code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import os,sys
sys.path.append('../../RL_lib/Agents')
sys.path.append('../../RL_lib/Policies/PPO')
sys.path.append('../../RL_lib/Policies/Common')
sys.path.append('../../RL_lib/Utils')
sys.path.append('../../Env')
sys.path.append('../../Imaging')
# %load_ext autoreload
# %load_ext autoreload
# %autoreload 2
# %matplotlib nbagg
import os
print(os.getcwd())
# + language="html"
# <style>
# .output_wrapper, .output {
# height:auto !important;
# max-height:1000px; /* your desired max-height here */
# }
# .output_scroll {
# box-shadow:none !important;
# webkit-box-shadow:none !important;
# }
# </style>
# -
# # Optimize Policy
# +
from env import Env
import env_utils as envu
from dynamics_model import Dynamics_model
from lander_model import Lander_model
from ic_gen_scene import Landing_icgen
import rl_utils
import attitude_utils as attu
from arch_policy_gtvf import Arch
from softmax_pd import Softmax_pd as PD
from policy_ppo import Policy
from value_function import Value_function
import policy_nets as policy_nets
import valfunc_nets as valfunc_nets
import cnn_nets
from agent import Agent
import torch.nn as nn
from flat_constraint import Flat_constraint
from glideslope_constraint import Glideslope_constraint
from rh_constraint import RH_constraint
from no_attitude_constraint import Attitude_constraint
from w_constraint import W_constraint
from reward_terminal_mdr import Reward
from asteroid_hfr_scene import Asteroid
from thruster_model import Thruster_model
asteroid_model = Asteroid(landing_site_override=None, omega_range=(1e-6,5e-4))
ap = attu.Quaternion_attitude()
from flash_lidar2 import Flash_lidar
import attitude_utils as attu
from triangle_ray_intersect import Triangle_ray_intersect
from isosphere import Isosphere
iso = Isosphere(recursion_level=2)
tri = Triangle_ray_intersect()
ap = attu.Quaternion_attitude()
P = 64
sensor = Flash_lidar(ap, tri, sqrt_pixels=int(np.sqrt(P)))
thruster_model = Thruster_model(pulsed=True, scale=1.0, offset=0.4)
lander_model = Lander_model(asteroid_model, thruster_model, ap, sensor, iso)
lander_model.get_state_agent = lander_model.get_state_agent_image_state_stab
logger = rl_utils.Logger()
dynamics_model = Dynamics_model(h=2)
obs_dim = 2*P
gt_dim = 13
action_dim = 12
actions_per_dim = 2
action_logit_dim = action_dim * actions_per_dim
recurrent_steps = 60
reward_object = Reward(landing_coeff=10.0, landing_rlimit=2, landing_vlimit=0.1, tracking_bias=0.01,
dalt_coeff=0.02, fuel_coeff=-0.01, use_gt=True)
glideslope_constraint = Glideslope_constraint(gs_limit=-1.0)
shape_constraint = Flat_constraint()
attitude_constraint = Attitude_constraint(ap)
w_constraint = W_constraint(w_limit=(0.1,0.1,0.1), w_margin=(0.05,0.05,0.05))
rh_constraint = RH_constraint(rh_limit=150)
wi=0.02
ic_gen = Landing_icgen(position_r=(50,600),
p_engine_fail=0.0,
p_scale=(0.01, 0.02),
engine_fail_scale=(1.0,1.0),
asteroid_axis_low=(300,300,300),
asteroid_axis_high=(600,600,600),
#position_theta=(0,np.pi/4),
lander_wll=(-wi,-wi,-wi),
lander_wul=(wi,wi,wi),
attitude_parameterization=ap,
attitude_error=(0,np.pi/16),
min_mass=450, max_mass=500,
debug=False,
inertia_uncertainty_diag=10.0,
inertia_uncertainty_offdiag=1.0)
env = Env(ic_gen, lander_model, dynamics_model, logger,
debug_done=False,
reward_object=reward_object,
glideslope_constraint=glideslope_constraint,
attitude_constraint=attitude_constraint,
w_constraint=w_constraint,
rh_constraint=rh_constraint,
tf_limit=600.0,print_every=10,nav_period=6)
env.ic_gen.show()
arch = Arch(gt_func=lander_model.get_state_agent_gt)
cnn = cnn_nets.CNN_layer(8,2,8)
policy = Policy(policy_nets.GRU_CNN2(7, action_logit_dim, cnn, recurrent_steps=recurrent_steps),
PD(action_dim, actions_per_dim),
shuffle=False, servo_kl=False, max_grad_norm=30,
init_func=rl_utils.xn_init, scale_image_obs=True, scale_vector_obs=True)
value_function = Value_function(valfunc_nets.GRU1(gt_dim, recurrent_steps=recurrent_steps), rollout_limit=3,
shuffle=False, batch_size=9999999, max_grad_norm=30, obs_key='gt_observes')
agent = Agent(arch, policy, value_function, None, env, logger,
policy_episodes=60, policy_steps=3000, gamma1=0.95, gamma2=0.995,
recurrent_steps=recurrent_steps, monitor=env.rl_stats)
fname = "optimize-RPT2"
policy.load_params(fname)
# -
# # Test Policy
# +
env.test_policy_batch(agent,5000,print_every=100,keys=lander_model.get_engagement_keys())
# -
| Experiments/Test/Test_50_600.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Exploring Ensemble Methods
# In this assignment, we will explore the use of boosting. We will use the pre-implemented gradient boosted trees in GraphLab Create. You will:
#
# * Use SFrames to do some feature engineering.
# * Train a boosted ensemble of decision-trees (gradient boosted trees) on the LendingClub dataset.
# * Predict whether a loan will default along with prediction probabilities (on a validation set).
# * Evaluate the trained model and compare it with a baseline.
# * Find the most positive and negative loans using the learned model.
# * Explore how the number of trees influences classification performance.
#
# Let's get started!
#
# ## Fire up Graphlab Create
import graphlab
# # Load LendingClub dataset
#
# We will be using the [LendingClub](https://www.lendingclub.com/) data. As discussed earlier, the [LendingClub](https://www.lendingclub.com/) is a peer-to-peer leading company that directly connects borrowers and potential lenders/investors.
#
# Just like we did in previous assignments, we will build a classification model to predict whether or not a loan provided by lending club is likely to default.
#
# Let us start by loading the data.
loans = graphlab.SFrame('lending-club-data.gl/')
# Let's quickly explore what the dataset looks like. First, let's print out the column names to see what features we have in this dataset. We have done this in previous assignments, so we won't belabor this here.
loans.column_names()
# ## Modifying the target column
#
# The target column (label column) of the dataset that we are interested in is called `bad_loans`. In this column **1** means a risky (bad) loan **0** means a safe loan.
#
# As in past assignments, in order to make this more intuitive and consistent with the lectures, we reassign the target to be:
# * **+1** as a safe loan,
# * **-1** as a risky (bad) loan.
#
# We put this in a new column called `safe_loans`.
loans['safe_loans'] = loans['bad_loans'].apply(lambda x : +1 if x==0 else -1)
loans = loans.remove_column('bad_loans')
# ## Selecting features
#
# In this assignment, we will be using a subset of features (categorical and numeric). The features we will be using are **described in the code comments** below. If you are a finance geek, the [LendingClub](https://www.lendingclub.com/) website has a lot more details about these features.
#
# The features we will be using are described in the code comments below:
target = 'safe_loans'
features = ['grade', # grade of the loan (categorical)
'sub_grade_num', # sub-grade of the loan as a number from 0 to 1
'short_emp', # one year or less of employment
'emp_length_num', # number of years of employment
'home_ownership', # home_ownership status: own, mortgage or rent
'dti', # debt to income ratio
'purpose', # the purpose of the loan
'payment_inc_ratio', # ratio of the monthly payment to income
'delinq_2yrs', # number of delinquincies
'delinq_2yrs_zero', # no delinquincies in last 2 years
'inq_last_6mths', # number of creditor inquiries in last 6 months
'last_delinq_none', # has borrower had a delinquincy
'last_major_derog_none', # has borrower had 90 day or worse rating
'open_acc', # number of open credit accounts
'pub_rec', # number of derogatory public records
'pub_rec_zero', # no derogatory public records
'revol_util', # percent of available credit being used
'total_rec_late_fee', # total late fees received to day
'int_rate', # interest rate of the loan
'total_rec_int', # interest received to date
'annual_inc', # annual income of borrower
'funded_amnt', # amount committed to the loan
'funded_amnt_inv', # amount committed by investors for the loan
'installment', # monthly payment owed by the borrower
]
# ## Skipping observations with missing values
#
# Recall from the lectures that one common approach to coping with missing values is to **skip** observations that contain missing values.
#
# We run the following code to do so:
# +
loans, loans_with_na = loans[[target] + features].dropna_split()
# Count the number of rows with missing data
num_rows_with_na = loans_with_na.num_rows()
num_rows = loans.num_rows()
print 'Dropping %s observations; keeping %s ' % (num_rows_with_na, num_rows)
# -
# Fortunately, there are not too many missing values. We are retaining most of the data.
# ## Make sure the classes are balanced
# We saw in an earlier assignment that this dataset is also imbalanced. We will undersample the larger class (safe loans) in order to balance out our dataset. We used `seed=1` to make sure everyone gets the same results.
# +
safe_loans_raw = loans[loans[target] == 1]
risky_loans_raw = loans[loans[target] == -1]
# Undersample the safe loans.
percentage = len(risky_loans_raw)/float(len(safe_loans_raw))
safe_loans = safe_loans_raw.sample(percentage, seed = 1)
risky_loans = risky_loans_raw
loans_data = risky_loans.append(safe_loans)
print "Percentage of safe loans :", len(safe_loans) / float(len(loans_data))
print "Percentage of risky loans :", len(risky_loans) / float(len(loans_data))
print "Total number of loans in our new dataset :", len(loans_data)
# -
# **Checkpoint:** You should now see that the dataset is balanced (approximately 50-50 safe vs risky loans).
# **Note:** There are many approaches for dealing with imbalanced data, including some where we modify the learning algorithm. These approaches are beyond the scope of this course, but some of them are reviewed in this [paper](http://ieeexplore.ieee.org/xpl/login.jsp?tp=&arnumber=5128907&url=http%3A%2F%2Fieeexplore.ieee.org%2Fiel5%2F69%2F5173046%2F05128907.pdf%3Farnumber%3D5128907 ). For this assignment, we use the simplest possible approach, where we subsample the overly represented class to get a more balanced dataset. In general, and especially when the data is highly imbalanced, we recommend using more advanced methods.
# ## Split data into training and validation sets
# We split the data into training data and validation data. We used `seed=1` to make sure everyone gets the same results. We will use the validation data to help us select model parameters.
train_data, validation_data = loans_data.random_split(.8, seed=1)
# # Gradient boosted tree classifier
# Gradient boosted trees are a powerful variant of boosting methods; they have been used to win many [Kaggle](https://www.kaggle.com/) competitions, and have been widely used in industry. We will explore the predictive power of multiple decision trees as opposed to a single decision tree.
#
# **Additional reading:** If you are interested in gradient boosted trees, here is some additional reading material:
# * [GraphLab Create user guide](https://dato.com/learn/userguide/supervised-learning/boosted_trees_classifier.html)
# * [Advanced material on boosted trees](http://homes.cs.washington.edu/~tqchen/pdf/BoostedTree.pdf)
#
#
# We will now train models to predict `safe_loans` using the features above. In this section, we will experiment with training an ensemble of 5 trees. To cap the ensemble classifier at 5 trees, we call the function with **max_iterations=5** (recall that each iterations corresponds to adding a tree). We set `validation_set=None` to make sure everyone gets the same results.
model_5 = graphlab.boosted_trees_classifier.create(train_data, validation_set=None,
target = target, features = features, max_iterations = 5)
# # Making predictions
#
# Just like we did in previous sections, let us consider a few positive and negative examples **from the validation set**. We will do the following:
# * Predict whether or not a loan is likely to default.
# * Predict the probability with which the loan is likely to default.
# +
# Select all positive and negative examples.
validation_safe_loans = validation_data[validation_data[target] == 1]
validation_risky_loans = validation_data[validation_data[target] == -1]
# Select 2 examples from the validation set for positive & negative loans
sample_validation_data_risky = validation_risky_loans[0:2]
sample_validation_data_safe = validation_safe_loans[0:2]
# Append the 4 examples into a single dataset
sample_validation_data = sample_validation_data_safe.append(sample_validation_data_risky)
sample_validation_data
# -
# ### Predicting on sample validation data
#
# For each row in the **sample_validation_data**, write code to make **model_5** predict whether or not the loan is classified as a **safe loan**.
#
# **Hint:** Use the `predict` method in `model_5` for this.
predictions = model_5.predict(sample_validation_data)
print predictions
# **Quiz question:** What percentage of the predictions on `sample_validation_data` did `model_5` get correct?
#
# ### Prediction probabilities
#
# For each row in the **sample_validation_data**, what is the probability (according **model_5**) of a loan being classified as **safe**?
#
# **Hint:** Set `output_type='probability'` to make **probability** predictions using `model_5` on `sample_validation_data`:
predictions_probability = model_5.predict(sample_validation_data, output_type='probability')
print predictions_probability
# **Quiz Question:** According to **model_5**, which loan is the least likely to be a safe loan?
#
# **Checkpoint:** Can you verify that for all the predictions with `probability >= 0.5`, the model predicted the label **+1**?
# ## Evaluating the model on the validation data
# Recall that the accuracy is defined as follows:
# $$
# \mbox{accuracy} = \frac{\mbox{# correctly classified examples}}{\mbox{# total examples}}
# $$
#
# Evaluate the accuracy of the **model_5** on the **validation_data**.
#
# **Hint**: Use the `.evaluate()` method in the model.
print model_5.evaluate(validation_data)['accuracy']
# Calculate the number of **false positives** made by the model.
predictions_validation_data = model_5.predict(validation_data)
print len(predictions_validation_data)
# **Quiz question**: What is the number of **false positives** on the **validation_data**?
# Calculate the number of **false negatives** made by the model.
# +
false_positive =0
false_negative =0
correct_predictions=0
for i in range(len(predictions_validation_data)):
if predictions_validation_data[i]== -1 and validation_data['safe_loans'][i]==1:
false_negative = false_negative +1
elif predictions_validation_data[i]== 1 and validation_data['safe_loans'][i]==-1:
false_positive = false_positive +1
elif predictions_validation_data[i]==validation_data['safe_loans'][i]:
correct_predictions = correct_predictions + 1
print false_positive
print false_negative
print correct_predictions
# -
# ## Comparison with decision trees
#
# In the earlier assignment, we saw that the prediction accuracy of the decision trees was around **0.64** (rounded). In this assignment, we saw that **model_5** has an accuracy of **0.67** (rounded).
#
# Here, we quantify the benefit of the extra 3% increase in accuracy of **model_5** in comparison with a single decision tree from the original decision tree assignment.
#
# As we explored in the earlier assignment, we calculated the cost of the mistakes made by the model. We again consider the same costs as follows:
#
# * **False negatives**: Assume a cost of \$10,000 per false negative.
# * **False positives**: Assume a cost of \$20,000 per false positive.
#
# Assume that the number of false positives and false negatives for the learned decision tree was
#
# * **False negatives**: 1936
# * **False positives**: 1503
#
# Using the costs defined above and the number of false positives and false negatives for the decision tree, we can calculate the total cost of the mistakes made by the decision tree model as follows:
#
# ```
# cost = $10,000 * 1936 + $20,000 * 1503 = $49,420,000
# ```
#
# The total cost of the mistakes of the model is $49.42M. That is a **lot of money**!.
#
# **Quiz Question**: Using the same costs of the false positives and false negatives, what is the cost of the mistakes made by the boosted tree model (**model_5**) as evaluated on the **validation_set**?
cost = 10000 * 1463 + 20000 * 1618
print cost
# **Reminder**: Compare the cost of the mistakes made by the boosted trees model with the decision tree model. The extra 3% improvement in prediction accuracy can translate to several million dollars! And, it was so easy to get by simply boosting our decision trees.
# ## Most positive & negative loans.
#
# In this section, we will find the loans that are most likely to be predicted **safe**. We can do this in a few steps:
#
# * **Step 1**: Use the **model_5** (the model with 5 trees) and make **probability predictions** for all the loans in the **validation_data**.
# * **Step 2**: Similar to what we did in the very first assignment, add the probability predictions as a column called **predictions** into the validation_data.
# * **Step 3**: Sort the data (in descreasing order) by the probability predictions.
#
# Start here with **Step 1** & **Step 2**. Make predictions using **model_5** for examples in the **validation_data**. Use `output_type = probability`.
predictions_probability_validation_data = model_5.predict(validation_data, output_type='probability')
validation_data['predictions']=predictions_probability_validation_data
# **Checkpoint:** For each row, the probabilities should be a number in the range **[0, 1]**. We have provided a simple check here to make sure your answers are correct.
print "Your loans : %s\n" % validation_data['predictions'].head(4)
print "Expected answer : %s" % [0.4492515948736132, 0.6119100103640573,
0.3835981314851436, 0.3693306705994325]
# Now, we are ready to go to **Step 3**. You can now use the `prediction` column to sort the loans in **validation_data** (in descending order) by prediction probability. Find the top 5 loans with the highest probability of being predicted as a **safe loan**.
top5_loans = validation_data.sort('predictions', ascending=False)
top5_loans[0:5]
bottom5_loans = validation_data.sort('predictions')
bottom5_loans
# ** Quiz question**: What grades are the top 5 loans?
#
# Let us repeat this excercise to find the top 5 loans (in the **validation_data**) with the **lowest probability** of being predicted as a **safe loan**:
positive_words = validation_data.topk('predictions', k=5, reverse=True)['grade']
print positive_words
# **Checkpoint:** You should expect to see 5 loans with the grade ['**D**', '**C**', '**C**', '**C**', '**B**'].
# ## Effect of adding more trees
# In this assignment, we will train 5 different ensemble classifiers in the form of gradient boosted trees. We will train models with 10, 50, 100, 200, and 500 trees. We use the **max_iterations** parameter in the boosted tree module.
#
# Let's get sarted with a model with **max_iterations = 10**:
model_10 = graphlab.boosted_trees_classifier.create(train_data, validation_set=None,
target = target, features = features, max_iterations = 10, verbose=False)
# Now, train 4 models with **max_iterations** to be:
# * `max_iterations = 50`,
# * `max_iterations = 100`
# * `max_iterations = 200`
# * `max_iterations = 500`.
#
# Let us call these models **model_50**, **model_100**, **model_200**, and **model_500**. You can pass in `verbose=False` in order to suppress the printed output.
#
# **Warning:** This could take a couple of minutes to run.
model_50 = graphlab.boosted_trees_classifier.create(train_data, validation_set=None,
target = target, features = features, max_iterations = 50, verbose=False)
model_100 = graphlab.boosted_trees_classifier.create(train_data, validation_set=None,
target = target, features = features, max_iterations = 100, verbose=False)
model_200 = graphlab.boosted_trees_classifier.create(train_data, validation_set=None,
target = target, features = features, max_iterations = 200, verbose=False)
model_500 = graphlab.boosted_trees_classifier.create(train_data, validation_set=None,
target = target, features = features, max_iterations = 500, verbose=False)
# ## Compare accuracy on entire validation set
# Now we will compare the predicitve accuracy of our models on the validation set. Evaluate the **accuracy** of the 10, 50, 100, 200, and 500 tree models on the **validation_data**. Use the `.evaluate` method.
print model_10.evaluate(validation_data)['accuracy']
print model_50.evaluate(validation_data)['accuracy']
print model_100.evaluate(validation_data)['accuracy']
print model_200.evaluate(validation_data)['accuracy']
print model_500.evaluate(validation_data)['accuracy']
# **Quiz Question:** Which model has the **best** accuracy on the **validation_data**?
#
# **Quiz Question:** Is it always true that the model with the most trees will perform best on test data?
# ## Plot the training and validation error vs. number of trees
# Recall from the lecture that the classification error is defined as
#
# $$
# \mbox{classification error} = 1 - \mbox{accuracy}
# $$
#
# In this section, we will plot the **training and validation errors versus the number of trees** to get a sense of how these models are performing. We will compare the 10, 50, 100, 200, and 500 tree models. You will need [matplotlib](http://matplotlib.org/downloads.html) in order to visualize the plots.
#
# First, make sure this block of code runs on your computer.
import matplotlib.pyplot as plt
# %matplotlib inline
def make_figure(dim, title, xlabel, ylabel, legend):
plt.rcParams['figure.figsize'] = dim
plt.title(title)
plt.xlabel(xlabel)
plt.ylabel(ylabel)
if legend is not None:
plt.legend(loc=legend, prop={'size':15})
plt.rcParams.update({'font.size': 16})
plt.tight_layout()
# In order to plot the classification errors (on the **train_data** and **validation_data**) versus the number of trees, we will need lists of these accuracies, which we get by applying the method `.evaluate`.
#
# **Steps to follow:**
#
# * **Step 1:** Calculate the classification error for model on the training data (**train_data**).
# * **Step 2:** Store the training errors into a list (called `training_errors`) that looks like this:
# ```
# [train_err_10, train_err_50, ..., train_err_500]
# ```
# * **Step 3:** Calculate the classification error of each model on the validation data (**validation_data**).
# * **Step 4:** Store the validation classification error into a list (called `validation_errors`) that looks like this:
# ```
# [validation_err_10, validation_err_50, ..., validation_err_500]
# ```
# Once that has been completed, the rest of the code should be able to evaluate correctly and generate the plot.
#
#
# Let us start with **Step 1**. Write code to compute the classification error on the **train_data** for models **model_10**, **model_50**, **model_100**, **model_200**, and **model_500**.
train_err_10=1-model_10.evaluate(train_data)['accuracy']
train_err_50=1-model_50.evaluate(train_data)['accuracy']
train_err_100=1-model_200.evaluate(train_data)['accuracy']
train_err_200=1-model_200.evaluate(train_data)['accuracy']
train_err_500=1-model_500.evaluate(train_data)['accuracy']
# Now, let us run **Step 2**. Save the training errors into a list called **training_errors**
training_errors = [train_err_10, train_err_50, train_err_100,
train_err_200, train_err_500]
# Now, onto **Step 3**. Write code to compute the classification error on the **validation_data** for models **model_10**, **model_50**, **model_100**, **model_200**, and **model_500**.
validation_err_10=1-model_10.evaluate(validation_data)['accuracy']
validation_err_50=1-model_50.evaluate(validation_data)['accuracy']
validation_err_100=1-model_200.evaluate(validation_data)['accuracy']
validation_err_200=1-model_200.evaluate(validation_data)['accuracy']
validation_err_500=1-model_500.evaluate(validation_data)['accuracy']
# Now, let us run **Step 4**. Save the training errors into a list called **validation_errors**
validation_errors = [validation_err_10, validation_err_50, validation_err_100,
validation_err_200, validation_err_500]
# Now, we will plot the **training_errors** and **validation_errors** versus the number of trees. We will compare the 10, 50, 100, 200, and 500 tree models. We provide some plotting code to visualize the plots within this notebook.
#
# Run the following code to visualize the plots.
# +
plt.plot([10, 50, 100, 200, 500], training_errors, linewidth=4.0, label='Training error')
plt.plot([10, 50, 100, 200, 500], validation_errors, linewidth=4.0, label='Validation error')
make_figure(dim=(10,5), title='Error vs number of trees',
xlabel='Number of trees',
ylabel='Classification error',
legend='best')
# -
# **Quiz question**: Does the training error reduce as the number of trees increases?
#
# **Quiz question**: Is it always true that the validation error will reduce as the number of trees increases?
| Machine_Learning_Classification/module-8-boosting-assignment-1-blank.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:covid_reddit_behaviour]
# language: python
# name: conda-env-covid_reddit_behaviour-py
# ---
# +
import pandas as pd
import altair as alt
alt.data_transformers.enable('data_server')
alt.renderers.enable('mimetype')
# -
# # Import Data
lonely_pre = pd.read_csv('../../data/lonely_pre_features_tfidf_256.csv')
lonely_post = pd.read_csv('../../data/lonely_post_features_tfidf_256.csv')
lonely_pre.head(3)
lonely_pre.tail(3)
# ## Features
#
# 1. The published paper, from which the dataset is cited, has an in depth breakdown of the features which can be found here: [https://www.jmir.org/2020/10/e22635/](https://www.jmir.org/2020/10/e22635/).
#
# 2. The question we are asking is: How has the substance use increased over the pandemic?
#
# - Feature of interest: `substance_use_total`
#
# - In order to accomplish our EDA task for Sunday 21st November, we will have to filter our dataset - and associated EDA tasks - to focus exclusively on this feature.
# ## High level analysis
# The question we are asking is *How has the substance use increased over the pandemic?*.
# For this project we have selected _one_ feature of interest: `substance_use_total`. In order to accomplish our EDA task for Sunday 21st November, we will have to filter our dataset - and associated EDA tasks - to focus exclusively on this feature.
# +
columns_of_interest = ['subreddit', 'author', 'date', 'post', 'substance_use_total']
lonely_pre = lonely_pre.loc[:, columns_of_interest]
lonely_post = lonely_post.loc[:, columns_of_interest]
# -
lonely_pre.head(5)
lonely_pre.info()
lonely_post.head(5)
lonely_post.info()
# ### Composition
# > There are no missing values from the dataset.
print(f'Total number of records in the pre-pandemic dataset: {len(lonely_pre)}')
lonely_pre.describe()
print(f'Total number of records in the post-pandemic dataset: {len(lonely_post)}')
lonely_post.describe()
# Combining the pre and post into one dataset with a new feature column allows us to compare the data side by side
# +
lonely_pre['period'] = 'pre'
lonely_post['period'] = 'post'
lonely_df = pd.concat([lonely_post, lonely_pre])
lonely_df
# -
# ### Visualization
alt.Chart(lonely_df).mark_bar(opacity=0.8).encode(
x=alt.X('substance_use_total', bin=alt.Bin(maxbins=30)),
y='count()',
color='period'
)
print(f"Number of unique authors (posters) in 'pre' dataset: {len(lonely_pre.author.unique())}")
print(f"Number of unique authors (posters) in 'post' dataset: {len(lonely_post.author.unique())}")
# **From above figures, it can be summarized that:**
#
# 1. Each observation in each dataset is associated with a unique reddit user.
# 2. There are less unique authors posting in this subreddit post pandemic. It is likely because the `post` dataset covers a 4-month period while the `pre` data set covers a whole year period.
# 3. The `substance_use_total` `post` pandemic is higher as shown in the plot.
| eda/subreddit/eda-lonely.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="MfBg1C5NB3X0"
# # Ungraded lab: Distributed Strategies with TF and Keras
# ------------------------
#
# + [markdown] id="8yL0-KcLPwtk"
#
# Welcome, during this ungraded lab you are going to perform a distributed training strategy using TensorFlow and Keras, specifically the [`tf.distribute.MultiWorkerMirroredStrategy`](https://www.tensorflow.org/api_docs/python/tf/distribute/MultiWorkerMirroredStrategy).
#
# With the help of this strategy, a Keras model that was designed to run on single-worker can seamlessly work on multiple workers with minimal code change. In particular you will:
#
#
# 1. Perform training with a single worker.
# 2. Understand the requirements for a multi-worker setup (`tf_config` variable) and using context managers for implementing distributed strategies.
# 3. Use magic commands to simulate different machines.
# 4. Perform a multi-worker training strategy.
#
# This notebook is based on the official [Multi-worker training with Keras](https://www.tensorflow.org/tutorials/distribute/multi_worker_with_keras) notebook, which covers some additional topics in case you want a deeper dive into this topic.
#
# [Distributed Training with TensorFlow](https://www.tensorflow.org/guide/distributed_training) guide is also available for an overview of the distribution strategies TensorFlow supports for those interested in a deeper understanding of `tf.distribute.Strategy` APIs.
#
# Let's get started!
# + [markdown] id="MUXex9ctTuDB"
# ## Setup
#
# First, some necessary imports.
# + id="bnYxvfLD-LW-"
import os
import sys
import json
import time
# + [markdown] id="Zz0EY91y3mxy"
# Before importing TensorFlow, make a few changes to the environment.
#
# - Disable all GPUs. This prevents errors caused by the workers all trying to use the same GPU. **For a real application each worker would be on a different machine.**
#
#
# - Add the current directory to python's path so modules in this directory can be imported.
# + id="685pbYEY3jGC"
# Disable GPUs
os.environ["CUDA_VISIBLE_DEVICES"] = "-1"
# Add current directory to path
if '.' not in sys.path:
sys.path.insert(0, '.')
# + [markdown] id="Rd4L9Ii77SS8"
# The previous step is important since this notebook relies on writting files using the magic command `%%writefile` and then importing them as modules.
#
# Now that the environment configuration is ready, import TensorFlow.
#
# + id="vHNvttzV43sA"
import tensorflow as tf
# Ignore warnings
tf.get_logger().setLevel('ERROR')
# + [markdown] id="0S2jpf6Sx50i"
# ### Dataset and model definition
# + [markdown] id="fLW6D2TzvC-4"
# Next create an `mnist.py` file with a simple model and dataset setup. This python file will be used by the worker-processes in this tutorial.
#
# The name of this file derives from the dataset you will be using which is called [mnist](https://keras.io/api/datasets/mnist/) and consists of 60,000 28x28 grayscale images of the first 10 digits.
# + colab={"base_uri": "https://localhost:8080/"} id="dma_wUAxZqo2" outputId="ba7dfe40-0b7e-4d67-b8fa-f33192e3afce"
# %%writefile mnist.py
# import os
import tensorflow as tf
import numpy as np
def mnist_dataset(batch_size):
# Load the data
(x_train, y_train), _ = tf.keras.datasets.mnist.load_data()
# Normalize pixel values for x_train and cast to float32
x_train = x_train / np.float32(255)
# Cast y_train to int64
y_train = y_train.astype(np.int64)
# Define repeated and shuffled dataset
train_dataset = tf.data.Dataset.from_tensor_slices((x_train, y_train)).shuffle(60000).repeat().batch(batch_size)
return train_dataset
def build_and_compile_cnn_model():
# Define simple CNN model using Keras Sequential
model = tf.keras.Sequential([
tf.keras.layers.InputLayer(input_shape=(28, 28)),
tf.keras.layers.Reshape(target_shape=(28, 28, 1)),
tf.keras.layers.Conv2D(32, 3, activation='relu'),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.Dense(10)
])
# Compile model
model.compile(
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
optimizer=tf.keras.optimizers.SGD(learning_rate=0.001),
metrics=['accuracy'])
return model
# + [markdown] id="CbDEKpGowcyT"
# Check that the file was succesfully created:
# + colab={"base_uri": "https://localhost:8080/"} id="IxsnfpVurQ1g" outputId="1865da85-bb2c-4b24-f15c-f26caafb0c56"
# !ls *.py
# + [markdown] id="2UL3kisMO90X"
# Import the mnist module you just created and try training the model for a small number of epochs to observe the results of a single worker to make sure everything works correctly.
# + colab={"background_save": true, "base_uri": "https://localhost:8080/"} id="6Qe6iAf5O8iJ" outputId="28ede9c6-cc28-4890-8893-ea25af385346"
# Import your mnist model
import mnist
# Set batch size
batch_size = 64
# Load the dataset
single_worker_dataset = mnist.mnist_dataset(batch_size)
# Load compiled CNN model
single_worker_model = mnist.build_and_compile_cnn_model()
# As training progresses, the loss should drop and the accuracy should increase.
single_worker_model.fit(single_worker_dataset, epochs=3, steps_per_epoch=70)
# + [markdown] id="ZpLPWFJh1CAK"
# Everything is working as expected!
#
# Now you will see how multiple workers can be used as a distributed strategy.
# + [markdown] id="JmgZwwymxqt5"
# ## Multi-worker Configuration
#
# Now let's enter the world of multi-worker training. In TensorFlow, the `TF_CONFIG` environment variable is required for training on multiple machines, each of which possibly has a different role. `TF_CONFIG` is a JSON string used to specify the cluster configuration on each worker that is part of the cluster.
#
# There are two components of `TF_CONFIG`: `cluster` and `task`.
#
# Let's dive into how they are used:
#
# `cluster`:
# - **It is the same for all workers** and provides information about the training cluster, which is a dict consisting of different types of jobs such as `worker`.
#
# - In multi-worker training with `MultiWorkerMirroredStrategy`, there is usually one `worker` that takes on a little more responsibility like saving checkpoint and writing summary file for TensorBoard in addition to what a regular `worker` does.
# -Such a worker is referred to as the `chief` worker, and it is customary that the `worker` with `index` 0 is appointed as the chief `worker` (in fact this is how `tf.distribute.Strategy` is implemented).
#
# `task`:
# - Provides information of the current task and is different on each worker. It specifies the `type` and `index` of that worker.
#
# Here is an example configuration:
# + colab={"background_save": true} id="XK1eTYvSZiX7"
tf_config = {
'cluster': {
'worker': ['localhost:12345', 'localhost:23456']
},
'task': {'type': 'worker', 'index': 0}
}
# + [markdown] id="JjgwJbPKZkJL"
# Here is the same `TF_CONFIG` serialized as a JSON string:
# + colab={"background_save": true, "base_uri": "https://localhost:8080/", "height": 35} id="yY-T0YDQZjbu" outputId="fd1e15b3-8e9e-4d2e-a112-422ea3390eff"
json.dumps(tf_config)
# + [markdown] id="8YFpxrcsZ2xG"
# ### Explaining the TF_CONFIG example
#
# In this example you set a `TF_CONFIG` with 2 workers on `localhost`. In practice, users would create multiple workers on external IP addresses/ports, and set `TF_CONFIG` on each worker appropriately.
#
# Since you set the task `type` to `"worker"` and the task `index` to `0`, **this machine is the first worker and will be appointed as the chief worker**.
#
# Note that other machines will need to have the `TF_CONFIG` environment variable set as well, and it should have the same `cluster` dict, but different task `type` or task `index` depending on what the roles of those machines are. For instance, for the second worker you would set `tf_config['task']['index']=1`.
#
# + [markdown] id="f83FVYqDX3aX"
# ### Quick Note on Environment variables and subprocesses in notebooks
#
# Above, `tf_config` is just a local variable in python. To actually use it to configure training, this dictionary needs to be serialized as JSON, and placed in the `TF_CONFIG` environment variable.
#
# In the next section, you'll spawn new subprocesses for each worker using the `%%bash` magic command. Subprocesses inherit environment variables from their parent, so they can access `TF_CONFIG`.
#
# You would never really launch your jobs this way (as subprocesses of an interactive Python runtime), but it's how you will do it for the purposes of this tutorial.
# + [markdown] id="UhNtHfuxCGVy"
# ## Choose the right strategy
#
# In TensorFlow there are two main forms of distributed training:
#
# * Synchronous training, where the steps of training are synced across the workers and replicas, and
# * Asynchronous training, where the training steps are not strictly synced.
#
# `MultiWorkerMirroredStrategy`, which is the recommended strategy for synchronous multi-worker training is the one you will be using.
#
# To train the model, use an instance of `tf.distribute.MultiWorkerMirroredStrategy`.
#
#
# + colab={"background_save": true} id="1uFSHCJXMrQ-"
strategy = tf.distribute.MultiWorkerMirroredStrategy()
# + [markdown] id="N0iv7SyyAohc"
# `MultiWorkerMirroredStrategy` creates copies of all variables in the model's layers on each device across all workers. It uses `CollectiveOps`, a TensorFlow op for collective communication, to aggregate gradients and keep the variables in sync. The [official TF distributed training guide](https://www.tensorflow.org/guide/distributed_training) has more details about this.
#
# + [markdown] id="H47DDcOgfzm7"
# ### Implement Distributed Training via Context Managers
#
# To distribute the training to multiple-workers all you need to do is to enclose the model building and `model.compile()` call inside `strategy.scope()`.
#
# The distribution strategy's scope dictates how and where the variables are created, and in the case of `MultiWorkerMirroredStrategy`, the variables created are `MirroredVariable`s, and they are replicated on each of the workers.
#
# + colab={"background_save": true} id="wo6b9wX65glL"
# Implementing distributed strategy via a context manager
with strategy.scope():
multi_worker_model = mnist.build_and_compile_cnn_model()
# + [markdown] id="jfYpmIxO6Jck"
# Note: `TF_CONFIG` is parsed and TensorFlow's GRPC servers are started at the time `MultiWorkerMirroredStrategy()` is called, so the `TF_CONFIG` environment variable must be set before a `tf.distribute.Strategy` instance is created.
#
# **Since `TF_CONFIG` is not set yet the above strategy is effectively single-worker training**.
#
#
# + [markdown] id="JxzYhF0dL6qQ"
# ## Train the model
#
# ### Create training script
#
# To actually run with `MultiWorkerMirroredStrategy` you'll need to run worker processes and pass a `TF_CONFIG` to them.
#
# Like the `mnist.py` file written earlier, here is the `main.py` that each of the workers will run:
# + colab={"background_save": true, "base_uri": "https://localhost:8080/"} id="BcsuBYrpgnlS" outputId="4fafa1e2-7ecf-4a7e-89f9-18b08e7b8925"
# %%writefile main.py
import os
import json
import tensorflow as tf
import mnist # Your module
# Define batch size
per_worker_batch_size = 64
# Get TF_CONFIG from the env variables and save it as JSON
tf_config = json.loads(os.environ['TF_CONFIG'])
# Infer number of workers from tf_config
num_workers = len(tf_config['cluster']['worker'])
# Define strategy
strategy = tf.distribute.MultiWorkerMirroredStrategy()
# Define global batch size
global_batch_size = per_worker_batch_size * num_workers
# Load dataset
multi_worker_dataset = mnist.mnist_dataset(global_batch_size)
# Create and compile model following the distributed strategy
with strategy.scope():
multi_worker_model = mnist.build_and_compile_cnn_model()
# Train the model
multi_worker_model.fit(multi_worker_dataset, epochs=3, steps_per_epoch=70)
# + [markdown] id="Aom9xelvJQ_6"
# In the code snippet above note that the `global_batch_size`, which gets passed to `Dataset.batch`, is set to `per_worker_batch_size * num_workers`. This ensures that each worker processes batches of `per_worker_batch_size` examples regardless of the number of workers.
# + [markdown] id="lHLhOii67Saa"
# The current directory should now contain both Python files:
# + colab={"background_save": true, "base_uri": "https://localhost:8080/"} id="bi6x05Sr60O9" outputId="f798f814-588a-4eb2-df68-b93858fd3810"
# !ls *.py
# + [markdown] id="qmEEStPS6vR_"
# ### Set TF_CONFIG environment variable
#
# Now json-serialize the `TF_CONFIG` and add it to the environment variables:
# + colab={"background_save": true} id="9uu3g7vV7Bbt"
# Set TF_CONFIG env variable
os.environ['TF_CONFIG'] = json.dumps(tf_config)
# + [markdown] id="-WDqwMPNneON"
# And terminate all background processes:
# + colab={"background_save": true, "base_uri": "https://localhost:8080/"} id="txMXaq8d8N_S" outputId="11ba6b9f-de12-408e-f043-4373359e5aff"
# first kill any previous runs
# %killbgscripts
# + [markdown] id="MsY3dQLK7jdf"
# ### Launch the first worker
#
# Now, you can launch a worker process that will run the `main.py` and use the `TF_CONFIG`:
# + colab={"background_save": true, "base_uri": "https://localhost:8080/"} id="qnSma_Ck7r-r" outputId="4728c637-3f04-482e-bfaf-210f7f923ad9" magic_args="--bg" language="bash"
# python main.py &> job_0.log
# + [markdown] id="ZChyazqS7v0P"
# There are a few things to note about the above command:
#
# 1. It uses the `%%bash` which is a [notebook "magic"](https://ipython.readthedocs.io/en/stable/interactive/magics.html) to run some bash commands.
# 2. It uses the `--bg` flag to run the `bash` process in the background, because this worker will not terminate. It waits for all the workers before it starts.
#
# The backgrounded worker process won't print output to this notebook, so the `&>` redirects its output to a file, so you can see what happened.
#
# So, wait a few seconds for the process to start up:
# + colab={"background_save": true} id="Hm2yrULE9281"
# Wait for logs to be written to the file
time.sleep(10)
# + [markdown] id="ZFPoNxg_9_Mx"
# Now look what's been output to the worker's logfile so far using the `cat` command:
# + colab={"background_save": true, "base_uri": "https://localhost:8080/"} id="vZEOuVgQ9-hn" outputId="3b64814e-faa8-43f7-d101-20e20b0c5636" language="bash"
# cat job_0.log
# + [markdown] id="RqZhVF7L_KOy"
# The last line of the log file should say: `Started server with target: grpc://localhost:12345`. The first worker is now ready, and is waiting for all the other worker(s) to be ready to proceed.
# + [markdown] id="Pi8vPNNA_l4a"
# ### Launch the second worker
#
# Now update the `tf_config` for the second worker's process to pick up:
# + colab={"background_save": true} id="lAiYkkPu_Jqd"
tf_config['task']['index'] = 1
os.environ['TF_CONFIG'] = json.dumps(tf_config)
# + [markdown] id="0AshGVO0_x0w"
# Now launch the second worker. This will start the training since all the workers are active (so there's no need to background this process):
# + colab={"background_save": true, "base_uri": "https://localhost:8080/"} id="_ESVtyQ9_xjx" outputId="9a3413b2-8c6d-405f-922e-3468fe57e190" language="bash"
# python main.py
# + [markdown] id="hX4FA2O2AuAn"
# Now if you recheck the logs written by the first worker you'll see that it participated in training that model:
# + colab={"background_save": true, "base_uri": "https://localhost:8080/"} id="rc6hw3yTBKXX" outputId="09e56ea2-5994-46d8-f3d4-07857816ca42" language="bash"
# cat job_0.log
# + [markdown] id="zL79ak5PMzEg"
# Unsurprisingly this ran _slower_ than the the test run at the beginning of this tutorial. **Running multiple workers on a single machine only adds overhead**. The goal here was not to improve the training time, but only to give an example of multi-worker training.
# + [markdown] id="xckY28bOV_p8"
# -----------------------------
# **Congratulations on finishing this ungraded lab!** Now you should have a clearer understanding of how to implement distributed strategies with Tensorflow and Keras.
#
# Although this tutorial didn't show the true power of a distributed strategy since this will require multiple machines operating under the same network, you now know how this process looks like at a high level.
#
# In practice and especially with very big models, distributed strategies are commonly used as they provide a way of better managing resources to perform time-consuming tasks, such as training in a fraction of the time that it will take without the strategy.
#
# **Keep it up!**
| 3_Machine Learning Modeling Pipelines in Production/Week3/C3_W3_Lab_1_Distributed_Training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="HW777fm6jEK2"
# # **CNN for multi-class classification**
# + id="rGMN4kwwjJXw"
import keras,os
from keras.models import Sequential
from keras.layers import Dense, Conv2D, MaxPool2D , Flatten
from keras.preprocessing.image import ImageDataGenerator
import numpy as np
# + colab={"base_uri": "https://localhost:8080/"} id="20GjyE6WQOz7" outputId="f07e2563-3b0d-4c90-a9c6-b36de4e85a23"
from google.colab import drive
drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/"} id="yk7GEeTmjmbX" outputId="079d7056-1179-44e9-8ec5-91cd9c6218fb"
dir = 'drive/MyDrive/Fish_Dataset'
image_generator = ImageDataGenerator(rescale=1/255, validation_split=0.2)
train_dataset = image_generator.flow_from_directory(batch_size=32,
directory=dir,
shuffle=True,
target_size=(224, 224),
subset="training",
class_mode='categorical')
validation_dataset = image_generator.flow_from_directory(batch_size=32,
directory=dir,
shuffle=True,
target_size=(224, 224),
subset="validation",
class_mode='categorical')
# + [markdown] id="egvxPCSmKdXz"
# ### VGG16
# + id="ie2B9Jn7kwLl"
# model = Sequential()
# model.add(Conv2D(input_shape=(224,224,3),filters=64,kernel_size=(3,3),padding="same", activation="relu"))
# model.add(Conv2D(filters=32,kernel_size=(3,3),padding="same", activation="relu"))
# model.add(MaxPool2D(pool_size=(2,2),strides=(2,2)))
# model.add(Conv2D(filters=32, kernel_size=(3,3), padding="same", activation="relu"))
# model.add(Conv2D(filters=32, kernel_size=(3,3), padding="same", activation="relu"))
# model.add(MaxPool2D(pool_size=(2,2),strides=(2,2)))
# model.add(Conv2D(filters=32, kernel_size=(3,3), padding="same", activation="relu"))
# model.add(Conv2D(filters=32, kernel_size=(3,3), padding="same", activation="relu"))
# model.add(Conv2D(filters=32, kernel_size=(3,3), padding="same", activation="relu"))
# model.add(MaxPool2D(pool_size=(2,2),strides=(2,2)))
# model.add(Conv2D(filters=32, kernel_size=(3,3), padding="same", activation="relu"))
# model.add(Conv2D(filters=32, kernel_size=(3,3), padding="same", activation="relu"))
# model.add(Conv2D(filters=32, kernel_size=(3,3), padding="same", activation="relu"))
# model.add(MaxPool2D(pool_size=(2,2),strides=(2,2)))
# model.add(Conv2D(filters=32, kernel_size=(3,3), padding="same", activation="relu"))
# model.add(Conv2D(filters=32, kernel_size=(3,3), padding="same", activation="relu"))
# model.add(Conv2D(filters=32, kernel_size=(3,3), padding="same", activation="relu"))
# model.add(MaxPool2D(pool_size=(2,2),strides=(2,2)))
# + id="-xIUjMlvlWWc"
# model.add(Flatten())
# model.add(Dense(units=4096,activation="relu"))
# model.add(Dense(units=4096,activation="relu"))
# model.add(Dense(units=9, activation="softmax"))
# + [markdown] id="FehD4IriomXi"
# ### Custom CNN Architecture
# + id="QtIExbW4olYb"
model = Sequential()
model.add(Conv2D(input_shape=(224,224,3),filters=64,kernel_size=(3,3),padding="same", activation="relu"))
model.add(Conv2D(filters=32,kernel_size=(3,3),padding="same", activation="relu"))
model.add(MaxPool2D(pool_size=(2,2),strides=(2,2)))
model.add(Conv2D(filters=32, kernel_size=(3,3), padding="same", activation="relu"))
model.add(Conv2D(filters=32, kernel_size=(3,3), padding="same", activation="relu"))
model.add(MaxPool2D(pool_size=(2,2),strides=(2,2)))
model.add(Flatten())
model.add(Dense(units=9, activation="softmax"))
# + id="-iFpX32nlikT" colab={"base_uri": "https://localhost:8080/"} outputId="87bddc1c-de91-4fca-a7f7-fdc16609aec8"
from keras.optimizers import Adam
opt = Adam(lr=0.001)
model.compile(optimizer=opt, loss=keras.losses.categorical_crossentropy, metrics=['accuracy'])
# + id="1eoQBQzFll-D" colab={"base_uri": "https://localhost:8080/"} outputId="099374bb-3a4c-43c2-8e10-88a822f310b3"
model.summary()
# + id="vhbmdY6Almog" colab={"base_uri": "https://localhost:8080/"} outputId="345e6880-2fb4-42bf-dbcd-ba9e82457c86"
from keras.callbacks import ModelCheckpoint, EarlyStopping
checkpoint = ModelCheckpoint("my_model.h5", monitor='val_acc', verbose=1, save_best_only=True, save_weights_only=False, mode='auto', period=1)
early = EarlyStopping(monitor='val_acc', min_delta=0, patience=20, verbose=1, mode='auto')
hist = model.fit_generator(steps_per_epoch=100,generator=train_dataset, validation_data= validation_dataset, validation_steps=20, epochs=25 , callbacks=[checkpoint,early])
# + id="Ye2I7FW7lz3A"
model.save('my_model.h5')
model.save('drive/MyDrive/Models/my_model.h5')
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="yEonDnBD1v3A" outputId="02f6db97-372e-4cf3-f85c-90827bb50401"
import matplotlib.pyplot as plt
plt.plot(hist.history["accuracy"])
plt.plot(hist.history['loss'])
plt.title("Train data accuracy and loss")
plt.ylabel("Accuracy")
plt.xlabel("Epoch")
plt.legend(["Accuracy","loss"])
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="c1r0pvkdTNs4" outputId="11ba7b5c-eb91-4850-def6-e10243dce5da"
plt.plot(hist.history['val_accuracy'])
plt.plot(hist.history['val_loss'])
plt.title("Test data accuracy and loss")
plt.ylabel("Accuracy")
plt.xlabel("Epoch")
plt.legend(["Validation Accuracy","Validation Loss"])
plt.show()
# + id="wAkryqrbl3-e" colab={"base_uri": "https://localhost:8080/", "height": 302} outputId="d7bc1cdd-23fd-4b68-f1a1-93d5bf8c67c9"
from keras.preprocessing import image
from keras.models import load_model
import random as rd
i = rd.randint(0, len(validation_dataset.filenames))
path = dir + "/" + validation_dataset.filenames[i]
print("Input: ",validation_dataset.filenames[i].split('/')[0])
img = image.load_img(path,target_size=(224,224))
img = np.asarray(img)
plt.imshow(img)
img = np.expand_dims(img, axis=0)
class_names = ['Black_sea_sprat', 'Gilt-Head-Bream', 'Horse Mackrel', 'Red Mullet', 'Red Sea Bream', 'Sea Bass', 'Shrimp', 'Striped Red Mullet', 'Trout']
saved_model = load_model("my_model.h5")
output = saved_model.predict(img)
ind = np.argmax(output[0])
print("Prediction: ", class_names[ind])
# + id="B6Yl5MXL4gI2"
| Problem2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Create dynamic table for Top 50 interactions for COAD
import ipywidgets as widgets
from ipywidgets import interact, interact_manual
import pandas as pd
import numpy as np
# get data
database = "../db/dbCOAD-DRD.csv"
df = pd.read_csv(database)
df
# use other order of columns
df_repurposing = df[['AE', 'HGNC_symbol', 'DrugName', 'ProteinID', 'DrugCID', 'Drug']]
df_repurposing
# saving as HTML file the same result
fout = open("../extras/full_table.html","w")
fout.write(df_repurposing.to_html(index=False))
fout.close()
# counting the elements
print('No of genes:', len(list(set(df_repurposing['HGNC_symbol']))))
print('No of PDBs:', len(list(set(df_repurposing['ProteinID']))))
print('No of drug names:', len(list(set(df_repurposing['DrugName']))))
print('No of drug compounds:', len(list(set(df_repurposing['DrugCID']))))
| COAD-DRD/scripts/full_Table.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="2EU2eYXohNR5" colab_type="text"
# ## Ribbon CNN Training Notebook
#
# This notebook contains the code to train a simple CNN to classify different types of flare ribbons visible in 1600Å AIA (*Atmospheric Imaging Assembly*) images.
#
# To use this notebook the following packages are needed:
#
# (To train the network)
# 1. numpy
#
# 2. pickle (for reading in training data)
#
# 3. keras
#
# 4. scikit-learn
#
# (To create training/test plots)
# 5. matplotlib
#
# 6. pandas
#
# 7. seaborn
#
# Note that with the training data included in *4class_data.pickle* a GPU is currently not needed to train this model, however if the amount of data is increased this would have to change.
#
# The parameters chosen (epochs, batch size etc) are selected to optimize the network performance that corresponds to the training set, again if the training set is altered these may also have to be modified.
#
# Note that in this notebook k-fold cross-validation has also been implemented (where k = 5), this is to ensure a more vigorus training of the model with varying validation sets used throughout training. For more information on cross validation please see [here](https://machinelearningmastery.com/k-fold-cross-validation/).
#
# + id="7Jj7Xu2Wq5Bh" colab_type="code" colab={}
#training packages
import numpy as np
import pickle
from keras.utils import to_categorical
from sklearn.utils import shuffle
from sklearn.model_selection import KFold
from keras.models import Sequential, load_model
from keras.layers import Convolution2D, MaxPooling2D, Flatten, Dense, Dropout
from keras import optimizers
#plotting packages
import seaborn as sn
import pandas as pd
import matplotlib.pyplot as plt
from scipy import interp
from itertools import cycle
import sklearn
# + [markdown] id="kq75hST_m9qE" colab_type="text"
# We will initially read in the training and test data, with the model parameters also defined.
# + id="Xj6miZGGrQqE" colab_type="code" colab={}
#Import data
with open('4class_data_training.pickle','rb') as t:
input_train, target_train = pickle.load(t)
with open('4class_data_test.pickle','rb') as t:
input_test, target_test = pickle.load(t)
# + id="ff6p8G28rso1" colab_type="code" colab={}
# Model configuration
batch_size = 32
img_width, img_height, img_num_channels = 250, 250, 1
no_classes = 4
no_epochs = 10
validation_split = 0.4
verbosity = 1
num_folds = 5
input_shape = (img_width, img_height, img_num_channels)
# + id="pzc5yVF-ryGd" colab_type="code" colab={}
# Define per-fold acc/loss lists
acc_per_fold = []
loss_per_fold = []
histories =[]
model_history = []
#Define training set
inputs = input_train
targets = target_train
# + [markdown] id="RZvVGGA_ns8q" colab_type="text"
# K-fold cross-validation implemented below (the model code can be extracted from here is cross-validation is too computationally expensive).
# + id="LPko-ETrsDw_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="47996cc4-22a2-4126-b297-<KEY>"
# Define the K-fold Cross Validator
kfold = KFold(n_splits=num_folds, shuffle=True)
# K-fold Cross Validation model evaluation
fold_no = 1
for train, test in kfold.split(inputs, targets):
# Define the model architecture
model = Sequential()
model.add(Convolution2D(32, kernel_size=(3, 3), activation='relu', input_shape=input_shape))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Convolution2D(64, kernel_size=(3, 3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Flatten())
model.add(Dropout(0.5))
model.add(Dense(128, activation='relu'))
model.add(Dense(4, activation='softmax'))
sgd = optimizers.sgd(lr=0.001, clipvalue=0.5)
model.compile(loss='categorical_crossentropy', optimizer=sgd, metrics=['accuracy'])
# Fit data to model
history = model.fit(inputs[train], targets[train], batch_size=batch_size, epochs=no_epochs, verbose=verbosity, validation_split=validation_split)
histories.append(history)
model_history.append(model)
# Generate generalization metrics
scores = model.evaluate(inputs[test], targets[test], verbose=0)
print(f'Score for fold {fold_no}: {model.metrics_names[0]} of {scores[0]}; {model.metrics_names[1]} of {scores[1]*100}%')
acc_per_fold.append(scores[1] * 100)
loss_per_fold.append(scores[0])
# Increase fold number
fold_no = fold_no + 1
# == Provide average scores ==
print('------------------------------')
print('Score per fold')
for i in range(0, len(acc_per_fold)):
print('------------------------------')
print(f'> Fold {i+1} - Loss: {loss_per_fold[i]} - Accuracy: {acc_per_fold[i]}%')
print('------------------------------')
print('Average scores for all folds:')
print(f'> Accuracy: {np.mean(acc_per_fold)} (+- {np.std(acc_per_fold)})')
print(f'> Loss: {np.mean(loss_per_fold)}')
print('-------------------------------')
# + [markdown] id="jx3ZdXp4oA7r" colab_type="text"
# The model is now sufficently trained - the plotting routines listed below are just some examples of how the model can be tested and results plotted.
#
# First, the results from the cross-validation are plotted.
# + id="Nqc_fOo6pMXh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 383} outputId="da0f14b9-0157-44fa-f323-11ed4223fd50"
for i in range(len(histories)):
# plot loss
plt.subplot(211)
plt.title('Cross Entropy Loss')
plt.plot(range(1, 11), histories[i].history['loss'], color='blue', label='train')
plt.plot(range(1, 11), histories[i].history['val_loss'],color='orange', label='validation')
plt.xlabel('Epoch')
plt.ylabel('Loss')
if i==0:
plt.legend()
# plot accuracy
plt.subplot(212)
plt.title('Classification Accuracy')
plt.plot(range(1, 11), histories[i].history['accuracy'], color='blue', label='train')
plt.plot(range(1, 11), histories[i].history['val_accuracy'], color='orange', label='validation')
plt.xlabel('Epoch')
plt.ylabel('Accuracy')
if i ==0:
plt.legend()
plt.subplots_adjust(hspace = 0.6)
# + [markdown] id="9oLyjJyVufPm" colab_type="text"
# A confusion matrix using the test data set is created below.
# + id="T1PfopfJt69j" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 312} outputId="12f7a1b1-e4b5-4b44-8a40-0003bd2a9f95"
#test trained model.
testout = model.predict(input_test)
matrix = sklearn.metrics.confusion_matrix(testout.argmax(axis=1), np.array(target_test))
normmatrix = matrix.astype('float') / matrix.sum(axis=1)[:, np.newaxis]
classes = ['background', '2 ribbon', 'limb', 'compact']
df_cm = pd.DataFrame(normmatrix, index = classes,columns = classes)
ax= plt.subplot()
sn.heatmap(df_cm, annot=True)
ax.set_ylim(len(matrix), -0.5)
plt.title('Confusion Matrix')
plt.xlabel('Predicted Class')
plt.ylabel('True Class')
| CNN_k_fold.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import s3fs
import numpy as np
import pandas as pd
from pandas import set_option
from matplotlib import pyplot as plt
from pandas import read_csv
#view data (Providers_Updated_430.txt)
df = pd.read_csv('s3://filtered-datasets/Providers_Updated_430.txt', sep='\t')
from pandas import set_option
set_option('display.max_columns', None)
df.head(5)
df.shape
#display null value counts for each column
pd.set_option('display.max_rows', 120)
df.isnull().sum()
# +
# imputation for total_claim_count_ge65
# total_claim_count_ge65 (If ge65_suppress_flag = *=>total_claim_count_ge65 = 5)
df['total_claim_count_ge65'] = np.where((df['ge65_suppress_flag'] == '*'), 5, df['total_claim_count_ge65'])
#If ge65_suppress_flag = #=>total_claim_count_ge65 = total_claim_count – 5
df['total_claim_count_ge65'] = np.where((df['ge65_suppress_flag'] == '#'), df['total_claim_count'] - 5,
df['total_claim_count_ge65'])
df.head(5)
# +
# imputation for bene_count_ge65
# Bene_count_ge65 ( If bene_count_ge65_suppress_flag = * =>bene_count_ge65 = 5)
df['bene_count_ge65'] = np.where((df['bene_count_ge65_suppress_flag'] == '*'), 5, df['bene_count_ge65'])
# If bene_count_ge65_suppress_flag = # =>bene_count_ge65 = total_bene_count – 5
df['bene_count_ge65'] = np.where((df['bene_count_ge65_suppress_flag'] == '#'), df['bene_count'] - 5,
df['bene_count_ge65'])
df.head(5)
# +
# imputations for brand claim count,generic_claim_count, and other claim count
# brand claim count (If brand_suppress_flag = *=>brand_claim_count = 5)
df['brand_claim_count'] = np.where((df['brand_suppress_flag'] == '*'), 5, df['brand_claim_count'])
# generic_claim_count (If generic_suppress_flag = *=>generic_claim_count = 5)
df['generic_claim_count'] = np.where((df['generic_suppress_flag'] == '*'), 5, df['generic_claim_count'])
# other_claim_count(If other_suppress_flag = *=>other_claim_count = 5)
df['other_claim_count'] = np.where((df['other_suppress_flag'] == '*'), 5, df['other_claim_count'])
# brand_claim_count(If brand_suppress_flag = #=>brand_claim_count = total_claim_count - other_claim_count – generic_claim_count)
df['brand_claim_count'] = np.where((df['brand_suppress_flag'] == '#'), df['total_claim_count'] - df['other_claim_count'] -df['generic_claim_count'],
df['brand_claim_count'])
# generic claim count (If generic_suppress_flag = #=>generic_claim_count = total_claim_count – brand_claim_count – other_claim_count)
df['generic_claim_count'] = np.where((df['generic_suppress_flag'] == '#'), df['total_claim_count'] - df['brand_claim_count'] -df['other_claim_count'],
df['generic_claim_count'])
# other claim count (If other_suppress_flag = #=>other_claim_count = total_claim_count – brand_claim_count – generic_claim_count)
df['other_claim_count'] = np.where((df['other_suppress_flag'] == '#'), df['total_claim_count'] - df['brand_claim_count'] -df['generic_claim_count'],
df['other_claim_count'])
df.head(5)
# +
# imputations for mapd claim count and Pdp_claim_count
# mapd claim count (If mapd_suppress_flag = *=>Mapd_claim_count = 5)
df['mapd_claim_count'] = np.where((df['mapd_suppress_flag'] == '*'), 5, df['mapd_claim_count'])
# Pdp_claim_count (If pdp_suppress_flag = *=>Pdp_claim_count = 5)
df['pdp_claim_count'] = np.where((df['pdp_suppress_flag'] == '*'), 5, df['pdp_claim_count'])
# mapd claim count (If mapd_suppress_flag = #=>Mapd_claim_count = total_claim_count – pdp_claim_count)
df['mapd_claim_count'] = np.where((df['mapd_suppress_flag'] == '#'), df['total_claim_count'] - df['pdp_claim_count'],
df['mapd_claim_count'])
# pdp claim count (If pdp_suppress_flag = #=>Pdp_suppress_flag = total_claim_count – mapd_claim_count)
df['pdp_claim_count'] = np.where((df['pdp_suppress_flag'] == '#'), df['total_claim_count'] - df['mapd_claim_count'],
df['pdp_claim_count'])
df.head(10)
# +
# imputations for lis claim count and nonlis claim count
# lis claim count (If lis_suppress_flag = * =>lis_claim_count = 5)
df['lis_claim_count'] = np.where((df['lis_suppress_flag'] == '*'), 5, df['lis_claim_count'])
# nonlis claim count (If nonlis_suppress_flag = *=>ononlis_claim_count = 5)
df['nonlis_claim_count'] = np.where((df['nonlis_suppress_flag'] == '*'), 5, df['nonlis_claim_count'])
# lis claim count if lis_suppress_flag = #=> lis_claim_count = total_claim_count – non_claim_count
# non_claim_count is missing in the data
#imputing remaining null values in lis claim count with the median for the lis claim count column
df['lis_claim_count']=df['lis_claim_count'].fillna(df['lis_claim_count'].median())
# nonlis claim count (If nonlis_suppress_flag = # =>non_suppress_flag = total_claim_count – lis_claim_count)
df['nonlis_claim_count'] = np.where((df['nonlis_suppress_flag'] == '#'), df['total_claim_count'] - df['lis_claim_count'],
df['nonlis_claim_count'])
df.head(15)
# +
# imputations for antipsych claim count ge65
# antipsych claim count ge65 (If antipsych_ge65_suppress_flag = *=>antipsych_claim_count_ge65 = 5)
df['antipsych_claim_count_ge65'] = np.where((df['antipsych_ge65_suppress_flag'] == '*'), 5, df['antipsych_claim_count_ge65'])
# antipsych claim count ge65 (If antipsych_ge65_suppress_flag = #)
#impute remaining null values with the median for the antipsych claim count ge65
df['antipsych_claim_count_ge65']=df['antipsych_claim_count_ge65'].fillna(df['antipsych_claim_count_ge65'].median())
df.head(10)
# +
# imputations for antipsych_bene_count_ge65
# antipsych_bene_count_ge65 (If antipsych_bene_ge65_suppress_flag = * =>antipsych_bene_count_ge65 = 5)
df['antipsych_bene_count_ge65'] = np.where((df['antipsych_bene_ge65_suppress_flg'] == '*'), 5, df['antipsych_bene_count_ge65'])
# antipsych_bene_count_ge65 (If antipsych_bene_ge65_suppress_flag = #)
# impute remaining null values with the median for the antipsych_bene_count_ge65 column
df['antipsych_bene_count_ge65']=df['antipsych_bene_count_ge65'].fillna(df['antipsych_bene_count_ge65'].median())
df.head(10)
# -
#display all remaining null value counts
pd.set_option('display.max_rows', 120)
df.isnull().sum()
from pandas import set_option
set_option('display.max_columns', None)
df.head(10)
# +
# impute other null values
# total_30_day_fill_count_ge65
df['total_30_day_fill_count_ge65']=df['total_30_day_fill_count_ge65'].fillna(df['total_30_day_fill_count_ge65'].median())
# bene_count
df['bene_count']=df['bene_count'].fillna(df['bene_count'].median())
# total_drug_cost_ge65
df['total_drug_cost_ge65']=df['total_drug_cost_ge65'].fillna(df['total_drug_cost_ge65'].median())
# total_day_supply_ge65
df['total_day_supply_ge65']=df['total_day_supply_ge65'].fillna(df['total_day_supply_ge65'].median())
# bene_count_ge65
df['bene_count_ge65']=df['bene_count_ge65'].fillna(df['bene_count_ge65'].median())
# brand_drug_cost
df['brand_drug_cost']=df['brand_drug_cost'].fillna(df['brand_drug_cost'].median())
# generic_drug_cost
df['generic_drug_cost']=df['generic_drug_cost'].fillna(df['generic_drug_cost'].median())
# other_drug_cost
df['other_drug_cost']=df['other_drug_cost'].fillna(df['other_drug_cost'].median())
# mapd_drug_cost
df['mapd_drug_cost']=df['mapd_drug_cost'].fillna(df['mapd_drug_cost'].median())
# pdp_drug_cost
df['pdp_drug_cost']=df['pdp_drug_cost'].fillna(df['pdp_drug_cost'].median())
# lis_claim_count
df['lis_claim_count']=df['lis_claim_count'].fillna(df['lis_claim_count'].median())
# lis_drug_cost
df['lis_drug_cost']=df['lis_drug_cost'].fillna(df['lis_drug_cost'].median())
# nonlis_claim_count
df['nonlis_claim_count']=df['nonlis_claim_count'].fillna(df['nonlis_claim_count'].median())
# nonlis_drug_cost
df['nonlis_drug_cost']=df['nonlis_drug_cost'].fillna(df['nonlis_drug_cost'].median())
# opioid_claim_count
df['opioid_claim_count']=df['opioid_claim_count'].fillna(df['opioid_claim_count'].median())
# opioid_drug_cost
df['opioid_drug_cost']=df['opioid_drug_cost'].fillna(df['opioid_drug_cost'].median())
# opioid_day_supply
df['opioid_day_supply']=df['opioid_day_supply'].fillna(df['opioid_day_supply'].median())
# opioid_bene_count
df['opioid_bene_count']=df['opioid_bene_count'].fillna(df['opioid_bene_count'].median())
# opioid_prescriber_rate
df['opioid_prescriber_rate']=df['opioid_prescriber_rate'].fillna(df['opioid_prescriber_rate'].median())
# la_opioid_claim_count
df['la_opioid_claim_count']=df['la_opioid_claim_count'].fillna(df['la_opioid_claim_count'].median())
# la_opioid_drug_cost
df['la_opioid_drug_cost']=df['la_opioid_drug_cost'].fillna(df['la_opioid_drug_cost'].median())
# la_opioid_day_supply
df['la_opioid_day_supply']=df['la_opioid_day_supply'].fillna(df['la_opioid_day_supply'].median())
# la_opioid_bene_count
df['la_opioid_bene_count']=df['la_opioid_bene_count'].fillna(df['la_opioid_bene_count'].median())
# la_opioid_prescriber_rate
df['la_opioid_prescriber_rate']=df['la_opioid_prescriber_rate'].fillna(df['la_opioid_prescriber_rate'].median())
# antibiotic_claim_count
df['antibiotic_claim_count']=df['antibiotic_claim_count'].fillna(df['antibiotic_claim_count'].median())
# antibiotic_drug_cost
df['antibiotic_drug_cost']=df['antibiotic_drug_cost'].fillna(df['antibiotic_drug_cost'].median())
# antibiotic_bene_count
df['antibiotic_bene_count']=df['antibiotic_bene_count'].fillna(df['antibiotic_bene_count'].median())
# antipsych_drug_cost_ge65
df['antipsych_drug_cost_ge65']=df['antipsych_drug_cost_ge65'].fillna(df['antipsych_drug_cost_ge65'].median())
# average_age_of_beneficiaries
df['average_age_of_beneficiaries']=df['average_age_of_beneficiaries'].fillna(df['average_age_of_beneficiaries'].median())
# beneficiary_female_count
df['beneficiary_female_count']=df['beneficiary_female_count'].fillna(df['beneficiary_female_count'].median())
# beneficiary_male_count
df['beneficiary_male_count']=df['beneficiary_male_count'].fillna(df['beneficiary_male_count'].median())
# beneficiary_nondual_count
df['beneficiary_nondual_count']=df['beneficiary_nondual_count'].fillna(df['beneficiary_nondual_count'].median())
# beneficiary_dual_count
df['beneficiary_dual_count']=df['beneficiary_dual_count'].fillna(df['beneficiary_dual_count'].median())
# beneficiary_average_risk_score
df['beneficiary_average_risk_score']=df['beneficiary_average_risk_score'].fillna(df['beneficiary_average_risk_score'].median())
# -
#display all remaining null value counts
pd.set_option('display.max_rows', 120)
df.isnull().sum()
# convert EXCL Year and Excl type to objects
df["EXCLYear"]= df["EXCLYear"].astype(object)
#convert nppes provider zip to object
df["REINYear"]= df["REINYear"].astype(object)
# Further analysis will be performed to ascertain needed further processing including transformations or column (atrribute) dropping.
| Cleaning_and_Merging/Providers_updated_430_txt null value imputations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 정규 표현식 살펴보기
#
# 정규 표현식(Regular Expressions)은 복잡한 문자열을 처리할 때 사용하는 기법으로, 파이썬만의 고유 문법이 아니라 문자열을 처리하는 모든 곳에서 사용한다. 정규 표현식을 배우는 것은 파이썬을 배우는 것과는 또 다른 영역의 과제이다.
#
# ※ 정규 표현식은 줄여서 간단히 "정규식"이라고도 말한다.
# ## 정규 표현식은 왜 필요한가?
#
#
# 다음과 같은 문제가 주어졌다고 가정해 보자.
#
# - 주민등록번호를 포함하고 있는 텍스트가 있다. 이 텍스트에 포함된 모든 주민등록번호의 뒷자리를 * 문자로 변경해 보자.
#
# 우선 정규식을 전혀 모르면 다음과 같은 순서로 프로그램을 작성해야 할 것이다.
#
# 1. 전체 텍스트를 공백 문자로 나눈다(split).
# 2. 나뉜 단어가 주민등록번호 형식인지 조사한다.
# 3. 단어가 주민등록번호 형식이라면 뒷자리를 *로 변환한다.
# 4. 나뉜 단어를 다시 조립한다.
# 5. 이를 구현한 코드는 아마도 다음과 같을 것이다.
data="""
park 800905-1049118
kim 700905-1059119
"""
result=[]
# +
for line in data.split("\n"): # data의 데이터를 line별로 split 해서 line에 저장.
word_result=[]
for word in line.split(" "): # line에 저장된 데이터 하나 씩을 공백을 기준으로 word에 저장.
if len(word)==14 and word[:6].isdigit( ) and word[7:].isdigit():
# 문자열 형태이더라도 숫자가 저장된 word[:6]과 word[7:]의 경우에 isdigit에서 True값이 리턴
# word[:6].isdigit()은
# 문자열 형태에 알파벳이 저장된 경우에는 isalpha 함수에서 True 가 리턴
# 따라서 if문은 word중 14자리인 주민번호data이면서 숫자로이루어진 word[:6]과 word[7:]인 경우에 실행.
word=word[:6] + '-' +'*******' # 해당 word에 word[:6]만 그대로 넣어주고 뒤는 -와 *로 저장해줌.
word_result.append(word) # 이를 word_result에 확장하여 넣어줌.
result.append(" ".join(word_result))
print("\n".join(result))
#print(result)
# list = str.split() : 문자열 => 리스트, 공백시 스페이스 기준
# ” ".join( list ) : 리스트에서 문자열으로 index마다 공백을 줌.
# -
# 결과값:
# park 800905-*******
# kim 700905-*******
# - 반면에 정규식을 사용하면 다음처럼 훨씬 간편하고 직관적인 코드를 작성할 수 있다. 아직 정규식 사용 방법을 배우지 않았으니 눈으로만 살펴보자.
# +
import re
data = """
park 800905-1049118
kim 700905-1059119
"""
pat = re.compile("(\d{6})[-]\d{7}")
print(pat.sub("\g<1>-*******", data))
# -
# 정규 표현식을 사용하면 이렇게 간단한 예제에서도 코드가 상당히 간결해진다.
# 만약 찾으려는 문자열 또는 바꾸어야 할 문자열의 규칙이 매우 복잡하다면 정규식의 효용은 더 커지게 된다.
#
# 이제부터 정규 표현식의 기초부터 심화 부분까지 차근차근 알아보자.
| JumptoPython/Part_7_Regular Expression(Regex)/07-1_Regular_Expressions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/navidroo/image_classification/blob/master/Untitled0.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="JM9iY8lcKuWs" colab_type="code" outputId="7ebaf73b-b00e-4771-f694-4d8ac50ff620" colab={"base_uri": "https://localhost:8080/", "height": 158}
pip install keras
# + [markdown] id="GFCzr8hHmize" colab_type="toc"
#
# + id="Q9bLq1JeK4-u" colab_type="code" colab={}
from keras.datasets import cifar10
# + id="nH5n-IVZKwzf" colab_type="code" colab={}
(x_train,y_train), (x_test,y_test) = cifar10.load_data()
# + id="z8AnTbjBK2nm" colab_type="code" outputId="4c656abf-ade6-481e-eedc-72813f32ea9f" colab={"base_uri": "https://localhost:8080/", "height": 34}
str_pri = '(x_train,y_train), (x_test,y_test)'.replace('(','').replace(')','').replace(" ",'').split(',')
str_pri
# + id="yVm08IuQLVRK" colab_type="code" outputId="f4bb4edd-1f25-49f5-9f14-ed412778478d" colab={"base_uri": "https://localhost:8080/", "height": 158}
for i in str_pri:
print(i)
print(type(globals()[i]))
# + id="hwmBBq8FLhml" colab_type="code" outputId="6d0d5f63-4d6b-41d2-b3fb-54083ecd1a15" colab={"base_uri": "https://localhost:8080/", "height": 87}
for i in str_pri:
print(f'{i} shape:',globals()[i].shape)
# + id="85QQL_2tN299" colab_type="code" outputId="9bd39ff6-ac83-4508-ec2b-e224dadb0c75" colab={"base_uri": "https://localhost:8080/", "height": 879}
x_train[0]
# + id="IqklcM-gOfoU" colab_type="code" colab={}
import matplotlib.pyplot as plt
# + id="cQVH3fm2OnaL" colab_type="code" outputId="ed22ba7e-0599-4d1f-b65a-f3e10a453273" colab={"base_uri": "https://localhost:8080/", "height": 284}
plt.imshow(x_train[0])
# + id="JI7ok-PVOqHD" colab_type="code" colab={}
| Untitled0.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <div style="color:#303030;font-family:'arial blACK', sans-serif,monospace; text-align: center; padding: 50px 0; vertical-align:middle;" > <img src="https://github.com/PIA-Group/ScientIST-notebooks/blob/master/_Resources/Images/Lightbulb.png?raw=true" style=" background:linear-gradient(to right,#FDC86E,#fbb144);border-radius:10px;width:150px;text-align:left; margin-left:10%" /> <span style="position:relative; bottom:70px; margin-left:5%;font-size:170%;"> Digital to Analog Converter - DAC </span> </div>
# ## <span style="color:#fbb144;"> Keywords: </span>
# ```Digital to Analog Converter (DAC)```, ```Digital Signal Processing (DSP)```, ```Arduino```
# # I. Introduction
# <br>
# <div class="title"style="width:100%; background:linear-gradient(to right,#FDC86E,#fbb144);font-family:'arial black',monospace; text-align: center; padding: 7px 0; border-radius: 5px 50px;margin-top:-15px" > </div>
# ## <div style="color:#fbb144"> 1. Background </div>
# ...
# ## <div style="color:#fbb144"> 2. Objectives</div>
# * Objetive 1 ...
# * Objetive 2 ...
# * ...
# ## <div style="color:#fbb144"> 3. Materials (optional) </div>
# * Material 1 ...
# * Material 2 ...
# * ...
# # II. Experimental
# <br>
# <div style="width:100%; background:linear-gradient(to right,#FDC86E,#fbb144);color:#282828;font-family:'arial black'; text-align: center; padding: 7px 0; border-radius: 5px 50px; margin-top:-15px" > </div>
# ### <div style="color:#fbb144"> 1. Include a Digital to Anaolg Converter (DAC TDA8702) in the Digital Signal Processing (DSP) system </div>
# Structure examples of DACs:
#
# <img src="https://github.com/PIA-Group/ScientIST-notebooks/blob/master/_Resources/Images/A.Signal_Acquisition_IMG/a010/DAC.png?raw=true" width="500" border="0">
# ### <div style="color:#fbb144"> 2. Generate the following analog output wave </div>
#
# <img src="https://github.com/PIA-Group/ScientIST-notebooks/blob/master/_Resources/Images/A.Signal_Acquisition_IMG/a010/wave.png?raw=true" width="500" border="0">
# The following code is based on the example BitMask from Arduino learning
# ```C
#
# byte data=0; // output value
# int Ts=100; // Sampling period
# int pinOff =2; //LSB in
#
#
# void setup()
# {
# for(pin=2; pin<=9;pin++) pintMode(pin, OUTPUT);
# }
#
#
# void loop()
# {
# byte mask=1;
# byte pin=pinOff;
# data=data + 1;
# for(mask=00000001; mask>0; mask<<=1)
# { //iterate through bit mask
# if (data & mask) digitalWrite(pin,HIGH);
# else digitalWrite(pin,LOW);
# pin=pin+1;
# }
# delay(Ts); //delay
# }
#
# ```
# <div style="background:#48ba57;font-family:'arial', monospace; text-align: center; padding: 10px 0; border-radius:10px; width:70%; margin:auto " >
# <span style="font-size:20px;position:relative;color:white; "> Note </span> <br>
# <div style="background:#9de3a6;font-size:12px">
# You can consult the example BitMask from Arduino learning here:
# https://www.arduino.cc/en/Tutorial/BitMask
# </div>
# </div>
# ### <div style="color:#fbb144"> 3. Alternative Example - Writing an entire byte to the output ports </div>
#
#
# ```C
# void setup(){
# // set pins 1 (serial transmit) and 2..7 as output,
# // but leave pin 0 (serial receive) as input
# // (otherwise serial port will stop working!) ...
# DDRD = B11111110; // digital pins 7,6,5,4,3,2,1,0
# // set pins 8..13 as output...
# DDRB = B00111111; // digital pins -,-,13,12,11,10,9,8
# // Turn off digital output pins 2..7 ...
# PORTD &= B00000011; // turns off 2..7, but leaves pins 0 and 1 alone
# // Write simultaneously to pins 8..13...
# PORTB = B00111000; // turns on 13,12,11; turns off 10,9,8
# }
#
# ```
# # III. Explore
# <br>
# <div class='h1' style="width:100%; background:linear-gradient(to right,#FDC86E,#fbb144);color:#282828;font-family:'arial black'; text-align: center; padding: 7px 0; border-radius: 5px 50px;margin-top:-15px" > </div>
# <div style="background:#946db2;font-family:'arial', monospace; text-align: center; padding: 10px 0; border-radius:10px; width:70%; margin:auto " >
# <span style="font-size:20px;position:relative;color:white; "> Explore </span> <br>
# <div style="background:#d0b3e6;font-size:12px">
# Read the “Port Registers” here:
# https://www.arduino.cc/en/Reference/PortManipulation
# </div>
# </div>
# <div style="background:#946db2;font-family:'arial', monospace; text-align: center; padding: 10px 0; border-radius:10px; width:70%; margin:auto " >
# <span style="font-size:20px;position:relative;color:white; "> Explore </span> <br>
# <div style="background:#d0b3e6;font-size:12px">
# Read the “Bit Math Tutorial by CosineKitty” here:
# https://playground.arduino.cc/Code/BitMath/
# </div>
# </div>
# <div style="background:#946db2;font-family:'arial', monospace; text-align: center; padding: 10px 0; border-radius:10px; width:70%; margin:auto " >
# <span style="font-size:20px;position:relative;color:white; "> Explore </span> <br>
# <div style="background:#d0b3e6;font-size:12px">
# Read the “DIRECT CONTROL OF ARDUINO UNO DIGITAL INPUT/OUTPUT PINS USING PORT REGISTERS” here:
# https://www.google.com/url?sa=D&q=http://www.fiz-ix.com/2013/02/direct-control-of-arduino-uno-digital-inputoutput-pins-using-port-registers/&ust=1602691500000000&usg=AOvVaw3jYM6F1D-5Ctyx4DxTClCG&hl=pt-BR
# </div>
# </div>
# <div style="height:100px; background:white;border-radius:10px;text-align:center">
#
# <a> <img src="https://github.com/PIA-Group/ScientIST-notebooks/blob/master/_Resources/Images/IT.png?raw=true" alt="it" style=" bottom: 0; width:250px;
# display: inline;
# left: 250px;
# position: absolute;"/> </a>
# <img src="https://github.com/PIA-Group/ScientIST-notebooks/blob/master/_Resources/Images/IST.png?raw=true"
# alt="alternate text"
# style="position: relative; width:250px; float: left;
# position: absolute;
# display: inline;
# bottom: 0;
# right: 100;"/>
# </div>
# <div style="width: 100%; ">
# <div style="background:linear-gradient(to right,#FDC86E,#fbb144);color:white;font-family:'arial', monospace; text-align: center; padding: 50px 0; border-radius:10px; height:10px; width:100%; float:left " >
# <span style="font-size:12px;position:relative; top:-25px"> Please provide us your feedback <span style="font-size:14px;position:relative;COLOR:WHITE"> <a href="https://forms.gle/C8TdLQUAS9r8BNJM8">here</a>.</span></span>
# <br>
# <span style="font-size:17px;position:relative; top:-20px"> Suggestions are welcome! </span>
# </div>
# ```Contributors: Prof. <NAME>; Prof. <NAME>; <NAME>```
| A.Signal_Acquisition/A010 Digital to Analog Converter - DAC.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.1.0
# language: julia
# name: julia-1.1
# ---
# # Baseline examples
#
# <NAME>
#
# August 2017; updated February 2019
#
# Examples of using the baseline function with various algorithms (splines, polynomials, ALS, arPLS...)
#
# ## Documentation
#
# See http://charlesll.github.io/Spectra.jl/stable/ for information about spectra.
#
# ## References
#
# a good read about the ALS and arPLS algorithms is
#
# <NAME>., <NAME>, <NAME>, and <NAME> (2015), Baseline correction using asymmetrically reweighted penalized least squares smoothing, Analyst, 140(1), 250–257, doi:10.1039/C4AN01061B.
#
# ## Importing the relevant libraries
# +
# The Julia library to treat Raman spectroscopy data
using Spectra
# to plot stuffs
using PyPlot
# we need the Random library
using Random
# -
# ## Creating a fake signal to know the ground truth
# +
x = collect(50:1.0:500)
# 5 gaussian peaks
p1 = 20.0 .* exp.(-log(2) .* ((x .-150.0)./15.0).^2)
p2 = 100.0 .* exp.(-log(2) .* ((x .-250.0)./5.0).^2)
p3 = 50.0 .* exp.(-log(2) .* ((x .-450.0)./1.0).^2)
p4 = 20.0 .* exp.(-log(2) .* ((x .-350.0)./30.0).^2)
p5 = 30.0 .* exp.(-log(2) .* ((x .-460.0)./5.0).^2)
# some background: a large gaussian + linear
bkg = 60.0 .* exp.(-log(2) .* ((x .-250.0)./200.0).^2) .+ 0.1.*x
# some noise
noise = 2.0 * randn!(ones(size(x,1)))
# the observation
y = p1 + p2 + p3 + p4 + p5 + noise +bkg
# making a plot
plot(x,y,"k-",label="signal")
legend()
xlabel("X",fontname="Arial",fontsize=18)
ylabel("Y",fontname="Arial",fontsize=18)
# -
# ## Calling the baseline() function to remove the background
#
# see documentation at http://charlesll.github.io/Spectra.jl/stable/PreProcessing/#baseline-subtraction
# +
# We define the portions of the spectra where we want to fit the signal background.
# roi should be an n x 2 array, see documentation at
roi = [0 100.;200 220;280 290;420 430; 480 500]
# caling the baseline function with a natural spline that will fit the signal in ROIs
y_gcvspl, bas_gcvspl = baseline(x,y,roi,"gcvspline",s=1.)
# using the ALS algorithm (Baek et al. 2015), 10^2-10^5 lambda and 0.001-0.1 p values are recommended
y_als, bas_als = baseline(x,y,roi,"als",p=0.01,lam=10^6,niter=10)
# using the arPLS algorithm (Baek et al. 2015)
y_arpls, bas_arpls = baseline(x,y,roi,"arPLS",p=0.1,lam=10.0^6)
figure()
# plotting the initial signal and the roi
plot(x,y,"black",label="signal")
# plotting the baselines
plot(x,bas_gcvspl,"cyan",label="gcvspline")
plot(x,bas_als,"purple",linestyle="--",label="ALS")
plot(x,bas_arpls,"orange",linestyle="-.",label="arPLS")
legend()
xlabel("X",fontname="Arial",fontsize=18)
ylabel("Y",fontname="Arial",fontsize=18)
# -
| examples/Baseline_examples_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# In this notebook, we are going discuss about methods for evaluating how a model performs - **evaluation metrics**
#
# ### Confusion matrix
#
# Below, we have an example of what is called a **confusion matrix**, it represents a table that describes how the model performed in terms of classfying the data points. For example, instead of only knowing that a point is incorrectly classified we would like to know where does it actually fall compared to the line. We would like to know what **type** of error are we dealing with.
#
# 
#
# Let's think about the graph for a minute. How many True Positives, True Negatives, False Positives, and False Negatives, are in the model above? What we have is:
# - 6 True Positives (correctly classified)
# - 5 True Negatives (correctly classified)
# - 2 False Positives
# - 1 False Negatives
#
# Sometimes in the literature, you'll see False Positives and False Negatives as Type 1 and Type 2 errors. Here is the correspondence:
#
# 
#
# - **Type 1 Error** (Error of the first kind, or False Positive): In the medical example, this is when we misdiagnose a healthy patient as sick.
# - **Type 2 Error** (Error of the second kind, or False Negative): In the medical example, this is when we misdiagnose a sick patient as healthy.
#
# ### Accuracy
#
# This is one of the ways of measuring how good a model is. To better visualize how it works, let's imagine the confusion table again, besides the types we also have the counting for each category. The way accuracy is essentially as a **ratio** of all correctly classified points and total points:
#
# $$ Accuracy = \frac{TP + TN}{TP + TN + FP + FN}$$
#
# where: TP = True positive; FP = False positive; TN = True negative; FN = False negative
#
# 
#
# ### When accuracy is not enough?
#
# While accuracy is a simple way of evaluating the performance it tends to work very poorly when the data is highly skewed failing to capture the incorrectly classified points in a useful manner.
#
# ### Precision
#
# "Precision refers to the closeness of two or more measurements to each other. Using the example above, if you weigh a given substance five times, and get 3.2 kg each time, then your measurement is very precise. Precision is independent of accuracy. You can be very precise but inaccurate, as described above. You can also be accurate but imprecise." $P = \frac{TP}{TP + FP}$
#
# 
#
#
# ### Recall
#
# Recall is the opposite measure of precision, out of the points labelled as positive how many of them were actually did we correctly predicted? $R = \frac{TP}{TP + FN}$
#
# 
#
# ### Receiver operating characteristic (ROC)
#
# From [Wikipedia](https://en.wikipedia.org/wiki/Receiver_operating_characteristic): *An ROC space is defined by FPR and TPR as x and y axes, respectively, which depicts relative trade-offs between true positive (benefits) and false positive (costs)*.
#
# False positive rate: $FPR = \frac{FP}{FP + TN}$
#
# True positive rate: $TPR = \frac{TP}{TP + FN}$
#
# 
#
# Area under ROC (AUC) can also go below 0.5 all the way to 0. Rule of thumb, the closer to 1, the better.
| extra/Evaluation_Metrics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Import Libaries
# +
import os
import shutil
import folium
import rioxarray
import numpy as np
import plotly.graph_objects as go
from glob import glob
from osgeo import gdal
from pyproj import Transformer
from natsort import natsorted, ns
import geopy
from geopy.geocoders import Nominatim
import rasterio
from rasterio.plot import show
# %matplotlib inline
# -
# search_address = "Steenplein 1, 2000 Antwerpen"
# search_address = "Bolivarplaats 20, 2000 Antwerpen"
print("Format example: Steenplein 1, 2000 Antwerpen")
search_address = input("Enter the adress: ")
# # Generete directory proper management
# +
main_dir = ['3D Image',"search address data", 'DSM', 'DTM']
for i in main_dir:
if not os.path.exists(i):
os.makedirs(i)
# -
# # Geo Location of Single Address
# ---
# * "User_Agent" is an http request header that is sent with each request. Nominatim requires this value to be set to your application name. The goal is to be able to limit the number of requests per application. "geocode" method, for resolving a location from a string, and may define a reverse method, which resolves a pair of coordinates to an address.
# * Folium makes it easy to visualize map data it is interactive 'map' Classes for drawing maps. can save in image or interactive html. "marker" set market type.
# * "pyproj.Transformer" has the capabilities of performing 2D, 3D, and 4D (time) transformations. Transform Geo to Bel Geo cordinates
# ---
# * Recieve X and Y value EPSG:31370 to compare with .tif file to find the what file does the location of the x and y is
# ## Function to create Map Location using GeoPy
# +
# to get the longtitude and latitude of the address entered & plot address on a map
def lat_long(address):
# GeoPy to get longtitude and latitude
geolocator = Nominatim(user_agent="Address_GeoLocator")
location = geolocator.geocode(address)
house_lat_long = [location.latitude, location.longitude]
return house_lat_long
def house_locate(func):
# to plot address
house_locate = folium.Map(location=func,zoom_start=18)
folium.Marker(location=func,
popup=list(func),
icon=folium.Icon(color='green', icon='location-arrow', prefix='fa') # Customize Icon
).add_to(house_locate)
#house_locate.save(address+".html")
return house_locate
def EPSG_Bel(lon, lat):
# transform to Belgium 'EPSG:31370' coordinate
transformer = Transformer.from_crs("EPSG:4326",
crs_to = 'EPSG:31370',
always_xy=True) #output coordinates using the traditional GIS order
x, y = transformer.transform(lon, lat)
return x,y
# +
lat,lon = lat_long(search_address)
x, y = EPSG_Bel(lon, lat)
print((x, y) , (lat, lon))
# -
# # Search all ".tif" files in working dir and sort arrage the files
def search_tif(path):
tif_files =[]
# using glob library to get all the file with .tif
files = glob(path,recursive = True)
for file in files:
tif_files.append(file)
# sort files with number in the file
tif_files = natsorted(tif_files, alg=ns.IGNORECASE)
return tif_files
# # Getting Longitute and Latitude form the ".tif" file
# ___
# * GIS raster dataset every pixels of a dataset is contained within a spatial bounding box
# +
# create all bounding box from tifs
def bounding_box(tifs):
bounds = []
for i in tifs:
src = rasterio.open(i) # open the source file
bounds.append(src.bounds) # grab the bounding box corordinates
return bounds
# Locate tif that contains the location and geting the location
def check_tif(x,y):
for i,b_box in enumerate(bounding_box_cordinates,1): # number the bounding box
if (x >= b_box[0] and x <= b_box[2]) & (y >= b_box[1] and y <= b_box[3]): # condition to filter the corordinates
if i in range(1,10): # add '0' for signle digit number to get correct files
i = "0" + str(i)
else:
i = str(i)
dsm_path = f'./DSM/DHMVIIDSMRAS1m_k{i}/GeoTIFF/DHMVIIDSMRAS1m_k{i}.tif'
dtm_path = f'./DTM/DHMVIIDTMRAS1m_k{i}/GeoTIFF/DHMVIIDTMRAS1m_k{i}.tif'
print('DSM File :', f'DHMVIIDSMRAS1m_k{i}.tif')
print('DTM File :', f'DHMVIIDTMRAS1m_k{i}.tif')
return dsm_path, dtm_path
# +
DSM_vla_tif = search_tif('.\\DSM\\**\\*.tif')
DTM_vla_tif = search_tif('.\\DTM\\**\\*.tif')
bounding_box_cordinates = bounding_box(DSM_vla_tif)
tif_path = check_tif(x,y)
print()
dsm_location = tif_path[0]
print(dsm_location)
dtm_location = tif_path[1]
print(dtm_location)
# -
# # Function to clip the house model
# +
"""
Detail documentaion: xarray - Clip
https://corteva.github.io/rioxarray/stable/examples/clip_geom.html?highlight=rio%20clip
"""
def clip_tif(path,window_size=30):
# work with any file that rasterio can open, generate 2D coordinates from the file’s attributes
da = rioxarray.open_rasterio(path,masked=True) # masked=True will convert from integer to float64 and fill with NaN
# Filter which file is function working with 'DSM' or 'DTM'
data_file = path[2:5].lower()
# set window size
ws = window_size
# create coordinates and geometries
c_1 = [(x-ws),(y+ws)]
c_2 = [(x+ws),(y+ws)]
c_3 = [(x+ws),(y-ws)]
c_4 = [(x-ws),(y-ws)]
geometries = [{'type': 'Polygon', 'coordinates': [[c_1,c_2,c_3,c_4,c_1]]}]
# clip the image as per the geometries size
clipped = da.rio.clip(geometries)
# save clip for Canopy Height Model
clip = clipped.rio.to_raster(f"{search_address}_clipped_{data_file}.tif", dtype="int32", tiled=True)
shutil.move(f"{search_address}_clipped_{data_file}.tif", f"./search address data/{search_address}_clipped_{data_file}.tif")
return clipped.plot();
# processing speed
# +
from time import time
t_start = time()
clip_tif(dsm_location,50);
t_end = time()
print(t_end - t_start)
# +
from time import time
t_start = time()
clip_tif(dtm_location,50);
t_end = time()
print(t_end - t_start)
# -
# # Function for Canopy Height Model
"""
Detail documentaion: Numpy masked arrays
https://rasterio.readthedocs.io/en/latest/topics/masks.html?highlight=read(1%2C%20masked%3DTrue)#numpy-masked-arrays
"""
def chm_tif():
# open the digital terrain model
with rasterio.open(f'search address data/{search_address}_clipped_dtm.tif') as src_dataset:
dtm_narr = src_dataset.read(1, masked=True) #read dataset bands as numpy masked arrays
dtm_profile = src_dataset.profile
src_dataset.close()
# open the digital surface model
with rasterio.open(f'search address data/{search_address}_clipped_dsm.tif') as src_dataset:
dsm_narr = src_dataset.read(1, masked=True) # read dataset bands as numpy masked arrays
dsm_profile = src_dataset.profile
src_dataset.close()
# calculate canopy height model
chm_narr = dsm_narr - dtm_narr
# save chm clipped
with rasterio.open(f'search address data/{search_address}_clipped_chm.tif', 'w', **dsm_profile) as dst_dataset:
# Write data to the destination dataset.
dst_dataset.write(chm_narr,1)
dst_dataset.close()
chm_tif = f'search address data/{search_address}_clipped_chm.tif'
return chm_tif
ds = gdal.Open(chm_tif())
data = ds.ReadAsArray()
data = data.astype(np.float32)
ds = None # Close gdal.open
house_locate(lat_long(search_address))
# +
colorscale = [[0, 'gold'], [0.5, 'mediumturquoise'], [1, 'lightsalmon']]
fig = go.Figure(data=[go.Surface(z=data,colorscale=colorscale)])
fig.update_traces(contours_z=dict(show=True, usecolormap=True, highlightcolor="limegreen", project_z=True))
fig.update_layout(title=search_address)
fig.show()
fig.write_image(f"./{main_dir[0]}/{search_address}.png")
# -
fig = go.Figure(data=[go.Surface(z=data)])
fig.update_traces(contours_z=dict(show=True, usecolormap=True, highlightcolor="limegreen", project_z=True))
fig.update_layout(title=search_address)
fig.show()
# to view actual house on google map
import webbrowser
url = f'https://www.google.com.my/maps/place/{str(lat)},{str(lon)}'
webbrowser.open(url)
| 3D_House.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: clouds113_kernel
# language: python
# name: clouds113_kernel
# ---
# ## Multiple linear regression
#
# **For Table 3 of the paper**
#
# Column-based QUBICC R2B5 model
# +
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_squared_error
from sklearn.preprocessing import StandardScaler
from tensorflow.keras import backend as K
from tensorflow.keras.regularizers import l1_l2
import tensorflow.nn as nn
import tensorflow as tf
import gc
import numpy as np
import os
from tensorflow.keras.optimizers import Nadam
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, BatchNormalization
import matplotlib.pyplot as plt
# -
# Prevents crashes of the code
gpus = tf.config.list_physical_devices('GPU')
tf.config.set_visible_devices(gpus[0], 'GPU')
# Allow the growth of memory Tensorflow allocates (limits memory usage overall)
for gpu in gpus:
tf.config.experimental.set_memory_growth(gpu, True)
import time
path = '/pf/b/b309170'
path_data = path + '/my_work/icon-ml_data/cloud_cover_parameterization/grid_column_based_QUBICC_R02B05/based_on_var_interpolated_data'
# +
# order_of_vars_narval = ['qv', 'qc', 'qi', 'temp', 'pres', 'zg', 'fr_land', 'clc']samples_total
# -
input_data = np.transpose(np.load(path_data + '/cloud_cover_input_qubicc.npy', mmap_mode='r'))
output_data = np.transpose(np.load(path_data + '/cloud_cover_output_qubicc.npy', mmap_mode='r'))
(samples_total, no_of_features) = input_data.shape
assert no_of_features == 163
# Remove columns that are constant in at least one of the training folds
# These features correspond to qc_4, qc_5, qc_6, qc_7, qc_8, qc_9, zg_4, zg_5, zg_6
remove_fields = [27, 28, 29, 30, 31, 32, 135, 136, 137]
input_data = np.delete(input_data, remove_fields, axis=1)
no_of_features = no_of_features - len(remove_fields)
# ### Training the multiple linear model on the entire data set
scaler = StandardScaler()
scaler.fit(input_data)
input_data_scaled = scaler.transform(input_data)
# +
t0 = time.time()
# The optimal multiple linear regression model
lin_reg = LinearRegression()
lin_reg.fit(input_data_scaled, output_data)
print(time.time() - t0)
# -
# Loss of this optimal multiple linear regression model
clc_predictions = lin_reg.predict(input_data_scaled)
lin_mse = mean_squared_error(output_data, clc_predictions)
print('The mean squared error of the linear model is %.2f.'%lin_mse)
# ### Zero Output Model
np.mean(output_data**2, dtype=np.float64)
# ### Constant Output Model
output_data.shape
mean = np.mean(output_data, axis=0)
# mean = np.mean(output_data, axis=0)
np.mean(((output_data - mean)**2), dtype=np.float64)
# ### Randomly initialized neural network
model = Sequential()
model.add(Dense(256, activation='relu', input_dim = no_of_features))
model.add(Dense(256, activation='relu'))
model.add(Dense(27, activation='linear'))
model.compile(loss='mse', optimizer=Nadam())
# +
# model_fold_3 is implemented in ICON-A
batch_size = 2**20
for i in range(1 + input_data_scaled.shape[0]//batch_size):
if i == 0:
clc_predictions = model.predict_on_batch(input_data_scaled[i*batch_size:(i+1)*batch_size])
else:
clc_predictions = np.concatenate((clc_predictions, model.predict_on_batch(input_data_scaled[i*batch_size:(i+1)*batch_size])), axis=0)
K.clear_session()
gc.collect()
# -
lin_mse = mean_squared_error(output_data, clc_predictions)
print('The mean squared error of the randomly initialized neural network is %.2f.'%lin_mse)
| additional_content/baselines/multiple_linear_regression_column_based-R2B5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="v7L8ZT2almwl"
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib as mplt
import matplotlib.pyplot as plt
# %matplotlib inline
from sklearn.metrics import accuracy_score, precision_score, recall_score, roc_curve, roc_auc_score, f1_score
from sklearn.model_selection import train_test_split
from sklearn.model_selection import StratifiedKFold
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import GridSearchCV
from sklearn.preprocessing import LabelEncoder
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import MinMaxScaler
from sklearn.preprocessing import RobustScaler
from sklearn.pipeline import Pipeline
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import Dropout
from keras.wrappers.scikit_learn import KerasClassifier
import tensorflow as tf
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.optimizers import Adamax
# + id="lbqW8TTGp4hP"
np.random.seed(42)
tf.random.set_seed(42)
# + id="NBgD1CYTqTsR"
train_data = pd.read_csv('Data/train.csv')
test_data = pd.read_csv('Data/test.csv')
submission_file = pd.read_csv('Data/sample_submission.csv')
# + id="uQl7kDa8qvxK"
train_data.drop(columns=['id'], axis=1, inplace=True)
# + colab={"base_uri": "https://localhost:8080/"} id="WlErXZC-qx2w" outputId="a6a3b783-a4b2-4d56-a2ae-d5933c79fdce"
print(train_data.shape)
print(test_data.shape)
# + id="FKLnnrkmq0OG"
X, y = train_data.drop(columns = ['target']), train_data['target']
# + colab={"base_uri": "https://localhost:8080/"} id="9Ilm-eJVq3Sz" outputId="dc1c720b-d5a9-49e5-dbf3-1fa190ddf9a0"
dtypes = train_data.dtypes
dtypes = dtypes[dtypes != 'object']
features = list(set(dtypes.index) - set(['target']))
len(features)
# + colab={"base_uri": "https://localhost:8080/", "height": 256} id="YTGSonwyq6fu" outputId="87363f93-8c1f-492e-a2c4-225c4596344f"
X.head()
# + colab={"base_uri": "https://localhost:8080/"} id="YyGhcp2StNcg" outputId="43fcef47-42c5-40e8-ef57-fdd86de2ce55"
y.describe()
# + id="jMx41cxq0q25"
X = X.astype(float)
y = y.astype(int)
X.fillna(X.mean(), inplace=True)
y.fillna(y.mean(), inplace=True)
print(f'check for null value in X: {X.isnull().sum().sum()}')
print(f'check for null value in y: {y.isnull().sum().sum()}')
# -
# ## Define Common Functions
def report_results(model_name, y_test, y_train, grid_search_model, grid_search_results):
# summarize results
print("Best: %f using %s" % (grid_search_results.best_score_, grid_search_results.best_params_))
means = grid_search_results.cv_results_['mean_test_score']
stds = grid_search_results.cv_results_['std_test_score']
params = grid_search_results.cv_results_['params']
for mean, stdev, param in zip(means, stds, params):
print("%f (%f) with: %r" % (mean, stdev, param))
y_hat_train = grid_search_model.predict(X_train)
y_hat_test = grid_search_model.predict(X_test)
train_score = accuracy_score(y_train, y_hat_train, normalize=False)
print(f'trian score: {train_score / y_train.shape[0]}')
test_score = accuracy_score(y_test, y_hat_test, normalize=False)
print(f'test score: {test_score / y_test.shape[0]}')
precision_train_score = precision_score(y_train, y_hat_train) * 100
precision_test_score = precision_score(y_test, y_hat_test) * 100
recall_train_score = recall_score(y_train, y_hat_train) * 100
recall_test_score = recall_score(y_test, y_hat_test) * 100
f1_train_score = f1_score(y_train, y_hat_train) * 100
f1_test_score = f1_score(y_test, y_hat_test) * 100
auc_train_score = roc_auc_score(y_train, y_hat_train) * 100
auc_test_score = roc_auc_score(y_test, y_hat_test) * 100
print("Precision = {:.2f}% , recall = {:.2f}% and f1_score={:.2f}% of the % model on the training data.".format(precision_train_score, recall_train_score, f1_train_score, model_name))
print("Precision = {:.2f}% , recall = {:.2f}% and f1_score={:.2f}% of the % model on the validation data.".format(precision_test_score, recall_test_score, f1_test_score, model_name))
print("ROC_AUC Score = {:.2f}% of the % model on the training data.".format(auc_train_score, model_name))
print("ROC_AUC Score = {:.2f}% of the % model on the validation data.".format(auc_test_score, model_name))
# ## Initial Baseline Implementation using KerasClassifier
# +
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.25, random_state=42)
X_train = X_train.reset_index(drop=True)
X_test = X_test.reset_index(drop=True)
scaler = StandardScaler()
num_cols = X_train.select_dtypes(['integer', 'float']).columns
X_train = pd.DataFrame(scaler.fit_transform(X_train[num_cols]), columns=num_cols)
X_test = pd.DataFrame(scaler.fit_transform(X_test[num_cols]), columns=num_cols)
def create_model(optimizer='adam', init='glorot_uniform', learning_rate=0.001):
# create model
model = Sequential()
model.add(Dense(128, input_dim=100, kernel_initializer=init, activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(64, kernel_initializer=init, activation='relu'))
model.add(Dropout(0.2))
#model.add(Dense(32, kernel_initializer=init, activation='relu'))
#model.add(Dropout(0.2))
model.add(Dense(16, kernel_initializer=init, activation='relu'))
model.add(Dropout(0.2))
#model.add(Dense(8, kernel_initializer=init, activation='relu'))
#model.add(Dropout(0.2))
model.add(Dense(1, kernel_initializer=init, activation='sigmoid'))
# Compile model
model.compile(loss='binary_crossentropy', optimizer=Adam(learning_rate=learning_rate), metrics=['accuracy'])
return model
# create model
# KerasClassifier is deprecated, use scikeras instead
# https://github.com/adriangb/scikeras
# https://www.adriangb.com/scikeras/stable/quickstart.html#training-a-model
model = KerasClassifier(build_fn=create_model, verbose=0)
# grid search epochs, batch size and optimizer
optimizers = ['adam']
init = ['glorot_uniform']
epochs = [140]
batches = [2048]
learning_rate = [0.001]
param_grid = dict(optimizer=optimizers, epochs=epochs, batch_size=batches, init=init, learning_rate=learning_rate)
grid = GridSearchCV(estimator=model, param_grid=param_grid, n_jobs=4)
grid_result = grid.fit(X_train, y_train)
report_results('KerasClassifier', y_test, y_train, grid, grid_result)
# +
# re-train with best parameter
X_train, y_train = X, y
X_train = X_train.reset_index(drop=True)
X_test = X_test.reset_index(drop=True)
scaler = StandardScaler()
num_cols = X_train.select_dtypes(['integer', 'float']).columns
X_train = pd.DataFrame(scaler.fit_transform(X_train[num_cols]), columns=num_cols)
# create model
model = KerasClassifier(build_fn=create_model, verbose=0)
optimizers = ['adam']
init = ['glorot_uniform']
epochs = [140]
batches = [2048]
learning_rate = [0.001]
param_grid = dict(optimizer=optimizers, epochs=epochs, batch_size=batches, init=init)
grid = GridSearchCV(estimator=model, param_grid=param_grid, n_jobs=-1)
grid_result = grid.fit(X_train, y_train)
# public score: 0.74687 (epoch=120) => 0.74720 (epoch=140)
# +
# re-train with best parameter
X_train, y_train = X, y
X_train = X_train.reset_index(drop=True)
X_test = X_test.reset_index(drop=True)
scaler = StandardScaler()
num_cols = X_train.select_dtypes(['integer', 'float']).columns
X_train = pd.DataFrame(scaler.fit_transform(X_train[num_cols]), columns=num_cols)
# create model
model = KerasClassifier(build_fn=create_model, verbose=1)
optimizers = ['adam']
init = ['glorot_uniform']
epochs = [130]
batches = [2048]
learning_rate = [0.001]
param_grid = dict(optimizer=optimizers, epochs=epochs, batch_size=batches, init=init)
grid = GridSearchCV(estimator=model, param_grid=param_grid, n_jobs=-1)
grid_result = grid.fit(X_train, y_train)
report_results('KerasClassifier', y_test, y_train, grid, grid_result)
# public score: 0.74712
# -
# ## Kaggle Submission
# +
data_test_norm = pd.DataFrame(scaler.transform(test_data[num_cols]), columns = num_cols)
test_predict = grid.predict_proba(data_test_norm)[::,1]
test_predict = test_predict.astype(float)
array = np.array(test_predict).tolist()
df = pd.DataFrame(test_data['id'])
df['id'] = df['id'].astype(int)
df['target'] = np.array(array)
df.to_csv('results/keras_results.csv', sep=',', encoding='utf-8', index=False)
# +
# %cd results/
# !kaggle competitions submit -c tabular-playground-series-nov-2021 -f keras_results.csv -m "keras implementation"
# %cd ..
# -
# !kaggle competitions submissions -c tabular-playground-series-nov-2021 -q
# ##
| tabular-playground-series-nov-2021/keras_playground_nov_2021.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
filename = '../../data/processed/reshaped.csv'
df = pd.read_csv(filename)
df.head()
# +
# Compute the correlation matrix
corr = df.drop(['zipcode','start_year'], axis=1).corr()
# Generate a mask for the upper triangle
mask = np.triu(np.ones_like(corr, dtype=bool))
# Set up the matplotlib figure
f, ax = plt.subplots(figsize=(11, 9))
# Generate a custom diverging colormap
cmap = sns.diverging_palette(230, 20, as_cmap=True)
# Draw the heatmap with the mask and correct aspect ratio
sns.heatmap(corr, mask=mask, cmap=cmap, vmax=.3, center=0,
square=True, linewidths=.5, cbar_kws={"shrink": .5})
# -
corr['avg_eqi_year_5-10'].drop(['avg_eqi_year_5-10']).sort_values(ascending=True)
df = df[df.columns.drop(list(df.filter(regex='SFR')))]
df['eqi_zip_abs_change'] = df.apply(lambda row: row.year_5_EQI_zip - row.year_1_EQI_zip, axis=1)
df['recpi_zip_abs_change'] = df.apply(lambda row: row.year_5_RECPI_zip - row.year_1_RECPI_zip, axis=1)
df['eqi_zip_pct_change'] = df.apply(lambda row: row.year_5_EQI_zip / row.year_1_EQI_zip, axis=1)
df['recpi_zip_pct_change'] = df.apply(lambda row: row.year_5_RECPI_zip / row.year_1_RECPI_zip, axis=1)
df['eqi_zip_change_1'] = df.apply(lambda row: row.year_2_EQI_zip / row.year_1_EQI_zip, axis=1)
df['eqi_zip_change_2'] = df.apply(lambda row: row.year_3_EQI_zip / row.year_2_EQI_zip, axis=1)
df['eqi_zip_change_3'] = df.apply(lambda row: row.year_4_EQI_zip / row.year_3_EQI_zip, axis=1)
df['eqi_zip_change_4'] = df.apply(lambda row: row.year_5_EQI_zip / row.year_4_EQI_zip, axis=1)
df['eqi_msa_change_1'] = df.apply(lambda row: row.year_2_EQI_MSA / row.year_1_EQI_MSA, axis=1)
df['eqi_msa_change_2'] = df.apply(lambda row: row.year_3_EQI_MSA / row.year_2_EQI_MSA, axis=1)
df['eqi_msa_change_3'] = df.apply(lambda row: row.year_4_EQI_MSA / row.year_3_EQI_MSA, axis=1)
df['eqi_msa_change_4'] = df.apply(lambda row: row.year_5_EQI_MSA / row.year_4_EQI_MSA, axis=1)
df['eqi_state_change_1'] = df.apply(lambda row: row.year_2_EQI_state / row.year_1_EQI_state, axis=1)
df['eqi_state_change_2'] = df.apply(lambda row: row.year_3_EQI_state / row.year_2_EQI_state, axis=1)
df['eqi_state_change_3'] = df.apply(lambda row: row.year_4_EQI_state / row.year_3_EQI_state, axis=1)
df['eqi_state_change_4'] = df.apply(lambda row: row.year_5_EQI_state / row.year_4_EQI_state, axis=1)
df['recpi_zip_change_1'] = df.apply(lambda row: row.year_2_RECPI_zip / row.year_1_RECPI_zip, axis=1)
df['recpi_zip_change_2'] = df.apply(lambda row: row.year_3_RECPI_zip / row.year_2_RECPI_zip, axis=1)
df['recpi_zip_change_3'] = df.apply(lambda row: row.year_4_RECPI_zip / row.year_3_RECPI_zip, axis=1)
df['recpi_zip_change_4'] = df.apply(lambda row: row.year_5_RECPI_zip / row.year_4_RECPI_zip, axis=1)
df['recpi_msa_change_1'] = df.apply(lambda row: row.year_2_RECPI_MSA / row.year_1_RECPI_MSA, axis=1)
df['recpi_msa_change_2'] = df.apply(lambda row: row.year_3_RECPI_MSA / row.year_2_RECPI_MSA, axis=1)
df['recpi_msa_change_3'] = df.apply(lambda row: row.year_4_RECPI_MSA / row.year_3_RECPI_MSA, axis=1)
df['recpi_msa_change_4'] = df.apply(lambda row: row.year_5_RECPI_MSA / row.year_4_RECPI_MSA, axis=1)
df['recpi_state_change_1'] = df.apply(lambda row: row.year_2_RECPI_state / row.year_1_RECPI_state, axis=1)
df['recpi_state_change_2'] = df.apply(lambda row: row.year_3_RECPI_state / row.year_2_RECPI_state, axis=1)
df['recpi_state_change_3'] = df.apply(lambda row: row.year_4_RECPI_state / row.year_3_RECPI_state, axis=1)
df['recpi_state_change_4'] = df.apply(lambda row: row.year_5_RECPI_state / row.year_4_RECPI_state, axis=1)
df['avg_eqi_year_1-5'] = df.apply(lambda row: ((row.year_1_EQI_zip + row.year_2_EQI_zip + row.year_3_EQI_zip + row.year_4_EQI_zip + row.year_5_EQI_zip)/5), axis=1)
# Compute new correlation matrix
corr = df.drop(['zipcode','start_year'], axis=1).corr()
corr['avg_eqi_year_5-10'].drop(['avg_eqi_year_5-10']).sort_values(ascending=True)
# +
# Generate a mask for the upper triangle
mask = np.triu(np.ones_like(corr, dtype=bool))
# Set up the matplotlib figure
f, ax = plt.subplots(figsize=(11, 9))
# Generate a custom diverging colormap
cmap = sns.diverging_palette(230, 20, as_cmap=True)
# Draw the heatmap with the mask and correct aspect ratio
sns.heatmap(corr, mask=mask, cmap=cmap, vmax=.3, center=0,
square=True, linewidths=.5, cbar_kws={"shrink": .5})
# -
df.head()
path = '../../data/processed/feature-eng.csv'
df.to_csv(path, index=False)
| notebooks/preprocessing/3.2-tjc-feature-engineering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"name": "#%%\n"}
from sys import modules
IN_COLAB = 'google.colab' in modules
if IN_COLAB:
# !pip install -q ir_axioms[examples] python-terrier
# + pycharm={"name": "#%%\n"}
# Start/initialize PyTerrier.
from pyterrier import started, init
if not started():
init(tqdm="auto")
# + pycharm={"name": "#%%\n"}
edition = 28
track = "deep.documents"
dataset_name = "msmarco-document/trec-dl-2019/judged"
contents_field = "body"
depth = 10
# + pycharm={"name": "#%%\n"}
from pyterrier.datasets import get_dataset
from ir_datasets import load
dataset = get_dataset(f"irds:{dataset_name}")
ir_dataset = load(dataset_name)
# + pycharm={"name": "#%%\n"}
from pathlib import Path
cache_dir = Path("cache/")
index_dir = cache_dir / "indices" / dataset_name.split("/")[0]
result_dir = Path(
"/mnt/ceph/storage/data-in-progress/data-research/"
"web-search/web-search-trec/trec-system-runs"
) / f"trec{edition}" / track
result_files = list(result_dir.iterdir())
# + pycharm={"name": "#%%\n"}
from pyterrier.index import IterDictIndexer
if not index_dir.exists():
indexer = IterDictIndexer(str(index_dir.absolute()))
indexer.index(
dataset.get_corpus_iter(),
fields=[contents_field]
)
# + pycharm={"name": "#%%\n"}
from pyterrier.io import read_results
from pyterrier import Transformer
from tqdm.auto import tqdm
results = [
Transformer.from_df(read_results(result_file))
for result_file in tqdm(result_files, desc="Load results")
]
results_names = [result_file.stem.replace("input.", "") for result_file in result_files]
# + pycharm={"name": "#%%\n"}
from ir_axioms.axiom import (
ArgUC, QTArg, QTPArg, aSL, PROX1, PROX2, PROX3, PROX4, PROX5, TFC1, TFC3, RS_TF, RS_TF_IDF, RS_BM25, RS_PL2, RS_QL,
AND, LEN_AND, M_AND, LEN_M_AND, DIV, LEN_DIV, M_TDC, LEN_M_TDC, STMC1, STMC1_f, STMC2, STMC2_f, LNC1, TF_LNC, LB1,
REG, ANTI_REG, ASPECT_REG, REG_f, ANTI_REG_f, ASPECT_REG_f
)
axioms = [
~ArgUC(), # Very slow due to network access.
