utils.to_categorical(y_test,10)
return y_train, y_test
# + id="5OYwjxIORNuq" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="60605e74-7feb-4f36-dff3-5a743b8178c0"
x_train, y_train, x_test, y_test = get_datasets('mnist')
x_image = x_train
x_train, x_test = preprocess_input(x_train, x_test)
y_train, y_test = preprocess_label(y_train, y_test)
_, features = x_train.shape
print("Size of training input:", x_train.shape)
print("Size of testing input:", x_test.shape)
print("Size of training label:", y_train.shape)
print("Size of testing label:", y_test.shape)
# + [markdown] id="5zW3rZUpo_um"
# **ดูตัวอย่างภาพใน datasets**
# + id="wtxt9c17qztY" colab={"base_uri": "https://localhost:8080/", "height": 244} outputId="393f7079-552b-4813-b750-aee196e3bb05"
X_image = restoreImg(x_train)
Ydigits = decode(y_train)
plotExampleImg("Show image examples", X_image, Ydigits)
# + [markdown] id="wIdX0nidp3k6"
# เอาไว้เทสโมเดลตั้งแต่ตัวอย่างที่ 4-9
# + id="b0tBhFyCrCjw"
from sklearn import metrics
def test_model(model, X_testImage, x_test, y_test):
Y_testDigits = decode(y_test) # convert binary to digits 0-9
Y_predict = model.predict(x_test, verbose=0)
Y_predictDigits = decode(Y_predict) # convert binary to digits 0-9
print("++++++++++++++++++++++")
print("Classification report")
print(metrics.classification_report(Y_testDigits, Y_predictDigits))
Y_prob = get_prob(Y_predict)
plotExampleImg("Show image examples", X_testImage, Y_testDigits, Y_prob)
# + [markdown] id="d3OUptnXEwhZ"
# # ตัวอย่างและการใช้งาน
#
# + [markdown] id="3KtSbzuVQy3A"
# ## เตรียม Datasets สำหรับตัวอย่างที่ 1 - 3
# + id="A_mlU9BAQ26T"
x_train, y_train, x_test, y_test = get_datasets('mnist')
x_train, x_test = preprocess_input(x_train, x_test)
# + [markdown] id="g-8K4x0mIbLZ"
# ## ตัวอย่างที่ 1
#
# <h3>Nearest neighbors</h3>
#
# เนื่องจากตัวอย่างข้อมูลเยอะ เลยรันนานหน่อย
# <hr>
#
# + id="7a9E9ysTIof0" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="7634aedf-82d5-4d07-f30d-7b849ba4967b"
train_nearest_neighbors(x_train, y_train, x_test, y_test)
# + [markdown] id="2qofr--zIaZO"
# ## ตัวอย่างที่ 2
#
# <h3>Support vector</h3>
#
# เนื่องจากตัวอย่างข้อมูลเยอะ เลยรันนานหน่อย
# <hr>
# + id="juq3p5rQZVpA" colab={"base_uri": "https://localhost:8080/", "height": 351} outputId="2a2e7ea6-f3f5-444b-8990-8b0b6313b0b6"
import random
x_train = random.sample(x_train,10000)
y_train = random.sample(y_train,10000)
x_test = random.sample(x_test,10000)
y_test = random.sample(y_test,10000)
train_support_vector(x_train, y_train, x_test, y_test)
# + [markdown] id="wabIiVeFFW7D"
# ## ตัวอย่างที่ 3
#
# <h3>Logistic regression (1 neural)</h3>
#
#
# ---
#
#
# + id="Tvkvl-bgZWWp"
# + [markdown] id="tyBIwI1RQqcd"
# ## เตรียม Datasets สำหรับตัวอย่างที่ 4 - 9
# + id="1fW8bIZjQui8"
x_train, y_train, x_test, y_test = get_datasets('mnist')
x_image = x_train
x_train, x_test = preprocess_input(x_train, x_test)
y_train, y_test = preprocess_label(y_train, y_test)
# + [markdown] id="x1j0N_XKRE3H"
# **ดูตัวอย่างภาพใน datasets**
# + id="XSrL1dC5Q91P"
X_image = restoreImg(x_train)
Ydigits = decode(y_train)
plotExampleImg("Show image examples", X_image, Ydigits)
# + [markdown] id="tutrAc247o90"
# ## ตัวอย่างที่ 4
#
# <h3> Neural network (Multilayer Perceptron: MLP)</h3>
#
#
# ---
# + id="ryRZTuqCYIkt"
model = build_MLP(features)
model.summary()
#model.fit(x_train, y_train, batch_size=32, epochs=10, verbose=1)
model.fit(x_train, y_train,
batch_size=32, epochs=101,
validation_data=(x_test, y_test),
callbacks=[plot],
verbose=0)
# + id="XI8JuLJgfTju"
import random
index = random.randint(0, 100)
Y_predict = model.predict(x_test, verbose=0)
print(y_test[index])
print(Y_predict[index])
# + id="PSPdh3ZBorGx"
X_testImage = restoreImg(x_test)
test_model(model, X_testImage, x_test, y_test)
# + [markdown] id="x8FaANhTAOmT"
# ## ตัวอย่างที่ 5
#
# <h3> Convolutional Neural Networks (CNN) with Conv2D </h3>
#
#
# ---
#
# + id="14FQ0gL7yXZy"
# reshape to Theano: (batchsize, chanel, row, colum) or Tensorflow: (batchsize, row, column, chanel)
XtrainCNN = reshapeCNN2D_Input(x_train)
XtestCNN = reshapeCNN2D_Input(x_test)
image_shape = XtrainCNN.shape[1:] # select (chanel, row, column) or (row, column, chanel)
model = build_CNN_2D(image_shape)
model.summary()
#model.fit(x_train, y_train, batch_size=32, epochs=10, verbose=1)
model.fit(XtrainCNN, y_train,
batch_size=32, epochs=61,
validation_data=(XtestCNN, y_test),
callbacks=[plot],
verbose=0)
# + id="dYvsoFex0WUa"
X_testImage = restoreImg(x_test)
test_model(model, X_testImage, XtestCNN, y_test)
# + [markdown] id="z4zTRknWbhMh"
# ## ตัวอย่างที่ 6
#
# <h3>Convolutional Neural Networks (CNN) with Convolution1D</h3
#
#
# ---
#
# + id="lPTqpn_MIviw"
# reshape to Theano: (batchsize, chanel, row, colum) or Tensorflow: (batchsize, row, column, chanel)
XtrainCNN = reshapeCNN1D_Input(x_train)
XtestCNN = reshapeCNN1D_Input(x_test)
image_shape = XtrainCNN.shape[1:] # select (chanel, row, column) or (row, column, chanel)
model = build_CNN_1D(image_shape)
model.summary()
#model.fit(x_train, y_train, batch_size=32, epochs=10, verbose=1)
model.fit(XtrainCNN, y_train,
batch_size=32, epochs=101,
validation_data=(XtestCNN, y_test),
callbacks=[plot],
verbose=0)
# + id="W1wIwkRFbMqj"
X_testImage = restoreImg(x_test)
test_model(model, X_testImage, XtestCNN, y_test)
# + [markdown] id="EE41cKDpbm9y"
# ## ตัวอย่างที่ 7
#
# <h3> Recurrent Neural Networks (RNN) </h3>
#
#
#
# ---
#
# + id="ncXmiqVAbNFF"
print(np.shape(x_train))
# reshape to sequences for Recurrent Neural Networks
XtrainSeq = getSequenceInput(x_train)
XtestSeq = getSequenceInput(x_test)
image_shape = XtrainSeq.shape[1:] # select (row, column)
model = build_RNN(image_shape)
model.summary()
#model.fit(x_train, y_train, batch_size=32, epochs=10, verbose=1)
model.fit(XtrainSeq, y_train,
batch_size=32, epochs=31,
validation_data=(XtestSeq, y_test),
callbacks=[plot],
verbose=0)
# + id="_EVU7e7NgGuE"
X_testImage = restoreImg(x_test)
test_model(model, X_testImage, XtestSeq, y_test)
# + [markdown] id="wZX7fCXNFDsF"
# ## ตัวอย่างที่ 8
#
# <h3>Long short-term memory (LSTM)</h3>
#
#
#
# ---
#
# + id="wxsHi9dLRw00"
print(np.shape(x_train))
# reshape to sequences for Recurrent Neural Networks
XtrainSeq = getSequenceInput(x_train)
XtestSeq = getSequenceInput(x_test)
image_shape = XtrainSeq.shape[1:] # select (row, column)
model = build_LSTM(image_shape)
model.summary()
#model.fit(x_train, y_train, batch_size=32, epochs=10, verbose=1)
model.fit(XtrainSeq, y_train,
batch_size=32, epochs=11,
validation_data=(XtestSeq, y_test),
callbacks=[plot],
verbose=0)
# + id="QAa1f1xjFRvH"
X_testImage = restoreImg(x_test)
test_model(model, X_testImage, XtestSeq, y_test)
# + [markdown] id="TVARoCmGRcGZ"
# ## ตัวอย่างที่ 9
#
# <h3>Gated Recurrent Unit (GRU)</h3>
#
#
#
# ---
#
# + id="U6lXh9G0MDay"
print(np.shape(x_train))
# reshape to sequences for Recurrent Neural Networks
XtrainSeq = getSequenceInput(x_train)
XtestSeq = getSequenceInput(x_test)
image_shape = XtrainSeq.shape[1:] # select (row, column)
model = build_LSTM(image_shape)
model.summary()
#model.fit(x_train, y_train, batch_size=32, epochs=10, verbose=1)
model.fit(XtrainSeq, y_train,
batch_size=32, epochs=11,
validation_data=(XtestSeq, y_test),
callbacks=[plot],
verbose=0)
# + id="A9Lt3K6On5nc"
X_testImage = restoreImg(x_test)
test_model(model, X_testImage, XtestSeq, y_test)
# + id="J5sJ721GYy-n"
#accuracy = model.evaluate(x=x_test, y=y_test, batch_size=32)
#print("Accuracy: ",accuracy[1])
#model.predict(x, batch_size=None, verbose=0, steps=None, callbacks=None)
# + [markdown] id="18HSMklg3dYp"
# # ฝากติดตาม
#
# <img height="20px" src="https://www.patanasongsivilai.com/blog/wp-content/uploads/2017/06/logo_facebook2.png" align="left" hspace="10px" vspace="0px"> https://www.facebook.com/programmerthai/
#
# <img height="14px" src="https://www.patanasongsivilai.com/blog/wp-content/uploads/2019/03/youtube.png" align="left" hspace="10px" vspace="3px"> https://www.youtube.com/channel/UCvhCKtG5cRE6K5--RJPql_w
#
# <img height="14px" src="https://www.patanasongsivilai.com/blog/wp-content/uploads/2019/09/github-2.jpg" align="left" hspace="10px" vspace="3px">https://github.com/adminho/machine-learning
# + [markdown] id="vkTHggXeSSyR"
# # ฝากประชาสัมพันธ์
# + id="i7SSybi6SZf3" cellView="form"
#@title หนังสือ AI (ปัญญาประดิษฐ์) ไม่ยาก เรียนรู้ได้ด้วยเลขม. ปลาย ราคา 295 บาท
# %%HTML
<iframe src="https://www.facebook.com/plugins/post.php?href=https%3A%2F%2Fwww.facebook.com%2Fprogrammerthai%2Fphotos%2Fa.2410667485889755%2F2555677101388792%2F%3Ftype%3D3&width=500" width="500" height="588" style="border:none;overflow:hidden" scrolling="no" frameborder="0" allowTransparency="true" allow="encrypted-media"></iframe>
| 24_example_deep_learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sklearn
# !python -m pip install sklearn
# +
# %load_ext autoreload
# %autoreload 2
# -
import pandas as pd
# !dir .\files\auto-mpg.csv
pd_data = pd.read_csv('./files/auto-mpg.csv', header=None)
pd_data.info()
pd_data.columns = ['mpg','cylinders','displacement','horsepower','weight',
'acceleration','model year','origin','name']
pd_data.shape
pd_data[['mpg','weight']]
# 원 파일에 헤더가 없으므로 dataFrame을 만들면서 header=None으로 헤더가 없다는 것을 알려줌
#
# pd_data.info() -> 결측치 확인
# Dtype이 object인 것은 문자도 숫자도 될 수 있으므로 확인 (3,8 column 제외)
#
# pd_data의 column명 지정
# x축은 weight y축은 mpg 지정
x = pd_data[['weight']]
y = pd_data[['mpg']]
x.shape, y.shape
# split 기능을 가져옴 (두 집단으로 나눠서 하나는 식을 만들고 나머지는 만들어진 식에 대입하여 확인)
from sklearn.model_selection import train_test_split
X_train, X_test, Y_train, Y_test = train_test_split(x,y)
X_train.shape, X_test.shape, Y_train.shape, Y_test.shape
# sklearn.linear_model에서 선형회귀 기능을 import
from sklearn.linear_model import LinearRegression
lr = LinearRegression()
lr.fit(X_train, Y_train)
# lr에 직선방정식
# total data
lr.fit(x,y)
# split data with 1 column
lr.coef_, lr.intercept_
# x의 계수
lr.coef_
# y 절편
lr.intercept_
# weight * mpg의 선형식 : y = - 0.00767661x + 46.31736442
# 선형식이 얼마나 정확한지 확인하는 방법 : score (정확도)
lr.score(x,y)
| autompg_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Automating Regression with Scikit-Learn
# TO DO
# - Explain batch, mini-batch, and stochastic gradient descent.
# ## Introduction
#
# [Scikit-Learn](http://scikit-learn.org/stable/) is a Python library for machine learning. It is powerful, easy to use, and contains a lot of functionality. Perhaps for these reasons, it is also very widely used. Instead of programming models from scratch, we'll plug into Scikit-Learn's functionality. This will make our models more powerful, easier to build and experiment with, faster to deploy, and much more maintainable.
#
# In so doing we'll also be able to peer into some of the details of machine learning that will help us understand it better.
# Load up the packages to investigate the data
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.cm as cm
# %matplotlib inline
import seaborn as sns
import os
# OS-independent way to navigate the file system
# Data directory is one directory up in relation to directory of this notebook
data_dir_root = os.path.normpath(os.getcwd() + os.sep + os.pardir)
data_dir = data_dir_root + os.sep + "Data" + os.sep
# Where the file is
file_url = data_dir + "food-truck-profits.txt"
# Load the data into a dataframe
data = pd.read_csv(file_url, header=None, names=['Population', 'Profit'])
# When there's only a single column of data as input, SciKit likes it reshaped into a column vector
X_food = data['Population'].values.reshape(-1,1)
# y = data['Profit'].values.reshape(-1,1)
# SciKit likes the output vector as an array of values
y_food = data['Profit'].values
X_food.shape, y_food.shape
X_food[0:5]
y_food[0:5]
# ## Regression with One Feature
#
# Let's take the restaurant profit data and use Scikit-Learn to create a regression.
# Check version
import sklearn
sklearn.__version__
# Load up the package
from sklearn.linear_model import LinearRegression
# Linear regression model with all the Scikit defaults
lin_reg = LinearRegression()
lin_reg.fit(X_food,y_food)
# Get the optimal values of the parameters
[w_0, w_1] = np.array([lin_reg.intercept_, lin_reg.coef_]).flatten()
[w_0, w_1]
# +
# Make predictions -- it's as easy as calling '.predict' on the model
# The multiplier of 10000 puts the profits back in terms of the actual values
# Here are the profits for the first 5 rows of the dataset
profits = (lin_reg.predict(X_food[0:5]) * 10000).flatten()
# Format the values to make them easy to read
print(['${:5,.0f}'.format(profit) for profit in profits])
# +
# What does the regression line look like?
# Create 100 equally spaced values going from the minimum value of population
# to the maximum value of the population in the dataset.
x = np.linspace(data.Population.min(), data.Population.max(), 100)
f = (w_0 * 1) + (w_1 * x)
fig, ax = plt.subplots(figsize=(8,5))
ax.plot(x, f, 'g', label='Prediction')
ax.scatter(data.Population, data.Profit, label='Training Data')
ax.legend(loc='upper left')
ax.set_xlabel('Population')
ax.set_ylabel('Profit')
ax.set_title('Predicted Profit vs. Population Size')
# -
# This is the famous R-squared value -- not one we'll use directly in this course.
lin_reg.score(X_food,y_food)
# ## A Closer Look at Machine Learning
# The linear regression package we used is simple an clean. But let's dive deeper into how a machine learning algorithm works. By doing so we'll learn more about what it takes to find a model's optimal parameter values.
# The 'SGD' in SGDRegressor below stands for *Stochastic Gradient Descent*. That's a cool but somewhat scary-sounding name. But fear not, the concept of it is easy to understand. (Please see the lecture notes titled 'Batch, Mini-Batch, and Stochastic Gradient Descent' for a complete explanation of these terms.)
#
# We're using the SGDRegressor instead of the linear regressor because it allows us to examine the learning algorithm frame by frame if you will. We can play the algorithm like a movie reel and see how it goes about finding the optimal value(s) of the parameters of the model. We'll learn some weird things that
# The SGDRegressor (stochastic gradient descent) regressor gives us
# control over the number of iterations and amount over the learning rate
# and the number of iterations.
from sklearn.linear_model import SGDRegressor
# when learning_rate='constant' eta0 -- the starting learning rate -- stays the same for each iteration
# learning_rate='invscaling' progressively lowers the learning rate as the number of iterations increase
lin_sgd = SGDRegressor(max_iter=500, eta0=0.00001, learning_rate='constant')
lin_sgd.fit(X_food,y_food)
# There are a lot of inputs to SGDRegressor -- the package that does stochastic gradient descent. These inputs determine how the learning algorithm goes about finding the optimal weights for the parameters. The two inputs we're concered with are:
# - eta0 which is the learning rate -- the size of the step taken in each iteration of gradient descent
# - max_iter -- the number of times that next step is taken
#
# By altering the values of these *hyper-parameters*, we can alter the result that the learning algorithm arrives at.
[w_0, w_1] = np.array([lin_sgd.intercept_, lin_sgd.coef_]).flatten()
[w_0, w_1]
profits = lin_sgd.predict(X_food[0:1]) * 10000
print(['${:5,.0f}'.format(profit) for profit in profits])
# ### EXERCISE
#
# 1. Change eta0 from 0.0001 to 0.1 in a set number of steps. Describe what happens to the predicted profit of the first restaurant in the dataset. Hold the number of iterations at 1000 throughout. What do you conclude about the SGDRegressor model?
#
# 2. Change the number of iterations from 1 to 1000 in a set number of steps. Describe what happens to the profit of the first restaurant in the dataset. Hold eta0 at 0.0001 throughout. What do you conclude about the SGDRegressor model?
# Doing the exercise
# The model is fit each time with the values of eta_0 and num_iterations
from sklearn.linear_model import SGDRegressor
def get_results(input_array, output_array, eta_0, num_iterations, num_results=1):
# num_results are the number of rows of the dataset for which predictions are needed
# e.g., num_results=3 will give predictions for the first 3 rows of the dataset
# Load up the SGDRegressor model with the given parameters
sgdR = SGDRegressor(penalty='none', max_iter=num_iterations, learning_rate='constant', eta0=eta_0)
# Fit the model to the inputs and the outputs (the training data)
sgdR.fit(input_array, output_array)
# Get the parameter values
W = np.array([sgdR.intercept_, sgdR.coef_]).flatten()
# Get the predicted price(s)
results = sgdR.predict(input_array[0:num_results])
return [W, results]
# Test it out
[W, results] = get_results(X_food, y_food, 0.01, 1, num_results=1)
print(W)
print(['${:5,.0f}'.format(result) for result in results])
# Run the function with the SAME values multiple times
# Notice how the profit fluctuates for the exact same input values to the function
# This is very worrisome! What's happening???
# Hint: It has to do with the "Stochastic" in "Stochastic Gradient Descent"
for i in range(5):
[W, results] = get_results(X_food, y_food, eta_0=0.03, num_iterations=10, num_results=1)
print(W)
print(['${:5,.0f}'.format(result) for result in results])
# ### Profit versus Learning Rate for a Fixed Number of Iterations
eta0_values = [0.01, 0.003, 0.001, 0.0003, 0.0001]
eta0_axis = [str(eta0_val) for eta0_val in eta0_values] # axis labels for the plot
results = [get_results(X_food, y_food, eta0_val , 1000, num_results=1) for eta0_val in eta0_values]
# Get the w0, w1, and profit values
w0_values = [results[i][0][0] for i in range(len(results))]
w1_values = [results[i][0][1] for i in range(len(results))]
profit_values = list([results[i][1] for i in range(len(results))])
print(w0_values)
print(w1_values)
print(profit_values)
profit_values
# How the profit changes as the learning rate increases
# Learning faster is not necessarily learning better!
fig, ax = plt.subplots(figsize=(8,5))
ax.plot(eta0_values, profit_values, 'g')
ax.set_xlabel('Learning Rate')
ax.set_ylabel('Profit')
ax.set_title('Predicted Profit vs. Learning Rate (# Iterations Fixed at 1000)')
# How the w0 and w1 values change as the learning rate increases
fig, ax = plt.subplots(figsize=(8,5))
ax.plot(eta0_values, w0_values, 'g', label="w0")
ax.plot(eta0_values, w1_values, 'y', label="w1")
ax.set_xlabel('Learning Rate')
ax.set_ylabel('Parameter Values')
ax.set_title('Parameter Values vs. Learning Rate (# Iterations Fixed at 1000)')
ax.legend()
# ### Profit versus Number of Iterations (Learning Rate is Fixed)
iter_values = [1, 10, 50, 100, 250, 500, 1000, 5000]
results = [get_results(X_food, y_food, 0.0001 , iter_val) for iter_val in iter_values]
# Get the w0, w1, and profit values
w0_values = [results[i][0][0] for i in range(len(results))]
w1_values = [results[i][0][1] for i in range(len(results))]
profit_values = [results[i][1] for i in range(len(results))]
# How the profit changes as the num of iterations increase
# More iterations are always better (up to a point)
fig, ax = plt.subplots(figsize=(8,5))
ax.plot(iter_values, profit_values, 'g')
ax.set_xlabel('Number of Iterations')
ax.set_ylabel('Profit')
ax.set_title('Predicted Profit vs. Number of Iterations (Learning Rate Fixed at 0.0001)')
# How the w0 and w1 values change as the number of iterations increase
fig, ax = plt.subplots(figsize=(8,5))
ax.plot(iter_values, w0_values, 'g', label="w0")
ax.plot(iter_values, w1_values, 'y', label="w1")
ax.set_xlabel('Number of Iterations')
ax.set_ylabel('Parameter Values')
ax.set_title('Parameter Values vs. Number of Iterations (Learning Rate Fixed at 0.0001)')
ax.legend()
# ## Regression with Multiple Features
# Load the housing prices dataset
file_url = data_dir_root + os.sep + "Data" + os.sep + "portland-house-prices.txt"
# Load the data into a dataframe
data2 = pd.read_csv(file_url, header=None, names=['Size', 'Bedrooms', 'Price'])
data2.head()
# Because the number of bedrooms is three orders of magnitude lower than the sizes of the houses in square feet, we need to scale the features so they are comparable. We do this using Scikit-Learn's preprocessing functions.
X = data2.iloc[:,0:2] # Don't have to explicitly reshape it
y = data2['Price'].values
X.shape, y.shape
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
X_scaled = scaler.fit_transform(X)
X_scaled[0:3]
lin_reg_m = LinearRegression()
lin_reg_m.fit(X_scaled,y)
# Get the optimal values of the parameters
w0 = lin_reg_m.intercept_
[w1, w2] = lin_reg_m.coef_
[w0, w1, w2]
# Make some predictions
prices = lin_reg_m.predict(X_scaled[0:5])
print(['${:5,.0f}'.format(price) for price in prices.flatten()])
# The traditional R-squared score
lin_reg_m.score(X_scaled,y)
# ### SGDRegressor for the housing prices dataset
# Run the SGD Regression and get the results
# Vary eta_0 and num_iterations to get a feel for how the learning algorithm behaves
[[[w0],[w1,w2]],prices] = get_results(X_scaled, y, eta_0=0.0009, num_iterations=1, num_results=1)
# Get the w0, w1, w2 parameter values and the price predictions
print(w0, w1, w2)
print(['${:5,.0f}'.format(price) for price in prices])
# ### Profit versus Learning Rate for a Fixed Number of Iterations
eta0_values = [0.01, 0.003, 0.001, 0.0003, 0.0001]
eta0_axis = [str(eta0_val) for eta0_val in eta0_values] # axis labels for the plot (not necessary)
results = [get_results(X_scaled, y, eta0_val , 1000) for eta0_val in eta0_values]
# Get the w0, w1, w2 parameter values and the price value(s)
w0_values = [results[i][0][0] for i in range(len(results))]
w1_values = [results[i][0][1][0] for i in range(len(results))]
w2_values = [results[i][0][1][1] for i in range(len(results))]
price_values = [results[i][1] for i in range(len(results))]
# How the price changes as the learning rate increases
# Learning faster is not necessarily learning better!
fig, ax = plt.subplots(figsize=(8,5))
ax.plot(eta0_values, price_values, 'g')
ax.set_xlabel('Learning Rate')
ax.set_ylabel('House Price')
ax.set_title('Predicted House Price vs. Learning Rate (# Iterations Fixed at 1000)')
# How the w0, w1, and w2 values change as the learning rate increases
fig, ax = plt.subplots(figsize=(8,5))
ax.plot(eta0_values, w0_values, 'g', label="w0")
ax.plot(eta0_values, w1_values, 'y', label="w1")
ax.plot(eta0_values, w2_values, 'b', label="w2")
ax.set_xlabel('Learning Rate')
ax.set_ylabel('Parameter Values')
ax.set_title('Parameter Values vs. Learning Rate (# Iterations Fixed at 1000)')
ax.legend()
# ### House Price versus Number of Iterations (Learning Rate is Fixed)
iter_values = [1, 10, 50, 100, 250, 500, 1000, 3000]
results = [get_results(X_scaled, y, 0.0001 , iter_val) for iter_val in iter_values]
# Get the w0, w1, w2 parameter values and the price value(s)
w0_values = [results[i][0][0] for i in range(len(results))]
w1_values = [results[i][0][1][0] for i in range(len(results))]
w2_values = [results[i][0][1][1] for i in range(len(results))]
price_values = [results[i][1] for i in range(len(results))]
# How the house price changes as the num of iterations increase
# More iterations are always better (up to a point)
fig, ax = plt.subplots(figsize=(8,5))
ax.plot(iter_values, price_values, 'g')
ax.set_xlabel('Number of Iterations')
ax.set_ylabel('House Price')
ax.set_title('Predicted House Price vs. Number of Iterations (Learning Rate Fixed at 0.0001)')
# How the w0, w1, and w2 values change as the number of iterations increase
fig, ax = plt.subplots(figsize=(8,5))
ax.plot(iter_values, w0_values, 'g', label="w0")
ax.plot(iter_values, w1_values, 'y', label="w1")
ax.plot(iter_values, w2_values, 'b', label="w2")
ax.set_xlabel('Number of Iterations')
ax.set_ylabel('Parameter Values')
ax.set_title('Parameter Values vs. Number of Iterations (Learning Rate Fixed at 0.0001)')
ax.legend()
# ### QUESTION
#
# Why do the iterations look much better (more stable) for the multi-feature regression?
# ## Summary and Looking Ahead
#
# 1. Hyperparameter settings such as the learning rate and the number of iterations control and influence how a learning algorithm arrives at its results.
#
# 2. Machine learning models depend on these learning algorithms. Therefore, the predictions of machine learning models depend on the particular choices made for the hyperparameter values. As we've seen, model predictions are sensitive to these settings.
#
# 3. So what are the right settings? It would be those settings that result in the best model.
#
# 4. What's the best model? It's the one with the best performance.
#
# 5. So how do we define the performance of a model and how do we measure it? That's what we'll turn to next.
| Notebooks/Regression-with-Scikit-Learn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] nbsphinx="hidden"
# # Spectral Analysis of Deterministic Signals
#
# *This jupyter notebook is part of a [collection of notebooks](../index.ipynb) on various topics of Digital Signal Processing. Please direct questions and suggestions to [<EMAIL>](mailto:<EMAIL>).*
# -
# ## Introduction
#
# The analysis of the spectral properties of a signal plays an important role in digital signal processing. Some application examples are the
#
# * [Spectrum analyzer](https://en.wikipedia.org/wiki/Spectrum_analyzer)
# * Detection of (harmonic) signals
# * [Estimation of fundamental frequency and harmonics](https://en.wikipedia.org/wiki/Modal_analysis)
# * Spectral suppression: acoustic echo suppression, noise reduction, ...
#
# In the practical realization of spectral analysis techniques, the [discrete Fourier transform](https://en.wikipedia.org/wiki/Discrete_Fourier_transform) (DFT) is applied to discrete finite-length signals in order to gain insights into their spectral composition. A basic task in spectral analysis is to determine the amplitude (and phase) of dominant harmonic contributions in a signal mixture. The properties of the DFT with respect to the analysis of an harmonic exponential signal are discussed in the following.
# ## The Leakage Effect
#
# [Spectral leakage](https://en.wikipedia.org/wiki/Spectral_leakage) is a fundamental effect of the DFT. It limits the ability to detect harmonic signals in signal mixtures and hence the performance of spectral analysis. In order to discuss this effect, the DFT of a discrete exponential signal is revisited starting from the Fourier transform of the continuous exponential signal. The connections between the Fourier transform, the [discrete-time Fourier transform](https://en.wikipedia.org/wiki/Discrete-time_Fourier_transform) (DTFT) and the DFT for a uniformly sampled signal are illustrated below.
#
# 
#
# Consequently, the leakage effect is discussed in the remainder of this section by considering the following four steps:
#
# 1. Fourier transform of an harmonic exponential signal,
# 2. discrete-time Fourier transform (DTFT) of a discrete harmonic exponential signal, and
# 3. DTFT of a finite-length discrete harmonic exponential signal
# 4. sampling of the DTFT.
# ### Fourier Transform of an Exponential Signal
#
# The harmonic exponential signal is defined as
#
# \begin{equation}
# x(t) = \mathrm{e}^{\,\mathrm{j}\, \omega_0 \, t}
# \end{equation}
#
# where $\omega_0 = 2 \pi f$ denotes the angular frequency of the signal. The Fourier transform of the exponential signal is
#
# \begin{equation}
# X(\mathrm{j}\, \omega) = \int\limits_{-\infty}^{\infty} x(t) \,\mathrm{e}^{\,- \mathrm{j}\, \omega \,t} \mathrm{d}t = 2\pi \; \delta(\omega - \omega_0)
# \end{equation}
#
# The spectrum consists of a single shifted Dirac impulse located at the angular frequency $\omega_0$ of the exponential signal. Hence the spectrum $X(\mathrm{j}\, \omega)$ consists of a clearly isolated and distinguishable event. In practice, it is not possible to compute the Fourier transform of a continuous signal by means of digital signal processing.
# ### Discrete-Time Fourier Transform of a Discrete Exponential Signal
#
# Now lets consider sampled signals. The discrete exponential signal $x[k]$ is derived from its continuous counterpart $x(t)$ above by equidistant sampling $x[k] := x(k T)$ with the sampling interval $T$
#
# \begin{equation}
# x[k] = \mathrm{e}^{\,\mathrm{j}\, \Omega_0 \,k}
# \end{equation}
#
# where $\Omega_0 = \omega_0 T$ denotes the normalized angular frequency. The DTFT is the Fourier transform of a sampled signal. For the exponential signal it is given as (see e.g. [reference card discrete signals and systems](../reference_cards/RC_discrete_signals_and_systems.pdf))
#
# \begin{equation}
# X(\mathrm{e}^{\,\mathrm{j}\, \Omega}) = \sum_{k = -\infty}^{\infty} x[k]\, \mathrm{e}^{\,-\mathrm{j}\, \Omega \,k} = 2\pi \sum_{n = -\infty}^{\infty} \delta((\Omega-\Omega_0) - 2\,\pi\,n)
# \end{equation}
#
# The spectrum of the DTFT is $2\pi$-periodic due to sampling. As a consequence, the transform of the discrete exponential signal consists of a series Dirac impulses. For the region of interest $-\pi < \Omega \leq \pi$ the spectrum consists of a clearly isolated and distinguishable event, as for the continuous case.
#
# The DTFT cannot be realized in practice, since is requires the knowledge of the signal $x[k]$ for all time instants $k$. In general, a measured signal is only known within a finite time-interval. The DFT of a signal of finite length can be derived from the DTFT in two steps:
#
# 1. truncation (windowing) of the signal and
# 2. sampling of the DTFT spectrum of the windowed signal.
#
# The consequences of these two steps are investigated in the following.
# ### Discrete-Time Fourier Transform of a Truncated Discrete Exponential Signal
#
# In general, truncation of a signal $x[k]$ to a length of $N$ samples is modeled by multiplying the signal with a window function $w[k]$ of length $N$
#
# \begin{equation}
# x_N[k] = x[k] \cdot w[k]
# \end{equation}
#
# where $x_N[k]$ denotes the truncated signal and $w[k] = 0$ for $\{k: k < 0 \wedge k \geq N \}$. The spectrum $X_N(\mathrm{e}^{\,\mathrm{j}\, \Omega})$ can be derived from the multiplication theorem of the DTFT as
#
# \begin{equation}
# X_N(\mathrm{e}^{\,\mathrm{j}\, \Omega}) = \frac{1}{2 \pi} X(\mathrm{e}^{\,\mathrm{j}\, \Omega}) \circledast_N W(\mathrm{e}^{\,\mathrm{j}\, \Omega})
# \end{equation}
#
# where $\circledast_N$ denotes the cyclic/[circular convolution](https://en.wikipedia.org/wiki/Circular_convolution) of length $N$. A hard truncation of the signal to $N$ samples is modeled by the rectangular signal
#
# \begin{equation}
# w[k] = \text{rect}_N[k] = \begin{cases}
# 1 & \mathrm{for} \; 0\leq k<N \\
# 0 & \mathrm{otherwise}
# \end{cases}
# \end{equation}
#
# Its spectrum is given as
#
# \begin{equation}
# W(\mathrm{e}^{\,\mathrm{j}\, \Omega}) = \mathrm{e}^{\,-\mathrm{j} \, \Omega \,\frac{N-1}{2}} \cdot \frac{\sin(\frac{N \,\Omega}{2})}{\sin(\frac{\Omega}{2})}
# \end{equation}
#
# The DTFT $X_N(\mathrm{e}^{\,\mathrm{j}\, \Omega})$ of the truncated exponential signal is derived by introducing the DTFT of the exponential signal and the window function into above cyclic convolution. Since, both the DTFT of the exponential signal and the window function are periodic with a period of $2 \pi$, the cyclic convolution with period $2 \pi$ is given by linear convolution of both spectra within $-\pi < \Omega \leq \pi$
#
# \begin{equation}
# X_N(\mathrm{e}^{\,\mathrm{j}\, \Omega}) = \delta(\Omega-\Omega_0) * \mathrm{e}^{\,-\mathrm{j} \, \Omega \,\frac{N-1}{2}} \cdot \frac{\sin(\frac{N \,\Omega}{2})}{\sin(\frac{\Omega}{2})} =
# \mathrm{e}^{\,-\mathrm{j}\, (\Omega-\Omega_0) \, \frac{N-1}{2}} \cdot \frac{\sin(\frac{N\, (\Omega-\Omega_0)}{2})}{\sin(\frac{(\Omega-\Omega_0)}{2})}
# \end{equation}
#
# Note that $X_N(\mathrm{e}^{\,\mathrm{j}\, \Omega})$ is periodic with a period of $2 \pi$. Clearly the DTFT of the truncated harmonic exponential signal $x_N[k]$ is not given by a series of Dirac impulses. Above equation is evaluated numerically in order to illustrate the properties of $X_N(\mathrm{e}^{\,\mathrm{j}\, \Omega})$.
# +
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
Om0 = 1 # frequency of exponential signal
N = 32 # length of signal
# DTFT of finite length exponential signal (analytic)
Om = np.linspace(-np.pi, np.pi, num=1024)
XN = np.exp(-1j * (Om-Om0) * (N-1) / 2) * (np.sin(N * (Om-Om0) / 2)) / (np.sin((Om-Om0) / 2))
# plot spectrum
plt.figure(figsize = (10, 8))
plt.plot(Om, abs(XN), 'r')
plt.title(r'Absolute value of the DTFT of a truncated exponential signal $e^{j \Omega_0 k}$ with $\Omega_0=$%2.2f' %Om0)
plt.xlabel(r'$\Omega$')
plt.ylabel(r'$|X_N(e^{j \Omega})|$')
plt.axis([-np.pi, np.pi, -0.5, N+5])
plt.grid()
# -
# **Exercise**
#
# * Change the frequency `Om0` of the signal and rerun the example. How does the magnitude spectrum change?
# * Change the length `N` of the signal and rerun the example. How does the magnitude spectrum change?
#
# Solution: The maximum of the absolute value of the spectrum is located at the frequency $\Omega_0$. It should become clear that truncation of the exponential signal leads to a broadening of the spectrum. The shorter the signal, the wider the mainlobe becomes.
# ### The Leakage Effect of the Discrete Fourier Transform
#
# The DFT is derived from the DTFT $X_N(\mathrm{e}^{\,\mathrm{j}\, \Omega})$ of the truncated signal $x_N[k]$ by sampling the DTFT equidistantly at $\Omega = \mu \frac{2 \pi}{N}$
#
# \begin{equation}
# X[\mu] = X_N(\mathrm{e}^{\,\mathrm{j}\, \Omega})\big\vert_{\Omega = \mu \frac{2 \pi}{N}}
# \end{equation}
#
# For the DFT of the exponential signal we finally get
#
# \begin{equation}
# X[\mu] = \mathrm{e}^{\,\mathrm{j}\, (\Omega_0 - \mu \frac{2 \pi}{N}) \frac{N-1}{2}} \cdot \frac{\sin(\frac{N \,(\Omega_0 - \mu \frac{2 \pi}{N})}{2})}{\sin(\frac{\Omega_0 - \mu \frac{2 \pi}{N}}{2})}
# \end{equation}
#
# The sampling of the DTFT is illustrated in the following example. Note that the normalized angular frequency $\Omega_0$ has been expressed in terms of the periodicity $P$ of the exponential signal $\Omega_0 = P \; \frac{2\pi}{N}$.
# +
N = 32 # length of the signal
P = 10.33 # periodicity of the exponential signal
Om0 = P * (2*np.pi/N) # frequency of exponential signal
# truncated exponential signal
k = np.arange(N)
x = np.exp(1j*Om0*k)
# DTFT of finite length exponential signal (analytic)
Om = np.linspace(0, 2*np.pi, num=1024)
Xw = np.exp(-1j*(Om-Om0)*(N-1)/2)*(np.sin(N*(Om-Om0)/2))/(np.sin((Om-Om0)/2))
# DFT of the exponential signal by FFT
X = np.fft.fft(x)
mu = np.arange(N) * 2*np.pi/N
# plot spectra
plt.figure(figsize = (10, 8))
ax1 = plt.gca()
plt.plot(Om, abs(Xw), 'r', label=r'$|X_N(e^{j \Omega})|$')
plt.stem(mu, abs(X), label=r'$|X_N[\mu]|$', basefmt=' ')
plt.ylim([-0.5, N+5]);
plt.title(r'Absolute value of the DTFT/DFT of a truncated exponential signal $e^{j \Omega_0 k}$ with $\Omega_0=$%2.2f' %Om0, y=1.08)
plt.legend()
ax1.set_xlabel(r'$\Omega$')
ax1.set_xlim([Om[0], Om[-1]])
ax1.grid()
ax2 = ax1.twiny()
ax2.set_xlim([0, N])
ax2.set_xlabel(r'$\mu$', color='C0')
ax2.tick_params('x', colors='C0')
# -
# **Exercise**
#
# * Change the periodicity `P` of the exponential signal and rerun the example. What happens if the periodicity is an integer? Why?
# * Change the length `N` of the DFT? How does the spectrum change?
# * What conclusions can be drawn for the analysis of a single exponential signal by the DFT?
# Solution: You should have noticed that for an exponential signal whose periodicity is an integer $P \in \mathbb{N}$, the DFT consists of a discrete Dirac pulse $X[\mu] = N \cdot \delta[\mu - P]$. In this case, the sampling points coincide with the maximum of the main lobe or the zeros of the DTFT. For non-integer $P$, hence non-periodic exponential signals with respect to the signal length $N$, the DFT has additional contributions. The shorter the length $N$, the wider these contributions are spread in the spectrum. This smearing effect is known as *leakage effect* of the DFT. It limits the achievable frequency resolution of the DFT when analyzing signal mixtures consisting of more than one exponential signal or exponential signals under additive noise. This is illustrated by the following numerical examples.
# ### Analysis of Signal Mixtures by the Discrete Fourier Transform
#
# In order to discuss the implications of the leakage effect when analyzing signal mixtures, the superposition of two exponential signals with different amplitudes and frequencies is considered
#
# \begin{equation}
# x_N[k] = A_1 \cdot e^{\mathrm{j} \Omega_1 k} + A_2 \cdot e^{\mathrm{j} \Omega_2 k}
# \end{equation}
#
# where $A_1, A_2 \in \mathbb{R}$. For convenience, a function is defined that calculates and plots the magnitude spectrum of $x_N[k]$.
def dft_signal_mixture(N, A1, P1, A2, P2):
# N: length of signal/DFT
# A1, P1, A2, P2: amplitude and periodicity of 1st/2nd complex exponential
# generate the signal mixture
Om0_1 = P1 * (2*np.pi/N) # frequency of 1st exponential signal
Om0_2 = P2 * (2*np.pi/N) # frequency of 2nd exponential signal
k = np.arange(N)
x = A1 * np.exp(1j*Om0_1*k) + A2 * np.exp(1j*Om0_2*k)
# DFT of the signal mixture
mu = np.arange(N)
X = np.fft.fft(x)
# plot spectrum
plt.figure(figsize = (10, 8))
plt.stem(mu, abs(X), basefmt=' ')
plt.title(r'Absolute value of the DFT of a signal mixture')
plt.xlabel(r'$\mu$')
plt.ylabel(r'$|X[\mu]|$')
plt.axis([0, N, -0.5, N+5]);
plt.grid()
# Lets first consider the case that the frequencies of the two exponentials are rather far apart in terms of normalized angular frequency
dft_signal_mixture(32, 1, 10.3, 1, 15.2)
# Investigating the magnitude spectrum one could conclude that the signal consists of two major contributions at the frequencies $\mu_1 = 10$ and $\mu_2 = 15$. Now lets take a look at a situation where the frequencies are closer together
dft_signal_mixture(32, 1, 10.3, 1, 10.9)
# From visual inspection of the spectrum it is rather unclear if the mixture consists of one or two exponential signals. So far the levels of both signals where chosen equal.
#
# Lets consider the case where the second signal has a much lower level that the first one. The frequencies have been chosen equal to the first example
dft_signal_mixture(32, 1, 10.3, 0.1, 15.2)
# Now the contribution of the second exponential is almost hidden in the spread spectrum of the first exponential. From these examples it should have become clear that the leakage effect limits the spectral resolution of the DFT.
# + [markdown] nbsphinx="hidden"
# **Copyright**
#
# This notebook is provided as [Open Educational Resource](https://en.wikipedia.org/wiki/Open_educational_resources). Feel free to use the notebook for your own purposes. The text is licensed under [Creative Commons Attribution 4.0](https://creativecommons.org/licenses/by/4.0/), the code of the IPython examples under the [MIT license](https://opensource.org/licenses/MIT). Please attribute the work as follows: *<NAME>, Digital Signal Processing - Lecture notes featuring computational examples, 2016-2018*.
| spectral_analysis_deterministic_signals/leakage_effect.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import seaborn as sns
from matplotlib import pyplot as plt
import pandas as pd
filename = ("housing.csv")
names = ['CRIM', 'ZN', 'INDUS', 'CHAS', 'NOX', 'RM', 'AGE', 'DIS', 'RAD', 'TAX', 'PTRATIO', 'B', 'LSTAT', 'MEDV']
dataset = pd.read_csv(filename, delim_whitespace=True, names=names)
prices = dataset['MEDV']
dataset = dataset.drop(['CRIM','ZN','INDUS','NOX','AGE','DIS','RAD', 'CHAS','PTRATIO','TAX','B'], axis = 1)
features = dataset.drop('MEDV', axis = 1)
dataset.head()
x1 = features['RM'].values
x2 = features['LSTAT'].values
Y= prices.values
m = len(x1)
x0 = np.ones(m)
#we assume x0 to be 1
X = np.array([x0, x1, x2]).T
#.T is used to obtain transpose
# Initial Coefficients of B is assumed to be 0
#our eqn will be in form of b0 + b1*x1 +b2*x2
B = np.zeros(3)
Y = np.array(Y)
alpha = 0.00012
#alpha is learning rate
def cost_function(X, Y, B):
m = len(Y)
J = np.sum((X.dot(B) - Y) ** 2)/(2 * m)
return J
inital_cost = cost_function(X, Y, B)
print(inital_cost)
def grad_desc(X, Y, B, a, iteration):
cost_iter = [0] * iteration
#is used to obtain a plot of how gradient descent works
m = len(Y)
for i in range(iteration):
# value obtained from hypothesis
h = X.dot(B)
# Difference between Hypothesis and Actual Y
#note that loss is calculate for all values so we divide it by m to obtain average loss
loss = h - Y
# Gradient Calculation
gradient = X.T.dot(loss) / m
# Changing Values of B using Gradient
B = B - a * gradient
# New Cost Value
cost = cost_function(X, Y, B)
cost_iter[i] = cost
return B, cost_iter
newB, cost_history = grad_desc(X, Y, B, alpha, 3000)
Ypred=X.dot(newB)
from mpl_toolkits.mplot3d import Axes3D
x = np.linspace(0,35,30)
y = np.linspace(0,10,30)
e,r = np.meshgrid(x,y)
Z=newB[0]+newB[1]*r + newB[2]*e
threedee = plt.figure().gca(projection='3d')
threedee.scatter(dataset['LSTAT'],dataset['RM'],dataset['MEDV'], color='r')
threedee.plot_surface(e, r, Z, color='y')
threedee.set_xlabel('LSTAT')
threedee.set_ylabel('RM')
threedee.set_zlabel('MEDV')
plt.figure(figsize=(500,400))
plt.show()
x=np.arange(1,3001)
plt.plot(x, cost_history)
plt.xlabel('iteration')
plt.ylabel('cost function')
plt.show()
def rmse(Y, Y_pred):
rmse = np.sqrt(sum((Y - Y_pred) ** 2) / len(Y))
return rmse
print(rmse(Y,Ypred))
| Regression and Classification from Scratch/linear regression from scratch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# %matplotlib inline
import numpy as np
import pandas as pd
import scipy.stats as stats
import matplotlib.pyplot as plt
import sklearn
import statsmodels.api as sm
from sklearn.linear_model import LinearRegression
from sklearn.linear_model import Lasso
from sklearn import metrics
from sklearn.grid_search import GridSearchCV
from sklearn.cross_validation import train_test_split
from sklearn.metrics import confusion_matrix
import seaborn as sns
sns.set_style("whitegrid")
sns.set_context("poster")
import warnings
warnings.filterwarnings('ignore')
from sklearn import cross_validation
from matplotlib import rcParams
from bs4 import BeautifulSoup
from pyquery import PyQuery as pq
from collections import defaultdict
from imdb import IMDb
import pandas as pd
import cPickle as pickle
import io
import time
import requests
from imdb import IMDb
ia = IMDb()
def get_mpaa(movieobj):
try:
mpaa = str(movieobj.data['mpaa']).split("Rated ", 1)[1].split(" ")[0]
except:
mpaa = np.nan
return mpaa
# -
# ##1. Background, Motivation and Related Work
#
# If one were to ask most movie executives what the most important quality of their movies should be, they would probably jump to the obvious answer: "Artistic depth and quality"
#
# ...Well, perhaps obvious to the die-hard movie afficianado, but in reality, the bottom line is probably what's most pressingly on the minds of any movie-making machine.
#
# So what strategies are most likely to make a film a box office smash? Over the coming analyses, we will explore the levers that a would-be producer might pull to maximize their sales.
#
# Slightly differently from the Oscar analysis, there has been limited work on the questions around Movie Gross. We did consider a paper out of Stanford http://cs229.stanford.edu/proj2013/vanderMerweEimon-MaximizingMovieProfit.pdf that delves into some of the concepts we are looking to address here.
#
# This Process Notebook outlines the following exercises performed to arrive at a results for the analysis:
#
# 1. Background, Motivation and Related Work
# 2. Initial Questions
# 3. Data Transformation: Dataframes, standardization and collation
# 4. Exploratory Data Analysis
# 5. Model Building using StatsModel
# 6. a. Leveraging sklearn as an alternative to StatsModel
# 7. b. Hyperparameter tuning using Lasso Regression in sklearn
# 8. Results and Conclusion
#
#
#
# ##2. Initial Questions
#
# - **Movie Budget**: Wealth begets wealth, goes the old adage. For studios looking to make a splash, how important are the investment dollars put against a production?
# - **Opening Weekend**: Film industry experts love the 'Opening Weekend' phenomenon, but just how much does the first 4 days of a movie's release truly matter?
# - **Opening Theaters**: Does the scale of release during the opening weekend affect a movie?
# - **IMDb rating**: Does how 'good' the movie is actually matter? We leverage the IMDb rating as a proxy for the 'inherent level of quality' of the movie.
# - **Seasonality**: How does a release around Christmas or the summer vacation affect the outcomes of a movie?
# - **MPAA Rating**: For the kids, the parents, or the young adults?
# - **Power Studios**: Do movie production houses like Warner Bros. or Universal have a power of their own, besides the factors listed above?
# ##3. Data Transformation
BOdict = pickle.load(io.open('BOdict.p', 'rb'))
BOdf = pd.DataFrame(BOdict).transpose()
# +
##Culling the dataset down to ensure we have non-null responses in our keys variables
limiteddf = BOdf.dropna(subset=['budget', 'season', 'mpaa', 'opening'])
## Ensuring that the number values are not in text format
limiteddf['gross'].replace(regex=True,inplace=True,to_replace=r'\D',value=r'')
limiteddf['opening'].replace(regex=True,inplace=True,to_replace=r'\D',value=r'')
limiteddf['opening theaters'].replace(regex=True,inplace=True,to_replace=r'\D',value=r'')
##Replacing empty values
limiteddf.loc[limiteddf['opening']=='', 'opening'] = 0
limiteddf.loc[limiteddf['opening theaters']=='', 'opening theaters'] = 0
##Converting to float values for numerical variables
limiteddf['gross'] = limiteddf['gross'].astype(float)
limiteddf['opening'] = limiteddf['opening'].astype(float)
limiteddf['opening theaters'] = limiteddf['opening theaters'].astype(float)
limiteddf['budget'] = limiteddf['budget'].astype(float)
limiteddf['rating'] = limiteddf['rating'].astype(float)
##Converting to season (as necessary)
#limiteddf.loc[limiteddf['season']==0, 'season'] = 'Jan-May'
#limiteddf.loc[limiteddf['season']==1, 'season'] = 'June-Sep'
#limiteddf.loc[limiteddf['season']==2, 'season'] = 'Oct-Nov'
#limiteddf.loc[limiteddf['season']==3, 'season'] = 'Dec'
#Creating dummy variables for the various seasons
seasonlist = limiteddf.season.unique()
for season in seasonlist:
limiteddf[season] = limiteddf['season']==season
# Invoking a procedure similar to get_mpaa in order to process the MPAA rating
for i in limiteddf.index:
try:
limiteddf.loc[i, 'mpaa_new'] = limiteddf.loc[i, 'mpaa'].split("Rated ", 1)[1].split(" ")[0]
except:
limiteddf.loc[i, 'mpaa_new'] = 'PG-13'
limiteddf.loc[limiteddf['mpaa_new']=='PG-', 'mpaa_new'] = 'PG'
limiteddf.loc[limiteddf['mpaa_new']=='NC-17', 'mpaa_new'] = 'R'
#Creating dummy variables for the various MPAA Ratings
mpaalist = limiteddf.mpaa_new.unique()
for mpaa in mpaalist:
limiteddf[mpaa] = limiteddf['mpaa_new']==mpaa
#Creating a list of prolific studios
studiodf = limiteddf.groupby('studio')
studioslist = studiodf['title'].count()
studioslist.sort(ascending=False)
#Identifying the top-5 studios
limiteddf['prol_studio'] = False
for i in studioslist.index[:5]:
limiteddf.loc[limiteddf['studio']==i,'prol_studio'] = True
#Identifying the next 5 top studios
limiteddf['Tier_2'] = False
for i in studioslist.index[6:12]:
limiteddf.loc[limiteddf['studio']==i,'Tier_2'] = True
#Renaming the columns for use later
limiteddf.rename(columns={'opening theaters': 'opening_theaters', 'opening': 'opening_gross'}, inplace=True)
# -
# ##4. Exploratory Data Analysis
##This first plot examines the relationship between Gross Revenue and Opening Weekend Revenue
sns.regplot(y="gross", x="opening_gross", data=limiteddf, fit_reg = True)
# +
# The next plot examines the relationship between the gross income and # of opening theaters
sns.regplot(y="gross", x="opening_theaters", data=limiteddf, fit_reg = True)
# The plot indicates that the relationship is positive, but definitely not linear.
# It appears to have a sharp upward curve at around 3000 theaters, which is an important
# finding for movie houses to consider
# +
#This plot looks at the relationship between a movie's gross revenue and its allocated budget
sns.regplot(y="gross", x="budget", data=limiteddf, fit_reg = True)
# The relationship looks like, although there appears to be a wide disperion, especially
# the further one goes out in revenue/budget
# +
# The next plot looks at how the gross revenue income increases with IMDb rating
sns.regplot(y="gross", x="rating", data=limiteddf, fit_reg = True)
# While there is definitely an upward trend, it is a very flat, indicating that the revenue
# is not strongly dependent on the rating - and perhaps the quality of the movie therein
# -
# ##5. Modeling Using Statsmodel
#
# Having explored the data to some degree, this section delves into multiple models that look at how the Gross Revenue of a movie might be associated with the factors posited at the start of this notebook.
#
# Given the limited number of features, we believe that using Statsmodel would be the ideal tool to leverage. We do analyze whether there are any differences from the sklearn analysis.
# Creating train and test datasets to leverage later
itrain, itest = train_test_split(xrange(limiteddf.shape[0]), train_size=0.6)
mask=np.ones(limiteddf.shape[0], dtype='int')
mask[itrain]=1
mask[itest]=0
mask = (mask==1)
# +
from statsmodels.formula.api import ols
# The first model leverages only three variables (counting season as a single variable)
m1 = ols('gross ~ budget + opening_theaters + season',limiteddf[mask]).fit()
print m1.summary()
#The R-squared associated with this analysis is about average, with plenty of unexplained variation in the data
# As an inital model, it appears to answer important questions around the role of budget
# -
m2 = ols('gross ~ budget + opening_theaters + opening_gross + season',limiteddf[mask]).fit()
print m2.summary()
#Model 2 adds in the gross revenue for the opening weekend. Once we do so,
# The coefficient on the opening theaters variable is reversed, which means that
# adding in the gross opening weekend has exposed the non-linear relationship
## Additionally, there is likely correlation between the opening weekend gross and theaters
m3 = ols('gross ~ budget + opening_theaters + opening_gross + season + prol_studio + Tier_2',limiteddf[mask]).fit()
print m3.summary()
# Model 3 incorporates whether the movie was part of a big studio
# According to the results, there appears to be little effect
m5 = ols('gross ~ budget + opening_gross + season + mpaa_new + rating + opening_theaters',limiteddf[mask]).fit()
print m5.summary()
#Model 5 (Model 4 was deprecated) appears to have the best results, with the minimum required
# variables.
# +
#We compare the results of our Model 1 to Model 5 by testing the observed vs. Predicted fits.
# The green line is a non-parametric lowess curve that attempts to be a comparison
# to help us determine how well the line of best fit describes the true nature of the data.
model1fit = zip(limiteddf[mask]['gross'],m1.fittedvalues)
m1df = pd.DataFrame(model1fit, columns=['observed gross','predicted gross'])
sns.regplot(y="predicted gross", x="observed gross", data=m1df, fit_reg = True, robust=True)
sns.regplot(y="predicted gross", x="observed gross", data=m1df, fit_reg = True, lowess=True)
plt.scatter(limiteddf[mask]['gross'], m1.fittedvalues)
plt.xlabel("Observed Gross: $Y_i$")
plt.ylabel("Predicted Gross: $\hat{Y}_i$")
plt.title("Observed Gross vs Predicted Gross: $Y_i$ vs $\hat{Y}_i$ \n Model 1: Budget, # Opening Theaters, Season")
# -
#Our residuals plot indicates that there is a strong 'fan' effect of heteroskedasticity
# where the error terms increase as we move further out
plt.scatter(m1.fittedvalues, limiteddf[mask]['gross'] - m1.fittedvalues)
plt.xlabel("Fitted Values")
plt.ylabel("Residuals")
plt.title("Residuals vs. Fitted values \n Model 1: Budget, # Opening Theaters, Season")
# +
#Model 5 appears to be a much better fit , where we are explaining the variation far better
model5fit = zip(limiteddf[mask]['gross'],m5.fittedvalues)
m5df = pd.DataFrame(model5fit, columns=['observed gross','predicted gross'])
sns.regplot(y="predicted gross", x="observed gross", data=m5df, fit_reg = True, robust=True)
sns.regplot(y="predicted gross", x="observed gross", data=m5df, fit_reg = True, lowess=True)
plt.scatter(limiteddf[mask]['gross'], m5.fittedvalues)
plt.xlabel("Gross: $Y_i$")
plt.ylabel("Predicted Gross: $\hat{Y}_i$")
plt.title("Gross vs Predicted Gross: $Y_i$ vs $\hat{Y}_i$ \n Model 5: Budget, Opening Weekend, Season, \n # of Opening Theaters, Age Rating and IMDb Rating")
# -
# The residuals have shrunk considerably in Model 5.
# There is also far less exhibited heteroskedasticity in the data.
plt.scatter(m5.fittedvalues, limiteddf[mask]['gross'] - m5.fittedvalues)
plt.xlabel("Fitted Values")
plt.ylabel("Residuals")
plt.title("Residuals vs. Fitted values \n Model 5: Budget, Opening Weekend, Season, \n # of Opening Theaters, Age Rating and IMDb Rating")
# ##5. Using SKlearn as an analysis alternative
# +
from sklearn.linear_model import LinearRegression
# Our X design matrix contains all the covariates that we believe might be interesting to explore
X = limiteddf[['Jan-May','June-Sep','Dec', 'budget', 'opening_gross', 'opening_theaters', 'PG-13', 'R', 'rating']]
# This creates a LinearRegression object
lm = LinearRegression()
# -
#We fit the outcome gross revenue variable on this model
lm.fit(X, limiteddf.gross)
print 'Estimated intercept coefficient:', lm.intercept_
print 'Number of coefficients:', len(lm.coef_)
# The coefficients
pd.DataFrame(zip(X.columns, lm.coef_), columns = ['features', 'estimatedCoefficients'])
# ###Creating a train/test set to rework lm and calculate MSE
## Creating a set of training and test data
X_train, X_test, Y_train, Y_test = sklearn.cross_validation.train_test_split(
X, limiteddf.gross, test_size=0.33, random_state = 5)
print X_train.shape
print X_test.shape
print Y_train.shape
print Y_test.shape
#fitting our training data to the model specified above.
lm = LinearRegression()
lm.fit(X_train, Y_train)
pred_train = lm.predict(X_train)
pred_test = lm.predict(X_test)
## Considering the MSE of our model on the fit versus train data
print "Fit a model X_train, and calculate MSE with Y_train:", np.mean((Y_train - lm.predict(X_train)) ** 2)
print "Fit a model X_train, and calculate MSE with X_test, Y_test:", np.mean((Y_test - lm.predict(X_test)) ** 2)
## The results do indicate that there is some difference between them, but we will
## officially check out models in the section below
##Plotting the results of our model against the residuals
plt.scatter(lm.predict(X_train), lm.predict(X_train) - Y_train, c='b', s=40, alpha=0.5)
plt.scatter(lm.predict(X_test), lm.predict(X_test) - Y_test, c='g', s=40)
plt.hlines(y = 0, xmin=0, xmax = 50)
plt.title('Residual Plot using training (blue) and test (green) data')
plt.ylabel('Residuals')
## Aside from the strong clustering of data points towards the left (lower)
## end of the spectrum, there generally appears to be equally distributed residuals
# ##5b. Using Sklearn to analyze models of increasing complexity while continuing to tune parameters
# +
## We have altered the CV_optimize functions and the do_classify functions to optimize
## the models for our analysis
## Here do_classify is modified to be 'do_regression', but performs similar actions.
## The outcome variable has been defined for us already as Gross Revenue
## The score function employed here is Mean Squared Error, as that is commonly used in
## linear regression models
## There are limited parameters for linear regression, and we will therefore mostly
## be employing a Lasso regression technique, which will eliminate any parameters
## that have negligible effects on the Gross Revenue outcome
def cv_optimize(regmodel, parameters, X, y, n_jobs=1, n_folds=5, score_func=None):
if score_func:
gs = GridSearchCV(regmodel, param_grid=parameters, cv=n_folds, n_jobs=n_jobs, scoring=score_func)
else:
gs = GridSearchCV(regmodel, param_grid=parameters, n_jobs=n_jobs, cv=n_folds)
gs.fit(X, y)
best = gs.best_estimator_
##Note we do not print all possible iterations of the best estimator
return best
def do_regression(regmodel, parameters, indf, featurenames, mask=None,
reuse_split=None, score_func=None, n_folds=5, n_jobs=1):
subdf=indf[featurenames]
X=subdf.values
y=indf['gross']
print "The features being evaluated in this model are:"
print featurenames
if mask !=None:
print "using mask"
Xtrain, Xtest, ytrain, ytest = X[mask], X[~mask], y[mask], y[~mask]
if reuse_split !=None:
print "using reuse split"
Xtrain, Xtest, ytrain, ytest = reuse_split['Xtrain'], reuse_split['Xtest'], reuse_split['ytrain'], reuse_split['ytest']
if parameters:
regmodel = cv_optimize(regmodel, parameters, Xtrain, ytrain, n_jobs=n_jobs, n_folds=n_folds, score_func=score_func)
regmodel=regmodel.fit(Xtrain, ytrain)
training_accuracy = regmodel.score(Xtrain, ytrain)
test_accuracy = regmodel.score(Xtest, ytest)
print "############# based on standard predict ################"
print "Accuracy on training data: %0.2f" % (training_accuracy)
print "Accuracy on test data: %0.2f" % (test_accuracy)
print "########################################################"
return regmodel, Xtrain, ytrain, Xtest, ytest
# +
##Using a budget-only model as the baseline
reg_1 = Lasso()
# The parameters being tuned are those specific to Lasso, in particular we expect only the
# Tolerance parameter to have any effect on the outcomes
parameters = {"fit_intercept": [True, False], "normalize":[True, False],
"copy_X":[True, False], "max_iter":[10,100,1000],
"tol": [0.0001,0.001, 0.00001,0.01]}
reg_1, Xtrain, ytrain, Xtest, ytest=do_regression(reg_1, parameters, limiteddf, ['budget'], mask=mask, score_func = 'r2')
# +
##Incorporating the # of Openings theaters as an additional predictor
reg_2 = Lasso()
parameters = {"fit_intercept": [True, False], "normalize":[True, False], "copy_X":[True, False], "max_iter":[10,100,1000],
"tol": [0.0001,0.001, 0.00001,0.01]}
reg_2, Xtrain, ytrain, Xtest, ytest=do_regression(reg_2, parameters, limiteddf, ['budget', 'opening_theaters'], mask=mask, score_func = 'mean_squared_error')
# +
##Incorporating Opening Weekend Gross
reg_3 = Lasso()
parameters = {"fit_intercept": [True, False], "normalize":[True, False], "copy_X":[True, False], "max_iter":[10,100,1000],
"tol": [0.0001,0.001, 0.00001,0.01]}
reg_3, Xtrain, ytrain, Xtest, ytest=do_regression(reg_3, parameters, limiteddf, ['budget', 'opening_theaters', 'opening_gross'], mask=mask, score_func = 'mean_squared_error')
# +
#This version incorporates the season that the movie was released in
reg_4 = Lasso()
parameters = {"fit_intercept": [True, False], "normalize":[True, False], "copy_X":[True, False], "max_iter":[10,100,1000],
"tol": [0.0001,0.001, 0.01]}
reg_4, Xtrain, ytrain, Xtest, ytest=do_regression(reg_4, parameters, limiteddf, ['Jan-May', 'June-Sep', 'Dec', 'budget', 'opening_gross', 'opening_theaters'], mask=mask, score_func = 'mean_squared_error')
# +
# This includes the studio, in the form of 'prol_studio' which measures if the studio is
# in the top 5 prolific studios in the country
reg_5 = Lasso()
parameters = {"fit_intercept": [True, False], "normalize":[True, False], "copy_X":[True, False], "max_iter":[10,100,1000],
"tol": [0.0001,0.001, 0.01]}
reg_5, Xtrain, ytrain, Xtest, ytest=do_regression(reg_5, parameters, limiteddf, ['prol_studio', 'Jan-May', 'June-Sep', 'Dec', 'budget', 'opening_gross', 'opening_theaters'], mask=mask, score_func = 'mean_squared_error')
# +
#With Tier_2 studio added in there to see if the movie belonging to any
# of the top 10 studios has an effect
reg_6 = Lasso()
parameters = {"fit_intercept": [True, False], "normalize":[True, False], "copy_X":[True, False], "max_iter":[10,100,1000],
"tol": [0.0001,0.001, 0.01]}
reg_6, Xtrain, ytrain, Xtest, ytest=do_regression(reg_6, parameters, limiteddf, ['prol_studio','Tier_2', 'Jan-May', 'June-Sep', 'Oct-Nov', 'budget', 'opening_gross', 'opening_theaters'], mask=mask, score_func = 'mean_squared_error')
# +
#With MPAA Rating included in the model
reg_7 = Lasso()
parameters = {"fit_intercept": [True, False], "normalize":[True, False], "copy_X":[True, False], "max_iter":[10,100,1000],
"tol": [0.0001,0.001, 0.01]}
reg_7, Xtrain, ytrain, Xtest, ytest=do_regression(reg_7, parameters, limiteddf, [ 'Jan-May', 'June-Sep', 'Oct-Nov', 'budget', 'opening_gross', 'opening_theaters','prol_studio','Tier_2', 'PG-13', 'R'], mask=mask, score_func = 'mean_squared_error')
# +
#With IMDb Movie Rating included
reg_8 = Lasso()
parameters = {"fit_intercept": [True, False], "normalize":[True, False], "copy_X":[True, False], "max_iter":[10,100,1000],
"tol": [0.0001,0.001, 0.01]}
reg_8, Xtrain, ytrain, Xtest, ytest=do_regression(reg_8, parameters, limiteddf, [ 'Jan-May', 'June-Sep', 'Oct-Nov', 'budget', 'opening_gross', 'opening_theaters', 'PG-13', 'R', 'rating'], mask=mask, score_func = 'mean_squared_error')
# -
## Having attained the model we believe to be the best fit of our data, we can
## compare the output to that received from the Statsmodel analysis
from sklearn.feature_selection import chi2
scores, pvalues = chi2(Xtrain, ytrain)
## This compiles the results from the sklearn variable
print 'Estimated intercept coefficient:', reg_8.intercept_
print 'Number of coefficients:', len(reg_8.coef_)
# The coefficients
print reg_8.intercept_
pd.DataFrame(zip(['Jan-May', 'June-Sep', 'Oct-Nov', 'budget', 'opening_gross', 'opening_theaters', 'PG-13','R', 'rating'], reg_8.coef_, scores, pvalues), columns = ['features', 'estimatedCoefficients', 'scores', 'pvalues'])
#Reporting the results from our last Statsmodel version, m5.
m5.summary()
# #Results and Conclusions
# The series of analyses conducted over the course of this section of the project have yielded interesting results, and we discuss the ramifications of those outputs here:
#
# - The 'budget' variable is both statistically signficant and meaningful. The coefficient of about 0.3 indicates that every dollar in the budget entails 30 cents of revenue. This means that simply throwing money at a movie in the hopes that it works is not a wise strategy, and the other factors in the model are very important in ensuring that a movie does financially well.
#
# - The coefficients on all the season variables indicates are negative, which means that the holdout month - December - appears to be associated with the best results for a movie's release, with movies being released during that month having an average 20M dollar boost compared to other seasons.
#
# - The movie's MPAA rating also seems to matter, with both PG-13 and R-rated movies performing worse than their G-rated counterparts. The magnitude of the difference, controlling for all other factors, is about 18M dollars.
#
# - The gross during the opening weekend is highly associated with the amount that the movie ends up making, with a coefficient of 2.7 this would emphasize the need to start off with a bang.
#
# - The opening theaters variable is only moderately statistically significant, and it would actually decrease the gross revenue if there were more theaters. We do not believe this variable to be accurate, not only because of the significance, but also because it likely exhibits multicollinearity with other covariates in the model.
#
# - The IMDb rating of the movie has a distinct, but muted, effect on the fiscal outcomes of a movie. According to the results, increasing an entire rating-point on the IMDb scale is associated with only a 10 million dollar increase in revenue. Inasmuch as our rating variable captures the 'true' quality of a movie, that is not a particularly encouraging sign for where the focus of the movie production houses lie.
#
| box_office_process_notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
import os
import torch as t
from utils.config import opt
from model import FasterRCNNVGG16
from trainer import FasterRCNNTrainer
from data.util import read_image
from utils.vis_tool import vis_bbox
from utils import array_tool as at
import cv2
import numpy
from numpy import array
from labels import *
# %matplotlib inline
faster_rcnn = FasterRCNNVGG16()
trainer = FasterRCNNTrainer(faster_rcnn).cuda()
#
# in this machine the cupy isn't install correctly...
# so it's a little slow
trainer.load('/home/ml/Downloads/chainer_best_model_converted_to_pytorch_0.7053.pth')
opt.caffe_pretrain=True # this model was trained from caffe-pretrained model
# +
cap = cv2.VideoCapture("test.mp4")
print ("capture starts")
no_p=0
count = 0
list=['bike','bus','car','motorbike']
while (cap.isOpened()):
ret,image_np = cap.read()
if count%20 == 0 :
no_p=no_p+1
image_np = cv2.resize(image_np, (720,360))
image_cv=image_np
image_np = image_np.transpose((2, 0, 1))
img = t.from_numpy(image_np)[None]
_bboxes, _labels, _scores = trainer.faster_rcnn.predict(img,visualize=True)
X = numpy.array(_labels)
Y = numpy.array(_bboxes)
Y=Y.astype(int)
i=0
j=0
x1=[]
y1=[]
x2=[]
y2=[]
for j in range (X.shape[1]):
x1=(Y[0][i][1])
y1=(Y[0][i][0])
x2=(Y[0][i][3])
y2=(Y[0][i][2])
for i in range (X.shape[1]):
if LABEL_NAMES[X[0,i]] in list:
print ('found a vehicle!')
print (LABEL_NAMES[X[0,i]])
cv2.rectangle(image_cv,(x1,y1),(x2,y2),(0,255,0),2)
cv2.putText(image_cv,LABEL_NAMES[X[0,i]],(x1,y1),cv2.FONT_HERSHEY_SIMPLEX,2,(255,0,0))
cv2.imshow("Show",image_cv)
print (no_p)
if cv2.waitKey(25) & 0xFF == ord('q'):
cv2.destroyAllWindows()
cap.release()
break
count+=1
# -
| vehicle-detection-in-video.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <table> <tr>
# <td style="background-color:#ffffff;">
# <a href="http://qworld.lu.lv" target="_blank"><img src="..\images\qworld.jpg" width="25%" align="left"> </a></td>
# <td style="background-color:#ffffff;vertical-align:bottom;text-align:right;">
# prepared by <a href="http://abu.lu.lv" target="_blank"><NAME></a> (<a href="http://qworld.lu.lv/index.php/qlatvia/" target="_blank">QLatvia</a>)
# </td>
# </tr></table>
# <table width="100%"><tr><td style="color:#bbbbbb;background-color:#ffffff;font-size:11px;font-style:italic;text-align:right;">This cell contains some macros. If there is a problem with displaying mathematical formulas, please run this cell to load these macros. </td></tr></table>
# $ \newcommand{\bra}[1]{\langle #1|} $
# $ \newcommand{\ket}[1]{|#1\rangle} $
# $ \newcommand{\braket}[2]{\langle #1|#2\rangle} $
# $ \newcommand{\dot}[2]{ #1 \cdot #2} $
# $ \newcommand{\biginner}[2]{\left\langle #1,#2\right\rangle} $
# $ \newcommand{\mymatrix}[2]{\left( \begin{array}{#1} #2\end{array} \right)} $
# $ \newcommand{\myvector}[1]{\mymatrix{c}{#1}} $
# $ \newcommand{\myrvector}[1]{\mymatrix{r}{#1}} $
# $ \newcommand{\mypar}[1]{\left( #1 \right)} $
# $ \newcommand{\mybigpar}[1]{ \Big( #1 \Big)} $
# $ \newcommand{\sqrttwo}{\frac{1}{\sqrt{2}}} $
# $ \newcommand{\dsqrttwo}{\dfrac{1}{\sqrt{2}}} $
# $ \newcommand{\onehalf}{\frac{1}{2}} $
# $ \newcommand{\donehalf}{\dfrac{1}{2}} $
# $ \newcommand{\hadamard}{ \mymatrix{rr}{ \sqrttwo & \sqrttwo \\ \sqrttwo & -\sqrttwo }} $
# $ \newcommand{\vzero}{\myvector{1\\0}} $
# $ \newcommand{\vone}{\myvector{0\\1}} $
# $ \newcommand{\stateplus}{\myvector{ \sqrttwo \\ \sqrttwo } } $
# $ \newcommand{\stateminus}{ \myrvector{ \sqrttwo \\ -\sqrttwo } } $
# $ \newcommand{\myarray}[2]{ \begin{array}{#1}#2\end{array}} $
# $ \newcommand{\X}{ \mymatrix{cc}{0 & 1 \\ 1 & 0} } $
# $ \newcommand{\I}{ \mymatrix{rr}{1 & 0 \\ 0 & 1} } $
# $ \newcommand{\Z}{ \mymatrix{rr}{1 & 0 \\ 0 & -1} } $
# $ \newcommand{\Htwo}{ \mymatrix{rrrr}{ \frac{1}{2} & \frac{1}{2} & \frac{1}{2} & \frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & \frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} & \frac{1}{2} } } $
# $ \newcommand{\CNOT}{ \mymatrix{cccc}{1 & 0 & 0 & 0 \\ 0 & 1 & 0 & 0 \\ 0 & 0 & 0 & 1 \\ 0 & 0 & 1 & 0} } $
# $ \newcommand{\norm}[1]{ \left\lVert #1 \right\rVert } $
# $ \newcommand{\pstate}[1]{ \lceil \mspace{-1mu} #1 \mspace{-1.5mu} \rfloor } $
# $ \newcommand{\greenbit}[1] {\mathbf{{\color{green}#1}}} $
# $ \newcommand{\bluebit}[1] {\mathbf{{\color{blue}#1}}} $
# $ \newcommand{\redbit}[1] {\mathbf{{\color{red}#1}}} $
# $ \newcommand{\brownbit}[1] {\mathbf{{\color{brown}#1}}} $
# <h2> Основы Python: Условия </h2>
#
# Мы рассмотрим использование условий в Python.
#
# Пожалуйста, запустите каждую ячейку и проверьте результаты.
# +
# let's randomly pick a number between 0 and 9, and print its value if it is greater than 5
from random import randrange
r = randrange(10)
if r > 5:
print(r)
else:
print(-r)
# when the condition (r > 5) is valid/true, the code (print(r)) will be executed
# you may need to execute your code more than once to see an outcome
# +
# repeat the same task four times, and also print the value of iteration variable (i)
for i in range(4):
r = randrange(10) # this code belongs to for-loop, and so it is indented
if r > 5: # this code also belongs to for-loop, and so it is indented as well
print("i =",i,"r =",r) # this code belongs to if-statement, and so it is indented with respect to if-statement
# if you are unlucky (with probabability less than 13/100), you may not see any outcome after a single run
# +
# do the same task 100 times, and find the percentage of successful iterations (attempts)
# an iteration (attempt) is successful if the randomly picked number is greater than 5
# the expected percentage is 40, because, out of 10 numbers, there are 4 numbers greater than 5
# but the experimental results differ
success = 0
for i in range(100):
r = randrange(10)
if r > 5:
success = success + 1
print(success,"%")
# each experiment most probably will give different percentage value
# +
# let's randomly pick a number between 0 and 9, and print whether it is less than 6 or not
# we use two conditionals here
r = randrange(10)
print("the picked number is ",r)
if r < 6:
print("it is less than 6")
if r >= 6:
print("it is greater than or equal to 6")
# +
# let's write the same algorithm by using if-else structure
r = randrange(10)
print("the picked number is ",r)
if r < 6:
print("it is less than 6")
else: # if the above condition (r<6) is False
print("it is greater than or equal to 6")
# -
# <h3> Task 1 </h3>
#
# Произвольно (Рандомно) выберите число в интервале между 10 и 50 и выведите четность/нечетность полученного числа.
#
# Напомним, что оператор получения остатка от деления (mod) в Python обозначается знаком %.
#
# Число является четным, если делится на 2 без остатка.
#
# Число является нечетным, если при делении числа на 2 получается остаток, равный 1.
#
# your solution is here
#
# <a href="Python16_Basics_Conditionals_Solutions.ipynb#task1">click for our solution</a>
# +
#
# when there are many related conditionals, we can use if-elif-else structure
#
# let's randomly pick an even number between 1 and 99
# then determine whether it is less than 25, between 25 and 50, between 51 and 75, or greater than 75.
r = randrange(2,100,2) # randonmly pick a number in range {2,4,6,...,98}, which satisfies our condition
# let's print this range to verify our claim
print(list(range(2,100,2)))
print() # print an empty line
print("the picked number is",r)
if r < 25:
print("it is less than 25")
elif r<=50: # if the above condition is False and the condition here is True
print("it is between 25 and 50")
elif r<=75: # if both conditions above are False and the condition here is True
print("it is between 51 and 75")
else: # if none of the above conditions is True
print("it is greater than 75")
# -
# <h3> Task 2 </h3>
#
# Произвольно (Рандомно) выберите число в интервале от 0 до 99.
#
# С одинаковой вероятностью $\left( 0.5 = \dfrac{1}{2} \right)$, число окажется из интервала от 0 до 49 или от 50 до 99.
#
# "Найдите эту вероятность экспериментально".
#
# N=100 раз произвольно выберите число в интервале от 0 до 99, вычислите частоты в обоих случаях, далее каждый поделите на 100.