~QTArg(), # Very slow due to network access.
~QTPArg(), # Very slow due to network access.
~aSL(),
~LNC1(),
~TF_LNC(),
~LB1(),
~PROX1(),
~PROX2(),
~PROX3(),
~PROX4(),
~PROX5(),
~REG(),
~REG_f(),
~ANTI_REG(),
~ANTI_REG_f(),
~ASPECT_REG(),
~ASPECT_REG_f(),
~AND(),
~LEN_AND(),
~M_AND(),
~LEN_M_AND(),
~DIV(),
~LEN_DIV(),
~RS_TF(),
~RS_TF_IDF(),
~RS_BM25(),
~RS_PL2(),
~RS_QL(),
~TFC1(),
~TFC3(),
~M_TDC(),
~LEN_M_TDC(),
~STMC1(), # Rather slow due many similarity calculations.
~STMC1_f(), # Rather slow due many similarity calculations.
~STMC2(),
~STMC2_f(),
]
axiom_names = [axiom.axiom.name for axiom in axioms]
# + pycharm={"name": "#%%\n"}
from ir_axioms.backend.pyterrier.experiment import AxiomaticExperiment
experiment = AxiomaticExperiment(
retrieval_systems=results,
topics=dataset.get_topics(),
qrels=dataset.get_qrels(),
index=index_dir,
dataset=ir_dataset,
contents_accessor=contents_field,
axioms=axioms,
axiom_names=axiom_names,
depth=depth,
filter_by_qrels=False,
filter_by_topics=False,
verbose=True,
cache_dir=cache_dir,
)
# + pycharm={"name": "#%%\n"}
preferences = experiment.preferences
# + pycharm={"name": "#%%\n"}
preferences.to_csv(f"trec-{edition}-{track}-preferences-all-axioms-depth-{depth}.csv")
| examples/trec_28_deep_documents_preferences_depth_10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Assignment 3 Naive bayes classifier
# ### <NAME> 1001778272
#
# platform:
# I7-9700k GTX-1080ti
import os
import re
import string
import csv
# from bs4 import BeautifulSoup
import math
import random
from queue import PriorityQueue as PQueue
# ## Get data from txt
# My thoughts of this part come from my own homework in my Machine learning class.
# +
# constant
train_positive_file_dir = 'aclImdb/train/pos'
train_negitive_file_dir = 'aclImdb/train/neg'
test_positive_file_dir = 'aclImdb/test/pos'
test_negitive_file_dir = 'aclImdb/test/neg'
train_unsup_file_dir = 'aclImdb/train/unsup'
# -
def file_name(file_dir):
files_name = os.listdir(file_dir)
return files_name
# save filename in txt
def save_name(file_dir, name):
f = open(name + '.txt' ,'w') # 'a' add not reset.
files_name = file_name(file_dir)
for i in files_name:
f.write(i) # string
f.write("\n")
# +
save_name(train_positive_file_dir, 'train_positive_file_dir')
save_name(train_negitive_file_dir, 'train_negitive_file_dir')
save_name(test_positive_file_dir, 'test_positive_file_dir')
save_name(test_negitive_file_dir, 'test_negitive_file_dir')
save_name(train_unsup_file_dir, 'train_unsup_file_dir')
# -
def get_data(filename):
f = open(filename +'.txt')
file_names = []
for i in f.readlines():
file_names.append(i.replace("\n", ""))
return file_names
def load_data(fileList, url):
sentenseList = []
pa = string.punctuation
for file in fileList:
with open(url +"/"+ file,errors='ignore') as f:
ori_data = f.read().lower()
data1 = re.sub('\n{2,6}',' ',ori_data)
data2 = re.sub('\n',' ',data1)
data3 = re.sub(' ','yxw ',data2)
data4 = re.sub("[%s]+"%('"|#|$|%|&|\|(|)|*|+|-|/|<|=|>|@|^|`|{|}|~'), "", data3)
sentense = re.sub("[%s]+"%('.|?|!|:|;'),' ',data4)
sentenseList.append(sentense)
return sentenseList
# +
file_names_train_pos = get_data('train_positive_file_dir')
file_names_train_neg = get_data('train_negitive_file_dir')
file_names_test_pos = get_data('test_positive_file_dir')
file_names_test_neg = get_data('test_negitive_file_dir')
file_names_train_unsup = get_data('train_unsup_file_dir')
# +
train_sentenseList1 = load_data(file_names_train_pos, train_positive_file_dir)
train_sentenseList2 = load_data(file_names_train_neg, train_negitive_file_dir)
test_sentenseList1 = load_data(file_names_test_pos, test_positive_file_dir)
test_sentenseList2 = load_data(file_names_test_neg, test_negitive_file_dir)
train_unsup_sentenseList = load_data(file_names_train_unsup, train_unsup_file_dir)
# +
train_target1 = [1]*len(train_sentenseList1)
train_target2 = [0]*len(train_sentenseList2)
train_target = train_target1 + train_target2
train_text1 = train_sentenseList1
train_text2 = train_sentenseList2
train_text = train_text1 + train_text2
test_target1 = [1]*len(test_sentenseList1)
test_target2 = [0]*len(test_sentenseList2)
test_target = test_target1 + test_target2
test_text1 = test_sentenseList1
test_text2 = test_sentenseList2
test_text = test_text1 + test_text2
train_unsup_target = [0]*len(train_unsup_sentenseList)
train_unsup_text = train_unsup_sentenseList
# +
train_to_dict = {'content':train_text, 'target':train_target}
test_to_dict = {'content':test_text, 'target':test_target}
train_unsup_to_dict = {'content':train_unsup_text, 'target':train_unsup_target}
# -
# ## Remove stopwords and useless symbol
# +
# Copy the stopwords from wordcloud.
# My thoughts of this part come from my own homework in my Machine learning class.
stopSet = set({'did', 'such', 'doing', 'down', 'me', 'just', 'very', 'shan', 'against', 't', "you're",
'only', "haven't", 'yours', 'you', 'its', 'other', 'we', 'where', 'then', 'they', 'won', "you've",
'some', 've', 'y', 'each', "you'll", 'them', 'to', 'was', 'once', 'and', 'ain', 'under', 'through',
'for', "won't", 'mustn', 'a', 'are', 'that', 'at', 'why', 'any', 'nor', 'these', 'yourselves',
'has', 'here', "needn't", 'm', 'above', 'up', 'more', 'if', 'ma', 'didn', 'whom', 'can', 'have',
'an', 'should', 'there', 'couldn', 'her', 'how', 'of', 'doesn', "shouldn't", 'further',
"wasn't", 'between', 'd', 'wouldn', 'his', 'being', 'do', 'when', 'hasn', "she's", 'by', "should've",
'into', 'aren', 'weren', 'as', 'needn', 'what', "it's", 'hadn', 'with', 'after', 'he', 'off', 'not',
'does', 'own', "weren't", "isn't", 'my', 'too', "wouldn't", 'been', 'again', 'same', 'few', "don't",
'our', 'myself', 'your', 'before', 'about', 'most', 'during', 'll', 'on', 'shouldn', 'is', 'out',
"shan't", 'below', 'which', 'from', 'she', 'were', 'those', 'over', 'until', 'theirs', 'mightn',
'yourself', 'i', 'am', 'so', 'himself', 'it', 'had', 'or', 'all', 'while', "aren't", 'ours',
"that'll", 'but', 'because', 'in', 'now', 'themselves', 'him', "doesn't", 'both', 're', 'wasn',
's', "hasn't", "didn't", 'their', "mustn't", 'herself', 'the', 'this', 'will', 'isn', "you'd",
'haven', 'itself', "couldn't", 'o', 'be', 'don', 'hers', "mightn't", 'having', "hadn't", 'ourselves',
'who', 'than'})
# # Remove html characters. I used BeautifulSoup at this part but it's ok to remove this function.
# # So I just put it here as a comment but not use it.
# def strip_html(text):
# soup = BeautifulSoup(text, "html.parser")
# return soup.get_text()
#Removing the square brackets
def remove_between_square_brackets(text):
return re.sub('\[[^]]*\]', '', text)
def remove_special_characters(text, remove_digits=True):
pattern=r'[^a-zA-z0-9\s]'
text=re.sub(pattern,'',text)
return text
def remove_stopwords(text, is_lower_case=False):
# print(text)
tokens = text.split();
if is_lower_case:
filtered_tokens = [token for token in tokens if token not in stopSet]
else:
filtered_tokens = [token for token in tokens if token.lower() not in stopSet]
filtered_text = ' '.join(filtered_tokens)
return filtered_text
#Removing the noisy text
def denoise_text(text):
# text = strip_html(text)
text = remove_between_square_brackets(text)
text = remove_special_characters(text)
text = remove_stopwords(text)
return text
# +
# make a copy of data dict
train_to_dict_tmp = {'content':[], 'target':[]}
test_to_dict_tmp = {'content':[], 'target':[]}
unsup_to_dict_tmp = {'content':[], 'target':[]}
for i in range(len(train_to_dict['content'])):
train_to_dict_tmp['content'].append(denoise_text(train_to_dict['content'][i]))
train_to_dict_tmp['target'] = train_to_dict['target']
for i in range(len(test_to_dict['content'])):
test_to_dict_tmp['content'].append(denoise_text(test_to_dict['content'][i]))
test_to_dict_tmp['target'] = test_to_dict['target']
for i in range(len(train_unsup_to_dict['content'])):
unsup_to_dict_tmp['content'].append(denoise_text(train_unsup_to_dict['content'][i]))
unsup_to_dict_tmp['target'] = train_unsup_to_dict['target']
print(test_to_dict_tmp['target'][0])
print(test_to_dict['content'][0])
# -
# ## Build dictionary
# Define storage of word dictionary and occurrence time
wordSet = [] # store word as a set
word_dictionary = {} #dictionary, {Integer->index: String->word: }
word_count = {} #count the frequency of each word, {Integer->index: Integer->frequency}
word_occurrence = {} # count num of documents containing current word.
word_occurrence_pos = {}
word_occurrence_neg = {}
def get_word_set(wordSet, word_dict):
for text in word_dict['content']:
text_words = text.split()
for word in text_words:
wordSet.append(word);
return wordSet
wordSet = get_word_set(wordSet, train_to_dict_tmp)
wordSet = get_word_set(wordSet, unsup_to_dict_tmp)
wordSet = set(wordSet)
# wordSet is a set contains all words.
# build dictionary
count = 0 # index
for i in wordSet:
word_dictionary[i] = count
count = count + 1
word_dictionary
len(word_dictionary)
# +
def get_count(word_dictionary, word_count, wordSet, word_dict):
for text in word_dict['content']:
text_words = text.split()
for word in text_words:
# print(word)
if word in word_dictionary.keys():
if word_dictionary[word] not in word_count.keys():
word_count[word_dictionary[word]] = 1
else:
word_count[word_dictionary[word]] = word_count[word_dictionary[word]] + 1
return word_count
# -
word_count = get_count(word_dictionary, word_count, wordSet, train_to_dict_tmp)
word_count = get_count(word_dictionary, word_count, wordSet, unsup_to_dict_tmp)
# +
# omit rare words for example if the occurrence is less than five times
for word in wordSet:
if word_count[word_dictionary[word]] <= 5:
word_count.pop(word_dictionary[word])
word_dictionary
len(word_count)
# -
word_count
# ## calculate probability and conditional probability
# cauculate probability of each word. calculate word_occurrence_pos{} and word_occurrence_neg{}
def get_probability(word_dict, word_dictionary, wordSet, word_occurrence, word_occurrence_pos, word_occurrence_neg):
doc_number = 0
for i in range(len(word_dict['content'])):
text = word_dict['content'][i]
target = word_dict['target'][i]
# print(target)
doc_number += 1
text_word_set = set(text.split())
for word in text_word_set:
tmp = word_dictionary[word]
if tmp in word_occurrence.keys():
word_occurrence[tmp] += 1
else:
word_occurrence[tmp] = 1
if target == 0:
if tmp in word_occurrence_neg.keys():
word_occurrence_neg[tmp] += 1
else:
word_occurrence_neg[tmp] = 1
if target == 1:
if tmp in word_occurrence_pos.keys():
word_occurrence_pos[tmp] += 1
else:
word_occurrence_pos[tmp] = 1
return word_occurrence, word_occurrence_pos, word_occurrence_neg, doc_number
# get conditional probability with laplace smoothing
def get_condition_laplace(word, word_dictionary, word_occurrence, word_occurrence_pos, word_occurrence_neg, size_of_data):
tmp = word_dictionary[word]
conditional_probability_pos = 0
conditional_probability_neg = 0
if tmp in word_occurrence_pos.keys():
# Laplace Smoothing
conditional_probability_pos = float(word_occurrence_pos[tmp] + 1)/ float(size_of_data + 2)
else: conditional_probability_pos = float(1)/ float(size_of_data + 2)
if tmp in word_occurrence_neg.keys():
conditional_probability_neg = float(word_occurrence_neg[tmp] + 1)/ float(size_of_data + 2)
else: conditional_probability_neg = float(1)/ float(size_of_data + 2)
return conditional_probability_pos, conditional_probability_neg
# get conditional probability with m estimate smoothing
def get_condition_m_estimate(word, word_dictionary, word_occurrence, word_occurrence_pos, word_occurrence_neg, size_of_data):
tmp = word_dictionary[word]
conditional_probability_pos = 0
conditional_probability_neg = 0
if tmp in word_occurrence_pos.keys():
conditional_probability_pos = float(word_occurrence_pos[tmp] + 1.5)/ float(size_of_data + 3)
else: conditional_probability_pos = float(1.5)/ float(size_of_data + 3)
if tmp in word_occurrence_neg.keys():
conditional_probability_neg = float(word_occurrence_neg[tmp] + 1.5)/ float(size_of_data + 3)
else: conditional_probability_neg = float(1.5)/ float(size_of_data + 3)
return conditional_probability_pos, conditional_probability_neg
def naive_bayes(text, word_dictionary, word_occurrence, word_occurrence_pos, word_occurrence_neg, size_of_data):
pro_pos = 0
pro_neg = 0
for word in text.split():
if word in word_dictionary.keys():
tmp = word_dictionary[word]
if tmp in word_count.keys() and tmp in word_occurrence.keys():
# laplace
# conditional_probability_pos, conditional_probability_neg \
# = get_condition_laplace(word, word_dictionary, word_occurrence, word_occurrence_pos, word_occurrence_neg, size_of_data)
# m estimate
conditional_probability_pos, conditional_probability_neg \
= get_condition_m_estimate(word, word_dictionary, word_occurrence, word_occurrence_pos, word_occurrence_neg, size_of_data)
pro_pos += math.log(conditional_probability_pos)
pro_neg += math.log(conditional_probability_neg)
# print(pro_neg, pro_pos)
if pro_neg > pro_pos: return 0
return 1
def get_accuracy(word_dict, word_dictionary, word_occurrence, word_occurrence_pos, word_occurrence_neg):
count = 0
score = 0
for i in range(len(word_dict['content'])):
count += 1
text = word_dict['content'][i]
target = word_dict['target'][i]
res = naive_bayes(text, word_dictionary, word_occurrence, word_occurrence_pos, word_occurrence_neg, len(word_dict['content']))
if target == res:
score += 1
return float(score)/ float(count)
# ## Calculate accuracy using dev dataset
# ### compare smoothing methods: Laplace and m estimate smoothing.
def fold_validation(word_dict, n_fold):
dataset_split = []
text = word_dict['content']
fold_size = len(text)// n_fold
index_list = random.sample(range(0,len(text)),len(text))
# print(index)
for i in range (n_fold):
tmp_dict = {}
tmp_content = []
tmp_target = []
for index in index_list[i * fold_size: (i + 1)* fold_size]:
tmp_content.append(text[index])
tmp_target.append(word_dict['target'][index])
tmp_dict = {'content':tmp_content, 'target': tmp_target}
dataset_split.append(tmp_dict)
return dataset_split
# 5 folds validation
n_fold = 5
data_after_n_fold = fold_validation(train_to_dict_tmp, n_fold)
# using development dataset
# compare m-estimate and laplace smoothing. edit naive_bayes() line 8-13
score = []
for i in range(n_fold):
train_dict_final = {}
for tmp in range(n_fold):
tmp_content = []
tmp_target = []
if tmp != i:
tmp_content = tmp_content + data_after_n_fold[tmp]['content']
tmp_target = tmp_target + data_after_n_fold[tmp]['target']
train_dict_final = {'content':tmp_content, 'target': tmp_target}
test_dict_final = data_after_n_fold[i]
word_occurrence, word_occurrence_pos, word_occurrence_neg, doc_number \
= get_probability(train_dict_final, word_dictionary, wordSet, word_occurrence,\
word_occurrence_pos, word_occurrence_neg)
score.append(get_accuracy(test_dict_final, word_dictionary, word_occurrence, word_occurrence_pos, word_occurrence_neg))
print(score)
mean_score = 0
for i in score:
mean_score += i
print(float(mean_score)/ len(score))
# ### result of laplace smoothing
# round 1:
# [0.9, 0.8866, 0.8894, 0.888, 0.931]
# 0.899
# round 2
# [0.9, 0.8988, 0.8964, 0.8902, 0.9338]
# 0.90384
#
# ### result of m estimate smoothing
# round 1
# [0.905, 0.8964, 0.9014, 0.8958, 0.9268]
# 0.9050799999999999
# round 2
# [0.9022, 0.901, 0.903, 0.905, 0.9238]
# 0.907
#
# m estimate smoothing have better result
# ## Derive Top 10 words that predicts positive and negative class
# +
def get_top_10(word_dict, word_dictionary, word_occurrence, word_occurrence_pos, word_occurrence_neg):
top_10_pos = PQueue()
top_10_neg = PQueue()
for word in word_dictionary.keys():
if word in word_dictionary.keys():
tmp = word_dictionary[word]
if tmp in word_count.keys() and tmp in word_occurrence.keys():
conditional_probability_pos, conditional_probability_neg \
= get_condition_laplace(word, word_dictionary, word_occurrence, word_occurrence_pos, word_occurrence_neg, len(word_dict['content']))
top_10_pos.put([conditional_probability_pos * -1, word])
# if top_10_pos.qsize() > 10: top_10_pos.get()[-1]
top_10_neg.put([conditional_probability_neg * -1, word])
# if top_10_neg.qsize() > 10: top_10_neg.get()[-1]
return top_10_pos, top_10_neg
top_10_pos, top_10_neg = get_top_10(train_to_dict ,word_dictionary, word_occurrence, word_occurrence_pos, word_occurrence_neg)
# -
print('Top 10 words that predicts positive')
i = 10
while i > 0:
print(top_10_pos.get(-1)[1])
i -= 1
print('\n')
print('Top 10 words that predicts negative')
i = 10
while i > 0:
print(top_10_neg.get(-1)[1])
i -= 1
# ## Using the test dataset calculate the final accuracy.
# +
# final accuracy using m estimate smoothing
word_occurrence, word_occurrence_pos, word_occurrence_neg, doc_number \
= get_probability(train_to_dict_tmp, word_dictionary, wordSet, word_occurrence,\
word_occurrence_pos, word_occurrence_neg)
score = get_accuracy(train_to_dict_tmp, word_dictionary, word_occurrence, word_occurrence_pos, word_occurrence_neg)
score
# -
# final accuracy using m estimate smoothing = 0.91176
| static/files/.ipynb_checkpoints/NaiveBayesClassifier-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from __future__ import absolute_import
import sys
sys.path.append("..")
import numpy as np
np.seterr(invalid='raise')
import pandas as pd
import datetime
import csv
import os
import uuid
import matplotlib.pyplot as plt
from utils.data import read_h5_fx_history, read_csv_history
from utils.globals import EPS, DATASETS_DIR, OUTPUTS_DIR, CAPITAL_BASE_MULTIPLIER, MAX_WEIGHT, RISK
# +
#currency is the account currency
#curent_prices is a 3d array: 1dim is list of instuments, 2dim is window, 3dim is price
def _calculate_pip_value_in_account_currency_(currency, current_prices):
pip_values = []
if currency == account_currency.USD:
m = 0
print(instruments)
for instrument in instruments:
print(instrument)
current_prices_for_conversion = current_prices[m]
#if instrument == 'EUR/USD':
# EUR_USD = current_prices[m]
#elif instrument == 'USD/JPY':
# USD_JPY = current_prices[m]
#elif instrument == 'AUD/USD':
# AUD_USD = current_prices[m]
#elif instrument == 'GBP/USD':
# GBP_USD = current_prices[m]
first_currency = instrument[0:3]
second_currency = instrument[3:6]
print(first_currency + ' ' + second_currency)
if second_currency == 'USD':
pip_value = 0.0001
elif
elif first_currency == 'USD' and second_currency != 'JPY':
pip_value = 0.0001/current_prices[m]
elif first_currency == 'USD' and second_currency == 'JPY':
pip_value = 0.01/current_prices[m]
elif instrument == 'GBP/JPY':
pip_value = GBP_USD * 0.01/current_prices[m]
elif instrument == 'EUR/JPY':
pip_value = EUR_USD * 0.01/current_prices[m]
elif instrument == 'AUD/JPY':
pip_value = AUD_USD * 0.01/current_prices[m]
elif instrument == 'EUR/GBP':
pip_value = EUR_USD * 0.0001/current_prices[m]
pip_values.append(pip_value)
m += 1
return pip_values
def _calculate_pip_value_in_account_currency(self, currency, current_prices):
pip_values = []
if currency == account_currency.USD:
m = 0
print(self.instruments)
for instrument in self.instruments:
print(instrument)
if instrument == 'EUR/USD':
EUR_USD = current_prices[m]
elif instrument == 'USD/JPY':
USD_JPY = current_prices[m]
elif instrument == 'AUD/USD':
AUD_USD = current_prices[m]
elif instrument == 'GBP/USD':
GBP_USD = current_prices[m]
first_currency = instrument[0:3]
second_currency = instrument[3:6]
print(first_currency + ' ' + second_currency)
if second_currency == 'USD':
pip_value = 0.0001
elif first_currency == 'USD' and second_currency != 'JPY':
pip_value = 0.0001/current_prices[m]
elif first_currency == 'USD' and second_currency == 'JPY':
pip_value = 0.01/current_prices[m]
elif instrument == 'GBP/JPY':
pip_value = GBP_USD * 0.01/current_prices[m]
elif instrument == 'EUR/JPY':
pip_value = EUR_USD * 0.01/current_prices[m]
elif instrument == 'AUD/JPY':
pip_value = AUD_USD * 0.01/current_prices[m]
elif instrument == 'EUR/GBP':
pip_value = EUR_USD * 0.0001/current_prices[m]
pip_values.append(pip_value)
m += 1
return pip_values
# -
data, bid, ask, instruments = read_h5_fx_history(filepath=DATASETS_DIR +'fxcm_11_H4_2015_2018_test_with_dates_1000.h5', replace_zeros=True)
print('after read instuments {}', instruments)
# +
def instruments = ['EUR/USD', 'USD/JPY', 'AUD/USD', 'USD/CAD', 'GBP/USD', 'NZD/USD', 'GBP/JPY', 'EUR/JPY', 'AUD/JPY', 'EUR/GBP', 'USD/CHF']
def current_prices = [['USD/GBP',[1.30035, 1.30061, 1.29652]]]
| test_cases/.ipynb_checkpoints/Pip_converion_tester-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
#execfile(os.path.join(os.environ["SPARK_HOME"], 'python/pyspark/shell.py'))
from pyspark.sql import SparkSession
sparkSession = SparkSession.builder.enableHiveSupport().master("local").getOrCreate()
playHistoryPath = '/data/sample264'
playHistoryGraph = sparkSession.read.parquet(playHistoryPath)
playHistoryGraph.printSchema()
metaDataPath = '/data/meta'
metaDataGraph = sparkSession.read.parquet(metaDataPath)
metaDataGraph.printSchema()
playHistoryGraph.createTempView("history1")
playHistoryGraph.createTempView("history2")
# +
from pyspark.sql.functions import count, col
consecutiveTracksForUser = sparkSession.sql( \
"select h1.trackId as track1, h2.trackId as track2, h1.userId as user " \
"from history1 h1, history2 h2 " \
"where h1.userId = h2.userId " \
"and h1.trackId != h2.trackId " \
"and abs(h2.timestamp - h1.timestamp) <= 420 " \
).groupBy(col("track1"), col("track2")) \
.count().alias("count") \
.orderBy(col("track1"), col("track2")) \
.cache()
# +
#consecutiveTracksForUser.show()
# +
from pyspark.sql import Window
from pyspark.sql.functions import col, row_number, sum
window = Window.partitionBy("track1").orderBy(col("count").desc())
topsDF = consecutiveTracksForUser.withColumn("row_number", row_number().over(window)) \
.filter(col("row_number") <= 40) \
.drop(col("row_number")) \
.orderBy(col("track1"), col("track2")) \
.cache()
# +
#topsDF.show()
# +
sumsDF = topsDF.groupBy(col("track1")) \
.agg(sum(col("count")).alias("sum_weights")) \
.orderBy("track1") \
.cache()
# +
#sumsDF.show()
# -
normalized_count = topsDF.join(sumsDF, "track1", "inner") \
.withColumn("weight", col("count") / col("sum_weights")) \
.cache()
# +
#normalized_count.show()
# -
results = normalized_count.orderBy(col("weight").desc(), col("track1"), col("track2")).limit(40)
#results.show()
results = results.select(col("track1"), col("track2"))
for t1, t2 in results.collect():
print("{}\t{}".format(t1,t2))
| Week5-Graph-based-music-recommender-task-1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
from math import sqrt
from xgboost import XGBRegressor
from lightgbm import LGBMRegressor, LGBMClassifier
from xgboost import plot_importance
from sklearn.pipeline import Pipeline
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import OneHotEncoder
from sklearn.preprocessing import LabelEncoder
from sklearn.preprocessing import FunctionTransformer
from sklearn.preprocessing import StandardScaler
from sklearn.compose import ColumnTransformer
from sklearn.ensemble import RandomForestRegressor
from sklearn.ensemble import VotingRegressor
from sklearn.metrics import mean_absolute_error, mean_squared_error, roc_auc_score
from sklearn.model_selection import train_test_split
from sklearn.model_selection import GridSearchCV
from sklearn.model_selection import ParameterGrid
from sklearn.model_selection import cross_val_score
from sklearn.feature_selection import VarianceThreshold
from optuna.integration import OptunaSearchCV
from optuna.distributions import *
import operator
from functools import reduce
import sys
sys.path.append('../../../src')
from pipeline_utils import LGBMClassifierEarlyStopping, auc
data_dir = '../data'
RANDOM_STATE = 2021
df = pd.read_csv(f"{data_dir}/interim/train_for_David.csv", index_col='id')
# df = pd.read_parquet(f"{data_dir}/interim/train.parq", engine='pyarrow').convert_dtypes()
display(df.shape)
df.head(2)
numeric_features = [col for col in df.columns if col.startswith('cont')]
categorical_features = [col for col in df.columns if col.startswith('cat')]
def feature_engineering(df):
numeric_features = [col for col in df.columns if col.startswith('cont')]
categorical_features = [col for col in df.columns if col.startswith('cat')]
df[numeric_features] = df[numeric_features].astype('float')
df[categorical_features] = df[categorical_features].apply(lambda i: [sum(map(ord, x)) for x in i], axis='rows').astype('int')
df['gt_0.1'] = df[numeric_features].apply(lambda x: len([i for i in x if i > 0.1]), axis=1)
df['gt_0.5'] = df[numeric_features].apply(lambda x: len([i for i in x if i > 0.5]), axis=1)
df['mul_gt_o.1'] = 0
df.loc[(df['gt_0.1'] >=1), 'mul_gt_o.1'] = df[numeric_features].apply(lambda x: reduce(operator.mul, x), axis=1)
return df
df['target'] = df['target'].astype('int')
# df[categorical_features] = df[categorical_features].apply(lambda x: x.cat.codes).astype('int').astype('category')
X = df.copy()
y = X.pop('target')
X = feature_engineering(X)
X_train, X_valid, y_train, y_valid = train_test_split(
X, y, train_size=0.8, test_size=0.2, random_state=RANDOM_STATE,
)
# Numeric Pipeline
numeric_transformer = Pipeline(steps=[
('imputer', SimpleImputer(strategy='mean')),
('log', FunctionTransformer(np.log1p)),
('scaler', StandardScaler()),
])
# Categorical Pipeline
class MultiColumnLabelEncoder:
def __init__(self):
pass
def fit(self,X,y=None):
return self
def transform(self, X):
output = []
for x in X:
output.append(LabelEncoder().fit_transform(x))
return np.array(output)
def fit_transform(self,X,y=None):
return self.fit(X, y).transform(X)
categorical_transformer = Pipeline(steps=[
('imputer', SimpleImputer(strategy='most_frequent')),
('encoder', OneHotEncoder(handle_unknown='ignore', sparse=False)),
# ('encoder', MultiColumnLabelEncoder())
])
# Preprocess Pipeline
# - merge cateogrical & numeric into one pipeline
preprocessor = ColumnTransformer(
transformers=[
('num', numeric_transformer, X.columns),
# ('cat', categorical_transformer, categorical_features),
# ('num', numeric_transformer, numeric_features),
]
)
# Pipeline
# - merge preprocess & model into one pipeline
pipeline = Pipeline([
# ('preprocessor', preprocessor),
('model', LGBMClassifier(
# categorical_feature=list(range(len(categorical_features))),
# early_stopping_rounds=300,
# test_size=0.2,
# eval_metric='auc',
# objective='binary',
random_state=RANDOM_STATE,
)),
])
# Parameters for `OptunaSearchCV`
parameters = {
# "model__is_unbalance": CategoricalDistribution([True, False]),
"model__objective": CategoricalDistribution(["binary"]),
"model__metric": CategoricalDistribution(["auc"]),
"model__learning_rate": LogUniformDistribution(1e-3, 1.0),
'model__n_estimators': CategoricalDistribution(range(2000, 5001, 500)),
'model__reg_alpha': LogUniformDistribution(1e-3, 10.0),
'model__reg_lambda': LogUniformDistribution(1e-3, 10.0),
'model__colsample_bytree': CategoricalDistribution(np.arange(0.1, 1.01, 0.1)),
'model__subsample': CategoricalDistribution(np.arange(0.1, 1.01, 0.1)),
'model__subsample_freq': IntUniformDistribution(1, 10),
'model__max_depth': IntUniformDistribution(1, 32),
'model__num_leaves' : IntUniformDistribution(2, 256),
'model__min_child_samples': IntUniformDistribution(1, 256),
'model__cat_smooth' : IntUniformDistribution(1, 128),
'model__max_bin' : IntUniformDistribution(512, 2048),
'model__cat_l2': IntUniformDistribution(1, 32),
}
# +
# parameters = {'model__is_unbalance': [False], 'model__objective': ['binary'], 'model__metric': ['auc'], 'model__learning_rate': [0.08], 'model__n_estimators': [4000], 'model__reg_alpha': [6.25], 'model__reg_lambda': [0.025], 'model__colsample_bytree': [0.2], 'model__subsample': [0.8], 'model__subsample_freq': [10], 'model__max_depth': [16], 'model__num_leaves': [128], 'model__min_child_samples': [100], 'model__cat_smooth': [88], 'model__max_bin': [666], 'model__cat_l2': [20]}
# -
grid_search = OptunaSearchCV(
pipeline,
param_distributions=parameters,
cv=5,
random_state=RANDOM_STATE,
n_jobs=-1,
scoring=auc,
)
grid_search.fit(X_train, y_train, model__categorical_feature=list(range(len(categorical_features))))
# preds = grid_search.best_estimator_.predict(X_valid)
preds = grid_search.best_estimator_.predict_proba(X_valid)[:, 1]
roc_auc_score(y_valid, preds)
# +
# onehot + variance_drop 0.8915464060319749
# onehot 0.8970355121797735
# codes 0.8826400343415913
# with label encoder + categorical_feature 0.8843085347397801
# with label codes + categorical_feature 0.8895766476594142
# -
abs(grid_search.best_score_)
grid_search.best_params_
# ### Submission
X_test = pd.read_csv(f"{data_dir}/interim/test_for_David.csv", index_col='id')
# X_test =pd.read_parquet(f"{data_dir}/interim/test.parq", engine='pyarrow').convert_dtypes()
display(X_test.shape)
X_test.head(2)
X_test = feature_engineering(X_test)
# X_test[categorical_features] = X_test[categorical_features].apply(lambda x: x.cat.codes).astype('int').astype('category')
preds_test = grid_search.best_estimator_.predict_proba(X_test)[:, 1]
output = pd.DataFrame(
{'Id': X_test.index, 'target': preds_test})
output.to_csv(f"{data_dir}/processed/submission.csv", index=False)
| tabular-playground-series/mar-2021/notebooks/pipeline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: wentao
# language: python
# name: wentao
# ---
# +
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torchvision import datasets, transforms
# torchvision是独立于pytorch的关于图像操作的一些方便工具库,主要包括一下几个包:
# vision.datasets : 几个常用视觉数据集,可以下载和加载
# vision.models : 流行的模型,例如 AlexNet, VGG, and ResNet以及训练好的参数。
# vision.transforms : 常用的图像操作,例如:随机切割,旋转等。
# vision.utils : 用于把形似 (3 x H x W) 的张量保存到硬盘中,给一个mini-batch的图像可以产生一个图像格网。
print("PyTorch Version: ",torch.__version__)
torch.manual_seed(53113)
use_cuda = torch.cuda.is_available()
device = torch.device("cuda:1" if use_cuda else "cpu")
# -
# 首先我们定义一个基于ConvNet的简单神经网络
# ## 1、加载数据
batch_size = test_batch_size = 32
kwargs = {'num_workers': 1, 'pin_memory': True} if use_cuda else {}
mnist_data = datasets.MNIST("./mnist_data", train=True, download=True,
transform=transforms.Compose([
transforms.ToTensor(),
]))
mnist_data
import numpy as np
data = [d[0].data.cpu().numpy() for d in mnist_data]
print(np.mean(data))
print(np.std(data))
#torch.utils.data.DataLoader在训练模型时使用到此函数,用来把训练数据分成多个batch(iterator)
#transform 接受一个图像返回变换后的图像的函数,相当于图像预处理
#常用的操作如 ToTensor, RandomCrop,Normalize等.
#他们可以通过transforms.Compose被组合在一起
#.ToTensor()将shape为(H, W, C)的nump.ndarray或img转为shape为(C, H, W)的tensor,
#其将每一个数值归一化到[0,1],其归一化方法比较简单,直接除以255即可。
#.Normalize作用就是.ToTensor将输入归一化到(0,1)后,再使用公式”(x-mean)/std”,将每个元素分布到(-1,1)
#shuffle随机打乱数据
#kwargs是上面gpu的设置
train_loader = torch.utils.data.DataLoader(
datasets.MNIST('./mnist_data',
train=True, #如果true,从training.pt创建数据集
transform=transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.1307,), (0.3081,)) # 所有图片像素均值和方差
])),
batch_size=batch_size,
shuffle=True,
**kwargs)
test_loader = torch.utils.data.DataLoader(
datasets.MNIST('./mnist_data',
train=False, #如果False,从test.pt创建数据集
transform=transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.1307,), (0.3081,))
])),
batch_size=test_batch_size,
shuffle=True,
**kwargs)
len(train_loader)
train_loader.dataset[0][0].shape
# # 2、定义CNN模型
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
#torch.nn.Conv2d(in_channels, out_channels, kernel_size, stride=1)
#in_channels:输入图像通道数,手写数字图像为1,一般彩色图像为3
#out_channels:输出通道数,等于卷积核的数量
#kernel_size:卷积核大小
#stride:步长
self.conv1 = nn.Conv2d(1, 20, 5, 1)
#上个卷积网络的out_channels,就是下一个网络的in_channels,所以这里是20
#out_channels:卷积核数量50
self.conv2 = nn.Conv2d(20, 50, 5, 1)
#全连接层torch.nn.Linear(in_features, out_features)
#in_features:输入特征维度,4*4*50是自己算出来的,跟输入图像维度有关
#out_features;输出特征维度
self.fc1 = nn.Linear(4*4*50, 500)
#输出维度10,10分类
self.fc2 = nn.Linear(500, 10)
def forward(self, x):
x = F.relu(self.conv1(x)) # x = (32,50,24,24)
x = F.max_pool2d(x, 2, 2) # x = (32,50,12,12)
x = F.relu(self.conv2(x)) # x = (32,50,8,8)
x = F.max_pool2d(x, 2, 2) # x = (32,50,4,4)
x = x.view(-1, 4*4*50) # x = (32,4*4*50)
x = F.relu(self.fc1(x)) # x = (32,4*4*50)*(4*4*50, 500)=(32,500)
x = self.fc2(x) # x = (32,500)*(500, 10)=(32,10)
return F.log_softmax(x, dim=1) #log probability
model = Net()
batch = next(iter(train_loader))
data = batch[0]
batch[1].shape
batch[0].shape
data = F.relu(model.conv1(data))
data = F.max_pool2d(data,2,2)
data = F.relu(model.conv2(data))
data = F.max_pool2d(data,2,2)
data = data.view(-1,4*4*50)
data = F.relu(model.fc1(data))
data = model.fc2(data)
data = F.log_softmax(data, dim=1)
data.shape
data.argmax(dim=1, keepdim=True)
pred = data.argmax(dim=1, keepdim=True)
target = batch[1]
correct = pred.eq(target.view_as(pred)).sum().item()
correct
# # 3、初始化模型和定义优化函数
lr = 0.01
momentum = 0.5
model = Net().to(device) #模型初始化
optimizer = optim.SGD(model.parameters(), lr=lr, momentum=momentum) #定义优化器
# # 4、定义训练和测试模型
# NLL loss的定义
#
# $\ell(x, y) = L = \{l_1,\dots,l_N\}^\top, \quad
# l_n = - w_{y_n} x_{n,y_n}, \quad
# w_{c} = \text{weight}[c] \cdot \mathbb{1}\{c \not= \text{ignore_index}\}$
def train(model, device, train_loader, optimizer, epoch, log_interval=10000):
model.train() #进入训练模式
for batch_idx, (data, target) in enumerate(train_loader):
#data:[32,1,28,28] target:[32]
data, target = data.to(device), target.to(device)
output = model(data) #[32,10]
#F.nll_loss(F.log_softmax(input), target) :
#单分类交叉熵损失函数,一张图片里只能有一个类别,输入input的需要softmax
#还有一种是多分类损失函数,一张图片有多个类别,输入的input需要sigmoid
loss = F.nll_loss(output, target) #这里loss求的是平均数,除以了batch
optimizer.zero_grad() #梯度归零
loss.backward()
optimizer.step()
if batch_idx % log_interval == 0:
print("Train Epoch: {} [{}/{} ({:0f}%)]\tLoss: {:.6f}".format(
epoch,
batch_idx * len(data),
len(train_loader.dataset), #60000
100. * batch_idx / len(train_loader), #len(train_loader)=60000/32=1875
loss.item()
))
def test(model, device, test_loader):
model.eval() #进入测试模式
test_loss = 0
correct = 0
with torch.no_grad():
for data, target in test_loader:
data, target = data.to(device), target.to(device)
output = model(data)
test_loss += F.nll_loss(output, target, reduction='sum').item()
#reduction='sum'代表batch内的每个元素loss累加求和,默认是mean求平均
pred = output.argmax(dim=1, keepdim=True)
correct += pred.eq(target.view_as(pred)).sum().item()
test_loss /= len(test_loader.dataset)
print('\nTest set: Average loss: {:.4f}, Accuracy: {}/{} ({:.0f}%)\n'.format(
test_loss, correct, len(test_loader.dataset),
100. * correct / len(test_loader.dataset)))
# # 5、查看运行结果
# + tags=[]
epochs = 2
for epoch in range(1, epochs + 1):
train(model, device, train_loader, optimizer, epoch)
test(model, device, test_loader)
save_model = True
if save_model:
torch.save(model.state_dict(),"mnist_cnn.pt")
#词典格式,model.state_dict()只保存模型参数
# -
# # CNN模型的迁移学习
# - 很多时候当我们需要训练一个新的图像分类任务,我们不会完全从一个随机的模型开始训练,而是利用预训练的模型来加速训练的过程。我们经常使用在`ImageNet`上的预训练模型。
# - 这是一种transfer learning的方法。我们常用以下两种方法做迁移学习。
# - fine tuning: 从一个预训练模型开始,我们改变一些模型的架构,然后继续训练整个模型的参数。
# - feature extraction: 我们不再改变与训练模型的参数,而是只更新我们修改的部分模型(fc)参数。我们之所以叫它feature extraction是因为我们把预训练的CNN模型当做一个特征提取模型,利用提取出来的特征做来完成我们的训练任务。
#
# 以下是构建和训练迁移学习模型的基本步骤:
# - 初始化预训练模型
# - 把最后一层的输出层改变成我们想要分的类别总数
# - 定义一个optimizer来更新参数
# - 模型训练
# +
import numpy as np
import torchvision
from torchvision import datasets, transforms, models
import matplotlib.pyplot as plt
import time
import os
import copy
print("Torchvision Version: ",torchvision.__version__)
# -
# 数据
# ------
#
# 我们会使用*hymenoptera_data*数据集,[下载](https://download.pytorch.org/tutorial/hymenoptera_data.zip).
#
# 这个数据集包括两类图片, **bees** 和 **ants**, 这些数据都被处理成了可以使用`ImageFolder <https://pytorch.org/docs/stable/torchvision/datasets.html#torchvision.datasets.ImageFolder>`来读取的格式。我们只需要把``data_dir``设置成数据的根目录,然后把``model_name``设置成我们想要使用的与训练模型:
# ::
# [resnet, alexnet, vgg, squeezenet, densenet, inception]
#
# 其他的参数有:
# - ``num_classes``表示数据集分类的类别数
# - ``batch_size``
# - ``num_epochs``
# - ``feature_extract``表示我们训练的时候使用fine tuning还是feature extraction方法。如果``feature_extract = False``,整个模型都会被同时更新。如果``feature_extract = True``,只有模型的最后一层被更新。
# 1、查看数据,只是查看作用
# ---------
#
# +
# Top level data directory. Here we assume the format of the directory conforms to the ImageFolder structure
data_dir = "./hymenoptera_data"
# Batch size for training (change depending on how much memory you have)
batch_size = 32
# 输入图像维度
input_size = 224
#os.path.join() 连接路径,相当于.../data_dir/train
all_imgs = datasets.ImageFolder(os.path.join(data_dir, "train"),
transforms.Compose([
transforms.RandomResizedCrop(input_size), #把每张图片截成resnet需要输入的维度224
transforms.RandomHorizontalFlip(),#水平翻转
transforms.ToTensor(),
]))
loader = torch.utils.data.DataLoader(all_imgs, batch_size=batch_size, shuffle=True, num_workers=4)
# -
len(next(iter(loader)))
img = next(iter(loader))[0] #这个img是一个batch的tensor
img.shape
# +
unloader = transforms.ToPILImage() # reconvert into PIL image
def imshow(tensor, title=None):
image = tensor.cpu().clone() # we clone the tensor to not do changes on it
image = unloader(image) #tensor转换成图像
plt.imshow(image)
if title is not None:
plt.title(title)
plt.show()
plt.figure()
imshow(img[8], title='Image')
# -
# 3、把训练集和验证集分batch转换成迭代器
# ---------
#
# 现在我们知道了模型输入的size,我们就可以把数据预处理成相应的格式。
#
# +
data_transforms = {
"train": transforms.Compose([
transforms.RandomResizedCrop(input_size),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225])
]),
"val": transforms.Compose([
transforms.Resize(input_size),
transforms.CenterCrop(input_size),#中间截取
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225])
]),
}
#train,val datasets dataloaders合为一个dict
# Create training and validation datasets
image_datasets = {x: datasets.ImageFolder(os.path.join(data_dir, x), data_transforms[x])
for x in ['train', 'val']}
# Create training and validation dataloaders
dataloaders_dict = {x: torch.utils.data.DataLoader(image_datasets[x],
batch_size=batch_size, shuffle=True, num_workers=4) for x in ['train', 'val']}
#把迭代器存放到字典里作为value,key是train和val,后面调用key即可。
print("Datasets and Dataloaders Initialized")
# -
inputs, labels=next(iter(dataloaders_dict["train"]))
print(inputs.shape)
print(labels)
#其实最后一个batch不足32
# ## 4、加载resnet模型并修改全连接层
num_classes = 2
num_epochs = 2
# Flag for feature extracting.
# When False, we finetune the whole model, when True we only update the reshaped layer params
feature_extract = True #只更新修改的层
def set_parameter_requires_grad(model, feature_extracting):
if feature_extracting:
for param in model.parameters():
param.requires_grad = False
# + jupyter={"outputs_hidden": true} tags=[]
def initialize_model(num_classes, feature_extract, use_pretrained=True):#如果True,返回预训练好的模型和参数
model_ft = models.resnet18(pretrained=use_pretrained)
set_parameter_requires_grad(model_ft, feature_extract)
num_ftrs = model_ft.fc.in_features
#model_ft.fc是resnet的最后全连接层
#(fc): Linear(in_features=512, out_features=1000, bias=True)
#in_features 是全连接层的输入特征维度
model_ft.fc = nn.Linear(num_ftrs, num_classes)
input_size = 224 #resnet网络输入图片维度是224
return model_ft, input_size
model_ft, input_size = initialize_model(num_classes, feature_extract, use_pretrained=True)
print(model_ft)
# -
model_ft.fc.weight.requires_grad
model_ft.layer1[0].conv1.weight.requires_grad
models.resnet18().fc
model_ft.fc
# ## 5、查看需要更新的参数、定义优化器
# + jupyter={"outputs_hidden": true} tags=[]
next(iter(model_ft.named_parameters()))
# -
len(next(iter(model_ft.named_parameters()))) #是元组,只有两个值
# + jupyter={"outputs_hidden": true}
for name,param in model_ft.named_parameters():
print(name) #看下都有哪些参数
# + tags=[]
model_ft = model_ft.to(device)
# Gather the parameters to be optimized/updated in this run. If we are finetuning
# we will be updating all parameters. However, if we are doing
# feature extract method, we will only update the parameters
# that we have just initialized, i.e. the parameters with requires_grad is True.
params_to_update = model_ft.parameters() #需要更新的参数
print("Params to learn:")
if feature_extract:
params_to_update = [] #需要更新的参数存放在此
for name,param in model_ft.named_parameters():
if param.requires_grad == True:
params_to_update.append(param)
print("\t",name)
else:
for name,param in model_ft.named_parameters():
if param.requires_grad == True:
print("\t",name)
# -
optimizer_ft = optim.SGD(params_to_update, lr=0.001, momentum=0.9)
criterion = nn.CrossEntropyLoss()
data = next(iter(dataloaders_dict['train']))
inputs = data[0]
lables = data[1]
inputs = inputs.cuda(1)
model_ft(inputs).shape
# ## 6、定义训练函数
def train_eval_model(model, dataloaders, criterion, optimizer, num_epochs=5):
start = time.time()
val_acc_history = []
best_model_wts = copy.deepcopy(model.state_dict())
best_acc = 0.
for epoch in range(num_epochs):
print("Epoch {}/{}".format(epoch, num_epochs-1))
print("-"*10)
for phase in ["train", "val"]:
running_loss = 0.
running_corrects = 0.
if phase == "train":
model.train()
elif phase == 'val':
model.eval()
for inputs, labels in dataloaders[phase]:
inputs = inputs.to(device)
labels = labels.to(device)
with torch.autograd.set_grad_enabled(phase=="train"):
#torch.autograd.set_grad_enabled梯度管理器,可设置为打开或关闭
#phase=="train"返回值True和False,双等号要注意
outputs = model(inputs)
loss = criterion(outputs, labels)
_, preds = torch.max(outputs, 1)
#返回每一行最大的数和索引,preds的位置是索引的位置
#也可以preds = outputs.argmax(dim=1)
if phase == "train":
optimizer.zero_grad()
loss.backward()
optimizer.step()
running_loss += loss.item() * inputs.size(0) #交叉熵损失函数是平均过的
running_corrects += torch.sum(preds.view(-1) == labels.view(-1)).item()
epoch_loss = running_loss / len(dataloaders[phase].dataset)
epoch_acc = running_corrects / len(dataloaders[phase].dataset)
print("{} Loss: {} Acc: {}".format(phase, epoch_loss, epoch_acc))
if phase == "val" and epoch_acc > best_acc:
best_acc = epoch_acc
best_model_wts = copy.deepcopy(model.state_dict())
#模型变好,就拷贝更新后的模型参数
if phase == "val":
val_acc_history.append(epoch_acc) #记录每个epoch验证集的准确率
time_elapsed = time.time() - start
print("Training compete in {}m {}s".format(time_elapsed // 60, time_elapsed % 60))
print("Best val Acc: {}".format(best_acc))
model.load_state_dict(best_model_wts) #把最新的参数复制到model中
return model, val_acc_history
# ## 7、运行模型
# Train and evaluate
model_ft, ohist = train_eval_model(model_ft, dataloaders_dict, criterion, optimizer_ft, num_epochs=num_epochs)
ohist
# + jupyter={"outputs_hidden": true}
model_ft
# -
# Initialize the non-pretrained version of the model used for this run
scratch_model,_ = initialize_model(model_name,
num_classes,
feature_extract=False, #所有参数都训练
use_pretrained=False)# 不要imagenet的参数
scratch_model = scratch_model.to(device)
scratch_optimizer = optim.SGD(scratch_model.parameters(),
lr=0.001, momentum=0.9)
scratch_criterion = nn.CrossEntropyLoss()
_,scratch_hist = train_eval_model(scratch_model,
dataloaders_dict,
scratch_criterion,
scratch_optimizer,
num_epochs=num_epochs)
# +
# Plot the training curves of validation accuracy vs. number
# of training epochs for the transfer learning method and
# the model trained from scratch
# ohist = []
# shist = []
# ohist = [h.cpu().numpy() for h in ohist]
# shist = [h.cpu().numpy() for h in scratch_hist]
plt.title("Validation Accuracy vs. Number of Training Epochs")
plt.xlabel("Training Epochs")
plt.ylabel("Validation Accuracy")
plt.plot(range(1,num_epochs+1),ohist,label="Pretrained")
plt.plot(range(1,num_epochs+1),scratch_hist,label="Scratch")
plt.ylim((0,1.))
plt.xticks(np.arange(1, num_epochs+1, 1.0))
plt.legend()
plt.show()
| 5/CNN-Image-Classification-comments.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={} tags=["awesome-notebooks/Naas/Naas_Notification_demo.ipynb"]
# <img width="10%" alt="Naas" src="https://landen.imgix.net/jtci2pxwjczr/assets/5ice39g4.png?w=160"/>
#
# + [markdown] papermill={} tags=["awesome-notebooks/Naas/Naas_Notification_demo.ipynb"]
# # Naas - Notification demo
# <a href="https://app.naas.ai/user-redirect/naas/downloader?url=https://raw.githubusercontent.com/jupyter-naas/awesome-notebooks/master/Naas/Naas_Notification_demo.ipynb" target="_parent"><img src="https://naasai-public.s3.eu-west-3.amazonaws.com/open_in_naas.svg"/></a>
# + [markdown] papermill={} tags=["awesome-notebooks/Naas/Naas_Notification_demo.ipynb"]
# **Tags:** #naas #notification #snippet #operations
# + [markdown] papermill={} tags=["awesome-notebooks/Naas/Naas_Notification_demo.ipynb"]
# **Author:** [<NAME>](https://www.linkedin.com/in/ACoAAAJHE7sB5OxuKHuzguZ9L6lfDHqw--cdnJg/)
# + [markdown] papermill={} tags=["awesome-notebooks/Naas/Naas_Notification_demo.ipynb"]
# Read the doc: https://docs.naas.ai/features/scheduler
# <br>
# Note: use ''' around your content for HTML
# + [markdown] papermill={} tags=["awesome-notebooks/Naas/Naas_Notification_demo.ipynb"]
# ## Input
# + [markdown] papermill={} tags=["awesome-notebooks/Naas/Naas_Notification_demo.ipynb"]
# ### Import library
# + papermill={} tags=["awesome-notebooks/Naas/Naas_Notification_demo.ipynb"]
import naas
# + [markdown] papermill={} tags=["awesome-notebooks/Naas/Naas_Notification_demo.ipynb"]
# ## Model
# + [markdown] papermill={} tags=["awesome-notebooks/Naas/Naas_Notification_demo.ipynb"]
# ### Notification informations
# + papermill={} tags=["awesome-notebooks/Naas/Naas_Notification_demo.ipynb"]
email_to = "<EMAIL>"
subject = "🛎️ Naas Notification Test 🚨"
content ='''<p>If i can see an image below this text... </p>
<p><img src="https://specials-images.forbesimg.com/imageserve/5f1f37a40a5db2c8275972c0/960x0.jpg?fit=scale" alt="" width="959" height="663" /></p><br>
...it means everything goes well.'''
# + [markdown] papermill={} tags=["awesome-notebooks/Naas/Naas_Notification_demo.ipynb"]
# ## Output
# + [markdown] papermill={} tags=["awesome-notebooks/Naas/Naas_Notification_demo.ipynb"]
# ### Send the notification
# + papermill={} tags=["awesome-notebooks/Naas/Naas_Notification_demo.ipynb"]
naas.notification.send(email_to, subject, content)
| Naas/Naas_Notification_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Code for the midway progress report. Should accomplish the following objectives:
#
# - Load data and identify useful features
# - Use decision trees to find optimal bins for all continuous features
# - Apply various transformations to continuous variables
# - Run linear regression model on full dataset with all engineered features
# - Visualize model performance
# +
# %matplotlib inline
import dataset
import numpy as np
import pandas as pd
import sklearn.linear_model
import sklearn.pipeline
import sklearn.preprocessing
import sklearn.model_selection
import sklearn.tree
import pydotplus
from IPython.display import Image
from io import StringIO
from tqdm import tqdm
# -
# # Load data and identify useful features
# Connect to the database and query the data.
db = dataset.connect('postgres:///steam')
# +
query = '''
SELECT
steam_app_id,
game_name,
reviews_last_30_days,
pct_positive_reviews_last_30_days,
reviews_all_time,
pct_positive_reviews_all_time,
release_date,
title,
developer,
publisher,
num_achievements,
full_price,
metacritic_score,
genres,
details,
tags
FROM
game_crawl_view
WHERE
is_dlc = FALSE
AND pct_positive_reviews_all_time IS NOT NULL
AND short_description IS NOT NULL
AND short_description != ''
AND long_description IS NOT NULL
AND long_description != '';
'''
data = [r for r in db.query(query)]
df = pd.DataFrame(data)
df.index = df['steam_app_id']
df.drop('steam_app_id', axis=1, inplace=True)
display(df.head())
display(df.shape)
# -
# Create dummies for list variables (such as genre; each game can have multiple genres, where we really just need one-hot encodings for all genres).
# +
def create_dummies_from_list(df, col, db_table):
all_values = {r['descr'] for r in db[db_table].find()}
agg_series = df[col].apply(lambda x: [] if x is None else x)
df.drop(col, axis=1, inplace=True)
for val in all_values:
df['{}|{}'.format(col, val)] = agg_series.apply(lambda x: val in x)
create_dummies_from_list(df, 'genres', 'steam_genre')
create_dummies_from_list(df, 'details', 'steam_game_detail')
create_dummies_from_list(df, 'tags', 'steam_tag')
display(df.head())
display(df.shape)
# -
# Create dummies for regular categorical variables. However, these have way too many distinct values, so only save the one-hot encodings for values more frequent than 10 (i.e., only make a specific publisher a feature if they've published >= 10 games in our dataset).
# +
def create_dummies(df, col, min_num=10):
series = df[col]
df.drop(col, axis=1, inplace=True)
counts = series.value_counts()
filtered_counts = counts[counts >= min_num]
filtered_series = series.apply(lambda x: x if x in filtered_counts.index else np.NaN)
return df.merge(pd.get_dummies(filtered_series, prefix=col, prefix_sep='|'), left_index=True, right_index=True)
df = create_dummies(df, 'publisher')
df = create_dummies(df, 'developer')
# -
# Some variables have missing values that should represent 0. Make that replacement here.
for col in ['reviews_last_30_days', 'num_achievements', 'full_price']:
df[col].fillna(value=0, inplace=True)
# The timestamp probably won't be a useful feature, but convert it to a float so it's at least usable in case it becomes one. There are only a few games with null release date (38); drop them to avoid fitting the model on a null timestamp.
# +
df = df[~pd.isnull(df['release_date'])]
# Divide by 10^9 to get seconds since the epoch instead of nanoseconds
df.loc[:, 'release_date'] = (pd.to_datetime(df['release_date']).astype(np.int64) / 10**9).astype(np.float64)
# -
# Convert integer continuous variables to floats so they work well with our models later.
df.loc[:, 'reviews_all_time'] = df['reviews_all_time'].astype(np.float64)
# Remove the outcome and other related vars from the set of features to prepare for analysis.
# +
y = df['pct_positive_reviews_all_time']
for col in ('pct_positive_reviews_last_30_days', 'pct_positive_reviews_all_time',
'game_name', 'title', 'metacritic_score'):
df.drop(col, axis=1, inplace=True)
# -
df_train, df_test, y_train, y_test = sklearn.model_selection.train_test_split(df, y, test_size=0.2,
random_state=1337)
# Final feature set for analysis (until text features are added):
display(df_train.shape)
display(df_test.shape)
# # Use decision trees to find optimal bins for all continuous predictors
# When dealing with continuous variables that have a nonlinear relationship with the outcome, sometimes setting up discrete bins is the best way to utilize the feature in the model. Deciding how to best bin the variables can be challenging; unsupervised methods, such as equal intervals or equal interval width, can easily miss patterns in the data.
#
# One systematic way to develop optimal bins is to train a decision tree for each predictor. The split criteria of the decision tree will empirically determine the best way to divide up the variable values. We do that here for all continuous variables (and will use this technique again when we add word counts, which are technically discrete but will be treated as continuous due to their many possible values).
# +
def display_cv_results(grid_search, max_rows=10):
display(pd.DataFrame(grid_search.cv_results_)
.sort_values('mean_test_score', ascending=False)[:max_rows])
def find_optimal_bins(df_train, y_train, variable, display_report=False):
'''
Given our data and a variable, do the following:
1) Train a cross-validated decision tree using that variable and the outcome only
2) Return the trained tree, which can be used to apply the same bins onto new data.
'''
pipeline = sklearn.pipeline.Pipeline((
('scaler', sklearn.preprocessing.MinMaxScaler()),
('clf', sklearn.tree.DecisionTreeRegressor(presort=True))
))
param_grid = {
'clf__criterion': ('mse', 'friedman_mse', 'mae'),
'clf__min_impurity_decrease': (8e-1, 7e-1, 6.5e-1, 6.25e-1, 6e-1, 5.75e-1, 5.5e-1,
5e-1, 4.75e-1, 4.5e-1, 4.25e-1, 4e-1, 3.75e-1, 3.5e-1,
3e-1, 2.5e-1, 2.25e-1, 2e-1, 1.75e-1,
1.5e-1, 1e-1, 5e-2, 1e-2, 5e-3, 1e-3),
}
grid_search = sklearn.model_selection.GridSearchCV(
pipeline, param_grid=param_grid, scoring='neg_mean_squared_error',
n_jobs=-1, cv=3, return_train_score=True)
X = df_train[[variable]].as_matrix()
y = y_train.as_matrix()
grid_search.fit(X, y)
if display_report:
display_cv_results(grid_search)
return grid_search.best_estimator_
def apply_bins(pipeline, variable, df):
'''
Given a trained model, apply its bins to the column identified in the given dataset and
return the result.
'''
transformed_values = pipeline.named_steps.scaler.transform(df[[variable]])
return pipeline.named_steps.clf.apply(transformed_values)
def draw_tree(clf, variable):
dot_data = StringIO()
sklearn.tree.export_graphviz(clf, out_file=dot_data,
filled=True, rounded=True,
special_characters=True,
feature_names=['variable'],
node_ids=True)
graph = pydotplus.graph_from_dot_data(dot_data.getvalue())
display(Image(graph.create_png()))
# -
# Run the bin-finding for all the continuous vars. Output the CV results so we can expand the parameter grid if needed, and visualize each tree so we can confirm they make sense.
# +
continuous_vars = (
'full_price',
'reviews_last_30_days',
'reviews_all_time',
'release_date',
'num_achievements',
)
pipelines = {}
for var in continuous_vars:
print('Training tree for {}'.format(var))
pipeline = find_optimal_bins(df_train, y_train, var, display_report=True)
pipelines[var] = pipeline
clf = pipeline.named_steps.clf
draw_tree(clf, var)
for df in (df_train, df_test):
df['{}_binned'.format(var)] = apply_bins(pipeline, var, df)
# -
# # Apply various transformations to continuous variables
#
# Apply some common transformations to continuous variables in case they have a nonlinear relationship with the outcome.
for var in continuous_vars:
for df in (df_train, df_test):
df['{}_squared'.format(var)] = np.square(df[var])
# Add two for log transform to prevent numeric warnings
df['{}_log'.format(var)] = np.log(df[var]+2)
df['{}_loglog'.format(var)] = np.log(df['{}_log'.format(var)])
df['{}_sqrt'.format(var)] = np.sqrt(df[var])
df['{}_indicator'.format(var)] = df[var].apply(lambda x: 1 if x > 0
else 0)
for winsor_pctile in (90, 95, 99):
winsor_threshold = np.percentile(df[var], winsor_pctile)
df['{}_winsor{}'.format(var, winsor_pctile)] = df[var].apply(
lambda x: winsor_threshold if x > winsor_threshold
else x)
# # Run linear regression model on full dataset with all engineered features
#
# Use the engineered features in a regression model.
# +
pipeline = sklearn.pipeline.Pipeline((
('scaler', sklearn.preprocessing.MinMaxScaler()),
('clf', sklearn.linear_model.Lasso(random_state=7, max_iter=100000)),
))
param_grid = {
'clf__alpha': [0.1, 0.05, 0.025, 0.01, 0.0075, 0.006, 0.005, 0.004, 0.003, 0.0025]
}
grid_search = sklearn.model_selection.GridSearchCV(
pipeline, param_grid=param_grid, scoring='neg_mean_squared_error',
n_jobs=-1, cv=3, return_train_score=True)
grid_search.fit(df_train, y_train)
display_cv_results(grid_search)
# -
# # Visualize model performance
#
# Visualizing regression results in a high-dimensional space is tricky, so we'll try to find a few ways to look at the predictions.
# Plot the residuals to see if their distribution appears to be random.
# +
predictions = grid_search.predict(df_test)
residuals = predictions - y_test
residual_df = pd.DataFrame({'prediction': predictions, 'residual': residuals})
# There are a few (3) extreme outliers due to some highly inflated coefficients; ignore them or
# the graphs are too zoomed out to see anything
# Hopefully these extreme cases will go away when we use a more powerful model
display(residual_df[(residual_df['prediction'] >= 150) | (residual_df['prediction'] < 0)])
residual_df = residual_df[(residual_df['prediction'] <= 150) & (residual_df['prediction'] >= 0)]
residual_df.plot(x='prediction', y='residual', kind='scatter')
# -
# There are sharp lines along the boundaries of the actual data (since a proportion is bounded between 0 and 1), but the residuals otherwise look randomly distributed.
#
# Plot the distribution of predicted values and distribution of actual values to see whether the model is closely approximating the real distribution.
y_test.plot(kind='hist', title='Actual Distribution of Positive Review Proportion',
ylim=(0, 700), xlim=(0,120))
residual_df['prediction'].plot(kind='hist', title='Predicted Distribution of Positive Review Proportion',
ylim=(0, 700), xlim=(0, 120))
# The distributions peak around the same area, but the assumption of a normal distribution inherent in ordinary least-squares regression is clearly incorrect. We hope to more accurately match the true data distribution using more advanced techniques later.
#
# Finally, check the R^2 value for our model on our test dataset.
grid_search.best_estimator_.named_steps.clf.score(df_test, y_test)
# The value is huge because of a ton of variance in the test set that wasn't present in the training set. Hopefully, we can improve on this with a better model.
| Midway Code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Function Practice Exercises
#
# Problems are arranged in increasing difficulty:
# * Warmup - these can be solved using basic comparisons and methods
# * Level 1 - these may involve if/then conditional statements and simple methods
# * Level 2 - these may require iterating over sequences, usually with some kind of loop
# * Challenging - these will take some creativity to solve
# ## WARMUP SECTION:
# #### LESSER OF TWO EVENS: Write a function that returns the lesser of two given numbers *if* both numbers are even, but returns the greater if one or both numbers are odd
# lesser_of_two_evens(2,4) --> 2
# lesser_of_two_evens(2,5) --> 5
def lesser_of_two_evens(a,b):
if a%2 == 0 and b%2 == 0:
# Both numbers are even
# easier to use the min and max functions
# result = min(a,b)
# make it more tidy use return already here
return min(a,b)
# if a < b: option 1
# result = a
# else:
# result = b
else:
# One or both numbers are odd
# easier to use the min and max functions
# result = max(a,b)
# make it more tidy use return already here
return max(a,b)
# if a > b: option 1
# result = a
# else:
# result = b
#return result
# return a my code
# else:
# if a%2 != 0 or b%2 != 0:
# return b
# Check
lesser_of_two_evens(2,4)
# Check
lesser_of_two_evens(2,5)
# #### ANIMAL CRACKERS: Write a function takes a two-word string and returns True if both words begin with same letter
# animal_crackers('Levelheaded Llama') --> True
# animal_crackers('Crazy Kangaroo') --> False
# +
def animal_crackers(text):
# We can use the lower function if the both letters are the same but camel case
wordlist = text.lower().split()
#print(wordlist)
# first = wordlist[0]
# second = wordlist[1]
# make it more tidy move the wordlist indexing down
return wordlist[0][0] == wordlist[1][0]
# for letter in text.split(): my attempt
# print(letter)
# if text[0] = letter[0]:
# return True
# else:
# return False
# -
# Check
animal_crackers('Levelheaded Llama')
# Check
animal_crackers('Crazy Kangaroo')
# #### MAKES TWENTY: Given two integers, return True if the sum of the integers is 20 *or* if one of the integers is 20. If not, return False
#
# makes_twenty(20,10) --> True
# makes_twenty(12,8) --> True
# makes_twenty(2,3) --> False
def makes_twenty(n1,n2):
# First pass with Jose
# if n1 + n2 == 20:
# return True
# elif n1 == 20:
# return True
# elif n2 == 20:
# return True
# else:
# return False
# Second pass all on a single pline when using booleans
return (n1+n2) == 20 or n1 == 20 or n2 == 20
# Check
makes_twenty(20,10)
# Check
makes_twenty(2,3)
# # LEVEL 1 PROBLEMS
# #### OLD MACDONALD: Write a function that capitalizes the first and fourth letters of a name
#
# old_macdonald('macdonald') --> MacDonald
#
# Note: `'macdonald'.capitalize()` returns `'Macdonald'`
# +
def old_macdonald(name):
first_half = name[:3]
second_half = name[3:]
return first_half.capitalize() + second_half.capitalize()
# first attempt
# first_letter = name[0]
# inbetween = name[1:3]
# fourth_letter= name [3]
# rest = name[4:]
# return first_letter.upper() + inbetween + fourth_letter.upper() + rest
# -
# Check
old_macdonald('macdonald')
# #### MASTER YODA: Given a sentence, return a sentence with the words reversed
#
# master_yoda('I am home') --> 'home am I'
# master_yoda('We are ready') --> 'ready are We'
#
# Note: The .join() method may be useful here. The .join() method allows you to join together strings in a list with some connector string. For example, some uses of the .join() method:
#
# >>> "--".join(['a','b','c'])
# >>> 'a--b--c'
#
# This means if you had a list of words you wanted to turn back into a sentence, you could just join them with a single space string:
#
# >>> " ".join(['Hello','world'])
# >>> "Hello world"
def master_yoda(text):
wordlist = text.split()
reverse_word_list = wordlist[::-1]
return ' '.join(reverse_word_list)
# Check
master_yoda('I am home')
# Check
master_yoda('We are ready')
# to make a string we use the join method
mylist = ['a','b','c']
# it concatenates the list into a string
''.join(mylist)
# #### ALMOST THERE: Given an integer n, return True if n is within 10 of either 100 or 200
#
# almost_there(90) --> True
# almost_there(104) --> True
# almost_there(150) --> False
# almost_there(209) --> True
#
# NOTE: `abs(num)` returns the absolute value of a number
def almost_there(n):
return (abs(100-n) <=10) or (abs(200-n) <=10)
# Check
almost_there(104)
# Check
almost_there(150)
# Check
almost_there(209)
# # LEVEL 2 PROBLEMS
# #### FIND 33:
#
# Given a list of ints, return True if the array contains a 3 next to a 3 somewhere.
#
# has_33([1, 3, 3]) → True
# has_33([1, 3, 1, 3]) → False
# has_33([3, 1, 3]) → False
def has_33(nums):
# Jose first example
for i in range(0,len(nums)-1):
# another option is to add slices and compare, not as readable but a bit more slick
# if nums[i:i+2] == [3,3]
if nums[i] == 3 and nums[i+1] == 3:
return True
return False
# Check
has_33([1, 3, 3])
# Check
has_33([1, 3, 1, 3])
# Check
has_33([3, 1, 3])
# #### PAPER DOLL: Given a string, return a string where for every character in the original there are three characters
# paper_doll('Hello') --> 'HHHeeellllllooo'
# paper_doll('Mississippi') --> 'MMMiiissssssiiippppppiii'
def paper_doll(text):
return ''.join([letter*3 for letter in text])
# jose examples
# result = ''
# for char in text:
# result+= char*3
# return result
# Check
paper_doll('Hello')
# Check
paper_doll('Mississippi')
# #### BLACKJACK: Given three integers between 1 and 11, if their sum is less than or equal to 21, return their sum. If their sum exceeds 21 *and* there's an eleven, reduce the total sum by 10. Finally, if the sum (even after adjustment) exceeds 21, return 'BUST'
# blackjack(5,6,7) --> 18
# blackjack(9,9,9) --> 'BUST'
# blackjack(9,9,11) --> 19
def blackjack(a,b,c):
if sum([a,b,c]) <= 21:
return sum([a,b,c])
elif 11 in [a,b,c] and sum([a,b,c]) <= 31:
return sum([a,b,c])-10
else:
return "BUST"
# Check
blackjack(5,6,7)
# Check
blackjack(9,9,9)
# Check
blackjack(9,9,11)
# #### SUMMER OF '69: Return the sum of the numbers in the array, except ignore sections of numbers starting with a 6 and extending to the next 9 (every 6 will be followed by at least one 9). Return 0 for no numbers.
#
# summer_69([1, 3, 5]) --> 9
# summer_69([4, 5, 6, 7, 8, 9]) --> 9
# summer_69([2, 1, 6, 9, 11]) --> 14
def summer_69(arr):
total = 0
# by def I will add all the numbers if its true
add = True
for num in arr:
while add:
if num != 6:
total += num
break
else:
add = False
while not add:
if num != 9:
break
else:
add = True
return total
# Check
summer_69([1, 3, 5])
# Check
summer_69([4, 5, 6, 7, 8, 9])
# Check
summer_69([2, 1, 6, 9, 11])
# # CHALLENGING PROBLEMS
# #### SPY GAME: Write a function that takes in a list of integers and returns True if it contains 007 in order
#
# spy_game([1,2,4,0,0,7,5]) --> True
# spy_game([1,0,2,4,0,5,7]) --> True
# spy_game([1,7,2,0,4,5,0]) --> False
#
def spy_game(nums):
code = [0,0,7,'x']
for num in nums:
# Iterate through the code list and pop off index at zero each time
# [0,7,'x']
#[7,'x']
# ['x'] length=1
if num == code[0]:
code.pop(0)
return len(code) == 1
# Check
spy_game([1,2,4,0,0,7,5])
# Check
spy_game([1,0,2,4,0,5,7])
# Check
spy_game([1,7,2,0,4,5,0])
# #### COUNT PRIMES: Write a function that returns the *number* of prime numbers that exist up to and including a given number
# count_primes(100) --> 25
#
# By convention, 0 and 1 are not prime.
def count_primes(num):
# check for 0 or 1 input
# this is a challenge math problem and programming
# euler project website
if num < 2:
return 0
##############
# 2 or greater
##############
# Store our prime numbers
primes = [2]
# counter going upto input num
x = 3
# x is going through every number upto input
while x <= num:
# check if x is prime
#for y in range(3,x,2):
# another check is use the prime numbers we have collected to check them
for y in primes:
if x%y == 0:
# Skipping over the even numbers
x += 2
break
else:
primes.append(x)
x += 2
print(primes)
return len(primes)
# Check
count_primes(100)
# ### Just for fun:
# #### PRINT BIG: Write a function that takes in a single letter, and returns a 5x5 representation of that letter
# print_big('a')
#
# out: *
# * *
# *****
# * *
# * *
# HINT: Consider making a dictionary of possible patterns, and mapping the alphabet to specific 5-line combinations of patterns. <br>For purposes of this exercise, it's ok if your dictionary stops at "E".
def print_big(letter):
patterns = {1:' * ',2:' * * ',3:'* *',4:'*****',5:'**** ',6:' * ',7:' * ',8:'* * ',9:'* '}
alphabet = {'A':[1,2,4,3,3],'B':[5,3,5,3,5],'C':[4,9,9,9,4],'D':[5,3,3,3,5],'E':[4,9,4,9,4]}
for pattern in alphabet[letter.upper()]:
print(patterns[pattern])
print_big('a')
# ## Great Job!
| 03-Methods and functions/03-Function Practice Exercises.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + active=""
# .. _fodselsnummer_userguide:
#
# fodselsnummer Strings
# ============
# + active=""
# Introduction
# ------------
#
# The function :func:`clean_no_fodselsnummer() <dataprep.clean.clean_no_fodselsnummer.clean_no_fodselsnummer>` cleans a column containing Norwegian birth number (fodselsnummer) strings, and standardizes them in a given format. The function :func:`validate_no_fodselsnummer() <dataprep.clean.clean_no_fodselsnummer.validate_no_fodselsnummer>` validates either a single fodselsnummer strings, a column of fodselsnummer strings or a DataFrame of fodselsnummer strings, returning `True` if the value is valid, and `False` otherwise.
# -
# fodselsnummer strings can be converted to the following formats via the `output_format` parameter:
#
# * `compact`: only number strings without any seperators or whitespace, like "15108695088"
# * `standard`: fodselsnummer strings with proper whitespace in the proper places, like "151086 95088"
# * `birthdate`: get the person's birthdate, like "1986-10-15".
# * `gender`: get the person's birth gender ('M' or 'F').
#
# Invalid parsing is handled with the `errors` parameter:
#
# * `coerce` (default): invalid parsing will be set to NaN
# * `ignore`: invalid parsing will return the input
# * `raise`: invalid parsing will raise an exception
#
# The following sections demonstrate the functionality of `clean_no_fodselsnummer()` and `validate_no_fodselsnummer()`.
# ### An example dataset containing fodselsnummer strings
import pandas as pd
import numpy as np
df = pd.DataFrame(
{
"fodselsnummer": [
'15108695088',
'15108695077',
"999 999 999",
"004085616",
"002 724 334",
"hello",
np.nan,
"NULL",
],
"address": [
"123 Pine Ave.",
"main st",
"1234 west main heights 57033",
"apt 1 789 s maple rd manhattan",
"robie house, 789 north main street",
"1111 S Figueroa St, Los Angeles, CA 90015",
"(staples center) 1111 S Figueroa St, Los Angeles",
"hello",
]
}
)
df
# ## 1. Default `clean_no_fodselsnummer`
#
# By default, `clean_no_fodselsnummer` will clean fodselsnummer strings and output them in the standard format with proper separators.
from dataprep.clean import clean_no_fodselsnummer
clean_no_fodselsnummer(df, column = "fodselsnummer")
# ## 2. Output formats
# This section demonstrates the output parameter.
# ### `standard` (default)
clean_no_fodselsnummer(df, column = "fodselsnummer", output_format="standard")
# ### `compact`
clean_no_fodselsnummer(df, column = "fodselsnummer", output_format="compact")
# ### `birthdate`
clean_no_fodselsnummer(df, column = "fodselsnummer", output_format="birthdate")
# ### `gender`
clean_no_fodselsnummer(df, column = "fodselsnummer", output_format="gender")
# ## 3. `inplace` parameter
#
# This deletes the given column from the returned DataFrame.
# A new column containing cleaned fodselsnummer strings is added with a title in the format `"{original title}_clean"`.
clean_no_fodselsnummer(df, column="fodselsnummer", inplace=True)
# ## 4. `errors` parameter
# ### `coerce` (default)
clean_no_fodselsnummer(df, "fodselsnummer", errors="coerce")
# ### `ignore`
clean_no_fodselsnummer(df, "fodselsnummer", errors="ignore")
# ## 4. `validate_no_fodselsnummer()`
# `validate_no_fodselsnummer()` returns `True` when the input is a valid fodselsnummer. Otherwise it returns `False`.
#
# The input of `validate_no_fodselsnummer()` can be a string, a Pandas DataSeries, a Dask DataSeries, a Pandas DataFrame and a dask DataFrame.
#
# When the input is a string, a Pandas DataSeries or a Dask DataSeries, user doesn't need to specify a column name to be validated.
#
# When the input is a Pandas DataFrame or a dask DataFrame, user can both specify or not specify a column name to be validated. If user specify the column name, `validate_no_fodselsnummer()` only returns the validation result for the specified column. If user doesn't specify the column name, `validate_no_fodselsnummer()` returns the validation result for the whole DataFrame.
from dataprep.clean import validate_no_fodselsnummer
print(validate_no_fodselsnummer("15108695088"))
print(validate_no_fodselsnummer("15108695077"))
print(validate_no_fodselsnummer("999 999 999"))
print(validate_no_fodselsnummer("51824753556"))
print(validate_no_fodselsnummer("004085616"))
print(validate_no_fodselsnummer("hello"))
print(validate_no_fodselsnummer(np.nan))
print(validate_no_fodselsnummer("NULL"))
# ### Series
validate_no_fodselsnummer(df["fodselsnummer"])
# ### DataFrame + Specify Column
validate_no_fodselsnummer(df, column="fodselsnummer")
# ### Only DataFrame
validate_no_fodselsnummer(df)
| docs/source/user_guide/clean/clean_no_fodselsnummer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
##
# Advent of code 2019, AoC day 16 puzzle 1
# This solution (python3.7 jupyter notebook) by kannix68, @ 2020-01-06.
import sys
sys.path.insert(0, '..') # allow import from parent dir
from typing import Dict, List, Tuple
import lib.aochelper as aoc
from lib.aochelper import map_e
# -
import logging
logging.basicConfig(stream=sys.stdout, level=logging.INFO)
log = logging.getLogger(__name__)
## PROBLEM DOMAIN code
import itertools
from timeit import default_timer as timer # performance timing measurement
def phase_transform(ins: str, ptrn: List[int], num_phases = 1) -> str:
tmp = ins
for iter in range(num_phases):
if iter > 0:
tmp = outs
outdigs = []
for digidx in range(len(ins)):
digptrn = []
for i in ptrn:
for fac in range(digidx+1):
digptrn.append(i)
#log.debug(f"digid={digidx}, digptrn={digptrn}")
outdig_compositor = []
offset = 1 # number of elements to consume from (cycled) pattern before acting on input
for idx, val in enumerate(itertools.cycle(digptrn), -offset):
if idx == len(tmp):
break
if idx < 0:
continue
#log.debug(f" {idx}, ins-dig={ins[idx]} ptrn:{val}")
outdig_compositor.append( int(tmp[idx])*val )
dig = abs(sum(outdig_compositor))%10
outdigs.append(dig)
outs = ''.join(map_e(str, outdigs))
#log.debug(f"phase_transform #{iter+1}: outdigs={outs} via {len(ins)} iters from ptrn={aoc.cl(ptrn)}, input#={tmp}")
log.info(f"phase_transform iter={iter+1} out={outs} via {len(ins)} iters from ptrn={aoc.cl(ptrn)}, input0={ins[:40]}...")
return outs
# +
## MAIN
# +
### tests
# +
## example 0-a from text
ins = "9, 8, 7, 6, 5"
ins = map_e(int, ins.split(', '))
ptrn = "1, 2, 3"
ptrn = map_e(int, ptrn.split(', '))
res = phase_transform(ins, ptrn)
log.info(f"result={res}")
#result = solve(ins)
#expected = 31
#aoc.assert_msg(f"input={ins} expects output={expected}, got {result}", expected == result)
# +
## example 1
ins = "12345678"
ptrn = [0, 1, 0, -1]
res = phase_transform(ins, ptrn)
log.info(f"result iter#1={res}")
res = phase_transform(res, ptrn)
log.info(f"result iter#1={res}")
result = phase_transform(ins, ptrn, num_phases=4)
expected = "01029498"
aoc.assert_msg(f"input={ins} expects output={expected}, got {result}", expected == result)
# -
# example 2
ins = "80871224585914546619083218645595"
ptrn = [0, 1, 0, -1]
result = phase_transform(ins, ptrn, num_phases=100)
result = result[:8]
expected = "24176176"
aoc.assert_msg(f"input={ins} expects output={expected}, got {result}", expected == result)
# example 3
ins = "19617804207202209144916044189917"
ptrn = [0, 1, 0, -1]
result = phase_transform(ins, ptrn, num_phases=100)
result = result[:8]
expected = "73745418"
aoc.assert_msg(f"input={ins} expects output={expected}, got {result}", expected == result)
# example 4
ins = "69317163492948606335995924319873"
ptrn = [0, 1, 0, -1]
result = phase_transform(ins, ptrn, num_phases=100)
result = result[:8]
expected = "52432133"
aoc.assert_msg(f"input={ins} expects output={expected}, got {result}", expected == result)
# +
### personal input solution
# +
log.setLevel(logging.INFO)
data = aoc.read_file_to_str("day16.in").strip()
ptrn = [0, 1, 0, -1]
log.info(f"data-len={len(data)}, data={data[:40]}...")
tm_start = timer()
result = phase_transform(data, ptrn, num_phases=100)
tm_end = timer()
result = result[:8]
print(f"result={result} needed tm={tm_end-tm_start}")
| day16/day16a.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline
#
# # Explore labels
#
#
# In this example, we load in a single subject example, load a model, and predict activity at all
# model locations. We then plot locations, which are colored labels 'observed' and 'reconstructed'.
#
#
#
# +
# Code source: <NAME> & <NAME>
# License: MIT
import supereeg as se
# load example data
bo = se.load('example_data')
# plot original locations
bo.plot_locs()
# load example model
model = se.load('example_model')
# the default will replace the electrode location with the nearest voxel and reconstruct at all other locations
reconstructed_bo = model.predict(bo, nearest_neighbor=False)
# plot the all reconstructed locations
reconstructed_bo.plot_locs()
| docs/auto_examples/plot_labels.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/VintageGold/Outages_Prediction/blob/master/WeatherStation_Criteria_RawAllNOAA.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="VUtZxdUqAQir" colab_type="text"
# #This notebook defines the criteria for all NOAA weather stations
# + [markdown] id="pXwsfFw6AZNk" colab_type="text"
# ## Imports
# + id="6mTyqgHpjCeX" colab_type="code" colab={}
import os
from os import listdir
import pandas as pd
import seaborn as sns
# + id="sdNh7K0VjJus" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 86} outputId="663b85c9-aa04-404a-f399-7dd8547fee66"
from google.colab import drive
drive.mount('/content/drive')
# %cd drive/My\ Drive/
os. getcwd()
# + [markdown] id="e2U3juLLAbf1" colab_type="text"
# ## Load Data
# + [markdown] id="RqV3dLMMBAoI" colab_type="text"
# ### Validation of old data source to new. Old data source had null values filled in as 0
# + id="3RDCd5V6jKOe" colab_type="code" colab={}
df_NOAA = pd.read_csv('UMBC_Energy/Use/All_NOAA.csv',sep='|')
# + id="9e4LzbkcjUlq" colab_type="code" colab={}
df_NOAA
df_NOAA['state_station_dt'] = df_NOAA['state'] + df_NOAA['station'] + df_NOAA['date']
# + id="EaNs72z7jwcr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 589} outputId="b13fdf73-343e-4887-d7c0-a31d7b888bf7"
df_NOAA
# + id="6W_y-58AjV75" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 346} outputId="3110c7f6-95ec-48cb-9af8-5d853f911cb7"
df_NOAA.nunique()
# + [markdown] id="SqJne4VyBMRS" colab_type="text"
# ## New Data Source for Weather Stations
# + id="OtE5ADo8jbnd" colab_type="code" colab={}
df_NOAA_Raw_2020 = pd.read_csv('UMBC_Energy/Raw/Raw_2020_NOAA.csv',sep='|')
df_NOAA_Raw_2019 = pd.read_csv('UMBC_Energy/Raw/Raw_2019_NOAA.csv',sep='|')
df_NOAA_Raw_2018 = pd.read_csv('UMBC_Energy/Raw/Raw_2018_NOAA.csv',sep='|')
df_NOAA_Raw_2017 = pd.read_csv('UMBC_Energy/Raw/Raw_2017_NOAA.csv',sep='|')
df_NOAA_Raw_2016 = pd.read_csv('UMBC_Energy/Raw/Raw_2016_NOAA.csv',sep='|')
df_NOAA_Raw_2015 = pd.read_csv('UMBC_Energy/Raw/Raw_2015_NOAA.csv',sep='|')
# + id="_obkbp0Cj7lc" colab_type="code" colab={}
df_NOAA_Raw = pd.concat([df_NOAA_Raw_2020,df_NOAA_Raw_2019,df_NOAA_Raw_2018,df_NOAA_Raw_2017,df_NOAA_Raw_2016,df_NOAA_Raw_2015])
df_NOAA_Raw = df_NOAA_Raw.drop_duplicates('station_dt_key')
# + id="Y9yOYx7ZsSvJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 190} outputId="31d33c2f-898a-4465-d42b-e33826a700a5"
df_NOAA_Raw.nunique()
# + [markdown] id="R4AK2SmvBR4Y" colab_type="text"
# ## Load dataset that has the state of each weather station
# + id="HgkhA-swrMHy" colab_type="code" colab={}
df_station_details = pd.read_csv('UMBC_Energy/Raw/station_details.csv',sep='|')
# + id="iJppurIjwwf-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 416} outputId="c479ea74-e404-459f-aadd-0e2cd8690970"
df_station_details
# + id="gxu9GSalwAlN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 242} outputId="b542a9fb-a351-4b77-afcf-e58a6ea04eea"
df_station_details.nunique()
# + [markdown] id="ysxYygUrBtoQ" colab_type="text"
# ##### Some ID's appeared in multiple states so had to keep one of them. There wasn't a value gain or loss by dropping duplicates after the first id was found
# + id="je7h6NrmxWja" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="094ac0ad-236b-4c29-b2cc-309f0c1a45fb"
ids = df_station_details["id"]
df_station_details[ids.isin(ids[ids.duplicated()])].sort_values("id").reset_index()
# + id="620DE2KkynvK" colab_type="code" colab={}
df_station_details = df_station_details.drop_duplicates('id')
# + id="6_o1SYiMytwX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 416} outputId="39d62e46-3092-4b93-ea7f-784ab408db40"
df_station_details
# + [markdown] id="qHpd9VLRCW50" colab_type="text"
# ### Merge to pair station with State
# + id="0HBIHJS6wBfK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 644} outputId="2297ce5f-0e7a-4c6a-f3cf-7462b58a0e7c"
df_NOAA_Raw.merge(df_station_details,left_on='station',right_on='id',how='left').sort_values('date')
# + id="mcRlbs5WwOFb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 644} outputId="720fa5f9-7baf-4f67-d98d-1e20a6445fd4"
df_NOAA_Raw.merge(df_station_details,left_on='station',right_on='id',how='left').sort_values('date').reset_index()
# + id="q6NeSSNJ14RL" colab_type="code" colab={}
df_NOAA_Raw_merge = df_NOAA_Raw.merge(df_station_details,left_on='station',right_on='id',how='left').sort_values('date').reset_index().drop(columns=['index'])
# + [markdown] id="CGX4sYNlH_SL" colab_type="text"
# ## **Load All State and Abbreviations**
# + id="um6Ruh84Uz7m" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 202} outputId="f935f50c-6142-4ac9-dd0b-3a2922e52b2c"
df_state_abbv = pd.read_csv('UMBC_Energy/Universal_Data/state_abb.csv',sep=',')
df_state_abbv = df_state_abbv.drop(columns='Abbrev')
df_state_abbv.head()
# + [markdown] id="1enPBjTZCjQu" colab_type="text"
# ####Merge two datasets to get state abbreviations
# + id="xSYfuwd1VGQo" colab_type="code" colab={}
df_NOAA_merge_state = df_NOAA_Raw_merge.merge(df_state_abbv,left_on = 'state', right_on = 'State',how='left')
# + id="08oLhUIXVnN3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 644} outputId="d1072285-e77b-41e8-88a6-26175fa375cc"
df_NOAA_merge_state
# + id="jN5-rULF2DA6" colab_type="code" colab={}
#df_NOAA_merge_state.to_csv('UMBC_Energy/Raw/Raw_All_NOAA.csv',sep='|',index=False)
# + id="O16outIITgFa" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 325} outputId="e31b6dd5-7826-449c-d7c1-76097cfec8aa"
df_NOAA_merge_state.groupby('Code').count()
# + [markdown] id="OqkoDZSOQ_5X" colab_type="text"
# # ** Below: Selection Criteria = Sort on amount of occurences. Search for stations with all features filled with values similar to top occurence stations. More features with a value that is similar to the station who appear most frequent is preferred and selected for analysis**
# + [markdown] id="dzZNZ3EiQgUP" colab_type="text"
# ## DC
# + id="ngLZdKugIW9Y" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 233} outputId="846cdc21-6a41-49aa-ac90-ae46a1ef47a9"
#Selected Top 2 selected
df_NOAA_state_group = df_NOAA_merge_state[df_NOAA_merge_state.datacoverage >= .99]
df_NOAA_state_group.query('Code == "DC"').groupby('id').count().sort_values('station',ascending=False).head(6)
# + [markdown] id="rhgbzP8fQjDi" colab_type="text"
# ## DE
# + id="sEejwT9wL81d" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 356} outputId="9b0e3726-4ed5-4678-ea48-1c6b6c5ffe48"
#Selected First 2 and Tail id
df_NOAA_state_group = df_NOAA_merge_state[df_NOAA_merge_state.datacoverage >= .99]
df_NOAA_state_group.query('Code == "DE"').groupby('id').count().sort_values('station',ascending=False).head(9)
# + [markdown] id="1MfI44tUQnqZ" colab_type="text"
# ## MD
# + id="iH7C2aR_OUYr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 264} outputId="7c5ec641-477b-4ba9-e528-bafddcd1e9e0"
#Selected Top 5
df_NOAA_state_group = df_NOAA_merge_state[df_NOAA_merge_state.datacoverage >= .99]
df_NOAA_state_group.query('Code == "MD"').groupby('id').count().sort_values('station',ascending=False).head(6)
# + [markdown] id="X7qpqgZ0Qp3c" colab_type="text"
# ## NJ
# + id="yGjG1kY5PAoT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 264} outputId="d0442734-6aae-47cc-a1ea-b242c1a014e0"
#Selected Top 5
df_NOAA_state_group = df_NOAA_merge_state[df_NOAA_merge_state.datacoverage >= .99]
df_NOAA_state_group.query('Code == "NJ"').groupby('id').count().sort_values('station',ascending=False).head(6)
# + [markdown] id="Jkg5_UMhQsEP" colab_type="text"
# ## OH
# + id="FdKFrpisPEel" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 264} outputId="af7d38b2-30c1-46f7-9115-474ea735b2d4"
#Selected Top 5
df_NOAA_state_group = df_NOAA_merge_state[df_NOAA_merge_state.datacoverage >= .99]
df_NOAA_state_group.query('Code == "OH"').groupby('id').count().sort_values('station',ascending=False).head(6)
# + [markdown] id="KzXRKn_pQu1t" colab_type="text"
# ## PA
# + id="JBDtoCTgPFdM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 264} outputId="e586d255-633f-465a-b9bc-b95e1d0b4c73"
# Selected
df_NOAA_state_group = df_NOAA_merge_state[df_NOAA_merge_state.datacoverage >= .99]
df_NOAA_state_group.query('Code == "PA"').groupby('id').count().sort_values('station',ascending=False).head(6)
# + [markdown] id="_aNWXG11QxPV" colab_type="text"
# ## VA
# + id="bVnsM2wjPHTU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 264} outputId="b0fc0b8d-52de-4aee-b673-4a3803d28d75"
df_NOAA_state_group = df_NOAA_merge_state[df_NOAA_merge_state.datacoverage >= .99]
df_NOAA_state_group.query('Code == "VA"').groupby('id').count().sort_values('station',ascending=False).head(6)
# + [markdown] id="lGy3PEj2Qzo1" colab_type="text"
# ##WV
# + id="8HzHaHCEPIX3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 264} outputId="2291d581-08fa-4ac8-c39e-6cd1ec59d4e7"
df_NOAA_state_group = df_NOAA_merge_state[df_NOAA_merge_state.datacoverage >= .99]
df_NOAA_state_group.query('Code == "WV"').groupby('id').count().sort_values('station',ascending=False).head(6)
# + id="73wYgNbDPJhL" colab_type="code" colab={}
| WeatherStation_Criteria_RawAllNOAA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" deletable=true editable=true id="xu2SVpFJjmJr"
# # DeepDreaming with TensorFlow
# + [markdown] colab_type="toc" deletable=true editable=true id="hupz2hrZjdnC"
# >[Loading and displaying the model graph](#loading)
#
# >[Classifying Images](#classify)
#
# >[Naive feature visualization](#naive)
#
# >[Multiscale image generation](#multiscale)
#
# >[Laplacian Pyramid Gradient Normalization](#laplacian)
#
# >[Playing with feature visualzations](#playing)
#
# >[DeepDream](#deepdream)
#
# >[More Fun!](#fun)
#
# + [markdown] colab_type="text" deletable=true editable=true id="-PLC9SvcQgkG"
# # This tutorial will show how to work with TensorFlow, and how to use image classification models.
# You will benefit most from this if you have some working knowledge of python, and if you have a rough idea what a neural network is and how it works.
#
# You will learn about deep neural networks, in particular convolutional neural networks, and how they are used for image classification tasks. We will also gain
# an intuitive understanding how neural network represent information they have learned.
#
# Specifically, you will learn how to:
#
# - Load and a pre-trained TensorFlow model and inspect it.
# - Classify images using TensorFlow using a pre-trained model.
# - Visualize feature channels from a convolutional network to understand what it has learned.
# - Enhance what the network sees in a given image to produce dream-like images.
#
# The network we'll examine is [Inception-v3](http://arxiv.org/abs/1512.00567). It's trained to classify an image into 1 of the 1000 categories from the [ImageNet](http://image-net.org/) dataset. For a good introduction to neural networks, see the book [Neural Networks and Deep Learning](http://neuralnetworksanddeeplearning.com/) by <NAME>. For background on convolutional networks, see <NAME>'s excellent [blog post](http://colah.github.io/posts/2014-07-Conv-Nets-Modular/).
#
# As discussed in [Inceptionism: Going Deeper into Neural Networks](http://googleresearch.blogspot.com/2015/06/inceptionism-going-deeper-into-neural.html), our goal is to visualize the internal image representations learned by a network trained to classify images. We'll make these visualizations both efficient to generate, and even beautiful.
#
# Impatient readers can start with exploring the full galleries of images generated by the method described here for [GoogLeNet](http://storage.googleapis.com/deepdream/visualz/tensorflow_inception/index.html) and [VGG16](http://storage.googleapis.com/deepdream/visualz/vgg16/index.html) architectures.
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": []} colab_type="code" deletable=true editable=true executionInfo={"elapsed": 371, "status": "ok", "timestamp": 1457963606294, "user": {"color": "#1FA15D", "displayName": "<NAME>", "isAnonymous": false, "isMe": true, "permissionId": "12341152118244997759", "photoUrl": "https://lh3.googleusercontent.com/-XdUIqdMkCWA/AAAAAAAAAAI/AAAAAAAAAAA/4252rscbv5M/s128/photo.jpg", "sessionId": "1269ead540f76ce5", "userId": "108092561333339272254"}, "user_tz": -60} id="jtD9nb-2QgkY" outputId="b935629b-8608-45c1-942f-612b7dbb13d3"
# boilerplate code
import os
import re
from io import StringIO
import numpy as np
from functools import partial
import PIL.Image
import PIL.ImageOps
from IPython.display import clear_output, Image, display, HTML
import codecs
import tensorflow as tf
print("Done")
# + [markdown] colab_type="text" deletable=true editable=true id="ILvNKvMvc2n5"
# <a id='loading'></a>
# ## Loading and displaying the model graph
#
# The pretrained network can be downloaded [here](http://download.tensorflow.org/models/inception5h.zip). If it is not already here, download it and unpack the archive. The actual network is stored in the file `tensorflow_inception_graph.pb`. Set the `model_fn` variable to its path.
# + [markdown] deletable=true editable=true
# We will now load the network and prepare it for input. TensorFlow maintains a computation graph and a session, which maintains state for running computations and which can be executed remotely. We will first make a fresh graph and a session which uses that graph. The session will be used in the rest of the tutorial.
#
# We then load the model. The model consists of a computation graph which happens to have a node called "input", into which we need to feed a batch of input images. The input node in the graph expects images that are normalized by subtracting the average brightness of all images in the imagenet dataset.
#
# Because we will use single, unnormalized images, we will make a small graph that takes an image, subtracts the imagenet mean, and expands it to look like a batch of images.
#
# We then load the graph from file and import it into the default graph for our session. The little importer graph is now connected to the input node in the loaded graph, and we can feed regular images into the graph.
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": []} colab_type="code" deletable=true editable=true executionInfo={"elapsed": 2264, "status": "ok", "timestamp": 1457962713799, "user": {"color": "#1FA15D", "displayName": "<NAME>", "isAnonymous": false, "isMe": true, "permissionId": "12341152118244997759", "photoUrl": "https://lh3.googleusercontent.com/-XdUIqdMkCWA/AAAAAAAAAAI/AAAAAAAAAAA/4252rscbv5M/s128/photo.jpg", "sessionId": "761b412462cda2d0", "userId": "108092561333339272254"}, "user_tz": -60} id="1kJuJRLiQgkg" outputId="d2aaf8cc-91e1-4864-8cf8-0aef612db1d6"
# creating fresh Graph and TensorFlow session
graph = tf.Graph()
sess = tf.InteractiveSession(graph=graph)
# Prepare input for the format expected by the graph
t_input = tf.placeholder(np.float32, name='our_input') # define the input tensor
imagenet_mean = 117.0
t_preprocessed = tf.expand_dims(t_input-imagenet_mean, 0)
# Load graph and import into graph used by our session
model_fn = 'tensorflow_inception_graph.pb'
with open(model_fn, "rb") as datafile:
graph_def = tf.GraphDef.FromString(datafile.read())
tf.import_graph_def(graph_def, {'input':t_preprocessed})
sw = tf.train.SummaryWriter('./logs3', graph=tf.get_default_graph())
print("imported")
# + [markdown] deletable=true editable=true
# Let's first count how many layers there are in this graph (we'll only count the convolutional layers), and how many total features this graph uses internally. We'll look at what those features look like later, we have enough to choose from.