#
# Попробуйте повторить эксперимент для N=1,000, N=10,000, и N=100,000.
#
# Экспериментальные результаты будут ближе к идеальному соотношению при увеличении N.
#
# your solution is here
#
# <a href="Python16_Basics_Conditionals_Solutions.ipynb#task2">click for our solution</a>
# <h3> Простые числа </h3>
#
# Число простое, если оно больше 1 и делится без остатка только на 1 и на само себя.
#
# Например, любое отрицательное число, 0, или 1 не являются простыми.
#
# 2, 3, 5, 7 и 11 - первые пять простых чисел.
#
# 31 - еще одно простое число, так как оно не делится без остатка на любое другое число, кроме 1 и 31.
# <br>
# Иначе говоря, любое число в отрезке {2,3,4,...,30} не делит 31 без остатка.
#
# 4 - не простое число, так как оно также делится на 2 без остатка.
# <br>
# 9 - не простое число, так как оно также делится на 3 без остатка.
# +
# let's determine whether a randomly picked number between -10 and 100 is prime or not.
# this is a good example for using more than one conditional in different parts of the program
# this is also an example for "break" command, which terminates any loop immediately
r = randrange(-10,101) # pick a number between -10 and 100
print(r) # print is value
if r < 2: print("it is NOT a prime number") # this is by definition
elif r == 2: print("it is a PRIME number") # we already know this
else:
prime=True # we assume that r is prime, and try to falsify this assumption by looking for a divisor in the following loop
for i in range(2,r): # check all integers between 2 and r-1
if r % i ==0: # if i divides r without any reminder (or reminder is zero), then r is not a prime number
print("it is NOT a prime number")
prime=False # our assumption is falsifed
break # TERMINATE the iteration immediately
# we are out of if-scope
# we are out of for-loop-scope
if prime == True: # if our assumption is still True (if it was not falsified inside for-loop)
print("it is a PRIME number")
# -
# <h3> Написание функций в Python </h3>
#
# Наш код определяет, является ли целое число, переданное в качестве аргумента, простым или нет.
#
# Мы можем сохранить эту функцию в отдельном модуле и использовать по мере необходимости.
# +
# this is an example to write a function
# our function will return a Boolean value True or False
def prime(number): # our function takes one parameter (has one argument)
if number < 2: return False # once return command is executed, we exit the function
if number == 2: return True # because of return command, we can use "if" instead of "elif"
if number % 2 == 0: return False # any even number greater than 2 is not prime, because it is divisible by 2
for i in range(3,number,2): # we can skip even integers
if number % i == 0: return False # once we find a divisor of the number, we return False and exit the function
return True # the number has passed all checks until now
# by using "return" command appropriately, the programs can be shortened
# remark that this might not be a good choice everytime for readibility of codes
# let's test our program by printing all prime numbers between -10 and 30
for i in range(-10,30):
# we pass i to the function prime
if prime(i): # the function prime(i) returns True or False
print(i) # this code will be executed if i is prime, i.e., prime(i) returns True
# -
| python/Python16_Basics_Conditionals.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Comprehensions
# ### The list comprehension
# If you write a for loop **inside** a pair of square brackets for a list, you magic up a list as defined.
# This can make for concise but hard to read code, so be careful.
[2 ** x for x in range(10)]
# Which is equivalent to the following code without using comprehensions:
# +
result = []
for x in range(10):
result.append(2 ** x)
result
# -
# You can do quite weird and cool things with comprehensions:
[len(str(2 ** x)) for x in range(10)]
# ### Selection in comprehensions
# You can write an `if` statement in comprehensions too:
[2 ** x for x in range(30) if x % 3 == 0]
# Consider the following, and make sure you understand why it works:
"".join([letter for letter in "<NAME>"
if letter.lower() not in 'aeiou'])
# ### Comprehensions versus building lists with `append`:
# This code:
result = []
for x in range(30):
if x % 3 == 0:
result.append(2 ** x)
result
# Does the same as the comprehension above. The comprehension is generally considered more readable.
# Comprehensions are therefore an example of what we call 'syntactic sugar': they do not increase the capabilities of the language.
# Instead, they make it possible to write the same thing in a more readable way.
# Almost everything we learn from now on will be either syntactic sugar or interaction with something other than idealised memory, such as a storage device or the internet. Once you have variables, conditionality, and branching, your language can do anything. (And this can be proved.)
# ### Nested comprehensions
# If you write two `for` statements in a comprehension, you get a single array generated over all the pairs:
[x - y for x in range(4) for y in range(4)]
# You can select on either, or on some combination:
[x - y for x in range(4) for y in range(4) if x >= y]
# If you want something more like a matrix, you need to do *two nested* comprehensions!
[[x - y for x in range(4)] for y in range(4)]
# Note the subtly different square brackets.
# Note that the list order for multiple or nested comprehensions can be confusing:
[x+y for x in ['a', 'b', 'c'] for y in ['1', '2', '3']]
[[x+y for x in ['a', 'b', 'c']] for y in ['1', '2', '3']]
# ### Dictionary Comprehensions
# You can automatically build dictionaries, by using a list comprehension syntax, but with curly brackets and a colon:
{(str(x)) * 3: x for x in range(3)}
# ### List-based thinking
# Once you start to get comfortable with comprehensions, you find yourself working with containers, nested groups of lists
# and dictionaries, as the 'things' in your program, not individual variables.
# Given a way to analyse some dataset, we'll find ourselves writing stuff like:
#
# analysed_data = [analyze(datum) for datum in data]
# There are lots of built-in methods that provide actions on lists as a whole:
any([True, False, True])
all([True, False, True])
max([1, 2, 3])
sum([1, 2, 3])
# My favourite is `map`, which, similar to a list comprehension, applies one function to every member of a list:
[str(x) for x in range(10)]
list(map(str, range(10)))
# So I can write:
#
# analysed_data = map(analyse, data)
#
# We'll learn more about `map` and similar functions when we discuss functional programming later in the course.
# ### Classroom Exercise: Occupancy Dictionary
# Take your maze data structure. First write an expression to print out a new dictionary, which holds, for each room, that room's capacity. The output should look like:
{'bedroom': 1, 'garden': 3, 'kitchen': 1, 'living': 2}
# Now, write a program to print out a new dictionary, which gives,
# for each room's name, the number of people in it. Don't add in a zero value in the dictionary for empty rooms.
# The output should look similar to:
{'garden': 1, 'living': 1}
| ch00python/037comprehensions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
#import matplotlib.pyplot as plt
#import seaborn as sns
import pandas as pd
import numpy as np
import random as rnd
from sklearn.cross_validation import KFold, cross_val_score
# machine learning
from sklearn.linear_model import LogisticRegression
from sklearn.svm import SVC, LinearSVC
from sklearn.ensemble import RandomForestClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.naive_bayes import GaussianNB
from sklearn.linear_model import Perceptron
from sklearn.linear_model import SGDClassifier
from sklearn.tree import DecisionTreeClassifier
train_=pd.read_csv('../train_allcols.csv')
validate_=pd.read_csv('../validate_allcols.csv')
#test=pd.read_csv('../testwDSM.csv')
characters = pd.read_csv('chars.csv', sep=',')
train_.shape, validate_.shape, #test.shape
# -
characters.shape
train = train_.query('DSMCRIT > 13 and DSMCRIT < 19')
validate = validate_.query('DSMCRIT > 13 and DSMCRIT < 19')
#print train['DSMCRIT'].value_counts()
print train.shape
# +
#alcohol
#print train['DSMCRIT'].value_counts() / train['DSMCRIT'].count()
#print train['SUB1'].value_counts() / train['SUB1'].count()
# +
#train.query('SUB1 == 4')['DSMCRIT'].value_counts() / train.query('SUB1 == 4')['DSMCRIT'].count()
# -
#train.describe()
#train = train.sample(20000)
#validate = validate.sample(6000)
train.shape, #validate.shape, #validate.head(2)
# +
#train = train.query('SUB1 <= 10').query('SUB2 <= 10')
#validate = validate.query('SUB1 <= 10').query('SUB2 <= 10')
drop_list = ['DSMCRIT', #'NUMSUBS'
]
drop_list_select = ['RACE', 'PREG', 'ARRESTS', 'PSYPROB', 'DETNLF', 'ETHNIC', 'MARSTAT', 'GENDER', 'EDUC'
,'LIVARAG', 'EMPLOY', 'SUB3']
retain_list = ['RACE','PCPFLG','PRIMINC','LIVARAG','BENZFLG','HLTHINS','GENDER','ROUTE3','PRIMPAY',
'MARSTAT','PSYPROB','ROUTE2','EMPLOY','SUB2','FRSTUSE3','FREQ3','FRSTUSE2','OTHERFLG',
'EDUC','FREQ2','FREQ1','YEAR',
'PSOURCE','DETCRIM','DIVISION','REGION','NOPRIOR','NUMSUBS','ALCDRUG',
'METHUSE','FRSTUSE1','AGE','COKEFLG','OPSYNFLG','IDU','SERVSETA','ROUTE1','MARFLG',
'MTHAMFLG','HERFLG',
'ALCFLG','SUB1']
X_train = train[retain_list]
X_c = characters[retain_list]
Y_c = characters["DSMCRIT"]
#X_train = train.drop(drop_list + drop_list_select, axis=1)
Y_train = train["DSMCRIT"]
#X_validate = validate.drop(drop_list + drop_list_select, axis=1)
#Y_validate = validate["DSMCRIT"]
#X_test = test.drop(drop_list, axis=1)
X_train.shape, #X_validate.shape, #X_test.shape
# +
#one hot
from sklearn import preprocessing
# 1. INSTANTIATE
enc = preprocessing.OneHotEncoder()
# 2. FIT
enc.fit(X_train)
# 3. Transform
onehotlabels = enc.transform(X_train).toarray()
X_train = onehotlabels
#onehotlabels = enc.transform(X_validate).toarray()
#X_validate = onehotlabels
X_char = enc.transform(X_c).toarray()
print X_train.shape, X_char.shape#X_validate.shape
# -
#kfold
kf = 3
# +
# Logistic Regression
logreg = LogisticRegression(n_jobs=-1)
logreg.fit(X_train, Y_train)
#Y_pred = logreg.predict(X_test)
l_acc_log = cross_val_score(logreg, X_train, Y_train, cv=kf)
acc_log = round(np.mean(l_acc_log), 3)
l_acc_log = ['%.3f' % elem for elem in l_acc_log]
print l_acc_log
print acc_log
# -
yp_char = logreg.predict_proba(X_char)
print yp_char
#print Y_c
# +
# Random Forest (slow)
random_forest = RandomForestClassifier(n_estimators=200, max_depth=20, n_jobs=-1)
random_forest.fit(X_train, Y_train)
#Y_pred = random_forest.predict(X_test)
l_acc_random_forest = cross_val_score(random_forest, X_train, Y_train, cv=kf)
acc_random_forest = round(np.mean(l_acc_random_forest), 3)
l_acc_random_forest = ['%.3f' % elem for elem in l_acc_random_forest]
print l_acc_random_forest
print acc_random_forest
# -
yp_char = random_forest.predict_proba(X_char)
print yp_char
#print Y_c
| week9/mental-dsm-predict_prob.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1 - Convert Data from VED
# This short notebook converts the Vehicle Energy Dataset from the original multi-file CSV format to a more convenient single file parquet format. Parquet files do not require parsing, as CSV files do, so they are inherently faster to load.
#
# Start by downloading the data from https://github.com/gsoh/VED into the `data` folder (please create it if it is not there). After expanding all the CSV files, please run the code below.
#
# Note: Please install the `pyarrow` package before running this notebook.
import numpy as np
import pandas as pd
import os
# Set the data path and target file name.
data_path = "./data"
parquet_file = os.path.join(data_path, "ved.parquet")
# The `read_data_frame` function reads a single VED CSV file into its own DataFrame object. It is meant to be used with the `map` function in a comprehension expression (see below).
def read_data_frame(filename):
columns = ['DayNum', 'VehId', 'Trip', 'Timestamp(ms)', 'Latitude[deg]', 'Longitude[deg]',
'Vehicle Speed[km/h]']
types = {'VehId': np.int64,
'Trip': np.int64,
'Timestamp(ms)': np.int64}
df = pd.read_csv(filename, usecols=columns, dtype=types)
return df
# Read all the files into the same DataFrame and dump it into a single parquet file.
files = [os.path.join(data_path, file) for file in os.listdir(data_path) if file.endswith(".csv")]
df = pd.concat(map(read_data_frame, files), ignore_index=True)
df = df.sort_values(by=['VehId', 'DayNum', 'Timestamp(ms)'])
df.to_parquet(parquet_file)
| 1-convert-ved.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from IPython.display import Image
import requests
import base64
from snet_cli.sdk import Session, Client, AutoFundingFundingStrategy
from snet_cli.config import Config
# Load config from ~/.snet/config
session = Session(Config())
# Create a client
client = Client(session, "snet", "siggraph-colorization", AutoFundingFundingStrategy(amount_cogs=1, expiration="+10days"))
# Checking account address
client.ident.get_address()
# -
# Checking the input Image
r = requests.get("https://snet-models.s3.amazonaws.com/bh/Colorize/carnaval.jpg")
file_name = "carnaval.jpg"
with open(file_name, "wb") as fd:
fd.write(r.content)
Image(filename='carnaval.jpg')
# Create a request
request = client.classes.Input(img_input="https://snet-models.s3.amazonaws.com/bh/Colorize/carnaval.jpg")
# Get the response
response = client.stub.colorize(request)
# +
# Convert base64 bytes to JPG image
img_data = base64.b64decode(response.img_colorized)
file_name = "carnaval_colorized.jpg"
with open(file_name, "wb") as fd:
fd.write(img_data)
# Checking the image that was returned from service
Image(filename='carnaval_colorized.jpg')
# -
| Let_there_be_Color.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
import os
os.environ["CUDA_DEVICE_ORDER"]="PCI_BUS_ID";
os.environ["CUDA_VISIBLE_DEVICES"]="0";
import ktrain
from ktrain import vision as vis
from ktrain import text as txt
# # Additional Features
#
# In this notebook, we will demonstrated some additional features of *ktrain*. To demonstrate these features, we will first load some sample data and define a simple model that trains fast. As usual, we will wrap the model and data in a *ktrain* Learner object.
# load and prepare data as you normally would in Keras
from keras.preprocessing import sequence
from keras.datasets import imdb
NUM_WORDS = 20000
MAXLEN = 400
def load_data():
(x_train, y_train), (x_test, y_test) = imdb.load_data(num_words=NUM_WORDS)
x_train = sequence.pad_sequences(x_train, maxlen=MAXLEN)
x_test = sequence.pad_sequences(x_test, maxlen=MAXLEN)
return (x_train, y_train), (x_test, y_test)
(x_train, y_train), (x_test, y_test) = load_data()
# build a model as you normally would in Keras
from keras.models import Sequential
from keras.layers import Dense, Embedding, GlobalAveragePooling1D
def get_model():
model = Sequential()
model.add(Embedding(NUM_WORDS, 50, input_length=MAXLEN))
model.add(GlobalAveragePooling1D())
model.add(Dense(1, activation='sigmoid'))
model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
return model
model = get_model()
learner = ktrain.get_learner(model, train_data=(x_train, y_train), val_data = (x_test, y_test))
# ## Debugging Keras Models
#
# When debugging neural networks, it is sometimes useful to [inspect the results of intermediate layers](https://stackoverflow.com/questions/42233963/debugging-keras-intermediate-layer-or-objective-variables-with-tensorflow). *ktrain* makes this easy with the ```print_layers``` and ```layer_output``` methods. The ```print_layers``` method prints the layers of your network with each layer being assigned an index. The output of ```print_layers``` also shows which layers are trainable (unfrozen) and untrainable (frozen).
learner.print_layers()
# The ```layer_output``` method accepts a layer ID as input. By default, it will print the output of the layer associated with the layer ID using the first example in the training set. The example_id and use_val arguments can control which example is used to generate the output. In the example below, we print the output of the Dense layer of our network using the sixth example from our validation set (i.e., the example with index 5 in the x_test array).
learner.layer_output(2, example_id=5, use_val=True)
# Since the model has not yet been trained, it returns 0.5.
# Here, we print the output of the GlobalAveragePooling1D layer (layer_id=1) implicitly using the first example from our training set.
learner.layer_output(1)
# ## Setting Global Weight Decay in Keras
#
# [Weight decay](https://machinelearningmastery.com/how-to-reduce-overfitting-in-deep-learning-with-weight-regularization/) is a form of regularization to reduce overfitting. In Keras, this is typically done by [setting the weight decay for individual layers within the network](https://github.com/keras-team/keras/issues/2717). The global weight decay for a neural model can easily be set by the ```learner.set_weight_decay``` method.
#
learner.print_layers(show_wd=True)
learner.set_weight_decay(1e-3)
learner.get_weight_decay()
learner.print_layers(show_wd=True)
learner.autofit(0.005, 6)
# ## Saving the Current Model
#
# When issuing calls to ```autofit```, ```fit_onecycle```, and ```fit```, it is sometimes useful to periodically save the model in case additional training results in overfitting. This can be accomplished with the ```learner.save_model``` and ```learner.load_model``` methods, which are simply wrappers to ```model.save``` and ```load_model``` in Keras. Example usage is shown below:
learner.save_model('/tmp/mymodel')
# train further here
# if overfitting, then do this:
learner.load_model('/tmp/mymodel')
# ## Built-In Callbacks
#
# *ktrain* enables easy access to [EarlyStopping](https://keras.io/callbacks/#earlystopping), [ModelCheckpoint](https://keras.io/callbacks/#modelcheckpoint), and [ReduceLROnPlateau](https://keras.io/callbacks/#reducelronplateau) callbacks. All of the **fit** methods of *ktrain* have both an ```early_stopping``` argument and a ```checkpoint_folder``` argument. When setting ```early_stopping=3```, for instance, the training will disconinue automatically when the validation loss fails to improve after a 3 epochs. When suppying the path to a folder using ```checkpoint_folder``` argument, files containing the weights obtained after each epoch will be saved to the folder. In the example below, we will train our model using the ```fit``` method using both arguments.
learner.reset_weights()
learner.fit(0.005, 10, cycle_len=1, cycle_mult=2, early_stopping=3, checkpoint_folder='/tmp')
# The weights obtained after each of the six epochs will be available as files stored in the checkpoint_folder:
#
# ```
# /tmp/weights-01.hdf5 /tmp/weights-03.hdf5 /tmp/weights-05.hdf5
# /tmp/weights-02.hdf5 /tmp/weights-04.hdf5 /tmp/weights-06.hdf5
# ```
#
# These weights can be easily loaded into the model as one would normally do:
# ```
# learner.model.load_weights('/tmp/weights-02.hdfs')
# ```
#
# The ```checkpoint_folder``` argument can be used with any "fit" method (i.e., ```autofit```, ```fit_onecycle```, and ```fit```). It is particularly useful when needing to rollback to an earlier epoch in case the model overfits.
#
# In addition, the ```autofit``` method includes a **reduce_on_plateau** parameter, which can be used to automatically reduce the maximum (and base) learning rates in the [triangular learning rate policy](https://arxiv.org/abs/1506.01186) when the validation loss no longer improves by a factor specified by the **reduce_factor** argument. The criterion can be changed to validation accuracy using the **monitor** argument. When ```autofit``` is called without an **epochs** parameter, both the **early_stopping** and **reduce_on_plateau** are automatically enabled.
# ## Custom Callbacks
# The ```fit```, ```autofit```, and ```fit_onecycle``` methods of the Learner object can also accept custom callbacks, just as you can in standard calls to ```model.fit```. For instance, here we re-train our model and print the ROC-AUC score after each epoch:
# +
# define a custom callback for ROC-AUC
from keras.callbacks import Callback
from sklearn.metrics import roc_auc_score
class RocAucEvaluation(Callback):
def __init__(self, validation_data=(), interval=1):
super(Callback, self).__init__()
self.interval = interval
self.X_val, self.y_val = validation_data
def on_epoch_end(self, epoch, logs={}):
if epoch % self.interval == 0:
y_pred = self.model.predict(self.X_val, verbose=0)
score = roc_auc_score(self.y_val, y_pred)
print("\n ROC-AUC - epoch: %d - score: %.6f \n" % (epoch+1, score))
RocAuc = RocAucEvaluation(validation_data=(x_test, y_test), interval=1)
# -
# re-create our model from scratch and train using our custom ROC-AUC callback
learner = ktrain.get_learner(get_model(), train_data=(x_train, y_train), val_data = (x_test, y_test))
learner.autofit(0.005, 2, callbacks=[RocAuc])
# ## Using TensorBoard with *ktrain*
#
# Since *ktrain* is a lightweight wrapper around Keras, you can use TensorBoard callbacks with all `fit` methods in *ktrain* (e.g., `autofit`, `fit_onecycyle`, `fit`) just as you would normally do with `model.fit` in Keras:
# +
learner.reset_weights()
import keras
# create TensorBoard calback
tbCallBack = keras.callbacks.TensorBoard(log_dir='/tmp/Graph', histogram_freq=0, write_graph=True, write_images=True)
# supply it as custom callback to any fit method of Learner
learner.autofit(0.005, 2, callbacks=[tbCallBack])
# -
# When training is complete, TensorBoard can be started with:
#
# ```tensorboard --logdir /tmp/Graph```
# ## Previewing Data Augmentation
#
# *ktrain* allows you to preview a data augmentation prior to use in training. We will re-use the Dogs vs. Cats dataset from above. Let's set the location of the data and instantiate a data augmentation scheme using the ```ktrain.vision.get_data_aug``` function.
DATADIR = 'data/dogscats'
data_aug = vis.get_data_aug(horizontal_flip=True)
# Let's examine the data augmentaiton parameters, which will be the parameters enabled by default plus horizontal_flip=True. Note that the data_aug object is simply a Keras **ImageDataGenerator** object.
data_aug.__dict__
# Finally, let's run the data augmentation scheme for four images on a selected cat photo:
vis.preview_data_aug(DATADIR+'/train/cats/cat.10000.jpg', data_aug, n=8, rows=2)
| tutorial-A1-additional-tricks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/hzwer/LearningToPaint/blob/master/LearningToPaint.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="TFN3oT1Hkjfs" colab_type="code" colab={}
# !git clone https://github.com/hzwer/LearningToPaint.git
# + id="Dp7N29tGkwQs" colab_type="code" colab={}
# cd LearningToPaint/
# + [markdown] id="qTbhmFyawzhO" colab_type="text"
# Testing
# + id="z0wTTzOEbvps" colab_type="code" colab={}
# !wget "https://drive.google.com/uc?export=download&id=1-7dVdjCIZIxh8hHJnGTK-RA1-jL1tor4" -O renderer.pkl
# + id="Pfd53Hw2cfaY" colab_type="code" colab={}
# !wget "https://drive.google.com/uc?export=download&id=1a3vpKgjCVXHON4P7wodqhCgCMPgg1KeR" -O actor.pkl
# + id="QZpb3_3QiMZw" colab_type="code" colab={}
# !wget -U NoSuchBrowser/1.0 -O image/test.png https://raw.githubusercontent.com/hzwer/LearningToPaint/master/image/Trump.png
# + id="brX4ZlQoc9ss" colab_type="code" colab={}
# !python3 baseline/test.py --max_step=80 --actor=actor.pkl --renderer=renderer.pkl --img=image/test.png --divide=5
# + id="tLM4U6F0_yjV" colab_type="code" colab={}
# !ffmpeg -r 30 -f image2 -i output/generated%d.png -s 512x512 -c:v libx264 -pix_fmt yuv420p video.mp4 -q:v 0 -q:a 0
# + id="ekY7HcBeh8zl" colab_type="code" colab={}
from IPython.display import display, Image
import moviepy.editor as mpy
display(mpy.ipython_display('video.mp4', height=256, max_duration=100.))
display(Image('output/generated399.png'))
# + [markdown] id="d2mAkgRjwwuf" colab_type="text"
# Training
# + id="_-p0NhqyTqO_" colab_type="code" colab={}
# !mkdir data
# + id="XXAV9RwkTwKh" colab_type="code" colab={}
# cd data
# + colab_type="code" id="IzZUVjdrET2G" colab={}
# !gdown https://drive.google.com/uc?id=0B7EVK8r0v71pZjFTYXZWM3FlRnM
# + colab_type="code" id="zgguAW3eETVd" colab={}
# !unzip img_align_celeba.zip
# + id="XBH--DY-sK8V" colab_type="code" colab={}
# !rm img_align_celeba.zip
# + id="u6mVpjvBvzrb" colab_type="code" colab={}
# cd ..
# + id="-PYJVt8pc6BP" colab_type="code" colab={}
# !python3 baseline/train_renderer.py
# + id="VZWjNmD23gKm" colab_type="code" colab={}
# !pip install tensorboardX
# + id="ehnzhWn9GG4I" colab_type="code" colab={}
# %%writefile baseline/env.py
import sys
import json
import torch
import numpy as np
import argparse
import torchvision.transforms as transforms
import cv2
from DRL.ddpg import decode
from utils.util import *
from PIL import Image
from torchvision import transforms, utils
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
aug = transforms.Compose(
[transforms.ToPILImage(),
transforms.RandomHorizontalFlip(),
])
width = 128
convas_area = width * width
img_train = []
img_test = []
train_num = 0
test_num = 0
class Paint:
def __init__(self, batch_size, max_step):
self.batch_size = batch_size
self.max_step = max_step
self.action_space = (13)
self.observation_space = (self.batch_size, width, width, 7)
self.test = False
def load_data(self):
# CelebA
global train_num, test_num
for i in range(200000):
img_id = '%06d' % (i + 1)
try:
img = cv2.imread('./data/img_align_celeba/' + img_id + '.jpg', cv2.IMREAD_UNCHANGED)
img = cv2.resize(img, (width, width))
if i > 2000:
train_num += 1
img_train.append(img)
else:
test_num += 1
img_test.append(img)
finally:
if (i + 1) % 10000 == 0:
print('loaded {} images'.format(i + 1))
print('finish loading data, {} training images, {} testing images'.format(str(train_num), str(test_num)))
def pre_data(self, id, test):
if test:
img = img_test[id]
else:
img = img_train[id]
if not test:
img = aug(img)
img = np.asarray(img)
return np.transpose(img, (2, 0, 1))
def reset(self, test=False, begin_num=False):
self.test = test
self.imgid = [0] * self.batch_size
self.gt = torch.zeros([self.batch_size, 3, width, width], dtype=torch.uint8).to(device)
for i in range(self.batch_size):
if test:
id = (i + begin_num) % test_num
else:
id = np.random.randint(train_num)
self.imgid[i] = id
self.gt[i] = torch.tensor(self.pre_data(id, test))
self.tot_reward = ((self.gt.float() / 255) ** 2).mean(1).mean(1).mean(1)
self.stepnum = 0
self.canvas = torch.zeros([self.batch_size, 3, width, width], dtype=torch.uint8).to(device)
self.lastdis = self.ini_dis = self.cal_dis()
return self.observation()
def observation(self):
# canvas B * 3 * width * width
# gt B * 3 * width * width
# T B * 1 * width * width
ob = []
T = torch.ones([self.batch_size, 1, width, width], dtype=torch.uint8) * self.stepnum
return torch.cat((self.canvas, self.gt, T.to(device)), 1) # canvas, img, T
def cal_trans(self, s, t):
return (s.transpose(0, 3) * t).transpose(0, 3)
def step(self, action):
self.canvas = (decode(action, self.canvas.float() / 255) * 255).byte()
self.stepnum += 1
ob = self.observation()
done = (self.stepnum == self.max_step)
reward = self.cal_reward() # np.array([0.] * self.batch_size)
return ob.detach(), reward, np.array([done] * self.batch_size), None
def cal_dis(self):
return (((self.canvas.float() - self.gt.float()) / 255) ** 2).mean(1).mean(1).mean(1)
def cal_reward(self):
dis = self.cal_dis()
reward = (self.lastdis - dis) / (self.ini_dis + 1e-8)
self.lastdis = dis
return to_numpy(reward)
# + id="0kwVmo6yv1w3" colab_type="code" colab={}
# !python3 baseline/train.py --max_step=200 --debug --batch_size=96
| LearningToPaint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Frequentism, Samples, and the Bootstrap.
# New Latex commands are defined here. Doubleclick to see.
#
# $\newcommand{\Ex}{\mathbb{E}}$
# $\newcommand{\Var}{\mathrm{Var}}$
# $\newcommand{\Cov}{\mathrm{Cov}}$
# $\newcommand{\SampleAvg}{\frac{1}{N({S})} \sum_{s \in {S}}}$
# $\newcommand{\indic}{\mathbb{1}}$
# $\newcommand{\avg}{\overline}$
# $\newcommand{\est}{\hat}$
# $\newcommand{\trueval}[1]{#1^{*}}$
# $\newcommand{\Gam}[1]{\mathrm{Gamma}#1}$
# The %... is an iPython thing, and is not part of the Python language.
# In this case we're just telling the plotting library to draw things on
# the notebook, instead of on a separate window.
# %matplotlib inline
# See all the "as ..." contructs? They're just aliasing the package names.
# That way we can call methods like plt.plot() instead of matplotlib.pyplot.plot().
import numpy as np
import scipy as sp
import matplotlib as mpl
import matplotlib.cm as cm
import matplotlib.pyplot as plt
import pandas as pd
import time
pd.set_option('display.width', 500)
pd.set_option('display.max_columns', 100)
pd.set_option('display.notebook_repr_html', True)
import seaborn as sns
sns.set_style("whitegrid")
sns.set_context("poster")
# ## DATA AND MODELS
# ### Why do we do this? Lets get some data...
#
# Forty-four babies -- a new record -- were born in one 24-hour period at
# the Mater Mothers' Hospital in Brisbane, Queensland, Australia, on
# December 18, 1997. For each of the 44 babies, _The Sunday Mail_
# recorded the time of birth, the sex of the child, and the birth weight
# in grams. Also included is the number of minutes since midnight for
# each birth.
#
# REFERENCE:
# <NAME>. (December 21, 1997), "Babies by the Dozen for Christmas:
# 24-Hour Baby Boom," _The Sunday Mail_ (Brisbane), p. 7.
#
# "Datasets
# and Stories" article "A Simple Dataset for Demonstrating Common
# Distributions" in the _Journal of Statistics Education_ (Dunn 1999).
#
# Columns
#
# 1 - 8 Time of birth recorded on the 24-hour clock
# 9 - 16 Sex of the child (1 = girl, 2 = boy)
# 17 - 24 Birth weight in grams
# 25 - 32 Number of minutes after midnight of each birth
df = pd.read_table("babyboom.dat.txt", header=None, sep='\s+',
names=['24hrtime','sex','weight','minutes'])
df.head()
df.minutes.mean()
# ### What is data?
#
# In labs before, you have seen datasets. As so in the example above. You have seen probability distributions of this data. Calculated means. Calculated standard deviations.
# #### Pandas code for the week
#
# We'll keep showing some different aspects of Pandas+Seaborn each week. For example, you can very easily calculate correlations
df.corr()
# Remember that this correlation is a statistic calculated only on this data...this sample of babies. I have not asked the question: what does this mean for the population of babies.
#
# I'd thought that there would be a greater correlation between weight and sex, but apparently its not at all big for babies. Telles you Idont know much about babies :-). Here's a plot more tohelp you in future homework:
g = sns.FacetGrid(col="sex", data=df, size=8)
g.map(plt.hist, "weight")
# #### Samples vs population
#
# But we have never aked ourselves the philosophical question: what is data? **Frequentist statistics** is one answer to this philosophical question. It treats data as a **sample** from an existing **population**.
#
# This notion is probably clearest to you from elections, where some companies like Zogby or CNN take polls. The sample in these polls maybe a 1000 people, but they "represent" the electoral population at large. We attempt to draw inferences about how the population will vote based on these samples.
# ### Choosing a model
#
# Let us characterize our particular sample statistically then, using a *probability distribution*
#
#
# #### The Exponential Distribution
#
# The exponential distribution occurs naturally when describing the lengths of the inter-arrival times in a homogeneous Poisson process.
#
# It takes the form:
# $$
# f(x;\lambda) = \begin{cases}
# \lambda e^{-\lambda x} & x \ge 0, \\
# 0 & x < 0.
# \end{cases}
# $$
#
# From Wikipedia: *In probability theory, a Poisson process is a stochastic process which counts the number of events and the time that these events occur in a given time interval. The time between each pair of consecutive events has an exponential distribution with parameter $\lambda$ and each of these inter-arrival times is assumed to be independent of other inter-arrival times. The process is named after the French mathematician <NAME> and is a good model of radioactive decay, telephone calls and requests for a particular document on a web server, among many other phenomena.*
#
# In our example above, we have the arrival times of the babies. There is no reason to expect any specific clustering in time, so one could think of modelling the arrival of the babies via a poisson process.
#
# Furthermore, the Poisson distribution can be used to model the number of births each hour over the 24-hour period.
f = lambda x, l: l*np.exp(-l*x)*(x>0)
xpts=np.arange(-2,3,0.1)
plt.plot(xpts,f(xpts, 2),'o');
plt.xlabel("x")
plt.ylabel("exponential pdf")
# Note: **some of the code, and ALL of the visual style for the distribution plots below was shamelessly stolen from https://gist.github.com/mattions/6113437/ **.
# +
from scipy.stats import expon
x = np.linspace(0,4, 100)
colors=sns.color_palette()
lambda_ = [0.5, 1, 2, 4]
plt.figure(figsize=(12,4))
for l,c in zip(lambda_,colors):
plt.plot(x, expon.pdf(x, scale=1./l), lw=2,
color=c, label = "$\lambda = %.1f$"%l)
plt.fill_between(x, expon.pdf(x, scale=1./l), color=c, alpha = .33)
plt.legend()
plt.ylabel("PDF at $x$")
plt.xlabel("$x$")
plt.title("Probability density function of an Exponential random variable;\
differing $\lambda$");
# -
# ### How would we draw from this distribution?
#
# Lets use the built in machinery in `scipy.stats`:
from scipy.stats import expon
plt.plot(xpts,expon.pdf(xpts, scale=1./2.),'o')
plt.hist(expon.rvs(size=1000, scale=1./2.), normed=True, alpha=0.5, bins=30);
plt.xlabel("x")
plt.title("exponential pdf and samples(normalized)");
# In `scipy.stats`, you can alternatively create a frozen object, which holds values of things like the scale
rv = expon(scale=0.5)
plt.plot(xpts,rv.pdf(xpts),'o')
plt.hist(rv.rvs(size=1000), normed=True, alpha=0.5, bins=30);
plt.plot(xpts, rv.cdf(xpts));
plt.xlabel("x")
plt.title("exponential pdf, cdf and samples(normalized)");
# ### Understanding our data using a distribution
#
# Lets play with our data a bit to understand it:
#
# The first birth occurred at 0005, and the last birth in the 24-hour period at 2355. Thus the 43 inter-birth times happened over a 1430-minute period, giving a theoretical mean of 1430/43 = 33.26 minutes between births.
#
# Lets plot a histogram of the inter-birth times
timediffs = df.minutes.diff()[1:]
timediffs.hist(bins=20);
# The mean or of an exponentially distributed random variable X with rate parameter $\lambda$ can be analytically calculated as
#
# $$\Ex[X] = \frac{1}{\lambda}.$$
#
# This makes intuitive sense: if you get babies at an average rate of 2 per hour, then you can expect to wait half an hour for every baby.
#
# The variance of X is given by
#
# $$\Var[X] = \frac{1}{\lambda^2}.$$
#
# so the standard deviatiation is equal to the mean, just as in the discrete Poisson distribution.
lambda_from_mean = 1./timediffs.mean()
print lambda_from_mean, 1./lambda_from_mean
minutes=np.arange(0, 160, 5)
rv = expon(scale=1./lambda_from_mean)
plt.plot(minutes,rv.pdf(minutes),'o')
timediffs.hist(normed=True, alpha=0.5);
plt.xlabel("minutes");
plt.title("Normalized data and model for estimated $\hat{\lambda}$");
# What did we just do? We made a 'point estimate' of the scale or rate parameter as a compression of our data. But what does it mean to make such a point estimate? The next section on **Frequentist Statistics** tells us. But first, lets see the Poisson Distribution.
# #### An aside: The Poisson Distribution
#
# The *Poisson Distribution* is defined for all positive integers:
#
# $$P(Z=k)=\frac{\lambda^k e^{−\lambda}}{k!}, k=0,1,2,... $$
from scipy.stats import poisson
k = np.arange(15)
plt.figure(figsize=(12,8))
for i, lambda_ in enumerate([1, 2, 4, 6]):
plt.plot(k, poisson.pmf(k, lambda_), '-o', label=lambda_, color=colors[i])
plt.fill_between(k, poisson.pmf(k, lambda_), color=colors[i], alpha=0.5)
plt.legend()
plt.title("Poisson distribution")
plt.ylabel("PDF at $k$")
plt.xlabel("$k$");
per_hour = df.minutes // 60
num_births_per_hour=df.groupby(per_hour).minutes.count()
num_births_per_hour
num_births_per_hour.mean()
k = np.arange(5)
plt.figure(figsize=(12,8))
tcount=num_births_per_hour.sum()
plt.hist(num_births_per_hour, alpha=0.4, lw=3, normed=True, label="normed hist")
sns.kdeplot(num_births_per_hour, label="kde")
plt.plot(k, poisson.pmf(k, num_births_per_hour.mean()), '-o',label="poisson")
plt.title("Baby births")
plt.xlabel("births per hour")
plt.ylabel("rate")
plt.legend();
# ### Maximum Likelihood Estimation
#
# how did we know that the sample mean was a good thing to use?
#
# One of the techniques used to estimate such parameters in frequentist statistics is **maximum likelihood estimation**. Briefly, the idea behind it is:
#
# The product
#
# $$
# L(\lambda) = \prod_{i=1}^n P(x_i | \lambda)
# $$
#
# gives us a measure of how likely it is to observe values $x_1,...,x_n$ given the parameters $\lambda$. Maximum likelihood fitting consists of choosing the appropriate "likelihood" function $L=P(X|\lambda)$ to maximize for a given set of observations. How likely are the observations if the model is true?
#
# Often it is easier and numerically more stable to maximise the log likelyhood:
#
# $$
# \ell(\lambda) = \sum_{i=1}^n ln(P(x_i | \lambda))
# $$
#
# In the case of the exponential distribution we have:
#
# $$
# \ell(lambda) = \sum_{i=1}^n ln(\lambda e^{-\lambda x_i}) = \sum_{i=1}^n \left( ln(\lambda) - \lambda x_i \right).
# $$
#
# Maximizing this:
#
# $$
# \frac{d \ell}{d\lambda} = \frac{n}{\lambda} - \sum_{i=1}^n x_i = 0
# $$
#
# and thus:
#
# $$
# \est{\lambda_{MLE}} = \frac{1}{n}\sum_{i=1}^n x_i,
# $$
#
# which is identical to the simple estimator we used above. Usually one is not so lucky and one must use numerical optimization techniques.
#
# A crucial property is that, for many commonly occurring situations, maximum likelihood parameter estimators have an approximate normal distribution when n is large.
# ## FREQUENTIST STATISTICS
#
# In frequentist statistics, the data we have in hand, is viewed as a **sample** from a population. So if we want to estimate some parameter of the population, like say the mean, we estimate it on the sample.
#
# This is because we've been given only one sample. Ideally we'd want to see the population, but we have no such luck.
#
# The parameter estimate is computed by applying an estimator $F$ to some data $D$, so $\est{\lambda} = F(D)$.
#
#
# **The parameter is viewed as fixed and the data as random, which is the exact opposite of the Bayesian approach which you will learn later in this class. **
#
# For the babies, lets assume that an exponential distribution is a good description of the baby arrival process. Then we consider some larger population of babies from which this sample is drawn, there is some true $\trueval{\lambda}$ which defines it. We dont know this. The best we can do to start with is to estimate a lambda from the data set we have, which we denote $\est{\lambda}$.
#
# Now, imagine that I let you peek at the entire population in this way: I gave you some M data sets **drawn** from the population, and you can now find the mean on each such dataset, of which the one we have here is one.
# So, we'd have M means. You can think of these means as coming from some fixed parameter by some data drawing process
#
# Now if we had many replications of this data set: that is, data from other days, an **ensemble** of data sets, for example, we can compute other $\est{\lambda}$, and begin to construct the **sampling distribution** of $\lambda$.
# ### Segue: many samples on the binomial
from scipy.stats.distributions import bernoulli
def throw_a_coin(n):
brv = bernoulli(0.5)
return brv.rvs(size=n)
# The function below returns the mean for each sample in an ensemble of samples
def make_throws(number_of_samples, sample_size):
start=np.zeros((number_of_samples, sample_size), dtype=int)
for i in range(number_of_samples):
start[i,:]=throw_a_coin(sample_size)
return np.mean(start, axis=1)
# Let us now do 200 replications, each of which has a sample size of 1000 flips, and store the 200 means for each sample zise from 1 to 1000 in `sample_means`. This will rake some time to run as I am doing it for 200 replications at 1000 different sample sizes.
sample_sizes=np.arange(1,1001,1)
sample_means = [make_throws(number_of_samples=200, sample_size=i) for i in sample_sizes]
# So remember that for eachsample size, i am getting 200 means. Lets get the mean of the means at each sample size.
mean_of_sample_means = [np.mean(means) for means in sample_means]
plt.plot(sample_sizes, mean_of_sample_means);
plt.ylim([0.480,0.520]);
# Not surprisingly, the mean of the sample means converges to the distribution mean as the sample size N gets very large.
#
# #### The notion of a Sampling Distribution
#
# (some text is quoted from Murphy's machine learning book)
#
# In data science, we are always interested in understanding the world from incomplete data, in other words from a sample or a few samples of a population at large. Our experience with the world tells us that even if we are able to repeat an experiment or process, we will get more or less different answers the next time. If all of the answers were very different each time, we would never be able to make any predictions.
#
# But some kind of answers differ only a little, especially as we get to larger sample sizes. So the important question then becomes one of the distribution of these quantities from sample to sample, also known as a **sampling distribution**.
#
# Since, in the real world, we see only one sample, this distribution helps us do **inference**, or figure the uncertainty of the estimates of quantities we are interested in. If we can somehow cook up samples just somewhat different from the one we were given, we can calculate quantities of interest, such as the mean on each one of these samples. By seeing how these means vary from one sample to the other, we can say how typical the mean in the sample we were given is, and whats the uncertainty range of this quantity. This is why the mean of the sample means is an interesting quantity; it characterizes the **sampling distribution of the mean**, or the distribution of sample means.
#
# So, in the babies case, the uncertainty in the parameter estimate can be measured by computing the **sampling distribution** of the estimator.
# What you are doing is sampling many Data Sets $D_i$ from the true population (which we are not given you will argue, and you are right, but just wait a bit), say M of them, each of size N, from some true model $p(\cdot|\trueval{\lambda})$. We will now calculate M $\est{\lambda}_i$, one for each dataset. As we let $M \rightarrow \infty$, the distribution induced on $\est{\lambda}$ is the sampling distribution of the estimator.
# ## Inference
#
# Just having an estimate is no good. We will want to put confidence intervals on the estimation of the parameters. This presents a conundrum: we have access to only one sample, but want to compute a error estimate over multiple samples, using an estimator such as the standard deviation.
#
# At this point we are wishing for the Lord to have given us those other samples drawn from the population that we talked about above. But alas, no such luck...
#
# In the last two decades, **resampling** the ONE dataset we have has become computationally feasible. Resampling involves making new samples from the observations, each of which is analysed in the same way as out original dataset. One way to do this is the Bootstrap.
# ### Bootstrap
#
# Bootstrap tries to approximate our sampling distribution. If we knew the true parameters of the population, we could generate M fake datasets. Then we could compute the parameter (or another estimator) on each one of these, to get a empirical sampling distribution of the parameter or estimator, and which will give us an idea of how typical our sample is, and thus, how good our parameter estimations from our sample are.
# (again from murphy)
#
# But we dont have the true parameter. So we generate these samples, using the parameter we calculated. Or, alteratively, we sample with replacement the X from our original sample D, generating many fake datasets, and then compute the distribution on the parameters as before.
#
# We do it here for the mean of the time differences. We could also do it for its inverse, $\lambda$.
#
# #### Non Parametric bootstrap
#
# Resample the data! We can then plot the distribution of the mean time-difference.
M_samples=10000
N_points = timediffs.shape[0]
bs_np = np.random.choice(timediffs, size=(M_samples, N_points))
sd_mean=np.mean(bs_np, axis=1)
sd_std=np.std(bs_np, axis=1)
plt.hist(sd_mean, bins=30, normed=True, alpha=0.5,label="samples");
sns.kdeplot(sd_mean, label="inferred distribution")
plt.axvline(timediffs.mean(), 0, 1, color='r', label='Our Sample')
plt.legend()
# #### Parametric Bootstrap
#
# And here we do it in a parametric way. We get an "estimate" of the parameter from our sample, and them use the exponential distribution to generate many datasets, and then fir the parameter on each one of those datasets. We can then plot the distribution of the mean time-difference.
rv = expon(scale=1./lambda_from_mean)
M_samples=10000
N_points = timediffs.shape[0]
bs_p = rv.rvs(size=(M_samples, N_points))
sd_mean_p=np.mean(bs_p, axis=1)
sd_std_p=np.std(bs_p, axis=1)
plt.hist(sd_mean_p, bins=30, normed=True, alpha=0.5);
sns.kdeplot(sd_mean_p);
plt.axvline(timediffs.mean(), 0, 1, color='r', label='Our Sample')
| Labs/Resources/stats_probs/Freq.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### New to Plotly?
# Plotly's Python library is free and open source! [Get started](https://plotly.com/python/getting-started/) by downloading the client and [reading the primer](https://plotly.com/python/getting-started/).
# <br>You can set up Plotly to work in [online](https://plotly.com/python/getting-started/#initialization-for-online-plotting) or [offline](https://plotly.com/python/getting-started/#initialization-for-offline-plotting) mode, or in [jupyter notebooks](https://plotly.com/python/getting-started/#start-plotting-online).
# <br>We also have a quick-reference [cheatsheet](https://images.plot.ly/plotly-documentation/images/python_cheat_sheet.pdf) (new!) to help you get started!
# #### Version Check
# Plotly's python package is updated frequently. Run `pip install plotly --upgrade` to use the latest version.
import plotly
plotly.__version__
# #### NYC Flights Database
# +
import datetime
import numpy as np
import pandas as pd
import plotly.graph_objs as go
import plotly.plotly as py
from ipywidgets import widgets
# -
# We'll be making an application to take a look at delays from all flights out of NYC in the year 2013.
df = pd.read_csv(
'https://raw.githubusercontent.com/yankev/testing/master/datasets/nycflights.csv')
df = df.drop(df.columns[[0]], axis=1)
df.sample(3)
# Let's get the set of all the `airlines`, so that we can type the right things into the search box later.
df['carrier'].unique()
# Let's assign the widgets that we're going to be using in our app. In general all these widgets will be used to filter the data set, and thus what we visualize.
# +
month = widgets.IntSlider(
value=1.0,
min=1.0,
max=12.0,
step=1.0,
description='Month:',
continuous_update=False
)
use_date = widgets.Checkbox(
description='Date: ',
value=True,
)
container = widgets.HBox(children=[use_date, month])
textbox = widgets.Dropdown(
description='Airline: ',
value='DL',
options=df['carrier'].unique().tolist()
)
origin = widgets.Dropdown(
options=list(df['origin'].unique()),
value='LGA',
description='Origin Airport:',
)
# Assign an emptry figure widget with two traces
trace1 = go.Histogram(x=df['arr_delay'], opacity=0.75, name='Arrival Delays')
trace2 = go.Histogram(x=df['dep_delay'], opacity=0.75, name='Departure Delays')
g = go.FigureWidget(data=[trace1, trace2],
layout=go.Layout(
title=dict(
text='NYC FlightDatabase'
),
barmode='overlay'
))
# -
# Let now write a function that will handle the input from the widgets, and alter the state of the graph.
# +
def validate():
if origin.value in df['origin'].unique() and textbox.value in df['carrier'].unique():
return True
else:
return False
def response(change):
if validate():
if use_date.value:
filter_list = [i and j and k for i, j, k in
zip(df['month'] == month.value, df['carrier'] == textbox.value,
df['origin'] == origin.value)]
temp_df = df[filter_list]
else:
filter_list = [i and j for i, j in
zip(df['carrier'] == 'DL', df['origin'] == origin.value)]
temp_df = df[filter_list]
x1 = temp_df['arr_delay']
x2 = temp_df['dep_delay']
with g.batch_update():
g.data[0].x = x1
g.data[1].x = x2
g.layout.barmode = 'overlay'
g.layout.xaxis.title = 'Delay in Minutes'
g.layout.yaxis.title = 'Number of Delays'
origin.observe(response, names="value")
textbox.observe(response, names="value")
month.observe(response, names="value")
use_date.observe(response, names="value")
# -
# Time to try the app out!!
container2 = widgets.HBox([origin, textbox])
widgets.VBox([container,
container2,
g])
# + language="html"
# <img src = 'https://cloud.githubusercontent.com/assets/12302455/16637308/4e476280-43ac-11e6-9fd3-ada2c9506ee1.gif' >
# -
# #### Reference
help(go.FigureWidget)
# +
from IPython.display import display, HTML
display(HTML('<link href="//fonts.googleapis.com/css?family=Open+Sans:600,400,300,200|Inconsolata|Ubuntu+Mono:400,700" rel="stylesheet" type="text/css" />'))
display(HTML('<link rel="stylesheet" type="text/css" href="http://help.plot.ly/documentation/all_static/css/ipython-notebook-custom.css">'))
# !pip install git+https://github.com/plotly/publisher.git --upgrade
import publisher
publisher.publish(
'figurewidget_app.ipynb', 'python/figurewidget-app/', 'IPython FigureWidget',
'Interactive Data Analysis with Plotly',
title='Interactive Data Analysis with FigureWidget ipywidgets',
name='Interactive Data Analysis with FigureWidget ipywidgets',
has_thumbnail='true', thumbnail='thumbnail/multi-widget.jpg',
language='python', page_type='example_index',
display_as='chart_events', order=23,
ipynb='~notebook_demo/231')
# -
| _posts/python-v3/javascript-controls/figureWidgets/figurewidget_app.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
from csd import CSD
from csd.typings.typing import MeasuringTypes, CSDConfiguration, Backends
import numpy as np
alphas = list(np.arange(0.05, 1.55, 0.05))
csd = CSD(csd_config=CSDConfiguration({
'steps': 100,
'cutoff_dim': 10,
'batch_size': 10,
'architecture': {
'displacement': True,
'squeezing': False,
},
'save_results': False,
'save_plots': False
}))
results = csd.execute_all_backends_and_measuring_types(
alphas=alphas,
backends=[Backends.TENSORFLOW],
measuring_types=[MeasuringTypes.PROBABILITIES]
)
csd.plot_success_probabilities(measuring_types=[MeasuringTypes.PROBABILITIES])
csd.plot_success_probabilities(measuring_types=[MeasuringTypes.PROBABILITIES])
results = csd.execute_all_backends_and_measuring_types(
alphas=alphas,
measuring_types=[MeasuringTypes.SAMPLING]
)
csd.plot_success_probabilities(measuring_types=[MeasuringTypes.SAMPLING])
| examples/csd_optimization_with_plots.ipynb |
// -*- coding: utf-8 -*-
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .cpp
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: C++17
// language: C++17
// name: xcpp17
// ---
// # Aktualisieren der Materialien
//
// Sind Sie **nicht** auf der [Metis](https://metis.informatik.uos.de) müssen Sie die Materialien regelmäßig aktualisieren.
// Das geschieht automatisch, wenn Jupyter Notebook über die Links oder das Skript `aud` gestartet wird. Anonsten
// gibt es in der [**Baum-Ansicht**](http://localhost:8888/tree) den Button **Download AuD Material**. Wird dieser angeklickt, werden die aktuellen Materialien heruntergeladen. Das dauert unter Umständen etwas. Es sollten zwei Meldungen am unteren Bildschirmrand erscheinen. Als erstes
//
// > Downloading material from `http://abbozza.informatik.uos.de/aud/jupyter/aud-materila.zip`
//
// Nach einer gewissen Zeit (etwas gedulden!) sollte die Meldung
//
// > Material successfully updated!
//
// erscheinen. Dann müsste sich das Verzeichnis automatisch erneuern. Im Zweifelsfall führen sie einen **Reload** der Seite durch (F5).
//
// Das wars auch schon.
| lessons/00_Organisatorisches/01_Aktualisierung.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc=true
# # Symmetric Planar Waveguides
#
# **<NAME>**
#
# **Aug 2021**
#
# Planar waveguides are a strange abstraction. These are waveguides that are sandwiches with a specified thickness but are infinite in extent in the other directions. Studying planar waveguides before cylindrical waveguides is done because the math is simpler (solutions are trignometric functions instead of Bessel functions) and therefore it is a bit less likely that one will get lost in the math.
#
# ---
# *If* `` ofiber `` *is not installed, uncomment the following cell (i.e., delete the initial #) and execute it with* `` shift-enter ``. *Afterwards, you may need to restart the kernel/runtime before the module will import successfully.*"
# +
# #!pip install --user ofiber
# +
# Jupyterlite support for ofiber
try:
import micropip
await micropip.install("ofiber")
except ModuleNotFoundError:
pass
try:
import numpy as np
import matplotlib.pyplot as plt
import scipy
import ofiber
except ModuleNotFoundError:
print('ofiber is not installed. To install, uncomment and run the cell above.')
print('Once installation is successful, rerun this cell again.')
# to make graphs a bit better
# %config InlineBackend.figure_format='retina'
plt.style.use('seaborn-talk')
# -
# ## Modes in planar waveguides
#
# ### V=3.15
# +
V=3.15
xx = ofiber.TE_crossings(V)
aplt = ofiber.TE_mode_plot(V)
yy = np.sqrt((V / 2)**2 - xx[0::2]**2)
aplt.scatter(xx[0::2],yy,s=50)
yy = np.sqrt((V / 2)**2 - xx[1::2]**2)
aplt.scatter(xx[1::2],yy,s=50)
aplt.show()
# -
# ### V=4.77
# +
n1=1.503
n2=1.5
lambda0 = 0.5e-6
k=2*np.pi/lambda0
NA = np.sqrt(n1**2-n2**2)
d = 4e-6
V = k * d * NA
xx = ofiber.TE_crossings(V)
b = 1-(2*xx/V)**2
beta = np.sqrt((n1**2-n2**2)*b+n2**2)
theta = np.arccos(beta/n1)*180/np.pi
aplt = ofiber.TE_mode_plot(V)
yy = np.sqrt((V / 2)**2 - xx[0::2]**2)
aplt.scatter(xx[0::2],yy,s=50)
yy = np.sqrt((V / 2)**2 - xx[1::2]**2)
aplt.scatter(xx[1::2],yy,s=50)
aplt.show()
print(xx)
print('b =',b)
print('beta hat=',beta)
print('theta =',theta,' degrees')
# -
# ### V=5.5
# +
V=5.5
xx = ofiber.TE_crossings(V)
aplt = ofiber.TE_mode_plot(V)
yy = np.sqrt((V / 2)**2 - xx[0::2]**2)
aplt.scatter(xx[0::2],yy,s=50)
yy = np.sqrt((V / 2)**2 - xx[1::2]**2)
aplt.scatter(xx[1::2],yy,s=50)
aplt.show()
print('cutoff wavelength = %.0f nm'%(2*d*NA*1e9))
# -
# ### V=16
# +
V=16
n1=1.5
n2=1.49
xx = ofiber.TM_crossings(V,n1,n2)
aplt = ofiber.TM_mode_plot(V,n1,n2)
yy = np.sqrt((V / 2)**2 - xx[0::2]**2)
aplt.scatter(xx[0::2],yy,s=50)
yy = np.sqrt((V / 2)**2 - xx[1::2]**2)
aplt.scatter(xx[1::2],yy,s=50)
aplt.show()
# -
# ## Internal field inside waveguide
# +
V= 15
d = 1
x = np.linspace(-1,1,100)
m=1
plt.plot(x,ofiber.TE_field(V,d,x,m))
plt.annotate('m=%d'%m,xy=(0.25,0.7))
m=0
plt.plot(x,ofiber.TE_field(V,d,x,m))
plt.annotate('m=%d'%m,xy=(0.25,-0.7))
m=2
plt.plot(x,ofiber.TE_field(V,d,x,m))
plt.annotate('m=%d'%m,xy=(0.25,-0.7))
plt.plot(x,np.exp(-x**2/0.01),':b')
plt.plot([-1,1],[0,0],'k')
plt.plot([-0.5,-0.5],[-1,1],':k')
plt.plot([0.5,0.5],[-1,1],':k')
plt.annotate('planar bottom ',xy=(-0.5,-1),ha='right')
plt.annotate(' planar top',xy=(0.5,-1),ha='left')
plt.xlabel('Position (x/d)')
plt.ylabel('$|E_y(x)|^2$ [Normalized]')
plt.title('Modal Fields in symmetric planar waveguide V=%.2f'%V)
plt.show()
# -
# ## TE propagation constants for first five modes
# +
V = np.linspace(0.1,25,50)
for mode in range(5):
b = ofiber.TE_propagation_constant(V,mode)
plt.plot(V,b)
plt.text(25.5,b[-1],"mode=%d"%mode,va='center')
plt.xlabel("V")
plt.ylabel('b')
plt.title('Normalized TE Propagation Constants for Planar Waveguide')
plt.xlim(0,30)
plt.show()
# -
# ## TE & TM propagation constants for first five modes
# +
n1=1.5
n2=1.0
V = np.linspace(0.1,30,50)
for mode in range(7):
b = ofiber.TM_propagation_constant(V,n1,n2,mode)
plt.annotate(' mode=%d'%mode,xy=(30,b[-1]))
plt.plot(V,b,':b')
b = ofiber.TE_propagation_constant(V,mode)
plt.plot(V,b,'r')
plt.xlabel("V")
plt.ylabel('b')
plt.title('Normalized Propagation Constant for Planar Waveguide')
plt.xlim(0,35)
plt.show()
# -
| docs/3-Planar-Waveguide-Modes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Belajar memahami data dengan melihat visualisasinya
# Pie Chart
# +
import matplotlib.pyplot as plt
rasa = ('coklat', 'strawberry', 'vanila', 'blueberry', 'taro')
data = (12, 15, 3, 16, 5)
plt.pie(data, labels = rasa,)
plt.show()
# +
import matplotlib.pyplot as plt
rasa = ('coklat', 'strawberry', 'vanila', 'blueberry', 'taro')
data = (12, 15, 3, 16, 5)
plt.pie(data, labels = rasa, autopct='%1.2f%%')
plt.show()
# +
import matplotlib.pyplot as plt
rasa = ('coklat', 'strawberry', 'vanila', 'blueberry', 'taro')
data = (12, 15, 3, 16, 5)
warna = ('#A52A2A', '#FF0000', '#FFF5EE', '#800080', '#4682B4')
plt.pie(data, labels = rasa, autopct='%1.2f%%', colors=warna)
plt.show()
# +
import matplotlib.pyplot as plt
rasa = ('coklat', 'strawberry', 'vanila', 'blueberry', 'taro')
data = (12, 15, 3, 16, 5)
warna = ('#A52A2A', '#FF0000', '#FFF5EE', '#800080', '#4682B4')
highlight = (0,0,0.1,0,0)
plt.pie(data, labels = rasa, autopct='%1.2f%%', colors=warna, explode = highlight)
plt.show()
# +
import matplotlib.pyplot as plt
rasa = ('coklat', 'strawberry', 'vanila', 'blueberry', 'taro')
data = (12, 15, 3, 16, 5)
warna = ('#A52A2A', '#FF0000', '#FFF5EE', '#800080', '#4682B4')
highlight = (0,0,0.1,0,0)
plt.pie(data, labels = rasa, autopct='%1.2f%%', colors=warna, explode = highlight, shadow = True)
plt.show()
# +
import matplotlib.pyplot as plt
rasa = ('coklat', 'strawberry', 'vanila', 'blueberry', 'taro')
data = (12, 15, 3, 16, 5)
warna = ('#A52A2A', '#FF0000', '#FFF5EE', '#800080', '#4682B4')
highlight = (0,0,0.3,0,0)
plt.title('Penjualan Es Krim yang Kurang Diminati')
plt.pie(data, labels = rasa, autopct='%1.2f%%', colors=warna, explode = highlight, shadow = True)
plt.show()
# +
#challenge: buat supaya visualisasi menampilkan data eskrim yang paling diminati
import matplotlib.pyplot as plt
rasa = ('coklat', 'strawberry', 'vanila', 'blueberry', 'taro')
data = (12, 15, 3, 16, 5)
warna = ('#A52A2A', '#FF0000', '#FFF5EE', '#800080', '#4682B4')
highlight = (0,0.1,0,0,0)
plt.title('Penjualan Es Krim yang Paling Diminati')
plt.pie(data, labels = rasa, autopct='%1.2f%%', colors=warna, explode = highlight, shadow = True)
plt.show()
# -
# Bar Chart
# +
import matplotlib.pyplot as plt
import numpy as np
negara = ('Argentina', 'Brazil', 'Chile', 'Denmark', 'Finlandia', 'Germany', 'Hongaria', 'Indonesia', 'Japan', 'Kamerun')
populasi = (45380000, 212600000, 19120000, 5831000, 5531000, 83240000, 9750000, 273500000, 12580000, 2655000)
koordinat_x = np.arange(len(negara))
plt.bar(koordinat_x, populasi)
plt.show()
# +
import matplotlib.pyplot as plt
import numpy as np
negara = ('Argentina', 'Brazil', 'Chile', 'Denmark', 'Finlandia', 'Germany', 'Hongaria', 'Indonesia', 'Japan', 'Kamerun')
populasi = (45380000, 212600000, 19120000, 5831000, 5531000, 83240000, 9750000, 273500000, 12580000, 2655000)
koordinat_x = np.arange(len(negara))
plt.bar(koordinat_x, populasi, tick_label = negara)
plt.xticks(rotation = 90)
plt.ylabel('Populasi ( juta)')
plt.title('populasi 10 negara')
plt.show()
# +
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
negara = ('Argentina', 'Brazil', 'Chile', 'Denmark', 'Finlandia', 'Germany', 'Hongaria', 'Indonesia', 'Japan', 'Kamerun')
populasi = (45380000, 212600000, 19120000, 5831000, 5531000, 83240000, 9750000, 273500000, 12580000, 2655000)
df = pd.DataFrame({'Country': negara, 'Population': populasi,})
df.sort_values(by='Population', inplace = True)
koordinat_x = np.arange(len(data_frame))
plt.bar(koordinat_x, df['Population'], tick_label = df['Country'])
plt.xticks(rotation = 90)
plt.ylabel('Populasi (juta)')
plt.title('populasi 10 negara')
plt.show()
# +
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
negara = ('Argentina', 'Brazil', 'Chile', 'Denmark', 'Finlandia', 'Germany', 'Hongaria', 'Indonesia', 'Japan', 'Kamerun')
populasi = (45380000, 212600000, 19120000, 5831000, 5531000, 83240000, 9750000, 273500000, 12580000, 2655000)
df = pd.DataFrame({'Country': negara, 'Population': populasi,})
df.sort_values(by='Population', inplace = True)
koordinat_x = np.arange(len(df))
warna = ['#0000FF' for _ in range(len(df))]
warna [-3] = '#FF0000'
plt.figure(figsize=(20,10))
plt.bar(koordinat_x, df['Population'], tick_label = df['Country'], color = warna)
plt.xticks(rotation = 90)
plt.ylabel('Populasi (juta)')
plt.title('populasi 10 negara (rendah ke tinggi)')
plt.show()
# +
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
negara = ('Argentina', 'Brazil', 'Chile', 'Denmark', 'Finlandia', 'Germany', 'Hongaria', 'Indonesia', 'Japan', 'Kamerun')
populasi = (45380000, 212600000, 19120000, 5831000, 5531000, 83240000, 9750000, 273500000, 12580000, 2655000)
df = pd.DataFrame({'Country': negara, 'Population': populasi,})
df.sort_values(by='Population', inplace = True, ascending = False)
koordinat_x = np.arange(len(df))
warna = ['#0000FF' for _ in range(len(df))]
warna [-3] = '#FF0000'
plt.figure(figsize=(20,10))
plt.bar(koordinat_x, df['Population'], tick_label = df['Country'], color = warna)
plt.xticks(rotation = 90)
plt.ylabel('Populasi (juta)')
plt.title('populasi 10 negara (tinggi ke rendah)')
plt.show()
# -
# Line Graph
# +
import matplotlib.pyplot as plt
temperatur_c = [2,1,0,0,1,5,8,9,8,5,3,2,2]
jam = [0,2,4,6,8,10,12,14,16,18,20,22,24]
plt.plot(jam, temperatur_c, marker = 'o')
plt.title('suhu salatiga 24 Februari 2022')
plt.ylabel('suhu (celcius)')
plt.xlabel('jam')
plt.show()
# +
import matplotlib.pyplot as plt
temperatur_c = [2,1,0,0,1,5,8,9,8,5,3,2,2]
temperatur_c_prediksi = [2,2,3,1,0,4,6,10,9,3,1,3,3]
jam = [0,2,4,6,8,10,12,14,16,18,20,22,24]
plt.plot(jam, temperatur_c, marker = 'o')
plt.plot(jam, temperatur_c_prediksi, linestyle = '--')
plt.title('suhu salatiga 24 Februari 2022')
plt.ylabel('suhu (celcius)')
plt.xlabel('jam')
plt.show()
# -
# Scatter Plot
# +
import matplotlib.pyplot as plt
#import numpy as np
#import pandas as pd
negara = ['Argentina', 'Brazil', 'Chile', 'Denmark', 'Finlandia', 'Germany', 'Hongaria', 'Indonesia', 'Japan', 'Kamerun']
populasi = [45380000, 212600000, 19120000, 5831000, 5531000, 83240000, 9750000, 273500000, 12580000, 2655000]
gdp = [383, 1445, 252, 355, 271, 3806, 155, 1508, 5065, 39]
plt.scatter(populasi, gdp)
plt.show()
# -
# Heatmap
# +
import seaborn as sns
kota = ['Jakarta', 'Surakarta', 'Semarang', 'Surabaya', 'Pati', 'Salatiga', 'Ungaran', 'Tokyo', 'Kyoto', 'Delhi', 'Shanghai', 'New York', 'Antananarivo', 'Ghuangzou', 'Muntilan', 'Merakmati']
bulan = ['Januari', 'Februari', 'Maret', 'April', 'Mei', 'Juni', 'Juli', 'Agustus', 'September', 'Oktober', 'November', 'Desember']
temperatur = [
[20, 32, 12, 19, 22, 25, 15, 19, 24, 21, 11, 32], #Jakarta
[23, 22, 32, 29, 42, 21, 11, 18, 34, 27, 13, 24],#Surakarta
[18, 22, 10, 23, 54, 23, 27, 34, 12, 34, 18, 41],#Semarang
[10, 12, 14, 33, 24, 33, 19, 24, 32, 24, 17, 21],#Surabaya
[18, 27, 24, 23, 34, 13, 29, 44, 12, 29, 22, 39],#Pati
[20, 21, 33, 21, 16, 12, 24, 23, 35, 10, 32, 82],#Salatiga
[34, 12, 32, 18, 34, 52, 41, 23, 65, 26, 44, 12],#Ungaran
[1, 12, 4, 2, 4, 5, 2, 8, 6, 6, 4, 1],#Tokyo
[3, 18, 20, 19, 17, 11, 12, 18, 16, 16, 14, 11],#Kyoto
[24, 22, 22, 28, 32, 22, 21, 23, 25, 26, 24, 12],#Delhi
[24, 32, 34, 38, 32, 32, 31, 33, 35, 36, 34, 22],#Shanghai
[4, 22, 1, 21, 32, 22, 11, 33, 25, 56, 14, 12],#New York
[4, 22, 1, 21, 32, 22, 11, 33, 25, 56, 14, 12],#Antananarivo
[23, 22, 32, 29, 42, 21, 11, 18, 34, 27, 13, 24],#Ghuangzou
[10, 12, 14, 33, 24, 33, 19, 24, 32, 24, 17, 21],#Muntilan
[34, 12, 32, 18, 34, 52, 41, 23, 65, 26, 44, 12],#Merakmati
]
sns.heatmap(temperatur, yticklabels=kota, xticklabels= bulan,)
# +
import seaborn as sns
kota = ['Jakarta', 'Surakarta', 'Semarang', 'Surabaya', 'Pati', 'Salatiga', 'Ungaran', 'Tokyo', 'Kyoto', 'Delhi', 'Shanghai', 'New York', 'Antananarivo', 'Ghuangzou', 'Muntilan', 'Merakmati']
bulan = ['Januari', 'Februari', 'Maret', 'April', 'Mei', 'Juni', 'Juli', 'Agustus', 'September', 'Oktober', 'November', 'Desember']
temperatur = [
[20, 32, 12, 19, 22, 25, 15, 19, 24, 21, 11, 32], #Jakarta
[23, 22, 32, 29, 42, 21, 11, 18, 34, 27, 13, 24],#Surakarta
[18, 22, 10, 23, 54, 23, 27, 34, 12, 34, 18, 41],#Semarang
[10, 12, 14, 33, 24, 33, 19, 24, 32, 24, 17, 21],#Surabaya
[18, 27, 24, 23, 34, 13, 29, 44, 12, 29, 22, 39],#Pati
[20, 21, 33, 21, 16, 12, 24, 23, 35, 10, 32, 82],#Salatiga
[34, 12, 32, 18, 34, 52, 41, 23, 65, 26, 44, 12],#Ungaran
[1, 12, 4, 2, 4, 5, 2, 8, 6, 6, 4, 1],#Tokyo
[3, 18, 20, 19, 17, 11, 12, 18, 16, 16, 14, 11],#Kyoto
[24, 22, 22, 28, 32, 22, 21, 23, 25, 26, 24, 12],#Delhi
[24, 32, 34, 38, 32, 32, 31, 33, 35, 36, 34, 22],#Shanghai
[4, 22, 1, 21, 32, 22, 11, 33, 25, 56, 14, 12],#New York
[4, 22, 1, 21, 32, 22, 11, 33, 25, 56, 14, 12],#Antananarivo
[23, 22, 32, 29, 42, 21, 11, 18, 34, 27, 13, 24],#Ghuangzou
[10, 12, 14, 33, 24, 33, 19, 24, 32, 24, 17, 21],#Muntilan
[34, 12, 32, 18, 34, 52, 41, 23, 65, 26, 44, 12],#Merakmati
]
sns.heatmap(temperatur, yticklabels=kota, xticklabels= bulan, cmap = 'coolwarm',)
| pertemuan6_metdat/data_understanding_visualisasi.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction to `cf_xarray`
#
# This notebook is a brief introduction to `cf_xarray`'s current capabilities.
#
import cf_xarray as cfxr
import numpy as np
import xarray as xr
# `cf_xarray` works best when `xarray` keeps attributes by default.
#
xr.set_options(keep_attrs=True)
# Lets read two datasets.
#
ds = xr.tutorial.load_dataset("air_temperature")
ds.air.attrs["standard_name"] = "air_temperature"
ds
# This one is inspired by POP model output and illustrates how the coordinates
# attribute is interpreted. It also illustrates one way of tagging curvilinear
# grids for convenient use of `cf_xarray`
#
# +
from cf_xarray.datasets import popds as pop
pop
# -
# This synthetic dataset has multiple `X` and `Y` coords. An example would be
# model output on a staggered grid.
#
# +
from cf_xarray.datasets import multiple
multiple
# -
# This dataset has ancillary variables
#
# +
from cf_xarray.datasets import anc
anc
# -
# ## What attributes have been discovered?
#
# The criteria for identifying variables using CF attributes are listed
# [here](../criteria.rst).
#
ds.lon
# `ds.lon` has attributes `axis: X`. This means that `cf_xarray` can identify the
# `'X'` axis as being represented by the `lon` variable.
#
# It can also use the `standard_name` and `units` attributes to infer that `lon`
# is "Longitude". To see variable names that `cf_xarray` can infer, use `ds.cf`
#
ds.cf
# For `pop`, only `latitude` and `longitude` are detected, not `X` or `Y`. Please
# comment here: https://github.com/xarray-contrib/cf-xarray/issues/23 if you have
# opinions about this behaviour.
#
pop.cf
# For `multiple`, multiple `X` and `Y` coordinates are detected
#
multiple.cf
# ## Feature: Accessing coordinate variables
#
# `.cf` implements `__getitem__` to allow easy access to coordinate and axis
# variables.
#
ds.cf["X"]
# Indexing with a scalar key raises an error if the key maps to multiple variables
# names
#
# + tags=["raises-exception"]
multiple.cf["X"]
# + tags=["raises-exception"]
pop.cf["longitude"]
# -
# To get back all variables associated with that key, pass a single element list
# instead.
#
multiple.cf[["X"]]
pop.cf[["longitude"]]
# DataArrays return DataArrays
#
pop.UVEL.cf["longitude"]
# `Dataset.cf[...]` returns a single `DataArray`, parsing the `coordinates`
# attribute if present, so we correctly get the `TLONG` variable and not the
# `ULONG` variable
#
pop.cf["TEMP"]
# `Dataset.cf[...]` also interprets the `ancillary_variables` attribute. The
# ancillary variables are returned as coordinates of a DataArray
#
anc.cf["q"]
# ## Feature: Accessing variables by standard names
#
pop.cf[["sea_water_potential_temperature", "UVEL"]]
# Note that ancillary variables are included as coordinate variables
#
anc.cf["specific_humidity"]
# ## Feature: Utility functions
#
# There are some utility functions to allow use by downstream libraries
#
pop.cf.keys()
# You can test for presence of these keys
#
"sea_water_x_velocity" in pop.cf
# You can also get out the available Axis names
#
pop.cf.axes
# or available Coordinate names. Same for cell measures (`.cf.cell_measures`) and
# standard names (`.cf.standard_names`).
#
pop.cf.coordinates
# **Note:** Although it is possible to assign additional coordinates,
# `.cf.coordinates` only returns a subset of
# `("longitude", "latitude", "vertical", "time")`.
#
# ## Feature: Rewriting property dictionaries
#
# `cf_xarray` will rewrite the `.sizes` and `.chunks` dictionaries so that one can
# index by a special CF axis or coordinate name
#
ds.cf.sizes
# Note the duplicate entries above:
#
# 1. One for `X`, `Y`, `T`
# 2. and one for `longitude`, `latitude` and `time`.
#
# An error is raised if there are multiple `'X'` variables (for example)
#
# + tags=["raises-exception"]
multiple.cf.sizes
# -
multiple.v1.cf.sizes
# ## Feature: Renaming variables
#
# `cf_xarray` lets you rewrite variables in one dataset to like variables in
# another dataset.
#
# In this example, a one-to-one mapping is not possible and the coordinate
# variables are not renamed.
#
da = pop.cf["TEMP"]
da.cf.rename_like(ds)
# If we exclude all axes (variables with `axis` attribute), a one-to-one mapping
# is possible. In this example, `TLONG` and `TLAT` are renamed to `lon` and `lat`
# i.e. their counterparts in `ds`. Note the the `coordinates` attribute is
# appropriately changed.
#
da.cf.rename_like(ds, skip="axes")
# ## Feature: Rewriting arguments
#
# `cf_xarray` can rewrite arguments for a large number of xarray functions. By
# this I mean that instead of specifing say `dim="lon"`, you can pass `dim="X"` or
# `dim="longitude"` and `cf_xarray` will rewrite that to `dim="lon"` based on the
# attributes present in the dataset.
#
# Here are a few examples
#
# ### Slicing
#
ds.air.cf.isel(T=1)
# Slicing works will expand a single key like `X` to multiple dimensions if those
# dimensions are tagged with `axis: X`
#
multiple.cf.isel(X=1, Y=1)
# ### Reductions
#
ds.air.cf.mean("X")
# Expanding to multiple dimensions is also supported
#
# takes the mean along ["x1", "x2"]
multiple.cf.mean("X")
# ### Plotting
#
ds.air.cf.isel(time=1).cf.plot(x="X", y="Y")
ds.air.cf.isel(T=1, Y=[0, 1, 2]).cf.plot(x="longitude", hue="latitude")
# `cf_xarray` can facet
#
seasonal = (
ds.air.groupby("time.season")
.mean()
.reindex(season=["DJF", "MAM", "JJA", "SON"])
)
seasonal.cf.plot(x="longitude", y="latitude", col="season")
# ### Resample & groupby
#
ds.cf.resample(T="D").mean()
# `cf_xarray` also understands the "datetime accessor" syntax for groupby
#
ds.cf.groupby("T.month").mean("longitude")
# ### Rolling & coarsen
#
ds.cf.rolling(X=5).mean()
# `coarsen` works but everything later will break because of xarray bug
# https://github.com/pydata/xarray/issues/4120
#
# `ds.isel(lon=slice(50)).cf.coarsen(Y=5, X=10).mean()`
#
# ## Feature: mix "special names" and variable names
#
ds.cf.groupby("T.month").mean(["lat", "X"])
# ## Feature: Weight by Cell Measures
#
# `cf_xarray` can weight by cell measure variables if the appropriate attribute is
# set
#
# Lets make some weights (not sure if this is right)
ds.coords["cell_area"] = (
np.cos(ds.air.cf["latitude"] * np.pi / 180)
* xr.ones_like(ds.air.cf["longitude"])
* 105e3
* 110e3
)
# and set proper attributes
ds["cell_area"].attrs = dict(standard_name="cell_area", units="m2")
ds.air.attrs["cell_measures"] = "area: cell_area"
ds.air.cf.weighted("area").mean(["latitude", "time"]).cf.plot(x="longitude")
ds.air.mean(["lat", "time"]).cf.plot(x="longitude")
# ## Feature: Cell boundaries and vertices
#
# `cf_xarray` can infer cell boundaries (for rectilinear grids) and convert
# CF-standard bounds variables to vertices.
#
ds_bnds = ds.cf.add_bounds(["lat", "lon"])
ds_bnds
# We can also convert each bounds variable independently with the top-level
# functions
#
# +
lat_bounds = ds_bnds.cf.get_bounds("latitude")
lat_vertices = cfxr.bounds_to_vertices(lat_bounds, bounds_dim="bounds")
lat_vertices
# -
# Or we can convert _all_ bounds variables on a dataset
ds_crns = ds_bnds.cf.bounds_to_vertices()
ds_crns
# ## Feature: Add canonical CF attributes
#
# `cf_xarray` can add missing canonical CF attributes consistent with the official
# [CF standard name table](https://cfconventions.org/standard-names.html).
#
ds_canonical = ds.cf.add_canonical_attributes(verbose=True)
ds_canonical
| doc/examples/introduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="V58rxea0HqSa" colab={"base_uri": "https://localhost:8080/"} outputId="a9b62101-4f2e-4fc6-86ae-167b5754294d"
import os
# Find the latest version of spark 3.0 from http://www.apache.org/dist/spark/ and enter as the spark version
# For example:
# spark_version = 'spark-3.0.3'
spark_version = 'spark-3.0.3'
os.environ['SPARK_VERSION']=spark_version
# Install Spark and Java
# !apt-get update
# !apt-get install openjdk-11-jdk-headless -qq > /dev/null
# !wget -q http://www.apache.org/dist/spark/$SPARK_VERSION/$SPARK_VERSION-bin-hadoop2.7.tgz
# !tar xf $SPARK_VERSION-bin-hadoop2.7.tgz
# !pip install -q findspark
# Set Environment Variables
import os
os.environ["JAVA_HOME"] = "/usr/lib/jvm/java-11-openjdk-amd64"
os.environ["SPARK_HOME"] = f"/content/{spark_version}-bin-hadoop2.7"
# Start a SparkSession
import findspark
findspark.init()
# + id="_xKwTpATHqSe" colab={"base_uri": "https://localhost:8080/"} outputId="5ab64a10-f498-438c-a341-ff5fc079e004"
# Download the Postgres driver that will allow Spark to interact with Postgres.
# !wget https://jdbc.postgresql.org/download/postgresql-42.2.16.jar
# + id="MMqDAjVS0KN9"
from pyspark.sql import SparkSession
spark = SparkSession.builder.appName("M16-Amazon-Challenge").config("spark.driver.extraClassPath","/content/postgresql-42.2.16.jar").getOrCreate()
# + [markdown] id="cyBsySGuY-9V"
# ### Load Amazon Data into Spark DataFrame
# + id="CtCmBhQJY-9Z" colab={"base_uri": "https://localhost:8080/"} outputId="15da4527-c3a2-451f-b3c3-28b1ad88b7f8"
from pyspark import SparkFiles
url = "https://s3.amazonaws.com/amazon-reviews-pds/tsv/amazon_reviews_us_Software_v1_00.tsv.gz"
spark.sparkContext.addFile(url)
df = spark.read.option("encoding", "UTF-8").csv(SparkFiles.get(""), sep="\t", header=True, inferSchema=True)
df.show()
# + [markdown] id="2yUSe55VY-9t"
# ### Create DataFrames to match tables
# + id="C8REmY1aY-9u"
from pyspark.sql.functions import to_date
# Read in the Review dataset as a DataFrame
# + id="B0TESUDRY-90" colab={"base_uri": "https://localhost:8080/"} outputId="28675d19-d8c9-44df-d6be-1f5cd5c5f1e9"
# Create the customers_table DataFrame
customers_df = df.groupby("customer_id").count().withColumnRenamed("count", "customer_count")
customers_df.show()
# + id="4FwXA6UvY-96" colab={"base_uri": "https://localhost:8080/"} outputId="4d349584-bede-424f-eef9-002960e7f5a3"
# Create the products_table DataFrame and drop duplicates.
products_df = df.select(["product_id","product_title"]).drop_duplicates()
products_df.show()
# + id="MkqyCuNQY-9-" colab={"base_uri": "https://localhost:8080/"} outputId="d80103ca-3d17-4d9e-f1c1-a37b12359f44"
# Create the review_id_table DataFrame.
# Convert the 'review_date' column to a date datatype with to_date("review_date", 'yyyy-MM-dd').alias("review_date")
review_id_df = df.select(["review_id","customer_id","product_id","product_parent", to_date("review_date", 'yyyy-MM-dd').alias("review_date")])
review_id_df.show()
# + id="lzMmkdKmY--D" colab={"base_uri": "https://localhost:8080/"} outputId="950a299f-56ed-434a-9def-93b27cb6e9d6"
# Create the vine_table. DataFrame
vine_df = df.select(["review_id","star_rating","helpful_votes","total_votes","vine","verified_purchase"])
vine_df.show()
# + id="7jiUvs1aY--L"
from pyspark.sql.functions import col
# + id="T2zgZ-aKY--Q" colab={"base_uri": "https://localhost:8080/"} outputId="6dc964a4-c7dc-4351-db27-6abb30c91825"
filtered_df = df.filter(col("total_votes") >=10 )
filtered_df.show()
# + id="1m3yzn-LY--U" colab={"base_uri": "https://localhost:8080/"} outputId="ada5da7c-7b95-4515-9bc1-6d184f0f34a8"
vote_ratio_df = filtered_df.filter(col("helpful_votes")/col("total_votes")>=0.5)
vote_ratio_df.show()
# + id="KbXri15fY--Z" colab={"base_uri": "https://localhost:8080/"} outputId="5e42636e-24b5-4302-ae98-327e58f656ad"
paid_review_df = vote_ratio_df.filter(col("vine") == "Y")
paid_review_df.show()
# + id="XdQknSHLY--e" colab={"base_uri": "https://localhost:8080/"} outputId="d937a98e-065c-48bb-e435-829c53b95d5f"
unpaid_review_df = vote_ratio_df.filter(col("vine") == "N")
unpaid_review_df.show()
# + id="Exuo6ebUsCqW" colab={"base_uri": "https://localhost:8080/"} outputId="49342143-40e6-4c41-ce3a-7b565a00f103"
# PAID
# total reviews:
total_paid_reviews = paid_review_df.count()
print(total_paid_reviews)
# + colab={"base_uri": "https://localhost:8080/"} id="q4a_4tyb6etb" outputId="8ded3b4a-de29-4665-eceb-1ed7cfe6e975"
# 5 star reviews:
paid_five_star_reviews = paid_review_df.filter(col("star_rating")==5).count()
print(paid_five_star_reviews)
# + colab={"base_uri": "https://localhost:8080/"} id="o3hDLlOd9btf" outputId="c8554863-b26c-4d62-bed3-79b2bc7c742e"
# % 5 star reviews
paid_ratio = paid_five_star_reviews/total_paid_reviews
print(paid_ratio)
# + colab={"base_uri": "https://localhost:8080/"} id="A7ahCnlr9dHF" outputId="bf102078-50f3-4bdf-a6ca-6bcc8b52435f"
# UNPAID
# total reviews:
total_unpaid_reviews = unpaid_review_df.count()
print(total_unpaid_reviews)
# + colab={"base_uri": "https://localhost:8080/"} id="2JS_ahhB9ev9" outputId="7864a4b5-64ac-443b-ae83-5196ab68c705"
# 5 star reviews:
unpaid_five_star_reviews = unpaid_review_df.filter(col("star_rating")==5).count()
print(unpaid_five_star_reviews)
# + id="rwPtdrEc9gEd"
# % 5 star reviews
unpaid_ratio = unpaid_five_star_reviews/total_unpaid_reviews
print(unpaid_ratio)
| Vine_Review_Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] nbgrader={}
# # Project Euler: Problem 59
# + [markdown] nbgrader={}
# https://projecteuler.net/problem=59
#
# Each character on a computer is assigned a unique code and the preferred standard is ASCII (American Standard Code for Information Interchange). For example, uppercase A = 65, asterisk (*) = 42, and lowercase k = 107.