# + deletable=true editable=true
layers = [op.name for op in graph.get_operations() if op.type=='Conv2D' and 'import/' in op.name]
feature_nums = [int(graph.get_tensor_by_name(name+':0').get_shape()[-1]) for name in layers]
print 'Number of layers', len(layers)
print 'Total number of feature channels:', sum(feature_nums)
# + [markdown] deletable=true editable=true
# Now we'll look at what the graph looks like. We use tensorboard to visualize the graph, first stripping large constants (containing the pre-trained network weights) to speed things up. We can use the names shown in the diagram to identify layers we'd like to look into. Be sure to expand the "mixed" node, which contains the bulk of the graph.
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}, {"item_id": 2}]} colab_type="code" deletable=true editable=true executionInfo={"elapsed": 1198, "status": "ok", "timestamp": 1457962715078, "user": {"color": "#1FA15D", "displayName": "<NAME>", "isAnonymous": false, "isMe": true, "permissionId": "12341152118244997759", "photoUrl": "https://lh3.googleusercontent.com/-XdUIqdMkCWA/AAAAAAAAAAI/AAAAAAAAAAA/4252rscbv5M/s128/photo.jpg", "sessionId": "761b412462cda2d0", "userId": "108092561333339272254"}, "user_tz": -60} id="LrucdvgyQgks" outputId="5936270b-5da8-4825-b2e9-145c494d36e6"
# Helper functions for TF Graph visualization
def strip_consts(graph_def, max_const_size=32):
"""Strip large constant values from graph_def."""
strip_def = tf.GraphDef()
for n0 in graph_def.node:
n = strip_def.node.add()
n.MergeFrom(n0)
if n.op == 'Const':
tensor = n.attr['value'].tensor
size = len(tensor.tensor_content)
if size > max_const_size:
tensor.tensor_content = "<stripped %d bytes>"%size
return strip_def
def rename_nodes(graph_def, rename_func):
res_def = tf.GraphDef()
for n0 in graph_def.node:
n = res_def.node.add()
n.MergeFrom(n0)
n.name = rename_func(n.name)
for i, s in enumerate(n.input):
n.input[i] = rename_func(s) if s[0]!='^' else '^'+rename_func(s[1:])
return res_def
def show_graph(graph_def, max_const_size=32):
"""Visualize TensorFlow graph."""
if hasattr(graph_def, 'as_graph_def'):
graph_def = graph_def.as_graph_def()
strip_def = strip_consts(graph_def, max_const_size=max_const_size)
code = """
<script>
function load() {{
document.getElementById("{id}").pbtxt = {data};
}}
</script>
<link rel="import" href="https://tensorboard.appspot.com/tf-graph-basic.build.html" onload=load()>
<div style="height:600px">
<tf-graph-basic id="{id}"></tf-graph-basic>
</div>
""".format(data=repr(str(strip_def)), id='graph'+str(np.random.rand()))
iframe = """
<iframe seamless style="width:800px;height:620px;border:0" srcdoc="{}"></iframe>
""".format(code.replace('"', '"'))
display(HTML(iframe))
# Visualizing the network graph. Be sure expand the "mixed" nodes to see their
# internal structure. We are going to visualize "Conv2D" nodes.
tmp_def = rename_nodes(graph_def, lambda s:"/".join(s.split('_',1)))
show_graph(tmp_def)
# + [markdown] colab_type="text" deletable=true editable=true id="eJZVMSmiQgkp"
# To take a glimpse into the kinds of patterns that the network learned to recognize, we will try to generate images that maximize the sum of activations of particular channel of a particular convolutional layer of the neural network. The network we explore contains many convolutional layers, each of which outputs tens to hundreds of feature channels, so we have plenty of patterns to explore.
# + [markdown] deletable=true editable=true
# <a id='classify'></a>
# ## Classifying Images
#
# Let's first classify some images using this graph. The softmax layer contains the network's predictions in form of numerical IDs. To translate those to human-readable values, we have to load some translation files.
# + deletable=true editable=true
label_lookup_path = 'imagenet_2012_challenge_label_map_proto.pbtxt'
uid_lookup_path = 'imagenet_synset_to_human_label_map.txt'
# The id translation goes via id strings -- find translation between UID string and human-friendly names
proto_as_ascii_lines = open(uid_lookup_path).readlines()
uid_to_human = {}
p = re.compile(r'[n\d]*[ \S,]*')
for line in proto_as_ascii_lines:
parsed_items = p.findall(line)
uid = parsed_items[0]
human_string = parsed_items[2]
uid_to_human[uid] = human_string
# Get node IDs to UID strings map
proto_as_ascii_lines = open(label_lookup_path).readlines()
node_id_to_uid = {}
for line in proto_as_ascii_lines:
if line.startswith(' target_class:'):
target_class = int(line.split(': ')[1])
if line.startswith(' target_class_string:'):
target_class_string = line.split(': ')[1]
node_id_to_uid[target_class] = target_class_string[1:-2]
# Make node ID to human friendly names map
node_id_to_name = {}
for key, val in node_id_to_uid.iteritems():
name = uid_to_human[val]
node_id_to_name[key] = name
# make sure we have a name for each possible ID
for i in range(graph.get_tensor_by_name('import/softmax2:0').get_shape()[1]):
if i not in node_id_to_name:
node_id_to_name[i] = '???'
# + [markdown] deletable=true editable=true
# Now, you can find out what any neuron stands for. Imagenet has 1000 different classes, some of them pretty obscure.
# + deletable=true editable=true
node_id_to_name[438]
# + [markdown] deletable=true editable=true
# The predictions of the network are contained in the output of the softmax layer. In the network we loaded, the relevant layer is called "softmax2". We'll make a small function which feeds an input image, reads the result from the graph and translates it, plus an additional function that will create an image from an array.
# + deletable=true editable=true
# Helper function to get a named layer from the graph
def T(layer_name):
return graph.get_tensor_by_name("import/%s:0" % layer_name)
softmax = T('softmax2')
def prep_img(filename):
size = (224, 224)
img = PIL.Image.open(filename)
img.thumbnail(size, PIL.Image.ANTIALIAS)
thumb = PIL.ImageOps.fit(img, size, PIL.Image.ANTIALIAS, (0.5, 0.5))
return np.float32(thumb)
# Print the 5 top predictions for a given image
def prediction(filename, k=5):
img = prep_img(filename)
# Load, resize, and central square crop the image.
# Compute predictions
predictions = sess.run(softmax, {t_input: img})
predictions = np.squeeze(predictions)
top_k = predictions.argsort()[-k:][::-1]
for node_id in top_k:
human_string = node_id_to_name[node_id]
score = predictions[node_id]
print '%s (score = %.5f)' % (human_string, score)
# Helper function: Display an image
def showarray(a, fmt='jpeg', size=None):
a = np.uint8(np.clip(a, 0, 1)*255)
f = StringIO()
img = PIL.Image.fromarray(a)
if size is not None:
img = img.resize((size,size))
img.save(f, fmt)
display(Image(data=f.getvalue()))
print "defined"
# + [markdown] deletable=true editable=true
# Now we can classify a panda. You should also try other images.
# + deletable=true editable=true
showarray(prep_img('testimages/deer.jpg')/255., size=500)
prediction('testimages/deer.jpg')
# + [markdown] colab_type="text" deletable=true editable=true id="Nv2JqNLBhy1j"
# <a id='naive'></a>
# ## Naive feature visualization
# + [markdown] colab_type="text" deletable=true editable=true id="6LXaGEJkQgk4"
# This tells us what the network thinks is contained in an image, but we still don't know why. So let's start to look at the features the network has learned to recognize.
#
# First, we'll define some helper functions to show images, and an input image with a bit of random noise, which is always useful.
# + deletable=true editable=true
# Normalize an image for visualization
def visstd(a, s=0.1):
return (a-a.mean())/max(a.std(), 1e-4)*s + 0.5
img_noise = np.random.uniform(size=(224,224,3)) + 100.0
# + [markdown] deletable=true editable=true
# The first layer operates directly on the image. It contains a weight tensor taking a 7x7 patch of the image and computing 64 features from it. We can look at the learned weights in this layer to see what kinds of features these are.
# + deletable=true editable=true
# Uncomment the following line to see all the possible weights
print '\n'.join([op.name[7:] for op in graph.get_operations() if op.name.endswith('w')])
w = T('conv2d0_w')
shape = w.get_shape().as_list()
print shape
feature_id = 14
weights = tf.squeeze(tf.slice(w, [0, 0, 0, feature_id], [-1, -1, -1, 1])).eval()
showarray(visstd(weights), size=200)
# + [markdown] deletable=true editable=true
# If you're trained to look at filter kernels, this may be useful to you. You can also look at what the output of the convolution looks like, by running the first layer on an image.
# + deletable=true editable=true
a = T('conv2d0')
feature = tf.squeeze(tf.slice(a, [0, 0, 0, feature_id], [-1, -1, -1, 1]))
img = prep_img('testimages/deer.jpg')
feature = sess.run(feature, {t_input: img})
showarray(visstd(feature), size=500)
# + [markdown] deletable=true editable=true
# Even in the first layer, the weights are not easy to interpret. For all layers except the first, it is almost impossible. We have to find another way of understanding what the network has learned.
#
# We use a simple visualization technique to show what any given feature looks like: Image space gradient ascent. This works as follows: We pick a feature plane from any layer in the network. This feature plane recognizes the presence of a specific feature in the image. We will try to generate an image that maximizes this feature signal. We start with an image that is just noise, and compute the gradient of the feature signal (averaged over the whole image) with respect to the input image. We then modify the input image to increase the feature signal. This generates an image that this specific network layer thinks is full of whatever feature it is meant to detect.
# + deletable=true editable=true
def render_naive(t_obj, img0=img_noise, iter_n=20, step=1.0):
t_score = tf.reduce_mean(t_obj) # defining the optimization objective
t_grad = tf.gradients(t_score, t_input)[0] # behold the power of automatic differentiation!
img = img0.copy()
for i in xrange(iter_n):
g, score = sess.run([t_grad, t_score], {t_input:img})
# normalizing the gradient, so the same step size should work
g /= g.std()+1e-8 # for different layers and networks
img += g*step
print score,
clear_output()
showarray(visstd(img), size=448)
# + [markdown] deletable=true editable=true
# Now we have to pick a layer and a feature channel to visualize. The following cell will enumerate all the available layers. Refer to the graph above to see where they are.
#
# Earlier layers (closer to the bottom, i.e. the input) have lower level features, later layers have higher level features. Note that we use layer outputs before applying the ReLU nonlinearity in order to have non-zero gradients for features with negative initial activations (hence the "pre_relu").
# + deletable=true editable=true
# Run this cell if you'd like a list of all layers to pick from
print '\n'.join([op.name[7:] for op in graph.get_operations() if op.name.endswith('pre_relu')])
# + [markdown] deletable=true editable=true
# ## Rendering
#
# At this point, we can render one of these layers.
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}, {"item_id": 2}]} colab_type="code" deletable=true editable=true executionInfo={"elapsed": 3, "status": "ok", "timestamp": 1457962479327, "user": {"color": "#1FA15D", "displayName": "<NAME>", "isAnonymous": false, "isMe": true, "permissionId": "12341152118244997759", "photoUrl": "https://lh3.googleusercontent.com/-XdUIqdMkCWA/AAAAAAAAAAI/AAAAAAAAAAA/4252rscbv5M/s128/photo.jpg", "sessionId": "761b412462cda2d0", "userId": "108092561333339272254"}, "user_tz": -60} id="ZxC_XGGXQgk7" outputId="1c971a74-bf65-4069-cfd0-1473aa909a83"
# Pick any internal layer.
layer = 'mixed4d_3x3_bottleneck_pre_relu'
print '%d channels in layer.' % T(layer).get_shape()[-1]
# Pick a feature channel to visualize
channel = 139
render_naive(T(layer)[:,:,:,channel])
# + [markdown] colab_type="text" deletable=true editable=true id="ZroBKE5YiDsb"
# <a id="multiscale"></a>
# ## Multiscale image generation
#
# Looks like the network wants to show us something interesting! Let's help it. We are going to apply gradient ascent on multiple scales. Details formed on smaller scale will be upscaled and augmented with additional details on the next scale.
#
# Basically, instead of starting from a random noise image, we start only the first iteration (octave) from random noise, and each octave after we start from the upsampled result of the previous optimization on the smaller image.
#
# With multiscale image generation it may be tempting to set the number of octaves to some high value to produce wallpaper-sized images. Storing network activations and backprop values will quickly run out of GPU memory in this case. There is a simple trick to avoid this: split the image into smaller tiles and compute each tile gradient independently. Applying random shifts to the image before every iteration helps avoid tile seams and improves the overall image quality.
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}]} colab_type="code" deletable=true editable=true executionInfo={"elapsed": 464, "status": "ok", "timestamp": 1457963844162, "user": {"color": "#1FA15D", "displayName": "<NAME>", "isAnonymous": false, "isMe": true, "permissionId": "12341152118244997759", "photoUrl": "https://lh3.googleusercontent.com/-XdUIqdMkCWA/AAAAAAAAAAI/AAAAAAAAAAA/4252rscbv5M/s128/photo.jpg", "sessionId": "1269ead540f76ce5", "userId": "108092561333339272254"}, "user_tz": -60} id="2iwWSOgsQglG" outputId="221dae81-914b-4167-eb49-26ef2d431a66"
def tffunc(*argtypes):
'''Helper that transforms TF-graph generating function into a regular one.
See "resize" function below.
'''
placeholders = map(tf.placeholder, argtypes)
def wrap(f):
out = f(*placeholders)
def wrapper(*args, **kw):
return out.eval(dict(zip(placeholders, args)), session=kw.get('session'))
return wrapper
return wrap
# Helper function that uses TF to resize an image
def resize(img, size):
img = tf.expand_dims(img, 0)
return tf.image.resize_bilinear(img, size)[0,:,:,:]
resize = tffunc(np.float32, np.int32)(resize)
def calc_grad_tiled(img, t_grad, tile_size=512):
'''Compute the value of tensor t_grad over the image in a tiled way.
Random shifts are applied to the image to blur tile boundaries over
multiple iterations.'''
sz = tile_size
h, w = img.shape[:2]
sx, sy = np.random.randint(sz, size=2)
img_shift = np.roll(np.roll(img, sx, 1), sy, 0)
grad = np.zeros_like(img)
for y in xrange(0, max(h-sz//2, sz),sz):
for x in xrange(0, max(w-sz//2, sz),sz):
sub = img_shift[y:y+sz,x:x+sz]
g = sess.run(t_grad, {t_input:sub})
grad[y:y+sz,x:x+sz] = g
return np.roll(np.roll(grad, -sx, 1), -sy, 0)
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}, {"item_id": 2}]} colab_type="code" deletable=true editable=true executionInfo={"elapsed": 127, "status": "ok", "timestamp": 1457963487829, "user": {"color": "#1FA15D", "displayName": "<NAME>", "isAnonymous": false, "isMe": true, "permissionId": "12341152118244997759", "photoUrl": "https://lh3.googleusercontent.com/-XdUIqdMkCWA/AAAAAAAAAAI/AAAAAAAAAAA/4252rscbv5M/s128/photo.jpg", "sessionId": "1269ead540f76ce5", "userId": "108092561333339272254"}, "user_tz": -60} id="GRCJdG8gQglN" outputId="7e21352d-9131-4f81-a52f-912b2e299475"
def render_multiscale(t_obj, img0=img_noise, iter_n=10, step=1.0, octave_n=2, octave_scale=1.4):
t_score = tf.reduce_mean(t_obj) # defining the optimization objective
t_grad = tf.gradients(t_score, t_input)[0] # behold the power of automatic differentiation!
img = img0.copy()
for octave in xrange(octave_n):
if octave > 0:
hw = np.float32(img.shape[:2])*octave_scale
img = resize(img, np.int32(hw))
for i in xrange(iter_n):
g = calc_grad_tiled(img, t_grad)
# normalizing the gradient, so the same step size should work
g /= g.std()+1e-8 # for different layers and networks
img += g*step
print '.',
clear_output()
showarray(visstd(img))
print 'Octave %d' % octave
render_multiscale(T(layer)[:,:,:,channel])
# + [markdown] colab_type="text" deletable=true editable=true id="mDSZMtVYQglV"
# <a id="laplacian"></a>
# ## Laplacian Pyramid Gradient Normalization
#
# This looks better, but the resulting images mostly contain high frequencies. Can we improve it? One way is to add a smoothness prior into the optimization objective. This will effectively blur the image a little every iteration, suppressing the higher frequencies, so that the lower frequencies can catch up. This will require more iterations to produce a nice image. Why don't we just boost lower frequencies of the gradient instead? One way to achieve this is through the Laplacian pyramid decomposition. We call the resulting technique _Laplacian Pyramid Gradient Normalization_.
#
# We therefore split the image into its various frequency bands by repeatedly blurring it and extracting the highest frequencies by subtracting the blurred image (`lap_split` and `lap_split_n` below). We then normalize each frequency band separately (`normalize_std`), and then merge them back together by basically just adding them up (`lap_merge`).
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}]} colab_type="code" deletable=true editable=true executionInfo={"elapsed": 512, "status": "ok", "timestamp": 1457963876373, "user": {"color": "#1FA15D", "displayName": "<NAME>", "isAnonymous": false, "isMe": true, "permissionId": "12341152118244997759", "photoUrl": "https://lh3.googleusercontent.com/-XdUIqdMkCWA/AAAAAAAAAAI/AAAAAAAAAAA/4252rscbv5M/s128/photo.jpg", "sessionId": "1269ead540f76ce5", "userId": "108092561333339272254"}, "user_tz": -60} id="Do3WpFSUQglX" outputId="99835b80-ed6f-47a5-85c3-c77bd55d7b17"
k = np.float32([1,4,6,4,1])
k = np.outer(k, k)
k5x5 = k[:,:,None,None]/k.sum()*np.eye(3, dtype=np.float32)
def lap_split(img):
'''Split the image into lo and hi frequency components'''
with tf.name_scope('split'):
lo = tf.nn.conv2d(img, k5x5, [1,2,2,1], 'SAME') # Blurred image -- low frequencies only
lo2 = tf.nn.conv2d_transpose(lo, k5x5*4, tf.shape(img), [1,2,2,1])
hi = img-lo2 # hi is img with low frequencies removed
return lo, hi
def lap_split_n(img, n):
'''Build Laplacian pyramid with n splits'''
levels = []
for i in xrange(n):
img, hi = lap_split(img)
levels.append(hi)
levels.append(img)
return levels[::-1] # List of images with lower and lower frequencies
def lap_merge(levels):
'''Merge Laplacian pyramid'''
img = levels[0]
for hi in levels[1:]:
with tf.name_scope('merge'):
img = tf.nn.conv2d_transpose(img, k5x5*4, tf.shape(hi), [1,2,2,1]) + hi
return img # Reconstructed image, all frequencies added back together
def normalize_std(img, eps=1e-10):
'''Normalize image by making its standard deviation = 1.0'''
with tf.name_scope('normalize'):
std = tf.sqrt(tf.reduce_mean(tf.square(img)))
return img/tf.maximum(std, eps)
def lap_normalize(img, scale_n=4):
'''Perform the Laplacian pyramid normalization.'''
img = tf.expand_dims(img,0)
tlevels = lap_split_n(img, scale_n) # Split into frequencies
tlevels = map(normalize_std, tlevels) # Normalize each frequency band
out = lap_merge(tlevels) # Put image back together
return out[0,:,:,:]
# + [markdown] deletable=true editable=true
# We did all this in TensorFlow, so it generated a computation graph that we can inspect.
# + deletable=true editable=true
# Showing the lap_normalize graph with TensorBoard
lap_graph = tf.Graph()
with lap_graph.as_default():
lap_in = tf.placeholder(np.float32, name='lap_in')
lap_out = lap_normalize(lap_in)
show_graph(lap_graph)
# + [markdown] deletable=true editable=true
# We now hav everything to render another image. The algorithm is unchanged, except that in each iteration, the gradient image is normalized using the Laplacian normalization graph.
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}, {"item_id": 2}]} colab_type="code" deletable=true editable=true executionInfo={"elapsed": 17273, "status": "ok", "timestamp": 1457964054088, "user": {"color": "#1FA15D", "displayName": "<NAME>", "isAnonymous": false, "isMe": true, "permissionId": "12341152118244997759", "photoUrl": "https://lh3.googleusercontent.com/-XdUIqdMkCWA/AAAAAAAAAAI/AAAAAAAAAAA/4252rscbv5M/s128/photo.jpg", "sessionId": "1269ead540f76ce5", "userId": "108092561333339272254"}, "user_tz": -60} id="zj8Ms-WqQgla" outputId="aa54c6c3-bf38-4054-f3f4-a5c82218e251"
def render_lapnorm(t_obj, img0=img_noise, visfunc=visstd,
iter_n=10, step=1.0, octave_n=3, octave_scale=1.4, lap_n=4):
t_score = tf.reduce_mean(t_obj) # defining the optimization objective
t_grad = tf.gradients(t_score, t_input)[0] # behold the power of automatic differentiation!
# Build the Laplacian normalization graph
lap_norm_func = tffunc(np.float32)(partial(lap_normalize, scale_n=lap_n))
img = img0.copy()
for octave in xrange(octave_n):
if octave>0:
hw = np.float32(img.shape[:2])*octave_scale
img = resize(img, np.int32(hw))
for i in xrange(iter_n):
g = calc_grad_tiled(img, t_grad)
g = lap_norm_func(g) # New!
img += g*step
print '.',
clear_output()
showarray(visfunc(img))
render_lapnorm(T(layer)[:,:,:,channel])
# + [markdown] colab_type="text" deletable=true editable=true id="YzXJUF2lQgln"
# <a id="playing"></a>
# ## Playing with feature visualizations
#
# We got a nice smooth image using only 10 iterations per octave. In case of running on GPU this takes just a few seconds. Let's try to visualize another channel from the same layer. The network can generate wide diversity of patterns.
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}, {"item_id": 2}]} colab_type="code" deletable=true editable=true executionInfo={"elapsed": 35857, "status": "ok", "timestamp": 1457964089937, "user": {"color": "#1FA15D", "displayName": "<NAME>", "isAnonymous": false, "isMe": true, "permissionId": "12341152118244997759", "photoUrl": "https://lh3.googleusercontent.com/-XdUIqdMkCWA/AAAAAAAAAAI/AAAAAAAAAAA/4252rscbv5M/s128/photo.jpg", "sessionId": "1269ead540f76ce5", "userId": "108092561333339272254"}, "user_tz": -60} id="a6jfiWqZQglq" outputId="40ebf3c4-a262-4e11-fca9-d42eb9306edb"
render_lapnorm(T(layer)[:,:,:,65])
# + [markdown] colab_type="text" deletable=true editable=true id="ka6RyOMEnrB5"
# Lower layers produce features of lower complexity.
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}, {"item_id": 2}]} colab_type="code" deletable=true editable=true executionInfo={"elapsed": 25, "status": "ok", "timestamp": 1457967232229, "user": {"color": "#1FA15D", "displayName": "<NAME>", "isAnonymous": false, "isMe": true, "permissionId": "12341152118244997759", "photoUrl": "https://lh3.googleusercontent.com/-XdUIqdMkCWA/AAAAAAAAAAI/AAAAAAAAAAA/4252rscbv5M/s128/photo.jpg", "sessionId": "1269ead540f76ce5", "userId": "108092561333339272254"}, "user_tz": -60} id="KYOtrJxMnlws" outputId="8ec79dd8-259e-4bca-8115-705e76d1bc74"
render_lapnorm(T('mixed3b_1x1_pre_relu')[:,:,:,101])
# + [markdown] colab_type="text" deletable=true editable=true id="wuP8a4FlQglx"
# There are many interesting things one may try. For example, optimizing a linear combination of features gives a "mixture" pattern.
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}, {"item_id": 2}]} colab_type="code" deletable=true editable=true executionInfo={"elapsed": 14408, "status": "ok", "timestamp": 1457964104157, "user": {"color": "#1FA15D", "displayName": "<NAME>", "isAnonymous": false, "isMe": true, "permissionId": "12341152118244997759", "photoUrl": "https://lh3.googleusercontent.com/-XdUIqdMkCWA/AAAAAAAAAAI/AAAAAAAAAAA/4252rscbv5M/s128/photo.jpg", "sessionId": "1269ead540f76ce5", "userId": "108092561333339272254"}, "user_tz": -60} id="ozN-nH2yQgl0" outputId="a890305e-7bed-4011-8535-5882d6b27482"
render_lapnorm(T(layer)[:,:,:,65] + T(layer)[:,:,:,139], octave_n=4)
# + [markdown] colab_type="text" deletable=true editable=true id="lcPe-ZMv0dYR"
# <a id="deepdream"></a>
# ## DeepDream
#
# Now let's reproduce the [DeepDream algorithm](https://github.com/google/deepdream/blob/master/dream.ipynb) with TensorFlow.
#
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": []} colab_type="code" deletable=true editable=true executionInfo={"elapsed": 465, "status": "ok", "timestamp": 1457967388369, "user": {"color": "#1FA15D", "displayName": "<NAME>", "isAnonymous": false, "isMe": true, "permissionId": "12341152118244997759", "photoUrl": "https://lh3.googleusercontent.com/-XdUIqdMkCWA/AAAAAAAAAAI/AAAAAAAAAAA/4252rscbv5M/s128/photo.jpg", "sessionId": "1269ead540f76ce5", "userId": "108092561333339272254"}, "user_tz": -60} id="qM2U_96hyUwN" outputId="3725acc2-51cc-4894-e726-87bfe5727342"
def render_deepdream(t_obj, img0=img_noise,
iter_n=10, step=1.5, octave_n=4, octave_scale=1.4):
t_score = tf.reduce_mean(t_obj) # defining the optimization objective
t_grad = tf.gradients(t_score, t_input)[0] # behold the power of automatic differentiation!
# Build the Laplacian normalization graph
lap_norm_func = tffunc(np.float32)(partial(lap_normalize, scale_n=4))
# split the image into a number of octaves
img = img0
octaves = []
for i in xrange(octave_n-1):
hw = img.shape[:2]
lo = resize(img, np.int32(np.float32(hw)/octave_scale))
hi = img-resize(lo, hw)
img = lo
octaves.append(hi)
# generate details octave by octave
for octave in xrange(octave_n):
if octave>0:
hi = octaves[-octave]
img = resize(img, hi.shape[:2])+hi
for i in xrange(iter_n):
g = calc_grad_tiled(img, t_grad)
g = lap_norm_func(g)
img += g*(step / (np.abs(g).mean()+1e-7))
print '.',
clear_output()
showarray(img/255.0)
return img
# + [markdown] deletable=true editable=true
# Let's load some image and populate it with DogSlugs (in case you've missed them).
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}]} colab_type="code" deletable=true editable=true executionInfo={"elapsed": 600, "status": "ok", "timestamp": 1457967452116, "user": {"color": "#1FA15D", "displayName": "<NAME>", "isAnonymous": false, "isMe": true, "permissionId": "12341152118244997759", "photoUrl": "https://lh3.googleusercontent.com/-XdUIqdMkCWA/AAAAAAAAAAI/AAAAAAAAAAA/4252rscbv5M/s128/photo.jpg", "sessionId": "1269ead540f76ce5", "userId": "108092561333339272254"}, "user_tz": -60} id="M9_vOh_2Qgl-" outputId="eef01469-fb9b-4242-f249-e81383bf0433"
img0 = PIL.Image.open('testimages/mountains.jpg')
img0 = np.float32(img0)
showarray(img0/255.0)
# + [markdown] deletable=true editable=true
# Btw., this picture apparently contains a headland. Probably some sort of valley, alp or cliff.
# + deletable=true editable=true
prediction('testimages/mountains.jpg')
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}, {"item_id": 2}]} colab_type="code" deletable=true editable=true executionInfo={"elapsed": 79, "status": "ok", "timestamp": 1457967471615, "user": {"color": "#1FA15D", "displayName": "<NAME>", "isAnonymous": false, "isMe": true, "permissionId": "12341152118244997759", "photoUrl": "https://lh3.googleusercontent.com/-XdUIqdMkCWA/AAAAAAAAAAI/AAAAAAAAAAA/4252rscbv5M/s128/photo.jpg", "sessionId": "1269ead540f76ce5", "userId": "108092561333339272254"}, "user_tz": -60} id="k0oggbGEeC3U" outputId="c7258412-9cb1-4a94-d4f5-e6120a728c85"
_ = render_deepdream(tf.square(T('mixed4c')), img0)
# + [markdown] colab_type="text" deletable=true editable=true id="IJzvhEFxpB7E"
# Recall how the first parameter to `render_*` was the optimization objecting. In the above case, we are optimizing for `square(mixed4c)`, or in english: "Make an image in which layer `mixed4c` recognizes a lot of stuff. We're not optimizing for a specific feature any more, the network is just trying to overinterpret the image best it can.
#
# The network seems to like dogs and animal-like features due to the nature of the ImageNet dataset.
#
# However, we can force it to dream about a specific topic by changing the objective function to a specific feature or a combination of specific features. More castles!
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}, {"item_id": 2}]} colab_type="code" deletable=true editable=true executionInfo={"elapsed": 114, "status": "ok", "timestamp": 1457967665541, "user": {"color": "#1FA15D", "displayName": "<NAME>", "isAnonymous": false, "isMe": true, "permissionId": "12341152118244997759", "photoUrl": "https://lh3.googleusercontent.com/-XdUIqdMkCWA/AAAAAAAAAAI/AAAAAAAAAAA/4252rscbv5M/s128/photo.jpg", "sessionId": "1269ead540f76ce5", "userId": "108092561333339272254"}, "user_tz": -60} id="4GexZuwJdDmu" outputId="f140b073-7129-4889-f240-3d0e00530ada"
_ = render_deepdream(T(layer)[:,:,:,65], img0)
# + [markdown] colab_type="text" deletable=true editable=true id="mYsY6_Ngpfwl"
# Now, have fun! Upload your own images, and make the computer dream about the prettiest, creepiest, or most surreal things.
#
# Don't hesitate to use higher resolution inputs (also increase the number of octaves)! Here is an [example](http://storage.googleapis.com/deepdream/pilatus_flowers.jpg) of running the flower dream over a bigger image.
#
# The DeepDream [notebook](https://github.com/google/deepdream/blob/master/dream.ipynb) contains code with many more options to explore. You can guide the dreaming towards a specific image, or repeat it endlessly to produce dreamier dreams. If you're very patient, you can even make videos.
# + deletable=true editable=true
frame = img0
h, w = frame.shape[:2]
s = 0.05 # scale coefficient
for i in xrange(100):
frame = render_deepdream(tf.square(T('mixed4c')), img0=frame)
img = PIL.Image.fromarray(np.uint8(np.clip(frame, 0, 255)))
img.save("dream-%04d.jpg"%i)
# Zoom in while maintaining size
img = img.resize(np.int32([w*(1+s), h*(1+s)]))
t, l = np.int32([h*(1+s) * s / 2, w*(1+s) * s / 2])
img = img.crop([l, t, w-l, h-t])
img.load()
print img.size
frame = np.float32(img)
# + [markdown] colab_type="text" deletable=true editable=true id="mENNVQd3eD-h"
# <a id="fun"></a>
# ## More Fun
#
# For more more things to do, check out the [TensorFlow tutorials](http://tensorflow.org/tutorials). If you enjoyed this tutorial, you will probably like the [retraining example](https://github.com/tensorflow/tensorflow/tree/master/tensorflow/examples/image_retraining).
#
# In this tutorial, we used the Inception v3, trained on imagenet. We have recently released the [source code to Inception](https://github.com/tensorflow/models/tree/master/inception), allowing you to train an Inception network on your own data.
# + deletable=true editable=true
| 3_deepdream.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Sawppy Ackermann Math
# This Jupyter notebook works through the process of calculating the angle and velocity for every Sawppy wheel in response to a command to move the entire rover at a particular angular and linear velocity.
#
# ## Setup
# Sawppy chassis is represented by a list of six wheels, each listing a human-readable name and the X,Y coordinates relative to rover's center of motion.
# Sawppy chassis geometry (in meters)
SAWPPY_WHEELBASE_FRONT = 0.285
SAWPPY_WHEELBASE_MID = 0
SAWPPY_WHEELBASE_REAR = -0.257
SAWPPY_TRACK_FRONT = 0.23
SAWPPY_TRACK_MID = 0.26
SAWPPY_TRACK_REAR = 0.23
class ChassisWheel(object):
"""Information needed to calculate angle and speed for a specific wheel.
Axis orientation conforms to REP103. (+X is forward, +Y is left, +Z is up)
https://www.ros.org/reps/rep-0103.html
"""
def __init__(self, name, offset_front, offset_left):
"""Initialize a chassis wheel instance with X & Y relative to center.
Args:
name : name for this wheel
offset_front : front/back relative to center, positive forward.
offset_left : left/right relative to center, positive left.
"""
self.name = name
self.offset_front = offset_front
self.offset_left = offset_left
class ChassisWheelAngleSpeed(object):
"""Results of chassis geometry calculation for the named wheel."""
def __init__(self, name, angle, velocity):
"""Initialize a chassis wheel with desired angle and velocity.
Args:
name : name for this wheel
angle : steering angle for this wheel in radians.
velocity : rolling velocity for this wheel in meters/second
"""
self.name = name
self.angle = angle
self.velocity = velocity
test_chassis = []
test_chassis.append(
ChassisWheel(
'front_left', SAWPPY_WHEELBASE_FRONT, SAWPPY_TRACK_FRONT,
),
)
test_chassis.append(
ChassisWheel(
'front_right', SAWPPY_WHEELBASE_FRONT, -SAWPPY_TRACK_FRONT,
),
)
test_chassis.append(
ChassisWheel(
'mid_left', SAWPPY_WHEELBASE_MID, SAWPPY_TRACK_MID,
),
)
test_chassis.append(
ChassisWheel(
'mid_right', SAWPPY_WHEELBASE_MID, -SAWPPY_TRACK_MID,
),
)
test_chassis.append(
ChassisWheel(
'rear_left', SAWPPY_WHEELBASE_REAR, SAWPPY_TRACK_REAR,
),
)
test_chassis.append(
ChassisWheel(
'rear_right', SAWPPY_WHEELBASE_REAR, -SAWPPY_TRACK_REAR,
),
)
# ## Input
# +
import math
# Commanded inputs
velocityAngular = -0.2*math.pi # radians/sec
velocityLinear = 0 # meters/sec
# -
# ## Calculation
# ### Center of turn
#
# Consider the state of the robot after one second of traveling at commanded `velocityAngular` and `velocityLinear`. It would be pointing at `velocityAngular` radians off of +X axis (marking forward) and have covered `velocityLinear` distance. This describes an arc. Where is the center of the arc?
#
# One way to think about this problem is to consider the case where angular velocity is `2*pi`. After one second, the robot has traveled in a complete circle and `velocityLinear` is the circumference of that circle. Distance to the center of this circle would therefore be the radius, or `velocityLinear/2*pi`
#
# This formula holds for other values of `velocityAngular`. The linear distance traveled is some fraction of a circle, and dividing by the angular velocity returns the center of that circle.
#
# If `velocityAngular` is zero, we are traveling straight forward which is mathematically equivalent to a circle with infinite radius. This is set as our default value.
turnCenter = math.inf
if velocityAngular != 0:
turnCenter = velocityLinear / velocityAngular
print(turnCenter)
# ### Wheel Angle and Velocity
# Once the center of turn has been calculated, we can calculate the angle and distance from that point to each wheel.
# The angle becomes the [Ackermann steering angle](https://en.wikipedia.org/wiki/Ackermann_steering_geometry) for that wheel.
# The distance is compared with the center of turn, and the ratio determines velocity for that wheel.
calculated_results = []
for wheel in test_chassis:
if turnCenter == math.inf:
# Heading directly forward or back
wheel_angle = 0
wheel_velocity = velocityLinear
else:
# Dimensions of a trialge representing the wheel relative to center of turn.
opposite = wheel.offset_front
adjacent = turnCenter - wheel.offset_left
hypotenuse = math.sqrt(pow(opposite,2)+pow(adjacent,2))
if wheel.offset_front == 0:
wheel_angle = 0
else:
wheel_angle = math.atan(opposite / adjacent)
if velocityLinear == 0:
wheel_velocity = velocityAngular * hypotenuse
else:
wheel_velocity = math.copysign(velocityAngular * hypotenuse, velocityLinear)
# If center of turn is inside the wheel, we need to reverse direction.
if (turnCenter >= 0 and wheel.offset_left > 0 and wheel.offset_left > turnCenter) or \
(turnCenter < 0 and wheel.offset_left < 0 and wheel.offset_left < turnCenter):
wheel_velocity = wheel_velocity * -1
calculated_results.append(ChassisWheelAngleSpeed(wheel.name, wheel_angle, wheel_velocity))
# + tags=[]
import matplotlib.pyplot as plt
# if using a Jupyter notebook, include:
# %matplotlib inline
plot_x = []
plot_y = []
plot_u = []
plot_v = []
wheel_state = dict()
for wheel in test_chassis:
wheel_state[wheel.name] = dict()
wheel_state[wheel.name]['x'] = wheel.offset_front
wheel_state[wheel.name]['y'] = wheel.offset_left
wheel_state[wheel.name]['hypotenuse'] = math.sqrt(pow(wheel.offset_front,2)+pow(wheel.offset_left,2))
for wheel in calculated_results:
plot_x.append(wheel_state[wheel.name]['x'])
plot_y.append(wheel_state[wheel.name]['y'])
wheel_state[wheel.name]['angle'] = wheel.angle
wheel_state[wheel.name]['velocity'] = wheel.velocity
plot_u.append(math.cos(wheel.angle)*wheel.velocity)
plot_v.append(math.sin(wheel.angle)*wheel.velocity)
fig, ax = plt.subplots()
ax.set_title('Sawppy wheel angle and velocity')
ax.quiver(plot_x, plot_y, plot_u, plot_v)
ax.axis([-0.4, 0.4, -0.4, 0.4])
# -
# Print the input and output of this particular scenario in a format suitable for the unit test data file `chassis_wheel_calculator_tests.csv`
print('{: .6f},{: .6f},{: .6f},{: .6f},{: .6f},{: .6f},{: .6f},{: .6f},{: .6f},{: .6f},{: .6f},{: .6f},{: .6f},{: .6f}'.format(
velocityAngular, velocityLinear,
wheel_state['front_left']['angle'],
wheel_state['front_left']['velocity'],
wheel_state['front_right']['angle'],
wheel_state['front_right']['velocity'],
wheel_state['mid_left']['angle'],
wheel_state['mid_left']['velocity'],
wheel_state['mid_right']['angle'],
wheel_state['mid_right']['velocity'],
wheel_state['rear_left']['angle'],
wheel_state['rear_left']['velocity'],
wheel_state['rear_right']['angle'],
wheel_state['rear_right']['velocity']))
| jupyter/Sawppy Ackermann Math.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Gamma ray spectroscopy
# ## Functions
# +
# # %load ../setup.py
"""
Packages for plotting and other stuff
version: 1.0
author: Riasat
"""
# # %matplotlib widget
# data loading
import pandas as pd
# data maipulation
import pwlf
import numpy as np
from scipy.interpolate import interp1d, UnivariateSpline
from scipy.signal import find_peaks
# plotting tools
import matplotlib.pyplot as plt
# extra tweaks
import warnings
warnings.filterwarnings("ignore")
# plot tweaks
plt.style.use("seaborn-poster")
pd.options.display.max_columns = None
pd.options.display.float_format = "{:.5f}".format
# function for extrapolation
def extrapolate1d(x, y):
f = interp1d(x, y, kind="linear", fill_value="extrapolate")
a = np.arange(0, x[len(x) - 1], 0.001)
b = f(a)
return a, b
# function for interpolation
def interpolate1d(x, y):
f = interp1d(x, y, kind="linear", fill_value="extrapolate")
a = np.arange(x[0], x[len(x) - 1], 0.001)
b = f(a)
return a, b
# function for interpolation
def interpolate2d(x, y):
f = interp1d(x, y, kind="quadratic", fill_value="extrapolate")
a = np.arange(x[0], x[len(x) - 1], 0.001)
b = f(a)
return a, b
# function for interpolation
def interpolate3d(x, y):
f = interp1d(x, y, kind="cubic", fill_value="extrapolate")
a = np.arange(x[0], x[len(x) - 1], 0.001)
b = f(a)
return a, b
# funciton for polynomial fitting
def polfit(a, b, c):
z = np.polyfit(a, b, c)
f = np.poly1d(z)
x = np.arange(a[0], a[len(a) - 1], 0.001)
y = f(x)
return x, y
# function for picewise linear fit
def picewise_linear_fit(x, y, segments):
my_pwlf = pwlf.PiecewiseLinFit(x, y) # fit my data
res = my_pwlf.fit(segments) # fit the data for n line segments
# slopes = myPWLF.calc_slopes() # calculate slopes
# predict for the determined points
xHat = np.linspace(min(x), max(x), num=10000)
yHat = my_pwlf.predict(xHat)
# calculate statistics
# p = myPWLF.p_values(method="non-linear", step_size=1e-4) # p-values
# se = myPWLF.se # standard errors
return xHat, yHat
# -
# ## Data
# +
file_name = "data_gamma_spec.xlsx"
bg_count = 42
# calibration data
data_cesium_calib = pd.read_excel(file_name, sheet_name="cs calibration")
cs_channel_og = data_cesium_calib["cs_channel"]
cs_counts_og = data_cesium_calib["cs_counts"]
data_cobalt_calib = pd.read_excel(file_name, sheet_name="co calibration")
co_channel_og = data_cobalt_calib["co_channel"]
co_counts_og = data_cobalt_calib["co_counts"]
# distance data
data_dist = pd.read_excel(file_name, sheet_name="distance")
distance_og = data_dist["distance"]
dist_counts_og = data_dist["counts"]
net_dist_counts_og = dist_counts_og - bg_count
data_dist["net counts pm"] = net_dist_counts_og
print(f"{data_cesium_calib}, \n{data_cobalt_calib}, \n{data_dist}")
# -
# ## Spectrum
# cesium
channel_interpolated_cs, counts_interpolated_cs = interpolate3d(cs_channel_og, cs_counts_og)
# cobalt
channel_interpolated_co, counts_interpolated_co = interpolate3d(co_channel_og, co_counts_og)
# naming the elements
element_name = ["Cesium-137", "Cobalt-60", "Cobalt-60 Lower", "Cobalt-60 Higher"]
channel_interpolated = [channel_interpolated_cs, channel_interpolated_co]
counts_interpolated = [counts_interpolated_cs, counts_interpolated_co]
channel_original = [cs_channel_og, co_channel_og]
counts_original = [cs_counts_og, co_counts_og]
# ### Peak determination
#
# +
res_name = ["Cesium-137", "Cobalt-60 lower peak", "Cobalt-60 upper peak"]
for i in range(2):
peak_id_max = find_peaks(counts_interpolated[i], height=np.max(counts_interpolated[i]) - 500)
heights = peak_id_max[1]["peak_heights"]
pos = channel_interpolated[i][peak_id_max[0]]
print(f"{element_name[i]}: \n\t channel = {pos} and peak = {heights}")
peak_counts = [14173.38421456, 1567.36215049, 1344.06124333]
peak_channel = [27.2, 48, 54]
known_energy = [0.662, 1.171, 1.332]
# -
# ### Cesium spectrum
# +
plt.style.use("seaborn-poster")
plt.figure(figsize=(15, 8))
# plt.axvspan(vi[6], vi[7], alpha=0.2)
# for i in range(6, 8):
# plt.annotate(f"{vi[i]:.2f}", xy=(vi[i]-0.5, 0), fontsize=14)
# plt.annotate(f"43029", xy=(24 + 0.5, 43029), fontsize=14)
plt.title(f"{element_name[0]} Spectrum")
plt.xlabel("channel number (V)")
plt.ylabel("counts per minute")
plt.plot(channel_interpolated_cs, counts_interpolated_cs, "--", label="interpolated points")
plt.plot(cs_channel_og, cs_counts_og, "o", markersize=9, label="original points")
plt.legend(loc="upper left")
plt.grid(alpha=0.3, which="major")
plt.minorticks_on()
plt.grid(alpha=0.2, which="minor", ls="--")
# -
# ### Cobalt-60 Spectrum
# +
plt.style.use("seaborn-poster")
plt.figure(figsize=(15, 8))
# plt.axvspan(vi[2], vi[3], alpha=0.2)
# plt.axvspan(vi[4], vi[5], alpha=0.2)
# for i in range(2, 6):
# plt.annotate(f"{vi[i]:.2f}", xy=(vi[i]-1, 300), fontsize=14)
# for i in range(1,3):
# plt.annotate(f"{peak_counts[i]}", xy=(peak_channel[i] + 0.5, peak_counts[i]), fontsize=14)
plt.title(f"{element_name[1]} spectrum")
plt.xlabel("channel number (V)")
plt.ylabel("counts per minute")
plt.plot(channel_interpolated_co, counts_interpolated_co, "--", label="interpolated points")
plt.plot(co_channel_og, co_counts_og, "o", markersize=9, label="original points")
plt.legend(loc="upper left")
plt.grid(alpha=0.3, which="major")
plt.minorticks_on()
plt.grid(alpha=0.2, which="minor", ls="--")
plt.show()
# -
# ## Calibration
# +
# extrapolated points
peak_channel_fit, known_energy_fit = polfit(peak_channel, known_energy, 1)
cal_chan_ext, cal_eng_ext = extrapolate1d(peak_channel_fit, known_energy_fit)
ckt = [item for item in cal_eng_ext if item >= 0]
plt.style.use("seaborn-poster")
plt.figure(figsize=(15, 8))
plt.title(f"Calibaration curve")
plt.xlabel("Channel Number(V)")
plt.ylabel("Energy of element(MeV)")
# plt.plot(peak_channel, known_energy)
plt.plot(cal_chan_ext[634:], ckt, "-", label="fitted curve")
for i in range(len(res_name)):
plt.plot(peak_channel[i], known_energy[i], "o", label=res_name[i])
plt.annotate(f"({peak_channel[i]}, {known_energy[i]:.3f})", xy=(peak_channel[i]+0.5,known_energy[i]-0.025), fontsize=14)
plt.legend(loc="upper left")
plt.grid(alpha=0.3, which="major")
plt.minorticks_on()
plt.grid(alpha=0.2, which="minor", ls="--")
plt.show()
# -
scatter_peaks = [8.26, 17.10]
cs_scatter_energy = np.interp(scatter_peaks, cal_chan_ext, cal_eng_ext)
print(f"\n Back-scattering peak: {cs_scatter_energy[0]:.3f} MeV\n Compton edge peak energy: {cs_scatter_energy[1]:.3f} MeV")
# +
# converting counts per minute to per second
net_counts_ps = net_dist_counts_og/60
constant_k = net_counts_ps*distance_og*distance_og
data_dist["counts ps"] = net_counts_ps
data_dist["constant k"] = constant_k
dist_fitted, dist_counts_fitted = polfit(distance_og, net_dist_counts_og, 6)
plt.style.use("seaborn-poster")
plt.figure(figsize=(15, 8))
plt.title(f"Inverse square law")
plt.xlabel("distance(cm)")
plt.ylabel("counts per minute")
plt.plot(dist_fitted,dist_counts_fitted, "--", label="interpolated points")
plt.plot(distance_og,dist_counts_og, "o", markersize=9, label="original points")
plt.legend(loc="upper right")
plt.grid(alpha=0.3, which="major")
plt.minorticks_on()
plt.grid(alpha=0.2, which="minor", ls="--")
plt.show()
print(f"{data_dist}")
| 4th sem practicals/gamma spectroscopy/gamma_spec.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/dunkelweizen/DS-Unit-1-Sprint-2-Data-Wrangling-and-Storytelling/blob/master/Cai_Nowicki_Make_Explanatory_Visualizations_Assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="NMEswXWh9mqw"
# # ASSIGNMENT
#
# ### 1) Replicate the lesson code. I recommend that you [do not copy-paste](https://docs.google.com/document/d/1ubOw9B3Hfip27hF2ZFnW3a3z9xAgrUDRReOEo-FHCVs/edit).
#
# Get caught up to where we got our example in class and then try and take things further. How close to "pixel perfect" can you make the lecture graph?
#
# Once you have something that you're proud of, share your graph in the cohort channel and move on to the second exercise.
#
# ### 2) Reproduce another example from [FiveThityEight's shared data repository](https://data.fivethirtyeight.com/).
#
# **WARNING**: There are a lot of very custom graphs and tables at the above link. I **highly** recommend not trying to reproduce any that look like a table of values or something really different from the graph types that we are already familiar with. Search through the posts until you find a graph type that you are more or less familiar with: histogram, bar chart, stacked bar chart, line chart, [seaborn relplot](https://seaborn.pydata.org/generated/seaborn.relplot.html), etc. Recreating some of the graphics that 538 uses would be a lot easier in Adobe photoshop/illustrator than with matplotlib.
#
# - If you put in some time to find a graph that looks "easy" to replicate you'll probably find that it's not as easy as you thought.
#
# - If you start with a graph that looks hard to replicate you'll probably run up against a brick wall and be disappointed with your afternoon.
#
#
#
#
#
#
#
#
#
#
#
# + [markdown] id="EsKWzuqf8A_h" colab_type="text"
# ###Make Prototypes
# + id="7SY1ZHawyZvz" colab_type="code" outputId="67eac46e-991c-42d2-888e-1b618713bc78" colab={"base_uri": "https://localhost:8080/", "height": 285}
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
plt.style.use('fivethirtyeight')
fake = pd.Series([38,3,2,1,2,4,6,5,5,33],
index=range(1,11))
fake.plot.bar(color='#ed713a', width=0.9);
# + id="_Bu6u3jN8zI6" colab_type="code" outputId="1909a371-fb4b-410b-f5f5-28815ff87c96" colab={"base_uri": "https://localhost:8080/", "height": 508}
style_list = ['default', 'classic'] + sorted(
style for style in plt.style.available if style != 'classic')
style_list
# + id="_C2uwBX59CaD" colab_type="code" outputId="2990f47d-dee8-4edb-a1c2-3660c89a9d4b" colab={"base_uri": "https://localhost:8080/", "height": 126}
fake2 = pd.Series(
[1, 1, 1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,1,
2,2,2,
3,3,3,
4,4,
5,5,5,
6,6,6,6,
7,7,7,7,7,
8,8,8,8,
9,9,9,9,
10,10,10,10,10,10,10,10,10,10,10,10,10,10,10,10])
fake2.head()
# + id="-JB02j-B9dUF" colab_type="code" outputId="eb3951e5-408c-431e-9e41-2a7d325ed519" colab={"base_uri": "https://localhost:8080/", "height": 303}
plt.style.use('fivethirtyeight')
fake2.value_counts().sort_index().plot.bar(color='#ed713a', width=0.9)
# + [markdown] id="fNhRt6E-9qOQ" colab_type="text"
# ###Annotate with text
# + id="qMy8zubk9sO_" colab_type="code" outputId="4a0476a8-690e-4d67-a837-fc91a074df39" colab={"base_uri": "https://localhost:8080/", "height": 355}
from IPython.display import display, Image
url = 'https://fivethirtyeight.com/wp-content/uploads/2017/09/mehtahickey-inconvenient-0830-1.png'
example = Image(url=url, width=400)
display(example)
# + id="2U5vDeLu95mD" colab_type="code" outputId="0a17a37e-2f00-4d43-fd0d-8f890648015d" colab={"base_uri": "https://localhost:8080/", "height": 325}
fig = plt.figure(facecolor='black')
ax = fake2.value_counts().sort_index().plot.bar(color="#ed713a", width=0.9);
ax.set(facecolor='black')
plt.xlabel('Rating', color='white')
plt.ylabel('Percent of total votes', color='white')
# + id="UUANk-I7-UkA" colab_type="code" outputId="751572fe-d5f8-42b6-f710-a8795a03b555" colab={"base_uri": "https://localhost:8080/", "height": 35}
list(range(0,50,10))
# + id="hcK15YbJ-XJQ" colab_type="code" outputId="5b84362a-7ebe-4606-8825-d291303ae9fd" colab={"base_uri": "https://localhost:8080/", "height": 453}
fig = plt.figure(facecolor='white', figsize=(5,4))
ax = fake.plot.bar(color='#ed713a', width=0.9)
ax.set(facecolor='white')
ax.patch.set_alpha(0.1)
plt.xlabel('Rating', fontweight='bold')
plt.ylabel('Percent of total votes', fontweight='bold')
plt.title('`An Inconvenient Sequel: Truth to Power` is divisive',
fontsize=12,
loc='left',
x=-0.1,
y=1.1,
fontweight= 'bold')
plt.text(x=-1.7, y=fake.max() + 4, s='IMDb ratings for the film as of Aug. 29', fontsize=10)
plt.xticks(rotation=0, color='#a7a7a7')
plt.yticks(range(0,50,10), labels=[f'{i}' if i != 40 else f'{i}%' for i in range(0,50,10)], color='#a7a7a7')
# + [markdown] id="yan3cosIALoc" colab_type="text"
# ###Reproduce with real data
# + id="_7CsJyMeAmgu" colab_type="code" outputId="378dfbbb-03f1-4367-a412-38f4fa7c6fc0" colab={"base_uri": "https://localhost:8080/", "height": 305}
df = pd.read_csv('https://raw.githubusercontent.com/fivethirtyeight/data/master/inconvenient-sequel/ratings.csv')
df.head()
# + id="5YNghW8MA3xf" colab_type="code" outputId="411057db-b59b-42a9-d9a8-b07ec7208f29" colab={"base_uri": "https://localhost:8080/", "height": 526}
df.dtypes
# + id="RY4X7DsKA8_N" colab_type="code" outputId="670014f3-2b04-458e-8067-8039fc4f65a0" colab={"base_uri": "https://localhost:8080/", "height": 308}
df['timestamp'] = pd.to_datetime(df['timestamp'])
df.describe()
# + id="DfjUyH-jBGh2" colab_type="code" outputId="cebf73a6-5fb1-4fc6-fe41-f8805079454b" colab={"base_uri": "https://localhost:8080/", "height": 35}
df['timestamp'].min()
# + id="Alm8wTsrBMWj" colab_type="code" outputId="359b2b0c-e6f3-4516-f11e-912d8dd6dcc0" colab={"base_uri": "https://localhost:8080/", "height": 35}
df['timestamp'].max()
# + id="jP_u4hKVBOwr" colab_type="code" outputId="0199393a-f6dc-42d8-85ef-ff7226b231e4" colab={"base_uri": "https://localhost:8080/", "height": 335}
df = df.set_index('timestamp')
df.head()
# + id="7o3C2luCBSnE" colab_type="code" outputId="01e67fa6-c77a-40b9-925b-8f8ae4f255c9" colab={"base_uri": "https://localhost:8080/", "height": 1000}
df['2017-08-29']
# + id="dZzZDkbIBYCY" colab_type="code" outputId="6fb663fe-cf93-4d1f-afa2-af6a96c2c67a" colab={"base_uri": "https://localhost:8080/", "height": 335}
lastday = df['2017-08-29']
lastday_filtered = lastday[lastday['category'] == 'IMDb users']
lastday_filtered.head()
# + id="i3RxZ27TBmds" colab_type="code" outputId="d8034cc9-b8aa-4049-c1c6-832218452a9d" colab={"base_uri": "https://localhost:8080/", "height": 315}
lastday_filtered['respondents'].plot()
# + id="Y8RDbAGuBrwC" colab_type="code" outputId="ac8af7c7-38d4-4f96-f9d9-058863211b0a" colab={"base_uri": "https://localhost:8080/", "height": 54}
lastday_filtered['category'].value_counts()
# + id="xuJkHy_PBy2B" colab_type="code" outputId="177db95d-78b8-4aa5-82f8-491eefbdf493" colab={"base_uri": "https://localhost:8080/", "height": 199}
pct_columns = [f'{i}_pct' for i in range(1,11)]
pct_columns
# + id="TCAsO0jYBygb" colab_type="code" colab={}
final = lastday_filtered.tail(1)
# + id="2-1LjPwvB_C0" colab_type="code" outputId="12d593a9-463d-4c13-f3f5-d5c97f195552" colab={"base_uri": "https://localhost:8080/", "height": 145}
final
# + id="-RSLzOK4CCxb" colab_type="code" outputId="c388734f-b170-4c00-dee0-cd1e5f6185f3" colab={"base_uri": "https://localhost:8080/", "height": 348}
final[pct_columns].T
# + id="PJNnqGomCG7W" colab_type="code" colab={}
plot_data = final[pct_columns].T
plot_data.index = range(1,11)
# + id="9ylMSh8xCOP_" colab_type="code" outputId="cfe4e8ac-155a-4a00-c4bd-67b78631fdf4" colab={"base_uri": "https://localhost:8080/", "height": 348}
plot_data
# + id="PItaLcFhCSu8" colab_type="code" outputId="25796a09-04ce-492e-bcf3-03774525d8d2" colab={"base_uri": "https://localhost:8080/", "height": 366}
plt.style.use('fivethirtyeight')
ax = plot_data.plot.bar(color='#ed713a', width=0.9)
plt.xlabel('Rating', fontsize=9, fontweight='bold')
plt.ylabel('Percent of total votes', fontsize=9, fontweight='bold')
plt.title('`An Inconvenient Sequel: Truth to Power` is divisive',
fontsize=12,
x=-0.1, y=1.1, loc='left',
fontweight='bold',
fontname='Tahoma')
plt.text(x=-1.7, y=plot_data.max() + 4, s='IMDb ratings for the film as of Aug. 29',
fontsize=11)
plt.xticks(rotation=0, color='#a7a7a7', fontsize=8, clip_on='false')
plt.yticks(range(0,50,10), labels=[f'{i}' if i != 40 else f'{i}%' for i in range(0,50,10)],
color='#a7a7a7', fontsize=8, clip_on='false')
ax.set(facecolor='white')
fig = plt.figure(facecolor='white', edgecolor='white')
fig.patch.set_facecolor('white')
legend = ax.legend()
legend.remove()
display(fig)
# + id="GiC-_BeSFIzI" colab_type="code" outputId="2569b024-b540-437f-bd4b-81bd429b2f6a" colab={"base_uri": "https://localhost:8080/", "height": 355}
display(example)
# + [markdown] id="Lk0Fn4c1IpPV" colab_type="text"
# ## Reproduce Another Graph
# + id="5maGmwACJRLz" colab_type="code" outputId="3a206167-33c0-4faa-e3bf-7f2c3c7a688a" colab={"base_uri": "https://localhost:8080/", "height": 400}
url = 'https://fivethirtyeight.com/wp-content/uploads/2017/04/roeder-scrabble-1.png'
example = Image(url=url, width=400)
display(example)
# + id="_dtx8OzWJz7F" colab_type="code" colab={}
scrabble = pd.read_csv('https://media.githubusercontent.com/media/fivethirtyeight/data/master/scrabble-games/scrabble_games.csv')
# + id="J000zVPiJ4p5" colab_type="code" outputId="d96c430f-c543-4dbb-ee1e-d293ffe1b327" colab={"base_uri": "https://localhost:8080/", "height": 305}
scrabble.head()
# + id="EDow2SiMJ9Kt" colab_type="code" colab={}
#first need to remove tie games
#scrabble = scrabble[~scrabble['tie']]
# + id="W_S0k76zKV7r" colab_type="code" colab={}
#scrabble['tie'].sample(20)
# + id="3hpYGaN_Kl1u" colab_type="code" colab={}
#all I need is the winning score and the losing score for each game
columns = ['gameid', 'winnerscore', 'loserscore']
scrabble = scrabble[columns]
# + id="oLTS2dURK3fJ" colab_type="code" outputId="b40a9eca-c7ec-4566-d6c8-204a0df96ac7" colab={"base_uri": "https://localhost:8080/", "height": 348}
scrabble.sample(10)
# + id="e5H5z9dELQaq" colab_type="code" outputId="1ee319d9-ae24-489e-b029-b7ee4972535c" colab={"base_uri": "https://localhost:8080/", "height": 288}
scrabble.describe()
# + id="w_c5jmwDLb7k" colab_type="code" colab={}
scrabble = scrabble.set_index('gameid')
# + id="gd_IoFxTN5OV" colab_type="code" outputId="bc606a68-f534-4689-feb0-3c33105a4eb7" colab={"base_uri": "https://localhost:8080/", "height": 378}
scrabble.sample(10)
# + id="VwSCO-AaN-GK" colab_type="code" outputId="4d022e8f-204f-4d3e-dc3e-69cd2c5cc499" colab={"base_uri": "https://localhost:8080/", "height": 288}
scrabble.describe
#there are...negative scores? I should remove those, nothing in the original graph goes below zero
# + id="Pa7pry7POHF3" colab_type="code" colab={}
condition = scrabble['winnerscore'] >= 1
scrabble = scrabble[condition]
condition = scrabble['loserscore'] >= 1
scrabble = scrabble[condition]
#initial histogram showed that most of the scores were 0/0 (I guess forfeits?) and that ruined the graph
# + id="ZQmfPpC4Rjb0" colab_type="code" outputId="9d5d1367-6e56-4969-f4dd-26e5babbba14" colab={"base_uri": "https://localhost:8080/", "height": 299}
plt.hist(scrabble['loserscore'], color='red', bins=100)
plt.hist(scrabble['winnerscore'], color='green', bins=200)
plt.title('700,000 games of Scrabble', fontweight='bold', fontsize=14, loc='left')
plt.show()
# + id="KPL2hL73Zbp_" colab_type="code" outputId="da7cbe30-ff46-480c-a572-3973a9296251" colab={"base_uri": "https://localhost:8080/", "height": 300}
scrabble['winnerscore'].plot.kde()
# + id="k_hlKPRhWG0m" colab_type="code" outputId="574ddc0f-a8a0-4b59-c779-c9582b38e621" colab={"base_uri": "https://localhost:8080/", "height": 400}
display(example)
# + id="P8G_v8dTWUep" colab_type="code" colab={}
#why are the numbers in my graph not the same?
# + id="93go5lM-PCWS" colab_type="code" outputId="501601fa-95a1-4d1f-b67b-f2d103ff66f6" colab={"base_uri": "https://localhost:8080/", "height": 534}
import seaborn as sns
sns.distplot(sample['loserscore'], bins=40, hist=True, color='green', kde=False).set_title('70,000 games of Scrabble')
sns.distplot(sample['winnerscore'], bins=75, hist=True, color='red', kde=False)
ax.set_xlabel('Score')
ax.set_yticks(range(0,10000, 2500));
# + [markdown] id="xC2OBVBpK2-a" colab_type="text"
#
# + [markdown] id="0wSrBzmJyWaV" colab_type="text"
# # STRETCH OPTIONS
#
# ### 1) Reproduce one of the following using the matplotlib or seaborn libraries:
#
# - [thanksgiving-2015](https://fivethirtyeight.com/features/heres-what-your-part-of-america-eats-on-thanksgiving/)
# - [candy-power-ranking](https://fivethirtyeight.com/features/the-ultimate-halloween-candy-power-ranking/)
# - or another example of your choice!
#
# ### 2) Make more charts!
#
# Choose a chart you want to make, from [Visual Vocabulary - Vega Edition](http://ft.com/vocabulary).
#
# Find the chart in an example gallery of a Python data visualization library:
# - [Seaborn](http://seaborn.pydata.org/examples/index.html)
# - [Altair](https://altair-viz.github.io/gallery/index.html)
# - [Matplotlib](https://matplotlib.org/gallery.html)
# - [Pandas](https://pandas.pydata.org/pandas-docs/stable/visualization.html)
#
# Reproduce the chart. [Optionally, try the "Ben Franklin Method."](https://docs.google.com/document/d/1ubOw9B3Hfip27hF2ZFnW3a3z9xAgrUDRReOEo-FHCVs/edit) If you want, experiment and make changes.
#
# Take notes. Consider sharing your work with your cohort!
# + id="dRJkKftiy5BJ" colab_type="code" colab={}
# More Work Here
| Cai_Nowicki_Make_Explanatory_Visualizations_Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# source: http://www.scipy-lectures.org/intro/language/reusing_code.html#scripts
# # 1. Getting started with Python for science
# ### 1.2.5. Reusing code: scripts and modules
#
# %run test.py
# %run test.py aa sdf.p sdfe
import os
os.listdir('.')
os.getcwd()
import numpy as np
np.linspace(0,10,20)
# ##### Creating modules
import demo
demo.printa()
demo.c
# +
# demo?
# -
who
whos
dir(demo)
demo.__name__
demo.__doc__
import importlib
importlib.import_module('demo')
reload(demo)
# %run demo.py
# #### 1.2.5.5 Scripts or modules? How to organize your code
# **Note**: Rule of thumb
# * Sets of instructions that are called several times should be written inside **functions** for better code reusability.
# * Functions (or other bits of code) that are called from several scripts should be written inside a **module**, so that the module is imported in the different scripts (do not copy-and-paste your functions in the different scripts!)
#
# **PYTHONPATH**
# ```
# export PYTHONPATH=$PYTHONPATH:/home/michael/lib
# ```
import sys
new_path = "/home/michael/user_defined_modules"
if new_path not in sys.path:
sys.path.append(new_path)
import scipy
scipy.__file__
import scipy.version
import scipy.ndimage.morphology
from scipy.ndimage import morphology
os.getcwd()
os.listdir(os.curdir)
os.mkdir('junkdir')
os.rename('junkdir', 'foodir')
'junkdir' in os.listdir(os.curdir)
'foodir' in os.listdir(os.curdir)
os.rmdir('foodir')
a = os.path.abspath("demo.py/")
os.path.dirname(a)
os.path.split(a)
os.path.basename(a)
os.path.splitext(os.path.basename(a))
os.path.exists(a)
os.path.isfile(a)
os.path.isdir(a)
os.path.expanduser('~/local')
os.path.join(os.path.expanduser('~'), 'local','bin')
os.system('ls')
import sh
com = sh.ls()
print(com)
type(com)
for dirpath, dirnames, filenames in os.walk(os.curdir):
print(dirpath, dirnames)
for fp in filenames:
print(os.path.abspath(fp))
os.environ.keys()
os.environ['PYTHONPATH']
os.getenv('PATHONPATH')
import shutil
import glob
glob.glob('*.py')
sys.platform
sys.version
sys.prefix
sys.argv
sys.path
import pickle
l = [1, None, 'Stan']
pickle.dump(l, file('test.pkl', 'w'))
pickle.load(file('test.pkl'))
# #### 1.2.8.2 Catching exceptions
# ### try/ except
while True:
try:
x = int(raw_input('Please enter a number:'))
break
except ValueError:
print('That was no valid number. Try again...')
# ### Easier to ask for forgiveness than for permission
def print_sorted(collection):
try:
collection.sort()
except AttributeError:
pass
print(collection)
print_sorted([1,3,2])
print_sorted((1,3,2))
# #### 1.2.48.3 Raising exceptions
# * Capturing and reraising an exception:
def filter_name(name):
try:
name = name.encode('ascii')
except UnicodeError as e:
if name == 'Gaël':
print('OK, Gaël!')
else:
raise e
return name
filter_name('Gaël')
# * Exceptions to pass messages between parts of the code
def achilles_arrow(x):
if abs(x-1) < 1e-3:
raise StopIteration
x = 1 - (1-x)/2.
return x
x = 0
while True:
try:
x = achilles_arrow(x)
except StopIteration:
break
x
# ## 1.3 NumPy: creating and manipulating numerical data
# an overview of NumPy, the core tool for performant numerical computing with Python.
# ### 1.3.1 The NumPy array object
# NumPy arrays
# --------------
#
# ---
#
# 1. **Python** object:
# - high-level number objects: integers, floating point
# - container: lists(costless insertion and append), dictionaries(fast lookup)
#
# 2. **NumPy** provides:
# * extension package to Python for multi-dimensional arrays
# * closer to hardware(efficiency)
# * designed for scientificcomputation(convenience)
# * Also known as *array oriented computing*
import numpy as np
a = np.array([0,1,2,3])
a
# For example, An array contraining:
# * values of an experiments/simulation at discrete time steps
# * signal recorded by a measurement device, e.g. sound wave
# * pixels of an images, grey-level or colour
# * 3-D data measured at different X-Y-Z positions, e.g. MRI scan
# **Why it is useful**: Memory-efficient container that provides fast numerical operations.
L = range(1000)
# %timeit [i**2 for i in L]
a = np.arange(1000)
# %timeit a**2
# ### NumPy Reference documentation
# * On the web: http://docs.scipy.org/
# * Interactiv help:
# +
# np.array?
# -
# np.lookfor('create array')
# +
# np.con*?
# -
# Exercise: Simple arrays
# * Create a simple two dimensional array. First, redo the examples from above. And then create your own: how about odd numbers counting backwards on the first row, and even numbers on the second?
# * Use the functions len(), numpy.shape() on these arrays. How do they relate to each other? And to the ndim attribute of the arrays?
#
# ### <u>Functions for creating arrays</u>
#
# ---
# In practice, we rarely enter items one by one
#
# * Evenly spaced:
#
# ---
#
a = np.arange(10)
a
b = np.arange(1, 9, 2)
b
# * or by number of points:
c = np.linspace(0, 1, 6)
c
d = np.linspace(0, 1, 5, endpoint=False)
d
# Common arrays:
a = np.ones((3,3))
a
b = np.zeros((2,2))
b
c = np.eye(3)
c
d = np.diag(np.array([1, 2, 3, 4]))
d
# * np.random: random numbers
a = np.random.rand(4) # uniform in [0, 1]
a
b = np.random.randn(4) # Gaussian
b
np.random.seed(1234) # setting the random seed
np.random.rand(2,2)
# #### Exercise: Creating arrays using functions
# * Experiment with arange, linspace, ones, zeros, eye and diag
# * Create different kinds of arrays with numbers
# * Try setting the seed before creating an array with random values
# * Look at the function np.empty. What does it do? When might this be useful?
np.arange(5)
np.linspace(1,5,5)
np.ones((3,4))
np.zeros((3,5))
np.eye(4)
np.empty(3)
np.empty((2,3))
np.empty((1,9))
np.random.choice(5, 3, p=[0.1, 0, 0.3, 0.6, 0])
np.array([1+2j,3+4j,5+6*1j]).dtype
np.array([True, False, False]).dtype
f = np.array(['Bonjour', 'Hello', 'Hallo'])
f.dtype
import matplotlib.pyplot as plt
# %matplotlib inline
x = np.linspace(0, 3,20)
y = np.linspace(0,9,20)
plt.plot(x,y,'o')
plt.plot(x,y)
x = np.linspace(0,3,20)
y = np.linspace(0, 9, 20)
plt.plot(x,y)
a = np.arange(10)
a[::-1]
b = np.diag(np.arange(3))
b
b[2,1]
b[0]
a[2:9:3]
np.arange(0,51,10)[:, np.newaxis]
is_prime = np.ones((100,), dtype=bool)
N_max = int(np.sqrt(len(is_prime) - 1))
np.random.seed(3)
a = np.random.randint(0,21,15)
a
mask = (a % 3 ==0)
extract_from_a = a[mask]
extract_from_a
a[a % 3 ==0] =-1
a
a[[9,7]] = -100
a
a = np.arange(10)
idx = np.array([[3,4], [9,7]])
idx.shape, a[idx]
# ### 1.3.2 Numerical operations on arrays
# #### 1.3.2.1 Elementwise operations
# Basic operations
# * with scalars:
a = np.array([1,2,3,4])
a+1
2**a
b = np.ones(4) + 1
a = np.arange(10000)
# %timeit a +1
l = range(10000)
# %timeit [i+1 for i in l]
c = np.ones((3,3))
c.dot(c)
2**c
c**2
a = np.array([1,2,3,4])
b = np.array([4, 2, 2, 4])
a ==b
a = np.array([1, 2,3,4,])
b = np.array([ 4, 2, 2, 4])
c = np.array([1, 2, 3, 4])
np.array_equal(a, b), np.array_equal(a,c)
a = np.array([1,1,0,0], dtype=bool)
b = np.array([1,0,1,0], dtype=bool)
np.logical_or(a,b)
np.logical_and(a,b)
np.logical_xor(a,b)
# Transcendental functions:
a = np.arange(5)
np.sin(a), np.log(a), np.exp(a)
# +
# Shape mismatches
#a + np.array([1,2])
# Transpositon:
a = np.triu(np.ones((3,3)), 1)
np.triu(a)
# -
a.T
a += a.T
np.allclose(c,b)
# 1.3.2.2 Basic reductions
# computing sums
x = np.array([1,2,3,4])
np.sum(x)
x.sum()
x = np.array([[1, 1], [2,2]])
x.sum(axis=0)
x[:,0].sum(), x[:,1].sum()
x.sum(axis=1)
x[0,:].sum(), x[1,:].sum()
x = np.random.rand(2,2,2)
x.sum(axis=2), x.sum(axis=0)
x, x[0,1,:].sum()
# other reductions
# - works the same way (and take axis=)
x = np.array([1,3,2])
x.min(),x.max()
x.argmin(), x.argmax() # index of minimum, maxmum
np.all([True, True, False]), np.any([True, True, False])
a = np.zeros((100,100))
np.any(a !=0)
np.all(a==a)
a = np.array([1,2,3,2])
b = np.array([2,2,3,2])
c = np.array([6,4,4,5])
((a<=b) & (b<=c)).any()
# Statistics
x = np.array([1, 2,3,1])
y = np.array([[1,2,3],[5,6,1]])
x.mean()
np.median(x), np.mean(x)
np.median(y, axis=-1)
x.std()
# +
np.std(x)
# +
# np.cumsum?
# -
np.sum(y,axis=0)
np.cumsum(y, axis=0)
# ls
data = np.loadtxt('populations.txt')
year, hares, lynxes, carrots = data.T
plt.axes([0.2, 0.5, 0.5, 0.8])
plt.plot(year, hares,year, lynxes, year, carrots)
plt.axes([0.3,0.1,0.6,0.7])
plt.plot(year, hares,year, lynxes, year, carrots)
plt.legend(('Hares', 'Lynx','Carrot'), loc=(1.05,0.5))
pop = data[:,1:]
pop.mean(axis=0)
pop.std(axis=0)
np.argmax(pop, axis=1)
n_stories = 1000 # shuffled jumps
t_max = 200 # Position: cumulated jumps sum
t = np.arange(t_max)
steps = 2 * np.random.randint(0, 1+1, (n_stories, t_max))-1\
# we build the walks by summing steps along the time
pos = np.cumsum(steps, axis=1)
sq_distance = pos ** 2
pos, sq_distance
mean_sq_distance = np.mean(sq_distance, axis=0)
mean_sq_distance
plt.figure(figsize=(4,3))
plt.plot(t, np.sqrt(mean_sq_distance), 'g.', t, np.sqrt(t), 'y-')
plt.xlabel(r"$t$")
plt.ylabel(r"$\sqrt{\langle (\delta x)^2 \rangle}$")
plt.tight_layout()
# ### 1.3.2.3 Broadcasting
# * Basic operations on numpy arrays(addition, etc.) are elementwise
# * This works on arrays of the same size
# if arrays have different sizes Numpy can transform these arrays so that they all have the same size: this conversion is called Broadcasting.
#
# The image below gives an example of broadcasting:
a = np.tile(np.arange(0, 40, 10), (3,1)).T
a
b = np.array([0,1,2])
a + b
a = np.ones((4,5))
a[0] = 2
a = np.arange(0, 40, 10)
a.shape
a = a[:, np.newaxis]
a + b
mileposts = np.array([0, 198, 303, 736, 871, 1175, 1475, 1544, 1913, 2448])
distance_array = np.abs(mileposts - mileposts[:,np.newaxis])
#distance_array
x, y = np.arange(5), np.arange(5)[:, np.newaxis]
distance = np.sqrt(x **2 + y ** 2)
distance
plt.pcolor(distance)
plt.colorbar()
# Remark: the numpy.ogrid() function allows to directly create vectors x and y of the previous example, with two "significat dimensions":
x, y = np.ogrid[0:5, 0:5]
x, y, x.shape, y.shape
x, y = np.mgrid[0:4, 0:4]
x, y
# #### 1.3.2.4 Array shape manipulation
# Flattening
a = np.arange(1,7).reshape(2,3)
a.ravel()
a.T
a.T.ravel()
b =a.reshape(3,2)
b[0,0] = 99
a
a = np.zeros((3,2))
b = a.T.reshape(3*2)
b[0] = 9
a, b
z = np.array([1,2,3])
z
z[:, np.newaxis]
a = np.arange(4*3*2).reshape(4,3,2)
a
a[0,2,1]
b = a.transpose(1,2, 0)
b.shape
b[2,1,0] = -1
a[0,2,1]
# ## Resizing
# Size of an array can be changed with ndarray.resize:
#
a = np.arange(4)
#a.resize((8,))
a
# #### 1.3.2.5 Sorting data
# Sorting along an axis:
a = np.array([[4,3,5], [1,2,1]])
b = np.sort(a, axis=1)
b
a = np.array([4,3,1,2,])
j = np.argsort(a)
a = np.array([4,3,2,5])
a
j_max = np.argmax(a)
j_min = np.argmin(a)
j_max, j_min
# +
#ravel, sort and reshape
#array_equal, np.random.shuffle out-of-place
# -
# #### 1.3.2.6 Summary
# what do you need to know to get started?
# * Know how to create arrays: array, arange, ones, zeros
# * Know the shape of the array.shape, then use slicing to obtain different views of the array: array[::2],etc. Adjust the shape of the array using reshape or flatten it with ravel.
# * Obtain a subset of the elements of an array and/or modify their values with masks
# ```
# a[a < 0 ] = 0
# ```
# * Know miscellaneous operations on arrays, such as finding the mean or max(array.max()), array.mean(). No need to retain everything, but have the reflex to search in the documentation(online docs, help(), lookfor()) !!!
# * For advanced use: master the indexing with arrays of integers, as c
# +
a[a<0] = 0
# -
a
| scipy_lecture/1.2_scipy_lecture_note.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="http://hilpisch.com/tpq_logo.png" alt="The Python Quants" width="35%" align="right" border="0"><br>
# # Python for Finance (2nd ed.)
#
# **Mastering Data-Driven Finance**
#
# © Dr. <NAME> | The Python Quants GmbH
#
# <img src="http://hilpisch.com/images/py4fi_2nd_shadow.png" width="300px" align="left">
# # Model Calibration
# ## The Data
# + uuid="f087ed8b-b432-48a9-a791-451ac477c046"
import numpy as np
import pandas as pd
import datetime as dt
# -
from pylab import mpl, plt
plt.style.use('seaborn')
mpl.rcParams['font.family'] = 'serif'
# %matplotlib inline
# + uuid="1d45b83a-110e-4c94-9404-859554d3f2d7"
import sys
sys.path.append('../')
sys.path.append('../dx')
# -
dax = pd.read_csv('../../source/tr_eikon_option_data.csv',
index_col=0)
for col in ['CF_DATE', 'EXPIR_DATE']:
dax[col] = dax[col].apply(lambda date: pd.Timestamp(date))
dax.info()
dax.set_index('Instrument').head(7)
initial_value = dax.iloc[0]['CF_CLOSE']
calls = dax[dax['PUTCALLIND'] == 'CALL'].copy()
puts = dax[dax['PUTCALLIND'] == 'PUT '].copy()
calls.set_index('STRIKE_PRC')[['CF_CLOSE', 'IMP_VOLT']].plot(
secondary_y='IMP_VOLT', style=['bo', 'rv'], figsize=(10, 6));
# plt.savefig('../../images/ch21/dx_cal_01.png');
ax = puts.set_index('STRIKE_PRC')[['CF_CLOSE', 'IMP_VOLT']].plot(
secondary_y='IMP_VOLT', style=['bo', 'rv'], figsize=(10, 6))
ax.get_legend().set_bbox_to_anchor((0.25, 0.5));
# plt.savefig('../../images/ch21/dx_cal_02.png');
# ## Model Calibration
# ### Relevant Market Data
limit = 500
option_selection = calls[abs(calls['STRIKE_PRC'] - initial_value) < limit].copy()
option_selection.info()
option_selection.set_index('Instrument').tail()
option_selection.set_index('STRIKE_PRC')[['CF_CLOSE', 'IMP_VOLT']].plot(
secondary_y='IMP_VOLT', style=['bo', 'rv'], figsize=(10, 6));
# plt.savefig('../../images/ch21/dx_cal_03.png');
# ### Option Modeling
# + uuid="b2f6b10c-bf01-46f6-958a-e0e6266adbe8"
import dx
# + uuid="c93f5398-8620-48d1-9c6b-ae0c79653751"
pricing_date = option_selection['CF_DATE'].max()
# + uuid="c191a3e9-286e-4c84-8044-aaf05f88377b"
me_dax = dx.market_environment('DAX30', pricing_date)
# -
maturity = pd.Timestamp(calls.iloc[0]['EXPIR_DATE'])
# + uuid="52879659-97e7-4c0f-a14f-b29bea60b3c8"
me_dax.add_constant('initial_value', initial_value)
me_dax.add_constant('final_date', maturity)
me_dax.add_constant('currency', 'EUR')
# + uuid="f1862ff5-ef88-4364-beba-502872ac5450"
me_dax.add_constant('frequency', 'B')
me_dax.add_constant('paths', 10000)
# + uuid="f0aedd9b-a2c9-4713-a02d-cc5a8959408e"
csr = dx.constant_short_rate('csr', 0.01)
me_dax.add_curve('discount_curve', csr)
# + uuid="11c8a827-c54c-451a-93c3-9f8db6df970c"
me_dax.add_constant('volatility', 0.2)
me_dax.add_constant('lambda', 0.8)
me_dax.add_constant('mu', -0.2)
me_dax.add_constant('delta', 0.1)
# + uuid="fd70a940-895e-43a4-a66c-d2c3bcc7c285"
dax_model = dx.jump_diffusion('dax_model', me_dax)
# + uuid="fe6d0c61-4907-4466-98ae-3ca782f83964"
me_dax.add_constant('strike', initial_value)
me_dax.add_constant('maturity', maturity)
# + uuid="8e36b826-9439-49f2-b4fa-e35928b8df41"
payoff_func = 'np.maximum(maturity_value - strike, 0)'
# + uuid="d33efcec-e027-4b1a-8fa1-c13696779de3"
dax_eur_call = dx.valuation_mcs_european('dax_eur_call',
dax_model, me_dax, payoff_func)
# + uuid="cedcee4e-1135-4b38-9381-d74b306de63e"
dax_eur_call.present_value()
# + uuid="a4a9ab6f-0810-403f-b172-d08deb80c582"
option_models = {}
for option in option_selection.index:
strike = option_selection['STRIKE_PRC'].loc[option]
me_dax.add_constant('strike', strike)
option_models[strike] = dx.valuation_mcs_european(
'eur_call_%d' % strike,
dax_model,
me_dax,
payoff_func)
# + uuid="89ec1029-091b-4ab1-8d60-b7a604c02f69"
def calculate_model_values_old(p0):
''' Returns all relevant option values.
Parameters
===========
p0: tuple/list
tuple of kappa, theta, volatility
Returns
=======
model_values: dict
dictionary with model values
'''
volatility, lamb, mu, delta = p0
dax_model.update(volatility=volatility, lamb=lamb, mu=mu, delta=delta)
model_values = {}
for strike in option_models:
model_values[strike] = option_models[strike].present_value(fixed_seed=True)
return model_values
# + uuid="89ec1029-091b-4ab1-8d60-b7a604c02f69"
def calculate_model_values(p0):
''' Returns all relevant option values.
Parameters
===========
p0: tuple/list
tuple of kappa, theta, volatility
Returns
=======
model_values: dict
dictionary with model values
'''
volatility, lamb, mu, delta = p0
dax_model.update(volatility=volatility, lamb=lamb,
mu=mu, delta=delta)
return {
strike: model.present_value(fixed_seed=True)
for strike, model in option_models.items()
}
# + uuid="d16e0a95-8543-4b08-b056-3d4f83d05e51"
calculate_model_values((0.1, 0.1, -0.4, 0.0))
# -
# ### Calibration Procedure
# + uuid="6d94c077-ebf9-46e3-8185-d4c80d12116d"
i = 0
def mean_squared_error(p0):
''' Returns the mean-squared error given
the model and market values.
Parameters
===========
p0: tuple/list
tuple of kappa, theta, volatility
Returns
=======
MSE: float
mean-squared error
'''
global i
model_values = np.array(list(calculate_model_values(p0).values()))
market_values = option_selection['CF_CLOSE'].values
option_diffs = model_values - market_values
MSE = np.sum(option_diffs ** 2) / len(option_diffs)
if i % 75 == 0:
if i == 0:
print('%4s %6s %6s %6s %6s --> %6s' %
('i', 'vola', 'lambda', 'mu', 'delta', 'MSE'))
print('%4d %6.3f %6.3f %6.3f %6.3f --> %6.3f' %
(i, p0[0], p0[1], p0[2], p0[3], MSE))
i += 1
return MSE
# + uuid="489f4f60-5237-4eff-be12-19abc6583ecb"
mean_squared_error((0.1, 0.1, -0.4, 0.0))
# + uuid="d4d06a9e-929b-4a02-95c1-433529015988"
import scipy.optimize as spo
# + uuid="15b93990-3228-4330-b5df-10915827ebcc"
# %%time
i = 0
opt_global = spo.brute(mean_squared_error,
((0.10, 0.201, 0.025), # range for volatility
(0.10, 0.80, 0.10), # range for jump intensity
(-0.40, 0.01, 0.10), # range for average jump size
(0.00, 0.121, 0.02)), # range for jump variability
finish=None)
# + uuid="359c3f5f-4f47-4e53-a916-85d3c745ed1b"
mean_squared_error(opt_global)
# + uuid="9fd46baf-28af-4276-a1c1-b3e521550cdd"
# %%time
i = 0
opt_local = spo.fmin(mean_squared_error, opt_global,
xtol=0.00001, ftol=0.00001,
maxiter=200, maxfun=550)
# + uuid="d7c22ea7-b3dd-4408-9ef2-3a986354ba26"
i = 0
mean_squared_error(opt_local)
# + uuid="70b888ee-8d31-46b2-86be-ad95dedd347a"
calculate_model_values(opt_local)
# + uuid="d6cf96da-5139-435e-8ed9-6be593fa7a15"
option_selection['MODEL'] = np.array(list(calculate_model_values(opt_local).values()))
option_selection['ERRORS_EUR'] = (option_selection['MODEL'] -
option_selection['CF_CLOSE'])
option_selection['ERRORS_%'] = (option_selection['ERRORS_EUR'] /
option_selection['CF_CLOSE']) * 100
# + uuid="716cd81a-0e06-405c-ade7-7fa407ce19cb"
option_selection[['MODEL', 'CF_CLOSE', 'ERRORS_EUR', 'ERRORS_%']]
# + uuid="91c24e54-e18a-4cfe-88b2-82ef82947b9c"
round(option_selection['ERRORS_EUR'].mean(), 3)
# + uuid="91c24e54-e18a-4cfe-88b2-82ef82947b9c"
round(option_selection['ERRORS_%'].mean(), 3)
# + uuid="9d2912c2-9ab3-4423-95d5-984df6f9f31e"
fix, (ax1, ax2, ax3) = plt.subplots(3, sharex=True, figsize=(10, 10))
strikes = option_selection['STRIKE_PRC'].values
ax1.plot(strikes, option_selection['CF_CLOSE'], label='market quotes')
ax1.plot(strikes, option_selection['MODEL'], 'ro', label='model values')
ax1.set_ylabel('option values')
ax1.legend(loc=0)
wi = 15
ax2.bar(strikes - wi / 2., option_selection['ERRORS_EUR'], width=wi)
ax2.set_ylabel('errors [EUR]')
ax3.bar(strikes - wi / 2., option_selection['ERRORS_%'], width=wi)
ax3.set_ylabel('errors [%]')
ax3.set_xlabel('strikes');
# plt.savefig('../../images/ch21/dx_cal_04.png');
# -
# ## Market-Based Valuation
# ### Modeling Option Positions
# + uuid="40decf0f-1908-48e8-b5c3-0fa5b667575c"
me_dax = dx.market_environment('me_dax', pricing_date)
me_dax.add_constant('initial_value', initial_value)
me_dax.add_constant('final_date', pricing_date)
me_dax.add_constant('currency', 'EUR')
# + uuid="88c93f7b-7944-4724-8b7d-a8f9d231d926"
me_dax.add_constant('volatility', opt_local[0])
me_dax.add_constant('lambda', opt_local[1])
me_dax.add_constant('mu', opt_local[2])
me_dax.add_constant('delta', opt_local[3])
# + uuid="8d92c037-79db-4663-9ad4-863dc720d160"
me_dax.add_constant('model', 'jd')
# + uuid="ee26163e-cfc2-4bd4-99ef-d9d877dc9592"
payoff_func = 'np.maximum(strike - instrument_values, 0)'
# + uuid="f4882acb-2157-4073-b2e7-b9c79a428a2c"
shared = dx.market_environment('share', pricing_date)
shared.add_constant('maturity', maturity)
shared.add_constant('currency', 'EUR')
# + uuid="e48619d3-c12d-4387-bc0d-51dccd00d19e"
option_positions = {}
option_environments = {}
for option in option_selection.index:
option_environments[option] = dx.market_environment(
'am_put_%d' % option, pricing_date)
strike = option_selection['STRIKE_PRC'].loc[option]
option_environments[option].add_constant('strike', strike)
option_environments[option].add_environment(shared)
option_positions['am_put_%d' % strike] = \
dx.derivatives_position(
'am_put_%d' % strike,
quantity=np.random.randint(10, 50),
underlying='dax_model',
mar_env=option_environments[option],
otype='American',
payoff_func=payoff_func)
# -
# ### The Options Portfolio
# + uuid="f885f822-7d78-4841-99c5-f30f1bf23ebb"
val_env = dx.market_environment('val_env', pricing_date)
val_env.add_constant('starting_date', pricing_date)
val_env.add_constant('final_date', pricing_date)
val_env.add_curve('discount_curve', csr)
val_env.add_constant('frequency', 'B')
val_env.add_constant('paths', 25000)
# + uuid="d9a3a473-3cfb-49ab-be94-848349683a9a"
underlyings = {'dax_model' : me_dax}
# + uuid="590ae8ed-39b9-42a3-84f2-35b2b400d8b8"
portfolio = dx.derivatives_portfolio('portfolio', option_positions,
val_env, underlyings)
# + uuid="11539da8-35b0-4daf-94ee-52aae7d9fca8"
# %time results = portfolio.get_statistics(fixed_seed=True)
# + uuid="883fa311-3410-4572-a8d1-a526e2914f69"
results.round(1)
# + uuid="8ceae186-64e2-405d-8563-b6349c0f13b2"
results[['pos_value','pos_delta','pos_vega']].sum().round(1)
# -
# <img src="http://hilpisch.com/tpq_logo.png" alt="The Python Quants" width="35%" align="right" border="0"><br>
#
# <a href="http://tpq.io" target="_blank">http://tpq.io</a> | <a href="http://twitter.com/dyjh" target="_blank">@dyjh</a> | <a href="mailto:<EMAIL>"><EMAIL></a>
| code/ch21/21_market_valuation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.10 64-bit (''base'': conda)'
# name: python3
# ---
# + id="aTuyBfHqzAcN" executionInfo={"status": "ok", "timestamp": 1629215281083, "user_tz": -540, "elapsed": 3534, "user": {"displayName": "\uae40\ud604\uc6b0", "photoUrl": "", "userId": "06560543018646300359"}}
# 딥러닝을 구동하는 데 필요한 케라스 함수를 불러옵니다.
# 필요한 라이브러리를 불러옵니다.
import pandas as pd
import tensorflow as tf
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
gpus = tf.config.experimental.list_physical_devices('GPU')
if gpus:
try:
# Currently, memory growth needs to be the same across GPUs
for gpu in gpus:
tf.config.experimental.set_memory_growth(gpu, True)
logical_gpus = tf.config.experimental.list_logical_devices('GPU')
print(len(gpus), "Physical GPUs,", len(logical_gpus), "Logical GPUs")
except RuntimeError as e:
# Memory growth must be set before GPUs have been initialized
print(e)
# + colab={"base_uri": "https://localhost:8080/", "height": 460} id="V0IXVU84zCql" executionInfo={"status": "error", "timestamp": 1629215282626, "user_tz": -540, "elapsed": 479, "user": {"displayName": "\uae40\ud604\uc6b0", "photoUrl": "", "userId": "06560543018646300359"}} outputId="59dad213-744a-423f-9088-6cb518b7891f"
# 준비된 수술 환자 데이터를 불러들입니다.
Data_set = pd.read_csv("../dataset/ThoraricSurgery.csv")
data_vale=Data_set.values
# 환자의 기록과 수술 결과를 X와 Y로 구분하여 저장합니다.
X = data_vale[:,0:17]
Y = data_vale[:,17]
# + colab={"base_uri": "https://localhost:8080/", "height": 223} id="RRm80LL-zFGT" executionInfo={"status": "error", "timestamp": 1629215300173, "user_tz": -540, "elapsed": 256, "user": {"displayName": "\uae40\ud604\uc6b0", "photoUrl": "", "userId": "06560543018646300359"}} outputId="e39e7103-a91a-46fd-efa4-224450bf5d1a"
# 전체 데이터에서 학습 데이터와 테스트 데이터(0.2)로 구분
X_train1, X_test, Y_train1, Y_test = train_test_split(X, Y, test_size=0.2)
X_train, X_valid, Y_train, Y_valid = train_test_split(X_train1, Y_train1, test_size=0.3)
# + colab={"base_uri": "https://localhost:8080/", "height": 525} id="6TjtDHjbzJeS" executionInfo={"status": "error", "timestamp": 1629215339821, "user_tz": -540, "elapsed": 701, "user": {"displayName": "\uae40\ud604\uc6b0", "photoUrl": "", "userId": "06560543018646300359"}} outputId="741d91a7-a4df-464b-fd77-a72d05cf16ad"
# 딥러닝 구조를 결정합니다(모델을 설정하고 실행하는 부분입니다).
input_Layer = tf.keras.layers.Input(shape=(17,))
x = tf.keras.layers.Dense(10, activation='sigmoid',kernel_initializer=tf.keras.initializers.he_normal())(input_Layer) ## he 초기화 방법
x = tf.keras.layers.Dense(10, activation='sigmoid',kernel_initializer=tf.keras.initializers.glorot_uniform())(x) ## xavier 초기화 방법
Out_Layer= tf.keras.layers.Dense(1, activation='sigmoid')(x)
model = tf.keras.models.Model(inputs=[input_Layer], outputs=[Out_Layer])
model.summary()
# 딥러닝을 실행합니다.
loss=tf.keras.losses.binary_crossentropy
optimizer = tf.keras.optimizers.SGD(learning_rate=0.01)
metrics=tf.keras.metrics.binary_accuracy
model.compile(loss=loss, optimizer=optimizer, metrics=[metrics])
result=model.fit(X_train, Y_train, epochs=100, batch_size=10, validation_data=(X_valid,Y_valid))
print(result.history.keys())
# + id="smzZi6UDzTC1"
### result에서 loss와 val_loss의 key를 가지는 값들만 추출
loss = result.history['loss']
val_loss = result.history['val_loss']
### loss와 val_loss를 그래프화
epochs = range(1, len(loss) + 1)
plt.subplot(211) ## 2x1 개의 그래프 중에 1번째
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
### result에서 binary_accuracy와 val_binary_accuracy key를 가지는 값들만 추출
rmse = result.history['binary_accuracy']
val_rmse = result.history['val_binary_accuracy']
epochs = range(1, len(rmse) + 1)
### binary_accuracy와 val_binary_accuracy key를 그래프화
plt.subplot(212) ## 2x1 개의 그래프 중에 2번째
plt.plot(epochs, rmse, 'ro', label='Training binary_accuracy')
plt.plot(epochs, val_rmse, 'r', label='Validation binary_accuracy')
plt.title('Training and validation binary_accuracy')
plt.xlabel('Epochs')
plt.ylabel('binary_accuracy')
plt.legend()
# 결과를 출력합니다.
print("-----")
print(model.evaluate(X_test, Y_test))
print("\n Accuracy: %.4f" % (model.evaluate(X, Y)[1]))
plt.show()
| tensorflow/day3/practice/P_03_07_ThoraricSurgery_prediction_weight_init.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # JIT Exercise
# Use `jit` (either in function or decorator form) to speed up the Mandelbrot code below, then time and compare the results
# +
from numba import jit
import numpy
from matplotlib import pyplot, cm
# %matplotlib inlin
# %load_ext line_profiler
# -
@jit(nopython=True)
def mandel(x, y, max_iters):
i = 0
c = complex(x, y)
z = 0.0j
for i in range(max_iters):
z = z * z + c
if (z.real * z.real + z.imag * z.imag) >= 4:
return i
return 255
@jit(nopython=True)
def create_fractal(min_x, max_x, min_y, max_y, image, iters):
height = image.shape[0]
width = image.shape[1]
pixel_size_x = (max_x - min_x) / width
pixel_size_y = (max_y - min_y) / height
for x in range(width):
real = min_x + x * pixel_size_x
for y in range(height):
imag = min_y + y * pixel_size_y
color = mandel(real, imag, iters)
image[y, x] = color
return image
# %%time
image = numpy.zeros((500 * 2, 750 * 2), dtype=numpy.uint8)
image = create_fractal(-2.0, 1.0, -1.0, 1.0, image, 20)
# %%time
pyplot.figure(figsize=(10,8))
pyplot.imshow(image, cmap=cm.viridis)
pyplot.colorbar();
| exercices/JIT.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Homework and bake-off: Word relatedness
__author__ = "<NAME>"
__version__ = "CS224u, Stanford, Spring 2021"
# ## Contents
#
# 1. [Overview](#Overview)
# 1. [Set-up](#Set-up)
# 1. [Development dataset](#Development-dataset)
# 1. [Vocabulary](#Vocabulary)
# 1. [Score distribution](#Score-distribution)
# 1. [Repeated pairs](#Repeated-pairs)
# 1. [Evaluation](#Evaluation)
# 1. [Error analysis](#Error-analysis)
# 1. [Homework questions](#Homework-questions)
# 1. [PPMI as a baseline [0.5 points]](#PPMI-as-a-baseline-[0.5-points])
# 1. [Gigaword with LSA at different dimensions [0.5 points]](#Gigaword-with-LSA-at-different-dimensions-[0.5-points])
# 1. [t-test reweighting [2 points]](#t-test-reweighting-[2-points])
# 1. [Pooled BERT representations [1 point]](#Pooled-BERT-representations-[1-point])
# 1. [Learned distance functions [2 points]](#Learned-distance-functions-[2-points])
# 1. [Your original system [3 points]](#Your-original-system-[3-points])
# 1. [Bake-off [1 point]](#Bake-off-[1-point])
# 1. [Submission Instruction](#Submission-Instruction)
# ## Overview
#
# Word similarity and relatedness datasets have long been used to evaluate distributed representations. This notebook provides code for conducting such analyses with a new word relatedness datasets. It consists of word pairs, each with an associated human-annotated relatedness score.
#
# The evaluation metric for each dataset is the [Spearman correlation coefficient $\rho$](https://en.wikipedia.org/wiki/Spearman%27s_rank_correlation_coefficient) between the annotated scores and your distances, as is standard in the literature.
#
# This homework ([questions at the bottom of this notebook](#Homework-questions)) asks you to write code that uses the count matrices in `data/vsmdata` to create and evaluate some baseline models. The final question asks you to create your own original system for this task, using any data you wish. This accounts for 9 of the 10 points for this assignment.
#
# For the associated bake-off, we will distribute a new dataset, and you will evaluate your original system (no additional training or tuning allowed!) on that datasets and submit your predictions. Systems that enter will receive the additional homework point, and systems that achieve the top score will receive an additional 0.5 points.
# ## Set-up
# +
from collections import defaultdict
import csv
import itertools
import numpy as np
import os
import pandas as pd
import random
from scipy.stats import spearmanr
import vsm
import utils
# -
utils.fix_random_seeds()
# +
VSM_HOME = os.path.join('data', 'vsmdata')
DATA_HOME = os.path.join('data', 'wordrelatedness')
# -
# ## Development dataset
# You can use development dataset freely, since our bake-off evalutions involve a new test set.
dev_df = pd.read_csv(
os.path.join(DATA_HOME, "cs224u-wordrelatedness-dev.csv"))
# The dataset consists of word pairs with scores:
dev_df.head()
# This gives the number of word pairs in the data:
dev_df.shape[0]
# The test set will contain 1500 word pairs with scores of the same type. No word pair in the development set appears in the test set, but some of the individual words are repeated in the test set.
# ### Vocabulary
# The full vocabulary in the dataframe can be extracted as follows:
dev_vocab = set(dev_df.word1.values) | set(dev_df.word2.values)
len(dev_vocab)
# The vocabulary for the bake-off test is different – it is partly overlapping with the above. If you want to be sure ahead of time that your system has a representation for every word in the dev and test sets, then you can check against the vocabularies of any of the VSMs in `data/vsmdata` (which all have the same vocabulary). For example:
# +
task_index = pd.read_csv(
os.path.join(VSM_HOME, 'yelp_window5-scaled.csv.gz'),
usecols=[0], index_col=0)
full_task_vocab = list(task_index.index)
# -
len(full_task_vocab)
# If you can process every one of those words, then you are all set. Alternatively, you can wait to see the test set and make system adjustments to ensure that you can process all those words. This is fine as long as you are not tuning your predictions.
# ### Score distribution
# All the scores fall in $[0, 1]$, and the dataset skews towards words with low scores, meaning low relatedness:
ax = dev_df.plot.hist().set_xlabel("Relatedness score")
# ### Repeated pairs
# The development data has some word pairs with multiple distinct scores in it. Here we create a `pd.Series` that contains these word pairs:
# +
repeats = dev_df.groupby(['word1', 'word2']).apply(lambda x: x.score.var())
repeats = repeats[repeats > 0].sort_values(ascending=False)
repeats.name = 'score variance'
# -
repeats.shape[0]
# The `pd.Series` is sorted with the highest variance items at the top:
repeats.head()
# Since this is development data, it is up to you how you want to handle these repeats. The test set has no repeated pairs in it.
# ## Evaluation
# Our evaluation function is `vsm.word_relatedness_evaluation`. Its arguments:
#
# 1. A relatedness dataset `pd.DataFrame` – e.g., `dev_df` as given above.
# 1. A VSM `pd.DataFrame` – e.g., `giga5` or some transformation thereof, or a GloVe embedding space, or something you have created on your own. The function checks that you can supply a representation for every word in `dev_df` and raises an exception if you can't.
# 1. Optionally a `distfunc` argument, which defaults to `vsm.cosine`.
#
# The function returns a tuple:
#
# 1. A copy of `dev_df` with a new column giving your predictions.
# 1. The Spearman $\rho$ value (our primary score).
#
# Important note: Internally, `vsm.word_relatedness_evaluation` uses `-distfunc(x1, x2)` as its score, where `x1` and `x2` are vector representations of words. This is because the scores in our data are _positive_ relatedness scores, whereas we are assuming that `distfunc` is a _distance_ function.
#
# Here's a simple illustration using one of our count matrices:
count_df = pd.read_csv(
os.path.join(VSM_HOME, "giga_window5-scaled.csv.gz"), index_col=0)
count_pred_df, count_rho = vsm.word_relatedness_evaluation(dev_df, count_df)
count_rho
count_pred_df.head()
# It's instructive to compare this against a truly random system, which we can create by simply having a custom distance function that returns a random number in [0, 1] for each example, making no use of the VSM itself:
def random_scorer(x1, x2):
"""`x1` and `x2` are vectors, to conform to the requirements
of `vsm.word_relatedness_evaluation`, but this function just
returns a random number in [0, 1]."""
return random.random()
# +
random_pred_df, random_rho = vsm.word_relatedness_evaluation(
dev_df, count_df, distfunc=random_scorer)
random_rho
# -
# This is a truly baseline system!
# ## Error analysis
#
# For error analysis, we can look at the words with the largest delta between the gold score and the distance value in our VSM. We do these comparisons based on ranks, just as with our primary metric (Spearman $\rho$), and we normalize both rankings so that they have a comparable number of levels.
# +
def error_analysis(pred_df):
pred_df = pred_df.copy()
pred_df['relatedness_rank'] = _normalized_ranking(pred_df.prediction)
pred_df['score_rank'] = _normalized_ranking(pred_df.score)
pred_df['error'] = abs(pred_df['relatedness_rank'] - pred_df['score_rank'])
return pred_df.sort_values('error')
def _normalized_ranking(series):
ranks = series.rank(method='dense')
return ranks / ranks.sum()
# -
# Best predictions:
error_analysis(count_pred_df).head()
# Worst predictions:
error_analysis(count_pred_df).tail()
# ## Homework questions
#
# Please embed your homework responses in this notebook, and do not delete any cells from the notebook. (You are free to add as many cells as you like as part of your responses.)
# ### PPMI as a baseline [0.5 points]
# The insight behind PPMI is a recurring theme in word representation learning, so it is a natural baseline for our task. This question asks you to write code for conducting such experiments.
#
# Your task: write a function called `run_giga_ppmi_baseline` that does the following:
#
# 1. Reads the Gigaword count matrix with a window of 20 and a flat scaling function into a `pd.DataFrame`, as is done in the VSM notebooks. The file is `data/vsmdata/giga_window20-flat.csv.gz`, and the VSM notebooks provide examples of the needed code.
# 1. Reweights this count matrix with PPMI.
# 1. Evaluates this reweighted matrix using `vsm.word_relatedness_evaluation` on `dev_df` as defined above, with `distfunc` set to the default of `vsm.cosine`.
# 1. Returns the return value of this call to `vsm.word_relatedness_evaluation`.
#
# The goal of this question is to help you get more familiar with the code in `vsm` and the function `vsm.word_relatedness_evaluation`.
#
# The function `test_run_giga_ppmi_baseline` can be used to test that you've implemented this specification correctly.
# +
def run_giga_ppmi_baseline():
pass
##### YOUR CODE HERE
# -
def test_run_giga_ppmi_baseline(func):
"""`func` should be `run_giga_ppmi_baseline"""
pred_df, rho = func()
rho = round(rho, 3)
expected = 0.586
assert rho == expected, \
"Expected rho of {}; got {}".format(expected, rho)
if 'IS_GRADESCOPE_ENV' not in os.environ:
test_run_giga_ppmi_baseline(run_giga_ppmi_baseline)
# ### Gigaword with LSA at different dimensions [0.5 points]
# We might expect PPMI and LSA to form a solid pipeline that combines the strengths of PPMI with those of dimensionality reduction. However, LSA has a hyper-parameter $k$ – the dimensionality of the final representations – that will impact performance. This problem asks you to create code that will help you explore this approach.
#
# Your task: write a wrapper function `run_ppmi_lsa_pipeline` that does the following:
#
# 1. Takes as input a count `pd.DataFrame` and an LSA parameter `k`.
# 1. Reweights the count matrix with PPMI.
# 1. Applies LSA with dimensionality `k`.
# 1. Evaluates this reweighted matrix using `vsm.word_relatedness_evaluation` with `dev_df` as defined above. The return value of `run_ppmi_lsa_pipeline` should be the return value of this call to `vsm.word_relatedness_evaluation`.
#
# The goal of this question is to help you get a feel for how LSA can contribute to this problem.
#
# The function `test_run_ppmi_lsa_pipeline` will test your function on the count matrix in `data/vsmdata/giga_window20-flat.csv.gz`.
# +
def run_ppmi_lsa_pipeline(count_df, k):
pass
##### YOUR CODE HERE
# -
def test_run_ppmi_lsa_pipeline(func):
"""`func` should be `run_ppmi_lsa_pipeline`"""
giga20 = pd.read_csv(
os.path.join(VSM_HOME, "giga_window20-flat.csv.gz"), index_col=0)
pred_df, rho = func(giga20, k=10)
rho = round(rho, 3)
expected = 0.545
assert rho == expected,\
"Expected rho of {}; got {}".format(expected, rho)
if 'IS_GRADESCOPE_ENV' not in os.environ:
test_run_ppmi_lsa_pipeline(run_ppmi_lsa_pipeline)
# ### t-test reweighting [2 points]
# The t-test statistic can be thought of as a reweighting scheme. For a count matrix $X$, row index $i$, and column index $j$:
#
# $$\textbf{ttest}(X, i, j) =
# \frac{
# P(X, i, j) - \big(P(X, i, *)P(X, *, j)\big)
# }{
# \sqrt{(P(X, i, *)P(X, *, j))}
# }$$
#
# where $P(X, i, j)$ is $X_{ij}$ divided by the total values in $X$, $P(X, i, *)$ is the sum of the values in row $i$ of $X$ divided by the total values in $X$, and $P(X, *, j)$ is the sum of the values in column $j$ of $X$ divided by the total values in $X$.
#
# Your task: implement this reweighting scheme. You can use `test_ttest_implementation` below to check that your implementation is correct. You do not need to use this for any evaluations, though we hope you will be curious enough to do so!
# +
def ttest(df):
pass
##### YOUR CODE HERE
# -
def test_ttest_implementation(func):
"""`func` should be `ttest`"""
X = pd.DataFrame([
[1., 4., 3., 0.],
[2., 43., 7., 12.],
[5., 6., 19., 0.],
[1., 11., 1., 4.]])
actual = np.array([
[ 0.04655, -0.01337, 0.06346, -0.09507],
[-0.11835, 0.13406, -0.20846, 0.10609],
[ 0.16621, -0.23129, 0.38123, -0.18411],
[-0.0231 , 0.0563 , -0.14549, 0.10394]])
predicted = func(X)
assert np.array_equal(predicted.round(5), actual), \
"Your ttest result is\n{}".format(predicted.round(5))
if 'IS_GRADESCOPE_ENV' not in os.environ:
test_ttest_implementation(ttest)
# ### Pooled BERT representations [1 point]
# The notebook [vsm_04_contextualreps.ipynb](vsm_04_contextualreps.ipynb) explores methods for deriving static vector representations of words from the contextual representations given by models like BERT and RoBERTa. The methods are due to [Bommasani et al. 2020](https://www.aclweb.org/anthology/2020.acl-main.431). The simplest of these methods involves processing the words as independent texts and pooling the sub-word representations that result, using a function like mean or max.
#
# Your task: write a function `evaluate_pooled_bert` that will enable exploration of this approach. The function should do the following:
#
# 1. Take as its arguments (a) a word relatedness `pd.DataFrame` `rel_df` (e.g., `dev_df`), (b) a `layer` index (see below), and (c) a `pool_func` value (see below).
# 1. Set up a BERT tokenizer and BERT model based on `'bert-base-uncased'`.
# 1. Use `vsm.create_subword_pooling_vsm` to create a VSM (a `pd.DataFrame`) with the user's values for `layer` and `pool_func`.
# 1. Return the return value of `vsm.word_relatedness_evaluation` using this new VSM, evaluated on `rel_df` with `distfunc` set to its default value.
#
# The function `vsm.create_subword_pooling_vsm` does the heavy-lifting. Your task is really just to put these pieces together. The result will be the start of a flexible framework for seeing how these methods do on our task.
#
# The function `test_evaluate_pooled_bert` can help you obtain the design we are seeking.
# +
from transformers import BertModel, BertTokenizer
def evaluate_pooled_bert(rel_df, layer, pool_func):
bert_weights_name = 'bert-base-uncased'
# Initialize a BERT tokenizer and BERT model based on
# `bert_weights_name`:
##### YOUR CODE HERE
# Get the vocabulary from `rel_df`:
##### YOUR CODE HERE
# Use `vsm.create_subword_pooling_vsm` with the user's arguments:
##### YOUR CODE HERE
# Return the results of the relatedness evalution:
##### YOUR CODE HERE
# -
def test_evaluate_pooled_bert(func):
import torch
rel_df = pd.DataFrame([
{'word1': 'porcupine', 'word2': 'capybara', 'score': 0.6},
{'word1': 'antelope', 'word2': 'springbok', 'score': 0.5},
{'word1': 'llama', 'word2': 'camel', 'score': 0.4},
{'word1': 'movie', 'word2': 'play', 'score': 0.3}])
layer = 2
pool_func = vsm.max_pooling
pred_df, rho = evaluate_pooled_bert(rel_df, layer, pool_func)
rho = round(rho, 2)
expected_rho = 0.40
assert rho == expected_rho, \
"Expected rho={}; got rho={}".format(expected_rho, rho)
if 'IS_GRADESCOPE_ENV' not in os.environ:
test_evaluate_pooled_bert(evaluate_pooled_bert)
# ### Learned distance functions [2 points]
# The presentation thus far leads one to assume that the `distfunc` argument used in the experiments will be a standard vector distance function like `vsm.cosine` or `vsm.euclidean`. However, the framework itself simply requires that this function map two fixed-dimensional vectors to a real number. This opens up a world of possibilities. This question asks you to dip a toe in these waters.
#
# Your task: write a function `run_knn_score_model` for models in this class. The function should:
#
# 1. Take as its arguments (a) a VSM dataframe `vsm_df`, (b) a relatedness dataset (e.g., `dev_df`), and (c) a `test_size` value between 0.0 and 1.0 that can be passed directly to `train_test_split` (see below).
# 1. Create a feature matrix `X`: each word pair in `dev_df` should be represented by the concatenation of the vectors for word1 and word2 from `vsm_df`.
# 1. Create a score vector `y`, which is just the `score` column in `dev_df`.
# 1. Split the dataset `(X, y)` into train and test portions using [sklearn.model_selection.train_test_split](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.train_test_split.html).
# 1. Train an [sklearn.neighbors.KNeighborsRegressor](https://scikit-learn.org/stable/modules/generated/sklearn.neighbors.KNeighborsRegressor.html#sklearn.neighbors.KNeighborsRegressor) model on the train split from step 4, with default hyperparameters.
# 1. Return the value of the `score` method of the trained `KNeighborsRegressor` model on the test split from step 4.
#
# The functions `test_knn_feature_matrix` and `knn_represent` will help you test the crucial representational aspects of this.
#
# Note: if you decide to apply this approach to our task as part of an original system, recall that `vsm.create_subword_pooling_vsm` returns `-d` where `d` is the value computed by `distfunc`, since it assumes that `distfunc` is a distance value of some kind rather than a relatedness/similarity value. Since most regression models will return positive scores for positive associations, you will probably want to undo this by having your `distfunc` return the negative of its value.
# +
from sklearn.model_selection import train_test_split
from sklearn.neighbors import KNeighborsRegressor
def run_knn_score_model(vsm_df, dev_df, test_size=0.20):
pass
# Complete `knn_feature_matrix` for this step.
##### YOUR CODE HERE
# Get the values of the 'score' column in `dev_df`
# and store them in a list or array `y`.
##### YOUR CODE HERE
# Use `train_test_split` to split (X, y) into train and
# test protions, with `test_size` as the test size.
##### YOUR CODE HERE
# Instantiate a `KNeighborsRegressor` with default arguments:
##### YOUR CODE HERE
# Fit the model on the training data:
##### YOUR CODE HERE
# Return the value of `score` for your model on the test split
# you created above:
##### YOUR CODE HERE
def knn_feature_matrix(vsm_df, rel_df):
pass
# Complete `knn_represent` and use it to create a feature
# matrix `np.array`:
##### YOUR CODE HERE
def knn_represent(word1, word2, vsm_df):
pass
# Use `vsm_df` to get vectors for `word1` and `word2`
# and concatenate them into a single vector:
##### YOUR CODE HERE
# +
def test_knn_feature_matrix(func):
rel_df = pd.DataFrame([
{'word1': 'w1', 'word2': 'w2', 'score': 0.1},
{'word1': 'w1', 'word2': 'w3', 'score': 0.2}])
vsm_df = pd.DataFrame([
[1, 2, 3.],
[4, 5, 6.],
[7, 8, 9.]], index=['w1', 'w2', 'w3'])
expected = np.array([
[1, 2, 3, 4, 5, 6.],
[1, 2, 3, 7, 8, 9.]])
result = func(vsm_df, rel_df)
assert np.array_equal(result, expected), \
"Your `knn_feature_matrix` returns: {}\nWe expect: {}".format(
result, expected)
def test_knn_represent(func):
vsm_df = pd.DataFrame([
[1, 2, 3.],
[4, 5, 6.],
[7, 8, 9.]], index=['w1', 'w2', 'w3'])
result = func('w1', 'w3', vsm_df)
expected = np.array([1, 2, 3, 7, 8, 9.])
assert np.array_equal(result, expected), \
"Your `knn_represent` returns: {}\nWe expect: {}".format(
result, expected)
# -
if 'IS_GRADESCOPE_ENV' not in os.environ:
test_knn_represent(knn_represent)
test_knn_feature_matrix(knn_feature_matrix)
# ### Your original system [3 points]
#
# This question asks you to design your own model. You can of course include steps made above (ideally, the above questions informed your system design!), but your model should not be literally identical to any of the above models. Other ideas: retrofitting, autoencoders, GloVe, subword modeling, ...
#
# Requirements:
#
# 1. Your system must work with `vsm.word_relatedness_evaluation`. You are free to specify the VSM and the value of `distfunc`.
#
# 1. Your code must be self-contained, so that we can work with your model directly in your homework submission notebook. If your model depends on external data or other resources, please submit a ZIP archive containing these resources along with your submission.
#
# In the cell below, please provide a brief technical description of your original system, so that the teaching team can gain an understanding of what it does. This will help us to understand your code and analyze all the submissions to identify patterns and strategies. We also ask that you report the best score your system got during development, just to help us understand how systems performed overall.
# +
# PLEASE MAKE SURE TO INCLUDE THE FOLLOWING BETWEEN THE START AND STOP COMMENTS:
# 1) Textual description of your system.
# 2) The code for your original system.
# 3) The score achieved by your system in place of MY_NUMBER.
# With no other changes to that line.
# You should report your score as a decimal value <=1.0
# PLEASE MAKE SURE NOT TO DELETE OR EDIT THE START AND STOP COMMENTS
# NOTE: MODULES, CODE AND DATASETS REQUIRED FOR YOUR ORIGINAL SYSTEM
# SHOULD BE ADDED BELOW THE 'IS_GRADESCOPE_ENV' CHECK CONDITION. DOING
# SO ABOVE THE CHECK MAY CAUSE THE AUTOGRADER TO FAIL.
# START COMMENT: Enter your system description in this cell.
# My peak score was: MY_NUMBER
if 'IS_GRADESCOPE_ENV' not in os.environ:
pass
# STOP COMMENT: Please do not remove this comment.
# -
# ## Bake-off [1 point]
#
# For the bake-off, you simply need to evaluate your original system on the file
#
# `wordrelatedness/cs224u-wordrelatedness-test-unlabeled.csv`
#
# This contains only word pairs (no scores), so `vsm.word_relatedness_evaluation` will simply make predictions without doing any scoring. Use that function to make predictions with your original system, store the resulting `pred_df` to a file, and then upload the file as your bake-off submission.
#
# The following function should be used to conduct this evaluation:
def create_bakeoff_submission(
vsm_df,
distfunc,
output_filename="cs224u-wordrelatedness-bakeoff-entry.csv"):
test_df = pd.read_csv(
os.path.join(DATA_HOME, "cs224u-wordrelatedness-test-unlabeled.csv"))
pred_df, _ = vsm.word_relatedness_evaluation(test_df, vsm_df, distfunc=distfunc)
pred_df.to_csv(output_filename)
# For example, if `count_df` were the VSM for my system, and I wanted my distance function to be `vsm.euclidean`, I would do
create_bakeoff_submission(count_df, vsm.euclidean)
# This creates a file `cs224u-wordrelatedness-bakeoff-entry.csv` in the current directory. That file should be uploaded as-is. Please do not change its name.
#
# Only one upload per team is permitted, and you should do no tuning of your system based on what you see in `pred_df` – you should not study that file in anyway, beyond perhaps checking that it contains what you expected it to contain. The upload function will do some additional checking to ensure that your file is well-formed.
#
# People who enter will receive the additional homework point, and people whose systems achieve the top score will receive an additional 0.5 points. We will test the top-performing systems ourselves, and only systems for which we can reproduce the reported results will win the extra 0.5 points.
#
# Late entries will be accepted, but they cannot earn the extra 0.5 points.
# ## Submission Instruction
#
# Submit the following files to gradescope submission
#
# - Please do not change the file name as described below
# - `hw_wordrelatedness.ipynb` (this notebook)
# - `cs224u-wordrelatedness-bakeoff-entry.csv` (bake-off output)
#
| hw_wordrelatedness.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import pymc3 as pm
import numpy as np
import scipy.stats as stats
import matplotlib.pyplot as plt
import seaborn as sns
import matplotlib
palette = 'muted'
sns.set_palette(palette); sns.set_color_codes(palette)
# # Overfitting vs underfitting
# +
x = np.array([4.,5.,6.,9.,12, 14.])
y = np.array([4.2, 6., 6., 9., 10, 10.])
order = [0, 1, 2, 5]
plt.plot(x, y, 'o')
for i in order:
x_n = np.linspace(x.min(), x.max(), 100)
coeffs = np.polyfit(x, y, deg=i)
ffit = np.polyval(coeffs, x_n)
p = np.poly1d(coeffs)
yhat = p(x)
ybar = np.mean(y)
ssreg = np.sum((yhat-ybar)**2)
sstot = np.sum((y - ybar)**2)
r2 = ssreg / sstot
plt.plot(x_n, ffit, label='order {}, $R^2$= {:.2f}'.format(i, r2))
plt.legend(loc=2, fontsize=14);
plt.xlabel('$x$', fontsize=14);
plt.ylabel('$y$', fontsize=14, rotation=0);
fig = matplotlib.pyplot.gcf()
fig.set_size_inches(16,10)
# +
plt.figure(figsize=(8, 6))
x_values = np.linspace(-10, 10, 300)
for df in [1, 2, 5, 15]:
distri = stats.laplace(scale=df)
x_pdf = distri.pdf(x_values)
plt.plot(x_values, x_pdf, label='Laplace with scale ($b$) = {}'.format(df))
x_pdf = stats.norm.pdf(x_values)
plt.plot(x_values, x_pdf, label='Gaussian')
plt.xlabel('x')
plt.ylabel('p(x)', rotation=0)
plt.legend(loc=0, fontsize=14)
plt.xlim(-7, 7);
fig = matplotlib.pyplot.gcf()
fig.set_size_inches(16,10)
# -
x_1 = np.array([ 10., 8., 13., 9., 11., 14., 6., 4., 12., 7., 5.])
y_1 = np.array([ 8.04, 6.95, 7.58, 8.81, 8.33, 9.96, 7.24, 4.26,
10.84, 4.82, 5.68])
np.random.seed(1)
real_alpha = 4.25
real_beta = [8.7, -1.2]
data_size = 20
noise = np.random.normal(0, 2, size=data_size)
x_1 = np.linspace(0, 5, data_size)
y_1 = real_alpha + real_beta[0] * x_1 + real_beta[1] * x_1**2 + noise
order = 2#5
x_1p = np.vstack([x_1**i for i in range(1, order+1)])
x_1s = (x_1p - x_1p.mean(axis=1, keepdims=True))/x_1p.std(axis=1, keepdims=True)
y_1s = (y_1 - y_1.mean())/y_1.std()
plt.scatter(x_1s[0], y_1s);
plt.xlabel('$x$', fontsize=16);
plt.ylabel('$y$', fontsize=16, rotation=0);
# #### model_l: Not including polynomial term
with pm.Model() as model_l:
alpha = pm.Normal('alpha', mu=0, sd=10)
beta = pm.Normal('beta', mu=0, sd=1)
epsilon = pm.HalfCauchy('epsilon', 5)
mu = alpha + beta * x_1s[0]
y_l = pm.Normal('y_l', mu=mu, sd=epsilon, observed=y_1s)
trace_l = pm.sample(2100,chains=1,njobs=1)
chain_l = trace_l[100:]
pm.traceplot(chain_l);
pm.summary(chain_l)
# #### model_p: Iincluding polynomial term
with pm.Model() as model_p:
alpha = pm.Normal('alpha', mu=0, sd=10)
beta = pm.Normal('beta', mu=0, sd=1, shape=x_1s.shape[0])
epsilon = pm.HalfCauchy('epsilon', 5)
mu = alpha + pm.math.dot(beta, x_1s)
y_l = pm.Normal('y_l', mu=mu, sd=epsilon, observed=y_1s)
trace_p = pm.sample(2100,chains=1,njobs=1)
chain_p = trace_p[100:]
pm.traceplot(chain_p);
pm.summary(chain_p)
# +
alpha_l_post = chain_l['alpha'].mean()
betas_l_post = chain_l['beta'].mean(axis=0)
idx = np.argsort(x_1s[0])
y_l_post = alpha_l_post + betas_l_post * x_1s[0]
plt.plot(x_1s[0][idx], y_l_post[idx], label='Linear')
alpha_p_post = chain_p['alpha'].mean()
betas_p_post = chain_p['beta'].mean(axis=0)
y_p_post = alpha_p_post + np.dot(betas_p_post, x_1s)
plt.plot(x_1s[0][idx], y_p_post[idx], label='Pol order {}'.format(order))
plt.scatter(x_1s[0], y_1s)
plt.xlabel('$x$', fontsize=16)
plt.ylabel('$y$', fontsize=16, rotation=0);
plt.legend();
# -
# # Information criteria
model_p.name='Polynomial'
model_l.name='Linear'
dic_l = pm.compare({model_p: trace_p, model_l: trace_l})
dic_l
waic_l = pm.waic(trace=trace_l, model=model_l)
waic_l
waic_p = pm.waic(trace=trace_p, model=model_p)
waic_p
loo_l = pm.loo(trace=trace_l, model=model_l)
loo_l
loo_p = pm.loo(trace=trace_p, model=model_p)
loo_p
# ### Lower is better
# +
plt.figure(figsize=(8, 4))
plt.subplot(121)
for idx, ic in enumerate((waic_l, waic_p)):
plt.errorbar(ic[0], idx, xerr=ic[1], fmt='bo')
plt.title('WAIC')
plt.yticks([0, 1], ['linear', 'quadratic'])
plt.ylim(-1, 2)
plt.subplot(122)
for idx, ic in enumerate((loo_l, loo_p)):
plt.errorbar(ic[0], idx, xerr=ic[1], fmt='go')
plt.title('LOO')
plt.yticks([0, 1], ['linear', 'quadratic'])
plt.ylim(-1, 2)
plt.tight_layout()
# -
# ## Posterior predictive checks
# +
plt.figure(figsize=(12,6))
plt.subplot(121)
plt.scatter(x_1s[0], y_1s, c='r');
plt.ylim(-3, 3)
plt.xlabel('x')
plt.ylabel('y', rotation=0)
plt.title('Linear')
for i in range(0, len(chain_l['alpha']), 50):
plt.scatter(x_1s[0], chain_l['alpha'][i] + chain_l['beta'][i]*x_1s[0], c='g',
edgecolors='g', alpha=0.5);
plt.plot(x_1s[0], chain_l['alpha'].mean() + chain_l['beta'].mean()*x_1s[0], c='g', alpha=1)
plt.subplot(122)
plt.scatter(x_1s[0], y_1s, c='r');
plt.ylim(-3, 3)
plt.xlabel('x')
plt.ylabel('y', rotation=0)
plt.title('Order {}'.format(order))
for i in range(0, len(chain_p['alpha']), 50):
plt.scatter(x_1s[0], chain_p['alpha'][i] + np.dot(chain_p['beta'][i], x_1s), c='g',
edgecolors='g', alpha=0.5)
idx = np.argsort(x_1)
plt.plot(x_1s[0][idx], alpha_p_post + np.dot(betas_p_post, x_1s)[idx], c='g', alpha=1);
plt.tight_layout()
# -
# # Bayes factors
coins = 30 # 300
heads = 9 # 90
y = np.repeat([0, 1], [coins-heads, heads])
print('These are the coin tosses we are modelling:\n {}'.format(y))
with pm.Model() as model_BF:
p = np.array([0.5, 0.5])
# model_index is a stochastic variable governed by the Categorical distribution
# returning 0 or 1 for each model respectively
model_index = pm.Categorical('model_index', p=p)
# there are two models with different priors
# one alpha=4,beta=8 and another alpha=8,beta=4
m_0 = (4, 8)
m_1 = (8, 4)
# m returns the alpha,betas based on whether model index ==0
# or not (`pm.math.eq(model_index, 0)`)
m = pm.math.switch(pm.math.eq(model_index, 0), m_0, m_1)
# prior on theta of the Bernouli
theta = pm.Beta('theta', m[0], m[1])
# likelihood, y-> 1 heads, 0 tails
y_pred = pm.Bernoulli('y', theta, observed=y)
trace_BF = pm.sample(5000,chains=1,njobs=1)
# # $BayesFactor=\frac{p(y|M_{0})}{p(y|M_{1})}=\frac{p(M_{0}|y) \times p(M_{1})}{p(M_{1}|y) \times p(M_{0})}$
chain_BF = trace_BF[500:]
pm.traceplot(chain_BF)
pM1 = chain_BF['model_index'].mean()
pM0 = 1 - pM1
print('Prior of selecting model 0 = {:.2f}'.format(p[0]))
print('Prior of selecting model 1 = {:.2f}'.format(p[1]))
print('Posterior mean of selecting model 0 = {:.2f}'.format(pM1))
print('Posterior mean of selecting model 1 = {:.2f}'.format(pM0))
print('Bayes factor = {:.2f}, thus model 0 is more likely'.format((pM0/pM1)*(p[1]/p[0])))
# ### Also model 0 $\alpha=4, \beta=8$ is more compatible with the observations of mostly tails and $\theta<0.5$
# ## Comparison of models using Information Criteria
with pm.Model() as model_BF_0:
theta = pm.Beta('theta', 4, 8)
y_pred = pm.Bernoulli('y', theta, observed=y)
trace_BF_0 = pm.sample(5000,chains=1,njobs=1)
chain_BF_0 = trace_BF_0[500:]
pm.traceplot(trace_BF_0);
with pm.Model() as model_BF_1:
theta = pm.Beta('theta', 8, 4)
y_pred = pm.Bernoulli('y', theta, observed=y)
trace_BF_1 = pm.sample(5000,chains=1,njobs=1)
chain_BF_1 = trace_BF_1[500:]
pm.traceplot(chain_BF_1);
waic_0 = pm.waic(chain_BF_0, model_BF_0)
waic_0
waic_1 = pm.waic(chain_BF_1, model_BF_1)
waic_1
loo_0 = pm.loo(chain_BF_0, model_BF_0)
loo_0
loo_1 = pm.loo(chain_BF_1, model_BF_1)
loo_1
# +
est = [((38.02, 4.17), (39.41, 2.04)), ((36.69, 3.96), (38.09, 1.94)),
((368.41, 13.40), (368.76, 12.48)) , ((366.61, 13.31), (366.87, 12.34))]
title = ['WAIC 30_9', 'LOO 30_9', 'WAIC 300_90', 'LOO 300_90']
for i in range(4):
plt.subplot(2,2,i+1)
for idx, ic in enumerate(est[i]):
plt.errorbar(ic[0], idx, xerr=ic[1], fmt='bo')
plt.title(title[i])
plt.yticks([0, 1], ['model_0', 'model_1'])
plt.ylim(-1, 2)
plt.tight_layout()
# -
# ### Bayes factors are sensitive to the selection of priors and this selection becomes less relevant as we increase the data. As you can see from using the Information Criteria there isn't much of a difference between the models, and these differences are going to become less and less as we increase the number of data. This sensitivity of Bayes factors to the selection of priors makes people not want to use them as much.
import sys, IPython, scipy, matplotlib, platform
print("This notebook was created on a %s computer running %s and using:\nPython %s\nIPython %s\nPyMC3 %s\nNumPy %s\nSciPy %s\nMatplotlib %s\nSeaborn %s" % (platform.machine(), ' '.join(platform.linux_distribution()[:2]), sys.version[:5], IPython.__version__, pm.__version__, np.__version__, scipy.__version__, matplotlib.__version__, sns.__version__))
| Chapter 6/Bayesian Data Analysis Chapter 6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="sXRKWNln4TDu" colab_type="text"
# # This notebook depicts spark streaming process, it listens to topic and persists events in mongodb atlas(hosted over aws).
#
#
# + id="Lo1G0ZoA37-v" colab_type="code" colab={}
from pyspark import SparkConf, SparkContext
import sys
import math
assert sys.version_info >= (3, 5) # make sure we have Python 3.5+
import re, datetime, uuid
from pyspark.sql import SQLContext, Row, SparkSession, functions, types
from pyspark.sql.types import StructType, StructField, StringType, FloatType, TimestampType
# + [markdown] id="kGxfRPlkm5Yh" colab_type="text"
# # Spark Streaming process - Feature is available from spark 2.4 version
#
# + [markdown] id="HaIjtmEqnMkL" colab_type="text"
# **Streaming process handling etl layer, it captures events from kafka topic and transform data so that it can be loaded into mongodb**
#
# **Mongo atlas is used here, which is hosted on AWS, dependencies provided with spark.jars.packages parameter**
# + id="xHli3HpeTnAA" colab_type="code" colab={}
def save_batch(df, epoch_id):
df.write.format("com.mongodb.spark.sql.DefaultSource").mode("append") \
.option("database",'iot_prediction') \
.option("collection", 'battery_1') \
.save()
pass
def main(topic, freq, database_name, collection_name, bootstrap_server):
#spark = SparkSession.builder.appName('Read_Stream').config("spark.jars.packages",'org.apache.spark:spark-sql-kafka-0-10_2.11:2.4.0,org.mongodb.spark:mongo-spark-connector_2.11:2.3.0').config('spark.mongodb.input.uri', 'mongodb+srv://falcon:vancouver@cmpt733-stzkw.mongodb.net/test?retryWrites=true').config('spark.mongodb.output.uri', 'mongodb+srv://falcon:vancouver@cmpt733-stzkw.mongodb.net/test?retryWrites=true').getOrCreate()
spark = SparkSession.builder.appName('Read_Stream') \
.config("spark.jars.packages",'org.apache.spark:spark-sql-kafka-0-10_2.11:2.4.0,org.mongodb.spark:mongo-spark-connector_2.11:2.3.0') \
.config('spark.mongodb.input.uri', 'mongodb+srv://falcon:vancouver@cmpt733-stzkw.mongodb.net/test?retryWrites=true') \
.config('spark.mongodb.output.uri', 'mongodb+srv://falcon:vancouver@cmpt733-stzkw.mongodb.net/test?retryWrites=true') \
.getOrCreate()
messages = spark.readStream.format('kafka') \
.option('kafka.bootstrap.servers', bootstrap_server) \
.option('subscribe', topic).load()
spark.sparkContext.setLogLevel('WARN')
values = messages.select(messages['value'].cast('string'))
split_val = functions.split(values['value'], ',')
#Values are specific to business requirement
values = values.withColumn('Langitude', split_val.getItem(0))
values = values.withColumn('Battery_Level', split_val.getItem(1))
values = values.withColumn('Latitude', split_val.getItem(2))
values = values.withColumn('Battery_Cycle_No', split_val.getItem(3))
values = values.withColumn('Location', split_val.getItem(4))
values = values.withColumn('u_id', split_val.getItem(5))
values = values.withColumn('Battery_Type', split_val.getItem(6))
values = values.withColumn('Battery_Status', split_val.getItem(7))
values = values.withColumn('User_Type', split_val.getItem(8))
values = values.withColumn('DateTime', split_val.getItem(9))
values = values.withColumn('full_name', split_val.getItem(10))
stream = values.writeStream.foreachBatch(save_batch).start()
stream.awaitTermination(freq)
# + [markdown] id="PziKCE7I4wFY" colab_type="text"
# **Below main method is used for invocation, all relevant information should be passed in this method.**
#
# + id="YB2WRGv2nvGN" colab_type="code" colab={}
if __name__ == "__main__":
topic = "battery-1"
termination_time = 6000
database_name = 'iot_prediction'
collection_name = 'battery_1'
bootstrap_server = '192.168.127.12:9092'
main(topic, termination_time, database_name, collection_name, bootstrap_server)
# + id="wrFgykMwTrAM" colab_type="code" colab={}
| 7_data_listener.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Priority Comparison
# Goal: During the questionnarie HH members were directly asked what was their first/second/third priority. We compare whether the first sector priority as obtained by our indexes is the same as the one directly reported by HH.
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import math
sns.set_style('whitegrid')
# %matplotlib inline
severity = pd.read_csv('../../data/processed/severity.csv')
#severity.drop('Unnamed', axis = 1)
severity.head()
severity.drop('Unnamed: 0', axis = 1, inplace = True)
severity.head()
# ##### Extract first n Priorities from Data
#Explore General properties of the severity data
severity.describe()
# +
#Find first, second and third priority from the data
temp = pd.Series
temp = severity.idxmax(axis=1)
compare = pd.DataFrame()
#def get_n_best(x,n):
# return sorted(x)[-n]
compare['Data First Priority']= temp
compare['Data Second Priority'] = severity.T.apply(lambda x: x.nlargest(2).idxmin())
compare['Data Third Priority'] = severity.T.apply(lambda x: x.nlargest(3).idxmin())
compare.head()
compare['Data First Priority'].value_counts()
# -
# ##### Import Data from the Questionnarie
#
##### Group Data in PiN
df = pd.read_excel('../../data/raw/reach_nga_msna_clean_dataset_final.xlsx',sheet_name='clean_hh_data')
# Remove respondents who refuse to take the interview
df = df[df['Consent'] == 'Yes, agrees to interview']
# ##### Group answers to 'What is your xth priority' in PiN
# +
def group(element):
if (element == 'Water') or (element == 'Latrines'):
return 'WASH'
elif(element == 'Education / Schools'):
return 'Education'
elif(element == 'Nutrition'): #Notice that this is in principle wrong. It's just for the purpose of comparison
return 'Food'
elif(element == 'Livelihoods / Income'):
return 'ERL'
elif(element == 'Shelter support'):
return 'Shelter/NFI'
elif(element == 'Safety and security'):
return 'Protection'
elif(element == 'Transportation/roads') or (element == 'Legal documentation') or (element == 'Communication') or (element == 'Psychosocial support services'):
return 'Other'
elif(element == 'Non-food items (blankets, jerry cans, buckets, etc)') or (element == 'Reunification with family') or (element == 'Energy/Fuel'):
return 'Other'
else:
return element
# +
compare['Answer First Priority'] = df['What is the first priority need for your household overall?'].apply(group)
compare['Answer Second Priority'] = df['What is the second priority need for your household overall?'].apply(group)
compare['Answer Third Priority'] = df['What is the third priority need for your household overall?'].apply(group)
compare['Data First Priority'] = compare['Data First Priority'].apply(group)
compare['Data Second Priority'] = compare['Data Second Priority'].apply(group)
compare['Data Third Priority'] = compare['Data Third Priority'].apply(group)
# Check that the categories are the same
print(np.unique(list(compare['Answer First Priority'])))
print(np.unique(list(compare['Data First Priority'])))
# -
# ##### First Priority Comparison
compare['Compare First'] = compare['Answer First Priority'] == compare['Data First Priority']
compare['Compare First'].sum()/len(compare['Compare First'])*100
#Only 29% of the time we are able to correctly understand the main need of the household
# +
plotdf = compare[['Answer First Priority','Data First Priority']]
sns.set(rc={'figure.figsize':(11.7,8.27)})
sns_plot = sns.countplot(x="variable", hue="value", data=pd.melt(plotdf))
#sns_plot.savefig('Comparison_First.png')
#plt.savefig(sns_plot)
sns_plot.figure.savefig('First_Priority.png', bbox_inches = 'tight',
pad_inches = 0)
#sns_plot = sns.pairplot(df, hue='species', size=2.5)
#sns_plot.savefig("Firs_Priority.png")
# -
# ##### Second Priority Comparison
compare['Compare Second'] = compare['Answer Second Priority'] == compare['Data Second Priority']
compare['Compare Second'].sum()/len(compare['Compare Second'])*100
#Only 16% of the time we are able to correctly understand the Second Priority of the household
plotdf = compare[['Answer Second Priority','Data Second Priority']]
sns.set(rc={'figure.figsize':(11.7,8.27)})
sns.countplot(x="variable", hue="value", data=pd.melt(plotdf))
# ##### Third Priority Comparison
compare['Compare Third'] = compare['Answer Third Priority'] == compare['Data Third Priority']
compare['Compare Third'].sum()/len(compare['Compare Third'])*100
#Only 17% of the time we are able to correctly understand the third need of the household
plotdf = compare[['Answer Third Priority','Data Third Priority']]
sns.set(rc={'figure.figsize':(11.7,8.27)})
sns.countplot(x="variable", hue="value", data=pd.melt(plotdf))
# ### Try different metrics from Classification Problems: Sensitivity is the one that gives more information
# +
# TRUE print((compare == sector)['Answer First Priority'].sum()) 6191
# FALSE print((compare != sector)['Answer First Priority'].sum()) 4187
# POSITIVE print((compare == sector)['Data First Priority'].sum()) 4020
# NEGATIVE print((compare != sector)['Data First Priority'].sum()) 6358
from sklearn.metrics import cohen_kappa_score
def compute_TP(sector):
TP = 0
TP = (((compare == sector)['Answer First Priority']*1 + (compare == sector)['Data First Priority']*1) >1).sum()
return TP
def compute_TN(sector):
TN = 0
TN = ((((compare == sector)['Answer First Priority']==False)*1 + ((compare == sector)['Data First Priority']==False)*1) >1).sum()
return TN
def compute_FP(sector):
FP = 0
FP = ((((compare == sector)['Answer First Priority']==False)*1 + (compare == sector)['Data First Priority']*1) >1).sum()
return FP
def compute_FN(sector):
FN = 0
FN = ((((compare == sector)['Answer First Priority']==True)*1 + ((compare == sector)['Data First Priority']==False)*1) >1).sum()
return FN
def compute_P(sector):
P = 0
P = (compare == sector)['Answer First Priority'].sum()
return P
def compute_N(sector):
N = 0
N = (compare != sector)['Answer First Priority'].sum()
return N
def compute_metric(sector):
TP = compute_TP(sector)
TN = compute_TN(sector)
FP = compute_FP(sector)
FN = compute_FN(sector)
P = compute_P(sector)
N = compute_N(sector)
Precision = TP/(TP+FP)
Recall = TP/(TP+FN)
Sensitivity = TP/(P)
Specificity = TN/N
MCC = (TP*TN - FP*FN)/(math.sqrt((TP+FP)*(TP+FN)*(TN+FP)*(TN+FN))) #Matthews correlation coefficient
CK = cohen_kappa_score((compare == sector)['Answer First Priority']*1, (compare == sector)['Data First Priority']*1)
F1 = 2*(Precision*Recall)/(Precision+Recall)
return Precision, Recall, Sensitivity,Specificity,MCC,CK, F1
# -
sectors = ['Food','ERL','WASH','Health','Shelter/NFI','Protection','Education']
l = compute_metric('Food')
print(l)
metric = []
pd.Series(l)
metric = pd.DataFrame(index='Precision Recall Sensitivity Specificity MCC CK F1'.split())
for i in sectors:
metric[i] = compute_metric(i)
metric
#The metrics which seems to give the highest information is the Sensitivity
# ### Plot confusion matrix
# +
#Notice that the results used in this confusion matrix are the Sensitivities for each sector
from sklearn.metrics import confusion_matrix
from sklearn.utils.multiclass import unique_labels
def plot_confusion_matrix(y_true, y_pred, classes,
normalize=False,
title=None,
cmap=plt.cm.Blues,
display=False):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
if not title:
if normalize:
title = 'Normalized confusion matrix'
else:
title = 'Confusion matrix, without normalization'
# Compute confusion matrix
cm = confusion_matrix(y_true, y_pred)
# Only use the labels that appear in the data
classes = classes[unique_labels(y_true, y_pred)]
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
if display:
print(cm)
fig, ax = plt.subplots()
im = ax.imshow(cm, interpolation='nearest', cmap=cmap)
ax.figure.colorbar(im, ax=ax)
# We want to show all ticks...
ax.set(xticks=np.arange(cm.shape[1]),
yticks=np.arange(cm.shape[0]),
# ... and label them with the respective list entries
xticklabels=classes, yticklabels=classes,
title=title,
ylabel='True label',
xlabel='Predicted label')
# Rotate the tick labels and set their alignment.
plt.setp(ax.get_xticklabels(), rotation=45, ha="right",
rotation_mode="anchor")
# Loop over data dimensions and create text annotations.
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i in range(cm.shape[0]):
for j in range(cm.shape[1]):
ax.text(j, i, format(cm[i, j], fmt),
ha="center", va="center",
color="white" if cm[i, j] > thresh else "black")
fig.tight_layout()
return ax
### Plot confusion matrix
# +
sector_names = list(severity.columns.values)
sector_names.append('Other')
sector_names = np.array(sector_names)
# Build y_test and transform to categorical vector
y_test = compare["Answer First Priority"].values
for i, item in enumerate(y_test):
if item not in sector_names:
y_test[i] = 'Other'
for i, c in enumerate(sector_names):
c_idx = np.where(y_test==c)
y_test[c_idx] = i
# Build y_pred and transform to categorical vector
y_pred = compare["Data First Priority"].values
for i, c in enumerate(sector_names):
c_idx = np.where(y_pred==c)
y_pred[c_idx] = i
y_test = list(y_test)
y_pred = list(y_pred)
# +
y_pred = compare["Data First Priority"].values
for i, c in enumerate(sector_names):
c_idx = np.where(y_pred==c)
y_pred[c_idx] = i
y_test = list(y_test)
y_pred = list(y_pred)
len(y_pred)
confusion_matrix(y_test, y_pred)
# +
# Plot non-normalized confusion matrix
plot_confusion_matrix(y_test, y_pred, classes=sector_names,
title='Confusion matrix, without normalization')
# Plot normalized confusion matrix
plot_confusion_matrix(y_test, y_pred, classes=sector_names, normalize=True,
title='Normalized confusion matrix')
plt.savefig('ConfusionMatrix.png', bbox_inches = 'tight',
pad_inches = 0)
plt.show()
# -
# ### Compute Sensitivity and Specificity Including answer to second and third priority
def compute_Sensitivity(sector):
S1 = 0
S2 = 0
S3 = 0
P = compute_P(sector)
S1 = compute_TP(sector)/P
S2 = compute_TP_IncludingSecond(sector)/P
S3 = compute_TP_IncludingSecondThird(sector)/P
return S1,S2, S3
def compute_Specificity(sector):
S1 = 0
S2 = 0
S3 = 0
N = compute_N(sector)
S1 = compute_TN(sector)/N
S2 = compute_TNIncludingSecond(sector)/N
S3 = compute_TNIncludingThird(sector)/N
return S1,S2, S3
def compute_metricExtended(sector):
P = 0
N = 0
S1 = 0
S2 = 0
S3 = 0
P = compute_P(sector)
N = compute_N(sector)
S1 = compute_TP(sector)/P
S2 = compute_TP_IncludingSecond(sector)/P
S3 = compute_TP_IncludingSecondThird(sector)/P
Spec1 = compute_TN(sector)/N
Spec2 = compute_TNIncludingSecond(sector)/N
Spec3 = compute_TNIncludingThird(sector)/N
return S1,Spec1,S2,Spec2,S3,Spec3
sectors = ['Food','ERL','WASH','Health','Shelter/NFI','Protection','Education']
l = compute_metric('Food')
print(l)
metric2 = []
pd.Series(l)
metric2 = pd.DataFrame(index='Sensitivity1 Specificity1 Sensitivity2 Specificity2 Sensitivity3 Specificity3'.split())
for i in sectors:
metric2[i] = compute_metricExtended(i)
#metric.set_index(['A','B','C','D','E','F'])
metric2
#from numpy.random import randn
#np.random.seed(101)
#randn(5,4)
#df = pd.DataFrame(randn(5,4),index='A B C D E'.split(),columns='W X Y Z'.split()) #DataFrame(data,index,columns)
Sensitivity_Specificity = sns.heatmap(data = metric2, annot = True )
Sensitivity_Specificity.figure.savefig('Sensitivity_Specificity.png', bbox_inches = 'tight',
pad_inches = 0)
| notebooks/7) Compare_SeverityPriorities.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # なぜ量子コンピューティングなのか?
# ## コンピューターとは何か?
#
# このページにアクセスされた方は、コンピューターがどのようなものかをご存知でしょう。今日、コンピューターは、ノートパソコンや携帯電話から、信号機を制御するシステムまで、様々な形で利用されています。コンピューターは何でもできるようです。これらのシステムは非常に複雑で特殊なものですが、共通しているのは、コンピューターがある入力情報に対して一連の命令を実行し、何らかの新しい(出力)情報を与えてくれることです。
#
# コンピューターに与える命令は、非常に具体的で曖昧さのないものである必要があります。このような命令の集合を「アルゴリズム」と呼び、コンピュータの研究の多くは、異なるアルゴリズムの挙動を研究しています。このコースでは、キーボードやマウス、画面は使わず、情報とアルゴリズムだけで構成される、最も単純なコンピューターについて考えます。
#
# 
# + [markdown] formulas={"T": {"meaning": "This is the average time our search algorithm takes to run.", "type": "Locally defined variable"}, "exp": {"meaning": "This is the number of digits in our secret number. Because this is a superscript, this means we are doing 10 to the power of d.", "type": "Locally defined variable"}, "prop_to": {"meaning": "'Proportional to': Everything to the left of this symbol is <a href='https://en.wikipedia.org/wiki/Proportionality_(mathematics)'>proportional to</a> the things on the right.", "type": "Universal notation"}} gloss={"resources": {"text": "A resource is anything the algorithm needs to run. In computer science, this usually means either the time needed by the algorithm, or the space (e.g. computer memory).", "title": "Resources"}}
# ## コンピューターのアルゴリズムの分類
#
# 従来のコンピューターの中で量子コンピューターが果たす役割を理解するためには、まず、さまざまなアルゴリズムの性能をどのように測定するかを知る必要があります。
#
# コンピューターサイエンスでは、入力の大きさに応じて使用する[リソース](gloss:resources)がどのように大きくなるかでアルゴリズムを分類します。これをアルゴリズムの*複雑さ*と呼びます。例えば、ある数字が偶数かどうかを判断するアルゴリズムは、その数字の最後の一桁を見るだけでよいわけです。この場合、「入力」は数字で、「出力」は「偶数」か「奇数」のどちらかです。アルゴリズムが完了するまでの時間は、入力された数の大きさに依存しないので、これを*定数時間*アルゴリズムと呼びます。コンピューターによってこの結果を得るのにかかる時間は異なるかもしれませんが、それは他の要因によるもので、入力の長さによるものではありません。
#
# 
#
# 別の例を見てみましょう。今回は、入力は同じ長さの2つの数で、問題はそれらを足すことです。この場合、出力は新しい数字になります。2つの複数桁の数字を足す場合、学校で習ったであろう一般的なアルゴリズムでは、それぞれの数字の右端の桁から始めて、それらを足し合わせます。そして、1桁左に移動して(結果が9より大きい場合は「1」を繰り越す)、この処理を繰り返します。コンピューターは、足すべき桁がなくなるまでこれを繰り返し、アルゴリズムを終了します。
#
# 
#
# <!-- ::: q-block.exercise -->
#
# ### 足し算はどのくらい複雑なのか?
#
# <!-- ::: q-quiz(goal="intro-why-qc-0") -->
#
# <!-- ::: .question -->
#
# この足し算アルゴリズムが完了するまでにかかる時間は...
#
# <!-- ::: -->
#
# <!-- ::: .option(correct) -->
#
# 1. ...入力された数値の長さに線形に(比例して)増大する(線形時間)。
#
# <!-- ::: -->
#
# <!-- ::: .option -->
#
# 1. ...入力された数値の長さに影響されない(定数時間)
#
# <!-- ::: -->
#
# <!-- ::: .option -->
#
# 1. ...入力数の長さの2乗で大きくなる(2次時間)
#
# <!-- ::: -->
#
# <!-- ::: -->
#
# <!-- ::: -->
#
# 繰り返しになりますが、コンピューターによってこのアルゴリズムの実行速度は異なります。ノートパソコンでは、人間の何百万倍もの速さで足し算を実行できます。しかし、1秒間に100万回の演算ができても、1回しかできなくても、増加率は同じです。
#
# 
#
# ここで最後に、私たちにとって非常に興味深い例を一つ紹介しましょう。秘密の番号(暗証番号など)を持っていて、それを当てるという問題があるとします。この場合、問題の大きさは番号の長さです。
#
# 答えが正しいかどうかを確認する唯一の方法が、キーパッドに数字を打ち込むことだとしましょう。その数字が何であるかについての情報はないので、この秘密の数字を見つけるための最適なアルゴリズムは「総当り」方式を使用します。つまり、巧妙なことは何もせず、単に可能な限りの数字を試してみるということです。
#
# どれくらいの時間がかかるのでしょうか?さて、理論的には運が良ければ一回で答えを当てることができますが、これは非常に低い確率です。平均すると、可能な入力の約半分を試さなければならないので、アルゴリズムの実行時間は可能な組み合わせの数に比例します。そこで問題です。可能な組み合わせの数は、秘密番号の長さに応じてどのように増加するのでしょうか?
#
# 
#
# 秘密の番号に桁を追加するごとに、可能な組み合わせの数が10倍されます。例えば、1桁の秘密の数字には10の可能性があり (0, 1, 2, 3, 4, 5, 6, 7, 8, 9)、2桁の秘密の数字には100の可能性があるのです。各桁を当てるのにかかる時間が(長さに関係なく)同じだと仮定すると、次のように数学的に表すことができます。
#
# $$ \cssId{T}{T} \cssId{prop_to}{\propto} 10^\cssId{exp}{d}$$
#
# この式では桁数(d)が指数であることにお気づきでしょう。このように、これは*指数時間*アルゴリズムであり、実行時間は入力の長さに対して指数関数的に増加する、と言います。
#
# 
# + [markdown] gloss={"intractable": {"text": "An intractable problem is one which can be solved in theory, but requires too many resources in practice.", "title": "Intractable"}}
# ## なぜ、このようにアルゴリズムを測定するのか?
#
# コンピューターによって得意分野が異なるため、あるコンピューター上では別のコンピューター上よりもある演算が速くなることがあります。入力サイズに対する時間の増加を研究することによって、アルゴリズムとコンピューターの特定の組み合わせではなく、デバイス固有の詳細を無視し、実際に「アルゴリズム」を測定することができます。重要なことは、アルゴリズムが入力サイズに対してどのように変化するかを知ることで、アルゴリズムが管理可能な大きさになるかどうかもわかるということです。
#
# 上で見た線形時間足し算のアルゴリズムについて考えてみましょう。10桁の数字を1秒につき2つ足すことができたとしたら、直線的な伸び率から、20桁の数字を2秒で足すことができるはずです。10桁増えるごとに、計算時間はおよそ1秒ずつ増えていくはずです。
#
# 一方、上記の指数時間探索アルゴリズムを使って、10桁の暗証番号を1秒で見つけることができたとします。これは、あなたのコンピューターが1秒間に~5,000,000,000の組み合わせを試せるほど高速であることを意味します。このアルゴリズムを使ったコンピューターが20桁の暗証番号を見つけるには、およそ5,000,000,000秒(~150 年)かかると予想されます。さらに10桁増えると、約150,000,000,000年(宇宙の年齢の120倍)にもなります。指数関数時間のアルゴリズムは、わずかな入力桁数(この場合は~30桁)でも、実行するのが難しいどころか、文字通り不可能になってしまうのです。
#
# この暗証番号探索問題は、できるだけ単純にするための人工的な例ですが、コンピューターサイエンスには、非効率なアルゴリズムしかない現実の問題が数多く存在します。今日のコンピューターは驚くほど高速ですが、これらの[困難な](gloss:intractable)問題は、最大のスーパーコンピューターでも難しすぎることがあります。
#
# しかし、より効率的に時間増加するアルゴリズムが見つかれば、比較的遅いコンピューターや信頼性の低いコンピューターでも、これらの困難な問題を突然処理できるようになるかもしれません。そこで登場するのが、量子コンピューターです。
#
# ## 量子コンピューターはどのように役立つのか?
#
# これまで、アルゴリズムというものを非常に抽象的に考えてきましたが、そのアルゴリズムを実行するコンピューターは現実の世界に存在しなければなりません。そのコンピューターが高性能なマイクロチップであろうと、紙とペンを持った人間であろうと、すべてのコンピューターは最終的に物理法則に支配されており、その演算によって私たちが作れるアルゴリズムが限定されてしまうのです。
#
# 物理学は、宇宙に存在するすべてのものが従う一連のルールを解明しようとするものです。20世紀初頭、実験室での繊細な実験を通して、物理学者たちは、現在の物理学では説明できないような奇妙な振る舞いを目にしました。このことは、物理法則が正確でないことを意味します。そこで彼らは、より完全な「量子」物理学を開発し、この挙動を非常にうまく説明することに成功しました。
#
# 物理学者は、これまで見たこともないような振る舞いを説明するために量子物理学を生み出し、コンピューター科学者は、新しく発見されたこの振る舞いを利用することで、より効率的なアルゴリズムが作成することが(理論的には)できることを発見したのです。その結果、従来のコンピューターでは解決不可能な問題でも、この振る舞いを利用できる「量子」コンピューターであれば解決できると考えられるものがあります。そのひとつが*整数の因数分解*です。
#
# 「$x$」と呼ぶ整数があるとします。因数分解アルゴリズムでは、$p×q = x$となるような整数 $p$ と $q$ を求めます。これは簡単な場合もあります。$2000 = 2 × 1000$と一目でわかりますが、$x$が2つの大きな素数の積の場合はこの問題は非常に難しくなります。整数の因数分解について語るとき、最も困難な(最悪の)シナリオを想定することになります。下のコードセルでは、変数<code>x</code>に250桁の数字を代入しています。
# -
x = 2140324650240744961264423072839333563008614715144755017797754920881418023447140136643345519095804679610992851872470914587687396261921557363047454770520805119056493106687691590019759405693457452230589325976697471681738069364894699871578494975937497937
# + [markdown] gloss={"coreyears": {"text": "Conventional computer chips are often made from processors called <a href=\"https://en.wikipedia.org/wiki/Multi-core_processor\">\"cores\"</a>. A <i>core-year</i> is the equivalent of using one of these cores continuously for a year. For reference, a modern laptops have around 2-4 cores. The meaning of this number depends on how powerful the core is, but this should give you a rough idea of the computing power involved.", "title": "Core Year"}}
# 2020年に、研究者は古典的なスーパーコンピューターと~2700[コア年](gloss:coreyears)の処理能力を用いてこの数を因数分解しました。これは大規模な作業であり、本稿執筆時点では記録破りである。彼らの結果は、以下のコードセルで確認することができます(幸運なことに、私たちには乗算の効率的なアルゴリズムがあります!)。
# +
p = 64135289477071580278790190170577389084825014742943447208116859632024532344630238623598752668347708737661925585694639798853367
q = 33372027594978156556226010605355114227940760344767554666784520987023841729210037080257448673296881877565718986258036932062711
p*q == x # 「True」と評価される
# + [markdown] gloss={"RSA": {"text": "RSA numbers are numbers taken from the RSA factoring challenge. These numbers are intentionally chosen to be difficult to factor.<p>'RSA' are the initials of three of the people that invented the protocol that uses these large numbers to encrypt information.", "title": "RSA Number"}}
# 表示される出力は、セルの最終行の値です。この場合、<code>p*q == x</code>は<code>True</code>と評価されることがわかります。数学的に証明されているわけではありませんが、従来のコンピューターでこのような数の因数分解を行う効率的なアルゴリズムが存在しないことは確かです。実際、インターネットの暗号化の多くは、この問題が解決不可能であるという仮定に依存しており、617桁の[RSA](gloss:RSA)数の因数分解は不可能であるとされています。一方、量子コンピューターでは、十分な大きさの量子コンピューターができれば、1日以内にこれらの数字を因数分解することができると推定される効率的な因数分解アルゴリズムがわかっています。
# + [markdown] gloss={"noise": {"text": "Noise is useless information that's difficult to distinguish from useful information. For example, it's hard to hear someone talking to you if there are lots of other people talking loudly nearby.", "title": "Noise"}, "qiskit": {"text": "Qiskit is a software development kit for working with quantum computers.", "title": "Qiskit"}, "qubits": {"text": "A 'qubit' is a 'quantum bit'. We will study these later in this course.", "title": "Qubit"}, "transistor": {"text": "A transistor is an electronic device. They can be used to switch electric currents on and off, and can be used to build a computer processor.", "title": "Transistor"}}
# ## 今、私たちはどこにいるのか?
#
# 量子コンピューターは、より効率的なアルゴリズムを実行できることが分かっていますが、現在ある量子コンピューターは小さくて不安定なため、従来のコンピューターと比較して優位性を発揮することはできません。
#
# 量子コンピューターが解決できる問題の大きさを制限する要因は、ごく単純に考えても2つあります。1つ目は、量子コンピューターが保存・処理できるデータの量です。これは通常、[*量子ビット*](gloss:qubits)で測定されます。もし十分な量子ビットがなければ、あるサイズ以上の問題を保存・処理することはできません。2つ目は、量子コンピューターのエラーレートです。量子的な振る舞いは、実験室での繊細な実験でしか見ることができないため、量子コンピューターを作るのは繊細な作業です。今ある量子コンピューターはノイズが多いので、よく間違いますし、結果に「[ノイズ](gloss:noise)」が入ります。ノイズが多すぎると、結果が意味のないものになってしまうのです!
#
# 今のところ、量子コンピューターは実験的なものです。量子ビットの数やエラーレートに制限があるため、現在解決できる最大の問題は、従来のコンピューターでも容易に解決できるものです。
#
# 未来のある時点で、これは変わるでしょう。量子コンピューターを使った方が、従来のコンピューターよりも経済的に有利に問題を解決できるという「量子アドバンテージ」に到達するのです。なぜそれがわかるのでしょうか?*なぜなら、私たちはアルゴリズムをその増加率で測っているからです!*量子コンピューターが順調に発展し続ける限り、いずれは古典的なコンピューターを追い抜くことが分かっているのです。
#
# 
#
# 617桁のRSA数を1日以内に因数分解するには、~2,000万のノイズのある量子ビットが必要であるとされています。本稿執筆時点で、IBMは現在65量子ビットの量子コンピューターを保有しており、2023年までに1000量子ビットを超えるシステムの構築を目指しています。このマイルストーンのずっと前に、量子アドバンテージをもたらすと思われるアルゴリズムが他にもありますが、まだまだ先の話と思われるかもしれません。
#
# 従来のコンピューターがどこから来たのか、思い起こしてみる必要があります。下の写真は、1947年に作られた最初の[トランジスター](gloss:transistor)の写真です。トランジスターは、現代のコンピュータープロセッサーの構成要素です。
#
# 画像クレジット: 連邦職員<a href="https://clintonwhitehouse4.archives.gov/Initiatives/Millennium/capsule/mayo.html">リンク</a>、<a href="https://commons.wikimedia.org/w/index.php?curid=554340">パブリックドメイン</a>。
#
# それから70年、現代のコンピューターチップには、何十億というトランジスターが搭載されています。
#
# このコースの残りの部分では、より効率的なアルゴリズムを作成することを可能にする量子効果を探ります。このコースの終わりには、ソフトウェアパッケージである[Qiskit](gloss:qiskit)を使って、これらのアルゴリズムの1つを実行する量子コンピューターをプログラムすることができるようになるはずです。
# -
# <!-- ::: q-block.exercise -->
#
# ### クイッククイズ
#
# <!-- ::: q-quiz(goal="intro-why-qc-1") -->
#
# <!-- ::: .question -->
#
# 量子コンピューターは、いずれ...
#
# <!-- ::: -->
#
# <!-- ::: .option(correct) -->
#
# 1. ...従来のコンピューターでは困難な計算を行うことができます。
#
# <!-- ::: -->
#
# <!-- ::: .option -->
#
# 1. ...従来のコンピューターに置き換わります。
#
# <!-- ::: -->
#
# <!-- ::: .option -->
#
# 1. ...従来のコンピューターの速度を向上させます。
#
# <!-- ::: -->
#
# <!-- ::: -->
#
# <!-- ::: -->
| translations/ja/intro/why-quantum-computing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# +
# default_exp datasets.bases.interactions
# -
# # Interactions Dataset
# > Implementation of base modules for interactions dataset.
# +
#export
from typing import Any, Iterable, List, Optional, Tuple, Union, Callable
import sys
import os
from os import path as osp
import collections
import random
import numpy as np
import pandas as pd
from scipy.sparse import coo_matrix
import torch
from torch.utils.data import DataLoader, Dataset
from pytorch_lightning import LightningDataModule
from recohut.utils.common_utils import *
from recohut.datasets import base
from recohut.utils.splitting import random_split, stratified_split
import warnings
warnings.filterwarnings('ignore')
# -
# ## Interactions
#export
class Interactions(torch.utils.data.Dataset):
"""
Hold data in the form of an interactions matrix.
Typical use-case is like a ratings matrix:
- Users are the rows
- Items are the columns
- Elements of the matrix are the ratings given by a user for an item.
"""
def __init__(self, mat):
if isinstance(mat, np.ndarray):
mat = coo_matrix(mat)
self.mat = mat.astype(np.float32).tocoo()
self.n_users = self.mat.shape[0]
self.n_items = self.mat.shape[1]
def __getitem__(self, index):
row = self.mat.row[index]
col = self.mat.col[index]
val = self.mat.data[index]
return (row, col), val
def __len__(self):
return self.mat.nnz
matrix = np.random.randint(1,5,(5,5)).astype(np.float32)
matrix
interactions = Interactions(matrix)
interactions.__getitem__(1)
# ## InteractionsDataset
#export
class InteractionsDataset(torch.utils.data.Dataset, base.Dataset):
def __init__(self,
data_dir,
min_rating=None,
min_uc=5,
min_sc=5,
num_negative_samples=100,
max_samples=200,
data_type=None,
split_type='random',
val_p=0.2,
test_p=0.2,
seed=42,
*args,
**kwargs):
"""
Args:
data_dir: Where to save/load the data
min_uc: minimum user count to keep in the data
min_sc: minimum item count to keep in the data
min_rating: minimum rating threshold to convert explicit feedback into implicit
num_negative_samples: number of negative samples for each positive one
max_samples: max samples limit
data_type: train/valid/test
split_type: data split method - stratified/random
val_p: Percent (float) or number (int) of samples to use for the validation split
test_p: Percent (float) or number (int) of samples to use for the test split
seed: Random seed to be used for train/val/test splits
"""
self.min_rating = min_rating
self.min_uc = min_uc
self.min_sc = min_sc
self.num_negative_samples = num_negative_samples
self.max_samples = 200
self.data_type = data_type
self.val_p = val_p if val_p is not None else 0.2
self.test_p = test_p if test_p is not None else 0.2
self.seed = seed
self.split_type = split_type
super().__init__(data_dir)
assert self.min_uc >= 2, 'Need at least 2 ratings per user for validation and test'
self._process()
if self.data_type is not None:
self.load()
@property
def raw_file_names(self):
raise NotImplementedError
@property
def processed_file_names(self):
return ['data_train.pt',
'data_valid_pos.pt',
'data_valid_neg.pt',
'data_test_pos.pt',
'data_test_neg.pt']
def download(self):
raise NotImplementedError
def make_implicit(self, df):
"convert the explicit data to implicit by only keeping interactions with a rating >= min_rating"
print('Turning into implicit ratings')
df = df[df['rating'] >= self.min_rating].reset_index(drop=True)
df['rating'] = 1
return df
def filter_triplets(self, df):
print('Filtering triplets')
if self.min_sc > 0 or self.min_uc > 0:
item_sizes = df.groupby('sid').size()
good_items = item_sizes.index[item_sizes >= self.min_sc]
user_sizes = df.groupby('uid').size()
good_users = user_sizes.index[user_sizes >= self.min_uc]
while len(good_items) < len(item_sizes) or len(good_users) < len(user_sizes):
if self.min_sc > 0:
item_sizes = df.groupby('sid').size()
good_items = item_sizes.index[item_sizes >= self.min_sc]
df = df[df['sid'].isin(good_items)]
if self.min_uc > 0:
user_sizes = df.groupby('uid').size()
good_users = user_sizes.index[user_sizes >= self.min_uc]
df = df[df['uid'].isin(good_users)]
item_sizes = df.groupby('sid').size()
good_items = item_sizes.index[item_sizes >= self.min_sc]
user_sizes = df.groupby('uid').size()
good_users = user_sizes.index[user_sizes >= self.min_uc]
return df
def densify_index(self, df):
print('Densifying index')
umap = {u: i for i, u in enumerate(set(df['uid']))}
smap = {s: i for i, s in enumerate(set(df['sid']))}
df['uid'] = df['uid'].map(umap)
df['sid'] = df['sid'].map(smap)
return df, umap, smap
def load_ratings_df(self):
raise NotImplementedError
@staticmethod
def _convert_to_torch_sparse(mat):
values = mat.data
indices = np.vstack((mat.row, mat.col))
i = torch.LongTensor(indices)
v = torch.FloatTensor(values)
shape = mat.shape
return torch.sparse.FloatTensor(i, v, torch.Size(shape))
def process(self):
df = self.load_ratings_df()
if self.min_rating:
df = self.make_implicit(df)
df = self.filter_triplets(df)
df, umap, smap = self.densify_index(df)
self.num_users = max(df.uid) + 1 # df.uid.nunique()
self.num_items = max(df.sid) + 1 # df.sid.nunique()
mat = coo_matrix((np.array(df.rating),
(np.array(df.uid), np.array(df.sid))),
shape=(self.num_users, self.num_items))
if self.split_type == 'random':
mat_train, mat_valid, mat_test = random_split(mat = mat,
val_p = self.val_p,
test_p = self.test_p,
seed = self.seed)
elif self.split_type == 'stratified':
mat_train, mat_valid, mat_test = stratified_split(mat = mat,
val_p = self.val_p,
test_p = self.test_p,
seed = self.seed)
mat_train = self._convert_to_torch_sparse(mat_train)
torch.save(mat_train, self.processed_paths[0])
mat_valid_pos = self._convert_to_torch_sparse(mat_valid)._indices().T
_, indices = np.unique(mat_valid_pos[:, 0], return_index=True)
mat_valid_pos = mat_valid_pos[indices, :]
torch.save(mat_valid_pos, self.processed_paths[1])
pos_items = set(zip(mat_valid.row, mat_valid.col))
mat_valid_neg = self._negative_sample(np.arange(mat_valid.shape[0]), pos_items)
mat_valid_neg = torch.tensor(mat_valid_neg, dtype=torch.int)
torch.save(mat_valid_neg, self.processed_paths[2])
mat_test_pos = self._convert_to_torch_sparse(mat_test)._indices().T
_, indices = np.unique(mat_test_pos[:, 0], return_index=True)
mat_test_pos = mat_test_pos[indices, :]
torch.save(mat_test_pos, self.processed_paths[3])
pos_items = set(zip(mat_test.row, mat_test.col))
mat_test_neg = self._negative_sample(np.arange(mat_test.shape[0]), pos_items)
mat_test_neg = torch.tensor(mat_test_neg, dtype=torch.int)
torch.save(mat_test_neg, self.processed_paths[4])
return mat
def todense(self) -> np.matrix:
"""Transforms sparse matrix to np.matrix, 2-d."""
return self.mat.todense()
def toarray(self) -> np.array:
"""Transforms sparse matrix to np.array, 2-d."""
return self.mat.toarray()
def head(self, n: int = 5) -> np.array:
"""Return the first ``n`` rows of the dense matrix as a np.array, 2-d."""
n = self._prep_head_tail_n(n=n)
return self.mat.tocsr()[range(n), :].toarray()
def tail(self, n: int = 5) -> np.array:
"""Return the last ``n`` rows of the dense matrix as a np.array, 2-d."""
n = self._prep_head_tail_n(n=n)
return self.mat.tocsr()[range(-n, 0), :].toarray()
def _prep_head_tail_n(self, n: int) -> int:
"""Ensure we don't run into an ``IndexError`` when using ``head`` or ``tail`` methods."""
if n < 0:
n = self.num_users + n
if n > self.num_users:
n = self.num_users
return n
def _negative_sample(self, user_id: Union[int, np.array], positive_items) -> np.array:
"""Generate negative samples for a ``user_id``."""
if self.max_samples > 0:
# if we are here, we are doing true negative sampling
negative_item_ids_list = list()
if not isinstance(user_id, collections.abc.Iterable):
user_id = [user_id]
for specific_user_id in user_id:
# generate true negative samples for the ``user_id``
samples_checked = 0
temp_negative_item_ids_list = list()
while len(temp_negative_item_ids_list) < self.num_negative_samples:
negative_item_id = random.choice(range(self.num_items))
# we have a negative sample, make sure the user has not interacted with it
# before, else we resample and try again
while (
(specific_user_id, negative_item_id) in positive_items
or negative_item_id in temp_negative_item_ids_list
):
if samples_checked >= self.max_samples:
num_samples_left_to_generate = (
self.num_negative_samples - len(temp_negative_item_ids_list) - 1
)
temp_negative_item_ids_list += random.choices(
range(self.num_items), k=num_samples_left_to_generate
)
break
negative_item_id = random.choice(range(self.num_items))
samples_checked += 1
temp_negative_item_ids_list.append(negative_item_id)
negative_item_ids_list += [np.array(temp_negative_item_ids_list)]
if len(user_id) > 1:
negative_item_ids_array = np.stack(negative_item_ids_list)
else:
negative_item_ids_array = negative_item_ids_list[0]
else:
# if we are here, we are doing approximate negative sampling
if isinstance(user_id, collections.abc.Iterable):
size = (len(user_id), self.num_negative_samples)
else:
size = (self.num_negative_samples,)
negative_item_ids_array = np.random.randint(
low=0,
high=self.num_items,
size=size,
)
return negative_item_ids_array
def load(self):
if self.data_type=='train':
self.train = torch.load(self.processed_paths[0])
self.train_pos = self.train._indices().T
self.n_users, self.n_items = self.train.size()
self.train_score = torch.sparse.sum(self.train, dim=0).to_dense().repeat((self.n_users, 1))
self.train_score[self.train_pos[:, 0], self.train_pos[:, 1]] = 0
elif self.data_type=='valid':
self.valid_pos = torch.load(self.processed_paths[1])
self.valid_neg = torch.load(self.processed_paths[2])
self.n_users = self.valid_pos.shape[0]
valid_items = []
for u in range(self.n_users):
items = torch.cat((self.valid_pos[u, 1].view(1), self.valid_neg[u]))
valid_items.append(items)
self.valid_items = torch.vstack(valid_items)
self.valid_labels = torch.zeros(self.valid_items.shape)
self.valid_labels[:, 0] += 1
else:
self.test_pos = torch.load(self.processed_paths[3])
self.test_neg = torch.load(self.processed_paths[4])
self.n_users = self.test_pos.shape[0]
test_items = []
for u in range(self.n_users):
items = torch.cat((self.test_pos[u, 1].view(1), self.test_neg[u]))
test_items.append(items)
self.test_items = torch.vstack(test_items)
self.test_labels = torch.zeros(self.test_items.shape)
self.test_labels[:, 0] += 1
def __len__(self):
return self.n_users
def __train__(self, index):
return self.train_pos[index], self.train_score[self.train_pos[index][0]]
def __valid__(self, index):
return self.valid_pos[index], self.valid_items[index], self.valid_labels[index]
def __test__(self, index):
return self.test_pos[index], self.test_items[index], self.test_labels[index]
def __getitem__(self, index):
if self.data_type=='train':
return self.__train__(index)
elif self.data_type=='valid':
return self.__valid__(index)
else:
return self.__test__(index)
# Example:
class ML1mDataset(InteractionsDataset):
url = "http://files.grouplens.org/datasets/movielens/ml-1m.zip"
@property
def raw_file_names(self):
return 'ratings.dat'
def download(self):
path = download_url(self.url, self.raw_dir)
extract_zip(path, self.raw_dir)
from shutil import move, rmtree
move(osp.join(self.raw_dir, 'ml-1m', self.raw_file_names), self.raw_dir)
rmtree(osp.join(self.raw_dir, 'ml-1m'))
os.unlink(path)
def load_ratings_df(self):
df = pd.read_csv(self.raw_paths[0], sep='::', header=None, engine='python')
df.columns = ['uid', 'sid', 'rating', 'timestamp']
# drop duplicate user-item pair records, keeping recent ratings only
df.drop_duplicates(subset=['uid', 'sid'], keep='last', inplace=True)
return df
# ## InteractionsDataModule
#export
class InteractionsDataModule(LightningDataModule):
dataset_cls: str = ""
def __init__(self,
data_dir: Optional[str] = None,
num_workers: int = 0,
normalize: bool = False,
batch_size: int = 32,
shuffle: bool = True,
pin_memory: bool = True,
drop_last: bool = False,
*args,
**kwargs) -> None:
"""
Args:
data_dir: Where to save/load the data
num_workers: How many workers to use for loading data
normalize: If true applies rating normalize
batch_size: How many samples per batch to load
shuffle: If true shuffles the train data every epoch
pin_memory: If true, the data loader will copy Tensors into CUDA pinned memory before
returning them
drop_last: If true drops the last incomplete batch
"""
super().__init__(data_dir)
self.data_dir = data_dir if data_dir is not None else os.getcwd()
self.num_workers = num_workers
self.normalize = normalize
self.batch_size = batch_size
self.shuffle = shuffle
self.pin_memory = pin_memory
self.drop_last = drop_last
self.kwargs = kwargs
def prepare_data(self, *args: Any, **kwargs: Any) -> None:
"""Saves files to data_dir."""
self.data = self.dataset_cls(self.data_dir, **self.kwargs)
def setup(self, stage: Optional[str] = None) -> None:
"""Creates train, val, and test dataset."""
if stage == "fit" or stage is None:
self.dataset_train = self.dataset_cls(self.data_dir, **self.kwargs, data_type='train')
self.dataset_val = self.dataset_cls(self.data_dir, **self.kwargs, data_type='valid')
if stage == "test" or stage is None:
self.dataset_test = self.dataset_cls(self.data_dir, **self.kwargs, data_type='test')
def train_dataloader(self, *args: Any, **kwargs: Any) -> DataLoader:
"""The train dataloader."""
return self._data_loader(self.dataset_train, shuffle=self.shuffle)
def val_dataloader(self, *args: Any, **kwargs: Any) -> Union[DataLoader, List[DataLoader]]:
"""The val dataloader."""
return self._data_loader(self.dataset_val)
def test_dataloader(self, *args: Any, **kwargs: Any) -> Union[DataLoader, List[DataLoader]]:
"""The test dataloader."""
return self._data_loader(self.dataset_test)
def _data_loader(self, dataset: Dataset, shuffle: bool = False) -> DataLoader:
return DataLoader(
dataset,
batch_size=self.batch_size,
shuffle=shuffle,
num_workers=self.num_workers,
drop_last=self.drop_last,
pin_memory=self.pin_memory,
)
# Example:
class ML1mDataModule(InteractionsDataModule):
dataset_cls = ML1mDataset
#hide
# %reload_ext watermark
# %watermark -a "<NAME>." -m -iv -u -t -d -p recohut,pytorch_lightning
| nbs/datasets/bases/datasets.bases.interactions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.10.4 ('wilds')
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
# +
import sys
import os
sys.path.append(os.path.join("..", "helpers"))
from unet import UNet
import torch
device = torch.device("cuda" if torch.cuda.is_available() else "cpu") # PyTorch v0.4.0
# Unet has skip connections removed
model = UNet(in_channels=3,
out_channels=3,
n_blocks=3,
start_filters=32,
activation='relu',
normalization='batch',
conv_mode='same',
dim=2)
model = model.to(device)
input = torch.randn(size=(8, 3, 224, 224), dtype=torch.float32)
input = input.to(device)
with torch.no_grad():
out = model(input, encode_only=False)
features = model(input, encode_only=True)
print(f'Input: {input.shape}')
print(f'Out: {out.shape}')
print(f'Features: {features.shape}')
# +
from torchsummary import summary
device = torch.device("cuda" if torch.cuda.is_available() else "cpu") # PyTorch v0.4.0
model = model.to(device)
summary = summary(model, (3, 224, 224))
# +
shape = 224
def compute_max_depth(shape, max_depth=10, print_out=True):
shapes = []
shapes.append(shape)
for level in range(1, max_depth):
if shape % 2 ** level == 0 and shape / 2 ** level > 1:
shapes.append(shape / 2 ** level)
if print_out:
print(f'Level {level}: {shape / 2 ** level}')
else:
if print_out:
print(f'Max-level: {level - 1}')
break
return shapes
out = compute_max_depth(shape, print_out=True, max_depth=10)
| experiments/unet_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### Поиск минимального числа.
# #### Search for the minimum number.
# + pycharm={"name": "#%%\n", "is_executing": false}
a = [int(i) for i in input().split()]
print(a)
# + pycharm={"name": "#%%\n", "is_executing": false}
m = a[0]
for x in a:
if m > x:
m = x
print(m)
| stepik/programming_on_python/2_6_0_tasks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Welcome
# -------
#
# Source: https://github.com/piotrszul/spark-tutorial (great tutorials to get up to speed)
#
# Welcome to the Apache Spark tutorial notebooks.
#
# This very simple notebook is designed to test that your environment is setup correctly.
#
# Please `Run All` cells.
#
# The notebook should run without errors and you should see a histogram plot at the end.
#
# (You can also check the expected output [here](https://piotrszul.github.io/spark-tutorial/notebooks/0.1_Welcome.html))
#
#
# #### Let's go
#
# Let's check that there are some input data available:
# + language="sh"
#
# # All the test data sets are located in the `data` directory.
# # You can preview them using unix command such as `cat`, `head`, `tail`, `ls`, etc.
# # in `shell` cells marked with the `%%sh` magic, e.g.:
#
# head -n 10 prince_by_machiavelli.txt
# -
# Let's check if spark is available and what version are we using (should be 2.1+):
# +
# `spark` is the main entry point for all spark related operations.
# It is an instance of SparkSession and pyspark automatically creates one for you.
# Another one is `sc` an instance of SparkContext, which is used for low lever RRD API.
spark.version
# -
# Let's try to run a simple `Spark` program to compute the number of occurences of words in Machiavelli's "Prince", and display ten most frequent ones:
# +
import operator
import re
# Here we use Spark RDD API to split a text file into invividual words,
# to count the number of occurences of each word and to take top 10 most frequent words.
wordCountRDD = sc.textFile('prince_by_machiavelli.txt') \
.flatMap(lambda line: re.split(r'[^a-z\-\']+', line.lower())) \
.filter(lambda word: len(word) > 0 ) \
.map(lambda word: (word, 1)) \
.reduceByKey(operator.add)
# `take()` function takes the first n elements of an RDD
# and returns them in a python `list` object,
top10Words = wordCountRDD \
.map(lambda x: (x[1],x[0])) \
.sortByKey(False) \
.take(10)
# which can then be printed out
print(top10Words)
# -
# Spark SQL is a higer level API for structured data. The data are represented in `data frames` - table like object with columns and rows concenptully similar to `panadas` or `R` data fames.
#
# Let's use Spark SQL to display a table with the 10 least frequent words:
# +
# A data frame can be created from an RDD;
# schema defines the names (and types) of columns.
wordCountDF = spark.createDataFrame(wordCountRDD, schema = ['word', 'count'])
# it just means: sort by column `count` and take the first ten elements
bottom10Words = wordCountDF.sort('count').limit(10)
# `display` function can be used to display data frames (and also all other sorts of objects)
bottom10Words.show()
# -
# Let's save the results to a csv file.
#
# For the tutorial all the output files are saved in the `output` directory:
# +
# data frames can be saved in many common 'table' formats, for example `csv`.
# the `mode='overwrite'` tells Spark to overwite the output file is it exists
wordCountDF.write.csv('output/prince-word-count.csv', mode='overwrite', header=True)
# -
# Let's preview the output:
# + language="sh"
#
# # Same as with the input data sets, we can use the `%%sh` cells to preview the
# # files produced to the `output` directory.
#
# # Please note that output we have produced above is actually a directory:
#
# ls -l output/prince-word-count.csv
#
# # The `part-*` files inside contain the actual data.
#
# echo
# echo "Content:"
#
# head -n 10 output/prince-word-count.csv/part-00000-*.csv
# -
# Finally we can use python `matplotlib` to visualise the result.
#
# Let's plot the histogram of the distribution of word counts:
# +
import matplotlib.pyplot as plt
# we can convert (small) Spark data frames to `pandas`
wordCountPDF = wordCountDF.toPandas()
# and then use pyplot (plt) to display the results
# Please note that we call `plt.close()` first - this is needed for Databricks
# to start a new plot.
plt.close()
plt.hist(wordCountPDF['count'], bins = 20, log = True)
plt.show()
# -
# You can now play around modifyging some pieces of the code.
#
# When you are done and you are running off the local machine remeber **to close the notebook with `File/Close and Halt`**
| 07-bigdata-spark-databricks/pyspark-local-training/pyspark_welcome.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] run_control={"frozen": false, "read_only": false}
# # Tests on PDA
# + run_control={"frozen": false, "read_only": false}
from jove.SystemImports import *
from jove.DotBashers import *
from jove.Def_md2mc import *
from jove.Def_PDA import *
# + run_control={"frozen": false, "read_only": false}
a1b2_s = md2mc(src="File", fname="machines/pdafiles/a1b2_accept_s.pda")
dotObj_pda(a1b2_s)
# + run_control={"frozen": false, "read_only": false}
dotObj_pda(a1b2_s, FuseEdges=True)
# + run_control={"frozen": false, "read_only": false}
help(explore_pda)
# + run_control={"frozen": false, "read_only": false}
explore_pda("abb", a1b2_s, acceptance='ACCEPT_S')
# + run_control={"frozen": false, "read_only": false}
explore_pda("bab", a1b2_s, acceptance='ACCEPT_S')
# + run_control={"frozen": false, "read_only": false}
explore_pda("bba", a1b2_s, acceptance='ACCEPT_S')
# + run_control={"frozen": false, "read_only": false}
explore_pda("bbaabbbabaabbabbbb", a1b2_s, acceptance='ACCEPT_S')
# + run_control={"frozen": false, "read_only": false}
explore_pda("babaababbbaabbbbbb", a1b2_s, acceptance='ACCEPT_S')
# + run_control={"frozen": false, "read_only": false}
explore_pda("abbaababbbabbbbbba", a1b2_s, acceptance='ACCEPT_S')
# + run_control={"frozen": false, "read_only": false}
a1b2_f = md2mc(src="File", fname="machines/pdafiles/a1b2_accept_f.pda")
dotObj_pda(a1b2_f)
# + run_control={"frozen": false, "read_only": false}
dotObj_pda(a1b2_f, FuseEdges=True)
# + run_control={"frozen": false, "read_only": false}
explore_pda("abbaababbbabbbbbba", a1b2_f) # Default is accept_f
# + run_control={"frozen": false, "read_only": false}
explore_pda("babaababbbaabbbbbb", a1b2_f, acceptance='ACCEPT_F') # default acceptance
# + run_control={"frozen": false, "read_only": false}
explore_pda("bbaabbbabaabbabbbb", a1b2_f, STKMAX=2)
# + run_control={"frozen": false, "read_only": false}
pdaDyck = md2mc('''PDA
IF : (, #; (# -> A
A : (, (; (( -> A
A : ), (; '' -> A
A : '',#; # -> IF
''')
DOpdaDyck = dotObj_pda(pdaDyck, FuseEdges=True)
DOpdaDyck
# + run_control={"frozen": false, "read_only": false}
explore_pda("", pdaDyck)
# + run_control={"frozen": false, "read_only": false}
explore_pda("()", pdaDyck)
# + run_control={"frozen": false, "read_only": false}
explore_pda("()()(())", pdaDyck)
# + run_control={"frozen": false, "read_only": false}
explore_pda("()()(()", pdaDyck)
# + run_control={"frozen": false, "read_only": false}
DOpdaDyck.source
# + run_control={"frozen": false, "read_only": false}
pda1 = md2mc('''PDA
I : a, b; c -> F
''')
DOpda1 = dotObj_pda(pda1, FuseEdges=True)
DOpda1
# + run_control={"frozen": false, "read_only": false}
DOpda1.source
# + run_control={"frozen": false, "read_only": false}
pda2 = md2mc('''PDA
I : a, b ; c -> F
I : '', ''; d -> A
A : '', d ; '' -> F
''')
DOpda2 = dotObj_pda(pda2, FuseEdges=True)
DOpda2
# + run_control={"frozen": false, "read_only": false}
DOpda2.source
# + run_control={"frozen": false, "read_only": false}
explore_pda("a", pda1)
# + run_control={"frozen": false, "read_only": false}
explore_pda("a", pda2)
# + run_control={"frozen": false, "read_only": false}
pda3 = md2mc('''PDA
I : a, b ; c -> F
I : '', ''; d -> A
A : a, d ; '' -> F
''')
DOpda3 = dotObj_pda(pda3, FuseEdges=True)
DOpda3
# + run_control={"frozen": false, "read_only": false}
DOpda3.source
# + run_control={"frozen": false, "read_only": false}
explore_pda("a", pda3)
# + run_control={"frozen": false, "read_only": false}
pda4 = md2mc('''PDA
I : a, # ; c -> F
I : '', ''; d -> A
A : a, d ; '' -> F
''')
DOpda4 = dotObj_pda(pda4, FuseEdges=True)
DOpda4
# + run_control={"frozen": false, "read_only": false}
DOpda4.source
# + run_control={"frozen": false, "read_only": false}
explore_pda("a", pda4)
# + run_control={"frozen": false, "read_only": false}
pda5 = md2mc('''PDA
I : a, # ; c -> F
I : '', ''; d -> A
A : '', ''; '' -> A
A : a, d ; '' -> F
''')
DOpda5 = dotObj_pda(pda5, FuseEdges=True)
DOpda5
# + run_control={"frozen": false, "read_only": false}
DOpda5.source
# + run_control={"frozen": false, "read_only": false}
explore_pda("a", pda5)
# + run_control={"frozen": false, "read_only": false}
pda6 = md2mc('''PDA
I : a, # ; c -> F
I : '', ''; d -> A
A : '', ''; z -> A
A : '', z ; '' -> B
B : '', z ; '' -> C
C : '', z ; '' -> C
C : '', # ; '' | a, d; '' -> F
A : a, d ; '' -> F
''')
DOpda6 = dotObj_pda(pda6, FuseEdges=True)
DOpda6
# + run_control={"frozen": false, "read_only": false}
DOpda6.source
# + run_control={"frozen": false, "read_only": false}
explore_pda("a", pda6)
# + run_control={"frozen": false, "read_only": false}
explore_pda("a", pda6, chatty=True)
# + run_control={"frozen": false, "read_only": false}
explore_pda("a", pda6, STKMAX = 3, chatty=True)
# + run_control={"frozen": false, "read_only": false}
| notebooks/driver/Drive_BasicPDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Purpose
#
# This shows the improvement because of our thing. It's the big cahuna.
#
# ### Method
#
# I did 3 very large runs. Pretraining for 200 for each, and then training against
# critic with one, other, and both critic-counts missing.
#
# We'll show the grahs over epochs. We'll also show the training critic-loss, to see how well it can fit that curve.
#
# ### Data structure
#
# Each of the files is a JSON object. Its keys are the 3 training phases: ACTOR, CRITIC, and AC.
# Underneath this, it has subfields for what it was recording. For example, NDCG, or TRAINING_CRITIC_ERROR.
#
# ### Conclusions
#
# It seems as though BOTH_COUNTS is BEST, NO_COUNTS is WORST, WITHOUT_SEEN is pretty good, and WITHOUT_UNSEEN is pretty bad.
#
# My guess at reasons:
#
# * The magnitude of the NLL is probably a pretty good proxy for num_seen, because
#
# +
import matplotlib
import numpy as np
import matplotlib.pyplot as plt
import json
import seaborn as sns
sns.set()
DATA = {}
DATA['ml-20m'] = {}
DATA['ml-20m']['VAE'] = {}
DATA['netflix-prize'] = {}
DATA['netflix-prize']['VAE'] = {}
# DATA['without_seen'] = {}
# DATA['without_unseen'] = {}
# DATA['tentative_with_both'] = {}
print("Now, loading data")
with open("./data/ml-20m/VAE/actor_training.json", "r") as f:
DATA['ml-20m']['VAE']['actor'] = json.loads(f.read())
with open("./data/ml-20m/VAE/critic_training.json", "r") as f:
DATA['ml-20m']['VAE']['critic'] = json.loads(f.read())
with open("./data/ml-20m/VAE/warp_critic_training.json", "r") as f:
DATA['ml-20m']['VAE']['warp_critic'] = json.loads(f.read())
with open("./data/ml-20m/VAE/lambdarank_critic_training.json", "r") as f:
DATA['ml-20m']['VAE']['lambdarank_critic'] = json.loads(f.read())
with open("./data/netflix-prize/VAE/actor_training.json", "r") as f:
DATA['netflix-prize']['VAE']['actor'] = json.loads(f.read())
with open("./data/netflix-prize/VAE/critic_training.json", "r") as f:
DATA['netflix-prize']['VAE']['critic'] = json.loads(f.read())
with open("./data/netflix-prize/VAE/warp_critic_training.json", "r") as f:
DATA['netflix-prize']['VAE']['warp_critic'] = json.loads(f.read())
with open("./data/netflix-prize/VAE/lambdarank_critic_training.json", "r") as f:
DATA['netflix-prize']['VAE']['lambdarank_critic'] = json.loads(f.read())
# with open("./data/without_seen/actor_training.json", "r") as f:
# DATA['without_seen']['actor'] = json.loads(f.read())
# with open("./data/without_seen/critic_training.json", "r") as f:
# DATA['without_seen']['critic'] = json.loads(f.read())
# with open("./data/without_unseen/actor_training.json", "r") as f:
# DATA['without_unseen']['actor'] = json.loads(f.read())
# with open("./data/without_unseen/critic_training.json", "r") as f:
# DATA['without_unseen']['critic'] = json.loads(f.read())
# with open("./data/tentative_with_both/actor_training.json", "r") as f:
# DATA['tentative_with_both']['actor'] = json.loads(f.read())
# with open("./data/tentative_with_both/critic_training.json", "r") as f:
# DATA['tentative_with_both']['critic'] = json.loads(f.read())
print("Data loaded")
# +
print("First, we'll plot just the actor")
# https://stackoverflow.com/questions/20618804/how-to-smooth-a-curve-in-the-right-way
plt.clf()
plt.plot(range(200), DATA['ml-20m']['VAE']['actor']['ACTOR']['ndcg'], color="black")
# plt.plot(range(200), DATA['without_seen']['actor']['ACTOR']['ndcg'], color="blue")
# plt.plot(range(200), DATA['without_unseen']['actor']['ACTOR']['ndcg'], color="green")
# plt.plot(range(200), DATA['tentative_with_both']['actor']['ACTOR']['ndcg'], color="orange")
plt.ylim(0.40, 0.44)
plt.show()
# +
print("Now we'll add the critic to the end")
plt.clf()
plt.plot(range(200), DATA['ml-20m']['VAE']['actor']['ACTOR']['ndcg'], color="black")
plt.plot(range(200, 300), DATA['ml-20m']['VAE']['critic']['AC']['ndcg'], color="red")
plt.ylim(0.40, 0.45)
plt.show()
# +
print("Now we'll add the critic coming off the center")
stride = 12 # max( int(len(accuracy_a) / 8), 1)
fig=plt.figure(figsize=(4,3))
LAMBDARANK_DATA = DATA['ml-20m']['VAE']['lambdarank_critic']['ALTERNATIVE_ERROR']['ndcg']
plt.axvline(x=149, linewidth=1.5, color='gray', linestyle="--")
plt.plot(range(149, 200), [DATA['ml-20m']['VAE']['actor']['ACTOR']['ndcg'][150]] + DATA['ml-20m']['VAE']['critic']['AC']['ndcg'][:50], label='RaCT', marker= 's', color='r', markevery=stride,lw=2, mec='k', mew=1 , markersize=8)
plt.plot(range(200), DATA['ml-20m']['VAE']['actor']['ACTOR']['ndcg'], '-', label='VAE', color='k', markevery=stride,lw=1.6, mec='k', mew=1 , markersize=10)
### COMMENT THIS OUT TO GET NO WARP DATA IN PLOT ###
plt.plot(range(149, 200), [DATA['ml-20m']['VAE']['actor']['ACTOR']['ndcg'][150]] + DATA['ml-20m']['VAE']['warp_critic']['ALTERNATIVE_ERROR']['ndcg'][:50], label='WARP', marker= 's', color='g', markevery=stride,lw=2, mec='k', mew=1 , markersize=8)
plt.plot(range(149, 150+len(LAMBDARANK_DATA)), [DATA['ml-20m']['VAE']['actor']['ACTOR']['ndcg'][150]] + LAMBDARANK_DATA, label='LambdaRank', marker= 's', color='b', markevery=stride,lw=2, mec='k', mew=1 , markersize=8)
plt.title("ML-20M", fontsize=13)
# CHANGE THIS IF WE REMOVE SOME OF THE LEGEND
# leg = plt.legend(fontsize=13, shadow=True, loc=(0.002, 0.69))
leg = plt.legend(fontsize=13, shadow=True, loc=(0.002, 0.51))
plt.grid('on')
plt.xlabel('# Epoch of actor', fontsize=13)
plt.ylabel('Validation NDCG@100', fontsize=13)
### SWITCH "YTICKS" line TO GET NON-WARP AXES ###
plt.yticks((0.424, 0.428, 0.432, 0.436, 0.440), ('0.424', '0.428', '0.432', '0.436', '0.440'), color='k', size=10)
# plt.yticks((0.300, 0.333, 0.366, 0.400, 0.433), ('0.300', '0.333', '0.366', '0.400', '0.433'), color='k', size=10)
plt.xlim(50, 200)
### SWITCH "YLIM" LINE TO GET NON-WARP AXES ###
plt.ylim(0.424, 0.441)
# plt.ylim(0.300, 0.451)
plt.show()
fig.savefig('improve_ndcg_ml-20m.pdf', bbox_inches='tight')
# +
print("Now we'll add the critic coming off the center")
fig=plt.figure(figsize=(4,3))
plt.axvline(x=74, linewidth=1.5, color='gray', linestyle="--")
plt.plot(range(74, 100), [DATA['netflix-prize']['VAE']['actor']['ACTOR']['ndcg'][75]] + DATA['netflix-prize']['VAE']['critic']['AC']['ndcg'][:25], label='RaCT', marker= 's', color='r', markevery=stride,lw=2, mec='k', mew=1 , markersize=8)
plt.plot(range(100), DATA['netflix-prize']['VAE']['actor']['ACTOR']['ndcg'], '-', label='VAE', color='k', markevery=stride,lw=1.6, mec='k', mew=1 , markersize=10)
### COMMENT THIS OUT TO GET NO WARP DATA IN PLOT ###
plt.plot(range(74, 100), [DATA['netflix-prize']['VAE']['actor']['ACTOR']['ndcg'][75]] + DATA['netflix-prize']['VAE']['warp_critic']['ALTERNATIVE_ERROR']['ndcg'][:25], label='warp', marker= 's', color='g', markevery=stride,lw=2, mec='k', mew=1 , markersize=8)
lambdarank_netflix = DATA['netflix-prize']['VAE']['lambdarank_critic']['ALTERNATIVE_ERROR']['ndcg']
plt.plot(range(74, 75+len(lambdarank_netflix)), [DATA['netflix-prize']['VAE']['actor']['ACTOR']['ndcg'][75]] + lambdarank_netflix, label='LambdaRank', marker= 's', color='b', markevery=stride,lw=2, mec='k', mew=1 , markersize=8)
print(f"Number of lambdarank netflixes: {len(lambdarank_netflix)}")
plt.title("Netflix", fontsize=13)
# plt.title("NDCG@100 on netflix-prize, with/without critic", fontsize=20)
leg = plt.legend(fontsize=13, shadow=True, loc=(0.002, 0.69))
plt.grid('on')
plt.xlabel('# Epoch of actor', fontsize=13)
plt.ylabel('Validation NDCG@100', fontsize=13)
plt.yticks((0.38, 0.383, 0.386, 0.389, 0.392), ('0.38', '0.383', '0.386', '0.389', '0.392'), color='k', size=10)
### SWITCH "YTICKS" line TO GET NON-WARP AXES ###
plt.yticks((0.200, 0.250, 0.300, 0.350, 0.400), ('0.200', '0.250', '0.300', '0.350', '0.400'), color='k', size=10)
plt.xlim(15, 100)
### SWITCH "YLIM" LINE TO GET NON-WARP AXES ###
# plt.ylim(0.379, 0.392)
plt.ylim(0.200, 0.400)
plt.show()
fig.savefig('improve_ndcg_netflix.pdf', bbox_inches='tight')
# +
# MSD IS DIFFERENT, BECAUSE ITS DOWNLOADED FROM TENSORBOARD....
with open("./data/msd/VAE/actor_training.json", "r") as f:
first_part = json.loads(f.read())
with open("./data/msd/VAE/critic_training.json", "r") as f:
second_part = json.loads(f.read())
with open("./data/msd/VAE/warp_critic_training.json", "r") as f:
warp_critic_msd = json.loads(f.read())['ALTERNATIVE_ERROR']['ndcg']
with open("./data/msd/VAE/lambdarank_critic_training.json", "r") as f:
lambdarank_critic_msd = json.loads(f.read())['ALTERNATIVE_ERROR']['ndcg']
first_part = [e[2] for e in first_part]
second_part = [e[2] for e in second_part]
print(len(warp_critic_msd))
print(len(lambdarank_critic_msd))
# +
print("Now we'll add the critic coming off the center")
fig=plt.figure(figsize=(4,3))
plt.axvline(x=74, linewidth=1.5, color='gray', linestyle="--")
plt.plot(range(74, 100), [first_part[75]] + second_part[50:75], label='RaCT', marker= 's', color='r', markevery=stride,lw=2, mec='k', mew=1 , markersize=8)
plt.plot(range(100), first_part, label='VAE', color='k', markevery=stride,lw=1.6, mec='k', mew=1 , markersize=10)
plt.plot(range(74, 75 + len(warp_critic_msd)), [first_part[75]] + warp_critic_msd, label='WARP', marker= 's', color='g', markevery=stride,lw=2, mec='k', mew=1 , markersize=8)
plt.plot(range(74, 75 + len(lambdarank_critic_msd)), [first_part[75]] + lambdarank_critic_msd, label='WARP', marker= 's', color='b', markevery=stride,lw=2, mec='k', mew=1 , markersize=8)
# plt.ylim(0.37, 0.40)
# plt.title("NDCG@100 on MSD, with/without critic", fontsize=20)
plt.title("MSD", fontsize=13)
leg = plt.legend(fontsize=13, shadow=True, loc=(0.002, 0.69))
plt.grid('on')
plt.xlabel('# Epoch of actor', fontsize=13)
plt.ylabel('Validation NDCG@100', fontsize=13)
plt.xlim(15, 100)
# plt.ylim(0.30, 0.32)
plt.ylim(0.25, 0.32)
plt.show()
fig.savefig('improve_ndcg_msd.pdf', bbox_inches='tight')
# -
| paper_plots/MAIN_effect_of_critic/plot_improvement_vae.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Unified XKCD Colors
#
# 
#
# A few years ago, <NAME>, the creator of the brilliant [XKCD](https://www.xkcd.com/) webcomic
# was sucked into the named color swamp. His adventures in the swamp are described in
# [this entertaining blog post](https://blog.xkcd.com/2010/05/03/color-survey-results/). The "product"
# that emerged from his time in the swamp is a long list of "named colors."
#
# Named color lists are common. I used a list of 147 web colors for my Python `SmugPyter` [dominant image color program](https://github.com/bakerjd99/smugpyter). When putting together `SmugPyter` I wondered
# if 147 colors were enough and if they were evenly distributed over the RGB color space.
# I looked around for longer lists but didn't come up with anything until I stumbled on XKCD colors.
#
# XKCD colors were derived for a crowdsourcing experiment. Munroe asked web volunteers to "name" colors
# and then compiled the names. The result is both comprehensive and delightful. Some of the color names are:
# "poo", "baby shit brown", "barney purple", "booger", "diarrhea", "milk chocolate", "shit brown", and "toxic green."
# I don't know about you but given a choice between a politically correct inoffensive color list like
# [CSS3](https://www.cssportal.com/css3-color-names/) that reads like it was composed by transgendered
# EU bureaucrats on loan to the Canadian approved euphemism agency, and a manly up yours list that considers "diarrhea" a color
# well, this mainly manly man knows which list he'll choose.
#
# However, before we dump [X11](https://www.w3schools.com/colors/colors_x11.asp),
# [CSS3](https://www.w3schools.com/cssref/css_colors.asp), [HTML4](https://www.w3schools.com/html/html_colors.asp) and other girly-man color lists we'll have to do a little hacking.
# ### Let's cannibalize `webcolors.py`
#
# Our hack begins by tearing apart some open source named color code. I want
# `SmugPyter` to use XKCD colors instead of standard lists so I am starting with
# the Python module that `SmugPyter` imports to process named colors. I briefly considered
# forking `webcolors.py`, making my changes, and then issuing a pull request to have
# XKCD colors added as an official part of `webcolors.py` but that sounds like work and
# this is for fun. Besides, in every large program there's a small program screaming to get out.
# Let's just grab the germane bits of `webcolors.py` and XKCD'ify.
#
# From my poking around the following parts of `webcolors.py` are need to process XKCD colors.
# +
# XKCD color handling
import collections
import re
from math import sqrt
# tests and displays
import numpy as np
from PIL import Image
import random
# kmeans clustering
import scipy
import scipy.misc
import scipy.cluster
from sklearn.cluster import KMeans
# paths & files
import glob
import os
import csv
# call J from python
import jbase as j
print(j.__doc__)
# +
# NOTE: the code in this cell has been modified from the original source:
# https://github.com/ubernostrum/webcolors
HEX_COLOR_RE = re.compile(r'^#([a-fA-F0-9]{3}|[a-fA-F0-9]{6})$')
IntegerRGB = collections.namedtuple('IntegerRGB', ['red', 'green', 'blue'])
def _normalize_integer_rgb(value):
"""
Internal normalization function for clipping integer values into
the permitted range (0-255, inclusive).
"""
return 0 if value < 0 \
else 255 if value > 255 \
else value
def normalize_integer_triplet(rgb_triplet):
"""
Normalize an integer ``rgb()`` triplet so that all values are
within the range 0-255 inclusive.
"""
return IntegerRGB._make(
_normalize_integer_rgb(value) for value in rgb_triplet
)
def _reversedict(d):
"""
Internal helper for generating reverse mappings; given a
dictionary, returns a new dictionary with keys and values swapped.
"""
return {value: key for key, value in d.items()}
def normalize_hex(hex_value):
"""
Normalize a hexadecimal color value to 6 digits, lowercase.
"""
match = HEX_COLOR_RE.match(hex_value)
if match is None:
raise ValueError(
u"'{}' is not a valid hexadecimal color value.".format(hex_value)
)
hex_digits = match.group(1)
if len(hex_digits) == 3:
hex_digits = u''.join(2 * s for s in hex_digits)
return u'#{}'.format(hex_digits.lower())
def hex_to_rgb(hex_value):
"""
Convert a hexadecimal color value to a 3-tuple of integers
suitable for use in an ``rgb()`` triplet specifying that color.
"""
hex_value = normalize_hex(hex_value)
hex_value = int(hex_value[1:], 16)
return IntegerRGB(
hex_value >> 16,
hex_value >> 8 & 0xff,
hex_value & 0xff
)
def normalize_integer_triplet(rgb_triplet):
"""
Normalize an integer ``rgb()`` triplet so that all values are
within the range 0-255 inclusive.
"""
return IntegerRGB._make(
_normalize_integer_rgb(value) for value in rgb_triplet
)
def hex_to_name(hex_value):
"""
Convert a hexadecimal color value to its corresponding normalized
color name, if any such name exists.
When no color name for the value is found the result is None.
"""
normalized = normalize_hex(hex_value)
name = XKCD_HEX_TO_NAMES.get(normalized)
return name
def rgb_to_hex(rgb_triplet):
"""
Convert a 3-tuple of integers, suitable for use in an ``rgb()``
color triplet, to a normalized hexadecimal value for that color.
"""
return u'#{:02x}{:02x}{:02x}'.format(
*normalize_integer_triplet(
rgb_triplet
)
)
def rgb_to_name(rgb_triplet):
"""
Convert a 3-tuple of integers, suitable for use in an ``rgb()``
color triplet, to its corresponding normalized color name, if any
such name exists.