#
# A modern encryption method is to take a text file, convert the bytes to ASCII, then XOR each byte with a given value, taken from a secret key. The advantage with the XOR function is that using the same encryption key on the cipher text, restores the plain text; for example, 65 XOR 42 = 107, then 107 XOR 42 = 65.
#
# For unbreakable encryption, the key is the same length as the plain text message, and the key is made up of random bytes. The user would keep the encrypted message and the encryption key in different locations, and without both "halves", it is impossible to decrypt the message.
#
# Unfortunately, this method is impractical for most users, so the modified method is to use a password as a key. If the password is shorter than the message, which is likely, the key is repeated cyclically throughout the message. The balance for this method is using a sufficiently long password key for security, but short enough to be memorable.
#
# Your task has been made easy, as the encryption key consists of three lower case characters. Using cipher.txt (in this directory), a file containing the encrypted ASCII codes, and the knowledge that the plain text must contain common English words, decrypt the message and find the sum of the ASCII values in the original text.
# + [markdown] nbgrader={}
# The following cell shows examples of how to perform XOR in Python and how to go back and forth between characters and integers:
# + nbgrader={}
assert 65 ^ 42 == 107
assert 107 ^ 42 == 65
assert ord('a') == 97
assert chr(97) == 'a'
# + [markdown] nbgrader={}
# Certain functions in the `itertools` module may be useful for computing permutations:
# + nbgrader={}
import itertools
# + deletable=false nbgrader={"checksum": "6cff4e8e53b15273846c3aecaea84a3d", "solution": true}
#this cell brings the numbers from cipher.txt and appends them to a list as integers
stuff = open('cipher.txt', 'r')
i = stuff.read().split(',')
numbers=[]
for j in i:
numbers.append(int(j))
# -
#Made this encode function as prilimiary/test for my decode function
#works basically the same way as decode funtion
def encode(text, key):
encrypted=[]
count=0
for n in range(len(text)):
if len(key)<=count:
count=0
k=key[count]
n= text[n]
count+=1
encrypted.append(ord(n)^ord(k))
return (encrypted)
def decode(text, key):
message=[]
count=0
for n in range(len(text)):
#count starts at 0 and goes up by 1 for every character in key
#when count is greater than the length of key resets to 0
#will use count to call a specific letter in key
if len(key)<=count:
count=0
k = key[count]
n = text[n]
count += 1
#appends the corresponding character of: n XOR with correspoding the number of key
message.append(chr(n^ord(k)))
return (''.join(message))
#returns possible keys for the decryption
def find_key(text, keywords):
words=[]
#goes through all letters in alphebet
#uses chr to turn numbers to letters
#appends three letter words to list words
for x in range(97,123):
for y in range(97,123):
for z in range(97,123):
words.append(chr(x)+chr(y)+chr(z))
pos=[]
#takes the words in list words and puts them in the decode function
for key in words:
d=decode(text, key)
#goes through the input keywords
#if not in the decoded message break, if is append to pos
for w in keywords:
if w not in d:
break
else:
pos.append(key)
return (pos)
find_key(numbers, [' the ',])
TEXT = decode(numbers, 'god')
TEXT
#sums all the numbers corresponding to each character in the decoded message
sum(ord(i) for i in TEXT)
stuff.close()
# +
# YOUR CODE HERE
# + deletable=false nbgrader={"checksum": "dcdf6792a88c661545d3ca651212dba8", "grade": true, "grade_id": "projecteuler59", "points": 10}
# This cell will be used for grading, leave it at the end of the notebook.
| assignments/assignment02/ProjectEuler59.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !pip install opencv-python
# !pip install pytest-shutil
# ### Import libraries
import numpy as np
import pandas as pd
import cv2
import os
import tensorflow as tf
import keras
from keras.preprocessing.image import ImageDataGenerator
from keras.callbacks import TensorBoard, ModelCheckpoint
from keras.models import model_from_json
import random
from shutil import copyfile
tf.__version__
# ### Number of images in our Dataset
print("Number of images with mask used: ", len(os.listdir('../dataset/with_mask')))
print("Number of images without mask used: ", len(os.listdir('../dataset/without_mask')))
# ### Spliting the data into training and test set
def split_data(source, train, test, test_size):
data = os.listdir(source)
train_size = int(len(data)*(1 - test_size))
shuffled_data = random.sample(data, len(data))
training_data = shuffled_data[:train_size]
testing_data = shuffled_data[train_size:]
# make new folders train and test
for img in training_data:
temp_image = source+img
train_image = train+img
copyfile(temp_image, train_image)
for img in testing_data:
temp_image = source+img
test_image = test+img
copyfile(temp_image, test_image)
# +
MASK_SOURCE = '../dataset/with_mask/'
TRAIN_MASK_SOURCE = '../dataset/train/yes/'
TEST_MASK_SOURCE = '../dataset/test/yes/'
NO_MASK_SOURCE = '../dataset/without_mask/'
TEST_NO_MASK_SOURCE = '../dataset/test/no/'
TRAIN_NO_MASK_SOURCE = '../dataset/train/no/'
split_data(MASK_SOURCE, TRAIN_MASK_SOURCE, TEST_MASK_SOURCE, 0.2)
split_data(NO_MASK_SOURCE, TRAIN_NO_MASK_SOURCE, TEST_NO_MASK_SOURCE, 0.2)
# -
print('Number of training with mask: ', len(os.listdir(TRAIN_MASK_SOURCE)))
print('Number of training without mask: ', len(os.listdir(TRAIN_NO_MASK_SOURCE)))
print('Number of testing with mask: ', len(os.listdir(TEST_MASK_SOURCE)))
print('Number of testing without mask: ', len(os.listdir(TEST_NO_MASK_SOURCE)))
# ### Preprocessing data
# +
TRAIN_DIR = '../dataset/train/'
TEST_DIR = '../dataset/test/'
train_datagen = ImageDataGenerator(rescale = 1./255,
rotation_range=40,
width_shift_range=0.2,
height_shift_range=0.2,
shear_range = 0.2,
zoom_range = 0.2,
horizontal_flip = True)
training_set = train_datagen.flow_from_directory(TRAIN_DIR,
target_size = (150,150),
batch_size = 32,
class_mode = 'binary')
test_datagen = ImageDataGenerator(rescale = 1./255)
test_set = test_datagen.flow_from_directory(TEST_DIR,
target_size = (150,150),
batch_size = 32,
class_mode = 'binary')
# -
# ### Building CNN
# +
cnn = tf.keras.models.Sequential()
cnn.add(tf.keras.layers.Conv2D(filters=64, kernel_size=3, activation='relu', input_shape=[150, 150, 3]))
cnn.add(tf.keras.layers.MaxPool2D(pool_size=2, strides=2))
cnn.add(tf.keras.layers.Conv2D(filters=64, kernel_size=3, activation='relu'))
cnn.add(tf.keras.layers.MaxPool2D(pool_size=2, strides=2))
cnn.add(tf.keras.layers.Flatten())
cnn.add(tf.keras.layers.Dropout(0.5))
cnn.add(tf.keras.layers.Dense(units=128, activation='relu'))
cnn.add(tf.keras.layers.Dense(units=1, activation='sigmoid'))
# -
# ### Training the CNN
cnn.compile(optimizer = 'adam', loss = 'binary_crossentropy', metrics = ['accuracy'])
# Used to get best weights with monitored with validation loss
checkpoint = ModelCheckpoint('.ipynb_checkpoints/models_checkpoints/model-{epoch:03d}.model', monitor='val_loss', verbose=0, save_best_only=True, mode='auto')
# To train the modek on your computer and dataset
model = cnn.fit(x = training_set, validation_data = test_set, epochs = 30, callbacks=[checkpoint])
training_set.class_indices
# To save the model in your disk
'''
# serialize model to JSON
model_json = cnn.to_json()
with open("model.json", "w") as json_file:
json_file.write(model_json)
# serialize weights to HDF5
cnn.save_weights("model.h5")
print("Saved model to disk")
'''
# +
# Uncomment this to directly load the trained model if you don't wish to train it on your computer
# load json and create model
# json_file = open('model.json', 'r')
# loaded_model_json = json_file.read()
# json_file.close()
# loaded_model = model_from_json(loaded_model_json)
# load weights into new model
cnn.load_weights("model.h5")
print("Loaded model from disk")
# -
# ### Deploy the model on webcam input
# +
labels = ['No Mask :/', 'Mask On!', 'By <NAME>']
grid_color = [(0, 0, 255), (255, 0, 0)]
print("Press esc or q to quit")
# If you have multiple webcameras, feel free to change the source
# 0 is for default camera...
# 1 is for secondary camera...
webcam = cv2.VideoCapture(0)
classifier = cv2.CascadeClassifier('C:/Users/Shashank/anaconda3/Lib/site-packages/cv2/data/haarcascade_frontalface_default.xml')
try:
while webcam.isOpened():
_, frame = webcam.read()
frame = cv2.flip(frame, 1, 1)
faces = classifier.detectMultiScale(frame, 1.1, 4)
for (x,y,w,h) in faces:
face = frame[y-40:y+h+20, x-20:x+w+20]
try:
resized_data = (cv2.resize(face, (150, 150)))/255.0
except Exception as e:
resized_data = (cv2.resize(frame, (150, 150)))/255.0
final_data = np.expand_dims(resized_data, axis = 0)
prediction = cnn.predict(final_data)
answer = prediction[0][0] #Binary answer 0 = no, 1 = yes
# print(answer)
answer = int(answer < 0.2) #Can be changed according to your picture
# display the answer
cv2.rectangle(frame, (x,y), (x+w, y+h), grid_color[answer], 3)
cv2.rectangle(frame, (x-2, y-45), (x+w,y), grid_color[answer], -1)
cv2.putText(frame, labels[answer], (x,y-10), cv2.FONT_HERSHEY_SIMPLEX, 1, (255, 255, 255), 1)
cv2.putText(frame, labels[2], (250 ,470), cv2.FONT_HERSHEY_DUPLEX, 1, (0, 0, 0), 1)
cv2.imshow("MASK DETECTOR - by <NAME>", frame)
key = cv2.waitKey(1)
if key == 27 or (key & 0xFF == ord('q')):
break
except:
raise
finally:
webcam.release()
cv2.destroyAllWindows()
# -
# ### Test Without Webcam
# +
from keras.preprocessing import image
test_image = image.load_img('test_data/img4.jpg', target_size = (150, 150))
test_image = image.img_to_array(test_image)
test_image = np.expand_dims(test_image, axis = 0)
result = cnn.predict(test_image)
print(result[0][0])
res = "Mask on" if (result[0][0]>0.4) else "Mask off"
print(res)
| detector.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# *Accompanying code examples of the book "Introduction to Artificial Neural Networks and Deep Learning: A Practical Guide with Applications in Python" by [<NAME>](https://sebastianraschka.com). All code examples are released under the [MIT license](https://github.com/rasbt/deep-learning-book/blob/master/LICENSE). If you find this content useful, please consider supporting the work by buying a [copy of the book](https://leanpub.com/ann-and-deeplearning).*
#
# Other code examples and content are available on [GitHub](https://github.com/rasbt/deep-learning-book). The PDF and ebook versions of the book are available through [Leanpub](https://leanpub.com/ann-and-deeplearning).
# %load_ext watermark
# %watermark -a '<NAME>' -v -p torch
# - Runs on CPU or GPU (if available)
# # Model Zoo -- Convolutional Autoencoder with Deconvolutions (without pooling operations)
# A convolutional autoencoder using deconvolutional layers that compresses 768-pixel MNIST images down to a 7x7x8 (392 pixel) representation without using pooling operations but increasing the stride in convolutional layers.
# ## Imports
import numpy as np
import torch
import torch.nn.functional as F
from torch.utils.data import DataLoader
from torchvision import datasets
from torchvision import transforms
# +
##########################
### SETTINGS
##########################
# Device
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
print('Device:', device)
# Hyperparameters
random_seed = 123
learning_rate = 0.05
num_epochs = 10
batch_size = 128
##########################
### MNIST DATASET
##########################
# Note transforms.ToTensor() scales input images
# to 0-1 range
train_dataset = datasets.MNIST(root='data',
train=True,
transform=transforms.ToTensor(),
download=True)
test_dataset = datasets.MNIST(root='data',
train=False,
transform=transforms.ToTensor())
train_loader = DataLoader(dataset=train_dataset,
batch_size=batch_size,
shuffle=True)
test_loader = DataLoader(dataset=test_dataset,
batch_size=batch_size,
shuffle=False)
# Checking the dataset
for images, labels in train_loader:
print('Image batch dimensions:', images.shape)
print('Image label dimensions:', labels.shape)
break
# -
# ## Model
# +
##########################
### MODEL
##########################
class ConvolutionalAutoencoder(torch.nn.Module):
def __init__(self):
super(ConvolutionalAutoencoder, self).__init__()
# calculate same padding:
# (w - k + 2*p)/s + 1 = o
# => p = (s(o-1) - w + k)/2
### ENCODER
# 28x28x1 => 14x14x4
self.conv_1 = torch.nn.Conv2d(in_channels=1,
out_channels=4,
kernel_size=(3, 3),
stride=(2, 2),
# floor((2(14-1) - 28 + 3) / 2) = 0
padding=0)
# 14x14x4 => 7x7x8
self.conv_2 = torch.nn.Conv2d(in_channels=4,
out_channels=8,
kernel_size=(3, 3),
stride=(2, 2),
# ceil((2(7-1) - 14 + 3) / 2) = 1
padding=1)
### DECODER
# 7x7x8 => 15x15x4
self.deconv_1 = torch.nn.ConvTranspose2d(in_channels=8,
out_channels=4,
kernel_size=(3, 3),
stride=(2, 2),
padding=0)
# 15x15x4 => 29x29x1
self.deconv_2 = torch.nn.ConvTranspose2d(in_channels=4,
out_channels=1,
kernel_size=(3, 3),
stride=(2, 2),
padding=1)
def forward(self, x):
### ENCODER
x = self.conv_1(x)
x = F.leaky_relu(x)
x = self.conv_2(x)
x = F.leaky_relu(x)
### DECODER
x = self.deconv_1(x)
x = F.leaky_relu(x)
x = self.deconv_2(x)
x = F.leaky_relu(x)
x = x[:, :, :-1, :-1]
x = F.sigmoid(x)
return x
torch.manual_seed(random_seed)
model = ConvolutionalAutoencoder()
model = model.to(device)
##########################
### COST AND OPTIMIZER
##########################
cost_fn = torch.nn.BCELoss() # torch.nn.MSELoss()
optimizer = torch.optim.Adam(model.parameters(), lr=learning_rate)
# -
# ## Training
for epoch in range(num_epochs):
for batch_idx, (features, targets) in enumerate(train_loader):
# don't need labels, only the images (features)
features = features.to(device)
### FORWARD AND BACK PROP
decoded = model(features)
cost = cost_fn(decoded, features)
optimizer.zero_grad()
cost.backward()
### UPDATE MODEL PARAMETERS
optimizer.step()
### LOGGING
if not batch_idx % 50:
print ('Epoch: %03d/%03d | Batch %03d/%03d | Cost: %.4f'
%(epoch+1, num_epochs, batch_idx,
len(train_dataset)//batch_size, cost))
# ## Evaluation
# +
# %matplotlib inline
import matplotlib.pyplot as plt
##########################
### VISUALIZATION
##########################
n_images = 15
image_width = 28
fig, axes = plt.subplots(nrows=2, ncols=n_images,
sharex=True, sharey=True, figsize=(20, 2.5))
orig_images = features[:n_images]
decoded_images = decoded[:n_images]
for i in range(n_images):
for ax, img in zip(axes, [orig_images, decoded_images]):
ax[i].imshow(img[i].detach().reshape((image_width, image_width)), cmap='binary')
| code/model_zoo/pytorch_ipynb/autoencoder-deconv-2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/mrk-W2D1/tutorials/W2D1_BayesianStatistics/W2D1_Tutorial4.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="ICwW1AANyqVc"
# # NMA 2020 W2D1 -- (Bonus) Tutorial 4: Bayesian Decision Theory & Cost functions
# __Content creators:__ <NAME>, <NAME>, with help from <NAME>
#
# __Content reviewers:__ <NAME>, <NAME>, <NAME>, <NAME>
# + [markdown] colab_type="text" id="LqkrbIEeyqVl"
# # Tutorial Objectives
#
# *This tutorial is optional! Please do not feel pressured to finish it!*
#
# In the previous tutorials, we investigated the posterior, which describes beliefs based on a combination of current evidence and prior experience. This tutorial focuses on Bayesian Decision Theory, which combines the posterior with **cost functions** that allow us to quantify the potential impact of making a decision or choosing an action based on that posterior. Cost functions are therefore critical for turning probabilities into actions!
#
# In Tutorial 3, we used the mean of the posterior $p(x | \tilde x)$ as a proxy for the response $\hat x$ for the participants. What prompted us to use the mean of the posterior as a **decision rule**? In this tutorial we will see how different common decision rules such as the choosing the mean, median or mode of the posterior distribution correspond to minimizing different cost functions.
#
# In this tutorial, you will
# 1. Implement three commonly-used cost functions: mean-squared error, absolute error, and zero-one loss
# 2. Discover the concept of expected loss, and
# 3. Choose optimal locations on the posterior that minimize these cost functions. You will verify that it these locations can be found analytically as well as empirically.
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 516} colab_type="code" id="D_VO3CyRFCZm" outputId="fc1312ab-18b4-4e7e-8902-3afc7cdabb00"
#@title Video 1: Introduction
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id='BV1Tv411q77s', width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
video
# + [markdown] colab_type="text" id="jBNUP5ReEiug"
# ---
# Please execute the cell below to initialize the notebook environment
#
# + [markdown] colab_type="text" id="RgiqE5SlEl69"
# ---
# ### Setup
# + cellView="both" colab={} colab_type="code" id="JkdIcrE1yqVd"
# Imports
import numpy as np
import matplotlib.pyplot as plt
# + cellView="form" colab={} colab_type="code" id="Ba1nILlhFcwu"
#@title Figure Settings
import ipywidgets as widgets
plt.style.use("/share/dataset/COMMON/nma.mplstyle.txt")
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
# + cellView="form" colab={} colab_type="code" id="WwWZ3iPfE0Mi"
# @title Helper Functions
def my_gaussian(x_points, mu, sigma):
"""Returns un-normalized Gaussian estimated at points `x_points`
DO NOT EDIT THIS FUNCTION !!!
Args :
x_points (numpy array of floats) - points at which the gaussian is evaluated
mu (scalar) - mean of the Gaussian
sigma (scalar) - std of the gaussian
Returns:
(numpy array of floats): un-normalized Gaussian (i.e. without constant) evaluated at `x`
"""
return np.exp(-(x_points-mu)**2/(2*sigma**2))
def visualize_loss_functions(mse=None, abse=None, zero_one=None):
"""Visualize loss functions
Args:
- mse (func) that returns mean-squared error
- abse: (func) that returns absolute_error
- zero_one: (func) that returns zero-one loss
All functions should be of the form f(x, x_hats). See Exercise #1.
Returns:
None
"""
x = np.arange(-3, 3.25, 0.25)
fig, ax = plt.subplots(1)
if mse is not None:
ax.plot(x, mse(0, x), linewidth=2, label="Mean Squared Error")
if abse is not None:
ax.plot(x, abse(0, x), linewidth=2, label="Absolute Error")
if zero_one_loss is not None:
ax.plot(x, zero_one_loss(0, x), linewidth=2, label="Zero-One Loss")
ax.set_ylabel('Cost')
ax.set_xlabel('Predicted Value ($\hat{x}$)')
ax.set_title("Loss when the true value $x$=0")
ax.legend()
plt.show()
def moments_myfunc(x_points, function):
"""Returns the mean, median and mode of an arbitrary function
DO NOT EDIT THIS FUNCTION !!!
Args :
x_points (numpy array of floats) - x-axis values
function (numpy array of floats) - y-axis values of the function evaluated at `x_points`
Returns:
(tuple of 3 scalars): mean, median, mode
"""
# Calc mode of an arbitrary function
mode = x_points[np.argmax(function)]
# Calc mean of an arbitrary function
mean = np.sum(x_points * function)
# Calc median of an arbitrary function
cdf_function = np.zeros_like(x_points)
accumulator = 0
for i in np.arange(x.shape[0]):
accumulator = accumulator + posterior[i]
cdf_function[i] = accumulator
idx = np.argmin(np.abs(cdf_function - 0.5))
median = x_points[idx]
return mean, median, mode
def loss_plot(x, loss, min_loss, loss_label, show=False, ax=None):
if not ax:
fig, ax = plt.subplots()
ax.plot(x, loss, '-r', linewidth=2, label=loss_label)
ax.axvline(min_loss, ls='dashed', color='red', label='Minimum')
ax.set_ylabel('Expected Loss')
ax.set_xlabel('Orientation (Degrees)')
ax.legend()
if show:
plt.show()
def loss_plot_subfigures(x,
MSEloss, min_MSEloss, loss_MSElabel,
ABSEloss, min_ABSEloss, loss_ABSElabel,
ZeroOneloss, min_01loss, loss_01label):
fig_w, fig_h = plt.rcParams.get('figure.figsize')
fig, ax = plt.subplots(nrows=2, ncols=3, figsize=(fig_w*2, fig_h*2), sharex=True)
ax[0, 0].plot(x, MSEloss, '-r', linewidth=2, label=loss_MSElabel)
ax[0, 0].axvline(min_MSEloss, ls='dashed', color='red', label='Minimum')
ax[0, 0].set_ylabel('Expected Loss')
ax[0, 0].set_xlabel('Orientation (Degrees)')
ax[0, 0].set_title("Mean Squared Error")
ax[0, 0].legend()
pmoments_plot(x, posterior, ax=ax[1,0])
ax[0, 1].plot(x, ABSEloss, '-b', linewidth=2, label=loss_ABSElabel)
ax[0, 1].axvline(min_ABSEloss, ls='dashdot', color='blue', label='Minimum')
ax[0, 1].set_ylabel('Expected Loss')
ax[0, 1].set_xlabel('Orientation (Degrees)')
ax[0, 1].set_title("Absolute Error")
ax[0, 1].legend()
pmoments_plot(x, posterior, ax=ax[1,1])
ax[0, 2].plot(x, ZeroOneloss, '-g', linewidth=2, label=loss_01label)
ax[0, 2].axvline(min_01loss, ls='dotted', color='green', label='Minimum')
ax[0, 2].set_ylabel('Expected Loss')
ax[0, 2].set_xlabel('Orientation (Degrees)')
ax[0, 2].set_title("0-1 Loss")
ax[0, 2].legend()
pmoments_plot(x, posterior, ax=ax[1,2])
plt.show()
def pmoments_plot(x, posterior,
prior=None, likelihood=None, show=False, ax=None):
if not ax:
fig, ax = plt.subplots()
if prior:
ax.plot(x, prior, '-r', linewidth=2, label='Prior')
if likelihood:
ax.plot(x, likelihood, '-b', linewidth=2, label='Likelihood')
ax.plot(x, posterior, '-g', linewidth=4, label='Posterior')
mean, median, mode = moments_myfunc(x, posterior)
ax.axvline(mean, ls='dashed', color='red', label='Mean')
ax.axvline(median, ls='dashdot', color='blue', label='Median')
ax.axvline(mode, ls='dotted', color='green', label='Mode')
ax.set_ylabel('Probability')
ax.set_xlabel('Orientation (Degrees)')
ax.legend()
if show:
plt.show()
def generate_example_pdfs():
"""Generate example probability distributions as in T2"""
x=np.arange(-5, 5, 0.01)
prior_mean = 0
prior_sigma1 = .5
prior_sigma2 = 3
prior1 = my_gaussian(x, prior_mean, prior_sigma1)
prior2 = my_gaussian(x, prior_mean, prior_sigma2)
alpha = 0.05
prior_combined = (1-alpha) * prior1 + (alpha * prior2)
prior_combined = prior_combined / np.sum(prior_combined)
likelihood_mean = -2.7
likelihood_sigma = 1
likelihood = my_gaussian(x, likelihood_mean, likelihood_sigma)
likelihood = likelihood / np.sum(likelihood)
posterior = prior_combined * likelihood
posterior = posterior / np.sum(posterior)
return x, prior_combined, likelihood, posterior
def plot_posterior_components(x, prior, likelihood, posterior):
with plt.xkcd():
fig = plt.figure()
plt.plot(x, prior, '-r', linewidth=2, label='Prior')
plt.plot(x, likelihood, '-b', linewidth=2, label='Likelihood')
plt.plot(x, posterior, '-g', linewidth=4, label='Posterior')
plt.legend()
plt.title('Sample Output')
plt.show()
# + [markdown] colab_type="text" id="6MuqNavvFU-3"
# ### The Posterior Distribution
#
# This notebook will use a model similar to the puppet & puppeteer sound experiment developed in Tutorial 2, but with different probabilities for $p_{common}$, $p_{independent}$, $\sigma_{common}$ and $\sigma_{independent}$. Specifically, our model will consist of these components, combined according to Bayes' rule:
# $$
# \begin{eqnarray}
# \textrm{Prior} &=& \begin{cases} \mathcal{N_{common}}(0, 0.5) & 95\% \textrm{ weight}\\
# \mathcal{N_{independent}}(0, 3.0) & 5\% \textrm{ weight} \\
# \end{cases}\\\\
# \textrm{Likelihood} &= &\mathcal{N}(-2.7, 1.0)
# \end{eqnarray}
# $$
#
# We will use this posterior as an an example through this notebook. Please run the cell below to import and plot the model. You do not need to edit anything. These parameter values were deliberately chosen for illustration purposes: there is nothing intrinsically special about them, but they make several of the exercises easier.
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 288} colab_type="code" id="ePqZJTqwLkbV" outputId="115c977f-41f9-4de3-9639-ddcec695eb73"
x, prior, likelihood, posterior = generate_example_pdfs()
plot_posterior_components(x, prior, likelihood, posterior)
# + [markdown] colab_type="text" id="W9-0xA8du5-N"
# # Section 1: The Cost Functions
#
# Next, we will implement the cost functions.
# A cost function determines the "cost" (or penalty) of estimating $\hat{x}$ when the true or correct quantity is really $x$ (this is essentially the cost of the error between the true stimulus value: $x$ and our estimate: $\hat x$ -- Note that the error can be defined in different ways):
#
# $$\begin{eqnarray}
# \textrm{Mean Squared Error} &=& (x - \hat{x})^2 \\
# \textrm{Absolute Error} &=& \big|x - \hat{x}\big| \\
# \textrm{Zero-One Loss} &=& \begin{cases}
# 0,& \text{if } x = \hat{x} \\
# 1, & \text{otherwise}
# \end{cases}
# \end{eqnarray}
# $$
#
# In the cell below, fill in the body of these cost function. Each function should take one single value for $x$ (the true stimulus value : $x$) and one or more possible value estimates: $\hat{x}$.
#
# Return an array containing the costs associated with predicting $\hat{x}$ when the true value is $x$. Once you have written all three functions, uncomment the final line to visulize your results.
#
# _Hint:_ These functions are easy to write (1 line each!) but be sure *all* three functions return arrays of `np.float` rather than another data type.
# + cellView="code" colab={} colab_type="code" id="qGJxyXba0EwP"
def mse(x, x_hats):
"""Mean-squared error cost function
Args:
x (scalar): One true value of $x$
x_hats (scalar or ndarray): Estimate of x
Returns:
same shape/type as x_hats): MSE costs associated with
predicting x_hats instead of x$
"""
##############################################################################
# Complete the MSE cost function
#
### Comment out the line below to test your function
raise NotImplementedError("You need to complete the MSE cost function!")
##############################################################################
my_mse = ...
return my_mse
def abs_err(x, x_hats):
"""Absolute error cost function
Args:
x (scalar): One true value of $x$
x_hats (scalar or ndarray): Estimate of x
Returns:
(same shape/type as x_hats): absolute error costs associated with
predicting x_hats instead of x$
"""
##############################################################################
# Complete the absolute error cost function
#
### Comment out the line below to test your function
raise NotImplementedError("You need to complete the absolute error function!")
##############################################################################
my_abs_err = ...
return my_abs_err
def zero_one_loss(x, x_hats):
"""Zero-One loss cost function
Args:
x (scalar): One true value of $x$
x_hats (scalar or ndarray): Estimate of x
Returns:
(same shape/type as x_hats) of the 0-1 Loss costs associated with predicting x_hat instead of x
"""
##############################################################################
# Complete the zero-one loss cost function
#
### Comment out the line below to test your function
raise NotImplementedError("You need to complete the 0-1 loss cost function!")
##############################################################################
my_zero_one_loss = ...
return my_zero_one_loss
## When you are done with the functions above, uncomment the line below to
## visualize them
# visualize_loss_functions(mse, abs_err, zero_one_loss)
# + colab={"base_uri": "https://localhost:8080/", "height": 288} colab_type="code" id="-Hfi7F135Uku" outputId="653b40a4-ef6e-4b64-e76b-33ced28c7464"
# to_remove solution
def mse(x, x_hats):
"""Mean-squared error cost function
Args:
x (scalar): One true value of $x$
x_hats (scalar or ndarray): Estimate of x
Returns:
same shape/type as x_hats): MSE costs associated with
predicting x_hats instead of x$
"""
##############################################################################
# Complete the MSE cost function
#
### Comment out the line below to test your function
#raise NotImplementedError("You need to complete the MSE cost function!")
##############################################################################
my_mse = (x - x_hats)**2
return my_mse
def abs_err(x, x_hats):
"""Absolute error cost function
Args:
x (scalar): One true value of $x$
x_hats (scalar or ndarray): Estimate of x
Returns:
(same shape/type as x_hats): absolute error costs associated with
predicting x_hats instead of x$
"""
##############################################################################
# Complete the absolute error cost function
#
### Comment out the line below to test your function
#raise NotImplementedError("You need to complete the absolute error function!")
##############################################################################
my_abs_err = np.abs(x - x_hats)
return my_abs_err
def zero_one_loss(x, x_hats):
"""Zero-One loss cost function
Args:
x (scalar): One true value of $x$
x_hats (scalar or ndarray): Estimate of x
Returns:
(same shape/type as x_hats) of the 0-1 Loss costs associated with predicting x_hat instead of x
"""
##############################################################################
# Complete the zero-one loss cost function
#
### Comment out the line below to test your function
#raise NotImplementedError("You need to complete the 0-1 loss cost function!")
##############################################################################
my_zero_one_loss = (x != x_hats).astype(np.float)
return my_zero_one_loss
## When you are done with the functions above, uncomment the line below to
## visualize them
with plt.xkcd():
visualize_loss_functions(mse, abs_err, zero_one_loss)
# + [markdown] colab_type="text" id="I0um5kYNJ53x"
# # Section 2: Expected Loss
#
#
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 516} colab_type="code" id="1bJb-K-TKlq9" outputId="2ffa96fa-5dca-4bc2-a483-73dde742c8af"
#@title Video 2: Expected Loss
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id='BV1av411q7iK', width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
video
# + [markdown] colab_type="text" id="QEsxcge9KkxZ"
# A posterior distribution tells us about the confidence or credibility we assign to different choices. A cost function describes the penalty we incur when choosing an incorrect option. These concepts can be combined into an *expected loss* function. Expected loss is defined as:
#
# $$
# \begin{eqnarray}
# \mathbb{E}[\text{Loss} | \hat{x}] = \int L[\hat{x},x] \odot p(x|\tilde{x}) dx
# \end{eqnarray}
# $$
#
# where $L[ \hat{x}, x]$ is the loss function, $p(x|\tilde{x})$ is the posterior, and $\odot$ represents the [Hadamard Product](https://en.wikipedia.org/wiki/Hadamard_product_(matrices)) (i.e., elementwise multiplication), and $\mathbb{E}[\text{Loss} | \hat{x}]$ is the expected loss.
#
# In this exercise, we will calculate the expected loss for the: means-squared error, the absolute error, and the zero-one loss over our bimodal posterior $p(x | \tilde x)$.
#
# **Suggestions:**
# * We already pre-completed the code (commented-out) to calculate the mean-squared error, absolute error, and zero-one loss between $x$ and an estimate $\hat x$ using the functions you created in exercise 1
# * Calculate the expected loss ($\mathbb{E}[MSE Loss]$) using your posterior (imported above as `posterior`) & each of the loss functions described above (MSELoss, ABSELoss, and Zero-oneLoss).
# * Find the x position that minimizes the expected loss for each cost function and plot them using the `loss_plot` function provided (commented-out)
# + [markdown] colab_type="text" id="tKuLHqaS6GAX"
# ## Exercise 2: Finding the expected loss empirically via integration
# + cellView="code" colab={} colab_type="code" id="oVJnNYDBJ6rB"
def expected_loss_calculation(x, posterior):
ExpectedLoss_MSE = np.zeros_like(x)
ExpectedLoss_ABSE = np.zeros_like(x)
ExpectedLoss_01 = np.zeros_like(x)
for idx in np.arange(x.shape[0]):
estimate = x[idx]
###################################################################
## Insert code below to find the expected loss under each loss function
##
## remove the raise when the function is complete
raise NotImplementedError("Calculate the expected loss over all x values!")
###################################################################
MSELoss = mse(estimate, x)
ExpectedLoss_MSE[idx] = ...
ABSELoss = abs_err(estimate, x)
ExpectedLoss_ABSE[idx] = ...
ZeroOneLoss = zero_one_loss(estimate, x)
ExpectedLoss_01[idx] = ...
###################################################################
## Now, find the `x` location that minimizes expected loss
##
## remove the raise when the function is complete
raise NotImplementedError("Finish the Expected Loss calculation")
###################################################################
min_MSE = ...
min_ABSE = ...
min_01 = ...
return (ExpectedLoss_MSE, ExpectedLoss_ABSE, ExpectedLoss_01,
min_MSE, min_ABSE, min_01)
## Uncomment the lines below to plot the expected loss as a function of the estimates
#ExpectedLoss_MSE, ExpectedLoss_ABSE, ExpectedLoss_01, min_MSE, min_ABSE, min_01 = expected_loss_calculation(x, posterior)
#loss_plot(x, ExpectedLoss_MSE, min_MSE, f"Mean Squared Error = {min_MSE:.2f}")
#loss_plot(x, ExpectedLoss_ABSE, min_ABSE, f"Absolute Error = {min_ABSE:.2f}")
#loss_plot(x, ExpectedLoss_01, min_01, f"Zero-One Error = {min_01:.2f}")
# + colab={"base_uri": "https://localhost:8080/", "height": 889} colab_type="code" id="jwNbpZo7J6rF" outputId="eb8c2f16-5092-48de-9d02-5d07c683f27a"
# to_remove solution
def expected_loss_calculation(x, posterior):
ExpectedLoss_MSE = np.zeros_like(x)
ExpectedLoss_ABSE = np.zeros_like(x)
ExpectedLoss_01 = np.zeros_like(x)
for idx in np.arange(x.shape[0]):
estimate = x[idx]
###################################################################
## Insert code below to find the expected loss under each loss function
##
## remove the raise when the function is complete
#raise NotImplementedError("Calculate the expected loss over all x values!")
###################################################################
MSELoss = mse(estimate, x)
ExpectedLoss_MSE[idx] = np.sum(MSELoss * posterior)
ABSELoss = abs_err(estimate, x)
ExpectedLoss_ABSE[idx] = np.sum(ABSELoss * posterior)
ZeroOneLoss = zero_one_loss(estimate, x)
ExpectedLoss_01[idx] = np.sum(ZeroOneLoss * posterior)
###################################################################
## Now, find the `x` location that minimizes expected loss
##
## remove the raise when the function is complete
# raise NotImplementedError("Finish the Expected Loss calculation")
###################################################################
min_MSE = x[np.argmin(ExpectedLoss_MSE)]
min_ABSE = x[np.argmin(ExpectedLoss_ABSE)]
min_01 = x[np.argmin(ExpectedLoss_01)]
return (ExpectedLoss_MSE, ExpectedLoss_ABSE, ExpectedLoss_01,
min_MSE, min_ABSE, min_01)
## Uncomment the lines below to plot the expected loss as a function of the estimates
ExpectedLoss_MSE, ExpectedLoss_ABSE, ExpectedLoss_01, min_MSE, min_ABSE, min_01 = expected_loss_calculation(x, posterior)
with plt.xkcd():
loss_plot(x, ExpectedLoss_MSE, min_MSE, f"Mean Squared Error = {min_MSE:.2f}")
loss_plot(x, ExpectedLoss_ABSE, min_ABSE, f"Absolute Error = {min_ABSE:.2f}")
loss_plot(x, ExpectedLoss_01, min_01, f"Zero-One Error = {min_01:.2f}")
# + [markdown] colab_type="text" id="Drp7PBsZJ6q_"
# # Section 3: Analytical Solutions
#
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 516} colab_type="code" id="OfOI-83LYkf1" outputId="57ed04b5-e5c1-4b05-cc13-b8a4fa574e23"
#@title Video 3: Analytical Solutions
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id='BV1aa4y1a7Ex', width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
video
# + [markdown] colab_type="text" id="doazsK-CYleY"
# In the previous exercise, we found the minimum expected loss via brute-force: we searched over all possible values of $x$ and found the one that minimized each of our loss functions. This is feasible for our small toy example, but can quickly become intractable.
#
# Fortunately, the three loss functions examined in this tutorial have are minimized at specific points on the posterior, corresponding to the itss mean, median, and mode. To verify this property, we have replotted the loss functions from Exercise 2 below, with the posterior on the same scale beneath. The mean, median, and mode are marked on the posterior.
#
# Which loss form corresponds to each summary statistics?
#
# + colab={"base_uri": "https://localhost:8080/", "height": 583} colab_type="code" id="ZygmcypY7xzY" outputId="df84bd3d-92d2-46fe-c496-1986155db1ba"
loss_plot_subfigures(x,
ExpectedLoss_MSE, min_MSE, f"Mean Squared Error = {min_MSE:.2f}",
ExpectedLoss_ABSE, min_ABSE, f"Absolute Error = {min_ABSE:.2f}",
ExpectedLoss_01, min_01, f"Zero-One Error = {min_01:.2f}")
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" id="oYDbzbTYPEO0" outputId="80fc99eb-3290-4c5b-ada3-58ace07f30d8"
#to_remove explanation
"""
As you might recall from W1D3, the mean minimizes the mean-squared error.
Absolute error is minimized by the median, while zero-one loss is minimized
at the posterior's mode.
"""
# + [markdown] colab_type="text" id="lzYSxCRuyqVw"
# # Section 4: Conclusion
#
# + cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 516} colab_type="code" id="uQvIJQ7Xb1CD" outputId="46885803-b2ad-4ec2-c7b8-ed6d1eef999b"
#@title Video 4: Outro
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id='BV1kh411o7cu', width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
video
# + [markdown] colab_type="text" id="2TjHQTj5b2rJ"
# In this tutorial, we learned about three kinds of cost functions: mean-squared error, absolute error, and zero-one loss. We used expected loss to quantify the results of making a decision, and showed that optimizing under different cost functions led us to choose different locations on the posterior. Finally, we found that these optimal locations can be identified analytically, sparing us from a brute-force search.
#
# Here are some additional questions to ponder:
# * Suppose your professor offered to grade your work with a zero-one loss or mean square error.
# * When might you choose each?
# * Which would be easier to learn from?
# * All of the loss functions we considered are symmetrical. Are there situations where an asymmetrical loss function might make sense? How about a negative one?
#
#
#
#
#
#
#
#
| tutorials/W2D1_BayesianStatistics/W2D1_Tutorial4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Week 6 Homework Notebook
# This is to use for python homework practice
#
# ### Description
# Use plotly to produce examples or solve homework problems for week 6.
#
# pip install plotly first if you havent already done so.
import plotly.plotly
from plotly.graph_objs import Scatter, Layout
from numpy import arange, cos
import numpy as np
print('imports completed')
# +
# Copied from example Module 1
def g(x):
return x*x # This is the function in Lial Section 11.1 Example 1.
'''
The numbers chosen below here are somewhat arbitrary to show what the limit is. Think about the chart on
page 266 and graph 267. Break it into discrete confided steps. Focus on learning the idea of a limit, getting closer
and closer without actually reaching a number.
'''
n = 5
powers = arange(0, n+1)
print("powers: {}".format(powers))
denominator = 2.0**powers
print("denominator: {}".format(denominator))
delta = 2.0
'''
This determines the number of values calculated on each side of x=0.
denominator contains exponentiated values of 2.0. # This is the interval used on either side of x=2.0.
The following are values of x and f(x) trending to the limit at x=2.0.
Delta is being divided by powers of 2 to reduce the distance from the limit.
'''
x_r = 2.0+delta/denominator # Approaching from the right.
print("x_r: {}".format(x_r))
y_r = g(x_r)
print("y_r: {}".format(y_r))
x_l = 2.0-delta/denominator # Approaching from the left.
print("x_l: {}".format(x_l))
y_l = g(x_l)
print("y_l: {}".format(y_l))
# The following determine the vertical boundaries of the resulting plot.
# +
#Required for displaying plotly in jupyter notebook
plotly.offline.init_notebook_mode(connected=True)
# Create traces
trace1 = Scatter(x=x_r, y=y_r, name='right', line=dict(color='#bc42f4'))
trace2 = Scatter(x=x_l, y=y_l, name='left', line=dict(color='#41f1f4'))
point = Scatter(x=[2.0], y=[g(2.0)], name='point', mode='markers')
plotly.offline.iplot({
"data": [trace1, trace2, point],
"layout": Layout(title="Convergence Example")
})
# +
# Copied directly from example Modeule 2
'''
Math for Modelers Session #6 Python Module #2
Reading assignment:
"Think Python" 2nd Edition Chapter 7 (7.1-7.7)
“Think Python” 3rd Edition Chapter 7 (pages 75-81)
Module #2 objectives: 1) demonstrate numerical differentialtion,
and 2) illustrate results graphically.
A general function for calculating the slope between two points: x and
x+delta. See Lial Section 11.3 dealing with instantaneous rates of change.
'''
# Define a function for demonstration. This function may be changed.
def f(x):
return cos(x)
point = 1.0 # This is a point at which a derivative will be calculated.
# Calculate values for the tangent.
w = arange(point - 1.0, point + 1.1, 0.1)
t = f(point) + limit * (w - point)
# Now we are going to plot the original function over a wider range.
# Define a domain for the function.
domain = 3.14
# Calculate values for the function on both sides of x=1.0.
u = arange(point - domain, point + domain + 0.1, 0.1)
z = f(u)
#Required for displaying plotly in jupyter notebook
plotly.offline.init_notebook_mode(connected=True)
# Create traces
tangent = Scatter(x=w, y=t, name='tangent')
curve = Scatter(x=u, y=z, name='curve')
plotly.offline.iplot({
"data": [tangent, curve],
"layout": Layout(title="Tangent at a point")
})
# -
| Wk6/Wk6 Homework.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Auto Encoder with MNIST (or Fashion MNIST)
#
# * This code is available to tensorflow version 2.0
# * Implemented by [`tf.keras.layers`](https://www.tensorflow.org/versions/r2.0/api_docs/python/tf/keras/layers) [`tf.losses`](https://www.tensorflow.org/versions/r2.0/api_docs/python/tf/losses)
# ## Import modules
# +
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
from __future__ import unicode_literals
import os
import sys
import time
import glob
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import PIL
import imageio
from IPython import display
import tensorflow as tf
from tensorflow.keras import layers
sys.path.append(os.path.dirname(os.path.abspath('.')))
from utils.image_utils import *
from utils.ops import *
os.environ["CUDA_VISIBLE_DEVICES"]="0"
# -
# ## Setting hyperparameters
# +
# Training Flags (hyperparameter configuration)
model_name = 'ae'
train_dir = os.path.join('train', model_name, 'exp1')
dataset_name = 'fashion_mnist'
assert dataset_name in ['mnist', 'fashion_mnist']
max_epochs = 30
save_model_epochs = 5
print_steps = 1000
save_images_epochs = 1
batch_size = 128
learning_rate = 1e-4
num_examples_to_generate = 16
x_dim = 784
latent_dim = 200
# -
# ## Load the MNIST dataset
# +
# Load training and eval data from tf.keras
if dataset_name == 'mnist':
(train_images, train_labels), (test_images, test_labels) = \
tf.keras.datasets.mnist.load_data()
else:
(train_images, train_labels), (test_images, test_labels) = \
tf.keras.datasets.fashion_mnist.load_data()
train_images = train_images.reshape(-1, x_dim).astype('float32')
#train_images = train_images / 255. # Normalize the images to [0, 1]
train_images = (train_images - 127.5) / 127.5 # Normalize the images to [-1, 1]
test_images = test_images.reshape(-1, x_dim).astype('float32')
#test_images = test_images / 255. # Normalize the images to [0, 1]
test_images = (test_images - 127.5) / 127.5 # Normalize the images to [-1, 1]
# -
# ## Set up dataset with `tf.data`
#
# ### create input pipeline with `tf.data.Dataset`
# +
#tf.set_random_seed(219)
# for train
N = len(train_images)
train_dataset = tf.data.Dataset.from_tensor_slices(train_images)
train_dataset = train_dataset.shuffle(buffer_size=N)
train_dataset = train_dataset.batch(batch_size=batch_size)
print(train_dataset)
test_dataset = tf.data.Dataset.from_tensor_slices(test_images)
test_dataset = test_dataset.shuffle(buffer_size=N)
test_dataset = test_dataset.batch(batch_size=num_examples_to_generate)
print(test_dataset)
# -
# ## Create the AutoEncoder models
class AutoEncoder(tf.keras.Model):
"""Build a autoencoder model."""
def __init__(self, latent_dim=latent_dim, x_dim=x_dim):
super(AutoEncoder, self).__init__()
self.fc1 = layers.Dense(units=latent_dim, activation='relu')
self.fc2 = layers.Dense(units=x_dim, activation='tanh')
def call(self, inputs):
"""Run the model."""
fc1 = self.fc1(inputs)
fc2 = self.fc2(fc1)
return fc2
ae = AutoEncoder()
# ## Define the loss functions and the optimizer
def mse_loss(targets, predictions):
mse_loss = tf.reduce_mean(tf.reduce_sum((predictions - targets)**2, axis=1))
return mse_loss
optimizer = tf.keras.optimizers.Adam(learning_rate)
# ## Checkpoints (Object-based saving)
checkpoint_dir = train_dir
if not tf.io.gfile.exists(checkpoint_dir):
tf.io.gfile.makedirs(checkpoint_dir)
checkpoint_prefix = os.path.join(checkpoint_dir, "ckpt")
checkpoint = tf.train.Checkpoint(optimizer=optimizer, ae=ae)
# ## Training
# keeping the random image constant for generation (prediction) so
# it will be easier to see the improvement of the autoencoder.
for inputs in test_dataset.take(1):
constant_test_input = inputs
# ### Define training one step function
# Notice the use of `tf.function`
# This annotation causes the function to be "compiled".
@tf.function
def train_step(images):
with tf.GradientTape() as tape:
reconstruction = ae(images)
loss = mse_loss(images, reconstruction)
gradients = tape.gradient(loss, ae.trainable_variables)
optimizer.apply_gradients(zip(gradients, ae.trainable_variables))
return loss
# ### Train full steps
# +
print('Start Training.')
num_batches_per_epoch = int(N / batch_size)
global_step = tf.Variable(0, trainable=False)
for epoch in range(max_epochs):
for step, images in enumerate(train_dataset):
start_time = time.time()
loss = train_step(images)
global_step.assign_add(1)
if global_step.numpy() % print_steps == 0:
epochs = epoch + step / float(num_batches_per_epoch)
duration = time.time() - start_time
examples_per_sec = batch_size / float(duration)
display.clear_output(wait=True)
print("Epochs: {:.2f} global_step: {} loss: {:.3f} ({:.2f} examples/sec; {:.3f} sec/batch)".format(
epochs, global_step.numpy(), loss, examples_per_sec, duration))
for images in test_dataset.take(1):
sample_images = ae(images)
print_or_save_sample_images_two(images.numpy(),
sample_images.numpy(),
num_examples_to_generate)
if (epoch + 1) % save_images_epochs == 0:
display.clear_output(wait=True)
print("This images are saved at {} epoch".format(epoch+1))
sample_images = ae(constant_test_input)
print_or_save_sample_images_two(constant_test_input.numpy(),
sample_images.numpy(),
num_examples_to_generate,
is_save=True, epoch=epoch+1,
checkpoint_dir=checkpoint_dir)
# saving (checkpoint) the model every save_epochs
if (epoch + 1) % save_model_epochs == 0:
checkpoint.save(file_prefix=checkpoint_prefix)
print('Training Done.')
# -
# generating after the final epoch
display.clear_output(wait=True)
sample_images = ae(constant_test_input)
print_or_save_sample_images_two(constant_test_input.numpy(),
sample_images.numpy(),
num_examples_to_generate,
is_save=True, epoch=epoch+1,
checkpoint_dir=checkpoint_dir)
# ## Restore the latest checkpoint
# restoring the latest checkpoint in checkpoint_dir
checkpoint.restore(tf.train.latest_checkpoint(checkpoint_dir))
# ## Display an image using the epoch number
display_image(max_epochs, checkpoint_dir=checkpoint_dir)
# ## Generate a GIF of all the saved images.
filename = model_name + '_' + dataset_name + '.gif'
generate_gif(filename, checkpoint_dir)
display.Image(filename=filename + '.png')
| latentvariable/autoencoder.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from sklearn import datasets
import numpy as np
mat = datasets.make_spd_matrix(10)
mat
masking_array=np.random.binomial(1, .1, mat.shape).astype(bool)
mat[masking_array] = np.nan
mat[:4, :4]
from sklearn import preprocessing
impute = preprocessing.Imputer()
scaler = preprocessing.StandardScaler()
mat_imputed = impute.fit_transform(mat)
mat_imputed[:4,:4]
mat_imp_and_scaled=scaler.fit_transform(mat_imputed)
mat_imp_and_scaled[:4, :4]
mat[:4,:4]
from sklearn import pipeline
pipe=pipeline.Pipeline([('impute',impute),('scaler',scaler)])
pipe
new_mat = pipe.fit_transform(mat)
new_mat[:4, :4]
from sklearn.decomposition import FactorAnalysis
iris = datasets.load_iris()
fa = FactorAnalysis(n_components=2)
iris_two_dim=fa.fit_transform(iris.data)
from sklearn.cluster import KMeans
from scipy.spatial.distance import cdist
import numpy as np
import matplotlib.pyplot as plt
K = range(1, 10)
meandistortions = []
for k in K:
kmeans=KMeans(n_clusters=k)
kmeans.fit(iris_two_dim)
meandistortions.append(sum(np.min(
cdist(iris_two_dim,kmeans.cluster_centers_,
'euclidean'),axis=1))/iris_two_dim.shape[0])
plt.plot(K,meandistortions,'bx-')
plt.xlabel('k')
from sklearn import metrics
colors = ['b', 'g', 'r', 'c', 'm', 'y', 'k', 'b']
kmeans_model=KMeans(n_clusters=2).fit(iris_two_dim)
for i,l in enumerate(kmeans_model.labels_):
plt.plot(iris_two_dim[:,0][i],iris_two_dim[:,1][i],
color=colors[l],marker='o',ls='None')
from sklearn.datasets import load_iris
import numpy as np
iris=load_iris()
iris_data=iris.data
mask=np.random.binomial(1, .25, iris_data.shape).astype(bool)
iris_data[mask] = np.nan
iris_data[:5]
from sklearn import pipeline, preprocessing, decomposition
pca = decomposition.PCA()
imputer = preprocessing.Imputer()
pipe = pipeline.Pipeline([('imputer',imputer),('pca', pca)])
iris_data_transformed=pipe.fit_transform(iris_data)
iris_data_transformed[:5]
| Mathine_Learning/.ipynb_checkpoints/pipelines-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# #### A NLP project that estimates the sentiment scores of ABC news headlines over years using Affinn lexicon
#import libraries
library(tidyverse)
library(ggplot2)
library(reshape2)
library(quanteda) # install.packages("quanteda")
library(wordcloud2)
library(RColorBrewer)
library(tidytext)
library(grid)
#import data
raw_data <- read_csv('./abc_news_headlines.csv')
head(raw_data)
#have a look at the feature statistics
summary(raw_data)
#check if there is any missing values
colnames(raw_data)[apply(raw_data, 2, anyNA)]
# ##### As can be observed that the data contains number of count of different reaction to the headlines, this will be used to pre-determine people's emotion to the news.
# ##### 1. Positive emotion will include: likes, love, and thankful
# ##### 2. Negative emotion will inlude: sad and angry
# ##### 3. Neutral emotion will include: wow and haha
# +
#remove all the null rows
new_data <- raw_data[complete.cases(raw_data),]
#remove useless columns
new_data <- new_data[c(-12,-13)]
#add column for year and month
new_data$year_month <- substring(new_data$publish_date, 1, 6)
#add column for year
new_data$year <- substring(new_data$publish_date, 1, 4)
#add column for month
new_data$month <- substring(new_data$publish_date, 5, 6)
#define postivie emotion
new_data$positive_emotion <- new_data$likes_count + new_data$love_count + new_data$thankful_count
#define negative emotion
new_data$negative_emotion <- new_data$angry_count + new_data$sad_count
#define minimum value to be 1
new_data$negative_emotion[new_data$negative_emotion == 0] <- 1
#define nuetral emotion
new_data$neutral_emotion <- new_data$wow_count + new_data$haha_count
head(new_data)
# +
#define function to plot the barchart of number of emotion counts over year
emotion_df <- new_data %>% group_by(year) %>% summarise(
'positive emotion' = sum(positive_emotion),
'negative emotion' = sum(negative_emotion),
'neutral emotion' = sum(neutral_emotion))
emotion_df <- melt(emotion_df, id.vars = 'year')
emotion_df <- emotion_df %>% rename('reaction' = 'variable')
plot_all_emotion <- ggplot(emotion_df, aes(year, value/1e6, fill = reaction)) +
geom_bar(stat = 'identity', position = 'dodge') +
ylab('Number of Emotion Counts (Million)') +
theme_light() + theme(axis.text.x = element_text(angle = 90, vjust = 0.5, hjust=1))
positive_emotion_df <- new_data %>% group_by(year) %>% summarise(
'like' = sum(likes_count), 'love' = sum(love_count), 'thankful' = sum(thankful_count))
positive_emotion_df <- melt(positive_emotion_df, id.vars = 'year')
positive_emotion_df <- positive_emotion_df %>% rename('reaction' = 'variable')
plot_positive_emotion <- ggplot(positive_emotion_df, aes(year, value/1e6, fill =reaction)) +
geom_bar(stat = 'identity', position = 'dodge') +
ylab('Number of Emotion Counts (Million)') + theme_light() + theme(axis.text.x = element_blank())
#remove all the count columns
new_data <- new_data[c(-3:-11, -15:-17)]
# -
#combine the plots together
grid.newpage()
grid.draw(rbind(ggplotGrob(plot_all_emotion), ggplotGrob(plot_positive_emotion), size = 'first'))
# ##### The first barchart shows that the defined positive emotion outnumbers the other two emotions for all years. This can be explained by the second barchart as 'like' significantly outnumbers 'love' and 'thankful'. This may imply that when people react to news or even other post using 'like', it does not necessarily mean that they actually like the content of the news. The assumption of positivity of 'like' reaction above is wrong. Hence, the analysis below will use a more proper approach to investigate the sentiment.
#define function to generate corpus from the headline_text column
create_corpus <- function(dataframe){
#create initial corpus
corpus <- corpus(dataframe, text_field = 'headline_text') #some unused parameters: remove_twitter = TRUE, tolower = TRUE
#create cleaned corpus using quanteda
clean_corpus <- corpus %>%
tokens(remove_symbols = TRUE, remove_punct = TRUE, remove_numbers = TRUE) %>%
tokens_remove(stopwords("en")) %>%
tokens_wordstem()
return(clean_corpus)
}
#define a function to convert quanteda corpus to dataframe for word frequency
corpus_to_frame <- function(corpus, min_termfreq, min_docfreq){
#convert corpus to a sparse document-feature matrix
dfmat <- dfm(corpus)
#reduce the size of dfm
trimmed_dfm <- dfm_trim(dfmat, min_termfreq = min_termfreq, min_docfreq = min_docfreq)
#compute the total frequency of each word within the columns
frequency <- sort(colSums(as.matrix(trimmed_dfm)), decreasing=TRUE)
#construct the dataframe
df <- data.frame(word = names(frequency), freq=frequency, row.names = NULL)
return(df)
}
#define a smaller dataframe for news headlines
trimmed_data <- new_data[new_data$year>=2019,]
head(trimmed_data)
# ##### The data is trimmed to show only headlines after 2019 in order to reduce computational complexity for creating corpus
#call the function
small_corpus <- create_corpus(trimmed_data)
new_word_freq_df <- corpus_to_frame(small_corpus, 3, NULL)
head(new_word_freq_df)
#construct the word cloud
wordcloud2(data = new_word_freq_df, minRotation = 0, maxRotation = 0, ellipticity = 0.6, size = 0.5)
# ##### The result of the generated wordcloud is expected as the term 'coronavirus' dominates the headlines between 2019 and 2020. In comparison to the second term 'australia', the frequency of 'coronavirus' is 3000 higher.
#use afin setimental score for this analysis
afin_score <- as.data.frame(get_sentiments("afin"))
head(afin_score)
afin_score[afin_score$word=='coronavirus'|afin_score$word=='covid',]
# ##### However, when using Afinn lexicon to match the words, there is no result for 'coronavirus' or 'covid', meaning that the sentimental analysis for year 2019 and 2020 will be affected, which is also expected as the lexicon has not been updated yet.
#define a function to output the score for matching headline key words
get_score <- function(dataframe){
#separate the words in each row into separate row
expanded_data <- dataframe %>% separate_rows(headline_text) %>% rename(word = headline_text)
#store the score of matching words
score_df <- merge(expanded_data, afin_score, by = 'word', all.y = TRUE)
#remove missing values
score_df <- score_df[complete.cases(score_df), ]
return(score_df)
}
#count the number of headlines each month over the years
year_month_count <- new_data %>% count(year_month)
#call the function to output sentiment score
score_df <- get_score(new_data)
#estimate the total sentiment score each month over the eyars
score <- score_df %>% group_by(year_month) %>% summarise(sum = sum(score))
#compute the mean score
score$mean <- score$sum/year_month_count$n
#plot the totoal number of headlines in thousand units each month over the year
plot_all_headlines <- ggplot(year_month_count, aes(year_month, n/1000, group = 1)) + geom_line(col = 'chartreuse4', size = 1) +
theme_light() + ylab('Monthly Number of Headlines (Thousands)') +
scale_x_discrete(breaks = seq(200302, 202012, by = 100)) +
theme(axis.text.y = element_text(vjust = 0.5, hjust = 0.3),
axis.text.x = element_text(angle = 60, vjust = 0.5, hjust = 0.5),
axis.title.x = element_blank(),
legend.position = "none")
#plot the mean sentiment score each month over the year
plot_all_score <- ggplot(score, aes(year_month, mean, group = 1)) + geom_line(col = 'orange', size = 1) + theme_light() +
scale_x_discrete(breaks = seq(200302, 202012, by = 100)) + ylab('Monthly Mean Sentiment Score') +
theme(axis.text.x = element_blank(),
axis.title.x = element_blank(),
legend.position = "none")
#combine the plots together
grid.newpage()
grid.draw(rbind(ggplotGrob(plot_all_headlines), ggplotGrob(plot_all_score), size = "last"))
# ##### The changes in number of headlines experiences 3 local minima in 2003, 2006 and 2015. While there are no other evidence provided, the reason can be hardly interpreted, especially for 2003 when they whole world was experiencing SARS pandemic. Moreover, the plot shows decreasing trend after 2015 in comparison to the fluctuation in the previous years in spite of the 3 sharp declines.
# ##### It is very intersting to observe that all the mean sentiment scores negative. It does not necessarily imply that most of the news were negative but most of the key words used were negative. As people tend to pay more attention to negative things, the media uses such negativity theory to attract more readers.
| nlp_estimate_sentiment_score.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import os
#import umap
import math
import h5py
import scipy
import pickle
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from helper import *
np.set_printoptions(precision=2)
plt.rcParams['axes.labelsize'] = 25
plt.rc('xtick',labelsize=20)
plt.rc('ytick',labelsize=20)
plt.style.use('seaborn-whitegrid')
#plt.style.use('science')
# %matplotlib inline
# -
def get_distributions(arr):
if len(arr)!=2:
raise ValueError('Please enter only two arrays')
temp_arr = np.hstack((arr[0],arr[1]))
l_bound,u_bound = np.min(temp_arr),np.max(temp_arr)
bins = np.arange(math.floor(l_bound),math.ceil(u_bound));
p,_ = np.histogram(arr[0], bins = bins, density = True)
q,_ = np.histogram(arr[1], bins = bins, density = True)
return p,q
# ### Load training, test and control data
# +
met_model = 'varma_ecoli_small'
exp_id = 'fdp1'
# Create control dataset
'''
all_data_km = np.load(f'gan_input/{met_model}/{exp_id}/all_km_fdp1.npy')
control_samples = 5000
range_km = [np.min(all_data_km), np.max(all_data_km)]
control_data = np.random.uniform(range_km[0],range_km[1],(control_samples,all_data_km.shape[1]))
np.save('control_data_{self.exp_id}.npy',control_data)
'''
X_train = np.load(f'gan_input/{met_model}/{exp_id}/X_train_fdp1.npy')
X_test = np.load(f'gan_input/{met_model}/{exp_id}/X_val_fdp1.npy')
X_control = np.log(np.load(f'gan_input/{met_model}/{exp_id}/control_data_fdp1.npy'))
# -
# ### Calculate KL divergences
def calculate_distance(ref_data, path_to_folder, epochs, repeats):
X_mean = ref_data.mean(axis=0)
KL = []
for rep in range(repeats):
this_KL = []
for j in range(0,epochs,10):
this_set = np.load(f'{path_to_gan_data}repeat_{rep}/{j}_-1.npy')
gan_mean = this_set.mean(axis=0)
p,q = get_distributions([X_mean,gan_mean])
this_KL.append(KL_div(q,p))
KL.append(this_KL)
return KL
# +
epochs = 1000
repeats = 5
path_to_gan_data = 'gan_output_data/N-72000/'
KL_train = calculate_distance(X_train, path_to_gan_data, epochs, repeats)
KL_test = calculate_distance(X_test, path_to_gan_data, epochs, repeats)
KL_control = calculate_distance(X_control, path_to_gan_data, epochs, repeats)
# +
x_plot = np.arange(0,1000,10)
plt.figure(figsize=(20,10))
plt.plot(x_plot, np.array(KL_test).mean(axis = 0), color = '#477998', linewidth = 3, label = 'Test Data')
plt.plot(x_plot, np.array(KL_test).max(axis = 0), '--',color = '#477998', linewidth = 3, alpha = 1)
plt.plot(x_plot, np.array(KL_test).min(axis = 0), '--',color = '#477998' , linewidth = 3, alpha = 1)
plt.plot(x_plot, np.array(KL_train).mean(axis = 0), color = '#C4D6B0', linewidth = 3, label = 'Training Data')
plt.plot(x_plot, np.array(KL_train).max(axis = 0), '--',color = '#C4D6B0', linewidth = 3, alpha = 0.8)
plt.plot(x_plot, np.array(KL_train).min(axis = 0), '--',color = '#C4D6B0' , linewidth = 3, alpha = 0.8)
plt.legend(fontsize = 20)
plt.xlabel('Epochs')
plt.ylabel('KL divergence')
plt.savefig(f'figure_output/test_training.svg', dpi=300,
transparent=False, bbox_inches='tight')
# +
x_plot = np.arange(0,1000,10)
plt.figure(figsize=(20,10))
plt.plot(x_plot, np.array(KL_test).mean(axis = 0), color = '#477998', linewidth = 3, label = 'Test Data')
plt.plot(x_plot, np.array(KL_test).max(axis = 0), '--',color = '#477998', linewidth = 3, alpha = 1)
plt.plot(x_plot, np.array(KL_test).min(axis = 0), '--',color = '#477998' , linewidth = 3, alpha = 1)
plt.plot(x_plot, np.array(KL_train).mean(axis = 0), color = '#C4D6B0', linewidth = 3, label = 'Training Data')
plt.plot(x_plot, np.array(KL_train).max(axis = 0), '--',color = '#C4D6B0', linewidth = 3, alpha = 0.8)
plt.plot(x_plot, np.array(KL_train).min(axis = 0), '--',color = '#C4D6B0' , linewidth = 3, alpha = 0.8)
plt.plot(x_plot, np.array(KL_control).mean(axis = 0), color = '#291F1E', linewidth = 3, label = 'Control Data')
plt.plot(x_plot, np.array(KL_control).max(axis = 0), '--',color = '#291F1E', linewidth = 3, alpha = 0.8)
plt.plot(x_plot, np.array(KL_control).min(axis = 0), '--',color = '#291F1E' , linewidth = 3, alpha = 0.8)
plt.legend(fontsize = 20)
plt.xlabel('Epochs')
plt.ylabel('KL divergence')
plt.savefig(f'figure_output/test_training_control.svg', dpi=300,
transparent=False, bbox_inches='tight')
# -
| venv/distance_check.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/JRPateno/OOP-58002/blob/main/OOP_2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="O4z0h9o70CUy"
#
# + colab={"base_uri": "https://localhost:8080/"} id="iyuo0KD_0CoQ" outputId="00732008-8c6a-4370-c435-9be4e6b84589"
class Birds:
def __init__(self,bird_name):
self.bird_name = bird_name
def flying_birds(self):
print(f"{self.bird_name} flies above clouds")
def non_flying_birds(self):
print(f"{self.bird_name} is the national bird of Philippines")
vulture = Birds("Griffon Vulture")
crane = Birds("Common Crane")
emu = Birds("Emu")
vulture.flying_birds()
crane.flying_birds()
emu.non_flying_birds()
# + [markdown] id="8f35FvS40Ds5"
# Encapsulation (mangling w/ double underscore)
# + colab={"base_uri": "https://localhost:8080/"} id="3tR0ztJm0FaA" outputId="c8914914-c376-459e-8236-a8b311147bb7"
class foo:
def __init__(self,a,b):
self.a=a
self.b=b
def add(self):
return self.a + self.b
def sub(self):
return self.a - self.b
foo_object=foo(3,4)
foo_object.add()
foo_object.sub()
#Or
#print(foo_object.add())
#print(foo_object.sub())
foo_object.b = 5
print(foo_object.b)
print(foo_object.add())
print(foo_object.sub())
# + colab={"base_uri": "https://localhost:8080/"} id="odUlYPR72iAf" outputId="283b2103-4c78-4010-f1b1-e84446a74f8c"
class foo:
def __init__(self,a,b):
self.__a=a
self.__b=b
def add(self):
return self.__a + self.__b
def sub(self):
return self.__a - self.__b
foo_object=foo(3,4)
print(foo_object.add())
print(foo_object.sub())
foo_object.__b = 5
foo_object.__a = 7
print(foo_object.add())
print(foo_object.sub())
# + [markdown] id="uHIWuvrX6GIP"
# Inheritance
# + colab={"base_uri": "https://localhost:8080/"} id="AXCynSi76H3H" outputId="ddc60443-1f76-4856-f687-3e0d00238467"
class Person:
def __init__(self,firstname,surname):
self.firstname = firstname
self.surname = surname
def printname(self):
print(self.firstname,self.surname)
person = Person("<NAME>","Pateno")
person.printname()
class Student(Person):
pass
person = Student("Ash","Pikachu")
person.printname()
# + [markdown] id="7x29RHxE8sHG"
# Polymorphism
# + colab={"base_uri": "https://localhost:8080/"} id="xgjjooOQ8svR" outputId="84798fe3-39ad-4862-c9ee-3e40c6b26bdb"
class RegularPolygon:
def __init__(self,side):
self.side = side
class Square(RegularPolygon):
def area(self):
return self.side * self.side
class EquilateralTriangle(RegularPolygon):
def area(self):
return self.side * self.side * 0.433
x = Square(4)
y = EquilateralTriangle(3)
print(x.area())
print(y.area())
# + [markdown] id="GW2VQS3mAGbd"
# Application 1
# + [markdown] id="wJzEWlWmANOV"
# 1. create a python program that displays the name of three students(Student 1,Student 2,and Student 3) and their term grades
# 2. create a class name Person and attributes - std1,std2,std3,pre,mid,fin
# 3. compute the average of each term grade using Grade() method
# 4. information about student's grades must be hidden from others
# + colab={"base_uri": "https://localhost:8080/"} id="kSq95klXAqHN" outputId="c2660ab8-80a9-44ac-9edc-ce11e0f320dd"
class Student:
def __init__(self,name,pre,mid,fin):
self.name = name
self.pre = pre
self.mid = mid
self.fin = fin
def display(self):
print(f"Name:{self.name}")
print(f"Prelim Grade: {self.pre}")
print(f"Midterm Grade: {self.mid}")
print(f"Final Grade: {self.fin}")
return"\t"
Student1 = Student("<NAME>",84,85,86)
Student2 = Student("<NAME>",82,84,85)
Student3 = Student("<NAME>",86,86,89)
print(Student1.display())
print(Student2.display())
print(Student3.display())
# + id="9WSIiu8iG19c"
class Person:
def __init__(self,std1,std2,std3,pre,mid,fin):
self.std1 = std1
self.std2 = std2
self.std3 = std3
self.pre = pre
self.mid = mid
self.fin = fin
# + id="g1IDzFzbEqK9" colab={"base_uri": "https://localhost:8080/"} outputId="cdd60740-af8d-41a8-e180-d4ec3fdbd9c9"
class Compute:
def __init__(self,class_standing,exam):
self.class_standing = class_standing
self.exam = exam
def Grade(self):
return (self.class_standing * 0.60) + (self.exam * 0.40)
pregrade = Compute(85,75)
print(pregrade.Grade())
midgrade = Compute(80,70)
print(midgrade.Grade())
fingrade = Compute(83,74)
print(fingrade.Grade())
| OOP_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Dirichlet process mixtures for density estimation
# Author: [<NAME>](https://github.com/AustinRochford/)
#
# ## Dirichlet processes
#
# The [Dirichlet process](https://en.wikipedia.org/wiki/Dirichlet_process) is a flexible probability distribution over the space of distributions. Most generally, a probability distribution, $P$, on a set $\Omega$ is a [measure](https://en.wikipedia.org/wiki/Measure_(mathematics%29) that assigns measure one to the entire space ($P(\Omega) = 1$). A Dirichlet process $P \sim \textrm{DP}(\alpha, P_0)$ is a measure that has the property that, for every finite [disjoint](https://en.wikipedia.org/wiki/Disjoint_sets) partition $S_1, \ldots, S_n$ of $\Omega$,
#
# $$(P(S_1), \ldots, P(S_n)) \sim \textrm{Dir}(\alpha P_0(S_1), \ldots, \alpha P_0(S_n)).$$
#
# Here $P_0$ is the base probability measure on the space $\Omega$. The precision parameter $\alpha > 0$ controls how close samples from the Dirichlet process are to the base measure, $P_0$. As $\alpha \to \infty$, samples from the Dirichlet process approach the base measure $P_0$.