If there is no matching name None is returned
"""
return hex_to_name(
rgb_to_hex(
normalize_integer_triplet(
rgb_triplet
)
)
)
# -
# ### Unified XKCD Color Dictionary
#
# `webcolors.py` also contains long Python dictionary definitions that map names to colors coded as hexidecimals.
# +
CSS3_NAMES_TO_HEX = {
u'aliceblue': u'#f0f8ff',
u'antiquewhite': u'#faebd7',
u'aqua': u'#00ffff',
u'aquamarine': u'#7fffd4',
u'azure': u'#f0ffff',
u'beige': u'#f5f5dc',
u'bisque': u'#ffe4c4',
u'black': u'#000000',
u'blanchedalmond': u'#ffebcd',
u'blue': u'#0000ff',
u'blueviolet': u'#8a2be2',
u'brown': u'#a52a2a',
u'burlywood': u'#deb887',
u'cadetblue': u'#5f9ea0',
u'chartreuse': u'#7fff00',
u'chocolate': u'#d2691e',
u'coral': u'#ff7f50',
u'cornflowerblue': u'#6495ed',
u'cornsilk': u'#fff8dc',
u'crimson': u'#dc143c',
u'cyan': u'#00ffff',
u'darkblue': u'#00008b',
u'darkcyan': u'#008b8b',
u'darkgoldenrod': u'#b8860b',
u'darkgray': u'#a9a9a9',
u'darkgrey': u'#a9a9a9',
u'darkgreen': u'#006400',
u'darkkhaki': u'#bdb76b',
u'darkmagenta': u'#8b008b',
u'darkolivegreen': u'#556b2f',
u'darkorange': u'#ff8c00',
u'darkorchid': u'#9932cc',
u'darkred': u'#8b0000',
u'darksalmon': u'#e9967a',
u'darkseagreen': u'#8fbc8f',
u'darkslateblue': u'#483d8b',
u'darkslategray': u'#2f4f4f',
u'darkslategrey': u'#2f4f4f',
u'darkturquoise': u'#00ced1',
u'darkviolet': u'#9400d3',
u'deeppink': u'#ff1493',
u'deepskyblue': u'#00bfff',
u'dimgray': u'#696969',
u'dimgrey': u'#696969',
u'dodgerblue': u'#1e90ff',
u'firebrick': u'#b22222',
u'floralwhite': u'#fffaf0',
u'forestgreen': u'#228b22',
u'fuchsia': u'#ff00ff',
u'gainsboro': u'#dcdcdc',
u'ghostwhite': u'#f8f8ff',
u'gold': u'#ffd700',
u'goldenrod': u'#daa520',
u'gray': u'#808080',
u'grey': u'#808080',
u'green': u'#008000',
u'greenyellow': u'#adff2f',
u'honeydew': u'#f0fff0',
u'hotpink': u'#ff69b4',
u'indianred': u'#cd5c5c',
u'indigo': u'#4b0082',
u'ivory': u'#fffff0',
u'khaki': u'#f0e68c',
u'lavender': u'#e6e6fa',
u'lavenderblush': u'#fff0f5',
u'lawngreen': u'#7cfc00',
u'lemonchiffon': u'#fffacd',
u'lightblue': u'#add8e6',
u'lightcoral': u'#f08080',
u'lightcyan': u'#e0ffff',
u'lightgoldenrodyellow': u'#fafad2',
u'lightgray': u'#d3d3d3',
u'lightgrey': u'#d3d3d3',
u'lightgreen': u'#90ee90',
u'lightpink': u'#ffb6c1',
u'lightsalmon': u'#ffa07a',
u'lightseagreen': u'#20b2aa',
u'lightskyblue': u'#87cefa',
u'lightslategray': u'#778899',
u'lightslategrey': u'#778899',
u'lightsteelblue': u'#b0c4de',
u'lightyellow': u'#ffffe0',
u'lime': u'#00ff00',
u'limegreen': u'#32cd32',
u'linen': u'#faf0e6',
u'magenta': u'#ff00ff',
u'maroon': u'#800000',
u'mediumaquamarine': u'#66cdaa',
u'mediumblue': u'#0000cd',
u'mediumorchid': u'#ba55d3',
u'mediumpurple': u'#9370db',
u'mediumseagreen': u'#3cb371',
u'mediumslateblue': u'#7b68ee',
u'mediumspringgreen': u'#00fa9a',
u'mediumturquoise': u'#48d1cc',
u'mediumvioletred': u'#c71585',
u'midnightblue': u'#191970',
u'mintcream': u'#f5fffa',
u'mistyrose': u'#ffe4e1',
u'moccasin': u'#ffe4b5',
u'navajowhite': u'#ffdead',
u'navy': u'#000080',
u'oldlace': u'#fdf5e6',
u'olive': u'#808000',
u'olivedrab': u'#6b8e23',
u'orange': u'#ffa500',
u'orangered': u'#ff4500',
u'orchid': u'#da70d6',
u'palegoldenrod': u'#eee8aa',
u'palegreen': u'#98fb98',
u'paleturquoise': u'#afeeee',
u'palevioletred': u'#db7093',
u'papayawhip': u'#ffefd5',
u'peachpuff': u'#ffdab9',
u'peru': u'#cd853f',
u'pink': u'#ffc0cb',
u'plum': u'#dda0dd',
u'powderblue': u'#b0e0e6',
u'purple': u'#800080',
u'red': u'#ff0000',
u'rosybrown': u'#bc8f8f',
u'royalblue': u'#4169e1',
u'saddlebrown': u'#8b4513',
u'salmon': u'#fa8072',
u'sandybrown': u'#f4a460',
u'seagreen': u'#2e8b57',
u'seashell': u'#fff5ee',
u'sienna': u'#a0522d',
u'silver': u'#c0c0c0',
u'skyblue': u'#87ceeb',
u'slateblue': u'#6a5acd',
u'slategray': u'#708090',
u'slategrey': u'#708090',
u'snow': u'#fffafa',
u'springgreen': u'#00ff7f',
u'steelblue': u'#4682b4',
u'tan': u'#d2b48c',
u'teal': u'#008080',
u'thistle': u'#d8bfd8',
u'tomato': u'#ff6347',
u'turquoise': u'#40e0d0',
u'violet': u'#ee82ee',
u'wheat': u'#f5deb3',
u'white': u'#ffffff',
u'whitesmoke': u'#f5f5f5',
u'yellow': u'#ffff00',
u'yellowgreen': u'#9acd32',
}
# reverse dictionary
CSS3_HEX_TO_NAMES = _reversedict(CSS3_NAMES_TO_HEX)
len(CSS3_HEX_TO_NAMES)
# -
# To use XKCD colors add a similiar dictionary. The following defines what I call
# **Unified XKCD Colors.**. It was derived by merging `webcolors`, `CSS3` colors
# and XKCD colors. I removed duplicate names and duplicate codes. The result is
# a list of 970 uniquely named colors.
# +
# License: http://creativecommons.org/publicdomain/zero/1.0/
XKCD_NAMES_TO_HEX = {
u'acidgreen': u'#8ffe09',
u'adobe': u'#bd6c48',
u'algae': u'#54ac68',
u'algaegreen': u'#21c36f',
u'aliceblue': u'#f0f8ff',
u'almostblack': u'#070d0d',
u'amber': u'#feb308',
u'amethyst': u'#9b5fc0',
u'antiquewhite': u'#faebd7',
u'apple': u'#6ecb3c',
u'applegreen': u'#76cd26',
u'apricot': u'#ffb16d',
u'aqua': u'#00ffff',
u'aquablue': u'#02d8e9',
u'aquagreen': u'#12e193',
u'aquamarine': u'#7fffd4',
u'armygreen': u'#4b5d16',
u'asparagus': u'#77ab56',
u'aubergine': u'#3d0734',
u'auburn': u'#9a3001',
u'avocado': u'#90b134',
u'avocadogreen': u'#87a922',
u'azul': u'#1d5dec',
u'azure': u'#f0ffff',
u'babyblue': u'#a2cffe',
u'babygreen': u'#8cff9e',
u'babypink': u'#ffb7ce',
u'babypoo': u'#ab9004',
u'babypoop': u'#937c00',
u'babypoopgreen': u'#8f9805',
u'babypukegreen': u'#b6c406',
u'babypurple': u'#ca9bf7',
u'babyshitbrown': u'#ad900d',
u'babyshitgreen': u'#889717',
u'banana': u'#ffff7e',
u'bananayellow': u'#fafe4b',
u'barbiepink': u'#fe46a5',
u'barfgreen': u'#94ac02',
u'barney': u'#ac1db8',
u'barneypurple': u'#a00498',
u'battleshipgrey': u'#6b7c85',
u'beige': u'#f5f5dc',
u'berry': u'#990f4b',
u'bile': u'#b5c306',
u'bisque': u'#ffe4c4',
u'black': u'#000000',
u'blanchedalmond': u'#ffebcd',
u'bland': u'#afa88b',
u'blood': u'#770001',
u'bloodorange': u'#fe4b03',
u'bloodred': u'#980002',
u'blue': u'#0000ff',
u'blueberry': u'#464196',
u'blueblue': u'#2242c7',
u'bluegreen': u'#017a79',
u'bluegrey': u'#607c8e',
u'bluepurple': u'#5729ce',
u'blueviolet': u'#8a2be2',
u'bluewithahintofpurple': u'#533cc6',
u'blueygreen': u'#2bb179',
u'blueygrey': u'#89a0b0',
u'blueypurple': u'#6241c7',
u'bluish': u'#2976bb',
u'bluishgreen': u'#10a674',
u'bluishgrey': u'#748b97',
u'bluishpurple': u'#703be7',
u'blurple': u'#5539cc',
u'blush': u'#f29e8e',
u'blushpink': u'#fe828c',
u'booger': u'#9bb53c',
u'boogergreen': u'#96b403',
u'bordeaux': u'#7b002c',
u'boringgreen': u'#63b365',
u'bottlegreen': u'#044a05',
u'brick': u'#a03623',
u'brickorange': u'#c14a09',
u'brickred': u'#8f1402',
u'brightaqua': u'#0bf9ea',
u'brightblue': u'#0165fc',
u'brightcyan': u'#41fdfe',
u'brightgreen': u'#01ff07',
u'brightlavender': u'#c760ff',
u'brightlightblue': u'#26f7fd',
u'brightlightgreen': u'#2dfe54',
u'brightlilac': u'#c95efb',
u'brightlime': u'#87fd05',
u'brightlimegreen': u'#65fe08',
u'brightmagenta': u'#ff08e8',
u'brightolive': u'#9cbb04',
u'brightorange': u'#ff5b00',
u'brightpink': u'#fe01b1',
u'brightpurple': u'#be03fd',
u'brightred': u'#ff000d',
u'brightseagreen': u'#05ffa6',
u'brightskyblue': u'#02ccfe',
u'brightteal': u'#01f9c6',
u'brightturquoise': u'#0ffef9',
u'brightviolet': u'#ad0afd',
u'brightyellow': u'#fffd01',
u'brightyellowgreen': u'#9dff00',
u'britishracinggreen': u'#05480d',
u'bronze': u'#a87900',
u'brown': u'#a52a2a',
u'browngreen': u'#706c11',
u'browngrey': u'#8d8468',
u'brownish': u'#9c6d57',
u'brownishgreen': u'#6a6e09',
u'brownishgrey': u'#86775f',
u'brownishorange': u'#cb7723',
u'brownishpink': u'#c27e79',
u'brownishpurple': u'#76424e',
u'brownishred': u'#9e3623',
u'brownishyellow': u'#c9b003',
u'brownorange': u'#b96902',
u'brownred': u'#922b05',
u'brownyellow': u'#b29705',
u'brownygreen': u'#6f6c0a',
u'brownyorange': u'#ca6b02',
u'bruise': u'#7e4071',
u'bubblegum': u'#ff6cb5',
u'bubblegumpink': u'#fe83cc',
u'buff': u'#fef69e',
u'burgundy': u'#610023',
u'burlywood': u'#deb887',
u'burntorange': u'#c04e01',
u'burntred': u'#9f2305',
u'burntsiena': u'#b75203',
u'burntsienna': u'#b04e0f',
u'burntumber': u'#a0450e',
u'burntyellow': u'#d5ab09',
u'burple': u'#6832e3',
u'butter': u'#ffff81',
u'butterscotch': u'#fdb147',
u'butteryellow': u'#fffd74',
u'cadetblue': u'#5f9ea0',
u'camel': u'#c69f59',
u'camo': u'#7f8f4e',
u'camogreen': u'#526525',
u'camouflagegreen': u'#4b6113',
u'canary': u'#fdff63',
u'canaryyellow': u'#fffe40',
u'candypink': u'#ff63e9',
u'caramel': u'#af6f09',
u'carmine': u'#9d0216',
u'carnation': u'#fd798f',
u'carnationpink': u'#ff7fa7',
u'carolinablue': u'#8ab8fe',
u'celadon': u'#befdb7',
u'celery': u'#c1fd95',
u'cement': u'#a5a391',
u'cerise': u'#de0c62',
u'cerulean': u'#0485d1',
u'ceruleanblue': u'#056eee',
u'charcoal': u'#343837',
u'charcoalgrey': u'#3c4142',
u'chartreuse': u'#7fff00',
u'cherry': u'#cf0234',
u'cherryred': u'#f7022a',
u'chestnut': u'#742802',
u'chocolate': u'#d2691e',
u'chocolatebrown': u'#411900',
u'cinnamon': u'#ac4f06',
u'claret': u'#680018',
u'clay': u'#b66a50',
u'claybrown': u'#b2713d',
u'clearblue': u'#247afd',
u'cloudyblue': u'#acc2d9',
u'cobalt': u'#1e488f',
u'cobaltblue': u'#030aa7',
u'cocoa': u'#875f42',
u'coffee': u'#a6814c',
u'coolblue': u'#4984b8',
u'coolgreen': u'#33b864',
u'coolgrey': u'#95a3a6',
u'copper': u'#b66325',
u'coral': u'#ff7f50',
u'coralpink': u'#ff6163',
u'cornflower': u'#6a79f7',
u'cornflowerblue': u'#6495ed',
u'cornsilk': u'#fff8dc',
u'cranberry': u'#9e003a',
u'cream': u'#ffffc2',
u'creme': u'#ffffb6',
u'crimson': u'#dc143c',
u'custard': u'#fffd78',
u'dandelion': u'#fedf08',
u'dark': u'#1b2431',
u'darkaqua': u'#05696b',
u'darkaquamarine': u'#017371',
u'darkbeige': u'#ac9362',
u'darkblue': u'#00008b',
u'darkbluegreen': u'#005249',
u'darkbluegrey': u'#1f3b4d',
u'darkbrown': u'#341c02',
u'darkcoral': u'#cf524e',
u'darkcream': u'#fff39a',
u'darkcyan': u'#008b8b',
u'darkforestgreen': u'#002d04',
u'darkfuchsia': u'#9d0759',
u'darkgold': u'#b59410',
u'darkgoldenrod': u'#b8860b',
u'darkgrassgreen': u'#388004',
u'darkgray': u'#a9a9a9',
u'darkgreen': u'#006400',
u'darkgreenblue': u'#1f6357',
u'darkgrey': u'#363737',
u'darkgreyblue': u'#29465b',
u'darkhotpink': u'#d90166',
u'darkindigo': u'#1f0954',
u'darkishblue': u'#014182',
u'darkishgreen': u'#287c37',
u'darkishpink': u'#da467d',
u'darkishpurple': u'#751973',
u'darkishred': u'#a90308',
u'darkkhaki': u'#bdb76b',
u'darklavender': u'#856798',
u'darklilac': u'#9c6da5',
u'darklime': u'#84b701',
u'darklimegreen': u'#7ebd01',
u'darkmagenta': u'#8b008b',
u'darkmaroon': u'#3c0008',
u'darkmauve': u'#874c62',
u'darkmint': u'#48c072',
u'darkmintgreen': u'#20c073',
u'darkmustard': u'#a88905',
u'darknavy': u'#000435',
u'darknavyblue': u'#00022e',
u'darkolive': u'#373e02',
u'darkolivegreen': u'#556b2f',
u'darkorange': u'#ff8c00',
u'darkorchid': u'#9932cc',
u'darkpastelgreen': u'#56ae57',
u'darkpeach': u'#de7e5d',
u'darkperiwinkle': u'#665fd1',
u'darkpink': u'#cb416b',
u'darkplum': u'#3f012c',
u'darkpurple': u'#35063e',
u'darkred': u'#8b0000',
u'darkrose': u'#b5485d',
u'darkroyalblue': u'#02066f',
u'darksage': u'#598556',
u'darksalmon': u'#e9967a',
u'darksand': u'#a88f59',
u'darkseafoam': u'#1fb57a',
u'darkseafoamgreen': u'#3eaf76',
u'darkseagreen': u'#8fbc8f',
u'darkskyblue': u'#448ee4',
u'darkslateblue': u'#483d8b',
u'darkslategray': u'#2f4f4f',
u'darktan': u'#af884a',
u'darktaupe': u'#7f684e',
u'darkteal': u'#014d4e',
u'darkturquoise': u'#00ced1',
u'darkviolet': u'#9400d3',
u'darkyellow': u'#d5b60a',
u'darkyellowgreen': u'#728f02',
u'deepaqua': u'#08787f',
u'deepblue': u'#040273',
u'deepbrown': u'#410200',
u'deepgreen': u'#02590f',
u'deeplavender': u'#8d5eb7',
u'deeplilac': u'#966ebd',
u'deepmagenta': u'#a0025c',
u'deeporange': u'#dc4d01',
u'deeppink': u'#ff1493',
u'deeppurple': u'#36013f',
u'deepred': u'#9a0200',
u'deeprose': u'#c74767',
u'deepseablue': u'#015482',
u'deepskyblue': u'#00bfff',
u'deepteal': u'#00555a',
u'deepturquoise': u'#017374',
u'deepviolet': u'#490648',
u'denim': u'#3b638c',
u'denimblue': u'#3b5b92',
u'desert': u'#ccad60',
u'diarrhea': u'#9f8303',
u'dimgray': u'#696969',
u'dirt': u'#8a6e45',
u'dirtbrown': u'#836539',
u'dirtyblue': u'#3f829d',
u'dirtygreen': u'#667e2c',
u'dirtyorange': u'#c87606',
u'dirtypink': u'#ca7b80',
u'dirtypurple': u'#734a65',
u'dirtyyellow': u'#cdc50a',
u'dodgerblue': u'#1e90ff',
u'drab': u'#828344',
u'drabgreen': u'#749551',
u'driedblood': u'#4b0101',
u'duckeggblue': u'#c3fbf4',
u'dullblue': u'#49759c',
u'dullbrown': u'#876e4b',
u'dullgreen': u'#74a662',
u'dullorange': u'#d8863b',
u'dullpink': u'#d5869d',
u'dullpurple': u'#84597e',
u'dullred': u'#bb3f3f',
u'dullteal': u'#5f9e8f',
u'dullyellow': u'#eedc5b',
u'dusk': u'#4e5481',
u'duskblue': u'#26538d',
u'duskyblue': u'#475f94',
u'duskypink': u'#cc7a8b',
u'duskypurple': u'#895b7b',
u'duskyrose': u'#ba6873',
u'dust': u'#b2996e',
u'dustyblue': u'#5a86ad',
u'dustygreen': u'#76a973',
u'dustylavender': u'#ac86a8',
u'dustyorange': u'#f0833a',
u'dustypink': u'#d58a94',
u'dustypurple': u'#825f87',
u'dustyred': u'#b9484e',
u'dustyrose': u'#c0737a',
u'dustyteal': u'#4c9085',
u'earth': u'#a2653e',
u'eastergreen': u'#8cfd7e',
u'easterpurple': u'#c071fe',
u'ecru': u'#feffca',
u'eggplant': u'#380835',
u'eggplantpurple': u'#430541',
u'eggshell': u'#fffcc4',
u'eggshellblue': u'#c4fff7',
u'electricblue': u'#0652ff',
u'electricgreen': u'#21fc0d',
u'electriclime': u'#a8ff04',
u'electricpink': u'#ff0490',
u'electricpurple': u'#aa23ff',
u'emerald': u'#01a049',
u'emeraldgreen': u'#028f1e',
u'evergreen': u'#05472a',
u'fadedblue': u'#658cbb',
u'fadedgreen': u'#7bb274',
u'fadedorange': u'#f0944d',
u'fadedpink': u'#de9dac',
u'fadedpurple': u'#916e99',
u'fadedred': u'#d3494e',
u'fadedyellow': u'#feff7f',
u'fawn': u'#cfaf7b',
u'fern': u'#63a950',
u'ferngreen': u'#548d44',
u'firebrick': u'#b22222',
u'fireenginered': u'#fe0002',
u'flatblue': u'#3c73a8',
u'flatgreen': u'#699d4c',
u'floralwhite': u'#fffaf0',
u'fluorescentgreen': u'#08ff08',
u'flurogreen': u'#0aff02',
u'foamgreen': u'#90fda9',
u'forest': u'#0b5509',
u'forestgreen': u'#228b22',
u'forrestgreen': u'#154406',
u'frenchblue': u'#436bad',
u'freshgreen': u'#69d84f',
u'froggreen': u'#58bc08',
u'fuchsia': u'#ff00ff',
u'gainsboro': u'#dcdcdc',
u'ghostwhite': u'#f8f8ff',
u'gold': u'#ffd700',
u'golden': u'#f5bf03',
u'goldenbrown': u'#b27a01',
u'goldenrod': u'#daa520',
u'goldenyellow': u'#fec615',
u'grape': u'#6c3461',
u'grapefruit': u'#fd5956',
u'grapepurple': u'#5d1451',
u'grass': u'#5cac2d',
u'grassgreen': u'#3f9b0b',
u'grassygreen': u'#419c03',
u'gray': u'#808080',
u'green': u'#008000',
u'greenapple': u'#5edc1f',
u'greenblue': u'#01c08d',
u'greenbrown': u'#544e03',
u'greengrey': u'#77926f',
u'greenish': u'#40a368',
u'greenishbeige': u'#c9d179',
u'greenishblue': u'#0b8b87',
u'greenishbrown': u'#696112',
u'greenishcyan': u'#2afeb7',
u'greenishgrey': u'#96ae8d',
u'greenishtan': u'#bccb7a',
u'greenishteal': u'#32bf84',
u'greenishturquoise': u'#00fbb0',
u'greenishyellow': u'#cdfd02',
u'greenteal': u'#0cb577',
u'greenyblue': u'#42b395',
u'greenybrown': u'#696006',
u'greenyellow': u'#adff2f',
u'greenygrey': u'#7ea07a',
u'greenyyellow': u'#c6f808',
u'grey': u'#929591',
u'greyblue': u'#647d8e',
u'greybrown': u'#7f7053',
u'greygreen': u'#789b73',
u'greyish': u'#a8a495',
u'greyishblue': u'#5e819d',
u'greyishbrown': u'#7a6a4f',
u'greyishgreen': u'#82a67d',
u'greyishpink': u'#c88d94',
u'greyishpurple': u'#887191',
u'greyishteal': u'#719f91',
u'greypink': u'#c3909b',
u'greypurple': u'#826d8c',
u'greyteal': u'#5e9b8a',
u'grossgreen': u'#a0bf16',
u'gunmetal': u'#536267',
u'hazel': u'#8e7618',
u'heather': u'#a484ac',
u'heliotrope': u'#d94ff5',
u'highlightergreen': u'#1bfc06',
u'honeydew': u'#f0fff0',
u'hospitalgreen': u'#9be5aa',
u'hotgreen': u'#25ff29',
u'hotmagenta': u'#f504c9',
u'hotpink': u'#ff69b4',
u'hotpurple': u'#cb00f5',
u'huntergreen': u'#0b4008',
u'ice': u'#d6fffa',
u'iceblue': u'#d7fffe',
u'ickygreen': u'#8fae22',
u'indianred': u'#cd5c5c',
u'indigo': u'#4b0082',
u'indigoblue': u'#3a18b1',
u'iris': u'#6258c4',
u'irishgreen': u'#019529',
u'ivory': u'#fffff0',
u'jade': u'#1fa774',
u'jadegreen': u'#2baf6a',
u'junglegreen': u'#048243',
u'kelleygreen': u'#009337',
u'kellygreen': u'#02ab2e',
u'kermitgreen': u'#5cb200',
u'keylime': u'#aeff6e',
u'khaki': u'#f0e68c',
u'khakigreen': u'#728639',
u'kiwi': u'#9cef43',
u'kiwigreen': u'#8ee53f',
u'lavender': u'#e6e6fa',
u'lavenderblue': u'#8b88f8',
u'lavenderblush': u'#fff0f5',
u'lavenderpink': u'#dd85d7',
u'lawngreen': u'#7cfc00',
u'leaf': u'#71aa34',
u'leafgreen': u'#5ca904',
u'leafygreen': u'#51b73b',
u'leather': u'#ac7434',
u'lemon': u'#fdff52',
u'lemonchiffon': u'#fffacd',
u'lemongreen': u'#adf802',
u'lemonlime': u'#bffe28',
u'lemonyellow': u'#fdff38',
u'lichen': u'#8fb67b',
u'lightaqua': u'#8cffdb',
u'lightaquamarine': u'#7bfdc7',
u'lightbeige': u'#fffeb6',
u'lightblue': u'#add8e6',
u'lightbluegreen': u'#7efbb3',
u'lightbluegrey': u'#b7c9e2',
u'lightbluishgreen': u'#76fda8',
u'lightbrightgreen': u'#53fe5c',
u'lightbrown': u'#ad8150',
u'lightburgundy': u'#a8415b',
u'lightcoral': u'#f08080',
u'lightcyan': u'#e0ffff',
u'lighteggplant': u'#894585',
u'lightergreen': u'#75fd63',
u'lighterpurple': u'#a55af4',
u'lightforestgreen': u'#4f9153',
u'lightgold': u'#fddc5c',
u'lightgoldenrodyellow': u'#fafad2',
u'lightgrassgreen': u'#9af764',
u'lightgray': u'#d3d3d3',
u'lightgreen': u'#90ee90',
u'lightgreenblue': u'#56fca2',
u'lightgreenishblue': u'#63f7b4',
u'lightgrey': u'#d8dcd6',
u'lightgreyblue': u'#9dbcd4',
u'lightgreygreen': u'#b7e1a1',
u'lightindigo': u'#6d5acf',
u'lightishblue': u'#3d7afd',
u'lightishgreen': u'#61e160',
u'lightishpurple': u'#a552e6',
u'lightishred': u'#fe2f4a',
u'lightkhaki': u'#e6f2a2',
u'lightlavendar': u'#efc0fe',
u'lightlavender': u'#dfc5fe',
u'lightlightblue': u'#cafffb',
u'lightlightgreen': u'#c8ffb0',
u'lightlilac': u'#edc8ff',
u'lightlime': u'#aefd6c',
u'lightlimegreen': u'#b9ff66',
u'lightmagenta': u'#fa5ff7',
u'lightmaroon': u'#a24857',
u'lightmauve': u'#c292a1',
u'lightmint': u'#b6ffbb',
u'lightmintgreen': u'#a6fbb2',
u'lightmossgreen': u'#a6c875',
u'lightmustard': u'#f7d560',
u'lightnavy': u'#155084',
u'lightnavyblue': u'#2e5a88',
u'lightneongreen': u'#4efd54',
u'lightolive': u'#acbf69',
u'lightolivegreen': u'#a4be5c',
u'lightorange': u'#fdaa48',
u'lightpastelgreen': u'#b2fba5',
u'lightpeach': u'#ffd8b1',
u'lightpeagreen': u'#c4fe82',
u'lightperiwinkle': u'#c1c6fc',
u'lightpink': u'#ffb6c1',
u'lightplum': u'#9d5783',
u'lightpurple': u'#bf77f6',
u'lightred': u'#ff474c',
u'lightrose': u'#ffc5cb',
u'lightroyalblue': u'#3a2efe',
u'lightsage': u'#bcecac',
u'lightsalmon': u'#ffa07a',
u'lightseafoam': u'#a0febf',
u'lightseafoamgreen': u'#a7ffb5',
u'lightseagreen': u'#20b2aa',
u'lightskyblue': u'#87cefa',
u'lightslategray': u'#778899',
u'lightsteelblue': u'#b0c4de',
u'lighttan': u'#fbeeac',
u'lightteal': u'#90e4c1',
u'lightturquoise': u'#7ef4cc',
u'lighturple': u'#b36ff6',
u'lightviolet': u'#d6b4fc',
u'lightyellow': u'#ffffe0',
u'lightyellowgreen': u'#ccfd7f',
u'lightyellowishgreen': u'#c2ff89',
u'lilac': u'#cea2fd',
u'liliac': u'#c48efd',
u'lime': u'#00ff00',
u'limegreen': u'#32cd32',
u'limeyellow': u'#d0fe1d',
u'linen': u'#faf0e6',
u'lipstick': u'#d5174e',
u'lipstickred': u'#c0022f',
u'macaroniandcheese': u'#efb435',
u'magenta': u'#c20078',
u'mahogany': u'#4a0100',
u'maize': u'#f4d054',
u'mango': u'#ffa62b',
u'manilla': u'#fffa86',
u'marigold': u'#fcc006',
u'marine': u'#042e60',
u'marineblue': u'#01386a',
u'maroon': u'#800000',
u'mauve': u'#ae7181',
u'mediumaquamarine': u'#66cdaa',
u'mediumblue': u'#0000cd',
u'mediumbrown': u'#7f5112',
u'mediumgreen': u'#39ad48',
u'mediumgrey': u'#7d7f7c',
u'mediumorchid': u'#ba55d3',
u'mediumpink': u'#f36196',
u'mediumpurple': u'#9370db',
u'mediumseagreen': u'#3cb371',
u'mediumslateblue': u'#7b68ee',
u'mediumspringgreen': u'#00fa9a',
u'mediumturquoise': u'#48d1cc',
u'mediumvioletred': u'#c71585',
u'melon': u'#ff7855',
u'merlot': u'#730039',
u'metallicblue': u'#4f738e',
u'midblue': u'#276ab3',
u'midgreen': u'#50a747',
u'midnight': u'#03012d',
u'midnightblue': u'#191970',
u'midnightpurple': u'#280137',
u'militarygreen': u'#667c3e',
u'milkchocolate': u'#7f4e1e',
u'mint': u'#9ffeb0',
u'mintcream': u'#f5fffa',
u'mintgreen': u'#8fff9f',
u'mintygreen': u'#0bf77d',
u'mistyrose': u'#ffe4e1',
u'moccasin': u'#ffe4b5',
u'mocha': u'#9d7651',
u'moss': u'#769958',
u'mossgreen': u'#658b38',
u'mossygreen': u'#638b27',
u'mud': u'#735c12',
u'mudbrown': u'#60460f',
u'muddybrown': u'#886806',
u'muddygreen': u'#657432',
u'muddyyellow': u'#bfac05',
u'mudgreen': u'#606602',
u'mulberry': u'#920a4e',
u'murkygreen': u'#6c7a0e',
u'mushroom': u'#ba9e88',
u'mustard': u'#ceb301',
u'mustardbrown': u'#ac7e04',
u'mustardgreen': u'#a8b504',
u'mustardyellow': u'#d2bd0a',
u'mutedblue': u'#3b719f',
u'mutedgreen': u'#5fa052',
u'mutedpink': u'#d1768f',
u'mutedpurple': u'#805b87',
u'nastygreen': u'#70b23f',
u'navajowhite': u'#ffdead',
u'navy': u'#000080',
u'navyblue': u'#001146',
u'navygreen': u'#35530a',
u'neonblue': u'#04d9ff',
u'neongreen': u'#0cff0c',
u'neonpink': u'#fe019a',
u'neonpurple': u'#bc13fe',
u'neonred': u'#ff073a',
u'neonyellow': u'#cfff04',
u'niceblue': u'#107ab0',
u'nightblue': u'#040348',
u'ocean': u'#017b92',
u'oceanblue': u'#03719c',
u'oceangreen': u'#3d9973',
u'ocher': u'#bf9b0c',
u'ochre': u'#bf9005',
u'ocre': u'#c69c04',
u'offblue': u'#5684ae',
u'offgreen': u'#6ba353',
u'offwhite': u'#ffffe4',
u'offyellow': u'#f1f33f',
u'oldlace': u'#fdf5e6',
u'oldpink': u'#c77986',
u'oldrose': u'#c87f89',
u'olive': u'#808000',
u'olivebrown': u'#645403',
u'olivedrab': u'#6b8e23',
u'olivegreen': u'#677a04',
u'oliveyellow': u'#c2b709',
u'orange': u'#ffa500',
u'orangebrown': u'#be6400',
u'orangeish': u'#fd8d49',
u'orangepink': u'#ff6f52',
u'orangered': u'#ff4500',
u'orangeybrown': u'#b16002',
u'orangeyellow': u'#ffad01',
u'orangeyred': u'#fa4224',
u'orangeyyellow': u'#fdb915',
u'orangish': u'#fc824a',
u'orangishbrown': u'#b25f03',
u'orangishred': u'#f43605',
u'orchid': u'#da70d6',
u'pale': u'#fff9d0',
u'paleaqua': u'#b8ffeb',
u'paleblue': u'#d0fefe',
u'palebrown': u'#b1916e',
u'palecyan': u'#b7fffa',
u'palegold': u'#fdde6c',
u'palegoldenrod': u'#eee8aa',
u'palegreen': u'#98fb98',
u'palegrey': u'#fdfdfe',
u'palelavender': u'#eecffe',
u'palelightgreen': u'#b1fc99',
u'palelilac': u'#e4cbff',
u'palelime': u'#befd73',
u'palelimegreen': u'#b1ff65',
u'palemagenta': u'#d767ad',
u'palemauve': u'#fed0fc',
u'paleolive': u'#b9cc81',
u'paleolivegreen': u'#b1d27b',
u'paleorange': u'#ffa756',
u'palepeach': u'#ffe5ad',
u'palepink': u'#ffcfdc',
u'palepurple': u'#b790d4',
u'palered': u'#d9544d',
u'palerose': u'#fdc1c5',
u'palesalmon': u'#ffb19a',
u'paleskyblue': u'#bdf6fe',
u'paleteal': u'#82cbb2',
u'paleturquoise': u'#afeeee',
u'paleviolet': u'#ceaefa',
u'palevioletred': u'#db7093',
u'paleyellow': u'#ffff84',
u'papayawhip': u'#ffefd5',
u'parchment': u'#fefcaf',
u'pastelblue': u'#a2bffe',
u'pastelgreen': u'#b0ff9d',
u'pastelorange': u'#ff964f',
u'pastelpink': u'#ffbacd',
u'pastelpurple': u'#caa0ff',
u'pastelred': u'#db5856',
u'pastelyellow': u'#fffe71',
u'pea': u'#a4bf20',
u'peach': u'#ffb07c',
u'peachpuff': u'#ffdab9',
u'peachypink': u'#ff9a8a',
u'peacockblue': u'#016795',
u'peagreen': u'#8eab12',
u'pear': u'#cbf85f',
u'peasoup': u'#929901',
u'peasoupgreen': u'#94a617',
u'periwinkle': u'#8e82fe',
u'periwinkleblue': u'#8f99fb',
u'perrywinkle': u'#8f8ce7',
u'peru': u'#cd853f',
u'petrol': u'#005f6a',
u'pigpink': u'#e78ea5',
u'pine': u'#2b5d34',
u'pinegreen': u'#0a481e',
u'pink': u'#ffc0cb',
u'pinkish': u'#d46a7e',
u'pinkishbrown': u'#b17261',
u'pinkishgrey': u'#c8aca9',
u'pinkishorange': u'#ff724c',
u'pinkishpurple': u'#d648d7',
u'pinkishred': u'#f10c45',
u'pinkishtan': u'#d99b82',
u'pinkpurple': u'#db4bda',
u'pinkred': u'#f5054f',
u'pinky': u'#fc86aa',
u'pinkypurple': u'#c94cbe',
u'pinkyred': u'#fc2647',
u'pissyellow': u'#ddd618',
u'pistachio': u'#c0fa8b',
u'plum': u'#dda0dd',
u'plumpurple': u'#4e0550',
u'poisongreen': u'#40fd14',
u'poo': u'#8f7303',
u'poobrown': u'#885f01',
u'poop': u'#7f5e00',
u'poopbrown': u'#7a5901',
u'poopgreen': u'#6f7c00',
u'powderblue': u'#b0e0e6',
u'powderpink': u'#ffb2d0',
u'primaryblue': u'#0804f9',
u'prussianblue': u'#004577',
u'puce': u'#a57e52',
u'puke': u'#a5a502',
u'pukebrown': u'#947706',
u'pukegreen': u'#9aae07',
u'pukeyellow': u'#c2be0e',
u'pumpkin': u'#e17701',
u'pumpkinorange': u'#fb7d07',
u'pureblue': u'#0203e2',
u'purple': u'#800080',
u'purpleblue': u'#5d21d0',
u'purplebrown': u'#673a3f',
u'purplegrey': u'#866f85',
u'purpleish': u'#98568d',
u'purpleishblue': u'#6140ef',
u'purpleishpink': u'#df4ec8',
u'purplepink': u'#d725de',
u'purplered': u'#990147',
u'purpley': u'#8756e4',
u'purpleyblue': u'#5f34e7',
u'purpleygrey': u'#947e94',
u'purpleypink': u'#c83cb9',
u'purplish': u'#94568c',
u'purplishblue': u'#601ef9',
u'purplishbrown': u'#6b4247',
u'purplishgrey': u'#7a687f',
u'purplishpink': u'#ce5dae',
u'purplishred': u'#b0054b',
u'purply': u'#983fb2',
u'purplyblue': u'#661aee',
u'purplypink': u'#f075e6',
u'putty': u'#beae8a',
u'racinggreen': u'#014600',
u'radioactivegreen': u'#2cfa1f',
u'raspberry': u'#b00149',
u'rawsienna': u'#9a6200',
u'rawumber': u'#a75e09',
u'reallylightblue': u'#d4ffff',
u'red': u'#ff0000',
u'redbrown': u'#8b2e16',
u'reddish': u'#c44240',
u'reddishbrown': u'#7f2b0a',
u'reddishgrey': u'#997570',
u'reddishorange': u'#f8481c',
u'reddishpink': u'#fe2c54',
u'reddishpurple': u'#910951',
u'reddybrown': u'#6e1005',
u'redorange': u'#fd3c06',
u'redpink': u'#fa2a55',
u'redpurple': u'#820747',
u'redviolet': u'#9e0168',
u'redwine': u'#8c0034',
u'richblue': u'#021bf9',
u'richpurple': u'#720058',
u'robin': u'#6dedfd',
u'robineggblue': u'#8af1fe',
u'rosa': u'#fe86a4',
u'rose': u'#cf6275',
u'rosepink': u'#f7879a',
u'rosered': u'#be013c',
u'rosybrown': u'#bc8f8f',
u'rosypink': u'#f6688e',
u'rouge': u'#ab1239',
u'royal': u'#0c1793',
u'royalblue': u'#4169e1',
u'royalpurple': u'#4b006e',
u'ruby': u'#ca0147',
u'russet': u'#a13905',
u'rust': u'#a83c09',
u'rustbrown': u'#8b3103',
u'rustorange': u'#c45508',
u'rustred': u'#aa2704',
u'rustyorange': u'#cd5909',
u'rustyred': u'#af2f0d',
u'saddlebrown': u'#8b4513',
u'saffron': u'#feb209',
u'sage': u'#87ae73',
u'sagegreen': u'#88b378',
u'salmon': u'#fa8072',
u'salmonpink': u'#fe7b7c',
u'sand': u'#e2ca76',
u'sandbrown': u'#cba560',
u'sandstone': u'#c9ae74',
u'sandy': u'#f1da7a',
u'sandybrown': u'#f4a460',
u'sandyellow': u'#fce166',
u'sandyyellow': u'#fdee73',
u'sapgreen': u'#5c8b15',
u'sapphire': u'#2138ab',
u'scarlet': u'#be0119',
u'sea': u'#3c9992',
u'seablue': u'#047495',
u'seafoam': u'#80f9ad',
u'seafoamblue': u'#78d1b6',
u'seafoamgreen': u'#7af9ab',
u'seagreen': u'#2e8b57',
u'seashell': u'#fff5ee',
u'seaweed': u'#18d17b',
u'seaweedgreen': u'#35ad6b',
u'sepia': u'#985e2b',
u'shamrock': u'#01b44c',
u'shamrockgreen': u'#02c14d',
u'shit': u'#7f5f00',
u'shitbrown': u'#7b5804',
u'shitgreen': u'#758000',
u'shockingpink': u'#fe02a2',
u'sickgreen': u'#9db92c',
u'sicklygreen': u'#94b21c',
u'sicklyyellow': u'#d0e429',
u'sienna': u'#a0522d',
u'silver': u'#c0c0c0',
u'sky': u'#82cafc',
u'skyblue': u'#87ceeb',
u'slate': u'#516572',
u'slateblue': u'#6a5acd',
u'slategray': u'#708090',
u'slategreen': u'#658d6d',
u'slategrey': u'#59656d',
u'slimegreen': u'#99cc04',
u'snot': u'#acbb0d',
u'snotgreen': u'#9dc100',
u'snow': u'#fffafa',
u'softblue': u'#6488ea',
u'softgreen': u'#6fc276',
u'softpink': u'#fdb0c0',
u'softpurple': u'#a66fb5',
u'spearmint': u'#1ef876',
u'springgreen': u'#00ff7f',
u'spruce': u'#0a5f38',
u'squash': u'#f2ab15',
u'steel': u'#738595',
u'steelblue': u'#4682b4',
u'steelgrey': u'#6f828a',
u'stone': u'#ada587',
u'stormyblue': u'#507b9c',
u'straw': u'#fcf679',
u'strawberry': u'#fb2943',
u'strongblue': u'#0c06f7',
u'strongpink': u'#ff0789',
u'sunflower': u'#ffc512',
u'sunfloweryellow': u'#ffda03',
u'sunnyyellow': u'#fff917',
u'sunshineyellow': u'#fffd37',
u'sunyellow': u'#ffdf22',
u'swamp': u'#698339',
u'swampgreen': u'#748500',
u'tan': u'#d2b48c',
u'tanbrown': u'#ab7e4c',
u'tangerine': u'#ff9408',
u'tangreen': u'#a9be70',
u'taupe': u'#b9a281',
u'tea': u'#65ab7c',
u'teagreen': u'#bdf8a3',
u'teal': u'#008080',
u'tealblue': u'#01889f',
u'tealgreen': u'#25a36f',
u'tealish': u'#24bca8',
u'tealishgreen': u'#0cdc73',
u'terracota': u'#cb6843',
u'terracotta': u'#c9643b',
u'thistle': u'#d8bfd8',
u'tiffanyblue': u'#7bf2da',
u'tomato': u'#ff6347',
u'tomatored': u'#ec2d01',
u'topaz': u'#13bbaf',
u'toupe': u'#c7ac7d',
u'toxicgreen': u'#61de2a',
u'treegreen': u'#2a7e19',
u'trueblue': u'#010fcc',
u'truegreen': u'#089404',
u'turquoise': u'#40e0d0',
u'turquoiseblue': u'#06b1c4',
u'turquoisegreen': u'#04f489',
u'turtlegreen': u'#75b84f',
u'twilight': u'#4e518b',
u'twilightblue': u'#0a437a',
u'uglyblue': u'#31668a',
u'uglybrown': u'#7d7103',
u'uglygreen': u'#7a9703',
u'uglypink': u'#cd7584',
u'uglypurple': u'#a442a0',
u'uglyyellow': u'#d0c101',
u'ultramarine': u'#2000b1',
u'ultramarineblue': u'#1805db',
u'umber': u'#b26400',
u'velvet': u'#750851',
u'vermillion': u'#f4320c',
u'verydarkblue': u'#000133',
u'verydarkbrown': u'#1d0200',
u'verydarkgreen': u'#062e03',
u'verydarkpurple': u'#2a0134',
u'verylightblue': u'#d5ffff',
u'verylightbrown': u'#d3b683',
u'verylightgreen': u'#d1ffbd',
u'verylightpink': u'#fff4f2',
u'verylightpurple': u'#f6cefc',
u'verypaleblue': u'#d6fffe',
u'verypalegreen': u'#cffdbc',
u'vibrantblue': u'#0339f8',
u'vibrantgreen': u'#0add08',
u'vibrantpurple': u'#ad03de',
u'violet': u'#ee82ee',
u'violetblue': u'#510ac9',
u'violetpink': u'#fb5ffc',
u'violetred': u'#a50055',
u'viridian': u'#1e9167',
u'vividblue': u'#152eff',
u'vividgreen': u'#2fef10',
u'vividpurple': u'#9900fa',
u'vomit': u'#a2a415',
u'vomitgreen': u'#89a203',
u'vomityellow': u'#c7c10c',
u'warmblue': u'#4b57db',
u'warmbrown': u'#964e02',
u'warmgrey': u'#978a84',
u'warmpink': u'#fb5581',
u'warmpurple': u'#952e8f',
u'washedoutgreen': u'#bcf5a6',
u'waterblue': u'#0e87cc',
u'watermelon': u'#fd4659',
u'weirdgreen': u'#3ae57f',
u'wheat': u'#f5deb3',
u'white': u'#ffffff',
u'whitesmoke': u'#f5f5f5',
u'windowsblue': u'#3778bf',
u'wine': u'#80013f',
u'winered': u'#7b0323',
u'wintergreen': u'#20f986',
u'wisteria': u'#a87dc2',
u'yellow': u'#ffff00',
u'yellowbrown': u'#b79400',
u'yellowgreen': u'#9acd32',
u'yellowish': u'#faee66',
u'yellowishbrown': u'#9b7a01',
u'yellowishgreen': u'#b0dd16',
u'yellowishorange': u'#ffab0f',
u'yellowishtan': u'#fcfc81',
u'yellowochre': u'#cb9d06',
u'yelloworange': u'#fcb001',
u'yellowtan': u'#ffe36e',
u'yellowybrown': u'#ae8b0c',
u'yellowygreen': u'#bff128',
}
XKCD_HEX_TO_NAMES = _reversedict(XKCD_NAMES_TO_HEX)
len(XKCD_HEX_TO_NAMES)
# -
# convert some hex XKCD colors to RGB triples
barney, poop, shit = "#a00498", "#937c00", "#7b5804"
print(hex_to_rgb(barney)) # barney purple
print(hex_to_rgb(poop)) # baby poop
print(hex_to_rgb(shit)) # shit brown
print(hex_to_name(barney))
print(hex_to_name("#aaaaaa")) # None - for no match
# convert XKCD color triples back to names
[
XKCD_HEX_TO_NAMES.get(normalize_hex(barney)),
XKCD_HEX_TO_NAMES.get(normalize_hex(poop)),
XKCD_HEX_TO_NAMES.get(normalize_hex(shit))
]
# The following code snippets are used in `SmugPyter` to map arbitrary RGB color triples
# to the closest named color where "closest" is defined by Euclidean distance. I'm aware
# that this RGB "distance" may not be the best metric. I've been planning on
# testing other color spaces and other metrics to determine if they conform better
# to photographer expections. Maybe one day I will get around to it.
# +
# slight variations on colorkeys.py code
def closest_color(requested_color):
min_colors = {}
for key, name in XKCD_HEX_TO_NAMES.items():
r_c, g_c, b_c = hex_to_rgb(key)
rd = (r_c - requested_color[0]) ** 2
gd = (g_c - requested_color[1]) ** 2
bd = (b_c - requested_color[2]) ** 2
min_colors[(rd + gd + bd)] = name
return (min_colors[min(min_colors.keys())], sqrt(min(min_colors.keys())))
def get_xkcd_color_name(requested_color):
rgb_distance = 0
closest_name = actual_name = rgb_to_name(normalize_integer_triplet(requested_color))
if actual_name == None:
closest_name, rgb_distance = closest_color(requested_color)
return (actual_name, closest_name, rgb_distance)
# exact and nearest color
requested_color = hex_to_rgb(barney)
actual_name, closest_name , rgb_distance = get_xkcd_color_name(requested_color)
print("Actual color name:", actual_name, ", closest color name:", closest_name, " rgb distance:", rgb_distance)
requested_color = (119, 172, 152)
actual_name, closest_name , rgb_distance = get_xkcd_color_name(requested_color)
print("Actual color name:", actual_name, ", closest color name:", closest_name, " rgb distance:", rgb_distance)
# -
# ### Generate Some Random Colors
for a,b in 15 * [(0,255)]:
rgb = (random.randint(a,b), random.randint(a,b), random.randint(a,b))
_, xkcd_name, rgb_distance = get_xkcd_color_name(rgb)
print((xkcd_name, rgb_distance))
print(hex_to_rgb(XKCD_NAMES_TO_HEX.get(xkcd_name)))
# ### Fun with XKCD Colors
#
# Now that we can compute the nearest XKCD color given an arbitrary RGB triplet let's explore the *deep issues*
# like what exactly does "diarreha" and "baby shit green" look like.
diarrhea = u'#9f8303'
r, g, b = hex_to_rgb(diarrhea)
patch = Image.new("RGB", (100, 100), (int(r),int(g),int(b)))
display(patch)
babyshitgreen = u'#889717'
r, g, b = hex_to_rgb(babyshitgreen)
patch = Image.new("RGB", (100, 100), (int(r),int(g),int(b)))
display(patch)
# ### Load Test Images
#
# To compare XKCD colors to `webcolors` let's load the same test images used by the notebook
# [Finding Dominant Color Names from Color Space Coordinates and Images](https://github.com/bakerjd99/smugpyter/blob/master/notebooks/Finding%20Dominant%20Color%20Names%20from%20Color%20Space%20Coordinates%20and%20Images.ipynb).
#
image1 = Image.open('C:/SmugMirror/Mirror/Themes/Manipulations/ImageHacking/5NB7dXP-1f-green-gray-dragon-eggs.jpg')
image2 = Image.open('C:/SmugMirror/Mirror/Themes/Manipulations/ImageHacking/hjbftwN-1-your-grainy-hell-awaits-[409595101].jpg')
image3 = Image.open('C:/SmugMirror/Mirror/People/GreatandGreaterForebearers/LX8HmDV-2z-william-evert-baker-portrait-1950.jpg')
image4 = Image.open('C:/SmugMirror/Mirror/Themes/Manipulations/Panoramas/b36sc3H-1-norris-porcelain-basin-yellowstone-panorma.jpg')
# +
def resize_image(image, factor):
"""Resize PIL image maintaining aspect ratio."""
imcopy = image.copy()
width, height = imcopy.size
width = int(factor * width)
height = int(factor * height)
return imcopy.resize((width, height))
def show_xkcd_image_colors(image, colors, *, factor=0.4, patch_size=(50,50), spacing=10):
"""
Display resized image with adjacent color patches.
The top patch row shows raw colors and the second
shows nearest named XKCD colors.
"""
imnew = resize_image(image, factor)
# NIMP check that image is 2 * larger than patch size.
height = imnew.size[1]
width = imnew.size[0] + (spacing + patch_size[0]) * len(colors)
background = Image.new('RGBA', (width,height), (255, 255, 255, 255))
background.paste(imnew, (0,0))
rgb_distance = []
xkcd_color = []
for i, color in enumerate(colors):
r, g, b = color
patch1 = Image.new("RGB", patch_size, (int(r),int(g),int(b)))
#patch1 = Image.new("RGB", patch_size, color)
background.paste(patch1, (imnew.size[0] + spacing + i * (spacing + patch_size[0]), 0))
_, xkcd_name, rgb_metric = get_xkcd_color_name((int(r),int(g),int(b)))
xkcd_color.append(xkcd_name)
rgb_distance.append(rgb_metric)
# need pixel values for XKCD named colors as PIL images do not support XKCD names
r, g, b = hex_to_rgb(XKCD_NAMES_TO_HEX.get(xkcd_name))
patch2 = Image.new("RGB", patch_size, (r,g,b))
background.paste(patch2, (imnew.size[0] + spacing + i * (spacing + patch_size[0]), spacing + patch_size[1]))
display(background)
print([i for i in zip(xkcd_color, rgb_distance)])
print("mean: %s" % np.mean(rgb_distance))
print("variance: %s" % np.var(rgb_distance))
print("standard dev: %s" % np.std(rgb_distance))
# random color patches
rcolors0 = []
for a,b in 8 * [(0,255)]:
rgb0 = (random.randint(a,b), random.randint(a,b), random.randint(a,b))
print(get_xkcd_color_name(rgb0))
patch = Image.new("RGB", (75, 75), rgb0)
display(patch)
rcolors0.append(rgb0)
# patches beside main image
show_xkcd_image_colors(image2, rcolors0)
# -
# ### XKCD Dominant Color Calculations
#
# My primary reason for hacking around with XKCD colors is to find a better
# list of colors for assigning dominant image color keys. I use [`SmugPyter`](https://github.com/bakerjd99/smugpyter)
# to compute dominant image color keys for my [SmugMug pictures](https://conceptcontrol.smugmug.com/). If you
# browse my pictures you will see many keywords like: `0_burlywood`, `0_rosybrown`, `0_gainsboro` and `0_paleturquoise`.
# These color keywords are computed by `SmugPyter`.
#
# `SmugPyter` uses `webcolors` and `webcolors` contains 147 colors. Standard RGB supports sixteen million colors.
# Obviously, when you reduce sixteen million choices to 147 there are going to be gaps. The gaps show as
# questionable color shifts from raw RGB to named colors and in the clustering of named colors.
#
# When I computed dominant colors for all my SmugMug images I found:
#
# 1. Only 99 of 147 possible colors appear: about 67%.
#
# 2. The first six colors are assigned to over 50% of all images.
#
# 3. The first color is assigned more often than the last 79 colors.
#
# 4. *kmeans* dominant colors are not particularily meaningful for photographs.
#
# Expanding the list of named colors should result in more colors being used.
#
# The following code cell uses this [algorithm from Stack Overflow](https://stackoverflow.com/questions/3241929/python-find-dominant-most-common-color-in-an-image) to compute dominant colors.
# +
def array_from_image(image, *, factor=0.4):
image = resize_image(image, factor)
ar = np.asarray(image)
shape = ar.shape
ar = ar.reshape(scipy.product(shape[:2]), shape[2]).astype(float)
return ar
def calc1_colors(ar, *, num_clusters=8, factor=0.4):
colors, dist = scipy.cluster.vq.kmeans(ar, num_clusters)
return colors
def calc1_peak(ar, colors):
vecs, dist = scipy.cluster.vq.vq(ar, colors) # assign codes
counts, bins = scipy.histogram(vecs, len(colors)) # count occurrences
index_max = scipy.argmax(counts) # find most frequent
peak = colors[index_max]
return peak
def calc1_colors_peak(image, *, num_clusters=8, factor=0.4):
ar = array_from_image(image, factor=factor)
colors = calc1_colors(ar, num_clusters=num_clusters)
peak = calc1_peak(ar, colors)
return (colors, peak)
NUM_CLUSTERS = 8
calc1_colors1, calc1_peak1 = calc1_colors_peak(image1, num_clusters=NUM_CLUSTERS)
calc1_colors2, calc1_peak2 = calc1_colors_peak(image2, num_clusters=NUM_CLUSTERS)
calc1_colors3, calc1_peak3 = calc1_colors_peak(image3, num_clusters=NUM_CLUSTERS)
calc1_colors4, calc1_peak4 = calc1_colors_peak(image4, num_clusters=NUM_CLUSTERS)
# -
# Applying the dominant color calculation to the test images produces.
show_xkcd_image_colors(image1, calc1_colors1)
show_xkcd_image_colors(image2, calc1_colors2)
show_xkcd_image_colors(image3, calc1_colors3)
show_xkcd_image_colors(image4, calc1_colors4)
# ### More colors reduces mean RGB distance
#
# The mean distance between raw RGB colors and named XKCD colors is lower than the corresponding
# mean distance between raw RGB colors and `webcolors`. This is expected. The XKCD colors cover
# the RGB space better and you don't need to shift the raw RGB values as much. The longer list
# doesn't prevent all bad shifts but it reduces their frequency.
#
# The following J calculations show the difference in RGB space coverage between `webcolors`
# and unified XKCD colors.
# start J assumes jbase.py on Python's search path
j.init()
# load JOD addon and get smugpyter hacks
j.dor("load 'general/jod'")
j.dor("od ;:'smugpyter utils' [ 3 od '' ")
# generate and load the XKCDcolors group
j.dor("lg 'XKCDcolors'")
# display short help for group words
j.dor("hg 'XKCDcolors'")
# display web colors coverage verb
# a similar verb computes coverage for XKCD colors
j.dor("disp 'webdist'")
# The next calculation inserts the color sets into a `(256 256 256)` RGB color array.
# Each array coordinate represents one of the `256^3` possible eight bit RGB colors. The result
# is a `(4 4 4)` integer array that counts the number of colors that fall in each `(64 64 64)`
# subarray. The subarrays partition the larger array into sixty-four nonoverlapping bins.
#
# If the named colors are evenly distributed over the RGB cube you would expect the
# `138` unique webcolors to result in about `2.15` colors per cell and the `970` XKCD colors
# should be around `15.15.`
print(138 / 64)
print(970 / 64)
# run webdist , xkcdist and show counts
j.dor("'webcnt wednames'=.webdist 0")
j.dor("'xkcdcnt xkcdnames'=. xkcddist 0")
j.dor("webcnt;xkcdcnt")
j.dor("load 'bstats'")
j.dor("dstat&.> ,&.> webcnt;xkcdcnt")
# `webcolors` is not as smoothly distributed as you would like. Many cells have no colors
# and one has 30. There are a lot of `webcolor` names for bright colors. XKCD colors cover
# all the bins but the count variance is high.
# ### Basic `SmugPyter` Dominant Color Calculation with XKCD Colors
# +
def dualsort(a, b, *, reverse=False):
"""
Sort lists (a) and (b) using (a) to grade (b).
"""
temp = sorted(zip(a, b), key=lambda x: x[0], reverse=reverse)
return list(map(list, zip(*temp)))
def most_common(lst):
""" Pick most common item in a list - ok for small lists."""
return max(set(lst), key=lst.count)
def cluster_name_freq_dist(image, *, num_clusters=8, factor=0.4):
"""
Returns a tuple of sorted nearest named colors, cluster frequencies, and
distances from cluster raw colors. Items are sorted by decreasing
cluster frequency.
"""
km = KMeans(n_clusters=num_clusters)
imcopy = resize_image(image, factor=factor)
ar = np.asarray(imcopy)
pixels = ar.reshape((-1, 3))
km.fit(pixels)
colors = np.asarray(km.cluster_centers_, dtype=np.uint8)
frequencies = np.asarray(np.unique(km.labels_, return_counts = True)[1], dtype=np.int32)
names = []
distances = []
for color in colors:
_ , name , rgb_distance = get_xkcd_color_name(color)
names.append(name)
distances.append(rgb_distance)
# order by decreasing frequency
_ , names = dualsort(frequencies, names, reverse=True)
frequencies , distances = dualsort(frequencies, distances, reverse=True)
return (names, frequencies, distances)
def dominant_color_key(names_freqs_dists):
"""
Return a single dominant color key.
"""
names , frequencies , distances = names_freqs_dists
if len(names) > len(list(set(names))):
# most frequent repeated named color
key = most_common(names)
else:
# distances less greatest outlier
dist_sample = sorted(distances, reverse=True)[1:]
threshold = np.mean(dist_sample) + np.std(dist_sample)
# default color choice
key = names[0]
# return first color from sorted names that is no more
# than one standard deviation from the sample mean. If
# no such choice is made retain the default selection.
for name, distance in zip(names, distances):
if distance <= threshold:
key = name
break
return key.lower().strip()
names_freq_dist = cluster_name_freq_dist(image1)
print(dominant_color_key(names_freq_dist))
# -
# gather unique list of sample image files
# the same image may occur in many gallery directories
unique_files = set()
unique_full_paths = []
for i, full_path in enumerate(glob.iglob('c:\SmugMirror\Mirror\**\*.jpg', recursive=True)):
file_name = os.path.basename(full_path)
if file_name in unique_files:
#print("duplicate -> %s, %s" % (i,file_name))
continue
unique_files.add(file_name)
unique_full_paths.append(full_path)
print(len(unique_files))
print(len(unique_full_paths))
# ### Compute XKCD color for all SmugMug sample images
#
# The next cell computes the dominant color for all my SmugMug sample images. There are
# are about 4000 images. This calculation takes over an hour on my machine.
# +
# collect sample of XKCD dominant colors
rows = []
file_count = len(unique_full_paths)
for i, file_name in enumerate(unique_full_paths):
image = Image.open(file_name)
names_freq_dist = cluster_name_freq_dist(image)
xkcd_color = dominant_color_key(names_freq_dist)
color_info = {'XKCDColor': xkcd_color, 'SampleFile': os.path.basename(file_name)}
rows.append(color_info)
# every n rows print message
if 0 == (i % 20):
print("%s/%s %s - %s" % (i + 1, file_count, xkcd_color, os.path.basename(file_name)))
# write TAB delimited file
sample_file = "c:/temp/xkcd_dominant.txt"
keys = rows[0].keys()
with open(sample_file, 'w', newline='') as output_file:
dict_writer = csv.DictWriter(output_file, keys, dialect='excel-tab')
dict_writer.writeheader()
dict_writer.writerows(rows)
# -
# ### Frequencies of computed colors
#
# Applying the following J verb to the file computed in the previous cell.
j.dor('xkcdcolorfreq')
j.dor("cf=. xkcdcolorfreq 'c:/temp/xkcd_dominant.txt'")
j.dor("(0{cf) ,: 30 {.&.> 1{cf")
# ### Final Remarks
#
# Increasing the number of named colors does not significantly improve
# the quality of dominant color selection. The main effect is to
# increase the number of grays and near grays. This is not unexpected. There
# is a reason that photographs have been calibrated against [gray cards](https://en.wikipedia.org/wiki/Gray_card) for
# almost a century. If you average out all the colors in typical scenes
# in well exposed pictures you usually end up with a neutral tone or gray.
# Here the top ten dominant colors are mostly gray.
# +
topten = ['almostblack','silver','black','gainsboro','lightgray','lavender',
'darkgray','charcoalgrey','darkgrey','charcoal']
for color in topten:
r, g, b = hex_to_rgb(XKCD_NAMES_TO_HEX.get(color))
print(color)
patch = Image.new("RGB", (75, 75), (int(r),int(g),int(b)))
display(patch)
| notebooks/Unified XKCD Colors.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
from decision_tree import *
from data import my_data
tree = build_tree(my_data)
# +
def print_tree(tree, indent=''):
if tree.results != None:
print str(tree.results)
else:
print str(tree.col) + ':' + str(tree.value) +'? '
print indent + 'T->',
print_tree(tree.tb, indent + " ")
print indent + 'F->',
print_tree(tree.fb, indent + " ")
print_tree(tree)
# -
# 处理数值型的结果
def variance(rows):
if len(rows) == 0:
return 0
data = [float(row[-1]) for row in rows]
mean = sum(data) / len(data)
variance = sum([(d - mean) ** 2 for d in data]) / len(data)
return variance
| decide-tree/decision_tree_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
rfile = open("testlog.csv", "r")
lines = rfile.readlines()
len(lines)
lines[0]
np.arange(len(lines))
nparr = np.empty(len(lines) - 1)
from collections import Counter
for i in range(1, len(lines)):
l = lines[i]
l = l.strip("\n")
intV = int(l)
nparr[i-1] = intV
counter=Counter(nparr)
top = 100
print(np.arange(top))
print(counter.most_common(top))
plt.scatter(counter.most_common(top).keys, counter.most_common(top).values)
for i in range(1, 10):
print(i)
| mofanpython/analyze_log.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# Copyright (c) Microsoft Corporation. All rights reserved.
#
# Licensed under the MIT License.
# This notebook demonstrates how to run batch scoring job. __[Inception-V3 model](https://arxiv.org/abs/1512.00567)__ and unlabeled images from __[ImageNet](http://image-net.org/)__ dataset will be used. It registers a pretrained inception model in model registry then uses the model to do batch scoring on images in a blob container.
# +
import os
from azureml.core import Workspace, Run, Experiment
ws = Workspace.from_config()
print('Workspace name: ' + ws.name,
'Azure region: ' + ws.location,
'Subscription id: ' + ws.subscription_id,
'Resource group: ' + ws.resource_group, sep = '\n')
# Also create a Project and attach to Workspace
project_folder = "sample_projects"
run_history_name = project_folder
if not os.path.isdir(project_folder):
os.mkdir(project_folder)
# -
from azureml.core.compute import BatchAiCompute, ComputeTarget
from azureml.core.datastore import Datastore
from azureml.data.data_reference import DataReference
from azureml.pipeline.core import Pipeline, PipelineData
from azureml.pipeline.steps import PythonScriptStep
from azureml.core.runconfig import CondaDependencies, RunConfiguration
# ## Create and attach Compute targets
# Use the below code to create and attach Compute targets.
# Batch AI compute
cluster_name = "gpu_cluster"
try:
cluster = BatchAiCompute(ws, cluster_name)
print("found existing cluster.")
except:
print("creating new cluster")
provisioning_config = BatchAiCompute.provisioning_configuration(vm_size = "STANDARD_NC6",
autoscale_enabled = True,
cluster_min_nodes = 0,
cluster_max_nodes = 1)
# create the cluster
cluster = ComputeTarget.create(ws, cluster_name, provisioning_config)
cluster.wait_for_completion(show_output=True)
# # Python scripts to run
# Python scripts that run the batch scoring. `batchai_score.py` takes input images in `dataset_path`, pretrained models in `model_dir` and outputs a `results-label.txt` to `output_dir`.
# +
# %%writefile $project_folder/batchai_score.py
import os
import argparse
import datetime,time
import tensorflow as tf
from math import ceil
import numpy as np
import shutil
from tensorflow.contrib.slim.python.slim.nets import inception_v3
from azureml.core.model import Model
slim = tf.contrib.slim
parser = argparse.ArgumentParser(description="Start a tensorflow model serving")
parser.add_argument('--model_name', dest="model_name", required=True)
parser.add_argument('--label_dir', dest="label_dir", required=True)
parser.add_argument('--dataset_path', dest="dataset_path", required=True)
parser.add_argument('--output_dir', dest="output_dir", required=True)
parser.add_argument('--batch_size', dest="batch_size", type=int, required=True)
args = parser.parse_args()
image_size = 299
num_channel = 3
# create output directory if it does not exist
os.makedirs(args.output_dir, exist_ok=True)
def get_class_label_dict(label_file):
label = []
proto_as_ascii_lines = tf.gfile.GFile(label_file).readlines()
for l in proto_as_ascii_lines:
label.append(l.rstrip())
return label
class DataIterator:
def __init__(self, data_dir):
self.file_paths = []
image_list = os.listdir(data_dir)
total_size = len(image_list)
self.file_paths = [data_dir + '/' + file_name.rstrip() for file_name in image_list ]
self.labels = [1 for file_name in self.file_paths]
@property
def size(self):
return len(self.labels)
def input_pipeline(self, batch_size):
images_tensor = tf.convert_to_tensor(self.file_paths, dtype=tf.string)
labels_tensor = tf.convert_to_tensor(self.labels, dtype=tf.int64)
input_queue = tf.train.slice_input_producer([images_tensor, labels_tensor], shuffle=False)
labels = input_queue[1]
images_content = tf.read_file(input_queue[0])
image_reader = tf.image.decode_jpeg(images_content, channels=num_channel, name="jpeg_reader")
float_caster = tf.cast(image_reader, tf.float32)
new_size = tf.constant([image_size, image_size], dtype=tf.int32)
images = tf.image.resize_images(float_caster, new_size)
images = tf.divide(tf.subtract(images, [0]), [255])
image_batch, label_batch = tf.train.batch([images, labels], batch_size=batch_size, capacity=5 * batch_size)
return image_batch
def main(_):
start_time = datetime.datetime.now()
label_file_name = os.path.join(args.label_dir, "labels.txt")
label_dict = get_class_label_dict(label_file_name)
classes_num = len(label_dict)
test_feeder = DataIterator(data_dir=args.dataset_path)
total_size = len(test_feeder.labels)
count = 0
# get model from model registry
model_path = Model.get_model_path(args.model_name)
with tf.Session() as sess:
test_images = test_feeder.input_pipeline(batch_size=args.batch_size)
with slim.arg_scope(inception_v3.inception_v3_arg_scope()):
input_images = tf.placeholder(tf.float32, [args.batch_size, image_size, image_size, num_channel])
logits, _ = inception_v3.inception_v3(input_images,
num_classes=classes_num,
is_training=False)
probabilities = tf.argmax(logits, 1)
sess.run(tf.global_variables_initializer())
sess.run(tf.local_variables_initializer())
coord = tf.train.Coordinator()
threads = tf.train.start_queue_runners(sess=sess, coord=coord)
saver = tf.train.Saver()
saver.restore(sess, model_path)
out_filename = os.path.join(args.output_dir, "result-labels.txt")
with open(out_filename, "w") as result_file:
i = 0
while count < total_size and not coord.should_stop():
test_images_batch = sess.run(test_images)
file_names_batch = test_feeder.file_paths[i*args.batch_size: min(test_feeder.size, (i+1)*args.batch_size)]
results = sess.run(probabilities, feed_dict={input_images: test_images_batch})
new_add = min(args.batch_size, total_size-count)
count += new_add
i += 1
for j in range(new_add):
result_file.write(os.path.basename(file_names_batch[j]) + ": " + label_dict[results[j]] + "\n")
result_file.flush()
coord.request_stop()
coord.join(threads)
# copy the file to artifacts
shutil.copy(out_filename, "./outputs/")
# Move the processed data out of the blob so that the next run can process the data.
if __name__ == "__main__":
tf.app.run()
# -
# ## Prepare Model and Input data
# create directory for model
model_dir = 'models'
if not os.path.isdir(model_dir):
os.mkdir(model_dir)
# ### Download Model
# <font color=red>This manual step is required to register the model to the workspace</font>
#
# Download and extract model from http://download.tensorflow.org/models/inception_v3_2016_08_28.tar.gz to model_dir
# ### Get samples images and upload to Datastore
# <font color=red>This manual step is required to run batchai_score.py</font>
#
# Download and extract sample images from ImageNet evaluation set and **upload** to a blob that will be registered as a Datastore in the next step
#
# A copy of sample images from ImageNet evaluation set can be found at __[BatchAI Samples Blob](https://batchaisamples.blob.core.windows.net/samples/imagenet_samples.zip?st=2017-09-29T18%3A29%3A00Z&se=2099-12-31T08%3A00%3A00Z&sp=rl&sv=2016-05-31&sr=c&sig=PmhL%2BYnYAyNTZr1DM2JySvrI12e%2F4wZNIwCtf7TRI%2BM%3D)__
#
# There are multiple ways to create folders and upload files into Azure Blob Container - you can use __[Azure Portal](https://ms.portal.azure.com/)__, __[Storage Explorer](http://storageexplorer.com/)__, __[Azure CLI2](https://render.githubusercontent.com/azure-cli-extension)__ or Azure SDK for your preferable programming language.
account_name = "batchscoringdata"
sample_data = Datastore.register_azure_blob_container(ws, "sampledata", "sampledata",
account_name=account_name,
overwrite=True)
# # Output datastore
# We write the outputs to the default datastore
default_ds = "workspaceblobstore"
# # Specify where the data is stored or will be written to
from azureml.core.conda_dependencies import CondaDependencies
from azureml.data.data_reference import DataReference
from azureml.pipeline.core import Pipeline, PipelineData
from azureml.core import Datastore
from azureml.core import Experiment
input_images = DataReference(datastore=sample_data,
data_reference_name="input_images",
path_on_datastore="batchscoring/images",
mode="download"
)
model_dir = DataReference(datastore=sample_data,
data_reference_name="input_model",
path_on_datastore="batchscoring/models",
mode="download"
)
label_dir = DataReference(datastore=sample_data,
data_reference_name="input_labels",
path_on_datastore="batchscoring/labels",
mode="download"
)
output_dir = PipelineData(name="scores",
datastore_name=default_ds,
output_path_on_compute="batchscoring/results")
# ## Register the model with Workspace
# +
import shutil
from azureml.core.model import Model
# register downloaded model
model = Model.register(model_path = "models/inception_v3.ckpt",
model_name = "inception", # this is the name the model is registered as
tags = {'pretrained': "inception"},
description = "Imagenet trained tensorflow inception",
workspace = ws)
# remove the downloaded dir after registration if you wish
shutil.rmtree("models")
# -
# # Specify environment to run the script
# +
cd = CondaDependencies.create(pip_packages=["tensorflow-gpu==1.4.0", "azureml-defaults"])
# Runconfig
batchai_run_config = RunConfiguration(conda_dependencies=cd)
batchai_run_config.environment.docker.enabled = True
batchai_run_config.environment.docker.gpu_support = True
batchai_run_config.environment.docker.base_image = "microsoft/mmlspark:gpu-0.12"
batchai_run_config.environment.spark.precache_packages = False
# -
# # Steps to run
# A subset of the parameters to the python script can be given as input when we re-run a `PublishedPipeline`. In the current example, we define `batch_size` taken by the script as such parameter.
from azureml.pipeline.core.graph import PipelineParameter
batch_size_param = PipelineParameter(name="param_batch_size", default_value=20)
# +
inception_model_name = "inception_v3.ckpt"
batch_score_step = PythonScriptStep(
name="batch ai scoring",
script_name="batchai_score.py",
arguments=["--dataset_path", input_images,
"--model_name", "inception",
"--label_dir", label_dir,
"--output_dir", output_dir,
"--batch_size", batch_size_param],
target=cluster,
inputs=[input_images, label_dir],
outputs=[output_dir],
runconfig=batchai_run_config,
source_directory=project_folder
)
# -
pipeline = Pipeline(workspace=ws, steps=[batch_score_step])
pipeline_run = Experiment(ws, 'batch_scoring').submit(pipeline, pipeline_params={"param_batch_size": 20})
# # Monitor run
from azureml.train.widgets import RunDetails
RunDetails(pipeline_run).show()
pipeline_run.wait_for_completion(show_output=True)
# # Download and review output
step_run = list(pipeline_run.get_children())[0]
step_run.download_file("./outputs/result-labels.txt")
import pandas as pd
df = pd.read_csv("result-labels.txt", delimiter=":", header=None)
df.columns = ["Filename", "Prediction"]
df.head()
# # Publish a pipeline and rerun using a REST call
# ## Create a published pipeline
# +
published_pipeline = pipeline_run.publish_pipeline(
name="Inception v3 scoring", description="Batch scoring using Inception v3 model", version="1.0")
published_id = published_pipeline.id
# -
# ## Rerun using REST call
# ## Get AAD token
# +
from azureml.core.authentication import AzureCliAuthentication
import requests
cli_auth = AzureCliAuthentication()
aad_token = cli_auth.get_authentication_header()
# -
# ## Run published pipeline using its REST endpoint
# +
from azureml.pipeline.core import PublishedPipeline
rest_endpoint = PublishedPipeline.get_endpoint(published_id, ws)
# specify batch size when running the pipeline
response = requests.post(rest_endpoint, headers=aad_token, json={"param_batch_size": 50})
run_id = response.json()["Id"]
# -
# ## Monitor the new run
# +
from azureml.pipeline.core.run import PipelineRun
published_pipeline_run = PipelineRun(ws.experiments()["batch_scoring"], run_id)
RunDetails(published_pipeline_run).show()
| pipeline/pipeline-batch-scoring.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from scipy import stats
import pandas as pd
import seaborn as sns
import statsmodels.api as sm
# +
def rename_cols_and_save(xls_name):
df = pd.read_excel("../../data/{0}.xls".format(xls_name), index_col=None, header=None)
if xls_name == 'hprice1':
names_dict = {0:'price',
1:'assess',
2:'bdrms',
3:'lotsize',
4:'sqrft',
5:'colonial',
6:'lprice',
7:'lassess',
8:'llotsize',
9:'lsqrft',
}
df.rename(columns = names_dict, inplace = True)
df.to_csv("../../data/{0}.csv".format(xls_name), index=False)
return df
df = rename_cols_and_save(xls_name='hprice1')
# -
# ## Estimating a Simple Regression Model
sns_plot = sns.lmplot(x="sqrft", y="price", data=df)
sns_plot.savefig("images/correlation_price_sqrft.png")
# +
X = df["sqrft"]
X = sm.add_constant(X)
y = df["price"]
# Note the difference in argument order
model = sm.OLS(y, X).fit()
df['predictions_linear'] = predictions = model.predict(X) # make the predictions by the model
# Print out the statistics
model.summary()
# -
# ### Plotting Predicted vs. Actual
sns_plot = sns.lmplot(x="predictions_linear", y="price", data=df)
sns_plot.savefig("images/correlation_predictions_vs_actual.png")
# ### Using Diagnostic Residual Plots
# #### Examining Linear Relationship
df['residual'] = df["price"] - df ["predictions_linear"]
def dist_plot(df, var, color):
sns_plot = sns.distplot(df[var], color=color).get_figure()
sns_plot.savefig("images/dist_plot_{0}.png".format(var))
return None
dist_plot(df=df, var='residual', color='b')
# +
def normality_test(df, var):
k2, p = stats.normaltest(df[var])
alpha = 1e-3
print("p = {:g}".format(p))
print("null hypothesis: x comes from a normal distribution")
if p < alpha: # null hypothesis: x comes from a normal distribution
print("The null hypothesis can be rejected")
else:
print("The null hypothesis cannot be rejected")
return None
# -
normality_test(df=df, var='residual')
sns_plot = sns.lmplot(x="sqrft", y="residual", data=df)
sns_plot.savefig("images/correlation_sqrft_vs_residual.png")
# #### Examining Quadratic Relationship
sns_plot = sns.lmplot(x="sqrft", y="price", data=df, order=2)
sns_plot.savefig("images/correlation_price_sqrft_squared.png")
# +
df["squared_sqrft"] = df["sqrft"]**2
X = df["squared_sqrft"]
X = sm.add_constant(X)
y = df["price"]
# Note the difference in argument order
model = sm.OLS(y, X).fit()
df['predictions_nonlinear_sqr'] = predictions = model.predict(X) # make the predictions by the model
# Print out the statistics
model.summary()
# -
df['residual_nonlinear_sqr'] = df["price"] - df ["predictions_nonlinear_sqr"]
dist_plot(df=df, var='predictions_nonlinear_sqr', color='b')
normality_test(df=df, var='predictions_nonlinear_sqr')
sns_plot = sns.lmplot(x="sqrft", y="residual_nonlinear_sqr", data=df)
sns_plot.savefig("images/correlation_sqrft_vs_residual_nonlinear_sqr.png")
| notebooks/1b_simple_linear_regression_goodness_of_fit/1b_simple_linear_regression_goodness_of_fit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.2 64-bit (''.env'': venv)'
# language: python
# name: python37
# ---
# # Benchmarking Thinc layers with a custom `benchmark` layer
#
# This notebook shows how to write a `benchmark` layer that can wrap any layer(s) in your network and that **logs the execution times** of the initialization, forward pass and backward pass. The benchmark layer can also be mapped to an operator like `@` to make it easy to add debugging to your network.
# !pip install "thinc>=8.0.0a0" ml_datasets
# To log the results, we first set up a custom logger using Python's `logging` module. You could also just print the stats instead, but using `logging` is cleaner, since it lets other users modify the logger's behavior more easily, and separates the logs from other output and write it to a file (e.g. if you're benchmarking several layers during training). The following logging config will output the date and time, the name of the logger and the logged results.
# +
import logging
logger = logging.getLogger("thinc:benchmark")
if not logger.hasHandlers(): # prevent Jupyter from adding multiple loggers
formatter = logging.Formatter('%(asctime)s %(name)s %(message)s', datefmt="%Y-%m-%d %H:%M:%S")
handler = logging.StreamHandler()
handler.setFormatter(formatter)
logger.addHandler(handler)
logger.setLevel(logging.DEBUG)
# -
# Here's a minimalistic time logger that can be initialized with the name of a given layer, and can track several events (e.g. `"forward"` and `"backward"`). When the `TimeLogger.end` method is called, the output is formatted nicely and the elapsed time is logged with the logger name and colored label.
# +
from timeit import default_timer
from wasabi import color
class TimeLogger:
def __init__(self, name):
self.colors = {"forward": "green", "backward": "blue"}
self.name = name
self.timers = {}
def start(self, name):
self.timers[name] = default_timer()
def end(self, name):
result = default_timer() - self.timers[name]
label = f"{name.upper():<8}"
label = color(label, self.colors.get(name), bold=True)
logger.debug(f"{self.name:<12} | {label} | {result:.6f}")
# -
# The `benchmark` layer now has to wrap the forward pass, backward pass and initialization of the layer it wraps and log the execution times. It then returns a Thinc model instance with the custom `forward` function and a custom `init` function. We'll also allow setting a custom `name` to make it easier to tell multiple wrapped benchmark layers apart.
# +
from thinc.api import Model
def benchmark(layer, name=None):
name = name if name is not None else layer.name
t = TimeLogger(name)
def init(model, X, Y):
t.start("init")
result = layer.initialize(X, Y)
t.end("init")
return result
def forward(model, X, is_train):
t.start("forward")
layer_Y, layer_callback = layer(X, is_train=is_train)
t.end("forward")
def backprop(dY):
t.start("backward")
result = layer_callback(dY)
t.end("backward")
return result
return layer_Y, backprop
return Model(f"benchmark:{layer.name}", forward, init=init)
# -
# ---
#
# ## Usage examples
#
# ### Using the `benchmark` layer as a function
#
# We can now wrap one or more layers (including nested layers) with the `benchmark` function. This is the original model:
#
# ```python
# model = chain(Linear(1), Linear(1))
# ```
# +
import numpy
from thinc.api import chain, Linear
X = numpy.zeros((1, 2), dtype="f")
model = benchmark(chain(benchmark(Linear(1)), Linear(1)), name="outer")
model.initialize(X=X)
Y, backprop = model(X, is_train=False)
dX = backprop(Y)
# -
# ### Using the `benchmark` layer as an operator
#
# Alternatively, we can also use `Model.define_operators` to map `benchmark` to an operator like `@`. The left argument of the operator is the first argument passed into the function (the layer) and the right argument is the second argument (the name). The following example wraps the whole network (two chained `Linear` layers) in a benchmark layer named `"outer"`, and the first `Linear` layer in a benchmark layer named `"first"`.
# +
from thinc.api import Model
with Model.define_operators({">>": chain, "@": benchmark}):
model = (Linear(1) @ "first" >> Linear(1)) @ "outer"
model.initialize(X=X)
Y, backprop = model(X, is_train=True)
dX = backprop(Y)
# -
# ### Using the `benchmark` layer during training
# +
from thinc.api import Model, chain, Relu, Softmax, Adam
import ml_datasets
n_hidden = 32
dropout = 0.2
with Model.define_operators({">>": chain, "@": benchmark}):
model = (
Relu(nO=n_hidden, dropout=dropout) @ "relu1"
>> Relu(nO=n_hidden, dropout=dropout) @ "relu2"
>> Softmax()
)
train_X = numpy.zeros((5, 784), dtype="f")
train_Y = numpy.zeros((540, 10), dtype="f")
model.initialize(X=train_X[:5], Y=train_Y[:5])
optimizer = Adam(0.001)
for i in range(10):
for X, Y in model.ops.multibatch(8, train_X, train_Y, shuffle=True):
Yh, backprop = model.begin_update(X)
backprop(Yh - Y)
model.finish_update(optimizer)
| examples/05_benchmarking_layers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Week 2
# ## Introduction to Solid State
# +
import numpy as np
import matplotlib.pyplot as plt
import os
import subprocess
from polypy.read import History
from polypy.msd import MSD
from polypy import plotting
def get_diffusion(file, atom):
with open(file) as f:
y = False
for line in f:
if str("atom D ") in line:
y = True
if y == True and str(atom) in line:
d = line.split()
break
return d
# -
# Now that you are familiar with molecular dynamics, you are now going to use it to tackle some real world problems. In the next three weeks you will investigate the transport properties of a simple fluorite material - Ca$F_2$. The transport properties of a material determine many properties that are utilised for modern technological applications. For example, solid oxide fuel cell (SOFCs - Alternative to batteries) materials are dependent on the movement of charge carriers through the solid electrolyte and nuclear fuel materials oxidise and fall apart and this corrosive behaviour is dependent on the diffusion of oxygen into the lattice. Due to the importance of the transport properties of these materials, scientists and engineers spend large amounts of their time tring to optomise these properties using different stoichiometries, introducing defects and by using different syntheisis techniques. Over the next three weeks you will investigate how the transport properties of Ca$F_2$ are affected by temperature, structural defects (Schottky and Frenkel) and by chemcial dopants (e.g. different cations). A rough breakdown looks as follows
# - Week 2
# - Introduction to DL_POLY
# - Tutorial on the calculation of diffusion coefficients
# - Tutorial on the Arhennius equation
# - Molecular dynamics simulations of stoichiomteric Ca$F_2$
# - Week 3
# - Frenkel and Schottky defects
# - Week 4
# - Dopants
#
#
# ## Introduction to DL_POLY
#
# DL_POLY is a molecular dynamics program maintained by Daresbury laboratories. In contrast to pylj, DL_POLY is a three dimensional molecular dynamics code that is used worldwide by computational scientists for molecular simulation, but it should be noted that the theory is exactly the same and any understanding gained from pylj is completely applicable to DL_POLY. For the next three weeks you will use DL_POLY to run short molecular dynamics simulations on Ca$F_2$. You first need to understand the input files required for DL_POLY.
# - CONTROL - This is the file that contains all of the simulation parameters, e.g. simulation temperature, pressure, number of steps e.t.c
# - CONFIG - This is the file that contains the structure - i.e. the atomic coordinates of each atom.
# - FIELD - This is the file that contains the force field or potential model e.g. Lennard Jones.
#
# Contained within the folder "Input" you will find a file called input.txt. This is the main file that you will interact with over the next three weeks and is used to generate the FIELD, CONTROL and CONFIG. Essentially it is easier to meddle with input.txt than it is to meddle with the 3 DL_POLY files everytime you want to change something. To run metadise we will use the subprocess python module. You specify what program you want to run and the file that you want to run it in, you will need to ensure the file path is correct.
#
# #### It is essential that the codes that were downloaded from [here](https://people.bath.ac.uk/chsscp/teach/adv.bho/progs.zip) are in the Codes/ folder in the parent directory, or this following cell will crash.
subprocess.call('../Codes/metadise.exe', cwd='Input/')
os.rename('Input/control_o0001.dlp', 'Input/CONTROL')
os.rename('Input/config__o0001.dlp', 'Input/CONFIG')
os.rename('Input/field___o0001.dlp', 'Input/FIELD')
# Now you should have a CONFIG, CONTROL and FIELD file within the input directory. In theory you could just call the DL_POLY program on this directory and your simulation would run. However we need to tweak the CONTROL file in order to set up our desired simulation. Make a new subdirectory in the week 2 directory named "Example" and copy CONFIG, CONTROL and FIELD to that subdirectory. Now edit the CONTROL file.
#
# We want to change the following
#
# `Temperature 300 ---> Temperature 1500`
# `Steps 5001 ---> Steps 40000`
# `ensemble nve ---> ensemble npt hoover 0.1 0.5`
# `trajectory nstraj= 1 istraj= 250 keytrj=0 ---> trajectory nstraj= 0 istraj= 100 keytrj=0`
#
# Now your simulation is ready. As a point of interest it is always good to check your structure before and after the simulation. You can view the CONFIG file in three dimensions using the VESTA program. It is available for free at http://www.jp-minerals.org/vesta/en/download.html . Download it and use it to view your CONFIG, a demonstrator can help if necessary. VESTA can generate nice pictures which will look very good in a lab report.
#
# <center>
# <br>
# <img src="./figures/vesta.png\" width=\"400px\">
# <i>Figure 1. Fluorite Ca$F_2$ unit cell visualised in VESTA.</i>
# <br>
# </center>
#
#
# To run DL_POLY from within a notebook use the below command. Keep in mind that this simulation will take 20 or so minutes so be patient.
#
# If you are not comfortable with running things through this notebook then you can copy and paste the dlpoly_classic.exe executable into the Example/ sub directory and then double click the .exe file
subprocess.call('../Codes/dlpoly_classic.exe', cwd='Example/')
# Once DL_POLY has completed you will find several files relating to your simulaton.
# - HISTORY - This file contains the configuration of your system at each step during the simulation. You can view this as a movie using the VMD program - Ask a demonstrator for details
# - REVCON - This is the configuration at the end of the simulation - Can be viewed in VESTA - why not check to see how it has changed.
# - STATIS - Contains the stats at each step in the simulation.
# - OUTPUT - Contains properties
#
# It is now important to understand how we can actually use the details of the simulation to get some information on the properties of the material, e.g. Diffusion coefficients and activation energies.
# ## Mean Squared Displacements - Calculating diffusion coefficients
#
# As we have seen molecules in liquds, gases and solids do not stay in the same place and move constantly. Think about a drop of dye in a glass of water, as time passes the dye distributes throughout the water. This process is called diffusion and is common throughout nature.
#
# Using the dye as an example, the motion of a dye molecule is not simple. As it moves it is jostled by collisions with other molecules, preventing it from moving in a straight path. If the path is examined in close detail, it will be seen to be a good approximation to a random walk. In mathmatics a random walk is a series of steps, each taken in a random direction. This was analysed by <NAME> in a study of Brownian motion and he showed that the mean square of the distance travelled by a particle following a random walk is proportional to the time elapsed.
# \begin{align}
# \Big \langle r^2 \big \rangle & = 6 D_t + C
# \end{align}
#
# where $\Big \langle r^2 \big \rangle$ is the mean squared distance, t is time, D is the diffusion rate and C is a constant.
#
# ## What is the mean squared displacement
#
# Going back to the example of the dye in water, lets assume for the sake of simplicity that we are in one dimension. Each step can either be forwards or backwards and we cannot predict which. From a given starting position, what distance is our dye molecule likely to travel after 1000 steps? This can be determined simply by adding together the steps, taking into account the fact that steps backwards subtract from the total, while steps forward add to the total. Since both forward and backward steps are equally probable, we come to the surprising conclusion that the probable distance travelled sums up to zero.
#
# By adding the square of the distance we will always be adding positive numbers to our total which now increases linearly with time. Based upon equation 1 it should now be clear that a plot of $\Big \langle r^2 \big \rangle$ vs time with produce a line, the gradient of which is equal to 6D. Giving us direct access to the diffusion coefficient of the system.
#
# Lets try explore this with an example. Run a short DL_POLY simulation on the input files provided.
# You will a small MSD program called MSD.py to analyse your simulation results. First you need to read in the data, the HISTORY file contains a list of the atomic coordiantes held by the atoms during the simulation.
# +
# Read in the HISTORY file
## Provide the path to the simulation and the atom that you want data for.
data = History("Example/HISTORY", "F")
# -
# data is a dictionary variable containing the atomic trajectories, lattice vectors, total number of atoms, and total number of timesteps.
#
# data = {'trajectories':trajectories, 'lv':lv, 'timesteps':timesteps, 'natoms':natoms}
#
# The next step is to calculate the MSD.
# +
# Run the MSD calculation
f_msd = MSD(data.trajectory, sweeps=2)
output = f_msd.msd()
# -
# run_msd returns a dictionary containing the total MSD, the dimensional MSD values and the time.
# msd_data = {'msd': msd, 'xmsd': xmsd, 'ymsd': ymsd, 'zmsd': zmsd, 'time': time}
#
# This can then be plotted to give a nice linear relationship.
ax = plotting.msd_plot(output)
plt.show()
print("Three Dimensional Diffusion Coefficient", output.xyz_diffusion_coefficient())
print("One Dimensional Diffusion Coefficient in X", output.x_diffusion_coefficient())
print("One Dimensional Diffusion Coefficient in Y", output.y_diffusion_coefficient())
print("One Dimensional Diffusion Coefficient in Z", output.z_diffusion_coefficient())
# ## Simulation Length
#
# It is important to consider the lenght of your simulation (Number of steps). Create a new folder called "Example_2", copy the CONFIG, FIELD and CONTROL files from your previous simulation but this time change the number of steps to 10000. Now rerun the simulation.
subprocess.call('../Codes/dlpoly_classic.exe', cwd='Example_2/')
# +
data = History("Example_2/HISTORY", "F")
# Run the MSD calculation
f_msd = MSD(data.trajectory, sweeps=2)
output = f_msd.msd()
ax = plotting.msd_plot(output)
plt.show()
# -
print("Three Dimensional Diffusion Coefficient", output.xyz_diffusion_coefficient())
print("One Dimensional Diffusion Coefficient in X", output.x_diffusion_coefficient())
print("One Dimensional Diffusion Coefficient in Y", output.y_diffusion_coefficient())
print("One Dimensional Diffusion Coefficient in Z", output.z_diffusion_coefficient())
# You will hopefully see that your MSD plot has become considerably less linear. This shows that your simulation has not run long enough and your results will be unrealiable. You will hopefully also see a change to the value of your diffusion coefficient. The length of your simulation is something that you should keep in mind for the next 3 weeks.
# ## Arrhenius
# The next thing is to use the diffusion coefficients to calcaulte the activation energy for F diffusion. This rea=quires diffusion coefficients from a temperature range. Common sense and chemical intuition suggest that the higher the temperature, the faster a given chemical reaction will proceed. Quantitatively this relationship between the rate a reaction proceeds and its temperature is determined by the Arrhenius Equation. At higher temperatures, the probability that two molecules will collide is higher. This higher collision rate results in a higher kinetic energy, which has an effect on the activation energy of the reaction. The activation energy is the amount of energy required to ensure that a reaction happens.
#
# \begin{align}
# k = A * e^{(-Ea / RT)}
# \end{align}
#
# where k is the rate coefficient, A is a constant, Ea is the activation energy, R is the universal gas constant, and T is the temperature (in kelvin).
#
#
# ## Week 2 Exercise
#
# Using what you have learned over the last 45 mins your task this week is to calculate the activation energy of F diffusion in Ca$F_2$. You will need to select a temperature range and carry out simulations at different temperatures within that range.
# #### Questions to answer
# - In what temperature range is Ca$F_2$ completely solid i.e. no diffusion?
# - In what range is fluorine essentially liquid i.e. fluorine diffusion with no calcium diffusion?
# - What is the melting temperature?
# - Plot an Arrhenius plot and determine the activation energies in temperature range - You will need to rearange the equation.
#
#
# You are encouraged to split the work up within your group and to learn how to view the simulation "movie" using VMD (Ask a demonstrator). VMD is a fantastic program that allows you to visualise your simulation, included below is a video showing a short snippet of an MD simulation of Ca$F_2$. A single F atom has been highlighted to show that diffusion is occuring.
# %%HTML
<div align="middle">
<video width="80%" controls>
<source src="./figures/VMD_example.mp4" type="video/mp4">
</video></div>
# Furthermore, VMD can also be used to generate images showing the entire trajectory of the simulation, e.g.
#
#
#
# <center>
# <br>
# <img src="./figures/CaF2.png\" width=\"400px\">
# <i>Figure 2. A figure showing all positions occupied by F during an MD simulation at 1500 K. F positions are shown in orange and Ca atoms are shown in green.</i>
# <br>
# </center>
#
#
# To save you the time you can use the function declared at the start of this notebook to pull out a diffusion coefficient directly from the simulation output file. MSD.py is a small code to allow visualisation of the MSD plot but it is not neccesary every time you want the diffusion coefficient.
#
# It is up to you how you organise/create your directories but it is reccomended that you start a new notebook. Use the commands/functions used in this notebook to generate your input files, run DL_POLY and extract the diffusion coefficients. The write your own code to generate an Arrhenius plot and calculate the activation energies.
# If you finish early then feel free to start week 3.
| Week_3/.ipynb_checkpoints/week_3-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Pull in one grid cell from a CMIP6 model and run through rolling cdf test**
import intake
import xarray as xr
import os
import pandas as pd
import numpy as np
import zarr
import rhg_compute_tools.kubernetes as rhgk
import warnings
warnings.filterwarnings("ignore")
write_direc = '../figures/'
client, cluster = rhgk.get_big_cluster()
cluster.scale(30)
client
cluster.close()
# get some CMIP6 data from GCS.
#
# here we're going to get daily `tmax` from `IPSL` for historical and SSP370 runs. The ensemble member `r1i1p1f1` isn't available in GCS so we're using `r4i1p1f1` instead.
#
# Note that the `activity_id` for historical runs is `CMIP`, not `ScenarioMIP` as it is for the ssp-rcp scenarios.
activity_id = 'ScenarioMIP'
experiment_id = 'ssp370'
table_id = 'day'
variable_id = 'tasmax'
source_id = 'ACCESS-ESM1-5' # 'IPSL-CM6A-LR'
institution_id = 'NCAR'
member_id = 'r1i1p1f1'
# look at options first
df_cmip6 = pd.read_csv('https://cmip6.storage.googleapis.com/cmip6-zarr-consolidated-stores-noQC.csv', dtype={'version': 'unicode'})
len(df_cmip6)
df_subset_future = df_cmip6.loc[(df_cmip6['activity_id'] == activity_id) & (df_cmip6['experiment_id'] == experiment_id)
& (df_cmip6['table_id'] == table_id) & (df_cmip6['variable_id'] == variable_id)
& (df_cmip6['source_id'] == source_id) & (df_cmip6['member_id'] == member_id)]
df_subset_future
df_subset_hist = df_cmip6.loc[(df_cmip6['experiment_id'] == 'historical')
& (df_cmip6['table_id'] == table_id) & (df_cmip6['variable_id'] == variable_id)
& (df_cmip6['source_id'] == source_id) & (df_cmip6['member_id'] == member_id)]
df_subset_hist
# Now actually pull the data
# +
# search the cmip6 catalog
col = intake.open_esm_datastore("https://storage.googleapis.com/cmip6/pangeo-cmip6.json")
cat = col.search(activity_id=['CMIP', activity_id],
experiment_id=['historical', experiment_id], table_id=table_id, variable_id=variable_id,
source_id=source_id, member_id=member_id)
# -
# cat
cat['CMIP.CSIRO.ACCESS-ESM1-5.historical.day.gn']
ds_model = {}
ds_model['historical'] = cat['CMIP.CSIRO.ACCESS-ESM1-5.historical.day.gn'].to_dask().isel(member_id=0
).squeeze(drop=True).drop(['member_id',
'height'])
# Define the historical and simulated datasets
hist_ds = ds_model['historical']
hist_ds
ds_model['ssp370'] = cat['ScenarioMIP.CSIRO.ACCESS-ESM1-5.ssp370.day.gn'].to_dask().isel(member_id=0
).squeeze(drop=True).drop(['member_id',
'height'])
sim_ds = ds_model['ssp370']
sim_ds
# Pull in tasmax from ERA-5 for the reference dataset from 1995-2014
# +
# Per decade analysis
filenames = []
yrs = np.arange(1995,2015)
for num_yrs in range(len(yrs)):
filename = '/gcs/impactlab-data/climate/source_data/ERA-5/day/tmax/v1.1/tmax_daily_{}-{}.nc'.format(yrs[num_yrs], yrs[num_yrs])
filenames.append(filename)
era5_tmax = xr.open_mfdataset(filenames, concat_dim='time', combine='by_coords')
print('Loading ERA5 files...')
print(era5_tmax['tmax'])
# -
ref_ds = era5_tmax
ref_ds
# +
# Select a single grid point for analysis -- Miami, FL
lat = 25.78
lon = 280.0
ref_da = ref_ds['tmax'].sel(latitude=lat, longitude=lon, method='nearest')
hist_da = hist_ds['tasmax'].sel(lat=lat, lon=lon, method='nearest') # narrow the time
sim_da = sim_ds['tasmax'].sel(lat=lat, lon=lon, method='nearest')
# -
hist_da = hist_da.sel(time=slice('1995-01-01','2014-12-31'))
hist_da
ref_da
def remove_leap_days(da):
noleap = da.where(~((da.time.dt.month == 2) &
(da.time.dt.day == 29)), drop=True)
years = np.arange(da.time.dt.year.min(),da.time.dt.year.max()+1)
if len(da.time) == len(np.arange(1,366))*len(years):
noleap = noleap.assign_coords(dayofyear=xr.DataArray(np.array([np.arange(1,366)]*len(years)).flatten(),
dims=('time'),
coords={'time':noleap.time})).persist()
else:
# adjust years to leave off last (not-full) year
years = np.arange(da.time.dt.year.min(),da.time.dt.year.max())
# create full year time array
time_array = np.array([np.arange(1,366)]*len(years))
# creat datenum for last year (if first year, change to min)
last_yr = [np.arange(1,len(da['time'].sel(time=str(da.time.dt.year.max().values)))+1)]*1
# combine to one new time array
all_time = np.append(time_array, last_yr)
noleap = noleap.assign_coords(dayofyear=xr.DataArray(all_time.flatten(),
dims=('time'),
coords={'time':noleap.time})).persist()
return noleap
ref = remove_leap_days(ref_da)
hist = remove_leap_days(hist_da)
sim = remove_leap_days(sim_da)
ref = ref.load()
hist = hist.load()
sim = sim.load()
ref
# +
#chunks = {'time': -1}
#ref = ref.chunk(chunks).persist()
#hist = hist.chunk(chunks).persist()
#sim = sim.chunk(chunks).persist()
# -
# Load necessary packages, etc. for QDM implementation
# ! pip install git+https://github.com/ClimateImpactLab/xclim.git@qdm_add_year_selection
# +
# %matplotlib inline
import scipy as sp
import matplotlib.pyplot as plt
import dask
import dask.array as da
import dask.distributed as dd
import xclim
from xclim import sdba
from scipy.stats import scoreatpercentile
from scipy.stats.kde import gaussian_kde
from xclim.testing.tests.test_sdba import utils as tu
# -
write_direc = '../figures/'
# +
nquantiles = 100 #620 is empirical
kind = "+" #"*" #"+"
if kind=='*':
kindstr='mult'
else:
kindstr='add'
# +
grouper = "time.dayofyear"
window = 31
grouper = sdba.Grouper(grouper, window=window)
QMdg = sdba.adjustment.EmpiricalQuantileMapping(kind=kind, group=grouper, nquantiles=nquantiles)
QMdg.train(ref, hist) #(ref, hist)
DQMdg = sdba.adjustment.DetrendedQuantileMapping(kind=kind, group=grouper, nquantiles=nquantiles)
DQMdg.train(ref, hist)
QDMdg = sdba.adjustment.QuantileDeltaMapping(kind=kind, group=grouper, nquantiles=nquantiles)
QDMdg.train(ref, hist)
# -
hist_qdm = QDMdg.adjust(hist)
sim_qdm = QDMdg.adjust(sim)
fig,ax = plt.subplots(1,1,figsize=(20,10))
ref.plot(ax=ax, linewidth=2, color="r", label="Obs hist")
hist.plot(ax=ax, color="k", label="GCM hist")
sim.plot(ax=ax, color="blue", label="GCM future", alpha=.5)
#simtrend.plot(ax=ax, color="cyan", label="GCM future trend", alpha=.5)
sim_qdm.plot(ax=ax, linestyle=':', color="blue", label="QDM future", alpha=.5)
#simtrend_qdm.plot(ax=ax, linestyle=':', color="cyan", label="QDM future trend", alpha=.5)
ax.legend(frameon=False)
print('all days linear trend')
print(' sim ', sp.stats.linregress(np.arange(len(sim.time)), sim.values).slope*len(sim.time)) # total change
print(' QDM sim', sp.stats.linregress(np.arange(len(sim_qdm.time)), sim_qdm.values).slope*len(sim_qdm.time)) # total change
# **Rolling CDF**
# +
simy_qdm_list = []
#simtrendy_qdm_list = []
roll = 10
# because we're rolling, we can't use the first and last 10 years
bc_years = np.arange(sim.time.dt.year.values[0]+roll,sim.time.dt.year.values[-1]-roll)
for yr in bc_years:
print(yr)
timeslice = slice(str(yr-roll),str(yr+roll))
# testing this on a future with a trend and one without
simy_qdm_list.append(QDMdg.adjust(sim.sel(time=timeslice), year=yr))
# simtrendy_qdm_list.append(QDMdg.adjust(simtrend.sel(time=timeslice), year=yr))
simy_qdm = xr.concat(simy_qdm_list, dim='time')
#simtrendy_qdm = xr.concat(simtrendy_qdm_list, dim='time')
# -
print('all days linear trend')
print(' sim ', sp.stats.linregress(np.arange(len(sim.time)), sim.values).slope*len(sim.time)) # total change
print(' QDM sim', sp.stats.linregress(np.arange(len(sim_qdm.time)), sim_qdm.values).slope*len(sim_qdm.time)) # total change
# **Are the trends preserved across quantiles?**
from plotting import quantile_compare_plot, compare_quantile_deltas_scatter_hist, compare_gcm_qdm_quantile_deltas
# what are the best keys to use? 'GCM hist', 'QDM hist', 'GCM future','QDM future' or what is here?
raw_dict = {'ref':ref,
'hist':hist,
'sim':sim}
adjusted_dict = {'hist':hist_qdm,
'sim':sim_qdm}
bcgrouper = QDMdg.group
| notebooks/qdm_validation/single_gridcell_cmip6_rollingcdf_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="XdSyX7LEnCa3"
# The second major Python data structure is the dictionary. As you probably recall, dictionaries differ from lists in that you can access items in a dictionary by a key rather than a position. Later in this book you will see that there are many ways to implement a dictionary. The thing that is most important to notice right now is that the get item and set item operations on a dictionary are O(1). Another important dictionary operation is the contains operation. Checking to see whether a key is in the dictionary or not is also O(1).
# + [markdown] id="PV5CWu5BnV69"
# For our last performance experiment we will compare the performance of the contains operation between lists and dictionaries. In the process we will confirm that the contains operator for lists is O(n) and the contains operator for dictionaries is O(1).
#
# The experiment we will use to compare the two is simple. We’ll make a list with a range of numbers in it. Then we will pick numbers at random and check to see if the numbers are in the list. If our performance tables are correct the bigger the list the longer it should take to determine if any one number is contained in the list.
#
# We will repeat the same experiment for a dictionary that contains numbers as the keys. In this experiment we should see that determining whether or not a number is in the dictionary is not only much faster, but the time it takes to check should remain constant even as the dictionary grows larger.
# + [markdown] id="s6rTSf-moYiC"
# ## Contains
# + id="Z-T-wgmQofQi"
import timeit
import random
# + colab={"base_uri": "https://localhost:8080/"} id="0ZqeUJldo4D3" outputId="b6743981-6888-44b2-b2eb-141fa732eeb6"
print("Size\tList Time\t Dictionary Time")
for i in range(10000,1000001,20000):
t = timeit.Timer("random.randrange(%d) in x"%i,
"from __main__ import random,x")
x = list(range(i))
lst_time = t.timeit(number=1000)
x = {j:None for j in range(i)}
d_time = t.timeit(number=1000)
print("%d,%10.3f,%10.3f" % (i, lst_time, d_time))
# + [markdown] id="6D00g6e8o6w4"
# Figure 4 summarizes the results of running Listing 6. You can see that the dictionary is consistently faster. For the smallest list size of 10,000 elements a dictionary is 89.4 times faster than a list. For the largest list size of 990,000 elements the dictionary is 11,603 times faster! You can also see that the time it takes for the contains operator on the list grows linearly with the size of the list. This verifies the assertion that the contains operator on a list is O(n). It can also be seen that the time for the contains operator on a dictionary is constant even as the dictionary size grows. In fact for a dictionary size of 10,000 the contains operation took 0.004 milliseconds and for the dictionary size of 990,000 it also took 0.004 milliseconds.
# + [markdown] id="hWIfaOAPsjV-"
# <img src="https://runestone.academy/runestone/books/published/pythonds/_images/listvdict.png">
# + [markdown] id="--tuD5ixsn74"
# Since Python is an evolving language, there are always changes going on behind the scenes. The latest information on the performance of Python data structures can be found on the Python website. As of this writing the Python wiki has a nice time complexity page that can be found at the <a href="http://wiki.python.org/moin/TimeComplexity">Time Complexity Wiki.</a>
# + id="77FrAVoes9C_"
| Dictionary.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from __future__ import print_function
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
import torchvision
from torchvision import datasets, transforms
import matplotlib.pyplot as plt
import numpy as np
import torchvision.models as models
import os, sys, json
from models.track_net import TrackNet
from utils.get_dataloader import get_dataloader
import cv2
from PIL import Image
import datetime
from models.unet import UNet
from utils.detector import detect
# +
cuda0 = torch.device('cuda:0')
#model = TrackNet().to(cuda0)
model = UNet(9).to(cuda0)
model.load_state_dict(torch.load('weight/epoch_12_4237'))
model.eval()
def open_image(path, shape):
image = Image.open(path).convert("RGB")
return np.asarray(image.resize(shape))
# -
image3 = open_image('../data/DJI_0014/img_10697.jpg', (640,360))
image2 = open_image('../data/DJI_0014/img_10696.jpg', (640,360))
image1 = open_image('../data/DJI_0014/img_10695.jpg', (640,360))
image = np.dstack([image1, image2, image3])
print(image.shape)
image = image.transpose((2, 0, 1))
image = image[np.newaxis, :]
print(image.shape)
image = torch.from_numpy(image).float().to(cuda0)
print(image.shape)
output = model(image) * 255
import matplotlib.pyplot as plt
print(output.shape)
output_numpy = output.squeeze().cpu().detach().numpy()
plt.imshow(output_numpy, cmap = "gray")
plt.show()
plt.imshow(image1)
plt.show()
cap = cv2.VideoCapture('../data/DJI_0016.MP4')
count = -1
image1 = image2 = image3 = None
while True:
count += 1
ret, frame = cap.read()
if ret:
image = cv2.resize(frame, (640, 360))
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
if count == 0:
image1 = image
elif count == 1:
image2 = image
elif count == 2:
image3 = image
else:
image1, image2 = image2, image3
image3 = image
image = np.dstack([image3, image2, image1])
image = image.transpose((2, 0, 1))
image = image[np.newaxis, :]
image = torch.from_numpy(image).float().to(cuda0)
output = model(image)
output_numpy = output.squeeze().cpu().detach().numpy()
write_image = cv2.addWeighted(src1=cv2.cvtColor(image3, cv2.COLOR_RGB2GRAY),alpha=0.2,src2=output_numpy,beta=0.8,gamma=0, dtype=cv2.CV_32F)
#write_image = cv2.bitwise_and(src1=image_numpy, src2=output_numpy)
cv2.imwrite('output/{:05d}.jpg'.format(count), write_image)
if count % 100 == 0:
print(datetime.datetime.now(), count)
# +
def displayCircle(image, ballList, thickness=10, radius=25, color=(0, 0, 255)):
for i in range(len(ballList)):
x = int(ballList[i][0])
y = int(ballList[i][1])
if x == y == 0:
continue
cv2.circle(image, (x, y), radius, color, thickness)
return image
cap = cv2.VideoCapture('../data/DJI_0013.MP4')
count = -1
image1 = image2 = image3 = None
ball_list = []
all_ball_list = []
while True:
count += 1
ret, frame = cap.read()
if ret:
image = cv2.cvtColor(frame, cv2.COLOR_BGR2RGB)
if count == 0:
image1 = image
elif count == 1:
image2 = image
elif count == 2:
image3 = image
else:
image1, image2 = image2, image3
image3 = image
image = np.dstack([cv2.resize(image3, (640, 360)), cv2.resize(image2, (640, 360)), cv2.resize(image1, (640, 360))])
image = image.transpose((2, 0, 1))
image = image[np.newaxis, :]
image = torch.from_numpy(image).float().to(cuda0)
output = model(image)
output_numpy = output.squeeze().cpu().detach().numpy()
max_y, max_x = detect(output_numpy)
max_y, max_x = max_y * 1080 // 360, max_x * 1920 // 640
ball_list.append((max_x, max_y))
#all_ball_list.append((max_x, max_y))
if len(ball_list) > 200:
ball_list = ball_list[1:]
write_image = displayCircle(cv2.cvtColor(image2, cv2.COLOR_RGB2BGR), ball_list, 5, 5, (0,120,243))
#write_image = cv2.addWeighted(src1=cv2.cvtColor(image2, cv2.COLOR_RGB2GRAY),alpha=0.2,src2=output_numpy,beta=0.8,gamma=0, dtype=cv2.CV_32F)
cv2.imwrite('output/{:05d}.jpg'.format(count), write_image)
if count % 100 == 0:
print(datetime.datetime.now(), count)
| Untitled1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"is_executing": false}
# Source: https://stackoverflow.com/questions/2896179/fitting-a-gamma-distribution-with-python-scipy
import scipy.stats as stats
import seaborn as sns
alpha = 1
loc = 0
beta = 100
tetha = 1/beta
size=20
# data = stats.gamma.rvs(alpha, loc=loc, scale=beta, size=20)
data = stats.gamma.rvs(alpha, scale=tetha, size=size)
print(data)
ax = sns.distplot(data,
kde=True,
bins=size,
color='black',
# color='skyblue',
hist_kws={"linewidth": 15,'alpha':1})
ax.set(xlabel='Gamma Distribution', ylabel='Frequency')
# + pycharm={"name": "#%%\n", "is_executing": false}
fit_alpha, fit_loc, fit_beta=stats.gamma.fit(data)
print(fit_alpha, fit_loc, fit_beta)
print(alpha, loc, beta)
| hw1/others/gamma-dist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/nabiila-29/data-fellowship-5/blob/main/Nabiila_Practice_Case_Statistics_.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="LNTjkYhh2oOm"
# # Step 1: Data Preprocessing
# + id="7JgcisZYlyXE" colab={"base_uri": "https://localhost:8080/", "height": 493} outputId="f0e579ea-45d2-4f68-b873-d55ca7a1cf53"
#import the library, read file from github, dataoverview
import pandas as pd
df_raw1 = pd.read_csv("https://raw.githubusercontent.com/Syukrondzeko/Fellowship-5/main/Seasons_Stats.csv")
df_raw1
# + [markdown] id="06c9Ofa8fgKE"
# There are 24.691 rows and 53 column in this data type.
# + colab={"base_uri": "https://localhost:8080/", "height": 493} id="sxkKbY3MobUN" outputId="97efaa1d-3837-4d8d-b1cf-8163df01d97a"
#filter data with year 2017 only
df_raw2 = df_raw1[df_raw1['Year'] == 2017]
df_raw2
# + [markdown] id="HxBPrIlVfoM3"
# The rows decreasing from 24.691 to 595 rows
# + colab={"base_uri": "https://localhost:8080/"} id="0WhgVWaWk3yU" outputId="aead122a-5e08-4b9d-e7e9-6e973d6613a3"
#check data type
df_raw2.dtypes
# + [markdown] id="84peAiu8f1_3"
# The date type has been match. We don’t need to do data transformation
# + colab={"base_uri": "https://localhost:8080/"} id="ZSoC7fgIoT5L" outputId="977755b9-b89b-4f82-b50e-a70e46480ec5"
#Check duplicate values
df_raw2.duplicated(['Player'])
# + [markdown] id="f0rAASuIf-TY"
# 'True' indicates there are duplicate values
# + colab={"base_uri": "https://localhost:8080/", "height": 526} id="_Y_hpzoyoDKj" outputId="9ce67e1c-2333-4f38-9177-7b7bacfe580a"
#remove duplicate values
df_raw3 = df_raw2.drop_duplicates(subset=['Player'])
df_raw3
# + [markdown] id="xtKuqOwRgHvR"
# The rows decrease from 595 to 486 rows.
# + colab={"base_uri": "https://localhost:8080/"} id="NAF8B6Nbm3JR" outputId="ddfd1344-fa86-4d19-a048-6fdee2ab383f"
#check the null column
df_raw3.info()
# + [markdown] id="gHlonS76gUaO"
# 0 indicates there is no value on the column. We will drop them.
# + colab={"base_uri": "https://localhost:8080/", "height": 526} id="P9I6dHWeoDqF" outputId="5972b6cd-24db-4dc0-d1d3-a673c156cc2c"
#drop null columns
df = df_raw3.drop(columns=['blanl', 'blank2'])
df
# + [markdown] id="8ZVESdUYgoBT"
# The columns decrease from 53 to 51
# + [markdown] id="Gllf0Wy1qiwG"
# We have done our data reprocessing.
# After this reprocessing, the data rows are 486 and columns is 51.
# + [markdown] id="efV8n0xH4Zo-"
# # Step 2: Answering Question
# + [markdown] id="cFWqlXgJ4ybg"
# ## 1. Who is the youngest and oldest player in the NBA in 2017 for each team (Tm) ?
#
# + colab={"base_uri": "https://localhost:8080/"} id="f6KmrGxPsfcE" outputId="f6324d0d-c160-4092-86c5-10c52f80f5a7"
#Chi-Square Normality test
from scipy import stats
import scipy as scipy
stats.chisquare(df['Age'])
# + [markdown] id="1JqOT4tQs91O"
# If the p-value> 0.05, the distribution of the data is normal. Above we have 0.9 **The data distribution is normal**
# + colab={"base_uri": "https://localhost:8080/", "height": 197} id="dUvlRglqBulk" outputId="2a51e897-6ed8-42af-b682-79db4ee789ac"
df[["Tm", "Player", "Age"]].head()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="rb3xmRXPEzAb" outputId="b8abf043-abf6-4def-d6f0-51554003243e"
#fing the youngest player on every team
youngest = df.groupby('Tm')['Age', 'Player'].agg(min)
youngest
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="x4Qt8gNMY-nT" outputId="9c828ff7-79ee-49c6-e4d2-be14c459d875"
#find the oldest player on every team.
oldest = df.groupby('Tm')['Age', 'Player'].agg(max)
oldest
# + colab={"base_uri": "https://localhost:8080/", "height": 227} id="jEUlX_FlLxGJ" outputId="bcbd7d36-6200-4575-eb80-3b1e0fbc345c"
#optinal. Join the data
no1 = pd.merge(youngest, oldest, on='Tm', how = 'inner')
no1.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 227} id="Qf5rm0hcZysG" outputId="9e854715-1fa7-4abd-92e9-55ab81e7ec6a"
#rename the column name
no1.rename(columns ={'Age_x' : 'YoungestAge', 'Age_y' : 'OldestAge', 'Player_x': 'YoungestPlayer', 'Player_y' : 'OldestPlayer'}, inplace=True )
no1.head()
# + [markdown] id="jh53ZWvb42ro"
# ## 2. Which player has the most minutes played (MP) in each position (Pos)?
# + colab={"base_uri": "https://localhost:8080/"} id="c71MOXR9xTCM" outputId="8229c54d-ce3f-4a67-81e8-cebbc344f6c4"
#shapiro vilk test (normality test)
stats.shapiro(df['MP'])
# + [markdown] id="rqVXgDXRxpzj"
# We see the p-value. The p-value is the second index of the result above. The normal data has p-value 0> 0.05
#
# the p-value of the data is 3.89. **The data has normal distirbution**
# + colab={"base_uri": "https://localhost:8080/", "height": 313} id="aysPKxrsfd_K" outputId="f333e3df-8df0-46bf-ef3c-402645482146"
#player with most minutes played on every position
no2 = df.groupby('Pos')['MP', 'Player'].agg(max)
no2
# + [markdown] id="Hhukopfs4-wP"
# ## 3. Which team has the highest average total rebound percentage (TRB%), assist percentage (AST%), steal percentage (STL%), and block percentage (BLK%)?
#
# + [markdown] id="Wp8kHipCt4BW"
# ### TRB%
# + colab={"base_uri": "https://localhost:8080/"} id="yvy8-9NBrsDw" outputId="f868cb25-af6d-48ee-e047-26c6041edc1d"
#team with highest average of TRB%
trb = df.groupby(['Tm']).mean()['TRB%'].sort_values(ascending=False)
trb.head(1)
# + [markdown] id="6QU2Ze0Ds19n"
# Team with highest TRB% is WAS
# + [markdown] id="NebsCzmvt9Mo"
# ### AST%
# + colab={"base_uri": "https://localhost:8080/"} id="AaFVYSuigkgX" outputId="12d22e92-b153-415b-9975-5872ce4152e9"
#team with highest average of AST%
ast = df.groupby(['Tm']).mean()['AST%'].sort_values(ascending=False)
ast.head(1)
# + [markdown] id="v1UcZ736tO8B"
# Team with highest AST% is DEN
# + [markdown] id="MqIa7Q4DuE3F"
# ### STL%
# + colab={"base_uri": "https://localhost:8080/"} id="Hv91F0sCtbg2" outputId="155ecf05-38b7-43ba-eece-4c1733b79296"
#team with highest average of STL%
stl = df.groupby(['Tm']).mean()['STL%'].sort_values(ascending=False)
stl.head(1)
# + [markdown] id="0xBrlb0buIwr"
# ### BLK%
# + colab={"base_uri": "https://localhost:8080/"} id="Kr8_S1TJtrMA" outputId="1e79dd7e-e81b-4c8e-ec22-a40366f6dcc7"
#Team with highest average of BLK%
blk = df.groupby(['Tm']).mean()['BLK%'].sort_values(ascending=False)
blk.head(1)
# + [markdown] id="sh3gVOcftlaa"
# Team with highest BLK% is GSW
# + [markdown] id="-nPHfyuX5NuU"
# ## 4. Who is the best player in your opinion based on his record stats? note: you can refer to variables point (PTS), assists, rebounds, or anything else. A combination of several variables would be nice.
# + [markdown] id="JejrzlHyz-yi"
# Here, we will use WS as the indicator of best player
# + colab={"base_uri": "https://localhost:8080/", "height": 107} id="5c7BIVJrygoZ" outputId="1fcdae67-0494-4765-a479-a27a70716cb2"
#best player according to Win Share
best_player= df.sort_values('WS', ascending=False)
best_player[['Player', 'WS']].head(2)
# + [markdown] id="8L7n4lhr59Z8"
# <NAME> is the best player according to WS values
# + [markdown] id="RC7Rnb8-5V67"
# ## 5. Which team has the best average stat record of their players? Note: you can refer to points,assists, rebounds, or anything else. A combination of several variables would be nice
# + colab={"base_uri": "https://localhost:8080/"} id="rxhv9Nrj6hax" outputId="2b25b4f8-799f-4de1-86df-cbc98fa2f556"
#best team according to Win Share
best_team = df.groupby(['Tm']).mean()['WS'].sort_values(ascending=False)
best_team.head(2)
# + [markdown] id="uHj6UqGr6u0x"
# We use WS as the indicator of the best stat record. We group it by team. And found that GSW is the best team.
| Nabiila_Practice_Case_Statistics_.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Copyright 2016 The TensorFlow Authors. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
"""A factory-pattern class which returns classification image/label pairs."""
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
from datasets import cifar10
from datasets import flowers
from datasets import imagenet
from datasets import mnist
from datasets import kitti
datasets_map = {
'cifar10': cifar10,
'flowers': flowers,
'imagenet': imagenet,
'mnist': mnist,
'kitti': kitti,
}
def get_dataset(name, split_name, dataset_dir, file_pattern=None, reader=None):
"""Given a dataset name and a split_name returns a Dataset.
Args:
name: String, the name of the dataset.
split_name: A train/test split name.
dataset_dir: The directory where the dataset files are stored.
file_pattern: The file pattern to use for matching the dataset source files.
reader: The subclass of tf.ReaderBase. If left as `None`, then the default
reader defined by each dataset is used.
Returns:
A `Dataset` class.
Raises:
ValueError: If the dataset `name` is unknown.
"""
if name not in datasets_map:
raise ValueError('Name of dataset unknown %s' % name)
return datasets_map[name].get_split(
split_name,
dataset_dir,
file_pattern,
reader)
| datasets/dataset_factory.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 1. Write a Python program to print "Hello Python"?
# 2. Write a Python program to do arithmetical operations addition and division.?
# 3. Write a Python program to find the area of a triangle?
# 4. Write a Python program to swap two variables?
# 5. Write a Python program to generate a random number?
#
# +
## 1. Write a Python program to print "Hello Python"?
print("Hello Python")
# -
## 2. Write a Python program to do arithmetical operations addition and division.?
a = 10
b = 20
print(a+b)
print(a/b)
## 3. Write a Python program to find the area of a triangle?
a = 20
b = 10
area_of_triangle = (1/2)*a*b
print(area_of_triangle)
# +
## 4. Write a Python program to swap two variables?
a = 5
b = 4
c = a
a = b
b = c
print(a,b)
# -
## 5. Write a Python program to generate a random number?
import random
number = random.random()
print(number)
| Python_Programming Assignment_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="https://githubtocolab.com/giswqs/geemap/blob/master/examples/notebooks/15_convert_js_to_py.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab"/></a>
#
# Uncomment the following line to install [geemap](https://geemap.org) if needed.
# +
# # !pip install geemap
# -
import ee
import geemap
geemap.show_youtube('nAzZjKKd4w0')
# You can simply copy and paste your GEE JavaScripts into a code block wrapped with trip quotes and pass it to a variable.
#
# For example, you can grap GEE JavaScripts from [GEE Documentation](https://developers.google.com/earth-engine/image_visualization).
js_snippet = """
// Load an image.
var image = ee.Image('LANDSAT/LC08/C01/T1_TOA/LC08_044034_20140318');
// Define the visualization parameters.
var vizParams = {
bands: ['B5', 'B4', 'B3'],
min: 0,
max: 0.5,
gamma: [0.95, 1.1, 1]
};
// Center the map and display the image.
Map.setCenter(-122.1899, 37.5010, 10); // San Francisco Bay
Map.addLayer(image, vizParams, 'false color composite');
"""
geemap.js_snippet_to_py(js_snippet, add_new_cell=True, import_ee=True, import_geemap=True, show_map=True)
js_snippet = """
// Load an image.
var image = ee.Image('LANDSAT/LC08/C01/T1_TOA/LC08_044034_20140318');
// Create an NDWI image, define visualization parameters and display.
var ndwi = image.normalizedDifference(['B3', 'B5']);
var ndwiViz = {min: 0.5, max: 1, palette: ['00FFFF', '0000FF']};
Map.addLayer(ndwi, ndwiViz, 'NDWI', false);
"""
geemap.js_snippet_to_py(js_snippet)
js_snippet = """
// Load 2012 MODIS land cover and select the IGBP classification.
var cover = ee.Image('MODIS/051/MCD12Q1/2012_01_01')
.select('Land_Cover_Type_1');
// Define a palette for the 18 distinct land cover classes.
var igbpPalette = [
'aec3d4', // water
'152106', '225129', '369b47', '30eb5b', '387242', // forest
'6a2325', 'c3aa69', 'b76031', 'd9903d', '91af40', // shrub, grass
'111149', // wetlands
'cdb33b', // croplands
'cc0013', // urban
'33280d', // crop mosaic
'd7cdcc', // snow and ice
'f7e084', // barren
'6f6f6f' // tundra
];
// Specify the min and max labels and the color palette matching the labels.
Map.setCenter(-99.229, 40.413, 5);
Map.addLayer(cover,
{min: 0, max: 17, palette: igbpPalette},
'IGBP classification');
"""
geemap.js_snippet_to_py(js_snippet)
| examples/notebooks/15_convert_js_to_py.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import modelseedpy
from modelseedpy.helpers import get_classifier, get_template, get_file
get_file('template_core.json', 'data', 'template_folder')
get_file('template_gram_neg.json', 'data', 'template_folder')
get_file('template_gram_pos.json', 'data', 'template_folder')
get_file('knn_filter.pickle', 'data', 'classifier_folder')
get_file('knn_filter_features.json', 'data', 'classifier_folder')
# ### load classifier
# options
# * knn_filter
genome_cls = get_classifier('knn_filter')
# ### load template
# options
# * template_core
# * template_gram_neg
# * template_gram_pos
core_template = get_template('template_core')
core_template['id']
| examples/Others/download_templates.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # [教學目標]
# - 以下程式碼將示範在 python 如何利用 numpy 計算出兩組數據之間的相關係數,並觀察散佈圖
# - 藉由觀察相關矩陣與散佈圖的關係, 希望同學對 弱相關 / 正相關 的變數分布情形有比較直覺的理解
# # [範例重點]
# - 弱相關的相關矩陣 (Out[2]) 與散佈圖 (Out[3]) 之間的關係
# - 正相關的相關矩陣 (Out[4]) 與散佈圖 (Out[5]) 之間的關係
# +
# 載入基礎套件
import numpy as np
np.random.seed(1)
import matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# ### 弱相關
# +
# 隨機生成兩組 1000 個介於 0~50 的數的整數 x, y, 看看相關矩陣如何
x = np.random.randint(0, 50, 1000)
y = np.random.randint(0, 50, 1000)
# 呼叫 numpy 裡的相關矩陣函數 (corrcoef)
np.corrcoef(x, y)
# -
# 將分布畫出來看看吧
plt.scatter(x, y)
# ### 正相關
# +
# 隨機生成 1000 個介於 0~50 的數 x
x = np.random.randint(0, 50, 1000)
# 這次讓 y 與 x 正相關,再增加一些雜訊
y = x + np.random.normal(0, 10, 1000)
# 再次用 numpy 裡的函數來計算相關係數
np.corrcoef(x, y)
# -
# 再看看正相關的 x,y 分布
plt.scatter(x, y)
# ### 作業
# 參考範例程式碼,模擬一組負相關的資料,並計算出相關係數以及畫出 scatter plot
| 2nd-ML100Days/homework/D-009/Day_009_correlation_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Casting Classification as Regression, Regressing to Probabilities
# 1. We can turn classification labels into a one-hot vector.
# 2. We can regress to the vector.
# 3. To produce output classes, we can take the element with highest weight.
# 4. The regressed value can be interpreted as an (approximate) probability.
#
# Regressing to probabilities is a useful trick, especially when we start thinking about confidences and unsupervised data analysis.
#
# [Link to Fish Dataset Details](https://www.kaggle.com/aungpyaeap/fish-market)
# +
import numpy as np
import csv
rows = []
with open('Fish.csv') as csv_file:
csv_reader = csv.reader(csv_file, delimiter=',')
for row in csv_reader:
rows.append(row)
print(len(rows))
print(rows[0]) # first row is a header
print(rows[1])
rows = rows[1:]
labels = {} # Create a dictionary of label strings to numeric values
for row in rows:
if row[0] not in labels:
labels[row[0]]=len(labels)
print(labels)
inputs = np.array([row[1:] for row in rows])
#print(inputs)
real_outputs = np.array([labels[row[0]] for row in rows])
#print(outputs)
# -
def output_to_one_hot(categories, max_val):
data = np.zeros((len(categories), max_val))
data[np.arange(len(categories)), categories] = 1
return data
# +
def output_to_one_hot(categories, max_val):
data = np.zeros((len(categories), max_val))
data[np.arange(len(categories)), categories] = 1
return data
encodings = output_to_one_hot(real_outputs, len(labels))
print(encodings[:10])
print(encodings[-10:])
# -
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(inputs, encodings)
# # Assignment:
# 1. Define a network class that regresses to the 7 outputs.
# 2. Train a sufficiently large network to perform the categorization.
# 3. Measure the test accuracy of the model by counting the number of accurate labels
#
# # Stretch Goals:
# - Test out different network architectures (depth, breadth) and examine training performance.
# +
from __future__ import print_function
import torch; print(torch.__version__)
from torch.autograd import Variable
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
import torch.utils.data as utils
import matplotlib.pyplot as plt
# %matplotlib inline
t_inputs=Variable(torch.from_numpy(inputs.astype(np.float32)))
t_encodings=Variable(torch.from_numpy(encodings.astype(np.float32)))
#t_inputs=Variable(torch.from_numpy(x_train.astype(np.float32)))
#t_encodings=Variable(torch.from_numpy(y_train.astype(np.float32)))
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
# an affine operation: y = Wx + b
# 2 hidden layers?
self.fc1 = nn.Linear(6, 15)
self.fc2 = nn.Linear(15, 15)
self.fc3 = nn.Linear(15, 7)
def forward(self, x):
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.fc3(x)
return x
net = Net()
#print(net)
#t_inputs=net(Variable(torch.Tensor(inputs)))
net.zero_grad()
outputs = net(Variable(torch.Tensor([0,0,0,0,0,0])))
outputs.backward(torch.randn(7)) # Use random gradients to break symmetry?
learning_rate = 1 # Need to initialize carefully
for f in net.parameters():
f.data.sub_(f.grad.data * learning_rate)
from tqdm import trange # Used to provide progress bar
losses = []
# create your optimizer
optimizer = optim.Adam(net.parameters())
criterion = nn.MSELoss()
num_epochs = 100000
t = trange(num_epochs)
for epoch in t: # loop over the dataset multiple times
running_loss = 0.0
# wrap them in Variable
#reshaped_inputs = t_inputs.view(-1, 1) # Structure with each input in its own row
#reshaped_outputs = true_vals.view(-1, 1) # Neglecting to have outputs and true vals to match dimension is a common mistake.
# forward + backward + optimize
outputs = net(t_inputs)
#print(outputs)
#print(reshaped_outputs)
#encodings = np.zeros((len(outputs), 7))
#encodings[np.arange(len(outputs)), outputs] = 1
#encodings = output_to_one_hot(outputs, len(labels))
#loss = criterion(outputs, t_real_outputs)
#print(outputs.shape)
#print(t_real_outputs.shape)
#loss = criterion(outputs, t_real_outputs)
loss = criterion(outputs, t_encodings)
losses.append(loss)
loss.backward()
optimizer.step()
# zero the parameter gradients
optimizer.zero_grad()
t.set_description('ML (loss=%g)' % loss.item()) # Updates Loss information
#t.set_description('ML Loss: ' + str(loss.item())) # Updates Loss information
print('Finished Training')
print(outputs)
# +
real_categories = [max(range(len(vector)), key=lambda index:vector[index]) for vector in encodings]
categories = [max(range(len(vector)), key=lambda index:vector[index]) for vector in outputs]
score=0
for i in range(0, len(categories)):
if categories[i]==real_categories[i]:
score = score + 1
print('score =',score)
print('possible score=',len(categories))
| Section 2/Part 2/2-2 Supervised Learning - Carl.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
from selenium import webdriver
from selenium.webdriver.chrome.service import Service
from selenium.webdriver.common.by import By
from selenium.webdriver.support.ui import WebDriverWait
from selenium.webdriver.support import expected_conditions as EC
"""This script was written by <NAME>, January 2022."""
def pendputaway_automation(boxid,new_pallet,new_location,s,driver):
"""Handles the Putaway Receiving portion of putting a unit away. Specifically Pendputaway status."""
# Make the Pendputaway URL and go to it.
pend_url_prefix = ('https://w16kcst2.int.hp.com/Rec/Putaway?status=7&projectID=&boxID=')
pend_url = pend_url_prefix + str(boxid)
driver.get(pend_url)
settings_button = driver.find_element(By.CLASS_NAME, 'mdi-pencil-outline')
settings_button.click()
# Find the textboxes for filling in location and pallet as well as the save button.
location_field = driver.find_element(By.ID, 'SingleEdit_MDM_Location_LocationNumber')
pallet_field = driver.find_element(By.ID, 'txtPalletID')
save_button = driver.find_element(By.ID, 'btnSavePendPutaway')
# Fill in the textboxes and save the information.
location_field.clear()
location_field.send_keys(new_location)
pallet_field.clear()
pallet_field.send_keys(new_pallet)
save_button.click()
# Wait til the new information saves (or time out after 60 seconds)
try:
WebDriverWait(driver, 60).until(EC.url_to_be('https://w16kcst2.int.hp.com/Rec/Putaway?firstload=True'))
except:
print("This has taken to long. Internet might be down.")
input("Press ENTER to continue once the page has loaded (or quit and restart from " + grab_id + "): ")
| CST_Automation/pendputaway_automation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
from keras.models import load_model
# +
x_train = np.load('../Data/bird_bicycle_train.npy')
x_test = np.load('../Data/bird_bicycle_test.npy')
x_train = x_train / 255
x_test = x_test / 255
y_train = np.zeros(1000, dtype='int8')
y_train[:500] = 1
y_test = np.zeros(1000, dtype='int8')
y_test[:500] = 1
# -
plt.axis('off')
plt.imshow(x_test[500])
model = load_model('../Saved_models/birds_vgg_augment_model.016.h5')
model.evaluate(x_test, y_test)
model = load_model('../Saved_models/birds_xception.h5')
model.evaluate(x_test, y_test)
f, axs = plt.subplots(2, 6, figsize=(13, 6))
for i in range(6):
images = x_train[y_tmp == i]
axs[i // 5, i % 5].imshow(images[i])
axs[i // 5, i % 5].set_title(label_names[i])
axs[i // 5, i % 5].axis('off')
| EDA/bird_bicycle.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # The Wat
# For this project we will be using the jokes on /r/jokes subreddit as our dataset.
#
# Only "funny" jokes will be saved, since the last thing we want from our model is to make lame jokes... although I just realized that a dad jokes bot would be pretty hilarious too :D. Here, we'll select all the joke submissions with a number of upvotes (score) higher than a threshold of 420 (nice).
#
#
# There are 2 (afaik) ways to retrieve this data:
#
# - Using the praw lib (Python Reddit API Wrapper).
# - Pushshift Reddit API, which is a RESTful API.
#
# ## PRAW
#
# This is the easiest way of accessing reddit content with python. However, ever since reddit introduced a limit of use for its API, only 100 submissions can be retrieved per query.
# +
import praw
import pandas as pd
from tqdm import tqdm
reddit = praw.Reddit(client_id = 'b1pL2McByh6fAQ',
client_secret = '<KEY>',
username = 'yourejokeguy',
password = '<PASSWORD>}',
user_agent = 'joker')
rjokes = reddit.subreddit('jokes')
# +
top = rjokes.top(limit=100000)
jokeset = pd.DataFrame(columns=['Title', 'Text'])
pbar = tqdm(top, desc='Getting the jokes...')
for submission in pbar:
jokeset = jokeset.append({'Title': submission.title, 'Text': submission.selftext}, ignore_index=True)
# -
jokeset.head()
# +
jokeset = pd.DataFrame(columns=['Title', 'Text'])
p_bar = tqdm(psraw.submission_search(reddit, subreddit='jokes', limit=10000))
for item in p_bar:
jokeset = jokeset.append({'Title': item.title, 'Text': item.selftext}, ignore_index=True)
# -
jokeset
# ## Pushshift RESTful API
#
# With this approach we can overcome the rather stringent limitation of the previous method.
#
# Code adapted from: https://www.reddit.com/r/pushshift/comments/i8dlzs/how_to_download_all_posts_from_a_subreddit/
# +
import requests, json
import time
subreddit = 'jokes'
min_score = 420
maxThings = -1
printWait = 2
requestSize = 100
def requestJSON(url):
while True:
try:
r = requests.get(url)
if r.status_code != 200:
print('error code', r.status_code)
time.sleep(5)
continue
else:
break
except Exception as e:
print(e)
time.sleep(5)
continue
return json.loads(r.text)
meta = requestJSON('https://api.pushshift.io/meta')
limitPerMinute = meta['server_ratelimit_per_minute']
requestWait = 60 / limitPerMinute
print('server_ratelimit_per_minute', limitPerMinute)
things = ['submission']
num = 0
for thing in things:
i = 0
df = pd.DataFrame(columns=['title', 'text'])
print('\n[starting', thing + 's]')
if maxThings < 0:
url = 'https://api.pushshift.io/reddit/search/'\
+ thing + '/?subreddit=' + subreddit\
+ '&score=>' + str(min_score) \
+ '&metadata=true&size=0'
meta_json = requestJSON(url)
totalResults = meta_json['metadata']['total_results']
print('total ' + thing + 's', 'in', subreddit,':', totalResults)
else:
totalResults = maxThings
print('downloading most recent', maxThings)
created_utc = ''
startTime = time.time()
timePrint = startTime
while True:
url = 'http://api.pushshift.io/reddit/search/'\
+ thing \
+ '?subreddit=' + subreddit \
+ '&score=>' + str(min_score) \
+ '&size=' + str(requestSize)\
+ '&before=' + str(created_utc)
res_json = requestJSON(url)
if len(res_json['data']) == 0:
break
doneHere = False
for post in res_json['data']:
created_utc = post["created_utc"]
if 'selftext' not in post.keys():
print('\n #No text#')
print(post)
continue
joke = {'title': str(post['title']), 'text': str(post['selftext'])}
df = df.append(joke, ignore_index=True)
i += 1
if i >= totalResults:
doneHere = True
break
if doneHere:
break
if time.time() - timePrint > printWait:
timePrint = time.time()
percent = i / totalResults * 100
timePassed = time.time() - startTime
print('{:.2f}'.format(percent) + '%', '|',
time.strftime("%H:%M:%S", time.gmtime(timePassed)))
time.sleep(requestWait)
df.to_csv(f'data/{subreddit}_set.csv', index=False)
num += 1
# -
df
| jokes_generator/jokes_scraper.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline
from cgcglib import *
from fenics import *
import numpy as np
import matplotlib.pyplot as plt
plt.rcParams["figure.figsize"] = [24, 6]
# Standing wave
k = 3.0
w = 3.0
u0 = Expression("0.0", degree=3)
u1 = Expression("w*pi*sin(k*pi*x[1])", degree=3, k=w, w=w)
u_bc = Constant(0.0)
f = Constant(0.0)
def test_mesh_generator(T, mesh_generator, nx=50, prefix=None):
"""Solving the CG-CG wave equation on the mesh produced by mesh_generator.
Args:
T: Terminal time.
mesh_generator: a function which generates the mesh from (domain, nt, nx)
nx: number of nodes in the spatial direction
data: a quadruple (u0, u1, u_bc, f) as the initial, boundary, and right-hand side
file_prefix: A string if not None. The numerical solution is saved as file_prefix+".pvd".
While nx is specified by the user. nt, the number of nodes per temporal direction,
is always chosen so that for the uniforma mesh, dt/dx=0.5 (CFL safe).
"""
# Plot mesh for show
mesh = mesh_generator(SpaceTimeDomain(t0=0.0, t1=5.0, x0=0.0, x1=1.0), 50, 5)
plt.figure(figsize=(24, 10))
plot(mesh)
plt.axis("off")
plt.savefig("{}_mesh.svg".format(prefix), transparent=True, bbox_inches="tight", pad_inches=0.0)
plt.title("Mesh (down sampled for plotting)")
# Space-time domain
domain = SpaceTimeDomain(t0=0, t1=T, x0=0.0, x1=1.0)
# Computational mesh for solving PDE
nt = int(T / (domain.x1 - domain.x0)) * nx * 2 # dt/dx=0.5 for the unperturbed case
mesh = mesh_generator(domain, nt, nx)
# Solve
uh = solve_wave_equation(u0, u1, u_bc, f, domain=domain, mesh=mesh, degree=1)
# Plot the solution
plt.figure()
plot(uh)
plt.xlabel("Time")
plt.ylabel("Space")
plt.savefig("{}_sol.svg".format(prefix), transparent=True, bbox_inches="tight", pad_inches=0.0)
plt.title("Numerical solution")
# Plot the L2 norm in time
plt.figure()
ts = np.linspace(domain.t0, domain.t1, 2000)
xs = np.linspace(domain.x0, domain.x1, 100)
u_vals = np.vectorize(uh)(*np.meshgrid(ts, xs))
plt.plot(ts, np.linalg.norm(u_vals, axis=0))
plt.xlabel("Time")
plt.ylabel("Spatial L2 norm")
plt.savefig("{}_norm.svg".format(prefix), transparent=True, bbox_inches="tight", pad_inches=0.0)
plt.title("Sptial L2 norm in time")
# Uniform mesh (no problem)
def mesh_gen(domain, nt, nx):
return domain.get_uniform_mesh(nt, nx)
uh = test_mesh_generator(15, mesh_gen, prefix="uniform")
# Randomized mesh
def randomly_perturbed_mesh(percentage=0.0):
"""Randomly perturbed every non-boundary mesh node by a percentage of the mesh size."""
def mesh_gen(domain, nt, nx):
mesh = domain.get_uniform_mesh(nt, nx)
mesh = mesh_randomizer_2d(mesh, percentage=percentage, preserve_boundary=True)
return mesh
return mesh_gen
test_mesh_generator(15, randomly_perturbed_mesh(0.14), prefix="random")
# Structured deterministic perturbation
# In this example, nodes are perturbed spatially every 2 spatial layers.
# From the von Neumann analysis, the magnification in one layer can be cancelled by the magnification
# in the next layer. So the overall effect is small.
#
# Perturbation effect
# [0, 0.25] no visible effect
# [0.25, 0.5] still stable for long time with visible artifacts
# [0.5, 1) unstable
#
# In particular, the critical value 0.5 is CFL-like, in the sense that the method is fine below it but
# blows up rapidly above it.
def gapped_perturbation(pt, px):
def mesh_gen(domain, nt, nx):
mesh = domain.get_uniform_mesh(nt, nx)
x_shift = (domain.x1 - domain.x0) / nx * px
t_shift = (domain.t1 - domain.t0) / nt * pt
for i in range(1, nt):
for j in range(1, nx):
k = i + j * (nt + 1)
if i % 2 == 1:
mesh.coordinates()[k] += np.array([t_shift, x_shift])
return mesh
return mesh_gen
test_mesh_generator(20, gapped_perturbation(0.0, 0.2), prefix="two")
# Structured deterministic perturbation (gapped)
# In this example, nodes are perturbed spatially every 3 spatial layers.
# Becaues the perturbations are slightly further apart than the every other layer case,
# the effect of the perturbation is much stronger.
#
# The blow up time depends quite smoothly on the perturbation, in contrast with the previous
# case.
#
# Perturbation T to observe it (rough)
# 4% 100
# 3% 130
# 2% 200
# 1% 400
#
# My conjecture is that the method is unstable for any nonzero perturbation of this structre.
# This might be provable using the von Neumann analysis.
#
def gapped_perturbation(pt, px):
def mesh_gen(domain, nt, nx):
mesh = domain.get_uniform_mesh(nt, nx)
x_shift = (domain.x1 - domain.x0) / nx * px
t_shift = (domain.t1 - domain.t0) / nt * pt
for i in range(1, nt):
for j in range(1, nx):
k = i + j * (nt + 1)
if i % 3 == 1:
mesh.coordinates()[k] += np.array([t_shift, x_shift])
return mesh
return mesh_gen
test_mesh_generator(20, gapped_perturbation(0.0, 0.04), prefix="three")
| spacetime_wave_equation/cg_cg_wave.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import sklearn
import numpy as np
from sklearn.ensemble import RandomForestClassifier
from sklearn.tree import DecisionTreeClassifier
appRecord = pd.read_csv("application_record.csv");
creditRecord = pd.read_csv("credit_record.csv");
appRecord.head()
creditRecord.head()
df = pd.merge(
appRecord,
creditRecord,
how="inner"
)
df.head()
df.columns
y = df['STATUS'].map({"0":0,"1":1,'2':2,'3':3,'4':4,'5':5,"C":-1,"X":-2})
from sklearn.preprocessing import LabelEncoder,OneHotEncoder
label_encoder =LabelEncoder();
X = df.copy().drop(['ID','MONTHS_BALANCE','STATUS','OCCUPATION_TYPE'],axis=1)
X.head()
for col in ['CODE_GENDER','FLAG_OWN_CAR','FLAG_OWN_REALTY','NAME_INCOME_TYPE','NAME_EDUCATION_TYPE','NAME_FAMILY_STATUS','NAME_HOUSING_TYPE']:
X[col] = label_encoder.fit_transform(X[col])
X.columns
# as a test, make a forest model with just a single feature.
clf = RandomForestClassifier(max_depth=5, random_state=0);
clf.fit(X, y)
# export as dot file
# on the command line, type dot tree.dot -Tpng -o tree.png to conver into png.
from sklearn.tree import export_graphviz;
import os;
localFilePath = "C:/Users/yun91/Documents/GitHub/vis/data"
export_graphviz(
clf.estimators_[0],
out_file = "tree2.dot",
rounded = True,
filled = True)
| data/.ipynb_checkpoints/Ben_EDA-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.2 64-bit
# metadata:
# interpreter:
# hash: 31f2aee4e71d21fbe5cf8b01ff0e069b9275f58929596ceb00d14d90e3e16cd6
# name: python3
# ---
# # Lab Four
# ---
#
# For this lab we're going to be messing around with Markdown!
#
# Our Goals are:
# - Using Markdown!
#
# I want you to describe to me your process of converting the assignment and making it DRY.
#
# Please show the ability to:
# - Make a header
# - Make a list (ordered or unordered)
# - Link the Markdown Guide from the lesson.
# - Artuculate your thought process.
# <NAME>
# Professor Kippins
# Software Dev 1
# 3/23/20
#
# - The first thing I did to dry off the code of the lemonade stand was create two more universal variables.
# - I added a variable for the price of the lemonade and a variable for the price of the pretzel.
# - This is helpful because now the price of the pretzel and the price of the lemonade can change easily without going throguhout the entire code.
# - I also combined the two commands for the cash and combine them into one command.
# - I did this by add the amount of money from the pretzels and the amount of money from the lemonade together in the same line.
# - to make writing each customer's order easier, I created five different parameters within five different functions.
# - I made a function for the cash, tips, lemonade and pretzels.
# - In each function I made a parameter linking each piece I have to print at the end to a variable.
# - I then called each function into another function "serve_Customer"
# - Putting them all into on function made it efficent because for the actual code, I only need to call one funcion.
# - in the code, I assigned pretzels to x, lemonade to y, and tips to z.
# - I assgined each of the orders from the customers into the called function.
# - Each customer had one line of code instead of six like the original.
# - after filling in all 15 orders I ran the function and the output was the same as the wet code.
#
#
#
#
#
#
#
#
| JupyterNotebooks/Labs/Lab 4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 663} colab_type="code" id="jGwXGIXvFhXW" outputId="b8ed4be8-1a6e-4c93-a94f-245e28c70df4"
import json
import tensorflow as tf
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
# !wget --no-check-certificate \
# https://storage.googleapis.com/laurencemoroney-blog.appspot.com/sarcasm.json \
# -O /tmp/sarcasm.json
vocab_size = 1000
embedding_dim = 16
max_length = 120
trunc_type='post'
padding_type='post'
oov_tok = "<OOV>"
training_size = 20000
with open("/tmp/sarcasm.json", 'r') as f:
datastore = json.load(f)
sentences = []
labels = []
urls = []
for item in datastore:
sentences.append(item['headline'])
labels.append(item['is_sarcastic'])
training_sentences = sentences[0:training_size]
testing_sentences = sentences[training_size:]
training_labels = labels[0:training_size]
testing_labels = labels[training_size:]
tokenizer = Tokenizer(num_words=vocab_size, oov_token=oov_tok)
tokenizer.fit_on_texts(training_sentences)
word_index = tokenizer.word_index
training_sequences = tokenizer.texts_to_sequences(training_sentences)
training_padded = pad_sequences(training_sequences, maxlen=max_length, padding=padding_type, truncating=trunc_type)
testing_sequences = tokenizer.texts_to_sequences(testing_sentences)
testing_padded = pad_sequences(testing_sequences, maxlen=max_length, padding=padding_type, truncating=trunc_type)
model = tf.keras.Sequential([
tf.keras.layers.Embedding(vocab_size, embedding_dim, input_length=max_length),
tf.keras.layers.Bidirectional(tf.keras.layers.LSTM(32)),
tf.keras.layers.Dense(24, activation='relu'),
tf.keras.layers.Dense(1, activation='sigmoid')
])
model.compile(loss='binary_crossentropy',optimizer='adam',metrics=['accuracy'])
model.summary()
num_epochs = 5
history = model.fit(training_padded, training_labels, epochs=num_epochs, validation_data=(testing_padded, testing_labels), verbose=1)
# + colab={"base_uri": "https://localhost:8080/", "height": 549} colab_type="code" id="g9DC6dmLF8DC" outputId="539b5fa7-076d-421c-fab4-f4863f24912c"
import matplotlib.pyplot as plt
def plot_graphs(history, string):
plt.plot(history.history[string])
plt.plot(history.history['val_'+string])
plt.xlabel("Epochs")
plt.ylabel(string)
plt.legend([string, 'val_'+string])
plt.show()
plot_graphs(history, 'acc')
plot_graphs(history, 'loss')
# + colab={} colab_type="code" id="7ZEZIUppGhdi"
model.save("test.h5")
# + [markdown] colab_type="text" id="NMkkp0OLVZW6"
# ## Now try replacing bidirectional LSTM layer with conv1d
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="KM0mmtWwU80g" outputId="3b0ec99f-6073-4116-aadc-bd9d85803e61"
model = tf.keras.Sequential([
tf.keras.layers.Embedding(vocab_size, embedding_dim, input_length=max_length),
tf.keras.layers.Conv1D(128, 5, activation='relu'),
tf.keras.layers.GlobalMaxPooling1D(),
tf.keras.layers.Dense(24, activation='relu'),
tf.keras.layers.Dense(1, activation='sigmoid')
])
model.compile(loss='binary_crossentropy',optimizer='adam',metrics=['accuracy'])
model.summary()
num_epochs = 30
history = model.fit(training_padded, training_labels, epochs=num_epochs, validation_data=(testing_padded, testing_labels), verbose=1)
# + colab={"base_uri": "https://localhost:8080/", "height": 549} colab_type="code" id="kSe_CkX_VAD4" outputId="83303fe7-addd-40a3-b2af-2b394d95df29"
plot_graphs(history, 'acc')
plot_graphs(history, 'loss')
# + [markdown] colab_type="text" id="tePzbmKLVFqd"
# #### Using conv 1d is significantly faster than lstm while still gives similar result.
| 3. Natural Language Processing in TensorFlow/Week 1.1 - Detecting sarcasm in news headlines with LSTM and CNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# HIDDEN
from datascience import *
from prob140 import *
import numpy as np
import matplotlib.pyplot as plt
plt.style.use('fivethirtyeight')
# %matplotlib inline
import math
from scipy import stats
from scipy import misc
# ## Reversibility ##
# The reflecting random walk of the previous section has states 0, 1, 2, 3, 4 arranged in sequence clockwise on a circle. At each step the chain stays in place with probability $s$, moves to its clockwise neighbor with probability $r$ and to its counterclockwise neighbor with probability $p$. The stationary distribution of the chain assigns chance 0.2 to each state.
#
# If $r > p$, then the chain is more likely to be moving clockwise than counterclockwise. For example, in steady state, the probability of the path $0, 1, 2, 3$ is
#
# $$
# P(X_0 = 0)P(0, 1)P(1, 2)P(2, 3) = 0.2r^3
# $$
#
# The probability of the *reversed* path $3, 2, 1, 0$ is
#
# $$
# P(X_0 = 3)P(3, 2)P(2, 1)P(1, 0) = 0.2p^3
# $$
#
# If $r > p$, then the original path has higher chance.
#
# But if $r = p$, the chance of the original path is the same as that of the reversed path; in steady state, the chain is just as likely to be running in either direction. If someone simulates the chain in steady state and shows you the original path as well as the reversed path, you will not be able to tell which is which.
#
# In this section we define what it means for a Markov Chain to be *reversible* in this way.
# ### Reversed Process ###
# Let $X_0, X_1, \ldots $ be an irreducible Markov Chain with a finite state space and stationary distribution $\pi$. Start the chain off with this stationary distribution; that is, let $X_0$ have distribution $\pi$. Then for all $n \ge 1$, the distribution of $X_n$ is also $\pi$.
#
# Fix $n > 0$ and consider the *reversed* sequence $Y_0, Y_1, \ldots, Y_n$ defined by $Y_k = X_{n-k}$ for $k = 0, 1, \ldots, n$. Call $X_0, X_1, \ldots, X_n$ the *forwards* sequence.
#
# It is a wonderful fact that the reversed sequence is a time homogenous Markov Chain. To see why, we will check that the Markov property holds.
#
# Before we prove the general fact, let's make some exploratory calculations. Start with $n = 1$, so that $Y_0 = X_1$ and $Y_1 = X_0$. For states $i$ and $j$.
#
# \begin{align*}
# P(Y_1 = j \mid Y_0 = i) ~ &= ~ \frac{P(Y_1 = j, Y_0 = i)}{P(Y_0 = i)} \\
# &= ~ \frac{P(X_0 = j, X_1 = i)}{P(X_1 = i)} \\
# &= ~ \frac{\pi(j)P(j, i)}{\pi(i)}
# \end{align*}
#
# because the forwards sequence is in steady state. We have found a transition probability for the reversed sequence using the transition matrix and stationary distribution of the forwards sequence.
#
# For $n = 2$, we have $Y_0 = X_2$, $Y_1 = X_1$, and $Y_2 = X_0$. For states $k$, $i$, and $j$,
#
# \begin{align*}
# P(Y_2 = j \mid Y_0 = k, Y_1 = i) ~ &= ~ \frac{P(Y_2 = j, Y_1 = i, Y_0 = k)}{P(Y_1 = i, Y_0 = k)} \\
# &= ~ \frac{P(X_0 = j, X_1 = i, X_2 = k)}{P(X_1 = i, X_2 = k)} \\
# &= ~ \frac{\pi(j)P(j, i)P(i, k)}{\pi(i)P(i, k)} \\
# &= ~ \frac{\pi(j)P(j, i)}{\pi(i)}
# \end{align*}
#
# The answer doesn't depend on $k$. That's consistent with the Markov property. Also, put together the two facts we have just proved to notice that the transition probabilities are time homogenous.
#
# For general $n$, fix states $i$ and $j$ and an integer $m$ in the range 0 through $n-1$.
#
# \begin{align*}
# & P(Y_{m+1} = j \mid Y_0 = i_0, Y_1 = i_1, \ldots, Y_{m-1} = i_{m-1}, Y_m = i) \\ \\
# &=
# \frac{P(Y_0 = i_0, Y_1 = i_1 \ldots, Y_{m-1} = i_{m-1}, Y_m = i, Y_{m+1} = j)}
# {P(Y_0 = i_0, Y_1 = i_1 \ldots, Y_{m-1} = i_{m-1}, Y_m = i)} \\ \\
# &= \frac{P(X_n = i_0, X_{n-1} = i_1, \ldots, X_{n-m+1} = i_{m-1}, X_{n-m} = i, X_{n-m-1} = j)}
# {P(X_n = i_0, X_{n-1} = i_1, \ldots, X_{n-m+1)} = i_{m-1}, X_{n-m} = i)} \\ \\
# &= \frac{\pi(j)P(j, i)P(i, i_{m-1}) \cdots P(i_1, i_0)}
# {\pi(i)P(i, i_{m-1}) \cdots P(i_1, i_0)} \\ \\
# &= \frac{\pi(j)P(j, i)}{\pi(i)}
# \end{align*}
#
# This involves only $i$ and $j$, and not on $i_0, i_1, \ldots, i_{m-1}$ nor on $m$. So the Markov property is satisfied and the transition probabilities are time homogenous. The one-step "$i$ to $j$" transition probability for the reversed sequence is
#
# $$
# P(Y_1 = j \mid Y_0 = i) = \frac{\pi(j)P(j, i)}{\pi(i)}
# $$
# ### Reversible Chains ###
# The original "forwards" Markov Chain $X_0, X_1, \ldots $ is called *reversible* if for every $n$, the reversed sequence $Y_0, Y_1, \ldots Y_n$
# has *the same one-step transition probabilities as the original*; that is, if
#
# $$
# \frac{\pi(j)P(j, i)}{\pi(i)} = P(i, j) ~~~ \text{for all } i, j
# $$
#
# That is, the chain is reversible if
#
# $$
# \pi(i)P(i, j) = \pi(j)P(j, i) ~~~ \text{for all } i, j
# $$
#
# In other words:
#
# **The chain is reversible if the detailed balance equations have a positive solution.** This is consistent with our image of particles moving according to this chain in steady state: at each instant, the proportion of particles moving from $i$ to $j$ is exactly the same as the proportion moving from $j$ to $i$, for every pair of states $i$ and $j$.
# At the start of this section we looked at a random walk on a circle. Let's see what the definition of reversibility implies for this chain.
#
# - In the previous section we showed that when $p \ne r$, the detailed balance equations have no positive solution. Therefore, when $p \ne r$, the chain is not reversible. This is consistent with our earlier analysis.
#
# - When $p = r$, we found a solution to the detailed balance equations, and therefore the chain is reversible. This formalizes our idea that if $p = r$ then in steady state the chain "looks the same run forwards or backwards."
# ### Reversibility of Birth and Death Chains ###
# Recall that a *birth and death chain* is a Markov Chain on the integers, with one-step transitions restricted to going up by 1, going down by 1, or staying in place. It is not hard to check that every irreducible birth and death chain with a finite state space is reversible. You can simply solve the detailed balance equations just as we did for the Ehrenfest chain in the previous section.
#
# Go back and look through the examples in the text and exercises. The switching chains, the reflecting random walks (both lazy and not), both of the Ehrenfest chains, and the Bernoulli-Laplace chain are all irreducible birth and death chains, and hence are reversible.
# Let's confirm this in the case of a birth and death chain which at first glance seems not to be reversible. Here is the transition diagram of a Markov Chain $X_0, X_1, \ldots $.
#
# 
#
# This chain moves right (that is, has births) with high probability, so it seems as though we should be able to tell whether it's moving forwards or backwards. But remember that **time reversal happens in the steady state**. In the steady state, the chain is overwhelmingly likely to be shuttling between states 3 and 4. You can see this by solving the detailed balance equations.
#
# \begin{align*}
# \pi(1)\cdot 1 &= \pi(2) \cdot 0.1 ~~~~ \implies \pi(2) = 10\pi(1) \\
# \pi(2) \cdot 0.9 &= \pi(3) \cdot 0.1 ~~~~ \implies \pi(3) = 90\pi(1) \\
# \pi(3) \cdot 0.9 &= \pi(4) \cdot 1 ~~~~~~~ \implies \pi(4) = 81\pi(1)
# \end{align*}
#
# It will visit states 2 and 1 as well, but rarely, state 1 being particularly rare. These vists will intersperse the sojourns in 3 and 4, and the paths will be indistinguishable forwards and backwards.
#
# Let's simulate paths of this process. First, we construct the transition matrix and confirm our calculations of $\pi$.
# +
s = np.arange(1, 5)
def trans(i, j):
if i == 1:
if j == 2:
return 1
else:
return 0
elif i == 4:
if j == 3:
return 1
else:
return 0
elif j == i+1:
return 0.9
elif j == i-1:
return 0.1
else:
return 0
bnd = MarkovChain.from_transition_function(s, trans)
# -
pi = bnd.steady_state()
pi
# We can use `simulate_path` to plot a path of the chain. Notice that unlike our previous uses of this method, we are now passing an initial distribution as the first argument, not a particular state. The second argument is the number of steps, as before.
#
# The graph below shows one path of length 200. Run the cell a few times and look at each path forwards as well as backwards. You won't find a systematic difference between the two.
# +
plt.figure(figsize=(10,5))
n = 200 # the number of steps
x = np.arange(n+1) # the steps
y = bnd.simulate_path(pi, n, plot_path=True) # the simulated state at each step
# Axis labels and title
plt.xlabel('$n$')
plt.ylabel('$X_n$', rotation=0)
plt.title('Reversibility: Path of Birth and Death Chain in Steady State');
# -
| notebooks/Chapter_11/02_Reversibility.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tutorial 5: CH<sub>4</sub> Adsorption in a Zeolite
# import everything that is needed here
import matplotlib.pyplot as plt
import numpy as np
# ## Introduction
#
# In the previous session, we introduced the Grand Canonical (GC) ensemble, where the total number of particles, *N*, is allowed to vary but the chemical potential, $\mu$, the system volume, *V* and the temperature, *T* are fixed ($\mu$VT). We explored the advantages of operating under the GC ensemble in accurately simulating the thermal behaviour of a material as opposed to using 'fixed *N*' ensembles. In this session, we will again be using the $\mu$VT ensemble, but this time we will be applying it to adsorption of gas molecules into a zeolite.
#
# A zeolite is a solid (typically silica or alumina) that has a porous crystal structure. This creates channels and pores within the solid that are often large enough to allow small molecules like H<sub>2</sub>O and CO<sub>2</sub> to enter the solid. This gives zeolites a very large surface area for interactions with other molecular species. This high surface area allows zeolites to function as catalysts and 'molecular sieves', where they can separate components of a mixture by molecule size and/or its affinity to adsorb onto the zeolite surface. Any molecule that is too large simply won't be able to enter the zeolite and molecules that are too small or have low affinity for the surface will pass through the material with little adsorption. Zeolites are also very resistant to environmental conditions due to their chemical inertness; they have high melting points, resist high pressures, are insoluble in water and many other solvents and do not readily undergo redox reactions.
#
# There are a wide array of applications for zeolites, such as high-density storage of gases like hydrogen for fuel cells, separating mixtures like crude oil and the products of 'cracking', and 'carbon scrubbers' for greenhouse gas emissions from power stations. For a more comprehensive overview of zeolite materials and applications, see [1], [2] and [3].
#
# One potential application of zeolites that has been the subject of much research is use as 'carbon scrubbers', where greenhouse gases like CO<sub>2</sub> and CH<sub>4</sub> are removed from the flue gases from power stations before they are released into the atmosphere. Carbon scrubbers are just one of the many ways that greenhouse gas emissions can be reduced thereby limiting the effects of climate change. However, more work is needed to find the optimal zeolite structure that can preferentially adsorb, and therefore separate each greenhouse gas (or any other gas of interest) from a gaseous mixture. This can be done relatively quickly by using computational modelling using different (and often hypothetical) zeolite structures and determining how much gas is adsorbed as the partial pressure and temperature change. This is where Grand Canonical Monte Carlo (GCMC) techniques are particularly useful.
#
# In this session, we will be conducting a GCMC simulation of a bulk zeolite structure containing a variable number of CH<sub>4</sub> molecules. In this simulation, our Monte Carlo moves will be insert/delete moves which insert and remove CH<sub>4</sub> molecules from the system. Unlike the previous session, we will allow translational moves for CH<sub>4</sub> so that each molecule can explore its local environment. We will also define a new move type known as rotation moves, these are similar to translation moves but instead propose a rotation by a random value between zero and a pre-determined maximum number of degrees. This move type only applies to molecules or other objects that lack full rotational symmetry, *i.e.* Lennard-Jones particles used in previous sessions have been individual featureless spheres, which are rotationally symmetrical, so rotation moves would not change the configurational energy of the system.
#
# First, we shall give a breakdown of the DL_MONTE input files for this system, now that we have real molecules and structures to deal with in our system. An example of each one are detailed below:
#
# ### CONFIG
#
# As always the CONFIG file contains the starting structure, the beginning of the CONFIG file used in this session:
# # %load -r 1-12 inputs/Tut_4/main/init/CONFIG
Zeolite (Si and O, with some Xe)
0 0
24.4750735 0.0000000 0.0000000
0.0000000 24.4750735 0.0000000
0.0000000 0.0000000 24.4750735
NUMMOL 1 1 200
MOLECULE zeolite 584 584
Si c
0.4513898 -0.3668470 -0.4576217 0
Si c
-0.1875927 -0.3694940 -0.4576217 0
Si c
# With the rest of the file defining the remaining atoms in the molecule 'zeolite'. As you can see, the CONFIG has the same format as the previous CONFIG files. Our system is contained within a cube with dimensions of 24.4750735 Angstroms. There is one molecule present: the ‘zeolite’ molecule, containing 584 atoms (up to a maximum of 584) which are either silicon, ‘Si’, oxygen, ‘O\_’, or xenon, ‘Xe’.
#
# ### CONTROL
#
# The CONTROL file is shown below:
# # %load inputs/Tut_4/main/init/CONTROL
GCMC simulation of CO2 in zeolite
use gaspressure # use the partial pressure in GCMC moves (as opposed to chemical potential)
use orthonormal
finish
temperature 273.0
acceptmolmoveupdate 200 # Period (in moves) at which the maximum move size is recalculated
acceptmolrotupdate 200 # Period (in moves) at which the maximum rotation angle is updated
steps 1000000 # Number of moves to perform in simulation
equilibration 50000 # Equilibration time before statistics are gathered (in moves)
print 1000 # Information is output every 'print' moves
revconformat dlmonte # REVCON file is in DL_POLY CONFIG format
stack 10000 # Size of blocks (in moves) for block averaging
maxmolrot 0.005 # Initial maximum rotation angle (degrees)
move molecule 1 20 # Perform translation moves for 1 molecule type (ch4) 20% of the time
ch4
move gcinsertmol 1 60 0.5 # Perform insertion/removal moves for ch4 60% of the time, with a min. distance of 0.5 from atoms for inserts
ch4 0.0001 # Use a partial pressure of 0.0001 (katm) for ch4
start
# The CONTROL looks a little different to what you're used to, this is primarily because we are now trying to move and insert/delete real molecules in our simulation, rather than simple spherical particles as we have used in previous sessions. The 'use gaspressure' directive specified at the beginning of the CONTROL file means that the partial pressure of the gas, rather than the activity are specified.
#
# $$a = \gamma \frac{P}{P_0}$$
#
# where *a* is the activity, $\gamma$ is the fraction of the component within the gaseous mixture and is assumed to be 1 in this case, as we are dealing with pure CH<sub>4</sub>, and *P*, $P_0$ the pressure and reference pressure respectively.
#
# The activity relates to chemical potential according to
#
# $$a = \exp(\frac{\mu - \mu_0}{RT})$$
#
# where $\mu$ and $\mu_0$ are the chemical potential and reference chemical potential (usually that of an ideal gas), *R* gas constant and *T* temperature.
#
# The 'use orthonormal' directive tells DL_MONTE to keep our coordinates in each dimension (x, y and z) 90<sup>o</sup> from each other. Lines 6 and 7 state how often to update the maximum move distance for translational moves and maximum rotation angle for rotation moves, respectively. Lines 8-13 have the same function as in the previous CONTROL files. 'maxmolrot' states the initial maximum rotation angle for CH<sub>4</sub> in the system. The four lines proceeding this line define the translational and rotation moves for CH<sub>4</sub>, the first number states how many molecule types the move applies to and the second number states the relative weight at which the moves are conducted. 'move gcinsertmol' defines the insert/delete moves for CH<sub>4</sub>, it applies to just the one (CH<sub>4</sub>) molecule type with a weight of 60 like the other move types specified. The third number defines the minimum distance that you can insert a CH<sub>4</sub> molecule from any other atoms already present in the system, any insertions below this distance are automatically rejected moves. The final line states the partial pressure of CH<sub>4</sub>.
#
# The CH<sub>4</sub> molecules are considered to be rigid during the simulation, this restriction typically has to be in place for standard GCMC in order to satisfy detailed balance.
#
# ### FIELD
#
# The FIELD file is shown below:
# # %load inputs/Tut_4/main/init/FIELD
Force fields and bond constraints for for CH4 in a zeolite (use EPM2 for VDW)
CUTOFF 12.0
UNITS kcal
NCONFIGS 1
ATOMS 4
Si core 28 0.0
O_ core 16 0.0
CH core 16 0.0
Xe core 1 0.0
MOLTYPES 2
zeolite
MAXATOM 584
ch4
ATOMS 1 1
CH core 0.00000000 0.0000000 0.0000000
FINISH
VDW 4
CH core CH core lj 0.31494 3.72
O_ core CH core lj 0.224466 3.3765
CH core Xe core 12-6 16777216 0.0
CH core Si core 12-6 16777216 0.0
CLOSE
# The cutoff distance in this system is 12 Angstroms and the units of energy are in kcal. There are four atom types: silicon, Si, atoms with mass of 28 amu, oxygen, O\_, atoms with mass = 16 amu, CH 'atoms' with mass = 16 amu and xenon, Xe, with mass = 1 amu. All atoms have no net charge for the sake of simplicity. As you may have noticed, the mass of the Xe atoms is not the same as its atomic mass because, in this simulation, the actual mass of Xe has no impact on the course of the simulation.
#
# You will have also noticed that the methane molecules only have one CH 'atom', which might be unexpected given that a methane molecule actually contains one carbon and four hydrogen atoms with four C-H single covalent bonds. This alternative description is used because, in computational simulations, calculations should be as efficient as possible. One way of doing this is to reduce the system to the simplest representation possible while attempting to retain as much accuracy in the results as possible. Consider the CH<sub>4</sub> molecule: a heteroatomic, tetrahedral, spherically-symmetrical molecule, containing (roughly speaking) non-polar C-H bonds. This means that it has no net dipole moment and can be adequately described by one CH unit or 'atom' with the molecular mass of CH<sub>4</sub>; 16. This approximation of the full CH<sub>4</sub> structure and bonding is adequate for the purposes of this simulation. More intuitive representations of CH<sub>4</sub> that more accurately describe CH<sub>4</sub> behaviour and properties exist, but these would add unnecessary complexity to our simulation.
#
# At the end of the FIELD file, there are four defined interactions: one between two CH<sub>4</sub> molecules, one between CH<sub>4</sub> and the oxygen atoms in the zeolite, one between CH<sub>4</sub> and xenon and the final one between CH<sub>4</sub> and the silicon atoms in the zeolite. You will see two different interaction types: the familiar 'lj' potential and the '12-6' potential. 12-6 is the name given to an alternative form of the Lennard-Jones potential:
#
# $$\phi(r_{ij}) = \frac{A}{r_{ij}^{12}} - \frac{B}{r_{ij}^6}$$
#
# where $\phi(r_{ij})$ is the potential energy between two particles, i and j, separated by a distance, $r_{ij}$, *A* and *B* are constants. The first term therefore represent the repulsive part of the Lennard-Jones potential and the second term represents the attractive part of the potential. The two numbers specified in the lines for the '12-6' interactions are *A* and *B*, respectively. For more information, please refer to the DL_POLY manual.
#
# By visualising the structure, or otherwise, identify why the zeolite contains Xe atoms.
#
# HINT: The zeolite contains two different-sized pores in its unit cell, and experiments show that only one of these is involved in gas adsorption.
a = input()
# ## Exercise 1)
#
# In this exercise, you will be running simulations of the zeolite solid with the potential to add/remove CH<sub>4</sub> over the course of the calculations. Each of these calculations will be run at a constant temperature but with increasing partial pressure of CH<sub>4</sub>. From the output of these calculations, you will be able to plot an *adsorption isotherm* of CH<sub>4</sub> in this zeolite. An adsorption isotherm is a graph of the amount of gas adsorbed onto a surface plotted against partial pressure of the gas. These are used to find the partial pressure at which maximum adsorption is obtained (the saturation pressure).
#
# First, we need to load the inputs for the first adsorption calculations:
# +
# data input for exercises
# -
# Now let's run the calculation using the inputs specified in the cell above:
# +
# this calculation should take around x minutes to complete
# -
# Once the calculation is complete, let us first look at how the number of adsorbed CH<sub>4</sub> molecules changes over the course of the simulation. Executing the following cell should give produce this graph for the current set of inputs/outputs:
# +
# plot the time sequence of the number of methane present
# -
# Now we will run the simulation at a range of partial pressures of CH<sub>4</sub> at this temperature. To do this, simply input a new value for partial pressure into the cell below:
# +
# accept user input for new partial pressure, create a new directory, copy the input files into it and change the partial pressure in the CONTROL file
# -
# Executing the above cell will have created a new directory in the inputs folder. Repeat the above exercise by changing the filepath in the 'data input' cell and repeat the exercise for a range of partial pressures.
#
# How does the shape of the plot of the number of adsorbed CH<sub>4</sub> molecules over the course of the simulation change with partial pressure?
#
# To begin constructing your adsorption isotherm at this first temperature, execute the following cell:
# +
# extract average number of methane and append to an array of partial pressures, then plot the array.
# -
# This will extract the average number of adsorbed CH<sub>4</sub> taken over the course of the simulation from the OUTPUT.000 file. It then plots this value at each partial pressure calculation that you have run.
#
# Execute the above cell for each partial pressure you choose to gradually build the adsorption isotherm.
#
# Run the simulations until you have constructed a full isotherm and estimate the value of the saturation pressure, $P_s$, from it and enter it into the following cell:
b = input()
# By looking at the time sequences, what do you need to consider to ensure the accuracy of your calculation?
#
# HINT: Remember, the equilibration time in the CONTROL file tells DL_MONTE how much of the output data is used to calculate final averages.
# ## Exercise 2)
#
# Now that you understand the procedure of estimating an adsorption isotherm from these simulations, this exercise will focus on obtaining isotherms for a range of temperatures to see how varying the temperature changes the adsorption behaviour of the zeolite.
#
# Consider how temperature may affect the number of molecules adsorbed onto the zeolite surface.
#
# To create a set of inputs at a different temperature, type a new temperature value into the following cell and execute it:
# +
# accept user input, create a directory named after the input, copy DLMONTE input files from the first calculation into it and change the temperature in the CONTROL file to the value specified by the user
# -
# This will create a new directory named after the value you choose, copy the DL_MONTE files into it and change the 'temperature' line in the CONTROL file to the value you specify.
#
# Now repeat Exercise 1 for this new temperature to create a new adsorption isotherm at your new temperature. Record your estimates of $P_s$.
#
# However, you will first need to ensure that your system has sufficient time to equilibrate at each temperature before analysing any averaged data. Use the time sequences to assist you with this.
#
# Once you have a suitable estimate of the equilibration time, run the following cell to change the 'equilibration' value in your CONTROL file:
# +
# change the value of equilibration in CONTROL file
# -
# You will only need to estimate an equilibration time for your first partial pressure calculation at that temperature. Once you have changed the equilibration time in your first CONTROL file, it will be copied over to any and all subsequent partial pressures that you run at that temperature.
#
# Once you have created a couple of isotherms, you can plot them all on the same graph by running the following cell:
# +
# Plot all the isotherms on one graph here
# -
# We can now also plot your estimates of $P_s$ against temperature. This will help us to determine the ideal conditions to achieve maximum adsorption on this particular zeolite:
# +
# plot Ps vs T here
# -
# From your graph, identify under what conditions will you get the maximum adsorption of CH<sub>4</sub> into this zeolite.
#
# Given that this particular zeolite is thermally-stable up to around *z* K, are the conditions for maximum adsorption feasible?
#
# ## Conclusions:
#
# In this session, you will have appreciated the application of GCMC in the wider context of computational chemistry research and used GCMC to model the adsorption properties of methane onto a siliceous zeolite. You will have considered how to modify the simulation to improve the accuracy of the results. You will also have compared the results of your model with those from experiments and thence considered ways to improve upon the existing model. The next and final session of this course will encourage you to apply all that you have learned in this and previous sessions to solve problems.
# ## References:
#
# [1] <NAME>., <NAME>., <NAME>., <NAME>., *Introduction to Zeolite Science and Practice*, Elsevier Science, Burlington, 2007, **168**, 1-1058.
#
# [2] <NAME>., *Zeolite molecular sieves: structure, chemistry, and use*, Wiley, 1973.
#
# [3] <NAME>., *Zeolite Chemistry and Catalysis*, Springer Netherlands, Dordrecht, 2009.
| tutorials/Jupyter/Tutorial 4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import re
import json
df = pd.read_csv('data.csv')
event_names = list(df['event_name'])
stopwords = ['The', 'And', 'In', 'For', 'At', 'To', 'On', 'A', 'An', 'Of', 'With', 'By', 'Its', 'Can', 'Be', 'It',
'Too', 'Are', 'Day', 'As', 'My', 'Your', '']
words = []
for name in event_names:
# Save spaces for compound words like San_Francisco
name = re.sub(r'((?:^|\W)san)\s|((?:^|\W)de)\s|((?:^|\W)los)\s', r'\1\2\3_', name.lower())
# Remove extraneous non-alphanumeric chars
name = re.sub(r'[^0-9a-zA-Z\s_]+', '', name)
# Remove dates and time references
name = re.sub(r'[0-9]+(pm|am|th)', '', name).strip().title()
for word in name.split(' '):
if word not in stopwords and len(word) > 1 and not word.isdigit():
words.append(word)
with open('words.json', 'w') as outfile:
json.dump({'words': words}, outfile)
| data/.ipynb_checkpoints/Untitled-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Lesson 15 - Pandas Time Series
#
# *Adapted from Pandas documentation (esoteric/financial applications left out).*
#
# In working with time series data, we will frequently seek to:
#
# * generate sequences of fixed-frequency dates and time spans
# * conform or convert time series to a particular frequency
# * compute “relative” dates based on various non-standard time increments (e.g. 5 business days before the last business day of the year), or “roll” dates forward or backward
#
# Pandas provides a relatively compact and self-contained set of tools for performing the above tasks.
# modules used in this tutorial
import pandas as pd
import numpy as np
import pytz
import dateutil
# set max row and random seed
pd.set_option("display.max_rows", 20)
np.random.seed(12)
# ## Examples
#
# Create a range of dates:
rng = pd.date_range('1/1/2011', periods=72, freq='H')
rng[:5]
len(rng)
# Index Pandas objects with dates:
ts = pd.Series(np.random.randn(len(rng)), index=rng)
ts.head()
# Change frequency and fill gaps:
# to 45 minute frequency and forward fill
converted = ts.asfreq('45Min', method='pad')
converted.head()
# Frequency conversion and resampling of a time series:
ts.resample('D', how='mean')
ts.resample('D').mean()
ts.resample('T').mean().dropna()
# ## Overview
#
# The following table shows the type of time-related classes pandas can handle and how to create them:
#
# Class | Remarks | How to create
# ------|---------|--------------
# `Timestamp` | Represents a single time stamp | `to_datetime`, `Timestamp`
# `DatetimeIndex` | Index (~array) of `Timestamp` | `to_datetime`, `date_range`, `DatetimeIndex`
# `Period` | Represents a single time span | `Period`
# `PeriodIndex` | Index (~array) of `Period` | `period_range`, `PeriodIndex`
# ## Timestamps vs. Periods (time spans)
#
# Time-stamped data is the most basic type of timeseries data that associates values with points in time. For pandas objects it means using the points in time.
pd.datetime(2012, 5, 1)
pd.Timestamp(pd.datetime(2012, 5, 1))
pd.Timestamp('2012-05-01')
# However, in many cases it is more natural to associate things like change variables with a time span instead. The span represented by Period can be specified explicitly, or inferred from datetime string format.
#
# For example:
pd.Period('2011-01')
pd.Period('2012-05', freq='D')
# Timestamp and Period can be the index. Lists of Timestamp and Period are automatically coerced to DatetimeIndex and PeriodIndex respectively.
dates = [pd.Timestamp('2012-05-01'), pd.Timestamp('2012-05-02'), pd.Timestamp('2012-05-03')]
ts = pd.Series(np.random.randn(3), dates)
ts
ts.index
type(ts.index)
periods = [pd.Period('2012-01'), pd.Period('2012-02'), pd.Period('2012-03')]
ts = pd.Series(np.random.randn(3), periods)
ts
ts.index
type(ts.index)
# Pandas allows you to capture both representations and convert between them. Under the hood, Pandas represents timestamps using instances of Timestamp and sequences of timestamps using instances of DatetimeIndex. For regular time spans, Pandas uses Period objects for scalar values and PeriodIndex for sequences of spans.
# ## Converting to Timestamps
#
# To convert a Series or list-like object of date-like objects e.g. strings, epochs, or a mixture, you can use the to_datetime function. When passed a Series, this returns a Series (with the same index), while a list-like is converted to a DatetimeIndex:
pd.to_datetime(pd.Series(['Jul 31, 2009', '2010-01-10', None]))
pd.to_datetime(['2005/11/23', '2010.12.31'])
# If you use dates which start with the day first (i.e. European style), you can pass the dayfirst flag.
#
# **Warning:** You see in the second example below that dayfirst isn’t strict, so if a date can’t be parsed with the day being first it will be parsed as if dayfirst were False.
pd.to_datetime(['04-01-2012 10:00'], dayfirst=True)
pd.to_datetime(['04-01-2012 10:00'], dayfirst=False)
# default behavior is dayfirst=False
pd.to_datetime(['04-01-2012 10:00'])
pd.to_datetime(['14-01-2012', '01-14-2012'], dayfirst=True)
# **Note:** Specifying a format argument will potentially speed up the conversion considerably and on versions later than 0.13.0 explicitly specifying a format string of ‘%Y%m%d’ takes a faster path still.
#
# If you pass a single string to to_datetime, it returns single Timestamp. Also, Timestamp can accept the string input. Note that Timestamp doesn’t accept string parsing option like dayfirst or format: use to_datetime if these are required.
#
# The first option below would work with a list of dates, but the second would not.
#pd.to_datetime('11/12/2010', format='%m/%d/%Y')
pd.to_datetime('11/12/2010', format='%d/%m/%Y')
pd.to_datetime('11/12/2010')
pd.Timestamp('2010-11-12')
# ### Invalid data
#
# In version 0.17.0, the default for to_datetime is now errors='raise', rather than errors='ignore'. This means that invalid parsing will raise rather that return the original input as in previous versions.
#
# Pass errors='coerce' to convert invalid data to NaT (not a time):
# + active=""
# # this is the default, raise when unparseable -- convert cell type to Code to see error
# pd.to_datetime(['2009/07/31', 'asd'], errors='raise')
# + active=""
# # again, default is to generate an error
# pd.to_datetime(['2009/07/31', 'asd'])
# -
# don't convert anything and return the original input when unparseable
pd.to_datetime(['2009/07/31', 'asd'], errors='ignore')
# return NaT for input when unparseable
pd.to_datetime(['2009/07/31', 'asd'], errors='coerce')
# ### Epoch timestamps
#
# It’s also possible to convert integer or float epoch times. The default unit for these is nanoseconds (since these are how Timestamps are stored). However, often epochs are stored in another unit which can be specified:
#
# Typical epoch stored units (nanoseconds):
pd.to_datetime(0)
pd.to_datetime(1)
pd.to_datetime(3e9)
pd.to_datetime([1349720105, 1349806505, 1349892905, 1349979305, 1350065705], unit='s')
pd.to_datetime([1349720105100, 1349720105200, 1349720105300, 1349720105400, 1349720105500 ], unit='ms')
# **Note:** Epoch times will be rounded to the nearest nanosecond.
# ## Generating ranges of timestamps
#
# To generate an index with time stamps, you can use either the DatetimeIndex or Index constructor and pass in a list of datetime objects
# ### The slow way
dates = [pd.datetime(2012, 5, 1), pd.datetime(2012, 5, 2), pd.datetime(2012, 5, 3)]
index = pd.DatetimeIndex(dates)
index # Note the frequency information
index = pd.Index(dates)
index # Automatically converted to DatetimeIndex
# ### The quick way - data_range by periods or start-end
# Practically, this becomes very cumbersome because we often need a very long index with a large number of timestamps.
#
# If we need timestamps on a regular frequency, we can use the pandas function date_range to create timestamp indexes. (A similar function and bdate_range does the same thing for business days; the 'b' stands for 'business'.)
#
# Information needed to specify date ranges:
#
# * start, number of periods, AND frequency
# * start, end, AND frequency
#
# Frequency defaults to 'D' for date_range and 'B' for bdate_range.
# freq='M' uses last day of month by default
index = pd.date_range('2018-3-1', periods=1000, freq='M')
index
# freq='B' is business days
index = pd.date_range('2018-3-1', periods=1000, freq='B')
index
# default is freq='D' for date_range
index = pd.date_range('2018-3-1', periods=1000)
index
# default is freq='B' for bdate_range
index = pd.bdate_range('2018-3-1', periods=1000)
index
# ### Commonly used of offset aliases
#
# Alias | Description
# ------|------------
# D | day frequency
# B | business day frequency
# W | weekly frequency
# BW | business week end frequency
# M | month end frequency
# BM | business month end frequency
# MS | month start frequency
# A | year end frequency
# BA | business year end frequency
# AS | year start frequency
# H | hourly frequency
# T, min | minutely frequency
# S | secondly frequency
# L, ms | milliseonds
# U, us | microseconds
# N | nanoseconds
t1 = pd.datetime(2011, 1, 1)
t2 = pd.datetime(2012, 1, 1)
rng = pd.date_range(t1, t2, freq='D')
rng
pd.date_range(t1, t2, freq='W-SUN')
pd.date_range(t1, t2, freq='B')
pd.date_range(t1, t2, freq='BM')
pd.bdate_range(end=t2, periods=20)
pd.bdate_range(start=t1, periods=20)
# ## DatetimeIndex
#
# One of the main uses for DatetimeIndex is as an index for Pandas objects. The DatetimeIndex class contains many timeseries related optimizations:
#
# * A large range of dates for various offsets are pre-computed and cached under the hood in order to make generating subsequent date ranges very fast (just have to grab a slice)
# * Fast shifting using the shift and tshift method on pandas objects
# * Unioning of overlapping DatetimeIndex objects with the same frequency is very fast (important for fast data alignment)
# * Quick access to date fields via properties such as year, month, etc.
# * Regularization functions like snap and very fast asof logic
#
# DatetimeIndex objects has all the basic functionality of regular Index objects and a smorgasbord of advanced timeseries-specific methods for easy frequency processing.
#
# See also documentation for *Reindexing methods*.
#
# **Note:** While Pandas does not force you to have a sorted date index, some of these methods may have unexpected or incorrect behavior if the dates are unsorted. So please be careful.
#
# DatetimeIndex can be used like a regular index and offers all of its intelligent functionality like selection, slicing, etc.
start = pd.datetime(2011, 1, 1)
end = pd.datetime(2012, 1, 1)
rng = pd.date_range(start, end, freq='BM')
ts = pd.Series(np.random.randn(len(rng)), index=rng)
ts.index
ts[:5].index
ts[::2].index
# ### Slicing revisited
# In Pandas, **label-based slicing is inclusive**, whereas **index-based slicing is NOT inclusive** (like index-based slicing in Python generally).
x = pd.Series([0.0, 0.1, 0.2], index=['a', 'b', 'c'])
x[0:2]
x['a':'c']
# DatetimeIndex-based slicing is effectively label-based, so it is inclusive. It will not generate any dates outside of those dates if specified.
# ### DatetimeIndex partial string 'smart' indexing
#
# You can pass in dates and strings that parse to dates as indexing parameters:
ts['1/31/2011']
ts[pd.datetime(2011, 12, 25):]
ts['10/31/2011':'12/31/2011']
# To provide convenience for accessing longer time series, you can also pass in the year or year and month as strings:
ts['2011']
ts['2011-6']
# This type of slicing will work on a DataFrame with a DateTimeIndex as well. Since the partial string selection is a form of label slicing, the endpoints will be included. This would include matching times on an included date. Here’s an example:
dft = pd.DataFrame(np.random.randn(100000,1), columns=['A'],
index=pd.date_range('20130101', periods=100000, freq='T'))
dft.shape
dft
dft['2013']
# This starts on the very first time in the month, and includes the last date & time for the month:
dft['2013-1':'2013-2']
# This specifies a stop time that includes all of the times on the last day:
dft['2013-1':'2013-2-28']
# This specifies an exact stop time (and is not the same as the above):
dft['2013-1':'2013-2-28 00:00:00']
# We are stopping on the included end-point as it is part of the index:
dft['2013-1-15':'2013-1-15 12:30:00']
# To select a single row, use .loc:
dft.loc['2013-1-15 12:30:00']
# ### Datetime indexing
#
# Indexing a DateTimeIndex with a partial string depends on the “accuracy” of the period, in other words how specific the interval is in relation to the frequency of the index. In contrast, indexing with datetime objects is exact, because the objects have exact meaning. These also follow the semantics of including both endpoints.
#
# These datetime objects are specific hours, minutes, and seconds even though they were not explicitly specified (they are 0).
dft[pd.datetime(2013, 1, 1):pd.datetime(2013, 2, 28)]
# With no defaults:
dft[pd.datetime(2013, 1, 1, 10, 12, 0):pd.datetime(2013, 2, 28, 10, 12, 0)]
# ### Truncating & fancy indexing
#
# A truncate convenience function is provided that is equivalent to slicing:
ts.truncate(before='10/31/2011', after='12/31/2011')
# Even complicated fancy indexing that breaks the DatetimeIndex’s frequency regularity will result in a DatetimeIndex (but frequency is lost):
ts[[0, 2, 6]].index
# ### Time/Date Components
#
# There are several time/date properties that one can access from Timestamp or a collection of timestamps like a DateTimeIndex.
#
#
# Property | Description
# ---------|------------
# year | The year of the datetime
# month | The month of the datetime
# day | The days of the datetime
# hour | The hour of the datetime
# minute | The minutes of the datetime
# second | The seconds of the datetime
# microsecond | The microseconds of the datetime
# nanosecond | The nanoseconds of the datetime
# date | Returns datetime.date
# time | Returns datetime.time
# dayofyear | The ordinal day of year
# weekofyear | The week ordinal of the year
# week | The week ordinal of the year
# dayofweek | The day of the week with Monday=0, Sunday=6
# weekday | The day of the week with Monday=0, Sunday=6
# quarter | Quarter of the date: Jan=Mar = 1, Apr-Jun = 2, etc.
# days_in_month | The number of days in the month of the datetime
# is_month_start | Logical indicating if first day of month (defined by frequency)
# is_month_end | Logical indicating if last day of month (defined by frequency)
# is_quarter_start | Logical indicating if first day of quarter (defined by frequency)
# is_quarter_end | Logical indicating if last day of quarter (defined by frequency)
# is_year_start | Logical indicating if first day of year (defined by frequency)
# is_year_end | Logical indicating if last day of year (defined by frequency)
#
# Furthermore, if you have a Series with datetimelike values, then you can access these properties via the .dt accessor, see the docs.
ts
ts.index.month
# ## DateOffset objects
#
# In the preceding examples, we created DatetimeIndex objects at various frequencies by passing in frequency strings like 'M', 'W', and 'BM' to the freq keyword. Under the hood, these frequency strings are being translated into an instance of pandas DateOffset, which represents a regular frequency increment. Specific offset logic like “month”, “business day”, or “one hour” is represented in its various subclasses.
#
# Class name | Description
# -----------|------------
# DateOffset | Generic offset class, defaults to 1 calendar day
# BDay | business day (weekday)
# CDay | custom business day (experimental)
# Week | one week, optionally anchored on a day of the week
# WeekOfMonth | the x-th day of the y-th week of each month
# LastWeekOfMonth | the x-th day of the last week of each month
# MonthEnd | calendar month end
# MonthBegin | calendar month begin
# BMonthEnd | business month end
# BMonthBegin | business month begin
# CBMonthEnd | custom business month end
# CBMonthBegin | custom business month begin
# QuarterEnd | calendar quarter end
# QuarterBegin | calendar quarter begin
# BQuarterEnd | business quarter end
# BQuarterBegin | business quarter begin
# FY5253Quarter | retail (aka 52-53 week) quarter
# YearEnd | calendar year end
# YearBegin | calendar year begin
# BYearEnd | business year end
# BYearBegin | business year begin
# FY5253 | retail (aka 52-53 week) year
# BusinessHour | business hour
# Hour | one hour
# Minute | one minute
# Second | one second
# Milli | one millisecond
# Micro | one microsecond
# Nano | one nanosecond
# Basic function of DateOffset:
d = pd.datetime(2018, 2, 27, 9, 0)
d
pd.Timestamp(d)
d + pd.tseries.offsets.DateOffset(months=2, days=1)
# The key features of a DateOffset object are:
#
# * it can be added / subtracted to/from a datetime object to obtain a shifted date
# * it can be multiplied by an integer (positive or negative) so that the increment will be applied multiple times
# * it has rollforward and rollback methods for moving a date forward or backward to the next or previous “offset date”
#
# Subclasses of DateOffset define the apply function which dictates custom date increment logic, such as adding business days:
#
# class BDay(DateOffset):
# """DateOffset increments between business days"""
# def apply(self, other):
# ...
d - 5 * pd.tseries.offsets.BDay()
d + pd.tseries.offsets.BMonthEnd()
# The rollforward and rollback methods do exactly what you would expect:
d
offset = pd.tseries.offsets.BMonthEnd()
offset.rollforward(d)
offset.rollback(d)
# If you expect to use these functions, explore the pandas.tseries.offsets module and the various docstrings for the classes.
#
# These operations (apply, rollforward and rollback) preserves time (hour, minute, etc) information by default. To reset time, use normalize=True keyword when creating the offset instance. If normalize=True, result is normalized after the function is applied.
day = pd.tseries.offsets.Day()
day.apply(pd.Timestamp('2014-01-01 09:00'))
day = pd.tseries.offsets.Day(normalize=True)
day.apply(pd.Timestamp('2014-01-01 09:00'))
hour = pd.tseries.offsets.Hour()
hour.apply(pd.Timestamp('2014-01-01 22:00'))
pd.Timestamp('2014-01-01 23:00:00')
hour = pd.tseries.offsets.Hour(normalize=True)
hour.apply(pd.Timestamp('2014-01-01 22:00'))
hour.apply(pd.Timestamp('2014-01-01 23:00'))
# ### Parametric offsets
#
# Some of the offsets can be “parameterized” when created to result in different behaviors. For example, the Week offset for generating weekly data accepts a weekday parameter which results in the generated dates always lying on a particular day of the week:
d
d + pd.tseries.offsets.Week()
d + pd.tseries.offsets.Week(weekday=4)
(d + pd.tseries.offsets.Week(weekday=4)).weekday()
d - pd.tseries.offsets.Week()
# normalize option will be effective for addition and subtraction.
d + pd.tseries.offsets.Week(normalize=True)
d - pd.tseries.offsets.Week(normalize=True)
# Another example is parameterizing YearEnd with the specific ending month:
d + pd.tseries.offsets.YearEnd()
d + pd.tseries.offsets.YearEnd(month=6)
# ### Using offsets with Series / DatetimeIndex
#
# Offsets can be used with either a Series or DatetimeIndex to apply the offset to each element.
rng = pd.date_range('2012-01-01', '2012-01-03')
s = pd.Series(rng)
rng
rng + pd.tseries.offsets.DateOffset(months=2)
s + pd.tseries.offsets.DateOffset(months=2)
s - pd.tseries.offsets.DateOffset(months=2)
# If the offset class maps directly to a Timedelta (Day, Hour, Minute, Second, Micro, Milli, Nano) it can be used exactly like a Timedelta - see the Timedelta section for more examples.
s - pd.tseries.offsets.Day(2)
td = s - pd.Series(pd.date_range('2011-12-29', '2011-12-31'))
td
td + pd.tseries.offsets.Minute(15)
# Note that some offsets (such as BQuarterEnd) do not have a vectorized implementation. They can still be used but may calculate signficantly slower and will raise a PerformanceWarning.
rng + pd.tseries.offsets.BQuarterEnd()
# ### Offset Aliases
#
# A number of string aliases are given to useful common time series frequencies. We will refer to these aliases as offset aliases (referred to as time rules prior to v0.8.0).
#
# Alias | Description
# ------|------------
# B | business day frequency
# C | custom business day frequency (experimental)
# D | calendar day frequency
# W | weekly frequency
# M | month end frequency
# BM | business month end frequency
# CBM | custom business month end frequency
# MS | month start frequency
# BMS | business month start frequency
# CBMS | custom business month start frequency
# Q | quarter end frequency
# BQ | business quarter endfrequency
# QS | quarter start frequency
# BQS | business quarter start frequency
# A | year end frequency
# BA | business year end frequency
# AS | year start frequency
# BAS | business year start frequency
# BH | business hour frequency
# H | hourly frequency
# T, min | minutely frequency
# S | secondly frequency
# L, ms | milliseonds
# U, us | microseconds
# N | nanoseconds
# ### Combining Aliases
#
# As we have seen previously, the alias and the offset instance are fungible in most functions:
pd.date_range(start, periods=5, freq='B')
pd.date_range(start, periods=5, freq=pd.tseries.offsets.BDay())
# You can combine together day and intraday offsets:
pd.date_range(start, periods=10, freq='2h20min')
pd.date_range(start, periods=10, freq='1D10U')
# ## Time series-related instance methods
#
# ### Shifting / lagging
#
# One may want to shift or lag the values in a time series back and forward in time. The method for this is shift, which is available on all of the pandas objects.
ts = pd.Series(['a', 'b', 'c', 'd', 'e'], index=pd.date_range(start='3/1/18', periods=5))
ts
ts.shift(3)
# Notes: If freq is specified then the index values are shifted but the data
# is not realigned. That is, use freq if you would like to extend the
# index when shifting and preserve the original data.
ts.shift(3, freq='D')
ts.shift(3, freq='M')
# The same thing can be accomplisehd with the tshift convenience method that changes all the dates in the index by a specified number of offsets, rather than changing the alignment of the data and the index:
ts.tshift(3)
# Note that with tshift (and with shift+freq), the leading entry is no longer NaN because the data is not being realigned.
# ### Frequency conversion
#
# The primary function for changing frequencies is the asfreq function. For a DatetimeIndex, this is a convenient wrapper around reindex which generates a date_range and calls reindex.
dr = pd.date_range('1/1/2010', periods=3, freq='3B')
ts = pd.Series(np.random.randn(3), index=dr)
ts
ts.asfreq('B')
# asfreq provides a further convenience so you can specify an interpolation method (e.g. 'pad', i.e. forward fill) for any gaps that may appear after the frequency conversion.
ts.asfreq(pd.tseries.offsets.BDay(), method='pad')
# ### Filling forward / backward
#
# Related to asfreq and reindex is the fillna function, which can be used with any Pandas object.
ts.asfreq('B').fillna(0)
# ## Time Span Representation
#
# Regular intervals of time are represented by Period objects in pandas while sequences of Period objects are collected in a PeriodIndex, which can be created with the convenience function period_range.
#
# ### Period
#
# A Period represents a span of time (e.g., a day, a month, a quarter, etc). You can specify the span via freq keyword using a frequency alias like below. Because freq represents a span of Period, it cannot be negative like "-3D". ('A-DEC' is annual frequency, anchored end of December; same as 'A'.)
pd.Period('2012', freq='A')
pd.Period('2012-1-1', freq='D')
pd.Period('2012-1-1 19:00', freq='H')
pd.Period('2012-1-1 19:00', freq='5H')
# Adding and subtracting integers from periods shifts the period by its own frequency. Arithmetic is not allowed between Period with different freq (span).
p = pd.Period('2012', freq='A')
p + 1
p - 3
p = pd.Period('2012-01', freq='2M')
p + 2
p - 1
# If Period freq is daily or higher (D, H, T, S, L, U, N), offsets and timedelta-like can be added if the result can have the same freq.
p = pd.Period('2014-07-01 09:00', freq='H')
p + pd.tseries.offsets.Hour(2)
p + pd.tseries.offsets.timedelta(minutes=120)
p + np.timedelta64(7200, 's')
# If Period has other freqs, only the same offsets can be added.
p = pd.Period('2014-07', freq='M')
p + pd.tseries.offsets.MonthEnd(3)
pd.Period('2012', freq='A') - pd.Period('2002', freq='A')
# ### Periods vs. Timestamps
#
# Notice the difference between Timestamps and Periods in determining differences in times.
pd.datetime.today() - pd.Timestamp('1/1/1970')
pd.Period(pd.datetime.today(), 'D') - pd.Period('1/1/1970', 'D')
pd.Period(pd.datetime.today(), 'A') - pd.Period('1/1/1970', 'A')
t = pd.datetime.today() - pd.Timestamp('1/1/1970')
t.days
# ### PeriodIndex and period_range
#
# Regular sequences of Period objects can be collected in a PeriodIndex, which can be constructed using the period_range convenience function:
prng = pd.period_range('1/1/2011', '1/1/2012', freq='M')
prng
# The PeriodIndex constructor can also be used directly:
pd.PeriodIndex(['2011-1', '2011-2', '2011-3'], freq='M')
# Passing multiplied frequency outputs a sequence of Period which has multiplied span.
pd.PeriodIndex(start='2014-01', freq='3M', periods=4)
# Just like DatetimeIndex, a PeriodIndex can also be used to index pandas objects:
ps = pd.Series(np.random.randn(len(prng)), prng)
ps
# PeriodIndex supports addition and subtraction with the same rule as Period.
idx = pd.period_range('2014-07-01 09:00', periods=5, freq='H')
idx
idx + pd.tseries.offsets.Hour(2)
idx = pd.period_range('2014-07', periods=5, freq='M')
idx
idx + pd.tseries.offsets.MonthEnd(3)
# ### PeriodIndex partial string indexing
#
# You can pass in dates and strings to Series and DataFrame with PeriodIndex, in the same manner as DatetimeIndex. For details, refer to DatetimeIndex Partial String Indexing.
ps
ps['2011-01']
ps[pd.datetime(2011, 12, 25):]
ps['10/31/2011':'12/31/2011']
# Passing a string representing a lower frequency than PeriodIndex returns partial sliced data.
ps['2011']
dfp = pd.DataFrame(np.random.randn(600,1), columns=['A'], index=pd.period_range('2013-01-01 9:00', periods=600, freq='T'))
dfp
dfp['2013-01-01 10H']
# As with DatetimeIndex, the endpoints will be included in the result. The example below slices data starting from 10:00 to 11:59.
dfp['2013-01-01 10H':'2013-01-01 11H']
# ### Frequency conversion and resampling with PeriodIndex
#
# The frequency of Period and PeriodIndex can be converted via the asfreq method. Let’s start with the fiscal year 2011, ending in December:
p = pd.Period('2011', freq='A-DEC')
p
# We can convert it to a monthly frequency. Using the how parameter, we can specify whether to return the starting or ending month:
p.asfreq('M', how='start')
p.asfreq('M', how='end')
# ## Converting between representations
#
# Timestamped data can be converted to PeriodIndex-ed data using to_period and vice-versa using to_timestamp:
rng = pd.date_range('1/1/2012', periods=5, freq='M')
ts = pd.Series(np.random.randn(len(rng)), index=rng)
ts
ps = ts.to_period()
ps
ps.to_timestamp()
ps.to_timestamp('D', how='start')
ps.to_timestamp('D', how='end')
# ## Time zone handling
#
# Pandas provides rich support for working with timestamps in different time zones using pytz and dateutil libraries. dateutil support is new in 0.14.1 and currently only supported for fixed offset and tzfile zones. The default library is pytz. Support for dateutil is provided for compatibility with other applications e.g. if you use dateutil in other python packages.
#
# ### Working with time zones
#
# By default, pandas objects are time zone unaware:
rng = pd.date_range('3/6/2012 00:00', periods=15, freq='D')
rng.tz is None
# To supply the time zone, you can use the tz keyword to date_range and other functions. There are two options for time zone formats:
#
# * pytz - pytz brings the Olson tz database into Python. This library allows accurate and cross platform timezone calculations using Python 2.4 or higher. You can find a list of common (and less common) time zones using from pytz import common_timezones, all_timezones. For UTC, there is a special case tzutc.
# * dateutil - The dateutil module provides powerful extensions to the datetime module available in the Python standard library. It uses the OS timezones so there isn’t a fixed list available. For common zones, the names are the same as pytz. Dateutil time zone strings are distinguished from pytz time zones by starting with "dateutil/".
# pytz
rng_pytz = pd.date_range('3/6/2012 00:00', periods=10, freq='D', tz='Europe/London')
rng_pytz.tz
# dateutil
rng_dateutil = pd.date_range('3/6/2012 00:00', periods=10, freq='D', tz='dateutil/Europe/London')
rng_dateutil.tz
# dateutil - utc special case
rng_utc = pd.date_range('3/6/2012 00:00', periods=10, freq='D', tz=dateutil.tz.tzutc())
rng_utc.tz
# Note that the UTC timezone is a special case in dateutil and should be constructed explicitly as an instance of dateutil.tz.tzutc. You can also construct other timezones explicitly first, which gives you more control over which time zone is used:
# pytz
tz_pytz = pytz.timezone('Europe/London')
rng_pytz = pd.date_range('3/6/2012 00:00', periods=10, freq='D', tz=tz_pytz)
rng_pytz.tz == tz_pytz
# dateutil
tz_dateutil = dateutil.tz.gettz('Europe/London')
rng_dateutil = pd.date_range('3/6/2012 00:00', periods=10, freq='D', tz=tz_dateutil)
rng_dateutil.tz == tz_dateutil
# Timestamps, like Python’s datetime.datetime object can be either time zone naive or time zone aware. Naive time series and DatetimeIndex objects can be localized using tz_localize:
ts = pd.Series(np.random.randn(len(rng)), rng)
ts
ts_utc = ts.tz_localize('UTC')
ts_utc
# Again, you can explicitly construct the timezone object first. You can use the tz_convert method to convert pandas objects to convert tz-aware data to another time zone:
ts_utc.tz_convert('US/Eastern')
# Warnings:
#
# * Be wary of conversions between libraries. For some zones pytz and dateutil have different definitions of the zone. This is more of a problem for unusual timezones than for ‘standard’ zones like US/Eastern.
# * Be aware that a timezone definition across versions of timezone libraries may not be considered equal. This may cause problems when working with stored data that is localized using one version and operated on with a different version. See here for how to handle such a situation.
# * It is incorrect to pass a timezone directly into the datetime.datetime constructor (e.g., datetime.datetime(2011, 1, 1, tz=timezone('US/Eastern')). Instead, the datetime needs to be localized using the the localize method on the timezone.
# Under the hood, all timestamps are stored in UTC. Scalar values from a DatetimeIndex with a time zone will have their fields (day, hour, minute) localized to the time zone. However, timestamps with the same UTC value are still considered to be equal even if they are in different time zones:
rng_eastern = rng_utc.tz_convert('US/Eastern')
rng_berlin = rng_utc.tz_convert('Europe/Berlin')
rng_eastern[5]
rng_berlin[5]
rng_eastern[5] == rng_berlin[5]
# Like Series, DataFrame, and DatetimeIndex, Timestamps can be converted to other time zones using tz_convert:
rng_eastern[5]
rng_berlin[5]
rng_eastern[5].tz_convert('Europe/Berlin')
# Localization of Timestamps functions just like DatetimeIndex and Series:
rng[5]
rng[5].tz_localize('Asia/Shanghai')
# Operations between Series in different time zones will yield UTC Series, aligning the data on the UTC timestamps:
eastern = ts_utc.tz_convert('US/Eastern')
berlin = ts_utc.tz_convert('Europe/Berlin')
result = eastern + berlin
result
result.index
# ### Other time zones
# List all the time zones available from pytz:
for tz in pytz.all_timezones:
print(tz)
# For example, Berlin is GMT+1 (flip the sign):
rng.tz_localize('Europe/Berlin')
rng.tz_localize('Etc/GMT-1')
# We can check timezones by converting the current time 'now' to another time zone:
pd.Timestamp('now', tz='US/Eastern')
pd.Timestamp('now', tz='UTC')
pd.Timestamp('now', tz='Asia/Riyadh')
pd.Timestamp('now', tz='Etc/GMT-3')
# ### Example: Day of week corresponding to a future date
def future_day_of_week(date, years_offset):
# dictionary to rename day of week
day_dict = {0: 'Monday', 1: 'Tuesday', 2: 'Wednesday', 3: 'Thursday',
4: 'Friday', 5: 'Saturday', 6: 'Sunday'}
# covert date to timestamp
ts = pd.Timestamp(date)
# offset date by X years
ts_offset = ts + pd.tseries.offsets.DateOffset(years=years_offset)
# print with strftime to make date pretty
print('%s in %s years will be %s which is a %s.' % (
ts.strftime('%Y-%m-%d'),
years_offset,
ts_offset.strftime('%Y-%m-%d'),
day_dict[ts_offset.dayofweek]))
future_day_of_week('1/1/00', 21)
# days from today until a given day
pd.Period('1/1/21', 'D') - pd.Period(pd.datetime.today(), 'D')
| lessons/lesson15.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: fastai
# language: python
# name: fastai
# ---
# +
# !pip install -Uqq fastbook
from itertools import islice
import gzip
import fastbook
fastbook.setup_book()
# -
from fastai.text.core import WordTokenizer, Tokenizer
from fastai.text.learner import TextLearner
from fastbook import *
def get_dls(limit, bs, sl):
path = download_data("https://github.com/tianhuil/number-generator/blob/main/data/en.txt.gz?raw=true")
with gzip.open(path, 'rt') as fh:
lines = L([line.strip() for line in islice(fh, 1, limit+1)])
text = ' . '.join([l.strip() for l in lines])
text = text.replace(', ', ' , ').replace('-', ' - ')
tokens = text.split(' ')
vocab = L(tokens).unique()
word2idx = {w:i for i,w in enumerate(vocab)}
nums = L(word2idx[i] for i in tokens)
def group_chunks(ds, bs):
m = len(ds) // bs
new_ds = L()
for i in range(m): new_ds += L(ds[i + m*j] for j in range(bs))
return new_ds
seqs = L((tensor(nums[i:i+sl]), tensor(nums[i+1:i+sl+1]))
for i in range(0,len(nums)-sl-1,sl))
cut = int(len(seqs) * 0.8)
dls = DataLoaders.from_dsets(group_chunks(seqs[:cut], bs),
group_chunks(seqs[cut:], bs),
bs=bs, drop_last=True, shuffle=False)
return dls, len(vocab)
class LMModel7(Module):
def __init__(self, vocab_sz, n_hidden, n_layers, p):
self.i_h = nn.Embedding(vocab_sz, n_hidden)
self.rnn = nn.LSTM(n_hidden, n_hidden, n_layers, batch_first=True)
self.drop = nn.Dropout(p)
self.h_o = nn.Linear(n_hidden, vocab_sz)
self.h_o.weight = self.i_h.weight
self.h = [torch.zeros(n_layers, bs, n_hidden) for _ in range(2)]
def forward(self, x):
raw,h = self.rnn(self.i_h(x), self.h)
out = self.drop(raw)
self.h = [h_.detach() for h_ in h]
return self.h_o(out),raw,out
def reset(self):
for h in self.h: h.zero_()
# +
limit = 10000
sl = 16
df_results = []
for bs in (16, 32, 64, 128):
dls, vocab_sz = get_dls(limit, bs, sl)
model = LMModel7(vocab_sz, bs, 2, 0.4)
# learn = Learner(dls, model,
# loss_func=CrossEntropyLossFlat(), metrics=accuracy,
# cbs=[ModelResetter, RNNRegularizer(alpha=2, beta=1)])
learn = TextLearner(dls, model,
loss_func=CrossEntropyLossFlat(), metrics=accuracy)
df = learn.fit_one_cycle(15, 1e-2, wd=0.1)
df_results.append(pd.DataFrame(
[[k] + vals for [k, vals] in enumerate(learn.recorder.values)],
columns=learn.recorder.metric_names[:-1]
))
df_results[-1]['bs'] = bs
df_result = pd.concat(df_results)
# -
df_result.pivot(index='epoch', columns='bs', values='accuracy').plot();
| analysis/Learn_Numbers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# what do i need to do?
#
# I need to:
# * make a basic gan out of biodeg
# * make a basic gan out of druglike
# * figure out the predictors that are being used for biodegradability
# * try out those descriptors for predictive power on the druglike set
# * (also, do I want to make a bigger druglike set?)
# +
import numpy as np
import pandas as pd
import tensorflow as tf
import matplotlib.pyplot as plt
import seaborn as sns
from math import floor, ceil
from pylab import rcParams
from sklearn.utils import shuffle
from sklearn.feature_selection import VarianceThreshold
# %matplotlib inline
# -
def encode(series):
return pd.get_dummies(series.astype(str))
train_x = pd.read_csv('../../big datasets/drugml/x_train_res.csv')
train_y = pd.read_csv('../../big datasets/drugml/y_train_res.csv')
test_x = pd.read_csv('../../big datasets/drugml/x_test.csv')
test_y = pd.read_csv('../../big datasets/drugml/y_test.csv')
from sklearn.preprocessing import StandardScaler
xscaler = StandardScaler().fit(train_x)
train_x = xscaler.transform(train_x)
testscaler = StandardScaler().fit(test_x)
test_x = testscaler.transform(test_x)
x_train = pd.DataFrame(train_x)
train_y = pd.DataFrame(train_y)
x_test = pd.DataFrame(test_x)
test_y = pd.DataFrame(test_y)
# +
#x_train = x_train.drop(['Unnamed: 0'], axis=1)
train_y = train_y.drop(['Unnamed: 0'], axis=1)
y_train = encode(train_y)
#y_train = encode(train_y)
#x_test = test_x.drop(['Unnamed: 0'], axis=1)
test_y = test_y.drop(['Unnamed: 0'], axis=1)
y_test = encode(test_y)
#y_test = encode(test_y)
# -
x_train.shape
x_train.head()
random_state = 42
np.random.seed(random_state)
tf.set_random_seed(random_state)
# +
def multilayer_perceptron(x, weights, biases, keep_prob):
layer_1 = tf.add(tf.matmul(x, weights['h1']), biases['b1'])
layer_1 = tf.nn.relu(layer_1)
layer_1 = tf.nn.dropout(layer_1, keep_prob)
out_layer = tf.matmul(layer_1, weights['out']) + biases['out']
return out_layer
# -
param_dist = {'hidden' : [10, 20, 30, 40, 50],
'prob' : [0.2, 0.4, 0.6, 0.8],
'size': [32, 45, 60],
'rate' : [0.05, 0.01, 0.001, 0.001]}
y_test = pd.DataFrame(y_test)
y_test.head()
# +
#### use meeeeee !!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!
n_hidden_1 = 40
n_input = train_x.shape[1]
n_classes = y_train.shape[1]
weights = {
'h1': tf.Variable(tf.random_normal([n_input, n_hidden_1])),
'out': tf.Variable(tf.random_normal([n_hidden_1, n_classes]))
}
biases = {
'b1': tf.Variable(tf.random_normal([n_hidden_1])),
'out': tf.Variable(tf.random_normal([n_classes]))
}
keep_prob = tf.placeholder("float")
training_epochs = 3
display_step = 1
batch_size = 60
x = tf.placeholder("float", [None, n_input])
y = tf.placeholder("float", [None, n_classes])
predictions = multilayer_perceptron(x, weights, biases, keep_prob)
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits=predictions, labels=y))
optimizer = tf.train.AdamOptimizer(learning_rate=0.1).minimize(cost)
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
for epoch in range(training_epochs):
avg_cost = 0.0
total_batch = int(len(x_train) / batch_size)
x_batches = np.array_split(x_train, total_batch)
y_batches = np.array_split(y_train, total_batch)
for i in range(total_batch):
batch_x, batch_y = x_batches[i], y_batches[i]
_, c = sess.run([optimizer, cost],
feed_dict={
x: batch_x,
y: batch_y,
keep_prob: 0.35
})
avg_cost += c / total_batch
if epoch % display_step == 0:
print("Epoch:", '%04d' % (epoch+1), "cost=", \
"{:.9f}".format(avg_cost))
print("Optimization Finished!")
correct_prediction = tf.equal(tf.argmax(y, 1), tf.argmax(predictions, 1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, "float"))
print("Accuracy:", accuracy.eval({x: x_batches[3], y: y_batches[3], keep_prob: 1.0}))
confusion = tf.confusion_matrix(labels=tf.argmax(y, 1), predictions=tf.argmax(predictions, 1), num_classes=2)
print(confusion.eval({x: x_batches[3], y: y_batches[3], keep_prob: 1.0}))
correct_prediction = tf.equal(tf.argmax(y, 1), tf.argmax(predictions, 1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, "float"))
print("Accuracy:", accuracy.eval({x: x_test, y: y_test, keep_prob: 1.0}))
confusion = tf.confusion_matrix(labels=tf.argmax(y, 1), predictions=tf.argmax(predictions, 1), num_classes=2)
print(confusion.eval({x: x_test, y: y_test, keep_prob: 1.0}))
# -
what is the accuracy and confusion matrix for my testing data?
from sklearn.utils import shuffle
# +
#### use meeeeee
#### use meeeeee
n_hidden_1 = 60
n_input = train_x.shape[1]
n_classes = train_y.shape[1]
weights = {
'h1': tf.Variable(tf.random_normal([n_input, n_hidden_1])),
'out': tf.Variable(tf.random_normal([n_hidden_1, n_classes]))
}
biases = {
'b1': tf.Variable(tf.random_normal([n_hidden_1])),
'out': tf.Variable(tf.random_normal([n_classes]))
}
keep_prob = tf.placeholder("float")
training_epochs = 10
display_step = 1
batch_size = 60
x = tf.placeholder("float", [None, n_input])
y = tf.placeholder("float", [None, n_classes])
predictions = multilayer_perceptron(x, weights, biases, keep_prob)
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits=predictions, labels=y))
optimizer = tf.train.AdamOptimizer(learning_rate=0.1).minimize(cost)
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
for epoch in range(training_epochs):
avg_cost = 0.0
total_batch = int(len(x_train) / batch_size)
x_train, y_train = shuffle(x_train, y_train, random_state=12)
x_batches = np.array_split(x_train, total_batch)
y_batches = np.array_split(y_train, total_batch)
for i in range(total_batch):
batch_x, batch_y = x_batches[i], y_batches[i]
_, c = sess.run([optimizer, cost],
feed_dict={
x: batch_x,
y: batch_y,
keep_prob: 0.35
})
avg_cost += c / total_batch
if epoch % display_step == 0:
print("Epoch:", '%04d' % (epoch+1), "cost=", \
"{:.9f}".format(avg_cost))
x_test, y_test = shuffle(x_test, y_test, random_state=12)
print("Optimization Finished!")
correct_prediction = tf.equal(tf.argmax(y, 1), tf.argmax(predictions, 1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, "float"))
print("Accuracy:", accuracy.eval({x: x_test, y: y_test, keep_prob: 1.0}))
confusion = tf.confusion_matrix(labels=tf.argmax(y, 1), predictions=tf.argmax(predictions, 1), num_classes=2)
print(confusion.eval({x: x_test, y: y_test, keep_prob: 1.0}))
# -
sa
from imblearn.over_sampling import SMOTE
estimator = run_model(setup_model)
param_dist = {'hidden' : [10, 20, 30, 40, 50],
'prob' : [0.2, 0.4, 0.6, 0.8],
'size': [32, 45, 60],
'rate' : [0.05, 0.01, 0.001, 0.001]}
hid = [30, 35, 40, 45, 50, 55, 60, 65, 70]
pro = [0.3, 0.35, 0.4]
rat = [0.1]
hid = [30, 40, 50]
pro = [0.3, 0.5, 0.7]
rat = [0.1]
siz = [60]
acc = []
for hidden in hid:
for prob in pro:
for rate in rat:
mod = []
n_hidden_1 = hidden
n_input = train_x.shape[1]
n_classes = y_train.shape[1]
weights = {
'h1': tf.Variable(tf.random_normal([n_input, n_hidden_1])),
'out': tf.Variable(tf.random_normal([n_hidden_1, n_classes]))
}
biases = {
'b1': tf.Variable(tf.random_normal([n_hidden_1])),
'out': tf.Variable(tf.random_normal([n_classes]))
}
keep_prob = tf.placeholder("float", None)
training_epochs = 20
display_step = 2
batch_size = 60
x = tf.placeholder("float", [None, n_input])
y = tf.placeholder("float", [None, n_classes])
predictions = multilayer_perceptron(x, weights, biases, keep_prob)
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits=predictions, labels=y))
optimizer = tf.train.AdamOptimizer(learning_rate=rate).minimize(cost)
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
for epoch in range(training_epochs):
avg_cost = 0.0
total_batch = int(len(x_train) / batch_size)
x_train, y_train = shuffle(x_train, y_train, random_state=12)
x_batches = np.array_split(x_train, total_batch)
y_batches = np.array_split(y_train, total_batch)
for i in range(total_batch):
batch_x, batch_y = x_batches[i], y_batches[i]
_, c = sess.run([optimizer, cost],
feed_dict={
x: batch_x,
y: batch_y,
keep_prob: prob})
avg_cost += c / total_batch
if epoch % display_step == 0:
print("Epoch:", '%04d' % (epoch+1), "cost=", \
"{:.9f}".format(avg_cost))
correct_prediction = tf.equal(tf.argmax(y, 1), tf.argmax(predictions, 1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, "float"))
mod.append(accuracy.eval({x: x_test, y: y_test, keep_prob: 1.0}))
confusion = tf.confusion_matrix(labels=tf.argmax(y, 1), predictions=tf.argmax(predictions, 1), num_classes=2)
mod.append(confusion.eval({x: x_test, y: y_test, keep_prob: 1.0}))
mod.append(hidden)
mod.append(prob)
mod.append(rate)
acc.append(mod)
sorted(acc, key=lambda x: x[0])
[0.800813, array([[ 41, 1], [ 48, 156]]), 30, 20, 0.1],
[0.800813, array([[ 42, 0], [ 49, 155]]), 70, 40, 0.1],
[0.8130081, array([[ 42, 0], [ 46, 158]]), 70, 60, 0.1],
[0.81707317, array([[ 33, 9], [ 36, 168]]), 70, 40, 0.001],
[0.8292683, array([[ 41, 1], [ 41, 163]]), 50, 20, 0.1]]
[0.7195122, array([[ 42, 0], [ 69, 135]]), 30, 60, 0.1],
[0.7357724, array([[ 42, 0], [ 65, 139]]), 50, 45, 0.1],
[0.7479675, array([[ 42, 0], [ 62, 142]]), 70, 32, 0.1],
[0.7479675, array([[ 20, 22], [ 40, 164]]), 100, 32, 0.001],
[0.7723577, array([[ 42, 0], [ 56, 148]]), 50, 32, 0.1],
[0.7723577, array([[ 42, 0], [ 56, 148]]), 100, 32, 0.1],
[0.7804878, array([[ 42, 0], [ 54, 150]]), 30, 45, 0.1],
[0.7804878, array([[ 17, 25], [ 29, 175]]), 50, 60, 0.001],
[0.7886179, array([[ 42, 0], [ 52, 152]]), 50, 60, 0.1],
[0.7886179, array([[ 42, 0], [ 52, 152]]), 100, 45, 0.1],
[0.7886179, array([[ 29, 13], [ 39, 165]]), 100, 45, 0.001],
[0.7886179, array([[ 42, 0], [ 52, 152]]), 100, 60, 0.1],
[0.796748, array([[ 41, 1], [ 49, 155]]), 30, 32, 0.1],
[0.796748, array([[ 18, 24], [ 26, 178]]), 70, 45, 0.001],
[0.8130081, array([[ 41, 1], [ 45, 159]]), 70, 45, 0.1],
[0.8211382, array([[ 42, 0], [ 44, 160]]), 70, 60, 0.1], # explore same batch sizes, explore hidden less than 100
[0.85365856, array([[ 32, 10], [ 26, 178]]), 70, 32, 0.001]] # explore 0.1 versus 0.01 versus 0.001
[0.7706422, array([[110, 36],
[ 14, 58]]), 10, 0.7, 60, 0.1],
[0.77522933, array([[117, 29],
[ 20, 52]]), 30, 0.5, 60, 0.1],
[0.77522933, array([[127, 19],
[ 30, 42]]), 70, 0.7, 60, 0.1],
[0.7844037, array([[131, 15],
[ 32, 40]]), 50, 0.3, 60, 0.1],
[0.7844037, array([[133, 13],
[ 34, 38]]), 50, 0.7, 60, 0.1], # best 0.1 learning rate, dropout 0.5 or 0.3, best hidden unknown...