#
# Dirichlet processes have several properties that make them quite suitable to [MCMC](https://en.wikipedia.org/wiki/Markov_chain_Monte_Carlo) simulation.
#
# 1. The posterior given [i.i.d.](https://en.wikipedia.org/wiki/Independent_and_identically_distributed_random_variables) observations $\omega_1, \ldots, \omega_n$ from a Dirichlet process $P \sim \textrm{DP}(\alpha, P_0)$ is also a Dirichlet process with
#
# $$P\ |\ \omega_1, \ldots, \omega_n \sim \textrm{DP}\left(\alpha + n, \frac{\alpha}{\alpha + n} P_0 + \frac{1}{\alpha + n} \sum_{i = 1}^n \delta_{\omega_i}\right),$$
#
# where $\delta$ is the [Dirac delta measure](https://en.wikipedia.org/wiki/Dirac_delta_function)
#
# $$\begin{align*}
# \delta_{\omega}(S)
# & = \begin{cases}
# 1 & \textrm{if } \omega \in S \\
# 0 & \textrm{if } \omega \not \in S
# \end{cases}
# \end{align*}.$$
#
# 2. The posterior predictive distribution of a new observation is a compromise between the base measure and the observations,
#
# $$\omega\ |\ \omega_1, \ldots, \omega_n \sim \frac{\alpha}{\alpha + n} P_0 + \frac{1}{\alpha + n} \sum_{i = 1}^n \delta_{\omega_i}.$$
#
# We see that the prior precision $\alpha$ can naturally be interpreted as a prior sample size. The form of this posterior predictive distribution also lends itself to Gibbs sampling.
#
# 2. Samples, $P \sim \textrm{DP}(\alpha, P_0)$, from a Dirichlet process are discrete with probability one. That is, there are elements $\omega_1, \omega_2, \ldots$ in $\Omega$ and weights $w_1, w_2, \ldots$ with $\sum_{i = 1}^{\infty} w_i = 1$ such that
#
# $$P = \sum_{i = 1}^\infty w_i \delta_{\omega_i}.$$
#
# 3. The [stick-breaking process](https://en.wikipedia.org/wiki/Dirichlet_process#The_stick-breaking_process) gives an explicit construction of the weights $w_i$ and samples $\omega_i$ above that is straightforward to sample from. If $\beta_1, \beta_2, \ldots \sim \textrm{Beta}(1, \alpha)$, then $w_i = \beta_i \prod_{j = 1}^{n - 1} (1 - \beta_j)$. The relationship between this representation and stick breaking may be illustrated as follows:
# 1. Start with a stick of length one.
# 2. Break the stick into two portions, the first of proportion $w_1 = \beta_1$ and the second of proportion $1 - w_1$.
# 3. Further break the second portion into two portions, the first of proportion $\beta_2$ and the second of proportion $1 - \beta_2$. The length of the first portion of this stick is $\beta_2 (1 - \beta_1)$; the length of the second portion is $(1 - \beta_1) (1 - \beta_2)$.
# 4. Continue breaking the second portion from the previous break in this manner forever. If $\omega_1, \omega_2, \ldots \sim P_0$, then
#
# $$P = \sum_{i = 1}^\infty w_i \delta_{\omega_i} \sim \textrm{DP}(\alpha, P_0).$$
#
# [Suggested Further Reading]: (http://mlg.eng.cam.ac.uk/tutorials/07/ywt.pdf) and (https://www.stats.ox.ac.uk/~teh/research/npbayes/Teh2010a.pdf) for a brief introduction to other flavours of Dirichlet Processes, and their applications.
#
# We can use the stick-breaking process above to easily sample from a Dirichlet process in Python. For this example, $\alpha = 2$ and the base distribution is $N(0, 1)$.
# %matplotlib inline
from __future__ import division
from matplotlib import pyplot as plt
import numpy as np
import pymc3 as pm
import scipy as sp
import seaborn as sns
from theano import tensor as tt
import pandas as pd
blue, *_ = sns.color_palette()
# +
SEED = 5132290 # from random.org
np.random.seed(SEED)
# +
N = 20
K = 30
alpha = 2.
P0 = sp.stats.norm
# -
# We draw and plot samples from the stick-breaking process.
# +
beta = sp.stats.beta.rvs(1, alpha, size=(N, K))
w = np.empty_like(beta)
w[:, 0] = beta[:, 0]
w[:, 1:] = beta[:, 1:] * (1 - beta[:, :-1]).cumprod(axis=1)
omega = P0.rvs(size=(N, K))
x_plot = np.linspace(-3, 3, 200)
sample_cdfs = (w[..., np.newaxis] * np.less.outer(omega, x_plot)).sum(axis=1)
# +
fig, ax = plt.subplots(figsize=(8, 6))
ax.plot(x_plot, sample_cdfs[0], c='gray', alpha=0.75,
label='DP sample CDFs');
ax.plot(x_plot, sample_cdfs[1:].T, c='gray', alpha=0.75);
ax.plot(x_plot, P0.cdf(x_plot), c='k', label='Base CDF');
ax.set_title(r'$\alpha = {}$'.format(alpha));
ax.legend(loc=2);
# -
# As stated above, as $\alpha \to \infty$, samples from the Dirichlet process converge to the base distribution.
# +
fig, (l_ax, r_ax) = plt.subplots(ncols=2, sharex=True, sharey=True, figsize=(16, 6))
K = 50
alpha = 10.
beta = sp.stats.beta.rvs(1, alpha, size=(N, K))
w = np.empty_like(beta)
w[:, 0] = beta[:, 0]
w[:, 1:] = beta[:, 1:] * (1 - beta[:, :-1]).cumprod(axis=1)
omega = P0.rvs(size=(N, K))
sample_cdfs = (w[..., np.newaxis] * np.less.outer(omega, x_plot)).sum(axis=1)
l_ax.plot(x_plot, sample_cdfs[0], c='gray', alpha=0.75,
label='DP sample CDFs');
l_ax.plot(x_plot, sample_cdfs[1:].T, c='gray', alpha=0.75);
l_ax.plot(x_plot, P0.cdf(x_plot), c='k', label='Base CDF');
l_ax.set_title(r'$\alpha = {}$'.format(alpha));
l_ax.legend(loc=2);
K = 200
alpha = 50.
beta = sp.stats.beta.rvs(1, alpha, size=(N, K))
w = np.empty_like(beta)
w[:, 0] = beta[:, 0]
w[:, 1:] = beta[:, 1:] * (1 - beta[:, :-1]).cumprod(axis=1)
omega = P0.rvs(size=(N, K))
sample_cdfs = (w[..., np.newaxis] * np.less.outer(omega, x_plot)).sum(axis=1)
r_ax.plot(x_plot, sample_cdfs[0], c='gray', alpha=0.75,
label='DP sample CDFs');
r_ax.plot(x_plot, sample_cdfs[1:].T, c='gray', alpha=0.75);
r_ax.plot(x_plot, P0.cdf(x_plot), c='k', label='Base CDF');
r_ax.set_title(r'$\alpha = {}$'.format(alpha));
r_ax.legend(loc=2);
# -
# ## Dirichlet process mixtures
#
# For the task of density estimation, the (almost sure) discreteness of samples from the Dirichlet process is a significant drawback. This problem can be solved with another level of indirection by using Dirichlet process mixtures for density estimation. A Dirichlet process mixture uses component densities from a parametric family $\mathcal{F} = \{f_{\theta}\ |\ \theta \in \Theta\}$ and represents the mixture weights as a Dirichlet process. If $P_0$ is a probability measure on the parameter space $\Theta$, a Dirichlet process mixture is the hierarchical model
#
# $$
# \begin{align*}
# x_i\ |\ \theta_i
# & \sim f_{\theta_i} \\
# \theta_1, \ldots, \theta_n
# & \sim P \\
# P
# & \sim \textrm{DP}(\alpha, P_0).
# \end{align*}
# $$
#
# To illustrate this model, we simulate draws from a Dirichlet process mixture with $\alpha = 2$, $\theta \sim N(0, 1)$, $x\ |\ \theta \sim N(\theta, (0.3)^2)$.
# +
N = 5
K = 30
alpha = 2
P0 = sp.stats.norm
f = lambda x, theta: sp.stats.norm.pdf(x, theta, 0.3)
# +
beta = sp.stats.beta.rvs(1, alpha, size=(N, K))
w = np.empty_like(beta)
w[:, 0] = beta[:, 0]
w[:, 1:] = beta[:, 1:] * (1 - beta[:, :-1]).cumprod(axis=1)
theta = P0.rvs(size=(N, K))
dpm_pdf_components = f(x_plot[np.newaxis, np.newaxis, :], theta[..., np.newaxis])
dpm_pdfs = (w[..., np.newaxis] * dpm_pdf_components).sum(axis=1)
# +
fig, ax = plt.subplots(figsize=(8, 6))
ax.plot(x_plot, dpm_pdfs.T, c='gray');
ax.set_yticklabels([]);
# -
# We now focus on a single mixture and decompose it into its individual (weighted) mixture components.
# +
fig, ax = plt.subplots(figsize=(8, 6))
ix = 1
ax.plot(x_plot, dpm_pdfs[ix], c='k', label='Density');
ax.plot(x_plot, (w[..., np.newaxis] * dpm_pdf_components)[ix, 0],
'--', c='k', label='Mixture components (weighted)');
ax.plot(x_plot, (w[..., np.newaxis] * dpm_pdf_components)[ix].T,
'--', c='k');
ax.set_yticklabels([]);
ax.legend(loc=1);
# -
# Sampling from these stochastic processes is fun, but these ideas become truly useful when we fit them to data. The discreteness of samples and the stick-breaking representation of the Dirichlet process lend themselves nicely to Markov chain Monte Carlo simulation of posterior distributions. We will perform this sampling using `PyMC3`.
#
# Our first example uses a Dirichlet process mixture to estimate the density of waiting times between eruptions of the [Old Faithful](https://en.wikipedia.org/wiki/Old_Faithful) geyser in [Yellowstone National Park](https://en.wikipedia.org/wiki/Yellowstone_National_Park).
old_faithful_df = pd.read_csv(pm.get_data('old_faithful.csv'))
# For convenience in specifying the prior, we standardize the waiting time between eruptions.
old_faithful_df['std_waiting'] = (old_faithful_df.waiting - old_faithful_df.waiting.mean()) / old_faithful_df.waiting.std()
old_faithful_df.head()
# +
fig, ax = plt.subplots(figsize=(8, 6))
n_bins = 20
ax.hist(old_faithful_df.std_waiting, bins=n_bins, color=blue, lw=0, alpha=0.5);
ax.set_xlabel('Standardized waiting time between eruptions');
ax.set_ylabel('Number of eruptions');
# -
# Observant readers will have noted that we have not been continuing the stick-breaking process indefinitely as indicated by its definition, but rather have been truncating this process after a finite number of breaks. Obviously, when computing with Dirichlet processes, it is necessary to only store a finite number of its point masses and weights in memory. This restriction is not terribly onerous, since with a finite number of observations, it seems quite likely that the number of mixture components that contribute non-neglible mass to the mixture will grow slower than the number of samples. This intuition can be formalized to show that the (expected) number of components that contribute non-negligible mass to the mixture approaches $\alpha \log N$, where $N$ is the sample size.
#
# There are various clever [Gibbs sampling](https://en.wikipedia.org/wiki/Gibbs_sampling) techniques for Dirichlet processes that allow the number of components stored to grow as needed. [Stochastic memoization](http://danroy.org/papers/RoyManGooTen-ICMLNPB-2008.pdf) is another powerful technique for simulating Dirichlet processes while only storing finitely many components in memory. In this introductory example, we take the much less sophistocated approach of simply truncating the Dirichlet process components that are stored after a fixed number, $K$, of components. [Ohlssen, et al.](http://fisher.osu.edu/~schroeder.9/AMIS900/Ohlssen2006.pdf) provide justification for truncation, showing that $K > 5 \alpha + 2$ is most likely sufficient to capture almost all of the mixture weight ($\sum_{i = 1}^{K} w_i > 0.99$). In practice, we can verify the suitability of our truncated approximation to the Dirichlet process by checking the number of components that contribute non-negligible mass to the mixture. If, in our simulations, all components contribute non-negligible mass to the mixture, we have truncated the Dirichlet process too early.
#
# Our (truncated) Dirichlet process mixture model for the standardized waiting times is
#
# $$
# \begin{align*}
# \alpha
# & \sim \textrm{Gamma}(1, 1) \\
# \beta_1, \ldots, \beta_K
# & \sim \textrm{Beta}(1, \alpha) \\
# w_i
# & = \beta_i \prod_{j = i - 1}^i (1 - \beta_j) \\
# \\
# \lambda_1, \ldots, \lambda_K
# & \sim U(0, 5) \\
# \tau_1, \ldots, \tau_K
# & \sim \textrm{Gamma}(1, 1) \\
# \mu_i\ |\ \lambda_i, \tau_i
# & \sim N\left(0, (\lambda_i \tau_i)^{-1}\right) \\
# \\
# x\ |\ w_i, \lambda_i, \tau_i, \mu_i
# & \sim \sum_{i = 1}^K w_i\ N(\mu_i, (\lambda_i \tau_i)^{-1})
# \end{align*}
# $$
#
# Note that instead of fixing a value of $\alpha$, as in our previous simulations, we specify a prior on $\alpha$, so that we may learn its posterior distribution from the observations.
#
# We now construct this model using `pymc3`.
# +
N = old_faithful_df.shape[0]
K = 30
# -
def stick_breaking(beta):
portion_remaining = tt.concatenate([[1], tt.extra_ops.cumprod(1 - beta)[:-1]])
return beta * portion_remaining
with pm.Model() as model:
alpha = pm.Gamma('alpha', 1., 1.)
beta = pm.Beta('beta', 1., alpha, shape=K)
w = pm.Deterministic('w', stick_breaking(beta))
tau = pm.Gamma('tau', 1., 1., shape=K)
lambda_ = pm.Uniform('lambda', 0, 5, shape=K)
mu = pm.Normal('mu', 0, tau=lambda_ * tau, shape=K)
obs = pm.NormalMixture('obs', w, mu, tau=lambda_ * tau,
observed=old_faithful_df.std_waiting.values)
# We sample from the model 1,000 times using NUTS initialized with ADVI.
with model:
trace = pm.sample(1000, random_seed=SEED)
# The posterior distribution of $\alpha$ is highly concentrated between 0.25 and 1.
pm.traceplot(trace, varnames=['alpha']);
# To verify that truncation is not biasing our results, we plot the posterior expected mixture weight of each component.
# +
fig, ax = plt.subplots(figsize=(8, 6))
plot_w = np.arange(K) + 1
ax.bar(plot_w - 0.5, trace['w'].mean(axis=0), width=1., lw=0);
ax.set_xlim(0.5, K);
ax.set_xlabel('Component');
ax.set_ylabel('Posterior expected mixture weight');
# -
# We see that only three mixture components have appreciable posterior expected weights, so we conclude that truncating the Dirichlet process to forty components has not appreciably affected our estimates.
#
# We now compute and plot our posterior density estimate.
# +
post_pdf_contribs = sp.stats.norm.pdf(np.atleast_3d(x_plot),
trace['mu'][:, np.newaxis, :],
1. / np.sqrt(trace['lambda'] * trace['tau'])[:, np.newaxis, :])
post_pdfs = (trace['w'][:, np.newaxis, :] * post_pdf_contribs).sum(axis=-1)
post_pdf_low, post_pdf_high = np.percentile(post_pdfs, [2.5, 97.5], axis=0)
# +
fig, ax = plt.subplots(figsize=(8, 6))
n_bins = 20
ax.hist(old_faithful_df.std_waiting.values, bins=n_bins, normed=True,
color=blue, lw=0, alpha=0.5);
ax.fill_between(x_plot, post_pdf_low, post_pdf_high,
color='gray', alpha=0.45);
ax.plot(x_plot, post_pdfs[0],
c='gray', label='Posterior sample densities');
ax.plot(x_plot, post_pdfs[::100].T, c='gray');
ax.plot(x_plot, post_pdfs.mean(axis=0),
c='k', label='Posterior expected density');
ax.set_xlabel('Standardized waiting time between eruptions');
ax.set_yticklabels([]);
ax.set_ylabel('Density');
ax.legend(loc=2);
# -
# As above, we can decompose this density estimate into its (weighted) mixture components.
# +
fig, ax = plt.subplots(figsize=(8, 6))
n_bins = 20
ax.hist(old_faithful_df.std_waiting.values, bins=n_bins, normed=True,
color=blue, lw=0, alpha=0.5);
ax.plot(x_plot, post_pdfs.mean(axis=0),
c='k', label='Posterior expected density');
ax.plot(x_plot, (trace['w'][:, np.newaxis, :] * post_pdf_contribs).mean(axis=0)[:, 0],
'--', c='k', label='Posterior expected mixture\ncomponents\n(weighted)');
ax.plot(x_plot, (trace['w'][:, np.newaxis, :] * post_pdf_contribs).mean(axis=0),
'--', c='k');
ax.set_xlabel('Standardized waiting time between eruptions');
ax.set_yticklabels([]);
ax.set_ylabel('Density');
ax.legend(loc=2);
# -
# The Dirichlet process mixture model is incredibly flexible in terms of the family of parametric component distributions $\{f_{\theta}\ |\ f_{\theta} \in \Theta\}$. We illustrate this flexibility below by using Poisson component distributions to estimate the density of sunspots per year. This dataset can be downloaded from http://www.sidc.be/silso/datafiles. Source: WDC-SILSO, Royal Observatory of Belgium, Brussels.
sunspot_df = pd.read_csv(pm.get_data('sunspot.csv'), sep=';', names=['time', 'sunspot.year'], usecols=[0, 1])
sunspot_df.head()
# For this example, the model is
#
# $$
# \begin{align*}
# \alpha
# & \sim \textrm{Gamma}(1, 1) \\
# \beta_1, \ldots, \beta_K
# & \sim \textrm{Beta}(1, \alpha) \\
# w_i
# & = \beta_i \prod_{j = i - 1}^i (1 - \beta_j) \\
# \\
# \lambda_i, \ldots, \lambda_K
# & \sim U(0, 300)
# \\
# x\ |\ w_i, \lambda_i
# & \sim \sum_{i = 1}^K w_i\ \textrm{Poisson}(\lambda_i).
# \end{align*}
# $$
K = 50
N = sunspot_df.shape[0]
with pm.Model() as model:
alpha = pm.Gamma('alpha', 1., 1.)
beta = pm.Beta('beta', 1, alpha, shape=K)
w = pm.Deterministic('w', stick_breaking(beta))
mu = pm.Uniform('mu', 0., 300., shape=K)
obs = pm.Mixture('obs', w, pm.Poisson.dist(mu), observed=sunspot_df['sunspot.year'])
with model:
step = pm.Metropolis()
trace = pm.sample(1000, step=step, random_seed=SEED)
# For the sunspot model, the posterior distribution of $\alpha$ is concentrated between 0.6 and 1.2, indicating that we should expect more components to contribute non-negligible amounts to the mixture than for the Old Faithful waiting time model.
pm.traceplot(trace, varnames=['alpha']);
# Indeed, we see that between ten and fifteen mixture components have appreciable posterior expected weight.
# +
fig, ax = plt.subplots(figsize=(8, 6))
plot_w = np.arange(K) + 1
ax.bar(plot_w - 0.5, trace['w'].mean(axis=0), width=1., lw=0);
ax.set_xlim(0.5, K);
ax.set_xlabel('Component');
ax.set_ylabel('Posterior expected mixture weight');
# -
# We now calculate and plot the fitted density estimate.
x_plot = np.arange(250)
# +
post_pmf_contribs = sp.stats.poisson.pmf(np.atleast_3d(x_plot),
trace['mu'][:, np.newaxis, :])
post_pmfs = (trace['w'][:, np.newaxis, :] * post_pmf_contribs).sum(axis=-1)
post_pmf_low, post_pmf_high = np.percentile(post_pmfs, [2.5, 97.5], axis=0)
# +
fig, ax = plt.subplots(figsize=(8, 6))
ax.hist(sunspot_df['sunspot.year'].values, bins=40, normed=True, lw=0, alpha=0.75);
ax.fill_between(x_plot, post_pmf_low, post_pmf_high,
color='gray', alpha=0.45)
ax.plot(x_plot, post_pmfs[0],
c='gray', label='Posterior sample densities');
ax.plot(x_plot, post_pmfs[::200].T, c='gray');
ax.plot(x_plot, post_pmfs.mean(axis=0),
c='k', label='Posterior expected density');
ax.set_xlabel('Yearly sunspot count');
ax.set_yticklabels([]);
ax.legend(loc=1);
# -
# Again, we can decompose the posterior expected density into weighted mixture densities.
# +
fig, ax = plt.subplots(figsize=(8, 6))
ax.hist(sunspot_df['sunspot.year'].values, bins=40, normed=True, lw=0, alpha=0.75);
ax.plot(x_plot, post_pmfs.mean(axis=0),
c='k', label='Posterior expected density');
ax.plot(x_plot, (trace['w'][:, np.newaxis, :] * post_pmf_contribs).mean(axis=0)[:, 0],
'--', c='k', label='Posterior expected\nmixture components\n(weighted)');
ax.plot(x_plot, (trace['w'][:, np.newaxis, :] * post_pmf_contribs).mean(axis=0),
'--', c='k');
ax.set_xlabel('Yearly sunspot count');
ax.set_yticklabels([]);
ax.legend(loc=1);
# -
# An earlier version of this example first appeared [here](http://austinrochford.com/posts/2016-02-25-density-estimation-dpm.html).
| docs/source/notebooks/dp_mix.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + active=""
# 说明:
# 实现一个 MapSum 类,支持两个方法,insert 和 sum:
# 1、MapSum() 初始化 MapSum 对象
# 2、void insert(String key, int val) 插入 key-val 键值对,字符串表示键 key ,整数表示值 val 。
# 3、如果键 key 已经存在,那么原来的键值对将被替代成新的键值对。
#
# 示例:
# 输入:
# ["MapSum", "insert", "sum", "insert", "sum"]
# [[], ["apple", 3], ["ap"], ["app", 2], ["ap"]]
# 输出:
# [null, null, 3, null, 5]
#
# 解释:
# MapSum mapSum = new MapSum();
# mapSum.insert("apple", 3);
# mapSum.sum("ap"); // return 3 (apple = 3)
# mapSum.insert("app", 2);
# mapSum.sum("ap"); // return 5 (apple + app = 3 + 2 = 5)
#
# 提示:
# 1 <= key.length, prefix.length <= 50
# key 和 prefix 仅由小写英文字母组成
# 1 <= val <= 1000
# 最多调用 50 次 insert 和 sum
# +
class Node:
def __init__(self):
self.children = {}
self.isWord = False
class Trie:
def __init__(self):
self.root = Node()
def insert(self, word, val):
node = self.root
for char in word:
if char not in node.children:
node.children[char] = [val, Node()]
else:
node.children[char][0] += val
node = node.children[char][1]
node.isWord = True
class MapSum:
def __init__(self):
self.trie = Trie()
self.words = {}
def insert(self, key: str, val: int) -> None:
self.words[key] = val
if key not in self.words:
self.trie.insert(key, val)
else:
self.trie = Trie()
for key, val in self.words.items():
self.trie.insert(key, val)
def sum(self, prefix: str) -> int:
node = self.trie.root
res = float('inf')
for w in prefix:
if w in node.children:
res = min(res, node.children[w][0])
node = node.children[w][1]
else:
return 0
return res
# +
class TrieNode:
def __init__(self):
self.children = collections.defaultdict(TrieNode)
self.end_word = None
self.val = 0
class MapSum:
def __init__(self):
"""
Initialize your data structure here.
"""
self.root = TrieNode()
def insert(self, key: str, val: int) -> None:
# Insert into trie, increment by val at each node
node = self.root
for char in key:
node = node.children[char]
node.val += val
# For existing words, replace val by walking the trie
# again and decrementing each node by the old val
if node.end_word is not None:
old_val = node.end_word
node = self.root
for char in key:
node = node.children[char]
node.val -= old_val
# Mark end of word with the inserted value
node.end_word = val
def sum(self, prefix: str) -> int:
# Walk trie and return val at final node of prefix
node = self.root
for char in prefix:
node = node.children[char]
return node.val
# -
obj = MapSum()
obj.insert('apple', 3)
obj.insert('app', 2)
obj.insert('apple', 2)
param_2 = obj.sum('app')
print(param_2)
["MapSum", "insert", "sum", "insert", "insert", "sum"]
[[], ["apple",3], ["ap"], ["app",2], ["apple", 2], ["ap"]]
class MapSum:
def __init__(self):
"""
Initialize your data structure here.
"""
MapSum.lst = {}
def insert(self, key: str, val: int) -> None:
MapSum.lst.update({key: val})
def sum(self, prefix: str) -> int:
ans = 0
for i in MapSum.lst.keys():
if i.startswith(prefix):
ans+=MapSum.lst[i]
return ans
136 + 32 + 584 + 32 + 136 + 584 + 32
72 + 32 +584 + 288+2336 +128+128+32
| Trie/1110/677. Map Sum Pairs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="ZdW7nQ-cbIqo"
# ## Important links
#
# * https://nasa-impact.github.io/etci2021/
# * https://competitions.codalab.org/competitions/30440
# + [markdown] id="nZo0QYmDnyb7"
# ## Data Collection
# + colab={"base_uri": "https://localhost:8080/"} id="of12DJY_a2jj" outputId="bcad2c7c-9816-473f-ca7b-559790bc7d98"
# !gdown --id 14HqNW5uWLS92n7KrxKgDwUTsSEST6LCr
# + id="uXhuqjLrbD-H"
# !unzip -qq train.zip
# !rm -rf train.zip
# + colab={"base_uri": "https://localhost:8080/"} id="WGjp5GS9bzNn" outputId="f1dbe379-9381-4095-931d-54882369bd6f"
# !ls -lh train | head -10
# + colab={"base_uri": "https://localhost:8080/"} id="PX00Kq1YcLQh" outputId="0e0ded97-91fd-47bf-9711-43a452087aed"
# !ls -lh train/bangladesh_20170314t115609/tiles | head -10
# + colab={"base_uri": "https://localhost:8080/"} id="gCYM6RQ4cXlm" outputId="7cd939de-1bf0-4777-9c43-b8a42e261ab8"
# !ls -lh train/bangladesh_20170314t115609/tiles/flood_label | head -10
# + colab={"base_uri": "https://localhost:8080/"} id="bZva1Js7qfyO" outputId="b990c269-4eca-4d64-ff46-0512cbe6dd2b"
# !ls -lh train/bangladesh_20170314t115609/tiles/vh | head -10
# + [markdown] id="MpN2AOPKhKzn"
# From [here](https://nasa-impact.github.io/etci2021/#semantic-labels):
#
# > The provided training data is split across 29 root folders named \<region>\_\<datetime>*, region being the region and datetime being the date and time of the flood event. Each root folder includes 4 sub-folders: vv, vh, water_body_label and flood_label with 2,068 files each. vv and vh correspond to the satellite images listed earlier and images in the flood_label and water_body_label folder provide reference ground truth.
# + [markdown] id="ICOG2m4yn1gY"
# ## Imports
# + id="k9SNgHAnhilI"
from imutils import paths
from tqdm import tqdm
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import tensorflow as tf
import pandas as pd
import numpy as np
import cv2
import re
import os
# + [markdown] id="H5yXXB0mn2wI"
# ## Investigation
# + colab={"base_uri": "https://localhost:8080/"} id="V-saXYh5hmMr" outputId="2898b3c0-f947-4c4d-8fbf-0aec07143e2c"
all_image_paths = list(paths.list_images("train"))
print(f"Total images: {int(len(all_image_paths)/2)}")
# + [markdown] id="qo9sYuf7iDb0"
# So, we have 33,406 satellite images and the rest are binary segmentation maps.
#
# For a given image id (e.g. `nebraska_20170309t002110`), its correspnding ground-truths i.e. the segmentation maps are present in either of these two folders: `water_body_label` and `flood_label`. Let's write a few utility functions for knowing the dataset in a better way.
# + [markdown] id="oti_hOWBmI26"
# **How many unique image IDs are there?**
# + colab={"base_uri": "https://localhost:8080/"} id="TO4sCY85mMwV" outputId="3ca32213-f5e4-4da6-e2b7-22fa22b3ef94"
image_ids = {path.split("/")[1] for path in all_image_paths}
print(len(image_ids))
# + [markdown] id="gXah9TzHnILz"
# Now, let's investigate how are these IDs distributed? **Do all the IDs have the same amount of images present?**
# + id="wVGpVal5j2J0"
def get_image_paths(image_id):
flood_image_root = os.path.join("train", image_id, "tiles", "flood_label")
water_body_root = os.path.join("train", image_id, "tiles", "water_body_label")
vh_root = os.path.join("train", image_id, "tiles", "vh")
vv_root = os.path.join("train", image_id, "tiles", "vv")
flood_image_paths = list(paths.list_images(flood_image_root))
water_body_paths = list(paths.list_images(water_body_root))
vh_image_paths = list(paths.list_images(vh_root))
vv_image_paths = list(paths.list_images(vv_root))
return flood_image_paths, water_body_paths,\
vh_image_paths, vv_image_paths
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="5dZk7BiimDvC" outputId="3731e75e-1b7b-461c-deb4-8afa6a1a9202"
distribution_dict = {}
for id in tqdm(image_ids):
distribution_dict[id] = {}
flood_image_paths, water_body_paths, vh_image_paths, vv_image_paths = \
get_image_paths(id)
distribution_dict[id]["flood_images"] = len(flood_image_paths)
distribution_dict[id]["water_body_images"] = len(water_body_paths)
distribution_dict[id]["vh_images"] = len(vh_image_paths)
distribution_dict[id]["vv_images"] = len(vv_image_paths)
distribution_df = pd.DataFrame.from_dict(distribution_dict).T
assert len(distribution_df) == len(image_ids)
distribution_df
# + [markdown] id="Z7-1q6k_pmVg"
# No huge distribution skews noticed. But for **`bangladesh_20170314t115609`** there is a mismatch between the number of flood image maps and the number of VV images.
# + [markdown] id="Ly1feNzrw9x_"
# ## Visualization
#
# Now, let's write a utility function that would return the images belonging to the format - `<region>_<datetime>*_x-*_y-*.png`.
#
# It seems like the VV images should be used for predicting flood levels and VH images should be used for predicting water body levels.
#
# <p align="center">
# <img src=https://i.ibb.co/mCZp6X4/image.png></ing>
# </p>
#
# However,
#
# > We expect participants to provide a binary segmentation of the region of interest (ROI), (i.e. 256x256 pixels) as a numpy array with the byte (uint8) data type:
# **1: Flood region, 0: Not flood region**.
# + id="6GGeTAT6xgEd"
# https://stackoverflow.com/a/2669120/7636462
def sorted_nicely(l):
convert = lambda text: int(text) if text.isdigit() else text
alphanum_key = lambda key: [ convert(c) for c in re.split('([0-9]+)', key) ]
return sorted(l, key = alphanum_key)
# + colab={"base_uri": "https://localhost:8080/"} id="ybMpR4Lr1XwU" outputId="0bf007f7-1f4d-4b38-82da-753c8ed58b66"
all_image_paths = sorted_nicely(all_image_paths)
vv_image_paths = [path for path in all_image_paths if ("vv" in path) and ("ipynb_checkpoints" not in path)]
flood_image_paths = [path for path in all_image_paths if ("flood" in path) and ("ipynb_checkpoints" not in path)]
vh_image_paths = [path for path in all_image_paths if ("vh" in path) and ("ipynb_checkpoints" not in path)]
water_body_label_paths = [path for path in all_image_paths if ("water" in path) and ("ipynb_checkpoints" not in path)]
len(flood_image_paths), len(vv_image_paths), len(vh_image_paths), len(water_body_label_paths)
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="56YkE9eO-tV2" outputId="cf9e881c-5afb-4304-f0d8-5a6d5e3f5aaa"
all_image_paths[0]
# + [markdown] id="uSjPRl7h_3fq"
# What is `.ipynb_checkpoints` doing here? 😨
# + colab={"base_uri": "https://localhost:8080/"} id="EgyNRBv5-77w" outputId="ab93dfa3-5757-40ab-dbf6-5723cb505979"
# Verify if we have maintained the order
flood_image_paths[:5], vv_image_paths[:5]
# + colab={"base_uri": "https://localhost:8080/"} id="nMo5X1YC-u-I" outputId="c33991ad-674b-4ce4-e2ba-5a1140c7bbb8"
water_body_label_paths[:5], vh_image_paths[:5]
# + id="wWnesEnH_Myh"
def get_image_id(filename):
return filename.split("/")[1]
# + id="E594rhX5-162"
def show_all_four_images(filenames, titles):
plt.figure(figsize=(20, 10))
images = []
for filename in filenames:
images.append(mpimg.imread(filename))
plt.suptitle(get_image_id(filenames[0]), size=16)
columns = 4
for i, image in enumerate(images):
ax = plt.subplot(len(images)/ columns + 1, columns, i + 1)
ax.set_title(titles[i])
plt.imshow(image)
plt.show()
# + id="wo1PLOaHHnTF"
regex = r"_x-\d+_y-\d+"
compiler = re.compile(regex)
def get_intensity(path):
return compiler.search(path).group()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="kGZrcDtv-_0G" outputId="f900fbc2-7c6b-4ae5-b839-d0cfcb2ea45b"
import random
titles = ["V V","V H" , "Land or water before flood/Water body image" ,"After Flood/flood image"]
random_index = random.sample(range(0, len(vv_image_paths)), 10)
for i in random_index:
# The assertions make sure we are operating on the right pairs
assert get_intensity(vv_image_paths[i]) == get_intensity(flood_image_paths[i])
assert get_intensity(vh_image_paths[i]) == get_intensity(water_body_label_paths[i])
show_all_four_images([vv_image_paths[i], vh_image_paths[i],
water_body_label_paths[i], flood_image_paths[i] ] , titles)
# + [markdown] id="b9xG2zqRMDL6"
# **Some noise found (from an earlier iteration)**:
#
# * https://ibb.co/m6x9f1S
# * https://ibb.co/rfWtJy7
#
# How in an all-white image, any segmentation map is present?
# + [markdown] id="G6auRGru_Xmy"
# ### Displaying the RGB composite
#
# From [here](https://sentinel.esa.int/web/sentinel/user-guides/sentinel-1-sar/product-overview/polarimetry):
#
# > The composite RGB (colour) image on the right was created using the VV channel for red, VH channel for green and the ratio $|VV| / |VH|$ for blue.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="em9A7OGT_cQr" outputId="5a5c6e0f-4fb3-4ed2-8440-30974886fba6"
from PIL import Image
def show_all_combined_images(i, titles):
columns = 3
red, _ , _ = Image.open(vv_image_paths[i]).split()
red = np.asarray(red)
_, green, _ = Image.open(vh_image_paths[i]).split()
green = np.asarray(green)
blue = np.abs(red) / np.abs(green)
blue = (blue * 255).astype(np.uint8)
rgb = Image.fromarray(np.dstack((red,green,blue)))
images = [rgb]
images.append(mpimg.imread(water_body_label_paths[i]))
images.append(mpimg.imread(flood_image_paths[i]))
plt.figure(figsize=(20, 10))
plt.suptitle(get_image_id(vv_image_paths[i]), size=16)
for i, image in enumerate(images):
ax = plt.subplot(len(images)/ columns + 1, columns, i + 1)
ax.set_title(titles[i])
plt.imshow(image)
titles = ["Combined" , "Land or water before flood/Water body image" ,"After Flood/flood image"]
for i in random_index:
show_all_combined_images(i , titles)
# + [markdown] id="3m2FFZNgEoH6"
# ## Observations
#
# * We need to be careful about the way we would shuffle the samples. We likely wouldn't want to just randomly shuffle them. Because if we do so then the continual order of samples for a particular region and timestamp would get broken.
# * We also cannot randomly sample data points for our local validation set. It's much like predicting the next frame for a given sequence of frames. We would want to train models on a sequence of *ordered* frames and use that to infer the next one.
# * Can we simply discard the blank images (all white ones under `Combined` and their respective labels)? I don't see any point in keeping them.
# + [markdown] id="8hfNf05tMk-0"
# ## Some preprocessing
#
# Referred from this [video](https://youtu.be/derOXkPCH80). A PDF is present [here](http://step.esa.int/docs/tutorials/S1TBX%20SAR%20Basics%20Tutorial.pdf).
# + [markdown] id="i-d9KEDlMxrq"
# ### Speckle removal
# + id="5CsEObUJMqOc"
# https://stackoverflow.com/a/39786527/7636462
from scipy.ndimage.filters import uniform_filter
from scipy.ndimage.measurements import variance
def lee_filter(img, size=20):
img_mean = uniform_filter(img, (size, size, size))
img_sqr_mean = uniform_filter(img**2, (size, size, size))
img_variance = img_sqr_mean - img_mean**2
overall_variance = variance(img)
img_weights = img_variance / (img_variance + overall_variance)
img_output = img_mean + img_weights * (img - img_mean)
return img_output
# + id="oPCtW9E-PSNX"
random_index = random.sample(range(0, len(vv_image_paths)), 10)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="HkXQBuSVN_Vn" outputId="cf79231b-09b4-40b6-cab0-b0f550f213f0"
# With Speckle Removal
def show_all_four_images(filenames, titles, speckle=False):
plt.figure(figsize=(20, 10))
images = []
for filename in filenames:
image = mpimg.imread(filename)
if speckle:
lee_filter(image)
images.append(image)
plt.suptitle(get_image_id(filenames[0]), size=16)
columns = 4
for i, image in enumerate(images):
ax = plt.subplot(len(images)/ columns + 1, columns, i + 1)
ax.set_title(titles[i])
plt.imshow(image)
plt.show()
titles = ["V V","V H" , "Land or water before flood/Water body image" ,"After Flood/flood image"]
for i in random_index:
show_all_four_images([vv_image_paths[i], vh_image_paths[i],
water_body_label_paths[i], flood_image_paths[i] ] , titles, True)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="LZDu2KwYOhKa" outputId="7fdc41f0-bf4e-458a-dac5-50ed8e5892ad"
# Without Speckle
for i in random_index:
show_all_four_images([vv_image_paths[i], vh_image_paths[i],
water_body_label_paths[i], flood_image_paths[i] ] , titles, False)
# + [markdown] id="qWZSFxwbP0ZF"
# Seems like the Sentinel-1 images have gone through some speckle removal already. We can confirm this by examining the distribution of the histograms.
| notebooks/Data_Viz.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: hw3
# language: python
# name: hw3
# ---
import pandas as pd
import numpy as np
import os
from sklearn.metrics import roc_curve, auc
import matplotlib.pyplot as plt
from random import uniform
import time
import random
os.chdir('/Users/elizabeth/Desktop/BMI203_HW3/data')
# +
def read_pairs(file):
#Open the file
f = open ( file , 'r')
#Read all the lines
lines = f.read().splitlines()
#Create a list of the paired filenames
files = [line.split()for line in lines]
return files
def read_sequence(file):
#Store protein name
protein_name = file.split('/')[-1][:-3]
#Open the file
f = open ( file , 'r')
#Read all the lines
lines = f.read().splitlines()
#Remove the header
lines = lines[1:]
#Initialize array to store sequence
sequence = ''
for line in lines:
sequence = sequence + line
sequence = sequence.upper()
return protein_name, sequence
def read_scoring_matrix(file):
#Open the file
f = open ( file , 'r')
#Read all the lines
lines = f.read().splitlines()
#Find where the header ends
aa_names_index = 0
while lines[aa_names_index][0] == '#':
aa_names_index = aa_names_index +1
#Remove header
lines = lines[aa_names_index:]
#Initialize DataFrame to store scoring matrix
amino_acids = lines[0].split()
scoring_df = pd.DataFrame(index=amino_acids, columns=amino_acids)
#Store scores into dataframe
for line_index in range(1,len(lines)):
scoring_df.iloc[line_index-1] = lines[line_index].split()
#Convert scores to numbers from strings
scoring_df = scoring_df.apply(pd.to_numeric)
return scoring_df
def smith_waterman_alignment(path_seq_a, path_seq_b, path_scoring_matrix, gap_open, gap_extend):
#Read sequences and scoring matrix
name_a, seq_a = read_sequence(path_seq_a)
name_b, seq_b = read_sequence(path_seq_b)
cost_matrix = read_scoring_matrix(path_scoring_matrix)
#Store value for -Inf
MIN = -float("inf")
"""
We will initialize the values for three matrices:
A - score for aligning seq_a and seq_b up to i,j with a gap in seq_a at position i
B - score for aligning seq_a and seq_b up to i,j with a gap in seq_b at position j
M - score for aligning seq_a and seq_b with a alignment at position i,j
"""
dim_a = len(seq_a)+1
dim_b = len(seq_b)+1
####Initialize matrices
A = np.zeros((dim_a, dim_b))
B = np.zeros((dim_a, dim_b))
M = np.zeros((dim_a, dim_b))
A_direction = {}
B_direction = {}
M_direction = {}
for i in range(dim_a):
A_direction[i,0] = "end"
B_direction[i,0] = "end"
M_direction[i,0] = "end"
for j in range(dim_b):
A_direction[0,j] = "end"
B_direction[0,j] = "end"
M_direction[0,j] = "end"
#Fill in top row and left row for each gap matrix with -Inf because we will not allow
#a gap at the start for our local alignment
for i in range(1,dim_a):
A[i,0] = MIN
B[i,0] = MIN
for j in range(1,dim_b):
A[0,j] = MIN
B[0,j] = MIN
#Make list to keep track of direction
event_gap_a = ["open_gap_a", "extend_gap_a", "open_gap_a_from_b"]
event_gap_b = ["open_gap_b", "extend_gap_b", "open_gap_b_from_a"]
event_match = ["match", "close_gap_a","close_gap_b","end"]
"""
Now we will fill in the values for these three matrices
"""
for i in range(1,dim_a):
for j in range(1,dim_b):
#For A (putting a gap in seq_a at position i), we have three possibilities for how seq_a up to i
#and seq_b up to j-1 (since j is aligning to the gap in seq_a) could have been aligned. We can come from:
# 1) a previous alignment
# 2) a previous gap in seq_a
# 3) a previous gap in seq_b (so a new gap in seq_a)
#dic_A["M":M[i,j-1] + gap_open + gap_extend, "A":gap_extend + A[i,j-1]]
values = [M[i,j-1] - gap_open - gap_extend , A[i,j-1] - gap_extend, B[i,j-1] - gap_open - gap_extend]
A[i,j] = max(values)
A_direction[i,j] = event_gap_a[values.index(A[i,j])]
#For B (putting a gap in seq_b at position j), we have three possibilities for how seq_a up to i-1 (since
# i is aligning with a gap) and seq_b up to j could have been aligned. We can come from:
# 1) a previous alignment
# 2) a previous gap in seq_a (so a new gap in seq_b)
# 3) a previous gap in seq_b
values = [M[i-1,j] - gap_open - gap_extend , B[i-1,j] - gap_extend, A[i-1,j] - gap_open - gap_extend, ]
B[i,j] = max(values)
B_direction[i,j] = event_gap_b[values.index(B[i,j])]
#For M alinging position i and j from seq_a and seq_b respectively we can come from:
# 1) a previous alignment
# 2) a previous gap in seq_a
# 3) a previous gap in seq_b
#Cost for aligning seq_a and seq_b at position i,j (need to account for zero indexing)
#Let 0 be the minimum score in order to create local rather than global optimum alignments
values = [cost_matrix.loc[seq_a[i-1],seq_b[j-1]] + M[i-1,j-1], cost_matrix.loc[seq_a[i-1],seq_b[j-1]] + A[i-1,j-1], cost_matrix.loc[seq_a[i-1],seq_b[j-1]] + B[i-1,j-1],0]
M[i,j] = max(values)
M_direction[i,j] = event_match[values.index(max(values))]
if M.max() == 0:
return "","",0
#Do traceback to get aligned sequence
#Initalize strings to contan alignment
#Find index of max score position in Z
indices = np.where(M == M.max())
#If there are multiple alignments take the first position
index_a = indices[0][0]
index_b = indices[1][0]
#Store first traceback value and initial direction
alignment_seqa, alignment_seqb = seq_a[index_a-1], seq_b[index_b-1]
direction = M_direction[index_a,index_b]
#Do traceback and store sequence
while direction != "end":
#Move in recorded direction and update alligned sequence
if index_a == 1 or index_b == 1:
break
elif direction == 'close_gap_a':
index_a, index_b = index_a-1, index_b-1
alignment_seqa, alignment_seqb = "-"+ alignment_seqa, seq_b[index_b-1]+ alignment_seqb
direction = A_direction[index_a, index_b]
elif direction == 'close_gap_b':
index_a, index_b = index_a-1, index_b-1
alignment_seqa, alignment_seqb = seq_a[index_a-1]+ alignment_seqa, "-"+ alignment_seqb
direction = B_direction[index_a, index_b]
elif direction == 'match':
index_a, index_b = index_a-1, index_b-1
alignment_seqa, alignment_seqb = seq_a[index_a-1] + alignment_seqa, seq_b[index_b-1]+ alignment_seqb
direction = M_direction[index_a, index_b]
elif direction == 'end':
break
elif direction == 'open_gap_a':
index_a, index_b = index_a, index_b-1
alignment_seqa, alignment_seqb = seq_a[index_a-1]+ alignment_seqa,seq_b[index_b-1]+ alignment_seqb
direction = M_direction[index_a, index_b]
elif direction == 'extend_gap_a':
index_a, index_b = index_a, index_b-1
alignment_seqa, alignment_seqb = "-"+ alignment_seqa,seq_b[index_b-1]+ alignment_seqb
direction = A_direction[index_a, index_b]
elif direction == 'open_gap_a_from_b':
index_a, index_b = index_a, index_b-1
alignment_seqa, alignment_seqb = seq_a[index_a-1]+ alignment_seqa,"-"+ alignment_seqb
direction = B_direction[index_a, index_b]
elif direction == 'open_gap_b':
index_a, index_b = index_a-1, index_b
alignment_seqa, alignment_seqb = seq_a[index_a-1]+ alignment_seqa,seq_b[index_b-1]+ alignment_seqb
direction = M_direction[index_a, index_b]
elif direction == 'extend_gap_b':
index_a, index_b = index_a-1, index_b
alignment_seqa, alignment_seqb = seq_a[index_a-1]+ alignment_seqa,"-"+ alignment_seqb
direction = B_direction[index_a, index_b]
elif direction == 'open_gap_b_from_a':
index_a, index_b = index_a-1, index_b
alignment_seqa, alignment_seqb = "-"+ alignment_seqa,seq_b[index_b-1]+ alignment_seqb
direction = A_direction[index_a, index_b]
#Need to remove last match if it actually was making the alignment worse
if cost_matrix.loc[seq_a[index_a-1],seq_b[index_b-1]] < 0:
return alignment_seqa[1:], alignment_seqb[1:], M.max()
else:
return alignment_seqa, alignment_seqb, M.max()
def smith_waterman(path_seq_a, path_seq_b, path_scoring_matrix, gap_open, gap_extend):
#Read sequences and scoring matrix
name_a, seq_a = read_sequence(path_seq_a)
name_b, seq_b = read_sequence(path_seq_b)
cost_matrix = read_scoring_matrix(path_scoring_matrix)
#Store value for -Inf
MIN = -float("inf")
"""
We will initialize the values for three matrices:
A - score for aligning seq_a and seq_b up to i,j with a gap in seq_a at position i
B - score for aligning seq_a and seq_b up to i,j with a gap in seq_b at position j
M - score for aligning seq_a and seq_b with a alignment at position i,j
"""
dim_a = len(seq_a)+1
dim_b = len(seq_b)+1
####Initialize matrices
A = np.zeros((dim_a, dim_b))
B = np.zeros((dim_a, dim_b))
M = np.zeros((dim_a, dim_b))
#Fill in top row and left row for each gap matrix with -Inf because we will not allow
#a gap at the start for our local alignment
for i in range(1,dim_a):
A[i,0] = MIN
B[i,0] = MIN
for j in range(1,dim_b):
A[0,j] = MIN
B[0,j] = MIN
"""
Now we will fill in the values for these three matrices
"""
for i in range(1,dim_a):
for j in range(1,dim_b):
#For A (putting a gap in seq_a at position i), we have three possibilities for how seq_a up to i
#and seq_b up to j-1 (since j is aligning to the gap in seq_a) could have been aligned. We can come from:
# 1) a previous alignment
# 2) a previous gap in seq_a
# 3) a previous gap in seq_b (so a new gap in seq_a)
#dic_A["M":M[i,j-1] + gap_open + gap_extend, "A":gap_extend + A[i,j-1]]
A[i,j] = max(M[i,j-1] - gap_open - gap_extend , A[i,j-1] - gap_extend, B[i,j-1] - gap_open - gap_extend)
#For B (putting a gap in seq_b at position j), we have three possibilities for how seq_a up to i-1 (since
# i is aligning with a gap) and seq_b up to j could have been aligned. We can come from:
# 1) a previous alignment
# 2) a previous gap in seq_a (so a new gap in seq_b)
# 3) a previous gap in seq_b
B[i,j] = max(M[i-1,j] - gap_open - gap_extend , B[i-1,j] - gap_extend, A[i-1,j] - gap_open - gap_extend)
#For M alinging position i and j from seq_a and seq_b respectively we can come from:
# 1) a previous alignment
# 2) a previous gap in seq_a
# 3) a previous gap in seq_b
#Cost for aligning seq_a and seq_b at position i,j (need to account for zero indexing)
#Let 0 be the minimum score in order to create local rather than global optimum alignments
M[i,j] = max(cost_matrix.loc[seq_a[i-1],seq_b[j-1]] + M[i-1,j-1], cost_matrix.loc[seq_a[i-1],seq_b[j-1]] + A[i-1,j-1], cost_matrix.loc[seq_a[i-1],seq_b[j-1]] + B[i-1,j-1],0)
return np.max(M)
def smith_waterman_opt_matrix(path_seq_a, path_seq_b, scoring_matrix, gap_open, gap_extend):
#Read sequences and scoring matrix
name_a, seq_a = read_sequence(path_seq_a)
name_b, seq_b = read_sequence(path_seq_b)
cost_matrix = scoring_matrix
#Store value for -Inf
MIN = -float("inf")
"""
We will initialize the values for three matrices:
A - score for aligning seq_a and seq_b up to i,j with a gap in seq_a at position i
B - score for aligning seq_a and seq_b up to i,j with a gap in seq_b at position j
M - score for aligning seq_a and seq_b with a alignment at position i,j
"""
dim_a = len(seq_a)+1
dim_b = len(seq_b)+1
####Initialize matrices
A = np.zeros((dim_a, dim_b))
B = np.zeros((dim_a, dim_b))
M = np.zeros((dim_a, dim_b))
#Fill in top row and left row for each gap matrix with -Inf because we will not allow
#a gap at the start for our local alignment
for i in range(1,dim_a):
A[i,0] = MIN
B[i,0] = MIN
for j in range(1,dim_b):
A[0,j] = MIN
B[0,j] = MIN
"""
Now we will fill in the values for these three matrices
"""
for i in range(1,dim_a):
for j in range(1,dim_b):
#For A (putting a gap in seq_a at position i), we have three possibilities for how seq_a up to i
#and seq_b up to j-1 (since j is aligning to the gap in seq_a) could have been aligned. We can come from:
# 1) a previous alignment
# 2) a previous gap in seq_a
# 3) a previous gap in seq_b (so a new gap in seq_a)
#dic_A["M":M[i,j-1] + gap_open + gap_extend, "A":gap_extend + A[i,j-1]]
values = [M[i,j-1] - gap_open - gap_extend , A[i,j-1] - gap_extend, B[i,j-1] - gap_open - gap_extend]
A[i,j] = max(values)
#For B (putting a gap in seq_b at position j), we have three possibilities for how seq_a up to i-1 (since
# i is aligning with a gap) and seq_b up to j could have been aligned. We can come from:
# 1) a previous alignment
# 2) a previous gap in seq_a (so a new gap in seq_b)
# 3) a previous gap in seq_b
values = [M[i-1,j] - gap_open - gap_extend , B[i-1,j] - gap_extend, A[i-1,j] - gap_open - gap_extend, ]
B[i,j] = max(values)
#For M alinging position i and j from seq_a and seq_b respectively we can come from:
# 1) a previous alignment
# 2) a previous gap in seq_a
# 3) a previous gap in seq_b
#Cost for aligning seq_a and seq_b at position i,j (need to account for zero indexing)
#Let 0 be the minimum score in order to create local rather than global optimum alignments
values = [cost_matrix.loc[seq_a[i-1],seq_b[j-1]] + M[i-1,j-1], cost_matrix.loc[seq_a[i-1],seq_b[j-1]] + A[i-1,j-1], cost_matrix.loc[seq_a[i-1],seq_b[j-1]] + B[i-1,j-1],0]
M[i,j] = max(values)
return np.max(M)
def smith_waterman_len_adj(path_seq_a, path_seq_b, path_scoring_matrix, gap_open, gap_extend):
#Read sequences and scoring matrix
name_a, seq_a = read_sequence(path_seq_a)
name_b, seq_b = read_sequence(path_seq_b)
cost_matrix = read_scoring_matrix(path_scoring_matrix)
#Store value for -Inf
MIN = -float("inf")
"""
We will initialize the values for three matrices:
A - score for aligning seq_a and seq_b up to i,j with a gap in seq_a at position i
B - score for aligning seq_a and seq_b up to i,j with a gap in seq_b at position j
M - score for aligning seq_a and seq_b with a alignment at position i,j
"""
dim_a = len(seq_a)+1
dim_b = len(seq_b)+1
####Initialize matrices
A = np.zeros((dim_a, dim_b))
B = np.zeros((dim_a, dim_b))
M = np.zeros((dim_a, dim_b))
#Fill in top row and left row for each gap matrix with -Inf because we will not allow
#a gap at the start for our local alignment
for i in range(1,dim_a):
A[i,0] = MIN
B[i,0] = MIN
for j in range(1,dim_b):
A[0,j] = MIN
B[0,j] = MIN
"""
Now we will fill in the values for these three matrices
"""
for i in range(1,dim_a):
for j in range(1,dim_b):
#For A (putting a gap in seq_a at position i), we have three possibilities for how seq_a up to i
#and seq_b up to j-1 (since j is aligning to the gap in seq_a) could have been aligned. We can come from:
# 1) a previous alignment
# 2) a previous gap in seq_a
# 3) a previous gap in seq_b (so a new gap in seq_a)
#dic_A["M":M[i,j-1] + gap_open + gap_extend, "A":gap_extend + A[i,j-1]]
A[i,j] = max(M[i,j-1] - gap_open - gap_extend , A[i,j-1] - gap_extend, B[i,j-1] - gap_open - gap_extend)
#For B (putting a gap in seq_b at position j), we have three possibilities for how seq_a up to i-1 (since
# i is aligning with a gap) and seq_b up to j could have been aligned. We can come from:
# 1) a previous alignment
# 2) a previous gap in seq_a (so a new gap in seq_b)
# 3) a previous gap in seq_b
B[i,j] = max(M[i-1,j] - gap_open - gap_extend , B[i-1,j] - gap_extend, A[i-1,j] - gap_open - gap_extend)
#For M alinging position i and j from seq_a and seq_b respectively we can come from:
# 1) a previous alignment
# 2) a previous gap in seq_a
# 3) a previous gap in seq_b
#Cost for aligning seq_a and seq_b at position i,j (need to account for zero indexing)
#Let 0 be the minimum score in order to create local rather than global optimum alignments
M[i,j] = max(cost_matrix.loc[seq_a[i-1],seq_b[j-1]] + M[i-1,j-1], cost_matrix.loc[seq_a[i-1],seq_b[j-1]] + A[i-1,j-1], cost_matrix.loc[seq_a[i-1],seq_b[j-1]] + B[i-1,j-1],0)
return np.max(M)/min(len(seq_a),len(seq_b))
def score_alignment(seq_a, seq_b, scoring_matrix,gap,ext):
if len(seq_a) != len(seq_b):
return "Please return valid local alignments"
else:
i = 0
score = 0
while i in range(len(seq_a)):
if seq_a[i] != "-" and seq_b[i] != "-":
score = scoring_matrix.loc[seq_a[i],seq_b[i]] + score
elif seq_a[i] == "-" and seq_b[i] != "-":
if seq_a[i-1] != "-":
score = -gap -ext + score
else:
score = -ext + score
elif seq_b[i] == "-": #and seq_a[i] != "-":
if seq_b[i-1] != "-":
score = -gap -ext + score
else:
score = -ext + score
elif seq_b[i] == "-" and seq_a[i] == "-":
if seq_b[i-1] == "-" and seq_a[i-1] == "-":
score = -ext*2 + score
elif seq_b[i-1] != "-" and seq_a[i-1] != "-":
score = -ext*2 + -gap*2 + score
else:
score = -ext*2 + -gap + score
#print(score)
i += 1
return score
def score_performance(pos_sequences,neq_sequences,scoring_matrix, gap,ext):
##Caluclate the performance for the current scoring matrix
pos_scores = [score_alignment(x[0], x[1], scoring_matrix, gap, ext) for x in pos_sequences]
neg_scores = [score_alignment(x[0], x[1], scoring_matrix, gap, ext) for x in neq_sequences]
#Find thresholds for 0, 0.1, 0.2, and 0.3 false positive rate. (there are 50 Pos_pairs)
thresholds = []
thresholds.append(sorted(neg_scores)[-1])
thresholds.append(sorted(neg_scores)[-6])
thresholds.append(sorted(neg_scores)[-11])
thresholds.append(sorted(neg_scores)[-16])
#Calculate true_pos rate at each score
false_pos = []
true_pos = []
for value in thresholds:
false_pos.append(np.sum(np.array(neg_scores) > value)/len(neg_scores))
true_pos.append(np.sum(np.array(pos_scores) > value)/len(pos_scores))
overall_score = np.sum(true_pos)
return overall_score
def score_alignment(seq_a, seq_b, scoring_matrix,gap,ext):
if len(seq_a) != len(seq_b):
return "Please return valid local alignments"
else:
i = 0
score = 0
while i in range(len(seq_a)):
if seq_a[i] != "-" and seq_b[i] != "-":
score = scoring_matrix.loc[seq_a[i],seq_b[i]] + score
elif seq_a[i] == "-" and seq_b[i] != "-":
if seq_a[i-1] != "-":
score = -gap -ext + score
else:
score = -ext + score
elif seq_b[i] == "-": #and seq_a[i] != "-":
if seq_b[i-1] != "-":
score = -gap -ext + score
else:
score = -ext + score
elif seq_b[i] == "-" and seq_a[i] == "-":
if seq_b[i-1] == "-" and seq_a[i-1] == "-":
score = -ext*2 + score
elif seq_b[i-1] != "-" and seq_a[i-1] != "-":
score = -ext*2 + -gap*2 + score
else:
score = -ext*2 + -gap + score
#print(score)
i += 1
return score
# +
pos_pairs = read_pairs('pairs/Pospairs.txt')
neg_pairs = read_pairs('pairs/Negpairs.txt')
gap_test = 2
ext_test = 3
test_a, test_b, score= smith_waterman_alignment(pos_pairs[0][0], pos_pairs[0][1], 'scoring/BLOSUM50', gap_test, ext_test)
print(score)
score_mat = read_scoring_matrix('scoring/BLOSUM50')
score_alignment(test_a, test_b, score_mat,gap_test,ext_test)
# -
smith_waterman(pos_pairs[0][0], pos_pairs[0][1], 'scoring/BLOSUM50', 4, 1)
#Test alignment sequences from EMBOSS for pos_scores[0]
test = 'SLEAAQKSNVTSSWAKASAAWGTAGPEFFMALFDAHDDVFAKFSGLFSGAAKGT---VKNTPEMAAQAQSFKGLVSNWVDNLDNAGALEGQC------KTFAA-------NHKARGISAGQLEAAFKVLSGFMK---SYGG-----DE---GAWTAVAG-ALMGEI'
test_2 = 'SLEHA-KVD-TSNEARQD------GIDLYKHMFENYPP-LRKY---FKSREEYTAEDVQNDPFFAKQGQ--KILLACHV--------L---CATYDDRETFNAYTRELLDRH-AR----DHVHMPPEVWTDFWKLFEEYLGKKTTLDEPTKQAWHEI-GREFAKEI'
score_mat = read_scoring_matrix('scoring/BLOSUM50')
score_alignment(test, test_2, score_mat,4,1)
# Part 1.1
# +
pos_pairs = read_pairs('pairs/Pospairs.txt')
neg_pairs = read_pairs('pairs/Negpairs.txt')
fp_result = []
print(time.time())
for gap in range(1,21):
for ext in range(1,6):
pos_scores = [smith_waterman(x[0], x[1], 'scoring/BLOSUM50', gap, ext) for x in pos_pairs]
neg_scores = [smith_waterman(x[0], x[1], 'scoring/BLOSUM50', gap, ext) for x in neg_pairs]
#Threshold for 0.7 True Positive Rate. (there are 50 Pos_pairs)
threshold = sorted(pos_scores)[14]
false_pos = np.sum(np.array(neg_scores) > threshold)/len(neg_scores)
true_pos = np.sum(np.array(pos_scores) > threshold)/len(pos_scores)
fp_result.append([gap,ext,false_pos,true_pos])
print("finished for ",gap,ext,time.time())
# -
len(fp_result)
# +
pos_pairs = read_pairs('pairs/Pospairs.txt')
neg_pairs = read_pairs('pairs/Negpairs.txt')
start = time.time()
for gap in range(6,21):
for ext in range(1,6):
pos_scores = [smith_waterman(x[0], x[1], 'scoring/BLOSUM50', gap, ext) for x in pos_pairs]
neg_scores = [smith_waterman(x[0], x[1], 'scoring/BLOSUM50', gap, ext) for x in neg_pairs]
#Threshold for 0.7 True Positive Rate. (there are 50 Pos_pairs)
threshold = sorted(pos_scores)[14]
false_pos = np.sum(np.array(neg_scores) > threshold)/len(neg_scores)
true_pos = np.sum(np.array(pos_scores) > threshold)/len(pos_scores)
fp_result.append([gap,ext,false_pos,true_pos])
print("finished for ",gap,ext,time.time()-start)
# -
df = pd.DataFrame(fp_result, columns=["gap_open","gap_ext","FP","TP"])
#Show the best gap-opening and gap_extend penalty
df[df.loc[:,"FP"] == min(df.loc[:,"FP"])]
opt_gap = 5
opt_ext = 3
plt.scatter(df.loc[df["gap_ext"] == 1,:]['gap_open'],df.loc[df["gap_ext"] == 1,:]['FP'], c='g', label='gap_ext = 1')
plt.scatter(df.loc[df["gap_ext"] == 2,:]['gap_open'],df.loc[df["gap_ext"] == 2,:]['FP'], c='b', label='gap_ext = 2')
plt.scatter(df.loc[df["gap_ext"] == 3,:]['gap_open'],df.loc[df["gap_ext"] == 3,:]['FP'], c='yellow', label='gap_ext = 3')
plt.scatter(df.loc[df["gap_ext"] == 4,:]['gap_open'],df.loc[df["gap_ext"] == 4,:]['FP'], c='darkorange', label='gap_ext = 4')
plt.scatter(df.loc[df["gap_ext"] == 5,:]['gap_open'],df.loc[df["gap_ext"] == 5,:]['FP'], c='deeppink', label='gap_ext = 5')
plt.legend(loc='lower right')
plt.xlabel("Gap Open Penalty")
plt.ylabel("False Positive Rate")
plt.show()
df.to_csv('false_positives.csv')
# Part 1.2
# +
#ROC
#Load list of pos_pairs and neg_pairs
pos_pairs = read_pairs('pairs/Pospairs.txt')
neg_pairs = read_pairs('pairs/Negpairs.txt')
#Load list of scoring matrices
scoring_files = ["scoring/BLOSUM50","scoring/BLOSUM62","scoring/PAM100","scoring/PAM250"]
#Usng optimized gap opening penatlty (gap) and gap extension penalty (ext)
gap = opt_gap
ext = opt_ext
scoring_results = {}
fp_result_scoring = {}
#Calculate fpr (for TP of 0.7) and scoring predictions for each scoring matrix
for path in scoring_files:
pos_scores = [smith_waterman(x[0], x[1], path, gap, ext) for x in pos_pairs]
neg_scores = [smith_waterman(x[0], x[1], path, gap, ext) for x in neg_pairs]
#Threshold for 0.7 True Positive Rate. (there are 50 Pos_pairs)
threshold = sorted(pos_scores)[14]
false_pos = np.sum(np.array(neg_scores) > threshold)/len(neg_scores)
true_pos = np.sum(np.array(pos_scores) > threshold)/len(pos_scores)
fp_result_scoring[path] = [false_pos,true_pos]
overall_scores = pos_scores + neg_scores
scoring_results[path] = overall_scores
#Plot ROC curve
# Compute ROC curve and ROC area for each scoring matrix
y_true = [1]*len(pos_pairs) + [0]*len(neg_pairs)
fpr = dict()
tpr = dict()
roc_auc = dict()
for path in scoring_files:
fpr[path], tpr[path], _ = roc_curve(y_true, scoring_results[path])
roc_auc[path] = auc(fpr[path], tpr[path])
colors = ['deeppink','aqua', 'darkorange', 'cornflowerblue']
for path, color in zip(scoring_files, colors):
plt.plot(fpr[path], tpr[path], color=color, lw=2,
label='ROC curve of class {0} (area = {1:0.2f})'
''.format(path, roc_auc[path]))
plt.plot([0, 1], [0, 1], 'k--', lw=2)
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic to all scoring matrices')
plt.legend(loc="lower right")
plt.show()
# -
print(fp_result_scoring)
print(sorted(scoring_results['scoring/PAM100'][0:50])[13:18])
threshold = sorted(scoring_results['scoring/PAM100'][0:50])[13]
# #False positive rate for true positive rate of 0.72
np.sum(np.array(scoring_results['scoring/PAM100'][50:]) > threshold)/len(neg_scores)
# Part 1.3
# +
#ROC
#Load list of pos_pairs and neg_pairs
pos_pairs = read_pairs('pairs/Pospairs.txt')
neg_pairs = read_pairs('pairs/Negpairs.txt')
#Load list of scoring matrices
scoring_files = ["scoring/BLOSUM50","scoring/BLOSUM62","scoring/PAM100","scoring/PAM250"]
#Usng optimized gap opening penatlty (gap) and gap extension penalty (ext)
gap = opt_gap
ext = opt_ext
scoring_results_adj = {}
fp_result_scoring_adj = {}
#Calculate fpr (for TP of 0.7) and scoring predictions for each scoring matrix with the length adjusted score
for path in scoring_files:
pos_scores = [smith_waterman_len_adj(x[0], x[1], path, gap, ext) for x in pos_pairs]
neg_scores = [smith_waterman_len_adj(x[0], x[1], path, gap, ext) for x in neg_pairs]
#Threshold for 0.7 True Positive Rate. (there are 50 Pos_pairs)
threshold = sorted(pos_scores)[14]
false_pos = np.sum(np.array(neg_scores) > threshold)/len(neg_scores)
true_pos = np.sum(np.array(pos_scores) > threshold)/len(pos_scores)
fp_result_scoring_adj[path] = [false_pos,true_pos]
overall_scores = pos_scores + neg_scores
scoring_results_adj[path] = overall_scores
# Compute ROC curve and ROC area for each scoring matrix with the length adjusted score
y_true = [1]*len(pos_pairs) + [0]*len(neg_pairs)
fpr_adj = dict()
tpr_adj = dict()
roc_auc_adj = dict()
for path in scoring_files:
fpr_adj[path], tpr_adj[path], _ = roc_curve(y_true, scoring_results_adj[path])
roc_auc_adj[path] = auc(fpr_adj[path], tpr_adj[path])
#Plot ROC for best matrix
colors = ['orange','yellow']
opt_path = 'scoring/PAM100'
plt.plot(fpr[opt_path], tpr[opt_path], color=colors[0], lw=2,
label='ROC curve of class {0} (area = {1:0.2f})'
''.format(opt_path + " Orig", roc_auc[opt_path]))
plt.plot(fpr_adj[opt_path], tpr_adj[opt_path], color=colors[1], lw=2,
label='ROC curve of class {0} (area = {1:0.2f})'
''.format(opt_path + " Adj", roc_auc_adj[opt_path]))
plt.plot([0, 1], [0, 1], 'k--', lw=2)
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic for original and length adjusted PAM100 matrix')
plt.legend(loc="lower right")
plt.show()
# -
print(roc_auc_adj)
print(fp_result_scoring_adj)
# +
plt.hist(scoring_results['scoring/PAM100'][0:50], bins='auto', alpha = 0.5, label = "Positive Pairs")
plt.hist(scoring_results['scoring/PAM100'][50:], bins='auto', alpha = 0.5, label = "Negative Pairs")
plt.title("Histogram of scores for original algorithm")
plt.legend(loc='upper right')
plt.show()
plt.hist(scoring_results_adj['scoring/PAM100'][0:50], bins='auto', alpha = 0.5, label = "Positive Pairs")
plt.hist(scoring_results_adj['scoring/PAM100'][50:], bins='auto', alpha = 0.5, label = "Negative Pairs")
plt.title("Histogram of scores for length adjusted algorithm")
plt.legend(loc='upper right')
plt.show()
# +
#Determine optimal local alginments for the optimum matrix from Part 1
pos_pairs = read_pairs('pairs/Pospairs.txt')
neg_pairs = read_pairs('pairs/Negpairs.txt')
gap = opt_gap
ext = opt_ext
opt_matrix = 'scoring/PAM100'
opt_matrix_df = read_scoring_matrix(opt_matrix)
pos_seq = []
neg_seq = []
pos_scores = []
neg_scores =[]
for x in pos_pairs:
opt_seq_a,opt_seq_b,score = smith_waterman_alignment(x[0], x[1], opt_matrix, gap, ext)
if score_alignment(opt_seq_a, opt_seq_b, opt_matrix_df, gap, ext) != score:
print (x[0], x[1])
break
pos_seq.append([opt_seq_a,opt_seq_b])
pos_scores.append(score)
print("Finished", x[0], x[1])
print("finished positives")
for x in neg_pairs:
opt_seq_a,opt_seq_b,score = smith_waterman_alignment(x[0], x[1], opt_matrix, gap, ext)
if score_alignment(opt_seq_a, opt_seq_b, opt_matrix_df, gap, ext) != score:
print (x[0], x[1])
break
neg_seq.append([opt_seq_a,opt_seq_b])
neg_scores.append(score)
print("Finished", x[0], x[1])
# -
start = time.time()
path = 'scoring/PAM100'
#i = np.random.randint(0,high=50)
test_a, test_b, score= smith_waterman_alignment(pos_pairs[0][0], pos_pairs[0][1], path, 5, 3)
print(score)
score_mat = read_scoring_matrix(path)
print(score_alignment(test_a, test_b, score_mat, 5, 3))
print(time.time()-start)
test_a
test_b
#EMBOSS test case
#['sequences/prot-0031.fa', 'sequences/prot-0034.fa']
a = "VIEKAD---NAAQVK---DALTKMRAAALDAQKATP-----PKLEDKSPDS-PEM-KD---FR---HGFDILVGQIDDALKL---ANEGKVKEAQAA-AEQLKTTRNAYHQKYR"
b = "VV-KGQAPYDAAQIKANVEVLKTL--TAL------PWAAFGPGTE--GGDARPEIWSDAASFKQKQQAF-----Q-DNIVKLSAAADAGDLDKLRAAFGD-VGASCKACHDAYR"
score_mat = read_scoring_matrix(path)
score_alignment(a, b, score_mat, 4, 1)
#EMBOSS test case
#['sequences/prot-0102.fa', 'sequences/prot-0098.fa']
a = "IDRDAVLNMWQ-QGLGASHISKTMNIARSTVYK"
b = "IQR-GV-SPSQAQGLGSNLVT---EV-R--VYN"
score_mat = read_scoring_matrix(path)
score_alignment(a, b, score_mat, 4, 1)
# Part 2.1 and 2.2
test = read_scoring_matrix('scoring/PAM100')
score_performance(pos_seq, neg_seq,test, 5,3)
test = read_scoring_matrix('scoring/BLOSUM50')
score_performance(pos_seq, neg_seq,test, 5,3)
# +
def optimize_scoring_matrix(alignments_pos, alignments_neg, starting_matrix_path, gap_open, gap_ext, num_iterations):
#Record start time
start = time.time()
#Load scoring matrix
score_mat = read_scoring_matrix(starting_matrix_path)
score_mat = score_mat.astype(np.float64)
#Store column names
residues = score_mat.columns.tolist()
#Intialize the iteration matrices and iteration score lists
iteration_mat = [score_mat]*10
#Initialize best iteration scores
starting_score = score_performance(alignments_pos, alignments_neg, score_mat, gap_open, gap_ext)
iteration_scores = [starting_score]*10
print("Loaded initial matrix and scores")
#Perform optimization with pool of matrices
#Take top 10 scoring matrices
#Mutate each one nine times to create pool of 100 matrices
#Repeat
iteration_counter = [0]
iteration_score_counter = []
iteration_score_counter.append([np.mean(iteration_scores),np.std(iteration_scores),np.max(iteration_scores)])
for i in range(num_iterations):
#Find top 10 matrices
best_mat_scores = sorted(iteration_scores)[-10:]
best_mat_indices = [iteration_scores.index(x) for x in best_mat_scores]
#Make list of 100 matrices with each of the best ten repeated ten times
new_matrices = []
new_scores = []
for j in range(10):
new_matrices = new_matrices + [iteration_mat[best_mat_indices[j]]]*10
new_scores = new_scores + [iteration_scores[best_mat_indices[j]]]*10
iteration_mat = new_matrices
iteration_scores = new_scores
#Mutate the 9 copies but keep one of each of the originals
for k in list(set(range(100))-set([0,10,20,30,40,50,60,70,80,90])):
rand_adj = pd.DataFrame(np.zeros((24,24)), columns = residues, index = residues)
for b in range(24):
for c in range(b,24):
rand_adj.iloc[b,c] += uniform(-1,1)
rand_adj.iloc[c,b] = rand_adj.iloc[b,c]
iteration_mat[k] = iteration_mat[k] + rand_adj
iteration_scores[k] = score_performance(alignments_pos, alignments_neg, iteration_mat[k], gap_open, gap_ext)
iteration_counter.append(i+1)
iteration_score_counter.append([np.mean(iteration_scores),np.std(iteration_scores), np.max(iteration_scores)])
print("Finished iteration", i+1)
new_matrix_score = max(iteration_scores)
new_matrix = iteration_mat[iteration_scores.index(max(iteration_scores))]
end = time.time()
total_time = (end-start)/60
return iteration_mat, iteration_scores, new_matrix, new_matrix_score, starting_score, iteration_counter, iteration_score_counter, total_time
# -
start = time.time()
output = optimize_scoring_matrix(pos_seq, neg_seq, 'scoring/PAM100', opt_gap, opt_ext, 100)
print(time.time()-start)
output[6]
print(score_performance(pos_seq,neg_seq,output[2],5,3) == output[3])
print(output[3])
def score_performance(pos_sequences,neq_sequences,scoring_matrix, gap,ext):
##Caluclate the performance for the current scoring matrix
pos_scores = [score_alignment(x[0], x[1], scoring_matrix, gap, ext) for x in pos_sequences]
neg_scores = [score_alignment(x[0], x[1], scoring_matrix, gap, ext) for x in neq_sequences]
#Find thresholds for 0, 0.1, 0.2, and 0.3 false positive rate. (there are 50 Pos_pairs)
thresholds = []
thresholds.append(sorted(neg_scores)[-1])
thresholds.append(sorted(neg_scores)[-6])
thresholds.append(sorted(neg_scores)[-11])
thresholds.append(sorted(neg_scores)[-16])
#Calculate true_pos rate at each score
false_pos = []
true_pos = []
for value in thresholds:
false_pos.append(np.sum(np.array(neg_scores) > value)/len(neg_scores))
true_pos.append(np.sum(np.array(pos_scores) > value)/len(pos_scores))
overall_score = np.sum(true_pos)
return overall_score
df_pos = pd.DataFrame({'pos':pos_seq})
df_pos.to_csv('tests/pos_seq.csv')
df_neg = pd.DataFrame({'neg':neg_seq})
df_neg.to_csv('tests/neg_seq.csv')
def write_optimal_matrix(filename, k, path, opt_matrix, opt_matrix_score, original_score,time):
"""
Write the optimal scoring matrix out to a file.
Input: a filename and the output from optimize_scoring_matrix
Output: none
"""
out = open(filename, 'w')
out.write("# Optimal Matrix after %d iterations on %s scoring matrix\n" % (k, path))
out.write("# Time (min) %d \n" % time)
out.write("# Original matrix score %d \n" % original_score)
out.write("# New matrix score %d \n" % opt_matrix_score)
out.close()
opt_matrix.to_csv(filename, index=None, sep=' ', mode='a')
# +
write_optimal_matrix('/Users/elizabeth/Desktop/BMI203_HW3/data/optimization/opt_PAM100',100,'scoring/PAM100',output[2],output[3],output[4],output[-1])
# -
write_optimal_matrix('/Users/elizabeth/Desktop/BMI203_HW3/data/optimization/opt_PAM100_rounded',100,'scoring/PAM100',output[2].round(2),output[3],output[4],output[-1])
# +
print(score_performance(pos_seq,neg_seq,output[2],5,3) == output[3])
print(output[3])
pd.set_option('display.max_columns', None) # or 1000
pd.set_option('display.max_rows', None) # or 1000
pd.set_option('display.max_colwidth', -1)
pd.set_option('display.width', 4)
output[2].round(2)
# +
pd.set_option('display.max_columns', None) # or 1000
pd.set_option('display.max_rows', None) # or 1000
pd.set_option('display.max_colwidth', -1)
pd.set_option('display.width', 4)
path = 'scoring/PAM100'
scoring_matrix = read_scoring_matrix(path)
scoring_matrix
# -
def score_performance_all(pos_sequences,neq_sequences,scoring_matrix, gap,ext):
##Caluclate the performance for the current scoring matrix
pos_scores = [score_alignment(x[0], x[1], scoring_matrix, gap, ext) for x in pos_sequences]
neg_scores = [score_alignment(x[0], x[1], scoring_matrix, gap, ext) for x in neq_sequences]
#Find thresholds for 0, 0.1, 0.2, and 0.3 false positive rate. (there are 50 Pos_pairs)
thresholds = []
thresholds.append(sorted(neg_scores)[-1])
thresholds.append(sorted(neg_scores)[-6])
thresholds.append(sorted(neg_scores)[-11])
thresholds.append(sorted(neg_scores)[-16])
#Calculate true_pos rate at each score
false_pos = []
true_pos = []
for value in thresholds:
false_pos.append(np.sum(np.array(neg_scores) > value)/len(neg_scores))
true_pos.append(np.sum(np.array(pos_scores) > value)/len(pos_scores))
overall_score = np.sum(true_pos)
return overall_score,false_pos,true_pos
test = read_scoring_matrix('optimization/opt_PAM100')
print(score_performance_all(pos_seq,neg_seq,test,5,3))
# Part 2.3
# +
#ROC
#Plot ROC for initial alignments for original matrix and optimized matrix
scoring_results = {}
fp_result_scoring = {}
#Original
path = 'scoring/PAM100'
scoring_matrix = read_scoring_matrix(path)
pos_scores = [score_alignment(x[0], x[1], scoring_matrix, 5, 3) for x in pos_seq]
neg_scores = [score_alignment(x[0], x[1], scoring_matrix, 5, 3) for x in neg_seq]
threshold = sorted(pos_scores)[14]
false_pos = np.sum(np.array(neg_scores) > threshold)/len(neg_scores)
true_pos = np.sum(np.array(pos_scores) > threshold)/len(pos_scores)
fp_result_scoring[path] = [false_pos,true_pos]
overall_scores = pos_scores + neg_scores
scoring_results[path] = overall_scores
#Optimized
path = 'optimization/opt_PAM100'
scoring_matrix = read_scoring_matrix(path)
pos_scores = [score_alignment(x[0], x[1], scoring_matrix, 5, 3) for x in pos_seq]
neg_scores = [score_alignment(x[0], x[1], scoring_matrix, 5, 3) for x in neg_seq]
threshold = sorted(pos_scores)[14]
false_pos = np.sum(np.array(neg_scores) > threshold)/len(neg_scores)
true_pos = np.sum(np.array(pos_scores) > threshold)/len(pos_scores)
fp_result_scoring[path] = [false_pos,true_pos]
overall_scores = pos_scores + neg_scores
scoring_results[path] = overall_scores
#Optimized New alignments
path = 'optimization/opt_PAM100'
pos_pairs = read_pairs('pairs/Pospairs.txt')
neg_pairs = read_pairs('pairs/Negpairs.txt')
pos_scores = [smith_waterman(x[0], x[1], path, 5, 3) for x in pos_pairs]
neg_scores = [smith_waterman(x[0], x[1], path, 5, 3) for x in neg_pairs]
threshold = sorted(pos_scores)[14]
false_pos = np.sum(np.array(neg_scores) > threshold)/len(neg_scores)
true_pos = np.sum(np.array(pos_scores) > threshold)/len(pos_scores)
fp_result_scoring[path + 'calc'] = [false_pos,true_pos]
overall_scores = pos_scores + neg_scores
scoring_results[path + 'calc'] = overall_scores
y_true = [1]*len(pos_pairs) + [0]*len(neg_pairs)
fpr = dict()
tpr = dict()
roc_auc = dict()
for path in list(scoring_results.keys()):
fpr[path], tpr[path], _ = roc_curve(y_true, scoring_results[path])
roc_auc[path] = auc(fpr[path], tpr[path])
colors = ['deeppink','aqua', 'darkorange']
i = 0
for path in list(scoring_results.keys()):
plt.plot(fpr[path], tpr[path], color=colors[i], lw=2,
label='ROC curve of class {0} (area = {1:0.2f})'
''.format(path, roc_auc[path]))
i = i+1
plt.plot([0, 1], [0, 1], 'k--', lw=2)
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('ROC for original,optimized, and re-aligned sequences with optimized matrix scoring for PAM100 matrix')
plt.legend(loc="lower right")
plt.show()
# +
y_true = [1]*len(pos_pairs) + [0]*len(neg_pairs)
fpr = dict()
tpr = dict()
roc_auc = dict()
for path in list(scoring_results.keys()):
fpr[path], tpr[path], _ = roc_curve(y_true, scoring_results[path])
roc_auc[path] = auc(fpr[path], tpr[path])
colors = ['deeppink','aqua', 'darkorange']
i = 0
for path in list(scoring_results.keys()):
plt.plot(fpr[path], tpr[path], color=colors[i], lw=2,
label='ROC curve of class {0} (area = {1:0.2f})'
''.format(path, roc_auc[path]))
i = i+1
plt.plot([0, 1], [0, 1], 'k--', lw=2)
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('ROC for original,optimized, and re-aligned sequences with optimized matrix scoring for PAM100 matrix')
plt.legend(loc="lower right")
plt.show()
# -
print(fp_result_scoring)
# +
#Plot Optimization Performance
mean_values = [output[6][i][0] for i in range(101)]
std_values = [output[6][i][1] for i in range(101)]
max_values = [output[6][i][2] for i in range(101)]
iteration_values = list(range(101))
plt.errorbar(x=iteration_values, y=mean_values, yerr = std_values, marker='^', ecolor = "black")
plt.xlabel('Iteration Number')
plt.ylabel('Mean of the scores of matrices for optimizing PAM100')
plt.show()
# -
# Part 2.3
# +
start = time.time()
output_MATIO = optimize_scoring_matrix(pos_seq, neg_seq, 'optimization/MATIO', opt_gap, opt_ext, 100)
print(time.time()-start)
# -
print(score_performance(pos_seq,neg_seq,output_MATIO[2],5,3) == output_MATIO[3])
print(output_MATIO[3])
# +
pd.set_option('display.max_columns', None) # or 1000
pd.set_option('display.max_rows', None) # or 1000
pd.set_option('display.max_colwidth', -1)
pd.set_option('display.width', 4)
output_MATIO[2].round(2)
# +
pd.set_option('display.max_columns', None) # or 1000
pd.set_option('display.max_rows', None) # or 1000
pd.set_option('display.max_colwidth', -1)
pd.set_option('display.width', 4)
path = 'optimization/MATIO'
scoring_matrix = read_scoring_matrix(path)
scoring_matrix
# +
write_optimal_matrix('/Users/elizabeth/Desktop/BMI203_HW3/data/optimization/opt_MATIO',100,'optimization/MATIO',output_MATIO[2],output_MATIO[3],output_MATIO[4],output_MATIO[-1])
# +
#ROC
#Plot ROC for initial alignments for original matrix and optimized matrix
scoring_results = {}
fp_result_scoring = {}
#Original
path = 'optimization/MATIO'
scoring_matrix = read_scoring_matrix(path)
pos_scores = [score_alignment(x[0], x[1], scoring_matrix, 5, 3) for x in pos_seq]
neg_scores = [score_alignment(x[0], x[1], scoring_matrix, 5, 3) for x in neg_seq]
threshold = sorted(pos_scores)[14]
false_pos = np.sum(np.array(neg_scores) > threshold)/len(neg_scores)
true_pos = np.sum(np.array(pos_scores) > threshold)/len(pos_scores)
fp_result_scoring[path] = [false_pos,true_pos]
overall_scores = pos_scores + neg_scores
scoring_results[path] = overall_scores
#Optimized
path = 'optimization/opt_MATIO'
scoring_matrix = read_scoring_matrix(path)
pos_scores = [score_alignment(x[0], x[1], scoring_matrix, 5, 3) for x in pos_seq]
neg_scores = [score_alignment(x[0], x[1], scoring_matrix, 5, 3) for x in neg_seq]
threshold = sorted(pos_scores)[14]
false_pos = np.sum(np.array(neg_scores) > threshold)/len(neg_scores)
true_pos = np.sum(np.array(pos_scores) > threshold)/len(pos_scores)
fp_result_scoring[path] = [false_pos,true_pos]
overall_scores = pos_scores + neg_scores
scoring_results[path] = overall_scores
#Optimized New alignments
path = 'optimization/opt_MATIO'
pos_pairs = read_pairs('pairs/Pospairs.txt')
neg_pairs = read_pairs('pairs/Negpairs.txt')
pos_scores = [smith_waterman(x[0], x[1], path, 5, 3) for x in pos_pairs]
neg_scores = [smith_waterman(x[0], x[1], path, 5, 3) for x in neg_pairs]
threshold = sorted(pos_scores)[14]
false_pos = np.sum(np.array(neg_scores) > threshold)/len(neg_scores)
true_pos = np.sum(np.array(pos_scores) > threshold)/len(pos_scores)
fp_result_scoring[path + ' calc'] = [false_pos,true_pos]
overall_scores = pos_scores + neg_scores
scoring_results[path + ' calc'] = overall_scores
y_true = [1]*len(pos_pairs) + [0]*len(neg_pairs)
fpr = dict()
tpr = dict()
roc_auc = dict()
for path in list(scoring_results.keys()):
fpr[path], tpr[path], _ = roc_curve(y_true, scoring_results[path])
roc_auc[path] = auc(fpr[path], tpr[path])
colors = ['deeppink','aqua', 'darkorange']
i = 0
for path in list(scoring_results.keys()):
plt.plot(fpr[path], tpr[path], color=colors[i], lw=2,
label='ROC curve of class {0} (area = {1:0.2f})'
''.format(path, roc_auc[path]))
i = i+1
plt.plot([0, 1], [0, 1], 'k--', lw=2)
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('ROC for original,optimized, and re-aligned sequences with optimized matrix scoring for MATIO matrix')
plt.legend(loc="lower right")
plt.show()
# -
print(fp_result_scoring)
print(output_MATIO[3])
print(output_MATIO[4])
# +
#Plot Optimization Performance
mean_values = [output_MATIO[6][i][0] for i in range(101)]
std_values = [output_MATIO[6][i][1] for i in range(101)]
max_values = [output_MATIO[6][i][2] for i in range(101)]
iteration_values = list(range(101))
plt.errorbar(x=iteration_values, y=mean_values, yerr = std_values, marker='^', ecolor = "black")
plt.xlabel('Iteration Number')
plt.ylabel('Mean of the scores of matrices for optimizing MATIO')
plt.show()
# -
| HW_3_in_progress.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tensorflow_gpuenv
# language: python
# name: tensorflow_gpuenv
# ---
# +
# here we first need to implement the teacher,
# a Wide Res Net in Keras that is able to reach more
# than 92% of accuracy on CIFAR10
# but first let's learn how keras works...
# +
# from the official keras documentation: https://keras.io/examples/mnist_cnn/
from __future__ import print_function
import keras
from keras.datasets import mnist
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten
from keras.layers import Conv2D, MaxPooling2D
from keras import backend as K
batch_size = 128
num_classes = 10
epochs = 12
# input image dimensions
img_rows, img_cols = 28, 28
# the data, split between train and test sets
(x_train, y_train), (x_test, y_test) = mnist.load_data()
if K.image_data_format() == 'channels_first':
x_train = x_train.reshape(x_train.shape[0], 1, img_rows, img_cols)
x_test = x_test.reshape(x_test.shape[0], 1, img_rows, img_cols)
input_shape = (1, img_rows, img_cols)
else:
x_train = x_train.reshape(x_train.shape[0], img_rows, img_cols, 1)
x_test = x_test.reshape(x_test.shape[0], img_rows, img_cols, 1)
input_shape = (img_rows, img_cols, 1)
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
x_train /= 255
x_test /= 255
print('x_train shape:', x_train.shape)
print(x_train.shape[0], 'train samples')
print(x_test.shape[0], 'test samples')
# convert class vectors to binary class matrices
y_train = keras.utils.to_categorical(y_train, num_classes)
y_test = keras.utils.to_categorical(y_test, num_classes)
#model definition
model = Sequential()
model.add(Conv2D(32, kernel_size=(3, 3),
activation='relu',
input_shape=input_shape))
model.add(Conv2D(64, (3, 3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(num_classes, activation='softmax'))
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adadelta(),
metrics=['accuracy'])
model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(x_test, y_test))
score = model.evaluate(x_test, y_test, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
# +
# if i use the last for CIFAR10
from __future__ import print_function
import keras
from keras.datasets import mnist
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten
from keras.layers import Conv2D, MaxPooling2D
from keras import backend as K
from keras.datasets import cifar10
from keras.callbacks import LearningRateScheduler
import numpy as np
def lr_schedule(epoch):
lrate = 0.001
if epoch > 75:
lrate = 0.0005
if epoch > 100:
lrate = 0.0003
return lrate
batch_size = 128
num_classes = 10
epochs = 6
# input image dimensions
img_rows, img_cols = 32, 32
# the data, split between train and test sets
(x_train, y_train), (x_test, y_test) = cifar10.load_data()
if K.image_data_format() == 'channels_first':
x_train = x_train.reshape(x_train.shape[0], 3, img_rows, img_cols)
x_test = x_test.reshape(x_test.shape[0], 3, img_rows, img_cols)
input_shape = (3, img_rows, img_cols)
else:
x_train = x_train.reshape(x_train.shape[0], img_rows, img_cols, 3)
x_test = x_test.reshape(x_test.shape[0], img_rows, img_cols, 3)
input_shape = (img_rows, img_cols, 3)
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
#old normalization
#x_train /= 255
#x_test /= 255
# new normalization with z-score
mean = np.mean(x_train,axis=(0,1,2,3))
std = np.std(x_train,axis=(0,1,2,3))
x_train = (x_train-mean)/(std+1e-7)
x_test = (x_test-mean)/(std+1e-7)
print('x_train shape:', x_train.shape)
print(x_train.shape[0], 'train samples')
print(x_test.shape[0], 'test samples')
# convert class vectors to binary class matrices
y_train = keras.utils.to_categorical(y_train, num_classes)
y_test = keras.utils.to_categorical(y_test, num_classes)
#model definition
model = Sequential()
model.add(Conv2D(32, kernel_size=(5, 5),
activation='relu',
input_shape=input_shape))
model.add(Conv2D(64, (3, 3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(num_classes, activation='softmax'))
# define optimizer
opt_rms = keras.optimizers.rmsprop(lr=0.001,decay=1e-6)
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=opt_rms,
metrics=['accuracy'])
model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(x_test, y_test),
callbacks=[LearningRateScheduler(lr_schedule)])
#save to disk
model_json = model.to_json()
with open('model.json', 'w') as json_file:
json_file.write(model_json)
model.save_weights('model.h5')
# final evaluation on test
score = model.evaluate(x_test, y_test, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
# +
# load the model we just trained
# if i use the last for CIFAR10
from __future__ import print_function
import keras
from keras.datasets import mnist
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten
from keras.layers import Conv2D, MaxPooling2D
from keras import backend as K
from keras.datasets import cifar10
from keras.callbacks import LearningRateScheduler
from keras.models import model_from_json
from keras.models import load_model
import numpy as np
batch_size = 128
num_classes = 10
epochs = 6
# input image dimensions
img_rows, img_cols = 32, 32
# the data, split between train and test sets
(x_train, y_train), (x_test, y_test) = cifar10.load_data()
if K.image_data_format() == 'channels_first':
x_train = x_train.reshape(x_train.shape[0], 3, img_rows, img_cols)
x_test = x_test.reshape(x_test.shape[0], 3, img_rows, img_cols)
input_shape = (3, img_rows, img_cols)
else:
x_train = x_train.reshape(x_train.shape[0], img_rows, img_cols, 3)
x_test = x_test.reshape(x_test.shape[0], img_rows, img_cols, 3)
input_shape = (img_rows, img_cols, 3)
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
#old normalization
#x_train /= 255
#x_test /= 255
# new normalization with z-score
mean = np.mean(x_train,axis=(0,1,2,3))
std = np.std(x_train,axis=(0,1,2,3))
x_train = (x_train-mean)/(std+1e-7)
x_test = (x_test-mean)/(std+1e-7)
print('x_train shape:', x_train.shape)
print(x_train.shape[0], 'train samples')
print(x_test.shape[0], 'test samples')
# convert class vectors to binary class matrices
y_train = keras.utils.to_categorical(y_train, num_classes)
y_test = keras.utils.to_categorical(y_test, num_classes)
# Model reconstruction from JSON file
with open('model.json', 'r') as f:
model = model_from_json(f.read())
# Load weights into the new model
model.load_weights('model.h5')
# define optimizer
opt_rms = keras.optimizers.rmsprop(lr=0.001,decay=1e-6)
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=opt_rms,
metrics=['accuracy'])
# final evaluation on test
score = model.evaluate(x_test, y_test, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
# -
| Our_code/Keras(not working)/older failed tests/old_train_teacher.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: dev
# kernelspec:
# display_name: py365_for_class
# language: python
# name: py365_for_class
# ---
# UCSD Data Science Bootcamp, HW 21 ML
# <NAME>, 4/28/20
# Update sklearn to prevent version mismatches
# !pip install sklearn --upgrade
# install joblib. This will be used to save your model.
# Restart your kernel after installing
# !pip install joblib
import pandas as pd
import numpy as np
# # Read the CSV and Perform Basic Data Cleaning
df = pd.read_csv("exoplanet_data.csv")
# Drop the null columns where all values are null
df = df.dropna(axis='columns', how='all')
# Drop the null rows
df = df.dropna()
df.head()
# Remove "CANDIDATE" outcomes from the dataset
print(len(df))
# df = df.loc[((df['koi_disposition'] == 'CONFIRMED') | (df['koi_disposition'] == 'FALSE POSITIVE')) , :]
print(len(df))
df.columns
# # Select your features (columns)
# Set features. This will also be used as your x values.
# selected_features = df[['names', 'of', 'selected', 'features', 'here']]
selected_features = df[ ['koi_fpflag_nt', 'koi_fpflag_ss', 'koi_fpflag_co',
'koi_fpflag_ec', 'koi_period', 'koi_time0bk', 'koi_impact','koi_duration', 'koi_depth', 'koi_prad',
'koi_teq', 'koi_insol', 'koi_model_snr', 'koi_tce_plnt_num', 'koi_steff', 'koi_slogg',
'koi_srad', 'ra', 'dec', 'koi_kepmag'] ]
# # Create a Train Test Split
#
# Use `koi_disposition` for the y values
# +
# y = df["koi_disposition"].map({'CANDIDATE':0.0, 'CONFIRMED':1.0, 'FALSE POSITIVE':0.0}).values.reshape(-1, 1)
y = df["koi_disposition"]
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(selected_features, y, random_state=1)
y
# -
print("X_train length: ", len(X_train))
print("X_test length: ", len(X_test))
print("y_train length: ", len(y_train))
print("y_test length: ", len(y_test))
X_train.head()
# # Pre-processing
#
# Scale the data using the MinMaxScaler and perform some feature selection
# Scale your data
from sklearn.preprocessing import StandardScaler
X_scaler = StandardScaler().fit(X_train)
# y_scaler = StandardScaler().fit(y_train)
X_train_scaled = X_scaler.transform(X_train)
X_test_scaled = X_scaler.transform(X_test)
# y_train_scaled = y_scaler.transform(y_train)
# y_test_scaled = y_scaler.transform(y_test)
# # Logistic Regression
from sklearn.linear_model import LogisticRegression
classifier = LogisticRegression(random_state=99)
classifier
y_train = y_train.ravel()
model2 = classifier.fit(X_train_scaled, y_train)
# # Train the Model
#
#
print(f"Training Data Score: {model2.score(X_train_scaled, y_train)}")
print(f"Testing Data Score: {model2.score(X_test_scaled, y_test)}")
# # Hyperparameter Tuning
#
# Use `GridSearchCV` to tune the model's parameters
# Create the GridSearchCV model
from sklearn.model_selection import GridSearchCV
param_grid = {'C': [0.1, 0.5, 1, 1.5, 2, 3, 5, 7, 10],
'penalty': ["l1", "l2"]}
grid = GridSearchCV(model2, param_grid, verbose=3)
# Train the model with GridSearch
grid.fit(X_train_scaled, y_train)
print(grid.best_params_)
print(grid.best_score_)
# # Save the Model
# save your model by updating "your_name" with your name
# and "your_model" with your model variable
# be sure to turn this in to BCS
# if joblib fails to import, try running the command to install in terminal/git-bash
import joblib
filename = 'alexis-logistic_regression.sav'
joblib.dump(grid, filename)
# +
# models = [logistic_regression, random_forest, svm]
# for model in models:
# model.fit()
# model.score()
# -
from sklearn.ensemble import RandomForestClassifier
# +
lr = LogisticRegression(random_state=99)
rf = RandomForestClassifier(random_state=99)
models = [lr, rf]
names = ['Logistic Regression', 'Random Forest']
# -
for name, model in zip(names, models):
model.fit(X_train_scaled, y_train)
score = model.score(X_test_scaled, y_test)
print(name, score)
| model_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # Verify Source and Target Databases and the Ingestion Pipeline <a name="top"></a>
#
# ## Table of Contents:
#
# 1. [Overview](#Overview)
# 2. [Aurora MySQL as Source Database](#Aurora-MySQL-as-Source-Database)
# 3. [Amazon Redshift as Data Warehouse](#Amazon-Redshift-as-Data-Warehouse)
# 4. [AWS DMS as Near Real-Time Ingestion Pipeline](#AWS-DMS-as-Near-Real-Time-Ingestion-Pipeline)
# 5. [Simulate Inserts](#Simulate-Inserts)
# ## Overview
# [(Back to the top)](#top)
#
# We will use this notebook to verify that our Aurora MySQL and Amazon Redshift Databases are up and running.
#
# The Aurora MySQL database will serve as the source of transactions, and the Amazon Redshift database will serve as the target Data Warehouse. We will execute inserts from this notebook as well to simulate new transactions.
#
# <img src="../resources/module2_architecture_diagram.png" alt="Module2 Architecture Diagram]" style="width: 1000px;"/>
#
#
# ## Aurora MySQL as Source Database
# [(Back to the top)](#top)
#
# Let's first test connectivity to our database:
# +
import MySQLdb,random,time
from tqdm.notebook import trange, tqdm
host = '###mysql_host###'
user = 'master'
password = '<PASSWORD>'
port = 3306
db = 'salesdb'
conn = MySQLdb.Connection(
host=host,
user=user,
passwd=password,
port=port,
db=db
)
# -
# Let's run some SQL statements. We will use the following helper functions to execute SQL statements:
# +
def execute_sql(sql):
results=[]
conn.query(sql)
result = conn.store_result()
for i in range(result.num_rows()):
r = result.fetch_row()
#print(r)
results.append(r)
return results
def execute_dml(sql):
conn.query(sql)
rowcount = conn.affected_rows()
print ("Rows updated: %d"%rowcount)
conn.commit()
# -
execute_sql("show tables")
# This is a generic SALES OLTP schema. Of the tables above, the SALES_ORDER_DETAIL is the one for which we will be inserting records.
# ## Amazon Redshift as Data Warehouse
# [(Back to the top)](#top)
#
# Let's test connectivity to our target datawarehouse:
# +
import psycopg2
import warnings
warnings.filterwarnings('ignore')
rs_host='###redshift_host###'
rs_dbname='sales_analytics_dw'
rs_user = 'awsuser'
rs_password = '<PASSWORD>'
rs_port = 5439
con=psycopg2.connect(dbname=rs_dbname, host=rs_host, port=rs_port, user=rs_user, password=rs_password)
# -
# Now that the connectivity to the Amazon Redshift database is working fine, let's load the schema to the Amazon Redshift database:
# +
def load_redshift_schemas(conn, scriptFileName):
with open (scriptFileName, "r") as scriptfile:
contents=scriptfile.read()
commands=[]
for sql in str(contents).split(";")[:-1]:
commands.append(sql)
for sql in commands:
cursor = con.cursor()
cursor.execute(sql)
cursor.close()
# commit the changes
con.commit()
cursor = con.cursor()
cursor.execute("Select distinct tablename from PG_TABLE_DEF where schemaname = 'public'")
rows = cursor.fetchall()
for row in rows:
print (" ", row)
cursor.close()
load_redshift_schemas(con,'redshift-schema.sql')
# -
# ## AWS DMS as Near Real-Time Ingestion Pipeline
# [(Back to the top)](#top)
#
# In this step we will execute a full load of data from this database to Amazon S3 using AWS DMS:
#
# - Navigate to the DMS Console by clicking on Services -> DMS.
# - Locate the menu item Conversion & migration->Database migration tasks from the left-hand panel of the DMS Console.
# - Select the only Replication Task item and click on the button Actions -> Restart/Resume to start this task.
# - You can monitor the progress of this task by clicking on the task link and viewing the 'Table Statistics' tab.
# ## Simulate Inserts
# [(Back to the top)](#top)
#
# Let's perform some Inserts to our data. We will use the helper function below to perform the inserts.
# +
def insert_orders(order_id,new_order_id,max_line_id):
print (new_order_id)
execute_dml(f"insert into SALES_ORDER( ORDER_ID, SITE_ID,ORDER_DATE,SHIP_MODE ) select {new_order_id}, SITE_ID,ORDER_DATE,SHIP_MODE from SALES_ORDER where ORDER_ID={order_id}")
execute_dml(f"insert into SALES_ORDER_DETAIL( ORDER_ID, LINE_ID,LINE_NUMBER,PRODUCT_ID,QUANTITY,UNIT_PRICE,DISCOUNT,SUPPLY_COST,TAX,ORDER_DATE ) select {new_order_id}, {max_line_id}+LINE_ID,LINE_NUMBER,PRODUCT_ID,QUANTITY,UNIT_PRICE,DISCOUNT,SUPPLY_COST,TAX,ORDER_DATE from SALES_ORDER_DETAIL where ORDER_ID={order_id}")
def generate_orders(n):
new_order_id=execute_sql('select max(order_id) FROM SALES_ORDER')[0][0][0]
max_line_id=execute_sql('select max(line_id) FROM SALES_ORDER_DETAIL')[0][0][0]
print (f"max_line_id : {max_line_id}")
for i in tqdm(range(n)):
order_id=random.randint(1,29000)
new_order_id +=1
insert_orders(order_id,new_order_id,max_line_id)
# -
generate_orders(100)
# We can view the updates that AWS DMS has pushed through using the 'Table Statistics' tab for the Replication task within the AWS DMS Console.
#
# <div class="alert alert-block alert-info"><b>Note:</b> Please keep this notebook open as we move to the 2nd notebook in this Module to execute the AWS Glue incremental ETL jobs. We will execute the cell above again when we need to insert more data.</div>
| notebooks/module2/1_Verify_Source_and_Target_Databases_and_the_Ingestion_Pipeline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:PythonData]
# language: python
# name: conda-env-PythonData-py
# ---
import os
from os import chdir
import numpy as np
import pandas as pd
import csv
file_path = "citi_Consolid/"
file_path
newdir = os.listdir(fileYear)
newdir
target_file = '2017_Output.csv'
path_and_target = file_path + target_file
path_and_target
books_df = pd.read_csv(path_and_target, encoding="utf-8")
books_df.head()
| python_tools/column_Data_CleanUp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import lhsmdu
import matplotlib.pyplot as plt
import numpy as np
import time
def simpleaxis(axes, every=False):
if not isinstance(axes, (list, np.ndarray)):
axes = [axes]
for ax in axes:
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
if every:
ax.spines['bottom'].set_visible(False)
ax.spines['left'].set_visible(False)
ax.get_xaxis().tick_bottom()
ax.get_yaxis().tick_left()
ax.set_title('')
dimensions = [2**x for x in xrange(1,6)]
samples = [4**x for x in xrange(1,6)]
timeMatrix = np.zeros(shape=(len(samples),len(dimensions)))
for i, sample in enumerate(samples):
for j, dims in enumerate(dimensions):
print(sample, dims)
start = time.time()
a = lhsmdu.sample(dims, sample)
end = time.time()
timeMatrix[i,j] = end - start
timeMatrix
for j, row in enumerate(timeMatrix):
print(row, dimensions)
fig, ax = plt.subplots()
ax.plot(dimensions, row, '.-')
ax.set_xlabel("Dimensions")
ax.set_ylabel("Time (s)")
simpleaxis(ax)
ax.set_title("Samples = {}".format(samples[j]))
plt.savefig("Runtimes_samples={}".format(samples[j]))
plt.close()
for j, row in enumerate(timeMatrix.T):
print(row)
fig, ax = plt.subplots()
ax.loglog(samples, row, '.-')
ax.set_xlabel("Samples")
ax.set_ylabel("Time (s)")
simpleaxis(ax)
ax.set_title("Dims = {}".format(dimensions[j]))
plt.savefig("Runtimes_dims={}".format(dimensions[j]))
plt.close()
| lhsmdu/benchmark/Testing lhsmdu times.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# -*- coding: utf-8 -*-
"""
Created on Sat Apr 25 22:26:08 2020
@author: <NAME>
"""
import numpy as np
import sklearn
import sklearn.datasets
import sklearn.linear_model
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# 
# 
# #### Forward Computation (Vectorized Equations) :
#
#
#
# $$ Z^l = (W^l)^T . a^{l-1} + b^l$$
# Here $ a^0 = X^T ; a^l = \sigma '(Z^l) $
#
#
# Loss for $ X_n$ $$ Loss_n = \frac{1}{2} \sum_{k=1}^C (y_k - a_k^L)^2$$
#
# #### Backprogation (Vectorized Equations) :
# $$\delta^ l = W^{l+1} . \delta^{l+1} \oplus \sigma'(Z^{l}) $$ where $ \delta^ l = \frac{\delta Loss}{\delta Z^{l}}$ and $\oplus $ denotes elementwise multiplication
#
#
# $$ \delta^L = (a^{L} - y^{L}) \oplus \sigma'(Z^{L})$$ where $L$ is the last layer
#
#
# $$ \frac{ \delta Loss}{\delta b^l} = \delta^l$$
#
#
# $$ \frac{\delta Loss}{\delta W^l} = a^{l-1} . (\delta ^l)^T $$
#
#
# 
# +
def dsigmoid(x):
return np.exp(x)/ (1.0 + np.exp(x))**2
def sigmoid(x):
return 1.0/ (1.0 + np.exp(-x))
class MLP:
def __init__(self):
self.W = [None]
self.A = [None]
self.Z = [None]
self.b = [None]
self.layers = 0
self.i_nodes = 0
def add_input_layer(self,n_nodes):
self.A[0] = np.empty(n_nodes,dtype=object)
self.i_nodes = n_nodes
def add_layer(self,n_nodes):
if(self.layers == 0 ):
self.W.append(np.random.randn(self.i_nodes,n_nodes))
else:
self.W.append(np.random.randn(self.W[self.layers].shape[1],n_nodes))
self.b.append(np.zeros((n_nodes, 1)))
self.layers += 1
def forward(self,X):
self.A = [None]
self.Z = [None]
self.A[0] = X
L = self.layers
for l in range(1,L+1): # 1 tp L
self.Z.append(np.dot(self.W[l].T,self.A[l-1]) + self.b[l]) #Z[l] created
self.A.append(sigmoid(self.Z[l])) #A[l] created
def back_prop(self,X,Y):
self.A[0] = X
L = self.layers
m = X.shape[1]
self.dZ = [None for _ in range(L+1)]
self.dW = [None for _ in range(L+1)]
self.db = [None for _ in range(L+1)]
self.dZ[L] = np.multiply((self.A[L] - Y),dsigmoid(self.Z[L]))
self.dW[L] = (1/m) * np.dot(self.A[L-1],self.dZ[L].T)
self.db[L] = (1/m) * np.sum(self.dZ[L], axis=1, keepdims=True)
for l in range(L-1,0,-1):
self.dZ[l] = np.multiply(np.dot(self.W[l+1],self.dZ[l+1]),
dsigmoid(self.Z[l]))
self.dW[l] = (1/m) * np.dot(self.A[l-1],self.dZ[l].T)
self.db[l] = (1/m) * np.sum(self.dZ[l], axis=1, keepdims=True)
def train(self, X, Y, epochs=10000, learning_rate=1.2):
""" Complete process of learning, alternates forward pass,
backward pass and parameters update """
self.losses = []
m = X.shape[0]
for e in range(epochs):
L = self.layers
self.forward(X.T)
loss = np.sum((Y.T - self.A[L])**2)/ m
self.back_prop(X.T, Y.T)
self.losses.append(loss)
for l in range(1,L+1):
self.W[l] -= learning_rate * self.dW[l]
self.b[l] -= learning_rate * self.db[l]
if e % 1000 == 0:
print("Loss ", e+1, " = ", loss)
# +
def load_data():
N = 200
gq = sklearn.datasets.make_gaussian_quantiles(mean=None, cov=0.7,
n_samples=N, n_features=2,
n_classes=2, shuffle=True,
random_state=None)
return gq
gaussian_quantiles = load_data()
X, Y = gaussian_quantiles
# Input Data
plt.figure("Input Data")
plt.scatter(X[:, 0], X[:, 1], c=Y, s=40, cmap=plt.cm.Spectral)
plt.show()
Y = Y.reshape(len(Y),1)
# +
nn = MLP()
nn.add_input_layer(X.shape[1])
nn.add_layer(4)
nn.add_layer(Y.shape[1])
nn.train(X, Y)
plt.plot(range(len(nn.losses)),nn.losses)
plt.xlabel("Epochs")
plt.ylabel("Loss Function")
plt.title("Loss v/s Epoch")
plt.show()
| Deep-Learning/GeneralizedMLP.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import cadquery as cq
# These can be modified rather than hardcoding values for each dimension.
circle_radius = 50.0 # Radius of the plate
thickness = 13.0 # Thickness of the plate
rectangle_width = 13.0 # Width of rectangular hole in cylindrical plate
rectangle_length = 19.0 # Length of rectangular hole in cylindrical plate
# Extrude a cylindrical plate with a rectangular hole in the middle of it.
# 1. Establishes a workplane that an object can be built on.
# 1a. Uses the named plane orientation "front" to define the workplane, meaning
# that the positive Z direction is "up", and the negative Z direction
# is "down".
# 2. The 2D geometry for the outer circle is created at the same time as the
# rectangle that will create the hole in the center.
# 2a. The circle and the rectangle will be automatically centered on the
# workplane.
# 2b. Unlike some other functions like the hole(), circle() takes
# a radius and not a diameter.
# 3. The circle and rectangle are extruded together, creating a cylindrical
# plate with a rectangular hole in the center.
# 3a. circle() and rect() could be changed to any other shape to completely
# change the resulting plate and/or the hole in it.
result = cq.Workplane("front").circle(circle_radius) \
.rect(rectangle_width, rectangle_length) \
.extrude(thickness)
# Displays the result of this script
show_object(result)
| examples/Ex004_Extruded_Cylindrical_Plate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="xBF9RPBhw2we"
# ### Dataset Reading
# + id="pN4tMIn1w2wg" executionInfo={"status": "ok", "timestamp": 1603710119565, "user_tz": -330, "elapsed": 2528, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="4d03416e-2531-4353-cfa6-deb1d75dcca2" colab={"base_uri": "https://localhost:8080/", "height": 289}
import pandas as pd
data = pd.read_excel('drive/My Drive/Constraint_Competition_Dataset/Constraint_Covid-19_English_Train.xlsx')
pd.set_option('display.max_colwidth',150)
data.head()
# + id="O9ABoWjOw2wl" executionInfo={"status": "ok", "timestamp": 1603710120260, "user_tz": -330, "elapsed": 3202, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="ef039553-665c-4bbb-b9cf-f55328b9c0c1" colab={"base_uri": "https://localhost:8080/", "height": 35}
data.shape
# + id="JSKI3CX6w2wp" executionInfo={"status": "ok", "timestamp": 1603710120262, "user_tz": -330, "elapsed": 3186, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="396e503b-9d56-4428-9d7e-4a45b0cb863f" colab={"base_uri": "https://localhost:8080/", "height": 87}
print(data.dtypes)
# + [markdown] id="XNsif5VGw2ws"
# ### Making of "label" Variable
# + id="gwE60IAxw2ws" executionInfo={"status": "ok", "timestamp": 1603710120264, "user_tz": -330, "elapsed": 3171, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="477443de-b88b-49d9-d049-855065401982" colab={"base_uri": "https://localhost:8080/", "height": 121}
label = data['label']
label.head()
# + [markdown] id="ShrD5Y7ew2wv"
# ### Checking Dataset Balancing
# + id="kFui_Mz3w2wv" executionInfo={"status": "ok", "timestamp": 1603710121127, "user_tz": -330, "elapsed": 4018, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="679dcc0a-e796-4415-85a3-0bb5355796b2" colab={"base_uri": "https://localhost:8080/", "height": 349}
print(label.value_counts())
import matplotlib.pyplot as plt
label.value_counts().plot(kind='bar', color='blue')
# + [markdown] id="MRSdKLNiw2wx"
# ### Convering label into "0" or "1"
# + id="0ESnvF3Vw2wy" executionInfo={"status": "ok", "timestamp": 1603710121128, "user_tz": -330, "elapsed": 4003, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="a7807220-ee46-4728-9d3d-df8431d867ac" colab={"base_uri": "https://localhost:8080/", "height": 35}
import numpy as np
classes_list = ["fake","real"]
label_index = data['label'].apply(classes_list.index)
final_label = np.asarray(label_index)
print(final_label[:10])
# + id="NSuVpENKGBWU" executionInfo={"status": "ok", "timestamp": 1603710121999, "user_tz": -330, "elapsed": 4869, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
from keras.utils.np_utils import to_categorical
label_twoDimension = to_categorical(final_label, num_classes=2)
# + id="GtlQzqdpGMBM" executionInfo={"status": "ok", "timestamp": 1603710122000, "user_tz": -330, "elapsed": 4841, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="6e2bbdb3-b77f-4750-a45a-5d8f520bf530" colab={"base_uri": "https://localhost:8080/", "height": 191}
print(label_twoDimension[:10])
# + [markdown] id="s2JSVKo3w2w0"
# ### Making of "text" Variable
# + id="-VK4ScnGw2w1" executionInfo={"status": "ok", "timestamp": 1603710122001, "user_tz": -330, "elapsed": 4815, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="a8f0672d-0928-4971-c8b7-aa168c44eb0c" colab={"base_uri": "https://localhost:8080/", "height": 228}
text = data['tweet']
text.head(10)
# + [markdown] id="tbKm17HIw2w3"
# ### Dataset Pre-processing
# + id="_Sf_xhO6w2w7" executionInfo={"status": "ok", "timestamp": 1603710122002, "user_tz": -330, "elapsed": 4808, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
import re
def text_clean(text):
''' Pre process and convert texts to a list of words '''
text=text.lower()
# Clean the text
text = re.sub(r"[^A-Za-z0-9^,!.\/'+-=]", " ", text)
text = re.sub(r"what's", "what is ", text)
text = re.sub(r"I'm", "I am ", text)
text = re.sub(r"\'s", " ", text)
text = re.sub(r"\'ve", " have ", text)
text = re.sub(r"can't", "cannot ", text)
text = re.sub(r"wouldn't", "would not ", text)
text = re.sub(r"shouldn't", "should not ", text)
text = re.sub(r"shouldn", "should not ", text)
text = re.sub(r"didn", "did not ", text)
text = re.sub(r"n't", " not ", text)
text = re.sub(r"i'm", "i am ", text)
text = re.sub(r"\'re", " are ", text)
text = re.sub(r"\'d", " would ", text)
text = re.sub(r"\'ll", " will ", text)
text = re.sub('https?://\S+|www\.\S+', "", text)
text = re.sub(r",", " ", text)
text = re.sub(r"\.", " ", text)
text = re.sub(r"!", " ! ", text)
text = re.sub(r"\/", " ", text)
text = re.sub(r"\^", " ^ ", text)
text = re.sub(r"\+", " + ", text)
text = re.sub(r"\-", " - ", text)
text = re.sub(r"\=", " = ", text)
text = re.sub(r"'", " ", text)
text = re.sub(r"(\d+)(k)", r"\g<1>000", text)
text = re.sub(r":", " : ", text)
text = re.sub(r" e g ", " eg ", text)
text = re.sub(r" b g ", " bg ", text)
text = re.sub(r" u s ", " american ", text)
text = re.sub(r"\0s", "0", text)
text = re.sub(r" 9 11 ", "911", text)
text = re.sub(r"e - mail", "email", text)
text = re.sub(r"j k", "jk", text)
text = re.sub(r"\s{2,}", " ", text)
text = re.sub(r"[0-9]", "", text)
# text = re.sub(r"rt", " ", text)
return text
# + id="5_JQL5rRw2xA" executionInfo={"status": "ok", "timestamp": 1603710122463, "user_tz": -330, "elapsed": 5263, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
clean_text = text.apply(lambda x:text_clean(x))
# + id="A_uqquBZw2xE" executionInfo={"status": "ok", "timestamp": 1603710122465, "user_tz": -330, "elapsed": 5245, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="3cc14c6d-8c89-4080-e51f-9655e632deeb" colab={"base_uri": "https://localhost:8080/", "height": 228}
clean_text.head(10)
# + [markdown] id="AGYA06eJw2xJ"
# ### Removing stopwords
# + id="JBLDOKifw2xK" executionInfo={"status": "ok", "timestamp": 1603710123565, "user_tz": -330, "elapsed": 6325, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="401ee3a3-a21e-41bd-8387-dab328a4b9ed" colab={"base_uri": "https://localhost:8080/", "height": 52}
import nltk
from nltk.corpus import stopwords
nltk.download('stopwords')
def stop_words_removal(text1):
text1=[w for w in text1.split(" ") if w not in stopwords.words('english')]
return " ".join(text1)
# + id="dwSLSw3Nw2xN" executionInfo={"status": "ok", "timestamp": 1603710140499, "user_tz": -330, "elapsed": 23256, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
clean_text_ns=clean_text.apply(lambda x: stop_words_removal(x))
# + id="OFjJCsd_w2xQ" executionInfo={"status": "ok", "timestamp": 1603710140505, "user_tz": -330, "elapsed": 23246, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="10d7df6f-7332-4579-b8a9-c89f7a176700" colab={"base_uri": "https://localhost:8080/", "height": 228}
print(clean_text_ns.head(10))
# + [markdown] id="Vxq3KDt4w2xS"
# ### Lemmatization
# + id="FlGoDlLmw2xT" executionInfo={"status": "ok", "timestamp": 1603710140506, "user_tz": -330, "elapsed": 23230, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="ea927bc3-c9a7-48c8-82f7-da321ae09bf3" colab={"base_uri": "https://localhost:8080/", "height": 70}
"""# Lemmatization
import nltk
nltk.download('wordnet')
from nltk.stem import WordNetLemmatizer
lemmatizer = WordNetLemmatizer()
def word_lemmatizer(text):
lem_text = "".join([lemmatizer.lemmatize(i) for i in text])
return lem_text"""
# + id="desz-r2qw2xW" executionInfo={"status": "ok", "timestamp": 1603710140507, "user_tz": -330, "elapsed": 23215, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="07e427da-925d-4c1b-c3df-e14e378e1b88" colab={"base_uri": "https://localhost:8080/", "height": 35}
"""clean_text_lem = clean_text_ns.apply(lambda x : word_lemmatizer(x))"""
# + id="OuhsiibOw2xY" executionInfo={"status": "ok", "timestamp": 1603710140508, "user_tz": -330, "elapsed": 23200, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="c1e05e2d-96b9-44cd-b44c-af41f172a93a" colab={"base_uri": "https://localhost:8080/", "height": 35}
"""print(clean_text_lem.head(10))"""
# + [markdown] id="96IyUsaow2xa"
# ### Stemming
# + id="2TuWAy4bw2xb" executionInfo={"status": "ok", "timestamp": 1603710140509, "user_tz": -330, "elapsed": 23198, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
# Stemming
from nltk.stem import PorterStemmer
stemmer = PorterStemmer()
def word_stemmer(text):
stem_text = "".join([stemmer.stem(i) for i in text])
return stem_text
# + id="ivl__lJWw2xe" executionInfo={"status": "ok", "timestamp": 1603710140510, "user_tz": -330, "elapsed": 23196, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
clean_text_stem = clean_text_ns.apply(lambda x : word_stemmer(x))
# + id="qoMbymPmw2xf" executionInfo={"status": "ok", "timestamp": 1603710140511, "user_tz": -330, "elapsed": 23175, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="d7b44579-d0bd-4eb2-c2e0-4f7cbccec77d" colab={"base_uri": "https://localhost:8080/", "height": 228}
print(clean_text_stem.head(10))
# + id="0UFWzUEcw2xh" executionInfo={"status": "ok", "timestamp": 1603710140512, "user_tz": -330, "elapsed": 23173, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
# final_text = [x for x in clean_text_lem if len(x) > 3]
# + id="15kD9mAWw2xj" executionInfo={"status": "ok", "timestamp": 1603710140513, "user_tz": -330, "elapsed": 23171, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
#print(final_text)
# + [markdown] id="LyORidvKw2xl"
# ### Tokenization using "keras"
# + id="feW2fI8Dw2xl" executionInfo={"status": "ok", "timestamp": 1603710141450, "user_tz": -330, "elapsed": 24104, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
import keras
import tensorflow
from keras.preprocessing.text import Tokenizer
tok_all = Tokenizer(filters='!"#$%&\'()*+,-./:;<=>?@[\\]^_`{|}~', lower=True, char_level = False)
tok_all.fit_on_texts(clean_text_stem)
# + [markdown] id="pVf7lAKJw2xo"
# ### Making Vocab for words
# + id="LtBxjGZKw2xo" executionInfo={"status": "ok", "timestamp": 1603710141452, "user_tz": -330, "elapsed": 24084, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="26125a3f-6918-4051-947c-5c92c19add2e" colab={"base_uri": "https://localhost:8080/", "height": 35}
vocabulary_all = len(tok_all.word_counts)
print(vocabulary_all)
# + id="PKAhcecYw2xr" executionInfo={"status": "ok", "timestamp": 1603710141454, "user_tz": -330, "elapsed": 24067, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="d52d1d1d-dee4-46a5-de8b-8a5551046e42" colab={"base_uri": "https://localhost:8080/", "height": 55}
l = tok_all.word_index
print(l)
# + [markdown] id="wLKyeIYHw2xu"
# ### encoding or sequencing
# + id="5tTNFeyrw2xu" executionInfo={"status": "ok", "timestamp": 1603710141457, "user_tz": -330, "elapsed": 24050, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="db971d76-d053-49de-b245-3dbae9518529" colab={"base_uri": "https://localhost:8080/", "height": 52}
encoded_clean_text_stem = tok_all.texts_to_sequences(clean_text_stem)
print(clean_text_stem[1])
print(encoded_clean_text_stem[1])
# + [markdown] id="ao425zSrw2xw"
# ### Pre-padding
# + id="mJB28ImAw2xw" executionInfo={"status": "ok", "timestamp": 1603710141459, "user_tz": -330, "elapsed": 24048, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
from keras.preprocessing import sequence
max_length = 100
padded_clean_text_stem = sequence.pad_sequences(encoded_clean_text_stem, maxlen=max_length, padding='pre')
# + [markdown] id="lEigFn5fWFAs"
# # Test Data Pre-processing
# + [markdown] id="4zQ1QbtFWX_J"
# # Data test Reading
# + id="F0wlDEHwWOlx" executionInfo={"status": "ok", "timestamp": 1603710142028, "user_tz": -330, "elapsed": 24599, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="211dcf2d-239b-4a34-f9d7-6c42b294611b" colab={"base_uri": "https://localhost:8080/", "height": 254}
data_t = pd.read_excel('drive/My Drive/Constraint_Competition_Dataset/Constraint_Covid-19_English_Val.xlsx')
pd.set_option('display.max_colwidth',150)
data_t.head()
# + id="W5bwz_-dWyui" executionInfo={"status": "ok", "timestamp": 1603710142029, "user_tz": -330, "elapsed": 24582, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="67224520-bdac-4419-95e4-0df2f8874ae4" colab={"base_uri": "https://localhost:8080/", "height": 35}
data_t.shape
# + id="ntkVP_FiW4vn" executionInfo={"status": "ok", "timestamp": 1603710142030, "user_tz": -330, "elapsed": 24567, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="2b72d4bb-e613-4e9a-ae89-069480b9f492" colab={"base_uri": "https://localhost:8080/", "height": 87}
print(data_t.dtypes)
# + [markdown] id="Ocyn5IEDXAr7"
# # Making of "label" Variable
# + id="bAglc2pzXDpJ" executionInfo={"status": "ok", "timestamp": 1603710142032, "user_tz": -330, "elapsed": 24553, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="9ba03232-67df-4594-b563-ef3c7771cc2a" colab={"base_uri": "https://localhost:8080/", "height": 121}
label_t = data_t['label']
label_t.head()
# + [markdown] id="VVxcyv1uYhUV"
# # Checking Dataset Balancing
# + id="2GJE9j_OW5kG" executionInfo={"status": "ok", "timestamp": 1603710142033, "user_tz": -330, "elapsed": 24535, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="e0c98463-665e-48b9-eae4-b86d5a924445" colab={"base_uri": "https://localhost:8080/", "height": 347}
print(label_t.value_counts())
import matplotlib.pyplot as plt
label_t.value_counts().plot(kind='bar', color='red')
# + [markdown] id="Kq3obUM1Y3v3"
# # Convering label into "0" or "1"
# + id="0V7LGxK_ZA4S" executionInfo={"status": "ok", "timestamp": 1603710142034, "user_tz": -330, "elapsed": 24516, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="56660bfc-3995-45d8-e1ce-030f7b8c6aa3" colab={"base_uri": "https://localhost:8080/", "height": 35}
import numpy as np
classes_list_t = ["fake","real"]
label_t_index = data_t['label'].apply(classes_list_t.index)
final_label_t = np.asarray(label_t_index)
print(final_label_t[:10])
# + id="4Ve8y_srZA75" executionInfo={"status": "ok", "timestamp": 1603710142035, "user_tz": -330, "elapsed": 24513, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
from keras.utils.np_utils import to_categorical
label_twoDimension_t = to_categorical(final_label_t, num_classes=2)
# + id="3rmVyCfKZSxz" executionInfo={"status": "ok", "timestamp": 1603710142035, "user_tz": -330, "elapsed": 24495, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="809996cc-3acb-40dc-a50c-f111b9b0df84" colab={"base_uri": "https://localhost:8080/", "height": 191}
print(label_twoDimension_t[:10])
# + [markdown] id="R5NMHXF6ZZJj"
# # Making of "text" Variable
# + id="BFFgaFBHZomG" executionInfo={"status": "ok", "timestamp": 1603710142036, "user_tz": -330, "elapsed": 24480, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="3eebf62c-1c98-4c77-e7f0-f631b833db9a" colab={"base_uri": "https://localhost:8080/", "height": 228}
text_t = data_t['tweet']
text_t.head(10)
# + [markdown] id="wdok08rOZwro"
# # **Dataset Pre-processing**
# 1. Remove unwanted words
# 2. Stopwords removal
# 3. Stemming
# 4. Tokenization
# 5. Encoding or Sequencing
# 6. Pre-padding
# + [markdown] id="QrxT9sK5bUs3"
# ### 1. Removing Unwanted Words
# + id="eapxovvvavlO" executionInfo={"status": "ok", "timestamp": 1603710142037, "user_tz": -330, "elapsed": 24477, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
import re
def text_clean(text):
''' Pre process and convert texts to a list of words '''
text=text.lower()
# Clean the text
text = re.sub(r"[^A-Za-z0-9^,!.\/'+-=]", " ", text)
text = re.sub(r"what's", "what is ", text)
text = re.sub(r"I'm", "I am ", text)
text = re.sub(r"\'s", " ", text)
text = re.sub(r"\'ve", " have ", text)
text = re.sub(r"can't", "cannot ", text)
text = re.sub(r"wouldn't", "would not ", text)
text = re.sub(r"shouldn't", "should not ", text)
text = re.sub(r"shouldn", "should not ", text)
text = re.sub(r"didn", "did not ", text)
text = re.sub(r"n't", " not ", text)
text = re.sub(r"i'm", "i am ", text)
text = re.sub(r"\'re", " are ", text)
text = re.sub(r"\'d", " would ", text)
text = re.sub(r"\'ll", " will ", text)
text = re.sub('https?://\S+|www\.\S+', "", text)
text = re.sub(r",", " ", text)
text = re.sub(r"\.", " ", text)
text = re.sub(r"!", " ! ", text)
text = re.sub(r"\/", " ", text)
text = re.sub(r"\^", " ^ ", text)
text = re.sub(r"\+", " + ", text)
text = re.sub(r"\-", " - ", text)
text = re.sub(r"\=", " = ", text)
text = re.sub(r"'", " ", text)
text = re.sub(r"(\d+)(k)", r"\g<1>000", text)
text = re.sub(r":", " : ", text)
text = re.sub(r" e g ", " eg ", text)
text = re.sub(r" b g ", " bg ", text)
text = re.sub(r" u s ", " american ", text)
text = re.sub(r"\0s", "0", text)
text = re.sub(r" 9 11 ", "911", text)
text = re.sub(r"e - mail", "email", text)
text = re.sub(r"j k", "jk", text)
text = re.sub(r"\s{2,}", " ", text)
text = re.sub(r"[0-9]", "", text)
# text = re.sub(r"rt", " ", text)
return text
# + id="ZKXhURU5a0q-" executionInfo={"status": "ok", "timestamp": 1603710142038, "user_tz": -330, "elapsed": 24474, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
clean_text_t = text_t.apply(lambda x:text_clean(x))
# + id="4R6Paqqia0y_" executionInfo={"status": "ok", "timestamp": 1603710142039, "user_tz": -330, "elapsed": 24457, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="6a42b6b6-2d6b-4cb9-ca18-d39d8bbe632e" colab={"base_uri": "https://localhost:8080/", "height": 228}
clean_text_t.head(10)
# + [markdown] id="lyxeJ7xtbB5-"
# ### 2. Removing Stopwords
# + id="yfdc4WLNbIYP" executionInfo={"status": "ok", "timestamp": 1603710142040, "user_tz": -330, "elapsed": 24440, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="d2917ddc-fea3-4f4a-dfa7-f64ef2f1a5f0" colab={"base_uri": "https://localhost:8080/", "height": 52}
import nltk
from nltk.corpus import stopwords
nltk.download('stopwords')
def stop_words_removal(text1):
text1=[w for w in text1.split(" ") if w not in stopwords.words('english')]
return " ".join(text1)
# + id="7lH4FtPtbfmc" executionInfo={"status": "ok", "timestamp": 1603710148308, "user_tz": -330, "elapsed": 30704, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
clean_text_t_ns=clean_text_t.apply(lambda x: stop_words_removal(x))
# + id="xSzxQQE0bfpw" executionInfo={"status": "ok", "timestamp": 1603710148317, "user_tz": -330, "elapsed": 30692, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="a52f6b8d-6faf-4db8-9515-ddc928b0406d" colab={"base_uri": "https://localhost:8080/", "height": 228}
print(clean_text_t_ns.head(10))
# + [markdown] id="9VkXLxaMbpqb"
# ### 3. Stemming
# + id="2gEVoc0fbu1m" executionInfo={"status": "ok", "timestamp": 1603710148318, "user_tz": -330, "elapsed": 30689, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
# Stemming
from nltk.stem import PorterStemmer
stemmer = PorterStemmer()
def word_stemmer(text):
stem_text = "".join([stemmer.stem(i) for i in text])
return stem_text
# + id="RnIAjbL7bvon" executionInfo={"status": "ok", "timestamp": 1603710148320, "user_tz": -330, "elapsed": 30687, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
clean_text_t_stem = clean_text_t_ns.apply(lambda x : word_stemmer(x))
# + id="hywyHMQ8bz9B" executionInfo={"status": "ok", "timestamp": 1603710148321, "user_tz": -330, "elapsed": 30665, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="b3a55775-c476-49af-e503-32686c1e1942" colab={"base_uri": "https://localhost:8080/", "height": 228}
print(clean_text_t_stem.head(10))
# + [markdown] id="gNW4AywXb4ZL"
# ### 4. Tokenization
# + id="F-79JOmgb_io" executionInfo={"status": "ok", "timestamp": 1603710148323, "user_tz": -330, "elapsed": 30663, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
import keras
import tensorflow
from keras.preprocessing.text import Tokenizer
tok_test = Tokenizer(filters='!"#$%&\'()*+,-./:;<=>?@[\\]^_`{|}~', lower=True, char_level = False)
tok_test.fit_on_texts(clean_text_t_stem)
# + id="4YCYJtTKcKe-" executionInfo={"status": "ok", "timestamp": 1603710148324, "user_tz": -330, "elapsed": 30643, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="6bcaddd8-0023-46c8-b1de-1e01b1efe48e" colab={"base_uri": "https://localhost:8080/", "height": 35}
vocabulary_all_test = len(tok_test.word_counts)
print(vocabulary_all_test)
# + id="9UCJEGCMcOri" executionInfo={"status": "ok", "timestamp": 1603710148325, "user_tz": -330, "elapsed": 30623, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="657c102c-0c1f-4f17-c4b6-067e1b73e38a" colab={"base_uri": "https://localhost:8080/", "height": 55}
test_list = tok_test.word_index
print(test_list)
# + [markdown] id="qZeXZbM5cPm5"
# ### 5. Encoding or Sequencing
# + id="88IUoE2tcavl" executionInfo={"status": "ok", "timestamp": 1603710148327, "user_tz": -330, "elapsed": 30604, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="7b8027ec-662c-4287-b807-b680922a846d" colab={"base_uri": "https://localhost:8080/", "height": 52}
encoded_clean_text_t_stem = tok_all.texts_to_sequences(clean_text_t_stem)
print(clean_text_t_stem[0])
print(encoded_clean_text_t_stem[0])
# + [markdown] id="2qg4xgewcjLG"
# ### 6. Pre-padding
# + id="arj7T2r1coOw" executionInfo={"status": "ok", "timestamp": 1603710148328, "user_tz": -330, "elapsed": 30598, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
from keras.preprocessing import sequence
max_length = 100
padded_clean_text_t_stem = sequence.pad_sequences(encoded_clean_text_t_stem, maxlen=max_length, padding='pre')
# + [markdown] id="QfhyZliqgYTb"
# # fastText Embedding
# + id="G4S7PI9cw2xy" executionInfo={"status": "ok", "timestamp": 1603710315786, "user_tz": -330, "elapsed": 198036, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="1dafacf8-ee47-4605-ca87-69a2c533622a" colab={"base_uri": "https://localhost:8080/", "height": 35}
# fastText Embedding link - https://fasttext.cc/docs/en/crawl-vectors.html
import os
import numpy as np
embeddings_index = {}
f = open('drive/My Drive/ML Internship IIIT Dharwad/Copy of cc.en.300.vec',encoding='utf-8',errors='ignore')
for line in f:
values = line.split()
word = values[0]
coefs = np.asarray(values[1:], dtype='float32')
embeddings_index[word] = coefs
f.close()
print('Loaded %s word vectors.' % len(embeddings_index))
# + id="7-9fLmPZzlP_" executionInfo={"status": "ok", "timestamp": 1603710315789, "user_tz": -330, "elapsed": 198034, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
embedding_matrix = np.zeros((vocabulary_all+1, 300))
for word, i in tok_all.word_index.items():
embedding_vector = embeddings_index.get(word)
if embedding_vector is not None:
embedding_matrix[i] = embedding_vector
# + [markdown] id="oM5OmlqZgrLy"
# # **BiLSTM Model**
# + id="r2VGeKXv0vOz" executionInfo={"status": "ok", "timestamp": 1603710315790, "user_tz": -330, "elapsed": 198030, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
from keras.preprocessing import sequence
from keras.preprocessing import text
import numpy as np
from keras.models import Sequential
from keras.layers import Dense, Dropout, Activation
from keras.layers import Embedding, LSTM , Bidirectional
from keras.layers import Conv1D, Flatten
from keras.preprocessing import text
from keras.models import Sequential,Model
from keras.layers import Dense ,Activation,MaxPool1D,Conv1D,Flatten,Dropout,Activation,Dropout,Input,Lambda,concatenate
from keras.utils import np_utils
from nltk.corpus import stopwords
from nltk.tokenize import RegexpTokenizer
from nltk.stem.porter import PorterStemmer
import nltk
import csv
import pandas as pd
from keras.preprocessing import text as keras_text, sequence as keras_seq
# + id="qr8uLf-q0lPJ" executionInfo={"status": "ok", "timestamp": 1603710323052, "user_tz": -330, "elapsed": 205289, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
Bi_lstm3_network = Sequential()
# Adding Embedding layer
Bi_lstm3_network.add(Embedding(vocabulary_all+1,300,weights = [embedding_matrix], input_length=max_length, trainable= False))
# Adding 3 Bi-Lstm layers
Bi_lstm3_network.add(Bidirectional(LSTM(128, return_sequences=True)))
Bi_lstm3_network.add(Dropout(0.2))
Bi_lstm3_network.add(Bidirectional(LSTM(64, return_sequences=False)))
Bi_lstm3_network.add(Dropout(0.2))
# Adding Dense layer
Bi_lstm3_network.add(Dense(64,activation="relu"))
Bi_lstm3_network.add(Dropout(0.3))
Bi_lstm3_network.add(Dense(2,activation="sigmoid"))
# + id="iqV6VLZ83HH6" executionInfo={"status": "ok", "timestamp": 1603710323056, "user_tz": -330, "elapsed": 205281, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="fbf3dcc4-0218-4d2b-de3c-ded512b6dc24" colab={"base_uri": "https://localhost:8080/", "height": 433}
Bi_lstm3_network.summary()
# + id="80QTgAc6BMJ1" executionInfo={"status": "ok", "timestamp": 1603710323057, "user_tz": -330, "elapsed": 205279, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
from keras.optimizers import Adam
Bi_lstm3_network.compile(loss = "binary_crossentropy", optimizer=Adam(lr=0.00003), metrics=["accuracy"])
# + id="h9c9ECvp7P9f" executionInfo={"status": "ok", "timestamp": 1603710323745, "user_tz": -330, "elapsed": 205956, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="4d7a44ec-b53d-4713-d704-7ec67f396b61" colab={"base_uri": "https://localhost:8080/", "height": 976}
from keras.utils.vis_utils import plot_model
plot_model(Bi_lstm3_network, to_file='Bi_lstm3_network.png', show_shapes=True, show_layer_names=True)
# + id="LR0JsV_kAcRY" executionInfo={"status": "ok", "timestamp": 1603710323747, "user_tz": -330, "elapsed": 205955, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
from keras.callbacks import EarlyStopping, ReduceLROnPlateau,ModelCheckpoint
earlystopper = EarlyStopping(patience=8, verbose=1)
reduce_lr = ReduceLROnPlateau(monitor='val_loss', factor=0.9,
patience=2, min_lr=0.00001, verbose=1)
# + [markdown] id="fMnqgj6rhDVR"
# ### **Model Fitting or Training**
# + id="5nbnfnRZAv1M" executionInfo={"status": "ok", "timestamp": 1603710487517, "user_tz": -330, "elapsed": 369715, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="bd06ff9f-4527-4f6f-d09a-de0f15ef8f14" colab={"base_uri": "https://localhost:8080/", "height": 1000}
hist = Bi_lstm3_network.fit(padded_clean_text_stem,label_twoDimension,validation_data=(padded_clean_text_t_stem,label_twoDimension_t),epochs=150,batch_size=32,callbacks=[earlystopper, reduce_lr])
# + [markdown] id="T5W_uxCThTLl"
# # log loss
# + id="X9DBoQg8Cf1G" executionInfo={"status": "ok", "timestamp": 1603710489571, "user_tz": -330, "elapsed": 371766, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
Bi_lstm3_network_predictions = Bi_lstm3_network.predict(padded_clean_text_t_stem)
# + id="bJQznoSlJ5bT" executionInfo={"status": "ok", "timestamp": 1603710489580, "user_tz": -330, "elapsed": 371762, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="3e65ed65-0800-47e0-dde1-5e147b58623e" colab={"base_uri": "https://localhost:8080/", "height": 35}
from sklearn.metrics import log_loss
log_loss_test= log_loss(label_twoDimension_t,Bi_lstm3_network_predictions)
log_loss_test
# + [markdown] id="MryQdO5YhdEz"
# # Classification Report
# + id="3UhoxZljKBVs" executionInfo={"status": "ok", "timestamp": 1603710489582, "user_tz": -330, "elapsed": 371759, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
predictions = np.zeros_like(Bi_lstm3_network_predictions)
predictions[np.arange(len(Bi_lstm3_network_predictions)), Bi_lstm3_network_predictions.argmax(1)] = 1
# + id="pNAHulQqKP80" executionInfo={"status": "ok", "timestamp": 1603710489587, "user_tz": -330, "elapsed": 371750, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="f1f41419-c394-427f-9aeb-34bc266be777" colab={"base_uri": "https://localhost:8080/", "height": 35}
predictionInteger=(np.argmax(predictions, axis=1))
predictionInteger
# + id="p4zH_CHRSkji" executionInfo={"status": "ok", "timestamp": 1603710489590, "user_tz": -330, "elapsed": 371741, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="15c53ecf-c60b-4cdc-97e4-f4cc83c4bd88" colab={"base_uri": "https://localhost:8080/", "height": 35}
'''pred_label = np.array(predictionInteger)
df = pd.DataFrame(data=pred_label , columns=["task1"])
print(df)'''
# + id="gMcD5cG7XLL9" executionInfo={"status": "ok", "timestamp": 1603710489592, "user_tz": -330, "elapsed": 371740, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
# df.to_csv("submission_EN_A.csv", index=False)
# + id="HE-j9PERKXBE" executionInfo={"status": "ok", "timestamp": 1603710489594, "user_tz": -330, "elapsed": 371730, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="027abe4a-0b30-49a9-c1cb-4e3633b12ab4" colab={"base_uri": "https://localhost:8080/", "height": 191}
from sklearn.metrics import classification_report
print(classification_report(label_twoDimension_t,predictions))
# + [markdown] id="WqNX-4ljhwsu"
# # Epoch v/s Loss Plot
# + id="Dk322X4pKjEQ" executionInfo={"status": "ok", "timestamp": 1603710489596, "user_tz": -330, "elapsed": 371720, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="0cfbeb12-c9d8-4644-c5dd-c6c5245c7b3a" colab={"base_uri": "https://localhost:8080/", "height": 295}
from matplotlib import pyplot as plt
plt.plot(hist.history["loss"],color = 'red', label = 'train_loss')
plt.plot(hist.history["val_loss"],color = 'blue', label = 'val_loss')
plt.title('Loss Visualisation')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.savefig('Bi_lstm3_HASOC_Eng_lossPlot.pdf',dpi=1000)
from google.colab import files
files.download('Bi_lstm3_HASOC_Eng_lossPlot.pdf')
# + [markdown] id="A5eYuEVbh0Qi"
# # Epoch v/s Accuracy Plot
# + id="BSDEzNM1LKmp" executionInfo={"status": "ok", "timestamp": 1603710490595, "user_tz": -330, "elapsed": 372707, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="e2ec74d8-2ba2-4047-9740-6bb4465b7721" colab={"base_uri": "https://localhost:8080/", "height": 295}
plt.plot(hist.history["accuracy"],color = 'red', label = 'train_accuracy')
plt.plot(hist.history["val_accuracy"],color = 'blue', label = 'val_accuracy')
plt.title('Accuracy Visualisation')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.legend()
plt.savefig('Bi_lstm3_HASOC_Eng_accuracyPlot.pdf',dpi=1000)
files.download('Bi_lstm3_HASOC_Eng_accuracyPlot.pdf')
# + [markdown] id="5v-PNBwUh6fK"
# # Area under Curve-ROC
# + id="rIga22ZbL5Lg" executionInfo={"status": "ok", "timestamp": 1603710493133, "user_tz": -330, "elapsed": 375241, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
pred_train = Bi_lstm3_network.predict(padded_clean_text_stem)
pred_test = Bi_lstm3_network.predict(padded_clean_text_t_stem)
# + id="rWKVJtN1Mz_d" executionInfo={"status": "ok", "timestamp": 1603710493134, "user_tz": -330, "elapsed": 375240, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
import numpy as np
import matplotlib.pyplot as plt
from itertools import cycle
from sklearn import svm, datasets
from sklearn.metrics import roc_curve, auc
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import label_binarize
from sklearn.multiclass import OneVsRestClassifier
from scipy import interp
def plot_AUC_ROC(y_true, y_pred):
n_classes = 2 #change this value according to class value
# Compute ROC curve and ROC area for each class
fpr = dict()
tpr = dict()
roc_auc = dict()
for i in range(n_classes):
fpr[i], tpr[i], _ = roc_curve(y_true[:, i], y_pred[:, i])
roc_auc[i] = auc(fpr[i], tpr[i])
# Compute micro-average ROC curve and ROC area
fpr["micro"], tpr["micro"], _ = roc_curve(y_true.ravel(), y_pred.ravel())
roc_auc["micro"] = auc(fpr["micro"], tpr["micro"])
############################################################################################
lw = 2
# Compute macro-average ROC curve and ROC area
# First aggregate all false positive rates
all_fpr = np.unique(np.concatenate([fpr[i] for i in range(n_classes)]))
# Then interpolate all ROC curves at this points
mean_tpr = np.zeros_like(all_fpr)
for i in range(n_classes):
mean_tpr += interp(all_fpr, fpr[i], tpr[i])
# Finally average it and compute AUC
mean_tpr /= n_classes
fpr["macro"] = all_fpr
tpr["macro"] = mean_tpr
roc_auc["macro"] = auc(fpr["macro"], tpr["macro"])
# Plot all ROC curves
plt.figure()
plt.plot(fpr["micro"], tpr["micro"],
label='micro-average ROC curve (area = {0:0.2f})'
''.format(roc_auc["micro"]),
color='deeppink', linestyle=':', linewidth=4)
plt.plot(fpr["macro"], tpr["macro"],
label='macro-average ROC curve (area = {0:0.2f})'
''.format(roc_auc["macro"]),
color='navy', linestyle=':', linewidth=4)
colors = cycle(['aqua', 'darkorange'])
#classes_list1 = ["DE","NE","DK"]
classes_list1 = ["Non-duplicate","Duplicate"]
for i, color,c in zip(range(n_classes), colors,classes_list1):
plt.plot(fpr[i], tpr[i], color=color, lw=lw,
label='{0} (AUC = {1:0.2f})'
''.format(c, roc_auc[i]))
plt.plot([0, 1], [0, 1], 'k--', lw=lw)
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic curve')
plt.legend(loc="lower right")
#plt.show()
plt.savefig('Bi_lstm3_HASOC_Eng_Area_RocPlot.pdf',dpi=1000)
files.download('Bi_lstm3_HASOC_Eng_Area_RocPlot.pdf')
# + id="i3tsqxDENNB6" executionInfo={"status": "ok", "timestamp": 1603710493135, "user_tz": -330, "elapsed": 375230, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}} outputId="df9ad210-c703-4baa-e86f-6578ffb6882c" colab={"base_uri": "https://localhost:8080/", "height": 333}
plot_AUC_ROC(label_twoDimension_t,pred_test)
# + id="6boPbARON83n" executionInfo={"status": "ok", "timestamp": 1603710493137, "user_tz": -330, "elapsed": 375228, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggzx2UtLRgfAA9F990pxIS41WDo9WctnzL8mXg3LQ=s64", "userId": "17758832831689054457"}}
| BiLSTM Models/With fastText/BiLSTM2 Covid-19.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# An LSF file is a binary file of concatenated IMC messages.