[0.78899086, array([[129, 17],
[ 29, 43]]), 30, 0.7, 60, 0.1],
[0.79816514, array([[113, 33],
[ 11, 61]]), 10, 0.5, 60, 0.1],
[0.79816514, array([[138, 8],
[ 36, 36]]), 50, 0.5, 60, 0.1],
[0.8027523, array([[119, 27],
[ 16, 56]]), 30, 0.3, 60, 0.1],
[0.8027523, array([[137, 9],
[ 34, 38]]), 70, 0.3, 60, 0.1],
[0.8348624, array([[128, 18],
[ 18, 54]]), 70, 0.5, 60, 0.1]
[0.83027524, array([[132, 14],
[ 23, 49]]), 50, 0.3, 60, 0.1]] # narrow in on 50 and 0.3
[0.8211009, array([[126, 20],
[ 19, 53]]), 60, 0.35, 60, 0.1], winner
hyperparameters: 60, .35, 60, 0.1
from random import randint
from sklearn.model_selection import RandomizedSearchCV
# +
# Load the dataset
X, Y = x_train, y_train
# Create model for KerasClassifier
def create_model(hparams1,
hparams2,
hparams3,
hparams4):
n_hidden_1 = hparams1
n_input = train_x.shape[1]
n_classes = train_y.shape[1]
weights = {
'h1': tf.Variable(tf.random_normal([n_input, n_hidden_1])),
'out': tf.Variable(tf.random_normal([n_hidden_1, n_classes]))
}
biases = {
'b1': tf.Variable(tf.random_normal([n_hidden_1])),
'out': tf.Variable(tf.random_normal([n_classes]))
}
keep_prob = tf.placeholder("float")
training_epochs = 200
display_step = 90
batch_size = hparams3
x = tf.placeholder("float", [None, n_input])
y = tf.placeholder("float", [None, n_classes])
predictions = multilayer_perceptron(x, weights, biases, keep_prob)
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits=predictions, labels=y))
optimizer = tf.train.AdamOptimizer(learning_rate=hparams4).minimize(cost)
return
with tf.Session() as sess:
model = sess.run(tf.global_variables_initializer())
for epoch in range(training_epochs):
avg_cost = 0.0
total_batch = int(len(x_train) / batch_size)
x_batches = np.array_split(x_train, total_batch)
y_batches = np.array_split(y_train, total_batch)
for i in range(total_batch):
batch_x, batch_y = x_batches[i], y_batches[i]
_, c = sess.run([optimizer, cost],
feed_dict={
x: batch_x,
y: batch_y,
keep_prob: hparams2
})
avg_cost += c / total_batch
if epoch % display_step == 0:
print("Epoch:", '%04d' % (epoch+1), "cost=", \
"{:.9f}".format(avg_cost))
print("Optimization Finished!")
correct_prediction = tf.equal(tf.argmax(y, 1), tf.argmax(predictions, 1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, "float"))
print("Accuracy:", accuracy.eval({x: x_test, y: y_test, keep_prob: 1.0}))
confusion = tf.confusion_matrix(labels=tf.argmax(y, 1), predictions=tf.argmax(predictions, 1), num_classes=2)
print(confusion.eval({x: x_test, y: y_test, keep_prob: 1.0}))
# Specify parameters and distributions to sample from
hparams1 = randint(10, 100)
hparams2 = randint(0,10)*0.1
hparams3 = randint(30,60)
hparams4 = [0.1, 0.05, 0.01, 0.001]
# Prepare the Dict for the Search
param_dist = dict(hparams1=hparams1,
hparams2=hparams2,
hparams3=hparams3,
hparams4=hparams4)
# Search in action!
n_iter_search = 16 # Number of parameter settings that are sampled.
random_search = RandomizedSearchCV(estimator=model,
param_distributions=param_dist,
n_iter=n_iter_search)
random_search.fit(X, Y)
# Show the results
print("Best: %f using %s" % (random_search.best_score_, random_search.best_params_))
means = random_search.cv_results_['mean_test_score']
stds = random_search.cv_results_['std_test_score']
params = random_search.cv_results_['params']
for mean, stdev, param in zip(means, stds, params):
print("%f (%f) with: %r" % (mean, stdev, param))
# +
# Load the dataset
# Create model for KerasClassifier
n_hidden_1 = hparams1
n_input = train_x.shape[1]
n_classes = train_y.shape[1]
weights = {
'h1': tf.Variable(tf.random_normal([n_input, n_hidden_1])),
'out': tf.Variable(tf.random_normal([n_hidden_1, n_classes]))
}
biases = {
'b1': tf.Variable(tf.random_normal([n_hidden_1])),
'out': tf.Variable(tf.random_normal([n_classes]))
}
keep_prob = tf.placeholder("float")
training_epochs = 200
display_step = 90
batch_size = hparams3
x = tf.placeholder("float", [None, n_input])
y = tf.placeholder("float", [None, n_classes])
predictions = multilayer_perceptron(x, weights, biases, keep_prob)
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits=predictions, labels=y))
optimizer = tf.train.AdamOptimizer(learning_rate=hparams4).minimize(cost)
with tf.Session() as sess:
model = sess.run(tf.global_variables_initializer())
for epoch in range(training_epochs):
avg_cost = 0.0
total_batch = int(len(x_train) / batch_size)
x_batches = np.array_split(x_train, total_batch)
y_batches = np.array_split(y_train, total_batch)
for i in range(total_batch):
batch_x, batch_y = x_batches[i], y_batches[i]
_, c = sess.run([optimizer, cost],
feed_dict={
x: batch_x,
y: batch_y,
keep_prob: hparams2
})
avg_cost += c / total_batch
if epoch % display_step == 0:
print("Epoch:", '%04d' % (epoch+1), "cost=", \
"{:.9f}".format(avg_cost))
print("Optimization Finished!")
correct_prediction = tf.equal(tf.argmax(y, 1), tf.argmax(predictions, 1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, "float"))
print("Accuracy:", accuracy.eval({x: x_test, y: y_test, keep_prob: 1.0}))
confusion = tf.confusion_matrix(labels=tf.argmax(y, 1), predictions=tf.argmax(predictions, 1), num_classes=2)
print(confusion.eval({x: x_test, y: y_test, keep_prob: 1.0}))
# Specify parameters and distributions to sample from
hparams1 = randint(10, 100)
hparams2 = randint(0,10)*0.1
hparams3 = randint(30,60)
hparams4 = [0.1, 0.05, 0.01, 0.001]
# Prepare the Dict for the Search
param_dist = dict(hparams1=hparams1,
hparams2=hparams2,
hparams3=hparams3,
hparams4=hparams4)
# Search in action!
n_iter_search = 16 # Number of parameter settings that are sampled.
random_search = RandomizedSearchCV(estimator=model,
param_distributions=param_dist,
n_iter=n_iter_search)
random_search.fit(X, Y)
# Show the results
print("Best: %f using %s" % (random_search.best_score_, random_search.best_params_))
means = random_search.cv_results_['mean_test_score']
stds = random_search.cv_results_['std_test_score']
params = random_search.cv_results_['params']
for mean, stdev, param in zip(means, stds, params):
print("%f (%f) with: %r" % (mean, stdev, param))
# -
def model(X_train, Y_train, X_val, Y_val):
model = Sequential()
model.add(Dense({{choice([10, 20, 40, 104])}}))
model.add(Activation({{choice(['relu', 'sigmoid'])}}))
model.add(Dropout({{uniform(0, 1)}}))
model.add(Dense({{choice([10, 20, 40, 104])}}))
model.add(Activation({{choice(['relu', 'sigmoid'])}}))
model.add(Dropout({{uniform(0, 1)}}))
if conditional({{choice(['two', 'three'])}}) == 'three':
model.add(Dense({{choice([10, 20, 40, 104])}}))
model.add(Activation({{choice(['relu', 'sigmoid'])}}))
model.add(Dropout({{uniform(0, 1)}}))
model.add(Dense(10))
model.add(Activation('softmax'))
adam = keras.optimizers.Adam(lr={{choice([10**-3, 10**-2, 10**-1])}})
rmsprop = keras.optimizers.RMSprop(lr={{choice([10**-3, 10**-2, 10**-1])}})
sgd = keras.optimizers.SGD(lr={{choice([10**-3, 10**-2, 10**-1])}})
choiceval = {{choice(['adam', 'sgd', 'rmsprop'])}}
if choiceval == 'adam':
optim = adam
elif choiceval == 'rmsprop':
optim = rmsprop
else:
optim = sgd
model.compile(loss='categorical_crossentropy', metrics=['accuracy'],optimizer=optim)
model.fit(X_train, Y_train,
batch_size={{choice([128,256,512])}},
nb_epoch=20,
verbose=2,
validation_data=(X_val, Y_val))
score, acc = model.evaluate(X_val, Y_val, verbose=0)
print('Test accuracy:', acc)
return {'loss': -acc, 'status': STATUS_OK, 'model': model}
X_train, Y_train, X_val, Y_val = x_train, y_train, x_test, y_test
from hyperopt import Trials, STATUS_OK, tpe
from hyperas import optim
from hyperas.distributions import choice, uniform
import keras
data = X_train, Y_train, X_val, Y_val
best_run, best_model = optim.minimize(model=model,
data=data,
algo=tpe.suggest,
max_evals=30,
trials=Trials(),
notebook_name='Neural Network-Copy1')
# need to make this different... this needs to have a different train/test layout because the test data is not getting called properly. try https://towardsdatascience.com/how-to-use-dataset-in-tensorflow-c758ef9e4428 this method here
# _________________________________
# correct_prediction = tf.equal(tf.argmax(y, 1), tf.argmax(predictions, 1))
# accuracy = tf.reduce_mean(tf.cast(correct_prediction, "float"))
# print("Accuracy:", accuracy.eval({x: x_test, y: y_test, keep_prob: 1.0}))
#
#
# confusion = tf.confusion_matrix(labels=tf.argmax(y, 1), predictions=tf.argmax(predictions, 1), num_classes=2)
# print(confusion.eval({x: x_test, y: y_test, keep_prob: 1.0}))
# total_error = tf.reduce_sum(tf.square(tf.subtract(y, tf.reduce_mean(y))))
# unexplained_error = tf.reduce_sum(tf.square(tf.subtract(y, predictions)))
# R_squared = tf.subtract(1.0, tf.divide(unexplained_error, total_error))
# print(R_squared.eval({x: x_test, y: y_test, keep_prob: 1.0}))
# sklearn naive random oversampling (imbalanced data)
# https://imbalanced-learn.readthedocs.io/en/stable/over_sampling.html
# https://imbalanced-learn.readthedocs.io/en/stable/generated/imblearn.tensorflow.balanced_batch_generator.html
# * look for network architecture from paper that used the QSAR
# * lasso
# * fix for sparse data
# * find any columns that are uniform (or very low variation)
# * normalize
# * tensorboard
# * early stopping - ask rainie if i need help
# * put layer in after dropout
# * if oversampling, up the dropout (is there a ratio)
# * test set needs to be balanced but not oversampled
# * use their train/test split, then shuffle the data
# * combine the set up cells and do a for loop for the number of nodes (like 5-50 at 5 or 10 node increments)
# * use on waffle?
#
#
# create environment (homoganize)
# install everything i need into it
# * do the wget thing
# MSE = tf.metrics.mean_squared_error(tf.cast(y_test, tf.float32),
# predictions,
# weights=None,
# metrics_collections=None,
# updates_collections=None,
# name=None)
# print("MSE:", MSE)
# fn = tf.metrics.false_negatives(
# tf.cast(y_test, tf.float32),
# predictions,
# weights=None,
# metrics_collections=None,
# updates_collections=None,
# name=None)
# tn = tf.metrics.true_negatives(
# tf.cast(y_test, tf.float32),
# predictions,
# weights=None,
# metrics_collections=None,
# updates_collections=None,
# name=None)
# fp = tf.metrics.false_positives(
# tf.cast(y_test, tf.float32),
# predictions,
# weights=None,
# metrics_collections=None,
# updates_collections=None,
# name=None)
# tp = tf.metrics.true_positives(
# tf.cast(y_test, tf.float32),
# predictions,
# weights=None,
# metrics_collections=None,
# updates_collections=None,
# name=None)
# print("FN:", fn, "TN:", tn, "FP:", fp, "TP:", tp)
# total_error = tf.reduce_sum(tf.square(tf.subtract(tf.cast(y_test, tf.float32), tf.reduce_mean(tf.cast(y_test, tf.float32)))))
# unexplained_error = tf.reduce_sum(tf.square(tf.subtract(tf.cast(y_test, tf.float32), tf.cast(predictions, tf.float32))))
# R_squared = tf.subtract(1.0, tf.divide(unexplained_error, total_error))
# print(R_squared)
# df = pd.DataFrame({'num_legs': [2, 4, 8, 0],
# ... 'num_wings': [2, 0, 0, 0],
# ... 'num_specimen_seen': [10, 2, 1, 8]},
# ... index=['falcon', 'dog', 'spider', 'fish'])
# df
# df1 = df.iloc[:,1:3]
# df1.head()
| notebooks/bdrug_workingNeural Network-Copy1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="MBLabwK1LkZd" cellView="form"
#@title <center><img src='https://i.imgur.com/Vz0uo7C.png' width='275'>
OMDB_API = "" #@param {type: 'string'}
TMDB_API = "" #@param {type: 'string'}
REMOTE_NAME = "" #@param {type: 'string'}
SHARED_DRIVE = "" #@param {type: 'string'}
RCLONE_PASSWORD = "" #@param {type: 'string'}
HIDE_POST_COUNT = 25 #@param {type: 'integer'}
HIDE_REACT_SCORE = 25 #@param {type: 'integer'}
if OMDB_API == '':
OMDB_API = 'c42fc7df'
if TMDB_API == '':
TMDB_API = '6ed050a1118db3bc2c12a4ed13236430'
from google.colab import files
from IPython.display import clear_output
## Install FFmpeg, MediaInfo, rClone
# ##!wget -q 'https://github.com/InspectorWilliamHenderson/rFFmedia/releases/latest/download/rFFmedia.zip' && unzip -o -qq './rFFmedia.zip' -d '/' && rm -rf '/content/sample_data' './rFFmedia.zip
# ##!apt update
# !apt-get install mediainfo
# !curl https://rclone.org/install.sh | sudo bash
# !apt install ffmpeg
## Configure rClone
# !echo 'Upload the rClone.conf'
uploaded = files.upload()
clear_output()
# !rclone --config '' move './rclone.conf' '/root/.config/rclone/'
## rClone Mount
if SHARED_DRIVE != '':
SHARED_DRIVE = f'--drive-root-folder-id={SHARED_DRIVE}'
if RCLONE_PASSWORD != '':
# %env RCLONE_CONFIG_PASS=$RCLONE_PASSWORD
clear_output()
# !mkdir '/content/{REMOTE_NAME}/'
# !rclone mount '{REMOTE_NAME}': '/content/{REMOTE_NAME}/' --buffer-size 32M --daemon $SHARED_DRIVE --max-read-ahead 128M --poll-interval 30s --read-only --use-mmap --vfs-cache-mode full --vfs-read-chunk-size-limit 32M --vfs-read-chunk-size 16M -v
# + id="EHDvTWpgLbVr" cellView="form"
#@title <h3><b><center>Movie and TV Shows Template
import base64, os, re, requests, subprocess, threading
def GetMediaInfo(PATH):
global MEDIAINFO
MEDIAINFO = subprocess.check_output(['mediainfo', PATH]).decode().strip()
MEDIAINFO = MEDIAINFO.replace(PATH, os.path.basename(PATH))
def CreateScreenshots(PATH, TIME):
global SCREENSHOTS
LINK = subprocess.Popen(['ffmpeg', '-ss', TIME, '-i', PATH, '-frames:v', '1', '-y', f'./frame_{TIME}.jpeg'], stdin=open(os.devnull, 'wb'), stdout=open(os.devnull, 'wb')).communicate()
if os.path.exists(f'./frame_{TIME}.jpeg'):
LINK = requests.post('https://api.imgbb.com/1/upload', {'key': '<KEY>', 'image': base64.b64encode(open(f'./frame_{TIME}.jpeg', 'rb').read())})
if str(LINK.status_code) == '200':
SCREENSHOTS += f"[URL='{LINK.json()['data']['url']}'][IMG]{LINK.json()['data']['url']}[/IMG][/URL]\n"
def InitiateScreenshots():
global SCREENSHOTS
if ('.DoVi.' in os.path.basename(PATH)) or ('.DV.' in os.path.basename(PATH)):
SCREENSHOTS = "[HR][/HR][INDENT][SIZE=6][FORUMCOLOR][B]Media Screenshots :[/B][/FORUMCOLOR][/SIZE][/INDENT]\n[SPOILER='Media Screenshots'][JUSTIFY]Screenshots Can't be Made for This Release Because There is No Tool to Make Screenshots Out of a Muxed Dolby Vision File. There is a Very Limited Amount of Devices that can Play Dolby Vision Muxed this Way so Keep that in Mind Before Downloading.[/JUSTIFY][/SPOILER]"
elif SCREENSHOTS == True:
SCREENSHOTS = "[HR][/HR][INDENT][SIZE=6][FORUMCOLOR][B]Media Screenshots :[/B][/FORUMCOLOR][/SIZE][/INDENT]\n[SPOILER='Media Screenshots'][CENTER]"
SCREENSHOTS_01 = threading.Thread(target=CreateScreenshots(PATH, '02:00.00'))
SCREENSHOTS_02 = threading.Thread(target=CreateScreenshots(PATH, '04:00.00'))
SCREENSHOTS_03 = threading.Thread(target=CreateScreenshots(PATH, '06:00.00'))
SCREENSHOTS_01.start()
SCREENSHOTS_02.start()
SCREENSHOTS_03.start()
SCREENSHOTS_01.join()
SCREENSHOTS_02.join()
SCREENSHOTS_03.join()
SCREENSHOTS += '[/CENTER][/SPOILER]'
if SCREENSHOTS == "[HR][/HR][INDENT][SIZE=6][FORUMCOLOR][B]Media Screenshots :[/B][/FORUMCOLOR][/SIZE][/INDENT]\n[SPOILER='Media Screenshots'][CENTER][/CENTER][/SPOILER]":
SCREENSHOTS = ''
else:
SCREENSHOTS = ''
def GenerateLink(PATH, FOLDER):
global LINK
if LINK == "":
LINK = PATH.replace(f'/content/{REMOTE_NAME}/', '')
if FOLDER != 'Disabled':
LINK = os.path.dirname(LINK)
if FOLDER == 'GRANDPARENT FOLDER':
LINK = os.path.dirname(LINK)
if SHARED_DRIVE == '':
LINK = subprocess.check_output(['rclone', 'link', f'{REMOTE_NAME}:{LINK}']).decode().strip()
else:
LINK = subprocess.check_output(['rclone', 'link', f'{REMOTE_NAME}:{LINK}', SHARED_DRIVE]).decode().strip()
LINK = ''.join(LINK)
LINK = f"[HIDEREACT=1,2,3,4,5,6,7,8][DOWNCLOUD]{LINK}[/DOWNCLOUD][/HIDEREACT]"
if HIDE_REACT_SCORE != 0:
LINK = f'[HIDEREACTSCORE={HIDE_REACT_SCORE}]{LINK}[/HIDEREACTSCORE]'
if HIDE_POST_COUNT != 0:
LINK = f'[HIDEPOSTS={HIDE_POST_COUNT}]{LINK}[/HIDEPOSTS]'
def GetIMDb(PATH):
global IMDB
if IMDB == '':
try:
YEAR = '&y=' + re.findall(r'(19\d{2}|20\d{2})', PATH)[-1]
except:
YEAR = ''
TITLE = re.sub('[^a-zA-Z0-9]', '+', os.path.basename(PATH)).split('+' + YEAR.replace('&y=', '') + '+')[0].split('+S0')[0].split('+S1')[0].split('+S2')[0].split('+1080p')[0].split('+2160p')[0].split('+UHD')[0].split('+REPACK')[0].split('+HYBRID')[0].split('+EXTENDED')[0]
IMDB = requests.get(f'http://www.omdbapi.com/?t={TITLE}{YEAR}&apikey={OMDB_API}&r=json').json()
else:
IMDB = re.search(r'tt([0-9]+)', IMDB)[1].lstrip('0').zfill(7)
IMDB = requests.get(f'http://www.omdbapi.com/?i=tt{IMDB}&apikey={OMDB_API}&r=json').json()
def GetYouTube(QUERY):
global YOUTUBE
try:
if YOUTUBE == '':
YOUTUBE = 'https://www.youtube.com/watch?v=' + re.search(r'watch\?v=(\S{11})', requests.get(f'https://www.youtube.com/results?search_query={QUERY}+Official+Trailer').text)[1]
else:
YOUTUBE = 'https://www.youtube.com/watch?v=' + re.findall(r'([A-Za-z0-9-_]{11})', YOUTUBE)[-1]
except:
YOUTUBE = ''
def BlackPearl_Posting_Template():
global FOLDER, IMDB, LINK, PATH, SCREENSHOTS, YOUTUBE
# !rm -f ./frame*.jpeg
GetMediaInfo_THREAD = threading.Thread(target=GetMediaInfo(PATH))
InitiateScreenshots_THREAD = threading.Thread(target=InitiateScreenshots())
GenerateLink_THREAD = threading.Thread(target=GenerateLink(PATH, FOLDER))
GetIMDb_THREAD = threading.Thread(target=GetIMDb(PATH))
GetMediaInfo_THREAD.start()
InitiateScreenshots_THREAD.start()
GenerateLink_THREAD.start()
GetIMDb_THREAD.start()
GetIMDb_THREAD.join()
if IMDB['Response'] == 'False':
print('Give IMDb ID')
return None
GetYouTube_THREAD = threading.Thread(target=GetYouTube((IMDB['Title'] + " " + IMDB['Year'][0:4]).replace(' ', '+')))
GetYouTube_THREAD.start()
BBCODE = ''
try:
IMDB['Poster'] = "https://image.tmdb.org/t/p/original" + requests.get(f"http://api.themoviedb.org/3/movie/{IMDB['imdbID']}/images?api_key={TMDB_API}&language=en").json()['posters'][0]['file_path']
BBCODE = f"[CENTER][URL='{IMDB['Poster']}'][IMG WIDTH='350px']{IMDB['Poster']}[/IMG][/URL][/CENTER]\n"
except:
if ('Poster' in IMDB and IMDB['Poster'] != 'N/A'):
IMDB['Poster'] = re.sub('_V1_SX\d+.jpg', '_V1_SX1000.png', IMDB['Poster'])
BBCODE = f"[CENTER][URL='{IMDB['Poster']}'][IMG WIDTH='350px']{IMDB['Poster']}[/IMG][/URL][/CENTER]\n"
else:
BBCODE = '[CENTER][SIZE=160px][B]\n\n\n\n\nPOSTER\nMISSING\n\n\n\n\n\n[/B][/SIZE][/CENTER]\n'
BBCODE += f"[CENTER][URL='https://blackpearl.biz/search/1/?q={IMDB['imdbID']}&o=date'][FORUMCOLOR][B][SIZE=6]{IMDB['Title']} ({IMDB['Year'][0:4]})[/SIZE][/B][/FORUMCOLOR][/URL][/CENTER]\n"
BBCODE += f"[CENTER][URL='https://imdb.com/title/{IMDB['imdbID']}'][IMG WIDTH='46px']https://ia.media-imdb.com/images/M/MV5BMTk3ODA4Mjc0NF5BMl5BcG5nXkFtZTgwNDc1MzQ2OTE@.png[/IMG][/URL][/CENTER]"
BBCODE += f"[HR][/HR][INDENT][SIZE=6][FORUMCOLOR][B]Plot Summary :[/B][/FORUMCOLOR][/SIZE]\n\n[JUSTIFY]{IMDB['Plot']}[/JUSTIFY][/INDENT]" if ('Plot' in IMDB and IMDB['Plot'] != 'N/A') else ''
if ('Type' in IMDB and IMDB['Type'] == 'movie'):
IMDB['Type'] = 'Movie'
elif ('Type' in IMDB and IMDB['Type'] == 'series'):
IMDB['Type'] = 'TV Show'
else:
IMDB['Type'] = 'IMDb'
BBCODE += f"[HR][/HR][INDENT][SIZE=6][FORUMCOLOR][B]{IMDB['Type']} Info :[/B][/FORUMCOLOR][/SIZE][/INDENT]\n[LIST]"
BBCODE += f"[*][FORUMCOLOR][B]IMDb :[/B][/FORUMCOLOR] {IMDB['imdbRating']} ({IMDB['imdbVotes']})\n" if ('imdbRating' in IMDB and IMDB['imdbRating'] != 'N/A' and 'imdbVotes' in IMDB and IMDB['imdbVotes'] != 'N/A') else ''
BBCODE += f"[*][FORUMCOLOR][B]Rated :[/B][/FORUMCOLOR] {IMDB['Rated']}\n" if ('Rated' in IMDB and IMDB['Rated'] != 'N/A') else ''
BBCODE += f"[*][FORUMCOLOR][B]Genres :[/B][/FORUMCOLOR] {IMDB['Genre']}\n" if ('Genre' in IMDB and IMDB['Genre'] != 'N/A') else ''
BBCODE += f"[*][FORUMCOLOR][B]Awards :[/B][/FORUMCOLOR] {IMDB['Awards']}\n" if ('Awards' in IMDB and IMDB['Awards'] != 'N/A') else ''
BBCODE += f"[*][FORUMCOLOR][B]Runtime :[/B][/FORUMCOLOR] {IMDB['Runtime']}\n" if ('Runtime' in IMDB and IMDB['Runtime'] != 'N/A') else ''
BBCODE += f"[*][FORUMCOLOR][B]Starring :[/B][/FORUMCOLOR] {IMDB['Actors']}\n" if ('Actors' in IMDB and IMDB['Actors'] != 'N/A') else ''
BBCODE += f"[*][FORUMCOLOR][B]Countries :[/B][/FORUMCOLOR] {IMDB['Country']}\n" if ('Country' in IMDB and IMDB['Country'] != 'N/A') else ''
BBCODE += f"[*][FORUMCOLOR][B]Languages :[/B][/FORUMCOLOR] {IMDB['Language']}\n" if ('Language' in IMDB and IMDB['Language'] != 'N/A') else ''
BBCODE += f"[*][FORUMCOLOR][B]Written By :[/B][/FORUMCOLOR] {IMDB['Writer']}\n" if ('Writer' in IMDB and IMDB['Writer'] != 'N/A') else ''
BBCODE += f"[*][FORUMCOLOR][B]Directed By :[/B][/FORUMCOLOR] {IMDB['Director']}\n" if ('Director' in IMDB and IMDB['Director'] != 'N/A') else ''
BBCODE += f"[*][FORUMCOLOR][B]Release Date :[/B][/FORUMCOLOR] {IMDB['Released']}\n" if ('Released' in IMDB and IMDB['Released'] != 'N/A') else ''
BBCODE += f"[*][FORUMCOLOR][B]Production By :[/B][/FORUMCOLOR] {IMDB['Production']}\n" if ('Production' in IMDB and IMDB['Production'] != 'N/A') else ''
BBCODE += f"[*][FORUMCOLOR][B]DVD Release Date:[/B][/FORUMCOLOR] {IMDB['DVD']}\n" if ('DVD' in IMDB and IMDB['DVD'] != 'N/A') else ''
BBCODE += f"[*][FORUMCOLOR][B]Official Website :[/B][/FORUMCOLOR] {IMDB['Website']}\n" if ('Website' in IMDB and IMDB['Website'] != 'N/A') else ''
BBCODE += f"[*][FORUMCOLOR][B]Box Office Collection :[/B][/FORUMCOLOR] {IMDB['BoxOffice']}[/LIST]" if ('BoxOffice' in IMDB and IMDB['BoxOffice'] != 'N/A') else '[/LIST]'
BBCODE = BBCODE.replace('[HR][/HR][INDENT][SIZE=6][FORUMCOLOR][B]IMDb Info :[/B][/FORUMCOLOR][/SIZE][/INDENT]\n[LIST][/LIST]', '')
GetYouTube_THREAD.join()
BBCODE += f'[HR][/HR][INDENT][SIZE=6][FORUMCOLOR][B]Official Trailer :[/B][/FORUMCOLOR][/SIZE][/INDENT]\n\n[CENTER]{YOUTUBE}[/CENTER]' if YOUTUBE != '' else ''
BBCODE += '[HR][/HR][INDENT][SIZE=6][FORUMCOLOR][B]Media Info :[/B][/FORUMCOLOR][/SIZE][/INDENT]'
GetMediaInfo_THREAD.join()
InitiateScreenshots_THREAD.join()
BBCODE += f"[SPOILER='Media Info'][CODE TITLE='Media Info']{MEDIAINFO}[/CODE][/SPOILER]\n{SCREENSHOTS}"
GenerateLink_THREAD.join()
BBCODE += f'[HR][/HR][INDENT][SIZE=6][FORUMCOLOR][B]Download Link :[/B][/FORUMCOLOR][/SIZE][/INDENT]\n[CENTER]{LINK}[/CENTER]'
print(BBCODE)
# !rm -f ./frame*.jpeg
# + id="Qc221ZFqP9gx" cellView="form"
IMDB = "" #@param {type: 'string'}
YOUTUBE = "" #@param {type: 'string'}
PATH = "" #@param {type: 'string'}
LINK = "" #@param {type: 'string'}
FOLDER = "Disabled" #@param ['Disabled', 'PARENT FOLDER', 'GRANDPARENT FOLDER']
SCREENSHOTS = False #@param {type: 'boolean'}
try:
BlackPearl_Posting_Template()
except NameError:
print("Run Second Cell Before Running The Third Cell")
| BlackPearl Template Posters.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
from explore import *
pandas_vertices_from_plyfile('data/cube.ply')
pptk_plot_df(pandas_vertices_from_plyfile('data/cube.ply'))
plydata = load_example_cube()
pptk_plot_plydata(plydata)
vaex_vertices_from_plyfile('data/cube.ply')
ipv_plot_df(vaex_vertices_from_plyfile('data/cube.ply'))
_df = vaex_vertices_from_plyfile('data/cube.ply')
ipv_plot_df(_df)
_df['x']
ipv_plot_df(_df.to_pandas_df())
ipv.show()
# Ok, some weird issue using vaex dataframes in ipyvolume, I'll just go with pandas for the time being.
| req1.2_import_functions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Error Analysis and Visualisation for XGBoost Models
# ### Check running python kernel
# !which python
# ### Imports
import matplotlib.pyplot as plt
import pandas as pd
import glob
import os
from sklearn.metrics import r2_score
# ### Top Level Constants
drop_columns = ['lon', 'lat', 'APE']
HIGH_THRESHOLD = 1e10
result_dir = '../../data/midlats/results'
model_type = 'midlats'
# ### Inference Files
FILE_PATHS = os.path.join(result_dir, '*'+model_type+'*.csv')
FILE_NAMES = glob.glob(FILE_PATHS)
print(FILE_NAMES)
# ### Make Dataframe
fli_df = pd.concat((pd.read_csv(f, index_col=False).reset_index(drop=True) for f in FILE_NAMES), ignore_index=True)
fli_df
# ### Stats
actual_mean = fli_df['actual_load'].mean()
actual_median = fli_df['actual_load'].median()
print("Actual Fuel Load mean - {actual_mean} and median - {actual_median}".format(actual_mean = actual_mean, actual_median = actual_median))
Predicted_mean = fli_df['predicted_load'].mean()
Predicted_median = fli_df['predicted_load'].median()
print("Predicted Fuel Load mean - {Predicted_mean} and median - {Predicted_median}".format(Predicted_mean = Predicted_mean, Predicted_median = Predicted_median))
# ### $R^2$ Score
# More details [here](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.r2_score.html)
r2_score(fli_df['actual_load'], fli_df['predicted_load'])
# ### General Trend of Actual and Predicted Values
axes = fli_df.drop(columns=drop_columns).plot(subplots = True, figsize=(12, 12))
for ax in axes:
ax.set_ylim(0, 1e11)
# ### Distribution of high values (> HIGH_THRESHOLD)
fli_df_high = fli_df[fli_df['actual_load'] >= HIGH_THRESHOLD].reset_index(drop=True)
axes = fli_df_high.drop(columns=drop_columns).plot.hist(bins=40,
subplots=True,
figsize=(12, 12),
cumulative=True)
# ### Error Stats
fli_err = fli_df.drop(columns=['lat', 'lon', 'predicted_load', 'actual_load'])
fli_err.max()
fli_err.mean()
fli_err.median()
| notebooks/results/error_analysis_catboost_midlats.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: LSST
# language: python
# name: lsst
# ---
# # LSST Camera Geometry
#
# <br>Owner(s): **<NAME>** ([@kadrlica](https://github.com/LSSTScienceCollaborations/StackClub/issues/new?body=@kadrlica))
# <br>Last Verified to Run: **2021-09-11**
# <br>Verified Stack Release: **w_2021_33**
#
# This notebook demonstrates how to interact with the Stack representation of the LSST camera (`lsstCam`). We use it to get the geometry of the various nested components of the camere focal plane -- i.e., amps, detectors, and rafts. We then produce a labeled figure of the LSST focal plane geometry.
#
# ### Learning Objectives:
#
# After working through this tutorial you should be able to:
#
# 1. Access the geometry of specific LSST amps, detectors, and rafts.
# 2. Plot the geometry of these camera components.
# 3. Create a labeled plot of the LSST focal plane geometry.
#
# ### Logistics
# This notebook is intended to be run at `lsst-lsp-stable.ncsa.illinois.edu` or `data.lsst.cloud` from a local git clone of the [StackClub](https://github.com/LSSTScienceCollaborations/StackClub) repo.
# ## Setup
# You can find the Stack version by using `eups list -s` on the terminal command line.
# Site, host, and stack version
# ! echo $EXTERNAL_INSTANCE_URL
# ! echo $HOSTNAME
# ! eups list -s | grep lsst_distrib
# +
import os
from matplotlib.patches import Polygon
from matplotlib.collections import PatchCollection
import matplotlib.pyplot as plt
from lsst.daf.butler import Butler
from lsst.afw.cameraGeom import utils as cgUtils
from lsst.obs.lsst.cameraTransforms import LsstCameraTransforms
from lsst.obs.lsst.cameraTransforms import ampPixelToCcdPixel
import lsst.afw.cameraGeom as cameraGeom
import lsst.afw.geom as afwGeom
from lsst.afw.cameraGeom import FIELD_ANGLE, FOCAL_PLANE
# -
# ## Existing Tools
#
# The stack provides a pre-wrapped visualization of the camera geometry. However, this figure has a few limitations: the detector labels are very small, rafts are only indicated in the sensor names, and amplifier geometry is not shown. A more visually apealing version of the LSST camera geometry can be found [here](https://confluence.lsstcorp.org/display/LSWUG/Representation+of+a+Camera?preview=/4129064/10190878/LSST_FocalPlane.png#/); however, it has a different orientation than is used by the DM Stack.
# +
URL = os.getenv('EXTERNAL_INSTANCE_URL')
if URL.endswith('data.lsst.cloud'): # IDF
repo = "s3://butler-us-central1-dp01"
elif URL.endswith('ncsa.illinois.edu'): # NCSA
repo = "/repo/dc2"
else:
raise Exception(f"Unrecognized URL: {URL}")
dataset='DC2'
collection='2.2i/runs/DP0.1'
butler = Butler(repo,collections=collection)
camera = butler.get('camera',instrument='LSSTCam-imSim')
cgUtils.plotFocalPlane(camera)
# -
# The source code for `cgUtils.plotFocalPlane` found [here](https://github.com/lsst/afw/blob/01e196b6519ef91d51c61435065e16477972b897/python/lsst/afw/cameraGeom/utils.py#L87) gives us a very good place to start our investigation of the LSST camera geometry.
# ## Plot CCDs on Focal Plane
#
# To get started, it's helpful to understand how to reproduce the figure above. The basic idea is to build an instance of the LSST camera object, and then loop through the detectors that it contains. For each detector, we plot the detector location and label the detector. We make a slight change to the code above by labeling with the CCD ID (a unique integer for each detector) rather than the `name` of the detector (which is built from the raft/sensor information)
# +
fig= plt.figure(figsize=(10,10))
ax = plt.gca()
xvals,yvals = [],[]
colors,patches = [],[]
for det in camera:
corners = [(c.getX(), c.getY()) for c in det.getCorners(FOCAL_PLANE)]
for corner in corners:
xvals.append(corner[0])
yvals.append(corner[1])
colors.append('b')
patches.append(Polygon(corners, True))
center = det.getOrientation().getFpPosition()
text = det.getId()
rot = 90 if text in (195,196,199,200) else 0
ax.text(center.getX(), center.getY(), text,
ha='center', va='center', size=12, rotation=rot)
patchCollection = PatchCollection(patches, alpha=0.6, facecolor=colors)
ax.add_collection(patchCollection)
ax.set_xlim(min(xvals) - abs(0.1*min(xvals)),
max(xvals) + abs(0.1*max(xvals)))
ax.set_ylim(min(yvals) - abs(0.1*min(yvals)),
max(yvals) + abs(0.1*max(yvals)))
ax.set_xlabel('Focal Plane X (mm)')
ax.set_ylabel('Focal Plane Y (mm)')
# -
# ## Plot Amps on a CCD
#
# We'll start by grabbing the amps from a specific detector (the central detector R22,S11). We can then plot the extents of the amps in pixel coordinates.
# +
# Create the camera transformation
transform = LsstCameraTransforms(camera)
# Get the central detector
det = transform.getDetector('R22_S11')
# Get the amps for this detector
amps = det.getAmplifiers()
fig,ax = plt.subplots(1,figsize=(10,10))
patches,colors = [],[]
for amp in amps:
corners = [(c.getX(), c.getY()) for c in amp.getBBox().getCorners()]
patches.append(Polygon(corners, True))
colors.append('b')
center = amp.getBBox().getCenter()
text = amp.getName()
ax.text(center.getX(), center.getY(), text, color='k',
ha='center', va='center', size=14)
# Add the patch collection
patchCollection = PatchCollection(patches, alpha=0.6, facecolor=colors,edgecolor='k')
ax.add_collection(patchCollection)
# Set some labels and extent
ax.set_xlim(-200,4250)
ax.set_ylim(-200,4250)
ax.set_xlabel('CCD X (pix)')
ax.set_ylabel('CCD Y (pix)')
# -
# Ok, so this is all well and good, but what if we want to plot the physical positions of the amps in focal plane coordinates? We should be able to do this with a transformation.
# +
def plotFancyDetector(camera, detectorName='R22_S11', figsize=(10.,10.)):
# Create the camera transformation
transform = LsstCameraTransforms(camera)
# Get the central detector
det = transform.getDetector(detectorName)
# Get the amps for this detector
amps = det.getAmplifiers()
fig,ax = plt.subplots(1,figsize=figsize)
patches,colors = [],[]
xvals,yvals = [],[]
for amp in amps:
points = [transform.ccdPixelToFocalMm(c.getX(), c.getY(), det.getName()) for c in amp.getBBox().getCorners()]
corners = [(p.getX(),p.getY()) for p in points]
for corner in corners:
xvals.append(corner[0])
yvals.append(corner[1])
patches.append(Polygon(corners, True))
colors.append('skyblue')
# Center in pixels
center_pix = amp.getBBox().getCenter()
# center in mm
center = transform.ccdPixelToFocalMm(center_pix.getX(),center_pix.getY(),det.getName())
text = amp.getName()
ax.text(center.getX(), center.getY(), text, color='k',
ha='center', va='center', size=14)
# Add the patch collection
patchCollection = PatchCollection(patches, alpha=0.6, facecolor=colors,edgecolor='k')
ax.add_collection(patchCollection)
ax.set_xlim(min(xvals) - abs(0.02*min(xvals)),
max(xvals) + abs(0.02*max(xvals)))
ax.set_ylim(min(yvals) - abs(0.02*min(yvals)),
max(yvals) + abs(0.02*max(yvals)))
ax.set_xlabel('Focal Plane X (mm)')
ax.set_ylabel('Focal Plane Y (mm)')
ax.set_title(det.getName())
# Set some labels and extent
plotFancyDetector(camera,'R11_S11')
# -
# # Plot CCDs on a Raft
#
# It looks like the LSST Camera object doesn't have the concept of a raft (just a list of detectors). If we want to assemble a raft, we can do so directly from the CCDs.
# +
def plotFancyRaft(camera, raftName='R22', figsize=(10.,10.)):
colorMap = {0: 'skyblue', 1: 'y', 2: 'g', 3: 'r'}
transform = LsstCameraTransforms(camera)
plt.figure(figsize=figsize)
ax = plt.gca()
patches, colors = [],[]
xvals, yvals = [],[]
for det in camera:
if not det.getName().startswith(raftName): continue
corners = [(c.getX(), c.getY()) for c in det.getCorners(FOCAL_PLANE)]
for corner in corners:
xvals.append(corner[0])
yvals.append(corner[1])
colors.append(colorMap[int(det.getType())])
patches.append(Polygon(corners, True))
center = det.getOrientation().getFpPosition()
name = det.getName()
# Label central raft
text = '(%s,%s)'%tuple(name.split('_')[1].strip('S'))
ax.text(center.getX(), center.getY(), text, color='0.3',
ha='center', va='center',size=18)
patchCollection = PatchCollection(patches, alpha=0.6, facecolor=colors)
ax.add_collection(patchCollection)
ax.set_xlim(min(xvals) - abs(0.1*min(xvals)),
max(xvals) + abs(0.1*max(xvals)))
ax.set_ylim(min(yvals) - abs(0.1*min(yvals)),
max(yvals) + abs(0.1*max(yvals)))
ax.set_xlabel('Focal Plane X (mm)')
ax.set_ylabel('Focal Plane Y (mm)')
ax.set_title(raftName)
plotFancyRaft(camera)
# -
# ## Plot Focal Plane
#
# Now we put together what we've learned to try to replicate the image [here](https://confluence.lsstcorp.org/display/LSWUG/Representation+of+a+Camera?preview=/4129064/10190878/LSST_FocalPlane.png#/). Note that the corner rafts (containing the focus and guiding CCDs) are not yet included in the `lsstCamera` model. Also, since the concept of a raft doesn't really exist, we don't plot outlines around the rafts (we could hack this if we wanted).
# +
def plotFancyFocalPlane(camera, figsize=(10., 10.), showFig=True, savePath=None):
"""Make a plot of the focal plane along with a set points that sample
the field of view.
Parameters
----------
camera : `lsst.afw.cameraGeom.Camera`
A camera object
figsize : `tuple` containing two `float`
Matplotlib style tuple indicating the size of the figure in inches
showFig : `bool`
Display the figure on the screen?
savePath : `str` or `None`
If not `None`, save a copy of the figure to this name.
"""
try:
from matplotlib.patches import Polygon
from matplotlib.collections import PatchCollection
import matplotlib.pyplot as plt
except ImportError:
raise ImportError(
"Can't run plotFocalPlane: matplotlib has not been set up")
colorMap = {0: 'skyblue', 1: 'y', 2: 'g', 3: 'r'}
transform = LsstCameraTransforms(camera)
plt.figure(figsize=figsize)
ax = plt.gca()
patches, colors = [],[]
xvals, yvals = [],[]
for det in camera:
corners = [(c.getX(), c.getY()) for c in det.getCorners(FOCAL_PLANE)]
for corner in corners:
xvals.append(corner[0])
yvals.append(corner[1])
colors.append(colorMap[int(det.getType())])
patches.append(Polygon(corners, True))
center = det.getOrientation().getFpPosition()
name = det.getName()
# Label central raft
if name.startswith('R22'):
if not name.endswith('S11'):
# Label CCDs for central raft
text = '(%s,%s)'%tuple(name.split('_')[-1].strip('S'))
ax.text(center.getX(), center.getY(), text,
ha='center', va='center', size=10)
else:
# Draw the amps for the central CCD
amp_patches = []
for amp in det.getAmplifiers():
points = [transform.ccdPixelToFocalMm(c.getX(), c.getY(), det.getName()) for c in amp.getBBox().getCorners()]
corners = [(p.getX(),p.getY()) for p in points]
amp_patches.append(Polygon(corners, True))
# Add the amp patch collection
patchCollection = PatchCollection(amp_patches, alpha=0.6, facecolor='none',edgecolor='k')
ax.add_collection(patchCollection)
elif name.endswith('S11'):
text = '(%s,%s)'%tuple(name.split('_')[0].strip('R'))
ax.text(center.getX(), center.getY(), text, color='0.3',
ha='center', va='center',size=22)
for raft in ('R00', 'R40', 'R44', 'R04'):
# These rafts don't have an S11 sensor, so need to figure out the raft center from the other rafts around them
_, y, x = list(raft)
column = camera[f'R1{x}_S11'].getOrientation().getFpPosition() # Just needs to be in the column, could have used R2{x}
row = camera[f'R{y}1_S11'].getOrientation().getFpPosition() # Same for rows
text = f'({y},{x})'
ax.text(column.getX(), row.getY(), text, color='0.3',
ha='center', va='center',size=22)
patchCollection = PatchCollection(patches, alpha=0.6, facecolor=colors)
ax.add_collection(patchCollection)
ax.set_xlim(min(xvals) - abs(0.1*min(xvals)),
max(xvals) + abs(0.1*max(xvals)))
ax.set_ylim(min(yvals) - abs(0.1*min(yvals)),
max(yvals) + abs(0.1*max(yvals)))
ax.set_xlabel('Focal Plane X (mm)',fontsize=18)
ax.set_ylabel('Focal Plane Y (mm)',fontsize=18)
if savePath is not None:
plt.savefig(savePath)
if showFig:
plt.show()
# Plot the focal plane
plotFancyFocalPlane(camera)
| Visualization/LsstCameraGeometry.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from sklearn import preprocessing
import math
import random
import matplotlib.pyplot as plt
optimum_val = []
def Dejon_fun(a,b):
obj_val = []
tmp = 0
for i in range(len(a)):
tmp = a[i]**2 + b[i]**2
obj_val.append(tmp)
print(obj_val)
return obj_val
def Rosenbrock_fun(a, b):
f = 0
obj_val = []
for i in range(len(a)):
f = 100*np.power(b[i]-np.power(a[i],2),2) + np.power(a[i]-1,2)
obj_val.append(f)
print(obj_val)
return obj_val
def Rastrigin_fun(a,b):
import math
obj_val = []
tem2 = []
for i in range(len(a)):
# x = 20 + (np.subtract(np.square(a), 10 * math.cos(2*np.pi* a[i]))) + (np.subtract(np.square(b), 10 * math.cos(2*np.pi* b[i])))
tmp2 = 20 + np.power(a[i],2)-10*np.cos(2*np.pi*a[i]) + np.power(b[i],2)-10*np.cos(2*np.pi*b[i])
# print(tmp2)
obj_val.append(tmp2)
# print(obj_val)
return (obj_val)
def Schwefel_function(a,b):
obj_val = []
tmp = 0
for i in range(len(a)):
# print(i)
tmp = -a[i]*np.sin(np.sqrt(np.absolute(a[i]))) - b[i]*np.sin(np.sqrt(np.absolute(b[i])))
# print(tmp)
obj_val.append(tmp)
print(obj_val)
return obj_val
def Ackley_fun(a,b):
tmp1 = 0
tmp2 = 0
obj_val = []
for i in range(len(a)):
if (a[i]* ((np.square(np.absolute(a[i])) + np.square(np.absolute(b[i])))) == 0):
tmp1 =0
else:
tmp1 = 20.-20.*np.exp(-0.2*np.sqrt(np.absolute(1./a[i]* (np.square(np.absolute(a[i])) + np.square(np.absolute(b[i]))))))
if (a[i]*((np.cos(a[i]*2.*np.pi)) + (np.cos(b[i]*2.*np.pi)))) == 0:
tmp2 = 0
else:
tmp2 = np.e-np.exp(1./(a[i]*((np.cos(a[i]*2.*np.pi)) + (np.cos(b[i]*2.*np.pi)))))
x = tmp1 + tmp2
obj_val.append(x)
print("obj_aukley", obj_val)
return obj_val
def Griewangk_fun(a,b):
tmp1 = 0
tmp2 = tmp3 = 0
obj_val = []
for i in range(len(a)):
tmp1 = np.power(a[i],2) + np.power(b[i],2)
tmp2 = np.cos(a[i]/np.sqrt(2))*np.cos(b[i]/np.sqrt(2))
tmp3 - tmp1/4000
x = tmp3 - tmp2 + 1
obj_val.append(x)
return obj_val
def Lilun_fun(a,b):
tmp = 0
sume = 0
obj_val = []
for i in range(len(a)):
sume = a[i] + b[i]
tmp = np.power(a[i], 4) + np.power(b[i], 4) + np.power(sume, 2)
obj_val.append(tmp)
print(obj_val)
return obj_val
| Particle Swarm Optimization/Objective_Functions-Copy1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Tutorial demonstrating the basic functionality of the `iwatlas` package
#
# In this tutorial we will learn how to:
#
# - Download the data netcdf4 file
# - Inspect the data
# - Plot a variable
# - Interpolate a scalar onto a point
# - Extract a time-series of internal tide sea surface height amplitude at a point
# - Extract the stratification at a point
#
# ---
#
#
# +
# These are the sub-modules in the iwatlas package that we will use
from iwatlas import sshdriver
from iwatlas import uvdriver
from iwatlas import harmonics
from iwatlas import stratification as strat
from iwatlas import iwaves
import xarray as xr
import pandas as pd
import numpy as np
from scipy.interpolate import interp1d
import matplotlib.pyplot as plt
# +
# Uncomment this option to allow for interactive plot windows (e.g. zooming)
# # %matplotlib notebook
# -
# Set where you want to download the 200 MB data file
# basedir = '/home/jupyter-ubuntu/data/iwatlas'
basedir = '../DATA'
# +
# %%time
# Download the data if it does not exist
import urllib, os
# Link to a 200 MB data file on cloudstor
# publicurl = 'https://cloudstor.aarnet.edu.au/plus/s/vdksw5WKFOTO0nD/download'
publicurl = 'https://research-repository.uwa.edu.au/files/93942498/NWS_2km_GLORYS_hex_2013_2014_InternalWave_Atlas.nc'
atlasfile = '{}/NWS_2km_GLORYS_hex_2013_2014_InternalWave_Atlas.nc'.format(basedir)
if os.path.exists(basedir):
print('Folder exists.')
else:
print('Making folder {}'.format(basedir))
os.mkdir(basedir)
if os.path.exists(atlasfile):
print('File exists.')
else:
print('Downloading file...')
urllib.request.urlretrieve (publicurl, atlasfile)
print('Done. Saved to {}'.format(atlasfile))
atlasfile
# -
# # Example 1: Open the dataset
#
# For this we will use the function `sshdriver.load_ssh_clim` method. This method wraps the `sfoda.ugrid.sunxray.Sunxray` class that is basically an unstructured grid `xarray.Dataset` object.
# +
atlasfile
ssh = sshdriver.load_ssh_clim(atlasfile)
ssh = sshdriver.load_ssh_clim(ssh)
ssh
# -
# We can see from the printout above the *coordinate* and *data variables* plus some *global attributes*. Many of the coordinate and data variables are associated with the unstructured grid topology (e.g., $xv$, $cells$, $nfaces$). The data variables of interest are stored as follows:
#
# - Sea surface height harmonics: `SSH_BC_*`
# - Stratification variables: `N2_*`
#
# # Example 2: Inspect an individual variable
#
# The `xarray.Dataset` class is the `._ds` attribute. To access a variable (an `xarray.DataArray` object) we call `ssh._ds[varname]`.
ssh._ds['SSH_BC_var']
# # Example 3: Plot a spatial variable
#
# The data is on an unstructured grid so we call either `plotcelldata` or `contourf` methods in the object.
# +
# To show help for a method within an object use the ? symbol e.g.,
# ssh.plotcelldata?
# -
# Plot the Sea surface height signal variance
plt.figure(figsize=(10,6))
ssh.plotcelldata(ssh._ds['SSH_BC_var'], vmax=1e-2, cmap='Reds')
plt.title(ssh._ds['SSH_BC_var'].attrs['long_name'])
# This is nice but it is hard to see any landmarks. Let's add some bathymetry contours...
plt.figure(figsize=(10,6))
ax=plt.subplot(111,facecolor='0.5')
ssh.plotcelldata(ssh._ds['SSH_BC_var'], vmax=1e-2, cmap='Reds')
ssh.contourf(ssh._ds['dv'], [100, 200, 500, 1000, 2000], colors='k', linewidths=0.2, filled=False, colorbar=False)
plt.title(ssh._ds['SSH_BC_var'].attrs['long_name'])
# # Example 4: Interpolate a scalar onto a point
#
# The `sunxray` object has a convenient `.interpolate` method to extract a scalar from the unstructured grid.
#
# WA-IMOS locations (August 2019)
sites = {
'NIN100':{'y':-21.84986667,'x':113.9064667},
'NWSBAR':{'y':-20.76128333,'x':114.7586167},
'NWSROW':{'y':-17.75801667,'x':119.9061},
'NWSBRW':{'y':-14.23543333,'x':123.1623833},
'NWSLYN':{'y':-9.939416667,'x':130.3490833},
'PIL200':{'x': 115.9154, 'y':-19.435333} ,
'KIM200':{'x':121.243217 , 'y':-15.534517} ,
'KIM400':{'x': 121.114967, 'y':-15.22125} ,
'ITFTIS':{'x': 127.5577, 'y':-9.819217} ,
'BB250':{'x':123.34613 , 'y':-13.75897} ,
'Prelude':{'x':123.3506, 'y':-13.7641} ,
}
# +
# Spatial coordinates of point
sitename = 'ITFTIS'
xpt = sites[sitename]['x']
ypt = sites[sitename]['y']
# Call to the interpolation method
mydata = ssh.interpolate(ssh._ds['SSH_BC_var'].values, xpt, ypt, kind='linear')
print('The SSH variance at X: {} Y: {} is {}'.format(\
xpt, ypt, mydata ) )
# -
# # Example 5: Extract a time-series of baroclinic sea surface height anomaly
#
# To do this we use the driver functions in the `iwatlas.sshdriver` submodule. In particular the `predict_ssh` method
#
#
# +
# sshdriver.predict_ssh?
# +
# Create a time vector
dt = 1800
numdays = 365
tstart = np.datetime64('2013-07-01 00:00:00')
nsteps = numdays*86400//dt
timeout = np.array([tstart+np.timedelta64(ii*dt,'s') for ii in range(nsteps)])
# Call the prediction funciton
ssh_ts = sshdriver.predict_ssh(ssh, xpt, ypt, timeout)
# Plot the time series
plt.figure()
plt.plot(timeout, ssh_ts, lw=0.25)
plt.xticks(rotation=17)
plt.ylabel('$SSH_{BC}$ [m]')
# -
# Maybe we also want to look at the contributions from the individual harmonics to this signal and how they vary. To do this use the `extract_amp_nonstat` function.
#
# This outputs two arrays, the real and imaginary amplitude, for each harmonic
# +
ssh_ns_re, ssh_ns_im = sshdriver.extract_amp_nonstat(ssh, np.array([xpt]), np.array([ypt]), timeout)
# To plot the amplitude of the first harmonic (M2) use the
plt.figure()
plt.plot(timeout, ssh_ts, '0.5', lw=0.25)
plt.plot(timeout, np.abs(ssh_ns_re[0,...] + 1j*ssh_ns_im[0,...]), 'r', ) # M2
plt.plot(timeout, np.abs(ssh_ns_re[3,...] + 1j*ssh_ns_im[3,...]), 'k', ) # K1
plt.xticks(rotation=17)
plt.ylabel('$SSH_{BC}$ [m]')
# -
# # Example: Extract the density stratification at a point
#
# Use the `iwatlas.stratification` module to extract density (buoyancy frequency) profile
# +
# strat.predict_N2?
# +
#
nz = 80 # Number of vertical layers to output
# Call the predict_N2 method. Note that this function expects numpy arrays, not scalars, for all inputs
# Only compute N^2 at a few time steps
tsteps = [0,7200,13200]
N2_z, zout = strat.predict_N2(ssh, np.array([xpt]), np.array([ypt]), timeout[tsteps], nz)
# -
# Convert N^2 to units of cycles per day
cpd = 2*np.pi/86400.
N_cpd = np.sqrt(N2_z)/cpd
86400/600.
# +
plt.figure()
plt.plot(N_cpd[:,0,0], -zout[:,0,0], )
plt.plot(N_cpd[:,0,1], -zout[:,0,1], )
plt.plot(N_cpd[:,0,2], -zout[:,0,2], )
plt.legend(timeout[tsteps])
plt.xlabel('$N$ [cpd]')
# -
# # Calculate the internal wave parameters
#
# +
# Calculate the internal wave parameters
mode= 0
# Mode shapes
phi_n, cn = iwaves.calc_modes(N2_z, zout, mode=mode)
# KdV nonlinearity parameter
alpha = iwaves.calc_alpha(phi_n, cn, zout)
# KdV dispersion parameter
beta = iwaves.calc_beta(phi_n, cn, zout)
# Factor to convert SSH to isotherm displacement amplitude
sshratio = iwaves.amp_to_ssh_ratio(N2_z, phi_n, zout)
print(cn[0,0], alpha[0,0], beta[0,0], sshratio[0,0])
# -
# # Example: convert SSH prediction to isotherm displacement amplitude
#
# This is simply the sshratio multiplied by the SSH
amp_ts = ssh_ts*sshratio[0,0]
plt.figure()
plt.plot(timeout, amp_ts, '0.5', lw=0.25)
plt.xticks(rotation=17)
plt.ylabel('Displacement Amp. [m]')
# # Example: convert SSH to a baroclinic velocity prediciton
# +
# Predict the time series of amplitude (note this needs to be expanded in the vertical direction)
ut, vt = uvdriver.predict_uv(ssh, np.array([xpt]), np.array([ypt]), timeout)
plt.figure(figsize=(12,6))
plt.plot(timeout, ut, lw=0.2)
plt.plot(timeout, vt, lw=0.2)
plt.ylabel('Velocity [m/s]')
# -
# %%time
# Compute the velocity as a function of z
# this requires calculating vertical mode function for every time step so may take a minute or two
uz, vz, zout = uvdriver.predict_uv_z(ssh, np.array([xpt]), np.array([ypt]), timeout)
# +
# Plot the surface velocity
usurf = uz[0,0,...]
vsurf = vz[0,0,...]
plt.figure(figsize=(12,6))
plt.plot(timeout, usurf,lw=0.2)
plt.plot(timeout, vsurf,lw=0.2)
plt.ylabel('Velocity [m/s]')
# -
| sandpit/tutorial_iwatlas_basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Example 1b: Linear convection in 2D, revisited
#
# We will now revisit the first example of this tutorial with an example that is better suited to the numerical scheme used in Devito. As a reminder, the governing equation is:
#
# $$\frac{\partial u}{\partial t}+c\frac{\partial u}{\partial x} + c\frac{\partial u}{\partial y} = 0$$
#
# We then discretized this using forward differences in time and backward differences in space:
#
# $$u_{i,j}^{n+1} = u_{i,j}^n-c \frac{\Delta t}{\Delta x}(u_{i,j}^n-u_{i-1,j}^n)-c \frac{\Delta t}{\Delta y}(u_{i,j}^n-u_{i,j-1}^n)$$
#
# In the previous example, the system was initialised with a hat function. As easy as this example seems, it actually highlights a few limitations of finite differences and related methods:
# - The governing equation above contains spatial derivatives ($\frac{\partial u}{\partial x}$ and $\frac{\partial u}{\partial y}$). The hat, with its sharp corners, is discontinuous and therefore non-smooth, meaning that the derivatives do not exist at the corners of the hat. This means that the governing equation has no solution in the strict sense for this problem. Mathematically, this problem can be overcome by introducing weak solutions, which still exist in the presence of discontinuities, as long as the problem is smooth almost everywhere. The Finite Volume (FV), Finite Element (FEM) and related schemes are based on this weak form.
# - The finite differences method only works well if finite differences are a good approximation of the derivatives. With the chosen discretization above, this requires that $\frac{u_{i,j}^n-u_{i-1,j}^n}{\Delta x} \approx \frac{\partial u}{\partial x}$ and $\frac{u_{i,j}^n-u_{i,j-1}^n}{\Delta y} \approx \frac{\partial u}{\partial y}$. This is the case for systems with a smooth solution if $\Delta x$ and $\Delta y$ are sufficiently small. But if the solution is non-smooth, as in this example, then we can't expect much regardless of the grid size.
# - First-order methods, such as the backward differences that we have used in this exampe, are known to create artificial diffusion. Higher-order schemes, such as central differences, avoid this problem. However, in the presence of discontinuities these methods introduce so-called spurious oscillations. These oscillations may even build up (grow infinitely) and cause the computation to diverge.
# - Discontinuities can appear by themselves for some equations (such as the Burgers equation that we discuss next), even if the intial condition is smooth. In CFD, discontinuities appear for example as shocks in the simulation of transonic flow. For this reason, numerical schemes that behave well in the presence of discontinuities have been a research subject for a long time. A thorough discussion is beyond the scope of this tutorial, but can be found in [<NAME> (1992): Numerical Methods for Conservation Laws, 2nd ed., <NAME>, pp. 8-13].
#
# In the remainder of this example, we will reproduce the results from the previous example, only this time with a smooth initial condition. This lets us observe Devito in a setting for which it is better equipped.
#
# +
from examples.cfd import plot_field, init_hat, init_smooth
import numpy as np
# %matplotlib inline
# Some variable declarations
nx = 81
ny = 81
nt = 100
c = 1.
dx = 2. / (nx - 1)
dy = 2. / (ny - 1)
sigma = .2
dt = sigma * dx
# -
# Let us now initialise the field with an infinitely smooth bump, as given by [<NAME> (2012): Unsteady Adjoint Analysis for Output Sensitivity
# and Mesh Adaptation, PhD thesis, p. 68] as $$
# f(r)=
# \begin{cases}
# \frac{1}{A}e^{-1/(r-r^2)} &\text{ for } 0 < r < 1,\\
# 0 &\text{ else.}
# \end{cases}
# $$
# We use this with $A=100$, and define the initial condition in two dimensions as $$u^0(x,y)=1+f\left(\frac{2}{3}x\right)*f\left(\frac{2}{3}y\right).$$
# +
#NBVAL_IGNORE_OUTPUT
# Create field and assign initial conditions
u = np.empty((nx, ny))
init_smooth(field=u, dx=dx, dy=dy)
# Plot initial condition
plot_field(u, zmax=4)
# -
# Solving this will move the bump again.
# +
# Repeat initialisation, so we can re-run the cell
init_smooth(field=u, dx=dx, dy=dy)
for n in range(nt + 1):
# Copy previous result into a new buffer
un = u.copy()
# Update the new result with a 3-point stencil
u[1:, 1:] = (un[1:, 1:] - (c * dt / dx * (un[1:, 1:] - un[1:, :-1])) -
(c * dt / dy * (un[1:, 1:] - un[:-1, 1:])))
# Apply boundary conditions
u[0, :] = 1.
u[-1, :] = 1.
u[:, 0] = 1.
u[:, -1] = 1.
# +
#NBVAL_IGNORE_OUTPUT
# A small sanity check for auto-testing
assert (u[45:55, 45:55] > 1.8).all()
u_ref = u.copy()
plot_field(u, zmax=4.)
# -
# Hooray, the wave moved! It looks like the solver works much better for this example: The wave has not noticeably changed its shape.
# #### Devito implementation
# Again, we can re-create this via a Devito operator. Let's fill the initial buffer with smooth data and look at it:
# +
#NBVAL_IGNORE_OUTPUT
from devito import Grid, TimeFunction
grid = Grid(shape=(nx, ny), extent=(2., 2.))
u = TimeFunction(name='u', grid=grid)
init_smooth(field=u.data[0], dx=dx, dy=dy)
plot_field(u.data[0])
# -
# We create again the discretized equation as shown below. Note that the equation is still the same, only the initial condition has changed.
# +
from devito import Eq
eq = Eq(u.dt + c*u.dxl + c*u.dyl)
print(eq)
# -
# SymPy can re-organise this equation just like in the previous example.
# +
from devito import solve
stencil = solve(eq, u.forward)
print(stencil)
# -
# We can now use this stencil expression to create an operator to apply to our data object:
# +
#NBVAL_IGNORE_OUTPUT
from devito import Operator
# Reset our initial condition in both buffers.
# This is required to avoid 0s propagating into
# our solution, which has a background value of 1.
init_smooth(field=u.data[0], dx=dx, dy=dy)
init_smooth(field=u.data[1], dx=dx, dy=dy)
# Apply boundary conditions
u.data[:, 0, :] = 1.
u.data[:, -1, :] = 1.
u.data[:, :, 0] = 1.
u.data[:, :, -1] = 1.
# Create an Operator that updates the forward stencil
# point in the interior subdomain only.
op = Operator(Eq(u.forward, stencil, subdomain=grid.interior))
# Apply the operator for a number of timesteps
op(time=nt, dt=dt)
plot_field(u.data[0, :, :])
# Some small sanity checks for the testing framework
assert (u.data[0, 45:55, 45:55] > 1.8).all()
assert np.allclose(u.data[0], u_ref, rtol=3.e-2)
# -
# Again, this looks just like the result from NumPy. Since this example is just like the one before, the low-level treatment of boundaries is also unchanged.
# +
#NBVAL_IGNORE_OUTPUT
# Reset our data field and ICs in both buffers
init_smooth(field=u.data[0], dx=dx, dy=dy)
init_smooth(field=u.data[1], dx=dx, dy=dy)
# For defining BCs, we generally to explicitly set rows/columns
# in our field using an expression. We can use Devito's "indexed"
# notation to do this:
x, y = grid.dimensions
t = grid.stepping_dim
bc_left = Eq(u[t + 1, 0, y], 1.)
bc_right = Eq(u[t + 1, nx-1, y], 1.)
bc_top = Eq(u[t + 1, x, ny-1], 1.)
bc_bottom = Eq(u[t + 1, x, 0], 1.)
# Now combine the BC expressions with the stencil to form operator.
expressions = [Eq(u.forward, stencil, subdomain=grid.interior)]
expressions += [bc_left, bc_right, bc_top, bc_bottom]
op = Operator(expressions=expressions, dle=None, dse=None) # <-- Turn off performance optimisations
op(time=nt, dt=dt)
plot_field(u.data[0, :, :])
# Some small sanity checks for the testing framework
assert (u.data[0, 45:55, 45:55] > 1.8).all()
assert np.allclose(u.data[0], u_ref, rtol=3.e-2)
# -
# The C code of the Kernel is also still the same.
print(op.ccode)
| examples/cfd/01_convection_revisited.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:anaconda]
# language: python
# name: conda-env-anaconda-py
# ---
# +
# %matplotlib inline
import networkx as nx
#import pygraphviz
import pyparsing
import numpy as np
import matplotlib.pylab as plt
from IPython.display import Math
# +
A = np.array([[0,1,1],[0,0,1],[1,0,0]])
G = nx.Graph(A)
labels = {0: "a", 1:"b", 2:"c"}
pos = [(0,0),(3,1),(1,0)]
plt.figure(figsize=(12,2.5))
nx.draw(G, pos, cmap='jet', edge_color=[0.1,0.7,0.9], node_color="white", node_size=500, labels=labels, font_size=10, arrows=True)
#nx.draw(G, pos, node_color="white", node_size=500, arrows=False)
#nx.draw_graphviz(G,node_size=500, labels=labels, font_size=24, arrows=True)
plt.show()
#nx.draw_networkx()
# +
from itertools import product
M = 6;
N = 6
Z = 1.7
NN = M*N
A = np.zeros((NN,NN))
X = np.zeros((NN))
Y = np.zeros((NN))
sig = 0.1;
coords = []
#cols = ("blue","red","yellow","black")
cols = ("black","black")
col = []
for i,j in product(range(N),range(M)):
ex = np.random.randn(1)*sig
ey = np.random.randn(1)*sig
coords.append((j,i))
X[i*M+j] = i+ex
Y[i*M+j] = j+ey
col.append(np.random.choice(cols))
for k,r in product(range(NN),range(NN)):
if k != r:
d = (X[k]-X[r])**2 + (Y[k]-Y[r])**2
A[k,r] = 1 if d < Z else 0
G = nx.Graph(A)
plt.figure(figsize=(M,N))
#nx.draw(G, pos, node_color="white", node_size=500, labels=labels, font_size=10, arrows=True)
nx.draw(G, coords, node_color='black', node_size=200, arrows=False, linewidths=14.)
nx.draw_networkx_nodes(G, coords, node_color='white', node_size=200, arrows=False, linewidths=11., linecolors='black')
#nx.draw_graphviz(G,node_size=500, labels=labels, font_size=24, arrows=True)
plt.show()
# +
#t = nx.dfs_tree(G,17)
t = nx.bfs_tree(G,1)
#t = nx.prim_mst(G)
plt.figure(figsize=(M,N))
nx.draw(t, coords, node_size=200,node_color="black",linewidths=14.)
nx.draw_networkx_nodes(t, coords, node_color="white", node_size=200,linewidths=11.)
plt.show()
# +
#nx.view_pygraphviz(G)
N = 20
#H = nx.random_graphs.watts_strogatz_graph(N,5,0.1)
H = nx.random_graphs.random_regular_graph(3,N)
lbl = {e:e for e in range(N)}
#nx.view_pygraphviz(H)
nx.draw_networkx_nodes(H,node_color="black",alpha=1, node_size=500, pos=nx.spectral_layout(H))
nx.draw(H,labels=lbl,node_color="white",alpha=1, node_size=400, pos=nx.spectral_layout(H))
# +
G = nx.Graph()
d = 10
G.add_node(0)
coord = [(0.5,0)]
depth = [0]
for n in range(2,256*4):
G.add_node(n-1)
p = int(np.floor(n/2))
depth.append(depth[p-1]+1)
if 2*p==n: # left child
ep = -(1.0/(2**(depth[p-1]+2)))
else:
ep = 1.0/(2**(depth[p-1]+2))
coord.append((coord[p-1][0]+ep,-(depth[p-1]+1)))
G.add_edge(n-1,p-1)
plt.figure(figsize=(35,6))
nx.draw(G, coord, node_size=50, node_color='black')
#nx.draw_shell
# +
import heapq
import numpy as np
N = 50
thr = 0.35
lb = 0.1
X = np.random.rand(N,2)
D = np.zeros((N,N))
for i,j in product(range(N),range(N)):
D[i,j] = np.sqrt((X[i,0]-X[j,0])**2 + (X[i,1]-X[j,1])**2)
if D[i,j]>thr or D[i,j]<lb :
D[i,j] = np.Inf
visited = np.empty(N,dtype=bool); visited.fill(False)
root = 0
visited[root] = True
numvis = 1;
spt = np.empty(N,dtype=int)
spt.fill(-1)
spt[root] = -1
q = []
for j in range(N):
if np.isfinite(D[root,j]):
heapq.heappush(q, (D[root,j], root, j))
while numvis<N:
if len(q)==0:
break;
d,i,j = heapq.heappop(q)
while len(q)>0 and visited[j]:
d,i,j = heapq.heappop(q)
spt[j] = i
visited[j] = True
numvis+=1
for k in range(N):
if np.isfinite(D[j,k]) and not visited[k]:
heapq.heappush(q, (D[j,k], j, k))
print(spt)
plt.figure(figsize=(10,10))
plt.plot(X[:,0],X[:,1],'o')
for i,j in product(range(N),range(N)):
if not np.isinf(D[i,j]):
plt.plot(X[[i,j],0],X[[i,j],1],'k:')
for u in range(N):
if spt[u]!=-1:
plt.plot(X[[u,spt[u]],0],X[[u,spt[u]],1],'r-')
plt.show()
# +
import matplotlib.pyplot as plt
import matplotlib.tri as tri
import numpy as np
import math
min_radius = 0.1
N = 100
x = np.random.rand(N)
y = np.random.rand(N)
# Create the Triangulation; no triangles so Delaunay triangulation created.
triang = tri.Triangulation(x, y)
# Mask off unwanted triangles.
xmid = x[triang.triangles].mean(axis=1)
ymid = y[triang.triangles].mean(axis=1)
mask = np.where(xmid*xmid + ymid*ymid < min_radius*min_radius, 1, 0)
triang.set_mask(mask)
# Plot the triangulation.
plt.figure(figsize=(5,5))
plt.gca().set_aspect('equal')
plt.triplot(triang, 'bo-')
plt.title('triplot of Delaunay triangulation')
plt.show()
# -
# * Traveling Salesman
# * Complete Binary tree
# * Spanning Tree
# * Bipartite Graph
# * Shortest Path Tree
import pygraphviz as pgv
from IPython.display import Image
from IPython.display import display
# +
def random_alphabet(N=20, first_letter='A'):
"""Generates unique strings to be used as index_names"""
if N<27:
alphabet = [chr(i+ord(first_letter)) for i in range(N)]
else:
alphabet = ['X'+str(i) for i in range(N)]
return alphabet
def random_parents(alphabet, max_indeg=3):
"""Random DAG generation"""
N = len(alphabet)
print(alphabet)
indeg = lambda: np.random.choice(range(1,max_indeg+1))
parents = {a:[b for b in np.random.choice(alphabet[0:(1 if i==0 else i)], replace=False, size=min(indeg(),i))] for i,a in enumerate(alphabet)}
return parents
def show_dag_image(index_names, parents, imstr='_BJN_tempfile.png', prog='dot'):
name2idx = {name: i for i,name in enumerate(index_names)}
A = pgv.AGraph(directed=True)
for i_n in index_names:
A.add_node(name2idx[i_n], label=i_n)
for j_n in parents[i_n]:
A.add_edge(name2idx[j_n], name2idx[i_n])
A.layout(prog=prog)
A.draw(imstr)
display(Image(imstr))
return
index_names = random_alphabet(10)
parents = random_parents(index_names, 3)
show_dag_image(index_names, parents, prog='neato')
# -
#
# Road Network
#
# We will build a 2D square grid where neighbors are connected
#
# Remove Random junctions for a more realistic view
# Compute a smooth height z by a linear dynamics
# Transform x,y,z and print
#
#
# +
import numpy as np
import scipy as sc
import pandas as pd
from itertools import product
def ind2idx(i,j, M, N):
return i + M*j
def idx2ind(k, M, N):
return k % M, k//M
def neigh(i,j, M, N):
ng = {'n': None, 's': None, 'w': None, 'e': None}
# north
if i>0:
ng['n'] = ind2idx(i-1,j,M,N)
# south
if i<M-1:
ng['s'] = ind2idx(i+1,j,M,N)
# west
if j>0:
ng['w'] = ind2idx(i,j-1,M,N)
#east
if j<N-1:
ng['e'] = ind2idx(i,j+1,M,N)
return ng
# Build a grid of junctions
M, N = 12,15
#ng = neigh(0,0,M,N)
#print(ng)
## Build the Adjecency list of the undirected graph
Adj = [[] for i in range(M*N)]
for j in range(N):
for i in range(M):
k = ind2idx(i,j,M,N)
ng = neigh(i,j,M,N)
south = ng['s']
if south is not None:
Adj[k].append(south)
Adj[south].append(k)
if np.random.rand()<0.8:
east = ng['e']
if east is not None:
Adj[k].append(east)
Adj[east].append(k)
# print(k,Adj[k])
# Kill a fraction of nodes randomly
kill = np.random.choice(range(M*N), size=M*N//10)
for k in kill:
for u in Adj[k]:
Adj[u].remove(k)
Adj[k] = []
## Place nodes on a perturbed grid
X = 0.9*np.random.rand(N) + np.arange(0, N)
Y = 0.9*np.random.rand(M) + np.arange(0, M)
Coords = np.zeros((M*N, 3))
for k in range(M*N):
i, j = idx2ind(k, M, N)
Coords[k, 0] = X[j]+0.1*np.random.randn()
Coords[k, 1] = Y[i]+0.1*np.random.randn()
Coords[k, 2] = np.random.rand()
## Iterate to get a smooth terrain
EPOCHS = 30
for e in range(EPOCHS):
perm = np.random.permutation(M*N)
for k in perm:
if Adj[k]:
Coords[k,2] = 0.9*Coords[k,2] + 0.1*np.mean(Coords[Adj[k],2])
# -
plot_topology(Adj, M, N)
# +
merge = np.random.choice(range(M*N), replace=False, size=30)
for u in merge:
if Adj[u]:
v = np.random.choice(Adj[u])
# Disconnect v from u
Adj[v].remove(u)
Adj[u].remove(v)
## transfer all the remaining edges to v
for w in Adj[u]:
if w not in Adj[v]:
Adj[v].append(w)
Adj[w].append(v)
Adj[w].remove(u)
Adj[u] = []
# -
plot_topology(Adj, M, N)
# +
## Print node coordinates
for k in range(M*N):
print("%2.1f, %2.1f, %2.1f" % (Coords[k, 0], Coords[k, 1], Coords[k, 0]))
# Print Edges
for k in range(M*N):
for u in Adj[k]:
print('%d,%d' % (k,u))
# +
import matplotlib.pylab as plt
def plot_topology(Adj, M, N):
plt.figure(figsize=(10,10))
for k,ls in enumerate(Adj):
i,j = idx2ind(k, M,N)
for u in ls:
i_target, j_target = idx2ind(u, M,N)
plt.plot([j, j_target ],[i, i_target],'k')
if Adj[k]:
plt.plot(j, i,'ro')
plt.show()
plot_topology(Adj, M, N)
# +
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
fig = plt.figure(figsize=(10,10))
ax = fig.add_subplot(111, projection='3d')
for k,ls in enumerate(Adj):
for u in ls:
ax.plot([Coords[k,0], Coords[u,0] ],[Coords[k,1], Coords[u,1] ], [Coords[k,2], Coords[u,2] ],'k')
if Adj[k]:
ax.plot([Coords[k,0]], [Coords[k,1]], [Coords[k,2]], 'ro')
ax.set_zlim([0, 1])
plt.show()
# +
for k in range(M*N):
print("%2.1f, %2.1f, %2.1f" % (Coords[k, 0], Coords[k, 1], Coords[k, 0]))
for k in range(M*N):
for u in Adj[k]:
print('%d,%d' % (k,u))
# +
G = nx.Graph(A)
plt.figure(figsize=(M,N))
#nx.draw(G, pos, node_color="white", node_size=500, labels=labels, font_size=10, arrows=True)
nx.draw(G, coords, node_color='black', node_size=200, arrows=False, linewidths=14.)
nx.draw_networkx_nodes(G, coords, node_color='white', node_size=200, arrows=False, linewidths=11., linecolors='black')
#nx.draw_graphviz(G,node_size=500, labels=labels, font_size=24, arrows=True)
plt.show()
# +
import itertools
import numpy as np
import matplotlib.pylab as plt
import daft
# Instantiate the PGM.
pgm = daft.PGM([3.6, 3.6], origin=[0.7, 0.7], node_unit=0.4, grid_unit=1,
directed=False)
for i, (xi, yi) in enumerate(itertools.product(range(1, 5), range(1, 5))):
pgm.add_node(daft.Node(str(i), "", xi, yi))
for e in [(4, 9), (6, 7), (3, 7), (10, 11), (10, 9), (10, 14),
(10, 6), (10, 7), (1, 2), (1, 5), (1, 0), (1, 6), (8, 12), (12, 13),
(13, 14), (15, 11)]:
pgm.add_edge(str(e[0]), str(e[1]))
# Render and save.
pgm.render()
#pgm.figure.savefig("mrf.pdf")
#pgm.figure.savefig("mrf.png", dpi=150)
plt.show(pgm.ax)
# +
from matplotlib import rc
rc("font", family="serif", size=12)
rc("text", usetex=True)
import daft
pgm = daft.PGM([3.6, 2.4], origin = [1.15, 0.8], node_ec="none")
pgm.add_node(daft.Node("cloudy", r"cloudy", 3, 3))
pgm.add_node(daft.Node("rain", r"rain", 2, 2))
pgm.add_node(daft.Node("sprinkler", r"sprinkler", 4, 2))
pgm.add_node(daft.Node("wet", r"grass wet", 3, 1))
pgm.add_edge("cloudy", "rain")
pgm.add_edge("cloudy", "sprinkler")
pgm.add_edge("rain", "wet")
pgm.add_edge("sprinkler", "wet")
pgm.render()
plt.show(pgm.ax)
# +
from matplotlib import rc
ff = "comic sans ms"
# ff = "impact"
# ff = "times new roman"
rc("font", family=ff, size=12)
rc("text", usetex=False)
import daft
pgm = daft.PGM([3.6, 1.8], origin=[2.2, 1.6], aspect=2.1)
pgm.add_node(daft.Node("confused", r"confused", 3.0, 3.0))
pgm.add_node(daft.Node("ugly", r"ugly font", 3.0, 2.0, observed=True))
pgm.add_node(daft.Node("bad", r"bad talk", 5.0, 2.0, observed=True))
pgm.add_edge("confused", "ugly")
pgm.add_edge("ugly", "bad")
pgm.add_edge("confused", "bad")
pgm.render()
plt.show(pgm.ax)
# +
from matplotlib import rc
rc("font", family="serif", size=12)
rc("text", usetex=True)
import daft
# Instantiate the PGM.
pgm = daft.PGM([2.3, 2.05], origin=[0.3, 0.3])
# Hierarchical parameters.
pgm.add_node(daft.Node("alpha", r"$\alpha$", 0.5, 2, fixed=True))
pgm.add_node(daft.Node("beta", r"$\beta$", 1.5, 2))
# Latent variable.
pgm.add_node(daft.Node("w", r"$w_n$", 1, 1))
# Data.
pgm.add_node(daft.Node("x", r"$x_n$", 2, 1, observed=True))
# Add in the edges.
pgm.add_edge("alpha", "beta")
pgm.add_edge("beta", "w")
pgm.add_edge("w", "x")
pgm.add_edge("beta", "x")
# And a plate.
pgm.add_plate(daft.Plate([0.5, 0.5, 2, 1], label=r"$n = 1, \cdots, N$",
shift=-0.1))
# Render and save.
pgm.render()
plt.show(pgm.ax)
# +
from matplotlib import rc
rc("font", family="serif", size=12)
rc("text", usetex=True)
import daft
# Colors.
p_color = {"ec": "#46a546"}
s_color = {"ec": "#f89406"}
pgm = daft.PGM([3.6, 3.5], origin=[0.7, 0])
n = daft.Node("phi", r"$\phi$", 1, 3, plot_params=s_color)
n.va = "baseline"
pgm.add_node(n)
pgm.add_node(daft.Node("speckle_coeff", r"$z_i$", 2, 3, plot_params=s_color))
pgm.add_node(daft.Node("speckle_img", r"$x_i$", 2, 2, plot_params=s_color))
pgm.add_node(daft.Node("spec", r"$s$", 4, 3, plot_params=p_color))
pgm.add_node(daft.Node("shape", r"$g$", 4, 2, plot_params=p_color))
pgm.add_node(daft.Node("planet_pos", r"$\mu_i$", 3, 3, plot_params=p_color))
pgm.add_node(daft.Node("planet_img", r"$p_i$", 3, 2, plot_params=p_color))
pgm.add_node(daft.Node("pixels", r"$y_i ^j$", 2.5, 1, observed=True))
# Edges.
pgm.add_edge("phi", "speckle_coeff")
pgm.add_edge("speckle_coeff", "speckle_img")
pgm.add_edge("speckle_img", "pixels")
pgm.add_edge("spec", "planet_img")
pgm.add_edge("shape", "planet_img")
pgm.add_edge("planet_pos", "planet_img")
pgm.add_edge("planet_img", "pixels")
# And a plate.
pgm.add_plate(daft.Plate([1.5, 0.2, 2, 3.2], label=r"exposure $i$",
shift=-0.1))
pgm.add_plate(daft.Plate([2, 0.5, 1, 1], label=r"pixel $j$",
shift=-0.1))
# Render and save.
pgm.render()
plt.show(pgm.ax)
# -
# %connect_info
| DrawGraphs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# [View in Colaboratory](https://colab.research.google.com/github/raahatg21/MNIST-Dataset-using-TensorFlow/blob/master/1_Logistic_Regression.ipynb)
# + [markdown] id="HE7G0z5BROXl" colab_type="text"
# # MNIST Dataset: Logistic Regression
# + [markdown] id="2yNY4JD9BjYy" colab_type="text"
# Classification on MNIST Dataset using basic Logistic Regression using TensorFlow. Test Accuracy: 90.9%
# + id="PH0a9VEuPO9R" colab_type="code" colab={}
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
# + id="X0Jw2scmTUtN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 419} outputId="bd3904fc-3a2a-4da3-ac68-f7df7aeb221c"
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets("MNIST_data/", one_hot = True)
# + id="-JPr0QKzTtgo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 104} outputId="da58fd15-acac-471d-8348-368b385cdb79"
# Setup to use TensorBoard in Google Colab
# !wget https://bin.equinox.io/c/4VmDzA7iaHb/ngrok-stable-linux-amd64.zip
# !unzip ngrok-stable-linux-amd64.zip
# + id="gW09tEzKUWVX" colab_type="code" colab={}
LOG_DIR = './log'
get_ipython().system_raw(
'tensorboard --logdir {} --host 0.0.0.0 --port 6006 &'
.format(LOG_DIR)
)
# + id="nvc-RdZYUb2H" colab_type="code" colab={}
get_ipython().system_raw('./ngrok http 6006 &')
# + id="oD2uer6IUlbq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="0c18feb1-a185-47ff-c6e3-13f1277d0f60"
# ! curl -s http://localhost:4040/api/tunnels | python3 -c \
# "import sys, json; print(json.load(sys.stdin)['tunnels'][0]['public_url'])"
# + id="auPastjDUqUR" colab_type="code" colab={}
# Defining Hyperparameters
learning_rate = 0.05
batch_size = 100
max_epochs = 100
# + id="n0LwQe6wU46c" colab_type="code" colab={}
# Defining Placeholders
X = tf.placeholder(tf.float32, [None, 784])
Y = tf.placeholder(tf.float32, [None, 10])
# + id="XT8eCDx8VFpn" colab_type="code" colab={}
# Defining weights and bias variables
w = tf.Variable(tf.zeros([784, 10]))
b = tf.Variable(tf.zeros([10]))
# + id="gxusg6hhVRIV" colab_type="code" colab={}
# Defining Logistic Regression function
with tf.name_scope("wx_b") as scope:
Y_hat = tf.nn.softmax(tf.matmul(X, w) + b)
# + id="CXsg7SKcVgkF" colab_type="code" colab={}
# Defining Summary Ops
w_h = tf.summary.histogram("weights", w)
b_h = tf.summary.histogram("bias", b)
# + id="MiOIkkBgV0K7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 176} outputId="2b6b7b68-cd61-4695-c808-b6ab41ec2e43"
# Defining Cross-entropy Loss
with tf.name_scope("Cross-entropy") as scope:
loss = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(labels = Y, logits = Y_hat))
tf.summary.scalar("cross-entropy", loss)
# + id="1QZUEcWeWUT_" colab_type="code" colab={}
# Defining the Optimiser
with tf.name_scope("Train") as scope:
optimizer = tf.train.GradientDescentOptimizer(0.01).minimize(loss)
# + id="lr9tGjzvWutp" colab_type="code" colab={}
# Defining the Metric
correct_preds = tf.equal(tf.argmax(Y_hat, 1), tf.argmax(Y, 1))
accuracy = tf.reduce_mean(tf.cast(correct_preds, tf.float32))
# + id="lpuuZ8cBXBSb" colab_type="code" colab={}
init = tf.global_variables_initializer()
# + id="uOb2kdjgXInQ" colab_type="code" colab={}
merged_summary_op = tf.summary.merge_all()
# + id="3wCyMK8NXNOj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1771} outputId="c0ac2a4c-7bc6-4b24-c9cb-aa1d6068e0fa"
with tf.Session() as sess:
sess.run(init)
summary_writer = tf.summary.FileWriter('./log', sess.graph)
for epoch in range(max_epochs):
loss_avg = 0
batches = int(mnist.train.num_examples/batch_size)
for i in range(batches):
batch_X, batch_y = mnist.train.next_batch(100)
_, l, summary_str = sess.run([optimizer, loss, merged_summary_op], feed_dict = {X: batch_X, Y: batch_y})
loss_avg += l
summary_writer.add_summary(summary_str, epoch*batches + i)
loss_avg /= batches
print("Epoch {0}: Loss {1}".format(epoch, loss_avg))
print(sess.run(accuracy, feed_dict = {X: mnist.test.images, Y: mnist.test.labels})) # Testing
# + id="2fJO34rKZLjG" colab_type="code" colab={}
| 1_Logistic_Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Pascal's Triangle II
# + active=""
# Given a non-negative index k where k ≤ 33, return the kth index row of the Pascal's triangle.
# Note that the row index starts from 0.
# In Pascal's triangle, each number is the sum of the two numbers directly above it.
# Example:
# Input: 3
# Output: [1,3,3,1]
#
# Follow up:
# Could you optimize your algorithm to use only O(k) extra space?
# -
# https://upload.wikimedia.org/wikipedia/commons/0/0d/PascalTriangleAnimated2.gif
# 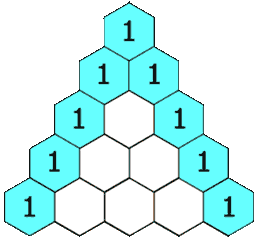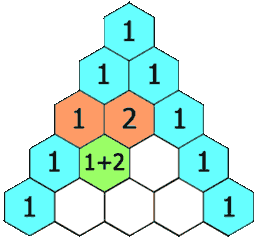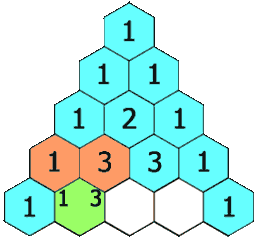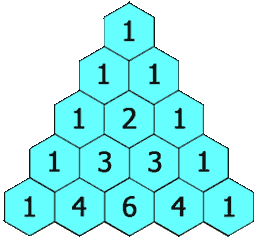
# In Pascal's triangle, each number is the sum of the two numbers directly above it
class Solution:
def getRow(self, rowIndex): # 50.05%
"""
:type rowIndex: int
:rtype: List[int]
"""
if rowIndex == 0:
return [1]
elif rowIndex == 1:
return [1, 1]
else:
newlst = [1] * (rowIndex+1)
lst = self.getRow(rowIndex-1)
for i in range(rowIndex//2):
newlst[i+1] = lst[i] + lst[i+1]
newlst[(rowIndex+1)//2:] = newlst[:(rowIndex+1)-(rowIndex+1)//2][::-1]
return newlst
rowIndex = 5
ans = Solution()
ans.getRow(rowIndex)
| 119. Pascal_s Triangle II.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # North Atlantic Basin Hurricane Predictor Model
# 
# <center>(Figure 1) Hurricane Zones</center>
# Import modules
import pandas as pd
import numpy as np
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier, AdaBoostClassifier
from sklearn.svm import SVC
from sklearn.neighbors import KNeighborsClassifier
from sklearn.model_selection import train_test_split, cross_val_score, GridSearchCV
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
from sklearn import metrics
from imblearn.over_sampling import RandomOverSampler
import matplotlib.pyplot as plt
# Import data
df = pd.read_csv('./data_preprocessed.csv')
df.drop('Unnamed: 0', axis=1, inplace=True)
# Set independent and dependent variables
# df = df.sample(5000)
X = df.drop(['A', 'B', 'C', 'D', 'E', 'F', 'G', 'H', 'I', 'J', 'K'], axis=1)
y = df[['A', 'B', 'C', 'D', 'E', 'F', 'G', 'H', 'I', 'J', 'K']]
# Split data into training and test sets
X_train, X_test, y_train, y_test = train_test_split(X,
y,
test_size=0.25,
random_state=42,
shuffle=True)
# ### Determining Our Approach
# Balancing classes and standardize independent variables
# Oversamping 'hits' to balance classes
# Let's look at zone "G"
ros = RandomOverSampler(random_state=42)
X_trainG, y_trainG = ros.fit_sample(X_train, y_train.G)
scaler = StandardScaler()
X_trainG = scaler.fit_transform(X_trainG)
X_test = scaler.transform(X_test)
# Balancing classes and standarding features greatly improved F1 score
# Baseline for a storm striking zone "G" (Southern Florida)
1 - y.G.sum() / y.G.count()
# The classes aren't balanced with 90.2% of storm events missing zone "G"
# #### Preliminary models
clf = LogisticRegression()
print(cross_val_score(clf, X_trainG, y_trainG, cv=3, n_jobs=-1).mean())
print(cross_val_score(clf, X_trainG, y_trainG, scoring='f1', cv=3, n_jobs=-1).mean())
clf = RandomForestClassifier()
print(cross_val_score(clf, X_trainG, y_trainG, cv=3, n_jobs=-1).mean())
print(cross_val_score(clf, X_trainG, y_trainG, scoring='f1', cv=5, n_jobs=-1).mean())
clf = SVC()
print(cross_val_score(clf, X_trainG, y_trainG, cv=3, n_jobs=-1).mean())
print(cross_val_score(clf, X_trainG, y_trainG, scoring='f1', cv=3, n_jobs=-1).mean())
clf = KNeighborsClassifier()
print(cross_val_score(clf, X_trainG, y_trainG, cv=3, n_jobs=-1).mean())
print(cross_val_score(clf, X_trainG, y_trainG, scoring='f1', cv=3, n_jobs=-1).mean())
clf = AdaBoostClassifier()
print(cross_val_score(clf, X_trainG, y_trainG, cv=3, n_jobs=-1).mean())
print(cross_val_score(clf, X_trainG, y_trainG, scoring='f1', cv=3, n_jobs=-1).mean())
# After at glance at the different classifiers, Random Forest jumps out as a strong candidate
# ## Hypertuning and Evaluating Models
# #### Random Forest
parms = {
'n_estimators': [46],
'max_depth': [97],
}
model = RandomForestClassifier()
clf = GridSearchCV(model, parms, cv=5, scoring='f1', n_jobs=-1, verbose=1, error_score=0)
clf.fit(X_trainG, y_trainG)
print(clf.best_score_,
clf.best_estimator_)
print('Test evaluation: ',
metrics.f1_score(y_test.G, clf.predict(X_test)))
# Overfit the random forest model
# #### K-Nearest Neighbors
parms = {
'n_neighbors': [1],
'p': [2]
}
model = KNeighborsClassifier()
clf = GridSearchCV(model, parms, cv=5, scoring='f1', n_jobs=-1, verbose=1, error_score=0)
clf.fit(X_trainG, y_trainG)
print(clf.best_score_,
clf.best_estimator_)
# print('Test evaluation: ',
# metrics.f1_score(y_test.G, clf.predict(X_test)))
model = KNeighborsClassifier(n_neighbors=1, p=2)
clf = model.fit(X_trainG, y_trainG)
print('Test evaluation: ',
metrics.f1_score(y_test.G, clf.predict(X_test)))
# Overfit KNN model
# #### Logistic Regression
parms = {
'C': np.linspace(28, 40, 2)
}
model = LogisticRegression()
clf = GridSearchCV(model, parms, cv=5, scoring='f1', n_jobs=-1, verbose=1, error_score=0)
clf.fit(X_trainG, y_trainG)
print(clf.best_score_,
clf.best_estimator_)
# print('Test evaluation: ',
# metrics.f1_score(y_test.G, clf.predict(X_test)))
model = LogisticRegression(C=28)
clf = model.fit(X_trainG, y_trainG)
print('Test evaluation: ',
metrics.f1_score(y_test.G, clf.predict(X_test)))
# Logistic regression did not perform well
# #### Support Vector Machine
parms = {
'kernel': ['rbf'],
'C': range(1, 25, 5)
}
model = SVC()
clf = GridSearchCV(model, parms, cv=3, scoring='f1', n_jobs=-1, verbose=1, error_score=0)
clf.fit(X_trainG, y_trainG)
print(clf.best_score_,
clf.best_estimator_)
# Long runtime to fit SVM model
(12140 * 338) / df.shape[0]
# ##### PCA
# Using PCA to reduce dataset in order to run support vector machine
pca = PCA(110)
X_trainG_pca = pca.fit_transform(X_trainG)
X_test_pca = pca.transform(X_test)
pca.explained_variance_ratio_.sum()
parms = {
'kernel': ['rbf'],
'C': range(1, 25, 5)
}
model = SVC()
clf = GridSearchCV(model, parms, cv=3, scoring='f1', n_jobs=-1, verbose=1, error_score=0)
clf.fit(X_trainG_pca, y_trainG)
print(clf.best_score_,
clf.best_estimator_)
# %%time
model = SVC(kernel='rbf', C=11)
clf = model.fit(X_trainG_pca, y_trainG)
print('Test evaluation: ',
metrics.f1_score(y_test.G, clf.predict(X_test_pca)))
# SVC takes too long to train
# PCA + kNN
# %%time
for n in range(10, 220, 10):
pca = PCA(n)
X_trainG_pca = pca.fit_transform(X_trainG)
X_test_pca = pca.transform(X_test)
print('PCA: ', n)
print('Explained variance: ', pca.explained_variance_ratio_.sum())
clf = KNeighborsClassifier(1)
clf.fit(X_trainG_pca, y_trainG)
print('Test evaluation: ',
metrics.f1_score(y_test.G, clf.predict(X_test_pca)))
print('***')
# Curse of dimensionality does not have a significant effect
# %%time
model = KNeighborsClassifier(n_neighbors=1, p=2)
clf = model.fit(X_trainG, y_trainG)
preds = clf.predict(X_test)
print('Test evaluation: ',
metrics.f1_score(y_test.G, preds))
print(metrics.classification_report(y_test.G, preds, target_names=['Miss', 'Hit']))
print(metrics.confusion_matrix(y_test.G, preds))
# # Training all Zones with kNN (k=1)
model = {}
Xtrain = {}
Xtest = {}
ytrain = {}
ytest = {}
baseline = {}
preds = {}
f1_scores = {}
for zone in 'ABCDEFGHIJK':
print('*** Zone: ', zone, ' ***')
df = pd.read_csv('./data_preprocessed.csv')
df.drop('Unnamed: 0', axis=1, inplace=True)
# df = df.sample(5000)
X = df.drop(['A', 'B', 'C', 'D', 'E', 'F', 'G', 'H', 'I', 'J', 'K'], axis=1)
y = df[['A', 'B', 'C', 'D', 'E', 'F', 'G', 'H', 'I', 'J', 'K']]
X_train, X_test, y_train, y_test = train_test_split(X,
y,
test_size=0.2,
random_state=42,
shuffle=True)
ros = RandomOverSampler(random_state=42)
X_train_resampled, y_train_resampled = ros.fit_sample(X_train, y_train[zone])
scaler = StandardScaler()
X_train_resampled_ss = scaler.fit_transform(X_train_resampled)
X_test_ss = scaler.transform(X_test)
# model[zone] = KNeighborsClassifier(1).fit(X_train_resampled_ss, y_train_resampled)
Xtrain[zone] = X_train_resampled_ss
ytrain[zone] = y_train_resampled
Xtest[zone] = X_test_ss
ytest[zone] = y_test[zone]
baseline[zone] = 1 - y[zone].sum() / y[zone].count()
# preds[zone] = model[zone].predict(Xtest[zone])
# f1_scores[zone] = metrics.f1_score(ytest[zone], preds[zone])
# print(zone, f1_scores[zone])
# Let's see if neural nets improve our results
# Import keras modules
from keras.models import Sequential
from keras.layers.core import Dense, Dropout, Activation, Flatten
from keras.layers import Convolution1D, MaxPooling1D
from keras import regularizers
from keras.layers.normalization import BatchNormalization
from keras.layers.advanced_activations import *
from keras.optimizers import Adam
from keras.initializers import *
from keras.models import model_from_json
for zone in 'ABCDEFGHIJK':
''' For each zone, training a deep neural net
topredict the zone that storm will hit '''
# Initialize model
model = Sequential()
input_units = Xtrain[zone].shape[1]
hidden_units = input_units
# Topology: n(relu, 40% dropout) --> n+30(relu, 20% dropout)
# --> 12(relu) --> 1 output(sigmoid)
model.add(Dense(hidden_units,
input_dim=input_units,
activation='relu'))
model.add(Dropout(0.4))
model.add(Dense(hidden_units + 30,
activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(12,
activation='relu'))
model.add(Dense(1, activation='sigmoid'))
model.compile(loss='binary_crossentropy',
optimizer='Adam',
metrics=['accuracy'])
hist = model.fit(Xtrain[zone],
ytrain[zone],
validation_data=(Xtest[zone], ytest[zone]),
epochs=125,
batch_size=256,
shuffle=True,
verbose=0)
# Saving model and weights to .json and .h5
# https://machinelearningmastery.com/save-load-keras-deep-learning-models/
model_json = model.to_json()
with open('./neural-models/' + zone + '.json', 'w') as json_file:
json_file.write(model_json)
model.save_weights('./neural-models/' + zone + '.h5')
# Plotting loss funciton vs epochs
plt.figure()
plt.plot(hist.history['loss'])
plt.plot(hist.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='best') ## I love this loc = 'best' command.
plt.show()
# Plotting accuracy vs epochs
plt.figure()
plt.plot(hist.history['acc'])
plt.plot(hist.history['val_acc'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='best')
plt.show()
hist.history['val_acc'][-1]
# F1 score on test set
print('Test F1 Score: ', zone, metrics.f1_score(ytest[zone],
np.round(model.predict(Xtest[zone]))))
# Zone "K" results
metrics.confusion_matrix(ytest['K'], np.round(model.predict(Xtest['K'])))
metrics.accuracy_score(ytest['K'], np.round(model.predict(Xtest['K'])))
| -bak/Model.ipynb |
/ ---
/ jupyter:
/ jupytext:
/ text_representation:
/ extension: .q
/ format_name: light
/ format_version: '1.5'
/ jupytext_version: 1.14.4
/ kernelspec:
/ display_name: SQL
/ language: sql
/ name: SQL
/ ---
/ + [markdown] azdata_cell_guid="94dc5a40-ef57-4733-b53a-1dfcbe2ead58"
/ # SQL Server 2019 Data Virtualization - Using Polybase to query Azure CosmosDB
/ This notebook contains an example of how to use external tables to query data in Azure CosmosDB (using MongoDB API) without moving data. You may need to change identity, secret, connection, database, schema, and remote table names to work with your Azure CosmosDB.
/
/ This notebook also assumes you are using SQL Server 2019 Release Candidate or later and that the Polybase feature has been installed and enabled.
/
/ This notebook uses the sample WideWorldImporters sample database but can be used with any user database.
/ + [markdown] azdata_cell_guid="678fd495-dc2c-4986-94ff-49d4d3e80d8f"
/ ## Step 0: Create the CosmosDB database, document, collection, and add data
/
/ Create a new database, collection, and document with CosmosDB in Azure. You can the Azure portal to create a new Azure CosmosDB database (choose for Mongo API). Use the Data Explorer tool from the portal to create a database called **WideWorldImporters** with a collection called **Orders**. Then create a new document with field names and values like the following (Note: the _id field was created by Data Explorer and the id field was a default value already provided by the tool)
/
/
/ ```json
/ {
/ "_id" : ObjectId("5c54aa72dd13c70f445745bf"),
/ "id" : "1",
/ "OrderID" : 1,
/ "SalesPersonPersonID" : 2,
/ "CustomerName" : "<NAME>",
/ "CustomerContact" : "<NAME>",
/ "OrderDate" : "2018-05-14",
/ "CustomerPO" : "20180514",
/ "ExpectedDeliveryDate" : "2018-05-21"
/ }
/ ```
/ + [markdown] azdata_cell_guid="15cffd28-c5d1-428c-b1b7-9312c0bfc4a5"
/ ## Step 1: Create a master key
/ Create a master key to encrypt the database credential
/ + azdata_cell_guid="52585c08-4689-4b5f-bcf8-9434e7853053"
USE [WideWorldImporters]
GO
CREATE MASTER KEY ENCRYPTION BY PASSWORD = '<password>'
GO
/ + [markdown] azdata_cell_guid="62ab9558-5fe0-4fb9-8046-e4631795d0a7"
/ ## Step 2: Create a database credential
/ Create the database scoped credentials with the Azure CosmosDB user and password. You can get the IDENTITY (user) and secret (password) from the Connection String option in the Azure portal
/ + azdata_cell_guid="e42b9c0f-1875-47cc-b8f1-2de488413692"
CREATE DATABASE SCOPED CREDENTIAL CosmosDBCredentials
WITH IDENTITY = '<user>', Secret = '<password>'
GO
/ + [markdown] azdata_cell_guid="390ae534-1b64-44a2-9a41-68c1915e5ba1"
/ ## Step 3: Create an EXTERNAL DATA SOURCE
/ The EXTERNAL DATA SOURCE indicates what type of data source, the connection "string", where PUSHDOWN predicates should be used (if possible), and the name of the database credential.
/
/ The LOCATION syntax is <datasourcetype>:<connection string>.
/
/ datasourcetype can be sqlserver, oracle, teradata, mongodb, or odbc (Windows only)
/ The connection string depends on the datasourcetype
/
/ For this example, create a data source for the Azure CosmoDB sderver using the host URI and port. The LOCATION is built from <HOST>:<PORT> from the Connection String in the Azure Portal
/ + azdata_cell_guid="85e81b03-6570-4cd1-ac59-bbe06154d6a1"
CREATE EXTERNAL DATA SOURCE CosmosDB
WITH (
LOCATION = 'mongodb://<uri>:<port>',
PUSHDOWN = ON,
CREDENTIAL = CosmosDBCredentials
)
GO
/ + [markdown] azdata_cell_guid="312f8c6e-f364-4cae-be9d-329dfe88bff2"
/ ## Step 4: Create a schema for the EXTERNAL TABLE
/ Schemas provide convenient methods to secure and organize objects
/ + azdata_cell_guid="1e134575-6506-4b19-835b-c226653b8ae9"
CREATE SCHEMA cosmosdb
GO
/ + [markdown] azdata_cell_guid="d8b61725-50e9-43c3-8da2-e9f4d99fac79"
/ ## Step 5: Create an EXTERNAL TABLE
/ An external table provides metadata so SQL Server knows how to map columns to the remote table. The name of the table for the external table can be your choice. But the columns must be specified with the same name as they are defined in the remote table. Furthermore, local data types must be compatible with the remote table.