# This notebook demonstrates some of the included functionality for parsing and exporting LSF files.
#
# **Warning:** *Pyimc parses the LSF files under the assumption that the IMC specification is the same as the one it was compiled against.*
# +
import pyimc
from pyimc.lsf import LSFReader, LSFExporter
lsf_path ='./Data.lsf'
# -
# ## LSFReader usage
# The LSFReader class can be instantiated using the 'with' statement to automatically handle file opening/closing.
# Print first three messages in LSF file
# save_index: a message index is generated, speeds up subsequent reads
with LSFReader(lsf_path, save_index=True) as lsf:
for i, msg in zip(range(3), lsf.read_message()):
# Print outputs the message type, destination/source imc address, timestamp and fields
print('Message', i)
print(msg)
print()
# The convenience function "read" returns a generator of messages of a certain type instead of using the "with" command.
# Can be iterated over or converted to a list
msgs = list(LSFReader.read(lsf_path, types=[pyimc.EstimatedState]))
last_msg = msgs[-1]
print(last_msg)
# ## LSFExporter usage
# This class reads LSF files and exports it to a Pandas DataFrame.
# +
import pandas as pd
from IPython.display import display
pd.set_option('display.max_columns', 30)
exp = LSFExporter(lsf_path)
df = exp.export_messages(imc_type=pyimc.EstimatedState)
# Print first five rows
display(df.head())
# +
import numpy as np
# Index by timestamp instead
df_t = df.set_index('timestamp')
# Get all rows between two timestamps
t_start = np.datetime64('2020-02-11 08:36:00')
t_end = np.datetime64('2020-02-11 08:36:10')
df_slice = df_t[t_start:t_end]
display(df_slice)
# -
# Output data to a csv file
df_slice.to_csv('output.csv')
# +
# Read another message type using the same exporter (re-use metadata)
df2 = exp.export_messages(imc_type=pyimc.Announce)
# Remove all messages not originating from lauv-simulator-1
df2 = df2[df2['src'] == 'lauv-simulator-1']
display(df2.head(2))
# Alternatively, supply a lambda predicate to the export_messages function (avoids converting unnecessary messages to pandas)
imc_id = exp.get_node_id('lauv-simulator-1')
df3 = exp.export_messages(imc_type=pyimc.Announce, condition=lambda msg: msg.src == imc_id)
display(df3.head(2))
| examples/lsf_parsing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Let's see if you need to unzip the data again.
# !ls ../data
# +
# Unzip the data if needed
# Replace PASSWORD with the password to unzip
# !unzip -P PASSWORD ../data.zip -d ../
# +
import sys
sys.path.append("..")
from data.preparer import load_youtube_dataset
from babble import Explanation
from babble import BabbleStream
from babble.Candidate import Candidate
from analyzer import upload_data
from metal.analysis import lf_summary
from metal.analysis import label_coverage
from metal import LabelModel
from metal.tuners import RandomSearchTuner
from babble.utils import ExplanationIO
from snorkel.labeling import filter_unlabeled_dataframe
import nltk
nltk.download("punkt")
import pandas as pd
from datetime import datetime
stat_history = pd.DataFrame()
# -
DELIMITER = "#"
df_train, df_dev, df_valid, df_test, _ = load_youtube_dataset(delimiter=DELIMITER)
print("{} training examples".format(len(df_train)))
print("{} development examples".format(len(df_dev)))
print("{} validation examples".format(len(df_valid)))
print("{} test examples".format(len(df_test)))
# Transform the data into a format compatible with Babble Labble:
# +
dfs = [df_train, df_dev]
dfs[0]['label'] = -1
for df in dfs:
df["id"] = range(len(df))
df["label"] += 1
Cs = [df.apply(lambda x: Candidate(x), axis=1) for df in dfs]
# babble labble uses 1 and 2 for labels, while our data uses 0 and 1
# add 1 to convert
Ys = [df.label.values for df in dfs]
Ys[0] -= 1 # no label (training set) should be set to -1
# -
# # Youtube Spam Classification Task with Babble
#
# ### For this task, you will work with comments from 5 different YouTube videos, and classify comments as either spam or legitimate comments by writing labeling explanations with Babble Labble.
#
# Spam can be defined as irrelevant or unsolicited messages sent over the Internet.
#
# Start the timer!
stat_history = stat_history.append({
"time": datetime.now(),
"num_lfs": 0,
"f1": 0.0,
"precision": 0.0,
"recall": 0.0,
"training_label_coverage": 0.0,
"training_label_size": 0.0
}, ignore_index=True)
#define labels
ABSTAIN = 0
NOT_SPAM = 1
SPAM = 2
babbler = BabbleStream(Cs, Ys, balanced=True, shuffled=True, seed=456)
# Recall that aliases are a way to refer to a set of words in a rule.
#
# For example, with
# `aliases = {"couples": ["girlfriend", "boyfriend", "wife", "husband"]}`
#
# --> now you can refer to "couples" in a rule, and the parser will know you mean any of these terms.
#
#
aliases = {}
babbler.add_aliases(aliases)
def prettyprint(candidate):
# just a helper function to print the candidate nicely
print("MENTION ID {}".format(candidate.mention_id))
print()
print(candidate.text)
# Let's see an example candidate!
candidate = babbler.next()
prettyprint(candidate)
#
# Your task is to __create labeling functions__ by writing natural language descriptions of labeling rules. Try to write them as quickly and accurately as possible.
#
# You may consult the internet at any time.
# ## Create Explanations
#
# Creating explanations generally happens in five steps:
# 1. View candidates
# 2. Write explanations
# 3. Get feedback
# 4. Update explanations
# 5. Apply label aggregator
#
# Steps 3-5 are optional; explanations may be submitted without any feedback on their quality. However, in our experience, observing how well explanations are being parsed and what their accuracy/coverage on a dev set are (if available) can quickly lead to simple improvements that yield significantly more useful labeling functions. Once a few labeling functions have been collected, you can use the label aggregator to identify candidates that are being mislabeled and write additional explanations targeting those failure modes.
# ### Collection
#
# Use `babbler` to show candidates
candidate = babbler.next()
print(candidate)
# If you don't know whether it's spam or not, it's okay to make your best guess or skip an example.
# For a candidate you decide to label, write an explanation of why you chose that label.
#
# You can consult the internet or refer to the babble tutorial notebook.
e0 = Explanation(
# feel free to change the name to something that describes your rule better.
name = "e0",
label = ABSTAIN,
condition = "",
# remember that is argument (candidate) is optional.
# You can use it to make sure the explanation applies to the candidate you pass as an argument.
candidate = candidate.mention_id
)
e1 = Explanation(
name = "e1",
label = ABSTAIN,
condition = "",
candidate = candidate.mention_id
)
e2 = Explanation(
name = "e2",
label = ABSTAIN,
condition = "",
candidate = candidate.mention_id
)
e3 = Explanation(
name = "e3",
label = ABSTAIN,
condition = "",
candidate = candidate.mention_id
)
e4 = Explanation(
name = "e4",
label = ABSTAIN,
condition = "",
candidate = candidate.mention_id
)
e5 = Explanation(
name = "e5",
label = ABSTAIN,
condition = "",
candidate = candidate.mention_id
)
e6 = Explanation(
name = "e6",
label = ABSTAIN,
condition = "",
candidate = candidate.mention_id
)
e7 = Explanation(
name = "e7",
label = ABSTAIN,
condition = "",
candidate = candidate.mention_id
)
e8 = Explanation(
name = "e8",
label = ABSTAIN,
condition = "",
candidate = candidate.mention_id
)
e9 = Explanation(
name = "e9",
label = ABSTAIN,
condition = "",
candidate = candidate.mention_id
)
# Babble will parse your explanations into functions, then filter out functions that are duplicates, incorrectly label their given candidate, or assign the same label to all examples.
# +
# Add any explanations that you haven't committed yet
explanations = [e0, e1, e2, e3, e4, e5, e6, e7, e8, e9]
parses, filtered = babbler.apply(explanations)
stat_history = stat_history.append({
"time": datetime.now(),
"num_lfs": len(parses),
"num_explanations": len(explanations),
"num_filtered": len(filtered)
}, ignore_index=True)
# -
# ### Analysis
# See how your explanations were parsed and filtered
try:
dev_analysis = babbler.analyze(parses)
display(dev_analysis)
dev_analysis['time'] = datetime.now()
dev_analysis['eval'] = "dev"
dev_analysis["lf_id"] = dev_analysis.index
stat_history = stat_history.append(dev_analysis, sort=False, ignore_index=True)
except ValueError as e:
print("It seems as though none of your labeling functions were parsed. See the cells above and below for more information.")
print("ERROR:")
print(e)
babbler.filtered_analysis(filtered)
babbler.commit()
# ### Evaluation
# Get feedback on the performance of your explanations
# +
from metal.analysis import lf_summary
Ls = [babbler.get_label_matrix(split) for split in [0,1,2]]
lf_names = [lf.__name__ for lf in babbler.get_lfs()]
lf_summary(Ls[1], Ys[1], lf_names=lf_names)
# +
search_space = {
'n_epochs': [50, 100, 500],
'lr': {'range': [0.01, 0.001], 'scale': 'log'},
'show_plots': False,
}
tuner = RandomSearchTuner(LabelModel, seed=123)
label_aggregator = tuner.search(
search_space,
train_args=[Ls[0]],
X_dev=Ls[1], Y_dev=Ys[1],
max_search=20, verbose=False, metric='f1')
# record statistics over time
pr, re, f1, acc = label_aggregator.score(Ls[1], Ys[1], metric=['precision', 'recall', 'f1', 'accuracy'])
stats = {
"precision": pr,
"recall": re,
"f1": f1,
"accuracy": acc,
"eval": "dev",
"model": "label_aggregator",
"time": datetime.now(),
"training_label_coverage": label_coverage(Ls[0]),
"training_label_size": label_coverage(Ls[0])*len(dfs[0])
}
stat_history = stat_history.append(stats, ignore_index=True)
# -
# view some incorrectly labeled examples for a given LF
j = 0
print(lf_names[j])
# set j to match the value of the LF you're interested in
L_dev = Ls[1].todense()
display(df_dev[L_dev[:,j].A1==abs(df_dev["label"]-3)])
# ## Train Model
# We can train a simple bag of words model on these labels, and see test accuracy.
#
# (This step may take a while).
L_train = Ls[0].todense()
probs_train = label_aggregator.predict_proba(L=L_train)
mask = (L_train != 0).any(axis=1).A1
df_train_filtered = df_train.iloc[mask]
probs_train_filtered = probs_train[mask]
print("{} out of {} examples used for training data".format(len(df_train_filtered), len(df_train)))
from analyzer import train_model_from_probs
stats = train_model_from_probs(df_train_filtered, probs_train_filtered, df_valid, df_test)
stats["time"] = datetime.now()
stat_history = stat_history.append(stats, ignore_index=True)
# ## Save
# When your time is up, please save your explanations and model!
# Enter your name (for file naming)
YOUR_NAME = ""
# +
# save statistics history
stat_history.to_csv("babble_youtube_statistics_history.csv")
# save explanations
FILE = "babble_youtube_explanations.tsv"
from types import SimpleNamespace
exp_io = ExplanationIO()
for exp in explanations:
if exp.candidate is None:
exp.candidate = SimpleNamespace(mention_id = None)
exp_io.write(explanations, FILE)
explanations = exp_io.read(FILE)
# save label model
label_aggregator.save("babble_youtube_lfmodel.pkl")
# -
# !mkdir babble_youtube
stat_history.to_csv("babble_youtube/statistics_history.csv")
# %history -p -o -f babble_youtube/history.log
# !cp babble_youtube_task.ipynb babble_youtube/notebook.ipynb
# +
# save explanations
FILE = "babble_youtube/explanations.tsv"
from types import SimpleNamespace
exp_io = ExplanationIO()
for exp in explanations:
if exp.candidate is None:
exp.candidate = SimpleNamespace(mention_id = None)
exp_io.write(explanations, FILE)
explanations = exp_io.read(FILE)
# save label model
label_aggregator.save("babble_youtube/lfmodel.pkl")
# +
# zip and upload the data
import shutil
shutil.make_archive(YOUR_NAME + "_babble_youtube", 'zip', "babble_youtube")
assert len(YOUR_NAME) > 0
upload_data(YOUR_NAME + "_babble_youtube.zip")
# -
# ...And you're done with this spam classification task!
#
# ## THANK YOU :]
| notebooks/babble_youtube_task.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Example of a 1D Channel driven flow
#
# Model Setup
# -----------
#
# 2D, Stokes Equation with noslip BC at top and bottom boundary and a lateral pressure gradient driving the flow, a.k.a. Poiseuille Flow.
#
# \\[
# \frac{\partial \tau}{\partial y} = \mu \frac{\partial^{2} \mathbf{u}}{\partial{y}^{2}} = \frac{\partial p}{\partial x}
# \\]
#
# \\[
# \nabla \cdot \mathbf{u} = 0
# \\]
#
# with $ x_{a} \leqslant x \leqslant x_{b} $ and $ 0.0 \leqslant y \leqslant h $
#
# Boundary conditions:
#
# * $\mathbf{u}(x,y=h) = \mathbf{u}(x,y=0) = \left[0,0 \right]$
# * $P(x_a) = P_a$
# * $P(x_b) = P_b $
#
# ------
#
# A 1D solution in $y$-axis, described by
#
# $ \mathbf{u}(x,y) = \left[ \frac{1}{2 \mu} \frac{\partial p }{\partial x} ( y^{2} - h y ), 0.0 \right]$
# We implement the above boundary conditions using:
# * a `DirichletCondition` object for $\mathbf{u}(x,y=1) = \mathbf{u}(x,y=0) = \left[0,0 \right]$
# * a `NeumannCondition` object for $P(x_a) = P_a$ & $P(x_b) = P_b $
#
# The `NeumannCondition` object, used with the `Stokes` object, defines a stress along a boundary such that:
# * $ \sigma_{ij} n_{j} = \phi_{i} $ on $ \Gamma_{\phi} $
#
# where
# * $n$ is the surface normal pointing outwards,
# * $ \sigma_{ij} = \tau_{ij} - \delta_{ij} P$ is the prescribed stress tensor, which is multiplied my $ n $ at $ \Gamma_{\phi} $ to produce $\phi_{i}$, a surface traction on the given boundary.
# +
import underworld as uw
from underworld import function as fn
import underworld.visualisation as vis
import math
import numpy as np
# for machines without matplotlib #
make_graphs = False
if uw.utils.is_kernel():
make_graphs = True
try:
import matplotlib
except ImportError:
make_graphs=False
# -
# Setup parameters
# -----
# +
# domain height
h = 1.0
# Set a constant viscosity.
viscosity = 1.4
# position of walls and associated pressure on walls
xa = -1.0
pa = 4.0
xb = 1.0
pb = 3.0
# +
mesh = uw.mesh.FeMesh_Cartesian( elementType = ("Q1/dQ0"),
elementRes = (128, 128),
minCoord = (xa, 0.),
maxCoord = (xb, h))
velocityField = uw.mesh.MeshVariable( mesh=mesh, nodeDofCount=2 )
pressureField = uw.mesh.MeshVariable( mesh=mesh.subMesh, nodeDofCount=1 )
appliedTractionField = uw.mesh.MeshVariable( mesh=mesh, nodeDofCount=2 )
# initialise velocity, pressure field
velocityField.data[:] = [0.,0.]
pressureField.data[:] = 0.
# -
jWalls = mesh.specialSets["MinJ_VertexSet"] + mesh.specialSets["MaxJ_VertexSet"]
iWalls = mesh.specialSets["MinI_VertexSet"] + mesh.specialSets["MaxI_VertexSet"]
allWalls = iWalls + jWalls
# +
vBC = uw.conditions.DirichletCondition( variable = velocityField,
indexSetsPerDof = (jWalls, allWalls) )
dp_dx = (pb-pa)/(xb-xa)
# This stress is multiplied by the wall normal to produce a traction force.
#############
# Remember total stress = deviatoric - isotropic.
# Thus +pressure is a negative stress.
#############
# The left wall normal unit vector is (-1,0)
# The right wall normal unit vector is (1,0)
# (-press) * normal_j = surface_force
appliedTractionField.data[mesh.specialSets["MinI_VertexSet"].data] = (pa,0.0)
appliedTractionField.data[mesh.specialSets["MaxI_VertexSet"].data] = (-1*pb,0.0)
nbc = uw.conditions.NeumannCondition( fn_flux=appliedTractionField,
variable=velocityField,
indexSetsPerDof=(iWalls, None) )
print(dp_dx)
# +
stokes = uw.systems.Stokes( velocityField = velocityField,
pressureField = pressureField,
conditions = [vBC, nbc],
fn_viscosity = viscosity,
fn_bodyforce = 0.0 )
solver = uw.systems.Solver( stokes )
solver.solve()
# -
fig = vis.Figure()
velmagfield = uw.function.math.sqrt( uw.function.math.dot( velocityField, velocityField ) )
fig.append( vis.objects.VectorArrows(mesh, velocityField, arrowHead=0.2, scaling=0.9) )
# fig.append( vis.objects.Mesh(mesh) )
fig.append( vis.objects.Surface( mesh, pressureField ) )
fig.show()
# +
ids = mesh.specialSets["MaxI_VertexSet"]
coords = mesh.data[ids.data] # xcoords
print("Ids, ", ids.size)
V = velocityField.evaluate(ids)
gradV = velocityField.fn_gradient.evaluate(ids)
from mpi4py import MPI
comm = MPI.COMM_WORLD
# assuming order in the allgather is the same
# some squashing of arrays required for parallel comms.
V = np.concatenate(
comm.allgather( V.ravel() )
).reshape(-1,mesh.dim)
gradV = np.concatenate(
comm.allgather( gradV.ravel() )
).reshape(-1,mesh.dim*velocityField.data.shape[1])
u = V[:,0] ; v = V[:,1]
du_dx = gradV[:,0] ; du_dy = gradV[:,1]
dv_dx = gradV[:,2] ; dv_dy = gradV[:,3]
# -
def exact_vx(y):
ana_u = 1.0 / (2.0 * viscosity)* dp_dx * (y**2 - h*y)
return ana_u
if make_graphs:
uw.utils.matplotlib_inline()
import matplotlib.pyplot as pyplot
import matplotlib.pylab as pylab
pyplot.ion() # needed to ensure pure python jobs do now hang on show()
pylab.rcParams[ 'figure.figsize'] = 12, 6
pyplot.plot(coords[:,1], u, 'o', color = 'green', label='vx')
pyplot.plot(coords[:,1], v, 'o', color = 'red', label='vy')
big = np.linspace(0.0,h)
pyplot.plot(big, exact_vx(big), 'D', color = 'purple', label='exact_vx', markersize=2)
pyplot.legend()
pyplot.xlabel('y coords at x=xmax')
pyplot.ylabel('velocity')
pyplot.show()
# +
ana_u = exact_vx(coords[:,1])
if uw.mpi.rank == 0:
abserr = np.linalg.norm(ana_u - u)
mag = np.linalg.norm(ana_u)
relerr = abserr / mag
threshold = 1.0e-4
print("Relative error against analytic solution: {}".format(relerr))
if relerr > threshold:
raise RuntimeError("The numerical solution is outside the error threshold of the analytic solution." \
"The Relative error was ", relerr," the threshold is ", threshold)
# +
# Further analytics
# pyplot.plot(coords[:,1], du_dy, 'o', color = 'purple', label='du_dy')
# pyplot.plot(coords[:,1], du_dx, '+', color = 'black', label='du_dx')
# pyplot.plot(coords[:,1], dv_dy, 'x', color = 'orange', label='dv_dy')
# pyplot.plot(coords[:,1], dv_dx, '.', color = 'red', label='dv_dx')
# pyplot.legend()
# pyplot.xlabel('y coords at x=xmax')
# pyplot.ylabel('velocity gradients')
# pyplot.show()
# +
## Further analytics
# def exact_shearSR(y):
# shearSR = dp_dx / (2.0*viscosity) * (y - h/2 )
# return shearSR
# strainRate = fn.tensor.symmetric( velocityField.fn_gradient )
# devstress = 2.0 * viscosity * strainRate
# pyplot.plot(coords[:,1], sr, '-', label='exact_shearSR')
# pyplot.plot(coords[:,1], strainRate.evaluate(ids)[:,2], 'o', color = 'purple', label='sr_shear')
# pyplot.plot(coords[:,1], devstress.evaluate(ids)[:,2], '+', label='tau_shear')
# pyplot.legend()
# pyplot.xlabel('y coords at x=xmax')
# pyplot.ylabel('strain rate')
# pyplot.show()
| docs/test/StokesEq_PoiseuilleFlow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/jazzathoth/DS-Unit-1-Sprint-4-Statistical-Tests-and-Experiments/blob/master/module2-sampling-confidence-intervals-and-hypothesis-testing/LS_DS_142_Sampling_Confidence_Intervals_and_Hypothesis_Testing.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="838Dmw1kM2LK" colab_type="text"
# # Lambda School Data Science Module 142
# ## Sampling, Confidence Intervals, and Hypothesis Testing
# + [markdown] id="dbcPKIo5M6Ny" colab_type="text"
# ## Prepare - examine other available hypothesis tests
#
# If you had to pick a single hypothesis test in your toolbox, t-test would probably be the best choice - but the good news is you don't have to pick just one! Here's some of the others to be aware of:
# + id="uu-41Y1XDDM1" colab_type="code" colab={}
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
from scipy.stats import chisquare # One-way chi square test
# + id="tlBel8j9M6tB" colab_type="code" outputId="b61bc5de-d1f7-4bb2-edf7-5ae5bbd2eaea" colab={"base_uri": "https://localhost:8080/", "height": 187}
# Chi square can take any crosstab/table and test the independence of rows/cols
# The null hypothesis is that the rows/cols are independent -> low chi square
# The alternative is that there is a dependence -> high chi square
# Be aware! Chi square does *not* tell you direction/causation
ind_obs = np.array([[1, 1], [2, 2]]).T
print(ind_obs)
print(chisquare(ind_obs, axis=None))
dep_obs = np.array([[16, 18, 16, 14, 12, 12], [32, 24, 16, 28, 20, 24]]).T
print(dep_obs)
print(chisquare(dep_obs, axis=None))
# + id="nN0BdNiDPxbk" colab_type="code" outputId="c53a4568-47c6-4c10-fb9b-8be46b40e6ee" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Distribution tests:
# We often assume that something is normal, but it can be important to *check*
# For example, later on with predictive modeling, a typical assumption is that
# residuals (prediction errors) are normal - checking is a good diagnostic
from scipy.stats import normaltest
# Poisson models arrival times and is related to the binomial (coinflip)
sample = np.random.poisson(5, 1000)
print(normaltest(sample)) # Pretty clearly not normal
# + id="P5t0WhkDReFO" colab_type="code" outputId="218c2369-b627-4b11-cb84-68b56c535308" colab={"base_uri": "https://localhost:8080/", "height": 51}
# Kruskal-Wallis H-test - compare the median rank between 2+ groups
# Can be applied to ranking decisions/outcomes/recommendations
# The underlying math comes from chi-square distribution, and is best for n>5
from scipy.stats import kruskal
x1 = [1, 3, 5, 7, 9]
y1 = [2, 4, 6, 8, 10]
print(kruskal(x1, y1)) # x1 is a little better, but not "significantly" so
x2 = [1, 1, 1]
y2 = [2, 2, 2]
z = [2, 2] # Hey, a third group, and of different size!
print(kruskal(x2, y2, z)) # x clearly dominates
# + [markdown] id="7pT3IP36Rh0b" colab_type="text"
# And there's many more! `scipy.stats` is fairly comprehensive, though there are even more available if you delve into the extended world of statistics packages. As tests get increasingly obscure and specialized, the importance of knowing them by heart becomes small - but being able to look them up and figure them out when they *are* relevant is still important.
# + [markdown] id="L1_KRuHCM7BW" colab_type="text"
# ## Live Lecture - let's explore some more of scipy.stats
#
# Candidate topics to explore:
#
# - `scipy.stats.chi2` - the Chi-squared distribution, which we can use to reproduce the Chi-squared test
# - Calculate the Chi-Squared test statistic "by hand" (with code), and feed it into `chi2`
# - Build a confidence interval with `stats.t.ppf`, the t-distribution percentile point function (the inverse of the CDF) - we can write a function to return a tuple of `(mean, lower bound, upper bound)` that you can then use for the assignment (visualizing confidence intervals)
# + id="qW6k0dorM7Lz" colab_type="code" colab={}
# Taking requests! Come to lecture with a topic or problem and we'll try it.
# + id="n8M8yuQwDozV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="24928e6b-29e8-43dd-aec8-ccf2636510a6"
gender = ['male', 'male','male','female','female','female']
eats_outside = ['outside', 'inside', 'inside', 'inside', 'outside', 'outside']
df = pd.DataFrame({'gender': gender, 'preference': eats_outside})
df.head()
# + id="qslIT0QZDowW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 142} outputId="7ea6dd35-1bf6-4199-aa01-6f72516511d9"
table = pd.crosstab(df.gender, df.preference)
table
# + id="08O31232Dose" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 173} outputId="935f3ad4-e0d3-4f1c-cf7f-a17a6b1e141e"
pd.crosstab(df.gender, df.preference, margins=True)
# + id="f2NO3_dADog3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="bd9cad5d-be7f-40fb-b220-f70de47df6c5"
df = df.replace('male', 0)
df = df.replace('female', 1)
df = df.replace('outside', 0)
df = df.replace('inside', 1)
df.head()
# + id="03UwEbfQDob7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 170} outputId="cdd9f975-4a34-4cd0-b6a1-74eae28bd059"
pd.crosstab(df.gender, df.preference, margins=True)
expected = [[1.5, 1.5],
[1.5, 1.5]]
pd.crosstab(df.gender, df.preference, margins=True, normalize='all')
observed = np.array([[.5, .5],[.5, .5]])
deviation = numerator = observed - expected
print(numerator)
deviation_squared = deviation**2
print('deviation squared: ', deviation_squared, '\n')
fraction = (deviation_squared / expected)
print('\nfraction: ', fraction, '\n')
# + id="RJG1Qit9KTFf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9a5c6c85-603a-4e80-deed-90e9ad990d53"
expected_values = [[1.5, 1.5], [1.5, 1.5]]
deviation = (((.5)**2) / 1.5) * 4 # 0.5^2 deviation per cell, scaled and added
print(deviation)
# + id="KDpWvXFHDoY9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="7c4cbe00-42f1-477e-d15e-74d2430a19f5"
chi_data = [[1,2],
[2,1]]
from scipy.stats import chisquare # one way chi square
chisquare(chi_data, axis=None)
# + id="v3mFGsu2DoVq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 187} outputId="19905a59-567a-43e2-e345-a56c6ff16093"
from scipy.stats import chi2_contingency
chi2stat, pvalue, dof, observed = chi2_contingency(table)
print('chi2 stat: ', chi2stat, '\n')
print('p-value: ', pvalue, '\n')
print('degrees of freedom: ', dof, '\n')
print('Contingency Table: \n', observed, '\n')
# + id="W4ivaWoeDoSn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="faf0890c-3d68-4b9d-ccbe-2c311480dbba"
def lazy_chisquare(observed, expected):
chisquare = 0
for row_obs, row_exp in zip(observed, expected):
for obs, exp in zip(row_obs, row_exp):
chisquare += (obs - exp)**2 / exp
return chisquare
chi_data = [[1, 2], [2, 1]]
expected_values = [[1.5, 1.5], [1.5, 1.5]]
chistat = lazy_chisquare(chi_data, expected_values)
chistat
# + [markdown] id="uJUMRGoYMAWn" colab_type="text"
# # Confidence Intervals
# + id="7Ch2OQEnDn7S" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="9bd7fcdd-a9a7-4156-c201-7ebeffbd3574"
#confidence_interval = [lower_bound, upper_bound]
coinflips = np.random.binomial(n=1, p=.7, size=100)
print(coinflips)
# + id="UI0SsTJCMD57" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="3c4bc48e-ef6f-4172-8048-1a84735e3179"
import scipy.stats as stats
stats.ttest_1samp(coinflips, 0.5)
# + id="sjSj3zTtMD26" colab_type="code" colab={}
def confidence_interval(data, confidence=.95):
data = np.array(data)
mean = np.mean(data)
n = len(data)
stderr = stats.sem(data)
interval = stderr * stats.t.ppf((1 + confidence) / 2., n-1)
return(mean, mean - interval, mean + interval)
def report_confidence_interval(confidence_interval):
s = "our mean lies in the interval [{:.2}, {:.2}]".format(
confidence_interval[1], confidence_interval[2])
return s
# + id="D0xET8tTMD0H" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="22d34062-7da6-481b-de15-0e72336e9b65"
coinflip_interval = confidence_interval(coinflips)
coinflip_interval
# + id="bV8nTjNnMDxB" colab_type="code" colab={}
# + id="RDeqLEmZMDt7" colab_type="code" colab={}
# + id="qsnSCdvoMDqa" colab_type="code" colab={}
# + id="mlSQEWzoMDn0" colab_type="code" colab={}
# + [markdown] id="11OzdxWTM7UR" colab_type="text"
# ## Assignment - Build a confidence interval
#
# A confidence interval refers to a neighborhood around some point estimate, the size of which is determined by the desired p-value. For instance, we might say that 52% of Americans prefer tacos to burritos, with a 95% confidence interval of +/- 5%.
#
# 52% (0.52) is the point estimate, and +/- 5% (the interval $[0.47, 0.57]$) is the confidence interval. "95% confidence" means a p-value $\leq 1 - 0.95 = 0.05$.
#
# In this case, the confidence interval includes $0.5$ - which is the natural null hypothesis (that half of Americans prefer tacos and half burritos, thus there is no clear favorite). So in this case, we could use the confidence interval to report that we've failed to reject the null hypothesis.
#
# But providing the full analysis with a confidence interval, including a graphical representation of it, can be a helpful and powerful way to tell your story. Done well, it is also more intuitive to a layperson than simply saying "fail to reject the null hypothesis" - it shows that in fact the data does *not* give a single clear result (the point estimate) but a whole range of possibilities.
#
# How is a confidence interval built, and how should it be interpreted? It does *not* mean that 95% of the data lies in that interval - instead, the frequentist interpretation is "if we were to repeat this experiment 100 times, we would expect the average result to lie in this interval ~95 times."
#
# For a 95% confidence interval and a normal(-ish) distribution, you can simply remember that +/-2 standard deviations contains 95% of the probability mass, and so the 95% confidence interval based on a given sample is centered at the mean (point estimate) and has a range of +/- 2 (or technically 1.96) standard deviations.
#
# Different distributions/assumptions (90% confidence, 99% confidence) will require different math, but the overall process and interpretation (with a frequentist approach) will be the same.
#
# Your assignment - using the data from the prior module ([congressional voting records](https://archive.ics.uci.edu/ml/datasets/Congressional+Voting+Records)):
#
# 1. Generate and numerically represent a confidence interval
# 2. Graphically (with a plot) represent the confidence interval
# 3. Interpret the confidence interval - what does it tell you about the data and its distribution?
#
# Stretch goals:
#
# 1. Write a summary of your findings, mixing prose and math/code/results. *Note* - yes, this is by definition a political topic. It is challenging but important to keep your writing voice *neutral* and stick to the facts of the data. Data science often involves considering controversial issues, so it's important to be sensitive about them (especially if you want to publish).
# 2. Apply the techniques you learned today to your project data or other data of your choice, and write/discuss your findings here.
# 3. Refactor your code so it is elegant, readable, and can be easily run for all issues.
# + [markdown] id="nyJ3ySr7R2k9" colab_type="text"
# ## Resources
#
# - [Interactive visualize the Chi-Squared test](https://homepage.divms.uiowa.edu/~mbognar/applets/chisq.html)
# - [Calculation of Chi-Squared test statistic](https://en.wikipedia.org/wiki/Pearson%27s_chi-squared_test)
# - [Visualization of a confidence interval generated by R code](https://commons.wikimedia.org/wiki/File:Confidence-interval.svg)
# - [Expected value of a squared standard normal](https://math.stackexchange.com/questions/264061/expected-value-calculation-for-squared-normal-distribution) (it's 1 - which is why the expected value of a Chi-Squared with $n$ degrees of freedom is $n$, as it's the sum of $n$ squared standard normals)
# + id="jNDaew_MD_WD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="77964527-f608-4506-c402-57c82f478ba0"
# TODO - your code!
votes_url = 'https://archive.ics.uci.edu/ml/machine-learning-databases/voting-records/house-votes-84.data'
votes = pd.read_csv(votes_url, header=None, names=['party',
'infant_bill',
'water_project',
'budget',
'physician_bill',
'aid_elsalvador',
'church_school',
'anti_satellite',
'contra_aid',
'mx_missile',
'immigration',
'synfuels',
'edu_spending',
'superfund_litigation',
'crime',
'duty_free',
's_africa_export'])
votes.shape
# + id="CNHkNNcmEA25" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 193} outputId="6a8cc038-c78d-4bb7-9b97-c0e1af802c2d"
votes.describe()
# + id="hp74QRilEAoh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="99f77fed-3856-407d-d241-66fe6825c3e6"
votes['party'].value_counts()
# + id="MFeQpF9PEJdl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 323} outputId="29f7dd63-63b0-403f-fc7b-66c736ebade2"
votes_nan = pd.DataFrame()
for i in votes:
votes_nan[i] = votes[i].replace('?', np.nan)
print(votes_nan.isnull().sum())
# + id="7908JXkbEO7V" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="1ad1d4d3-0ce1-41fe-b1c5-702b7d9f7753"
democrats = votes.loc[votes['party'] == 'democrat']
republicans = votes.loc[votes['party'] == 'republican']
democrats.shape, republicans.shape
# + id="GnPOMe2CEY1F" colab_type="code" colab={}
counts_party = pd.DataFrame({'party vote':['democrats yes', 'democrats no', 'republicans yes', 'republicans no']})
dyes, dno, ryes, rno = int(), int(), int(), int()
for bill in votes_nan.drop(['party'], axis=1):
for ind in votes_nan[bill].index:
vote = votes_nan[bill].loc[ind]
party = votes_nan['party'].loc[ind]
if party == 'democrat':
if vote == 'y':
dyes += 1
elif vote == 'n':
dno += 1
elif party == 'republican':
if vote == 'y':
ryes += 1
elif vote == 'n':
rno += 1
counts_party[bill] = [dyes, dno, ryes, rno]
counts_party.set_index('party vote')
#democrats = votes.loc[votes['party'] == 'democrat']
#republicans = votes.loc[votes['party'] == 'republican']
# + id="nmqdNc7vNUzF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 278} outputId="85d25150-fd40-45c1-f7aa-ece1618f9434"
counts_party
# + id="T2djhvh2NLy0" colab_type="code" colab={}
counts_party.hist(figsize=(12,12));
# + id="KD2BSsh8O-Q1" colab_type="code" colab={}
dem_enc = democrats.drop(['party'], axis=1).replace('?', np.nan).replace({'y':1,'n':0})
rep_enc = republicans.drop(['party'], axis=1).replace('?', np.nan).replace({'y':1,'n':0})
vote_enc = votes.drop(['party'], axis=1).replace('?', np.nan).replace({'y':1,'n':0})
# + id="TioKEQSpPX3l" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 306} outputId="d2ea40bb-72e1-438f-c301-19372cebb0fe"
dem_enc.isnull().sum()
# + id="AhQacqPqNhLQ" colab_type="code" colab={}
dem_total, rep_total, dem_rep = pd.DataFrame(), pd.DataFrame(), pd.DataFrame()
for bill in dem_enc:
res, prob = stats.ttest_ind(dem_enc[bill], vote_enc[bill], nan_policy='omit')
dem_total[bill] = [res,prob]
for bill in rep_enc:
res, prob = stats.ttest_ind(vote_enc[bill], rep_enc[bill], nan_policy='omit')
rep_total[bill] = [res,prob]
for bill in rep_enc:
res, prob = stats.ttest_ind(dem_enc[bill], rep_enc[bill], nan_policy='omit', equal_var=False)
dem_rep[bill] = [res,prob]
# + id="To1P3RWWWIzI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 131} outputId="44236da3-8d10-4b71-b5fe-5bd1750acedc"
dem_total
# + id="JddpVc5WXyJX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 131} outputId="8f9776dc-1ee8-4c20-d9fb-6e01d6bdd4ac"
rep_total
# + id="PCfbHUSdX0b7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 131} outputId="0883ca17-4dab-4148-d72c-281cd451bd3f"
dem_rep
# + id="D6W_CGhSX_gn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 231} outputId="d63766ee-e2c0-48bd-9f2d-e76ae2f161c0"
dem_t1, rep_t1 = pd.DataFrame(), pd.DataFrame()
for bill in dem_enc:
res, prob = stats.ttest_1samp(dem_enc[bill], nan_policy='omit')
dem_t1[bill] = [res,prob]
for bill in rep_enc:
res, prob = stats.ttest_ind(rep_enc[bill], nan_policy='omit')
rep_t1[bill] = [res,prob]
# + id="txkelB3pZAPL" colab_type="code" colab={}
# + id="dNN9MGvJZAGB" colab_type="code" colab={}
# + id="ahpKMlaqUl3O" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="ba3efc25-c36d-4371-b45b-c14e9935bb88"
tv, prob = stats.ttest_ind(vote_enc['infant_bill'], dem_enc['infant_bill'])
print(tv, prob)
print(type(tv), type(prob))
# + id="ic0wdQFSGDJG" colab_type="code" colab={}
import seaborn as sns
# + id="7k3yD_BTJG9e" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 224} outputId="bce5ecf0-ceed-47e3-c997-dae08293e8e6"
# + id="4ReCcVIvUsuE" colab_type="code" colab={}
| module2-sampling-confidence-intervals-and-hypothesis-testing/LS_DS_142_Sampling_Confidence_Intervals_and_Hypothesis_Testing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Multi-label classification
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
from fastai.conv_learner import *
PATH = 'data/planet/'
# +
# Data preparation steps if you are using Crestle:
os.makedirs('data/planet/models', exist_ok=True)
os.makedirs('/cache/planet/tmp', exist_ok=True)
# !ln -s /datasets/kaggle/planet-understanding-the-amazon-from-space/train-jpg {PATH}
# !ln -s /datasets/kaggle/planet-understanding-the-amazon-from-space/test-jpg {PATH}
# !ln -s /datasets/kaggle/planet-understanding-the-amazon-from-space/train_v2.csv {PATH}
# !ln -s /cache/planet/tmp {PATH}
# -
# ls {PATH}
# + [markdown] heading_collapsed=true
# ## Multi-label versus single-label classification
# + hidden=true
from fastai.plots import *
# + hidden=true
def get_1st(path): return glob(f'{path}/*.*')[0]
# + hidden=true
dc_path = "data/dogscats/valid/"
list_paths = [get_1st(f"{dc_path}cats"), get_1st(f"{dc_path}dogs")]
plots_from_files(list_paths, titles=["cat", "dog"], maintitle="Single-label classification")
# + [markdown] hidden=true
# In single-label classification each sample belongs to one class. In the previous example, each image is either a *dog* or a *cat*.
# + hidden=true
list_paths = [f"{PATH}train-jpg/train_0.jpg", f"{PATH}train-jpg/train_1.jpg"]
titles=["haze primary", "agriculture clear primary water"]
plots_from_files(list_paths, titles=titles, maintitle="Multi-label classification")
# + [markdown] hidden=true
# In multi-label classification each sample can belong to one or more clases. In the previous example, the first images belongs to two clases: *haze* and *primary*. The second image belongs to four clases: *agriculture*, *clear*, *primary* and *water*.
# -
# ## Multi-label models for Planet dataset
# +
from planet import f2
metrics=[f2]
f_model = resnet34
# -
label_csv = f'{PATH}train_v2.csv'
n = len(list(open(label_csv)))-1
val_idxs = get_cv_idxs(n)
# We use a different set of data augmentations for this dataset - we also allow vertical flips, since we don't expect vertical orientation of satellite images to change our classifications.
def get_data(sz):
tfms = tfms_from_model(f_model, sz, aug_tfms=transforms_top_down, max_zoom=1.05)
return ImageClassifierData.from_csv(PATH, 'train-jpg', label_csv, tfms=tfms,
suffix='.jpg', val_idxs=val_idxs, test_name='test-jpg')
data = get_data(256)
x,y = next(iter(data.val_dl))
y
list(zip(data.classes, y[0]))
plt.imshow(data.val_ds.denorm(to_np(x))[0]*1.4);
sz=64
data = get_data(sz)
data = data.resize(int(sz*1.3), 'tmp')
learn = ConvLearner.pretrained(f_model, data, metrics=metrics)
lrf=learn.lr_find()
learn.sched.plot()
lr = 0.2
learn.fit(lr, 3, cycle_len=1, cycle_mult=2)
lrs = np.array([lr/9,lr/3,lr])
learn.unfreeze()
learn.fit(lrs, 3, cycle_len=1, cycle_mult=2)
learn.save(f'{sz}')
learn.sched.plot_loss()
sz=128
learn.set_data(get_data(sz))
learn.freeze()
learn.fit(lr, 3, cycle_len=1, cycle_mult=2)
learn.unfreeze()
learn.fit(lrs, 3, cycle_len=1, cycle_mult=2)
learn.save(f'{sz}')
sz=256
learn.set_data(get_data(sz))
learn.freeze()
learn.fit(lr, 3, cycle_len=1, cycle_mult=2)
learn.unfreeze()
learn.fit(lrs, 3, cycle_len=1, cycle_mult=2)
learn.save(f'{sz}')
multi_preds, y = learn.TTA()
preds = np.mean(multi_preds, 0)
f2(preds,y)
# ### End
| Courses/DeepLearning_1/lesson2-image_models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Formação Cientista de Dados - Prof. <NAME>
# Funções
# +
def imprime():
print("esta é uma função")
imprime()
# +
#com parametro
def imprime(n):
print(n)
imprime("Impressão deste texto")
# +
#com retorno
def potencia(n):
return n * n
x = potencia(3)
print(x)
# +
#com valor default
def intervalo(inic=1,fim=10):
for n in range(inic, fim+1):
print(n)
x = intervalo(1,10)
y = intervalo()
# -
| 1.Pratica em Python/scripts/6.Funcoes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Recruitment limit simulations
# required libraries
import numpy as np
from scipy import stats
import pandas as pd
import os
def cellsCycling(ts,cells,rl,tau):
# a cell divides when it completes its cell cycle,
# meaning that the time remaining to cell division ('tr') reachs zero.
cells_that_divide = []
for cell_id in cells.keys():
# go through each cell
if not cells[cell_id]['dormant']:
cells[cell_id]['tc'] += 1 # updates cell cycle position (simulation time dependent)
if cells[cell_id]['td'] == cells[cell_id]['tc']: # checks if cell cycle is completed
# if the cell completes its cell cycle
cells_that_divide.append(cell_id)
else:
if cells[cell_id]['recruited']:
cells[cell_id]['awakeningDelay'] -= 1
if cells[cell_id]['awakeningDelay'] == 0:
cells[cell_id]['dormant'] = False
if cells[cell_id]['position'] >= rl and ts < tau and not cells[cell_id]['recruited']:
# signal pulse at time tau in lambda microns
cells = cycleShortening(cells,cell_id) # cell recruitment
cells = updatePositions(cells,cells_that_divide) # cell pushing mechanism
cells = cellsDivision(cells,cells_that_divide) # cell division
return cells
def cycleShortening(cells,cell_id):
# cell cycle shortening implementation
cycle_position = cells[cell_id]['tc']
cycle_length = cells[cell_id]['td']
g1_length = int(cycle_length*long_g1_proportion)
g1_reduction = int(cycle_length*g1_reduction_proportion)
s_length = int(cycle_length*long_s_proportion)
s_reduction = int(cycle_length*s_reduction_proportion)
g2m_length = int(cycle_length*long_g2m_proportion)
if 0 <= cycle_position <= g1_reduction:
# cell in the G1 skip
# G1 skipping part 1 (partial synchronization implementation part 1)
cells[cell_id]['tc'] = cycle_position-cycle_position
elif g1_reduction < cycle_position <= g1_length:
# cell in the rest of G1
# G1 skipping part 2 (partial synchronization implementation part 2)
cells[cell_id]['tc'] = cycle_position-g1_reduction
elif g1_length < cycle_position <= g1_length+s_length:
# cell in S phase
# S mapping (proportional mapping implementation)
cells[cell_id]['tc'] = int((cycle_position-g1_length)*((s_length-s_reduction)/s_length)+(g1_length-g1_reduction))
elif g1_length+s_length < cycle_position <= g1_length+s_length+g2m_length+2:
# cell in G2/M
cells[cell_id]['tc'] = cycle_position-g1_reduction-s_reduction
cells[cell_id]['td'] = cycle_length-g1_reduction-s_reduction
cells[cell_id]['recruited'] = True
cells[cell_id]['dormant'] = False
return cells
def updatePositions(cells,cells_that_divide):
# cell pushing mechanism implementation
movements = {}
for cell_id in cells.keys():
cell_movement = 0
for divided_cell in cells_that_divide:
if cells[cell_id]['position'] >= cells[divided_cell]['position']:
cell_movement += 1
movements[cell_id] = cell_movement
for cell_id in cells.keys():
cells[cell_id]['position'] = cells[cell_id]['position']+movements[cell_id]*cell_diameter
return cells
def cellsDivision(cells,cells_that_divide):
# creates new cells based on mothers properties
for cell_id in cells_that_divide:
cells[cell_id]['tc'] = 0
daughter_id = len(cells)
if cells[cell_id]['recruited']:
# daughters of recruited cells are also recruited cells
cells[cell_id]['td'] = lognormal(short_cycle_mean,short_cycle_std)
cells[daughter_id] = {'tc':0,
'td':lognormal(short_cycle_mean,short_cycle_std),
'recruited':True,
'position':cells[cell_id]['position']-cell_diameter,
'dormant':False,
'awakeningDelay':0}
else:
# daughters of non-recruited cells are also non-recruited cells
cells[cell_id]['td'] = lognormal(long_cycle_mean,long_cycle_std)
cells[daughter_id] = {'tc':0,
'td':lognormal(long_cycle_mean,long_cycle_std),
'recruited':False,
'position':cells[cell_id]['position']-cell_diameter,
'dormant':False,
'awakeningDelay':0}
return cells
def tc_distribution(td):
x = np.arange(0,td+1)
f = 2**(1-(x*p)/td)
fn = f/f.sum() # normalization
tc = np.random.choice(x, p=fn)
return tc
def lognormal(mu_x,dt_x,size=1,integer=True):
# Draw one value (or more if size > 1) from a discretized lognormal distribution
mu = np.log(mu_x**2/np.sqrt(mu_x**2+dt_x**2))
sigma = np.sqrt(np.log(1+dt_x**2/mu_x**2))
shape = sigma # Scipy's shape parameter
scale = np.exp(mu) # Scipy's scale parameter
distribution = stats.lognorm.rvs(scale=scale,s=shape,size=size)
if len(distribution) == 1:
if integer:
return int(distribution[0])
else:
return distribution[0]
else:
return distribution
def run():
# simulation run
# initial conditions
cells = {}
for cell_id in range(0,n0):
cell_key = cell_id
td = lognormal(long_cycle_mean,long_cycle_std)
tc = tc_distribution(td)
cells[cell_key] = {'td':td, # cell cycle length
'tc':tc, # cell cycle position
'position':(cell_key+1-n0)*cell_diameter,
'recruited':False,
'dormant':False,
'awakeningDelay':0}
g0_cells_number = int(n0*g0_prop)
cells_df = pd.DataFrame.from_dict(cells, orient='index')
g0_cells = cells_df[cells_df['tc'] <= long_g1].sample(g0_cells_number).index
cells_df.loc[g0_cells,'dormant'] = True
cells_df.loc[g0_cells,'awakeningDelay'] = awakeningDelay
cells = cells_df.to_dict(orient='index')
# time iteration
rl = {} # empty simulation output
ts = 0 # simulation time = 0
for ts in range(0,steps):
signal_pos = ts*(-l/tau)
cells = cellsCycling(ts,cells,signal_pos,tau)
cells_df = pd.DataFrame.from_dict(cells, orient='index')
rl[ts] = cells_df[cells_df['recruited']==False]['position'].max()
return rl
# +
# run parameters
n0_mean,n0_std = 196,2 # n0 mean and standar deviation
l_mean,l_std = 828,30 # lambda mean and standar deviation
tau_mean,tau_std = 85,12 # tau mean and standar deviation
steps = 1+24*8 # number of steps (in hours)
np.random.seed(0) # random seed
seeds_number = 1000 # number of simulations
# constants
cell_diameter = 13.2 # cell diameter
long_g1 = 152 # G1 length in long cycle
long_s = 179 # S length in long cycle
short_g1 = 22 # G1 length in short cycle
short_s = 88 # S length in short cycle
long_g2m = short_g2m = 9 # G2/M length in both, long and short cycle
long_cycle_mean = long_g1+long_s+long_g2m # long cell cycle mean
long_cycle_std = 32 # long cell cycle standar deviation
short_cycle_mean = short_g1+short_s+short_g2m # short cell cycle mean
short_cycle_std = 10 # short cell cycle standar deviation
long_g1_proportion = long_g1/long_cycle_mean # G1 proportion in the long cell cycle
long_s_proportion = long_s/long_cycle_mean # S proportion in the long cell cycle
long_g2m_proportion = long_g2m/long_cycle_mean # G2/M proportion in the long cell cycle
short_g1_proportion = short_g1/short_cycle_mean # G1 proportion in the short cell cycle
short_s_proportion = short_s/short_cycle_mean # S proportion in the short cell cycle
short_g2m_proportion = short_g2m/short_cycle_mean # G2/M proportion in the short cell cycle
g1_reduction_proportion = (long_g1-short_g1)/long_cycle_mean # proportion of G1 reduction in the long cell cycle
s_reduction_proportion = (long_s-short_s)/long_cycle_mean # proportion of S reduction in the long cell cycle
g0_prop = 0.12 # G0 cells proportion
awakeningDelay = 72 # G0 cells activation delay
p = 2 # tc_distribution perturbation
# directory name
root = './simulations/'
model = 'switchpoint/'
parameters = 'n0='+str(n0_mean)+'\n'+'l='+str(l_mean)+'\n'+'tau='+str(tau_mean)+'/'
path = root+model+parameters
if not os.path.isdir(path):
os.makedirs(path)
# simulations
for seed in range(1,seeds_number+1):
# parameters drawing
n0 = int(np.random.normal(n0_mean,n0_std))
l = int(np.random.normal(l_mean,l_std))
tau = int(np.random.normal(tau_mean,tau_std))
# simulation run
simulation = run()
# output file for each seed
parameters = 'seed='+str(seed)+'_n0='+str(n0)+'_'+'l='+str(l)+'_'+'tau='+str(tau)
data = pd.DataFrame.from_dict(simulation, orient='index')
outfile = open(path+parameters+'.csv', 'a')
data.to_csv(outfile, sep=',')
outfile.close()
| main/Simulating_recruitment_limit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:100% !important; }</style>"))
# #### This notebook was created to visualize homoplasic Single Nucleotide Variants using the t-SNE embeddings derived from the pairwise SNP distance matrix
# +
# %matplotlib inline
import os
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import matplotlib as mpl
import matplotlib.ticker as ticker
from pylab import MaxNLocator
import seaborn as sns; sns.set()
from matplotlib.colors import LogNorm
from matplotlib import gridspec
import itertools
#for exporting to Adobe Illustrator
mpl.rcParams['pdf.fonttype'] = 42
mpl.rcParams['ps.fonttype'] = 42
# -
# ## Load SNP genotype matrix and Annotation Files
# +
#load isolate annotation file (columns of Genotype Matrix)
isolate_annotation_DF = pd.read_pickle('/n/data1/hms/dbmi/farhat/Roger/inhost_TB_dynamics_project/genotypes_matrix_and_tSNE/Genotype_Filtered_2/genotypes_isolate_annotation.pkl')
#load SNP annotation file (rows of Genotype Matrix) with gene annotation information
SNP_annotation_DF = pd.read_pickle('/n/data1/hms/dbmi/farhat/Roger/inhost_TB_dynamics_project/genotypes_matrix_and_tSNE/Genotype_Filtered_2/genotypes_SNP_annotation.pkl')
#load Genotypes Matrix
genotypes_array = np.load('/n/data1/hms/dbmi/farhat/Roger/inhost_TB_dynamics_project/genotypes_matrix_and_tSNE/Genotype_Filtered_2/genotypes_matrix.npy')
# -
allele_code_map_dict = {'A':0 , 'C':1 , 'G':2 , 'T':3}
allele_code_map_dict_r = {0:'A' , 1:'C' , 2:'G' , 3:'T'}
isolate_annotation_DF.head()
np.shape(isolate_annotation_DF)
SNP_annotation_DF.head()
np.shape(SNP_annotation_DF)
np.shape(genotypes_array)
# ## t-SNE
# Import t-SNE embeddings
# - perplex = 175
# - lr = 1500
t_SNE_coords = np.loadtxt('/n/data1/hms/dbmi/farhat/Roger/inhost_TB_dynamics_project/genotypes_matrix_and_tSNE/tb_output/tSNE_embeddings_from_pairwise_SNP_dist_perplexity175_lr1500.txt')
t_SNE_coords
np.shape(t_SNE_coords)
# ### *Function* to plot den-SNE & label isolates with a mutant allele
plt.style.use('ggplot')
plt.rcParams['lines.linewidth']=0
plt.rcParams['axes.facecolor']='1.0'
plt.rcParams['xtick.color']='black'
plt.rcParams['axes.grid']=False
plt.rcParams['axes.edgecolor']='black'
plt.rcParams['grid.color']= '1.0'
plt.rcParams.update({'font.size': 9})
plt.rc('font', family='serif')
def t_SNE_label_isolates_with_SNP(pos_i , mut_allele , SNP_label , fig , ax):
#Get Genotypes for SNP
#get row index of genotypes matrix that corresponds to this row
SNP_site_row_index = SNP_annotation_DF[SNP_annotation_DF.pos == pos_i].index.values[0]
#get the genotypes for all isolates
SNP_site_genotypes = genotypes_array[SNP_site_row_index , :]
#construct a boolean array that is TRUE if isolate's genotype matches the mutant allele and FALSE otherwise
isolate_mutant_allele_match_bool = SNP_site_genotypes == allele_code_map_dict[mut_allele]
#construct a boolean array that is FALSE if isolate's genotype matches the mutant allele and TRUE otherwise
isolate_mutant_allele_no_match_bool = SNP_site_genotypes != allele_code_map_dict[mut_allele]
#Plot coordinates for each isolate
#isolates without mutant allele
other_isolates = ax.scatter( t_SNE_coords[: , 0][isolate_mutant_allele_no_match_bool] , t_SNE_coords[: , 1][isolate_mutant_allele_no_match_bool] , color = 'xkcd:golden yellow' , edgecolor = 'white' , linewidth = 0.15 , alpha = 0.40 , s = 13)
#isolates with mutant allele
mut_allele_isolates = ax.scatter( t_SNE_coords[: , 0][isolate_mutant_allele_match_bool] ,t_SNE_coords[: , 1][isolate_mutant_allele_match_bool] , color = 'xkcd:black' , edgecolor = 'white' , linewidth = 0.15 , alpha = 1.0 , s = 13)
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
for axis in ['bottom','left']:
ax.spines[axis].set_linewidth(0.5)
ax.tick_params(axis='y', which='major', labelsize=7.5, pad = 0.0, labelcolor = 'k', rotation = 90)
ax.tick_params(axis='x', which='major', labelsize=7.5, pad = 0.0, labelcolor = 'k')
#add legend
ax.legend((mut_allele_isolates, other_isolates), (SNP_label + ' (N = ' + str(sum(isolate_mutant_allele_match_bool)) + ')', 'isolates without mutant allele'), bbox_to_anchor=(0,0.97,1,0.2) , scatterpoints=1, loc='lower left' , ncol=1,fontsize=8 , mode="expand", borderaxespad=0, frameon=False, handletextpad=0.01)
#add axis labels
ax.set_xlabel('t-SNE 1' , fontsize = '7.5' , color = 'k')
ax.tick_params(axis='x', pad=0.0)
ax.set_ylabel('t-SNE 2', fontsize = '7.5', color = 'k')
ax.tick_params(axis='y', pad=0.0)
# #### Load the table of in-host SNPs that were detected to be *phylogentically convergent*
phy_conv_in_host_SNPs = pd.read_csv('/n/data1/hms/dbmi/farhat/Roger/inhost_TB_dynamics_project/CSV_files/rolling_DB_scrape/SNPs_in_multiple_global_lineages.csv')
phy_conv_in_host_SNPs = phy_conv_in_host_SNPs.loc[: , ['ref_position_and_alt_allele' , 'ref_position' , 'genomic_coord' , 'ref_allele' , 'alt_allele' , 'gene_id' , 'Gene_Category' , 'symbol' , 'SNP_type' , 'AA_change']]
phy_conv_in_host_SNPs.head()
# #### Create the gene labels & the SNP labels for each mutation
#manually enter the gene symbols for the intergenic regions
phy_conv_in_host_SNPs.loc[13 , 'symbol'] = 'promoter fabG1-inhA'
phy_conv_in_host_SNPs.loc[16 , 'symbol'] = 'int Rv2828c_Rv2829c'
phy_conv_in_host_SNPs.loc[21 , 'symbol'] = 'promoter embA-embB'
#manually enter the SNP labels for the intergenic regions
phy_conv_in_host_SNPs.rename(columns={"AA_change": "SNP_label"} , inplace = True)
phy_conv_in_host_SNPs.loc[13 , 'SNP_label'] = phy_conv_in_host_SNPs.loc[13 , 'ref_allele'] + str(phy_conv_in_host_SNPs.loc[13 , 'ref_position']) + phy_conv_in_host_SNPs.loc[13 , 'alt_allele']
phy_conv_in_host_SNPs.loc[16 , 'SNP_label'] = phy_conv_in_host_SNPs.loc[16 , 'ref_allele'] + str(phy_conv_in_host_SNPs.loc[16 , 'ref_position']) + phy_conv_in_host_SNPs.loc[16 , 'alt_allele']
phy_conv_in_host_SNPs.loc[21 , 'SNP_label'] = phy_conv_in_host_SNPs.loc[21 , 'ref_allele'] + str(phy_conv_in_host_SNPs.loc[21 , 'ref_position']) + phy_conv_in_host_SNPs.loc[21 , 'alt_allele']
phy_conv_in_host_SNPs
# ## Main Figure 7
# #### SNP_indices: 2, 6, 12, 17, 18
# +
figM7 = plt.figure(constrained_layout=True)
spec2 = gridspec.GridSpec(ncols=2, nrows=3, figure=figM7 , wspace=0.155, hspace=0.265)
figM7_ax1 = figM7.add_subplot(spec2[0, 0])
figM7_ax2 = figM7.add_subplot(spec2[0, 1])
figM7_ax3 = figM7.add_subplot(spec2[1, 0])
figM7_ax4 = figM7.add_subplot(spec2[1, 1])
figM7_ax5 = figM7.add_subplot(spec2[2, 0])
figM7_ax6 = figM7.add_subplot(spec2[2, 1])
################################################################################################################
circ_size = 13
alpha = 0.65
#Modern
#lineage 2 (N = 7,000)
L2 = figM7_ax1.scatter( t_SNE_coords[: , 0][isolate_annotation_DF.lineage_1 == '2'] , t_SNE_coords[: , 1][isolate_annotation_DF.lineage_1 == '2'] , color = 'xkcd:bright blue' , edgecolor = 'white' , linewidth = 0.15 , alpha = alpha , s = circ_size)
#lineage 4 (N = 7,000)
L4 = figM7_ax1.scatter( t_SNE_coords[: , 0][isolate_annotation_DF.lineage_1 == '4'] , t_SNE_coords[: , 1][isolate_annotation_DF.lineage_1 == '4'] , color = 'xkcd:red' , edgecolor = 'white' , linewidth = 0.15 , alpha = alpha , s = circ_size)
#Ancestral
#lineage 3 (N = 3,352)
L3 = figM7_ax1.scatter( t_SNE_coords[: , 0][isolate_annotation_DF.lineage_1 == '3'] , t_SNE_coords[: , 1][isolate_annotation_DF.lineage_1 == '3'] , color = 'xkcd:purple' , edgecolor = 'white' , linewidth = 0.15 , alpha = alpha , s = circ_size)
#lineage 1 (N = 2,802)
L1 = figM7_ax1.scatter( t_SNE_coords[: , 0][isolate_annotation_DF.lineage_1 == '1'] , t_SNE_coords[: , 1][isolate_annotation_DF.lineage_1 == '1'] , color = 'xkcd:pink' , edgecolor = 'white' , linewidth = 0.15 , alpha = alpha , s = circ_size)
#lineage 5 (N = 101)
L5 = figM7_ax1.scatter( t_SNE_coords[: , 0][isolate_annotation_DF.lineage_1 == '5'] , t_SNE_coords[: , 1][isolate_annotation_DF.lineage_1 == '5'] , color = 'xkcd:brown' , edgecolor = 'white' , linewidth = 0.15 , alpha = alpha , s = circ_size)
#lineage 6 (N = 97)
L6 = figM7_ax1.scatter( t_SNE_coords[: , 0][isolate_annotation_DF.lineage_1 == '6'] , t_SNE_coords[: , 1][isolate_annotation_DF.lineage_1 == '6'] , color = 'xkcd:green' , edgecolor = 'white' , linewidth = 0.15 , alpha = alpha , s = circ_size)
figM7_ax1.spines['right'].set_visible(False)
figM7_ax1.spines['top'].set_visible(False)
for axis in ['bottom','left']:
figM7_ax1.spines[axis].set_linewidth(0.5)
figM7_ax1.tick_params(axis='y', which='major', labelsize=7.5, pad = 0.0, labelcolor = 'k', rotation = 90)
figM7_ax1.tick_params(axis='x', which='major', labelsize=7.5, pad = 0.0, labelcolor = 'k')
#add legend
figM7_ax1.legend((L1, L2, L3, L4, L5, L6), ('1 (N = 2,802)', '2 (N = 7,000)', '3 (N = 3,352)', '4 (N = 7,000)', '5 (N = 101)', '6 (N = 97)') , bbox_to_anchor=(0,0.97,1,0.2) , scatterpoints=1 , loc='lower left', ncol=3, fontsize=8 ,mode="expand", borderaxespad=0 , frameon=False , handletextpad=0.01)
#add axis labels
figM7_ax1.set_xlabel('t-SNE 1' , fontsize = '7.5' , color = 'k')
figM7_ax1.tick_params(axis='x', pad=0.0)
figM7_ax1.set_ylabel('t-SNE 2', fontsize = '7.5', color = 'k')
figM7_ax1.tick_params(axis='y', pad=0.0)
################################################################################################################
SNP_i_index = 2
t_SNE_label_isolates_with_SNP(phy_conv_in_host_SNPs.loc[SNP_i_index , 'ref_position'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'alt_allele'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'symbol'] + ' ' + phy_conv_in_host_SNPs.loc[SNP_i_index , 'SNP_label'] , figM7 , figM7_ax2)
SNP_i_index = 6
t_SNE_label_isolates_with_SNP(phy_conv_in_host_SNPs.loc[SNP_i_index , 'ref_position'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'alt_allele'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'symbol'] + ' ' + phy_conv_in_host_SNPs.loc[SNP_i_index , 'SNP_label'] , figM7 , figM7_ax3)
SNP_i_index = 12
t_SNE_label_isolates_with_SNP(phy_conv_in_host_SNPs.loc[SNP_i_index , 'ref_position'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'alt_allele'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'symbol'] + ' ' + phy_conv_in_host_SNPs.loc[SNP_i_index , 'SNP_label'] , figM7 , figM7_ax4)
SNP_i_index = 17
t_SNE_label_isolates_with_SNP(phy_conv_in_host_SNPs.loc[SNP_i_index , 'ref_position'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'alt_allele'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'symbol'] + ' ' + phy_conv_in_host_SNPs.loc[SNP_i_index , 'SNP_label'] , figM7 , figM7_ax5)
SNP_i_index = 18
t_SNE_label_isolates_with_SNP(phy_conv_in_host_SNPs.loc[SNP_i_index , 'ref_position'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'alt_allele'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'symbol'] + ' ' + phy_conv_in_host_SNPs.loc[SNP_i_index , 'SNP_label'] , figM7 , figM7_ax6)
figM7.text(0.10, 0.89, 'A', fontsize=12 , fontweight='bold' , color = 'k')
figM7.text(0.51, 0.89, 'B', fontsize=12 , fontweight='bold' , color = 'k')
figM7.text(0.10, 0.62, 'C', fontsize=12 , fontweight='bold' , color = 'k')
figM7.text(0.51, 0.62, 'D', fontsize=12 , fontweight='bold' , color = 'k')
figM7.text(0.10, 0.35, 'E', fontsize=12 , fontweight='bold' , color = 'k')
figM7.text(0.51, 0.35, 'F', fontsize=12 , fontweight='bold' , color = 'k')
figM7 = plt.gcf()
figM7.set_size_inches(8.5, 14.0)
figM7.tight_layout()
file_name = '/n/data1/hms/dbmi/farhat/Roger/inhost_TB_dynamics_project/genotypes_matrix_and_tSNE/figures/t_SNE_plots/SNP_homoplasy_tSNE_plots/Figure 7.tif'
plt.savefig(file_name, bbox_inches='tight', dpi = 300 , transparent = False)
plt.show()
# -
# ## Supplementary Figure 10
def t_SNE_label_isolates_with_SNP(pos_i , mut_allele , SNP_label , fig , ax):
#Get Genotypes for SNP
#get row index of genotypes matrix that corresponds to this row
SNP_site_row_index = SNP_annotation_DF[SNP_annotation_DF.pos == pos_i].index.values[0]
#get the genotypes for all isolates
SNP_site_genotypes = genotypes_array[SNP_site_row_index , :]
#construct a boolean array that is TRUE if isolate's genotype matches the mutant allele and FALSE otherwise
isolate_mutant_allele_match_bool = SNP_site_genotypes == allele_code_map_dict[mut_allele]
#construct a boolean array that is FALSE if isolate's genotype matches the mutant allele and TRUE otherwise
isolate_mutant_allele_no_match_bool = SNP_site_genotypes != allele_code_map_dict[mut_allele]
#Plot coordinates for each isolate
#isolates without mutant allele
other_isolates = ax.scatter( t_SNE_coords[: , 0][isolate_mutant_allele_no_match_bool] , t_SNE_coords[: , 1][isolate_mutant_allele_no_match_bool] , color = 'xkcd:golden yellow' , edgecolor = 'white' , linewidth = 0.1 , alpha = 0.40 , s = 10.5)
#isolates with mutant allele
mut_allele_isolates = ax.scatter( t_SNE_coords[: , 0][isolate_mutant_allele_match_bool] ,t_SNE_coords[: , 1][isolate_mutant_allele_match_bool] , color = 'xkcd:black' , edgecolor = 'white' , linewidth = 0.1 , alpha = 1.0 , s = 10.5)
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
for axis in ['bottom','left']:
ax.spines[axis].set_linewidth(0.5)
ax.set_xticks([])
ax.set_yticks([])
#add legend
ax.legend([mut_allele_isolates], [SNP_label + ' (N = ' + str(sum(isolate_mutant_allele_match_bool)) + ')'], bbox_to_anchor=(0,0.97,1,0.2) , scatterpoints=1, loc='lower left' , ncol=1,fontsize=6.5 , mode="expand", borderaxespad=0, frameon=False, handletextpad=0.01)
# #### SNP_indices: 0, 1, 3, 4, 5, 7, 8, 9, 10, 11, 13
# +
figS10 = plt.figure(constrained_layout=True)
spec2 = gridspec.GridSpec(ncols=3, nrows=4, figure=figS10 , wspace=0.075, hspace=0.10)
figS10_ax1 = figS10.add_subplot(spec2[0, 0])
figS10_ax2 = figS10.add_subplot(spec2[0, 1])
figS10_ax3 = figS10.add_subplot(spec2[0, 2])
figS10_ax4 = figS10.add_subplot(spec2[1, 0])
figS10_ax5 = figS10.add_subplot(spec2[1, 1])
figS10_ax6 = figS10.add_subplot(spec2[1, 2])
figS10_ax7 = figS10.add_subplot(spec2[2, 0])
figS10_ax8 = figS10.add_subplot(spec2[2, 1])
figS10_ax9 = figS10.add_subplot(spec2[2, 2])
figS10_ax10 = figS10.add_subplot(spec2[3, 0])
figS10_ax11 = figS10.add_subplot(spec2[3, 1])
figS10_ax12 = figS10.add_subplot(spec2[3, 2])
################################################################################################################
circ_size = 10.5
alpha = 0.65
#Modern
#lineage 2 (N = 7,000)
L2 = figS10_ax1.scatter( t_SNE_coords[: , 0][isolate_annotation_DF.lineage_1 == '2'] , t_SNE_coords[: , 1][isolate_annotation_DF.lineage_1 == '2'] , color = 'xkcd:bright blue' , edgecolor = 'white' , linewidth = 0.1 , alpha = alpha , s = circ_size)
#lineage 4 (N = 7,000)
L4 = figS10_ax1.scatter( t_SNE_coords[: , 0][isolate_annotation_DF.lineage_1 == '4'] , t_SNE_coords[: , 1][isolate_annotation_DF.lineage_1 == '4'] , color = 'xkcd:red' , edgecolor = 'white' , linewidth = 0.1 , alpha = alpha , s = circ_size)
#Ancestral
#lineage 3 (N = 3,352)
L3 = figS10_ax1.scatter( t_SNE_coords[: , 0][isolate_annotation_DF.lineage_1 == '3'] , t_SNE_coords[: , 1][isolate_annotation_DF.lineage_1 == '3'] , color = 'xkcd:purple' , edgecolor = 'white' , linewidth = 0.1 , alpha = alpha , s = circ_size)
#lineage 1 (N = 2,802)
L1 = figS10_ax1.scatter( t_SNE_coords[: , 0][isolate_annotation_DF.lineage_1 == '1'] , t_SNE_coords[: , 1][isolate_annotation_DF.lineage_1 == '1'] , color = 'xkcd:pink' , edgecolor = 'white' , linewidth = 0.1 , alpha = alpha , s = circ_size)
#lineage 5 (N = 101)
L5 = figS10_ax1.scatter( t_SNE_coords[: , 0][isolate_annotation_DF.lineage_1 == '5'] , t_SNE_coords[: , 1][isolate_annotation_DF.lineage_1 == '5'] , color = 'xkcd:brown' , edgecolor = 'white' , linewidth = 0.1 , alpha = alpha , s = circ_size)
#lineage 6 (N = 97)
L6 = figS10_ax1.scatter( t_SNE_coords[: , 0][isolate_annotation_DF.lineage_1 == '6'] , t_SNE_coords[: , 1][isolate_annotation_DF.lineage_1 == '6'] , color = 'xkcd:green' , edgecolor = 'white' , linewidth = 0.1 , alpha = alpha , s = circ_size)
figS10_ax1.spines['right'].set_visible(False)
figS10_ax1.spines['top'].set_visible(False)
for axis in ['bottom','left']:
figS10_ax1.spines[axis].set_linewidth(0.5)
figS10_ax1.set_xticks([])
figS10_ax1.set_yticks([])
#add legend
figS10_ax1.legend((L1, L2, L3, L4, L5, L6), ('1 (N = 2,802)', '2 (N = 7,000)', '3 (N = 3,352)', '4 (N = 7,000)', '5 (N = 101)', '6 (N = 97)') , bbox_to_anchor=(0,0.97,1,0.2) , scatterpoints=1 , loc='lower left', ncol=2, fontsize=6.5 ,mode="expand", borderaxespad=0 , frameon=False , handletextpad=0.01)
################################################################################################################
SNP_i_index = 0
t_SNE_label_isolates_with_SNP(phy_conv_in_host_SNPs.loc[SNP_i_index , 'ref_position'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'alt_allele'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'symbol'] + ' ' + phy_conv_in_host_SNPs.loc[SNP_i_index , 'SNP_label'] , figS10 , figS10_ax2)
SNP_i_index = 1
t_SNE_label_isolates_with_SNP(phy_conv_in_host_SNPs.loc[SNP_i_index , 'ref_position'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'alt_allele'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'symbol'] + ' ' + phy_conv_in_host_SNPs.loc[SNP_i_index , 'SNP_label'] , figS10 , figS10_ax3)
SNP_i_index = 3
t_SNE_label_isolates_with_SNP(phy_conv_in_host_SNPs.loc[SNP_i_index , 'ref_position'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'alt_allele'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'symbol'] + ' ' + phy_conv_in_host_SNPs.loc[SNP_i_index , 'SNP_label'] , figS10 , figS10_ax4)
SNP_i_index = 4
t_SNE_label_isolates_with_SNP(phy_conv_in_host_SNPs.loc[SNP_i_index , 'ref_position'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'alt_allele'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'symbol'] + ' ' + phy_conv_in_host_SNPs.loc[SNP_i_index , 'SNP_label'] , figS10 , figS10_ax5)
SNP_i_index = 5
t_SNE_label_isolates_with_SNP(phy_conv_in_host_SNPs.loc[SNP_i_index , 'ref_position'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'alt_allele'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'symbol'] + ' ' + phy_conv_in_host_SNPs.loc[SNP_i_index , 'SNP_label'] , figS10 , figS10_ax6)
SNP_i_index = 7
t_SNE_label_isolates_with_SNP(phy_conv_in_host_SNPs.loc[SNP_i_index , 'ref_position'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'alt_allele'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'symbol'] + ' ' + phy_conv_in_host_SNPs.loc[SNP_i_index , 'SNP_label'] , figS10 , figS10_ax7)
SNP_i_index = 8
t_SNE_label_isolates_with_SNP(phy_conv_in_host_SNPs.loc[SNP_i_index , 'ref_position'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'alt_allele'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'symbol'] + ' ' + phy_conv_in_host_SNPs.loc[SNP_i_index , 'SNP_label'] , figS10 , figS10_ax8)
SNP_i_index = 9
t_SNE_label_isolates_with_SNP(phy_conv_in_host_SNPs.loc[SNP_i_index , 'ref_position'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'alt_allele'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'symbol'] + ' ' + phy_conv_in_host_SNPs.loc[SNP_i_index , 'SNP_label'] , figS10 , figS10_ax9)
SNP_i_index = 10
t_SNE_label_isolates_with_SNP(phy_conv_in_host_SNPs.loc[SNP_i_index , 'ref_position'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'alt_allele'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'symbol'] + ' ' + phy_conv_in_host_SNPs.loc[SNP_i_index , 'SNP_label'] , figS10 , figS10_ax10)
SNP_i_index = 11
t_SNE_label_isolates_with_SNP(phy_conv_in_host_SNPs.loc[SNP_i_index , 'ref_position'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'alt_allele'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'symbol'] + ' ' + phy_conv_in_host_SNPs.loc[SNP_i_index , 'SNP_label'] , figS10 , figS10_ax11)
SNP_i_index = 13
t_SNE_label_isolates_with_SNP(phy_conv_in_host_SNPs.loc[SNP_i_index , 'ref_position'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'alt_allele'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'symbol'] + ' ' + phy_conv_in_host_SNPs.loc[SNP_i_index , 'SNP_label'] , figS10 , figS10_ax12)
figS10 = plt.gcf()
figS10.set_size_inches(9.0, 13.0)
figS10.tight_layout()
file_name = '/n/data1/hms/dbmi/farhat/Roger/inhost_TB_dynamics_project/genotypes_matrix_and_tSNE/figures/t_SNE_plots/SNP_homoplasy_tSNE_plots/Figure 7 - figure supplement 1.tif'
plt.savefig(file_name, bbox_inches='tight', dpi = 300 , transparent = False)
plt.show()
# -
# ## Supplementary Figure 11
# #### SNP_indices: 14, 15, 16, 19, 20, 21, 22, 23, 24, 25
# +
figS11 = plt.figure(constrained_layout=True)
spec2 = gridspec.GridSpec(ncols=3, nrows=4, figure=figS11 , wspace=0.075, hspace=0.10)
figS11_ax1 = figS11.add_subplot(spec2[0, 0])
figS11_ax2 = figS11.add_subplot(spec2[0, 1])
figS11_ax3 = figS11.add_subplot(spec2[0, 2])
figS11_ax4 = figS11.add_subplot(spec2[1, 0])
figS11_ax5 = figS11.add_subplot(spec2[1, 1])
figS11_ax6 = figS11.add_subplot(spec2[1, 2])
figS11_ax7 = figS11.add_subplot(spec2[2, 0])
figS11_ax8 = figS11.add_subplot(spec2[2, 1])
figS11_ax9 = figS11.add_subplot(spec2[2, 2])
figS11_ax10 = figS11.add_subplot(spec2[3, 0])
figS11_ax11 = figS11.add_subplot(spec2[3, 1])
#figS11_ax12 = figS11.add_subplot(spec2[3, 2])
################################################################################################################
circ_size = 10.5
alpha = 0.65
#Modern
#lineage 2 (N = 7,000)
L2 = figS11_ax1.scatter( t_SNE_coords[: , 0][isolate_annotation_DF.lineage_1 == '2'] , t_SNE_coords[: , 1][isolate_annotation_DF.lineage_1 == '2'] , color = 'xkcd:bright blue' , edgecolor = 'white' , linewidth = 0.1 , alpha = alpha , s = circ_size)
#lineage 4 (N = 7,000)
L4 = figS11_ax1.scatter( t_SNE_coords[: , 0][isolate_annotation_DF.lineage_1 == '4'] , t_SNE_coords[: , 1][isolate_annotation_DF.lineage_1 == '4'] , color = 'xkcd:red' , edgecolor = 'white' , linewidth = 0.1 , alpha = alpha , s = circ_size)
#Ancestral
#lineage 3 (N = 3,352)
L3 = figS11_ax1.scatter( t_SNE_coords[: , 0][isolate_annotation_DF.lineage_1 == '3'] , t_SNE_coords[: , 1][isolate_annotation_DF.lineage_1 == '3'] , color = 'xkcd:purple' , edgecolor = 'white' , linewidth = 0.1 , alpha = alpha , s = circ_size)
#lineage 1 (N = 2,802)
L1 = figS11_ax1.scatter( t_SNE_coords[: , 0][isolate_annotation_DF.lineage_1 == '1'] , t_SNE_coords[: , 1][isolate_annotation_DF.lineage_1 == '1'] , color = 'xkcd:pink' , edgecolor = 'white' , linewidth = 0.1 , alpha = alpha , s = circ_size)
#lineage 5 (N = 101)
L5 = figS11_ax1.scatter( t_SNE_coords[: , 0][isolate_annotation_DF.lineage_1 == '5'] , t_SNE_coords[: , 1][isolate_annotation_DF.lineage_1 == '5'] , color = 'xkcd:brown' , edgecolor = 'white' , linewidth = 0.1 , alpha = alpha , s = circ_size)
#lineage 6 (N = 97)
L6 = figS11_ax1.scatter( t_SNE_coords[: , 0][isolate_annotation_DF.lineage_1 == '6'] , t_SNE_coords[: , 1][isolate_annotation_DF.lineage_1 == '6'] , color = 'xkcd:green' , edgecolor = 'white' , linewidth = 0.1 , alpha = alpha , s = circ_size)
figS11_ax1.spines['right'].set_visible(False)
figS11_ax1.spines['top'].set_visible(False)
for axis in ['bottom','left']:
figS11_ax1.spines[axis].set_linewidth(0.5)
figS11_ax1.set_xticks([])
figS11_ax1.set_yticks([])
#add legend
figS11_ax1.legend((L1, L2, L3, L4, L5, L6), ('1 (N = 2,802)', '2 (N = 7,000)', '3 (N = 3,352)', '4 (N = 7,000)', '5 (N = 101)', '6 (N = 97)') , bbox_to_anchor=(0,0.97,1,0.2) , scatterpoints=1 , loc='lower left', ncol=2, fontsize=6.5 ,mode="expand", borderaxespad=0 , frameon=False , handletextpad=0.01)
################################################################################################################
SNP_i_index = 14
t_SNE_label_isolates_with_SNP(phy_conv_in_host_SNPs.loc[SNP_i_index , 'ref_position'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'alt_allele'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'symbol'] + ' ' + phy_conv_in_host_SNPs.loc[SNP_i_index , 'SNP_label'] , figS11 , figS11_ax2)
SNP_i_index = 15
t_SNE_label_isolates_with_SNP(phy_conv_in_host_SNPs.loc[SNP_i_index , 'ref_position'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'alt_allele'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'symbol'] + ' ' + phy_conv_in_host_SNPs.loc[SNP_i_index , 'SNP_label'] , figS11 , figS11_ax3)
SNP_i_index = 16
t_SNE_label_isolates_with_SNP(phy_conv_in_host_SNPs.loc[SNP_i_index , 'ref_position'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'alt_allele'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'symbol'] + ' ' + phy_conv_in_host_SNPs.loc[SNP_i_index , 'SNP_label'] , figS11 , figS11_ax4)
SNP_i_index = 19
t_SNE_label_isolates_with_SNP(phy_conv_in_host_SNPs.loc[SNP_i_index , 'ref_position'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'alt_allele'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'symbol'] + ' ' + phy_conv_in_host_SNPs.loc[SNP_i_index , 'SNP_label'] , figS11 , figS11_ax5)
SNP_i_index = 20
t_SNE_label_isolates_with_SNP(phy_conv_in_host_SNPs.loc[SNP_i_index , 'ref_position'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'alt_allele'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'symbol'] + ' ' + phy_conv_in_host_SNPs.loc[SNP_i_index , 'SNP_label'] , figS11 , figS11_ax6)
SNP_i_index = 21
t_SNE_label_isolates_with_SNP(phy_conv_in_host_SNPs.loc[SNP_i_index , 'ref_position'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'alt_allele'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'symbol'] + ' ' + phy_conv_in_host_SNPs.loc[SNP_i_index , 'SNP_label'] , figS11 , figS11_ax7)
SNP_i_index = 22
t_SNE_label_isolates_with_SNP(phy_conv_in_host_SNPs.loc[SNP_i_index , 'ref_position'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'alt_allele'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'symbol'] + ' ' + phy_conv_in_host_SNPs.loc[SNP_i_index , 'SNP_label'] , figS11 , figS11_ax8)
SNP_i_index = 23
t_SNE_label_isolates_with_SNP(phy_conv_in_host_SNPs.loc[SNP_i_index , 'ref_position'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'alt_allele'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'symbol'] + ' ' + phy_conv_in_host_SNPs.loc[SNP_i_index , 'SNP_label'] , figS11 , figS11_ax9)
SNP_i_index = 24
t_SNE_label_isolates_with_SNP(phy_conv_in_host_SNPs.loc[SNP_i_index , 'ref_position'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'alt_allele'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'symbol'] + ' ' + phy_conv_in_host_SNPs.loc[SNP_i_index , 'SNP_label'] , figS11 , figS11_ax10)
SNP_i_index = 25
t_SNE_label_isolates_with_SNP(phy_conv_in_host_SNPs.loc[SNP_i_index , 'ref_position'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'alt_allele'] , phy_conv_in_host_SNPs.loc[SNP_i_index , 'symbol'] + ' ' + phy_conv_in_host_SNPs.loc[SNP_i_index , 'SNP_label'] , figS11 , figS11_ax11)
figS11 = plt.gcf()
figS11.set_size_inches(9.0, 13.0)
figS11.tight_layout()
file_name = '/n/data1/hms/dbmi/farhat/Roger/inhost_TB_dynamics_project/genotypes_matrix_and_tSNE/figures/t_SNE_plots/SNP_homoplasy_tSNE_plots/Figure 7 - figure supplement 2.tif'
plt.savefig(file_name, bbox_inches='tight', dpi = 300 , transparent = False)
plt.show()
| SNPs from Public Samples & tSNE Homoplasy Visualization/(F) t-SNE SNV Homoplasy Visualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.0.1
# language: julia
# name: julia-1.0
# ---
a = falses(3,4)
a
a[2,2] = 1
a
i = (3,3)
x = rand(3,4)#*[10,3,4]
#convert(Array{Int,2}, ceil.(x))
N = 50
b = [4;3;5]
r = [10;3;4]
D = length(b)
C = prod(r)
N, C
x = convert(Array{Int,2}, floor.(broadcast(+, broadcast(*, rand(D,N), r), b)))
x = convert(Array{Int,2}, ceil.(broadcast(*, rand(D,N), r)))
M = falses(r...)
size(M)
for i=1:size(x,2)
M[x[:,i]...] = 1
end
M
ndims(x)
a[i...] = 1
a
X = Set{Int64}()
for i=1:10
append(X, )
| factorize_sets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # PyTorch `nn` package
# ### The Learning Process of a Neural Network
#
# 
#
# <span class="fn"><i>Source:</i> [1] - _Deep Learning with PyTorch_ </span>
# ## Torch `autograd`
#
# **TL;DR**:
#
# (doc. reference: [torch autograd](https://pytorch.org/docs/stable/autograd.html))
#
# `torch.autograd` provides classes and functions implementing automatic differentiation of arbitrary scalar valued functions.
#
# It requires minimal changes to the existing code - you only need to declare `Tensor`s for which gradients should be computed with the `requires_grad=True` keyword.
# #### A quick look at the backend: Automatic Differentiation
# **TL; DR**: Automatic Differentiation lets you compute **exact** derivatives in **constant time**
#
# ###### Teaser
#
# Automatic Differentiation is the secret sauce that powers most of the existing Deep Learning frameworks (e.g. Pytorch or TensorFlow).
#
# In a nutshell, Deep learnin frameworks provide the (technical) infrastructure in which computing the derivative of a function takes as much time as evaluating the function. In particular, the design idea is: "you define a network with a loss function, and you get a gradient *for free*".
#
#
# **Differentiation** in general is becoming a **first class citizen** in programming languages with early work started by <NAME> of LLVM famework — see the [Differentiable Programming Manifesto](https://github.com/apple/swift/blob/master/docs/DifferentiableProgramming.md) for more detail.
# ###### Example
#
# Rather than talking about large neural networks, we will seek to understand automatic differentiation via a small problem borrowed from the book of *<NAME> (2008)*.
#
# In the following we will adopt their very same **three-part** notation.
#
# A function $f: \mathbb{R^n} \mapsto \mathbb{R^m}$ is constructed using intermediate variables $v_i$ such that:
#
# - variables $v_{i-n} = x_i$, $i = 1,\ldots,n$ are the input variables;
# - variables $v_i$, $i = 1,\ldots,l$ are the working **intermediate** variables;
# - variables $y_{m-i} = v_{l-i}$, $i = m-1,\ldots,0$ are the output variables.
#
# <img src="ad_example.png" class="maxw80" />
# The **traversal** of the graph and the **direction** in which gradients are actually computed defines the two modalities of AD:
#
# * **forward mode** AD;
# * **backward mode** AD.
# ###### Backward Mode
#
# AD in the reverse accumulation mode corresponds to a generalized backpropagation algorithm, in that it propagates derivatives backward from a given output. This is done by complementing each intermediate variable $v_i$ with an **adjoint**:
# $$
# \bar{v}_{i} = \frac{\partial y_i}{\partial v_i} = \displaystyle{\sum_{j:\text{child of i}} \bar{vj} \frac{\partial v_j}{\partial v_i}}
# $$
# <img src="backward_ad.png" class="maxw85" />
# <img src="https://github.com/google/tangent/raw/master/docs/toolspace.png" />
# There are various ways to implement this abstraction in its full generality, but an implementation requires more code than can easily appear here. The method we are interested here is the **tape-based** method:
#
# **tape-based**: This approach tracks the actual composed functions as they are called during execution of the forward pass. One name for this data structure is the *Wengert list*.
#
# $\rightarrow$ With the ordered sequence of computations in hand, it is then possible to walk backward through the list to compute the gradient.
#
# $\rightarrow$ The advantage of this is that it can more easily use all the features of the host language and the imperative execution is easier to understand.
#
# $\rightarrow$ The downside is that it can be more difficult to optimize the code and reuse computations across executions.
#
# The automatic differentiation in [PyTorch](https://pytorch.org/) roughly follows this model.
# ###### References and Futher Reading
# 1. [(*Paper*) Automatic Differentiation in Machine Learning: a Survey](https://arxiv.org/abs/1502.05767)
# ---
# ### `torch.nn` in a Nutshell
#
# Computational graphs and autograd are a very powerful paradigm for defining complex operators and automatically taking derivatives; however for large neural networks raw autograd can be a bit too low-level.
#
# When building neural networks we frequently think of arranging the computation into layers, some of which
# have learnable parameters which will be optimized during learning.
#
# In TensorFlow, packages like **Keras**, (old **TensorFlow-Slim**, and **TFLearn**) provide higher-level abstractions over raw computational graphs that are useful for building neural networks.
#
# In PyTorch, the `nn` package serves this same purpose.
#
# The `nn` package defines a set of `Module`s, which are roughly equivalent to neural network layers.
#
# A `Module` receives input `Tensor`s and computes output `Tensor`s, but may also hold internal state such as `Tensor`s containing learnable parameters.
#
# The `nn` package also defines a set of useful `loss` functions that are commonly used when
# training neural networks.
# ##### PyTorch Examples
#
# The following examples have been extracted from the [PyTorch Examples Repository](https://github.com/jcjohnson/pytorch-examples) by `@jcjohnson`
# In this example we use the `nn` package to implement our two-layer network:
import torch
# +
# N is batch size; D_in is input dimension;
# H is hidden dimension; D_out is output dimension.
N, D_in, H, D_out = 64, 1000, 100, 10
# Create random Tensors to hold inputs and outputs
x = torch.randn(N, D_in)
y = torch.randn(N, D_out)
# Use the nn package to define our model as a sequence of layers. nn.Sequential
# is a Module which contains other Modules, and applies them in sequence to
# produce its output. Each Linear Module computes output from input using a
# linear function, and holds internal Tensors for its weight and bias.
model = torch.nn.Sequential(
torch.nn.Linear(D_in, H),
torch.nn.ReLU(),
torch.nn.Linear(H, D_out),
)
# +
# The nn package also contains definitions of popular loss functions; in this
# case we will use Mean Squared Error (MSE) as our loss function.
loss_fn = torch.nn.MSELoss(reduction='sum')
learning_rate = 1e-4
# -
for t in range(500):
# Forward pass: compute predicted y by passing x to the model. Module objects
# override the __call__ operator so you can call them like functions. When
# doing so you pass a Tensor of input data to the Module and it produces
# a Tensor of output data.
y_pred = model(x)
# Compute and print loss. We pass Tensors containing the predicted and true
# values of y, and the loss function returns a Tensor containing the
# loss.
loss = loss_fn(y_pred, y)
if t % 50 == 0:
print(t, loss.item())
# Zero the gradients before running the backward pass.
model.zero_grad()
# Backward pass: compute gradient of the loss with respect to all the learnable
# parameters of the model. Internally, the parameters of each Module are stored
# in Tensors with requires_grad=True, so this call will compute gradients for
# all learnable parameters in the model.
loss.backward()
# Update the weights using gradient descent. Each parameter is a Tensor, so
# we can access its gradients like we did before.
with torch.no_grad():
for param in model.parameters():
param -= learning_rate * param.grad
# ### `torch.optim`
# Up to this point we have updated the weights of our models by manually mutating the Tensors holding learnable parameters (**using `torch.no_grad()` or `.data` to avoid tracking history in autograd**).
#
# This is not a huge burden for simple optimization algorithms like stochastic gradient descent, but in practice we often train neural networks using more sophisticated optimizers like `AdaGrad`, `RMSProp`,
# `Adam`.
#
# The optim package in PyTorch abstracts the idea of an optimization algorithm and provides implementations of commonly used optimization algorithms.
#
# Let's finally modify the previous example in order to use `torch.optim` and the `Adam` algorithm:
# ##### Model and Optimiser (w/ Parameters) at a glance
#
# 
#
# <span class="fn"><i>Source:</i> [1] - _Deep Learning with PyTorch_
# +
# N is batch size; D_in is input dimension;
# H is hidden dimension; D_out is output dimension.
N, D_in, H, D_out = 64, 1000, 100, 10
# Create random Tensors to hold inputs and outputs
x = torch.randn(N, D_in)
y = torch.randn(N, D_out)
# Use the nn package to define our model and loss function.
model = torch.nn.Sequential(
torch.nn.Linear(D_in, H),
torch.nn.ReLU(),
torch.nn.Linear(H, D_out),
)
loss_fn = torch.nn.MSELoss(reduction='sum')
# -
# Use the optim package to define an Optimizer that will update the weights of
# the model for us. Here we will use Adam; the optim package contains many other
# optimization algoriths. The first argument to the Adam constructor tells the
# optimizer which Tensors it should update.
learning_rate = 1e-4
optimizer = torch.optim.Adam(model.parameters(), lr=learning_rate)
for t in range(500):
# Forward pass: compute predicted y by passing x to the model.
y_pred = model(x)
# Compute and print loss.
loss = loss_fn(y_pred, y)
if t % 50 == 0:
print(t, loss.item())
# Before the backward pass, use the optimizer object to zero all of the
# gradients for the variables it will update (which are the learnable
# weights of the model). This is because by default, gradients are
# accumulated in buffers( i.e, not overwritten) whenever .backward()
# is called. Checkout docs of torch.autograd.backward for more details.
optimizer.zero_grad()
# Backward pass: compute gradient of the loss with respect to model
# parameters
loss.backward()
# Calling the step function on an Optimizer makes an update to its
# parameters
optimizer.step()
# ### Can we do better ?
# Possible scenario:
#
# - Specify models that are more complex than a sequence of existing (pre-defined) modules;
# - Customise the learning procedure (e.g. _weight sharing_ ?)
# - ?
#
# For these cases, **PyTorch** allows to define our own custom modules by subclassing `nn.Module` and defining a `forward` method which receives the input data (i.e. `Tensor`) and returns the output (i.e. `Tensor`).
#
# It is in the `forward` method that **all** the _magic_ of Dynamic Graph and `autograd` operations happen!
# ### PyTorch: Custom Modules
# Let's implement our **two-layers** model as a custom `nn.Module` subclass
class TwoLayerNet(torch.nn.Module):
def __init__(self, D_in, H, D_out):
"""
In the constructor we instantiate two nn.Linear modules and assign them as
member variables.
"""
super(TwoLayerNet, self).__init__()
self.linear1 = torch.nn.Linear(D_in, H)
self.hidden_activation = torch.nn.ReLU()
self.linear2 = torch.nn.Linear(H, D_out)
def forward(self, x):
"""
In the forward function we accept a Tensor of input data and we must return
a Tensor of output data. We can use Modules defined in the constructor as
well as arbitrary operators on Tensors.
"""
l1 = self.linear1(x)
h_relu = self.hidden_activation(l1)
y_pred = self.linear2(h_relu)
return y_pred
# +
# N is batch size; D_in is input dimension;
# H is hidden dimension; D_out is output dimension.
N, D_in, H, D_out = 64, 1000, 100, 10
# Create random Tensors to hold inputs and outputs
x = torch.randn(N, D_in)
y = torch.randn(N, D_out)
# -
# Construct our model by instantiating the class defined above
model = TwoLayerNet(D_in, H, D_out)
# Construct our loss function and an Optimizer. The call to model.parameters()
# in the SGD constructor will contain the learnable parameters of the two
# nn.Linear modules which are members of the model.
criterion = torch.nn.MSELoss(reduction='sum')
optimizer = torch.optim.SGD(model.parameters(), lr=1e-4)
for t in range(500):
# Forward pass: Compute predicted y by passing x to the model
y_pred = model(x)
# Compute and print loss
loss = criterion(y_pred, y)
if t % 50 == 0:
print(t, loss.item())
# Zero gradients, perform a backward pass, and update the weights.
optimizer.zero_grad()
loss.backward()
optimizer.step()
# ##### Last but not least
#
# ```python
# >>> optimizer = torch.optim.SGD(model.parameters(), lr=1e-4)
# ```
#
# Being `model` a subclass of `nn.Module`, `model.parameters()` will automatically capture all the `Layers/OP/Tensors/Parameters` that require gradient computation, so to feed to the `autograd` engine during the *backward* (optimisation) step.
# ###### `model.named_parameters`
for name_str, param in model.named_parameters():
print("{:21} {:19} {}".format(name_str, str(param.shape), param.numel()))
# **WAIT**: What happened to `hidden_activation` ?
# ```python
# self.hidden_activation = torch.nn.ReLU()
# ```
#
# So, it looks that we are registering in the constructor a submodule (`torch.nn.ReLU`) that has no parameters.
#
# Generalising, if we would've had **more** (hidden) layers, it would have required the definition of one of these submodules for each pair of layers (at least).
# ---
# ##### Introducing the Functional API
#
# PyTorch has functional counterparts of every `nn` module.
#
# By _functional_ here we mean "having no internal state", or, in other words, "whose output value is solely and fully determined by the value input arguments".
#
# Indeed, `torch.nn.functional` provides the many of the same modules we find in `nn`, but with all eventual parameters moved as an argument to the function call.
#
# For instance, the functional counterpart of `nn.Linear` is `nn.functional.linear`, which is a function that has signature `linear(input, weight, bias=None)`.
#
# The `weight` and `bias` parameters are **arguments** to the function.
# $\rightarrow$ For the curious minds: [The difference and connection between torch.nn and torch.nn.function from relu's various implementations](https://programmer.group/5d5a404b257d7.html)
# ###### Loss Functions
#
# ```python
# CrossEntropyLoss == LogSoftmax + NLLLoss
# BCEWithLogits == LogSigmoid + NLLLoss
# MSELoss(reduce=sum) == SSE
# ```
# ---
# ### References and Futher Reading:
#
# 1. [Deep Learning with PyTorch, Luca Antiga et. al.](https://www.manning.com/books/deep-learning-with-pytorch)
# 2. [(**Terrific**) PyTorch Examples Repo](https://github.com/jcjohnson/pytorch-examples) (*where most of the examples in this notebook have been adapted from*)
| tutorial_pytorch/2_torch_nn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="Tce3stUlHN0L"
# ##### Copyright 2019 The TensorFlow Authors.
#
# + cellView="form" colab={} colab_type="code" id="tuOe1ymfHZPu"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="MfBg1C5NB3X0"
# # Multi-worker training with Estimator
#
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/tutorials/distribute/multi_worker_with_estimator"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/tutorials/distribute/multi_worker_with_estimator.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/tutorials/distribute/multi_worker_with_estimator.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/tutorials/distribute/multi_worker_with_estimator.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="xHxb-dlhMIzW"
# ## Overview
#
# Note: While you can use Estimators with `tf.distribute` API, it's recommended to use Keras with `tf.distribute`, see [multi-worker training with Keras](multi_worker_with_keras.ipynb). Estimator training with `tf.distribute.Strategy` has limited support.
#
#
# This tutorial demonstrates how `tf.distribute.Strategy` can be used for distributed multi-worker training with `tf.estimator`. If you write your code using `tf.estimator`, and you're interested in scaling beyond a single machine with high performance, this tutorial is for you.
#
# Before getting started, please read the [distribution strategy](../../guide/distributed_training.ipynb) guide. The [multi-GPU training tutorial](./keras.ipynb) is also relevant, because this tutorial uses the same model.
#
# + [markdown] colab_type="text" id="MUXex9ctTuDB"
# ## Setup
#
# First, setup TensorFlow and the necessary imports.
# + colab={} colab_type="code" id="bnYxvfLD-LW-"
import tensorflow_datasets as tfds
import tensorflow as tf
tfds.disable_progress_bar()
import os, json
# + [markdown] colab_type="text" id="hPBuZUNSZmrQ"
# ## Input function
#
# This tutorial uses the MNIST dataset from [TensorFlow Datasets](https://www.tensorflow.org/datasets). The code here is similar to the [multi-GPU training tutorial](./keras.ipynb) with one key difference: when using Estimator for multi-worker training, it is necessary to shard the dataset by the number of workers to ensure model convergence. The input data is sharded by worker index, so that each worker processes `1/num_workers` distinct portions of the dataset.
# + colab={} colab_type="code" id="dma_wUAxZqo2"
BUFFER_SIZE = 10000
BATCH_SIZE = 64
def input_fn(mode, input_context=None):
datasets, info = tfds.load(name='mnist',
with_info=True,
as_supervised=True)
mnist_dataset = (datasets['train'] if mode == tf.estimator.ModeKeys.TRAIN else
datasets['test'])
def scale(image, label):
image = tf.cast(image, tf.float32)
image /= 255
return image, label
if input_context:
mnist_dataset = mnist_dataset.shard(input_context.num_input_pipelines,
input_context.input_pipeline_id)
return mnist_dataset.map(scale).cache().shuffle(BUFFER_SIZE).batch(BATCH_SIZE)
# + [markdown] colab_type="text" id="4BlcVXMhB59T"
# Another reasonable approach to achieve convergence would be to shuffle the dataset with distinct seeds at each worker.
# + [markdown] colab_type="text" id="8YFpxrcsZ2xG"
# ## Multi-worker configuration
#
# One of the key differences in this tutorial (compared to the [multi-GPU training tutorial](./keras.ipynb)) is the multi-worker setup. The `TF_CONFIG` environment variable is the standard way to specify the cluster configuration to each worker that is part of the cluster.
#
# There are two components of `TF_CONFIG`: `cluster` and `task`. `cluster` provides information about the entire cluster, namely the workers and parameter servers in the cluster. `task` provides information about the current task. The first component `cluster` is the same for all workers and parameter servers in the cluster, and the second component `task` is different on each worker and parameter server and specifies its own `type` and `index`. In this example, the task `type` is `worker` and the task `index` is `0`.
#
# For illustration purposes, this tutorial shows how to set a `TF_CONFIG` with 2 workers on `localhost`. In practice, you would create multiple workers on an external IP address and port, and set `TF_CONFIG` on each worker appropriately, i.e. modify the task `index`.
#
# Warning: *Do not execute the following code in Colab.* TensorFlow's runtime will attempt to create a gRPC server at the specified IP address and port, which will likely fail.
#
# ```
# os.environ['TF_CONFIG'] = json.dumps({
# 'cluster': {
# 'worker': ["localhost:12345", "localhost:23456"]
# },
# 'task': {'type': 'worker', 'index': 0}
# })
# ```
#
# + [markdown] colab_type="text" id="qDreJzTffAP5"
# ## Define the model
#
# Write the layers, the optimizer, and the loss function for training. This tutorial defines the model with Keras layers, similar to the [multi-GPU training tutorial](./keras.ipynb).
# + colab={} colab_type="code" id="WNvOn_OeiUYC"
LEARNING_RATE = 1e-4
def model_fn(features, labels, mode):
model = tf.keras.Sequential([
tf.keras.layers.Conv2D(32, 3, activation='relu', input_shape=(28, 28, 1)),
tf.keras.layers.MaxPooling2D(),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(64, activation='relu'),
tf.keras.layers.Dense(10)
])
logits = model(features, training=False)
if mode == tf.estimator.ModeKeys.PREDICT:
predictions = {'logits': logits}
return tf.estimator.EstimatorSpec(labels=labels, predictions=predictions)
optimizer = tf.compat.v1.train.GradientDescentOptimizer(
learning_rate=LEARNING_RATE)
loss = tf.keras.losses.SparseCategoricalCrossentropy(
from_logits=True, reduction=tf.keras.losses.Reduction.NONE)(labels, logits)
loss = tf.reduce_sum(loss) * (1. / BATCH_SIZE)
if mode == tf.estimator.ModeKeys.EVAL:
return tf.estimator.EstimatorSpec(mode, loss=loss)
return tf.estimator.EstimatorSpec(
mode=mode,
loss=loss,
train_op=optimizer.minimize(
loss, tf.compat.v1.train.get_or_create_global_step()))
# + [markdown] colab_type="text" id="P94PrIW_kSCE"
# Note: Although the learning rate is fixed in this example, in general it may be necessary to adjust the learning rate based on the global batch size.
# + [markdown] colab_type="text" id="UhNtHfuxCGVy"
# ## MultiWorkerMirroredStrategy
#
# To train the model, use an instance of `tf.distribute.experimental.MultiWorkerMirroredStrategy`. `MultiWorkerMirroredStrategy` creates copies of all variables in the model's layers on each device across all workers. It uses `CollectiveOps`, a TensorFlow op for collective communication, to aggregate gradients and keep the variables in sync. The [`tf.distribute.Strategy` guide](../../guide/distributed_training.ipynb) has more details about this strategy.
# + colab={} colab_type="code" id="1uFSHCJXMrQ-"
strategy = tf.distribute.experimental.MultiWorkerMirroredStrategy()
# + [markdown] colab_type="text" id="H47DDcOgfzm7"
# ## Train and evaluate the model
#
# Next, specify the distribution strategy in the `RunConfig` for the estimator, and train and evaluate by invoking `tf.estimator.train_and_evaluate`. This tutorial distributes only the training by specifying the strategy via `train_distribute`. It is also possible to distribute the evaluation via `eval_distribute`.
# + colab={} colab_type="code" id="BcsuBYrpgnlS"
config = tf.estimator.RunConfig(train_distribute=strategy)
classifier = tf.estimator.Estimator(
model_fn=model_fn, model_dir='/tmp/multiworker', config=config)
tf.estimator.train_and_evaluate(
classifier,
train_spec=tf.estimator.TrainSpec(input_fn=input_fn),
eval_spec=tf.estimator.EvalSpec(input_fn=input_fn)
)
# + [markdown] colab_type="text" id="XVk4ftYx6JAO"
# ## Optimize training performance
#
# You now have a model and a multi-worker capable Estimator powered by `tf.distribute.Strategy`. You can try the following techniques to optimize performance of multi-worker training:
#
# * *Increase the batch size:* The batch size specified here is per-GPU. In general, the largest batch size that fits the GPU memory is advisable.
# * *Cast variables:* Cast the variables to `tf.float` if possible. The official ResNet model includes [an example](https://github.com/tensorflow/models/blob/8367cf6dabe11adf7628541706b660821f397dce/official/resnet/resnet_model.py#L466) of how this can be done.
# * *Use collective communication:* `MultiWorkerMirroredStrategy` provides multiple [collective communication implementations](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/python/distribute/cross_device_ops.py).
# * `RING` implements ring-based collectives using gRPC as the cross-host communication layer.
# * `NCCL` uses [Nvidia's NCCL](https://developer.nvidia.com/nccl) to implement collectives.
# * `AUTO` defers the choice to the runtime.
#
# The best choice of collective implementation depends upon the number and kind of GPUs, and the network interconnect in the cluster. To override the automatic choice, specify a valid value to the `communication` parameter of `MultiWorkerMirroredStrategy`'s constructor, e.g. `communication=tf.distribute.experimental.CollectiveCommunication.NCCL`.
#
# Visit the [Performance section](../../guide/function.ipynb) in the guide to learn more about other strategies and [tools](../../guide/profiler.md) you can use to optimize the performance of your TensorFlow models.
#
# + [markdown] colab_type="text" id="AW0Hb2xM6EGX"
# ## Other code examples
#
# 1. [End to end example](https://github.com/tensorflow/ecosystem/tree/master/distribution_strategy) for multi worker training in tensorflow/ecosystem using Kubernetes templates. This example starts with a Keras model and converts it to an Estimator using the `tf.keras.estimator.model_to_estimator` API.
# 2. [Official models](https://github.com/tensorflow/models/tree/master/official), many of which can be configured to run multiple distribution strategies.
#
| site/en-snapshot/tutorials/distribute/multi_worker_with_estimator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
## What The Heck Is The Latent Position Matrix, Anyway?
# -
# ## When Should You Use ASE vs LSE?
# You'll note that both Adjacency Spectral Embedding - which embeds using only the adjacency matrix - and Laplacian Spectral Embedding - which embeds using the Laplacian - are both reasonable options to embed your network. When should you use one compared to the other?
# +
# Generate a network from an SBM
B = np.array([[0.02, 0.044, .002, .009],
[0.044, 0.115, .010, .042],
[.002, .010, .020, .045],
[.009, .042, .045, .117]])
n = [100, 100, 100, 100]
A, labels = sbm(n=n, p=B, return_labels=True)
# Instantiate an ASE model and find the embedding
ase = ASE(n_components=2)
embedding_ase = ase.fit_transform(A)
# LSE
lse = LSE(n_components=2)
embedding_lse = lse.fit_transform(A)
# plot
from graphbook_code import draw_layout_plot
fig, axs = plt.subplots(1, 2, figsize=(10, 5))
plot_latents(embedding_ase, labels=labels, ax=axs[0],
title="Adjacency Spectral Embedding");
plot_latents(embedding_lse, labels=labels, ax=axs[1],
title="Laplacian Spectral Embedding");
| network_machine_learning_in_python/representations/ch6/spectral_.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # "Learning for Gold"
# > "Machine learning with multiple targets"
#
# - toc: true
# - badges: true
# - image: images/int2/int2-gold.jpg
# - comments: true
# - author: <NAME>
# - categories: [machine learning, python, eda, cleaning, multi-target, regression]
# ## Introduction
#
# Let's say you've been hired by a gold extraction company.
#
# They're interested in developing a model that will predict the recovery of gold in their gold extraction process. We will explore the dataset, determine an appropriate error metric, and find a model that minimizes this error.
# 
# +
# collapse
from pathlib import Path
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from joblib import dump, load
sns.set()
from sklearn.metrics import make_scorer
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.impute import SimpleImputer
from sklearn.ensemble import RandomForestRegressor
from sklearn.multioutput import MultiOutputRegressor, RegressorChain
from sklearn.linear_model import SGDRegressor
from sklearn.metrics import mean_absolute_error
from sklearn.dummy import DummyRegressor
from sklearn.model_selection import RandomizedSearchCV, GridSearchCV, cross_validate
STATE = 10
# -
model_path = Path("../models")
file_paths = [
Path("../datasets") / ("gold_recovery_" + name + ".csv")
for name in ["train", "test", "full"]
]
train, X_test, full = (pd.read_csv(f, index_col='date') for f in file_paths)
# ## Exploratory Data Analysis
# The following transform allows us to separate categories into individual state, statetype and mineral to better understand how share value changes in each step.
# ### Filtration process
train.columns[train.columns.str.endswith("au")]
# ### Target Leakage
#
# We need to make sure we don't include features that will not be available at prediction time.
#
# We can determine which features these are by comparing our training set with the given test set that omits certain features.
# Examining the difference in columns between the training and testing set.
features = X_test.columns
train.loc[:, sorted(set(train.columns).difference(set(features)))].info()
# 
test = full.loc[X_test.index]
print(f"Training set shape: {train.shape}, Testing set shape: {test.shape}")
# ### Target Variable Creation
# In order to predict recovery, recovery must be generated as a target variable from rougher input, output and tails.
# 
# Where:
# - C - share of gold in concentrate right after flotation - `rougher.output.concentrate_au`
# - F - share of gold in the feed before flotation - `rougher.input.feed_au`
# - T - share of gold in rougher tails right after flotation - `rougher.output.tail_au`
# We're going to calculate gold recovery from the first and second process using the following function.
#
# I've added some filtration steps if any value is zero or negative.
def calc_recovery(C, F, T):
rec = ((C * (F - T)) / (F * (C - T))) * 100
return rec
# +
datasets = [train, test]
types = {
"recovery_first": [
"rougher.output.concentrate_au",
"rougher.input.feed_au",
"rougher.output.tail_au",
]
}
for d in datasets:
for name, cols in types.items():
d[name] = calc_recovery(*[d[t] for t in cols])
# -
# ### Missing and Unusual Values in Targets
# Strategy:
#
# - Drop NA values if contained in either target variables.
# - Drop values that are between 0 and 100 in the training and testing set.
# - EDA shows that values in the range of 40 - 98 % recovery helps reduce outlying values.
# - These will only be performed on the training set.
#
#
def clean_target(df, targets, lower=0, upper=100):
clean_na = ~df[targets].isna().any(axis=1)
clean_outrange = df[targets].apply(lambda x: x.between(lower, upper, inclusive=False)).all(axis=1)
return df.loc[clean_na & clean_outrange]
targets = ['recovery_first', 'final.output.recovery']
train, test = [clean_target(_, targets) for _ in [train, test]]
X_train, y_train = (
train[features],
train[["recovery_first", "final.output.recovery"]],
)
X_test, y_test = (
test[features],
test[["recovery_first", "final.output.recovery"]],
)
# There's a long tail distribtion, though I'm uncertain if these values are a true phenomenon.
# collapse
with sns.plotting_context('notebook', font_scale=1.5):
fig, axes = plt.subplots(nrows=2, sharex=True, figsize=[10,10])
sns.boxplot(train["recovery_first"], ax=axes[0])
sns.boxplot(train["final.output.recovery"], ax=axes[1])
fig.suptitle("Comparing Recovery Between First and Final Filtration Steps")
#collapse
with sns.plotting_context("notebook", font_scale=1.5):
fig, ax = plt.subplots(figsize=[10,10])
sns.scatterplot(data=train, x='recovery_first', y='final.output.recovery', alpha=0.25, ax=ax)
ax.set(title='The relationship between first and final steps of filteration recovery percentage')
# Here we can observe how the process creating a higher share of gold as it is processed, while the other minerals have a higher share in the tails.
#
# There also seems to be some long tails with the output.concentrate of au.
#
# The final stage is much more effective at increasing the share of gold in our solution.
#
# Because the second process is so much more effective, We can give more weight to the model that can give us a stronger prediction of a high yield at the final stage.
# +
# collapse
dfs = []
for group, col_names in types.items():
vis_df = pd.melt(
train[col_names], var_name="Extract Stage", value_name="Gold Concentration"
)
vis_df["group"] = group
dfs.append(vis_df)
vis_df = pd.concat(dfs)
vis_df["Extract Stage"] = (
vis_df["Extract Stage"]
.str.replace("final.output.concentrate_au", "output")
.str.replace("rougher.output.tail_au", "tails")
.str.replace("final.output.tail_au", "tails")
.str.replace("rougher.input.feed_au", "input")
)
vis_df.loc[
(vis_df["group"] == "final.output.recovery")
& (vis_df["Extract Stage"] == "rougher.output.concentrate_au"),
"Extract Stage",
] = (
"input"
)
vis_df["Extract Stage"] = vis_df["Extract Stage"].str.replace(
"rougher.output.concentrate_au", "output"
)
order = ["input", "output", "tails"]
with sns.plotting_context("notebook", font_scale=1.5):
g = sns.catplot(
x="group",
y="Gold Concentration",
data=vis_df,
col="Extract Stage",
kind="box",
col_order=order,
)
# -
# We can now compare metal shares throughout the purification process.
# +
#collapse
cats = [
"rougher.input.feed",
"rougher.output.concentrate",
"rougher.output.tail",
"final.output.concentrate",
"final.output.tail",
]
ele_df = train.loc[:, train.columns.str.match("|".join(cats))].melt(
var_name="name", value_name="share"
)
pattern = r"(?P<stage>\w+)\.(?P<statetype>\w+\.\w+)_(?P<mineral>\w\w)"
ele_df = ele_df.join(ele_df["name"].str.extract(pattern)).drop("name", axis=1)
with sns.plotting_context("notebook", font_scale=1.5):
g = sns.catplot(
kind="box",
hue="stage",
y="share",
data=ele_df,
col_order=["au", "ag", "pb"],
x="statetype",
col="mineral",
hue_order=["rougher", "final"],
order=["input.feed", "output.concentrate", "output.tail"],
)
g.set_xticklabels(rotation=30)
# -
# collapse
ele_df.loc[ele_df["mineral"] == "au"].groupby(
["statetype", "stage", "mineral"]
).describe()
#collapse
scatter_dat = train.loc[:, train.columns.str.match("|".join(cats))]
with sns.plotting_context("notebook", font_scale=1.5):
fig, axes = plt.subplots(nrows=2, figsize=[10,10])
sns.scatterplot(data=scatter_dat, x="rougher.input.feed_au", y="rougher.output.concentrate_au", ax=axes[0], alpha=0.25)
sns.scatterplot(data=scatter_dat, x="rougher.output.concentrate_au", y="final.output.concentrate_au", ax=axes[1], alpha=0.25)
# In order to determine if the training set is very representative, we can compare our features between training and testing sets. Here's and example of how we can do that.
#collapse
comp_feed = pd.concat(
[train["rougher.input.feed_size"], test["rougher.input.feed_size"]], axis=1
)
comp_feed.columns = ["train", "test"]
comp_feed = pd.melt(comp_feed, var_name="rougher input feed size", value_name="share")
sns.boxplot(data=comp_feed, x="rougher input feed size", y="share")
plt.show()
# ## Machine Learning
# 
# ### Preprocessing
# Based on exploring the dataset, I'm going to trim target training values to be in between 40 and 98 % recovery.
#
# Ideally this will yield a higher test score.
# +
targets = ['recovery_first', 'final.output.recovery']
train_c = clean_target(train, targets, lower=40, upper=98)
test_c = clean_target(test, targets)
print(f"After filtering, we are retaining {(train_c.shape[0]/train.shape[0]):.2%} of the training data")
print(f"After filtering, we are retaining {(test_c.shape[0]/test.shape[0]):.2%} of the testing data")
train = train_c
test = test_c
train[targets].describe()
# -
# ### Scoring
# 
# Because there are multiple levels of filtration, It's important to factor in both the rougher and final recovery for predictions.
# +
def calc_smape(y: np.array, y_pred: np.array) -> float:
"""Symmetric Mean Absolute Percentage Error"""
smape_s = (
np.mean(
(
np.abs(y_pred - y)
/ ((np.abs(y) + np.abs(y_pred)) / 2)
)
)
* 100
)
return smape_s
def smape_score(y: np.array, y_pred: np.array) -> float:
"""Combine the """
smape_r = calc_smape(y[:, 0], y_pred[:, 0])
smape_f = calc_smape(y[:, 1], y_pred[:, 1])
final_score = smape_r * 0.25 + smape_f * 0.75
return final_score
# -
# Here's what our target and training data look like.
y_train.head()
X_train.head()
# Now that we have a preprocessing plan, lets experiment with two algorithms.
#
#
# - A stochastic gradient decent algorithm with a grid search of reasonable hyperparameters.
# - A random forest with 200 estimators.
# Here's what the grid search looks like. Luckly, SGD can be very quick and we can rapidly search through our parameters.
# +
#collapse
prep = make_pipeline(SimpleImputer(), StandardScaler())
lin = make_pipeline(prep, MultiOutputRegressor(SGDRegressor(random_state=STATE)))
lhparams = {
"multioutputregressor__estimator__alpha": 10.0 ** -np.arange(1, 4),
"multioutputregressor__estimator__penalty": ["l2", "l1", "elasticnet"],
"multioutputregressor__estimator__loss": [
"huber",
"squared_loss",
"epsilon_insensitive",
],
"pipeline__simpleimputer__strategy": ["mean", "median"],
}
try:
lin_search = load(model_path / "lin_search.joblib")
except:
lin_search = GridSearchCV(
lin,
lhparams,
scoring=make_scorer(smape_score, greater_is_better=False),
cv=5,
n_jobs=-1,
)
lin_search.fit(X_train, y_train.values)
dump(lin_search, model_path / "lin_search.joblib")
print(f"Our best linear model produced a symmetric mean absolute error of {-lin_search.best_score_:.2f} percent on the mean validation score")
# -
# We can see the params from our search as well:
#collapse
lin_search.best_params_
# To see if it's worth experimenting with a random forest algorithm, let's try using one and seeing how well it compares to our best linear model.
rf_pipe = make_pipeline(
SimpleImputer(), RandomForestRegressor(n_estimators=200, random_state=STATE)
)
rf_cv = cross_validate(
rf_pipe,
X=X_train,
y=y_train.values,
cv=5,
scoring=make_scorer(smape_score, greater_is_better=False),
n_jobs=-1,
)
# The random forest regressor isn't producing very good validation scores. Let's stick with our best linear model.
rf_cv
# We can also determine a baseline to see how much more effective our best predictor is.
# +
#collapse
dummy_score = DummyRegressor(strategy='median')
dummy_score.fit(X=X_train, y=y_train.values)
dum_pred = dummy_score.predict(X_test)
print(f"""Our baseline using the median score of our training data
yields a SMAPE of: {smape_score(y=y_test.values, y_pred=dum_pred):.2f}% on the test dataset""")
# -
# ## Conclusion
#collapse
test_score = lin_search.score(X_test, y_test.values)
print(f"""Our best linear model produced a symmetric mean absolute error of:
{-test_score:.2f}% on our testing dataset""")
# We're able to get a slight improvement on recovery using our linear model, but not by much.
#
# If we're not satisfied with this score, it may be worth experimenting with more advanced imputation techniques, more complex models, and feature engineering.
| _notebooks/2020-07-30-int-project-gold.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="oS3PLcFyx2qV"
# Tratamiento de datos
# ==============================================================================
import numpy as np
import pandas as pd
# Gráficos
# ==============================================================================
import matplotlib.pyplot as plt
# Preprocesado y modelado
# ==============================================================================
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.metrics import accuracy_score
from sklearn.metrics import confusion_matrix
from sklearn.compose import ColumnTransformer
from sklearn.preprocessing import OneHotEncoder
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import train_test_split
from sklearn.model_selection import RepeatedKFold
from sklearn.model_selection import GridSearchCV
from sklearn.model_selection import ParameterGrid
from sklearn.inspection import permutation_importance
import multiprocessing
from sklearn.metrics import f1_score
# + [markdown] id="5Z5IC7jH4eQF"
# # PREPARO CSV
#
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="Ycop49sR0N19" outputId="77c7eae4-7ae5-4ad6-a8a6-b50d7274d779"
labels=pd.read_csv("../../csv/train_labels.csv")
labels.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 224} id="lvRAfRJx24Pv" outputId="2650951d-aa3f-45ae-bf7a-4ee22fed0428"
values=pd.read_csv("../../csv/train_values.csv")
values.head()
# + colab={"base_uri": "https://localhost:8080/"} id="qmLkwqs63Fhc" outputId="cad8ecc1-ac29-4577-f797-6b9252665827"
values.isnull().values.any()
# + id="6oEG2A77aVdk"
values = values.dropna()
# + colab={"base_uri": "https://localhost:8080/"} id="YFkxsAFa3PWR" outputId="1cf7131a-eb51-4790-8e49-5f127530f444"
labels.isnull().values.any()
# + colab={"base_uri": "https://localhost:8080/"} id="WT-SXWgn3Slp" outputId="3511c1bc-2674-470a-e059-3d7b76f55c7a"
values.info()
# + colab={"base_uri": "https://localhost:8080/"} id="2K9gA5gx3_va" outputId="2d0707d5-220a-4def-b923-9957799f7f5b"
to_be_categorized = ["land_surface_condition", "foundation_type", "roof_type",\
"position", "ground_floor_type", "other_floor_type",\
"plan_configuration", "legal_ownership_status", "geo_level_1_id"]
for row in to_be_categorized:
values[row] = values[row].astype("category")
values.info()
# + colab={"base_uri": "https://localhost:8080/"} id="AiuCbCfy4Dqw" outputId="249a7fe9-d3c7-44c6-e46c-412a75caef7c"
datatypes = dict(values.dtypes)
for row in values.columns:
if datatypes[row] != "int64" and datatypes[row] != "int32" and \
datatypes[row] != "int16" and datatypes[row] != "int8":
continue
if values[row].nlargest(1).item() > 32767 and values[row].nlargest(1).item() < 2**31:
values[row] = values[row].astype(np.int32)
elif values[row].nlargest(1).item() > 127:
values[row] = values[row].astype(np.int16)
else:
values[row] = values[row].astype(np.int8)
values.info()
# + colab={"base_uri": "https://localhost:8080/"} id="Q5n6W6gL4H48" outputId="8e62a5ef-c287-4e94-f3ce-23663da7b090"
labels.info()
# + colab={"base_uri": "https://localhost:8080/"} id="4GclKble4QMz" outputId="990b1764-a26b-455f-f70f-7eb666cfc75f"
labels["building_id"] = labels["building_id"].astype(np.int32)
labels["damage_grade"] = labels["damage_grade"].astype(np.int8)
labels.info()
# + id="DPJa5KvIm9lb"
important_values = values\
.merge(labels, on="building_id")
important_values.drop(columns=["building_id"], inplace = True)
# + colab={"base_uri": "https://localhost:8080/"} id="DEmFjcDa8DZL" outputId="9d38d283-c93e-4fd7-9bcd-90d6eec67ee1"
important_values.info()
# + colab={"base_uri": "https://localhost:8080/"} id="IK81aWD-7fZp" outputId="8f19e98d-137f-463c-b1fa-534d267b6bb4"
X_train, X_test, y_train, y_test = train_test_split(important_values.drop(columns = 'damage_grade'),
important_values['damage_grade'], test_size = 0.15, random_state = 123)
def feature_engieniere(df):
df["volume_percentage"] = df["height_percentage"] * df["area_percentage"]
df["less_than_25_years"] = df["age"] >= 25
df["more_than_20_metres"] = df["height_percentage"] <= 20
for i in [1,2,3]:
df[f"geo_{i}_height_avg"] = df.groupby(f"geo_level_{i}_id")["height_percentage"].transform(np.average)
df[f"geo_{i}_area_avg"] = df.groupby(f"geo_level_{i}_id")["area_percentage"].transform(np.average)
df[f"geo_{i}_age_avg"] = df.groupby(f"geo_level_{i}_id")["age"].transform(np.average)
df[f"geo_{i}_families_avg"] = df.groupby(f"geo_level_{i}_id")["count_families"].transform(np.average)
df[f"geo_{i}_volume_avg"] = df.groupby(f"geo_level_{i}_id")["volume_percentage"].transform(np.average)
df[f"geo_{i}_floors_avg"] = df.groupby(f"geo_level_{i}_id")["count_floors_pre_eq"].transform(np.average)
return df
#X_train = feature_engieniere(X_train)
#X_test = feature_engieniere(X_test)
category_cols = X_train.select_dtypes(include=['category', "bool"]).columns.to_list()
numeric_cols = X_train.select_dtypes(include=['int16', 'int8', "float64"]).columns.to_list()
preprocessor = ColumnTransformer([('onehot', OneHotEncoder(handle_unknown='ignore'), category_cols)],
remainder='passthrough')
X_train_prep = preprocessor.fit_transform(X_train)
X_test_prep = preprocessor.transform(X_test)
encoded_cat = preprocessor.named_transformers_['onehot'].get_feature_names(category_cols)
labels = np.concatenate([numeric_cols, encoded_cat])
X_train_prep = pd.DataFrame(X_train_prep, columns=labels)
X_test_prep = pd.DataFrame(X_test_prep, columns=labels)
X_train_prep.info()
# +
from sklearn.utils import class_weight
classes_weights = class_weight.compute_sample_weight(
class_weight='balanced',
y=y_train
)
classes_weights
# + colab={"base_uri": "https://localhost:8080/"} id="hXQVJXvVXBSi" outputId="1d0f6cf2-1164-4659-81fe-1b32873e702a"
param_grid = {'n_estimators' : [350,300],
'max_depth' : [9,8],
'subsample' : [0.6, 0.8],
'learning_rate' : [0.15, 0.45]
}
grid = GridSearchCV(
estimator = GradientBoostingClassifier(random_state=123),
param_grid = param_grid,
scoring = 'accuracy',
n_jobs = -1,
cv = RepeatedKFold(n_splits=3, n_repeats=1, random_state=123),
refit = True,
verbose = 1,
return_train_score = True
)
grid.fit(X = X_train_prep, y = y_train)
resultados = pd.DataFrame(grid.cv_results_)
resultados.filter(regex = '(param*|mean_t|std_t)') \
.drop(columns = 'params') \
.sort_values('mean_test_score', ascending = False) \
.head(4)
modelo_final = grid.best_estimator_
# + id="CIzewnimXJgM"
# Mejores hiperparámetros por validación cruzada
# ==============================================================================
print("----------------------------------------")
print("Mejores hiperparámetros encontrados (cv)")
print("----------------------------------------")
print(grid.best_params_, ":", grid.best_score_, grid.scoring)
# + id="lbIHxIF3XONH"
modelo_final = GradientBoostingClassifier(n_estimators = 305,
max_depth = 9,
min_samples_split = 2,
min_samples_leaf = 3,
subsample=0.6,
verbose=True,
learning_rate=0.15)
modelo_final.fit(X = X_train_prep, y = y_train, sample_weight = classes_weights)
predicciones = modelo_final.predict(X = X_test_prep)
mat_confusion = confusion_matrix(
y_true = y_test,
y_pred = predicciones
)
accuracy = accuracy_score(
y_true = y_test,
y_pred = predicciones,
normalize = True
)
print("Matriz de confusión")
print("-------------------")
print(mat_confusion)
print("")
print(f"El accuracy de test es: {100 * accuracy} %")
# + id="-IYEzyZqqFOm"
y_preds = modelo_final.predict(X_test_prep)
f1_score(y_test, y_preds, average='micro')
# + id="Xx6ZvE8ms78f"
test_values = pd.read_csv('test_values.csv', index_col = "building_id")
test_values
# + id="ljvF19v7tOLa"
for row in to_be_categorized:
test_values[row] = test_values[row].astype("category")
datatypes = dict(test_values.dtypes)
for row in test_values.columns:
if datatypes[row] != "int64" and datatypes[row] != "int32" and \
datatypes[row] != "int16" and datatypes[row] != "int8":
continue
if test_values[row].nlargest(1).item() > 32767 and test_values[row].nlargest(1).item() < 2**31:
test_values[row] = test_values[row].astype(np.int32)
elif test_values[row].nlargest(1).item() > 127:
test_values[row] = test_values[row].astype(np.int16)
else:
test_values[row] = test_values[row].astype(np.int8)
cat_cols = test_values.select_dtypes(include=['category', "bool"]).columns.to_list()
num_cols = test_values.select_dtypes(include=['int16', 'int8', "float64"]).columns.to_list()
preprocessor_2 = ColumnTransformer([('onehot', OneHotEncoder(handle_unknown='ignore'), cat_cols)],
remainder='passthrough')
test_values_reduce = preprocessor_2.fit_transform(test_values)
en_cat = preprocessor_2.named_transformers_['onehot'].get_feature_names(cat_cols)
labels_2 = np.concatenate([num_cols, en_cat])
test_values_reduce = pd.DataFrame(test_values_reduce, columns=labels_2)
test_values_reduce.info()
# + id="iBa8hG9Kvsed"
preds = modelo_final.predict(test_values_reduce)
submission_format = pd.read_csv("submission_format.csv", index_col = "building_id")
my_submission = pd.DataFrame(data=preds,
columns=submission_format.columns,
index=submission_format.index)
# + id="dO68hXsEwGRO"
my_submission.to_csv('gradient_boosting_model_1.csv')
# !head gradient_boosting_model_1.csv
| src/GradientBoosting/jf-model-1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.3 64-bit (''base'': conda)'
# metadata:
# interpreter:
# hash: bc017789cbaa633e1b24bc52b8280b5aad5b4cc01b03a9facb312052cb83bb85
# name: python3
# ---
# +
# mounting Google Drive to Colab VM.
from google.colab import drive
drive.mount('/content/drive', force_remount=True)
# Python interpreter of the Colab VM has to be able to load py files from within it
import sys
FOLDERPATH = "image-classification/"
sys.path.append('/content/drive/My Drive/{}'.format(FOLDERPATH))
# Import needed modules
from src.models.train_model import *
# -
| notebooks/classification-example-colab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="H8ThuCTjEjuG" colab_type="text"
# # Notebook destinado a segunda parte do PP1 da disciplina de RNA 2020.1 - Colaboradores <NAME>, <NAME> e <NAME>.
# + [markdown] id="EkFE5yTuNiuZ" colab_type="text"
# ## Importando as bibliotecas necessárias
# + id="vQrxap_dHVT5" colab_type="code" colab={}
import warnings
warnings.filterwarnings('ignore')
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
plt.style.use('ggplot')
plt.style.use('grayscale')
# + [markdown] id="WK5yHD74JW4b" colab_type="text"
# ## Importando o dataset limpo
# + [markdown] id="UL4Vu8BPDeEK" colab_type="text"
# > Formas alternativas de leitura do dataset do github
# + id="1IoqEcVSCIiU" colab_type="code" colab={}
#pd.read_csv('https://raw.githubusercontent.com/userddssilva/Analise-COVID19-PP1-RNA2020.1/master/dataset/dataset_limpo_covid19_manaus.csv?token=<KEY>')
#.read_csv('https://raw.githubusercontent.com/userddssilva/Analise-COVID19-PP1-RNA2020.1/master/dataset/dataset_covid19_manaus.csv?token=AMHWGQPMBW2HL4RB3VNPKOC7H7CHE', encoding='ISO-8859-1', sep=';')
# + id="FO36eUxqJck8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 419} outputId="68336afa-1ad3-430e-d624-88eb95ef1563"
df_dataset = pd.read_csv('/content/dataset_limpo_covid19_manaus.csv')
df_dataset
# + [markdown] id="JxLI9EUkOWUB" colab_type="text"
# **1. Construa um histograma denotando a quantidade de casos nos 10 bairros em que houve mais casos registrados. Inclua todos os bairros remanescentes em uma categoria denominada “Outros.” Denote as informações de maneira percentual.**
# + id="WN4EBBh9wv7g" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="2a31efb9-b1f3-45e8-85eb-e4037ddef8d0"
x = df_dataset.bairro
plt.figure(figsize=(30,10))
plt.hist(x, bins=300, density=100)
plt.ylabel('quantidade de casos')
plt.xlabel('bairros')
plt.show()
values_by_bairro = df_dataset.bairro.value_counts()
y = values_by_bairro[0:10].sort_values().values
x = list(values_by_bairro[0:10].sort_values().keys())
n_outros = sum(values_by_bairro[11:].sort_values().values)
y = np.append(y, [n_outros])
y = (y/sum(y))*100
x.append('OUTROS')
# Bring some raw data.
frequencies = y
# In my original code I create a series and run on that,
# so for consistency I create a series from the list.
freq_series = pd.Series(frequencies)
x_labels = x
# Plot the figure.
plt.figure(figsize=(12, 8))
ax = freq_series.plot(kind='bar')
ax.set_title('Distribuição percentual dos bairros com mais casos')
ax.set_xlabel('Bairros')
ax.set_ylabel('Percentual')
ax.set_xticklabels(x_labels)
rects = ax.patches
# Make some labels.
labels = ["%.2f%%" % i for i in y]
for rect, label in zip(rects, labels):
height = rect.get_height()
ax.text(rect.get_x() + rect.get_width() / 2, height + 5, label,
ha='center', va='bottom')
plt.show()
# + [markdown] id="s3iaDA84OwFA" colab_type="text"
# **2. Denote, por sexo, o boxplot da idade dos casos confirmados. Há outliers?**
# + id="S-PD1oxCw1Ia" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 592} outputId="1142d037-cc54-47f3-e3bd-8fed98f24fcd"
fame = df_dataset[df_dataset['sexo'] == 'F'].idade.values
male = df_dataset[df_dataset['sexo'] == 'M'].idade.values
plt.figure(figsize=(10, 10))
plt.boxplot([fame, male], labels=('Femino', 'Masculino'))
plt.show()
# + [markdown] id="3HHlHVltO31n" colab_type="text"
# **3. Denote em um gráfico de barras o número de novos casos por dia, considerando os 10 últimos dias na base de dados.**
# + id="SRtOqFZBy_XV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="e4279eb4-fd10-496d-9fa5-61b6a2dbee60"
dates = df_dataset
#datas transformadas pro formato Ano-Mês-Dia
dates['dt_notificacao'] = pd.to_datetime(dates['dt_notificacao'], dayfirst=True)
#datas ordenadas
dates = dates.sort_values(by='dt_notificacao')
#obetenção do número de casos
dates_cases = dates.groupby(['dt_notificacao']).size()
n = dates_cases.size
n = n-10
cases = dates_cases[n:].values
casos = []
for i in cases:
casos.append(i)
#salvando as 10 datas
dates = dates.groupby(['dt_notificacao']).head(1)
date = dates.dt_notificacao.values
datas = []
for i in range(len(date)-10,len(date)):
datas.append(date[i])
#Transformando as datas para o formato dia/mês/ano
for i in range(0,len(datas)):
data = pd.to_datetime(str(datas[i]))
datas[i] = data.strftime('%d/%m/%Y')
#Plotando o gráfico
plt.barh(datas, casos)
plt.ylabel('Data da Notificação')
plt.xlabel('Casos')
plt.show()
# + id="eputw4GpXRN1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 359} outputId="95ce3b85-5d59-4ec4-8805-94c32ebb41ad"
df_dataset.groupby(by=['dt_notificacao','conclusao']).count().reset_index().sort_values(by='dt_notificacao').tail(10)
# + [markdown] id="kw8lsMM7O9pn" colab_type="text"
# **4. Repita o gráfico anterior considerando o número de casos recuperado.**
# + id="GjEeqru68zVO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="1c4e55a7-2f48-4150-dc89-b3350528bd42"
dates = df_dataset
#datas transformadas pro formato Ano-Mês-Dia
dates['dt_notificacao'] = pd.to_datetime(dates['dt_notificacao'], dayfirst=True)
#datas ordenadas
dates = dates.sort_values(by='dt_notificacao')
#obetenção do número de casos
dates_cases = dates.groupby(['dt_notificacao','conclusao']).size()
n = dates_cases.size
n = n-10
cases = dates_cases[n:].values
casos = []
for i in cases:
casos.append(i)
#obtendo status da recuperacao
conclusao = []
for i in dates.conclusao:
conclusao.append(i)
tam = len(conclusao)-10
conclusao = conclusao[tam:]
#obter quantidade de recuperados
df_casos_recuperados = dates.groupby(['dt_notificacao','conclusao']).size()
df_casos_recuperados
qtde_conclusoes = []
for i in df_casos_recuperados:
qtde_conclusoes.append(i)
qtde_conclusoes = qtde_conclusoes[len(qtde_conclusoes)-10:]
#salvando as 10 datas
dates = dates.groupby(['dt_notificacao','conclusao']).head(1)
date = dates.dt_notificacao.values
#salvando as 10 datas
dates = dates.groupby(['dt_notificacao']).head(1)
date = dates.dt_notificacao.values
datas = []
for i in range(len(date)-10,len(date)):
datas.append(date[i])
#Transformando as datas para o formato dia/mês/ano
for i in range(0,len(datas)):
data = pd.to_datetime(str(datas[i]))
datas[i] = data.strftime('%d/%m/%Y')
df_index_conclusoes = dates.conclusao
index_casos_recuperados = []
for i in df_index_conclusoes:
index_casos_recuperados.append(i)
tam2 = len(index_casos_recuperados)-10
index_casos_recuperados = index_casos_recuperados[tam2:]
label_recuperado = []
for i in range(0,10):
if (index_casos_recuperados[i]=="Recuperado"):
label_recuperado.append(qtde_conclusoes[i])
else:
label_recuperado.append(0)
#Plotando o gráfico
plt.barh(datas, label_recuperado)
plt.ylabel('Notificação do Caso')
plt.xlabel('Casos Recuperados')
plt.show()
# + [markdown] id="zBBwiYZdPKqw" colab_type="text"
# **5. Construa um histograma que denote a quantidade percentual de casos por grupo etário, considerando que cada grupo contempla uma década (0 a 10 anos, 11 a 20 anos, etc.).**
# + id="0ia50QTP80Xu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 337} outputId="e96d39f4-68f1-49fb-8c78-7dac830a398b"
idades = df_dataset.idade.values
max_val = idades.max()
min_val = idades.min()
amount_intervals = []
a = 0
k = []
for intervalo in range(10, max_val+10, 10):
b = intervalo
amount_interval = df_dataset[(df_dataset['idade'] >= a) & (df_dataset['idade'] <= b)].shape[0]
amount_intervals.append(amount_interval)
k.append(b)
a = intervalo +1
amount_intervals = np.array(amount_intervals)
amount_intervals = (amount_intervals/sum(amount_intervals))*100
x_labels = ['0-10'] + [str(i-10)+'-'+str(i) for i in k[1:]]
plt.figure(figsize=(10, 5))
plt.bar(x_labels, amount_intervals, width=1)
plt.xlabel('Intervalo das idades')
plt.ylabel('Porcentagem de pessoas')
plt.show()
# + [markdown] id="3hb6fw2hPU3W" colab_type="text"
# **6. Elabore um gráfico que mostra o cumulativo de casos notificados ao longo do tempo.**
# + id="R8J23tAz_JHi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 391} outputId="1eee249c-65d5-4506-95ff-e687fc1c10f9"
#obter as datas
dates = df_dataset
#datas transformadas pro formato Ano-Mês-Dia
dates['dt_notificacao'] = pd.to_datetime(dates['dt_notificacao'], dayfirst=True)
#datas ordenadas
dates = dates.sort_values(by='dt_notificacao')
date = dates.dt_notificacao.values
# casos
datas = []
for i in range(0,len(date)):
datas.append(date[i])
casos = []
for i in range(0,len(datas)):
casos.append(1)
for i in range(0,len(datas)):
if (i+1<len(datas)):
casos[i+1] = casos[i] + casos[i+1]
fig, ax = plt.subplots(figsize=(10,6))
ax.plot(datas,casos,label="casos")
ax.set_xlabel('Tempo (Dias)')
ax.set_ylabel('Quantidade de Casos')
plt.show()
# + [markdown] id="Cjz4icH8PbgB" colab_type="text"
# **7. Faça um gráfico do tipo scatterplot que denote a idade versus o número total de casos registrado para aquela idade. Aproveite o processamento efetuado para o cálculo da correlação. É possível observar alguma tendência?**
# + id="vDXCnArq_KzM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="81473639-0c1f-4bc7-d1dd-22d536e61635"
data = df_dataset.groupby(['idade']).size()
y = data.index
x = data.values
plt.scatter(x,y)
plt.ylabel('Idade')
plt.xlabel('Total de casos')
plt.show()
| visualizacao_de_dados.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.5 64-bit
# name: python3
# ---
# + id="QVOHeIOgD_4X" outputId="948f7f53-9498-4938-b944-2841991245b6" colab={"base_uri": "https://localhost:8080/"}
from bs4 import BeautifulSoup
import requests
r = requests.get('https://bama.ir/car/all-brands/all-models/all-trims?price=30-40&page=1')
soup = BeautifulSoup(r.text,'html.parser')
# val = soup.find_all('h2')
val = soup.find('div', attrs = {'itemprop':'name'})
print(val)
| Chapter 4/test2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# %reload_ext autoreload
# %autoreload 2
# %matplotlib qt5
# +
import numpy as np
import matplotlib.pyplot as plt
from skimage import io
from skimage.transform import hough_circle, hough_circle_peaks, hough_ellipse
from skimage.feature import canny
from skimage.morphology import remove_small_objects
from skimage.draw import circle_perimeter, circle
from skimage.util import img_as_ubyte
from pathlib import Path
from skimage import data, color
plt.rcParams["figure.figsize"] = (20, 10)
import torch
from narsil.segmentation.network import basicUnet, smallerUnet
from narsil.liverun.utils import padTo32
import math
from datetime import datetime
import time
from scipy.signal import find_peaks
from skimage.filters import gaussian
# +
#imgpath = Path('/home/pk/Documents/realtimeData/hetero40x/Pos103/phaseFast/img_000000017.tiff')
#imgpath = Path('/home/pk/Documents/EXP-21-BY1006/therun/Pos11/phase/img_000000008.tiff')
# -
# device = torch.device("cuda:1" if torch.cuda.is_available() else "cpu")
# #cellSegModelPath = Path('/home/pk/Documents/models/mixed10epochs_betterscale_contrastAdjusted1.pth')
#
# modelPath = Path('/home/pk/Documents/models/channels.pth')
# pad = padTo32()
# with torch.no_grad():
# cellNetState = torch.load(modelPath)
#
# if cellNetState['modelParameters']['netType'] == 'big':
# cellSegNet = basicUnet(cellNetState['modelParameters']['transposeConv'])
# elif cellNetState['modelParameters']['netType'] == 'small':
# cellSegNet = smallerUnet(cellNetState['modelParameters']['transposeConv'])
#
# cellSegNet.load_state_dict(cellNetState['model_state_dict'])
# cellSegNet.to(device)
# cellSegNet.eval()
def imgFilenameFromNumber(number):
if number == 0:
num_digits = 1
else:
num_digits = int(math.log10(number)) + 1
imgFilename = 'img_' + '0' * (9 - num_digits) + str(number) + '.tiff'
return imgFilename
def barcodesFromPeaks(one_img, minLengthOfChannel= 200,
minPeaksDistance=25, gapWidth=48,
numChannels=21):
hist = np.sum(one_img, axis = 0) > minLengthOfChannel
peaks, _ = find_peaks(hist, distance=minPeaksDistance, plateau_size=15)
indices_with_larger_gaps = np.where(np.ediff1d(peaks) > gapWidth)[0]
locations_before_barcode = peaks[indices_with_larger_gaps]
locations_after_barcode = peaks[indices_with_larger_gaps + 1]
locations_barcode = np.rint(np.mean((locations_before_barcode,
locations_after_barcode), axis = 0)).astype('int')
num_barcodes = len(locations_barcode)
# there are 5 barcodes seen in the image
if num_barcodes == 5:
# count the number of channels before the first barcode and after the
# last barcode and include them upto numChannels channels
y_channels = []
# channels before first barcode
indices_before_first = np.where(peaks < locations_barcode[0])[0]
y_channels.extend(list(peaks[indices_before_first]))
for i in range(num_barcodes):
indices = np.where(np.logical_and(peaks > locations_barcode[i-1],
peaks < locations_barcode[i]))[0]
y_channels.extend(list(peaks[indices]))
# number of channels to count after the last
number_to_include = numChannels - len(indices_before_first)
indices_after_last = np.where(peaks > locations_barcode[-1])[0]
y_channels.extend(list(peaks[indices_after_last][:number_to_include]))
elif num_barcodes == 6:
y_channels = []
# count only the channels between barcodes and
# grab the (x, y) locations to cut,
# x will be the top of the channel, row number
# y will be the peak picked up in the histogram, between the barcodes
# count 21 channels after calculating
for i in range(num_barcodes):
indices = np.where(np.logical_and(peaks > locations_barcode[i-1],
peaks < locations_barcode[i]))[0]
#if len(indices) == 21:
# all good pick them up
y_channels.extend(list(peaks[indices]))
else:
# detection failure, since it is ambiguous skipp the position
y_channels = []
print(f"Detection failure, {num_barcodes} detected")
# locations of the barcode and locations of channels to cut.
return locations_barcode, y_channels
def apply(dirname, minLengthOfChannel = 200, minPeaksDistance = 25, gapWidth=48):
device = torch.device("cuda:1" if torch.cuda.is_available() else "cpu")
modelPath = Path('/home/pk/Documents/models/channels.pth')
pad = padTo32()
with torch.no_grad():
cellNetState = torch.load(modelPath)
if cellNetState['modelParameters']['netType'] == 'big':
cellSegNet = basicUnet(cellNetState['modelParameters']['transposeConv'])
elif cellNetState['modelParameters']['netType'] == 'small':
cellSegNet = smallerUnet(cellNetState['modelParameters']['transposeConv'])
cellSegNet.load_state_dict(cellNetState['model_state_dict'])
cellSegNet.to(device)
cellSegNet.eval()
start = time.time()
count = 0
for i in range(102, 841):
imgfilename = dirname + "Pos" + str(i) + '/phaseFast/img_000000000.tiff'
imgpath = Path(imgfilename)
image = io.imread(imgpath)
image = pad(image)
imgTensor = torch.from_numpy(image.astype('float32')).unsqueeze(0).unsqueeze(0).to(device)
imgTensor = (imgTensor - torch.mean(imgTensor))/torch.std(imgTensor)
out = torch.sigmoid(cellSegNet(imgTensor))
out_cpu = out.detach().cpu().numpy().squeeze(0).squeeze(0)
#print(imgTensor.shape)
out_cpu = gaussian(out_cpu, sigma = 3)
out_cpu = out_cpu > 0.5
out_cpu = remove_small_objects(out_cpu.astype('bool'), min_size=100)
locations_barcode, locations_channels = barcodesFromPeaks(out_cpu)
print(f"Barcodes: {len(locations_barcode)} --- no channels: {len(locations_channels)}")
if len(locations_channels) != 105:
count += 1
#plt.figure()
#plt.imshow(out_cpu)
#plt.plot(hist)
#plt.plot(peaks, hist[peaks], 'r*')
#plt.plot(peaks[indices_with_larger_gaps],
# hist[peaks[indices_with_larger_gaps]],'o', markersize=10,
# scalex=False, scaley=False, fillstyle='none')
#plt.show()
print(imgpath)
duration = time.time() - start
print(f"Duration: {duration/i}s")
print(f"Failed count: {count}")
del cellSegNet
del cellNetState
torch.cuda.empty_cache()
return None
# +
#one_img = apply('/home/pk/Documents/realtimeData/hetero40x/Pos103/phaseFast/')
# +
#one_img = apply("/mnt/sda1/Praneeth/EXP-20-BP0361 75k imaging 6ugml/dry run/")
# -
one_img = apply("/mnt/sda1/Praneeth/EXP-20-BP0361 75k imaging 6ugml/after loading/")
def barcodesFromPeaks(one_img, minLengthOfChannel= 200,
minPeaksDistance=25, gapWidth=48,
numChannels=21):
hist = np.sum(one_img, axis = 0) > minLengthOfChannel
peaks, _ = find_peaks(hist, distance=minPeaksDistance, plateau_size=15)
indices_with_larger_gaps = np.where(np.ediff1d(peaks) > gapWidth)[0]
locations_before_barcode = peaks[indices_with_larger_gaps]
locations_after_barcode = peaks[indices_with_larger_gaps + 1]
locations_barcode = np.rint(np.mean((locations_before_barcode,
locations_after_barcode), axis = 0)).astype('int')
num_barcodes = len(locations_barcode)
# there are 5 barcodes seen in the image
if num_barcodes == 5:
# count the number of channels before the first barcode and after the
# last barcode and include them upto numChannels channels
y_channels = []
# channels before first barcode
indices_before_first = np.where(peaks < locations_barcode[0])[0]
y_channels.extend(list(peaks[indices_before_first]))
for i in range(num_barcodes):
indices = np.where(np.logical_and(peaks > locations_barcode[i-1],
peaks < locations_barcode[i]))[0]
y_channels.extend(list(peaks[indices]))
# number of channels to count after the last
number_to_include = numChannels - len(indices_before_first)
indices_after_last = np.where(peaks > locations_barcode[-1])[0]
y_channels.extend(list(peaks[indices_after_last][:number_to_include]))
elif num_barcodes == 6:
y_channels = []
# count only the channels between barcodes and
# grab the (x, y) locations to cut,
# x will be the top of the channel, row number
# y will be the peak picked up in the histogram, between the barcodes
# count 21 channels after calculating
for i in range(num_barcodes):
indices = np.where(np.logical_and(peaks > locations_barcode[i-1],
peaks < locations_barcode[i]))[0]
#if len(indices) == 21:
# all good pick them up
y_channels.extend(list(peaks[indices]))
else:
# detection failure, since it is ambiguous skipp the position
print("Detection failure")
# locations of the barcode and locations of channels to cut.
return locations_barcode, y_channels
barcodes, y_channels = barcodesFromPeaks(one_img)
barcodes
len(y_channels)
# +
#one_img = apply("/home/pk/Documents/EXP-21-BY1006/therun/")
# -
plt.imshow(one_img)
minLengthOfChannel = 200
hist = np.sum(one_img, axis = 0) > minLengthOfChannel
plt.plot(hist)
plt.figure()
plt.imshow(one_img)
plt.show()
b = np.ones((10,))
b
np.insert(b, 0, 12)
3.29
image = io.imread(imgpath)
image = pad(image)
imgTensor = torch.from_numpy(image.astype('float32')).unsqueeze(0).unsqueeze(0).to(device)
imgTensor = (imgTensor - torch.mean(imgTensor))/torch.std(imgTensor)
out = torch.sigmoid(cellSegNet(imgTensor))
out_cpu = out.detach().cpu().numpy().squeeze(0).squeeze(0)
out_cpu
plt.imshow(out_cpu)
from skimage.measure import regionprops, label
from datetime import datetime
print(datetime.now())
props = regionprops(label(out_cpu > 0.9))
print(datetime.now())
removed_labels = []
labeled_img = label(out_cpu > 0.5)
plt.imshow(labeled_img, cmap='gnuplot2')
image = io.imread(path).astype('float32')
image.shape
image.dtype
plt.imshow(image)
image = (image - np.mean(image))/np.std(image)
from skimage import filters
edges = canny(image, sigma=7)
plt.imshow(edges)
from scipy.ndimage import binary_fill_holes, binary_erosion
image = binary_fill_holes(edges)
plt.imshow(image)
image = binary_erosion(image)
plt.imshow(image)
def detect_circles(in_img):
edges = canny(in_img, sigma=2)
hough_radii = np.arange(10, , 2)
hough_res = hough_circle(edges, hough_radii)
accums, cx, cy, radii = hough_circle_peaks(hough_res, hough_radii, total_num_peaks=300)
img1 = np.zeros(in_img.shape)
img1 = color.gray2rgb(img1)
for center_y, center_x, radius, (r, g, b, _) in zip(cy, cx, radii,
plt.cm.nipy_spectral(np.linspace(0,1, len(radii))) # color map
):
circy, circx = circle(center_y, center_x, radius)
img1[circy, circx] = (r*255, g*255, b*255)
return img1
img = detect_circles(image)
plt.imshow(img)
radii
cx
# +
cy
# -
| notebooks/Testing barcode extraction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import psycopg2
import sqlalchemy
import matplotlib.pyplot as plt
import scipy.stats as sts
from config import user, password
# source reference
#https://blog.panoply.io/connecting-jupyter-notebook-with-postgresql-for-python-data-analysis
from sqlalchemy import create_engine
# Postgres username, password, and database name
POSTGRES_ADDRESS = 'localhost'
POSTGRES_PORT = '5432'
POSTGRES_USERNAME = user
POSTGRES_PASSWORD = password
POSTGRES_DBNAME = '09-SQL-Homework'
# A long string that contains the necessary Postgres login information
postgres_str = ('postgresql://{username}:{password}@{ipaddress}:{port}/{dbname}'
.format(username=POSTGRES_USERNAME,password=<PASSWORD>,ipaddress=POSTGRES_ADDRESS,port=POSTGRES_PORT,dbname=POSTGRES_DBNAME))
# Create the connection
cnx = create_engine(postgres_str)
salaries_df = pd.read_sql_query('''SELECT salary FROM salaries;''', cnx)
# examine the sql return
salaries_df.head()
# +
# examine the salaries
# Get the measures of central tendency
count_salary = salaries_df['salary'].count()
print(f"The salary count is {count_salary}")
# Get the measures of central tendency
min_salary = np.min(salaries_df['salary'])
print(f"The min salary is {min_salary}")
# Get the max salary
max_salary = np.max(salaries_df['salary'])
print(f"The max salary is {max_salary}")
# Get the median salary
median_salary= np.median(salaries_df['salary'])
print(f"The median salary is {median_salary}")
mode_salary = sts.mode(salaries_df['salary'])
print(f"The mode salary is {mode_salary}")
# -
# examine the salaries
salaries_df['salary'].value_counts().sort_values(ascending=False)
# +
# Display the distribution of the salaries
# source - UCF Bootcamp stats training :)
# and to help me understand the result of normaltest
# https://datascienceplus.com/normality-tests-in-python/
# draw an average line in histogram
#https://stackoverflow.com/questions/16180946/drawing-average-line-in-histogram-matplotlib
#adjust the figure size
fig_size = plt.rcParams["figure.figsize"]
fig_size[0] = 20
fig_size[1] = 10
plt.rcParams["figure.figsize"] = fig_size
#set tick font size and plot font size
plt.tick_params(labelsize=15)
plt.rcParams.update({'font.size': 15})
#histogram, labels and mean line
plt.hist(salaries_df['salary'])
plt.title("Distribution of salaries")
plt.xlabel('Salaries')
plt.ylabel('Counts')
plt.axvline(salaries_df['salary'].mean(), color='r', linestyle='dashed', linewidth=2,label='Mean salary '+str(round(salaries_df['salary'].mean())))
plt.legend()
x_axis = np.arange(min_salary,max_salary,10000)
# Create the ticks for our bar chart's x axis
tick_locations = [value for value in x_axis]
plt.xticks(tick_locations, x_axis)
# save then show
plt.savefig("./Images/distribution_salaryp.png")
plt.show()
# normality test
# provide the p-value
stat, p = sts.normaltest(salaries_df['salary'].sample(50))
# interpret
alpha = 0.05
if p > alpha:
print(f'Sample looks Gaussian/normal (fail to reject NULL hypothesis p-value = {p}')
else:
print(f'Sample does not look Gaussian/normal (reject NULL hypothesis p-value = {p}')
# -
# query db to get the average salaries by title
avg_title_salaries_df = (pd.read_sql_query(
'''select t.title,round(avg(s.salary)) average_title_salary
from salaries s
join titles t on
t.emp_no = s.emp_no
group by t.title;
''', cnx)
)
# examine the saverafe salaries vy title
avg_title_salaries_df.head(7)
# +
# create a bar chart of average salaries by title
#Source reference on fig size
# https://stackabuse.com/python-data-visualization-with-matplotlib/
titles = avg_title_salaries_df['title']
x_axis = np.arange(len(titles))
y_axis = avg_title_salaries_df['average_title_salary']
#set tick font size and plot font size
plt.tick_params(labelsize=15)
plt.rcParams.update({'font.size': 15})
#set figue size
fig_size = plt.rcParams["figure.figsize"]
fig_size[0] = 20
fig_size[1] = 10
plt.rcParams["figure.figsize"] = fig_size
# Labels and titles
plt.title("Average Salary by Title")
plt.xlabel("Title")
plt.ylabel("Average Salary")
# Create a bar chart based upon the above data
plt.bar(x_axis, y_axis, color="b", align="center")
# Create the ticks for our bar chart's x axis
tick_locations = [value for value in x_axis]
plt.xticks(tick_locations, titles,rotation=45)
#save our chart
plt.tight_layout()
plt.savefig("./Images/bar_average_salary_by_title.png")
# -
# ## Epilogue
#
# This data is very skewed. Far too many employees have a salary of 40k .
#
# Evidence in hand, you march into your boss's office and present the visualization.
# With a sly grin, your boss thanks you for your work. On your way out of the office,
# you hear the words, "Search your ID number."
# You look down at your badge to see that your employee ID number is 499942.
#
# HAHAHAHA - My name is <NAME>!!!!!!
| query_db.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# Check nvcc version
# !nvcc -V
# Check GCC version
# !gcc --version
# +
# Check Pytorch installation
import torch, torchvision
print(torch.__version__, torch.cuda.is_available())
# Check MMSegmentation installation
import mmseg
print(mmseg.__version__)
# +
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:100% !important; }</style>"))
# Python ≥3.5 is required
import sys
assert sys.version_info >= (3, 5)
# Scikit-Learn ≥0.20 is required
import sklearn
assert sklearn.__version__ >= "0.20"
# Common imports
import numpy as np
import os
# to make this notebook's output stable across runs
np.random.seed(42)
# To plot pretty figures
# %matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rc('axes', labelsize=14)
mpl.rc('xtick', labelsize=12)
mpl.rc('ytick', labelsize=12)
plt.rc('font', size=12)
plt.rc('figure', figsize = (12, 5))
# Settings for the visualizations
import seaborn as sns
sns.set_style("whitegrid")
sns.set_context("notebook", font_scale=1, rc={"lines.linewidth": 2,'font.family': [u'times']})
import pandas as pd
pd.set_option('display.max_rows', 25)
pd.set_option('display.max_columns', 500)
pd.set_option('display.max_colwidth', 50)
# Ignore useless warnings (see SciPy issue #5998)
import warnings
warnings.filterwarnings(action="ignore", message="^internal gelsd")
# +
from torchvision import datasets, models, transforms
import torch.optim as optim
if torch.cuda.is_available():
torch.cuda.init()
print(torch.cuda.current_device())
print(torch.cuda.device_count())
print(torch.cuda.memory_allocated())
print(torch.cuda.memory_cached())
cuda = torch.device('cuda')
cuda0 = torch.device('cuda:0')
print(cuda0)
# -
torch.cuda.memory_summary(device=None, abbreviated=False)
torch.cuda.empty_cache()
# + pycharm={"is_executing": true}
from mmseg.apis import init_segmentor, inference_segmentor, show_result_pyplot
from mmseg.core.evaluation import get_palette
# -
import os.path as osp
import numpy as np
from PIL import Image
import skimage
from skimage import io
print(skimage.__version__)
import mmcv
classes = ['background', 'human']
palette = [[0,0,0], [255,0,0]]
class_weight = [1.0, 6.0] # 0.05 human pixels
# +
import os.path as osp
import mmcv
import numpy as np
from PIL import Image
from mmseg.datasets.builder import DATASETS
from mmseg.datasets.custom import CustomDataset
@DATASETS.register_module()
class ADE20KDataset01(CustomDataset):
"""ADE20K dataset.