/
/ Create the external table to match the Azure CosmosDB document. The WITH clause specifies a LOCATION. This LOCATION is different than the EXTERNAL DATA SOURCE. The LOCATION is the CosmosDB database and collection.
/ + azdata_cell_guid="50c7ad4f-bded-4ede-8404-9a0d8ef8c253"
CREATE EXTERNAL TABLE cosmosdb.Orders
(
[_id] NVARCHAR(100) COLLATE Latin1_General_100_CI_AS NOT NULL,
[id] NVARCHAR(100) COLLATE Latin1_General_100_CI_AS NOT NULL,
[OrderID] int NOT NULL,
[SalesPersonPersonID] int NOT NULL,
[CustomerName] NVARCHAR(100) COLLATE Latin1_General_100_CI_AS NOT NULL,
[CustomerContact] NVARCHAR(100) COLLATE Latin1_General_100_CI_AS NOT NULL,
[OrderDate] NVARCHAR(100) COLLATE Latin1_General_100_CI_AS NOT NULL,
[CustomerPO] NVARCHAR(100) COLLATE Latin1_General_100_CI_AS NULL,
[ExpectedDeliveryDate] NVARCHAR(100) COLLATE Latin1_General_100_CI_AS NOT NULL
)
WITH (
LOCATION='WideWorldImporters.Orders',
DATA_SOURCE=CosmosDB
)
GO
/ + [markdown] azdata_cell_guid="56e86b58-f275-48bf-ae46-bcf98a432e7c"
/ ## Step 6: Create statistics
/ SQL Server allows you to store local statistics about specific columns from the remote table. This can help the query processing to make more efficient plan decisions.
/ + azdata_cell_guid="aaa8950d-da30-4691-a10c-5645f688688c"
CREATE STATISTICS CosmosDBOrderSalesPersonStats ON cosmosdb.Orders ([SalesPersonPersonID]) WITH FULLSCAN
GO
/ + [markdown] azdata_cell_guid="2a0cd685-98c4-4a30-84c8-106ad7882264"
/ ## Step 7: Try to scan the remote table
/ Run a simple query on the EXTERNAL TABLE to scan all rows.
/ + azdata_cell_guid="947368be-f370-4f9a-a39f-ca440a32f8f4"
SELECT * FROM cosmosdb.Orders
GO
/ + [markdown] azdata_cell_guid="3409e8ac-49b4-4e97-85bc-349f67aaf042"
/ ## Step 8: Query the remote table with a WHERE clause
/ Even though the table may be small SQL Server will "push" the WHERE clause filter to the remote table
/ + azdata_cell_guid="2b3a9bfa-7269-452a-b968-545cf0314a30"
SELECT * FROM cosmosdb.Orders WHERE SalesPersonPersonID = 2
GO
/ + [markdown] azdata_cell_guid="735f001b-8067-4465-a80a-d1a34324c6b7"
/ ## Step 9: Join with local SQL Server tables
/ Find out the name of the salesperson and which customer they worked with to test out the new mobile app experience.
/ + azdata_cell_guid="dbf869ac-1969-4c50-bd82-215038da238f"
SELECT FullName, o.CustomerName, o.CustomerContact, o.OrderDate
FROM cosmosdb.Orders o
JOIN [Application].[People] p
ON o.SalesPersonPersonID = p.PersonID
GO
/ + [markdown] azdata_cell_guid="8d86be35-d207-434a-9cdb-06962ee44dc8"
/
| sql2019workshop/sql2019wks/08_DataVirtualization/sqldatahub/cosmosdb/cosmosdbexternaltable.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#hide
from intel_invest.core import *
# # intel_invest
#
# > A package to obtain the relevant figures for a company and apply some rules of thumb precribed in the book Intelligent Investor, by <NAME>.
# ## Install
# `pip install intel_invest`
# ## How to use
# To obtain Graham's opinion on `Abbott Laboratories` do the following:
stock = Stock("ABT")
# Ben will ask two questions:
#
# - is price < 25 * average 5 year earning per share
# - is price to earning ratio * price to book value < 22.5
# question 1 is answered by
stock.will_ben_buy_1()
# question 2 is answered by
stock.will_ben_buy_2()
# to obtain the current price
stock.get_price()
# obtain the last 5 years of earning per share
stock.get_esp()
# obtain Graham's recommended maximum stock price
stock.max_price()
# In this case one will be advised not to buy this Stock as it is too expensive.
| index.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Getting Started with Accelerated Computing
#
# In this self-paced, hands-on lab, we will briefly explore some methods for accelerating applications on a GPU.
#
# Lab created by <NAME> (Follow [@CUDAHamster](https://twitter.com/@cudahamster) on Twitter)
# The following timer counts down to a five minute warning before the lab instance shuts down. You should get a pop up at the five minute warning reminding you to save your work!
# <iframe id="timer" src="timer/timer.html" width="100%" height="120px"></iframe>
# ---
# Before we begin, let's verify [WebSockets](http://en.wikipedia.org/wiki/WebSocket) are working on your system. To do this, execute the cell block below by giving it focus (clicking on it with your mouse), and hitting Ctrl-Enter, or pressing the play button in the toolbar above. If all goes well, you should see get some output returned below the grey cell. If not, please consult the [Self-paced Lab Troubleshooting FAQ](https://developer.nvidia.com/self-paced-labs-faq#Troubleshooting) to debug the issue.
print "The answer should be three: " + str(1+2)
# Let's execute the cell below to display information about the GPUs running on the server.
# !nvidia-smi
# ---
# If you have never before taken an IPython Notebook based self-paced lab from NVIDIA, please watch this video. It will explain the infrastructure we are using for this lab, as well as give some tips on it's usage. If you've never taken a lab on this sytem before, its highly recommended you watch this short video first.<br><br>
# <div align="center"><iframe width="640" height="390" src="http://www.youtube.com/embed/ZMrDaLSFqpY" frameborder="0" allowfullscreen></iframe></div>
# ## Introduction to GPU Computing
#
# You may not realize it, but GPUs (GPU is short for Graphics Processing Unit) are good for much more than displaying great graphics in video games. In fact, there is a good chance that your daily life is being affected by GPU-accelerated computing.
#
# GPU-accelerated computing is the use of a graphics processing unit (GPU) together with a CPU to accelerate scientific, engineering, mobile and enterprise applications. Pioneered by NVIDIA, GPUs now power energy-efficient datacenters in government labs, universities, enterprises, and small-and-medium businesses around the world.
#
# ### How Applications Accelerate With GPUs
# GPU-accelerated computing offers unprecedented application performance by offloading compute-intensive portions of the application to the GPU, while the remainder of the code still runs on the CPU. From a user's perspective, applications simply run significantly faster.
#
# 
#
# ### CPU Versus GPU
# A simple way to understand the difference between a CPU and GPU is to compare how they process tasks. A CPU consists of a few cores optimized for sequential serial processing while a GPU consists of thousands of smaller, more efficient cores designed for handling multiple tasks simultaneously.
#
# GPUs have thousands of cores to process parallel workloads efficiently
#
# 
#
# There are hundreds of industry-leading applications already GPU-accelerated. Find out if the applications you use are GPU-accelerated by looking in NVIDIA's [application catalog](http://www.nvidia.com/content/tesla/pdf/gpu-apps-catalog-sept13-digital-fnl-hr.pdf).
#
# ### How to Accelerate Applications
#
# If GPU-acceleration is not already available for your application, you may be interested in developing GPU-accelerated code yourself. There are three main methods methods to achieve GPU-acceleration in your code, and that is what the rest of this lab attempts to demonstrate. The methods are summarized below.
#
# <img src="files/images/three_methods.png" />
#
#
# Enough introduction, let's start the hands-on work!
# ## Libraries
#
# As with any type of computer programming, libraries give access to many different types of functions and algorithms that you do not have to directly implement in your software. Libraries are typically highly-optimized and are accessed through a set of Application Programming Interfaces (APIs). Making use of GPU-accelerated libraries is typically the quickest way to add acceleration to your application. In fact, there are a number of GPU-accelerated libraries that are API compatible with the CPU version. This means you simply change the library you are compiling against - no code changes neccesary!
#
# There is an ever growing number of [libraries available](https://developer.nvidia.com/gpu-accelerated-libraries) for GPU-acclerated computing, both from NVIDIA and 3rd party developers. They range from basic building block libraries, to incredibly complex and dense. You can take full advantage of their capabilities without having to write any of that GPU-accelerated code yourself.
#
# One example of a library that contains GPU-accelerated functions is the open-source computer vision package called [OpenCV](http://opencv.org/). To quote the OpenCV site, "OpenCV was built to provide a common infrastructure for computer vision applications and to accelerate the use of machine perception in the commercial products."
#
# ### Task #1
#
# Your first task in this lab is to compile and run some simple OpenCV code to generate a line drawing of a given image. You'll then see how calling the GPU-accelerated versions of the OpenCV functions results in the same image, but generated in less time.
#
# You are not required to modify any code in this task, but a text editor is present below if you wish to experiment with different values in the code. The source image we are going to work with looks like this:
#
# <img src="files/task1/images/shield.jpg" width=500 />
#
# Let's first run the CPU-only version of this program to see what the output should look like. To do this, execute the following cell block to compile the CPU version. **You execute a cell in this lab by first selecting it with your mouse and then pressing either Ctrl+Enter** (keeps focus on the cell), or Shift+Enter or clicking the play button in the toolbar (moves focus to next cell after execution). Try that now. You should see `Compiled Successfully` printed out if everything works.
# !g++ task1/lines_cpu.cpp -lopencv_core -lopencv_highgui -lopencv_imgproc -o lines_cpu && echo "Compiled Successfully"
# Next we'll execute the compiled program and time how long it takes. Execute the below cell to do this.
#
# **NOTE:** You may notice that the `lines_cpu` program is being executed twice below, but only timed once. This is because the first time this program is run on the system some time is spent loading the OpenCV libraries. By only timing the second run, we remove this load time.
# + language="bash"
# ./lines_cpu
# time ./lines_cpu
# -
# The `real` value should indicate the program took around 2.9 seconds to run. At this point, an output image has been generated and written to the file `out_cpu.png`. Let's use some Python code to display this image in your browser. After executing the cell below, you should see a line drawing of the original image shown above.
from IPython.core.display import Image, display
cpu = Image('out_cpu.png', width=700)
display(cpu)
# Now let's compile and run the GPU version of this program. The compile & run steps have been combined into a single cell which you can execute below.
# + language="bash"
# g++ task1/lines_gpu.cpp -lopencv_core -lopencv_highgui -lopencv_imgproc -lopencv_gpu -o lines_gpu
# ./lines_gpu
# time ./lines_gpu
# -
# By moving the computationally intensive portions of this program to the GPU, we were able to achieve a 5.3x speed-up (from 2.9s to 0.5s), even for this very simple application. This includes the time required to move the image data to GPU memory, process it, and then copy the result back to the CPU's memory in order to write the image to a file.
#
# Use the below cell to confirm the same image was created by the GPU version of the functions.
from IPython.core.display import Image, display
gpu = Image('out_gpu.png', width=800)
display(gpu)
# You can compare the CPU and GPU versions of the application by executing the line below to show the differences. The GPU code will be on the right, and the CPU on the left. Changed lines are marked by a `|` and new lines are marked by a `>` on the GPU side.
# !sdiff task1/lines_cpu.cpp task1/lines_gpu.cpp
# You can see by the above `sdiff` only a few lines of code were added or modified in order to accelerage the functions on the GPU. This really shows the power of using libraries in your code - no need to reinvent the wheel!
#
# Finally, if you wish to modify the code, simply click on the `task1` folder on the left, and select either the `lines_cpu.cpp` or `lines_gpu.cpp` file. If you modify and save either file, you can reuse the corresponding cells above to compile & run the new code.
#
# **Note** You are encouraged to finish the other tasks before coming back and experimenting with this code. This way you are less likely to run out of time before the lab ends.
# <iframe id="task1" src="task1" width="100%" height="400px">
# <p>Your browser does not support iframes.</p>
# </iframe>
# ## Compiler Directives
#
# Now that we've seen how libraries can be used to help accelerate your code, let's move on to a more flexible approach; using compiler directives. Here we will provide hints to the compiler and let it accelerate the code for us. So while this is not quite as easy as using a library, it is more flexible and yet does not require you to modify the underlying source code.
#
# ### Task #2
# Open-specification [OpenACC](http://www.openacc.org/) directives are a straightforward way to accelerate existing Fortran, C and C++ applications. With OpenACC directives, you provide hints via compiler directives (or 'pragmas') to tell the compiler where - and how - it should parallelize compute-intensive code for execution on an accelerator.
#
# If you've done parallel programming using OpenMP, OpenACC is very similar: using directives, applications can be parallelized *incrementally*, with little or no change to the Fortran, C or C++ source. Debugging and code maintenance are easier. OpenACC directives are designed for *portability* across operating systems, host CPUs, and accelerators. You can use OpenACC directives with GPU accelerated libraries, explicit parallel programming languages (e.g., CUDA), MPI, and OpenMP, *all in the same program.*
#
# Watch the following short video introduction to OpenACC:
#
# <div align="center"><iframe width="640" height="390" style="margin: 0 auto;" src="http://www.youtube.com/embed/c9WYCFEt_Uo" frameborder="0" allowfullscreen></iframe></div>
# To demonstrate the power of OpenACC, we're going to look at a very simple Matrix [Transpose](http://en.wikipedia.org/wiki/Transpose) code. This task just involves compiling and running the source to show the differences in performance between an 8-thread [OpenMP](http://openmp.org/wp/) version of the code running on the CPU, and the OpenACC version running on the GPU.
#
# The source code found below is broken up into these functions:
#
# * `referenceTranspose` - the naive transpose function executed in parallel on the CPU using OpenMP
# * `openACCTranspose` - the naive transpose function executed on the massively parallel GPU using OpenACC
# * `time_kernel` - a helper function used to measure the bandwidth of running the `referenceTranpose` function
# * `time_kernel_acc` - a helper function used to measure the bandwidth of running the `openACCTranpose` function
#
# While it's not important to understand all this code, there are a few important take aways.
#
# 1. The OpenACC version of the transpose is compared against the OpenMP version to check for correctness
# 2. In order to get an accurate bandwidth measurement, each version of the transpose is run 500 times and the average is taken from those runs.
# 3. There is no GPU-specific code being used in this example. All acceleration is implemented by the OpenACC [PGI](http://www.pgroup.com/) compiler for you.
#
# Before executing the code, you should look for the following OpenACC directives and see if you can understand their purpose in the code:
#
# * `#pragma acc parallel loop collapse(2) present(in,out)` (line 28) - The `parallel` OpenACC directive tells the compiler that it should offload the code in the structured code block following the `#pragma` (in our case the nested for-loops) following our further instructions and execute it on the GPU. The `loop` tells the compiler to parallelize the next loop. `collapse(2)` says to apply this directive to the next two loops. And finally the `present(in,out)` tells the compiler we've already copied the `in` and `out` data to the device.
# * `#pragma acc data copyin(in[0:rows*cols]) copyout(out[0:rows*cols])` (line 94) - The `data` directive is used to tell the compiler how and when to move data between the CPU (host) memory and the GPU memory. Since we are executing each transpose function 500 times, it doesn't make sense to copy the input and output data across the PCI-Express bus for each iteration as this would severely skew the timing results. This directive says "At the beginning of this pragma, copy the input data to the device. At the end of the structured code block, copy the output data from the device to the host memory."
# * `#pragma acc wait` (line 102) - The `wait` directive tells the compiler that it should wait at this point for all the work on the device to complete. Since the CPU and GPU are two separate processors, they are able to execute code independently. If this `wait` was not there, the timing code would be incorrect as the CPU would not wait for the GPU to finish its work before executing the next line of code.
#
# To look at the code, click on the `task2.c` filename below. If you decide to make changes to the code, make sure to click the Save button in the text editor box (not the tool bar at the top of the browser tab).
# <iframe id="task2" src="task2" width="100%" height="550px">
# <p>Your browser does not support iframes.</p>
# </iframe>
# To compile the task2.c file, simply execute the below cell. Information about the accelerated portions of the code will be printed out, and you can learn more about what these mean by taking the other OpenACC labs available on [nvidia.qwiklab.com](https://nvidia.qwiklab.com/tags/openacc) or the more immersive [OpenACC course](https://developer.nvidia.com/openacc-course).
# !pgcc -o task2_out -acc -Minfo=accel task2/task2.c && echo "Compiled Successfully"
# To run the task after you have successfully compiled, execute the next cell. You should see the GPU is about 3.7x faster than the 8-thread OpenMP verison of the code. Not bad for only adding three `#pragma`'s.
# + language="bash"
# export OMP_NUM_THREADS=8
# ./task2_out
# -
# The high-level flow recommended to take with OpenACC is as follows:
#
# 1. Identify the computationally intensive portions of your code - these are usually good targets for OpenACC. Use any popular CPU profiling tool, the [nvprof tool](http://docs.nvidia.com/cuda/profiler-users-guide/index.html#nvprof-overview) provided in the CUDA toolkit from NVIDIA, and see which functions take up the most amount of time.
# 2. Accelerate the code on the GPU using `kernels` or the `parallel` OpenACC directives. It's very important to verify accuracy of the results at this stage. Don't focus on performance yet.
# 3. Once the code is correctly accelerated, optimize data movement with the various data directives. **This is where you will usually begin to see increases in performance**. Often people get discouraged when they don't see any speedups, or even slowdowns, after step #2. It's important to continue to step #3.
# 4. Perform any additional optimizations if needed and repeat the steps.
# ### Task #3
#
# While compiler directives generally are associated with C, C++ and Fortran, let's see how we can use a similar approach in Python with the `@vectorize` decorator and the [Continuum Analytics](http://continuum.io/) [Numba](http://numba.pydata.org/) compiler.
#
# First let's execute a CPU-only version of a Monte Carlo Options Pricer simulation code. It's not important to understand exactly what this code is doing, only that we get a similar stock price between the two versions. We also want to look at the `time elapsed` value in the text output.
#
# Execute the next cell to load the common code into our namespace. You can download this code at the end of the lab if you wish to look at it in more detail.
# %run -i monte.py
# Execute the following cell to run the CPU version of the `step` function. This should generate text output and a graph in about 15 seconds.
# +
# %matplotlib inline
def step_cpu(prices, dt, c0, c1, noises):
return prices * np.exp(c0 * dt + c1 * noises)
driver(step_cpu, do_plot=True)
# -
# Now, let's accelerate the `step` function on the GPU. To do this, we're going to use a Python decorator. Using the `@vectorize` decorator, numba can compile the `step` function into a [ufunc (universal function)](http://docs.scipy.org/doc/numpy/reference/ufuncs.html) that operates over [NumPy](http://www.numpy.org/) arrays as fast as traditional ufuncs written in C!
#
# `@vectorize` in numba works by running through all of the elements of the input arrays executing the scalar function on each set. This means that our `step_gpu` function needs to be a scalar function - taking scalar inputs and returning a scalar output. To accomplish this, the only thing we have to modify is to use `math.exp` which operates on scalars instead of `np.exp` which expects a NumPy array.
#
# Since a compiler is trying to turn the `step_gpu` function into machine code, we need to provide it with some information about the data types being passed in. That's the first parameter you see being passed to `@vectorize`.
#
# Finally, we are targeting the GPU with this decorator (the second parameter). And that's it! The compiler handles the work of generating the GPU code, performing any data movement required, and launching the work. Go ahead and execute the below cell to see what kind of speed up we get.
# +
from numba import vectorize
import math # needed for the math.exp function
@vectorize(['float64(float64, float64, float64, float64, float64)'], target='cuda')
def step_gpu(prices, dt, c0, c1, noises):
return prices * math.exp(c0 * dt + c1 * noises)
driver(step_gpu, do_plot=True)
# -
# You should see about a 27% increase in speed.
#
# In the interest of transperency, if you change the `target` to `parallel` instead of `cuda`, the compiler will target the multi-core CPU availalbe on this instance and you will get similar performance to what you just got on the GPU. The reason for this is we're only porting a very small amount of computation the GPU, and therefore not hiding the latency of transferring data around. If you decide to take the Python labs on [nvidia.qwiklab.com](https://nvidia.qwiklab.com/), you will see how we can achieve much greater increases in performance of this algorithm by moving more computation to the GPU with both library calls and some CUDA code, and hiding the cost of trasnferring data.
# ## CUDA
#
# Programming for the GPU in a CUDA-enabled language is the most flexible of the three approaches. While CUDA was initially just a C compiler when it was first released, it has grown into the parallel computing platform for accessing the general purpose, massively parallel compute power of an NVIDIA GPU.
#
# There is a growing list of languages that understand how to speak CUDA and target the GPU including but not limited to C, C++, Fortran, R, and Python. In this lab, you will write some CUDA code directly in Python. This code will be compiled using [Continuum Analytics](http://continuum.io/) [Numba](http://numba.pydata.org/numba-doc/0.13.4/CUDAJit.html) compiler which contains CUDA Python support.
#
# ### Task #4
#
# This task does not require any modifications to get working and will be generating the [Mandelbrot Set](http://en.wikipedia.org/wiki/Mandelbrot_set). It is designed to show you the speed-up you can get using CUDA Python to move computational intensive portions of code to the GPU.
#
# Executing the below cell will run the same algorithm on first the GPU and then again on the CPU. Both of these examples are using code compiled from Python using the Numba compiler. The timing of the GPU includes all data transfers between the CPU memory and GPU memory in order to make a fair comparison. While it's not explicitly coded, the Numba compiler is able to recognize and handle the need for the `gimage` data to be tranferred to the GPU before `create_fractal_gpu` is called, and back when it's complete. The `cuda.synchronize` is there to ensure the timing information is accurate.
#
# Feel free to change the `numIters` variable to decrease or increase the number of iterations performed. In addition you can modify the fractal grid points (starting values of `-2.0, 1.0, -1.0, 1.0`) to change the area of the fractal processed. As you increase the number of iterations, you should see the gap in performance between the GPU and CPU increasing as the amount of computation hides the data transfer latency.
#
# You will notice that the GPU version adds `[griddim, blockdim]` before the parameter list. These values control how the parallel work is spread across the GPU and will be described in more detail in the next task. **You should run the next cell twice, the first time may be slower due to the one-time compilation of the `create_fractal_*` functions**
# +
from mymandel import *
numIters = 20
# Run the GPU Version first
gimage = np.zeros((1024, 1536), dtype = np.uint8)
blockdim = (32, 8)
griddim = (32, 16)
with mytimer('Mandelbrot created on GPU'):
create_fractal_gpu[griddim, blockdim](-2.0, 1.0, -1.0, 1.0, gimage, numIters)
cuda.synchronize
imshow(gimage)
show()
# Run the CPU Version last
image = np.zeros_like(gimage)
with mytimer('Mandelbrot created on CPU'):
create_fractal_cpu(-2.0, 1.0, -1.0, 1.0, image, numIters)
imshow(image)
show()
# -
# You should see around a 9x speed increase when moving from the GPU to the CPU when using the original parameters.
#
# If you are interested in seeing the rest of the code used in the above example, please execute the next cell. This is not a requirement for the lab, but you may find it insightful after you perform the next task. In addition, at the end of this lab, you are presented with a section on downloading the code for offline viewing - but be careful you don't run out of time!
# +
# # %load mymandel.py
from contextlib import contextmanager
import time
import numpy as np
from pylab import imshow, show
from numba import *
@autojit
def mandel(x, y, max_iters):
"""
Given the real and imaginary parts of a complex number,
determine if it is a candidate for membership in the Mandelbrot
set given a fixed number of iterations.
"""
c = complex(x, y)
z = 0.0j
for i in xrange(max_iters):
z = z*z + c
if (z.real*z.real + z.imag*z.imag) >= 4:
return i
return max_iters
# The compiled CPU version of the fractal code
@autojit
def create_fractal_cpu(min_x, max_x, min_y, max_y, image, iters):
height = image.shape[0]
width = image.shape[1]
pixel_size_x = (max_x - min_x) / width
pixel_size_y = (max_y - min_y) / height
for x in xrange(width):
real = min_x + x * pixel_size_x
for y in xrange(height):
imag = min_y + y * pixel_size_y
color = mandel(real, imag, iters)
image[y, x] = color
# create a GPU accelerated version of the madel function
#so it can be called from other device functions like mandel_kernel
mandel_gpu = cuda.jit(restype=uint32, argtypes=[f8, f8, uint32], device=True)(mandel)
# The compiled GPU version of the fractal code
@cuda.jit(argtypes=[f8, f8, f8, f8, uint8[:,:], uint32])
def create_fractal_gpu(min_x, max_x, min_y, max_y, image, iters):
height = image.shape[0]
width = image.shape[1]
pixel_size_x = (max_x - min_x) / width
pixel_size_y = (max_y - min_y) / height
startX, startY = cuda.grid(2) # startX = cuda.threadIdx.x + cuda.blockDim.x * cuda.BlockIdx.x
# startX = cuda.threadIdx.x + cuda.blockDim.x * cuda.BlockIdx.x
gridX = cuda.gridDim.x * cuda.blockDim.x; # Number of threads, or size of image in X direction
gridY = cuda.gridDim.y * cuda.blockDim.y; # Number of threads, or size of image in Y direction
for x in xrange(startX, width, gridX):
real = min_x + x * pixel_size_x
for y in xrange(startY, height, gridY):
imag = min_y + y * pixel_size_y
image[y, x] = mandel_gpu(real, imag, iters)
# Used for timing sections of code
@contextmanager
def mytimer(name):
startTime = time.time()
yield
elapsedTime = time.time() - startTime
print('{} in {} ms'.format(name, int(elapsedTime * 1000)))
# -
# ### Task 5 - Hello World
#
# For this task, you get to try your hand and writing some CUDA Python code. We are going to be using the following concepts:
#
# * <code style="color:green">@cuda.autojit</code> - this decorator is used to tell the CUDA compiler that the function is to be compiled for the GPU. With `autojit`, the compiler will try and determine the type information of the variables being passed in. You can create your own signatures manually by using the `jit` decorator.
# * <code style="color:green">cuda.blockIdx.x</code> - this is a read-only variable that is defined for you. It is used within a GPU kernel to determine the ID of the block which is currently executing code. Since there will be many blocks running in parallel, we need this ID to help determine which chunk of data that particular block will work on.
# * <code style="color:green">cuda.threadIdx.x</code> - this is a read-only variable that is defined for you. It is used within a GPU kernel to determine the ID of the thread which is currently executing code in the active block.
# * <code style="color:green">myKernel\[number_of_blocks, threads_per_block\](...)</code> - this is the syntax used to launch a kernel on the GPU. Inside the list (the square brackets `[...]`), the first number is the total number of blocks we want to run on the GPU, and the second is the number of threads there are per block. It's possible, and in fact recommended, for one to schedule more blocks than the GPU can actively run in parallel. The system will just continue executing blocks until they have all completed. The following video addresses grids, blocks, and threads in more detail.
#
# <div align="center"><iframe width="640" height="390" src="http://www.youtube.com/embed/KM-zbhyz9f4" frameborder="0" allowfullscreen></iframe></div>
#
# Let's explore the above concepts by doing a simple "Hello World" example.
#
# Most of the code in this example has already been written for you. Your task is to modify the single line in the `hello` function such that the data printed out at the end looks like:
#
# `[[0 0 0 0 0]]`
#
# What's happening is that all the threads in block 0 are writing the block ID into their respective place in the array. Remember that this function is being run in parallel by the threads in block 0, each with their own unique thread ID. Since we're launching a single block with 5 threads, the following is happening parallel:
#
# data[0,0] = 0
# data[0,1] = 0
# data[0,2] = 0
# data[0,3] = 0
# data[0,4] = 0
#
# If you get stuck, click on the link below the code to see the solution.
# +
from numba import *
import numpy as np
@cuda.jit
def hello(data):
data[ cuda.blockIdx.x , cuda.threadIdx.x ] = cuda.blockIdx.x
numBlocks = 1
threadsPerBlock = 5
data = np.ones((numBlocks, threadsPerBlock), dtype=np.uint8)
hello[numBlocks,threadsPerBlock](data)
print data
# -
# See the solution [below](#Task-#5-Solution)
# Once you have a solution generating the correct output, try increasing the number of blocks by a few and see if you understand the output you get. Be careful about making the number of blocks too big, as it may take a while to print out all those values! In addition, there is a limit on the number of overall threads, the number of blocks, and the number of threads per block you can request.
# ## Learn More
#
# If you are interested in learning more, you can use the following resources:
#
# * More labs are available at [nvidia.qwiklab.com](https://nvidia.qwiklab.com)
# * CUDA/GPU Registered Developers with NVIDIA will periodically receive free Credits for use on nvidia.qwiklab.com. [Sign up today](https://developer.nvidia.com/registered-developer-programs)!
# * Learn more at the [CUDA Developer Zone](https://developer.nvidia.com/category/zone/cuda-zone).
# * If you have an NVIDIA GPU in your system, you can download and install the [CUDA toolkit](https://developer.nvidia.com/cuda-toolkit) which comes packed with lots of sample code to look at. Otherwise you can go to [docs.nvidia.com/cuda](http://docs.nvidia.com/cuda) and explore the samples there.
# * Take the fantastic online and **free** Udacity [Intro to Parallel Programming](https://www.udacity.com/course/cs344) course which uses CUDA C.
# * Search or ask questions on [Stackoverflow](http://stackoverflow.com/questions/tagged/cuda) using the cuda tag
# * NVIDIA provided hands-on training at major conferences such as SuperComputer and its own GPU Technology Conference.
# <a id="post-lab"></a>
# ## Post-Lab
#
# Finally, don't forget to save your work from this lab before time runs out and the instance shuts down!!
#
# 1. Save this IPython Notebook by going to `File -> Download as -> IPython (.ipynb)` at the top of this window. *Please note that the in-browser text editors, and likely the executable cells, will not work if you run this IPython Notebook locally.*
# 2. You can execute the following cell block to create a zip-file of the files you've been working on, and download it with the link below.
# + language="bash"
# rm -f gpu_computing.zip
# zip -r gpu_computing task*/* *.py
# -
# **After** executing the above cell, you should be able to download the zip file [here](files/gpu_computing.zip)
# <a id="FAQ"></a>
# ---
# # Lab FAQ
#
# Q: I'm encountering issues executing the cells, or other technical problems?<br>
# A: Please see [this](https://developer.nvidia.com/self-paced-labs-faq#Troubleshooting) infrastructure FAQ.
#
# Q: I'm getting unexpected behavior (i.e., incorrect output) when running any of the tasks.<br>
# A: It's possible that one or more of the CUDA Runtime API calls are actually returning an error, but the code above is not checking for this (due to the time constraints of this lab). Try checking the return value of any CUDA Runtime API call and see if the value is non-zero indicating an error.
# ## Solutions
#
# ### Task #5 Solution
#
# ```
# from numba import *
# import numpy as np
#
# @cuda.jit
# def hello(data):
# data[cuda.blockIdx.x,cuda.threadIdx.x] = cuda.blockIdx.x
#
# numBlocks = 1
# threadsPerBlock = 5
#
# data = np.ones((numBlocks, threadsPerBlock), dtype=np.uint8)
#
# hello[numBlocks,threadsPerBlock](data)
#
# print data
# ```
#
# [Return](#Task-5---Hello-World)
# <style>
# p.hint_trigger{
# margin-bottom:7px;
# margin-top:-5px;
# background:#64E84D;
# }
# .toggle_container{
# margin-bottom:0px;
# }
# .toggle_container p{
# margin:2px;
# }
# .toggle_container{
# background:#f0f0f0;
# clear: both;
# font-size:100%;
# }
# </style>
# <script>
# $("p.hint_trigger").click(function(){
# $(this).toggleClass("active").next().slideToggle("normal");
# });
#
# $(".toggle_container").hide();
# </script>
| cuda_quiklabs/Accelerated Computing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Audience Splitting in A/B Experiments
# > A tutorial on how to split audience in a deterministic way using hashing.
#
# - toc: false
# - badges: true
# - comments: true
# - categories: [altair, python]
# - image: images/chart-preview.png
# # About
#
# One key element in running a A/B experiment is splitting of audience based on the unit of diversion. Most of the experiment platforms does the splitting of audience for us. But there are situation in which analyst need to run an A/B experiment and splitting of audience need to performed by the analyst. In most of the organizations data is stored in a database and it would be nice if we can perform treatment assignment in SQL . Also, we need the audience split to perform post-hoc analysis of the experiment. In this blog, I will show how to perform audience splitting in spark and Hive using an example.
# ### Data Preparation
# 1. Lets create a spark session in local.
# 2. Lets create a dummy dataset with 100,000 customers along with gender information.
# 3. Add uuid column to the dataframe to uniquely identify a user.
# 4. Convert pandas dataframe to a spark dataframe
# 5. Register the spark dataframe as "user_table" to be accessed in Hive
# +
import pyspark
import altair as alt
import numpy as np
import pandas as pd
import uuid
import scipy.stats as sc
from vega_datasets import data
from pyspark.sql import SparkSession
spark = SparkSession \
.builder \
.enableHiveSupport() \
.getOrCreate()
customers = (pd.DataFrame({'user': np.arange(100000),
'gender':[np.random.choice(['m','f'], p=[0.55,0.45]) for _ in np.arange(100000)]})
.assign(user_uuid=[uuid.uuid4() for _ in range(100000)])
)
customers.head()
# -
sdf=spark.createDataFrame(customers.astype(str))
sdf.createOrReplaceTempView("user_table")
sdf.toPandas().head()
# ### Audience splitting
# > Cool hashing trick to perform audience splitting
#
# 1. Select the unit of diversion key : user_uuid in our case (or the ID field we want to split on).
# 2. And a salt('new_widget' in our example), unique value to identify our experiment.
# 3. Concatenate car_uuid with the salt selected.
# 4. Apply a hashing algorithm like md5 hash to split audience into treatment and control
query="""select
user_uuid,
if(
conv(
substr(
md5(concat(user_uuid, '-','new_widget')),
1, 6),
16,10)/conv('ffffff',16,10) > 0.50, 'treatment', 'control') as treatment
,gender
from user_table
"""
df_audience=spark.sql(query).toPandas()
# ### Validation of assignment
# > Chi-Square test of indepence is our friend
# Lets visualize the split and looks like assignment is 50-50. But how do we validate this with statistically rigor ?
(df_audience
.groupby('treatment')
.agg(users=('user_uuid','count'))
.reset_index()
.assign(percent_users=lambda x:(x['users']/x['users'].sum())*100)
.style.format({'percent_users':'{0:.2f}%'.format})
)
# One way to validate this is see if distribution of gender is random across treatment and control. This can be translated in to a chi square test with the following hypothesis:
#
# **Null Hypothesis H<sub>0</sub>**: Gender is independent of treatment assignment
# **Alternate Hypothesis H<sub>a</sub>**: Gender is not independent of treatment assignment
# Let's run an chi-square test. P-value of 0.14 indicates we can't reject the null hypothesis - gender is independent of the treatment assignment
chi2, p, dof, expected=sc.chi2_contingency(pd.crosstab(df_audience.treatment,
df_audience.gender,
values=df_audience.user_uuid,
aggfunc='count'))
print ("p-value is {}".format(p))
# ### Conclusion
#
# Hashing is very useful technique to assign users to treatment and control in a deterministic way. Using the user_uuid and salt we can get the experiment assignment back. This can also be done easily in any SQL database.
#
| _notebooks/2020-01-15-audience-hashing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Mask R-CNN - Train on Shapes Dataset
#
#
# This trains Mask R-CNN on our pig dataset. You'll need a GPU, though, because the network backbone is a Resnet101, which would be too slow to train on a CPU. On a GPU, you can start to get okay-ish results in a few minutes, and good results in less than an hour.
#
# +
import os
import sys
import random
import math
import re
import time
import numpy as np
import cv2
import matplotlib
import matplotlib.pyplot as plt
import glob
import skimage.io
import pandas as pd
# Root directory of the project is the repo folder,
# needed for external modules etc
ROOT_DIR = os.path.abspath("../../")
PARENT_DIR = os.path.abspath("../../../")
#parent dir is where the repo was cloned, and where /data should be located
DATA_DIR = os.path.join(PARENT_DIR,"data")
TRAIN_DATA_DIR = os.path.join(DATA_DIR,"train")
VAL_DATA_DIR = os.path.join(DATA_DIR,"validation")
ANNOTATION_FILE = "batch1_coco_2019-09-30_1132.json"
annotations_file_path = os.path.join(DATA_DIR,ANNOTATION_FILE)
#PIGS_ONLY_FOLDER = "/data/open_datasets/open_images_v5/images/pigsonly" #pigs from open image v5
# Import Mask RCNN
sys.path.append(ROOT_DIR) # To find local version of the mrcnn library
from mrcnn.config import Config
from mrcnn import utils
import mrcnn.model as modellib
from mrcnn import visualize
from mrcnn.model import log
sys.path.append(os.path.join(ROOT_DIR,"digipig/lbbx2coco"))
import olatools
import json
from PIL import Image, ImageDraw
# %matplotlib inline
# Directory to save logs and trained model
MODEL_DIR = os.path.join(ROOT_DIR, "logs")
# Local path to trained weights file
COCO_MODEL_PATH = os.path.join(ROOT_DIR, "mask_rcnn_coco.h5")
# Download COCO trained weights from Releases if needed
if not os.path.exists(COCO_MODEL_PATH):
print("downloading")
utils.download_trained_weights(COCO_MODEL_PATH)
# -
# ## Configurations
# +
class ShapesConfig(Config):
"""
Edit: adopting example to train on pig images
Configuration for training on the toy shapes dataset.
Derives from the base Config class and overrides values specific
to the toy shapes dataset.
"""
# Give the configuration a recognizable name
NAME = "pigs"
# Train on 1 GPU and 8 images per GPU. We can put multiple images on each
# GPU because the images are small. Batch size is 8 (GPUs * images/GPU).
# seems to ba an issue with more than 1 gpu see issue no 921 in repo
# fix : https://github.com/yoninachmany/spacenet-rio-buildings-mask-rcnn/commit/5a2a68e830b6878a9a06175ac9be4c3bfa22f1bc
GPU_COUNT = 3
IMAGES_PER_GPU = 1
# Number of classes (including background)
# start with pigs only
NUM_CLASSES = 1 + 3 # background + 3 pig_classes
# Use small images for faster training. Set the limits of the small side
# the large side, and that determines the image shape.
IMAGE_MIN_DIM = 128
IMAGE_MAX_DIM = 1280
# Use smaller anchors because our image and objects are small
RPN_ANCHOR_SCALES = (8, 16, 32, 64, 128) # anchor side in pixels // added 256, 512 -> got: IndexError: index 5 is out of bounds for axis 0 with size 5
# Reduce training ROIs per image because the images are small and have
# few objects. Aim to allow ROI sampling to pick 33% positive ROIs.
TRAIN_ROIS_PER_IMAGE = 128 #org sugg val 32
# Use a small epoch since the data is simple
#STEPS_PER_EPOCH = 100
#"If you have a training set of fixed size you can ignore it.." [1]
# use small validation steps since the epoch is small
VALIDATION_STEPS = 5
#similar to steps_per_epoch but on the validation data set instead on the training data. If you have the time to go through your whole validation data set I recommend to skip this parameter. [1]
#ref [1] https://datascience.stackexchange.com/questions/29719/how-to-set-batch-size-steps-per-epoch-and-validation-steps
config = ShapesConfig()
config.display()
# -
# ## Notebook Preferences
def get_ax(rows=1, cols=1, size=8):
"""Return a Matplotlib Axes array to be used in
all visualizations in the notebook. Provide a
central point to control graph sizes.
Change the default size attribute to control the size
of rendered images
"""
_, ax = plt.subplots(rows, cols, figsize=(size*cols, size*rows))
return ax
# ## Dataset
#
#
#
# Extend the Dataset class and add a method to load the shapes dataset, `load_shapes()`, and override the following methods:
#
# * load_image()
# * load_mask()
# * image_reference()
# +
class PigsDataset(utils.Dataset):
def __init__(self, annotations, images_dir, class_map=None):
super().__init__(class_map=class_map)
self.annotations_file = annotations
self.image_folder = images_dir
def load_pigs(self, count, height, width, datadir):
"""
load pig files into dataset object from mrcnn
"""
# Add classes
self.add_class("pigs", 1, "pig_body")
self.add_class("pigs", 2, "pig_head")
self.add_class("pigs", 3, "pig_tail")
# Add images to image list (see parent class)
counter = 0
for file in os.listdir(datadir):
# Todo: check for ".png" extention to exclude irrelevant files e.g. os-generated files
if counter>=count:
break
else: counter = counter + 1
tmp_id = int(olatools.intify_filename(file))
test = self.add_image("pigs", image_id=tmp_id, path=file) #width=width, height=height, bg_color=bg_color, shapes="pig")
#test is an un-used return value for debuging
def load_image(self, image_id):
"""Load the specified image and return a [H,W,3] Numpy array.
"""
datadir = self.image_folder #"/home/blaise/code/grisehale_nmbu/data/train" #omg!!!!
# Load image
image = skimage.io.imread(datadir+"/"+self.image_info[image_id]['path'])
# If grayscale. Convert to RGB for consistency.
if image.ndim != 3:
image = skimage.color.gray2rgb(image)
# If has an alpha channel, remove it for consistency
if image.shape[-1] == 4:
image = image[..., :3]
return image
# def image_reference(self, image_id):
# """Return the shapes data of the image."""
def load_mask(self, image_id):
"""instance masks for shapes of the given image ID.
"""
with open(annotations_file_path, 'r') as rf:
annotations_file_object = json.load(rf)
use_static_dimentions = True
width = 1280
height = 720
count = 0
mask = np.zeros([height, width, count], dtype=np.uint8)
class_ids = np.zeros(0)
#check if id is in map
map_keys = self.image_from_source_map.keys()
if (image_id in self.image_from_source_map.values()):
for annotation in annotations_file_object["annotations"]:
# todo: add check to see if id exists in train folder images
id_x = annotation["image_id"]
#image_id is sequential number of images 0/1 - 582/583
#annotations_file_object contains id on form 54171016201710171313390240015840
#the map function uses this form to map between the two: pigs.54171016201710171313390240015840 maps 0-500
if 'pigs.'+str(annotation["image_id"]) in self.image_from_source_map:
if (image_id == self.image_from_source_map['pigs.'+str(annotation["image_id"])]) :
#find img dimentions (there must be a bettery way to do this)
if not(use_static_dimentions):
for image in annotations_file_object["images"]:
if image["id"]==image_id:
height = image["height"]
width = image["width"]
class_ids= np.append(class_ids,annotation["category_id"])
polygon = annotation["segmentation"][0]
polypoints = [np.array(olatools.get_xy_pairs_vector_from_xyxy_list(polygon))]
img = np.zeros([height, width, 1], np.uint8)
img = cv2.fillPoly(img, polypoints,(255,255,255))
mask = np.append(mask,img,2)
else:
print("did not found id ")
print(self.image_from_source_map)
return mask.astype(np.bool), class_ids.astype(np.int32)
# +
# Training dataset
dataset_train = PigsDataset(annotations_file_path, TRAIN_DATA_DIR)
dataset_train.load_pigs(531, config.IMAGE_SHAPE[0], config.IMAGE_SHAPE[1], TRAIN_DATA_DIR)
dataset_train.prepare()
# Validation dataset
dataset_val = PigsDataset(annotations_file_path, VAL_DATA_DIR)
dataset_val.load_pigs(52, config.IMAGE_SHAPE[0], config.IMAGE_SHAPE[1], VAL_DATA_DIR)
dataset_val.prepare()
# +
# Load and display random samples from validation
image_ids = np.random.choice(dataset_val.image_ids, 5)
for image_id in image_ids:
image = dataset_val.load_image(image_id)
mask, class_ids = dataset_val.load_mask(image_id)
visualize.display_top_masks(image, mask, class_ids, dataset_val.class_names)
# -
# Create model in training mode
model = modellib.MaskRCNN(mode="training", config=config,
model_dir=MODEL_DIR)
# +
# Which weights to start with?
init_with = "coco" # "imagenet", "coco", or "last"
if init_with == "imagenet":
model.load_weights(model.get_imagenet_weights(), by_name=True)
elif init_with == "coco":
# Load weights trained on MS COCO, but skip layers that
# are different due to the different number of classes
# See README for instructions to download the COCO weights
model.load_weights(COCO_MODEL_PATH, by_name=True, exclude=["mrcnn_class_logits", "mrcnn_bbox_fc", "mrcnn_bbox", "mrcnn_mask"])
elif init_with == "last":
# Load the last model you trained and continue training
model.load_weights(model.find_last(), by_name=True)
# -
# ## Training
#
# Train in two stages:
# 1. Only the heads. Here we're freezing all the backbone layers and training only the randomly initialized layers (i.e. the ones that we didn't use pre-trained weights from MS COCO). To train only the head layers, pass `layers='heads'` to the `train()` function.
#
# 2. Fine-tune all layers. For this simple example it's not necessary, but we're including it to show the process. Simply pass `layers="all` to train all layers.
# +
# Train the head branches
# Passing layers="heads" freezes all layers except the head
# layers. You can also pass a regular expression to select
# which layers to train by name pattern.
model.train(dataset_train, dataset_val,
learning_rate=config.LEARNING_RATE,
epochs=1,
layers='heads')
# delete below after training 2020-01-27
# Fine tune all layers
# Passing layers="all" trains all layers. You can also
# pass a regular expression to select which layers to
# train by name pattern.
model.train(dataset_train, dataset_val,
learning_rate=config.LEARNING_RATE / 10,
epochs=2,
layers="all")
#model_path = os.path.join(MODEL_DIR, "mask_rcnn_pig1-t500-bigimg-train_data_only_deleteme.h5")
#model.keras_model.save_weights(model_path)
# -
model_path = os.path.join(MODEL_DIR, "mask_rcnn_pig1-t500-bigimg-train_data_deletethis.h5")
model.keras_model.save_weights(model_path)
# +
model.train(dataset_train, dataset_val,
learning_rate=config.LEARNING_RATE,
epochs=1,
layers='heads')
# Fine tune all layers
# Passing layers="all" trains all layers. You can also
# pass a regular expression to select which layers to
# train by name pattern.
model.train(dataset_train, dataset_val,
learning_rate=config.LEARNING_RATE / 10,
epochs=2,
layers="all")
model_path = os.path.join(MODEL_DIR, "mask_rcnn_pig1-t500-bigimg-train_data_only.h5")
model.keras_model.save_weights(model_path)
# -
# Save weights
# Typically not needed because callbacks save after every epoch
# Uncomment to save manually
model_path = os.path.join(MODEL_DIR, "mask_rcnn_pig1-t500-bigimg.h5")
model.keras_model.save_weights(model_path)
# ## Detection
# +
class InferenceConfig(ShapesConfig):
GPU_COUNT = 1
IMAGES_PER_GPU = 1
inference_config = InferenceConfig()
# Recreate the model in inference mode
model = modellib.MaskRCNN(mode="inference",
config=inference_config,
model_dir=MODEL_DIR)
# Get path to saved weights
# Either set a specific path or find last trained weights
# model_path = os.path.join(MODEL_DIR, "mask_rcnn_pig1-t500-bigimg-train_data_deletethis.h5")
# model_path = model.find_last()
model_path = os.path.join(PARENT_DIR, "mask_rcnn_pig1-t500-bigimg-train_data_only.h5")
# Load trained weights
print("Loading weights from ", model_path)
model.load_weights(model_path, by_name=True)
# +
# Test on a random image
image_id = random.choice(dataset_val.image_ids)
original_image, image_meta, gt_class_id, gt_bbox, gt_mask =\
modellib.load_image_gt(dataset_val, inference_config,
image_id, use_mini_mask=False)
log("original_image", original_image)
log("image_meta", image_meta)
log("gt_class_id", gt_class_id)
log("gt_bbox", gt_bbox)
log("gt_mask", gt_mask)
visualize.display_instances(original_image, gt_bbox, gt_mask, gt_class_id,
dataset_train.class_names, figsize=(8, 8))
# +
results = model.detect([original_image], verbose=1)
r = results[0]
visualize.display_instances(original_image, r['rois'], r['masks'], r['class_ids'],
dataset_val.class_names, r['scores'], ax=get_ax())
# -
# ## Evaluation
# +
# Compute VOC-Style mAP @ IoU=0.5
# Will fail with inputs not countaining expected classes
number_of_eval_images = 3
image_ids = np.random.choice(dataset_val.image_ids, number_of_eval_images)
APs = []
gt_match_df = pd.DataFrame( columns=["gt_class_id","gt_match"])
performance_summary_df = pd.DataFrame( columns=["img","num_gt_pigs","num_gt_heads","num_gt_tails",\
"num_pred_pigs", "num_pred_heads", "num_pred_tails",\
"num_match_pigs", "num_match_heads", "num_match_tails"])
start = time.time()
for image_id in image_ids:
# Load image and ground truth data
image, image_meta, gt_class_id, gt_bbox, gt_mask =\
modellib.load_image_gt(dataset_val, inference_config,
image_id, use_mini_mask=False)
molded_images = np.expand_dims(modellib.mold_image(image, inference_config), 0)
# Run object detection
results = model.detect([image], verbose=0)
r = results[0]
# Compute AP
AP, precisions, recalls, overlaps =\
utils.compute_ap(gt_bbox, gt_class_id, gt_mask,
r["rois"], r["class_ids"], r["scores"], r['masks'])
APs.append(AP)
unique_pred, counts_pred = np.unique(r['class_ids'], return_counts=True)
num_pred_pigs = counts_pred[0]
num_pred_heads = counts_pred[1]
num_pred_tails = counts_pred[2]
unique_gt, counts_gt = np.unique(gt_class_id, return_counts=True)
num_gt_pigs = counts_gt[0]
num_gt_heads = counts_gt[1]
num_gt_tails = counts_gt[2]
gt_match, pred_match, overlaps = utils.compute_matches(
gt_bbox, gt_class_id, gt_mask,
r["rois"], r["class_ids"], r["scores"], r['masks'],
0.5)
tmp_data = pd.DataFrame({"gt_class_id":gt_class_id, "gt_match":gt_match})
gt_match = tmp_data.query("gt_match > -1").groupby("gt_class_id").count()
num_match_pigs = gt_match.query("gt_class_id == 1").values[0][0]
num_match_heads = gt_match.query("gt_class_id == 2").values[0][0]
num_match_tails = gt_match.query("gt_class_id == 3").values[0][0]
tmp_summary = pd.DataFrame({
"img":image_id,
"num_gt_pigs": num_gt_pigs,
"num_gt_heads": num_gt_heads,
"num_gt_tails": num_gt_tails,
"num_pred_pigs": num_pred_pigs,
"num_pred_heads": num_pred_heads,
"num_pred_tails": num_pred_tails,
"num_match_pigs": num_match_pigs,
"num_match_heads": num_match_heads,
"num_match_tails": num_match_tails,},index=[image_id])
performance_summary_df = performance_summary_df.append(tmp_summary) #ignore_index = True)
gt_match_df = gt_match_df.append(tmp_data, ignore_index = True)
if num_gt_pigs<num_match_pigs or num_gt_heads<num_match_heads or num_gt_tails<num_match_tails:
print('inconsistent number of matches: more matches than ground truths')
end = time.time()
print(f"elapsed time for {number_of_eval_images} images: {end - start} equals {(end - start)/number_of_eval_images} time per image")
# -
performance_summary_df
performance_summary_df.to_csv("performance_summary_evaluation.csv")
# +
gt_pigs = performance_summary_df['num_gt_pigs'].sum()
gt_heads = performance_summary_df['num_gt_heads'].sum()
gt_tails = performance_summary_df['num_gt_tails'].sum()
pred_pigs = performance_summary_df['num_pred_pigs'].sum()
pred_heads = performance_summary_df['num_pred_heads'].sum()
pred_tails = performance_summary_df['num_pred_tails'].sum()
tp_pigs = performance_summary_df['num_match_pigs'].sum()
tp_heads = performance_summary_df['num_match_heads'].sum()
tp_tails = performance_summary_df['num_match_tails'].sum()
fp_pigs = pred_pigs - tp_pigs
fp_heads = pred_heads - tp_heads
fp_tails = pred_tails - tp_tails
fn_pigs = gt_pigs - tp_pigs
fn_heads = gt_heads - tp_heads
fn_tails = gt_tails - tp_tails
#precision = tp / (tp+fp)
pigs_precision = tp_pigs / (tp_pigs + fp_pigs)
heads_precision = tp_heads / (tp_heads + fp_heads)
tails_precision = tp_tails / (tp_tails + fp_tails)
#recall = tp / (fn+tp)
pigs_recall = tp_pigs / (fn_pigs + tp_pigs)
heads_recall = tp_heads / (fn_heads + tp_heads)
tails_recall = tp_tails / (fn_tails + tp_tails)
# -
print(f" pigs precision: {pigs_precision} \n pigs recall: {pigs_recall} \n")
print(f" heads precision: {heads_precision} \n heads recall: {heads_recall} \n")
print(f" tails precision: {tails_precision} \n tails recall: {tails_recall} \n")
print(f"pigs: \ntp: {tp_pigs}, \nfp: {fp_pigs}, \nfn: {fn_pigs} \n")
print(f"heads: \ntp: {tp_heads}, \nfp: {fp_heads}, \nfn: {fn_heads} \n")
print(f"tails: \ntp: {tp_tails}, \nfp: {fp_tails}, \nfn: {fn_tails} \n")
| digipig/pigs_shapes/01_train_evaluate_pigs_heads_tails_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.10 64-bit (''Braket'': conda)'
# name: python3
# ---
# # Simulating noise on Amazon Braket
#
# This notebook gives a detailed overview of noise simulations on Amazon Braket. Amazon Braket provides two noise simulators: a local noise simulator that you can use for free as part of the Braket SDK and a fully managed, high-performing noise simulator, DM1. Both simulators are based on the density matrix formalism. After this tutorial, you will be able to define noise channels, apply noise to new or existing circuits, and run those circuits on the Braket noise simulators.
#
# ### Table of contents:
# * [Background](#Background)
# * [Noise simulation based on the density matrix formalism](#density_matrix)
# * [Quantum channel and Kraus representation](#quantum_channel)
# * [General imports](#imports)
# * [Quick start](#start)
# * [Defining noise channels](#noise_channels)
# * [Pre-defined noise channels](#pre-defined)
# * [Defining custom noise channels](#self-defined)
# * [Adding noise to a circuit](#apply_noise)
# * [Build noisy circuits bottom-up](#apply_noise_directly)
# * [Applying noise to existing circuits with global methods](#apply_noise_globally)
# * [Applying gate noise to the circuit](#gate-noise)
# * [Applying initialization noise to the circuit](#initialization-noise)
# * [Applying readout noise to the circuit](#readout-noise)
# * [Using both the direct and global methods to apply noise](#both)
# * [Running a noisy circuit](#run)
# ## Background <a class="anchor" id="Background"></a>
#
# ### Noise simulation based on the density matrix formalism <a class="anchor" id="density_matrix"></a>
# In an ideal case, a quantum state prepared by a noise-free circuit can be described by a state vector $|\psi\rangle$ -- we call it a 'pure state'. However, the presence of noise in realistic quantum devices will introduce classical uncertainty to the quantum state. For example, a bit flip error with 50% probability acting on a qubit flips the $|0\rangle$ state into either $|0\rangle$ or $|1\rangle$ with a 50-50 chance. Note that this is different from an Hadamard-gate acting on $|0\rangle$: The latter results in a coherent superposition of $|0\rangle$ and $|1\rangle$, whereas the former is a classical, so-called mixture of $|0\rangle$ and $|1\rangle$. The most general way of describing a quantum state in the presence of noise is through the so-called density matrix: $\rho = \sum_i p_i|\psi_i\rangle\langle\psi_i|$. It can be understood as a classical mixture of a series of pure states $|\psi_i\rangle$ (each of which could be highly entangled), where $p_i$ is the probability of the state being in $|\psi_i\rangle$. Because the $p_i$ are classical probabilities they have to sum up to 1: $\sum_i p_i = 1$. The density matrix of a pure state is simply $\rho = |\psi\rangle\langle\psi|$ and, in the bit-flip example from above, the density matrix would be $\rho = 0.5|0\rangle\langle 0| + 0.5|1\rangle\langle 1|$.
#
# The density matrix formalism is a very useful way to describe a noisy system with probabilistic outcomes. It gives an exact description of a quantum system going through a quantum channel with noise. Besides, the expectation value of an observable $\langle O\rangle$ can be easily calculated by $\rm{Tr}(O\rho)$, where "$\rm{Tr}$" is the trace operator.
#
# ### Quantum channel and Kraus representation <a class="anchor" id="quantum_channel"></a>
#
# A [quantum channel](https://en.wikipedia.org/wiki/Quantum_channel) describes the time evolution of a quantum state which is expressed as a density matrix. For instance, to understand what a series of noisy gates does to the state of a quantum computer, you can apply a quantum channel corresponding to the different gate and noise operations.
# Mathematically speaking, a quantum channel is a completely positive and trace-preserving (CPTP) linear map acting on a density matrix. Completely positive means the channel maps positive operators into positive operators (even if the operator is applied to part of a larger system) to make sure the density matrix describes a proper quantum state after the map. Trace-preserving means the trace of the density matrix remains unchanged during the mapping process (this is so that after the map the classical probabilities $p_i$ still sum to 1).
#
# The so-called _Kraus representation_ is a commonly used representation for CPTP maps. [Kraus's theorem](https://en.wikipedia.org/wiki/Quantum_operation#Kraus_operators) states that any quantum operation acting on a quantum state $\rho$ can be expressed as a map $\varepsilon(\rho) = \sum_i K_i\rho K_i^{\dagger}$, and it satisfies: $\sum_i K_i^{\dagger}K_i = \mathbb{1}$, where $\mathbb{1}$ is the Identity operator.
#
# Let's get started and have a look how you can define and simulate noisy circuits on Amazon Braket.
# ## General imports <a class="anchor" id="imports"></a>
# Let's begin with the usual imports and setting our s3 location where we want to persist results.
from braket.circuits import Circuit, Observable, Gate, Noise
from braket.devices import LocalSimulator
from braket.aws import AwsDevice
import numpy as np
from scipy.stats import unitary_group
# <div class="alert alert-block alert-info">
# <b>Note</b> Enter your desired S3 location (bucket and prefix). Remember that bucket names for Amazon Braket always begin with "amazon-braket-".
# </div>
#
# enter the S3 bucket you created during onboarding (or any other bucket starting with "amazon-braket-")
my_bucket = "amazon-braket-Your-Bucket-Name" # the name of the bucket
my_prefix = "Your-Folder-Name" # the name of the folder in the bucket
s3_folder = (my_bucket, my_prefix)
# ## Quick start <a class="anchor" id="start"></a>
#
# Let's start with a simple example of running a noisy circuit on Amazon Braket.
# +
# build a simple circuit
circ = Circuit().h(0).cnot(0,1)
# define a noise channel
noise = Noise.BitFlip(probability=0.1)
# add noise to every gate in the circuit
circ.apply_gate_noise(noise)
# select the local noise simulator
device = LocalSimulator('braket_dm')
# run the circuit on the local simulator
task = device.run(circ, shots = 1000)
# visualize the results
result = task.result()
measurment = result.measurement_counts
print('measurement results:', measurment)
# -
# Ideally, in the noise-free case, the circuit we defined prepares a Bell-state, and we would expect to measure only '00' and '11' outcomes. However, the presence of noise, in our case a bit flip error, means that sometimes we find the state in '01' and '10' instead.
#
# The local simulator is suitable for fast prototyping on small circuits. If you want to run a noisy circuit with more than 10~12 qubits, we recommend using the managed simulator DM1. Using DM1, you can run circuits with up to 17 qubits, and benefit from parallel execution for a group of circuits. The code below shows an example of preparing a 13-qubit GHZ state in the presence of noise.
# +
def ghz_circuit(n_qubits: int) -> Circuit:
"""
Function to return simple GHZ circuit ansatz. Assumes all qubits in range(0, n_qubits-1)
are entangled.
"""
circuit = Circuit().h(0)
for ii in range(0, n_qubits-1):
circuit.cnot(control=ii, target=ii+1)
return circuit
# build a 13-qubit GHZ circuit
circ = ghz_circuit(13)
# define a noise channel
noise = Noise.Depolarizing(probability=0.1)
# add noise to every gate in the circuit
circ.apply_gate_noise(noise)
# select the managed density matrix simulator DM1
device = AwsDevice("arn:aws:braket:::device/quantum-simulator/amazon/dm1")
# run the circuit on DM1
task = device.run(circ, s3_folder, shots = 10)
# visualize the results
result = task.result()
measurment = result.measurement_counts
print('measurement results:', measurment)
# -
# We now start exploring the detailed instructions and use cases of each step in the following sections.
# ## Defining noise channels <a class="anchor" id="noise_channels"></a>
#
# To apply noise to a quantum circuit, first, you need to define the noise channel, which is defined in Kraus representation. We offer many commonly-used noise channels in the `Noise` class of the [Amazon Braket SDK](https://amazon-braket-sdk-python.readthedocs.io/en/latest/_apidoc/braket.circuits.html). In addition, you can also define your own custom noise channel as a list of Kraus operators.
#
# ### Pre-defined noise channels <a class="anchor" id="pre-defined"></a>
#
# The pre-defined single-qubit noise channels include `BitFlip`, `PhaseFlip`, `Depolarizing`, `AmplitudeDamping`, `GeneralizedAmplitudeDamping`, `PhaseDamping` and `PauliChannel`.
# The pre-defined two-qubit noise channels include `TwoQubitDepolarizing` and `TwoQubitDephasing`. The Kraus representations for all of the pre-defined channels are summarized in the following table.
#
# __single-qubit noise channels__
#
# | Noise channel | <div style="width:290px">Kraus representation</div> | Parameter |
# |:-------------- |:-------------------------------------------------- |:------------|
# | `BitFlip` | $(1-p)\rho$ + $pX\rho X$| $p$ is the probability of the bit flip noise. |
# | `PhaseFlip` | $(1-p)\rho$ + $pZ\rho Z$| $p$ is the probability of the phase flip noise. |
# | `Depolarizing` |$(1-p)\rho$ + $p/3(X\rho X$ + $Y\rho Y$ + $Z\rho Z)$|$p$ is the probability of the depolarizing noise (the three possible error cases share the same probability of $p/3$).|
# |`AmplitudeDamping`|$K_0\rho K_0^\dagger$ + $K_1\rho K_1^\dagger$|$K_0=[1,0;0,\sqrt{1-\gamma}]$, $K_1=[0,\sqrt{\gamma};0,0]$, where $\gamma$ is the rate of amplitude damping.|
# |`GeneralizedAmplitudeDamping`|$K_0\rho K_0^\dagger$ + $K_1\rho K_1^\dagger$ + $K_2\rho K_2^\dagger$ + $K_3 \rho K_3^\dagger$|$K_0=\sqrt{p}[1,0;0,\sqrt{1-\gamma}]$, $K_1=\sqrt{p}[0,\sqrt{\gamma};0,0]$, $K_2=\sqrt{1-p}[\sqrt{1-\gamma},0;0,1]$, $K_3=\sqrt{1-p}[0,0;\sqrt{\gamma},0]$, where $\gamma$ is the rate of amplitude damping, and $p$ is the probability of the system been excited by the environment [1].|
# |`PhaseDamping`|$K_0\rho K_0^\dagger$ + $K_1 \rho K_1^\dagger$|$K_0=[1,0;0,\sqrt{1-\gamma}]$, $K_1=[0,0;0,\sqrt{\gamma}]$, where $\gamma$ is the rate of phase damping.|
# |`PauliChannel`|$(1-p_x-p_y-p_z)\rho$ + $p_xX\rho X$ + $p_yY\rho Y$ + $p_zZ\rho Z$|$p_x$, $p_y$ and $p_z$ are probabilities for the Pauli X, Y, Z noise respectively.|
#
#
# __two-qubit noise channels__
#
# |<div style="width:160px">Noise channel</div>| <div style="width:290px">Kraus representation</div> | Parameter |
# |:----------------------- |:-------------------------------------------------- |:------------|
# | `TwoQubitDepolarizing`| $(1-p)\rho$ + $p/15(IX\rho IX$ + $IY\rho IY$ + $IZ\rho IZ$ + $XI\rho XI$ +....+ $ZZ\rho ZZ)$| $p$ is the probability of the two-qubit depolarizing noise (the 15 possible error combinations share the same probability of $p/15$).|
# | `TwoQubitDephasing` | $(1-p)\rho$ + $p/3(IZ\rho IZ$ + $ZI\rho ZI$ + $ZZ\rho ZZ)$| $p$ is the probability of the two-qubit dephasing noise (the three possible error combinations share the same probability of $p/3$). |
# The following code block takes the example of the bit flip noise channel: $\rho\rightarrow(1-p)\rho$ + $pX\rho X$, where $p$ corresponds to the `probability` parameter when defining the noise. This noise channel is equivalent to applying a bit flip error (applying an X gate) with probability $p$ and doing nothing with probability $1-p$. You can check the target qubit count and the Kraus operators of the noise channel defined.
# +
# define a bit flip noise channel with probability = 0.1
noise = Noise.BitFlip(probability=0.1)
print('name: ', noise.name)
print('qubit count: ', noise.qubit_count)
print('Kraus operators: ')
for matrix in noise.to_matrix():
print(matrix, '\n')
# -
# Other pre-defined noise channels can be used in a similar way:
# define a phase flip noise channel
noise = Noise.PhaseFlip(probability=0.1)
# define a single-qubit depolarizing noise channel
noise = Noise.Depolarizing(probability=0.1)
# define a two-qubit depolarizing noise channel
noise = Noise.TwoQubitDepolarizing(probability=0.1)
# define a two-qubit dephasing noise channel
noise = Noise.TwoQubitDephasing(probability=0.1)
# define an amplitude damping noise channel
noise = Noise.AmplitudeDamping(gamma=0.1)
# define a generalized amplitude damping noise, where gamma is the amplitude damping rate, and
# probability is the probability of the system being excited by the environment.
noise = Noise.GeneralizedAmplitudeDamping(gamma=0.1, probability=0.1)
# define a phase damping noise channel
noise = Noise.PhaseDamping(gamma=0.1)
# define a Pauli noise channel
noise = Noise.PauliChannel(probX=0.1, probY=0.2, probZ=0.3)
# ### Defining custom noise channels <a class="anchor" id="self-defined"></a>
# Apart from the pre-defined noise models, you can also define your own noise model by specifying a list of Kraus operators. The following code shows an example of defining a two-qubit Kraus channel with randomly generated unitary operators.
# +
# creat an arbitrary 2-qubit Kraus matrix
E0 = unitary_group.rvs(4) * np.sqrt(0.2)
E1 = unitary_group.rvs(4) * np.sqrt(0.8)
K = [E0, E1]
# define a two-qubit noise channel with Kraus operators
noise = Noise.Kraus(K)
# -
# Note that the noise channel you define needs to form a CPTP map. If the input matrices do not define a CPTP map, an error will be raised.
# +
K_invalid = [np.random.randn(2,2), np.random.randn(2,2)]
try:
noise = Noise.Kraus(K_invalid)
pass
except ValueError as err:
print(err)
# -
# ## Adding noise to a circuit <a class="anchor" id="apply_noise"></a>
#
# There are two methods to build a 'noisy' circuit. First, you can add noise to the circuit 'bottom-up', by using the noise operations in the same way as you would add a gate to the circuit. Second, you can use the methods `apply_gate_noise()`, `apply_initialization_noise()` and `apply_readout_noise()` to apply gate error, qubit initialization error and measurement error globally to existing circuits.
#
# The direct method is more flexible as you can apply noise to any place in a circuit. But for an existing large circuit with lots of gates, you may want to use the global methods to conveniently apply noise to the circuit.
#
#
# ### Build noisy circuits bottom-up <a class="anchor" id="apply_noise_directly"></a>
# Noise channels can be applied to the circuit the same way as gates. The following example shows how to apply single- and two-qubit noise channels directly to a circuit. The noise applied can be visualized in the circuit diagram with the `print()` method.
# apply depolarizing noise
circ = Circuit().x(0).x(1).cnot(0,1).depolarizing(1, probability=0.2).x(0).two_qubit_dephasing(target1=0, target2=1, probability=0.1)
print(circ)
# ### Applying noise to existing circuits with global methods<a class="anchor" id="apply_noise_globally"></a>
#
# We offer three methods to apply noise globally to the circuit: `apply_gate_noise()`, `apply_initialization_noise()` and `apply_readout_noise()`. In the following, we explain in detail the usage of these three methods.
#
# #### Applying gate noise to the circuit <a class="anchor" id="gate-noise"></a>
#
# `apply_gate_noise()` is the method to conveniently apply gate-noise to the circuit. It accepts the following input parameters:
#
# - __noise__: A single or a list of noise channel in `Noise` type.
# - __target_unitary__: A single unitary gate in the form of a matrix in `numpy.ndarray` type. The noise will be applied to that unitary gate.
# - __target_gates__: A single or a list of gates in `Gate` type. Note that `target_gates` and `target_unitary` can not be provided at the same time. If none of `target_gates` and `target_unitary` is given, noise will be applied to all the gates in the circuit.
# - __target_qubits__: A single or a list of qubit indexes. If not given, noise will be applied to all the qubits in the circuit.
#
# When calling the method, the noise channel(s) will be applied right after all `target_gates` in `target_qubits`.
# <div class="alert alert-block alert-info">
# <b>Note</b> When you call this method, noise will be inserted right after the gate. If you like to apply more than one noise operation, be aware of the order. Alternatively, you can provide a list of noise operations in one call, and the noise will be applied in forward order.
# </div>
# The code below is an example of applying phase damping noise to all gates in the circuit.
# +
noise = Noise.PhaseDamping(gamma=0.1)
# the noise channel is applied to every gate in the circuit
circ = Circuit().x(0).bit_flip(0,0.1).cnot(0,1)
circ.apply_gate_noise(noise)
print('Noise is applied to every gate in the circuit:\n')
print(circ)
# -
# If you want to apply noise to some particular gates in the circuit, you can specify them as `target_gates`. Below is an example in which noise is applied to all X gates in the circuit.
# <div class="alert alert-block alert-info">
# <b>Note</b> The <code>target_gates</code> must be a <code>Gate</code> type. You can find all available gates with the following commands:
#
# <code>
# from braket.circuits import Gate
# gate_set = [attr for attr in dir(Gate) if attr[0] in string.ascii_uppercase]
# print(gate_set)
# </code>
# </div>
# the noise channel is applied to all the X gates in the circuit
circ = Circuit().x(0).y(1).cnot(0,2).x(1).z(2)
circ.apply_gate_noise(noise, target_gates = Gate.X)
print('Noise is applied to every X gate:\n')
print(circ)
# If you define custom unitary gates as part of your circuit, and you want to apply noise to them, you can use the `target_unitary` criterion.
U1=unitary_group.rvs(4)
U2=unitary_group.rvs(4)
circ = Circuit().x(0).y(1).unitary((0,1),U1).cnot(0,2).x(1).z(2).unitary((1,2),U2)
circ.apply_gate_noise(noise, target_unitary = U2)
print('Noise is applied to U2:\n')
print(circ)
# If you want to apply noise to some particular qubits in the circuit, you can specify them as `target_qubits`. Below is an example to apply noise to all gates in qubits 0 and 2 in the circuit.
# the noise channel is applied to every gate on qubits 0 and 2
circ = Circuit().x(0).y(1).cnot(0,2).x(1).z(2)
circ.apply_gate_noise(noise, target_qubits = [0,2])
print('Noise is applied to every gate in qubits 0 and 2:\n')
print(circ)
# The `target_qubits` and `target_gates` criteria can be used at the same time. The code block below applies the gate noise to all X gates in qubit 0.
# the noise channel is applied to X gate on qubits 0
circ = Circuit().x(0).y(1).cnot(0,2).x(0).x(1).z(2)
circ.apply_gate_noise(noise, target_gates = Gate.X, target_qubits = 0)
print('Noise is applied to X gates in qubits 0:\n')
print(circ)
# If a list of noise channels is provided, the first noise channel in the list will be applied first, then the second.
# +
# define two noise channels
noise1 = Noise.Depolarizing(probability=0.1)
noise2 = Noise.BitFlip(probability=0.2)
# apply a list of noise channels
circ = Circuit().x(0).y(1).cnot(0,2).x(1).z(2)
circ.apply_gate_noise([noise1, noise2], target_qubits = [0,1])
print('Noise channels are applied to every gate in qubits 0 and 1:\n')
print(circ)
# -
# If you want to apply multi-qubit noise channels to a gate, the number of qubits associated with the gate must equal to the number of qubits defined by the noise channel, or otherwise the noise will not be applied. Below shows an example.
# +
# define a two-qubit noise channel
noise = Noise.TwoQubitDephasing(probability=0.1)
# apply the noise to the circuit
circ = Circuit().x(0).y(1).cnot(0,2).x(1).z(2).swap(1,0)
circ.apply_gate_noise(noise)
print('The two-qubit noise channel is applied to all the two-qubit gates in the circuit:\n')
print(circ)
# -
# #### Applying initialization noise to the circuit <a class="anchor" id="initialization-noise"></a>
#
# `apply_initialization_noise()` is the method to apply initialization noise to the circuit. By using the method, the noise will be applied to every qubit at the beginning of a circuit. It accepts the following input parameters:
#
# - __noise__: a single or a list of noise channel in `Noise` type.
# - __target_qubits__: a single or a list of qubit indexes. If not given, noise will be applied to all the qubits in the circuit.
#
# If you want to apply the initialization noise to an empty circuit, you need to provide `target_qubits` to the method.
# <div class="alert alert-block alert-info">
# <b>Note</b> When you call this method, noise will be inserted at the very beginning of the circuit. If you like to apply more than one noise operation, be aware of the order. Alternatively, you can provide a list of noise operations in one call, and the noise will be applied in forward order.
# </div>
# +
# define a noise channel
noise = Noise.Depolarizing(probability=0.1)
# the noise channel is applied as the initialization noise to the circuit
circ = Circuit().x(0).y(1).cnot(0,2).x(1).z(2)
circ.apply_initialization_noise(noise)
print('Initialization noise is applied to the circuit:\n')
print(circ)
# -
# If you want to apply a multi-qubit noise channel as the initialization noise to a circuit and if the number of the qubits in the existing circuit doesn't match the number of qubits as defined by the noise channel, you need to provide `target_qubits` with the number of qubits matching the noise channel.
# +
# define a two-qubit noise channel
noise = Noise.TwoQubitDephasing(probability=0.1)
# the noise channel is applied as the initialization noise to the circuit
circ = Circuit().x(0).y(1).cnot(0,1).x(1).z(0)
circ.apply_initialization_noise(noise)
print('Initialization noise is applied to the circuit:\n')
print(circ)
# -
# #### Applying readout noise to the circuit <a class="anchor" id="readout-noise"></a>
#
# The method of `apply_readout_noise()` is very similar to the method to apply initialization noise, except that the noise channel is applied to every qubit in the end of a circuit. It accepts the following input parameters:
#
# - __noise__: a single or a list of noise channel in `Noise` type.
# - __target_qubits__: a single or a list of qubit indexes. If not given, noise will be applied to all the qubits in the circuit.
#
# If you want to apply the readout noise to an empty circuit, you need to provide `target_qubits` to the method.
# <div class="alert alert-block alert-info">
# <b>Note</b> When you call this method, noise will be inserted at the very end of the circuit. If you like to apply more than one noise operation, be aware of the order. You can also provide a list of noise operations in the one call, and the noise will be applied in forward order.
# </div>
# +
# define a noise channel
noise = Noise.Depolarizing(probability=0.1)
# the noise channel is applied as the readout noise to the circuit
circ = Circuit().x(0).y(1).cnot(0,2).x(1).z(2)
circ.apply_readout_noise(noise)
print('Read-out noise is applied to the circuit:\n')
print(circ)
# -
# If you want to apply a multi-qubit noise channel as the readout noise to a circuit and if the number of the qubits in the existing circuit doesn't match the number of qubits as defined by the noise channel, you need to provide `target_qubits` with the number of qubits matching the noise channel.
# ### Using both the direct and global methods to apply noise <a class="anchor" id="both"></a>
# You can apply noise to the circuit using both the direct and global methods.
# +
# define a noise channel
noise = Noise.PhaseFlip(probability=0.2)
# create a circuit and add noise directly to the circuit
circ = Circuit().x(0).y(1).bit_flip(0,0.1).cnot(1,2).two_qubit_depolarizing(1, 2, probability=0.1).z(2)
circ.apply_gate_noise(noise, target_qubits=0)
print('Noise channels are applied to the circuit:\n')
print(circ)
# -
# ## Running a noisy circuit <a class="anchor" id="run"></a>
#
# Running a noisy circuit is like running any other task on Amazon Braket. In the example below we will pick the local simulator to run our circuit.
#
# With shots = 0, you can obtain the exact values of probability, density matrix and expectation values of the mixed state by attaching the corresponding result type. The reduced density matrix is also available if providing the targets qubits. If no target qubit is provided, the full density matrix will be returned.
#
# An example is shown in the code block below.
# +
# define the noise channel
noise = Noise.AmplitudeDamping(gamma=0.1)
# create a circuit
circ = Circuit().x(0).y(1).cnot(0,2).x(1).z(2)
# apply the noise to qubits 0 and 2 in the circuit
circ.apply_gate_noise(noise, target_qubits = [0,2])
# attach the result types
circ.probability()
circ.expectation(observable = Observable.Z(),target=0)
# attach the density matrix with target=[0,1], and the reduced density matrix of qubits 0,1 will be returned
circ.density_matrix(target=[0,1])
print(circ)
# choose the noise simulator, which is called "braket_dm"
device = LocalSimulator("braket_dm")
# run the circuit
task = device.run(circ, shots=0)
result = task.result()
print('- Probability is: ')
print(result.values[0])
print('- Expectation value <Z_0> is: ')
print(result.values[1])
print('- The reduced Density Matrix is: ')
print(result.values[2])
# -
# With shots > 0, the results are sampled from the probability distributions. The result type `density_matrix` is not available for shots > 0.
#
# The code below shows the expectation value $\langle Z_0\rangle$ and the probability that the mixed state collapsing into different states. We see those values here are different from the exact values obtained in the shots = 0 case.
# +
# create a circuit
circ = Circuit().x(0).y(1).cnot(0,2).x(1).z(2)
circ.apply_gate_noise(noise, target_qubits = [0,2])
circ.probability()
circ.expectation(observable = Observable.Z(),target=0)
print(circ)
# run the circuit
task = device.run(circ, shots=100)
result = task.result()
print('- Probability is: ')
print(result.values[0])
print('- Expectation value <Z_0> is: ')
print(result.values[1])
# -
# ## Reference
# [1] <NAME>, <NAME>. "Squeezed generalized amplitude damping channel", Physical Review A, 2008, 77(1): 012318.
| examples/braket_features/Simulating_Noise_On_Amazon_Braket.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# +
from BA_kernel_function import polynomial_kernel
from BA_kernel_function import linear_kernel
from BA_kernel_function import rbf_kernel
import numpy as np
def kernel_matrix(X, kernel, coef0=1.0, degree=3, gamma=0.1):
X = np.array(X,dtype=np.float64)
mat = []
for i in X:
row = []
for j in X:
if kernel=='poly':
row.append(polynomial_kernel(i,j,coef0,degree))
elif kernel=='linear':
row.append(linear_kernel(i,j,coef0))
elif kernel=='rbf':
row.append(rbf_kernel(i,j,gamma))
else:
row.append(np.dot(i,j))
mat.append(row)
return mat
| 02 Kernel-based Learning/Tutorial 05 - Support Vector Regression/BA_gram.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/snehalsandy/examples/blob/master/Untitled26.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="qoQiY0lWLwob"
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
import pandas_profiling as pp
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.decomposition import PCA
from sklearn.metrics import roc_auc_score
from sklearn.metrics import roc_curve
from sklearn.metrics import f1_score, precision_score, recall_score
# + id="_gI8WekOQGgs"
df=pd.read_csv("/content/sample_data/musk_csv.csv")
# + colab={"base_uri": "https://localhost:8080/", "height": 247} id="3nsD1shRQPPx" outputId="2633b803-ed56-4dba-917d-2440ac6e245c"
df.head()
# + colab={"base_uri": "https://localhost:8080/"} id="s59s-kcSQTeT" outputId="1cd5056c-29ab-42b3-9a8c-399c549e3c80"
df.shape
# + colab={"base_uri": "https://localhost:8080/"} id="lwa8LemxQVja" outputId="47c8bd72-422f-404b-8340-88fad642eeb9"
df.isna().sum()
# + id="E5wMua-oQ42B"
corr_matrix = df.corr().abs()
# + id="QIFuzB64RfsD"
upper = corr_matrix.where(np.triu(np.ones(corr_matrix.shape), k=1).astype(np.bool))
to_drop = [column for column in upper.columns if any(upper[column] > 0.92)]
# + id="FBbl5a__RmDk"
dd = df.drop(columns = to_drop)
# + colab={"base_uri": "https://localhost:8080/", "height": 514} id="054ygFGiRt-9" outputId="f89d618c-a92a-4226-9c24-57ac3d59861a"
dd
# + colab={"base_uri": "https://localhost:8080/"} id="Edf2pa3cRyG1" outputId="8d746c24-d5dc-4186-a0c3-e161c32de6b5"
dd.shape
# + colab={"base_uri": "https://localhost:8080/"} id="TXtZbOVCR5TO" outputId="eff372d8-5a13-4ba2-9173-5520093eff25"
train,test = train_test_split(dd, random_state=30, test_size = 0.2)
Xtrain = train.iloc[:,3:-1]
Ytrain = train.iloc[:,-1:]
Xtest = test.iloc[:,3:-1]
Ytest = test.iloc[:,-1:]
Xtrain.shape
# + id="408asfbxSIKt"
import tensorflow as tf
# + colab={"base_uri": "https://localhost:8080/"} id="xUrWcgSdSqRE" outputId="6531f94a-272e-4a32-8999-a9160b7b83a3"
a=[1]*Xtrain.shape[0]
Xtrain["demo"]=a
Xtrain.shape
# + colab={"base_uri": "https://localhost:8080/"} id="2No_89QbSzTc" outputId="00bb41d3-b678-459f-ce97-2d478192330b"
b=[1]*Xtest.shape[0]
Xtest["demo"]=b
Xtest.shape
# + id="m9axD1MvS1yT"
x_train=Xtrain.values.reshape(Xtrain.shape[0],19,6,1)
x_test=Xtest.values.reshape(Xtest.shape[0],19,6,1)
# + colab={"base_uri": "https://localhost:8080/"} id="_zSD1oBQS6XM" outputId="9654cea4-5112-43a0-cb54-2a0d3cfdf17b"
x_train.shape
# + colab={"base_uri": "https://localhost:8080/"} id="Y0uPNMdfTBBU" outputId="08cc34ae-0c3e-4d35-a665-25dbc04cd147"
x_test.shape
# + id="7X4hr8JJTDIx"
import keras
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten
from keras.layers import Conv2D, MaxPooling2D
# + id="B4Tf7cUBULlN"
model=Sequential()
model.add(Conv2D(32,kernel_size=(3,3),activation='relu',input_shape=(19,6,1)))
model.add(Conv2D(64,(3,3),activation='relu'))
model.add(MaxPooling2D(pool_size=(2,2)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(128,activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(1,activation='sigmoid'))
# + id="zvIt9G9pUVg9"
model.compile(loss=keras.losses.binary_crossentropy,optimizer=keras.optimizers.Adadelta(),metrics=['accuracy'])
# + colab={"base_uri": "https://localhost:8080/"} id="ad3X0mg7Uaz-" outputId="3a23328c-ee16-4be4-c438-f939d5c407b2"
history = model.fit(x_train,Ytrain,batch_size=128,epochs=150,validation_data=(x_test,Ytest))
score=model.evaluate(x_test,Ytest,verbose=0)
print(score)
# + colab={"base_uri": "https://localhost:8080/", "height": 579} id="1icvcE7QUyIt" outputId="81bb2795-8713-4311-963c-7b5a1d555168"
# %matplotlib inline
# summarize history for accuracy
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
# summarize history for loss
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper right')
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="_VuOcTFWVCv4" outputId="0d95016b-800d-45a6-c7c7-792492056b56"
print("f1_score:",f1_score(Ytest,model.predict_classes(x_test),))
print("recall:",recall_score(Ytest,model.predict_classes(x_test),))
print("Validation Loss:",score[0])
print("Validation Accuracy:",score[1])
# + id="lzIPLSMcV1KX"
model.save('/content/sample_data/model2.h5')
# + id="PQZ08JSQXduY"
| Untitled26.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/HarshZ26/Object-Detection/blob/master/Raccoon_ML.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="AZ4e08MTyhCa" outputId="166225ba-8672-4e77-c660-4730c2361d18"
# !ls
# ! git clone "https://github.com/datitran/raccoon_dataset.git"
# + id="Oc8DWeCc4P8C" colab={"base_uri": "https://localhost:8080/"} outputId="a3a8e21f-eaaf-453d-e025-779d938eaa0a"
# %matplotlib inline
import os
import glob
import pandas as pd
import xml.etree.ElementTree as ET
import csv
import matplotlib.pyplot as plt
import torch
import torch.utils.data
import torchvision
import numpy as np
from PIL import Image
import torchvision.transforms as transforms
from random import randrange
import torch.nn as nn
import torch.nn.functional as F
import imutils
import cv2
def xml_to_csv(path):
xml_list = []
for xml_file in glob.glob(path + '/*.xml'):
tree = ET.parse(xml_file)
root = tree.getroot()
for member in root.findall('object'):
value = (root.find('filename').text,
int(root.find('size')[0].text),
int(root.find('size')[1].text),
member[0].text,
int(member[4][0].text),
int(member[4][1].text),
int(member[4][2].text),
int(member[4][3].text)
)
xml_list.append(value)
column_name = ['filename', 'width', 'height', 'class', 'xmin', 'ymin', 'xmax', 'ymax']
xml_df = pd.DataFrame(xml_list, columns=column_name)
return xml_df
def main():
image_path = '/content/raccoon_dataset/annotations'
xml_df = xml_to_csv(image_path)
xml_df.to_csv('raccoon_labels.csv', index=None)
print(xml_df)
print('Successfully converted xml to csv.')
main()
# + colab={"base_uri": "https://localhost:8080/"} id="MB9JiM3j6y3s" outputId="1cd75f6f-b474-48bb-b506-bf1e5b436c51"
# opening the CSV file
with open('/content/raccoon_labels.csv', mode ='r')as file:
# reading the CSV file
csvFile = csv.reader(file)
# displaying the contents of the CSV file
for lines in csvFile:
print(lines)
# + id="xB8LVmk67rnm"
class myOwnDataset(torch.utils.data.Dataset):
def __init__(self, root_img, root_label, transforms=None,id = None):
self.root_img = root_img
self.root_label = root_label
# the list of filename
self.transforms = transforms
self.file = open(root_label)
self.reader = csv.reader(self.file)
temp_lis1 = list(self.reader)
self.lis = temp_lis1[1:] + temp_lis1[1:90]
self.id = id
# the list of label
def __getitem__(self, index):
# obtain filenames from list
image_filename = self.lis[index][0]
# Load data and label
image = Image.open(os.path.join(self.root_img, image_filename),'r')
# print(type(image))
width,height,cls,xmin,ymin,xmax,ymax = self.lis[index][1:]
if self.id==1:
if index >170:
cls=0
im1 = image.crop((0,0,64,64))
else:
cls = 1
im1 = image.crop((int(xmin),int(ymin), int(xmax), int(ymax)))
if self.id==0:
if index >44:
cls=0
im1 = image.crop((0,0,64,64))
else:
cls = 1
im1 = image.crop((int(xmin),int(ymin), int(xmax), int(ymax)))
temp_var1 ,temp_var2 = image.size
im1 = im1.resize((256, 256))
mulx = 256/temp_var1
muly = 256/temp_var2
box = [[round(int(xmin)*mulx),round(int(ymin)*muly), round(int(xmax)*mulx), round(int(ymax)*muly)]]
# Bounding boxes for objects
# In pytorch, the input should be [xmin, ymin, xmax, ymax]
cls = torch.tensor(int(cls), dtype=torch.int64)
box = torch.tensor(box).squeeze()
# Size of bbox (Rectangular)
if self.transforms is not None:
image = self.transforms(im1)
return image,cls,box
def __len__(self):
return len(self.lis)
# + id="VT5o16LU8L67"
def get_transform():
custom_transforms = []
custom_transforms.append(torchvision.transforms.ToTensor())
return torchvision.transforms.Compose(custom_transforms)
# + colab={"base_uri": "https://localhost:8080/"} id="YVkX5MzMkzZu" outputId="03cce5f5-96f2-4fd7-9c4f-a934e96ea2c5"
# to remove grayscale images from directory and updating the labels
path_img = '/content/raccoon_dataset/images'
path_label = '/content/raccoon_dataset/data/train_labels.csv'
df = pd.read_csv("/content/raccoon_dataset/data/train_labels.csv")
# updating the column value/data
print(df.loc[df['filename']=='raccoon-161.jpg'])
print(df.loc[df['filename']=='raccoon-150.jpg'])
print(df.loc[df['filename']=='raccoon-152.jpg'])
df.drop(index = 124,inplace = True)
df.drop(index = 135,inplace = True)
df.drop(index = 172,inplace = True)
df.to_csv("AllDetails.csv", index=False)
# writing into the file
# + id="jQqLYk628dIS"
# path to your own data and csv file
train_data_dir = '/content/raccoon_dataset/images'
train_label = '/content/AllDetails.csv'
test_data_dir = '/content/raccoon_dataset/images'
test_label = '/content/raccoon_dataset/data/test_labels.csv'
batchsize = 32
# create own Dataset
train_dataset = myOwnDataset(root_img = train_data_dir,root_label = train_label ,
transforms = get_transform(),id = 1
)
test_dataset = myOwnDataset(root_img = test_data_dir,root_label = test_label ,
transforms = get_transform(),id = 0
)
def collate_fn(batch):
data = [item[0] for item in batch]
target = [item[1] for item in batch]
box = [item[2] for item in batch]
target = torch.LongTensor(target)
box = torch.stack(box,dim=0)
data = torch.stack(data,dim=0)
return [data, target,box]
# own DataLoader
train_loader = torch.utils.data.DataLoader(train_dataset,
batch_size=batchsize,
shuffle=True,
collate_fn = collate_fn)
test_loader = torch.utils.data.DataLoader(test_dataset,
batch_size=batchsize,
shuffle=True,
collate_fn = collate_fn)
# + colab={"base_uri": "https://localhost:8080/", "height": 338} id="FQ0HRK6KETmm" outputId="ac1b3f7e-304d-4da3-f13b-cad731dc4fb1"
train_features, train_labels,train_box= next(iter(test_loader))
print(f"Feature batch shape: {train_features.size()}")
print(f"Labels batch shape: {len(train_labels)}")
print('box',train_box.size())
img = train_features[0].squeeze()
img1 = train_features[0].squeeze()
img = torchvision.transforms.functional.convert_image_dtype(image= img,dtype=torch.uint8)
img = img.numpy()
im2display = img.transpose((1,2,0))
plt.imshow(im2display, interpolation='nearest')
# def imshow(img):
# npimg = img.numpy()
# plt.imshow(np.transpose(npimg, (1, 2, 0)))
# plt.show()
# #Get some random training images
# dataiter = iter(train_loader)
# images, labels,box = dataiter.next()
# #Show images
# imshow(torchvision.utils.make_grid(images))
# + id="ukVRJEessH39"
num_epochs = 20
num_classes = 2
batch_size = 32
learning_rate = 0.001
class ConvNet(nn.Module):
def __init__ (self):
super(ConvNet,self).__init__()
self.features = nn.Sequential(
nn.Conv2d(3,32, kernel_size =11, stride=1, padding =2),
nn.ReLU(),
nn.MaxPool2d(kernel_size =4,stride = 2),
nn.Conv2d(32, 64, kernel_size=5, stride=1, padding=2),
nn.ReLU(),
nn.MaxPool2d(kernel_size=4, stride=2),
nn.Conv2d(64, 128, kernel_size=3, stride=1, padding=2),
nn.ReLU(),
nn.MaxPool2d(kernel_size=3, stride=2),
nn.Conv2d(128, 256, kernel_size=3, stride=1, padding=1),
nn.ReLU(),
nn.MaxPool2d(kernel_size=3, stride=2))
self.classifier = nn.Sequential(
nn.Conv2d(256,128, kernel_size=15, stride=1, padding=0),
nn.ReLU(),
nn.Conv2d(128,64,kernel_size = 1,stride = 1,padding = 0),
nn.ReLU(),
nn.Conv2d(64,num_classes,kernel_size = 1,stride = 1,padding = 0)
)
self.regressor = nn.Sequential(
nn.Conv2d(256,128, kernel_size=15, stride=1, padding=0, bias=False),
nn.ReLU(),
nn.Conv2d(128,64,kernel_size = 1,stride = 1,padding = 0, bias=False),
nn.ReLU(),
nn.Conv2d(64,4,kernel_size = 1,stride = 1,padding = 0,bias=False)
)
def forward(self,x):
out = self.features(x)
out1 = self.classifier(out)
out2 = self.regressor(out)
return out1,out2
# + id="1E8DQw-xsMUE" colab={"base_uri": "https://localhost:8080/"} outputId="a2cbaeda-a9d6-481e-d04e-9fdbbf62401f"
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
model = ConvNet().to(device)
criterion1 = nn.CrossEntropyLoss()
criterion2 = nn.MSELoss()
optimizer = torch.optim.Adam(model.parameters(),lr = learning_rate)
model.train()
# + id="JSYxX4kBsZur" colab={"base_uri": "https://localhost:8080/"} outputId="4b11cd0a-480d-4075-df86-b1cec6ce05e9"
#train featurizer and classifier
total_step = len(train_loader)
correct_epoch = []
loss_lis1 = []
acc_lis = []
for epoch in range(num_epochs):
num_correct = 0
num_total = 0
for i,(images,labels,box) in enumerate(train_loader):
images, labels,box = images.to(device), labels.to(device), box.to(device)
#Run the forward pass
outputs,_= model(images)
outputs = outputs.reshape(outputs.size(0),-1)
#print("labels",labels)
#print(outputs.size(),labels)
loss = criterion1(outputs,labels)
loss_lis1.append(loss.item())
#Backprop
optimizer.zero_grad()
loss.backward()
optimizer.step()
#track accuracy
total = labels.size(0)
_ , predicted = torch.max(outputs.data,1)
correct = (predicted ==labels).sum().item()
acc_lis.append(correct/total)
num_correct += correct
num_total += total
if (i+1)%4 == 0:
print('Epoch [{}/{}],Step [{}/{}], Loss {: .4f}, Accuracy: {:.2f}%'
.format(epoch + 1, num_epochs, i + 1, total_step, loss.item(),
(correct / total) * 100))
correct_epoch.append((num_correct/num_total)*100)
# + colab={"base_uri": "https://localhost:8080/", "height": 637} id="XjsUnyBCvjOh" outputId="f95b56af-5c99-4da0-e98b-dda74201c38c"
# testing the trained classifier
model.eval()
with torch.no_grad():
correct = 0
total = 0
for images, labels,_ in test_loader:
images, labels = images.to(device), labels.to(device)
outputs,_ = model(images)
outputs = outputs.reshape(outputs.size(0),-1)
print(outputs[0])
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Test Accuracy of the model test images: {} %'.format((correct / total) * 100))
print('correct: '+str(correct)+'\t'+ 'total:'+str(total))
print(correct_epoch)
plt.plot(loss_lis1)
plt.show()
plt.plot(acc_lis)
plt.show()
# + id="6AcbgFWbv0LD"
# another dataset to train regressor
class myOwnDataset(torch.utils.data.Dataset):
def __init__(self, root_img, root_label, transforms=None):
self.root_img = root_img
self.root_label = root_label
# the list of filename
self.transforms = transforms
self.file = open(root_label)
self.reader = csv.reader(self.file)
temp_lis1 = list(self.reader)
self.lis = temp_lis1[1:]
# the list of label
def __getitem__(self, index):
# obtain filenames from list
image_filename = self.lis[index][0]
# Load data and label
image = Image.open(os.path.join(self.root_img, image_filename),'r')
# print(type(image))
width,height,cls,xmin,ymin,xmax,ymax = self.lis[index][1:]
cls = 1
temp_var1 ,temp_var2 = image.size
im1 = image.resize((256, 256))
mulx = 256/temp_var1
muly = 256/temp_var2
box = [[round(int(xmin)*mulx),round(int(ymin)*muly), round(int(xmax)*mulx), round(int(ymax)*muly)]]
box = torch.tensor(box).squeeze()
# Bounding boxes for objects
# In pytorch, the input should be [xmin, ymin, xmax, ymax]
cls = torch.tensor(int(cls), dtype=torch.int64)
if self.transforms is not None:
image = self.transforms(im1)
return image,cls,box
def __len__(self):
return len(self.lis)
# + id="y6OUWsQ1v0_Q"
valid_data_dir = '/content/raccoon_dataset/images'
valid_label = '/content/AllDetails.csv'
batchsize = 32
# create own Dataset
valid_dataset = myOwnDataset(root_img = valid_data_dir,root_label = valid_label,
transforms = get_transform()
)
def collate_fn(batch):
data = [item[0] for item in batch]
target = [item[1] for item in batch]
box = [item[2] for item in batch]
target = torch.LongTensor(target)
box = torch.stack(box,dim=0)
data = torch.stack(data,dim=0)
return [data, target,box]
# own DataLoader
valid_loader = torch.utils.data.DataLoader(valid_dataset,
batch_size=batchsize,
shuffle=True,
collate_fn = collate_fn)
# + colab={"base_uri": "https://localhost:8080/", "height": 356} id="y-Jqhms9VA9M" outputId="26f3086c-a4a7-4b0f-a787-bffd92eafaa0"
valid_features, valid_labels,bbox= next(iter(valid_loader))
print(f"Feature batch shape: {valid_features.size()}")
print('box',bbox[0])
# print(f"Labels batch shape: {len(train_labels)}")
img = valid_features[0].squeeze()
img2 = valid_features
print(img2.size())
img1 = valid_features
# img1 = torchvision.transforms.functional.convert_image_dtype(image= img2,dtype=torch.uint8)
img1 = img1.numpy()
img = torchvision.transforms.functional.convert_image_dtype(image= img,dtype=torch.uint8)
img = img.numpy()
im2display = img.transpose((1,2,0))
plt.imshow(im2display, interpolation='nearest')
print(valid_labels)
# + id="yGRCcWoHXjgC" colab={"base_uri": "https://localhost:8080/"} outputId="d5074314-7788-4793-ac27-2727df2f5e8f"
# freezing featurizer and classifier parameters
learning_rate = 0.005
params = model.state_dict()
key = list(params.keys())
model.features[0].weight.requires_grad = False
model.features[0].bias.requires_grad = False
model.features[3].weight.requires_grad = False
model.features[3].bias.requires_grad = False
model.features[6].weight.requires_grad = False
model.features[6].bias.requires_grad = False
model.features[9].weight.requires_grad = False
model.features[9].bias.requires_grad = False
model.classifier[0].weight.requires_grad = False
model.classifier[0].bias.requires_grad = False
model.classifier[2].weight.requires_grad = False
model.classifier[2].bias.requires_grad = False
model.classifier[4].weight.requires_grad = False
model.classifier[4].bias.requires_grad = False
# model.features.requires_grad = True
# model.classifier.requires_grad = True
for name, param in model.named_parameters():
if param.requires_grad:print(name)
optimizer2 = torch.optim.Adam(filter(lambda p: p.requires_grad, model.parameters()),lr = learning_rate)
model.train()
# + colab={"base_uri": "https://localhost:8080/"} id="P17UEjECXBL1" outputId="d924443d-1858-49a2-cebe-793fee8ccdd2"
#train regressor
total_step = len(valid_loader)
num_epochs = 100
correct_epoch = []
loss_lis2 = []
for epoch in range(num_epochs):
num_correct = 0
num_total = 0
for i,(images,_,box) in enumerate(valid_loader):
images,box = images.to(device),box.to(device,dtype=torch.float32)
#Run the forward pass
_,outputs= model(images)
outputs = outputs.reshape(outputs.size(0),-1)
#print("labels",labels)
#print(outputs.size(),labels)
loss = criterion2(outputs,box)
loss_lis2.append(loss.item())
#Backprop
optimizer2.zero_grad()
loss.backward()
optimizer2.step()
if (i+1)%4 == 0:
print('Epoch [{}/{}],Step [{}/{}], Loss {: .4f}'.format(epoch + 1, num_epochs, i + 1, total_step, loss.item()))
# + colab={"base_uri": "https://localhost:8080/", "height": 318} id="v0IHFSUBrn4T" outputId="f80a97d8-377e-4fcb-d85c-c438a4813fa2"
# testing regressor
model.eval()
with torch.no_grad():
correct = 0
total = 0
for images,_,box in test_loader:
images ,box= images.to(device), box.to(device)
_,outputs = model(images)
outputs = outputs.reshape(outputs.size(0),-1)
print(outputs[0],box[0])
plt.plot(loss_lis2)
plt.show()
# + id="g_BnOm1E_aOC"
# for large images
class myOwnDataset(torch.utils.data.Dataset):
def __init__(self, root_img, root_label, transforms=None):
self.root_img = root_img
self.root_label = root_label
# the list of filename
self.transforms = transforms
self.file = open(root_label)
self.reader = csv.reader(self.file)
temp_lis1 = list(self.reader)
self.lis = temp_lis1[1:]
# the list of label
def __getitem__(self, index):
# obtain filenames from list
image_filename = self.lis[index][0]
# Load data and label
image = Image.open(os.path.join(self.root_img, image_filename),'r')
# print(type(image))
width,height,cls,xmin,ymin,xmax,ymax = self.lis[index][1:]
cls = 1
temp_var1 ,temp_var2 = image.size
im1 = image.resize((512, 512))
mulx = 512/temp_var1
muly = 512/temp_var2
box = [[round(int(xmin)*mulx),round(int(ymin)*muly), round(int(xmax)*mulx), round(int(ymax)*muly)]]
box = torch.tensor(box).squeeze()
# number of objects in the image
# Bounding boxes for objects
cls = torch.tensor(int(cls), dtype=torch.int64)
if self.transforms is not None:
image = self.transforms(im1)
return image,cls,box
def __len__(self):
return len(self.lis)
# + id="QaAjLRZ03qdJ"
valid_data_dir = '/content/raccoon_dataset/images'
valid_label = '/content/raccoon_dataset/data/test_labels.csv'
batchsize = 1
# create own Dataset
valid_dataset = myOwnDataset(root_img = valid_data_dir,root_label = valid_label,
transforms = get_transform()
)
def collate_fn(batch):
data = [item[0] for item in batch]
target = [item[1] for item in batch]
box = [item[2] for item in batch]
target = torch.LongTensor(target)
box = torch.stack(box,dim=0)
data = torch.stack(data,dim=0)
return [data, target,box]
# own DataLoader
valid_loader = torch.utils.data.DataLoader(valid_dataset,
batch_size=batchsize,
shuffle=True,
collate_fn = collate_fn)
# + id="I5Ru7cRVv9_E"
def pyramid(image,scale,miniSize = (256,256)):
yield image
while True:
w = int(image.shape[1]/scale)
image = imutils.resize(image,width = w)
if image.shape[0] < miniSize[1] or image.shape[1] < miniSize[0]:
break
yield image
def sliding_window(image,stepSize,windowSize):
t_img = image.transpose((2,0,1))
test_img = np.expand_dims(t_img, axis=0)
test_img = torch.tensor(test_img)
return mod(test_img)
# yield (x,y,image[y:y+ windowSize[1],x:x + windowSize[0]])
def mod(ig):
model.eval()
with torch.no_grad():
ig = ig.to(device)
outputs,bux = model(ig)
print('result',outputs.size())
probabilities = torch.nn.functional.softmax(outputs[0], dim=0)
bux = bux.cpu()
bux = bux.numpy()
result = probabilities.cpu()
result = result.numpy()
print(np.shape(result),np.shape(bux[0]))
return result,bux[0]
# + colab={"base_uri": "https://localhost:8080/", "height": 338} id="QXNTjFr24KJG" outputId="4abd0728-3932-41e1-a13c-20ab64b45bb7"
valid_features, valid_labels,bbox= next(iter(valid_loader))
print(f"Feature batch shape: {valid_features.size()}")
print('box',bbox[0])
# print(f"Labels batch shape: {len(train_labels)}")
img = valid_features[0].squeeze()
img2 = valid_features
print(img2.size())
img1 = valid_features
# img1 = torchvision.transforms.functional.convert_image_dtype(image= img2,dtype=torch.uint8)
img1 = img1.numpy()
img = torchvision.transforms.functional.convert_image_dtype(image= img,dtype=torch.uint8)
img = img.numpy()
im2display = img.transpose((1,2,0))
plt.imshow(im2display, interpolation='nearest')
print(valid_labels)
# + id="oq6qV2HKVXK5" colab={"base_uri": "https://localhost:8080/", "height": 478} outputId="9c5f840b-1f81-49fe-f8a7-d457cc763c6c"
print(np.shape(img1))
image = img1[0].transpose((1,2,0))
print(np.shape(image))
print(bbox)
winH = 256
winW = 256
cord_lis = []
font = cv2.FONT_HERSHEY_SIMPLEX
imgs = image.copy()
m = 1
for resized in pyramid(image, scale=1.2):
blocks,cords = sliding_window(resized,1,(256,256))
lengt = np.shape(blocks)[1]
for y in range(lengt):
for x in range(lengt):
if blocks[1][x][y]>0.999976 and m>1.5 :
tx =8*x*m
ty = 8*y*m
cord_lis.append((round(cords[0][x][y]*m + tx),round(cords[1][x][y]*m+ ty),round(cords[2][x][y]*m + tx),round(cords[3][x][y]*m+ ty)))
# cv2.rectangle(imgs, (8*x, 8*y), (8*x + winW, 8*y + winH), (255, 0, 0), 2)
# cv2.putText(imgs, "Raccoon", (8*x,8*y), font, 1, (255,0,0), 3, cv2.LINE_AA)
m = m*1.2
# storing coordinates into list and plotting them at end
cv2.rectangle(imgs, (bbox[0][0], bbox[0][1]), (bbox[0][2], bbox[0][3]), (0, 255, 0), 2)
for item in cord_lis:
cv2.rectangle(imgs, (item[0], item[1]), (item[2], item[3]), (255, 0, 0), 3)
plt.imshow(imgs)
plt.show()
# t_img = image[y:y+winH,x:x+winW]
# # t_img = t_img.transpose((2,0,1))
# # print("test",np.shape(t_img))
# test_img = np.expand_dims(t_img, axis=0)
# # print("test",np.shape(test_img))
# test_img = torch.tensor(test_img)
# tet = Image.fromarray(t_img,'RGB')
# result = foo(tet)
# result = result.cpu()
# result = result.numpy()
# # with open("imagenet_classes.txt", "r") as f:
# # categories = [s.strip() for s in f.readlines()]
# # Show top categories per image
# font = cv2.FONT_HERSHEY_SIMPLEX
# txt = cls_lis[np.argmax(result)]
# cv2.rectangle(clone, (x, y), (x + winW, y + winH), (255, 0, 0), 2)
# cv2.rectangle(clone, (bbox[0][0], bbox[0][1]), (bbox[0][2], bbox[0][3]), (0, 255, 0), 2)
# cv2.putText(clone, txt, (x,y), font, 1, (255,0,0), 3, cv2.LINE_AA)
# plt.imshow(clone)
# plt.show()
# plt.clf()
| Overfeat/Raccoon_ML.ipynb |