In segmentation map annotation for ADE20K, 0 stands for background, which
is not included in 150 categories. ``reduce_zero_label`` is fixed to True.
The ``img_suffix`` is fixed to '.jpg' and ``seg_map_suffix`` is fixed to
'.png'.
"""
CLASSES = classes
PALETTE = palette
def __init__(self, **kwargs):
super(ADE20KDataset01, self).__init__(
img_suffix='.jpg',
seg_map_suffix='.png',
reduce_zero_label=False,
**kwargs)
assert osp.exists(self.img_dir)
def results2img(self, results, imgfile_prefix, to_label_id, indices=None):
if indices is None:
indices = list(range(len(self)))
mmcv.mkdir_or_exist(imgfile_prefix)
result_files = []
for result, idx in zip(results, indices):
filename = self.img_infos[idx]['filename']
basename = osp.splitext(osp.basename(filename))[0]
png_filename = osp.join(imgfile_prefix, f'{basename}.png')
output = Image.fromarray(result.astype(np.uint8))
output.save(png_filename)
result_files.append(png_filename)
return result_files
def format_results(self,
results,
imgfile_prefix,
to_label_id=True,
indices=None):
if indices is None:
indices = list(range(len(self)))
assert isinstance(results, list), 'results must be a list.'
assert isinstance(indices, list), 'indices must be a list.'
result_files = self.results2img(results, imgfile_prefix, to_label_id,
indices)
return result_files
# -
from mmcv import Config
cfg = Config.fromfile('../configs/bisenetv1/bisenetv1_r18-d32_lr5e-3_4x4_512x512_160k_coco-stuff164k.py')
data_root = '../data/ade/ADEChallengeData2016'
img_dir = 'images-binary'
ann_dir = 'binary-labels'
context_channels=(128, 256, 512)
spatial_channels=(64, 64, 64, 128)
decode_head_loss_weight = 0.3
auxiliary_head_loss_weight = decode_head_loss_weight*0.4
norm_config_type = 'BN'
from mmseg.apis import set_random_seed
# Since we use ony one GPU, BN is used instead of BN
cfg.norm_cfg = dict(type=norm_config_type, requires_grad=True) # Segmentation usually uses SyncBN
# model settings
cfg.model = dict(
type='EncoderDecoder', # Name of segmentor
pretrained=None,
backbone=dict(
type='BiSeNetV1', # The type of backbone.
in_channels=3,
context_channels=context_channels,
spatial_channels=spatial_channels,
out_indices=(0, 1, 2),
out_channels=256,
backbone_cfg=dict(
type='ResNet',
in_channels=3,
depth=18,
num_stages=4,
out_indices=(0, 1, 2, 3),
dilations=(1, 1, 1, 1),
strides=(1, 2, 2, 2),
norm_cfg=cfg.norm_cfg,
norm_eval=False,
style='pytorch',
contract_dilation=True),
norm_cfg=cfg.norm_cfg,
align_corners=False,
init_cfg=None),
decode_head=dict(
type='FCNHead',
in_channels=256,
in_index=0,
channels=256,
num_convs=1,
concat_input=False,
dropout_ratio=0.1,
num_classes=len(classes),
norm_cfg=cfg.norm_cfg,
align_corners=False,
loss_decode=[
dict(
type='CrossEntropyLoss',
loss_name='loss_ce',
use_sigmoid=False,
loss_weight=decode_head_loss_weight,
#class_weight=class_weight
),
dict(
type='DiceLoss',
loss_name='loss_dice',
use_sigmoid=False,
loss_weight=decode_head_loss_weight,
class_weight=class_weight
)
]
),
auxiliary_head=[
dict(
type='FCNHead', # Type of auxiliary head. Please refer to mmseg/models/decode_heads for available options.
in_channels=128, # Input channel of auxiliary head.
channels=64, # The intermediate channels of decode head.
num_convs=1, # Number of convs in FCNHead. It is usually 1 in auxiliary head.
num_classes=len(classes), # Number of segmentation class. Usually 19 for cityscapes, 21 for VOC, 150 for ADE20k.
in_index=1, # The index of feature map to select.
norm_cfg=cfg.norm_cfg, # The configuration of norm layer.
concat_input=False, # The configuration of norm layer.
align_corners=False, # The align_corners argument for resize in decoding.
loss_decode=[
dict( # Config of loss function for the decode_head.
type='CrossEntropyLoss', # Type of loss used for segmentation.
loss_name='loss_ce',
use_sigmoid=False, # Whether use sigmoid activation for segmentation.
loss_weight=auxiliary_head_loss_weight, # Loss weight of auxiliary head, which is usually 0.4 of decode head.
#class_weight=class_weight
),
dict( # Config of loss function for the decode_head.
type='DiceLoss', # Type of loss used for segmentation.
loss_name='loss_dice',
use_sigmoid=False, # Whether use sigmoid activation for segmentation.
loss_weight=auxiliary_head_loss_weight, # Loss weight of auxiliary head, which is usually 0.4 of decode head.
class_weight=class_weight
)
]
),
dict(
type='FCNHead',
in_channels=128,
channels=64,
num_convs=1,
num_classes=len(classes),
in_index=2,
norm_cfg=cfg.norm_cfg,
concat_input=False,
align_corners=False,
loss_decode=[
dict( # Config of loss function for the decode_head.
type='CrossEntropyLoss', # Type of loss used for segmentation.
loss_name='loss_ce',
use_sigmoid=False, # Whether use sigmoid activation for segmentation.
loss_weight=auxiliary_head_loss_weight, # Loss weight of auxiliary head, which is usually 0.4 of decode head.
#class_weight=class_weight
),
dict( # Config of loss function for the decode_head.
type='DiceLoss', # Type of loss used for segmentation.
loss_name='loss_dice',
use_sigmoid=False, # Whether use sigmoid activation for segmentation.
loss_weight=auxiliary_head_loss_weight, # Loss weight of auxiliary head, which is usually 0.4 of decode head.
class_weight=class_weight
)
]
)
],
# model training and testing settings
train_cfg=dict(), # train_cfg is just a place holder for now.
test_cfg=dict(mode='whole') # The test mode, options are 'whole' and 'sliding'. 'whole': whole image fully-convolutional test. 'sliding': sliding crop window on the image.
)
# +
############### start dataset
cfg.dataset_type = 'ADE20KDataset01' # Dataset type, this will be used to define the dataset.
cfg.data_root = data_root # Root path of data.
cfg.img_norm_cfg = dict( # Image normalization config to normalize the input images.
mean=[123.675, 116.28, 103.53], # Mean values used to pre-training the pre-trained backbone models.
std=[58.395, 57.12, 57.375], # Standard variance used to pre-training the pre-trained backbone models.
to_rgb=True) # The channel orders of image used to pre-training the pre-trained backbone models.
cfg.crop_size = (512, 512) # The crop size during training.
cfg.train_pipeline = [ # Training pipeline.
dict(type='LoadImageFromFile'), # First pipeline to load images from file path.
dict(type='LoadAnnotations', reduce_zero_label=False), # Second pipeline to load annotations for current image.
dict(type='Resize', # Augmentation pipeline that resize the images and their annotations.
img_scale=(2048, 512), # The largest scale of image.
ratio_range=(0.5, 2.0)), # The augmented scale range as ratio.
dict(type='RandomCrop', # Augmentation pipeline that randomly crop a patch from current image.
crop_size=cfg.crop_size, # The crop size of patch.
cat_max_ratio=0.75), # The max area ratio that could be occupied by single category.
dict(
type='RandomFlip', # Augmentation pipeline that flip the images and their annotations
flip_ratio=0.5), # The ratio or probability to flip
dict(type='PhotoMetricDistortion'), # Augmentation pipeline that distort current image with several photo metric methods.
dict(
type='Normalize', # Augmentation pipeline that normalize the input images
**cfg.img_norm_cfg),
dict(type='Pad', # Augmentation pipeline that pad the image to specified size.
size=cfg.crop_size, # The output size of padding.
pad_val=0, # The padding value for image.
seg_pad_val=255), # The padding value of 'gt_semantic_seg'.
dict(type='DefaultFormatBundle'), # Default format bundle to gather data in the pipeline
dict(type='Collect', # Pipeline that decides which keys in the data should be passed to the segmentor
keys=['img', 'gt_semantic_seg'])
]
cfg.test_pipeline = [
dict(type='LoadImageFromFile'), # First pipeline to load images from file path
dict(
type='MultiScaleFlipAug', # An encapsulation that encapsulates the test time augmentations
img_scale=(2048, 512), # Decides the largest scale for testing, used for the Resize pipeline
flip=False, # Whether to flip images during testing
transforms=[
dict(type='Resize', # Use resize augmentation
keep_ratio=True), # Whether to keep the ratio between height and width, the img_scale set here will be suppressed by the img_scale set above.
dict(type='RandomFlip'), # Thought RandomFlip is added in pipeline, it is not used when flip=False
dict(
type='Normalize', # Normalization config, the values are from img_norm_cfg
**cfg.img_norm_cfg),
dict(type='ImageToTensor', # Convert image to tensor
keys=['img']),
dict(type='Collect', # Collect pipeline that collect necessary keys for testing.
keys=['img'])
])
]
cfg.data = dict(
samples_per_gpu=16,
workers_per_gpu=16,
train=dict(
type=cfg.dataset_type,
data_root=cfg.data_root,
img_dir=f'{img_dir}/training',
ann_dir=f'{ann_dir}/training',
pipeline=cfg.train_pipeline),
val=dict(
type=cfg.dataset_type,
data_root=cfg.data_root,
img_dir=f'{img_dir}/validation',
ann_dir=f'{ann_dir}/validation',
pipeline=cfg.test_pipeline),
test=dict(
type=cfg.dataset_type,
data_root=cfg.data_root,
img_dir=f'{img_dir}/validation',
ann_dir=f'{ann_dir}/validation',
pipeline=cfg.test_pipeline))
############ end dataset
# +
from mmseg.datasets import build_dataset
from mmseg.models import build_segmentor
from mmseg.apis import train_segmentor
# Build the dataset
datasets = [build_dataset(cfg.data.train)]
#print(datasets[0].CLASSES)
print(len(datasets[0].CLASSES))
# -
# modify num classes of the model in decode/auxiliary head
cfg.model.decode_head.num_classes = len(datasets[0].CLASSES)
cfg.model.auxiliary_head[0].num_classes = len(datasets[0].CLASSES)
cfg.model.auxiliary_head[1].num_classes = len(datasets[0].CLASSES)
# +
# Set up working dir to save files and logs.
cfg.work_dir = './work_dirs/testv1.Training_Tricks.160000iter'
############ start default runtime
cfg.log_config = dict( # config to register logger hook
interval=50, # Interval to print the log
hooks=[
dict(type='TensorboardLoggerHook'), # The Tensorboard logger is also supported
dict(type='TextLoggerHook', by_epoch=False)
])
cfg.dist_params = dict(backend='nccl') # Parameters to setup distributed training, the port can also be set.
cfg.log_level = 'INFO' # The level of logging.
cfg.load_from = './work_dirs/testv1.Training_Tricks.160000iter/iter_160000.pth' # load models as a pre-trained model from a given path. This will not resume training.
cfg.resume_from = './work_dirs/testv1.Training_Tricks.160000iter/iter_160000.pth' # Resume checkpoints from a given path, the training will be resumed from the iteration when the checkpoint's is saved.
cfg.workflow = [('train', 1)] # Workflow for runner. [('train', 1)] means there is only one workflow and the workflow named 'train' is executed once. The workflow trains the model by 40000 iterations according to the `runner.max_iters`.
cfg.cudnn_benchmark = True # Whether use cudnn_benchmark to speed up, which is fast for fixed input size.
############ end default runtime
############ start schedule
cfg.optimizer = dict( # Config used to build optimizer, support all the optimizers in PyTorch whose arguments are also the same as those in PyTorch
type='SGD', # Type of optimizers, refer to https://github.com/open-mmlab/mmcv/blob/master/mmcv/runner/optimizer/default_constructor.py#L13 for more details
lr=0.005, # Learning rate of optimizers, see detail usages of the parameters in the documentation of PyTorch
momentum=0.9, # Momentum
weight_decay=0.0005,
paramwise_cfg=dict(
custom_keys=dict(head=dict(lr_mult=10.0)))
)
cfg.optimizer_config = dict() # Config used to build the optimizer hook, refer to https://github.com/open-mmlab/mmcv/blob/master/mmcv/runner/hooks/optimizer.py#L8 for implementation details.
lr_config = dict(
policy='poly', # The policy of scheduler, also support Step, CosineAnnealing, Cyclic, etc. Refer to details of supported LrUpdater from https://github.com/open-mmlab/mmcv/blob/master/mmcv/runner/hooks/lr_updater.py#L9.
power=0.9, # The power of polynomial decay.
min_lr=0.00001, # The minimum learning rate to stable the training.
by_epoch=False,
warmup='linear',
warmup_iters=1000
) # Whether count by epoch or not.
cfg.runner=dict(
type='IterBasedRunner', # Type of runner to use (i.e. IterBasedRunner or EpochBasedRunner)
max_iters=160000) # Total number of iterations. For EpochBasedRunner use `max_epochs`
cfg.checkpoint_config = dict( # Config to set the checkpoint hook, Refer to https://github.com/open-mmlab/mmcv/blob/master/mmcv/runner/hooks/checkpoint.py for implementation.
by_epoch=False, # Whether count by epoch or not.
interval=10000,
meta=dict(
CLASSES=classes,
PALETTE=palette
)
)
cfg.evaluation = dict( # The config to build the evaluation hook. Please refer to mmseg/core/evaluation/eval_hook.py for details.
interval=5000, # The interval of evaluation.
metric='mIoU',
pre_eval=True) # The evaluation metric.
############ end schedule
# +
# Set seed to facitate reproducing the result
cfg.seed = 0
set_random_seed(0, deterministic=False)
cfg.gpu_ids = range(1)
# Let's have a look at the final config used for training
print(f'Config:\n{cfg.pretty_text}')
# -
# Build the detector
model = build_segmentor(cfg.model, train_cfg=cfg.get('train_cfg'), test_cfg=cfg.get('test_cfg'))
# +
# Add an attribute for visualization convenience
model.CLASSES = datasets[0].CLASSES
model.PALETTE = datasets[0].PALETTE
# Create work_dir
mmcv.mkdir_or_exist(osp.abspath(cfg.work_dir))
# -
train_segmentor(model, datasets, cfg, distributed=False, validate=True, meta=dict())
# human IoU above 70%
img = mmcv.imread('../data/ade/ADEChallengeData2016/images-binary/validation/ADE_val_00000011.jpg')
plt.imshow(img)
# +
img = mmcv.imread('../data/ade/ADEChallengeData2016/images-binary/validation/ADE_val_00000011.jpg')
model.cfg = cfg
result = inference_segmentor(model, img)
# -
np.asarray(result).max()
plt.figure(figsize=(8, 6))
show_result_pyplot(model, img, result, palette)
img = mmcv.imread('../data/ade/ADEChallengeData2016/binary-labels/validation/ADE_val_00000011.png')
img[img == 1] = 255
plt.imshow(img, cmap="gray")
img.max()
img.min()
# print('show video')
# test a video and show the results
video = mmcv.VideoReader('../data/ramp.mp4')
import time
video[1].shape
x = 100
# +
# %%time
start = time.time()
for frame in video[:x]: # time - how many frame per sec - metrics acc - IoU
result = inference_segmentor(model, frame)
#model.show_result(frame, result)
end = time.time()
# -
from humanfriendly import format_timespan
aux0 = (end-start)
print("Time:", format_timespan(aux0))
aux1 = aux0/x
print("Time/frame:", format_timespan(aux1))
print("FPS: ", 1/aux1)
# display the training loss changes
| comparisons/trash/inference_ADE_2_BiSeNetv1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 2020년 6월 26일 금요일
# ### BaekJoon - 11055 : 가장 큰 증가 부분 수열 (Python)
# ### 문제 : https://www.acmicpc.net/problem/11055
# ### 블로그 : https://somjang.tistory.com/entry/BaekJoon-11055%EB%B2%88-%EA%B0%80%EC%9E%A5-%ED%81%B0-%EC%A6%9D%EA%B0%80-%EB%B6%80%EB%B6%84-%EC%88%98%EC%97%B4-Python
# ### 첫번째 시도
# +
inputNum = int(input())
inputNums = input()
inputNums = inputNums.split()
inputNums = [int(num) for num in inputNums]
nc = [0] * (inputNum)
maxNum = 0
for i in range(0, inputNum):
nc[i] = nc[i] + inputNums[i]
for j in range(0, i):
if inputNums[j] < inputNums[i] and nc[i] < nc[j] + inputNums[i]:
nc[i] = nc[j] + inputNums[i]
if maxNum < nc[i]:
maxNum = nc[i]
print(max(nc))
| DAY 101 ~ 200/DAY141_[BaekJoon] 가장 큰 증가 부분 수열 (Python).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # About this notebook
# ### The machine learning model trained in this notebook forecasts the daily # of available hospital beds in the Dallas-Forth Worth region based on past data.
#
# The data was obtained from [Combined Hospital Data over Time by Trauma Service Area (TSA)](https://dshs.texas.gov/coronavirus/additionaldata.aspx). **When given a list of future dates as input, the model forecasts the expected # of hospital beds that will be available on those days and also provides a plausible range of available beds.**
#
# ### There are 2 parts to this notebook
# 1. **[Part 1:](https://render.githubusercontent.com/view/ipynb?commit=e0b9c646ae851237e40aee17a1da21e454f4cd20&enc_url=68747470733a2f2f7261772e67697468756275736572636f6e74656e742e636f6d2f494a2d417070732f44616c6c61732d446174612d4368616c6c656e67652f653062396336343661653835313233376534306165653137613164613231653435346634636432302f444657253230417661696c61626c65253230486f73706974616c253230426564253230466f726563617374696e672e6970796e623f746f6b656e3d414d4e5a57323551574447435550324d4f4d575a5a5843374e564e5157&nwo=IJ-Apps%2FDallas-Data-Challenge&path=DFW+Available+Hospital+Bed+Forecasting.ipynb&repository_id=298444278&repository_type=Repository#Part-1)** Training the model on dates before September, then evaluating its accuracy at forecasting the # of hospital beds available on September dates *(for which data is available)*.
# 2. [**Part 2:**](https://render.githubusercontent.com/view/ipynb?commit=2411b5458460c2e529e3bc7fb68196f38fce176c&enc_url=68747470733a2f2f7261772e67697468756275736572636f6e74656e742e636f6d2f494a2d417070732f44616c6c61732d446174612d4368616c6c656e67652f323431316235343538343630633265353239653362633766623638313936663338666365313736632f444657253230417661696c61626c65253230486f73706974616c253230426564253230466f726563617374696e672e6970796e623f746f6b656e3d414d4e5a5732354b4b42354d36584651365054584b3553374e564e5336&nwo=IJ-Apps%2FDallas-Data-Challenge&path=DFW+Available+Hospital+Bed+Forecasting.ipynb&repository_id=298444278&repository_type=Repository#Part-2---The-Future) Training the model on all available data *(including September)* and **forecasting the # of beds available for future dates: the next 30 days.**
# # Part 1
import pandas as pd
from datetime import datetime
from pandas import DataFrame
from pandas import to_datetime
from pandas import read_csv
from pandas import to_datetime
import datetime
import os
# fbprophet is Facebook's open source library for time series prediction
from fbprophet import Prophet
# ## Processing the Data
# The below data was extracted from the Texas DSHS Coronavirus website:
# [Combined Hospital Data over Time by Trauma Service Area (TSA)](https://dshs.texas.gov/coronavirus/additionaldata.aspx)
#
# - In the code below, I am reading in the data from a CSV. It isn't ready in an immediately-readable format to be stored in the list so I have to split it by commas, read each number as an int, and store it in a list.
# - I also create a dataframe, `df` to store each date from April 12 and the # of hospital beds available on that day.
# - I am training my model on these dates before September so that after it is trained, I can make it forecast the # of available beds on each day in September *(data it hasn't seen before)*, compare it to the actual # of beds, and calculate the accuracy of my model.
# +
# read in the data from the available_beds.csv file for the # of available hospital beds in DFW
# from April 12 to Sept 22 (as of the time of coding this)
data = pd.read_csv(os.path.join("available_beds.csv"), engine='python')
data = data.transpose() # switch rows and columns
# Given a string, this function returns a number. The reason for this is because when splitting the data
#. by spaces, it contains commas and new-line characters.
def getNumberFromString(number):
s = ""
for pos in range(0, len(bed_number)):
if(not bed_number[pos].isdigit()):
break
s += bed_number[pos]
return s
# -
# x is a list of the number of available hospital beds each day in DFW.
x = []
# The "Dallas/Ft.Worth" can be switched out with any city in the dataset
for bed_number in str(data["Dallas/Ft. Worth"]).split(" "):
if(not bed_number[:1].isdigit()):
continue
x.append(int(getNumberFromString(bed_number)))
# +
# Generate dates from April 12th to last date in the dataset
end_date = pd.to_datetime("2020-04-12") + pd.DateOffset(days=len(x) - 1)
dates = pd.date_range(start="2020-04-12",end=end_date)
# List comprehension that crops out the hour, minutes, and seconds in the time
dates = [str(i)[:10] for i in dates]
# This code slices the last 22 days off the training data and saves it for later for the model to forecast
# At the time of coding this, the testing data spans from September 1st to 22nd
testD = dates[-22:]
testX = x[-22:]
dates = dates[:-22]
x = x[:-22]
print("# of Dates:", len(dates), "\nFrom", dates[0], "to", dates[len(dates) - 1])
print("# of Dates to Forecast:", len(testD), "\nFrom", testD[0], "to", testD[len(testD) - 1])
# -
# Create a pandas DataFrame object from the list of dates and hospital beds
df = pd.DataFrame({'Dates' : dates, 'Beds' : x})
df
# ## Plotting the data
# The code below plots the data and saves the graph in a high-quality 1200x1200 png
# +
# Plot the data
import matplotlib.pyplot as plt
df.plot()
plt.title("# of Hospital Beds Available in DFW", fontsize=18)
plt.xlabel('Day since April 13',fontsize=15)
plt.ylabel('# of Beds Available',fontsize=18)
# plt.savefig('Hospital Beds Available.png', dpi=1200)
plt.show()
# -
# ## Training the Model
# Now that I have the data, I train the Prophet model on the data *(all dates before September)*.
#
# In the second block, I create a `DataFrame` from the list of test dates I had set aside in the beginning. My model will predict the # of beds available on these test dates - *(September dates)* - based on the historical data it has been fed.
# prophet requires the column names to be "ds" and "y"
df.columns = ['ds', 'y']
# make the model
model = Prophet()
# train the model on the dataset (in this case dataframe df)
model.fit(df)
# Generate a list of future dates for the model to predict
future = DataFrame(testD)
future.columns = ['ds']
# Printing the September dates to forecast
future
# ## Testing the Model
# In this step, I test the model and use it to predict the # of available hospital beds for the dates I generated above.
#
# I then plot a graph showing the # of available beds since April 13th and a blue line that shows the # of beds my model predicted would be available on future dates.
# +
# forecast contains the predictions for the dates we provided
forecast = model.predict(future)
# This prints the date, expected # of beds available, and the lower and upper bounds of the forecasted interval
# print(forecast[['ds', 'yhat', 'yhat_lower', 'yhat_upper']])
# plot forecast
model.plot(forecast)
plt.title("Forecasted # of Available Hospital Beds in DFW for September", fontsize=17)
plt.xlabel('Dates',fontsize=17)
plt.ylabel('# of Beds Available',fontsize=23)
# plt.savefig('September Forecast.png', dpi=1200)
plt.show()
# -
# ## Evaluating the Model
# After having trained the model and having it predict the daily # of available beds from September 1st to 22nd, in this step I will evaluate the model's accuracy.
#
# Among the useful information the model provides is the **expected # of available beds** and **an expected range of available beds** *(interval)*.
#
# Below are some of the metrics I am examining
#
# - Percent Error
# - Mean Absolute Error (MAE)
# - Percentage of Dates within predicted *interval*
#
# ### 1. Visualizing Forecasts vs Actual # of Available Beds
# +
intervals = [] # Intervals of bed predictions
predicted = [] # Predicted # of available beds
differences = [] # Difference (actual - predicted)
# For each date that the model had to forecast, I get the predicted number of beds, and the confidence interval (or range of beds for that day)
for count, (pred, lower, upper) in enumerate(zip(forecast['yhat'], forecast['yhat_lower'], forecast['yhat_upper'])):
intervals.append("{:.0f} - {:.0f}".format(lower, upper))
differences.append(testX[count] - int(round(pred)))
predicted.append(int(round(pred)))
# data frame of the summary between actual # and forecasted # of available beds
summary = pd.DataFrame({'Dates' : testD, 'Actual': testX, 'Predicted' : predicted,
'Difference' : differences, 'Intervals' : intervals})
# Displaying the dataframe
print(" # of Beds Forecasted vs Actual")
summary
# -
# ### 2. Percent Error & Percent of accurate forecasted intervals
# After getting the important data I need (forecasts and forecast range), I calculate the percent error of the forecast for each date in September and average them.
#
# I also count the # of intervals that correctly captured the actual # of available beds on that day. ***Example:*** *For September 21st, the model forecasted that between 1504 to 2071 beds would be available. The actual number was 1782 which lies inside the interval, near the center, so the model's forecast contained the actual number.*
# +
error = []
absolute_error = 0 # Mean Absolute Error
contains = [] # True or False based on whether a forecasted interval contains the actual # of available beds
average = 0
for i in range (0, len(predicted)):
# Calculate percent error for each date. testX is a list of the actual # of beds
average += (testX[i] - predicted[i])/testX[i] * 100
error.append(round(((testX[i] - predicted[i])/testX[i] * 100)))
absolute_error += abs(testX[i] - predicted[i])
# Determining whether forecast interval contains actual # of beds
lower_bound = forecast['yhat_lower'][i]
upper_bound = forecast['yhat_upper'][i]
contains.append((lower_bound <= testX[i] and testX[i] <= upper_bound))
# calculate average percent error
average = round(average/len(error),1)
# calculate mean absolute error
absolute_error = round(absolute_error / len(error))
# Count # of forecasted intervals that contained the actual # of available beds
numberCorrect = sum(interval for interval in contains)
percentCorrect = int((numberCorrect * 100)/len(contains))
print("---------------------------------")
print("AVERAGE PERCENT ERROR:", average, "%")
print("MEAN ABSOLUTE ERROR:", absolute_error, "beds\n")
print("On {0}% of the days ({1} out of 22 days), the model's forecast interval of the # of available beds contained the actual # of beds available on that day.".format(percentCorrect, numberCorrect))
print("---------------------------------")
results = pd.DataFrame({'Dates' : testD, 'Percent Error': [str(e) + "%" for e in error],
'Accurate Forecast Interval' : contains})
results
# -
# ## Results/Conclusion
# - The model had an **average percent of error of about 10%.** Considering that the # of available beds in April was very different from how it is now, these are good results.
# - Additionally, when predicting the # of beds available for the first 22 days in September, the model also provided an expected range of available beds. For **17 out of the 22 days (77%)**, this expected range accurately captured the correct # of available beds, meaning that the model can forecast the ballpark of number of beds available
# - **The model's performance will improve as more time passes by and more data is available.**
# # Part 2 - The Future
# ### Forecasting future dates - the next 31 days
# In this part, I will train a model to forecast the daily # of available hospital beds in Dallas/Fort Worth for dates that have not yet arrived *(late September and October)*.
#
# This time, I will include data from September when training the model.
# +
x = []
# Reading & storing the # of available beds in DFW
for bed_number in str(data["Dallas/Ft. Worth"]).split(" "):
if(not bed_number[:1].isdigit()):
continue
x.append(int(getNumberFromString(bed_number)))
# Generate dates from April 12th to last date in the dataset
end_date = pd.to_datetime("2020-04-12") + pd.DateOffset(days=len(x) - 1)
dates = pd.date_range(start="2020-04-12",end=end_date)
dates = [str(i)[:10] for i in dates]
print("# of Dates:", len(dates), "\nFrom", dates[0], "to", dates[len(dates) - 1])
# -
# Create a pandas DataFrame object from the list of dates and beds
df = pd.DataFrame({'Dates' : dates, 'Beds' : x})
# prophet requires the column names to be "ds" and "y"
df.columns = ['ds', 'y']
# make the model
model = Prophet()
# train the model on the dataset (in this case dataframe df)
model.fit(df)
# ### Forecasting future dates
# The code below forecasts the # of available beds from **September 23rd to October 23rd**
# Generating a list of future dates for the model to forecast
future_dates = pd.date_range(start="2020-09-23",end="2020-10-23")
future = DataFrame(future_dates)
future.columns = ['ds']
# forecast contains the predictions for the dates we provided
forecast = model.predict(future)
# +
alternate = forecast[:]
model.plot(alternate)
plt.title("Forecasted # of Available Hospital Beds in DFW", fontsize=17)
plt.xlabel('Dates',fontsize=17)
plt.ylabel('# of Beds Available',fontsize=23)
# plt.savefig('Future Forecast.png', dpi=1200)
plt.show()
# -
# ### Zoomed-in graph
# The code below creates a graph that zooms in on the graph above. **It shows the last 31 days where the # of available beds was known next to the forecasted # of beds in Dallas from September 23rd to October 23rd.**
toplot = []
before = 31
for i in x[-before:]:
toplot.append(i)
for i in alternate['yhat']:
toplot.append(int(i))
# +
# total available beds
import matplotlib.patches as mpatches
from matplotlib.pyplot import figure
alternate
plt.title("Forecasted # of Available Hospital Beds in DFW", fontsize=17)
plt.xlabel('Days since September 3',fontsize=17)
plt.ylabel('# of Beds Available',fontsize=18)
# Generate the before and after lists for x values
x1 = [i for i in range(0,before)]
x2 = [i for i in range(before, len(alternate) + before)]
# Create the legend
blue_patch = mpatches.Patch(color='blue', label='Last known # of beds')
red_patch = mpatches.Patch(color='red', label='Forecasted # of beds')
plt.legend(handles=[blue_patch, red_patch])
plt.plot(x1, toplot[:before], color='blue')
plt.plot(x2, toplot[before:], color='red')
# plt.savefig('Future Forecast Zoomed.png', dpi=1200)
plt.show()
# -
# # End of Notebook
| DFW Available Hospital Bed Forecasting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="HLU-f5CAapbg" executionInfo={"status": "ok", "timestamp": 1637316697249, "user_tz": -330, "elapsed": 411, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjc2nHKs9TxBjI8Rdr48x2ktCUgrltQi-Ca-pdomA=s64", "userId": "12752491426441843216"}}
import csv
import fastai
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from pandas.plotting import scatter_matrix
from sklearn import preprocessing
from sklearn.preprocessing import MinMaxScaler
from datetime import datetime
from fastai.tabular import add_datepart
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_squared_error, r2_score, mean_absolute_error, accuracy_score
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="s9c0w6lPazUF" executionInfo={"status": "ok", "timestamp": 1637316700035, "user_tz": -330, "elapsed": 636, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjc2nHKs9TxBjI8Rdr48x2ktCUgrltQi-Ca-pdomA=s64", "userId": "12752491426441843216"}} outputId="248ade10-1491-4384-a869-4b7eb006ea0a"
df = pd.read_csv('/content/drive/MyDrive/MLproject/DRREDDY.csv')
df.describe()
# + colab={"base_uri": "https://localhost:8080/", "height": 311} id="9bEUSmdqGL5J" executionInfo={"status": "ok", "timestamp": 1637316704776, "user_tz": -330, "elapsed": 2594, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjc2nHKs9TxBjI8Rdr48x2ktCUgrltQi-Ca-pdomA=s64", "userId": "12752491426441843216"}} outputId="d6383cab-9c51-4fba-fae7-a63365ffc515"
#Visualization
#histogram
fig = plt.figure(figsize =(20,8))
df.hist()
fig.show()
# + id="lCVmgYwib6-r" executionInfo={"status": "ok", "timestamp": 1637316706598, "user_tz": -330, "elapsed": 447, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjc2nHKs9TxBjI8Rdr48x2ktCUgrltQi-Ca-pdomA=s64", "userId": "12752491426441843216"}}
df['Date'] = pd.to_datetime(df.Date,format='%Y-%m-%d')
df.index = df['Date']
#sorting
data = df.sort_index(ascending=True, axis=0)
#creating a separate dataset
new_data = pd.DataFrame(index=range(0,len(df)),columns=['Date', 'Close','Open','High','Volume'])
new_data['Date'] = df['Date'].values
new_data['Close'] = df['Close'].values
new_data['Open'] = df['Open'].values
new_data['High'] = df['High'].values
new_data['Volume'] = df['Volume'].values
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="4hKlqHmdcD23" executionInfo={"status": "ok", "timestamp": 1637306374523, "user_tz": -330, "elapsed": 1271, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjc2nHKs9TxBjI8Rdr48x2ktCUgrltQi-Ca-pdomA=s64", "userId": "12752491426441843216"}} outputId="5e1df670-44ee-4664-8ff9-1bf66f22a297"
new_data
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="u9vtre72GARC" executionInfo={"status": "ok", "timestamp": 1637316721129, "user_tz": -330, "elapsed": 11316, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjc2nHKs9TxBjI8Rdr48x2ktCUgrltQi-Ca-pdomA=s64", "userId": "12752491426441843216"}} outputId="c52e69ff-fa29-41f2-8c9f-b3f0a02e7747"
#Visualization
#histogram
#data Close, High, Open
plt.figure(figsize=(16,8))
plt.plot(df['Date'],df['Close'], label='Close Price history')
plt.plot(df['Date'],df['Open'], label='open Price history')
plt.plot(df['Date'],df['High'], label='High Price history')
plt.legend()
plt.show()
plt.figure(figsize=(16,8))
plt.plot(df['Date'],df['Volume'], label='Volume history')
plt.legend()
plt.show()
#subplots
plt.figure(figsize=(16,9))
fig, ax = plt.subplots(figsize=(10,10))
dataplot = sns.heatmap(df.corr(), cmap="YlGnBu", annot=True)
# Horizontal Bar Plot
df1 = df
df1.index=df1['Date']
df1['year'] = df1.index.year
fig = plt.figure(figsize =(20,8))
plt.bar(df1['year'],df1['Close'])
plt.show()
# + id="DL_dyci5eg8G" executionInfo={"status": "ok", "timestamp": 1637306390603, "user_tz": -330, "elapsed": 675, "user": {"displayName": "te<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjc2nHKs9TxBjI8Rdr48x2ktCUgrltQi-Ca-pdomA=s64", "userId": "12752491426441843216"}}
import fastai
# + colab={"base_uri": "https://localhost:8080/"} id="9P-WUHglcDoP" executionInfo={"status": "ok", "timestamp": 1637316721130, "user_tz": -330, "elapsed": 7, "user": {"displayName": "tej<NAME>uru", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjc2nHKs9TxBjI8Rdr48x2ktCUgrltQi-Ca-pdomA=s64", "userId": "12752491426441843216"}} outputId="920a5a04-f2d4-4526-a7ed-08e385864a31"
from fastai.tabular import add_datepart
add_datepart(new_data, 'Date')
new_data.drop('Elapsed', axis=1, inplace=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="kwPrJi3ALDf1" executionInfo={"status": "ok", "timestamp": 1637306396444, "user_tz": -330, "elapsed": 594, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjc2nHKs9TxBjI8Rdr48x2ktCUgrltQi-Ca-pdomA=s64", "userId": "12752491426441843216"}} outputId="3ff941b2-e049-4af4-95a3-b458f5f1650b"
new_data
# + colab={"base_uri": "https://localhost:8080/"} id="8w9UUpSyFa0y" executionInfo={"status": "ok", "timestamp": 1637316731132, "user_tz": -330, "elapsed": 381, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjc2nHKs9TxBjI8Rdr48x2ktCUgrltQi-Ca-pdomA=s64", "userId": "12752491426441843216"}} outputId="a18e68ab-951f-4f50-fc60-51d88464c7c9"
new_data.shape
# + id="Pss1VD8Usyd9" executionInfo={"status": "ok", "timestamp": 1637316733290, "user_tz": -330, "elapsed": 424, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjc2nHKs9TxBjI8Rdr48x2ktCUgrltQi-Ca-pdomA=s64", "userId": "12752491426441843216"}}
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler(feature_range=(0, 1))
scal = scaler.fit_transform(new_data)
# + id="vGUktGsaLNRt" executionInfo={"status": "ok", "timestamp": 1637316735519, "user_tz": -330, "elapsed": 503, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjc2nHKs9TxBjI8Rdr48x2ktCUgrltQi-Ca-pdomA=s64", "userId": "12752491426441843216"}}
train = scal[:3714]
test = scal[3714:]
# + id="W4fbGKExGgSX" executionInfo={"status": "ok", "timestamp": 1637316853635, "user_tz": -330, "elapsed": 375, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjc2nHKs9TxBjI8Rdr48x2ktCUgrltQi-Ca-pdomA=s64", "userId": "12752491426441843216"}}
x_train = train[:,1:]
y_train = train[:,0]
x_test = test[:, 1:]
y_test = test[:, 0]
# + [markdown] id="vRvfoIJPIxbv"
# #Linear Regression
# + colab={"base_uri": "https://localhost:8080/", "height": 568} id="P_B89kjiD5xz" executionInfo={"status": "ok", "timestamp": 1637317202117, "user_tz": -330, "elapsed": 1648, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjc2nHKs9TxBjI8Rdr48x2ktCUgrltQi-Ca-pdomA=s64", "userId": "12752491426441843216"}} outputId="ef2b9a67-6e55-4a75-c81e-7ba201ea8ade"
class LinearRegression:
def __init__(self,lr=0.00005,itr=1500):
self.lr = lr
self.itr = itr
def fit(self,X,y):
self.m, self.n = X.shape
self.X = np.append(np.ones((self.m, 1)), X, axis=1)
self.y = y
# h(x) = theta . X
self.theta = np.zeros(self.n + 1)
for i in range(self.itr):
self.gradient_descent()
return self
def gradient_descent(self):
y_pred = np.dot(self.X, self.theta)
# calculate gradient
dtheta = (1.0/self.m) * np.dot(self.X.T,(y_pred - self.y))
# update weights
self.theta -= self.lr * dtheta
return self
def predict(self,X):
m,n = X.shape
X = np.append(np.ones((m, 1)), X, axis=1)
return np.dot(X,self.theta)
def score(self,X,y):
y_pred = self.predict(X)
u = ((y-y_pred)**2).sum()
v = ((y-y.mean())**2).sum()
return (1-(u/v))
model = LinearRegression(lr = 0.0000002,itr=1000)
model.fit(x_train, y_train)
y_pred = model.predict(x_test)
y_pred
#vizulization predictions Vs actual
fig = plt.figure(figsize =(20,8))
plt.plot(preds_linear,label = 'prediction')
plt.plot(y_test, label = 'actual')
plt.show()
ac_lr = lr_model.score(x_test,y_test)
print('R^2: ',ac_lr)
print("Mean Absolute Error:", mean_absolute_error(y_test, y_pred))
print("Coefficient of Determination:", r2_score(y_test, preds_linear))
print("MSE: ",mean_squared_error(y_test, preds_linear))
print("RMSE: ",np.sqrt(mean_squared_error(y_test, preds_linear)))
# + [markdown] id="GUWaZip2qD_G"
# #Linear Regss
# + colab={"base_uri": "https://localhost:8080/"} id="at0ceKaYp-fJ" executionInfo={"status": "ok", "timestamp": 1637317589410, "user_tz": -330, "elapsed": 991, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AO<KEY>", "userId": "12752491426441843216"}} outputId="0cd76c1d-699a-4371-b754-baf89f467f3d"
def cost_function(X, y, w, b):
cost = np.sum((((X.dot(w) + b) - y) ** 2) / (2*len(y)))
return cost
def gradient_descent_function(X, y, w, b, alpha=0.001, epochs=1000):
m = len(y)
costs = [0] * epochs
for epoch in range(epochs):
# Calculate the value -- Forward Propagation
z = X.dot(w) + b
# Calculate the losses
loss = z - y
# Calculate gradient descent
weight_gradient = X.T.dot(loss) / m
bias_gradient = np.sum(loss) / m
# Update weights and bias
w = w - alpha*weight_gradient
b = b - alpha*bias_gradient
# Store current lost
cost = cost_function(X, y, w, b)
costs[epoch] = cost
return w, b, costs
# Initial random weights
w = np.random.randn(x_train.shape[1])
# Initial bias
bias = 0
weights, bias, costs = gradient_descent_function(x_train, y_train, w, bias, epochs=1500);
def predict(X, w, b):
return X.dot(w) + b
ydfg = predict(x_test, weights, bias)
mean_squared_error(y_test, ydfg)*100
# + id="zUh9cv_0qBjy"
#vizulization predictions Vs actual
fig = plt.figure(figsize =(20,8))
plt.plot(preds_linear,label = 'prediction')
plt.plot(y_test, label = 'actual')
plt.show()
ac_lr = lr_model.score(x_test,y_test)
print('R^2: ',ac_lr)
print("Mean Absolute Error:", mean_absolute_error(y_test, preds_linear))
print("Coefficient of Determination:", r2_score(y_test, preds_linear))
print("MSE: ",mean_squared_error(y_test, preds_linear))
print("RMSE: ",np.sqrt(mean_squared_error(y_test, preds_linear)))
# + [markdown] id="Gqe74eWOI3xx"
# #Decision tree
# + id="d1eQCazvj5wI" executionInfo={"status": "ok", "timestamp": 1637306566442, "user_tz": -330, "elapsed": 3, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjc2nHKs9TxBjI8Rdr48x2ktCUgrltQi-Ca-pdomA=s64", "userId": "12752491426441843216"}}
#Decision Tree
#Building Decision Tree Model
from sklearn.tree import DecisionTreeRegressor
dtree = DecisionTreeRegressor(random_state=0)
dtree.fit(x_train,y_train)
pred_Dtree = dtree.predict(x_test)
# + colab={"base_uri": "https://localhost:8080/", "height": 483} id="XtLPA9YPkGOz" executionInfo={"status": "ok", "timestamp": 1637308028951, "user_tz": -330, "elapsed": 1190, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjc2nHKs9TxBjI8Rdr48x2ktCUgrltQi-Ca-pdomA=s64", "userId": "12752491426441843216"}} outputId="2401fea3-4b55-4889-b7f0-7f0ae24b1ec7"
fig = plt.figure(figsize =(20,8))
plt.plot(pred_Dtree,label = 'prediction')
plt.plot(y_test, label = 'actual')
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="Ju_AYkk4E6cV" executionInfo={"status": "ok", "timestamp": 1637308139471, "user_tz": -330, "elapsed": 403, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjc2nHKs9TxBjI8Rdr48x2ktCUgrltQi-Ca-pdomA=s64", "userId": "12752491426441843216"}} outputId="3d11abe4-2152-4cb3-9805-7ce2d5bd2b83"
dscore = dtree.score(x_test,y_test)
dscore
# + [markdown] id="NwZFZN6vI-YT"
# #Random Forest
# + colab={"base_uri": "https://localhost:8080/"} id="eenSnKW0ke6F" executionInfo={"status": "ok", "timestamp": 1637306757753, "user_tz": -330, "elapsed": 2365, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjc2nHKs9TxBjI8Rdr48x2ktCUgrltQi-Ca-pdomA=s64", "userId": "12752491426441843216"}} outputId="5077ba6a-0910-41a7-c788-6aab044eddcf"
from sklearn.ensemble import RandomForestRegressor
rnd= RandomForestRegressor()
c = rnd.fit(x_train, y_train)
rc = rnd.predict(x_test)
# + colab={"base_uri": "https://localhost:8080/", "height": 500} id="DPvSVC0Sxp5Z" executionInfo={"status": "ok", "timestamp": 1637306775788, "user_tz": -330, "elapsed": 829, "user": {"displayName": "tejareddy piduru", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjc2nHKs9TxBjI8Rdr48x2ktCUgrltQi-Ca-pdomA=s64", "userId": "12752491426441843216"}} outputId="be07fed9-2009-457b-f8c9-a91061897df0"
fig = plt.figure(figsize =(20,8))
plt.plot(rc)
plt.plot(y_test)
# + colab={"base_uri": "https://localhost:8080/"} id="n6liUTFhv0Bs" executionInfo={"status": "ok", "timestamp": 1637307724847, "user_tz": -330, "elapsed": 702, "user": {"displayName": "te<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjc2nHKs9TxBjI8Rdr48x2ktCUgrltQi-Ca-pdomA=s64", "userId": "12752491426441843216"}} outputId="1f08efa6-d491-4aea-ba29-b753a5a21632"
rnd_mod = rnd.score(x_test,y_test)
rnd_mod
# + [markdown] id="J_aSJ196MSMp"
# #Final
#
# + colab={"base_uri": "https://localhost:8080/", "height": 483} id="lLBXV9XVK-yp" executionInfo={"status": "ok", "timestamp": 1637308471618, "user_tz": -330, "elapsed": 1016, "user": {"displayName": "tejareddy piduru", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjc2nHKs9TxBjI8Rdr48x2ktCUgrltQi-Ca-pdomA=s64", "userId": "12752491426441843216"}} outputId="66107b36-9e8a-4f95-cad3-f3cb3b180b22"
fig = plt.figure(figsize =(20,8))
plt.plot(y_test, label='actual')
plt.plot(preds_linear,label = 'linear_prediction')
plt.plot(pred_Dtree, label = 'Dtree prediction')
plt.plot(rc,label = 'Dtree prediction')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 349} id="n9pGEHF8JTLS" executionInfo={"status": "ok", "timestamp": 1637310189357, "user_tz": -330, "elapsed": 764, "user": {"displayName": "te<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjc2nHKs9TxBjI8Rdr48x2ktCUgrltQi-Ca-pdomA=s64", "userId": "12752491426441843216"}} outputId="e103db8c-e974-475d-c13c-4df14c3a4239"
sns.barplot(['LR','Decision tree','Random Forest'],[ac_lr,dscore,rnd_mod])
plt.title('Model comparison')
plt.xlabel('Model')
plt.ylabel('Accuracy')
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="JDoZgBKyPNhc" executionInfo={"status": "ok", "timestamp": 1637309601347, "user_tz": -330, "elapsed": 403, "user": {"displayName": "tejareddy piduru", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjc2nHKs9TxBjI8Rdr48x2ktCUgrltQi-Ca-pdomA=s64", "userId": "12752491426441843216"}} outputId="92c91c54-a3f0-4003-e9aa-f3f6ce34647a"
print("R^2 scores\nlinear regression: ",ac_lr,"\ndecission tree regressor: ",dscore,'\nRandom forest regressor: ',rnd_mod)
# + colab={"base_uri": "https://localhost:8080/"} id="XbcW8eGlQfKY" executionInfo={"status": "ok", "timestamp": 1637309762178, "user_tz": -330, "elapsed": 401, "user": {"displayName": "tejareddy piduru", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjc2nHKs9TxBjI8Rdr48x2ktCUgrltQi-Ca-pdomA=s64", "userId": "12752491426441843216"}} outputId="71ad4e6f-6ea9-4628-f257-a923a1a6068b"
print("Rmse scores\nlinear regression: ",np.sqrt(mean_squared_error(y_test, preds_linear)),"\ndecission tree regressor: ",np.sqrt(mean_squared_error(y_test, pred_Dtree)),'\nRandom forest regressor: ',np.sqrt(mean_squared_error(y_test, rc)))
# + colab={"base_uri": "https://localhost:8080/", "height": 349} id="C5m-I4geRFQE" executionInfo={"status": "ok", "timestamp": 1637310196939, "user_tz": -330, "elapsed": 1122, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjc2nHKs9TxBjI8Rdr48x2ktCUgrltQi-Ca-pdomA=s64", "userId": "12752491426441843216"}} outputId="be3eb9f7-aa49-493f-913d-e0f3c2152299"
sns.barplot(['LR','Decision tree','RandomForest'],[np.sqrt(mean_squared_error(y_test, preds_linear)),np.sqrt(mean_squared_error(y_test, pred_Dtree)),np.sqrt(mean_squared_error(y_test, rc))])
plt.title('Model comparison')
plt.xlabel('Model')
plt.ylabel('Accuracy')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 500} id="Aqz2XKscWfDN" executionInfo={"status": "ok", "timestamp": 1637311219824, "user_tz": -330, "elapsed": 807, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gjc2nHKs9TxBjI8Rdr48x2ktCUgrltQi-Ca-pdomA=s64", "userId": "12752491426441843216"}} outputId="b71af8f2-2720-48b0-e4c7-244e413058a5"
fig = plt.figure(figsize =(20,8))
plt.plot(df['Date'],df['Close'], label='Close Price history')
| Ml/MLproject/project.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <NAME>
# patterns
n=int(input())
i=1
while i<= n:
j=1
while j <= n:
print('*', end='')
j=j+1
print()
i=i+1
n=int(input())
i=1
while i<= n:
j=1
while j <= i:
print(j, end='')
j=j+1
print()
i=i+1
# ##### n= int(input())
# i=1
# while i<=n :
# j= 1
# p = i
# while j <=i :
# print(p, end='')
# j=j+1
# p=p+1
# print()
# i= i+1
#
# +
n=int(input())
i=1
p=1
while i<= n:
j=1
while j <= i:
print(p, end='')
j=j+1
p=p+1
print()
i=i+1
# -
n=int(input())
i=1
j=1
k=1
while i<= n:
j=j+1
p=1
while p <= i:
k=j-1
print(k, end='')
p=p+1
print()
i=i+1
a=66
print(chr(a))
n= int(input())
i=1
while i<=n :
j= 1
k=65
p = k
while j <=i :
print(chr(p), end='a')
j=j+1
p=p+1
print()
k=k+1
i= i+1
n=int(input())
for i in range(n+1, 1, -1):
for j in range(1, i):
print(j, end="")
print()
n = int(input())
i=1
while i<=n:
j=0
while j<i:
if i==1:
print(1, end="")
elif j and j<i-1:
print(2, end="")
else:
print(1, end='')
j=j+1
print()
i=i+1
n = int(input("Enter a number of n: "))
for i in range(n):
print((chr(65+i)+" ")*(i+1))
n=int(input())
for i in range(65,65+n):
k=i
# Inner loop
for j in range(65,i+1):
print(chr(k),end="")
k=k+1
print()
# +
n = int(input())
for i in range(n, 0, -1):
for j in range(i, n + 1, 1):
print(chr(ord('A') + j - 1),
end = " ")
print("")
# -
# +
Number = int(input())
i = 0
First_Value = 1
Second_Value = 1
while(i < Number):
if(i <= 1):
Next = 1
else:
Next = First_Value + Second_Value
First_Value = Second_Value
Second_Value = Next
i = i + 1
print(Next)
# -
| Patterns 1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Wildfire statistics
#
# PAIRS layer `50035` contains information on _burned areas_. I.e. a pixel has the value 1 on the timestamp a burn occured. (The data is derived from http://modis-fire.umd.edu/ba.html.) Leveraging the spatial and temporal aggregation features in PAIRS allows us to calculate burned areas for each state in the US and year.
# +
import os, numpy as np, pandas as pd, logging, matplotlib.pyplot as plt, re, requests, sklearn
from datetime import datetime, timedelta
from ibmpairs import paw
os.environ['PAW_PAIRS_DEFAULT_PASSWORD_FILE_NAME'] = '<path to ibmpairspass.txt>'
os.environ['PAW_PAIRS_DEFAULT_USER'] = '<username>'
os.environ['PAW_LOG_LEVEL_ENV'] = 'WARNING'
paw.load_environment_variables()
logging.basicConfig(level = logging.WARNING)
pawLogger = logging.getLogger('ibmpairs.paw')
pawLogger.setLevel(logging.ERROR)
# -
iso8601 = '%Y-%m-%dT%H:%M:%SZ'
yearStart, yearEnd = 2001, 2019
burnableLand = [62, 63, 64, 141, 142, 143, 152, 176]
# The following JSON defines the PAIRS query. We are requesting years 2010 to 2018. Since we are interested in the aggregate annual burned areas, we are performing a `Max` aggregation. (Since pixels in this layer have the value 1 exclusively, `Min` and `Mean` would give the same result. `Sum` would not for pixels that were affected by multiple fires.)
#
# Note that spatial aggregation is always performed _last_ in PAIRS. In the case at hand, we aggregate across US states.
queryJson = {
"layers": [
{
"alias" : "B{y}".format(y = y),
"id": '50035',
"output": True,
"temporal" : {"intervals" : [{
"start" : datetime(y, 1, 1).strftime(iso8601), "end" : datetime(y+1, 1, 1).strftime(iso8601)
}]},
"aggregation" : "Max",
"type": "raster"
}
for y in range(yearStart, yearEnd)],
"name": "Burned areas",
"spatial" : {
"type" : "poly", "aoi" : "24",
"aggregation": {
"aoi": [
i for i in range(121, 172)
]
}
},
"temporal": {"intervals": [{"start" : datetime(2019, 1, 1).strftime(iso8601), "end" : datetime(2020, 1, 1).strftime(iso8601)}]},
}
# However, we are not only interested in the wildfire data. Since PAIRS offers a multitude of additional data layers, we can look for statistically significant dependencies. Thus we add layers `92` and `49069`. The former is the daily maximum temperature as reported by PRISM, the latter daily averaged soil moisture at a depth of 0-7 cm in m3/m3. The soil moisture data is calculated from the ECMWF interim reanalysis.
queryJson["layers"].extend([
{
"alias" : "TMax{y}".format(y = y),
"id": '92',
"output": True,
"temporal" : {"intervals" : [{
"start" : datetime(y, 1, 1).strftime(iso8601), "end" : datetime(y+1, 1, 1).strftime(iso8601)
}]},
"aggregation" : "Max",
"type": "raster"
}
for y in range(yearStart, yearEnd)
])
queryJson["layers"].extend([
{
"alias" : "SW{y}".format(y = y),
"id": '49069',
"output": True,
"temporal" : {"intervals" : [{
"start" : datetime(y, 1, 1).strftime(iso8601), "end" : datetime(y+1, 1, 1).strftime(iso8601)
}]},
"aggregation" : "Sum",
"type": "raster"
}
for y in range(yearStart, yearEnd)
])
query = paw.PAIRSQuery(queryJson)
query.submit()
# Note: Spatial aggregation is not the fastest process. This might take an hour to run.
query.poll_till_finished()
query.download()
query.create_layers()
# Since all the data is aggregated by state, PAIRS essentially returns a number of data frames in the form of csv files.
# # Analysis
# The first step in our analysis is some simple data munging. We take the various data frames returned by PAIRS and wrapped in `query.data` and extract the features we are interested in. For the burned area data this is the area, which can be found in the `count()[unit: km^2]` statistic. For the maximum temperature we choose the spatial maximum, for the soil water content the spatial mean.
from sklearn import linear_model
tmaxData = list()
burnData = list()
swData = list()
for k in query.data.keys():
swYear = re.match('Daily global weather\-Daily SWVL 1 History\[SW([0-9]{4})\]\-Sum', k)
tmaxYear = re.match('Daily US weather \(PRISM\)\-Daily maximum temperature\[TMax([0-9]{4})\]\-Max', k)
burnedYear = re.match('Burned area \(MODIS\)\-Burned area\[B([0-9]{4})\]\-Max', k)
if tmaxYear:
query.data[k]['Year'] = int(tmaxYear.group(1))
tmaxData.append(query.data[k])
elif burnedYear:
query.data[k]['Year'] = int(burnedYear.group(1))
burnData.append(query.data[k])
elif swYear:
query.data[k]['Year'] = int(swYear.group(1))
swData.append(query.data[k])
else:
raise Exception('No match for key {}.'.format(k))
tmaxData = pd.concat(tmaxData, 0).reset_index(drop = True)
burnData = pd.concat(burnData, 0).reset_index(drop = True)
swData = pd.concat(swData, 0).reset_index(drop = True)
# Later on we will analyze the data by state. To do so, we load information on the various AOIs from PAIRS. Strictly speaking we are not only aggregating by state since we treat the District of Columbia separately.
aoiDetails = [requests.get('https://pairs.res.ibm.com/ws/queryaois/aoi/{aoi}'.format(aoi = aoi), auth = query.auth).json() for aoi in range(121, 172)]
aoiDetails = pd.DataFrame(aoiDetails)
allTheData = pd.merge(
burnData[['PAIRS polygon ID', 'Year', 'count()[unit: km^2]']],
tmaxData[['PAIRS polygon ID', 'Year', 'max()']],
on = ['PAIRS polygon ID', 'Year'], how = 'outer'
).rename(columns = {'count()[unit: km^2]' : 'BurnedArea', 'max()' : 'TMax'})
allTheData['BurnedArea'].fillna(0, inplace = True)
allTheData = pd.merge(allTheData, swData[['PAIRS polygon ID', 'Year', 'mean()']], on = ['PAIRS polygon ID', 'Year'], how = 'outer').rename(columns = {'mean()' : 'SoilWater'})
allTheData = pd.merge(allTheData, aoiDetails[['id', 'name']], left_on = 'PAIRS polygon ID', right_on = 'id', how = 'inner').drop(['PAIRS polygon ID', 'id'], axis = 1)
# ## Conterminous USA
#
# To start, we take a look at the conterminous USA. (If we wanted to analyze Hawaii or Alaska, we could not use the PRISM data. An obvious replacement would be datasets 190 or 157, the _Global weather (ERA5)_ and _Current and historical weather (IBM TWC)_ respectively.
conusData = allTheData.groupby('Year').aggregate({'BurnedArea' : np.sum, 'TMax' : np.mean, 'SoilWater' : np.mean})
conusData.corr()
# Having take a look at correlations, we fit 3 models -- one for each combination of independent variables.
conusModel = linear_model.LinearRegression()
conusModel.fit(conusData[['TMax']] - conusData[['TMax']].mean(), conusData['BurnedArea'])
conusModel.score(conusData[['TMax']] - conusData[['TMax']].mean(), conusData['BurnedArea'])
conusModel2 = linear_model.LinearRegression()
conusModel2.fit(conusData[['TMax', 'SoilWater']] - conusData[['TMax', 'SoilWater']].mean(), conusData['BurnedArea'])
conusModel2.score(conusData[['TMax', 'SoilWater']] - conusData[['TMax', 'SoilWater']].mean(), conusData['BurnedArea'])
conusModel3 = linear_model.LinearRegression()
conusModel3.fit(conusData[['SoilWater']] - conusData[['SoilWater']].mean(), conusData['BurnedArea'])
conusModel3.score(conusData[['SoilWater']] - conusData[['SoilWater']].mean(), conusData['BurnedArea'])
# As one might expect the model containing both maximum temperature and soil water is best in terms of $R^2$ score. However, the improvement is marginal. For simplicity we continue our analysis focussing on a simple linear dependence between burned area and maximum temperature.
X = np.linspace(
conusData['TMax'].min() - conusData['TMax'].mean(),
conusData['TMax'].max() - conusData['TMax'].mean(), 100
).reshape(-1, 1)
y = conusModel.predict(X)
plt.figure(figsize = (12, 8))
plt.scatter(conusData['TMax'] - conusData['TMax'].mean(), conusData['BurnedArea'])
plt.plot(X, y, color = 'red')
plt.xlabel('Maximum temperature (deviation from mean) [K]')
plt.ylabel('Burned area [Km2]')
plt.title('Conterminous USA - Burned area vs. annual maximum temperature')
plt.savefig('linearDependence.png', bbox_inches = 'tight')
plt.show()
# To evaluate our model, we calculate some statistical scores. Specifically, we calculate
#
# $
# \hat{\beta} = (X^T X)^{-1} X^T y \\
# \hat{y} = X \hat{\beta} \\
# \hat{\sigma}^2 = \frac{1}{N-p-1} \sum (y - \hat{y})^2 \\
# Var(\hat{\beta}) = (X^T X)^{-1} \hat{\sigma}^2 \\
# v_j = (X^T X)^{-1}_{jj} \\
# z_j = \frac{\hat{\beta}_j}{\hat{\sigma} \sqrt{v_j}}
# $
# +
X = (conusData[['TMax']] - conusData[['TMax']].mean())
X['1'] = 1
X = X[['1', 'TMax']].values
y = conusData['BurnedArea'].values
betaHat = np.matmul(np.matmul(np.linalg.inv(np.matmul(X.transpose(), X)), X.transpose()), y)
yHat = np.matmul(X, betaHat)
sigmaHat2 = np.power(y - yHat, 2).sum() / (len(y) - 1 - 1)
VarBeta = np.linalg.inv(np.matmul(X.transpose(), X)) * sigmaHat2
v = np.array([np.linalg.inv(np.matmul(X.transpose(), X))[i, i] for i in range(2)])
zScore = betaHat / (np.sqrt(sigmaHat2 * v))
# -
zScore
# ## Per-state analysis
#
# To proceed, we would like to evaluate and compare different states. This poses a problem however: How do we best compare areas affected by wildfires across states with vastly different spatial coverage? An additional $1 Km^2$ in Rhode Island is a very different situation to an additional $1 Km^2$ in Alaska. One possibility is to normalize the burned area we are considering before by the _burnable area_; i.e. the fraction of a state's area that is covered by forests, shrubland or other vegetation that is susceptible to wildfires. To do so, we will query layer `48522`, which contains land use data from the US Department of Agriculture. The original data has 30 m resolution. `48522` is a convenience product that contains the same information coarse-grained to 250 m.
#
# Note: One can certainly argue whether our methodology is correct. From an ecological perspective, every square kilometer of burned area is problematic, no matter where. Having said that, note that the mathematics of our linear fits will not be affected by a simple rescaling of the dependent variable.
#
# ### Normalizing the area
#
# In either case, we proceed by grabbing the data from layer `48522`. In principle one could do this for each year or aggregate over multiple years. In what follows we simply use 2016 as a representative.
burnableAreaQueryJson = {
"layers": [
{
"alias" : "crop2016",
"id": '48522',
"output": False,
"temporal" : {"intervals" : [{
"snapshot" : datetime(2016, 1, 1).strftime(iso8601)
}]},
"type": "raster"
},
{
"alias" : "burnable2016",
"output": True,
"expression" : "0 + (" + " || ".join(["($crop2016 == {})".format(crop) for crop in burnableLand]) + ")"
}
],
"name": "BurnedAreas",
"spatial" : {
"type" : "poly", "aoi" : "24",
"aggregation" : {"aoi" : list(range(121, 172))}
},
"temporal": {"intervals": [{"start" : datetime(2019, 1, 1).strftime(iso8601), "end" : datetime(2020, 1, 1).strftime(iso8601)}]},
}
burnableAreaQuery = paw.PAIRSQuery(burnableAreaQueryJson)
burnableAreaQuery.submit()
burnableAreaQuery.poll_till_finished()
burnableAreaQuery.download()
burnableAreaQuery.create_layers()
# Having completed the query, we join the data ...
burnableArea = pd.merge(
burnableAreaQuery.data['Expression-burnable2016[burnable2016]-Exp'],
aoiDetails[['id', 'name']], left_on = 'PAIRS polygon ID', right_on = 'id', how = 'inner'
).drop(['PAIRS polygon ID', 'id', 'min()', 'max()', 'mean()', '2nd moment'], 1).rename(columns = {'count()[unit: km^2]' : 'BurnableArea'})
# ... and calculate the aforementioned fraction.
evenMoreData = pd.merge(allTheData, burnableArea, on = 'name', how = 'outer')
evenMoreData['BurnedFraction'] = evenMoreData['BurnedArea'] / evenMoreData['BurnableArea'] * 100
# We are now in the position to fit a linear model for each state. As we are ignoring Alaska and Hawaii and are treating D.C. as a state, we have exactly 49 states which we can nicely arrange in a grid:
# +
fig, ax = plt.subplots(7, 7, figsize = (25, 25), sharex = True, sharey = True)
betaHats, zScores, varBetaHats = dict(), dict(), dict()
for i, state in evenMoreData.groupby('name').aggregate({'BurnedArea' : np.sum}).sort_values('BurnedArea', ascending = False).index.to_series().reset_index(drop = True).iteritems():
stateData = evenMoreData[evenMoreData['name'] == state]
# We fit a model
stateModel = linear_model.LinearRegression()
stateModel.fit(stateData[['TMax']] - stateData[['TMax']].mean(), stateData['BurnedFraction'])
# Evaluating the model
X = (stateData[['TMax']] - stateData[['TMax']].mean())
X['1'] = 1
X = X[['1', 'TMax']].values
y = stateData['BurnedFraction'].values
betaHat = np.matmul(np.matmul(np.linalg.inv(np.matmul(X.transpose(), X)), X.transpose()), y)
yHat = np.matmul(X, betaHat)
sigmaHat2 = np.power(y - yHat, 2).sum() / (len(y) - 1 - 1)
varBetaHat = np.linalg.inv(np.matmul(X.transpose(), X)) * sigmaHat2
v = np.array([np.linalg.inv(np.matmul(X.transpose(), X))[i, i] for i in range(2)])
zScore = betaHat / (np.sqrt(sigmaHat2 * v))
betaHats[state] = betaHat[1]
zScores[state] = zScore[1]
varBetaHats[state] = varBetaHat[1, 1]
# Plotting
X = np.linspace(-3.5, 5.0, 100).reshape(-1, 1)
y = stateModel.predict(X)
row = i//7
column = i % 7
ax[row, column].scatter(stateData['TMax'] - stateData['TMax'].mean(), stateData['BurnedFraction'])
ax[row, column].plot(X, y, color = 'red')
ax[row, column].set_title(state)
for a in ax.flat:
a.set(xlabel='ΔTemperature [K]', ylabel='Area [%]')
a.label_outer()
plt.savefig('linarDependenceByState.png', bbox_inches = 'tight')
plt.show()
# -
# To conclude, we pick the states with significant z-scores. Interestingly, this does not include California, which shows us how noisy this data is.
scoreSummary = pd.merge(pd.DataFrame({'z' : zScores, 'beta' : betaHats, 'Var(beta)' : varBetaHats}).sort_values('z', ascending = False), evenMoreData.groupby('name').aggregate({'BurnedArea' : np.sum, 'BurnedFraction' : np.mean}), left_index = True, right_index = True)
scoreSummary[scoreSummary['z'].abs() > 2.0].sort_values('BurnedArea', ascending = False)
# ### Visualization
#
# Finally, we can visualize our results
from shapely.geometry import shape, multipolygon
import geojson, geopandas
polygons = [(aoi, shape(geojson.loads(requests.get('https://pairs.res.ibm.com/ws/queryaois/geojson/{aoi}'.format(aoi = aoi), auth = query.auth).json()))) for aoi in range(121, 172)]
polygonsDF = pd.DataFrame(polygons, columns = ['id', 'Geometry'])
polygonsDF = pd.merge(polygonsDF, aoiDetails[['id', 'name']]).drop(['id'], 1)
scoredPolygons = geopandas.GeoDataFrame(pd.merge(scoreSummary, polygonsDF, left_index = True, right_on = 'name'), geometry = 'Geometry')
# +
fig, ax = plt.subplots(2, 2, figsize = (24, 8), sharex = True, sharey = True)
scoredPolygons.plot(column = 'beta', cmap = 'RdYlGn_r', ax = ax[0][0], legend = True, vmax = scoredPolygons['beta'].abs().max(), vmin = -scoredPolygons['beta'].abs().max())
ax[0][0].set_ylabel('Latitude')
ax[0][0].set_title('beta - burned fraction vs. temperature [% area / K]')
scoredPolygons.dropna().plot(column = 'BurnedFraction', cmap = 'Reds', ax = ax[0][1], legend = True)
ax[0][1].set_title('Burned area fraction')
scoredPolygons.plot(column = 'Var(beta)', cmap = 'Blues_r', ax = ax[1][0], legend = True)
ax[1][0].set_ylabel('Latitude')
ax[1][0].set_xlabel('Longitude')
ax[1][0].set_title('Var(beta)')
scoredPolygons.dropna().plot(column = 'z', cmap = 'coolwarm', ax = ax[1][1], legend = True, vmin = -2.2, vmax = 2.2)
ax[1][1].set_xlabel('Longitude')
ax[1][1].set_title('z-scores')
plt.savefig('maps.png', dpi = 60, bbox_inches = 'tight')
plt.show()
# -
| examples/WildfireStatisticsUSA/WildfireAnalytics.ipynb |